code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (PyTorch 1.8 Python 3.6 GPU Optimized)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:ap-southeast-2:452832661640:image/pytorch-1.8-gpu-py36
# ---
# + [markdown] id="SKQ4bH7qMGrA"
# # Check Colab Resources
#
# > Mainly use this to check on Colab environment and it also can be used in others.
#
# + [markdown] id="QMMqmdiYMkvi"
# ## What's the GPU Type
#
# With diff Colab subscription, you may get different GPU type: such as T4, P100, or even slower K80. It's good to understand your GPU capacities before running Deep Learning model training.
#
# If the execution result of running the code cell below is 'Not connected to a GPU', you can change the runtime by going to Runtime > Change runtime type in the menu to enable a GPU accelerator, and then re-execute the code cell.
# + id="23TOba33L4qf"
# gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
if gpu_info.find('failed') >= 0:
print('Not connected to a GPU')
else:
print(gpu_info)
# + [markdown] id="Sa-IrJS1aRVJ"
# In order to use a GPU with your notebook, select the Runtime > Change runtime type menu, and then set the hardware accelerator drop-down to GPU.
# + [markdown] id="65MSuHKqNeBZ"
# ## More memory
#
# <p>With Colab Pro, you have the option to access high-memory VMs when they are available, and with Pro+ even more so. To set your notebook preference to use a high-memory runtime, select the runtime > 'Change runtime type' menu, and then select High-RAM in the runtime shape drop-down.</p>
# <p>You can see how much memory you have available at any time by running the following code cell.</p>
# If the execution result of running the code cell below is 'Not using a high-RAM runtime', then you can enable a high-RAM runtime via Runtime > Change runtime type in the menu. Then select High-RAM in the Runtime shape drop-down. After, re-execute the code cell.
# + id="V1G82GuO-tez"
from psutil import virtual_memory
ram_gb = virtual_memory().total / 1e9
print('Your runtime has {:.1f} gigabytes of available RAM\n'.format(ram_gb))
if ram_gb < 20:
print('Not using a high-RAM runtime')
else:
print('You are using a high-RAM runtime!')
# -
| notebooks/colab_resource_check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %run ../../main.py
# %matplotlib inline
# +
import pandas as pd
from cba.algorithms import M1Algorithm, M2Algorithm, top_rules, createCARs
from cba.data_structures import TransactionDB
# +
#
#
# =========================
# Oveření běhu v závislosti na vložených pravidlech / instancích
# =========================
#
#
#
import time
rule_count = 100
benchmark_data = {
"input rows": [],
"input rules": [],
"output rules M1 pyARC": [],
"output rules M1 pyARC unique": [],
"output rules M2 pyARC": [],
"time M1 pyARC": [],
"time M1 pyARC unique": [],
"time M2 pyARC": []
}
stop_m2 = False
number_of_iterations = 30
directory = "c:/code/python/machine_learning/assoc_rules"
dataset_name_benchmark = "lymph0"
pd_ds = pd.read_csv("c:/code/python/machine_learning/assoc_rules/train/{}.csv".format(dataset_name_benchmark))
for i in range(11):
dataset_name_benchmark = "lymph0"
pd_ds = pd.concat([pd_ds, pd_ds])
txns = TransactionDB.from_DataFrame(pd_ds, unique_transactions=True)
txns_unique = TransactionDB.from_DataFrame(pd_ds, unique_transactions=False)
rules = top_rules(txns.string_representation, appearance=txns.appeardict, target_rule_count=rule_count)
cars = createCARs(rules)
if len(cars) > rule_count:
cars = cars[:rule_count]
m1t1 = time.time()
m1clf_len = []
for _ in range(number_of_iterations):
m1 = M1Algorithm(cars, txns)
clf = m1.build()
m1clf_len.append(len(clf.rules) + 1)
m1t2 = time.time()
m1t1_unique = time.time()
m1clf_len_unique = []
for _ in range(number_of_iterations):
m1 = M1Algorithm(cars, txns_unique)
clf = m1.build()
m1clf_len_unique.append(len(clf.rules) + 1)
m1t2_unique = time.time()
if not stop_m2:
m2t1 = time.time()
m2clf_len = []
for _ in range(number_of_iterations):
m2 = M2Algorithm(cars, txns)
clf = m2.build()
m2clf_len.append(len(clf.rules) + 1)
m2t2 = time.time()
m1duration = (m1t2 - m1t1) / number_of_iterations
m1duration_unique = (m1t2_unique - m1t1_unique) / number_of_iterations
outputrules_m1 = sum(m1clf_len) / len(m1clf_len)
outputrules_m1_unique = sum(m1clf_len_unique) / len(m1clf_len_unique)
if not stop_m2:
m2duration = (m2t2 - m2t1) / number_of_iterations
outputrules_m2 = sum(m2clf_len) / len(m2clf_len)
if m2duration > 0.5:
stop_m2 = True
benchmark_data["input rows"].append(len(txns))
benchmark_data["input rules"].append(rule_count)
benchmark_data["output rules M1 pyARC"].append(outputrules_m1)
benchmark_data["output rules M1 pyARC unique"].append(outputrules_m1_unique)
benchmark_data["output rules M2 pyARC"].append(None if stop_m2 else outputrules_m2)
benchmark_data["time M1 pyARC"].append(m1duration)
benchmark_data["time M1 pyARC unique"].append(m1duration_unique)
benchmark_data["time M2 pyARC"].append(None if stop_m2 else m2duration)
print("data_count:", len(txns))
print("M1 duration:", m1duration)
print("M1 unique duration", m1duration_unique)
print("M1 output rules", outputrules_m1)
if not stop_m2:
print("M2 duration:", m2duration)
print("M2 output rules", outputrules_m2)
print("\n\n")
# +
#benchmark_data.pop("M2_duration")
benchmark_df = pd.DataFrame(benchmark_data)
benchmark_df.plot(x=["input rows"], y=["time M1 pyARC", "time M2 pyARC"])
#benchmark_df.to_csv("../data/data_sensitivity.csv")
# -
benchmark_df
# +
R_benchmark = pd.read_csv("../data/arc-data-size.csv")
R_benchmark[["input rows"]] = R_benchmark[["input rows"]].astype(str)
R_benchmark.set_index("input rows", inplace=True)
# -
R_benchmark.head()
benchmark_df[["input rows"]] = benchmark_df[["input rows"]].astype(str)
benchmark_df = benchmark_df.set_index("input rows")
benchmark_all = benchmark_df.join(R_benchmark, lsuffix="_py", rsuffix="_R")
benchmark_all
# +
import matplotlib.pyplot as plt
labels = ["pyARC - m1", "pyARC - m2", "arc", "rCBA", "arulesCBA"]
ax = benchmark_all.plot(y=["time M1 pyARC", "time M2 pyARC", "time_arc", "time_acba", "time_rcba"])
ax.legend(labels)
plt.savefig("../data/data_size_sensitivity_plot.png")
# -
benchmark_all.plot(y=["time M1 pyARC", "time M2 pyARC", "time_arc", "time_acba", "time_rcba"])
| notebooks/benchmark/speed_benchmark_datacase_count.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # First ML Pipeline
#
# Analysis by <NAME>
#
# Just a first pass modeling of the data using the cumulant feature engineering. The models tested were:
#
# 1. Linear SVC
# 1. Random Forest
# 1. Gradient Boosted Classifier
# 1. rbfSVC
#
# All models were chosen with their default parameters. Evaluations were just the confusion matrix plots and classfication reports built into sklearn. This was not performed through cross-validation
#
# ## Primary Insights
#
# 1. Ensemble trees performed the best, in particular the gradient boosted ones.
# 1. The models' performance was not uniform along classes.
# 1. The adipose and background tissues were easily classified.
# 1. The rest of the classes were classifed with ~70 accuracy
# 1. Considering the "Multi-Texture.." paper, I was surprised by the relatively poor performance of the rbfSVC.
# 1. I suspect this is due to poor hyperparameter selection due to my lack of scaling? If I recall correctly, they didn't say anything preprocessing/hyperparameter selection in their paper.
# 1. It's also possible that my choice of cumulants in lieu of moments was mistaken?
#
# 1. My wrangle was a bit off, I'll have to go back and rewrite a lot of the code
# +
import xarray as xr
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.base import BaseEstimator, TransformerMixin
from scipy.stats import kstat
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split
from sklearn.metrics import (
accuracy_score,
classification_report,
confusion_matrix,
plot_confusion_matrix,
)
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingClassifier, RandomForestClassifier
from sklearn.svm import LinearSVC, SVC
# +
def load_X_train(filepath):
X = xr.open_dataarray(filepath).values
X = X.reshape(X.shape[0], -1)
return X
class cumulants_extractor(BaseEstimator, TransformerMixin):
'''
returns a numpy array of all k-th cumulants less than
highest_cumulant (which must be less than 4)
'''
def __init__(self, highest_cumulant):
self.highest_cumulant = highest_cumulant
def fit(self, X, y = None):
return self
def get_cumulants(self, v):
kstats = np.array([kstat(data = v, n = k)
for k in range(1, self.highest_cumulant + 1)])
return kstats
def transform(self, X):
# X =
cumulants = np.apply_along_axis(func1d = self.get_cumulants,
axis = 1,
arr = X,
)
return cumulants
# +
yfilepath = "../../data/clean_data/train_data/y_64_L_clean_train.nc"
y = xr.open_dataarray(yfilepath).values
y = np.arange(1, 9) * y
y = y.sum(axis = 1)
Xfilepath = "../../data/clean_data/train_data/X_64_L_clean_train.nc"
X = load_netcdf(Xfilepath)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.20, random_state=42
)
# +
class_names = [
"Tumor",
"Stroma",
"Complex",
"Lympho",
"Debris",
"Mucosa",
"Adipose",
"Empty",
]
models = [
('Linear SVC', LinearSVC(max_iter = 10**5, dual=False)),
("Gradient-Boosted DTs", GradientBoostingClassifier(random_state = 42)),
("Random Forest", RandomForestClassifier(random_state = 42)),
("rbfSVC", SVC(kernel="rbf")),
]
pipelines = [
Pipeline([("cumulant extractor", cumulants_extractor(4)), model])
for model in models
]
# +
def plot_confusion_matrices(pipelines, X_train, y_train, X_test, y_test):
n_of_models = len(models)
fig, ax = plt.subplots(1, n_of_models, figsize=(30, 8))
fig.suptitle("Confusion Matrices of Models", fontsize=30)
for i, pipeline in enumerate(pipelines):
pipeline.fit(X_train, y_train)
plot_confusion_matrix(
pipeline,
X_test,
y_test,
display_labels=class_names,
normalize="true",
xticks_rotation="vertical",
ax=ax[i],
)
ax[i].set(title=pipeline.steps[-1][0])
plt.show()
plot_confusion_matrices(pipeplines, X_train, y_train, X_test, y_test)
# -
model.steps[-1][0]
for pipeline in pipelines:
pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_test)
print("Performance of " + pipeline.steps[-1][0] + "\n")
print(
classification_report(
y_test,
y_pred,
labels=range(1, 9),
target_names=class_names,
zero_division=0,
)
)
| develop/2020-5-15-jpm-first-ml-pipeline/2020-5-15-jpm-first-ml-pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import numpy as np
import pandas as pd
from sklearn import metrics
from sklearn.preprocessing import LabelEncoder
from sklearn.cross_validation import train_test_split
from unidecode import unidecode
# +
permulaan = [
'bel',
'se',
'ter',
'men',
'meng',
'mem',
'memper',
'di',
'pe',
'me',
'ke',
'ber',
'pen',
'per',
]
hujung = ['kan', 'kah', 'lah', 'tah', 'nya', 'an', 'wan', 'wati', 'ita']
def naive_stemmer(word):
assert isinstance(word, str), 'input must be a string'
hujung_result = re.findall(r'^(.*?)(%s)$' % ('|'.join(hujung)), word)
word = hujung_result[0][0] if len(hujung_result) else word
permulaan_result = re.findall(r'^(.*?)(%s)' % ('|'.join(permulaan[::-1])), word)
permulaan_result.extend(re.findall(r'^(.*?)(%s)' % ('|'.join(permulaan)), word))
mula = permulaan_result if len(permulaan_result) else ''
if len(mula):
mula = mula[1][1] if len(mula[1][1]) > len(mula[0][1]) else mula[0][1]
return word.replace(mula, '')
def classification_textcleaning(string):
string = re.sub(
'http\S+|www.\S+',
'',
' '.join(
[i for i in string.split() if i.find('#') < 0 and i.find('@') < 0]
),
)
string = unidecode(string).replace('.', ' . ').replace(',', ' , ')
string = re.sub('[^A-Za-z ]+', ' ', string)
string = re.sub(r'[ ]+', ' ', string).strip()
string = ' '.join(
[i for i in re.findall('[\\w\']+|[;:\-\(\)&.,!?"]', string) if len(i)]
)
string = string.lower().split()
string = [(naive_stemmer(word), word) for word in string]
return (
' '.join([word[0] for word in string if len(word[0]) > 1]),
' '.join([word[1] for word in string if len(word[0]) > 1]),
)
# -
df = pd.read_csv('dataset/sentiment-data-v2.csv')
Y = LabelEncoder().fit_transform(df.label)
df.head()
# +
with open('dataset/polarity-negative-translated.txt','r') as fopen:
texts = fopen.read().split('\n')
labels = [0] * len(texts)
with open('dataset/polarity-positive-translated.txt','r') as fopen:
positive_texts = fopen.read().split('\n')
labels += [1] * len(positive_texts)
texts += positive_texts
texts += df.iloc[:,1].tolist()
labels += Y.tolist()
assert len(labels) == len(texts)
# -
for i in range(len(texts)):
texts[i] = classification_textcleaning(texts[i])[0]
from sklearn.feature_extraction.text import TfidfVectorizer
import xgboost as xgb
from malaya.text_functions import STOPWORDS
target = LabelEncoder().fit_transform(labels)
tfidf = TfidfVectorizer(ngram_range=(1, 3),min_df=2).fit(texts)
vectors = tfidf.transform(texts)
vectors.shape
train_X, test_X, train_Y, test_Y = train_test_split(vectors, target, test_size = 0.2)
train_d = xgb.DMatrix(train_X, train_Y)
test_d = xgb.DMatrix(test_X, test_Y)
params_xgd = {
'min_child_weight': 10.0,
'max_depth': 7,
'objective': 'multi:softprob',
'max_delta_step': 1.8,
'num_class': 2,
'colsample_bytree': 0.4,
'subsample': 0.8,
'learning_rate': 0.1,
'gamma': 0.65,
'silent': True,
'eval_metric': 'mlogloss'
}
model = xgb.train(params_xgd, train_d, 10000, evals=[(test_d, 'validation')],
early_stopping_rounds=100, verbose_eval=5)
predicted = np.argmax(model.predict(xgb.DMatrix(test_X),ntree_limit=model.best_ntree_limit),axis=1)
print(metrics.classification_report(test_Y, predicted, target_names = ['negative','positive']))
text = (
'kerajaan sebenarnya sangat bencikan rakyatnya, minyak naik dan segalanya'
)
model.predict(
xgb.DMatrix(tfidf.transform([classification_textcleaning(text)[0]])),
ntree_limit = model.best_ntree_limit,
)
# +
import pickle
with open('xgboost-sentiment.pkl','wb') as fopen:
pickle.dump(model,fopen)
with open('tfidf-xgboost-sentiment.pkl','wb') as fopen:
pickle.dump(tfidf,fopen)
# -
| session/sentiment/xgboost.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Anomaly Detection Tutorial
//
// This guide will show how to use Tribuo’s anomaly detection models to find anomalous events in a toy dataset drawn from a mixture of Gaussians. We'll discuss the options in the LibSVM anomaly detection algorithm (using a one-class nu-SVM) and discuss evaluations for anomaly detection tasks.
//
// ## Setup
//
// We'll load in a jar and import a few packages.
// %jars ./tribuo-anomaly-libsvm-4.0.0-jar-with-dependencies.jar
import org.tribuo.*;
import org.tribuo.util.Util;
import org.tribuo.anomaly.*;
import org.tribuo.anomaly.evaluation.*;
import org.tribuo.anomaly.example.AnomalyDataGenerator;
import org.tribuo.anomaly.libsvm.*;
import org.tribuo.common.libsvm.*;
var eval = new AnomalyEvaluator();
// ## Dataset
// Tribuo's anomaly detection package comes with a simple data generator that emits pairs of datasets containing 5 features. The training data is free from anomalies, and each example is sampled from a 5 dimensional gaussian with fixed mean and diagonal covariance. The test data is sampled from a mixture of two distributions, the first is the same as the training distribution, and the second uses a different mean for the gaussian (keeping the same covariance for simplicity). All the data points sampled from the second distribution are marked `ANOMALOUS`, and the other points are marked `EXPECTED`. These form the two classes for Tribuo's anomaly detection system. You can also use any of the standard data loaders to pull in anomaly detection data.
//
// The libsvm anomaly detection algorithm requires there are no anomalies in the training data, but this is not required in general for Tribuo's anomaly detection infrastructure.
//
// We'll sample 2000 points for each dataset, and 20% of the test set will be anomalies.
var pair = AnomalyDataGenerator.gaussianAnomaly(2000,0.2);
var data = pair.getA();
var test = pair.getB();
// ## Model Training
// We'll fit a one-class SVM to our training data, and then use that to determine what things in our test set are anomalous. We'll use an [RBF Kernel](https://en.wikipedia.org/wiki/Radial_basis_function_kernel), and set the kernel width to 1.0.
var params = new SVMParameters<>(new SVMAnomalyType(SVMAnomalyType.SVMMode.ONE_CLASS), KernelType.RBF);
params.setGamma(1.0);
params.setNu(0.1);
var trainer = new LibSVMAnomalyTrainer(params);
// Training is the same as other Tribuo prediction tasks, just call train and pass the training data.
var startTime = System.currentTimeMillis();
var model = trainer.train(data);
var endTime = System.currentTimeMillis();
System.out.println();
System.out.println("Training took " + Util.formatDuration(startTime,endTime));
// Unfortunately the LibSVM implementation is a little chatty and insists on writing to standard out, but after that we can see it took about 140ms to run (on my 2020 16" Macbook Pro, you may get slightly different runtimes). We can check how many support vectors are used by the SVM, from the training set of 2000:
((LibSVMAnomalyModel)model).getNumberOfSupportVectors()
// So we used 301 datapoints to model the density of the expected data.
// ## Model evaluation
// Tribuo's infrastructure treats anomaly detection as a binary classification problem with the fixed label set {`EXPECTED`,`ANOMALOUS`}. When we have ground truth labels we can thus measure the true positives (anomalous things predicted as anomalous), false positives (expected things predicted as anomalous), false negatives (anomalous things predicted as expected) and true negatives (expected things predicted as expected), though the latter number is not usually that useful. We can also calculate the usual summary statistics: precision, recall and F1 of the anomalous class. We're going to compare against the ground truth labels from the data generator.
var testEvaluation = eval.evaluate(model,test);
System.out.println(testEvaluation.toString());
System.out.println(testEvaluation.confusionString());
// We can see that the model has no false negatives, and so perfect recall, but has a precision of 0.62, so approximately 62% of the positive predictions are true anomalies. This can be tuned by changing the width of the gaussian kernel which changes the range of values which are considered to be expected. The confusion matrix presents the same results in a more common form for classification tasks.
//
// We plan to further expand Tribuo's anomaly detection functionality to incorporate other algorithms in the future.
| tutorials/anomaly-tribuo-v4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming and Database Fundamentals for Data Scientists - EAS503
# # Visualization with Seaborn
# Matplotlib has proven to be an incredibly useful and popular visualization tool, but even avid users will admit it often leaves much to be desired.
# There are several valid complaints about Matplotlib that often come up:
#
# - Prior to version 2.0, Matplotlib's defaults are not exactly the best choices. It was based off of MATLAB circa 1999, and this often shows.
# - Matplotlib's API is relatively low level. Doing sophisticated statistical visualization is possible, but often requires a *lot* of boilerplate code.
# - Matplotlib predated Pandas by more than a decade, and thus is not designed for use with Pandas ``DataFrame``s. In order to visualize data from a Pandas ``DataFrame``, you must extract each ``Series`` and often concatenate them together into the right format. It would be nicer to have a plotting library that can intelligently use the ``DataFrame`` labels in a plot.
#
# An answer to these problems is [Seaborn](http://seaborn.pydata.org/). Seaborn provides an API on top of Matplotlib that offers sane choices for plot style and color defaults, defines simple high-level functions for common statistical plot types, and integrates with the functionality provided by Pandas ``DataFrame``s.
#
#
# ## Seaborn Versus Matplotlib
#
# Here is an example of a simple random-walk plot in Matplotlib, using its classic plot formatting and colors.
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
# We create some random walk data:
# Create some data
rng = np.random.RandomState(0)
x = np.linspace(0, 10, 500)
y = np.cumsum(rng.randn(500, 6), 0)
# And do a simple plot:
# Plot the data with Matplotlib defaults
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# Although the result contains all the information we'd like it to convey, it does so in a way that is not all that aesthetically pleasing, and even looks a bit old-fashioned in the context of 21st-century data visualization.
#
# Now let's take a look at how it works with Seaborn.
# As we will see, Seaborn has many of its own high-level plotting routines, but it can also overwrite Matplotlib's default parameters and in turn get even simple Matplotlib scripts to produce vastly superior output.
# We can set the style by calling Seaborn's ``set()`` method.
# By convention, Seaborn is imported as ``sns``:
import seaborn as sns
sns.set()
# Now let's rerun the same two lines as before:
# same plotting code as above!
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# Ah, much better!
# ## Exploring Seaborn Plots
#
# The main idea of Seaborn is that it provides high-level commands to create a variety of plot types useful for statistical data exploration, and even some statistical model fitting.
#
# Let's take a look at a few of the datasets and plot types available in Seaborn. Note that all of the following *could* be done using raw Matplotlib commands (this is, in fact, what Seaborn does under the hood) but the Seaborn API is much more convenient.
# ### Histograms, KDE, and densities
#
# Often in statistical data visualization, all you want is to plot histograms and joint distributions of variables.
# We have seen that this is relatively straightforward in Matplotlib:
# +
data = np.random.multivariate_normal([0, 0], [[5, 2], [2, 2]], size=2000)
data = pd.DataFrame(data, columns=['x', 'y'])
for col in 'xy':
plt.hist(data[col], normed=True, alpha=0.5)
# -
# ### Pair plots
#
# When you generalize joint plots to datasets of larger dimensions, you end up with *pair plots*. This is very useful for exploring correlations between multidimensional data, when you'd like to plot all pairs of values against each other.
#
# We'll demo this with the well-known Iris dataset, which lists measurements of petals and sepals of three iris species:
iris = sns.load_dataset("iris")
iris.head()
# Visualizing the multidimensional relationships among the samples is as easy as calling ``sns.pairplot``:
sns.pairplot(iris, hue='species', size=2.5);
# ### Faceted histograms
#
# Sometimes the best way to view data is via histograms of subsets. Seaborn's ``FacetGrid`` makes this extremely simple.
# We'll take a look at some data that shows the amount that restaurant staff receive in tips based on various indicator data:
tips = sns.load_dataset('tips')
tips.head()
# +
tips['tip_pct'] = 100 * tips['tip'] / tips['total_bill']
grid = sns.FacetGrid(tips, row="sex", col="time", margin_titles=True)
grid.map(plt.hist, "tip_pct", bins=np.linspace(0, 40, 15));
# -
# ### Factor plots
#
# Factor plots can be useful for this kind of visualization as well. This allows you to view the distribution of a parameter within bins defined by any other parameter:
with sns.axes_style(style='ticks'):
g = sns.factorplot("day", "total_bill", "sex", data=tips, kind="box")
g.set_axis_labels("Day", "Total Bill");
# ### Joint distributions
#
# Similar to the pairplot we saw earlier, we can use ``sns.jointplot`` to show the joint distribution between different datasets, along with the associated marginal distributions:
with sns.axes_style('white'):
sns.jointplot("total_bill", "tip", data=tips, kind='hex')
# ### Bar plots
#
# Time series can be plotted using ``sns.factorplot``. In the following example, we'll use the Planets data that we first saw in [Aggregation and Grouping](03.08-Aggregation-and-Grouping.ipynb):
planets = sns.load_dataset('planets')
planets.head()
with sns.axes_style('white'):
g = sns.factorplot("year", data=planets, aspect=2,
kind="count", color='steelblue')
g.set_xticklabels(step=5)
# We can learn more by looking at the *method* of discovery of each of these planets:
with sns.axes_style('white'):
g = sns.factorplot("year", data=planets, aspect=4.0, kind='count',
hue='method', order=range(2001, 2015))
g.set_ylabels('Number of Planets Discovered')
# ## Example: Exploring Marathon Finishing Times
#
# Here we'll look at using Seaborn to help visualize and understand finishing results from a marathon.
#
data = pd.read_csv('marathon-data.csv')
data.head()
# By default, Pandas loaded the time columns as Python strings (type ``object``); we can see this by looking at the ``dtypes`` attribute of the DataFrame:
data.dtypes
import datetime as datetime
# Let's fix this by providing a converter for the times:
# +
def convert_time(s):
h, m, s = map(int, s.split(':'))
return datetime.timedelta(hours=h, minutes=m, seconds=s)
data = pd.read_csv('marathon-data.csv',
converters={'split':convert_time, 'final':convert_time})
data.head()
# -
data.dtypes
# That looks much better. For the purpose of our Seaborn plotting utilities, let's next add columns that give the times in seconds:
data['split_sec'] = data['split'].astype(int) / 1E9
data['final_sec'] = data['final'].astype(int) / 1E9
data.head()
# To get an idea of what the data looks like, we can plot a ``jointplot`` over the data:
with sns.axes_style('white'):
g = sns.jointplot("split_sec", "final_sec", data, kind='hex')
g.ax_joint.plot(np.linspace(4000, 16000),
np.linspace(8000, 32000), ':k')
# The dotted line shows where someone's time would lie if they ran the marathon at a perfectly steady pace. The fact that the distribution lies above this indicates (as you might expect) that most people slow down over the course of the marathon.
# If you have run competitively, you'll know that those who do the opposite—run faster during the second half of the race—are said to have "negative-split" the race.
#
# Let's create another column in the data, the split fraction, which measures the degree to which each runner negative-splits or positive-splits the race:
data['split_frac'] = 1 - 2 * data['split_sec'] / data['final_sec']
data.head()
# Where this split difference is less than zero, the person negative-split the race by that fraction.
# Let's do a distribution plot of this split fraction:
sns.distplot(data['split_frac'], kde=False);
plt.axvline(0, color="k", linestyle="--");
sum(data.split_frac < 0)
# Out of nearly 40,000 participants, there were only 250 people who negative-split their marathon.
#
# Let's see whether there is any correlation between this split fraction and other variables. We'll do this using a ``pairgrid``, which draws plots of all these correlations:
g = sns.PairGrid(data, vars=['age', 'split_sec', 'final_sec', 'split_frac'],
hue='gender', palette='RdBu_r')
g.map(plt.scatter, alpha=0.8)
g.add_legend();
# It looks like the split fraction does not correlate particularly with age, but does correlate with the final time: faster runners tend to have closer to even splits on their marathon time.
# (We see here that Seaborn is no panacea for Matplotlib's ills when it comes to plot styles: in particular, the x-axis labels overlap. Because the output is a simple Matplotlib plot, however, the methods in [Customizing Ticks](04.10-Customizing-Ticks.ipynb) can be used to adjust such things if desired.)
#
# The difference between men and women here is interesting. Let's look at the histogram of split fractions for these two groups:
sns.kdeplot(data.split_frac[data.gender=='M'], label='men', shade=True)
sns.kdeplot(data.split_frac[data.gender=='W'], label='women', shade=True)
plt.xlabel('split_frac');
# The interesting thing here is that there are many more men than women who are running close to an even split!
# This almost looks like some kind of bimodal distribution among the men and women. Let's see if we can suss-out what's going on by looking at the distributions as a function of age.
#
# A nice way to compare distributions is to use a *violin plot*
sns.violinplot("gender", "split_frac", data=data,
palette=["lightblue", "lightpink"]);
# This is yet another way to compare the distributions between men and women.
#
# Let's look a little deeper, and compare these violin plots as a function of age. We'll start by creating a new column in the array that specifies the decade of age that each person is in:
data['age_dec'] = data.age.map(lambda age: 10 * (age // 10))
data.head()
# +
men = (data.gender == 'M')
women = (data.gender == 'W')
with sns.axes_style(style=None):
sns.violinplot("age_dec", "split_frac", hue="gender", data=data,
split=True, inner="quartile",
palette=["lightblue", "lightpink"]);
# -
# Looking at this, we can see where the distributions of men and women differ: the split distributions of men in their 20s to 50s show a pronounced over-density toward lower splits when compared to women of the same age (or of any age, for that matter).
#
# Also surprisingly, the 80-year-old women seem to outperform *everyone* in terms of their split time. This is probably due to the fact that we're estimating the distribution from small numbers, as there are only a handful of runners in that range:
(data.age > 80).sum()
# Back to the men with negative splits: who are these runners? Does this split fraction correlate with finishing quickly? We can plot this very easily. We'll use ``regplot``, which will automatically fit a linear regression to the data:
g = sns.lmplot('final_sec', 'split_frac', col='gender', data=data,
markers=".", scatter_kws=dict(color='c'))
g.map(plt.axhline, y=0.1, color="k", ls=":");
# Apparently the people with fast splits are the elite runners who are finishing within ~15,000 seconds, or about 4 hours. People slower than that are much less likely to have a fast second split.
| notebooks/Matplotlib_Seaborn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Pyro에서 독립 차원 선언: plate"
# > "plate를 이용하여 데이터의 각 사례(example)는 독립적이라고 선언한다."
#
# - toc: true
# - badges: true
# - author: 단호진
# - categories: [ppl]
# 시계열 데이터가 아닌 데이터 셋의 각 사례(example)를 생각해보자. 그 사례는 어떤 파라미터를 가진 확률 변수의 실현값이라고 생각할 수 있다. 고정된 파라미터라고 해도 확률 과정이므로 표본 사례의 값은 서로 다를 수 있다. 한편 사례와 사례의 차이는 파라이터의 차이에서도 나올 수 있다. 데이터의 사례뿐만 아니라 숨은 변수가 숨은 확률 변수들의 결합 확률 분포를 따르는 상황도 있을 수 있다. 어는 경우든 데이터의 독립성을 선언해야 할 필요가 생긴다. 이 포스트에서는 plate 컨텍스트를 이용하여 데이터의 각 사례가 독립적이라고 선언하는 방법을 정리해 보겠다.
#
# [Pyro의 확률적 통계 추론 입문](https://pyro.ai/examples/svi_part_i.html) 튜토리얼의 동전 편형성 문제를 plate를 사용하여 재 구성해보겠다.
# ## 동전 편향 문제
#
# 동전을 열번 던져서 앞면이 7회 나왔다고 할 때 빈도주의에서는 p=0.5로 가정하고 분석을 진행한다. 이항 분포를 따른다고 할 때 관측의 확률은 $\binom{10}{3} \frac{1}{2^{10}} = 0.117$이다. 3개보다 더 적게 나오거나 7개 이상 나오는 경우를 따져보면 다음과 같다.
# +
from math import comb
from functools import reduce
pr = map(lambda k: comb(10, k) / 2 ** 10, [0, 1, 2, 3])
pr = reduce(lambda a, x: a + x, pr) * 2 # two-tail
pr
# -
# 빈도주의 관점에서 p=0.5라는 것을 기각하기 어렵다. 베이즈 통계라면 관측 데이터에서 최선의 p를 추정해 보고자 한다. 사전 분포를 B(15.0, 15.0)인 베타 함수로 정의하여 p=0.5 중심에서 약간의 편향이 존재할 수 있다고 보았다. 관측 데이터를 얻은 후 사후 분포를 pyro로 구해 보자.
# +
import numpy as np
import pandas as pd
import seaborn as sns
from scipy.stats import beta
import matplotlib.pyplot as plt
def plot(a, b, ax):
rv = beta(a, b)
p = np.linspace(0.1, 0.9, 41)
df = pd.DataFrame(dict(p=p, pdf=rv.pdf(p)))
return sns.lineplot(x='p', y='pdf', data=df, ax=ax)
fig, ax = plt.subplots()
a, b = 15.0, 15.0
ax = plot(a, b, ax)
ax.legend(['prior']);
# +
import torch
from torch.distributions import constraints
import pyro
import pyro.distributions as dist
pyro.enable_validation(True)
# -
# ## ELBO
#
# ELBO(Evidence Lower BOund)의 최소화 하고자 한다. $ELBO \equiv \mathbb{E}_{q_{\phi} (z)} [\log p_{\theta} (x, z) - \log q_{\phi} (z)]$ 식으로 주어 진다면 $\log p_{\theta} (x, z)$는 model 함수에서 $\log q_{\phi} (z)$는 guide 함수를 통해서 구하게 된다. 배경 이론에 대해서는 [Pyro의 확률적 통계 추론 입문](https://pyro.ai/examples/svi_part_i.html) 튜토리얼을 참고하자. pyro.sample 함수가 정의되면 내부적으로 log_prob 함수를 통하여 로그 확률을 평가하게 된다.
#
# 이 문제에서 guide에 베이즈 분석의 사전 분포를 정의하였다. 알고자 하는 분포의 근사로 생각해도 좋다.
# ## 데이터
data = [1.0] * 7 + [0.0] * 3
data = torch.tensor(data)
data
# ## 모델
# model1 + guide1은 숨은 확률 변수 z에서 하나의 값을 실현하고 그 값에 대하여 관측치를 평가한다. 동일 동전으로 10번의 동전 던지기를 수행했다고 볼 수 있다.
# +
def guide1(data):
alpha_q = pyro.param(
'alpha_q', torch.tensor(15.0), constraint=constraints.positive)
beta_q = pyro.param(
'beta_q', torch.tensor(15.0), constraint=constraints.positive)
pyro.sample('z', dist.Beta(alpha_q, beta_q)) # shape: []
def model1(data):
alpha0 = torch.tensor(10.0)
beta0 = torch.tensor(10.0)
z = pyro.sample('z', dist.Beta(alpha0, beta0))
with pyro.plate('data', len(data)):
pyro.sample(
'obs',
dist.Bernoulli(z),
obs=data)
# -
# ## 훈련
# +
import numpy as np
svi = pyro.infer.SVI(
model1, guide1, pyro.optim.Adam({'lr': 0.0005}), pyro.infer.Trace_ELBO())
steps = 2000
for step in range(steps):
l = svi.step(data)
if step % 100 == 0:
alpha_q, beta_q = pyro.param('alpha_q').item(), pyro.param('beta_q').item()
print(f'loss: {l:.2f}, alpha_q: {alpha_q:.2f}, beta_q: {beta_q:.2f}')
inferred_mean = alpha_q / (alpha_q + beta_q)
# compute inferred standard deviation
factor = beta_q / (alpha_q * (1.0 + alpha_q + beta_q))
inferred_std = inferred_mean * np.sqrt(factor)
print("\nbased on the data and our prior belief, the fairness " +
"of the coin is %.3f +- %.3f" % (inferred_mean, inferred_std))
# -
# ## 사후 분포
fig, ax = plt.subplots()
ax = plot(15.0, 15.0, ax)
a, b = pyro.param('alpha_q').item(), pyro.param('beta_q').item()
ax = plot(a, b, ax)
ax.legend(['prior', 'model/guide 1']);
# * 사후 분포가 사전 분포에서 관측에 의한 p=7/10 방향으로 이동하였다.
# * MAP로 동전의 편향을 점추정하거나 범위를 추정할 수 있다.
# * 추가로 동전 던지기를 한다면 새로운 prior로 이 모델을 이용할 수도 있다.
| _notebooks/2021-01-22-pyro-plate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import sys
import logging
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3' # FATAL
logging.getLogger('tensorflow').setLevel(logging.DEBUG)
try:
from google.colab import drive
drive.mount('/content/drive')
# !pip install -q ruamel.yaml
# !pip install -q tensorboard-plugin-profile
project_path = '/content/drive/MyDrive/Colab Projects/QuantumFlow'
except:
project_path = os.path.expanduser('~/QuantumFlow')
# +
os.chdir(project_path)
sys.path.append(project_path)
import numpy as np
import tensorflow as tf
import tree
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
import quantumflow
experiment = 'xdiff'
run_name = 'default'
epoch = None
preview = 5
base_dir = os.path.join(project_path, "experiments", experiment)
params = quantumflow.utils.load_yaml(os.path.join(base_dir, 'hyperparams.yaml'))[run_name]
run_dir = os.path.join(base_dir, run_name)
# -
dataset_validate = quantumflow.instantiate(params['dataset_validate'], run_dir=run_dir)
dataset_validate.build()
_ = quantumflow.get_class(params['model']['class'])
model = tf.keras.models.load_model(os.path.join(run_dir, 'saved_model'))
if epoch is not None: _ = model.load_weights(os.path.join(run_dir, params['checkpoint']['filename'].format(epoch=epoch)))
model.summary()
# +
@tf.function
def predict_fn(features_batch):
density = tf.nest.flatten(features_batch)
with tf.GradientTape() as tape:
tape.watch(density)
outputs_batch = model(density)
outputs_batch['derivative'] = 1/dataset_validate.h*tape.gradient(outputs_batch['kinetic_energy'], density)[0]
return outputs_batch
def predict(features, batch_size=None):
if batch_size is None:
return tree.map_structure(lambda out: out.numpy(), predict_fn(features))
else:
outputs = []
dataset_size = tree.flatten(features)[0].shape[0]
steps = -(-dataset_size//batch_size)
print_steps = max(1, steps//100)
print('/', dataset_size)
for i in range(steps):
if i % print_steps == 0: print(i*batch_size, end=' ')
features_batch = tree.map_structure(lambda inp: inp[i*batch_size:(i+1)*batch_size], features)
outputs.append(predict_fn(features_batch))
print()
return tree.map_structure(lambda *outs: np.concatenate(outs, axis=0), *outputs)
# -
targets_pred = predict(dataset_validate.features, params['dataset_validate'].get('max_batch_size', 1))
targets_pred['kinetic_energy'][:preview]
dataset_validate.targets['kinetic_energy'][:preview]
kinetic_energy_err = targets_pred['kinetic_energy'] - dataset_validate.targets['kinetic_energy'][:len(targets_pred['kinetic_energy'])]
kcalmol_per_hartree = 627.5094738898777
np.mean(np.abs(kinetic_energy_err))*kcalmol_per_hartree
plt.figure(figsize=(20, 3))
plt.plot(dataset_validate.x, dataset_validate.targets['kinetic_energy_density'][:preview, :].transpose(), 'k:')
plt.plot(dataset_validate.x, targets_pred['kinetic_energy_density'][:preview, :].transpose())
plt.show()
plt.figure(figsize=(20, 3))
plt.plot(dataset_validate.x, dataset_validate.targets['kinetic_energy_density'][:preview, :].transpose() - targets_pred['kinetic_energy_density'][:preview, :].transpose())
plt.show()
targets_pred['derivative'].shape
plt.figure(figsize=(20, 3))
plt.plot(dataset_validate.x, dataset_validate.derivative[:preview, :].transpose(), 'k:')
plt.plot(dataset_validate.x, targets_pred['derivative'][:preview, :].transpose())
plt.show()
tf.config.experimental.get_memory_info('GPU:0')['peak']/1024**3
| notebooks/test_dft_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:untitled1]
# language: python
# name: conda-env-untitled1-py
# ---
# # Plotting a pourbaix diagram using the Materials Project API
# *If you use this infrastructure, please consider citing the following work:*
#
# [<NAME>, <NAME>, <NAME> and <NAME>, Phys. Rev. B, 2012, 85, 235438.](https://journals.aps.org/prb/abstract/10.1103/PhysRevB.85.235438)
#
# [<NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME> and <NAME>, Chem. Mater., 2017, DOI: acs.chemmater.7b03980.](http://pubs.acs.org/doi/10.1021/acs.chemmater.7b03980)
#
# **Notebook Author:** <NAME>
# ----------
#
# The Materials Project REST interface includes functionality to construct pourbaix diagrams from computed entries. Note that the Pourbaix infrastructure is still undergoing revision, but now includes a simplified interface that enables MP and pymatgen users to fetch entries that have been processed according to the [Materials Project Aqueous Compatibility scheme](http://pymatgen.org/_modules/pymatgen/entries/compatibility.html). Thus, users can reproduce web Pourbaix diagrams in two or three steps in pymatgen.
# +
# Import necessary tools from pymatgen
from pymatgen import MPRester
from pymatgen.analysis.pourbaix_diagram import PourbaixDiagram, PourbaixPlotter
# %matplotlib inline
# Initialize the MP Rester
mpr = MPRester()
# -
# To retrieve entries necessary to construct a Pourbaix Diagram use `MPRester.get_pourbaix_entries(LIST_OF_ENTRIES)` with a list of entries comprising your chemical system. It is not necessary to include 'O' and 'H' in your list, as they are added automatically. This function also contains all of necessary preprocessing to ensure the PourbaixEntries are compatible with the pymatgen PourbaixDiagram constructor.
# Get all pourbaix entries corresponding to the Cu-O-H chemical system.
entries = mpr.get_pourbaix_entries(["Cu"])
# Pourbaix diagrams can be constructed using `PourbaixDiagram(RETRIEVED_ENTRIES)` as below. Note that a `comp_dict` keyword argument may also be supplied to the `PourbaixDiagram` constructor if a fixed composition for a multi-element pourbaix diagram is desired.
# Construct the PourbaixDiagram object
pbx = PourbaixDiagram(entries)
# The `PourbaixAnalyzer` includes a number of useful functions for determining stable species and stability of entries relative to a given pourbaix facet (i.e. as a function of pH and V).
# Get an entry stability as a function of pH and V
entry = [e for e in entries if e.entry_id == 'mp-1692'][0]
print("CuO's potential energy per atom relative to the most",
"stable decomposition product is {:0.2f} eV/atom".format(
pbx.get_decomposition_energy(entry, pH=7, V=-0.2)))
# This suggests that CuO, for example, has a large driving force for decomposition at neutral pH and mildly reducing conditions.
# To see this in more detail, we can plot the pourbaix diagram. The `PourbaixPlotter` object is also initialized using an instance of the `PourbaixDiagram` object.
plotter = PourbaixPlotter(pbx)
plotter.get_pourbaix_plot().show()
# The PourbaixPlotter object can also plot the relative stability of an entry across the pourbaix diagram. To do this, use the PourbaixPlotter.plot_entry_stability method.
plt = plotter.plot_entry_stability(entry)
plt.show()
# # Plotting k-nary systems
# Pymatgen also supports binary/ternary pourbaix diagrams with fixed compositions of the non-H or O elements. This is achieved by finding all possible combinations of entries that fulfill the composition constraint and treating them as individual entries in pourbaix space. Note that you can supply a composition dictionary and to further tune the pourbaix diagram.
# Get all pourbaix entries corresponding to the Fe-O-H chemical system.
entries = mpr.get_pourbaix_entries(["Bi", "V"])
# Construct the PourbaixDiagram object
pbx = PourbaixDiagram(entries, comp_dict={"Bi": 0.5, "V": 0.5},
conc_dict={"Bi": 1e-8, "V": 1e-8}, filter_solids=True)
# Note that the `filter_solids` keyword argument in the `PourbaixDiagram` instantiation above tells the constructor whether to filter solids by phase stability on the compositional phase diagram. Note that Pourbaix Diagrams generated with and without this argument may look significantly different in the OER and HER regions, since highly oxidized materials (e. g. Bi$_2$O$_5$) or highly reduced materials (e. g. most hydrides) may not be stable on the compositional phase diagram. The filtering process significantly reduces the time it takes to generate all of the combined entries for a binary or ternary pourbaix diagram though, so it may be prudent to use in those cases.
# Construct the pourbaix analyzer
plotter = PourbaixPlotter(pbx)
plt = plotter.get_pourbaix_plot()
plt.show()
# Getting the heatmaps for a solid entry in these cases is a bit more involved, because many of the regions of the pourbaix diagram include multiphase entries. Here's an example for this case.
bivo4_entry = [entry for entry in entries if entry.entry_id=="mp-613172"][0]
plt = plotter.plot_entry_stability(bivo4_entry)
# If a compound is provided that doesn't meet the constraints of the composition (in this case, equal parts Bi and V), the plotter will attempt to find the most stable entry containing that solid and any spectating ions. In this case, it estimates the stability of BiO$_2$(s) + VO$_4$-.
bio2_entry = [entry for entry in entries if entry.entry_id=="mp-557993"][0]
plt = plotter.plot_entry_stability(bio2_entry)
| notebooks/2017-12-15-Plotting a Pourbaix Diagram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Lm49B6yjpdl9" colab_type="code" colab={}
# #!pip install eli5
# + id="I27nlp81pg2x" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
import eli5
from eli5.sklearn import PermutationImportance
from ast import literal_eval
from tqdm import tqdm_notebook
# + id="EMW9hgV1plfK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9046fa8c-43a1-48f0-ab9e-38610a5c1f70" executionInfo={"status": "ok", "timestamp": 1581892279623, "user_tz": -60, "elapsed": 793, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix"
# + id="jeQ-7rWeptXD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="02189c63-49f1-429e-c372-0320b8e989de" executionInfo={"status": "ok", "timestamp": 1581892284635, "user_tz": -60, "elapsed": 2226, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
# ls
# + id="LgxmO6ePpvd7" colab_type="code" colab={}
# !git add matrix_one/day5.ipynb
# + id="EaxRLMYjp1ol" colab_type="code" colab={}
df = pd.read_csv('data/men_shoes.csv', low_memory=False)
# + id="-unDoCmdp6x8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="510cd3b2-f29f-4f5f-d1b5-83a308322f3b" executionInfo={"status": "ok", "timestamp": 1581884363060, "user_tz": -60, "elapsed": 645, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df.columns
# + id="om5zHtPyp82z" colab_type="code" colab={}
def run_model(feautures, model=DecisionTreeRegressor(max_depth=5)):
X = df[ feautures ].values
y = df['prices_amountmin'].values
scores = cross_val_score(model, X, y, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="L3KLuzzuqDih" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0c96f6bb-f6b9-4155-851e-f00bde11858c" executionInfo={"status": "ok", "timestamp": 1581884403729, "user_tz": -60, "elapsed": 544, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df['brand_cat'] = df['brand'].map(lambda x: str(x).lower()).factorize()[0]
run_model(['brand_cat'])
# + id="SRgCVNBtqG02" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c25bd55a-63c2-4d81-af85-5efefb2b6471" executionInfo={"status": "ok", "timestamp": 1581884416938, "user_tz": -60, "elapsed": 3605, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
model = RandomForestRegressor(max_depth=5, n_estimators=100, random_state=0)
run_model(['brand_cat'], model)
# + id="RDQQH_cdqJRd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="df199541-3c92-46dd-ab25-fda2de7f6645" executionInfo={"status": "ok", "timestamp": 1581884728709, "user_tz": -60, "elapsed": 571, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df.features.head().values
# + id="DI9kKjcUrPZw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c5032d9c-2c17-48f8-b94f-822e6fd2105f" executionInfo={"status": "ok", "timestamp": 1581884808540, "user_tz": -60, "elapsed": 736, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
str_dict = '[{"key":"Gender","value":["Men"]},{"key":"Color","value":["Black"]},{"key":"Shipping Weight (in pounds)","value":["0.45"]},{"key":"Condition","value":["New"]},{"key":"Brand","value":["SERVUS BY HONEYWELL"]},{"key":"manufacturer_part_number","value":["ZSR101BLMLG"]}]'
literal_eval(str_dict)[0]['value'][0]
# + id="cTDbPI80ridH" colab_type="code" colab={}
def parse_features(x):
output_dict = {}
if str(x) == 'nan': return output_dict
features = literal_eval(x.replace('\\"','"'))
for item in features:
key = item['key'].lower().strip()
value = item['value'][0].lower().strip()
output_dict[key] = value
return output_dict
#{'key': 'Gender', 'value': ['Men']}
df['features_parsed'] = df['features'].map(parse_features)
# + id="VRdHc6AYuLeS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fbb41e91-ed29-4972-89b1-110d879f9d03" executionInfo={"status": "ok", "timestamp": 1581885498541, "user_tz": -60, "elapsed": 527, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
keys = set()
df['features_parsed'].map(lambda x: keys.update(x.keys()))
len(keys)
# + id="Adg7w2NruSHw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="3d7e9308-b80f-4c41-d85a-befb63eee343" executionInfo={"status": "ok", "timestamp": 1581885943925, "user_tz": -60, "elapsed": 565, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "14751556941823709984"}}
df.features_parsed.head().values
# + id="1LLOScK7v-0F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["9dc0b65f1cc646eb9ca4c3f7ec191c2d", "900071003a704156adfa734940b823cd", "cd0e06be352b4421b61bf3114c0e7311", "c33606aed7434c268bd38f7ebfe0098b", "d7248799bbef4439a98598ef3b4955a8", "938292e1a76148d5b55273b09db2001c", "7dd28bc014d74b938540056033ed17a0", "049d3c5e271f4e9c8eaa1332c2eea4f4"]} outputId="f3701a22-95ea-43f6-cbcc-5eb4c10192dc" executionInfo={"status": "ok", "timestamp": 1581885996379, "user_tz": -60, "elapsed": 4117, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
def get_name_feat(key):
return 'feat_' + key
for key in tqdm_notebook(keys):
df[get_name_feat(key)] = df.features_parsed.map(lambda feats: feats[key] if key in feats else np.nan)
# + id="YBGrygqiwKyG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="a0acd38d-4a35-43c1-a452-a67853765e27" executionInfo={"status": "ok", "timestamp": 1581886008469, "user_tz": -60, "elapsed": 680, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df.columns
# + id="RZb_qTqYwOcQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="810b8ac2-e9e5-4d85-e6df-2328ce76c14e" executionInfo={"status": "ok", "timestamp": 1581886045074, "user_tz": -60, "elapsed": 652, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df[ df['feat_athlete'].isnull()].shape
# + id="K6AKFNX_wXhK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cab05cbe-0f70-48c6-bfea-7004b6790081" executionInfo={"status": "ok", "timestamp": 1581886303755, "user_tz": -60, "elapsed": 513, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df.shape[0]
# + id="StGxN7HWwZnx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1b14cc1e-b8b6-4604-c818-750cd223819f" executionInfo={"status": "ok", "timestamp": 1581886418235, "user_tz": -60, "elapsed": 2088, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df [ False == df['feat_athlete'].isnull()].shape[0]
# + id="AFI2fAxwxyIY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="348f20d4-f297-496a-bbbe-ea88070fa2ac" executionInfo={"status": "ok", "timestamp": 1581886449846, "user_tz": -60, "elapsed": 542, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df [ False == df['feat_athlete'].isnull()].shape[0]/df.shape[0] * 100
# + id="GI8npN6Ux3Ov" colab_type="code" colab={}
keys_stat = {}
for key in keys:
keys_stat[key] = df [ False == df[get_name_feat(key)].isnull()].shape[0]/df.shape[0] * 100
# + id="qpsLFiTuyPLg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="7a9d9dee-03d9-402f-a535-966990a3d420" executionInfo={"status": "ok", "timestamp": 1581886824708, "user_tz": -60, "elapsed": 483, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
{k: v for k,v in keys_stat.items() if v > 30}
# + id="9NGhm6A7yTFG" colab_type="code" colab={}
df['feat_brand_cat'] = df['feat_brand'].factorize()[0]
df['feat_color_cat'] = df['feat_color'].factorize()[0]
df['feat_gender_cat'] = df['feat_gender'].factorize()[0]
df['feat_manufacturer part number_cat'] = df['feat_manufacturer part number'].factorize()[0]
df['feat_material_cat'] = df['feat_material'].factorize()[0]
df['feat_sport_cat'] = df['feat_sport'].factorize()[0]
df['feat_style_cat'] = df['feat_style'].factorize()[0]
for key in keys:
df[get_name_feat(key) + '_cat'] = df[get_name_feat(key)].factorize()[0]
# + id="aCJl8k1azqdn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="b30db18e-91b6-4484-f67c-33e5c43baf2c" executionInfo={"status": "ok", "timestamp": 1581887210472, "user_tz": -60, "elapsed": 465, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df[df.brand == df.feat_brand][['brand', 'feat_brand']].head()
# + id="3I_IjFDm096O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="6316dc2c-18f6-40a9-e48c-a182e994a910" executionInfo={"status": "ok", "timestamp": 1581887256124, "user_tz": -60, "elapsed": 644, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df[df.brand != df.feat_brand][['brand', 'feat_brand']].head()
# + id="P2HOLuxq1AT2" colab_type="code" colab={}
df['brand'] = df['brand'].map(lambda x:str(x).lower())
# + id="IkQ3f5bN1MU5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f2ec7e6f-27f1-4090-81df-f009e54a8b6c" executionInfo={"status": "ok", "timestamp": 1581887388909, "user_tz": -60, "elapsed": 483, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df[df.brand == df.feat_brand].shape
# + id="G9ic8-Mg0gQk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="825be0ca-f535-4618-9608-e5b3192c223e" executionInfo={"status": "ok", "timestamp": 1581887447305, "user_tz": -60, "elapsed": 3576, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
model = RandomForestRegressor(max_depth=5, n_estimators=100)
run_model(['brand_cat'], model)
# + id="ofsIqsBw6pj8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f34e0628-9bc0-4fc7-e18d-58943cbd2f09" executionInfo={"status": "ok", "timestamp": 1581888836023, "user_tz": -60, "elapsed": 486, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
feats_cat = [x for x in df.columns if 'cat' in x ]
feats_cat
# + id="kQkDiwgl1tGx" colab_type="code" colab={}
feats = ['brand_cat', 'feat_metal type_cat', 'feat_brand_cat', 'feat_gender_cat', 'feat_material_cat', 'feat_adjustable_cat', 'feat_case thickness_cat']
#feats += feats_cat
#feats = list(set(feats))
model = RandomForestRegressor(max_depth=5, n_estimators=100)
result = run_model(feats, model)
# + id="5ZBtQt1N27l6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="253526fc-d9dc-4e7a-b5db-0a6f4a425c0f" executionInfo={"status": "ok", "timestamp": 1581891489842, "user_tz": -60, "elapsed": 3785, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
X = df[ feats ].values
y = df['prices_amountmin'].values
m = RandomForestRegressor(max_depth=5, n_estimators=100, random_state=0)
m.fit(X, y)
print(result)
perm = PermutationImportance(m, random_state=1).fit(X, y);
eli5.show_weights(perm, feature_names=feats)
# + id="SLQ0eEAB4DLx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="1eb5888e-3aed-42b7-d116-5c5fdf059824" executionInfo={"status": "ok", "timestamp": 1581888234400, "user_tz": -60, "elapsed": 2393, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df["brand"].value_counts(normalize=True)
# + id="Vj-yBYW34oT_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="163c96c7-b6f4-4ef4-b300-4f2d1e3eb9a0" executionInfo={"status": "ok", "timestamp": 1581888530441, "user_tz": -60, "elapsed": 1196, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df[df['brand'] == 'nike'].features_parsed.sample(5).values
# + id="h7KmgiPT49Z4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="eb62e8ef-467b-4185-c5d9-70b9137e418c" executionInfo={"status": "ok", "timestamp": 1581888583778, "user_tz": -60, "elapsed": 910, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBuV0ZYs4G1JcHJGjxAAh0kSuBYyHDVQ1UA7oSoxg=s64", "userId": "14751556941823709984"}}
df['feat_age group'].value_counts()
| matrix_one/day5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Installs
pip install pymongo==3.11.0
pip install matplotlib
# # Imports
from os import environ
from pprint import pprint as pp
from pymongo import MongoClient
import matplotlib.pyplot as plt
# # Database Link
client = MongoClient(environ["MONGO_PORT_27017_TCP_ADDR"], 27017)
database = client.polyplot
collection = database["indicators"]
# # Basic Functions
# +
def drop_collection():
collection.drop()
def find_indicator(query={}, filter={"_id": 0}):
return dict(collection.find_one(query, filter))
def find_indicators(query={}, filter={"_id": 0}, sort=[("name", 1)], limit=0):
collection.create_index(sort)
return list(collection.find(query, filter).sort(sort).limit(limit))
def update_indicator(indicator):
return collection.update_one({"code": indicator["code"]}, {"$set": indicator})
def update_indicators():
for indicator in find_indicators():
update_indicator(indicator)
# -
# # Indicator Completeness
#
# Indicator completeness is a measure of how complete the dataset is within the given scope.
#
# The cell below displays a simple histogram to help visualize the distribution of indicator completeness.
# +
completeness = []
for indicator in find_indicators(filter={"_id": 0, "completeness": 1, "size": 1}):
if "completeness" in indicator: completeness.append(indicator["completeness"])
plt.hist(completeness, bins=20)
plt.show()
# -
# # Indicator Size
#
# Indicator size is a measure of the approximate byte size of the given indicator.
#
# The cell below displays a simple histogram to help visualize the distribution of indicator size.
# +
size = []
for indicator in find_indicators(filter={"_id": 0, "completeness": 1, "size": 1}):
if "size" in indicator: size.append(indicator["size"])
plt.hist(size, bins=50)
plt.show()
| notes/collections/indicators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="LrELv42oqtC-" outputId="88236aab-4036-4764-8d70-b402a6ce99c2"
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
from __future__ import print_function
import tensorflow as tf
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
from keras.layers.normalization import BatchNormalization
# + colab={} colab_type="code" id="9H3JvVkScrZT"
import matplotlib.pyplot as plt
# + colab={} colab_type="code" id="18s9TXnNaVXl"
config = tf.ConfigProto( device_count = {'GPU': 1 , 'CPU': 56} )
sess = tf.Session(config=config)
keras.backend.set_session(sess)
# + colab={} colab_type="code" id="sFg0N_oqaFOf"
def plt_dynamic(fig, x, vy, ty, ax, colors=['b']):
ax.plot(x, vy, 'b', label="Validation Loss")
ax.plot(x, ty, 'r', label="Train Loss")
plt.legend()
plt.grid()
fig.canvas.draw()
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="iYrfygMRq3e6" outputId="62f851ec-0a7e-45de-ec8d-4a57204dfb5b"
batch_size = 128
num_classes = 10
epochs = 12
# input image dimensions
img_rows, img_cols = 28, 28
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="MpJ-0UZNq57Y" outputId="ccd3c84d-d873-4c73-c4c2-9945d0766add"
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# + [markdown] colab_type="text" id="2ysrt2LEjrMF"
# ## [1] 3x3 convolution kernel
# + [markdown] colab_type="text" id="qonLJFqxjzEC"
# ### [1.1] Conv (32-64) | MaxPool | Dropout x2 | Dense
# + colab={"base_uri": "https://localhost:8080/", "height": 442} colab_type="code" id="ZZmb_ZsRq_wB" outputId="178ea1d8-e564-408c-b085-ca2dcb401d5a"
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 408} colab_type="code" id="sZ4FNweldu1u" outputId="8f632c76-25c6-44a1-d920-6435a8e9da78"
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="WSGIRXo3bqrx" outputId="fe10eb9b-01de-4794-ec42-c1e486affca5"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="vGUO_xp9kkG7"
# ### [1.2] Conv (64-32) | MaxPool | Dropout x2 | Dense
# + colab={"base_uri": "https://localhost:8080/", "height": 833} colab_type="code" id="OgrHc0O2cPNo" outputId="a0c4d795-b3f0-4469-d5b2-bbe6bd52c6a9"
model = Sequential()
model.add(Conv2D(
64, kernel_size=(3, 3), activation='relu', input_shape=input_shape
))
model.add(Conv2D(32, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="wVB8Vwhsi-yX" outputId="7978f621-8a45-43ff-ed11-caf9b378a004"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="6cRREIRdkqNZ"
# ### [1.3] Conv (64-32) | MaxPool | Dropout x2 | Dense | Padding (same)
# + colab={"base_uri": "https://localhost:8080/", "height": 833} colab_type="code" id="ICGP10lVjDSk" outputId="9ff6e732-f332-46d8-f0c6-dc70a1005924"
model = Sequential()
model.add(Conv2D(
64, kernel_size=(3, 3), activation='relu', input_shape=input_shape,
padding='same'
))
model.add(Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="SWjkRecKkajC" outputId="4c83d15d-f71c-4d91-9357-a6bdf15c2e29"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="3htdZDsxk9U_"
# ### [1.4] Conv (64-32) | MaxPool | Dropout x2 | Dense | Padding (same) | BN
# + colab={"base_uri": "https://localhost:8080/", "height": 1040} colab_type="code" id="GikKrnD1MTY5" outputId="e2287bf6-fef2-4072-b6c8-4b3aa976d476"
model = Sequential()
model.add(Conv2D(
64, kernel_size=(3, 3), activation='relu', input_shape=input_shape,
padding='same'
))
model.add(Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="ZtsenGVZMbLW" outputId="5b03354a-23e6-489d-bf5f-957fb40916c3"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="W2y7QnfylDlK"
# ### [1.5] Conv (64-32) | MaxPool | Dropout x2 | Dense (256-128) | Padding (same) | BN
# + colab={"base_uri": "https://localhost:8080/", "height": 935} colab_type="code" id="Jd0Yp4t3McMJ" outputId="f4f377a3-6fb4-45c8-be65-e91458516336"
model = Sequential()
model.add(Conv2D(
64, kernel_size=(3, 3), activation='relu', input_shape=input_shape,
padding='same'
))
model.add(Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(rate=0.5))
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 316} colab_type="code" id="GnvSSxTGN_BA" outputId="9c1f96fa-1a23-4891-f70b-415c1a0a2e01"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="bDa8tpfClJVw"
# ## [2] Convolution 5x5 kernel
# + [markdown] colab_type="text" id="i0RwKRadlOYQ"
# ### [2.1] Conv (128-64-32) | MaxPool | Dropout x2 | Dense
# + colab={"base_uri": "https://localhost:8080/", "height": 867} colab_type="code" id="3wevi7MeOAF6" outputId="9130e3cf-2deb-4288-f6a6-0a4c2612de2e"
model = Sequential()
model.add(Conv2D(
128, kernel_size=(5, 5), activation='relu', input_shape=input_shape
))
model.add(Conv2D(64, (5, 5), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Conv2D(32, (5, 5), activation='relu'))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="Tmc_rDVMPSFO" outputId="4370e257-1b7e-4cf5-be1c-cb9935c540aa"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="_s13tlualaHG"
# ### [2.2] Conv (128-64-32) | MaxPool x2 | Dropout x2 | Dense
# + colab={"base_uri": "https://localhost:8080/", "height": 901} colab_type="code" id="vPks7otbPVNH" outputId="0a9d906f-4928-4a34-ed9b-451dfe9541b4"
model = Sequential()
model.add(Conv2D(
128, kernel_size=(5, 5), activation='relu', input_shape=input_shape
))
model.add(Conv2D(64, (5, 5), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Conv2D(32, (5, 5), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="bTDrEz6RQorK" outputId="a7e6c369-a22f-491e-babd-3d06895f7aa4"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="lW927xDHlw3t"
# ### [2.3] Conv (128-64-32) | MaxPool x2 | Dropout x3 | Dense | BN x2
# + colab={"base_uri": "https://localhost:8080/", "height": 1037} colab_type="code" id="RBk13dFcQz0s" outputId="8e09d42c-e249-4e6c-a52f-03003119d6ad"
model = Sequential()
model.add(Conv2D(
128, kernel_size=(5, 5), activation='relu', input_shape=input_shape
))
model.add(Conv2D(64, (5, 5), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Conv2D(32, (5, 5), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(rate=0.5))
model.add(Dense(64, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="n0FjpP8sR8jL" outputId="3edcbf87-936b-46a2-ca6e-7cfb9318b7c8"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="hpA2r5COl5hh"
# ### [2.4] Conv (128-64-32) | MaxPool x2 | Dropout x3 | Dense (128-64) | Padding (same) | BN
# + colab={"base_uri": "https://localhost:8080/", "height": 1071} colab_type="code" id="lasLlWEsR9f5" outputId="af772955-9d46-41b5-ec90-1a3b42b16103"
model = Sequential()
model.add(Conv2D(
128, kernel_size=(5, 5), activation='relu', input_shape=input_shape
))
model.add(Conv2D(64, (5, 5), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Conv2D(32, (5, 5), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(16, (5, 5), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(64, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 315} colab_type="code" id="ZxGKgm0AT49C" outputId="12f07b75-dd19-4927-c1a8-03ccff86492c"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="7KWUG2ximGku"
# ## [3] Convolution 7x7 kernel
# + [markdown] colab_type="text" id="vQsJ0fAQmKXA"
# ### [3.1] Conv (128-64-32) | MaxPool x2 | Dropout x3 | Dense (128-64) | Padding (same) | BN
# + colab={"base_uri": "https://localhost:8080/", "height": 1003} colab_type="code" id="yb2QCHmwUGqF" outputId="8b233867-8b1f-42d8-bc1b-4a92052adcaa"
model = Sequential()
model.add(Conv2D(
128, kernel_size=(7, 7), activation='relu', input_shape=input_shape,
padding='same'
))
model.add(Conv2D(64, (7, 7), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Conv2D(32, (7, 7), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(64, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="iCBpwhWxVbWl" outputId="54f4e85a-e540-49e9-a600-112f6fb70fdf"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="KBMC926JmXsA"
# ### [3.2] Conv (256-128-64) | MaxPool x2 | Dropout x3 | Dense (128-64) | Padding (same) | BN
# + colab={"base_uri": "https://localhost:8080/", "height": 1003} colab_type="code" id="P8D2_VPGVk8q" outputId="c8f60025-4a31-4f82-f2cf-fe476ee40567"
model = Sequential()
model.add(Conv2D(
256, kernel_size=(7, 7), activation='relu', input_shape=input_shape,
padding='same'
))
model.add(Conv2D(128, (7, 7), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Conv2D(64, (7, 7), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(64, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="byvanralX3lA" outputId="3f30c49a-93eb-4e12-b959-2cf4dc33424c"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab_type="text" id="Y3efzjZomiHZ"
# ### [3.3] Conv (256-128-64) | MaxPool x2 | Dropout x3 | Dense (256-128-64) | Padding (same) | BN
# + colab={"base_uri": "https://localhost:8080/", "height": 1071} colab_type="code" id="SlvL_ihHX4gB" outputId="1be284d0-3734-4b1c-c8a6-2842dc37aa62"
model = Sequential()
model.add(Conv2D(
256, kernel_size=(7, 7), activation='relu', input_shape=input_shape,
padding='same'
))
model.add(Conv2D(128, (7, 7), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(rate=0.75))
model.add(Conv2D(64, (7, 7), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(128, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(rate=0.5))
model.add(Dense(64, activation='relu'))
model.add(Dropout(rate=0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 313} colab_type="code" id="KzeYB5i4c6Xs" outputId="2f5783eb-9538-48eb-e9b4-5e29bbd29ac1"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
# list of epoch numbers
x = list(range(1,epochs+1))
vy = history.history['val_loss']
ty = history.history['loss']
plt_dynamic(fig, x, vy, ty, ax)
# + [markdown] colab={} colab_type="code" id="niK4jJ2lc7V3"
# ## [4] Conclusion
# -
from prettytable import PrettyTable
x = PrettyTable()
x.field_names = ["convolution kernel", "architecture", "Test Accuracy"]
x.add_row(["3x3", "Conv(32-64) | MaxPool | Dropout x2 | Dense", 0.9903])
x.add_row(["3x3", "Conv (64-32) | MaxPool | Dropout x2 | Dense", 0.9893])
x.add_row(["3x3", "Conv (64-32) | MaxPool | Dropout x2 | Dense | Padding (same)", 0.9888])
x.add_row(["3x3", "Conv (64-32) | MaxPool | Dropout x2 | Dense | Padding (same) | BN", 0.9896])
x.add_row(["3x3", "Conv (64-32) | MaxPool | Dropout x2 | Dense (256-128) | Padding (same) | BN", 0.9902])
x.add_row(["5x5", "Conv (128-64-32) | MaxPool | Dropout x2 | Dense", 0.9943])
x.add_row(["5x5", "Conv (128-64-32) | MaxPool x2 | Dropout x2 | Dense", 0.9948])
x.add_row(["5x5", "Conv (128-64-32) | MaxPool x2 | Dropout x3 | Dense | BN x2", 0.993])
x.add_row(["5x5", "Conv (128-64-32) | MaxPool x2 | Dropout x3 | Dense (128-64) | Padding (same) | BN", 0.9942])
x.add_row(["7x7", "Conv (128-64-32) | MaxPool x2 | Dropout x3 | Dense (128-64) | Padding (same) | BN", 0.9940])
x.add_row(["7x7", "Conv (256-128-64) | MaxPool x2 | Dropout x3 | Dense (128-64) | Padding (same) | BN", 0.9963])
x.add_row(["7x7", "Conv (256-128-64) | MaxPool x2 | Dropout x3 | Dense (256-128-64) | Padding (same) | BN", 0.9948])
print(x)
| 13.CNN_mnist/.ipynb_checkpoints/13_cnn_mnist-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 给定一个数字字符串S,例如S = “123456579”,我们可以将其拆分为类似于Fibonacci的序列[123,456,579]。
# 形式上,类似于斐波那契的序列是非负整数的列表F,使得:
# 1、0 <= F [i] <= 2^31-1(即,每个整数都适合32位有符号整数类型);
# 2、F.length> = 3;
# 3、F[i] + F[i+1] = F[i+2]的所有0 <= i <F.length-2。
# 另外,请注意,将字符串分割成段时,每个段都不能有多余的前导零,除了如果该部分本身是数字0。
# 返回从S拆分的任何类似于Fibonacci的序列,如果无法完成,则返回[]。
#
# Example 1:
# Input: "123456579"
# Output: [123,456,579] 123+456 = 579
#
# Example 2:
# Input: "11235813"
# Output: [1,1,2,3,5,8,13]
#
# Example 3:
# Input: "112358130"
# Output: []
# Explanation: The task is impossible.
#
# Example 4:
# Input: "0123"
# Output: []
# Explanation: Leading zeroes are not allowed, so "01", "2", "3" is not valid.
#
# Example 5:
# Input: "1101111"
# Output: [110, 1, 111]
# Explanation: The output [11, 0, 11, 11] would also be accepted.
# -
class Solution:
def splitIntoFibonacci(self, S: str):
self.res = []
self.dfs(S, [], 0)
return self.res
def dfs(self, S, temp, count):
if not S and count >= 3:
self.res = temp[:]
return
for step in range(1, min(len(S)+1, 11)):
val = S[:step]
if str(int(val)) == val and 0 <= int(val) <= pow(2, 31) - 1: # 防止前导 0 的出现
if count < 2:
temp.append(int(val))
self.dfs(S[step:], temp, count+1)
temp.pop()
elif temp[count-2] + temp[count-1] == int(val):
temp.append(int(val))
self.dfs(S[step:], temp, count+1)
temp.pop()
class Solution:
def splitIntoFibonacci(self, S: str):
for i in range(min(10, len(S))):
pass
S_ = "539834657215398346785398346991079669377161950407626991734534318677529701785098211336528511"
solution = Solution()
solution.splitIntoFibonacci(S_)
class Solution(object):
def splitIntoFibonacci(self, S):
for i in range(min(10, len(S))):
x = S[:i+1]
if x != '0' and x.startswith('0'):
break
a = int(x)
for j in range(i+1, min(i+10, len(S))):
y = S[i+1: j+1]
if y != '0' and y.startswith('0'): break
b = int(y)
fib = [a, b]
k = j + 1
while k < len(S):
nxt = fib[-1] + fib[-2]
nxtS = str(nxt)
if nxt <= 2**31 - 1 and S[k:].startswith(nxtS):
k += len(nxtS)
fib.append(nxt)
else:
break
else:
if len(fib) >= 3:
return fib
return []
| Back Tracking/0907/842. Split Array into Fibonacci Sequence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Install requirements
# !pip install tensorflow_addons numpy pandas tensorflow sklearn nltk spacy textblob gensim scipy seaborn matplotlib minio mlflow wordcloud boto3
# +
import os
cwd = os.getcwd()
cwd
# -
# +
import sys
# sys.path is a list of absolute path strings
sys.path.append('/opt/app-root/src/anz_ml_project/')
from src.loadingdata.read_dataset import ReadData
from src.features.build_features import BuildFeatures
from src.modules.build_model import BuildModel
from src.modules.train_model import MLflow, TrainModel
from src.modules.predict_model import Predictor,Transformer,DownloadArtifact
# from src.modules.predict_model import BuildModel
# from src.modules.train_model import BuildModel
# -
# # Load Libraries
# +
# import sys
# # sys.path is a list of absolute path strings
# sys.path.append('/opt/app-root/src/anz_ml_project/')
# from src.features.build_features import BuildFeatures
import os
import numpy as np
import pandas as pd
import tensorflow as tf
import tensorflow_addons as tfa
from sklearn.feature_extraction.text import TfidfVectorizer
import sklearn.feature_extraction.text as text
from sklearn import model_selection, preprocessing, linear_model, naive_bayes, metrics, svm
# from sklearn.naive_bayes import MultinomialNB
# from sklearn.linear_model import LogisticRegression
# from sklearn.ensemble import RandomForestClassifier
# from sklearn.svm import LinearSVC
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from textblob import TextBlob
from nltk.stem import PorterStemmer,SnowballStemmer
from textblob import Word
from sklearn.feature_extraction.text import CountVectorizer,TfidfVectorizer
from nltk.corpus import stopwords
from nltk.tokenize import sent_tokenize, word_tokenize
from nltk.tokenize.toktok import ToktokTokenizer
# from wordcloud import WordCloudfrom wordcloud import WordCloud
from io import StringIO
import string
import gensim
from gensim.models import Word2Vec
import itertools
import scipy
from scipy import spatial
import seaborn as sns
import matplotlib.pyplot as plt
import re
import nltk
import joblib
import mlflow
import warnings
from minio import Minio
import subprocess
import ipynbname
warnings.filterwarnings("ignore")
import absl.logging
absl.logging.set_verbosity(absl.logging.ERROR)
tokenizer = ToktokTokenizer()
# stopword_list = nltk.download('stopwords')
# -
# # Define a class to read the dataset
# # Define a class to preprocess the data and make them ready for modeling
# # Define a class for building the Deep learning based model for NLP
# # Define a class to configur MLFLOW
# # Define a class for training the model and tracking it with MLflow
# # Define classes for Deploy simulations
# # download artifacts for testing
# + [markdown] tags=[]
# # Initialize the config file for mlflow and Minio
# +
HOST = "http://mlflow:5500"
PROJECT_NAME = "NlpTc"
EXPERIMENT_NAME = "NlpLstm"
os.environ['MLFLOW_S3_ENDPOINT_URL']='http://minio-ml-workshop:9000'
os.environ['AWS_ACCESS_KEY_ID']='minio'
os.environ['AWS_SECRET_ACCESS_KEY']='minio123'
os.environ['AWS_REGION']='us-east-1'
os.environ['AWS_BUCKET_NAME']='raw-data-saeed'
# -
#
# ## Define a Function to read from Minio S3 Bucket
def get_s3_server():
minioClient = Minio('minio-ml-workshop:9000',
access_key='minio',
secret_key='minio123',
secure=False)
return minioClient
client = get_s3_server()
# ## SetUp MLFlow to track the model
mlflow = MLflow(mlflow, HOST,EXPERIMENT_NAME).SetUp_Mlflow()
# # Readinng the data
#
train_data, test_data, train_labels, test_labels,enc = ReadData(S3BucketName = "raw-data-saeed",FILE_NAME="data.csv").ReadDataFrameData()
joblib.dump(enc, 'labelencoder.pkl')
# +
# from src.features.build_features import BuildFeatures
train_data.shape
test_labels.shape
# -
# # Prepare data for modeling
BFCLASS = BuildFeatures(TRAIN_DATA=train_data,TEST_DATA=test_data,TRAIN_LABELS=train_labels,TEST_LABELS=test_labels, GloveData="glove.6B.50d.txt",EMBEDDING_DIM=50, WEIGHT_FLAG = False,MLFLOW_S3_ENDPOINT_URL = "minio-ml-workshop:9000",AWS_ACCESS_KEY_ID='minio',AWS_SECRET_ACCESS_KEY = 'minio123',SECURE = False)
train_data,test_data,train_labels , test_labels,word_index,tokenizer,MAX_SEQUENCE_LENGTH = BFCLASS.PreProcessingTextData()
joblib.dump(tokenizer, 'tokenizer.pkl')
# + [markdown] pycharm={"name": "#%% md\n"}
# # Deep Learning define, train and test model
#
# -
# ## Define the Model
model = BuildModel(WORD_INDEX=word_index, MAX_SEQUENCE_LENGTH=MAX_SEQUENCE_LENGTH, EMWEIGHTS=[]).SetupModel()
# ## Train the Model
model, history= TrainModel(model, tokenizer, enc,train_data, train_labels,test_data, test_labels,HOST, EXPERIMENT_NAME, BATCH_SIZE=64,EPOCHS=1).ModelTraining()
# ### Plot the training and testing Loss
fig1 = plt.figure()
plt.plot(history.history['loss'],'r',linewidth=3.0)
plt.plot(history.history['val_loss'],'b',linewidth=3.0)
plt.legend(['Training loss', 'Validation Loss'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('Loss',fontsize=16)
plt.title('Loss Curves :RNN - LSTM',fontsize=16)
plt.show()
# #### Plot the training and testing Accuracy
fig1 = plt.figure()
plt.plot(history.history['acc'],'r',linewidth=3.0)
plt.plot(history.history['val_acc'],'b',linewidth=3.0)
plt.legend(['Training acc', 'Validation acc'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('Accuracy',fontsize=16)
plt.title('Accuracy Curves :RNN - LSTM',fontsize=16)
plt.show()
# ## Test the model
#predictions on test data
predicted=model.predict(test_data)
predicted
#model evaluation
import sklearn
from sklearn.metrics import precision_recall_fscore_support as score
precision, recall, fscore, support = score(test_labels, predicted.round())
print('precision: \n{}'.format(precision))
print('recall: \n{}'.format(recall))
print('fscore: \n{}'.format(fscore))
print('support: \n{}'.format(support))
print("############################")
# +
from sklearn.metrics import confusion_matrix
conf_mat = confusion_matrix(test_labels.argmax(axis=1), predicted.argmax(axis=1))
fig, ax = plt.subplots(figsize=(8,6))
sns.heatmap(conf_mat, annot=True, fmt="d", cmap="BuPu",xticklabels=enc.classes_,yticklabels=enc.classes_)
plt.ylabel('Actual')
plt.xlabel('Predicted')
plt.show()
# -
# ### Download the artifacts
# +
DownloadArtifact(mlflow, MODEL_NAME='lstmt1', MODEL_VRSION='1').download_artifacts()
# -
# ### Test with Actual data
# #### Define a sample
sample_data = {"data":
{
"names":
[
"Debt collection"
],
"ndarray": ["could longer pay enormous charge hired company nl take either nothing pay day loan company accept term get several letter week threatened take civil action get check"]
}
}
# #### Transform the data
#
# + tags=[]
ready_data = Transformer().transform_input(sample_data,"name","meta")
# -
# ### Test the prediction
output = Predictor().predict(ready_data,"features")
output = Predictor().predict(ready_data,ready_data)
model.predict(ready_data)
from sklearn.metrics import classification_report
print(classification_report(test_labels, predicted.round(),target_names=enc.classes_))
# After hours of training we get good results with LSTM(type of recurrent neural network) compared to CNN. From the learning curves it is clear the model needs to be tuned for overfitting by selecting hyperparameters such as no of epochs via early stopping and dropout for regularization.
#
# We could further improve our final result by ensembling our xgboost and Neural network models by using Logistic Regression as our base model.
#
#
| notebooks/text_classification_notebooks/text_classification_notebooks/NLPConsumerComplaintsClassificationDL-LSTM-clean-v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solução de sistemas lineares através da eliminação de Gauss
# ## License
#
# All content can be freely used and adapted under the terms of the
# [Creative Commons Attribution 4.0 International License](http://creativecommons.org/licenses/by/4.0/).
#
# 
# ## Imports
#
# Coloque **todos** os `import` na célula abaixo. Não se esqueça do `%matplotlib inline` para que os gráficos apareçam no notebook.
import numpy as np
# %matplotlib inline
# ## Tarefa
#
# Reduza o sistema linear $\bar{\bar{A}}\bar{x} = \bar{b}$ para sua forma escalonada (row echelon form).
N = 11 # Número de linhas
M = 11 # Número de colunas
sistema = [[25, 108, 58, 102, 61, 24, 39, 40, 67, 11, 5],
[51, 28, 17, 98, 105, 43, 86, 2, 71, 20, 95],
[56, 70, 109, 45, 117, 52, 97, 92, 1, 42, 78],
[59, 6, 50, 37, 120, 118, 104, 64, 53, 100, 66],
[82, 74, 68, 80, 16, 115, 63, 103, 60, 30, 93],
[79, 113, 65, 72, 22, 35, 101, 10, 83, 89, 85],
[34, 57, 90, 9, 14, 47, 76, 44, 26, 48, 114],
[41, 7, 21, 75, 33, 49, 32, 119, 46, 0, 87],
[91, 77, 55, 18, 106, 54, 116, 12, 13, 31, 38],
[27, 81, 29, 107, 84, 15, 3, 112, 88, 69, 19],
[110, 96, 23, 73, 36, 8, 99, 4, 94, 111, 62]]
lado_direito = [3323, 2869, 4085, 3211, 4181, 3831, 2699, 2891, 3188, 3586, 3598]
list(range(2, 5, 1))
# +
for k in range(0, N - 1, 1):
for i in range(k + 1, M, 1):
for j in range(k, M, 1):
sistema[i][j] = sistema[i][j] - ((sistema[k][j]*sistema[i][j])/sistema[k][j])
# -
a = [[1, 2], [3, 4]]
a[0][0]
# ### Resultado esperado
#
# Quando executada, a célula abaixo deverá imprimir exatamente:
#
# A | b
# 25.000 108.000 58.000 102.000 61.000 24.000 39.000 40.000 67.000 11.000 5.000 | 3323.000
# 0.000 -192.320 -101.320 -110.080 -19.440 -5.960 6.440 -79.600 -65.680 -2.440 84.800 | -3909.920
# -0.000 0.000 69.653 -85.077 -2.262 3.568 3.883 73.557 -90.367 19.541 -9.005 | 136.662
# 0.000 0.000 0.000 -7.233 2.634 66.807 1.160 25.893 37.269 64.787 -49.820 | 341.724
# 0.000 0.000 0.000 0.000 -177.918 -539.479 -85.844 -165.046 -356.419 -575.159 391.184 | -4053.756
# 0.000 0.000 0.000 0.000 -0.000 -539.656 43.030 -267.157 -271.025 -378.300 357.879 | -3235.329
# 0.000 0.000 0.000 0.000 0.000 0.000 55.468 -69.824 59.860 57.791 78.381 | 141.706
# 0.000 0.000 0.000 0.000 0.000 0.000 0.000 70.381 24.471 60.345 -29.950 | 741.794
# 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 -16.843 -126.224 62.584 | 57.797
# 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 -291.983 268.919 | 537.838
# -0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 -71.547 | -143.094
#
print(' A | b ')
for i in range(N):
for j in range(M):
print('{:8.3f} '.format(sistema[i][j]), end='')
print('| {:8.3f}'.format(lado_direito[i]))
# ## Tarefa
#
# Utilize a matriz escalonada acima para encontrar a solução do sistema através da substituição retrocedida (back substitution). Coloque a solução em uma lista chamada `solucao`.
#
# **Dica**:
#
# * Para criar uma lista com um mesmo valor repetido N vezes: `[numero]*N`. Por exemplo, `[0]*4` resulta em `[0, 0, 0, 0]`.
# ### Resultado esperado
#
# Quando executada, a célula abaixo deverá imprimir exatamente:
#
# 6.000000 9.000000 5.000000 7.000000 3.000000 1.000000 8.000000 10.000000 4.000000 0.000000 2.000000
for i in range(N):
print('{:8.6f} '.format(solucao[i]), end='')
# ## Tarefa Bônus
# Sua tarefa será aplicar a eliminação de Gauss para resolver o sistema linear gerado abaixo.
# Para tornar a vida mais interessante, vamos gerar um sistema linear aleatório
N, M = 13, 13
A = np.random.random_integers(10, 50, size=(N, M)).tolist()
for i in range(N):
A[i][i] = 0
b = np.dot(A, np.random.random_integers(1, 20, size=N)).tolist()
print(' A | b ')
for i in range(N):
for j in range(M):
print('{:7.1f} '.format(A[i][j]), end='')
print('| {:7.1f}'.format(b[i]))
# Porém, há um problema. Você não poderá fazer isso diretamente pois $A_{ii} == 0$ (a diagonal da matriz é zero). Logo, não há como dividir pelo elemento pivo e não há como aplicar a eliminação de Gauss nesse sistema. **Experimente aplicar a eliminação convencional que você implementou acima nesse sistema e veja o que acontece.**
#
#
# ### Condensação pivotal (partial pivoting)
#
# Existe um método para contornar esse problema chamado **condensação pivotal** (partial pivoting). A ideia é que podemos trocar duas linhas do sistema de lugar sem alterar a solução. Logo, podemos tirar os elementos 0 da diagonal da matriz trocando por outra linha sem o 0 na posição do pivo. Um jeito melhor ainda de fazer isso é colocando a linha com o **maior elemento** possível na posição do pivo (veja [aqui uma ótima explicação disso](http://www.math.iitb.ac.in/~neela/partialpivot.pdf)).
#
# O método da eliminação de Gauss com condensação pivotal é:
#
# Para cada linha k do sistema:
# Ache a linha de A com o maior elemento na coluna k (só precisa considerar de k para baixo)
# Troque a linha k com a linha determinada acima (não esqueça de trocar o lado direito também)
# pivo = A[k][k]
# Escalonamento para zerar a coluna k (igual ao feita na eliminação convencional)
#
#
# Resolva o sistema linear definido acima utilizando eliminação de Gauss com condensação pivotal. Coloque a solução em uma variável chamada `solucao`.
# ### Resultado esperado
#
# Quando executada, a célula abaixo deverá imprimir `True`.
print(np.allclose(np.linalg.solve(A, b), solucao))
| sistemas-lineares.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:571]
# language: python
# name: conda-env-571-py
# ---
# ### Import modules
# +
import numpy as np
import pandas as pd
import torch
import torch.nn as nn
from sklearn.model_selection import train_test_split, cross_validate, cross_val_score
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.compose import ColumnTransformer, make_column_transformer
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.pipeline import Pipeline, make_pipeline
from sklearn.metrics import make_scorer
from sklearn.dummy import DummyRegressor
from xgboost import XGBRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.linear_model import Ridge
import statsmodels.api as sm
import statsmodels.api as sm
import matplotlib.pyplot as plt
import altair as alt
# -
# Save a vega-lite spec and a PNG blob for each plot in the notebook
alt.renderers.enable('mimetype')
# Handle large data sets without embedding them in the notebook
alt.data_transformers.enable('data_server')
# ## Utility Functions and Classes
# Function to plot the raw data
def plot_data(data, x_axis, y_axis):
"""
Plots line chart of the time series data (y-axis) against time (x-axis)
Parameters
----------
data: pd.DataFrame
input dataframe
x_axis: str
the name of the time(datetime) column
y_axis: str
the name of the time series data column
Returns
----------
lines: altair.vegalite.v4.api.Chart
altair chart object
"""
data = data.reset_index()
lines = alt.Chart(data).mark_line().encode(
x=alt.X(x_axis),
y=alt.Y(y_axis)
).properties(
width=700,
height=250
)
return lines
# Generate lags
def generate_lags(df, num_lags):
"""
Creates new features which are lags of the target variable
Parameters
----------
df: pd.DataFrame
input dataframe
num_lags: int
the number of lags
Returns
----------
df_tmp: pd.DataFrame
dataframe with lagged features created
"""
df_tmp = df.copy()
for n in range(1, num_lags+1):
df_tmp[f'lag{n}'] = df_tmp['target'].shift(n)
df_tmp = df_tmp.iloc[num_lags:]
return df_tmp
# Custom transformer for automated feature extraction
class UniveriateTimeSeriesAddFeatures(BaseEstimator, TransformerMixin):
def __init__(self, num_feats=4):
self.num_feats = num_feats
def __extract_features(self, X):
"""
Creates new features from timestamp
Parameters
----------
X: pd.DataFrame
input dataframe
Returns
----------
df_features: pd.DataFrame
dataframe with new features created
"""
df_features = X.copy()
df_features = (
df_features
.assign(hour = df_features.index.hour)
.assign(day = df_features.index.day)
.assign(month = df_features.index.month)
.assign(day_of_week = df_features.index.dayofweek)
# .assign(week_of_year = df.index.isocalendar().week)
)
df_features = df_features.reset_index()
return df_features
def fit(self, X, y=None):
return self
def transform(self, X, y=None):
df_features = X.copy()
df_features = self.__extract_features(df_features)
df_features = df_features.iloc[:,1:]
return df_features
# Function to collate results
def mean_std_cross_val_scores(model, X_train, y_train, **kwargs):
"""
Returns mean and std of cross validation
Parameters
----------
model :
scikit-learn model
X_train : numpy array or pandas DataFrame
X in the training data
y_train :
y in the training data
Returns
----------
pandas Series with mean scores from cross_validation
"""
scores = cross_validate(model, X_train, y_train, **kwargs)
mean_scores = pd.DataFrame(scores).mean()
std_scores = pd.DataFrame(scores).std()
out_col = []
for i in range(len(mean_scores)):
out_col.append((f"%0.3f (+/- %0.3f)" % (mean_scores[i], std_scores[i])))
return pd.Series(data=out_col, index=mean_scores.index)
# Function to calculate MAPE
def mape(true, pred):
return 100.0 * np.mean(np.abs((pred - true) / true))
# ## Read the data
# +
# Read the data
df = pd.read_csv('data/PJME_hourly.csv')
# Remove spaces from column names
df.columns = df.columns.str.replace(' ', '')
# Convert Datetime column to datetime data type
df['Datetime'] = pd.to_datetime(df['Datetime'])
# Set index to datetime column
df = df.set_index('Datetime')
# Sort the index to ascending order
df = df.sort_index()
# Rename the PJME_MW column to target column
df = df.rename(columns={'PJME_MW': 'target'})
# Check head of the resulting dataframe
df.head()
# -
# ## Split into train and test
df_train, df_test = train_test_split(df, test_size=0.2, shuffle=False, random_state=123)
print(df_train.shape)
print(df_test.shape)
# ## Visualize the data - Basic EDA
plot_data(df_train, x_axis='Datetime', y_axis='target')
# ## ACF and PACF plots
# +
ts = df_train.target
fig,ax = plt.subplots(2,1,figsize=(8,8))
fig = sm.graphics.tsa.plot_acf(ts, lags=100, ax=ax[0])
fig = sm.graphics.tsa.plot_pacf(ts, lags=100, ax=ax[1])
plt.show()
# -
# ## Separate train and test
# #### Total Lags = 10
# Total lags defined
tot_lags = 10
df_train = generate_lags(df=df_train, num_lags=tot_lags)
df_test = generate_lags(df=df_test, num_lags=tot_lags)
df_test.head()
# +
X_train, y_train = (
df_train.drop(columns=['target']),
df_train['target']
)
X_test, y_test = (
df_test.drop(columns=['target']),
df_test['target']
)
# Print shapes of the resulting train and test data sets
print(f'Shape of X_train and y_train: {X_train.shape, y_train.shape}')
print(f'Shape of X_test and y_test: {X_test.shape, y_test.shape}')
# -
# ## Feature Engineering Using Pipeline Transformations
preprocessor = Pipeline(
[
('add_lag_step', UniveriateTimeSeriesAddFeatures()),
('stdscaler', StandardScaler())
]
)
# ## Models
# +
# make a scorer function that we can pass into cross-validation
mape_scorer = make_scorer(mape, greater_is_better=False)
scoring_metrics = {
"neg RMSE": "neg_root_mean_squared_error",
"r2": "r2",
"mape": mape_scorer,
}
# -
results = {}
# #### Dummy Regressor
# +
dummy = DummyRegressor()
results["Dummy"] = mean_std_cross_val_scores(
dummy, X_train, y_train, return_train_score=True, scoring=scoring_metrics
)
pd.DataFrame(results)
# -
# #### Other Models
# +
pipe_ridge = make_pipeline(preprocessor, Ridge(random_state=123))
pipe_xgb = make_pipeline(preprocessor, XGBRegressor(random_state=123))
pipe_rf = make_pipeline(preprocessor, RandomForestRegressor(random_state=123))
models = {
'ridge': pipe_ridge,
'xgboost': pipe_xgb,
'rf_regressor': pipe_rf
}
# +
for (name, model) in models.items():
results[name] = mean_std_cross_val_scores(
model, X_train, y_train, return_train_score=True, scoring=scoring_metrics
)
pd.DataFrame(results)
# -
# #### Best Model (Example of XGBRegressor provided below)
pipe_xgb.fit(X_train, y_train);
preds = pipe_xgb.predict(X_test)
actual_vs_pred_df = y_test.reset_index()
actual_vs_pred_df['predicted'] = preds
actual_vs_pred_df.head()
# +
actual_vs_pred_df_long = pd.melt(actual_vs_pred_df, id_vars='Datetime')
actual_vs_pred_df_long.tail()
# -
# Visualize the actual vs predicted chart for the last 2 days of the test data period
alt.Chart(actual_vs_pred_df_long[actual_vs_pred_df_long.Datetime > '2018-07-24'],
title='Actual vs Predicted on Test Data').mark_line().encode(
x=alt.X('Datetime', type='temporal', title='Datetime'),
y=alt.Y('value', type='quantitative', title='Actual vs Predicted'),
color=alt.Color('variable', title='Type')
).properties(
width=700
).interactive()
print(f'MAPE for test set: {mape(actual_vs_pred_df["target"], actual_vs_pred_df["predicted"]):.2f}%')
# #### Total Lags = 50
# +
df_train, df_test = train_test_split(df, test_size=0.2, shuffle=False, random_state=123)
# Total lags defined
tot_lags = 50
df_train = generate_lags(df=df_train, num_lags=tot_lags)
df_test = generate_lags(df=df_test, num_lags=tot_lags)
X_train, y_train = (
df_train.drop(columns=['target']),
df_train['target']
)
X_test, y_test = (
df_test.drop(columns=['target']),
df_test['target']
)
# Print shapes of the resulting train and test data sets
print(f'Shape of X_train and y_train: {X_train.shape, y_train.shape}')
print(f'Shape of X_test and y_test: {X_test.shape, y_test.shape}')
# -
preprocessor = Pipeline(
[
('add_lag_step', UniveriateTimeSeriesAddFeatures()),
('stdscaler', StandardScaler())
]
)
results = {}
# +
dummy = DummyRegressor()
results["Dummy"] = mean_std_cross_val_scores(
dummy, X_train, y_train, return_train_score=True, scoring=scoring_metrics
)
pd.DataFrame(results)
# +
pipe_ridge = make_pipeline(preprocessor, Ridge(random_state=123))
pipe_xgb = make_pipeline(preprocessor, XGBRegressor(random_state=123))
pipe_rf = make_pipeline(preprocessor, RandomForestRegressor(random_state=123))
models = {
'ridge': pipe_ridge,
'xgboost': pipe_xgb,
'rf_regressor': pipe_rf
}
# +
for (name, model) in models.items():
results[name] = mean_std_cross_val_scores(
model, X_train, y_train, return_train_score=True, scoring=scoring_metrics
)
pd.DataFrame(results)
# +
pipe_xgb.fit(X_train, y_train);
preds = pipe_xgb.predict(X_test)
actual_vs_pred_df = y_test.reset_index()
actual_vs_pred_df['predicted'] = preds
# actual_vs_pred_df.head()
actual_vs_pred_df_long = pd.melt(actual_vs_pred_df, id_vars='Datetime')
# actual_vs_pred_df_long.tail()
# -
# Visualize the actual vs predicted chart for the last 2 days of the test data period
alt.Chart(actual_vs_pred_df_long[actual_vs_pred_df_long.Datetime > '2018-07-24'],
title='Actual vs Predicted on Test Data').mark_line().encode(
x=alt.X('Datetime', type='temporal', title='Datetime'),
y=alt.Y('value', type='quantitative', title='Actual vs Predicted'),
color=alt.Color('variable', title='Type')
).properties(
width=700
).interactive()
print(f'MAPE for test set: {mape(actual_vs_pred_df["target"], actual_vs_pred_df["predicted"]):.2f}%')
# #### Total Lags = 5
# +
df_train, df_test = train_test_split(df, test_size=0.2, shuffle=False, random_state=123)
# Total lags defined
tot_lags = 5
df_train = generate_lags(df=df_train, num_lags=tot_lags)
df_test = generate_lags(df=df_test, num_lags=tot_lags)
X_train, y_train = (
df_train.drop(columns=['target']),
df_train['target']
)
X_test, y_test = (
df_test.drop(columns=['target']),
df_test['target']
)
# Print shapes of the resulting train and test data sets
print(f'Shape of X_train and y_train: {X_train.shape, y_train.shape}')
print(f'Shape of X_test and y_test: {X_test.shape, y_test.shape}')
# -
preprocessor = Pipeline(
[
('add_lag_step', UniveriateTimeSeriesAddFeatures()),
('stdscaler', StandardScaler())
]
)
results = {}
# +
dummy = DummyRegressor()
results["Dummy"] = mean_std_cross_val_scores(
dummy, X_train, y_train, return_train_score=True, scoring=scoring_metrics
)
pd.DataFrame(results)
# +
pipe_ridge = make_pipeline(preprocessor, Ridge(random_state=123))
pipe_xgb = make_pipeline(preprocessor, XGBRegressor(random_state=123))
pipe_rf = make_pipeline(preprocessor, RandomForestRegressor(random_state=123))
models = {
'ridge': pipe_ridge,
'xgboost': pipe_xgb,
'rf_regressor': pipe_rf
}
# +
for (name, model) in models.items():
results[name] = mean_std_cross_val_scores(
model, X_train, y_train, return_train_score=True, scoring=scoring_metrics
)
pd.DataFrame(results)
# +
pipe_xgb.fit(X_train, y_train);
preds = pipe_xgb.predict(X_test)
actual_vs_pred_df = y_test.reset_index()
actual_vs_pred_df['predicted'] = preds
# actual_vs_pred_df.head()
actual_vs_pred_df_long = pd.melt(actual_vs_pred_df, id_vars='Datetime')
# actual_vs_pred_df_long.tail()
# -
# Visualize the actual vs predicted chart for the last 2 days of the test data period
alt.Chart(actual_vs_pred_df_long[actual_vs_pred_df_long.Datetime > '2018-07-24'],
title='Actual vs Predicted on Test Data').mark_line().encode(
x=alt.X('Datetime', type='temporal', title='Datetime'),
y=alt.Y('value', type='quantitative', title='Actual vs Predicted'),
color=alt.Color('variable', title='Type')
).properties(
width=700
).interactive()
print(f'MAPE for test set: {mape(actual_vs_pred_df["target"], actual_vs_pred_df["predicted"]):.2f}%')
# #### Total Lags = 30
# +
df_train, df_test = train_test_split(df, test_size=0.2, shuffle=False, random_state=123)
# Total lags defined
tot_lags = 30
df_train = generate_lags(df=df_train, num_lags=tot_lags)
df_test = generate_lags(df=df_test, num_lags=tot_lags)
X_train, y_train = (
df_train.drop(columns=['target']),
df_train['target']
)
X_test, y_test = (
df_test.drop(columns=['target']),
df_test['target']
)
# Print shapes of the resulting train and test data sets
print(f'Shape of X_train and y_train: {X_train.shape, y_train.shape}')
print(f'Shape of X_test and y_test: {X_test.shape, y_test.shape}')
# -
preprocessor = Pipeline(
[
('add_lag_step', UniveriateTimeSeriesAddFeatures()),
('stdscaler', StandardScaler())
]
)
results = {}
# +
dummy = DummyRegressor()
results["Dummy"] = mean_std_cross_val_scores(
dummy, X_train, y_train, return_train_score=True, scoring=scoring_metrics
)
pd.DataFrame(results)
# +
pipe_ridge = make_pipeline(preprocessor, Ridge(random_state=123))
pipe_xgb = make_pipeline(preprocessor, XGBRegressor(random_state=123))
pipe_rf = make_pipeline(preprocessor, RandomForestRegressor(random_state=123))
models = {
'ridge': pipe_ridge,
'xgboost': pipe_xgb,
'rf_regressor': pipe_rf
}
# +
for (name, model) in models.items():
results[name] = mean_std_cross_val_scores(
model, X_train, y_train, return_train_score=True, scoring=scoring_metrics
)
pd.DataFrame(results)
# +
pipe_xgb.fit(X_train, y_train);
preds = pipe_xgb.predict(X_test)
actual_vs_pred_df = y_test.reset_index()
actual_vs_pred_df['predicted'] = preds
# actual_vs_pred_df.head()
actual_vs_pred_df_long = pd.melt(actual_vs_pred_df, id_vars='Datetime')
# actual_vs_pred_df_long.tail()
# -
# Visualize the actual vs predicted chart for the last 2 days of the test data period
alt.Chart(actual_vs_pred_df_long[actual_vs_pred_df_long.Datetime > '2018-07-24'],
title='Actual vs Predicted on Test Data').mark_line().encode(
x=alt.X('Datetime', type='temporal', title='Datetime'),
y=alt.Y('value', type='quantitative', title='Actual vs Predicted'),
color=alt.Color('variable', title='Type')
).properties(
width=700
).interactive()
print(f'MAPE for test set: {mape(actual_vs_pred_df["target"], actual_vs_pred_df["predicted"]):.2f}%')
| timeseries_using_lags.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/python
import pickle
import sys
import matplotlib.pyplot
sys.path.append("../tools/")
from feature_format import featureFormat, targetFeatureSplit
### read in data dictionary, convert to numpy array
data_dict = pickle.load( open("../final_project/final_project_dataset.pkl", "rb") )
data_dict.pop( 'TOTAL', 0 )
features = ["salary", "bonus"]
data = featureFormat(data_dict, features)
#print(data_dict)
### your code below
for point in data:
salary = point[0]
bonus = point[1]
matplotlib.pyplot.scatter( salary, bonus )
matplotlib.pyplot.xlabel("salary")
matplotlib.pyplot.ylabel("bonus")
matplotlib.pyplot.show()
| outliers/Outliers.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# ```
# --- Day 7: Amplification Circuit ---
#
# Based on the navigational maps, you're going to need to send more power to your ship's thrusters to reach Santa in time. To do this, you'll need to configure a series of amplifiers already installed on the ship.
#
# There are five amplifiers connected in series; each one receives an input signal and produces an output signal. They are connected such that the first amplifier's output leads to the second amplifier's input, the second amplifier's output leads to the third amplifier's input, and so on. The first amplifier's input value is 0, and the last amplifier's output leads to your ship's thrusters.
#
# O-------O O-------O O-------O O-------O O-------O
# 0 ->| Amp A |->| Amp B |->| Amp C |->| Amp D |->| Amp E |-> (to thrusters)
# O-------O O-------O O-------O O-------O O-------O
# The Elves have sent you some Amplifier Controller Software (your puzzle input), a program that should run on your existing Intcode computer. Each amplifier will need to run a copy of the program.
#
# When a copy of the program starts running on an amplifier, it will first use an input instruction to ask the amplifier for its current phase setting (an integer from 0 to 4). Each phase setting is used exactly once, but the Elves can't remember which amplifier needs which phase setting.
#
# The program will then call another input instruction to get the amplifier's input signal, compute the correct output signal, and supply it back to the amplifier with an output instruction. (If the amplifier has not yet received an input signal, it waits until one arrives.)
#
# Your job is to find the largest output signal that can be sent to the thrusters by trying every possible combination of phase settings on the amplifiers. Make sure that memory is not shared or reused between copies of the program.
