code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python385jvsc74a57bd04e4b1c0eba25b43fb0828087952fe1bfe39b80460d2a5f528a72e68cadaca174
# ---
# # Visualizations
# +
# %matplotlib qt
from matplotlib import patches as mpatches
from collections import Counter
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_context("paper")
sns.set(font="Times New Roman", font_scale=1.50)
sns.set_style("whitegrid")
# load
csv_path = "./data/anonymized/tracks.csv"
df: pd.DataFrame = pd.read_csv(csv_path, sep=",")
df: pd.DataFrame = df.drop(
df[df["user"] == "participant_7"].index) # ignore participant 7
# -
# ## Motivation
# ### box__consumption_phase_participant
# +
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df["user"] = df["user"].map(lambda user: translation[user])
column_csv = "consumption"
column_csv_translation = "Priemerná spotreba paliva [L/100km]"
# picked data
df_boxplot: pd.DataFrame = df[["user", "phase", column_csv]]
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(
color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
sns.boxplot(x="user", y=column_csv, hue="phase", data=df_boxplot, palette="Set1", boxprops=dict(alpha=.3),
showfliers=False)
ax = sns.stripplot(x="user", y=column_csv, hue="phase",
data=df_boxplot, dodge=True, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Účastníci")
plt.ylabel(column_csv_translation)
fig.legend(title='Fázy', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### box__score_phase_participant
# +
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df["user"] = df["user"].map(lambda user: translation[user])
column_csv = "score"
column_csv_translation = "Ekologické skóre [-]"
# picked data
df_boxplot: pd.DataFrame = df[["user", "phase", column_csv]]
df_boxplot: pd.DataFrame = df_boxplot.drop(df_boxplot[df_boxplot["phase"] == 1].index)
# legend colours and names
legend_names = ["2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(
color='#477ca8')]
# plotting
fig = plt.figure()
sns.boxplot(x="user", y=column_csv, hue="phase", data=df_boxplot, palette="Set1", boxprops=dict(alpha=.3),
showfliers=False)
ax = sns.stripplot(x="user", y=column_csv, hue="phase",
data=df_boxplot, dodge=True, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Účastníci")
plt.ylabel(column_csv_translation)
fig.legend(title='Fázy', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### box__consumption_phase_strategy
# +
column_csv = "consumption"
column_csv_translation = "Priemerná spotreba paliva [L/100km]"
df_strategy = df[[column_csv, "phase", "strategy"]]
# df_strategy = df_strategy.drop(df_strategy[df_strategy["phase"] == 1].index)
# legend colours and names
legend_names = ["Odmeny", "Gamifikácia"]
legend_patches = [mpatches.Patch(
color='#cb3335'), mpatches.Patch(color='#477ca8')]
# plotting
fig = plt.figure()
sns.boxplot(x="phase", y=column_csv, hue="strategy", data=df_strategy, palette="Set1", boxprops=dict(alpha=.3),
showfliers=False)
ax = sns.stripplot(x="phase", y=column_csv, hue="strategy",
data=df_strategy, dodge=True, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Fáza")
plt.ylabel(column_csv_translation)
fig.legend(title='Motivačná stratégia', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### box__score_phase_strategy
# +
column_csv = "score"
column_csv_translation = "Ekologické skóre [-]"
df_strategy = df[[column_csv, "phase", "strategy"]]
df_strategy = df_strategy.drop(df_strategy[df_strategy["phase"] == 1].index)
# legend colours and names
legend_names = ["Odmeny", "Gamifikácia"]
legend_patches = [mpatches.Patch(
color='#cb3335'), mpatches.Patch(color='#477ca8')]
# plotting
fig = plt.figure()
sns.boxplot(x="phase", y=column_csv, hue="strategy", data=df_strategy, palette="Set1", boxprops=dict(alpha=.3),
showfliers=False)
ax = sns.stripplot(x="phase", y=column_csv, hue="strategy",
data=df_strategy, dodge=True, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Fáza")
plt.ylabel(column_csv_translation)
fig.legend(title='Motivačná stratégia', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### bar__consumption_phase_participant
# +
# picked data
column_csv = "consumption"
df_consumption: pd.DataFrame = df[["user", "phase", column_csv]]
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df_consumption["user"] = df_consumption["user"].map(lambda user: translation[user])
# grouping by user and phase -> mean
grouped = df_consumption.groupby(by=["user", "phase"], as_index=False).mean()
# sorting it, so the users are in order on chart
sort_keys = {f"P{i}": i for i in range(12)}
grouped["user_id"] = [sort_keys[user] for user in grouped.user]
grouped = grouped.sort_values(by="user_id")
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
ax = sns.barplot(x="user", y="consumption", hue="phase", data=grouped, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Participanti")
plt.ylabel("Priemerná spotreba paliva [L/100km]")
fig.legend(title='Fáza', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### bar__speed_phase_participant
# +
# picked data
column_csv = "speed"
df_speed: pd.DataFrame = df[["user", "phase", column_csv]]
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df_speed["user"] = df_speed["user"].map(lambda user: translation[user])
# grouping by user and phase -> mean
grouped = df_speed.groupby(by=["user", "phase"], as_index=False).mean()
# sorting it, so the users are in order on chart
sort_keys = {f"P{i}": i for i in range(12)}
grouped["user_id"] = [sort_keys[user] for user in grouped.user]
grouped = grouped.sort_values(by="user_id")
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
ax = sns.barplot(x="user", y="speed", hue="phase", data=grouped, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Participanti")
plt.ylabel("Priemerná rýchlosť [km/h]")
fig.legend(title='Fáza', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### box__time_phase_participant
# +
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df["user"] = df["user"].map(lambda user: translation[user])
# picked data
column_csv = "duration"
column_csv_translation = "Čas presunu [min]"
df_times: pd.DataFrame = df[["user", "phase", column_csv]]
df_times["duration"] /= 60
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(
color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
sns.boxplot(x="user", y=column_csv, hue="phase", data=df_times, palette="Set1", boxprops=dict(alpha=.3),
showfliers=False)
ax = sns.stripplot(x="user", y=column_csv, hue="phase",
data=df_times, dodge=True, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Participanti")
plt.ylabel(column_csv_translation)
fig.legend(title='Fázy', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### bar__time_phase_participant
# +
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df["user"] = df["user"].map(lambda user: translation[user])
# picked data
column_csv = "duration"
df_times: pd.DataFrame = df[["user", "phase", column_csv]]
df_times["duration"] /= 60
# grouping by user and phase -> mean
grouped = df_times.groupby(by=["user", "phase"], as_index=False).mean()
# sorting it, so the users are in order on chart
sort_keys = {f"P{i}": i for i in range(12)}
grouped["user_id"] = [sort_keys[user] for user in grouped.user]
grouped = grouped.sort_values(by="user_id")
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
ax = sns.barplot(x="user", y="duration", hue="phase", data=grouped, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Participanti")
plt.ylabel("Priemerný čas presunu [min]")
fig.legend(title='Fáza', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### bar__fuelconsumed_phase_participant
# +
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df["user"] = df["user"].map(lambda user: translation[user])
# picked data
column_csv = "fuelConsumed"
df_fuel: pd.DataFrame = df[["user", "phase", column_csv]]
# grouping by user and phase -> mean
grouped = df_fuel.groupby(by=["user", "phase"], as_index=False).sum()
# sorting it, so the users are in order on chart
sort_keys = {f"P{i}": i for i in range(12)}
grouped["user_id"] = [sort_keys[user] for user in grouped.user]
grouped = grouped.sort_values(by="user_id")
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
ax = sns.barplot(x="user", y="fuelConsumed", hue="phase", data=grouped, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Participanti")
plt.ylabel("Celkové spotrebované množstvo paliva [L]")
fig.legend(title='Fáza', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ## Rules
# ### bar__tracks_phase_participant
# +
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df["user"] = df["user"].map(lambda user: translation[user])
column_csv = "user"
df_counting = df[[column_csv, "phase"]]
counts_for_phases = {i: Counter(
df_counting[df_counting["phase"] == i][column_csv]) for i in range(1, 4)}
data = {
"Participant": [],
"Fáza": [],
"Počet": []
}
for phase in counts_for_phases.keys():
for user in counts_for_phases[phase].keys():
data["Participant"].append(user)
data["Fáza"].append(phase)
data["Počet"].append(counts_for_phases[phase][user])
df_counting = pd.DataFrame.from_dict(data)
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(
color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
ax = sns.barplot(x="Participant", y="Počet", hue="Fáza",
data=df_counting, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Účastníci")
plt.ylabel("Počet jázd [-]")
fig.legend(title='Fáza', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### bar__length_phase_participant
# +
# picked data
column_csv = "length"
df_length: pd.DataFrame = df[["user", "phase", column_csv]]
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df_length["user"] = df_length["user"].map(lambda user: translation[user])
# grouping by user and phase -> mean
grouped = df_length.groupby(by=["user", "phase"], as_index=False).mean()
# sorting it, so the users are in order on chart
sort_keys = {f"P{i}": i for i in range(12)}
grouped["user_id"] = [sort_keys[user] for user in grouped.user]
grouped = grouped.sort_values(by="user_id")
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
ax = sns.barplot(x="user", y="length", hue="phase", data=grouped, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Participanti")
plt.ylabel("Priemerná vzdialenosť presunu [min]")
fig.legend(title='Fáza', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ### box__length_phase_participant
# +
# translation of xticks
translation = {f"participant_{i}": f"P{i}" for i in range(12)}
df["user"] = df["user"].map(lambda user: translation[user])
column_csv = "length"
column_csv_translation = "Prejdená vzdialenosť [km]"
# picked data
df_boxplot: pd.DataFrame = df[["user", "phase", column_csv]]
# legend colours and names
legend_names = ["1", "2", "3"]
legend_patches = [mpatches.Patch(color='#cb3335'), mpatches.Patch(
color='#477ca8'), mpatches.Patch(color='#59a257')]
# plotting
fig = plt.figure()
sns.boxplot(x="user", y=column_csv, hue="phase", data=df_boxplot, palette="Set1", boxprops=dict(alpha=.3),
showfliers=False)
ax = sns.stripplot(x="user", y=column_csv, hue="phase",
data=df_boxplot, dodge=True, palette="Set1")
ax.get_legend().remove()
plt.xlabel("Účastníci")
plt.ylabel(column_csv_translation)
fig.legend(title='Fázy', labels=legend_names,
handles=legend_patches,
fancybox=True, shadow=True, loc='upper center', ncol=len(legend_patches))
plt.show()
# -
# ## Misc
# ### corr_matrix
# +
df_corr: pd.DataFrame = df[['consumption', 'duration', 'fuelConsumed', 'length', 'score', 'speed']]
f = plt.figure(figsize=(19, 15))
ax = sns.heatmap(df_corr.corr(), annot=True)
ax.set_title('Korelačná matica', fontsize=16)
del f, ax
# -
# ### tt__parametric
# +
from scipy.stats import ttest_ind, shapiro
import seaborn as sns
# control groups
rewards = df[df['strategy'] == 'rewards']
gamification = df[df['strategy'] == 'gamification']
# assumption of t-test - normal distribution of samples
print(shapiro(rewards['consumption']))
print(shapiro(gamification['consumption']))
# plot distribution
sns.displot(df, x="consumption", hue="strategy", multiple="dodge")
plt.legend(labels=["Gamifikácia", "Odmeny"], loc='upper center', fancybox=True)
plt.xlabel("Priemerná spotreba paliva [L/100km]")
plt.ylabel("Počet")
plt.show()
# ttest - independent samples of scores
print(ttest_ind(rewards['consumption'], gamification['consumption']))
# -
# ### avg_consumptions
# +
# control groups
rewards = df[df['strategy'] == 'rewards']
gamification = df[df['strategy'] == 'gamification']
print(rewards[rewards["phase"] == 1]['consumption'].mean())
print(rewards[rewards["phase"] == 1]['consumption'].std())
print(gamification[gamification["phase"] == 1]['consumption'].mean())
print(gamification[gamification["phase"] == 1]['consumption'].std())
print()
print(rewards[rewards["phase"] == 2]['consumption'].mean())
print(rewards[rewards["phase"] == 2]['consumption'].std())
print(gamification[gamification["phase"] == 2]['consumption'].mean())
print(gamification[gamification["phase"] == 2]['consumption'].std())
print()
print(rewards[rewards["phase"] == 3]['consumption'].mean())
print(rewards[rewards["phase"] == 3]['consumption'].std())
print(gamification[gamification["phase"] == 3]['consumption'].mean())
print(gamification[gamification["phase"] == 3]['consumption'].std())
# -
# ### avg_scores
# +
# control groups
rewards = df[df['strategy'] == 'rewards']
gamification = df[df['strategy'] == 'gamification']
print(rewards[rewards["phase"] == 1]['score'].mean())
print(rewards[rewards["phase"] == 1]['score'].std())
print(gamification[gamification["phase"] == 1]['score'].mean())
print(gamification[gamification["phase"] == 1]['score'].std())
print()
print(rewards[rewards["phase"] == 2]['score'].mean())
print(rewards[rewards["phase"] == 2]['score'].std())
print(gamification[gamification["phase"] == 2]['score'].mean())
print(gamification[gamification["phase"] == 2]['score'].std())
print()
print(rewards[rewards["phase"] == 3]['score'].mean())
print(rewards[rewards["phase"] == 3]['score'].std())
print(gamification[gamification["phase"] == 3]['score'].mean())
print(gamification[gamification["phase"] == 3]['score'].std())
# -
| data/dataset_mining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Author: <NAME>.
# # Assignment: Time Series
# In this task I have to make ARIMA model over shampoo salesdata and check the MSE between predicted and actual value.
#Load the required libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from statsmodels.tsa.arima_model import ARIMA
from sklearn.metrics import mean_squared_error
import datetime
def parser(x):
return datetime.strptime('190'+x, '%Y-%m')
series = pd.read_csv('sales-of-shampoo-over-a-three-ye.csv', header=0,parse_dates=True,index_col='Month')
series.head()
series.info()
series.isnull().any(), series.isna().sum()
#Delete null values
series = series.dropna()
#Visualization of actual data
series.plot()
plt.show()
#Converting to numpy array
sales = series.values
sales
size = int(len(sales) * 0.60)
print(f"Total length: {len(sales)}")
print(f"Training size: {size}")
#Train test split
train, test = sales[0:size], sales[size:len(sales)]
#ARIMA model
history = [x for x in train]
predictions = list()
for t in range(len(test)):
model = ARIMA(history,order=(5,1,0))
model_fit = model.fit(disp=0)
output = model_fit.forecast()
yhat = output[0]
predictions.append(yhat)
obs = test[t]
history.append(obs)
print(f'Predicted={yhat},Expected ={obs}')
error = mean_squared_error(test,predictions)
print("------------------------------------------")
print(f"TEST Mean Square Error :{error}")
#Visualization of Expected vs predicted data
plt.plot(test)
plt.plot(predictions, color='red')
plt.show()
| Time_Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import nltk
import numpy as np
import re
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english'))
from nltk.stem import WordNetLemmatizer
from nltk.stem import PorterStemmer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
import tensorflow as tf
from tensorflow.keras.layers import *
from tensorflow.keras.models import Sequential
from sklearn.model_selection import train_test_split
import sys
sys.setrecursionlimit(10000)
sys.getrecursionlimit()
train = pd.read_csv('train.csv')
# Splitting the data and creating randomness
x_train, x_test, y_train, y_test = train_test_split(train, train.iloc[:,-6:], shuffle = True,test_size=0.50, random_state = 123)
# dividing the data according to my ram size you may increase it accordint to your need
#my pc is crashing and raising memory error if i use full data
x_train = x_train[:5000]
y_train = y_train[:5000]
x_test = x_test[:1000]
y_test = y_test[:1000]
x_train.reset_index(inplace=True,drop=True)
x_test.reset_index(inplace=True,drop=True)
x_train.head()
# Stemming
stemmer = PorterStemmer()
for i,text in enumerate(x_train['comment_text']):
text = text.lower()
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "can not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub('\W', ' ', text)
text = re.sub('[^A-Za-z\' ]+', '',text)
text = re.sub('\s+', ' ', text)
words = text.split()
words = [stemmer.stem(word) for word in words if word not in set(stopwords.words('english'))]
x_train['comment_text'][i] = ' '.join(words)
x_train.head()
# Stemming
for i,text in enumerate(x_test['comment_text']):
text = text.lower()
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "can not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub('\W', ' ', text)
text = re.sub('[^A-Za-z\' ]+', '',text)
text = re.sub('\s+', ' ', text)
words = text.split()
words = [stemmer.stem(word) for word in words if word not in set(stopwords.words('english'))]
x_test['comment_text'][i] = ' '.join(words)
x_test.head()
# +
# # if pc is not good this will couse memory error
# ps = PorterStemmer()
# def clean_text(text):
# text = text.lower()
# text = re.sub(r"what's", "what is ", text) ### conversion of contraction words to expanded words
# text = re.sub(r"\'s", " ", text)
# text = re.sub(r"\'ve", " have ", text)
# text = re.sub(r"can't", "can not ", text)
# text = re.sub(r"n't", " not ", text)
# text = re.sub(r"i'm", "i am ", text)
# text = re.sub(r"\'re", " are ", text)
# text = re.sub(r"\'d", " would ", text)
# text = re.sub(r"\'ll", " will ", text)
# text = re.sub(r"\'scuse", " excuse ", text)
# text = re.sub('\W', ' ', text) ### removing non-word characters
# text = re.sub('[^A-Za-z\' ]+', '',text) ### removing all non-alphanumeric values(Except single quotes)
# text = re.sub('\s+', ' ', text)
# text = text.strip(' ')
# text = ' '.join([ps.stem(word) for word in text.split() if word not in (stop_words)]) ### Stopwords removal
# return text
# x_train["comment_text"] = train["comment_text"].apply(clean_text)
# # x_test["comment_text"] = test["comment_text"].apply(clean_text)
# -
# # Bag of Words with ngrams
cv = CountVectorizer(ngram_range=(2, 2),max_features=3000)
X = cv.fit_transform(x_train['comment_text']).toarray()
X = X.reshape(5000,3000,1)
Y = y_train.values
# preparing data for testing+
x_test = cv.transform(x_test['comment_text']).toarray()
x_test = x_test.reshape(1000,3000,1)
y_test = y_test.values
model = Sequential()
model.add(LSTM(units = 64, dropout = 0.2,input_shape=(3000,1),return_sequences=True))
model.add(LSTM(units = 64, dropout = 0.2))
model.add(Dense(units = 6, activation = 'sigmoid'))
model.summary()
model.compile(loss = "binary_crossentropy", optimizer = "adam", metrics = ["AUC"])
model.fit(X, Y, batch_size = 32, epochs = 1)
y_pred = model.predict(x_test)
y_pred
# +
#the output of y_pred is producing the 6 element array which is the probability of belong to particular label
# In this file we used bag of words with n grams range = 2
# train test split is used to divide the dataset with some randomness
# the reasono i choose less data is because my not have much memory to handle that amount of data
# you may use full data if your pc is good enought to handle the data..
# for further we can use word embeddings to improve the model (word2vec,embedding layer etc)
# -
# # MLP
nn = Sequential()
nn.add(Dense(16, input_dim=3000, activation='relu'))
nn.add(Dense(12, activation='relu'))
nn.add(Dense(6, activation='softmax'))
nn.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
nn.summary()
X = X.reshape(5000,3000)
tf.config.run_functions_eagerly(True)
nn.fit(X, Y, batch_size = 32, epochs = 1)
x_test = x_test.reshape(1000,3000)
pred_nn = nn.predict(x_test)
pred_nn
# # Question and Answer
# Q. Experimentation with different experiment setups:
#
# a. data pre-processing techniques – tokenise (e.g. will you use n-grams?), normalise text, apply stopwords, and so on
#
# Answer: For preprocessing first the regular expression is used to remove the special characters and symbols from the dataset then stopwords is applied to remove the unnessary words, after that the texxt is now ready for vectorization for this bag of word technique is used with bigram.
# Q. NLP algorithms and techniques – explain choice
#
# Answer: For classification of text first we experiment with LSTM model which is widely known for text classsification as it stores the memory state after it a simple neural network is used for classification which is very accurate. As after applying the bag of words (countvectorizer) the input feature become 3000 long so these two model are best for this much feature
# Q. text featurisation/transformation into numerical vectors – justify choices like one hot encoding, and other relevant methods
#
# Answer: the one hot encoding is not possible for this amount of data because we have huge corpus and using one hot vectors is not efficient in for this task. so we used bag of word for vectorization
# Q. training/text/validate dataset splitting – how did you split and why
#
# Answer: for this we use train_test_split from sklearn which provide randomness in the data after splitting
# Q. choices of loss functions and optimisers (if appropriate/relevant) – explain your choices with facts from the results
#
# Answer: As our problem is multi label classification problem so it is obvious to use categorical_crossentropy loss function because it is always used for this type of tassks and adam optimizer is best because it finds the convergence in very effictive manner.
| CommentClassification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Minimize beamwidth of an array with arbitrary 2-D geometry
#
# A derivative work by <NAME>, 5/14/2014.<br>
# Adapted (with significant changes) from the CVX example of the same name, by <NAME>, 2/2/2006.
#
#
# Topic References:
#
# * "Convex optimization examples" lecture notes (EE364) by <NAME>
# * "Antenna array pattern synthesis via convex optimization" by <NAME> and <NAME>
#
# ## Introduction
#
# This algorithm designs an antenna array such that:
#
# * it has unit sensitivity at some target direction
# * it obeys a constraint on a minimum sidelobe level outside the beam
# * it minimizes the beamwidth of the pattern.
#
# This is a quasiconvex problem. Define the target direction as $\theta_{\mbox{tar}}$, and a beamwidth of $\Delta \theta_{\mbox{bw}}$. The beam occupies the angular interval
#
# $$\Theta_b = \left(\theta_{\mbox{tar}}
# -\frac{1}{2}\Delta \theta_{\mbox{bw}},\; \theta_{\mbox{tar}}
# + \frac{1}{2}\Delta \theta_{\mbox{bw}}\right).
# $$
#
# Solving for the minimum beamwidth $\Delta \theta_{\mbox{bw}}$ is performed by bisection, where the interval which contains the optimal value is bisected according to the result of the following feasibility problem:
#
# \begin{array}{ll}
# \mbox{minimize} & 0 \\
# \mbox{subject to} & y(\theta_{\mbox{tar}}) = 1 \\
# & \left|y(\theta)\right| \leq t_{\mbox{sb}}
# \quad \forall \theta \notin \Theta_b.
# \end{array}
#
# $y$ is the antenna array gain pattern (a complex-valued function), $t_{\mbox{sb}}$ is the maximum allowed sideband gain threshold, and the variables are $w$ (antenna array weights or shading coefficients). The gain pattern is a linear function of $w$: $y(\theta) = w^T a(\theta)$ for some $a(\theta)$ describing the antenna array configuration and specs.
#
# Once the optimal beamwidth is found, the solution $w$ is refined with the following optimization:
#
# \begin{array}{ll}
# \mbox{minimize} & \|w\| \\
# \mbox{subject to} & y(\theta_{\mbox{tar}}) = 1 \\
# & \left|y(\theta)\right| \leq t_{\mbox{sb}}
# \quad \forall \theta \notin \Theta_b.
# \end{array}
#
# The implementation below discretizes the angular quantities and their counterparts, such as $\theta$.
#
# ## Problem specification and data
#
# ### Antenna array selection
#
# Choose either:
#
# * A random 2D positioning of antennas.
# * A uniform 1D positioning of antennas along a line.
# * A uniform 2D positioning of antennas along a grid.
# +
import cvxpy as cvx
import numpy as np
# Select array geometry:
ARRAY_GEOMETRY = '2D_RANDOM'
#ARRAY_GEOMETRY = '1D_UNIFORM_LINE'
#ARRAY_GEOMETRY = '2D_UNIFORM_LATTICE'
# -
# ## Data generation
# +
#
# Problem specs.
#
lambda_wl = 1 # wavelength
theta_tar = 60 # target direction
min_sidelobe = -20 # maximum sidelobe level in dB
max_half_beam = 50 # starting half beamwidth (must be feasible)
#
# 2D_RANDOM:
# n randomly located elements in 2D.
#
if ARRAY_GEOMETRY == '2D_RANDOM':
# Set random seed for repeatable experiments.
np.random.seed(1)
# Uniformly distributed on [0,L]-by-[0,L] square.
n = 36
L = 5
loc = L*np.random.random((n,2))
#
# 1D_UNIFORM_LINE:
# Uniform 1D array with n elements with inter-element spacing d.
#
elif ARRAY_GEOMETRY == '1D_UNIFORM_LINE':
n = 30
d = 0.45*lambda_wl
loc = np.hstack(( d * np.matrix(range(0,n)).T, \
np.zeros((n,1)) ))
#
# 2D_UNIFORM_LATTICE:
# Uniform 2D array with m-by-m element with d spacing.
#
elif ARRAY_GEOMETRY == '2D_UNIFORM_LATTICE':
m = 6
n = m**2
d = 0.45*lambda_wl
loc = np.matrix(np.zeros((n, 2)))
for x in range(m):
for y in range(m):
loc[m*y+x,:] = [x,y]
loc = loc*d
else:
raise Exception('Undefined array geometry')
#
# Construct optimization data.
#
# Build matrix A that relates w and y(theta), ie, y = A*w.
theta = np.mat(range(1, 360+1)).T
A = np.kron(np.cos(np.pi*theta/180), loc[:, 0].T) \
+ np.kron(np.sin(np.pi*theta/180), loc[:, 1].T)
A = np.exp(2*np.pi*1j/lambda_wl*A)
# Target constraint matrix.
ind_closest = np.argmin(np.abs(theta - theta_tar))
Atar = A[ind_closest,:]
# -
# ## Solve using bisection algorithm
# +
# Bisection range limits. Reduce by half each step.
halfbeam_bot = 1
halfbeam_top = max_half_beam
print 'We are only considering integer values of the half beam-width'
print '(since we are sampling the angle with 1 degree resolution).'
print
# Iterate bisection until 1 angular degree of uncertainty.
while halfbeam_top - halfbeam_bot > 1:
# Width in degrees of the current half-beam.
halfbeam_cur = np.ceil( (halfbeam_top + halfbeam_bot)/2.0 )
# Create optimization matrices for the stopband,
# i.e. only A values for the stopband angles.
ind = np.nonzero(np.squeeze(np.array(np.logical_or( \
theta <= (theta_tar-halfbeam_cur), \
theta >= (theta_tar+halfbeam_cur) ))))
As = A[ind[0],:]
#
# Formulate and solve the feasibility antenna array problem.
#
# As of this writing (2014/05/14) cvxpy does not do complex valued math,
# so the real and complex values must be stored seperately as reals
# and operated on as follows:
# Let any vector or matrix be represented as a+bj, or A+Bj.
# Vectors are stored [a; b] and matrices as [A -B; B A]:
# Atar as [A -B; B A]
Atar_R = Atar.real
Atar_I = Atar.imag
neg_Atar_I = -Atar_I
Atar_RI = np.bmat('Atar_R neg_Atar_I; Atar_I Atar_R')
# As as [A -B; B A]
As_R = As.real
As_I = As.imag
neg_As_I = -As_I
As_RI = np.bmat('As_R neg_As_I; As_I As_R')
As_RI_top = np.bmat('As_R neg_As_I')
As_RI_bot = np.bmat('As_I As_R')
# 1-vector as [1; 0] since no imaginary part
realones_ri = np.mat( np.vstack( \
(np.ones(Atar.shape[0]),
np.zeros(Atar.shape[0])) ))
# Create cvxpy variables and constraints
w_ri = cvx.Variable(shape=(2*n,1))
constraints = [ Atar_RI*w_ri == realones_ri]
# Must add complex valued constraint
# abs(As*w <= 10**(min_sidelobe/20)) row by row by hand.
# TODO: Future version use norms() or complex math
# when these features become available in cvxpy.
for i in range(As.shape[0]):
#Make a matrix whos product with w_ri is a 2-vector
#which is the real and imag component of a row of As*w
As_ri_row = np.vstack((As_RI_top[i, :], As_RI_bot[i, :]))
constraints.append( \
cvx.norm(As_ri_row*w_ri) <= 10**(min_sidelobe/20) )
# Form and solve problem.
obj = cvx.Minimize(0)
prob = cvx.Problem(obj, constraints)
prob.solve(solver=cvx.CVXOPT)
# Bisection (or fail).
if prob.status == cvx.OPTIMAL:
print ('Problem is feasible for half beam-width = {}'
' degress').format(halfbeam_cur)
halfbeam_top = halfbeam_cur
elif prob.status == cvx.INFEASIBLE:
print ('Problem is not feasible for half beam-width = {}'
' degress').format(halfbeam_cur)
halfbeam_bot = halfbeam_cur
else:
raise Exception('CVXPY Error')
# Optimal beamwidth.
halfbeam = halfbeam_top
print 'Optimum half beam-width for given specs is {}'.format(halfbeam)
# Compute the minimum noise design for the optimal beamwidth
ind = np.nonzero(np.squeeze(np.array(np.logical_or( \
theta <= (theta_tar-halfbeam), \
theta >= (theta_tar+halfbeam) ))))
As = A[ind[0],:]
# As as [A -B; B A]
# See earlier calculations for real/imaginary representation
As_R = As.real
As_I = As.imag
neg_As_I = -As_I
As_RI = np.bmat('As_R neg_As_I; As_I As_R')
As_RI_top = np.bmat('As_R neg_As_I')
As_RI_bot = np.bmat('As_I As_R')
constraints = [ Atar_RI*w_ri == realones_ri]
# Same constraint as a above, on new As (hense different
# actual number of constraints). See comments above.
for i in range(As.shape[0]):
As_ri_row = np.vstack((As_RI_top[i, :], As_RI_bot[i, :]))
constraints.append( \
cvx.norm(As_ri_row*w_ri) <= 10**(min_sidelobe/20) )
# Form and solve problem.
# Note the new objective!
obj = cvx.Minimize(cvx.norm(w_ri))
prob = cvx.Problem(obj, constraints)
prob.solve(solver=cvx.SCS)
#if prob.status != cvx.OPTIMAL:
# raise Exception('CVXPY Error')
# -
# ## Result plots
# +
import matplotlib.pyplot as plt
# Show plot inline in ipython.
# %matplotlib inline
# Plot properties.
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
#
# First Figure: Antenna Locations
#
plt.figure(figsize=(6, 6))
plt.scatter(np.array(loc[:, 0]), np.array(loc[:, 1]), \
s=30, facecolors='none', edgecolors='b')
plt.title('Antenna Locations', fontsize=16)
plt.tight_layout()
plt.show()
#
# Second Plot: Array Pattern
#
# Complex valued math to calculate y = A*w_im;
# See comments in code above regarding complex representation as reals.
A_R = A.real
A_I = A.imag
neg_A_I = -A_I
A_RI = np.bmat('A_R neg_A_I; A_I A_R');
y = A_RI*w_ri.value
y = y[0:y.shape[0]/2] + 1j*y[y.shape[0]/2:] #now native complex
plt.figure(figsize=(6,6))
ymin, ymax = -40, 0
plt.plot(np.arange(360)+1, np.array(20*np.log10(np.abs(y))))
plt.plot([theta_tar, theta_tar], [ymin, ymax], 'g--')
plt.plot([theta_tar+halfbeam, theta_tar+halfbeam], [ymin, ymax], 'r--')
plt.plot([theta_tar-halfbeam, theta_tar-halfbeam], [ymin, ymax], 'r--')
plt.xlabel('look angle', fontsize=16)
plt.ylabel(r'mag $y(\theta)$ in dB', fontsize=16)
plt.ylim(ymin, ymax)
plt.tight_layout()
plt.show()
#
# Third Plot: Polar Pattern
#
plt.figure(figsize=(6,6))
zerodB = 50
dBY = 20*np.log10(np.abs(y)) + zerodB
plt.plot(np.array(dBY)*np.array(np.cos(np.pi*theta/180)), \
np.array(dBY)*np.array(np.sin(np.pi*theta/180)))
plt.xlim(-zerodB, zerodB)
plt.ylim(-zerodB, zerodB)
plt.axis('off')
# 0 dB level.
plt.plot(zerodB*np.array(np.cos(np.pi*theta/180)), \
zerodB*np.array(np.sin(np.pi*theta/180)), 'k:')
plt.text(-zerodB,0,'0 dB', fontsize=16)
# Max sideband level.
m=min_sidelobe + zerodB
plt.plot(m*np.array(np.cos(np.pi*theta/180)), \
m*np.array(np.sin(np.pi*theta/180)), 'k:')
plt.text(-m,0,'{:.1f} dB'.format(min_sidelobe), fontsize=16)
#Lobe center and boundaries angles.
theta_1 = theta_tar+halfbeam
theta_2 = theta_tar-halfbeam
plt.plot([0, 55*np.cos(theta_tar*np.pi/180)], \
[0, 55*np.sin(theta_tar*np.pi/180)], 'k:')
plt.plot([0, 55*np.cos(theta_1*np.pi/180)], \
[0, 55*np.sin(theta_1*np.pi/180)], 'k:')
plt.plot([0, 55*np.cos(theta_2*np.pi/180)], \
[0, 55*np.sin(theta_2*np.pi/180)], 'k:')
#Show plot.
plt.tight_layout()
plt.show()
| examples/notebooks/WWW/ant_array_min_beamwidth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Wasserstein Discriminant Analysis
#
#
# This example illustrate the use of WDA as proposed in [11].
#
#
# [11] <NAME>., <NAME>., <NAME>., & <NAME>. (2016).
# Wasserstein Discriminant Analysis.
#
# +
# Author: <NAME> <<EMAIL>>
#
# License: MIT License
import numpy as np
import matplotlib.pylab as pl
from ot.dr import wda, fda
# -
# Generate data
# -------------
#
#
# +
n = 1000 # nb samples in source and target datasets
nz = 0.2
# generate circle dataset
t = np.random.rand(n) * 2 * np.pi
ys = np.floor((np.arange(n) * 1.0 / n * 3)) + 1
xs = np.concatenate(
(np.cos(t).reshape((-1, 1)), np.sin(t).reshape((-1, 1))), 1)
xs = xs * ys.reshape(-1, 1) + nz * np.random.randn(n, 2)
t = np.random.rand(n) * 2 * np.pi
yt = np.floor((np.arange(n) * 1.0 / n * 3)) + 1
xt = np.concatenate(
(np.cos(t).reshape((-1, 1)), np.sin(t).reshape((-1, 1))), 1)
xt = xt * yt.reshape(-1, 1) + nz * np.random.randn(n, 2)
nbnoise = 8
xs = np.hstack((xs, np.random.randn(n, nbnoise)))
xt = np.hstack((xt, np.random.randn(n, nbnoise)))
# -
# Plot data
# ---------
#
#
# +
pl.figure(1, figsize=(6.4, 3.5))
pl.subplot(1, 2, 1)
pl.scatter(xt[:, 0], xt[:, 1], c=ys, marker='+', label='Source samples')
pl.legend(loc=0)
pl.title('Discriminant dimensions')
pl.subplot(1, 2, 2)
pl.scatter(xt[:, 2], xt[:, 3], c=ys, marker='+', label='Source samples')
pl.legend(loc=0)
pl.title('Other dimensions')
pl.tight_layout()
# -
# Compute Fisher Discriminant Analysis
# ------------------------------------
#
#
# +
p = 2
Pfda, projfda = fda(xs, ys, p)
# -
# Compute Wasserstein Discriminant Analysis
# -----------------------------------------
#
#
# +
p = 2
reg = 1e0
k = 10
maxiter = 100
Pwda, projwda = wda(xs, ys, p, reg, k, maxiter=maxiter)
# -
# Plot 2D projections
# -------------------
#
#
# +
xsp = projfda(xs)
xtp = projfda(xt)
xspw = projwda(xs)
xtpw = projwda(xt)
pl.figure(2)
pl.subplot(2, 2, 1)
pl.scatter(xsp[:, 0], xsp[:, 1], c=ys, marker='+', label='Projected samples')
pl.legend(loc=0)
pl.title('Projected training samples FDA')
pl.subplot(2, 2, 2)
pl.scatter(xtp[:, 0], xtp[:, 1], c=ys, marker='+', label='Projected samples')
pl.legend(loc=0)
pl.title('Projected test samples FDA')
pl.subplot(2, 2, 3)
pl.scatter(xspw[:, 0], xspw[:, 1], c=ys, marker='+', label='Projected samples')
pl.legend(loc=0)
pl.title('Projected training samples WDA')
pl.subplot(2, 2, 4)
pl.scatter(xtpw[:, 0], xtpw[:, 1], c=ys, marker='+', label='Projected samples')
pl.legend(loc=0)
pl.title('Projected test samples WDA')
pl.tight_layout()
pl.show()
| _downloads/65e6bca00e3e928055316164cb5aede0/plot_WDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" deletable=false editable=false id="JdFUQNA5XUbn"
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS109B Data Science 2: Advanced Topics in Data Science
#
# ## Homework 4: CNN
#
# **Harvard University**<br/>
# **Spring 2020**<br/>
# **Instructors:** <NAME>, <NAME>, <NAME><br/>
#
# <hr style="height:2pt">
# + colab={"base_uri": "https://localhost:8080/", "height": 17} colab_type="code" deletable=false editable=false id="cKWDlL0JXUbs" outputId="642bd4a8-ebb6-415f-e935-337ee5ccaef6"
#RUN THIS CELL
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
# + deletable=false editable=false
#RUN THIS CELL
import os
import pathlib
working_dir = pathlib.Path().absolute()
# Uncomment the line below to help debug if the path to included images don't show
#print(working_dir)
os.chdir(working_dir)
# + [markdown] deletable=false editable=false
# <hr style="height:2pt">
#
# ### INSTRUCTIONS
#
# - To submit your assignment follow the instructions given in Canvas.
#
# - This homework can be submitted in pairs.
#
# - If you submit individually but you have worked with someone, please include the name of your **one** partner below.
# - Please restart the kernel and run the entire notebook again before you submit. (Exception - you may skip the cells where you train neural networks, running the cells which load previously saved weights instead. However, **don't delete/overwrite the output that model.fit produced during training!**)
#
# **Names of person you have worked with goes here:**
# <br><BR>
#
# <hr style="height:2pt">
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" deletable=false editable=false id="hakVc8z8aGmt" outputId="8ad99d3a-22a6-4509-a417-c679a16a50ca"
import numpy as np
from PIL import Image
from matplotlib import pyplot
import matplotlib.pylab as plt
from scipy.signal import convolve2d
# %matplotlib inline
import tensorflow as tf
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import Dense, Dropout, Flatten, Activation
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.optimizers import Adam, SGD
## Additional Packages required if you don't already have them
# While in your conda environment,
# imageio
# Install using "conda install imageio"
# pillow
# Install using "conda install pillow"
# tensorflow-datasets
# Install using "conda install tensorflow-datasets"
# tf-keras-vis
# Install using "pip install tf-keras-vis"
# tensorflow-addons
# Install using "pip install tensorflow-addons"
# + [markdown] colab_type="text" deletable=false editable=false id="rUkgUGwJXUcH"
# <div class="theme"> Overview </div>
#
#
# In this homework, we will explore Convolutional Neural Networks (CNNs). We'll explore the mechanics of convolutional operations and how filters can extract certain features of images, increasing in abstraction with depth. Then we will build a CNN to classify CIFAR-10 images, a standard pedagogical problem, and use saliency maps to understand what the network is paying attention to. Finally, we will see that CNNs aren't just for classifying. They can serve as image input processing for a variety of tasks, as we will show by training a network to rotate faces upright.
# + [markdown] deletable=false editable=false
# <div class='exercise'> <b> Question 1: Convolutional Neural Network Mechanics [10pts total] </b></div>
#
#
# As you know from lecture, in convolutional neural networks, a convolution is a multiplicative operation on a local region of values. Convolutional layers have shown themselves to have been very useful in image classification, as they allows the network to retain local spatial information for feature extraction.
#
#
# **1.1** Calculate Convolutions. [5pts]
#
#
#
#
# For the following 2D matrix:
#
# $$
# \left( \begin{array}{cccc}
# 2 & 3 & 2 & 4 \\
# 3 & 1 & 2 & 2 \\
# 4 & 1 & 0 & 1 \\
# 7 & 2 & 1 & 3
# \end{array} \right)
# $$
#
# you will use the following 2x2 filter to perform a 2D convolution operation.
#
# $$
# \left( \begin{array}{cc}
# 2 & 1 \\
# 1 & 3
# \end{array} \right)
# $$
#
# Compute this operation by hand assuming a vertical and horizontal stride of 1 as well as a) valid, b) same, and c) full padding modes.
# + [markdown] deletable=false editable=false
# **You may answer question 1.1 in this markdown cell by replacing the '?' marks with the correct value.**
#
#
#
# A) Valid
#
# $$
# \left( \begin{array}{cccc}
# ? & ? & ? \\
# ? & ? & ? \\
# ? & ? & ?
# \end{array} \right)
# $$
#
# B) Same padding. We will accept solutions for all combinations (top & left, top & right, bottom & left, bottom & right).
#
# $$
# \left( \begin{array}{cccc}
# ? & ? & ? & ? \\
# ? & ? & ? & ? \\
# ? & ? & ? & ? \\
# ? & ? & ? & ?
# \end{array} \right)
# $$
#
# C) full padding
#
# $$
# \left( \begin{array}{cccc}
# ? & ? & ? & ? & ? \\
# ? & ? & ? & ? & ? \\
# ? & ? & ? & ? & ? \\
# ? & ? & ? & ? & ? \\
# ? & ? & ? & ? & ?
# \end{array} \right)
# $$
# + [markdown] deletable=false editable=false
# **1.2** Understanding Pooling Operations. [5pts]
#
# Pooling operations are often used in convolutional neural networks to reduce the dimensionality of the feature maps as well as overall network complexity. Two main types of pooling are used: AveragePooling and MaxPooling.
#
# Using the matrix below, write the output of the AveragePooling and MaxPooling operations with pool size 2x2 and stride 2x2. Repeat with a stride of 1x1.
#
# $$
# \left( \begin{array}{cccc}
# 1 & 2 & 2 & 4 \\
# 3 & 1 & 2 & 1 \\
# 4 & 1 & 0 & 2 \\
# 5 & 2 & 2 & 1
# \end{array} \right)
# $$
# + [markdown] deletable=false editable=false
# **You may answer question 1.2 in this markdown cell by replacing the '?' marks with the correct value.**
#
# **A) size 2x2 and stride 2x2**
#
# MaxPooling:
# $$
# \left( \begin{array}{cccc}
# ? & ? \\
# ? & ?
# \end{array} \right)
# $$
# AveragePooling:
# $$
# \left( \begin{array}{cccc}
# ? & ? \\
# ? & ?
# \end{array} \right)
# $$
#
# **B) size 2x2 and stride 1x1**
#
# MaxPooling:
# $$
# \left( \begin{array}{cccc}
# ? & ? & ? \\
# ? & ? & ? \\
# ? & ? & ?
# \end{array} \right)
# $$
# AveragePooling:
# $$
# \left( \begin{array}{cccc}
# ? & ? & ? \\
# ? & ? & ? \\
# ? & ? & ?
# \end{array} \right)
# $$
# + [markdown] deletable=false editable=false
# ## Answers
# + [markdown] autograde="1.1" deletable=false editable=false
# **1.1** Calculate Convolutions. [5pts]
#
#
#
#
# For the following 2D matrix:
#
# $$
# \left( \begin{array}{cccc}
# 2 & 3 & 2 & 4 \\
# 3 & 1 & 2 & 2 \\
# 4 & 1 & 0 & 1 \\
# 7 & 2 & 1 & 3
# \end{array} \right)
# $$
#
# you will use the following 2x2 filter to perform a 2D convolution operation.
#
# $$
# \left( \begin{array}{cc}
# 2 & 1 \\
# 1 & 3
# \end{array} \right)
# $$
#
# Compute this operation by hand assuming a vertical and horizontal stride of 1 as well as a) valid, b) same, and c) full padding modes.
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] autograde="1.2" deletable=false editable=false
# **1.2** Understanding Pooling Operations. [5pts]
#
# Pooling operations are often used in convolutional neural networks to reduce the dimensionality of the feature maps as well as overall network complexity. Two main types of pooling are used: AveragePooling and MaxPooling.
#
# Using the matrix below, write the output of the AveragePooling and MaxPooling operations with pool size 2x2 and stride 2x2. Repeat with a stride of 1x1.
#
# $$
# \left( \begin{array}{cccc}
# 1 & 2 & 2 & 4 \\
# 3 & 1 & 2 & 1 \\
# 4 & 1 & 0 & 2 \\
# 5 & 2 & 2 & 1
# \end{array} \right)
# $$
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] colab_type="text" deletable=false editable=false id="JCGL_ur7VIbj"
# <div class='exercise'> <b> Question 2: CNNs at Work [30pts total] </b></div>
# + [markdown] colab_type="text" deletable=false editable=false id="YARDL5cVVIbl"
# Consider the following image of Widener Library:
#
# 
#
# **2.1** [2pts] Load the image as a 2D Numpy array into the variable `library_image_data`. Normalize the image data so that values within `library_image_data` fall within [0., 1.]. The image is located at 'data/Widener_Library.jpg'.
#
# **2.2** Filters for image processing. [5pts] Peform sharpening and normalized box blurring using 3x3 convolution kernels (see https://en.wikipedia.org/wiki/Kernel_(image_processing) for example), and apply each of these kernels to the image (separately on each color channel) with same padding mode (you may want to implement your own convolution function or try using [scipy.signal.convolve2d](https://docs.scipy.org/doc/scipy/reference/generated/scipy.signal.convolve2d.html)). Show the resulting 3-channel color images, using `np.clip()` to clip values to the range [0., 1.] for use with `imshow`.
#
# **2.3** What a CNN sees. [5pts] Normalize `library_image_data` to the range [-0.5, 0.5], saving the resulting array to `norm_img`. Consider the following 3x3x3 kernel $A_{ijk}$ where
# $$
# A_{:,:,1} = A_{:,:,2} = A_{:,:,3} = \frac{1}{3} \left( \begin{array}{ccc}
# -1 & -1 & -1 \\
# -1 & 8 & -1 \\
# -1 & -1 & -1 \\
# \end{array} \right),
# $$
# i.e. it has the same 3x3 depth slice throughout its depth (3 channels for the RGB channels of the input image). Apply $A_{ijk}$ to `norm_img` like a CNN would, using same padding mode (hint: what should the resulting shape of the feature map be?).
#
# After the convolution part, CNNs next need an activation function. We now *rectify* the feature map by applying the ReLU activation function:
#
# `
# if x < 0:
# x = 0
# else:
# x = x
# `
#
# or equivalently, $\textrm{ReLU}(x) = \textrm{max}(0,x)$ as it is often written. You can use `np.clip(x, a_min=0., a_max=None)` as a rectifier. Plot the rectified feature map using `imshow`, using the option `cmap='gray'` to produce a nice black & white image. What is this kernel doing?
#
#
# **2.4** [5pts] Look up or come up with a 3x3x3 kernel for performing vertical edge detection, and another for performing horizontal edge detection. Apply the vertical edge kernel to `norm_img`, and then through a ReLU, saving the result as `vedges`. Apply the horizontal edge kernel to `norm_img`, pass it through a ReLU and save the result as `hedges`. Plot `vedges` and `hedges` with `imshow` in black & white. Don't worry too much about what kernels you end up using, or what overall normalization factor you use. As long as in the plot vertical(horizontal) edges are clearly emphasized while horizontal(vertical) edges are suppressed, you've done it correctly.
#
# Together, `vedges` and `hedges` could be the output of the first layer of a CNN. Now we will investigate what can happen when we stack CNNs.
#
# **2.5** [8pts] Concatenate `vedges` and `hedges` in a third dimension, calling the output `feature_map`. `feature_map` should have dimensions (267, 400, 2). Take the following 3x3x2 kernel $B_{ijk}$:
#
# $$
# B_{:,:,1} = B_{:,:,2} = \left( \begin{array}{ccc}
# 0 & 0 & 0 \\
# 0 & 1 & 0 \\
# 0 & 0 & 0 \\
# \end{array} \right),
# $$
# and apply it to `feature_map`. This time, before we pass it through the activation, we will add a bias. For now, start with a bias of `bias = -2`, and pass the the result through a ReLU, saving the output in variable `outmap`. Plot `outmap` in black & white. Depending on the normalization of your vertical/horizontal kernels, you will have to play with the `bias` until most of the image is black, except for some shapes that should pop out (you'll still have dots of white in other places). Now that the image has passed through 2 CNN layers, what feature(s) does this latest layer seem to be picking out? (Open-ended question, but there are wrong answers. Think about what $B_{ijk}$ is doing, in combination with the bias and rectifier)
#
# **2.6** [5pts] Take a moment to think about the results of question 2.5. What seems to be the purpose of adding more CNN layers to increase the depth of a network? Why might it be useful to have multiple kernels/filters in a layer? Answer in 3-4 sentences.
#
# + [markdown] deletable=false editable=false
# ## Answers
# + [markdown] autograde="2.1" deletable=false editable=false
# **2.1** [2pts] Load the image as a 2D Numpy array into the variable `library_image_data`. Normalize the image data so that values within `library_image_data` fall within [0., 1.]. The image is located at 'data/Widener_Library.jpg'.
#
# + deletable=false
# your code here
# + deletable=false editable=false
# + [markdown] autograde="2.2" deletable=false editable=false
# **2.2** Filters for image processing. [5pts] Peform sharpening and normalized box blurring using 3x3 convolution kernels (see https://en.wikipedia.org/wiki/Kernel_(image_processing) for example), and apply each of these kernels to the image (separately on each color channel) with same padding mode (you may want to implement your own convolution function or try using [scipy.signal.convolve2d](https://docs.scipy.org/doc/scipy/reference/generated/scipy.signal.convolve2d.html)). Show the resulting 3-channel color images, using `np.clip()` to clip values to the range [0., 1.] for use with `imshow`.
#
# + deletable=false
# your code here
# + [markdown] autograde="2.3" deletable=false editable=false
# **2.3** What a CNN sees. [5pts] Normalize `library_image_data` to the range [-0.5, 0.5], saving the resulting array to `norm_img`. Consider the following 3x3x3 kernel $A_{ijk}$ where
# $$
# A_{:,:,1} = A_{:,:,2} = A_{:,:,3} = \frac{1}{3} \left( \begin{array}{ccc}
# -1 & -1 & -1 \\
# -1 & 8 & -1 \\
# -1 & -1 & -1 \\
# \end{array} \right),
# $$
# i.e. it has the same 3x3 depth slice throughout its depth (3 channels for the RGB channels of the input image). Apply $A_{ijk}$ to `norm_img` like a CNN would, using same padding mode (hint: what should the resulting shape of the feature map be?).
#
# After the convolution part, CNNs next need an activation function. We now *rectify* the feature map by applying the ReLU activation function:
#
# `
# if x < 0:
# x = 0
# else:
# x = x
# `
#
# or equivalently, $\textrm{ReLU}(x) = \textrm{max}(0,x)$ as it is often written. You can use `np.clip(x, a_min=0., a_max=None)` as a rectifier. Plot the rectified feature map using `imshow`, using the option `cmap='gray'` to produce a nice black & white image. What is this kernel doing?
#
#
# + deletable=false
# your code here
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] autograde="2.4" deletable=false editable=false
# **2.4** [5pts] Look up or come up with a 3x3x3 kernel for performing vertical edge detection, and another for performing horizontal edge detection. Apply the vertical edge kernel to `norm_img`, and then through a ReLU, saving the result as `vedges`. Apply the horizontal edge kernel to `norm_img`, pass it through a ReLU and save the result as `hedges`. Plot `vedges` and `hedges` with `imshow` in black & white. Don't worry too much about what kernels you end up using, or what overall normalization factor you use. As long as in the plot vertical(horizontal) edges are clearly emphasized while horizontal(vertical) edges are suppressed, you've done it correctly.
#
# Together, `vedges` and `hedges` could be the output of the first layer of a CNN. Now we will investigate what can happen when we stack CNNs.
#
# + deletable=false
# your code here
# + [markdown] autograde="2.5" deletable=false editable=false
# **2.5** [8pts] Concatenate `vedges` and `hedges` in a third dimension, calling the output `feature_map`. `feature_map` should have dimensions (267, 400, 2). Take the following 3x3x2 kernel $B_{ijk}$:
#
# $$
# B_{:,:,1} = B_{:,:,2} = \left( \begin{array}{ccc}
# 0 & 0 & 0 \\
# 0 & 1 & 0 \\
# 0 & 0 & 0 \\
# \end{array} \right),
# $$
# and apply it to `feature_map`. This time, before we pass it through the activation, we will add a bias. For now, start with a bias of `bias = -2`, and pass the the result through a ReLU, saving the output in variable `outmap`. Plot `outmap` in black & white. Depending on the normalization of your vertical/horizontal kernels, you will have to play with the `bias` until most of the image is black, except for some shapes that should pop out (you'll still have dots of white in other places). Now that the image has passed through 2 CNN layers, what feature(s) does this latest layer seem to be picking out? (Open-ended question, but there are wrong answers. Think about what $B_{ijk}$ is doing, in combination with the bias and rectifier)
#
# + deletable=false
# your code here
# + deletable=false
# your code here
# + [markdown] autograde="2.6" deletable=false editable=false
# **2.6** [5pts] Take a moment to think about the results of question 2.5. What seems to be the purpose of adding more CNN layers to increase the depth of a network? Why might it be useful to have multiple kernels/filters in a layer? Answer in 3-4 sentences.
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] colab_type="text" deletable=false editable=false id="hMCaFlMTVIbo"
# <div class='exercise'> <b> Question 3: Building a Basic CNN Model [30pts total] </b></div>
#
# In this question, you will use Keras to create a convolutional neural network for predicting the type of object shown in images from the [CIFAR-10](https://keras.io/datasets/#cifar10-small-image-classification) dataset, which contains 50,000 32x32 training images and 10,000 test images of the same size, with a total of 10 sizes.
#
# + [markdown] deletable=false editable=false
# <span class='sub-q'> Loading CIFAR-10 and Constructing the Model. </span>
#
# Load CIFAR-10 and use a combination of the following layers: Conv2D, MaxPooling2D, Dense, Dropout and Flatten Layers (not necessarily in this order, and you can use as many layers as you'd like) to build your classification model. You may use an existing architecture like AlexNet or VGG16, or create one of your own design. However, you should construct the network yourself and not use a pre-written implementation. At least one of your Conv2D layers should have at least 9 filters to be able to do question 3.3.
#
# Convolutional neural networks are very computationally intensive. We highly recommend that you train your model on a system using GPUs. On CPUs, this training can take over an hour. On GPUs, it can be done within minutes. If you become frustrated having to rerun your model every time you open your notebook, take a look at how to save your model weights as explicitly detailed in **question 4**, where it is required to save your weights.
#
# You can approach the problems in this question by first creating a model assigning 32 filters to each Conv2D layer recreate the model with 64 filters/layer, 128, etc. For each generated model, keep track of the total number of parameters.
#
# **3.1** [6pts] Report the total number of parameters in your model. How does the number of total parameters change (linearly, exponentially) as the number of filters per layer increases (your model should have at least 2 Conv layers)? You can find this empirically by constructing multiple models with the same type of architecture, increasing the number of filters. Generate a plot showing the relationship and explain why it has this relationship.
# + [markdown] colab_type="text" deletable=false editable=false id="SNY2YPTWVIbr"
# **3.2** Choosing a Model, Training and Evaluating It. [7pts total]
# **[5pts]** Take your model from above and train it. You can choose to train your model for as long as you'd like, but you should aim for at least 10 epochs. Your validation accuracy should exceed 70%. Training for 10 epochs on a CPU should take about 30-60 minutes. **[2pts]** Plot the loss and accuracy (both train and test) for your chosen architecture.
# + [markdown] colab_type="text" deletable=false editable=false id="SEgT5bNOVIbu"
# **Techniques to Visualize the Model.**
#
# We will gain an intuition into how our model is processing the inputs in two ways. First we'll ask you to use feature maps to visualize the activations in the intermediate layers of the network. We've provided a helper function `get_feature_maps` to aid in extracting feature maps from layer outputs in your model network. Feel free to take advantage of it if you'd like. We'll also ask you to use [saliency maps](https://arxiv.org/abs/1312.6034) to visualize the pixels that have the largest impact on the classification of an input (image in this case), as well as a more recent development,[Grad-CAM](https://arxiv.org/abs/1610.02391), which has been shown to better indicate the attention of CNNs.
#
# **3.3** [5pts] For a given input image from the test set that is correctly classified, use your model and extract 9 feature maps from an intermediate convolutional layer of your choice and plot the images in a 3x3 grid (use `imshow`'s `cmap='gray'` to show the feature maps in black & white). Make sure to plot (and clearly label) your original input image as well. You may use the provided `get_feature_maps` function and the `cifar10dict` dictionary to convert class index to the correct class name.
#
# **3.4** [5pts] For the same input image generate and plot a (SmoothGrad) saliency map to show the pixels in the image most pertinent to classification, and a Grad-CAM heatmap. This is most easily done with the [tf-keras-vis](https://pypi.org/project/tf-keras-vis/) package. Take a look at the "Usage" examples; it will be straightforward to apply to our model. Feel free to pick your own [colormap](https://matplotlib.org/3.1.0/tutorials/colors/colormaps.html); the `jet` colormap is harder to interpret than sequential ones. Arrange the three plots in a row using subplots: Original Image, Saliency Map, GradCAM. Which visualization is easier to understand in your case, and what does the network seem to be focusing on?
#
# **3.5** [7pts] Repeat `3.4` for an image from the test set that is *incorrectly classified*, indicating both the incorrect label and what the correct label should be, and from the visualizations of network attention, hypothesize why the network arrived at its answer. (Make sure you pass a new loss to the visualizers that uses the *incorrect* class index, because we want to see what caused the network to think the image was in that category!) If you had control over what images go in the training dataset, how could you modify it to avoid this particular network failure?
# + [markdown] colab_type="text" deletable=false editable=false id="N9lpw9ivXUck"
# *Some code that will help you generate feature maps*
# + colab={} colab_type="code" deletable=false editable=false id="nFxRTGLyVIbx"
def get_feature_maps(model, layer_id, input_image):
"""Returns intermediate output (activation map) from passing an image to the model
Parameters:
model (tf.keras.Model): Model to examine
layer_id (int): Which layer's (from zero) output to return
input_image (ndarray): The input image
Returns:
maps (List[ndarray]): Feature map stack output by the specified layer
"""
model_ = Model(inputs=[model.input], outputs=[model.layers[layer_id].output])
return model_.predict(np.expand_dims(input_image, axis=0))[0,:,:,:].transpose((2,0,1))
# + [markdown] colab_type="text" deletable=false editable=false id="7Beiv7ULXUcw"
# *A dictionary to turn class index into class labels for CIFAR-10*
# + colab={} colab_type="code" deletable=false editable=false id="6hvLFUROXUc2"
cifar10dict = {0 : 'airplane', 1 : 'automobile', 2 : 'bird', 3 : 'cat', 4 : 'deer', 5 : 'dog', 6 : 'frog', 7 : 'horse', 8 : 'ship', 9 : 'truck'}
# + [markdown] deletable=false editable=false
# *Some imports for getting the CIFAR-10 dataset and for help with visualization*
# + deletable=false editable=false
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.utils import to_categorical
from tf_keras_vis.saliency import Saliency
from tf_keras_vis.utils import normalize
from matplotlib import cm
from tf_keras_vis.gradcam import Gradcam
# + [markdown] deletable=false editable=false
# ## Answers
# + [markdown] autograde="3.1" deletable=false editable=false
# **3.1** [6pts] Report the total number of parameters in your model. How does the number of total parameters change (linearly, exponentially) as the number of filters per layer increases (your model should have at least 2 Conv layers)? You can find this empirically by constructing multiple models with the same type of architecture, increasing the number of filters. Generate a plot showing the relationship and explain why it has this relationship.
# + deletable=false
# your code here
# + deletable=false
# your code here
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] autograde="3.2" deletable=false editable=false
# **3.2** Choosing a Model, Training and Evaluating It. [7pts total]
# **[5pts]** Take your model from above and train it. You can choose to train your model for as long as you'd like, but you should aim for at least 10 epochs. Your validation accuracy should exceed 70%. Training for 10 epochs on a CPU should take about 30-60 minutes. **[2pts]** Plot the loss and accuracy (both train and test) for your chosen architecture.
# + deletable=false
# your code here
# + deletable=false
# your code here
# + deletable=false
# your code here
# + deletable=false
# plotting
# your code here
# -
# Save weights
# your code here
# + deletable=false
# Load saved weights
# your code here
# + [markdown] autograde="3.3" deletable=false editable=false
# **3.3** [5pts] For a given input image from the test set that is correctly classified, use your model and extract 9 feature maps from an intermediate convolutional layer of your choice and plot the images in a 3x3 grid (use `imshow`'s `cmap='gray'` to show the feature maps in black & white). Make sure to plot (and clearly label) your original input image as well. You may use the provided `get_feature_maps` function and the `cifar10dict` dictionary to convert class index to the correct class name.
#
# + deletable=false
# your code here
# + [markdown] autograde="3.4" deletable=false editable=false
# **3.4** [5pts] For the same input image generate and plot a (SmoothGrad) saliency map to show the pixels in the image most pertinent to classification, and a Grad-CAM heatmap. This is most easily done with the [tf-keras-vis](https://pypi.org/project/tf-keras-vis/) package. Take a look at the "Usage" examples; it will be straightforward to apply to our model. Feel free to pick your own [colormap](https://matplotlib.org/3.1.0/tutorials/colors/colormaps.html); the `jet` colormap is harder to interpret than sequential ones. Arrange the three plots in a row using subplots: Original Image, Saliency Map, GradCAM. Which visualization is easier to understand in your case, and what does the network seem to be focusing on?
#
# + deletable=false
# your code here
# -
# *Your answer here*
#
# + [markdown] autograde="3.5" deletable=false editable=false
# **3.5** [7pts] Repeat `3.4` for an image from the test set that is *incorrectly classified*, indicating both the incorrect label and what the correct label should be, and from the visualizations of network attention, hypothesize why the network arrived at its answer. (Make sure you pass a new loss to the visualizers that uses the *incorrect* class index, because we want to see what caused the network to think the image was in that category!) If you had control over what images go in the training dataset, how could you modify it to avoid this particular network failure?
# + deletable=false
# your code here
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] deletable=false editable=false
# <div class='exercise'> <b> Question 4: Image Orientation Estimation [30pts] </b></div>
# + [markdown] deletable=false editable=false
# In this problem we will construct a neural network to predict how far a face is from being "upright". Image orientation estimation with convolutional networks was first implemented in 2015 by Fischer, Dosovitskiy, and Brox in a paper titled ["Image Orientation Estimation with Convolutional Networks"](https://lmb.informatik.uni-freiburg.de/Publications/2015/FDB15/image_orientation.pdf), where the authors trained a network to straighten a wide variety of images using the Microsoft COCO dataset. In order to have a reasonable training time for a homework, we will be working on a subset of the problem where we just straighten images of faces. To do this, we will be using the [CelebA](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) dataset of celebrity faces, where we assume that professional photographers have taken level pictures. The training will be supervised, with a rotated image (up to $\pm 60^\circ$) as an input, and the amount (in degrees) that the image has been rotated as a target.
#
# The network training for this question can be long (even using a GPU on the JupyterHub, it can take 1-2 hours to reach peak network performance), but deep learning generally requires substantial training times on the order of days or weeks. One aim of this problem is to give you a gentle introduction to some techniques for prototyping such networks before a full training.
# + [markdown] deletable=false editable=false
# <span class='sub-q'> Loading CelebA and Rotating Images. [5 pts] </span>
#
# **4.1** Loading CelebA and Thinking about Datasets. [2pts] Run the cells provided below to automatically download the CelebA dataset. It is about 1.3GB, which can take 10-20 minutes to download. This happens only once; in the future when you rerun the cell, it will use the dataset stored on your machine. The creation of the normalization/rotation/resize pipeline has been done for you, resulting in train dataset `train_rot_ds` and test dataset `test_rot_ds`. [TensorFlow Datasets](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) are objects that behave like Python generators, allowing you to take elements (either input/target tuples or feature dictionaries) until you have gone through the entire dataset. Note how this is different from Question 3 where the entire dataset was loaded in as an array. Datasets also allow you to pipeline transformations to be applied to the elements, resulting in a new transformed Dataset (like `train_rot_ds`). **Question: Aside from pipelining, what is an important practical reason to use Datasets over simply loading all the data in X and Y arrays?**
#
# **4.2** Taking a look. [3pts] In a grid of subplots, plot at least 4 rotated images from `train_rot_ds` with the titles being the amount the images have been rotated. The floating point numbers in the titles should have a reasonable number of digits (read about formatting floats using Python f-strings if you're unfamiliar). Hint: one way to get a few image+label tuples from the Dataset is with `train_rot_ds.take(4)`. Check the [TensorFlow Datasets documentation](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) for more.
# + [markdown] deletable=false editable=false
# <span class='sub-q'> Building a Model. [11pts total]</span>
#
# **4.3** Conceptual Question. [2pts] Dropout layers have been shown to work well for regularizing deep neural networks, and can be used for very little computational cost. For our network, is it a good idea to use dropout layers? Explain, being sure to explicitly discuss how a dropout layer works, and what that would mean for our model.
#
# **4.4** Compile a Model. [4pts] Construct a model with multiple Conv layers and any other layers you think would help. Be sure to output `<yourmodelname>.summary()` as always. Feel free to experiment with architectures and number of parameters if you wish to get better performance or better training speed. You certainly don't need more than a few million parameters; we were able to it with substantially fewer. Any working setup is acceptable though.
#
# **4.5** Training the Model. [5pts] Train your model using `<yourmodelname>.fit()`. The syntax is a little different when working with Datasets instead of numpy arrays; take a look at the [tf.keras.Model](https://www.tensorflow.org/api_docs/python/tf/keras/Model#fit) documentation. Be sure to also pass the test data as validation data. When passing `train_rot_ds` to `fit()`, you will find it useful to use pipelines to [batch](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch) the data. You can also experiment with [prefetching](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch) batches/elements from the dataset, which may allow you to speed up iterations by a few percent. Finally, while dry-running and prototyping your model, you may find it useful to [take](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#take) a subset of the data to speed up experimentation. Your final model should be trained on all the available training data though. You should achieve a validation loss of less than 9, corresponding to $\pm 3^\circ$ accuracy in predicting the rotations on the test set. This can be achieved in just 2-3 epochs, though you are free to train as long as you want.
# + [markdown] deletable=false editable=false
# <span class='sub-q'> Saving a Model. [3pts total]</span>
#
# There are various ways to save a model, to various degrees. You can simply save the weights, you can save just the architecture, or you can save the whole model, including the state of the optimizer. The final way allows you to restart training as if it never stopped. For our purposes, we will only be saving the weights for simplicity. This way tends to be the most robust, and in general you'll usually have fewer problems running your model on other machines, or interfacing with other packages like tf-keras-vis.
#
# **4.6** Conceptual Question.[2pt] Suppose you save just the weights after training for a while. If you were to load the weights again and continue training, would it work? How will it be different than continuing from a full-model save? Answer in a few sentences.
#
# **4.7** Save and load your weights. [1pt] Save your model weights to the path 'model/*somename*' where *somename* is whatever filename prefix you want. Then load weights from the same path.
#
# Note: If you don't intend to use it, you may leave your line of code commented out. Nothing should change if you run it after saving it though, since it will load the same weights and everything else about the model will still be in memory. If you close your notebook or restart your kernel in the future, run all the cells required to compile the model, but skip the cells that performs the fit and the save. After running the load weights cell, your previously trained model will be restored.
# + [markdown] deletable=false editable=false
# <span class='sub-q'> Testing your model. [11pts total]</span>
#
# **4.8** Checking out performance on the Test Set.[5pts] Create a subplots grid with 4 rows and 3 columns. Each row will be a separate image from the test set (of your choice) and each column will consist of: Original Image, Predicted Straightened Image, Target Straightened Image. The title of the latter two should be the predicted rotation and the actual rotation. For example, a row should look something like this:
# 
# This can be achieved using the provided function `rot_resize` to correct for the rotation predicted by your network.
#
# **4.9** Visualizing Attention. [5pts] Like in question 3, we will use the saliency map and GradCAM to see what the network was looking at to determine the orientation of a testset image. The code will be very similar to what you used in question 3, but there are two important modifications. In defining the new `model_modifier(m)` function, simply replace the contents with `pass`. This is because your model does not (should not) have a softmax activation on the last layer, so we don't need this function to do anything. The other modification is to change the loss function (that was defined as a Python lambda function) to an MSE, so it should now be `tf.keras.backend.mean((output - label)**2)` where label is the actual rotation of the image. Pick any image from the test set, and like before, make a row of 3 subplots showing the original image, the saliency map, and the GradCAM output. __Question: What types of features does the network appear to use to determine orientation?__
#
# **4.10** Correct an image of your choosing. [1pt] Find an image or image(s) (not from the provided test/training sets), or make your own. You may rotate it yourself up to $\pm60^\circ$, or the face can already be naturally rotated. Resize and crop the image to 140px by 120px, load it here, and normalize it to [0.,1.] (you may use the provided `normalize_image` function) and use your network to correct it. I found that my network was a very effective "un-confuser":
# 
# + [markdown] deletable=false editable=false
# ## Answers
# + [markdown] autograde="4.1" deletable=false editable=false
# **4.1** Loading CelebA and Thinking about Datasets. [2pts] Run the cells provided below to automatically download the CelebA dataset. It is about 1.3GB, which can take 10-20 minutes to download. This happens only once; in the future when you rerun the cell, it will use the dataset stored on your machine. The creation of the normalization/rotation/resize pipeline has been done for you, resulting in train dataset `train_rot_ds` and test dataset `test_rot_ds`. [TensorFlow Datasets](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) are objects that behave like Python generators, allowing you to take elements (either input/target tuples or feature dictionaries) until you have gone through the entire dataset. Note how this is different from Question 3 where the entire dataset was loaded in as an array. Datasets also allow you to pipeline transformations to be applied to the elements, resulting in a new transformed Dataset (like `train_rot_ds`). **Question: Aside from pipelining, what is an important practical reason to use Datasets over simply loading all the data in X and Y arrays?**
#
# + deletable=false
import certifi
import urllib3 # For handling https certificate verification
import scipy.ndimage as ndimage
import tensorflow_datasets as tfds
import tensorflow_addons as tfa
# This line will download the CelebA dataset. The download will only happen the first time you ever run this cell.
train_celeb, test_celeb = tfds.load('celeb_a', split=['train', 'test'], shuffle_files=False)
# + deletable=false editable=false
# You may use the following two functions
def normalize_image(img):
return tf.cast(img, tf.float32)/255.
def rot_resize(img, deg):
rotimg = ndimage.rotate(img, deg, reshape=False, order=3)
rotimg = np.clip(rotimg, 0., 1.)
rotimg = tf.image.resize_with_crop_or_pad(rotimg,140,120)
return rotimg
################################################################
# Don't manually invoke these functions; they are for Dataset
# pipelining that is already done for you.
################################################################
def tf_rot_resize(img, deg):
"""Dataset pipe that rotates an image and resizes it to 140x120"""
rotimg = tfa.image.rotate(img, deg/180.*np.pi, interpolation="BILINEAR")
rotimg = tf.image.resize_with_crop_or_pad(rotimg,140,120)
return rotimg
def tf_random_rotate_helper(image):
"""Dataset pipe that normalizes image to [0.,1.] and rotates by a random
amount of degrees in [-60.,60.], returning an (input,target) pair consisting
of the rotated and resized image and the degrees it has been rotated by."""
image = normalize_image(image)
deg = tf.random.uniform([],-60.,60.)
return (tf_rot_resize(image,deg), deg) # (data, label)
def tf_random_rotate_image(element):
"""Given an element drawn from the CelebA dataset, this returns a rotated
image and the amount it has been rotated by, in degrees."""
image = element['image']
image, label = tf_random_rotate_helper(image)
image.set_shape((140,120,3))
return image, label
################################################################
# + deletable=false
# Pipeline for creating randomly rotated images with their target labels being
# the amount they were rotated, in degrees.
train_rot_ds = train_celeb.map(tf_random_rotate_image)
test_rot_ds = test_celeb.map(tf_random_rotate_image)
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] autograde="4.2" deletable=false editable=false
# **4.2** Taking a look. [3pts] In a grid of subplots, plot at least 4 rotated images from `train_rot_ds` with the titles being the amount the images have been rotated. The floating point numbers in the titles should have a reasonable number of digits (read about formatting floats using Python f-strings if you're unfamiliar). Hint: one way to get a few image+label tuples from the Dataset is with `train_rot_ds.take(4)`. Check the [TensorFlow Datasets documentation](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) for more.
# + deletable=false
# your code here
# + [markdown] autograde="4.3" deletable=false editable=false
# **4.3** Conceptual Question. [2pts] Dropout layers have been shown to work well for regularizing deep neural networks, and can be used for very little computational cost. For our network, is it a good idea to use dropout layers? Explain, being sure to explicitly discuss how a dropout layer works, and what that would mean for our model.
#
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] autograde="4.4" deletable=false editable=false
# **4.4** Compile a Model. [4pts] Construct a model with multiple Conv layers and any other layers you think would help. Be sure to output `<yourmodelname>.summary()` as always. Feel free to experiment with architectures and number of parameters if you wish to get better performance or better training speed. You certainly don't need more than a few million parameters; we were able to it with substantially fewer. Any working setup is acceptable though.
#
# + deletable=false
# your code here
# + deletable=false
# your code here
# + [markdown] autograde="4.5" deletable=false editable=false
# **4.5** Training the Model. [5pts] Train your model using `<yourmodelname>.fit()`. The syntax is a little different when working with Datasets instead of numpy arrays; take a look at the [tf.keras.Model](https://www.tensorflow.org/api_docs/python/tf/keras/Model#fit) documentation. Be sure to also pass the test data as validation data. When passing `train_rot_ds` to `fit()`, you will find it useful to use pipelines to [batch](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch) the data. You can also experiment with [prefetching](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch) batches/elements from the dataset, which may allow you to speed up iterations by a few percent. Finally, while dry-running and prototyping your model, you may find it useful to [take](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#take) a subset of the data to speed up experimentation. Your final model should be trained on all the available training data though. You should achieve a validation loss of less than 9, corresponding to $\pm 3^\circ$ accuracy in predicting the rotations on the test set. This can be achieved in just 2-3 epochs, though you are free to train as long as you want.
# + deletable=false
# your code here
# + [markdown] autograde="4.6" deletable=false editable=false
# **4.6** Conceptual Question.[2pt] Suppose you save just the weights after training for a while. If you were to load the weights again and continue training, would it work? How will it be different than continuing from a full-model save? Answer in a few sentences.
#
# + [markdown] deletable=false
# *Your answer here*
# + [markdown] autograde="4.7" deletable=false editable=false
# **4.7** Save and load your weights. [1pt] Save your model weights to the path 'model/*somename*' where *somename* is whatever filename prefix you want. Then load weights from the same path.
#
# Note: If you don't intend to use it, you may leave your line of code commented out. Nothing should change if you run it after saving it though, since it will load the same weights and everything else about the model will still be in memory. If you close your notebook or restart your kernel in the future, run all the cells required to compile the model, but skip the cells that performs the fit and the save. After running the load weights cell, your previously trained model will be restored.
# + deletable=false
# your code here
# + [markdown] autograde="4.8" deletable=false editable=false
# **4.8** Checking out performance on the Test Set.[5pts] Create a subplots grid with 4 rows and 3 columns. Each row will be a separate image from the test set (of your choice) and each column will consist of: Original Image, Predicted Straightened Image, Target Straightened Image. The title of the latter two should be the predicted rotation and the actual rotation. For example, a row should look something like this:
# 
# This can be achieved using the provided function `rot_resize` to correct for the rotation predicted by your network.
#
# + deletable=false
# your code here
# + [markdown] autograde="4.9" deletable=false editable=false
# **4.9** Visualizing Attention. [5pts] Like in question 3, we will use the saliency map and GradCAM to see what the network was looking at to determine the orientation of a testset image. The code will be very similar to what you used in question 3, but there are two important modifications. In defining the new `model_modifier(m)` function, simply replace the contents with `pass`. This is because your model does not (should not) have a softmax activation on the last layer, so we don't need this function to do anything. The other modification is to change the loss function (that was defined as a Python lambda function) to an MSE, so it should now be `tf.keras.backend.mean((output - label)**2)` where label is the actual rotation of the image. Pick any image from the test set, and like before, make a row of 3 subplots showing the original image, the saliency map, and the GradCAM output. __Question: What types of features does the network appear to use to determine orientation?__
#
# + deletable=false
# your code here
# -
# *Your answer here*
#
# + [markdown] autograde="4.10" deletable=false editable=false
# **4.10** Correct an image of your choosing. [1pt] Find an image or image(s) (not from the provided test/training sets), or make your own. You may rotate it yourself up to $\pm60^\circ$, or the face can already be naturally rotated. Resize and crop the image to 140px by 120px, load it here, and normalize it to [0.,1.] (you may use the provided `normalize_image` function) and use your network to correct it. I found that my network was a very effective "un-confuser":
# 
# + deletable=false
# your code here
# + deletable=false editable=false
| content/HW/hw4/109/cs109b_hw4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="_-GR0EDHM1SO"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="R3yYtBPkM2qZ"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="6Y8E0lw5eYWm"
# # Post-training dynamic range quantization
# + [markdown] colab_type="text" id="CIGrZZPTZVeO"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/lite/performance/post_training_quant"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/performance/post_training_quant.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/performance/post_training_quant.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/tensorflow/lite/g3doc/performance/post_training_quant.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="BTC1rDAuei_1"
# ## Overview
#
# [TensorFlow Lite](https://www.tensorflow.org/lite/) now supports
# converting weights to 8 bit precision as part of model conversion from
# tensorflow graphdefs to TensorFlow Lite's flat buffer format. Dynamic range quantization achieves a 4x reduction in the model size. In addition, TFLite supports on the fly quantization and dequantization of activations to allow for:
#
# 1. Using quantized kernels for faster implementation when available.
# 2. Mixing of floating-point kernels with quantized kernels for different parts
# of the graph.
#
# The activations are always stored in floating point. For ops that
# support quantized kernels, the activations are quantized to 8 bits of precision
# dynamically prior to processing and are de-quantized to float precision after
# processing. Depending on the model being converted, this can give a speedup over
# pure floating point computation.
#
# In contrast to
# [quantization aware training](https://github.com/tensorflow/tensorflow/tree/r1.14/tensorflow/contrib/quantize)
# , the weights are quantized post training and the activations are quantized dynamically
# at inference in this method.
# Therefore, the model weights are not retrained to compensate for quantization
# induced errors. It is important to check the accuracy of the quantized model to
# ensure that the degradation is acceptable.
#
# This tutorial trains an MNIST model from scratch, checks its accuracy in
# TensorFlow, and then converts the model into a Tensorflow Lite flatbuffer
# with dynamic range quantization. Finally, it checks the
# accuracy of the converted model and compare it to the original float model.
# + [markdown] colab_type="text" id="2XsEP17Zelz9"
# ## Build an MNIST model
# + [markdown] colab_type="text" id="dDqqUIZjZjac"
# ### Setup
# + colab={} colab_type="code" id="gyqAw1M9lyab"
import logging
logging.getLogger("tensorflow").setLevel(logging.DEBUG)
import tensorflow as tf
from tensorflow import keras
import numpy as np
import pathlib
# + [markdown] colab_type="text" id="eQ6Q0qqKZogR"
# ### Train a TensorFlow model
# + colab={"height": 51} colab_type="code" id="hWSAjQWagIHl" outputId="961899f8-1597-4417-b21d-cae94a330ecc"
# Load MNIST dataset
mnist = keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# Normalize the input image so that each pixel value is between 0 to 1.
train_images = train_images / 255.0
test_images = test_images / 255.0
# Define the model architecture
model = keras.Sequential([
keras.layers.InputLayer(input_shape=(28, 28)),
keras.layers.Reshape(target_shape=(28, 28, 1)),
keras.layers.Conv2D(filters=12, kernel_size=(3, 3), activation=tf.nn.relu),
keras.layers.MaxPooling2D(pool_size=(2, 2)),
keras.layers.Flatten(),
keras.layers.Dense(10)
])
# Train the digit classification model
model.compile(optimizer='adam',
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model.fit(
train_images,
train_labels,
epochs=1,
validation_data=(test_images, test_labels)
)
# + [markdown] colab_type="text" id="5NMaNZQCkW9X"
# For the example, since you trained the model for just a single epoch, so it only trains to ~96% accuracy.
#
# + [markdown] colab_type="text" id="xl8_fzVAZwOh"
# ### Convert to a TensorFlow Lite model
#
# Using the Python [TFLiteConverter](https://www.tensorflow.org/lite/convert/python_api), you can now convert the trained model into a TensorFlow Lite model.
#
# Now load the model using the `TFLiteConverter`:
# + colab={} colab_type="code" id="_i8B2nDZmAgQ"
converter = tf.lite.TFLiteConverter.from_keras_model(model)
tflite_model = converter.convert()
# + [markdown] colab_type="text" id="F2o2ZfF0aiCx"
# Write it out to a tflite file:
# + colab={} colab_type="code" id="vptWZq2xnclo"
tflite_models_dir = pathlib.Path("/tmp/mnist_tflite_models/")
tflite_models_dir.mkdir(exist_ok=True, parents=True)
# + colab={"height": 34} colab_type="code" id="Ie9pQaQrn5ue" outputId="046db0bc-1745-4e94-9f21-f7e91bdaebda"
tflite_model_file = tflite_models_dir/"mnist_model.tflite"
tflite_model_file.write_bytes(tflite_model)
# + [markdown] colab_type="text" id="7BONhYtYocQY"
# To quantize the model on export, set the `optimizations` flag to optimize for size:
# + colab={"height": 34} colab_type="code" id="g8PUvLWDlmmz" outputId="d79b45d3-babf-4890-8036-de2f497da88a"
converter.optimizations = [tf.lite.Optimize.DEFAULT]
tflite_quant_model = converter.convert()
tflite_model_quant_file = tflite_models_dir/"mnist_model_quant.tflite"
tflite_model_quant_file.write_bytes(tflite_quant_model)
# + [markdown] colab_type="text" id="PhMmUTl4sbkz"
# Note how the resulting file, is approximately `1/4` the size.
# + colab={"height": 119} colab_type="code" id="JExfcfLDscu4" outputId="d1fda4c2-343e-40fb-f90f-b6bde00c523e"
# !ls -lh {tflite_models_dir}
# + [markdown] colab_type="text" id="L8lQHMp_asCq"
# ## Run the TFLite models
#
# Run the TensorFlow Lite model using the Python TensorFlow Lite
# Interpreter.
#
# + [markdown] colab_type="text" id="Ap_jE7QRvhPf"
# ### Load the model into an interpreter
# + colab={} colab_type="code" id="Jn16Rc23zTss"
interpreter = tf.lite.Interpreter(model_path=str(tflite_model_file))
interpreter.allocate_tensors()
# + colab={} colab_type="code" id="J8Pztk1mvNVL"
interpreter_quant = tf.lite.Interpreter(model_path=str(tflite_model_quant_file))
interpreter_quant.allocate_tensors()
# + [markdown] colab_type="text" id="2opUt_JTdyEu"
# ### Test the model on one image
# + colab={} colab_type="code" id="AKslvo2kwWac"
test_image = np.expand_dims(test_images[0], axis=0).astype(np.float32)
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
interpreter.set_tensor(input_index, test_image)
interpreter.invoke()
predictions = interpreter.get_tensor(output_index)
# + colab={"height": 281} colab_type="code" id="XZClM2vo3_bm" outputId="0fa4155b-01f8-4fea-f586-d9044d73572e"
import matplotlib.pylab as plt
plt.imshow(test_images[0])
template = "True:{true}, predicted:{predict}"
_ = plt.title(template.format(true= str(test_labels[0]),
predict=str(np.argmax(predictions[0]))))
plt.grid(False)
# + [markdown] colab_type="text" id="LwN7uIdCd8Gw"
# ### Evaluate the models
# + colab={} colab_type="code" id="05aeAuWjvjPx"
# A helper function to evaluate the TF Lite model using "test" dataset.
def evaluate_model(interpreter):
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
# Run predictions on every image in the "test" dataset.
prediction_digits = []
for test_image in test_images:
# Pre-processing: add batch dimension and convert to float32 to match with
# the model's input data format.
test_image = np.expand_dims(test_image, axis=0).astype(np.float32)
interpreter.set_tensor(input_index, test_image)
# Run inference.
interpreter.invoke()
# Post-processing: remove batch dimension and find the digit with highest
# probability.
output = interpreter.tensor(output_index)
digit = np.argmax(output()[0])
prediction_digits.append(digit)
# Compare prediction results with ground truth labels to calculate accuracy.
accurate_count = 0
for index in range(len(prediction_digits)):
if prediction_digits[index] == test_labels[index]:
accurate_count += 1
accuracy = accurate_count * 1.0 / len(prediction_digits)
return accuracy
# + colab={"height": 34} colab_type="code" id="DqXBnDfJ7qxL" outputId="78f393f8-c4a5-41e0-abe4-ab6a5c394e51"
print(evaluate_model(interpreter))
# + [markdown] colab_type="text" id="Km3cY9ry8ZlG"
# Repeat the evaluation on the dynamic range quantized model to obtain:
#
# + colab={"height": 34} colab_type="code" id="-9cnwiPp6EGm" outputId="d82552d7-8a2c-49dc-a19a-56010a013102"
print(evaluate_model(interpreter_quant))
# + [markdown] colab_type="text" id="L7lfxkor8pgv"
# In this example, the compressed model has no difference in the accuracy.
# + [markdown] colab_type="text" id="M0o1FtmWeKZm"
# ## Optimizing an existing model
#
# Resnets with pre-activation layers (Resnet-v2) are widely used for vision applications.
# Pre-trained frozen graph for resnet-v2-101 is available on
# [Tensorflow Hub](https://tfhub.dev/google/imagenet/resnet_v2_101/classification/4).
#
# You can convert the frozen graph to a TensorFLow Lite flatbuffer with quantization by:
#
# + colab={} colab_type="code" id="jrXZxSJiJfYN"
import tensorflow_hub as hub
resnet_v2_101 = tf.keras.Sequential([
keras.layers.InputLayer(input_shape=(224, 224, 3)),
hub.KerasLayer("https://tfhub.dev/google/imagenet/resnet_v2_101/classification/4")
])
converter = tf.lite.TFLiteConverter.from_keras_model(resnet_v2_101)
# + colab={"height": 34} colab_type="code" id="LwnV4KxwVEoG" outputId="7d50f90d-6104-43a3-863c-28db9465d483"
# Convert to TF Lite without quantization
resnet_tflite_file = tflite_models_dir/"resnet_v2_101.tflite"
resnet_tflite_file.write_bytes(converter.convert())
# + colab={"height": 34} colab_type="code" id="2qkZD0VoVExe" outputId="76a47590-fa91-49b9-f568-4e00b46c9537"
# Convert to TF Lite with quantization
converter.optimizations = [tf.lite.Optimize.DEFAULT]
resnet_quantized_tflite_file = tflite_models_dir/"resnet_v2_101_quantized.tflite"
resnet_quantized_tflite_file.write_bytes(converter.convert())
# + colab={"height": 102} colab_type="code" id="vhOjeg1x9Knp" outputId="c643a660-f815-49f0-ac4b-ac48af3c1203"
# !ls -lh {tflite_models_dir}/*.tflite
# + [markdown] colab_type="text" id="qqHLaqFMCjRZ"
# The model size reduces from 171 MB to 43 MB.
# The accuracy of this model on imagenet can be evaluated using the scripts provided for [TFLite accuracy measurement](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/lite/tools/accuracy/ilsvrc).
#
# The optimized model top-1 accuracy is 76.8, the same as the floating point model.
| tensorflow/lite/g3doc/performance/post_training_quant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
# http://media.daum.net/economic/
# path = 'https://www.genie.co.kr/chart/top200/'
path = 'http://media.daum.net/economic/'
req = requests.get(path)
# import time
# time.sleep(5)
req.status_code
from bs4 import BeautifulSoup
soup = BeautifulSoup(req.content, 'html.parser')
type(soup)
result = soup.select('div > strong.tit_thumb > a[href].link_txt')
result[0]
tag = result[0]
result[0].text
tag['href']
# [
# [title. link],
# [title01, link01],
# [title02, link02],
# ...
# ]
contents = []
for tag in result:
# print(tag.text.strip(),tag['href'])
# strip()으로 공백 제거
title = tag.text.strip()
link = tag['href'].strip()
content = [title, link]
contents.append(content)
len(contents)
import pandas
pd = pandas.DataFrame(contents, columns=['Title', 'Link'])
pd
pd.to_excel('./saves/economic01.xls', index=False)
| scraping_news.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp config
# -
# # config
#
# > API details.
#hide
from nbdev.showdoc import *
# +
#export
import urllib
PATH_CHROME_DRIVER = '/home/danph/Repos/run/drivers/chromedriver'
# PATH_PROJECT = '/home/danph/Repos/myprojects/AussieDataScience'
PATH_PROJECT = 'gs://australia-southeast1-dsjobs-72227026-bucket'
LINKEDIN_JOBLIST = PATH_PROJECT + '/data/linkedin_jobs_list_full.csv'
LINKEDIN_JOBLIST_TEMP = PATH_PROJECT + '/data/linkedin_jobs_list_temp'
LINKEDIN_JOBLIST_LATEST_DATE = PATH_PROJECT + "/data/linkedin_jobs_list_latest_date.txt"
LINKEDIN_JOBS = PATH_PROJECT + '/data/linkedin_jobs.csv'
LINKEDIN_JOBS_CLEAN = PATH_PROJECT + '/data/linkedin_jobs_clean.csv'
LINKEDIN_JOBS_DTM = PATH_PROJECT + '/data/linkedin_jobs_dtm.csv'
SEEK_JOBLIST = PATH_PROJECT + '/data/seek_jobs_list.csv'
SEEK_JOBS = PATH_PROJECT + '/data/seek_jobs.csv'
SEEK_JOBS_CLEAN = PATH_PROJECT + '/data/seek_jobs_clean.csv'
SEEK_JOBS_DTM = PATH_PROJECT + '/data/seek_jobs_dtm.csv'
KEYWORDS = ["data scientist", "data analyst", "data engineer",
"machine learning", "machine learning engineer"]
LINKEDIN_BASE = 'https://www.linkedin.com/jobs/search/?f_TPR=r604800&'
LINKEDIN_URLS = [LINKEDIN_BASE+f'keywords={urllib.parse.quote(keyword)}&location=Australia&sortBy=R' for keyword in KEYWORDS]
# -
| 00_config.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# +
# This part is from https://mne.tools/stable/auto_tutorials/machine-learning/plot_sensors_decoding.html
import numpy as np
import matplotlib.pyplot as plt
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
import mne
from mne.datasets import sample
from mne.decoding import (SlidingEstimator, GeneralizingEstimator, Scaler,
cross_val_multiscore, LinearModel, get_coef,
Vectorizer, CSP)
data_path = sample.data_path()
subjects_dir = data_path + '/subjects'
raw_fname = data_path + '/MEG/sample/sample_audvis_raw.fif'
tmin, tmax = -0.200, 0.500
event_id = {'Auditory/Left': 1, 'Visual/Left': 3} # just use two
raw = mne.io.read_raw_fif(raw_fname, preload=True)
# The subsequent decoding analyses only capture evoked responses, so we can
# low-pass the MEG data. Usually a value more like 40 Hz would be used,
# but here low-pass at 20 so we can more heavily decimate, and allow
# the examlpe to run faster. The 2 Hz high-pass helps improve CSP.
raw.filter(2, 20)
events = mne.find_events(raw, 'STI 014')
# Set up pick list: EEG + MEG - bad channels (modify to your needs)
raw.info['bads'] += ['MEG 2443', 'EEG 053'] # bads + 2 more
# Read epochs
epochs = mne.Epochs(raw, events, event_id, tmin, tmax, proj=True,
picks=('mag', 'grad', 'eog'), baseline=(None, 0.), preload=True,
reject=dict(grad=4000e-13, eog=150e-6), decim=10)
epochs.pick_types(meg=True, exclude='bads') # remove stim and EOG
del raw
X = epochs.get_data() # MEG signals: n_epochs, n_meg_channels, n_times
y = epochs.events[:, 2] # target: Audio left or right
# -
epochs.info
y
y[y==3] = 0
y
from sklearn.model_selection import StratifiedShuffleSplit
cv = StratifiedShuffleSplit(n_splits=5, test_size=0.2, random_state=124)
clf = make_pipeline(StandardScaler(), LogisticRegression())
time_decod = SlidingEstimator(clf, scoring='roc_auc')
X.shape
scores = cross_val_multiscore(time_decod, X, y, cv=cv)
scores.shape
plt.plot(epochs.times, scores.mean(axis=0));
clf = make_pipeline(StandardScaler(),
LinearModel(LogisticRegression()))
time_decod = SlidingEstimator(clf, scoring='roc_auc')
time_decod.fit(X, y)
coef = get_coef(time_decod, 'filters_', inverse_transform=True)
coef.shape
evoked_coef = mne.EvokedArray(coef,
epochs.info,
tmin=epochs.times[0])
evoked_coef.plot_joint();
from sklearn.naive_bayes import GaussianNB
clf_nb = make_pipeline(StandardScaler(), GaussianNB())
time_decod_nb = SlidingEstimator(clf_nb, scoring='roc_auc')
scores_nb = cross_val_multiscore(time_decod_nb, X, y, cv=cv)
plt.plot(epochs.times, scores.mean(axis=0), 'b', label='LR')
plt.plot(epochs.times, scores_nb.mean(axis=0), 'r', label='GNB');
plt.legend();
plt.grid();
scores.mean()
scores_nb.mean()
plt.plot(epochs.times ,scores.std(axis=0), 'b', label='LR')
plt.plot(epochs.times ,scores_nb.std(axis=0), 'r', label='GNB')
plt.legend();
plt.grid();
| mne_decoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 需要先安裝 gym[atari]
# headless 執行: xvfb-run -a jupyter notebook
import gym
env = gym.make('Pong-ram-v0')
import numpy as np
import ipywidgets as W
from PIL import Image
from io import BytesIO
def to_png(a):
with BytesIO() as bio:
Image.fromarray(a).save(bio, 'png')
return bio.getvalue()
# Q learning 的骨架
#
# * 用 `compute_s(observation)` 來計算 state
# * 用 `Qupdate(s, a, v)` 來 update `Q(s,a)`
# * 用 `Qfunc(s)` 來算 `Q(s, ...)`
#
# +
from time import sleep
from random import randint, random, shuffle, choice
actions = [0,2,3]
def Qlearn(test=False, screen=None, T=40):
observation = env.reset()
for i in range(50):
observation, reward, done, info =env.step(choice(actions))
s2 = compute_s(observation)
total_r = 0
for i in range(T):
s = s2
if not test and random()< ϵ:
a = choice(actions)
elif s is None:
a = choice(actions)
else:
a = actions[np.argmax(Qfunc(s))]
observation, reward, done, info = env.step(a)
s2 = compute_s(observation)
r = reward
total_r+=r
if not test and s is not None:
if s2 is None:
r=1.
if r:
v = r
else:
v = γ*Qfunc(s2).max()
a = max(0, a-1)
Qupdate(s, a, v)
if screen is not None:
img = env.render(mode='rgb_array')
screen.value = to_png(img)
sleep(1/60)
return total_r
# -
# 最簡單的是把整個 observation 當成 state,但是很慢
# +
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, LeakyReLU
from keras.optimizers import Adam, SGD
Q = Sequential()
Q.add(Dense(2048, input_shape=(256,))) # 輸入是 i, j
Q.add(LeakyReLU(0.2))
Q.add(Dense(3)) # 因為輸出是 +-1
Q.compile(loss='mse',optimizer=Adam(1e-4), metrics=['accuracy']) # 輸出 a
def Qfunc(s):
return Q.predict(s)[0]
def Qupdate(s, a, v):
Y = Q.predict(s)
Y[0][a] = v
return Q.train_on_batch(s, Y)
def compute_s(observation):
dx = (observation[58]+128)%256-128
if dx>=0:
return None
ob = list(observation)
return np.array([ob+[(v+128)%256-128 for v in ob]], dtype='float32')/255
# -
# %matplotlib inline
from matplotlib import pyplot as plt
# 很久,可以小跑一下看看
screen = W.Image()
display(screen)
txt =W.Text()
display(txt)
r = 0
γ = 1
ϵ = 0.1
rr= -1
rate = []
for j in range(101):
if j%100==99:
r=sum(Qlearn(test=True, T=40) for i in range(20))
print(j, r/20)
if r>=-2:
break
rr = rr*0.95 + 0.05*Qlearn(T=40)
rate.append(rr)
txt.value="j={} r={}".format(j,rr)
plt.clf()
plt.plot(rate)
with BytesIO() as bio:
plt.savefig(bio)
screen.value = bio.getvalue()
# 之前 tabular 的 state, 用二次函數近似
# +
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, LeakyReLU, LocallyConnected1D, Lambda
from keras.initializers import Constant, RandomNormal
from keras.activations import selu
from keras.optimizers import Adam, SGD, RMSprop
Q = Sequential()
Q.add(Dense(3, input_shape=(2,),
kernel_initializer=RandomNormal(stddev=0.001) )) # 輸入是 i, j
Q.compile(loss='mse',optimizer=SGD(1e-3)) # 輸出 a
def Qfunc(s):
X = np.array([[s[0][0], s[0][0]**2]])
return Q.predict(X)[0]
def Qupdate(s, a, v):
X = np.array([[s[0][0], s[0][0]**2]])
Y = Q.predict(X)
Y[0][a] = v
return Q.train_on_batch(X, Y)
def compute_s(observation):
dx = (observation[58]+127)%256-127
if dx>=0:
return None
dy = (observation[56]+127)%256-127
x,y0 = observation[[49,54]]
y2 = observation[60]
y = (int(y0 - (186-x)*dy/dx)-44)%(326)
if y>163:
y=326-y
y+=38
s = (y-y2)/2
return np.float32([[s/50]])
# -
screen = W.Image()
display(screen)
txt =W.Text()
display(txt)
γ, ϵ, rr = 1, 3., -8
for j in range(1001):
if j%100==99:
r=sum(Qlearn(test=True, T=400) for i in range(20))
print(j, r/20)
rr = rr*0.95 + 0.05*Qlearn(T=400)
txt.value="j={} r={} ϵ={}".format(j,rr, ϵ)
plt.clf()
Qvalue = np.array([Qfunc(np.float32([[i/50]])) for i in range(-50,50)])
plt.plot(Qvalue[:,0], 'r')
plt.plot(Qvalue[:,1], 'g')
plt.plot(Qvalue[:,2], 'b')
with BytesIO() as bio:
plt.savefig(bio)
screen.value = bio.getvalue()
ϵ = max(0.1, ϵ*0.99)
# 測試
screen = W.Image()
display(screen)
Qlearn(test=True, screen=screen, T=400)
# 用常態分佈曲線來逼近
# +
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, LeakyReLU, LocallyConnected1D, Lambda,Reshape
from keras.initializers import Constant, RandomNormal
from keras.activations import selu
from keras.optimizers import Adam, SGD, RMSprop
import keras.backend as K
Q = Sequential()
Q.add(Dense(3, input_shape=(2,) , use_bias=False)) # 輸入是 i, j
Q.add(Lambda(lambda x: K.exp(x)))
Q.add(Reshape((3,1)))
Q.add(LocallyConnected1D(filters=1,kernel_size=1,
kernel_initializer="zeros"))
Q.add(Reshape( (3,) ))
Q.compile(loss='mse',optimizer=SGD(1e-3)) # 輸出 a
Q.layers[0].set_weights([np.array([[0,-1., 1.],[-3., -3., -3.]])])