#
# For example, suppose you want to try the phase setting sequence 3,1,2,4,0, which would mean setting amplifier A to phase setting 3, amplifier B to setting 1, C to 2, D to 4, and E to 0. Then, you could determine the output signal that gets sent from amplifier E to the thrusters with the following steps:
#
# Start the copy of the amplifier controller software that will run on amplifier A. At its first input instruction, provide it the amplifier's phase setting, 3. At its second input instruction, provide it the input signal, 0. After some calculations, it will use an output instruction to indicate the amplifier's output signal.
# Start the software for amplifier B. Provide it the phase setting (1) and then whatever output signal was produced from amplifier A. It will then produce a new output signal destined for amplifier C.
# Start the software for amplifier C, provide the phase setting (2) and the value from amplifier B, then collect its output signal.
# Run amplifier D's software, provide the phase setting (4) and input value, and collect its output signal.
# Run amplifier E's software, provide the phase setting (0) and input value, and collect its output signal.
# The final output signal from amplifier E would be sent to the thrusters. However, this phase setting sequence may not have been the best one; another sequence might have sent a higher signal to the thrusters.
#
# Here are some example programs:
#
# Max thruster signal 43210 (from phase setting sequence 4,3,2,1,0):
#
# 3,15,3,16,1002,16,10,16,1,16,15,15,4,15,99,0,0
# Max thruster signal 54321 (from phase setting sequence 0,1,2,3,4):
#
# 3,23,3,24,1002,24,10,24,1002,23,-1,23,
# 101,5,23,23,1,24,23,23,4,23,99,0,0
# Max thruster signal 65210 (from phase setting sequence 1,0,4,3,2):
#
# 3,31,3,32,1002,32,10,32,1001,31,-2,31,1007,31,0,33,
# 1002,33,7,33,1,33,31,31,1,32,31,31,4,31,99,0,0,0
# Try every combination of phase settings on the amplifiers. What is the highest signal that can be sent to the thrusters?
# ```
# +
pos(p) = p + 1;
ref(a,p) = a[pos(a[pos(p)])];
@enum Op plus=1 mul=2 stor=3 out=4 halt=99
@enum Mode posi=0 imm=1
struct Operation
op::Op
mode1::Mode
mode2::Mode
mode3::Mode
end
getOps(str::String) = getOps(parse(Int,str))
function getOps(num)
op = Op(num % 100)
mode1 = Mode((num ÷ 100) % 10)
mode2 = Mode((num ÷ 1000) % 10)
mode3 = Mode((num ÷ 10000) % 10)
Operation(op,mode1,mode2,mode3)
end
function op1(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
prog[pos(prog[pos(p+3)])] = p1 + p2
p + 4
end
function op2(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
prog[pos(prog[pos(p+3)])] = p1 * p2
p + 4
end
function op3(operation,p,prog)
# println("entering stor ","p ",p," prog ",prog, " input ",input)
p1 = prog[pos(p+1)]
#println("storage position",p1)
prog[pos(p1)] = input
#println("exiting stor")
p + 2
end
function op4(operation,p,prog,input)
output = []
#println("entering out")
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
#println("output:",p1)
push!(output,p1)
#println("exiting out $output")
(p + 2,output)
end
function execute(op::Operation, p, prog)
if op.op == plus
return op1(op,p,prog)
end
if op.op == mul
return op2(op,p,prog)
end
if op.op == stor
return op3(op,p,prog,1)
end
if op.op == out
return op4(op,p,prog)
end
error("panic executing unknown op")
end
function execute(prog)
p = 0
op = getOps(prog[pos(p)])
#println("op: ", op)
while op.op != halt
#println(prog)
p = execute(op,p,prog)
#println("instruction: ",p, " -> ",prog[pos(p)])
op = getOps(prog[pos(p)])
end
prog
end;
println(execute([1,9,10,3,2,3,11,0,99,30,40,50]) == [3500,9,10,70,2,3,11,0,99,30,40,50])
println(execute([1,0,0,0,99]) == [2,0,0,0,99])
println(execute([2,3,0,3,99]) == [2,3,0,6,99])
println(execute([2,4,4,5,99,0]) == [2,4,4,5,99,9801])
println(execute([1,1,1,4,99,5,6,0,99]) == [ 30,1,1,4,2,5,6,0,99])
# Opcode 5 is jump-if-true:
# if the first parameter is non-zero,
# it sets the instruction pointer to the value from the second parameter.
# Otherwise, it does nothing.
function op5(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
if p1 != 0
return p2
else
return p + 3
end
end
# Opcode 6 is jump-if-false:
# if the first parameter is zero,
# it sets the instruction pointer to the value from the second parameter. Otherwise, it does nothing.
function op6(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
if p1 == 0
return p2
else
return p + 3
end
end
# Opcode 7 is less than:
# if the first parameter is less than the second parameter,
# it stores 1 in the position given by the third parameter. Otherwise, it stores 0.
function op7(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
if p1 < p2
prog[pos(prog[pos(p+3)])] = 1
else
prog[pos(prog[pos(p+3)])] = 0
end
p + 4
end
# Opcode 8 is equals:
# if the first parameter is equal to the second parameter,
# it stores 1 in the position given by the third parameter. Otherwise, it stores 0.
function op8(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
if p1 == p2
prog[pos(prog[pos(p+3)])] = 1
else
prog[pos(prog[pos(p+3)])] = 0
end
p + 4
end
@enum Op plus=1 mul=2 stor=3 out=4 jnz=5 jez=6 lt=7 eq=8 halt=99
@enum Mode posi=0 imm=1
struct Operation
op::Op
mode1::Mode
mode2::Mode
mode3::Mode
end
function execute(op::Operation, p, prog, input,output)
if op.op == plus
return (op1(op,p,prog),input,output)
end
if op.op == mul
return (op2(op,p,prog),input,output)
end
if op.op == stor
#println(length(input))
next = input[1]
input=input[2:end]
#println(next)
#println(length(input))
return (op3(op,p,prog,next),input,output)
end
if op.op == out
(p1,output) = op4(op,p,prog,output)
#println(output)
return (p1,input,output)
end
if op.op == jnz
return (op5(op,p,prog),input,output)
end
if op.op == jez
return (op6(op,p,prog),input,output)
end
if op.op == lt
return (op7(op,p,prog),input,output)
end
if op.op == eq
return (op8(op,p,prog),input,output)
end
error("panic executing unknown op")
end
function execute(prog,p,input,output)
op = getOps(prog[pos(p)])
#println("op: ", op)
while op.op != halt
#println(prog)
(p,input,output) = execute(op,p,prog,input,output)
#println("instruction: ",p, " -> ",prog[pos(p)])
op = getOps(prog[pos(p)])
end
(prog,p,output)
end
# The above example program uses an input instruction to ask for a single number.
# The program will then
# output 999 if the input value is below 8,
# output 1000 if the input value is equal to 8,
# or output 1001 if the input value is greater than 8.
execute(
[3,21,1008,21,8,20,1005,20,22,107,8,21,20,1006,20,31,
1106,0,36,98,0,0,1002,21,125,20,4,20,1105,1,46,104,
999,1105,1,46,1101,1000,1,20,4,20,1105,1,46,98,99]
,0,[-8],[]);
execute(
[3,21,1008,21,8,20,1005,20,22,107,8,21,20,1006,20,31,
1106,0,36,98,0,0,1002,21,125,20,4,20,1105,1,46,104,
999,1105,1,46,1101,1000,1,20,4,20,1105,1,46,98,99]
,0,[8]);
a = execute(
[3,21,1008,21,8,20,1005,20,22,107,8,21,20,1006,20,31,
1106,0,36,98,0,0,1002,21,125,20,4,20,1105,1,46,104,
999,1105,1,46,1101,1000,1,20,4,20,1105,1,46,98,99]
,0,[16],[]);
#example usage
#using DelimitedFiles
#prog = readdlm("input.txt", '\t', Int, ',')
#execute(prog,5);
a
# -
using DelimitedFiles
get_amp() = readdlm("input.txt", '\t', Int, ',')
typeof(get_amp())
# Max thruster signal 43210 (from phase setting sequence 4,3,2,1,0):
function circuit(a,prog)
(amp1,p,output) = execute(prog,0,[a[1],0],[]);
(amp2,p,output) = execute(prog,0,[a[2],output[1]],[]);
(amp3,p,output) = execute(prog,0,[a[3],output[1]],[]);
(amp4,p,output) = execute(prog,0,[a[4],output[1]],[]);
(amp5,p,output) = execute(prog,0,[a[5],output[1]],[]);
output[1]
end
test_amp() = [3,15,3,16,1002,16,10,16,1,16,15,15,4,15,99,0,0]
circuit([4,3,2,1,0],test_amp())
#65210
test_amp() = [3,31,3,32,1002,32,10,32,1001,31,-2,31,1007,31,0,33,1002,33,7,33,1,33,31,31,1,32,31,31,4,31,99,0,0,0]
circuit([1,0,4,3,2],test_amp())
import Pkg; Pkg.add("Combinatorics")
using Combinatorics
all_inputs = permutations([0,1,2,3,4]) |> collect;
all_values = []
for x ∈ all_inputs
y = circuit(x,test_amp())
push!(all_values,y)
end
maximum(all_values)
# should return 359142
prog = readdlm("input.txt", '\t', Int, ',')
all_values = []
for x ∈ all_inputs
y = circuit(x,prog)
push!(all_values,y)
end
maximum(all_values)
# ```
# --- Part Two ---
#
# It's no good - in this configuration, the amplifiers can't generate a large enough output signal to produce the thrust you'll need. The Elves quickly talk you through rewiring the amplifiers into a feedback loop:
#
# O-------O O-------O O-------O O-------O O-------O
# 0 -+->| Amp A |->| Amp B |->| Amp C |->| Amp D |->| Amp E |-.
# | O-------O O-------O O-------O O-------O O-------O |
# | |
# '--------------------------------------------------------+
# |
# v
# (to thrusters)
# Most of the amplifiers are connected as they were before; amplifier A's output is connected to amplifier B's input, and so on. However, the output from amplifier E is now connected into amplifier A's input. This creates the feedback loop: the signal will be sent through the amplifiers many times.
#
# In feedback loop mode, the amplifiers need totally different phase settings: integers from 5 to 9, again each used exactly once. These settings will cause the Amplifier Controller Software to repeatedly take input and produce output many times before halting. Provide each amplifier its phase setting at its first input instruction; all further input/output instructions are for signals.
#
# Don't restart the Amplifier Controller Software on any amplifier during this process. Each one should continue receiving and sending signals until it halts.
#
# All signals sent or received in this process will be between pairs of amplifiers except the very first signal and the very last signal. To start the process, a 0 signal is sent to amplifier A's input exactly once.
#
# Eventually, the software on the amplifiers will halt after they have processed the final loop. When this happens, the last output signal from amplifier E is sent to the thrusters. Your job is to find the largest output signal that can be sent to the thrusters using the new phase settings and feedback loop arrangement.
#
# Here are some example programs:
#
# Max thruster signal 139629729 (from phase setting sequence 9,8,7,6,5):
#
# 3,26,1001,26,-4,26,3,27,1002,27,2,27,1,27,26,
# 27,4,27,1001,28,-1,28,1005,28,6,99,0,0,5
# Max thruster signal 18216 (from phase setting sequence 9,7,8,5,6):
#
# 3,52,1001,52,-5,52,3,53,1,52,56,54,1007,54,5,55,1005,55,26,1001,54,
# -5,54,1105,1,12,1,53,54,53,1008,54,0,55,1001,55,1,55,2,53,55,53,4,
# 53,1001,56,-1,56,1005,56,6,99,0,0,0,0,10
# Try every combination of the new phase settings on the amplifier feedback loop. What is the highest signal that can be sent to the thrusters?
#
# Your puzzle answer was 4374895.
#
#
#
# +
pos(p) = p + 1;
ref(a,p) = a[pos(a[pos(p)])];
@enum Op plus=1 mul=2 stor=3 out=4 halt=99
@enum Mode posi=0 imm=1
struct Operation
op::Op
mode1::Mode
mode2::Mode
mode3::Mode
end
getOps(str::String) = getOps(parse(Int,str))
function getOps(num)
op = Op(num % 100)
mode1 = Mode((num ÷ 100) % 10)
mode2 = Mode((num ÷ 1000) % 10)
mode3 = Mode((num ÷ 10000) % 10)
Operation(op,mode1,mode2,mode3)
end
function op1(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
prog[pos(prog[pos(p+3)])] = p1 + p2
p + 4
end
function op2(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
prog[pos(prog[pos(p+3)])] = p1 * p2
p + 4
end
function op3(operation,p,prog,input)
# println("entering stor ","p ",p," prog ",prog, " input ",input)
p1 = prog[pos(p+1)]
#println("storage position",p1)
prog[pos(p1)] = input
#println("exiting stor")
p + 2
end
function op4(operation,p,prog,input)
output = []
#println("entering out")
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
#println("output:",p1)
push!(output,p1)
#println("exiting out $output")
(p + 2,output)
end
function execute(op::Operation, p, prog)
if op.op == plus
return op1(op,p,prog)
end
if op.op == mul
return op2(op,p,prog)
end
if op.op == stor
return op3(op,p,prog,1)
end
if op.op == out
return op4(op,p,prog)
end
error("panic executing unknown op")
end
function execute(prog)
p = 0
op = getOps(prog[pos(p)])
#println("op: ", op)
while op.op != halt
#println(prog)
p = execute(op,p,prog)
#println("instruction: ",p, " -> ",prog[pos(p)])
op = getOps(prog[pos(p)])
end
prog
end;
println(execute([1,9,10,3,2,3,11,0,99,30,40,50]) == [3500,9,10,70,2,3,11,0,99,30,40,50])
println(execute([1,0,0,0,99]) == [2,0,0,0,99])
println(execute([2,3,0,3,99]) == [2,3,0,6,99])
println(execute([2,4,4,5,99,0]) == [2,4,4,5,99,9801])
println(execute([1,1,1,4,99,5,6,0,99]) == [ 30,1,1,4,2,5,6,0,99])
# Opcode 5 is jump-if-true:
# if the first parameter is non-zero,
# it sets the instruction pointer to the value from the second parameter.
# Otherwise, it does nothing.
function op5(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
if p1 != 0
return p2
else
return p + 3
end
end
# Opcode 6 is jump-if-false:
# if the first parameter is zero,
# it sets the instruction pointer to the value from the second parameter. Otherwise, it does nothing.
function op6(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
if p1 == 0
return p2
else
return p + 3
end
end
# Opcode 7 is less than:
# if the first parameter is less than the second parameter,
# it stores 1 in the position given by the third parameter. Otherwise, it stores 0.
function op7(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
if p1 < p2
prog[pos(prog[pos(p+3)])] = 1
else
prog[pos(prog[pos(p+3)])] = 0
end
p + 4
end
# Opcode 8 is equals:
# if the first parameter is equal to the second parameter,
# it stores 1 in the position given by the third parameter. Otherwise, it stores 0.
function op8(operation,p,prog)
if operation.mode1 == posi
p1 = ref(prog,p+1)
elseif operation.mode1 == imm
p1 = prog[pos(p+1)]
end
if operation.mode2 == posi
p2 = ref(prog,p+2)
elseif operation.mode2 == imm
p2 = prog[pos(p+2)]
end
if p1 == p2
prog[pos(prog[pos(p+3)])] = 1
else
prog[pos(prog[pos(p+3)])] = 0
end
p + 4
end
@enum Op plus=1 mul=2 stor=3 out=4 jnz=5 jez=6 lt=7 eq=8 halt=99
@enum Mode posi=0 imm=1
struct Operation
op::Op
mode1::Mode
mode2::Mode
mode3::Mode
end
function execute(op::Operation, p, prog, input,output)
if op.op == plus
return (op1(op,p,prog),input,output,false)
end
if op.op == mul
return (op2(op,p,prog),input,output,false)
end
if op.op == stor
if length(input) == 0
return (p,input,output,true)
end
next = input[1]
input=input[2:end]
return (op3(op,p,prog,next),input,output,false)
end
if op.op == out
(p1,output) = op4(op,p,prog,output)
#println(output)
return (p1,input,output,false)
end
if op.op == jnz
return (op5(op,p,prog),input,output,false)
end
if op.op == jez
return (op6(op,p,prog),input,output,false)
end
if op.op == lt
return (op7(op,p,prog),input,output,false)
end
if op.op == eq
return (op8(op,p,prog),input,output,false)
end
error("panic executing unknown op")
end
function execute(prog,p,input,output)
op = getOps(prog[pos(p)])
while op.op != halt
(p,input,output,halt) = execute(op,p,prog,input,output)
if halt
return (prog,p,output,:pause)
end
op = getOps(prog[pos(p)])
end
(prog,p,output,:over)
end
# The above example program uses an input instruction to ask for a single number.
# The program will then
# output 999 if the input value is below 8,
# output 1000 if the input value is equal to 8,
# or output 1001 if the input value is greater than 8.
println(execute(
[3,21,1008,21,8,20,1005,20,22,107,8,21,20,1006,20,31,
1106,0,36,98,0,0,1002,21,125,20,4,20,1105,1,46,104,
999,1105,1,46,1101,1000,1,20,4,20,1105,1,46,98,99]
,0,[-8],[]))
println(execute(
[3,21,1008,21,8,20,1005,20,22,107,8,21,20,1006,20,31,
1106,0,36,98,0,0,1002,21,125,20,4,20,1105,1,46,104,
999,1105,1,46,1101,1000,1,20,4,20,1105,1,46,98,99]
,0,[8],[]))
a = execute(
[3,21,1008,21,8,20,1005,20,22,107,8,21,20,1006,20,31,
1106,0,36,98,0,0,1002,21,125,20,4,20,1105,1,46,104,
999,1105,1,46,1101,1000,1,20,4,20,1105,1,46,98,99]
,0,[16],[]);
#example usage
#using DelimitedFiles
#prog = readdlm("input.txt", '\t', Int, ',')
#execute(prog,5);
a
# -
# Max thruster signal 139629729 (from phase setting sequence 9,8,7,6,5):
function feedback(a,prog)
state = :start
p1,p2,p3,p4,p5=0,0,0,0,0
amp1,amp2,amp3,amp4,amp5=copy(prog),copy(prog),copy(prog),copy(prog),copy(prog)
output1,output2,output3,output4,output5=[0],[0],[0],[0],[0]
(amp1,p1,output1,state) = execute(amp1,p1,[a[1]],[]);
(amp2,p2,output2,state) = execute(amp2,p2,[a[2]],[]);
(amp3,p3,output3,state) = execute(amp3,p3,[a[3]],[]);
(amp4,p4,output4,state) = execute(amp4,p4,[a[4]],[]);
(amp5,p5,output5,state) = execute(amp5,p5,[a[5]],[]);
output5=[0]
while state != :over
(amp1,p1,output1,state) = execute(amp1,p1,[output5[1]],[]);
(amp2,p2,output2,state) = execute(amp2,p2,[output1[1]],[]);
(amp3,p3,output3,state) = execute(amp3,p3,[output2[1]],[]);
(amp4,p4,output4,state) = execute(amp4,p4,[output3[1]],[]);
(amp5,p5,output5,state) = execute(amp5,p5,[output4[1]],[]);
end
output5[1]
end
test_amp() = [3,26,1001,26,-4,26,3,27,1002,27,2,27,1,27,26,27,4,27,1001,28,-1,28,1005,28,6,99,0,0,5]
feedback([9,8,7,6,5],test_amp())
#Max thruster signal 18216 (from phase setting sequence 9,7,8,5,6):
test_amp() = [3,52,1001,52,-5,52,3,53,1,52,56,54,1007,54,5,55,1005,55,26,1001,54,-5,54,1105,1,12,1,53,54,53,1008,54,0,55,1001,55,1,55,2,53,55,53,4,53,1001,56,-1,56,1005,56,6,99,0,0,0,0,10]
feedback([9,7,8,5,6],test_amp())
#65210
test_amp() = [3,31,3,32,1002,32,10,32,1001,31,-2,31,1007,31,0,33,1002,33,7,33,1,33,31,31,1,32,31,31,4,31,99,0,0,0]
feedback([1,0,4,3,2],test_amp())
#echo
test_amp() = [3,15,3,16,1002,16,10,16,1,16,15,15,4,15,99,0,0]
feedback([4,3,2,1,0],test_amp())
# should return 4374895
using Combinatorics, DelimitedFiles
all_inputs = permutations([5,6,7,8,9]) |> collect;
prog = readdlm("input.txt", '\t', Int, ',')
all_values = []
for x ∈ all_inputs
y = feedback(x,prog)
push!(all_values,y)
end
maximum(all_values)
| julia/7/day7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
from PyQt5.QtWidgets import (QApplication, QWidget, QToolTip, QFileDialog,QButtonGroup,
QLabel, QRadioButton, QComboBox, QLineEdit, QPushButton, QGridLayout)
from PyQt5.QtGui import QIcon
from PyQt5.QtGui import QFont
from playsound import playsound
from scipy import stats
import scipy.io
from scipy.io import wavfile
import sounddevice as sd
import numpy as np
import pandas as pd
import pygame
import time
from numpy.lib import stride_tricks
from PyQt5 import QtGui
import python_speech_features
import librosa
import csv
from keras.models import load_model
from keras.preprocessing.image import ImageDataGenerator
from PIL import Image
import math
import matplotlib.pyplot as plt
from matplotlib.backends.backend_qt5agg import FigureCanvasQTAgg as FigureCanvus
from matplotlib.backends.backend_qt5agg import NavigationToolbar2QT as NavigationToolbar
from sklearn.externals import joblib
import os
import glob,os.path
'''
#QtCore module contains the core non-GUI functionality. This module is used for working with time, files and directories, various data types, streams, URLs, mime types, threads or processes
#QtGui contains classes for windowing system integration, event handling, 2D graphics, basic imaging, fonts and text
#QtWidgets module contains classes that provide a set of UI elements to create classic desktop-style user interfaces
#QtNetwork module contains the classes for network programming.
#QtWebSockets module contains classes that implement the WebSocket protocol.
#QtSql module provides classes for working with databases.
#QtXml contains classes for working with XML files
#QtMultimedia contains classes to handle multimedia content and APIs to access camera and radio functionality.
'''
class Window(QWidget): #inherits from the QWidget class. QWidget widget is the base class of all user interface objects in PyQt5.
def __init__(self):
super().__init__() #super() method returns the parent object of the Window class and we call its constructor.
self.initUI()
def initUI(self):
#Naming The Widgets
audioType = QLabel('Audio Input (.wav file): ')
#audioType.setFont(QFont.setBold(self,True))
fs = QLabel('Sampling Freq.(Hz)')
time = QLabel('Duration(Sec)')
predictionPart = QLabel('Prediction Part-50% Overlap')
myFont = QtGui.QFont()
myFont.setBold(True)
predictionPart.setFont(myFont)
modelSelect = QLabel('Select The Model')
frameSize = QLabel('Frame Size (in ms)')
windowLenEdit = QLabel('Window Length(ms)')
predicitonResult = QLabel('Total Prediction')
self.lbl = QLabel(self)
self.Index = 0
self.modelIndex = 0
pygame.init()
#Implementing Those Widgets
self.nameOfAudio = QLineEdit()
self.uploadAudio = QPushButton("Upload Audio")
self.fsEdit = QComboBox()
##### Reording Button #####
self.timeDuration = QLineEdit()
self.loadAudio = QPushButton("Load Data")
self.loadAudio.setEnabled(False)
self.plotEdit = QPushButton("Plot Data")
self.plotEdit.setEnabled(False)
self.playFrame = QPushButton('Play Frame')
self.playFrame.setEnabled(False)
self.figure = plt.figure(figsize=(5,5),dpi=120)
self.canvas = FigureCanvus(self.figure)
self.toolbar = NavigationToolbar(self.canvas,self)
self.importModel = QPushButton("Import Model")
self.importModel.setEnabled(False)
self.processStart = QPushButton("Process")
self.processStart.setEnabled(False)
self.predictStart = QPushButton("Predict")
self.predictStart.setEnabled(False)
##### Model Selection #####
self.modelSelect = QComboBox()
##### Frame Size Selection #####
self.frameSizeEdit = QComboBox()
##### Window Length Selection for Prediction #####
self.windowLenEdit = QComboBox()
self.modelGraph = QComboBox()
self.Show = QPushButton("Show")
self.Show.setEnabled(False)
self.predictionRecord = QPushButton("Result")
self.predictionRecord.setEnabled(False)
self.totalPredictionResult = QLineEdit()
self.cancelEdit = QPushButton("CANCEL")
self.cancelEdit.setEnabled(False)
self.back = QPushButton("<<")
self.back.setEnabled(False)
self.front = QPushButton(">>")
self.front.setEnabled(False)
self.showFrame = QLineEdit()
self.startTime = QLineEdit()
self.endTime = QLineEdit()
self.reset = QPushButton('Reset')
self.reset.setEnabled(False)
#Filling In Details
self.fsEdit.addItem('16000')
self.fsEdit.addItem('44100')
self.frameSizeEdit.addItem('100')
self.frameSizeEdit.addItem('250')
self.frameSizeEdit.addItem('500')
#self.frameSizeEdit.addItem('500')
#self.modelSelect.addItem('SVM')
self.modelSelect.addItem('FNN')
self.modelSelect.addItem('SVM')
self.modelSelect.addItem('1D CNN')
self.modelSelect.addItem('2D CNN')
#self.modelSelect.addItem('1D CNN Without MFCC')
self.windowLenEdit.addItem('200')
self.windowLenEdit.addItem('500')
self.windowLenEdit.addItem('1000')
self.windowLenEdit.addItem('2000')
self.windowLenEdit.addItem('5000')
#self.modelGraph.addItem('Model Plot')
self.modelGraph.addItem('K-fold Accuracy')
self.modelGraph.addItem('K-fold Loss')
self.modelGraph.addItem('Confusion-Matrix')
#Setting Layout
grid = QGridLayout()
grid.setSpacing(5)
#1st Row
#grid.addWidget(audioType, 1, 0, 1, 1)
grid.addWidget(self.nameOfAudio,1,1,1,2)
grid.addWidget(self.uploadAudio,1,0,1,1)
grid.addWidget(fs , 1, 3, 1, 1)
grid.addWidget(self.fsEdit,1,4,1,1)
#2nd Row
grid.addWidget(self.loadAudio,2,1,1,1)
grid.addWidget(self.plotEdit, 2, 2, 1, 1)
grid.addWidget(time, 2, 3, 1, 1)
grid.addWidget(self.timeDuration, 2, 4, 1, 1)
#3rd Row
grid.addWidget(self.playFrame,3,0,1,1)
grid.addWidget(self.toolbar, 3, 1, 1, 4)
#4th Row
grid.addWidget(self.canvas, 4, 0, 4, 4)
grid.addWidget(self.lbl, 4,4,1,2)
#5th Row
grid.addWidget(predictionPart, 8, 2, 1, 1)
#6th Row
grid.addWidget(modelSelect, 9, 0, 1, 1)
grid.addWidget(self.modelSelect, 9, 1, 1, 1)
grid.addWidget(frameSize , 9, 2, 1, 1)
grid.addWidget(self.frameSizeEdit, 9, 3, 1, 1)
grid.addWidget(self.modelGraph, 9, 4, 1, 1)
grid.addWidget(self.Show, 9, 5, 1, 1)
#7th Row
grid.addWidget(windowLenEdit , 10, 0, 1, 1)
grid.addWidget(self.windowLenEdit, 10, 1, 1, 1)
grid.addWidget(self.importModel, 10, 2, 1, 1)
grid.addWidget(self.processStart,10, 3, 1, 1)
grid.addWidget(self.predictStart,10, 4, 1, 1)
#8th Row
grid.addWidget(predicitonResult, 11, 0, 1, 1)
grid.addWidget(self.totalPredictionResult, 11,1, 1, 3)
self.totalPredictionResult.resize(220,80)
grid.addWidget(self.predictionRecord, 11, 4, 1, 1)
#9th Row
grid.addWidget(self.back,12,0,1,1)
grid.addWidget(self.startTime,12,1,1,1)
grid.addWidget(self.showFrame,12,2,1,1)
grid.addWidget(self.endTime,12,3,1,1)
grid.addWidget(self.front,12,4,1,1)
#10th row
grid.addWidget(self.reset,13,4,1,1)
grid.addWidget(self.cancelEdit,13,5,1,1)
self.setLayout(grid)
self.uploadAudio.clicked.connect(self.showDialog)
self.loadAudio.clicked.connect(self.load)
self.plotEdit.clicked.connect(self.plot)
self.playFrame.clicked.connect(self.playframe)
self.importModel.clicked.connect(self.importmodel)
self.processStart.clicked.connect(self.process)
self.predictStart.clicked.connect(self.predict)
self.predictionRecord.clicked.connect(self.record)
self.Show.clicked.connect(self.modelShow)
self.back.clicked.connect(self.left)
self.front.clicked.connect(self.right)
self.reset.clicked.connect(self.Reset)
self.cancelEdit.clicked.connect(self.cancel)
self.setGeometry(300, 300, 500, 400) #locates the window on the screen and sets it size(x,y,x+w,y+d)
self.setWindowTitle('GUI for Audio Scene Prediction')
#self.show(QIcon('FileName.png'))
self.show()
def str2int_fs(self):
b = [int(y) for y in self.fsEdit.currentText()]
c = 0;
for i in b:
c = c*10 + i
return c
def str2int_framesize(self):
b = [int(y) for y in self.frameSizeEdit.currentText()]
c = 0;
for i in b:
c = c*10 + i
return c
def str2int_winlen(self):
b = [int(y) for y in self.windowLenEdit.currentText()]
c = 0;
for i in b:
c = c*10 + i
return c
def showDialog(self):
self.fname = QFileDialog.getOpenFileName(self,
'Open Recorded Audio',
'C:\\Users\\ASUS\\AppData\\Local\\Programs\\Python\\Python36\\BTP',
'Audio files (*.wav *.mp3)')
self.nameOfAudio.setText(self.fname[0])
self.loadAudio.setEnabled(True)
def load(self):
fs1 = self.str2int_fs()
(self.wavFile,self.rate) = librosa.load(self.fname[0],sr=int(fs1),mono=True)
time_duration = self.wavFile.size/self.rate
pr = str(time_duration) + " Sec"
self.timeDuration.setText(pr)
self.plotEdit.setEnabled(True)
self.importModel.setEnabled(True)
self.Show.setEnabled(True)
self.reset.setEnabled(True)
self.cancelEdit.setEnabled(True)
def Identify(self):
a = ['AC & FAN','CRYING','MUSIC','SPEECH']
return (a[self.mode[self.Index]])
def Identify_wav(self):
a = ['AC & FAN.wav','CRYING.wav','MUSIC.wav','SPEECH.wav']
return (a[self.mode[self.Index]])
def left(self):
self.front.setEnabled(True)
self.Index -= 1
if(self.Index<=0):
self.back.setEnabled(False)
self.frameplot()
start = "<< "+"{:.1f}".format(self.Index*self.time)+' sec.'
self.startTime.setText(start)
end = "{:.1f}".format((self.Index+1)*self.time)+' sec. >>'
self.endTime.setText(end)
show = self.Identify()
show_wav = self.Identify_wav()
p = "Frame " + str(self.Index+1) + " || " + show
self.showFrame.setText(p)
pygame.mixer.music.load(show_wav)
pygame.mixer.music.play()
def right(self):
self.back.setEnabled(True)
self.Index += 1
if (self.Index>=self.mode.size-1):
self.front.setEnabled(False)
self.frameplot()
start = "<< "+"{:.1f}".format(self.Index*self.time)+' sec.'
self.startTime.setText(start)
end = "{:.1f}".format((self.Index+1)*self.time)+' sec. >>'
self.endTime.setText(end)
show = self.Identify()
show_wav = self.Identify_wav()
p = "Frame " + str(self.Index+1) + " || " + show
self.showFrame.setText(p)
pygame.mixer.music.load(show_wav)
pygame.mixer.music.play()
def plot(self):
self.figure.clear()
ax = self.figure.add_subplot(111)
x = np.arange(1,self.wavFile.size+1)
x = np.divide(x,self.rate)
ax.plot(x,self.wavFile,'b-')
#ax.set_title('Uploaded Audio')
self.canvas.draw()
self.playFrame.setEnabled(True)
self.passWavFile = self.wavFile
def frameplot(self):
self.playFrame.setEnabled(True)
self.figure.clear()
start = int(self.Index*self.time*self.rate)
end = int((self.Index+1)*self.time*self.rate)-1
wave = self.wavFile[start:end]
x = np.arange(1,wave.size+1)
x = np.divide(x,self.rate)
x = np.add(x,self.Index*self.time)
ax = self.figure.add_subplot(111)
ax.plot(x,wave,'b-')
ax.set_title('Frame Number '+str(self.Index+1))
self.canvas.draw()
self.passWavFile = wave
def playframe(self):
sd.play(self.passWavFile,self.rate)
""" short time fourier transform of audio signal """
def stft(self,sig, frameSize, overlapFac=0.9, window=np.hanning):
win = window(frameSize)
hopSize = int(frameSize - np.floor(overlapFac * frameSize))
# zeros at beginning (thus center of 1st window should be for sample nr. 0)
samples = np.append(np.zeros(int(np.floor(frameSize/2.0))), sig)
# cols for windowing
cols = np.ceil( (len(samples) - frameSize) / float(hopSize)) + 1
# zeros at end (thus samples can be fully covered by frames)
samples = np.append(samples, np.zeros(frameSize))
frames = stride_tricks.as_strided(samples, shape=(int(cols), frameSize), strides=(samples.strides[0]*hopSize, samples.strides[0])).copy()
frames *= win
return np.fft.rfft(frames)
def logscale_spec(self,spec, sr=44100, factor=20.):
timebins, freqbins = np.shape(spec)
scale = np.linspace(0, 1, freqbins) ** factor
scale *= (freqbins-1)/max(scale)
scale = np.unique(np.round(scale))
# create spectrogram with new freq bins
newspec = np.complex128(np.zeros([timebins, len(scale)]))
for i in range(0, len(scale)):
if i == len(scale)-1:
newspec[:,i] = np.sum(spec[:,int(scale[i]):], axis=1)
else:
newspec[:,i] = np.sum(spec[:,int(scale[i]):int(scale[i+1])], axis=1)
# list center freq of bins
allfreqs = np.abs(np.fft.fftfreq(freqbins*2, 1./sr)[:freqbins+1])
freqs = []
for i in range(0, len(scale)):
if i == len(scale)-1:
freqs += [np.mean(allfreqs[int(scale[i]):])]
else:
freqs += [np.mean(allfreqs[int(scale[i]):int(scale[i+1])])]
return newspec, freqs
def plotstft(self, samples, samplerate, binsize=400, plotpath=None, colormap="jet"):
plt.close('all')
s = self.stft(samples, binsize)
sshow, freq = self.logscale_spec(s, factor=1.0, sr=samplerate)
ims = 20.*np.log10(np.abs(sshow)/10e-6) # amplitude to decibel
w = ims.shape[0]/1250.0
h = ims.shape[1]/1250.0
plt.figure(figsize=(w,h))
plt.axis('off')
fig = plt.imshow(ims.T, origin="lower", aspect="auto", cmap=colormap, interpolation="nearest")
fig.axes.get_xaxis().set_visible(False)
fig.axes.get_yaxis().set_visible(False)
if plotpath:
plt.savefig(plotpath, bbox_inches="tight", pad_inches=0., dpi=1000)
else:
plt.show()
plt.clf()
return ims
def importmodel(self):
if (self.modelSelect.currentText()=='FNN'):
if (self.frameSizeEdit.currentText()=='100'):
self.my_model=load_model('manikanta_FeedforwardNN_4class_100ms.h5')
self.modelIndex = 0
elif (self.frameSizeEdit.currentText()=='250'):
self.my_model=load_model('manikanta_FeedforwardNN_4class_250ms.h5')
self.modelIndex = 0
elif (self.frameSizeEdit.currentText()=='500'):
self.my_model=load_model('manikanta_FeedforwardNN_4class_500ms.h5')
self.modelIndex = 0
elif (self.modelSelect.currentText()=='1D CNN'):
if (self.frameSizeEdit.currentText()=='100'):
self.my_model=load_model('my_CNN_100ms_mani.h5')
self.modelIndex = 1
elif (self.frameSizeEdit.currentText()=='250'):
self.my_model=load_model('my_CNN_250ms_mani.h5')
self.modelIndex = 1
else :
self.my_model=load_model('my_CNN_500ms_mani.h5')
self.modelIndex = 1
elif(self.modelSelect.currentText()=='SVM'):
if (self.frameSizeEdit.currentText()=='100'):
self.filename = 'SVM_100ms_Rbf_model.save'
self.scaler_file = "my_scaler_100ms.save"
self.modelIndex = 2
elif (self.frameSizeEdit.currentText()=='250'):
self.filename = 'SVM_250ms_Rbf_model.save'
self.scaler_file = "my_scaler_250ms.save"
self.modelIndex = 2
else :
self.filename = 'SVM_500ms_Rbf_model.save'
self.scaler_file = "my_scaler_500ms.save"
self.modelIndex = 2
else:
if (self.frameSizeEdit.currentText()=='100'):
self.my_model=load_model('mani_spectrogrammodel_cnn_100ms.h5')
self.modelIndex = 1
elif (self.frameSizeEdit.currentText()=='250'):
self.my_model=load_model('mani_spectrogrammodel_cnn_250ms.h5')
self.modelIndex = 1
else :
self.my_model=load_model('mani_spectrogrammodel_cnn_500ms.h5')
self.modelIndex = 1
self.processStart.setEnabled(True)
print(self.modelIndex)
def process(self):
self.frameSize = self.str2int_framesize()
if(self.modelIndex!=3):
print(self.frameSize)
self.overLap = self.frameSize/2
print(self.overLap)
print(self.rate*self.frameSize/1000)
print(math.log2(self.rate*self.frameSize/1000))
print(math.ceil(math.log2(self.rate*self.frameSize/1000)))
self.nfft = 2**(math.ceil(math.log2(self.rate*self.frameSize/1000)))
self.mfcc = python_speech_features.base.mfcc(self.wavFile, samplerate=self.rate, winlen=self.frameSize/1000, winstep=self.overLap/1000, numcep=13, nfilt=26,
nfft=self.nfft, lowfreq=0, highfreq=None, preemph=0.97, ceplifter=22, appendEnergy=True)
self.csvData = self.mfcc
with open('prediction.csv','w') as csvFile:
writer = csv.writer(csvFile)
writer.writerows(self.csvData)
csvFile.close()
else:
self.classes = ['Spectogram']
fs = self.rate
fsize = int(fs/2)
hop = fsize // 10
hop_length = fsize // 40
num_classes = len(self.classes)
y = self.wavFile
rng = y.shape[0]//hop - 1
for i in range(0, rng):
data = y[i*hop:i*hop+fsize]
path = './Test_2D_CNN/Spectogram/' + str(i+1) + '.png'
ims = self.plotstft(data, fs, plotpath=path)
self.img_height,self.img_width = 128,128
self.test_datagen = ImageDataGenerator(rescale=1./255)
self.test_generator = self.test_datagen.flow_from_directory('./Test_2D_CNN',target_size=(self.img_height,self.img_width),
batch_size=21,
shuffle=False,
classes=self.classes,
class_mode='categorical')
self.predictStart.setEnabled(True)
def predict(self):
if(self.modelIndex==2):
df1=pd.read_csv("prediction.csv",na_values=['NA','?'])
df1.columns=['MFCC0', 'MFCC1','MFCC2','MFCC3','MFCC4','MFCC5','MFCC6','MFCC7','MFCC8',
'MFCC9', 'MFCC10' ,'MFCC11', 'MFCC12']
loaded_model_rbf1 = joblib.load(open(self.filename,'rb'))
scaler = joblib.load(self.scaler_file)
X = scaler.transform(df1)
holdout_pred_rbf1 = loaded_model_rbf1.predict(X)
self.test_pred1 = np.empty(holdout_pred_rbf1.size,dtype='int')
c = 0
for i in holdout_pred_rbf1:
if(i=='AC & FAN'):
self.test_pred1[c] = 0
elif(i=='CRYING'):
self.test_pred1[c] = 1
elif(i=='MUSIC'):
self.test_pred1[c] = 2
else:
self.test_pred1[c] = 3
c += 1
print(holdout_pred_rbf1)
print(self.test_pred1)
elif(self.modelIndex==3):
self.test_generator.reset()
self.test_steps = self.test_generator.n//self.test_generator.batch_size
self.y_test = self.my_model.predict_generator(self.test_generator,
steps=self.test_steps,
verbose=1)
self.test_pred1 = np.argmax(self.y_test, axis=1).astype('int8')
print(self.test_pred1)
filelist = glob.glob(os.path.join('./Test_2D_CNN/Spectogram/', "*.png"))
for f in filelist:
os.remove(f)
else:
df1=pd.read_csv("prediction.csv")
df1.columns=['MFCC0', 'MFCC1','MFCC2','MFCC3','MFCC4','MFCC5','MFCC6','MFCC7','MFCC8', 'MFCC9', 'MFCC10' ,'MFCC11', 'MFCC12']
if (self.modelIndex==1):
df1 = np.expand_dims(df1,axis=2)
my_his1=self.my_model.predict(df1)
self.test_pred1 = np.argmax(my_his1, axis=1)
print(self.test_pred1)
self.size = self.test_pred1.size
self.winLen = self.str2int_winlen()
self.predictionFrame = int(2*(self.winLen/self.frameSize))
NPadding = (-self.test_pred1.size)%self.predictionFrame
random = -np.random.randint(1,20000000,size=NPadding)
self.test_pred1 = np.concatenate((self.test_pred1,random),axis=0)
#self.test_pred1 = np.pad(self.test_pred1, (0,NPadding), 'constant', constant_values=(0,-1))
self.test_pred2 = librosa.util.frame(self.test_pred1, frame_length=self.predictionFrame, hop_length=self.predictionFrame)
self.test_pred2 = self.test_pred2.T
self.mode = stats.mode(self.test_pred2, axis=1)
self.mode = self.mode[0][:, 0]
print(self.mode)
self.time = self.winLen/1000
if(self.modelIndex!=3):
os.remove('prediction.csv')
self.predictionRecord.setEnabled(True)
def record(self):
self.Index = 0
self.front.setEnabled(True)
c_acfan=sum(self.test_pred1==0)
c_crying=sum(self.test_pred1==1)
c_music=sum(self.test_pred1==2)
c_speech=sum(self.test_pred1==3)
print('AC & FAN Predicted : ',100*c_acfan/self.size)
print('CRYING Predicted : ',100*c_crying/self.size)
print('MUSIC Predicted : ',100*c_music/self.size)
print('SPEECH Predicted : ',100*c_speech/self.size)
pr = 'AF: '+"{:.2f}".format(100*c_acfan/self.size) + '||' + 'C: ' + "{:.2f}".format(100*c_crying/self.size) + '||' + 'M: '+"{:.2f}".format(100*c_music/self.size)+ '||' + 'S: '+"{:.2f}".format(100*c_speech/self.size)
self.totalPredictionResult.setText(pr)
self.frameplot()
show = self.Identify()
show_wav = self.Identify_wav()
p = "Frame "+str(self.Index+1) + " || " + show
self.startTime.setText('<< 0 sec')
self.showFrame.setText(p)
self.endTime.setText(str(self.time)+' sec >>')
pygame.mixer.music.load(show_wav)
pygame.mixer.music.play()
def modelShow(self):
img_name = self.modelGraph.currentText()
frameS = self.frameSizeEdit.currentText()
modelN = self.modelSelect.currentText()
image_name = frameS+'_'
if(modelN=='FNN'):
image_name += 'FNN_'
elif(modelN=='1D CNN'):
image_name += 'CNN_'
else:
image_name += 'SVM_'
if(img_name=='K-fold Accuracy'):
image_name += 'acc'
elif(img_name=='K-fold Loss'):
image_name += 'loss'
else:
image_name += 'cm'
pixmap = QtGui.QPixmap(image_name+'.png')
self.lbl.setPixmap(pixmap)
def cancel(self):
print('Cancelled')
self.close()
def Reset(self):
self.figure.clear()
self.loadAudio.setEnabled(False)
self.plotEdit.setEnabled(False)
self.playFrame.setEnabled(False)
self.processStart.setEnabled(False)
self.Show.setEnabled(False)
self.back.setEnabled(False)
self.front.setEnabled(False)
self.predictionRecord.setEnabled(False)
self.predictStart.setEnabled(False)
self.Index = 0
self.figure.clear()
self.nameOfAudio.setText(' ')
self.timeDuration.setText('')
self.totalPredictionResult.setText('')
self.showFrame.setText('')
self.startTime.setText('')
self.endTime.setText('')
self.lbl.clear()
self.reset.setEnabled(False)
if __name__ == '__main__':
##
app = QApplication(sys.argv) #Every PyQt5class Window(QWidget): #inherits from the QWidget class. QWidget widget is the base class of all user interface objects in PyQt5.
window = Window()
sys.exit(app.exec_()) #enters the mainloop of the application. The event handling starts from this point.
| GUI for Audio Scene Recognition (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **<NAME>**
# *January 13, 2021*
#
# # Download GOES Data: Timerange
# For all options, refer to the GOES-2-go Reference Guide: [goes2go.data.goes_timerange](https://blaylockbk.github.io/goes2go/_build/html/reference_guide/index.html#goes2go.data.goes_timerange)
# +
from goes2go.data import goes_timerange
from datetime import datetime
import pandas as pd
# -
# ---
# ### Example 1:
# Download an ABI file from GOES-East for an hour period. Data is returned as a file list.
# +
## Dates may be specified as datetime, pandas datetimes, or string dates
## that pandas can interpret.
## Specify start/end time with datetime object
#start = datetime(2021, 1, 1, 0, 30)
#end = datetime(2021, 1, 1, 1, 30)
## Specify start/end time as a panda-parsable string
start = '2021-01-01 00:30'
end = '2021-01-01 01:30'
g = goes_timerange(start, end,
satellite='goes16',
product='ABI',
return_as='filelist')
# -
g
g.attrs
# ### Show the files on my home drive...
# + language="bash"
# tree ~/data
# -
| docs/user_guide/notebooks/DEMO_download_goes_timerange.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 3.6.8
# language: python
# name: 3.6.8
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import lightgbm as lgb
import joblib
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import f1_score
# -
df = pd.read_csv("data/one_hot_data.csv")
df.head()
df2 = pd.read_csv("data/train_labels.csv")
df2.head()
test = df['building_id']
test2 = df2['building_id']
for i in range(len(test)):
if test[i] != test2[i]:
print(i)
df = df.drop(['Unnamed: 0', 'building_id'], axis=1)
df.head()
columns = df.columns
X = df.to_numpy()
Y = df2['damage_grade'].to_numpy()
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.2)
d_train = lgb.Dataset(X_train, label=y_train-1)
params = {'objective': 'multiclass',
'num_class': 3,
'metric': 'multi_logloss',
'boosting_type': 'gbdt', # dart
'max_bin': 4095,
'learning_rate': 0.001,
'num_threads': 6,
'num_iterations': 5000,
'num_leaves': 1200
}
lite = lgb.train(params, d_train)
y_pred = lite.predict(X_test)
y_preds = np.argmax(y_pred, axis=1)
y_preds += 1
y_train_pred = lite.predict(X_train)
y_train_preds = np.argmax(y_train_pred, axis=1)
y_train_preds += 1
print(f1_score(y_train, y_train_preds, average='micro'))
print(f1_score(y_test, y_preds, average='micro'))
joblib.dump(lite, "decision_models/lite-7.model")
# 1 gdbt - 100, 2 dart - 100, 3 dart-leaves-500-bin-1023, 3 (72.26) gdbt-leaves-500-bin-1023, 4 (73.33) gdbt-leaves-500-bin-4095,
# 5 (73.87) gdbt-leaves-1000-bin-4095, 6 (73.48) dart-leaves-1000-bin-4095, 7 (74.65) gbdt-leaves-1200-bin-4095
# +
def get_leaves_vals(x):
accuracies = [None] * len(x)
accuracies2 = [None] * len(x)
for i in range(len(x)):
leaves = lgb.LGBMClassifier(n_estimators=1000, n_jobs=6, num_leaves=x[i], objective='multiclass',
boosting_type='gbdt')
leaves.fit(X_train, y_train)
accuracies[i] = f1_score(y_test, leaves.predict(X_test), average='micro')
leaves = lgb.LGBMClassifier(n_estimators=1000, n_jobs=6, num_leaves=x[i], objective='multiclass',
boosting_type='dart')
leaves.fit(X_train, y_train)
accuracies2[i] = f1_score(y_test, leaves.predict(X_test), average='micro')
return accuracies, accuracies2
xs3000 = [30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, 90, 95, 100]
ys3000, ysdart = get_leaves_vals(xs3000)
# -
print(xs3000[:-4])
print(xs3000[-4:])
morexs = [81, 83, 87, 89]
moreys3000, moreysdart = get_leaves_vals(morexs)
print(xs3000[:-4] + morexs[:2] + [xs3000[-4]] + morexs[2:] + xs3000[-3:])
# +
actualxs = xs3000[:-4] + morexs[:2] + [xs3000[-4]] + morexs[2:] + xs3000[-3:]
actuays = ys3000[:-4] + moreys3000[:2] + [ys3000[-4]] + moreys3000[2:] + ys3000[-3:]
actualysdart = ysdart[:-4] + moreysdart[:2] + [ysdart[-4]] + moreysdart[2:] + ysdart[-3:]
plt.figure(figsize=(12,6))
plt.plot(actualxs, actuays, '-ok', label='gbdt', color='black');
plt.plot(actualxs, actualysdart, '-ok', label='dart', color='brown');
plt.legend()
plt.xlabel('Number of Leaves')
plt.ylabel('F1 Score')
plt.title('Lite Gradient Boosting Ensemble Validation accuracy for Number of Leaves')
# plt.savefig('source_images/num_leaves_analysis.png')
# -
plt.figure(figsize=(12,6))
plt.plot(xs3000, ys3000, '-ok', label='gbdt', color='black');
plt.plot(xs3000, ysdart, '-ok', label='dart', color='brown');
plt.legend()
plt.xlabel('Number of Leaves')
plt.ylabel('F1 Score')
plt.title('Lite Gradient Boosting Ensemble Validation Accuracy for Number of Leaves')
plt.savefig('source_images/num_leaves_analysis.png')
# +
def getcounts(x):
accuracies = [None] * len(x)
accuracies2 = [None] * len(x)
for i in range(len(x)):
leaves = lgb.LGBMClassifier(n_estimators=x[i], n_jobs=6, num_leaves=80, objective='multiclass',
boosting_type='gbdt')
leaves.fit(X_train, y_train)
accuracies[i] = f1_score(y_test, leaves.predict(X_test), average='micro')
leaves = lgb.LGBMClassifier(n_estimators=x[i], n_jobs=6, num_leaves=80, objective='multiclass',
boosting_type='dart')
leaves.fit(X_train, y_train)
accuracies2[i] = f1_score(y_test, leaves.predict(X_test), average='micro')
return accuracies
xs3000 = [100, 1000, 2000, 3000, 4000]
ys3000, ysdart = get_leaves_vals(xs3000)
# +
plt.figure(figsize=(12,6))
plt.plot(xs3000, ys3000, '-ok', label='gbdt', color='black');
plt.plot(xs3000, ysdart, '-ok', label='dart', color='brown');
plt.legend()
plt.xlabel('Number of Estimators')
plt.ylabel('F1 Score')
plt.title('Lite Gradient Boosting Ensemble Validation accuracy for Number of Estimators')
plt.savefig('source_images/num_estimator_analysis.png')
# -
# lite_ensemble = lgb.LGBMClassifier(n_estimators=3000, n_jobs=6, num_leaves=1000)
lite_ensemble = lgb.LGBMClassifier(n_estimators=4000, n_jobs=6, num_leaves=65, objective='multiclass',
boosting_type='dart')
lite_ensemble.fit(X_train, y_train)
print(f1_score(y_train, lite_ensemble.predict(X_train), average='micro'))
print(f1_score(y_test, lite_ensemble.predict(X_test), average='micro'))
joblib.dump(lite_ensemble, "decision_models/lite-ensemble-13.model")
# 1 (74.75) ensemble-leaves-31-n_estimators-3000, 2 (74.3) ensemble-leaves-100-n_estimators-3000, 3 (74.82) ensemble-leaves-32-n_estimators-3000,
# 4 (74.70) ensemble-leaves-33-n_estimators-3000, 5 (74.79) ensemble-leaves-34-n_estimators-3000, 6 (74.68) ensemble-leaves-35-n_estimators-3000,
# 7 (74.70) ensemble-leaves-30-n_estimators-3000, 8 (74.67) ensemble-leaves-32-n_estimators-5000, 9 (74.68) ensemble-leaves-32-n_estimators-4000,
# 10 (74.42) dart-ensemble-leaves-32-n_estimators-5000, 11 (74.70) dart-ensemble-leaves-60-n_estimators-4000,
# 12 (74.61) gbdt-ensemble-leaves-60-n_estimators-4000, 13 (74.55) dart-ensemble-leaves-50-n_estimators-4000,
# 14 (74.53) dart-ensemble-leaves-40-n_estimators-4000,
litef_ensemble = lgb.LGBMClassifier(n_estimators=3000, n_jobs=6, num_leaves=32, objective='multiclass')
litef_ensemble.fit(X, Y)
test = pd.read_csv('data/test_values.csv', index_col='building_id')
test.head()
# +
def oneHot(df, column_name):
one_hot = pd.get_dummies(df[column_name])
df = df.drop(column_name,axis = 1)
df = df.join(one_hot)
for letter in one_hot.columns:
df[column_name + "_" + letter] = df[letter]
df.drop(letter, axis=1, inplace=True)
return df
categorical_vars = ["foundation_type", "land_surface_condition", "roof_type", "ground_floor_type", "other_floor_type", "position", "plan_configuration", "legal_ownership_status"]
for var in categorical_vars:
test = oneHot(test, var)
# +
print(len(test.columns), len(columns))
for i in range (len(test.columns)):
if test.columns[i] != columns[i]:
print(test.columns[i], columns[i])
# -
test = test.reindex(columns=columns)
test1 = test.to_numpy()
test_pred = litef_ensemble.predict(test1)
submission_format = pd.read_csv('data/submission_format.csv', index_col='building_id')
my_submission = pd.DataFrame(data=test_pred,
columns=submission_format.columns,
index=submission_format.index)
my_submission.head()
my_submission.to_csv('data/submission2.csv')
param_leaves = {
'num_leaves':[32, 50, 90]
}
gsearch = GridSearchCV(estimator = lgb.LGBMClassifier(n_estimators=3000, n_jobs=6),
param_grid = param_leaves, scoring='f1_micro', n_jobs=6, cv=5)
gsearch.fit(X_train, y_train)
print(f1_score(y_train, gsearch.predict(X_train), average='micro'))
print(f1_score(y_test, gsearch.predict(X_test), average='micro'))
gsearch.best_estimator_
joblib.dump(gsearch, "decision_models/gsearch-1.model")
# 'num_leaves':[32, 50, 90], n_estimators=3000
# in the future, will use fewer estimators for figuring out optimal parameter, then run full training
| lightgbmtree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parser Example
#
# A brief tutorial for the `ctwrap.Parser` object, which parses YAML defined quantities within simulation modules (and also handles unit conversions).
from ruamel import yaml
from ctwrap import Parser
# ### 1. Load a YAML File
# +
# load configuration from a yaml file
with open('ignition.yaml') as yml:
data = yaml.load(yml, Loader=yaml.SafeLoader)
config = data['defaults']
# -
# this is a conventional dictionary (two levels)
config
# ### 2. Create Parser Object
# a parser object takes a dictionary as input
initial = Parser(config['initial'])
initial
# and still acts much like a dictionary
initial.keys()
# ### 3. Access Values with Units
#
# Unit manipulations are based on the Python package `pint`
# as dictionary items
initial['T'], initial['P'], initial['phi']
# as attributes
initial.T, initial.P, initial.phi
# convert to desired unit
initial.P.to('pascal')
# convert to desired unit (magnitude)
initial.P.m_as('pascal')
# ### 4. Access Values without Units
# values that do not have units
initial.fuel, initial.oxidizer
# ### 5. Access Raw Values
initial.raw['P']
initial.raw['oxidizer']
| examples/parser_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/baranikannan/python-scripts/blob/master/colab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="mqnBoY0Vargk" colab_type="code" colab={}
# Install useful stuff
# ! apt install --yes ssh screen nano htop ranger git > /dev/null
# SSH setting
# ! echo "root:carbonara" | chpasswd
# ! echo "PasswordAuthentication yes" > /etc/ssh/sshd_config
# ! echo "PermitUserEnvironment yes" >> /etc/ssh/sshd_config
# ! echo "PermitRootLogin yes" >> /etc/ssh/sshd_config
# ! service ssh restart > /dev/null
# Download ngrok
# ! wget -q -c -nc https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
# ! unzip -qq -n ngrok-stable-linux-amd64.zip
# ! mkdir ~/.ssh
# ! echo "ssh-rsa AAAAB3NzaC1yc2EAAAADAQABAAACAQDFREI5+XPdlcA+Rd+YG4a6vQTFu3ltXdo//tQP/R3b3sXnGEKOfhoYjOug7ubC0VQ7IaA8pZ/rt3uB/ACPqTSjIdL02B8c8//BoPKhv8XSTiW+9WWAH4EtWoFav4TPn1GMGes+AvwiDzv5YXqxkHYXaiK3Dgai76wKQJQ9SBp2zbHEC+O2B0Yi9j0hz9DOFWiNpLMqb2EbhA5pexGN0isQtyKm2UnVS0vh5YnTSRb5eRItD6QqGSXmbsNr7tAYqjgAPW9ZK1mdxu0xMTIq/cZycIFvkQzPDA44AUU1bzFQZMT3lF74HptO7Ns4lrdaUKys25+jECbLqQIml0O9olSGue+gsz6LBj3lPxwpKy2+xWirf9TFs32la2f8nRAxRPtxJnP5TNiMpRhe2epR1NsGRQEhIhZMWSfQFUBGLeei1mYYVnNN5St+6uGpFvFOoJmgJycazB7ijd641l4V9BXFPi8+v8WAvSkjIjONlBfjLXLFmZ3JZ0aySVB5w1kY+H3+FOs5PqYa/lD/pw2XCsrjtyn7u+xw/xJ3cwFPMHBDHi5KGeLcSIG0BA/7Tm0iD5JNj4QLQXMPDE+zZaENNnrFPL7Ol+9RWxsdTBEuzkYfUGigIP7h0JU69pmZnbEBfXKzoFZ4rODunFzyKD6yXkPmPhI0t7LSRj2ixfT5op1MEw== root" > ~/.ssh/authorized_keys
# Run ngrok
authtoken = ""
get_ipython().system_raw('./ngrok authtoken $authtoken && ./ngrok tcp 22 &')
# ! sleep 3
# Get the address for SSH
import requests
from re import sub
r = requests.get('http://localhost:4040/api/tunnels')
str_ssh = r.json()['tunnels'][0]['public_url']
str_ssh = sub("tcp://", "", str_ssh)
str_ssh = sub(":", " -p ", str_ssh)
str_ssh = "ssh root@" + str_ssh
print(str_ssh)
# + id="Le3dTSw4a0GU" colab_type="code" colab={}
import time
import sys
for i in range(20000):
time.sleep(0.5)
print(f"\rNumber {i}",end="")
sys.stdout.flush()
# + id="Rx9wOf1SuHR2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="6fc77c05-3b4d-4849-d56c-c099d8a21ab1"
# !ps auxwww | grep ngrok
# + id="teSBC_J10XY5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="801a7659-f52a-4a68-e87b-17b0b439ac5f"
# !netstat -tnap | grep -i listen
# + id="a2m0lvQl0crV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="0c656abe-b275-4ea7-f98e-15707b7c7cec"
# !shutdown
# + id="k2MgASDr0gtB" colab_type="code" colab={}
# !apt update
# + id="BieA0aoC0i4z" colab_type="code" colab={}
| colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unsupervised learning
# ### AutoEncoders
# An autoencoder, is an artificial neural network used for learning efficient codings.
#
# The aim of an autoencoder is to learn a representation (encoding) for a set of data, typically for the purpose of dimensionality reduction.
# <img src="imgs/autoencoder.png" width="25%">
# Unsupervised learning is a type of machine learning algorithm used to draw inferences from datasets consisting of input data without labeled responses. The most common unsupervised learning method is cluster analysis, which is used for exploratory data analysis to find hidden patterns or grouping in data.
# +
# based on: https://blog.keras.io/building-autoencoders-in-keras.html
encoding_dim = 32
input_img = Input(shape=(784,))
encoded = Dense(encoding_dim, activation='relu')(input_img)
decoded = Dense(784, activation='sigmoid')(encoded)
autoencoder = Model(input=input_img, output=decoded)
encoder = Model(input=input_img, output=encoded)
encoded_input = Input(shape=(encoding_dim,))
decoder_layer = autoencoder.layers[-1]
decoder = Model(input=encoded_input, output=decoder_layer(encoded_input))
autoencoder.compile(optimizer='adadelta', loss='binary_crossentropy')
(x_train, _), (x_test, _) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
x_train = x_train.reshape((len(x_train), np.prod(x_train.shape[1:])))
x_test = x_test.reshape((len(x_test), np.prod(x_test.shape[1:])))
#note: x_train, x_train :)
autoencoder.fit(x_train, x_train,
nb_epoch=50,
batch_size=256,
shuffle=True,
validation_data=(x_test, x_test))
# -
# #### Testing the Autoencoder
# +
encoded_imgs = encoder.predict(x_test)
decoded_imgs = decoder.predict(encoded_imgs)
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# original
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# reconstruction
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# -
# #### Sample generation with Autoencoder
# +
encoded_imgs = np.random.rand(10,32)
decoded_imgs = decoder.predict(encoded_imgs)
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# generation
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# -
# #### Pretraining encoders
# One of the powerful tools of auto-encoders is using the encoder to generate meaningful representation from the feature vectors.
# +
# Use the encoder to pretrain a classifier
# -
# ---
# # Natural Language Processing using Artificial Neural Networks
# > “In God we trust. All others must bring data.” – <NAME>, statistician
# # Word Embeddings
#
# ### What?
# Convert words to vectors in a high dimensional space. Each dimension denotes an aspect like gender, type of object / word.
#
# "Word embeddings" are a family of natural language processing techniques aiming at mapping semantic meaning into a geometric space. This is done by associating a numeric vector to every word in a dictionary, such that the distance (e.g. L2 distance or more commonly cosine distance) between any two vectors would capture part of the semantic relationship between the two associated words. The geometric space formed by these vectors is called an embedding space.
#
#
# ### Why?
# By converting words to vectors we build relations between words. More similar the words in a dimension, more closer their scores are.
#
# ### Example
# _W(green) = (1.2, 0.98, 0.05, ...)_
#
# _W(red) = (1.1, 0.2, 0.5, ...)_
#
# Here the vector values of _green_ and _red_ are very similar in one dimension because they both are colours. The value for second dimension is very different because red might be depicting something negative in the training data while green is used for positiveness.
#
# By vectorizing we are indirectly building different kind of relations between words.
# ## Example of `word2vec` using gensim
from gensim.models import word2vec
from gensim.models.word2vec import Word2Vec
# ### Reading blog post from data directory
import os
import pickle
DATA_DIRECTORY = os.path.join(os.path.abspath(os.path.curdir), 'data', 'word_embeddings')
male_posts = []
female_post = []
# +
with open(os.path.join(DATA_DIRECTORY,"male_blog_list.txt"),"rb") as male_file:
male_posts= pickle.load(male_file)
with open(os.path.join(DATA_DIRECTORY,"female_blog_list.txt"),"rb") as female_file:
female_posts = pickle.load(female_file)
# -
print(len(female_posts))
print(len(male_posts))
filtered_male_posts = list(filter(lambda p: len(p) > 0, male_posts))
filtered_female_posts = list(filter(lambda p: len(p) > 0, female_posts))
posts = filtered_female_posts + filtered_male_posts
print(len(filtered_female_posts), len(filtered_male_posts), len(posts))
# ## Word2Vec
w2v = Word2Vec(size=200, min_count=1)
w2v.build_vocab(map(lambda x: x.split(), posts[:100]), )
w2v.vocab
w2v.similarity('I', 'My')
print(posts[5])
w2v.similarity('ring', 'husband')
w2v.similarity('ring', 'housewife')
w2v.similarity('women', 'housewife') # Diversity friendly
# ## Doc2Vec
#
# The same technique of word2vec is extrapolated to documents. Here, we do everything done in word2vec + we vectorize the documents too
import numpy as np
# 0 for male, 1 for female
y_posts = np.concatenate((np.zeros(len(filtered_male_posts)),
np.ones(len(filtered_female_posts))))
len(y_posts)
# # Convolutional Neural Networks for Sentence Classification
# Train convolutional network for sentiment analysis.
#
# Based on
# "Convolutional Neural Networks for Sentence Classification" by <NAME>
# http://arxiv.org/pdf/1408.5882v2.pdf
#
# For 'CNN-non-static' gets to 82.1% after 61 epochs with following settings:
# embedding_dim = 20
# filter_sizes = (3, 4)
# num_filters = 3
# dropout_prob = (0.7, 0.8)
# hidden_dims = 100
#
# For 'CNN-rand' gets to 78-79% after 7-8 epochs with following settings:
# embedding_dim = 20
# filter_sizes = (3, 4)
# num_filters = 150
# dropout_prob = (0.25, 0.5)
# hidden_dims = 150
#
# For 'CNN-static' gets to 75.4% after 7 epochs with following settings:
# embedding_dim = 100
# filter_sizes = (3, 4)
# num_filters = 150
# dropout_prob = (0.25, 0.5)
# hidden_dims = 150
#
# * it turns out that such a small data set as "Movie reviews with one
# sentence per review" (<NAME> Lee, 2005) requires much smaller network
# than the one introduced in the original article:
# - embedding dimension is only 20 (instead of 300; 'CNN-static' still requires ~100)
# - 2 filter sizes (instead of 3)
# - higher dropout probabilities and
# - 3 filters per filter size is enough for 'CNN-non-static' (instead of 100)
# - embedding initialization does not require prebuilt Google Word2Vec data.
# Training Word2Vec on the same "Movie reviews" data set is enough to
# achieve performance reported in the article (81.6%)
#
# ** Another distinct difference is slidind MaxPooling window of length=2
# instead of MaxPooling over whole feature map as in the article
# +
import numpy as np
import word_embedding
from word2vec import train_word2vec
from keras.models import Sequential, Model
from keras.layers import (Activation, Dense, Dropout, Embedding,
Flatten, Input,
Conv1D, MaxPooling1D)
from keras.layers.merge import Concatenate
np.random.seed(2)
# -
# ### Parameters
#
# Model Variations. See <NAME>'s Convolutional Neural Networks for
# Sentence Classification, Section 3 for detail.
model_variation = 'CNN-rand' # CNN-rand | CNN-non-static | CNN-static
print('Model variation is %s' % model_variation)
# Model Hyperparameters
sequence_length = 56
embedding_dim = 20
filter_sizes = (3, 4)
num_filters = 150
dropout_prob = (0.25, 0.5)
hidden_dims = 150
# Training parameters
batch_size = 32
num_epochs = 100
val_split = 0.1
# Word2Vec parameters, see train_word2vec
min_word_count = 1 # Minimum word count
context = 10 # Context window size
# ### Data Preparation
# +
# Load data
print("Loading data...")
x, y, vocabulary, vocabulary_inv = word_embedding.load_data()
if model_variation=='CNN-non-static' or model_variation=='CNN-static':
embedding_weights = train_word2vec(x, vocabulary_inv,
embedding_dim, min_word_count,
context)
if model_variation=='CNN-static':
x = embedding_weights[0][x]
elif model_variation=='CNN-rand':
embedding_weights = None
else:
raise ValueError('Unknown model variation')
# -
# Shuffle data
shuffle_indices = np.random.permutation(np.arange(len(y)))
x_shuffled = x[shuffle_indices]
y_shuffled = y[shuffle_indices].argmax(axis=1)
print("Vocabulary Size: {:d}".format(len(vocabulary)))
# ### Building CNN Model
# +
graph_in = Input(shape=(sequence_length, embedding_dim))
convs = []
for fsz in filter_sizes:
conv = Conv1D(filters=num_filters,
filter_length=fsz,
padding='valid',
activation='relu',
strides=1)(graph_in)
pool = MaxPooling1D(pool_length=2)(conv)
flatten = Flatten()(pool)
convs.append(flatten)
if len(filter_sizes)>1:
out = Concatenate()(convs)
else:
out = convs[0]
graph = Model(input=graph_in, output=out)
# main sequential model
model = Sequential()
if not model_variation=='CNN-static':
model.add(Embedding(len(vocabulary), embedding_dim, input_length=sequence_length,
weights=embedding_weights))
model.add(Dropout(dropout_prob[0], input_shape=(sequence_length, embedding_dim)))
model.add(graph)
model.add(Dense(hidden_dims))
model.add(Dropout(dropout_prob[1]))
model.add(Activation('relu'))
model.add(Dense(1))
model.add(Activation('sigmoid'))
# +
model.compile(loss='binary_crossentropy', optimizer='rmsprop',
metrics=['accuracy'])
# Training model
# ==================================================
model.fit(x_shuffled, y_shuffled, batch_size=batch_size,
nb_epoch=num_epochs, validation_split=val_split, verbose=2)
# -
# # Another Example
#
# Using Keras + [**GloVe**](http://nlp.stanford.edu/projects/glove/) - **Global Vectors for Word Representation**
# ## Using pre-trained word embeddings in a Keras model
#
# **Reference:** [https://blog.keras.io/using-pre-trained-word-embeddings-in-a-keras-model.html]()
| day03/3.1 AutoEncoders and Embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Joint
from thinkbayes2 import *
import thinkplot
import numpy as np
from matplotlib import pyplot as plt
# +
from math import cos, atan2
# 改善后的模型,增加对射击者的擅长距离建模
class Position(Suite, Joint):
def __init__(self, x_length, y_length, sample_step, best = 20, std = 10):
self.best = best
self.std = std
Pmf.__init__(self)
# Joint.__init__(self)
xs = np.linspace(0, x_length, num=int((x_length)/sample_step+1), endpoint=True)
ys = np.linspace(sample_step, y_length, num=int((y_length)/sample_step), endpoint=True)
for i in xs:
for j in ys:
self.Set((i,j), 1)
self.Normalize()
def Likelihood(self, data, hypo):
x_hit = data
x, y = hypo
theta = atan2(abs(x_hit-x), y)
strafing_speed = y / cos(theta)**2
return (1/strafing_speed) * scipy.stats.norm.pdf(np.linalg.norm((x_hit-x, y)), self.best, self.std)
# -
shooter = Position(50,50,1)
shooter.UpdateSet((5,15,16,18,18.1,21,26.4, 45))
# shooter.Print()
thinkplot.Contour(shooter, contour=False, pcolor=True)
plt.show()
# shooter.Print()
# thinkplot.Contour(shooter, pcolor=True)
# plt.show()
x_margin = shooter.Marginal(0, label="X")
thinkplot.Pdf(x_margin)
y_margin = shooter.Marginal(1, label="Y")
thinkplot.Pdf(y_margin)
thinkplot.Config(legend = True)
plt.show()
# # 边缘概率
thinkplot.Cdfs((x_margin.MakeCdf("X Position Margin Distribute"),
y_margin.MakeCdf("Y Position Margin Distribute")))
print(x_margin.CredibleInterval())
print(y_margin.CredibleInterval())
thinkplot.Config(legend=True)
plt.show()
# # 条件概率
for y in [5,15,45]:
con = shooter.Conditional(0,1,y)
con.label = str(y)
thinkplot.Pdf(con)
thinkplot.Config(legend = True)
plt.show()
# +
def MakeCrediblePlot(suite):
"""Makes a plot showing several two-dimensional credible intervals.
suite: Suite
"""
d = dict((pair, 0) for pair in suite.Values())
percentages = [75, 50, 25, 10, 5]
for p in percentages:
interval = suite.MaxLikeInterval(p)
for pair in interval:
d[pair] += 1
thinkplot.Contour(d, contour=False, pcolor=True)
plt.show()
MakeCrediblePlot(shooter)
| code/.ipynb_checkpoints/0Joint-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import sys
sys.executable
# + pycharm={"is_executing": true}
import torch
from utils import set_seed, set_plt_style
set_seed(42)
set_plt_style()
device = torch.device("cuda" if torch.cuda.is_available else "cpu")
# +
from dataset import make_loaders
batch_size = 64
test_size = val_size = 0.2
train_loader, val_loader, test_loader = make_loaders(batch_size, val_size, test_size)
# +
from models import *
from utils import train, plot_confusion_matrix
model = MalConvPlus(8, 4096, 128, 32, device).to(device)
train(model, train_loader, val_loader, device, "malconv_plus")
plot_confusion_matrix(model, test_loader, "malconv_plus", device)
# -
model = MalConvBase(8, 4096, 128, 32).to(device)
train(model, train_loader, val_loader, device, "malconv_base")
plot_confusion_matrix(model, test_loader, "malconv_base", device)
model = MalConvBase(16, 4096, 128, 32).to(device)
train(model, train_loader, val_loader, device, "malconv_base_emb16")
plot_confusion_matrix(model, test_loader, "malconv_base_emb16", device)
model = MalConvBase(8, 4096, 256, 32).to(device)
train(model, train_loader, val_loader, device, "malconv_base_conv256")
plot_confusion_matrix(model, test_loader, "malconv_base_conv256", device)
model = MalConvBase(8, 4096, 128, 64).to(device)
train(model, train_loader, val_loader, device, "malconv_base_win64")
plot_confusion_matrix(model, test_loader, "malconv_base_win64", device)
# Original: 92 (33, 59),
# Conv256: 93 (34, 59),
# Emb16: 94 (34, 60),
# Win64: 94 (34, 60)
model = MalConvBase(16, 4096, 128, 64).to(device)
train(model, train_loader, val_loader, device, "malconv_base_emb16win64", patience=5)
plot_confusion_matrix(model, test_loader, "malconv_base_emb16win64", device)
model = MalConvPlus(16, 4096, 128, 32).to(device)
train(model, train_loader, val_loader, device, "malconv_plus_emb16")
plot_confusion_matrix(model, test_loader, "malconv_plus_emb16", device)
model = RCNN(8, 128, 32, torch.nn.GRU, 256, 1, False, False).to(device)
train(model, train_loader, val_loader, device, "gru_base")
plot_confusion_matrix(model, test_loader, "gru_base", device)
model = RCNN(8, 128, 32, torch.nn.GRU, 256, 1, True, False).to(device)
train(model, train_loader, val_loader, device, "gru_bidir")
plot_confusion_matrix(model, test_loader, "gru_bidir", device)
model = RCNN(8, 128, 32, torch.nn.GRU, 256, 1, False, True).to(device)
train(model, train_loader, val_loader, device, "gru_res")
plot_confusion_matrix(model, test_loader, "gru_res", device)
model = RCNN(8, 128, 64, torch.nn.GRU, 256, 1, False, True).to(device)
train(model, train_loader, val_loader, device, "gru_wind64res")
plot_confusion_matrix(model, test_loader, "gru_win64res", device)
model = RCNN(8, 128, 32, torch.nn.GRU, 128, 1, False, False).to(device)
train(model, train_loader, val_loader, device, "gru_hid128")
plot_confusion_matrix(model, test_loader, "gru_hid128", device)
| run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Density of States Analysis Example
#
# This example demonatrates a routine procedure of calculating phonon density of states
# from an experimental NeXus data file for a powder vanadium sample measured at ARCS,
# a direct-geometry neutron chopper spectrometer at the Spallation Neutron Source (SNS),
# Oak Ridge National Lab.
#
# ## Summary of processing steps
#
# * Gather experimental information and experimental raw data
# * Reduce raw data to S(Q,E), the experimental dynamical structure factor, and inspect
# * Convert S(Q,E) to phonon DOS
# ## Preparation
# Get python tools ready. This may takes a while
import os, numpy as np
import histogram.hdf as hh, histogram as H
from matplotlib import pyplot as plt
# %matplotlib notebook
# # %matplotlib inline
import mantid
from multiphonon import getdos
from multiphonon.sqe import plot as plot_sqe
# Create a new working directory and change into it. All inputs, intermediate results and final outputs will be in this new directory.
projectdir = os.path.abspath('./V_Ei120meV-noUI')
# !mkdir -p {projectdir}
# %cd {projectdir}
# ## Get experimental data
#
# For SNS users, experimental data are available in /SNS/"instrument_name"/IPTS-#### folders at the SNS analysis cluster.
# Here we will download the required data file from web.
# Build download command
dest = 'ARCS_V_annulus.nxs'
url = "https://mcvine.ornl.gov/multiphonon/ARCS_V_annulus.nxs"
cmd = 'wget %r -O %r' % (url, dest)
print cmd
# Download: this will take a while (can be a few minutes to an hour, depending on internet speed)
# %%time
# !{cmd} >log.download 2>err.download
# The following command should show the downloaded file "ARCS_V_annulus.nxs"
# ls
# ## Experimental data and condition
# To start, we need to set the locations of the data files measured for the sample and empty can (for background correction), as well as the experimental conditions such as incident neutron energy (Ei, in meV) and sample temperature (T, in Kelvin).
# The example inputs explained:
#
# * samplenxs: ARCS_V_annulus.nxs we just downloaded
# * mtnxs: None. This means we will skip the empty can background correction for this example.
# * Ei: 120. This is set by Fermi chopper settings during the experiment. An approximate number is fine. The actual Ei will be caculated from the experimental NeXus file.
# * T: 300. This is set by sample environment. For room temperature measurement, use 300 (K).
samplenxs = './ARCS_V_annulus.nxs'
mtnxs = None
Ei = 120
T = 300
# ## Obtain S(Q,E)
#
# Now we are ready to reduce the experimental data to obtain the dynamical structure factor, S(Q,E).
#
# S(Q,E) spectra for both the sample and the empty can is the starting point for getdos processing.
#
# The Q and E axes need to be define:
#
# * E axis
# - Emin: -115. Usually -Ei
# - Emax: 115. Usually slightly smaller than Ei
# - dE: 1. Usually Ei/100
# * Q axis
# - Qmin: 0. Usually 0
# - Qmax: 17. Usually 2 X E2Q(Ei)
# - dQ: 0.1. Usually Emax/100
Qaxis = Qmin, Qmax, dQ = 0, 17, 0.1
Eaxis = Emin, Emax, dE = -115., 115., 1.
workdir = 'work'
iqe_h5 = 'iqe.h5'
from multiphonon import getdos
# %%time
for m in getdos.reduce2iqe(samplenxs,
Emin=Emin, Emax=Emax, dE=dE, Qmin=Qmin, Qmax=Qmax, dQ=dQ,
iqe_h5=iqe_h5, workdir=workdir):
print m
# ls -tl {workdir}/{iqe_h5}
# Plot sample IQE
# +
iqe = hh.load(os.path.join(workdir, iqe_h5))
plt.figure(figsize=(6,4))
plot_sqe(iqe)
# plt.xlim(0, 11)
plt.clim(0, 3e-3)
# -
# This is a plot of vanadium S(Q, E) histogram.
# * The colored region is within the dynamical range of the measurement
# * Vanadium is incoherent, therefore the intensity is mostly momentum-independent
# * Make sure the energy and momentum transfer axes are reasonable so that the S(Q,E) spectrum looks reasonable
# * You can improve the Q,E axis parameters if you like, by re-executing the relevant cells above
# Now integreate over the Q (momentum transfer) axis to obtain energy spectrum I(E)
iqe2 = iqe.copy()
I = iqe2.I; I[I!=I] = 0 # remove NaNs
IE = iqe2.sum('Q') # sum over Q
plt.figure(figsize=(6,4))
plt.plot(IE.energy, IE.I)
# * At the center of this plot there is an enormous peak that is due to elastic scattering, which should be excluded from the phonon DOS calculation
# * Zoom in to see the rough range of the elastic peak and take notes. We need them in the analysis below.
# ## Run GetDOS
#
# Phonon DOS will be obtained from SQE histogram by an iterative procedure where multiphonon and multiple scattering corrections are applied to the measured SQE spectrum, assuming
# incoherent approximation, and the corrected spectrum
# is then converted to DOS.
#
# Input parameters
# - Emin, Emax of elastic peak: -15, 7. Make an estimate from the I(E) spectrum
# - Average atomic mass: 50.94. Atomic mass of vanadium
# - mt_fraction: 0.9. Depends on the geometrical property of the sample and the empty can. Usually between 0.9 and 1.
# - Ecutoff: Max phonon energy. 40meV. This is also used as the "stiching point" if multiple Ei datasets are combined.
# - C_ms: 0.26: Ratio of multiple scattering to multiphon scattering. Depends on sample shape.
# - const_bg_fraction: 0.004: Background noise level.
# - initdos: leave it as None for standard DOS analysis. If working with multiple Ei datasets, this should be the DOS histgram obtained from larger Ei.
#
for msg in getdos.getDOS(
samplenxs, mt_fraction=0.9, const_bg_fraction=0.004,
Emin=Emin, Emax=Emax, dE=dE, Qmin=Qmin, Qmax=Qmax, dQ=dQ,
T=300., Ecutoff=40.,
elastic_E_cutoff=(-15, 7.),
M=50.94,
C_ms = 0.26,
Ei = 120,
initdos=None,
workdir = workdir,
):
print msg
# ## Check output
# Results are saved in "work" directory
# ls {workdir}/
# Plot the final result for DOS
dos = hh.load(os.path.join(workdir, 'final-dos.h5'))
plt.figure(figsize=(5,3))
plt.plot(dos.E, dos.I)
plt.xlabel('Energy (meV)')
plt.xlim(0, 50)
plt.tight_layout()
# More plotting utils are available
from multiphonon.backward import plotutils as pu
plt.figure(figsize=(5,3))
pu.plot_dos_iteration(workdir)
plt.xlim(0, 50)
plt.figure(figsize=(6,4))
pu.plot_residual(workdir)
plt.figure(figsize=(8, 4))
pu.plot_intermediate_result_se(os.path.join(workdir, 'round-4'))
| examples/getdos2-V_Ei120meV-noUI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../scripts/')
from dp_policy_agent import *
import random, copy
class StateInfo: #q4stateinfo
def __init__(self, action_num, epsilon=0.3):
self.q = np.zeros(action_num)
self.epsilon = epsilon
def greedy(self):
return np.argmax(self.q)
def epsilon_greedy(self, epsilon):
if random.random() < epsilon:
return random.choice(range(len(self.q)))
else:
return self.greedy()
def pi(self):
return self.epsilon_greedy(self.epsilon)
def max_q(self): #追加
return max(self.q)
class SarsaAgent(PuddleIgnoreAgent): #名前を変えましょう ###sarsa1
def __init__(self, time_interval, particle_pose, envmap, puddle_coef=100, alpha=0.5, \
motion_noise_stds={"nn":0.19, "no":0.001, "on":0.13, "oo":0.2}, widths=np.array([0.2, 0.2, math.pi/18]).T, \
lowerleft=np.array([-4, -4]).T, upperright=np.array([4, 4]).T): #alpha追加
super().__init__(time_interval, particle_pose, envmap, None, puddle_coef, motion_noise_stds)
###DynamicProgrammingから持ってくる###
self.pose_min = np.r_[lowerleft, 0]
self.pose_max = np.r_[upperright, math.pi*2]
self.widths = widths
self.index_nums = ((self.pose_max - self.pose_min)/self.widths).astype(int)
nx, ny, nt = self.index_nums
self.indexes = list(itertools.product(range(nx), range(ny), range(nt)))
###PuddleIgnorePolicyの方策と価値関数の読み込み###
self.actions, self.ss = self.set_action_value_function()
###強化学習用変数### #追加
self.alpha = alpha
self.s, self.a = None, None
self.update_end = False
self.step = 0
def set_action_value_function(self):
policy = np.zeros(np.r_[self.index_nums,2])
for line in open("puddle_ignore_policy.txt", "r"):
d = line.split()
policy[int(d[0]), int(d[1]), int(d[2])] = [float(d[3]), float(d[4])]
actions = list(set([tuple(policy[i]) for i in self.indexes]))
action_num = len(actions)
ss = {}
for line in open("puddle_ignore_values.txt", "r"):
d = line.split()
index, value = (int(d[0]), int(d[1]), int(d[2])), float(d[3])
ss[index] = StateInfo(action_num)
for i, a in enumerate(actions):
ss[index].q[i] = value if tuple(policy[index]) == a else value - 0.1
ss[index].q[i] *= 10 #大きくしておく
return actions, ss
def policy(self, pose): ###q4policy
index = np.floor((pose - self.pose_min)/self.widths).astype(int)
index[2] = (index[2] + self.index_nums[2]*1000)%self.index_nums[2]
for i in [0,1]:
if index[i] < 0: index[i] = 0
elif index[i] >= self.index_nums[i]: index[i] = self.index_nums[i] - 1
s = tuple(index) #Q学習の式で使う記号に変更
a = self.ss[s].pi()
return s, a #行動は値でなくインデックスで返す
def decision(self, observation=None):###q4decision
if self.step%100000 == 0:
with open("tmp/sarsa0_policy"+ str(self.step)+ ".txt", "w") as f: ###
for index in self.indexes:
p = self.actions[self.ss[index].greedy()]
f.write("{} {} {} {} {}\n".format(index[0], index[1], index[2], p[0], p[1]))
f.flush()
with open("tmp/sarsa0_value"+ str(self.step)+ ".txt", "w") as f: ###
for index in self.indexes:
f.write("{} {} {} {}\n".format(index[0], index[1], index[2], self.ss[index].max_q()))
f.flush()
###終了処理###
if self.update_end:
return 0.0, 0.0
if self.in_goal:
self.update_end = True #ゴールに入った後も一回だけ更新があるので即終了はしない
###カルマンフィルタの実行###
self.kf.motion_update(self.prev_nu, self.prev_omega, self.time_interval)
self.kf.observation_update(observation)
###行動決定と報酬の処理###
s_, a_ = self.policy(self.kf.belief.mean)
r = self.time_interval*self.reward_per_sec()
self.total_reward += r
###Q学習と現在の状態と行動の保存###
self.q_update(r, s_, a_) #a_も必要になる
self.s, self.a = s_, a_
###出力###
self.prev_nu, self.prev_omega = self.actions[a_]
self.step += 1
return self.actions[a_]
def q_update(self, r, s_, a_):###sarsa1
if self.s == None:
return
q = self.ss[self.s].q[self.a]
q_ = self.final_value if self.in_goal else self.ss[s_].q[a_] #max_qからQ(s_,a_)に書き換え
self.ss[self.s].q[self.a] = (1.0 - self.alpha)*q + self.alpha*(r + q_)
class WarpRobot(Robot): ###q5warprobot
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self.init_agent = copy.deepcopy(self.agent) #エージェントのコピーを残しておく
def choose_pose(self): #初期位置をランダムに決めるメソッド(雑)
xy = random.random()*8-4
t = random.random()*2*math.pi
return np.array([4, xy, t]).T if random.random() > 0.5 else np.array([xy, 4, t]).T
def reset(self):
#ssだけ残してエージェントを初期化
tmp = self.agent.ss
t = self.agent.step
self.agent = copy.deepcopy(self.init_agent)
self.agent.ss = tmp
self.agent.step = t
#初期位置をセット(ロボット、カルマンフィルタ)
self.pose = self.choose_pose()
self.agent.kf.belief = multivariate_normal(mean=self.pose, cov=np.diag([1e-10, 1e-10, 1e-10]))
#軌跡の黒い線が残らないように消す
self.poses = []
def one_step(self, time_interval):
if self.agent.update_end:
with open("log.txt", "a") as f:
f.write("{}\n".format(self.agent.total_reward + self.agent.final_value))
self.reset()
return
super().one_step(time_interval)
if __name__ == '__main__':
time_interval = 0.1
world = PuddleWorld(400000000, time_interval) #時間をほぼ無限に
m = Map()
m.append_landmark(Landmark(-4,2))
m.append_landmark(Landmark(2,-3))
m.append_landmark(Landmark(4,4))
m.append_landmark(Landmark(-4,-4))
world.append(m)
###ゴールの追加###
goal = Goal(-3,-3)
world.append(goal)
###水たまりの追加###
world.append(Puddle((-2, 0), (0, 2), 0.1))
world.append(Puddle((-0.5, -2), (2.5, 1), 0.1))
###ロボットを1台登場させる###
init_pose = np.array([3, 3, 0]).T
sa = SarsaAgent(time_interval, init_pose, m) #エージェント変更
r = WarpRobot(init_pose, sensor=Camera(m, distance_bias_rate_stddev=0, direction_bias_stddev=0),
agent=sa, color="red", bias_rate_stds=(0,0))
world.append(r)
world.draw()
#r.one_step(0.1) #デバッグ時
# +
p = np.zeros(sa.index_nums[0:2])
for x in range(sa.index_nums[0]):
for y in range(sa.index_nums[1]):
a = sa.ss[(x,y,22)].greedy()
p[x,y] = sa.actions[a][0] + sa.actions[a][1]
import seaborn as sns
sns.heatmap(np.rot90(p), square=False)
plt.show()
# +
v = np.zeros(r.agent.index_nums[0:2])
for x in range(r.agent.index_nums[0]):
for y in range(r.agent.index_nums[1]):
v[x,y] = r.agent.ss[(x,y,18)].max_q()
import seaborn as sns
sns.heatmap(np.rot90(v), square=False)
plt.show()
# -
with open("sarsa_policy0.txt", "w") as f: ###
for index in sa.indexes:
p = sa.actions[sa.ss[index].greedy()]
f.write("{} {} {} {} {}\n".format(index[0], index[1], index[2], p[0], p[1]))
# #
| section_reinforcement_learning/sarsa1_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DAT257x: Reinforcement Learning Explained
#
# ## Lab 2: Bandits
#
# ### Exercise 2.1B: Round Robin
# +
import numpy as np
import sys
if "../" not in sys.path:
sys.path.append("../")
from lib.envs.bandit import BanditEnv
from lib.simulation import Experiment
# -
#Policy interface
class Policy:
#num_actions: (int) Number of arms [indexed by 0 ... num_actions-1]
def __init__(self, num_actions):
self.num_actions = num_actions
def act(self):
pass
def feedback(self, action, reward):
pass
#Greedy policy
class Greedy(Policy):
def __init__(self, num_actions):
Policy.__init__(self, num_actions)
self.name = "Greedy"
self.total_rewards = np.zeros(num_actions, dtype = np.longdouble)
self.total_counts = np.zeros(num_actions, dtype = np.longdouble)
def act(self):
current_averages = np.divide(self.total_rewards, self.total_counts, where = self.total_counts > 0)
current_averages[self.total_counts <= 0] = 0.5 #Correctly handles Bernoulli rewards; over-estimates otherwise
current_action = np.argmax(current_averages)
return current_action
def feedback(self, action, reward):
self.total_rewards[action] += reward
self.total_counts[action] += 1
# We have seen in the previous exercise that a greedy policy can lock into sub-optimal action. Could it be worse than a simple round-robin selection?
# Let's implement a round robin policy: that is "pulling" the arms in round robin fashion. So for example, if you have three arms, the sequence will be arm 1, arm 2, arm 3 and then back to arm 1, and so on, until the trials finishes.
# We have given you some boiler plate code, you only need to modify the part as indicated.
class RoundRobin(Policy):
def __init__(self, num_actions):
Policy.__init__(self, num_actions)
self.name = "Round Robin"
self.total_rewards = np.zeros(num_actions, dtype = np.longdouble)
self.total_counts = np.zeros(num_actions, dtype = np.longdouble)
self.previous_action = 0 #keep track of previous action
def act(self):
"""Implement Round Robin here"""
if self.previous_action < (num_actions - 1):
current_action = self.previous_action + 1
self.previous_action = current_action
else:
current_action = 0
self.previous_action = current_action
return current_action
def feedback(self, action, reward):
self.total_rewards[action] += reward
self.total_counts[action] += 1
# Now let's run the same simulation and keep the parameters as the previous exercise.
evaluation_seed = 8026
num_actions = 5
trials = 10000
distribution = "bernoulli"
env = BanditEnv(num_actions, distribution, evaluation_seed)
agent = RoundRobin(num_actions)
experiment = Experiment(env, agent)
experiment.run_bandit(trials)
# Observe the above results. Did the round-robin beat the greedy algorithm in this case?
#
#
# Once you have answered the questions in this lab, play around with different evaluation_seed and/or num_actions. Essentially creating a different version of the BanditEnv environment. Run the simulation and observe the results.
#
| Module 2/Ex2.1B Round Robin-completed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import matplotlib.pyplot as plt
import matplotlib as mpl
import numpy as np
import pickle
import boto
import pandas as pd
import seaborn as sns
import mpld3
from bokeh import *
from ggplot import *
# %matplotlib inline
# ## Get Data
# +
# # %%timeit
# import pickle
# inFile=open('DUMP_stuff.dat','r')
# mentionCounter=pickle.load(inFile)
# hashTagCounter=pickle.load(inFile)
# taggedHashTagCounter=pickle.load(inFile)
# domainCounter=pickle.load(inFile)
# topicCounter=pickle.load(inFile)
# timeCounter=pickle.load(inFile)
# rawDomainCounter=pickle.load(inFile)
# -
# cPickle currently approx 3 times faster than pickle
import cPickle
inFile=open('DUMP_stuff.dat','r')
mentionCounter=cPickle.load(inFile)
hashTagCounter=cPickle.load(inFile)
domainCounter=cPickle.load(inFile)
topicCounter=cPickle.load(inFile)
timeCounter=cPickle.load(inFile)
rawDomainCounter=cPickle.load(inFile)
mentionCounterFrame = pd.DataFrame(mentionCounter)
hashTagCounterFrame= pd.DataFrame(hashTagCounter)
domainCounterFrame= pd.DataFrame(domainCounter)
topicCounterFrame= pd.DataFrame(topicCounter)
timeCounterFrame= pd.DataFrame(timeCounter)
rawDomainCounterFrame= pd.DataFrame(rawDomainCounter)
mentionCounterFrame.columns = ['Account', 'Number of tweets']
hashTagCounterFrame.columns = ['Hashtag', 'Number of tweets']
domainCounterFrame.columns = ['Domain', 'Number of tweets']
topicCounterFrame.columns = ['Topic', 'Number of tweets']
timeCounterFrame.columns = ['Time', 'Number of tweets']
rawDomainCounterFrame.columns = ['URL', 'Number of tweets']
rawDomainCounterFrame = rawDomainCounterFrame.replace(to_replace='', value=np.nan)
rawDomainCounterFrame = rawDomainCounterFrame.dropna()
#rawDomainCounterFrame = rawDomainCounterFrame[pd.notnull(rawDomainCounterFrame['URL'])]
rawDomainCounterFrame.sort(columns='Number of tweets', ascending=False, inplace=True)
rawDomainCounterFrame.head()
# # Topics and Categories
from matplotlib.ticker import ScalarFormatter
formatter = ScalarFormatter()
formatter.set_scientific(False)
sns.set_context("poster")
sns.set_style("darkgrid", {'font.size': 14, 'axes.labelsize': 16, 'legend.fontsize': 14.0, 'axes.titlesize': 12, 'xtick.labelsize': 14,
'ytick.labelsize': 14})
# ## Number of Tweets by Hashtag
fig, ax = plt.subplots()
hashTagCounter.reverse()
ax.barh(range(10),[v[1] for v in hashTagCounter[-10:]],log=False,linewidth=0,alpha=0.7,color="#00aeef")
ax.set_axis_bgcolor('#efefef')
ax.xaxis.set_major_formatter(formatter)
ax.xaxis.set_major_locator(plt.MaxNLocator(4))
ax.set_yticks([i+0.5 for i in range(10)])
ax.set_xlabel('Number of Tweets')
ax.set_yticklabels(['#'+v[0] for v in hashTagCounter[-10:]]);
plt.savefig('../web/charts/hashtags.png', bbox_inches='tight',dpi=200)
# ## Number of Tweets by URL
fig, ax = plt.subplots()
rawDomainCounter.reverse()
ax.barh(range(10),[v[1] for v in rawDomainCounter[-12:-2]],log=False,linewidth=0,alpha=0.7,color="#00aeef")
ax.set_axis_bgcolor('#efefef')
ax.xaxis.set_major_formatter(formatter)
ax.xaxis.set_major_locator(plt.MaxNLocator(4))
ax.set_yticks([i+0.5 for i in range(10)])
ax.set_xlabel('Number of Tweets')
ax.set_yticklabels([v[0] for v in rawDomainCounter[-12:-2]]);
plt.savefig('../web/charts/rawdomains.png', bbox_inches='tight',dpi=200)
rawDomainFig, ax = plt.subplots()
rawDomainCounter.reverse()
ax.barh(range(10),[v[1] for v in rawDomainCounter[-12:-2]],log=False,linewidth=0,alpha=0.7,color="#00aeef")
ax.set_axis_bgcolor('#efefef')
ax.xaxis.set_major_formatter(formatter)
ax.xaxis.set_major_locator(plt.MaxNLocator(4))
ax.set_yticks([i+0.5 for i in range(10)])
ax.set_xlabel('Number of Tweets')
ax.set_yticklabels([v[0] for v in rawDomainCounter[-12:-2]]);
plt.savefig('../web/charts/rawdomains.png', bbox_inches='tight',dpi=200)
mpld3.save_html(rawDomainFig, '../charts/rawdomains.php', figid="taggedHashtagsFig")
mpld3.display(rawDomainFig)
# ## Number of Tweets by Link Domain
fig, ax = plt.subplots()
domainCounter.reverse()
ax.barh(range(10),[v[1] for v in domainCounter[-10:]],log=False,linewidth=0,alpha=0.7,color="#00aeef")
ax.set_axis_bgcolor('#efefef')
ax.xaxis.set_major_formatter(formatter)
ax.xaxis.set_major_locator(plt.MaxNLocator(4))
ax.set_yticks([i+0.5 for i in range(10)])
ax.set_xlabel('Number of Tweets')
ax.set_yticklabels([v[0] for v in domainCounter[-10:]]);
plt.savefig('../web/charts/domains.png', bbox_inches='tight',dpi=200)
# ## Number of Tweets by Tagged Topic
topicCounter
len(topicCounter[-10:])
range(9)
[v[1] for v in topicCounter[-10:]]
fig, ax = plt.subplots()
topicCounter.reverse()
ax.barh(range(4),[v[1] for v in topicCounter[-4:]],log=False,linewidth=0,alpha=0.7,color="#00aeef")
ax.set_axis_bgcolor('#efefef')
ax.xaxis.set_major_formatter(formatter)
ax.xaxis.set_major_locator(plt.MaxNLocator(4))
ax.set_yticks([i+0.5 for i in range(4)])
ax.set_xlabel('Number of Tweets')
ax.set_yticklabels([v[0] for v in topicCounter[-4:]]);
plt.savefig('../web/charts/topics.png', bbox_inches='tight',dpi=200)
sns.set_context("poster")
sns.despine()
sns.set(style="whitegrid")
sns.barplot(topicCounterFrame["Topic"],topicCounterFrame["Number of tweets"], color="#00aeef")
plt.savefig('../charts/topics_seaborn.png')
# ggplot
ggplot(aes(x="Topic", weight="Number of tweets"), topicCounterFrame) + geom_bar(fill='#00aeef')
# ## Number of Tweets by @Mentioned Account
fig, ax = plt.subplots()
mentionCounter.reverse()
ax.barh(range(10),[v[1] for v in mentionCounter[-10:]],log=False,linewidth=0,alpha=0.7,color="#00aeef")
ax.set_axis_bgcolor('#efefef')
ax.xaxis.set_major_formatter(formatter)
ax.xaxis.set_major_locator(plt.MaxNLocator(4))
ax.set_yticks([i+0.5 for i in range(10)])
ax.set_xlabel('Number of Tweets')
ax.set_yticklabels(['@'+v[0] for v in mentionCounter[-10:]]);
plt.savefig('../web/charts/mentions.png', bbox_inches='tight',dpi=200)
from IPython.core.display import HTML
styles = open("../css/custom.css", "r").read()
HTML(styles)
| ipynb/plot_stuff-RCN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle
import os
import sys
src_dir = os.path.join(os.getcwd(), '..', '..', '03-src')
sys.path.append(src_dir)
import decisionclass.decision_functions as hmd
# import pandas as pd
# from itertools import combinations
# %matplotlib inline
# -
# ## Here is an example of how to use the decision function
example = hmd.Decision(example=True)
example.option_value_df
example.print_results()
example.plot_radar2()
# example.plot_venn3()
# ## Here is an example of using the cereal decision object
f = open('../../01-data/03-decisions/cereal_decision.pkl', 'rb')
cereal_decision = pickle.load(f)
f.close
print(cereal_decision.feature_list)
important_features = ['low_sugars_per_cal', 'low_sodium_per_cal', 'protein_per_cal']
cereal_decision.feature_list_keep(important_features)
cereal_decision.print_results()
| 02-notebooks/02-decision-function/cmb_decision_function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sqlalchemy import create_engine
from sklearn.ensemble import GradientBoostingClassifier
from sklearn import metrics
# from sklearn.grid_search import GridSearchCV
# +
username = "privateuser"
password = "<PASSWORD>"
port = 7777
engine = create_engine('mysql+mysqldb://%s:%s@localhost:%i/Shutterfly'%(username, password, port))
# -
def load_dataset(split="trn_set", limit=None, ignore_categorical=False):
sql = """
SELECT o.*, f1.*, f2.*, f3.*, f4.*,
EXTRACT(MONTH FROM o.dt) AS month
FROM %s AS t
JOIN Online AS o
ON t.index = o.index
JOIN features_group_1 AS f1
ON t.index = f1.index
JOIN features_group_2 AS f2
ON t.index = f2.index
JOIN features_group_3 AS f3
ON t.index = f3.index
JOIN features_group_4 AS f4
ON t.index = f4.index
"""%split
if limit:
sql += " LIMIT %i"%limit
df = pd.read_sql_query(sql.replace('\n', " ").replace("\t", " "), engine)
df.event1 = df.event1.fillna(0)
X = df.drop(["index", "event2", "dt", "day", "session", "visitor", "custno"], axis=1)
if ignore_categorical:
X = X.drop(["last_category", "event1", "month", "last_event1",
"last_event2", "last_prodcat1", "last_prodcat2"], axis=1)
Y = df.event2
return X, Y
X_trn, Y_trn = load_dataset("tst_set", limit=10000)
print(X_trn.head(5).T)
import matplotlib.pyplot as plt
for col in ["last_category", "event1", "month", "last_event1", "last_event2", "last_prodcat1", "last_prodcat2"]:
if col in X_trn.columns:
print(col)
# +
plt.figure(figsize=(15, 15))
for i, col in enumerate(X_trn.columns):
ax = plt.subplot(8, 7, i + 1)
ax.hist(X_trn[col].dropna())
ax.set_title(col)
plt.tight_layout()
plt.savefig("figs/features.png")
| case_study/shutterfly/feature_engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
train=pd.read_csv('train.csv')
train.head(10)
# # Conditition on City Group , TYpe and P1 - cause they look like categorical
# * First issue will be to define the labels and the mappings , and the join table , this will not be hard , real issue will be to macke prob spae
group=np.unique(train['City Group'],return_counts=True)
group
types=np.unique(train['Type'],return_counts=True)
types
p1=np.unique(train['P1'],return_counts=True)
p1
# ### The condititional prob model where will be :
# * P group,type,p1
# * Meaning what is prob of p1 being some value givin the fact that group and type has some values
# * This will be first derived feature
group_labels=list(np.unique(train['City Group']))
group_labels
types_labels=list(np.unique(train['Type']))
types_labels
p1_labels=list(np.unique(train['P1']))
p1_labels
group_mapping={label: index
for index,label in enumerate(group_labels)}
group_mapping
types_mapping={label: index
for index,label in enumerate(types_labels)}
types_mapping
p1_mapping={label: index
for index,label in enumerate(p1_labels)}
p1_mapping
| ConditioningRealData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <img src="http://xarray.pydata.org/en/stable/_static/dataset-diagram-logo.png" align="right" width="30%">
#
# # Dask e Xarray para computação paralela
#
# Este notebook demonstra um dos recursos mais poderosos do xarray: a capacidade
# de trabalhar em sintonia com matrizes dask e facilmente permitir que os usuários executem o código de análise em paralelo.
#
# Até o final deste notebook, veremos:
#
# 1. Que as estruturas de dados Xarray `DataArray` e `Dataset` são parte das coleções Dask, isso é, podemos executar as funções de alto nível Dask como `dask.visualize(xarray_object)`;
# 2. Que todas as operações integradas do xarray podem usar o dask de forma transparente;
# 3. Que o Xarray fornece ferramentas para paralelizar facilmente funções personalizadas em blocos de objetos xarray apoiados em dask.
#
# ## Conteúdo
#
# 1. [Lendo dados com Dask e Xarray](#Lendo-dados-com-Dask-e-Xarray)
# 2. [Computação paralela/streaming/lazy usando dask.array com Xarray](#Computação-paralela/streaming/lazy-usando-dask.array-com-Xarray)
# 3. [Paralelização automática com apply_ufunc e map_blocks](#Paralelização-automática-com-apply_ufunc-e-map_blocks)
#
# Primeiro, vamos fazer as importações necessárias, iniciar um cluster dask e testar o painel
#
import expectexception
import numpy as np
import xarray as xr
# Primeiro, vamos configurar um `LocalCluster` usando` dask.distributed`.
#
# Você pode usar qualquer tipo de cluster dask. Esta etapa é completamente independente de
# xarray.
# +
from dask.distributed import Client
client = Client()
client
# -
# <p>👆</p> Clique no link Dashboard acima.
#
# Vamos testar se o painel está funcionando.
#
# +
import dask.array
dask.array.ones(
(1000, 4), chunks=(2, 1)
).compute() # devemos ver a atividade no painel
# -
# <a id='readwrite'></a>
#
# ## Lendo dados com Dask e Xarray
#
# O argumento `chunks` para `open_dataset` e `open_mfdataset` permite que você leia conjuntos de dados como matrizes dask. Veja https://xarray.pydata.org/en/stable/dask.html#reading-and-writing-data para mais
# detalhes.
#
ds = xr.tutorial.open_dataset(
"air_temperature",
chunks={
"lat": 25,
"lon": 25,
"time": -1,
}, # isso diz ao xarray para abrir o conjunto de dados como um array dask
)
ds
# A representação para o DataArray `air` inclui agora também a representação dask.
ds.air
ds.air.chunks
# **Dica**: Todas as variáveis em um `Dataset` _não_ necessariamente precisam ter o mesmo tamanho de blocos ao longo dimensões comuns.
#
mean = ds.air.mean("time") # nenhuma atividade no painel
mean # contém uma matriz dask
# Isso é verdadeiro para todas as operações de xarray, incluindo *slicing*
#
ds.air.isel(lon=1, lat=20)
# e operações mais complicadas...
#
# <a id='compute'></a>
#
# ## Computação paralela/*streaming*/*lazy* usando dask.array com Xarray
#
# O Xarray envolve o dask perfeitamente para que todos os cálculos sejam adiados até que explicitamente requeridos:
#
mean = ds.air.mean("time") # nenhuma atividade no painel
mean # contém uma matriz dask
# Isso é verdadeiro para todas as operações de xarray, incluindo seleção em fatias
#
timeseries = (
ds.air.rolling(time=5).mean().isel(lon=1, lat=20)
) # nenhuma atividade no painel
timeseries # contém uma matriz dask
timeseries = ds.air.rolling(time=5).mean() # nenhuma atividade no painel
timeseries # contém uma matriz dask
# ### Obtendo valores concretos de arrays dask
#
# Em algum ponto, você desejará realmente obter valores concretos do dask.
#
# Existem duas maneiras de calcular valores em matrizes dask. Esses valores concretos são
# geralmente matrizes NumPy, mas podem ser uma matriz `pydata/sparse`, por exemplo.
#
# 1. `.compute()` retorna um objeto xarray;
# 2. `.load()` substitui a matriz dask no objeto xarray por uma matriz numpy. Isso é equivalente a `ds = ds.compute()`.
#
computed = mean.compute() # atividade no painel
computed # contém agora valores reais NumPy
# Observe que `mean` ainda contém uma matriz dask
#
mean
# Mas se chamarmos `.load()`, `mean` agora conterá uma matriz numpy
mean.load()
# Vamos verificar outra vez...
#
mean
# **Dica:** `.persist()` carrega os valores na RAM distribuída. Isso é útil se
# você usará repetidamente um conjunto de dados para computação, mas é muito grande para
# carregar na memória local. Você verá uma tarefa persistente no painel.
#
# Veja https://docs.dask.org/en/latest/api.html#dask.persist para mais detalhes.
#
# ### Extraindo dados subjacentes: `.values` vs` .data`
#
# Existem duas maneiras de extrair os dados subjacentes em um objeto xarray.
#
# 1. `.values` sempre retornará uma matriz NumPy. Para objetos xarray apoiados em dask,
# isso significa que compute sempre será chamado;
# 2. `.data` retornará uma matriz Dask.
#
# #### Exercício
#
# Tente extrair um array dask de `ds.air`.
#
# +
# Seu código aqui
# -
# Agora extraia um array NumPy de `ds.air`. Você vê atividade de computação em seu
# painel de controle?
#
# ## Estruturas de dados Xarray são coleções dask de primeira classe.
#
# Isso significa que você pode fazer coisas como `dask.compute(xarray_object)`,
# `dask.visualize(xarray_object)`, `dask.persist(xarray_object)`. Isso funciona para
# DataArrays e Datasets.
#
# #### Exercício
#
# Visualize o gráfico de tarefas para `média`.
#
# +
# Seu código aqui
# -
# Visualize o gráfico de tarefas para `mean.data`. É igual ao gráfico ao acima?
#
# +
# Seu código aqui
# -
# ## Paralelização automática com apply_ufunc e map_blocks
#
# Quase todas as operações integradas do xarray funcionam em arrays Dask.
#
# Às vezes, a análise exige funções que não estão na API do xarray (por exemplo, scipy).
# Existem três maneiras de aplicar essas funções em paralelo em cada bloco de seu
# objeto xarray:
#
# 1. Extraia arrays Dask de objetos xarray (`.data`) e use Dask diretamente, por exemplo,
# (`Apply_gufunc`, `map_blocks`,` map_overlap` ou `blockwise`);
#
# 2. Use `xarray.apply_ufunc()` para aplicar funções que consomem e retornam matrizes NumPy;
#
# 3. Use `xarray.map_blocks()`, `Dataset.map_blocks()` ou `DataArray.map_blocks()` para aplicar funções que consomem e retornam objetos xarray.
#
# O método que você usa depende basicamente do tipo de objetos de entrada esperados pela função que você está envolvendo e o nível de desempenho ou conveniência que você deseja.
# ### `map_blocks`
#
# `map_blocks` é inspirado na função `dask.array` de mesmo nome e permite você mapear uma função em blocos do objeto xarray (incluindo Datasets).
#
# No tempo de _computação_, sua função receberá um objeto Xarray com valores concretos
# (calculados) junto com os metadados apropriados. Esta função deve retornar um objeto xarray.
#
# Aqui está um exemplo:
# +
def time_mean(obj):
# use a conveniente API do xarray aqui
# você pode converter para um dataframe do pandas e usar a API extensa do pandas
# ou use .plot() e plt.savefig para salvar visualizações em disco em paralelo.
return obj.mean("lat")
ds.map_blocks(time_mean) # isso é lazy!
# -
# isto irá calcular os valores e devolverá True se o cálculo funcionar como esperado
ds.map_blocks(time_mean).identical(ds.mean("lat"))
# #### Exercise
#
# Tente aplicar a seguinte função com `map_blocks`. Especifique `scale` como um
# argumento e `offset` como um kwarg.
#
# A docstring pode ajudar:
# https://xarray.pydata.org/en/stable/generated/xarray.map_blocks.html
#
# ```python
# def time_mean_scaled(obj, scale, offset):
# return obj.mean("lat") * scale + offset
# ```
#
# #### Funções mais avançadas
#
# `map_blocks` precisa saber _exatamente_ como o objeto retornado se parece.
# A função faz isso passando um objeto xarray de formato "0" para a função e examinando o
# resultado. Essa abordagem pode não funcionar em todos os casos. Para esses casos de uso avançados, `map_blocks` permite um kwarg` template`.
# Veja
# https://xarray.pydata.org/en/latest/dask.html#map-blocks para mais detalhes.
#
# ### apply_ufunc
#
# `Apply_ufunc` é um wrapper mais avançado que é projetado para aplicar funções
# que esperam e retornam NumPy (ou outras matrizes). Por exemplo, isso incluiria
# toda a API do SciPy. Uma vez que `apply_ufunc` opera em NumPy ou objetos Dask, ele ignora a sobrecarga de usar objetos Xarray, tornando-o uma boa escolha para funções de desempenho crítico.
#
# `Apply_ufunc` pode ser um pouco complicado de acertar, pois opera em um nível mais baixo
# nível do que `map_blocks`. Por outro lado, o Xarray usa `apply_ufunc` internamente
# para implementar muito de sua API, o que significa que é bastante poderoso!
#
# ### Um exemplo simples
#
# Funções simples que atuam independentemente em cada valor devem funcionar sem qualquer
# argumentos adicionais. No entanto, o manuseio do `dask` precisa ser explicitamente habilitado
#
# + tags=["raises-exception"]
# %%expect_exception
squared_error = lambda x, y: (x - y) ** 2
xr.apply_ufunc(squared_error, ds.air, 1)
# -
# Existem duas opções para o kwarg `dask`:
#
# 1. `dask = "allowed"` (permitido): Arrays Dask são passados para a função do usuário. Essa é uma boa escolha se sua função pode lidar com arrays dask e não chamará compute explicitamente.
# 2. `dask = "paralelizado"` (paralelizado). Isso aplica a função do usuário sobre os blocos do dask array usando `dask.array.blockwise`. Isso é útil quando sua função não pode lidar com matrizes dask nativamente (por exemplo, API scipy).
#
# Uma vez que `squared_error` pode lidar com arrays dask sem computá-los, especificamos
# `dask = "permitido"`.
sqer = xr.apply_ufunc(
squared_error,
ds.air,
1,
dask="allowed",
)
sqer # DataArray apoiado por dask! com bons metadados!
# ### Um exemplo mais complicado com uma função compatível com dask
#
# Para usar operações mais complexas que consideram alguns valores de matriz coletivamente,
# é importante entender a ideia de **dimensões centrais** do NumPy ao generalizar ufuncs. As dimensões principais são definidas como dimensões que não devem ser
# propagadas. Normalmente, eles correspondem às dimensões fundamentais sobre
# as quais uma operação é definida, por exemplo, o eixo somado em `np.sum`. Uma boa pista sobre a necessidade de dimensões centrais é a presença de um argumento do `axis` na
# função NumPy correspondente.
#
# Com `apply_ufunc`, as dimensões principais são reconhecidas pelo nome e, em seguida, movidas para a última dimensão de quaisquer argumentos de entrada antes de aplicar a função fornecida.
# Isso significa que para funções que aceitam um argumento de `axis`, você geralmente precisa para definir `axis = -1`.
#
# Vamos usar `dask.array.mean` como um exemplo de uma função que pode lidar com o dask
# arrays e usa um kwarg `axis`:
#
# +
def time_mean(da):
return xr.apply_ufunc(
dask.array.mean,
da,
input_core_dims=[["time"]],
dask="allowed",
kwargs={"axis": -1}, # core dimensions are moved to the end
)
time_mean(ds.air)
# -
ds.air.mean("time").identical(time_mean(ds.air))
# ### Paralelizando funções que desconhecem dask
#
# Um recurso muito útil do `apply_ufunc` é a capacidade de aplicar funções arbitrárias
# em paralelo a cada bloco. Esta habilidade pode ser ativada usando `dask = "parallelized"`. Novamente, o Xarray precisa de muitos metadados extras, dependendo da função, argumentos extras como `output_dtypes` e `output_sizes` podem ser necessários.
#
# Usaremos `scipy.integrate.trapz` como um exemplo de uma função que não consegue
# lidar com matrizes dask e requer uma dimensão central:
#
# +
import scipy as sp
import scipy.integrate
sp.integrate.trapz(ds.air.data) # NÃO retorna uma matriz dask
# -
# #### Exercício
#
# Use `apply_ufunc` para aplicar `sp.integrate.trapz` ao longo do eixo do `tempo` para que
# você obtenha o retorno de um array dask. Você precisará especificar `dask = "parallelized"` e `output_dtypes` (uma lista de `dtypes` por variável retornada).
# +
# Seu código aqui
# -
# ## Veja mais detalhes
#
# 1. https://xarray.pydata.org/en/stable/examples/apply_ufunc_vectorize_1d.html#
# 2. https://docs.dask.org/en/latest/array-best-practices.html
#
| python-brasil-2021/06_xarray_e_dask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Lf7huAiYp-An"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" id="YHz2D-oIqBWa"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="x44FFES-r6y0"
# # Federated Learning for Text Generation
# + [markdown] id="iPFgLeZIsZ3Q"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/federated/tutorials/federated_learning_for_text_generation"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/federated/blob/master/docs/tutorials/federated_learning_for_text_generation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/federated/blob/master/docs/tutorials/federated_learning_for_text_generation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] id="KbNz2tuvsAFB"
# **NOTE**: This colab has been verified to work with the [latest released version](https://github.com/tensorflow/federated#compatibility) of the `tensorflow_federated` pip package, but the Tensorflow Federated project is still in pre-release development and may not work on `master`.
#
# This tutorial builds on the concepts in the [Federated Learning for Image Classification](federated_learning_for_image_classification.ipynb) tutorial, and demonstrates several other useful approaches for federated learning.
#
# In particular, we load a previously trained Keras model, and refine it using federated training on a (simulated) decentralized dataset. This is practically important for several reasons . The ability to use serialized models makes it easy to mix federated learning with other ML approaches. Further, this allows use of an increasing range of pre-trained models --- for example, training language models from scratch is rarely necessary, as numerous pre-trained models are now widely available (see, e.g., [TF Hub](https://www.tensorflow.org/hub)). Instead, it makes more sense to start from a pre-trained model, and refine it using Federated Learning, adapting to the particular characteristics of the decentralized data for a particular application.
#
# For this tutorial, we start with a RNN that generates ASCII characters, and refine it via federated learning. We also show how the final weights can be fed back to the original Keras model, allowing easy evaluation and text generation using standard tools.
# + id="9LcC1AwjoqfR"
#@test {"skip": true}
# !pip install --quiet --upgrade tensorflow_federated_nightly
# !pip install --quiet --upgrade nest_asyncio
import nest_asyncio
nest_asyncio.apply()
# + id="ZjDQysatrc2S"
import collections
import functools
import os
import time
import numpy as np
import tensorflow as tf
import tensorflow_federated as tff
np.random.seed(0)
# Test the TFF is working:
tff.federated_computation(lambda: 'Hello, World!')()
# + [markdown] id="lyICXwVAxvW9"
# ## Load a pre-trained model
#
# We load a model that was pre-trained following the TensorFlow tutorial
# [Text generation using a RNN with eager execution](https://www.tensorflow.org/tutorials/sequences/text_generation). However,
# rather than training on [The Complete Works of Shakespeare](http://www.gutenberg.org/files/100/100-0.txt), we pre-trained the model on the text from the <NAME>'
# [A Tale of Two Cities](http://www.ibiblio.org/pub/docs/books/gutenberg/9/98/98.txt)
# and
# [A Christmas Carol](http://www.ibiblio.org/pub/docs/books/gutenberg/4/46/46.txt).
#
# Other than expanding the vocabulary, we didn't modify the original tutorial, so this initial model isn't state-of-the-art, but it produces reasonable predictions and is sufficient for our tutorial purposes. The final model was saved with `tf.keras.models.save_model(include_optimizer=False)`.
#
# We will use federated learning to fine-tune this model for Shakespeare in this tutorial, using a federated version of the data provided by TFF.
#
# + [markdown] id="XgF8e2Ksyq1F"
# ### Generate the vocab lookup tables
# + id="IlCgQBRVymwR"
# A fixed vocabularly of ASCII chars that occur in the works of Shakespeare and Dickens:
vocab = list('dhlptx@DHLPTX $(,048cgkoswCGKOSW[_#\'/37;?bfjnrvzBFJNRVZ"&*.26:\naeimquyAEIMQUY]!%)-159\r')
# Creating a mapping from unique characters to indices
char2idx = {u:i for i, u in enumerate(vocab)}
idx2char = np.array(vocab)
# + [markdown] id="2EH6MFRdzAwd"
# ### Load the pre-trained model and generate some text
# + id="iIK674SrtCTm"
def load_model(batch_size):
urls = {
1: 'https://storage.googleapis.com/tff-models-public/dickens_rnn.batch1.kerasmodel',
8: 'https://storage.googleapis.com/tff-models-public/dickens_rnn.batch8.kerasmodel'}
assert batch_size in urls, 'batch_size must be in ' + str(urls.keys())
url = urls[batch_size]
local_file = tf.keras.utils.get_file(os.path.basename(url), origin=url)
return tf.keras.models.load_model(local_file, compile=False)
# + id="WvuwZBX5Ogfd"
def generate_text(model, start_string):
# From https://www.tensorflow.org/tutorials/sequences/text_generation
num_generate = 200
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
text_generated = []
temperature = 1.0
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
predictions = tf.squeeze(predictions, 0)
predictions = predictions / temperature
predicted_id = tf.random.categorical(
predictions, num_samples=1)[-1, 0].numpy()
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2char[predicted_id])
return (start_string + ''.join(text_generated))
# + id="MGAdStJ5wDPV"
# Text generation requires a batch_size=1 model.
keras_model_batch1 = load_model(batch_size=1)
print(generate_text(keras_model_batch1, 'What of TensorFlow Federated, you ask? '))
# + [markdown] id="kKMUn-TlgxuP"
# ## Load and Preprocess the Federated Shakespeare Data
#
# The `tff.simulation.datasets` package provides a variety of datasets that are split into "clients", where each client corresponds to a dataset on a particular device that might participate in federated learning.
#
# These datasets provide realistic non-IID data distributions that replicate in simulation the challenges of training on real decentralized data. Some of the pre-processing of this data was done using tools from the [Leaf project](https://arxiv.org/abs/1812.01097) ([github](https://github.com/TalwalkarLab/leaf)).
# + id="di3nStTDg0qc"
train_data, test_data = tff.simulation.datasets.shakespeare.load_data()
# + [markdown] id="_iiY65Vv4QNK"
# The datasets provided by `shakespeare.load_data()` consist of a sequence of
# string `Tensors`, one for each line spoken by a particular character in a
# Shakespeare play. The client keys consist of the name of the play joined with
# the name of the character, so for example `MUCH_ADO_ABOUT_NOTHING_OTHELLO` corresponds to the lines for the character Othello in the play *Much Ado About Nothing*. Note that in a real federated learning scenario
# clients are never identified or tracked by ids, but for simulation it is useful
# to work with keyed datasets.
#
# Here, for example, we can look at some data from King Lear:
# + id="FEKiy1ntmmnk"
# Here the play is "The Tragedy of King Lear" and the character is "King".
raw_example_dataset = train_data.create_tf_dataset_for_client(
'THE_TRAGEDY_OF_KING_LEAR_KING')
# To allow for future extensions, each entry x
# is an OrderedDict with a single key 'snippets' which contains the text.
for x in raw_example_dataset.take(2):
print(x['snippets'])
# + [markdown] id="kUnbI5Hp4sXg"
# We now use `tf.data.Dataset` transformations to prepare this data for training the char RNN loaded above.
#
# + id="9kDkmGe-7No7"
# Input pre-processing parameters
SEQ_LENGTH = 100
BATCH_SIZE = 8
BUFFER_SIZE = 100 # For dataset shuffling
# + id="W95Of6Bwsrfc"
# Construct a lookup table to map string chars to indexes,
# using the vocab loaded above:
table = tf.lookup.StaticHashTable(
tf.lookup.KeyValueTensorInitializer(
keys=vocab, values=tf.constant(list(range(len(vocab))),
dtype=tf.int64)),
default_value=0)
def to_ids(x):
s = tf.reshape(x['snippets'], shape=[1])
chars = tf.strings.bytes_split(s).values
ids = table.lookup(chars)
return ids
def split_input_target(chunk):
input_text = tf.map_fn(lambda x: x[:-1], chunk)
target_text = tf.map_fn(lambda x: x[1:], chunk)
return (input_text, target_text)
def preprocess(dataset):
return (
# Map ASCII chars to int64 indexes using the vocab
dataset.map(to_ids)
# Split into individual chars
.unbatch()
# Form example sequences of SEQ_LENGTH +1
.batch(SEQ_LENGTH + 1, drop_remainder=True)
# Shuffle and form minibatches
.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
# And finally split into (input, target) tuples,
# each of length SEQ_LENGTH.
.map(split_input_target))
# + [markdown] id="Jw98HnKmEhuh"
# Note that in the formation of the original sequences and in the formation of
# batches above, we use `drop_remainder=True` for simplicity. This means that any
# characters (clients) that don't have at least `(SEQ_LENGTH + 1) * BATCH_SIZE`
# chars of text will have empty datasets. A typical approach to address this would
# be to pad the batches with a special token, and then mask the loss to not take
# the padding tokens into account.
#
# This would complicate the example somewhat, so for this tutorial we only use full batches, as in the
# [standard tutorial](https://www.tensorflow.org/tutorials/sequences/text_generation).
# However, in the federated setting this issue is more significant, because many
# users might have small datasets.
#
# Now we can preprocess our `raw_example_dataset`, and check the types:
# + id="7rTal7bksWwc"
example_dataset = preprocess(raw_example_dataset)
print(example_dataset.element_spec)
# + [markdown] id="ePT8Oawm8SRP"
# ## Compile the model and test on the preprocessed data
# + [markdown] id="vEgDsz-48cAq"
# We loaded an uncompiled keras model, but in order to run `keras_model.evaluate`, we need to compile it with a loss and metrics. We will also compile in an optimizer, which will be used as the on-device optimizer in Federated Learning.
# + [markdown] id="RsuVZ5KMWnn8"
# The original tutorial didn't have char-level accuracy (the fraction
# of predictions where the highest probability was put on the correct
# next char). This is a useful metric, so we add it.
# However, we need to define a new metric class for this because
# our predictions have rank 3 (a vector of logits for each of the
# `BATCH_SIZE * SEQ_LENGTH` predictions), and `SparseCategoricalAccuracy`
# expects only rank 2 predictions.
# + id="gOUiDBvmWlM9"
class FlattenedCategoricalAccuracy(tf.keras.metrics.SparseCategoricalAccuracy):
def __init__(self, name='accuracy', dtype=tf.float32):
super().__init__(name, dtype=dtype)
def update_state(self, y_true, y_pred, sample_weight=None):
y_true = tf.reshape(y_true, [-1, 1])
y_pred = tf.reshape(y_pred, [-1, len(vocab), 1])
return super().update_state(y_true, y_pred, sample_weight)
# + [markdown] id="U2X9eFgt94PM"
# Now we can compile a model, and evaluate it on our `example_dataset`.
# + id="c3Xd-52-9zGa"
BATCH_SIZE = 8 # The training and eval batch size for the rest of this tutorial.
keras_model = load_model(batch_size=BATCH_SIZE)
keras_model.compile(
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[FlattenedCategoricalAccuracy()])
# Confirm that loss is much lower on Shakespeare than on random data
loss, accuracy = keras_model.evaluate(example_dataset.take(5), verbose=0)
print(
'Evaluating on an example Shakespeare character: {a:3f}'.format(a=accuracy))
# As a sanity check, we can construct some completely random data, where we expect
# the accuracy to be essentially random:
random_guessed_accuracy = 1.0 / len(vocab)
print('Expected accuracy for random guessing: {a:.3f}'.format(
a=random_guessed_accuracy))
random_indexes = np.random.randint(
low=0, high=len(vocab), size=1 * BATCH_SIZE * (SEQ_LENGTH + 1))
data = collections.OrderedDict(
snippets=tf.constant(
''.join(np.array(vocab)[random_indexes]), shape=[1, 1]))
random_dataset = preprocess(tf.data.Dataset.from_tensor_slices(data))
loss, accuracy = keras_model.evaluate(random_dataset, steps=10, verbose=0)
print('Evaluating on completely random data: {a:.3f}'.format(a=accuracy))
# + [markdown] id="lH0WzL5L8Lm4"
# ## Fine-tune the model with Federated Learning
# + [markdown] id="NCao4M3L_tsA"
# TFF serializes all TensorFlow computations so they can potentially be run in a
# non-Python environment (even though at the moment, only a simulation runtime implemented in Python is available). Even though we are running in eager mode, (TF 2.0), currently TFF serializes TensorFlow computations by constructing the
# necessary ops inside the context of a "`with tf.Graph.as_default()`" statement.
# Thus, we need to provide a function that TFF can use to introduce our model into
# a graph it controls. We do this as follows:
# + id="5KadIvFp7m6y"
# Clone the keras_model inside `create_tff_model()`, which TFF will
# call to produce a new copy of the model inside the graph that it will
# serialize. Note: we want to construct all the necessary objects we'll need
# _inside_ this method.
def create_tff_model():
# TFF uses an `input_spec` so it knows the types and shapes
# that your model expects.
input_spec = example_dataset.element_spec
keras_model_clone = tf.keras.models.clone_model(keras_model)
return tff.learning.from_keras_model(
keras_model_clone,
input_spec=input_spec,
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[FlattenedCategoricalAccuracy()])
# + [markdown] id="ZJF_yhJxAi2l"
# Now we are ready to construct a Federated Averaging iterative process, which we will use to improve the model (for details on the Federated Averaging algorithm, see the paper [Communication-Efficient Learning of Deep Networks from Decentralized Data](https://arxiv.org/abs/1602.05629)).
#
# We use a compiled Keras model to perform standard (non-federated) evaluation after each round of federated training. This is useful for research purposes when doing simulated federated learning and there is a standard test dataset.
#
# In a realistic production setting this same technique might be used to take models trained with federated learning and evaluate them on a centralized benchmark dataset for testing or quality assurance purposes.
# + id="my3PW3qhAMDA"
# This command builds all the TensorFlow graphs and serializes them:
fed_avg = tff.learning.build_federated_averaging_process(
model_fn=create_tff_model,
client_optimizer_fn=lambda: tf.keras.optimizers.SGD(lr=0.5))
# + [markdown] id="qVOkzs9C9kmv"
# Here is the simplest possible loop, where we run federated averaging for one round on a single client on a single batch:
# + id="lrjUrkjq9jYk"
state = fed_avg.initialize()
state, metrics = fed_avg.next(state, [example_dataset.take(5)])
train_metrics = metrics['train']
print('loss={l:.3f}, accuracy={a:.3f}'.format(
l=train_metrics['loss'], a=train_metrics['accuracy']))
# + [markdown] id="o2CjvVg0FZpS"
# Now let's write a slightly more interesting training and evaluation loop.
#
# So that this simulation still runs relatively quickly, we train on the same three clients each round, only considering two minibatches for each.
#
# + id="wE386-rbMCve"
def data(client, source=train_data):
return preprocess(source.create_tf_dataset_for_client(client)).take(5)
clients = [
'ALL_S_WELL_THAT_ENDS_WELL_CELIA', 'MUCH_ADO_ABOUT_NOTHING_OTHELLO',
]
train_datasets = [data(client) for client in clients]
# We concatenate the test datasets for evaluation with Keras by creating a
# Dataset of Datasets, and then identity flat mapping across all the examples.
test_dataset = tf.data.Dataset.from_tensor_slices(
[data(client, test_data) for client in clients]).flat_map(lambda x: x)
# + [markdown] id="cU3FuY00MOoX"
# The initial state of the model produced by `fed_avg.initialize()` is based
# on the random initializers for the Keras model, not the weights that were loaded,
# since `clone_model()` does not clone the weights. To start training
# from a pre-trained model, we set the model weights in the server state
# directly from the loaded model.
# + id="vm_-PU8OFXpY"
NUM_ROUNDS = 5
# The state of the FL server, containing the model and optimization state.
state = fed_avg.initialize()
# Load our pre-trained Keras model weights into the global model state.
state = tff.learning.state_with_new_model_weights(
state,
trainable_weights=[v.numpy() for v in keras_model.trainable_weights],
non_trainable_weights=[
v.numpy() for v in keras_model.non_trainable_weights
])
def keras_evaluate(state, round_num):
# Take our global model weights and push them back into a Keras model to
# use its standard `.evaluate()` method.
keras_model = load_model(batch_size=BATCH_SIZE)
keras_model.compile(
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[FlattenedCategoricalAccuracy()])
state.model.assign_weights_to(keras_model)
loss, accuracy = keras_model.evaluate(example_dataset, steps=2, verbose=0)
print('\tEval: loss={l:.3f}, accuracy={a:.3f}'.format(l=loss, a=accuracy))
for round_num in range(NUM_ROUNDS):
print('Round {r}'.format(r=round_num))
keras_evaluate(state, round_num)
state, metrics = fed_avg.next(state, train_datasets)
train_metrics = metrics['train']
print('\tTrain: loss={l:.3f}, accuracy={a:.3f}'.format(
l=train_metrics['loss'], a=train_metrics['accuracy']))
print('Final evaluation')
keras_evaluate(state, NUM_ROUNDS + 1)
# + [markdown] id="SoshvcHhXVa6"
# With the default changes, we haven't done enough training to make a big difference, but if you train longer on more Shakespeare data, you should see a difference in the style of the text generated with the updated model:
# + id="NTUig7QmXavy"
# Set our newly trained weights back in the originally created model.
keras_model_batch1.set_weights([v.numpy() for v in keras_model.weights])
# Text generation requires batch_size=1
print(generate_text(keras_model_batch1, 'What of TensorFlow Federated, you ask? '))
# + [markdown] id="4DA1Fkf5mN0s"
# ## Suggested extensions
#
# This tutorial is just the first step! Here are some ideas for how you might try extending this notebook:
# * Write a more realistic training loop where you sample clients to train on randomly.
# * Use "`.repeat(NUM_EPOCHS)`" on the client datasets to try multiple epochs of local training (e.g., as in [McMahan et. al.](https://arxiv.org/abs/1602.05629)). See also [Federated Learning for Image Classification](federated_learning_for_image_classification.ipynb) which does this.
# * Change the `compile()` command to experiment with using different optimization algorithms on the client.
# * Try the `server_optimizer` argument to `build_federated_averaging_process` to try different algorithms for applying the model updates on the server.
# * Try the `client_weight_fn` argument to to `build_federated_averaging_process` to try different weightings of the clients. The default weights client updates by the number of examples on the client, but you can do e.g. `client_weight_fn=lambda _: tf.constant(1.0)`.
| site/en-snapshot/federated/tutorials/federated_learning_for_text_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading DFlowFM Map Output
#
# Demonstrate basic loading and plotting of map output from DFlowFM. This
# example uses sample output which is not part of the stompy
# example data.
import matplotlib.pyplot as plt
import xarray as xr
from stompy.model.delft import dfm_grid
from stompy.grid import unstructured_grid
# %matplotlib notebook
# # Open the NetCDF Map file
# +
map_nc='/home/emma/test_run/r14_map.nc'
# This pulls the grid topology/geometry out of the netcdf.
# The netcdf files aren't quite ugrid compliant, so we have
# to use DFMGrid, specific to Flow FM's version of netcdf
grid=dfm_grid.DFMGrid(map_nc)
# The xarray dataset is used to access the variables which
# will be plotted on the grid
ds=xr.open_dataset(map_nc)
# +
fig,axs=plt.subplots(1,3,sharex=True,sharey=True,figsize=(12,6))
# To make it a bit faster, supply a clipping box:
clip=(543535., 553384.,4176205., 4202129.)
grid.plot_edges(ax=axs[0],clip=clip)
grid.plot_cells(ax=axs[1],clip=clip)
grid.plot_nodes(ax=axs[2],clip=clip)
# +
# Select the second timestep, top layer of output.
# .values returns a numpy array, dropping the XArray wrapping
surf_salt=ds.sa1.isel(time=1,laydim=0).values
fig,ax=plt.subplots(figsize=(10,8))
# plot_* commands typically return a collection
coll=grid.plot_cells(values=surf_salt)
# Doctor up the presentation a bit:
coll.set_edgecolor('face')
plt.colorbar(coll,label='Salinity (ppt)')
| examples/plot_dflow_map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ---
# description: Continual Learning Algorithms Prototyping Made Easy
# ---
# # Training
#
# Welcome to the "_Training_" tutorial of the "_From Zero to Hero_" series. In this part we will present the functionalities offered by the `training` module.
# + pycharm={"name": "#%%\n"}
# !pip install git+https://github.com/ContinualAI/avalanche.git
# -
# ## 💪 The Training Module
#
# The `training` module in _Avalanche_ is designed with modularity in mind. Its main goals are to:
#
# 1. provide a set of popular **continual learning baselines** that can be easily used to run experimental comparisons;
# 2. provide simple abstractions to **create and run your own strategy** as efficiently and easily as possible starting from a couple of basic building blocks we already prepared for you.
#
# At the moment, the `training` module includes two main components:
#
# * **Strategies**: these are popular baselines already implemented for you which you can use for comparisons or as base classes to define a custom strategy.
# * **Plugins**: these are classes that allow to add some specific behaviour to your own strategy. The plugin system allows to define reusable components which can be easily combined together (e.g. a replay strategy, a regularization strategy). They are also used to automatically manage logging and evaluation.
#
# Keep in mind that many Avalanche components are independent from Avalanche strategies. If you already have your own strategy which does not use Avalanche, you can use Avalanche's benchmarks, models, data loaders, and metrics without ever looking at Avalanche's strategies.
#
# ## 📈 How to Use Strategies & Plugins
#
# If you want to compare your strategy with other classic continual learning algorithm or baselines, in _Avalanche_ you can instantiate a strategy with a couple lines of code.
#
# ### Strategy Instantiation
# Most strategies require only 3 mandatory arguments:
# - **model**: this must be a `torch.nn.Module`.
# - **optimizer**: `torch.optim.Optimizer` already initialized on your `model`.
# - **loss**: a loss function such as those in `torch.nn.functional`.
#
# Additional arguments are optional and allow you to customize training (batch size, epochs, ...) or strategy-specific parameters (buffer size, regularization strength, ...).
# + pycharm={"name": "#%%\n"}
from torch.optim import SGD
from torch.nn import CrossEntropyLoss
from avalanche.models import SimpleMLP
from avalanche.training.strategies import Naive, CWRStar, Replay, GDumb, Cumulative, LwF, GEM, AGEM, EWC
model = SimpleMLP(num_classes=10)
optimizer = SGD(model.parameters(), lr=0.001, momentum=0.9)
criterion = CrossEntropyLoss()
cl_strategy = Naive(
model, optimizer, criterion,
train_mb_size=100, train_epochs=4, eval_mb_size=100
)
# -
# ### Training & Evaluation
#
# Each strategy object offers two main methods: `train` and `eval`. Both of them, accept either a _single experience_(`Experience`) or a _list of them_, for maximum flexibility.
#
# We can train the model continually by iterating over the `train_stream` provided by the scenario.
# + pycharm={"name": "#%%\n"}
from avalanche.benchmarks.classic import SplitMNIST
# scenario
benchmark = SplitMNIST(n_experiences=5, seed=1)
# TRAINING LOOP
print('Starting experiment...')
results = []
for experience in benchmark.train_stream:
print("Start of experience: ", experience.current_experience)
print("Current Classes: ", experience.classes_in_this_experience)
cl_strategy.train(experience)
print('Training completed')
print('Computing accuracy on the whole test set')
results.append(cl_strategy.eval(benchmark.test_stream))
# -
# ### Adding Plugins
#
# Most continual learning strategies follow roughly the same training/evaluation loops, i.e. a simple naive strategy (a.k.a. finetuning) augmented with additional behavior to counteract catastrophic forgetting. The plugin systems in Avalanche is designed to easily augment continual learning strategies with custom behavior, without having to rewrite the training loop from scratch. Avalanche strategies accept an optional list of `plugins` that will be executed during the training/evaluation loops.
#
# For example, early stopping is implemented as a plugin:
# + pycharm={"name": "#%%\n"}
from avalanche.training.plugins import EarlyStoppingPlugin
strategy = Naive(
model, optimizer, criterion,
plugins=[EarlyStoppingPlugin(patience=10, val_stream_name='train')])
# + [markdown] pycharm={"name": "#%% md\n"}
# In Avalanche, most continual learning strategies are implemented using plugins, which makes it easy to combine them together. For example, it is extremely easy to create a hybrid strategy that combines replay and EWC together by passing the appropriate `plugins` list to the `BaseStrategy`:
# + pycharm={"name": "#%%\n"}
from avalanche.training.strategies import BaseStrategy
from avalanche.training.plugins import ReplayPlugin, EWCPlugin
replay = ReplayPlugin(mem_size=100)
ewc = EWCPlugin(ewc_lambda=0.001)
strategy = BaseStrategy(
model, optimizer, criterion,
plugins=[replay, ewc])
# + [markdown] pycharm={"name": "#%% md\n"}
# Beware that most strategy plugins modify the internal state. As a result, not all the strategy plugins can be combined together. For example, it does not make sense to use multiple replay plugins since they will try to modify the same strategy variables (mini-batches, dataloaders), and therefore they will be in conflict.
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 📝 A Look Inside Avalanche Strategies
#
# If you arrived at this point you already know how to use Avalanche strategies and are ready to use it. However, before making your own strategies you need to understand a little bit the internal implementation of the training and evaluation loops.
#
# In _Avalanche_ you can customize a strategy in 2 ways:
#
# 1. **Plugins**: Most strategies can be implemented as additional code that runs on top of the basic training and evaluation loops (`Naive` strategy, or `BaseStrategy`). Therefore, the easiest way to define a custom strategy such as a regularization or replay strategy, is to define it as a custom plugin. The advantage of plugins is that they can be combined together, as long as they are compatible, i.e. they do not modify the same part of the state. The disadvantage is that in order to do so you need to understand the `BaseStrategy` loop, which can be a bit complex at first.
# 2. **Subclassing**: In _Avalanche_, continual learning strategies inherit from the `BaseStrategy`, which provides generic training and evaluation loops. Most `BaseStrategy` methods can be safely overridden (with some caveats that we will see later).
#
# Keep in mind that if you already have a working continual learning strategy that does not use _Avalanche_, you can use most Avalanche components such as `benchmarks`, `evaluation`, and `models` without using _Avalanche_'s strategies!
#
# ### Training and Evaluation Loops
#
# As we already mentioned, _Avalanche_ strategies inherit from `BaseStrategy`. This strategy provides:
#
# 1. Basic _Training_ and _Evaluation_ loops which define a naive (finetuning) strategy.
# 2. _Callback_ points, which are used to call the plugins at a specific moments during the loop's execution.
# 3. A set of variables representing the state of the loops (current model, data, mini-batch, predictions, ...) which allows plugins and child classes to easily manipulate the state of the training loop.
#
# The training loop has the following structure:
# ```text
# train
# before_training
#
# before_train_dataset_adaptation
# train_dataset_adaptation
# after_train_dataset_adaptation
# make_train_dataloader
# model_adaptation
# make_optimizer
# before_training_exp # for each exp
# before_training_epoch # for each epoch
# before_training_iteration # for each iteration
# before_forward
# after_forward
# before_backward
# after_backward
# after_training_iteration
# before_update
# after_update
# after_training_epoch
# after_training_exp
# after_training
# ```
#
# The evaluation loop is similar:
# ```text
# eval
# before_eval
# before_eval_dataset_adaptation
# eval_dataset_adaptation
# after_eval_dataset_adaptation
# make_eval_dataloader
# model_adaptation
# before_eval_exp # for each exp
# eval_epoch # we have a single epoch in evaluation mode
# before_eval_iteration # for each iteration
# before_eval_forward
# after_eval_forward
# after_eval_iteration
# after_eval_exp
# after_eval
# ```
#
# Methods starting with `before/after` are the methods responsible for calling the plugins.
# Notice that before the start of each experience during training we have several phases:
# - *dataset adaptation*: This is the phase where the training data can be modified by the strategy, for example by adding other samples from a separate buffer.
# - *dataloader initialization*: Initialize the data loader. Many strategies (e.g. replay) use custom dataloaders to balance the data.
# - *model adaptation*: Here, the dynamic models (see the `models` tutorial) are updated by calling their `adaptation` method.
# - *optimizer initialization*: After the model has been updated, the optimizer should also be updated to ensure that the new parameters are optimized.
#
# ### Strategy State
# The strategy state is accessible via several attributes. Most of these can be modified by plugins and subclasses:
# - `self.clock`: keeps track of several event counters.
# - `self.experience`: the current experience.
# - `self.adapted_dataset`: the data modified by the dataset adaptation phase.
# - `self.dataloader`: the current dataloader.
# - `self.mbatch`: the current mini-batch. For classification problems, mini-batches have the form `<x, y, t>`, where `x` is the input, `y` is the label, and `t` is the target.
# - `self.mb_output`: the current model's output.
# - `self.loss`: the current loss.
# - `self.is_training`: `True` if the strategy is in training mode.
#
# ## How to Write a Plugin
# Plugins provide a simple solution to define a new strategy by augmenting the behavior of another strategy (typically a naive strategy). This approach reduces the overhead and code duplication, **improving code readability and prototyping speed**.
#
# Creating a plugin is straightforward. You create a class which inherits from `StrategyPlugin` and implements the callbacks that you need. The exact callback to use depend on your strategy. You can use the loop shown above to understand what callbacks you need to use. For example, we show below a simple replay plugin that uses `after_training_exp` to update the buffer after each training experience, and the `before_training_exp` to customize the dataloader. Notice that `before_training_exp` is executed after `make_train_dataloader`, which means that the `BaseStrategy` already updated the dataloader. If we used another callback, such as `before_train_dataset_adaptation`, our dataloader would have been overwritten by the `BaseStrategy`. Plugin methods always receive the `strategy` as an argument, so they can access and modify the strategy's state.
# + pycharm={"name": "#%%\n"}
from avalanche.benchmarks.utils.data_loader import ReplayDataLoader
from avalanche.training.plugins import StrategyPlugin
from avalanche.training.storage_policy import ReservoirSamplingBuffer
class ReplayP(StrategyPlugin):
def __init__(self, mem_size):
""" A simple replay plugin with reservoir sampling. """
super().__init__()
self.buffer = ReservoirSamplingBuffer(max_size=mem_size)
def before_training_exp(self, strategy: "BaseStrategy",
num_workers: int = 0, shuffle: bool = True,
**kwargs):
""" Use a custom dataloader to combine samples from the current data and memory buffer. """
if len(self.buffer.buffer) == 0:
# first experience. We don't use the buffer, no need to change
# the dataloader.
return
strategy.dataloader = ReplayDataLoader(
strategy.adapted_dataset,
self.buffer.buffer,
oversample_small_tasks=True,
num_workers=num_workers,
batch_size=strategy.train_mb_size,
shuffle=shuffle)
def after_training_exp(self, strategy: "BaseStrategy", **kwargs):
""" Update the buffer. """
self.buffer.update(strategy, **kwargs)
benchmark = SplitMNIST(n_experiences=5, seed=1)
model = SimpleMLP(num_classes=10)
optimizer = SGD(model.parameters(), lr=0.01, momentum=0.9)
criterion = CrossEntropyLoss()
strategy = Naive(model=model, optimizer=optimizer, criterion=criterion, train_mb_size=128,
plugins=[ReplayP(mem_size=2000)])
strategy.train(benchmark.train_stream)
strategy.eval(benchmark.test_stream)
# + [markdown] pycharm={"name": "#%% md\n"}
# Check `StrategyPlugin`'s documentation for a complete list of the available callbacks.
# -
# ## How to Write a Custom Strategy
#
# You can always define a custom strategy by overriding `BaseStrategy` methods.
# However, There is an important caveat to keep in mind. If you override a method, you must remember to call all the callback's handlers (the methods starting with `before/after`) at the appropriate points. For example, `train` calls `before_training` and `after_training` before and after the training loops, respectively. The easiest way to avoid mistakes is to start from the `BaseStrategy` method that you want to override and modify it to your own needs without removing the callbacks handling.
#
# Notice that even though you don't use plugins, `BaseStrategy` implements some internal components as plugins. Also, the `EvaluationPlugin` (see `evaluation` tutorial) uses the strategy callbacks.
#
# `BaseStrategy` provides the global state of the loop in the strategy's attributes, which you can safely use when you override a method. As an example, the `Cumulative` strategy trains a model continually on the union of all the experiences encountered so far. To achieve this, the cumulative strategy overrides `adapt_train_dataset` and updates `self.adapted_dataset' by concatenating all the previous experiences with the current one.
# + pycharm={"name": "#%%\n"}
from avalanche.benchmarks.utils import AvalancheConcatDataset
from avalanche.training import BaseStrategy
class Cumulative(BaseStrategy):
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self.dataset = None # cumulative dataset
def train_dataset_adaptation(self, **kwargs):
super().train_dataset_adaptation(**kwargs)
curr_data = self.experience.dataset
if self.dataset is None:
self.dataset = curr_data
else:
self.dataset = AvalancheConcatDataset([self.dataset, curr_data])
self.adapted_dataset = self.dataset.train()
strategy = Cumulative(model=model, optimizer=optimizer, criterion=criterion, train_mb_size=128)
strategy.train(benchmark.train_stream)
# -
# Easy, isn't it? :-\)
#
# In general, we recommend to _implement a Strategy via plugins_, if possible. This approach is the easiest to use and requires a minimal knowledge of the `BaseStrategy`. It also allows other people to use your plugin and facilitates interoperability among different strategies.
#
# For example, replay strategies can be implemented as a custom strategy of the `BaseStrategy` or as plugins. However, creating a plugin is allows to use the replay strategy in conjunction with other strategies.
# This completes the "_Training_" chapter for the "_From Zero to Hero_" series. We hope you enjoyed it!
#
# ## 🤝 Run it on Google Colab
#
# You can run _this chapter_ and play with it on Google Colaboratory: [](https://colab.research.google.com/github/ContinualAI/colab/blob/master/notebooks/avalanche/3.-training.ipynb)
| notebooks/from-zero-to-hero-tutorial/04_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 420-A52-SF - Algorithmes d'apprentissage supervisé - Hiver 2020 - Spécialisation technique en Intelligence Artificielle - <NAME>, M.Sc.
# <br/>
# 
# <br/>
# **Objectif:** cette séance de travaux pratiques est consacrée à la transformation des variables explicatives catégorielles en **variables indicatrices**
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# ### 0 - Chargement des bibliothèques
# +
# Manipulation de données
import numpy as np
import pandas as pd
# Visualisation de données
import matplotlib.pyplot as plt
import seaborn as sns
# Machine Learning
from sklearn.decomposition import PCA
# -
# Configuration de la visualisation
sns.set(style="darkgrid", rc={'figure.figsize':(11.7,8.27)})
sns.set_context("notebook", font_scale=1.5, rc={"lines.linewidth": 2.5})
# ### 1 - Lecture du jeu de données *Heart*
# **Exercice 1: lire le fichier `Heart.csv`**
# Compléter le code ci-dessous ~ 1 ligne
HRT = None
# On supprime les données manquantes. Ceci sera vu plus en détail plus tard dans le cours
HRT = HRT.dropna()
# **Exercice 2: afficher les dix premières lignes de la trame de données**
# Compléter le code ci-dessous ~ 1 ligne
None
# **Exercice 3: quel est le type de données (dtype) du ndarray sous-jacent ?**
# Compléter le code ci-dessous ~ 1 ligne
None
# **Exercice 4: trouver tous les niveaux des variables catégorielles**<br/>
# Indice: vous devrez utilisez la méthode `unique()` de pandas
# Compléter le code ci-dessous ~ 3 lignes
None
# ### 2 - Création des variables indicatrices
# **Exercice 5: créer les variables indicatrices**<br/>
# Compléter le code ci-dessous ~ 7 lignes
None
# **Exercice 6: quel est maintenant le type de données (dtype) du ndarray sous-jacent ?**
# Compléter le code ci-dessous ~ 1 ligne
None
# ### 3 - Affichages des composante principales (contenu non inclus dans le cours)
X = HRT.drop(['AHD'], axis=1).values
y = HRT.AHD.values
pca = PCA(n_components=2)
pca.fit(X)
X_pca = pca.transform(X)
ax = sns.scatterplot(x=X_pca[:,0], y=X_pca[:,1], hue=HRT['AHD'], s = 60)
ax.set_xlabel('PC1')
ax.set_ylabel('PC2')
# ### Fin du TP
| nbs/07-variables-explicatives-categorielles/07-TP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Solution Notebook
# ## Problem: Create a class with an insert method to insert an int to a list. It should also support calculating the max, min, mean, and mode in O(1).
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Can we assume the inputs are valid?
# * No
# * Is there a range of inputs?
# * 0 <= item <= 100
# * Should mean return a float?
# * Yes
# * Should the other results return an int?
# * Yes
# * If there are multiple modes, what do we return?
# * Any of the modes
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * None -> TypeError
# * [] -> ValueError
# * [5, 2, 7, 9, 9, 2, 9, 4, 3, 3, 2]
# * max: 9
# * min: 2
# * mean: 55
# * mode: 9 or 2
# ## Algorithm
#
# Return the input, val
#
# Complexity:
# * Time: O(1)
# * Space: O(1)
# ## Code
# +
from __future__ import division
class Solution(object):
def __init__(self, upper_limit=100):
self.max = None
self.min = None
# Mean
self.num_items = 0
self.running_sum = 0
self.mean = None
# Mode
self.array = [0] * (upper_limit+1)
self.mode_ocurrences = 0
self.mode = None
def insert(self, val):
if val is None:
raise TypeError('val cannot be None')
if self.max is None or val > self.max:
self.max = val
if self.min is None or val < self.min:
self.min = val
# Calculate the mean
self.num_items += 1
self.running_sum += val
self.mean = self.running_sum / self.num_items
# Calculate the mode
self.array[val] += 1
if self.array[val] > self.mode_ocurrences:
self.mode_ocurrences = self.array[val]
self.mode = val
# -
# ## Unit Test
# +
# %%writefile test_math_ops.py
import unittest
class TestMathOps(unittest.TestCase):
def test_math_ops(self):
solution = Solution()
self.assertRaises(TypeError, solution.insert, None)
solution.insert(5)
solution.insert(2)
solution.insert(7)
solution.insert(9)
solution.insert(9)
solution.insert(2)
solution.insert(9)
solution.insert(4)
solution.insert(3)
solution.insert(3)
solution.insert(2)
self.assertEqual(solution.max, 9)
self.assertEqual(solution.min, 2)
self.assertEqual(solution.mean, 5)
self.assertTrue(solution.mode in (2, 9))
print('Success: test_math_ops')
def main():
test = TestMathOps()
test.test_math_ops()
if __name__ == '__main__':
main()
# -
# %run -i test_math_ops.py
| online_judges/math_ops/math_ops_solution.ipynb |
from azure.eventhub import EventData
from azure.eventhub import EventHubClient
from pyspark.sql.functions import *
from pyspark.sql.types import *
import json
from datetime import datetime
# ## Connection details
# +
kv_scope = 'key-vault-secret'
# Variables
eventhubs_namespace = dbutils.secrets.get(scope =kv_scope, key = 'traffic-eventhubs-namespace')
eventhubs_accesskey = dbutils.secrets.get(scope =kv_scope, key = 'traffic-eventhubs-accesskey')
eventhubs_accessid = dbutils.secrets.get(scope =kv_scope, key = 'traffic-eventhubs-accessid')
eventhubs_name = dbutils.secrets.get(scope =kv_scope, key = 'traffic-eventhubs-name')
# Build connection string with the above information
cameraHubConnectionString = 'Endpoint=sb://{}.servicebus.windows.net/;SharedAccessKeyName={};SharedAccessKey={};EntityPath={}'.format(
eventhubs_namespace,
eventhubs_accessid,
eventhubs_accesskey,
eventhubs_name)
print(cameraHubConnectionString)
# -
# ## Event schema definition
# 1. Everything is defined as a string, otherwise we get null values
# +
# Define schema and create incoming camera eventstream
cameraEventSchema = StructType([ StructField('TrajectId', StringType(), True),
StructField('CameraId', StringType(), True),
StructField('EventTime', StringType(), True),
StructField('Lane', StringType(), True),
StructField('Country', StringType(), True),
StructField('LicensePlate', StringType(), True),
StructField('Make', StringType(), True),
StructField('Color', StringType(), True)])
# -
# ## Configure event hub reader
# +
# Starting position
startingEventPosition = {
'offset': '@latest',
'seqNo': -1, #not in use
'enqueuedTime': None, #not in use
'isInclusive': True
}
# Source with default settings
ehConf = {
'eventhubs.connectionString' : cameraHubConnectionString,
'eventhubs.consumerGroup': 'db-ingestion',
'eventhubs.startingPosition': json.dumps(startingEventPosition),
'maxEventsPerTrigger': 5
}
incomingStream = spark \
.readStream \
.format('eventhubs') \
.options(**ehConf) \
.load()
# -
# ## Transform streams to readable dataframes
# 1. First we define the 4 event hubs properties (Offset, Time, Timestamp and Body)
# 1. Then, by using the from_json method on the Body property, we apply the above defined schema
# 1. After this, we select from the deserialized json the 5 properties which we will ingest
#
# This is the json we'll get
# ```json
# {"TrajectId":"01","CameraId":"Camera1","EventTime":"2019-12-09T09:59:58.2710792+00:00","Lane":"2","Country":"BE","LicensePlate":"1-KHC-729","Make":"Renault","Color":"Gray"}
# ```
# Define parsing query selecting the required properties from the incoming telemetry data
cameraData = \
incomingStream \
.withColumn('Offset', col('offset')) \
.withColumn('Body', col('body')) \
.withColumn('CameraEvents', from_json(col('Body').cast(StringType()), cameraEventSchema)) \
.withColumn('Time (readable)', col('CameraEvents.EventTime').cast(TimestampType())) \
.withColumn('Timestamp', col('enqueuedTime')) \
.withColumn('TrajectId', col('CameraEvents.TrajectId').cast(StringType())) \
.withColumn('CameraId', col('CameraEvents.CameraId').cast(StringType())) \
.withColumn('EventTime', col('CameraEvents.EventTime').cast(TimestampType())) \
.withColumn('Lane', col('CameraEvents.Lane').cast(IntegerType())) \
.withColumn('Country', col('CameraEvents.Country').cast(StringType())) \
.withColumn('LicensePlate', col('CameraEvents.LicensePlate').cast(StringType())) \
.withColumn('Make', col('CameraEvents.Make').cast(StringType())) \
.withColumn('Color', col('CameraEvents.Color').cast(StringType())) \
.select('TrajectId', 'CameraId', 'EventTime', 'Lane', 'Country', 'LicensePlate', 'Make', 'Color') \
# ## Stream all iot telemetry to Spark table
#
# This is needed as having multiple queries on the same EventHub stream would result in epoch issues.
#
# For this, the [following stackoverflow post](https://stackoverflow.com/questions/54750779/reusing-an-event-hub-stream-for-multiple-queries-in-azure-data-bricks/54761116#54761116) gives more details.
delta_table_name = 'CameraTelemetry' + datetime.today().strftime('%Y%m%d')
print('Saving all data in table', delta_table_name)
cameraData.writeStream \
.format('delta') \
.outputMode('append') \
.option('checkpointLocation', '/data/' + delta_table_name + '/_checkpoints/data_file') \
.table(delta_table_name)
| src/databricks/notebooks/traffic-camera-ingestion.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## All stations models
# +
import os
import datetime
import IPython
import IPython.display
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
import math
from scipy import stats
import re
import json
mpl.rcParams['figure.figsize'] = (8, 6)
mpl.rcParams['axes.grid'] = False
# %matplotlib inline
# # %run model_utils.ipynb
# +
### Set fixed variables
input_path = 'data/PROCESSED/STATIONS_CLEAN'
path = 'data/PROCESSED/MODEL_OUTPUTS'
## sampling frequency
sample_freq = 60 #(time in minutes)
steps=int(sample_freq/5)
time_steps = int(60/sample_freq)
#60min - 12 steps
#30min - 6 steps
#15min - 3 steps
#5min - 1 step
MAX_EPOCHS = 20
# -
files_ids = {'guadalupe':'guadalupe_validation.csv',
'banos':'banos_validation.csv',
'aeropuerto': 'aeropuerto_validation.csv',
'quisapincha':'quisapincha_validation.csv',
'chiquiurco':'chiquiurco_validation.csv',
'AJSucre':'AJSucre_validation.csv',
'JAlvarez':'JAlvarez_validation.csv',
'pfc-hgpt':'pfc-hgpt_validation.csv',
'calamaca':'calamaca_validation.csv',
'mulaCorral':'mulaCorral_validation.csv',
'pampasSalasaca':'pampasSalasaca_validation.csv',
'tasinteo':'tasinteo_validation.csv',
'pisayambo':'pisayambo_validation.csv',
'cunchibamba':'cunchibamba-hgpt_validation.csv'
}
## cuchimbamba??
def selectCols(df):
d = dict(zip(df.columns[1:20],df.columns[1:20]+f'_{station[0:3]}'))
#df = df[['Date_Time', 'ATAvg', 'RHAvg','WDAvg','WSAvg','WAvgx', 'WAvgy','Day_sin', 'Day_cos', 'Year_sin', 'Year_cos']]
df = df[['Date_Time', 'ATAvg', 'RHAvg','WSAvg','WAvgx','Day_sin', 'Day_cos']]
df.rename(columns=d, inplace= True)
df.set_index('Date_Time', inplace =True)
df.index = pd.to_datetime(df.index)
return df
### combine all stations in one df
for idx, station in enumerate(files_ids.keys()):
if idx == 0:
df = pd.read_csv(f'{input_path}/{station}_validation.csv')
merged = selectCols(df)
else:
df1 = pd.read_csv(f'{input_path}/{station}_validation.csv')
df1 = selectCols(df1)
merged=pd.merge(merged,df1, how='inner', left_index=True, right_index=True)
merged.columns
merged.head()
print(f'len = {len(merged)}')
merged.drop_duplicates(keep = 'first',inplace=True)
print(f'len (after duplicate drop = {len(merged)}')
merged.isnull().values.any()
#delete outliers an dfill with the meadian value
Q1 = merged.quantile(0.25)
Q3 = merged.quantile(0.75)
IQR = Q3 - Q1
print(IQR)
for i,col in enumerate(merged.columns):
merged[col] = np.where((merged[col] < (Q1[i] - 1.5 * IQR[i]))|(merged[col] > (Q3[i] + 1.5 * IQR[i])), np.nan, merged[col])
merged.isnull().values.any()
merged['month']= merged.index.month
merged['hour']= merged.index.hour
for v in merged.columns:
merged[v] = merged.groupby(['hour','month'])[v].transform(lambda x: x.fillna(x.median()))
merged.drop(columns=['month','hour'],inplace=True)
merged.isnull().values.any()
### Subsample to get data for every hour (starting from index 0, get 12 steps)
df = merged[0::steps]
print(f'starts at = {df.index.min().date()}')
print(f'ends at = {df.index.max().date()}')
# +
## Select variables of interest
r = re.compile(f'ATAvg.*') # only csv files
temp = list(filter(r.match, list(df.columns)))
r = re.compile(f'RHAvg.*') # only csv files
rh = list(filter(r.match, list(df.columns)))
# +
### Split data into 70%, 20%, 10% split for the training, validation, and test sets
column_indices = {name: i for i, name in enumerate(df.columns)}
n = len(df)
train_df = df[0:int(n*0.7)]
val_df = df[int(n*0.7):int(n*0.9)]
test_df = df[int(n*0.9):]
num_features = df.shape[1]
# +
# ### Normalize the data ### ROOM TO MAKE TESTS (this is just an average)
# train_mean = train_df.mean()
# train_std = train_df.std()
# train_df = (train_df - train_mean) / train_std
# val_df = (val_df - train_mean) / train_std
# test_df = (test_df - train_mean) / train_std
# -
df.head()
date_time = pd.to_datetime(df.index, format='%Y-%m-%d %H:%M:%S')
train_df.to_csv('data/PROCESSED/TRAIN_TEST_VAL/all_train_df_raw.csv')
test_df.to_csv('data/PROCESSED/TRAIN_TEST_VAL/all_test_df_raw.csv')
val_df.to_csv('data/PROCESSED/TRAIN_TEST_VAL/all_val_df_raw.csv')
## read functions
# %run model_utils.ipynb
# ## MinMax
train_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_train_df_raw.csv')
test_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_test_df_raw.csv')
val_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_val_df_raw.csv')
train_df.set_index('Date_Time', inplace =True)
train_df.index = pd.to_datetime(train_df.index)
test_df.set_index('Date_Time', inplace =True)
test_df.index = pd.to_datetime(test_df.index)
val_df.set_index('Date_Time', inplace =True)
val_df.index = pd.to_datetime(val_df.index)
### make a copy of unscaled data
train_df_raw = train_df.copy()
test_df_raw = test_df.copy()
val_df_raw = val_df.copy()
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
train_df[train_df.columns] = scaler.fit_transform(train_df[train_df.columns])
test_df[test_df.columns] = scaler.fit_transform(test_df[test_df.columns])
val_df[val_df.columns] = scaler.fit_transform(val_df[val_df.columns])
scaler_type='minmax'
IPython.display.clear_output()
train_df.to_csv('data/PROCESSED/TRAIN_TEST_VAL/all_train_df_minmax.csv')
test_df.to_csv('data/PROCESSED/TRAIN_TEST_VAL/all_test_df_minmax.csv')
val_df.to_csv('data/PROCESSED/TRAIN_TEST_VAL/all_val_df_minmax.csv')
station = 'all'
num_features = train_df.shape[1]
vars_to_analize = temp+rh ## get temperature and rel humidity variables
batch_size = 32
MAX_EPOCHS = 20
input_width = 48*time_steps
OUT_STEPS =24*time_steps
scaler_type = 'minmax'
# +
multi_val_performance = {}
multi_performance = {}
r2 ={}
## window
window = WindowGenerator(
input_width=input_width, label_width=OUT_STEPS, shift=OUT_STEPS)
window.plot(plot_col=list(window.column_indices.keys())[0])
# +
# %%time
### RNN
print(f'RNN')
multi_lstm_model = tf.keras.Sequential([
# Shape [batch, time, features] => [batch, lstm_units]
# Adding more `lstm_units` just overfits more quickly.
tf.keras.layers.LSTM(batch_size, return_sequences=False),
# Shape => [batch, out_steps*features]
tf.keras.layers.Dense(OUT_STEPS*num_features,
kernel_initializer=tf.initializers.zeros()),
# Shape => [batch, out_steps, features]
tf.keras.layers.Reshape([OUT_STEPS, num_features])
])
history = compile_and_fit(multi_lstm_model, window)
IPython.display.clear_output()
multi_val_performance[f'MultiLSTM_model_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = multi_lstm_model.evaluate(window.val)
multi_performance[f'MultiLSTM_model_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = multi_lstm_model.evaluate(window.test, verbose=0)
losses = pd.DataFrame(history.history)
losses.plot()
plt.savefig(f'{path}/{station}_multi_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_losses.png',dpi=100)
per = pd.DataFrame.from_dict(multi_performance, orient='index',columns=['loss_test','mae_test'])
val= pd.DataFrame.from_dict(multi_val_performance, orient='index',columns=['loss_val','mae_val'])
pd.merge(per, val, how='inner',left_index=True, right_index =True).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_overall.csv')
# +
# %%time
accuracy={}
plot_col = vars_to_analize
scaler_type='minmax'
for col in range(len(plot_col)):
plot_col_index = window.column_indices[plot_col[col]]
all_preds=[]
all_labels =[]
n_batches = len(tuple(window.test))
for i in range(n_batches):
#print(f'i = {i}')
for inputs, labels in window.test.take(i): # iterate over batches
numpy_labels = labels.numpy() ### get labels
numpy_inputs = inputs.numpy() ### get inputs
preds = multi_lstm_model(numpy_inputs) ### make prediction from trined model
numpy_preds = preds.numpy() ### get predictions
all_preds_by_time = []
all_labels_by_time = []
scaler = MinMaxScaler()
obj = scaler.fit(test_df_raw)
for j in range(numpy_labels.shape[1]): ## number of time steps
### get values for each bacth and time and de-normalize
#print(f'j = {j}')
batch_pred = obj.inverse_transform(numpy_preds[:,j,:])[:,plot_col_index]
batch_label = obj.inverse_transform(numpy_labels[:,j,:])[:,plot_col_index]
all_preds_by_time.extend(batch_pred)
#print(f'all_preds_by_time = {len(all_preds_by_time)}')
all_labels_by_time.extend(batch_label)
all_preds.append(all_preds_by_time)
all_labels.append(all_labels_by_time)
if len(all_preds) >= i:
break
## covert to array (shape= i,time*batch_size)
multi_preds = np.vstack(all_preds)
multi_labels = np.vstack(all_labels)
mae_pred = []
r2_pred = []
mse_pred =[]
rmse_pred = []
for a in np.arange(0,multi_labels.shape[1],step=batch_size):
multi_labels[:,a:a+batch_size]= np.nan_to_num(multi_labels[:,a:a+batch_size],100) ## substitute NANwith 100 for RH
multi_preds[:,a:a+batch_size]= np.nan_to_num(multi_preds[:,a:a+batch_size],100)
mae = mean_absolute_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
mae_pred.append(mae)
mse = mean_squared_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
mse_pred.append(mse)
rmse = math.sqrt(mse)
rmse_pred.append(rmse)
r2 = round(r2_score(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size]),3)
r2_pred.append(r2)
# if a == 0:
# fig, ax = plt.subplots(1, 4, figsize=(20, 5))
# #plt.suptitle(f'{multi_lstm_model}, window: {window.input_width}_{window.shift}',fontsize = 14)
# ax[0].plot(df.index, df.mae, '-o',c='#ff5555')
# ax[0].set_xlabel(f'prediction times {plot_col[col]}')
# ax[0].set_ylabel(f'MAE {plot_col[col]} [de-normed]')
# ax[3].plot(df.index, df.r2,'-o', c='#0ca4b4')
# ax[3].set_xlabel(f'prediction times {plot_col[col]}')
# ax[3].set_ylabel(f'R2 {plot_col[col]} [de-normed]')
# ax[1].plot(df.index, df.mse,'-o', c='#ff5555')
# ax[1].set_xlabel(f'prediction times {plot_col[col]}')
# ax[1].set_ylabel(f'MSE {plot_col[col]} [de-normed]')
# ax[2].plot(df.index, df.rmse, '-o',c='#ff5555')
# ax[2].set_xlabel(f'prediction times {plot_col[col]}')
# ax[2].set_ylabel(f'RMSE {plot_col[col]} [de-normed]')
df = pd.DataFrame(mae_pred, columns=['mae'])
df['r2']=r2_pred
df['mse']=mse_pred
df['rmse']=rmse_pred
accuracy[plot_col[col]] = {'r2':r2_pred,
'mae':mae_pred,
'mse': mse_pred,
'rmse':rmse_pred}
r2 ={}
r2[f'MultiLSTM_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = accuracy
pd.concat({k: pd.DataFrame(v).T for k, v in r2.items()}, axis=0).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_times.csv')
# -
# ## Robust
## read functions
# %run model_utils.ipynb
train_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_train_df_raw.csv')
test_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_test_df_raw.csv')
val_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_val_df_raw.csv')
train_df.set_index('Date_Time', inplace =True)
train_df.index = pd.to_datetime(train_df.index)
test_df.set_index('Date_Time', inplace =True)
test_df.index = pd.to_datetime(test_df.index)
val_df.set_index('Date_Time', inplace =True)
val_df.index = pd.to_datetime(val_df.index)
### make a copy of unscaled data
train_df_raw = train_df.copy()
test_df_raw = test_df.copy()
val_df_raw = val_df.copy()
from sklearn.preprocessing import RobustScaler
scaler = RobustScaler()
train_df[train_df.columns] = scaler.fit_transform(train_df[train_df.columns])
test_df[test_df.columns] = scaler.fit_transform(test_df[test_df.columns])
val_df[val_df.columns] = scaler.fit_transform(val_df[val_df.columns])
scaler_type='robust'
IPython.display.clear_output()
train_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_train_df_{scaler_type}.csv')
test_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_test_df_{scaler_type}.csv')
val_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_val_df_{scaler_type}.csv')
station = 'all'
num_features = train_df.shape[1]
vars_to_analize = temp+rh ## get temperature and rel humidity variables
batch_size = 32
MAX_EPOCHS = 20
input_width = 48*time_steps
OUT_STEPS =24*time_steps
# +
multi_val_performance = {}
multi_performance = {}
r2 ={}
## window
window = WindowGenerator(
input_width=input_width, label_width=OUT_STEPS, shift=OUT_STEPS)
window.plot(plot_col=list(window.column_indices.keys())[0])
# +
# %%time
### RNN
print(f'RNN')
multi_lstm_model = tf.keras.Sequential([
# Shape [batch, time, features] => [batch, lstm_units]
# Adding more `lstm_units` just overfits more quickly.
tf.keras.layers.LSTM(batch_size, return_sequences=False),
# Shape => [batch, out_steps*features]
tf.keras.layers.Dense(OUT_STEPS*num_features,
kernel_initializer=tf.initializers.zeros()),
# Shape => [batch, out_steps, features]
tf.keras.layers.Reshape([OUT_STEPS, num_features])
])
history = compile_and_fit(multi_lstm_model, window)
IPython.display.clear_output()
multi_val_performance[f'MultiLSTM_model_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = multi_lstm_model.evaluate(window.val)
multi_performance[f'MultiLSTM_model_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = multi_lstm_model.evaluate(window.test, verbose=0)
losses = pd.DataFrame(history.history)
losses.plot()
plt.savefig(f'{path}/{station}_multi_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_losses.png',dpi=100)
per = pd.DataFrame.from_dict(multi_performance, orient='index',columns=['loss_test','mae_test'])
val= pd.DataFrame.from_dict(multi_val_performance, orient='index',columns=['loss_val','mae_val'])
pd.merge(per, val, how='inner',left_index=True, right_index =True).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_overall.csv')
r2[f'MultiLSTM_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = window.get_predictions(model=multi_lstm_model,plot_col =vars_to_analize, scaler_type = 'robust',plot=False)
pd.concat({k: pd.DataFrame(v).T for k, v in r2.items()}, axis=0).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_times.csv')
# +
accuracy={}
plot_col = vars_to_analize
scaler_type='robust'
col = 0
for col in range(len(plot_col)):
plot_col_index = window.column_indices[plot_col[col]]
all_preds=[]
all_labels =[]
n_batches = len(tuple(window.test))
for i in range(n_batches):
#print(f'i = {i}')
for inputs, labels in window.test.take(i): # iterate over batches
numpy_labels = labels.numpy() ### get labels
numpy_inputs = inputs.numpy() ### get inputs
preds = multi_lstm_model(numpy_inputs) ### make prediction from trined model
numpy_preds = preds.numpy() ### get predictions
all_preds_by_time = []
all_labels_by_time = []
scaler = RobustScaler()
obj = scaler.fit(test_df_raw)
for j in range(numpy_labels.shape[1]): ## number of time steps
### get values for each bacth and time and de-normalize
#print(f'j = {j}')
batch_pred = obj.inverse_transform(numpy_preds[:,j,:])[:,plot_col_index]
batch_label = obj.inverse_transform(numpy_labels[:,j,:])[:,plot_col_index]
all_preds_by_time.extend(batch_pred)
#print(f'all_preds_by_time = {len(all_preds_by_time)}')
all_labels_by_time.extend(batch_label)
all_preds.append(all_preds_by_time)
all_labels.append(all_labels_by_time)
if len(all_preds) >= i:
break
## covert to array (shape= i,time*batch_size)
multi_preds = np.vstack(all_preds)
multi_labels = np.vstack(all_labels)
mae_pred = []
r2_pred = []
mse_pred =[]
rmse_pred = []
for a in np.arange(0,multi_labels.shape[1],step=batch_size):
mae = mean_absolute_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
mae_pred.append(mae)
mse = mean_squared_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
mse_pred.append(mse)
rmse = math.sqrt(mse)
rmse_pred.append(rmse)
r2 = round(r2_score(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size]),3)
r2_pred.append(r2)
df = pd.DataFrame(mae_pred, columns=['mae'])
df['r2']=r2_pred
df['mse']=mse_pred
df['rmse']=rmse_pred
accuracy[plot_col[col]] = {'r2':r2_pred,
'mae':mae_pred,
'mse': mse_pred,
'rmse':rmse_pred}
if a == 0:
fig, ax = plt.subplots(1, 4, figsize=(20, 5))
#plt.suptitle(f'{multi_lstm_model}, window: {window.input_width}_{window.shift}',fontsize = 14)
ax[0].plot(df.index, df.mae, '-o',c='#ff5555')
ax[0].set_xlabel(f'prediction times {plot_col[col]}')
ax[0].set_ylabel(f'MAE {plot_col[col]} [de-normed]')
ax[3].plot(df.index, df.r2,'-o', c='#0ca4b4')
ax[3].set_xlabel(f'prediction times {plot_col[col]}')
ax[3].set_ylabel(f'R2 {plot_col[col]} [de-normed]')
ax[1].plot(df.index, df.mse,'-o', c='#ff5555')
ax[1].set_xlabel(f'prediction times {plot_col[col]}')
ax[1].set_ylabel(f'MSE {plot_col[col]} [de-normed]')
ax[2].plot(df.index, df.rmse, '-o',c='#ff5555')
ax[2].set_xlabel(f'prediction times {plot_col[col]}')
ax[2].set_ylabel(f'RMSE {plot_col[col]} [de-normed]')
r2[f'MultiLSTM_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = accuracy
pd.concat({k: pd.DataFrame(v).T for k, v in r2.items()}, axis=0).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_times.csv')
# -
r2 ={}
r2[f'MultiLSTM_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = accuracy
pd.concat({k: pd.DataFrame(v).T for k, v in r2.items()}, axis=0).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_times.csv')
# ## Power
## read functions
# %run model_utils.ipynb
train_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_train_df_raw.csv')
test_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_test_df_raw.csv')
val_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_val_df_raw.csv')
train_df.set_index('Date_Time', inplace =True)
train_df.index = pd.to_datetime(train_df.index)
test_df.set_index('Date_Time', inplace =True)
test_df.index = pd.to_datetime(test_df.index)
val_df.set_index('Date_Time', inplace =True)
val_df.index = pd.to_datetime(val_df.index)
### make a copy of unscaled data
train_df_raw = train_df.copy()
test_df_raw = test_df.copy()
val_df_raw = val_df.copy()
from sklearn.preprocessing import PowerTransformer
scaler = PowerTransformer()
train_df[train_df.columns] = scaler.fit_transform(train_df[train_df.columns])
test_df[test_df.columns] = scaler.fit_transform(test_df[test_df.columns])
val_df[val_df.columns] = scaler.fit_transform(val_df[val_df.columns])
scaler_type='power'
IPython.display.clear_output()
train_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_train_df_{scaler_type}.csv')
test_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_test_df_{scaler_type}.csv')
val_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_val_df_{scaler_type}.csv')
station = 'all'
num_features = train_df.shape[1]
vars_to_analize = temp+rh ## get temperature and rel humidity variables
batch_size = 32
MAX_EPOCHS = 20
input_width = 48*time_steps
OUT_STEPS =24*time_steps
# +
multi_val_performance = {}
multi_performance = {}
r2 ={}
## window
window = WindowGenerator(
input_width=input_width, label_width=OUT_STEPS, shift=OUT_STEPS)
window.plot(plot_col=list(window.column_indices.keys())[0])
# +
# %%time
### RNN
print(f'RNN')
multi_lstm_model = tf.keras.Sequential([
# Shape [batch, time, features] => [batch, lstm_units]
# Adding more `lstm_units` just overfits more quickly.
tf.keras.layers.LSTM(batch_size, return_sequences=False),
# Shape => [batch, out_steps*features]
tf.keras.layers.Dense(OUT_STEPS*num_features,
kernel_initializer=tf.initializers.zeros()),
# Shape => [batch, out_steps, features]
tf.keras.layers.Reshape([OUT_STEPS, num_features])
])
history = compile_and_fit(multi_lstm_model, window)
IPython.display.clear_output()
multi_val_performance[f'MultiLSTM_model_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = multi_lstm_model.evaluate(window.val)
multi_performance[f'MultiLSTM_model_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = multi_lstm_model.evaluate(window.test, verbose=0)
losses = pd.DataFrame(history.history)
losses.plot()
plt.savefig(f'{path}/{station}_multi_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_losses.png',dpi=100)
val= pd.DataFrame.from_dict(multi_val_performance, orient='index',columns=['loss_val','mae_val'])
#r2[f'MultiLSTM_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = window.get_predictions(model=multi_lstm_model,plot_col =vars_to_analize, scaler_type = 'power',plot=False)
#pd.merge(per, val, how='inner',left_index=True, right_index =True).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_overall.csv')
# +
accuracy={}
plot_col = vars_to_analize
scaler_type='power'
for col in range(len(plot_col)):
plot_col_index = window.column_indices[plot_col[col]]
all_preds=[]
all_labels =[]
n_batches = len(tuple(window.test))
for i in range(n_batches):
#print(f'i = {i}')
for inputs, labels in window.test.take(i): # iterate over batches
numpy_labels = labels.numpy() ### get labels
numpy_inputs = inputs.numpy() ### get inputs
preds = multi_lstm_model(numpy_inputs) ### make prediction from trined model
numpy_preds = preds.numpy() ### get predictions
all_preds_by_time = []
all_labels_by_time = []
scaler = PowerTransformer()
obj = scaler.fit(test_df_raw)
for j in range(numpy_labels.shape[1]): ## number of time steps
### get values for each bacth and time and de-normalize
#print(f'j = {j}')
batch_pred = obj.inverse_transform(numpy_preds[:,j,:])[:,plot_col_index]
batch_label = obj.inverse_transform(numpy_labels[:,j,:])[:,plot_col_index]
all_preds_by_time.extend(batch_pred)
#print(f'all_preds_by_time = {len(all_preds_by_time)}')
all_labels_by_time.extend(batch_label)
all_preds.append(all_preds_by_time)
all_labels.append(all_labels_by_time)
if len(all_preds) >= i:
break
## covert to array (shape= i,time*batch_size)
multi_preds = np.vstack(all_preds)
multi_labels = np.vstack(all_labels)
mae_pred = []
r2_pred = []
mse_pred =[]
rmse_pred = []
for a in np.arange(0,multi_labels.shape[1],step=batch_size):
multi_labels[:,a:a+batch_size]= np.nan_to_num(multi_labels[:,a:a+batch_size],100) ## substitute NANwith 100 for RH
multi_preds[:,a:a+batch_size]= np.nan_to_num(multi_preds[:,a:a+batch_size],100)
mae = mean_absolute_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
mae_pred.append(mae)
mse = mean_squared_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
mse_pred.append(mse)
rmse = math.sqrt(mse)
rmse_pred.append(rmse)
r2 = round(r2_score(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size]),3)
r2_pred.append(r2)
df = pd.DataFrame(mae_pred, columns=['mae'])
df['r2']=r2_pred
df['mse']=mse_pred
df['rmse']=rmse_pred
accuracy[plot_col[col]] = {'r2':r2_pred,
'mae':mae_pred,
'mse': mse_pred,
'rmse':rmse_pred}
if a == 0:
fig, ax = plt.subplots(1, 4, figsize=(20, 5))
#plt.suptitle(f'{multi_lstm_model}, window: {window.input_width}_{window.shift}',fontsize = 14)
ax[0].plot(df.index, df.mae, '-o',c='#ff5555')
ax[0].set_xlabel(f'prediction times {plot_col[col]}')
ax[0].set_ylabel(f'MAE {plot_col[col]} [de-normed]')
ax[3].plot(df.index, df.r2,'-o', c='#0ca4b4')
ax[3].set_xlabel(f'prediction times {plot_col[col]}')
ax[3].set_ylabel(f'R2 {plot_col[col]} [de-normed]')
ax[1].plot(df.index, df.mse,'-o', c='#ff5555')
ax[1].set_xlabel(f'prediction times {plot_col[col]}')
ax[1].set_ylabel(f'MSE {plot_col[col]} [de-normed]')
ax[2].plot(df.index, df.rmse, '-o',c='#ff5555')
ax[2].set_xlabel(f'prediction times {plot_col[col]}')
ax[2].set_ylabel(f'RMSE {plot_col[col]} [de-normed]')
r2 ={}
r2[f'MultiLSTM_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = accuracy
pd.concat({k: pd.DataFrame(v).T for k, v in r2.items()}, axis=0).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_times.csv')
# -
col
np.isnan(multi_preds[:,a:a+batch_size]).any()
multi_labels[:,a:a+batch_size]= np.nan_to_num(multi_labels[:,a:a+batch_size],100)
mean_absolute_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
# ## Standard
## read functions
# %run model_utils.ipynb
train_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_train_df_raw.csv')
test_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_test_df_raw.csv')
val_df= pd.read_csv('data/PROCESSED/TRAIN_TEST_VAL/all_val_df_raw.csv')
train_df.set_index('Date_Time', inplace =True)
train_df.index = pd.to_datetime(train_df.index)
test_df.set_index('Date_Time', inplace =True)
test_df.index = pd.to_datetime(test_df.index)
val_df.set_index('Date_Time', inplace =True)
val_df.index = pd.to_datetime(val_df.index)
### make a copy of unscaled data
train_df_raw = train_df.copy()
test_df_raw = test_df.copy()
val_df_raw = val_df.copy()
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
train_df[train_df.columns] = scaler.fit_transform(train_df[train_df.columns])
test_df[test_df.columns] = scaler.fit_transform(test_df[test_df.columns])
val_df[val_df.columns] = scaler.fit_transform(val_df[val_df.columns])
scaler_type='stand'
IPython.display.clear_output()
train_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_train_df_{scaler_type}.csv')
test_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_test_df_{scaler_type}.csv')
val_df.to_csv(f'data/PROCESSED/TRAIN_TEST_VAL/all_val_df_{scaler_type}.csv')
# +
## Select variables of interest
r = re.compile(f'ATAvg.*') # only csv files
temp = list(filter(r.match, list(train_df.columns)))
r = re.compile(f'RHAvg.*') # only csv files
rh = list(filter(r.match, list(train_df.columns)))
# -
station = 'all'
num_features = train_df.shape[1]
vars_to_analize = temp+rh ## get temperature and rel humidity variables
batch_size = 32
MAX_EPOCHS = 20
input_width = 48*time_steps
OUT_STEPS =24*time_steps
# +
multi_val_performance = {}
multi_performance = {}
r2 ={}
## window
window = WindowGenerator(
input_width=input_width, label_width=OUT_STEPS, shift=OUT_STEPS)
window.plot(plot_col=list(window.column_indices.keys())[0])
# +
# %%time
### RNN
print(f'RNN')
multi_lstm_model = tf.keras.Sequential([
# Shape [batch, time, features] => [batch, lstm_units]
# Adding more `lstm_units` just overfits more quickly.
tf.keras.layers.LSTM(batch_size, return_sequences=False),
# Shape => [batch, out_steps*features]
tf.keras.layers.Dense(OUT_STEPS*num_features,
kernel_initializer=tf.initializers.zeros()),
# Shape => [batch, out_steps, features]
tf.keras.layers.Reshape([OUT_STEPS, num_features])
])
history = compile_and_fit(multi_lstm_model, window)
IPython.display.clear_output()
multi_val_performance[f'MultiLSTM_model_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = multi_lstm_model.evaluate(window.val)
multi_performance[f'MultiLSTM_model_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = multi_lstm_model.evaluate(window.test, verbose=0)
losses = pd.DataFrame(history.history)
losses.plot()
plt.savefig(f'{path}/{station}_multi_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_losses.png',dpi=100)
per = pd.DataFrame.from_dict(multi_performance, orient='index',columns=['loss_test','mae_test'])
val= pd.DataFrame.from_dict(multi_val_performance, orient='index',columns=['loss_val','mae_val'])
pd.merge(per, val, how='inner',left_index=True, right_index =True).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_overall.csv')
# +
# %%time
accuracy={}
plot_col = vars_to_analize
scaler_type='stand'
for col in range(len(plot_col)):
plot_col_index = window.column_indices[plot_col[col]]
all_preds=[]
all_labels =[]
n_batches = len(tuple(window.test))
for i in range(n_batches):
#print(f'i = {i}')
for inputs, labels in window.test.take(i): # iterate over batches
numpy_labels = labels.numpy() ### get labels
numpy_inputs = inputs.numpy() ### get inputs
preds = multi_lstm_model(numpy_inputs) ### make prediction from trined model
numpy_preds = preds.numpy() ### get predictions
all_preds_by_time = []
all_labels_by_time = []
scaler = StandardScaler()
obj = scaler.fit(test_df_raw)
for j in range(numpy_labels.shape[1]): ## number of time steps
### get values for each bacth and time and de-normalize
#print(f'j = {j}')
batch_pred = obj.inverse_transform(numpy_preds[:,j,:])[:,plot_col_index]
batch_label = obj.inverse_transform(numpy_labels[:,j,:])[:,plot_col_index]
all_preds_by_time.extend(batch_pred)
#print(f'all_preds_by_time = {len(all_preds_by_time)}')
all_labels_by_time.extend(batch_label)
all_preds.append(all_preds_by_time)
all_labels.append(all_labels_by_time)
if len(all_preds) >= i:
break
## covert to array (shape= i,time*batch_size)
multi_preds = np.vstack(all_preds)
multi_labels = np.vstack(all_labels)
mae_pred = []
r2_pred = []
mse_pred =[]
rmse_pred = []
for a in np.arange(0,multi_labels.shape[1],step=batch_size):
multi_labels[:,a:a+batch_size]= np.nan_to_num(multi_labels[:,a:a+batch_size],100) ## substitute NANwith 100 for RH
multi_preds[:,a:a+batch_size]= np.nan_to_num(multi_preds[:,a:a+batch_size],100)
mae = mean_absolute_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
mae_pred.append(mae)
mse = mean_squared_error(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size])
mse_pred.append(mse)
rmse = math.sqrt(mse)
rmse_pred.append(rmse)
r2 = round(r2_score(multi_labels[:,a:a+batch_size], multi_preds[:,a:a+batch_size]),3)
r2_pred.append(r2)
# if a == 0:
# fig, ax = plt.subplots(1, 4, figsize=(20, 5))
# #plt.suptitle(f'{multi_lstm_model}, window: {window.input_width}_{window.shift}',fontsize = 14)
# ax[0].plot(df.index, df.mae, '-o',c='#ff5555')
# ax[0].set_xlabel(f'prediction times {plot_col[col]}')
# ax[0].set_ylabel(f'MAE {plot_col[col]} [de-normed]')
# ax[3].plot(df.index, df.r2,'-o', c='#0ca4b4')
# ax[3].set_xlabel(f'prediction times {plot_col[col]}')
# ax[3].set_ylabel(f'R2 {plot_col[col]} [de-normed]')
# ax[1].plot(df.index, df.mse,'-o', c='#ff5555')
# ax[1].set_xlabel(f'prediction times {plot_col[col]}')
# ax[1].set_ylabel(f'MSE {plot_col[col]} [de-normed]')
# ax[2].plot(df.index, df.rmse, '-o',c='#ff5555')
# ax[2].set_xlabel(f'prediction times {plot_col[col]}')
# ax[2].set_ylabel(f'RMSE {plot_col[col]} [de-normed]')
df = pd.DataFrame(mae_pred, columns=['mae'])
df['r2']=r2_pred
df['mse']=mse_pred
df['rmse']=rmse_pred
accuracy[plot_col[col]] = {'r2':r2_pred,
'mae':mae_pred,
'mse': mse_pred,
'rmse':rmse_pred}
r2 ={}
r2[f'MultiLSTM_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}'] = accuracy
pd.concat({k: pd.DataFrame(v).T for k, v in r2.items()}, axis=0).to_csv(f'{path}/{station}_lstm-{batch_size}-var-{scaler_type}_{sample_freq}m_w{input_width}_{OUT_STEPS}_{MAX_EPOCHS}_performance_times.csv')
# -
| All_stations_scalers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotting data from OMNIWeb Data
#
#
# +
#Import necessary libraries
import numpy as np
import pandas as pd
from datetime import datetime, timedelta
from matplotlib import pyplot as plt
from matplotlib import dates as mpl_dates
# %matplotlib inline
plt.style.use('seaborn')
# -
#Extracting data from text file
data= np.loadtxt('omm.txt')
size= np.shape(data)
df = pd.read_csv("omm.txt",
delim_whitespace=True,
usecols=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9],
names=["Year", "DOY", "Hour", "B", "Bx", "By","Bz","Pd","Kp","Sp"])
df.head()
#Change time
df.index = pd.to_datetime(df["Year"] * 100000 + df["DOY"] * 100 + df["Hour"], format="%Y%j%H")
df = df.drop(columns=["Year", "DOY", "Hour"])
df.head()
# # Generate Plots Directly from Data Frame
df.plot(figsize=(15,4),color =('#a6cee3','#1f78b4','#b2df8a','#33a02c','#fbb4b9','#765F2E'))
# ## Generating Subplots for each Variable
ax= df.plot(subplots=True, figsize=(15,10),color=('#AE9C45','#6073B1','#A7B8F8','#052955'))
plt.xlabel('Date')
plt.ylabel('Different Parameters')
# ## Alternative Way
# Extracting Data from file.
B = data[:,3]
Bx = data[:,4]
By = data[:,5]
Bz= data[:,6]
Pd = data[:,7] #Proton Density
Kp = data[:,8] #Kp Index*10
Sp= data[:,9] #sunspot no
# +
#Generating plots
plt.figsize=(20,20)
plt.plot(df.index,B,linestyle=':',linewidth = 1, color='k', label="B")
plt.plot(df.index,Bx,linestyle='-',linewidth = 1, color='#444444',label="Bx")
plt.plot(df.index,By,linestyle='--',linewidth = 1, color='#41b6c4',label="By")
plt.plot(df.index,Bz,linestyle='solid',linewidth = 1, color='#2c7fb8',label="Bz")
plt.plot(df.index,Pd,linestyle='dashdot',linewidth = 1, color='y',label="Plasma density")
plt.plot(df.index,Kp,linestyle='dashed',linewidth = 1, color='#018571',label="Kp Index")
plt.plot(df.index,Sp,linestyle='dotted',linewidth = 1, color='#253494',label="No of Sunspots")
plt.legend()
import matplotlib
matplotlib.rc('xtick', labelsize=10)
matplotlib.rc('ytick', labelsize=10)
plt.title('Solar Activity 1-11,September,2017')
plt.xlabel('Date')
plt.ylabel('Data')
#Saving the plot
plt.savefig("OP3.png",dpi=300)
# -
# ## Plot the data creating subplots
# +
#Generating Subplots
fig,(ax1,ax2,ax3,ax4,ax5,ax6,ax7) = plt.subplots(7,1,figsize=(50,50), sharex=True)
ax1.plot(df.index,B,linestyle=':',linewidth = 7, color='k', label="B",)
ax2.plot(df.index,Bx,linestyle='-',linewidth = 7, color='#a1dab4',label="Bx")
ax3.plot(df.index,By,linestyle='--',linewidth = 7, color='#41b6c4',label="By")
ax4.plot(df.index,Bz,linestyle='solid',linewidth = 7, color='#2c7fb8',label="Bz")
ax5.plot(df.index,Pd,linestyle='dashdot',linewidth = 7, color='#80cdc1',label="Plasma density")
ax6.plot(df.index,Kp,linestyle='dashed',linewidth = 7, color='#018571',label="Kp Index")
ax7.plot(df.index,Sp,linestyle='dotted',linewidth = 7, color='#253494',label="No of Sunspots")
#Add legend
fig.legend(fontsize=30) # using a size in points
#Set X,Y axis label for each subplots
plt.xlabel('Date',fontsize=40)
ax1.set_ylabel('IMF Magnitude, B', fontsize= 30)
ax2.set_ylabel( 'Bx', fontsize= 30)
ax3.set_ylabel( 'By', fontsize= 30)
ax4.set_ylabel( 'Bz', fontsize= 30)
ax5.set_ylabel( 'Plasma Density', fontsize= 30)
ax6.set_ylabel( 'Kp Index* 10', fontsize= 30)
ax7.set_ylabel( 'No. of Sunspots', fontsize= 30)
import matplotlib
matplotlib.rc('xtick', labelsize=50)
matplotlib.rc('ytick', labelsize=50)
#Saving the plot
plt.savefig("OP.png",dpi=300)
# -
| OMNIWEB Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# $$ \LaTeX \text{ command declarations here.}
# \newcommand{\R}{\mathbb{R}}
# \renewcommand{\vec}[1]{\mathbf{#1}}
# \newcommand{\X}{\mathcal{X}}
# \newcommand{\D}{\mathcal{D}}
# \newcommand{\G}{\mathcal{G}}
# \newcommand{\Parents}{\mathrm{Parents}}
# \newcommand{\NonDesc}{\mathrm{NonDesc}}
# \newcommand{\I}{\mathcal{I}}
# $$
# + slideshow={"slide_type": "skip"}
from __future__ import division
# scientific
# %matplotlib inline
from matplotlib import pyplot as plt;
import numpy as np;
# ipython
import IPython;
# python
import os;
#####################################################
# image processing
import PIL;
# trim and scale images
def trim(im, percent=100):
print("trim:", percent);
bg = PIL.Image.new(im.mode, im.size, im.getpixel((0,0)))
diff = PIL.ImageChops.difference(im, bg)
diff = PIL.ImageChops.add(diff, diff, 2.0, -100)
bbox = diff.getbbox()
if bbox:
x = im.crop(bbox)
return x.resize(((x.size[0]*percent)//100,
(x.size[1]*percent)//100),
PIL.Image.ANTIALIAS);
#####################################################
# daft (rendering PGMs)
import daft;
# set to FALSE to load PGMs from static images
RENDER_PGMS = False;
# decorator for pgm rendering
def pgm_render(pgm_func):
def render_func(path, percent=100, render=None, *args, **kwargs):
print("render_func:", percent);
# render
render = render if (render is not None) else RENDER_PGMS;
if render:
print("rendering");
# render
pgm = pgm_func(*args, **kwargs);
pgm.render();
pgm.figure.savefig(path, dpi=300);
# trim
img = trim(PIL.Image.open(path), percent);
img.save(path, 'PNG');
else:
print("not rendering");
# error
if not os.path.isfile(path):
raise("Error: Graphical model image %s not found."
+ "You may need to set RENDER_PGMS=True.");
# display
return IPython.display.Image(filename=path);#trim(PIL.Image.open(path), percent);
return render_func;
######################################################
# + [markdown] slideshow={"slide_type": "slide"}
# # EECS 445: Machine Learning
# ## Lecture 15: Exponential Families & Bayesian Networks
# * Instructor: **<NAME>**
# * Date: November 7, 2016
#
# *Lecture Exposition Credit:* <NAME> & <NAME>
# + [markdown] slideshow={"slide_type": "slide"}
# ## References
#
# - **[MLAPP]** Murphy, Kevin. [*Machine Learning: A Probabilistic Perspective*](https://mitpress.mit.edu/books/machine-learning-0). 2012.
# - **[Koller & Friedman 2009]** Koller, Daphne and <NAME>. [*Probabilistic Graphical Models*](https://mitpress.mit.edu/books/probabilistic-graphical-models). 2009.
# - **[Hero 2008]** Hero, Alfred O.. [*Statistical Methods for Signal Processing*](http://web.eecs.umich.edu/~hero/Preprints/main_564_08_new.pdf). 2008.
# - **[Blei 2011]** <NAME>. [*Notes on Exponential Families*](https://www.cs.princeton.edu/courses/archive/fall11/cos597C/lectures/exponential-families.pdf). 2011.
# - **[Jordan 2010]** Jordan, <NAME>.. [*The Exponential Family: Basics*](http://www.cs.berkeley.edu/~jordan/courses/260-spring10/other-readings/chapter8.pdf). 2008.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Outline
#
# - Exponential Families
# - Sufficient Statistics & Pitman-Koopman-Darmois Theorem
# - Mean and natural parameters
# - Maximum Likelihood estimation
# - Probabilistic Graphical Models
# - Directed Models (Bayesian Networks)
# - Conditional Independence & Factorization
# - Examples
# + [markdown] slideshow={"slide_type": "slide"}
# # Exponential Families
#
# > Uses material from **[MLAPP]** §9.2 and **[Hero 2008]** §3.5, §4.4.2
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exponential Family: Introduction
#
# We have seen many distributions.
# * Bernoulli
# * Gaussian
# * Exponential
# * Gamma
#
# Many of these belong to a more general class called the **exponential family**.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exponential Family: Introduction
#
# Why do we care?
# * only family of distributions with finite-dimensional **sufficient statistics**
# * only family of distributions for which **conjugate priors** exist
# * makes the least set of assumptions subject to some user-chosen constraints (**Maximum Entropy**)
# * core of generalized linear models and **variational inference**
# + [markdown] slideshow={"slide_type": "slide"}
# ### Sufficient Statistics
#
# **Recall:** A **statistic** $T(\D)$ is a function of the observed data $\D$.
# - Mean, $T(x_1, \dots, x_n) = \frac{1}{n}\sum_{k=1}^n x_k$
# - Variance, maximum, mode, etc.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Sufficient Statistics: Definition
#
# Suppose we have a model $P$ with parameters $\theta$. Then,
#
# > A statistic $T(\D)$ is **sufficient** for $\theta$ if no other statistic calculated from the same sample provides any additional information about the parameter.
#
# That is, if $T(\D_1) = T(\D_2)$, our estimate of $\theta$ given $\D_1$ or $\D_2$ will be the same.
# - Mathematically, $P(\theta | T(\D), \D) = P(\theta | T(\D))$ independently of $\D$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Sufficient Statistics: Example
#
# Suppose $X \sim \mathcal{N}(\mu, \sigma^2)$ and we observe $\mathcal{D} = (x_1, \dots, x_n)$. Let
# - $\hat\mu$ be the sample mean
# - $\hat{\sigma}^2$ be the sample variance
#
# Then $T(\mathcal{D}) = (\hat\mu, \hat{\sigma}^2)$ is sufficient for $\theta=(\mu, \sigma^2)$.
# - Two samples $\D_1$ and $\D_2$ with the same mean and variance give the same estimate of $\theta$
#
# <span style="color:gray">(we are sweeping some details under the rug)</span>
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exponential Family: Definition
#
# **INTUITION**: $p()$ has **exp family form** when density $p(x | \theta)$ can be written as $$\exp(\text{Linear combination of }\theta\text{ and features of }x)$$
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# **DEF:** $p(x | \theta)$ has **exponential family form** if:
# $$
# \begin{align}
# p(x | \theta)
# &= \frac{1}{Z(\theta)} h(x) \exp\left[ \eta(\theta)^T \phi(x) \right] \\
# &= h(x) \exp\left[ \eta(\theta)^T \phi(x) - A(\theta) \right]
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
#
# - $Z(\theta)$ is the **partition function** for normalization
# - $A(\theta) = \log Z(\theta)$ is the **log partition function**
# - $\phi(x) \in \R^d$ is a vector of **sufficient statistics**
# - $\eta(\theta)$ maps $\theta$ to a set of **natural parameters**
# - $h(x)$ is a scaling constant, usually $h(x)=1$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Bernoulli
#
# The Bernoulli distribution can be written as
# $$
# \begin{align}
# \mathrm{Ber}(x | \mu)
# &= \mu^x (1-\mu)^{1-x} \\
# &= \exp\left[ x \log \mu + (1-x) \log (1-\mu) \right] \\
# &= \exp\left[ \eta(\mu)^T \phi(x) \right]
# \end{align}
# $$
#
# where $\eta(\mu) = (\log\mu, \log(1-\mu))$ and $\phi(x) = (x, 1-x)$
# - There is a linear dependence between features $\phi(x)$
# - This representation is **overcomplete**
# - $\eta$ is not uniquely determined
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Bernoulli
#
# Instead, we can find a **minimal** parameterization:
# $$
# \begin{align}
# \mathrm{Ber}(x | \mu)
# &= (1-\mu) \exp\left[ x \log\frac{\mu}{1-\mu} \right]
# \end{align}
# $$
#
# This gives **natural parameters** $\eta = \log \frac{\mu}{1-\mu}$.
# - Now, $\eta$ is unique
# + [markdown] slideshow={"slide_type": "slide"}
# ### Other Examples
#
# Exponential Family Distributions:
# - Multivariate normal
# - Exponential
# - Dirichlet
#
# Non-examples:
# - Student t-distribution can't be written in exponential form
# - Uniform distribution support depends on the parameters $\theta$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Log-Partition Function
#
# Derivatives of the **log-partition function** $A(\theta)$ yield **cumulants** of the sufficient statistics *(Exercise!)*
# - $\nabla_\theta A(\theta) = E[\phi(x)]$
# - $\nabla^2_\theta A(\theta) = \text{Cov}[ \phi(x) ]$
#
# This guarantees that $A(\theta)$ is convex!
# - Its Hessian is the covariance matrix of $X$, which is positive-definite.
# - Later, this will guarantee a unique global maximum of the likelihood!
# + [markdown] slideshow={"slide_type": "skip"}
# #### <span style="color:gray">Proof of Convexity: First Derivative</span>
#
# $$
# \begin{align}
# \frac{dA}{d\theta}
# &= \frac{d}{d\theta} \left[ \log \int exp(\theta\phi(x))h(x)dx \right] \\
# &= \frac{\frac{d}{d\theta} \int exp(\theta\phi(x))h(x)dx)}{\int exp(\theta\phi(x))h(x)dx)} \\
# &= \frac{\int \phi(x)exp(\theta\phi(x))h(x)dx}{exp(A(\theta))} \\
# &= \int \phi(x) \exp[\theta\phi(x)-A(\theta)] h(x) dx \\
# &= \int \phi(x) p(x) dx \\
# &= E[\phi(x)]
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# #### <span style="color:gray">Proof of Convexity: Second Derivative</span>
#
# $$
# \begin{align}
# \frac{d^2A}{d\theta^2}
# & = \int \phi(x)\exp[\theta \phi(x) - A(\theta)] h(x) (\phi(x) - A'(\theta)) dx \\
# & = \int \phi(x) p(x) (\phi(x) - A'(\theta))dx \\
# & = \int \phi^2(x) p(X) dx - A'(\theta) \int \phi(x)p(x)dx \\
# & = E[\phi^2(x)] - E[\phi(x)]^2 \hspace{2em} (\because A'(\theta) = E[\phi(x)]) \\
# & = Var[\phi(x)]
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# #### <span style="color:gray">Proof of Convexity: Second Derivative</span>
#
# For multi-variate case, we have
#
# $$ \frac{\partial^2A}{\partial\theta_i \partial\theta_j} = E[\phi_i(x)\phi_j(x)] - E[\phi_i(x)] E[\phi_j(x)]$$
#
# and hence,
# $$ \nabla^2A(\theta) = Cov[\phi(x)] $$
#
# Since covariance is positive definite, we have $A(\theta)$ convex as required.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exponential Family: Likelihood for a *Set* of Data
#
# For data $\D = (x_1, \dots, x_N)$, the likelihood is
# $$
# p(\D|\theta)
# = \left[ \prod_{k=1}^N h(x_k) \right] Z(\theta)^{-N} \exp\left[ \eta(\theta)^T \left(\sum_{k=1}^N \phi(x_k) \right) \right]
# $$
#
# The sufficient statistics are now $\phi(\D) = \sum_{k=1}^N \phi(x_k)$.
# - **Bernoulli:** $\phi = \# Heads$
# - **Normal:** $\phi = [ \sum_k x_k, \sum_k x_k^2 ]$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exponential Family: MLE
#
# For natural parameters $\theta$ and data $\D = (x_1, \dots, x_N)$,
# $$
# \log p(\D|\theta) = \theta^T \phi(\D) - N A(\theta)
# $$
#
# Since $-A(\theta)$ is concave and $\theta^T\phi(\D)$ linear,
# - the log-likelihood is concave
# - under many conditions, there is a unique global maximum!
# > Strict convexity of the log-partition function requires that we are working with a "regular exponential family". More on this can be found in [These Notes](http://pages.cs.wisc.edu/~jerryzhu/cs731/expfamily.pdf).