Q.layers[3].set_weights([np.array([[[.11]],[[.1]],[[.1]]]), np.array([[-0.]]*3) ])
def Qfunc(s):
X = np.array([[s[0][0], s[0][0]**2]])#, s[0][0]**3, s[0][0]**4]])
return Q.predict(X)[0]
def Qupdate(s, a, v):
X = np.array([[s[0][0], s[0][0]**2]])#, s[0][0]**3, s[0][0]**4]])
Y = Q.predict(X)
Y[0][a] = v
return Q.train_on_batch(X, Y)
def compute_s(observation):
dx = (observation[58]+127)%256-127
if dx>=0:
return None
dy = (observation[56]+127)%256-127
x,y0 = observation[[49,54]]
y2 = observation[60]
y = (int(y0 - (186-x)*dy/dx)-44)%(326)
if y>163:
y=326-y
y+=38
s = (y-y2)/2
return np.float32([[s/50]])
# -
# %matplotlib notebook
# %matplotlib notebook
import matplotlib.pyplot as plt
plt.ion()
fig = plt.figure()
ax = fig.gca()
txt =W.Text()
display(txt)
γ, ϵ, rr = 1, .1, -8
for j in range(0, 1001):
if j%100==99:
r=sum(Qlearn(test=True, T=1400) for i in range(20))
print("{} {}\n".format(j, r/20))
if r>0:
break
rr = rr*0.95 + 0.05*Qlearn(T=1400)
txt.value="j={} r={} ϵ={}".format(j,rr, ϵ)
ax.clear()
Qvalue = np.array([Qfunc(np.float32([[i/50]])) for i in range(-50,50)])
for i, c in enumerate("rgb"):
ax.plot(Qvalue[:,i], c)
fig.canvas.draw()
# ## 增加兩個新的 action
# 如果之前移動了,則這次不移動。
#
# +
from time import sleep
from random import randint, random, shuffle, choice
actions2 = [0,2,3,4,5]
def Qlearn2(test=False, screen=None, T=40):
observation = env.reset()
for i in range(50):
a = choice(actions2)
observation, reward, done, info =env.step(a)
s2 = compute_s(observation)
last_a = a
total_r = 0
for i in range(T):
s = s2
if not test and random()< ϵ:
a = choice(actions2)
elif s is None:
a = choice(actions2)
else:
a = actions2[np.argmax(Qfunc(s))]
if a>=4:
if last_a==a-2:
observation, reward, done, info = env.step(0)
last_a = 0
else:
observation, reward, done, info = env.step(a-2)
last_a = a-2
else:
observation, reward, done, info = env.step(a)
last_a = a
s2 = compute_s(observation)
r = reward
total_r+=r
if not test and s is not None:
if s2 is None:
r=1.
if r:
v = r
else:
v = γ*Qfunc(s2).max()
a = max(0, a-1)
Qupdate(s, a, v)
if screen is not None:
img = env.render(mode='rgb_array')
screen.value = to_png(img)
sleep(1/60)
return total_r
# -
# 模型也一樣,只是多了兩個輸出。
# +
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, LeakyReLU, LocallyConnected1D, Lambda,Reshape
from keras.initializers import Constant, RandomNormal
from keras.activations import selu
from keras.optimizers import Adam, SGD, RMSprop
import keras.backend as K
Q = Sequential()
Q.add(Dense(5, input_shape=(2,) , use_bias=False)) # 輸入是 i, j
Q.add(Lambda(lambda x: K.exp(x)))
Q.add(Reshape((5,1)))
Q.add(LocallyConnected1D(filters=1,kernel_size=1,
kernel_initializer="zeros"))
Q.add(Reshape( (5,) ))
Q.compile(loss='mse',optimizer=SGD(1e-3)) # 輸出 a
Q.layers[0].set_weights([np.array([[0,-1., 1., -1, 1],
[-12.,-3.,-3.,-6.,-6.]])])
Q.layers[3].set_weights([np.array([[[.16]],[[.1]],[[.1]],[[.15]],[[.15]]]), np.array([[-0.]]*5) ])
def Qfunc(s):
X = np.array([[s[0][0], s[0][0]**2]])
return Q.predict(X)[0]
def Qupdate(s, a, v):
X = np.array([[s[0][0], s[0][0]**2]])
Y = Q.predict(X)
Y[0][a] = v
return Q.train_on_batch(X, Y)
def compute_s(observation):
dx = (observation[58]+127)%256-127
if dx>=0:
return None
dy = (observation[56]+127)%256-127
x,y0 = observation[[49,54]]
y2 = observation[60]
y = (int(y0 - (186-x)*dy/dx)-44)%(326)
if y>163:
y=326-y
y+=38
s = (y-y2)/2
return np.float32([[s/50]])
# -
# %matplotlib notebook
# %matplotlib notebook
import matplotlib.pyplot as plt
plt.ion()
fig = plt.figure()
ax = plt.gca()
txt =W.Text()
display(txt)
γ, ϵ, rr = 1, .1, -8
for j in range(0, 1001):
if j%100==99:
r=sum(Qlearn2(test=True, T=1400) for i in range(20))
print("{} {}\n".format(j, r/20))
if r>0:
break
rr = rr*0.95 + 0.05*Qlearn2(T=1400)
txt.value="j={} r={} ϵ={}".format(j,rr, ϵ)
ax.clear()
Qvalue = np.array([Qfunc(np.float32([[i/50]])) for i in range(-50,50)])
for i, c in enumerate("rgbyk"):
ax.plot(Qvalue[:,i], c)
fig.canvas.draw()
from keras.models import load_model
Q = load_model('Qlearn2_function.h5')
# 測試
screen = W.Image()
display(screen)
Qlearn2(test=True, screen=screen, T=1400)
# +
#Q.save('Qlearn2_function.h5')
| RL/Pong-Function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Table of Contents
# - **What is Python**
# - **Python environments**
# - **Setup Guide**
# # What is Python?
#
# Python is a modern, general-purpose, object-oriented, high-level programming language.
#
# General characteristics of Python:
#
# * *clean and simple language*:
# * easy to read code,
# * easy to learn, natural, syntax,
# * maintainability scales well with size of projects.
# * *expressive language*: fewer lines of code, fewer bugs, easier to maintain.
#
# Technical details:
#
# * *dynamically typed*: no need to define the type of variables, function arguments or return types,
# * *automatic memory management*: no need to explicitly allocate and deallocate memory for variables and data arrays,
# * *interpreted**: source code is not compiled to binary machine code but pre-compiled in an intermediate representation (named bytecode); the Python interpreter reads and executes the bytecode,
# * *object-oriented*: effective in tackling complexity for large programs.
#
# Advantages:
#
# * Ease of programming, minimizing the time required to develop, debug and maintain the code,
# * Well designed language that encourage many good programming practices:
# * “Blocks by Indentation” forces proper code structuring & readability,
# * Modular and object-oriented programming, good system for packaging and re-use of code,
# * Documentation tightly integrated with the code,
# * A large standard library, and a large collection of add-on packages.
#
# Disadvantages:
#
# * Since Python is an interpreted and dynamically typed programming language, the execution of python code can be slow compared to compiled statically typed programming languages, such as C/C++,
# * Somewhat decentralized, with different environment, packages and documentation spread out at different places.
# # Python environments
#
# There are many different *environments* through which the Python *interpreter* can be used. Each environment has different advantages and is suitable for different workflows. One strength of Python is that it is versatile and can be used in complementary ways, but it can be confusing for beginners so we will start with a brief survey of python environments.
#
#
# ## Python interpreter
#
# The standard way to use the Python programming language is to use the Python interpreter to run Python code. The **Python interpreter is a program that reads and execute the python code in files passed to it as arguments**. At the command prompt, the command `python` is used to invoke the Python interpreter.
# For example, to run a file `my-program.py` that contains Python code from the command prompt, use:
#
# ```shell
# $ python my-program.py
# ```
#
# We can also start the interpreter by simply typing `python` at the command line, and interactively type Python code into the interpreter (press `Ctrl+D` or type `exit()` to exit):
#
# ```python
# while True:
# code = input(">>> ") # prompt the user for some code
# exec(code) # execute it
# ```
#
# This model is often called a REPL, or Read-Eval-Print-Loop:
#
# The `>>>` symbols represent the *prompt* where you will type code expressions. The interpreter awaits our instructions step by step. This is often how we want to work when quick testing **small portions of code**, or when doing small calculations.
#
#
# While some Python programmers execute all of their Python code in this way, those doing data analysis or scientific computing make use of IPython (an enhanced Python interpreter) or Jupyter notebooks (web-based code notebooks originally created within the IPython project).
#
#
# ## IPython
# IPython (*Interactive Python*) is a **command shell for interactive computing in Python**.
#
# IPython offers a fully compatible replacement for the standard Python interpreter, with convenient shell features, special commands, command history mechanism and output results caching.
#
# We can start IPython by simply typing `ipython` at the command (press `Ctrl+D` or type `exit()` to exit).
#
# The original IPython interface runs an interactive shell built with Python. The name shell indicates that it is the outermost layer around the kernel (i.e. the engine capable of executing code) and allows user to access it through a user interface.
#
# The default IPython prompt adopts the numbered `In [1]:` style compared with the standard `>>>` prompt. Just as with the standard interpreter, you can execute arbitrary Python statements by typing them in and pressing Return (or Enter).
# ## Jupyter Notebook
#
# The "Project **Jupyter**" denotes an organization created with the aim of **supporting interactive data science and scientific computing** via the development of open-source software.
#
# The **Notebook** term may refer both to the notebook document and the web-based environment used to create it. Jupyter Notebook represents another interface to the IPython kernel and can connect to it to allow interactive programming in Python. It uses an internal library for converting the document in HTML and allows visualization and editing in the browser. Although using a web browser as graphical interface, Jupyter notebooks are usually run locally, from the same computer that runs the browser.
#
# 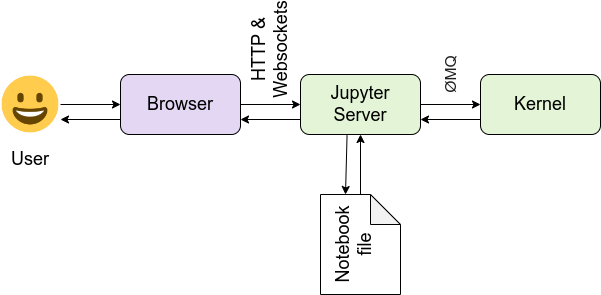
#
# The document you are reading, with extension `.ipynb`, is a notebook and consists in an ordered list of cells which can contatin code, text, mathematics, visualization of output and plots.
#
# Jupyter notebooks are particularly useful as scientific lab books when you are doing lots of data analysis using computational tools. This is because, with Jupyter notebooks, you can:
# * **Record the code you write in a notebook as you manipulate your data**. This is useful to remember what you've done, repeat it if necessary, etc.
# * **Graphs and other figures are rendered directly in the notebook**.
# * You can **update the notebook (or parts thereof) with new data by re-running cells**. You could also copy the cell and re-run the copy only if you want to retain a record of the previous attempt.
# There are two types of cell:
# - **Markdown cells**: contain explanatory text.
# - **Code cells**: contain executable code.
#
# #### Text cells
#
# Text cells (like this) use *markdown syntax*: it consists in plain text formatting syntax that enables the creation of rich text that can be converted to HTML!
#
# You can include well-formatted text, formulas, and images too.
# #### Code cells
# A code cell contains executable code and displays its output just below.
# The subsequent cells are examples of code cell: execute them clicking the play button or using Ctrl+Enter.
# this is an executable code cell.
a = 2
b = 4
a*b
# In *code cells* you can use question mark for accessing the documentation:
# +
# a?
# -
# In *code cells* you can also use system aliases. Use exclamation mark for terminal operation:
# !python --version
# !ls
# # Setup Guide: Installation of Python
#
# ## Versions of Python
#
# from [python wiki](https://wiki.python.org/moin/Python2orPython3):
#
# *Python 2.x is legacy, Python 3.x is the present and future of the language*
#
# >*Python 3.0 was released in 2008. The final 2.x version 2.7 release came out in mid-2010, with a statement of extended support for this end-of-life release. The 2.x branch will see no new major releases after that.*
#
# >*As of January 2020, Python 2 has reached End Of Life (EOL) status, meaning it will receive no further updates or bugfixes, including for security issues. Many frameworks and other add on projects are following a similar policy.*
#
#
# >*As such, we can only recommend learning and teaching Python 3.*
#
# To see which version of Python you have, run:
# ```shell
# $ python --version
# Python 3.9.7
# ```
#
# Several versions of Python can be installed in parallel.
#
# ### IMPORTANT: In this course we will use Python 3
#
# There are several differences between Python 2.7.x and Python 3.x; the most relevant ones are reported [here](https://sebastianraschka.com/Articles/2014_python_2_3_key_diff.html).
# New users are recommended to download and install **Anaconda**.
# It is a package manager, an environment manager, a Python distribution, and a collection of 1,000+ open source packages. Among the available packages:
# - **jupyter**;
# - **numpy**: fundamental package for scientific computing with Python;
# - **matplotlib**: a Python plotting library;
# - **pandas**: a Python library for data pre-processing and data analysis;
# - **scikit-learn**: a Machine Learning library in Python;
# - **NLTK** (Natural Language Toolkit): platform for building Python programs to work with human language data.
#
# Download Anaconda from the [official website](https://www.anaconda.com/products/individual) and follow the [instructions](https://docs.anaconda.com/anaconda/install/) for your OS.
#
# **conda** is the package and environment manager provided with Anaconda. From the [conda website](https://conda.io/en/latest/):
# > Conda is an open source **package management system** and **environment management system** that runs on Windows, macOS and Linux. Conda quickly installs, runs and updates packages and their dependencies. Conda easily creates, saves, loads and switches between environments on your local computer. It was created for Python programs, but it can package and distribute software for any language.
#
#
# ## Managing packages
#
# In programming, a module is a piece of software that has a specific functionality. In Java, the term *package* is often used as a synonym of module. In Python, a *package* is a collection of modules.
#
# - `conda` provides many commands for installing packages from the Anaconda repository and cloud.
# - `pip` is the Python Packaging Authority’s recommended tool for installing packages from the Python Package Index, PyPI.
#
# For example, you can install a given package by typing the following commands in the terminal:
# > ```conda install [package name]```
#
# or
#
# > ```pip install [package name]```
#
# ## Managing Environments
#
# Python has its own unique way of downloading, storing, and locating packages (or modules). There are a few different locations where these packages can be installed on your system.
#
# Third party packages, or **site packages**, installed using `pip` or `conda` are typically placed in one of the directories pointed to by `site.getsitepackages`:
# +
import site
print(site.getsitepackages())
# -
# By default, every project on your system will use these same directories to store and retrieve site packages.
# Site packages are stored according to just their name, there is no differentiation between versions.
#
# Consider the following scenario
# - you have two projects: *ProjectA* and *ProjectB*.
# - Both projects have a dependency on the same package, *ProjectC*.
# - *ProjectA* needs *ProjectC* `v1.0.0`.
# - *ProjectB* needs *ProjectC* `v2.0.0`.
#
# This is a real problem for Python since it can’t differentiate between versions in the `site-packages` directory. So both `v1.0.0` and `v2.0.0` would reside in the same directory with the same name.
#
# An **environment manager** allows to create lightweight "virtual environments" with their own site directories, optionally isolated from system site directories. This means that each project can have its own dependencies, regardless of what dependencies every other project has.
#
# In our example, we would just need to create a separate virtual environment for both *ProjectA* and *ProjectB*.
# Each environment, in turn, would be able to depend on whatever version of *ProjectC* they choose, independent of the other.
#
# Practically, **virtual environments are just directories containing a few scripts**.
#
# Availble tools:
# - `conda` provides many commands for managing environments
# - `venv` is the Python tool for managing virtual environments
#
# ## Differences between `conda`, `pip`, `virtualenv`
#
# Look at [the documentation](https://docs.conda.io/projects/conda/en/latest/commands.html#conda-vs-pip-vs-virtualenv-commands) for the full-size table, and at [this blog post](https://www.anaconda.com/blog/understanding-conda-and-pip).
# 
#
#
#
# ## Creating a conda environment
# 1. Create the environment named "aideLAB" using the following command (Unix shell or Anaconda prompt)
#
# ```bash
# $ conda create --name aideLAB python=3.9
# ```
#
# 2. Get the list of available environments
#
# ```bash
# $ conda env list
# ```
# 3. Activate your newly created environment
#
# ```bash
# $ conda activate aideLAB
# ```
#
# 4. Install `jupyter` (or any other needed package)
#
# ```bash
# (cyberLab)$ conda install jupyter
# ```
# or
#
# ```bash
# (cyberLab)$ pip install jupyter
# ```
#
# 5. If you need to go back to the "system context":
#
# ```bash
# (cyberLab)$ conda deactivate
# ```
#
#
# ## Jupyter Notebook Usage
#
# After installing jupyter using `conda` or `pip`, to start a Jupyter notebook server, navigate to a suitable working directory and type the following command (Unix shell or Anaconda prompt)
# ```bash
# (aideLAB)$ jupyter notebook
# ```
#
# This starts a Jupyter notebook server and automatically opens it in the browser. You should get something like this:
#
# 
#
# Here's a quick list of the main functionalities in Jupyter notebooks (have a look at `Help > Keyboard Shortcuts`)
# * Start a new notebook clicking `New` in the top-right corner
# * Type in some Python code in a **cell** and press `Shift` + `Enter` to execute.
# * Change the cell type from Code to Markdown using the drop-down box (top-middle) to write explanatory text. Markdown guidance is available [here](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet%20%22Markdown%20cheat-sheet%20on%20GitHub).
# * Add new cells using the `+` button (top-left).
# * Save the notebook using the disk button (top-left).
# * Run cells in various ways (run all, run selection, run all above, run all below, etc.) using the options in the `Cell` menu.
# * Interrupt the kernel, restart it, clear all output, etc. using the options in the `Kernel` menu.
# * Download as Python script, HTML, LaTeX, etc. using the options under `File > Download as`.
#
# An open notebook has exactly one interactive session connected to a kernel which will execute code sent by the user and communicate back results.
#
# This kernel remains active if the web browser window is closed, and reopening the same notebook from the dashboard will reconnect the web application to the same kernel.
# **To sum up**:
#
# You may want to use an integrated development environment (IDE), e.g. Spyder (comes with Anaconda installation), PyCharm, JupyterLab to name but a few, for the following reasons:
# - more complex projects (many files)
# - debug utilities
# - powerful code editing tools
#
# You may prefer Jupyter notebook:
# - for scripts and prototypes
# - for interactive devoloping
# - for visualization purpose
#
| 1_Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:fastai-cpu]
# language: python
# name: conda-env-fastai-cpu-py
# ---
# Eval scripts adapted from https://github.com/SpaceNetChallenge/utilities/tree/master/python
# +
import numpy as np
import geopandas as gpd
import rtree
from pathlib import Path
import matplotlib.pyplot as plt
import matplotlib as mpl
# %matplotlib inline
from tqdm import tqdm
# +
def create_rtree_from_poly(poly_list):
# create index
index = rtree.index.Index(interleaved=False)
for idx, building in enumerate(poly_list):
minx, miny, maxx, maxy = building.bounds
envelope = (minx, maxx, miny, maxy)
index.insert(idx, envelope)
return index
def search_rtree(test_building, index):
# input test poly ogr.Geometry and rtree index
if test_building.type == 'Polygon' or \
test_building.type == 'MultiPolygon':
minx, miny, maxx, maxy = test_building.bounds
envelope = (minx, maxx, miny, maxy)
fidlist = index.intersection(envelope)
else:
fidlist = []
return fidlist
# -
def iou(test_poly, truth_polys, truth_index=[]):
fidlistArray = []
iou_list = []
if truth_index:
fidlist = search_rtree(test_poly, truth_index)
for fid in fidlist:
if not test_poly.is_valid:
test_poly = test_poly.buffer(0.0)
intersection_result = test_poly.intersection(truth_polys[fid].buffer(0.0))
fidlistArray.append(fid)
if intersection_result.type == 'Polygon' or \
intersection_result.type == 'MultiPolygon':
intersection_area = intersection_result.area
union_area = test_poly.union(truth_polys[fid].buffer(0.0)).area
iou_list.append(intersection_area / union_area)
else:
iou_list.append(0)
else:
for idx, truth_poly in enumerate(truth_polys):
if not test_poly.is_valid or not truth_poly.is_valid:
test_poly = test_poly.buffer(0.0)
truth_poly = truth_poly.buffer(0.0)
# print(f'fixed geom error at {idx}')
intersection_result = test_poly.intersection(truth_poly)
#print(idx, intersection_result.type)
if intersection_result.type == 'Polygon' or \
intersection_result.type == 'MultiPolygon':
intersection_area = intersection_result.area
union_area = test_poly.union(truth_poly).area
iou_list.append(intersection_area / union_area)
# print(f'found intersect at test_poly {i} with truth poly {idx}')
# print(intersection_area/union_area)
else:
iou_list.append(0)
return iou_list, fidlistArray
# +
def score(test_polys, truth_polys, threshold=0.5, truth_index=[],
resultGeoJsonName = [],
imageId = []):
# Define internal functions
# Find detections using threshold/argmax/IoU for test polygons
true_pos_count = 0
false_pos_count = 0
truth_poly_count = len(truth_polys)
true_ids = []
false_ids = []
for idx, test_poly in tqdm(enumerate(test_polys)):
if truth_polys:
iou_list, fidlist = iou(test_poly, truth_polys, truth_index)
if not iou_list:
maxiou = 0
else:
maxiou = np.max(iou_list)
# print(maxiou, iou_list, fidlist)
if maxiou >= threshold:
true_pos_count += 1
true_ids.append(idx)
minx, miny, maxx, maxy = truth_polys[fidlist[np.argmax(iou_list)]].bounds
envelope = (minx, maxx, miny, maxy)
truth_index.delete(fidlist[np.argmax(iou_list)], envelope)
#del truth_polys[fidlist[np.argmax(iou_list)]]
else:
false_pos_count += 1
false_ids.append(idx)
else:
false_pos_count += 1
false_ids.append(idx)
false_neg_count = truth_poly_count - true_pos_count
return true_pos_count, false_pos_count, false_neg_count, true_ids, false_ids
# -
def evalfunction(image_id, test_polys, truth_polys, truth_index=[], resultGeoJsonName=[], threshold = 0.5):
if len(truth_polys)==0:
true_pos_count = 0
false_pos_count = len(test_polys)
false_neg_count = 0
else:
true_pos_count, false_pos_count, false_neg_count, true_ids, false_ids = score(test_polys, truth_polys,
truth_index=truth_index,
resultGeoJsonName=resultGeoJsonName,
imageId=image_id,
threshold=threshold
)
if (true_pos_count > 0):
precision = float(true_pos_count) / (float(true_pos_count) + float(false_pos_count))
recall = float(true_pos_count) / (float(true_pos_count) + float(false_neg_count))
F1score = 2.0 * precision * recall / (precision + recall)
else:
F1score = 0
return ((F1score, true_pos_count, false_pos_count, false_neg_count), true_ids, false_ids, image_id)
def precision_recall(true_pos_count, false_pos_count, false_neg_count):
precision = float(true_pos_count) / (float(true_pos_count) + float(false_pos_count))
recall = float(true_pos_count) / (float(true_pos_count) + float(false_neg_count))
return (precision, recall)
TRUTH = Path('znz-input')
TEST = Path('znz-20190118')
df_truth = gpd.read_file(f'{str(TRUTH)}/grid_042.geojson')
df_test = gpd.read_file(f'{str(TEST)}/grid_042_20190118_07_classes.geojson')
df_truth.head()
df_test.head()
df_truth.geometry.plot(figsize=(10,10))
df_test.geometry.plot(figsize=(10,10))
df_test['cat'].value_counts()
df_truth['condition'].value_counts()
cats = [('conf_foundation','Foundation'),('conf_unfinished','Incomplete'),('conf_completed','Complete')]
for (test_cat, truth_cat) in cats:
test_polys = [geom for geom in df_test[df_test['cat'] == test_cat].geometry]
truth_polys = [geom for geom in df_truth[df_truth['condition'] == truth_cat].geometry]
truth_index = create_rtree_from_poly(truth_polys)
scores = evalfunction(grid_num,test_polys, truth_polys, truth_index=truth_index)
print(truth_cat)
print(scores[0],precision_recall(*scores[0][1:]))
test_polys = [geom for geom in df_test.geometry]
truth_polys = [geom for geom in df_truth.geometry]
truth_index = create_rtree_from_poly(truth_polys)
scores = evalfunction(grid_num,test_polys, truth_polys, truth_index=truth_index)
scores[0],precision_recall(*scores[0][1:])
| archive/znz-eval-20190118.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A03. `TRIPOLpy` Tutorial
# Here I will reduce the 2019-05-03 TRIPOL observation.
#
# First, download and install [``TRIPOLpy`` package](https://github.com/ysBach/TRIPOLpy) (**please read the README** there).
#
# Refer to the [TRIPOL lecture note](https://github.com/ysBach/AO2019/blob/master/Notebooks/A02_TRIPOL.md) for how to access the data.
#
# Because there were some problems in the header of 2019-05-03 observation (``RET-ANG1``, which is the half-wave plate angle), you have to manually input the value. I chose this observation because it is rather different from usual TRIPOL data and you can experience how to overcome such problems within ``TRIPOLpy``.
#
# The contents below follow this format:
# ```
# 1. The code line is given.
# 2. Simple explanation of the code line.
# 3. The snippet of the original source code (for the full source code, please see TRIPOLpy)
# ```
#
# * **NOTE**: the ``self`` in the first arguments below are for the python ``class``. Please ignore it if you want to utilize the snippets I showed here to your ``function`` object.
# ## 1. Importing and Setting Up
# +
from astropy.io import fits
from pathlib import Path
from tripolpy import preprocessor
TOPPATH = Path('.')
RAWPATH = TOPPATH / "rawdata"
# -
# ## 2. Initiation
p = preprocessor.Preprocessor(topdir=TOPPATH, rawdir=RAWPATH)
# The above is identical to
#
# ```python
# USEFUL_KEYS = ["EXPTIME", "FILTER", "DATE-OBS", "RET-ANG1", "CCD_TEMP",
# "CCD_COOL", "OBJECT", "EPOCH", "RA", "DEC", "ALT", "AZ",
# "AIRMASS"]
# p = preprocessor.Preprocessor(topdir=TOPPATH, rawdir=RAWPATH,
# summary_keywords=USEFUL_KEYS)
# ```
#
# because the original source code (you can verify by yourself from the source code) is like this:
# ```python
# class Preprocessor():
# def __init__(self, topdir, rawdir, summary_keywords=USEFUL_KEYS):
# """
# Parameters
# ----------
# topdir : path-like
# The top directory of which all the other paths will be represented
# relative to.
#
# rawdir: path-like
# The directory where all the FITS files are stored (without any
# subdirectory)
#
# summary_keywords: list of str, optional
# The keywords of the header to be used for the summary table.
# """
# # code goes here
# ```
# ## 3. Organize the TRIPOL images.
p.organize_tripol(archive_dir=RAWPATH / "archive")
# This first **updates header** appropriately, and then **organizes** the files based on the header. It automatically finds bias, dark, flat, and object frames. For all such image types, it separates them first by filter and then by object name (``OBJECT`` key of the header).
#
# This is identical to use the following code:
# ```python
# rename_by = ["COUNTER", "FILTER", "OBJECT", "EXPOS", "RET-ANG1"]
# mkdir_by = ["FILTER", "OBJECT"]
# p.organize_tripol(rename_by=rename_by,
# mkdir_by=mkdir_by,
# delimiter='_',
# archive_dir=RAWPATH / "archive")
# ```
#
# This is because, by default, the ``.organize_tripol()`` method is defined as this:
#
# ```python
# def organize_tripol(self,
# rename_by=["COUNTER", "FILTER",
# "OBJECT", "EXPOS", "RET-ANG1"],
# mkdir_by=["FILTER", "OBJECT"], delimiter='_',
# archive_dir=None, verbose=False):
# ''' Rename FITS files after updating theur headers.
# Parameters
# ----------
# fpath: path-like
# The path to the target FITS file.
# rename_by: list of str
# The keywords in header to be used for the renaming of FITS files.
# Each keyword values are connected by ``delimiter``.
# mkdir_by: list of str, optional
# The keys which will be used to make subdirectories to classify
# files. If given, subdirectories will be made with the header value
# of the keys.
# delimiter: str, optional
# The delimiter for the renaming.
# archive_dir: path-like or None, optional
# Where to move the original FITS file. If ``None``, the original file
# will remain there. Deleting original FITS is dangerous so it is only
# supported to move the files. You may delete files manually if
# needed.
# '''
# # code goes here
# ```
#
# * **TIP**: Now see the raw data directory. You can see the data are re-arranged into many separate folders and now have meaningful file names. Also many folders will have been generated by now.
# ## 4. Bias Combine (Master Bias or Master Zero)
p.make_bias(savedir=TOPPATH / "calibration")
# This extracts only the bias frames from the previously organized files and do bias combine. Then the master bias is saved to ``savedir``.
#
# This is identical to the following code:
# ``` python
# MEDCOMB_KEYS = dict(overwrite=True,
# unit='adu',
# combine_method="median",
# reject_method=None,
# combine_uncertainty_function=None)
#
# p.make_bias(savedir=TOPPATH / "calibration",
# hdr_keys="OBJECT",
# hdr_vals="bias",
# group_by=["FILTER"],
# delimiter='_',
# dtype='float32',
# comb_kwargs=MEDCOMB_KEYS
# )
# ```
#
# Since the related source codes are too long, I only included core parts. If you are interested in, please check the original source code by yourself.
#
#
# ```python
# def make_bias(self, savedir=None, hdr_keys="OBJECT", hdr_vals="bias",
# group_by=["FILTER"], delimiter='_', dtype='float32',
# comb_kwargs=MEDCOMB_KEYS):
# ''' Finds and make bias frames.
# Parameters
# ----------
# savedir : path-like, optional.
# The directory where the frames will be saved.
#
# hdr_key : str or list of str, optional
# The header keys to be used for the identification of the bias
# frames. Each value should correspond to the same-index element of
# ``hdr_val``.
#
# hdr_val : str, float, int or list of such, optional
# The header key and values to identify the bias frames. Each value
# should correspond to the same-index element of ``hdr_key``.
#
# group_by : None, str or list str, optional.
# The header keywords to be used for grouping frames. For dark
# frames, usual choice can be ``['EXPTIME']``.
#
# delimiter : str, optional.
# The delimiter for the renaming.
#
# dtype : str or numpy.dtype object, optional.
# The data type you want for the final master bias frame. It is
# recommended to use ``float32`` or ``int16`` if there is no
# specific reason.
#
# comb_kwargs: dict or None, optional.
# The parameters for ``combine_ccd``.
# '''
# # code goes here
#
# ```
#
# ```python
# def combine_ccd(fitslist, trim_fits_section=None, output=None, unit='adu',
# subtract_frame=None, combine_method='median', reject_method=None,
# normalize_exposure=False, exposure_key='EXPTIME',
# combine_uncertainty_function=ccdproc_mad2sigma_func,
# extension=0, dtype=np.float32, type_key=None, type_val=None,
# output_verify='fix', overwrite=False,
# **kwargs):
# ''' Combining images
# # See original source code for the details.
# '''
# # code goes here
# ```
# ## 5. Dark Combine (Master Dark for Each Exposure Time)
p.make_dark(savedir=TOPPATH / "calibration")
# This code makes dark frames. The dark images are grouped by filter and exposure time.
#
# This is identical to the following code:
# ```python
# MEDCOMB_KEYS = dict(overwrite=True,
# unit='adu',
# combine_method="median",
# reject_method=None,
# combine_uncertainty_function=None)
#
# p.make_dark(savedir=TOPPATH / "calibration",
# hdr_keys="OBJECT",
# hdr_vals="dark",
# bias_sub=True,
# group_by=["FILTER", "EXPTIME"],
# bias_grouped_by=["FILTER"],
# exposure_key="EXPTIME",
# dtype='float32',
# delimiter='_',
# comb_kwargs=MEDCOMB_KEYS)
# ```
#
# The original method source code is like this:
#
# ```python
# def make_dark(self, savedir=None, hdr_keys="OBJECT", hdr_vals="dark",
# bias_sub=True,
# group_by=["FILTER", "EXPTIME"], bias_grouped_by=["FILTER"],
# exposure_key="EXPTIME", dtype='float32',
# delimiter='_', comb_kwargs=MEDCOMB_KEYS):
# """ Makes and saves dark (bias subtracted) images.
# Parameters
# ----------
# savedir: path-like, optional
# The directory where the frames will be saved.
#
# hdr_key : str or list of str, optional
# The header keys to be used for the identification of the bias
# frames. Each value should correspond to the same-index element of
# ``hdr_val``.
#
# hdr_val : str, float, int or list of such, optional
# The header key and values to identify the bias frames. Each value
# should correspond to the same-index element of ``hdr_key``.
#
# bias_sub: bool, optional
# If ``True``, subtracts bias from dark frames using self.biaspahts.
#
# group_by: None, str or list str, optional
# The header keywords to be used for grouping frames. For dark
# frames, usual choice can be ``['EXPTIME']``.
#
# bias_grouped_by: str or list of str, optional
# How the bias frames are grouped by.
#
# exposure_key: str, optional
# If you want to make bias from a list of dark frames, you need to
# let the function know the exposure time of the frames, so that the
# miniimum exposure time frame will be used as bias. Default is
# "EXPTIME".
#
# comb_kwargs: dict or None, optional
# The parameters for ``combine_ccd``.
# """
# # Code goes here
#
# ```
# ## 6. Flat Combine (Master Flat)
p.make_flat(savedir=TOPPATH / "calibration", group_by=["FILTER"])
# This makes flat images. First, it selects flat images for each filter. Then it again group by the half-wave plate angles.
#
# * **NOTE**: In 2019-05-03 observation, we took only half-wave plate angle of 0 degree. To use this flat throughout all the half-wave plate angles (which is not desired but the best way we can do given the data set), we forced the code **not** to distinguish flats based on the angle (header keyword is ``RET-ANG1``).
#
# The above is identical to the following code:
# ```python
# MEDCOMB_KEYS = dict(overwrite=True,
# unit='adu',
# combine_method="median",
# reject_method=None,
# combine_uncertainty_function=None)
#
# p.make_flat(savedir=TOPPATH / "calibration", group_by=["FILTER"])
# ```
#
# The original source code looks like below. As you can see, if you use ``.make_flat()`` without specifying ``group_by``, it will by default use ``group_by=["FILTER", "RET-ANG1"]``.
#
# ```python
# def make_flat(self, savedir=None,
# hdr_keys=["OBJECT"], hdr_vals=["flat"],
# group_by=["FILTER", "RET-ANG1"],
# bias_sub=True, dark_sub=True,
# bias_grouped_by=["FILTER"],
# dark_grouped_by=["FILTER", "EXPTIME"],
# exposure_key="EXPTIME",
# comb_kwargs=MEDCOMB_KEYS, delimiter='_', dtype='float32'):
# '''Makes and saves flat images.
# Parameters
# ----------
# savedir: path-like, optional
# The directory where the frames will be saved.
#
# hdr_key : str or list of str, optional
# The header keys to be used for the identification of the bias
# frames. Each value should correspond to the same-index element of
# ``hdr_val``.
#
# hdr_val : str, float, int or list of such, optional
# The header key and values to identify the bias frames. Each value
# should correspond to the same-index element of ``hdr_key``.
#
# bias_sub, dark_sub : bool, optional
# If ``True``, subtracts bias and dark frames using ``self.biaspahts``
# and ``self.darkpaths``.
#
# group_by: None, str or list str, optional
# The header keywords to be used for grouping frames. For dark
# frames, usual choice can be ``['EXPTIME']``.
#
# bias_grouped_by, dark_grouped_by : str or list of str, optional
# How the bias and dark frames are grouped by.
#
# exposure_key: str, optional
# If you want to make bias from a list of dark frames, you need to
# let the function know the exposure time of the frames, so that the
# miniimum exposure time frame will be used as bias. Default is
# "EXPTIME".
#
# comb_kwargs: dict or None, optional
# The parameters for ``combine_ccd``.
# '''
# # Code goes here
# ```
# ## 7. Preprocess at Once
summary_reduced = p.do_preproc(savedir=TOPPATH / "processed",
flat_grouped_by=["FILTER"])
# It now does the preprocessing and summarize the processed images as ``astropy Table`` object.
#
# It is identical to use
# ```python
# summary_reduced = p.do_preproc(savedir=TOPPATH / "processed",
# delimiter='_',
# dtype='float32',
# bias_grouped_by=["FILTER"],
# dark_grouped_by=["FILTER", "EXPTIME"],
# flat_grouped_by=["FILTER"],
# verbose_bdf=True,
# verbose_summary=False)
# ```
#
# Note that, the original source code (see below), ``flat_grouped_by=["FILTER", "RET-ANG1"]`` by default. As I did **NOT** use ``"RET-ANG1"`` (the half-wave plate angle) for the flat grouping, I had to use ``flat_grouped_by=["FILTER"]``.
#
# ```python
# def do_preproc(self, savedir=None, delimiter='_', dtype='float32',
# bias_grouped_by=["FILTER"],
# dark_grouped_by=["FILTER", "EXPTIME"],
# flat_grouped_by=["FILTER", "RET-ANG1"],
# verbose_bdf=True, verbose_summary=False):
# ''' Conduct the preprocessing using simplified ``bdf_process``.
# Parameters
# ----------
# savedir: path-like, optional
# The directory where the frames will be saved.
#
# delimiter : str, optional.
# The delimiter for the renaming.
#
# dtype : str or numpy.dtype object, optional.
# The data type you want for the final master bias frame. It is
# recommended to use ``float32`` or ``int16`` if there is no
# specific reason.
#
# bias_grouped_by, dark_grouped_by : str or list of str, optional
# How the bias, dark, and flat frames are grouped by.
# '''
# # Code goes here
# ```
#
# ## 8. View the Summary
# Finally, print out the results:
summary_reduced
# * **TIP**: You can view the resulting summary file with, e.g., Excel, by ``[your_RAWPATH]/summary_reduced.csv``.
| Notebooks/A03_TRIPOLpy_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from os import listdir
from os.path import isfile, join
# Read all results from txt files
grad_path = "./raw_files/GRAD/"
grad_files = [f for f in listdir(grad_path) if isfile(join(grad_path, f))]
gradl_path = "./raw_files/GRAD_L/"
gradl_files = [f for f in listdir(gradl_path) if isfile(join(gradl_path, f))]
gradh5_path = "./raw_files/GRAD_H5/"
gradh5_files = [f for f in listdir(gradh5_path) if isfile(join(gradh5_path, f))]
# GRAD Algorithm
column_names = ["Data-set", "Size", "Algorithm", "Support", "Run-time", "Memory", "Patterns"]
df_grad = pd.DataFrame(columns = column_names)
for file in grad_files:
f_path = join(grad_path,file)
res = pd.read_csv(f_path, sep = ': ', header=None, engine='python', nrows=8)
run = float(res[1][0][0:6]) # run-time
mem = float(res[1][1][0:5]) # memory
pat = float(res[1][7]) # patterns
alg = res[1][2] # algorithm
att = int(res[1][3]) # number of attributes in the data set
sup = float(res[1][5]) # minimum support
size = int(res[1][4]) # data set size
if att == 98:
col = "C2K"
elif att == 9:
col = "UCI"
else:
col = ""
df_grad = df_grad.append({"Data-set": col, "Size": size, "Algorithm":alg, "Support": sup, "Run-time":run, "Memory":mem, "Patterns":pat}, ignore_index=True)
# GRAD-H5 Algorithm
# column_names = ["Data-set", "Algorithm", "Support", "Run-time", "Memory", "Patterns"]
df_gradh5 = pd.DataFrame(columns = column_names)
for file in gradh5_files:
f_path = join(gradh5_path,file)
res = pd.read_csv(f_path, sep = ': ', header=None, engine='python', nrows=11)
run = float(res[1][0][0:6]) # run-time
mem = float(res[1][1][0:5]) # memory
pat = float(res[1][10]) # patterns
alg = res[1][2] # algorithm
att = int(res[1][5]) # number of attributes in the data set
sup = float(res[1][3]) # minimum support
size = int(res[1][6]) # data set size
if att == 98:
col = "C2K"
elif att == 9:
col = "UCI"
else:
col = ""
df_gradh5 = df_gradh5.append({"Data-set": col, "Size": size, "Algorithm":alg, "Support": sup, "Run-time":run, "Memory":mem, "Patterns":pat}, ignore_index=True)
#GRAD-L Algorithm
column_names = ["Data-set", "Size", "Algorithm", "Support", "Chunk-size", "Run-time", "Memory", "Patterns"]
df_gradl = pd.DataFrame(columns = column_names)
for file in gradl_files:
f_path = join(gradl_path,file)
res = pd.read_csv(f_path, sep = ': ', header=None, engine='python', nrows=12)
run = float(res[1][0][0:6]) # run-time
mem = float(res[1][1][0:5]) # memory
pat = float(res[1][11]) # patterns
alg = res[1][2] # algorithm
att = int(res[1][5]) # number of attributes in the data set
sup = float(res[1][3]) # minimum support
size = int(res[1][6]) # data set size
chk = int(res[1][7]) # chunk size
if att == 98:
col = "C2K"
elif att == 9:
col = "UCI"
else:
col = ""
df_gradl = df_gradl.append({"Data-set": col, "Size": size, "Algorithm":alg, "Support": sup, "Chunk-size": chk, "Run-time":run, "Memory":mem, "Patterns":pat}, ignore_index=True)
# +
# df_grad
# +
# df_gradh5
# +
# df_gradl
# +
# Combining all results into one data-frame
frames = [df_grad, df_gradh5, df_gradl]
df_res = pd.concat(frames)
df_res
# +
# Describing the results
# df_res.describe()
# df_res.groupby(["Data-set", "Support", "Algorithm"]).describe().to_excel("stats.xlsx", sheet_name="Stats")
df_res.groupby(["Data-set", "Size", "Algorithm"]).describe(percentiles=[])
# +
# Ignore these columns
# df_res = df_res.drop(['Support', 'Chunk-size'], axis=1)
df = df_res.groupby(["Data-set", "Size", "Algorithm"])#.describe(percentiles=[])
df2 = pd.concat([df.min(), df.mean(), df.max(), df.std()], keys=['min', 'mean', 'max', 'std'], axis=1)
#df2.filter(like="Run-time")
df2.columns = df2.columns.swaplevel(0, 1)
df2.sort_index(axis=1, level=0, inplace=True, ascending=False)
df2.round(3)
# -
| results/large_gps/.ipynb_checkpoints/analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Difference of Gaussians Application
# ----
#
# <div class="alert alert-box alert-info">
# Please use Jupyter labs http://<board_ip_address>/lab for this notebook.
# </div>
#
# This notebook shows how to download and play with the Difference of Gaussians Application
#
# ## Aims
# * Instantiate the application
# * Start the application
# * Play with the runtime parameters
# * Stop the application
#
# ## Table of Contents
# * [Download Composable Overlay](#download)
# * [Start Application](#start)
# * [Play with the Application](#play)
# * [Stop Application](#stop)
# * [Conclusion](#conclusion)
#
# ----
#
# ## Revision History
#
# * v1.0 | 30 March 2021 | First notebook revision.
#
# ----
# ## Download Composable Overlay <a class="anchor" id="download"></a>
#
# Download the Composable Overlay using the `DifferenceGaussians` class which wraps all the functionality needed to run this application
# +
from composable_pipeline import DifferenceGaussians
app = DifferenceGaussians("../overlay/cv_dfx_4_pr.bit")
# -
# ## Start Application <a class="anchor" id="start"></a>
#
# Start the application by calling the `.start()` method, this will:
#
# 1. Initialize the pipeline
# 1. Setup initial parameters
# 1. Display the implemented pipelined
# 1. Configure HDMI in and out
#
# The output image should be visible on the external screen at this point
#
# <div class="alert alert-heading alert-danger">
# <h4 class="alert-heading">Warning:</h4>
#
# Failure to connect HDMI cables to a valid video source and screen may cause the notebook to hang
# </div>
app.start()
# ## Play with the Application <a class="anchor" id="play"></a>
#
# The `.play` attribute exposes the $\sigma$ parameters of the Difference of Gaussians application to play with.
#
# Move the sliders and notice how the output video changes.
app.play
# ## Stop Application <a class="anchor" id="stop"></a>
#
# Finally stop the application to release the resources
#
# <div class="alert alert-heading alert-danger">
# <h4 class="alert-heading">Warning:</h4>
#
# Failure to stop the HDMI Video may hang the board
# when trying to download another bitstream onto the FPGA
# </div>
app.stop()
# ----
#
# ## Conclusion <a class="anchor" id="conclusion"></a>
#
# This notebook has presented the Difference of Gaussian application that leverages the Composable Overlay.
#
# The runtime parameters of such application can be modified using sliders from `ipywidgets`
#
# | | [Corner Detect Application ➡️](02_corner_detect_app.ipynb)
# Copyright © 2021 Xilinx, Inc
#
# SPDX-License-Identifier: BSD-3-Clause
#
# ----
| composable_pipeline/notebooks/applications/01_difference_gaussians_app.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Symmetrize
#
# In this notebook, we will use the _symmetrize_ function to create bi-directional edges in an undirected graph
#
# Notebook Credits
# * Original Authors: <NAME> and <NAME>
# * Created: 08/13/2019
# * Updated: 10/28/2019
#
# RAPIDS Versions: 0.10.0
#
# Test Hardware
#
# * GV100 32G, CUDA 10.0
#
#
# ## Introduction
# In many cases, an Undirected graph is saved as a single edge between vertex pairs. That saves a lot of space in the data file. However, in order to process that data in cuGraph, there needs to be an edge in each direction for undirected. Converting from a single edge to two edges, one in each direction, is called symmetrization.
#
# To symmerize an edge list (COO data) use:<br>
#
# **cugraph.symmetrize(source, destination, value)**
# * __source__: cudf.Series
# * __destination__: cudf.Series
# * __value__: cudf.Series
#
#
# Returns:
# * __triplet__: three variables are returned:
# * __source__: cudf.Series
# * __destination__: cudf.Series
# * __value__: cudf.Series
#
# ### Test Data
# We will be using an undirected unsymmetrized version of the Zachary Karate club dataset. The result of symmetrization shopuld be a dataset equal to to the version used in the PageRank notebook.
#
# *<NAME>, An information flow model for conflict and fission in small groups, Journal of
# Anthropological Research 33, 452-473 (1977).*
#
#
# 
#
# Import needed libraries
import cugraph
import cudf
# Read the unsymmetrized data
unsym_data ='../data/karate_undirected.csv'
gdf = cudf.read_csv(unsym_data, names=["src", "dst"], delimiter='\t', dtype=["int32", "int32"] )
# load the full symmetrized dataset for comparison
datafile='../data/karate-data.csv'
test_gdf = cudf.read_csv(datafile, names=["src", "dst"], delimiter='\t', dtype=["int32", "int32"] )
print("Unsymmetrized Graph")
print("\tNumber of Vertices: " + str(len(gdf)))
print("Baseline Graph")
print("\tNumber of Vertices: " + str(len(test_gdf)))
# _Since the unsymmetrized graph only has one edge between vertices, that underlying code treats that as a directed graph_
G = cugraph.Graph()
G.from_cudf_edgelist(gdf, source='src', destination='dst')
gdf_page = cugraph.pagerank(G)
# best PR score is
m = gdf_page['pagerank'].max()
df = gdf_page.query('pagerank == @m')
df
# ### Now Symmetrize the dataset
df = cugraph.symmetrize_df(gdf, 'src', 'dst')
print("Unsymmetrized Graph")
print("\tNumber of Vertices: " + str(len(gdf)))
print("Symmetrized Graph")
print("\tNumber of Vertices: " + str(len(df)))
print("Baseline Graph")
print("\tNumber of Vertices: " + str(len(test_gdf)))
| notebooks/structure/Symmetrize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from glm.families import Gaussian, Bernoulli, Poisson, Gamma
from glm.glm import GLM
from glm.simulation import Simulation
import statsmodels.api as sm
import statsmodels
# -
N = 1000
X = np.empty(shape=(N, 3))
X[:, 0] = 1.0
X[:, 1] = np.random.uniform(-1, 1, size=N)
X[:, 2] = np.random.uniform(-1, 1, size=N)
nu = 1 - 2*X[:, 1] + np.random.normal(0.0, 1.0, size=N)
# ## Linear Model
y = nu + np.random.normal(size=N)
model = GLM(family=Gaussian())
model.fit(X, y)
model.coef_
model.coef_covariance_matrix_
model.coef_standard_error_
model.p_values_
yhat = model.predict(X)
yhat[:10]
mod = sm.OLS(y, X)
res = mod.fit()
print(res.summary())
res.params
res.bse
# ## Linear Regression With Formula
df = pd.DataFrame(
np.concatenate([X[:, 1:], y.reshape(-1, 1)], axis=1),
columns=['Moshi', 'SwimSwim', 'y'])
df.shape
model = GLM(family=Gaussian())
model.fit(df, formula='y ~ Moshi + SwimSwim')
model.coef_
model.summary()
yhat = model.predict(df)
yhat[:10]
model.formula
# ### Simulation with Model Formula
from glm.simulation import Simulation
sim = Simulation(model)
df.shape
boots = sim.non_parametric_bootstrap(df, n_sim=5)
boots[0].X_names
# ## Run some simulations off the linear model.
s = Simulation(model)
s.sample(X)
models = s.parametric_bootstrap(X, n_sim=10)
for model in models:
print(model.coef_)
models = s.non_parametric_bootstrap(X, y, n_sim=10)
for model in models:
print(model.coef_)
# ## Linear Model with Sample Weights
sample_weights = np.random.uniform(0, 2, size=N)
model = GLM(family=Gaussian())
model = model.fit(X, y, sample_weights=sample_weights)
model.coef_
# ## Logistic Model
p = 1 / (1 + np.exp(-nu))
y_logistic = np.random.binomial(1, p=p, size=N)
model = GLM(family=Bernoulli())
model.fit(X, y_logistic)
model.coef_
model.dispersion_
model.coef_covariance_matrix_
model.coef_standard_error_
model.p_values_
mod = sm.Logit(y_logistic, X)
res = mod.fit()
print(res.summary())
s = Simulation(model)
s.sample(X, n_sim=10)
for model in s.parametric_bootstrap(X, n_sim=10):
print(model.coef_)
for model in s.non_parametric_bootstrap(X, y_logistic, n_sim=10):
print(model.coef_)
# ## Poission Model
mu = np.exp(nu)
y_poisson = np.random.poisson(lam=mu, size=N)
model = GLM(family=Poisson())
model.fit(X, y_poisson)
model.coef_
model.coef_covariance_matrix_
model.coef_standard_error_
mod = statsmodels.discrete.discrete_model.Poisson(y_poisson, X)
res = mod.fit()
print(res.summary())
s = Simulation(model)
s.sample(X, n_sim=10)
for model in s.parametric_bootstrap(X, n_sim=10):
print(model.coef_)
for model in s.non_parametric_bootstrap(X, y_poisson, n_sim=10):
print(model.coef_)
# ## Poisson with Exposures
mu = np.exp(nu)
expos = np.random.uniform(0, 10, size=N)
y_poisson = np.random.poisson(lam=(mu*expos), size=N)
model = GLM(family=Poisson())
model.fit(X, y_poisson, offset=np.log(expos))
model.coef_
model.coef_standard_error_
# ## Gamma Regression
mu = np.exp(nu)
y_gamma = np.random.gamma(shape=2.0, scale=(mu / 2.0), size=N)
gamma_model = GLM(family=Gamma())
gamma_model.fit(X, y_gamma)
gamma_model.coef_
gamma_model.coef_standard_error_
gamma_model.dispersion_
gamma_model = sm.GLM(y_gamma, X,
family=sm.families.Gamma(
link=statsmodels.genmod.families.links.log))
res = gamma_model.fit()
print(res.summary())
# ## Exponential Regression
mu = np.exp(nu)
y_exponential = np.random.exponential(scale=mu, size=N)
exponential_model = GLM(family=Gamma())
exponential_model.fit(X, y_exponential)
exponential_model.coef_
exponential_model.coef_standard_error_
exponential_model.dispersion_
exponential_model = sm.GLM(y_exponential, X,
family=sm.families.Gamma(
link=statsmodels.genmod.families.links.log))
res = exponential_model.fit()
print(res.summary())
# ## Linear Model with Correlated Predictors
N = 1000
X = np.empty(shape=(N, 3))
X[:, 0] = 1.0
X[:, 1] = np.random.uniform(size=N)
X[:, 2] = 0.5*X[:, 1] + np.random.uniform(-0.5, 0.5, size=N)
nu = 1 - 2*X[:, 1] + X[:, 2]
y = nu + np.random.normal(size=N)
model = GLM(family=Gaussian())
model.fit(X, y)
model.coef_
model.coef_covariance_matrix_
model.coef_standard_error_
mod = sm.OLS(y, X)
res = mod.fit()
print(res.summary())
| examples/glm-examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deploying a MedNIST Classifier with BentoML
#
# This notebook demos the process of packaging up a trained model using BentoML into an artifact which can be run as a local program performing inference, a web service doing the same, and a Docker containerized web service. BentoML provides various ways of deploying models with existing platforms like AWS or Azure but we'll focus on local deployment here since researchers are more likely to do this. This tutorial will train a MedNIST classifier like the [MONAI tutorial here](https://github.com/Project-MONAI/tutorials/blob/master/2d_classification/mednist_tutorial.ipynb) and then do the packaging as described in this [BentoML tutorial](https://github.com/bentoml/gallery/blob/master/pytorch/fashion-mnist/pytorch-fashion-mnist.ipynb).
# ## Setup environment
# !python -c "import monai" || pip install -q "monai-weekly[pillow, tqdm]"
# !python -c "import bentoml" || pip install -q bentoml
# ## Setup imports
# + tags=[]
# Copyright 2020 MONAI Consortium
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import os
import shutil
import tempfile
import glob
import PIL.Image
import torch
import numpy as np
from ignite.engine import Events
from monai.apps import download_and_extract
from monai.config import print_config
from monai.networks.nets import DenseNet121
from monai.engines import SupervisedTrainer
from monai.transforms import (
AddChannel,
Compose,
LoadImage,
RandFlip,
RandRotate,
RandZoom,
ScaleIntensity,
EnsureType,
)
from monai.utils import set_determinism
set_determinism(seed=0)
print_config()
# -
# ## Download dataset
#
# The MedNIST dataset was gathered from several sets from [TCIA](https://wiki.cancerimagingarchive.net/display/Public/Data+Usage+Policies+and+Restrictions),
# [the RSNA Bone Age Challenge](http://rsnachallenges.cloudapp.net/competitions/4),
# and [the NIH Chest X-ray dataset](https://cloud.google.com/healthcare/docs/resources/public-datasets/nih-chest).
#
# The dataset is kindly made available by [Dr. <NAME>., Ph.D.](https://www.mayo.edu/research/labs/radiology-informatics/overview) (Department of Radiology, Mayo Clinic)
# under the Creative Commons [CC BY-SA 4.0 license](https://creativecommons.org/licenses/by-sa/4.0/).
#
# If you use the MedNIST dataset, please acknowledge the source.
# + tags=[]
directory = os.environ.get("MONAI_DATA_DIRECTORY")
root_dir = tempfile.mkdtemp() if directory is None else directory
print(root_dir)
resource = "https://github.com/Project-MONAI/MONAI-extra-test-data/releases/download/0.8.1/MedNIST.tar.gz"
md5 = "0bc7306e7427e00ad1c5526a6677552d"
compressed_file = os.path.join(root_dir, "MedNIST.tar.gz")
data_dir = os.path.join(root_dir, "MedNIST")
if not os.path.exists(data_dir):
download_and_extract(resource, compressed_file, root_dir, md5)
# +
subdirs = sorted(glob.glob(f"{data_dir}/*/"))
class_names = [os.path.basename(sd[:-1]) for sd in subdirs]
image_files = [glob.glob(f"{sb}/*") for sb in subdirs]
image_files_list = sum(image_files, [])
image_class = sum(([i] * len(f) for i, f in enumerate(image_files)), [])
image_width, image_height = PIL.Image.open(image_files_list[0]).size
print(f"Label names: {class_names}")
print(f"Label counts: {list(map(len, image_files))}")
print(f"Total image count: {len(image_class)}")
print(f"Image dimensions: {image_width} x {image_height}")
# -
# ## Setup and Train
#
# Here we'll create a transform sequence and train the network, omitting validation and testing since we know this does indeed work and it's not needed here:
train_transforms = Compose(
[
LoadImage(image_only=True),
AddChannel(),
ScaleIntensity(),
RandRotate(range_x=np.pi / 12, prob=0.5, keep_size=True),
RandFlip(spatial_axis=0, prob=0.5),
RandZoom(min_zoom=0.9, max_zoom=1.1, prob=0.5),
EnsureType(),
]
)
# +
class MedNISTDataset(torch.utils.data.Dataset):
def __init__(self, image_files, labels, transforms):
self.image_files = image_files
self.labels = labels
self.transforms = transforms
def __len__(self):
return len(self.image_files)
def __getitem__(self, index):
return self.transforms(self.image_files[index]), self.labels[index]
# just one dataset and loader, we won't bother with validation or testing
train_ds = MedNISTDataset(image_files_list, image_class, train_transforms)
train_loader = torch.utils.data.DataLoader(train_ds, batch_size=300, shuffle=True, num_workers=10)
# -
device = torch.device("cuda:0")
net = DenseNet121(spatial_dims=2, in_channels=1, out_channels=len(class_names)).to(device)
loss_function = torch.nn.CrossEntropyLoss()
opt = torch.optim.Adam(net.parameters(), 1e-5)
max_epochs = 5
# +
def _prepare_batch(batch, device, non_blocking):
return tuple(b.to(device) for b in batch)
trainer = SupervisedTrainer(device, max_epochs, train_loader, net, opt, loss_function, prepare_batch=_prepare_batch)
@trainer.on(Events.EPOCH_COMPLETED)
def _print_loss(engine):
print(f"Epoch {engine.state.epoch}/{engine.state.max_epochs} Loss: {engine.state.output[0]['loss']}")
trainer.run()
# -
# The network will be saved out here as a Torchscript object but this isn't necessary as we'll see later.
torch.jit.script(net).save("classifier.zip")
# ## BentoML Setup
#
# BentoML provides it's platform through an API to wrap service requests as method calls. This is obviously similar to how Flask works (which is one of the underlying technologies used here), but on top of this is provided various facilities for storing the network (artifacts), handling the IO component of requests, and caching data. What we need to provide is a script file to represent the services we want, BentoML will take this with the artifacts we provide and store this in a separate location which can be run locally as well as uploaded to a server (sort of like Docker registries).
#
# The script below will create our API which includes MONAI code. The transform sequence needs a special read Transform to turn a data stream into an image, but otherwise the code like what was used above for training. The network is stored as an artifact which in practice is the stored weights in the BentoML bundle. This is loaded at runtime automatically, but instead we could load the Torchscript model instead if we wanted to, in particular if we wanted an API that didn't rely on MONAI code.
#
# The script needs to be written out to a file first:
# +
# %%writefile mednist_classifier_bentoml.py
from typing import BinaryIO, List
import numpy as np
from PIL import Image
import torch
from monai.transforms import (
AddChannel,
Compose,
Transform,
ScaleIntensity,
EnsureType,
)
import bentoml
from bentoml.frameworks.pytorch import PytorchModelArtifact
from bentoml.adapters import FileInput, JsonOutput
from bentoml.utils import cached_property
MEDNIST_CLASSES = ["AbdomenCT", "BreastMRI", "CXR", "ChestCT", "Hand", "HeadCT"]
class LoadStreamPIL(Transform):
"""Load an image file from a data stream using PIL."""
def __init__(self, mode=None):
self.mode = mode
def __call__(self, stream):
img = Image.open(stream)
if self.mode is not None:
img = img.convert(mode=self.mode)
return np.array(img)
@bentoml.env(pip_packages=["torch", "numpy", "monai", "pillow"])
@bentoml.artifacts([PytorchModelArtifact("classifier")])
class MedNISTClassifier(bentoml.BentoService):
@cached_property
def transform(self):
return Compose([LoadStreamPIL("L"), AddChannel(), ScaleIntensity(), EnsureType()])
@bentoml.api(input=FileInput(), output=JsonOutput(), batch=True)
def predict(self, file_streams: List[BinaryIO]) -> List[str]:
img_tensors = list(map(self.transform, file_streams))
batch = torch.stack(img_tensors).float()
with torch.no_grad():
outputs = self.artifacts.classifier(batch)
_, output_classes = outputs.max(dim=1)
return [MEDNIST_CLASSES[oc] for oc in output_classes]
# -
# Now the script is loaded and the classifier artifact is packed with the network's state. This is then saved to a repository directory on the local machine:
# +
from mednist_classifier_bentoml import MedNISTClassifier # noqa: E402
bento_svc = MedNISTClassifier()
bento_svc.pack('classifier', net.cpu().eval())
saved_path = bento_svc.save()
print(saved_path)
# -
# We can look at the contents of this repository, which includes code and setup scripts:
# !ls -l {saved_path}
# This repository can be run like a stored program where we invoke it by name and the API name ("predict") we want to use and provide the inputs as a file:
# !bentoml run MedNISTClassifier:latest predict --input-file {image_files[0][0]}
# The service can also be run off of a Flask web server. The following script starts the service, waits for it to get going, uses curl to send the test file as a POST request to get a prediction, then kill the server:
# + magic_args="-s {image_files[0][0]}" language="bash"
# # filename passed in as an argument to the cell
# test_file=$1
#
# # start the Flask-based server, sending output to /dev/null for neatness
# bentoml serve --port=8000 MedNISTClassifier:latest &> /dev/null &
#
# # recall the PID of the server and wait for it to start
# lastpid=$!
# sleep 5
#
# # send the test file using curl and capture the returned string
# result=$(curl -s -X POST "http://127.0.0.1:8000/predict" -F image=@$test_file)
# # kill the server
# kill $lastpid
#
# echo "Prediction: $result"
# -
# The service can be packaged as a Docker container to be started elsewhere as a server:
# !bentoml containerize MedNISTClassifier:latest -t mednist-classifier:latest
# !docker image ls
if directory is None:
shutil.rmtree(root_dir)
| deployment/bentoml/mednist_classifier_bentoml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# + [markdown] id="dmmvuIksue4c"
# - **Programmer:**
# - **<NAME>**
# - **<NAME>**
#
# ## **Assigment 3**
# **Brief overview of assignment**
#
# ***perform a penalized
# (regularized) logistic (multinomial) regression fit using ridge regression, with the model
# parameters obtained by batch gradient descent. Your predictions will be based on K=5
# continental ancestries (African, European, East Asian, Oceanian, or Native American). Ridge
# regression will permit you to provide parameter shrinkage (tuning parameter 𝜆=0) to mitigate
# overfitting. The tuning parameter 𝜆 will be chosen using five-fold cross validation, and the best-
# fit model parameters will be inferred on the training dataset conditional on an optimal tuning
# parameter. This trained model will be used to make predictions on new test data points***
# + [markdown] id="fZ2NSNZaue4h"
# > ***Table of Contents***
# <hr>
#
#
# * [Import Packages](#Import_Packages)
# * [Import packages for manipulating data](#Import_packages_for_manipulating_data)
# * [Import packages for splitting data](#Import_packages_for_splitting_data)
# * [Import packages for modeling data](#Import_packages_for_modeling_data)
# * [Import packages for Scaling and Centering data](#Import_packages_for_Scaling_and_Centering_data)
# * [Import packages for Measuring Model Perormance](#Import_packages_for_Measuring_Model_Perormance)
#
# * [Data Processing](#Data_Processing)
# * [Import Data](#Import_data)
# * [Lets change the categorical values](#Lets_change_the_categorical_values)
# * [Create Predictor and Target numpy array](#Create_Predictor_and_Target_numpy_array)
# * [Create a Normalize copy of variables](#Create_a_Normalize_copy_of_variables)
# * [Split Data](#Split_Data:)
# * [Regression Model](#Regression_Model)
# * [Define our learning rates:](#Define_our_learning_rates)
# * [Create the Regression Objects](#Create_the_Regression_Objects)
# * [LogisticRegression Library](#LogisticRegression_Library)
#
#
# + [markdown] id="QAiw7MS9ue4j"
# > ***Deliverables***
# <hr>
#
# * [**Deliverable 6.1**](#Deliverable_6.1)
# * [**Deliverable 6.2**](#Deliverable_6.2)
# * [**Deliverable 6.3**](#Deliverable_6.3)
# * [**Deliverable 6.4**](#Deliverable_6.4)
# * [**Deliverable 6. Reason for difference**](#Deliverable_6_Reason_for_difference)
# + [markdown] id="2qUFlnh1ue4j"
# # Import Packages <a class="anchor" id="Import_Packages"></a>
# + [markdown] id="PB-AwZVDue4k"
# ### Import packages for manipulating data <a class="anchor" id="Import_packages_for_manipulating_data"></a>
# + id="6cDGv8Khue4k"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
import matplotlib.mlab as mlab
import math
import csv
import random
# %matplotlib inline
from sklearn.preprocessing import LabelEncoder
import seaborn as sns
import math
# + [markdown] id="X0gW1nTsue4m"
# ### Import packages for splitting data <a class="anchor" id="Import_packages_for_splitting_data"></a>
# + id="hjWloQNnue4n"
from sklearn.model_selection import train_test_split, cross_val_score, KFold, StratifiedKFold
from sklearn.model_selection import GridSearchCV
# + [markdown] id="Tqgw1bVFue4n"
# ### Import packages for modeling data <a class="anchor" id="Import_packages_for_modeling_data"></a>
# + id="FvZMHjvmue4o"
# Import models:
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression as linearR_Model, Ridge as RidgeR_Model
from sklearn.linear_model import RidgeCV
from sklearn.linear_model import ElasticNet
from sklearn.linear_model import ElasticNetCV
from sklearn.linear_model import LogisticRegression
from sklearn.exceptions import ConvergenceWarning
#from sklearn.utils._testing import ignore_warnings
import warnings
warnings.filterwarnings('ignore', category=ConvergenceWarning) # To filter out the Convergence warning
warnings.filterwarnings('ignore', category=UserWarning)
from itertools import product
# + [markdown] id="Ee6QUGCSue4p"
# ### Import packages for Scaling and Centering data <a class="anchor" id="Import_packages_for_Scaling_and_Centering_data"></a>
# + id="ZZ3nOXd8ue4p"
from sklearn.preprocessing import StandardScaler
# + [markdown] id="awr1dcfxue4p"
# ### Import packages for Measuring Model Perormance <a class="anchor" id="Import_packages_for_Measuring_Model_Perormance"></a>
# + id="IO-wwpPMue4q"
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
from sklearn.metrics import make_scorer
# + [markdown] id="3owCV2k0ue4q"
# # Data Processing <a class="anchor" id="Data_Processing"></a>
# + [markdown] id="SfaotWjJue4q"
# ### Import Data <a class="anchor" id="Import_data"></a>
# + [markdown] id="x86tPdyCue4r"
# ***Traing Dataset***
# + colab={"base_uri": "https://localhost:8080/", "height": 144} id="Pd6-H6lfue4r" outputId="9a4c562d-f7dd-4df4-a1e0-a2548d9d90a8"
Train_dataset = pd.read_csv ('TrainingData_N183_p10.csv')
Train_dataset.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="csjaBU1Cue4s" outputId="6315056e-792c-49c7-9c5c-76d3c0346bed"
# What are the datatypes of each observation:
print(Train_dataset.dtypes)
# Shape of my data
print('The size of our data are: ',Train_dataset.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="DLVGuKQ5ue4s" outputId="f016833e-31d1-48f2-cdc4-3ef2fa9ba841"
print('Training Dataset Missing Values: \n',Train_dataset.isnull().sum())
# + [markdown] id="Te0ntT70ue4s"
# ***Test Dataset***
# + colab={"base_uri": "https://localhost:8080/", "height": 144} id="mi1k5xQYue4t" outputId="f3301882-4e58-4b32-fdc1-1a59bfcde48e"
Test_dataset = pd.read_csv ('TestData_N111_p10.csv')
Test_dataset.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="C3aqSdNLue4t" outputId="0fdb0452-1db6-4a23-9ffa-dd3d9956dcfc"
# What are the datatypes of each observation:
print(Test_dataset.dtypes)
# Shape of my data
print('The size of our data are: ',Test_dataset.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="L207lbaiue4t" outputId="345b2589-b1c5-4b10-ae21-3dd0000b6fbc"
# Are there any null or missing values
print('Test Dataset Missing Values: \n',Test_dataset.isnull().sum())
# + [markdown] id="QuHKDRLUue4u"
# ### Lets change the categorical values <a class="anchor" id="Lets_change_the_categorical_values"></a>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="sd7nJVOHue4u" outputId="c4b397ba-3946-434b-f73e-ab9adfe17032"
# recode the categories
Training_Class = Train_dataset['Ancestry'].unique().tolist()
Test_Class = Test_dataset['Ancestry'].unique().tolist()
num_features = len(Training_Class)
print("Unique Values for Train Ancestry: ", Training_Class)
print("Unique Values for Test Ancestry: ", Test_Class)
# + id="CIIc0esJue4u"
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
Train_dataset['Ancestry_Encoded'] = le.fit_transform(Train_dataset.iloc[:,-1:])
Test_dataset['Ancestry_Encoded'] = le.fit_transform(Test_dataset.iloc[:,-1:])
# + colab={"base_uri": "https://localhost:8080/", "height": 144} id="_CjuIej-ue4u" outputId="8018312d-1283-4d9a-866a-8dfeeec2f00c"
Train_dataset.head(3)
# + [markdown] id="4VSof3lAue4u"
# ### Create Predictor and Target numpy array <a class="anchor" id="Create_Predictor_and_Target_numpy_array"></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="z9hAQmYcue4v" outputId="84f06175-971e-4984-f52f-77675aaef924"
# Target:
Y_Train= Train_dataset['Ancestry_Encoded'].to_numpy()
Y_Test= Test_dataset['Ancestry_Encoded'].to_numpy()
Y_Train.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="iQ9uafZ4ue4v" outputId="54fc05a5-ed04-4c2c-8355-dc829ad44e93"
# Convert the Pandas dataframe to numpy ndarray for computational improvement
X_Train = Train_dataset.iloc[:,:-2].to_numpy()
X_Test = Test_dataset.iloc[:,:-2].to_numpy()
print(type(X_Train), X_Train[:1], "Shape = ", X_Train.shape)
# + [markdown] id="hFDnLTEDue4v"
# ### Create a Normalize copy of variables <a class="anchor" id="Create_a_Normalize_copy_of_variables"></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="49-K0j34ue4v" outputId="29cf1183-754f-4bee-b1fc-ae99567e5c0b"
# Create Standarizing ObjectPackages:
standardization = StandardScaler()
# Strandardize
n_observations = len(Train_dataset)
variables = Train_dataset.columns
# Standardize the Predictors (X)
X_Train = standardization.fit_transform(X_Train)
# Add a constanct to the predictor matrix
#X_Train = np.column_stack((np.ones(n_observations),X_Train))
# Save the original M and Std of the original data. Used for unstandardize
original_means = standardization.mean_
# we chanced standardization.std_ to standardization.var_**.5
originanal_stds = standardization.var_**.5
print("observations :", n_observations)
print("variables :", variables[:2])
print('original_means :', original_means)
print('originanal_stds :', originanal_stds)
# + [markdown] id="bm9rE-r9ue4w"
# ### Split Data: <a class="anchor" id="Split_Data:"></a>
# + [markdown] id="1tDsmt9Cue4w"
# #let's first split it into train and test part
# X_train, X_out_sample, y_train, y_out_sample = train_test_split(Xst, y_Centered, test_size=0.40, random_state=101) # Training and testing split
#
# X_validation, X_test, y_validation, y_test = train_test_split(X_out_sample, y_out_sample, test_size=0.50, random_state=101) # Validation and test split
#
# # Print Data size
# print ("Train dataset sample size: {}".format(len(X_train)))
# print ("Validation dataset sample size: {}".format(len(X_validation)))
# print ("Test dataset sample size: {}".format( len(X_test)))
# + [markdown] id="A90wcawSue4w"
# # Regression Model <a class="anchor" id="Regression_Model"></a>
# <hr>
# + [markdown] id="X0VLOqIgue4w"
# ### Define our learning rates <a class="anchor" id="Define_our_learning_rates"></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="NfjpUAslue4w" outputId="6c6096c7-eea7-4526-9639-ce259b212343"
# Define my tuning parameter values 𝜆:
learning_rates_λ = [1.e-04, 1.e-03, 1.e-02, 1.e-01, 1.e+00, 1.e+01, 1.e+02, 1.e+03, 1.e+04]
print(learning_rates_λ)
# learning rate
α = 1e-4
# K-folds
k = 5
# Itterations
n_iters = 10000
# + [markdown] id="7RJC_4Q_ue4x"
# ### Create the Regression Objects <a class="anchor" id="Create_the_Regression_Objects"></a>
# + [markdown] id="arh5UEjbue4x"
# **LogisticRegression Library** <a class="anchor" id="LogisticRegression_Library"></a>
# + id="OkjHDb6Eue4x"
# LogisticRegression
from sklearn.linear_model import LogisticRegression
Library_LogisticRegression = LogisticRegression(max_iter = 10000, multi_class='multinomial', solver='lbfgs', penalty='l2', C=1)
# + [markdown] id="oxabTSELue4x"
# ## **Deliverable 7.1** <a class="anchor" id="Deliverable_6.1"></a>
# <h>
#
# > Deliverable 1: Illustrate the effect of the tuning parameter on the inferred ridge regression coefficients by generating five plots (one for each of the 𝐾=5 ancestry classes) of 10 lines (one for each of the 𝑝=10 features), with the 𝑦-axis as 𝛽̂
# 𝑗𝑘, 𝑗=1,2,…,10 for the graph of class 𝑘, and 𝑥-axis the corresponding log-scaled tuning parameter value log10(𝜆) that
# 7
# generated the particular 𝛽̂
# 𝑗𝑘. Label both axes in all five plots. Without the log scaling of the tuning parameter, the plot will look distorted.
# + [markdown] id="_dpSgiabue4y"
# **LogisticRegression with Library**
# + id="ZS8XcUaEue4y"
L𝛽_per_λ=[] # set empty list
# Evaluate tuning parameters with LogisticRegression penalty
for tuning_param in learning_rates_λ:
Library_LogisticRegression = LogisticRegression(max_iter = 10000, multi_class = 'multinomial', solver = 'lbfgs', penalty ='l2', C = tuning_param)
Library_LogisticRegression.fit(X_Train, Y_Train)
c = np.array(Library_LogisticRegression.coef_)
# c = np.append(tuning_param,c)
L𝛽_per_λ.append(Library_LogisticRegression.coef_)
# print(c)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="_IcmDKwpue4y" outputId="1e9bd9e9-a0be-4cc7-955c-aede101f87d7"
L𝛽_per_λ[0]
# + id="nf2e0HTDue4y"
# Loop throught the betas, by class generated by each lamda
temp_df = []
for l in range(np.array(L𝛽_per_λ).shape[0]):
for c in range(np.array(L𝛽_per_λ).shape[1]):
temp_df.append(np.append(L𝛽_per_λ[l][c],(learning_rates_λ[l],c)))
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="XoeUHdWkue4y" outputId="ef0d0f74-2de9-4d6e-fe72-e3aae14c7409"
TunnedL𝛽_df=pd.DataFrame(np.array(temp_df))
TunnedL𝛽_df.columns=['PC1', 'PC2', 'PC3', 'PC4', 'PC5', 'PC6', 'PC7', 'PC8', 'PC9', 'PC10', 'Lambda', 'Class']
#TunnedL𝛽_df['Class_Name'] = TunnedL𝛽_df['Class_Name'].apply(lambda x: Training_Class[int(x)])
TunnedL𝛽_df.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="FKFHJ0Auue4z" outputId="b5d1436b-0b08-47e2-8c72-b926ec77ffc2"
Training_Class
# + colab={"base_uri": "https://localhost:8080/", "height": 331} id="JUBoYcoBue4z" outputId="3a2ea178-e9f0-498c-c98b-709da2a8a20e"
TunnedL𝛽_df[TunnedL𝛽_df.Class.eq(0)]
# + colab={"base_uri": "https://localhost:8080/", "height": 1876} id="xmvJFP2bue4z" outputId="a8845168-a486-4fe6-ab5e-5dfc614ee01a"
# Plot tuning parameter on the inferred ridge regression coefficients
sns.set(rc = {'figure.figsize':(15,8)})
for i, c in enumerate(Training_Class):
sns.set_theme(style="whitegrid")
sns.set_palette("mako")
for j in range(1, 1 + X_Train.shape[1]):
sns.lineplot( x = TunnedL𝛽_df[TunnedL𝛽_df.Class.eq(i)]['Lambda'], y = TunnedL𝛽_df[TunnedL𝛽_df.Class.eq(i)]['PC{}'.format(j)], palette='mako', label = 'PC{}'.format(j) )
sns.set()
plt.xscale('log')
plt.legend(bbox_to_anchor=(1.09, 1), loc='upper left')
plt.xlabel('Log Lambda')
plt.ylabel('Coefficient Values')
plt.suptitle('Inferred Ridge Regression Coefficient Tuning Parameters of' + ' ' + c + ' ' + 'Class')
plt.show()
# + [markdown] id="X7f_U3pDue4z"
# # **Deliverable 7.2** <a class="anchor" id="Deliverable_6.2"></a>
# Illustrate the effect of the tuning parameter on the cross validation error by generating a plot with the 𝑦-axis as CV(5) error, and the 𝑥-axis the corresponding log-scaled tuning parameter value log10(𝜆) that generated the particular CV(5) error. Label both axes in the plot. Without the log scaling of the tuning parameter 𝜆, the plots will look distorted.
# + [markdown] id="y3lr1rFYue4z"
# **CV Elastic Net with Library**
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="McWV5Efrue40" outputId="b4ca00b6-c95b-43fd-aa8e-cd0e53d28ce7"
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
#Define the model
Library_LogisticRegression = LogisticRegression(max_iter = 10000, multi_class = 'multinomial', solver = 'lbfgs', penalty ='l2')
# Create the Kfold:
cv_iterator = KFold(n_splits = 5, shuffle=True, random_state=101)
cv_score = cross_val_score(Library_LogisticRegression, X_Train, Y_Train, cv=cv_iterator, scoring='neg_mean_squared_error', n_jobs=1)
print (cv_score)
print ('Cv score: mean %0.3f std %0.3f' % (np.mean(cv_score), np.std(cv_score)))
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="wsk37cY6ue40" outputId="5d83e531-2312-4bc1-dcb1-442352003017"
# define grid
Parm_grid = dict()
Parm_grid['C'] = learning_rates_λ
Parm_grid
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="rNGvOeWhue40" outputId="3baf5043-3083-4f56-eb7a-5fa78e51f2eb"
# Lets define search
GsearchCV = GridSearchCV(estimator = Library_LogisticRegression, param_grid = Parm_grid, scoring = 'neg_mean_squared_error', n_jobs=1, refit=True, cv=cv_iterator)
GsearchCV.fit(X_Train, Y_Train)
# + colab={"base_uri": "https://localhost:8080/", "height": 331} id="psatnUqAue40" outputId="fe126e22-3540-4038-ad5e-789b607dce07"
GCV_df = pd.concat([pd.DataFrame(GsearchCV.cv_results_["params"]),pd.DataFrame(GsearchCV.cv_results_["mean_test_score"], columns=["mean_test_score"])],axis=1)
#GCV_df.index=GCV_df['alpha']
GCV_df.rename(columns={"C": "learning_rates_λ"}, inplace=True)
GCV_df
# + colab={"base_uri": "https://localhost:8080/", "height": 566} id="ZTiKO6UAue40" outputId="8e52025d-71a8-411d-fa60-60036fc95b52"
sns.set_theme(style="whitegrid")
sns.set_palette("mako")
plt.plot(GCV_df["learning_rates_λ"] , GCV_df["mean_test_score"])
sns.set_palette("mako")
sns.set()
plt.suptitle('Effect of the uning parameter on the cross validation error log10(lambda)')
plt.xscale('log')
plt.xlabel('Lambda')
plt.ylabel('loss from cross valiadation')
plt.show()
# + [markdown] id="_LHbyoNRue41"
# # **Deliverable 7.3** <a class="anchor" id="Deliverable_6.3"></a>
# Indicate the value of 𝜆 that generated the smallest CV(5) error
# + [markdown] id="_XKTKUG5ue41"
# **Smallest CV with Library**
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="oHGAC_95ue41" outputId="f3a60450-467b-4169-ef9f-c7f7fd21990d"
print ('Best: ',GsearchCV.best_params_)
print ('Best CV mean squared error: %0.3f' % np.abs(GsearchCV.best_score_))
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="MfHtU6EVue41" outputId="5db86ff7-17ff-433d-f3a7-6fe656969975"
GCV_df.sort_values(by=['mean_test_score'], ascending=False)[:1]
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="-hSF7cC5ue41" outputId="1b65d27a-0251-4bea-db48-48861e77c05e"
# Alternative: sklearn.linear_model.ElasticNetCV
from sklearn.linear_model import LogisticRegressionCV
auto_LR = LogisticRegressionCV(Cs = learning_rates_λ, cv=5, max_iter = 10000, multi_class = 'multinomial', solver = 'lbfgs', penalty ='l2', n_jobs= 1 )
auto_LR.fit(X_Train, Y_Train)
#print ('Best alpha: %0.5f' % auto_LR.alpha_)
print ('Best 𝜆: ' , auto_LR.C_)
# + [markdown] id="eM4N_2jCue42"
# # **Deliverable 7.4** <a class="anchor" id="Deliverable_6.4"></a>
#
# Given the optimal 𝜆, retrain your model on the entire dataset of 𝑁=183 observations to obtain an estimate of the (𝑝+1)×𝐾 model parameter matrix as 𝐁̂ and make predictions of the probability for each of the 𝐾=5 classes for the 111 test individuals located in TestData_N111_p10.csv. That is, for class 𝑘, compute
# 𝑝𝑘(𝑋;𝐁̂)=exp(𝛽̂0𝑘+Σ𝑋𝑗𝛽̂ 𝑗𝑘𝑝𝑗=1)
# / Σexp(𝛽̂0ℓ+Σ𝑋𝑗𝛽̂ 𝑗ℓ𝑝𝑗=1)
#
# - for each of the 111 test samples 𝑋, and also predict the most probable ancestry label as
# - 𝑌̂(𝑋)=arg max𝑘∈{1,2,…,𝐾}𝑝𝑘(𝑋;𝐁̂)
# - Report all six values (probability for each of the 𝐾=5 classes and the most probable ancestry label) for all 111 test individuals.
# + [markdown] id="jo55OlKnue42"
# **Tunned with best λ with Library**
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="PErXslROue42" outputId="406d9af4-09b1-4bfb-bfe6-98db22976086"
Library_LogisticRegression_best= LogisticRegression(max_iter = 10000, multi_class='multinomial', solver='lbfgs', penalty='l2', C= auto_LR.C_[0])
Library_LogisticRegression_best.fit( X_Train, Y_Train )
y_predM_best = Library_LogisticRegression_best.predict(X_Test)
print ("Betas= ", np.mean(Library_LogisticRegression_best.coef_, 0))
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="ZkOnUiR0ue42" outputId="5b75546f-9953-4bec-c779-52212d5e23be"
yhat = Library_LogisticRegression_best.predict_proba(X_Test)
# summarize the predicted probabilities
print('Predicted Probabilities: %s' % yhat[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="F9H88sTBue42" outputId="35cf8500-163e-418e-97a9-d3184c4a89c0"
ŷ_test = Library_LogisticRegression_best.predict_proba(X_Test)
ŷ_test[:3]
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="9Ep_IF3Uue42" outputId="72f74182-fb9e-4839-a1a9-f097c3ed0d07"
Y_class = Library_LogisticRegression_best.predict(X_Test)
Y_class
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="FoZCP092ue43" outputId="7bb11d8d-f021-40ca-f77c-3abf077f5e9f"
# Re-lable feature headers and add new class prediction index column
new_colNames = ['{}_Probability'.format(c_name) for c_name in Training_Class] + ['ClassPredInd']
new_colNames
# + id="x_bDg2MXue43"
# Implemnt index array of probabilities
i_prob = np.concatenate((ŷ_test, Y_class[:, None]), 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="QA2fz66Que43" outputId="58c0b070-3be4-47b3-e621-4980fa9eec34"
# Create New dataframe for probality indeces
df2 = pd.DataFrame(i_prob, columns = new_colNames)
df2
# + id="hRAT2uc0ue43"
# Concat dependant Ancestory features to dataframe
dep_preds = pd.concat([Test_dataset['Ancestry'], df2], axis = 1)
# + id="ItsgJBuRue43"
# Add new
dep_preds['ClassPredName'] = dep_preds['ClassPredInd'].apply(lambda x: Training_Class[int(x)])
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="anKsZHWxue43" outputId="14439f99-ab9f-4734-8e6e-29e6a9b5db4c"
# Validate Probability predictions dataframe
dep_preds.head()
# + id="32XEV_57ue44"
# Slice prediction and set new feature vector column variable
prob_1 = dep_preds.loc[:, 'Ancestry':'NativeAmerican_Probability']
# + id="XLG5JABPue44"
# Unpivot convert dataFrame to long format
prob_2 = pd.melt(prob_1, id_vars = ['Ancestry'], var_name = 'Ancestry_Predictions', value_name = 'Probability')
# + id="XpWUBAD4ue44"
# Test for true probability
prob_2['Ancestry_Predictions'] = prob_2['Ancestry_Predictions'].apply(lambda x: x.split('Prob')[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="SKGdbaDFue44" outputId="cb1da972-49c8-4c7b-bb46-2298b49e26e1"
# Validate dataframe
prob_2.head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="5vnc8FsRue44" outputId="1cc68e12-d923-4f87-c9aa-9a6d7ea6c6f1"
# Validate dataframe features
print('Describe Columns:=', prob_2.columns, '\n')
print('Data Index values:=', prob_2.index, '\n')
print('Describe data:=', prob_2.describe(), '\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 560} id="efHFVS3zue44" outputId="7a91d840-8740-45b0-dcdc-a153e5c02a52"
# Plot Probality prediction matrix
sns.set(rc = {'figure.figsize':(15,8)})
sns.set_theme(style="whitegrid")
fig, ax = plt.subplots()
sns.barplot(data = prob_2[prob_2['Ancestry'] != 'Unknown'],color = 'r', x = 'Ancestry', y = 'Probability', hue = 'Ancestry_Predictions', palette = 'mako')
plt.xlabel('Ancestory Classes')
plt.ylabel('Probability')
plt.suptitle('Probabilty of Ancestor classes')
#plt.savefig("Assignment3_Deliverable4.png")
plt.show()
# + [markdown] id="3cHf0cSAue44"
# # **Deliverable 7.5** <a class="anchor" id="Deliverable_6.5"></a>
# How do the class label probabilities differ for the Mexican and African American samples when compared to the class label probabilities for the unknown samples? Are these class probabilities telling us something about recent history? Explain why these class probabilities are reasonable with respect to knowledge of recent history?
# + [markdown] id="mnZWnz6Be_1A"
# - In comparison to the class label probabilities for the unknown samples, those with unknown ancestry show a probability close to or equal to one while the other classes show a probability close to zero or less than one. African American samples showed similar results. The model assigned high probabilities to the African ancestry class for each of these samples. However, both Native American and European ancestry contribute high probabilities to the Mexican population on average with Native American slightly higher than European.
| Assignment 3/From Libaries Implementation/_CAP5625_Assigment_3_libaries_LogisticRidgeRegresion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
from classMVA import MVA
import numpy as np
import matplotlib.pyplot as plt
# + deletable=true editable=true
X = np.random.randn(5000,7) * np.array([1, 1, np.sqrt(0.5), np.sqrt(0.1), np.sqrt(0.01), 1e-4, 1e-4])
#Regression coefficients
R = np.array([[0, 0.3, 0],[0, 0, 0],[0, 0, 0], [0.4, 0.3, 0], [0.5, 0.3, 100], [0, 0, 0], [0, 0, 0]])
print R
Y = X.dot(R) + np.sqrt(1e-3)*np.random.randn(5000,3)
plt.imshow(X.T.dot(X), interpolation='none')
plt.show()
# + deletable=true editable=true
#Finally, we mix a little bit some coordinates:
W = np.eye(7,7)
W[0,3] = .3
W[2,5] = .8
print W
X2 = X.dot(W)
plt.imshow(X2.T.dot(X2), interpolation='none')
plt.show()
# + deletable=true editable=true
RDD_X = sc.parallelize(X2.tolist())
RDD_Y = sc.parallelize(Y.tolist())
RDD_labeled = RDD_Y.zip(RDD_X)
print RDD_X.count()
print RDD_Y.count()
print RDD_labeled.count()
print RDD_X.first()
print RDD_Y.first()
print RDD_labeled.first()
# + deletable=true editable=true
#PCA example
prueba = MVA('PCA','l1','None',1e-5,7,0.0000001,1e-3,100)
RDD_PCA = RDD_labeled.map(lambda x: x[1])
prueba.prepareData(RDD_PCA)
prueba.normalizer()
# + deletable=true editable=true
print prueba._typeMVA
print prueba._typeReg
print prueba._typeNorm
print prueba._tol
print prueba._numVariables
print prueba._M
print prueba._data.take(2)
print prueba._normdata.take(2)
print prueba._scaler
print prueba._U
print prueba._max_Ustep
# + deletable=true editable=true
prueba = MVA('PCA','l1','None',1e-5,7,0.001,1e-3,100)
prueba.fit(RDD_PCA)
print prueba._U
# + deletable=true editable=true
#RDD_NEW= RDD_PCA.map(lambda x: x.dot(((self._U).T))).collect()
RDD_NEW=prueba.predict(RDD_PCA)
print RDD_NEW.count()
print RDD_NEW.take(2)
print RDD_NEW.first()
# + deletable=true editable=true
RDD2 = prueba._scaler.transform(RDD_PCA)
U = prueba._U
sc.broadcast(U)
RDD2 = RDD2.map(lambda x: x.dot(((U).T)))
print RDD2.first()
#print RDD2.count()
#print RDD2.take(2)
# + deletable=true editable=true
def sum_matrix(matriz,n,m):
for i in range(n):
suma=0.00000000e+00
for j in range(m):
suma=suma+np.abs(matriz[i][j])
print np.abs(matriz[i][j])
return suma
# + deletable=true editable=true
#BLANQUEADO DE DATOS
X_2=(X2).dot(U.T)
Cov=X_2.T.dot(X_2)
MM=np.diagonal(Cov).reshape(1,6)
from scipy import sparse
diag=sparse.spdiags(MM,0,6,6).toarray()
#print diag
fin=Cov-diag
print fin
#print fin[0][1]
suma=sum_matrix(fin,5,5)
print 'suma= ' + str(suma)
plt.imshow(np.abs(fin), interpolation='none')
plt.show()
# + deletable=true editable=true
matrix = RDD2.map(lambda x : np.dot(x[:,np.newaxis],x[:,np.newaxis].T)).mean()
print matrix
#X_2=(X2).dot(U.T)
plt.imshow(matrix, interpolation='none')
plt.show()
# + deletable=true editable=true
from operator import add
def count_0s (array):
cuenta=0
for i in range(len(array)):
if array[i] > 1e-4 :
cuenta=cuenta+1
return cuenta
#array= [ 1.02782977e-03 , 1.54021761e-04 , -1.67806562e-03 , -4.35857479e-05, -2.32505213e-06, 0.00000000e+00]
#cuenta=count_0s(array)
#print cuenta
number_0s=RDD2.map(lambda x: ('zero',count_0s(x)))
number=number_0s.reduceByKey(add)
print number_0s.take(8)
print number.collect()
# + deletable=true editable=true
#self, typeMVA, typeReg,typeNorm, tol, regParam=0.01, step=1e-3, iterations=100, max_Ustep=10):
prueba = MVA('PCA','l1','None',1e-5,7,0.00000001,1e-3,100)
prueba.fit(RDD_PCA)
print prueba._U
# + deletable=true editable=true
from sklearn import preprocessing
plt.imshow(np.abs(preprocessing.normalize(prueba._U, norm='l2')), interpolation='none')
plt.show()
# + deletable=true editable=true
from sklearn import decomposition
pca = decomposition.PCA(n_components=6)
pca.fit(RDD_PCA.collect())
plt.imshow(np.abs(pca.components_), interpolation='none')
plt.show()
# + deletable=true editable=true
prueba1 = MVA('PCA','l1','None',1e-5,7,0.0001,1e-2,100)
prueba1.fit(RDD_PCA)
# + deletable=true editable=true
plt.imshow(np.abs(preprocessing.normalize(prueba._U, norm='l2')), interpolation='none')
plt.show()
# + deletable=true editable=true
prueba2 = MVA('PCA','l1','None',1e-5,7,0.01,1e-2,100)
prueba2.fit(RDD_PCA)
# + deletable=true editable=true
prueba3 = MVA('PCA','l1','None',1e-5,7,0.1,1e-2,100)
prueba3.fit(RDD_PCA)
# + deletable=true editable=true
import matplotlib.pyplot as plt
plt.figure()
plt.plot(abs(prueba1._U.T[:,5]),'b',label='0.0001')
plt.hold(True)
plt.plot(abs(prueba2._U.T[:,5]),'r',label='0.01')
plt.hold(True)
plt.plot(abs(prueba3._U.T[:,5]),'g',label='0.1')
plt.legend(loc = 1)
plt.xlabel('k', fontsize=14)
plt.ylabel('U(k)', fontsize=14)
plt.show()
# + deletable=true editable=true
from sklearn import decomposition
from sklearn import preprocessing
plt.imshow(np.abs(preprocessing.normalize(prueba._U, norm='l2')), interpolation='none')
plt.show()
# + deletable=true editable=true
# + deletable=true editable=true
plt.plot(np.abs(preprocessing.normalize(prueba._U, norm='l2')).T)
plt.show()
# + deletable=true editable=true
plt.plot(abs(pca.components_.T))
plt.show()
# + deletable=true editable=true
prueba = MVA('PCA','l1','None',1e-5,7,0.1,1e-3,100)
prueba.fit(RDD_PCA)
# + deletable=true editable=true
from sklearn import preprocessing
print prueba._U
plt.imshow(np.abs(preprocessing.normalize(prueba._U, norm='l2')), interpolation='none')
plt.show()
# + deletable=true editable=true
#self, typeMVA, typeReg,typeNorm, tol, regParam=0.01, step=1e-3, iterations=100, max_Ustep=10):
prueba = MVA('OPLS','l1','norm',1e-5,7)
prueba.fit(RDD_labeled)
# + deletable=true editable=true
plt.imshow(np.abs(prueba._U), interpolation='none')
plt.show()
# + deletable=true editable=true
plt.plot(abs(prueba._U.T[:,0]))
plt.show()
# + deletable=true editable=true
import matplotlib.pyplot as plt
import numpy as np
x=[0.00001, 0.001,0.1,0.5]
y=[3.080e-7,3.137e-7,4.438e-5,5.858e-5]
plt.semilogx()
plt.semilogy()
#plt.legend(loc = 2)
plt.xlabel('Reg', fontsize=14)
plt.ylabel('Sum', fontsize=14)
plt.plot(x,y,'g')
plt.show()
# -
| synthetic_data2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (tensorflow)
# language: python
# name: tensorflow
# ---
arr = [1,2,3,4,5,6,7]
min(arr)
def index_prod(lst):
# Create an empty output list
output = [None] * len(lst)
# Set initial product and index for greedy run forward
product = 1
i = 0
while i < len(lst):
# Set index as cumulative product
output[i] = product
# Cumulative product
product *= lst[i]
# Move forward
i +=1
# Now for our Greedy run Backwards
product = 1
# Start index at last (taking into account index 0)
i = len(lst) - 1
# Until the beginning of the list
while i >=0:
# Same operations as before, just backwards
output[i] *= product
product *= lst[i]
i -= 1
return output
index_prod([1,2,3,4])
def overlap(r1, r2):
r1_x, r1_y = r1['x'], r1['y']
r2_x, r2_y = r2['x'], r2['y']
r1_x_range, r1_y_range = r1['h'], r1['w']
r2_x_range, r2_y_range = r2['h'], r2['w']
if r2_x in range(r1_x + r1_x_range) and \
r2_y in range(r1_y + r1_y_range):
return True
else:
return False
r1 = {'x': 2 , 'y': 4,'w':5,'h':12}
r2 = {'x': 1 , 'y': 5,'w':7,'h':14}
overlap(r1,r2)
# +
from random import randint
def dice():
return randint(1, 5)
def roll_7():
while True:
roll_1 = dice()
roll_2 = dice()
roll = ((roll_1 - 1)*5) + (roll_2 )
if roll > 21:
continue
return roll %7 + 1
# -
s = 'Shivam'
print(s)
len(s)
def name(a):
if len(a) == 0 or len(a) == 1:
return a
else:
return name(a[1:]) + a[0]
name("hello")
0 == 0.00
for i in range(20,30):
print (i ** (1/2))
| interviewQuestions/E-CommerceCompany/min_max.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (tunnel)
# language: python
# name: python3
# ---
# + [markdown] id="vhe1yX4AMckE"
# # DSE Course 2, Session 1: Model Selection and Validation
#
# **Instructor**: <NAME>
#
# **Contact**: <EMAIL>
#
# <br>
#
# ---
#
# <br>
#
# At the end of the last course, we saw the basic recipe for creating a supervised machine learning model:
#
# 1. Environment setup and importing data
# 2. Rudimentary exploratory data analysis
# 3. Feature engineering
# 4. Choosing and training a model:
# 1. choose model
# 2. choose hyperparameters
# 3. fit using training data
# 4. predict using validation data
#
# In C1 S7, I chose our model and hyperparameters preemptively. How did I do that? In the real world, you won't necessarily have the best intution about how to make these choices. In today's session, we will algorithmize the way we approach choosing and training a model
#
# Note: I will import libraries at the beginning of this notebook, as is good practice, but will reimport them as they are used to remind ourselves where each method came from!
#
# <br>
#
# <p align=center>
# <img src="https://raw.githubusercontent.com/wesleybeckner/ds_for_engineers/main/assets/C2/schedule.png"></img>
# </p>
#
# ---
#
# <br>
#
# <a name='top'></a>
#
# # Contents
#
# * 1.0 [Preparing Environment and Importing Data](#x.0)
# * 1.0.1 [Import Packages](#x.0.1)
# * 1.0.2 [Load Dataset](#x.0.2)
# * 1.1 [Model Validation](#1.1)
# * 1.1.1 [Holdout Sets](#1.1.1)
# * 1.1.2 [Data Leakage and Cross-Validation](#1.1.2)
# * 1.1.3 [Bias-Variance Tradeoff](#1.1.3)
# * 1.1.4 [Learning Curves](#1.1.4)
# * 1.1.4.1 [Considering Model Complexity](#x.1.4.1)
# * 1.1.4.1 [Considering Training Set Size](#x.1.4.2)
# * 1.2 [Model Validation in Practice](#1.2)
# * 1.2.1 [Grid Search](#1.2.1)
#
# <br>
#
# [References](#reference)
#
# ---
# + [markdown] id="mNtJitcRW51Y"
# <a name='x.0'></a>
#
# ## 1.0 Preparing Environment and Importing Data
#
# [back to top](#top)
# + [markdown] id="5PcjXaRjJCOi"
# <a name='x.0.1'></a>
#
# ### 1.0.1 Import Packages
#
# [back to top](#top)
# + id="q80G6fFCJkjt" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1620423863818, "user_tz": 300, "elapsed": 1696, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="ce044e20-733b-468f-95f7-d638034ab9cd"
# Pandas library for the pandas dataframes
import pandas as pd
import numpy as np
import datetime
import matplotlib.pyplot as plt
import seaborn as sns
import random
import scipy.stats as stats
from patsy import dmatrices
from statsmodels.stats.outliers_influence import variance_inflation_factor
from sklearn.neighbors import KNeighborsRegressor
from sklearn.preprocessing import LabelEncoder, StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression, LinearRegression
from sklearn import metrics
from sklearn.metrics import r2_score
# + [markdown] id="UrVE2RBuJEwg"
# <a name='x.0.2'></a>
#
# ### 1.0.2 Load Dataset
#
# [back to top](#top)
#
# In course 1 we cursorily discussed why we may need strategies for validating our model. Here we'll discuss it more in depth.
#
# I'm going to take a simple example. In the following, I have a dataset that contains some data about a piece of equipment
# + id="fOeegZuwJnEh" executionInfo={"status": "ok", "timestamp": 1620423864030, "user_tz": 300, "elapsed": 1906, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}}
df = pd.read_csv("https://raw.githubusercontent.com/wesleybeckner/"\
"ds_for_engineers/main/data/misc/knife_in_dataset.csv")
# + [markdown] id="nbzsVbOST9vJ"
# You can see that this is a multindexed dataset, so we will have to do some data preprocessing acrobatics to get our training and test sets to look the way we want
# + colab={"base_uri": "https://localhost:8080/", "height": 253} id="KF1Gy8STPvkP" executionInfo={"status": "ok", "timestamp": 1620423864031, "user_tz": 300, "elapsed": 1808, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="1cb400e3-cb52-4577-9d6d-3017563805f0"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 628} id="NvDxXgcqPtf5" executionInfo={"status": "ok", "timestamp": 1620057475878, "user_tz": 300, "elapsed": 2453, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="d8f51e92-bab4-4c31-f4fe-08d26f94e719"
fig, ax = plt.subplots(1, 1, figsize=(10,10))
sns.heatmap(df.iloc[:,3:], ax=ax)
# + colab={"base_uri": "https://localhost:8080/"} id="5jQahdaYUG5_" executionInfo={"status": "ok", "timestamp": 1620057477158, "user_tz": 300, "elapsed": 3721, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="cdabdabb-248c-40bf-d361-46a8f2cca6af"
# Here I'm parsing this dataframe so that each one of my training values
# contains the correct information
ls = []
for row in df.index:
for col in df.columns[3:]:
stuff = list(np.append(df.iloc[row,:3].values, col))
stuff.append(df.iloc[row][col])
ls.append(stuff)
data = np.array(ls)
print(data.shape)
X = data[:,:-1]
y = data[:,-1]
# + [markdown] id="2mRJz9__1smm"
# We check that the dimensions of our x and y data make sense:
# + colab={"base_uri": "https://localhost:8080/"} id="ATwy5miJ1mQ5" executionInfo={"status": "ok", "timestamp": 1620057477161, "user_tz": 300, "elapsed": 3715, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="d48d965c-c8eb-4944-b525-cf2a496dfd43"
print("X shape: {}".format(X.shape))
print("y shape: {}".format(y.shape))
# + [markdown] id="7B3YvBprQ3il"
# let's go ahead and load our wine dataset as well...
# + id="hR3zCNbwO-D_"
wine = pd.read_csv("https://raw.githubusercontent.com/wesleybeckner/"\
"ds_for_engineers/main/data/wine_quality/winequalityN.csv")
# + id="GivwSc_7JoUU"
wine.dropna(inplace=True)
wine['quality_label'] = wine['quality'].apply(lambda x: 'low' if x <=5 else
'med' if x <= 7 else 'high')
class_tp = {'red': 0, 'white': 1}
y_tp = wine['type'].map(class_tp)
wine['type_encoding'] = y_tp
class_ql = {'low':0, 'med': 1, 'high': 2}
y_ql = wine['quality_label'].map(class_ql)
wine['quality_encoding'] = y_ql
wine.columns = wine.columns.str.replace(' ', '_')
# + id="267JW5JP6P3z"
# df = pd.read_csv("https://raw.githubusercontent.com/wesleybeckner/"\
# "ds_for_engineers/main/data/wine_quality/winequalityN.csv")
# df['quality_label'] = df['quality'].apply(lambda x: 'low' if x <=5 else
# 'med' if x <= 7 else 'high')
# df.columns = df.columns.str.replace(' ', '_')
# df.dropna(inplace=True)
# class_tp = {'red': 0, 'white': 1}
# y_tp = df['type'].map(class_tp)
# df['type_encoding'] = y_tp
# class_ql = {'low':0, 'med': 1, 'high': 2}
# y_ql = df['quality_label'].map(class_ql)
# df['quality_encoding'] = y_ql
# features = list(df.columns[1:-1].values)
# features.remove('type_encoding')
# features.remove('quality_label')
# features.remove('quality')
# features
# + [markdown] id="062Czp9-rBOi"
# <a name='1.1'></a>
#
# ## 1.1 Model Validation
#
# [back to top](#top)
#
# *doing it the wrong way*<br>
#
# While we're here, I'm going to introduce a VERY SIMPLE supervised learning method called K-Nearest Neighbors.
# + [markdown] id="fa_N3uQFn-mB"
# <a name='x.1.0'></a>
#
# ## 1.1.0 K-Nearest Neighbors
#
# [back to top](#top)
#
# K-Nearest Neighbors is perhaps the simplest algorithm of them all. It is essentially a lookup table: We select the hyperparameter K, and when assigning a new value a data label, assign it according to, the majority label in the vicinity of the new datapoint. The vicinity being determined by K, the number of nearest neighbors we are going to assess.
# + id="hQVbpeY9UtVx"
X = X.astype(int)
y = y.astype(int)
# + colab={"base_uri": "https://localhost:8080/"} id="nwI8pcOUTe2G" executionInfo={"status": "ok", "timestamp": 1620058563748, "user_tz": 300, "elapsed": 171, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="95662a32-80b2-46f1-ceb7-f444a459e580"
knn = KNeighborsRegressor(n_neighbors=1)
knn.fit(X,y)
# + colab={"base_uri": "https://localhost:8080/"} id="wUfxGTuRSC1u" executionInfo={"status": "ok", "timestamp": 1620058564136, "user_tz": 300, "elapsed": 158, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="8440b312-c9ec-424f-a561-adb0d6059e8b"
knn.score(X,y)
# + [markdown] id="7MZcVZ3OU16F"
# Wow! we achieved a model with a perfect score! But is this really how we would expect the model to perform on data it had never seen before? Probably not. How do we actually check for this?
#
# While we're at it, let's do the same for our wine dataset. Last week we left out a portion of the data for testing with the following cell
#
# ```
# cols = wine.columns
# cols = list(cols.drop(['type', 'type_encoding', 'quality', 'quality_encoding',
# 'quality_label']))
# X_train, X_test, y_train, y_test = train_test_split(wine.loc[:, cols], y_tp,
# test_size=0.4, random_state=42)
# ```
#
#
# This time we'll train our model on the entire dataset
# + id="uJsw6wqhMofr"
cols = wine.columns
cols = list(cols.drop(['type', 'type_encoding', 'quality', 'quality_encoding',
'quality_label']))
X_wine, y_wine = wine.loc[:, cols], wine['quality']
# + id="wHLJcFrNVfN8"
model = LogisticRegression(penalty='l2',
tol=.001,
C=.003,
class_weight='balanced',
solver='liblinear',
max_iter=1e6)
# + colab={"base_uri": "https://localhost:8080/"} id="taPFa7m4VgfS" executionInfo={"status": "ok", "timestamp": 1620057485021, "user_tz": 300, "elapsed": 403, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="8c50df5c-5a8f-418d-ed3e-e9ffb8d07834"
model.fit(X_wine, y_wine)
# + id="GT9p9qUtVn2E"
y_pred = model.predict(X_wine)
# + colab={"base_uri": "https://localhost:8080/"} id="W97d6iQq772M" executionInfo={"status": "ok", "timestamp": 1620057497493, "user_tz": 300, "elapsed": 249, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="b511b84f-b405-4c53-f6fe-a87d95d1755d"
print('Accuracy: {:2.2%} '.format(metrics.accuracy_score(y_wine,
y_pred)))
# + [markdown] id="a7zdrdPfrW1s"
# <a name='1.1.1'></a>
#
# ### 1.1.1 Holdout Sets
#
# [back to top](#top)
#
# The way we account for unseen data, in practice, is to leave a portion of the dataset out for testing. This way, we can estimate how our model will perform on entirely new data it may come across in application.
# + colab={"base_uri": "https://localhost:8080/"} id="L6J-ZiMhY2Rb" executionInfo={"status": "ok", "timestamp": 1620058566884, "user_tz": 300, "elapsed": 211, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="aea2ebf3-57c4-45aa-fc1a-4bfdf853d8cd"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
knn = KNeighborsRegressor(n_neighbors=1)
knn.fit(X_train,y_train)
print(knn.score(X_test, y_test))
# + [markdown] id="ElRZqQdRZGWs"
# We see that we get a more reasonable value for our performance!
# + id="2uIEDsTBZFhm"
cols = wine.columns
cols = list(cols.drop(['type', 'type_encoding', 'quality', 'quality_encoding',
'quality_label']))
X_train, X_test, y_train, y_test = train_test_split(wine.loc[:, cols], wine['quality'],
test_size=0.2, random_state=42)
# + colab={"base_uri": "https://localhost:8080/"} id="T16H18oucuFV" executionInfo={"status": "ok", "timestamp": 1620058489772, "user_tz": 300, "elapsed": 338, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="b2d69f04-db5f-41be-eadd-e8a81c48d351"
model.fit(X_train, y_train)
# + id="YS2UizTxcutO"
y_pred = model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="T8zI9dGs20oN" executionInfo={"status": "ok", "timestamp": 1620057525968, "user_tz": 300, "elapsed": 210, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="e9681d21-cf25-40c3-d98f-b1b2636c6d81"
print('Accuracy: {:2.2%} '.format(metrics.accuracy_score(y_test,
y_pred)))
# + colab={"base_uri": "https://localhost:8080/"} id="LNva_uMm9SxM" executionInfo={"status": "ok", "timestamp": 1620057625689, "user_tz": 300, "elapsed": 216, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="b58672c8-d2bb-48ba-94a9-2f105002c101"
target_names = wine['quality'].unique
cm = metrics.confusion_matrix(y_true=y_test, y_pred=y_pred)
cm
# + [markdown] id="fcTkufrvrWv3"
# <a name='1.1.2'></a>
#
# ### 1.1.2 Data Leakage and Cross-Validation
#
# [back to top](#top)
#
# An even more rigorous method to leaving out a single test set, is to perform cross validation. Imagine a situation where we are trying to estimate the best value of K in our KNN algorithm. If we continually train our model with new values of K on our training set, and test with our testing set, "knowledge" of our test set values with leak into our model, as we choose the best value for K based on how it performs on our test set (even though we did not train on this test set). We call this phenomenon *data leakage*. CV or Cross Validation overcomes this by only evaluating our parameters with our training set.
#
# <p align="center">
# <img src='https://scikit-learn.org/stable/_images/grid_search_workflow.png' width=500px></img>
#
# <small>[image src](https://scikit-learn.org/stable/modules/cross_validation.html)</small>
# <p/>
#
# In this scheme, we don't evaluate our model on the test set until the very end. Rather, we estimate our hyperparameter performances by slicing the training set into cross folds
#
# <p align="center">
# <img src='https://scikit-learn.org/stable/_images/grid_search_cross_validation.png' width=500px></img>
#
# <small>[image src](https://scikit-learn.org/stable/modules/cross_validation.html)</small>
# </p>
#
#
# + id="gMUdmyQaiY6L"
from sklearn.model_selection import cross_val_score
scores = cross_val_score(knn, X_train, y_train, cv=5)
# + colab={"base_uri": "https://localhost:8080/"} id="OXE1NNecie6Q" executionInfo={"status": "ok", "timestamp": 1620058573898, "user_tz": 300, "elapsed": 207, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="ab359438-460d-4610-f195-c7d7b4be8a10"
scores
# + colab={"base_uri": "https://localhost:8080/"} id="IXES3jUtihv1" executionInfo={"status": "ok", "timestamp": 1620058593032, "user_tz": 300, "elapsed": 270, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="2303592f-267f-4f9a-f5cf-f8f8af676fd8"
print("%0.2f accuracy with a standard deviation of %0.3f" % (scores.mean(), scores.std()))
# + [markdown] id="tvLpUxd3i3L5"
# More information on the cross_val_score method in sklearn can be found [here](https://scikit-learn.org/stable/modules/cross_validation.html)
#
# An additional topic on cross validation is the extreme leave-one-out validation, you can read more about that [here](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html#Model-validation-via-cross-validation)
# + [markdown] id="1Y-bhAb0rWpm"
# <a name='1.1.3'></a>
#
# ### 1.1.3 Bias-Variance Tradeoff
#
# [back to top](#top)
#
# This next concept will be most easily understood (imo) if we go ahead an make up some data ourselves, I'm going to do that now.
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="sgq-OrwG0G5T" executionInfo={"status": "ok", "timestamp": 1620058676029, "user_tz": 300, "elapsed": 498, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="17ef5173-5f73-46c3-a5d9-e7f004cb1b4b"
# we can throttle the error rate
err = .5
random.seed(42)
# our data has a KNOWN underlying functional form (log(x))
def func(x, err):
return np.log(x) + err * random.randint(-1,1) * random.random()
x = np.arange(20,100)
y = [func(t, err) for t in x]
plt.plot(x,y, ls='', marker='.')
plt.xlabel('X')
plt.ylabel('Y')
# + [markdown] id="VZUc4fEBWFR4"
# Let's fit to just a portion of this data
# + id="H8mPD-xzWHFt"
random.seed(42)
X_train = random.sample(list(x), 10)
indices = [list(x).index(i) for i in X_train]
# we could also do it this way: np.argwhere([i in X_train for i in x])
y_train = [y[i] for i in indices]
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="mwsSWyitQ7kA" executionInfo={"status": "ok", "timestamp": 1620058694716, "user_tz": 300, "elapsed": 756, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="8f552d45-7e91-431e-ac82-07f282d0fe3b"
plt.plot(X_train,y_train, ls='', marker='.')
# + [markdown] id="JqF5qNQLTBd8"
# Now let's take two extreme scenarios, fitting a linear line and a high order polynomial, to these datapoints. Keeping in mind the larger dataset, as well as the error we introduced in our data generating function, this will really illustrate our point!
# + colab={"base_uri": "https://localhost:8080/", "height": 306} id="chkTfkJFULzm" executionInfo={"status": "ok", "timestamp": 1620058758871, "user_tz": 300, "elapsed": 827, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="2afb20bf-213c-47cd-d4a3-d0d0fcc03560"
# solving our training data with a n-degree polynomial
coefs = np.polyfit(X_train, y_train, 9)
# solve the slope and intercept of our 1-degree polynomial ;)
model = LinearRegression()
model.fit(np.array(X_train).reshape(-1,1), y_train)
# create some x data to plot our functions
X_seq = np.linspace(min(X_train),max(X_train),300).reshape(-1,1)
fig, ax = plt.subplots(1,2,figsize=(15,5))
ax[0].plot(X_seq, model.predict(X_seq), c='grey', ls='--')
ax[0].plot(X_train, y_train, ls='', marker='.')
ax[0].set_ylim(min(y_train), max(y_train))
ax[0].set_title("High Bias Model")
ax[1].plot(X_seq, np.polyval(coefs, X_seq), c='grey', ls='--')
ax[1].plot(X_train, y_train, ls='', marker='.')
ax[1].set_ylim(min(y_train), max(y_train))
ax[1].set_title("High Variance Model")
# + [markdown] id="q6a2moL6bud6"
# We've demonstrated two extreme cases. On the left, we limit our regression to only two parameters, a slope and a y-intercept. We say that this model has *high bias* because we are forcing the functional form without much consideration to the underlying data — we are saying this data is generated by a linear function, and no matter what data I train on, my final model will still be a straight line that more or less appears the same. Put another way, it has *low variance* with respect to the underlying data.
#
# On the right, we've allowed our model just as many polynomials it needs to perfectly fit the training data! We say this model has *low bias* because we don't introduce many constraints on the final form of the model. it is *high variance* because depending on the underlying training data, the final outcome of the model can change quite drastically!
#
# In reality, the best model lies somewhere between these two cases. In the next few paragraphs we'll explore this concept further:
#
# 1. what happens when we retrain these models on different samples of the data population
# * and let's use this to better understand what we mean by *bias* and *variance*
# 2. what happens when we tie this back in with the error we introduced to the data generator?
# * and let's use this to better understand irreducible error
# + colab={"base_uri": "https://localhost:8080/", "height": 289} id="_PYu7H13nHSk" executionInfo={"status": "ok", "timestamp": 1620058847831, "user_tz": 300, "elapsed": 834, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="3e6764c7-f00d-4842-afa7-de1e9608d255"
random.seed(42)
fig, ax = plt.subplots(1,2,figsize=(15,5))
for samples in range(5):
X_train = random.sample(list(x), 10)
indices = [list(x).index(i) for i in X_train]
y_train = [y[i] for i in indices]
# solving our training data with a n-degree polynomial
coefs = np.polyfit(X_train, y_train, 9)
# solve the slope and intercept of our 1-degree polynomial ;)
model = LinearRegression()
model.fit(np.array(X_train).reshape(-1,1), y_train)
# create some x data to plot our functions
X_seq = np.linspace(min(X_train),max(X_train),300).reshape(-1,1)
ax[0].plot(X_seq, model.predict(X_seq), alpha=0.5, ls='--')
ax[0].plot(X_train, y_train, ls='', marker='.')
ax[0].set_ylim(min(y_train), max(y_train))
ax[0].set_title("High Bias Model")
ax[1].plot(X_seq, np.polyval(coefs, X_seq), alpha=0.5, ls='--')
ax[1].plot(X_train, y_train, ls='', marker='.')
ax[1].set_ylim(min(y_train), max(y_train))
ax[1].set_title("High Variance Model")
# + [markdown] id="er3Vvoqyn8zH"
# As we can see, depending on what data we train our model on, the *high bias* model changes relatively slightly, while the *high variance* model changes a whole awful lot!
#
# The *high variance* model is prone to something we call *overfitting*. It fits the training data very well, but at the expense of creating a good, generalizable model that does well on unseen data. Let's take our last models, and plot them along the rest of the unseen data, what we'll call the *population*
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="b8TubrVIqTAD" executionInfo={"status": "ok", "timestamp": 1619022018315, "user_tz": 300, "elapsed": 9611, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="da8c3815-c641-419e-99a3-6a7ac6ded22f"
# solving our training data with a n-degree polynomial
coefs = np.polyfit(X_train, y_train, 9)
# solve the slope and intercept of our 1-degree polynomial ;)
model = LinearRegression()
model.fit(np.array(X_train).reshape(-1,1), y_train)
# create some x data to plot our functions
X_seq = np.linspace(min(X_train),max(X_train),300).reshape(-1,1)
fig, ax = plt.subplots(1,2,figsize=(15,5))
ax[0].plot(X_seq, model.predict(X_seq), c='grey', ls='--')
ax[0].plot(x, y, ls='', marker='*', alpha=0.6)
ax[0].plot(X_train, y_train, ls='', marker='.')
ax[0].set_ylim(min(y), max(y))
ax[0].set_title("High Bias Model")
ax[1].plot(X_seq, np.polyval(coefs, X_seq), c='grey', ls='--')
ax[1].plot(x, y, ls='', marker='*', alpha=0.6)
ax[1].plot(X_train, y_train, ls='', marker='.')
ax[1].set_ylim(min(y), max(y))
ax[1].set_title("High Variance Model")
# + [markdown] id="9mtZ-VTNronN"
# In particular, we see that the high variance model is doing very wacky things, demonstrating behaviors in the model where the underlying population data really gives no indication of such behavior. We say that these high variance model are particuarly prone to the phenomenon of *over fitting* and this is generally due to the fact that there is irreducible error in the underlying data. Let's demonstrate this.
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="MUMgvt2pt-Wp" executionInfo={"status": "ok", "timestamp": 1619022018507, "user_tz": 300, "elapsed": 9790, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="f692c0f4-150a-4438-c794-10ce5e92ec6a"
x = np.arange(20,100)
y = [func(t, err=0) for t in x]
plt.plot(x,y, ls='', marker='.')
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="aG_Th8lxu6-Z" executionInfo={"status": "ok", "timestamp": 1619022019674, "user_tz": 300, "elapsed": 10948, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="16727b71-0661-45b5-ee57-6f8b281433c6"
random.seed(42)
X_train = random.sample(list(x), 10)
indices = [list(x).index(i) for i in X_train]
# we could also do it this way: np.argwhere([i in X_train for i in x])
y_train = [y[i] for i in indices]
# solving our training data with a n-degree polynomial
coefs = np.polyfit(X_train, y_train, 9)
# solve the slope and intercept of our 1-degree polynomial ;)
model = LinearRegression()
model.fit(np.array(X_train).reshape(-1,1), y_train)
# create some x data to plot our functions
X_seq = np.linspace(min(X_train),max(X_train),300).reshape(-1,1)
fig, ax = plt.subplots(1,2,figsize=(15,5))
ax[0].plot(X_seq, model.predict(X_seq), c='grey', ls='--')
ax[0].plot(x, y, ls='', marker='o', alpha=0.2)
ax[0].plot(X_train, y_train, ls='', marker='.')
ax[0].set_ylim(min(y), max(y))
ax[0].set_title("High Bias Model")
ax[1].plot(X_seq, np.polyval(coefs, X_seq), c='grey', ls='--')
ax[1].plot(x, y, ls='', marker='o', alpha=0.2)
ax[1].plot(X_train, y_train, ls='', marker='.')
ax[1].set_ylim(min(y), max(y))
ax[1].set_title("High Variance Model")
# + [markdown] id="H-psbne6vKXa"
# This time, our high variance model really *gets it*! And this is because the data we trained on actually *is* a good representation of the entire population. But this, in reality, almost never, ever happens. In the real world, we have irreducible error in our data samples, and we must account for this when choosing our model.
#
# I'm summary, we call this balance between error in our model functional form, and error from succumbing to irreducible error in our training data, the *bias variance tradeoff*
# + [markdown] id="QdPtUGUJt-Ib"
# #### 1.1.3.1 Exercise: Quantitatively Define Performance
#
# Up until now, we've explored this idea of bias variance tradeoff from a qualitative standpoint. As an exercise, continue with this idea, this time calculating the mean squared error (MSE) and R-square between the model and UNSEEN (non-training data) population data.
#
# Do this for a 9th order polynomial and repeat for population data with low, med, and high degrees of error. Be sure to seed any random number generator to ensure reproducibility
# + id="P-iju-_vKAEX" executionInfo={"status": "ok", "timestamp": 1620423882272, "user_tz": 300, "elapsed": 269, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}}
# Code Cell for Exercise 1.1.3.1
# + [markdown] id="O9u-hhgErWiU"
# <a name='1.1.4'></a>
#
# ### 1.1.4 Learning Curves
#
# [back to top](#top)
#
# To move from qualitative to quantitative understanding of bias-variance tradeoff we need to introduce some metric for model performance. A good one to use here is R-square, a measure of the degree to which predictions match actual values. We can import a tool from sklearn to calculate this for us.
#
#
# + id="dXj-4YAYzbmo"
from sklearn.metrics import r2_score
# + [markdown] id="4JEEWq3x00X0"
# <a name='x.1.4.1'></a>
#
# #### 1.1.4.1 Considering Model Complexity
#
# [back to top](#top)
#
# In a learning curve, we will typically plot the training and testing scores together, to give a sense of when we have either too much bias or too much variance in our model.
#
# I'm going to go ahead and recreate the original data distribution we introduced in 1.1.3
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="G6BGqnPc0wdE" executionInfo={"status": "ok", "timestamp": 1620059182055, "user_tz": 300, "elapsed": 430, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="4c385855-4487-490f-9421-27d32fad745f"
# we can throttle the error rate
err = .5
random.seed(42)
# our data has a KNOWN underlying functional form (log(x))
def func(x, err):
return np.log(x) + err * random.randint(-1,1) * random.random()
x = np.arange(20,100)
y = [func(t, err) for t in x]
plt.plot(x,y, ls='', marker='.')
# + [markdown] id="VJsU_dU71xla"
# Now let's itteratively introduce more complexity into our model
# + colab={"base_uri": "https://localhost:8080/", "height": 353} id="ba5YUVmP1zwO" executionInfo={"status": "ok", "timestamp": 1620059189225, "user_tz": 300, "elapsed": 805, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="c6afba86-af14-4111-ccb6-bad5560ce694"
random.seed(42)
fig, ax = plt.subplots(1,2,figsize=(10,5))
X_train = random.sample(list(x), 10)
indices = [list(x).index(i) for i in X_train]
y_train = [y[i] for i in indices]
ax[0].plot(X_train, y_train, ls='', marker='.', color='black')
for complexity in range(1,10):
# solving our training data with a n-degree polynomial
coefs = np.polyfit(X_train, y_train, complexity)
# create some x data to plot our functions
X_seq = np.linspace(min(X_train),max(X_train),300).reshape(-1,1)
score = r2_score(np.polyval(coefs, X_train), y_train)
ax[0].plot(X_seq, np.polyval(coefs, X_seq), alpha=0.5, ls='--')
ax[0].set_ylim(min(y_train), max(y_train))
ax[0].set_title("Predictions with Increasing Model Complexity")
ax[1].plot(complexity, score, ls='', marker='.',
label='{}-poly, {:.2f}-score'.format(complexity, score))
ax[1].set_title("Scores with Increasing Model Complexity")
ax[1].legend()
# + [markdown] id="4-ikwp8HtIUz"
# As we see from both plots, the score on the training data increases with added model complexity. Giving us the expected perfect fit when the order is the same as the number of data points! This is part I of our learning curve. Part II consists of plotting the training data score with the testing data score.
#
# Something else I'm going to do, is define the training portion of the data as a fraction of the overall population size. To keep the comparisons the same as up until now, I will keep this training fraction low at .2
# + colab={"base_uri": "https://localhost:8080/", "height": 353} id="2GWQnzfStW_H" executionInfo={"status": "ok", "timestamp": 1620059992589, "user_tz": 300, "elapsed": 1077, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="6f71316e-17bb-4c5c-8d9b-6f059b19473c"
random.seed(42)
# defining my training fraction
training_frac = .2
# create test and training data
X_train = random.sample(list(x), int(int(len(x))*training_frac))
train_indices = [list(x).index(i) for i in X_train]
y_train = [y[i] for i in train_indices]
test_indices = [i for i in range(len(x)) if i not in train_indices]
X_test = [x[i] for i in test_indices]
y_test = [y[i] for i in test_indices]
# initialize the plot and display the data
fig, ax = plt.subplots(1,2,figsize=(10,5))
ax[0].plot(X_train, y_train, ls='', marker='.', color='black')
ax[0].plot(X_test, y_test, ls='', marker='.', color='grey', alpha=0.5)
for complexity in range(1,10):
# solving our training data with a n-degree polynomial
coefs = np.polyfit(X_train, y_train, complexity)
# create some x data to plot our functions
X_seq = np.linspace(min(X_train),max(X_train),300).reshape(-1,1)
score1 = r2_score(np.polyval(coefs, X_train), y_train)
score2 = r2_score(np.polyval(coefs, X_test), y_test)
ax[0].plot(X_seq, np.polyval(coefs, X_seq), alpha=0.5, ls='--',
label='{}-poly, {:.2f}-score'.format(complexity, score2))
ax[0].set_ylim(min(y_train), max(y_train))
ax[0].set_title("Predictions with Increasing Model Complexity")
ax[1].plot(complexity, score1, ls='', marker='.', color='blue',
label='{}-poly, {:.2f}-score'.format(complexity, score1))
ax[1].plot(complexity, score2, ls='', marker='o', color='red',
label='{}-poly, {:.2f}-score'.format(complexity, score2))
ax[1].set_title("Scores with Increasing Model Complexity")
ax[1].legend(['Train $R^2$', 'Test $R^2$'])
ax[0].legend()
# + [markdown] id="oJuVDgBs0RD-"
# As we can see, The 2nd order polynomial achieves the greatest best test set data $R^2$, while the highest order polynomial achieves the best training set data $R^2$. This learning curve is explanative of what we see generally, namely a divergence after some degree of complexity between training and test set performances. In this case, we would resolve to choose the 2nd order polynomial as the best model for our data.
#
# <img src="https://jakevdp.github.io/PythonDataScienceHandbook/figures/05.03-validation-curve.png" width=500px></img>
#
# <small>[img src](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html)</small>
#
#
# + [markdown] id="kSIXqSgS3jdB"
# <a name='x.1.4.2'></a>
#
# #### 1.1.4.2 Considering Training Set Size
#
# [back to top](#top)
#
# The last piece of the puzzle we require, to fully cover learning curves, is the effect of training data size on the model. This is why I introduced the 'fraction of training data' parameter earlier. Let's explore.
# + id="_7Qd0jUS4Qan" colab={"base_uri": "https://localhost:8080/", "height": 455} executionInfo={"status": "ok", "timestamp": 1620060176926, "user_tz": 300, "elapsed": 773, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="436e31e9-98d1-4091-cefa-cfa81f351012"
random.seed(42)
# initialize the plot and display the data
fig, ax = plt.subplots(1,1,figsize=(10,5))
for training_frac in np.linspace(0.1,.9,50):
# create test and training data
X_train = random.sample(list(x), int(int(len(x))*training_frac))
indices = [list(x).index(i) for i in X_train]
y_train = [y[i] for i in indices]
test_indices = [i for i in range(len(x)) if i not in indices]
X_test = [x[i] for i in test_indices]
y_test = [y[i] for i in test_indices]
# solving our training data with a n-degree polynomial
coefs = np.polyfit(X_train, y_train, 9)
score1 = r2_score(np.polyval(coefs, X_train), y_train)
score2 = r2_score(np.polyval(coefs, X_test), y_test)
ax.plot(training_frac, score1, ls='', marker='.', color='blue',
label='{}-poly, {:.2f}-score'.format(training_frac, score1))
ax.plot(training_frac, score2, ls='', marker='o', color='red',
label='{}-poly, {:.2f}-score'.format(training_frac, score2))
ax.set_title("9th-order Polynomial Score with Increasing Training Set Size")
ax.legend(['Train','Test'])
ax.set_xlabel('Training Fraction')
ax.set_ylabel('$R^2$')
# + [markdown] id="YpQog5q6kJyI"
# What we see here is a trend that happens generally, as our amount of training data increases, our models handle more complexity. This is illustrated below.
#
# <img src="https://jakevdp.github.io/PythonDataScienceHandbook/figures/05.03-learning-curve.png" width=500px></img>
#
# <small>[img src](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html)
# + [markdown] id="_HNtge3hXHXF"
# #### 1.1.4.3 Exercise: Visualization
#
# Starting with the code from block 1.1.4.2, make a side-by-side plot of a 3rd degree polynomial and a 12th degree polynomial. On the x axis slowly increase the training set size, on the y axis plot the scores for the training and test sets.
# + id="I4GvFJ51KfC4" executionInfo={"status": "ok", "timestamp": 1620423908328, "user_tz": 300, "elapsed": 263, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}}
# Code Cell for Exercise 1.1.4.3
# + [markdown] id="X4T8i4CHl7bf"
# > As a visualization exercise, how would you attempt to combine the ideas of model performance with increasing training set size and increasing model complexity? Could you create this visualization with something other than a polynomial model?
# + [markdown] id="ponptvjBsA-a"
# <a name='1.2'></a>
#
# ## 1.2 Model Validation in Practice
#
# [back to top](#top)
#
# We will now turn our attention to practical implementation.
#
# In practice, there are a wide number of variables (called hyperparameters) to consider when choosing a model. Scikit learn has a useful method called Grid Search that will iterate through every possible combination of a range of hyperparameter settings you provide as input.
#
# Before we get started with grid search, we'll need to switch over from our numpy polynomial fit method to one in sklearn. Here, the caveat is our actual model will solve for the *coefficients* infront of the polynomials. We will *engineer* the polynomial features ourselves. This is an example of *feature engineering* which we will revisit in depth in a later session.
# + id="SsSjSzjHm_gD"
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import make_pipeline
def PolynomialRegression(degree=2, **kwargs):
return make_pipeline(PolynomialFeatures(degree),
LinearRegression(**kwargs))
# + [markdown] id="w1bOUNYpsA3T"
# <a name='1.2.1'></a>
#
# ### 1.2.1 Grid Search
#
# [back to top](#top)
#
#
# + id="oj0_BOSEpAR8"
from sklearn.model_selection import GridSearchCV
param_grid = {'polynomialfeatures__degree': np.arange(10),
'linearregression__fit_intercept': [True, False],
'linearregression__normalize': [True, False]}
grid = GridSearchCV(PolynomialRegression(), param_grid, cv=7)
# + id="NhQvExqTpZ58"
# create test and training data
random.seed(42)
X_train = random.sample(list(x), int(int(len(x))*.8))
indices = [list(x).index(i) for i in X_train]
y_train = [y[i] for i in indices]
test_indices = [i for i in range(len(x)) if i not in indices]
X_test = [x[i] for i in test_indices]
y_test = [y[i] for i in test_indices]
# + colab={"base_uri": "https://localhost:8080/"} id="JKBIvPSkpQX1" executionInfo={"status": "ok", "timestamp": 1620060629325, "user_tz": 300, "elapsed": 989, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="889a6458-0f6f-4f15-ac1e-16ad9e5e3565"
grid.fit(np.array(X_train).reshape(-1,1), y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="jr8BoNbWppOA" executionInfo={"status": "ok", "timestamp": 1620060639318, "user_tz": 300, "elapsed": 193, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="9828b587-2dd5-4294-aecf-b1e7cd1ee19f"
grid.best_params_
# + [markdown] id="x0wP5Q0WqIfA"
# to grab the best model from the CV/search outcome. we use grid.best_estimator
# + colab={"base_uri": "https://localhost:8080/", "height": 304} id="gIF1LEQvqHvN" executionInfo={"status": "ok", "timestamp": 1620060649196, "user_tz": 300, "elapsed": 526, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gis2tewog0nYcz7REtNxkAs58_fKdVn5wvb3mXkPQ=s64", "userId": "17051665784581118920"}} outputId="2d40ce36-30fe-4f0c-abbd-6f73f2c9c14e"
model = grid.best_estimator_
# create some x data to plot our functions
X_seq = np.linspace(min(X_train),max(X_train),300).reshape(-1,1)
fig, ax = plt.subplots(1,1,figsize=(15,5))
ax.plot(X_seq, model.predict(X_seq), c='grey', ls='--')
ax.plot(x, y, ls='', marker='*', alpha=0.6)
ax.plot(X_train, y_train, ls='', marker='.')
ax.set_ylim(min(y), max(y))
ax.set_title("Best Grid Search CV Model")
# + [markdown] id="BhS-p_SSvIRK"
# <a name='reference'></a>
#
# # References
#
# [back to top](#top)
#
# ## Model Validation
#
# * [cross_val_score](https://scikit-learn.org/stable/modules/cross_validation.html)
# * [leave-one-out](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html#Model-validation-via-cross-validation)
# + id="DoLMulhuk5Xk"
| courses/C2_ Build Your Model/DSE C2 S1_ Model Selection and Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # ICESat-2's Nested Variables
#
# This notebook ({nb-download}`download <IS2_data_variables.ipynb>`) illustrates the use of icepyx for managing lists of available and wanted ICESat-2 data variables.
# The two use cases for variable management within your workflow are:
# 1. During the data access process, whether that's via order and download (e.g. via NSIDC DAAC) or remote (e.g. via the cloud).
# 2. When reading in data to a Python object (whether from local files or the cloud).
#
# A given ICESat-2 product may have over 200 variable + path combinations.
# icepyx includes a custom `Variables` module that is "aware" of the ATLAS sensor and how the ICESat-2 data products are stored.
# The module can be accessed independently, but is optimally used as a component of a `Query` object (Case 1) or `Read` object (Case 2).
#
# This notebook illustrates in detail how the `Variables` module behaves using a `Query` data access example.
# However, module usage is analogous through an icepyx ICESat-2 `Read` object.
# More detailed example workflows specifically for the [query](https://icepyx.readthedocs.io/en/latest/example_notebooks/IS2_data_access.html) and [read](https://icepyx.readthedocs.io/en/latest/example_notebooks/IS2_data_read-in.html) tools within icepyx are available as separate Jupyter Notebooks.
#
# Questions? Be sure to check out the FAQs throughout this notebook, indicated as italic headings.
# ### _Why do ICESat-2 products need a custom variable manager?_
#
# _It can be confusing and cumbersome to comb through the 200+ variable and path combinations contained in ICESat-2 data products._
# _The icepyx `Variables` module makes it easier for users to quickly find and extract the specific variables they would like to work with across multiple beams, keywords, and variables and provides reader-friendly formatting to browse variables._
# _A future development goal for `icepyx` includes developing an interactive widget to further improve the user experience._
# _For data read-in, additional tools are available to target specific beam characteristics (e.g. strong versus weak beams)._
# #### Some technical details about the Variables module
# For those eager to push the limits or who want to know more implementation details...
#
# The only required input to the `Variables` module is `vartype`.
# `vartype` has two acceptible string values, 'order' and 'file'.
# If you use the module as shown in icepyx examples (namely through a `Read` or `Query` object), then this flag will be passed automatically.
# It simply tells the software how to generate the list of possible variable values - either by pinging NSIDC for a list of available variables (`query`) or from the user-supplied file (`read`).
# Import packages, including icepyx
import icepyx as ipx
from pprint import pprint
# ## Interacting with ICESat-2 Data Variables
#
# Each variables instance (which is actually an associated Variables class object) contains two variable list attributes.
# One is the list of possible or available variables (`avail` attribute) and is unmutable, or unchangeable, as it is based on the input product specifications or files.
# The other is the list of variables you'd like to actually have (in your downloaded file or data object) from all the potential options (`wanted` attribute) and is updateable.
#
# Thus, your `avail` list depends on your data source and whether you are accessing or reading data, while your `wanted` list may change for each analysis you are working on or depending on what variables you want to see.
#
# The variables parameter has methods to:
# * get a list of all available variables, either available from the NSIDC or the file (`avail()` method).
# * append new variables to the wanted list (`append()` method).
# * remove variables from the wanted list (`remove()` method).
#
# We'll showcase the use of all of these methods and attributes below using an `icepyx.Query` object.
# Usage is identical in the case of an `icepyx.Read` object.
# More detailed example workflows specifically for the [query](https://icepyx.readthedocs.io/en/latest/example_notebooks/IS2_data_access.html) and [read](https://icepyx.readthedocs.io/en/latest/example_notebooks/IS2_data_read-in.html) tools within icepyx are available as separate Jupyter Notebooks.
#
# Create a query object and log in to Earthdata
#
# For this example, we'll be working with a land ice product (ATL06) for an area along West Greenland (Disko Bay).
# A second option for an atmospheric product (ATL09) that uses profiles instead of the ground track (gt) categorization is also provided.
region_a = ipx.Query('ATL06',[-55, 68, -48, 71],['2019-02-22','2019-02-28'], \
start_time='00:00:00', end_time='23:59:59')
# +
# Uncomment and run the code in this cell to use the second variable subsetting suite of examples,
# with the beam specifier containing "profile" instead of "gt#l"
# region_a = ipx.Query('ATL09',[-55, 68, -48, 71],['2019-02-22','2019-02-28'], \
# start_time='00:00:00', end_time='23:59:59')
# -
region_a.earthdata_login('icepyx_devteam','<EMAIL>')
# ### ICESat-2 data variables
#
# ICESat-2 data is natively stored in a nested file format called hdf5.
# Much like a directory-file system on a computer, each variable (file) has a unique path through the heirarchy (directories) within the file.
# Thus, some variables (e.g. `'latitude'`, `'longitude'`) have multiple paths (one for each of the six beams in most products).
#
# #### Determine what variables are available
# `region_a.order_vars.avail` will return a list of all valid path+variable strings.
region_a.order_vars.avail()
# To increase readability, you can use built in functions to show the 200+ variable + path combinations as a dictionary where the keys are variable names and the values are the paths to that variable.
# `region_a.order_vars.parse_var_list(region_a.order_vars.avail())` will return a dictionary of variable:paths key:value pairs.
region_a.order_vars.parse_var_list(region_a.order_vars.avail())
# By passing the boolean `options=True` to the `avail` method, you can obtain lists of unique possible variable inputs (var_list inputs) and path subdirectory inputs (keyword_list and beam_list inputs) for your data product. These can be helpful for building your wanted variable list.
region_a.order_vars.avail(options=True)
# ### Building your wanted variable list
#
# Now that you know which variables and path components are available, you need to build a list of the ones you'd like included.
# There are several options for generating your initial list as well as modifying it, giving the user complete control.
#
# The options for building your initial list are:
# 1. Use a default list for the product (not yet fully implemented across all products. Have a default variable list for your field/product? Submit a pull request or post it as an issue on [GitHub](https://github.com/icesat2py/icepyx)!)
# 2. Provide a list of variable names
# 3. Provide a list of profiles/beams or other path keywords, where "keywords" are simply the unique subdirectory names contained in the full variable paths of the product. A full list of available keywords for the product is displayed in the error message upon entering `keyword_list=['']` into the `append` function (see below for an example) or by running `region_a.order_vars.avail(options=True)`, as above.
#
# **Note: all products have a short list of "mandatory" variables/paths (containing spacecraft orientation and time information needed to convert the data's `delta_time` to a readable datetime) that are automatically added to any built list. If you have any recommendations for other variables that should always be included (e.g. uncertainty information), please let us know!**
#
# Examples of using each method to build and modify your wanted variable list are below.
region_a.order_vars.wanted
region_a.order_vars.append(defaults=True)
pprint(region_a.order_vars.wanted)
# The keywords available for this product are shown in the error message upon entering a blank keyword_list, as seen in the next cell.
region_a.order_vars.append(keyword_list=[''])
# ### Modifying your wanted variable list
#
# Generating and modifying your variable request list, which is stored in `region_a.order_vars.wanted`, is controlled by the `append` and `remove` functions that operate on `region_a.order_vars.wanted`. The input options to `append` are as follows (the full documentation for this function can be found by executing `help(region_a.order_vars.append)`).
# * `defaults` (default False) - include the default variable list for your product (not yet fully implemented for all products; please submit your default variable list for inclusion!)
# * `var_list` (default None) - list of variables (entered as strings)
# * `beam_list` (default None) - list of beams/profiles (entered as strings)
# * `keyword_list` (default None) - list of keywords (entered as strings); use `keyword_list=['']` to obtain a list of available keywords
#
# Similarly, the options for `remove` are:
# * `all` (default False) - reset `region_a.order_vars.wanted` to None
# * `var_list` (as above)
# * `beam_list` (as above)
# * `keyword_list` (as above)
region_a.order_vars.remove(all=True)
pprint(region_a.order_vars.wanted)
# ### Examples (Overview)
# Below are a series of examples to show how you can use `append` and `remove` to modify your wanted variable list.
# For clarity, `region_a.order_vars.wanted` is cleared at the start of many examples.
# However, multiple `append` and `remove` commands can be called in succession to build your wanted variable list (see Examples 3+).
#
# There are two example tracks.
# The first is for land ice (ATL06) data that is separated into beams.
# The second is for atmospheric data (ATL09) that is separated into profiles.
# Both example tracks showcase the same functionality and are provided for users of both data types.
# ------------------
# ### Example Track 1 (Land Ice - run with ATL06 dataset)
#
# #### Example 1.1: choose variables
# Add all `latitude` and `longitude` variables across all six beam groups. Note that the additional required variables for time and spacecraft orientation are included by default.
region_a.order_vars.append(var_list=['latitude','longitude'])
pprint(region_a.order_vars.wanted)
# #### Example 1.2: specify beams and variable
# Add `latitude` for only `gt1l` and `gt2l`
region_a.order_vars.remove(all=True)
pprint(region_a.order_vars.wanted)
var_dict = region_a.order_vars.append(beam_list=['gt1l', 'gt2l'], var_list=['latitude'])
pprint(region_a.order_vars.wanted)
# #### Example 1.3: add/remove selected beams+variables
# Add `latitude` for `gt3l` and remove it for `gt2l`
region_a.order_vars.append(beam_list=['gt3l'],var_list=['latitude'])
region_a.order_vars.remove(beam_list=['gt2l'], var_list=['latitude'])
pprint(region_a.order_vars.wanted)
# #### Example 1.4: `keyword_list`
# Add `latitude` and `longitude` for all beams and with keyword `land_ice_segments`
region_a.order_vars.append(var_list=['latitude', 'longitude'],keyword_list=['land_ice_segments'])
pprint(region_a.order_vars.wanted)
# #### Example 1.5: target a specific variable + path
# Remove `gt1r/land_ice_segments/longitude` (but keep `gt1r/land_ice_segments/latitude`)
region_a.order_vars.remove(beam_list=['gt1r'], var_list=['longitude'], keyword_list=['land_ice_segments'])
pprint(region_a.order_vars.wanted)
# #### Example 1.6: add variables not specific to beams/profiles
# Add `rgt` under `orbit_info`.
region_a.order_vars.append(keyword_list=['orbit_info'],var_list=['rgt'])
pprint(region_a.order_vars.wanted)
# #### Example 1.7: add all variables+paths of a group
# In addition to adding specific variables and paths, we can filter all variables with a specific keyword as well. Here, we add all variables under `orbit_info`. Note that paths already in `region_a.order_vars.wanted`, such as `'orbit_info/rgt'`, are not duplicated.
region_a.order_vars.append(keyword_list=['orbit_info'])
pprint(region_a.order_vars.wanted)
# #### Example 1.8: add all possible values for variables+paths
# Append all `longitude` paths and all variables/paths with keyword `land_ice_segments`.
#
# Similarly to what is shown in Example 4, if you submit only one `append` call as `region_a.order_vars.append(var_list=['longitude'], keyword_list=['land_ice_segments'])` rather than the two `append` calls shown below, you will only add the variable `longitude` and only paths containing `land_ice_segments`, not ALL paths for `longitude` and ANY variables with `land_ice_segments` in their path.
region_a.order_vars.append(var_list=['longitude'])
region_a.order_vars.append(keyword_list=['land_ice_segments'])
pprint(region_a.order_vars.wanted)
# #### Example 1.9: remove all variables+paths associated with a beam
# Remove all paths for `gt1l` and `gt3r`
region_a.order_vars.remove(beam_list=['gt1l','gt3r'])
pprint(region_a.order_vars.wanted)
# #### Example 1.10: generate a default list for the rest of the tutorial
# Generate a reasonable variable list prior to download
region_a.order_vars.remove(all=True)
region_a.order_vars.append(defaults=True)
pprint(region_a.order_vars.wanted)
# ------------------
# ### Example Track 2 (Atmosphere - run with ATL09 dataset commented out at the start of the notebook)
#
# #### Example 2.1: choose variables
# Add all `latitude` and `longitude` variables
region_a.order_vars.append(var_list=['latitude','longitude'])
pprint(region_a.order_vars.wanted)
# #### Example 2.2: specify beams/profiles and variable
# Add `latitude` for only `profile_1` and `profile_2`
region_a.order_vars.remove(all=True)
pprint(region_a.order_vars.wanted)
var_dict = region_a.order_vars.append(beam_list=['profile_1','profile_2'], var_list=['latitude'])
pprint(region_a.order_vars.wanted)
# #### Example 2.3: add/remove selected beams+variables
# Add `latitude` for `profile_3` and remove it for `profile_2`
region_a.order_vars.append(beam_list=['profile_3'],var_list=['latitude'])
region_a.order_vars.remove(beam_list=['profile_2'], var_list=['latitude'])
pprint(region_a.order_vars.wanted)
# #### Example 2.4: `keyword_list`
# Add `latitude` for all profiles and with keyword `low_rate`
region_a.order_vars.append(var_list=['latitude'],keyword_list=['low_rate'])
pprint(region_a.order_vars.wanted)
# #### Example 2.5: target a specific variable + path
# Remove `'profile_1/high_rate/latitude'` (but keep `'profile_3/high_rate/latitude'`)
region_a.order_vars.remove(beam_list=['profile_1'], var_list=['latitude'], keyword_list=['high_rate'])
pprint(region_a.order_vars.wanted)
# #### Example 2.6: add variables not specific to beams/profiles
# Add `rgt` under `orbit_info`.
region_a.order_vars.append(keyword_list=['orbit_info'],var_list=['rgt'])
pprint(region_a.order_vars.wanted)
# #### Example 2.7: add all variables+paths of a group
# In addition to adding specific variables and paths, we can filter all variables with a specific keyword as well. Here, we add all variables under `orbit_info`. Note that paths already in `region_a.order_vars.wanted`, such as `'orbit_info/rgt'`, are not duplicated.
region_a.order_vars.append(keyword_list=['orbit_info'])
pprint(region_a.order_vars.wanted)
# #### Example 2.8: add all possible values for variables+paths
# Append all `longitude` paths and all variables/paths with keyword `high_rate`.
# Simlarly to what is shown in Example 4, if you submit only one `append` call as `region_a.order_vars.append(var_list=['longitude'], keyword_list=['high_rate'])` rather than the two `append` calls shown below, you will only add the variable `longitude` and only paths containing `high_rate`, not ALL paths for `longitude` and ANY variables with `high_rate` in their path.
region_a.order_vars.append(var_list=['longitude'])
region_a.order_vars.append(keyword_list=['high_rate'])
pprint(region_a.order_vars.wanted)
# #### Example 2.9: remove all variables+paths associated with a profile
# Remove all paths for `profile_1` and `profile_3`
region_a.order_vars.remove(beam_list=['profile_1','profile_3'])
pprint(region_a.order_vars.wanted)
# #### Example 2.10: generate a default list for the rest of the tutorial
# Generate a reasonable variable list prior to download
region_a.order_vars.remove(all=True)
region_a.order_vars.append(defaults=True)
pprint(region_a.order_vars.wanted)
# ### Using your wanted variable list
#
# Now that you have your wanted variables list, you need to use it within your icepyx object (`Query` or `Read`) will automatically use it.
# #### With a `Query` object
# In order to have your wanted variable list included with your order, you must pass it as a keyword argument to the `subsetparams()` attribute or the `order_granules()` or `download_granules()` (which calls `order_granules` under the hood if you have not already placed your order) functions.
region_a.subsetparams(Coverage=region_a.order_vars.wanted)
# Or, you can put the `Coverage` parameter directly into `order_granules`:
# `region_a.order_granules(Coverage=region_a.order_vars.wanted)`
#
# However, then you cannot view your subset parameters (`region_a.subsetparams`) prior to submitting your order.
region_a.order_granules()# <-- you do not need to include the 'Coverage' kwarg to
# order if you have already included it in a call to subsetparams
region_a.download_granules('/home/jovyan/icepyx/dev-notebooks/vardata') # <-- you do not need to include the 'Coverage' kwarg to
# download if you have already submitted it with your order
# #### With a `Read` object
# Calling the `load()` method on your `Read` object will automatically look for your wanted variable list and use it.
# Please see the [read-in example Jupyter Notebook](https://icepyx.readthedocs.io/en/latest/example_notebooks/IS2_data_read-in.html) for a complete example of this usage.
#
# #### Credits
# * based on the subsetting notebook by: <NAME> and <NAME>
| doc/source/example_notebooks/IS2_data_variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import camb
import pyhmcode
def massfunction_params(plot=False):
camb.set_feedback_level(0)
# Cosmological parameters for CAMB
h = 0.7
omc = 0.25
omb = 0.048
mnu = 0.06
w = -1.0
wa = 0.0
ns = 0.97
As = 2.1e-9
n_z = 4
k_max = 20.0
z_lin = np.linspace(0, 2.0, n_z)
# Get linear power spectrum
# Set up CAMB
p = camb.CAMBparams(WantTransfer=True,
WantCls=False,
Want_CMB_lensing=False,
DoLensing=False,
NonLinearModel=camb.nonlinear.Halofit(halofit_version="mead"))
p.set_cosmology(H0=h*100, omch2=omc*h**2, ombh2=omb*h**2, mnu=mnu)
p.set_dark_energy(w=w, wa=wa)
p.set_initial_power(camb.InitialPowerLaw(As=As, ns=ns))
p.set_matter_power(redshifts=z_lin, kmax=k_max, nonlinear=True)
# Compute CAMB results
r = camb.get_results(p)
k_lin, z_lin, pofk_lin_camb = r.get_linear_matter_power_spectrum(nonlinear=False)
Pk_nl_CAMB_interpolator = r.get_matter_power_interpolator()
pofk_nonlin_camb = Pk_nl_CAMB_interpolator.P(z_lin, k_lin, grid=True)
sigma8 = r.get_sigma8()[-1]
omv = r.omega_de + r.get_Omega("photon") + r.get_Omega("neutrino")
omm = p.omegam
# Setup HMCode
c = pyhmcode.Cosmology()
c.om_m = omm
c.om_b = omb
c.om_v = omv
c.h = h
c.ns = ns
c.sig8 = sigma8
c.m_nu = mnu
c.set_linear_power_spectrum(k_lin, z_lin, pofk_lin_camb)
pofk_hmc = {"logT" : [], "A" : []}
hmod = pyhmcode.Halomodel(pyhmcode.HMcode2016, verbose=False)
A = np.linspace(2.0, 3.13, 7)
print("Varying halo A")
for i, As in enumerate(A):
hmod.As = As
#hmod.eta0 = 0.98 - 0.12*As
pofk_hmc["A"].append(pyhmcode.calculate_nonlinear_power_spectrum(c, hmod, verbose=False)[0])
print("Varying TAGN")
hmod = pyhmcode.Halomodel(pyhmcode.HMcode2020_feedback, verbose=False)
logTAGN = np.linspace(7.3, 8.3, 7)
for i, logT in enumerate(logTAGN):
c.theat = 10**logT
pofk_hmc["logT"].append(pyhmcode.calculate_nonlinear_power_spectrum(c, hmod, verbose=True)[0])
if plot:
import matplotlib.pyplot as plt
import matplotlib.colorbar
cmap = plt.get_cmap("magma_r")
fig, ax = plt.subplots(2, 1, figsize=(5, 4))
fig.subplots_adjust(left=0.2, hspace=0.3, right=0.95, bottom=0.1)
for param_name, ls, param in [("logT", "-", logTAGN), ("A", "--", A)]:
cb_ax = matplotlib.colorbar.make_axes(ax)
norm = matplotlib.colors.Normalize(vmin=param[0], vmax=param[-1])
cb1 = matplotlib.colorbar.ColorbarBase(cb_ax[0], cmap=cmap,
norm=norm, **cb_ax[1])
cb1.set_label(param_name)
_ = [ax[0].loglog(k_lin, pofk_hmc[param_name][i],
c=cmap(i/len(param))) for i in range(len(param))]
_ = ax[0].loglog(k_lin, pofk_nonlin_camb[0], ls="--", c="k")
ax[0].set_title(f"Non-linear power spectrum, vary {param_name}")
ax[0].set_xlabel("k [h/Mpc]")
ax[0].set_ylabel("P(k) [Mpc^3 h^-3]")
_ = [ax[1].semilogx(k_lin, pofk_hmc[param_name][i]/pofk_nonlin_camb[0] - 1,
ls=ls, c=cmap(i/len(param))) for i in range(len(param))]
ax[1].set_xlabel("k [h/Mpc]")
ax[1].set_ylabel("HMCode/CAMB-1")
fig.dpi = 300
fig.savefig(f"plots/vary_baryon.png")
plt.show()
if __name__ == "__main__":
massfunction_params(plot=True)
# -
| notebooks/vary_baryon_parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load the modules
import cupy as cp
import numpy as np
import cudf
import dask_cudf
import cugraph
import cuspatial
import datetime as dttm
from cuml.preprocessing import (LabelEncoder, OneHotEncoder, train_test_split)
from dask_cuda import LocalCUDACluster
from dask.distributed import Client
from blazingsql import BlazingContext
from os import listdir
# +
cluster = LocalCUDACluster()
client = Client(cluster)
bc = BlazingContext(dask_client=client)
# -
# # Create the tables
# +
transactions_path = '../../data/seattle_parking/parking_MayJun2019.parquet'
transactions_parq = [f'{transactions_path}/{d}' for d in listdir(transactions_path) if d != '.ipynb_checkpoints']
locations_path = '../../data/seattle_parking/parking_locations.parquet'
locations_parq = [f'{locations_path}/{d}' for d in listdir(locations_path) if d != '.ipynb_checkpoints']
# -
bc.create_table('parking_transactions', transactions_parq)
bc.create_table('parking_locations', locations_parq)
# # Featurize parking locations
# +
parking_locations = bc.sql('SELECT * FROM parking_locations').compute()
parking_locations['ParkingArea_concat'] = (
parking_locations['PaidParkingArea']
.str.replace(' ', '_')
.str.cat(parking_locations['PaidParkingSubArea'], sep='__')
)
parking_locations['ParkingArea_concat'].head()
# +
le = LabelEncoder()
ohe = OneHotEncoder(sparse=False, output_type='cudf')
encoded = le.fit_transform(parking_locations['ParkingArea_concat']).to_frame()
encoded = ohe.fit_transform(encoded)
encoded_col_names = [f'parkingArea_cat_{i}' for i in range(encoded.shape[1])]
encoded = cudf.DataFrame(encoded, columns=encoded_col_names)
for col in encoded_col_names:
encoded[col] = encoded[col].astype('float32')
parking_locations = parking_locations.join(encoded)
# -
parking_locations = dask_cudf.from_cudf(parking_locations, npartitions=8)
bc.create_table('parking_locations', parking_locations)
# # Featurize parking transactions
# +
parking_transactions = (
bc.sql('''
SELECT A.*
, TIMESTAMPADD(HOUR, -1, A.OccupancyDateTime) AS time_prior_1h
, TIMESTAMPADD(DAY, -1, A.OccupancyDateTime) AS time_prior_1d
, TIMESTAMPADD(DAY, -7, A.OccupancyDateTime) AS time_prior_7d
FROM parking_transactions AS A
WHERE OccupancyDateTime >= '2019-06-01'
''')
)
bc.create_table('parking_transactions', parking_transactions)
# +
parking_transactions_agg = bc.sql('''
SELECT SourceElementKey
, transaction_year
, transaction_month
, transaction_day
, transaction_hour
, AVG(CAST(PaidOccupancy AS FLOAT)) AS average_occupancy
FROM (
SELECT A.*
, YEAR(OccupancyDateTime) AS transaction_year
, MONTH(OccupancyDateTime) AS transaction_month
, DAYOFMONTH(OccupancyDateTime) AS transaction_day
, HOUR(OccupancyDateTime) AS transaction_hour
FROM parking_transactions AS A
) AS outer_query
GROUP BY SourceElementKey
, transaction_year
, transaction_month
, transaction_day
, transaction_hour
''')
bc.create_table('parking_transactions_agg', parking_transactions_agg)
# -
dataset_for_training = bc.sql('''
WITH temp_query_prior_1h AS (
SELECT A.SourceElementKey
, A.PaidOccupancy AS Label
, A.OccupancyDateTime
, B.average_occupancy as AvgOccupancy_prior_1h
FROM (
SELECT SourceElementKey
, PaidOccupancy
, OccupancyDateTime
, YEAR(time_prior_1h) AS transaction_year
, MONTH(time_prior_1h) AS transaction_month
, DAYOFMONTH(time_prior_1h) AS transaction_day
, HOUR(time_prior_1h) AS transaction_hour
FROM parking_transactions) AS A
LEFT OUTER JOIN parking_transactions_agg AS B
ON A.SourceElementKey = B.SourceElementKey
AND A.transaction_year = B.transaction_year
AND A.transaction_month = B.transaction_month
AND A.transaction_day = B.transaction_day
AND A.transaction_hour = B.transaction_hour
)
, temp_query_prior_1d AS (
SELECT A.SourceElementKey
, A.PaidOccupancy AS Label
, A.OccupancyDateTime
, B.average_occupancy as AvgOccupancy_prior_1d
FROM (
SELECT SourceElementKey
, PaidOccupancy
, OccupancyDateTime
, YEAR(time_prior_1d) AS transaction_year
, MONTH(time_prior_1d) AS transaction_month
, DAYOFMONTH(time_prior_1d) AS transaction_day
, HOUR(time_prior_1d) AS transaction_hour
FROM parking_transactions) AS A
LEFT OUTER JOIN parking_transactions_agg AS B
ON A.SourceElementKey = B.SourceElementKey
AND A.transaction_year = B.transaction_year
AND A.transaction_month = B.transaction_month
AND A.transaction_day = B.transaction_day
AND A.transaction_hour = B.transaction_hour
)
, temp_query_prior_7d AS (
SELECT A.SourceElementKey
, A.PaidOccupancy AS Label
, A.OccupancyDateTime
, B.average_occupancy as AvgOccupancy_prior_7d
FROM (
SELECT SourceElementKey
, PaidOccupancy
, OccupancyDateTime
, YEAR(time_prior_7d) AS transaction_year
, MONTH(time_prior_7d) AS transaction_month
, DAYOFMONTH(time_prior_7d) AS transaction_day
, HOUR(time_prior_7d) AS transaction_hour
FROM parking_transactions) AS A
LEFT OUTER JOIN parking_transactions_agg AS B
ON A.SourceElementKey = B.SourceElementKey
AND A.transaction_year = B.transaction_year
AND A.transaction_month = B.transaction_month
AND A.transaction_day = B.transaction_day
AND A.transaction_hour = B.transaction_hour
)
SELECT hr_1.*
, d_1.AvgOccupancy_prior_1d
, d_7.AvgOccupancy_prior_7d
FROM temp_query_prior_1h AS hr_1
LEFT OUTER JOIN temp_query_prior_1d AS d_1
ON hr_1.SourceElementKey = d_1.SourceElementKey
AND hr_1.Label = d_1.Label
AND hr_1.OccupancyDateTime = d_1.OccupancyDateTime
LEFT OUTER JOIN temp_query_prior_7d AS d_7
ON hr_1.SourceElementKey = d_7.SourceElementKey
AND hr_1.Label = d_7.Label
AND hr_1.OccupancyDateTime = d_7.OccupancyDateTime
''').repartition(npartitions=8)
dataset_for_training.head()
dataset_for_training = (
dataset_for_training[[
'SourceElementKey'
, 'Label'
, 'AvgOccupancy_prior_1h'
, 'AvgOccupancy_prior_1d'
, 'AvgOccupancy_prior_7d'
]]
.merge(parking_locations[['SourceElementKey'] + encoded_col_names], on=['SourceElementKey'])
.dropna()
.drop(columns=['SourceElementKey'])
)
dataset_for_training.head()
# +
def random_column(df):
frame_len = len(df)
df['random'] = cp.random.rand(frame_len)
return df
dataset_for_training['Label'] = dataset_for_training['Label'].astype('float32')
dataset_for_training = dataset_for_training.map_partitions(random_column).compute()
train_X = dataset_for_training.query('random < 0.7')[[
'AvgOccupancy_prior_1h'
, 'AvgOccupancy_prior_1d'
, 'AvgOccupancy_prior_7d'
] + encoded_col_names]
train_y = dataset_for_training.query('random < 0.7')['Label']
test_X = dataset_for_training.query('random >= 0.7')[[
'AvgOccupancy_prior_1h'
, 'AvgOccupancy_prior_1d'
, 'AvgOccupancy_prior_7d'
] + encoded_col_names]
test_y = dataset_for_training.query('random >= 0.7')['Label']
# -
print(f'Full size: {len(dataset_for_training):,}, size of training: {len(train_y):,}, size of testing: {len(test_y):,}')
train_X.head()
from cuml import RandomForestRegressor
from cuml.metrics.regression import r2_score
rfr = RandomForestRegressor(
n_estimators=10
, max_depth=20
, min_rows_per_node=50
, verbose=2
)
rfr.fit(train_X, train_y)
rfr.predict(test_X)
r2_score(test_y, rfr.predict(test_X))
# # Featurize parking locations
# +
def extractLon(location):
lon = location.str.extract('([0-9\.\-]+) ([0-9\.]+)')[0]
return lon
def extractLat(location):
lon = location.str.extract('([0-9\.\-]+) ([0-9\.]+)')[1]
return lon
parking_locations['longitude'] = extractLon(parking_locations['Location']).astype('float32')
parking_locations['latitude'] = extractLat(parking_locations['Location']).astype('float32')
parking_locations[['Location', 'longitude', 'latitude']].head()
# -
# ## As crow flies vs as people walk
# 
# ### Read in the graph data
# Thanks to <NAME> for analyzing the map of King County roads and producing the data we will now use.
# #### Download and unpack the data
# +
import os
directory = os.path.exists('../../data')
archive = os.path.exists('../../data/king_county_road_graph_20190909.tar.gz')
file_graph = os.path.exists('../../data/king_county_road_graph_20190909.csv')
file_nodes = os.path.exists('../../data/king_county_road_nodes_20190909.csv')
if not directory:
os.mkdir('../data')
if not archive:
import wget, shutil
wget.download('http://tomdrabas.com/data/seattle_parking/king_county_road_graph_20190909.tar.gz')
shutil.move('king_county_road_graph_20190909.tar.gz', '../../data/king_county_road_graph_20190909.tar.gz')
if not file_graph or not file_nodes:
import tarfile
tf = tarfile.open('../../data/king_county_road_graph_20190909.tar.gz')
tf.extractall(path='../../data/')
# -
# #### Let's read the King County road data
road_graph_data = cudf.read_csv('../../data/seattle_parking/king_county_road_graph_20190909.csv')
road_graph_data['node1'] = road_graph_data['node1'].astype('int32')
road_graph_data['node2'] = road_graph_data['node2'].astype('int32')
road_graph_data['LENGTH'] = road_graph_data['LENGTH'] * 3 # convert to feet as the LENGHT was given in yards
road_nodes = cudf.read_csv('../../data/seattle_parking/king_county_road_nodes_20190909.csv')
road_nodes['NodeID'] = road_nodes['NodeID'].astype('int32')
# Store the maximum of the `NodeID` so we can later append the additional nodes that will be perpendicular to the actual parking locations. We also specify the offset - this will be used to append parking nodes.
offset = 100000
nodeId = road_nodes['NodeID'].max() ## so we can number the parking nodes properly (since we'll be adding a perpendicular projections)
parking_nodes_idx = road_nodes['NodeID'].max() + offset ## retain it so we can later filter the results to only parking locations
nodeId
# Move all the parking locations to host (via `.to_pandas()` method) so we can loop through all the ~1500 parking locations. Here, we also create an empty DataFrame that will hold the parking location nodes.
locations = parking_locations.compute().to_pandas().to_dict('records')
parking_locations_nodes = cudf.DataFrame(columns=['NodeID', 'Lon', 'Lat', 'SourceElementKey'])
added_location_edges = cudf.DataFrame(columns=['node1', 'node2', 'LENGTH'])
# Let's process the parking data. The kernel below finds equations of two lines:
#
# 1. Line that goes through road intersections
# 2. Line that is perpendicular to (1) and goes through the parking location.
#
# 
#
# Ultimately, we are finind the intersection of these two lines -- we call it the `PROJ` point below.
def kernel_find_projection(Lon_x, Lat_x, Lon_y, Lat_y, Lon_PROJ, Lat_PROJ, Lon_REF, Lat_REF):
for i, (lon_x, lat_x, lon_y, lat_y) in enumerate(zip(Lon_x, Lat_x, Lon_y, Lat_y)):
# special case where A and B have the same LON i.e. vertical line
if lon_x == lon_y:
Lon_PROJ[i] = lon_x
Lat_PROJ[i] = Lat_REF
else:
# find slope
a_xy = (lat_x - lat_y) / float(lon_x - lon_y)
# special case where A and B have the same LAT i.e. horizontal line
if a_xy == 0:
Lon_PROJ[i] = Lon_REF
Lat_PROJ[i] = lat_x
else:
# if neither of the above special cases apply
# find the equation of the perpendicular line
a_R = -1 / a_xy ### SLOPE
# find intersections
b_xy = lat_x - a_xy * lon_x
b_R = Lat_REF - a_R * Lon_REF
# find the coordinates
Lon_PROJ[i] = (b_R - b_xy) / (a_xy - a_R)
Lat_PROJ[i] = a_R * Lon_PROJ[i] + b_R
# +
# %%time
parking_locations_cnt = len(locations)
print('Number of parking locations: {0:,}'.format(parking_locations_cnt))
for i, loc in enumerate(locations):
if i % 100 == 0:
print('Processed: {0:,} ({1:.2%}) nodes'.format(i, i/float(parking_locations_cnt)))
#### INCREASE THE COUNTER AND GET THE REFERENCE POINT
nodeId = nodeId + 1
lat_r = loc['latitude']
lon_r = loc['longitude']
#### APPEND GEO COORDINATES TO INTERSECTION AND SUBSET DOWN THE DATASET
#### TO POINTS WITHIN ~2000ft FROM PARKING SPOT
paths = (
road_graph_data
.rename(columns={'node1': 'NodeID'})
.merge(road_nodes[['NodeID', 'Lat', 'Lon']], on='NodeID', how='left')
.rename(columns={'NodeID': 'node1', 'node2': 'NodeID'})
.merge(road_nodes[['NodeID', 'Lat', 'Lon']], on='NodeID', how='left')
.rename(columns={'NodeID': 'node2'})
.query('Lat_x >= (@lat_r - 0.005) and Lat_x <= (@lat_r + 0.005)')
.query('Lon_x >= (@lon_r - 0.005) and Lon_x <= (@lon_r + 0.005)')
.query('Lat_y >= (@lat_r - 0.005) and Lat_y <= (@lat_r + 0.005)')
.query('Lon_y >= (@lon_r - 0.005) and Lon_y <= (@lon_r + 0.005)')
)
#### APPEND THE PARKING LOCATION SO WE CAN CALCULATE DISTANCES
paths['Lon_REF'] = loc['longitude']
paths['Lat_REF'] = loc['latitude']
paths = paths.apply_rows(
kernel_find_projection
, incols = ['Lon_x', 'Lat_x', 'Lon_y', 'Lat_y', 'Lon_REF', 'Lat_REF']
, outcols = {'Lon_PROJ': np.float64, 'Lat_PROJ': np.float64}
, kwargs = {'Lon_REF': loc['longitude'], 'Lat_REF': loc['latitude']}
)
#### CALCULATE THE DISTANCES SO WE CAN CHECK IF THE PROJ POINT IS BETWEEN ROAD NODES
paths['Length_x_PROJ'] = cuspatial.haversine_distance(
paths['Lon_x']
, paths['Lat_x']
, paths['Lon_PROJ']
, paths['Lat_PROJ'])# * 0.621371 * 5280
paths['Length_x_PROJ'] = paths['Length_x_PROJ'] * 0.621371 * 5280
paths['Length_y_PROJ'] = cuspatial.haversine_distance(
paths['Lon_y']
, paths['Lat_y']
, paths['Lon_PROJ']
, paths['Lat_PROJ'])
paths['Length_y_PROJ'] = paths['Length_y_PROJ'] * 0.621371 * 5280
paths['Length_REF_PROJ'] = cuspatial.haversine_distance(
paths['Lon_REF']
, paths['Lat_REF']
, paths['Lon_PROJ']
, paths['Lat_PROJ'])
paths['Length_REF_PROJ'] = paths['Length_REF_PROJ'] * 0.621371 * 5280
#### SELECT THE POINTS THAT A LESS THAN OR EQAL TO TOTAL LENGTH OF THE EDGE (WITHIN 1 ft)
paths['PROJ_between'] = (paths['Length_x_PROJ'] + paths['Length_y_PROJ']) <= (paths['LENGTH'] + 1)
#### SELECT THE CLOSEST
closest = (
paths
.query('PROJ_between')
.nsmallest(1, 'Length_REF_PROJ')
.to_pandas()
.to_dict('records')[0]
)
# add nodes
nodes = cudf.DataFrame({
'NodeID': [nodeId + offset, nodeId]
, 'Lon': [closest['Lon_REF'], closest['Lon_PROJ']]
, 'Lat': [closest['Lat_REF'], closest['Lat_PROJ']]
, 'SourceElementKey': [loc['SourceElementKey'], None]
})
parking_locations_nodes = cudf.concat([parking_locations_nodes, nodes])
# add edges (bi-directional)
edges = cudf.DataFrame({
'node1': [nodeId, nodeId, nodeId, closest['node1'], closest['node2'], nodeId + offset]
, 'node2': [closest['node1'], closest['node2'], nodeId + offset, nodeId, nodeId, nodeId]
, 'LENGTH': [
closest['Length_x_PROJ'], closest['Length_y_PROJ'], closest['Length_REF_PROJ']
, closest['Length_x_PROJ'], closest['Length_y_PROJ'], closest['Length_REF_PROJ']
]
})
added_location_edges = cudf.concat([added_location_edges, edges]) ## append to the temp DataFrame
print('Finished processing...')
# -
road_graph_data.head()
road_nodes = (
cudf
.concat([road_nodes[['NodeID', 'Lon', 'Lat']], parking_locations_nodes])
.reset_index(drop=True)
)
# Now we can find the nearest intersections from the Space Needle!
location = {'latitude': 47.620422, 'longitude': -122.349358}
date = dttm.datetime.strptime('2019-06-21 13:21:00', '%Y-%m-%d %H:%M:%S')
bc.create_table('parking_transactions'
, bc.sql(f'''
SELECT *
, YEAR(OccupancyDateTime) AS transaction_year
, MONTH(OccupancyDateTime) AS transaction_month
, DAYOFMONTH(OccupancyDateTime) AS transaction_day
, HOUR(OccupancyDateTime) AS transaction_hour
FROM parking_transactions
''')
)
# +
road_nodes['Lon_REF'] = location['longitude']
road_nodes['Lat_REF'] = location['latitude']
road_nodes['Distance'] = cuspatial.haversine_distance(
road_nodes['Lon']
, road_nodes['Lat']
, road_nodes['Lon_REF']
, road_nodes['Lat_REF'])
road_nodes['Distance'] = road_nodes['Distance'] * 0.621371 * 5280
space_needle_to_nearest_intersection = road_nodes.nsmallest(5, 'Distance') ### Space Needle is surrounded by around 5 road intersections hence we add 5
space_needle_to_nearest_intersection_dist = space_needle_to_nearest_intersection['Distance'].to_array()[0]
space_needle_to_nearest_intersection['node1'] = nodeId + 2
space_needle_to_nearest_intersection = (
space_needle_to_nearest_intersection
.rename(columns={'NodeID': 'node2', 'Distance': 'LENGTH'})
[['node1', 'node2', 'LENGTH']]
)
road_graph_data = cudf.concat([space_needle_to_nearest_intersection, added_location_edges, road_graph_data])
space_needle_to_nearest_intersection ### SHOW THE EDGES
# -
# ### The road graph
road_graph_data = road_graph_data.reset_index(drop=True)
road_graph_data['node1'] = road_graph_data['node1'].astype('int32')
road_graph_data['node2'] = road_graph_data['node2'].astype('int32')
g = cugraph.Graph()
g.from_cudf_edgelist(
road_graph_data
, source='node1'
, destination='node2'
, edge_attr='LENGTH'
, renumber=False
)
# Now we can use the `.sssp(...)` method from `cugraph` to find the shortest distances to parking spots from the Space Needle!
all_distances = cugraph.sssp(g, nodeId + 2)
distances = all_distances.query('vertex > @parking_nodes_idx and distance < 1000')
distances
# #### INFERENCE
inference = distances.merge(parking_locations_nodes, left_on='vertex', right_on='NodeID')
inference['OccupancyDateTime'] = date
inference['time_prior_1h'] = date + dttm.timedelta(hours=-1)
inference['time_prior_1d'] = date + dttm.timedelta(days=-1)
inference['time_prior_7d'] = date + dttm.timedelta(days=-7)
bc.create_table('inference', inference)
inference
inference = bc.sql(f'''
WITH temp_query_prior_1h AS (
SELECT A.SourceElementKey
, A.OccupancyDateTime
, B.average_occupancy as AvgOccupancy_prior_1h
FROM (
SELECT SourceElementKey
, OccupancyDateTime
, YEAR(time_prior_1h) AS transaction_year
, MONTH(time_prior_1h) AS transaction_month
, DAYOFMONTH(time_prior_1h) AS transaction_day
, HOUR(time_prior_1h) AS transaction_hour
FROM inference) AS A
LEFT OUTER JOIN parking_transactions_agg AS B
ON A.SourceElementKey = B.SourceElementKey
AND A.transaction_year = B.transaction_year
AND A.transaction_month = B.transaction_month
AND A.transaction_day = B.transaction_day
AND A.transaction_hour = B.transaction_hour
)
, temp_query_prior_1d AS (
SELECT A.SourceElementKey
, A.OccupancyDateTime
, B.average_occupancy as AvgOccupancy_prior_1d
FROM (
SELECT SourceElementKey
, OccupancyDateTime
, YEAR(time_prior_1d) AS transaction_year
, MONTH(time_prior_1d) AS transaction_month
, DAYOFMONTH(time_prior_1d) AS transaction_day
, HOUR(time_prior_1d) AS transaction_hour
FROM inference) AS A
LEFT OUTER JOIN parking_transactions_agg AS B
ON A.SourceElementKey = B.SourceElementKey
AND A.transaction_year = B.transaction_year
AND A.transaction_month = B.transaction_month
AND A.transaction_day = B.transaction_day
AND A.transaction_hour = B.transaction_hour
)
, temp_query_prior_7d AS (
SELECT A.SourceElementKey
, A.OccupancyDateTime
, B.average_occupancy as AvgOccupancy_prior_7d
FROM (
SELECT SourceElementKey
, OccupancyDateTime
, YEAR(time_prior_7d) AS transaction_year
, MONTH(time_prior_7d) AS transaction_month
, DAYOFMONTH(time_prior_7d) AS transaction_day
, HOUR(time_prior_7d) AS transaction_hour
FROM inference) AS A
LEFT OUTER JOIN parking_transactions_agg AS B
ON A.SourceElementKey = B.SourceElementKey
AND A.transaction_year = B.transaction_year
AND A.transaction_month = B.transaction_month
AND A.transaction_day = B.transaction_day
AND A.transaction_hour = B.transaction_hour
)
SELECT hr_1.*
, d_1.AvgOccupancy_prior_1d
, d_7.AvgOccupancy_prior_7d
FROM temp_query_prior_1h AS hr_1
LEFT OUTER JOIN temp_query_prior_1d AS d_1
ON hr_1.SourceElementKey = d_1.SourceElementKey
AND hr_1.OccupancyDateTime = d_1.OccupancyDateTime
LEFT OUTER JOIN temp_query_prior_7d AS d_7
ON hr_1.SourceElementKey = d_7.SourceElementKey
AND hr_1.OccupancyDateTime = d_7.OccupancyDateTime
ORDER BY SourceElementKey
''').compute()
inference_X = (
inference[[
'SourceElementKey'
, 'AvgOccupancy_prior_1h'
, 'AvgOccupancy_prior_1d'
, 'AvgOccupancy_prior_7d'
]]
.merge(parking_locations[['SourceElementKey'] + encoded_col_names].compute(), on=['SourceElementKey'])
.sort_values(by='SourceElementKey')
.dropna()
.drop(columns=['SourceElementKey'])
)
inference_X
inference.join(rfr.predict(inference_X).to_frame('prediction'))
# `cugraph` returns a DataFrame with vertex, distance to that vertex, and the total distance traveled to that vertex from the `nodeId + 1` node -- the Space Needle. Here, we unfold the full path.
# +
# unfold -- create the whole path
closest_node = nodeId + 2
parking_cnt = distances['vertex'].count()
for i in range(parking_cnt):
print('Processing record: {0}'.format(i))
parking_node = distances.iloc[i]
vertex = int(parking_node['vertex'])
predecessor = int(parking_node['predecessor'])
if i == 0:
paths = all_distances.query('vertex == @vertex')
else:
paths = cudf.concat([all_distances.query('vertex == @vertex'), paths])
while vertex != closest_node:
temp = all_distances.query('vertex == @predecessor')
paths = cudf.concat([temp, paths])
predecessor = temp['predecessor'].to_array()[0]
vertex = temp['vertex'].to_array()[0]
# -
paths
| codes/parking/blazing_seattleParking_old.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Problem Statement:
# Predict the survival of a horse based on various observed medical conditions. Load
# the data from “horses.csv” and observe whether it contains missing values. The dataset contains many
# categorical features; replace them with label encoding. Replace the missing values by the most frequent value
# in each column. Fit a decision tree classifier and random forest classifier, and observe the accuracy.
# ### Objective:
# Learn to fit a decision tree, and compare its accuracy with random forest classifier.
# +
import pandas as pd
import matplotlib.pyplot as plot
# %matplotlib inline
# -
animals = pd.read_csv('C:/data/horse.csv')
animals.info()
animals.shape
animals.head()
animals.describe()
animals.isnull().sum()
animals.columns
target = animals['outcome']
target.unique()
animals = animals.drop(['outcome'],axis=1)
# +
#Select all categorical variables and create dummies
category_variables=['surgery', 'age', 'temp_of_extremities', 'peripheral_pulse',
'mucous_membrane', 'capillary_refill_time', 'pain', 'peristalsis',
'abdominal_distention', 'nasogastric_tube', 'nasogastric_reflux',
'rectal_exam_feces', 'abdomen','abdomo_appearance','surgical_lesion',
'cp_data']
for category in category_variables:
animals[category] = pd.get_dummies(animals[category])
# +
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
x,y = animals.values, target.values
label_encoder = LabelEncoder()
y = label_encoder.fit_transform(y)
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size= 0.2,random_state=1)
# -
from sklearn.tree import DecisionTreeClassifier
print(x_train.shape)
# +
#handling missing value
#from sklearn.preprocessing import Imputer
from sklearn.impute import SimpleImputer
import numpy as np
#imp = Imputer(missing_values = 'NaN', strategy = "most_frequent",axis = 0)
imp = SimpleImputer(missing_values = np.nan, strategy = "most_frequent")
x_train = imp.fit_transform(x_train)
x_test = imp.fit_transform(x_test)
# -
classifier = DecisionTreeClassifier()
classifier.fit(x_train,y_train)
y_predict = classifier.predict(x_test)
from sklearn.metrics import accuracy_score
accuracy = accuracy_score(y_predict,y_test)
print(accuracy)
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier()
classifier.fit(x_train,y_train)
y_predict = classifier.predict(x_test)
accuracy = accuracy_score(y_predict,y_test)
print(accuracy)
| Machine Learning/13_Horse_Survival_Analysis_Decision_Tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Spark MLlib
# see: http://spark.apache.org/mllib/
#
# compare: https://github.com/krishnaik06/Pyspark-With-Python/blob/main/Tutorial%206-Example%20Of%20Pyspark%20ML.ipynb
from pyspark.sql import SparkSession
# + tags=[]
spark=SparkSession.builder.appName('MLlib').getOrCreate()
# -
df=spark.read.csv('test1.csv',header=True,inferSchema=True) #inferSchema --> age type turns from string to integer
df.show()
df.printSchema()
from pyspark.ml.feature import VectorAssembler
featureassembler=VectorAssembler(inputCols=["age","Experience"],outputCol="Independent Features")
output=featureassembler.transform(df)
output.show()
output.columns
finalized_data=output.select("Independent Features","Salary")
finalized_data.show()
# +
from pyspark.ml.regression import LinearRegression
##train test split
train_data,test_data=finalized_data.randomSplit([0.75,0.25])
# -
regressor=LinearRegression(featuresCol='Independent Features', labelCol='Salary')
regressor=regressor.fit(train_data)
### Coefficients
regressor.coefficients
### Intercepts
regressor.intercept
### Prediction
pred_results=regressor.evaluate(test_data)
pred_results.predictions.show()
pred_results.meanAbsoluteError,pred_results.meanSquaredError
| 5-MLlib Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sprintenv
# language: python
# name: sprintenv
# ---
# +
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from sklearn.utils import shuffle
from sklearn.utils import check_random_state
from sklearn.cluster import KMeans
from sklearn.preprocessing import normalize
from sklearn.metrics import pairwise_distances
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.decomposition import NMF, LatentDirichletAllocation
from sklearn.datasets import make_blobs
from scipy.sparse import csr_matrix
import pandas as pd
import numpy as np
import sys
import os
# %matplotlib inline
# -
# # Running kmeans in sci-kit learn
# +
n_samples = 1500
random_state = 170
X, y = make_blobs(n_samples=n_samples, random_state=random_state)
plt.scatter(X[:, 0], X[:, 1])
plt.title("Create random blobs")
# +
# Incorrect number of clusters
y_pred = KMeans(n_clusters=2, random_state=random_state).fit_predict(X)
plt.scatter(X[:, 0], X[:, 1], c=y_pred)
plt.title("Incorrect Number of Blobs")
# +
# Incorrect number of clusters
y_pred = KMeans(n_clusters=3, random_state=random_state).fit_predict(X)
plt.scatter(X[:, 0], X[:, 1], c=y_pred)
plt.title("correct Number of Blobs")
# -
# # Task1: Understanding initialization impact on convergence
# +
random_state = np.random.RandomState(0)
# Datasets generation parameters
n_samples_per_center = 100
grid_size = 3
scale = 0.1
n_clusters = grid_size ** 2
def make_data(random_state, n_samples_per_center, grid_size, scale):
random_state = check_random_state(random_state)
centers = np.array([[i, j]
for i in range(grid_size)
for j in range(grid_size)])
n_clusters_true, n_features = centers.shape
noise = random_state.normal(
scale=scale, size=(n_samples_per_center, centers.shape[1]))
X = np.concatenate([c + noise for c in centers])
y = np.concatenate([[i] * n_samples_per_center
for i in range(n_clusters_true)])
return shuffle(X, y, random_state=random_state)
# -
# ## Kmeans initialisation effect
# +
X, y = make_data(random_state, n_samples_per_center, grid_size, scale)
km = KMeans(n_clusters=n_clusters, init='random', n_init=1,
random_state=random_state).fit(X)
plt.figure()
for k in range(n_clusters):
my_members = km.labels_ == k
color = cm.nipy_spectral(float(k) / n_clusters, 1)
plt.plot(X[my_members, 0], X[my_members, 1], 'o', marker='.', c=color)
cluster_center = km.cluster_centers_[k]
plt.plot(cluster_center[0], cluster_center[1], 'o',
markerfacecolor=color, markeredgecolor='k', markersize=6)
plt.title("Example cluster allocation with a random init\n"
"with KMeans")
plt.show()
# +
X, y = make_data(random_state, n_samples_per_center, grid_size, scale)
km = KMeans(n_clusters=n_clusters, init='k-means++', n_init=1,
random_state=random_state).fit(X)
plt.figure()
for k in range(n_clusters):
my_members = km.labels_ == k
color = cm.nipy_spectral(float(k) / n_clusters, 1)
plt.plot(X[my_members, 0], X[my_members, 1], 'o', marker='.', c=color)
cluster_center = km.cluster_centers_[k]
plt.plot(cluster_center[0], cluster_center[1], 'o',
markerfacecolor=color, markeredgecolor='k', markersize=6)
plt.title("Example cluster allocation with a random init\n"
"with KMeans")
plt.show()
# -
# ## Inertia measure
# +
# Number of time the k-means algorithm will be run with different
# centroid seeds. The final results will be the best output of n_init consecutive runs in terms of inertia.
n_init_range = np.array([1, 5, 10, 15, 20])
# Number of run (with randomly generated dataset) for each strategy so as
# to be able to compute an estimate of the standard deviation
n_runs = 10
plt.figure()
plots = []
legends = []
cases = [
(KMeans, 'k-means++', {}),
(KMeans, 'random', {}),
]
for factory, init, params in cases:
print("Evaluation of %s with %s init" % (factory.__name__, init))
inertia = np.empty((len(n_init_range), n_runs))
for run_id in range(n_runs):
X, y = make_data(run_id, n_samples_per_center, grid_size, scale)
for i, n_init in enumerate(n_init_range):
km = factory(n_clusters=n_clusters, init=init, random_state=run_id,
n_init=n_init, **params).fit(X)
inertia[i, run_id] = km.inertia_
p = plt.errorbar(n_init_range, inertia.mean(axis=1), inertia.std(axis=1))
plots.append(p[0])
legends.append("%s with %s init" % (factory.__name__, init))
plt.xlabel('n_init')
plt.ylabel('inertia')
plt.legend(plots, legends)
plt.title("Mean inertia for various k-means init across %d runs" % n_runs)
# -
# # Task 2: Use k-means to cluster wikipedia articles
# ## Compute TF-IDF
# +
# import text documents from wikipedia abstracts
wiki_data=pd.read_csv('../people_wiki.txt',delimiter='\t', index_col='name')['text']
wiki_data.head()
# -
#Define the TFIDF vectorizer that will be used to process the data
tfidf_vectorizer = TfidfVectorizer()
#Apply this vectorizer to the full dataset to create normalized vectors
tf_idf = tfidf_vectorizer.fit_transform(wiki_data)
# ## Choose the right number of k
def plot_k_vs_heterogeneity(k_values, heterogeneity_values):
plt.figure(figsize=(7,4))
plt.plot(k_values, heterogeneity_values, linewidth=4)
plt.xlabel('K')
plt.ylabel('Inertia')
plt.title('K vs. Inertia')
plt.rcParams.update({'font.size': 16})
plt.tight_layout()
heterogeneity_values = []
k_list = [2, 10, 25, 50, 100, 300]
##### TAKES FOREVER TO RUN#####
for n_clusters in k_list:
km = KMeans(n_clusters=n_clusters, init='k-means++').fit(tf_idf)
heterogeneity_values.append(km.inertia_)
plot_k_vs_heterogeneity(k_list, heterogeneity_values)
# compute kmeans using random initialization
kmeans = KMeans(n_clusters=100, init='random').fit(tf_idf)
# predict the closest cluster for each point
labels= kmeans.predict(tf_idf)
# retrieve other pages from a cluster
def get_cluster(cluster_nb, labels, dataset):
curr_cluster=[]
abstract=[]
for index in range(len(labels)):
if labels[index]==cluster_nb:
curr_cluster.append(dataset.index.values[index,])
abstract.append(dataset.iloc[index,])
result = pd.DataFrame({'names': curr_cluster,
'abstract': abstract})
return result
# Find the cluster in which Dr. Dre is
pos = wiki_data.index.get_loc("Dr. Dre")
labels[pos]
cluster= get_cluster(labels[pos], labels, wiki_data)
cluster
# try to get a smarter initialization?
# # LDA modeling
# +
# import text documents from wikipedia abstracts
wiki_data=pd.read_csv('../people_wiki.txt',delimiter='\t', index_col='name')['text']
wiki_data.head()
# LDA can only use raw term counts for LDA because it is a probabilistic graphical model
tf_vectorizer = CountVectorizer(max_df=0.95, min_df=2, stop_words='english')
tf = tf_vectorizer.fit_transform(wiki_data)
tf_feature_names = tf_vectorizer.get_feature_names()
# -
no_topics = 20
# Run LDA
lda = LatentDirichletAllocation(n_topics=no_topics).fit(tf)
# # Display and evaluate topics
# +
def display_topics(model, feature_names, no_top_words):
for topic_idx, topic in enumerate(model.components_):
print("Topic %d:" % (topic_idx))
print(" ".join([feature_names[i]
for i in topic.argsort()[:-no_top_words - 1:-1]]))
no_top_words = 10
display_topics(lda, tf_feature_names, no_top_words)
# -
# transform method returns a matrix with one line per document, columns being topics weight
predict = lda.transform(tf)
result = pd.DataFrame(predict).set_index(wiki_data.reset_index()['name'])
# +
# get distribution of topics for Juliette Binoche
d = pd.Series(result.loc["Juliette Binoche"]).sort_values(ascending=False)
ax = d.plot(kind='bar', title='<NAME> Wikipedia Topic distribution',
figsize=(10,6), width=.8, fontsize=14, rot=45 )
ax.title.set_size(20)
# -
# Adjust (less or more topics) and rerun the LDA model
#display best word by topic
# transform method returns a matrix with one line per document, columns being topics weight
# get distribution of topics for Zendaya
| sprint4_clustering/sprint4_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ZvoneST/pytorch-labs/blob/master/lovro_overfitting.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="ycCVB6FOEe__" outputId="abffa856-63c7-4bdb-cdca-479f6de6cc59"
from __future__ import print_function
import argparse
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
from torch.autograd import Variable
from torchvision import transforms as T
import matplotlib.pyplot as plt
from torchvision.datasets import ImageFolder
# Training settings
batch_size = 64
# MNIST Dataset
train_dataset = datasets.MNIST(root='./data/',
train=True,
transform=transforms.ToTensor(),
download=True)
data_transforms = transforms.Compose([
transforms.Grayscale(num_output_channels=1),transforms.ToTensor()])
test_dataset = ImageFolder('./data/zvone/', data_transforms)
# Data Loader (Input Pipeline)
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size=5)
#print(self.conv1.weight.shape)
self.conv2 = nn.Conv2d(10, 20, kernel_size=5)
self.conv3 = nn.Conv2d(20, 20, kernel_size=3)
#print(self.conv2.weight.shape)
self.mp = nn.MaxPool2d(2)
self.fc = nn.Linear(320, 10)
def forward(self, x):
in_size = x.size(0)
x = F.relu(self.conv1(x))
#print(x.shape)
x = F.relu(self.mp(self.conv2(x)))
x = F.relu(self.mp(self.conv3(x)))
#print("2.", x.shape)
# x = F.relu(self.mp(self.conv3(x)))
x = x.view(in_size, -1) # flatten the tensor
#print("3.", x.shape)
x = self.fc(x)
return F.log_softmax(x)
model = Net()
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.5)
def train(epoch):
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
data, target = Variable(data), Variable(target)
optimizer.zero_grad()
output = model(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
if batch_idx % 10 == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
def test():
model.eval()
test_loss = 0
correct = 0
for data, target in test_loader:
data, target = Variable(data, volatile=True), Variable(target)
output = model(data)
# print(target, output)
# sum up batch loss
test_loss += F.nll_loss(output, target, size_average=False).data
# get the index of the max log-probability
pred = output.data.max(1, keepdim=True)[1]
correct += pred.eq(target.data.view_as(pred)).cpu().sum()
test_loss /= len(test_loader.dataset)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
for epoch in range(1, 2):
train(epoch)
test()
model_scripted = torch.jit.script(model) # Export to TorchScript
model_scripted.save('model.pt') # Save
# + colab={"base_uri": "https://localhost:8080/"} id="qtSXATmGe03a" outputId="7b2d0737-bb1b-4f9b-c6bc-b06a341410a0"
test_loader.dataset
# + id="poIUBCNOFzhu"
# !rm -rf `find -type d -name .ipynb_checkpoints`
# + colab={"base_uri": "https://localhost:8080/"} id="QIWnWe89xgSw" outputId="5ec9fbbe-26b4-4ae9-bf69-e0f3160ab681"
imgs = torch.stack([img_t for img_t, _ in test_dataset], dim=1)
imgs.view(1, -1).mean(dim=1)
# + colab={"base_uri": "https://localhost:8080/"} id="j7LMaIUOzWJ7" outputId="e54f9a9d-7c73-4d1b-83f6-c2abbdab1c22"
imgs.view(1, -1).std(dim=1)
# + colab={"base_uri": "https://localhost:8080/"} id="IZ1otSmu7Bzx" outputId="61c9e66a-f34a-4498-cafc-66b3e4ebf217"
imgs = torch.stack([img_t for img_t, _ in train_dataset], dim=1)
imgs.view(1, -1).mean(dim=1)
# + colab={"base_uri": "https://localhost:8080/"} id="lt4JIIrc7GYo" outputId="5453bb33-5f5d-4808-b0b3-fa663fcb91e4"
imgs.view(1, -1).std(dim=1)
# + id="OkF5SVPL0JUo"
max_degree=30
train_dataset = datasets.MNIST(root='./data/',
train=True,
transform=transforms.Compose([
transforms.RandomAffine((0, 10)),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
# transforms.Normalize((0.1307,), (0.3081,))
]),
download=False)
data_transforms = transforms.Compose([
transforms.Grayscale(num_output_channels=1),transforms.ToTensor()])
#test_dataset = ImageFolder('lovro/', data_transforms)
# Data Loader (Input Pipeline)
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="eEg4cKjGwz61" outputId="37c7fc1b-77af-4d3f-c477-ce263d002e32"
imgs, lbls = next(iter(train_loader))
imgs[7].data.shape
plt.imshow(imgs[7].data.reshape((28,28)), cmap="gray")
# + colab={"base_uri": "https://localhost:8080/"} id="c1LKbLLg0ZuM" outputId="e32884df-d7fe-44c9-b85b-ca840cdc8ceb"
model = Net()
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.5)
for epoch in range(1, 2):
train(epoch)
test()
# + [markdown] id="JocDEp7719Xs"
# ## Data Augmentation / Image transformation
#
# A deep network requires extensive data to achieve decent performance. To build a good classifier with small training data, image augmentation can solve the problem to a greater extend. Image augmentation generates images by different ways of processing, such as random shift, rotation, flips, etc.
#
# Below are the list of transformations that come pre-built with PyTorch:
# - ToTensor
# - ToPILImage
# - Normalize
# - Resize
# - Scale
# - CenterCrop
# - Pad
# - Lambda
# - RandomApply
# - RandomChoice
# - RandomOrder
# - RandomCrop
# - RandomHorizontalFlip
# - RandomVerticalFlip
# - RandomResizedCrop
# - RandomSizedCrop
# - FiveCrop
# - TenCrop
# - LinearTransformation
# - ColorJitter
# - RandomRotation
# - RandomAffine
# - Grayscale
# - RandomGrayscale
# - RandomPerspective
# - RandomErasing
#
#
#
# + [markdown] id="354SdU9c1_tx"
# ##Ex 1
# Consider which of the following data augmentation are useful in increasing the correct algorithm on the drawn examples. Implement the selected functions and check that you have increased the accuracy of the algorithm.
# + id="PVJqIS0Y3L9H"
#your code
# + id="_EjMV_Bd-QhZ"
# + [markdown] id="j1YCVYI8_fku"
# # Dropout
# Yet another way to prevent overfitting is to build many models, then average their predictions at test time. Each model might have a different set of initial weights.
#
# We won't show an example of model averaging here. Instead, we will show another idea that sounds drastically different on the surface.
#
# This idea is called dropout: we will randomly "drop out", "zero out", or "remove" a portion of neurons from each training iteration.
#
#
#
# In different iterations of training, we will drop out a different set of neurons.
#
# The technique has an effect of preventing weights from being overly dependent on each other: for example for one weight to be unnecessarily large to compensate for another unnecessarily large weight with the opposite sign. Weights are encouraged to be "more independent" of one another.
#
# During test time though, we will not drop out any neurons; instead we will use the entire set of weights. This means that our training time and test time behaviour of dropout layers are different. In the code for the function train and get_accuracy, we use model.train() and model.eval() to flag whether we want the model's training behaviour, or test time behaviour.
#
# While unintuitive, using all connections is a form of model averaging! We are effectively averaging over many different networks of various connectivity structures.
# + id="qqYcLVJJ-ZiL"
# MNIST Dataset
train_dataset = datasets.MNIST(root='./data/',
train=True,
transform=transforms.ToTensor(),
download=True)
data_transforms = transforms.Compose([
transforms.Grayscale(num_output_channels=1),transforms.ToTensor()])
test_dataset = ImageFolder('lovro/', data_transforms)
# Data Loader (Input Pipeline)
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size=5)
nn.Dropout(0.2)
#print(self.conv1.weight.shape)
self.conv2 = nn.Conv2d(10, 20, kernel_size=5)
nn.Dropout(0.2)
self.conv3 = nn.Conv2d(20, 20, kernel_size=3)
#print(self.conv2.weight.shape)
nn.Dropout(0.2)
self.mp = nn.MaxPool2d(2)
self.fc = nn.Linear(320, 10)
nn.Dropout(0.2)
def forward(self, x):
in_size = x.size(0)
x = F.relu(self.conv1(x))
#print(x.shape)
x = F.relu(self.mp(self.conv2(x)))
x = F.relu(self.mp(self.conv3(x)))
#print("2.", x.shape)
# x = F.relu(self.mp(self.conv3(x)))
x = x.view(in_size, -1) # flatten the tensor
#print("3.", x.shape)
x = self.fc(x)
return F.log_softmax(x)
model = Net()
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.5)
# + colab={"base_uri": "https://localhost:8080/"} id="_KGIrURv-s0S" outputId="870ed114-0bd6-4732-a406-6867b036b981"
for epoch in range(1, 2):
train(epoch)
test()
# + [markdown] id="CgJSntNaAwJI"
# # Ex2
# Try to combine data augmentation and dropout idea and compare result of convolutional neural network.
# + id="OgSSpaVSBgax"
#your code
# + [markdown] id="lizmmwXCBiMN"
# # Weight Decay
# A more interesting technique that prevents overfitting is the idea of weight decay. The idea is to **penalize large weights**. We avoid large weights, because large weights mean that the prediction relies a lot on the content of one pixel, or on one unit. Intuitively, it does not make sense that the classification of an image should depend heavily on the content of one pixel, or even a few pixels.
#
# Mathematically, we penalize large weights by adding an extra term to the loss function, the term can look like the following:<br>
#
# * $L^1$ regularization $\sum_{k} |w_k|$<br>
# Mathematically, this term encourages weights to be exactly 0.<br><br>
# * $L^2$ regularization $\sum_{k} w_k^2$<br>
# Mathematically, in each iteration the weight is pushed towards 0.<br><br>
# * Combination of $L^1$ and $L^2$ regularization: add a term $\sum_{k} |w_k| + w_k^2$ to the loss function.
#
#
#
#
#
# In PyTorch, weight decay can also be done automatically inside an optimizer. The parameter weight_decay of optim.SGD and most other optimizers uses L2 regularization for weight decay. The value of the weight_decay parameter is another tunable hyperparameter.
# + id="avD6AFsRGFHV"
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.5, weight_decay=1e-5)
# + [markdown] id="MA9rzKInJ-wT"
# # Ex3
# Try which combination of methods (or maybe all) makes the best possible result.
# + id="1zjt2H6dKgFt"
#your code
| lovro_overfitting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Importing the Libraries
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
# Filtering the Warnings
warnings.filterwarnings("ignore")
print('Libraries Loaded Sucessfully...!')