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exponential Family: MLE
#
# To find the maximum, recall $\nabla_\theta A(\theta) = E_\theta[\phi(x)]$, so
# \begin{align*}
# \nabla_\theta \log p(\D | \theta) & =
# \nabla_\theta(\theta^T \phi(\D) - N A(\theta)) \\
# & = \phi(\D) - N E_\theta[\phi(X)] = 0
# \end{align*}
# Which gives
# $$E_\theta[\phi(X)] = \frac{\phi(\D)}{N} = \frac{1}{N} \sum_{k=1}^N \phi(x_k)$$
#
# At the MLE $\hat\theta_{MLE}$, the empirical average of sufficient statistics equals their expected value.
# - this is called **moment matching**
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exponential Family: MLE for the Bernoulli
#
# As an example, consider the Bernoulli distribution
# - Sufficient statistic $N$, $\phi(\D) = \# Heads$
#
# $$
# \hat\mu_{MLE} = \frac{\# Heads}{N}
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# ### Bayes for Exponential Family
#
# Exact Bayesian analysis is considerably simplified if the prior is **conjugate** to the likelihood.
# - Simply, this means that prior $p(\theta)$ has the same form as the posterior $p(\theta|\mathcal{D})$.
#
# This requires likelihood to have finite sufficient statistics
# * Exponential family to the rescue!
#
# **Note**: We will release some notes on cojugate priors + exponential families. It's hard to learn from slides and needs a bit more description.
# + [markdown] slideshow={"slide_type": "skip"}
# ### Likelihood for exponential family
#
# Likelihood:
# $$ p(\mathcal{D}|\theta) \propto g(\theta)^N \exp[\eta(\theta)^T s_N]\\
# s_N = \sum_{i=1}^{N}\phi(x_i)$$
#
# In terms of canonical parameters:
# $$ p(\mathcal{D}|\eta) \propto \exp[N\eta^T \bar{s} -N A(\eta)] \\
# \bar s = \frac{1}{N}s_N $$
# + [markdown] slideshow={"slide_type": "skip"}
# ### Conjugate prior for exponential family
#
# * The prior and posterior for an exponential family involve two parameters, $\tau$ and $\nu$, initially set to $\tau_0, \nu_0$
#
# $$ p(\theta| \nu_0, \tau_0) \propto g(\theta)^{\nu_0} \exp[\eta(\theta)^T \tau_0] $$
#
# * Denote $ \tau_0 = \nu_0 \bar{\tau}_0$ to separate out the size of the **prior pseudo-data**, $\nu_0$ , from the mean of the sufficient statistics on this pseudo-data, $\tau_0$ . Hence,
#
# $$ p(\theta| \nu_0, \bar \tau_0) \propto \exp[\nu_0\eta^T \bar \tau_0 - \nu_0 A(\eta)] $$
#
# * Think of $\tau_0$ as a "guess" of the future sufficient statistics, and $\nu_0$ as the strength of this guess
# + [markdown] slideshow={"slide_type": "skip"}
# ### Prior: Example
#
# $$
# \begin{align}
# p(\theta| \nu_0, \tau_0)
# &\propto (1-\theta)^{\nu_0} \exp[\tau_0\log(\frac{\theta}{1-\theta})] \\
# &= \theta^{\tau_0}(1-\theta)^{\nu_0 - \tau_0}
# \end{align}
# $$
#
# Define $\alpha = \tau_0 +1 $ and $\beta = \nu_0 - \tau_0 +1$ to see that this is a **beta distribution**.
# + [markdown] slideshow={"slide_type": "skip"}
# ### Posterior
#
# Posterior:
# $$ p(\theta|\mathcal{D}) = p(\theta|\nu_N, \tau_N) = p(\theta| \nu_0 +N, \tau_0 +s_N) $$
#
# Note that we obtain **hyper-parameters** by adding. Hence,
#
# $$ \begin{align}
# p(\eta|\mathcal{D})
# &\propto \exp[\eta^T (\nu_0 \bar\tau_0 + N \bar s) - (\nu_0 + N) A(\eta) ] \\
# &= p(\eta|\nu_0 + N, \frac{\nu_0 \bar\tau_0 + N \bar s}{\nu_0 + N})
# \end{align}$$
# where $\bar s = \frac 1 N \sum_{i=1}^{N}\phi(x_i)$.
#
# * *posterior hyper-parameters are a convex combination of the prior mean hyper-parameters and the average of the sufficient statistics.*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Break time!
#
# <img src="images/122_yawn_cat_gifs.gif"/>
# + [markdown] slideshow={"slide_type": "slide"}
# # Probabilistic Graphical Models
#
# > Uses material from **[MLAPP]** §10.1, 10.2 and **[<NAME> 2009]**.
# + [markdown] slideshow={"slide_type": "skip"}
# > "I basically know of two principles for treating complicated systems in simple ways: the first is the principle of modularity and the second is the principle of abstraction. I am an apologist for computational probability in machine learning because I believe that probability theory implements these two principles in deep and intriguing ways — nameley through factorization and through averaging. Exploiting these two mechanisms as fully as possible seems to me to be the way forward in machine learning"  – <NAME> (qtd. in MLAPP)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Graphical Models: Motivation
#
# Suppose we observe multiple correlated variables $x=(x_1, \dots, x_n)$.
# - Words in a document
# - Pixels in an image
#
# How can we compactly represent the **joint distribution** $p(x|\theta)$?
# - How can we tractably *infer* one set of variables given another?
# - How can we efficiently *learn* the parameters?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Joint Probability Tables
#
# One (bad) choice is to write down a **Joint Probability Table**.
# - For $n$ binary variables, we must specify $2^n - 1$ probabilities!
# - Expensive to store and manipulate
# - Impossible to learn so many parameters
# - Very hard to interpret!
#
# > Can we be more concise?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Motivating Example: Coin Flips
#
# What is the joint distribution of three independent coin flips?
# - Explicitly specifying the JPT requires $2^3-1=7$ parameters.
#
# Assuming independence, $P(X_1, X_2, X_3) = P(X_1)P(X_2)P(X_3)$
# - Each marginal $P(X_k)$ only requires one parameter, the bias
# - This gives a total of $3$ parameters, compared to $8$.
#
# > Exploiting the **independence structure** of a joint distribution leads to more concise representations.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Motivating Example: Naive Bayes
#
# In Naive Bayes, we assumed the features $X_1, \dots, X_N$ were independent given the class label $C$:
# $$
# P(x_1, \dots, x_N, c) = P(c) \prod_{k=1}^N P(x_k | c)
# $$
#
# This greatly simplified the learning procedure:
# - Allowed us to look at each feature individually
# - Only need to learn $O(CN)$ probabilities, for $C$ classes and $N$ features
# + [markdown] slideshow={"slide_type": "slide"}
# ### Conditional Independence
#
# The key to efficiently representing large joint distributions is to make **conditional independence** assumptions of the form
# $$
# X \perp Y \mid Z
# \iff
# p(X,Y|Z) = p(X|Z)p(Y|Z)
# $$
#
# > Once $z$ is known, information about $x$ does not tell us any information about $y$ and vice versa.
#
# An effective way to represent these assumptions is with a **graph**.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Bayesian Networks: Definition
#
# A **Bayesian Network** $\mathcal{G}$ is a directed acyclic graph whose nodes represent random variables $X_1, \dots, X_n$.
# - Let $\Parents_\G(X_k)$ denote the parents of $X_k$ in $\G$
# - Let $\NonDesc_\G(X_k)$ denote the variables in $\G$ who are not descendants of $X_k$.
#
# > Examples will come shortly...
# + [markdown] slideshow={"slide_type": "slide"}
# ### Bayesian Networks: Local Independencies
#
# Every Bayesian Network $\G$ encodes a set $\I_\ell(\G)$ of **local independence assumptions**:
#
# > For each variable $X_k$, we have $(X_k \perp \NonDesc_\G(X_k) \mid \Parents_\G(X_k))$
#
# Every node $X_k$ is conditionally independent of its nondescendants given its parents.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Naive Bayes
#
# The graphical model for Naive Bayes is shown below:
# - $\Parents_\G(X_k) = \{ C \}$, $\NonDesc_\G(X_k) = \{ X_j \}_{j\neq k} \cup \{ C \}$
# - Therefore $X_j \perp X_k \mid C$ for any $j \neq k$
# + slideshow={"slide_type": "skip"}
@pgm_render
def pgm_naive_bayes():
pgm = daft.PGM([4,3], origin=[-2,0], node_unit=0.8, grid_unit=2.0);
# nodes
pgm.add_node(daft.Node("c", r"$C$", -0.25, 2));
pgm.add_node(daft.Node("x1", r"$X_1$", -1, 1));
pgm.add_node(daft.Node("x2", r"$X_2$", -0.5, 1));
pgm.add_node(daft.Node("dots", r"$\cdots$", 0, 1, plot_params={ 'ec' : 'none' }));
pgm.add_node(daft.Node("xN", r"$X_N$", 0.5, 1));
# edges
pgm.add_edge("c", "x1", head_length=0.08);
pgm.add_edge("c", "x2", head_length=0.08);
pgm.add_edge("c", "xN", head_length=0.08);
return pgm;
# + slideshow={"slide_type": "-"}
# %%capture
pgm_naive_bayes("images/naive-bayes.png");
# + [markdown] slideshow={"slide_type": "slide"}
# ### Subtle Point: Graphs & Distributions
#
# A Bayesian network $\G$ over variables $X_1, \dots, X_N$ encodes a set of **conditional independencies**.
# - Shows independence structure, nothing more.
# - Does **not** tell us how to assign probabilities to a configuration $(x_1, \dots x_N)$ of the variables.
#
# There are **many** distributions $P$ satisfying the independencies in $\G$.
# - Many joint distributions share a common structure, which we exploit in algorithms.
# - The distribution $P$ may satisfy other independencies **not** encoded in $\G$.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Subtle Point: Graphs & Distributions
#
# If $P$ satisfies the independence assertions made by $\G$, we say that
# - $\G$ is an **I-Map** for $P$
# - or that $P$ **satisfies** $\G$.
#
# Any distribution satisfying $\G$ shares common structure.
# - We will exploit this structure in our algorithms
# - This is what makes graphical models so **powerful**!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Review: Chain Rule for Probability
#
# We can factorize any joint distribution via the **Chain Rule for Probability**:
# $$
# \begin{align}
# P(X_1, \dots, X_N)
# &= P(X_1) P(X_2, \dots, X_N | X_1) \\
# &= P(X_1) P(X_2 | X_1) P(X_3, \dots, X_N | X_1, X_2) \\
# &= \prod_{k=1}^N P(X_k | X_1, \dots, X_{k-1})
# \end{align}
# $$
#
# > Here, the ordering of variables is arbitrary. This works for any permutation.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Bayesian Networks: Topological Ordering
#
# Every network $\G$ induces a **topological (partial) ordering** on its nodes:
# > Parents assigned a lower index than their children
# + slideshow={"slide_type": "skip"}
@pgm_render
def pgm_topological_order():
pgm = daft.PGM([4, 4], origin=[-4, 0])
# Nodes
pgm.add_node(daft.Node("x1", r"$1$", -3.5, 2))
pgm.add_node(daft.Node("x2", r"$2$", -2.5, 1.3))
pgm.add_node(daft.Node("x3", r"$3$", -2.5, 2.7))
pgm.add_node(daft.Node("x4", r"$4$", -1.5, 1.6))
pgm.add_node(daft.Node("x5", r"$5$", -1.5, 2.3))
pgm.add_node(daft.Node("x6", r"$6$", -0.5, 1.3))
pgm.add_node(daft.Node("x7", r"$7$", -0.5, 2.7))
# Add in the edges.
pgm.add_edge("x1", "x4", head_length=0.08)
pgm.add_edge("x1", "x5", head_length=0.08)
pgm.add_edge("x2", "x4", head_length=0.08)
pgm.add_edge("x3", "x4", head_length=0.08)
pgm.add_edge("x3", "x5", head_length=0.08)
pgm.add_edge("x4", "x6", head_length=0.08)
pgm.add_edge("x4", "x7", head_length=0.08)
pgm.add_edge("x5", "x7", head_length=0.08)
return pgm;
# + slideshow={"slide_type": "-"}
# %%capture
pgm_topological_order("images/topological-order.png")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Factorization Theorem: Statement
#
# **Theorem:** *(Koller & Friedman 3.1)* If $\G$ is an I-map for $P$, then $P$ **factorizes** as follows:
# $$
# P(X_1, \dots, X_N) = \prod_{k=1}^N P(X_k \mid \Parents_\G(X_k))
# $$
#
# > Let's prove it together!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Factorization Theorem: Proof
#
# First, apply the chain rule to any topological ordering:
# $$
# P(X_1, \dots, X_N) = \prod_{k=1}^N P(X_k \mid X_1, \dots, X_{k-1})
# $$
#
# Consider one of the factors $P(X_k \mid X_1, \dots, X_{k-1})$.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Factorization Theorem: Proof
#
# Since our variables $X_1,\dots,X_N$ are in topological order,
# - $\Parents_\G(X_k) \subseteq \{ X_1, \dots, X_{k-1} \}$
# - None of $X_k$'s descendants can possibly lie in $\{ X_1, \dots, X_{k-1} \}$
# + [markdown] slideshow={"slide_type": "fragment"}
# Therefore, $\{ X_1, \dots, X_{k-1} \} = \Parents_\G(X_k) \cup \mathcal{Z}$
# - for some $\mathcal{Z} \subseteq \NonDesc_\G(X_k)$.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Factorization Theorem: Proof
#
# Recall the following property of conditional independence:
# $$
# ( X \perp Y, W \mid Z ) \implies (X \perp Y \mid Z)
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Since $\G$ is an I-map for $P$ and $\mathcal{Z} \subseteq \NonDesc_\G(X_k)$, we have
# $$\begin{align}
# & (X_k \perp \NonDesc_\G(X_k) \mid \Parents_\G(X_k)) \\
# \implies & (X_k \perp \mathcal{Z} \mid \Parents_\G(X_k))
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Factorization Theorem: Proof
#
# We have just shown $(X_k \perp \mathcal{Z} \mid \Parents_\G(X_k))$, therefore
# $$
# P(X_k \mid X_1, \dots, X_{k-1}) = P(X_k \mid \Parents_\G(X_k))
# $$
#
# - Recall $\{ X_1, \dots, X_{k-1} \} = \Parents_\G(X_k) \cup \mathcal{Z}$.
#
# > **Remember:** $X_k$ is conditionally independent of its nondescendants given its parents!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Factorization Theorem: End of Proof
#
# Applying this to every factor, we see that
# $$
# \begin{align}
# P(X_1, \dots, X_N)
# &= \prod_{k=1}^N P(X_k \mid X_1, \dots, X_{k-1}) \\
# &= \prod_{k=1}^N P(X_k \mid \Parents_\G(X_k))
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Factorization Theorem: Consequences
#
# We just proved that for any $P$ satisfying $\G$,
# $$
# P(X_1, \dots, X_N) = \prod_{k=1}^N P(X_k \mid \Parents_\G(X_k))
# $$
#
# It suffices to store **conditional probability tables** $P(X_k | \Parents_\G(X_k))$!
# - Requires $O(N2^k)$ features if each node has $\leq k$ parents
# - Substantially more compact than **JPTs** for $N$ large, $\G$ sparse
# - We can also specify that a CPD is Gaussian, Dirichlet, etc.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Fully Connected Graph
#
# A **fully connected graph** makes no independence assumptions.
# $$
# P(A,B,C) = P(A) P(B|A) P(C|A,B)
# $$
# + slideshow={"slide_type": "skip"}
@pgm_render
def pgm_fully_connected_a():
pgm = daft.PGM([4, 4], origin=[0, 0])
# nodes
pgm.add_node(daft.Node("a", r"$A$", 2, 3.5))
pgm.add_node(daft.Node("b", r"$B$", 1.3, 2.5))
pgm.add_node(daft.Node("c", r"$C$", 2.7, 2.5))
# add in the edges
pgm.add_edge("a", "b", head_length=0.08)
pgm.add_edge("a", "c", head_length=0.08)
pgm.add_edge("b", "c", head_length=0.08)
return pgm;
# + slideshow={"slide_type": "-"}
# %%capture
pgm_fully_connected_a("images/fully-connected-a.png")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Fully Connected Graph
#
# There are many possible fully connected graphs:
# $$
# \begin{align}
# P(A,B,C)
# &= P(A) P(B|A) P(C|A,B) \\
# &= P(B) P(C|B) P(A|B,C)
# \end{align}
# $$
# + slideshow={"slide_type": "skip"}
@pgm_render
def pgm_fully_connected_b():
pgm = daft.PGM([8, 4], origin=[0, 0])
# nodes
pgm.add_node(daft.Node("a1", r"$A$", 2, 3.5))
pgm.add_node(daft.Node("b1", r"$B$", 1.5, 2.8))
pgm.add_node(daft.Node("c1", r"$C$", 2.5, 2.8))
# add in the edges
pgm.add_edge("a1", "b1", head_length=0.08)
pgm.add_edge("a1", "c1", head_length=0.08)
pgm.add_edge("b1", "c1", head_length=0.08)
# nodes
pgm.add_node(daft.Node("a2", r"$A$", 4, 3.5))
pgm.add_node(daft.Node("b2", r"$B$", 3.5, 2.8))
pgm.add_node(daft.Node("c2", r"$C$", 4.5, 2.8))
# add in the edges
pgm.add_edge("b2", "c2", head_length=0.08)
pgm.add_edge("b2", "a2", head_length=0.08)
pgm.add_edge("c2", "a2", head_length=0.08)
return pgm;
# + slideshow={"slide_type": "-"}
# %%capture
pgm_fully_connected_b("images/fully-connected-b.png")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Bayesian Networks & Causality
#
# The fully-connected example brings up a crucial point:
#
# > Directed edges do **not** necessarily represent causality.
#
# Bayesian networks encode **independence assumptions** only.
# - This representation is not unique.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Markov Chain
#
# State at time $t$ depends only on state at time $t-1$.
# $$
# P(X_0, X_1, \dots, X_N) = P(X_0) \prod_{t=1}^N P(X_t \mid X_{t-1})
# $$
# + slideshow={"slide_type": "skip"}
@pgm_render
def pgm_markov_chain():
pgm = daft.PGM([6, 6], origin=[0, 0])
# Nodes
pgm.add_node(daft.Node("x1", r"$\mathbf{x}_1$", 2, 2.5))
pgm.add_node(daft.Node("x2", r"$\mathbf{x}_2$", 3, 2.5))
pgm.add_node(daft.Node("ellipsis", r" . . . ", 3.7, 2.5, offset=(0, 0), plot_params={"ec" : "none"}))
pgm.add_node(daft.Node("ellipsis_end", r"", 3.7, 2.5, offset=(0, 0), plot_params={"ec" : "none"}))
pgm.add_node(daft.Node("xN", r"$\mathbf{x}_N$", 4.5, 2.5))
# Add in the edges.
pgm.add_edge("x1", "x2", head_length=0.08)
pgm.add_edge("x2", "ellipsis", head_length=0.08)
pgm.add_edge("ellipsis_end", "xN", head_length=0.08)
return pgm;
# + slideshow={"slide_type": "-"}
# %%capture
pgm_markov_chain("images/markov-chain.png")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Hidden Markov Model
#
# Noisy observations $X_k$ generated from hidden Markov chain $Y_k$.
# $$
# P(\vec{X}, \vec{Y}) = P(Y_1) P(X_1 \mid Y_1) \prod_{k=2}^N \left(P(Y_k \mid Y_{k-1}) P(X_k \mid Y_k)\right)
# $$
# + slideshow={"slide_type": "skip"}
@pgm_render
def pgm_hmm():
pgm = daft.PGM([7, 7], origin=[0, 0])
# Nodes
pgm.add_node(daft.Node("Y1", r"$Y_1$", 1, 3.5))
pgm.add_node(daft.Node("Y2", r"$Y_2$", 2, 3.5))
pgm.add_node(daft.Node("Y3", r"$\dots$", 3, 3.5, plot_params={'ec':'none'}))
pgm.add_node(daft.Node("Y4", r"$Y_N$", 4, 3.5))
pgm.add_node(daft.Node("x1", r"$X_1$", 1, 2.5, observed=True))
pgm.add_node(daft.Node("x2", r"$X_2$", 2, 2.5, observed=True))
pgm.add_node(daft.Node("x3", r"$\dots$", 3, 2.5, plot_params={'ec':'none'}))
pgm.add_node(daft.Node("x4", r"$X_N$", 4, 2.5, observed=True))
# Add in the edges.
pgm.add_edge("Y1", "Y2", head_length=0.08)
pgm.add_edge("Y2", "Y3", head_length=0.08)
pgm.add_edge("Y3", "Y4", head_length=0.08)
pgm.add_edge("Y1", "x1", head_length=0.08)
pgm.add_edge("Y2", "x2", head_length=0.08)
pgm.add_edge("Y4", "x4", head_length=0.08)
return pgm;
# + slideshow={"slide_type": "-"}
# %%capture
pgm_hmm("images/hmm.png")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Plate Notation
#
# We can represent (conditionally) iid variables using **plate notation**.
# + slideshow={"slide_type": "skip"}
@pgm_render
def pgm_plate_example():
pgm = daft.PGM([4,3], origin=[-2,0], node_unit=0.8, grid_unit=2.0);
# nodes
pgm.add_node(daft.Node("lambda", r"$\lambda$", -0.25, 2));
pgm.add_node(daft.Node("t1", r"$\theta_1$", -1, 1.3));
pgm.add_node(daft.Node("t2", r"$\theta_2$", -0.5, 1.3));
pgm.add_node(daft.Node("dots1", r"$\cdots$", 0, 1.3, plot_params={ 'ec' : 'none' }));
pgm.add_node(daft.Node("tN", r"$\theta_N$", 0.5, 1.3));
pgm.add_node(daft.Node("x1", r"$X_1$", -1, 0.6));
pgm.add_node(daft.Node("x2", r"$X_2$", -0.5, 0.6));
pgm.add_node(daft.Node("dots2", r"$\cdots$", 0, 0.6, plot_params={ 'ec' : 'none' }));
pgm.add_node(daft.Node("xN", r"$X_N$", 0.5, 0.6));
pgm.add_node(daft.Node("LAMBDA", r"$\lambda$", 1.5, 2));
pgm.add_node(daft.Node("THETA", r"$\theta_k$", 1.5,1.3));
pgm.add_node(daft.Node("XX", r"$X_k$", 1.5,0.6));
# edges
pgm.add_edge("lambda", "t1", head_length=0.08);
pgm.add_edge("lambda", "t2", head_length=0.08);
pgm.add_edge("lambda", "tN", head_length=0.08);
pgm.add_edge("t1", "x1", head_length=0.08);
pgm.add_edge("t2", "x2", head_length=0.08);
pgm.add_edge("tN", "xN", head_length=0.08);
pgm.add_edge("LAMBDA", "THETA", head_length=0.08);
pgm.add_edge("THETA", "XX", head_length=0.08);
pgm.add_plate(daft.Plate([1.1,0.4,0.8,1.2], label=r"$\qquad\quad\; K$",
shift=-0.1))
return pgm;
# + slideshow={"slide_type": "-"}
# %%capture
pgm_plate_example("images/plate-example.png")
| lecture15_exp_families_bayesian_networks/lecture15_exp_families_bayesian_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + tags=["parameters"]
"""
Update Parameters Here
"""
COLLECTION_NAME = "MekaVerse"
CONTRACT = "0x9a534628b4062e123ce7ee2222ec20b86e16ca8f"
LAST_N_EVENTS = 150
GRIFTER_ADDRESS = "" # optional overlay of grifter sales on map (grifter sales must have occured in last 'LAST_N_EVENTS' sales)
# +
"""
@author: mdigi14
"""
import datetime
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from honestnft_utils import opensea
from honestnft_utils import constants
from honestnft_utils import config
RARITY_DB = pd.read_csv(f"{config.RARITY_FOLDER}/{COLLECTION_NAME}_raritytools.csv")
sales = []
"""
Plot params
"""
plt.rcParams.update({"figure.facecolor": "white", "savefig.facecolor": "white"})
"""
Helper Functions
"""
def get_opensea_data(contract, continuous):
data = opensea.get_opensea_events(
contract_address=contract,
event_type="successful",
continuous=continuous,
)
return data
# +
"""
Generate Plot
"""
if LAST_N_EVENTS <= constants.OPENSEA_MAX_LIMIT:
events = get_opensea_data(CONTRACT, continuous=False)
else:
events = get_opensea_data(CONTRACT, continuous=True)
events = events[:LAST_N_EVENTS]
for event in events:
try:
token_id = int(event["asset"]["token_id"])
sale = dict()
sale["TOKEN_ID"] = token_id
sale["USER"] = event["transaction"]["from_account"]["address"]
sale["SELLER"] = event["seller"]["address"]
sale["DATE"] = event["created_date"]
sale["RANK"] = int(RARITY_DB[RARITY_DB["TOKEN_ID"] == token_id]["Rank"])
sale["PRICE"] = float(event["total_price"]) / constants.ETHER_UNITS
except:
continue
sales.append(sale)
df = pd.DataFrame(sales)
df = df[df["RANK"].notna()]
df.to_csv(f"{config.ROOT_DATA_FOLDER}/recent_sales.csv")
X = df["RANK"].values.reshape(-1, 1)
Y = df["PRICE"].values.reshape(-1, 1)
linear_regressor = LinearRegression()
linear_regressor.fit(X, Y)
Y_pred = linear_regressor.predict(X)
df = df.sort_values(by="RANK")
ax = df.plot.scatter(
x="RANK",
y="PRICE",
grid=True,
alpha=0.5,
title=COLLECTION_NAME,
figsize=(14, 7),
)
if GRIFTER_ADDRESS != "":
GRIFTER_DB = df[df["SELLER"] == GRIFTER_ADDRESS]
ranks = GRIFTER_DB["RANK"]
prices = GRIFTER_DB["PRICE"]
plt.scatter(x=ranks, y=prices, color="black", s=25)
plt.plot(X, Y_pred, color="red")
plt.xlabel("Rarity Rank (lower rank is better)")
plt.ylabel("Price (Ether)")
plt.title(
f"{COLLECTION_NAME} - Last {LAST_N_EVENTS} Sales (Before {datetime.datetime.now(datetime.timezone.utc).strftime('%Y-%m-%d %H:%M:%S %Z')})"
)
plt.savefig(f"{config.FIGURES_FOLDER}/{COLLECTION_NAME}_price_vs_rank.png")
plt.show()
| fair_drop/rarity_vs_lastprice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
# # Expressing Quantitative Relationships with Charts
#
# We use charts to express quantitative relationships in our data (or find them!) by giving shape to those relationships. Surprisingly, there's only eight (8) of those.
#
#
# 1. Time Series
# 2. Ranking
# 3. Part/Whole
# 4. Deviation
# 5. Distribution
# 6. Correlation
# 7. Geospatial
# 8. Nominal
#
# Note that these relationships are essentially *messages* or *stories* we want to convey about our data. We might want to tell several stories about a particular variable. This implies that you want *one chart per message*. Do not make charts tell more than one story. This is the problem with the Playfair chart. It is telling a Time Series story about the Exports and Imports of England and a Difference story about the Trade Balance. This should be two charts.
#
# Many of these concepts will be clearer when we start Exploratory Data Analysis. This section is mostly to plant seeds for later. We'll take up these different approaches when we start EDA.
#
# For each of these basic relationships we're going to look at the concepts we use around them and talk about some chart designs for each one. For each, we will consider the following options:
#
# 1. (horizontal|vertical) bars
# 2. (horizontal|vertical) dots
# 2. points
# 3. lines
# 4. points and lines
#
# These are combinations of the basic building blocks. We're going to look at the Nominal relationship first.
# ## Nominal
#
# The concepts we normally associate with Nominal relationships include:
#
# * categories
#
# Generally, the following options are good for Nominal relationships:
#
# 1. horizontal bars|dots
# 2. vertical bars|dots
# 3. points
#
# Sometimes horizontal bars are preferred to vertical bars if the labels are overly long. When plotting multiple Nominal relationships we will often add color or plot "small multiples" (to be discussed later). Do not add color to a single nominal relationship.
# ## Time Series
#
# The concepts we normally associated with Time Series include:
#
# * change
# * rise
# * increase
# * fluctuate
# * grow
# * decline
# * decrease
# * trend
#
# For Time Series data, you can use:
#
# 1. vertical bars
# 2. points and lines
#
# You can also use *boxes* which we have not yet shown. Boxes do not start at the x-axis as bars do. Instead they start at some minimum value and end at some maximum value. Boxes are good for showing low and high values for a particular observation (say, a stock market price per day).
#
# If there is an obvious continuity in the data, then it is best to use points *and* lines. If the continuity is broken, you may need to use just points or a combination of points and lines. Sometimes it's better to use vertical bars.
#
# *Never use horizontal bars or dots for Time Series*
# ## Ranking
#
# The concepts we normally associated with Ranking include:
#
# * larger than
# * smaller than
# * equal to
# * greater than
# * less than
#
# Generally speaking bar charts are the best chart for Rankings. You can either sort the data from highest to lowest (to emphasize the lowest value and ascending values) or lowest to highest (to emphasize the largest value and descending values).
# ## Part/Whole
#
# The concepts we normally associated with Part/Whole include:
#
# * rate or rate of total
# * percent or percentage of total
# * share
# * accounts for *x* percent
#
# We've already ruled out Pie Charts for showing Part/Whole relationships. Noting our problems with comparing non-aligned positions, we're going to suggest bar charts as the best way to represent Part/Whole relationships, either vertical or horizontal.
# ## Deviation
#
# The concepts we normally associated with Deviation include:
#
# * plus or minus
# * variance
# * difference
# * relative to
#
# Deviation stories should always be told *as the deviation* and not simply make the viewer calculate the deviation in their head.
#
# The effective displays for deviations are:
#
# * bars (either horizontal or vertical)
# * lines (when the deviations are from a Time Series)
# ## Distribution
#
# The concepts we normally associated with Distribution include:
#
# * frequency
# * distribution
# * range
# * concentration
# * normal curve/distribution (and others)
#
# We will have a lot to say about distributions later, in the EDA module. In general, we can use:
#
# * vertical bars (as a *histogram*)
# * lines (as a *frequency polygon*)
# * dots
# * boxes (as a *box and whiskers* plot)
#
# as well as combinations of the above to effectively display distributions.
# ## Correlation
#
# The concepts we normally associated with Correlation include:
#
# * increases with
# * decreases with
# * changes with
# * varies with
# * caused by
# * affected by
# * follows
#
# The canonical representation of some kind of correlation is to plot the numerical values on a scatter plot (XY-plot) and insert either a linear or LOESS trend line. Another approach would be to plot two side by side horizontal bar charts, called a *table lens*.
# ## Geospatial
#
# The concepts we normally associated with Geospatial relationships include:
#
# * geography
# * location
# * where
# * region
# * territory
# * country
# * state/province
# * count/borough/parish
# * city
#
# In this particular case, the forms (dots, lines, bars) are generally drawn or plotted on a map. When working with location data, you should consider whether or not the *location* is actually important the story. For example, do people already know where the 50 US states are in relation to each other? Is it really necessary to put this on a map rather than a table? Unless geography is part of the story, consider whether your visualization really requires a map.
#
# If you determine that a map is integral to the story, then possible solutions include:
#
# * points of varying size
# * points or areas of varying color *intensity*
# * color intensities applied directly to geographical areas (this was the option Cleveland disliked...notice that not everyone is in agreement!)
# * lines of varying thickness or color intensity.
#
# One type of chart that we haven't talked about is the *Heat Map*. A Heat Map can be considered a Geospatial chart in an abstract space where color is often used to represent a 3rd dimension.
| fundamentals_2018.9/visualization/relationships.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chemical Kinetics and Numerical Integration
#
# Here we will use methods of numerical integration to solve for the abundances of the H$_3^+$ isotopologues in the ion trap experiment from last week's notebook. After using integrated rate equations and curve fitting, we came up with this result:
#
# 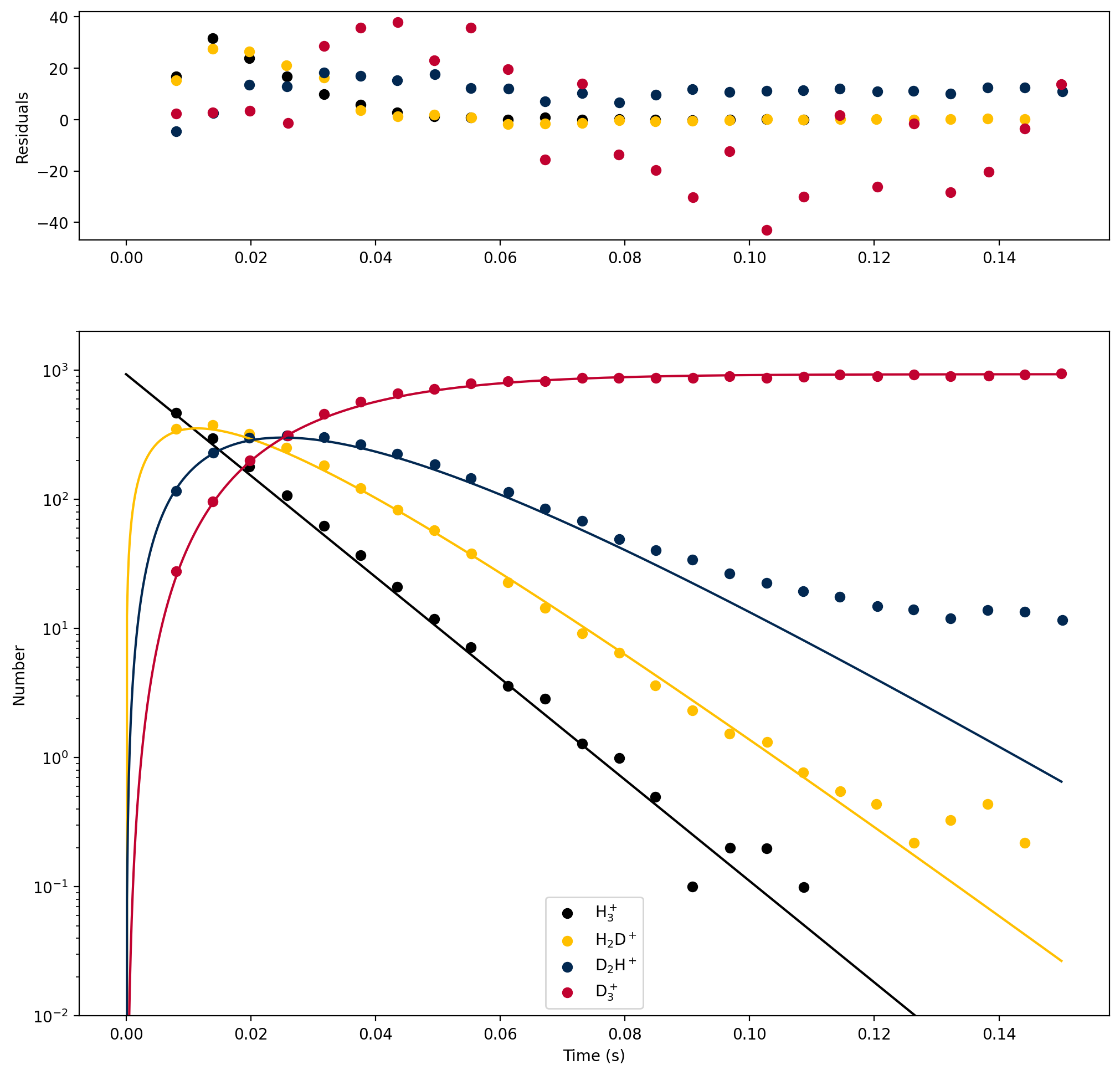
#
# The deviations, most notable in the D$_2$H$^+$ results, are because the reverse reactions were not included in our model. It would be very difficult to derive new rate equations, so we will use numerical methods instead.
#
# ## Forward Euler Method
#
# First, we will reimplement the exact same model as last time, but this time we will solve using the Forward Euler Method. First, load in the `deuteration.csv` file. It contains the same experimental data as last week, but the time field has been rounded and lined up so that all abundances for each molecule are given at the same time values. This will make comparisons with the numerical models easier down the road.
# +
import numpy as np
from matplotlib import pyplot as plt
import pandas as pd
df = pd.read_csv('deuteration.csv')
df
# -
# As a reminder, the model was defined by the equations:
#
# $$ \frac{\text{d}[\text{H}_3^+]}{\text{d}t} = -k_1[\text{H}_3^+] $$
#
# $$ \frac{\text{d}[\text{H}_2\text{D}^+]}{\text{d}t} = k_1[\text{H}_3^+] - k_2[\text{H}_2\text{D}^+] $$
#
# $$ \frac{\text{d}[\text{D}_2\text{H}^+]}{\text{d}t} = k_2[\text{H}_2\text{D}^+] - k_3[\text{D}_2\text{H}^+] $$
#
# $$ \frac{\text{d}[\text{H}_3^+]}{\text{d}t} = k_3[\text{D}_2\text{H}^+] $$
#
# We can express these in a simple form with the matrix equation:
#
# $$ \begin{bmatrix} \text{d}[\text{H}_3^+]/\text{d}t \\ \text{d}[\text{H}_2\text{D}^+]/\text{d}t \\ \text{d}[\text{D}_2\text{H}^+]/\text{d}t \\ \text{d}[\text{D}_3^+]/\text{d}t \end{bmatrix} = \begin{bmatrix} -k_1 & 0 & 0 & 0 \\ k_1 & -k_2 & 0 & 0 \\ 0 & k_2 & -k_3 & 0 \\ 0 & 0 & k_3 & 0 \end{bmatrix} \begin{bmatrix}[\text{H}_3^+] \\ [\text{H}_2\text{D}^+] \\ [\text{D}_2\text{H}^+] \\ [\text{D}_3^+] \end{bmatrix} $$
#
# Then, taking a time step $\Delta t$, we can compute new concentrations:
#
# $$ \begin{bmatrix}[\text{H}_3^+] \\ [\text{H}_2\text{D}^+] \\ [\text{D}_2\text{H}^+] \\ [\text{D}_3^+] \end{bmatrix}_{\,i+1} = \begin{bmatrix}[\text{H}_3^+] \\ [\text{H}_2\text{D}^+] \\ [\text{D}_2\text{H}^+] \\ [\text{D}_3^+] \end{bmatrix}_{\,i} + \begin{bmatrix} -k_1 & 0 & 0 & 0 \\ k_1 & -k_2 & 0 & 0 \\ 0 & k_2 & -k_3 & 0 \\ 0 & 0 & k_3 & 0 \end{bmatrix} \begin{bmatrix}[\text{H}_3^+] \\ [\text{H}_2\text{D}^+] \\ [\text{D}_2\text{H}^+] \\ [\text{D}_3^+] \end{bmatrix}_{\,i} \Delta t$$
#
# As of Python 3.5, matrix multiplication (and other types of dot products) can be done with the `@` operator. When used with `numpy.ndarray` objects, the [`numpy.matmul`](https://numpy.org/doc/stable/reference/generated/numpy.matmul.html) function is called. In our case, we will create a 4x4 matrix called `J` and a 1D array with 4 elements called `n` to store the abundances. When we call `J@n`, it multiplies each row of `J` by the 4 elements in `n`, and adds them up. Here we use the results from the curve fitting to ideally give us similar results as last time. We will set the step size `dt` to 0.1 ms, and take 1500 steps.
# +
#iniitialize rate constants
hd=6.3e10
k1=1.43e-9*hd
k2=1.33e-9*hd
k3=1.05e-9*hd
#H3+ at t=0 is 932, H2D+, D2H+, and D3+ start at 0.
n0 = np.array([932,0,0,0])
#initialize an empty 4x4 matrix, and plug in k values at the right places
J = np.zeros((4,4))
J[0,0] = -k1
J[1,1] = -k2
J[2,2] = -k3
J[1,0] = k1
J[2,1] = k2
J[3,2] = k3
#this array n will be updated with the new concentrations at each step. Initialize it at n0
n = n0
dt = 1e-4
steps = 1500
#this array will keep track of the values of n at each step
nt = np.zeros((steps+1,len(n0)))
nt[0] = n0
#take each steps, updating n at each one; store the results in the nt array
for i in range(0,steps):
n = n + J@n*dt
nt[i+1] = n
nt
# -
# Now we can plot the results and compare with the experimental data.
# +
fig,ax = plt.subplots()
t = np.linspace(0,150e-3,len(nt))
ax.scatter(df['time'],df['H3+'],color='#000000',label=r'H$_3^+$')
ax.scatter(df['time'],df['H2D+'],color='#ffbf00',label=r'H$_2$D$^+$')
ax.scatter(df['time'],df['D2H+'],color='#022851',label=r'D$_2$H$^+$')
ax.scatter(df['time'],df['D3+'],color='#c10230',label=r'D$_3^+$')
ax.set_xlabel("Time (s)")
ax.set_ylabel("Number")
lines = ax.plot(t,nt)
lines[0].set_color('#000000')
lines[1].set_color('#ffbf00')
lines[2].set_color('#022851')
lines[3].set_color('#c10230')
ax.set_yscale('log')
# -
# Note that the step size is a critical parameter! If we increase the step size too much, we can get some bad results.
# +
n = n0
dt = 5e-3
steps = round(.15/dt)+1
nt = np.zeros((steps+1,len(n0)))
nt[0] = n0
for i in range(0,steps):
n = n + J@n*dt
nt[i+1] = n
fig,ax = plt.subplots()
t = np.linspace(0,len(nt)*dt,len(nt))
ax.scatter(df['time'],df['H3+'],color='#000000',label=r'H$_3^+$')
ax.scatter(df['time'],df['H2D+'],color='#ffbf00',label=r'H$_2$D$^+$')
ax.scatter(df['time'],df['D2H+'],color='#022851',label=r'D$_2$H$^+$')
ax.scatter(df['time'],df['D3+'],color='#c10230',label=r'D$_3^+$')
ax.set_xlabel("Time (s)")
ax.set_ylabel("Number")
lines = ax.plot(t,nt)
lines[0].set_color('#000000')
lines[1].set_color('#ffbf00')
lines[2].set_color('#022851')
lines[3].set_color('#c10230')
ax.set_yscale('log')
# -
# ## Least Squares Fitting and Numerical Integration
#
# It is possible (though not very common) to implement least squares fitting together with the numerical integration in order to estimate the kinetics parameters. We'll walk through the process here. Last time we used `scipy.optimize.least_squares`, which required us to calculate the residuals vector between the model and the experimental data. When using integrated rate equations, this was straightforward because we could just plug in the time for each data point into the model and compute the model's prediction. With numerical integration; however, we do not have such a function!
#
# Instead, what we can do is save the model's outputs whenever the time matches the time at which an experimental data point is taken. If we choose time steps judiciously, we can make sure that we always sample the model at each time point needed. If we inspect the data frame, we can see that all of the time points are at a multiple of 0.1 ms.
df
# Therefore, a time step `dt` of 0.1 ms (or some integer factor smaller) will ensure that the model samples each time point we need to compare with the experimental data. The code below checks to see if `i` (the current time in units of `dt`) is in the array `tvals`, which is the time array converted to units of dt, and if so it stores the current model abundances in a list for later use. Importantly, this is chosen such that all of the time comparisons are between integers so that we don't have to worry about issues with floating point comparisons.
#
# At the end of the clode block, `nm` is a 2D numpy array where each row is a time point and each column is the abundance of one of the ions.
# +
n = n0
dt = 1e-4
steps = 1500
nmodel = []
tvals = df['time'].to_numpy()/dt
tvals = tvals.astype(int)
for i in range(0,steps+1):
n = n + J@n*dt
if i in tvals:
nmodel.append(n)
nm = np.array(nmodel)
nm
# -
tvals
# Now we'll plot the results. A quick side note here: we've been doing a lot of repetitive manual color changing. If you have a set of colors you want to consistently use, you can change matplotlib's default color cycling (see this [tutorial](https://matplotlib.org/tutorials/intermediate/color_cycle.html) for a quick example). Below I create a new `cycler` object that tells matplotlib to cycle between the 4 colors we have been using instead of its defaults. As the tutorial shows, you can either set the cycler on an `Axes` object like in the code below, which only affects that object, or you can apply the cycler to all subsequently created plots.
# +
from cycler import cycler
ucd_cycler = (cycler(color=['#000000','#ffbf00','#022851','#c10230','#266041','#8a532f']))
fig,ax = plt.subplots()
ax.set_prop_cycle(ucd_cycler)
ax.plot(df['time'],nm,'o')
# -
# Now let's turn that into a function that takes the kinetics parameters (`h30`, `k1`, `k2`, `k3`) as arguments. We also need to pass the time values at which the model should be sampled, the step size, and the number of steps.
# +
def runmodel(h30,k1,k2,k3,tvals,dt,steps):
n = np.asarray([h30,0,0,0])
nmodel = []
J = np.zeros((4,4))
J[0,0] = -k1
J[1,1] = -k2
J[2,2] = -k3
J[1,0] = k1
J[2,1] = k2
J[3,2] = k3
for i in range(0,steps+1):
n = n + J@n*dt
if i in tvals:
nmodel.append(n)
return(np.array(nmodel))
# -
# Test to make sure the `runmodel` function works as intended:
# +
tvals = df['time'].to_numpy()/dt
tvals = tvals.astype(int)
hd=6.3e10
k1=1.43e-9*hd
k2=1.33e-9*hd
k3=1.05e-9*hd
h30 = 932
runmodel(h30,k1,k2,k3,tvals,1e-4,1500)
# -
# To perform the `least_squares` optimization, we need to create a function that computes the residuals of the model. This function must have the signature `f(x,*args,**kwargs)` where `x` is an array containing the parameters that will be optimized (`h30`, `k1`, `k2`, and `k3`), `*args` contains any additional arguments that are needed, and `**kwargs` can contain any other information.
#
# Like last time, we'll use `**kwargs` to pass in the experimental data. `*args` will contain the `tvals`, `dt`, and `steps` parameters that need to be passed to `runmodel.` Ance we have the results of the model, we need to compute the residuals.
def total_fit(x,*args,**kwargs):
df = kwargs['df']
nm = runmodel(*x,*args)
#a naive algorithm using for loops; slow, but flexible!
# out = []
# for i,model in enumerate(nm):
# for j,mol in enumerate(['H3+','H2D+','D2H+','D3+']):
# n = df.at[i,mol]
# if np.isfinite(n):
# out.append(n-model[j])
# return out
#taking advantage of numpy's array routines: fast, but requires more work if anything changes
rh3 = df['H3+'] - nm[:,0]
rh3 = rh3[~np.isnan(rh3)] #remove NaNs... isnan returns an array of booleans, so we take the logical not and use it as a slice to extract only the finite values
rh2d = df['H2D+'] - nm[:,1]
rh2d = rh2d[~np.isnan(rh2d)]
#there are no NaNs in the experimental data for D2H+ or D3+
rd2h = df['D2H+'] - nm[:,2]
rd3 = df['D3+'] - nm[:,3]
#concatenate and return
return np.concatenate((rh3,rh2d,rd2h,rd3))
# Now we can use `least_squares` to compute optimal parameters, and we can see that we get almost exactly the same results as the integrated rate equation approach. Note, however, that there is no problem with us starting out with `k1` and `k2` being equal! There is no divide by 0 error with numerical integration like there was with the integrated rate equations.
# +
import scipy.optimize as opt
import numpy.linalg as la
data = {
'df' : df
}
tvals = df['time'].to_numpy()/dt
tvals = tvals.astype(int)
hd=6.3e10
result = opt.least_squares(total_fit,[950,1.3e-9*hd,1.3e-9*hd,1e-9*hd],
args=[tvals,1e-4,1500],kwargs=data,verbose=1)
pcov = la.inv(result.jac.T @ result.jac)
for i,x in enumerate(['[H3+]0','k1','k2','k3']):
den = hd
if i==0:
den = 1.
print(f'{x} = {result.x[i]/den:.2e} +/- {np.sqrt(pcov[i][i])/den:.2e}')
# -
# ## Integration with `scipy.integrate`
#
# Our manual implementation of the numerical integration uned the Forward Euler Method, whose total error is proportional to $(\Delta t)^{1}$. It is usually desirable to use a higher-order method to achieve either higher accuracy or obtain the same accuracy with fewer steps. The function we are going to explore is [`scipy.integrate.solve_ivp`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.integrate.solve_ivp.html), which is made to solve initial value problems.
# +
import scipy.integrate as spi
# spi.solve_ivp?
# -
# As we can see from the function description, we need to provide at least 3 arguments:
# - `fun` is a function that computes the vector of derivatives. Its function signature needs to be `f(t,y,*args)`. `t` is the current time, `y` is the current state array (in our case, the array containing the molecule abundances), and the remainder of the arguments can contain anything else needed to compute the derivatives (e.g., rate coefficients, etc)
# - `t_span` is a tuple that specifies the initial and final time for the integration
# - `y0` is a vector containing the initial conditions - the starting abundances for the molecules.
#
# In addition to those required parameters, there are three other optional arguments that are useful for us:
# - `method` selects which numerical integration method will be employed. The default, `'RK45'`, is the fourth-order Runge-Kutta method, but several others are available, including some implicit solvers that are important when problems are "stiff." A system of equations is stiff when the solutions are very sensitive to the step size even when the solution appears "smooth." Chemical kinetics problems are frequently stiff when there are some very slow reactions combined with others that are very fast, and you want to evaluate the system over a long time compared with the rate of the fast reactions. In the current example, all of the reactions have comparable rates, so we will stick with `'RK45'`, but often the `'Adams'` or `'Radau'` methods are more appropriate for kinetics problems.
# - `t_eval` is a list of times at which the model returns abundances. If this is None, the model only gives the results at the final time. If we pass an array of times, the results will contain the abundances at all of the time values specified in `t_eval` which fall within `t_span`
# - `dense_output` causes the solver to construct functions that interpolate between time steps. This allows you to (approximately) evaluate the model at any time, not just at the time steps that were used in the model.
#
# Note that nowhere do you need to specify the step size! All of the methods employ various algorithms to automatically determine the step size needed to bring the error down to a certain desired value. Some even include adaptive step sizes that can take smaller or larger steps depending on the magnitudes of the derivatives.
#
# Let's re-implement the same model, but this time perform the integration with `solve_ivp`. First we need to write a function that computes the derivative.
# function must take t and y as its first 2 arguments. Since our derivatives don't explicitly depend on t, that variable isn't used in the body of the function.
# to calculate the rates, we need the rate coefficients and abundances. The abundances are in y, so we need to pass the k values as arguments.
def calc_derivative(t,y,k1,k2,k3):
J = np.zeros((len(y),len(y)))
J[0,0] = -k1
J[1,1] = -k2
J[2,2] = -k3
J[1,0] = k1
J[2,1] = k2
J[3,2] = k3
return J@y
# With that, we can now use `solve_ivp` to compute the solution from 0 to 0.15 seconds. We'll use the default `RK45` integrator, and set the `dense_output` flag to allow us to generate a quasi-continuous model function. In addition, we'll pass our `df['time']` array to `t_eval` so that we have the exact model values at the experimental time points.
#
# Within the `result` object that is returned, we can access the dense solution with `result.sol`, which takes a time value as an argument. The solution values are in `result.y`, and the time points for each solution are in `result.t`. The plot that this cell creates shows both the dense output and the discrete solutions.
# +
hd=6.3e10
k1=1.43e-9*hd
k2=1.33e-9*hd
k3=1.05e-9*hd
h30 = 932
result = spi.solve_ivp(calc_derivative,(0,.15),y0=[h30,0,0,0],
t_eval=df['time'],method='RK45',
dense_output=True,args=(k1,k2,k3))
fig,ax = plt.subplots()
ax.set_prop_cycle(ucd_cycler)
t = np.linspace(0,160e-3,1000)
ax.plot(t,result.sol(t).T)
ax.plot(result.t,result.y.T,'o')
# -
# ## Extending the System
#
# We wish to add the reverse reactions to the system and also to explicitly model the reactions using the full second-order rates instead of the pseudo-first-order ones we have been using up until this points. So far our system has been explicitly hardcoded. This is fine for a small system like this, but if we start to have many more molecules and reactions, manually coding the rates is tedious and also error-prone.
#
# We will aim to improve the reliability and flexibility of the code by defining the model in terms of chemical reactions, and we will automatically generate the rate equations from the reactions themselves. First, let's create a list of molecules. We'll use a pandas dataframe for convenience, though we could implement this with lists or numpy arrays as well.
#
# We'll start by refactoring the existing pseudo-first-order system, and then show how we can easily convert to the full second order reaction network.
species = pd.DataFrame(['H3+','H2D+','D2H+','D3+'],columns=['name'])
species
# Now each molecule will be referred to by its index in this dataframe instead of by its name. We next need to define the chemical reactions that link these molecules together. We'll do this by creating a new class that contains the reactants, the products, and the rate coefficient. The class first extracts the unique reactants and products along with how many times that reactant/product appears, and stores those numbers as as numpy arrays, and uses We also make a `__str__` function for convenience that will print the reaction and its rate. We pass the `species` data into the constructor so that the reaction can get the names of the molecules.
class Reaction:
def __init__(self,species,reactants=[],products=[],k=0.0):
self.reactants, self.rcounts = np.unique(np.asarray(reactants),return_counts=True)
self.products, self.pcounts = np.unique(np.asarray(products),return_counts=True)
rnames = []
pnames = []
for r,c in zip(self.reactants,self.rcounts):
rnames.append(self.makename(species,c,r))
for p,c in zip(self.products,self.pcounts):
pnames.append(self.makename(species,c,p))
self.k = k
self.name = f'{" + ".join(rnames)} --> {" + ".join(pnames)}, k = {self.k:.2e}'
def __str__(self):
return self.name
def makename(self,species,c,n):
out = species.at[n,'name']
if c > 1:
out = f'{c}{out}'
return out
# To create a reaction, we call the `Reaction` constructor and give it the species list, then the list of the reactants' ID numbers, then the products' ID numbers, and then the rate coefficient. Since we're currently only considering the forward reactions and keeping \[HD\] constant, we can just include the ions.
r1 = Reaction(species,[0],[1],1.43e-9*hd)
r2 = Reaction(species,[1],[2],1.33e-9*hd)
r3 = Reaction(species,[2],[3],1.05e-9*hd)
reactions = pd.DataFrame([r1,r2,r3],columns=['reaction'])
reactions
# Note that we can make reactions that involve multiple of the same molecule. A silly example:
print(Reaction(species,[0,0,1],[3,3,2],1.))
# For computing the derivatives, we can use the definitions of the rate of an elementary reaction. For example, the elementary reaction A + B --> C + D has the following rates:
#
# $$ -\frac{\text{d}[A]}{\text{d}t} = -\frac{\text{d}[B]}{\text{d}t} = \frac{\text{d}[C]}{\text{d}t} = \frac{\text{d}[D]}{\text{d}t} = k[\text{A}][\text{B}] $$
#
# If the reaction has the form 2A --> C + D; this can also be written as A + A --> C + D, and the only difference is that the rate of change for \[A\] is twice as fast as the change for each product:
#
# $$ -\frac{1}{2}\frac{\text{d}[A]}{\text{d}t} = \frac{\text{d}[C]}{\text{d}t} = \frac{\text{d}[D]}{\text{d}t} = k[\text{A}]^2 = k[\text{A}][\text{A}] $$
#
# What this means is that for each molecule, we just need to loop over the reactions, and each time the molecule appears as a reactant, we subtract its rate coefficient times the product of the reactants, and each time it appears as a product, we add k times the product of the reactants. This will work even if the molecule appears twice in the same reaction (either as a reactant or a product, or even both!), becase we'll add the rate once for each time the molecule appears in the reaction.
#
# The code below is a new implementation of the derivative calculation that does this. It loops over the reactions, and for each reactant it subtracts the rate, and for each product it adds the rate.
def calc_derivative_2(t,y,rxns):
out = np.zeros_like(y)
for r in rxns['reaction']:
out[r.reactants] -= r.k*np.prod(np.power(y[r.reactants],r.rcounts))*r.rcounts
out[r.products] += r.k*np.prod(np.power(y[r.reactants],r.rcounts))*r.pcounts
return out
# This code takes advantage of numpy's advanced indexing capabilities. The lists of unique reactant and product IDs are used as indices to choose which concentrations to include in the rate calculations as well as which concentration derivatives to change. Note that the rates depend on the concentrations of the reactants, not the concentrations of the products. Below is some sample code showing how this works. The reaction is 3A + B --> 2C + D. Rate = k\[A\]^3\[B\] = (1e-4)(10)^3(3) = 0.3. So the concentration of A should change by -(3)(0.3) = -0.9, B should change by -0.3, C should change by +(2)(0.6) = 0.6, and D by +0.3
y = np.asarray([10.,3.,7.,2.])
out = np.zeros_like(y)
reactants = np.asarray([0,1,0,0])
products = np.asarray([2,2,3])
reactants, rcounts = np.unique(reactants,return_counts=True)
products, pcounts = np.unique(products,return_counts=True)
out[reactants] += -1e-4*np.prod(np.power(y[reactants],rcounts))*rcounts
out[products] += 1e-4*np.prod(np.power(y[reactants],rcounts))*pcounts
out
# And now, as a last sanity check, we should be able to plug in our reactions and initial conditions into the solver and get the same results.
r1 = Reaction(species,[0],[1],1.43e-9*hd)
r2 = Reaction(species,[1],[2],1.33e-9*hd)
r3 = Reaction(species,[2],[3],1.05e-9*hd)
reactions = pd.DataFrame([r1,r2,r3],columns=['reaction'])
result = spi.solve_ivp(calc_derivative_2,(0,.16),y0=[932,0,0,0],t_eval=df['time'],method='RK45',dense_output=True,args=[reactions])
fig,ax = plt.subplots()
ax.set_prop_cycle(ucd_cycler)
t = np.linspace(0,160e-3,1000)
ax.plot(t,result.sol(t).T)
ax.plot(result.t,result.y.T,'o')
# ## Second Order Kinetics and Reverse Reactions
#
# With our `Reaction` class and `calc_derivative_2` functions, it is now easy to include H2 and HD in the model, and do the second-order chemistry. The only addition is that we need to be careful about units. In the rate equations, the concentrations are given in molecules per cubic centimeter, so we need to divide the ion counts by the trap volume, which we do not exactly know. It may be listed in one of the many papers the group has published. However, the volume is likely on the order of 1 cubic centimeter. We can use that for now, and show that the final results in the end are not very sensitive to this number unless it's smaller than ~10-5, which seems physically impossible.
# +
species = pd.DataFrame(['H3+','H2D+',"D2H+","D3+",'HD','H2'],columns=['name'])
reactions = [Reaction(species,[0,4],[1,5],k=1.43e-9),
Reaction(species,[1,4],[2,5],k=1.33e-9),
Reaction(species,[2,4],[3,5],k=1.05e-9)]
reactions = pd.DataFrame(reactions,columns=['reaction'])
print(reactions)
volume = 1
result = spi.solve_ivp(calc_derivative_2,(0,.15),y0=[932/volume,0,0,0,6.3e10,0],t_eval=df['time'],method='RK45',dense_output=True,args=[reactions])
fig,ax = plt.subplots(figsize=(10,8))
ax.set_prop_cycle(ucd_cycler)
t = np.linspace(0,150e-3,1000)
lines = ax.plot(t,(result.sol(t)).T)
for l,n in zip(lines,species['name']):
l.set_label(n)
ax.scatter(df['time'],df['H3+']/volume,color='#000000')
ax.scatter(df['time'],df['H2D+']/volume,color='#ffbf00')
ax.scatter(df['time'],df['D2H+']/volume,color='#022851')
ax.scatter(df['time'],df['D3+']/volume,color='#c10230')
ax.set_xlim(0,150e-3)
ax.set_ylabel('Abundance (cm$^{-3}$)')
ax.set_yscale('log')
ax.legend()
# -
# We can see from this graph why the pseudo-first-order approximation is so good: if there are only ~1000 ions in a cubic centimeter, there are over 6e10 HD molecules. Even after all 1000 H$_3^+$ ions are converted to D$_3^+$, only 3000 of the HD molecules disappeared, which is negligible. However, eventually if we make the trap volume small enough, we can start to see an effect on the model. For instance, here we make the trap volume 1.5e-3, which means there are roughly as many H$_3^+$ ions as HD molecules. The chemistry is qualitatively different, yet we did not have to rederive any rate equations. Numerical integration is versatile.
# +
species = pd.DataFrame(['H3+','H2D+',"D2H+","D3+",'HD','H2'],columns=['name'])
reactions = [Reaction(species,[0,4],[1,5],k=1.43e-9),
Reaction(species,[1,4],[2,5],k=1.33e-9),
Reaction(species,[2,4],[3,5],k=1.05e-9)]
reactions = pd.DataFrame(reactions,columns=['reaction'])
print(reactions)
volume = 1.5e-8
result = spi.solve_ivp(calc_derivative_2,(0,.15),y0=[932/volume,0,0,0,6.3e10,0],t_eval=df['time'],method='RK45',dense_output=True,args=[reactions])
fig,ax = plt.subplots(figsize=(10,8))
ax.set_prop_cycle(ucd_cycler)
t = np.linspace(0,150e-3,1000)
lines = ax.plot(t,(result.sol(t)).T)
for l,n in zip(lines,species['name']):
l.set_label(n)
ax.scatter(df['time'],df['H3+']/volume,color='#000000')
ax.scatter(df['time'],df['H2D+']/volume,color='#ffbf00')
ax.scatter(df['time'],df['D2H+']/volume,color='#022851')
ax.scatter(df['time'],df['D3+']/volume,color='#c10230')
ax.set_xlim(0,150e-3)
ax.set_ylabel('Abundance (cm$^{-3}$)')
ax.set_yscale('log')
ax.legend()
# -
# Returning to more reasonable volumes, we can turn on the reverse reactions and see what happens. The paper says that the reverse reactions occur with rate coefficients that are of order 2e-10
# +
species = pd.DataFrame(['H3+','H2D+',"D2H+","D3+",'HD','H2'],columns=['name'])
reactions = [Reaction(species,[0,4],[1,5],k=1.43e-9),
Reaction(species,[1,4],[2,5],k=1.33e-9),
Reaction(species,[2,4],[3,5],k=1.05e-9),
Reaction(species,[1,5],[0,4],k=2e-10),
Reaction(species,[2,5],[1,4],k=2e-10),
Reaction(species,[3,5],[2,4],k=2e-10)]
reactions = pd.DataFrame(reactions,columns=['reaction'])
print(reactions)
volume = 1
result = spi.solve_ivp(calc_derivative_2,(0,.15),y0=[932/volume,0,0,0,6.3e10,0],t_eval=df['time'],method='RK45',dense_output=True,args=[reactions])
fig,ax = plt.subplots(figsize=(10,8))
ax.set_prop_cycle(ucd_cycler)
t = np.linspace(0,150e-3,1000)
lines = ax.plot(t,(result.sol(t)[0:4]).T)
for l,n in zip(lines,species['name'][0:4]):
l.set_label(n)
ax.scatter(df['time'],df['H3+']/volume,color='#000000')
ax.scatter(df['time'],df['H2D+']/volume,color='#ffbf00')
ax.scatter(df['time'],df['D2H+']/volume,color='#022851')
ax.scatter(df['time'],df['D3+']/volume,color='#c10230')
ax.set_xlim(0,150e-3)
ax.set_ylabel('Abundance (cm$^{-3}$)')
ax.set_yscale('log')
ax.legend()
# -
# It appears to make no difference! This is because in our model, the abundance of H$_2$ remains tiny. However, experimentally, the HD gas has a purity of only 97%. If we plug that in for the initial abundances, we can start to see something:
# +
species = pd.DataFrame(['H3+','H2D+',"D2H+","D3+",'HD','H2'],columns=['name'])
reactions = [Reaction(species,[0,4],[1,5],k=1.43e-9),
Reaction(species,[1,4],[2,5],k=1.33e-9),
Reaction(species,[2,4],[3,5],k=1.05e-9),
Reaction(species,[1,5],[0,4],k=2e-10),
Reaction(species,[2,5],[1,4],k=2e-10),
Reaction(species,[3,5],[2,4],k=2e-10)]
reactions = pd.DataFrame(reactions,columns=['reaction'])
print(reactions)
volume = 1
result = spi.solve_ivp(calc_derivative_2,(0,.15),y0=[930/volume,0,0,0,0.97*6.3e10,0.03*6.3e10],t_eval=df['time'],method='RK45',dense_output=True,args=[reactions])
fig,ax = plt.subplots(figsize=(10,8))
ax.set_prop_cycle(ucd_cycler)
t = np.linspace(0,150e-3,1000)
lines = ax.plot(t,(result.sol(t)[0:4]*volume).T)
ax.scatter(df['time'],df['H3+'],color='#000000',label=r'H$_3^+$')
ax.scatter(df['time'],df['H2D+'],color='#ffbf00',label=r'H$_2$D$^+$')
ax.scatter(df['time'],df['D2H+'],color='#022851',label=r'D$_2$H$^+$')
ax.scatter(df['time'],df['D3+'],color='#c10230',label=r'D$_3^+$')
ax.set_ylim(0.1,2000)
ax.set_xlim(0,150e-3)
ax.legend(loc='lower left',bbox_to_anchor=(0.1,0.3))
ax.set_xlabel("Time (s)")
ax.set_ylabel('Ion count')
ax.set_yscale('log')
# -
# After some manual adjustment of the rate coefficients, we can obtain good agreement with the experimental data. It is theoretically possible to improve this with `least_squares`, but there are now 6 rate coefficients and an extra parameter for the percentage of H$_2$ that would need to be optimized as well, which makes the process slow. Also, some parameters have only a tiny effect on the data, so a lot of care has to be taken to ensure the optimization works well.
# +
species = pd.DataFrame(['H3+','H2D+',"D2H+","D3+",'HD','H2'],columns=['name'])
reactions = [Reaction(species,[0,4],[1,5],k=1.4e-9),
Reaction(species,[1,4],[2,5],k=1.4e-9),
Reaction(species,[2,4],[3,5],k=1.1e-9),
Reaction(species,[1,5],[0,4],k=1e-10),
Reaction(species,[2,5],[1,4],k=2e-10),
Reaction(species,[3,5],[2,4],k=4e-10)]
reactions = pd.DataFrame(reactions,columns=['reaction'])
print(reactions)
volume = 1
result = spi.solve_ivp(calc_derivative_2,(0,.15),y0=[930/volume,0,0,0,0.97*6.3e10,0.03*6.3e10],t_eval=df['time'],method='RK45',dense_output=True,args=[reactions])
fig,ax = plt.subplots(figsize=(10,8))
ax.set_prop_cycle(ucd_cycler)
t = np.linspace(0,150e-3,1000)
lines = ax.plot(t,(result.sol(t)[0:4]*volume).T)
ax.scatter(df['time'],df['H3+'],color='#000000',label=r'H$_3^+$')
ax.scatter(df['time'],df['H2D+'],color='#ffbf00',label=r'H$_2$D$^+$')
ax.scatter(df['time'],df['D2H+'],color='#022851',label=r'D$_2$H$^+$')
ax.scatter(df['time'],df['D3+'],color='#c10230',label=r'D$_3^+$')
ax.set_ylim(0.1,2000)
ax.set_xlim(0,150e-3)
ax.legend(loc='lower left',bbox_to_anchor=(0.1,0.3))
ax.set_xlabel("Time (s)")
ax.set_ylabel('Ion count')
ax.set_yscale('log')
# -
# ## Using Implicit Solvers
#
# The implicit solvers that are good for stiff problems carry one additional complication: they require the Jacobian matrix in order to run efficiently. For a kinetics system with N molecules, the Jacobian matrix contains derivatives of the rates for each molecule with respect to every molecule:
#
# $$ J_{ij} = \frac{\partial}{\partial [\text{X}]_j} \text{Rate}_i $$
#
# For a reaction aA + bB --> cC + dD, we know the rates are:
#
# $$ \frac{\text{d}[\text{A}]}{\text{d}t} = -ak[\text{A}]^a[\text{B}]^b, \quad \frac{\text{d}[\text{B}]}{\text{d}t} = -bk[\text{A}]^a[\text{B}]^b, \quad \frac{\text{d}[\text{C}]}{\text{d}t} = ck[\text{A}]^a[\text{B}]^b, \quad \frac{\text{d}[\text{D}]}{\text{d}t} = -dk[\text{A}]^a[\text{B}]^b $$
#
# Taking the rate for A as an example, the derivatives with respect to each molecule are:
#
# $$ \frac{\partial}{\partial [\text{A}]} \text{Rate}_\text{A} = -aka[\text{A}]^{a-1}[\text{B}]^b, \quad \frac{\partial}{\partial [\text{B}]} \text{Rate}_\text{A} = -akb[\text{a}]^a[\text{B}]^{b-1}, \quad \frac{\partial}{\partial [\text{C}]} \text{Rate}_\text{A} = 0, \quad \frac{\partial}{\partial [\text{D}]} \text{Rate}_\text{A} = 0 $$
#
# If we apply this to each rate, the Jacobian matrix for this reaction is:
#
# $$ J = \begin{bmatrix} -aka[\text{A}]^{a-1}[\text{B}]^b & -akb[\text{A}]^a[\text{B}]^{b-1} & 0 & 0 \\ -bka[\text{A}]^{a-1}[\text{B}]^b & -bkb[\text{A}]^a[\text{B}]^{b-1} & 0 & 0 \\ cka[\text{A}]^{a-1}[\text{B}]^b & ckb[\text{A}]^a[\text{B}]^{b-1} & 0 & 0 \\ dka[\text{A}]^{a-1}[\text{B}]^b & dkb[\text{A}]^a[\text{B}]^{b-1} & 0 & 0\end{bmatrix} $$
#
# Assuming our system contains two other molecules E and F, the total contribution to the Jacobian matrix for this one reaction would have 0s in all of the extra rows and columns because the rate of this reaction does not depend on the concentrations of E or F:
#
# $$ J = \begin{bmatrix} -aka[\text{A}]^{a-1}[\text{B}]^b & -akb[\text{A}]^a[\text{B}]^{b-1} & 0 & 0 & 0 & 0 \\ -bka[\text{A}]^{a-1}[\text{B}]^b & -bkb[\text{A}]^a[\text{B}]^{b-1} & 0 & 0 & 0 & 0 \\ cka[\text{A}]^{a-1}[\text{B}]^b & ckb[\text{A}]^a[\text{B}]^{b-1} & 0 & 0 & 0 & 0 \\ dka[\text{A}]^{a-1}[\text{B}]^b & dkb[\text{A}]^a[\text{B}]^{b-1} & 0 & 0 & 0 & 0 \\ 0 & 0 & 0 & 0 & 0 & 0 \\ 0 & 0 & 0 & 0 & 0 & 0 \end{bmatrix} $$
#
# Then we can repeat the process for each reaction in the system, just adding to the appropriate elements of the Jacobian matrix. We can provide a function to calculate the Jacobian whose signature is `f(t,y,*args)` just like for `calc_derivative_2`.
# +
def calc_jacobian(t,y,rxns):
J = np.zeros((y.size,y.size)) #create an empty NxN matrix, where N = number of molecules in the system
for r in rxns['reaction']:
#loop over reactants; each loop computes one column of the Jacobian matrix
for i,(rc,ex) in enumerate(zip(r.reactants,r.rcounts)):
out = np.zeros(y.size)
#when we compute df/di, the power of reactant i is reduced by 1. So subtract 1 from the reactant counts at the ith position
#However, we don't want to modify the reaction itself, so make a copy of rcounts
ords = np.copy(r.rcounts)
ords[i] -= 1
#calculate the base rate = k * count * product (concentrations raised to correct powers)
rate = r.k*ex*np.prod(np.power(y[r.reactants],ords))
#rectants decrease by reactant count * base rate; products increase by product count * base rate
out[r.reactants] -= r.rcounts*rate
out[r.products] += r.pcounts*rate
#add to the correct column of the Jacobian matrix for this reactant
J[:,rc] += out
return J
#play around with the reaction definition to ensure jacobian is calculated correctly, using formulas above
r = Reaction(species,[0,1,1],[2,2,3],2.)
y = np.asarray([10.,20.,30.,40.])
calc_jacobian(0,y,pd.DataFrame([r],columns=['reaction']))
# -
# For large systems of reactions, we can also define the jacobian's sparsity structure. This is a matrix that has 1s in the positions where the Jacobian may be nonzero, and 0s where the Jacobian is always 0. The algorithms in the `solve_ivp` function can use that information to speed up the calculations because it can reduce the number of calculations it needs to perform. When the reaction network is large, there may be only a few reactions linked to some molecules, and the rows/columns corresponding to that element may contain many 0s. The sparsity structure depends only on the reaction network, not the state of the system, so we can precalculate it before running `solve_ivp`. For completeness, we'll do it here.
# +
def compute_sparsity(species,rxns):
out = np.zeros((species.size,species.size))
for rxn in rxns['reaction']:
for r in rxn.reactants:
out[r,rxn.reactants] = 1
for p in rxn.products:
out[p,rxn.products] = 1
out[r,p] = 1
return out
species = pd.DataFrame(['H3+','H2D+',"D2H+","D3+",'HD','H2'],columns=['name'])
reactions = [Reaction(species,[0,4],[1,5],k=1.4e-9),
Reaction(species,[1,4],[2,5],k=1.4e-9),
Reaction(species,[2,4],[3,5],k=1.1e-9),
Reaction(species,[1,5],[0,4],k=1e-10),
Reaction(species,[2,5],[1,4],k=2e-10),
Reaction(species,[3,5],[2,4],k=4e-10)]
reactions = pd.DataFrame(reactions,columns=['reaction'])
print(reactions)
compute_sparsity(species,reactions)
# -
# Of course, there are very few 0s in this matrix. H$_3^+$ is not directly linked to D$_2$H$^+$ or D$_3^+$, and H$_2$D$^+$ is not linked to D$_3^+$, but otherwise each molecule is connected by at least 1 reaction. Now that we have the Jacobian and the sparsity structure, we can use one of the implicit solvers. (Strictly speaking, it is possible to use an implicit solver without the Jacobian matrix, in which case the Jacobian can be estimated by finite differences. However, doing so is extremely slow and introduces additional error, so it should be avoided).
# +
species = pd.DataFrame(['H3+','H2D+',"D2H+","D3+",'HD','H2'],columns=['name'])
reactions = [Reaction(species,[0,4],[1,5],k=1.4e-9),
Reaction(species,[1,4],[2,5],k=1.4e-9),
Reaction(species,[2,4],[3,5],k=1.1e-9),
Reaction(species,[1,5],[0,4],k=1e-10),
Reaction(species,[2,5],[1,4],k=2e-10),
Reaction(species,[3,5],[2,4],k=4e-10)]
reactions = pd.DataFrame(reactions,columns=['reaction'])
volume = 1.
print(reactions)
sparse = compute_sparsity(species,reactions)
result = spi.solve_ivp(calc_derivative_2,(0,.15),y0=[930/volume,0,0,0,0.97*6.3e10,0.03*6.3e10],t_eval=df['time'],
method='Radau',dense_output=True,args=[reactions],jac=calc_jacobian,jac_sparsity=sparse)
fig,ax = plt.subplots(figsize=(10,8))
ax.set_prop_cycle(ucd_cycler)
t = np.linspace(0,150e-3,1000)
lines = ax.plot(t,(result.sol(t)[0:4]*volume).T)
ax.scatter(df['time'],df['H3+'],color='#000000',label=r'H$_3^+$')
ax.scatter(df['time'],df['H2D+'],color='#ffbf00',label=r'H$_2$D$^+$')
ax.scatter(df['time'],df['D2H+'],color='#022851',label=r'D$_2$H$^+$')
ax.scatter(df['time'],df['D3+'],color='#c10230',label=r'D$_3^+$')
ax.set_ylim(0.1,2000)
ax.set_xlim(0,150e-3)
ax.legend(loc='lower left',bbox_to_anchor=(0.1,0.3),framealpha=0.)
ax.set_xlabel("Time (s)")
ax.set_ylabel('Ion count')
ax.set_yscale('log')
# -
| _notebooks/week5/odes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:xtar-dev]
# language: python
# name: conda-env-xtar-dev-py
# ---
import xarray as xr
from pathlib import Path
import glob
# ## Create Synthetic Data from Xarray
#
# We will pull out some data from the Xarray tutorial and replicate this data `n_files` times in a new data directory.
def create_synthetic_data(n_files=10, data_dir='data/air'):
"""
Create netCDF files
Parameters
---------
n_files : int
Total number of files that will be created
data_dir : str
Directory to use when creating the netCDF files
Returns
-------
filenames : list
List of filepaths for the created netCDF files
"""
ds = xr.tutorial.open_dataset('air_temperature')
data_dir = Path(data_dir)
data_dir.mkdir(parents=True, exist_ok=True)
filenames = [f'{data_dir}/air_temperature_{i}.nc' for i in range(n_files)]
for filename in filenames:
ds.to_netcdf(filename)
return filenames
filepaths = create_synthetic_data()
filepaths
# Let's create a tarbar for the created netCDF files
# !tar Pcvf data/air_dataset.tar data/air
# ## ratarmount
#
# We will use [ratarmount](https://github.com/mxmlnkn/ratarmount) to create an index file with file names, ownership, permission flags, and offset information to be stored at the TAR file's location. Once the index is created, ratarmount then offers a FUSE mount integration for easy access to the files.
#
# **NOTE:** Since `ratarmount` uses FUSE to mount the TAR file as a "filesystem in user space", you will need FUSE installed. On OSX, you will need to install [osxfuse](https://osxfuse.github.io/) *by hand*. On Linux, you can install `libfuse` using `conda`, if it is not already installed on your system.
#
# **NOTE:** If you have `libfuse` on your system and it is *older* than October 19, 2018 (i.e., < 3.3.0 for `fuse3` or < 2.9.9 for `fuse2`), and you have either Lustre or GPFS filesystems, `ratarmount` will fail with an error saying that your filesystem is unsupported. The solution is to upgrade to a newer version of `libfuse`.
#
# **NOTE:** If you install the `libfuse` Conda-Forge package on a Linux system, then you need to set the `FUSE_LIBRARY_PATH` environment variable to the location of the `libfuse.so` library file (e.g., `export FUSE_LIBRARY_PATH=/path/to/libfuse3.so`). If you do not do this, then `fusepy` (another dependency of `ratarmount`) will use the system `libfuse.so` file, which might be old.
#
# **NOTE:** Currently, the Conda-Forge version of `libfuse` does *not* build the `libfuse` utilities such as `fusermount3`. However, `fusepy` uses these utility functions under the hood when trying to mount the userspace filesystem. If you install the most recent version of `libfuse` and properly set the location of `libfuse` so that `fusepy` can find it (i.e., `FUSE_LIBRARY_PATH`), you will get an error the `fusermount3` cannot be found.
# %%time
# !ratarmount --recreate-index data/air_dataset.tar mounted_air_dataset
# %%time
mounted_dir = Path("mounted_air_dataset/data/air/")
list(mounted_dir.iterdir())
# For comparison, this is how long it takes to `list` the original data directory.
# %%time
mounted_dir = Path("data/air/")
list(mounted_dir.iterdir())
# **Substantially slower to list the directory contents (~2x slower)!!!**
# ## Benchmarks
from dask.distributed import performance_report, Client
client = Client()
client
# ### Original netCDF files
ds_orig = xr.open_mfdataset("data/air/*.nc", combine='nested', concat_dim='member_id')
ds_orig
ds_orig.air.data.visualize()
# ### Mounted netCDF files from the tar archive
ds_mntd = xr.open_mfdataset("mounted_air_dataset/data/air/*.nc", combine='nested', concat_dim='member_id')
ds_mntd
ds_mntd.air.data.visualize()
# ### Benchmark: Yearly Averages
# %%time
ds_orig.groupby('time.year').mean(['time', 'member_id']).compute()
# %%time
ds_mntd.groupby('time.year').mean(['time', 'member_id']).compute()
# ### Dask Performance Reports
with performance_report(filename="dask-perf-report-original.html"):
ds_orig.groupby('time.year').mean(['time', 'member_id']).compute()
with performance_report(filename="dask-perf-report-mounted.html"):
ds_mntd.groupby('time.year').mean(['time', 'member_id']).compute()
from IPython.display import HTML
display(HTML("dask-perf-report-original.html"))
display(HTML("dask-perf-report-mounted.html"))
| notebooks/tar-netcdf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://pythonista.mx)
# # Completado de elementos en objetos que contengan iterables
#
# Python permite insertar código que genere una serie de objetos mediante iteradores y condicionales. Este "autollenado" de elementos se conoce por su nombre en inglés "comprehension" y se puede apicar entre otros a objetos de tipo.
#
# * ```list```.
# * ```tuple```.
# * ```dict```.
# * ```set```.
# * ```forzenset```.
# * ```bytes```.
# * ```bytearray```.
#
# ## Completado de elementos en objetos de tipo _list_.
#
# La sintaxis es la siguente para un objeto de tipo _list_.
# ```
# [<expresión> for <nombre> in <iterable> if <expresión lógica>]
# ```
# El proceso de completado es el siguiente:
# * Realiza la iteración definida por la expresión _for_ .. *in*.
# * A cada elemento iterado le aplica la condición lógica.
# * Si la condición lógica se cumple, añade el resultado de la expresión aplicada al elemento.
#
# Es válido no incluir una expresión condicional.
# **Ejemplos:**
[x for x in 'Hola']
list('Hola')
[5 * x for x in range(1, 21) if x % 2 == 0]
[letra.upper() for letra in 'Parangaricutirimicuaro' if letra.lower() not in ['a', 'e', 'i', 'o', 'u']]
# Usando el último ejemplo, el código sin utilizar completado de elementos sería algo similar a lo siguiente:
lista = []
for letra in 'Parangaricutirimicuaro':
if letra.lower() not in ['a', 'e', 'i', 'o', 'u']:
lista.append(letra.upper())
print(lista)
# ### Completado con expresiones ternarias.
#
# Es posible utilizar expresiones ternarias para el completado de elementos con la siguiente sintaxis:
#
# ```
# [<expresión ternaria> for <nombre> in <iterable>]
# ```
# **Ejemplos:**
[x if x % 2 == 0 else 5 * x for x in range(1, 21)]
[letra.upper() if letra.lower() not in ['a', 'e', 'i', 'o', 'u'] else letra.lower() for letra in 'chapultepec']
# ## Completado de elementos en objetos de tipo _tuple_.
#
# Cuando se utiliza el completado en un objeto tipo *tuple*, el objeto resultante es un generador.
#
# **Ejemplo:**
#
generador = (5 * x for x in range(1, 21) if x % 2 == 0)
type(generador)
print(generador)
for item in generador:
print(item)
generador = (5 * x for x in range(1, 21) if x % 2 == 0)
list(generador)
# ## Completado de elementos en objetos de tipo _dict_.
#
# En este caso, lo más común es que el elemento iterable sea el identificador.
#
# Sintaxis:
# ```
# {<expresión> for <nombre> in iterable> if <expresión lógica>}
# ```
# **Ejemplo:**
#
# * Las siguientes celdas definirán una serie de identificadores en *campos*, los cuales serán utilizados como el objeto iterable en el completado de objetos tipo *dict*.
# * En cada iteración se ejecutará la función *input()* y el texto ingresado será asociado al identificador correspondiente.
campos = ('nombre', 'primer apellido', 'segundo apellido', 'correo')
{campo: input('Ingrese {}: '.format(campo)) for campo in campos}
registro = {campo: input('Ingrese {}: '.format(campo)) for campo in campos}
registro
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2019.</p>
| 25_completado_de_elementos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# # SuSiE Purity Plot
# + [markdown] kernel="SoS"
# This is mostly the same as `20180516_Purity_Plot.ipynb` but using `lm_less` simulation model instead of `simple_lm`. The difference is that now PVE is confined to 0.2.
# + kernel="SoS"
# %revisions -n 10 --source
# + kernel="SoS"
# %cd ~/GIT/github/mvarbvs/dsc
# + [markdown] kernel="SoS"
# Other differences include:
#
# 1. Increase number of genes to 100 and use 2 replicates per analysis
# 2. Fix a minor bug in residual variance simulation
# 3. Use $|r|$ for LD filtering, not $r^2$
# 4. **Add histogram of purity**
#
# Therefore I chose an LD cutoff of 0.5 for now (comparable to 0.25 when squared). But we argue that in practice it does not matter.
# + [markdown] kernel="SoS"
# ## Results
#
# ### Purity plots
#
# Purity plots updated to [here](http://shiny.stephenslab.uchicago.edu/gaow/purity_20180620). In particular we focus on this one with 2 causal, prior 0.1 for PVE 0.2, and `null_weight` set to 0.5.
# + kernel="SoS"
# %preview susie_comparison/purity_20180620/12.png
# + [markdown] kernel="SoS"
# ### Purity filter vs capture rate
# + kernel="SoS"
# %preview susie_comparison/ld_20180620.png
# + [markdown] kernel="SoS"
# ### Histogram of purity
# + kernel="SoS"
# %preview susie_comparison/hist_0722.png
# + [markdown] kernel="SoS"
# ## Run benchmark
# + kernel="Bash"
dsc susie.dsc --target susie_comparison
# + kernel="SoS"
[global]
parameter: cwd = path('~/GIT/github/mvarbvs/dsc')
parameter: outdir = 'susie_comparison'
parameter: name = '1008'
parameter: source = "liter_data"
parameter: dataset = 'lm_less'
parameter: susie = "fit_susie"
parameter: null_weight = [0.0,0.5,0.9,0.95]
parameter: maxL=10
ld_cutoff = 0.25
lfsr_cutoff = 0.05
# + [markdown] kernel="SoS"
# ## Plot by purity
# + kernel="SoS"
[purity_1, ld_1]
target = f"{source}.dataset {dataset} {dataset}.pve {dataset}.n_signal {susie}.maxL {susie}.null_weight {susie}.prior_var {susie}"
output: f'{cwd}/{outdir}/purity_{name}_{source}_{dataset}_{susie}.rds'
R: expand = '${ }', workdir = cwd
out = dscrutils::dscquery("${outdir}", target = "${target}")
saveRDS(out, ${_output:r})
# + kernel="SoS"
[purity_2, ld_2]
parameter: pve = [0.2]
parameter: L = [1,2,3,4,5]
parameter: prior = [0, 0.1, 0.4]
ld_col = 1
combos = len(pve) * len(L) * len(null_weight) * len(prior)
output_files = [f'{_input:d}/{x+1}.rds' for x in range(combos)]
input: for_each = ['pve', 'L', 'null_weight', 'prior'], concurrent = True
output: output_files[_index]
R: expand = '${ }', workdir = cwd
options(warn=2)
get_combined = function(sub, dirname, ld_col) {
out_files = sub[,c("${susie}.output.file", "${dataset}.output.file")]
combined = list(purity = NULL, lfsr = NULL, size = NULL,
captures = NULL, total_captures = NULL)
for (i in 1:nrow(out_files)) {
fit_file = paste0(dirname, out_files[i,1], '.rds')
dat = readRDS(fit_file)$fit
truth = dscrutils::read_dsc(paste0(dirname, out_files[i,2]))$data$true_coef
L = sub[i,"${dataset}.n_signal"]
for (r in 1:2) {
signals = which(truth[,r]!=0)
if (is.null(dat[[r]]$sets$cs)) next
purity_r = as.matrix(dat[[r]]$sets$purity)
dm = dim(purity_r)
if (dm[1] < ${maxL}) {
purity_r = rbind(purity_r, matrix(0, ${maxL}-dm[1], dm[2]))
}
if (is.null(combined$purity)) combined$purity = purity_r[,ld_col]
else combined$purity = cbind(combined$purity, purity_r[,ld_col])
#
if (is.null(combined$size)) combined$size = susieR:::n_in_CS(dat[[r]])
else combined$size = cbind(combined$size, susieR:::n_in_CS(dat[[r]]))
#
if (is.null(combined$lfsr)) combined$lfsr = susieR::susie_get_lfsr(dat[[r]])
else combined$lfsr = cbind(combined$lfsr, susieR::susie_get_lfsr(dat[[r]]))
#
capture_status = unlist(lapply(1:length(dat[[r]]$sets$cs), function(i) sum(dat[[r]]$sets$cs[[i]] %in% signals)))
if (length(capture_status) < ${maxL})
capture_status = c(capture_status, rep(L, ${maxL} - length(capture_status)))
if (is.null(combined$captures)) combined$captures = capture_status
else combined$captures = cbind(combined$captures, capture_status)
#
detected = colSums(do.call(rbind, lapply(1:length(dat[[r]]$sets$cs), function(i) signals %in% dat[[r]]$sets$cs[[i]])))
if (length(detected) < L) {
detected = c(detected, rep(0, L - length(detected)))
}
if (is.null(combined$total_captures)) combined$total_captures = detected
else combined$total_captures = combined$total_captures + detected
}
}
return(combined)
}
out = readRDS(${_input:r})
sub = out[which(out$${dataset}.pve == ${_pve} & out$${dataset}.n_signal == ${_L} & out$${susie}.null_weight == ${_null_weight} & out$${susie}.prior_var == ${_prior}),]
combined = get_combined(sub, "${outdir}/", ${ld_col})
write(paste(${_pve}, ${_L}, ${_prior}, ${_null_weight}, "${_output:n}.png", sep=','), file='${_output:n}.log')
saveRDS(combined, ${_output:r})
# + kernel="SoS"
[purity_3, ld_3]
input: group_by = 1, concurrent = True
output: f"{_input:n}.pkl"
bash: expand = True, workdir = cwd
dsc-io {_input} {_output}
# + kernel="SoS"
[purity_4]
input: group_by = 1, concurrent = True
output: f"{_input:n}.pdf"
python: expand = '${ }', workdir = cwd
import numpy as np
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
COLORS = ['#348ABD', '#7A68A6', '#A60628', '#467821', '#FF0000', '#188487', '#E2A233',
'#A9A9A9', '#000000', '#FF00FF', '#FFD700', '#ADFF2F', '#00FFFF']
color_mapper = np.vectorize(lambda x: dict([(i,j) for i,j in enumerate(COLORS)]).get(x))
def plot_purity(data, output, lfsr_cutoff = 0.05, verbose = False, delta_l = 2):
purity = np.array(data['purity'])
lfsr = np.array(data['lfsr'])
size = np.array(data['size'])
capture = np.array(data['captures'])
capture_summary = [f"Signal {idx+1} captured {int(item)}/{purity.shape[1]}" for idx, item in enumerate([data['total_captures']] if isinstance(data['total_captures'], np.float64) else data['total_captures'])]
n_causal = len(capture_summary)
idx = 0
plt.figure(figsize=(8, 8))
L = purity.shape[0]
if delta_l > 0:
L = min(n_causal + delta_l, L)
cols = 2
rows = L // cols + L % cols
position = range(1,L + 1)
insig = []
for x, y, z, c in zip(size, purity, lfsr, capture):
# exclude previously marked insignificant positions
exclude = [i for i, item in enumerate(x) if (x[i], y[i]) in insig]
z_sig = [i for i, zz in enumerate(z) if zz <= lfsr_cutoff and i not in exclude]
z_nsig = [i for i, zz in enumerate(z) if zz > lfsr_cutoff and i not in exclude]
colors = [4 if i == 0 else 0 for i in c]
plt.subplot(rows,cols,position[idx])
idx += 1
if len(z_sig):
label = f'L{idx}: lfsr<={lfsr_cutoff}'
plt.scatter(np.take(x, z_sig),
np.take(y, z_sig),
c = color_mapper(np.take(colors, z_sig)),
label = label, marker = '*')
if len(z_nsig):
label = f'L{idx}: lfsr>{lfsr_cutoff}'
plt.scatter(np.take(x, z_nsig),
np.take(y, z_nsig),
c = color_mapper(np.take(colors, z_nsig)),
label = label, marker = 'x')
# mask colored (insig) sets
insig.extend([(x[i], y[i]) for i, item in enumerate(colors) if item > 0 and i not in exclude])
insig = list(set(insig))
plt.legend(bbox_to_anchor=(0,1.02,1,0.2), loc="lower left",
mode="expand", borderaxespad=0, ncol=2, handletextpad=0.1)
plt.axhline(y=${ld_cutoff}, color = '#FF0000', alpha=0.5)
plt.xlabel("CS Size")
plt.ylabel("CS Purity")
if idx >= L:
break
plt.subplots_adjust(hspace=0.5/3*rows, wspace = 0.25)
if verbose:
plt.suptitle(f"95% CI set sizes vs min(abs(LD)) | LD filter ${ld_cutoff}\n{'; '.join(capture_summary)}")
plt.savefig(output, dpi=500, bbox_inches='tight')
plt.gca()
import pickle, os
data = pickle.load(open('${_input}', 'rb'))
try:
plot_purity(data, '${_output}', lfsr_cutoff = ${lfsr_cutoff})
os.system("convert -density 120 ${_output} ${_output:n}.png")
except Exception as e:
print(e)
os.system("touch ${_output}")
# + kernel="SoS"
[purity_5]
header = 'PVE,N_Causal,susie_prior,null_weight,output'
output: f'{cwd}/{outdir}/purity_{name}/index.html'
bash: expand = True, workdir = cwd
echo {header} > {_output:n}.csv
cat {str(_input).replace('.pdf', '.log')} >> {_output:n}.csv
cd {_output:d}
dsc-io {_output:an}.csv {_output:a}
# + [markdown] kernel="SoS"
# ## Plot capture rate by purity filter
# + kernel="SoS"
[ld_4]
output: f'{cwd}/{outdir}/ld_{name}.pkl'
python: expand = '${ }', workdir = cwd
import numpy as np
import pickle
lds = dict([(x, []) for x in np.linspace(0, 1, num=21) if x < 1])
for f in [${_input:r,}]:
data = pickle.load(open(f, 'rb'))
if data['purity'] is None:
# failed to generate this dataset
continue
for k in lds:
cs_capture_status = np.array(data['captures'][np.where(data['purity'] > k)]).ravel()
if len(cs_capture_status):
lds[k].append((f, sum(cs_capture_status > 0) / len(cs_capture_status)))
with open(${_output:r}, 'wb') as f:
pickle.dump(lds, f)
# + kernel="SoS"
[ld_5]
output: f'{_input:n}.png'
python: expand = '${ }', workdir = cwd
import pickle
import pandas as pd
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(rc={'figure.figsize':(8,4), "font.size":10, "axes.titlesize":10,"axes.labelsize":10}, style = "whitegrid")
fig, ax = plt.subplots(figsize=(8,4))
data = pickle.load(open('${outdir}/ld_${name}.pkl', 'rb'))
plot_data = dict([('LD cutoff', []), ('CS capture rate', [])])
idx = 0
for k in sorted(data.keys()):
idx += 1
if idx % 2:
continue
for item in data[k]:
plot_data['LD cutoff'].append(f'{k:.2f}')
plot_data['CS capture rate'].append(item[1])
plot_data = pd.DataFrame(plot_data)
sns.violinplot(ax=ax, x="LD cutoff", y="CS capture rate", data=plot_data, cut=0.5, inner="box")
sns.despine()
plt.axhline(0.95, color='red')
ax.get_figure().savefig("${_output:n}.pdf")
import os
os.system("convert -density 120 ${_output:n}.pdf ${_output}")
# + [markdown] kernel="SoS"
# ## Histogram of purity
# + kernel="SoS"
[purity_6]
use_null_weight = 0.5
output: f'{cwd}/{outdir}/hist_{name}.png'
python: expand = '${ }', workdir = cwd
import numpy as np
import pandas as pd
import pickle, os
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(rc={'figure.figsize':(4,4), "font.size":20, "axes.titlesize":20,"axes.labelsize":20}, style = "white")
data = pd.read_csv("${_input:n}.csv").query('null_weight == ${use_null_weight} and susie_prior == 0.1')
files = data['output'].tolist()
ncausal = data['N_Causal'].tolist()
pngs = []
for f,n in zip(files,ncausal):
hist_dat = pickle.load(open(f[:-3] + 'pkl', 'rb'))['purity']
fig, ax = plt.subplots(figsize=(4,4))
sns.distplot(np.ravel(np.array(hist_dat)), ax=ax, bins=20, kde=False, color='#800000')
sns.despine()
ax.set_title(f'{n} effect variable{"s" if n > 1 else ""}')
ax.get_figure().savefig(f'${_output:n}_{n}.pdf')
os.system(f"convert -density 120 ${_output:n}_{n}.pdf ${_output:n}_{n}.png")
pngs.append(f'${_output:n}_{n}.png')
import os
os.system(f'convert +append {" ".join(pngs)} ${_output}')
| src/20180620_Purity_Plot_Lite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dialogue System Pepper
# The code aims to give Pepper basic conversation abilities, this includes a speech recognition module, a conversational engine to formulate the answers and the speech synthesis.
# The dialogue is purpose-less, this means that no task is pursued except a normal and pleasant interaction (this is the objective).
# It can be used as a fall-back system also for a task oriented interaction, often the counterpart tends to ramble or to test the IQ of the robot asking general questions. <br>
#
# Author: <NAME> <br>
# Email: <EMAIL>.lirussi(at)studio.unibo.it
#
# ## Table of Contents:
# - [Requirements](#req)
# - [CONVERSATIONAL ENGINE](#conv)
# - [PEPPER PART](#pepper)
# - [SPEECH SYNTHESIS](#synth)
# - [SPEECH RECOGNITION](#rec)
# - [Closing](#close)
# ## Requirements <a class="anchor" id="req"></a>
# The Speech Synthesis works with
# * **Python 2.7** , because it uses
# * [Pepper API (NAOqi 2.5) ](https://developer.softbankrobotics.com/pepper-naoqi-25/naoqi-developer-guide/naoqi-apis)
#
#
# The Conversational Engine works with
# * **Java** (because no AIML-2.0 systems in Python 2 were found)
#
# The Speech Recognition module was built to be able to run ON Pepper computer (in the head) it's only dependencies are
# * **Python 2.7** , because it uses
# * [Pepper API (NAOqi 2.5) ](https://developer.softbankrobotics.com/pepper-naoqi-25/naoqi-developer-guide/naoqi-apis)
# * **numpy**
#
# All of them are pre-installed on Pepper, if you want to run on your computer just create an environment that has all them.
#It has been used python 2.7.18, the cell will give you your current verison.
import sys
print("Python version:")
print (sys.version)
# ## CONVERSATIONAL ENGINE <a class="anchor" id="conv"></a>
# There should be a "lib" folder with the program Ab.jar, the files retrieved from the engine are in another folder "bots/en/" <br>
# It starts a process to which it can be passes a string in input to generate a response.
#
# +
import subprocess
from subprocess import Popen, PIPE, STDOUT
pobj = subprocess.Popen(['java', '-jar', 'lib/Ab.jar', 'Main', 'bot=en'],
stdin =subprocess.PIPE,
stdout=subprocess.PIPE,
stderr=subprocess.PIPE)
# +
import subprocess as sp
from threading import Thread
from Queue import Queue,Empty
import time
def getabit(o,q):
for c in iter(lambda:o.read(1),b''):
q.put(c)
o.close()
def getdata(q):
r = b''
while True:
try:
c = q.get(False)
except Empty:
break
else:
r += c
return r
q = Queue()
t = Thread(target=getabit,args=(pobj.stdout,q))
t.daemon = True
t.start()
while True:
print('Sleep for 2 seconds...')
time.sleep(2)#to ensure that the data will be processed completely
print('Data received:' + getdata(q).decode())
if not t.isAlive():
break
#in_dat = input('Your data to input:')
pobj.stdin.write(b'hello\n')
#when human says nothing
#pobj.stdin.write(b'\n')
pobj.stdin.flush()
break
# -
print('DATA RECEIVED:\n' + getdata(q).decode())
# ### Process response-string
# this function processes the data that has been received: it retrieves just the string of the answer
def processResponse(raw):
response = raw.replace("\n", " ") # changes new-line with space
#response = response[7:-7] # cuts beginning and end
temp = response.partition('Robot:')[-1].rpartition('Human:')[0] #takes response between "Robot:" and "Human:"
if not temp:
return response
return temp
#test
classic_response = "Robot: Hi nice to see you! \nHuman: "
error_response = "[Error string lenght can vary] Robot: I don't have an answer for that. \nHuman: "
print '-----RAW:-----'
print error_response
print '-----PROCESSED:-----'
print processResponse(error_response)
# ## PEPPER PART <a class="anchor" id="pepper"></a>
################### Adjusting IP and ports ###########################
IP_number = "192.168.0.118" #this is local one, use the real robot ip
port_number = 9559 #this is local one, use the real robot port number
#IMPORTS
import naoqi
from naoqi import ALProxy
import qi
import os
import time
from random import randint
#SESSION OPENING
session = qi.Session()
try:
session.connect("tcp://" + IP_number + ":" + str(port_number))
except RuntimeError:
print ("Can't connect to Naoqi at ip \"" + args.ip + "\" on port " + str(args.port) +".\n"
"Please check your script arguments. Run with -h option for help.")
sys.exit(1)
# ### SPEECH SYNTHESIS <a class="anchor" id="synth"></a>
# The text-to-speech is the one integrated in the robot to keep the Pepper-voice and to use the gestures at the same time. <br>
# We need the animated-speech service, but the parameters can be set in the normal text-to-speech service, it will influence the animated one.
# Multiple voices are available, "naoenu" is the best, "paola" for a "litte bit" of italianity, it sounds good while gesticulating and reflects more the author of the code :)
#
#
# +
#ASKING A SERVICE from the session
#we are using the animated speech, to set the parameters we need to set them in the text to speech service
aup = session.service("ALAnimatedSpeech") #aup = ALProxy("ALAnimatedSpeech", IP_number, port_number)
tts = session.service("ALTextToSpeech")
#available voices
print( "voices available: "+str(tts.getAvailableVoices()) )
# +
#PARAMETERS
tts.setVoice("naoenu")
#tts.setParameter("speed", 100) #Acceptable range is [50 - 400]. 100 default.
#tts.setParameter("pitchShift", 1.1) #Acceptable range is [0.5 - 4]. 0 disables the effect. 1 default.
tts.setParameter("volume", 70)#[0 - 100] 70 is ok if robot volume is 60
#reset Speed
#tts.resetSpeed()
# +
#test string
string1="Hello, I am Pepper robot! The speech synthesis is working fine."
string2="Hello! ^start(animations/Stand/Gestures/Hey_1) Nice to meet you ^wait(animations/Stand/Gestures/Hey_1)"
string2="Hello. Look I can stop moving ^mode(disabled) and after I can resume moving ^mode(contextual), you see ?"
wake="^pCall(ALMotion.wakeUp()) Ok, I wake up."
aup.say(string1)
# -
# ### SPEECH RECOGNITION <a class="anchor" id="rec"></a>
# For this part it's mandatory to use a service to record audio on pepper and process it with another method, the integrated speech recognition il limited to a bunch of words. The code of the service will analyse the level of sound intensity and, based on the parameters in the code below, decide when start recording and when stopping. <br>
# NOTE: since Pepper cannot process the recognition, but just the amount of noise in the environment this is a [really challenging problem of turn-taking.](https://en.wikipedia.org/wiki/Turn-taking) <br>
# Be careful changing the parameters because it could happen that the audio file is stopped too early for a long pause in the speech, the audio file is stopped after the initial silence cause it has been detected that nobody is speaking.