# +
# Loading the Dataset
matches_dataset = pd.read_csv('Datasets/matches.csv')
# Printing the first 5 rows
matches_dataset.head()
# +
# Describing the Dataset
matches_dataset.describe()
# +
# Shape of the Dataset
matches_dataset.shape
# -
matches_dataset.tail()
# +
# Droping the some columns from the Dataset
matches_dataset.drop('umpire3', inplace=True, axis=1)
print('Column Dropped Sucessfully...!')
matches_dataset.head(8)
# +
player_of_match = dict()
for player in matches_dataset['player_of_match']:
player_of_match[player] = player_of_match.get(player, 0) + 1
print(player_of_match)
# +
# Printing the results of Man of the Match
for player,number in player_of_match.items():
if number == 1:
print('{} got {} time Man of the Match Award.'.format(player,number))
else:
print('{} got {} times Man of the Match Award.'.format(player,number))
# -
# ### Plotting the results of Man Of the Match Award in IPL 2008 - 2018
# +
player_names = list(player_of_match.keys())
number_of_times = list(player_of_match.values())
manOfTheMatch = dict()
for Name,value in player_of_match.items():
if value >= 6:
manOfTheMatch[Name] = value
# Plotting the Graph
plt.bar(range(len(manOfTheMatch)), manOfTheMatch.values())
plt.xticks(range(len(manOfTheMatch)), list(manOfTheMatch.keys()), rotation='vertical')
plt.xlabel('Player Name')
plt.ylabel('No Of Times')
plt.title('Man Of the Match Award')
plt.show()
# -
# ### Number Of Wins Of Each Team
# +
teamWinCounts = dict()
for team in matches_dataset['winner']:
if team == None:
continue
else:
teamWinCounts[team] = teamWinCounts.get(team,0) + 1
for teamName, Count in teamWinCounts.items():
print(teamName,':',Count)
# -
# ### Plotting the Results Of Team Winning
numberOfWins = teamWinCounts.values()
teamName = teamWinCounts.keys()
plt.bar(range(len(teamWinCounts)), numberOfWins)
plt.xticks(range(len(teamWinCounts)), list(teamWinCounts.keys()), rotation='vertical')
plt.xlabel('Team Names')
plt.ylabel('Number Of Win Matches')
plt.title('Analysis Of Number Of Matches win by Each Team From 2008 - 2018', color="Orange")
plt.show()
# ### Total Matches Played by Each team From 2008 - 2018
# +
totalMatchesCount = dict()
# For Team1
for team in matches_dataset['team1']:
totalMatchesCount[team] = totalMatchesCount.get(team, 0) + 1
# For Team2
for team in matches_dataset['team2']:
totalMatchesCount[team] = totalMatchesCount.get(team, 0) + 1
# Printing the total matches played by each team
for teamName, count in totalMatchesCount.items():
print('{} : {}'.format(teamName,count))
# -
# ### Plotting the Total Matches Played by Each Team
# +
teamNames = totalMatchesCount.keys()
teamCount = totalMatchesCount.values()
plt.bar(range(len(totalMatchesCount)), teamCount)
plt.xticks(range(len(totalMatchesCount)), list(teamNames), rotation='vertical')
plt.xlabel('Team Names')
plt.ylabel('Number Of Played Matches')
plt.title('Total Number Of Matches Played By Each Team From 2008 - 2018')
plt.show()
# -
| IPL 2008 - 2018 Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from datetime import datetime
import bz2
import glob
# +
from datetime import datetime
startTime = datetime.now()
print("Started: " + str(startTime))
stepTime = datetime.now()
# -
def printStepTime(text):
global stepTime
endTime = datetime.now()
print("Finished " + text + ": " + str(endTime) + ", took " + str(endTime - stepTime))
stepTime = endTime
# +
# Iterate through data in repo and uncompress it
# Put in one giant file; we'll process better later
with open('data/all_data.ndjson','w') as out:
for div in glob.glob('./data/OpenAccess-master/metadata/objects/*'):
print('Working on: ',div)
for file in glob.glob(f'{div}/*'):
with bz2.open(file, "rb") as f:
out.write(f.read().decode())
printStepTime("uncompressing data")
| unzip_repo_contents.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Geodepy Tutorial: Time-Dependent Datum Transformations
#
# This tutorial demonstrates the use of GeodePy for transformations between time-dependent dynamic datums. We'll work through each of the steps required to tranform a coordinate in ITRF2005 at epoch 2000.0 to ITRF2014 at 2020.0 on the Australian Plate, then build a function containing these steps to simplify this process.
# Start by importing the following Functions and Modules (datetime is used for the representation of epochs):
from datetime import date
from geodepy.constants import itrf14togda20, itrf14to05
from geodepy.transform import *
# We'll define a point for transformation. For this exercise, we'll use a point called Bob which is in terms of ITRF2005 at epoch 2000.0 and is shown using a UTM projection. The parts below are (zone, easting, northing, ellipsoidal height):
bob = (53, 386353.2371, 7381852.2967, 587.5814)
# #### Part 1: Getting Things in the Right Format
# To convert between time-dependent datums, we need our coordinates to be in Cartesian (XYZ) format. To get Bob in this format, we start by converting the UTM part (zone, easting and northing) into Geographic (latitude and longitude) format:
lat, lon, psf, grid_conv = grid2geo(bob[0], bob[1], bob[2])
lat, lon
# We can now use Bob's Latitude, Longitude and Ellipsoidal Height to calculate Cartesian Coordinates for Bob. These are still in terms of ITRF2005 at epoch 2000.0:
x, y, z = llh2xyz(lat, lon, bob[3])
x, y, z
# #### Part 2: Moving to the Reference Epoch
# Transformations between datums in GeodePy use Transformation Objects which are contained in the Constants module. These contain parameters used in Conformal (Helmert) 7 and 15 parameter transformations. To go between ITRF2005 and ITRF2014, we use `itrf14to05`, but because we're going the other way we use `-itrf14to05`:
-itrf14to05
# Before we can use this, we need to get our coordinates into the Reference Epoch of the transformation we plan to use (in this case, 2010.0 or the 1st Jan 2010). Because Bob is in Australia, we need to use the Australian Plate Motion Model to do this. It's parameters are in `itrf14togda20`:
itrf14togda20
# It also has a reference epoch; for `itrf14togda20`, it's 2020.0. So to move coordinates from 2000.0 to 2010.0 (+10.0 years) using `itrf14togda20` we need to use `conform14` with a `to_epoch` 10.0 years past our reference epoch (2020.0 + 10.0 = 2030.0):
x, y, z
x2, y2, z2 = conform14(x, y, z, date(2030, 1, 1), itrf14togda20)
x2, y2, z2
# We now have Bob's Cartesian Coordinates in ITRF2005 at epoch 2010.0!
# #### Part 3: The Transformation
# Now that we have Bob's coordinates in the reference epoch of `itrf14to05`, we can perform this transformation. Because we're not changing epochs in this part, we can use the `conform7` function which ignores the time-dependent parameters of our transformation:
x3, y3, z3 = conform7(x2, y2, z2, -itrf14to05)
x3, y3, z3
# This gives us Bob's Cartesian Coordinates in ITRF2014 at epoch 2010.0.
# #### Part 4: Moving to the Final Epoch
# The final tranformation is moving Bob's epoch to it's final destination (2020.0) using `itrf14togda20`. As the period of movement (2010.0 to 2020.0) is the same (+10.0 years), we use the same `to_epoch` of 2030.0 as in Part 2:
x4, y4, z4 = conform14(x3, y3, z3, date(2030, 1, 1), itrf14togda20)
x4, y4, z4
# These coordinates are now in terms of ITRF2014 at epoch 2020.0
# #### Part 5: Getting Things in their Original Format
# To bring everything back into the format we started, we'll step through the process in Part 1 in reverse. So we'll start by converting our Cartesian Coordinates to Geographic format:
lat_end, lon_end, ell_ht_end = xyz2llh(x4, y4, z4)
lat_end, lon_end, ell_ht_end
# Next, we'll convert the latitude and longitude into UTM grid coordinates:
hem_end, zone_end, east_end, north_end, psf_end, grid_conv_end = geo2grid(lat_end, lon_end)
zone_end, east_end, north_end
# Combining this with our ellipsoidal height from above, we get the final coordinate:
zone_end, east_end, north_end, ell_ht_end
# We can then compare the starting coordinate to the final coordinate and see the difference:
bob_end = (zone_end, east_end, north_end, ell_ht_end)
bob_end
bob
# #### Part 6: Simplifying steps into a single function
# To wrap up this tutorial, we'll combine all of the steps above into a single function:
# +
def bobtransform(zone, east, north, ell_ht):
lat, lon, psf, grid_conv = grid2geo(zone, east, north)
x, y, z = llh2xyz(lat, lon, ell_ht)
x2, y2, z2 = conform14(x, y, z, date(2030, 1, 1), itrf14togda20)
x3, y3, z3 = conform7(x2, y2, z2, -itrf14to05)
x4, y4, z4 = conform14(x3, y3, z3, date(2030, 1, 1), itrf14togda20)
lat_end, lon_end, ell_ht_end = xyz2llh(x4, y4, z4)
hem_end, zone_end, east_end, north_end, psf_end, grid_conv_end = geo2grid(lat_end, lon_end)
return zone_end, east_end, north_end, ell_ht_end
newbob = bobtransform(bob[0], bob[1], bob[2], bob[3])
newbob
| docs/tutorials/GeodePy Tutorial - Time-Dependent Datum Transformations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env_pda_2022
# language: python
# name: env_pda_2022
# ---
# <center><font size="+4">Programming and Data Analytics 1 2021/2022</font></center>
# <center><font size="+2">Sant'Anna School of Advanced Studies, Pisa, Italy</font></center>
# <center><img src="https://github.com/EMbeDS-education/StatsAndComputing20212022/raw/main/PDA/jupyter/jupyterNotebooks/images/SSSA.png" width="700" alt="EMbeDS"></center>
#
# <center><font size="+2">Course responsible</font></center>
# <center><font size="+2"><NAME> <EMAIL></font></center>
#
# <center><font size="+2">Co-lecturer </font></center>
# <center><font size="+2"><NAME> <EMAIL></font></center>
#
# ---
# <center><font size="+4">Lecture 4: Control and Repetition Statements</font><br/></center>
# <center><font size="+2"> and CSV manipulation/visualization applied on COVID-19 data</font></center>
# ---
from IPython.display import Image, display
img=Image(filename='images/tentativeLecturePlan.png',width=700)
display(img)
# <font size="+2"> How good are we so far - Kahoot quiz on previous class </font>
# * Using your phone or a different display go to [https://kahoot.it/](https://kahoot.it/)
# * Type the given PIN
# 
from IPython.display import IFrame
IFrame("https://kahoot.it/", 500, 400)
# # Intro: we need to control at runtime the execution flow of our programs
# ## We don't precisely do cooking recipes
# At the beginning of the course we made a parallel among cooking recipes and software programs.
#
# * A recipe is a list of instructions to be executed in sequence. No decisions have to be taken.
# * When programming, it is very rare to deal with problems that require only such simple processing.
#
# Most of the times we need to write programs able to take decisions:
# * Depending on the input
# * Depending on the current values of certain variables in the program
# ## Today
# In this lecture we will discuss the specific statements (aka control and repetition structures) necessary to:
# * __IF__: Execute blocks of statements only if a certain condition holds
# * __LOOP__: Execute blocks of statements several times as long a certain condition holds
# Statements like `if` and loops (`while`, `for`) that contain _blocks of statements_
# * are known as [__compound statements__](https://docs.python.org/3/reference/compound_stmts.html)
# * affect/control the execution of those other statements in some way
# * blocks are built using the _indentation rule_, as explained below.
# * What matters are the spaces preceding a statement
# * No explicit block delimiters like `{...}` are needed
# # If statement
# This section presents the various forms in which we can use the [`if` statement](https://docs.python.org/3/reference/compound_stmts.html#the-if-statement)
# ## Choose whether executing a block or not
# In its simplest form, an `if` statement consists of
# * A guard: a Boolean expression (as presented in class 2 evaluated to either `True` or `False`
# * A body: block of statements executed only if the guards evaluates to `True`
# +
x=1
y=2
if x==y:
print('The guard was satisfied')
print('After the first if\n')
x=1
y=1
if x==y :
print('The guard was satisfied')
print('After the second if')
# -
# > Pay attention to tabs/indentation! The body of the `if` is defined by the indentation <br/>
# > The following prgram is equal to the first one above up to a tab, behaves differently!
x=1
y=2
if x==y :
print('The guard was satisfied')
print('Still in the if!')
print('Not in the if!')
# ## Choose which block to execute among two blocks
# What if we need to choose between executing two blocks of statements according to a condition?
# +
deposit = 10
withdraw = 6
if deposit >= withdraw :
deposit-=withdraw
print('Withdrawing',withdraw,'succeeded. ')
print('We are left with',deposit)
else:
missing = withdraw - deposit
actualWithdraw = deposit
deposit = 0
print('Withdrawing',withdraw,'failed. ')
print('We withdrew',actualWithdraw,'remaining with',deposit)
# -
if deposit >= withdraw :
deposit-=withdraw
print('Withdrawing',withdraw,'succeeded. ')
print('We are left with',deposit)
else:
missing = withdraw - deposit
actualWithdraw = deposit
deposit = 0
print('Withdrawing',withdraw,'failed. ')
print('We withdrew',actualWithdraw,'remaining with',deposit)
# > The copy-paste of code above is __bad practice__. We will see in the next class how to avoid it!
# ## Choose wich block to execute among several blocks
print('The cappuccino law.')
hour=int(input('What time is it (0-24)?'))
if hour < 5 :
print('No. But some people have hot milk to help their sleep.')
elif hour <= 11 :
print('You can safely have a cappuccino.')
elif hour == 12 :
print("That's shady. Drink it quickly.")
elif hour <= 15 :
print('No way!')
elif hour <= 20 :
print('No. But you can have hot milk if you are sick.')
else:
print("No. But some people have hot milk before going to sleep.")
# There can be:
# * 0, 1, or more `elif`
# * take this branch if previous guards failed and this one passes
# * 0 or 1 `else`
# * a sort of _default_ case taken if all other guards fail
#
# Of course, we can have nested `if`
# * The `else` and `elif` are matched to the correct `if` according to the indentation rule
n=int(input())
if n >= 2 :
if n <= 5 :
print(n,'belongs to the interval [2,5]')
else :
print(n,'belongs to the interval (5,infinity)')
else :
print(n,'belongs to the interval [0,2)')
# # Looping constructs
# A basic __`if`__ statement can be used to
# * decide whether executing a block __0 times or 1 time__
#
# A looping construct (__`while`__,__`for`__) is like a basic `if`, but can be used to
# * decide whether executing a block __0 times, 1 time, 2 times, 3 times, 4 times ...__
# ## While loops
# We begin from [while loops](https://docs.python.org/3/reference/compound_stmts.html#the-while-statement)
# * showing how we can use them to compute the factorial of a number: $n! = n\cdot(n-1)\cdot(n-2)\cdot\ldots\cdot1$
# +
nstr = input('Give me a (small) number. I will compute its factorial')
n=int(nstr)
fact=n
factstr=str(n)
while n >1:
n-=1
fact*=n
factstr = factstr + '*' + str(n)
print('The factorial of',nstr,'is',fact,'computed as',factstr)
# -
# Echo lines from console as long as the user does not type `'end'`
line = ''
while line != 'end':
line = input('Next line:')
print(line.upper())
print('Bye!')
# We can modify the execution flow within an iteration using
# * `break` interrupts the current iteration and exits the loop
# * `continue` interrupts the current iteration and go to the next iteration
#
# Let's print only non-numeric strings until we get `'end'`
from IPython.display import Image, display
img=Image(filename='images/breakContinue.png',width=500)
display(img)
#Echo without printing `'end'`
line = ''
while True :
line = input('Next line:')
if line == 'end':
break
print(line.upper())
print('Bye!')
#Echo without printing `'end'` and without printing numbers
line = ''
while True :
line = input('Next line:')
if line == 'end' :
break
elif line.isnumeric():
continue
print(line.upper())
print('Bye!')
# For pedagogical reasons, I gave solutions with `continue` and `break`.
# - In reality, I would have done it differently, using a more intuitive/simpler approach.
#
# For example, we can get 'echo without printing `end`' by
# * just anticipating the first `input()` before the loop
# * moving the input inside the loop at the end of the loop: each iteration reads the value for the next iteration
#
# > You should always think carefully on how to write your programs, always choosing the __simplest approach__.
#
#Echo without printing `'end'`
line = input('Next line:')
while line!='end' :
print(line.upper())
line = input('Next line:')
print('Bye!')
# While loops can have as well an `else` branch,
# * The block of the `else` branch is executed the first time the guard evaluates to `False`
# * but **it is not executed** if the cycle terminated due to a `break`
# * I __discourage__ the use of this because
# * It does not add much to the expressiveness
# * It is not particularly intuitive
# * Might lead to bugs: not executed if the loop terminates due to a `break`
#Echo without printing `'end'`
stop = False
while not stop :
print('Loop body')
stop = True
#break
else :
print('Else branch')
print('Bye!')
# ## For loops
# We now look at [`for` loops](https://docs.python.org/3/reference/compound_stmts.html#the-for-statement)
# * They have pretty much the same expressive power of while loops,
# * but make it easier the coding of certain computations
# * They are useful when
# * you have to iterate over a collection
# * you know in advance the number `n` of iterations you need
# * You obtain this using the collection `range(n)`
# ### Basic usage
lst2=[0]*10
for i in range(10):
lst2[i]=i*2
print(lst2)
for c in 'ciao':
print(c)
# Intuitively, **a for loop is not so different from a list comprehension**
# * But they are very different constructs.
# * List comprehensions can be used as an expression to be assigned.
# +
rng = range(9)
lst = [i*2 for i in rng]
print(lst)
lst2=[]
for i in rng:
lst2.append(i*2)
print(lst2)
# -
# Rule of thumb:
# * Whenever you do simple iterations on the elements of a collection to create a new one
# * __use list comprehensions__. They are more efficient!
# * Whenever you have to do more operations, e.g. execute more statements in each iteration
# * use for loops
for i in range(10):
print('Hello')
print('How are you?')
print('...')
print('The square of',i,'is',i**2)
# ### Nested loops
# Let's create a 3x3 matrix containing random values from 0 to 1
import random
for i in range(10):
print(random.random())
# +
import random
n_rows = 3
n_cols = 3
matrix = [ [ random.random() for c in range(n_cols) ] for r in range(n_rows) ]
print(matrix)
print()
#Let's pretty-print the created matrix
for r in range(n_rows) :
for c in range(n_cols) :
val = '{:.2f}'.format(matrix[r][c])
print(val,end="")
print()
#help(print)
# +
#Let's compute some quantities on the matrix
maxm=-1 #the maximum value
minm=2 #the minimum value
summ=0 #the sum of the values in the matrix
for r in range(n_rows) :
for c in range(n_cols) :
if matrix[r][c] > maxm :
maxm = matrix[r][c]
if matrix[r][c] < minm :
minm = matrix[r][c]
summ+=matrix[r][c]
print(f'Sum {summ:.2f}, Min {minm:.4f}, Max {maxm}')
print('Sum {:.2f}, Min {:.4f}, Max {}'.format(summ,minm,maxm))
# -
# Did you notice
# - `f'Sum {summ}, Min {minm}, Max {maxm}'`
# - `'Sum {}, Min {}, Max {}'.format(summ,minm,maxm)`
#
# These are examples of __string formatting__
# - It can be convenient. [Take a look at further options](https://realpython.com/python-string-formatting/)
# ### Iterating over more collections at once ...
numbers="12345"
vowels="aeiou"
for (n,v) in zip(numbers,vowels):
print(n,v)
# ### Two subtleties on the `for` loop
# Two subtleties on the `for` loop from [here](https://docs.python.org/3/reference/compound_stmts.html#the-for-statement)
# * In my opinion it is not good practice to use these two features below, but you have to know about them to defend against them ;)
# * In particular, you should __avoid 2__. It makes the code much more difficult to read and debug
print('Subtlety 1: update the value of the iterating variable')
for i in range(1,4):
print(i)
i = i*i
print('Square value is',i)
print()
# As you can see
# * We can modify the variable `i` in an iteration. But this does not affect the next iterations: a new value will be assigned to `i` by the loop.
# * You should better use a different ad-hoc variable
#
print('Subtlety 2: modifiy the list on which you are iterating')
#DON"T DO THIS!
lst = [1,2,3,4]
for i in lst :
print(i)
if i==2:
lst.remove(i)
print("NEVER DO SOMETHING LIKE THIS!!! \nOr you will regret it when spending hours trying to debug/understand your code.\n")
# Did you notice what happened?
# * The element `3` has been skipped. Why?
# * Because it is after the element we removed. Ok, but why?
# * Because:
# * The `for` loop has an internal counter to remember the current position in the list it is iterating.
# * When we removed `2`, `3` got the position in the list previously occupied by `2`
# * Therefore the `for` loop erroneously believed to have already processed `3` and went to the next element (`4`).
# Similarly to the `while` statement, the `for` supports
# * `continue`
# * `break`
# * `else`
# # How is it going with our remote setting?
# Meanwhile at last week G7...
from IPython.display import Audio,Image, YouTubeVideo
id='_g6jAnLD2r0' #https://www.youtube.com/watch?v=_g6jAnLD2r0&ab_channel=ITVNews
YouTubeVideo(id=id,width=600,height=300)
# # Application to the official Italian COVID-19 data
# You might know that Protezione Civile publishes everyday data on the status of the COVID-19 epidemy in Italy
from IPython.display import IFrame
IFrame("http://opendatadpc.maps.arcgis.com/apps/opsdashboard/index.html#/b0c68bce2cce478eaac82fe38d4138b1", 900, 700)
# By following the 'CSV' link in the bottom-right of the page, we can download several data in CSV format.
# * For example, [this link](https://github.com/pcm-dpc/COVID-19/blob/master/dati-andamento-nazionale/dpc-covid19-ita-andamento-nazionale.csv) points to the latest information at national level
#
# We are going to see next how `if` and `loops` can be used to perform easy manipulations on these data.
# * For easiness of distribution of this document, we downloaded a copy of this file updated on
# * Friday the 8th of May 2020
# * The file is available [here](csv/covid19/dpc-covid19-ita-andamento-nazionale.csv).
# * Friday the 26th of February 2021
# * The file is available [here](csv/covid19/dpc-covid19-ita-andamento-nazionale_26_02_2021.csv).
# * If you follow these links, jupyter will provide a nice rendering
# * For pedagogical reasons, we will not use yet advanced libraries for loading and manipulating this data
# * This will make more evident the role of `if` and `for`, and how to read files
# * This is good to help you understanding that there is no magic in programming
# ## Import the necessary modules
# First of all, we need to import two required libraries.
# * The first time you run this code, you might need to install these libraries
# +
#import sys
# #!{sys.executable} -m pip install matplotlib
# -
import csv
import matplotlib.pyplot as plt
# In particular
# * `csv` is a Python library offering functionalities to load CSV files. Find more [here](https://docs.python.org/3/library/csv.html)
# * `matlplotlib` is a Python library for creating plots. Find more [here](https://matplotlib.org/)
# * We already used this library in the previous class
# ## Load the data of interest
# +
#fileName='csv/covid19/dpc-covid19-ita-andamento-nazionale.csv'
fileName='csv/covid19/dpc-covid19-ita-andamento-nazionale_26_02_2021.csv'
#I want to 'open' a csvfile to 'r'ead it
# For the time being, ignore the with statement.
# Assume it does: csvfile = open(fileName, 'r')
# I will tell you more on this in two classes from now
with open(fileName, 'r') as csvfile:
#Intuitively,
# `csv` allows us to read csv files row by row.
# `rows` is a list of rows in the file.
# Each row is in turn a list of strings containing one entry per column
rows = csv.reader(csvfile, delimiter=',')
#We load the header of the file in the list `header`
header = next(rows,None)
#print(header)
#Let's get the data of interest ... using 3 approaches
labels_of_interest = ['terapia_intensiva', 'totale_casi' , 'totale_positivi' , 'tamponi' ]
labels_en = ['Intensive care' , 'Total infected', 'Currently infected', 'Swab/tests']
#1) Using a for loop
label_to_column = dict()
column = 0
for label in header :
if label in labels_of_interest:
#I have found a label of interest
label_to_column[label]=column
#We exit the loop as soon as we have considered all labels of interest
if len(label_to_column) == len(labels_of_interest) :
break
column += 1
#print(label_to_column)
#2) Using a while loop
label_to_column2 = dict()
column = 0
while len(label_to_column2) != len(labels_of_interest) :
label = header[column]
if label in labels_of_interest:
label_to_column2[label]=column
column+=1
#print(label_to_column2)
#3) Using dictionary comprehension
label_to_column3 = { label : header.index(label) for label in header if label in labels_of_interest }
#print(label_to_column3)
#print(label_to_column==label_to_column2==label_to_column3)
#We now iterate over all rows to load the data
intensive_care = []
total_infected = []
currently_infected = []
tests = []
#We put these lists in another list to simplify the for loop below
loadedData = [intensive_care,total_infected,currently_infected,tests]
for row in rows :
l=0
for label in labels_of_interest:
column = label_to_column[label]
data = row[column]
loadedData[l].append(int(data))
l+=1
#print(intensive_care)
#print(total_infected)
#print(tests)
# -
# ## Populate the plots
# +
#We now create three plots
x = range(len(intensive_care))
for i in range(len(labels_en)) :
plt.plot(x,loadedData[i], label=labels_en[i],linewidth=3)
#plt.plot(x,intensive_care, label='Intensive care',linewidth=3)
#plt.plot(x,total_infected, label='Total infected',linewidth=3)
#plt.plot(x,currently_infected, label='Currently infected',linewidth=3)
#plt.plot(x,tests, label='Swab/tests',linewidth=3)
plt.legend(fontsize=13)
plt.xlabel('Days')
plt.ylabel('Values')
#plt.savefig('Italian COVID-19 data',bbox_inches='tight')
plt.show()
#plt.show()
# -
# ## Navigate data to compute new data of interest
peak_infected = max(currently_infected)
day_of_peak = currently_infected.index(peak_infected)
print('This data spans',len(currently_infected),'days.')
print('On day',day_of_peak,'we had had the peak of infected:',peak_infected)
print('Therefore we reached the peak',len(currently_infected)-day_of_peak,'days ago')
print('\tActually',len(currently_infected)-day_of_peak,' days before the latest measured day in the CSV')
print('On the last measured day we have',currently_infected[len(currently_infected)-1],'currently infected people')
# Let's show this information **graphcially**
# +
plt.plot(x,currently_infected, label='Currently infected',linewidth=3)
plt.legend(fontsize=13)
plt.xlabel('Days')
plt.ylabel('Values')
#plt.savefig('Italian COVID-19 data',bbox_inches='tight')
msg='The peak on day\n'+str(x[day_of_peak])+" was "+str(currently_infected[day_of_peak])
plt.annotate(msg, xy=(x[day_of_peak], currently_infected[day_of_peak]),
xytext=(x[day_of_peak]-200, currently_infected[day_of_peak]-300000),
arrowprops=dict(facecolor='black', shrink=0.05))
last = len(currently_infected)-1
msg='The most recent\nvalue is '+str(currently_infected[last])
plt.annotate(msg, xy=(x[last], currently_infected[last]),
xytext=(x[last]-100, currently_infected[last]-300000),
arrowprops=dict(facecolor='black', shrink=0.2))
plt.show()
# -
# What if we are interested in the **daily variation of infected**?
# * In absolute value
# * In percentage
# +
prevDay=0
daily_variation=[]
daily_variation_perc=[]
for i in range(len(currently_infected)) :
difference = currently_infected[i] - prevDay
daily_variation.append(difference)
if(prevDay==0):
daily_variation_perc.append(0)
else:
daily_variation_perc.append(difference/prevDay*100)
prevDay=currently_infected[i]
#print(daily_variation)
#print(daily_variation_perc)
plt.plot(x,daily_variation, label='Daily variation',linewidth=3)
plt.plot(x,[0]*len(x))
plt.legend(fontsize=13)
plt.show()
plt.plot(x,daily_variation_perc, label='Daily variation %',linewidth=3)
plt.ylabel('% variation')
plt.legend(fontsize=13)
plt.show()
# -
# What if we are interested in the
# * **daily variation after the peak**?
#
# Well, luckily we know how to obtain subsuquences...
plt.plot(x[day_of_peak:],daily_variation[day_of_peak:], label='Daily variation after the peak',linewidth=3)
plt.legend(fontsize=13)
plt.show()
# # Next class...
# At the beginning of this class we have seen a clear example of __spaghetti code__
# +
deposit = 20
withdraw = 7
if deposit >= withdraw :
deposit-=withdraw
print('Withdrawing',withdraw,'succeeded. ')
print('We are left with',deposit)
else:
missing = withdraw - deposit
actualWithdraw = deposit
deposit = 0
print('Withdrawing',withdraw,'failed. ')
print('We withdrew',actualWithdraw,'remaining with',deposit)
print()
if deposit >= withdraw :
deposit-=withdraw
print('Withdrawing',withdraw,'succeeded. ')
print('We are left with',deposit)
else:
missing = withdraw - deposit
actualWithdraw = deposit
deposit = 0
print('Withdrawing',withdraw,'failed. ')
print('We withdrew',actualWithdraw,'remaining with',deposit)
print()
if deposit >= withdraw :
deposit-=withdraw
print('Withdrawing',withdraw,'succeeded. ')
print('We are left with',deposit)
else:
missing = withdraw - deposit
actualWithdraw = deposit
deposit = 0
print('Withdrawing',withdraw,'failed. ')
print('We withdrew',actualWithdraw,'remaining with',deposit)
print()
# -
# Did you notice that we copy-pasted several times the same `if-else` statement?
# This is very bad because:
# * It makes the code less readable
# * It makes the code less manutenable
# * What if you have to change something in the if-else statement?
# * You would have to modify each copy... which is very error-prone
#
# Solution: declare a new function!
# +
def withdraw_amoount(deposit,withdraw):
if deposit >= withdraw :
deposit-=withdraw
print('Withdrawing',withdraw,'succeeded. ')
print('We are left with',deposit)
else:
missing = withdraw - deposit
actualWithdraw = deposit
deposit = 0
print('Withdrawing',withdraw,'failed. ')
print('We withdrew',actualWithdraw,'remaining with',deposit)
print()
return deposit
deposit = 20
withdraw = 7
deposit = withdraw_amoount(deposit,withdraw)
deposit = withdraw_amoount(deposit,withdraw)
deposit = withdraw_amoount(deposit,withdraw)
# -
# Furthermore, functions, or libraries (collections of functions and types) can be declared in different files, maintaining your code
# * simple
# * modular
# * manutenable
| PDA/jupyter/jupyterNotebooks/_04ControlAndRepetitionStatements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparing model metrics with SageMaker Pipelines and SageMaker Model Registry
#
# ### SageMaker Model Registry
# SageMaker Model Registry is a central location where customers can manage their machine learning models, compare different model versions and visualize metrics. It's a registry with which data scientists can register machine learning models with relevant metadata and metrics, and from which machine learning engineers or DevOps engineers can deploy machine learning models. In larger MLOps systems, a model registry is usually where the teams in charge of deploying machine learning models, meet the teams in charge of developing and training machine learning models.
#
# 
#
#
# ### MLOps
#
# MLOps, or Machine Learning Operations, is the concept of applying DevOps practices on the lifecycle of a machine learning model. Among many other things, MLOps usually consists of two workflows that sits on either side of a machine learning model registry; one to train a model and one to deploy a model. A model registry is a central location to manage machine learning models, where ML engineers and data scientists can compare different model versions, visualize metrics, and decide which versions to accept and which to reject. Ideally, approving a new version of a model triggers a pipeline that ultimately deploys the model into production.
#
#
# From a high-level perspective, it can look like this.
#
# 
#
# The deployment pipeline is what is most similar to traditional CI/CD methods, propagating an artifact through a pipeline that performs testing and if the tests are successful, deploys them. Any CI/CD tools can be used for the deployment, but to automate the entire machine learning model lifecycle, a pipeline on "the other side" of the model registry is required as well. A pipeline that ultimately produces a new machine learning model and registers it with the model registry. A SageMaker Pipeline.
#
# 
#
#
# **This notebook**, demonstrate how SageMaker Pipelines can be used to create a reusable machine learning pipeline that preprocesses, trains, evaluates and registers a machine learning model with the SageMaker Model Registry for visualization and comparison of different model versions.
#
# ### Amazon SageMaker Pipelines
#
# Amazon SageMaker Pipelines is a purpose-built, easy-to-use CI/CD service for machine learning. With SageMaker Pipelines, customers can create machine learning workflows with an easy-to-use Python SDK, and then visualize and manage workflows using Amazon SageMaker Studio.
#
# #### SageMaker Pipeline steps and parameters
# SageMaker pipelines works on the concept of steps. The order steps are executed in is inferred from the dependencies each step have. If a step has a dependency on the output from a previous step, it's not executed until after that step has completed successfully.
#
# SageMaker Pipeline Parameters are input parameters specified when triggering a pipeline execution. They need to be explicitly defined when creating the pipeline and contain default values.
#
# To know more about the type of steps and parameters supported, check out the [SageMaker Pipelines Overview](https://docs.aws.amazon.com/sagemaker/latest/dg/pipelines-sdk.html).
#
# #### SageMaker Pipeline DAG
#
# When creating a SageMaker Pipeline, SageMaker creates a Direct Acyclic Graph, DAG, that customers can visualize in Amazon SageMaker Studio. The DAG can be used to track pipeline executions, outputs and metrics. In this notebook, a SageMaker pipeline with the following DAG is created:
#
# 
# ## Predict customer churn with XGboost
#
# ### Data
#
# This notebook uses a synthetic dataset.
#
# This dataset contains information about phone carrier customers, such as customer location, phone number, number of phone calls etc.
#
#
# ### Overview
# **Disclaimer** This notebook was created using [Amazon SageMaker Studio](https://aws.amazon.com/sagemaker/studio/) and the `Python3(DataScience) kernel`. SageMaker Studio is required for the visualizations of the DAG and model metrics to work.
#
# The purpose of this notebook is to demonstrate how SageMaker Pipelines can be used to preprocess, train, evaluate and push new machine learning models into the SageMaker model registry for version comparison. All scripts to preprocess the data and evaluate the trained model have been prepared in advance and are available here:
# - [preprocess.py](preprocess.py)
# - [evaluate.py](evaluate.py).
#
#
#
# #### Table of contents
#
# * [Define parameters](#parameters)
# In this section the parameters of the pipeline are defined.
# * [Preprocess step](#preprocess)
# In this section an SKLearnProcessor is created and used in a Preprocess step.
# * [Train step](#train)
# In this section the SageMaker managed XGboost container is downloaded and an Estimator object and Training step are created.
# * [Evaluate model step](#evaluate)
# In this section a ScriptProcessor is created, used in a Processing step to compute some evaluation metrics of the previously trained model.
# * [Condition step](#condition)
# In this section a condition step is defined, using the metrics from the evaluation step.
# * [Register model step](#register)
# In this section a register model step is created, where the trained model is registered with the SageMaker Model Registry.
# * [Create SageMaker Pipeline](#orchestrate)
# In the last section, the SageMaker pipeline is created and all steps orchestrated before executing the pipeline.
#
# !pip install -U sagemaker --quiet # Ensure latest version of SageMaker is installed
import sagemaker
import sagemaker.session
session = sagemaker.session.Session()
region = session.boto_region_name
role = sagemaker.get_execution_role()
bucket = session.default_bucket()
model_package_group_name = "Churn-XGboost" # Model name in model registry
prefix = "sagemaker/Churn-xgboost" # Prefix to S3 artifacts
pipeline_name = "ChurnPipeline" # SageMaker Pipeline name
# +
# Upload the raw datasets to S3
large_input_data_uri = session.upload_data(
path="dataset/large/churn-dataset.csv", key_prefix=prefix + "/data/large"
)
small_input_data_uri = session.upload_data(
path="dataset/small/churn-dataset.csv", key_prefix=prefix + "/data/small"
)
test_data_uri = session.upload_data(path="dataset/test/test.csv", key_prefix=prefix + "/data/test")
print("Large data set uploaded to ", large_input_data_uri)
print("Small data set uploaded to ", small_input_data_uri)
print("Test data set uploaded to ", test_data_uri)
# -
# <a id='parameters'></a>
#
# ### Pipeline input parameters
#
# Pipeline Parameters are input parameter when triggering a pipeline execution. They need to be explicitly defined when creating the pipeline and contain default values.
# +
from sagemaker.workflow.parameters import (
ParameterInteger,
ParameterString,
)
# How many instances to use when processing
processing_instance_count = ParameterInteger(name="ProcessingInstanceCount", default_value=1)
# What instance type to use for processing
processing_instance_type = ParameterString(
name="ProcessingInstanceType", default_value="ml.m5.large"
)
# What instance type to use for training
training_instance_type = ParameterString(name="TrainingInstanceType", default_value="ml.m5.xlarge")
# Where the input data is stored
input_data = ParameterString(
name="InputData",
default_value=small_input_data_uri,
)
# What is the default status of the model when registering with model registry.
model_approval_status = ParameterString(
name="ModelApprovalStatus", default_value="PendingManualApproval"
)
# -
# <a id='preprocess'></a>
#
# ## Preprocess data step
# In the first step an sklearn processor is created, used in the ProcessingStep.
# +
from sagemaker.sklearn.processing import SKLearnProcessor
from sagemaker.processing import ProcessingInput, ProcessingOutput
from sagemaker.workflow.steps import ProcessingStep
from sagemaker.workflow.functions import Join
from sagemaker.workflow.execution_variables import ExecutionVariables
# Create SKlearn processor object,
# The object contains information about what instance type to use, the IAM role to use etc.
# A managed processor comes with a preconfigured container, so only specifying version is required.
sklearn_processor = SKLearnProcessor(
framework_version="0.23-1",
role=role,
instance_type=processing_instance_type,
instance_count=processing_instance_count,
base_job_name="churn-processing-job",
)
# Use the sklearn_processor in a Sagemaker pipelines ProcessingStep
step_preprocess_data = ProcessingStep(
name="Preprocess-Churn-Data",
processor=sklearn_processor,
inputs=[
ProcessingInput(source=input_data, destination="/opt/ml/processing/input"),
],
outputs=[
ProcessingOutput(
output_name="train",
source="/opt/ml/processing/train",
destination=Join(
on="/",
values=[
"s3://{}".format(bucket),
prefix,
ExecutionVariables.PIPELINE_EXECUTION_ID,
"train",
],
),
),
ProcessingOutput(
output_name="validation",
source="/opt/ml/processing/validation",
destination=Join(
on="/",
values=[
"s3://{}".format(bucket),
prefix,
ExecutionVariables.PIPELINE_EXECUTION_ID,
"validation",
],
),
),
ProcessingOutput(
output_name="test",
source="/opt/ml/processing/test",
destination=Join(
on="/",
values=[
"s3://{}".format(bucket),
prefix,
ExecutionVariables.PIPELINE_EXECUTION_ID,
"test",
],
),
),
],
code="preprocess.py",
)
# -
# <a id='train'></a>
#
# ## Train model step
# In the second step, the train and validation output from the precious processing step are used to train a model. The XGBoost container is retrieved and then an XGBoost estimator is created, on which hyper parameters are specified before the training step is created.
# +
from sagemaker.inputs import TrainingInput
from sagemaker.workflow.steps import TrainingStep
from sagemaker.estimator import Estimator
# Fetch container to use for training
image_uri = sagemaker.image_uris.retrieve(
framework="xgboost",
region=region,
version="1.2-2",
py_version="py3",
instance_type=training_instance_type,
)
# Create XGBoost estimator object
# The object contains information about what container to use, what instance type etc.
xgb_estimator = Estimator(
image_uri=image_uri,
instance_type=training_instance_type,
instance_count=1,
role=role,
disable_profiler=True,
)
xgb_estimator.set_hyperparameters(
max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
objective="binary:logistic",
num_round=25,
)
# Use the xgb_estimator in a Sagemaker pipelines ProcessingStep.
# NOTE how the input to the training job directly references the output of the previous step.
step_train_model = TrainingStep(
name="Train-Churn-Model",
estimator=xgb_estimator,
inputs={
"train": TrainingInput(
s3_data=step_preprocess_data.properties.ProcessingOutputConfig.Outputs[
"train"
].S3Output.S3Uri,
content_type="text/csv",
),
"validation": TrainingInput(
s3_data=step_preprocess_data.properties.ProcessingOutputConfig.Outputs[
"validation"
].S3Output.S3Uri,
content_type="text/csv",
),
},
)
# -
# <a id='evaluate'></a>
#
# ## Evaluate model step
# When a model a trained, it's common to evaluate the model on unseen data before registering it with the model registry. This ensures the model registry isn't cluttered with poorly performing model versions. To evaluate the model, create a ScriptProcessor object and use it in a ProcessingStep.
#
# **Note** that a separate preprocessed test dataset is used to evaluate the model, and not the output of the processing step. This is only for demo purposes, to ensure the second run of the pipeline creates a model with better performance. In a real-world scenario, the test output of the processing step would be used.
#
# +
from sagemaker.processing import ScriptProcessor
from sagemaker.workflow.properties import PropertyFile
# Create ScriptProcessor object.
# The object contains information about what container to use, what instance type etc.
evaluate_model_processor = ScriptProcessor(
image_uri=image_uri,
command=["python3"],
instance_type=processing_instance_type,
instance_count=processing_instance_count,
base_job_name="script-churn-eval",
role=role,
)
# Create a PropertyFile
# A PropertyFile is used to be able to reference outputs from a processing step, for instance to use in a condition step.
# For more information, visit https://docs.aws.amazon.com/sagemaker/latest/dg/build-and-manage-propertyfile.html
evaluation_report = PropertyFile(
name="EvaluationReport", output_name="evaluation", path="evaluation.json"
)
# Use the evaluate_model_processor in a Sagemaker pipelines ProcessingStep.
step_evaluate_model = ProcessingStep(
name="Evaluate-Churn-Model",
processor=evaluate_model_processor,
inputs=[
ProcessingInput(
source=step_train_model.properties.ModelArtifacts.S3ModelArtifacts,
destination="/opt/ml/processing/model",
),
ProcessingInput(
source=test_data_uri, # Use pre-created test data instead of output from processing step
destination="/opt/ml/processing/test",
),
],
outputs=[
ProcessingOutput(
output_name="evaluation",
source="/opt/ml/processing/evaluation",
destination=Join(
on="/",
values=[
"s3://{}".format(bucket),
prefix,
ExecutionVariables.PIPELINE_EXECUTION_ID,
"evaluation-report",
],
),
),
],
code="evaluate.py",
property_files=[evaluation_report],
)
# -
# <a id='register'></a>
#
# ## Register model step
# If the trained model meets the model performance requirements a new model version is registered with the model registry for further analysis. To attach model metrics to the model version, create a [ModelMetrics](https://docs.aws.amazon.com/sagemaker/latest/dg/model-monitor-model-quality-metrics.html) object using the evaluation report created in the evaluation step. Then, create the RegisterModel step.
#
# +
from sagemaker.model_metrics import MetricsSource, ModelMetrics
from sagemaker.workflow.step_collections import RegisterModel
# Create ModelMetrics object using the evaluation report from the evaluation step
# A ModelMetrics object contains metrics captured from a model.
model_metrics = ModelMetrics(
model_statistics=MetricsSource(
s3_uri=Join(
on="/",
values=[
step_evaluate_model.arguments["ProcessingOutputConfig"]["Outputs"][0]["S3Output"][
"S3Uri"
],
"evaluation.json",
],
),
content_type="application/json",
)
)
# Crete a RegisterModel step, which registers the model with Sagemaker Model Registry.
step_register_model = RegisterModel(
name="Register-Churn-Model",
estimator=xgb_estimator,
model_data=step_train_model.properties.ModelArtifacts.S3ModelArtifacts,
content_types=["text/csv"],
response_types=["text/csv"],
inference_instances=["ml.t2.medium", "ml.m5.xlarge", "ml.m5.large"],
transform_instances=["ml.m5.xlarge"],
model_package_group_name=model_package_group_name,
approval_status=model_approval_status,
model_metrics=model_metrics,
)
# -
# <a id='condition'></a>
#
# ## Accuracy condition step
# Adding conditions to the pipeline is done with a ConditionStep.
# In this case, we only want to register the new model version with the model registry if the new model meets an accuracy condition.
# +
from sagemaker.workflow.conditions import ConditionGreaterThanOrEqualTo
from sagemaker.workflow.condition_step import (
ConditionStep,
JsonGet,
)
# Create accuracy condition to ensure the model meets performance requirements.
# Models with a test accuracy lower than the condition will not be registered with the model registry.
cond_gte = ConditionGreaterThanOrEqualTo(
left=JsonGet(
step=step_evaluate_model,
property_file=evaluation_report,
json_path="binary_classification_metrics.accuracy.value",
),
right=0.7,
)
# Create a Sagemaker Pipelines ConditionStep, using the condition above.
# Enter the steps to perform if the condition returns True / False.
step_cond = ConditionStep(
name="Accuracy-Condition",
conditions=[cond_gte],
if_steps=[step_register_model],
else_steps=[],
)
# -
# <a id='orchestrate'></a>
#
# ## Pipeline Creation: Orchestrate all steps
#
# Now that all pipeline steps are created, a pipeline is created.
# +
from sagemaker.workflow.pipeline import Pipeline
# Create a Sagemaker Pipeline.
# Each parameter for the pipeline must be set as a parameter explicitly when the pipeline is created.
# Also pass in each of the steps created above.
# Note that the order of execution is determined from each step's dependencies on other steps,
# not on the order they are passed in below.
pipeline = Pipeline(
name=pipeline_name,
parameters=[
processing_instance_type,
processing_instance_count,
training_instance_type,
model_approval_status,
input_data,
],
steps=[step_preprocess_data, step_train_model, step_evaluate_model, step_cond],
)
# -
# #### Submit pipeline, and start it.
# +
# Submit pipline
pipeline.upsert(role_arn=role)
# Execute pipeline using the default parameters.
execution = pipeline.start()
execution.wait()
# List the execution steps to check out the status and artifacts:
execution.list_steps()
# -
# ## Visualize SageMaker Pipeline DAG
# In SageMaker Studio, choose `SageMaker Components and registries` in the left pane and under `Pipelines`, click the pipeline that was created. Then all pipeline executions are shown and the one just created should have a status of `Executing`. Selecting that execution, the different pipeline steps can be tracked as they execute.
#
# 
#
# ## Visualize model performance metrics
# Once the pipeline has completed successfully, metrics attached to the model version can be visualized. In SageMaker Studio, choose `SageMaker Components and registries` in the left pane and under `Model registry`, select the model package that was created. If a new model package group was created, only one model version should be visible. Click that version and visualize the model performance metrics.
#
# 
#
#
#
#
#
#
#
# ## Compare model performance metrics
#
# When there are more than one model version, they can be visualized side by side.
#
#
# Run the pipeline again, but with a larger dataset.
#
#
# +
# Execute pipeline with explicit parameters
execution = pipeline.start(
parameters=dict(
InputData=large_input_data_uri,
)
)
execution.wait()
# -
# ### Visualize
# Select both versions and right-click. Choose `Compare model versions`.
#
#
# 
# ## Clean up (optional)
# Delete the model registry and the pipeline to keep the studio environment tidy.
# +
def delete_model_package_group(sm_client, package_group_name):
try:
model_versions = sm_client.list_model_packages(ModelPackageGroupName=package_group_name)
except Exception as e:
print("{} \n".format(e))
return
for model_version in model_versions["ModelPackageSummaryList"]:
try:
sm_client.delete_model_package(ModelPackageName=model_version["ModelPackageArn"])
except Exception as e:
print("{} \n".format(e))
time.sleep(0.5) # Ensure requests aren't throttled
try:
sm_client.delete_model_package_group(ModelPackageGroupName=package_group_name)
print("{} model package group deleted".format(package_group_name))
except Exception as e:
print("{} \n".format(e))
return
def delete_sagemaker_pipeline(sm_client, pipeline_name):
try:
sm_client.delete_pipeline(
PipelineName=pipeline_name,
)
print("{} pipeline deleted".format(pipeline_name))
except Exception as e:
print("{} \n".format(e))
return
# +
import boto3
import time
client = boto3.client("sagemaker")
delete_model_package_group(client, model_package_group_name)
delete_sagemaker_pipeline(client, pipeline_name)
# -
| sagemaker-pipeline-compare-model-versions/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Background Image {#background_image_example}
# ================
#
# Add a background image with
# `pyvista.Plotter.add_background_image`{.interpreted-text role="func"}.
#
import pyvista as pv
from pyvista import examples
# Plot an airplane with the map of the earth in the background
#
# +
earth_alt = examples.download_topo_global()
pl = pv.Plotter()
actor = pl.add_mesh(examples.load_airplane(), smooth_shading=True)
pl.add_background_image(examples.mapfile)
pl.show()
# -
# Plot several earth related plots
#
# +
pl = pv.Plotter(shape=(2, 2))
pl.subplot(0, 0)
pl.add_text('Earth Visible as Map')
pl.add_background_image(examples.mapfile, as_global=False)
pl.subplot(0, 1)
pl.add_text('Earth Altitude')
actor = pl.add_mesh(earth_alt, cmap='gist_earth')
pl.subplot(1, 0)
topo = examples.download_topo_land()
actor = pl.add_mesh(topo, cmap='gist_earth')
pl.add_text('Earth Land Altitude')
pl.subplot(1, 1)
pl.add_text('Earth Visible as Globe')
pl.add_mesh(examples.load_globe(), smooth_shading=True)
pl.show()
| examples/02-plot/background_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
np.random.seed(0)
from sklearn.model_selection import cross_val_score, StratifiedKFold, GroupKFold
from sklearn.metrics import mean_squared_error as mse, precision_score, recall_score
from sklearn import utils
from scikitplot.metrics import plot_confusion_matrix, plot_calibration_curve
from scikitplot.estimators import plot_learning_curve, plot_feature_importances
from sklearn.dummy import DummyClassifier, DummyRegressor
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
import xgboost as xgb
from collections import defaultdict
import gc
import eli5
from eli5.sklearn import PermutationImportance
# -
#loading the data
train = pd.read_hdf('train_online_retail.h5').reset_index(drop=True)
train.info()
train.head(20)
# ## Dane:
#
# - `invoice` - invoice number
# - `stock_code` - product ID
# - `description` - product description
# - `quantity` - quantity of product bought in specific transaction
# - `invoice_date` - date of invoice issue
# - `price_unit` - unit price of product in pounds
# - `price_total` - total price = `price_unit * quantity`
# - `customer_id` - customers ID
# - `country` - name of the country from where the product was bought
# - `is_canceled` - boolean value if product was retailed or not
# - `is_test` - boolean value whether it's a test dataset
train['is_canceled'].value_counts()
test = pd.read_hdf('test_online_retail.h5').reset_index(drop=True)
test.info()
test.head()
# +
def dist(df):
print('detailed')
print( df['is_canceled'].value_counts(normalize=True).values * 100)
df_agg = df.groupby('invoice').sum()
df_agg['is_canceled'] = (df_agg['is_canceled'] > 0).astype(np.int)
print('orders')
print(df_agg['is_canceled'].value_counts(normalize=True).values * 100)
dist(train)
# +
X = train[['invoice', 'is_canceled']].values
y = train['is_canceled'].values
groups = train['invoice'].values
X_shuffled, y_shuffled, groups_shuffled = utils.shuffle(X, y, groups)
group_kfold = GroupKFold(n_splits=10)
for train_idx, test_idx in group_kfold.split(X_shuffled, y_shuffled, groups_shuffled):
sel_df = train[ train.invoice.isin( groups_shuffled[test_idx]) ]
dist(sel_df)
print("=====\n\n")
# -
#function to get features that we will use for model training
def get_feats(df):
feats = df.select_dtypes(np.number, np.bool).columns
black_list = ['is_canceled', 'is_test', 'is_canceled_pred', 'total_return', 'total_return_pred']
feats = [feat for feat in feats if feat not in black_list]
return feats
# +
#class for cross validation with GroupKfold method
class CrossValidation:
def __init__(self, groups, index, shuffle=True, n_splits=5, random_state=8888):
self.n_splits = n_splits
self.groups = groups
self.index = index
self.shuffle = shuffle
self.random_state = random_state
def kfold_split(self, X, y, groups=None):
if self.shuffle:
X, y, self.groups, self.index = utils.shuffle(X, y, self.groups, self.index, random_state = self.random_state)
group_kfold = GroupKFold(n_splits = self.n_splits)
for train_idx, test_idx in group_kfold.split(X, y, self.groups):
X_train, y_train = X[train_idx], y[train_idx]
X_test, y_test = X[test_idx], y[test_idx]
yield train_idx, test_idx
def get_index(self):
return self.index
# +
#function for calculating mse metric on aggregated data
def group_and_calc_mse(train):
train_agg = train.groupby('invoice')[ ['price_total', 'is_canceled_pred', 'is_canceled'] ].sum()
train_agg['is_canceled'] = train_agg['is_canceled'] > 0
train_agg['is_canceled_pred'] = train_agg['is_canceled_pred'] > 0
train_agg['total_return'] = train_agg['price_total'] * train_agg['is_canceled']
train_agg['total_return_pred'] = train_agg['price_total'] * train_agg['is_canceled_pred']
return mse( train_agg['total_return'].values, train_agg['total_return_pred'].values)
# -
#function to update test fold
def update_test_fold(y_pred, df, index_map, test_idx, pred_feat='is_canceled_pred'):
sel_index = df.index.isin( index_map[test_idx] )
df.loc[ sel_index, [pred_feat]] = y_pred
#function to run cross validation with plot learning curve, confusion_matrix and feature importance
def run_cv(train, model_cls, model_params, n_splits=5, shuffle_rows=True, target='is_canceled',
is_plot_learning_curve=True, is_plot_confusion_matrix=True, is_plot_feature_importances=True):
train.fillna(-1, inplace=True)
feats = get_feats(train)
print(feats)
X = train[feats].values
y = train[target].values
pred_feature = '{}_pred'.format(target)
train[pred_feature] = np.nan
cv = CrossValidation(train['invoice'], train.index)
for train_idx, test_idx in cv.kfold_split(X, y):
X_train, y_train = X[train_idx], y[train_idx]
X_test, y_test = X[test_idx], y[test_idx]
model = model_cls(**model_params)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
update_test_fold(y_pred, train, cv.get_index(), test_idx)
train[pred_feature] = train[pred_feature].astype('str')
train[pred_feature] = train[pred_feature]=='True'
print('mse: ', group_and_calc_mse(train))
if is_plot_learning_curve:
title = 'Learning curve: {} {}'.format(model_cls.__name__, model_params)
cv = CrossValidation(train['invoice'], train.index)
plot_learning_curve(model_cls(**model_params), X, y, title=title, figsize=(15, 5), random_state=0, cv=n_splits, scoring='recall')
if is_plot_confusion_matrix:
plot_confusion_matrix(train['is_canceled'], train[pred_feature])
if is_plot_feature_importances:
model = model_cls(**model_params)
model.fit(X, y)
perm = PermutationImportance(model, random_state=8888).fit(X, y)
return eli5.show_weights(perm, feature_names=feats)
#checking dummy classifier
run_cv(train, DummyClassifier, {'random_state': 8888})
run_cv(train, DummyClassifier, {'strategy': 'constant', 'constant': True, 'random_state': 8888})
run_cv(train, DummyClassifier, {'strategy': 'constant', 'constant': False, 'random_state': 8888})
run_cv(train, DecisionTreeClassifier, {'max_depth': 10})
# FEATURE ENGINEERING
# +
#kaggle submission
#orders_test['total_return'] = y_pred
#orders_test[ ['invoice', 'total_return'] ].to_csv('submit_dummy_model.csv', index=False)
# -
| retail prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # cProfile
#
# Python自带了几个性能分析的模块:profile、cProfile和hotshot,使用方法基本都差不多,无非模块是纯Python还是用C写的。cProfile效率更高
#
# # pyinstrument
# pip install pyinstrument
#
# 能生成tree 状的调用,但是缺调用次数统计
# run profile
# !python -m cProfile -o profile_test.out profile_test.py
# !python -c "import pstats; p=pstats.Stats('profile_test.out'); p.print_stats()"
# 可以设置排序方式,例如以花费时间多少排序
# !python -c "import pstats; p=pstats.Stats('profile_test.out'); p.sort_stats('time').print_stats()"
# # sort_stats支持以下参数:
# * calls, cumulative, file, line, module, name, nfl, pcalls, stdname, time
# # pyinstrument
#
# https://pyinstrument.readthedocs.io/en/latest/guide.html#profile-a-python-script
#
#
# ## 常用命令
# * python -m pyinstrument --show-all -r json -o 1.json main_v1_ye.py
# * python -m pyinstrument --show-all -r html -o 1.html main_v1_ye.py
| pybasic/profile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv('/home/anderson/Desktop/Sao-Paulo-Crime-Study/output.csv', encoding = 'latin1')
df['RUBRICA'].value_counts()
(df['RUBRICA'] == 'HomicÃdio qualificado (art. 121, §2o.)' ).sum()
df.RUBRICA
# ### Vou selecionar homicídio qualificado, Lesão Corporal seguida de morte, que são os 2 crimes com dolo que resultam em morte.
list = ['HomicÃdio qualificado (art. 121, §2o.)']
list
df.head()
# ### Abaixo faço um for loop pra construir um dataframe apenas desses dois crimes:
# +
for i in list:
df = df[df['RUBRICA']==i]
# -
df.head(3)
df['DATA_OCORRENCIA_BO'] = pd.to_datetime(df['DATA_OCORRENCIA_BO'])
# ### Quais os meses, dias da semana e horários mais comuns de ocorrências de homicídio qualificado ?
# +
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set_style('darkgrid')
# -
df['dia_da_semana'] = df['DATA_OCORRENCIA_BO'].dt.weekday_name
df['mes'] = df['DATA_OCORRENCIA_BO'].dt.month
semana = pd.DataFrame(df['dia_da_semana'].value_counts())
semana
semana.plot(title = 'Distribuição dos Homicídios por dia da Semana' , kind='barh',color = 'red', use_index=True, legend=True, sort_columns=True)
mes = pd.DataFrame(df['mes'].value_counts())
mes.head()
mes.plot(title = 'Distribuição dos Homicídios por dia mês' , kind='barh',color = 'blue', use_index=True, legend=True, sort_columns=True)
top_10_horario_crimes = pd.DataFrame(df['HORA_OCORRENCIA_BO'].value_counts().head(10))
top_10_horario_crimes
top_10_horario_crimes.plot(title = 'Top 10 de horarios de maior ocorrencia de crimes' , kind='barh',color = 'purple', use_index=True, legend=False, sort_columns=True)
top_10_logradouros_mais_homicidios = pd.DataFrame(df['LOGRADOURO'].value_counts().head(10))
top_10_logradouros_mais_homicidios
top_10_logradouros_mais_homicidios.plot(title = 'Top 10 logradouros com maior ocorrencia de homicídios' , kind='bar',color = 'green', use_index=True, legend=False, sort_columns=True)
df.info()
motivacao_homicidios = pd.DataFrame(df['DESDOBRAMENTO'].value_counts())
motivacao_homicidios.plot(title = 'Motivação dos homicídios' , kind='bar',color = 'pink', use_index=True, legend=True)
top_10_idades_vitimas = pd.DataFrame(df['IDADE_PESSOA'].value_counts().head(10))
top_10_idades_vitimas
top_10_idades_vitimas.index.name = 'Idades'
# +
top_10_idades_vitimas.plot(title = 'As idades mais comuns das vítimas' , kind='bar',color = 'orange', use_index=True, legend=False)
top_10_idades_vitimas.index.name = 'Idades'
# +
import pandas as pd
values = [[1,2], [2,5]]
df = pd.DataFrame(values, columns=['Type A', 'Type B'], index=['Index 1','Index 2'])
df.columns.name = 'Type'
df.index.name = 'Index'
df.plot(lw=2, colormap='jet', marker='.', markersize=10, title='Video streaming dropout by category')
# -
sexo = pd.DataFrame(df['SEXO_PESSOA'].value_counts())
sexo
nd : str
‘line’ : line plot (default)
‘bar’ : vertical bar plot
‘barh’ : horizontal bar plot
‘hist’ : histogram
‘box’ : boxplot
‘kde’ : Kernel Density Estimation plot
‘density’ : same as ‘kde’
‘area’ : area plot
‘pie’ : pie plot
‘scatter’ : scatter plot
‘hexbin’ : hexbin plot
sexo.plot(title = 'Sexo das vítimas' , kind='pie', use_index=True, legend=False , subplots = True)
delegacia = pd.DataFrame(df['NOME_DELEGACIA'].value_counts())
delegacia.head()
top10_delegacias_homicidios = delegacia.head(10)
top10_delegacias_homicidios
top10_delegacias_homicidios.plot(title = 'As 10 delegacias com maior registro de homicidios' , kind='pie', use_index=True, legend=False , subplots = True)
# ### Agora, os estudos usando Time Series
df['NUM_BO'].plot(legend=True,figsize=(10,4))
timestamp = df[['DATA_OCORRENCIA_BO','HORA_OCORRENCIA_BO']]
# +
#allDays = pd.DataFrame(df.set_index('timestamp').groupby(pd.TimeGrouper('1H')).sum())
# -
timestamp.head()
timestamp['DATA_OCORRENCIA_BO'] = timestamp['DATA_OCORRENCIA_BO'].astype(str)
timestamp.info()
datetime = timestamp['DATA_OCORRENCIA_BO'] + ' ' + timestamp['HORA_OCORRENCIA_BO']
datetime = pd.DataFrame(datetime)
datetime.head()
datetime = pd.to_datetime(datetime[0])
datetime.head()
datetime = pd.DataFrame(datetime)
datetime = datetime.rename(columns={0: 'data_e_hora'})
datetime
df = pd.merge(df, datetime, left_index=True, right_index=True, how='outer')
df.info()
df
# ### Abaixo, só confirmando o datetime criado!
df[['HORA_OCORRENCIA_BO','data_e_hora']].head()
df.set_index(keys = 'indices', append=True, inplace=True)
df.sort(columns='DATA_OCORRENCIA_BO', axis=0, ascending=True, inplace=False)
df.isnull().sum()
df.info()
def count(num):
if num > 0:
return 1
else:
return 'nan'
df['quantidade'] = df['mes'].apply(count)
df
df_agrupado_por_dia = pd.DataFrame(df.set_index('data_e_hora').groupby(pd.TimeGrouper('24H')).sum())
df_agrupado_por_dia
df['data_e_hora'].max()
df_agrupado_por_dia['quantidade'].plot(legend=True,figsize=(16,8))
| Sao_Paulo_Homicidios_Dolosos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle
modelo = open('modelo_consumo_cerveja','rb')
lm_new = pickle.load(modelo)
modelo.close()
temp_max = 30.5
chuva = 12.2
fds = 0
entrada = [[temp_max, chuva, fds]]
print('{0:.2f} litros'.format(lm_new.predict(entrada)[0]))
# -
| reg-linear/Projeto/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="t_gnhLD0HvhU" toc-hr-collapsed=true
# ## Summary
#
# - *hidden_size = 162*.
# - *num_heads = 9*.
# - *dropout = 0*.
# - N=16.
# - Add node and edge features (node features as 81-dim. embedding in `hidden_size`-dim space).
# - Edgeconv: embed x and edge to half their size and keep row x only.
# - Embed attention with `model_size == 63` and add `outbed_x` parameter to `MyAttention`.
#
# ----
# -
# ## Install dependencies (Google Colab only)
try:
import google.colab
GOOGLE_COLAB = True
except ImportError:
GOOGLE_COLAB = False
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="hIbvKDk_HvhX" outputId="349cb6d3-ffbc-4550-d9dd-dd6288ed5356"
if GOOGLE_COLAB:
# !pip install --upgrade torch-scatter
# !pip install --upgrade torch-sparse
# !pip install --upgrade torch-cluster
# !pip install --upgrade torch-spline-conv
# !pip install torch-geometric
# + colab={"base_uri": "https://localhost:8080/", "height": 203} colab_type="code" id="tw-y9MmIHvha" outputId="0666864a-5017-40a1-a77c-d23388eb0b1c"
if GOOGLE_COLAB:
# !pip install git+https://gitlab.com/ostrokach/proteinsolver.git
# + [markdown] colab_type="text" id="eoBMUoW2Hvhp" toc-hr-collapsed=true
# ## Imports
# + colab={} colab_type="code" id="TbKxMUZWHvhq"
import atexit
import csv
import itertools
import tempfile
import time
import uuid
import warnings
from collections import deque
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pyarrow
import torch
import torch.nn as nn
import torch.utils.tensorboard
from torch import optim
from torch_geometric.data import DataLoader
# + colab={"base_uri": "https://localhost:8080/", "height": 334} colab_type="code" id="rczy7pPiHvhs" outputId="8e4673d7-4cc7-4f8e-ffc4-de94302e5fab"
import proteinsolver
import proteinsolver.datasets
# + colab={} colab_type="code" id="4fFnAyUOHvhv"
# %load_ext autoreload
# %autoreload 2
# -
assert torch.cuda.is_available()
# ## Parameters
device = torch.device("cuda:0")
DATA_ROOT = Path(tempfile.gettempdir())
DATA_ROOT = next(Path("/localscratch/").glob("strokach.*")).joinpath("sudoku")
DATA_ROOT.mkdir(exist_ok=True)
DATA_ROOT
UNIQUE_ID = "5f2949e5"
CONTINUE_PREVIOUS = True
try:
NOTEBOOK_PATH
UNIQUE_PATH
except NameError:
NOTEBOOK_PATH = Path("sudoku_train").resolve()
NOTEBOOK_PATH.mkdir(exist_ok=True)
if UNIQUE_ID is None:
UNIQUE_ID = uuid.uuid4().hex[:8]
exist_ok = False
else:
exist_ok = True
UNIQUE_PATH = NOTEBOOK_PATH.joinpath(UNIQUE_ID)
UNIQUE_PATH.mkdir(exist_ok=exist_ok)
NOTEBOOK_PATH, UNIQUE_PATH
DATAPKG_DATA_DIR = Path(f"~/datapkg_output_dir").expanduser().resolve()
DATAPKG_DATA_DIR
proteinsolver.settings.data_url = DATAPKG_DATA_DIR.as_posix()
proteinsolver.settings.data_url
# ## Datasets
datasets = {}
# ### `SudokuDataset`
# + colab={} colab_type="code" id="ZP3bLDO9PgT9"
for i in range(10):
dataset_name = f"sudoku_train_{i}"
datasets[dataset_name] = proteinsolver.datasets.SudokuDataset4(
root=DATA_ROOT.joinpath(dataset_name), subset=f"train_{i}"
)
# -
datasets["sudoku_valid_0"] = proteinsolver.datasets.SudokuDataset4(
root=DATA_ROOT.joinpath("sudoku_valid_0"), subset=f"valid_0"
)
datasets["sudoku_valid_old"] = proteinsolver.datasets.SudokuDataset2(
root=DATA_ROOT.joinpath("sudoku_valid_old"),
data_url=DATAPKG_DATA_DIR.joinpath(
"deep-protein-gen", "sudoku", "sudoku_valid.csv.gz"
).as_posix(),
)
# + [markdown] colab_type="text" id="e3kWuRaxr89h"
# # Models
# + colab={} colab_type="code" id="tNCaSlrFDGA6"
# %%file {UNIQUE_PATH}/model.py
import copy
import tempfile
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.nn.modules.container import ModuleList
from torch_geometric.nn.inits import reset
from torch_geometric.utils import (
add_self_loops,
remove_self_loops,
scatter_,
to_dense_adj,
to_dense_batch,
)
class EdgeConvMod(torch.nn.Module):
def __init__(self, nn, aggr="max"):
super().__init__()
self.nn = nn
self.aggr = aggr
self.reset_parameters()
def reset_parameters(self):
reset(self.nn)
def forward(self, x, edge_index, edge_attr=None):
""""""
row, col = edge_index
x = x.unsqueeze(-1) if x.dim() == 1 else x
# TODO: Try -x[col] instead of x[col] - x[row]
if edge_attr is None:
out = torch.cat([x[row], x[col]], dim=-1)
else:
out = torch.cat([x[row], x[col], edge_attr], dim=-1)
out = self.nn(out)
x = scatter_(self.aggr, out, row, dim_size=x.size(0))
return x, out
def __repr__(self):
return "{}(nn={})".format(self.__class__.__name__, self.nn)
class EdgeConvBatch(nn.Module):
def __init__(self, gnn, hidden_size, batch_norm=True, dropout=0.2):
super().__init__()
self.gnn = gnn
x_post_modules = []
edge_attr_post_modules = []
if batch_norm is not None:
x_post_modules.append(nn.LayerNorm(hidden_size))
edge_attr_post_modules.append(nn.LayerNorm(hidden_size))
if dropout:
x_post_modules.append(nn.Dropout(dropout))
edge_attr_post_modules.append(nn.Dropout(dropout))
self.x_postprocess = nn.Sequential(*x_post_modules)
self.edge_attr_postprocess = nn.Sequential(*edge_attr_post_modules)
def forward(self, x, edge_index, edge_attr=None):
x, edge_attr = self.gnn(x, edge_index, edge_attr)
x = self.x_postprocess(x)
edge_attr = self.edge_attr_postprocess(edge_attr)
return x, edge_attr
def get_graph_conv_layer(input_size, hidden_size, output_size):
mlp = nn.Sequential(
#
nn.Linear(input_size, hidden_size),
nn.ReLU(),
nn.Linear(hidden_size, output_size),
)
gnn = EdgeConvMod(nn=mlp, aggr="add")
graph_conv = EdgeConvBatch(gnn, output_size, batch_norm=True, dropout=0.2)
return graph_conv
class MyEdgeConv(torch.nn.Module):
def __init__(self, hidden_size):
super().__init__()
self.embed_x = nn.Linear(hidden_size, hidden_size // 2)
self.embed_edge = nn.Linear(hidden_size, hidden_size // 2)
self.nn = nn.Sequential(
#
nn.Linear(hidden_size, hidden_size * 2),
nn.ReLU(),
nn.Linear(hidden_size * 2, hidden_size),
)
self.reset_parameters()
def reset_parameters(self):
reset(self.nn)
def forward(self, x, edge_index, edge_attr=None):
""""""
row, col = edge_index
x = x.unsqueeze(-1) if x.dim() == 1 else x
x_in = self.embed_x(x)
edge_attr_in = self.embed_edge(edge_attr)
x_edge_attr_in = torch.cat([x_in[row], edge_attr_in], dim=-1)
edge_attr_out = self.nn(x_edge_attr_in)
# if edge_attr is None:
# out = torch.cat([x[row], x[col]], dim=-1)
# else:
# out = torch.cat([x[row], x[col], edge_attr], dim=-1)
# edge_attr_out = self.nn(out)
return edge_attr_out
def __repr__(self):
return "{}(nn={})".format(self.__class__.__name__, self.nn)
class MyAttn(torch.nn.Module):
def __init__(self, hidden_size):
super().__init__()
model_size = 63
self.embed_x = nn.Linear(hidden_size, model_size)
self.outbed_x = nn.Linear(model_size, hidden_size)
self.attn = MultiheadAttention(
embed_dim=model_size,
kdim=hidden_size,
vdim=hidden_size,
num_heads=9,
dropout=0,
bias=True,
)
self.reset_parameters()
def reset_parameters(self):
reset(self.attn)
def forward(self, x, edge_index, edge_attr, batch):
""""""
query = self.embed_x(x).unsqueeze(0)
key = to_dense_adj(edge_index, batch=batch, edge_attr=edge_attr).squeeze(0)
adjacency = to_dense_adj(edge_index, batch=batch).squeeze(0)
key_padding_mask = adjacency == 0
key_padding_mask[torch.eye(key_padding_mask.size(0)).to(torch.bool)] = 0
# attn_mask = torch.zeros_like(key)
# attn_mask[mask] = -float("inf")
x_out, _ = self.attn(query, key, key, key_padding_mask=key_padding_mask)
# x_out = torch.where(torch.isnan(x_out), torch.zeros_like(x_out), x_out)
x_out = x_out.squeeze(0)
x_out = self.outbed_x(x_out)
assert (x_out == x_out).all().item()
assert x.shape == x_out.shape, (x.shape, x_out.shape)
return x_out
def __repr__(self):
return "{}(nn={})".format(self.__class__.__name__, self.nn)
class Net(nn.Module):
def __init__(
self, x_input_size, adj_input_size, hidden_size, output_size, batch_size=1
):
super().__init__()
x_labels = torch.arange(81, dtype=torch.long)
self.register_buffer("x_labels", x_labels)
self.register_buffer("batch", torch.zeros(10000, dtype=torch.int64))
self.embed_x = nn.Sequential(nn.Embedding(x_input_size, hidden_size), nn.ReLU())
self.embed_x_labels = nn.Sequential(nn.Embedding(81, hidden_size), nn.ReLU())
self.finalize_x = nn.Sequential(
nn.Linear(hidden_size * 2, hidden_size), nn.LayerNorm(hidden_size)
)
if adj_input_size:
self.embed_adj = nn.Sequential(
nn.Linear(adj_input_size, hidden_size),
nn.ReLU(),
nn.Linear(hidden_size, hidden_size),
nn.LayerNorm(hidden_size),
# nn.ELU(),
)
else:
self.embed_adj = None
N = 16
self.N = N
norm = nn.LayerNorm(hidden_size)
self.x_norms_0 = _get_clones(norm, N)
self.adj_norms_0 = _get_clones(norm, N)
self.x_norms_1 = _get_clones(norm, N)
self.adj_norms_1 = _get_clones(norm, N)
edge_conv = MyEdgeConv(hidden_size)
self.edge_convs = _get_clones(edge_conv, N)
attn = MyAttn(hidden_size)
self.attns = _get_clones(attn, N)
self.dropout = nn.Dropout(0.1)
self.linear_out = nn.Linear(hidden_size, output_size)
def forward(self, x, edge_index, edge_attr):
x = self.embed_x(x)
x_labels = self.embed_x_labels(self.x_labels)
x_labels = x_labels.repeat(x.size(0) // x_labels.size(0), 1)
x = torch.cat([x, x_labels], dim=1)
x = self.finalize_x(x)
edge_attr = self.embed_adj(edge_attr)
for i in range(self.N):
edge_attr_out = self.edge_convs[i](x, edge_index, edge_attr)
edge_attr = edge_attr + self.dropout(edge_attr_out)
edge_attr = self.adj_norms_1[i](edge_attr)
x_out = self.attns[i](
x,
edge_index,
self.adj_norms_0[i](edge_attr_out),
self.batch[: x.size(0)],
)
x = x + self.dropout(x_out)
x = self.x_norms_1[i](x)
x = self.linear_out(x)
return x
class MultiheadAttention(nn.Module):
def __init__(
self,
embed_dim,
num_heads,
dropout=0.0,
bias=True,
add_bias_kv=False,
add_zero_attn=False,
kdim=None,
vdim=None,
):
super().__init__()
self.embed_dim = embed_dim
self.kdim = kdim if kdim is not None else embed_dim
self.vdim = vdim if vdim is not None else embed_dim
self._qkv_same_embed_dim = self.kdim == embed_dim and self.vdim == embed_dim
self.num_heads = num_heads
self.dropout = dropout
self.head_dim = embed_dim // num_heads
assert (
self.head_dim * num_heads == self.embed_dim
), "embed_dim must be divisible by num_heads"
if self._qkv_same_embed_dim is False:
self.q_proj_weight = nn.Parameter(torch.Tensor(embed_dim, embed_dim))
self.k_proj_weight = nn.Parameter(torch.Tensor(embed_dim, self.kdim))
self.v_proj_weight = nn.Parameter(torch.Tensor(embed_dim, self.vdim))
else:
self.in_proj_weight = nn.Parameter(torch.empty(3 * embed_dim, embed_dim))
if bias:
self.in_proj_bias = nn.Parameter(torch.empty(3 * embed_dim))
else:
self.register_parameter("in_proj_bias", None)
self.out_proj = nn.Linear(embed_dim, embed_dim, bias=bias)
if add_bias_kv:
self.bias_k = nn.Parameter(torch.empty(1, 1, embed_dim))
self.bias_v = nn.Parameter(torch.empty(1, 1, embed_dim))
else:
self.bias_k = self.bias_v = None
self.add_zero_attn = add_zero_attn
self._reset_parameters()
def _reset_parameters(self):
if self._qkv_same_embed_dim:
nn.init.xavier_uniform_(self.in_proj_weight)
else:
nn.init.xavier_uniform_(self.q_proj_weight)
nn.init.xavier_uniform_(self.k_proj_weight)
nn.init.xavier_uniform_(self.v_proj_weight)
if self.in_proj_bias is not None:
nn.init.constant_(self.in_proj_bias, 0.0)
nn.init.constant_(self.out_proj.bias, 0.0)
if self.bias_k is not None:
nn.init.xavier_normal_(self.bias_k)
if self.bias_v is not None:
nn.init.xavier_normal_(self.bias_v)
def forward(
self,
query,
key,
value,
key_padding_mask=None,
need_weights=True,
attn_mask=None,
):
if hasattr(self, "_qkv_same_embed_dim") and self._qkv_same_embed_dim is False:
return F.multi_head_attention_forward(
query,
key,
value,
self.embed_dim,
self.num_heads,
None, # set self.in_proj_weight = None
self.in_proj_bias,
self.bias_k,
self.bias_v,
self.add_zero_attn,
self.dropout,
self.out_proj.weight,
self.out_proj.bias,
training=self.training,
key_padding_mask=key_padding_mask,
need_weights=need_weights,
attn_mask=attn_mask,
use_separate_proj_weight=True,
q_proj_weight=self.q_proj_weight,
k_proj_weight=self.k_proj_weight,
v_proj_weight=self.v_proj_weight,
)
else:
if not hasattr(self, "_qkv_same_embed_dim"):
warnings.warn(
"A new version of MultiheadAttention module has been implemented. \
Please re-train your model with the new module",
UserWarning,
)
return F.multi_head_attention_forward(
query,
key,
value,
self.embed_dim,
self.num_heads,
self.in_proj_weight,
self.in_proj_bias,
self.bias_k,
self.bias_v,
self.add_zero_attn,
self.dropout,
self.out_proj.weight,
self.out_proj.bias,
training=self.training,
key_padding_mask=key_padding_mask,
need_weights=need_weights,
attn_mask=attn_mask,
)
def _get_clones(module, N):
return ModuleList([copy.deepcopy(module) for i in range(N)])
# -
# %run {UNIQUE_PATH}/model.py
# + colab={} colab_type="code" id="tNCaSlrFDGA6"
# %%file {UNIQUE_PATH}/stats.py
import atexit
import csv
import time
import warnings
import numpy as np
class Stats:
epoch: int
step: int
batch_size: int
echo: bool
total_loss: float
num_correct_preds: int
num_preds: int
num_correct_preds_missing: int
num_preds_missing: int
num_correct_preds_missing_valid: int
num_preds_missing_valid: int
num_correct_preds_missing_valid_old: int
num_preds_missing_valid_old: int
start_time: float
def __init__(
self, *, epoch=0, step=0, batch_size=1, filename=None, echo=True, tb_writer=None
):
self.epoch = epoch
self.step = step
self.batch_size = batch_size
self.echo = echo
self.tb_writer = tb_writer
self.prev = {}
self.init_parameters()
if filename:
self.filehandle = open(filename, "wt", newline="")
self.writer = csv.DictWriter(
self.filehandle, list(self.stats.keys()), dialect="unix"
)
atexit.register(self.filehandle.close)
else:
self.filehandle = None
self.writer = None
def init_parameters(self):
self.num_steps = 0
self.total_loss = 0
self.num_correct_preds = 0
self.num_preds = 0
self.num_correct_preds_missing = 0
self.num_preds_missing = 0
self.num_correct_preds_missing_valid = 0
self.num_preds_missing_valid = 0
self.num_correct_preds_missing_valid_old = 0
self.num_preds_missing_valid_old = 0
self.start_time = time.perf_counter()
def reset_parameters(self):
self.prev = self.stats
self.init_parameters()
@property
def header(self):
return "".join(to_fixed_width(self.stats.keys()))
@property
def row(self):
return "".join(to_fixed_width(self.stats.values(), 4))
@property
def stats(self):
with warnings.catch_warnings():
warnings.simplefilter("ignore")
return {
"epoch": self.epoch,
"step": self.step,
"datapoint": self.datapoint,
"avg_loss": np.float64(1) * self.total_loss / self.num_steps,
"accuracy": np.float64(1) * self.num_correct_preds / self.num_preds,
"accuracy_m": np.float64(1)
* self.num_correct_preds_missing
/ self.num_preds_missing,
"accuracy_mv": self.accuracy_mv,
"accuracy_mv_old": self.accuracy_mv_old,
"time_elapsed": time.perf_counter() - self.start_time,
}
@property
def accuracy_mv(self):
return (
np.float64(1)
* self.num_correct_preds_missing_valid
/ self.num_preds_missing_valid
)
@property
def accuracy_mv_old(self):
return (
np.float64(1)
* self.num_correct_preds_missing_valid_old
/ self.num_preds_missing_valid_old
)
@property
def datapoint(self):
return self.step * self.batch_size
def write_header(self):
if self.echo:
print(self.header)
if self.writer is not None:
self.writer.writeheader()
def write_row(self):
if self.echo:
print(self.row, end="\r")
if self.writer is not None:
self.writer.writerow(self.stats)
self.filehandle.flush()
if self.tb_writer is not None:
stats = self.stats
datapoint = stats.pop("datapoint")
for key, value in stats.items():
self.tb_writer.add_scalar(key, value, datapoint)
self.tb_writer.flush()
def to_fixed_width(lst, precision=None):
lst = [round(l, precision) if isinstance(l, float) else l for l in lst]
return [f"{l: <18}" for l in lst]
# -
# %run {UNIQUE_PATH}/stats.py
# +
def get_stats_on_missing(x, y, output):
mask = (x == 9).squeeze()
if not mask.any():
return 0.0, 0.0
output_missing = output[mask]
_, predicted_missing = torch.max(output_missing.data, 1)
return (predicted_missing == y[mask]).sum().item(), len(predicted_missing)
from contextlib import contextmanager
@contextmanager
def eval_net(net: nn.Module):
training = net.training
try:
net.train(False)
yield
finally:
net.train(training)
# +
batch_size = 6
info_size = 200
hidden_size = 162
checkpoint_size = 100_000
batch_size, info_size, hidden_size
# -
tensorboard_path = NOTEBOOK_PATH.joinpath("runs", UNIQUE_PATH.name)
tensorboard_path.mkdir(exist_ok=True)
tensorboard_path
# +
last_epoch = None
last_step = None
last_datapoint = None
last_state_file = None
if CONTINUE_PREVIOUS:
for path in UNIQUE_PATH.glob("*.state"):
e, s, d, amv = path.name.split("-")
datapoint = int(d.strip("d"))
if last_datapoint is None or datapoint >= last_datapoint:
last_datapoint = datapoint
last_epoch = int(e.strip("e"))
last_step = int(s.strip("s"))
last_state_file = path
last_epoch, last_step, last_datapoint, last_state_file
# + colab={"base_uri": "https://localhost:8080/", "height": 6380} colab_type="code" id="azxPKTSKwcE8" outputId="09ed9b8b-25b9-49ff-8d02-83a4ec0ecba1"
net = Net(
x_input_size=13,
adj_input_size=3,
hidden_size=hidden_size,
output_size=9,
batch_size=batch_size,
).to(device)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(net.parameters(), lr=0.0001)
scheduler = optim.lr_scheduler.ReduceLROnPlateau(optimizer, "max", verbose=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 6380} colab_type="code" id="azxPKTSKwcE8" outputId="09ed9b8b-25b9-49ff-8d02-83a4ec0ecba1"
if CONTINUE_PREVIOUS:
net.load_state_dict(torch.load(last_state_file))
print("Loaded network state file.")
# + colab={"base_uri": "https://localhost:8080/", "height": 6380} colab_type="code" id="azxPKTSKwcE8" outputId="09ed9b8b-25b9-49ff-8d02-83a4ec0ecba1"
stats = Stats(
epoch=last_epoch if CONTINUE_PREVIOUS else 0,
step=last_step if CONTINUE_PREVIOUS else 0,
batch_size=batch_size,
filename=UNIQUE_PATH.joinpath("training.log"),
echo=True,
tb_writer=torch.utils.tensorboard.writer.SummaryWriter(
log_dir=tensorboard_path.with_suffix(f".xxx"),
purge_step=(last_datapoint if CONTINUE_PREVIOUS else None),
),
)
stats.write_header()
# + colab={"base_uri": "https://localhost:8080/", "height": 6380} colab_type="code" id="azxPKTSKwcE8" outputId="09ed9b8b-25b9-49ff-8d02-83a4ec0ecba1"
datasets[f"sudoku_valid_0"].reset()
valid_0_data = list(itertools.islice(datasets[f"sudoku_valid_0"], 300))
valid_old_data = list(itertools.islice(datasets[f"sudoku_valid_old"], 300))
tmp_data = valid_0_data[0].to(device)
edge_index = tmp_data.edge_index
edge_attr = tmp_data.edge_attr
net = net.train()
for epoch in range(stats.epoch, 100_000):
stats.epoch = epoch
train_dataloader = DataLoader(
datasets[f"sudoku_train_{epoch}"],
shuffle=False,
num_workers=1,
batch_size=batch_size,
drop_last=True,
)
for data in train_dataloader:
stats.step += 1
if CONTINUE_PREVIOUS and stats.step <= last_step:
continue
optimizer.zero_grad()
data = data.to(device)
output = net(data.x, data.edge_index, data.edge_attr)
loss = criterion(output, data.y)
loss.backward()
stats.total_loss += loss.detach().item()
stats.num_steps += 1
# Accuracy for all
_, predicted = torch.max(output.data, 1)
stats.num_correct_preds += (predicted == data.y).sum().item()
stats.num_preds += len(predicted)
# Accuracy for missing only
num_correct, num_total = get_stats_on_missing(data.x, data.y, output)
stats.num_correct_preds_missing += num_correct
stats.num_preds_missing += num_total
optimizer.step()
if (stats.datapoint % info_size) < batch_size:
for j, data in enumerate(valid_0_data):
data = data.to(device)
with torch.no_grad() and eval_net(net):
output = net(data.x, data.edge_index, data.edge_attr)
num_correct, num_total = get_stats_on_missing(data.x, data.y, output)
stats.num_correct_preds_missing_valid += num_correct
stats.num_preds_missing_valid += num_total
for j, data in enumerate(valid_old_data):
data = data.to(device)
with torch.no_grad() and eval_net(net):
output = net(data.x, data.edge_index, edge_attr)
num_correct, num_total = get_stats_on_missing(data.x, data.y, output)
stats.num_correct_preds_missing_valid_old += num_correct
stats.num_preds_missing_valid_old += num_total
stats.write_row()
stats.reset_parameters()
if (stats.datapoint % checkpoint_size) < batch_size:
output_filename = (
f"e{stats.epoch}-s{stats.step}-d{stats.datapoint}"
f"-amv{str(round(stats.prev['accuracy_mv'], 4)).replace('.', '')}.state"
)
torch.save(net.state_dict(), UNIQUE_PATH.joinpath(output_filename))
scheduler.step(stats.prev["accuracy_mv"])
output_filename = (
f"e{stats.epoch}-s{stats.step}-d{stats.datapoint}"
f"-amv{str(round(stats.prev['accuracy_mv'], 4)).replace('.', '')}.state"
)
torch.save(net.state_dict(), UNIQUE_PATH.joinpath(output_filename))
# -
| notebooks/04_sudoku_train-5f2949e5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jorgejosht/daa_2021_1/blob/master/13enero.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="LqJ_YenThpX3"
class NodoArbol:
def __init__(self,dato, left=None, right=None):
self.dato=dato
self.left=left
self.right=right
# + id="Y0zwiSMpiZrk"
class BinarySearchTree:
def __init__(self):
self.__root=None
def insert(self,value):
if self.__root==None:
self.__root=NodoArbol(value,None,None)
else:
self.__insert_nodo__(self.__root,value)
def __insert_nodo__(self,nodo,value):
if nodo.dato==value:
pass
elif value<nodo.dato:
if nodo.left==None:
nodo.left=NodoArbol(value,None,None)
else:
self.__insert_nodo__(nodo.left,value)
else:
if nodo.right==None:
nodo.right=NodoArbol(value,None,None)
else:
return self.__busca_nodo(self.__root,value)
def __busca_nodo(self,nodo,value):
if nodo==None:
return None
elif nodo.dato==value:
return nodo.dato
elif value<nodo.dato:
return self.__busca_nodo(nodo.left,value)
else:
return self.__busca_nodo(nodo.right,value)
def transversal(self,format="inorden"):
if format=="inorden":
self.__recorrido_in(self.__root)
elif format=="preorden":
self.__recorrido_pre(self.__root)
elif format=="posorden":
self.__recorrido_pos(self.__root)
else:
print("Formato de recorrido no válido")
def __recorrido_pre(self,nodo):
if nodo!=None:
print(nodo.dato, end=",")
self.__recorrido_pre(nodo.left)
self.__recorrido_pre(nodo.right)
def __recorrido_in(self,nodo):
if nodo!=None:
self.__recorrido_in(nodo.left)
print(nodo.dato, end=",")
self.__recorrido_in(nodo.right)
def __recorrido_pos(self,nodo):
if nodo!=None:
self.__recorrido_pos(nodo.left)
self.__recorrido_pos(nodo.right)
print(nodo.dato, end=",")
# + id="haCGsZzUj8gd" colab={"base_uri": "https://localhost:8080/"} outputId="424477b0-941e-4c2a-dd65-ef9969148edf"
arbolb=BinarySearchTree()
arbolb.insert(50)
arbolb.insert(30)
arbolb.insert(20)
print("Recorrido preorden: ")
arbolb.transversal(format="preorden")
print("\nRecorrido inorden: ")
arbolb.transversal()
print("\nRecorrido posorden")
arbolb.transversal(format="posorden")
| 13enero.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Rinott's ranking and selection procedure
import numpy as np
import pandas as pd
import random as rd
df = pd.read_excel (r'rinott.xlsx')
#print (df)
# set parameters from file
Pstar = df.iloc[0,1]
n0 = df.iloc[1,1]
h = df.iloc[2,1]
deltastar = df.iloc[3,1]
# set data points
data = df.iloc[10:,:]
#data
def Rinott(Pstar, n0, h, deltastar, data):
nu = n0-1
h = h # already provided
# step (ii)
# define a frame that holds all samples
samples = np.empty([n0, data.shape[1]])
# collect samples
for i in np.arange(0, data.shape[1]-1, step = 1):
samples[:,i] = data.iloc[:n0,i+1]#data.iloc[:,i+1].sample(n=n0, random_state=1)
# step (iii)
# summary statistics calculations
stats = np.empty([data.shape[1]-1,4])
for i in np.arange(0, data.shape[1]-1, step = 1):
stats[i,0] = np.mean(samples[:,i])
stats[i,1] = np.var(samples[:,i], ddof = 1)
# Stage 2
# step (i)
for i in np.arange(0, data.shape[1]-1, step = 1):
stats[i,2] = max(n0, np.ceil(((h*np.sqrt(stats[i,1]))/deltastar)**2))
if stats[i,2] > n0:
sample_ext = data.iloc[:,i+1].sample(n = int(stats[i,2] - n0), random_state = 2)
# step (ii)
stats[i,3] = np.mean(np.concatenate((samples[:,i], sample_ext), axis = 0))
else:
stats[i,3] = np.mean(samples[:,i])
# step (iii)
istar = np.argmax(stats[:,3])+1
# step (iv)
print("Sample", istar, "is the rv associated with the largest mu of", round(max(stats[:,3]),3))
return(stats)
results = Rinott(Pstar, n0, h, deltastar, data)
results = np.round(results,3)
results = pd.DataFrame(results) # transform into data frame
results = results.rename(columns={0: "Ybar^0_i", 1: "S_i^2", 2: "n_i", 3: "Ybar_i"})
results = results.rename(index = {0: 1, 1: 2, 2: 3, 3: 4, 4: 5, 5: 6, 6: 7, 7: 8, 8: 9, 9: 10})
results.to_csv(index=True, path_or_buf = 'Rinott_results.csv')
results
| Rinotts ranking and selection procedure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def traverseDict(keyPtr, valList, searchVal):
global dictPath
print(valList)
for val in valList:
try:
if type(keyPtr[val]) == type([]):
for subVali in range(0,len(keyPtr[val])):
dictPath += "item {} ".format(subVali)
traverseDict(keyPtr[val][subVal], [], 0)
elif searchVal in keyPtr[val]:
return dictPath
else:
dictPath += val+" "
traverseDict(keyPtr[val], [v for k,v in keyPtr[val].items()], searchVal)
except KeyError:
dictPath = ""
continue
| resources/DataMining/trash.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cirq
# In this notebook, we will discuss the ideas behind the Variational-Quantum-Eigensolver. This algorithm relies on the variational principle and is used to find the ground state of a quantum system (and therefore, more generally speaking extracts the lowest eigenvalue of a hermitian matrix). However, contrary to the standard setting of computational physics, the wavefunction here is not parameterised on a classical computer but by a quantum circuit. We also note that many classical optimisation problems can be mapped onto quantum Hamiltonians, and then finding the ground state becomes equivalent to minimising the cost function.
#
# The quantum circuit is then used to estimate the expectation value of the energy, while the parameter optimisation is done classically. The main hope here is that, especially for many-body systems/ exponentially large matrices, the estimation of the energy can be done much more efficiently on a quantum computer than on a classical computer (wherein the setting of many-body physics one usually relies on Monte Carlo type estimations).
# We note that, in general, the problem is in QMA and, therefore, still hard for a quantum computer. Nevertheless, it is a promising idea that can also be combined with domain knowledge to chose a suitable trial wavefunction.
# We also note that this is an active research topic, and there are many details, which we will not cover to make this algorithm efficient.
#
# The intent of this notebook is instead to outline the idea behind the algorithm and show a simple working example.
#
# Reading: Quantum Sci. Technol. 3 030503, ncomms5213, arXiv:1407.7863
# Before describing the VQE let's very briefly review the variational principle in quantum mechanics.
#
# First, let $\mathcal{H}$ be a Hilbertspace (on a qubit system this will always be finite dimensional but in principle it doesn't have to be), $|\psi\rangle$ a normalised state $\in \mathcal{H}$ and $H$ a hermitian operator (in our case it's simply a hermitian matrix) over $\mathcal{H}$. We note that using the spectral theorem we can write $H = \sum_i E_i | i\rangle \langle i|$ (where we ordered the $E_i$ in ascending order)
# It is then easy to see that
# \begin{equation}
# \langle\psi| H |\psi \rangle = \sum_i E_i |\langle i|\psi \rangle|^2 \geq \sum_i E_0 |\langle i|\psi \rangle|^2 = E_0.
# \end{equation}
# This simply means that any expectation value of $H$ is an upper bound of the groundstate energy. The idea of the variational ansatz is now simple. One parameterises the wave-function by a set of paramters $\theta$ and minimises the resulting expecation value $\langle\psi(\theta)| H |\psi(\theta) \rangle$. Depending on the how well the wavefunction is chosen the result will be close to the true groundstate of the system. In fact for a finite dimensional Hilbertspace we could in theory span the whole Hilbertspace with a finite number of $\theta$ and then find the true groundstate, in practice this is of course untracable for larger systems.
# At this stage we can already see that domain knowledge can be a great benefit when chosing the trial wavefunction.
# \begin{theorem}
# The VQE algorithm works as follows
#
# 1. Prepare $|\psi(\theta)\rangle$ (or more generally $\rho(\theta)$)
# 2. Meassure $\langle H\rangle(\theta) = E(\theta)$.
# 3. Use a classical optimisation scheme to update $\theta$
# 4. Iterate 1-3 until desired convergence is achieved
# \end{theorem}
# We can directly see that this type of algorithm is extremly general and there are many possible ways to realise such a scheme. One does not even have to use a gate-based quantum computer for this.
#
# In the following we will go through a simple example to demonstrate how a simplified implementation could look like. This implementation is far from optimal but serves as a concrete example to illustrate the concept. We also note that the exact details of an efficient implementation of such an algorithm is an active research field and will depend on the hardware at hand.
# Before finally turning to the example we want to quickly note, that the tensor product of Pauli operators spans the space of Hermitian matricies and it is therfore in principle enough to restric ourselves to Hamiltonians made up of tensor products of Pauli operators. In quantum chemistry for example this mapping can be made explicit by a Jordan-Wigner (or Bravyi−Kitaev) transformation.
#
# Let's consider the two qubit Hamiltonian
# \begin{equation}
# H = X_1 + 0.1Z_1 - Z_2 + 0.5 X_1Z_2 + 0.3Z_1Y_2
# \end{equation}
# Before we get into the details on how to perform 2.
# Let's think about a suitable variational Ansatz.
# Here we already see two competing principles.
#
# On the one hand we want our Ansatz to be simple (meaning as few parameters as possible and keeping the circuit depth as shallow as possible, while also respecting the current hardware limitations) and on the other hand we want it to ideally span the Hilbert space or at least the relevant part of the Hilbert space. Again the optimal trade-off can vary from problem to problem and it is a priori far from trivial to chose a good variational Ansatz.
#
#
#
# \begin{remark}
# For the example at hand it we recall that state of a two-level system can be discribed as by two real numbers (since only the relative phase has a physical meaning and the state is normalised), one such choice (and perhaps the most common one) is the position on the Bloch sphere, which is given by $\phi, \theta$.
# In order to make use of the existing gates another approach is taken. We recall that any unitary transformation of a qubit state can be thought of as a rotation on the Bloch ssphere and we therefore have to consider only $SU(2)$. This can of course be generalised to $n$ qubits, where we would now have to consider $SU(2^n)$, to parametrise any unitary transformation on the Bloch sphere, realising that $\mathrm{dim}[SU(2^n)] =4^n-1$ we conclude that we need at least $4^n-1$ parameters, if we only have access to rotational and $CNOT$ gates. We also recall that using the Euler represntation of $SU(2)$ any element $U\in SU(2)$ can be written as $U= R_z(\phi)R_y(\theta)R_z(\psi)$.
# \end{remark}
# Let's now parametrise some circuits. In order to gain some more insight we will implement two-qubit circuits with increasing complexity, and one universal circuit
#implement universal SU(2) gate
def U_single_qubit(q,phi,theta,psi):
yield cirq.rz(phi)(q)
yield cirq.ry(theta)(q)
yield cirq.rz(psi)(q)
# Using the KAK decomposition of $SU(4)$ we can implement a universal $SU(4)$ unitary operator and using it as our variational Ansatz
def universal_va(q1,q2,parameters):
yield U_single_qubit(q1,*parameters[0])
yield U_single_qubit(q2,*parameters[1])
yield cirq.CNOT(q1,q2)
yield U_single_qubit(q1,*parameters[2])
yield U_single_qubit(q2,*parameters[3])
yield cirq.CNOT(q2,q1)
yield U_single_qubit(q1,*parameters[4])
yield cirq.CNOT(q1,q2)
yield U_single_qubit(q1,*parameters[5])
yield U_single_qubit(q2,*parameters[6])
# This has circuit has 21 free parameters let's also initialise a simpler circuit, that can not span the whole space (feel free to implement your own ciruit)
def non_uni_va(q1,q2,parameters):
yield U_single_qubit(q1,*parameters[0])
yield U_single_qubit(q2,*parameters[1])
yield cirq.CNOT(q1,q2)
yield U_single_qubit(q1,*parameters[2])
yield U_single_qubit(q2,*parameters[3])
# Let's get to step 2. Here we have to meassure the Hamoltonian again this step can be done in many different way and the most efficient way of doing so can depend on the underlying structure of the problem.
#
# When considering a general Hamlitonian one can now devise two different straightforward strategies
# \begin{enumerate}
# \item Measure the state in the computational basis calculate $E(\theta)$, repeat until the result is coverged
# \item Perform direct Pauli measurments using the quantum circuit, calculate $E(\theta)$ from those individual measurments
# \end{enumerate}
# The first option is cloesly related to standard Monte Carlo methods, the only difference here is that the cirucit naturally implements the improtance sampling according to the wave-function. Here at each step the whole Energy is estimated. A major drawback of this approach is, that to achieve convergence we might have so sample many times and in principle we still have to perform large matrix multiplications on a classical computer
#
# The second option has the major advantage, is that we measure the expectation values in a suitable basis and therefore the outcome of ony term is just the coefficient multiplied by $\pm 1$, to caculate the expectation value we now have to nothing but averaging over many measurments without having to perfrom any matrix multiplications. Here in each run a single expectation value is measured for each term in the Hamiltonian (using clever grouping it is possible to measure more than one term).
#
# Usually the second method is to be preferred since it does not involve large matricies (which is usually one of the bottlenecks on a classical computer).
# In order to measure the values of a pauli-matrix in a suitable basis we now define the functions, which meassure in the X, Y basis. The Z-basis is the computational one.
#
# In order to meassure with respect to the X (Y) basis we just need to find the operator that transforms from the eigenbasis to the computational basis. For X this is simply the Hadamard gate and for Y it is $S^\dagger H$
# +
def x_measurment(q):
yield cirq.H(q)
def y_measurment(q):
Sdagger = (cirq.Z**(-0.5))
yield Sdagger(q)
yield cirq.H(q)
# -
# Let's see how many measurments we have to make to get one estimate for the energy.
# We see that we can rewrite
# \begin{equation}
# H = X_1 + 0.1Z_1 - Z_2 + 0.5 X_1Z_2 + 0.3Z_1Y_2 = [X_1(1+0.5Z_2) -Z_2]+Z_1(0.1+0.3Y_2) = H_1+H_2,
# \end{equation}
# which shows us that in fact two measurments are enough
# +
#this implementation is not really efficent and could be improved but it shows the individual steps nicely
def H_1_measurment(q1,q2):
yield x_measurment(q1)
yield cirq.measure(q1,q2, key='H_1')
def H_2_measurment(q1,q2):
yield y_measurment(q2)
yield cirq.measure(q1,q2, key='H_2')
# +
def E1_curcuit(va,parameters):
q1, q2 =cirq.LineQubit.range(2)
c = cirq.Circuit()
c.append(va(q1,q2,parameters))
c.append(H_1_measurment(q1,q2))
return c
def E2_curcuit(va,parameters):
q1, q2 =cirq.LineQubit.range(2)
c = cirq.Circuit()
c.append(va(q1,q2,parameters))
c.append(H_2_measurment(q1,q2))
return c
# +
simulator = cirq.Simulator()
def transform_to_eigenvalue(measurements):
return [[1 if i == 0 else -1 for i in j ] for j in measurements]
def H_estimator(parameters,N,va):
parameters = [[parameters[i+j] for j in range(3)] for i in range(7)]
c1 = E1_curcuit(va,parameters)
c2 = E2_curcuit(va,parameters)
r1 = simulator.run(c1, repetitions=N)
r2 = simulator.run(c2, repetitions=N)
m1 =transform_to_eigenvalue(r1.measurements['H_1'])
m2 =transform_to_eigenvalue(r2.measurements['H_2'])
mean = 0
for i in zip(m1,m2):
mean += i[0][0]*(1+1/2*i[0][1])-i[0][1]+i[1][0]*(0.1+0.3*i[1][1])
return mean/N
# -
# We have now implemented step 2. Lastly it we want to perform step 3. and then we can compare our results to those obtained by exact diagonalisation
# Since 3. relies on a classical optimisation algorithm there are many algorithms to chose from. Maybe a very naive guess would be gradient decent. In pratice (for reasons not that we will not discuss here see arXiv:1509.04279) this peforms relatively poorly in practice.
#
# Here we will use the Nelder-Mead simplex method as it is a derivative free optimisation method again see for arXiv:1509.04279 a discussion.
import scipy
uni_va_opt = scipy.optimize.minimize(H_estimator
, x0=2*np.pi*np.random.rand(21), args=(5000, universal_va)
,method='Nelder-Mead')
uni_va_opt.fun
non_uni_va_opt= scipy.optimize.minimize(H_estimator
, x0=2*np.pi*np.random.rand(12), args=(5000, non_uni_va)
,method='Nelder-Mead')
non_uni_va_opt.fun
# Lastly let's check our result against the result obtained using exact diagonalisation.
# As a reminder we want to diagonalise the Hamiltonian follwoing
# \begin{equation}
# H = X_1 + 0.1Z_1 - Z_2 + 0.5 X_1Z_2 + 0.3Z_1Y_2
# \end{equation}
##Exact diagonalisation
import numpy as np
X = np.array([[0,1],[1,0]])
Z = np.array([[1,0],[0,-1]])
Y = np.array([[0,-1j],[1j,0]])
I = np.array([[1,0],[0,1]])
H = np.kron(X,I)+0.1*np.kron(Z,I)-np.kron(I,Z)+0.5*np.kron(X,Z)+0.3*np.kron(Z,Y)
H
##exact result
np.min(np.linalg.eigh(H)[0])
# We see that both of our variational wavefunctions performed fairly well. Some of the error is also attributed to the noise of evaluating the expectation value. It also shows us that in most cases it is not neccessary to chose a universal approximator of the wavefunction.
| src/.ipynb_checkpoints/A1 VQE-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AI Automation for AI Fairness
#
# When AI models contribute to high-impact decisions such as whether or
# not someone gets a loan, we want them to be fair.
# Unfortunately, in current practice, AI models are often optimized
# primarily for accuracy, with little consideration for fairness. This
# notebook gives a hands-on example for how AI Automation can help build AI
# models that are both accurate and fair.
# This notebook is written for data scientists who have some familiarity
# with Python. No prior knowledge of AI Automation or AI Fairness is
# required, we will introduce the relevant concepts as we get to them.
#
# Bias in data leads to bias in models. AI models are increasingly
# consulted for consequential decisions about people, in domains
# including credit loans, hiring and retention, penal justice, medical,
# and more. Often, the model is trained from past decisions made by
# humans. If the decisions used for training were discriminatory, then
# your trained model will be too, unless you are careful. Being careful
# about bias is something you should do as a data scientist.
# Fortunately, you do not have to grapple with this issue alone. You
# can consult others about ethics. You can also ask yourself how your AI
# model may affect your (or your institution's) reputation. And
# ultimately, you must follow applicable laws and regulations.
#
# _AI Fairness_ can be measured via several metrics, and you need to
# select the appropriate metrics based on the circumstances. For
# illustration purposes, this notebook uses one particular fairness
# metric called _disparate impact_. Disparate impact is defined as the
# ratio of the rate of favorable outcome for the unprivileged group to
# that of the privileged group. To make this definition more concrete,
# consider the case where a favorable outcome means getting a loan, the
# unprivileged group is women, and the privileged group is men. Then if
# your AI model were to let women get a loan in 30% of the cases and men
# in 60% of the cases, the disparate impact would be 30% / 60% = 0.5,
# indicating a gender bias towards men. The ideal value for disparate
# impact is 1, and you could define fairness for this metric as a band
# around 1, e.g., from 0.8 to 1.2.
#
# To get the best performance out of your AI model, you must experiment
# with its configuration. This means searching a high-dimensional space
# where some options are categorical, some are continuous, and some are
# even conditional. No configuration is optimal for all domains let
# alone all metrics, and searching them all by hand is impossible. In
# fact, in a high-dimensional space, even exhaustively enumerating all
# the valid combinations soon becomes impractical. Fortunately, you can
# use tools to automate the search, thus making you more productive at
# finding good models quickly. These productivity and quality
# improvements become compounded when you have to do the search over.
#
# _AI Automation_ is a technology that assists data scientists in
# building AI models by automating some of the tedious steps. One AI
# automation technique is _algorithm selection_ , which automatically
# chooses among alternative algorithms for a particular task. Another AI
# automation technique is _hyperparameter tuning_ , which automatically
# configures the arguments of AI algorithms. You can use AI automation
# to optimize for a variety of metrics. This notebook shows you how to use AI
# automation to optimize for both accuracy and for fairness as measured
# by disparate impact.
#
# This [Jupyter](https://jupyter.org/)
# notebook uses the following open-source Python libraries.
# [AIF360](https://aif360.mybluemix.net/)
# is a collection of fairness metrics and bias mitigation algorithms.
# The [pandas](https://pandas.pydata.org/) and
# [scikit-learn](https://scikit-learn.org/) libraries support
# data analysis and machine learning with data structures and a
# comprehensive collection of AI algorithms.
# The [hyperopt](http://hyperopt.github.io/hyperopt/) library
# implements both algorithm selection and hyperparameter tuning for
# AI automation.
# And [Lale](https://github.com/IBM/lale) is a library for
# semi-automated data science; this notebook uses Lale as the backbone
# for putting the other libraries together.
#
# Our starting point is a dataset and a task. For illustration
# purposes, we picked [credit-g](https://www.openml.org/d/31), also
# known as the German Credit dataset. Each row describes a person
# using several features that may help evaluate them as a potential
# loan applicant. The task is to classify people into either
# good or bad credit risks. We load the version of the dataset from
# OpenML along with some fairness metadata.
from lale.lib.aif360 import fetch_creditg_df
all_X, all_y, fairness_info = fetch_creditg_df()
# To see what the dataset looks like, we can use off-the-shelf
# functionality from pandas for inspecting a few
# rows. The creditg dataset has a single label column, `class`, to be
# predicted as the outcome, which can be `good` or `bad`. Some of the
# feature columns are numbers, others are categoricals.
import pandas as pd
pd.options.display.max_columns = None
pd.concat([all_y, all_X], axis=1)
# The `fairness_info` is a JSON object that specifies metadata you
# need for measuring and mitigating fairness. The `favorable_labels`
# attribute indicates that when the `class` column contains the value
# `good`, that is considered a positive outcome.
# A _protected attribute_ is a feature that partitions the population
# into groups whose outcome should have parity.
# Values in the `personal_status` column that indicate that the indidual
# is `male` are considered privileged, and so are values in the
# `age` column that indicate that hte individual is between 26 and 100
# years old.
import lale.pretty_print
lale.pretty_print.ipython_display(fairness_info)
# A best practice for any machine-learning experiments is to split
# the data into a training set and a hold-out set. Doing so helps
# detect and prevent over-fitting. The fairness information induces
# groups in the dataset by outcomes and by privileged groups. We
# want the distribution of these groups to be similar for the training
# set and the holdout set. Therefore, we split the data in a
# stratified way.
from lale.lib.aif360 import fair_stratified_train_test_split
train_X, test_X, train_y, test_y = fair_stratified_train_test_split(
all_X, all_y, **fairness_info, test_size=0.33, random_state=42)
# Let's use the `disparate_impact` metric to measure how biased the
# training data and the test data are. At 0.75 and 0.73, they are far
# from the ideal value of 1.0.
from lale.lib.aif360 import disparate_impact
disparate_impact_scorer = disparate_impact(**fairness_info)
print("disparate impact of training data {:.2f}, test data {:.2f}".format(
disparate_impact_scorer.scoring(X=train_X, y_pred=train_y),
disparate_impact_scorer.scoring(X=test_X, y_pred=test_y)))
# Before we look at how to train a classifier that is optimized for both
# accuracy and disparate impact, we will set a baseline, by training a
# pipeline that is only optimized for accuracy. For this purpose, we
# import a few algorithms from scikit-learn and Lale:
# `Project` picks a subset of the feature columns,
# `OneHotEncoder` turns categoricals into numbers,
# `ConcatFeatures` combines sets of feature columns,
# and the three interpretable classifiers `LR`, `Tree`, and `KNN`
# make predictions.
from lale.lib.lale import Project
from sklearn.preprocessing import OneHotEncoder
from lale.lib.lale import ConcatFeatures
from sklearn.linear_model import LogisticRegression as LR
from sklearn.tree import DecisionTreeClassifier as Tree
from sklearn.neighbors import KNeighborsClassifier as KNN
# To use AI Automation, we need to define a _search space_ ,
# which is a set of possible machine learning pipelines and
# their associated hyperparameters. The following code
# uses Lale to define a search space.
import lale
lale.wrap_imported_operators()
prep_to_numbers = (
(Project(columns={"type": "string"}) >> OneHotEncoder(handle_unknown="ignore"))
& Project(columns={"type": "number"})
) >> ConcatFeatures
planned_orig = prep_to_numbers >> (LR | Tree | KNN)
planned_orig.visualize()
# The call to `wrap_imported_operators` augments the algorithms
# that were imported from scikit-learn with metadata about
# their hyperparameters.
# The Lale combinator `>>` pipes the output from one operator to
# the next one, creating a dataflow edge in the pipeline.
# The Lale combinator `&` enables multiple sub-pipelines to run
# on the same data.
# Here, `prep_to_numbers` projects string columns and one-hot encodes them;
# projects numeric columns and leaves them unmodified; and
# finally concatenates both sets of columns back together.
# The Lale combinator `|` indicates
# algorithmic choice: `(LR | Tree | KNN)` indicates that
# it is up to the AI Automation to decide which of the three different
# classifiers to use. Note that the classifiers are
# not configured
# with concrete hyperparameters, since those will be left for the
# AI automation to choose instead.
# The search space is encapsulated in the object `planned_orig`.
#
# We will use hyperopt to select the algorithms and to tune their
# hyperparameters. Lale provides a `Hyperopt` that
# turns a search space such as the one specified above into an
# optimization problem for hyperopt. After 10 trials, we get back
# the model that performed best for the default optimization
# objective, which is accuracy.
from lale.lib.lale import Hyperopt
best_estimator = planned_orig.auto_configure(
train_X, train_y, optimizer=Hyperopt, cv=3, max_evals=10)
best_estimator.visualize()
# As shown by the visualization, the search found a pipeline
# with a PCA transformer and an LR classifier.
# Inspecting the hyperparameters reveals which values
# worked best for the 10 trials on the dataset at hand.
best_estimator.pretty_print(ipython_display=True, show_imports=False)
# We can use the accuracy score metric from scikit-learn to measure
# how well the pipeline accomplishes the objective for which it
# was trained.
import sklearn.metrics
accuracy_scorer = sklearn.metrics.make_scorer(sklearn.metrics.accuracy_score)
print(f'accuracy {accuracy_scorer(best_estimator, test_X, test_y):.1%}')
# However, we would like our model to be not just accurate but also fair.
# We can use the same `disparate_impact_scorer` from before to evaluate
# the fairness of `best_estimator`.
print(f'disparate impact {disparate_impact_scorer(best_estimator, test_X, test_y):.2f}')
# The model is biased, which is no surprise, since it was trained
# from biased data. We would prefer a
# model that is much more fair. The AIF360 toolkit provides several
# algorithms for mitigating fairness problems. One of them is
# `DisparateImpactRemover`, which modifies the features that are
# not the protected attribute in such a way that it is hard to
# predict the protected attribute from them. We use a Lale version
# of `DisparateImpactRemover` that wraps the corresponding AIF360
# algorithm for AI Automation. This algorithm has a hyperparameter
# `repair_level` that we will tune with hyperparameter optimization.
from lale.lib.aif360 import DisparateImpactRemover
lale.pretty_print.ipython_display(
DisparateImpactRemover.hyperparam_schema('repair_level'))
# We compose the bias mitigation algorithm in a pipeline with
# a choice of classifiers as before.
# In the visualization, light blue indicates trainable operators
# and dark blue indicates that automation must make a choice before
# the operators can be trained. Compared to the earlier pipeline,
# we pass the data preparation sub-pipeline as an argument to `DisparateImpactRemover`,
# since that fairness mitigator needs numerical data to work on.
di_remover = DisparateImpactRemover(
**fairness_info, preparation=prep_to_numbers)
planned_fairer = di_remover >> (LR | Tree | KNN)
planned_fairer.visualize()
# Besides changing the planned pipeline to use a fairness mitigation
# operator, we should also change the optimization objective. We need
# a scoring function that blends accuracy with disparate impact.
# While you could define this scorer yourself, Lale also provides a
# pre-defined version.
from lale.lib.aif360 import accuracy_and_disparate_impact
combined_scorer = accuracy_and_disparate_impact(**fairness_info)
# Fairness metrics can be more unstable than accuracy, because they depend
# not just on the distribution of labels, but also on the distribution of
# privileged and unprivileged groups as defined by the protected attributes.
# In AI Automation, k-fold cross validation helps reduce overfitting.
# To get more stable results, we will stratify these k folds by both labels
# and groups.
from lale.lib.aif360 import FairStratifiedKFold
fair_cv = FairStratifiedKFold(**fairness_info, n_splits=3)
# Now, we have all the pieces in place to use AI Automation
# on our `planned_fairer` pipeline for both accuracy and
# disparate impact.
trained_fairer = planned_fairer.auto_configure(
train_X, train_y, optimizer=Hyperopt, cv=fair_cv,
max_evals=10, scoring=combined_scorer, best_score=1.0)
# As with any trained model, we can evaluate and visualize the result.
print(f'accuracy {accuracy_scorer(trained_fairer, test_X, test_y):.1%}')
print(f'disparate impact {disparate_impact_scorer(trained_fairer, test_X, test_y):.2f}')
trained_fairer.visualize()
# As the result demonstrates, the best model found by AI Automation
# has similar accuracy and better disparate impact than the one we saw
# before. Also, it has tuned the repair level and
# has picked and tuned a classifier.
trained_fairer.pretty_print(ipython_display=True, show_imports=False)
# These results may vary by dataset and search space.
#
# In summary, this blog post showed you how to use AI Automation
# from Lale, while incorporating a fairness mitigation technique
# into the pipeline and a fairness metric into the objective.
# Of course, this blog post only scratches the surface of what can
# be done with AI Automation and AI Fairness. We encourage you to
# check out the open-source projects Lale and AIF360 and use them
# to build your own fair and accurate models!
#
# - Lale: https://github.com/IBM/lale
# - AIF360: https://aif360.mybluemix.net/
# - API documentation: [lale.lib.aif360](https://lale.readthedocs.io/en/latest/modules/lale.lib.aif360.html#module-lale.lib.aif360)
| examples/demo_aif360.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > 原文地址 https://mp.weixin.qq.com/s?__biz=MzIzNzA4NDk3Nw==&mid=2457739380&idx=1&sn=122f15af3520857314199127ca79cad4&chksm=ff44882ac833013cb52d848aa03f2547f973d05572a0e90f3f68e662f573076507853691e222&mpshare=1&scene=24&srcid=&sharer_sharetime=1590505368566&sharer_shareid=316859bf78c7a4dcfe65351f82355327&key=4a6324d6ed20374b2ee3f6b714ba4c639a666cd163fba9d5e94aa754ed867c7dc268b148c37973ba3f2d81307162049ef360b855e17126c1be1067fa76d178362807a2501f080fdf9b34e15cbe16781d&ascene=14&uin=MTk3NzE3NDc2Nw%3D%3D&devicetype=Windows+10+x64&version=62090070&lang=zh_CN&exportkey=AQcehB42P8T%2BSFIJUqHMMDM%3D&pass_ticket=<PASSWORD>4Ozk<PASSWORD>gfs<PASSWORD>czer<PASSWORD>
#
#
# 本文于网络整理,版权归原作者所有,如来源信息有误或侵犯权益,请联系我删除。
#
# 自己写 Python 也有四五年了,一直是用自己的 “强迫症” 在维持自己代码的质量。都有去看 Google 的 Python 代码规范,对这几年的工作经验,做个简单的笔记,如果你也在学 Python,准备要学习 Python,希望这篇文章对你有用。
# ## 1. 首先
#
# - 建议 1、理解 Pythonic 概念—- 详见 Python 中的《Python 之禅》
#
# - 建议 2、编写 Pythonic 代码
#
# (1)避免不规范代码,比如只用大小写区分变量、使用容易混淆的变量名、害怕过长变量名等。有时候长的变量名会使代码更加具有可读性。
#
# (2)深入学习 Python 相关知识,比如语言特性、库特性等,比如 Python 演变过程等。深入学习一两个业内公认的 Pythonic 的代码库,比如 Flask 等。
#
# - 建议 3:理解 Python 与 C 的不同之处,比如缩进与 {},单引号双引号,三元操作符?, Switch-Case 语句等。
#
# - 建议 4:在代码中适当添加注释
#
# - 建议 5:适当添加空行使代码布局更加合理
#
# - 建议 6:编写函数的 4 个原则
#
# (1)函数设计要尽量短小,嵌套层次不宜过深
#
# (2)函数声明应该做到合理、简单、易用
#
# (3)函数参数设计应该考虑向下兼容
#
# (4)一个函数只做一件事,尽量保证函数粒度的一致性
#
# - 建议 7:将常量集中在一个文件,且常量名尽量使用全大写字母
# ## 2. 编程惯用法
#
# - 建议 8:利用 assert 语句来发现问题,但要注意,断言 assert 会影响效率
#
# - 建议 9:数据交换值时不推荐使用临时变量,而是直接 a, b = b, a
#
# - 建议 10:充分利用惰性计算(Lazy evaluation)的特性,从而避免不必要的计算
#
# - 建议 11:理解枚举替代实现的缺陷(最新版 Python 中已经加入了枚举特性)
#
# - 建议 12:不推荐使用 type 来进行类型检查,因为有些时候 type 的结果并不一定可靠。如果有需求,建议使用 isinstance 函数来代替
#
# - 建议 13:尽量将变量转化为浮点类型后再做除法(Python3 以后不用考虑)
#
# - 建议 14:警惕 eval() 函数的安全漏洞,有点类似于 SQL 注入
#
# - 建议 15:使用 enumerate() 同时获取序列迭代的索引和值
#
# - 建议 16:分清 == 和 is 的适用场景,特别是在比较字符串等不可变类型变量时(详见评论)
#
# - 建议 17:尽量使用 Unicode。在 Python2 中编码是很让人头痛的一件事,但 Python3 就不用过多考虑了
#
# - 建议 18:构建合理的包层次来管理 Module
# ## 3. 基础用法
#
# - 建议 19:有节制的使用 from…import 语句,防止污染命名空间
#
# - 建议 20:优先使用 absolute import 来导入模块(Python3 中已经移除了 relative import)
#
# - 建议 21:i+=1 不等于 ++i,在 Python 中,++i 前边的加号仅表示正,不表示操作
#
# - 建议 22:习惯使用 with 自动关闭资源,特别是在文件读写中
#
# - 建议 23:使用 else 子句简化循环(异常处理)
#
# - 建议 24:遵循异常处理的几点基本原则
#
# (1)注意异常的粒度,try 块中尽量少写代码
#
# (2)谨慎使用单独的 except 语句,或 except Exception 语句,而是定位到具体异常
#
# (3)注意异常捕获的顺序,在合适的层次处理异常
#
# (4)使用更加友好的异常信息,遵守异常参数的规范
#
# - 建议 25:避免 finally 中可能发生的陷阱
#
# - 建议 26:深入理解 None,正确判断对象是否为空。
#
# - 建议 27:连接字符串应优先使用 join 函数,而不是 + 操作
#
# - 建议 28:格式化字符串时尽量使用 format 函数,而不是 % 形式
#
# - 建议 29:区别对待可变对象和不可变对象,特别是作为函数参数时
#
# - 建议 30:[], {} 和 ():一致的容器初始化形式。使用列表解析可以使代码更清晰,同时效率更高
#
# - 建议 31:函数传参数,既不是传值也不是传引用,而是传对象或者说对象的引用
#
# - 建议 32:警惕默认参数潜在的问题,特别是当默认参数为可变对象时
#
# - 建议 33:函数中慎用变长参数 **args 和** **kargs**
#
# (1)这种使用太灵活,从而使得函数签名不够清晰,可读性较差
#
# (2)如果因为函数参数过多而是用变长参数简化函数定义,那么一般该函数可以重构
#
# - 建议 34:深入理解 str() 和 repr() 的区别
#
# (1)两者之间的目标不同:str 主要面向客户,其目的是可读性,返回形式为用户友好性和可读性都比较高的字符串形式;而 repr 是面向 Python 解释器或者说 Python 开发人员,其目的是准确性,其返回值表示 Python 解释器内部的定义
#
# (2)在解释器中直接输入变量,默认调用 repr 函数,而 print(var) 默认调用 str 函数
#
# (3)repr 函数的返回值一般可以用 eval 函数来还原对象
#
# (4)两者分别调用对象的内建函数 **__str__** () 和 **__repr__** ()
#
# - 建议 35:分清静态方法 staticmethod 和类方法 classmethod 的使用场景
# ## 4. 库的使用
#
# - 建议 36:掌握字符串的基本用法
#
# - 建议 37:按需选择 sort() 和 sorted() 函数
#
# sort() 是列表在就地进行排序,所以不能排序元组等不可变类型。
#
# sorted() 可以排序任意的可迭代类型,同时不改变原变量本身。
#
# - 建议 38:使用 copy 模块深拷贝对象,区分浅拷贝(shallow copy)和深拷贝(deep copy)
#
# - 建议 39:使用 Counter 进行计数统计,Counter 是字典类的子类,在 collections 模块中
#
# - 建议 40:深入掌握 ConfigParse
#
# - 建议 41:使用 argparse 模块处理命令行参数
#
# - 建议 42:使用 pandas 处理大型 CSV 文件
#
# Python 本身提供一个 CSV 文件处理模块,并提供 reader、writer 等函数。
#
# Pandas 可提供分块、合并处理等,适用于数据量大的情况,且对二维数据操作更方便。
#
# - 建议 43:使用 ElementTree 解析 XML
#
# - 建议 44:理解模块 pickle 的优劣
#
# 优势:接口简单、各平台通用、支持的数据类型广泛、扩展性强
#
# 劣势:不保证数据操作的原子性、存在安全问题、不同语言之间不兼容
#
# - 建议 45:序列化的另一个选择 JSON 模块:load 和 dump 操作
#
# - 建议 46:使用 traceback 获取栈信息
#
# - 建议 47:使用 logging 记录日志信息
#
# - 建议 48:使用 threading 模块编写多线程程序
#
# - 建议 49:使用 Queue 模块使多线程编程更安全
# ## 5. 设计模式
#
# - 建议 50:利用模块实现单例模式
#
# - 建议 51:用 mixin 模式让程序更加灵活
#
# - 建议 52:用发布 - 订阅模式实现松耦合
#
# - 建议 53:用状态模式美化代码
# ## 6. 内部机制
#
# - 建议 54:理解 build-in 对象
#
# - 建议 55:**__init__** () 不是构造方法,理解 **__new__** () 与它之间的区别
#
# - 建议 56:理解变量的查找机制,即作用域
#
# - 局部作用域
#
# - 全局作用域
#
# - 嵌套作用域
# -
# - 内置作用域
#
# - 建议 57:为什么需要 self 参数
#
# - 建议 58:理解 MRO(方法解析顺序)与多继承
#
# - 建议 59:理解描述符机制
#
# - 建议 60:区别 **__getattr__** () 与 **__getattribute__** () 方法之间的区别
#
# - 建议 61:使用更安全的 property
#
# - 建议 62:掌握元类 metaclass
#
# - 建议 63:熟悉 Python 对象协议
#
# - 建议 64:利用操作符重载实现中缀语法
#
# - 建议 65:熟悉 Python 的迭代器协议
#
# - 建议 66:熟悉 Python 的生成器
#
# - 建议 67:基于生成器的协程和 greenlet,理解协程、多线程、多进程之间的区别
#
# - 建议 68:理解 GIL 的局限性
#
# - 建议 69:对象的管理和垃圾回收
# ## 7.使用工具辅助项目开发
#
# - 建议 70:从 PyPI 安装第三方包
#
# - 建议 71:使用 pip 和 yolk 安装、管理包
#
# - 建议 72:做 paster 创建包
#
# - 建议 73:理解单元测试的概念
#
# - 建议 74:为包编写单元测试
#
# - 建议 75:利用测试驱动开发(TDD)提高代码的可测性
#
# - 建议 76:使用 Pylint 检查代码风格
#
# - 代码风格审查
#
# - 代码错误检查
#
# - 发现重复以及不合理的代码,方便重构
#
# - 高度的可配置化和可定制化
#
# - 支持各种 IDE 和编辑器的集成
#
# - 能够基于 Python 代码生成 UML 图
#
# - 能够与 Jenkins 等持续集成工具相结合,支持自动代码审查
#
# - 建议 77:进行高效的代码审查
#
# - 建议 78:将包发布到 PyPI
# ## 8. 性能剖析与优化
#
# - 建议 79:了解代码优化的基本原则
#
# - 建议 80:借助性能优化工具
#
# - 建议 81:利用 cProfile 定位性能瓶颈
#
# - 建议 82:使用 memory_profiler 和 objgraph 剖析内存使用
#
# - 建议 83:努力降低算法复杂度
#
# - 建议 84:掌握循环优化的基本技巧
#
# - 减少循环内部的计算
#
# - 将显式循环改为隐式循环,当然这会牺牲代码的可读性
#
# - 在循环中尽量引用局部变量
#
# - 关注内层嵌套循环
#
# - 建议 85:使用生成器提高效率
#
# - 建议 86:使用不同的数据结构优化性能
#
# - 建议 87:充分利用 set 的优势
#
# - 建议 88:使用 multiprocessing 模块克服 GIL 缺陷
#
# - 建议 89:使用线程池提高效率
#
# - 建议 90:使用 Cython 编写扩展模块
| 基础/90 条写 Python 程序的建议.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (pygta5)
# language: python
# name: pygta5
# ---
# # Part 2 - Model creation
# In this part we are going to create the neural network model and train.
# ## Loding Libraries
# Here we load the common libraries
import numpy as np
from grabscreen import grab_screen
import cv2
#import tdqm
import os
import pandas as pd
#from tqdm import tqdm
from collections import deque
from models import inception_v3 as googlenet
from models import alexnet2
from random import shuffle
# # Analysis of the Input file
# We are going to analize the files that we have created in the part 1
# Let us first select the first created file.
#Preprocessed image rgb color - no image filters
file_name = "preprocessed_training_data-1.npy"
#file_name = "training_data-1.npy"
# +
#Processed image single color
#file_name = "processed_training_data-1.npy"
# -
# full file info
train_data = np.load(file_name,allow_pickle=True)
# This file has the following shape
train_data.shape
#(500, 2)
# +
#train_data
# -
type(train_data )
# The the first three input frames are presented as:
train_data[0][1]
train_data[1][1]
train_data[2][1]
# There are 29 input componentes for each frame, we can plot the histogram for each component
train_hist = train_data[:]
train_hist.shape
df = pd.DataFrame()
for i in range(len(train_hist)):
row=list(train_hist[i][1])
#print(row)
temp = pd.DataFrame([row])
# print(temp)
df = pd.concat([df, temp])
df
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# We can analize the train data
# +
#ax = df.hist(bins=29, grid=False, figsize=(15,18), color='#86bf91', zorder=5, rwidth=5)
# -
train = train_data[:-50]
test = train_data[-50:]
# +
#test
# -
test.shape
#(50, 2)
train.shape
#(450, 2)
# We begin the train part
# ## Train Image part ( 4 Dimensional)
X_image = np.array([i[0] for i in train])
X_image.shape
#(450, 270, 480, 3)
X_image[449][269][3]
# +
#(450, 270, 480, 3) We choose only the imagen part of the train data,
#There are 450 picturtes with resolutionn WIDTH = 480 and HEIGHT = 270 with 3 colors rgb
# +
# We perform the reshape
# -
# numpy.reshape(a, newshape, order='C')
#
# - a - Array to be reshaped.
# - newshape - The new shape should be compatible with the original shape.
#
# - order- Read the elements of a using this index order, and place the elements into the reshaped array using this index order.
#
# Gives a new shape to an array without changing its data.
WIDTH = 480
HEIGHT = 270
# Using arr.reshape() will give a new shape to an array without changing the data. Just remember that when you use the reshape method, the array you want to produce needs to have the same number of elements as the original array.
# If you start with an array with N elements, you’ll need to make sure that your new array also has a total of N elements
import numpy as np
X_image.size
X_image.ndim
X_image.shape
# You can use reshape() to reshape your array.
#
#
#
# 
#
# With np.reshape, you can specify a few optional parameters:
# np.reshape(a, newshape=(d, e), order='C')
#
# a is the array to be reshaped.
#
# newshape is the new shape you want. You can specify an integer or a tuple of integers. If you specify an integer, the result will be an array of that length. The shape should be compatible with the original shape.
#
# order: C means to read/write the elements using C-like index order, F means to read/write the elements using Fortran-like index order, A means to read/write the elements in Fortran-like index order if a is Fortran contiguous in memory, C-like order otherwise. (This is an optional parameter and doesn’t need to be specified.)
#
# If you want to learn more about C and Fortran order, you can read more about the internal organization of NumPy arrays here. Essentially, C and Fortran orders have to do with how indices correspond to the order the array is stored in memory. In Fortran, when moving through the elements of a two-dimensional array as it is stored in memory, the first index is the most rapidly varying index. As the first index moves to the next row as it changes, the matrix is stored one column at a time. This is why Fortran is thought of as a Column-major language. In C on the other hand, the last index changes the most rapidly. The matrix is stored by rows, making it a Row-major language. What you do for C or Fortran depends on whether it’s more important to preserve the indexing convention or not reorder the data.
X_image.size
# We will reshape 270, 480 to 480, 270
#
# (450, 270, 480, 3) -> (450, 480, 270, 3)
# What does -1 mean in numpy reshape? A numpy matrix can be reshaped into a vector using reshape function with parameter -1. The criterion to satisfy for providing the new shape is that 'The new shape should be compatible with the original shape'
#
# numpy allow us to give one of new shape parameter as -1 (eg: (-1,WIDTH,HEIGHT,3) . It simply means that it is an unknown dimension and we want numpy to figure it out. And numpy will figure this by looking at the 'length of the array and remaining dimensions' and making sure it satisfies the above mentioned criteria
# For preprocessed rgb
X=X_image.reshape(-1,WIDTH,HEIGHT,3)
X.shape
#(450, 480, 270, 3) For 3 colors
X.size
# +
# For processed unicolor
#X=X_image.reshape(-1,WIDTH,HEIGHT,1)
#X.shape
#(450, 480, 270, 1) For one color
# -
# ## Train Input part ( 1 Dimensional )
Y = [i[1] for i in train]
#Z = [i[2] for i in train]
Y[0]
type(Y[0])
len(Y)
type(Y)
# We begin the test part
# ## Test Image part ( 4 Dimensional)
test.shape
test_image = np.array([i[0] for i in test])
type(test_image)
test_image.ndim
test_image.shape
#(50, 270, 480, 3)
# numpy.reshape(a, newshape, order='C')
#
# - a - Array to be reshaped.
# - newshape - The new shape should be compatible with the original shape.
#
# - order- Read the elements of a using this index order, and place the elements into the reshaped array using this index order.
#
# Gives a new shape to an array without changing its data.
#For Preprocessed
test_x=test_image.reshape(-1,WIDTH,HEIGHT,3)
test_x.shape
#(50, 480, 270, 3)
# +
#For Processed
#test_x=test_image.reshape(-1,WIDTH,HEIGHT,1)
#test_x.shape
# -
# ## Test Input part
test_y = [i[1] for i in test]
# +
#FILE_I_END = 1860
FILE_I_END = 2
WIDTH = 480
HEIGHT = 270
LR = 1e-3
#EPOCHS = 30
EPOCHS = 1
MODEL_NAME = 'model/test'
PREV_MODEL = ''
LOAD_MODEL = True
wl = 0
sl = 0
al = 0
dl = 0
wal = 0
wdl = 0
sal = 0
sdl = 0
nkl = 0
w = [1,0,0,0,0,0,0,0,0]
s = [0,1,0,0,0,0,0,0,0]
a = [0,0,1,0,0,0,0,0,0]
d = [0,0,0,1,0,0,0,0,0]
wa = [0,0,0,0,1,0,0,0,0]
wd = [0,0,0,0,0,1,0,0,0]
sa = [0,0,0,0,0,0,1,0,0]
sd = [0,0,0,0,0,0,0,1,0]
nk = [0,0,0,0,0,0,0,0,1]
# -
len(nk )
# +
#model = googlenet(WIDTH, HEIGHT, 3, LR, output=9, model_name=MODEL_NAME)
# -
size=[0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0]
len(size)
model = googlenet(WIDTH, HEIGHT, 3, LR, output=29, model_name=MODEL_NAME)
model.fit({'input': X},
{'targets': Y},
n_epoch=1,
validation_set=({'input': test_x},{'targets': test_y}),
snapshot_step=2500,
show_metric=True,
run_id=MODEL_NAME)
from IPython.display import display_html
def restartkernel() :
display_html("<script>Jupyter.notebook.kernel.restart()</script>",raw=True)
restartkernel()
# ## Full code
# +
import numpy as np
from grabscreen import grab_screen
import cv2
#import tdqm
import os
import pandas as pd
#from tqdm import tqdm
from collections import deque
from models import inception_v3 as googlenet
from models import alexnet2
from random import shuffle
#FILE_I_END = 1860
FILE_I_END = 30
#FILE_I_END = 2
WIDTH = 480
HEIGHT = 270
LR = 1e-3
#EPOCHS = 32
EPOCHS = 1
MODEL_NAME = 'model/test'
PREV_MODEL = ''
LOAD_MODEL = False
wl = 0
sl = 0
al = 0
dl = 0
wal = 0
wdl = 0
sal = 0
sdl = 0
nkl = 0
w = [1,0,0,0,0,0,0,0,0]
s = [0,1,0,0,0,0,0,0,0]
a = [0,0,1,0,0,0,0,0,0]
d = [0,0,0,1,0,0,0,0,0]
wa = [0,0,0,0,1,0,0,0,0]
wd = [0,0,0,0,0,1,0,0,0]
sa = [0,0,0,0,0,0,1,0,0]
sd = [0,0,0,0,0,0,0,1,0]
nk = [0,0,0,0,0,0,0,0,1]
model = googlenet(WIDTH, HEIGHT, 3, LR, output=29, model_name=MODEL_NAME)
if LOAD_MODEL:
model.load(PREV_MODEL)
print('We have loaded a previous model!!!!')
# iterates through the training files
for e in range(EPOCHS):
data_order = [i for i in range(1,FILE_I_END+1)]
shuffle(data_order)
for count,i in enumerate(data_order):
try:
#Preprocessed image rgb color - no image filters
file_name = 'preprocessed_training_data-{}.npy'.format(i)
# full file info
train_data = np.load(file_name,allow_pickle=True)
train = train_data[:-50]
test = train_data[-50:]
X_image = np.array([i[0] for i in train])
# For preprocessed rgb
X=X_image.reshape(-1,WIDTH,HEIGHT,3)
Y = [i[1] for i in train]
test_image = np.array([i[0] for i in test])
#For Preprocessed
test_x=test_image.reshape(-1,WIDTH,HEIGHT,3)
test_y = [i[1] for i in test]
model.fit({'input': X},
{'targets': Y},
n_epoch=1,
validation_set=({'input': test_x},{'targets': test_y}),
snapshot_step=2500,
show_metric=True,
run_id=MODEL_NAME)
if count%10 == 0:
print('SAVING MODEL!')
model.save(MODEL_NAME)
except Exception as e:
print(str(e))
# -
# ## Alternative Approaches
# In this notebook we have used InceptionV3, however it is possible use differnet available popular neural networks.
# Taking into account Keras, In deep learning models there are some applications available alongside pre-trained weights. These models can be used for prediction, feature extraction, and fine-tuning.
#
# Weights are downloaded automatically when instantiating a model. They are stored at ~/.keras/models/.
#
# Upon instantiation, the models will be built according to the image data format set in your Keras configuration file at ~/.keras/keras.json. For instance, if you have set image_data_format=channels_last, then any model loaded from this repository will get built according to the TensorFlow data format convention, "Height-Width-Depth".
#
# Available models
# Model Size (MB) Top-1 Accuracy Top-5 Accuracy Parameters Depth Time (ms) per inference step (CPU) Time (ms) per inference step (GPU)
# - Xception 88 79.0% 94.5% 22.9M 81 109.4 8.1
# - VGG16 528 71.3% 90.1% 138.4M 16 69.5 4.2
# - VGG19 549 71.3% 90.0% 143.7M 19 84.8 4.4
# - ResNet50 98 74.9% 92.1% 25.6M 107 58.2 4.6
# - ResNet50V2 98 76.0% 93.0% 25.6M 103 45.6 4.4
# - ResNet101 171 76.4% 92.8% 44.7M 209 89.6 5.2
# - ResNet101V2 171 77.2% 93.8% 44.7M 205 72.7 5.4
# - ResNet152 232 76.6% 93.1% 60.4M 311 127.4 6.5
# - ResNet152V2 232 78.0% 94.2% 60.4M 307 107.5 6.6
# - InceptionV3 92 77.9% 93.7% 23.9M 189 42.2 6.9
| versions/0.01/2-train_model_preprocessed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-2019_rbig_ad]
# language: python
# name: conda-env-.conda-2019_rbig_ad-py
# ---
# # Information Theory Measures
#
# In this notebook, I will be demonstrating some of the aspects of information theory measures.
# ## Data - Climate Models
# +
import os, sys
cwd = os.getcwd()
source_path = f"{cwd}/../../../"
sys.path.insert(0, f'{source_path}')
import numpy as np
# Data Loaders
from src.data.climate.amip import DataDownloader, DataLoader
from src.data.climate.era5 import get_era5_data
from src.data.climate.ncep import get_ncep_data
from src.features.climate.build_features import (
get_time_overlap, check_time_coords, regrid_2_lower_res, get_spatial_cubes, normalize_data)
from src.experiments.climate.amip_global import (
experiment_loop_comparative,
experiment_loop_individual
)
# Stat Tools
from src.models.information.entropy import RBIGEstimator as RBIGENTEST
from src.models.information.mutual_information import RBIGEstimator as RBIGMIEST
from scipy import stats
import pandas as pd
import xarray as xr
from tqdm import tqdm
from sklearn import preprocessing
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
amip_data_path = f"/home/emmanuel/projects/2020_rbig_rs/data/climate/raw/amip/"
era5_path = f"/home/emmanuel/projects/2020_rbig_rs/data/climate/raw/era5/"
ncep_path = f"/home/emmanuel/projects/2020_rbig_rs/data/climate/raw/ncep/"
results_path = f"/home/emmanuel/projects/2020_rbig_rs/data/climate/results/"
fig_path = f"/home/emmanuel/projects/2020_rbig_rs/reports/figures/climate/"
# ## Demo Experiment
# ### Experimental Paams
# +
class DataArgs:
data_path = "/home/emmanuel/projects/2020_rbig_rs/data/climate/raw/amip/"
results_path = "/home/emmanuel/projects/2020_rbig_rs/data/climate/results/amip"
class CMIPArgs:
# Fixed Params
spatial_windows = [
1, 2, # Spatial Window for Density Cubes
3,4,5,6,7,8,9,10
]
# Free Params
variables = [
'psl' # Mean Surface Pressure
]
cmip_models = [
"inmcm4",
"access1_0",
"bcc_csm1_1",
"bcc_csm1_1_m",
"bnu_esm",
"giss_e2_r",
"cnrm_cm5",
"ipsl_cm5a_lr",
"ipsl_cm5a_mr",
"ipsl_cm5b_lr",
"mpi_esm_lr",
"mpi_esm_mr",
"noresm1_m",
]
base_models = [
'ncep',
"era5"
]
# -
# ### Part I - Grab Data
# +
from src.data.climate.amip import get_base_model
base_dat = get_base_model(CMIPArgs.base_models[0], CMIPArgs.variables[0])
# base_dat
# +
from src.data.climate.cmip5 import get_cmip5_model
cmip_dat = get_cmip5_model(CMIPArgs.cmip_models[0], CMIPArgs.variables[0])
# cmip_dat
# -
# ### Part II - Regrid Data
# +
base_dat, cmip_dat = regrid_2_lower_res(base_dat, cmip_dat)
assert(base_dat.shape[1] == cmip_dat.shape[1])
assert(base_dat.shape[2] == cmip_dat.shape[2])
# base_dat
# -
# ### Part III - Find Overlapping Times
base_dat.shape, cmip_dat.shape
base_dat, cmip_dat = get_time_overlap(base_dat, cmip_dat)
# ### Part IV - Get Density Cubes
base_df = get_spatial_cubes(base_dat, CMIPArgs.spatial_windows[3])
cmip_df = get_spatial_cubes(cmip_dat, CMIPArgs.spatial_windows[3])
base_df.shape
# ### Normalize
base_df = normalize_data(base_df)
cmip_df = normalize_data(cmip_df)
# ## Information Theory Measures
# ### Entropy, H($X$)
# +
subsample = 10_000
batch_size = None
bootstrap = False
ent_est = RBIGENTEST(
batch_size=batch_size,
bootstrap=bootstrap,
)
ent_est.fit(base_df[:subsample])
h = ent_est.score(base_df[:subsample])
h
# -
# #### with Bootstrap
# +
batch_size = 10_000
bootstrap = True
n_iterations = 100
ent_est = RBIGENTEST(
batch_size=batch_size,
bootstrap=bootstrap,
n_iterations=n_iterations
)
ent_est.fit(base_df)
h = ent_est.score(base_df)
h
# -
plt.hist(ent_est.raw_scores)
plt.hist(ent_est.raw_scores)
# #### W. Batches
# +
subsample = 40_000
ent_est = RBIGENTEST(batch_size=10_000)
ent_est.fit(base_df[:subsample])
h = ent_est.score(base_df[:subsample])
h
# -
ent_est.raw_scores
# ### Total Correlation, TC($X$)
# +
subsample = 40_000
tc_est = RBIGMIEST(batch_size=None)
tc_est.fit(base_df[:subsample])
tc = tc_est.score(base_df[:subsample])
tc
# -
# #### w. Batches
# +
subsample = 40_000
tc_est = RBIGMIEST(batch_size=10_000)
tc_est.fit(base_df[:subsample])
tc = tc_est.score(base_df[:subsample])
tc
# -
tc_est.raw_scores
# ### Mutual Information, MI($X$)
# +
subsample = 100_000
mi_est = RBIGMIEST(batch_size=None)
mi_est.fit(
base_df[:subsample],
cmip_df[:subsample]
)
mi = mi_est.score(base_df[:subsample])
mi
# -
# #### w. Batches
# +
subsample = 100_000
mi_est = RBIGMIEST(batch_size=50_000)
mi_est.fit(
base_df[:subsample],
cmip_df[:subsample]
)
mi = mi_est.score(base_df[:subsample])
mi
# -
mi_est.raw_values
# ### Mutual Information II, H(X) + H(Y) - H(X,Y)
# +
subsample = 100_000
batch_size = 25_000
# H(X)
print('H(X)')
x_ent_est = RBIGENTEST(batch_size=batch_size)
x_ent_est.fit(base_df.values[:subsample])
h_x = x_ent_est.score(base_df.values[:subsample])
# H(Y)
print('H(Y)')
y_ent_est = RBIGENTEST(batch_size=batch_size)
y_ent_est.fit(cmip_df.values[:subsample])
h_y = y_ent_est.score(cmip_df.values[:subsample])
# H(X,Y)
print('H(X,Y)')
xy_ent_est = RBIGENTEST(batch_size=50_000)
xy_ent_est.fit(
np.hstack(
(
base_df.values[:subsample],
cmip_df.values[:subsample]
)
),
)
h_xy = xy_ent_est.score(base_df.values[:subsample])
# +
# H(X,Y)
print('H(X,Y)')
xy_ent_est = RBIGENTEST(batch_size=50_000)
xy_ent_est.fit(
np.hstack(
(
base_df.values[:subsample],
cmip_df.values[:subsample]
)
),
)
h_xy = xy_ent_est.score(base_df.values[:subsample])
h_xy
# -
h_x, h_y, h_xy, h_x + h_y - h_xy
# ### Correlation: Pearson, Spearman, KendallTau
# +
pear = stats.pearsonr(
base_df[:subsample].ravel(),
cmip_df[:subsample].ravel(),
)
spear = stats.spearmanr(
base_df[:subsample].ravel(),
cmip_df[:subsample].ravel(),
)
kend = stats.kendalltau(
base_df[:subsample].ravel(),
cmip_df[:subsample].ravel(),
)
pear[0], spear[0], kend[0]
# -
| notebooks/climate/rcp/3.2_demo_it_measures-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# name: python390jvsc74a57bd063fd5069d213b44bf678585dea6b12cceca9941eaf7f819626cde1f2670de90d
# ---
# +
import pandas as pd
# Load the spam dataset (NB: "ham" is the label for non-spam messages.)
spam = pd.read_csv('data/spam.csv')
spam.head(10)
# +
import spacy
# Create an empty spaCy model (English language: en_core_web_sm)
nlp = spacy.blank("en")
# Create the TextCategorizer. "exclusive_classes" set as True and the "architecture" set to "bow" (Bag of words).
textcat = nlp.create_pipe(
"textcat",
config={
"exclusive_classes": True,
"architecture": "bow"})
# Add the TextCategorizer to the model:
nlp.add_pipe(textcat)
# +
# Add the 2 labels ("ham" and "spam") to the text classifier:
textcat.add_label("ham")
textcat.add_label("spam")
# +
# Convert labels and store in the correct dictionary:
train_texts = spam['text'].values
train_labels = [{'cats': {'ham': label == 'ham',
'spam': label == 'spam'}}
for label in spam['label']]
# +
# Combine both thte texts and the labels into a single list.
train_data = list(zip(train_texts, train_labels))
train_data[:3]
# +
from spacy.util import minibatch
spacy.util.fix_random_seed(1)
optimizer = nlp.begin_training()
# Create the batch generator: (Batch size of 8)
batches = minibatch(train_data, size=8)
# Iterate through minibatches:
for batch in batches:
texts, labels = zip(*batch)
nlp.update(texts, labels, sgd=optimizer)
# The above code transforms the batch from a list of (text, label)s into separate lists for texts and labels to update(). (This code is a useful way to split a list of tuples into lists.)
# +
# This uses the same code as above but with an additional loop for multiple epochs. The training data is also reshuffled at the beginning of each loop.
import random
random.seed(1)
spacy.util.fix_random_seed(1)
optimizer = nlp.begin_training()
losses = {}
for epoch in range(10): # 10 epochs
random.shuffle(train_data)
# Create the batch generator: (Batch size of 8)
batches = minibatch(train_data, size=8)
# Iterate through minibatches
for batch in batches:
texts, labels = zip(*batch)
nlp.update(texts, labels, sgd=optimizer, losses=losses)
print(losses)
# +
# Example text taken from Kaggle tutorial code
texts = ["Are you ready for the tea party????? It's gonna be wild",
"URGENT Reply to this message for GUARANTEED FREE TEA" ]
# tokenise the above text:
docs = [nlp.tokenizer(text) for text in texts]
# Use textcat to get the scores for each doc:
textcat = nlp.get_pipe('textcat')
scores, _ = textcat.predict(docs)
print(scores)
# +
# From the scores predicted by the model find the label with the highest score (probability).
predicted_labels = scores.argmax(axis=1)
print([textcat.labels[label] for label in predicted_labels])
# +
# Below is some code I thought was particularly useful from the exercise:
def evaluate(model, texts, labels):
""" Returns the accuracy of a TextCategorizer model.
Arguments
---------
model: ScaPy model with a TextCategorizer
texts: Text samples, from load_data function
labels: True labels, from load_data function
"""
# Get predictions from textcat model (using your predict method)
predicted_class = predict(models, texts)
# From labels, get the true class as a list of integers (POSITIVE -> 1, NEGATIVE -> 0)
true_class = [int(each['cats']['POSITIVE']) for each in labels]
# A boolean or int array indicating correct predictions
correct_predictions = predicted_class == true_class
# The accuracy, number of correct predictions divided by all predictions
accuracy = correct_predictions.mean()
return accuracy
# -
| Kaggle_ML_Tutorials/NaturalLanguageProcessing/TCtutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="168QPRizVFFg"
# <p style="font-size:32px;text-align:center"> <b>Social network Graph Link Prediction - Facebook Challenge</b> </p>
# + colab={} colab_type="code" id="Q8lS7fVyVFFl"
#Importing Libraries
# please do go through this python notebook:
import warnings
warnings.filterwarnings("ignore")
import csv
import pandas as pd#pandas to create small dataframes
import datetime #Convert to unix time
import time #Convert to unix time
# if numpy is not installed already : pip3 install numpy
import numpy as np#Do aritmetic operations on arrays
# matplotlib: used to plot graphs
import matplotlib
import matplotlib.pylab as plt
import seaborn as sns#Plots
from matplotlib import rcParams#Size of plots
from sklearn.cluster import MiniBatchKMeans, KMeans#Clustering
import math
import pickle
import os
# to install xgboost: pip3 install xgboost
import xgboost as xgb
import warnings
import networkx as nx
import pdb
import pickle
from pandas import HDFStore,DataFrame
from pandas import read_hdf
from scipy.sparse.linalg import svds, eigs
import gc
from tqdm import tqdm
# + [markdown] colab_type="text" id="1znHayNeVFFt"
# # 1. Reading Data
# + colab={} colab_type="code" id="Uq9HbHwEVFFv" outputId="b2aa525a-93d3-47c3-8216-416a811bc812"
if os.path.isfile('data/after_eda/train_pos_after_eda.csv'):
train_graph=nx.read_edgelist('data/after_eda/train_pos_after_eda.csv',delimiter=',',create_using=nx.DiGraph(),nodetype=int)
print(nx.info(train_graph))
else:
print("please run the FB_EDA.ipynb or download the files from drive")
# -
train_graph.in_degree(1), train_graph.out_degree(1)
# + [markdown] colab_type="text" id="HmlUa64tVFF7"
# # 2. Similarity measures
# + [markdown] colab_type="text" id="ivVMUMiWVFF9"
# ## 2.1 Jaccard Distance:
# http://www.statisticshowto.com/jaccard-index/
# + [markdown] colab_type="text" id="NoWCYuRBVFF_"
# \begin{equation}
# j = \frac{|X\cap Y|}{|X \cup Y|}
# \end{equation}
# + colab={} colab_type="code" id="Seo4z5SnVFGB"
#for followees
def jaccard_for_followees(a,b):
try:
if len(set(train_graph.successors(a))) == 0 | len(set(train_graph.successors(b))) == 0:
return 0
sim = (len(set(train_graph.successors(a)).intersection(set(train_graph.successors(b)))))/\
(len(set(train_graph.successors(a)).union(set(train_graph.successors(b)))))
except:
return 0
return sim
# + colab={} colab_type="code" id="Oa9FMlS8VFGF" outputId="426a6833-1631-4024-c24a-d21ae7686472"
#one test case
print(jaccard_for_followees(273084,1505602))
# + colab={} colab_type="code" id="Gf8njOv6VFGK" outputId="8ba07727-a0ab-498e-819f-0d310876191c"
#node 1635354 not in graph
print(jaccard_for_followees(273084,1635354))
# + colab={} colab_type="code" id="LO-a5ZkKVFGO"
#for followers
def jaccard_for_followers(a,b):
try:
if len(set(train_graph.predecessors(a))) == 0 | len(set(g.predecessors(b))) == 0:
return 0
sim = (len(set(train_graph.predecessors(a)).intersection(set(train_graph.predecessors(b)))))/\
(len(set(train_graph.predecessors(a)).union(set(train_graph.predecessors(b)))))
return sim
except:
return 0
# + colab={} colab_type="code" id="DlbX2t0jVFGQ" outputId="7e4b4536-442a-4b0c-ae02-fb442c1955db"
print(jaccard_for_followers(273084,470294))
# + colab={} colab_type="code" id="OgeBW2LMVFGU" outputId="1e12fabe-d990-4506-bb6b-c86b01d1b0af"
#node 1635354 not in graph
print(jaccard_for_followees(669354,1635354))
# + [markdown] colab_type="text" id="MnH2my2UVFGX"
# ## 2.2 Cosine distance
# + [markdown] colab_type="text" id="XNvdBGS2VFGY"
# \begin{equation}
# CosineDistance = \frac{|X\cap Y|}{|X|\cdot|Y|}
# \end{equation}
# + colab={} colab_type="code" id="Iznz67EdVFGZ"
#for followees
def cosine_for_followees(a,b):
try:
if len(set(train_graph.successors(a))) == 0 | len(set(train_graph.successors(b))) == 0:
return 0
sim = (len(set(train_graph.successors(a)).intersection(set(train_graph.successors(b)))))/\
(math.sqrt(len(set(train_graph.successors(a)))*len((set(train_graph.successors(b))))))
return sim
except:
return 0
# + colab={} colab_type="code" id="H55ALjkMVFGc" outputId="531fceba-60f4-4e6b-97f4-f37733dc468f"
print(cosine_for_followees(273084,1505602))
# + colab={} colab_type="code" id="q0RGKgJFVFGf" outputId="41202fc6-f4aa-4a1d-d8f6-84f960a3fbba"
print(cosine_for_followees(273084,1635354))
# + colab={} colab_type="code" id="KJ_yGxA0VFGj"
def cosine_for_followers(a,b):
try:
if len(set(train_graph.predecessors(a))) == 0 | len(set(train_graph.predecessors(b))) == 0:
return 0
sim = (len(set(train_graph.predecessors(a)).intersection(set(train_graph.predecessors(b)))))/\
(math.sqrt(len(set(train_graph.predecessors(a))))*(len(set(train_graph.predecessors(b)))))
return sim
except:
return 0
# + colab={} colab_type="code" id="75QrFJb6VFGm" outputId="f01e0558-f1e3-465f-ab14-0e4ca764f4aa"
print(cosine_for_followers(2,470294))
# + colab={} colab_type="code" id="-ut4k_F0VFGq" outputId="8bc9607a-7262-43e2-9de8-f71d276762fc"
print(cosine_for_followers(669354,1635354))
# + [markdown] colab_type="text" id="DaIHhWh6VFGv"
# ## 3. Ranking Measures
# + [markdown] colab_type="text" id="6nfV1SprVFGx"
# https://networkx.github.io/documentation/networkx-1.10/reference/generated/networkx.algorithms.link_analysis.pagerank_alg.pagerank.html
#
# PageRank computes a ranking of the nodes in the graph G based on the structure of the incoming links.
#
# <img src='PageRanks-Example.jpg'/>
#
# Mathematical PageRanks for a simple network, expressed as percentages. (Google uses a logarithmic scale.) Page C has a higher PageRank than Page E, even though there are fewer links to C; the one link to C comes from an important page and hence is of high value. If web surfers who start on a random page have an 85% likelihood of choosing a random link from the page they are currently visiting, and a 15% likelihood of jumping to a page chosen at random from the entire web, they will reach Page E 8.1% of the time. <b>(The 15% likelihood of jumping to an arbitrary page corresponds to a damping factor of 85%.) Without damping, all web surfers would eventually end up on Pages A, B, or C, and all other pages would have PageRank zero. In the presence of damping, Page A effectively links to all pages in the web, even though it has no outgoing links of its own.</b>
# + [markdown] colab_type="text" id="GkkfYYZ6VFGy"
# ## 3.1 Page Ranking
#
# https://en.wikipedia.org/wiki/PageRank
#
# + colab={} colab_type="code" id="AtvqwZ34VFGy"
if not os.path.isfile('data/fea_sample/page_rank.p'):
pr = nx.pagerank(train_graph, alpha=0.85)
pickle.dump(pr,open('data/fea_sample/page_rank.p','wb'))
else:
pr = pickle.load(open('data/fea_sample/page_rank.p','rb'))
# + colab={} colab_type="code" id="lXGKYYf6VFG2" outputId="bb3d1b7a-81f9-44ab-dbe7-3214ccd47179"
print('min',pr[min(pr, key=pr.get)])
print('max',pr[max(pr, key=pr.get)])
print('mean',float(sum(pr.values())) / len(pr))
# + colab={} colab_type="code" id="5xwlah4oVFG4" outputId="992fdfad-7ff6-4626-c9ee-d9bce220a680"
#for imputing to nodes which are not there in Train data
mean_pr = float(sum(pr.values())) / len(pr)
print(mean_pr)
# -
# + [markdown] colab_type="text" id="HhPbSL1tVFG7"
# # 4. Other Graph Features
# + [markdown] colab_type="text" id="AgsorCl7VFG8"
# ## 4.1 Shortest path:
# + [markdown] colab_type="text" id="E7teH2LCVFG9"
# Getting Shortest path between twoo nodes, if nodes have direct path i.e directly connected then we are removing that edge and calculating path.
# + colab={} colab_type="code" id="RA076ovzVFG9"
#if has direct edge then deleting that edge and calculating shortest path
def compute_shortest_path_length(a,b):
p=-1
try:
if train_graph.has_edge(a,b):
train_graph.remove_edge(a,b)
p= nx.shortest_path_length(train_graph,source=a,target=b)
train_graph.add_edge(a,b)
else:
p= nx.shortest_path_length(train_graph,source=a,target=b)
return p
except:
return -1
# + colab={} colab_type="code" id="AxnKId11VFG_" outputId="15ca223a-6a04-4549-d010-54619b472a9e"
#testing
compute_shortest_path_length(77697, 826021)
# + colab={} colab_type="code" id="0huWCNtRVFHC" outputId="6debfa4f-2067-48bc-84b3-ab86e2d9dea6"
#testing
compute_shortest_path_length(669354,1635354)
# + [markdown] colab_type="text" id="baE_95bzVFHF"
# ## 4.2 Checking for same community
# + colab={} colab_type="code" id="15CIQqAbVFHG"
#getting weekly connected edges from graph
wcc=list(nx.weakly_connected_components(train_graph))
def belongs_to_same_wcc(a,b):
index = []
if train_graph.has_edge(b,a):
return 1
if train_graph.has_edge(a,b):
for i in wcc:
if a in i:
index= i
break
if (b in index):
train_graph.remove_edge(a,b)
if compute_shortest_path_length(a,b)==-1:
train_graph.add_edge(a,b)
return 0
else:
train_graph.add_edge(a,b)
return 1
else:
return 0
else:
for i in wcc:
if a in i:
index= i
break
if(b in index):
return 1
else:
return 0
# + colab={} colab_type="code" id="fAzOHtCFVFHI" outputId="2b043a87-b460-42bf-f37e-4c04bbed6586"
belongs_to_same_wcc(861, 1659750)
# + colab={} colab_type="code" id="HMdYpPuGVFHK" outputId="2005e22c-b60f-48d7-839b-650bf97cae35"
belongs_to_same_wcc(669354,1635354)
# + [markdown] colab_type="text" id="q74nth0OVFHN"
# ## 4.3 Adamic/Adar Index:
# Adamic/Adar measures is defined as inverted sum of degrees of common neighbours for given two vertices.
# $$A(x,y)=\sum_{u \in N(x) \cap N(y)}\frac{1}{log(|N(u)|)}$$
# + colab={} colab_type="code" id="CeS98LI5VFHO"
#adar index
def calc_adar_in(a,b):
sum=0
try:
n=list(set(train_graph.successors(a)).intersection(set(train_graph.successors(b))))
if len(n)!=0:
for i in n:
sum=sum+(1/np.log10(len(list(train_graph.predecessors(i)))))
return sum
else:
return 0
except:
return 0
# + colab={} colab_type="code" id="KezFeRmyVFHQ" outputId="2f9c0e11-02d9-4f28-d67a-65e3d4943e99"
calc_adar_in(1,189226)
# + colab={} colab_type="code" id="vj_m89bBVFHV" outputId="68a0a099-2954-402f-c80f-6d436ffa1aba"
calc_adar_in(669354,1635354)
# + [markdown] colab_type="text" id="pBUudhFAVFHY"
# ## 4.4 Is persion was following back:
# + colab={} colab_type="code" id="j_mwmopLVFHZ"
def follows_back(a,b):
if train_graph.has_edge(b,a):
return 1
else:
return 0
# + colab={} colab_type="code" id="LdjUXIfbVFHb" outputId="ed3d8640-9834-4a95-e712-804292da70e9"
follows_back(1,189226)
# + colab={} colab_type="code" id="PmZtL65YVFHf" outputId="18ea6fe2-3f96-42c0-d116-ecb76ddba4b5"
follows_back(669354,1635354)
# + [markdown] colab_type="text" id="29Vrq2EXVFHi"
# ## 4.5 Katz Centrality:
# https://en.wikipedia.org/wiki/Katz_centrality
#
# https://www.geeksforgeeks.org/katz-centrality-centrality-measure/
# Katz centrality computes the centrality for a node
# based on the centrality of its neighbors. It is a
# generalization of the eigenvector centrality. The
# Katz centrality for node `i` is
#
# $$x_i = \alpha \sum_{j} A_{ij} x_j + \beta,$$
# where `A` is the adjacency matrix of the graph G
# with eigenvalues $$\lambda$$.
#
# The parameter $$\beta$$ controls the initial centrality and
#
# $$\alpha < \frac{1}{\lambda_{max}}.$$
# + colab={} colab_type="code" id="CN5OSqrkVFHj"
if not os.path.isfile('data/fea_sample/katz.p'):
katz = nx.katz.katz_centrality(train_graph,alpha=0.005,beta=1)
pickle.dump(katz,open('data/fea_sample/katz.p','wb'))
else:
katz = pickle.load(open('data/fea_sample/katz.p','rb'))
# + colab={} colab_type="code" id="gcU83vw7VFHm" outputId="05f49ad4-46fe-4cf6-f32a-2fe4846b0714"
print('min',katz[min(katz, key=katz.get)])
print('max',katz[max(katz, key=katz.get)])
print('mean',float(sum(katz.values())) / len(katz))
# + colab={} colab_type="code" id="qcboIksiVFHt" outputId="99f52422-9edb-479a-d5d9-e33401160da7"
mean_katz = float(sum(katz.values())) / len(katz)
print(mean_katz)
# + [markdown] colab_type="text" id="SRZqGFgYVFHx"
# ## 4.6 Hits Score
# The HITS algorithm computes two numbers for a node. Authorities estimates the node value based on the incoming links. Hubs estimates the node value based on outgoing links.
#
# https://en.wikipedia.org/wiki/HITS_algorithm
# + colab={} colab_type="code" id="WXNHRdzUVFHz"
if not os.path.isfile('data/fea_sample/hits.p'):
hits = nx.hits(train_graph, max_iter=100, tol=1e-08, nstart=None, normalized=True)
pickle.dump(hits,open('data/fea_sample/hits.p','wb'))
else:
hits = pickle.load(open('data/fea_sample/hits.p','rb'))
# + colab={} colab_type="code" id="PSUwSZBVVFH3" outputId="77448253-5409-4229-f0be-b8dbc14d7f46"
print('min',hits[0][min(hits[0], key=hits[0].get)])
print('max',hits[0][max(hits[0], key=hits[0].get)])
print('mean',float(sum(hits[0].values())) / len(hits[0]))
# -
# + [markdown] colab_type="text" id="ZZtowOLZVFH6"
# # 5. Featurization
# + [markdown] colab_type="text" id="o6NnRWmLVFH6"
# ## 5. 1 Reading a sample of Data from both train and test
# + colab={} colab_type="code" id="wgHje1UVVFH8"
import random
if os.path.isfile('data/after_eda/train_after_eda.csv'):
filename = "data/after_eda/train_after_eda.csv"
# you uncomment this line, if you dont know the lentgh of the file name
# here we have hardcoded the number of lines as 15100030
# n_train = sum(1 for line in open(filename)) #number of records in file (excludes header)
n_train = 15100028
s = 100000 #desired sample size
skip_train = sorted(random.sample(range(1,n_train+1),n_train-s))
#https://stackoverflow.com/a/22259008/4084039
# + colab={} colab_type="code" id="zOzuRFFlVFH-"
if os.path.isfile('data/after_eda/test_after_eda.csv'): # changed train to test
filename = "data/after_eda/test_after_eda.csv"
# you uncomment this line, if you dont know the lentgh of the file name
# here we have hardcoded the number of lines as 3775008
# n_test = sum(1 for line in open(filename)) #number of records in file (excludes header)
n_test = 3775006
s = 50000 #desired sample size
skip_test = sorted(random.sample(range(1,n_test+1),n_test-s))
#https://stackoverflow.com/a/22259008/4084039
# + colab={} colab_type="code" id="3D_SeUCOVFH_" outputId="322902a4-0420-4b99-8606-5fd0de4bbea4"
print("Number of rows in the train data file:", n_train)
print("Number of rows we are going to elimiate in train data are",len(skip_train))
print("Number of rows in the test data file:", n_test)
print("Number of rows we are going to elimiate in test data are",len(skip_test))
# + colab={} colab_type="code" id="pCisf6PpVFID" outputId="daf2af43-3f98-4466-ad99-03bc54464714"
df_final_train = pd.read_csv('data/after_eda/train_after_eda.csv', skiprows=skip_train, names=['source_node', 'destination_node'])
df_final_train['indicator_link'] = pd.read_csv('data/train_y.csv', skiprows=skip_train, names=['indicator_link'])
print("Our train matrix size ",df_final_train.shape)
df_final_train.head(2)
# + colab={} colab_type="code" id="tFn1RkdyVFIH" outputId="1ca99e70-6d2a-45f2-f51c-fd3b1211ad20"
df_final_test = pd.read_csv('data/after_eda/test_after_eda.csv', skiprows=skip_test, names=['source_node', 'destination_node'])
df_final_test['indicator_link'] = pd.read_csv('data/test_y.csv', skiprows=skip_test, names=['indicator_link'])
print("Our test matrix size ",df_final_test.shape)
df_final_test.head(2)
# -
# + [markdown] colab_type="text" id="gIaOWDaDVFIJ"
# ## 5.2 Adding a set of features
#
# __we will create these each of these features for both train and test data points__
# <ol>
# <li>jaccard_followers</li>
# <li>jaccard_followees</li>
# <li>cosine_followers</li>
# <li>cosine_followees</li>
# <li>num_followers_s</li>
# <li>num_followees_s</li>
# <li>num_followers_d</li>
# <li>num_followees_d</li>
# <li>inter_followers</li>
# <li>inter_followees</li>
# </ol>
# + colab={} colab_type="code" id="2qTkOiBcVFIJ"
if not os.path.isfile('data/fea_sample/storage_sample_stage1.h5'):
#mapping jaccrd followers to train and test data
df_final_train['jaccard_followers'] = df_final_train.apply(lambda row:
jaccard_for_followers(row['source_node'],row['destination_node']),axis=1)
df_final_test['jaccard_followers'] = df_final_test.apply(lambda row:
jaccard_for_followers(row['source_node'],row['destination_node']),axis=1)
#mapping jaccrd followees to train and test data
df_final_train['jaccard_followees'] = df_final_train.apply(lambda row:
jaccard_for_followees(row['source_node'],row['destination_node']),axis=1)
df_final_test['jaccard_followees'] = df_final_test.apply(lambda row:
jaccard_for_followees(row['source_node'],row['destination_node']),axis=1)
#mapping jaccrd followers to train and test data
df_final_train['cosine_followers'] = df_final_train.apply(lambda row:
cosine_for_followers(row['source_node'],row['destination_node']),axis=1)
df_final_test['cosine_followers'] = df_final_test.apply(lambda row:
cosine_for_followers(row['source_node'],row['destination_node']),axis=1)
#mapping jaccrd followees to train and test data
df_final_train['cosine_followees'] = df_final_train.apply(lambda row:
cosine_for_followees(row['source_node'],row['destination_node']),axis=1)
df_final_test['cosine_followees'] = df_final_test.apply(lambda row:
cosine_for_followees(row['source_node'],row['destination_node']),axis=1)
# + colab={} colab_type="code" id="fz2eZpSnVFIL"
def compute_features_stage1(df_final):
#calculating no of followers followees for source and destination
#calculating intersection of followers and followees for source and destination
num_followers_s=[]
num_followees_s=[]
num_followers_d=[]
num_followees_d=[]
inter_followers=[]
inter_followees=[]
for i,row in df_final.iterrows():
try:
s1=set(train_graph.predecessors(row['source_node']))
s2=set(train_graph.successors(row['source_node']))
except:
s1 = set()
s2 = set()
try:
d1=set(train_graph.predecessors(row['destination_node']))
d2=set(train_graph.successors(row['destination_node']))
except:
d1 = set()
d2 = set()
num_followers_s.append(len(s1))
num_followees_s.append(len(s2))
num_followers_d.append(len(d1))
num_followees_d.append(len(d2))
inter_followers.append(len(s1.intersection(d1)))
inter_followees.append(len(s2.intersection(d2)))
return num_followers_s, num_followers_d, num_followees_s, num_followees_d, inter_followers, inter_followees
# + colab={} colab_type="code" id="VFc60kcRVFIN"
if not os.path.isfile('data/fea_sample/storage_sample_stage1.h5'):
df_final_train['num_followers_s'], df_final_train['num_followers_d'], \
df_final_train['num_followees_s'], df_final_train['num_followees_d'], \
df_final_train['inter_followers'], df_final_train['inter_followees']= compute_features_stage1(df_final_train)
df_final_test['num_followers_s'], df_final_test['num_followers_d'], \
df_final_test['num_followees_s'], df_final_test['num_followees_d'], \
df_final_test['inter_followers'], df_final_test['inter_followees']= compute_features_stage1(df_final_test)
hdf = HDFStore('data/fea_sample/storage_sample_stage1.h5')
hdf.put('train_df',df_final_train, format='table', data_columns=True)
hdf.put('test_df',df_final_test, format='table', data_columns=True)
hdf.close()
else:
df_final_train = read_hdf('data/fea_sample/storage_sample_stage1.h5', 'train_df',mode='r')
df_final_test = read_hdf('data/fea_sample/storage_sample_stage1.h5', 'test_df',mode='r')
# + [markdown] colab_type="text" id="go_e8hxxVFIO"
# ## 5.3 Adding new set of features
#
# __we will create these each of these features for both train and test data points__
# <ol>
# <li>adar index</li>
# <li>is following back</li>
# <li>belongs to same weakly connect components</li>
# <li>shortest path between source and destination</li>
# </ol>
# + colab={} colab_type="code" id="LqB0Peg0VFIP"
if not os.path.isfile('data/fea_sample/storage_sample_stage2.h5'):
#mapping adar index on train
df_final_train['adar_index'] = df_final_train.apply(lambda row: calc_adar_in(row['source_node'],row['destination_node']),axis=1)
#mapping adar index on test
df_final_test['adar_index'] = df_final_test.apply(lambda row: calc_adar_in(row['source_node'],row['destination_node']),axis=1)
#--------------------------------------------------------------------------------------------------------
#mapping followback or not on train
df_final_train['follows_back'] = df_final_train.apply(lambda row: follows_back(row['source_node'],row['destination_node']),axis=1)
#mapping followback or not on test
df_final_test['follows_back'] = df_final_test.apply(lambda row: follows_back(row['source_node'],row['destination_node']),axis=1)
#--------------------------------------------------------------------------------------------------------
#mapping same component of wcc or not on train
df_final_train['same_comp'] = df_final_train.apply(lambda row: belongs_to_same_wcc(row['source_node'],row['destination_node']),axis=1)
##mapping same component of wcc or not on train
df_final_test['same_comp'] = df_final_test.apply(lambda row: belongs_to_same_wcc(row['source_node'],row['destination_node']),axis=1)
#--------------------------------------------------------------------------------------------------------
#mapping shortest path on train
df_final_train['shortest_path'] = df_final_train.apply(lambda row: compute_shortest_path_length(row['source_node'],row['destination_node']),axis=1)
#mapping shortest path on test
df_final_test['shortest_path'] = df_final_test.apply(lambda row: compute_shortest_path_length(row['source_node'],row['destination_node']),axis=1)
hdf = HDFStore('data/fea_sample/storage_sample_stage2.h5')
hdf.put('train_df',df_final_train, format='table', data_columns=True)
hdf.put('test_df',df_final_test, format='table', data_columns=True)
hdf.close()
else:
df_final_train = read_hdf('data/fea_sample/storage_sample_stage2.h5', 'train_df',mode='r')
df_final_test = read_hdf('data/fea_sample/storage_sample_stage2.h5', 'test_df',mode='r')
# + [markdown] colab_type="text" id="HJ8Dbma_VFIR"
# ## 5.4 Adding new set of features
#
# __we will create these each of these features for both train and test data points__
# <ol>
# <li>Weight Features
# <ul>
# <li>weight of incoming edges</li>
# <li>weight of outgoing edges</li>
# <li>weight of incoming edges + weight of outgoing edges</li>
# <li>weight of incoming edges * weight of outgoing edges</li>
# <li>2*weight of incoming edges + weight of outgoing edges</li>
# <li>weight of incoming edges + 2*weight of outgoing edges</li>
# </ul>
# </li>
# <li>Page Ranking of source</li>
# <li>Page Ranking of dest</li>
# <li>katz of source</li>
# <li>katz of dest</li>
# <li>hubs of source</li>
# <li>hubs of dest</li>
# <li>authorities_s of source</li>
# <li>authorities_s of dest</li>
# </ol>
# + [markdown] colab_type="text" id="iVHI2jtNVFIS"
# #### Weight Features
# + [markdown] colab_type="text" id="rXmUYF9FVFIT"
# In order to determine the similarity of nodes, an edge weight value was calculated between nodes. Edge weight decreases as the neighbor count goes up. Intuitively, consider one million people following a celebrity on a social network then chances are most of them never met each other or the celebrity. On the other hand, if a user has 30 contacts in his/her social network, the chances are higher that many of them know each other.
# `credit` - Graph-based Features for Supervised Link Prediction
# <NAME>, <NAME>, <NAME>
# + [markdown] colab_type="text" id="Qzbs2no7VFIV"
# \begin{equation}
# W = \frac{1}{\sqrt{1+|X|}}
# \end{equation}
# + [markdown] colab_type="text" id="kkzUPrWaVFIV"
# it is directed graph so calculated Weighted in and Weighted out differently
# + colab={} colab_type="code" id="FgNMzzTbVFIW" outputId="7e8e6d88-8bd6-45f6-f80e-82b093c18974"
#weight for source and destination of each link
Weight_in = {}
Weight_out = {}
for i in tqdm(train_graph.nodes()):
s1=set(train_graph.predecessors(i))
w_in = 1.0/(np.sqrt(1+len(s1)))
Weight_in[i]=w_in
s2=set(train_graph.successors(i))
w_out = 1.0/(np.sqrt(1+len(s2)))
Weight_out[i]=w_out
#for imputing with mean
mean_weight_in = np.mean(list(Weight_in.values()))
mean_weight_out = np.mean(list(Weight_out.values()))
# + colab={} colab_type="code" id="AF4yPhIOVFIY"
if not os.path.isfile('data/fea_sample/storage_sample_stage3.h5'):
#mapping to pandas train
df_final_train['weight_in'] = df_final_train.destination_node.apply(lambda x: Weight_in.get(x,mean_weight_in))
df_final_train['weight_out'] = df_final_train.source_node.apply(lambda x: Weight_out.get(x,mean_weight_out))
#mapping to pandas test
df_final_test['weight_in'] = df_final_test.destination_node.apply(lambda x: Weight_in.get(x,mean_weight_in))
df_final_test['weight_out'] = df_final_test.source_node.apply(lambda x: Weight_out.get(x,mean_weight_out))
#some features engineerings on the in and out weights
df_final_train['weight_f1'] = df_final_train.weight_in + df_final_train.weight_out
df_final_train['weight_f2'] = df_final_train.weight_in * df_final_train.weight_out
df_final_train['weight_f3'] = (2*df_final_train.weight_in + 1*df_final_train.weight_out)
df_final_train['weight_f4'] = (1*df_final_train.weight_in + 2*df_final_train.weight_out)
#some features engineerings on the in and out weights
df_final_test['weight_f1'] = df_final_test.weight_in + df_final_test.weight_out
df_final_test['weight_f2'] = df_final_test.weight_in * df_final_test.weight_out
df_final_test['weight_f3'] = (2*df_final_test.weight_in + 1*df_final_test.weight_out)
df_final_test['weight_f4'] = (1*df_final_test.weight_in + 2*df_final_test.weight_out)
# + colab={} colab_type="code" id="uhxzhQ9aVFIa"
if not os.path.isfile('data/fea_sample/storage_sample_stage3.h5'):
#page rank for source and destination in Train and Test
#if anything not there in train graph then adding mean page rank
df_final_train['page_rank_s'] = df_final_train.source_node.apply(lambda x:pr.get(x,mean_pr))
df_final_train['page_rank_d'] = df_final_train.destination_node.apply(lambda x:pr.get(x,mean_pr))
df_final_test['page_rank_s'] = df_final_test.source_node.apply(lambda x:pr.get(x,mean_pr))
df_final_test['page_rank_d'] = df_final_test.destination_node.apply(lambda x:pr.get(x,mean_pr))
#================================================================================
#Katz centrality score for source and destination in Train and test
#if anything not there in train graph then adding mean katz score
df_final_train['katz_s'] = df_final_train.source_node.apply(lambda x: katz.get(x,mean_katz))
df_final_train['katz_d'] = df_final_train.destination_node.apply(lambda x: katz.get(x,mean_katz))
df_final_test['katz_s'] = df_final_test.source_node.apply(lambda x: katz.get(x,mean_katz))
df_final_test['katz_d'] = df_final_test.destination_node.apply(lambda x: katz.get(x,mean_katz))
#================================================================================
#Hits algorithm score for source and destination in Train and test
#if anything not there in train graph then adding 0
df_final_train['hubs_s'] = df_final_train.source_node.apply(lambda x: hits[0].get(x,0))
df_final_train['hubs_d'] = df_final_train.destination_node.apply(lambda x: hits[0].get(x,0))
df_final_test['hubs_s'] = df_final_test.source_node.apply(lambda x: hits[0].get(x,0))
df_final_test['hubs_d'] = df_final_test.destination_node.apply(lambda x: hits[0].get(x,0))
#================================================================================
#Hits algorithm score for source and destination in Train and Test
#if anything not there in train graph then adding 0
df_final_train['authorities_s'] = df_final_train.source_node.apply(lambda x: hits[1].get(x,0))
df_final_train['authorities_d'] = df_final_train.destination_node.apply(lambda x: hits[1].get(x,0))
df_final_test['authorities_s'] = df_final_test.source_node.apply(lambda x: hits[1].get(x,0))
df_final_test['authorities_d'] = df_final_test.destination_node.apply(lambda x: hits[1].get(x,0))
#================================================================================
hdf = HDFStore('data/fea_sample/storage_sample_stage3.h5')
hdf.put('train_df',df_final_train, format='table', data_columns=True)
hdf.put('test_df',df_final_test, format='table', data_columns=True)
hdf.close()
else:
df_final_train = read_hdf('data/fea_sample/storage_sample_stage3.h5', 'train_df',mode='r')
df_final_test = read_hdf('data/fea_sample/storage_sample_stage3.h5', 'test_df',mode='r')
# + [markdown] colab_type="text" id="p6xkDfD-VFIb"
# ## 5.5 Adding new set of features
#
# __we will create these each of these features for both train and test data points__
# <ol>
# <li>SVD features for both source and destination</li>
# </ol>
# + colab={} colab_type="code" id="WQO6E65eVFIc"
def svd(x, S):
try:
z = sadj_dict[x]
return S[z]
except:
return [0,0,0,0,0,0]
# + colab={} colab_type="code" id="9sOyLwvNVFId"
#for svd features to get feature vector creating a dict node val and inedx in svd vector
sadj_col = sorted(train_graph.nodes())
sadj_dict = { val:idx for idx,val in enumerate(sadj_col)}
# + colab={} colab_type="code" id="zLSt8fGVVFIg"
Adj = nx.adjacency_matrix(train_graph,nodelist=sorted(train_graph.nodes())).asfptype()
# + colab={} colab_type="code" id="soq-VAHlVFIh" outputId="3f9bfb32-004f-4698-e415-469243250130"
U, s, V = svds(Adj, k = 6)
print('Adjacency matrix Shape',Adj.shape)
print('U Shape',U.shape)
print('V Shape',V.shape)
print('s Shape',s.shape)
# -
U[0,:]
np.dot(U[0,:],V[:,0])
# <h2 style="font-family:'Segoe UI';background-color:#a00;color:white"> 5.6 Feature: svd_dot (Assignment features)<br></h2><br>
# Dot product between source node svd and destination node svd features.
# https://storage.googleapis.com/kaggle-forum-message-attachments/2594/supervised_link_prediction.pdf
train_nodes = list(zip(df_final_train.source_node,df_final_train.destination_node))
test_nodes = list(zip(df_final_test.source_node, df_final_test.destination_node))
svd_dot_train_source = []
for v in tqdm(train_nodes):
s, d = v
svd_dot_train_source.append(np.dot(svd(s,U),np.array(svd(d,U)).T)) # svd(s, U) will return svd for source node for U**** svd(d,U).T => is for destination node for V
svd_dot_train_source = np.array(svd_dot_train_source)
df_final_train['svd_dot_source'] = svd_dot_train_source
df_final_train['svd_dot_source']
svd_dot_train_destination = []
for v in tqdm(train_nodes):
s, d = v
svd_dot_train_destination.append(np.dot(svd(s,V.T),np.array(svd(d,V.T)).T)) # svd(s, V) will return svd for source node of V**** svd(d,V.T).T => is destination node for V
svd_dot_train_destination = np.array(svd_dot_train_destination)
df_final_train['svd_dot_destination'] = svd_dot_train_destination
df_final_train['svd_dot_destination']
svd_dot_test_source = []
for v in tqdm(test_nodes):
s,d = v
svd_dot_test_source.append(np.dot(svd(s,U),np.array(svd(d,U)).T))
svd_dot_test_source = np.array(svd_dot_test_source)
df_final_test['svd_dot_source'] = svd_dot_test_source
df_final_test.svd_dot_source
svd_dot_test_destination = []
for v in tqdm(test_nodes):
s,d = v
svd_dot_test_destination.append(np.dot(svd(s,V.T),np.array(svd(d,V.T)).T))
svd_dot_test_destination = np.array(svd_dot_test_destination)
df_final_test['svd_dot_destination'] = svd_dot_test_destination
df_final_test.svd_dot_destinationtination
# <h2 style="font-family:'Segoe UI';background-color:#a00;color:white">5.7 Feature : Preferential Attachment (Assignment feature)<br></h2>
#
# **score(x,y) = |sqrt(x)|.|sqrt(y)|**
# +
def pa_score_followers(x,y):
if x in train_graph:
x_degree = train_graph.in_degree(x)
else:
x_degree=0
if y in train_graph:
y_degree = train_graph.in_degree(y)
else:
y_degree=0
# print(x_degree, y_degree) # for testing purpose
return np.abs(np.sqrt(x_degree))*np.abs(np.sqrt(y_degree))
def pa_score_followee(x,y):
if x in train_graph:
x_degree = train_graph.out_degree(x)
else:
x_degree=0
if y in train_graph:
y_degree = train_graph.out_degree(y)
else:
y_degree=0
# print(x_degree, y_degree) # for testing purpose
return np.abs(np.sqrt(x_degree))*np.abs(np.sqrt(y_degree))
# -
pa_score_followers(27,130) # Preferential attachment score of node no: 27, 30 of our training graph
pa_score_followee(27, 130)
train_nodes = list(zip(df_final_train.source_node,df_final_train.destination_node))
test_nodes = list(zip(df_final_test.source_node, df_final_test.destination_node))
from tqdm import tqdm
# +
train_pref_attach_score_followers = []
for v in tqdm(train_nodes):
i,j = v
train_pref_attach_score_followers.append(pa_score_followers(i,j))
train_pref_attach_score_followers = np.array(train_pref_attach_score_followers)
# -
df_final_train['pa_score_followers']= train_pref_attach_score_followers
# +
train_pref_attach_score_followee = []
for v in tqdm(train_nodes):
i,j = v
train_pref_attach_score_followee.append(pa_score_followee(i,j))
train_pref_attach_score_followee = np.array(train_pref_attach_score_followee)
# -
df_final_train['pa_score_followee'] = train_pref_attach_score_followee
# +
test_pref_attach_score_followers = []
for v in tqdm(test_nodes):
i,j = v
test_pref_attach_score_followers.append(pa_score_followers(i,j))
test_pref_attach_score_followers = np.array(test_pref_attach_score_followers)
# -
df_final_test['pa_score_followers']= test_pref_attach_score_followers
# +
test_pref_attach_score_followee = []
for v in tqdm(test_nodes):
i,j = v
test_pref_attach_score_followee.append(pa_score_followee(i,j))
test_pref_attach_score_followee = np.array(test_pref_attach_score_followee)
# -
df_final_test['pa_score_followee'] = test_pref_attach_score_followee
df_final_test.shape
df_final_test.columns
# + colab={} colab_type="code" id="ls5fqLFhVFIm"
if not os.path.isfile('data/fea_sample/assignment_storage_sample_stage4.h5'):
#===================================================================================================
df_final_train[['svd_u_s_1', 'svd_u_s_2','svd_u_s_3', 'svd_u_s_4', 'svd_u_s_5', 'svd_u_s_6']] = \
df_final_train.source_node.apply(lambda x: svd(x, U)).apply(pd.Series)
df_final_train[['svd_u_d_1', 'svd_u_d_2', 'svd_u_d_3', 'svd_u_d_4', 'svd_u_d_5','svd_u_d_6']] = \
df_final_train.destination_node.apply(lambda x: svd(x, U)).apply(pd.Series)
#===================================================================================================
df_final_train[['svd_v_s_1','svd_v_s_2', 'svd_v_s_3', 'svd_v_s_4', 'svd_v_s_5', 'svd_v_s_6',]] = \
df_final_train.source_node.apply(lambda x: svd(x, V.T)).apply(pd.Series)
df_final_train[['svd_v_d_1', 'svd_v_d_2', 'svd_v_d_3', 'svd_v_d_4', 'svd_v_d_5','svd_v_d_6']] = \
df_final_train.destination_node.apply(lambda x: svd(x, V.T)).apply(pd.Series)
#===================================================================================================
df_final_test[['svd_u_s_1', 'svd_u_s_2','svd_u_s_3', 'svd_u_s_4', 'svd_u_s_5', 'svd_u_s_6']] = \
df_final_test.source_node.apply(lambda x: svd(x, U)).apply(pd.Series)
df_final_test[['svd_u_d_1', 'svd_u_d_2', 'svd_u_d_3', 'svd_u_d_4', 'svd_u_d_5','svd_u_d_6']] = \
df_final_test.destination_node.apply(lambda x: svd(x, U)).apply(pd.Series)
#===================================================================================================
df_final_test[['svd_v_s_1','svd_v_s_2', 'svd_v_s_3', 'svd_v_s_4', 'svd_v_s_5', 'svd_v_s_6',]] = \
df_final_test.source_node.apply(lambda x: svd(x, V.T)).apply(pd.Series)
df_final_test[['svd_v_d_1', 'svd_v_d_2', 'svd_v_d_3', 'svd_v_d_4', 'svd_v_d_5','svd_v_d_6']] = \
df_final_test.destination_node.apply(lambda x: svd(x, V.T)).apply(pd.Series)
#===================================================================================================
hdf = HDFStore('data/fea_sample/assignment_storage_sample_stage4.h5')
hdf.put('train_df_assignment',df_final_train, format='table', data_columns=True)
hdf.put('test_df_assignment',df_final_test, format='table', data_columns=True)
hdf.close()
# + colab={} colab_type="code" id="0-hBtlkzVFIn"
# prepared and stored the data from machine learning models
# pelase check the FB_Models.ipynb
| Facebook challenge/FB_featurization_mdiqbalbajmi00786gmail.com.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Understanding Structured Point Clouds (SPCs)
# Structured Point Clouds (SPC) is a differentiable, GPU-compatible, spatial-data structure which efficiently organizes 3D geometrically sparse information in a very compressed manner.
#
# 
#
# <b> When should you use it? </b>
# * The SPC data structure is very general, which makes it <mark>a suitable building block for a variety of applications</mark>.
# * Examples include:
# * [Representation & rendering of implicit 3D surfaces](https://nv-tlabs.github.io/nglod/)
# * Convolutions on voxels, meshes and point clouds
# * and more..
#
# SPCs are easily convertible from point clouds and meshes, and can be optimized to represent encoded neural implicit fields.
#
# <b> In this tutorial you will learn to: </b>
# 1. Construct a SPC from triangular meshes and point clouds.
# 2. Visualize the SPC using ray-tracing functionality.
# 3. Become familiar with the internals of kaolin's SPC data structure
#
# Practitioners are encouraged to view the [documentation](https://kaolin.readthedocs.io/en/latest/modules/kaolin.ops.spc.html?highlight=spc#kaolin-ops-spc) for additional details about the internal workings of this data structure. <br>
# This tutorial is best run locally to observe the full output.
# ## Setup
# This tutorial assumes a minimal version of [kaolin v0.10.0](https://kaolin.readthedocs.io/en/latest/notes/installation.html). <br>
# In addition, the following libraries are required for this tutorial:
# !pip install -q termcolor
# !pip install -q ipywidgets
# +
from PIL import Image
import torch
import torch.nn.functional as F
import numpy as np
from matplotlib import pyplot as plt
import ipywidgets as widgets
from termcolor import colored
import kaolin as kal
from spc_formatting import describe_octree, color_by_level
# -
# To study the mechanics of the SPC structure, we'll need some auxilary functions (you may skip for now): <br>
# +
def describe_tensor(torch_tensor, tensor_label, with_shape, with_content):
if with_shape:
print(f'"{tensor_label}" is a {torch_tensor.dtype} tensor of size {tuple(torch_tensor.shape)}')
if with_content:
print(f'Raw Content: \n{torch_tensor.cpu().numpy()}')
def convert_texture_to_torch_sample_format(texture):
""" Convert to (1, C, Tex-H, Tex-W) format """
return texture.unsqueeze(0).type(sampled_uvs.dtype).permute(0, 3, 1, 2)
# -
# ### Preliminaries: Load Mesh and sample as Point Cloud
# Throughout this tutorial we'll be using a triangular mesh as an example. <br>
# First, we import the mesh using kaolin:
# +
# Path to some .obj file with textures
mesh_path = "../samples/colored_sphere.obj"
mesh = kal.io.obj.import_mesh(mesh_path, with_materials=True)
print(f'Loaded mesh with {len(mesh.vertices)} vertices, {len(mesh.faces)} faces and {len(mesh.materials)} materials.')
# -
# Next, we'll oversample the mesh faces to make sure our SPC structure is densely populated and avoids "holes"
# at the highest resolution level.
#
# Note that our mesh face-vertices are mapped to some texture coordinates.
# Luckily, kaolin has a `sample_points` function that will take care of interpolating these coordinates for us.
# The sampled vertices will be returned along with the interpolated uv coordinates as well:
# +
# Sample points over the mesh surface
num_samples = 1000000
# Load the uv coordinates per face-vertex like "features" per face-vertex,
# which sample_points will interpolate for new sample points.
# mesh.uvs is a tensor of uv coordinates of shape (#num_uvs, 2), which we consider as "features" here
# mesh.face_uvs_idx is a tensor of shape (#faces, 3), indexing which feature to use per-face-per-vertex
# Therefore, face_features will be of shape (#faces, 3, 2)
face_features = mesh.uvs[mesh.face_uvs_idx]
# Kaolin assumes an exact batch format, we make sure to convert from:
# (V, 3) to (1, V, 3)
# (F, 3, 2) to (1, F, 3, 2)
# where 1 is the batch size
batched_vertices = mesh.vertices.unsqueeze(0)
batched_face_features = face_features.unsqueeze(0)
# sample_points is faster on cuda device
batched_vertices = batched_vertices.cuda()
faces = mesh.faces.cuda()
batched_face_features = batched_face_features.cuda()
sampled_verts, _, sampled_uvs = kal.ops.mesh.trianglemesh.sample_points(batched_vertices,
faces,
num_samples=num_samples,
face_features=batched_face_features)
print(f'Sampled {sampled_verts.shape[1]} points over the mesh surface:')
print(f'sampled_verts is a tensor with batch size {sampled_verts.shape[0]},',
f'with {sampled_verts.shape[1]} points of {sampled_verts.shape[2]}D coordinates.')
print(f'sampled_uvs is a tensor with batch size {sampled_uvs.shape[0]},',
f'representing the corresponding {sampled_uvs.shape[1]} {sampled_uvs.shape[2]}D UV coordinates.')
# -
# To finish our setup, we'll want to use the UV coordinates to perform texture sampling and obtain the RGB color of each point we have:
# +
# Convert texture to sample-compatible format
diffuse_color = mesh.materials[0]['map_Kd'] # Assumes a shape with a single material
texture_maps = convert_texture_to_torch_sample_format(diffuse_color) # (1, C, Th, Tw)
texture_maps = texture_maps.cuda()
# Sample colors according to uv-coordinates
sampled_uvs = kal.render.mesh.utils.texture_mapping(texture_coordinates=sampled_uvs,
texture_maps=texture_maps,
mode='nearest')
# Unbatch
vertices = sampled_verts.squeeze(0)
vertex_colors = sampled_uvs.squeeze(0)
# Normalize to [0,1]
vertex_colors /= 255
print(f'vertices is a tensor of {vertices.shape}')
print(f'vertex_colors is a tensor of {vertices.shape}')
# -
# ## 1. Create & Visualize SPC
# ### Create the SPC
# We start by converting our Point Cloud of continuous 3D coordinates to a Structured Point Cloud. <br>
#
# `unbatched_pointcloud_to_spc` will return a `Spc` object, a data class holding all Structured Point Cloud related information. <br>
# At the core of this object, points are kept in quantized coordinates using a compressed octree. <br>
#
# The returned object contains multiple low-level data structures which we'll explore in details in the next section.
# For now keep in mind that its important fields: `octree`, `features`, `point_hierarchy`, `pyramid` and `prefix`, represent our data structure.
#
# When constructing a `Spc` object, the resolution of quantized coordinates can be controlled by the octree `level` arg, such that: $resolution=2^{level}$
# Our SPC will contain a hierarchy of multiple levels
level = 3
spc = kal.ops.conversions.pointcloud.unbatched_pointcloud_to_spc(vertices, level, features=vertex_colors)
# ### Set-up the camera
# The SPC data structure can be efficiently visualized using ray-tracing ops. <br>
#
# Note that SPC also supports differentiable rendering. In this tutorial we'll limit our demonstration to rendering this data structure efficiently. <br>
# Differentiable ray-tracing is beyond the scope of this guide, and will be covered in future tutorials.
# To begin our ray tracing implementation, we'll first need to set up our camera view and [generate some rays](https://www.scratchapixel.com/lessons/3d-basic-rendering/ray-tracing-generating-camera-rays). <br>
# We'll assume a pinhole camera model, and use the `look_at` function, which sets up a camera view originating at position `camera_from`, looking towards `camera_to`. <br>
# `width`, `height`, `mode` and `fov` will determine the dimensions of our view.
# +
def _normalized_grid(width, height, device='cuda'):
"""Returns grid[x,y] -> coordinates for a normalized window.
Args:
width, height (int): grid resolution
"""
# These are normalized coordinates
# i.e. equivalent to 2.0 * (fragCoord / iResolution.xy) - 1.0
window_x = torch.linspace(-1, 1, steps=width, device=device) * (width / height)
window_y = torch.linspace(1,- 1, steps=height, device=device)
coord = torch.stack(torch.meshgrid(window_x, window_y)).permute(1,2,0)
return coord
def look_at(camera_from, camera_to, width, height, mode='persp', fov=90.0, device='cuda'):
"""Vectorized look-at function, returns an array of ray origins and directions
URL: https://www.scratchapixel.com/lessons/mathematics-physics-for-computer-graphics/lookat-function
"""
camera_origin = torch.FloatTensor(camera_from).to(device)
camera_view = F.normalize(torch.FloatTensor(camera_to).to(device) - camera_origin, dim=0)
camera_right = F.normalize(torch.cross(camera_view, torch.FloatTensor([0,1,0]).to(device)), dim=0)
camera_up = F.normalize(torch.cross(camera_right, camera_view), dim=0)
coord = _normalized_grid(width, height, device=device)
ray_origin = camera_right * coord[...,0,np.newaxis] * np.tan(np.radians(fov/2)) + \
camera_up * coord[...,1,np.newaxis] * np.tan(np.radians(fov/2)) + \
camera_origin + camera_view
ray_origin = ray_origin.reshape(-1, 3)
ray_offset = camera_view.unsqueeze(0).repeat(ray_origin.shape[0], 1)
if mode == 'ortho': # Orthographic camera
ray_dir = F.normalize(ray_offset, dim=-1)
elif mode == 'persp': # Perspective camera
ray_dir = F.normalize(ray_origin - camera_origin, dim=-1)
ray_origin = camera_origin.repeat(ray_dir.shape[0], 1)
else:
raise ValueError('Invalid camera mode!')
return ray_origin, ray_dir
# -
# Now generate some rays using the functions we've just created:
# +
# ray_o and ray_d ~ torch.Tensor (width x height, 3)
# represent rays origin and direction vectors
ray_o, ray_d = look_at(camera_from=[-2.5,2.5,-2.5],
camera_to=[0,0,0],
width=1024,
height=1024,
mode='persp',
fov=30,
device='cuda')
print(f'Total of {ray_o.shape[0]} rays generated.')