# Nevertheless, the ideal thing is to minimize the parameters to reduce the amount of time to recognize the sentence of the person. <br>
# The service will send the audio file to Google speech recognition API and generate an event when it receives the response. <br><br>
# We create modules that subscribe to this event, the Base just writes the result recognized, the Dialogue one is a litte more complicated:
# When the result is received the module if there is nothing recognized forced the robot to ask multiple times to repeat, then it will just listen in loop. If the result is intelligible, it sends recognized string to the conversational engine, processes the response and it passes it to the speech synthesis. At the end it starts listening again.
#
# >**REMEMBER TO TURN ON THE RECOGNITION SERVICE** with a shell in the speech-recognition folder: <br>
# > use python 2.7 or activate the environment with: *conda activate python2* <br>
# > run the service: <br>
# > *python module_speechrecognition.py --pip (your robot IP)* <br>
#COMPUTER MICROPHONE?
'''
import speech_recognition as sr
with sr.Microphone() as source:
try:
r = sr.Recognizer()
audio = r.listen(source, timeout = 30)
catched = r.recognize_google(audio,key = None, language = "en-US", show_all = True)
print catched
except:
print("It didn't work")
'''
class BaseSpeechReceiverModule(naoqi.ALModule):
"""
Use this object to get call back from the ALMemory of the naoqi world.
Your callback needs to be a method with two parameter (variable name, value).
"""
def __init__( self, strModuleName ):
try:
naoqi.ALModule.__init__(self, strModuleName )
self.BIND_PYTHON( self.getName(),"callback" )
except BaseException, err:
print( "ERR: ReceiverModule: loading error: %s" % str(err) )
# __init__ - end
def __del__( self ):
print( "INF: ReceiverModule.__del__: cleaning everything" )
self.stop()
def start( self ):
memory = naoqi.ALProxy("ALMemory", IP_number, port_number)
memory.subscribeToEvent("SpeechRecognition", self.getName(), "processRemote")
print( "INF: ReceiverModule: started!" )
def stop( self ):
print( "INF: ReceiverModule: stopping..." )
memory = naoqi.ALProxy("ALMemory", IP_number, port_number)
memory.unsubscribe(self.getName())
print( "INF: ReceiverModule: stopped!" )
def version( self ):
return "1.1"
def processRemote(self, signalName, message):
# Do something with the received speech recognition result
print(message)
class DialogueSpeechReceiverModule(naoqi.ALModule):
"""
Use this object to get call back from the ALMemory of the naoqi world.
Your callback needs to be a method with two parameter (variable name, value).
"""
def __init__( self, strModuleName ):
self.misunderstandings=0
try:
naoqi.ALModule.__init__(self, strModuleName )
self.BIND_PYTHON( self.getName(),"callback" )
except BaseException, err:
print( "ERR: ReceiverModule: loading error: %s" % str(err) )
# __init__ - end
def __del__( self ):
print( "INF: ReceiverModule.__del__: cleaning everything" )
self.stop()
def start( self ):
memory = naoqi.ALProxy("ALMemory", IP_number, port_number)
memory.subscribeToEvent("SpeechRecognition", self.getName(), "processRemote")
print( "INF: ReceiverModule: started!" )
def stop( self ):
print( "INF: ReceiverModule: stopping..." )
memory = naoqi.ALProxy("ALMemory", IP_number, port_number)
memory.unsubscribe(self.getName())
print( "INF: ReceiverModule: stopped!" )
def version( self ):
return "2.0"
def processRemote(self, signalName, message):
if autodec:
#always disable to not detect its own speech
SpeechRecognition.disableAutoDetection()
#and stop if it was already recording another time
SpeechRecognition.pause()
# received speech recognition result
print("INPUT RECOGNIZED: \n"+message)
#computing answer
if message=='error':
self.misunderstandings +=1
if self.misunderstandings ==1:
answer="I didn't understand, can you repeat?"
elif self.misunderstandings ==0:
answer="Sorry I didn't get it, can you say it one more time?"
elif self.misunderstandings ==0:
answer="Today I'm having troubles uderstanding what you are saying, I'm sorry"
else:
answer=" "
print('ERROR, DEFAULT ANSWER:\n'+answer)
else:
self.misunderstandings = 0
#sending recognized input to conversational engine
pobj.stdin.write(b''+message+'\n')
pobj.stdin.flush()
#getting answer
time.sleep(1)#to ensure that the data will be processed completely
answer = getdata(q).decode()
answer = processResponse(answer)
print('DATA RECEIVED AS ANSWER:\n'+answer)
#text to speech the answer
aup.say(answer)
if autodec:
print("starting service speech-rec again")
SpeechRecognition.start()
print("autodec enabled")
SpeechRecognition.enableAutoDetection()
else:
#asking the Speech Recognition to LISTEN AGAIN
SpeechRecognition.startRecording()
# +
# We need this broker to be able to construct
# NAOqi modules and subscribe to other modules
# The broker must stay alive until the program exists
myBroker = naoqi.ALBroker("myBroker",
"0.0.0.0", # listen to anyone
0, # find a free port and use it
IP_number, # parent broker IP
port_number) # parent broker port
try:
p = ALProxy("DialogueSpeechReceiverModule", "192.168.0.118", 9559)
p.exit() # kill previous instance
except:
pass
# Reinstantiate module
# Warning: ReceiverModule must be a global variable
# The name given to the constructor must be the name of the
# variable
'''
global BaseSpeechReceiverModule
BaseSpeechReceiverModule = BaseSpeechReceiverModule("BaseSpeechReceiverModule")
BaseSpeechReceiverModule.start()
'''
global DialogueSpeechReceiverModule
DialogueSpeechReceiverModule = DialogueSpeechReceiverModule("DialogueSpeechReceiverModule")
DialogueSpeechReceiverModule.start()
SpeechRecognition = ALProxy("SpeechRecognition")
SpeechRecognition.start()
SpeechRecognition.calibrate()
#SpeechRecognition.setLanguage("de-de")
#autodetection
autodec=False #to know if we have to enable again after the robot speech
if autodec:
SpeechRecognition.enableAutoDetection()
print("waiting calibration to finish")
time.sleep(6)
SpeechRecognition.setAutoDetectionThreshold(20) #to avoid movement of the head to trigger the listening
#the human speech starts from 20, but the head movement sounds can reach 25, there is no perfect value
print("threshold updated successfully")
#NOTES for autodetection:
#1. for the autodetection the threshold should be high, or it recognizes
# the head movement as a sound high enough to start listening
# --> it will start to say that he does't understand
# --> it can be possible to deactivate the sententence if no words are recognized
# ----> but it will not react if something is not recognized
#2. the auto-detection should be deactivated when the robot speaks and activated again
# when the sentence is finished, or it will pick up his own speech and answer to himself
# /!\ IF THERE IS ERROR
# "Can't find service: SpeechRecognition"
# REMEMBER TO TURN ON THE SERVICE with a shell with python2:
# conda activate python2
# python module_speechrecognition.py --pip (your robot IP)
# "... object is not callable"
# execute again the cell of the module (es: the definition of the class 'DialogueSpeechReceiverModule' )
# -
SpeechRecognition.printInfo()
SpeechRecognition.setAutoDetectionThreshold(10)
SpeechRecognition.setLookaheadDuration(2)
#amount of seconds, before the threshold trigger, that will be included in the request
#default is 1
SpeechRecognition.setIdleReleaseTime(3)
#idle time (RMS below threshold) after which we stop recording
#default is 2
#NOTE: too short can cut the sentence in a pause between words
SpeechRecognition.setHoldTime(4)
#waits at least these sec to stop from the beginning
#default is 3
#NOTE: too short and while the person thinks what to say the recognition stops
#manual ask for start recording
SpeechRecognition.startRecording()
# ## Closing process <a class="anchor" id="close"></a>
#conversational engine closing
pobj.stdin.close()
pobj.terminate()
#speech recognition closing
SpeechRecognition.pause()
DialogueSpeechReceiverModule.stop()
myBroker.shutdown()
# TODO:
# bloccare la testa,
# mettere timer di risposta
| Dialogue-Pepper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 (''research'': conda)'
# language: python
# name: python3
# ---
# +
# For this coding problem, we are about to simulate a deep learning
# training process.
# In computer vision field, classification is a very important
# and traditional field. Assume that we have a some pseudo data input
# for a mlp (multi-layer perceptron) model.
# Q1: How should we arrange the sequence of fully-connected layer, activation
# layer, batch normalization layer? Why?
# x - relu - fc - bn
# x - relu - bn - fc
# x - bn - fc - relu
# x - bn - relu - fc
# x - fc - relu - bn
# x - fc - bn - relu
# Q2: What is the role for batch normalization playing for?
# Q3: Assume we need to define a 2-layer mlp.
# We have our pseudo data x in shape of [batch_size, input_size]
# We have our pseudo label y in shape of [batch_size, num_classes](one-hot vector)
# We need you to !!!only!!! use numpy to build a training pipeline,
# 1. Use Sigmoid as your activation function
# 2. Use cross entropy loss
# 3. No batch normalization
# 4. Write your own back propogation progess to update your weights of fc layers.
# 5. Using SGD optimizer algorithm
# 6. Don't forget to do softmax in final output
# 7. Finish the api we given to you
import numpy as np
# ============================== Question ==============================
class MyMLP():
def __init__(
self,
input_size: int,
hidden_size: int,
num_classes: int,
lr: float,
batch_size: int = 1,
):
self.input_size = input_size
self.num_classes = num_classes
self.batch_size = batch_size
self.lr = lr
self.w1 = np.random.randn(hidden_size, input_size)
self.w2 = np.random.randn(num_classes, hidden_size)
def generate_data(self):
x = np.random.randn(self.batch_size, self.input_size) # [bs, input_size]
y = np.random.randint(0, self.num_classes, (self.batch_size, )) # [bs,]
y = np.eye(self.num_classes)[y] # [bs, num_cls]
return x, y
def sigmoid(self, x: np.ndarray) -> np.ndarray:
# y = sigmoid(x)
return None
def partial_sigmoid(self, x: np.ndarray) -> np.ndarray:
# y' = d(sigmoid(x))/d(x)
return None
def softmax(self, x: np.ndarray) -> np.ndarray:
# x = [batch_size, dim] -> do softmax in column-wise dimension
return None
def forward(self, x: np.ndarray) -> np.ndarray:
# Two layer mlp
# x -> fc -> sigmoid -> fc -> softmax
# Notice that we use the formulation y = x W.T (without bias)
self.a1 = None# fc1 output
self.h1 = None# sigmoid output
self.a2 = None# fc2 output
self.out = None# softmax output
return None
def loss(self, y_hat: np.ndarray, y: np.ndarray) -> np.ndarray:
# yhat
# cross entropy loss
return None
def bp(self, x: np.ndarray, y:np.ndarray) -> np.ndarray:
# Update w1 and w2
self.dl_da2 = None # partial loss to partial a2
self.dl_dw2 = None# partial loss to partial w2
self.dl_dh1 = None# partial loss to partial h1
self.dl_da1 = None# partial loss to partial a1
self.d1_dw1 = None# partial loss to partial w1
# SGD Update w1 and w2:
self.w1 = None
self.w2 = None
return None
def matrix_differential_propogation(self, x: np.ndarray, y: np.ndarray) -> np.ndarray:
# Bonus
# Matrix-Matrix function derivatives
# Double check for gradient
return None
if __name__ == '__main__':
np.random.seed(2022)
mymlp = MyMLP(input_size=4, hidden_size=10, num_classes=3, lr=0.01, batch_size=1)
x, y = mymlp.generate_data()
# Step 1
# check your forward result:
out = mymlp.forward(x)
assert np.linalg.norm(out - np.array([[0.36235674, 0.40728367, 0.23035959]])) < 1e-7
# Step 2
# check your loss
loss = mymlp.loss(out, y)
assert np.linalg.norm(loss - np.array([[-1.01512607, -0. , -0., ]])) < 1e-7
w1, w2 = mymlp.bp(x, y)
# Step 2
# check your gradients:
assert np.linalg.norm(mymlp.dl_da2 - np.array([[-0.63764326, 0.40728367, 0.23035959]])) < 1e-7
assert np.linalg.norm(mymlp.dl_dw2 - np.array([[-0.40732181, -0.47931036, -0.15217421, -0.03860381, -0.09738246, -0.36582423,
-0.57306758, -0.36619898, -0.53017721, -0.32491802],
[ 0.2601698, 0.30615125, 0.09719866, 0.02465752, 0.06220137, 0.23366394,
0.36603706, 0.2339033, 0.33864158, 0.2075358 ],
[ 0.14715201, 0.17315911, 0.05497555, 0.01394629, 0.03518109, 0.13216029,
0.20703051, 0.13229568, 0.19153563, 0.11738222]])) < 1e-7
assert np.linalg.norm(mymlp.dl_dh1 - np.array([[-0.22072836, 0.54220196, 0.2838984, -0.24103344, -1.74668689, -0.65096623,
-0.33832413, -0.18948642, 0.76756801, 0.07541716]]))
assert np.linalg.norm(mymlp.dl_da1 - np.array([[-0.05093011, 0.10120303, 0.05158341, -0.01370905, -0.22601836, -0.15920446,
-0.03079302, -0.04632553, 0.1075607, 0.0188474 ]]))
assert np.linalg.norm(mymlp.dl_dw1 - np.array([[ 0.03771251, -0.04810758, -0.05592065, -0.02520829],
[-0.07493837, 0.09559438, 0.11111971, 0.05009129],
[-0.03819625, 0.04872466, 0.05663796, 0.02553164],
[ 0.01015122, -0.0129493, -0.01505237, -0.00678541],
[ 0.16736108, -0.21349247, -0.24816545, -0.11186968],
[ 0.11788702, -0.15038138, -0.17480459, -0.07879958],
[ 0.02280148, -0.02908648, -0.03381036, -0.01524126],
[ 0.03430292, -0.04375818, -0.05086487, -0.02292921],
[-0.07964608, 0.10159971, 0.11810036, 0.05323807],
[-0.01395604, 0.01780288, 0.02069421, 0.00932868]])) < 1e-7
# Step 3
# check your updated weight result
assert np.linalg.norm(w1 - np.array([[-0.00090502, -0.27442035, -0.13872636, 1.98493824],
[ 0.28285871, 0.75985271, 0.29987041, 0.53979636],
[ 0.37387925, 0.37732615, -0.09077957, -2.30619859],
[ 1.14265851, -1.53552479, -0.86360149, 1.01661279],
[ 1.03229027, -0.8223573, 0.02138651, -0.38222486],
[-0.30536435, 0.99879532, -0.12552579, -1.47509791],
[-1.94113434, 0.83393979, -0.56687978, 1.17463937],
[ 0.3187258, 0.19130801, 0.36977883, -0.10091857],
[-0.94101303, -1.40515771, 2.07946601, -0.12084862],
[ 0.75993144, 1.82725411, -0.66093403, -0.80789955]])) < 1e-7
assert np.linalg.norm(w2 - np.array([[ 0.89187333, -0.21265435, -0.93800277, 0.59992435, 2.22408652, 1.0036637,
1.15540522, -0.1519143, -1.64527408, -1.45186499],
[ 0.31815989, 0.80828812, -0.24177658, 0.16487451, -0.03412259, 0.08552794,
1.03048616, -1.06316559, -1.01697087, -0.42230754],
[ 0.93068173, 0.31559234, -0.943021, 0.32112784, -1.36990554, -0.21448964,
-0.11681657, 0.62104636, 0.55332894, -2.95860606]])) < 1e-7
dl_dw2 = mymlp.matrix_differential_propogation(x, y)
assert np.linalg.norm(dl_dw2 - np.array([[-0.40732181, -0.47931036, -0.15217421, -0.03860381, -0.09738246, -0.36582423,
-0.57306758, -0.36619898, -0.53017721, -0.32491802],
[ 0.2601698, 0.30615125, 0.09719866, 0.02465752, 0.06220137, 0.23366394,
0.36603706, 0.2339033, 0.33864158, 0.2075358 ],
[ 0.14715201, 0.17315911, 0.05497555, 0.01394629, 0.03518109, 0.13216029,
0.20703051, 0.13229568, 0.19153563, 0.11738222]])) < 1e-7
# -
# Some reference for matrix derivates:
# $$
# \begin{align}
# &d(X\pm Y) = dX \pm dY,d(XY)=d XY+X dY \\
# &d X\odot Y=d X \odot Y+X\odot dY \\
# &d(\sigma(X))=\sigma'(X)\odot dX \\
# &tr(AB) = tr(BA) \\
# &tr(A^T(B\odot C))=tr((A\odot B)^TC), A,B,C\in \mathbb R^{n\times n} \\
# &df=tr(\frac{\partial f}{\partial X}^T dX) \\
# \end{align}
# $$
#
# $$
# \begin{align}
# &\text{vec}(A + B)=\text{vec}(A)+\text{vec}(B) \\
# &\text{vec}(AXB) = (B^T \otimes A)\text{vec}(X) \\
# &\text{vec}(A^T)=K_{mn}\text{vec}(A),A\in \mathbb R^{m\times n},K_{mn}\in \mathbb R^{mn\times mn}, \quad K_{mn} \text{ is commutation matrix} \\
# &\text{vec}(A\odot X)=\text{dial}(A)\text{vec}(X), \text{\quad diag}(A)\in \mathbb R^{mn\times mn} \text{ is a diagonal use elements in A, oredered by columns} \\
# &(A\otimes B)^T=A^T\otimes B^T \\
# &\text{vec}(ab^T)=b\otimes a \\
# &(A\otimes B)(C\otimes D)=(AC)\otimes(BD) \\
# &K_{mn}=K^T_{nm}, K_{mn}K_{nm}=I \\
# &K_{pm}(A\otimes B)K_{nq}=B\otimes A, A\in \mathbb R^{m\times n}, B\in \mathbb R^{p\times q} \\
# \end{align}
# $$
#
#
| Coding/mlp_bp_question.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generalizing trajectories
#
#
#
# <img align="right" src="https://anitagraser.github.io/movingpandas/pics/movingpandas.png">
#
# [](https://mybinder.org/v2/gh/anitagraser/movingpandas-examples/main?filepath=1-tutorials/7-generalizing-trajectories.ipynb)
#
# To reduce the size of trajectory objects, we can generalize them, for example, using the Douglas-Peucker algorithm:
# +
import pandas as pd
import geopandas as gpd
from geopandas import GeoDataFrame, read_file
from shapely.geometry import Point, LineString, Polygon
from datetime import datetime, timedelta
import movingpandas as mpd
print(mpd.__version__)
# -
gdf = read_file('../data/geolife_small.gpkg')
gdf['t'] = pd.to_datetime(gdf['t'])
gdf = gdf.set_index('t').tz_localize(None)
traj_collection = mpd.TrajectoryCollection(gdf, 'trajectory_id')
original_traj = traj_collection.trajectories[1]
print(original_traj)
original_traj.plot(column='speed', linewidth=5, capstyle='round', figsize=(9,3), legend=True, vmax=20)
# ## DouglasPeuckerGeneralizer
# Try different tolerance settings and observe the results in line geometry and therefore also length:
help(mpd.DouglasPeuckerGeneralizer)
generalized_traj = mpd.DouglasPeuckerGeneralizer(original_traj).generalize(tolerance=0.001)
generalized_traj.plot(column='speed', linewidth=5, capstyle='round', figsize=(9,3), legend=True, vmax=20)
print('Original length: %s'%(original_traj.get_length()))
print('Generalized length: %s'%(generalized_traj.get_length()))
# ## MinTimeDeltaGeneralizer
#
# An alternative generalization method is to down-sample the trajectory to ensure a certain time delta between records:
help(mpd.MinTimeDeltaGeneralizer)
time_generalized = mpd.MinTimeDeltaGeneralizer(original_traj).generalize(tolerance=timedelta(minutes=1))
time_generalized.plot(column='speed', linewidth=5, capstyle='round', figsize=(9,3), legend=True, vmax=20)
time_generalized.to_point_gdf().head(10)
original_traj.to_point_gdf().head(10)
| 1-tutorials/7-generalizing-trajectories.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sentence Level Aspect-Based Sentiment Analysis with TreeLSTMs
#
# This notebook trains a Constituency Tree-LSTM model on the Laptop review dataset using Tensorflow Fold.
import tensorflow as tf
sess = tf.InteractiveSession()
import tensorflow_fold as td
import gensim
import numpy as np
import math
# ## Data Loading & Preprocessing
#
# We load the tree strings from the tree folder and convert them into tree objects. The list of tree objects is passed to the main model for training/evaluation.
#
# The code in the cell below creates tree objects and provides utilities to operate over them.
# +
class Node: # a node in the tree
def __init__(self, label = None, word=None):
self.label = label
self.word = word
self.parent = None # reference to parent
self.left = None # reference to left child
self.right = None # reference to right child
# true if I am a leaf (could have probably derived this from if I have
# a word)
self.isLeaf = False
# true if we have finished performing fowardprop on this node (note,
# there are many ways to implement the recursion.. some might not
# require this flag)
self.level = 0
#defeault intitialziation of depth
self.has_label = False
class Tree:
def __init__(self, treeString, openChar='(', closeChar=')', label_size = 18):
tokens = []
self.open = '('
self.close = ')'
for toks in treeString.strip().split():
tokens += list(toks)
self.root = self.parse(tokens, label_size = label_size)
self.self_binarize() #ensure binary parse tree - a node can have 0 or 2 child nodes
self.binary = check_for_binarization(self.root)
assert self.binary == True, "Tree is not binary"
self.depth = get_depth(self.root)
self.levels = max(math.floor(math.log(float(self.depth)) / math.log(float(2)))-1, 0)
self.labels = get_labels(self.root)
def parse(self, tokens, parent=None, label_size = 18):
assert tokens[0] == self.open, "Malformed tree"
assert tokens[-1] == self.close, "Malformed tree"
split = 1 # position after open
marker = 1
countOpen = countClose = 0
label = None
if (split + label_size) < len(tokens):
str1 = ''.join(tokens[split: (split + label_size)])
if str1.isdigit():
label = tokens[split: (split + label_size)]
label = np.asarray(label).astype(int)
split += label_size
marker += label_size
if tokens[split] == self.open:
countOpen += 1
split += 1
# Find where left child and right child split
while countOpen != countClose:
if tokens[split] == self.open:
countOpen += 1
if tokens[split] == self.close:
countClose += 1
split += 1
# New node
if isinstance(label, np.ndarray):
node = Node(label)
node.has_label = True
else:
node = Node()
if parent:
node.parent = parent
node.level = parent.level + 1
# leaf Node
if countOpen == 0:
node.word = ''.join(tokens[marker:-1]) # distinguish between lower and upper. Important for words like Apple
node.isLeaf = True
return node
node.left = self.parse(tokens[marker:split], parent=node)
if (tokens[split] == self.open) :
node.right = self.parse(tokens[split:-1], parent=node)
return node
def get_words(self):
def get_leaves(node):
if node is None:
return []
if node.isLeaf:
return [node]
else:
return getLeaves(node.left) + getLeaves(node.right)
leaves = getLeaves(self.root)
words = [node.word for node in leaves]
return words
def self_binarize(self):
def binarize_tree(node):
if node.isLeaf:
return
elif ((node.left is not None) & (node.right is not None)):
binarize_tree(node.left)
binarize_tree(node.right)
else:
#fuse parent node with child node
node.left.label = node.label
node.left.level -= 1
if (node.level != 0):
if (node.parent.right is node):
node.parent.right = node.left
else:
node.parent.left = node.left
node.left.parent = node.parent
else:
self.root = node.left
node.left.parent = None
self.root.has_label = True
binarize_tree(node.left)
binarize_tree(self.root)
#optional function to push labels to child nodes from root node, Not needed for LSTM trees
def propagate_label(node, levels, depth):
if node is None:
return
if (node.level > levels):
return
if node.parent:
node.label = node.parent.label
node.has_label = True
propagate_label(node.left, levels, depth)
propagate_label(node.right, levels, depth)
def get_depth(node):
if node is None:
return
if node.isLeaf:
return 0
return (1+ max(get_depth(node.left), get_depth(node.right)))
def get_labels(node):
if node is None:
return []
if node.has_label == False:
return []
return get_labels(node.left) + get_labels(node.right) + [node.label]
def check_for_binarization(node): #check whether we have a binary parse tree
if node.isLeaf:
return True
elif (node.right is None):
return False
else:
b1 = check_for_binarization(node.left)
b2 = check_for_binarization(node.right)
return (b1 & b2)
# -
# We load the strings and convert them into a list of tree objects.
def loadTrees(dataSet='train'):
"""
Loads training trees. Maps leaf node words to word ids.
"""
file = 'trees/%s.txt' % dataSet
print ("Loading %s trees.." % dataSet)
with open(file, 'r') as fid:
trees = [Tree(l) for l in fid.readlines()]
return trees
train_trees = loadTrees('train')
dev_trees = loadTrees('dev')
test_trees = loadTrees('test')
# Create a list of root nodes for each of the tree objects.
train_nodes = [t.root for t in train_trees]
dev_nodes = [t.root for t in dev_trees]
test_nodes = [t.root for t in test_trees]
# Load the entire Google Word2vec corpus into memory. This will take a few minutes.
def loadmodel():
print("Loading Google Word2vecs....")
model = gensim.models.KeyedVectors.load_word2vec_format('./GoogleNews-vectors-negative300.bin.gz', binary = True)
return model
model = loadmodel()
# Create a dictionary that maps a word to word2vec only for words in the training, dev and test set.
#only retrain words that are in train, dev and test sets
def filter_model(model):
filtered_dict = {}
trees = loadTrees('train') + loadTrees('dev') + loadTrees('test')
words = [t.get_words() for t in trees]
vocab = set()
for word in words:
vocab.update(word)
for word in vocab:
if word in model.vocab:
filtered_dict[word] = model[word]
return filtered_dict
filtered_model = filter_model(model)
# Loads embedings, returns weight matrix and dict from words to indices.
def load_embeddings(filtered_model):
print('loading word embeddings')
weight_vectors = []
word_idx = {}
for word, vector in filtered_model.items():
word_idx[word] = len(weight_vectors)
weight_vectors.append(np.array(vector, dtype=np.float32))
# Random embedding vector for unknown words.
weight_vectors.append(np.random.uniform(
-0.05, 0.05, weight_vectors[0].shape).astype(np.float32))
return np.stack(weight_vectors), word_idx
weight_matrix, word_idx = load_embeddings(filtered_model)
class BinaryTreeLSTMCell(tf.contrib.rnn.BasicLSTMCell):
"""LSTM with two state inputs.
This is the model described in section 3.2 of 'Improved Semantic
Representations From Tree-Structured Long Short-Term Memory
Networks' <http://arxiv.org/pdf/1503.00075.pdf>, with recurrent
dropout as described in 'Recurrent Dropout without Memory Loss'
<http://arxiv.org/pdf/1603.05118.pdf>.
"""
def __init__(self, num_units, keep_prob=1.0):
"""Initialize the cell.
Args:
num_units: int, The number of units in the LSTM cell.
keep_prob: Keep probability for recurrent dropout.
"""
super(BinaryTreeLSTMCell, self).__init__(num_units)
self._keep_prob = keep_prob
def __call__(self, inputs, state, scope=None):
with tf.variable_scope(scope or type(self).__name__):
lhs, rhs = state
c0, h0 = lhs
c1, h1 = rhs
concat = tf.contrib.layers.linear(
tf.concat([inputs, h0, h1], 1), 5 * self._num_units)
# i = input_gate, j = new_input, f = forget_gate, o = output_gate
i, j, f0, f1, o = tf.split(value=concat, num_or_size_splits=5, axis=1)
j = self._activation(j)
if not isinstance(self._keep_prob, float) or self._keep_prob < 1:
j = tf.nn.dropout(j, self._keep_prob)
new_c = (c0 * tf.sigmoid(f0 + self._forget_bias) +
c1 * tf.sigmoid(f1 + self._forget_bias) +
tf.sigmoid(i) * j)
new_h = self._activation(new_c) * tf.sigmoid(o)
new_state = tf.contrib.rnn.LSTMStateTuple(new_c, new_h)
return new_h, new_state
keep_prob_ph = tf.placeholder_with_default(1.0, [])
lstm_num_units = 300 # Tai et al. used 150, but our regularization strategy is more effective
tree_lstm = td.ScopedLayer(
tf.contrib.rnn.DropoutWrapper(
BinaryTreeLSTMCell(lstm_num_units, keep_prob=keep_prob_ph),
input_keep_prob=keep_prob_ph, output_keep_prob=keep_prob_ph),
name_or_scope='tree_lstm')
NUM_ASPECTS = 18 # number of aspects
NUM_POLARITY = 3 #number of polarity classes assicated with an aspect (1 = mildly +ve or -ve, 2 = -ve, 3 = +ve)
output_layer = td.FC(NUM_ASPECTS*(NUM_POLARITY+2), activation=None, name='output_layer')
word_embedding = td.Embedding(
*weight_matrix.shape, initializer=weight_matrix, name='word_embedding', trainable = False)
embed_subtree = td.ForwardDeclaration(name='embed_subtree')
def logits_and_state():
"""Creates a block that goes from tokens to (logits, state) tuples."""
unknown_idx = len(word_idx)
lookup_word = lambda word: word_idx.get(word, unknown_idx)
word2vec = (td.GetItem(0) >> td.InputTransform(lookup_word) >>
td.Scalar('int32') >> word_embedding)
pair2vec = (embed_subtree(), embed_subtree())
# Trees are binary, so the tree layer takes two states as its input_state.
zero_state = td.Zeros((tree_lstm.state_size,) * 2)
# Input is a word vector.
zero_inp = td.Zeros(word_embedding.output_type.shape[0])
word_case = td.AllOf(word2vec, zero_state)
pair_case = td.AllOf(zero_inp, pair2vec)
tree2vec = td.OneOf(len, [(1, word_case), (2, pair_case)])
return tree2vec >> tree_lstm >> (output_layer, td.Identity())
def tf_node_loss(logits, labels):
logits_ = tf.reshape(logits, [-1, NUM_ASPECTS, NUM_POLARITY + 2])
#compute loss related to task 1: aspect detection
binarized = tf.cast((labels > 0), tf.int32) #binarize the labels to compute loss for aspect detection
logits2 = tf.slice(logits_, [0,0,0], [-1,-1, 2])
loss2 = tf.reduce_sum(tf.nn.sparse_softmax_cross_entropy_with_logits(logits = logits2, labels=binarized), axis = 1)
# compute loss related to task 2: polarity prediction
padding = tf.constant([[0,0], [0,0], [1,0]])
logits3 = tf.pad(tf.log(tf.nn.softmax(tf.slice(logits_, [0, 0, 2], [-1,-1, -1]))), padding)
labels2 = tf.pad(tf.slice(tf.one_hot(labels, depth = 4, axis = -1), [0,0,1], [-1,-1,-1]), padding)
loss3 = tf.reduce_sum(tf.multiply(labels2, logits3), [1,2])
final_loss = loss2 + tf.scalar_mul(-1.05,loss3)
return final_loss
# +
#Task 2: compute true positives for aspect polarities
def task2_truepositives(logits, labels):
logits_ = tf.reshape(logits, [-1, NUM_ASPECTS, NUM_POLARITY + 2])
predictions = tf.cast(((( logits_[:,:, 2] ) > (logits_[:,:, 3] )) & (( logits_[:,:, 2] ) > (logits_[:,:, 4] ))), tf.float64)
actuals = tf.cast(((labels > 0) & (labels < 2)), tf.float64)
ones_like_actuals = tf.ones_like(actuals)
zeros_like_actuals = tf.zeros_like(actuals)
ones_like_predictions = tf.ones_like(predictions)
zeros_like_predictions = tf.zeros_like(predictions)
ans_1 = tf.reduce_sum(
tf.cast(
tf.logical_and(
tf.equal(actuals, ones_like_actuals),
tf.equal(predictions, ones_like_predictions)
),
tf.float64
), axis = 1
)
predictions = tf.cast(((( logits_[:,:, 3] ) > (logits_[:,:, 2] )) & (( logits_[:,:, 3] ) > (logits_[:,:, 4] ))), tf.float64)
actuals = tf.cast(((labels > 1) & (labels < 3)), tf.float64)
ones_like_actuals = tf.ones_like(actuals)
zeros_like_actuals = tf.zeros_like(actuals)
ones_like_predictions = tf.ones_like(predictions)
zeros_like_predictions = tf.zeros_like(predictions)
ans_2 = tf.reduce_sum(
tf.cast(
tf.logical_and(
tf.equal(actuals, ones_like_actuals),
tf.equal(predictions, ones_like_predictions)
),
tf.float64
), axis = 1
)
predictions = tf.cast(((( logits_[:,:, 4] ) > (logits_[:,:, 2] )) & (( logits_[:,:, 4] ) > (logits_[:,:, 3] ))), tf.float64)
actuals = tf.cast(((labels > 2) & (labels < 4)), tf.float64)
ones_like_actuals = tf.ones_like(actuals)
zeros_like_actuals = tf.zeros_like(actuals)
ones_like_predictions = tf.ones_like(predictions)
zeros_like_predictions = tf.zeros_like(predictions)
ans_3 = tf.reduce_sum(
tf.cast(
tf.logical_and(
tf.equal(actuals, ones_like_actuals),
tf.equal(predictions, ones_like_predictions)
),
tf.float64
), axis = 1
)
return ans_1 + ans_2 + ans_3
# -
#Task 2: compute total number of aspects
def task2_dem(logits, labels):
actuals = tf.cast(labels > 0, tf.float64)
ones_like_actuals = tf.ones_like(actuals)
return tf.reduce_sum(tf.cast(tf.equal(actuals, ones_like_actuals), tf.float64), axis = 1)
#Task 1: compute true positive rate
def tf_tpr(logits, labels):
logits_ = tf.nn.softmax(tf.reshape(logits, [-1, NUM_ASPECTS, NUM_POLARITY + 2]))
predictions = tf.cast(( logits_[:,:, 1] ) > (logits_[:,:, 0]), tf.float64)
actuals = tf.cast( labels > 0, tf.float64)
ones_like_actuals = tf.ones_like(actuals)
zeros_like_actuals = tf.zeros_like(actuals)
ones_like_predictions = tf.ones_like(predictions)
zeros_like_predictions = tf.zeros_like(predictions)
ans = tf.reduce_sum(
tf.cast(
tf.logical_and(
tf.equal(actuals, ones_like_actuals),
tf.equal(predictions, ones_like_predictions)
),
tf.float64
), axis = 1
)
return ans
#Task 1: compute true negative rate
def tf_tnr(logits, labels):
logits_ = tf.nn.softmax(tf.reshape(logits, [-1, NUM_ASPECTS, NUM_POLARITY + 2]))
predictions = tf.cast((logits_[:,:, 1] ) > (logits_[:,:, 0]), tf.float64)
actuals = tf.cast(labels > 0, tf.float64)
ones_like_actuals = tf.ones_like(actuals)
zeros_like_actuals = tf.zeros_like(actuals)
ones_like_predictions = tf.ones_like(predictions)
zeros_like_predictions = tf.zeros_like(predictions)
ans = tf.reduce_sum(
tf.cast(
tf.logical_and(
tf.equal(actuals, zeros_like_actuals),
tf.equal(predictions, zeros_like_predictions)
),
tf.float64
), axis = 1
)
return ans
#Task 1: compute false positive rate
def tf_fpr(logits, labels):
logits_ = tf.nn.softmax(tf.reshape(logits, [-1, NUM_ASPECTS, NUM_POLARITY + 2]))
predictions = tf.cast((logits_[:,:, 1] ) > (logits_[:,:, 0]), tf.float64)
actuals = tf.cast(labels > 0, tf.float64)
ones_like_actuals = tf.ones_like(actuals)
zeros_like_actuals = tf.zeros_like(actuals)
ones_like_predictions = tf.ones_like(predictions)
zeros_like_predictions = tf.zeros_like(predictions)
ans = tf.reduce_sum(
tf.cast(
tf.logical_and(
tf.equal(actuals, zeros_like_actuals),
tf.equal(predictions, ones_like_predictions)
),
tf.float64
), axis = 1
)
return ans
#Task 1: compute false negative rate
def tf_fnr(logits, labels):
logits_ = tf.nn.softmax(tf.reshape(logits, [-1, NUM_ASPECTS, NUM_POLARITY + 2]))
predictions = tf.cast((logits_[:,:, 1] ) > (logits_[:,:, 0]), tf.float64)
actuals = tf.cast(labels > 0, tf.float64)
ones_like_actuals = tf.ones_like(actuals)
zeros_like_actuals = tf.zeros_like(actuals)
ones_like_predictions = tf.ones_like(predictions)
zeros_like_predictions = tf.zeros_like(predictions)
ans = tf.reduce_sum(
tf.cast(
tf.logical_and(
tf.equal(actuals, ones_like_actuals),
tf.equal(predictions, zeros_like_predictions)
),
tf.float64
), axis = 1
)
return ans
def add_metrics(is_root, is_neutral):
"""A block that adds metrics for loss and hits; output is the LSTM state."""
c = td.Composition(
name='predict(is_root=%s, is_neutral=%s)' % (is_root, is_neutral))
with c.scope():
# destructure the input; (labels, neutral, (logits, state))
labels = c.input[0]
logits = td.GetItem(0).reads(c.input[2])
state = td.GetItem(1).reads(c.input[2])
loss = td.Function(tf_node_loss)
td.Metric('all_loss').reads(loss.reads(logits, labels))
if is_root: td.Metric('root_loss').reads(loss)
tpr = td.Function(tf_tpr)
tnr = td.Function(tf_tnr)
fpr = td.Function(tf_fpr)
fnr = td.Function(tf_fnr)
t2_acc = td.Function(task2_truepositives)
t2_dem = td.Function(task2_dem)
td.Metric('all_tpr').reads(tpr.reads(logits, labels))
td.Metric('all_tnr').reads(tnr.reads(logits, labels))
td.Metric('all_fpr').reads(fpr.reads(logits, labels))
td.Metric('all_fnr').reads(fnr.reads(logits, labels))
td.Metric('all_task2').reads(t2_acc.reads(logits, labels))
td.Metric('all_task2dem').reads(t2_dem.reads(logits, labels))
if is_root:
td.Metric('tpr').reads(tpr)
td.Metric('tnr').reads(tnr)
td.Metric('fpr').reads(fpr)
td.Metric('fnr').reads(fnr)
td.Metric('task2').reads(t2_acc)
td.Metric('task2dem').reads(t2_dem)
# output the state, which will be read by our by parent's LSTM cell
c.output.reads(state)
return c
def tokenize(node):
group = []
neutral = '2'
if node.has_label:
label = node.label
neutral = '1'
else:
label = np.zeros((NUM_ASPECTS,),dtype=np.int)
if node.isLeaf:
group = [node.word]
else:
group = [node.left, node.right]
return label, neutral, group
node = train_nodes[0]
label, neutral, group = tokenize(node)
print (len(group))
print (label.shape)
def embed_tree(logits_and_state, is_root):
"""Creates a block that embeds trees; output is tree LSTM state."""
return td.InputTransform(tokenize) >> td.OneOf(
key_fn=lambda pair: pair[1] == '2', # label 2 means neutral
case_blocks=(add_metrics(is_root, is_neutral=False),
add_metrics(is_root, is_neutral=True)),
pre_block=(td.Vector(NUM_ASPECTS, dtype = 'int32'), td.Scalar('int32'), logits_and_state))
model = embed_tree(logits_and_state(), is_root=True)
embed_subtree.resolve_to(embed_tree(logits_and_state(), is_root=False))
compiler = td.Compiler.create(model)
print('input type: %s' % model.input_type)
print('output type: %s' % model.output_type)
metrics = {k: tf.reduce_mean(v) for k, v in compiler.metric_tensors.items()}
LEARNING_RATE = 0.05
KEEP_PROB = 0.60
BATCH_SIZE = 32
EPOCHS = 80
global_step = tf.Variable(0, trainable=False)
starter_learning_rate = LEARNING_RATE
learning_rate = tf.train.exponential_decay(starter_learning_rate, global_step,
600, 0.90, staircase=True)
train_feed_dict = {keep_prob_ph: KEEP_PROB}
loss = tf.reduce_mean(compiler.metric_tensors['root_loss'])
opt = tf.train.AdagradOptimizer(LEARNING_RATE)
learning_step = (
tf.train.AdagradOptimizer(learning_rate)
.minimize(loss, global_step=global_step)
)
sess.run(tf.global_variables_initializer())
def train_step(batch):
train_feed_dict[compiler.loom_input_tensor] = batch
_, batch_loss = sess.run([learning_step, loss], train_feed_dict)
return batch_loss
def train_epoch(train_set):
list = [train_step(batch) for batch in td.group_by_batches(train_set, BATCH_SIZE)]
return sum(list)/ max(len(list), 1)
train_set = compiler.build_loom_inputs(train_nodes)
dev_feed_dict = compiler.build_feed_dict(dev_nodes)
def dev_eval(epoch, train_loss):
dev_metrics = sess.run(metrics, dev_feed_dict)
dev_loss = dev_metrics['root_loss']
tp = dev_metrics['tpr']
tn = dev_metrics['tnr']
fp = dev_metrics['fpr']
fn = dev_metrics['fnr']
tpr = float(tp)/(float(tp) + float(fn))
fpr = float(fp)/(float(tp) + float(fn))
t2_acc = float(dev_metrics['task2'])/ float(dev_metrics['task2dem'])
recall = tpr
if (float(tp) + float(fp)) > 0:
precision = float(tp)/(float(tp) + float(fp))
else: precision = 0.
if precision + recall > 0:
f1_score = (2 * (precision * recall)) / (precision + recall)
else: f1_score = 0.
print('epoch:%4d, train_loss: %.3e, dev_loss: %.3e,Task1 Precision: %.3e, Task1 Recall: %.3e, Task1 F1 score: %2.3e, Task2 Acc: %2.3e'
% (epoch, train_loss, dev_loss, precision, recall, f1_score, t2_acc))
return f1_score
best_accuracy = 0.0
save_path = 'weights/sentiment_model'
for epoch, shuffled in enumerate(td.epochs(train_set, EPOCHS), 1):
train_loss = train_epoch(shuffled)
f1_score = dev_eval(epoch, train_loss)
| LSTM_TreeModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Calculate Bulk Wind Shear for JSON Sounding Files
# +
import metpy.calc as mpcalc
from metpy.units import units
import pandas as pd
import numpy as np
from metpy.calc.tools import get_layer, _get_bound_pressure_height
from metpy.calc.tools import _less_or_close, _greater_or_close, log_interpolate_1d
# -
df = pd.read_json("data_IAD.json")
df
p = df['pres'].values[0][0] * units.hPa
t = df['tmpc'].values[0][0] * units.degC
td = df['dwpt'].values[0][0] * units.degC
dir = df['wdir'].values[0][0] * units.degrees
spd = df['wspd'].values[0][0] * units.knots
heights = df['hght'].values[0][0] * units.meter
u, v = mpcalc.wind_components(df["wspd"].values[0][0] * units.knot, df["wdir"].values[0][0] * units.deg)
# ## Quick function to get bulk shear at desired layer (meters) for JSON files
#
def get_bulk_shear(sound_file,depth):
"""Get bulk shear for desired layer depth based on JSON sounding file
Args
----
sound_file : str
JSON sounding file name
depth : int
layer depth desired in meters
Returns
-------
Prints u, v, speed, and direction for bilk shear values
u_bulk_shear : pint.quantity.build_quantity_class.<locals>.Quantity
u-component of layer bulk shear
v_bulk_shear : pint.quantity.build_quantity_class.<locals>.Quantity
v-component of layer bulk shear
bulk_shear_speed : pint.quantity.build_quantity_class.<locals>.Quantity
layer bulk shear wind speed
bulk_shear_dir : pint.quantity.build_quantity_class.<locals>.Quantity
layer bulk shear wind direction
"""
printmd(f"\n**Sounding Location: {sound_file}**")
print(f"Desired layer: {depth/1000}km\n"+\
"---------------------------------")
df = pd.read_json(sound_file)
p = df['pres'].values[0][0] * units.hPa
Z = df['hght'].values[0][0] * units.meter
def replace_empty_str(col):
for i in range(len(df[col][0][0][:])):
if df[col][0][0][i] == '':
df[col][0][0][i] = 0
return df
for i in df.columns:
replace_empty_str(i)
u, v = mpcalc.wind_components(df["wspd"].values[0][0] * units.knot, df["wdir"].values[0][0] * units.deg)
u_bulk_shear,v_bulk_shear = mpcalc.bulk_shear(p,u,v,heights=Z,depth=depth * units.meter)
print(f"u-bulk shear: {u_bulk_shear}\nv-bulk shear: {v_bulk_shear}")
bulk_shear_speed = np.sqrt(u_bulk_shear**2 + v_bulk_shear**2)
bulk_shear_dir = mpcalc.wind_direction(u_bulk_shear,v_bulk_shear)
print(f"bulk shear speed: {bulk_shear_speed}\nbulk shear direction: {bulk_shear_dir}")
return u_bulk_shear, v_bulk_shear, bulk_shear_speed, bulk_shear_dir
def get_layer(pressure, *args, heights=None, bottom=None, depth=100 * units.hPa,
interpolate=True):
r"""Return an atmospheric layer from upper air data with the requested bottom and depth.
This function will subset an upper air dataset to contain only the specified layer. The
bottom of the layer can be specified with a pressure or height above the surface
pressure. The bottom defaults to the surface pressure. The depth of the layer can be
specified in terms of pressure or height above the bottom of the layer. If the top and
bottom of the layer are not in the data, they are interpolated by default.
Parameters
----------
pressure : array-like
Atmospheric pressure profile
args : array-like
Atmospheric variable(s) measured at the given pressures
heights: array-like, optional
Atmospheric heights corresponding to the given pressures. Defaults to using
heights calculated from ``p`` assuming a standard atmosphere [NOAA1976]_.
bottom : `pint.Quantity`, optional
The bottom of the layer as a pressure or height above the surface pressure. Defaults
to the highest pressure or lowest height given.
depth : `pint.Quantity`, optional
The thickness of the layer as a pressure or height above the bottom of the layer.
Defaults to 100 hPa.
interpolate : bool, optional
Interpolate the top and bottom points if they are not in the given data. Defaults
to True.
Returns
-------
`pint.Quantity, pint.Quantity`
The pressure and data variables of the layer
"""
# If we get the depth kwarg, but it's None, set it to the default as well
if depth is None:
depth = 100 * units.hPa
# Make sure pressure and datavars are the same length
for datavar in args:
if len(pressure) != len(datavar):
raise ValueError('Pressure and data variables must have the same length.')
# If the bottom is not specified, make it the surface pressure
if bottom is None:
bottom = np.nanmax(pressure.m) * pressure.units
bottom_pressure, bottom_height = _get_bound_pressure_height(pressure, bottom,
heights=heights,
interpolate=interpolate)
# Calculate the top if whatever units depth is in
if depth.dimensionality == {'[length]': -1.0, '[mass]': 1.0, '[time]': -2.0}:
top = bottom_pressure - depth
elif depth.dimensionality == {'[length]': 1}:
top = bottom_height + depth
else:
raise ValueError('Depth must be specified in units of length or pressure')
top_pressure, _ = _get_bound_pressure_height(pressure, top, heights=heights,
interpolate=interpolate)
ret = [] # returned data variables in layer
# Ensure pressures are sorted in ascending order
sort_inds = np.argsort(pressure)
pressure = pressure[sort_inds]
# Mask based on top and bottom pressure
inds = (_less_or_close(pressure, bottom_pressure)
& _greater_or_close(pressure, top_pressure))
p_interp = pressure[inds]
# Interpolate pressures at bounds if necessary and sort
if interpolate:
# If we don't have the bottom or top requested, append them
if not np.any(np.isclose(top_pressure, p_interp)):
p_interp = np.sort(np.append(p_interp.m, top_pressure.m)) * pressure.units
if not np.any(np.isclose(bottom_pressure, p_interp)):
p_interp = np.sort(np.append(p_interp.m, bottom_pressure.m)) * pressure.units
ret.append(p_interp[::-1])
for datavar in args:
# Ensure that things are sorted in ascending order
datavar = datavar[sort_inds]
if interpolate:
# Interpolate for the possibly missing bottom/top values
datavar_interp = log_interpolate_1d(p_interp, pressure, datavar)
datavar = datavar_interp
else:
datavar = datavar[inds]
ret.append(datavar[::-1])
return ret
# +
from metpy import constants as mpconsts
t0 = 288. * units.kelvin
p0 = 1013.25 * units.hPa
def height_to_pressure_std(height):
r"""Convert height data to pressures using the U.S. standard atmosphere [NOAA1976]_.
The implementation inverts the formula outlined in [Hobbs1977]_ pg.60-61.
Parameters
----------
height : `pint.Quantity`
Atmospheric height
Returns
-------
`pint.Quantity`
The corresponding pressure value(s)
Notes
-----
.. math:: p = p_0 e^{\frac{g}{R \Gamma} \text{ln}(1-\frac{Z \Gamma}{T_0})}
"""
gamma = 6.5 * units('K/km')
return p0 * (1 - (gamma / t0) * height) ** (mpconsts.g / (mpconsts.Rd * gamma))
def pressure_to_height_std(pressure):
r"""Convert pressure data to heights using the U.S. standard atmosphere [NOAA1976]_.
The implementation uses the formula outlined in [Hobbs1977]_ pg.60-61.
Parameters
----------
pressure : `pint.Quantity`
Atmospheric pressure
Returns
-------
`pint.Quantity`
The corresponding height value(s)
Notes
-----
.. math:: Z = \frac{T_0}{\Gamma}[1-\frac{p}{p_0}^\frac{R\Gamma}{g}]
"""
gamma = 6.5 * units('K/km')
return (t0 / gamma) * (1 - (pressure / p0).to('dimensionless')**(
mpconsts.Rd * gamma / mpconsts.g))
# -
_get_bound_pressure_height(p, 6000*units.m, heights=heights, interpolate=True)
# +
p_layer, t_layer, u_layer, v_layer, dir_layer, spd_layer, hghts_layer = get_layer(
p, t, u, v, dir, spd, heights, heights=heights, depth=6000*units.meter)
printmd("**Top of 6km layer (from get_layer)**")
print("==================================")
print("Pressure: " + str(p_layer[-1]))
print("Height (from sounding): " + str(hghts_layer[-1]))
print("Height (from std atm): " + str(pressure_to_height_std(p_layer[-1])))
print()
printmd("**Expected top of 6km layer**")
print("==========================")
#baseHght = pressure_to_height_std(p[0])
baseHght = heights[0]
pressure_bound, height_bound = _get_bound_pressure_height(p, 6000*units.meter+baseHght)
stdPres = height_to_pressure_std(6000*units.meter+baseHght)
stdHght = pressure_to_height_std(stdPres)
print("Pressure bounds from _get_bound_pressure_height: " + str(pressure_bound))
print("Height bounds from _get_bound_pressure_height: " + str(height_bound))
print("Base Height (from std atm): " + str(baseHght))
print("6km Pressure (from std atm): " + str(stdPres))
print("6km-Pressure Height (from std atm): " + str(stdHght))
print()
printmd("**Bottom of 6km layer**")
print("=================")
print("Pressure: " + str(p_layer[0]))
print("Heights (from sounding): " + str(hghts_layer[0]))
print("Heights (from std atm): " + str(pressure_to_height_std(p_layer[0])))
print()
ushr = u_layer[-1] - u_layer[0]
vshr = v_layer[-1] - v_layer[0]
print("U-Shear component: " + str(ushr))
print("V-Shear component: " + str(vshr))
print("Bulk Shear Speed: " + str(np.sqrt(ushr**2 + vshr**2)))
u_bulk_shear, v_bulk_shear, bulk_shear_speed, bulk_shear_dir = get_bulk_shear("data_IAD.json", 6000)
# -
from IPython.display import Markdown, display
def printmd(string):
display(Markdown(string))
def calc_BRN(pressure, u, v, temp, cape, heights=None):
r"""Calculate Bulk Richardson Number and BRN shear.
Parameters
----------
pressure : `pint.Quantity`
Atmospheric pressure profile
u : `pint.Quantity`
U-component of wind.
v : `pint.Quantity`
V-component of wind.
temp : `pint.Quantity`
Atmospheric temperature profile (can be either temperature or virtual temperature)
cape : `pint.Quantity`
CAPE value to use as the numerator of the BRN calculation.
heights : `pint.Quantity`, optional
Heights in meters from sounding (not AGL)
Returns
-------
brn : `pint.Quantity`
Bulk Richadson Number
brn_shr : `pint.Quantity`
Bulk Richardson Shear (the denominator of the BRN)
"""
from metpy import constants as mpconsts
baseZ = heights[0]
sumZ = -baseZ
rho6km = 0
u6km = 0
v6km = 0
u500 = 0
v500 = 0
for i, p in enumerate(pressure):
rho = (p.to('Pa')/(temp[i].to('K')*mpconsts.Rd)).to('kg / m^3')
print(rho)
sumZ += heights[i]
u_weighted = u[i]*rho
v_weighted = v[i]*rho
u_weighted = u_weighted.magnitude
v_weighted = v_weighted.magnitude
if (sumZ >= 6000 * units.meter):
layerZ=heights[i] - heights[i-1]
fraction = (6000 * units.meter - layerZ)/sumZ
fraction = fraction.to('dimensionless')
u_frac = ((u_weighted - uprev) * fraction) + uprev
v_frac = ((v_weighted - vprev) * fraction) + vprev
rho6km += rho
u_weighted = u_frac*rho
v_weighted = v_frac*rho
u_weighted = u_weighted.magnitude
v_weighted = v_weighted.magnitude
u6km += u_weighted
v6km += v_weighted
break
rho6km += rho
u6km += u_weighted
v6km += v_weighted
if (sumZ < 500 * units.meter):
u500 += u_weighted
v500 += v_weighted
divisor = i+1
uprev = u_weighted
vprev = v_weighted
u6kmAvg = u6km/(i+1)
v6kmAvg = v6km/(i+1)
print(u6kmAvg, v6kmAvg)
u500Avg = u500/divisor
v500Avg = v500/divisor
print(u500Avg, v500Avg)
uDiff = (u6kmAvg-u500Avg)
vDiff = (v6kmAvg-v500Avg)
print(uDiff, vDiff)
mag = np.sqrt(uDiff**2+vDiff**2)
brnshr = (mag**2)*.5
brnshr = brnshr.to('m^2/s^2')
brn = cape/brnu
brn = brn.magnitude
return brn, brnshr
calc_BRN(p,u,v,t,5065 * units('J/kg'),heights=heights)
def calc_BRN_2(pressure, u, v, temp, cape, heights=None):
r"""Calculate Bulk Richardson Number and BRN shear.
Parameters
----------
pressure : `pint.Quantity`
Atmospheric pressure profile
u : `pint.Quantity`
U-component of wind.
v : `pint.Quantity`
V-component of wind.
temp : `pint.Quantity`
Atmospheric temperature profile (can be either temperature or virtual temperature)
cape : `pint.Quantity`
CAPE value to use as the numerator of the BRN calculation.
heights : `pint.Quantity`, optional
Heights in meters from sounding (not AGL)
Returns
-------
brn : `pint.Quantity`
Bulk Richadson Number
brn_shr : `pint.Quantity`
Bulk Richardson Shear (the denominator of the BRN)
"""
from metpy import constants as mpconsts
baseZ = heights[0]
sumZ = -baseZ
rho6km = 0
u6km = 0
v6km = 0
u500 = 0
v500 = 0
rho = (pressure[0].to('Pa')/(temp[0].to('K')*mpconsts.Rd)).to('kg / m^3')
u_weighted = u[0]*rho
v_weighted = v[0]*rho
u_weighted = u_weighted.magnitude
v_weighted = v_weighted.magnitude
ubase_weighted = u_weighted
vbase_weighted = v_weighted
for i, p in enumerate(pressure):
rho = (p.to('Pa')/(temp[i].to('K')*mpconsts.Rd)).to('kg / m^3')
print(rho)
sumZ += heights[i]
u_weighted = u[i]*rho
v_weighted = v[i]*rho
u_weighted = u_weighted.magnitude
v_weighted = v_weighted.magnitude
if (sumZ >= 6000 * units.meter):
layerZ=heights[i] - heights[i-1]
fraction = (6000 * units.meter - layerZ)/sumZ
fraction = fraction.to('dimensionless')
u_frac = ((u_weighted - uprev) * fraction) + uprev
v_frac = ((v_weighted - vprev) * fraction) + vprev
u_weighted = u_frac*rho
v_weighted = v_frac*rho
u_weighted = u_weighted.magnitude
v_weighted = v_weighted.magnitude
u6km += u_weighted
v6km += v_weighted
break
if (sumZ < 500 * units.meter):
u500 += u_weighted
v500 += v_weighted
divisor = i+1
uprev = u_weighted
vprev = v_weighted
u6kmAvg = u6km/(i+1)
v6kmAvg = v6km/(i+1)
print(u6kmAvg, v6kmAvg)
u500Avg = u500/divisor
v500Avg = v500/divisor
print(u500Avg, v500Avg)
ushr = (u6kmAvg-u500Avg)
vshr = (v6kmAvg-v500Avg)
print(ushr, vshr)
mag = np.sqrt(ushr**2+vshr**2)
print(mag)
# brnu = (mag)*.5
brnshr = (mag**2)*.5
brnshr = brnshr.to('m^2/s^2')
brn = cape/brnshr
brn = brn.magnitude
return brn, brnshr
calc_BRN_2(p,u,v,t,5065 * units('J/kg'),heights=heights)
def calc_BRN_3(pressure, u, v, cape, heights=None):
r"""Calculate Bulk Richardson Number and BRN shear.
Parameters
----------
pressure : `pint.Quantity`
Atmospheric pressure profile
u : `pint.Quantity`
U-component of wind.
v : `pint.Quantity`
V-component of wind.
cape : `pint.Quantity`
CAPE value to use as the numerator of the BRN calculation.
heights : `pint.Quantity`, optional
Heights in meters from sounding (not AGL)
Returns
-------
brn : `pint.Quantity`
Bulk Richadson Number
brn_shr : `pint.Quantity`
Bulk Richardson Shear (the denominator of the BRN)
"""
p_layer500, u_layer500, v_layer500 = get_layer(p, u, v, heights=None, depth=500*units.meter)
u_layer500 = u_layer500.to('m/s')
v_layer500 = v_layer500.to('m/s')
avgU500 = sum(u_layer500 * p_layer500) / sum(p_layer500)
avgV500 = sum(v_layer500 * p_layer500) / sum(p_layer500)
print(avgU500, avgV500)
p_layer6km, u_layer6km, v_layer6km = get_layer(p, u, v, heights=None, depth=6000*units.meter)
u_layer6km = u_layer6km.to('m/s')
v_layer6km = v_layer6km.to('m/s')
avgU6km = sum(u_layer6km * p_layer6km) / sum(p_layer6km)
avgV6km = sum(v_layer6km * p_layer6km) / sum(p_layer6km)
print(avgU6km, avgV6km)
ushr = avgU6km-avgU500
vshr = avgV6km-avgV500
print(ushr, vshr)
mag = np.sqrt(ushr**2+vshr**2)
print(mag)
brnshr = (mag**2)*.5
brn = cape/brnshr
brn = brn.magnitude
return brn, brnshr
calc_BRN_3(p,u,v,5065 * units('J/kg'),heights=heights)
def calc_BRN_4(pressure, spd, cape, heights=None):
r"""Calculate Bulk Richardson Number and BRN shear.
Parameters
----------
pressure : `pint.Quantity`
Atmospheric pressure profile
spd : `pint.Quantity`
Wind speed.
cape : `pint.Quantity`
CAPE value to use as the numerator of the BRN calculation.
heights : `pint.Quantity`, optional
Heights in meters from sounding (not AGL)
Returns
-------
brn : `pint.Quantity`
Bulk Richadson Number
brn_shr : `pint.Quantity`
Bulk Richardson Shear (the denominator of the BRN)
"""
p_layer500, spd_layer500 = get_layer(p, spd, heights=None, depth=500*units.meter)
avg500 = sum(spd_layer500 * p_layer500) / sum(p_layer500)
print(avg500)
p_layer6km, spd_layer6km = get_layer(p, spd, heights=None, depth=6000*units.meter)
avg6km = sum(spd_layer6km * p_layer6km) / sum(p_layer6km)
print(avg6km)
shr = avg6km-avg500
print(shr)
print(spd_layer6km)
brnshr = (shr**2)*.5
brnshr = brnshr.to('m^2/s^2')
brn = cape/brnshr
brn = brn.magnitude
return brn, brnshr
calc_BRN_4(p, spd, 5065 * units('J/kg'), heights=heights)
def calc_BRN_5(pressure, u, v, cape, heights=None):
r"""Calculate Bulk Richardson Number and BRN shear.
Parameters
----------
pressure : `pint.Quantity`
Atmospheric pressure profile
u : `pint.Quantity`
U-component of wind.
v : `pint.Quantity`
V-component of wind.
cape : `pint.Quantity`
CAPE value to use as the numerator of the BRN calculation.
heights : `pint.Quantity`, optional
Heights in meters from sounding (not AGL)
Returns
-------
brn : `pint.Quantity`
Bulk Richadson Number
brn_shr : `pint.Quantity`
Bulk Richardson Shear (the denominator of the BRN)
"""
p_layer500, u_layer500, v_layer500 = get_layer(p, u, v, heights=None, depth=500*units.meter)
print(u_layer500, v_layer500)
avgU500 = sum(u_layer500) / p_layer500.size
avgV500 = sum(v_layer500) / p_layer500.size
print(avgU500, avgV500)
p_layer6km, u_layer6km, v_layer6km = get_layer(p, u, v, heights=None, depth=6000*units.meter)
print(u_layer6km, v_layer6km, p_layer6km)
avgU6km = sum(u_layer6km) / p_layer6km.size
avgV6km = sum(v_layer6km) / p_layer6km.size
print(avgU6km, avgV6km)
ushr = avgU6km-avgU500
vshr = avgV6km-avgV500
print(ushr, vshr)
mag = np.sqrt(ushr**2+vshr**2)
print(mag)
brnshr = (mag**2)*.5
brn = cape/brnshr
brn = brn.magnitude
return brn, brnshr
calc_BRN_5(p,u,v,5534 * units('J/kg'),heights=None)
def calc_BRN_6(pressure, u, v, temp, cape, heights=None):
r"""Calculate Bulk Richardson Number and BRN shear.
Parameters
----------
pressure : `pint.Quantity`
Atmospheric pressure profile
u : `pint.Quantity`
U-component of wind.
v : `pint.Quantity`
V-component of wind.
temp : `pint.Quantity`
Atmospheric temperature profile
cape : `pint.Quantity`
CAPE value to use as the numerator of the BRN calculation.
heights : `pint.Quantity`, optional
Heights in meters from sounding (not AGL)
Returns
-------
brn : `pint.Quantity`
Bulk Richadson Number
brn_shr : `pint.Quantity`
Bulk Richardson Shear (the denominator of the BRN)
"""
p_layer500, u_layer500, v_layer500, t_layer500 = get_layer(p, u, v, t, heights=None, depth=500*units.meter)
rho_layer500 = (p_layer500.to('Pa')/(t_layer500.to('K')*mpconsts.Rd)).to('kg/m^3')
print(rho_layer500)
u_layer500 = u_layer500.to('m/s')
v_layer500 = v_layer500.to('m/s')
avgU500 = sum(u_layer500 * rho_layer500) / sum(rho_layer500)
avgV500 = sum(v_layer500 * rho_layer500) / sum(rho_layer500)
print(avgU500, avgV500)
p_layer6km, u_layer6km, v_layer6km, t_layer6km = get_layer(p, u, v, t, heights=None, depth=6000*units.meter)
rho_layer6km = (p_layer6km.to('Pa')/(t_layer6km.to('K')*mpconsts.Rd)).to('kg/m^3')
print(rho_layer6km)
u_layer6km = u_layer6km.to('m/s')
v_layer6km = v_layer6km.to('m/s')
avgU6km = sum(u_layer6km * rho_layer6km) / sum(rho_layer6km)
avgV6km = sum(v_layer6km * rho_layer6km) / sum(rho_layer6km)
print(avgU6km, avgV6km)
ushr = avgU6km-avgU500
vshr = avgV6km-avgV500
print(ushr, vshr)
mag = np.sqrt(ushr**2+vshr**2)
print(mag)
brnshr = (mag**2)*.5
brnshr = brnshr.to("m^2/s^2")
brn = cape/brnshr
brn = brn.magnitude
return brn, brnshr
calc_BRN_6(p,u,v,t,5534 * units('J/kg'),heights=None)
def calc_BRN_7(pressure, u, v, cape, heights=None):
r"""Calculate Bulk Richardson Number and BRN shear.
Parameters
----------
pressure : `pint.Quantity`
Atmospheric pressure profile
u : `pint.Quantity`
U-component of wind.
v : `pint.Quantity`
V-component of wind.
temp : `pint.Quantity`
Atmospheric temperature profile
cape : `pint.Quantity`
CAPE value to use as the numerator of the BRN calculation.
heights : `pint.Quantity`, optional
Heights in meters from sounding (not AGL)
Returns
-------
brn : `pint.Quantity`
Bulk Richadson Number
brn_shr : `pint.Quantity`
Bulk Richardson Shear (the denominator of the BRN)
"""
p_layer500, u_layer500, v_layer500 = get_layer(p, u, v, heights=None, depth=500*units.meter)
u_layer500 = u_layer500.to('m/s')
v_layer500 = v_layer500.to('m/s')
avgU500 = sum(u_layer500 * p_layer500) / sum(p_layer500)
avgV500 = sum(v_layer500 * p_layer500) / sum(p_layer500)
print(avgU500, avgV500)
p_layer6km, u_layer6km, v_layer6km = get_layer(p, u, v, heights=None, depth=6000*units.meter)
u_layer6km = u_layer6km.to('m/s')
v_layer6km = v_layer6km.to('m/s')
avgU6km = sum(u_layer6km * p_layer6km) / sum(p_layer6km)
avgV6km = sum(v_layer6km * p_layer6km) / sum(p_layer6km)
print(avgU6km, avgV6km)
ushr = avgU6km-avgU500
vshr = avgV6km-avgV500
print(ushr, vshr)
mag = np.sqrt(ushr**2+vshr**2)
print(mag)
brnshr = (mag**2)*.5
brn = cape/brnshr
brn = brn.magnitude
return brn, brnshr
calc_BRN_7(p,u,v,5534 * units('J/kg'),heights=None)
| bulk_shear_calc_issue.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
# Problem 1
bike_data = pd.read_csv('../data/seattle-bike-data.csv')
# +
# Problem 2
# convert to datetime data type
bike_data['Date'] = pd.to_datetime(bike_data['Date'])
# i.
bike_data['Total Bicycle Count'] = bike_data['Fremont Bridge East Sidewalk'] + bike_data['Fremont Bridge West Sidewalk']
# ii
bike_data['Hour of Day'] = bike_data['Date'].dt.hour
# iii
bike_data['Year'] = bike_data['Date'].dt.year
# -
# Problem 3
bike_data_2016 = bike_data[bike_data['Year'] == 2016]
# Problem 4
plt.scatter('Hour of Day', 'Total Bicycle Count', data=bike_data, alpha=0.2)
plt.title('Total Bicycle Count vs Hour of Day')
plt.xlabel('Hour of Day')
plt.ylabel('Total Bicycle Count')
plt.legend();
# Problem 5
bike_data.groupby(by=['Hour of Day'], as_index=False) \
.mean() \
.sort_values(by=['Total Bicycle Count'], ascending=[False]) \
.iloc[0,[0, 3]]
| analysis/data-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduccion a Pandas
import pandas as pd
pd.__version__
# http://pandas.pydata.org/
#
# Pandas es la extensión logica de numpy al mundo del Análisis de datos.
#
# De forma muy general, Pandas extrae la figura del Dataframe conocida por aquellos que usan R a python.
#
# Un pandas dataframe es una tabla, lo que es una hoja de excel, con filas y columnas.
#
# En pandas cada columna es una Serie que esta definida con un numpy array por debajo.
# ### Qué puede hacer pandas por ti?
# - Cargar datos de diferentes recursos
# - Búsqueda de una fila o columna en particular
# - Realización de calculos estadísticos
# - Processamiento de datos
# - Combinar datos de múltiples recursos
# # 1. Creación y Carga de Datos
# ---------------------------------
# pandas puede leer archivos de muchos tipos, csv, json, excel entre otros
# <img src='./img/pandas_resources.PNG'>
# ### Creación de dataframes
#
# A partir de datos almacenados en diccionarios o listas es posible crear dataframes
# +
dicx= {
"nombre": ["Rick", "Morty"],
"apellido": ["Sanchez", "Smith"],
"edad": [60, 14],
}
rick_morty = pd.DataFrame(dicx)
rick_morty
# +
lista = [["Rick", "Sanchez", 60],
["Morty", "Smith", 14]]
columnas= ["nombre", "apellido", "edad"]
df_rick_morty = pd.DataFrame(lista, columns = columnas)
df_rick_morty
# -
type(df_rick_morty.nombre)
# ### Carga de datos a partir de fuentes de información
#
# Pandas soporta múltiples fuentes de información entre los que estan csv , sql, json,excel, etc
df = pd.read_csv('./data/primary_results.csv')
df_tabla_muestra = pd.read_clipboard()
df_tabla_muestra.head()
# https://e-consulta.sunat.gob.pe/cl-at-ittipcam/tcS01Alias
df_sunat = pd.read_clipboard()
df_sunat.head()
# por convención cuando se analiza un solo dataframe se suele llamar df
df = votos_primarias_us
df
# # Exploración
# -----------------------------
# `shape` nos devuelve el número de filas y columnas
df.shape
# `head` retorna los primertos 5 resultados contenidos en el dataframe (df)
# head retorna los primeros 5 resultados del dataframe
df.head(10)
# `tail` retorna los 5 últimos resultados contenidos en el dataframe (df)
# tail -> retorna los últimos 5 resultados del df
df.tail()
df.dtypes
# Describe -> nos brinda un resumen de la cantidad de datos, promedio, desviación estandar, minimo, máximo, etc
# de los datos de las columnas posibles
df.describe()
# # Seleccion
# ----------------------------
df.head()
# La columna a la izquierda del state es el index. Un dataframe tiene que tener un index, que es la manera de organizar los datos.
df.index
#
# ### Seleccion de Columnas
df.columns # nos brinda los nombres de columna contenidas en la tabla
# Seleccionamos una columna mediante '[]' como si el dataframe fuese un diccionario
# Seleccion de una única columna
df['county'].head(5)
# Seleccion de más de una columna
columnas = ['state','state_abbreviation']
df[columnas].head()
df["state"][:100]
# Tambien podemos seleccionar una columna mediante '.'
df.state_abbreviation.head()
# ### Seleccion de Filas
# podemos seleccionar una fila mediante su index.
df.loc[0]
# Importante, df.loc selecciona por indice, no por posición. Podemos cambiar el indice a cualquier otra cosa, otra columna o una lista separada, siempre que el nuevo indice tenga la misma longitud que el Dataframe
df2 = df.set_index("county")
df2.head()
df2.index
# Esto va a fallar por que df2 no tiene un indice numérico.
df2.loc[0]
# Ahora podemos seleccionar por condado
df2.loc["Los Angeles"]
# Si queremos seleccionar por el numero de fila independientemente del índice, podemos usar `iloc`
df2.iloc[0]
df2 = df2.reset_index(drop=True)
df2.head()
# # Filtrando Información
# ------------------------------
# Podemos filtrar un dataframe de la misma forma que filtramos en numpy
df[df['votes']>=590502]
# podemos concatenar varias condiciones usando `&`
df[(df.county=="Manhattan") & (df.party=="Democrat")]
# alternativamente podemos usar el método `query`
df.query("county=='Manhattan' and party=='Democrat'")
county = 'Manhattan'
df.query("county==@county and party=='Democrat'")
# # Procesado
# ------------------------------
# podemos usar `sort_values` para orderar el dataframe acorde al valor de una columna
df_sorted = df.sort_values(by="votes", ascending=False)
df_sorted.head()
df.groupby(["state", "party"])
df.groupby(["state", "party"])["votes"].sum()
# podemos usar `apply` en una columna para obtener una nueva columna en función de sus valores
df['letra_inicial'] = df.state_abbreviation.apply(lambda s: s[0])
df.groupby("letra_inicial")["votes"].sum().sort_values()
# Podemos unir dos dataframes en funcion de sus columnas comunes usando `merge`
# Descargamos datos de pobreza por condado en US en https://www.ers.usda.gov/data-products/county-level-data-sets/county-level-data-sets-download-data/
df_pobreza = pd.read_csv("./data/PovertyEstimates.csv")
df_pobreza.head()
df = df.merge(df_pobreza, left_on="fips", right_on="FIPStxt")
df.head()
county_votes = df.groupby(["county","party"]).agg({
"fraction_votes":"mean",
"PCTPOVALL_2015": "mean"
}
)
county_votes
# # Exportar
# ----------------------------------
# podemos escribir a excel, necesitamos instalar el paquete `xlwt`
# <img src='https://pandas.pydata.org/docs/_images/02_io_readwrite1.svg'>
rick_morty.to_excel("rick_y_morty.xls", sheet_name="personajes")
rick_morty.to_excel('rick_y_morty.xlsx',sheet_name="personajes",index=False)
rick_morty.to_csv('rick_y_morty.csv',sep='|',encoding='utf-8',index=False)
# podemos leer de excel, necesitamos el paquete `xlrd`
rick_morty2 = pd.read_excel("./rick_y_morty.xls", sheet_name="personajes")
rick_morty2.head()
| Modulo4/1.1 Intro a Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2 - Learning Phrase Representations using RNN Encoder-Decoder for Statistical Machine Translation
#
# In this second notebook on sequence-to-sequence models using PyTorch and TorchText, we'll be implementing the model from [Learning Phrase Representations using RNN Encoder-Decoder for Statistical Machine Translation](https://arxiv.org/abs/1406.1078). This model will achieve improved test perplexity whilst only using a single layer RNN in both the encoder and the decoder.
#
# ## Introduction
#
# Let's remind ourselves of the general encoder-decoder model.
#
# 
#
# We use our encoder (green) over the source sequence to create a context vector (red). We then use that context vector with the decoder (blue) and a linear layer (purple) to generate the target sentence.
#
# In the previous model, we used an multi-layered LSTM as the encoder and decoder.
#
# 
#
# One downside of the previous model is that the decoder is trying to cram lots of information into the hidden states. Whilst decoding, the hidden state will need to contain information about the whole of the source sequence, as well as all of the tokens have been decoded so far. By alleviating some of this information compression, we can create a better model!
#
# We'll also be using a GRU (Gated Recurrent Unit) instead of an LSTM (Long Short-Term Memory). Why? Mainly because that's what they did in the paper (this paper also introduced GRUs) and also because we used LSTMs last time. If you want to understand how GRUs (and LSTMs) differ from standard RNNS, check out [this](https://colah.github.io/posts/2015-08-Understanding-LSTMs/) link. Is a GRU better than an LSTM? [Research](https://arxiv.org/abs/1412.3555) has shown they're pretty much the same, and both are better than standard RNNs.
#
# ## Preparing Data
#
# All of the data preparation will be (almost) the same as last time, so I'll very briefly detail what each code block does. See the previous notebook if you've forgotten.
#
# We'll import PyTorch, TorchText, spaCy and a few standard modules.
# +
import torch
import torch.nn as nn
import torch.optim as optim
from torchtext.datasets import TranslationDataset, Multi30k
from torchtext.data import Field, BucketIterator
import spacy
import random
import math
import time
# -
# Then set a random seed for deterministic results/reproducability.
# +
SEED = 1234
random.seed(SEED)
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
# -
# Instantiate our German and English spaCy models.
spacy_de = spacy.load('de')
spacy_en = spacy.load('en')
# Previously we reversed the source (German) sentence, however in the paper we are implementing they don't do this, so neither will we.
# +
def tokenize_de(text):
"""
Tokenizes German text from a string into a list of strings
"""
return [tok.text for tok in spacy_de.tokenizer(text)]
def tokenize_en(text):
"""
Tokenizes English text from a string into a list of strings
"""
return [tok.text for tok in spacy_en.tokenizer(text)]
# -
# Create our fields to process our data. This will append the "start of sentence" and "end of sentence" tokens as well as converting all words to lowercase.
# +
SRC = Field(tokenize=tokenize_de,
init_token='<sos>',
eos_token='<eos>',
lower=True)
TRG = Field(tokenize = tokenize_en,
init_token='<sos>',
eos_token='<<PASSWORD>>',
lower=True)
# -
# Load our data.
train_data, valid_data, test_data = Multi30k.splits(exts = ('.de', '.en'),
fields = (SRC, TRG))
# We'll also print out an example just to double check they're not reversed.
print(vars(train_data.examples[0]))
# Then create our vocabulary, converting all tokens appearing less than twice into `<unk>` tokens.
SRC.build_vocab(train_data, min_freq = 2)
TRG.build_vocab(train_data, min_freq = 2)
# Finally, define the `device` and create our iterators.
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# +
BATCH_SIZE = 128
train_iterator, valid_iterator, test_iterator = BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE,
device = device)
# -
# ## Building the Seq2Seq Model
#
# ### Encoder
#
# The encoder is similar to the previous one, with the multi-layer LSTM swapped for a single-layer GRU. We also don't pass the dropout as an argument to the GRU as that dropout is used between each layer of a multi-layered RNN. As we only have a single layer, PyTorch will display a warning if we try and use pass a dropout value to it.
#
# Another thing to note about the GRU is that it only requires and returns a hidden state, there is no cell state like in the LSTM.
#
# $$\begin{align*}
# h_t &= \text{GRU}(x_t, h_{t-1})\\
# (h_t, c_t) &= \text{LSTM}(x_t, (h_{t-1}, c_{t-1}))\\
# h_t &= \text{RNN}(x_t, h_{t-1})
# \end{align*}$$
#
# From the equations above, it looks like the RNN and the GRU are identical. Inside the GRU, however, is a number of *gating mechanisms* that control the information flow in to and out of the hidden state (similar to an LSTM). Again, for more info, check out [this](https://colah.github.io/posts/2015-08-Understanding-LSTMs/) excellent post.
#
# The rest of the encoder should be very familar from the last tutorial, it takes in a sequence, $X = \{x_1, x_2, ... , x_T\}$, recurrently calculates hidden states, $H = \{h_1, h_2, ..., h_T\}$, and returns a context vector (the final hidden state), $z=h_T$.
#
# $$h_t = \text{EncoderGRU}(x_t, h_{t-1})$$
#
# This is identical to the encoder of the general seq2seq model, with all the "magic" happening inside the GRU (green squares).
#
# 
class Encoder(nn.Module):
def __init__(self, input_dim, emb_dim, hid_dim, dropout):
super().__init__()
self.hid_dim = hid_dim
self.embedding = nn.Embedding(input_dim, emb_dim) #no dropout as only one layer!
self.rnn = nn.GRU(emb_dim, hid_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, src):
#src = [src sent len, batch size]
embedded = self.dropout(self.embedding(src))
#embedded = [src sent len, batch size, emb dim]
outputs, hidden = self.rnn(embedded) #no cell state!
#outputs = [src sent len, batch size, hid dim * n directions]
#hidden = [n layers * n directions, batch size, hid dim]
#outputs are always from the top hidden layer
return hidden
# ## Decoder
#
# The decoder is where the implementation differs significantly from the previous model and we alleviate some of the information compression.
#
# Instead of the GRU in the decoder taking just the target token, $y_t$ and the previous hidden state $s_{t-1}$ as inputs, it also takes the context vector $z$.
#
# $$s_t = \text{DecoderGRU}(y_t, s_{t-1}, z)$$
#
# Note how this context vector, $z$, does not have a $t$ subscript, meaning we re-use the same context vector returned by the encoder for every time-step in the decoder.
#
# Before, we predicted the next token, $\hat{y}_{t+1}$, with the linear layer, $f$, only using the top-layer decoder hidden state at that time-step, $s_t$, as $\hat{y}_{t+1}=f(s_t^L)$. Now, we also pass the current token, $\hat{y}_t$ and the context vector, $z$ to the linear layer.
#
# $$\hat{y}_{t+1} = f(y_t, s_t, z)$$
#
# Thus, our decoder now looks something like this:
#
# 
#
# Note, the initial hidden state, $s_0$, is still the context vector, $z$, so when generating the first token we are actually inputting two identical context vectors into the GRU.
#
# How do these two changes reduce the information compression? Well, hypothetically the decoder hidden states, $s_t$, no longer need to contain information about the source sequence as it is always available as an input. Thus, it only needs to contain information about what tokens it has generated so far. The addition of $y_t$ to the linear layer also means this layer can directly see what the token is, without having to get this information from the hidden state.
#
# However, this hypothesis is just a hypothesis, it is impossible to determine how the model actually uses the information provided to it (don't listen to anyone that tells you differently). Nevertheless, it is a solid intuition and the results seem to indicate that this modifications are a good idea!
#
# Within the implementation, we will pass $y_t$ and $z$ to the GRU by concatenating them together, so the input dimensions to the GRU are now `emb_dim + hid_dim` (as context vector will be of size `hid_dim`). The linear layer will take $y_t, s_t$ and $z$ also by concatenating them together, hence the input dimensions are now `emb_dim + hid_dim*2`. We also don't pass a value of dropout to the GRU as it only uses a single layer.
#
# `forward` now takes a `context` argument. Inside of `forward`, we concatenate $y_t$ and $z$ as `emb_con` before feeding to the GRU, and we concatenate $y_t$, $s_t$ and $z$ together as `output` before feeding it through the linear layer to receive our predictions, $\hat{y}_{t+1}$.
class Decoder(nn.Module):
def __init__(self, output_dim, emb_dim, hid_dim, dropout):
super().__init__()
self.hid_dim = hid_dim
self.output_dim = output_dim
self.embedding = nn.Embedding(output_dim, emb_dim)
self.rnn = nn.GRU(emb_dim + hid_dim, hid_dim)
self.out = nn.Linear(emb_dim + hid_dim * 2, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, input, hidden, context):
#input = [batch size]
#hidden = [n layers * n directions, batch size, hid dim]
#context = [n layers * n directions, batch size, hid dim]
#n layers and n directions in the decoder will both always be 1, therefore:
#hidden = [1, batch size, hid dim]
#context = [1, batch size, hid dim]
input = input.unsqueeze(0)
#input = [1, batch size]
embedded = self.dropout(self.embedding(input))
#embedded = [1, batch size, emb dim]
emb_con = torch.cat((embedded, context), dim = 2)
#emb_con = [1, batch size, emb dim + hid dim]
output, hidden = self.rnn(emb_con, hidden)
#output = [sent len, batch size, hid dim * n directions]
#hidden = [n layers * n directions, batch size, hid dim]
#sent len, n layers and n directions will always be 1 in the decoder, therefore:
#output = [1, batch size, hid dim]
#hidden = [1, batch size, hid dim]
output = torch.cat((embedded.squeeze(0), hidden.squeeze(0), context.squeeze(0)),
dim = 1)
#output = [batch size, emb dim + hid dim * 2]
prediction = self.out(output)
#prediction = [batch size, output dim]
return prediction, hidden
# ## Seq2Seq Model
#
# Putting the encoder and decoder together, we get:
#
# 
#
# Again, in this implementation we need to ensure the hidden dimensions in both the encoder and the decoder are the same.
#
# Briefly going over all of the steps:
# - the `outputs` tensor is created to hold all predictions, $\hat{Y}$
# - the source sequence, $X$, is fed into the encoder to receive a `context` vector
# - the initial decoder hidden state is set to be the `context` vector, $s_0 = z = h_T$
# - we use a batch of `<sos>` tokens as the first `input`, $y_1$
# - we then decode within a loop:
# - inserting the input token $y_t$, previous hidden state, $s_{t-1}$, and the context vector, $z$, into the decoder
# - receiving a prediction, $\hat{y}_{t+1}$, and a new hidden state, $s_t$
# - we then decide if we are going to teacher force or not, setting the next input as appropriate (either the ground truth next token in the target sequence or the highest predicted next token)
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder, device):
super().__init__()
self.encoder = encoder
self.decoder = decoder
self.device = device
assert encoder.hid_dim == decoder.hid_dim, \
"Hidden dimensions of encoder and decoder must be equal!"
def forward(self, src, trg, teacher_forcing_ratio = 0.5):
#src = [src sent len, batch size]
#trg = [trg sent len, batch size]
#teacher_forcing_ratio is probability to use teacher forcing
#e.g. if teacher_forcing_ratio is 0.75 we use ground-truth inputs 75% of the time
batch_size = trg.shape[1]
max_len = trg.shape[0]
trg_vocab_size = self.decoder.output_dim
#tensor to store decoder outputs
outputs = torch.zeros(max_len, batch_size, trg_vocab_size).to(self.device)
#last hidden state of the encoder is the context
context = self.encoder(src)
#context also used as the initial hidden state of the decoder
hidden = context
#first input to the decoder is the <sos> tokens
input = trg[0,:]
for t in range(1, max_len):
#insert input token embedding, previous hidden state and the context state
#receive output tensor (predictions) and new hidden state
output, hidden = self.decoder(input, hidden, context)
#place predictions in a tensor holding predictions for each token
outputs[t] = output
#decide if we are going to use teacher forcing or not
teacher_force = random.random() < teacher_forcing_ratio
#get the highest predicted token from our predictions
top1 = output.argmax(1)
#if teacher forcing, use actual next token as next input
#if not, use predicted token
input = trg[t] if teacher_force else top1
return outputs
# # Training the Seq2Seq Model
#
# The rest of this tutorial is very similar to the previous one.
#
# We initialise our encoder, decoder and seq2seq model (placing it on the GPU if we have one). As before, the embedding dimensions and the amount of dropout used can be different between the encoder and the decoder, but the hidden dimensions must remain the same.
# +
INPUT_DIM = len(SRC.vocab)
OUTPUT_DIM = len(TRG.vocab)
ENC_EMB_DIM = 256
DEC_EMB_DIM = 256
HID_DIM = 512
ENC_DROPOUT = 0.5
DEC_DROPOUT = 0.5
enc = Encoder(INPUT_DIM, ENC_EMB_DIM, HID_DIM, ENC_DROPOUT)
dec = Decoder(OUTPUT_DIM, DEC_EMB_DIM, HID_DIM, DEC_DROPOUT)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model = Seq2Seq(enc, dec, device).to(device)
# -
# Next, we initialize our parameters. The paper states the parameters are initialized from a normal distribution with a mean of 0 and a standard deviation of 0.01, i.e. $\mathcal{N}(0, 0.01)$.
#
# It also states we should initialize the recurrent parameters to a special initialization, however to keep things simple we'll also initialize them to $\mathcal{N}(0, 0.01)$.
# +
def init_weights(m):
for name, param in m.named_parameters():
nn.init.normal_(param.data, mean=0, std=0.01)
model.apply(init_weights)
# -
# We print out the number of parameters.
#
# Even though we only have a single layer RNN for our encoder and decoder we actually have **more** parameters than the last model. This is due to the increased size of the inputs to the GRU and the linear layer. However, it is not a significant amount of parameters and causes a minimal amount of increase in training time (~3 seconds per epoch extra).
# +
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
# -
# We initiaize our optimizer.
optimizer = optim.Adam(model.parameters())
# We also initialize the loss function, making sure to ignore the loss on `<pad>` tokens.
# +
PAD_IDX = TRG.vocab.stoi['<pad>']
criterion = nn.CrossEntropyLoss(ignore_index = PAD_IDX)
# -
# We then create the training loop...
def train(model, iterator, optimizer, criterion, clip):
model.train()
epoch_loss = 0
for i, batch in enumerate(iterator):
src = batch.src
trg = batch.trg
optimizer.zero_grad()
output = model(src, trg)
#trg = [trg sent len, batch size]
#output = [trg sent len, batch size, output dim]
output = output[1:].view(-1, output.shape[-1])
trg = trg[1:].view(-1)
#trg = [(trg sent len - 1) * batch size]
#output = [(trg sent len - 1) * batch size, output dim]
loss = criterion(output, trg)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), clip)
optimizer.step()
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# ...and the evaluation loop, remembering to set the model to `eval` mode and turn off teaching forcing.
def evaluate(model, iterator, criterion):
model.eval()
epoch_loss = 0
with torch.no_grad():
for i, batch in enumerate(iterator):
src = batch.src
trg = batch.trg
output = model(src, trg, 0) #turn off teacher forcing
#trg = [trg sent len, batch size]
#output = [trg sent len, batch size, output dim]
output = output[1:].view(-1, output.shape[-1])
trg = trg[1:].view(-1)
#trg = [(trg sent len - 1) * batch size]
#output = [(trg sent len - 1) * batch size, output dim]
loss = criterion(output, trg)
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# We'll also define the function that calculates how long an epoch takes.
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# Then, we train our model, saving the parameters that give us the best validation loss.
# +
N_EPOCHS = 10
CLIP = 1
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_iterator, optimizer, criterion, CLIP)
valid_loss = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut2-model.pt')
print(f'Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train PPL: {math.exp(train_loss):7.3f}')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. PPL: {math.exp(valid_loss):7.3f}')
# -
# Finally, we test the model on the test set using these "best" parameters.
# +
model.load_state_dict(torch.load('tut2-model.pt'))
test_loss = evaluate(model, test_iterator, criterion)
print(f'| Test Loss: {test_loss:.3f} | Test PPL: {math.exp(test_loss):7.3f} |')
# -
# Just looking at the test loss, we get better performance. This is a pretty good sign that this model architecture is doing something right! Relieving the information compression seems like the way forard, and in the next tutorial we'll expand on this even further with *attention*.
| 2 - Learning Phrase Representations using RNN Encoder-Decoder for Statistical Machine Translation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Imports
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# Local
import Neuron
import models as models
import train as train
import batch_utils
import data_transforms
import generate_training_data
# -
# ## Data
training_data = generate_training_data.y_shape(n_nodes=20,
data_size=1000,
first_length=10,
branching_node=6)
# ## Global parameters
# +
n_nodes = 20
input_dim = 100
n_epochs = 5
batch_size = 32
n_batch_per_epoch = np.floor(training_data['morphology']['n20'].shape[0]/batch_size).astype(int)
d_iters = 20
lr_discriminator = 0.001
lr_generator = 0.001
train_loss = 'binary_crossentropy'
#train_loss = 'wasserstein_loss'
rule = 'none'
d_weight_constraint = [-.03, .03]
g_weight_constraint = [-33.3, 33.3]
m_weight_constraint = [-33.3, 33.3]
# -
# # Run
geom_model, morph_model, disc_model, gan_model = \
train.train_model(training_data=training_data,
n_nodes=n_nodes,
input_dim=input_dim,
n_epochs=n_epochs,
batch_size=batch_size,
n_batch_per_epoch=n_batch_per_epoch,
d_iters=d_iters,
lr_discriminator=lr_discriminator,
lr_generator=lr_generator,
d_weight_constraint=d_weight_constraint,
g_weight_constraint=g_weight_constraint,
m_weight_constraint=m_weight_constraint,
rule=rule,
train_loss=train_loss,
verbose=True)
| BonsaiNet/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from nltk.tokenize import word_tokenize
ref_summary_path='../yelp_dataset/yelp_reference_summary.csv'
ref_summary=pd.read_csv(ref_summary_path)
cont=ref_summary.iloc[:,[1,2,3,4,5,6,7,8,9]]
stats=cont.applymap(word_tokenize).applymap(len)
stats['mean']=stats.iloc[:,0:7].apply(np.mean,axis=1)
stats['max']=stats.iloc[:,0:7].apply(np.max,axis=1)
stats['min']=stats.iloc[:,0:7].apply(np.min,axis=1)
stats['diff']=stats['mean']-stats['Answer.summary']
stats
abs(stats['diff']).mean()
# ## Meansum_hotel
hotel_summary_path='../../outputs/eval/hotel/n_docs_8/unsup_ndoc=8_fullText/summary_first_100.csv'
hotel_doc_path='../../outputs/eval/hotel/n_docs_8/unsup_ndoc=8_fullText/reviews_first_100.csv'
hotel_summary=pd.read_csv(hotel_summary_path,index_col=0)
hotel_doc=pd.read_csv(hotel_doc_path,index_col=0)
hotel_summary=pd.concat([hotel_summary,hotel_doc],axis=1)
hotel_summary['docs']=hotel_summary['docs'].apply(lambda x:x.split('</DOC>'))
# +
def docs_decompose(hotel_summary,num):
hotel_summary['doc'+str(num)]=hotel_summary['docs'].apply(lambda x:x[num])
return 0
for i in range(8):
docs_decompose(hotel_summary,i)
hotel_summary.drop(columns=['docs'],inplace=True)
hotel_summary_stats=hotel_summary.applymap(word_tokenize).applymap(len)
# -
hotel_summary_stats
hotel_summary_stats['mean']=hotel_summary_stats.iloc[:,1:8].apply(np.mean,axis=1)
hotel_summary_stats['max']=hotel_summary_stats.iloc[:,1:8].apply(np.max,axis=1)
hotel_summary_stats['min']=hotel_summary_stats.iloc[:,1:8].apply(np.min,axis=1)
hotel_summary_stats['diff']=hotel_summary_stats['mean']-hotel_summary_stats['summary']
hotel_summary_stats
abs(hotel_summary_stats['diff']).mean()
# ## Meansum_hotel_mask
hotel_mask_summary_path='../../outputs/eval/hotel_mask/n_docs_8/unsup_hotel_mask/summaries.json'
hotel_mask_summary=pd.read_json(hotel_mask_summary_path)[['docs','summary']].head(200)
hotel_mask_summary['docs']=hotel_mask_summary['docs'].apply(lambda x:x.split('</DOC>'))
for i in range(8):
docs_decompose(hotel_mask_summary,i)
# +
hotel_mask_summary.drop(columns=['docs'],inplace=True)
hotel_mask_summary_stats=hotel_mask_summary.applymap(word_tokenize).applymap(len)
# -
hotel_mask_summary_stats['mean']=hotel_mask_summary_stats.iloc[:,1:8].apply(np.mean,axis=1)
hotel_mask_summary_stats['max']=hotel_mask_summary_stats.iloc[:,1:8].apply(np.max,axis=1)
hotel_mask_summary_stats['min']=hotel_mask_summary_stats.iloc[:,1:8].apply(np.min,axis=1)
hotel_mask_summary_stats['diff']=hotel_mask_summary_stats['mean']-hotel_mask_summary_stats['summary']
abs(hotel_mask_summary_stats['diff']).mean()
hotel_mask_summary_stats.head(20)
hotel_mask_summary.iloc[8,0]
| datasets/original_csv_data/length_mask_review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit
# name: python39164bitcbdf1bbf6d0547a2abdb665663ed1f56
# ---
# # Build a cuisine recommender
# + tags=[]
import pandas as pd
# -
data = pd.read_csv('/Users/robbiewoolterton/Documents/GitHub/ML-For-Beginners/4-Classification/data/cleaned_cuisine.csv')
data.head()
X = data.iloc[:,2:]
X.head()
y = data[['cuisine']]
y.head()
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.model_selection import cross_val_score
from sklearn.metrics import accuracy_score,precision_score,confusion_matrix,classification_report
# + tags=[]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
# -
model = SVC(kernel='linear', C=10, probability=True, random_state=0)
model.fit(X_train, y_train.values.ravel())
y_pred = model.predict(X_test)
# + tags=[]
print(classification_report(y_test, y_pred))
# +
from skl2onnx import convert_sklearn
from skl2onnx.common.data_types import FloatTensorType
initial_type = [('float_input', FloatTensorType([None, 380]))]
options = {id(model): {'nocl': True, 'zipmap': False}}
# -
onx = convert_sklearn(model, initial_types=initial_type, options=options)
with open('./web-app/model.onx', 'wb') as f:
f.write(onx.SerializeToString())
| 4-Classification/4-Applied/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div align="right" style="text-align: right"><i><NAME>, Feb 2021</i></div>
#
# # CrossProduct Puzzle
#
# The 538 Riddler [poses a type of puzzle](https://fivethirtyeight.com/features/can-you-cross-like-a-boss/) called ***CrossProduct***, which works like this:
#
# >*Replace each "?" in the table with a single digit so that the product of the digits in each row equals the number to the right of the row, and the product of the digits in each column equals the number above the column.*
#
#
# Sample puzzle:
#
#
# | 6615 | 15552 | 420 | [6×3] |
# |-------|-------|-------|---|
# | ? | ? | ? |**210**|
# | ? | ? | ? |**144**|
# | ? | ? | ? |**54**|
# | ? | ? | ? |**135**|
# | ? | ? | ? |**4**|
# | ? | ? | ? |**49**|
#
#
# Solution:
#
# |6615|15552| 420| [6×3]|
# |---|---|---|---|
# |7|6|5|**210**|
# |9|8|2|**144**|
# |3|9|2|**54**|
# |5|9|3|**135**|
# |1|4|1|**4**|
# |7|1|7|**49**|
#
#
#
#
#
#
# We could solve CrossProduct puzzles by hand, but why not write a program to do it?
#
# # Data type definitions
#
# Here are the data types we will use in trying to solve CrossProduct puzzles:
# - `Digit`: a single digit, from 1 to 9 (but not 0).
# - `Row`: a tuple of digits that forms a row in the table, e.g. `(7, 6, 5)`.
# - `Table`: a table of digits that fill in all the "?"s; a list of rows, e.g. `[(7, 6, 5), (9, 8, 2), ...]`.
# - `Products`: a list of the numbers that corresponding digits must multiply to, e.g. in the puzzle above:
# - `[6615, 15552, 420]` for the column products;
# - `[210, 144, 54, 135, 4, 49]` for the row products.
# - `Puzzle`: a puzzle to be solved, as defined by the row products and column products.
# +
from typing import Tuple, List, Set, Iterable, Optional
from numpy import divide, prod, transpose
from collections import namedtuple
import random
Digit = int
Row = Tuple[Digit, ...]
Table = List[Row]
Product = int
Products = List[Product]
Puzzle = namedtuple('Puzzle', 'row_prods, col_prods')
# -
# # The puzzles
#
# Here are the puzzles given by 538 Riddler (they promised one a week for four weeks; so far we've seen three):
puzzles = (
Puzzle([135, 45, 64, 280, 70], [3000, 3969, 640]),
Puzzle([210, 144, 54, 135, 4, 49], [6615, 15552, 420]),
Puzzle([280, 168, 162, 360, 60, 256, 126], [183708, 245760, 117600]))
# # Strategy
#
# Here's my strategy:
# - To solve a puzzle, first find all ways to fill the first row, and for each way, solve the rest of the puzzle.
# - To fill a row, first find all ways to fill the first digit, and for each way, fill the rest of the row.
#
# So the first step is to define `fill_one_row(row_prod, col_prods)` to return a set of digit-tuples that can legally fill a row that has the given row product in a puzzle with the given column products.
# - If `col_prods` is empty, then there is one solution (the 0-length tuple) if `row_prod` is 1, and no solution otherwise.
# - Otherwise, try each digit `d` that divides both the `row_prod` and the first number in `col_prods`, and then try all ways to fill the rest of the row.
def fill_one_row(row_prod: Product, col_prods: Products) -> Set[Row]:
"All permutations of digits that multiply to `row_prod` and evenly divide `col_prods`."
if not col_prods:
return {()} if row_prod == 1 else set()
else:
return {(d, *rest) for d in range(1, 10)
if (row_prod / d).is_integer() and (col_prods[0] / d).is_integer()
for rest in fill_one_row(row_prod // d, col_prods[1:])}
# Some examples:
fill_one_row(210, [6615, 15552, 420]) # There are 2 ways to fill this row
fill_one_row(54, [6615, 15552, 420]) # There are 8 ways to fill this row
# Now we can solve the rest of a puzzle:
#
# - `solve(puzzle)` finds the first solution. (A well-formed puzzle has exactly one solution, but some might have more, or none.)
# - `solutions(puzzle)` yields all possible solutions to a puzzle. There are three main cases to consider:
# - A puzzle with no rows has the empty table, `[]`, as a solution, as long as the column products are all 1.
# - A puzzle with rows might have solutions, as long as the column products are all integers. Call `fill_row` to get all possible ways to fill the first row, and for each one recursively call `solutions` to get all the possible ways of filling the rest of the rows (making sure to pass in an altered `col_prods` where each element is divided by the corresponding element in the first row).
# - Otherwise there are no solutions.
# +
def solve(puzzle) -> Optional[Table]: return next(solutions(puzzle), None)
def solutions(puzzle) -> Iterable[Table]:
"""Yield all tables that solve the puzzle.
The product of the digits in row r must equal row_prods[r], for all r.
The product of the digits in column c must equal col_prods[c], for all c."""
row_prods, col_prods = puzzle
if not row_prods and all(c == 1 for c in col_prods):
yield []
elif row_prods and all(c == int(c) for c in col_prods):
yield from ([row1, *rows]
for row1 in fill_one_row(row_prods[0], col_prods)
for rows in solutions(Puzzle(row_prods[1:], list(divide(col_prods, row1)))))
# -
# # Solutions
#
# Here are solutions to the three puzzles posed by *The Riddler*:
[solve(p) for p in puzzles]
# Those are the correct solutions. However, we could make them look nicer.
#
# # Prettier solutions
# +
from IPython.display import Markdown, display
def pretty(puzzle) -> Markdown:
"""A puzzle and its solution in pretty Markdown format."""
row_prods, col_prods = puzzle
head = row(col_prods + [f'[{len(row_prods)}×{len(col_prods)}]'])
dash = row(['---'] * (1 + len(col_prods)))
rest = [row(r + (f'**{rp}**',))
for r, rp in zip(solve(puzzle), row_prods)]
return Markdown('\n'.join([head, dash, *rest]))
def row(items) -> str:
"""Make a markdown table row."""
return '|' + '|'.join(map(str, items)) + '|'
# -
for p in puzzles:
display(pretty(p))
# # Making new puzzles
#
# Can we make new puzzles? Can we make well-formed ones (those with exactly one solution)? Here is an approach:
# - Make a table filled with random digits (`random_table`).
# - Make a puzzle from the row and column products of the table (`table_puzzle`).
# - Repeat `N` times (`random_puzzles`).
# - Optionally, check if puzzles are `well-formed`.
#
# +
def random_table(nrows, ncols) -> Table:
"Make a table of random digits of the given size."
return [tuple(random.randint(1, 9) for c in range(ncols))
for r in range(nrows)]
def table_puzzle(table) -> Puzzle:
"Given a table, compute the puzzle it is a solution for."
return Puzzle([prod(row) for row in table],
[prod(col) for col in transpose(table)])
def random_puzzles(N, nrows, ncols, seed=42) -> List[Puzzle]:
"Return a list of `N` random puzzles."
random.seed(seed) # For reproducability
return [table_puzzle(random_table(nrows, ncols)) for _ in range(N)]
def well_formed(puzzle) -> bool:
"Does the puzzle have exactly one solution?"