# -
# ### Render
# We're now ready to perform the actual ray tracing. <br>
# kaolin will "shoot" the rays for us, and perform an efficient intersection test between each ray and cell within the SPC structure. <br>
# In kaolin terminology, <b>nuggets</b> are "ray-cell intersections" (or rather "ray-point" intersections).
#
# <b>nuggets </b> are of represented by a structure of two tensors: `nugs_ridx` and `nugs_pidx`, <br>which form together pairs of `(index_to_ray, index_to_points)`. <br>
# Both tensors are 1-dimensional tensors of shape (`#num_intersection`,).
octree, features = spc.octrees, spc.features
point_hierarchy, pyramid, prefix = spc.point_hierarchies, spc.pyramids[0], spc.exsum
nugs_ridx, nugs_pidx, depth = kal.render.spc.unbatched_raytrace(octree, point_hierarchy, pyramid, prefix, ray_o, ray_d, level)
print(f'Total of {nugs_ridx.shape[0]} nuggets were traced.\n')
# Since we're assuming here our surface is opaque, for each ray, we only care about the <b>nugget</b>
# closest to the camera. <br>
# Note that per "ray-pack", the returned <b>nuggets</b> are already sorted by depth. <br>
# The method below returns a boolean mask which specifies which <b>nuggets</b> represent a "first-hit".
masked_nugs = kal.render.spc.mark_pack_boundaries(nugs_ridx)
nugs_ridx = nugs_ridx[masked_nugs]
nugs_pidx = nugs_pidx[masked_nugs]
# Finally, for each ray that hit the surface, a corresponding "first-hit" nugget exists. <br>
# +
# 1. We initialize an empty canvas.
image = torch.ones_like(ray_o)
# 2. We'll query all first-hit nuggets to obtain their corresponding point-id (which cell they hit in the SPC).
ridx = nugs_ridx.long()
pidx = nugs_pidx.long() - pyramid[1,level]
# 3. We'll query the features auxilary structure to obtain the color.
# 4. We set each ray value as the corresponding nugget color.
image[ridx] = features[pidx]
image = image.reshape(1024, 1024, 3)
# -
# Putting it all together, we write our complete `render()` function and display the trace using matplotlib:
# +
import matplotlib.pyplot as plt
def render(level):
""" Create & render an image """
spc = kal.ops.conversions.pointcloud.unbatched_pointcloud_to_spc(vertices, level, vertex_colors)
octree, features, point_hierarchy, pyramid, prefix = spc.octrees, spc.features, spc.point_hierarchies, spc.pyramids[0], spc.exsum
nugs_ridx, nugs_pidx, depth = kal.render.spc.unbatched_raytrace(octree, point_hierarchy, pyramid, prefix, ray_o, ray_d, level)
masked_nugs = kal.render.spc.mark_pack_boundaries(nugs_ridx)
nugs_ridx = nugs_ridx[masked_nugs]
nugs_pidx = nugs_pidx[masked_nugs]
ridx = nugs_ridx.long()
pidx = nugs_pidx.long() - pyramid[1,level]
image = torch.ones_like(ray_o)
image[ridx] = features[pidx]
image = image.reshape(1024, 1024, 3)
return image
fig = plt.figure(figsize=(20,10))
# Render left image of level 3 SPC
image1 = render(level=3)
image1 = image1.cpu().numpy().transpose(1,0,2)
ax = fig.add_subplot(1, 2, 1)
ax.set_title("level 3", fontsize=26)
ax.axis('off')
plt.imshow(image1)
# Render right image of level 5 SPC
image2 = render(level=8)
image2 = image2.cpu().numpy().transpose(1,0,2)
ax = fig.add_subplot(1, 2, 2)
ax.set_title("level 5", fontsize=26)
ax.axis('off')
plt.imshow(image2)
plt.show()
# -
# Finally, putting it all together, we may also construct the following interactive demo:
# +
def update_demo(widget_spc_level):
image = render(widget_spc_level)
plt.figure(figsize=(10,10))
plt.axis('off')
plt.imshow(image.cpu().numpy().transpose(1,0,2))
plt.show()
def show_interactive_demo(max_level=10):
start_value = min(7, max_level)
widget_spc_level = widgets.IntSlider(value=start_value, min=1, max=max_level, step=1, orientation='vertical',
description='<h5>SPC Level:</h5>', disabled=False,
layout=widgets.Layout(height='100%',))
out = widgets.interactive_output(update_demo, {'widget_spc_level': widget_spc_level})
display(widgets.HBox([widgets.VBox([widget_spc_level]), out]))
show_interactive_demo()
# -
# ## 2. SPC internals
# In this section we'll explore the various components that make up the [SPC](https://kaolin.readthedocs.io/en/latest/modules/kaolin.ops.spc.html?highlight=spc#structured-point-clouds) we've just created. <br>
# We'll learn how data is stored, and how to view stored data.
# ### Boilerplate code
# Let's rebuild our SPC object with fewer levels, that will make the internals easier to study. <br>
# You may customize the number of levels and compare how the output changes.
level = 3
spc = kal.ops.conversions.pointcloud.unbatched_pointcloud_to_spc(vertices, level, features=vertex_colors)
# Ok, let's see what we've got.
# ### octree
# The first field we'll look into, `octrees`, keeps the entire geometric structure in a compressed manner. <br>
# This is a huge advantage, as this structure is now small enough to fit our sparse data, which makes it very efficient.
octree = spc.octrees
# +
describe_tensor(torch_tensor=octree, tensor_label='octree', with_shape=True, with_content=True)
print(f'\n"octrees" represents a hierarchy of {len(octree)} octree nodes.')
print(f"Let's have a look at the binary representation and what it means:\n")
describe_octree(octree, level)
text_out = widgets.Output(layout={'border': '0.2px dashed black'})
with text_out:
print('How to read the content of octrees?')
print('- Each entry represents a single octree of 8 cells --> 8 bits.')
print('- The bit position determines the cell index, in Morton Order.')
print('- The bit value determines if the cell is occupied or not.')
print(f'- If a cell is occupied, an additional octree may be generated in the next level, up till level {level}.')
print('For example, an entry of 00000001 is a single level octree, where only the bottom-left most cell is occupied.')
display(widgets.HBox([text_out]))
# -
# ##### Order of octants within partitioned cells
# 
# Notice the field is named in plural.
# That's because kaolin can batch multiple instances of octrees together within the same object. <br>
print(spc.batch_size)
# Pay attention that `octrees` uses [packed representation](https://kaolin.readthedocs.io/en/latest/modules/kaolin.ops.batch.html?highlight=packed#packed), meaning, there is no explicit batch dimension. <br>
# Instead, we track the length of each octree instance in a separate field:
# +
octrees_lengths = spc.lengths
describe_tensor(torch_tensor=octrees_lengths, tensor_label='lengths', with_shape=True, with_content=True)
text_out = widgets.Output(layout={'border': '0.2px dashed black'})
with text_out:
print('How to read the content of lengths?')
print(f'- This Spc stores a batch of {len(spc.lengths)} octrees.')
print(f'- The first octree is represented by {spc.lengths[0]} non-leaf cells.')
print(f'- Therefore the information of the first octree is kept in bytes 0-{spc.lengths[0]-1} of the octrees field.')
display(widgets.HBox([text_out]))
# -
# Advanced users who prefer a non object-oriented lower-level api can also use the following functionality which `kal.ops.conversions.pointcloud.unbatched_pointcloud_to_spc` employs under the hood:
# +
from kaolin.ops.spc.points import quantize_points, points_to_morton, morton_to_points, unbatched_points_to_octree
# Construct a batch of 2 octrees. For brevity, we'll use the same ocuupancy data for both.
# 1. Convert continous to quantized coordinates
# 2. Build the octrees
points1 = quantize_points(vertices.contiguous(), level=2)
octree1 = unbatched_points_to_octree(points1, level=2)
points2 = quantize_points(vertices.contiguous(), level=3)
octree2 = unbatched_points_to_octree(points2, level=3)
# Batch 2 octrees together. For packed representations, this is just concatenation.
octrees = torch.cat((octree1, octree2), dim=0)
lengths = torch.tensor([len(octree1), len(octree2)], dtype=torch.int32)
describe_tensor(torch_tensor=octrees, tensor_label='octrees', with_shape=True, with_content=True)
print('')
describe_tensor(torch_tensor=lengths, tensor_label='lengths', with_shape=True, with_content=True)
# -
# These structures form the bare minimum required to shift back to high-level api and construct a Spc object:
kal.rep.spc.Spc(octrees, lengths)
# ### features
# So far we've lookied into how Structured Point Clouds keep track of occupancy. <br>
# Next we'll study how they keep track of features.
# The `features` field contains features information per cell.
features = spc.features
# +
def paint_features(features):
plt.figure(figsize=(10,10))
plt.axis('off')
plt.imshow(features.cpu().numpy()[None])
plt.show()
print('In this tutorial, cell features are RGB colors:')
describe_tensor(torch_tensor=features, tensor_label='features', with_shape=True, with_content=False)
paint_features(features)
text_out = widgets.Output(layout={'border': '0.2px dashed black'})
with text_out:
print('How to read the content of features?')
print(f'- We keep features only for leaf cells, a total of {features.shape[0]}.')
print(f'- The number of leaf cells can be obtained by summarizing the "1" bits at level {spc.max_level},\n' \
' the last level of the octree.')
print(f'- The dimensionality of each attribute is {features.shape[1]} (e.g: RGB channels)')
print('\nReminder - the highest level of occupancy octree is:')
describe_octree(spc.octrees, level, limit_levels=[spc.max_level])
display(widgets.HBox([text_out]))
# -
# ### pyramid & exsum
# Since the occupancy information is [compressed](https://kaolin.readthedocs.io/en/latest/modules/kaolin.ops.spc.html?highlight=spc#octree) and [packed](https://kaolin.readthedocs.io/en/latest/modules/kaolin.ops.batch.html?highlight=packed#packed), accessing level-specific information consistently involves
# cumulative summarization of the number of "1" bits. <br>
# It makes sense to calculate this information once and then cache it. <br>
# The `pyramid` field does exactly that: it keeps summarizes the number of occupied cells per level, and their cumsum, for fast level-indexing.
# A pyramid is kept per octree in the batch.
# We'll study the pyramid of the first and only entry in the batch.
pyramid = spc.pyramids[0]
# +
describe_tensor(torch_tensor=pyramid, tensor_label='pyramid', with_shape=True, with_content=True)
out_left = widgets.Output(layout={'border': '0.2px dashed black'})
out_right = widgets.Output(layout={'border': '0.2px dashed black'})
print('\nHow to read the content of pyramids?')
with out_left:
print('"pyramid" summarizes the number of occupied \ncells per level, and their cumulative sum:\n')
for i in range(pyramid.shape[-1]):
if i ==0:
print(f'Root node (implicitly defined):')
elif i+1 < pyramid.shape[-1]:
print(f'Level #{i}:')
else:
print(f'Final entry for total cumsum:')
print(f'\thas {pyramid[0,i]} occupied cells')
print(f'\tstart idx (cumsum excluding current level): {pyramid[1,i]}')
with out_right:
print(f'"octrees" represents a hierarchy of {len(spc.octrees)} octree nodes.')
print(f"Each bit represents a cell occupancy:\n")
describe_octree(octree, level)
display(widgets.HBox([out_left, out_right]))
# -
# Similarly, kaolin keeps a complementary field, `exsum`, which tracks the cumulative summarization of bits per-octree to fast access parent-child information between levels:
exsum = spc.exsum
# +
describe_tensor(torch_tensor=exsum, tensor_label='exsum', with_shape=True, with_content=True)
out_left = widgets.Output(layout={'border': '0.2px dashed black'})
out_right = widgets.Output(layout={'border': '0.2px dashed black'})
print('\nHow to read the content of exsum?')
with out_left:
print('"exsum" summarizes the cumulative number of occupied \ncells per octree, e.g: exclusive sum of "1" bits:\n')
for i in range(exsum.shape[-1]):
print(f'Cells in Octree #{i} start from cell idx: {exsum[i]}')
with out_right:
print(f'"octrees" represents a hierarchy of {len(octree)} octree nodes.')
print(f"Each bit represents a cell occupancy:\n")
describe_octree(octree, level)
display(widgets.HBox([out_left, out_right]))
# -
# When using Spc objects, pyramids are implicitly created the first time they are needed so you don't have to worry about them. <br>
# For advanced users, the low-level api allows their explicit creation through `scan_octrees`:
# +
lengths = torch.tensor([len(octrees)], dtype=torch.int32)
max_level, pyramid, exsum = kal.ops.spc.spc.scan_octrees(octree, lengths)
print('max_level:')
print(max_level)
print('\npyramid:')
print(pyramid)
print('\nexsum:')
print(exsum)
# -
# ### point_hierarchies
# `point_hierarchies` is an auxilary field, which holds the *sparse* coordinates of each point / occupied cell within the octree, for easier access.
#
# Sparse coordinates are packed for all cells on all levels combined.
describe_tensor(torch_tensor=spc.point_hierarchies, tensor_label='point_hierarchies', with_shape=True, with_content=False)
# We can use the information stored in the pyramids field to color the coordinates by level:
# +
out_left = widgets.Output(layout={'border': '0.2px dashed black'})
out_right = widgets.Output(layout={'border': '0.2px dashed black', 'width': '60%'})
max_points_to_display = 17 # To avoid clutter
with out_left:
level_idx =0
point_idx = 0
remaining_cells_per_level = spc.pyramids[0,0].cpu().numpy().tolist()
for coord in spc.point_hierarchies:
if not remaining_cells_per_level[level_idx]:
level_idx += 1
point_idx = 0
else:
remaining_cells_per_level[level_idx] -= 1
if point_idx == max_points_to_display:
print(colored(f'skipping more..', level_color))
elif point_idx < max_points_to_display:
level_color = color_by_level(level_idx - 1)
print(colored(f'Level #{level_idx}, Point #{point_idx}, ' \
f'Coords: {coord.cpu().numpy().tolist()}', level_color))
point_idx += 1
with out_right:
print('How to read the content of point_hierarchies?')
print(f'- Each cell / point is represented by {spc.point_hierarchies.shape[-1]} indices (xyz).')
print('- Sparse coordinates are absolute: \n they are defined relative to the octree origin.')
print('- Compare the point coordinates with the demo below.\n\n Remember: unoccupied cells are not displayed!')
show_interactive_demo(max_level=spc.max_level)
display(widgets.HBox([out_left, out_right]))
# -
# ## Where to go from here
# Structured Point Clouds support other useful operators which were not covered by this tutorial:
#
# 1. [Convolutions](https://kaolin.readthedocs.io/en/latest/modules/kaolin.ops.spc.html?highlight=SPC#kaolin.ops.spc.Conv3d)
# 2. [Querying points by location](https://kaolin.readthedocs.io/en/latest/modules/kaolin.ops.spc.html?highlight=SPC#kaolin.ops.spc.unbatched_query)
# 3. [Differential ray-tracing ops](https://kaolin.readthedocs.io/en/latest/modules/kaolin.render.spc.html#kaolin-render-spc)
#
| examples/tutorial/understanding_spcs_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="oOxHNI7hdnvw" outputId="439fde24-41a9-4fbc-e20f-8aa745a267dd"
# !pip install d2l==0.17.2
# + id="PQQYzighdpBV"
# implement a dropout_layer function that drops out the elements in the tensor input X with probability dropout
import tensorflow as tf
from d2l import tensorflow as d2l
def dropout_layer(X, dropout):
assert 0 <= dropout <= 1
# In this case, all elements are dropped out
if dropout == 1:
return tf.zeros_like(X)
# In this case, all elements are kept
if dropout == 0:
return X
mask = tf.random.uniform(
shape=tf.shape(X), minval=0, maxval=1) < 1 - dropout
return tf.cast(mask, dtype=tf.float32) * X / (1.0 - dropout)
# + colab={"base_uri": "https://localhost:8080/"} id="AvJVC8xQdpDJ" outputId="ba049865-27ec-487d-f2c7-e8917a5ab9cc"
# pass the input X through the dropout operation, with probabilities 0, 0.5, and 1, respectively
X = tf.reshape(tf.range(16, dtype=tf.float32), (2, 8))
print(X)
print(dropout_layer(X, 0.))
print(dropout_layer(X, 0.5))
print(dropout_layer(X, 1.))
# + id="JA0lBmskdpFR"
# define an MLP with two hidden layers containing 256 units each
num_outputs, num_hiddens1, num_hiddens2 = 10, 256, 256
# + id="p_TavAK0dpG2"
# applies dropout to the output of each hidden layer (following the activation function)
dropout1, dropout2 = 0.2, 0.5
class Net(tf.keras.Model):
def __init__(self, num_outputs, num_hiddens1, num_hiddens2):
super().__init__()
self.input_layer = tf.keras.layers.Flatten()
self.hidden1 = tf.keras.layers.Dense(num_hiddens1, activation='relu')
self.hidden2 = tf.keras.layers.Dense(num_hiddens2, activation='relu')
self.output_layer = tf.keras.layers.Dense(num_outputs)
def call(self, inputs, training=None):
x = self.input_layer(inputs)
x = self.hidden1(x)
if training:
x = dropout_layer(x, dropout1)
x = self.hidden2(x)
if training:
x = dropout_layer(x, dropout2)
x = self.output_layer(x)
return x
net = Net(num_outputs, num_hiddens1, num_hiddens2)
# + colab={"base_uri": "https://localhost:8080/", "height": 655} id="0s_1UrlwdpIr" outputId="639a88f4-25f6-46ab-c53b-e89983075a37"
# !pip uninstall matplotlib
# !pip install --upgrade matplotlib
# + id="dy3mO_i8e_1j"
# + colab={"base_uri": "https://localhost:8080/", "height": 262} id="A6feBRlxdpKU" outputId="065af7d2-be24-4e13-82fc-77bea32e2ef6"
num_epochs, lr, batch_size = 10, 0.5, 256
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size)
trainer = tf.keras.optimizers.SGD(learning_rate=lr)
d2l.train_ch3(net, train_iter, test_iter, loss, num_epochs, trainer)
# + id="yMDfpHgSdpLr"
net = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(256, activation=tf.nn.relu),
# Add a dropout layer after the first fully connected layer
tf.keras.layers.Dropout(dropout1),
tf.keras.layers.Dense(256, activation=tf.nn.relu),
# Add a dropout layer after the second fully connected layer
tf.keras.layers.Dropout(dropout2),
tf.keras.layers.Dense(10),
])
# + colab={"base_uri": "https://localhost:8080/", "height": 262} id="RXy1CTGkdpNo" outputId="d3a4aa1f-a236-4d22-bdb6-f92919e6a491"
# train and test the model
trainer = tf.keras.optimizers.SGD(learning_rate=lr)
d2l.train_ch3(net, train_iter, test_iter, loss, num_epochs, trainer)
# + id="oXNkdZtodpPE"
# + id="VukHYu96dpQl"
# + id="jZh2EcU4dpSa"
# + id="Mv1J-6kMdpU-"
# + id="Q232E8RTdpW6"
# + id="Qqj07kMSdpYy"
# + id="fuonrv_6dpbv"
# + id="sW5YdAY9dpfl"
# + id="BfbSnIkYdphc"
# + id="3hocBcnjdpkz"
| Week 9/4_6_dropout.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python 基础练习3
# ## 用户输入——示例
# 请把最后一行 `run_adder` 从注释拿出来,这样才能运行这段代码。
# +
# 我们定义一个加法计算器
def run_adder():
total = 0
while True:
user_input = input("请输入下一个数。输入 'stop' 停止。")
if user_input == "stop":
break # break 语句会退出当前的循环(如果是多层循环的话只会跳出一层循环)
x = float(user_input) # 这一句可能会导致问题,比如当用户的输入包含字母
total = total + x
print(total)
# run_adder()
# -
# 你会发现,好不容易输入了一大堆数字,一不小心输入一个字母,程序就崩溃,之前累加的数值就全白费了。
# <br>
#
# 所以需要在 `float(user_input)` 这一句之前,判断一下用户输入到底能不能被转换成数字。
# ## 正则表达式——示例
# 注意,我们会用 `\` 来对一些特殊字符进行 `转义`,来去掉它们的特殊含义。
# 比如 `.` 在表达式中是 `通配符` 的意思,会匹配任何字符。加上斜杠后,就退化成字面上的 `.`。
# +
import re
third_decimals = ["3.142", "123.456", "125135316235623.123", "0.456"] # 精确到三位小数的数字
non_third_decimals = ["12", "1.234567", "abc", "123a.133", "ba12"] # 所有其它的字符串
expression = "^[0-9]*\.[0-9]{3}$" # 注意 '*' 后面的 '\.'这个斜杠是必要的,因为 `.` 是一个有特殊含义的字符,就和 `[` 一样
# 不是真的去匹配 `[` 这个字符。这种有特殊含义的字符前面需要加上斜杠来进行 `转义`。
for string in third_decimals:
print(bool(re.search(expression, string)))
print("以上全部为 True")
for string in non_third_decimals:
print(bool(re.search(expression, string)))
print("以上全部为 False")
# -
# 请把最后一行 `run_adder_safe` 从注释拿出来,这样才能运行这段代码。
# +
# 我们定义一个安全的加法计算器
integer_expression = "^[0-9]{1,}$"
def run_adder_safe():
total = 0
while True:
user_input = input("请输入下一个数。输入 'stop' 停止。")
if user_input == "stop":
break
if not re.search(integer_expression, user_input):
print("输入不合法!")
continue # continue 可以直接结束当前这一轮的循环,进入下一轮。
# 因为输入不合法,所以直接重新来一轮循环,从而跳过了下面那句会出错的句子。
x = float(user_input)
total = total + x
print("当前总和: " + str(total))
print(total)
# run_adder_safe()
# -
# ## 正则表达式——练习
#
# +
valid_sort_code = ["24-12-45", "55-34-12", "00-00-11", "55-66-77"] # 合法的 sort code(英国银行卡上用的)
invalid_sort_code = ["aa-22-33", "11 22 33", "15t23g", "314159", "a1-b2-c3"] # 所有其它的字符串
expression = "" # 在这里写表达式!
for string in valid_sort_code:
print(bool(re.search(expression, string)))
print("以上全部为 True")
for string in invalid_sort_code:
print(bool(re.search(expression, string)))
print("以上全部为 False")
# -
# 更多练习请点击[这里](https://regex.sketchengine.co.uk/)!
# ## 用户输入和正则匹配——练习
# +
# 先读取用户输入数字,再读取 '+', '-', '*', '/' 四种字符中的一个
# 然后像计算器那样进行运算,直到用户输入 `stop`
# 注意使用正则表达式检查用户输入是否合法
# 如果觉得困难,就只允许整数。喜欢挑战的话可以尝试一下匹配浮点数。
import re
def run_calculator():
while None:
return None
def isValidNumber(user_input):
return False
def isValidOp(user_input):
return False
run_calculator()
# -
# ## 字典——示例
# +
x = dict()
x["abc"] = 10
x["bcd"] = 20
x["cde"] = 30
print(x.keys())
print(x.values())
print(x.items())
# -
x["abc"] = 100
print(x.values())
print(20 in x.values())
print(200 in x.values())
print("abcd" in x.keys())
# ## 文本处理——示例
# 我们先下载一下《红楼梦》这本书。下面的代码之后要求大家都会写,现在可以先跳过。
# +
# 这一段代码请复制到本地运行,拿到我们需要处理的文本。
# 有兴趣的同学可以研究一下它做了啥。
# 不想看的同学可以先跳过。
# 不过这种代码之后要求会写的。
# 总之就是自动从服务器下载了一个 txt 文件。
import requests
import os
dir_path = 'txt_files'
file_name = 'hongloumeng.txt'
full_path = os.path.join(dir_path, file_name)
if not os.path.exists(dir_path):
os.mkdir(dir_path)
if not os.path.exists(full_path):
url = "http://icewould.com/m5-101/hongloumeng"
print("正在下载文件...")
result = requests.get(url)
with open(full_path, "wb") as f:
f.write(result.content)
print("下载完成")
# -
# 读取文件
lines = None
with open('./txt_files/hongloumeng.txt', 'r', encoding = 'utf-8') as f:
lines = f.readlines()
print(lines[:3]) # 看一下前 3 个段落
print()
print("全书一共 " + str(len(lines)) + " 个段落") # 看一下全书一共有多少段落
content = "".join(lines) # lines 是一个数组,每一个元素是一个段落。现在把它完整拼起来,成为一整个字符串,存放在 content 变量里。
print("全书一共 " + str(len(content)) + " 个字符")
# 通过字典,我们来统计一下,每一个字出现的次数。
# 大家可以把最后的 `print` 从注释里拿出来,打印一下看看。
counter = dict()
for character in content:
if not character in counter.keys():
counter[character] = 1
else:
counter[character] = counter[character] + 1
# print(counter)
# 其实,可以直接使用 `Python` 自带的 `Counter` 进行统计,但是为了让大家熟练掌握 `Dictionary` 的用法,还是建议大家先自己手写。
# 然后,我们看看《红楼梦》中,`贾xx` 或者 `贾x` 字符串共出现了多少次。
import re
expression = "贾[\u4e00-\u9fa5]{1,2}"
result = re.findall(expression, content)
print(len(result))
# 好奇它们都是啥么?我们打印出前几个看一下。
print(result[:100])
# 注意,这些不都是人名哦,比如 `贾政便`。
# 要提取出姓贾的人名,可能需要一番功夫了。
# <br>
#
# 此外,我们可以把重复的删去。这里要用到 `set` 这个数据结构,有兴趣的同学可以去搜一下 `python set`。`set` 其实就是数学里的 `集合`,它不允许有重复的元素。
print(set(result[:100]))
# 一共有多少个不重复的呢?
print(len(set(result)))
# ## 文本处理——练习
# ### 练习1
# 请把我们的《红楼梦》文本中所有不是汉字的东西去掉
#
#
# 匹配所有**非**汉字的正则表达式是:
# `[^\u4e00-\u9fa5]`
#
# 匹配所有汉字的正则表达式是:
# `[\u4e00-\u9fa5]`
#
# 请先不要看以下代码,自己尝试一下。提示,使用 `re.sub` 函数以及 `""`(空字符串)来替换,达到删除的效果。
# +
# 请遮住以下代码,自己先写
import re
def clear_content(content):
expression = '[^\u4e00-\u9fa5]'
result = re.sub(expression, "", content)
return result
result = clear_content(content)
print(result[:300]) # 展示处理完毕后的前 300 个字
# -
# ### 练习2
# 请统计《红楼梦》中出现次数最多的汉字(注意,请先把标点符号去除掉)
# ### 练习 3
# 请统计《红楼梦》中出现次数最多的前三个汉字。
# ### 练习 4
# 请统计《红楼梦》中,黛玉与宝玉两个人名在相隔 20 字内同时出现的次数。
| m5-101/content/python-basics-3-practice/python-basics-3-practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# name: python36964bit01763e9be29f414ba19e1fc9f5b6ec3a
# ---
# +
from __future__ import division
from datetime import datetime, timedelta,date
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# from pyramid.arima import auto_arima
import pmdarima as pm
from pmdarima import model_selection
import warnings
warnings.filterwarnings("ignore")
df_sales = pd.read_csv('/mnt/c/Users/gregoire.jan/OneDrive - Accenture/Documents/Projects/aip/salesforecastaip/data/raw/train.csv')
#represent month in date field as its first day
df_sales['date'] = pd.to_datetime(df_sales['date'])
df_sales['date'] = df_sales['date'].dt.year.astype('str') + '-' + df_sales['date'].dt.month.astype('str') + '-01'
df_sales['date'] = pd.to_datetime(df_sales['date'])
#groupby date and sum the sales
df_sales = df_sales.groupby('date').sales.sum().reset_index()
stepwise_model = pm.auto_arima(df_sales[:-6].sales.values, start_p=1, start_q=1,
max_p=3, max_q=3, m=12,
start_P=0, seasonal=True,
d=1, D=1, trace=True,
error_action='ignore',
suppress_warnings=True,
stepwise=True)
print(stepwise_model.aic())
stepwise_model.fit(df_sales[:-6].sales.values)
stepwise_model.predict(n_periods=6)
| notebooks/predict_arima.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Expressions and Arithmetic
# + [markdown] slideshow={"slide_type": "-"} tags=["remove-cell"]
# **CS1302 Introduction to Computer Programming**
# ___
# + [markdown] slideshow={"slide_type": "slide"}
# ## Operators
# + [markdown] slideshow={"slide_type": "fragment"}
# The followings are common operators you can use to form an expression in Python:
# + [markdown] slideshow={"slide_type": "-"}
# | Operator | Operation | Example |
# | --------: | :------------- | :-----: |
# | unary `-` | Negation | `-y` |
# | `+` | Addition | `x + y` |
# | `-` | Subtraction | `x - y` |
# | `*` | Multiplication | `x*y` |
# | `/` | Division | `x/y` |
# + [markdown] slideshow={"slide_type": "fragment"}
# - `x` and `y` in the examples are called the *left and right operands* respectively.
# - The first operator is a *unary operator*, which operates on just one operand.
# (`+` can also be used as a unary operator, but that is not useful.)
# - All other operators are *binary operators*, which operate on two operands.
# + [markdown] slideshow={"slide_type": "fragment"}
# Python also supports some more operators such as the followings:
# + [markdown] slideshow={"slide_type": "-"}
# | Operator | Operation | Example |
# | -------: | :--------------- | :-----: |
# | `//` | Integer division | `x//y` |
# | `%` | Modulo | `x%y` |
# | `**` | Exponentiation | `x**y` |
# + code_folding=[0] slideshow={"slide_type": "fragment"}
# ipywidgets to demonstrate the operations of binary operators
from ipywidgets import interact
binary_operators = {'+':' + ','-':' - ','*':'*','/':'/','//':'//','%':'%','**':'**'}
@interact(operand1=r'10',
operator=binary_operators,
operand2=r'3')
def binary_operation(operand1,operator,operand2):
expression = f"{operand1}{operator}{operand2}"
value = eval(expression)
print(f"""{'Expression:':>11} {expression}\n{'Value:':>11} {value}\n{'Type:':>11} {type(value)}""")
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** What is the difference between `/` and `//`?
# + [markdown] nbgrader={"grade": true, "grade_id": "integer-division", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
# - `/` is the usual division, and so `10/3` returns the floating-point number $3.\dot{3}$.
# - `//` is integer division, and so `10//3` gives the integer quotient 3.
# + [markdown] slideshow={"slide_type": "fragment"}
# **What does the modulo operator `%` do?**
# + [markdown] slideshow={"slide_type": "-"}
# You can think of it as computing the remainder, but the [truth](https://docs.python.org/3/reference/expressions.html#binary-arithmetic-operations) is more complicated than required for the course.
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** What does `'abc' * 3` mean? What about `10 * 'a'`?
# + [markdown] nbgrader={"grade": true, "grade_id": "concatenation", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
# - The first expression means concatenating `'abc'` three times.
# - The second means concatenating `'a'` ten times.
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** How can you change the default operands (`10` and `3`) for different operators so that the overall expression has type `float`.
# Do you need to change all the operands to `float`?
# + [markdown] nbgrader={"grade": true, "grade_id": "mixed-type", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
# - `/` already returns a `float`.
# - For all other operators, changing at least one of the operands to `float` will return a `float`.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Operator Precedence and Associativity
# + [markdown] slideshow={"slide_type": "fragment"}
# An expression can consist of a sequence of operations performed in a row such as `x + y*z`.
# + [markdown] slideshow={"slide_type": "fragment"}
# **How to determine which operation should be performed first?**
# + [markdown] slideshow={"slide_type": "fragment"}
# Like arithmetics, the order of operations is decided based on the following rules applied sequentially:
# 1. *grouping* by parentheses: inner grouping first
# 1. operator *precedence/priority*: higher precedence first
# 1. operator *associativity*:
# - left associativity: left operand first
# - right associativity: right operand first
# + [markdown] slideshow={"slide_type": "subslide"}
# **What are the operator precedence and associativity?**
# + [markdown] slideshow={"slide_type": "fragment"}
# The following table gives a concise summary:
# + [markdown] slideshow={"slide_type": "-"}
# | Operators | Associativity |
# | :--------------- | :-----------: |
# | `**` | right |
# | `-` (unary) | right |
# | `*`,`/`,`//`,`%` | left |
# | `+`,`-` | left |
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Play with the following widget to understand the precedence and associativity of different operators.
# In particular, explain whether the expression `-10 ** 2*3` gives $(-10)^{2\times 3}= 10^6 = 1000000$.
# + slideshow={"slide_type": "-"}
from ipywidgets import fixed
@interact(operator1={'None':'','unary -':'-'},
operand1=fixed(r'10'),
operator2=binary_operators,
operand2=fixed(r'2'),
operator3=binary_operators,
operand3=fixed(r'3')
)
def three_operators(operator1,operand1,operator2,operand2,operator3,operand3):
expression = f"{operator1}{operand1}{operator2}{operand2}{operator3}{operand3}"
value = eval(expression)
print(f"""{'Expression:':>11} {expression}\n{'Value:':>11} {value}\n{'Type:':>11} {type(value)}""")
# + [markdown] nbgrader={"grade": true, "grade_id": "precedence", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
# The expression evaluates to $(-(10^2))\times 3=-300$ instead because the exponentiation operator `**` has higher precedence than both the multiplication `*` and the negation operators `-`.
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** To avoid confusion in the order of operations, we should follow the [style guide](https://www.python.org/dev/peps/pep-0008/#other-recommendations) when writing expression.
# What is the proper way to write `-10 ** 2*3`?
# + nbgrader={"grade": true, "grade_id": "pep8", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
print(-10**2 * 3) # can use use code-prettify extension to fix incorrect styles
print((-10)**2 * 3)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Augmented Assignment Operators
# + [markdown] slideshow={"slide_type": "fragment"}
# - For convenience, Python defines the [augmented assignment operators](https://docs.python.org/3/reference/simple_stmts.html#grammar-token-augmented-assignment-stmt) such as `+=`, where
# - `x += 1` means `x = x + 1`.
# + [markdown] slideshow={"slide_type": "fragment"}
# The following widgets demonstrate other augmented assignment operators.
# + slideshow={"slide_type": "-"}
from ipywidgets import interact, fixed
@interact(initial_value=fixed(r'10'),
operator=['+=','-=','*=','/=','//=','%=','**='],
operand=fixed(r'2'))
def binary_operation(initial_value,operator,operand):
assignment = f"x = {initial_value}\nx {operator} {operand}"
_locals = {}
exec(assignment,None,_locals)
print(f"""Assignments:\n{assignment:>10}\nx: {_locals['x']} ({type(_locals['x'])})""")
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Can we create an expression using (augmented) assignment operators? Try running the code to see the effect.
# + slideshow={"slide_type": "-"}
3*(x = 15)
# + [markdown] nbgrader={"grade": true, "grade_id": "assignment-statement", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
# Assignment operators are used in assignment statements, which are not expressions because they cannot be evaluated.
| _build/jupyter_execute/Lecture2/Expressions and Arithmetic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
print('UbitName: aakashgu')
print('Person Number : 50289067')
import cv2
from matplotlib import pyplot as plt
#reading in rgb format
airplane = cv2.imread('task1.png', cv2.IMREAD_COLOR)
airplane = cv2.cvtColor(airplane, cv2.COLOR_RGB2GRAY)
cv2.namedWindow('image', cv2.WINDOW_NORMAL)
cv2.imshow('image', airplane)
cv2.waitKey(0)
cv2.destroyAllWindows()
print(type(airplane))
print('Shape :',airplane.shape)
print('Size : ',airplane.size)
print(airplane[0])
print(type(airplane))
print('Shape :',airplane.shape)
print('Rows :',airplane.shape[0])
print('Columns :',airplane.shape[1])
print('Size : ',airplane.size)
print(type(airplane[0][0]))
img = list(airplane)
print(type(img))
print(len(img))
print(len(img[0]))
# ## Project1
# 1)Edge Detection
# 2)Keypoint Detection
# 3)Cursor Detection
# ### Edge detection -
# Write programs to detect edges in Fig. 1 (along both x and y directions) using Sobel operator.
#
# #### What is sobel operator?
# Sobel operator or sobel filter is used for edge detection in computer vision or image processing.
# ### Formulation
# The operator uses two 3×3 kernels which are convolved with the original image to calculate approximations of the derivatives – one for horizontal changes, and one for vertical. If we define A as the source image, and Gx and Gy are two images which at each point contain the horizontal and vertical derivative approximations respectively, the computations are as follows:
plt.imshow(cv2.imread('sobel_wiki.JPG', cv2.IMREAD_COLOR))
plt.show()
plt.imshow(cv2.imread('convolution_operation.JPG', cv2.IMREAD_COLOR))
plt.show()
# +
sobel_x = [[1 , 0 , -1],[2 , 0 , -2],[1 , 0, -1]]
#testing array from http://www.songho.ca/dsp/convolution/convolution2d_example.html
#sobel_x = [[-1 , -2 , -1],[0 , 0 , 0],[1 , 2, 1]]
print(sobel_x)
print(type(sobel_x))
print('Sobel Kernel Rows : ',len(sobel_x))
print('Sobel Kernal Columns :',len(sobel_x[0]))
#Where h is the matrix on which convolution is being applied that airplane picture and
# F is operation matrix. In this case it is sobel matrix
# +
#img = [[1,2,3],[4,5,6],[7,8,9]]
#img = [[a,b,c],[d,e,f],[g,h,i]]
#img = np.array([[1,2,31,4,5],[6,0,81,9,10],[11,1,131,14,15],[16,17,181,19,20],[21,22,231,24,25]])
#img = [[0,81,9],[1,131,14],[17,181,19]]
u = len(img)#image rows
v = len(img[0])#image columns
len_k_row = len(sobel_x)
len_k_col = len(sobel_x[0])
print('row :',u,'\ncolumn :',v)
# -
print(sobel_x[0][0])
import numpy as np
#sobel_x_matrix = np.zeros_like(img)
#print(sobel_x_matrix)
offset = len(sobel_x)//2
sobel_matrix_list = []
row_list = []
sum_all = 0
import copy
for h_row in range(0,u):
for h_column in range(0,v):
sum_all = 0
#print('New col')
#print()
row_list.append(sum_all)
for k_row in range(0,len_k_row):
k_row_flip = len_k_row -1 -k_row
for k_column in range(0,len_k_col):
k_col_flip = len_k_col -1 - k_column
#to take care of kernel outside image index
if (k_row_flip-h_row > offset or k_col_flip-h_column > offset or h_row-k_row_flip+offset>=u or h_column-k_col_flip+offset >= v):
#print()
continue
else:
#sobel_x_matrix[h_row][h_column] += sobel_x[k_row_flip][k_col_flip]*img[h_row-k_row_flip+offset][h_column-k_col_flip+offset]
row_list[len(row_list)-1]+= sobel_x[k_row_flip][k_col_flip]*img[h_row-k_row_flip+offset][h_column-k_col_flip+offset]
sobel_matrix_list.append(copy.deepcopy(row_list))
row_list.clear()
#if sobel_x_matrix[h_row][h_column] <0 :
# print('-ve')
sobel_x_matrix = copy.deepcopy(np.asanyarray(sobel_matrix_list))
print(sobel_x_matrix)
# +
#print(sobel_x_matrix[0])
#print(sobel_matrix_list[0])
# -
print(type(sobel_x_matrix))
print(len(sobel_matrix_list))
print(len(sobel_matrix_list[0]))
# Eliminate zero values with method 1
pos_edge_x = (sobel_x_matrix - np.min(sobel_x_matrix)) / (np.max(sobel_x_matrix) - np.min(sobel_x_matrix))
cv2.namedWindow('pos_edge_x_dir', cv2.WINDOW_NORMAL)
cv2.imshow('pos_edge_x_dir', pos_edge_x)
cv2.waitKey(0)
cv2.destroyAllWindows()
# Eliminate zero values with method 2
pos_edge_x = np.abs(sobel_x_matrix) / np.max(np.abs(sobel_x_matrix))
cv2.namedWindow('pos_edge_x_dir', cv2.WINDOW_NORMAL)
cv2.imshow('pos_edge_x_dir', pos_edge_x)
cv2.waitKey(0)
cv2.destroyAllWindows()
# +
sobel_y = [[1 , 2, 1],[0 , 0 , 0],[-1 , -2 , -1]]
print(sobel_y)
print(type(sobel_y))
print('Sobel Kernel Rows : ',len(sobel_y))
print('Sobel Kernal Columns :',len(sobel_y[0]))
print(sobel_y[0][0])
offset = len(sobel_y)//2
sobel_y_matrix_list = []
row_list_sobel_y = []
sum_all = 0
for h_row in range(0,u):
for h_column in range(0,v):
sum_all = 0
#print('New col')
#print()
row_list_sobel_y.append(sum_all)
for k_row in range(0,len_k_row):
k_row_flip = len_k_row -1 -k_row
for k_column in range(0,len_k_col):
k_col_flip = len_k_col -1 - k_column
#to take care of kernel outside image index
if (k_row_flip-h_row > offset or k_col_flip-h_column > offset or h_row-k_row_flip+offset>=u or h_column-k_col_flip+offset >= v):
#print()
continue
else:
#sobel_x_matrix[h_row][h_column] += sobel_x[k_row_flip][k_col_flip]*img[h_row-k_row_flip+offset][h_column-k_col_flip+offset]
row_list_sobel_y[len(row_list_sobel_y)-1]+= sobel_y[k_row_flip][k_col_flip]*img[h_row-k_row_flip+offset][h_column-k_col_flip+offset]
sobel_y_matrix_list.append(copy.deepcopy(row_list_sobel_y))
row_list_sobel_y.clear()
#if sobel_x_matrix[h_row][h_column] <0 :
# print('-ve')
sobel_y_matrix = copy.deepcopy(np.asanyarray(sobel_y_matrix_list))
print(sobel_y_matrix)
# -
print(type(sobel_y_matrix))
print(len(sobel_y_matrix_list))
print(len(sobel_y_matrix_list[0]))
# +
# Eliminate zero values with method 1
pos_edge_y = (sobel_y_matrix - np.min(sobel_y_matrix)) / (np.max(sobel_y_matrix) - np.min(sobel_y_matrix))
cv2.namedWindow('pos_edge_y_dir', cv2.WINDOW_NORMAL)
cv2.imshow('pos_edge_y_dir', pos_edge_y)
cv2.waitKey(0)
cv2.destroyAllWindows()
# Eliminate zero values with method 2
pos_edge_y = np.abs(sobel_y_matrix) / np.max(np.abs(sobel_y_matrix))
cv2.namedWindow('pos_edge_y_dir', cv2.WINDOW_NORMAL)
cv2.imshow('pos_edge_y_dir', pos_edge_y)
cv2.waitKey(0)
cv2.destroyAllWindows()
# magnitude of edges (conbining horizontal and vertical edges)
edge_magnitude = np.sqrt(sobel_x_matrix ** 2 + sobel_y_matrix ** 2)
edge_magnitude /= np.max(edge_magnitude)
cv2.namedWindow('edge_magnitude', cv2.WINDOW_NORMAL)
cv2.imshow('edge_magnitude', edge_magnitude)
cv2.waitKey(0)
cv2.destroyAllWindows()
edge_direction = np.arctan(sobel_y_matrix / (sobel_x_matrix + 1e-3))
edge_direction = edge_direction * 180. / np.pi
edge_direction /= np.max(edge_direction)
cv2.namedWindow('edge_direction', cv2.WINDOW_NORMAL)
cv2.imshow('edge_direction', edge_magnitude)
cv2.waitKey(0)
cv2.destroyAllWindows()
print("Original image size: {:4d} x {:4d}".format(len(img), len(img[1])))
print("Resulting image size: {:4d} x {:4d}".format(edge_magnitude.shape[0], edge_magnitude.shape[1]))
# -
| Project1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="8irlm3BTEQVo" executionInfo={"status": "ok", "timestamp": 1632692679138, "user_tz": 360, "elapsed": 980, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}} outputId="92648d32-c267-4ec0-9567-d8b885b92f71"
# !pip list | grep soup
# + id="av5xYKiAEHwE" executionInfo={"status": "ok", "timestamp": 1632694990754, "user_tz": 360, "elapsed": 174, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}}
import requests
import pandas as pd
import dateutil
from tqdm import tqdm
from time import sleep
from bs4 import BeautifulSoup
# + id="WRW3MCBgEfSp" executionInfo={"status": "ok", "timestamp": 1632694991067, "user_tz": 360, "elapsed": 162, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}}
def scrape_creation_date(subreddit):
"""
Go visit the given subreddit
return the date the subreddit was created
like yyyy-mm-dd
"""
headers = {
'User-Agent': 'Mozilla/5.0'
}
url = 'https://www.reddit.com/r/' + subreddit
r = requests.get(url, headers=headers)
if r.status_code != 200:
raise ValueError('status code is ' + r.status_code)
soup = BeautifulSoup(r.text)
# This is the class of the div that holds the creation date
# as of September 26, 2021
cake_attr = {'class':'_2QZ7T4uAFMs_N83BZcN-Em'}
cake_div = soup.find_all(attrs=cake_attr)
if cake_div == []:
return 'Unable to locate cake div'
cake_txt = cake_div[0].getText().replace('Created ','')
return dateutil.parser.parse(cake_txt).strftime('%Y-%m-%d')
# + colab={"base_uri": "https://localhost:8080/"} id="D5ZuTbqoKIIt" executionInfo={"status": "ok", "timestamp": 1632708600079, "user_tz": 360, "elapsed": 13091701, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}} outputId="2b8f5b3f-7a81-4ed2-dddd-dd3a4264ee0a"
df = pd.read_csv('extra_clean.csv')
subs = df['subreddit']
cakes = {}
for sub in tqdm(subs, ascii=True):
try:
cakes[sub] = scrape_creation_date(sub)
sleep(1)
except ValueError:
sleep(2)
cakes[sub] = scrape_creation_date(sub)
except:
cakes[sub] = 'Some other error'
# + colab={"base_uri": "https://localhost:8080/"} id="j-6uRD1eMnY2" executionInfo={"status": "ok", "timestamp": 1632708622735, "user_tz": 360, "elapsed": 142, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}} outputId="c0b4b7a1-8d38-4ea2-85be-1f278b0bfc01"
len(cakes.keys())
# + colab={"base_uri": "https://localhost:8080/"} id="FJQ37RnUBHgz" executionInfo={"status": "ok", "timestamp": 1632708729951, "user_tz": 360, "elapsed": 121, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}} outputId="9b063a7b-3c1a-4510-fd8b-7e8ddf3c70c8"
errors = 0
for k, v in cakes.items():
if v == 'Some other error' or v == 'Unable to locate cake div':
errors += 1
errors
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="yzZbAUJ6BUpb" executionInfo={"status": "ok", "timestamp": 1632708952737, "user_tz": 360, "elapsed": 125, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}} outputId="76996d44-4a8e-4dce-fe6a-87865ac490f7"
new_df = pd.DataFrame.from_dict(cakes, orient='index', columns=['creation_date'])
new_df = new_df.reset_index()
new_df[['subreddit', 'creation_date']] = new_df[['index', 'creation_date']]
new_df = new_df.drop(columns=['index'])
new_df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="822MNH54BVis" executionInfo={"status": "ok", "timestamp": 1632709226391, "user_tz": 360, "elapsed": 120, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}} outputId="d68b5a01-c4db-4977-9622-abe698d7950c"
join_df = pd.merge(df, new_df, how='inner', on='subreddit')
join_df.head()
# + id="WFcITz2FDa5i" executionInfo={"status": "ok", "timestamp": 1632709258145, "user_tz": 360, "elapsed": 121, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16755363354285896480"}}
join_df.to_csv('plus_creation_date.csv')
# + id="lY_MTA7dDipf"
| scrape_creation_date.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import emoji
from glove_vec import read_glove_vectors
from keras.models import Sequential
from keras.layers import Dense, Input, Dropout, SimpleRNN, LSTM, Activation
from keras.utils import np_utils
import matplotlib.pyplot as plt
# -
train = pd.read_csv("train_emoji.csv", header=None)
test = pd.read_csv("test_emoji.csv", header=None)
train.head()
test.head()
emoji_dict = {0 : ":heart:", 1 : ":baseball:", 2 : ":smile:",
3 : ":disappointed:", 4 : ":fork_and_knife:"}
for ix in emoji_dict.keys():
print (ix, emoji.emojize(emoji_dict[ix],use_aliases=True))
# +
X_train = train[0]
Y_train = train[1]
X_test = test[0]
Y_test = test[1]
print (X_train.shape, Y_train.shape)
print (X_test.shape, Y_test.shape)
# -
print (X_train[0], Y_train[0])
# +
for ix in range(X_train.shape[0]):
X_train[ix] = X_train[ix].split()
for ix in range(X_test.shape[0]):
X_test[ix] = X_test[ix].split()
Y_train = np_utils.to_categorical(Y_train)
# -
print (X_train[0], Y_train[0])
np.unique(np.array([len(ix) for ix in X_train]), return_counts=True)
np.unique(np.array([len(ix) for ix in X_test]), return_counts=True)
emb_matrix = read_glove_vectors("/home/vasu/all_projects/CB/glove.6B.50d.txt")
print (emb_matrix["i"], emb_matrix["i"].shape)
# +
embeddings_train = np.zeros((X_train.shape[0], 10, 50))
embeddings_test = np.zeros((X_test.shape[0], 10, 50))
for ix in range(X_train.shape[0]):
for ij in range(len(X_train[ix])):
embeddings_train[ix][ij] = emb_matrix[X_train[ix][ij].lower()]
for ix in range(X_test.shape[0]):
for ij in range(len(X_test[ix])):
embeddings_test[ix][ij] = emb_matrix[X_test[ix][ij].lower()]
# -
print (embeddings_train.shape, embeddings_test.shape)
# +
model = Sequential()
model.add(LSTM(32, input_shape=(10,50), return_sequences=True))
model.add(Dropout(0.5))
model.add(LSTM(32, return_sequences=False))
model.add(Dropout(0.5))
model.add(Dense(5, activation='softmax'))
model.summary()
# -
model.compile(loss="categorical_crossentropy", optimizer="RMSprop", metrics=['accuracy'])
hist = model.fit(embeddings_train, Y_train, epochs=50, batch_size=12,shuffle=True, validation_split=0.05)
pred = model.predict_classes(embeddings_test)
acc = float(sum(pred==Y_test))/embeddings_test.shape[0]
print(acc)
def print_emoji(label):
return emoji.emojize(emoji_dict[label], use_aliases=True)
for ix in range(embeddings_test.shape[0]):
if pred[ix] != Y_test[ix]:
print (X_test[ix], print_emoji(Y_test[ix]), print_emoji(pred[ix]))
| RNN_LSTM_Emoji_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression: Using a Decomposition (Cholesky Method)
# --------------------------------
#
# This script will use TensorFlow's function, `tf.cholesky()` to decompose our design matrix and solve for the parameter matrix from linear regression.
#
# For linear regression we are given the system $A \cdot x = y$. Here, $A$ is our design matrix, $x$ is our parameter matrix (of interest), and $y$ is our target matrix (dependent values).
#
# For a Cholesky decomposition to work we assume that $A$ can be broken up into a product of a lower triangular matrix, $L$ and the transpose of the same matrix, $L^{T}$.
#
# Note that this is when $A$ is square. Of course, with an over determined system, $A$ is not square. So we factor the product $A^{T} \cdot A$ instead. We then assume:
#
# $$A^{T} \cdot A = L^{T} \cdot L$$
#
# For more information on the Cholesky decomposition and it's uses, see the following wikipedia link: [The Cholesky Decomposition](https://en.wikipedia.org/wiki/Cholesky_decomposition)
#
# Given that $A$ has a unique Cholesky decomposition, we can write our linear regression system as the following:
#
#
# \begin{align}
# A \cdot x &= y \\
# \Longleftrightarrow A^T \cdot A \cdot x &= A^T \cdot y \\
# \Longleftrightarrow L^{T} \cdot L \cdot x &= A^{T} \cdot y
# \end{align}
#
# Then we break apart the system as follows:
#
# $$L^{T} \cdot z = A^{T} \cdot y$$
#
# and
#
# $$L \cdot x = z$$
#
# The steps we will take to solve for $x$ are the following
#
# 1. Compute the Cholesky decomposition of $A$, where $A^{T} \cdot A = L^{T} \cdot L$.
#
# 2. Solve ($L^{T} \cdot z = A^{T} \cdot y$) for $z$.
#
# 3. Finally, solve ($L \cdot x = z$) for $x$.
#
# We start by loading the necessary libraries.
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow.python.framework import ops
ops.reset_default_graph()
# Next we create a graph session
sess = tf.Session()
# We use the same method of generating data as in the prior recipe for consistency.
# Create the data
x_vals = np.linspace(0, 10, 100)
y_vals = x_vals + np.random.normal(0, 1, 100)
# We generate the design matrix, $A$.
# Create design matrix
x_vals_column = np.transpose(np.matrix(x_vals))
ones_column = np.transpose(np.matrix(np.repeat(1, 100)))
A = np.column_stack((ones_column, x_vals_column))
# Next, we generate the
# +
# Create y matrix
y = np.transpose(np.matrix(y_vals))
# Create tensors
A_tensor = tf.constant(A)
y_tensor = tf.constant(y)
# -
# Now we calculate the square of the matrix $A$ and the Cholesky decomposition.
# Find Cholesky Decomposition
tA_A = tf.matmul(tf.transpose(A_tensor), A_tensor)
L = tf.cholesky(tA_A) # tf.cholesky returns L only. L is Lower triangular matrix
# We solve the first equation. (see step 2 in the intro paragraph above)
# Solve L*z=t(A)*y
tA_y = tf.matmul(tf.transpose(A_tensor), y)
z = tf.matrix_solve(L, tA_y)
# We finally solve for the parameter matrix by solving the second equation (see step 3 in the intro paragraph).
# +
# Solve L' * sol = z
sol = tf.matrix_solve(tf.transpose(L), z)
solution_eval = sess.run(sol) # shape=(2, 1)
# -
# Extract the coefficients and create the best fit line.
# +
# Extract coefficients
slope = solution_eval[1][0]
y_intercept = solution_eval[0][0]
print('slope: ' + str(slope))
print('y_intercept: ' + str(y_intercept))
# Get best fit line
best_fit = []
for i in x_vals:
best_fit.append(slope*i+y_intercept)
# -
# Finally, we plot the fit with Matplotlib.
# Plot the results
plt.style.use("ggplot")
plt.plot(x_vals, y_vals, 'o', label='Data')
plt.plot(x_vals, best_fit, 'b-', label='Best fit line', linewidth=3)
plt.legend(loc='upper left')
plt.show()
| 03_Linear_Regression/02_Implementing_a_Decomposition_Method/02_lin_reg_decomposition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Part 0: Jupyter Notebooks
#
# 1. What are they?
# 2. Why are they?
# 2. How do they work?
# + [markdown] slideshow={"slide_type": "subslide"}
# # What are they?
#
# https://jupyter.org/
# - Web-based interface to combine Markdown, Code inputs, Outputs
# - Use Python (or other languages) outside of the terminal
# - This has grown in the scientific Python community into a larger ecosystem (including JupyterHub and Binder)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Why are they?
#
# - Self-documenting workflows
# - Reproducible
# - With good Markdown/Outputs they ARE reports
# + [markdown] slideshow={"slide_type": "subslide"}
# # How do they work?
#
# - Enter code in *code* cells
# - Click `>Run` to execute a code cell
# - You can also hit `Ctrl+Enter` to execute the current cell
# - Or `Shift+Enter` to execute a cell and move down one
# - There are lots of keyboard shortcuts in the *Help* menu
# + [markdown] slideshow={"slide_type": "subslide"}
# # How do they work?
#
# - Enter Markdown in *markdown* cells
#
# ## Markdown
#
# - A **shortform** for `HTML`, we can use \*'s and \_'s to quickly emphasize things
# - Bullets are also easy
#
# If there are 3 things to remember, it would be:
#
# 1. Don't overthink it, just adding in some \#'s and \-'s looks pretty good
# 1. This is how you make numbered lists
# 6. The actual number out front doesn't matter
# + [markdown] slideshow={"slide_type": "slide"}
# # Is that it?
#
# Not really, there's lots of great features and extensions for Jupyter Notebooks that make them a great part of a Data Science workflow
#
# e.g. Jupyter RISE allows this to be a live code slideshow
#
# and --> *exercise* let's us do our exercises. Just enable this below (and refresh the page)!
# + slideshow={"slide_type": "-"}
# !jupyter contrib nbextension install --sys-prefix
# + slideshow={"slide_type": "-"}
# !jupyter nbextension enable exercise2/main
# + [markdown] slideshow={"slide_type": "slide"} solution2="hidden" solution2_first=true
# # EXERCISE
# + solution2="hidden"
# Hidden solution?
# + slideshow={"slide_type": "-"}
# Your Solution
a = 1
print(a * 2)
| Full/Part 1 - Jupyter Notebooks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Merkle's Puzzles
# In cryptography, Merkle's puzzles is an early construction for a public-key cryptosystem, a protocol devised by <NAME> in 1974 and published in 1978. It allows two parties to agree on a shared secret by exchanging messages, even if they have no secrets in common beforehand.
#
# ## High Level Description
# 1. Bob generates 2^N messages containing, "This is message X. This is the symmetrical key, Y", where X is an identifier, and Y is a secret key meant for symmetrical encryption. Both X and Y are unique to each message. All the messages are encrypted in a way such that a user may conduct a brute force attack on each message with some difficulty. Bob sends all the encrypted messages to Alice.
# 1. Alice receives all the encrypted messages, and randomly chooses a single message to brute force. After Alice discovers both the identifier X and the secret key Y inside that message, she encrypts her clear text with the secret key Y, and sends that identifier (in cleartext) with her cipher text to Bob.
# 1. Bob finds the secret key paired with that identifier, and deciphers Alice's cipher text with that secret key.
# ## Implementation
#
# Python implementation using [AES](https://en.wikipedia.org/wiki/Advanced_Encryption_Standard) for symmetric encryption and secret module for [CSPRNG](https://en.wikipedia.org/wiki/Cryptographically_secure_pseudorandom_number_generator). The key used for the encryption is just a 256bit random number. Also for simplicity AES is used in [ECB](https://en.wikipedia.org/wiki/Block_cipher_mode_of_operation) mode which is not recommended for secure systems.
import secrets
from Crypto.Cipher import AES
# Define some constants here. The puzzle difficulty is set to 8 bit which is very easy and in real system must be bigger.
# The complexity of the entire puzzle depends also on the number of messages generated because an adversary could be able to solve all the puzzles and to find the PID sent by Alice to Bob.
PUZZLE_ID = b'AES Puzzle'
PUZZLE_BYTES = 16 # 128 bits
PUZZLE_DIFFICULTY = 8 # bits
MESSAGES_NUM = 2 ** 10
HASH_BYTES = 32 # 256 bits
keys = {}
puzzles = []
# ## Functions
"""
This function pads a byte array to length of *16
"""
def pad16(b):
n = 16-len(b) % 16;
r = secrets.randbits(n*8).to_bytes(n, 'big')
return b + r
"""
Creates a puzzle using AES cipher. Generates a random PUZZLE_DIFFICULTY bits which is filled up to PUZZLE_BYTES with zeros.
Then PK is generated which will be the private key for the communication which only Alice and Bob will know. PID is generated
to be the index of PK.
"""
def createPuzzle():
key = secrets.randbits(PUZZLE_DIFFICULTY).to_bytes(PUZZLE_BYTES, 'big')
cipher = AES.new(key, AES.MODE_ECB)
pid = secrets.randbits(HASH_BYTES*8).to_bytes(HASH_BYTES, 'big')
pk = secrets.randbits(HASH_BYTES*8).to_bytes(HASH_BYTES, 'big')
msg = PUZZLE_ID + pid + pk
puzzle = cipher.encrypt(pad16(msg))
return puzzle, pid, pk
"""
Attempt to brute force a puzzle. Returns PID and PK, as well as bool if successful
"""
def bruteForce(puzzle):
for i in range(0, 2 ** PUZZLE_DIFFICULTY):
key = i.to_bytes(PUZZLE_BYTES, 'big')
cipher = AES.new(key, AES.MODE_ECB)
msg = cipher.decrypt(puzzle)
if msg.find(PUZZLE_ID) == 0:
offset = len(PUZZLE_ID)
pid = msg[offset:offset+HASH_BYTES]
pk = msg[offset+HASH_BYTES:offset+HASH_BYTES*2]
return True, pid, pk
return False
"""
Encrypts a message using private key PK. PID is appended to the message to be used as index from the other party.
Message length is also included since the padding changes the original message
"""
def encryptMessage(pid, pk, msg):
cipher = AES.new(pk, AES.MODE_ECB)
msg_len = len(msg).to_bytes(4, 'big')
return pid + msg_len + cipher.encrypt(pad16(msg))
"""
Decrypts a message. The message format must be PID + MESSAGE_LEN + AES(message)
"""
def descryptMessage(secret):
pid = secret[:HASH_BYTES]
msg_len = int.from_bytes(secret[HASH_BYTES:HASH_BYTES+4], 'big')
msg = secret[HASH_BYTES+4:]
pk = keys[pid]
cipher = AES.new(pk, AES.MODE_ECB)
decrypted = cipher.decrypt(msg)
return decrypted[:msg_len]
# ## Example
# 1. Bob generates 2^N messages
for i in range(0, MESSAGES_NUM):
puzzle, pid, pk = createPuzzle()
keys[pid] = pk
puzzles.append(puzzle)
'Bob has generated {} puzzles'.format(len(puzzles))
# 2. Alice receives all the encrypted messages, and randomly chooses a single message to brute force
try_puzzle = secrets.choice(puzzles)
'Alice has chosen a puzzle: ' + try_puzzle.hex()
# +
cracked, pid, pk = bruteForce(try_puzzle)
if not cracked:
raise Error('Hmm, something went wrong')
'Alice has cracked the puzzle and found: PID:{} and PK:{}'.format(pid.hex(), pk.hex())
# -
secret = encryptMessage(pid, pk, b'Hi Bob, this is secure channel')
'Alice has sent encrypted message to Bob sayng "Hi Bob, this is secure channel"'
# 3. Bob finds the secret key paired with that identifier, and deciphers Alice's cipher text with that secret key.
decrypted = descryptMessage(secret)
'Bob has decrypted the message and it was: "{}"'.format(str(decrypted, 'utf-8'))
# ## References
#
# 1. Wikipedia, "Merkle's Puzzles", https://en.wikipedia.org/wiki/Merkle%27s_Puzzles
| Cryptography/Merkle Puzzles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div class ="alert alert-warning">
#
#
# notebook consultable, exécutable, modifiable et téléchargeable en ligne :
#
#
# - se rendre à : https://github.com/nsi-acot/continuite_pedagogique_premiere
# - cliquer sur l'icone "launch binder" en bas de page
# - patienter quelques secondes que le serveur Jupyter démarre
# - naviguer dans le dossier `"./types_construits/dictionnaires/"`
# - cliquer sur le nom de ce notebook
# </div>
# # <center> Dictionnaires</center>
# ### Définition et création
# Les types de données construits que nous avons abordés jusqu’à présent (chaînes, listes et tuples) sont tous des séquences, c’est-à-dire des suites ordonnées d’éléments. Dans une séquence, il est facile
# d’accéder à un élément quelconque à l’aide d’un index (un nombre entier), mais encore faut-il le connaître.
#
# Les dictionnaires constituent un autre type construit. Ils ressemblent aux
# listes( ils sont modifiables comme elles), mais ce ne sont pas des séquences.
# Les éléments que nous allons y enregistrer ne seront pas disposés dans un ordre immuable. En revanche, nous pourrons accéder à n’importe lequel d’entre eux à l’aide d’un index que l’on appellera une clé, laquelle pourra être alphabétique ou numérique.
#
#
# Comme dans une liste, les éléments mémorisés dans un dictionnaire peuvent être de n’importe quel type( valeurs numériques, chaînes, listes ou encore des dictionnaires, et même aussi des fonctions).
#
# Exemple :
#
# +
dico1={}
dico1['nom']='Paris-Orly Airport'
dico1['ville']='Paris'
dico1['pays']='France'
dico1['code']='ORY'
dico1['gps']=(48.7233333,2.3794444)
print(dico1)
print(dico1['code'])
print(dico1['gps'])
# -
# Remarques :
#
# - Des accolades délimitent un dictionnaire.
# - Les éléments d'un dictionnaire sont séparés par une virgule.
# - Chacun des éléments est une paire d'objets séparés par deux points : une clé et une valeur.
# - La valeur de la clé `'code'` est `'ORY'`.
# - La valeur de la clé `'gps'` est un tuple de deux flottants.
#
# ### Exercice 1 :
# Voici des données concernant l'aéroport international de Los Angeles :
#
# * __"Los Angeles International Airport","Los Angeles","United States","LAX",33.94250107,-118.4079971__
#
#
# En utilisant les mêmes clés que dans l'exemple précédent, créer le dictionnaire 'dico2' qui contiendra les données ci-dessus
# +
#Réponse
dico2={}
dico2['nom']=' '
dico2['ville']=' '
dico2['pays']=' '
dico2['code']=' '
dico2['gps']=(0,0)
print(dico2)
# -
# ### Méthodes
# +
#Afficher les clés
print(dico1.keys())
#Affichées les valeurs
print(dico1.values())
#Afficher les éléments
print(dico1.items())
# -
# ### Parcours
# On peut traiter par un parcours les éléments contenus dans un dictionnaire, mais attention :
# * Au cours de l'itération, __ce sont les clés__ qui servent d'index
# * L'ordre dans lequel le parcours s'effectue est imprévisible
#
# Exemple :
for element in dico1:
print(element)
# ### Exercice 2 :
# Modifier le programme ci-dessus pour obtenir l'affichage ci-dessous :
for element in dico1:
print(element)
# ## 4. Exercices
# ### Exercice 3 :
# Lors d'une élection, entre 2 et 6 candidats se présentent. Il y a 100 votants, chacun glisse un bulletin avec le nom d'un candidat dans l'urne. Les lignes de code ci-dessous simulent cette expérience.
# +
from random import *
#liste des candidats potentiels
noms=['Alice','Bob','Charlie','Daniella','Eva','Fred']
#liste des candidats réels
candidats=list(set([choice(noms) for i in range(randint(2,len(noms)))]))
#nombre de votants
votants=100
#liste des bulletins dans l'urne
urne=[choice(candidats) for i in range(votants)]
print('Candidats : ',candidats)
#print("Contenu de l'urne :", urne)
# -
# 1. Vérifier que les candidats réels changent à chaque éxecution de la cellule ci-desssus ainsi que le contenu de l'urne.
# 2. Compléter la fonction `depouillement(urne)`. Elle prend en paramètre une liste (la liste des bulletins exprimés) et renvoie un dctionnaire. Les paires clés-valeurs sont respectivement constituées du noms d'un candidat réels et du nombre de bulletins exprimés en sa faveur. Par exemple, si la liste des candidats est `['Alice','Charlie,'Bob']`, le résultat pourra s'afficher sous la forme `{'Eva': 317, 'Charlie': 363, 'Alice': 320}`
# +
def depouillement(urne):
decompte={}
for bulletin in urne:
if bulletin in decompte:
pass
else:
pass
return decompte
depouillement(urne)
# -
# 3. Ecrire la fonction `vainqueur(election)` qui prend en paramètre un dictionnaire contenant le décompte d'une urne renvoyé par la fonction précédente et qui renvoie le nom du vainqueur.
# +
def vainqueur(election):
vainqueur=''
nmax = 0
for nom in election:
pass
return vainqueur
vainqueur(depouillement(urne))
# -
# ### Exercice 4
# <img style='float:center;' src='https://openflights.org/demo/openflights-routedb-2048.png' width=500>
#
# Sur le site https://openflights.org/data.html , on trouve des bases de données mondiales aéronautiques.Le fichier `airports.txt` présent dans le dossier de cette feuille contient des informations sur les aéroports.
#
#
# Chaque ligne de ce fichier est formatée comme l'exemple ci-dessous :
#
# `1989,"Bora Bora Airport","Bora Bora","French Polynesia","BOB","NTTB",-16.444400787353516,-151.75100708007812,10,-10,"U","Pacific/Tahiti","airport","OurAirports"`
#
# On souhaite extraire les informations suivantes pour chaque aéroport:
# * Sa référence unique, un entier.
# * Le nom de l'aéroport, une chaîne de caractères
# * La ville principale qu'il dessert, une chaîne de caractères
# * Le pays de cette ville,une chaîne de caractères
# * Le code IATA de l'aéroport composé de 3 lettres en majuscules
# * Ses coordonées gps (latitude puis longitude), un tuple de 2 flottants.
# **1. Compléter les champs ci-dessous pour l'aéroport cité en exemple:**
# * ref : 1989
# * nom : Bora Bora Airport
# * ville : Bora Bora
# * pays : French Polynesia
# * code : BOB
# * gps : (-16.444400787353516,-151.75100708007812)
# **2. La fonction `data_extract` doit parcourir le fichier et extraire les données demandées qu'elle renvoie sous forme d'une liste de dictionnaires.**
#
# * Chaque élément de la liste est donc un dictionnaire qui correspond à un aéroport.
# * Les clés sont les noms des champs que l'on souhaite extraire et les valeurs sont celles associées à chaque aéroport.
#
# Recopier , modifier et compléter cette fonction pour qu'elle éxécute la tâche demandée :
# +
#2.
def data_extract(chemin):
fichier = open(chemin, "r",encoding='utf-8')
res = [] # Pour contenir le résultat qui sera une liste de dictionnaires
for ligne in fichier:
datas = ligne.strip().split(",") # une ligne du fichier
res.append(
{ "ref": int(datas[0]),
"nom": datas[1][1:-1],
"ville": "A compléter",
"pays": datas[3][1:-1],
"A compléter": datas[4][1:-1],
"gps" : "A compléter"
})
fichier.close()
return res
airports=data_extract('airports.txt')
#nombre d'aéroports référencés
print("A compléter")
#un aéroport au hasard
print(choice(airports))
# -
# **3. A l'aide d'une liste en compréhension, récupérer la liste des villes françaises desservies par un aéroport**
#3. Réponse
country='France'
res=[]
print(res)
# **4. Ecrire la fonction `infos(airports,ville)`.**
#
# Elle prend en paramètres la liste des dictionnaires de données extraites et une chaîne de caractères(le nom d'une ville). Elle renvoie les informations du ou des aéroports de cette ville dans une liste de dictionnaires:
# +
#4.
def infos(airports,ville):
print(infos(airports,'Nice'))
# -
# **5. Compléter les listes du code ci-dessous pour représenter les points de coordonnées gps de chacun des aéroports de la base de données(liste `X` des longitudes et liste `Y` des latitudes)**
# +
#5.
from matplotlib import pyplot #bibliothèque pour tracer des graphiques
X=[] #liste des longitudes
Y=[] #liste des latitudes
pyplot.figure(figsize = (14,7))
pyplot.xlim(-180, 180)
pyplot.ylim(-90, 90)
pyplot.plot(X,Y,'rx') # g=green, x est une croix
# -
# **6. Recopier et modifier le code précédent pour faire apparaître en rouge les aéroports situés en zone tropicale (c'est à dire dont la latitude est comprise entre $-23$ et $+23$)et en bleu les autres**
# +
#5.Réponse
| types_construits/dictionnaires/dictionnaires.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# (Image Credit: Dimensionless Blog)
#
# # Business Understanding
#
# Custom Churn refers to the number of customers who are likely to cancel the subscription after a given interval of time. It has lot of business value, for an organization ranging from providing services such as SaaS, Entertainment Platform, or Telecom services. The various techniques through which the customers can be kept in check and ensure that they do not withdraw from the platform can possibly be as follows:
#
# 1. Fresh customer acquirement
# 2. Upsell the existing customer base
# 3. Customer Retention
#
# These strategies play a key role in deciding the future customer base and profit of an organization, and also the special emphasis is on the Return of Investment (ROI) which is the ratio of the revenue generated from these activities to the cost of conducting them which should be substantial. Individual attention cannot be given to each customer, and even if provided the revenue generated would not exceed the cost and time given for retention. Thus, we are left with the option of identifying the set of customers which are likely to leave or cancel the subscription, and hence provide an optimal strategy to retain those in that particular set. This can be done by predicting the customer churn in advance before cancellation of the subscription by their respective users.
#
# A simple concept adopted by the Telecom companies involved churn prediction as a binary classification task, where the the output or target variable is simply having two values 'Yes' or 'No' whether a customer is likely to be retained or not in the user subscription. It involves a simple three step approach:
#
# 1. Gathering the customer data, the factors which would affect their subscription in a .csv file
# 2. Designing a predictor or making use of third-party prediction service where the data has to be uploaded for modelling
# 3. Use the model on each current customer to predict whether they are at risk of leaving.
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 5GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
df = pd.read_csv('/kaggle/input/churn-modelling/Churn_Modelling.csv')
# -
df.head()
df.info()
print("Shape of Data: ", df.shape)
df.describe()
# We will be following the **Crisp-DM** Approach in this project starting with the following.
#
# # Data Understanding
#
# There are 14 features in the given dataset:
# 1. RowNumber (Primary ID): Unique row figure assigned to each row of the data
# 2. CustomerId (Primary ID) and Surname: The unique identification number assigned to each customer, and 2932 different user surnames
# 3. CreditScore (numerical): The credit scores are assigned to customer based on their subscription and loyalty and are ranging from 350-850
# 4. Geography (categorical): The data is for only three countries: France, Spain and Germany
# 5. Gender (categorical): It is either 'Male' or 'Female'
# 6. Age (numerical): It is from 18 years old adults to 92 years old senior citizens
# 7. Tenure (numerical): The number of years the customer has been using the platform ranging from 0 to 10 years
# 8. Balance (numerical: float): The amount of money in the customer's account
# 9. NumOfProducts (numerical): The number of product subscriptions active for a user (ranging from 1 to 4)
# 10. HasCrCard (categorical): Whether the user has a credit card or not (1:'Yes' or 0:'No')
# 11. IsActiveMember(categorical): Whether the user is an active member or not (1:'Yes' or 0:'No')
# 12. EstimatedSalary (numerical): An estimate of the user salary in the future
# 13. Exited (categorical): Whether the customer will cancel or continue the subscription
#
# Clearly, our **target** variable is the Exited feature which is defining whether the user will continue the subscription or not
#
# - Task: Identify the users that will be exiting the platform (Exited: 0)
# - Input Features: RowNumber, CustomerId, CreditScore, Geography, Gender, Age, Tenure, Balance, NumOfProducts, HasCrCard, IsActiveMember, EstimatedSalary
# - Target Variable: Exited
#
# There is no missing values in the data, and all columns are filled, hence we do not have to handle missing values, and the features seem quite independent too.
# The features RowNumber, CustomerID do not add much to the prediction and can be either dropped or simply filtered for processing purposes
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.express as px
plt.figure(figsize=(15,15))
sns.heatmap(df.corr(), annot=True, cmap='viridis')
# # Data Preparation
#
# We can clearly see that there is no significant correlation between any of the features, and the highest correlation seems to between Age, and the target variable Exited which infers that there is a certain age group which is more likely to cancel the subscription than others, however this is also not highly correlated.
#
# - There is a negative correlation between the CreditScore and Exited, which means that with increasing credit score, the chances of cancelling the subscription decreases and vice versa.
# - Similarly, there is a negative correlation between the tenure and the cancellation meaning that larger tenures are less likely to cancel the subscription indicating loyalty to the services, and can be used effectively to identify a loyal customer base that can be upselled.
# - The correlation between the Number of Products and the subscription cancellation is also negative, meaning with higher number of services, obviously the customer is less likely to unsubscribe and will be present on the platform.
# - The negative correaltion between the Credit Card holders and cancellation is an unusual relationship to be studied.
# +
# Create Input Features, and Output Feature
X = df.iloc[:, 3:-1].values
y = df.iloc[:, -1].values
print("Input Features: ", X)
print("Target Variable: ", y)
# -
# We can observe that the highest number of customers are from France, while Germany and Spain have almost equal number of customers, but France has almost double than them
country = df.groupby(['Geography']).count()
country.RowNumber
sns.countplot(data=df, x='Geography')
# +
# The number of males is greater than the number of females
sns.countplot(data=df, x='Gender')
# Nearly 1000 more males are present in the subscription than females
genders = df.groupby(['Gender']).count()
print(genders)
# -
# So, we can say that the users are almost equitably distributed except for new users and very loyal users (10 years)
sns.countplot(data=df, x='Tenure')
# The median of the data is clearly 37 years of age and the upper-quartile limit is 62 years, rest are outliers
fig = px.box(df, y="Age")
fig.show()
# So, there are only 359 users greater than age of 62 using the platform
total_aged = (df["Age"]>62)
total_aged.value_counts()
# The user base below 32 years old is substantial
total_young = (df["Age"]<32)
total_young.value_counts()
# The estimated salaries are within the range of nearly 1000000 for majority of cases
plt.figure(figsize=(20,20))
sns.catplot(x="Geography", y="EstimatedSalary", hue="Gender", kind="box", data=df)
plt.title("Geography VS Estimated Salary")
plt.xlabel("Geography")
plt.ylabel("Estimated Salary")
# Most users in all the countries are within the age of 65 at max
fig = px.box(df, x="Age", y="Geography", notched=True)
fig.show()
fig = px.parallel_categories(df, dimensions=['HasCrCard', 'IsActiveMember'],
color_continuous_scale=px.colors.sequential.Inferno,
labels={'Gender':'Sex', 'HasCrCard':'Credit Card Holder', 'IsActiveMember':'Activity Status'})
fig.show()
fig = px.parallel_categories(df, dimensions=['HasCrCard', 'Gender','IsActiveMember'],
color_continuous_scale=px.colors.sequential.Inferno,
labels={'Gender':'Sex', 'HasCrCard':'Credit Card Holder', 'IsActiveMember':'Activity Status'})
fig.show()
fig = px.scatter_matrix(df,
dimensions=["Age"],
color="Exited")
fig.show()
# # A Few Insights and data preparation
#
# - The maximum number of customers are from the country of France, almost double that of the user base present in Germany and Spain.
# - Most of the users, are in age range of 31-65 with only 359 users above the age of 62 in all three countries.
# - The estimated salaries of females is higher than males in Germany and Spain, while in France, that of males is higher than that of females.
# - There are almost equal number of credit card holders who are active and inactive as well. Hence, we cannot say just by their credit card status whether they will remain active (also inferring the hypothesis that they may not have the capability to pay by credit card for subscription)
# - Studying the gender with the credit card history, we can point out that males and females are in equal number almost contributing to being active and inactive users, and hence does not get affected much with their credit card status.
# - There are no non-subscribers below the age of 36, and the maximum unsubscriptions take place in age-group of 51-55 and above 61.
# - Most of the features are independent from each other and hence should not be combined to form newer features.
# Label Encoding the "Gender" column
from sklearn.preprocessing import LabelEncoder
label_encoder = LabelEncoder()
X[:, 2] = label_encoder.fit_transform(X[:, 2])
print(X)
# One Hot Encoding the "Geography" column
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
coltrans = ColumnTransformer(transformers=[('encoder', OneHotEncoder(), [1])], remainder='passthrough')
X = np.array(coltrans.fit_transform(X))
print(X.shape)
# Standardize all the values in order to make them comparable
from sklearn.preprocessing import StandardScaler
stand_sc = StandardScaler()
X = stand_sc.fit_transform(X)
print(X)
# To balance the number of males, and females in the data along with countries to prevent overfitting
from imblearn.over_sampling import SMOTE
k = 1
seed=100
sm = SMOTE(sampling_strategy='auto', k_neighbors=k, random_state=seed)
X_res, y_res = sm.fit_resample(X, y)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_res, y_res, test_size = 0.3, random_state = 0)
# # Modelling and Evaluation
# +
from sklearn.metrics import precision_score, recall_score, f1_score, roc_auc_score, accuracy_score, classification_report, roc_curve
def evaluation(y_test, clf, X_test):
"""
Method to compute the following:
1. Classification Report
2. F1-score
3. AUC-ROC score
4. Accuracy
Parameters:
y_test: The target variable test set
grid_clf: Grid classifier selected
X_test: Input Feature Test Set
"""
y_pred = clf.predict(X_test)
print('CLASSIFICATION REPORT')
print(classification_report(y_test, y_pred))
print('AUC-ROC')
print(roc_auc_score(y_test, y_pred))
print('F1-Score')
print(f1_score(y_test, y_pred))
print('Accuracy')
print(accuracy_score(y_test, y_pred))
# looking at the importance of each feature
def feature_importance(model):
importances=model.feature_importances_
# visualize to see the feature importance
indices=np.argsort(importances)[::-1]
plt.figure(figsize=(20,10))
plt.bar(range(X.shape[1]), importances[indices])
plt.show()
def plot_loss(model):
prob=model.predict_proba(X_test)[:,1]
fpr, tpr, thresholds=roc_curve(y_test, prob)
plt.plot(fpr, tpr)
auc=roc_auc_score(y_test, prob)
print(auc)
# -
from sklearn.tree import DecisionTreeClassifier
dt_model = DecisionTreeClassifier()
dt_model.fit(X_train, y_train)
evaluation(y_test, dt_model, X_test)
feature_importance(dt_model)
plot_loss(dt_model)
from sklearn.ensemble import RandomForestClassifier
rf_model = RandomForestClassifier(n_estimators=100)
rf_model.fit(X_train, y_train)
evaluation(y_test, rf_model, X_test)
feature_importance(rf_model)
plot_loss(rf_model)
from sklearn.svm import SVC
svc_model=SVC(probability=True)
svc_model.fit(X_train, y_train)
evaluation(y_test, svc_model, X_test)
plot_loss(svc_model)
from sklearn.linear_model import LogisticRegression
lr_model = LogisticRegression()
lr_model.fit(X_train, y_train)
evaluation(y_test, lr_model, X_test)
plot_loss(lr_model)
from xgboost import XGBClassifier
xg_model = XGBClassifier()
xg_model.fit(X_train, y_train)
evaluation(y_test, xg_model, X_test)
feature_importance(xg_model)
plot_loss(xg_model)
from sklearn.ensemble import GradientBoostingClassifier
gb_model = GradientBoostingClassifier()
gb_model.fit(X_train, y_train)
evaluation(y_test, gb_model, X_test)
feature_importance(gb_model)
plot_loss(gb_model)
# +
# Let us try a small neural network to perform the required classification
import tensorflow as tf
# Initializing the ANN
nn = tf.keras.models.Sequential()
# Adding the input layer and the first hidden layer
nn.add(tf.keras.layers.Dense(units=10, activation='relu'))
# Adding the second hidden layer
nn.add(tf.keras.layers.Dense(units=6, activation='relu'))
# Adding the output layer
nn.add(tf.keras.layers.Dense(units=1, activation='sigmoid'))
# Part 3 - Training the ANN
# Compiling the ANN
nn.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# -
nn.fit(X_train, y_train, batch_size = 128, steps_per_epoch = 64, epochs = 50)
y_pred = nn.predict(X_test)
for i in range(0, y_pred.size):
if y_pred[i] > 0.5:
y_pred[i] = 1
else:
y_pred[i] = 0
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print(cm)
accuracy = round(accuracy_score(y_test, y_pred) * 100, 2)
print(accuracy)
import seaborn as sns
import matplotlib.pyplot as plt
plt.figure(figsize = (15, 15))
sns.heatmap(cm, annot = True, fmt = '.0f', linewidths = .1, square = True, cmap='viridis')
plt.xlabel('Prediction')
plt.title('Accuracy: {0}'.format(round(accuracy, 2)))
plt.ylabel('Actual')
plt.show()
# # Summary
#
# - Dataset has no missing values and has mostly independent features
# - The imbalance present in the data in form of majority males and lesser females, and France dominating over Spain and Germany was solved using Synthetic Minority Oversampling
# - Age is the best feature used for predicting the customer churn closely followed by other indpendent features in the data
# - The credit card status does not seem to be much of a contributing feature and hence, credit card offers may not be a relevant business strategy to reduce customer churn.
# - The majority of subscribers were in age bracket of [32, 65].
# - There were very few subscribers above the age of 65 combined in all three countries.
# - The estimated salary of females was higher than that of males in Spain and Germany, but did not showed a direct contribution to active subscriptions.
# - The performance of XG-Boost Classifier was best among its counterparts with F1-Score of 90.09
| Day 62/model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Retrieve phase from center of mass
# - uses stempy functions
# +
# %matplotlib widget
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import imageio
import stempy.io as stio
import stempy.image as stim
# -
def phase_from_com(com, theta=0, flip=False, reg=1e-10):
"""Integrate 4D-STEM centre of mass (DPC) measurements to calculate
object phase. Assumes a three dimensional array com, with the final
two dimensions corresponding to the image and the first dimension
of the array corresponding to the y and x centre of mass respectively.
Note this version of the reconstruction is not quantitative.
author: <NAME>
Parameters
----------
com : ndarray, 3D
The center of mass for each frame as a 3D array of size [2, M, N]
theta : float
The angle between real space and reciprocal space in radians
flip : bool
Whether to flip the com direction to account for a mirror across the vertical axis.
reg : float
A regularization parameter
Returns
-------
: ndarray, 2D
A 2D ndarray of the DPC phase.
"""
# Perform rotation and flipping if needed (from py4dstem)
CoMx = com[0,]
CoMy = com[1,]
if not flip:
CoMx_rot = CoMx*np.cos(theta) - CoMy*np.sin(theta)
CoMy_rot = CoMx*np.sin(theta) + CoMy*np.cos(theta)
if flip:
CoMx_rot = CoMx*np.cos(theta) + CoMy*np.sin(theta)
CoMy_rot = CoMx*np.sin(theta) - CoMy*np.cos(theta)
# Get shape of arrays
ny, nx = com.shape[1:]
# Calculate Fourier coordinates for array
ky, kx = [np.fft.fftfreq(x) for x in [ny,nx]]
# Calculate numerator and denominator expressions for solution of
# phase from centre of mass measurements
numerator = ky[:,None]*np.fft.fft2(CoMx_rot)+kx[None,:]*np.fft.fft2(CoMy_rot)
denominator = 2*np.pi*1j*((kx**2)[None,:]+(ky**2)[:,None])+reg
# Avoid a divide by zero for the origin of the Fourier coordinates
numerator[0,0] = 0
denominator[0,0] = 1
# Return real part of the inverse Fourier transform
return np.real(np.fft.ifft2(numerator/denominator))
# +
# Load a sparse 4D camera data set
# Close all previous windows to avoid too many windows
plt.close('all')
scan_num = 18
threshold = 4.5
dPath = Path('/mnt/hdd1/2021.03.02')
fPath = Path('data_scan{}_th{}_electrons.h5'.format(scan_num, threshold))
fname = dPath / fPath
electron_events = stio.load_electron_counts(str(fname))
print('File: {}'.format(fname))
print('Initial scan dimensions = {}'.format(electron_events.scan_dimensions))
# +
# Calculate a summed diffraction pattern of frames
# And find the center
dp = stim.calculate_sum_sparse(electron_events.data[::10],
electron_events.frame_dimensions)
# Set the center of the pattern (use figure below for manual)
center = stim.com_dense(dp)
#center = (248, 284)
print(center)
fg,ax = plt.subplots(1, 1)
ax.imshow(np.log(dp+1))
ax.scatter(center[0], center[1], c='r')
_ = ax.legend(['center of pattern'])
# +
# Calculate a virtual bright field and dark field
outer_angle = 30 # in pixels
ims = stim.create_stem_images(electron_events, (0, 180), (50, 280), center=center) # here center is (col, row)
bf = ims[0,]
adf = ims[1,]
fg,ax = plt.subplots(1, 2, sharex=True, sharey=True)
ax[0].imshow(bf)
ax[0].set(title='vBF')
ax[1].imshow(adf)
ax[1].set(title='vADF')
# +
# Calculate the center of mass of every frame
com = stim.com_sparse(electron_events.data, electron_events.frame_dimensions)
# This can be removed in the future
com = com.reshape((2, electron_events.scan_dimensions[1], electron_events.scan_dimensions[0]))
fg,ax = plt.subplots(1,2,sharex=True,sharey=True)
axim0 = ax[0].imshow(com[0,], cmap='bwr',vmin=com[0,10:-10,:].min(),vmax=com[0,10:-10,].max())
axim1 = ax[1].imshow(com[1,], cmap='bwr',vmin=com[1,10:-10,:].min(),vmax=com[1,10:-10,].max())
# +
# Calculate the radius and angle for each COM measurement
com_mean = np.mean(com,axis=(1,2))
com_r = np.sqrt( (com[0,] - com_mean[0])**2 + (com[1,] - com_mean[1])**2 )
com_theta = np.arctan2((com[1,] - com_mean[1]), (com[0,] - com_mean[0]))
fg,ax = plt.subplots(1, 2,sharex=True,sharey=True)
ax[0].imshow(com_r,cmap='magma',vmin=com_r[10:-10,:].min(),vmax=com_r[10:-10,].max())
ax[1].imshow(com_theta, cmap='twilight')
# +
# Retrieve phase from center of mass
# 300kV: flip=True and theta=0 + STEM scan rotation
# 80 kV: flip=True and theta=35 works well.
flip = True
theta = 0 * np.pi / 180. # rotation between diffraction and real space scan directions
# Calculate the phase
ph = phase_from_com(com, flip=flip, theta=theta, reg=1e-1)
fg,ax = plt.subplots(1,2,sharex=True,sharey=True)
#ax[0].imshow(ph, vmin=ph[10:-10,10:-10].min(), vmax=ph[10:-10,10:-10].max())
ax[0].imshow(ph / np.std(ph), vmin=-2, vmax=2)
ax[0].set(title = 'DPC')
ax[1].imshow(adf)
ax[1].set(title = 'vADF')
fg,ax = plt.subplots(1,2,sharex=True,sharey=True)
ax[0].imshow(np.log(np.abs(np.fft.fftshift(np.fft.fft2(ph)))),vmin=1e-3)
ax[1].imshow(np.log(np.abs(np.fft.fftshift(np.fft.fft2(bf)))))
# -
fg,ax = plt.subplots(1, 3, sharex=True, sharey=True,figsize=(12,5))
ax[0].imshow(bf)
ax[0].set(title='BF')
ax[1].imshow(adf)
ax[1].set(title='ADF')
ax[2].imshow(ph,vmin=ph[10:-10,].min(),vmax=ph[10:-10,].max())
ax[2].set(title = 'DPC')
# Save the data
print('Saving COM and DPC for scan number {}'.format(scan_num))
imageio.imwrite(fname.with_name('scan{}_DPC'.format(scan_num)).with_suffix('.tif'), ph.astype(np.float32))
imageio.imwrite(fname.with_name('scan{}_comx'.format(scan_num)).with_suffix('.tif'), com[0,].astype(np.float32))
imageio.imwrite(fname.with_name('scan{}_comy'.format(scan_num)).with_suffix('.tif'), com[1,].astype(np.float32))
imageio.imwrite(fname.with_name('scan{}_BF'.format(scan_num)).with_suffix('.tif'), bf.astype(np.float32))
imageio.imwrite(fname.with_name('scan{}_ADF'.format(scan_num)).with_suffix('.tif'), adf.astype(np.float32))
| examples/center_of_mass_sparse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="j7CI8T-Oxv3a"
# ### Dependencies
# + colab_type="code" id="1RUl59iOxtSM" outputId="1ef1d907-e2de-4ba2-ebda-a2ad04cb97a9" colab={"base_uri": "https://localhost:8080/", "height": 121}
# !git clone "https://github.com/suyash/ContextualDecomposition.git" && mv ContextualDecomposition/cd ./cd
# + colab_type="code" id="5Q_p9EfZxQpx" outputId="314577d9-3ae3-4666-fb20-cf0023bc8bb7" colab={"base_uri": "https://localhost:8080/", "height": 86}
# !curl -L -o "job_dir.zip" "https://drive.google.com/uc?export=download&id=1wkygiEOc2T9LFbD4fcO__9tXGVkxu0qu" && unzip -q -d "job_dir" "job_dir.zip"
# + [markdown] colab_type="text" id="8eZJUKK70evz"
# ### Imports
# + colab_type="code" id="ADDYjJZq0rtZ" outputId="e67a4834-e2ce-4af0-c56c-34b24bf4fa13" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorflow_version 2.x
# + colab_type="code" id="wGhV7mrdxBpY" colab={}
import numpy as np
import tensorflow as tf
from tensorflow.keras import Model # pylint: disable=import-error
from cd.cd import cnn_net_decomposition
from cd.preprocess import create_table, create_inv_table
# + colab_type="code" id="aBQ73NDPxBpi" outputId="a39963a1-38b6-48a0-8935-8532c2add168" colab={"base_uri": "https://localhost:8080/", "height": 764}
model = tf.keras.models.load_model("job_dir/saved_model/best")
model.summary()
# + colab_type="code" id="eckB_JwkxBps" colab={}
tokens = np.load("job_dir/tokens.npy")
table = create_table(tokens)
inv_table = create_inv_table(tokens)
# + [markdown] colab_type="text" id="iVFs3an0xBpy"
# ### Process Input
# + id="WMw-sO_wKKmq" colab_type="code" colab={}
s = "the longer the movie goes , the worse it gets , but it 's actually pretty good in the first few minutes"
# + colab_type="code" id="NGgdMoEUxBp7" outputId="f0e81a2d-3537-4bb1-abe7-0318f37f0b21" colab={"base_uri": "https://localhost:8080/", "height": 86}
inp = table.lookup(tf.constant(s.split()))
inp = tf.expand_dims(inp, 0)
inp
# + [markdown] colab_type="text" id="0WX57WJYxBqC"
# ### Generate Overall Prediction
# + colab_type="code" id="dYcUo5ntxBqE" outputId="b5b65ec3-c322-4be2-f14f-ed0b9c9d0b7e" colab={"base_uri": "https://localhost:8080/", "height": 34}
x = model.predict(inp)
x = tf.math.softmax(x)
x
# + [markdown] colab_type="text" id="EuK66fAxxBqI"
# $P(neg) = 0.98$, $P(pos) = 0.01$
#
# Now, decomposing and getting predictions for subsections
# + colab_type="code" id="N1gvo4A1xBqJ" outputId="3bc7e4b1-410a-4a40-cf63-232ca23ebdd2" colab={"base_uri": "https://localhost:8080/", "height": 399}
t = inv_table.lookup(inp[0]).numpy()
list(enumerate(t))
# + [markdown] colab_type="text" id="x97SfTKlxBqe"
# ### Decomposing the prediction into the prediction for [0..15] and [16..26]
# + colab_type="code" id="6c2JiDkAxBqg" colab={}
weights = model.weights
# + colab_type="code" id="UxtXQoMtxBqm" colab={}
embed_inp = tf.nn.embedding_lookup(params=weights[0], ids=inp)
# + id="-aGT_ORkKKnL" colab_type="code" colab={}
conv_weights = []
for i in range((len(weights) - 3) // 2):
conv_weights.append([weights[2 * i + 1], weights[2 * i + 2]])
# + colab_type="code" id="NG-I_V1dxBqq" colab={}
pred_0_11, _ = cnn_net_decomposition(embed_inp, conv_weights, 0, 11)
pred_12_21, _ = cnn_net_decomposition(embed_inp, conv_weights, 12, 21)
# + id="d1Q-3LcwKKnR" colab_type="code" colab={}
dw, db = weights[-2], weights[-1]
# + colab_type="code" id="AAOiY_AaxBqu" outputId="02fe38b3-a910-4c3a-fe10-d4befbb3b1d0" colab={"base_uri": "https://localhost:8080/", "height": 34}
tf.math.softmax(tf.matmul(pred_0_11, dw) + db)
# + colab_type="code" id="Bd64gk9TxBqz" outputId="d0be51f5-bd3c-4a06-a461-f94efea25aa1" colab={"base_uri": "https://localhost:8080/", "height": 34}
tf.math.softmax(tf.matmul(pred_12_21, dw) + db)
# + [markdown] colab_type="text" id="AN9S5whwxBq4"
# decomposed prediction for __"the longer the movie goes , the worse it gets , but"__: $P(neg) = 0.999, P(pos) = 0.0001$
#
# decomposed prediction for __"it 's actually pretty good in the first few minutes"__: $P(neg) = 0.0008, P(pos) = 0.991$
# + [markdown] colab_type="text" id="Q9yTEzkh8Gcs"
# ### Individual Word Level Decomposition
# + colab_type="code" id="7pPBCCbZ8KGj" colab={}
preds = []
for i in range(22):
rel, _ = cnn_net_decomposition(embed_inp, conv_weights, i, i)
pred = tf.math.softmax(tf.matmul(rel, dw) + db)
preds.append(pred.numpy().tolist()[0])
# + colab_type="code" id="T9VmfVvk8lmi" outputId="d99ec7d2-6379-459c-d8d9-01e607dd649b" colab={"base_uri": "https://localhost:8080/", "height": 399}
list(zip(t, preds))
| conv1d_decomposition_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="1bWBoBl-_b3z"
# Importing necessary libs
# + id="tYEoncVA_0vu"
import pandas as pd
import tensorflow
import numpy
import sklearn
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import LabelEncoder, StandardScaler
# + [markdown] id="TUv5W0ze_kiY"
# I ran this code on Google Colab with mounted Drive so specify path of dataset if running on local. If using Kaggle specify kaggle library.
# + [markdown] id="eUQKODgz_0Pn"
# This Dataset is of a server log for week1 and classifies sessions as suspicious or not. We are trying to see what factors are responsible for malicious activities. I have done some basic preprocessing steps followed by EDA.
# + id="gnVymflIDG6g"
df = pd.read_csv("../input/server-logs-suspicious/CIDDS-001-external-week1.csv")
# + id="__nENwX0HKML" outputId="0a4b7023-c785-405d-a104-4bae6992e25b"
df.head()
# + id="FIjE7iTQHoZX" outputId="e24980bc-7ccf-4b4d-9a95-02f32cc9061d"
df.columns
# + [markdown] id="8ZO3E5JGAXzh"
# Checking for NULL values below. Some classification models on sklearn do not work well with missing values therefore it is necessary to remove or impute them with another well fitted statistical measure
# + id="EIo-vV7sHwWq" outputId="a26b7e21-113b-4f05-9189-901636fd7306"
df.isnull().sum()
# + id="nshdgCxTHzBD" outputId="a5d5e59e-42e2-4d0e-d91e-27e6fb9edf9f"
df.dtypes
# + [markdown] id="yGQElITKAuJP"
# We have the class variable which we need to predict later on. It has following unique values.
# + id="kuSHBt2BH2Ba" outputId="ebcdb332-ec08-424b-b5f6-b49f6fd9f942"
df["class"].unique()
# + id="67dlfclVH6EM" outputId="56e3b9c7-8052-4a08-9143-d7f2e70c93e9"
data = df.copy()
data.shape
# + [markdown] id="YBaTNLsPA47X"
# We are using seaborn plots to visualize our data
# + [markdown] id="JpNUKft9BKoS"
# We can see that majority of the data is suspicious sessions so our model will learn better on classifiying suspicious class.
# + id="T0DnbWx7IYrz" outputId="de7ec9ee-0d7d-44c7-eae7-d5496f2a5186"
sns.set(style = "darkgrid")
sns.countplot(x = "class",data=data)
# + [markdown] id="NSFZgXwSBcWA"
# The Flags field is a string of some flag variables which are necessary in the networking field. Wherever a certain flag is not set a "." is used. We need to break this columns into individual variables for flags as it will help the model learn faster and establish better relations.
# + id="TMRAnN5lIifc" outputId="0be9d8b1-987a-4978-8cb9-465e888777ad"
df["Flags"].unique()
# + id="SW3q73xHJR-a"
df["A"]=0
df["P"]=0
df["S"]=0
df["R"]=0
df["F"]=0
df["x"]=0
# + id="yd1nVyb_JVc4"
def set_flag(data,check):
val=0;
if(check in list(data["Flags"])):
val = 1 ;
return val;
# + id="1RqUebMuJYB3" outputId="799c4ecf-f021-41d1-a865-050caec0ffdc"
df.columns
# + id="iTEuJv8MJZ16"
df["A"] = df.apply(set_flag,check ="A", axis = 1)
df["P"] = df.apply(set_flag,check = "P" ,axis = 1)
df["S"] = df.apply(set_flag,check ="S",axis = 1)
df["R"] = df.apply(set_flag,check="R" ,axis = 1)
df["F"] = df.apply(set_flag,check ="F" ,axis = 1)
df["x"] = df.apply(set_flag,check ="x" ,axis = 1)
# + [markdown] id="F64xlKicB_co"
# Checking here the individual flag variables and impact of each variable on class. You can change the variable name in the below plot and see the impact.