S = solutions(puzzle)
first, second = next(S, None), next(S, None)
return first is not None and second is None
# -
random_table(nrows=5, ncols=3)
puz = table_puzzle(_)
well_formed(puz)
len(list(solutions(puz)))
pretty(puz)
# How likely are random puzzles (of various sizes) to be well-formed?
N = 200
for r, c in [(3, 3), (3, 4), (4, 3), (3, 5), (5, 3), (4, 4), (6, 3)]:
w = sum(map(well_formed, random_puzzles(N, r, c))) / N
print(f'{w:3.0%} of random puzzles with {r} rows and {c} cols ({r * c:2} cells) are well-formed')
# We see that most puzzles are not well-formed. Smaller sizes are more likely to yield well-formed puzzles.
#
# # Speed
#
# How long does it take to solve random puzzles? We can do a thousand small (5x3) puzzles in about two seconds:
# %time all(solve(p) for p in random_puzzles(1000, 5, 3))
# Puzzles that are even a little bit larger can be a lot slower, and there is huge variability in the time to solve. For example, a single 10 x 6 puzzle can take from a few milliseconds to tens of seconds:
[p10x6] = random_puzzles(1, 10, 6)
# %time pretty(p10x6)
# In general, the time to solve a puzzle can grow exponentially in the number of cells. Consider a row in a six-column puzzle, where the products are all 5040. There are 3,960 ways to fill this row:
n = 5040
len(fill_one_row(n, [n] * 6))
# If four rows all had a similar number of possibilities and didn't constrain each other, that would be hundreds of trillions of combinations to try—an infeasible number. We will need a faster algorithm for larger puzzles.
#
# # Faster Speed
#
# To speed things up, we could encode the puzzle as a constraint satisfaction problem (CSP), and use a highly-optimized [CSP solver](https://developers.google.com/optimization/cp/cp_solver). But even without going to a professional-grade CSP solver, we could borrow the heuristics they use. There are four main considerations in CSP solving:
# - **Variable definition**: In `solutions`, we are treating each **row** as a variable, and asking "which of the possible values returned by `fill_one_row` will work as the value of this row? An alternative would be to treat each **cell** as a variable, and fill in the puzzle one cell at a time rather than one row at a time. This has the advantage that each variable has only 9 possible values, not thousands of possibilities.
# - **Variable ordering**: In `solutions`, we consider the variables (the rows) in strict top-to-bottom order. It is usually more efficient to reorder the variables, filling in first the variable with the minimum number of possible values. The reasoning is that if you have a variable with only 2 possibilities, you have a 50% chance of guessing right the first time, whereas if there were 100 possibilities, you have only a 1% chance of guessing right.
# - **Value ordering**: The function `fill_one_row` returns values in sorted lexicographic order, lowest first. We could reorder the values to pick the one that imposes the least constraints first (that is, the value that allows the most possibilities for the other variables).
# - **Domain-specific heuristics**: CSP solvers are general, but sometimes knowledge that is specific to a problem can be helpful. One fact about CrossProduct is that the digits 5 and 7 are special in the sense that if a row (or column) product is divisible by 5 (or 7), then the digit 5 (or 7) must appear in the row (or column). That is not true for the other digits (for example, if a row product is divisible by 8, then an 8 may appear in the row, or it might be a 2 and a 4, or three 6s, etc.).
#
# Usually variable ordering is the most productive heuristic. Let's try it. The function `reorder` takes a puzzle and returns a version of the puzzle with the row products permuted so that the rows with the fewest possible fillers come first:
def reorder(puzzle) -> Puzzle:
"""Create a version of puzzle with the rows reordered so the rows with the fewest
number of possible fillers come first."""
def fillers(r): return len(fill_one_row(r, puzzle.col_prods))
rows = sorted(puzzle.row_prods, key=fillers)
return Puzzle(rows, puzzle.col_prods)
p2 = puzzles[2]
p2, reorder(p2)
# How many ways are there to fill each row?
{r: len(fill_one_row(r, p2.col_prods))
for r in reorder(p2).row_prods}
# Now I'll define a set of test puzzles and see how long it takes to solve them all, and compare that to the time to solve the reordered versions:
test_puzzles = random_puzzles(20, 10, 3)
# %time all(solve(p) for p in test_puzzles)
# %time all(solve(reorder(p)) for p in test_puzzles)
# That's a nice improvement—150 times faster on this small test set! I'm curious whether we would get even more speedup by treating each cell as a separate variable, or by considering value ordering, but I'll leave that as an exercise for the reader.
# # Tests
#
# A suite of unit tests:
# +
def test():
"Test suite for CrossProduct functions."
assert fill_one_row(1, []) == {()}
assert fill_one_row(2, []) == set()
assert fill_one_row(9, [9]) == {(9,)}
assert fill_one_row(10, [10]) == set()
assert fill_one_row(73, [360, 360, 360]) == set()
assert solve(Puzzle([], [])) == []
assert solve(Puzzle([], [1])) == []
assert solve(Puzzle([], [2])) == None
assert solve(Puzzle([5], [5])) == [(5,)]
assert solve(Puzzle([0], [0])) == None # Maybe should allow zero as a digit?
assert solve(Puzzle([2, 12], [3, 8])) == [(1, 2), (3, 4)]
assert fill_one_row(729, [90, 126, 81]) == {(9, 9, 9)} # Unique fill
assert fill_one_row(729, [90, 126, 81, 30]) == {
(3, 9, 9, 3), (9, 3, 9, 3), (9, 9, 3, 3), (9, 9, 9, 1)}
# 72 has the most ways to fill a 3-digit row
assert max(range(1, 100), key=lambda n: len(fill_one_row(n, [5*7*8*9]*3))) == 72
assert fill_one_row(72, [72, 72, 72]) == {
(1, 8, 9),
(1, 9, 8),
(2, 4, 9),
(2, 6, 6),
(2, 9, 4),
(3, 3, 8),
(3, 4, 6),
(3, 6, 4),
(3, 8, 3),
(4, 2, 9),
(4, 3, 6),
(4, 6, 3),
(4, 9, 2),
(6, 2, 6),
(6, 3, 4),
(6, 4, 3),
(6, 6, 2),
(8, 1, 9),
(8, 3, 3),
(8, 9, 1),
(9, 1, 8),
(9, 2, 4),
(9, 4, 2),
(9, 8, 1)}
assert fill_one_row(7**8, [7]*9) == {
(1, 7, 7, 7, 7, 7, 7, 7, 7),
(7, 1, 7, 7, 7, 7, 7, 7, 7),
(7, 7, 1, 7, 7, 7, 7, 7, 7),
(7, 7, 7, 1, 7, 7, 7, 7, 7),
(7, 7, 7, 7, 1, 7, 7, 7, 7),
(7, 7, 7, 7, 7, 1, 7, 7, 7),
(7, 7, 7, 7, 7, 7, 1, 7, 7),
(7, 7, 7, 7, 7, 7, 7, 1, 7),
(7, 7, 7, 7, 7, 7, 7, 7, 1)}
assert solve(Puzzle([210, 144, 54, 135, 4, 49], [6615, 15552, 420])) == [
(7, 6, 5),
(9, 8, 2),
(3, 9, 2),
(5, 9, 3),
(1, 4, 1),
(7, 1, 7)]
assert sorted(solutions(Puzzle([8, 8, 1], [8, 8, 1]))) == [ # Multi-solution puzzle
[(1, 8, 1),
(8, 1, 1),
(1, 1, 1)],
[(2, 4, 1),
(4, 2, 1),
(1, 1, 1)],
[(4, 2, 1),
(2, 4, 1),
(1, 1, 1)],
[(8, 1, 1),
(1, 8, 1),
(1, 1, 1)]]
assert not list(solutions(Puzzle([8, 8, 1], [8, 8, 2]))) # Unsolvable puzzle
assert solve(Puzzle([1470, 720, 270, 945, 12, 343],
[6615, 15552, 420, 25725])) == [ # 4 column puzzle
(7, 6, 5, 7),
(9, 8, 2, 5),
(3, 9, 2, 5),
(5, 9, 3, 7),
(1, 4, 1, 3),
(7, 1, 7, 7)]
puzz = Puzzle([6, 120, 504], [28, 80, 162])
table = [(1, 2, 3),
(4, 5, 6),
(7, 8, 9)]
assert solve(puzz) == table
assert table_puzzle(table) == puzz
assert well_formed(puzz)
assert not well_formed(Puzzle([7, 7], [7, 7]))
assert well_formed(Puzzle([64, 224, 189, 270, 405, 144, 105],
[308700, 12960, 1119744]))
assert row((1, 2, 3)) == '|1|2|3|'
col_prods = [193536, 155520, 793800]
assert (reorder(Puzzle([10, 48, 36, 7, 32, 81, 252, 160, 21, 90], col_prods)) ==
Puzzle([ 7, 10, 160, 21, 81, 252, 90, 32, 48, 36], col_prods))
return True
test()
| ipynb/CrossProduct.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd # Required for data management
import numpy as np # Required for some computations
import matplotlib.pyplot as plt # Required for plotting
import datetime as DT #Required to create SerDates for my hourly data
# +
df = pd.read_csv('Woodlands_Piezometer.csv', header=0)
DateArray = df.as_matrix()
DateArray = DateArray[:,0:4].astype(int)
print(DateArray)
SerDates = [DT.datetime(*x) for x in DateArray]
df['SerDates'] = SerDates
#df
# -
# Use the pandas plotting function, specifying 'SerDates' (the serial datas) on the x axis and 'Q' (discharge)
# on the y axis. I'm also using the 'figsize' option to increase the size of the plotted image
df.plot(df['SerDates'].values, y='DTW_m',figsize=(14,10))
# Use the pandas .values operator to load the values of discharge (the 'Q' column)
# from the pandas dataframe into a numpy array
DTW = df['DTW_m'].values
# +
# Use numpy array indexing to get discharge from October 2, 2000 to September 30, 2015.
# The 'end' of a numpy array can be indexed using -1: as if the index in the negative direction
# wraps back around to the end of the record
DTWt = DTW[1:-1]
# Similarly get discharge from October 1, 2000 to September 29, 2015 by getting data from the
# beginning of the record to the 'end' minus 1 using the -2 as the index.
DTWtm1 = DTW[0:-2]
# +
# Use Matplotlib to create a scatter plot of these two time series, create a title and label axes
plt.figure(figsize=(14,10))
plt.plot(DTWtm1,DTWt,'o')
plt.title('How correlated is this hour\'s DTW with previous hour\'s?')
plt.xlabel('$DTW_{t-1}$ m')
plt.ylabel('$DTW_t$ m')
plt.show()
#5. Save the dataframe as a pandas "pickle" file¶
df.to_pickle('WoodlandsDTW_WY2013-2017.pkl')
# -
| Module1 (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: LSST SIMS MAF (Py3, w.2021.14)
# language: python
# name: lsst_sims_maf-w.2021.14
# ---
# Getting started using this brown dwarf metric notebook as a guide: https://github.com/yoachim/LSST-BD-Cadence/blob/main/bd_dist_metric.ipynb
# +
from lsst.sims.maf.metrics.baseMetric import BaseMetric
import lsst.sims.maf.utils as mafUtils
import lsst.sims.utils as utils
from scipy import interpolate
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import healpy as hp
import astropy.units as u
from astropy.io import ascii, fits
import lsst.sims.maf.db as db
import lsst.sims.maf.utils as utils
import lsst.sims.maf.metrics as metrics
import lsst.sims.maf.slicers as slicers
import lsst.sims.maf.stackers as stackers
import lsst.sims.maf.metricBundles as metricBundles
import lsst.sims.maf.maps as maps
# -
from mafContrib.lssmetrics import depthLimitedNumGalMetric
from mafContrib.LSSObsStrategy.galaxyCountsMetric_extended import GalaxyCountsMetric_extended \
as GalaxyCountsMetric
lv_dat0 = fits.getdata('lsst_galaxies_1p25to9Mpc_table.fits')
lv_dat_cuts = (lv_dat['dec'] < 35.0) & (lv_dat['MStars'] > 1e7)
lv_dat = lv_dat0[lv_dat_cuts]
#dbFile = '/sims_maf/fbs_1.7/baseline/baseline_nexp2_v1.7_10yrs.db'
#dbFile = '/sims_maf/fbs_1.7/baseline/baseline_nexp1_v1.7_10yrs.db'
#dbFile = '/sims_maf/fbs_1.7/footprint_tune/footprint_7_v1.710yrs.db'
dbFile = '/sims_maf/fbs_1.7/rolling/rolling_scale0.8_nslice3_v1.7_10yrs.db'
runName = dbFile.replace('.db', '')
conn = db.OpsimDatabase(dbFile)
outDir='temp'
resultsDb = db.ResultsDb(outDir=outDir)
# +
#nside=4096 # ~approximate resolution 0.858872 arcminutes
nside = 16 # 64
bundleList = []
#sql=''
sql = 'filter="g" or filter="i"'
#sqlconstraint = 'filter = "r" and night < 365'
sql_i = 'filter = "i"'
sql_g = 'filter = "g"'
sql_r = 'filter = "r"'
#sql = 'night < 2000 and (filter="g" or filter="i")'
#slicer = slicers.HealpixSlicer(nside=nside)
slicer = slicers.UserPointsSlicer(lv_dat['ra'], lv_dat['dec'])
#plotDict = {}
plotDict = {}
metric_ngal = GalaxyCountsMetric(nside=nside, metricName='numGal')
bundleList.append(metricBundles.MetricBundle(metric_ngal,slicer,sql_i, plotDict=plotDict,
runName=runName))
#metric_ngal = depthLimitedNumGalMetric.DepthLimitedNumGalMetric(nfilters_needed=2, nside=nside, lim_ebv=0.2, metricName='numGal')
#bundleList.append(metricBundles.MetricBundle(metric_ngal,slicer,sql, plotDict=plotDict,
# runName=runName))
#okr5 = (r5 < 30) & (r5 > 20)
#np.median(r5[okr5])
# 26.865
mafMap = maps.StellarDensityMap()
metric_nstar = metrics.StarDensityMetric(rmagLimit=26.865, metricName='nstars_rmag<medianM5')
bundleList.append(metricBundles.MetricBundle(metric_nstar,slicer, sql, mapsList=[mafMap], plotDict=plotDict, runName=runName))
metric_nstar24 = metrics.StarDensityMetric(rmagLimit=24, metricName='nstars_rmag<24')
metric_nstar24p5 = metrics.StarDensityMetric(rmagLimit=24.5, metricName='nstars_rmag<24.5')
metric_nstar25 = metrics.StarDensityMetric(rmagLimit=25, metricName='nstars_rmag<25')
metric_nstar25p5 = metrics.StarDensityMetric(rmagLimit=25.5, metricName='nstars_rmag<25.5')
metric_nstar26 = metrics.StarDensityMetric(rmagLimit=26, metricName='nstars_rmag<26')
metric_nstar26p5 = metrics.StarDensityMetric(rmagLimit=26.5, metricName='nstars_rmag<26.5')
metric_nstar27 = metrics.StarDensityMetric(rmagLimit=27, metricName='nstars_rmag<27')
bundleList.append(metricBundles.MetricBundle(metric_nstar24, slicer, sql, mapsList=[mafMap], plotDict=plotDict, runName=runName))
bundleList.append(metricBundles.MetricBundle(metric_nstar24p5, slicer, sql, mapsList=[mafMap], plotDict=plotDict, runName=runName))
bundleList.append(metricBundles.MetricBundle(metric_nstar25, slicer, sql, mapsList=[mafMap], plotDict=plotDict, runName=runName))
bundleList.append(metricBundles.MetricBundle(metric_nstar25p5, slicer, sql, mapsList=[mafMap], plotDict=plotDict, runName=runName))
bundleList.append(metricBundles.MetricBundle(metric_nstar26, slicer, sql, mapsList=[mafMap], plotDict=plotDict, runName=runName))
bundleList.append(metricBundles.MetricBundle(metric_nstar26p5, slicer, sql, mapsList=[mafMap], plotDict=plotDict, runName=runName))
bundleList.append(metricBundles.MetricBundle(metric_nstar27, slicer, sql, mapsList=[mafMap], plotDict=plotDict, runName=runName))
metric_coadd = metrics.Coaddm5Metric()
bundleList.append(metricBundles.MetricBundle(metric_coadd, slicer, sql_i, plotDict=plotDict))
bundleList.append(metricBundles.MetricBundle(metric_coadd, slicer, sql_g, plotDict=plotDict))
bundleList.append(metricBundles.MetricBundle(metric_coadd, slicer, sql_r, plotDict=plotDict))
#metric_seeing = metrics.PercentileMetric(col='seeingFwhmGeom', percentile=33.3)
#bundleList.append(metricBundles.MetricBundle(metric_seeing, slicer, sql_i, plotDict=plotDict))
#metric_median_seeing = metrics.MedianMetric(col='seeingFwhmGeom')
#bundleList.append(metricBundles.MetricBundle(metric_median_seeing, slicer, sql_i, plotDict=plotDict))
# -
bd = metricBundles.makeBundlesDictFromList(bundleList)
bg = metricBundles.MetricBundleGroup(bd, conn, outDir=outDir, resultsDb=resultsDb)
bg.runAll()
#bg.plotAll(closefigs=False)
# +
# S/N = S/sqrt(S+N) ~ S/sqrt(N)
# +
root0 = str.replace(runName, '/', '_')
root = str.replace(root0, '.', '_')
i5 = bg.bundleDict['opsim_CoaddM5_i_USER'].metricValues
g5 = bg.bundleDict['opsim_CoaddM5_g_USER'].metricValues
r5 = bg.bundleDict['opsim_CoaddM5_r_USER'].metricValues
#iband_seeing33 = bg.bundleDict['opsim_33th_ile_seeingFwhmGeom_i_USER'].metricValues
#iband_seeing = bg.bundleDict['opsim_Median_seeingFwhmGeom_i_USER'].metricValues
ngal = bg.bundleDict[root+'_numGal_i_USER'].metricValues
#ngal = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_numGal_g_or_i_USER'].metricValues
nstar = bg.bundleDict[root+'_nstars_rmagltmedianM5_g_or_i_USER'].metricValues
#ngal = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_numGal_i_USER'].metricValues
##ngal = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_numGal_g_or_i_USER'].metricValues
#nstar = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_nstars_rmagltmedianM5_g_or_i_USER'].metricValues
nstar_r24 = bg.bundleDict[root+'_nstars_rmaglt24_g_or_i_USER'].metricValues
nstar_r24p5 = bg.bundleDict[root+'_nstars_rmaglt24_5_g_or_i_USER'].metricValues
nstar_r25 = bg.bundleDict[root+'_nstars_rmaglt25_g_or_i_USER'].metricValues
nstar_r25p5 = bg.bundleDict[root+'_nstars_rmaglt25_5_g_or_i_USER'].metricValues
nstar_r26 = bg.bundleDict[root+'_nstars_rmaglt26_g_or_i_USER'].metricValues
nstar_r26p5 = bg.bundleDict[root+'_nstars_rmaglt26_5_g_or_i_USER'].metricValues
nstar_r27 = bg.bundleDict[root+'_nstars_rmaglt27_g_or_i_USER'].metricValues
#nstar_r24 = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_nstars_rmaglt24_g_or_i_USER'].metricValues
#nstar_r24p5 = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_nstars_rmaglt24_5_g_or_i_USER'].metricValues
#nstar_r25 = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_nstars_rmaglt25_g_or_i_USER'].metricValues
#nstar_r25p5 = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_nstars_rmaglt25_5_g_or_i_USER'].metricValues
#nstar_r26 = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_nstars_rmaglt26_g_or_i_USER'].metricValues
#nstar_r26p5 = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_nstars_rmaglt26_5_g_or_i_USER'].metricValues
#nstar_r27 = bg.bundleDict['_sims_maf_fbs_1_7_baseline_baseline_nexp2_v1_7_10yrs_nstars_rmaglt27_g_or_i_USER'].metricValues
# -
plt.hist(r5, bins=np.arange(21, 30, 0.1), log=True)
plt.show()
# Calculate the factor to go from number per healpix to number per square arcminute or per square arcsec
pixarea_deg = hp.nside2pixarea(nside, degrees=True)*(u.degree**2)
pixarea_arcmin = pixarea_deg.to(u.arcmin**2)
pixarea_arcsec = pixarea_deg.to(u.arcsec**2)
# +
nstar_all = nstar*0.0
rbinvals = np.arange(24.0, 27.5, 0.5)
rbinnames = [nstar_r24, nstar_r24p5, nstar_r25, nstar_r25p5, nstar_r26, nstar_r26p5, nstar_r27]
for binval,bindensity in zip(rbinvals, rbinnames):
inbin = np.where(np.abs(r5-binval) < 0.25)
nstar_all[inbin] = bindensity[inbin]
outside_faint = (r5 >= 27.25)
outside_bright = (r5 <= 23.75)
nstar_all[outside_faint] = nstar[outside_faint]
nstar_all[outside_bright] = nstar_r24[outside_bright]
# Star density is number of stars per square arcsec. Convert to a total number per healpix, then number per sq. arcmin:
nstar_all_per_healpix = nstar_all*pixarea_arcsec
nstar_all_per_arcmin = nstar_all_per_healpix/pixarea_arcmin
# +
# Number of galaxies is the total in each healpix. Convert to number per sq. arcmin:
ngal_per_arcmin = ngal/pixarea_arcmin
# Star density is number of stars per square arcsec. Convert to a total number per healpix, then number per sq. arcmin:
nstar_per_healpix = nstar*pixarea_arcsec
nstar_per_arcmin = nstar_per_healpix/pixarea_arcmin
# -
# Compare the fractional difference between the different densities:
plt.hist((nstar_all_per_arcmin-nstar_per_arcmin)/nstar_per_arcmin, bins=np.arange(-1, 1, 0.05))
plt.show()
# +
# Account for:
# - cmd_frac: fraction of CMD area being selected
# - stargal_contamination: what fraction of objects in your "star" selection after star-galaxy separation are background galaxy contaminants?
cmd_frac = 0.1
stargal_contamination = 0.40
nsigma = 10.0
# Output: number of stars required to reach our requested S/N
#dum = hp.mollview(ngal_per_arcmin, title='ngxs')
#dum = hp.mollview(nstar_per_arcmin, title='nstars')
#dum = hp.mollview(nsigma*np.sqrt(ngal_per_arcmin*(cmd_frac*stargal_contamination)), title='nstars')
#dum = hp.mollview(nsigma*np.sqrt(ngal_per_arcmin*(cmd_frac*stargal_contamination)+(nstar_per_arcmin*cmd_frac)), title='nstars', max=30)
# -
np.nanmean(ngal_per_arcmin)
# ### Next apply this to simulated dwarf galaxy stellar populations:
#
# 1. Calculate the luminosity function of a simulated dwarf _of a given luminosity_,
# 2. Place it at the desired distance,
# 3. Derive the surface brightness of this dwarf at the applied distance.
# +
'''
#first: compute integrated mag in B:
LF = ascii.read('LF_-1.5_10Gyr_B.dat', header_start=12)
mags = LF['magbinc']
counts = LF['Bmag']
intBmag = -2.5*np.log10( np.sum(counts * np.power(10.0,-0.4*mags) ))
print ('intMag='+str(intBmag))
#result is 6.856379, store it for later
'''
# make fake LF for old galaxy of given integrated B, distance modulus mu, in any of filters ugrizY
def makeFakeLF(intB, mu, filtername):
if (filtername=='y'): filtername=='Y'
modelBmag = 6.856379 # integrated B mag of the model LF being read
LF = ascii.read('LF_-1.5_10Gyr.dat', header_start=12)
mags = LF['magbinc']
counts = LF[filtername+'mag']
# shift model LF to requested distance and dim it
mags = mags + mu
modelBmag = modelBmag + mu
# scale model counts up/down to reach the requested intB
factor = np.power(10.0,-0.4*(intB-modelBmag))
counts = factor * counts
# resample from Poisson:
return mags, counts
# countsPoisson = np.random.poisson(counts)
# return mags, countsPoisson
#test: now will make a fake LF for 47Tuc, with intB=5.78, mu0=13.32
#intB=5.78
#mu0=13.32
#LFmags,LFcounts = makeFakeLF(intB,mu0,'r')
#for i in range(len(LFmags)):
# print (LFmags[i],LFcounts[i])
# +
lf_dict_i = {}
lf_dict_g = {}
tmp_MB = -10.0
for i in range(101):
mbkey = f'MB{tmp_MB:.2f}'
iLFmags,iLFcounts = makeFakeLF(tmp_MB, 0.0, 'i')
lf_dict_i[mbkey] = (np.array(iLFmags), np.array(iLFcounts))
gLFmags,gLFcounts = makeFakeLF(tmp_MB, 0.0, 'g')
lf_dict_g[mbkey] = (np.array(gLFmags), np.array(gLFcounts))
tmp_MB += 0.1
# -
plt.plot(iLFmags, np.log10(iLFcounts))
lf_dict_i.keys()
# +
def sum_luminosity(LFmags, LFcounts):
magref = LFmags[0]
totlum = 0.0
for mag, count in zip(LFmags, LFcounts):
tmpmags = np.repeat(mag, count)
totlum += np.sum(10.0**((magref - tmpmags)/2.5))
mtot = magref-2.5*np.log10(totlum)
return mtot
def sblimit(mags_g, mags_i, nstars_req, distlim):
distance_limit = distlim*1e6 # distance limit in parsecs
distmod_limit = 5.0*np.log10(distance_limit) - 5.0
mg_lim = []
mi_lim = []
sbg_lim = []
sbi_lim = []
flag_lim = []
for glim, ilim, nstars, distmod_limit in zip(mags_g, mags_i, nstars_req, distmod_limit):
# for i in range(len(mags_g)):
if (glim > 15) and (ilim > 15):
# print(glim, ilim, nstars)
fake_MB = -10.0
ng = 1e6
ni = 1e6
while (ng > nstars) and (ni > nstars) and fake_MB<-2.0:
# B_fake = distmod_limit+fake_MB
mbkey = f'MB{fake_MB:.2f}'
iLFmags0,iLFcounts0 = lf_dict_i[mbkey]
gLFmags0,gLFcounts0 = lf_dict_g[mbkey]
iLFcounts = np.random.poisson(iLFcounts0)
gLFcounts = np.random.poisson(gLFcounts0)
iLFmags = iLFmags0+distmod_limit # Add the distance modulus to make it apparent mags
gLFmags = gLFmags0+distmod_limit # Add the distance modulus to make it apparent mags
# print(iLFcounts0-iLFcounts)
gsel = (gLFmags <= glim)
isel = (iLFmags <= ilim)
ng = np.sum(gLFcounts[gsel])
ni = np.sum(iLFcounts[isel])
# print('fake_MB: ',fake_MB, ' ng: ',ng, ' ni: ', ni, ' nstars: ', nstars)
fake_MB += 0.1
if fake_MB > -9.9:
gmag_tot = sum_luminosity(gLFmags[gsel], gLFcounts[gsel]) - distmod_limit
imag_tot = sum_luminosity(iLFmags[isel], iLFcounts[isel]) - distmod_limit
# S = m + 2.5logA, where in this case things are in sq. arcmin, so A = 1 arcmin^2 = 3600 arcsec^2
sbtot_g = distmod_limit + gmag_tot + 2.5*np.log10(3600.0)
sbtot_i = distmod_limit + imag_tot + 2.5*np.log10(3600.0)
mg_lim.append(gmag_tot)
mi_lim.append(imag_tot)
sbg_lim.append(sbtot_g)
sbi_lim.append(sbtot_i)
if (ng < ni):
flag_lim.append('g')
else:
flag_lim.append('i')
else:
mg_lim.append(999.9)
mi_lim.append(999.9)
sbg_lim.append(999.9)
sbi_lim.append(999.9)
flag_lim.append('none')
else:
mg_lim.append(999.9)
mi_lim.append(999.9)
sbg_lim.append(-999.9)
sbi_lim.append(-999.9)
flag_lim.append('none')
return mg_lim, mi_lim, sbg_lim, sbi_lim, flag_lim
# +
# Account for:
# - cmd_frac: fraction of CMD area being selected
# - stargal_contamination: what fraction of objects in your "star" selection after star-galaxy separation are background galaxy contaminants?
cmd_frac = 0.1
stargal_contamination = 0.40
nsigma = 10.0
nstars_required = nsigma*np.sqrt(ngal_per_arcmin*(cmd_frac*stargal_contamination)+(nstar_all_per_arcmin*cmd_frac))
#nstars_required = nsigma*np.sqrt(ngal_per_arcmin*(cmd_frac*stargal_contamination)+(nstar_per_arcmin*cmd_frac))
# +
# nstars_required
# -
mg_lim, mi_lim, sb_g_lim, sb_i_lim, flag_lim = sblimit(g5, i5, nstars_required, distlim=lv_dat['dist_Mpc'])
# +
#plt.plot(lv_dat['dist_Mpc'], mi_lim, 'k.')
#plt.ylim(-3, -12)
#plt.plot(lv_dat['dist_Mpc'], sb_i_lim, 'k.')
#plt.ylim(25, 35)
mg_lim = np.array(mg_lim)
sb_i_lim = np.array(sb_i_lim)
okgx = (np.abs(sb_i_lim) < 90) # & (lv_dat['dec'] < 20)
plt.scatter(lv_dat[okgx]['ra'], lv_dat[okgx]['dec'], c=mg_lim[okgx], cmap='Spectral', vmax=-6, vmin=-3)
#plt.scatter(lv_dat[okgx]['ra'], lv_dat[okgx]['dec'], c=sb_i_lim[okgx], cmap='Spectral', vmax=31, vmin=28)
plt.plot(lv_dat['ra'][~okgx], lv_dat['dec'][~okgx], '.', color='Gray', alpha=0.3)
plt.colorbar(label='limiting $M_g$')
plt.xlim(360,0)
plt.xlabel('RA (deg)')
plt.ylabel('Dec (deg)')
plt.show()
# -
plt.hist(lv_dat[okgx]['dist_Mpc'], bins=np.arange(1.2, 9.0, 0.2), histtype='step', color='Black', label='has limit')
plt.hist(lv_dat[~okgx]['dist_Mpc'], bins=np.arange(1.2, 9.0, 0.2), histtype='step', color='Gray', linestyle='--', label='unsuccessful')
plt.legend(loc='upper left')
plt.xlabel('dist (Mpc)')
plt.show()
np.sum((lv_dat['Mstars'] > 1e8) & (lv_dat['dec'] < 20))
print(np.sum(np.array(mi_lim) < 30), np.sum(np.array(mi_lim) < 30)/len(mi_lim), np.sum(np.array(mi_lim) > 30)/len(mi_lim))
#plt.hist(lv_dat[okgx]['M_B'], bins=np.arange(-22.0, -2.0, 0.25), histtype='step', color='Black')
plt.hist(np.log10(lv_dat[okgx]['MStars']), bins=np.arange(6.0, 12.0, 0.25), histtype='step', color='Black')
plt.xlabel('M_stars (M_Sun)')
plt.show()
# +
params = {
'axes.labelsize': 20,
'font.size': 20,
'legend.fontsize': 14,
# 'xtick.labelsize': 16,
'xtick.major.width': 3,
'xtick.minor.width': 2,
'xtick.major.size': 8,
'xtick.minor.size': 5,
'xtick.direction': 'in',
'xtick.top': True,
'lines.linewidth':3,
'axes.linewidth':3,
'axes.labelweight':3,
'axes.titleweight':3,
'ytick.major.width':3,
'ytick.minor.width':2,
'ytick.major.size': 8,
'ytick.minor.size': 5,
'ytick.direction': 'in',
'ytick.right': True,
# 'ytick.labelsize': 20,
# 'text.usetex': True,
# 'text.latex.preamble': r'\boldmath',
'figure.figsize': [9, 7],
'figure.facecolor': 'White'
}
plt.rcParams.update(params)
plt.plot(lv_dat['Dist_Mpc'], mi_lim, 'k.')
plt.ylim(-2.5, -10.5)
plt.xlim(1.2, 6.2)
plt.xlabel('distance (Mpc)')
plt.ylabel('limiting $M_i$')
plt.minorticks_on()
plt.show()
# -
plt.plot(lv_dat['Dist_Mpc'], sb_i_lim, 'k.')
plt.ylim(23, 35)
plt.xlim(1.2, 6.2)
plt.xlabel('distance (Mpc)')
plt.ylabel('limiting $\mu_i$ (mag/arcsec$^2$)')
plt.minorticks_on()
plt.show()
# +
# Use the conversion from Appendix A of Komiyama+2018, ApJ, 853, 29:
# V = g_hsc - 0.371*(gi_hsc)-0.068
mv = mg_lim-0.371*(mg_lim-mi_lim)-0.068
sbv = sb_g_lim-0.371*(sb_g_lim-sb_i_lim)-0.068
# -
#plt.hist(mv, bins=np.arange(-10.3, -3.3, 0.2), histtype='step', color='Black')
#plt.hist(mg_lim, bins=np.arange(-10.3, -3.3, 0.2), histtype='step', color='Blue')
#plt.hist(mi_lim, bins=np.arange(-10.3, -3.3, 0.2), histtype='step', color='Red')
plt.hist(sbv, bins=np.arange(27, 35, 0.2), histtype='step', color='Black')
plt.hist(sb_g_lim, bins=np.arange(27, 35, 0.2), histtype='step', color='Blue')
plt.hist(sb_i_lim, bins=np.arange(27, 35, 0.2), histtype='step', color='Red')
plt.show()
plt.plot(lv_dat['Dist_Mpc'], sbv, 'k.')
plt.ylim(23, 35)
plt.xlim(1.2, 6.2)
plt.xlabel('distance (Mpc)')
plt.ylabel('limiting $\mu_V$ (mag/arcsec$^2$)')
plt.minorticks_on()
plt.show()
plt.plot(lv_dat['Dist_Mpc'], mv, 'k.')
plt.ylim(-2.5, -8.5)
plt.xlim(1.2, 8.2)
plt.xlabel('distance (Mpc)')
plt.ylabel('limiting $M_V$')
plt.minorticks_on()
plt.show()
print(len(lv_dat), len(lv_dat[(mv > -7) & (mv < 0)]))
import glob
glob.glob('/sims_maf/fbs_1.7/rolling/*')
lv_dat['Dist_Mpc']
| notebooks/LV_metric_scratch_LVgxs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Market Basket Encoder
# This function takes in Transaction ID and Product and encodes into a matrix of ID's and Products
#modules
import pandas as pd
# +
df = pd.read_csv("https://raw.githubusercontent.com/simmieyungie/Bakery-Analytics/master/BreadBasket_DMS.csv")
df.head()
# -
df.groupby(["Transaction", "Item"]).size()
# +
#Configure os folder
import os
# Create a folder for the pipeline step files
experiment_folder = r'C:\Users\SIMIYOUNG\Documents\Python-Projects\RFM\PyRFM\RFM'
os.makedirs(experiment_folder, exist_ok=True)
print(experiment_folder)
# -
# %%writefile $experiment_folder/mba_encoder.py
def mba_encoder(data, id_col, product_col, quantity_col, summary_criteria = None):
'''
This function will preprocess and return a one-hot encode matrix of ID's and products.
data: Dataframe
A dataframe containing at least the customer ID, Transaction/Order Date
id_col: String
Column name containing ID
product_col: String
Column name containing product col
quantity_col: String
Column name containing quantity of product purchase
summary_criteria: Dict
A dictionary containing summary criteria
'''
#error handlers
#error instance to check if input is either a pandas dataframe and is also not none
if isinstance(data, pd.DataFrame) == False or data is None:
raise ValueError("data: Expecting a Dataframe or got 'None'")
#error instance to check if id column is an identified column name
if id_col not in data.columns:
raise ValueError("id: Expected a valid id column name in Dataframe 'data'")
#error instance to check if product column is an identified column name
if product_col not in data.columns:
raise ValueError("id: Expected a valid product column name in Dataframe 'data'")
#error handler if instance for summary_criteria if
if summary_criteria != None and isinstance(summary_criteria, dict) == False:
raise TypeError("summary_criteria: Expected a dict dtype if summary_criteria is not 'None'")
#if data is to be summarised
if summary_criteria != None and isinstance(summary_criteria, dict) == True:
summary = list(summary_criteria.keys())
cols = list(summary_criteria.values())
data.groupby([summary])
#Encoding
basket = data \
.unstack() \
.reset_index()
.set_index(id_col)
return basket
| Playground/Market Basket Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercise 05: Using pandas to Compute the Mean, Median, and Variance of a Dataset
# In this exercise, you will consolidate the skills you've acquired in the last exercise and use Pandas to do some very basic mathematical calculations on our `world_population.csv` dataset.
# Pandas have consistent API, so it should be rather easy to transfer your knowledge of the mean method to median and variance.
# Your already existing knowledge of NumPy will also help.
# #### Loading the dataset
# importing the necessary dependencies
import pandas as pd
# loading the Dataset
dataset = pd.read_csv('../../Datasets/world_population.csv', index_col=0)
# looking at the first two rows of the dataset
dataset[0:2]
# ---
# #### Mean
# calculate the mean of the third row
dataset.iloc[[2]].mean(axis=1)
# calculate the mean of the last row
dataset.iloc[[-1]].mean(axis=1)
# calculate the mean of the country Germany
dataset.loc[["Germany"]].mean(axis=1)
# **Note:**
# `.iloc()` and `.loc()` are two important methods when indexing with Pandas. They allow to make precise selections of data based on either the integer value index (`iloc`) or the index column (`loc`), which in our case is the country name column.
# ---
# #### Median
# calculate the median of the last row
dataset.iloc[[-1]].median(axis=1)
# calculate the median of the last 3 rows
dataset[-3:].median(axis=1)
# **Note:**
# Slicing can be done in the same way as with NumPy.
# `dataset[1:3]` will return the second and third row of our dataset.
# calculate the median of the first 10 countries
dataset.head(10).median(axis=1)
# **Note:**
# When handling larger datasets, the order in which methods get executed definitely matters.
# Think about what `.head(10)` does for a moment, it simply takes your dataset and returns the first 10 rows of it, cutting down your input to the `.mean()` method drastically.
# This will definitely have an impact when using more memory intensive calculations, so keep an eye on the order.
# ---
# #### Variance
# calculate the variance of the last 5 columns
dataset.var().tail()
# ---
# As mentioned in the introduction of Pandas, it's interoperable with several of NumPy's features.
# Here's an example of how to use NumPy's `mean` method with a Pandas dataFrame.
# +
# NumPy Pandas interoperability
import numpy as np
print("Pandas", dataset["2015"].mean())
print("NumPy", np.mean(dataset["2015"]))
| Exercise05/Exercise05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: DS-Unit-4-Sprint-3-Deep-Learning
# language: python
# name: ds-unit-4-sprint-3-deep-learning
# ---
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# ## *Data Science Unit 4 Sprint 3 Assignment 1*
#
# # Recurrent Neural Networks and Long Short Term Memory (LSTM)
#
# 
#
# It is said that [infinite monkeys typing for an infinite amount of time](https://en.wikipedia.org/wiki/Infinite_monkey_theorem) will eventually type, among other things, the complete works of Wiliam Shakespeare. Let's see if we can get there a bit faster, with the power of Recurrent Neural Networks and LSTM.
#
# This text file contains the complete works of Shakespeare: https://www.gutenberg.org/files/100/100-0.txt
#
# Use it as training data for an RNN - you can keep it simple and train character level, and that is suggested as an initial approach.
#
# Then, use that trained RNN to generate Shakespearean-ish text. Your goal - a function that can take, as an argument, the size of text (e.g. number of characters or lines) to generate, and returns generated text of that size.
#
# Note - Shakespeare wrote an awful lot. It's OK, especially initially, to sample/use smaller data and parameters, so you can have a tighter feedback loop when you're trying to get things running. Then, once you've got a proof of concept - start pushing it more!
# + colab={} colab_type="code" id="Ltj1je1fp5rO"
import tensorflow as tf
import numpy as np
import os
import time
import re
# -
path_to_shakespeare = tf.keras.utils.get_file('shakespeare.txt', 'https://www.gutenberg.org/files/100/100-0.txt')
text = open(path_to_shakespeare, 'rb').read().decode(encoding='utf-8')
# count number of chars
len(text)
def clean_shakespeare(text):
# get contents
contents = text[974:2893].replace("\r", "").replace("\n", "")
contents = re.sub("\s{2,}", ",,", contents)
contents = contents.split(",,")
for idx,book in enumerate(contents):
if book == "THE LIFE OF KING HENRY THE FIFTH":
contents[idx] = "THE LIFE OF KING HENRY V"
elif book == "THE TRAGEDY OF MACBETH":
contents[idx] = "MACBETH"
elif book == "THE TRAGEDY OF OTHELLO, MOOR OF VENICE":
contents[idx] = "OTHELLO, THE MOOR OF VENICE"
elif book == "TWELFTH NIGHT; OR, WHAT YOU WILL":
contents[idx] = "TWELFTH NIGHT: OR, WHAT YOU WILL"
# remove the project gutenberg info
text = text[2893:-21529]
shakespeare_dict = {}
for idx,book in enumerate(contents):
strpos = text.find(book) + len(book)
if idx + 1 != len(contents):
next_book = contents[idx + 1]
endpos = text.find(next_book, strpos)
shakespeare_dict[book] = text[strpos:endpos]
else:
shakespeare_dict[book] = text[strpos:]
return shakespeare_dict, text
books, text = clean_shakespeare(text)
vocab = sorted(set(text))
len(vocab)
# +
char2idx = {u:i for i, u in enumerate(vocab)}
idx2char = np.array(vocab)
text_as_int = np.array([char2idx[c] for c in text])
# -
seq_length = 100
examples_per_epoch = len(text)//(seq_length+1)
char_dataset = tf.data.Dataset.from_tensor_slices(text_as_int)
sequences = char_dataset.batch(seq_length+1, drop_remainder=True)
# +
def split_input_target(chunk):
input_text = chunk[:-1]
target_text = chunk[1:]
return input_text, target_text
dataset = sequences.map(split_input_target)
# -
for input_example, target_example in dataset.take(1):
print ('Input data: ', repr(''.join(idx2char[input_example.numpy()])))
print ('Target data:', repr(''.join(idx2char[target_example.numpy()])))
for i, (input_idx, target_idx) in enumerate(zip(input_example[:5], target_example[:5])):
print("Step {:4d}".format(i))
print(" input: {} ({:s})".format(input_idx, repr(idx2char[input_idx])))
print(" expected output: {} ({:s})".format(target_idx, repr(idx2char[target_idx])))
# +
BATCH_SIZE = 64
BUFFER_SIZE = 10000
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
# +
vocab_size = len(vocab)
embedding_dim = 256
rnn_units = 1024
# -
def build_model(vocab_size, embedding_dim, rnn_units, batch_size):
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim,
batch_input_shape=[batch_size, None]),
tf.keras.layers.GRU(rnn_units,
return_sequences=True,
stateful=True,
recurrent_initializer='glorot_uniform'),
tf.keras.layers.Dense(vocab_size)
])
return model
model = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
def loss(labels, logits):
return tf.keras.losses.sparse_categorical_crossentropy(labels, logits, from_logits=True)
model.compile(optimizer='adam', loss=loss)
# +
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt_{epoch}")
checkpoint_callback=tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_prefix,
save_weights_only=True)
# -
EPOCHS=30
history = model.fit(dataset, epochs=EPOCHS, callbacks=[checkpoint_callback])
# +
model = build_model(vocab_size, embedding_dim, rnn_units, batch_size=1)
model.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
model.build(tf.TensorShape([1, None]))
# -
model.summary()
def generate_text(model, start_string):
# Evaluation step (generating text using the learned model)
# Number of characters to generate
num_generate = 1000
# Converting our start string to numbers (vectorizing)
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
# Empty string to store our results
text_generated = []
# Low temperatures results in more predictable text.
# Higher temperatures results in more surprising text.
# Experiment to find the best setting.
temperature = 1.0
# Here batch size == 1
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
# remove the batch dimension
predictions = tf.squeeze(predictions, 0)
# using a categorical distribution to predict the character returned by the model
predictions = predictions / temperature
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
# We pass the predicted character as the next input to the model
# along with the previous hidden state
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2char[predicted_id])
return (start_string + ''.join(text_generated))
print(generate_text(model, start_string=u"A rose "))
# + [markdown] colab_type="text" id="zE4a4O7Bp5x1"
# # Resources and Stretch Goals
# + [markdown] colab_type="text" id="uT3UV3gap9H6"
# ## Stretch goals:
# - Refine the training and generation of text to be able to ask for different genres/styles of Shakespearean text (e.g. plays versus sonnets)
# - Train a classification model that takes text and returns which work of Shakespeare it is most likely to be from
# - Make it more performant! Many possible routes here - lean on Keras, optimize the code, and/or use more resources (AWS, etc.)
# - Revisit the news example from class, and improve it - use categories or tags to refine the model/generation, or train a news classifier
# - Run on bigger, better data
#
# ## Resources:
# - [The Unreasonable Effectiveness of Recurrent Neural Networks](https://karpathy.github.io/2015/05/21/rnn-effectiveness/) - a seminal writeup demonstrating a simple but effective character-level NLP RNN
# - [Simple NumPy implementation of RNN](https://github.com/JY-Yoon/RNN-Implementation-using-NumPy/blob/master/RNN%20Implementation%20using%20NumPy.ipynb) - Python 3 version of the code from "Unreasonable Effectiveness"
# - [TensorFlow RNN Tutorial](https://github.com/tensorflow/models/tree/master/tutorials/rnn) - code for training a RNN on the Penn Tree Bank language dataset
# - [4 part tutorial on RNN](http://www.wildml.com/2015/09/recurrent-neural-networks-tutorial-part-1-introduction-to-rnns/) - relates RNN to the vanishing gradient problem, and provides example implementation
# - [RNN training tips and tricks](https://github.com/karpathy/char-rnn#tips-and-tricks) - some rules of thumb for parameterizing and training your RNN
| module1-rnn-and-lstm/LS_DS_431_RNN_and_LSTM_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''sc_workshops'': conda)'
# metadata:
# interpreter:
# hash: 787ea478c22349cf73d867deb0a19fc58e75e9742a9aed6f48d06cc412ad6e3b
# name: python3
# ---
# # Combining Distance Lattices
#
# In this workshop, we will learn about how to combine different distance fields in advance to simulation.
# ## 0. Initialization
#
# ### 0.1. Load required libraries
import os
import topogenesis as tg
import pyvista as pv
import trimesh as tm
import numpy as np
import networkx as nx
np.random.seed(0)
# ### 0.2. Define the Neighborhood (Stencil)
# creating neighborhood definition
stencil = tg.create_stencil("von_neumann", 1, 1)
# setting the center to zero
stencil.set_index([0,0,0], 0)
print(stencil)
# ### 0.3. Load the envelope lattice as the avialbility lattice
# loading the lattice from csv
lattice_path = os.path.relpath('../data/voxelized_envelope.csv')
avail_lattice = tg.lattice_from_csv(lattice_path)
init_avail_lattice = tg.to_lattice(np.copy(avail_lattice), avail_lattice)
# ## 1. Distance Field Construction
#
# ### 1.1. Extract the connectivity graph from the lattice based on the defined stencil
# +
# find the number of all voxels
vox_count = avail_lattice.size
# initialize the adjacency matrix
adj_mtrx = np.zeros((vox_count,vox_count))
# Finding the index of the available voxels in avail_lattice
avail_index = np.array(np.where(avail_lattice == 1)).T
# fill the adjacency matrix using the list of all neighbours
for vox_loc in avail_index:
# find the 1D id
vox_id = np.ravel_multi_index(vox_loc, avail_lattice.shape)
# retrieve the list of neighbours of the voxel based on the stencil
vox_neighs = avail_lattice.find_neighbours_masked(stencil, loc = vox_loc)
# iterating over the neighbours
for neigh in vox_neighs:
# setting the entry to one
adj_mtrx[vox_id, neigh] = 1.0
# construct the graph
g = nx.from_numpy_array(adj_mtrx)
# -
# ### 1.2. Compute distances on the graph
# compute the distance of all voxels to all voxels using floyd warshal algorithm
dist_mtrx = nx.floyd_warshall_numpy(g)
# ### 1.3. Select the entrance voxel
# +
p = pv.Plotter(notebook=True)
# initialize the selection lattice
base_lattice = avail_lattice * 0 - 1
# init base flat
base_flat = base_lattice.flatten().astype(int)
# Set the grid dimensions: shape + 1 because we want to inject our values on the CELL data
grid = pv.UniformGrid()
grid.dimensions = np.array(base_lattice.shape) + 1
# The bottom left corner of the data set
grid.origin = base_lattice.minbound - base_lattice.unit * 0.5
# These are the cell sizes along each axis
grid.spacing = base_lattice.unit
# adding the boundingbox wireframe
p.add_mesh(grid.outline(), color="grey", label="Domain")
# adding the avilability lattice
init_avail_lattice.fast_vis(p)
# adding axes
p.add_axes()
p.show_bounds(grid="back", location="back", color="#aaaaaa")
def create_mesh(value):
i = int(value)
# init base flat
base_flat = base_lattice.flatten().astype(int)
base_flat = base_flat * 0 - 1
base_flat[i] = 0
base_new = base_flat.reshape(base_lattice.shape)
# Add the data values to the cell data
grid.cell_arrays["Selection"] = base_new.flatten(order="F").astype(int) # Flatten the array!
# filtering the voxels
threshed = grid.threshold([-0.1, 0.9])
# adding the voxels
p.add_mesh(threshed, name='sphere', show_edges=True, opacity=1.0, show_scalar_bar=False)
return
p.add_slider_widget(create_mesh, [0, len(base_flat)], title='1D Index', value=0, event_type="always", style="classic", pointa=(0.1, 0.1), pointb=(0.9, 0.1))
p.show(use_ipyvtk=True)
# -
# ### 1.4. Construct Distance to 1st Entrance Lattice
# +
# select the corresponding row in the matrix
ent_1_dist = dist_mtrx[51]
# find the maximum valid value
max_valid = np.ma.masked_invalid(ent_1_dist).max()
# set the infinities to one more than the maximum valid values
ent_1_dist[ent_1_dist == np.inf] = max_valid + 1
# -
# ### 1.5. Construct Distance to 2nd Entrance Lattice
# +
# select the corresponding row in the matrix
ent_2_dist = dist_mtrx[682]
# find the maximum valid value
max_valid = np.ma.masked_invalid(ent_2_dist).max()
# set the infinities to one more than the maximum valid values
ent_2_dist[ent_2_dist == np.inf] = max_valid + 1
# -
# ### 1.6. Combining different distance fields
# +
# finding the minimum distance between two entrance
# fields (aka finding the closest entrance and replacing
# the distance with the distance to that entrance)
ent_dist = np.minimum(ent_1_dist, ent_2_dist)
# mapping the values from (0, max) to (1, 0)
ent_flat = 1 - ent_dist / np.max(ent_dist)
# constructing the lattice
ent_acc_lattice = tg.to_lattice(ent_flat.reshape(avail_lattice.shape), avail_lattice)
# -
# ### 1.7. Visualize the distance lattice
# +
# convert mesh to pv_mesh
def tri_to_pv(tri_mesh):
faces = np.pad(tri_mesh.faces, ((0, 0),(1,0)), 'constant', constant_values=3)
pv_mesh = pv.PolyData(tri_mesh.vertices, faces)
return pv_mesh
# load the mesh from file
context_path = os.path.relpath('../data/immediate_context.obj')
context_mesh = tm.load(context_path)
# initiating the plotter
p = pv.Plotter(notebook=True)
# Create the spatial reference
grid = pv.UniformGrid()
# Set the grid dimensions: shape because we want to inject our values
grid.dimensions = ent_acc_lattice.shape
# The bottom left corner of the data set
grid.origin = ent_acc_lattice.minbound
# These are the cell sizes along each axis
grid.spacing = ent_acc_lattice.unit
# Add the data values to the cell data
grid.point_arrays["Entrance Access"] = ent_acc_lattice.flatten(order="F") # Flatten the Lattice
# adding the meshes
p.add_mesh(tri_to_pv(context_mesh), opacity=0.1, style='wireframe')
# adding the volume
opacity = np.array([0.0,0.6,0.6,0.6,0.6,0.6,0.6]) * 0.6
p.add_volume(grid, cmap="coolwarm", clim=[0.0, 1.0] ,opacity=opacity)
# plotting
p.show(use_ipyvtk=True)
# -
# ### 1.6. Save Entrance Access Lattice to CSV
# +
# save the sun access latice to csv
csv_path = os.path.relpath('../data/ent_access.csv')
ent_acc_lattice.to_csv(csv_path)
# -
# ### Credits
__author__ = "<NAME> and <NAME>"
__license__ = "MIT"
__version__ = "1.0"
__url__ = "https://github.com/shervinazadi/spatial_computing_workshops"
__summary__ = "Spatial Computing Design Studio Workshop on MCDA and Path Finding for Generative Spatial Relations"
| notebooks/w+3_combining_lattices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Abstractive
# <div class="alert alert-info">
#
# This tutorial is available as an IPython notebook at [Malaya/example/abstractive-summarization](https://github.com/huseinzol05/Malaya/tree/master/example/abstractive-summarization).
#
# </div>
# <div class="alert alert-warning">
#
# This module only trained on standard language structure, so it is not save to use it for local language structure.
#
# </div>
# <div class="alert alert-warning">
#
# This module trained heavily on news structure.
#
# </div>
# %%time
import malaya
from pprint import pprint
# +
import re
# minimum cleaning, just simply to remove newlines.
def cleaning(string):
string = string.replace('\n', ' ')
string = re.sub(r'[ ]+', ' ', string).strip()
return string
# -
# I am going to simply copy paste some local news into this notebook. I will search about `<NAME>` in google news, [link here](https://www.google.com/search?q=isu+mahathir&sxsrf=ALeKk02V_bAJC3sSrV38JQgGYWL_mE0biw:1589951900053&source=lnms&tbm=nws&sa=X&ved=2ahUKEwjapNmx2MHpAhVp_XMBHRt7BEQQ_AUoAnoECCcQBA&biw=1440&bih=648&dpr=2).
# **link**: https://www.hmetro.com.my/mutakhir/2020/05/580438/peletakan-jawatan-tun-m-ditolak-bukan-lagi-isu
#
# **Title**: Peletakan jawatan Tun M ditolak, bukan lagi isu.
#
# **Body**: PELETAKAN jawatan Tun Dr <NAME> sebagai Pengerusi Parti Pribumi Bersatu Malaysia (Bersatu) ditolak di dalam mesyuarat khas Majlis Pimpinan Tertinggi (MPT) pada 24 Februari lalu.
#
# Justeru, tidak timbul soal peletakan jawatan itu sah atau tidak kerana ia sudah pun diputuskan pada peringkat parti yang dipersetujui semua termasuk Presiden, Tan Sri Muhyiddin Yassin.
#
# Bekas Setiausaha Agung Bersatu Datuk Marzuki Yahya berkata, pada mesyuarat itu MPT sebulat suara menolak peletakan jawatan Dr Mahathir.
#
# "Jadi ini agak berlawanan dengan keputusan yang kita sudah buat. Saya tak faham bagaimana Jabatan Pendaftar Pertubuhan Malaysia (JPPM) kata peletakan jawatan itu sah sedangkan kita sudah buat keputusan di dalam mesyuarat, bukan seorang dua yang buat keputusan.
#
# "Semua keputusan mesti dibuat melalui parti. Walau apa juga perbincangan dibuat di luar daripada keputusan mesyuarat, ini bukan keputusan parti.
#
# "Apa locus standy yang ada pada Setiausaha Kerja untuk membawa perkara ini kepada JPPM. Seharusnya ia dibawa kepada Setiausaha Agung sebagai pentadbir kepada parti," katanya kepada Harian Metro.
#
# Beliau mengulas laporan media tempatan hari ini mengenai pengesahan JPPM bahawa Dr Mahathir tidak lagi menjadi Pengerusi Bersatu berikutan peletakan jawatannya di tengah-tengah pergolakan politik pada akhir Februari adalah sah.
#
# Laporan itu juga menyatakan, kedudukan <NAME> memangku jawatan itu juga sah.
#
# Menurutnya, memang betul Dr Mahathir menghantar surat peletakan jawatan, tetapi ditolak oleh MPT.
#
# "Fasal yang disebut itu terpakai sekiranya berhenti atau diberhentikan, tetapi ini mesyuarat sudah menolak," katanya.
#
# Marzuki turut mempersoal kenyataan media yang dibuat beberapa pimpinan parti itu hari ini yang menyatakan sokongan kepada Perikatan Nasional.
#
# "Kenyataan media bukanlah keputusan rasmi. Walaupun kita buat 1,000 kenyataan sekali pun ia tetap tidak merubah keputusan yang sudah dibuat di dalam mesyuarat. Kita catat di dalam minit apa yang berlaku di dalam mesyuarat," katanya.
# +
string = """
PELETAKAN jawatan Tun Dr <NAME> sebagai Pengerusi Parti Pribumi Bersatu Malaysia (Bersatu) ditolak di dalam mesyuarat khas Majlis Pimpinan Tertinggi (MPT) pada 24 Februari lalu.
Justeru, tidak timbul soal peletakan jawatan itu sah atau tidak kerana ia sudah pun diputuskan pada peringkat parti yang dipersetujui semua termasuk Presiden, Tan Sri <NAME>.
Bekas Setiausaha Agung Bersatu Datuk <NAME> berkata, pada mesyuarat itu MPT sebulat suara menolak peletakan jawatan Dr Mahathir.
"Jadi ini agak berlawanan dengan keputusan yang kita sudah buat. Saya tak faham bagaimana Jabatan Pendaftar Pertubuhan Malaysia (JPPM) kata peletakan jawatan itu sah sedangkan kita sudah buat keputusan di dalam mesyuarat, bukan seorang dua yang buat keputusan.
"Semua keputusan mesti dibuat melalui parti. Walau apa juga perbincangan dibuat di luar daripada keputusan mesyuarat, ini bukan keputusan parti.
"Apa locus standy yang ada pada Setiausaha Kerja untuk membawa perkara ini kepada JPPM. Seharusnya ia dibawa kepada Setiausaha Agung sebagai pentadbir kepada parti," katanya kepada Harian Metro.
Beliau mengulas laporan media tempatan hari ini mengenai pengesahan JPPM bahawa Dr Mahathir tidak lagi menjadi Pengerusi Bersatu berikutan peletakan jawatannya di tengah-tengah pergolakan politik pada akhir Februari adalah sah.
Laporan itu juga menyatakan, kedudukan <NAME> memangku jawatan itu juga sah.
Menurutnya, memang betul Dr Mahathir menghantar surat peletakan jawatan, tetapi ditolak oleh MPT.
"Fasal yang disebut itu terpakai sekiranya berhenti atau diberhentikan, tetapi ini mesyuarat sudah menolak," katanya.
Marzuki turut mempersoal kenyataan media yang dibuat beberapa pimpinan parti itu hari ini yang menyatakan sokongan kepada Perikatan Nasional.
"Kenyataan media bukanlah keputusan rasmi. Walaupun kita buat 1,000 kenyataan sekali pun ia tetap tidak merubah keputusan yang sudah dibuat di dalam mesyuarat. Kita catat di dalam minit apa yang berlaku di dalam mesyuarat," katanya.
"""
string = cleaning(string)
# -
# **Link**: https://www.malaysiakini.com/news/525953
#
# **Title**: Mahathir jangan hipokrit isu kes mahkamah Riza, kata Takiyuddin
#
# **Body**: Menteri undang-undang <NAME> berkata kerajaan berharap Dr <NAME> tidak bersikap hipokrit dengan mengatakan beliau tertanya-tanya dan tidak faham dengan keputusan mahkamah melepas tanpa membebaskan (DNAA) Riza Aziz, anak tiri bekas perdana menteri Najib Razak, dalam kes pengubahan wang haram membabitkan dana 1MDB.
#
# Pemimpin PAS itu berkata ini kerana keputusan itu dibuat oleh peguam negara dan dilaksanakan oleh timbalan pendakwa raya yang mengendalikan kes tersebut pada akhir 2019.
#
# “Saya merujuk kepada kenyataan Dr Mahathir tentang tindakan <NAME> memberikan pelepasan tanpa pembebasan (discharge not amounting to acquittal) kepada Riza Aziz baru-baru ini.
#
# “Kerajaan berharap Dr Mahathir tidak bersikap hipokrit dengan mengatakan beliau ‘tertanya-tanya’, keliru dan tidak faham terhadap suatu keputusan yang dibuat oleh <NAME> dan dilaksanakan oleh <NAME> yang mengendalikan kes ini pada akhir tahun 2019,” katanya dalam satu kenyataan hari ini.
#
# Riza pada Khamis dilepas tanpa dibebaskan daripada lima pertuduhan pengubahan wang berjumlah AS$248 juta (RM1.08 bilion).
#
# Dalam persetujuan yang dicapai antara pihak Riza dan pendakwaan, beliau dilepas tanpa dibebaskan atas pertuduhan itu dengan syarat memulangkan semula aset dari luar negara dengan nilai anggaran AS$107.3 juta (RM465.3 juta).
#
# Ekoran itu, Mahathir antara lain menyuarakan kekhuatirannya berkenaan persetujuan itu dan mempersoalkan jika pihak yang didakwa atas tuduhan mencuri boleh terlepas daripada tindakan jika memulangkan semula apa yang dicurinya.
#
# "Dia curi berbilion-bilion...Dia bagi balik kepada kerajaan. Dia kata kepada kerajaan, 'Nah, duit yang aku curi. Sekarang ini, jangan ambil tindakan terhadap aku.' Kita pun kata, 'Sudah bagi balik duit okey lah'," katanya.
#
# Menjelaskan bahawa beliau tidak mempersoalkan keputusan mahkamah, Mahathir pada masa sama berkata ia menunjukkan undang-undang mungkin perlu dipinda.
#
# Mengulas lanjut, Takiyuddin yang juga setiausaha agung PAS berkata
# kenyataan Mahathir tidak munasabah sebagai bekas perdana menteri.
#
# "Kerajaan berharap Dr Mahathir tidak terus bertindak mengelirukan rakyat dengan mengatakan beliau ‘keliru’.
#
# “Kerajaan PN akan terus bertindak mengikut undang-undang dan berpegang kepada prinsip kebebasan badan kehakiman dan proses perundangan yang sah,” katanya.
# +
string2 = """
Menteri undang-undang <NAME> berkata kerajaan berharap Dr <NAME> tidak bersikap hipokrit dengan mengatakan beliau tertanya-tanya dan tidak faham dengan keputusan mahkamah melepas tanpa membebaskan (DNAA) Riza Aziz, anak tiri bekas perdana menteri Najib Razak, dalam kes pengubahan wang haram membabitkan dana 1MDB.
Pemimpin PAS itu berkata ini kerana keputusan itu dibuat oleh peguam negara dan dilaksanakan oleh timbalan pendakwa raya yang mengendalikan kes tersebut pada akhir 2019.
“Saya merujuk kepada kenyataan Dr Mahathir tentang tindakan <NAME>yen memberikan pelepasan tanpa pembebasan (discharge not amounting to acquittal) kepada Riza Aziz baru-baru ini.
“Kerajaan berharap Dr Mahathir tidak bersikap hipokrit dengan mengatakan beliau ‘tertanya-tanya’, keliru dan tidak faham terhadap suatu keputusan yang dibuat oleh Peguam Negara dan dilaksanakan oleh Timbalan Pendakwa Raya yang mengendalikan kes ini pada akhir tahun 2019,” katanya dalam satu kenyataan hari ini.
Riza pada Khamis dilepas tanpa dibebaskan daripada lima pertuduhan pengubahan wang berjumlah AS$248 juta (RM1.08 bilion).
Dalam persetujuan yang dicapai antara pihak Riza dan pendakwaan, beliau dilepas tanpa dibebaskan atas pertuduhan itu dengan syarat memulangkan semula aset dari luar negara dengan nilai anggaran AS$107.3 juta (RM465.3 juta).
Ekoran itu, Mahathir antara lain menyuarakan kekhuatirannya berkenaan persetujuan itu dan mempersoalkan jika pihak yang didakwa atas tuduhan mencuri boleh terlepas daripada tindakan jika memulangkan semula apa yang dicurinya.
"Dia curi berbilion-bilion...Dia bagi balik kepada kerajaan. Dia kata kepada kerajaan, 'Nah, duit yang aku curi. Sekarang ini, jangan ambil tindakan terhadap aku.' Kita pun kata, 'Sudah bagi balik duit okey lah'," katanya.
Menjelaskan bahawa beliau tidak mempersoalkan keputusan mahkamah, Mahathir pada masa sama berkata ia menunjukkan undang-undang mungkin perlu dipinda.
Mengulas lanjut, Takiyuddin yang juga setiausaha agung PAS berkata
kenyataan Mahathir tidak munasabah sebagai bekas perdana menteri.
"Kerajaan berharap Dr Mahathir tidak terus bertindak mengelirukan rakyat dengan mengatakan beliau ‘keliru’.
“Kerajaan PN akan terus bertindak mengikut undang-undang dan berpegang kepada prinsip kebebasan badan kehakiman dan proses perundangan yang sah,” katanya.
"""
string2 = cleaning(string2)
# -
# ### List available Transformer models
malaya.summarization.abstractive.available_transformer()
# 1. `t2t` is multitasks Transformer using Tensor2Tensor library.
# 2. `t5` is multitasks Transformer from T5 paper.
# 3. `pegasus` is Sentence Gap Pegasus.
# 4. `bigbird` is Finetuning Sentence Gap Pegasus.
# ### Load T5 Transformer
#
# ```python
# def transformer(model: str = 't2t', quantized: bool = False, **kwargs):
#
# """
# Load Malaya transformer encoder-decoder model to generate a summary given a string.
#
# Parameters
# ----------
# model : str, optional (default='t2t')
# Model architecture supported. Allowed values:
#
# * ``'t2t'`` - Malaya Transformer BASE parameters.
# * ``'small-t2t'`` - Malaya Transformer SMALL parameters.
# * ``'t2t-distill'`` - Distilled Malaya Transformer BASE parameters.
# * ``'t5'`` - T5 BASE parameters.
# * ``'small-t5'`` - T5 SMALL parameters.
# * ``'bigbird'`` - BigBird + Pegasus BASE parameters.
# * ``'small-bigbird'`` - BigBird + Pegasus SMALL parameters.
# * ``'pegasus'`` - Pegasus BASE parameters.
# * ``'small-pegasus'`` - Pegasus SMALL parameters.
#
# quantized : bool, optional (default=False)
# if True, will load 8-bit quantized model.
# Quantized model not necessary faster, totally depends on the machine.
#
# Returns
# -------
# result: model
# List of model classes:
#
# * if `t2t` in model, will return `malaya.model.tf.Summarization`.
# * if `t5` in model, will return `malaya.model.t5.Summarization`.
# * if `bigbird` in model, will return `malaya.model.bigbird.Summarization`.
# * if `pegasus` in model, will return `malaya.model.pegasus.Summarization`.
# """
# ```
#
# For T5, I am going to use `quantized` model, `normal` model is too big, 1.25GB.
# +
# # !pip3 install install tensorflow-text==1.15.1
# -
t5 = malaya.summarization.abstractive.transformer(model = 't5', quantized = True)
# ### Load T2T Transformer
#
# ```python
# def transformer(model: str = 't2t', quantized: bool = False, **kwargs):
#
# """
# Load Malaya transformer encoder-decoder model to generate a summary given a string.
#
# Parameters
# ----------
# model : str, optional (default='t2t')
# Model architecture supported. Allowed values:
#
# * ``'t2t'`` - Malaya Transformer BASE parameters.
# * ``'small-t2t'`` - Malaya Transformer SMALL parameters.
# * ``'t2t-distill'`` - Distilled Malaya Transformer BASE parameters.
# * ``'t5'`` - T5 BASE parameters.
# * ``'small-t5'`` - T5 SMALL parameters.
# * ``'bigbird'`` - BigBird + Pegasus BASE parameters.
# * ``'small-bigbird'`` - BigBird + Pegasus SMALL parameters.
# * ``'pegasus'`` - Pegasus BASE parameters.
# * ``'small-pegasus'`` - Pegasus SMALL parameters.
#
# quantized : bool, optional (default=False)
# if True, will load 8-bit quantized model.
# Quantized model not necessary faster, totally depends on the machine.
#
# Returns
# -------
# result: model
# List of model classes:
#
# * if `t2t` in model, will return `malaya.model.tf.Summarization`.
# * if `t5` in model, will return `malaya.model.t5.Summarization`.
# * if `bigbird` in model, will return `malaya.model.bigbird.Summarization`.
# * if `pegasus` in model, will return `malaya.model.pegasus.Summarization`.
# """
# ```
model = malaya.summarization.abstractive.transformer(model = 'small-t2t')
# #### Load Quantized model
#
# To load 8-bit quantized model, simply pass `quantized = True`, default is `False`.
#
# We can expect slightly accuracy drop from quantized model, and not necessary faster than normal 32-bit float model, totally depends on machine.
quantized_model = malaya.summarization.abstractive.transformer(model = 'small-t2t', quantized = True)
# #### summarization mode
#
# in Malaya provided 2 different modes for summarization,
#
# 1. generate summary,
#
# ```python
# model.greedy_decoder(strings, mode = 'ringkasan')
# ```
#
# 2. generate title,
#
# ```python
# model.greedy_decoder(strings, mode = 'tajuk')
# ```
#
# default is `ringkasan`. **`pegasus` and `bigbird` do not have summarization mode**.
# #### Predict using greedy decoder
#
# ```python
# def greedy_decoder(
# self,
# strings: List[str],
# mode: str = 'ringkasan',
# postprocess: bool = True,
# **kwargs,
# ):
# """
# Summarize strings using greedy decoder.
#
# Parameters
# ----------
# strings: List[str]
# mode: str
# mode for summarization. Allowed values:
#
# * ``'ringkasan'`` - summarization for long sentence, eg, news summarization.
# * ``'tajuk'`` - title summarization for long sentence, eg, news title.
# postprocess: bool, optional (default=True)
# If True, will filter sentence generated using ROUGE score and removed international news publisher.
#
# Returns
# -------
# result: List[str]
# """
# ```
#
# **For T5, we only provided `greedy_decoder` method to predict.**
#
# **`pegasus` and `bigbird` do not have summarization mode**.
pprint(t5.greedy_decoder([string], mode = 'tajuk'))
pprint(t5.greedy_decoder([string2], mode = 'tajuk'))
pprint(model.greedy_decoder([string], mode = 'tajuk'))
pprint(quantized_model.greedy_decoder([string], mode = 'tajuk'))
pprint(t5.greedy_decoder([string], mode = 'ringkasan'))
pprint(t5.greedy_decoder([string2], mode = 'ringkasan'))
pprint(model.greedy_decoder([string], mode = 'ringkasan'))
pprint(quantized_model.greedy_decoder([string], mode = 'ringkasan'))
# #### Predict using beam decoder
#
# ```python
# def beam_decoder(
# self,
# strings: List[str],
# mode: str = 'ringkasan',
# postprocess: bool = True,
# **kwargs,
# ):
# """
# Summarize strings using beam decoder, beam width size 3, alpha 0.5 .
#
# Parameters
# ----------
# strings: List[str]
# mode: str
# mode for summarization. Allowed values:
#
# * ``'ringkasan'`` - summarization for long sentence, eg, news summarization.
# * ``'tajuk'`` - title summarization for long sentence, eg, news title.
# postprocess: bool, optional (default=True)
# If True, will filter sentence generated using ROUGE score and removed international news publisher.
#
# Returns
# -------
# result: List[str]
# """
# ```
#
# **`pegasus` and `bigbird` do not have summarization mode**.
pprint(model.beam_decoder([string], mode = 'tajuk'))
pprint(quantized_model.beam_decoder([string], mode = 'tajuk'))
pprint(model.beam_decoder([string], mode = 'ringkasan'))
pprint(quantized_model.beam_decoder([string], mode = 'ringkasan'))
# #### Predict using nucleus decoder
#
# ```python
# def nucleus_decoder(
# self,
# strings: List[str],
# mode: str = 'ringkasan',
# top_p: float = 0.7,
# postprocess: bool = True,
# **kwargs,
# ):
# """
# Summarize strings using nucleus sampling.
#
# Parameters
# ----------
# strings: List[str]
# mode: str
# mode for summarization. Allowed values:
#
# * ``'ringkasan'`` - summarization for long sentence, eg, news summarization.
# * ``'tajuk'`` - title summarization for long sentence, eg, news title.
# top_p: float, (default=0.7)
# cumulative distribution and cut off as soon as the CDF exceeds `top_p`.
# postprocess: bool, optional (default=True)
# If True, will filter sentence generated using ROUGE score and removed international news publisher.
#
# Returns
# -------
# result: List[str]
# """
# ```
#
# **`pegasus` and `bigbird` do not have summarization mode**.
pprint(model.nucleus_decoder([string], mode = 'tajuk'))
pprint(model.nucleus_decoder([string], mode = 'ringkasan'))
| example/abstractive-summarization/load-abstractive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv("Users.csv")
df.head()
df.shape
# Novice, Contributor, Expert, Master, and Grandmaster.
# 6 unique values representing user's level on Kaggle's progression system
df.PerformanceTier.nunique()
df.groupby("PerformanceTier")["DisplayName"].nunique().rename(index = {0 : "Novice"})
df[df.PerformanceTier == 5]["DisplayName"]
df[df.PerformanceTier == 4]["DisplayName"]
df[df.PerformanceTier == 3]["DisplayName"]
df[df.PerformanceTier == 2]["DisplayName"]
df[df.PerformanceTier == 1]["DisplayName"]
df[df.PerformanceTier == 0]["DisplayName"]
| DataScience365/Day 6/Kaggle user info.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example Angle Comparison
#
# At times it may be useful to confirm that the spatial transform solution returned by PINK is sensible. In this notebook we perform our derivation of the spatial transform for an image to its corresponding best matching neuron.
# +
from concurrent.futures import ProcessPoolExecutor
import matplotlib.pyplot as plt
plt.rcParams.update({'figure.max_open_warning': 0})
import numpy as np
from tqdm import tqdm
from scipy.ndimage import rotate
import pyink as pu
# -
# ## Datasets
#
# A set of SOMs were trained against ~24,000 images with images from EMU and WISE W1 surveys. The input image size was 5 arcminutes. Below are the base file names that will be used to load in:
# - preprocessed training images provided to PINK
# - the SOM, mapping and spatial transform data files produced by PINK
#
# The `PathHelper` is a simple utility class to help make folders and sub-folders without to much difficulty.
#
# +
path = pu.PathHelper('Example_Images', clobber=True)
suffix = 'B3Circular_h45_w45_emu.bin'
som = pu.SOM(f'../../EMU_WISE_E95E05_Aegean_Components_Complex_EMUWISE_IslandNorm_Log_Reprojected/SOMs/SOM_{suffix}')
transform = pu.Transform(f'../../EMU_WISE_E95E05_Aegean_Components_Complex_EMUWISE_IslandNorm_Log_Reprojected/SOMs/TRANSFORM_{suffix}')
mapper = pu.Mapping(f'../../EMU_WISE_E95E05_Aegean_Components_Complex_EMUWISE_IslandNorm_Log_Reprojected/SOMs/MAP_{suffix}')
image_binary = '../../EMU_WISE_E95E05_Aegean_Components_Complex_EMUWISE_IslandNorm_Log_Reprojected/EMU_WISE_E95E05_Aegean_Components_Complex_EMUWISE_IslandNorm_Log_Reprojected.bin'
imgs = pu.ImageReader(image_binary)
# -
# Note that the number of valid pixels in each of the masks is the same. This has to be done to ensure a correct pixelwise euclidean distance summation. At the moment a radius of 75 produces an inconsistent number of valid pixels. This is being looked into and is likely a rounding error (if you spot it please let me know :) )
# +
src_img = imgs.data[1, 0]
bmu_min = mapper.bmu(1)
bmu_img = som[bmu_min][0]
src_mask = pu.circular_mask(src_img, radius=74)
bmu_mask = pu.circular_mask(bmu_img, radius=74)
fig, (ax1, ax2) = plt.subplots(1,2)
ax1.imshow(src_mask)
ax2.imshow(bmu_mask)
print('Valid number of pixels: ', np.sum(src_mask), np.sum(bmu_mask))
# -
# # Angle distribution
#
# The spatial transforms return by `PINK` should be, essentially, random. Here a few quick tests are performed to ensure that they behave as expected. This includes manually finding the optimal spatial transform solution between a given image a neuron to see if it agrees with the solution found by `PINK`.
# +
fig, ax = plt.subplots(1,1, figsize=(10,5))
bmus = mapper.bmu()[:]
datas = transform.data[:][np.arange(transform.data.shape[0]), bmus[:, 0], bmus[:, 1]]
ax.hist(datas['angle'].flatten(), bins=100)
ax.set(xlabel='Angle (radians / float32)')
fig.savefig(f'{path}/Angles.png')
# -
def pink_rotation(src, angle):
"""A pure python translation of the C++ code used by PINK.
"""
width_margin = 0
height_margin = 0
rot = np.zeros_like(src)
x0 = src.shape[1] // 2
y0 = src.shape[0] // 2
cos_alpha = np.cos(angle)
sin_alpha = np.sin(angle)
for x2 in range(src.shape[1]):
for y2 in range(src.shape[0]):
x1 = (x2 + width_margin - x0) * cos_alpha + (y2 + height_margin - y0) * sin_alpha + x0 + 0.1
if x1 < 0 or int(x1) >= src.shape[1]:
rot[y2, x2] = 0
continue
y1 = (y2 + height_margin - y0) * cos_alpha - (x2 + width_margin - x0) * sin_alpha + y0 + 0.1
if y1 < 0 or int(y1) >= src.shape[0]:
rot[y2, x2] = 0
continue
ix1 = int(x1)
iy1 = int(y1)
ix1b = int(ix1 + 1)
iy1b = int(iy1 + 1)
rx1 = int(x1 - ix1)
ry1 = int(y1 - iy1)
cx1 = int(1. - rx1)
cy1 = int(1. - ry1)
if max(iy1, ix1, iy1b, ix1b) >= 150:
continue
rot[y2, x2] = cx1 * cy1 * src[iy1, ix1] +\
cx1 * ry1 * src[iy1b, ix1] +\
rx1 * cy1 * src[iy1, ix1b] +\
rx1 * ry1 * src[iy1b, ix1b]
return rot
# +
def euclidean_dist(img_a, img_b):
return np.sum((img_a - img_b)**2.)
def manual_transform(img_idx, imgs, mapper, transform, som, rot_axes=(1,0),
pink_rotate=False, verbose=True, plot=True):
"""Given a best matching neuron and an image, compute the best matching spatial transform outside of PINK.
"""
src_img = imgs.data[img_idx, 0]
bmu_img = som[bmu_min][0]
bmu_trans = transform.data[(img_idx, *bmu_min)]
bmu_ed = mapper.data[(img_idx, *bmu_min)]
src_mask = pu.square_mask(src_img, 105)
bmu_mask = pu.square_mask(bmu_img, 105)
src_mask = pu.circular_mask(src_img, radius=74)
bmu_mask = pu.circular_mask(bmu_img, radius=74)
no_rots = 360
do_flip = True
residuals = np.zeros(no_rots * (do_flip+1))
radian_step = 2*np.pi / no_rots
offset = no_rots // 4
for i in range(offset):
ang = radian_step * i
if pink_rotate:
rot_img = pink_rotation(src_img, ang)
else:
rot_img = rotate(src_img, -np.rad2deg(ang), reshape=False, order=1)
residuals[i] = euclidean_dist(rot_img[src_mask], bmu_img[bmu_mask])
rot_img_90 = np.rot90(rot_img, axes=rot_axes)
residuals[i+offset] = euclidean_dist(rot_img_90[src_mask], bmu_img[bmu_mask])
rot_img_180 = np.rot90(rot_img_90, axes=rot_axes)
residuals[i+(2*offset)] = euclidean_dist(rot_img_180[src_mask], bmu_img[bmu_mask])
rot_img_270 = np.rot90(rot_img_180, axes=rot_axes)
residuals[i+(3*offset)] = euclidean_dist(rot_img_270[src_mask], bmu_img[bmu_mask])
if do_flip:
flip_img = rot_img[::-1]
residuals[i+no_rots] = euclidean_dist(flip_img[src_mask], bmu_img[bmu_mask])
flip_img_90 = rot_img_90[::-1]
residuals[i+offset+no_rots] = euclidean_dist(flip_img_90[src_mask], bmu_img[bmu_mask])
flip_img_180 = rot_img_180[::-1]
residuals[i+(2*offset)+no_rots] = euclidean_dist(flip_img_180[src_mask], bmu_img[bmu_mask])
flip_img_270 = rot_img_270[::-1]
residuals[i+(3*offset)+no_rots] = euclidean_dist(flip_img_270[src_mask], bmu_img[bmu_mask])
arg_min = np.argmin(residuals)
flip = arg_min // 360
rot = (arg_min % no_rots) * radian_step
if verbose:
print(img_idx)
print("\tManual:", flip, rot)
print("\tPINK :", bmu_trans)
print("\tDiff :", flip - bmu_trans[0], rot - bmu_trans[1])
if plot:
fig, ax = plt.subplots(1,1)
idx = np.arange(no_rots)
ax.plot(idx * radian_step, residuals[idx], 'r-', label='No flip')
if do_flip:
idx = np.arange(no_rots, 2*no_rots)
ax.plot((idx-no_rots)*radian_step, residuals[idx], 'g:', label='With flip')
if flip:
ax.axvline(rot, ls=':', color='green', label='Manual Minimised ED Position')
else:
ax.axvline(rot, ls='-', color='red', label='Manual Minimised ED Position')
if bmu_trans[0] == 1:
ax.axvline(bmu_trans[1], ls=':', lw=5, color='Pink', label='PINK Minimised ED Position')
else:
ax.axvline(bmu_trans[1], ls='-', lw=5, color='Pink', label='PINK Minimised ED Position')
ax.legend(loc='upper right')
ax.set(xlabel="Angle (Radian)", ylabel="Residual")
fig.show()
cp_img = src_img.copy()
cp_img = rotate(cp_img, -np.rad2deg(rot), reshape=False)
if flip:
cp_img = cp_img[::-1, :]
pink_img = src_img.copy()
pink_img = rotate(pink_img, -np.rad2deg(bmu_trans[1]), reshape=False, order=2)
if bmu_trans[0] == 1:
pink_img = pink_img[::-1, :]
if plot:
fig, ax = plt.subplots(1,3, figsize=(15, 4))
ax[0].imshow(cp_img)
ax[0].set(title='Manual Transform')
ax[0].grid(which='major', axis='both')
ax[1].imshow(bmu_img[31:-31, 31:-31])
ax[1].set(title='BMU')
ax[1].grid(which='major', axis='both')
ax[2].imshow(pink_img)
ax[2].set(title='PINK Transform')
ax[2].grid(which='major', axis='both')
fig.show()
return flip, rot, rot - bmu_trans[1], bmu_trans[0], bmu_trans[1], bmu_ed
# +
def manual_transform_lambda(i):
return manual_transform(i, imgs, mapper, transform, som, verbose=False, plot=False)
samples = 1000
workers = 8
with ProcessPoolExecutor(max_workers=workers) as executor:
results = list(tqdm(executor.map(manual_transform_lambda, np.arange(samples), chunksize=samples//workers//4), total=samples))
# +
res = np.array(results)
fig, ax = plt.subplots(1,1)
ax.hist(res[:,2], bins=100)
tick_lines = np.deg2rad(30)*np.arange(-13, 13)
ax.set_xticks(tick_lines, minor=True)
ax.grid(True, axis='both', which='minor')
ax.set_xlabel('radians')
fig.tight_layout()
fig.savefig(f"{path}/residual_angle.png")
# -
# The spatial transform solutions derived between PINK and the python code for almost all sources should agree on a 1-to-1 level. You may notice that there are a set of sources that fall off this 1-to-1 one line. This can be tested below.
# +
res = np.array(results)
fig, ax = plt.subplots(1,1)
cim = ax.scatter(res[:, 1], res[:, 4], c=res[:,0], s=26)
ax.set(xlabel='Manual Rotation', ylabel='PINK Rotation')
tick_lines = np.deg2rad(30)*np.arange(-1, 13)
ax.set_xticks(tick_lines, minor=True)
ax.set_yticks(tick_lines, minor=True)
ax.grid(True, axis='both', which='minor')
one = np.linspace(0, 2*np.pi)
ax.plot(one, one, 'r-')
fig.colorbar(cim, label='Manual Flip')
fig.tight_layout()
fig.savefig(f'{path}/Manual_PINK_Angle.png')
# -
# You may also notice that this are largely consistent with whether an image was flipped or not flipped. Generally for these sources the euclidean distances are approximately the same between two competing spatial transform solutions, and what determines the minimum is influenced by numerical precision errors.
# +
outlier = np.argwhere(np.abs(res[:,1] - res[:,4]) > np.pi )
manual_transform(outlier[0][0], imgs, mapper, transform, som, verbose=False, plot=True)
# -
| Example_Notebooks/Example_Angle_Comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests # Библиотека для отправки запросов
import numpy as np # Библиотека для матриц, векторов и линала
import pandas as pd # Библиотека для табличек
import time # Библиотека для времени
from bs4 import BeautifulSoup #Библиотека для парсинга
from fake_useragent import UserAgent #Библиотека для фейковых данных
from tqdm.notebook import tqdm #Библиотека для красивого отслеживания цикла
def get_page_links(page_number):
page_link = 'https://realt.by/sale/flats/?search=eJxFjEEOwiAURE9D1x8otixcSLnHD5ZPJWpLKMbrC2zcTGbeTOYsrhBm2uKxY%2FRMGWLKXtVQju%2BOu3vXaF7x2Wh10EpmgemlqZm7WmY5m3n3twFPKp9Ux5lWTJQxua2fSvh3jwboIqTyq9I8AJEPIwSYtBiF5kJO4v4Dfn0tNw%3D%3D&page={}'.format(page_number)
response = requests.get(page_link, headers={'User-Agent': UserAgent().chrome})
if not response.ok:
return []
html = response.content
soup = BeautifulSoup(html,'html.parser')
flat = soup.findAll('div', {"class": "teaser-tile teaser-tile-right"})
links = [link.a["href"] for link in flat]
return links
def get_flat_data(flat_page):
response = requests.get(flat_page, headers={'User-Agent': UserAgent().chrome})
if not response.ok:
return response.status_code
html = response.content
soup = BeautifulSoup(html,'html.parser')
features = soup.findAll('table', {'class' : 'table-params table-borderless table-striped mb-0'})
table_df_first = pd.read_html(str(features))[0]
table_df_second = pd.read_html(str(features))[1]
combined_data = table_df_first.append(table_df_second).reset_index(drop=True)
price_class = soup.find('div', {'class' : 'd-flex align-items-center fs-giant'})
price = ['Цена', str(price_class.strong.contents[0])]
combined_data.loc[len(combined_data)] = price
combined_data = combined_data.T
combined_data.columns = combined_data.iloc[0]
combined_data.drop(0,axis=0,inplace=True)
return combined_data
def finish_data():
finish_data_df = get_flat_data(get_page_links(0)[0])
for j in tqdm(range(481)):
for i in tqdm(get_page_links(j)):
try:
data = get_flat_data(i)
finish_data_df = pd.concat([finish_data_df, data], ignore_index=True)
except:
print("Error! can't parsing this page or flat doesn't have a price:", i)
finish_data_df.drop(0,axis=0)
return finish_data_df
finish_data = finish_data()
finish_data.to_csv('flat_research_minsk.csv', index=False)
finish_data.head()
| flat_parser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# +
import matplotlib.pyplot as plt
import os
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import FunctionTransformer, LabelBinarizer
from sklearn.model_selection import StratifiedKFold
import tensorflow as tf
import pickle
import numpy as np
import pandas as pd
from random import choice
from annsa.template_sampling import *
# -
import tensorflow.contrib.eager as tfe
tf.enable_eager_execution()
# #### Import model, training function
from annsa.model_classes import (dnn_model_features,
DNN,
save_model,
train_earlystop)
# ## Dataset Construction
# #### Load dataset
background_dataset = pd.read_csv('../../source-interdiction/training_testing_data/background_template_dataset.csv')
source_dataset = pd.read_csv('../../source-interdiction/training_testing_data/shielded_templates_200kev_dataset.csv')
# #### Dataset details
print('sourcedist: ' + str(sorted(set(source_dataset['sourcedist']))))
print('sourceheight: ' + str(sorted(set(source_dataset['sourceheight']))))
print('alum shieldingdensity: ' + str(sorted(set(source_dataset[source_dataset['shielding']=='alum']['shieldingdensity']))))
print('iron shieldingdensity: ' + str(sorted(set(source_dataset[source_dataset['shielding']=='iron']['shieldingdensity']))))
print('lead shieldingdensity: ' + str(sorted(set(source_dataset[source_dataset['shielding']=='lead']['shieldingdensity']))))
print('fwhm: ' + str(sorted(set(source_dataset['fwhm']))))
# #### Customize dataset
# +
source_dataset = source_dataset[(source_dataset['fwhm']==7.0) |
(source_dataset['fwhm']==7.5) |
(source_dataset['fwhm']==8.0)]
source_dataset = source_dataset[(source_dataset['sourcedist']==50.5) |
(source_dataset['sourcedist']==175.0) |
(source_dataset['sourcedist']==300.0)]
source_dataset = source_dataset[(source_dataset['sourceheight']==50.0) |
(source_dataset['sourceheight']==100.0) |
(source_dataset['sourceheight']==150.0)]
# remove 80% shielding
source_dataset = source_dataset[source_dataset['shieldingdensity']!=13.16]
source_dataset = source_dataset[source_dataset['shieldingdensity']!=11.02]
source_dataset = source_dataset[source_dataset['shieldingdensity']!=1.61]
# -
# #### Remove empty spectra
# +
zero_count_indicies = np.argwhere(np.sum(source_dataset.values[:,6:],axis=1) == 0).flatten()
print('indicies dropped: ' +str(zero_count_indicies))
source_dataset.drop(source_dataset.index[zero_count_indicies], inplace=True)
# -
# #### Add empty spectra for background
# +
blank_spectra = []
for fwhm in set(source_dataset['fwhm']):
num_examples = source_dataset[(source_dataset['fwhm']==fwhm) &
(source_dataset['isotope']==source_dataset['isotope'].iloc()[0])].shape[0]
for k in range(num_examples):
blank_spectra_tmp = [0]*1200
blank_spectra_tmp[5] = fwhm
blank_spectra_tmp[0] = 'background'
blank_spectra_tmp[3] = 'background'
blank_spectra.append(blank_spectra_tmp)
source_dataset = source_dataset.append(pd.DataFrame(blank_spectra,
columns=source_dataset.columns))
# -
# #### Create dataset from spectra
spectra_dataset = source_dataset.values[:,5:].astype('float64')
all_keys = source_dataset['isotope'].values
# ## Define Training Parameters
# #### Define online data augmentation
# +
def integration_time():
return np.random.uniform(10,600)
def background_cps():
return np.random.poisson(200)
def signal_to_background():
return np.random.uniform(0.5,2)
def calibration():
return [np.random.uniform(0,20),
np.random.uniform(2500/3000,3500/3000),
0]
online_data_augmentation = online_data_augmentation_vanilla(background_dataset,
background_cps,
integration_time,
signal_to_background,
calibration,)
# -
# ## Load testing dataset
testing_dataset = np.load('../dataset_generation/testing_dataset_full_200keV_1000.npy')
# +
testing_spectra = np.add(testing_dataset.item()['sources'], testing_dataset.item()['backgrounds'])
testing_keys = testing_dataset.item()['keys']
mlb=LabelBinarizer()
all_keys_binarized = mlb.fit_transform(all_keys.reshape([all_keys.shape[0],1]))
testing_keys_binarized = mlb.transform(testing_keys)
training_keys_binarized = mlb.transform(all_keys)
# -
# # Train network
# ### Define hyperparameters
number_hyperparameters_to_search = 256
earlystop_errors_test = []
model_id='DNN-onlinedataaugfull'
def make_model():
'''
Makes a random model given some parameters.
'''
number_layers = choice([1, 2, 3])
dense_nodes = 2**np.random.randint(5, 10, number_layers)
dense_nodes = np.sort(dense_nodes)
dense_nodes = np.flipud(dense_nodes)
model_features = dnn_model_features(
learining_rate=10**np.random.uniform(-4,-1),
l2_regularization_scale=10**np.random.uniform(-2,0),
dropout_probability=np.random.uniform(0,1),
batch_size=2**np.random.randint(4,10),
output_size=training_keys_binarized.shape[1],
dense_nodes=dense_nodes,
activation_function=choice([tf.nn.tanh,tf.nn.relu,tf.nn.sigmoid]),
scaler=choice([make_pipeline(FunctionTransformer(np.log1p, validate=True)),
make_pipeline(FunctionTransformer(np.sqrt, validate=True))]))
model = DNN(model_features)
return model, model_features
# ### Search hyperparameters
# +
testing_errors = []
for network_id in range(number_hyperparameters_to_search):
# reset model on each iteration
model, model_features = make_model()
model_features.activation_function = tf.nn.sigmoid
model_features.batch_size = 32
model_features.dense_nodes = [256]
model_features.dropout_probability = 0.63
model_features.l2_regularization_scale = 0.04293
model_features.learining_rate = 0.0001212
model_features.scaler = make_pipeline(FunctionTransformer(np.sqrt, validate=True))
optimizer = tf.train.AdamOptimizer(model_features.learining_rate)
costfunction_errors_tmp, earlystop_errors_tmp = train_earlystop(
training_data=spectra_dataset,
training_keys=training_keys_binarized,
testing_data=testing_spectra,
testing_keys=testing_keys_binarized,
model=model,
optimizer=optimizer,
num_epochs=500,
obj_cost=model.cross_entropy,
earlystop_cost_fn=model.f1_error,
earlystop_patience=10,
verbose=True,
fit_batch_verbose=1,
data_augmentation=online_data_augmentation)
testing_errors.append(earlystop_errors_tmp)
# np.save('./final-models/final_test_errors_'+model_id, training_errors)
# model.save_weights('./final-models/'+model_id+'_checkpoint_'+str(network_id))
network_id += 1
# -
| examples/source-interdiction/hyperparameter-search/DNNHPSearch-OnlineDataAugmentation-Full.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Инициализация
import os
import numpy as np
import pymorphy2
from tqdm import tqdm_notebook
token_filenames = [f for f in os.listdir('./devset/') if '.tokens' in f]
# # Функции для работы с файлами
class Token:
def __init__(self, position, length, text):
self._position = position
self._length = length
self._text = text
self._pos = None
self._tag = None
class Span:
def __init__(self, token_id):
self._token_id = token_id
def load_tokens(token_filename, dev=True):
_set = './devset/' if dev else './testset/'
tokens = dict()
with open(_set + token_filename, encoding='utf8') as f:
for line in f:
split = line.split()
if len(split) > 0:
t = Token(split[1], split[2], split[3])
tokens[split[0]] = t
return tokens
def load_spans(token_filename, dev=True):
_set = './devset/' if dev else './testset/'
spans = dict()
with open(_set + token_filename.split('.')[0] + '.spans', encoding='utf8') as f:
for line in f:
split = line.split()
s = Span(split[4])
spans[split[0]] = s
return spans
def transform_base_tag(base_tag):
if base_tag == 'Person':
return 'PER'
if base_tag == 'Location' or base_tag == 'LocOrg':
return 'LOC'
if base_tag == 'Org':
return 'ORG'
else:
return 'MISC'
def load_objects(token_filename, tokens, spans, dev=True):
_set = './devset/' if dev else './testset/'
with open(_set + token_filename.split('.')[0] + '.objects', encoding='utf8') as f:
for line in f:
line = line.split(' # ')[0]
split = line.split()
base_tag = transform_base_tag(split[1])
span_ids = split[2:]
if len(span_ids) == 1:
tokens[spans[span_ids[0]]._token_id]._tag = 'U-' + base_tag
else:
for i, span_id in enumerate(span_ids):
if i == 0:
tokens[spans[span_ids[i]]._token_id]._tag = 'B-' + base_tag
if i == len(span_ids) - 1:
tokens[spans[span_ids[i]]._token_id]._tag = 'L-' + base_tag
else:
tokens[spans[span_ids[i]]._token_id]._tag = 'I-' + base_tag
return tokens
morph = pymorphy2.MorphAnalyzer()
def fill_pos(tokens):
for id, token in tokens.items():
pos = morph.parse(token._text)[0].tag.POS
if pos is None:
pos = 'None'
token._pos = pos
if token._tag is None:
token._tag = 'O'
return tokens
# +
def word2features(sent, i):
word = sent[i]._text
postag = sent[i]._pos
features = {
'bias': 1.0,
'word.lower()': word.lower(),
'word[-3:]': word[-3:],
'word[-2:]': word[-2:],
'word.isupper()': word.isupper(),
'word.istitle()': word.istitle(),
'word.isdigit()': word.isdigit(),
'postag': postag,
}
if i > 0:
word1 = sent[i-1]._text
postag1 = sent[i-1]._pos
features.update({
'-1:word.lower()': word1.lower(),
'-1:word.istitle()': word1.istitle(),
'-1:word.isupper()': word1.isupper(),
'-1:postag': postag1,
})
else:
features['BOS'] = True
if i < len(sent)-1:
word1 = sent[i+1]._text
postag1 = sent[i+1]._pos
features.update({
'+1:word.lower()': word1.lower(),
'+1:word.istitle()': word1.istitle(),
'+1:word.isupper()': word1.isupper(),
'+1:postag': postag1,
})
else:
features['EOS'] = True
return features
def sent2features(sent):
return [word2features(sent, i) for i in range(len(sent))]
def sent2labels(sent):
return [token._tag for token in sent]
def sent2tokens(sent):
return [token for token in sent]
# -
def split_tokens_by_sents(tokens):
sents = []
sent = []
for id, token in tokens.items():
if token._text != '.':
sent.append(token)
else:
sents.append(sent)
sent = []
return sents
# # Готовим тренировочную выборку
def generate_sents(token_filename):
tokens = load_tokens(token_filename)
spans = load_spans(token_filename)
tokens = load_objects(token_filename, tokens, spans)
tokens = fill_pos(tokens)
sents = split_tokens_by_sents(tokens)
return sents
sents = []
for token_filename in tqdm_notebook(token_filenames):
sents += generate_sents(token_filename)
len(sents)
# +
from sklearn.model_selection import train_test_split
import numpy as np
train_ids, test_ids = train_test_split(np.arange(len(sents)))
# -
train_sents = np.array(sents)[train_ids]
test_sents = np.array(sents)[test_ids]
# +
X_train = [sent2features(s) for s in train_sents]
y_train = [sent2labels(s) for s in train_sents]
X_test = [sent2features(s) for s in test_sents]
y_test = [sent2labels(s) for s in test_sents]
# -
# # Тренируем модель
# +
from sklearn_crfsuite import CRF
crf = CRF(algorithm='lbfgs', c1=0.1, c2=0.1, max_iterations=100, all_possible_transitions=True)
crf.fit(X_train, y_train)
# +
from sklearn_crfsuite.metrics import flat_classification_report
print(flat_classification_report(y_test, crf.predict(X_test)))
# -
# # Применяем модель
def get_entities(tokens):
rows = []
buffer = []
for id, token in tokens.items():
tag = token._tag
if tag.startswith('U'):
rows.append('%s %d %d\n' % (tag.split('-')[1], int(token._position), int(token._length)))
elif tag.startswith('B') or tag.startswith('I'):
buffer.append(token)
elif tag.startswith('L'):
buffer.append(token)
start = int(buffer[0]._position)
length = int(buffer[-1]._position) + int(buffer[-1]._length) - int(start)
rows.append('%s %d %d\n' % (tag.split('-')[1], start, length))
buffer = []
return rows
test_token_filenames = [filename for filename in os.listdir('./testset') if '.tokens' in filename]
for token_filename in tqdm_notebook(test_token_filenames):
tokens = load_tokens(token_filename, dev=False)
tokens = fill_pos(tokens)
sents = split_tokens_by_sents(tokens)
X = [sent2features(s) for s in sents]
y_pred = crf.predict(X)
for i in range(len(y_pred)):
for j in range(len(y_pred[i])):
sents[i][j]._tag = y_pred[i][j]
rows = get_entities(tokens)
with open('./results_crf/' + token_filename.split('.')[0] + '.task1', 'w') as f:
f.writelines(rows)
# # Проверяем результаты
# !python scripts\t1_eval.py -s .\testset -t .\results_crf -o .\output\
| NER CRF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Install dependencies for this example
# Note: This does not include itkwidgets, itself
import sys
# !{sys.executable} -m pip install --upgrade itk-io itk-cuberille itk-totalvariation
# +
try:
from urllib.request import urlretrieve
except ImportError:
from urllib import urlretrieve
import os
import numpy as np
import itk
from itkwidgets import view
import itkwidgets
# -
# Download data
# Source: https://data.broadinstitute.org/bbbc/BBBC024/
file_name = 'HL50_cell_line_c00_03_extraction.tif'
if not os.path.exists(file_name):
url = 'https://data.kitware.com/api/v1/file/5b61f16c8d777f06857c1949/download'
urlretrieve(url, file_name)
# +
image = itk.imread(file_name, itk.F)
view(image, gradient_opacity=0.4, cmap=itkwidgets.cm.BrBG, annotations=False, vmax=800, ui_collapsed=True)
# +
# Segment the cells
smoothed = itk.prox_tv_image_filter(image, weights=50, maximum_number_of_iterations=3)
LabelImageType = itk.Image[itk.UC, 3]
threshold_filter = itk.MomentsThresholdImageFilter.New(smoothed)
threshold_filter.Update()
threshold = threshold_filter.GetThreshold()
# -
interpolator = itk.BSplineInterpolateImageFunction.New(smoothed, spline_order=3)
mesh = itk.cuberille_image_to_mesh_filter(smoothed,
interpolator=interpolator,
iso_surface_value=threshold,
project_vertices_to_iso_surface=True,
project_vertex_surface_distance_threshold=0.5)
view(geometries=mesh)
| examples/VisualizeImageLabels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Quiz #0203
import pandas as pd
import numpy as np
import os
# #### Read in the data.
# +
# Go to the directory where the data file is located.
# os.chdir(r'~~') # Please, replace the path with your own.
# -
df = pd.read_csv('data_sales.csv', header='infer')
df.shape
df.head(5)
# #### Answer the following questions.
# 1). Append a new variable $Amount = UnitPric \times Units$.
# 2). Average unit price for each region. Use the groupby() method.
# 3). Average unit price for each region. Use the pivot_table() method.
# 4). Average unit price and units for each region in one code sentence. Use the groupby() method.
# 5). Average unit price and units for each region in one code sentence. Use the pivot_table() method.
# 6). Total units for each region and item type in one code sentence. Use the pivot_table() method. Fill the missing values with 0.