# + id="0edOQIrpJnN_" outputId="2a56af77-f2dc-427c-922c-32fa102f15ce"
sns.countplot(x="S",hue = "class",data=df)
# + id="02rsOpPKKG7z" outputId="8a6b23b7-eaeb-44e4-e8a6-215583205b64"
sns.countplot(x = "Proto",hue = "class",data = df)
# + [markdown] id="c7uq92T2CW-h"
# Dropping some unnecessary columns and columns having a single value like flows and tos
# + id="KtRpjWSYKIqP"
df=df.drop(columns = ["Date first seen","attackType","attackID","attackDescription","Flows","Tos","Flags"])
# + id="YEttqYvSKNX_" outputId="3e7ec970-f7cb-47a9-84c1-a80a51d8d86b"
df.head()
# + [markdown] id="9bHOQ0lfCh7e"
# The Bytes variable was an object as seen in head command and the model would not recognize it as a number. Therefore we convert the M in the number to a multiplication of 1M with the number part. This has been simply done using regex
# + id="A-HrbEZKKPE2"
import re
def convtonum(data):
num1=data["Bytes"]
if "M" in data["Bytes"]:
num=re.findall("[0-9.0-9]",data["Bytes"])
num1 = float("".join(num))*100000
num1 = float(num1)
return num1
# + id="ybYgbSPeK2bo"
df["Bytes"] = df.apply(convtonum,axis = 1)
# + id="OZvR7eLnK4hq" outputId="c8d8ca8c-4250-4dc4-c1d0-01f497f23ab7"
df.head()
# + [markdown] id="BJRjT5xCC1Sv"
# Label Encoding categorical values.
# + id="DlabnMXVK75q"
from sklearn.preprocessing import LabelEncoder
col = ["Proto","class","Src IP Addr","Dst IP Addr"]
enc = LabelEncoder()
for col_name in col:
df[col_name]=enc.fit_transform(df[col_name])
# + [markdown] id="QVCOsfemDBfI"
# Correlation Heatmap shows how each variable is correlated with class variable which we will try to predict.
# + id="QgO9oJHbK_RT" outputId="cbae4600-9493-46d2-ca7c-e7c52f1f1e0a"
data1 = df.copy()
plt.figure(figsize=(18,5))
sns.heatmap(data1.corr(),annot=True,cmap = "RdYlGn")
# + id="hjuIMn6JLBof"
data_y = data1["class"]
data_x = data1.drop(columns = ["class"])
# + [markdown] id="eECO8UZBDYMn"
# Breaking dataset into train and test sets randomly
# + id="RU98hlRcLGJl"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data_x, data_y, test_size=0.3, random_state=1)
# + id="PwNTNcQGLINv"
#decision-tree-classifier - single-tree-classifier // using all features
from sklearn.tree import DecisionTreeClassifier
clf = DecisionTreeClassifier(criterion="entropy", max_depth=10) # you can use GINI index also here as a critirion
clf = clf.fit(X_train,y_train)
y_pred = clf.predict(X_test)
# + [markdown] id="WqPWIcGIDtMS"
# We got a 99% accuracy on the first go we can check further if it is overfitting and also see important variable in the model
# + id="l6FDaQnKLc4d" outputId="13b85bcc-b1be-4e69-be0a-91d569d699f0"
from sklearn import metrics
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# + id="Jxndbn8nLoct" outputId="33bda12c-58bf-4321-f8a9-5172654308fa"
feat_importances = pd.Series(clf.feature_importances_, index=X_train.columns)
feat_importances.nlargest(15).plot(kind='barh')
plt.show()
# + id="4AJfj5cpX_YU" outputId="b1f85a67-5d2d-4ff5-b969-24eda5ba9660"
# if you want to select most important features from an algorithm use recursive feature elimination and run algorithm on that
from sklearn.feature_selection import RFE
m = DecisionTreeClassifier(criterion="entropy", max_depth=10)
rfe = RFE(m,8)
fit=rfe.fit(X_train,y_train)
print(X_train.columns)
print("Num Features: %d" % fit.n_features_)
print("Selected Features: %s" % fit.support_)
print("Feature Ranking: %s" % fit.ranking_)
# -
# Please upvote the kernel if it was helpful
# + id="7IAWPxe3asYI"
# + id="R5YVsoPktkX6"
| 02.EDA/01.eda-and-classification-for-beginners.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # 白盒攻击
# +
import os
import torch
from torch import nn, optim
from tqdm import tqdm
import utils
model_dict_path = "./white_model_1.pt"
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(device)
# -
# ## 1. 模型定义
net = utils.white_model()
net.to(device)
# ## 2. 加载数据
batch_size = 256
train_iter, test_iter = utils.load_data_fashion_mnist(batch_size=batch_size)
# ## 3. 训练或者加载模型
if os.path.exists(model_dict_path):
print("加载之前训练好的模型...", end="")
net.load_state_dict(torch.load(model_dict_path))
print("完成.")
print("test acc: ", utils.evaluate_accuracy(test_iter, net))
else:
print("训练模型...")
lr, num_epochs = 0.001, 10
optimizer = torch.optim.Adam(net.parameters(), lr=lr)
utils.train(net, train_iter, test_iter, batch_size, optimizer, device, num_epochs)
print("保存模型...", end="")
torch.save(net.state_dict(), model_dict_path)
print("完成.")
# ## 4. 选取1000个成功分类的样本
# +
net.eval()
X_test, y_test = utils.select_right_sample(net, test_iter)
assert (net(X_test.to(device)).argmax(dim=1) == y_test.to(device)).float().sum().cpu().item() == 1000.0
# -
# ## 5. 攻击
attack_lr, max_attack_step = 0.01, 50
before_atk_X, before_atk_y = [], []
after_atk_X, after_atk_y = [], []
for i in tqdm(range(1000)):
X_, y_, success = utils.white_box_attack(net, X_test[i:i+1], y_test[i:i+1], attack_lr, max_attack_step)
if success:
before_atk_X.append(X_test[i])
before_atk_y.append(y_test[i])
after_atk_X.append(X_)
after_atk_y.append(y_)
print("学习率%.3f,最大步数%d的白盒攻击成功率: %.2f%%" % (attack_lr, max_attack_step, len(after_atk_y) / 10))
print("成功样本举例:\n")
print("攻击前:")
utils.show_fashion_mnist(before_atk_X[:10], utils.get_fashion_mnist_labels(before_atk_y[:10]))
print("攻击后:")
utils.show_fashion_mnist(after_atk_X[:10], utils.get_fashion_mnist_labels(after_atk_y[:10]))
| white_box_attack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Understanding Tash's USMLE workload
# # Imports
# #### Standard libraries
# +
import os
import json
import pickle
import datetime
# -
# #### Third party imports
from moviepy.editor import VideoFileClip
# #### Local aplication imports
# # Initial exploration
# + [markdown] heading_collapsed="true" tags=[]
# ## Exploring video
# -
pth = "../../../../../Desktop/USMLE STEP 1/Kaplan USMLE Step 1 Videos 2020/01 Anatomy 34h27m/01 Embryology & Histology/"
vx = [vid for vid in os.listdir(pth) if "06" in vid][0]
vx = VideoFileClip(pth + vx)
vx_dur = vx.duration
vx_dur/60
# + [markdown] tags=[]
# ## Working with one dir
# -
pth = "../../../../../Desktop/USMLE STEP 1/Kaplan USMLE Step 1 Videos 2020/01 Anatomy 34h27m/02 Gross Anatomy/"
vx = [vid for vid in os.listdir(pth)]
# + tags=[]
dirx = {}
for vid in os.listdir(pth):
dirx[vid] = round(VideoFileClip(pth + vid).duration/60, 2)
# +
vsum = 0
for vid in dirx:
vsum += dirx[vid]
vsum/60
# + [markdown] tags=[]
# # Calculating work plan
# + [markdown] tags=[]
# ## How much time for all USMLE?
# + [markdown] heading_collapsed="true" tags=[]
# ### Calculating total time for each video with loop
# -
pth = "../../../../../Desktop/USMLE STEP 1/Kaplan USMLE Step 1 Videos 2020/"
main_topic_list = [mtl for mtl in os.listdir(pth) if "." not in mtl]
# + tags=[] active=""
# comp_dir = {}
# tot_t = 0
# for mtl in main_topic_list:
#
# comp_dir[mtl] = {}
#
#
# for stl in os.listdir(pth + mtl):
#
# comp_dir[mtl][stl] = {}
#
#
# for vid in [v for v in os.listdir(pth + mtl + "/" + stl) if ".ini" not in v]:
#
# comp_dir[mtl][stl][vid] = round(VideoFileClip(pth + mtl + "/" + stl + "/" + vid).duration/60, 2)
# tot_t += comp_dir[mtl][stl][vid]
#
# + active=""
# pickle.dump(comp_dir, open("time_dict.pkl", "wb"))
# + [markdown] heading_collapsed="true" tags=[]
# ### Importing results form pickle
# -
comp_dir = pickle.load(open("time_dict.pkl", "rb"))
# + [markdown] heading_collapsed="true" tags=[]
# ### Calculating total time in minutes for all videos
# -
tt = 0
for mtl in comp_dir:
for stl in comp_dir[mtl]:
for vid in comp_dir[mtl][stl]:
tt += comp_dir[mtl][stl][vid]
print("Total vide time in minutes: ", tt)
print("Total vide time in hours: ", tt/60)
print("Total vide time in days: ", tt/(60*24))
# + [markdown] heading_collapsed="true" tags=[]
# ### How many hours from today until goal day?
# -
time_i = datetime.date(2021, 5, 7)
time_f = datetime.date(2021, 9, 1)
days = time_f - time_i
days = days.days
days
(tt/60)/days
# + [markdown] tags=[]
# ## Table with organized information
# -
for lv1 in comp_dir["07 Physiology 34h33m"]:
print(lv1)
# + [markdown] tags=[]
# ---
# ---
| notebooks/tash_usmle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading data from Excel
#
# Let's get some data. Download [Sample Superstore Sales .xls file](https://community.tableau.com/docs/DOC-1236) or [my local copy](https://github.com/parrt/msan692/raw/master/data/SampleSuperstoreSales.xls) and open it in Excel to see what it looks like.
#
# Data of interest that we want to process in Python often comes in the form of an Excel spreadsheet, but the data is in a special non-text-based (binary) format that we can't read directly:
with open('data/SampleSuperstoreSales.xls', "rb") as f:
txt = f.read()
print(txt[0:100])
# ## Converting Excel files with csvkit
#
# There's a really useful tool kit called `csvkit`, which you can install with:
# ! pip install -q -U csvkit
# Then, the following command works without having to run or even own Excel on your laptop (but you might get lots of warnings):
# ! in2csv data/SampleSuperstoreSales.xls > /tmp/t.csv
# ! head -3 /tmp/t.csv # show 3 lines of t.csv
# You might be wondering why we would need a commandline version that converts Excel files to CSV. Certainly, as we see in the next section, we can use pandas. The difference is that using the command line means we don't have to write software, which is much more involved than simply executing a few lines on the commandline. For example, we could easily use a shell `for` loop to convert multiple Excel files to csv without starting a Python development environment.
# ## Reading Excel files with Pandas
#
# The easiest way to read Excel files with Python is to use Pandas:
import pandas as pd
table = pd.read_excel("data/SampleSuperstoreSales.xls") # XLS NOT CSV!
table.head(3)
# # Reading CSV data using built-in csv package
#
# Grab the CSV version of the Excel file [SampleSuperstoreSales.csv](https://github.com/parrt/msan692/raw/master/data/SampleSuperstoreSales.csv) we've been playing with. You already know how to load CSV with pandas, but sometimes it's reasonable to process the CSV at a lower level, without getting a big data frame back. For example, we might want to simply process the CSV file line by line if it is truly huge.
#
# In your project, you dealt with very simple CSV, but it can get much more complicated so let's take a quick look how to use the `csv` package. CSV files can get complicated when we need to embed a comma. One such case from the above file shows how fields with commas get quoted:
#
# ```
# "Eldon Base for stackable storage shelf, platinum"
# ```
#
# What happens when we want to encode a quote? Well, somehow people decided that `""` double quotes was the answer (not!) and we get fields encoded like this:
#
# ```
# "1.7 Cubic Foot Compact ""Cube"" Office Refrigerators"
# ```
#
# The good news is that Python's `csv` package knows how to read Excel-generated files that use such encoding. Here's a sample script that reads such a file into a list of lists:
# +
import sys
import csv
table_file = "data/SampleSuperstoreSales.csv"
with open(table_file, "r") as csvfile:
f = csv.reader(csvfile, dialect='excel')
data = []
for row in f:
data.append(row)
print(data[:6])
# -
# Or add to a numpy `array`:
import numpy as np
np.array(data)
# ## Reading CSV into Pandas Data frames
#
# *This should be review for you...*
#
# In the end, the easiest way to deal with loading CSV files is probably with [Pandas](http://pandas.pydata.org/), even if you need the data in a different data structure. For example, to load our sales CSV, we don't even have to manually open and close a file:
import pandas
df = pandas.read_csv("data/SampleSuperstoreSales.csv")
df.head(3)
# Pandas hides all of the details. I also find that pulling out columns is nice with pandas. Here's how to print the customer name column:
df['Customer Name'].head()
df.Profit.head()
# ### Exercise
#
# From the sales CSV file, use pandas to read in the data and multiply the `Order Quantity` and `Unit Price` columns to get a new column.
df = pandas.read_csv("data/SampleSuperstoreSales.csv")
(df['Order Quantity']*df['Unit Price']).head()
| notes/excel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: openmc-env
# language: python
# name: openmc-env
# ---
# # OpenMC introduction
#
# Please indicate your name below, since you will need to submit this notebook completed latest the day after the datalab.
#
# Don't forget to save your progress during the datalab to avoid any loss due to crashes.
name=''
# In our previous endeavours we experienced that performing an accurate simulation of particle transport is not trivial. There are several softwares developed to solve this task, and for someone who is more interested in performing the analysis of a reactor core, it is often better to use an already verified and validated code instead of developing a new one from scratch. There are several such codes solving neutron transport either relying on deterministic or stochastic methods. During this datalab we will use a rather young code called OpenMC, which was originally developed at MIT.
#
# OpenMC is a Monte Carlo neutron and photon transport simulation code. It can be used both for fixed source and for criticality calculations. The user can define complicated geometries with constructive solid geometries. OpenMC is written in C++, however it has a rich Python API, which will allow us to comminicate with the code through jupyter notebooks.
#
# OpenMC is not yet an industrial standard code, however our motivation to use it for this course was that it is
#
# 1. freely available
# 2. the important physics we could not tackle within this course ourselves (eg. thermal scattering and several reactions) are implemented in it
# 3. since we can interact with the code through Python, we can deepen our programming skills
# 4. for small problems it runs relatively fast on a single PC
#
# That said, today we are going to walk through a simple openMC input file. During this course we will only work with pincell models (ie. a single fuel pin infinite in the axial length with reflective boundaries, so neutrons are reflected back from the boundary in this way approximating an infinite core). This is of course a simplification of a complete reactor core, but it will still allow us to see a lot of cool physics. One could of course perform simulations at the assembly level or for a full-core, however such simulations would be more time consuming. There are several further examples at https://docs.openmc.org/.
#
# **Note that this notebook should be opened from an environment where openmc is available!**
#
# ## The input file
#
# During the datalab we will define the following problem which roughly matches a typical western-type pressurized water reactor's conditions:
#
# - PWR pincell with reflective boundaries
# - Fuel radius: 0.41 cm
# - Fuel material: enriched UO2 with density 10.5 g/cm3
# - Cladding inner/outer radius: 0.42/0.45 cm
# - Cladding material: zirconium with density 6.6 g/cm3
# - Coolant/moderator: pressurized water with density 0.75 g/cm3
# - Cell pitch: 1.26 cm
#
# Note that in practice, PWR fuel cladding is made of an alloy called Zircaloy, nevertheless it has little influence on the neutron economy, therefore we will approximate it as pure zirconium with its natural abundance of isotopes.
#
# So what do we need in order to define an openMC input?
#
# 1. Materials: which materials are included in our geometry (nuclide content, temperature, density)?
# 2. Geometry: how does the geometry look like?
# 3. Tally: what are the quantities of interest? For example flux, or fission rate?
# 4. Settings: how many neutrons we would like to include in the calculation?
#
# For each of these steps we will use the python API to export an xml file, which the code will use once we execute it. Note however that one could directly write xml files without using python, and run the code outside of Jupyter. For larger problems this is sometimes better. However we will want to analyse the output results in python, and sometimes we will script the input, so for us it is handy to use the python API.
#
# Let's get started!
#
# ### Import
#
# We will need first to import openmc, and of course anything else we might need.
import openmc
import numpy as np
import matplotlib.pyplot as plt
import os
# ### Materials
#
# Do you remember the example `Material()` class we developed before? That is pretty similar to the `Material()` class of openMC. We create an instance of `Material()`, with a numeric id of the material, and then we can add nuclides, and set the density. We can also set the temperature (which can be 294 K, 600 K, 900 K, 1200 K, 1500 K, the temperatures at which the cross sections are evaluated). There are other options as well (for example one can directly set the enrichment), what you can find in the documentation.
#
# If the nuclide inventory is not relevant for a material, we can add elements, in which case openMC will assume the natural abundance of isotopes.
#
# Notice that for water we will need to link to the $S(\alpha,\beta)$ laws, since we need to take into account the molecular bounds for scattering on hydrogen.
#
# Finally, we create an instance of `Materials()` (note, it is plural now!), and link the materials of the problem to this object. Then with the `export_to_xml()` method we export the information into an xml file.
# +
uo2 = openmc.Material(1, "uo2",temperature=1200)
# Add nuclides to uo2
uo2.add_nuclide('U235', 0.04)
uo2.add_nuclide('U238', 0.96)
uo2.add_nuclide('O16', 2.0)
uo2.set_density('g/cm3', 10.5)
#cladding
zirconium = openmc.Material(2, "zirconium",temperature=900)
zirconium.add_element('Zr', 1.0)
zirconium.set_density('g/cm3', 6.6)
#coolant
water = openmc.Material(3, "h2o")
water.add_nuclide('H1', 2.0)
water.add_nuclide('O16', 1.0)
water.set_density('g/cm3', 0.75)
water.add_s_alpha_beta('c_H_in_H2O')
#creating Materials() and exporting to xml
mats = openmc.Materials([uo2, zirconium, water])
mats.export_to_xml()
# -
# We can print the content of the created xml file, with the linux command `cat`. (The `!` sign let's the python interpreter know that this is a command of the operating system.
#
# You can see how all the information was structured in an xml (which is probably familiar from the previous datalab).
# !cat materials.xml
# ### Geometry
#
# Defining the geometry is only marginally more complicated. We will need to define surfaces (eg. `ZCylinder()`, `XPlane()`, and `YPlane()`), and regions bounded by the surfaces. Then we link the regions and the filling material to `Cell()` objects, which all have a numeric ID.
#
# Our geometry is infinite in the axial dimension. First we define three cylinders (the bounding surfaces of the fuel, and the cladding). Then we use constuctive solid geometry rules to define the regions between the surfaces, and create the fuel pin. Finally we will define the water region around the fuel.
#
# We will then need to define a `Universe()` (the usefulness of this is not apparent in our simple exercise, but when defining reactor cores with repetitive patterns of various types of fuel assemblies, then "universes" are the way to handle this in most neutron transport code. You do not need to worry about this in this course). And finally we create a `Geometry()`, and link the universe. We will then export the information into an xml file.
#
# We further commented the lines to explain them
# +
fuel_or = openmc.ZCylinder(r=0.41) #fuel cylinder with outer radius
clad_ir = openmc.ZCylinder(r=0.42) #clad inner cylinder with inner radius
clad_or = openmc.ZCylinder(r=0.45) #clad outer cylinder with outer radius
fuel_region = -fuel_or #inside the fuel cylinder
gap_region = +fuel_or & -clad_ir #outside of fuel cylinder and inside of clad inner cylinder
clad_region = +clad_ir & -clad_or #outside of clad inner cylinder and inside of clad outer cylinder
fuel = openmc.Cell(1, 'fuel')
fuel.fill = uo2
fuel.region = fuel_region
#notice that for the gap between the fuel and the clad we do not need to link a material
# we consider that there is void there (considering the low density of the filling gas this is a fair approximation)
gap = openmc.Cell(2, 'air gap')
gap.region = gap_region
clad = openmc.Cell(3, 'clad')
clad.fill = zirconium
clad.region = clad_region
pitch = 1.26
#we define the x and y planes with boundary condition
left = openmc.XPlane(x0=-pitch/2, boundary_type='reflective')
right = openmc.XPlane(x0=pitch/2, boundary_type='reflective')
bottom = openmc.YPlane(y0=-pitch/2, boundary_type='reflective')
top = openmc.YPlane(y0=pitch/2, boundary_type='reflective')
#outside of left and inside of right, outside of bottom, and inside of top and outside of clad outer cylinder
water_region = +left & -right & +bottom & -top & +clad_or
moderator = openmc.Cell(4, 'moderator')
moderator.fill = water
moderator.region = water_region
root = openmc.Universe(cells=[fuel, gap, clad, moderator])
geom = openmc.Geometry()
geom.root_universe = root
geom.export_to_xml()
# -
# We can again inspect the xml file.
# !cat geometry.xml
# ### Tally
#
# We can describe the physical quantities of interest with tallies. If no tally is defined the code will only calculate the k-effective.
#
# As we can see in the openMC documentation (in the user's guide) a tally is a quantity defined as
#
# \begin{equation}
# X = \underbrace{\int d\mathbf{r} \int d\mathbf{\Omega} \int
# dE}_{\text{filters}} \underbrace{f(\mathbf{r}, \mathbf{\Omega},
# E)}_{\text{scores}} \psi (\mathbf{r}, \mathbf{\Omega}, E)
# \end{equation}
#
# For example if function $f()$ would be unity, we would score the flux in a certain part of the phase space. If function $f()$ is a macroscopic cross section, we would score reaction rates.
#
# Therefore we have to specify two things for a tally, which is created with a `Tally()` object:
#
# 1. *filters*: which part of the phase space we would like score at (for example only a certain part of the geometry is of interest, or certain energy bins)?
# 2. *scores*: what is the physical quantity of interest (for example flux, fission rate etc)
#
# You can find a detailed description of the available scores and filters in the documentation.
#
# Once the tallies are defined, we will need to link them into a `Tallies()` object, and export an xml file.
#
# In the following code block we define three tallies.
#
# 1. tally for the neutron spectrum and the fission rate vs energy in the fuel
# 2. tally for the neutron spectrum in the moderator
# 3. a Mesh tally along the x-axis to tally the spatial dependence of the flux with three energy groups (thermal: 0-1eV, epithermal: 1eV-100keV, fast: 100keV-20MeV)
#
# The reason why we tally the fission rate as well in the 1st tally is to see how openMC will store the results.
# +
# Tally 1: energy spectrum in fuel
cell_filter1 = openmc.CellFilter(fuel)
energybins=np.logspace(-2,7,1001) #1000 bins between 1e-2 eV and 1e7 eV
energy_filter = openmc.EnergyFilter(energybins)
t1 = openmc.Tally(1)
t1.filters = [cell_filter1,energy_filter]
t1.scores = ['flux','fission']
# Tally 2: energy spectrum in moderator
# we can use the same energy_filter as before
cell_filter2 = openmc.CellFilter(moderator)
t2 = openmc.Tally(2)
t2.filters = [cell_filter2,energy_filter]
t2.scores = ['flux']
# Tally 3: mesh tally in dimension x in three energy groups
energy_filter2 = openmc.EnergyFilter([0., 1, 100e3, 20.0e6])
myMesh=openmc.Mesh(name='xmesh')
myMesh.dimension=[100] #number of spatial bins along x axis
myMesh.lower_left=[-pitch/2]
myMesh.upper_right=[pitch/2]
mesh_filter = openmc.MeshFilter(myMesh)
t3 = openmc.Tally(3)
t3.filters = [mesh_filter,energy_filter2]
t3.scores = ['flux']
tallies = openmc.Tallies([t1,t2,t3])
tallies.export_to_xml()
# -
# We can inspect the xml file.
# !cat tallies.xml
# ### Settings
#
# There are some parameters we will need to set: we have to include the original source location and specify the number of batches and particles per batches for the criticality mode calculation. We also have to specify the number of "inactive" batches (these cycles are used to spread the fission source over the geometry, but the scores are still biased due to the original source not sampling all the possible fission sites properly).
#
# Since we have no better guess for the moment, we will place the source first in the center of the geometry. The number of batches and particles are set so that the results are reasonably accurate. If you wanted to have more accurate results you could increase these numbers, if you wanted to have shorter calculations, you could decrease these numbers.
#
# Again, the instance of the `Setting()` object is exported to xml.
# +
point = openmc.stats.Point((0, 0, 0))
src = openmc.Source(space=point)
settings = openmc.Settings()
settings.source = src
settings.batches = 100
settings.inactive = 10
settings.particles = 5000
settings.export_to_xml()
# -
# !cat settings.xml
# ## Plotting the geometry
#
# There are several ways of plotting the geometry. The simplest inline plotting option is to call the `plot()` method of the `Universe()` object (in our case this was called `root`). This is a wrapper of matplotlib. However, there is a more advanced options by setting up an instance of the `Plot()` class, and running openMC directly to generate a plot. Since in this course we work with a rather simple geometry, we will not look at the advanced plotting option, but you are welcome to check the user's guide.
root.plot()
# ## Calculation
#
# Finally, we are done, and nothing left just to run openMC and wait for the results. For this we have to specify where the cross section files are (this we could do in the system as well). Then call the `openmc.run()` method. And you immediately see how the k-effective is being estimated.
#
# **Note 1**: The results are being stored in a file named 'statepoint.100.h5' (100 refers to the number of batches), also a 'summary.h5' and some other files are created. Sometimes openMC is complaining if there are already existing h5 files, so in this case we can remove (`rm`) these files. For this we make a system call with `os.system` to remove any file for which the name starts with 's' and ands with 'h5'. Nevertheless, be careful with removing files, not to loose data.
#
# **Note 2**: Sometimes the python API breaks for no apparent reason, you can just restart the kernel, if that happens.
# +
import os
os.system('rm s*h5')
openmc.run()
# -
# ## Post-processing
#
# Alright, we did our calculation, we even saw that some k-effective values were printed, but we still haven't seen any more results. For this we will need to read in the statepoint file. We will read this file into an object called `sp`. If we hit TAB while typing `sp.` we can review the available methods and attributes. For example we can get the final k-effective as below.
sp = openmc.StatePoint('statepoint.100.h5')
sp.k_combined
# And the values we can access for further processing or plotting:
keff=sp.k_combined.nominal_value
keff_error=sp.k_combined.std_dev
print(keff,keff_error)
# The tallies are stored in a dictionary where the keys are the numeric IDs we defined previously:
sp.tallies
# We can convert the tally results into pandas dataframes. So at the end we have a nice table, for each energy bin there is a row for the flux and a row for the fission rate. If we prefer we can split the dataframe based on conditions to store separately the flux and the fission rate. For this we can apply conditions as we have seen during the previous datalabs.
#
# The results are stored in the 'mean' column.
tallydf1=sp.tallies[1].get_pandas_dataframe()
tallydf1.head() #prints the first 5 rows
tallydf1flux=tallydf1[tallydf1['score']=='flux']
tallydf1fiss=tallydf1[tallydf1['score']=='fission']
tallydf1flux.head()
# Let's plot these results.
energy=(tallydf1flux['energy low [eV]']+tallydf1flux['energy high [eV]'])/2
plt.figure()
plt.loglog(energy,tallydf1flux['mean'])
plt.xlabel('energy (eV)')
plt.ylabel('Group flux per source particle')
plt.show()
# These of course does not look like the spectrum we saw during the lecture. But here actually each value is not the flux at a certain energy, but the integral of the flux between energies. If we divide the integral flux with the width of the bins (`deltaE`) we get the more familiar shape for the spectrum.
#
# We can clearly see the Maxwellien thermal component, the 1/E part with the self-shielding effect of the resonances, and the Watt-spectrum at high energies.
#
# The very first "negative" peak is due to the 6.67 eV resonance of U-238.
# +
deltaE=(tallydf1flux['energy high [eV]']-tallydf1flux['energy low [eV]'])
plt.figure()
plt.loglog(energy,tallydf1flux['mean']/deltaE,lw=2)
plt.ylabel('Spectrum per source particle (1/eV)')
plt.xlabel('Energy (eV)')
plt.show()
# -
# Now it is your turn to plot the spectrum in the moderator (preferably include both the fuel and the moderator spectrum in the same figure). You will need to `get_pandas_dataframe` from tally 2, and then plot the results. Compare with the spectrum in the fuel, what is the most noticable difference?
# +
# your code comes here
# -
# Your conclusion comes here.
# Finally, we can look at the spatial dependence of the flux. We read in the 3rd tally. Try to split the dataset according to the energy groups and plot the flux vs x-coordinate for the three energy groups separately. Normalize each curve by their maximum, so they are comparable.
#
# Conclude your findings!
tallydf3=sp.tallies[3].get_pandas_dataframe()
tallydf3.head()
# +
# your code comes here
# -
# Your conclusion comes here!
# ## Experiment time!
#
# If time permits, modify your input, and see how the results change. Points of interest:
#
# - Change the void content (ie. decrease the moderator density), and observe how the k-eff changes. What happens with the spectrum?
# - What is the impact on the k-eff if you change the UO2 temperature?
# - What is the impact of increasing the pitch of the pincell?
#
# You can also take a look at the third set of home assignments, where you will need to implement various geometries in openMC, and already give it a try.
| Datalabs/Datalab06/6-openMCIntro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # %matplotlib widget
# %load_ext autoreload
# %autoreload 2
# ## Imports
# +
# modified from test.ipynb and main.py
import time
from utils import plot_3d_boundary, plot_3d_observed_rewards
from pylab import *
from matplotlib import pyplot as plt
from matplotlib import cm
import numpy as np
import GPy
import warnings
warnings.filterwarnings('ignore')
import logging
logging.basicConfig(level=logging.INFO)
from emukit.model_wrappers import GPyModelWrapper
from emukit.model_wrappers.gpy_quadrature_wrappers import BaseGaussianProcessGPy, RBFGPy
from emukit.core import ParameterSpace, ContinuousParameter, DiscreteParameter
from emukit.core.loop import UserFunctionWrapper
from emukit.core import ParameterSpace, ContinuousParameter
from emukit.core.initial_designs import RandomDesign
from GPy.models import GPRegression
from skopt.benchmarks import branin as _branin
from emukit.test_functions import branin_function
from scse.api.simulation import run_simulation
from matplotlib.colors import LogNorm
from matplotlib import pyplot as plt
# Decision loops
from emukit.experimental_design import ExperimentalDesignLoop
from emukit.bayesian_optimization.loops import BayesianOptimizationLoop
from emukit.quadrature.loop import VanillaBayesianQuadratureLoop
# Acquisition functions
from emukit.bayesian_optimization.acquisitions import ExpectedImprovement
from emukit.experimental_design.acquisitions import ModelVariance
# from emukit.quadrature.acquisitions import IntegralVarianceReduction
from emukit.experimental_design.acquisitions import IntegratedVarianceReduction
# Acquistion optimizers
from emukit.core.optimization import GradientAcquisitionOptimizer
# Stopping conditions
from emukit.core.loop import FixedIterationsStoppingCondition
from emukit.core.loop import ConvergenceStoppingCondition
from emukit.bayesian_optimization.acquisitions.log_acquisition import LogAcquisition
# Constraints
from emukit.core.constraints import LinearInequalityConstraint
# -
# Libraries for multiprocessing
from multiprocess.pool import Pool
from tqdm.auto import tqdm
from loop import *
# ## Experimental Setup
# +
surge_modulator = 1.0
all_scenarios = {"Baseline (20 % Discount)": {"discharge_discount": 0.8,
"charging_discount": 1.05,
"surge_modulator": surge_modulator},
"25 % Discount,": {"discharge_discount": 0.75,
"charging_discount": 1.05,
"surge_modulator": surge_modulator},
"33 % Discount": {"discharge_discount": 0.67,
"charging_discount": 1.05,
"surge_modulator": surge_modulator},
"33 % Discount +": {"discharge_discount": 0.67,
"charging_discount": 1.0,
"surge_modulator": surge_modulator},
"50 % Discount": {"discharge_discount": 0.5,
"charging_discount": 1.05,
"surge_modulator": surge_modulator}}
scenario = "Baseline (20 % Discount)"
# scenario = "33 % Discount"sounds g
# +
timesteps_per_day = 48
num_days = 10
time_horizon_value = timesteps_per_day*num_days
def invoke_miniscot(x):
"""
Handling single API call to miniSCOT simulation given some inputs
x contains parameter configs x = [x0 x1 ...]
- The order of parameters in x should follow the order specified in the parameter_space declaration
- E.g. here we specify num_batteries = x[0]
"""
kwargs = {
'time_horizon': time_horizon_value,
'num_batteries': int(x[0])
}
kwargs.update(all_scenarios[scenario])
# kwargs["surge_modulator"] = 1.5
# kwargs["solar_surge_modulator"] = 0.5
# kwargs["surge_scenario"] = "wind+solar"
kwargs["surge_modulator"] = 1.0
kwargs["solar_surge_modulator"] = 1.0
kwargs["surge_scenario"] = "solar"
cum_reward = run_simulation(**kwargs)
return cum_reward[-1]
# -
def f(X):
"""
Handling multiple API calls to miniSCOT simulation given some inputs
X is a matrix of parameters
- Each row is a set of parameters
- The order of parameters in the row should follow the order specified in the parameter_space declaration
"""
Y = []
for x in X:
cum_reward = invoke_miniscot(x)
# Note that we negate the reward; want to find min
Y.append(-cum_reward[-1])
Y = np.reshape(np.array(Y), (-1, 1))
return Y
def f_multiprocess(X):
"""
Handling multiple API calls to miniSCOT simulation given some inputs using multiprocessing.
X is a matrix of parameters
- Each row is a set of parameters
- The order of parameters in the row should follow the order specified in the parameter_space declaration
"""
# Set to None to use all available CPU
max_pool = 5
with Pool(max_pool) as p:
Y = list(
tqdm(
p.imap(invoke_miniscot, X),
total=X.shape[0]
)
)
# Note that we negate the reward; want to find min
Y = -np.reshape(np.array(Y), (-1, 1))
return Y
# +
# Basic plotting function
def plot_reward(X, Y, labels):
"""
Plots reward against a maximum of two dimensions.
"""
plt.style.use('seaborn')
fig = plt.figure(figsize=(12, 12))
order = np.argsort(X[:,0])
if X.shape[1] == 1:
ax = plt.axes()
ax.plot(X[order,0], Y[order])
ax.set_xlabel(labels[0])
ax.set_ylabel("Cumulative reward")
elif X.shape[1] == 2:
ax = plt.axes(projection='3d')
im = ax.plot_trisurf(X[order,0].flatten(), X[order,1].flatten(), Y[order].flatten(), cmap=cm.get_cmap('autumn'))
fig.colorbar(im)
ax.view_init(90, 90)
ax.set_xlabel(labels[0])
ax.set_ylabel(labels[1])
ax.set_zlabel("Cumulative reward") # (£/MWh)
else:
raise ValueError('X has too many dimensions to plot - max 2 allowed')
return fig, ax
# -
# ## Run Experiments
from scse.default_run_parameters.national_grid_default_run_parameters import DEFAULT_RUN_PARAMETERS
# +
max_num_batteries = 1000
# units in £/
min_battery_penalty = DEFAULT_RUN_PARAMETERS.battery_penalty - 100000
max_battery_penalty = DEFAULT_RUN_PARAMETERS.battery_penalty + 200000
min_battery_capacity = 1
max_battery_capacity = 80
num_data_points = 30
# timesteps_per_day = 48
# num_days = 30
# time_horizon_value = timesteps_per_day*time_horizon_value
# min_time_horizon_value = timesteps_per_week * 1
# max_time_horizon_value = timesteps_per_week * 2
num_batteries = DiscreteParameter('num_batteries', np.linspace(0, max_num_batteries, num_data_points))
# max_battery_capacities = DiscreteParameter('max_battery_capacity', range(min_battery_capacity, max_battery_capacity+1))
# battery_penalty = DiscreteParameter(
# 'battery_penalty', range(min_battery_penalty, max_battery_penalty+1))
parameters = [num_batteries]
parameter_space = ParameterSpace(parameters)
# +
# Get and check the parameters of the intial values (X)
design = RandomDesign(parameter_space)
X = design.get_samples(num_data_points)
print(X.shape)
X = np.linspace(0, max_num_batteries+1, num_data_points).reshape([num_data_points, 1])
X, X.shape
# -
# ### Get initial data points
# +
design = RandomDesign(parameter_space)
start = time.time()
Y = f_multiprocess(X)
end = time.time()
print("Getting {} initial simulation points took {} seconds".format(
num_data_points, round(end - start, 0)))
# -
# ### Check the cum. reward from the initial points to ensure we are seeing reasonable behaviour before fitting
plot_reward(X, Y, parameter_space.parameter_names)
# ### Run BO
1e0
# +
successful_sample = False
num_tries = 0
max_num_tries = 3
use_default= False
use_ard=False
while not successful_sample and num_tries < max_num_tries:
print(f"CURRENT ATTEMPT #{num_tries}")
# emulator model
if use_default:
gpy_model = GPRegression(X, Y)
else:
kernel = GPy.kern.RBF(1, lengthscale=1e0, variance=1e4, ARD=use_ard)
gpy_model = GPy.models.GPRegression(X, Y, kernel, noise_var=1e-10)
try:
gpy_model.optimize()
print("okay to optimize")
model_emukit = GPyModelWrapper(gpy_model)
# Load core elements for Bayesian optimization
expected_improvement = ExpectedImprovement(model=model_emukit)
optimizer = GradientAcquisitionOptimizer(space=parameter_space)
# Create the Bayesian optimization object
batch_size = 3
bayesopt_loop = BayesianOptimizationLoop(model=model_emukit,
space=parameter_space,
acquisition=expected_improvement,
batch_size=batch_size)
# Run the loop and extract the optimum; we either complete 10 steps or converge
max_iters = 3#10
stopping_condition = (
FixedIterationsStoppingCondition(i_max=max_iters) | ConvergenceStoppingCondition(eps=0.01)
)
bayesopt_loop.run_loop(f_multiprocess, stopping_condition)
print("successfully ran loop")
successful_sample = True
except:
num_tries += 1
# -
# ### Get new points from BO
# +
new_X, new_Y = bayesopt_loop.loop_state.X, bayesopt_loop.loop_state.Y
new_order = np.argsort(new_X[:,0])
new_X = new_X[new_order,:]
new_Y = new_Y[new_order]
# -
bayesopt_loop.loop_state.X
1055995.3436095843
# ### Plot results from BO with the GP fit
# +
x_plot = np.reshape(np.array([i for i in range(0, max_num_batteries)]), (-1,1))
mu_plot, var_plot = model_emukit.predict(x_plot)
plt.figure(figsize=(12, 8))
#plt.figure(figsize=(7, 5))
LEGEND_SIZE = 15
plt.plot(new_X, new_Y, "ro", markersize=10, label="All observations")
plt.plot(X, Y, "bo", markersize=10, label="Initial observations")
# plt.plot(x_plot, y_plot, "k", label="Objective Function")
plt.plot(x_plot, mu_plot, "C0", label="Model")
plt.fill_between(x_plot[:, 0],
mu_plot[:, 0] + np.sqrt(var_plot)[:, 0],
mu_plot[:, 0] - np.sqrt(var_plot)[:, 0], color="C0", alpha=0.6)
plt.fill_between(x_plot[:, 0],
mu_plot[:, 0] + 2 * np.sqrt(var_plot)[:, 0],
mu_plot[:, 0] - 2 * np.sqrt(var_plot)[:, 0], color="C0", alpha=0.4)
plt.fill_between(x_plot[:, 0],
mu_plot[:, 0] + 3 * np.sqrt(var_plot)[:, 0],
mu_plot[:, 0] - 3 * np.sqrt(var_plot)[:, 0], color="C0", alpha=0.2)
plt.legend(prop={'size': 14})
plt.xlabel("Number of Batteries", fontsize=14)
plt.ylabel("Cumulative Reward (£)", fontsize=14)
plt.grid(True)
plt.yticks(fontsize=13)
plt.xticks(fontsize=13)
# plt.show()
plt.savefig("surge.png")
# -
results = bayesopt_loop.get_results()
results.minimum_location, results.minimum_value # , results.best_found_value_per_iteration
results.minimum_location[0]
| src/bayesian_optimization/single_run_surge_sims.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Forecast of packages delay on the basis of existing schedule and delay by Tecnimont on Amur GPP Project
# Lets import all required libraries
# +
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.ensemble import RandomForestRegressor
from sklearn.preprocessing import LabelEncoder
import numpy as np
import pandas as pd
import statsmodels.stats.api as sm
# %pylab inline
# -
# Read a CVS file that was prepared and cleaned in Excel
p3 = pd.read_csv('for_analysis1.csv', header=0, sep=';', decimal=',', engine='python')
p3.head(20)
# Lets look into the dataset first
p3.describe()
p3.plot(kind='box', subplots=True, layout=(2,2), sharex=False, sharey=False)
p3.hist('delay')
pd.plotting.scatter_matrix(p3, alpha=0.2)
# Process a DataFrame, remove NAs and uninformative columns
p3.drop(['plan_afc','fact_afc', 'pckg', 'qty', 'cat'], axis=1, inplace=True)
p3.dropna(inplace = True)
p3.isna().sum()
for column in p3.columns.values:
print(column)
if p3[column].dtype != np.int64 and p3[column].dtype != np.float64:
enc = LabelEncoder()
enc.fit(p3[column].astype(str))
p3[column] = enc.transform(p3[column])
# Split dataframe in Train and Test pieces
X_train, X_test, y_train, y_test = train_test_split(p3.iloc[:, p3.columns != 'delay'], p3['delay'], test_size=0.25)
p3.info()
# Lets train linear model first
lm = LinearRegression()
lm.fit(X_train, y_train)
sqrt(mean_squared_error(lm.predict(X_train), y_train))
sqrt(mean_squared_error(lm.predict(X_test), y_test))
# +
plt.figure(figsize(16,7))
plt.subplot(121)
pyplot.scatter(y_train, lm.predict(X_train), color="red", alpha=0.1)
plot(range(500), color='black')
grid()
pyplot.title('Train set', fontsize=20)
pyplot.xlabel('Quality', fontsize=14)
pyplot.ylabel('Estimated quality', fontsize=14)
plt.subplot(122)
pyplot.scatter(y_test, lm.predict(X_test), color="red", alpha=0.1)
plot(range(500), color='black')
grid()
pyplot.title('Test set', fontsize=20)
pyplot.xlabel('Quality', fontsize=14)
pyplot.ylabel('Estimated quality', fontsize=14)
# -
# Lets try a bunch of different methods.
from pandas import read_csv
from pandas.plotting import scatter_matrix
from matplotlib import pyplot
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
models = []
models.append(('LR', LogisticRegression(solver='liblinear', multi_class='ovr')))
models.append(('LDA', LinearDiscriminantAnalysis()))
models.append(('KNN', KNeighborsClassifier()))
models.append(('CART', DecisionTreeClassifier()))
models.append(('NB', GaussianNB()))
models.append(('SVM', SVC(gamma='auto')))
results = []
names = []
for name, model in models:
kfold = StratifiedKFold(n_splits=10, random_state=1, shuffle=True)
cv_results = cross_val_score(model, X_train, Y_train, cv=kfold, scoring='accuracy')
results.append(cv_results)
names.append(name)
print('%s: %f (%f)' % (name, cv_results.mean(), cv_results.std()))
# Lets try Random Forest
rf = RandomForestRegressor(n_estimators=100, min_samples_leaf=3)
rf.fit(X_train, y_train)
sqrt(mean_squared_error(rf.predict(X_train), y_train))
sqrt(mean_squared_error(rf.predict(X_test), y_test))
# +
plt.figure(figsize(16,7))
plt.subplot(121)
pyplot.scatter(y_train, rf.predict(X_train), color="red", alpha=0.1)
plot(range(500), color='black')
grid()
pyplot.title('Train set', fontsize=20)
pyplot.xlabel('Quality', fontsize=14)
pyplot.ylabel('Estimated quality', fontsize=14)
plt.subplot(122)
pyplot.scatter(y_test, rf.predict(X_test), color="red", alpha=0.1)
plot(range(500), color='black')
grid()
pyplot.title('Test set', fontsize=20)
pyplot.xlabel('Quality', fontsize=14)
pyplot.ylabel('Estimated quality', fontsize=14)
# -
rf.score(X_test, y_test)
importances = pd.DataFrame(zip(X_train.columns, rf.feature_importances_))
importances.columns = ['feature name', 'importance']
importances.sort_values(by='importance', ascending=False)
# We can see that most of the value is given to model by revision number, which is natural as each revision takes time to be issued.
# More methods need to be applied to the model to explore the possibility to forecast delay on the basis of package discipline and design institute.
| projects/Forecast of Tecnimont delays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.6-pre
# language: julia
# name: julia-0.4
# ---
# # #Validation of Flap model
include("../src/UNSflow.jl")
using UNSflow
# # # Laura's case C - flap only
#Kinematics
alpha_amp = 0.
alpha_mean = 0.
alpha_zl = 0. #Flat plate
h_amp = 0.
k = 3.93
beta_amp = 5*pi/180.
# +
#for UNSflow
w = 2*k
T = (2*pi/w)
ncyc = 4
t_tot = ncyc*T
dt = 0.015*0.2/(k*beta_amp)
nsteps =round(Int,t_tot/dt)+1
alphadef = ConstDef(alpha_amp)
hdef = ConstDef(h_amp)
udef = ConstDef(1.)
ndef = CosDef(0., beta_amp, w, 0.)
full_kinem = KinemDefwFlap(alphadef, hdef, udef, ndef)
pvt = 0.2 #Doesnt matter for flap only
lespcrit = [21;] #high value to turn off LEV shedding
x_b = [0.75;]
surf = TwoDSurfwFlap(1., 1., "FlatPlate", pvt, 70, 35, "Prescribed", full_kinem,x_b, lespcrit)
curfield = TwoDFlowField()
# -
ldvm(surf, curfield, nsteps, dt)
#for Theodorsen
theo_in = TheoDefwFlap(alpha_amp, h_amp, alpha_mean, alpha_zl, k, 0., pvt, beta_amp, x_b[1], 0.)
(t_theo, _, cl_h, cl_alpha, cl_beta, cl_theo) = theodorsen(theo_in)
# +
#Plots to compare
#UNSflow
data = readdlm("results.dat")
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (data[range,1]-data[range[1]])/T
plot(tbyT, data[range,6])
plot(t_theo, imag(cl_theo))
# -
# # It seems to be working but for some reason, the imaginary part of Theodorsen is giving the correct value. It should be the real part for a cosine function. Possibly a missing i.
# # Lets try a sine flap
ndef = SinDef(0., beta_amp, w, 0.)
# everything else is unchanged
full_kinem = KinemDefwFlap(alphadef, hdef, udef, ndef)
# +
surf = TwoDSurfwFlap(1., 1., "FlatPlate", pvt, 70, 35, "Prescribed", full_kinem,x_b, lespcrit)
curfield = TwoDFlowField()
# -
ldvm(surf, curfield, nsteps, dt)
# +
#Plots to compare
#UNSflow
data = readdlm("results.dat")
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (data[range,1]-data[range[1]])/T
plot(tbyT, data[range,6])
plot(t_theo, -real(cl_theo))
# -
# # Negative of the real value corresponds to the sine results. So it seems like a extra i, not a missing one. In any case, the LDVM implementation seems right
# # # Variation of Laura's case E - Pitch and flap
#Kinematics
alpha_amp = 5*pi/180
alpha_mean = 0.
alpha_zl = 0. #Flat plate
h_amp = 0.
k = 3.93
beta_amp = 5*pi/180.
# +
#for UNSflow
w = 2*k
T = (2*pi/w)
ncyc = 4
t_tot = ncyc*T
dt = 0.015*0.2/(k*alpha_amp)
nsteps =round(Int,t_tot/dt)+1
# +
#for UNSflow
w = 2*k
T = (2*pi/w)
ncyc = 4
t_tot = ncyc*T
dt = 0.015*0.2/(k*beta_amp)
nsteps =round(Int,t_tot/dt)+1
alphadef = CosDef(alpha_mean, alpha_amp, w, 0.)
hdef = ConstDef(h_amp)
udef = ConstDef(1.)
ndef = CosDef(0., beta_amp, w, 0.)
full_kinem = KinemDefwFlap(alphadef, hdef, udef, ndef)
pvt = 0.2
lespcrit = [21;] #high value to turn off LEV shedding
x_b = [0.75;]
surf = TwoDSurfwFlap(1., 1., "FlatPlate", pvt, 70, 35, "Prescribed", full_kinem,x_b, lespcrit)
curfield = TwoDFlowField()
# -
ldvm(surf, curfield, nsteps, dt)
#for Theodorsen
theo_in = TheoDefwFlap(alpha_amp, h_amp, alpha_mean, alpha_zl, k, 0., pvt, beta_amp, x_b[1], 0.)
(t_theo, _, cl_h, cl_alpha, cl_beta, cl_theo) = theodorsen(theo_in)
# +
#Plots to compare
#UNSflow
data = readdlm("results.dat")
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (data[range,1]-data[range[1]])/T
plot(tbyT, data[range,6])
plot(t_theo, real(cl_theo))
# -
# # Looks good. alpha=10 + flap=5 becomes large angle and we start seeing differences between LDVM and theodorsen
| Notebooks/.ipynb_checkpoints/Flap_validation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This material is copied (possibily with some modifications) from the [Python for Text-Analysis course](https://github.com/cltl/python-for-text-analysis/tree/master/Chapters).
# # Chapter 12: Functions and scope
# *We use an example from [this website](http://anh.cs.luc.edu/python/hands-on/3.1/handsonHtml/functions.html) to show you some of the basics of writing a function.
# We use some materials from [this other Python course](https://github.com/kadarakos/python-course).*
# We have seen that Python has several built-in functions (e.g. `print()` or `max()`). But you can also create your own function. A function is a reusable block of code that performs a specific task. Once you have defined a function, you can use it at any place in your Python script. You can even import a function from an external module (as we will see in the next chapter). Therefore, they are beneficial for tasks that you will perform more often. Plus, functions are a convenient way to order your code and make it more readable!
#
# ### At the end of this chapter, you will be able to:
# * write your own function
# * work with function inputs
# * understand the difference between (keyword and positional) arguments and parameters
# * return zero, one, or multiple values
# * write function docstrings
# * understand scope of variables
# * store your function in a Python module and call it
# * debug your functions
#
# ### If you want to learn more about these topics, you might find the following link useful:
# * [Tutorial: Defining Functions of your Own](http://anh.cs.luc.edu/python/hands-on/3.1/handsonHtml/functions.html)
# * [The docstrings main formats](http://daouzli.com/blog/docstring.html)
# * [PEP 287 -- reStructured Docstring Format](https://www.python.org/dev/peps/pep-0287/)
# * [Introduction to assert](https://www.programiz.com/python-programming/assert-statement)
#
# **Now let's get started!**
# If you have **questions** about this chapter, contact Cody in the Slack group.
# # 1. Writing a function
#
# A **function** is an isolated chunk of code, that has a name, gets zero or more parameters, and returns a value. In general, a function will do something for you, based the input parameters you pass it, and it will typically return a result. You are not limited to using functions available in the standard library or the ones provided by external parties. You can also write your own functions!
#
# Whenever you are writing a function, you need to think of the following things:
# * What is the purpose of the function?
# * How should I name the function?
# * What input does the function need?
# * What output should the function generate?
#
# ## 1.1. Why use a function?
#
# There are several good reasons why functions are a vital component of any non-ridiculous programmer:
#
# * encapsulation: wrapping a piece of useful code into a function so that it can be used without knowledge of the specifics
# * generalization: making a piece of code useful in varied circumstances through parameters
# * manageability: Dividing a complex program up into easy-to-manage chunks
# * maintainability: using meaningful names to make the program more readable and understandable
# * reusability: a good function may be useful in multiple programs
# * recursion!
#
# ## 1.2. How to define a function
#
# Let's say we want to sing a birthday song to Emily. Then we print the following lines:
print("Happy Birthday to you!")
print("Happy Birthday to you!")
print("Happy Birthday, dear Emily.")
print("Happy Birthday to you!")
# This could be the purpose of a function: to print the lines of a birthday song for Emily.
# Now, we define a function to do this. Here is how you define a function:
#
# * write `def`;
# * the name you would like to call your function;
# * a set of parentheses containing the argument(s) of your function;
# * a colon;
# * a docstring describing what your function does;
# * the function definition;
# * ending with a return statement
#
# Statements must be indented, so that Python knows what belongs in the function and what not. Functions are only executed when you call them. It is good practice to define your functions at the top of your program or in another Python module.
#
# We give the function a clear name, `happy_birthday_to_emily` and we define the function as shown below. Note that we specify exactly what it does in the docstring in the beginning of the function:
def happy_birthday_to_emily(): # Function definition
"""
Print a birthday song to Emily.
"""
print("Happy Birthday to you!")
print("Happy Birthday to you!")
print("Happy Birthday, dear Emily.")
print("Happy Birthday to you!")
# If we execute the code above, we don't get any output. That's because we only told Python: "Here's a function to do this, please remember it." If we actually want Python to execute everything inside this function, we have to *call* it:
# ## 1.3 How to call a function
# It is important to distinguish between a function **definition** and a function **call**. We illustrate this in 1.3.1. You can also call functions from within other functions. This will become useful when you split up your code into small chunks that can be combined to solve a larger problem. This is illustrated in 1.3.2.
#
#
# ### 1.3.1) A simple function call
# A function is **defined** once. After the definition, Python has remembered what this function does in its memory.
# A function is **executed/called** as many times as we like. When calling a function, you should always use parentheses.
#
# +
# function definition:
def happy_birthday_to_emily(): # Function definition
"""
Print a birthday song to Emily.
"""
print("Happy Birthday to you!")
print("Happy Birthday to you!")
print("Happy Birthday, dear Emily.")
print("Happy Birthday to you!")
# function call:
print('Function call 1')
happy_birthday_to_emily()
print()
print('Function call 2')
# We can call the function as many times as we want (but we define it only once)
happy_birthday_to_emily()
print()
print('Function call 3')
happy_birthday_to_emily()
print()
# This will not call the function
# (without the parenthesis, Python thinks that `happy_birthday_to_emily`
# is a variable and not a function!):
print('This is not a function call')
happy_birthday_to_emily
# -
# ### 1.3.2 Calling a function from within another function
#
# We can also define functions that call other functions, which is very helpful if we want to split our task, into smaller, more manageable subtasks:
# +
def new_line():
"""Print a new line."""
print()
def two_new_lines():
"""Print two new lines."""
new_line()
new_line()
print("Printing a single line...")
new_line()
print("Printing two lines...")
two_new_lines()
print("Printed two lines")
# -
# You can do the same tricks that we learned to apply on the built-in functions, like asking for `help` or for a function `type`:
help(happy_birthday_to_emily)
type(happy_birthday_to_emily)
# The help we get on a function will become more interesting once we learn about function inputs and outputs ;-)
# ## 1.4 Working with function input
#
# ### 1.4.1 Parameters and arguments
# We use parameters and arguments to make a function execute a task depending on the input we provide. For instance, we can change the function above so we can input the name of a person and print a birthday song using this name. This results in a more generic function.
#
# To understand how we use **parameters** and **arguments**, keep in mind the distinction between function *definition* and function *call*.
#
# **Parameter**: The variable `name` in the **function definition** below is a **parameter**. Variables used in **function definitions** are called **parameters**.
#
# **Argument**: The variable `my_name` in the function call below is a value for the parameter `name` at the time when the function is called. We denote such variables **arguments**. We use arguments so we can direct the function to do different kinds of work when we call it at different times.
# function definition with using the parameter `name'
def happy_birthday(name):
"""
Print a birthday song with the "name" of the person inserted.
"""
print("Happy Birthday to you!")
print("Happy Birthday to you!")
print("Happy Birthday, dear " + name + ".")
print("Happy Birthday to you!")
# function call using specifying the value of the argument
happy_birthday("James")
# We can also store the name in a variable:
my_name="James"
happy_birthday(my_name)
# If we forgot to specify the name, we get an error:
happy_birthday()
# Functions can have multiple parameters. We can for example multiply two numbers in a function (using the two parameters x and y) and then call the function by giving it two arguments:
# +
def multiply(x, y):
"""Multiply two numeric values."""
result = x * y
print(result)
multiply(2020,527)
multiply(2,3)
# -
# ### 1.4.2 Positional vs keyword parameters and arguments
# The function definition tells Python which parameters are positional and which are keyword. As you might remember, positional means that you have to give an argument for that parameter; keyword means that you can give an argument value, but this is not necessary because there is a default value.
#
# So, to summarize these two notes, we distinguish between the following four categories:
#
#
# 1) **positional parameters**: (we indicate these when defining a function, and they are compulsory when calling the function)
#
# 2) **keyword parameters**: (we indicate these when defining a function, but they have a default value - and are optional when calling the function)
#
# 3) **positional arguments**: (we MUST specify these when calling a function)
#
# 4) **keyword arguments**: (we CAN specify these when calling a function)
# To summarize:
#
# | - | parameters | arguments |
# |---|---|---|
# | positional | indicated in definition | compulsory in call |
# | keyword | indicated in definition | optional in call |
# For example, if we want to have a function that can either multiply two or three numbers, we can make the third parameter a keyword parameter with a default of 1 (remember that any number multiplied with 1 results in that number):
def multiply(x, y, third_number=1): # x and y are positional parameters, third_number is a keyword parameter
"""Multiply two or three numbers and print the result."""
result=x*y*third_number
print(result)
multiply(2,3) # We only specify the positional arguments
multiply(2,3,third_number=4) # We specify both the positional arguments, and the keyword argument
# If we do not specify a positional argument, the function call will fail (with a very helpful error message):
multiply(3)
# ## 1.5 Output: the `return` statement
# Functions can have a **return** statement. The `return` statement returns a value back to the caller and **always** ends the execution of the function. This also allows us to use the result of a function outside of that function by assigning it to a variable:
# +
def multiply(x, y):
"""Multiply two numbers and return the result."""
multiplied = x * y
return multiplied
#here we assign the returned value to variable z
result = multiply(2, 5)
print(result)
# -
# We can also print the result directly (without assigning it to a varible), which gives us the same effect as using the print statements we used before:
print(multiply(30,20))
# If we assign the result to a variable, but do not use the return statement, the function cannot return it. Instead, it returns `None` (as you can try out below).
#
# This is important to realize: even functions without a `return` statement do return a value, albeit a rather boring one. This value is called `None` (it’s a built-in name). You have seen this already with list methods - for example `list.append(val)` adds a value to a list, but does not return anything explicitly.
#
# +
def multiply_no_return(x, y):
"""Multiply two numbers and does not return the result."""
result = x * y
is_this_a_result = multiply_no_return(2,3)
print(is_this_a_result)
# -
# **Returning multiple values**
#
# Similarly as the input, a function can also return **multiple values** as output. We call such a collection of values a *tuple* (does this term sound familiar ;-)?).
#
# +
def calculate(x,y):
"""Calculate product and sum of two numbers."""
product = x * y
summed = x + y
#we return a tuple of values
return product, summed
# the function returned a tuple and we unpack it to var1 and var2
var1, var2 = calculate(10,5)
print("product:",var1,"sum:",var2)
# -
# Make sure you actually save your 2 values into 2 variables, or else you end up with errors or unexpected behavior:
# +
#this will assign `var` to a tuple:
var = calculate(10,5)
print(var)
#this will generate an error
var1, var2, var3 = calculate(10,5)
# -
# Saving the resulting values in different variables can be useful when you want to use them in different places in your code:
# +
def sum_and_diff_len_strings(string1, string2):
"""
Return the sum of and difference between the lengths of two strings.
"""
sum_strings = len(string1) + len(string2)
diff_strings = len(string1) - len(string2)
return sum_strings, diff_strings
sum_strings, diff_strings = sum_and_diff_len_strings("horse", "dog")
print("Sum:", sum_strings)
print("Difference:", diff_strings)
# -
# ## 1.6 Documenting your functions with docstrings
#
# **Docstring** is a string that occurs as the first statement in a function definition.
#
# For consistency, always use """triple double quotes""" around docstrings. Triple quotes are used even though the string fits on one line. This makes it easy to later expand it.
#
# There's no blank line either before or after the docstring.
#
# The docstring is a phrase ending in a period. It prescribes the function or method's effect as a command ("Do this", "Return that"), not as a description; e.g. don't write "Returns the pathname ...".
#
# In practice, there are several formats for writing docstrings, and all of them contain more information than the single sentence description we mention here. Probably the most well-known format is reStructured Text. Here is an example of a function description in reStructured Text (reST):
#
def my_function(param1, param2):
"""
This is a reST style.
:param param1: this is a first param
:param param2: this is a second param
:returns: this is a description of what is returned
"""
return
# You can see that this docstring describes the function goal, its parameters, its outputs, and the errors it raises.
#
# It is a good practice to write a docstring for your functions, so we will always do this! For now we will stick with single-sentence docstrings
#
# You can read more about this topic [here](http://daouzli.com/blog/docstring.html), [here](https://stackoverflow.com/questions/3898572/what-is-the-standard-python-docstring-format), and [here](https://www.python.org/dev/peps/pep-0287/).
# ## 1.7 Debugging a function
# Sometimes, it can hard to write a function that works perfectly. A common practice in programming is to check whether the function performs as you expect it to do. The `assert` statement is one way of debugging your function. The syntax is as follows:
#
# assert `code` == `your expected output`,`message to show when code does not work as you'd expected`
# Let's try this on our simple function.
def is_even(p):
"""Check whether a number is even."""
if p % 2 == 1:
return False
else:
return True
# If the function behaves as we expect, Python will show nothing.
input_value = 2
expected_output = True
actual_output = is_even(input_value)
assert actual_output == expected_output, f'expected {expected_output}, got {actual_output}'
# However, when the actual output is different from what we expected, we got an error. Let's say we made a mistake in writing the function.
def is_even(p):
"""Check whether a number is even."""
if p % 2 == 1:
return False
else:
return False
input_value = 2
expected_output = True
actual_output = is_even(input_value)
assert actual_output == expected_output, f'expected {expected_output}, got {actual_output}'
# ## 1.8 Storing a function in a Python module
# Since Python functions are nice blocks of code with a clear focus, wouldn't it be nice if we can store them in a file? By doing this, we make our code visually very appealing since we are only left with functions calls instead of function definitions.
# Please open the file **utils_chapter11.py** (is in the same folder as the notebook you are now reading). In it, you will find three of the functions that we've shown so far in this notebook. So, how can we use those functions? We can `import` the function using the following syntax:
#
# `from` `NAME OF FILE WITHOUT .PY` `import` `function name`
from utils_functions_ch import happy_birthday
happy_birthday('George')
from utils_functions_ch import multiply
multiply(1,2)
from utils_functions_ch import is_even
is_it_even = is_even(5)
print(is_it_even)
# # 2. Variable scope
# Please note: scope is a hard concept to grasp but we think it is important to introduce it here. We will do our best to repeat it during the course.
# Any variables you declare in a function, as well as the parameters that are passed to a function will only exist within the **scope** of that function, i.e. inside the function itself. The following code will produce an error, because the variable `x` does not exist outside of the function:
# +
def setx():
"""Set the value of a variable to 1."""
x = 1
setx()
print(x)
# -
# Even when we return x, it does not exist outside of the function:
# +
def setx():
"""Set the value of a variable to 1."""
x = 1
return x
setx()
print(x)
# -
# Also consider this:
x = 0
def setx():
"""Set the value of a variable to 1."""
x = 1
setx()
print(x)
# In fact, this code has produced two completely unrelated `x`'s!
#
# So, you can not read a local variable outside of the local context. Nevertheless, it is possible to read a global variable from within a function, in a strictly read-only fashion.
# +
x = 1
def getx():
"""Print the value of a variable x."""
print(x)
getx()
# -
# You can use two built-in functions in Python when you are unsure whether a variable is local or global. The function `locals()` returns a list of all local variables, and the function `globals()` - a list of all global variables. Note that there are many non-interesting system variables that these functions return, so in practice it is best to check for membership with the `in` operator. For example:
# +
a=3
b=2
def setb():
"""Set the value of a variable b to 11."""
b=11
c=20
print("Is 'a' defined locally in the function:", 'a' in locals())
print("Is 'b' defined locally in the function:", 'b' in locals())
print("Is 'b' defined globally:", 'b' in globals())
setb()
print("Is 'a' defined globally:", 'a' in globals())
print("Is 'b' defined globally:", 'b' in globals())
print("Is 'c' defined globally:", 'c' in globals())
# -
# Finally, note that the local context stays local to the function, and is not shared even with other functions called within a function, for example:
# +
def setb_again():
"""Set the value of a variable to 3."""
b=3
print("in 'setb_again' b =", b)
def setb():
"""Set the value of a variable b to 2."""
b=2
setb_again()
print("in 'setb' b =", b)
b=1
setb()
print("global b =", b)
# -
# We call the function `setb()` from the global context, and we call the function `setb_again()` from the context of the function `setb()`. The variable `b` in the function `setb_again()` is set to 3, but this does not affect the value of this variable in the function `setb()` which is still 2. And as we saw before, the changes in `setb()` do not influence the value of the global variable (`b=1`).
# # Exercises
# **Exercise 1:**
#
# Write a function that converts meters to centimeters and prints the resulting value.
# +
# you code here
# -
# **Exercise 2**:
#
# Add another keyword parameter `message` to the multiply function, which will allow a user to print a message. The default value of this keyword parameter should be an empty string. Test this with 2 messages of your choice. Also test it without specifying a value for the keyword argument when calling a function.
# +
# function to modify:
def multiply(x, y, third_number=1):
"""Multiply two or three numbers and print the result."""
result=x*y*third_number
print(result)
# -
# **Exercise 3:**
#
# Write a function called `multiple_new_lines` which takes as argument an integer and prints that many newlines by calling the function newLine.
# +
def new_line():
"""Return a new line."""
return '\n'
# your code here
# -
# **Exercise 4:**
#
# Let's refactor the happy birthday function to have no repetition. Note that previously we print "Happy birthday to you!" three times. Make another function `happy_birthday_to_you()` that only prints this line and call it inside the function `happy_birthday(name)` in place of the strings.
# +
def happy_birthday_to_you():
# your code here
# original function - replace the print statements by the happy_birthday_to_you() function:
def happy_birthday(name):
"""
Print a birthday song with the "name" of the person inserted.
"""
print("Happy Birthday to you!")
print("Happy Birthday to you!")
print("Happy Birthday, dear " + name + ".")
print("Happy Birthday to you!")
# -
# **Exercise 5:**
#
# Try to figure out what is going on in the following examples. Think about how Python deals with the order of calling functions. For each print statement, write a short explanation (in markdown or with `#` notes) of the order of events.
# +
def multiply(x, y, third_number=1):
"""Multiply two or three numbers and print the result."""
result=x*y*third_number
return result
print(multiply(1+1,6-2))
print(multiply(multiply(4,2),multiply(2,5)))
print(len(str(multiply(10,100))))
# -
# **Exercise 6:**
#
# Complete this code to switch the values of two variables:
# +
def switch_two_values(x,y):
# your code here
a='orange'
b='apple'
a,b = switch_two_values(a,b) # `a` should contain "apple" after this call, and `b` should contain "orange"
print(a,b)
| chapters/Chapter 12 - Functions and scope.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # AutoML 05 : Blacklisting models, Early termination and handling missing data
#
# In this example we use the scikit learn's [digit dataset](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) to showcase how you can use AutoML for handling missing values in data. We also provide a stopping metric indicating a target for the primary metric so that AutoML can terminate the run without necessarly going through all the iterations. Finally, if you want to avoid a certain pipeline, we allow you to specify a black list of algos that AutoML will ignore for this run.
#
# Make sure you have executed the [00.configuration](00.configuration.ipynb) before running this notebook.
#
# In this notebook you would see
# 1. Creating an Experiment using an existing Workspace
# 2. Instantiating AutoMLConfig
# 4. Training the Model
# 5. Exploring the results
# 6. Testing the fitted model
#
# In addition this notebook showcases the following features
# - **Blacklist** certain pipelines
# - Specify a **target metrics** to indicate stopping criteria
# - Handling **Missing Data** in the input
#
#
# ## Create Experiment
#
# As part of the setup you have already created a <b>Workspace</b>. For AutoML you would need to create an <b>Experiment</b>. An <b>Experiment</b> is a named object in a <b>Workspace</b>, which is used to run experiments.
# +
import logging
import os
import random
from matplotlib import pyplot as plt
from matplotlib.pyplot import imshow
import numpy as np
import pandas as pd
from sklearn import datasets
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
from azureml.train.automl.run import AutoMLRun
# +
ws = Workspace.from_config()
# choose a name for the experiment
experiment_name = 'automl-local-missing-data'
# project folder
project_folder = './sample_projects/automl-local-missing-data'
experiment=Experiment(ws, experiment_name)
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Project Directory'] = project_folder
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
pd.DataFrame(data=output, index=['']).T
# -
# ## Diagnostics
#
# Opt-in diagnostics for better experience, quality, and security of future releases
from azureml.telemetry import set_diagnostics_collection
set_diagnostics_collection(send_diagnostics=True)
# ### Creating Missing Data
# +
from scipy import sparse
digits = datasets.load_digits()
X_digits = digits.data[10:,:]
y_digits = digits.target[10:]
# Add missing values in 75% of the lines
missing_rate = 0.75
n_missing_samples = int(np.floor(X_digits.shape[0] * missing_rate))
missing_samples = np.hstack((np.zeros(X_digits.shape[0] - n_missing_samples, dtype=np.bool), np.ones(n_missing_samples, dtype=np.bool)))
rng = np.random.RandomState(0)
rng.shuffle(missing_samples)
missing_features = rng.randint(0, X_digits.shape[1], n_missing_samples)
X_digits[np.where(missing_samples)[0], missing_features] = np.nan
# -
df = pd.DataFrame(data=X_digits)
df['Label'] = pd.Series(y_digits, index=df.index)
df.head()
# ## Instantiate Auto ML Config
#
#
# This defines the settings and data used to run the experiment.
#
# |Property|Description|
# |-|-|
# |**task**|classification or regression|
# |**primary_metric**|This is the metric that you want to optimize.<br> Classification supports the following primary metrics <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>balanced_accuracy</i><br><i>average_precision_score_weighted</i><br><i>precision_score_weighted</i>|
# |**max_time_sec**|Time limit in seconds for each iteration|
# |**iterations**|Number of iterations. In each iteration Auto ML trains the data with a specific pipeline|
# |**n_cross_validations**|Number of cross validation splits|
# |**preprocess**| *True/False* <br>Setting this to *True* enables Auto ML to perform preprocessing <br>on the input to handle *missing data*, and perform some common *feature extraction*|
# |**exit_score**|*double* value indicating the target for *primary_metric*. <br> Once the target is surpassed the run terminates|
# |**blacklist_algos**|*Array* of *strings* indicating pipelines to ignore for Auto ML.<br><br> Allowed values for **Classification**<br><i>LogisticRegression</i><br><i>SGDClassifierWrapper</i><br><i>NBWrapper</i><br><i>BernoulliNB</i><br><i>SVCWrapper</i><br><i>LinearSVMWrapper</i><br><i>KNeighborsClassifier</i><br><i>DecisionTreeClassifier</i><br><i>RandomForestClassifier</i><br><i>ExtraTreesClassifier</i><br><i>LightGBMClassifier</i><br><br>Allowed values for **Regression**<br><i>ElasticNet<i><br><i>GradientBoostingRegressor<i><br><i>DecisionTreeRegressor<i><br><i>KNeighborsRegressor<i><br><i>LassoLars<i><br><i>SGDRegressor<i><br><i>RandomForestRegressor<i><br><i>ExtraTreesRegressor<i>|
# |**X**|(sparse) array-like, shape = [n_samples, n_features]|
# |**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]<br>Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers. |
# |**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder. |
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
primary_metric = 'AUC_weighted',
max_time_sec = 3600,
iterations = 20,
n_cross_validations = 5,
preprocess = True,
exit_score = 0.994,
blacklist_algos = ['KNeighborsClassifier','LinearSVMWrapper'],
verbosity = logging.INFO,
X = X_digits,
y = y_digits,
path=project_folder)
# ## Training the Model
#
# You can call the submit method on the experiment object and pass the run configuration. For Local runs the execution is synchronous. Depending on the data and number of iterations this can run for while.
# You will see the currently running iterations printing to the console.
local_run = experiment.submit(automl_config, show_output=True)
# ## Exploring the results
# #### Widget for monitoring runs
#
# The widget will sit on "loading" until the first iteration completed, then you will see an auto-updating graph and table show up. It refreshed once per minute, so you should see the graph update as child runs complete.
#
# NOTE: The widget will display a link at the bottom. This will not currently work, but will eventually link to a web-ui to explore the individual run details.
from azureml.train.widgets import RunDetails
RunDetails(local_run).show()
#
# #### Retrieve All Child Runs
# You can also use sdk methods to fetch all the child runs and see individual metrics that we log.
# +
children = list(local_run.get_children())
metricslist = {}
for run in children:
properties = run.get_properties()
metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}
metricslist[int(properties['iteration'])] = metrics
rundata = pd.DataFrame(metricslist).sort_index(1)
rundata
# -
# ### Retrieve the Best Model
#
# Below we select the best pipeline from our iterations. Each pipeline is a tuple of three elements. The first element is the score for the pipeline the second element is the string description of the pipeline and the last element are the pipeline objects used for each fold in the cross-validation.
best_run, fitted_model = local_run.get_output()
# #### Best Model based on any other metric
# +
# lookup_metric = "accuracy"
# best_run, fitted_model = local_run.get_output(metric=lookup_metric)
# -
# #### Model from a specific iteration
# +
# iteration = 3
# best_run, fitted_model = local_run.get_output(iteration=iteration)
# -
# ### Register fitted model for deployment
description = 'AutoML Model'
tags = None
local_run.register_model(description=description, tags=tags)
local_run.model_id # Use this id to deploy the model as a web service in Azure
# ### Testing the Fitted Model
# +
digits = datasets.load_digits()
X_digits = digits.data[:10, :]
y_digits = digits.target[:10]
images = digits.images[:10]
#Randomly select digits and test
for index in np.random.choice(len(y_digits), 2):
print(index)
predicted = fitted_model.predict(X_digits[index:index + 1])[0]
label = y_digits[index]
title = "Label value = %d Predicted value = %d " % ( label,predicted)
fig = plt.figure(1, figsize=(3,3))
ax1 = fig.add_axes((0,0,.8,.8))
ax1.set_title(title)
plt.imshow(images[index], cmap=plt.cm.gray_r, interpolation='nearest')
plt.show()
| automl/05.auto-ml-missing-data-Blacklist-Early-Termination.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# ## Single qubit annealing
# This tutorial will recreate the single-qubit example in this paper: [Decoherence in adiabatic quantum computation](https://arxiv.org/abs/1503.08767).
#
# The Hamiltonian of this example is
#
# $$H(s) = -\frac{1}{2}(1-s)\sigma_x - \frac{1}{2}s\sigma_z \ ,$$
#
# which can be constructed by the following code block
using OpenQuantumTools, OrdinaryDiffEq, Plots
H = DenseHamiltonian([(s)->1-s, (s)->s], -[σx, σz]/2)
# This package directly interacts with [Plots.jl](https://github.com/JuliaPlots/Plots.jl) by defining [recipes](https://github.com/JuliaPlots/RecipesBase.jl). We can visualize the spectrum of the Hamiltonian by directly plotting the object:
# this plot recipe is for conveniently plotting the spectrum of the Hamiltonian
# the first 3 arguments are: the Hamiltonian, the grid `s` and the levels to keep
plot(H, 0:0.01:1, 2, linewidth=2)
# ### Unit ($h$ or $\hbar$)
# A keyword argument `unit` whose default value is `:h` can be provided to any Hamiltonian type's constructor. This argument specifies the unit of other input arguments. For example, setting `unit` to `:h` means the other input arguments have the unit of $\mathrm{GHz}$, while setting it to `:ħ` means the other input arguments have the unit of $2\pi\mathrm{GHz}$. To evaluate the Hamiltonian at a given time, the user should use the `evaluate` function instead of directly calling it. `evaluate` will always return the Hamiltonian value in the unit system of $h=1$. The following code block shows the effects of different choices of `unit`:
H_h = DenseHamiltonian([(s)->1-s, (s)->s], -[σx, σz]/2, unit=:h)
H_ħ = DenseHamiltonian([(s)->1-s, (s)->s], -[σx, σz]/2, unit=:ħ)
println("Setting unit to :h")
@show evaluate(H_h, 0.5)
println("Setting unit to :ħ")
@show evaluate(H_ħ, 0.5);
# Internally, HOQST uses a unit system of $\hbar=1$. If we call `H_h` directly, its value is scaled by $2\pi$:
H_h(0.5)
# ### Annealing
# The total Hamiltonian presented in [Ref.](https://arxiv.org/abs/1503.08767) is
#
# $$H(s) = H_{\mathrm{S}}(s) + gS \otimes B + H_{\mathrm{B}} \ .$$
#
# We denote $S$ the coupling and $\{gB, H_{\mathrm{B}}\}$ the bath.
#
# #### Coupling
# For constant coupling operators, we can use the constructor `ConstantCouplings`. Like Hamiltonian's case, there is a keyword argument `unit` to specify the input unit.
coupling = ConstantCouplings(["Z"])
# #### Bath
# A bath instance can be any object which implements the following three methods:
# 1. Correlation function: `correlation(τ, bath)`
# 2. Spectrum density: `γ(ω, bath)`
# 3. Lamb shift: `S(ω, bath)`
#
# Redfield/Adiabatic ME solvers require those three methods. Currently, we have built-in support for the Ohmic bath. An Ohmic bath object can be created by :
η = 1e-4
fc = 4
T = 16
bath = Ohmic(η, fc, T)
# `info_freq` is a convenient function to convert each quantity into the same unit.
info_freq(bath)
# We can also directly plot the spectrum density of Ohmic bath:
p1 = plot(bath, :γ, range(0,20,length=200), label="", size=(800, 400), linewidth=2)
p2 = plot(bath, :S, range(0,20,length=200), label="", size=(800, 400), linewidth=2)
plot(p1, p2, layout=(1,2), left_margin=3Plots.Measures.mm)
# #### Annealing object
#
# Finally, we can assemble the annealing object by
# Hamiltonian
H = DenseHamiltonian([(s)->1-s, (s)->s], -[σx, σz]/2, unit=:ħ)
# initial state
u0 = PauliVec[1][1]
# coupling
coupling = ConstantCouplings(["Z"], unit=:ħ)
# bath
bath = Ohmic(1e-4, 4, 16)
annealing = Annealing(H, u0; coupling=coupling, bath=bath)
# Because we want to compare our results to the [Ref.](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.91.062320), we need to set the unit to $\hbar=1$.
#
# ### Closed system
# There are several interfaces in HOQST that might be handy. The first one is the Schrodinger equation solver:
tf = 10*sqrt(2)
@time sol = solve_schrodinger(annealing, tf, alg=Tsit5(), retol=1e-4)
# The following line of code is a convenient recipe to plot the instantaneous population during the evolution.
# It currently only supports Hamiltonian with annealing parameter s = t/tf from 0 to 1.
# The third argument can be either a list or a number. When it is a list, it specifies the energy levels to plot (starting from 0); when it is a number, it specifies the total number of levels to plot.
plot(sol, H, [0], 0:0.01:tf, linewidth=2, xlabel = "t (ns)", ylabel="\$P_G(t)\$")
# The solution is an `ODESolution` object in `DifferentialEquations.jl` package. More details for the interface can be found [here](http://docs.juliadiffeq.org/latest/basics/solution.html). The state vector's value at a given time can be obtained by directly calling the `ODESolution` object.
sol(0.5)
# Other interfaces include
# You need to solve the unitary first before trying to solve Redfield equation
@time U = solve_unitary(annealing, tf, alg=Tsit5(), abstol=1e-8, retol=1e-8);
@time solve_von_neumann(annealing, tf, alg=Tsit5(), abstol=1e-8, retol=1e-8);
# ### Open System
# #### Time-dependent Redfield equation
# The time-dependent Redfield equation solver needs
# 1. Annealing object
# 2. Total annealing time
# 3. Pre-calculated unitary
# to work. The following code block illustrates how to supply the above three objects to the Redfield solver. Besides, all the other keyword arguments in [DifferentialEquations.jl](https://diffeq.sciml.ai/stable/basics/common_solver_opts/) are supported.
tf = 10*sqrt(2)
U = solve_unitary(annealing, tf, alg=Tsit5(), abstol=1e-8, retol=1e-8);
sol = solve_redfield(annealing, tf, U; alg=Tsit5(), abstol=1e-8, retol=1e-8)
plot(sol, H, [0], 0:0.01:tf, linewidth=2, xlabel="t (ns)", ylabel="\$P_G(t)\$")
# #### Adiabatic master equation
# The adiabatic master equation solver needs
# 1. Annealing object
# 2. Total Annealing time
#
# Besides other keyword arguments supported in [DifferentialEquations.jl](https://diffeq.sciml.ai/stable/basics/common_solver_opts/), it is highly recommended to add the `ω_hint` keyword argument. By doing this, the solver will pre-compute the quantity $S(\omega)$ in Lambshift within the range specified by `ω_hint` to speed up the computation.
tf = 10*sqrt(2)
@time sol = solve_ame(annealing, tf; alg=Tsit5(), ω_hint=range(-6, 6, length=100), reltol=1e-4)
plot(sol, H, [0], 0:0.01:tf, linewidth=2, xlabel="t (ns)", ylabel="\$P_G(t)\$")
# We can also solve the AME for a longer annealing time:
tf = 5000
@time sol_ame = solve_ame(annealing, tf; alg=Tsit5(), ω_hint=range(-6, 6, length=100), reltol=1e-6)
plot(sol_ame, H, [0], 0:1:tf, linewidth=2, xlabel="t (ns)", ylabel="\$P_G(t)\$")
# The above results agree with Fig 2 of the [reference paper](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.91.062320).
#
# #### Trajectory method for adiabatic master equation
# The package also supports the trajectory method for AME. More details of this method can be found in this [paper](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.97.022116). The basic workflow is to create an ODE [EnsembleProblem](https://docs.juliadiffeq.org/dev/features/ensemble/) via `build_ensembles` interface. Then, the resulting `EnsembleProblem` object can be solved by the native [Parallel Ensemble Simulations](https://docs.juliadiffeq.org/dev/features/ensemble/) interface of `DifferentialEquations.jl`. The following code block solves the same annealing process described above($t_f = 5000(ns)$) using multithreading. To keep the running time reasonably short, we simulate only 3000 trajectories in this example. The result may not converge to the true solution yet. The user is encouraged to try more trajectories and see how the result converges.
#
# The codes can also be deployed on high-performance clusters using Julia's native [distributed computing](https://docs.julialang.org/en/v1/manual/distributed-computing/) module.
# +
tf = 5000
# total number of trajectories
num_trajectories = 3000
# construct the `EnsembleProblem`
# `safetycopy` needs to be true because the current trajectories implementation is not thread-safe.
prob = build_ensembles(annealing, tf, :ame, ω_hint=range(-6, 6, length=200), safetycopy=true)
# to use multi-threads, you need to start Julia kernel with multiple threads
# julia --threads 8
sol = solve(prob, Tsit5(), EnsembleThreads(), trajectories=num_trajectories, reltol=1e-6, saveat=range(0,tf,length=100))
t_axis = range(0,tf,length=100)
dataset = []
for t in t_axis
w, v = eigen_decomp(H, t/tf)
push!(dataset, [abs2(normalize(so(t))' * v[:, 1]) for so in sol])
end
# the following codes calculate the instantaneous ground state population and its error bar by averaging over all the trajectories
pop_mean = []
pop_sem = []
for data in dataset
p_mean = sum(data) / num_trajectories
p_sem = sqrt(sum((x)->(x-p_mean)^2, data)) / num_trajectories
push!(pop_mean, p_mean)
push!(pop_sem, p_sem)
end
scatter(t_axis, pop_mean, marker=:d, yerror=2*pop_sem, label="Trajectory", markersize=6)
plot!(sol_ame, H, [0], t_axis, linewidth=2, label="Non-trajectory")
xlabel!("t (ns)")
ylabel!("\$P_G(s)\$")
| notebook/introduction/02-single_qubit_ame.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome to the practicals of 'The Cryosphere in the Climate System'
# **Important: Check that you run the practicals in its own window (not as a tab) and do not forget to download your work!!!**
# Here are some notebooks with general information about the programming environment and an short introductions to notebooks and python:
#
# - [MyBinder and JupyterLab](MyBinder_and_JupyterLab.ipynb): an introduction to the programming environment
# - [Getting started with Jupyter Notebook](getting_started_with_notebooks.ipynb): an introduction to Jupyter Notebooks
# - [Getting started with Python](getting_started_with_python.ipynb): an introduction to Python
# Here you find the exercises in chronological order:
#
# - [E01 First Datacrunching](E01_First_Datacrunching.ipynb)
# - [E02 Available Tools](E02_Available_Tools.ipynb)
# - [E03 Meteorological Conditions](E03_Meteorological_Conditions.ipynb)
# - [E04 DegreeDayModel](E04_DegreeDayModel.ipynb)
# - [E05 SEB challenge](E05_SEB_challenge.ipynb)
#
# For the next exercises we use notebooks from OGGM Edu, to open the OGGM Edu-MyBinder environment you can go on the website [OGGM Edu](https://edu.oggm.org/en/latest/notebooks_howto.html#) and click on the 'launch Edu Notebooks' button or you can use the links below to direclty navigate to the notebooks (**do not forget to open the new MyBinder environment in a new tab**, and check out the other notebooks available there ;) ):
#
# - [Flowline Model](https://mybinder.org/v2/gh/OGGM/binder/master?urlpath=git-pull?repo=https://github.com/OGGM/oggm-edu-notebooks%26amp%3Bbranch=master%26amp%3Burlpath=lab/tree/oggm-edu-notebooks/oggm-edu/flowline_model.ipynb%3Fautodecode)
# - [Glacier Surging Experiment](https://mybinder.org/v2/gh/OGGM/binder/master?urlpath=git-pull?repo=https://github.com/OGGM/oggm-edu-notebooks%26amp%3Bbranch=master%26amp%3Burlpath=lab/tree/oggm-edu-notebooks/oggm-edu/surging_experiment.ipynb%3Fautodecode)
# - if the links do not navigate you directly to the notebooks click on *oggm-edu-notebooks* and *oggm-edu* in the left sidebar and choose the desired notebook
# Here you find completed notebooks:
#
# - [E01 First Datacrunching Completed](E01_First_Datacrunching_Completed.ipynb)
# - [E03 Meteorological Conditions Completed](E03_Meteorological_Conditions_Completed.ipynb)
# - [E04 DegreeDayModel Completed](E04_DegreeDayModel_completed.ipynb)
| exercises/welcome_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def sum1(n):
final_sum = 0
for x in range(n+1):
final_sum += x
return final_sum
sum1(10)
def sum2(n):
return (n * (n + 1))/2
sum2(10)
# %timeit sum1(100)
# %timeit sum2(100)
# +
from math import log
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('bmh')
#Setting up runtime comparisons
n = np.linspace(1, 10, 100)
labels = ['Constant', 'Logarithmic', 'Linear', 'Log Linear', 'Quadratic', 'Cubic', 'Exponential']
big_o = [np.ones(n.shape), np.log(n), n, n*np.log(n), n**2, n**3, 2**n]
#Plot Setup
plt.figure(figsize = (12,10))
plt.ylim(0, 50)
for i in range(len(big_o)):
plt.plot(n, big_o[i], label = labels[i])
plt.legend(loc = 0)
plt.ylabel('Relartive Runtime')
plt.xlabel('n')
# + active=""
# Big-O Examples
# +
#O(1) Constant
def func_constant(values):
print(values[0])
# -
lst = [1, 2, 3]
func_constant(lst)
# +
#O(n) Linear
def func_lin(lst):
for val in lst:
print(val)
# -
func_lin(lst)
# +
#O(n^2) Quadratic
def func_quad(lst):
for item_1 in lst:
for item_2 in lst:
print(item_1, item_2)
# -
lst = [1, 2, 3, 4, 5]
func_quad(lst)
def print_once(lst):
for val in lst:
print(val)
print_once(lst)
def print_2(lst):
for val in lst:
print(val)
for val in lst:
print(val)
print_2(lst)
def comp(lst):
print(lst[0]) #O(1)
##### O(n/2) . . . O( 1/2 * n)
midpoint = int(len(lst)/2)
for val in lst[:midpoint]:
print(val)
#####
for x in range(10): #O(10)
print('Hello World!')
comp(lst) #O(1 + n/2 + 10) or O(n)
def matcher(lst, match):
for item in lst:
if item == match:
return True
return False
matcher(lst, 1) #Best case is O(1)
matcher(lst, 11) #Worst case is O(n)
def create_list(n):
new_list = []
for num in range(n):
new_list.append('new')
return new_list
create_list(10)
def printer(n):
for x in range(10): #Time Complexity is O(n)
print('Hello World!') #Space Complexity is O(1)
printer(6)
| Python for Algorithms, Data Structures, & Interviews/Algorithm Analysis and Big O/Introduction to Algorithm Analysis and Big O.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(devtools)
devtools::install_github("ajijohn/microclimRapi")
library(microclimRapi)
api_token = getToken('0eda0dd88d1e6b417e8e9bebfc02cd95','<KEY>','http://microclim.org/')
print(api_token)
# API token received, now place the request for extraction
ma <- microclimRapi:::MicroclimAPI$new(token = api_token,url_mc='http://microclim.org/')
# +
# Eagle Nest Wilderness Area - Colorado
mr <- microclimRapi:::MicroclimRequest$new(
latS = "39.48178546986059",
latN="39.890772566959534",
lonW="-106.51519775390625",
lonE="-106.03317260742188",
variable="ALBEDO",
shadelevel=0,
hod=0,
interval=0,
aggregation=0,
stdate="19810101",
eddate="19810131",
file="csv")
# -
# Place a request to extract
ext_req= ma$request(mr)
print(ext_req)
print(ext_req$request_id)
ma$status(ext_req$request_id)
# If status is EMAILED, then files can be downloaded
ftch_req=''
#Pull the files
if(ma$status(ext_req$request_id) == "EMAILED")
{
# place a request to fetch the files
ftch_req= ma$fetch(ext_req$request_id)
}
ftch_req
ftch_req$files[[1]]$Key
ncD <- ma$download(ext_req$request_id,ftch_req$files[[1]]$Key)
#download the csv file
writeBin(ncD, strsplit(ftch_req$files[[1]]$Key, "/")[[1]][2])
file.exists(strsplit(ftch_req$files[[1]]$Key, "/")[[1]][2])
lv <- read.csv(strsplit(ftch_req$files[[1]]$Key, "/")[[1]][2],comment.char = "#")
str(lv)
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# name: python374jvsc74a57bd07945e9a82d7512fbf96246d9bbc29cd2f106c1a4a9cf54c9563dadf10f2237d4
# ---
# # Ejercicios Numpy I
# Sirvan estos ejercicios para que fortalezcáis aquellos conceptos más importantes.
# ## Ejercicio 1
# Importa el módulo de `numpy` y obtén su versión.
import numpy as np
np.__version__
# ## Ejercicio 2
# Crea un array de una dimensión que vaya de 0 a 9. Llamalo `my_array`
my_array=np.array(range(0,10))
my_array
# ## Ejercicio 3
# Crea un array 3x3, que sea todo a True
np.full((3, 3), True)
# ## Ejercicio 4
# Extrae todos los impares de `my_array`
for value in my_array:
for v in value:
if v%2!=0:
print(v)
# ## Ejercicio 5
# En un nuevo array, sustituye todos los impares de `my_array` por -1
for r,elem in enumerate(my_array):
if elem%2 == 1:
my_array[r]=-1
print(my_array)
# ## Ejercicio 6
# En un nuevo array, sustituye todos los impares de `my_array` por -1. Esta vez usa `where`
print(np.where(my_array %2==1, -1, my_array))
# ## Ejercicio 7
# Convierte `my_array` en un nuevo array de dos dimensiones mediante `reshape`.
my_array.reshape(5,2)
# ## Ejercicio 8
# Concatena los siguientes arrays verticamente
#
# ```Python
# a = np.arange(10).reshape(2,-1)
# b = np.repeat(1, 10).reshape(2,-1)
# ```
a = np.arange(10).reshape(2,-1)
b = np.repeat(1, 10).reshape(2,-1)
np.concatenate([a,b], axis=0)
# ## Ejercicio 9
# Concatena los siguientes arrays horizontamente
#
# ```Python
# a = np.arange(10).reshape(2,-1)
# b = np.repeat(1, 10).reshape(2,-1)
# ```
np.concatenate([a,b], axis=-1)
# ## Ejercicio 10
# Encuentra todos los elementos en común entre ambos arrays. [Para ello usa el método `intersect1d`](https://numpy.org/doc/stable/reference/generated/numpy.intersect1d.html)
#
# ```Python
# a = np.array([1,2,3,2,3,4,3,4,5,6])
# b = np.array([7,2,10,2,7,4,9,4,9,8])
# ```
a = np.array([1,2,3,2,3,4,3,4,5,6])
b = np.array([7,2,10,2,7,4,9,4,9,8])
np.intersect1d(a,b)
# ## Ejercicio 11
# Obtén la documentación acerca de la función `concatenate`
# +
# np.concatenate?
# -
# ## Ejercicio 12
# Crea un array unidimensional que vaya del 0 al 9. Llámalo `my_array`.
# 1. Multiplica cada uno de los elementos de `my_array` por 10, en un nuevo array
# 2. Accede al primer elemento del array
# 3. Accede al último elemento del array
# 4. Imprime por pantalla los numeros del 1 al 7
# +
my_array=np.array(range(0,10))
print(my_array)
my_array1=my_array*10
print(my_array1)
print(my_array[0])
print(my_array[-1])
print(my_array[0:8])
# -
# ## Ejercicio 13
# Crea un array manualmente de tres filas por tres columnas. Llámalo `my_matrix`
my_matrix=np.array([[1,2,3],[4,5,6],[7,8,9]])
my_matrix
# ## Ejercicio 14
# Obtén la siguiente secuencia:
#
# ```Python
# [ 1, 4, 7, 10, 13, 16, 19, 22, 25, 28, 31, 34, 37, 40, 43, 46, 49]
# ```
np.arange(1, 50, 3)
# ## Ejercicio 15
# Obtén la siguiente secuencia
#
# ```Python
# [4., 3.75, 3.5 , 3.25, 3. , 2.75, 2.5 , 2.25, 2., 1.75, 1.5 ,1.25]
# ```
np.arange(4, 1, -0.25)
# ## Ejercicio 16
# Lee la imagen `numpy.png` que encontrarás en la carpeta **img**
from skimage.io import imread
import matplotlib.pyplot as plt
imagen = imread("C:\\Users\\Administrator\\Desktop\\TheBridge\\Bootcamp-DataScience-2021\\week4_EDA_np_pd_json_apis_regex\\day1_numpy_pandas_I\\theory\\numpy\\img\\numpy.png") #no he podido sacar el relativo para hacerlo
plt.imshow(imagen)
# ## Ejercicio 17
# Crea un array de 4x1. A continuación convierte el array a un 2x2
array_estupida=np.full((4, 1),1)
np.reshape(array_estupida,(2,-1)) #el menos uno corta por la mitad
# ## Ejercicio 18
# Crea una secuencia aleatoria. La secuencia se tiene que componer de 4 matrices de 5x3. Serían 3 dimensiones diferentes.
# +
array1=np.random.randint(1,100, size=(5,3,4))
array1
# -
# ## Ejercicio 19
# Crea una matriz de 10x1, toda ella compuesta por `False`. Después transformalo en un 2x5
array_estupida=np.full((10, 1),False)
array_estupida
np.reshape(array_estupida,(2,5))
# ## Ejercicio 20
# Dado el siguiente array
#
# ```Python
# x =np.random.randint(10, 20, size = (5,2,6))
# ```
#
# 1. ¿Cuántos elementos tiene el array?
# 2. ¿Cuántas dimensiones?
# 3. ¿Cuántos elementos tiene cada dimensión?
x =np.random.randint(10, 20, size = (5,2,6))
print(x)
x.size #elementos
print(len(x)) #numero de matrices/
print(len(x[0])) #numero de columnas
#dimensiones ->
print(np.ndim(x)) #dimensiones
np.size(x[0]) #elementos de cada elemento
x = np.zeros((2,3,4))
x
| week4_EDA_np_pd_json_apis_regex/day2_numpy_pandas_II/exercises/Ejercicios Numpy I.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **[Fashion-MNIST](https://github.com/zalandoresearch/fashion-mnist)** is a dataset of Zalando's article images—consisting of a training set of 60,000 examples and a test set of 10,000 examples. Each example is a 28x28 grayscale image, associated with a label from 10 classes. The dataset serves as a direct drop-in replacement for the original [MNIST dataset](http://yann.lecun.com/exdb/mnist/) for benchmarking machine learning algorithms. It shares the same image size and structure of training and testing splits.
#
# In this work, I will use the pre-trained model **VGG19**, developed by <NAME> and <NAME> in 2014, a simple and widely used convnet architecture for ImageNet. The model is trained for 10 epochs with batch size of 256, compiled with `categorical_crossentropy` loss function and `Adam` optimizer.
#
# ## Data Processing
# Let's first load the training and test data from csv files.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from keras.utils import to_categorical
# Load training and test data into dataframes
data_train = pd.read_csv('data/fashion-mnist_train.csv')
data_test = pd.read_csv('data/fashion-mnist_test.csv')
# X forms the training images, and y forms the training labels
X = np.array(data_train.iloc[:, 1:])
y = to_categorical(np.array(data_train.iloc[:, 0]))
# X_test forms the test images, and y_test forms the test labels
X_test = np.array(data_test.iloc[:, 1:])
y_test = to_categorical(np.array(data_test.iloc[:, 0]))
# -
# The images of fashionMNIST are black and white, while the required input for VGG19 must be colored images. Thus, I convert the images into colored ones with 3 channels R, G, B.
# +
# Convert the training and test images into 3 channels
X = np.dstack([X] * 3)
X_test = np.dstack([X_test]*3)
# Display their new shapes
X.shape, X_test.shape
# -
# Let's reshape the images into tensor format as required by TensorFlow.
# +
# Reshape images as per the tensor format required by tensorflow
X = X.reshape(-1, 28,28,3)
X_test= X_test.reshape (-1,28,28,3)
# Display the new shape
X.shape, X_test.shape
# -
# VGG19 requires minimum input image's width and height of 48, but I'll resize my images from 28 x 28 to 150 x 150.
# +
# Resize the images as 150 * 150 as required by VGG19
from keras.preprocessing.image import img_to_array, array_to_img
X = np.asarray([img_to_array(array_to_img(im, scale=False).resize((150,150))) for im in X])
X_test = np.asarray([img_to_array(array_to_img(im, scale=False).resize((150,150))) for im in X_test])
# Display the new shape
X.shape, X_test.shape
# -
# Here, I preprocess the images by reshaping them into the shape the network expects and scaling them so that all values are in the [0, 1] interval. For instance, the training images were stored in an array of shape (60000, 150, 150, 3) of type uint8 with values in the [0, 255] interval. I transform them into a float32 array of shape (60000, 150, 150, 3) with values between 0 and 1.
# +
# Normalise the data and change data type
X = X.astype('float32')
X /= 255
X_test = X_test.astype('float32')
X_test /= 255
# -
# Here, I split the original training data (60,000 images) into 80% training (48,000 images) and 20% validation (12000 images) optimize the classifier, while keeping the test data (10,000 images) to finally evaluate the accuracy of the model on the data it has never seen. This helps to see whether I'm over-fitting on the training data and whether I should lower the learning rate and train for more epochs if validation accuracy is higher than training accuracy or stop over-training if training accuracy shift higher than the validation.
# +
from sklearn.model_selection import train_test_split
# Here I split original training data to sub-training (80%) and validation data (20%)
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=13)
# Check the data size whether it is as per tensorflow and VGG19 requirement
X_train.shape, X_val.shape, y_train.shape, y_val.shape
# -
# ## VGG19
# A common and highly effective approach to deep learning on small image datasets is to use a pretrained network. A **pretrained network** is a saved network that was previously trained on a large dataset, typically on a large-scale image-classification task. If this original dataset is large enough and general enough, then the spatial hierarchy of features learned by the pretrained network can effectively act as a generic model of the visual world, and hence its features can prove useful for many different computer-vision problems, even though these new problems may involve completely different classes than those of the original task.
#
# I'll use the [VGG19 architecture](https://arxiv.org/abs/1409.1556), developed by <NAME> and <NAME> in 2014. It’s a simple and widely used convnet architecture for ImageNet. The model comes pre-packaged with Keras. Let's instantiate it:
# +
from keras.applications import VGG19
# Create the base model of VGG19
vgg19 = VGG19(weights='imagenet', include_top=False, input_shape = (150, 150, 3), classes = 10)
# -
# I passed 4 arguments to the constructor:
# * `weights` specifies the weight checkpoint from which to initialize the model.
# * `include_top` refers to including (or not) the densely connected classifier on top of the network. By default, this densely connected classifier corresponds to the 1,000 classes from ImageNet. Because I intend to use my own densely connected classifier (with only 10 classes), I don’t need to include it.
# * `input_shape`: optional shape tuple, only to be specified if `include_top` is False.
# * `classes`: optional number of classes to classify images into, only to be specified if `include_top` is True, and if no `weights` argument is specified.
#
# Here's the detail of the architecture of the VGG19 convolutional base.
vgg19.summary()
# ## Feature Extraction
# Feature extraction consists of using the representations learned by a previous network to extract interesting features from new samples. These features are then run through a new classifier, which is trained from scratch.
#
# CNNs used for image classification comprise two parts: they start with a series of pooling and convolution layers, and they end with a densely-connected classifier. The first part is called the **"convolutional base"** of the model. In the case of convnets, **"feature extraction"** will simply consist of taking the convolutional base of a previously-trained network, running the new data through it, and training a new classifier on top of the output.