# 7). Total sales amount for each region and item type in one code sentence. Use the pivot_table() method. Fill the missing values with 0.
| SIC_AI_Quizzes/SIC_AI_Chapter_03_Quiz/problem_0203.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ppiont/tensor-flow-state/blob/master/custom_training.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="vSuzkeZcM2UE" colab_type="code" outputId="9338790f-3408-4708-ae81-f1b539a83130" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount("/gdrive", force_remount = True)
# + id="lBC4ZppZNC3c" colab_type="code" outputId="4a91494f-48f9-499f-fdc9-8ccf7fa5da0b" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd "/gdrive/My Drive/tensor-flow-state/tensor-flow-state"
# + id="NKD3KY8hNEqo" colab_type="code" colab={}
import pandas as pd
df = pd.read_csv("data/final_data.csv", index_col = 0, parse_dates = True)
# + id="YyqsebC1PLdm" colab_type="code" colab={}
df = df[df.index > "2016-06-02"]
df['density'] = (df.flow * 60) / df.speed
# + id="kzVAi95GfcBW" colab_type="code" colab={}
cols = ["speed", "flow", "density", "speed_limit", "holiday", "weekend"]
continuous_cols = ["speed", "flow", "density"]
discrete_cols = ["speed_limit", "holiday", "weekend"]
df = df[cols]
# + [markdown] id="Uh4evfBugwJa" colab_type="text"
# ### First resample to elmininate some noise
# + id="FVkBrcJDhVxA" colab_type="code" colab={}
import numpy as np
def resample_df(df, freq = "15T"):
r_df = df.copy()
r_df = r_df.resample(freq).agg({
"speed": np.mean, "flow": np.sum, "density": np.mean, "speed_limit": np.median, "holiday": np.median, "weekend": np.median})
return r_df
# + id="NJyIniCBhWIc" colab_type="code" colab={}
r_df = resample_df(df, freq = "15T")
# + [markdown] id="_M66yh3ektP7" colab_type="text"
# ### Train, val, test split
# + id="rSeGBQtNq2GC" colab_type="code" colab={}
def train_split(split_df):
train = split_df[split_df.index.year < 2019].copy()
val = split_df[len(train): -len(split_df[split_df.index > "2019-06"]) - 1].copy()
test = r_df[len(train) + len(val):].copy()
return train, val, test
train, val, test = train_split(r_df)
# + [markdown] id="KFm8wSGx5dm-" colab_type="text"
# ### MinMax Scaling
# + id="fRSHkyO85jw3" colab_type="code" colab={}
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
scaler.fit(train)
X_train = scaler.transform(train)
y_train = X_train[:, 0]
X_val = scaler.transform(val)
y_val = X_val[:, 0]
X_test = scaler.transform(test)
y_test = X_test[:, 0]
# + id="KVzDdHiTRIXA" colab_type="code" colab={}
# # Shifty! 30 min prediction
# from scipy.ndimage.interpolation import shift
# y_train = shift(y_train, -1, mode = "nearest")
# y_val = shift(y_val, -1, mode = "nearest")
# y_test = shift(y_test, -1, mode = "nearest")
# + id="ODFfW7yFcwlM" colab_type="code" outputId="9ce57d20-faa1-49a6-dc5f-0745d11c16f8" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorflow_version 2.x
from tensorflow.keras.preprocessing.sequence import TimeseriesGenerator
lookback = int(24 * (60 / 15))
batch_size = 512
train_gen = TimeseriesGenerator(data = X_train, targets = y_train, length = lookback, batch_size = batch_size)
val_gen = TimeseriesGenerator(data = X_val, targets = y_val, length = lookback, batch_size = batch_size)
test_gen = TimeseriesGenerator(data = X_test, targets = y_test, length = lookback, batch_size = batch_size)
# + id="RqJorVCYE9vJ" colab_type="code" colab={}
# ML
import tensorflow as tf
import pdb
import matplotlib.pyplot as plt
import datetime, os
tf.keras.backend.set_floatx('float64')
# + id="wPmUXqP0N3rl" colab_type="code" colab={}
############## Define Neural Network Class ##############
class neural_net(tf.keras.Model):
def __init__(self, lookback = 24 * 4, dropout = 0.5, r_dropout = 0.5):
super(neural_net, self).__init__()
# Define lookback
self.lookback = lookback
# Define dropout
self.dropout = dropout
# Define r_dropout
self.r_dropout = r_dropout
# Define discrete feature layers
self.discrete_flatten = tf.keras.layers.Flatten()
self.discrete_prelu1 = tf.keras.layers.PReLU()
self.discrete_bnorm1 = tf.keras.layers.BatchNormalization()
self.discrete_dense1 = tf.keras.layers.Dense(32)
self.discrete_prelu2 = tf.keras.layers.PReLU()
# Define time feature layers
# Time 1
self.time1_conv1 = tf.keras.layers.Conv1D(filters = 64, kernel_size = (1))
self.time1_prelu1 = tf.keras.layers.PReLU()
self.time1_lstm1 = tf.keras.layers.LSTM(units = 32, dropout = self.dropout, recurrent_dropout = self.r_dropout, input_shape = (self.lookback, 64))
self.time1_prelu2 = tf.keras.layers.PReLU()
# Time 2
self.time2_conv1 = tf.keras.layers.Conv1D(filters = 64, kernel_size = (1))
self.time2_prelu1 = tf.keras.layers.PReLU()
self.time2_lstm1 = tf.keras.layers.LSTM(units = 32, dropout = self.dropout, recurrent_dropout = self.r_dropout, input_shape = (self.lookback, 64))
self.time2_prelu2 = tf.keras.layers.PReLU()
# Define Merged layers
self.bnorm1 = tf.keras.layers.BatchNormalization()
self.dense1 = tf.keras.layers.Dense(16)
self.prelu1 = tf.keras.layers.PReLU()
self.bnorm2 = tf.keras.layers.BatchNormalization()
self.dense2 = tf.keras.layers.Dense(1, activation = 'linear')
# Define the forward propagation
def call(self, inputs):
# Split time and discrete inputs
x_time = inputs[:, :, :-3]
x_discrete = inputs[:, :, -3:]
# Run discrete layers
x_discrete = self.discrete_flatten(x_discrete)
x_discrete = self.discrete_prelu1(x_discrete)
x_discrete = self.discrete_bnorm1(x_discrete)
x_discrete = self.discrete_dense1(x_discrete)
x_discrete = self.discrete_prelu2(x_discrete)
# Run time layers
x_time1 = self.time1_conv1(x_time)
x_time1 = self.time1_prelu1(x_time1)
x_time1 = self.time1_lstm1(x_time1)
x_time1 = self.time1_prelu2(x_time1)
x_time2 = self.time2_conv1(x_time)
x_time2 = self.time2_prelu1(x_time2)
x_time2 = self.time2_lstm1(x_time2)
x_time2 = self.time2_prelu2(x_time2)
# Concat layers
x = tf.concat([x_time1, x_time2, x_discrete], axis = 1)
x = self.bnorm1(x)
x = self.dense1(x)
x = self.prelu1(x)
x = self.bnorm2(x)
x = self.dense2(x)
# Return output
return x
# Create an instance of neural network model
model = neural_net(lookback = lookback, dropout = 0.5, r_dropout = 0.5)
# Define optimizer
optimizer = tf.keras.optimizers.Adam(learning_rate = 1e-3)
# Define loss function
loss_object = tf.keras.losses.MeanSquaredError()
# # Compile model
# model.compile(optimizer = optimizer, loss = loss_fn)
# # Define callbacks
# early_stopping = tf.keras.callbacks.EarlyStopping(monitor = 'val_loss', patience = 6, restore_best_weights = True)
# learning_rate_reduce = tf.keras.callbacks.ReduceLROnPlateau(monitor = 'val_loss', factor = 0.1, patience = 4, verbose = 0, mode='auto', min_delta = 0.0001, cooldown = 0, min_lr = 0)
# + id="M9A4twuLTbU7" colab_type="code" colab={}
# Metrics
train_loss = tf.keras.metrics.Mean(name = 'train_loss')
val_loss = tf.keras.metrics.Mean(name = 'val_loss')
# + id="oDC_hjVBOcTf" colab_type="code" colab={}
# Training step
@tf.function
def train_step(x, y):
with tf.GradientTape() as tape:
predictions = model(x, training=True)
loss = loss_object(y, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss(loss)
# + id="TdKMVW2FV9Xc" colab_type="code" colab={}
@tf.function
def val_step(x, y):
predictions = model(x, training=False)
v_loss = loss_object(y, predictions)
val_loss(v_loss)
# + id="oo8aeCiKV9cq" colab_type="code" outputId="e3662b3a-cae8-4d9f-be8e-bcb04bd802f2" colab={"base_uri": "https://localhost:8080/", "height": 272}
EPOCHS = 15
for epoch in range(EPOCHS):
# Reset the metrics at the start of the next epoch
train_loss.reset_states()
val_loss.reset_states()
for x, y in train_gen:
train_step(x, y)
for val_x, val_y in val_gen:
val_step(val_x, val_y)
template = 'Epoch {}, Loss: {}, Val Loss: {}'
print(template.format(epoch + 1,
train_loss.result(),
val_loss.result()))
# + id="atlb4h3q4Ejv" colab_type="code" colab={}
# + id="rLahr8Pw4Eaf" colab_type="code" colab={}
# + id="u51JBteL4EP8" colab_type="code" colab={}
# + id="ZnrqLVEP4Dup" colab_type="code" colab={}
# + id="A_YjgA0ZD59Q" colab_type="code" outputId="166456f3-f909-4c1d-ad50-f7c4030d7d0b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Train model
history = model.fit(train_gen, validation_data = val_gen, epochs = 50, callbacks = [early_stopping, learning_rate_reduce]) #, tensorboard_callback])
train_loss, val_loss = history.history['loss'], history.history['val_loss']
# + id="hOvUVaS5LVzd" colab_type="code" outputId="2f2444a0-1ee4-4049-e4e5-acb4b66bf94c" colab={"base_uri": "https://localhost:8080/", "height": 904}
# Plot test and validation loss
plt.rc('font', family='serif')
plt.rc('xtick', labelsize='x-small')
plt.rc('ytick', labelsize='x-small')
plt.rc('legend', fontsize='x-small')
plt.rcParams["figure.dpi"] = 300
plt.rcParams["figure.figsize"] = (4, 3)
plt.figure()
plt.semilogy(train_loss, 'k-', lw = 0.5)
plt.semilogy(val_loss, 'k--', lw = 0.5)
plt.xlabel("Epochs", fontsize = 'x-small')
plt.ylabel("Loss", fontsize = 'x-small')
plt.legend(['Train loss', 'Val loss'])
plt.grid(True)
plt.title('Train Loss', fontsize = 'x-small')
plt.tight_layout()
plt.show()
# plt.savefig("plots/train_loss_3.pdf", format = "pdf")
# + id="gg96cQGwN3vl" colab_type="code" colab={}
val_pred = model.predict(val_gen)
# + id="ro9ZbLYMN3zV" colab_type="code" colab={}
# predictions = pd.DataFrame(data = np.exp(val_log.iloc[lookback:, 0].values), index = val_log[lookback:].index, columns = ['True speed'])
# predictions["Predicted speed"] = np.exp(val_pred)
# predictions["Speed limit"] = np.where(val.iloc[lookback:, val.columns.get_loc('speed_limit')] > 0.5, 130, 100)
# + id="YnskeH-k2V-c" colab_type="code" colab={}
predictions = pd.DataFrame(data = r_df.iloc[len(train) + lookback : -len(r_df[r_df.index > "2019-06"]) - 1, 0].values, index = val[lookback:].index, columns = ['True speed'])
# predictions = pd.DataFrame(data = val.iloc[lookback:, 0].values, index = val[lookback:].index, columns = ['True speed'])
fudge = val.copy()
fudge.iloc[lookback:, 0] = val_pred
predictions["Predicted speed"] = scaler.inverse_transform(fudge)[lookback:, 0]
predictions["Speed limit"] = np.where(val.iloc[lookback:, val.columns.get_loc('speed_limit')] > 0.5, 130, 100)
# + id="ndpr0s0WN33G" colab_type="code" outputId="f1699328-42bc-458d-80c8-f4b1c2e2f09d" colab={"base_uri": "https://localhost:8080/", "height": 887}
import matplotlib as mpl
start = 1104
end = 1296
y1 = predictions["Speed limit"]
y2 = predictions["True speed"]
y3 = predictions["Predicted speed"]
fig, ax = plt.subplots(figsize = (6, 3), sharey = True)
plt.rc('font', family='serif')
plt.rc('xtick', labelsize='x-small')
plt.rc('ytick', labelsize='x-small')
plt.rc('legend', fontsize='x-small')
y1[start:end].plot(ax = ax, style = 'k:', lw = 0.4, label = "Speed limit")
y2[start:end].plot(ax = ax, style = 'k--', lw = 0.4, label = "True speed")
y3[start:end].plot(ax = ax, style = 'k-', lw = 0.4, label = "Predicted speed")
_, labels = ax.get_legend_handles_labels()
plt.legend(labels)
plt.title("Speed prediction on validation set", fontsize = 'x-small')
ax.set_xlabel('Time (15m)', fontsize = 'x-small')
ax.set_ylabel('Speed (kph)', fontsize = 'x-small')
plt.tight_layout()
# plt.savefig("plots/prediction_example_3.pdf", format = "pdf")
# + id="xGw0tlmWN3-2" colab_type="code" outputId="f847a775-55ad-4673-a0ff-b941152700e4" colab={"base_uri": "https://localhost:8080/", "height": 51}
speed = predictions['True speed'].values
print(f"Naive MSE: {np.mean((speed[1:] - speed[:-1])**2)}")
print(f"Model MSE: {np.mean((predictions['True speed'] - predictions['Predicted speed'])**2)}")
# + id="C7-i4cN5pNl9" colab_type="code" colab={}
#Evaluate on test set
test_loss = model.evaluate(test_gen)
print('Test Loss: ' + str(test_loss))
# + id="5j3YoP6JW2tU" colab_type="code" colab={}
\hline
Model run & Val $R^2$ & Test $R^2$\\
\hline
\#1 & $22.04$ & $14.81$ \\
\hline
\#2 & $20.03$ & $14.90$ \\
\hline
\#3 & $20.97$ & $14.63$ \\
\hline
Average & $21.01$ & $14.78$ \\
\hline
Naive & $24.45$ & $22.22$ \\
\hline
\end{tabular}
\caption{Validation and test results.}
\label{tab:R2}
# + id="OlbN7BhqXIlM" colab_type="code" outputId="4a6b5e4d-774d-4b8e-d80b-6e8c298a5c67" colab={"base_uri": "https://localhost:8080/", "height": 54}
import numpy as np
nums = [24.45, 22.22]
for num in nums:
print(np.sqrt(num))
# + id="FcyCCqzmXk_s" colab_type="code" colab={}
$24.45$ & $22.22$
| custom_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn
import unicodedata
import requests
import urllib.request
import time
from bs4 import BeautifulSoup
url = 'https://www.bumeran.com.pe/empleos-busqueda-practicante-comercial-pagina-{}.html'.format("1")
response = requests.get(url)
response
soup = BeautifulSoup(response.text, 'html.parser')
lista = soup.findAll('a')
lista2 = []
for i in lista:
if 'indiceAviso' in str(i):
lista2.append(i['href'])
lista2
inicio = 'https://www.bumeran.com.pe'
data = []
urlpuesto = inicio + lista2[0]
responsepuesto = requests.get(urlpuesto)
souppuesto = BeautifulSoup(responsepuesto.text, 'html.parser')
titulo1 = souppuesto.find_all("title")[0].text
descripcion1 = souppuesto.find_all("div", {"class": "aviso_description"})[0].text
titulo1
diclimpiar = {"\xa0":" ","\n":" ","\t":" "}
for k,y in diclimpiar.items():
descripcion1 = descripcion1.replace(k,y)
descripcion1
for i in lista2:
urlpuesto = inicio + i
responsepuesto = requests.get(urlpuesto)
souppuesto = BeautifulSoup(responsepuesto.text, 'html.parser')
data.append(["",""])
data[-1][0] = souppuesto.find_all("title")[0].text
data[-1][1] = souppuesto.find_all("div", {"class": "aviso_description"})[0].text
for k,y in diclimpiar.items():
data[-1][1] = data[-1][1].replace(k,y)
data
df = pd.DataFrame(data)
df
df.to_csv('data_bumeran.csv',encoding='utf-8')
| Hackathon1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Your read_csv() call to import the CSV data didn't generate an error, but the output is not entirely what we wanted. The row labels were imported as another column without a name.
#
# Remember index_col, an argument of read_csv(), that you can use to specify which column in the CSV file should be used as a row label? Well, that's exactly what you need here!
#
# Python code that solves the previous exercise is already included; can you make the appropriate changes to fix the data import?
# Import pandas as pd
import pandas as pd
# ### Specify the index_col argument inside pd.read_csv(): set it to 0, so that the first column is used as row labels.
# Fix import by including index_col
cars = pd.read_csv('cars.csv',index_col=0)
# Print out cars
print(cars)
cars.set_index('drives_right')
# it replace the previous index
print(cars.columns)
cars = pd.read_csv('cars.csv')
cars
cars.columns=['short_from','cars_per_cap', 'country', 'drives_right']
cars
cars.rename(columns={'country':'COUNTRY'},inplace=True)
cars.columns
cars.rename(columns=lambda x: x[0:3], inplace=True)
cars
cars.rename(index={0:'zero',1:'one'}, inplace=True)
| Intermediate Python for Data Science/Dictionaries - Pandas/.ipynb_checkpoints/07-CSV to DataFrame (2)-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import astropy as ap
import galpy as gp #affiliated with astropy, might have more things that we want
import numpy as np
import matplotlib.pyplot as plt
from astropy.modeling import models, fitting
ap.version.version
g_init = models.Gaussian1D(amplitude=1,mean=1,stddev=1)
print(g_init)
# +
x = np.linspace(-10,10,100)
y = np.exp(-0.5*(x-1)**2)
fit_g = fitting.LevMarLSQFitter()
g = fit_g(g_init,x,y)
# -
plt.plot(x,y,'ko')
plt.plot(x,g_init(x),'b-')
plt.plot(x,g(x))
| binder/testing/Timing_and_Old_Tests/astropy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import mxnet as mx
from mxnet import nd, autograd, gluon
# 
# # Raw Data - FER2013
# * https://www.kaggle.com/c/challenges-in-representation-learning-facial-expression-recognition-challenge/data
# * 48 X 48 gray scale images
# * 28,709 training samples
# * 3,589 validation data
# * 3,589 test data
# * 7 emotion labels (0=Angry, 1=Disgust, 2=Fear, 3=Happy, 4=Sad, 5=Surprise, 6=Neutral)
# * ~60-65% human accuracy in labelling
#
# 
#
# # Data Preparation is Critical
# * State of the Art accuracy using a Convolutional Neural Network (CNN) model directly on this raw data is **63%** [1].
# * State of the Art accuracy using a slightly modified Convolutional Neural Network (CNN) model after processing the data is **83%** [2].
# # Processed Data - FER+
#
# Follow the instructions listed in the README file - https://github.com/sandeep-krishnamurthy/facial-emotion-recognition-gluon#step-1---data-preparation
#
# * FER+ has new corrected labels
# * FER+ has 8 emotions - (0: 'neutral', 1: 'happiness', 2: 'surprise', 3: 'sadness', 4: 'anger', 5: 'disgust', 6: 'fear',7: 'contempt')
# * Image augmentations:
# * Crop faces in the images – bounding box in the FER+ dataset
# * Scale image size from 48 X 48 -> 64 X 64
# * Shift image
# * Flip image
# * Rotate (angle) image
# * Normalize the pixels in the image
# | Emotion | Train | Val | Test |
# |---|---|---|---|
# | neutral |8733 | 1180 | 1083 |
# | happiness | 7284 | 862 | 892 |
# | surprise | 3136 | 411 | 394 |
# | sadness | 3022 | 348 | 382 |
# | anger | 2098 | 289 | 269 |
# | disgust | 116 | 25 | 16 |
# | fear | 536 | 60 | 86 |
# | comtempt | 120 | 16 | 15 |
# 8 Emotions we want to recognize
emotion_table = {0: 'neutral',
1: 'happiness',
2: 'surprise',
3: 'sadness',
4: 'anger',
5: 'disgust',
6: 'fear',
7: 'contempt'}
processed_train_images = np.load('../data/fer_train_processed_images.npy')
processed_train_labels = np.load('../data/fer_train_processed_labels.npy')
print(processed_train_images.shape, processed_train_labels.shape)
from matplotlib import pyplot as plt
plt.imshow(processed_train_images[987].reshape(64,64), cmap='gray')
processed_test_images = np.load('../data/fer_test_processed_images.npy')
processed_test_labels = np.load('../data/fer_test_processed_labels.npy')
processed_val_images = np.load('../data/fer_val_processed_images.npy')
processed_val_labels = np.load('../data/fer_val_processed_labels.npy')
print(processed_test_images.shape, processed_test_labels.shape)
print(processed_val_images.shape, processed_val_labels.shape)
# +
# Set this to ctx = mx.cpu() if running on CPU.
# However, please note, it takes approx. 1.1 min/epoch on 1 GPU => Can take longer time on cPU
ctx = mx.gpu()
# -
# 
# # Step 1 – Construct the Neural Network
# ### 13 layer VGGNet presented in the paper [2]
# 
# +
# We use HybridSequential network type to able to save the trained model as symbols and params.
# More Info - https://mxnet.incubator.apache.org/tutorials/gluon/save_load_params.html
net = gluon.nn.HybridSequential()
# Construct 13 layer VGGNet suggested in the paper
with net.name_scope():
net.add(gluon.nn.Conv2D(channels=64, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.Conv2D(channels=64, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=2, strides=2))
net.add(gluon.nn.Dropout(0.25))
net.add(gluon.nn.Conv2D(channels=128, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.Conv2D(channels=128, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=2, strides=2))
net.add(gluon.nn.Dropout(0.25))
net.add(gluon.nn.Conv2D(channels=256, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.Conv2D(channels=256, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.Conv2D(channels=256, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=2, strides=2))
net.add(gluon.nn.Dropout(0.25))
net.add(gluon.nn.Conv2D(channels=256, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.Conv2D(channels=256, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.Conv2D(channels=256, kernel_size=3, padding=(1,1), activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=2, strides=2))
net.add(gluon.nn.Dropout(0.25))
net.add(gluon.nn.Flatten())
net.add(gluon.nn.Dense(1024, activation='relu'))
net.add(gluon.nn.Dropout(0.5))
net.add(gluon.nn.Dense(1024, activation='relu'))
net.add(gluon.nn.Dropout(0.5))
net.add(gluon.nn.Dense(8))
# -
# We Hybridize the HybridSequential network to able to save the trained model as symbols and params.
# More Info - https://mxnet.incubator.apache.org/tutorials/gluon/save_load_params.html
net.hybridize()
# # Step 2 – Initialize the parameters in Neural Network
net.collect_params().initialize(mx.init.Xavier(), ctx=ctx)
nd.waitall()
# Use MXBOARD here to visualize network
x = mx.sym.var('data')
sym = net(x)
mx.viz.plot_network(sym)
# # Step 3 – Prepare the Trainer with optimizer
softmax_cross_entropy = gluon.loss.SoftmaxCrossEntropyLoss()
batch_size = 32
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': 0.0025, 'momentum': 0.9})
# # Step 4 – Prepare the model evaluation strategy
def evaluate_accuracy(data_iterator, net):
acc = mx.metric.Accuracy()
for i, (data, label) in enumerate(data_iterator):
data = data.as_in_context(ctx)
label = label.as_in_context(ctx)
output = net(data)
predictions = nd.argmax(output, axis=1)
acc.update(preds=predictions, labels=label)
return acc.get()[1]
# # Step 5 – Prepare data loaders
# +
train_labels = np.argmax(processed_train_labels, axis=1)
val_labels = np.argmax(processed_val_labels, axis=1)
train_data = gluon.data.DataLoader(gluon.data.ArrayDataset(processed_train_images, train_labels), batch_size = batch_size, shuffle=True)
val_data = gluon.data.DataLoader(gluon.data.ArrayDataset(processed_val_images, val_labels), batch_size = batch_size)
# -
#
# # Step 6 – Train the Neural Network
# +
epochs = 25
train_accuracies = []
losses = []
val_accuracies = []
for e in range(epochs):
batch = 0
for i, (data, label) in enumerate(train_data):
data = data.as_in_context(ctx)
label = label.as_in_context(ctx)
with autograd.record():
output = net(data)
loss = softmax_cross_entropy(output, label)
loss.backward()
trainer.step(data.shape[0])
curr_loss = nd.mean(loss).asscalar()
batch +=1
val_accuracy = evaluate_accuracy(val_data, net)
train_accuracy = evaluate_accuracy(train_data, net)
losses.append(curr_loss)
train_accuracies.append(train_accuracy)
val_accuracies.append(val_accuracy)
print("Epoch %s. Loss: %s, Train_acc %s, Val_acc %s" % (e, curr_loss, train_accuracy, val_accuracy))
# -
# # Step 7 - Evaluate on Test Data
# +
# Test accuracy
acc = mx.metric.Accuracy()
test_labels = np.argmax(processed_test_labels, axis=1)
data_iterator = gluon.data.DataLoader(gluon.data.ArrayDataset(processed_test_images, test_labels), batch_size = 32)
for i, (data, label) in enumerate(data_iterator):
data = data.as_in_context(ctx)
label = label.as_in_context(ctx)
output = net(data)
predictions = nd.argmax(output, axis=1)
acc.update(preds=predictions, labels=label)
print("Test Accuracy - ", acc.get()[1])
# +
# for plotting purposes
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
epochs = range(len(train_accuracies))
f = plt.figure(figsize=(12,6))
fg1 = f.add_subplot(121)
fg2 = f.add_subplot(122)
fg1.set_xlabel('epoch',fontsize=14)
fg1.set_title('Loss over Training')
fg1.grid(True, which="both")
fg1.plot(epochs, losses)
fg2.set_title('Comparing accuracy')
fg2.set_xlabel('epoch', fontsize=14)
fg2.grid(True, which="both")
p1, = fg2.plot(epochs, train_accuracies)
p2, = fg2.plot(epochs, val_accuracies)
fg2.legend([p1, p2], ['training accuracy', 'validation accuracy'],fontsize=14)
# -
# Example Inference
idx = 98
plt.imshow(processed_test_images[idx].reshape(64,64), cmap='gray')
print("Actual Emotion - ", emotion_table[test_labels[idx]])
# Perform Inference
output = net(mx.nd.array(processed_test_images[idx].reshape(1,1,64,64)).as_in_context(ctx))
print("Predicted Emotion - ", emotion_table[nd.argmax(output, axis=1).asnumpy()[0]])
# 
# # Step 8 - Export the model for Production
# Export the model for production deployment.
# There will be 2 files exported:
# 1) gluon_ferplus-symbol.json => Contains the network definition
# 2) gluon_ferplus-0000.params => Contains the weights in the network
net.export('gluon_ferplus')
# # (Optional) Get pre-trained model
#
# If you prefer to directly use a pre-trained model, you can download the same from:
# ```
# wget https://s3.amazonaws.com/mxnet-demo-models/models/fer/gluon_ferplus-0000.params
# wget https://s3.amazonaws.com/mxnet-demo-models/models/fer/gluon_ferplus-symbol.json
# ```
# # References
# 1. <NAME>, <NAME>, <NAME>, <NAME>,
# <NAME>, <NAME>, <NAME>, <NAME>,
# <NAME>, <NAME>, et al. Challenges in
# representation learning: A report on three machine
# learning contests. In Neural information processing,
# pages 117–124. Springer, 2013
#
# 2. Training Deep Networks for Facial Expression Recognition with Crowd-Sourced Label Distribution Emad Barsoum et. al. https://arxiv.org/abs/1608.01041
#
| notebooks/Gluon_FERPlus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Importing packages
import csv
import nytimesarticle
from nytimesarticle import articleAPI
#API Key
api = articleAPI('<KEY>')
# This function takes in a response to the NYT api and parses the articles into a list
def parse(articles):
news = []
if 'response' in articles :
for i in articles["response"]["docs"]:
dat = {}
dat['headline'] = i['headline']['main'].encode("utf8")
dat['source'] = i['source']
dat['type'] = i['type_of_material']
dat['url'] = i['web_url']
dat['word_count'] = i['word_count']
news.append(dat)
return(news)
#retrieve article urls and store it in a csv file
all_articles = []
for i in range(0,100):
articles = api.search(q = "basketball", begin_date = 20180401, end_date = 20180511, page = i)
articles = parse(articles)
all_articles = all_articles + articles
keys = all_articles[0].keys()
outputfile = open('Testing_Sports.csv', 'w',newline='')
dict_writer = csv.DictWriter(outputfile,keys)
dict_writer.writerows(all_articles)
outputfile.close()
# +
#Importing packages
import csv
import urllib.request
from bs4 import BeautifulSoup
from bs4.element import Comment
#Parsing HTML documents using BeautifulSoup
def extractbody(body):
soup = BeautifulSoup(body, 'html.parser')
texts = soup.findAll(text=True)
visible_texts = filter(parse, texts)
return u" ".join(t.strip() for t in visible_texts)
#Parsing through different tags
def parse(element):
if element.parent.name in ['style', 'script', 'head', 'title', 'meta', '[document]']:
return False
if isinstance(element, Comment):
return False
return True
#Storing the text from news articles into different files
iterator=1
csvDataFile = open('Testing_Politics.csv', 'r')
csvReader = csv.reader(csvDataFile)
for row in csvReader:
html = urllib.request.urlopen(row[3]).read()
outputfile = open("Testing_Politics_%s.txt" %iterator,"w",encoding="utf-8")
outputfile.write(extractbody(html))
outputfile.close()
iterator = iterator+1
# -
| 2_DataCollection/News_DataCollection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
import pandas as pd
parquet_file = "example.parquet"
df = pd.read_parquet(parquet_file, engine='pyarrow')
# show the first lines of a table
df.head()
# show selected column(s) of the table
df[["one", "two"]].head()
# filter rows by value
df[df["two"]=="foo"].head()
# filter rows by value
df[df["three"]==True].head()
# filter rows by value
df[df["one"]>0].head()
| coding/python/day02/03_filter_parquet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import skfuzzy as fuzz
from skfuzzy import control as ctrl
# +
front_dist = ctrl.Antecedent(np.arange(0, 256, 1), 'front_dist')
right_dist = ctrl.Antecedent(np.arange(0, 256, 1), 'right_dist')
left_dist = ctrl.Antecedent(np.arange(0, 256, 1), 'left_dist')
right_motor = ctrl.Consequent(np.arange(-101, 101, 1), 'right_motor')
left_motor = ctrl.Consequent(np.arange(-101, 101, 1), 'left_motor')
# +
front_dist['near']= fuzz.trapmf(front_dist.universe, [0,0,10,30])
front_dist['medium']= fuzz.trimf(front_dist.universe, [25, 50, 80])
front_dist['far']= fuzz.trapmf(front_dist.universe, [75,100,255,255])
right_dist['near']= fuzz.trapmf(right_dist.universe, [0,0,10,30])
right_dist['medium']= fuzz.trimf(right_dist.universe, [25, 50, 80])
right_dist['far']= fuzz.trapmf(right_dist.universe, [75,100,255,255])
left_dist['near']= fuzz.trapmf(left_dist.universe, [0,0,10,30])
left_dist['medium']= fuzz.trimf(left_dist.universe, [25, 50, 80])
left_dist['far']= fuzz.trapmf(left_dist.universe, [75,100,255,255])
# -
front_dist['near'].view()
# +
right_motor['stop'] = fuzz.trapmf(right_motor.universe, [-10,-5, 5, 10])
right_motor['low'] = fuzz.trapmf(right_motor.universe, [5,20, 30, 60])
right_motor['high'] = fuzz.trapmf(right_motor.universe, [50,60, 100, 100])
right_motor['min_low'] = fuzz.trapmf(right_motor.universe, [-60,-30, -20, -5])
right_motor['min_high'] = fuzz.trapmf(right_motor.universe, [-100,-100, -60, -50])
left_motor['stop'] = fuzz.trapmf(left_motor.universe, [-10,-5, 5, 10])
left_motor['low'] = fuzz.trapmf(left_motor.universe, [5,20, 30, 60])
left_motor['high'] = fuzz.trapmf(left_motor.universe, [50,60, 100, 100])
left_motor['min_low'] = fuzz.trapmf(left_motor.universe, [-60,-30, -20, -5])
left_motor['min_high'] = fuzz.trapmf(left_motor.universe, [-100,-100, -60, -50])
# -
left_motor['low'].view()
# +
# Implement the rules:
r00 = ctrl.Rule(front_dist['near'] | right_dist['near']| left_dist['near'] , left_motor['high'])
r01 = ctrl.Rule(front_dist['near'] | right_dist['near']| left_dist['near'] , right_motor['min_high'] )
r10 = ctrl.Rule(front_dist['far'] , left_motor['high'])
r11 = ctrl.Rule(front_dist['far'] , right_motor['high'])
r20 = ctrl.Rule(front_dist['far'] | right_dist['medium'], left_motor['low'])
r21 = ctrl.Rule(front_dist['far'] | right_dist['medium'], right_motor['high'])
r30 = ctrl.Rule(front_dist['far'] | left_dist['medium'], right_motor['low'])
r31 = ctrl.Rule(front_dist['far'] | left_dist['medium'], left_motor['high'])
r40 = ctrl.Rule(front_dist['near'] | right_dist['medium'], left_motor['low'])
r41 = ctrl.Rule(front_dist['near'] | right_dist['medium'], right_motor['min_low'])
r42 = ctrl.Rule(front_dist['near'] | right_dist['far'], left_motor['low'])
r43 = ctrl.Rule(front_dist['near'] | right_dist['far'], right_motor['min_low'])
r50 = ctrl.Rule(front_dist['near'] | left_dist['medium'], right_motor['low'])
r51 = ctrl.Rule(front_dist['near'] | left_dist['medium'], left_motor['min_low'])
r52 = ctrl.Rule(front_dist['near'] | left_dist['far'], right_motor['low'])
r53 = ctrl.Rule(front_dist['near'] | left_dist['far'], left_motor['min_low'])
r60 = ctrl.Rule(front_dist['medium'] | right_dist['medium'], left_motor['low'])
r61 = ctrl.Rule(front_dist['medium'] | right_dist['medium'], right_motor['low'])
r70 = ctrl.Rule(front_dist['medium'] | left_dist['medium'], right_motor['low'])
r71 = ctrl.Rule(front_dist['medium'] | left_dist['medium'], left_motor['low'])
# +
RAO = ctrl.ControlSystem([
r00, r01, r10, r11, r10, r11, r20, r21, r30, r31, r40, r41 , r42, r43, r50, r51 , r52, r53, r60, r61, r70, r71
])
rao = ctrl.ControlSystemSimulation(RAO)
rao.input['front_dist'] = 25
rao.input['right_dist'] = 25
rao.input['left_dist'] = 150
# Crunch the numbers
rao.compute()
print (rao.output)
# print (rao.output['left_motor'])
# print (rao.output['right_motor'])
# rao.view(sim=rao)
# +
rao = ctrl.ControlSystemSimulation(RAO)
rao.input['front_dist'] = 20
# rao.input['right_dist'] = 100
rao.input['left_dist'] = 10
# Crunch the numbers
rao.compute()
print (rao.output)
# print (rao.output['left_motor'])
# print (rao.output['right_motor'])
# rao.view(sim=rao)
# -
| fuzzy- Robot Avoiding Obstacles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AryanNayak/Muse/blob/main/generative_cb.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="QXpDe8v5H7Nr" colab={"base_uri": "https://localhost:8080/"} outputId="f630cd71-24df-4355-fa58-be95937c7982"
import re
import random
# !wget -P data/full_dataset/ https://storage.googleapis.com/gresearch/goemotions/data/full_dataset/goemotions_1.csv
# !wget -P data/full_dataset/ https://storage.googleapis.com/gresearch/goemotions/data/full_dataset/goemotions_2.csv
# !wget -P data/full_dataset/ https://storage.googleapis.com/gresearch/goemotions/data/full_dataset/goemotions_3.csv
data_path = "data/full_dataset/goemotions_1.csv"
data_path2 = "data/full_dataset/goemotions_2.csv"
data_path3 = "data/full_dataset/goemotions_3.csv"
# Defining lines as a list of each line
with open(data_path, 'r', encoding='utf-8') as f:
lines = f.read().split('\n')
with open(data_path2, 'r', encoding='utf-8') as f:
lines2 = f.read().split('\n')
with open(data_path3, 'r', encoding='utf-8') as f:
lines2 = f.read().split('\n')
lines = [re.sub(r"\[\w+\]",'hi',line) for line in lines]
lines = [" ".join(re.findall(r"\w+",line)) for line in lines]
lines2 = [re.sub(r"\[\w+\]",'',line) for line in lines2]
lines2 = [" ".join(re.findall(r"\w+",line)) for line in lines2]
lines3 = [re.sub(r"\[\w+\]",'',line) for line in lines2]
lines3 = [" ".join(re.findall(r"\w+",line)) for line in lines2]
# grouping lines by response pair
pairs = list(zip(lines,lines2,lines3))
#random.shuffle(pairs)
# + id="s0URAjuRKvWq"
import numpy as np
input_docs = []
target_docs = []
input_tokens = set()
target_tokens = set()
for line in pairs[:400]:
input_doc, target_doc = line[0], line[1]
# Appending each input sentence to input_docs
input_docs.append(input_doc)
# Splitting words from punctuation
target_doc = " ".join(re.findall(r"[\w']+|[^\s\w]", target_doc))
# Redefine target_doc below and append it to target_docs
target_doc = '<START> ' + target_doc + ' <END>'
target_docs.append(target_doc)
# Now we split up each sentence into words and add each unique word to our vocabulary set
for token in re.findall(r"[\w']+|[^\s\w]", input_doc):
if token not in input_tokens:
input_tokens.add(token)
for token in target_doc.split():
if token not in target_tokens:
target_tokens.add(token)
input_tokens = sorted(list(input_tokens))
target_tokens = sorted(list(target_tokens))
num_encoder_tokens = len(input_tokens)
num_decoder_tokens = len(target_tokens)
# + id="7ZQqs9tPMmEG"
input_features_dict = dict(
[(token, i) for i, token in enumerate(input_tokens)])
target_features_dict = dict(
[(token, i) for i, token in enumerate(target_tokens)])
reverse_input_features_dict = dict(
(i, token) for token, i in input_features_dict.items())
reverse_target_features_dict = dict(
(i, token) for token, i in target_features_dict.items())
# + id="glkkUFqoNskS"
#Maximum length of sentences in input and target documents
max_encoder_seq_length = max([len(re.findall(r"[\w']+|[^\s\w]", input_doc))
for input_doc in input_docs])
max_decoder_seq_length = max([len(re.findall(r"[\w']+|[^\s\w]", target_doc))
for target_doc in target_docs])
encoder_input_data = np.zeros(
(len(input_docs), max_encoder_seq_length, num_encoder_tokens),
dtype='float32')
decoder_input_data = np.zeros(
(len(input_docs), max_decoder_seq_length, num_decoder_tokens),
dtype='float32')
decoder_target_data = np.zeros(
(len(input_docs), max_decoder_seq_length, num_decoder_tokens),
dtype='float32')
for line, (input_doc, target_doc) in enumerate(zip(input_docs, target_docs)):
for timestep, token in enumerate(re.findall(r"[\w']+|[^\s\w]", input_doc)):
#Assign 1. for the current line, timestep, & word in encoder_input_data
encoder_input_data[line, timestep, input_features_dict[token]] = 1.
for timestep, token in enumerate(target_doc.split()):
decoder_input_data[line, timestep, target_features_dict[token]] = 1.
if timestep > 0:
decoder_target_data[line, timestep - 1, target_features_dict[token]] = 1.
# + id="uZJ4SWLXO_uh"
from tensorflow import keras
from keras.layers import Input, LSTM, Dense
from keras.models import Model
#Dimensionality
dimensionality = 256
#The batch size and number of epochs
batch_size = 10
epochs = 1000
#Encoder
encoder_inputs = Input(shape=(None, num_encoder_tokens))
encoder_lstm = LSTM(dimensionality, return_state=True)
encoder_outputs, state_hidden, state_cell = encoder_lstm(encoder_inputs)
encoder_states = [state_hidden, state_cell]
#Decoder
decoder_inputs = Input(shape=(None, num_decoder_tokens))
decoder_lstm = LSTM(dimensionality, return_sequences=True, return_state=True)
decoder_outputs, decoder_state_hidden, decoder_state_cell = decoder_lstm(decoder_inputs, initial_state=encoder_states)
decoder_dense = Dense(num_decoder_tokens, activation='softmax')
decoder_outputs = decoder_dense(decoder_outputs)
# + id="BRvb9nnNPMPV" colab={"base_uri": "https://localhost:8080/"} outputId="fadf8e82-51cd-499f-dd3e-d3b3b92835af"
#Model
training_model = Model([encoder_inputs, decoder_inputs], decoder_outputs)
#Compiling
training_model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'], sample_weight_mode='temporal')
#Training
training_model.fit([encoder_input_data, decoder_input_data], decoder_target_data, batch_size = batch_size, epochs = epochs, validation_split = 0.2)
training_model.save('training_model.h5')
# + id="GWuDFsrsPnTk"
from keras.models import load_model
training_model = load_model('training_model.h5')
encoder_inputs = training_model.input[0]
encoder_outputs, state_h_enc, state_c_enc = training_model.layers[2].output
encoder_states = [state_h_enc, state_c_enc]
encoder_model = Model(encoder_inputs, encoder_states)
# + id="W7eBGr7UPrmD"
latent_dim = 256
decoder_state_input_hidden = Input(shape=(latent_dim,))
decoder_state_input_cell = Input(shape=(latent_dim,))
decoder_states_inputs = [decoder_state_input_hidden, decoder_state_input_cell]
# + id="inTEpA9qPuOK"
decoder_outputs, state_hidden, state_cell = decoder_lstm(decoder_inputs, initial_state=decoder_states_inputs)
decoder_states = [state_hidden, state_cell]
decoder_outputs = decoder_dense(decoder_outputs)
# + id="osaSchXwP57A"
decoder_model = Model([decoder_inputs] + decoder_states_inputs, [decoder_outputs] + decoder_states)
# + id="fgjCXEVkQOy1"
def decode_response(test_input):
#Getting the output states to pass into the decoder
states_value = encoder_model.predict(test_input)
#Generating empty target sequence of length 1
target_seq = np.zeros((1, 1, num_decoder_tokens))
#Setting the first token of target sequence with the start token
target_seq[0, 0, target_features_dict['<START>']] = 1.
#A variable to store our response word by word
decoded_sentence = ''
stop_condition = False;
while not stop_condition:
#Predicting output tokens with probabilities and states
output_tokens, hidden_state, cell_state = decoder_model.predict([target_seq] + states_value)
#Choosing the one with highest probability
sampled_token_index = np.argmax(output_tokens[0, -1, :])
sampled_token = reverse_target_features_dict[sampled_token_index]
decoded_sentence += " " + sampled_token
#Stop if hit max length or found the stop token
if (sampled_token == '<END>' or len(decoded_sentence) > max_decoder_seq_length):
stop_condition = True
#Update the target sequence
target_seq = np.zeros((1, 1, num_decoder_tokens))
target_seq[0, 0, sampled_token_index] = 1.
#Update states
states_value = [hidden_state, cell_state]
return decoded_sentence
# + id="uKzuMJ5CQWt1" colab={"base_uri": "https://localhost:8080/", "height": 313} outputId="227c1a96-5108-47c0-fe6c-dad02d5ee797"
class ChatBot:
negative_responses = ("no", "nope", "nah", "naw", "not a chance","sorry")
exit_commands = ("quit", "pause", "exit", "goodbye", "bye","later", "stop")
#Method to start the conversation
def start_chat(self):
user_response = input("Hi, I'm a chatbot trained on random dialogs. Would you like to chat with me?\n")
if user_response in self.negative_responses:
print("Ok, have a great day!")
return
self.chat(user_response)
#Method to handle the conversation
def chat(self, reply):
while not self.make_exit(reply):
reply = input(self.generate_response(reply)+"\n")
#Method to convert user input into a matrix
def string_to_matrix(self, user_input):
tokens = re.findall(r"[\w']+|[^\s\w]", user_input)
user_input_matrix = np.zeros(
(1, max_encoder_seq_length, num_encoder_tokens),
dtype='float32')
for timestep, token in enumerate(tokens):
if token in input_features_dict:
user_input_matrix[0, timestep, input_features_dict[token]] =1.
return user_input_matrix
#Method that will create a response using seq2seq model we built
def generate_response(self, user_input):
input_matrix = self.string_to_matrix(user_input)
chatbot_response = decode_response(input_matrix)
#Remove <START> and <END> tokens from chatbot_response
chatbot_response = chatbot_response.replace("<START>",'')
chatbot_response = chatbot_response.replace("<END>",'')
return chatbot_response
#Method to check for exit commands
def make_exit(self, reply):
for exit_command in self.exit_commands:
if exit_command in reply:
print("Ok, have a great day!")
return True
return False
chatbot = ChatBot()
chatbot.start_chat()
chatbot.chat()
# + id="3cla2IOTui8J"
| Conversational chatbot/generative_cb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from time import time
import os
import sys
sys.path.append('../')
from utils.codification_cnn import CNNLayer, NNLayer, ChromosomeCNN, FitnessCNN, FitnessCNNParallel
from utils.datamanager import DataManager
from utils.lr_finder import LRFinder
from time import time
import numpy as np
# +
fitness_cnn = FitnessCNN()
# dataset params:
data_folder = '../../../../../../datasets/MNIST_variations'
classes = []
# Fitness params
epochs = 75
batch_size = 128
verbose = 1
redu_plat = False
early_stop = 0
warm_up_epochs= 1
base_lr = 0.001
smooth = 0.1
cosine_dec = True
lr_find = True
dataset = 'MRDBI'
dm = DataManager(dataset, clases=classes, folder_var_mnist=data_folder)
data = dm.load_data()
print(data[0][0].shape)
# +
fitness_cnn.set_params(data=data, verbose=verbose, batch_size=batch_size, reduce_plateau=redu_plat,
epochs=epochs, cosine_decay=cosine_dec, early_stop=early_stop,
warm_epochs=warm_up_epochs, base_lr=base_lr, smooth_label=smooth, find_lr=lr_find)
fitness_folder = '../../delete'
fitness_file = '../../delete/fitness_example'
fitness_cnn.save(fitness_file)
fitness = FitnessCNNParallel()
# -
l1_2 = CNNLayer(86, (3,5), 'leakyreLu', 0.262, 1)
l2_2 = CNNLayer(84, (5,3), 'leakyreLu', 0.319, 1)
l3_2 = CNNLayer(243, (1,3), 'prelu', 0.322, 1)
l4_2 = NNLayer(948, 'sigmoid', 0.467)
l5_2 = NNLayer(780, 'sigmoid', 0.441)
best_mrdbi_v2 = ChromosomeCNN([l1_2, l2_2, l3_2], [l4_2, l5_2], fitness)
# +
FPS = {16:'FP16 + BN16', 32:'FP32 + BN32', 160:'FP16', 320:'FP32', 3216:'FP32 + BN16'}
fp = 16
fitness_cnn.save(fitness_file)
fitness.set_params(chrom_files_folder=fitness_folder, fitness_file=fitness_file, max_gpus=1,
fp=fp, main_line='python /home/daniel/proyectos/Tesis/project/GA/NeuroEvolution/train_gen.py')
ti = time()
print("Evaluationg Second model with FP%d, cosine_decay %s" %(fp, str(cos)) )
score = fitness.calc(best_mrdbi_v2, test=True)
print("Score: %0.3f" % score)
print("Elapsed time: %0.3f " % (time() - ti))
# +
l1_2 = CNNLayer(86, (3,5), 'leakyreLu', 0.262, 1)
l2_2 = CNNLayer(84, (5,3), 'leakyreLu', 0.319, 1)
l3_2 = CNNLayer(243, (1,3), 'prelu', 0.322, 1)
l4_2 = NNLayer(948, 'sigmoid', 0.467)
l5_2 = NNLayer(780, 'sigmoid', 0.441)
best_mrdbi_v2 = ChromosomeCNN([l1_2, l2_2, l3_2], [l4_2, l5_2], fitness_cnn)
FPS = {16:'FP16 + BN16', 32:'FP32 + BN32', 160:'FP16', 320:'FP32', 3216:'FP32 + BN16'}
fp = 16
ti = time()
print("Evaluationg Second model with FP%d" % fp )
score = fitness_cnn.calc(best_mrdbi_v2, fp=fp)
print("Score: %0.3f" % score)
print("Elapsed time: %0.3f " % (time() - ti))
# -
fitness_cnn.find_lr = False
ti = time()
print("Evaluationg Second model with FP%d" % fp )
score = fitness_cnn.calc(best_mrdbi_v2, fp=fp)
print("Score: %0.3f" % score)
print("Elapsed time: %0.3f " % (time() - ti))
| experiments/LR_test_finder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:jupyter_env] *
# language: python
# name: conda-env-jupyter_env-py
# ---
# +
from __future__ import print_function
from __future__ import absolute_import
from ku import generators as gr
from ku import generic as gen
from ku import image_utils as iu
from ku import model_helper as mh
from ku import image_augmenter as aug
from munch import Munch
import pandas as pd, numpy as np
import pytest, shutil, os
from matplotlib import pyplot as plt
# %load_ext autoreload
# %autoreload 2
# +
gen_params = Munch(batch_size = 2,
data_path = 'images',
input_shape = (224,224,5),
inputs = ['filename'],
outputs = ['score'],
shuffle = False,
fixed_batches = True)
ids = pd.read_csv(u'ids.csv', encoding='latin-1')
np.all(ids.columns == ['filename', 'score'])
np.all(ids.score == range(1,5))
# +
def preproc(im, arg1, arg2):
return np.zeros(1) + arg1 + arg2
gen_params_local = gen_params.copy()
gen_params_local.process_fn = preproc
gen_params_local.process_args = {'filename': ['filename_args','filename_args']}
gen_params_local.batch_size = 4
ids_local = ids.copy()
ids_local['filename_args'] = range(len(ids_local))
g = gr.DataGeneratorDisk(ids_local, **gen_params_local)
x = g[0]
# gen.pretty(g)
assert np.array_equal(np.squeeze(x[0][0].T), np.arange(4)*2)
# +
g = gr.DataGeneratorDisk(ids, **gen_params)
print(isinstance(g[0][1], list))
print(np.all(g[0][1][0] == np.array([[1],[2]])))
gen.get_sizes(g[0])=='([array<2,224,224,3>], [array<2,1>])'
# +
# read_fn = lambda p: iu.resize_image(iu.read_image(p), (100,100))
# g = gr.DataGeneratorDisk(ids, read_fn=read_fn, **gen_params)
# gen.get_sizes(g[0]) =='([array<2,100,100,3>], [array<2,1>])'
# +
# # reload(gen)
# x = np.array([[1,2,3]])
# print(gen.get_sizes(([x.T],1,[4,5])))
# y = np.array([[1,[1,2]]])
# print(gen.get_sizes(y))
# z = [g[0],([2],)]
# print(gen.get_sizes(z[1]))
# +
gen_params.inputs = ['filename', 'filename']
g = gr.DataGeneratorDisk(ids, **gen_params)
assert gen.get_sizes(g[0]) == '([array<2,224,224,3>, array<2,224,224,3>], [array<2,1>])'
g.inputs_df = ['score', 'score']
g.inputs = []
g.outputs = []
gen.get_sizes(g[0])
g.inputs_df = [['score'], ['score','score']]
assert gen.get_sizes(g[0]) == '([array<2,1>, array<2,2>], [])'
g.inputs_df = []
g.outputs = ['score']
assert gen.get_sizes(g[0]) == '([], [array<2,1>])'
g.outputs = ['score',['score']]
with pytest.raises(AssertionError): g[0]
g.outputs = [['score'],['score']]
assert gen.get_sizes(g[0]) == '([], [array<2,1>, array<2,1>])'
# +
with gen.H5Helper('data.h5', overwrite=True) as h:
data = np.expand_dims(np.array(ids.score), 1)
h.write_data(data, list(ids.filename))
with gen.H5Helper('data.h5', 'r') as h:
data = h.read_data(list(ids.filename))
assert all(data == np.array([[1],[2],[3],[4]]))
# +
gen_params.update(data_path='data.h5',
inputs=['filename'],
batch_size=2)
gen.pretty(gen_params)
g = gr.DataGeneratorHDF5(ids, **gen_params)
assert gen.get_sizes(g[0]) == '([array<2,1>], [array<2,1>])'
g.inputs_df = ['score', 'score']
g.inputs = []
g.outputs = []
assert gen.get_sizes(g[0]) == '([array<2,2>], [])'
g.inputs_df = [['score'], ['score','score']]
assert gen.get_sizes(g[0]) == '([array<2,1>, array<2,2>], [])'
g.inputs_df = []
g.outputs = ['score']
assert gen.get_sizes(g[0]) == '([], [array<2,1>])'
g.outputs = ['score',['score']]
with pytest.raises(AssertionError): g[0]
g.outputs = [['score'],['score']]
assert gen.get_sizes(g[0]) == '([], [array<2,1>, array<2,1>])'
# +
d = {'features': [1, 2, 3, 4, 5], 'mask': [1, 0, 1, 1, 0]}
df = pd.DataFrame(data=d)
def filter_features(df):
return np.array(df.loc[df['mask']==1,['features']])
gen_params.update(data_path = None,
outputs = filter_features,
inputs = [],
inputs_df = ['features'],
shuffle = False,
batch_size= 5)
# gen.pretty(gen_params)
g = gr.DataGeneratorHDF5(df, **gen_params)
assert gen.get_sizes(g[0]) == '([array<5,1>], array<3,1>)'
assert all(np.squeeze(g[0][0]) == np.arange(1,6))
assert all(np.squeeze(g[0][1]) == [1,3,4])
# +
m = np.zeros((5,5,3))
c = np.zeros((5,5,3))
c[1:4,1:4,:] = 1
assert np.array_equal(aug.cropout_patch(m, patch_size=(3,3), patch_position=(0.5,0.5), fill_val=1), c)
m = np.zeros((256,256,3))
plt.imshow(aug.cropout_random_patch(m.copy(), patch_size=(128,128), fill_val=1))
plt.show()
plt.imshow(aug.cropout_random_patch(m.copy(), patch_size=(128,128), fill_val=1))
plt.show()
# +
from ku import image_utils as iu
assert isinstance(iu.ImageAugmenter(np.ones(1)), aug.ImageAugmenter)
m = np.zeros((5,5,3))
c = np.zeros((5,5,3))
c[1:4,1:4,:] = 1
assert np.array_equal(aug.cropout_patch(m, patch_size=(3,3), patch_position=(0.5,0.5), fill_val=1), c)
assert np.array_equal(aug.ImageAugmenter(c).cropout((3,3), crop_pos=(0.5,0.5), fill_val=1).result, c)
assert np.array_equal(aug.ImageAugmenter(c).cropout((3,3), crop_pos=(0.5,0.5), fill_val=0).result, m)
assert np.array_equal(aug.ImageAugmenter(c).crop((3,3), crop_pos=(0.5,0.5)).result, np.ones((3,3,3)))
# +
m = np.zeros((5,5,3))
ml, mr = [m]*2
ml[0:2,0:2,:] = 1
mr[0:2,-2:,:] = 1
assert np.array_equal(iu.ImageAugmenter(m).fliplr().result, m)
assert np.array_equal(iu.ImageAugmenter(ml).fliplr().result, mr)
# +
# reload(gr)
def preproc(im, *arg):
if arg:
return np.zeros(im.shape) + arg
else:
return im
gen_params_local = gen_params.copy()
gen_params_local.update(process_fn = preproc,
data_path = 'data.h5',
inputs = ['filename', 'filename1'],
process_args = {'filename' :'args'},
batch_size = 4,
shuffle = False)
ids_local = ids.copy()
ids_local['filename1'] = ids_local['filename']
ids_local['args'] = range(len(ids_local))
ids_local['args1'] = range(len(ids_local),0,-1)
g = gr.DataGeneratorHDF5(ids_local, **gen_params_local)
assert np.array_equal(np.squeeze(g[0][0][0]), np.arange(4))
assert np.array_equal(np.squeeze(g[0][0][1]), np.arange(1,5))
assert np.array_equal(np.squeeze(g[0][1]), np.arange(1,5))
# +
# np.stack is much faster on float32, and still faster for float16 data
data_elem = np.arange(100000, dtype=np.float32)
data = [data_elem.copy() for i in range(10000)]
with gen.Timer('stack, convert float32'):
data_new_stack = np.float32(np.stack(data))
with gen.Timer('iterate, init float32'):
data_new = None
for i, d in enumerate(data):
if data_new is None:
data_new = np.zeros((len(data),)+d.shape, dtype=np.float32)
data_new[i, ...] = d
assert np.array_equal(data_new, data_new_stack)
gen.print_sizes(data_new)
gen.print_sizes(data_new_stack)
# +
reload(gr)
gen_params_ = gen_params.copy()
gen_params_.process_fn = lambda im: [im, im+1]
g = gr.DataGeneratorDisk(ids, **gen_params_)
gen.print_sizes(g[0])
assert np.array_equal(g[0][0][0], g[0][0][1]-1)
assert np.array_equal(g[0][1][0], np.array([[1],[2]]))
# +
def read_fn(*args):
g = args[1]
score = np.float32(g.ids[g.ids.filename==args[0]].score)
return np.ones((3,3)) * score
gen_params_local = gen_params.copy()
gen_params_local.batch_size = 3
gen_params_local.read_fn = read_fn
gen_params_local.process_fn = lambda im: [im, im+1]
g = gr.DataGeneratorDisk(ids, **gen_params_local)
gen.print_sizes(g[0])
print(g[0][0][1])
assert np.array_equal(g[0][0][0], g[0][0][1]-1)
assert np.array_equal(g[0][0][1][0,...], np.ones((3,3))*2.)
# +
m = np.ones((4,4))
assert np.array_equal(aug.imshuffle(m, [2,2]), np.ones((4,4)))
m[:,0] = 0
assert np.sum(aug.imshuffle(m, [4,4])==0)==4
assert np.array_equal(aug.imshuffle(m, [1,1]), m)
m = np.zeros((2,2))
m[0,0] = 1
for _ in range(1000):
assert np.sum(aug.imshuffle_pair(m, m, [2,2]))<=2
assert np.sum(aug.imshuffle_pair(m, 1-m, [2,2]))>=1
# +
m1 = np.ones((4,4))
m2 = np.zeros((4,4))
for _ in range(1000):
for ratio in [0,0.25,0.5,0.75,1]:
assert np.sum(aug.imshuffle_pair(m1, m2, [4,4], ratio)) == ratio*16
assert np.sum(aug.imshuffle_pair(m1, m2, [1,4], 0.6)) == 8
assert np.sum(aug.imshuffle_pair(m1, m2, [1,4], 0.7)) == 12
assert np.sum(aug.imshuffle_pair(m1, m2, [1,4], 0.75)) == 12
assert np.sum(aug.imshuffle_pair(m1, m2, [1,4], 0.8)) == 12
assert np.sum(aug.imshuffle_pair(m1, m2, [1,4], 0.9)) == 16
assert np.sum(aug.imshuffle_pair(m1, m2, [1,4], 1)) == 16
# +
m1 = np.ones((4,4))
m2 = np.zeros((4,4))
mix1 = aug.imshuffle_pair(m1, m2, [2,2], flip=True)
mix2 = aug.imshuffle_pair(m1, m2, [2,2], flip=False)
assert np.sum(mix1) == np.sum(mix2)
# +
size = 10
ids_defa = pd.read_csv(u'ids.csv', encoding='latin-1')
fnames = np.concatenate([ids_defa.filename.values]*3)[:size]
ids = pd.DataFrame(dict(cats = ['cat{}'.format(i) for i in range(size)],
dogs = ['dog{}'.format(i) for i in range(size)],
image_name = fnames,
group = [i//4 for i in range(10)]))
gen_params = Munch(batch_size = 1,
inputs = ['image_name'],
outputs = ['dogs'],
data_path = 'images',
group_by = 'group',
shuffle = False,
fixed_batches = True)
for batch_size, len_g in zip(range(1, 5), [10, 5, 5, 3]):
gen_params.batch_size = batch_size
g = gr.DataGeneratorDisk(ids, **gen_params)
# gen.print_sizes(g[0])
# print('num batches:',len(g))
assert len(g)==len_g
a = g.ids_index.groupby('batch_index').group_by.mean().values
b = g.ids_index.groupby('batch_index').group_by.last().values
assert np.array_equal(a, b)
# +
gen_params.group_by = None
for batch_size, len_g in zip(range(1, 5), [10, 5, 3, 2]):
gen_params.batch_size = batch_size
g = gr.DataGeneratorDisk(ids, **gen_params)
# print('num batches:',len(g))
assert len(g)==len_g
# display(g.ids_index)
gen_params.fixed_batches = False
for batch_size, len_g in zip(range(1, 5), [10, 5, 4, 3]):
gen_params.batch_size = batch_size
g = gr.DataGeneratorDisk(ids, **gen_params)
# print('num batches:',len(g))
assert len(g)==len_g
# display(g.ids_index)
# g.ids_index.sort_values('batch_index')
# iu.view_stack(gen.mapmm(g[0][0][0]))
# +
iu.resize_folder('images/', 'images_temp/',
image_size_dst=(50,50), over_write=True)
image_list = iu.glob_images('images_temp', verbose=False)
assert image_list
ims = iu.read_image_batch(image_list)
assert ims.shape == (4, 50, 50, 3)
failed_images, all_images = iu.check_images('images_temp/')
assert len(failed_images)==0
assert len(all_images)==4
iu.save_images_to_h5('images_temp', 'images.h5',
overwrite=True)
with gr.H5Helper('images.h5') as h:
assert list(h.hf.keys()) == sorted(all_images)
shutil.rmtree('images_temp')
os.unlink('images.h5')
# +
path_src='images/'
path_dst='images_aug/'
def process_gen():
for num_patch in [(i,j) for i in [1,2,4,8] for j in [1,2,4,8]]:
fn = lambda im: aug.imshuffle(im, num_patch)
yield fn, dict(num_patch=num_patch)
ids_aug, errors = iu.augment_folder(path_src, path_dst,
process_gen, verbose=False)
assert len(errors)==0
assert len(ids_aug)==64
(image_path, ext) = os.path.split(ids_aug.iloc[0,:].image_path)
_, file_names = iu.glob_images('{}{}/'.format(path_dst,image_path), split=True)
first_group_names = list(ids_aug.groupby('num_patch'))[0][1].image_name
assert sorted(first_group_names) == sorted(file_names)
shutil.rmtree(path_dst)
# +
iu.resize_folder('images/', 'images1/', image_size_dst=(100,100), overwrite=True)
gp = gen_params.copy()
gp.inputs = ['filename']
gp.group_names = ['images/']
gp.data_path = ''
g = gr.DataGeneratorDisk(ids, **gp)
assert gen.get_sizes(g[0]) == '([array<2,224,224,3>], [array<2,1>])'
gp.group_names = ['images/', 'images1/']
g = gr.DataGeneratorDisk(ids, **gp)
assert gen.get_sizes(g[0]) == '([array<2,224,224,3>, array<2,100,100,3>], [array<2,1>])'
gp.group_names = [['images/'], ['images1/']]
sizes = []
for i in range(100):
g = gr.DataGeneratorDisk(ids, **gp)
sizes.append(g[0][0][0].shape[1])
assert np.unique(sizes).shape[0]>1
# !rm -R images1/
# +
iu.resize_folder('images/', 'base/images100/', image_size_dst=(100,100), overwrite=True)
iu.resize_folder('images/', 'base/images50/', image_size_dst=(50,50), overwrite=True)
gp = gen_params.copy()
gp.inputs = ['filename']
gp.data_path = ''
gp.group_names = ['base']
gp.random_group = True
g = gr.DataGeneratorDisk(ids, **gp)
assert np.array_equal(np.unique([x[0][0].shape[1]
for i in range(100) for x in g]), [50,100])
# !rm -R base/
# +
ids = pd.DataFrame(dict(a = range(10),
b = list(range(9,-1,-1))))
gen_params = Munch(batch_size = 4,
data_path = None,
input_shape = None,
inputs_df = ['a'],
outputs = ['b'],
shuffle = False,
fixed_batches = True)
# check fixed batches switch
g = gr.DataGeneratorDisk(ids, **gen_params)
assert np.array_equal([gen.get_sizes(x) for x in g],
['([array<4,1>], [array<4,1>])',
'([array<4,1>], [array<4,1>])'])
assert np.array_equal(g[0][0][0].squeeze(), range(4))
gen_params.fixed_batches = False
g = gr.DataGeneratorDisk(ids, **gen_params)
assert np.array_equal([gen.get_sizes(x) for x in g],
['([array<4,1>], [array<4,1>])',
'([array<4,1>], [array<4,1>])',
'([array<2,1>], [array<2,1>])'])
assert np.array_equal(g[2][0][0].squeeze(), [8, 9])
# check randomized
gen_params.shuffle = True
gen_params.fixed_batches = False # maintain
g = gr.DataGeneratorDisk(ids, **gen_params)
# check if it returns all items
data = list(zip(*list(g)))
data0 = np.concatenate([l[0] for l in data[0]], axis=0).squeeze()
data1 = np.concatenate([l[0] for l in data[1]], axis=0).squeeze()
assert np.array_equal(np.sort(data0), np.arange(10))
assert np.array_equal(np.sort(data1), np.arange(10))
# check if randomization is applied, consistently
num_randoms0 = 0
num_randoms1 = 0
for i in range(100):
g = gr.DataGeneratorDisk(ids, **gen_params)
data = list(zip(*list(g)))
data0 = np.concatenate([l[0] for l in data[0]], axis=0).squeeze()
data1 = np.concatenate([l[0] for l in data[1]], axis=0).squeeze()
# check consistency
ids_ = ids.copy()
ids_.index = ids_.a
np.array_equal(ids_.loc[data0].b, data1)
num_randoms0 += not np.array_equal(data0, np.arange(10))
num_randoms1 += not np.array_equal(data1, np.arange(10))
# check randomizatino, at least once
assert num_randoms0
assert num_randoms0
# +
from keras.layers import Input
from keras.models import Model
from ku import applications as apps
ids = pd.DataFrame(dict(a = np.arange(100),
b = np.flip(np.arange(100))))
ids = apps.get_train_test_sets(ids)
# display(ids)
X = Input(shape=(1,), dtype='float32')
y = apps.fc_layers(X, name = 'head',
fc_sizes = [5, 1],
dropout_rates = [0, 0],
batch_norm = 0)
model = Model(inputs=X, outputs=y)
gen_params = Munch(batch_size = 4,
data_path = '',
input_shape = (1,),
inputs_df = ['a'],
outputs = ['b'])
helper = mh.ModelHelper(model, 'test_model', ids,
loss = 'MSE',
metrics = ['mean_absolute_error'],
monitor_metric = 'val_mean_absolute_error',
multiproc = False, workers = 2,
logs_root = 'logs',
models_root= 'models',
gen_params = gen_params)
print('Model name:', helper.model_name(test='on'))
helper.update_name()
valid_gen = helper.make_generator(ids[ids.set == 'validation'],
shuffle = False)
valid_gen.batch_size = len(valid_gen.ids)
valid_gen.on_epoch_end()
assert valid_gen.ids_index.batch_index.unique().size == 1
helper.train(lr=1e-1, epochs=50, verbose=False, valid_in_memory=True);
assert path.exists(helper.params.logs_root + '/' + helper.model_name())
helper.load_model(); # best
valid_best1 = helper.validate(verbose=1)
helper.train(lr=1, epochs=10, verbose=False, valid_in_memory=True);
# validate final model
valid_res_fin = helper.validate(verbose=1)
helper.load_model(); # best
valid_best2 = helper.validate(verbose=1)
if valid_res_fin['loss'] > valid_best1['loss']:
assert valid_best1['loss'] == valid_best2['loss']
y_pred = helper.predict(valid_gen)
y_true = ids[ids.set=='validation'].b.values
_, _, val_mae, _ = apps.rating_metrics(y_true, y_pred, show_plot=False);
print(valid_best2)
assert np.abs(val_mae - valid_best2['mean_absolute_error']) < 1e-2
# +
import glob
res = (256, 192)
archive_url = "http://datasets.vqa.mmsp-kn.de/archives/koniq10k_{}x{}.tar".format(*res)
print('download URL:', archive_url)
gen.download_archive(archive_url,'./')
assert os.path.exists('256x192')
assert len(glob.glob('256x192/*')) == 10373
shutil.rmtree('256x192')
archive_url = "http://datasets.vqa.mmsp-kn.de/archives/koniq10k_{}x{}_test.zip".format(*res)
print('download URL:', archive_url)
gen.download_archive(archive_url,'./')
assert os.path.exists('256x192')
assert len(glob.glob('256x192/*')) == 2015
shutil.rmtree('256x192')
# +
gen_params_local = gen_params.copy()
ids_local = ids.copy()
def ids_fn():
ids_local.score = -ids_local.score
return ids_local
gen_params_local.ids_fn = ids_fn
gen_params_local.batch_size = 4
g = gr.DataGeneratorDisk(ids, **gen_params_local)
x = g[0][1][0]
g.on_epoch_end()
y = g[0][1][0]
assert np.array_equal(-x, y)
| Model/ku/tests/tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import requests
from bs4 import BeautifulSoup
from tqdm import tqdm
# +
url = 'https://www.hikingupward.com/'
data = requests.get(url)
data.status_code
# print(data.text)
soup = BeautifulSoup(data.text, 'html.parser')
# soup
# -
##TODO: convert to function/class
url_links = [url + url_link.attrs.get('href')
for nav in soup.find_all('div', {'class': 'navigation'})
for url_link in nav.find_all('a', href = True)]
print(len(url_links))
# url_links
# Scrape these features:
#
# hike_len_in_mi, difficulty_rating, streams_rating, views_rating, solitude_rating, camping_rating,
#
# hiking_time, elevation_gain
# +
## TODO: implement else statement for if there is more than one hike on a page
## TODO: scrape the geocoordinates of the parking lot
# +
hike = []
def get_hike_data(hiking_upward_url: str) -> pd.DataFrame:
hike_content = requests.get(hiking_upward_url)
if hike_content.status_code == 200:
try:
hike_soup = BeautifulSoup(hike_content.text, 'html.parser')
tables = hike_soup.find('table', {'class': 'hike_pages'})
if len(tables.find_all('tr')) == 5:
rating_table = tables.find_all('tr')[1].find_all('td')
time_elev = tables.find_all('tr')[2].find_all('td')[1]
try:
hike_len_in_mi = float(rating_table[0].text.replace("mls", "").strip())
except:
hike_len_in_mi = np.nan
try:
difficulty_rating = int(rating_table[1].find('img').get('src').replace("../../images/stars/Star", "").replace("/images/stars/Star", "").replace("_clear.gif", "").replace("_grey.gif", "").replace("_red.gif", ""))
except:
difficulty_rating = np.nan
try:
streams_rating = int(rating_table[2].find('img').get('src').replace("../../images/stars/Star", "").replace("/images/stars/Star", "").replace("_clear.gif", "").replace("_grey.gif", "").replace("_red.gif", ""))
except:
streams_rating = np.nan
try:
views_rating = int(rating_table[3].find('img').get('src').replace("../../images/stars/Star", "").replace("/images/stars/Star", "").replace("_clear.gif", "").replace("_grey.gif", "").replace("_red.gif", ""))
except:
views_rating = np.nan
try:
solitude_rating = int(rating_table[4].find('img').get('src').replace("../../images/stars/Star", "").replace("/images/stars/Star", "").replace("_clear.gif", "").replace("_grey.gif", "").replace("_red.gif", ""))
except:
solitude_rating = np.nan
try:
camping_rating = int(rating_table[5].find('img').get('src').replace("../../images/stars/Star", "").replace("/images/stars/Star", "").replace("_clear.gif", "").replace("_grey.gif", "").replace("_red.gif", ""))
except:
camping_rating = np.nan
##av.note: to be implemented later
# try:
# hiking_duration = str(time_elev.contents[0].strip())
# except:
# hiking_duration = np.nan
# try:
# elevation_gain_ft = float(time_elev.contents[2].replace('ft', '').replace(',', '').replace('with three different ascents', '').replace('with multiple ascents', '').replace('with two ascents', '').replace('with two different ascents', '').strip())
# except:
# elevation_gain_ft = np.nan
df = pd.DataFrame({'hike_url': str(hiking_upward_url),
'hike_name': str(hike_soup.title.text),
'park_abbreviation': hiking_upward_url.replace('https://www.hikingupward.com/', '').split('/', 1)[0],
'hike_len_in_mi': hike_len_in_mi,
'difficulty_rating': difficulty_rating,
'streams_rating': streams_rating,
'views_rating': views_rating,
'solitude_rating': solitude_rating,
'camping_rating': camping_rating,
# 'hiking_duration_str': hiking_duration,
# 'elevation_gain_ft': elevation_gain_ft
},
index = [0])
return df
except Exception:
pass
# +
# test_link= url_links[0]
# print(test_link)
# test_df = get_hike_data(test_link)
# test_df
# -
for url_link in tqdm(url_links):
one_hike_df = get_hike_data(url_link)
hike.append(one_hike_df)
hike_df = pd.concat(hike, ignore_index = True)
print(hike_df.shape)
# ## Data quality checks
hike_df.sample(10)
##av.notes:
## according to the sortable map:
## there are no hikes with difficulty 0
## streams, views,solitude, and camping should all have ratings from 0-6
hike_df.describe()
# +
# hike_df.info()
# -
print(hike_df.park_abbreviation.isna().sum())
# print(hike_df.park_abbreviation.value_counts())
## percent of hikes in each park
hike_df.park_abbreviation.value_counts(normalize = True)*100
# +
# hike_df[hike_df.hike_len_in_mi.isna()]
# +
##av.note: to be implemented later
# hike_df[hike_df.elevation_gain_ft.isna()].sample(10)
# -
hike_df[hike_df.streams_rating.isna()].sample(5)
hike_df[hike_df.camping_rating.isna()].sample(5)
hike_df[hike_df.views_rating.isna()].sample(5)
# +
## these are probably ones that have two hikes!
# +
##av.note: to be implemented later
## TODO: figure out how to scrape these (something to do with the font attribute, maybe?)
# print(hike_df[hike_df.elevation_gain_ft.isna() & hike_df.hiking_duration_str.isna()].shape)
# hike_df[hike_df.elevation_gain_ft.isna() & hike_df.hiking_duration_str.isna()]
# -
# ## Data cleaning
## will need to replace NaN for Brandywine Recreation Area Hike
hike_df.streams_rating.fillna(0, inplace = True)
hike_df.camping_rating.fillna(0, inplace = True)
hike_df.views_rating.fillna(0, inplace = True)
# +
##av.note: to be implemented later
## get the state based on park abbreviation
# hike_df['state'] = 'TBD'
# -
hike_df.describe()
# ## Write Data
hike_df.to_csv("../data/hiking_upward_data.csv", index = False)
# ### exploration that made the above function possible
hike_data = requests.get(url_links[0])
hike_soup = BeautifulSoup(hike_data.text, 'html.parser')
# print(hike_soup)
hike_soup.title.text
# +
## exploration
# for tables in hike_soup.find_all('table', {'class': 'hike_pages'}):
# for table in tables.find_all('tr'):
# print(table)
# print("\n")
# # for row in table.find_all('td'):
# # print(row)
# # print("\n")
# -
tables = hike_soup.find('table', {'class': 'hike_pages'})
print(len(tables.find_all('tr')))
tables
tables.find_all('tr')[0].find_all('td')
rating_table = tables.find_all('tr')[1].find_all('td')
rating_table
hike_len_in_mi = float(rating_table[0].text.replace(" mls", ""))
print(hike_len_in_mi) # str - convert to float
difficulty_rating = int(rating_table[1].find('img').get('src').replace("../../images/stars/Star", "").replace("_clear.gif", ""))
print(difficulty_rating) #str - convert to int
streams_rating = int(rating_table[2].find('img').get('src').replace("../../images/stars/Star", "").replace("_clear.gif", ""))
print(streams_rating) #str - convert to int
views_rating = int(rating_table[3].find('img').get('src').replace("../../images/stars/Star", "").replace("_clear.gif", ""))
print(views_rating) #str - convert to int
solitude_rating = int(rating_table[4].find('img').get('src').replace("../../images/stars/Star", "").replace("_clear.gif", ""))
print(solitude_rating) #str - convert to int
camping_rating = int(rating_table[5].find('img').get('src').replace("../../images/stars/Star", "").replace("_clear.gif", ""))
print(camping_rating) #str - convert to int
time_elev = tables.find_all('tr')[2].find_all('td')[1]
# print(time_elev)
hiking_duration = time_elev.contents[0]
print(hiking_duration) #str
elevation_gain_ft = time_elev.contents[2].replace(' ft', '').replace(',', '')
print(elevation_gain_ft) #str - convert to int/float
| notebooks/get_hike_urls_and_data.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ + [markdown] cell_id="00002-e5896fa3-893d-4d3a-8977-4fd139140eae" deepnote_cell_type="markdown" tags=[]
/ 1. Functions are first class values
/ - Functions can be defined with in other functions
/ - Functions can be passed as params to other functions
/ - Functions can be return types of functions
/ 2. Immutability
/ - operations on objects creates new objects rather than modifying original
/ 3. Pure functions
/
/ > Pure Function = O/p depends on I/p + No side effects
/ - A pure function cannot rely on input from files, databases, web services etc
/ - always produces same output with given input
/
/ <figure>
/ <img src="https://user-images.githubusercontent.com/8268939/79669398-0a754800-8170-11ea-943c-bc19a5b65506.jpeg" width="400" height="200"/>
/ <figcaption>Fig - Pure function </figcaption>
/ </figure>
/ + [markdown] cell_id="00003-6ca15ea2-a6ae-4d24-8c6a-0051a37c1d43" deepnote_cell_type="markdown" tags=[]
/
/ + cell_id="00002-d6e98fa6-280c-4d41-98dc-a9dfa6c9a9a0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1754 execution_start=1615144586912 source_hash="d8f15c16" tags=[]
// pure function
def add(a:Int, b:Int):Int = {
return a + b
}
/ + cell_id="00003-d84f5e34-bbd3-4fe5-b414-f2c2aa16b4f7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1895 execution_start=1615144613915 source_hash="76c43eec" tags=[]
// Impure function
// returns different result every time called
def getSysTime:Long = {
return System.currentTimeMillis()
}
/ + [markdown] cell_id="00004-1386356c-8e73-4767-8e5f-2c0673646778" deepnote_cell_type="markdown" tags=[]
/ ## Block in scala
/
/ Blocks are themselves expressions; a block may appear everywhere an expression can
/
/ ### Blocks and visibility
/ + cell_id="00004-d0fcfe0b-f81b-4c00-8a6d-6bfbd4d84a5f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3945 execution_start=1615145301593 source_hash="e3521cf3" tags=[]
val x = 0
def f(y: Int) = y + 1 // whatever defined outside block can be accessed from inside block below
val result = {
val x = f(3) // whatever defined inside block accessible only here, this overwrites the definition of x outside
x * x
}
| scala/introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tape
# language: python
# name: tape
# ---
# # Multitask Learning Experiments, and Q2 Classification
#
# This notebook includes experiments with multitask learning using Keras, as well as classification and metrics for the q2 membrane bound vs water soluble task that is included as part of the DeepLoc dataset
# +
import json
from Bio import SeqIO
from Bio.Alphabet import IUPAC
from Bio.SeqRecord import SeqRecord
from Bio.Seq import Seq
import keras
from keras import backend as K
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout, BatchNormalization
from keras.utils import to_categorical
from keras.callbacks import EarlyStopping
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import torch
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix, classification_report
import tensorflow as tf
# -
np.random.seed(42)
# ### Data Loading
# +
seq_id_to_q2 = {}
seq_id_to_q10 = {}
seq_list = []
for record in SeqIO.parse("../data/deeploc_data_6000.fasta", "fasta"):
description = record.description
seq = record.seq
seq_list.append(len(str(seq)))
desc_split = description.split(" ")
# Use ID for indexing
ID = desc_split[0]
# Split label into Q10 subcellular location and Q2 membrane bound vs water soluble
label = desc_split[1]
loc_label = label[:len(label)-2]
mem_label = label[-1]
seq_id_to_q10[ID] = loc_label
seq_id_to_q2[ID] = mem_label
# -
# Look at percentiles for sequence length
seq_df = pd.DataFrame(seq_list)
seq_df.describe(percentiles=[.90, .95, .99])
train_arrays = np.load('../data/output_deeploc_train_6000.npz', allow_pickle=True)
valid_arrays = np.load('../data/output_deeploc_valid_6000.npz', allow_pickle=True)
test_arrays = np.load('../data/output_deeploc_test_6000.npz', allow_pickle=True)
# +
id_dict = {'Cell.membrane': 0,
'Cytoplasm': 1,
'Endoplasmic.reticulum': 2,
'Golgi.apparatus': 3,
'Lysosome/Vacuole': 4,
'Mitochondrion': 5,
'Nucleus': 6,
'Peroxisome': 7,
'Plastid': 8,
'Extracellular': 9}
id_dict_mem = {
'M': 0,
'S': 1,
'U': 2
}
reverse_id_dict = {value: key for key, value in id_dict.items()}
reverse_id_dict_mem = {value: key for key, value in id_dict_mem.items()}
# -
def gen_df(embeddings):
"""
Iterate over all of the sequence IDs in the given subset of the dataset (embeddings),
as a nested numpy array. Produce a numpy array of the average embeddings for each
sequence, as will a list of the labels by looking up the sequence IDs in seq_id_to_label
Args:
embeddings (numpy.lib.npyio.NpzFile): Nested numpy array containing embeddings for each sequence ID
seq_id_to_label (dict[str,str]): Map from sequence ID to classification label
Returns:
output (pd.DataFrame): Average embeddings for each sequence
labels (list[str])
"""
keys = embeddings.files
output, loc_labels, mem_labels = [], [], []
for key in keys:
d = embeddings[key].item()["avg"]
loc_labels.append(seq_id_to_q10[key])
mem_labels.append(seq_id_to_q2[key])
output.append(d)
return pd.DataFrame(output), loc_labels, mem_labels
train_df, train_loc_labels, train_mem_labels = gen_df(train_arrays)
valid_df, valid_loc_labels, valid_mem_labels = gen_df(valid_arrays)
test_df, test_loc_labels, test_mem_labels = gen_df(test_arrays)
# Look at distribution of classes for binary classification task. U stands for unknown - we can skip those samples
pd.Series(train_mem_labels).value_counts()
# ### Logistic Regression (Q2)
#
# Applying Logistic Regression to the membrane bound vs water soluble q2 classification problem, we were able to obtain 87% accuracy on the test set.
train_valid_df = train_df.append(valid_df, ignore_index=True)
train_valid_mem_labels = np.array(train_mem_labels + valid_mem_labels)
train_valid_df = train_valid_df[train_valid_mem_labels != 'U']
train_valid_mem_labels = train_valid_mem_labels[train_valid_mem_labels != 'U']
test_mem_labels_q2 = np.array(test_mem_labels)
test_mem_df = test_df[test_mem_labels_q2 != 'U']
test_mem_labels_q2 = test_mem_labels_q2[test_mem_labels_q2 != 'U']
train_valid_df.shape
clf = LogisticRegression(random_state=42)
clf.fit(train_valid_df, train_valid_mem_labels)
clf.score(train_valid_df, train_valid_mem_labels)
clf.score(test_mem_df, test_mem_labels_q2)
logreg_q2_test_preds = clf.predict(test_mem_df)
# ### Keras DNN (Q2)
#
# Applying a Keras DNN with two hidden layers of 32 nodes to the membrane bound vs water soluble q2 classification problem, we were able to obtain 89% accuracy on the test set.
#
# The DeepLoc dataset has many sequences with an unknown label for the q2 task, so we are excluding those samples from training.
# +
train_mem_labels_q2 = np.array(train_mem_labels)
valid_mem_labels_q2 = np.array(valid_mem_labels)
test_mem_labels_q2 = np.array(test_mem_labels)
train_df_q2 = train_df[train_mem_labels_q2 != 'U']
valid_df_q2 = valid_df[valid_mem_labels_q2 != 'U']
test_df_q2 = test_df[test_mem_labels_q2 != 'U']
train_mem_labels_q2 = train_mem_labels_q2[train_mem_labels_q2 != 'U']
valid_mem_labels_q2 = valid_mem_labels_q2[valid_mem_labels_q2 != 'U']
test_mem_labels_q2 = test_mem_labels_q2[test_mem_labels_q2 != 'U']
train_lab_mem = [id_dict_mem[label] for label in train_mem_labels_q2]
valid_lab_mem = [id_dict_mem[label] for label in valid_mem_labels_q2]
test_lab_mem = [id_dict_mem[label] for label in test_mem_labels_q2]
# -
model = Sequential()
model.add(Dense(32, input_dim=768, activation='relu'))
model.add(Dense(32, input_dim=768, activation='relu'))
model.add(Dense(2, activation='sigmoid'))
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
keras_callbacks = [
EarlyStopping(monitor='val_loss', patience=30, mode='min', min_delta=0.0001)
]
history = model.fit(train_df_q2, to_categorical(train_lab_mem),
validation_data=(valid_df_q2, to_categorical(valid_lab_mem)),
epochs=500, batch_size=64, callbacks=keras_callbacks)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model Accuracy (Q2)')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Valid'], loc='upper left')
plt.show()
model.evaluate(test_df_q2, to_categorical(test_lab_mem))
dnn_q2_test_probs = model.predict(test_df_q2)
dnn_q2_test_preds_num = np.argmax(dnn_q2_test_probs, axis=1)
dnn_q2_test_preds = [reverse_id_dict_mem[pred] for pred in dnn_q2_test_preds_num]
# ### Logistic Regression (Q10)
#
# Applying Logistic Regression to the subcellular location (q10) classification task, we were able to obtain 66% accuracy on the test set.
train_valid_df = train_df.append(valid_df, ignore_index=True)
train_valid_loc_labels = train_loc_labels + valid_loc_labels
print(train_valid_df.shape)
print(len(train_loc_labels))
print(len(valid_loc_labels))
clf = LogisticRegression(random_state=42)
clf.fit(train_valid_df, train_valid_loc_labels)
clf.score(train_valid_df, train_valid_loc_labels)
clf.score(test_df, test_loc_labels)
logreg_test_preds = clf.predict(test_df)
# ### Keras DNN (Q10)
#
# Applying a Keras DNN with two hidden layers of 32 nodes each to the subcellular location (q10) classification task, we were able to obtain 65% accuracy on the test set.
train_lab = [id_dict[label] for label in train_loc_labels]
valid_lab = [id_dict[label] for label in valid_loc_labels]
test_lab = [id_dict[label] for label in test_loc_labels]
model = Sequential()
model.add(Dense(32, input_dim=768, activation='relu'))
model.add(Dense(32, activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
keras_callbacks = [
EarlyStopping(monitor='val_loss', patience=50, mode='min', min_delta=0.0001)
]
history = model.fit(train_df, to_categorical(train_lab),
validation_data=(valid_df, to_categorical(valid_lab)),
epochs=500, batch_size=64, callbacks=keras_callbacks)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Valid'], loc='upper left')
plt.show()
model.evaluate(test_df, to_categorical(test_lab))
dnn_test_probs = model.predict(test_df)
dnn_test_preds_num = np.argmax(dnn_test_probs, axis=1)
dnn_test_preds = [reverse_id_dict[pred] for pred in dnn_test_preds_num]
# ### Metrics
#
# Here, we take a look at the classification report (precision and recall) as well as the confusion matrix for all four models above (logistic regression q2, logistic regression q10, keras DNN q2, keras DNN q10).
#
# One of the major takeaways is that for the subcellular location task, the model performance is considerably worse on the classes that appear least frequently in the dataset, such as Golgi.apparatus, Lysosome/Vacuole, and Peroxisome. Also, the Cytoplasm and Nucleus classes are commonly confused, which is in line with our intuition that they have similar embedding spaces after looking at the PCA results in notebook 4.
# #### Logistic Regression (Q2)
print("Logistic Regression Classification Report (Q2)\n")
print(classification_report(test_mem_labels_q2, logreg_q2_test_preds))
logreg_cm = confusion_matrix(test_mem_labels_q2, logreg_q2_test_preds)
df_cm = pd.DataFrame(logreg_cm, index=sorted(id_dict_mem.keys())[:2], columns=sorted(id_dict_mem.keys())[:2])
plt.figure(figsize = (10,7))
fig = sns.heatmap(df_cm, annot=True, cmap='Blues', fmt='g')
plt.xlabel("True Labels")
plt.ylabel("Predicted Labels")
plt.title("Logistic Regression Confusion Matrix (Q2)")
plt.show(fig)
# #### Logistic Regression (Q10)
print("Logistic Regression Classification Report (Q10)\n")
print(classification_report(test_loc_labels, logreg_test_preds))
logreg_cm = confusion_matrix(test_loc_labels, logreg_test_preds)
df_cm = pd.DataFrame(logreg_cm, index=sorted(id_dict.keys()), columns=sorted(id_dict.keys()))
plt.figure(figsize = (10,7))
fig = sns.heatmap(df_cm, annot=True, cmap='Blues', fmt='g')
plt.xlabel("True Labels")
plt.ylabel("Predicted Labels")
plt.title("Logistic Regression Confusion Matrix")
plt.show(fig)
# #### Keras DNN (Q2)
print("Keras DNN Classification Report Q2\n")
print(classification_report(test_mem_labels_q2, dnn_q2_test_preds))
dnn_cm = confusion_matrix(test_mem_labels_q2, dnn_q2_test_preds)
df_cm = pd.DataFrame(dnn_cm, index=sorted(id_dict_mem.keys())[:2], columns=sorted(id_dict_mem.keys())[:2])
plt.figure(figsize = (10,7))
fig = sns.heatmap(df_cm, annot=True, cmap='Blues', fmt='g')
plt.xlabel("True Labels")
plt.ylabel("Predicted Labels")
plt.title("Keras DNN Confusion Matrix Q2")
plt.show(fig)
# #### Keras DNN (Q10)
print("Keras DNN Classification Report Q10\n")
print(classification_report(test_loc_labels, dnn_test_preds))
dnn_cm = confusion_matrix(test_loc_labels, dnn_test_preds)
df_cm = pd.DataFrame(dnn_cm, index=sorted(id_dict.keys()), columns=sorted(id_dict.keys()))
plt.figure(figsize = (10,7))
fig = sns.heatmap(df_cm, annot=True, cmap='Blues', fmt='g')
plt.xlabel("True Labels")
plt.ylabel("Predicted Labels")
plt.title("Keras DNN Confusion Matrix Q10")
plt.show(fig)
# ### Multitask Learning
# #### Q10 / Q3
#
# Treating membrane bound vs water soluble q2 classification as a three class problem by including the unknown class to prevent having missing samples for the q10 subcellular location task.
#
# The model is a Keras DNN w/ two hidden layers of 32 nodes each, and two output heads for the membrane bound vs water soluble vs unkonwn task, and the subcellular location task, each with a softmax activation function.
#
# The results are as follows:
# * 71% accuracy on membrane bound vs water soluble protein
# * 64% accuracy on subcellular location
# +
# Create arrays for all labels (loc=q10, mem=q3)
train_loc_labels_mtu = np.array(train_loc_labels)
valid_loc_labels_mtu = np.array(valid_loc_labels)
test_loc_labels_mtu = np.array(test_loc_labels)
train_mem_labels_mtu = np.array(train_mem_labels)
valid_mem_labels_mtu = np.array(valid_mem_labels)
test_mem_labels_mtu = np.array(test_mem_labels)
# Create new label lists with ints instead of strings
train_lab_loc_u = [id_dict[label] for label in train_loc_labels_mtu]
valid_lab_loc_u = [id_dict[label] for label in valid_loc_labels_mtu]
test_lab_loc_u = [id_dict[label] for label in test_loc_labels_mtu]
train_lab_mem_u = [id_dict_mem[label] for label in train_mem_labels_mtu]
valid_lab_mem_u = [id_dict_mem[label] for label in valid_mem_labels_mtu]
test_lab_mem_u = [id_dict_mem[label] for label in test_mem_labels_mtu]
# +
keras.backend.clear_session()
visible = keras.Input(shape=(768,))
h1 = Dense(32, activation='relu')(visible)
h2 = Dense(32, activation='relu')(h1)
output_q10 = Dense(10, activation='softmax', name='q10')(h2)
output_q3 = Dense(3, activation='softmax', name='q3')(h2)
model = keras.Model(inputs=visible, outputs=[output_q10, output_q3])
model.compile(loss=['categorical_crossentropy', 'categorical_crossentropy'],
optimizer='adam', metrics=['accuracy', 'accuracy'])
# -
keras_callbacks = [
EarlyStopping(monitor='val_loss', patience=30, mode='min', min_delta=0.0001)
]
history = model.fit(train_df,
[to_categorical(train_lab_loc_u), to_categorical(train_lab_mem_u)],
validation_data=(valid_df, [to_categorical(valid_lab_loc_u), to_categorical(valid_lab_mem_u)]),
epochs=200, batch_size=64, callbacks=keras_callbacks)
plt.plot(history.history['q10_accuracy'])
plt.plot(history.history['val_q10_accuracy'])
plt.plot(history.history['q3_accuracy'])
plt.plot(history.history['val_q3_accuracy'])
plt.title('Model Accuracy (Multitask w/ Unknown)')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train (q10)', 'Valid (q10)', 'Train (q3)', 'Valid (q3)'], loc='upper left')
plt.show()
plt.plot(history.history['q10_loss'])
plt.plot(history.history['val_q10_loss'])
plt.plot(history.history['q3_loss'])
plt.plot(history.history['val_q3_loss'])
plt.title('Model Loss (Multitask w/ Unknown)')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train (q10)', 'Valid (q10)', 'Train (q3)', 'Valid (q3)'], loc='upper left')
plt.show()
model.evaluate(test_df, [to_categorical(test_lab_loc_u), to_categorical(test_lab_mem_u)])
# #### Masked Loss Function
#
# Using a masked loss function to `mask` the loss from the unknown labels for the binary classification task, so that we can use all of the samples and still have q10 / q2 classification tasks.
id_dict_mem_masked = {
'M': 0,
'S': 1,
'U': -1
}
# +
# Create new label lists with ints instead of strings
train_lab_loc_u = [id_dict[label] for label in train_loc_labels_mtu]
valid_lab_loc_u = [id_dict[label] for label in valid_loc_labels_mtu]
test_lab_loc_u = [id_dict[label] for label in test_loc_labels_mtu]
train_lab_mem_u = [id_dict_mem[label] for label in train_mem_labels_mtu]
valid_lab_mem_u = [id_dict_mem[label] for label in valid_mem_labels_mtu]
test_lab_mem_u = [id_dict_mem[label] for label in test_mem_labels_mtu]
# +
# This is where we mask out the unknown labels
mask_value = -1
def masked_loss_function(y_true, y_pred):
mask = K.cast(K.not_equal(y_true, mask_value), K.floatx())
return K.binary_crossentropy(y_true * mask, y_pred * mask)
# +
keras.backend.clear_session()
visible = keras.Input(shape=(768,))
h1 = Dense(32, activation='relu')(visible)
hf = Dense(32, activation='relu')(h1)
# NOTE: These commented out layers duplicate the diagram from the DeepLoc paper,
# but did not perform as well as a DNN with two hidden layers of 32 nodes
# h1 = Dense(32, activation='relu')(visible)
# h2 = Dropout(0.25)(h1)
# hf = BatchNormalization()(h2)
output_q10 = Dense(10, activation='softmax', name='q10')(hf)
output_q2 = Dense(3, activation='softmax', name='q2')(hf)
model = keras.Model(inputs=visible, outputs=[output_q10, output_q2])
model.compile(loss=['categorical_crossentropy', masked_loss_function],
optimizer='adam', metrics=['accuracy', 'accuracy'])
keras_callbacks = [
EarlyStopping(monitor='val_loss', patience=20, mode='min', min_delta=0.0001)
]
history = model.fit(train_df,
[to_categorical(train_lab_loc_u), to_categorical(train_lab_mem_u)],
validation_data=(valid_df, [to_categorical(valid_lab_loc_u), to_categorical(valid_lab_mem_u)]),
epochs=200, batch_size=64, callbacks=keras_callbacks)
# -
plt.plot(history.history['q10_accuracy'])
plt.plot(history.history['val_q10_accuracy'])
plt.plot(history.history['q2_accuracy'])
plt.plot(history.history['val_q2_accuracy'])
plt.title('Model Accuracy (Masked Multitask)')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train (q10)', 'Valid (q10)', 'Train (q2)', 'Valid (q2)'], loc='upper left')
plt.show()
plt.plot(history.history['q10_loss'])
plt.plot(history.history['val_q10_loss'])
plt.plot(history.history['q2_loss'])
plt.plot(history.history['val_q2_loss'])
plt.title('Model Loss (Masked Multitask)')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train (q10)', 'Valid (q10)', 'Train (q2)', 'Valid (q2)'], loc='upper left')
plt.show()
# Masked Loss Function Results
# * 72% accuracy on membrane bound vs water soluble protein
# * 64% accuracy on subcellular location
model.evaluate(test_df, [to_categorical(test_lab_loc_u), to_categorical(test_lab_mem_u)])
| notebooks/6_multitask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (nerds_env)
# language: python
# name: nerds_env
# ---
import pandas as pd
import py2neo
import os
NEO4J_CONN_URL = "bolt://localhost:7687"
NEO4J_USER = "neo4j"
NEO4J_PASS = "<PASSWORD>"
pd.options.display.max_colwidth = 100
graph = py2neo.Graph(NEO4J_CONN_URL, auth=(NEO4J_USER, NEO4J_PASS))
# # Find Important Nodes
# for this to work you need to install and configure pagerank algorithm in Neoj4 (follow docs)
def get_nodes_by_pagerank(graph, ent_type):
query = """
CALL algo.pageRank.stream('%s', 'REL', {iterations:20, dampingFactor:0.85})
YIELD nodeId, score
RETURN algo.asNode(nodeId).ename AS page, score
ORDER BY score DESC
""" % (ent_type)
results = graph.run(query).data()
return pd.DataFrame(results)
important_entities_df = get_nodes_by_pagerank(graph, "ENTITY")
important_entities_df.head(10)
# # Find interesting neighbors
def get_neighbors_by_type(graph, src_name, src_type, neighbor_type):
query = """
MATCH (e1:%s {ename:"%s"})<-[r:REL]->(e2:%s)
RETURN e1.ename AS src, e2.ename AS dst
""" % (src_type, src_name, neighbor_type)
results = graph.run(query).data()
results_df = (pd.DataFrame(results)
.groupby(["src", "dst"])["dst"]
.count()
.reset_index(name="count")
.sort_values("count", ascending=False)
)
return results_df
djt_per_neighbors_df = get_neighbors_by_type(graph, "GZMB_protein_human", "ENTITY", "ENTITY")
djt_per_neighbors_df.head()
# +
# FIXME: change to postgres
# def build_sentence_dictionary(sent_file):
# sent_dict = {}
# fsent = open(sent_file, "r")
# for line in fsent:
# pid, sid, sent_text = line.strip().split('\t')
# sent_dict[sid] = sent_text
# fsent.close()
# return sent_dict
# sent_dict = build_sentence_dictionary(os.path.join(DATA_DIR, "sentences.tsv"))
# len(sent_dict)
# +
# FIXME: change use of sent dictionary
# def show_connecting_sentences(graph, src_name, src_type, dst_name, dst_type, sent_dict):
# query = """
# MATCH (e1:%s {ename:"%s"})<-[r:REL]->(e2:%s {ename:"%s"})
# RETURN e1.ename AS src, e2.ename AS dst, r.sid AS sid
# ORDER BY sid
# """ % (src_type, src_name, dst_type, dst_name)
# result = graph.run(query).data()
# result_df = pd.DataFrame(result)
# result_df["sent_text"] = result_df["sid"].apply(lambda x: sent_dict[x])
# return result_df
# +
# EXAMPLE:
# djt_db_rel_df = show_connecting_sentences(graph, "<NAME>", "PER", "Deutsche Bank", "ORG", sent_dict)
# djt_db_rel_df.head(10)
# -
# # Find path connecting a pair of nodes
# +
# def get_path_between(graph, src_name, src_type, dst_name, dst_type):
# query = """
# MATCH (start:%s {ename:'%s'}), (end:%s {ename:'%s'})
# CALL algo.shortestPath.stream(start, end)
# YIELD nodeId, cost
# RETURN algo.asNode(nodeId).ename AS name, cost
# """ % (src_type, src_name, dst_type, dst_name)
# results = graph.run(query).data()
# path = [x["name"] for x in results]
# return path
# +
# djt_putin_link = get_path_between(graph, "<NAME>", "PER", "<NAME>", "PER")
# print(djt_putin_link)
| old_code/ExploreGraph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set_style("whitegrid")
sns.set_context("notebook", font_scale=2.5, rc={"lines.linewidth": 3.5})
bench = pd.DataFrame(index=[1, 2, 3, 4])
for lang in ['cpp', 'java', 'rust']:
bench[lang] = pd.read_csv('nprimes/{}_bench.csv'.format(lang)).set_index('nthreads')
bench.plot(xticks=[1,2, 3, 4], figsize=(16, 8))
plt.xlabel('number of threads')
plt.ylabel('time, sec')
plt.savefig('bench.png', transparent=True)
echo_bench = pd.read_csv('echod/bench.csv', index_col='lang')
class KFormater(matplotlib.ticker.Formatter):
def __call__(self, x, pos):
if x == 0:
return "0"
return "{:0.0f}K".format(x/1000)
echo_bench.plot.barh(alpha=0.5)
plt.axes().xaxis.set_major_formatter(KFormater())
plt.legend().set_visible(False)
plt.axes().set_ylabel("")
plt.axes().set_xticks(range(0, 50000, 10000))
plt.savefig('rps.png', dpi=600, transparent=True, bbox_inches='tight')
| plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="sZGgKYM8-Wqk"
# ## Линейная регрессия
#
# Задача регрессии похожа на задачу классификации, только вместо категории мы предсказываем число. Например стоимость автомобиля по его пробегу, количество пробок по времени суток и т.д.
#
# Линейная регрессия ищет линейную зависимость между объектом и ответом.
#
# ### Пример
# * $\mathbb{X} = \mathbb{R}$
# * $\mathbb{Y} = \mathbb{R}$
# * Найти такие $w, b \in \mathbb{R}$, что $w\cdot x_i + b = y_i$ для всех $i$.
#
# Иногда такую задачу называют Simple Linear Regression из-за её одномерности.
# + colab={} colab_type="code" id="EtQraQNY-Wqq"
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
import warnings
warnings.simplefilter("ignore")
from IPython.display import Image
# %matplotlib inline
sns.set(style="darkgrid")
SEED=31415
np.random.seed(31415)
# + colab={} colab_type="code" id="brj3dwDY-Wqv" outputId="2e0544ae-26ff-4d88-ccbd-3eb6c1870426"
x = np.arange(0, 10, 0.5)[:,np.newaxis]
y = np.sin(x) + x
plt.scatter(x, y)
plt.xlabel("x")
plt.ylabel("y")
# + [markdown] colab_type="text" id="2TpTk0hA-Wq1"
# Наша задача найти $a\cdot x_i + b = y_i$ заметим, что это то же самое что провести прямую, проходящую максимально близко с этими точками. Проведём какую-нибудь прямую.
# + colab={} colab_type="code" id="4w4hfq4d-Wq2" outputId="a10d8155-d5b7-4b49-d289-f034653376c6"
w, b = 1, 1
plt.scatter(x, y)
plt.plot(x, w * x + b, color='red')
plt.xlabel("x")
plt.ylabel("y")
# + [markdown] colab_type="text" id="tCLyjyrF-Wq7"
# Выглядит так, как будто это не самая оптимальная прямая.
#
# 
#
# А как вообще определять насколько прямая оптимальна? Что будет функцией качества? Существует несколько функций качества регрессии. Давайте рассмотрим их:
#
# * mean absolute error: $MAE(X) = \frac{1}{n}\sum_{i=1}^{n}|a(x_i) - y_i|$
# * mean squared error: $MSE(X) = \frac{1}{n}\sum_{i=1}^{n}(a(x_i) - y_i)^2$
#
# Давайте изучим их свойства.
# 1. Интерпретируемость. Допустим, вы хотите предсказать стоимость автомобиля. Что будет означать MSE в тако случае? А MAE?
# 1. Оценка качества модели. Такие функции ошибок позваляют легко сравнивать модели между собой, но что они гооврят о том насколько хорошо работает сама модель? Будет ли хорошим результат MSE=10 или MAE=10 на выборке, где все $y_i$ лежат в отрезке $[0,1]$? А в отрезке $[10^5,10^6]$.
# 1. Дифференцируемость (а это тут при чём вообще?!).
# 1. Устойчивость к выбросам. Какая из функций наиболее устойчива к выбросам? (изучим этот вопрос далее)
#
# Рассмотрим вопрос устойчивости к выбросам чуть подробнее. Реализуйте функции MSE и MAE изменив не более одной строчки кода.
# + colab={} colab_type="code" id="vj7ymeYb-Wq8"
from sklearn.metrics import mean_absolute_error, mean_squared_error
def MSE(predicted, y):
'''
Compute MSE (scalar) as in the formula above
:param predicted: np.array of predicted values, [n]
:param y: np.array of targets, [n]
'''
mae = <YOUR CODE>
return mae # scalar
def MAE(predicted, y):
'''
Compute MAE (scalar) as in the formula above
:param predicted: np.array of predicted values, [n]
:param y: np.array of targets, [n]
'''
mse = <YOUR CODE>
return mse # scalar
# + colab={} colab_type="code" id="RsLlMeDJ-WrA"
assert np.allclose(MSE(w * x + b, y), 1.084, rtol=0.1, atol=0.1), "We're sorry for your MSE"
assert np.allclose(MAE(w * x + b, y), 0.806, rtol=0.1, atol=0.1), "We're sorry for your MAE"
# + [markdown] colab_type="text" id="ESm-vn-V-WrD"
# Добавим в наш датасет выброс:
# + colab={} colab_type="code" id="x9rgKRea-WrE"
y[5] = 10
y[7] = -1
# + colab={} colab_type="code" id="TRtInBBB-WrK" outputId="b4ffa40d-1d6a-4243-f673-9c4e8f9ba1fa"
print("MSE =", MSE(w * x + b, y))
print("MAE =", MAE(w * x + b, y))
# + [markdown] colab_type="text" id="2AHFCSjx-WrP"
# На какую из метрик выброс повлиял больше? Почему?
#
# ## Обучение линейной регрессии (одномерный случай)
# Чаще всего для обучения используется функция MSE. В этом случае получаем следующую задачу оптимизации.
#
# $$\frac1n\sum\limits_{i=1}^n(w \cdot x_i + b - y_i)^2\to\min_{w,b}$$
#
# ### Метод наименьших квадратов
# Но подождите? А нельзя ли получить точное решение этой задачи?
#
# Возьмём производную функции, которую оптимизируем, по $b$:
# $$\frac1n\sum\limits_{i=1}^n(w \cdot x_i + b - y_i)$$
# Приравняем к нулю:
# $$\frac1n\sum\limits_{i=1}^n(w \cdot x_i + b - y_i)=0$$
# $$b = \overline{y}-w\cdot\overline{x}$$
#
# Аналогичными рассуждениями вычислите формулу для $b$. И допишите код, который вычисляет значения $w, b$.
# + colab={} colab_type="code" id="LX6nrBRt-WrQ"
def lin_reg_weight(x, y):
w = <YOUR CODE>
return w
def lin_reg_bias(x, y):
return y.mean() - lin_reg_weight(x, y) * x.mean()
# + colab={} colab_type="code" id="FUd7gjTy-WrU"
y = np.sin(x) + x
w = lin_reg_weight(x, y)
b = lin_reg_bias(x, y)
# + colab={} colab_type="code" id="ohO0SpjL-WrY" outputId="4257779d-96dd-456e-a34e-9420e55af3e4"
plt.scatter(x, y)
plt.plot(x, w * x + b, color='red')
plt.xlabel("x")
plt.ylabel("y")
# + colab={} colab_type="code" id="gYbqV19V-Wrd"
assert np.allclose(MSE(w * x + b, y), 0.434, rtol=0.1, atol=0.1), "something's wrong with your least squares"
# + [markdown] colab_type="text" id="Hir5Nl8w-Wrh"
# А где же, собственно, машинное обучение?
#
# ## Обучение линейной регрессии (многомерный случай)
# В общем случае (когда размерность пространства объектов не обязательно один) формула для линейной регрессии выглядит как суммирование признаков с некоторыми коэффициентами:
# $$a(x) = b + \sum\limits_{i=1}^n w_i\cdot x_i$$
# * Параметры $w$ называются веса (weights) или коэффициенты.
# * Параметр $b$ называется сдвигом (bias) или свободным коэффициентом.
# Заметим, что сумма в формуле это скалярное произведение двух векторов: $w$ и $x$. Перепишем формулу.
# $$a(x) = b + \langle w, x\rangle$$
# Как можно упростить запись, чтобы избавиться от слагаемого $b$?
#
# Итак, будем решать задачу оптимизации в общем виде.
# $$\frac1n\sum\limits_{i=1}^n(\langle w, x_i\rangle - y_i)^2\to\min\limits_w$$
# + [markdown] colab_type="text" id="2_8RoWMD-Wri"
# Пусть у нас есть $m$ объектов, у каждого объекта $n$ признаков. Тогда можно записать задачу в матричном виде:
# $$\frac1n||X\cdot w-y||^2\to\min\limits_w$$
# * $X$ — объектов матрица $[m, n]$
# * $w$ — вектор весов $[n, 1]$
# * $y$ — вектор ответов $[m, 1]$
# * $||V||^2$ — сумма квадратов всех элементов вектора
# + [markdown] colab_type="text" id="kccvqrQI-Wrj"
# Для этой задачи так же можно получить точное решение, но оно имеет несколько проблем:
# * Сложность алгоритма
# * Решение не всегда можетсуществовать
# * Если поменять функцию ошибок, аналитическое решение может даже не найтись
# + [markdown] colab_type="text" id="sppk6wAM-Wrk"
# ### Градиентный спуск
# Напомним, что градиент функции это вектор всех её частных производных.
#
# Для функции $$f(x_1, x_2, \ldots, x_n)$$ градиентом будет $$\nabla f(x_1, x_2, \ldots, x_n)=\left(\frac{\partial f}{\partial x_1}, \frac{\partial f}{\partial x_2}, \ldots, \frac{\partial f}{\partial x_n}\right)$$
# Также, градиент указывает на направление роста функции, а антиградиент — на направление убывания. Этими свойствами мы и будем пользоваться.
#
# Идея градиентного спуска довольно проста:
# 1. Стартуем в какой-то точке $n$-мерного пространства.
# 1. Считаем функцию ошибок.
# 1. Считаем градиенты от функции ошибок по каждому весу.
# 1. Сдвигаем каждый из весов на маленький шаг в сторону антиградиента.
# 1. Продолжаем, пока градиент не станет близок к нулю.
#
# Запишем более формально.
#
# Пусть
#
# $lr$ (от learning rate) — размер шага, на котором мы сдвигаем градиент на каждой итерации.
#
# $w$ — вектор весов, изначальнонулевой, либо из какого-то распределения.
#
# $L(w) = \frac1n\sum\limits_{i=1}^n(\langle w, x_i\rangle - y_i)^2$
#
# Тогда каждый шаг выглядит так: $$w_{new}:=w-lr\cdot\nabla L(w)$$
#
# Теперь запишем градиенты для линейной регрессии. Давайте найдём производную для одного веса $w_j$:
#
# $$L(w)=\frac1n\sum\limits_{i=1}^n(\langle w, x_i\rangle - y_i)^2$$
# $$\frac{\partial L(w)}{\partial w_j}=\frac2n\sum\limits_{i=1}^nx_i^j\cdot(\langle w, x_i\rangle - y_i)$$
# В матричном виде это будет выглядеть так:
# $$\nabla L(w)=\frac2nX^T\cdot(X\cdot w-y)$$
#
# **Важное замечание:** обратите внимание, что когда мы считаем градиенты, мы умножаем исходные данные. Если их не отнормировать перед этим, коэффициенты могут быстро стать очень большими.
#
# Теперь мы можем написать свою реализацию линейной регресси. **homework**
# -
Image()
# + colab={} colab_type="code" id="af3bLQ_x-Wrl"
from sklearn.base import BaseEstimator, RegressorMixin
class MyLinearRegression(BaseEstimator, RegressorMixin):
def __init__(self, features_size, gradient_descending, max_iter=1000, l=0, lr=0.001):
self.lr = lr
self.l = l
self.max_iter = max_iter
self.w = np.random.normal(size=(features_size + 1, 1))
self.gradient_descending = gradient_descending
def fit(self, X, y):
""" This should fit classifier. All the "work" should be done here
Recompute weights on each step
"""
X = np.concatenate([X, np.ones((X.shape[0], 1))], axis=1)
for step in range(self.max_iter):
g = self.gradient_descending(self.w, X, y, self.l)
if np.isnan(g[:,0]).any(): # check if gradients is nan something went wrong
raise RuntimeError("Your gradient is nan")
prev_w = np.copy(self.w)
# пересчитайте градиенты
self.w = <YOUR CODE>
if ((prev_w - self.w) ** 2).sum() < 1e-10:
break
return self
def predict(self, X, y=None):
""" Predict target class for X """
# единичную фичу для сдвига
X = np.concatenate([X, np.ones((X.shape[0], 1))], axis=1)
# посчитайте результат
return <YOUR CODE>
# + colab={} colab_type="code" id="dlbagiI1-Wrp"
def gradient_descending(w, X, y, l):
# посчитайте градиент функции ошибок по весам. Переменная l=0, не используйте её.
g = <YOUR CODE>
return g
# + [markdown] colab_type="text" id="Dc87H74I-Wrt"
# Добавим константный признак для всех объектов х. Он будет характеризовать сдвиг.
# + colab={} colab_type="code" id="HrwYJ4u9-Wrv"
from sklearn.preprocessing import scale
x = np.arange(0, 10, 0.5).reshape(20, 1)
y = np.sin(x) + x
y = y.reshape(20, 1)
x = scale(x, axis=0, with_mean=True, with_std=True) # нормализуем данные так, чтобы среднее было 0, дисперсия 1
y = scale(y, axis=0, with_mean=True, with_std=True)
# + colab={} colab_type="code" id="xB6c2vPK-Wry"
r = MyLinearRegression(1, gradient_descending, lr=1e-2).fit(x, y)
# + colab={} colab_type="code" id="fiCL8mjT-Wr1" outputId="09cdbda9-aa88-4d02-c84b-4d19307bec4f"
plt.scatter(x, y)
plt.plot(x, r.predict(x), color='red')
plt.xlabel("x")
plt.ylabel("y")
# + colab={} colab_type="code" id="eXEHrle6-Wr5"
assert np.allclose(MSE(r.predict(x), y), 0.05, rtol=0.1, atol=0.1), "something's wrong with your linear regression"
# + [markdown] colab_type="text" id="wIV54Tfr-Wr9"
# #### Fine, let's try more data!
# + colab={} colab_type="code" id="VdO_Jv8s-WsB"
N = 10000
x = np.arange(0, N, 0.5).reshape(2 * N, 1)
y = np.sqrt(x[:,0]) + 2 * (x[:,0] - 100) * (x[:,0] - 500)
y = y.reshape(2 * N, 1)
x = scale(x, axis=0, with_mean=True, with_std=True)
y = scale(y, axis=0, with_mean=True, with_std=True)
# + colab={} colab_type="code" id="4j8zMRWx-WsG" outputId="b3db14a3-e974-476a-89a3-b3dda1414f12"
r = MyLinearRegression(1, gradient_descending, lr=1e-1).fit(x, y)
# + [markdown] colab_type="text" id="uJxstHRR-WsJ"
# В чём же дело? Проблема в том, что веса быстро стали слишком большими. А, как известно, $\infty-\infty=\mathrm{nan}$.
#
# Давайте, также, будем штрафовать модель за то что веса стали слишком большие.
#
# Теперь функция ошибок будет не $\frac1n||X\cdot w-y||^2$, а $$\frac1n||X\cdot w-y||^2+\frac\lambda2\cdot ||w||^2$$
# $\lambda$ называют коэффициент регуляризации. И действительно, если формула будет такая, то как только веса станут слишком большими, модели придётся их уменьшить, ведь это уменьшит функцию ошибок. Такая регуляризация называется $L_2$-regularization. **homework**
# + colab={} colab_type="code" id="UNubzU5s-WsM"
def gradient_descending_with_l2(w, X, y, l):
# посчитайте градиент функции ошибок по весам. Используйте регуляризацию.
# Коэффициент регуляризации l
g = <YOUR CODE>
return g
# + colab={} colab_type="code" id="YIRJ7P9X-WsP"
r = MyLinearRegression(1, gradient_descending_with_l2, l=100, lr=1e-5).fit(x, y)
# + colab={} colab_type="code" id="sREhSmtd-WsS" outputId="f63314ad-30c5-434c-a91d-4b3ec3725cda"
plt.scatter(x[:,0], y)
plt.plot(x[:,0], r.predict(x), color='red')
plt.xlabel("x")
plt.ylabel("y")
# + colab={} colab_type="code" id="Mq_pyYWs-WsX" outputId="14beba15-66ef-4185-c387-93964a12aa0f"
print("Your MSE is: ", MSE(r.predict(x), y))
# + [markdown] colab_type="text" id="d6R4MdQ9-Wsb"
# Если данных слишком много, то посчитать градиент по всем объектам не получается. Для этого есть стохаистический градиентный спуск. В нём случайно выбирается один объект и градиенты считаются только для него. Реализуйте это. **homework**
# + colab={} colab_type="code" id="1xEVcrzL-Wsc"
def stokhaistik_gradient_descending_with_l2(w, X, y, l):
# посчитайте градиент функции ошибок для случайного объекта по весам. Используйте регуляризацию.
# Коэффициент регуляризации l=0
g = <YOUR CODE>
return g
# + colab={} colab_type="code" id="y7365uiK-Wsg"
r = MyLinearRegression(1, stokhaistik_gradient_descending_with_l2, l=0.1, lr=1e-3).fit(x, y)
# + colab={} colab_type="code" id="VN3D5TUk-Wsi" outputId="f0468e07-51b5-4c0a-b5d6-b91ab3a62a0d"
plt.scatter(x[:,0], y)
plt.plot(x[:,0], r.predict(x), color='red')
plt.xlabel("x")
plt.ylabel("y")
# + colab={} colab_type="code" id="KFN0rH-b-Wsm" outputId="65b6c948-e71f-4a66-e034-899fe4533f4c"
print("Your MSE is: ", MSE(r.predict(x), y))
# + [markdown] colab_type="text" id="k1j6h3bu-Wsq"
# ### Полиномиальные фичи.
# К сожалению, наш мир нелинейный, поэтому чтобы линейная регрессия в чистом виде часто работает не очень хорошо. Однако, можно добавить новые фичи, чтобы исправить это. Рассмотрим пример.
# + colab={} colab_type="code" id="JSKO44uR-Wsu"
from sklearn.linear_model import LinearRegression
# + colab={} colab_type="code" id="tr5_nndQ-Wsx"
x = np.linspace(0, 1, 100)
y = np.cos(1.5 * np.pi * x)
x_train = np.random.uniform(0, 1, size=30)
y_train = np.cos(1.5 * np.pi * x_train) + np.random.normal(scale=0.1, size=x_train.shape)
# + [markdown] colab_type="text" id="hMMugDkc-Ws0"
# Обучим линейную регрессию со стандартными фичами, со всеми степенями до 4, со всеми степенями до 20.
# + colab={"base_uri": "https://localhost:8080/", "height": 301} colab_type="code" id="XVg-S9f9-Ws0" outputId="b5498392-cdca-4ad8-a623-c330536b05f9"
from sklearn.preprocessing import PolynomialFeatures
fig, axs = plt.subplots(figsize=(16, 4), ncols=3)
for i, degree in enumerate([1, 4, 20]):
X_train = PolynomialFeatures(degree).fit_transform(x_train[:, None])
X = PolynomialFeatures(degree).fit_transform(x[:, None])
regr = LinearRegression().fit(X_train, y_train)
y_pred = regr.predict(X)
axs[i].plot(x, y, label="Real function")
axs[i].scatter(x_train, y_train, label="Data")
axs[i].plot(x, y_pred, label="Prediction")
if i == 0:
axs[i].legend()
axs[i].set_title("Degree = %d" % degree)
axs[i].set_xlabel("$x$")
axs[i].set_ylabel("$f(x)$")
axs[i].set_ylim(-2, 2)
# + [markdown] colab_type="text" id="ynPwbC3S-Ws3"
# Почему первая модель не доучилась, а последняя переобучилась?
#
# --
#
# Оказывается, что линейная регрессия — это очень мощный инструмент, если мы можем подбирать коэффициенты степенного ряда. Таким образом мы можем приближать любую функцию, не зная её вид, а подбирая коэффициенты в ряде Тейлора.
# + [markdown] colab_type="text" id="99foWvvu7Iq0"
# ## Линейная классификация
#
# Давайте приступим к рассмотрению задачи классификации. Начнём с того, что вспомним её постановку.
#
# Пусть задана обучающая выборка $X = \left\{ \left( x_i, y_i \right) \right\}_{i=1}^l, x_i \in \mathbb{X}, y_i \in \mathbb{Y},$ —
# * $l$ пар объект-ответ, где
# * $\mathbb{X}$ — пространство объектов,
# * $\mathbb{Y}$ — пространство ответов.
#
# Задача классификации отличается о задачи восстановления регрессии тем, что в данном случае пространство ответов конечно:
#
#
#
# В случае классификации формула для ответа на объекте $x$ видоизменяется по сравнению с линейной регрессией (при этом мы предполагаем, что нулевой признак в выборке — константный):
# $$a(x) = sign(\langle w, x\rangle)$$
#
# Множество точек $\langle w, x\rangle = 0$ является гиперплоскостью и делит пространство на 2 части по разные стороны от этой самой гиперплоскости. При этом объекты, расположенные по разные стороны от неё, полученный классификатор относит к различным классам в зависимости от знака скалярного произведения.
#
# 
#
#
# Но получившийся функционал $a(x)$ не получается оптимизировать градиентными методами. Тем не менее, у нас есть функция, которая переводит объект в $[-\infty, +\infty]$. Давайте научимся получать из этого вероятность $P\in[0,1]$. Это число будет означать, с какой вероятностью объект принадлежит к классу $1$?
#
# В качестве такой функции будем использовать [сигмоиду](https://ru.wikipedia.org/wiki/%D0%A1%D0%B8%D0%B3%D0%BC%D0%BE%D0%B8%D0%B4%D0%B0) (иногда называется логистическая функция): $$\sigma(x)=\frac{1}{1-e^{x}}$$
# 
#
# Таким образом, у нас получилась следующая функция:
#
# $$\tilde L(x) = \frac1{1-e^{\langle w, x\rangle}}$$
#
# А в качестве функции потерь возьмём логарифм функции правдоподобия, для двух классов она будет выглядеть так:
# $$ \dfrac{1}{N}\sum_{i=1}^N \log(1 + \exp(-\langle w, x_i \rangle y_i)) \to \min_w$$
#
# Точно так же, в логистической регрессии нужно использовать регуляризацию. Задачу обучения регуляризованной логистической регрессии можно записать следующим образом:
# $$ \dfrac{1}{N}\sum_{i=1}^N \log(1 + \exp(-\langle w, x_i \rangle y_i)) + \dfrac{\lambda}{2}\lVert w \rVert^2 \to \min_w$$
#
# ### Обучение Логистической Регрессии
#
#
#
# + colab={} colab_type="code" id="b7LqHVIR-Ws5"
from sklearn.datasets import make_classification, make_blobs
from sklearn.linear_model import LogisticRegression
# + colab={} colab_type="code" id="Pd4Ls-hePyiT"
X, y = make_blobs(n_samples=300, centers=2)
y = 2 * y - 1
# + colab={"base_uri": "https://localhost:8080/", "height": 594} colab_type="code" id="orE8ebj1Pmqs" outputId="5a5cdaa0-4a12-4085-cf27-93137a45cd83"
ind = y == -1
plt.figure(figsize=(20,10))
plt.scatter(X[ind,0], X[ind,1], c='red')
plt.scatter(X[~ind,0], X[~ind,1], c='blue')
plt.legend(['y = -1', 'y = 1'])
plt.show()
# + colab={} colab_type="code" id="yKG9TNAkPt4j"
lr = LogisticRegression().fit(X, y)
w_1 = lr.coef_[0][0]
w_2 = lr.coef_[0][1]
w_0 = lr.intercept_[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 594} colab_type="code" id="_YxCumE7RUgH" outputId="953eea08-5b4c-4875-dfc9-e8ac05c5f6f8"
ind = y == -1
plt.figure(figsize=(20,10))
plt.scatter(X[ind,0], X[ind,1], c='red')
plt.scatter(X[~ind,0], X[~ind,1], c='blue')
x_arr = np.linspace(-10, 5, 1000)
plt.plot(x_arr, -(w_0 + w_1 * x_arr) / w_2)
plt.legend(['y = -1', 'y = 1'])
plt.show()
# -
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
import warnings
warnings.simplefilter("ignore")
from IPython.display import Image
# %matplotlib inline
# # Семинар №6. Линейные модели и Градиентный спуск.
# # Постановка задачи
# Пререквизиты:
# * $n$ - количество объектов в выборке
# * $m$ - количество признаков у объекта
# * все вектора - **столбцы**
# * $I_{a\times b}$ - единичная матрица
# * $O_{a\times b}$ - нулевая матрица
# * $E_{a\times b}$ - матрица из всех единиц
# + [markdown] colab={} colab_type="code" id="Exd-v0aSRhj6"
# Давайте вспомним, что такое задача линейной регрессии в рамках машинного обучения.
#
# У нас есть обучающая выборка $(X, y)$, c $n$ объектами и $m$ признаками. (Значит размер $X$ - $n\times m$)
#
# Алгоритм решения выбирается из простнанства линейных функций:
#
# $a(x^1) = w_0 + w_1x_1^1 + \ldots + w_mx_m^1$
#
# -
# ## Точное решение
# Нам нужно найти такое $a(\cdot)$, чтобы решилась система уравнений
# $$\begin{cases}
# a(x^1) = w_0 + w_1x_1^1 + \ldots + w_mx_m^1 = y^1\\
# \ldots \\
# a(x^i) = w_0 + w_1x_1^i + \ldots + w_mx_m^i = y^i\\
# \ldots \\
# a(x^n) = w_0 + w_1x_1^n + \ldots + w_mx_m^n = y^n
# \end{cases}$$
# Давайте запишем в матричном виде, предварительно добавив к $X$ единичный столбец слева. $X = (E_{n\times 1}|X)$. Получим:
#
# $$Xw = y$$
# С курса линейной алгебры мы помним, что система линейных уравнений имеет единственное решение, когда $X$ - квадратная, максимального ранга (нету линейно зависимых строк\столбцов). По сути, если у $X$ существует обратная, тогда:
#
# $$w = X^{-1}y$$
# Такое решение означает, что через $m = n$ точек прямой (из которых ни одна не лежит на плоскости) можно провести ровно одну плоскость.
Image('плоскость.png')
# Однако никто нам не гарантирует, что матрица $X$ будет удовлетворять нужным свойствам.
# ## MSE
# Без ограничений общности можем считать, что в $X$ **отсутствуют** линейно-зависимые строки\столбцы (будем считать что их заранее убрали).
#
# Необязательная сноска:
# * Более того, в практическом смысле у нас действительно не может быть линейно-зависимых строк, потому что объекты у нас берутся из некоторого пространства $\mathbb{X}$. Если оно непрерывно (хотя бы один признак - непрерывен), то вероятность попасть в одну точку пространства равна 0.
#
# * К тому же, к реально наблюдаемым данных добавляется нормальный шум, который представляет собой независимую случайную величину - значит все объекты независимы.
# Пусть $n > m$. Тогда мы не можем провести через $n$ точек в пространстве размерности $m$ гиперплоскость.
Image('плоскость2.png', width=500)
# В этом случае давайте проводить плоскость такую, что расстояния от точек до этой плоскости будет минимальным.
Image('расстояние.jpg')
# Тут у нас есть вариант взять метрику эвклидову (квадраты расстояний $MSE$) или манхетонову (модуль расстояний $MAE$). Берем $MSE$, потому что ее можно легко дифференцировать (да, это действительно единственное объяснение)
# У тут мы можем вспомнить, что мы уже видели раньше: функционал ошибки
# $$Q(a(x), y) = MSE = \frac{1}{n}\sum_{i=1}^{n} (a(x^i)-y^i)^2 \rightarrow min$$
# Который можно записать в более аккуратном матричном виде:
# $$L(w) = \lVert Xw - Y\rVert_2^2 \rightarrow min$$
#
# $\lVert x \rVert_2^2 = \sum_{i=1}^{n}x_1^2 + \ldots + x_n^2$ -квадрат нормы эвклидовой метрики
# Теперь чтобы найти $w$ - применим единственное возможно действие, когда мы видим минимизацию линейной функции - возьмем производную по аргументу вектору или (тоже самое) возьмем частные производные каждого $w_i$
# $$\nabla_{w}L = \begin{cases}
# \frac{\partial L}{\partial w_1} = \frac{1}{n}\sum_{i=1}^{n} 2x_1^{i}(a(x^i)-y^i)\\
# \ldots \\
# \frac{\partial L}{\partial w_j} = \frac{1}{n}\sum_{i=1}^{n} 2x_j^{i}(a(x^i)-y^i)\\
# \ldots \\
# \frac{\partial L}{\partial w_j} = \frac{1}{n}\sum_{i=1}^{n} 2x_m^{i}(a(x^i)-y^i)
# \end{cases}$$
# $\nabla_{w}L$ - называется **градиентом** линейной функции $L$, по парамеру вектору $w$
# Если на эту систему внимательно посмотреть (или изучить матричное дифференцирование), можно понять, что данная систему можно записать тоже в матричном виде:
#
# $$ \nabla_{w}L = 2X^{T}(Xw - Y)$$
# Осталось совершить последнее стандартное действие - приравнять производную к нулю вектору и решить матричное уравнение.
# $$ 0 = 2X^{T}(Xw - Y)$$
#
# $$ X^{T}Xw = X^{T}Y$$
#
# $$ w = (X^{T}X)^{-1}X^{T}Y$$
| 06-linear/.ipynb_checkpoints/linear-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from scipy.stats import shapiro,normaltest,anderson,boxcox,skew
import smogn
data = pd.read_csv("fepB_complete.csv")
data.describe()
data.info()
def target_var(data):
y = data[['R1','R2','R3','R4','R5','R6','R7','R8']].copy()
flu_level = [0.5,1.5,2.5,3.5,4.5,5.5,6.5,7.5]
y['total'] = y.sum(axis=1)
y = y[['R1','R2','R3','R4','R5','R6','R7','R8']].div(y.total,axis=0)
y *= flu_level
y['weighted_mean'] = y.sum(axis=1)
return y['weighted_mean']
y = target_var(data)
df = data[['SeqID','dG_pairing','dG_folding']].copy()
df['weighted'] = y
# ### Original
ax = sns.histplot(data=y,bins=20,stat='density',kde=True,edgecolor='white')
ax.get_lines()[0].set_color('black')
print(np.round(skew(y),4))
# ### Log Transformation
lg_target = np.log1p(y)
ax = sns.histplot(data=lg_target,bins=20,stat='density',kde=True,edgecolor='white')
ax.get_lines()[0].set_color('black')
print(np.round(skew(lg_target),4))
# ### Square Root Transformation
sqrrt_target = y**(1/2)
ax = sns.histplot(data=sqrrt_target,bins=20,stat='density',kde=True,edgecolor='white')
ax.get_lines()[0].set_color('black')
print(np.round(skew(sqrrt_target),4))
# ### Boxcox Transformation
bcx_target,lam = boxcox(y)
ax = sns.histplot(data=bcx_target,bins=20,stat='density',kde=True,edgecolor='white')
ax.get_lines()[0].set_color('black')
print(np.round(skew(bcx_target),4))
| fepb/fepb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Market Response Functions
#
# This notebooks provides simple illustrative examples of how individual customer price response aggregate into the total market response function.
# +
import numpy as np
import pandas as pd
from matplotlib import pylab as plt
plt.style.use('seaborn-white')
import seaborn as sns
pd.options.mode.chained_assignment = None
plt.rcParams.update({'pdf.fonttype': 'truetype'})
import platform
print(f'Python {platform.python_version()}')
# -
# # Market Response Function for Yes-No Customer Choice
#
# We first consider the case of yes-no customer choice. Each customer has a maximum price she is willing to pay for a certain product or service, and she buys exactly one unit when the offered price is below her willingness to pay. This is typcial for services and durable goods.
#
# We generate uniformly distributed willingness-to-pay points for a small population of customers, and aggregate them into the overall market response that approaches the linear price-demand function as the number of customers grows.
# +
n = 100
n_customers = 16
#
# Generate response functions
#
willingness_to_pay = np.random.randint(0, n, size=n_customers)
y = np.zeros((n_customers, n))
for i, wtp in enumerate(willingness_to_pay):
y[i, :] = np.hstack([np.ones(wtp), np.zeros(n - wtp)])
market_response = np.sum(y, axis=0)
#
# Visualization
#
x = np.linspace(1, n, n)
fig, ax = plt.subplots(len(willingness_to_pay) + 1, 1, figsize=(8, 16), gridspec_kw={'height_ratios': [1]*n_customers + [10]})
for i, wtp in enumerate(willingness_to_pay):
ax[i].plot(x, y[i])
ax[i].tick_params(axis='x', which='both', bottom=True, top=True, labelbottom=False)
ax[i].set_yticks([])
ax[i].set_ylim([-0.5, 1.5])
ax[i].grid(True)
ax[-1].plot(x, market_response)
ax[-1].grid(True)
ax[-1].set_xlabel('Price')
plt.show()
# -
# # Market Response Function for Variable Quantity Choice
#
# Second, we repeat the simulation with variable-quantity responses. Each customer has a randomly parametrized linear response function: she buys more units at lower prices, and fewer units at higher prices.
# +
n = 100
n_customers = 8
max_units = 10
#
# Generate response functions
#
max_units_customer = np.random.randint(0, max_units, size=n_customers)
willingness_to_pay = np.random.randint(0, n, size=n_customers)
y = np.zeros((n_customers, n))
for i in range(n_customers):
y[i, :] = np.hstack([np.linspace(max_units_customer[i], 0, willingness_to_pay[i]), np.zeros(n - willingness_to_pay[i])])
market_response = np.sum(y, axis=0)
#
# Visualization
#
x = np.linspace(1, n, n)
fig, ax = plt.subplots(n_customers + 1, 1, figsize=(8, 16), gridspec_kw={'height_ratios': [1]*n_customers + [10]})
for i in range(n_customers):
ax[i].plot(x, y[i])
ax[i].tick_params(axis='x', which='both', bottom=True, top=True, labelbottom=False)
ax[i].set_ylim([-0.5, max_units])
ax[i].set_yticks(np.arange(0, max_units, 2), minor=True)
ax[i].grid(which='both')
ax[-1].plot(x, market_response)
ax[-1].grid(True)
ax[-1].set_xlabel('Price')
plt.show()
# -
# # Standard Market-Response Functions
#
# In this section, we plot several standard market response functions and the corresponding price elasticities computed emperically.
# +
fig, ax = plt.subplots(1, 2, figsize=(10, 5))
#
# Calculate the empirical price elasticity of demand
#
def empirical_elasticity(p, q):
e = np.zeros(len(q))
for i in range(1, len(q)):
e[i] = -(q[i] - q[i-1])/(p[i] - p[i-1]) * (p[i] / q[i])
return e
#
# Linear function
#
a, b = 10, 0.99
demand_function = lambda p: a - b*p
p = np.linspace(0, 10, 100)
ax[0].plot(p, demand_function(p), label='demand')
ax[0].plot(p, empirical_elasticity(p, demand_function(p)), label='elasticity')
ax[0].grid(True)
ax[0].set_xlabel('p')
ax[0].set_ylabel('q')
ax[0].legend()
ax[0].set_ylim([-1, a+1])
#
# Constant-elasticity function
#
epsilon = -0.25
demand_function = lambda p: 5*np.power(p, epsilon)
p = np.linspace(0, 10, 100)
ax[1].plot(p, demand_function(p), label='demand')
ax[1].plot(p, empirical_elasticity(p, demand_function(p)), label='elasticity')
ax[1].grid(True)
ax[1].set_xlabel('p')
ax[1].set_ylabel('q')
ax[1].legend()
plt.savefig('price-response.pdf')
plt.show()
# -
| pricing/market-response-functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import seaborn as sns
titanic = sns.load_dataset('titanic')
titanic.head(10)
X = titanic.drop('survived',axis = 1)
Y = titanic['survived']
X.shape
Y.shape
| Chapter01/Activity1.01/Activity1_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Exploration 2
#
# ### Imports and getting set up
# +
import pandas as pd
import numpy as np
import scipy.stats as st
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
# -
# ### DB Connection & Call
confile = list(pd.read_csv('../../dbcon.csv'))
postgres_db = 'studentsperformance'
db_connection = 'postgresql://{}:{}@{}:{}/{}'.format(confile[0], confile[1], confile[2], confile[3], postgres_db)
# +
query = '''
SELECT *
FROM studentsperformance
;'''
students_df = pd.read_sql(query, db_connection)
# -
display(
students_df.head(),
students_df.info(),
students_df.isna().mean()
)
# #### 1. Are there any differences between the genders, ethnicities, and parental level of education with respect to their performances in exams?
students_df['mathbins'] = pd.cut(students_df['math score'], bins=[0, 60, 70, 80, 90, 101], labels=['F', 'D', 'C', 'B', 'A'], right=False)
students_df['readbins'] = pd.cut(students_df['reading score'], bins=[0, 60, 70, 80, 90, 101], labels=['F', 'D', 'C', 'B', 'A'], right=False)
students_df['writbins'] = pd.cut(students_df['writing score'], bins=[0, 60, 70, 80, 90, 101], labels=['F', 'D', 'C', 'B', 'A'], right=False)
features = ['gender', 'race/ethnicity', 'parental level of education', 'lunch', 'test preparation course']
depvars = ['mathbins', 'readbins', 'writbins']
for feat in features:
featdf = pd.DataFrame()
for var in depvars:
studentct = pd.crosstab(students_df[var], students_df[feat])
display(studentct)
# <span style="color:blue">There are significant differences in here most of them with respect to the performance of males vs females in computational vs non-computational areas</span>
# #### 2. Are there any differences between the lunch types with respect to their performances in exams? If there are, how do you explain this?
# <span style="color:blue">There is a large difference. The median exam scores for the free/reduced lunch bracked is an F, while the median exam score for standard lunch students is around a C. This can be explained by the economic hardship between the families of students that would require assistance for lunch vs not. Families that would require assistance for lunch will typically be poorer and the increased hardship on the child may prevent them from performing well.</span>
# #### 3. Does the test preparation course seem to have an effect on the exam performances?
# <span style="color:blue">The distribution is about the same as the distribution for the lunch where the prep students have a median score of a C while the other group has a median score of an F. I would put that less as something to do with the course since whether or not the student takes a prep course is also affected by the same familial factors above</span>
# #### 4. Which 2 exam scores are most correlated with each other?
# <span style="color:blue">As seen and stated in the data above, the reading/writing are the most correlated with one another being the two non-computational exams, while math is the only computational subject here with data.</span>
| Assignment_14-6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# ## Data-X: Titanic Survival Analysis
#
# Data from: https://www.kaggle.com/c/titanic/data
# **Authors:** Several public Kaggle Kernels, edits by <NAME> & <NAME>
#
# <img src="data/Titanic_Variable.png">
# # Note
#
# Install xgboost package in your pyhton enviroment:
#
# try:
# ```
# $ conda install py-xgboost
# ```
#
# +
'''
# You can also install the package by running the line below
# directly in your notebook
''';
# #!conda install py-xgboost --y
# -
# ## Import packages
# +
# No warnings
import warnings
warnings.filterwarnings('ignore') # Filter out warnings
# data analysis and wrangling
import pandas as pd
import numpy as np
import random as rnd
# visualization
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# machine learning
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB # Gaussian Naive Bayes
from sklearn.linear_model import Perceptron
from sklearn.linear_model import SGDClassifier #stochastic gradient descent
from sklearn.tree import DecisionTreeClassifier
import xgboost as xgb
# Plot styling
sns.set(style='white', context='notebook', palette='deep')
plt.rcParams[ 'figure.figsize' ] = 9 , 5
# -
# ### Define fancy plot to look at distributions
# Special distribution plot (will be used later)
def plot_distribution( df , var , target , **kwargs ):
row = kwargs.get( 'row' , None )
col = kwargs.get( 'col' , None )
facet = sns.FacetGrid( df , hue=target , aspect=4 , row = row , col = col )
facet.map( sns.kdeplot , var , shade= True )
facet.set( xlim=( 0 , df[ var ].max() ) )
facet.add_legend()
plt.tight_layout()
# ## References to material we won't cover in detail:
#
# * **Gradient Boosting:** http://blog.kaggle.com/2017/01/23/a-kaggle-master-explains-gradient-boosting/
#
# * **Naive Bayes:** http://scikit-learn.org/stable/modules/naive_bayes.html
#
# * **Perceptron:** http://aass.oru.se/~lilien/ml/seminars/2007_02_01b-Janecek-Perceptron.pdf
# ## Input Data
# +
train_df = pd.read_csv('data/train.csv')
test_df = pd.read_csv('data/test.csv')
combine = [train_df, test_df]
# NOTE! When we change train_df or test_df the objects in combine
# will also change
# (combine is only a pointer to the objects)
# combine is used to ensure whatever preprocessing is done
# on training data is also done on test data
# -
# # Exploratory Data Anlysis (EDA)
# We will analyze the data to see how we can work with it and what makes sense.
train_df
print(train_df.columns)
# preview the data
train_df.head(10)
# General data statistics
train_df.describe()
# Data Frame information (null, data type etc)
train_df.info()
# ### Comment on the Data
# <div class='alert alert-info'>
# `PassengerId` is a random number and thus does not contain any valuable information. `Survived, Passenger Class, Age Siblings Spouses, Parents Children` and `Fare` are numerical values -- so we don't need to transform them, but we might want to group them (i.e. create categorical variables). `Sex, Embarked` are categorical features that we need to map to integer values. `Name, Ticket` and `Cabin` might also contain valuable information.
# </div>
# # Preprocessing Data
# check dimensions of the train and test datasets
print("Shapes Before: (train) (test) = ", \
train_df.shape, test_df.shape)
# +
# Drop columns 'Ticket', 'Cabin', need to do it for both test
# and training
train_df = train_df.drop(['Ticket', 'Cabin'], axis=1)
test_df = test_df.drop(['Ticket', 'Cabin'], axis=1)
combine = [train_df, test_df]
print("Shapes After: (train) (test) =", train_df.shape, test_df.shape)
# +
# Check if there are null values in the datasets
print(train_df.isnull().sum())
print()
print(test_df.isnull().sum())
# -
# # Data Preprocessing
train_df.head(5)
# ### Hypothesis
# The Title of the person is a feature that can predict survival
# List example titles in Name column
train_df.Name[:5]
# +
# from the Name column we will extract title of each passenger
# and save that in a column in the dataset called 'Title'
# if you want to match Titles or names with any other expression
# refer to this tutorial on regex in python:
# https://www.tutorialspoint.com/python/python_reg_expressions.htm
# Create new column called title
for dataset in combine:
dataset['Title'] = dataset.Name.str.extract(' ([A-Za-z]+)\.',\
expand=False)
# +
# Double check that our titles makes sense (by comparing to sex)
pd.crosstab(train_df['Title'], train_df['Sex'])
# -
# same for test set
pd.crosstab(test_df['Title'], test_df['Sex'])
# +
# We see common titles like Miss, Mrs, Mr, Master are dominant, we will
# correct some Titles to standard forms and replace the rarest titles
# with single name 'Rare'
for dataset in combine:
dataset['Title'] = dataset['Title'].\
replace(['Lady', 'Countess','Capt', 'Col', 'Don', 'Dr',\
'Major', 'Rev', 'Sir', 'Jonkheer', 'Dona'], 'Rare')
dataset['Title'] = dataset['Title'].replace('Mlle', 'Miss') #Mademoiselle
dataset['Title'] = dataset['Title'].replace('Ms', 'Miss')
dataset['Title'] = dataset['Title'].replace('Mme', 'Mrs') #Madame
# +
# Now that we have more logical titles, and a few groups
# we can plot the survival chance for each title
train_df[['Title', 'Survived']].groupby(['Title']).mean()
# -
# We can also plot it
sns.countplot(x='Survived', hue="Title", data=train_df, order=[1,0])
plt.xticks(range(2),['Made it','Deceased']);
# +
# Title dummy mapping
for dataset in combine:
binary_encoded = pd.get_dummies(dataset.Title)
newcols = binary_encoded.columns
dataset[newcols] = binary_encoded
train_df.head()
# -
train_df = train_df.drop(['Name', 'Title', 'PassengerId'], axis=1)
test_df = test_df.drop(['Name', 'Title'], axis=1)
combine = [train_df, test_df]
train_df.shape, test_df.shape
# ## Gender column
# +
# Map Sex to binary categories
for dataset in combine:
dataset['Sex'] = dataset['Sex'] \
.map( {'female': 1, 'male': 0} ).astype(int)
train_df.head()
# -
# ### Handle missing values for age
# We will now guess values of age based on sex (male / female)
# and socioeconomic class (1st,2nd,3rd) of the passenger.
#
# The row indicates the sex, male = 0, female = 1
#
# More refined estimate than only median / mean etc.
guess_ages = np.zeros((2,3),dtype=int) #initialize
guess_ages
# +
# Fill the NA's for the Age columns
# with "qualified guesses"
for idx,dataset in enumerate(combine):
if idx==0:
print('Working on Training Data set\n')
else:
print('-'*35)
print('Working on Test Data set\n')
print('Guess values of age based on sex and pclass of the passenger...')
for i in range(0, 2):
for j in range(0,3):
guess_df = dataset[(dataset['Sex'] == i) \
&(dataset['Pclass'] == j+1)]['Age'].dropna()
# Extract the median age for this group
# (less sensitive) to outliers
age_guess = guess_df.median()
# Convert random age float to int
guess_ages[i,j] = int(age_guess)
print('Guess_Age table:\n',guess_ages)
print ('\nAssigning age values to NAN age values in the dataset...')
for i in range(0, 2):
for j in range(0, 3):
dataset.loc[ (dataset.Age.isnull()) & (dataset.Sex == i) \
& (dataset.Pclass == j+1),'Age'] = guess_ages[i,j]
dataset['Age'] = dataset['Age'].astype(int)
print()
print('Done!')
train_df.head()
# -
# Split into age bands and look at survival rates
train_df['AgeBand'] = pd.cut(train_df['Age'], 5)
train_df[['AgeBand', 'Survived']].groupby(['AgeBand'], as_index=False)\
.mean().sort_values(by='AgeBand', ascending=True)
# +
# Plot distributions of Age of passangers who survived
# or did not survive
plot_distribution( train_df , var = 'Age' , target = 'Survived' ,\
row = 'Sex' )
# +
# Change Age column to
# map Age ranges (AgeBands) to integer values of categorical type
for dataset in combine:
dataset.loc[ dataset['Age'] <= 16, 'Age'] = 0
dataset.loc[(dataset['Age'] > 16) & (dataset['Age'] <= 32), 'Age'] = 1
dataset.loc[(dataset['Age'] > 32) & (dataset['Age'] <= 48), 'Age'] = 2
dataset.loc[(dataset['Age'] > 48) & (dataset['Age'] <= 64), 'Age'] = 3
dataset.loc[ dataset['Age'] > 64, 'Age']=4
train_df.head()
# Note we could just run vvvvvvvvvvvvv
# dataset['Age'] = pd.cut(dataset['Age'], 5,labels=[0,1,2,3,4])
# -
# remove AgeBand from before
train_df = train_df.drop(['AgeBand'], axis=1)
combine = [train_df, test_df]
train_df.head()
# # Create variable for Family Size
#
# How did the number of people the person traveled with impact the chance of survival?
# +
# SibSp = Number of Sibling / Spouses
# Parch = Parents / Children
for dataset in combine:
dataset['FamilySize'] = dataset['SibSp'] + dataset['Parch'] + 1
# Survival chance with FamilySize
train_df[['FamilySize', 'Survived']].groupby(['FamilySize'], as_index=False).mean().sort_values(by='Survived', ascending=False)
# -
# Plot it, 1 is survived
sns.countplot(x='Survived', hue="FamilySize", data=train_df, order=[1,0]);
# +
# Binary variable if the person was alone or not
for dataset in combine:
dataset['IsAlone'] = 0
dataset.loc[dataset['FamilySize'] == 1, 'IsAlone'] = 1
train_df[['IsAlone', 'Survived']].groupby(['IsAlone'], as_index=False).mean()
# +
# We will only use the binary IsAlone feature for further analysis
train_df = train_df.drop(['Parch', 'SibSp', 'FamilySize'], axis=1)
test_df = test_df.drop(['Parch', 'SibSp', 'FamilySize'], axis=1)
combine = [train_df, test_df]
train_df.head()
# +
# We can also create new features based on intuitive combinations
for dataset in combine:
dataset['Age*Class'] = dataset.Age * dataset.Pclass
train_df.loc[:, ['Age*Class', 'Age', 'Pclass']].head(8)
# -
# # Port the person embarked from
# Let's see how that influences chance of survival
# +
# To replace Nan value in 'Embarked', we will use the mode
# in 'Embaraked'. This will give us the most frequent port
# the passengers embarked from
freq_port = train_df.Embarked.dropna().mode()[0]
freq_port
# +
# Fill NaN 'Embarked' Values in the datasets
for dataset in combine:
dataset['Embarked'] = dataset['Embarked'].fillna(freq_port)
train_df[['Embarked', 'Survived']].groupby(['Embarked'], as_index=False).mean().sort_values(by='Survived', ascending=False)
# -
# Let's plot it
sns.countplot(x='Survived', hue="Embarked", data=train_df, order=[1,0]);
# +
# Create categorical dummy variables for Embarked values
for dataset in combine:
binary_encoded = pd.get_dummies(dataset.Embarked)
newcols = binary_encoded.columns
dataset[newcols] = binary_encoded
train_df.head()
# -
# Drop Embarked
for dataset in combine:
dataset.drop('Embarked', axis=1, inplace=True)
# ## Handle continuous values in the Fare column
# Fill the NA values in the Fares column with the median
test_df['Fare'].fillna(test_df['Fare'].dropna().median(), inplace=True)
test_df.head()
# q cut will find ranges equal to the quantile of the data
train_df['FareBand'] = pd.qcut(train_df['Fare'], 4)
train_df[['FareBand', 'Survived']].groupby(['FareBand'], as_index=False).mean().sort_values(by='FareBand', ascending=True)
# +
for dataset in combine:
dataset.loc[ dataset['Fare'] <= 7.91, 'Fare'] = 0
dataset.loc[(dataset['Fare'] > 7.91) & \
(dataset['Fare'] <= 14.454), 'Fare'] = 1
dataset.loc[(dataset['Fare'] > 14.454) & \
(dataset['Fare'] <= 31), 'Fare'] = 2
dataset.loc[ dataset['Fare'] > 31, 'Fare'] = 3
dataset['Fare'] = dataset['Fare'].astype(int)
train_df = train_df.drop(['FareBand'], axis=1)
combine = [train_df, test_df]
# -
train_df.head
# ## Finished
train_df.head(7)
# All features are approximately on the same scale
# no need for feature engineering / normalization
test_df.head(7)
# Check correlation between features
# (uncorrelated features are generally more powerful predictors)
colormap = plt.cm.viridis
plt.figure(figsize=(12,12))
plt.title('Pearson Correlation of Features', y=1.05, size=15)
sns.heatmap(train_df.astype(float).corr().round(2)\
,linewidths=0.1,vmax=1.0, square=True, cmap=colormap, \
linecolor='white', annot=True);
# # Next Up: Machine Learning!
# Now we will Model, Predict, and Choose algorithm for conducting the classification
# Try using different classifiers to model and predict. Choose the best model from:
# * Logistic Regression
# * KNN
# * SVM
# * Naive Bayes
# * Decision Tree
# * Random Forest
# * Perceptron
# * XGBoost
# ## Setup Train and Validation Set
# +
X = train_df.drop("Survived", axis=1) # Training & Validation data
Y = train_df["Survived"] # Response / Target Variable
# Since we don't have labels for the test data
# this won't be used. It's only for Kaggle Submissions
X_submission = test_df.drop("PassengerId", axis=1).copy()
print(X.shape, Y.shape)
# +
# Split training and test set so that we test on 20% of the data
# Note that our algorithms will never have seen the validation
# data during training. This is to evaluate how good our estimators are.
np.random.seed(1337) # set random seed for reproducibility
from sklearn.model_selection import train_test_split
X_train, X_val, Y_train, Y_val = train_test_split(X, Y, test_size=0.2)
print(X_train.shape, Y_train.shape)
print(X_val.shape, Y_val.shape)
# -
# ## Scikit-Learn general ML workflow
# 1. Instantiate model object
# 2. Fit model to training data
# 3. Let the model predict output for unseen data
# 4. Compare predicitons with actual output to form accuracy measure
# # Logistic Regression
logreg = LogisticRegression() # instantiate
logreg.fit(X_train, Y_train) # fit
Y_pred = logreg.predict(X_val) # predict
acc_log = round(logreg.score(X_val, Y_val) * 100, 2) # evaluate
acc_log
# +
#testing confusion matrix
from sklearn.metrics import confusion_matrix
confusion_matrix(Y_val, Y_pred)
# +
# Support Vector Machines
svc = SVC()
svc.fit(X_train, Y_train)
Y_pred = svc.predict(X_val)
acc_svc = round(svc.score(X_val, Y_val) * 100, 2)
acc_svc
# -
knn = KNeighborsClassifier(n_neighbors = 3)
knn.fit(X_train, Y_train)
Y_pred = knn.predict(X_val)
acc_knn = round(knn.score(X_val, Y_val) * 100, 2)
acc_knn
# +
# Perceptron
perceptron = Perceptron()
perceptron.fit(X_train, Y_train)
Y_pred = perceptron.predict(X_val)
acc_perceptron = round(perceptron.score(X_val, Y_val) * 100, 2)
acc_perceptron
# +
# XGBoost
gradboost = xgb.XGBClassifier(n_estimators=1000)
gradboost.fit(X_train, Y_train)
Y_pred = gradboost.predict(X_val)
acc_perceptron = round(gradboost.score(X_val, Y_val) * 100, 2)
acc_perceptron
# +
# Random Forest
random_forest = RandomForestClassifier(n_estimators=1000)
random_forest.fit(X_train, Y_train)
Y_pred = random_forest.predict(X_val)
acc_random_forest = round(random_forest.score(X_val, Y_val) * 100, 2)
acc_random_forest
# +
# Look at importnace of features for random forest
def plot_model_var_imp( model , X , y ):
imp = pd.DataFrame(
model.feature_importances_ ,
columns = [ 'Importance' ] ,
index = X.columns
)
imp = imp.sort_values( [ 'Importance' ] , ascending = True )
imp[ : 10 ].plot( kind = 'barh' )
print ('Training accuracy Random Forest:',model.score( X , y ))
plot_model_var_imp(random_forest, X_train, Y_train)
# -
# How to create a Kaggle submission:
Y_submission = random_forest.predict(X_submission)
submission = pd.DataFrame({
"PassengerId": test_df["PassengerId"],
"Survived": Y_submission
})
submission.to_csv('titanic.csv', index=False)
# # Legacy code (not used anymore)
# ```python
# # Map title string values to numbers so that we can make predictions
#
# title_mapping = {"Mr": 1, "Miss": 2, "Mrs": 3, "Master": 4, "Rare": 5}
# for dataset in combine:
# dataset['Title'] = dataset['Title'].map(title_mapping)
# dataset['Title'] = dataset['Title'].fillna(0)
# # Handle missing values
#
# train_df.head()
# ```
#
# ```python
# # Drop the unnecessary Name column (we have the titles now)
#
# train_df = train_df.drop(['Name', 'PassengerId'], axis=1)
# test_df = test_df.drop(['Name'], axis=1)
# combine = [train_df, test_df]
# train_df.shape, test_df.shape
# ```
#
# ```python
# # Create categorical dummy variables for Embarked values
# for dataset in combine:
# dataset['Embarked'] = dataset['Embarked'].map( {'S': 0, 'C': 1, 'Q': 2} ).astype(int)
#
# train_df.head()
# ```
| 06-ML-titanic/titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 3
# +
# imports
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import unyt
from geopy import distance
# -
sns.set_theme()
sns.set_style('whitegrid')
sns.set_context('notebook')
# # Problem 2
#
# ## Dispersion Relation for internal waves:
#
# ## $\omega^2 = \left ( \frac{k_x^2 + k_y^2}{k^2} \right ) N^2$
#
# ## Introduce $\phi$ as the angle off the plane of $\vec k$, then
#
# ## $\omega^2 = \cos^2\phi \, N^2$
#
# ## Now the water parcel motions (ignoring $y$ motions for the moment) are given by:
#
# ## $w = \hat w \, {\rm e}^{i (k_x x + k_z z - \omega t)}$
#
# ## and
#
# ## $u = \frac{-k_z}{k_x} \hat w \, {\rm e}^{i (k_x x + k_z z - \omega t)}$
#
# ## So we can define an angle for the particle motion off the horizon as $\alpha$ and this means:
#
# ## $\tan \alpha = \frac{k_x}{k_z}$
#
# ## Last, we recognize that $\tan\phi = k_z/k_x$ so that $\cot\phi = k_x/k_z$ and
#
# ## $\alpha = \tan^{-1} (\cot\phi)$
#
# ## And for a given period $P$, $\omega = 2\pi/P$
def phi_P(P, N=0.01*unyt.rad/unyt.s):
omega = 2*np.pi/P
#
phiv = np.arccos(omega/N)
#print('phi = {}'.format(phiv*180./2/np.pi))
return float(phiv)*unyt.rad
def phi_to_alpha(phi):
return np.arctan(1./np.tan(phi))*unyt.rad
def P_to_alpha(P, N=0.01*unyt.rad/unyt.s):
phi = phi_P(P, N=N)
alpha = phi_to_alpha(phi)
return alpha
# ## (a) 11 minutes
P_to_alpha(11*unyt.min).to('deg')
# ### Nearly vertical motion
# ## (b) 2 hours
P_to_alpha(2*unyt.hour).to('deg')
# ### Nearly even-steven between horizontal and vertical
# ## (c) 12.4 hours
P_to_alpha(12.4*unyt.hour).to('deg')
# ### Very close to purely horizontal
# ## (d) 24 hours
P_to_alpha(24.*unyt.hour).to('deg')
# ### Even closer
# ----
# # (3) Group Velocity
#
# ## We start with the same dispersion relation as above:
#
# ## $\omega^2 = \left ( \frac{k_x^2 + k_y^2}{k^2} \right ) N^2$
# ## (a) Calculate the group velocity vector:
#
# ## $\frac{\partial \omega}{\partial k_x} = \frac{k_x k_z^2 N}{k^3 \sqrt{k_x^2 + k_y^2}}$
#
# ### and
#
# ## $\frac{\partial \omega}{\partial k_y} = \frac{k_y k_z^2 N}{k^3 \sqrt{k_x^2 + k_y^2}}$
#
# ### and
#
# ## $\frac{\partial \omega}{\partial k_z} = - \frac{k_z \sqrt{k_x^2 + k_y^2} N}{k^3}$
#
# ## Re-expressing
#
# ## $\vec c_g = [\frac{k_x k_z}{k_x^2 + k_y^2}, \frac{k_y k_z}{k_x^2 + k_y^2}, -1] \frac{k_z N \sqrt{k_x^2 + k_y^2}}{k^3}$
# ## (b) Compare to phase velocity
#
# ## $\vec c_p = \frac{\omega}{k^2} [k_x, k_y, k_z]$
#
# ## Clearly, the phase velocity is oriented parallel to $\vec k$
#
# ## Let's consider the dot product: $\vec c_p \cdot \vec c_g$
#
# ## The first two terms reduce to $k_z$ and the third term gives $-k_z$, i.e. the group velocity is orthogonal to the phase velocity!
#
# ## (c) Magnitude and direction in limiting cases
# ## As $\omega \to 0$, this requires $k_z \gg k_x, k_y$i.e. $\vec k$ points vertically.
#
# ## In this limit, $\vec c_g$ will point horizontally. Taking $k_y=0$ for simplicity:
#
# ## $\vec c_g = \frac{k_z^2 N}{k^3} \hat x$ which for $k \approx k_z$ has magnitude
#
# ## $|\vec c_g| = N/k_z$
# ## For $\omega \to N$, this implies $k_z \ll k_x, k_y$. For the group velocity, this yields $\vec c_g || \hat z$, i.e. vertical direction.
#
# ## Meanwhile, the magnitude will go (taking $k_y=0$ for simlicity) as:
#
# ## $\frac{k_z N k_x}{k^3}$ which reduces to
#
# ## $\frac{N k_z}{k_x^2}$ which will be very small.
# ----
# # (4) Earth's Rotation
#
# ## Momentum equations:
#
# ## $\frac{\partial u}{\partial t} = - \frac{1}{\rho_0} \frac{\partial p}{\partial x} + fv$
#
# ## $\frac{\partial v}{\partial t} = - \frac{1}{\rho_0} \frac{\partial p}{\partial y} - fu$
#
# ## $\frac{\partial w}{\partial t} = - \frac{1}{\rho_0} \frac{\partial p}{\partial z} - \frac{\rho'}{\rho_0} g$
#
#
# ## $\nabla \cdot \vec u = 0$
#
# ## $\frac{\partial \rho'}{\partial t} = \frac{\rho_0}{g} N^2 \omega^2$
# ## Following Kundu, we take the time derivative of the continuity equation:
#
# ## $\frac{\partial^2 u}{\partial t \partial x} + \frac{\partial^2 v}{\partial t \partial y} + \frac{\partial^2 w}{\partial t \partial z} = 0$
# ## And now replace the time derivatives for $u, v$:
#
# ## $\frac{1}{\rho_0} \nabla_H^2 p - f [\frac{\partial v}{\partial x} - \frac{\partial u}{\partial y}] = \frac{\partial^2 w}{\partial t \partial z}$
#
# ## where from GFD, we recognize the relative vorticity:
#
# ## $\zeta = \partial v / \partial x - \partial u / \partial y$
#
# ## $\frac{1}{\rho_0} \nabla_H^2 p - f \zeta = \frac{\partial^2 w}{\partial t \partial z}$
#
# ## Using the time derivative of our third momentum equation and our 5th equation:
#
# ## $\frac{1}{\rho_0} \frac{\partial^2 p}{\partial t \partial z} = -\frac{\partial^2 w}{\partial t^2} - N^2 w$
# ## Now apply $\nabla_H^2$ to this equation and use the equation two up that involves $\nabla_H^2 p$ to get:
#
# ## $\frac{\partial^2}{\partial t \partial z} \left [ \frac{\partial^2 w}{\partial t \partial z} - f \zeta \right ] =
# -\nabla_H^2 \left ( \frac{\partial^2 w}{\partial t^2} + N^2 w \right ) $
# ## Now time for a bit of magic to deal with the $f\zeta$ term. Take the z-component of the curl of our momentum equations. Here they are before we subtract:
#
# ## $\frac{\partial^2 u}{\partial t \partial y} = - \frac{1}{\rho_0} \frac{\partial^2 p}{\partial x \partial y} + f \frac{\partial v}{\partial y}$
#
# ## $\frac{\partial^2 v}{\partial t \partial x} = - \frac{1}{\rho_0} \frac{\partial^2 p}{\partial y \partial x} - f \frac{\partial u}{\partial x}$
#
# ## Now subtract 1 from 2 to complete the curl, noting the pressure terms vanish:
#
# ## $\frac{\partial \zeta}{\partial t} = - f (\frac{\partial u}{\partial x} + \frac{\partial v}{\partial y})$
# ## And use continuity to give:
#
# ## $\frac{\partial \zeta}{\partial t} = f \frac{\partial w}{\partial z}$
# ## So, applying the time dertivatie of $\partial^2 / \partial t \partial z$ to $f\zeta$, we wind up with:
#
# ## $\frac{\partial^2}{\partial t \partial z} \left [ \frac{\partial^2 w}{\partial t \partial z} \right ] - f^2 \frac{\partial^2 w}{\partial z^2} =
# -\nabla_H^2 \left ( \frac{\partial^2 w}{\partial t^2} + N^2 w \right ) $
# ## Massaging, we finish with:
#
# ## $\left [ \frac{\partial^2}{\partial t^2} \left (\nabla_H^2 + \frac{\partial^2}{\partial z^2} \right ) + f^2 \frac{\partial^2}{\partial z^2} + N^2 \nabla_H^2 \right ] w = 0$
# ## (b) Dispersion relation for solutions for
#
# ## $w = \hat w {\rm e}^{i (k_x x + k_y y + k_z z - \omega t)}$
#
# ## $-\omega^2 k^2 + f^2 k_z^2 + N^2 (k_x^2 + k_y^2) = 0$
#
# ## or
#
# ## $\omega^2 = \frac{f^2 k_z^2 + N^2 (k_x^2 + k_y^2)}{k^2} = f^2 \sin^2 \phi + N^2 \cos^2 \phi$
# ## So we have an additonal term in the numerator that sets a floor to the frequency for vertical moving waves, dependent on $f$
# ## (c) Recalculating answers for Problem 2
def phi_P_f(P, N=0.01/unyt.s, f=1e-4/unyt.s):
omega = 2*np.pi/P
# Branch
if omega > f:
phiv = np.arccos(np.sqrt(omega**2-f**2)/np.sqrt(N**2 - f**2))
else:
phiv = np.arcsin(np.sqrt(N**2-omega**2)/np.sqrt(N**2-f**2))
#
#print('phi = {}'.format(phiv*180./2/np.pi))
return float(phiv)*unyt.rad
def P_to_alpha_f(P, N=0.01/unyt.s, f=1e-4/unyt.s):
phi = phi_P_f(P, N=N, f=f)
alpha = phi_to_alpha(phi)
return alpha
# ### (i) 11 min
P_to_alpha(11*unyt.min).to('deg'), P_to_alpha_f(11*unyt.min).to('deg')
# ### The new angle is slightly smaller
# ### (ii) 2 hours
t = 2*unyt.hour
P_to_alpha(t).to('deg'), P_to_alpha_f(t).to('deg')
# ### Again a bit smaller
# ### (iii) 12.4 hours
t = 12.4*unyt.hour
P_to_alpha(t).to('deg'), P_to_alpha_f(t).to('deg')
# ### Notably smaller (30% or so)
# ### (iv) 24 hours
t = 24*unyt.hour
P_to_alpha(t).to('deg'), P_to_alpha_f(t).to('deg')
# ### Here the solution breaks as the frequency is smaller than the Coriolis parameter!
| SIO-211A/hw/Homework3_rest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Activity 3: How to Build a Sunflower 🌻
# ___
# In this activity, you will learn how to code the intricate, beautiful packing of florets in a sunflower head, explore the sensitivity of the arrangement of florets to the golden angle, and create animations of a growing sunflower.
#
# Your reading assignment before this activity is ["Chapter 4 - Phyllotaxis"](http://algorithmicbotany.org/papers/abop/abop-ch4.pdf) from [*The Algorithmic Beauty of Plants*](http://algorithmicbotany.org/papers/#abop). The inspiration and theory behind the coding you will be doing today comes from there. The formulas, too, also come from the chapter, but ultimately derive from the classic work by Vogel, "A better way to construct the sunflower head". Both of these works form the basis for today's activity. Skim over the reading assignment if you have the chance (there is no need to , to get a feel for some of the math we will be using in today's activity.
#
# > **References:** <br/><br/> <NAME> and <NAME>. The algorithmic beauty of plants. Springer-Verlag, Berlin, Heidelberg, 1990. ISBN:0-387-97297-8 <br/><br/> <NAME>. A better way to construct the sunflower head. *Mathematical Biosciences*, 44:179–189, 1979.
# ___
# ## The formula for a sunflower
#
# From *The Algorithmic Beauty of Plants* chapter you read, you learned that Vogel had proposed the following formulas to calculate the angle and radius of each floret on a sunflower head:
#
# ```python
# theta = n * phi
# r = sqrt(n)
# ```
#
# Where `n` is the ordered number of each floret on a sunflower head, `theta` is the angle of the floret, `phi` is the golden angle, and `r` is radius, the floret's distance from the center of the sunflower. You should think of each sequential floret as having one more "turn" of `phi` and having a longer radius than the previous, following a square root function.
#
# Also remember back to trigonometry (nobody's favorite class, but let's try to recall) that `cos` and `sin` are the adjacent and opposite sides, respectively, of a right triangle divided by the hypotenuse. In the context of a radial Cartesian plane centered at (0,0), to retrieve the `x` and `y` coordinates we use the following equations:
#
# ```python
# x = r * cos(theta)
# y = r * sin(theta)
# ```
#
# Finally, we will be working in radians, not degrees. Remember also from trigonometry that to convert degrees into radians, we multiply degrees by ($\pi$ / 180)
# ___
# ## Build a sunflower!
#
# Using the equations above, build a sunflower!
#
# But first, since we will be using the golden angle quite a lot, let's just make that a variable. In the cell below, the golden angle is provided. Create a new variable called `phi` that is the golden angle in radians.
#
# It would be convenient to have $\pi$. The cell below imports the `math` module, which you will often likely need! To retrieve the value of $\pi$, simply type `math.pi`. You are also going to need `math` for the `math.sqrt()`, `math.cos()`, and `math.sin()` functions in this lesson.
# +
import math
# From the online encyclopedia of integer sequences:
# https://oeis.org/A096627
golden_angle = 137.5077640500378546463487396283702776206886952699253696312384958261062333851951
# Convert golden_angle to radians here. Call the variable phi
# -
# Next, write a `for` loop to calculate the `x` and `y` coordinates for each floret.
#
# **Do the following:**
#
# * Create two lists for your coordinates outside of the loop: a list for the `x` values and a list for the `y` values
# * Your `for` loop should iterate 1000 times, calculating the `x` and `y` coordinate values for 1000 florets
# * Within your loop, for each floret calculate:
# * Radius
# * Theta
# * x coordinate value
# * y coordinate value
# * Append `x` and `y` coordinate values to their respective lists
# +
# Put your answer here
# -
# Next, plot your results using matplotlib!
#
# * Pay attention to the size of the points, not too much overlap but not too small, either
# * Pick an appropriate color
# * Adjust the alpha appropriately
# * Consider scaling the x and y axes equal and setting a fixed aspect ratio
# * Turn the axes off
# * Always make aesthetically pleasing figures, ***especially*** when plotting sunflowers
# +
# You must alway import matplotlib before using it and use the inline code
import matplotlib.pyplot as plt
# %matplotlib inline
# Put the code for your plot here
# -
# ___
# ## What a difference an angle makes!
# You learned in the *The Algorithmic Beauty of Plants* chapter that you read that the packing of florets in a sunflower is extremely sensitive to the exact value of the golden angle. A little bit more or less and the beautifully-spaced arrangement that is readily apparent to the eye is destroyed. This is because of the special mathematical properties of the golden angle.
#
# In this exercise, you will be taking the sunflower florets you just created using your `for` loop and creating an animation, where the value of the golden angle deviates from -1 to +1 degrees over 100 steps. Using this animation, you will observe the sensitivity of the packing of florets in a sunflower to the golden angle, and how this sensitivity varies closer to and farther from the center.
#
# Our strategy will be to create a loop within a loop! The logic is this: you have already created a loop that will calculate all the `x` & `y` coordinates in a sunflower. You put that loop into another loop! The outer loop will iterate over deviations of the golden angle. So, the flow will look like this:
#
# 1. Outer loop creates a new angle value,
# 2. Inner loop caclulates all the `x` and `y` coordinate values for the new angle value,
# 3. The inner loop is exited, you are back in the outer loop,
# 4. Your previous code to plot a sunflower plots the florets for the current angle,
# 5. The outer loop repeats with a new angle value and a new frame of the animation!
#
# Just one more thing. We will be iterating in 100 steps from -1 to +1 degrees of the golden angle. How do you specify a set number of steps between two values? One function to do this is `linspace()`, from the `numpy` module. `linspace()` takes a start value (inclusive), stop value (inclusive), and a number of steps to incrementally iterate over the interval. Use this for your outer loop to minutely adjust the angle value from +- 1 degree of the golden angle!
#
# In the cell below, pseudo-code is provided to help guide you in constructing your first loop of a loop! Remember, you can almost use the previous code (with some modifications) for your loop to calculate `x` and `y` coordinate values and to plot your sunflower in the loop of loops below.
#
# This exercise is meant for you to spend time thinking about how complex loops work and how to modify and adjust code for different purposes. So spend time thinking through this.
#
# If you're successful, you will create a beautiful sunflower animation!
#
# Follow the pseudo-code and fill out the missing parts!
# +
# Imports to get the animation to work
from IPython.display import display, clear_output
import time
# Import numpy to use linspace
import numpy as np
# Using linspace() to create 100 increments between -1 and +1
for i in np.linspace(-1, 1, 100):
# Here, create a "new_angle" variable that is golden_angle + i
# Remember the new angle must be in radians!
# This is where to insert your previous loop to calculate x and y coords for the new angle
# Remember, theta will be calculated from your new angle! So be sure to change that!
# Remember, properly indent your for loop within the other for loop: indents of indents!
# Remember, keep the x and y coordinate lists outside of this loop (but within the first loop)
# Still calculate x and y values for 1000 florets using your previous loop
# Your previous loop code goes in this space!
# You are now outside of the inner loop and back in the outer loops
# For animation, you need to call a figure
fig = plt.figure(figsize=(10,10))
# Put your code for your plot of the sunflower here
# This is the code that creates the animation
time.sleep(0.001)
clear_output(wait=True)
display(fig)
fig.clear()
# Closes the figure animation once complete
plt.close()
# -
# ___
# ## But sunflowers grow!
#
# Yes, it's nice to create an animation changing the angle of florets with a fixed radius. But in the real world a sunflower is growing! The florets arise from a meristem (a population of stem cells) at the center of the sunflower. At the periphery of the shoot apical meristem in the center of the sunflower, cells are determined to become a floret. We call the ability of cells, like meristems, to produce different types of tissues "pluripotent". We say that cells are "determined" or "fated" to become a particular type of tissue as they lose pluripotency. "Differentiated" cells have already become a particular tissue. After a floret has differentiated, it moves away from the center of the sunflower.
#
# In your previous plots, the florets at the periphery were the first to arise, and were "pushed" from the center by all the other florets that subsequently arose. Let's create an animation where we watch florets arise at the center and move outwards!
#
# You will:
#
# 1. Create pre-populated lists of `thetas` and `radii`. Simply use the code you already have to calculate the thetas and radii.
# 2. Once born, a floret ***always*** keeps the same theta. But the radius gets longer and longer.
# 3. Our first loop will create lists of theta and radii that increase in length with each iteration. Because of this, we will start with one floret, then two, and so on, and with each additional floret, the radii of the first florets will become longer and longer. We will start with no florets and grow our sunflower, adding more and more. This loop takes one more member of your pre-specified `thetas` and `radii` lists with each iteration.
# 4. There is a problem, though. Each time we add more `thetas` (which are associated with unique florets), the oldest/first `thetas` are at the ***beginning*** of the list. The oldest `radii`, too, are at the beginning of their list and they will be the shortest radii! The oldest/first `thetas` should have the longest radii, not the shortest!
# 5. We will use the `.reverse()` function, which reverses the order of elements of a list. What this will do is insure that the oldest/first `thetas` of the thetas list correspond to the longest radii after the radii list is reversed.
# 6. Think through the reasoning above, it's complicated. We are adding one more floret with each iteration. But the first floret would always have the shortest radius, unless we reversed the radii list. By reversing the radii list, the first floret will have an increasing radius.
#
# Follow the reasoning above and make sure you understand it. Pseudo-code with comments is provided in the cells below. Using the skeleton, fill out the rest of the code. The code is in two parts:
#
# 1. First, using a `for` loop, create two lists: `thetas` and `radii`. These lists should contain **750** theta and radius values for florets.
#
# 2. Using the pseudo-code provided, create an animation of a growing sunflower.
# +
# In this cell, create two emtpy lists: "thetas" and "radii"
# Then, using a for loop, populate your two lists with theta and radii values for 750 florets
# You can use the code you have already produced
# Put your answer here
# +
# Next, using the pseudo-code below, create an animation of growing florets in a sunflower
# Pay attention to what the two loops are doing and how they function in real time
# Pay attention to indexing of lists
# Pay attention to the use of the .reverse() function
# Understand how this code creates an animation of a growing sunflower, frame-by-frame
# Put your answer below
# Imports to get the animation to work
from IPython.display import display, clear_output
import time
for i in range(750):
xlist = []
ylist = []
# This selects the current set of thetas to use
current_thetas = thetas[0:(i+1)]
# This selects the current set of radii to use
# Everytime the loop runs, this list will be recreated in the "forward" direction
# When created, shortest radii are first, longest radii are last
current_radii = radii[0:(i+1)]
# After creating the list of radii, "reverse" its order
# Reversing the order of the radii means that the
# florets first specified (the first thetas) have the longest radius
# After reversing, longest radii are first, shortest radii are last
current_radii.reverse()
for n in range (i+1):
# Here, for each floret n caclulate r, theta, x, y, and append x & y to their lists
# You are now back in the outer loop
# For animation, you need to call a figure
fig = plt.figure()
# Put your code to make a plot of a sunflower here
# This is the code that creates the animation
time.sleep(0.001)
clear_output(wait=True)
display(fig)
fig.clear()
# Closes the figure animation once complete
plt.close()
# -
# That's all for this activity! Thank you for participating!
| STUDENT_Activity 3--How to build a sunflower.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # indicators
import vectorbt as vbt
# + Collapsed="false"
import numpy as np
import pandas as pd
from datetime import datetime, timedelta
from numba import njit
import itertools
import talib
import ta
# -
# Disable caching for performance testing
vbt.settings.caching['enabled'] = False
close = pd.DataFrame({
'a': [1., 2., 3., 4., 5.],
'b': [5., 4., 3., 2., 1.],
'c': [1., 2., 3., 2., 1.]
}, index=pd.DatetimeIndex([
datetime(2018, 1, 1),
datetime(2018, 1, 2),
datetime(2018, 1, 3),
datetime(2018, 1, 4),
datetime(2018, 1, 5)
]))
np.random.seed(42)
high = close * np.random.uniform(1, 1.1, size=close.shape)
low = close * np.random.uniform(0.9, 1, size=close.shape)
volume = close * 0 + np.random.randint(1, 10, size=close.shape).astype(float)
big_close = pd.DataFrame(np.random.randint(10, size=(1000, 1000)).astype(float))
big_close.index = [datetime(2018, 1, 1) + timedelta(days=i) for i in range(1000)]
big_high = big_close * np.random.uniform(1, 1.1, size=big_close.shape)
big_low = big_close * np.random.uniform(0.9, 1, size=big_close.shape)
big_volume = big_close * 0 + np.random.randint(10, 100, size=big_close.shape).astype(float)
close_ts = pd.Series([1, 2, 3, 4, 3, 2, 1], index=pd.DatetimeIndex([
datetime(2018, 1, 1),
datetime(2018, 1, 2),
datetime(2018, 1, 3),
datetime(2018, 1, 4),
datetime(2018, 1, 5),
datetime(2018, 1, 6),
datetime(2018, 1, 7)
]))
high_ts = close_ts * 1.1
low_ts = close_ts * 0.9
volume_ts = pd.Series([4, 3, 2, 1, 2, 3, 4], index=close_ts.index)
# ## IndicatorFactory
# +
def apply_func(i, ts, p, a, b=100):
return ts * p[i] + a + b
@njit
def apply_func_nb(i, ts, p, a, b):
return ts * p[i] + a + b # numba doesn't support **kwargs
# Custom function can be anything that takes time series, params and other arguments, and returns outputs
def custom_func(ts, p, *args, **kwargs):
return vbt.base.combine_fns.apply_and_concat_one(len(p), apply_func, ts, p, *args, **kwargs)
@njit
def custom_func_nb(ts, p, *args):
return vbt.base.combine_fns.apply_and_concat_one_nb(len(p), apply_func_nb, ts, p, *args)
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_custom_func(custom_func, var_args=True)
.run(close, [0, 1], 10, b=100).out)
print(F.from_custom_func(custom_func_nb, var_args=True)
.run(close, [0, 1], 10, 100).out)
# +
# Apply function is performed on each parameter individually, and each output is then stacked for you
# Apply functions are less customizable than custom functions, but are simpler to write
def apply_func(ts, p, a, b=100):
return ts * p + a + b
@njit
def apply_func_nb(ts, p, a, b):
return ts * p + a + b # numba doesn't support **kwargs
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(apply_func, var_args=True)
.run(close, [0, 1], 10, b=100).out)
print(F.from_apply_func(apply_func_nb, var_args=True)
.run(close, [0, 1], 10, 100).out)
# -
# test *args
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts, p, a: ts * p + a, var_args=True)
.run(close, [0, 1, 2], 3).out)
print(F.from_apply_func(njit(lambda ts, p, a: ts * p + a), var_args=True)
.run(close, [0, 1, 2], 3).out)
# test **kwargs
# Numba doesn't support kwargs out of the box
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts, p, a=1: ts * p + a)
.run(close, [0, 1, 2], a=3).out)
# test no inputs
F = vbt.IndicatorFactory(param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda p: np.full((3, 3), p))
.run([0, 1]).out)
print(F.from_apply_func(njit(lambda p: np.full((3, 3), p)))
.run([0, 1]).out)
# +
# test no inputs with input_shape, input_index and input_columns
F = vbt.IndicatorFactory(param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda input_shape, p: np.full(input_shape, p), require_input_shape=True)
.run((5,), 0).out)
print(F.from_apply_func(njit(lambda input_shape, p: np.full(input_shape, p)), require_input_shape=True)
.run((5,), 0).out)
print(F.from_apply_func(lambda input_shape, p: np.full(input_shape, p), require_input_shape=True)
.run((5,), [0, 1]).out)
print(F.from_apply_func(njit(lambda input_shape, p: np.full(input_shape, p)), require_input_shape=True)
.run((5,), [0, 1]).out)
print(F.from_apply_func(lambda input_shape, p: np.full(input_shape, p), require_input_shape=True)
.run((5, 3), [0, 1], input_index=close.index, input_columns=close.columns).out)
print(F.from_apply_func(njit(lambda input_shape, p: np.full(input_shape, p)), require_input_shape=True)
.run((5, 3), [0, 1], input_index=close.index, input_columns=close.columns).out)
# -
# test multiple inputs
F = vbt.IndicatorFactory(input_names=['ts1', 'ts2'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts1, ts2, p: ts1 * ts2 * p)
.run(close, high, [0, 1]).out)
print(F.from_apply_func(njit(lambda ts1, ts2, p: ts1 * ts2 * p))
.run(close, high, [0, 1]).out)
# test no params
F = vbt.IndicatorFactory(input_names=['ts'], output_names=['out'])
print(F.from_apply_func(lambda ts: ts)
.run(close).out)
print(F.from_apply_func(njit(lambda ts: ts))
.run(close).out)
# test no inputs and no params
F = vbt.IndicatorFactory(output_names=['out'])
print(F.from_apply_func(lambda: np.full((3, 3), 1))
.run().out)
print(F.from_apply_func(njit(lambda: np.full((3, 3), 1)))
.run().out)
# test multiple params
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2))
.run(close, np.asarray([0, 1]), np.asarray([2, 3])).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)))
.run(close, np.asarray([0, 1]), np.asarray([2, 3])).out)
# test param_settings array_like
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2),
param_settings={'p1': {'is_array_like': True}})
.run(close, np.asarray([0, 1, 2]), np.asarray([2, 3])).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)),
param_settings={'p1': {'is_array_like': True}})
.run(close, np.asarray([0, 1, 2]), np.asarray([2, 3])).out)
# test param_settings bc_to_input
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2),
param_settings={'p1': {'is_array_like': True, 'bc_to_input': True}})
.run(close, np.asarray([0, 1, 2]), np.asarray([2, 3])).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)),
param_settings={'p1': {'is_array_like': True, 'bc_to_input': True}})
.run(close, np.asarray([0, 1, 2]), np.asarray([2, 3])).out)
# test param product
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2))
.run(close, [0, 1], [2, 3], param_product=True).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)))
.run(close, [0, 1], [2, 3], param_product=True).out)
# test default params
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2), p2=2)
.run(close, [0, 1]).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)), p2=2)
.run(close, [0, 1]).out)
# test hide_params
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2), hide_params=['p2'])
.run(close, [0, 1], 2).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)), hide_params=['p2'])
.run(close, [0, 1], 2).out)
# test hide_default
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2), p2=2)
.run(close, [0, 1], hide_default=False).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)), p2=2)
.run(close, [0, 1], hide_default=False).out)
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2), p2=2)
.run(close, [0, 1], hide_default=True).out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)), p2=2)
.run(close, [0, 1], hide_default=True).out)
# test multiple outputs
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['o1', 'o2'])
print(F.from_apply_func(lambda ts, p: (ts * p, ts * p ** 2))
.run(close, [0, 1]).o1)
print(F.from_apply_func(lambda ts, p: (ts * p, ts * p ** 2))
.run(close, [0, 1]).o2)
print(F.from_apply_func(njit(lambda ts, p: (ts * p, ts * p ** 2)))
.run(close, [0, 1]).o1)
print(F.from_apply_func(njit(lambda ts, p: (ts * p, ts * p ** 2)))
.run(close, [0, 1]).o2)
# +
# test in-place outputs
def apply_func(ts, ts_out, p):
ts_out[:, 0] = p
return ts * p
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'], in_output_names=['ts_out'])
print(F.from_apply_func(apply_func)
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(njit(apply_func))
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(apply_func, in_output_settings={'ts_out': {'dtype': np.int_}})
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(njit(apply_func), in_output_settings={'ts_out': {'dtype': np.int_}})
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(apply_func, ts_out=-1)
.run(close, [0, 1]).ts_out)
print(F.from_apply_func(njit(apply_func), ts_out=-1)
.run(close, [0, 1]).ts_out)
# -
# test kwargs_to_args
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts, p, a, kw: ts * p + a + kw, kwargs_to_args=['kw'], var_args=True)
.run(close, [0, 1, 2], 3, kw=10).out)
print(F.from_apply_func(njit(lambda ts, p, a, kw: ts * p + a + kw), kwargs_to_args=['kw'], var_args=True)
.run(close, [0, 1, 2], 3, kw=10).out)
# test caching func
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p'], output_names=['out'])
print(F.from_apply_func(lambda ts, param, c: ts * param + c, cache_func=lambda ts, params: 100)
.run(close, [0, 1]).out)
print(F.from_apply_func(njit(lambda ts, param, c: ts * param + c), cache_func=njit(lambda ts, params: 100))
.run(close, [0, 1]).out)
# test run_combs
F = vbt.IndicatorFactory(input_names=['ts'], param_names=['p1', 'p2'], output_names=['out'])
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2))
.run_combs(close, [0, 1, 2], [3, 4, 5], short_names=['i1', 'i2'])[0].out)
print(F.from_apply_func(lambda ts, p1, p2: ts * (p1 + p2))
.run_combs(close, [0, 1, 2], [3, 4, 5], short_names=['i1', 'i2'])[1].out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)))
.run_combs(close, [0, 1, 2], [3, 4, 5], short_names=['i1', 'i2'])[0].out)
print(F.from_apply_func(njit(lambda ts, p1, p2: ts * (p1 + p2)))
.run_combs(close, [0, 1, 2], [3, 4, 5], short_names=['i1', 'i2'])[1].out)
# +
from collections import namedtuple
TestEnum = namedtuple('TestEnum', ['Hello', 'World'])(0, 1)
# test attr_settings
F = vbt.IndicatorFactory(
input_names=['ts'], output_names=['o1', 'o2'], in_output_names=['ts_out'],
attr_settings={
'ts': {'dtype': None},
'o1': {'dtype': np.float_},
'o2': {'dtype': np.bool_},
'ts_out': {'dtype': TestEnum}
}
)
dir(F.from_apply_func(lambda ts, ts_out: (ts + ts_out, ts + ts_out)).run(close))
# -
CustomInd = vbt.IndicatorFactory(
input_names=['ts1', 'ts2'],
param_names=['p1', 'p2'],
output_names=['o1', 'o2']
).from_apply_func(lambda ts1, ts2, p1, p2: (ts1 * p1, ts2 * p2))
dir(CustomInd) # you can list here all of the available tools
custom_ind = CustomInd.run(close, high, [1, 2], [3, 4])
big_custom_ind = CustomInd.run(big_close, big_high, [1, 2], [3, 4])
print(custom_ind.wrapper.index) # subclasses ArrayWrapper
print(custom_ind.wrapper.columns)
print(custom_ind.wrapper.ndim)
print(custom_ind.wrapper.shape)
print(custom_ind.wrapper.freq)
# not changed during indexing
print(custom_ind.short_name)
print(custom_ind.level_names)
print(custom_ind.input_names)
print(custom_ind.param_names)
print(custom_ind.output_names)
print(custom_ind.output_flags)
print(custom_ind.p1_list)
print(custom_ind.p2_list)
# ### Pandas indexing
# +
print(custom_ind._ts1)
print(custom_ind.ts1)
print(custom_ind.ts1.iloc[:, 0])
print(custom_ind.iloc[:, 0].ts1)
print(custom_ind.ts1.iloc[:, [0]])
print(custom_ind.iloc[:, [0]].ts1)
print(custom_ind.ts1.iloc[:2, :])
print(custom_ind.iloc[:2, :].ts1)
# +
print(custom_ind.o1.iloc[:, 0])
# %timeit big_custom_ind.o1.iloc[:, 0] # benchmark, 1 column
print(custom_ind.iloc[:, 0].o1) # performed on the object itself
# %timeit big_custom_ind.iloc[:, 0] # slower since it forwards the operation to each dataframe
# +
print(custom_ind.o1.iloc[:, np.arange(3)])
# %timeit big_custom_ind.o1.iloc[:, np.arange(1000)] # 1000 columns
print(custom_ind.iloc[:, np.arange(3)].o1)
# %timeit big_custom_ind.iloc[:, np.arange(1000)]
# +
print(custom_ind.o1.loc[:, (1, 3, 'a')])
# %timeit big_custom_ind.o1.loc[:, (1, 3, 0)] # 1 column
print(custom_ind.loc[:, (1, 3, 'a')].o1)
# %timeit big_custom_ind.loc[:, (1, 3, 0)]
# +
print(custom_ind.o1.loc[:, (1, 3)])
# %timeit big_custom_ind.o1.loc[:, 1] # 1000 columns
print(custom_ind.loc[:, (1, 3)].o1)
# %timeit big_custom_ind.loc[:, 1]
# +
print(custom_ind.o1.xs(1, axis=1, level=0))
# %timeit big_custom_ind.o1.xs(1, axis=1, level=0) # 1000 columns
print(custom_ind.xs(1, axis=1, level=0).o1)
# %timeit big_custom_ind.xs(1, axis=1, level=0)
# -
# ### Parameter indexing
# Indexing by parameter
print(custom_ind._p1_mapper)
print(custom_ind.p1_loc[2].o1)
print(custom_ind.p1_loc[1:2].o1)
print(custom_ind.p1_loc[[1, 1, 1]].o1)
# %timeit big_custom_ind.p1_loc[1] # 1000 columns
# %timeit big_custom_ind.p1_loc[np.full(10, 1)] # 10000 columns
print(custom_ind._tuple_mapper)
print(custom_ind.tuple_loc[(1, 3)].o1)
print(custom_ind.tuple_loc[(1, 3):(2, 4)].o1)
# %timeit big_custom_ind.tuple_loc[(1, 3)]
# %timeit big_custom_ind.tuple_loc[[(1, 3)] * 10]
# ### Comparison methods
# +
print(custom_ind.o1 > 2)
# %timeit big_custom_ind.o1.values > 2 # don't even try pandas
print(custom_ind.o1_above(2))
# %timeit big_custom_ind.o1_above(2) # slower than numpy because of constructing dataframe
# +
print(pd.concat((custom_ind.o1 > 2, custom_ind.o1 > 3), axis=1))
# %timeit np.hstack((big_custom_ind.o1.values > 2, big_custom_ind.o1.values > 3))
print(custom_ind.o1_above([2, 3]))
# %timeit big_custom_ind.o1_above([2, 3])
# -
# ## TA-Lib
ts = pd.DataFrame({
'a': [1, 2, 3, 4, np.nan],
'b': [np.nan, 4, 3, 2, 1],
'c': [1, 2, np.nan, 2, 1]
}, index=pd.DatetimeIndex([
datetime(2018, 1, 1),
datetime(2018, 1, 2),
datetime(2018, 1, 3),
datetime(2018, 1, 4),
datetime(2018, 1, 5)
]))
# +
SMA = vbt.talib('SMA')
print(SMA.run(close['a'], 2).real)
print(SMA.run(close, 2).real)
print(SMA.run(close, [2, 3]).real)
# -
# %timeit SMA.run(big_close)
# %timeit SMA.run(big_close, np.arange(2, 10))
# %timeit SMA.run(big_close, np.full(10, 2))
# %timeit SMA.run(big_close, np.full(10, 2), run_unique=True)
comb = itertools.combinations(np.arange(2, 20), 2)
fast_windows, slow_windows = np.asarray(list(comb)).transpose()
print(fast_windows)
print(slow_windows)
# %timeit SMA.run(big_close, fast_windows), SMA.run(big_close, slow_windows) # individual caching
# %timeit SMA.run_combs(big_close, np.arange(2, 20)) # mutual caching
# %timeit vbt.MA.run(big_close, fast_windows), vbt.MA.run(big_close, slow_windows) # the same using Numba
# %timeit vbt.MA.run_combs(big_close, np.arange(2, 20))
sma1, sma2 = SMA.run_combs(close, [2, 3, 4])
print(sma1.real_above(sma2, crossover=True))
print(sma1.real_below(sma2, crossover=True))
dir(vbt.talib('BBANDS'))
# ## MA
print(close.rolling(2).mean())
print(close.ewm(span=3, min_periods=3).mean())
print(vbt.talib('SMA').run(close, timeperiod=2).real)
print(vbt.MA.run(close, [2, 3], ewm=[False, True]).ma) # adjust=False
# +
# One window
# %timeit big_close.rolling(2).mean() # pandas
# %timeit vbt.talib('SMA').run(big_close, timeperiod=2)
# %timeit vbt.MA.run(big_close, 2, return_cache=True) # cache only
# %timeit vbt.MA.run(big_close, 2) # with pre+postprocessing and still beats pandas
print(vbt.MA.run(big_close, 2).ma.shape)
# +
# Multiple windows
# %timeit pd.concat([big_close.rolling(i).mean() for i in np.arange(2, 10)])
# %timeit vbt.talib('SMA').run(big_close, np.arange(2, 10))
# %timeit vbt.MA.run(big_close, np.arange(2, 10))
# %timeit vbt.MA.run(big_close, np.arange(2, 10), run_unique=True)
# %timeit vbt.MA.run(big_close, np.arange(2, 10), return_cache=True) # cache only
cache = vbt.MA.run(big_close, np.arange(2, 10), return_cache=True)
# %timeit vbt.MA.run(big_close, np.arange(2, 10), use_cache=cache) # using cache
print(vbt.MA.run(big_close, np.arange(2, 10)).ma.shape)
# +
# One window repeated
# %timeit pd.concat([big_close.rolling(i).mean() for i in np.full(10, 2)])
# %timeit vbt.talib('SMA').run(big_close, np.full(10, 2))
# %timeit vbt.MA.run(big_close, np.full(10, 2))
# %timeit vbt.MA.run(big_close, np.full(10, 2), run_unique=True) # slower for large inputs
# %timeit vbt.MA.run(big_close, np.full(10, 2), return_cache=True)
print(vbt.MA.run(big_close, np.full(10, 2)).ma.shape)
# +
# %timeit pd.concat([big_close.iloc[:, :10].rolling(i).mean() for i in np.full(100, 2)])
# %timeit vbt.talib('SMA').run(big_close.iloc[:, :10], np.full(100, 2))
# %timeit vbt.MA.run(big_close.iloc[:, :10], np.full(100, 2))
# %timeit vbt.MA.run(big_close.iloc[:, :10], np.full(100, 2), run_unique=True) # faster for smaller inputs
# %timeit vbt.MA.run(big_close.iloc[:, :10], np.full(100, 2), return_cache=True)
print(vbt.MA.run(big_close.iloc[:, :10], np.full(100, 2)).ma.shape)
# +
ma = vbt.MA.run(close, [2, 3], ewm=[False, True])
print(ma.ma)
# -
ma[(2, False, 'a')].plot().show_svg()
# ## MSTD
print(close.rolling(2).std(ddof=0))
print(close.ewm(span=3, min_periods=3).std(ddof=0))
print(vbt.talib('STDDEV').run(close, timeperiod=2).real)
print(vbt.MSTD.run(close, [2, 3], ewm=[False, True]).mstd) # adjust=False, ddof=0
# +
# One window
# %timeit big_close.rolling(2).std()
# %timeit vbt.talib('STDDEV').run(big_close, timeperiod=2)
# %timeit vbt.MSTD.run(big_close, 2)
print(vbt.MSTD.run(big_close, 2).mstd.shape)
# +
# Multiple windows
# %timeit pd.concat([big_close.rolling(i).std() for i in np.arange(2, 10)])
# %timeit vbt.talib('STDDEV').run(big_close, timeperiod=np.arange(2, 10))
# %timeit vbt.MSTD.run(big_close, np.arange(2, 10))
print(vbt.MSTD.run(big_close, np.arange(2, 10)).mstd.shape)
# +
# One window repeated
# %timeit vbt.talib('STDDEV').run(big_close, timeperiod=np.full(10, 2))
# %timeit vbt.MSTD.run(big_close, window=np.full(10, 2))
print(vbt.MSTD.run(big_close, window=np.full(10, 2)).close.shape)
# +
mstd = vbt.MSTD.run(close, [2, 3], [False, True])
print(mstd.mstd)
# -
mstd[(2, False, 'a')].plot().show_svg()
# ## BBANDS
# +
print(vbt.ta('BollingerBands').run(close['a'], window=2, window_dev=2).bollinger_hband)
print(vbt.ta('BollingerBands').run(close['a'], window=2, window_dev=2).bollinger_mavg)
print(vbt.ta('BollingerBands').run(close['a'], window=2, window_dev=2).bollinger_lband)
print(vbt.talib('BBANDS').run(close, timeperiod=2, nbdevup=2, nbdevdn=2).upperband)
print(vbt.talib('BBANDS').run(close, timeperiod=2, nbdevup=2, nbdevdn=2).middleband)
print(vbt.talib('BBANDS').run(close, timeperiod=2, nbdevup=2, nbdevdn=2).lowerband)
print(vbt.BBANDS.run(close, window=2, ewm=False, alpha=2).upper)
print(vbt.BBANDS.run(close, window=2, ewm=False, alpha=2).middle)
print(vbt.BBANDS.run(close, window=2, ewm=False, alpha=2).lower)
# +
# One window
# %timeit vbt.talib('BBANDS').run(big_close, timeperiod=2)
# %timeit vbt.BBANDS.run(big_close, window=2)
print(vbt.BBANDS.run(big_close).close.shape)
# +
# Multiple windows
# %timeit vbt.talib('BBANDS').run(big_close, timeperiod=np.arange(2, 10))
# %timeit vbt.BBANDS.run(big_close, window=np.arange(2, 10))
print(vbt.BBANDS.run(big_close, window=np.arange(2, 10)).close.shape)
# +
# One window repeated
# %timeit vbt.talib('BBANDS').run(big_close, timeperiod=np.full(10, 2))
# %timeit vbt.BBANDS.run(big_close, window=np.full(10, 2))
print(vbt.BBANDS.run(big_close, window=np.full(10, 2)).close.shape)
# +
bb = vbt.BBANDS.run(close, window=2, alpha=[1., 2.], ewm=False)
print(bb.middle)
print()
print(bb.upper)
print()
print(bb.lower)
print()
print(bb.percent_b)
print()
print(bb.bandwidth)
# -
print(bb.close_below(bb.upper) & bb.close_above(bb.lower)) # price between bands
bb[(2, False, 1., 'a')].plot().show_svg()
# ## RSI
print(vbt.ta('RSIIndicator').run(close=close['a'], window=2).rsi) # alpha=1/n
print(vbt.ta('RSIIndicator').run(close=close['b'], window=2).rsi)
print(vbt.ta('RSIIndicator').run(close=close['c'], window=2).rsi)
print(vbt.talib('RSI').run(close, timeperiod=2).real)
print(vbt.RSI.run(close, window=[2, 2], ewm=[True, False]).rsi) # span=n
# +
# One window
# %timeit vbt.talib('RSI').run(big_close, timeperiod=2)
# %timeit vbt.RSI.run(big_close, window=2)
print(vbt.RSI.run(big_close, window=2).rsi.shape)
# +
# Multiple windows
# %timeit vbt.talib('RSI').run(big_close, timeperiod=np.arange(2, 10))
# %timeit vbt.RSI.run(big_close, window=np.arange(2, 10))
print(vbt.RSI.run(big_close, window=np.arange(2, 10)).rsi.shape)
# +
# One window repeated
# %timeit vbt.talib('RSI').run(big_close, timeperiod=np.full(10, 2))
# %timeit vbt.RSI.run(big_close, window=np.full(10, 2))
print(vbt.RSI.run(big_close, window=np.full(10, 2)).rsi.shape)
# +
rsi = vbt.RSI.run(close, window=[2, 3], ewm=[False, True])
print(rsi.rsi)
# -
print(rsi.rsi_above(70))
rsi[(2, False, 'a')].plot().show_svg()
# ## STOCH
print(vbt.ta('StochasticOscillator').run(high=high['a'], low=low['a'], close=close['a'], window=2, smooth_window=3).stoch)
print(vbt.ta('StochasticOscillator').run(high=high['a'], low=low['a'], close=close['a'], window=2, smooth_window=3).stoch_signal)
print(vbt.talib('STOCHF').run(high, low, close, fastk_period=2, fastd_period=3).fastk)
print(vbt.talib('STOCHF').run(high, low, close, fastk_period=2, fastd_period=3).fastd)
print(vbt.STOCH.run(high, low, close, k_window=2, d_window=3).percent_k)
print(vbt.STOCH.run(high, low, close, k_window=2, d_window=3).percent_d)
# +
# One window
# %timeit vbt.talib('STOCHF').run(big_high, big_low, big_close, fastk_period=2)
# %timeit vbt.STOCH.run(big_high, big_low, big_close, k_window=2)
print(vbt.STOCH.run(big_high, big_low, big_close, k_window=2).percent_d.shape)
# +
# Multiple windows
# %timeit vbt.talib('STOCHF').run(big_high, big_low, big_close, fastk_period=np.arange(2, 10))
# %timeit vbt.STOCH.run(big_high, big_low, big_close, k_window=np.arange(2, 10))
print(vbt.STOCH.run(big_high, big_low, big_close, k_window=np.arange(2, 10)).percent_d.shape)
# +
# One window repeated
# %timeit vbt.talib('STOCHF').run(big_high, big_low, big_close, fastk_period=np.full(10, 2))
# %timeit vbt.STOCH.run(big_high, big_low, big_close, k_window=np.full(10, 2))
print(vbt.STOCH.run(big_high, big_low, big_close, k_window=np.full(10, 2)).percent_d.shape)
# +
stochastic = vbt.STOCH.run(high, low, close, k_window=[2, 4], d_window=2, d_ewm=[False, True])
print(stochastic.percent_k)
print(stochastic.percent_d)
# -
stochastic[(2, 2, False, 'a')].plot().show_svg()
# ## MACD
# +
print(vbt.ta('MACD').run(close['a'], window_fast=2, window_slow=3, window_sign=2).macd)
print(vbt.ta('MACD').run(close['a'], window_fast=2, window_slow=3, window_sign=2).macd_signal)
print(vbt.ta('MACD').run(close['a'], window_fast=2, window_slow=3, window_sign=2).macd_diff)
print(vbt.talib('MACD').run(close, fastperiod=2, slowperiod=3, signalperiod=2).macd) # uses sma
print(vbt.talib('MACD').run(close, fastperiod=2, slowperiod=3, signalperiod=2).macdsignal)
print(vbt.talib('MACD').run(close, fastperiod=2, slowperiod=3, signalperiod=2).macdhist)
print(vbt.MACD.run(close, fast_window=2, slow_window=3, signal_window=2, macd_ewm=True, signal_ewm=True).macd)
print(vbt.MACD.run(close, fast_window=2, slow_window=3, signal_window=2, macd_ewm=True, signal_ewm=True).signal)
print(vbt.MACD.run(close, fast_window=2, slow_window=3, signal_window=2, macd_ewm=True, signal_ewm=True).hist)
# +
# One window
# %timeit vbt.talib('MACD').run(big_close, fastperiod=2)
# %timeit vbt.MACD.run(big_close, fast_window=2)
print(vbt.MACD.run(big_close, fast_window=2).macd.shape)
# +
# Multiple windows
# %timeit vbt.talib('MACD').run(big_close, fastperiod=np.arange(2, 10))
# %timeit vbt.MACD.run(big_close, fast_window=np.arange(2, 10))
print(vbt.MACD.run(big_close, fast_window=np.arange(2, 10)).macd.shape)
# +
# One window repeated
# %timeit vbt.talib('MACD').run(big_close, fastperiod=np.full(10, 2))
# %timeit vbt.MACD.run(big_close, fast_window=np.full(10, 2))
print(vbt.MACD.run(big_close, fast_window=np.full(10, 2)).macd.shape)
# +
macd = vbt.MACD.run(close, fast_window=2, slow_window=3, signal_window=[2, 3], macd_ewm=True, signal_ewm=True)
print(macd.macd)
print(macd.signal)
print(macd.hist)
# -
macd[(2, 3, 2, True, True, 'a')].plot().show_svg()
# ## ATR
print(vbt.ta('AverageTrueRange').run(high['a'], low['a'], close['a'], window=2).average_true_range)
print(vbt.ta('AverageTrueRange').run(high['b'], low['b'], close['b'], window=2).average_true_range)
print(vbt.ta('AverageTrueRange').run(high['c'], low['c'], close['c'], window=2).average_true_range)
print(vbt.talib('ATR').run(high, low, close, timeperiod=2).real)
print(vbt.ATR.run(high, low, close, window=[2, 3], ewm=[False, True]).atr)
# +
# One window
# %timeit vbt.talib('ATR').run(big_high, big_low, big_close, timeperiod=2)
# %timeit vbt.ATR.run(big_high, big_low, big_close, window=2)
print(vbt.ATR.run(big_high, big_low, big_close, window=2).atr.shape)
# +
# Multiple windows
# %timeit vbt.talib('ATR').run(big_high, big_low, big_close, timeperiod=np.arange(2, 10))
# %timeit vbt.ATR.run(big_high, big_low, big_close, window=np.arange(2, 10)) # rolling min/max very expensive
print(vbt.ATR.run(big_high, big_low, big_close, window=np.arange(2, 10)).atr.shape)
# +
# One window repeated
# %timeit vbt.talib('ATR').run(big_high, big_low, big_close, timeperiod=np.full(10, 2))
# %timeit vbt.ATR.run(big_high, big_low, big_close, window=np.full(10, 2))
print(vbt.ATR.run(big_high, big_low, big_close, window=np.full(10, 2)).atr.shape)
# +
atr = vbt.ATR.run(high, low, close, window=[2, 3], ewm=[False, True])
print(atr.tr)
print(atr.atr)
# -
atr[(2, False, 'a')].plot().show_svg()
# ## OBV
print(vbt.ta('OnBalanceVolumeIndicator').run(close['a'], volume['a']).on_balance_volume)
print(vbt.ta('OnBalanceVolumeIndicator').run(close['b'], volume['b']).on_balance_volume)
print(vbt.ta('OnBalanceVolumeIndicator').run(close['c'], volume['c']).on_balance_volume)
print(vbt.talib('OBV').run(close, volume).real)
print(vbt.OBV.run(close, volume).obv)
# +
# %timeit vbt.talib('OBV').run(big_close, big_volume)
# %timeit vbt.OBV.run(big_close, big_volume)
print(vbt.OBV.run(big_close, big_volume).obv.shape)
# +
obv = vbt.OBV.run(close, volume)
print(obv.obv)
# -
print(obv.obv_above([0, 5]))
obv['a'].plot().show_svg()
| tests/notebooks/indicators.ipynb |