#
# 
#
# Why only reuse the convolutional base? Could we reuse the densely-connected classifier as well? In general, it should be avoided. The reason is simply that the representations learned by the convolutional base are likely to be more generic and therefore more reusable: the feature maps of a convnet are presence maps of generic concepts over a picture, which is likely to be useful regardless of the computer vision problem at hand. On the other end, the representations learned by the classifier will necessarily be very specific to the set of classes that the model was trained on -- they will only contain information about the presence probability of this or that class in the entire picture. Additionally, representations found in densely-connected layers no longer contain any information about where objects are located in the input image: these layers get rid of the notion of space, whereas the object location is still described by convolutional feature maps. For problems where object location matters, densely-connected features would be largely useless.
#
# Looking back at the VGG 19 architecture, the final feature map has shape `(4, 4, 512)`. That's the feature on top of which I will stick a densely-connected classifier. First, let's pre-process the data so that it's trainable using VGG19.
# +
from keras.applications.vgg19 import preprocess_input
# Preprocessing the input
X_train = preprocess_input(X_train)
X_val = preprocess_input(X_val)
X_test = preprocess_input(X_test)
# -
# Now, in order to extract features from Fashion-MNIST, I will:
# * Run the convolutional base over the dataset.
# * Record its output to a Numpy array on disk.
# * Use this data as input to a standalone densely-connected classifier.
#
# This solution is very fast and cheap to run, because it only requires running the convolutional base once for every input image, and the convolutional base is by far the most expensive part of the pipeline. However, for the exact same reason, this technique would not allow me to leverage data augmentation at all.
#
# Now I will extract features from the train, test, and validation data simply by calling the `predict` method of the `vgg19` model.
# Extracting features
train_features = vgg19.predict(np.array(X_train), batch_size=256, verbose=1)
test_features = vgg19.predict(np.array(X_test), batch_size=256, verbose=1)
val_features = vgg19.predict(np.array(X_val), batch_size=256, verbose=1)
# Saving the features so that they can be used for future
np.savez("train_features", train_features, y_train)
np.savez("test_features", test_features, y_test)
np.savez("val_features", val_features, y_val)
# Current shape of features
print(train_features.shape, "\n", test_features.shape, "\n", val_features.shape)
# The extracted features are currently of shape `(samples, 4, 4, 512)`. I will feed them to a densely-connected classifier, so first I must flatten them to `(samples, 8192)`:
# Flatten extracted features
train_features = np.reshape(train_features, (48000, 4*4*512))
test_features = np.reshape(test_features, (10000, 4*4*512))
val_features = np.reshape(val_features, (12000, 4*4*512))
# At this point, I can define the densely-connected classifier (note the use of dropout for regularization), and train it on the data and labels that I just recorded:
# +
from keras.layers import Dense, Dropout
from keras.models import Model
from keras import models
from keras import layers
from keras import optimizers
# Add Dense and Dropout layers on top of VGG19 pre-trained
model = models.Sequential()
model.add(layers.Dense(512, activation='relu', input_dim=4 * 4 * 512))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(10, activation="softmax"))
# -
# When compiling the model, I choose **categorical_crossentropy** as the loss function (which is relevent for multiclass, single-label classification problem) and **Adam** optimizer.
# * The cross-entropy loss calculates the error rate between the predicted value and the original value. The formula for calculating cross-entropy loss is given [here](https://en.wikipedia.org/wiki/Cross_entropy). Categorical is used because there are 10 classes to predict from. If there were 2 classes, I would have used binary_crossentropy.
# * The Adam optimizer is an improvement over SGD(Stochastic Gradient Descent). The optimizer is responsible for updating the weights of the neurons via backpropagation. It calculates the derivative of the loss function with respect to each weight and subtracts it from the weight. That is how a neural network learns.
# +
import keras
# Compile the model
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
# -
# ## Training the Model
# As previously mentioned, I train the model with batch size of 256 and 10 epochs on both training and validation data.
# Train the the model
history = model.fit(train_features, y_train,
batch_size=256,
epochs=50,
verbose=1,
validation_data=(val_features, y_val))
score = model.evaluate(test_features, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# ## Results
# Let's plot training and validation accuracy as well as training and validation loss.
# +
# plot the loss and accuracy
import matplotlib.pyplot as plt
# %matplotlib inline
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.title('Training and validation accuracy')
plt.plot(epochs, acc, 'red', label='Training acc')
plt.plot(epochs, val_acc, 'blue', label='Validation acc')
plt.legend()
plt.figure()
plt.title('Training and validation loss')
plt.plot(epochs, loss, 'red', label='Training loss')
plt.plot(epochs, val_loss, 'blue', label='Validation loss')
plt.legend()
plt.show()
# -
# ## Classification Report
# I can summarize the performance of my classifier as follows:
# +
# get the predictions for the test data
predicted_classes = model.predict_classes(test_features)
# get the indices to be plotted
y_true = data_test.iloc[:, 0]
correct = np.nonzero(predicted_classes==y_true)[0]
incorrect = np.nonzero(predicted_classes!=y_true)[0]
# -
from sklearn.metrics import classification_report
target_names = ["Class {}".format(i) for i in range(10)]
print(classification_report(y_true, predicted_classes, target_names=target_names))
test_features[correct].shape
| .ipynb_checkpoints/VGG19-GPU-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.1
# language: julia
# name: julia-1.3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Sequence classification model for IMDB Sentiment Analysis
# (c) <NAME>, 2019
# * Objectives: Learn the structure of the IMDB dataset and train a simple RNN model.
# * Prerequisites: [RNN models](60.rnn.ipynb)
# -
# Set display width, load packages, import symbols
ENV["COLUMNS"] = 72
using Pkg; for p in ("Knet","IterTools"); haskey(Pkg.installed(),p) || Pkg.add(p); end
using Statistics: mean
using IterTools: ncycle
using Knet: Knet, AutoGrad, RNN, param, dropout, minibatch, nll, accuracy, progress!, adam, save, load, gc
# + slideshow={"slide_type": "slide"}
# Set constants for the model and training
EPOCHS=3 # Number of training epochs
BATCHSIZE=64 # Number of instances in a minibatch
EMBEDSIZE=125 # Word embedding size
NUMHIDDEN=100 # Hidden layer size
MAXLEN=150 # maximum size of the word sequence, pad shorter sequences, truncate longer ones
VOCABSIZE=30000 # maximum vocabulary size, keep the most frequent 30K, map the rest to UNK token
NUMCLASS=2 # number of output classes
DROPOUT=0.5 # Dropout rate
LR=0.001 # Learning rate
BETA_1=0.9 # Adam optimization parameter
BETA_2=0.999 # Adam optimization parameter
EPS=1e-08 # Adam optimization parameter
# -
# ## Load and view data
# + slideshow={"slide_type": "fragment"}
include(Knet.dir("data","imdb.jl")) # defines imdb loader
# + slideshow={"slide_type": "skip"}
@doc imdb
# + slideshow={"slide_type": "slide"}
@time (xtrn,ytrn,xtst,ytst,imdbdict)=imdb(maxlen=MAXLEN,maxval=VOCABSIZE);
# + slideshow={"slide_type": "fragment"}
println.(summary.((xtrn,ytrn,xtst,ytst,imdbdict)));
# + slideshow={"slide_type": "slide"}
# Words are encoded with integers
rand(xtrn)'
# + slideshow={"slide_type": "slide"}
# Each word sequence is padded or truncated to length 150
length.(xtrn)'
# + slideshow={"slide_type": "slide"}
# Define a function that can print the actual words:
imdbvocab = Array{String}(undef,length(imdbdict))
for (k,v) in imdbdict; imdbvocab[v]=k; end
imdbvocab[VOCABSIZE-2:VOCABSIZE] = ["<unk>","<s>","<pad>"]
function reviewstring(x,y=0)
x = x[x.!=VOCABSIZE] # remove pads
"""$(("Sample","Negative","Positive")[y+1]) review:\n$(join(imdbvocab[x]," "))"""
end
# -
# Hit Ctrl-Enter to see random reviews:
r = rand(1:length(xtrn))
println(reviewstring(xtrn[r],ytrn[r]))
# + slideshow={"slide_type": "fragment"}
# Here are the labels: 1=negative, 2=positive
ytrn'
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Define the model
# -
struct SequenceClassifier; input; rnn; output; pdrop; end
SequenceClassifier(input::Int, embed::Int, hidden::Int, output::Int; pdrop=0) =
SequenceClassifier(param(embed,input), RNN(embed,hidden,rnnType=:gru), param(output,hidden), pdrop)
# +
function (sc::SequenceClassifier)(input)
embed = sc.input[:, permutedims(hcat(input...))]
embed = dropout(embed,sc.pdrop)
hidden = sc.rnn(embed)
hidden = dropout(hidden,sc.pdrop)
return sc.output * hidden[:,:,end]
end
(sc::SequenceClassifier)(input,output) = nll(sc(input),output)
# -
# ## Experiment
dtrn = minibatch(xtrn,ytrn,BATCHSIZE;shuffle=true)
dtst = minibatch(xtst,ytst,BATCHSIZE)
length.((dtrn,dtst))
# For running experiments
function trainresults(file,maker; o...)
if (print("Train from scratch? "); readline()[1]=='y')
model = maker()
progress!(adam(model,ncycle(dtrn,EPOCHS);lr=LR,beta1=BETA_1,beta2=BETA_2,eps=EPS))
Knet.save(file,"model",model)
Knet.gc() # To save gpu memory
else
isfile(file) || download("http://people.csail.mit.edu/deniz/models/tutorial/$file",file)
model = Knet.load(file,"model")
end
return model
end
maker() = SequenceClassifier(VOCABSIZE,EMBEDSIZE,NUMHIDDEN,NUMCLASS,pdrop=DROPOUT)
# model = maker()
# nll(model,dtrn), nll(model,dtst), accuracy(model,dtrn), accuracy(model,dtst)
# (0.69312066f0, 0.69312423f0, 0.5135817307692307, 0.5096153846153846)
model = trainresults("imdbmodel132.jld2",maker);
# ┣████████████████████┫ [100.00%, 1170/1170, 00:15/00:15, 76.09i/s]
# nll(model,dtrn), nll(model,dtst), accuracy(model,dtrn), accuracy(model,dtst)
# (0.05217469f0, 0.3827392f0, 0.9865785256410257, 0.8576121794871795)
# ## Playground
predictstring(x)="\nPrediction: " * ("Negative","Positive")[argmax(Array(vec(model([x]))))]
UNK = VOCABSIZE-2
str2ids(s::String)=[(i=get(imdbdict,w,UNK); i>=UNK ? UNK : i) for w in split(lowercase(s))]
# Here we can see predictions for random reviews from the test set; hit Ctrl-Enter to sample:
r = rand(1:length(xtst))
println(reviewstring(xtst[r],ytst[r]))
println(predictstring(xtst[r]))
# Here the user can enter their own reviews and classify them:
println(predictstring(str2ids(readline(stdin))))
| tutorial/70.imdb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 필요한 라이브러리 불러오기
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# ### Kaggle Survey 데이터 불러오기
data = pd.read_csv("data/kaggle_survey_2020_responses.csv")
data
# ### 데이터 전처리
#
# - 교육상태와 관련이 있는 column들을 고릅니다.
#
# - Data Science 실무 경력과 관련된 column들을 고릅니다.
# +
# edu_column
edu_columns = ["Q4","Q6","Q15"]
# ds_column
ds_columns = ["Q5","Q20","Q22"]
y = ["Q24"]
edit_data = data[edu_columns + ds_columns+y]
edit_data
# -
# null이 아닌 row만
edit_data[edit_data.isnull().any(axis=1)] #null 인 row 확인
# 결측치 제거
final_data = edit_data.dropna()
final_data
# ## 교육수준과 관련된 EDA
#
# - EDA에서는 다음과 같은 항목을 살펴봅니다.
#
# > 기본적인 빈도 분석
#
# > 히스토그램
#
# > Pie chart와 같은 plotting 기법들
# ### Q4 column
#
# **"What is the highest level of formal education that you have attained or plan to attain with the next 2 years?"**
#
# - 기본적인 matplotlib(pyplot)을 이용하여 시각화를 해봅니다.
# Q4 = final_data["Q4"] # 첫번째 row는 질문 -> 지우자
Q4 = final_data["Q4"][1:]
Q4
plt.figure(figsize=(8, 8))
plt.title("Histogram of Q4 column")
plt.hist(Q4)
plt.xticks(rotation=90)
plt.show()
plt.figure(figsize=(10, 10))
plt.pie(Q4.value_counts(),
labels=Q4.value_counts().index,
autopct='%d%%',
startangle=90,
textprops={'fontsize':20})
plt.axis('equal')
plt.title("Pie chart for Q4 column", fontsize=20)
# ### Q6 column
# **For how many years have you been writing code and/or programming?**
Q6 = final_data["Q6"][1:]
Q6
Q6.value_counts()
plt.figure(figsize=(8, 8))
plt.hist(Q6)
plt.xticks(rotation='vertical')
plt.title("Histogram for Q6 column")
plt.show()
plt.figure(figsize=(8, 8))
plt.pie(Q6.value_counts(),
labels=Q6.value_counts().index,
autopct='%d%%',
textprops={'fontsize':24})
plt.axis('equal')
plt.title("Pie chart for Q6 column", fontsize=48)
# ### Q15 column
#
# **For how many years have you used machine learning methods?**
#
# - seaborn을 이용해서 시각화를 해봅니다.
Q15 = final_data["Q15"][1:]
Q15
# countplot을 사용해봅니다
sns.countplot(y="Q15", data=final_data[1:])
plt.show()
plt.figure(figsize=(8,8))
plt.pie(Q15.value_counts(),
labels=Q15.value_counts().index,
autopct='%d%%',
colors=sns.color_palette('hls',len(Q15.value_counts().index)),
textprops={'fontsize':12})
plt.axis('equal')
plt.title("Pie chart for Q15 column", fontsize=16, pad=50)
plt.show()
# ## 직업과 관련된 EDA
#
# - EDA에서는 다음과 같은 항목을 살펴봅니다.
#
# > 기본적인 빈도 분석
#
# > 히스토그램
#
# > Pie chart와 같은 plotting 기법들
# ### Q5 column
#
# **Select the title most similar to your current role**
#
# - 기본적인 matplotlib(pyplot)을 이용하여 시각화를 해봅니다.
Q5 = final_data["Q5"][1:]
Q5
Q5.value_counts()
plt.figure(figsize=(8,8))
# plt.hist(Q5)
plt.barh(Q5.value_counts().index, Q5.value_counts().values)
# plt.bar(Q5.value_counts().index, Q5.value_counts().values)
plt.xticks(rotation='vertical')
plt.title("Histogram for Q5 column")
plt.show()
plt.figure(figsize=(8, 8))
plt.pie(Q5.value_counts(),
labels=Q5.value_counts().index,
autopct='%d%%',
textprops={'fontsize':24})
plt.axis('equal')
plt.title("Pie chart for Q5 column", fontsize=48, pad=50)
plt.show()
# ### Q20 column
#
# **What is the size of the compnay where are employed?**
# - seaborn을 이용해서 시각화를 해봅니다.
Q20 = final_data["Q20"][1:]
Q20
Q20.value_counts()
# sns.countplot을 사용해봅시다.
sns.countplot(y="Q20", data=final_data[1:])
plt.show()
plt.figure(figsize=(8,8))
plt.pie(Q20.value_counts(),
labels=Q20.value_counts().index,
autopct='%d%%',
colors=sns.color_palette('hls',len(Q20.value_counts().index)),
textprops={'fontsize':16})
plt.axis('equal')
plt.title("Pie chart for Q20 column", fontsize=32, pad=50)
plt.show()
# ### Q22 column
#
# **Does your current employer incorporate machine learning methods into their business?**
# - seaborn을 이용해서 시각화를 해봅니다.
Q22 = final_data["Q22"][1:]
Q22.value_counts()
plt.figure(figsize=(4, 4))
# To-Do
# plt.bar(Q22.value_counts().index, Q22.value_counts().values)
plt.barh(Q22.value_counts().index, Q22.value_counts().values)
# plt.xticks(rotation=90)
plt.show()
plt.figure(figsize=(8, 8))
plt.pie(Q22.value_counts(),
labels=Q22.value_counts().index,
autopct='%.2f%%',
colors=sns.color_palette('hls',len(Q22.value_counts().index)),
textprops={'fontsize':12})
plt.axis('equal')
plt.title("Pie chart for Q22 column", fontsize=32, pad=50)
plt.show()
final_data
set(data["Q3"]) # --> Republic of Korea , South Korea
skorea = data[data["Q3"].isin(["Republic of Korea" , "South Korea"])]
skorea.head()
sQ4 = skorea["Q4"]
sQ4.value_counts()
sns.countplot(y="Q4", data=skorea[1:])
plt.show()
plt.figure(figsize=(8, 8))
plt.pie(sQ4.value_counts(),
labels=sQ4.value_counts().index,
autopct='%.2f%%',
colors=sns.color_palette('hls',len(sQ4.value_counts().index)),
textprops={'fontsize':12})
plt.axis('equal')
plt.title("Pie chart for sQ4 column in S_Korea", fontsize=32, pad=50)
plt.show()
| python/210915-Python-practice-kaggle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple Linear Regression
# pandas provides excellent data reading and querying module, dataframe which allows you to import data like SQL- queries
# Below is imported house price records from Trulia.
import pandas
df = pandas.read_excel('house_price.xlsx')
df[:10]
# # Prepare the data
# We want to use the price as the dependent variable and the area as the independent variable, i.e., use the house areas to predict the house prices
X = df['area']
print (X[:10])
X_reshape = X.values.reshape(-1,1) # reshape the X to a 2D array
print (X_reshape[:10])
y = df['price']
# sklearn provides a split function that can split the data into training data and testing data.
# +
import sklearn
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_reshape,y, test_size = 0.3) # put 30% data as the testing data
print ('number of training data:',len(X_train),len(y_train))
print ('number of testing data:',len(X_test),len(y_test))
# -
# # Train the model
# use the Linear Regression to estimate parameters from the training data.
# +
from sklearn import linear_model
slr = linear_model.LinearRegression() #create an linear regression model objective
slr.fit(X_train,y_train) # estimate the patameters
print('beta',slr.coef_)
print('alpha',slr.intercept_)
# -
# # Evaluate the Model
# Let's calculate the mean squared error and the r square of the model
# +
from sklearn.metrics import mean_squared_error, r2_score
y_predict = slr.predict(X_test) # predict the Y based on the model
mean_squared_error = mean_squared_error(y_test,y_predict) # calculate mean square error
r2_score = r2_score(y_test,y_predict) #calculate r square
print ('mean square error:',mean_squared_error )
print ('r square:',r2_score )
# -
# # Visualize the model
# using matplotlib to visualize the data
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.scatter(X_test, y_test, color='black') # create a scatterplot to visualize the test data
plt.plot(X_test, y_predict, color='blue', linewidth=3) # add a line chart to visualize the model
plt.xlabel('area')
plt.ylabel('price')
plt.show()
# -
| Lab3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Giới thiệu phương pháp học Leitner
# > drafting
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [tutorial]
# - image: images/leitner_box.jpg
# - hide: true
# # Giới thiệu
# Những thứ không hiệu quả cho việc học như: bài giảng, nhồi nhét và đọc lại.
#
# Phương pháp này dựa trên việc học tập một kiến thức mới lặp đi lặp lại, chúng ta kiểm tra trí nhớ của mình về một kiến thức nào đó lặp đi lặp lại và trong những khoảng thời gian khác nhau. Phương pháp này không phải là một study trick hay life hack mà là một cách để điều khiển bộ não của mình, giúp não bộ nhớ lâu hơn.
# Phương pháp này được phát minh bở nhà tâm lý học người <NAME>. Ông đã tiến hành một thí nghiệm, theo dõi khả năng nhớ hàng nghìn từ vựng vô nghĩa của mình và ghi chép việc quên của mình. Ông khám phá ra rằng ông đã quên hầu hết mọi thứ đã học trong vòng 24h đầu tiên và những thứ còn xót lại tiếp tục bị quên dần những ngày sau đó. Tuy nhiên, nhìn chung tốc độ quên những thứ đã học giảm dần khi chúng ta được chủ động xem lại kiến thức đó (không phải bị động) - mặc dù khi cyahúng ta dừng thực hành, bộ nhớ tiếp tục giảm. Do vậy, để học bất cứ thứ gì, bạn cần phải xem lại nó ngay lúc bộ não bắt đầu quên và thời gian giữa các lần review phải tăng dần.
#
# [forgeting_curve](images.forgetting_curve.png)
#
# Điều thú vị trong cách học này là bạn không phải học 1 thứ lặp đi lặp lại mà bạn chỉ học thứ mới hoặc thứ bạn cứ quên quài.
# # Hộp Leitner
# Đầu tiên, chúng ta chia hộp thành 7 cấp độ, bạn có thể có nhiều hoặc ít hơn nếu thích.
# Một card mới sẽ được bỏ vào hộp thứ 1, khuyến khich bắt đầu với 5 card mơi mỗi ngày.
# Khi bạn review một card, nếu bạn nhớ đúng kiến thức trong đó, chuyển card đó up một level. Nếu card của bạn ở level cuối cùng, chúc mừng bạn, bây giờ bạn có thể vứt chiệc card đó đi, kiến thức bên trong chiếc card đó sẽ theo bạn suốt đời.
# Tuy nhiên nếu bạn quên một card nào đó, bạn phải chuyển nó về hộp ban đầu, hộp đầu tiên.
# Thời gian review hộp leitner:
# - level 1: hằng ngày
# - level 2: 2 ngày một lần
# - level 3: 4 ngày một lần
# ... cứ như vậy, mỗi lần lên một bậc thì gấp đôi thời gian review
# [](schedule)
# Mỗi lần review, chúng ta sẽ review level từ trên xuống, như vậy level 1 sẽ được review cuối cùng. Như vậy chúng ta sẽ biết được những card mình bị quên và những card mới bỏ vào.
# Hằng ngày, cố gắng không để sót lại bất kì card nào ở level 1, học đi học lại đến khi nào mình nhớ nó và move nó lên level 2.
#
# review first => add new card later
#
# mỗi ngày chúng ta cố dành khoảng 20-30 phút để học thay vì xem tivi - bạn có thể nhớ mọi thứ trên đời. Tuy nhiên để xây dựng được thới quen mới khá khó, nếu bạn khởi đầu lớn, có thể bạn sẽ kết thúc nó ngay ngày hôm sau. Nếu bạn khởi đầu nhỏ và lấy cảm hứng dần dần, bạn có thể học nhiều hơn mỗi ngày, vì vậy mà mình khuyến khích học 5 card mỗi ngày.
#
# Sau khi chúng ta đã quen với cách học này, chúng ta có thể có 10,15,20,25,30 card/ngày.
# Nếu bạn học 30 card mỗi ngày, một năm bạn sẽ học được 10.000+ điều mới
# # Những việc có thể dẫn tới học sai
# - Việc học sẽ fail nếu những card của bạn cồng kềnh, không liên quan tới nhau và không có nghĩa. Mặc khác nếu những card của bạn là những mảng nhỏ, kết nôi vơi nhau thì sẽ tốt hơn. Đó cũng là cách nao bộ hoạt động, lots of small and connected things. Vấn đề không phải ở chỗ collection mà ở chỗ connection.
#
# => card của bạn phải nhỏ, kết nối, có ý nghĩa.
# - Nhỏ: quá nhiều thông tin trên một card => hãy cắt nhỏ nó ra thành nhiều card, smaller and connected pieces. => rule of thumb: mỗi card chỉ nếu có một và chỉ một idea.
# - connected: nếu bạn vẽ hình, ghi hoàn cảnh hoặc thông tin cá nhân lên card, sẽ gợi nhớ tốt hơn có card.
# - ý nghĩa: hãy chọn một topic nào đó mà bạn đang theo đuổi, học piano, đọc truyện, chơi game và bắt đầu dùng leitner box để học mọi thứ về bộ môn đó. Mình tin rằng cách tốt nhất để giữ motivation cho việc học là mình đọc học thứ gì đó mình quan tâm
#
# - không quan trọng bạn học buổi sáng hay chiều, quan trọng là hằng ngày đều phải học (đôi khi có thể skip 1 ngày)
# - you cheat: correct answer, time,
# - you dont shuffle
# - you dont cart your victories
# - you dont educate yourself
# - your flashcard are wrong
# +
https://www.youtube.com/watch?v=HN0OUnLxFeU&list=PLdddsM1tHEe-I7QpcFmoNsmwP3dWH_3s_
https://www.youtube.com/watch?v=hs5qmTKBSU0&list=PLdddsM1tHEe-I7QpcFmoNsmwP3dWH_3s_&index=3
https://www.youtube.com/watch?v=ad1nHS-3stg
https://ncase.me/remember/
https://en.wikipedia.org/wiki/Leitner_system
https://www.ankiapp.com/
https://tinycards.duolingo.com/
# -
| _notebooks/draft/phuong-phap-hoc-tap-leitner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.147462, "end_time": "2021-10-23T12:27:11.408781", "exception": false, "start_time": "2021-10-23T12:27:11.261319", "status": "completed"} tags=[]
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + [markdown] papermill={"duration": 0.059106, "end_time": "2021-10-23T12:27:11.529726", "exception": false, "start_time": "2021-10-23T12:27:11.47062", "status": "completed"} tags=[]
# ### 1.1. Install Packages
# + papermill={"duration": 153.059377, "end_time": "2021-10-23T12:29:44.649056", "exception": false, "start_time": "2021-10-23T12:27:11.589679", "status": "completed"} tags=[]
# ! python -m pip install tf-models-nightly --no-deps -q
# ! python -m pip install tf-models-official==2.4.0 -q
# ! python -m pip install tensorflow-gpu==2.4.1 -q
# ! python -m pip install tensorflow-text==2.4.1 -q
# ! python -m spacy download en_core_web_sm -q
# ! python -m spacy validate
# + [markdown] papermill={"duration": 0.062903, "end_time": "2021-10-23T12:29:44.774993", "exception": false, "start_time": "2021-10-23T12:29:44.71209", "status": "completed"} tags=[]
# ### 1.2. Import Libraries
# + papermill={"duration": 3.919414, "end_time": "2021-10-23T12:29:48.756362", "exception": false, "start_time": "2021-10-23T12:29:44.836948", "status": "completed"} tags=[]
# Preprocessing
import spacy
from spacy.lang.en.stop_words import STOP_WORDS
from spacy.lang.en import English
from sklearn.model_selection import train_test_split
from sklearn.metrics.pairwise import cosine_similarity
import re
import string
from bs4 import BeautifulSoup as bs
# Model Training
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_text as text
from tensorflow.keras import layers, Model
from tensorflow.keras import regularizers
from tensorflow.keras.metrics import BinaryAccuracy
from tensorflow.keras.losses import BinaryCrossentropy
import official.nlp.optimization
from official.nlp.optimization import create_optimizer # AdamW optimizer
from sklearn.metrics import roc_curve, confusion_matrix
# Visualization
import seaborn as sns
from wordcloud import WordCloud
import matplotlib.pyplot as plt
from matplotlib import rcParams
# Version
from platform import python_version
print(f'TensorFlow Version: {tf.__version__}')
print(f'Python Version: {python_version()}')
# + [markdown] papermill={"duration": 0.087101, "end_time": "2021-10-23T12:29:48.954699", "exception": false, "start_time": "2021-10-23T12:29:48.867598", "status": "completed"} tags=[]
# ### 1.3. Configure Settings
# + papermill={"duration": 1.250732, "end_time": "2021-10-23T12:29:50.2683", "exception": false, "start_time": "2021-10-23T12:29:49.017568", "status": "completed"} tags=[]
RANDOM_SEED = 123
nlp = spacy.load('en_core_web_sm')
pd.set_option('display.max_colwidth', None) # Expand DataFrame column width
rcParams['figure.figsize'] = (10, 6) # Custom plot dimensions
sns.set_theme(palette='muted', style='whitegrid') # Seaborn plot theme
# + [markdown] papermill={"duration": 0.062644, "end_time": "2021-10-23T12:29:50.39735", "exception": false, "start_time": "2021-10-23T12:29:50.334706", "status": "completed"} tags=[]
# ### 1.4. Load the Training and Test Files
# + papermill={"duration": 0.134389, "end_time": "2021-10-23T12:29:50.593755", "exception": false, "start_time": "2021-10-23T12:29:50.459366", "status": "completed"} tags=[]
path = '../input/nlp-getting-started/train.csv'
df = pd.read_csv(path)
print(df.shape)
df.head()
# + papermill={"duration": 0.159885, "end_time": "2021-10-23T12:29:50.866055", "exception": false, "start_time": "2021-10-23T12:29:50.70617", "status": "completed"} tags=[]
path_test = '../input/nlp-getting-started/test.csv'
df_test = pd.read_csv(path_test)
print(df_test.shape)
df_test.head()
# + [markdown] papermill={"duration": 0.118222, "end_time": "2021-10-23T12:29:51.110677", "exception": false, "start_time": "2021-10-23T12:29:50.992455", "status": "completed"} tags=[]
# # 2. Raw Data Analysis
# + papermill={"duration": 0.082839, "end_time": "2021-10-23T12:29:51.280174", "exception": false, "start_time": "2021-10-23T12:29:51.197335", "status": "completed"} tags=[]
df.info(verbose=False)
# + papermill={"duration": 0.088896, "end_time": "2021-10-23T12:29:51.45156", "exception": false, "start_time": "2021-10-23T12:29:51.362664", "status": "completed"} tags=[]
df_test.info(verbose=False)
# + papermill={"duration": 0.100543, "end_time": "2021-10-23T12:29:51.637181", "exception": false, "start_time": "2021-10-23T12:29:51.536638", "status": "completed"} tags=[]
df['text'].describe()
# + papermill={"duration": 0.081865, "end_time": "2021-10-23T12:29:51.786424", "exception": false, "start_time": "2021-10-23T12:29:51.704559", "status": "completed"} tags=[]
df_test['text'].describe()
# + [markdown] papermill={"duration": 0.06496, "end_time": "2021-10-23T12:29:51.916575", "exception": false, "start_time": "2021-10-23T12:29:51.851615", "status": "completed"} tags=[]
# ### 2.1. Remove Duplicate Data
#
# + papermill={"duration": 0.09018, "end_time": "2021-10-23T12:29:52.072315", "exception": false, "start_time": "2021-10-23T12:29:51.982135", "status": "completed"} tags=[]
duplicates = df[df.duplicated(['text', 'target'], keep=False)]
print(f'Train Duplicate Entries (text, target): {len(duplicates)}')
duplicates.head()
# + papermill={"duration": 0.12414, "end_time": "2021-10-23T12:29:52.301973", "exception": false, "start_time": "2021-10-23T12:29:52.177833", "status": "completed"} tags=[]
df.drop_duplicates(['text', 'target'], inplace=True, ignore_index=True)
print(df.shape, df_test.shape)
# + [markdown] papermill={"duration": 0.105607, "end_time": "2021-10-23T12:29:52.526242", "exception": false, "start_time": "2021-10-23T12:29:52.420635", "status": "completed"} tags=[]
# Duplicates with the same keyword and text share both target classes. The count is low enough to manually review these tweets and drop those with the incorrect target label.
# + papermill={"duration": 0.153488, "end_time": "2021-10-23T12:29:52.784646", "exception": false, "start_time": "2021-10-23T12:29:52.631158", "status": "completed"} tags=[]
new_duplicates = df[df.duplicated(['keyword', 'text'], keep=False)]
print(f'Train Duplicate Entries (keyword, text): {len(new_duplicates)}')
new_duplicates[['text', 'target']].sort_values(by='text')
# + papermill={"duration": 0.121655, "end_time": "2021-10-23T12:29:53.017558", "exception": false, "start_time": "2021-10-23T12:29:52.895903", "status": "completed"} tags=[]
# Drop the target label that is false for each duplicate pair
df.drop([4253, 4193, 2802, 4554, 4182, 3212, 4249, 4259, 6535, 4319, 4239, 606, 3936, 6018, 5573], inplace=True)
# + papermill={"duration": 0.136121, "end_time": "2021-10-23T12:29:53.263792", "exception": false, "start_time": "2021-10-23T12:29:53.127671", "status": "completed"} tags=[]
# Reset the dataframe index to account for missing numbers
df = df.reset_index(drop=True)
df
# + [markdown] papermill={"duration": 0.067183, "end_time": "2021-10-23T12:29:53.402632", "exception": false, "start_time": "2021-10-23T12:29:53.335449", "status": "completed"} tags=[]
# ### 2.2. Examine the Target Data Balance
# + papermill={"duration": 0.077767, "end_time": "2021-10-23T12:29:53.547882", "exception": false, "start_time": "2021-10-23T12:29:53.470115", "status": "completed"} tags=[]
df['target'].value_counts() / len(df)
# + [markdown] papermill={"duration": 0.067554, "end_time": "2021-10-23T12:29:53.683992", "exception": false, "start_time": "2021-10-23T12:29:53.616438", "status": "completed"} tags=[]
# ### 2.3. Check for Null Values
# + papermill={"duration": 0.076547, "end_time": "2021-10-23T12:29:53.828483", "exception": false, "start_time": "2021-10-23T12:29:53.751936", "status": "completed"} tags=[]
def null_table(data):
# Bool is True if values are not null
null_list = []
for i in data:
if data[i].notnull().any():
null_list.append(data[i].notnull().value_counts())
return pd.DataFrame(pd.concat(null_list, axis=1).T)
# + papermill={"duration": 0.091269, "end_time": "2021-10-23T12:29:53.987964", "exception": false, "start_time": "2021-10-23T12:29:53.896695", "status": "completed"} tags=[]
null_table(df)
# + papermill={"duration": 0.088131, "end_time": "2021-10-23T12:29:54.148157", "exception": false, "start_time": "2021-10-23T12:29:54.060026", "status": "completed"} tags=[]
null_table(df_test)
# + [markdown] papermill={"duration": 0.069521, "end_time": "2021-10-23T12:29:54.289955", "exception": false, "start_time": "2021-10-23T12:29:54.220434", "status": "completed"} tags=[]
# A check for null values in the data shows that the `location` column is missing a significant amount information. This could interfere with model performance, so that data will be excluded. In contrast, the `keyword` column has a more acceptable count of missing values so we can fill these with extracted keywords using spaCy.
# + [markdown] papermill={"duration": 0.068377, "end_time": "2021-10-23T12:29:54.426769", "exception": false, "start_time": "2021-10-23T12:29:54.358392", "status": "completed"} tags=[]
# # 3. Text Preprocessing and EDA
# + papermill={"duration": 0.080225, "end_time": "2021-10-23T12:29:54.574956", "exception": false, "start_time": "2021-10-23T12:29:54.494731", "status": "completed"} tags=[]
text = df['text']
target = df['target']
test_text = df_test['text']
# Print random samples from the training text
for i in np.random.randint(500, size=5):
print(f'Tweet #{i}: ', text[i], '=> Target: ', target[i], end='\n' * 2)
# + [markdown] papermill={"duration": 0.068947, "end_time": "2021-10-23T12:29:54.713524", "exception": false, "start_time": "2021-10-23T12:29:54.644577", "status": "completed"} tags=[]
# The next task is to build a text standardization function that is specific to the content found in tweets. Aside from punctuation, there are urls, abbreviations, entities, retweets, digits, stopwords, and of course emojis. The words in each tweet will also be lemmatized or reduced to their root form using the spaCy library. We'll start by building a lookup dictionary with common twitter phrase abbreviations. Tweet terms that match keys in the lookup dictionary will be expanded to their non-abbreviated form.
# + papermill={"duration": 0.084387, "end_time": "2021-10-23T12:29:54.867078", "exception": false, "start_time": "2021-10-23T12:29:54.782691", "status": "completed"} tags=[]
lookup_dict = {
'abt' : 'about',
'afaik' : 'as far as i know',
'bc' : 'because',
'bfn' : 'bye for now',
'bgd' : 'background',
'bh' : 'blockhead',
'br' : 'best regards',
'btw' : 'by the way',
'cc': 'carbon copy',
'chk' : 'check',
'dam' : 'do not annoy me',
'dd' : 'dear daughter',
'df': 'dear fiance',
'ds' : 'dear son',
'dyk' : 'did you know',
'em': 'email',
'ema' : 'email address',
'ftf' : 'face to face',
'fb' : 'facebook',
'ff' : 'follow friday',
'fotd' : 'find of the day',
'ftw': 'for the win',
'fwiw' : 'for what it is worth',
'gts' : 'guess the song',
'hagn' : 'have a good night',
'hand' : 'have a nice day',
'hotd' : 'headline of the day',
'ht' : 'heard through',
'hth' : 'hope that helps',
'ic' : 'i see',
'icymi' : 'in case you missed it',
'idk' : 'i do not know',
'ig': 'instagram',
'iirc' : 'if i remember correctly',
'imho' : 'in my humble opinion',
'imo' : 'in my opinion',
'irl' : 'in real life',
'iwsn' : 'i want sex now',
'jk' : 'just kidding',
'jsyk' : 'just so you know',
'jv' : 'joint venture',
'kk' : 'cool cool',
'kyso' : 'knock your socks off',
'lmao' : 'laugh my ass off',
'lmk' : 'let me know',
'lo' : 'little one',
'lol' : 'laugh out loud',
'mm' : 'music monday',
'mirl' : 'meet in real life',
'mrjn' : 'marijuana',
'nbd' : 'no big deal',
'nct' : 'nobody cares though',
'njoy' : 'enjoy',
'nsfw' : 'not safe for work',
'nts' : 'note to self',
'oh' : 'overheard',
'omg': 'oh my god',
'oomf' : 'one of my friends',
'orly' : 'oh really',
'plmk' : 'please let me know',
'pnp' : 'party and play',
'qotd' : 'quote of the day',
're' : 'in reply to in regards to',
'rtq' : 'read the question',
'rt' : 'retweet',
'sfw' : 'safe for work',
'smdh' : 'shaking my damn head',
'smh' : 'shaking my head',
'so' : 'significant other',
'srs' : 'serious',
'tftf' : 'thanks for the follow',
'tftt' : 'thanks for this tweet',
'tj' : 'tweetjack',
'tl' : 'timeline',
'tldr' : 'too long did not read',
'tmb' : 'tweet me back',
'tt' : 'trending topic',
'ty' : 'thank you',
'tyia' : 'thank you in advance',
'tyt' : 'take your time',
'tyvw' : 'thank you very much',
'w': 'with',
'wtv' : 'whatever',
'ygtr' : 'you got that right',
'ykwim' : 'you know what i mean',
'ykyat' : 'you know you are addicted to',
'ymmv' : 'your mileage may vary',
'yolo' : 'you only live once',
'yoyo' : 'you are on your own',
'yt': 'youtube',
'yw' : 'you are welcome',
'zomg' : 'oh my god to the maximum'
}
# + [markdown] papermill={"duration": 0.068176, "end_time": "2021-10-23T12:29:55.003733", "exception": false, "start_time": "2021-10-23T12:29:54.935557", "status": "completed"} tags=[]
# ### 3.1. Text Standardization Functions
# + papermill={"duration": 0.085547, "end_time": "2021-10-23T12:29:55.162747", "exception": false, "start_time": "2021-10-23T12:29:55.0772", "status": "completed"} tags=[]
def lemmatize_text(text, nlp=nlp):
doc = nlp(text)
lemma_sent = [i.lemma_ for i in doc if not i.is_stop]
return ' '.join(lemma_sent)
def abbrev_conversion(text):
words = text.split()
abbrevs_removed = []
for i in words:
if i in lookup_dict:
i = lookup_dict[i]
abbrevs_removed.append(i)
return ' '.join(abbrevs_removed)
def standardize_text(text_data):
entity_pattern = re.compile(r'(@[A-Za-z0-9]+)|([^0-9A-Za-z \t])|(\w+:\/\/\S+)')
url_pattern = re.compile(r'(?:\@|http?\://|https?\://|www)\S+')
retweet_pattern = re.compile(r'^(RT|RT:)\s+')
digit_pattern = re.compile(r'[\d]+')
# From https://gist.github.com/slowkow/7a7f61f495e3dbb7e3d767f97bd7304
emoji_pattern = re.compile("["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
u"\U00002500-\U00002BEF" # chinese char
u"\U00002702-\U000027B0"
u"\U00002702-\U000027B0"
u"\U000024C2-\U0001F251"
u"\U0001f926-\U0001f937"
u"\U00010000-\U0010ffff"
u"\u2640-\u2642"
u"\u2600-\u2B55"
u"\u200d"
u"\u23cf"
u"\u23e9"
u"\u231a"
u"\ufe0f" # dingbats
u"\u3030"
"]+", flags=re.UNICODE)
# Remove urls
url_strip = text_data.apply(lambda x: re.sub(url_pattern, '', x) if pd.isna(x) != True else x)
# Parse the HTML
html_parse = url_strip.apply(lambda x: bs(x, 'html.parser').get_text() if pd.isna(x) != True else x)
# Remove rewteets
retweet_strip = html_parse.apply(lambda x: re.sub(retweet_pattern, '', x) if pd.isna(x) != True else x)
# Remove emojis
emoji_strip = retweet_strip.apply(lambda x: re.sub(emoji_pattern, '', x) if pd.isna(x) != True else x)
# Remove entities
entity_strip = emoji_strip.apply(lambda x: re.sub(entity_pattern, '', x) if pd.isna(x) != True else x)
# Lowercase the strings
lowercase = entity_strip.apply(lambda x: str.lower(x) if pd.isna(x) != True else x)
# Remove punctuation
punct_strip = lowercase.apply(lambda x: re.sub(f'[{re.escape(string.punctuation)}]', '', x) if pd.isna(x) != True else x)
# Convert abbreviations to words
abbrev_converted = punct_strip.apply(lambda x: abbrev_conversion(x) if pd.isna(x) != True else x)
# Remove digits
digit_strip = abbrev_converted.apply(lambda x: re.sub(digit_pattern, '', x) if pd.isna(x) != True else x)
# Lemmatize text and filter stopwords
lemma_and_stop = digit_strip.apply(lambda x: lemmatize_text(x) if pd.isna(x) != True else x)
return lemma_and_stop
# + papermill={"duration": 76.214584, "end_time": "2021-10-23T12:31:11.445619", "exception": false, "start_time": "2021-10-23T12:29:55.231035", "status": "completed"} tags=[]
clean_text = np.asarray(standardize_text(text))
test_clean_text = np.asarray(standardize_text(test_text))
# Print random samples from the cleaned training text
for i in np.random.randint(500, size=5):
print(f'Tweet #{i}: ', clean_text[i], '=> Target: ', target[i], end='\n' * 2)
# + [markdown] papermill={"duration": 0.069217, "end_time": "2021-10-23T12:31:11.621921", "exception": false, "start_time": "2021-10-23T12:31:11.552704", "status": "completed"} tags=[]
# The text is now in a cleaner format for training, but before taking the next step we should explore it with some visualizations.
# + papermill={"duration": 0.076629, "end_time": "2021-10-23T12:31:11.767155", "exception": false, "start_time": "2021-10-23T12:31:11.690526", "status": "completed"} tags=[]
df['clean_text'] = pd.DataFrame(clean_text)
df_test['clean_text'] = pd.DataFrame(test_clean_text)
# + [markdown] papermill={"duration": 0.068302, "end_time": "2021-10-23T12:31:11.904709", "exception": false, "start_time": "2021-10-23T12:31:11.836407", "status": "completed"} tags=[]
# ### 3.2. Plot Tweet Length Histogram
# + papermill={"duration": 0.363648, "end_time": "2021-10-23T12:31:12.337085", "exception": false, "start_time": "2021-10-23T12:31:11.973437", "status": "completed"} tags=[]
df['tweet_len'] = df['clean_text'].apply(lambda x: len(x))
count, bin_edges = np.histogram(df['tweet_len'])
sns.histplot(data=df, x=df['tweet_len'], bins=bin_edges, hue=df['target'])
plt.title('Tweet Length Frequency')
plt.xlabel('Length of Tweets')
plt.ylabel('Frequency')
plt.show()
# + [markdown] papermill={"duration": 0.07107, "end_time": "2021-10-23T12:31:12.492223", "exception": false, "start_time": "2021-10-23T12:31:12.421153", "status": "completed"} tags=[]
# A tweet length histogram shows that longer tweets at the end of the distribution have a greater frequency of disaster.
# + [markdown] papermill={"duration": 0.070109, "end_time": "2021-10-23T12:31:12.631903", "exception": false, "start_time": "2021-10-23T12:31:12.561794", "status": "completed"} tags=[]
# ### 3.3. Display Non-Disaster and Disaster WordClouds
# + papermill={"duration": 0.565623, "end_time": "2021-10-23T12:31:13.267295", "exception": false, "start_time": "2021-10-23T12:31:12.701672", "status": "completed"} tags=[]
word_cloud_0 = WordCloud(collocations=False, background_color='white').generate(' '.join(df['clean_text'][df['target']==0]))
plt.imshow(word_cloud_0, interpolation='bilinear')
plt.title('Non-Disaster Wordcloud (0)')
plt.axis('off')
plt.show()
# + papermill={"duration": 0.546877, "end_time": "2021-10-23T12:31:13.891239", "exception": false, "start_time": "2021-10-23T12:31:13.344362", "status": "completed"} tags=[]
word_cloud_1 = WordCloud(collocations=False, background_color='black').generate(' '.join(df['clean_text'][df['target']==1]))
plt.imshow(word_cloud_1, interpolation='bilinear')
plt.title('Disaster Wordcloud (1)')
plt.axis('off')
plt.show()
# + [markdown] papermill={"duration": 0.0835, "end_time": "2021-10-23T12:31:14.056675", "exception": false, "start_time": "2021-10-23T12:31:13.973175", "status": "completed"} tags=[]
# ### 3.4. Remove the Word *new* from Text
# + papermill={"duration": 0.108002, "end_time": "2021-10-23T12:31:14.245984", "exception": false, "start_time": "2021-10-23T12:31:14.137982", "status": "completed"} tags=[]
pattern_new = re.compile(r'\bnew\b')
print('Training Counts of \'new\': ', len(re.findall(pattern_new, ' '.join(df['clean_text']))))
print('Test Counts of \'new\': ', len(re.findall(pattern_new, ' '.join(df_test['clean_text']))))
# + papermill={"duration": 0.135099, "end_time": "2021-10-23T12:31:14.461981", "exception": false, "start_time": "2021-10-23T12:31:14.326882", "status": "completed"} tags=[]
# Clean the word 'new' from the training and test data
df['clean_text'] = df['clean_text'].apply(lambda x: re.sub(pattern_new, '', x) if pd.isna(x) != True else x)
df_test['clean_text'] = df_test['clean_text'].apply(lambda x: re.sub(pattern_new, '', x) if pd.isna(x) != True else x)
# + papermill={"duration": 0.106994, "end_time": "2021-10-23T12:31:14.650296", "exception": false, "start_time": "2021-10-23T12:31:14.543302", "status": "completed"} tags=[]
print('Training Counts of \'new\': ', len(re.findall(pattern_new, ' '.join(df['clean_text']))))
print('Test Counts of \'new\': ', len(re.findall(pattern_new, ' '.join(df_test['clean_text']))))
# + [markdown] papermill={"duration": 0.081615, "end_time": "2021-10-23T12:31:14.812795", "exception": false, "start_time": "2021-10-23T12:31:14.73118", "status": "completed"} tags=[]
# ### 3.5. Fill Missing Keywords
#
# There are fifty-six missing keywords in the training data set and twenty-six in the test set. We will use spaCy's part of speech tagging feature to fill the missing words. The process is as follows:
# 1. Create a list of potential keywords by tagging nouns, pronouns, and adjectives
# 2. Use a sentence encoder to embed the list of potential keywords and the comparison text
# 3. Calculate vector distances using the cosine similarity function
# 4. Sort the vectors and select the top keyword for each tweet
#
# The sentence encoder used here is loaded from TensorFlow Hub. It is a pre-trained universal sentence encoder published by Google for use in natural language tasks.
# + papermill={"duration": 22.849206, "end_time": "2021-10-23T12:31:37.744564", "exception": false, "start_time": "2021-10-23T12:31:14.895358", "status": "completed"} tags=[]
# Load the sentence encoder
sentence_enc = hub.load('https://tfhub.dev/google/universal-sentence-encoder/4')
# + [markdown] papermill={"duration": 0.085935, "end_time": "2021-10-23T12:31:37.91465", "exception": false, "start_time": "2021-10-23T12:31:37.828715", "status": "completed"} tags=[]
# ### 3.6. Keyword Extract and Fill Functions
# + papermill={"duration": 0.094613, "end_time": "2021-10-23T12:31:38.095018", "exception": false, "start_time": "2021-10-23T12:31:38.000405", "status": "completed"} tags=[]
def extract_keywords(text, nlp=nlp):
potential_keywords = []
TOP_KEYWORD = -1
# Create a list for keyword parts of speech
pos_tag = ['ADJ', 'NOUN', 'PROPN']
doc = nlp(text)
for i in doc:
if i.pos_ in pos_tag:
potential_keywords.append(i.text)
document_embed = sentence_enc([text])
potential_embed = sentence_enc(potential_keywords)
vector_distances = cosine_similarity(document_embed, potential_embed)
keyword = [potential_keywords[i] for i in vector_distances.argsort()[0][TOP_KEYWORD:]]
return keyword
def keyword_filler(keyword, text):
if pd.isnull(keyword):
try:
keyword = extract_keywords(text)[0]
except:
keyword = ''
return keyword
# + papermill={"duration": 2.702018, "end_time": "2021-10-23T12:31:40.882942", "exception": false, "start_time": "2021-10-23T12:31:38.180924", "status": "completed"} tags=[]
df['keyword_fill'] = pd.DataFrame(list(map(keyword_filler, df['keyword'], df['clean_text']))).astype(str)
df_test['keyword_fill'] = pd.DataFrame(list(map(keyword_filler, df_test['keyword'], df_test['clean_text']))).astype(str)
print('Null Training Keywords => ', df['keyword_fill'].isnull().any())
print('Null Test Keywords => ', df_test['keyword_fill'].isnull().any())
# + [markdown] papermill={"duration": 0.084255, "end_time": "2021-10-23T12:31:41.052563", "exception": false, "start_time": "2021-10-23T12:31:40.968308", "status": "completed"} tags=[]
# Now that the missing keywords are filled we should standardize them to ensure our keywords are clean and ready for training.
# + papermill={"duration": 57.112812, "end_time": "2021-10-23T12:32:38.248989", "exception": false, "start_time": "2021-10-23T12:31:41.136177", "status": "completed"} tags=[]
df['keyword_fill'] = pd.DataFrame(standardize_text(df['keyword_fill']))
df_test['keyword_fill'] = pd.DataFrame(standardize_text(df_test['keyword_fill']))
# + papermill={"duration": 0.16105, "end_time": "2021-10-23T12:32:38.584713", "exception": false, "start_time": "2021-10-23T12:32:38.423663", "status": "completed"} tags=[]
df.head()
# + papermill={"duration": 0.098134, "end_time": "2021-10-23T12:32:38.769796", "exception": false, "start_time": "2021-10-23T12:32:38.671662", "status": "completed"} tags=[]
df_test.head()
# + [markdown] papermill={"duration": 0.083744, "end_time": "2021-10-23T12:32:38.937307", "exception": false, "start_time": "2021-10-23T12:32:38.853563", "status": "completed"} tags=[]
# ### 3.7. Plot the Keyword Frequencies
# + papermill={"duration": 0.098544, "end_time": "2021-10-23T12:32:39.119393", "exception": false, "start_time": "2021-10-23T12:32:39.020849", "status": "completed"} tags=[]
keyword_count_0 = pd.DataFrame(df['keyword_fill'][df['target']==0].value_counts().reset_index())
keyword_count_1 = pd.DataFrame(df['keyword_fill'][df['target']==1].value_counts().reset_index())
# + papermill={"duration": 0.310201, "end_time": "2021-10-23T12:32:39.513371", "exception": false, "start_time": "2021-10-23T12:32:39.20317", "status": "completed"} tags=[]
sns.barplot(data=keyword_count_0[:10], x='keyword_fill', y='index')
plt.title('Non-Disaster Keyword Frequency (0)')
plt.xlabel('Frequency')
plt.ylabel('Top 10 Keywords')
plt.show()
# + papermill={"duration": 0.306676, "end_time": "2021-10-23T12:32:39.904912", "exception": false, "start_time": "2021-10-23T12:32:39.598236", "status": "completed"} tags=[]
sns.barplot(data=keyword_count_1[:10], x='keyword_fill', y='index')
plt.title('Disaster Keyword Frequency (1)')
plt.xlabel('Frequency')
plt.ylabel('Top 10 Keywords')
plt.show()
# + [markdown] papermill={"duration": 0.084722, "end_time": "2021-10-23T12:32:40.075268", "exception": false, "start_time": "2021-10-23T12:32:39.990546", "status": "completed"} tags=[]
# A look at the top keywords for each class demonstrates the importance of observing the tweet itself to find context. At a glance, these keywords could belong to either class. That being the case, keywords can still emphasize important parts of speech that may be overlooked by the model.
# + [markdown] papermill={"duration": 0.084538, "end_time": "2021-10-23T12:32:40.245417", "exception": false, "start_time": "2021-10-23T12:32:40.160879", "status": "completed"} tags=[]
# # 4. Prepare Data for Training
# + [markdown] papermill={"duration": 0.08453, "end_time": "2021-10-23T12:32:40.5843", "exception": false, "start_time": "2021-10-23T12:32:40.49977", "status": "completed"} tags=[]
# ### 4.1. Select the Clean Text and Filled Keyword Columns
# + papermill={"duration": 0.096994, "end_time": "2021-10-23T12:32:40.766536", "exception": false, "start_time": "2021-10-23T12:32:40.669542", "status": "completed"} tags=[]
train_features = df[['clean_text','keyword_fill']]
test_features = df_test[['clean_text', 'keyword_fill']]
# + papermill={"duration": 0.098983, "end_time": "2021-10-23T12:32:40.951773", "exception": false, "start_time": "2021-10-23T12:32:40.85279", "status": "completed"} tags=[]
train_features[:5]
# + papermill={"duration": 0.099334, "end_time": "2021-10-23T12:32:41.13773", "exception": false, "start_time": "2021-10-23T12:32:41.038396", "status": "completed"} tags=[]
test_features[:5]
# + papermill={"duration": 0.094271, "end_time": "2021-10-23T12:32:41.318218", "exception": false, "start_time": "2021-10-23T12:32:41.223947", "status": "completed"} tags=[]
print(train_features.shape)
print(test_features.shape)
# + [markdown] papermill={"duration": 0.086211, "end_time": "2021-10-23T12:32:41.491361", "exception": false, "start_time": "2021-10-23T12:32:41.40515", "status": "completed"} tags=[]
# ### 4.2. Create Data Splits
# + papermill={"duration": 0.101088, "end_time": "2021-10-23T12:32:41.678491", "exception": false, "start_time": "2021-10-23T12:32:41.577403", "status": "completed"} tags=[]
train_x, val_x, train_y, val_y = train_test_split(
train_features,
target,
test_size=0.2,
random_state=RANDOM_SEED,
)
print(train_x.shape)
print(train_y.shape)
print(val_x.shape)
print(val_y.shape)
# + [markdown] papermill={"duration": 0.086268, "end_time": "2021-10-23T12:32:41.851746", "exception": false, "start_time": "2021-10-23T12:32:41.765478", "status": "completed"} tags=[]
# ### 4.3. Create TensorFlow Datasets
# + papermill={"duration": 0.1074, "end_time": "2021-10-23T12:32:42.047027", "exception": false, "start_time": "2021-10-23T12:32:41.939627", "status": "completed"} tags=[]
# Create TensorFlow Datasets
train_ds = tf.data.Dataset.from_tensor_slices((dict(train_x), train_y))
val_ds = tf.data.Dataset.from_tensor_slices((dict(val_x), val_y))
test_ds = tf.data.Dataset.from_tensor_slices(dict(test_features))
# + papermill={"duration": 0.095147, "end_time": "2021-10-23T12:32:42.228344", "exception": false, "start_time": "2021-10-23T12:32:42.133197", "status": "completed"} tags=[]
AUTOTUNE = tf.data.experimental.AUTOTUNE
BUFFER_SIZE = 1000
BATCH_SIZE = 32
def configure_dataset(dataset, shuffle=False, test=False):
# Configure the tf dataset for cache, shuffle, batch, and prefetch
if shuffle:
dataset = dataset.cache()\
.shuffle(BUFFER_SIZE, seed=RANDOM_SEED, reshuffle_each_iteration=True)\
.batch(BATCH_SIZE, drop_remainder=True).prefetch(AUTOTUNE)
elif test:
dataset = dataset.cache()\
.batch(BATCH_SIZE, drop_remainder=False).prefetch(AUTOTUNE)
else:
dataset = dataset.cache()\
.batch(BATCH_SIZE, drop_remainder=True).prefetch(AUTOTUNE)
return dataset
# + papermill={"duration": 0.107724, "end_time": "2021-10-23T12:32:42.422402", "exception": false, "start_time": "2021-10-23T12:32:42.314678", "status": "completed"} tags=[]
# Configure the datasets
train_ds = configure_dataset(train_ds, shuffle=True)
val_ds = configure_dataset(val_ds)
test_ds = configure_dataset(test_ds, test=True)
# + papermill={"duration": 0.099372, "end_time": "2021-10-23T12:32:42.60802", "exception": false, "start_time": "2021-10-23T12:32:42.508648", "status": "completed"} tags=[]
# Print the dataset specifications
print(train_ds.element_spec)
print(val_ds.element_spec)
print(test_ds.element_spec)
# + [markdown] papermill={"duration": 0.086528, "end_time": "2021-10-23T12:32:42.785723", "exception": false, "start_time": "2021-10-23T12:32:42.699195", "status": "completed"} tags=[]
# # 5. Building the Classifier Model
#
# For this task we will be using a pre-trained BERT model loaded from TensorFlow Hub. This model has 12 hidden layers, a hidden unit size of 768, and 12 attention heads. It has a companion preprocessor that is loaded from the same repository. This preprocessor takes text segments and converts them to numeric token ids accepted by the BERT encoder. These token ids are:
# * input_word_ids
# - ids of the input sequences
# * input_mask
# - represents all pre-padded input tokens as 1, and padded tokens as 0
# * input_type_ids
# - contains indices for each input segment with padding locations indexed at 0
# + [markdown] papermill={"duration": 0.087986, "end_time": "2021-10-23T12:32:42.961039", "exception": false, "start_time": "2021-10-23T12:32:42.873053", "status": "completed"} tags=[]
# ### 5.1. Load the Pre-trained BERT Encoder
# + papermill={"duration": 24.516365, "end_time": "2021-10-23T12:33:07.565057", "exception": false, "start_time": "2021-10-23T12:32:43.048692", "status": "completed"} tags=[]
# BERT encoder w/ preprocessor
bert_preprocessor = hub.KerasLayer('https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3', name='BERT_preprocesser')
bert_encoder = hub.KerasLayer('https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/4', trainable=True, name='BERT_encoder')
# Keyword embedding layer
nnlm_embed = hub.KerasLayer('https://tfhub.dev/google/nnlm-en-dim50/2', name='embedding_layer')
# + [markdown] papermill={"duration": 0.08899, "end_time": "2021-10-23T12:33:07.7494", "exception": false, "start_time": "2021-10-23T12:33:07.66041", "status": "completed"} tags=[]
# ### 5.2. Model Composition
#
# To build our classifier we will be using the TensorFlow functional API, this will reduce constraints on our model design. Two input branches will be merged into a classification layer. The first branch is a text input layer that feeds into the BERT preprocessor. This layer is passed to the BERT encoder and is returned as a pooled output. This output is then regularized with a dropout layer.
#
# On the second branch, a keyword input is passed to a pre-trained word embedding layer. The embeddings are flattened and passed into a dense neural net, then fed into a dropout layer.
#
# The layer outputs from each model are concatenated, passed into a dense neural net with dropout, then sent to a single unit dense classification layer with a sigmoid activation. The sigmoid activation function will return class probabilities that we can use to plot a Receiver Operating Characteristic ([ROC](https://en.wikipedia.org/wiki/Receiver_operating_characteristic)) curve and a [Confusion Matrix](https://en.wikipedia.org/wiki/Confusion_matrix) to analyze our results.
# + papermill={"duration": 0.10073, "end_time": "2021-10-23T12:33:07.93693", "exception": false, "start_time": "2021-10-23T12:33:07.8362", "status": "completed"} tags=[]
def build_model():
# Construct text layers
text_input = layers.Input(shape=(), dtype=tf.string, name='clean_text') # Name matches df heading
encoder_inputs = bert_preprocessor(text_input)
encoder_outputs = bert_encoder(encoder_inputs)
# pooled_output returns [batch_size, hidden_layers]
pooled_output = encoder_outputs["pooled_output"]
bert_dropout = layers.Dropout(0.1, name='BERT_dropout')(pooled_output)
# Construct keyword layers
key_input = layers.Input(shape=(), dtype=tf.string, name='keyword_fill') # Name matches df heading
key_embed = nnlm_embed(key_input)
key_flat = layers.Flatten()(key_embed)
key_dense = layers.Dense(128, activation='elu', kernel_regularizer=regularizers.l2(1e-4))(key_flat)
key_dropout = layers.Dropout(0.5, name='dense_dropout')(key_dense)
# Merge the layers and classify
merge = layers.concatenate([bert_dropout, key_dropout])
dense = layers.Dense(128, activation='elu', kernel_regularizer=regularizers.l2(1e-4))(merge)
dropout = layers.Dropout(0.5, name='merged_dropout')(dense)
clf = layers.Dense(1, activation='sigmoid', name='classifier')(dropout)
return Model([text_input, key_input], clf, name='BERT_classifier')
# + papermill={"duration": 0.831378, "end_time": "2021-10-23T12:33:08.855511", "exception": false, "start_time": "2021-10-23T12:33:08.024133", "status": "completed"} tags=[]
bert_classifier = build_model()
bert_classifier.summary()
# + papermill={"duration": 0.575985, "end_time": "2021-10-23T12:33:09.521281", "exception": false, "start_time": "2021-10-23T12:33:08.945296", "status": "completed"} tags=[]
tf.keras.utils.plot_model(bert_classifier, show_shapes=False, dpi=96)
# + [markdown] papermill={"duration": 0.0878, "end_time": "2021-10-23T12:33:09.69797", "exception": false, "start_time": "2021-10-23T12:33:09.61017", "status": "completed"} tags=[]
# For training the BERT model we'll use an Adam optimizer with weight decay ([AdamW](https://arxiv.org/abs/1711.05101)). This method differs from the standard Adam algorithm with its use of decoupled weight decay regularization. This is the optimizer that BERT was originally trained with. It has a linear warm-up period over the first 10% of training steps paired with a lower learning rate. One of the simpler ways to implement this [optimizer](https://github.com/tensorflow/models/blob/master/official/nlp/optimization.py) is with the TensorFlow official models collection.
# + [markdown] papermill={"duration": 0.088118, "end_time": "2021-10-23T12:33:09.874013", "exception": false, "start_time": "2021-10-23T12:33:09.785895", "status": "completed"} tags=[]
# ### 5.3. Construct the AdamW Optimizer
# + papermill={"duration": 0.112748, "end_time": "2021-10-23T12:33:10.075207", "exception": false, "start_time": "2021-10-23T12:33:09.962459", "status": "completed"} tags=[]
EPOCHS = 5
LEARNING_RATE = 5e-5
STEPS_PER_EPOCH = int(train_ds.unbatch().cardinality().numpy() / BATCH_SIZE)
VAL_STEPS = int(val_ds.unbatch().cardinality().numpy() / BATCH_SIZE)
# Calculate the train and warmup steps for the optimizer
TRAIN_STEPS = STEPS_PER_EPOCH * EPOCHS
WARMUP_STEPS = int(TRAIN_STEPS * 0.1)
adamw_optimizer = create_optimizer(
init_lr=LEARNING_RATE,
num_train_steps=TRAIN_STEPS,
num_warmup_steps=WARMUP_STEPS
)
# + [markdown] papermill={"duration": 0.08739, "end_time": "2021-10-23T12:33:10.249988", "exception": false, "start_time": "2021-10-23T12:33:10.162598", "status": "completed"} tags=[]
# # 6. Train the Classifier
# + papermill={"duration": 236.542108, "end_time": "2021-10-23T12:37:06.891097", "exception": false, "start_time": "2021-10-23T12:33:10.348989", "status": "completed"} tags=[]
bert_classifier.compile(
loss=BinaryCrossentropy(from_logits=True),
optimizer= adamw_optimizer,
metrics=[BinaryAccuracy(name='accuracy')]
)
history = bert_classifier.fit(
train_ds,
epochs=EPOCHS,
steps_per_epoch=STEPS_PER_EPOCH,
validation_data= val_ds,
validation_steps=VAL_STEPS
)
# + [markdown] papermill={"duration": 0.184136, "end_time": "2021-10-23T12:37:07.262161", "exception": false, "start_time": "2021-10-23T12:37:07.078025", "status": "completed"} tags=[]
# # 7. Visualize Results
# + [markdown] papermill={"duration": 0.182765, "end_time": "2021-10-23T12:37:07.630594", "exception": false, "start_time": "2021-10-23T12:37:07.447829", "status": "completed"} tags=[]
# ### 7.1. Plot the Loss and Accuracy Metrics
# + papermill={"duration": 0.191166, "end_time": "2021-10-23T12:37:08.005126", "exception": false, "start_time": "2021-10-23T12:37:07.81396", "status": "completed"} tags=[]
# Assign the loss and accuracy metrics
train_loss = history.history['loss']
val_loss = history.history['val_loss']
train_acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
# + papermill={"duration": 0.773409, "end_time": "2021-10-23T12:37:08.96224", "exception": false, "start_time": "2021-10-23T12:37:08.188831", "status": "completed"} tags=[]
# Plot the training and validation metrics
fig, (ax1, ax2) = plt.subplots(2, 1, sharex=True)
sns.lineplot(ax=ax1, data = train_acc, label=f'Training Accuracy')
sns.lineplot(ax=ax1, data = val_acc, label=f'Validation Accuracy')
sns.lineplot(ax=ax2, data = train_loss, label=f'Training Loss')
sns.lineplot(ax=ax2, data = val_loss, label=f'Validation Loss')
ax1.set_ylabel('Accuracy')
ax1.set_xlim(xmin=0)
ax2.set_ylabel('Loss')
ax2.set_xlabel('Epochs')
ax2.set_xlim(xmin=0)
plt.suptitle('History')
plt.show()
# + [markdown] papermill={"duration": 0.185214, "end_time": "2021-10-23T12:37:09.348492", "exception": false, "start_time": "2021-10-23T12:37:09.163278", "status": "completed"} tags=[]
# ### 7.2. Plot the ROC Curve
# + papermill={"duration": 0.421802, "end_time": "2021-10-23T12:37:09.957115", "exception": false, "start_time": "2021-10-23T12:37:09.535313", "status": "completed"} tags=[]
# Get the array of labels from the validation Dataset
val_target = np.asarray([i[1] for i in list(val_ds.unbatch().as_numpy_iterator())])
print(val_target.shape)
val_target[:5]
# + papermill={"duration": 10.369488, "end_time": "2021-10-23T12:37:20.513182", "exception": false, "start_time": "2021-10-23T12:37:10.143694", "status": "completed"} tags=[]
# Get predictions from the validation Dataset
val_predict = bert_classifier.predict(val_ds)
# + papermill={"duration": 0.375277, "end_time": "2021-10-23T12:37:21.074619", "exception": false, "start_time": "2021-10-23T12:37:20.699342", "status": "completed"} tags=[]
# Get the false positive and true positive rates
fpr, tpr, _ = roc_curve(val_target, val_predict)
plt.plot(fpr, tpr, color='orange')
plt.plot([0,1], [0,1], linestyle='--')
plt.title('Validation ROC Curve')
plt.xlabel('False Positives (%)')
plt.ylabel('True Positives (%)')
plt.grid(True)
plt.show()
# + [markdown] papermill={"duration": 0.186593, "end_time": "2021-10-23T12:37:21.449078", "exception": false, "start_time": "2021-10-23T12:37:21.262485", "status": "completed"} tags=[]
# ### 7.3. Plot the Confusion Matrix
# + papermill={"duration": 0.443717, "end_time": "2021-10-23T12:37:22.079777", "exception": false, "start_time": "2021-10-23T12:37:21.63606", "status": "completed"} tags=[]
THRESHOLD = 0.5 # Default value
# Get the true negative, false positive, false negative, and true positive values
tn, fp, fn, tp = confusion_matrix(val_target, val_predict > THRESHOLD).flatten()
# Construct the dataframe
cm = pd.DataFrame(
[[tn, fp], [fn, tp]],
index=['No Disaster', 'Disaster'],
columns=['No Disaster', 'Disaster']
)
# Plot the matrix
sns.heatmap(cm, annot=True, fmt='g')
plt.title('Validation Confusion Matrix')
plt.ylabel('Actual')
plt.xlabel('Predicted')
plt.show()
# + [markdown] papermill={"duration": 0.19714, "end_time": "2021-10-23T12:37:22.465594", "exception": false, "start_time": "2021-10-23T12:37:22.268454", "status": "completed"} tags=[]
# # 8. Predictions on the Test Data
# + papermill={"duration": 21.818374, "end_time": "2021-10-23T12:37:44.471442", "exception": false, "start_time": "2021-10-23T12:37:22.653068", "status": "completed"} tags=[]
predictions = bert_classifier.predict(test_ds)
print(predictions.shape)
print(predictions[:5])
# + [markdown] papermill={"duration": 0.187719, "end_time": "2021-10-23T12:37:44.84976", "exception": false, "start_time": "2021-10-23T12:37:44.662041", "status": "completed"} tags=[]
# ### 8.1. Predictions Histogram
# + papermill={"duration": 0.48692, "end_time": "2021-10-23T12:37:45.52792", "exception": false, "start_time": "2021-10-23T12:37:45.041", "status": "completed"} tags=[]
count, bin_edges = np.histogram(predictions)
sns.histplot(predictions, bins=bin_edges, legend=False)
plt.axvline(x=THRESHOLD, linestyle='--', color='black', label='Threshold')
plt.title('Predicted Probability of Disaster')
plt.xlabel('Probabilities')
plt.ylabel('Frequency')
plt.legend()
plt.show()
# + [markdown] papermill={"duration": 0.189694, "end_time": "2021-10-23T12:37:45.908704", "exception": false, "start_time": "2021-10-23T12:37:45.71901", "status": "completed"} tags=[]
# This distribution represents the binary nature of the prediction values. The threshold marks the line by which predictions will be considered either non-disasters (below threshold) or disasters (above threshold). The default value for a binary accuracy threshold is `0.5`. Finally we use this threshold to label and submit the predictions.
# + [markdown] papermill={"duration": 0.191661, "end_time": "2021-10-23T12:37:46.290662", "exception": false, "start_time": "2021-10-23T12:37:46.099001", "status": "completed"} tags=[]
# ### 8.2. Label the Predictions
# + papermill={"duration": 0.204798, "end_time": "2021-10-23T12:37:46.689367", "exception": false, "start_time": "2021-10-23T12:37:46.484569", "status": "completed"} tags=[]
# Use the threshold to label predictions
predictions = np.where(predictions > THRESHOLD, 1, 0)
df_predictions = pd.DataFrame(predictions)
df_predictions.columns = ['target']
print(df_predictions.shape)
df_predictions.head()
# + [markdown] papermill={"duration": 0.189498, "end_time": "2021-10-23T12:37:47.070557", "exception": false, "start_time": "2021-10-23T12:37:46.881059", "status": "completed"} tags=[]
# ### 8.3. Submit the Predictions
# + papermill={"duration": 0.209913, "end_time": "2021-10-23T12:37:47.470916", "exception": false, "start_time": "2021-10-23T12:37:47.261003", "status": "completed"} tags=[]
# Concatenate the columns and convert submission to csv
submission = pd.concat([df_test['id'], df_predictions], axis=1)
submission.to_csv('submission.csv', index=False)
# + papermill={"duration": 0.194747, "end_time": "2021-10-23T12:37:47.854992", "exception": false, "start_time": "2021-10-23T12:37:47.660245", "status": "completed"} tags=[]
| BERT-with-TF_& _Spacy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Load Loan Prediction Dataset
import pandas as pd
loan_data = pd.read_csv('train_loan.csv')
loan_data.head()
# ### Feature 1 -Total Income
loan_data['Total_income'] = loan_data['ApplicantIncome'] + loan_data['CoapplicantIncome']
loan_data[['ApplicantIncome', 'CoapplicantIncome', 'Total_income']].head()
# ### Feature 2 - Loan amount and Income Ratio
loan_data['loan_income_ratio'] = loan_data['LoanAmount'] / loan_data['ApplicantIncome']
loan_data[['ApplicantIncome', 'LoanAmount', 'loan_income_ratio']].head()
| Feature Engineering/Feature Interaction/Feature Interaction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/PytorchLightning/pytorch-lightning/blob/master/notebooks/02-datamodules.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2O5r7QvP8-rt" colab_type="text"
# # PyTorch Lightning DataModules ⚡
#
# With the release of `pytorch-lightning` version 0.9.0, we have included a new class called `LightningDataModule` to help you decouple data related hooks from your `LightningModule`.
#
# This notebook will walk you through how to start using Datamodules.
#
# The most up to date documentation on datamodules can be found [here](https://pytorch-lightning.readthedocs.io/en/latest/datamodules.html).
#
# ---
#
# - Give us a ⭐ [on Github](https://www.github.com/PytorchLightning/pytorch-lightning/)
# - Check out [the documentation](https://pytorch-lightning.readthedocs.io/en/latest/)
# - Join us [on Slack](https://join.slack.com/t/pytorch-lightning/shared_invite/zt-f6bl2l0l-JYMK3tbAgAmGRrlNr00f1A)
# + [markdown] id="6RYMhmfA9ATN" colab_type="text"
# ### Setup
# Lightning is easy to install. Simply ```pip install pytorch-lightning```
# + id="lj2zD-wsbvGr" colab_type="code" colab={}
# ! pip install pytorch-lightning --quiet
# + [markdown] id="8g2mbvy-9xDI" colab_type="text"
# # Introduction
#
# First, we'll go over a regular `LightningModule` implementation without the use of a `LightningDataModule`
# + id="eg-xDlmDdAwy" colab_type="code" colab={}
import pytorch_lightning as pl
from pytorch_lightning.metrics.functional import accuracy
import torch
from torch import nn
import torch.nn.functional as F
from torch.utils.data import random_split, DataLoader
# Note - you must have torchvision installed for this example
from torchvision.datasets import MNIST, CIFAR10
from torchvision import transforms
# + [markdown] id="DzgY7wi88UuG" colab_type="text"
# ## Defining the LitMNISTModel
#
# Below, we reuse a `LightningModule` from our hello world tutorial that classifies MNIST Handwritten Digits.
#
# Unfortunately, we have hardcoded dataset-specific items within the model, forever limiting it to working with MNIST Data. 😢
#
# This is fine if you don't plan on training/evaluating your model on different datasets. However, in many cases, this can become bothersome when you want to try out your architecture with different datasets.
# + id="IQkW8_FF5nU2" colab_type="code" colab={}
class LitMNIST(pl.LightningModule):
def __init__(self, data_dir='./', hidden_size=64, learning_rate=2e-4):
super().__init__()
# We hardcode dataset specific stuff here.
self.data_dir = data_dir
self.num_classes = 10
self.dims = (1, 28, 28)
channels, width, height = self.dims
self.transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])
self.hidden_size = hidden_size
self.learning_rate = learning_rate
# Build model
self.model = nn.Sequential(
nn.Flatten(),
nn.Linear(channels * width * height, hidden_size),
nn.ReLU(),
nn.Dropout(0.1),
nn.Linear(hidden_size, hidden_size),
nn.ReLU(),
nn.Dropout(0.1),
nn.Linear(hidden_size, self.num_classes)
)
def forward(self, x):
x = self.model(x)
return F.log_softmax(x, dim=1)
def training_step(self, batch, batch_idx):
x, y = batch
logits = self(x)
loss = F.nll_loss(logits, y)
return pl.TrainResult(loss)
def validation_step(self, batch, batch_idx):
x, y = batch
logits = self(x)
loss = F.nll_loss(logits, y)
preds = torch.argmax(logits, dim=1)
acc = accuracy(preds, y)
result = pl.EvalResult(checkpoint_on=loss)
result.log('val_loss', loss, prog_bar=True)
result.log('val_acc', acc, prog_bar=True)
return result
def configure_optimizers(self):
optimizer = torch.optim.Adam(self.parameters(), lr=self.learning_rate)
return optimizer
####################
# DATA RELATED HOOKS
####################
def prepare_data(self):
# download
MNIST(self.data_dir, train=True, download=True)
MNIST(self.data_dir, train=False, download=True)
def setup(self, stage=None):
# Assign train/val datasets for use in dataloaders
if stage == 'fit' or stage is None:
mnist_full = MNIST(self.data_dir, train=True, transform=self.transform)
self.mnist_train, self.mnist_val = random_split(mnist_full, [55000, 5000])
# Assign test dataset for use in dataloader(s)
if stage == 'test' or stage is None:
self.mnist_test = MNIST(self.data_dir, train=False, transform=self.transform)
def train_dataloader(self):
return DataLoader(self.mnist_train, batch_size=32)
def val_dataloader(self):
return DataLoader(self.mnist_val, batch_size=32)
def test_dataloader(self):
return DataLoader(self.mnist_test, batch_size=32)
# + [markdown] id="K7sg9KQd-QIO" colab_type="text"
# ## Training the ListMNIST Model
# + id="QxDNDaus6byD" colab_type="code" colab={}
model = LitMNIST()
trainer = pl.Trainer(max_epochs=2, gpus=1, progress_bar_refresh_rate=20)
trainer.fit(model)
# + [markdown] id="dY8d6GxmB0YU" colab_type="text"
# # Using DataModules
#
# DataModules are a way of decoupling data-related hooks from the `LightningModule` so you can develop dataset agnostic models.
# + [markdown] id="eJeT5bW081wn" colab_type="text"
# ## Defining The MNISTDataModule
#
# Let's go over each function in the class below and talk about what they're doing:
#
# 1. ```__init__```
# - Takes in a `data_dir` arg that points to where you have downloaded/wish to download the MNIST dataset.
# - Defines a transform that will be applied across train, val, and test dataset splits.
# - Defines default `self.dims`, which is a tuple returned from `datamodule.size()` that can help you initialize models.
#
#
# 2. ```prepare_data```
# - This is where we can download the dataset. We point to our desired dataset and ask torchvision's `MNIST` dataset class to download if the dataset isn't found there.
# - **Note we do not make any state assignments in this function** (i.e. `self.something = ...`)
#
# 3. ```setup```
# - Loads in data from file and prepares PyTorch tensor datasets for each split (train, val, test).
# - Setup expects a 'stage' arg which is used to separate logic for 'fit' and 'test'.
# - If you don't mind loading all your datasets at once, you can set up a condition to allow for both 'fit' related setup and 'test' related setup to run whenever `None` is passed to `stage`.
# - **Note this runs across all GPUs and it *is* safe to make state assignments here**
#
#
# 4. ```x_dataloader```
# - `train_dataloader()`, `val_dataloader()`, and `test_dataloader()` all return PyTorch `DataLoader` instances that are created by wrapping their respective datasets that we prepared in `setup()`
# + id="DfGKyGwG_X9v" colab_type="code" colab={}
class MNISTDataModule(pl.LightningDataModule):
def __init__(self, data_dir: str = './'):
super().__init__()
self.data_dir = data_dir
self.transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])
# self.dims is returned when you call dm.size()
# Setting default dims here because we know them.
# Could optionally be assigned dynamically in dm.setup()
self.dims = (1, 28, 28)
self.num_classes = 10
def prepare_data(self):
# download
MNIST(self.data_dir, train=True, download=True)
MNIST(self.data_dir, train=False, download=True)
def setup(self, stage=None):
# Assign train/val datasets for use in dataloaders
if stage == 'fit' or stage is None:
mnist_full = MNIST(self.data_dir, train=True, transform=self.transform)
self.mnist_train, self.mnist_val = random_split(mnist_full, [55000, 5000])
# Assign test dataset for use in dataloader(s)
if stage == 'test' or stage is None:
self.mnist_test = MNIST(self.data_dir, train=False, transform=self.transform)
def train_dataloader(self):
return DataLoader(self.mnist_train, batch_size=32)
def val_dataloader(self):
return DataLoader(self.mnist_val, batch_size=32)
def test_dataloader(self):
return DataLoader(self.mnist_test, batch_size=32)
# + [markdown] id="H2Yoj-9M9dS7" colab_type="text"
# ## Defining the dataset agnostic `LitModel`
#
# Below, we define the same model as the `LitMNIST` model we made earlier.
#
# However, this time our model has the freedom to use any input data that we'd like 🔥.
# + id="PM2IISuOBDIu" colab_type="code" colab={}
class LitModel(pl.LightningModule):
def __init__(self, channels, width, height, num_classes, hidden_size=64, learning_rate=2e-4):
super().__init__()
# We take in input dimensions as parameters and use those to dynamically build model.
self.channels = channels
self.width = width
self.height = height
self.num_classes = num_classes
self.hidden_size = hidden_size
self.learning_rate = learning_rate
self.model = nn.Sequential(
nn.Flatten(),
nn.Linear(channels * width * height, hidden_size),
nn.ReLU(),
nn.Dropout(0.1),
nn.Linear(hidden_size, hidden_size),
nn.ReLU(),
nn.Dropout(0.1),
nn.Linear(hidden_size, num_classes)
)
def forward(self, x):
x = self.model(x)
return F.log_softmax(x, dim=1)
def training_step(self, batch, batch_idx):
x, y = batch
logits = self(x)
loss = F.nll_loss(logits, y)
return pl.TrainResult(loss)
def validation_step(self, batch, batch_idx):
x, y = batch
logits = self(x)
loss = F.nll_loss(logits, y)
preds = torch.argmax(logits, dim=1)
acc = accuracy(preds, y)
result = pl.EvalResult(checkpoint_on=loss)
result.log('val_loss', loss, prog_bar=True)
result.log('val_acc', acc, prog_bar=True)
return result
def configure_optimizers(self):
optimizer = torch.optim.Adam(self.parameters(), lr=self.learning_rate)
return optimizer
# + [markdown] id="G4Z5olPe-xEo" colab_type="text"
# ## Training the `LitModel` using the `MNISTDataModule`
#
# Now, we initialize and train the `LitModel` using the `MNISTDataModule`'s configuration settings and dataloaders.
# + id="kV48vP_9mEli" colab_type="code" colab={}
# Init DataModule
dm = MNISTDataModule()
# Init model from datamodule's attributes
model = LitModel(*dm.size(), dm.num_classes)
# Init trainer
trainer = pl.Trainer(max_epochs=3, progress_bar_refresh_rate=20, gpus=1)
# Pass the datamodule as arg to trainer.fit to override model hooks :)
trainer.fit(model, dm)
# + [markdown] id="WNxrugIGRRv5" colab_type="text"
# ## Defining the CIFAR10 DataModule
#
# Lets prove the `LitModel` we made earlier is dataset agnostic by defining a new datamodule for the CIFAR10 dataset.
# + id="1tkaYLU7RT5P" colab_type="code" colab={}
class CIFAR10DataModule(pl.LightningDataModule):
def __init__(self, data_dir: str = './'):
super().__init__()
self.data_dir = data_dir
self.transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
self.dims = (3, 32, 32)
self.num_classes = 10
def prepare_data(self):
# download
CIFAR10(self.data_dir, train=True, download=True)
CIFAR10(self.data_dir, train=False, download=True)
def setup(self, stage=None):
# Assign train/val datasets for use in dataloaders
if stage == 'fit' or stage is None:
cifar_full = CIFAR10(self.data_dir, train=True, transform=self.transform)
self.cifar_train, self.cifar_val = random_split(cifar_full, [45000, 5000])
# Assign test dataset for use in dataloader(s)
if stage == 'test' or stage is None:
self.cifar_test = CIFAR10(self.data_dir, train=False, transform=self.transform)
def train_dataloader(self):
return DataLoader(self.cifar_train, batch_size=32)
def val_dataloader(self):
return DataLoader(self.cifar_val, batch_size=32)
def test_dataloader(self):
return DataLoader(self.cifar_test, batch_size=32)
# + [markdown] id="BrXxf3oX_gsZ" colab_type="text"
# ## Training the `LitModel` using the `CIFAR10DataModule`
#
# Our model isn't very good, so it will perform pretty badly on the CIFAR10 dataset.
#
# The point here is that we can see that our `LitModel` has no problem using a different datamodule as its input data.
# + id="sd-SbWi_krdj" colab_type="code" colab={}
dm = CIFAR10DataModule()
model = LitModel(*dm.size(), dm.num_classes, hidden_size=256)
trainer = pl.Trainer(max_epochs=5, progress_bar_refresh_rate=20, gpus=1)
trainer.fit(model, dm)
| notebooks/02-datamodules.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# # Hyperparameter tuning with Raytune and visulization using Tensorboard and Weights & Biases
# ## Initial imports
# +
import os
import numpy as np
import pandas as pd
import torch
from torch.optim import SGD, lr_scheduler
from pytorch_widedeep import Trainer
from pytorch_widedeep.preprocessing import TabPreprocessor
from pytorch_widedeep.models import TabMlp, WideDeep
from torchmetrics import F1 as F1_torchmetrics
from torchmetrics import Accuracy as Accuracy_torchmetrics
from torchmetrics import Precision as Precision_torchmetrics
from torchmetrics import Recall as Recall_torchmetrics
from pytorch_widedeep.metrics import Accuracy, Recall, Precision, F1Score, R2Score
from pytorch_widedeep.initializers import XavierNormal
from pytorch_widedeep.callbacks import (
EarlyStopping,
ModelCheckpoint,
RayTuneReporter,
)
from pytorch_widedeep.datasets import load_bio_kdd04
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
from ray import tune
from ray.tune.schedulers import AsyncHyperBandScheduler
from ray.tune import JupyterNotebookReporter
from ray.tune.integration.wandb import WandbLoggerCallback, wandb_mixin
import wandb
import tracemalloc
tracemalloc.start()
# increase displayed columns in jupyter notebook
pd.set_option("display.max_columns", 200)
pd.set_option("display.max_rows", 300)
# -
df = load_bio_kdd04(as_frame=True)
df.head()
# imbalance of the classes
df['target'].value_counts()
# drop columns we won't need in this example
df.drop(columns=['EXAMPLE_ID', 'BLOCK_ID'], inplace=True)
df_train, df_valid = train_test_split(df, test_size=0.2, stratify=df['target'], random_state=1)
df_valid, df_test = train_test_split(df_valid, test_size=0.5, stratify=df_valid['target'], random_state=1)
# ## Preparing the data
continuous_cols = df.drop(columns=['target']).columns.values.tolist()
# +
# deeptabular
tab_preprocessor = TabPreprocessor(continuous_cols=continuous_cols, scale=True)
X_tab_train = tab_preprocessor.fit_transform(df_train)
X_tab_valid = tab_preprocessor.transform(df_valid)
X_tab_test = tab_preprocessor.transform(df_test)
# target
y_train = df_train["target"].values
y_valid = df_valid["target"].values
y_test = df_test["target"].values
# -
# ## Define the model
input_layer = len(tab_preprocessor.continuous_cols)
output_layer = 1
hidden_layers = np.linspace(
input_layer * 2, output_layer, 5, endpoint=False, dtype=int
).tolist()
deeptabular = TabMlp(
mlp_hidden_dims=hidden_layers,
column_idx=tab_preprocessor.column_idx,
continuous_cols=tab_preprocessor.continuous_cols,
)
model = WideDeep(deeptabular=deeptabular)
model
# Metrics from torchmetrics
accuracy = Accuracy_torchmetrics(average=None, num_classes=2)
precision = Precision_torchmetrics(average="micro", num_classes=2)
f1 = F1_torchmetrics(average=None, num_classes=2)
recall = Recall_torchmetrics(average=None, num_classes=2)
# +
# # Metrics from pytorch-widedeep
# accuracy = Accuracy(top_k=2)
# precision = Precision(average=False)
# recall = Recall(average=True)
# f1 = F1Score(average=False)
# +
config = {
"batch_size": tune.grid_search([1000, 5000]),
"wandb": {
"project": "test",
"api_key_file": os.getcwd() + "/wandb_api.key",
},
}
# Optimizers
deep_opt = SGD(model.deeptabular.parameters(), lr=0.1)
# LR Scheduler
deep_sch = lr_scheduler.StepLR(deep_opt, step_size=3)
@wandb_mixin
def training_function(config, X_train, X_val):
early_stopping = EarlyStopping()
model_checkpoint = ModelCheckpoint(save_best_only=True,
wb=wandb)
# Hyperparameters
batch_size = config["batch_size"]
trainer = Trainer(
model,
objective="binary_focal_loss",
callbacks=[RayTuneReporter, early_stopping, model_checkpoint],
lr_schedulers={"deeptabular": deep_sch},
initializers={"deeptabular": XavierNormal},
optimizers={"deeptabular": deep_opt},
metrics=[accuracy, precision, recall, f1],
verbose=0,
)
trainer.fit(X_train=X_train, X_val=X_val, n_epochs=5, batch_size=batch_size)
X_train = {"X_tab": X_tab_train, "target": y_train}
X_val = {"X_tab": X_tab_valid, "target": y_valid}
asha_scheduler = AsyncHyperBandScheduler(
time_attr="training_iteration",
metric="_metric/val_loss",
mode="min",
max_t=100,
grace_period=10,
reduction_factor=3,
brackets=1,
)
analysis = tune.run(
tune.with_parameters(training_function, X_train=X_train, X_val=X_val),
resources_per_trial={"cpu": 1, "gpu": 0},
progress_reporter=JupyterNotebookReporter(overwrite=True),
scheduler=asha_scheduler,
config=config,
callbacks=[WandbLoggerCallback(
project=config["wandb"]["project"],
api_key_file=config["wandb"]["api_key_file"],
log_config=True)],
)
# -
analysis.results
# Using Weights and Biases logging you can create [parallel coordinates graphs](https://docs.wandb.ai/ref/app/features/panels/parallel-coordinates) that map parametr combinations to the best(lowest) loss achieved during the training of the networks
#
# 
# local visualization of raytune reults using tensorboard
# %load_ext tensorboard
# %tensorboard --logdir ~/ray_results
| examples/12_HyperParameter_tuning_w_RayTune_n_WnB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
#
# # Create a 3D model of a Permo-Carboniferous Trough (PCT)
#
# Based on four seismic sections from the NAGRA report
# `NAGRA NTB 14-02 <https://www.nagra.ch/data/documents/database/dokumente/$default/Default\%20Folder/Publikationen/NTBs\%202014\%20-\%202015/d_ntb14-02\%20Dossier\%20I.pdf>`_ [1],
# we extracted interface and orientation points of main eras (paleozoic, mesozoic, cenozoic) and major graben faults.
# Data from these 2D sections are complemented with data from GeoMol 2019, e.g. base of the PCT, thrusts, and normal faults.
#
# The lithological units comprise the permo-carboniferous filling (paleozoic), Mesozoic, Tertiary strata, as well as the crystalline basement rocks. An important decision before building the geological model,
# is to define model units. Based on the purpose of the envisaged model, different units have to be defined. As the final result of this work will be an ensemble of advective heat-transport models,
# key paremeters for defining units are permeability, porosity, thermal conductivity of different geological layers. As part of the exploration work of nagra
# (National Cooperative for the Disposal of Radioactive Waste), regional and local hydrogeological models were constructed. The therein defined hydrostratigraphy provides the basis for defining the
# model units of this geological model. The regional hydrogeologic model is presented in the report
# `NAGRA NAB 13-23 <https://www.nagra.ch/data/documents/database/dokumente/$default/Default\%20Folder/Publikationen/NABs\%202004\%20-\%202015/e_nab13-023.pdf>`_ [2].
#
# With the regional model covering an area comprising all potential storage sites defined by nagra, local models were built as well. These models comprise a more detailed hydrostratigraphy.
#
# The potential storage site "<NAME>" is within our model area, thus we also consider the hydrostratigraphy defined in this local hydrogeological model presented in the report
# `NAGRA NAB 13-26 <https://www.nagra.ch/data/documents/database/dokumente/$default/Default\%20Folder/Publikationen/NABs\%202004%20-\%202015/e_nab13-026.pdf>`_ [3].
#
# The model comprises an area of 45 km x 32 km, in x- and y-direction, respectively. It extends down to a depth of 6 km, with reference sea level.
# This notebook demonstrates step-by-step how the model is generated within the open source modeling software `GemPy <https://www.gempy.org/>`_ [4].
# First, we will import libraries necessary to run this notebook:
#
# +
# Importing GemPy
import gempy as gp
# Import improved plotting features from GemPy
from gempy.plot import visualization_2d as vv
from gempy.plot import vista
# Importing auxilary libraries
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import matplotlib
matplotlib.rcParams['figure.figsize'] = (20.0, 10.0)
# -
# This example code was generated with Gempy-Version:
#
#
print(f"GemPy Version: {gp.__version__}")
# Initialize the model
# --------------------
# For modeling the PermoCarboniferous trough (**PCT**) in GemPy, we need to initialize a GemPy model object. This model object comprises multiple input data, such as interface points and orientations,
# which we previously stored in a `.csv` file. Further, we import the topography from a GeoTiff file.
# Conceptually, we create two models:
#
# . 1. With data of the the base of the PCT known
# . 2. With additional data where the base of the PCT is inferred
#
# The distinction of inferred vs. known locations of the PCT is based on GeoMol 2019, an update geological model of the Swiss Molasse Basin and adjacent areas. Known and inferred parts of the PCT in
# GeoMol can be seen `here <https://viewer.geomol.ch/webgui/gui2.php?viewHash=02171f57ee58a4082d3eb9cdc541c08b>`_.
#
# In this notebook, the user can choose whether only the "known" parts of the PCT base will be considered for modeling, or also the the inferred parts.
#
#
# string either "known" or "inferred" to switch between model data
#
#
# +
switch = "known"
if switch == 'known':
# Import data - NOT INFERRED
# Create a model instance
geo_model = gp.create_model('PCT_model')
# Initialize the model, set dimension and load interface and orientation data
gp.init_data(geo_model, [2640000, 2685000., 1240000., 1275000., -6000, 1000.], [50, 50, 50],
path_i = '../../../Editorial-Transitional-Heatflow/data/processed/GemPy/00_gempy_inputs/2021-06-02_interfaces_no_fault_horizon_reduced_graben_and_mandach.csv',
path_o = '../../../Editorial-Transitional-Heatflow/data/processed/GemPy/00_gempy_inputs/20201007_orientations_with_Jurathrust5_no_quat_meso_reduced2.csv')
geo_model.set_topography(source='gdal', filepath='../../../Editorial-Transitional-Heatflow/data/processed/GemPy/06_DTMs/DTM_200_for_GemPy_Model.tif')
elif switch == 'inferred':
# Import data - INFERRED
# Create a model instance
geo_model = gp.create_model('PCT_model_inferred')
# Initialize the model, set dimension and load interface and orientation data
gp.init_data(geo_model, [2640000, 2685000., 1240000., 1275000., -6000, 1000.], [50, 50, 50],
path_i = '../../data/processed/GemPy/00_gempy_inputs/20201005_interfaces_Jurathrust5_pct_inferred.csv',
path_o = '../../data/processed/GemPy/00_gempy_inputs/20201007_orientations_with_Jurathrust5_no_quat_meso_reduced2_pct_inferred.csv')
geo_model.set_topography(source='gdal', filepath='../../data/processed/GemPy/06_DTMs/DTM_200_for_GemPy_Model.tif')
# -
# To be coherent with existing geological models, e.g. geological cross-sections by nagra, we adapt the coloring for units according to
# `NTB 14-02 <https://www.nagra.ch/data/documents/database/dokumente/$default/Default\%20Folder/Publikationen/NTBs\%202014\%20-\%202015/d_ntb14-02\%20Dossier\%20I.pdf>`_ [5].
# For this, we create a color dictionary linking the units of the model to hex-color-codes.
#
#
# +
col_dict = {'basement': '#efad83',
'graben-fill': '#97ca68',
'Mittlerer-Muschelkalk': '#f9ee3a',
'Oberer-Muschelkalk': '#ffcf59',
'Keuper': '#ffe19f',
'Opalinuston': '#7f76b4',
'Dogger': '#b0ac67',
'Effinger-Schichten': '#47c4e2',
'Malm': '#92d2ec',
'USM': '#fbf379',
'OMM': '#fbf379',
'BIH-Basement-N': '#015482',
'Fault-south': '#4585a8',
'Fault_Basement_A': '#851515',
'Vorwald_Basement': '#b54343',
'Jurathrust5': '#5DA629',
'Mandach': '#408f09'}
geo_model.surfaces.colors.change_colors(col_dict)
# -
# ## Visualize the data distribution
# The following plot shows the different interface and orientation data loaded in the previous cell:
#
#
gp.plot_2d(geo_model, show_data=True, show_lith=False, show_results=False, direction='z', legend=False)
# The different colors in the plot represent the different model units. Circles represent the interface points, while arrows define the orientation of the respective surface in space.
#
# GemPy interpolates these input data in space using a universal co-kriging approach. Later on, we will set up the interpolator.
#
#
# ## Setting up Cross sections from the Nagra Report
#
# As stated before, next to GeoMol [6], we incorporate geological interpretations from four migrated seismic sections, the NAGRA report
# `NTB 14-02 <https://www.nagra.ch/data/documents/database/dokumente/$default/Default\%20Folder/Publikationen/NTBs\%202014\%20-\%202015/d_ntb14-02\%20Dossier\%20I.pdf>`_.
# For comparing the model results with the original interpretations, we define three cross sections in the model domain by specifying their start- and end-points and their resolution:
#
# set three sections which go roughly North South:
#
#
section_dict = {'section4_3':([2670826,1268793],[2676862,1255579],[100,100]),
'section4_4':([2649021,1267107],[2659842,1246715],[100,100]),
'section4_8':([2643284,1259358],[2680261,1268521],[100,100])}
geo_model.set_section_grid(section_dict)
# ## Display Model Information
#
# In the following, we will go through model construction step-by-step. As an overview, we display the different units (here called `surfaces`) included in the model.
# Note that also faults are surfaces within this model context. Currently, they are not marked as faults, and GemPy would treat them as the base of another geological model unit.
#
# To clarify, we model the base of a unit volume. That is, everything above the base surface is the respective unit, until the next base surface is reached.
# In total, our model comprises 17 `surfaces`. Everything beneath is filled with the 18th surface, called `basement`.
#
# ### Surfaces
# The majority of the structural features, i.e. normal- and thrust faults, are named following the names in GeoMol.
# Main features of the model is the asymetric graben system, with the major normal faults (:code:`Fault_Basement_A`, :code:`Fault-south`, :code:`BIH-Basement-N`),
# and the graben fill, which is not present beyond the graben shoulders, unless where it is inferred.
# This, as well as the stop of major normal faults beneath the mesozoic units (the base of :code:`Mittlerer-Muschelkalk`) are important considerations for the modeling process.
#
#
geo_model.surfaces
# ## Characteristics
# One characteristic seen in the table above, is that all surfaces are assigned to a :code:`series` called :code:`Default series`.
# A _series_ in GemPy indicates whether units should be interpolated using the same parameters. That is, all :code:`surfaces` within the same :code:`series` will be sub-parallel.
# Thus, surfaces have to be grouped into different series, depending on their geometry in space. For instance, sub-parallel layers of a sedimentary sequence should be grouped in the same series,
# while an unconformity, or a fault should be assorted to its own series.
#
# In this model, we group the majority of mesozoic and cenozoic units in one series, called :code:`Post_graben_series`. Only the mesozoic surface :code:`Mittlerer-Muschelkalk` will be assigned its own
# series, as it forms the basal detachement of the Jura Mountains. Palaeozoic graben sediments are also assigned its own series.
#
#
# Assign formations to series
gp.map_series_to_surfaces(geo_model,
{"Thrust_Mandach": 'Mandach',
"Thrust_Jura": 'Jurathrust5',
#"Thrust_Jura6": 'Jurathrust6', #('Jurathrust4', 'Jurathrust5', 'Jurathrust6'),
"Fault_north_series": 'Fault_Basement_A',
"Fault_south_series": 'Fault-south',
"Vorwald_series": 'Vorwald_Basement',
"BIH_series": 'BIH-Basement-N',
"Fault_north_series": 'Fault_Basement_A',
"Fault_south_series": 'Fault-south',
"Post_graben_series": ('OMM',
'USM',
'Malm',
'Effinger-Schichten',
'Dogger',
'Opalinuston',
'Keuper',
'Oberer-Muschelkalk'),
"Detachement": 'Mittlerer-Muschelkalk',
"Graben_series": 'graben-fill'},
remove_unused_series=True)
geo_model.surfaces
# ## Define Faults
# To distinguish between lithological units and faults, we have to assign which series are faults. Faults can be infinite, i.e. have the same displacement throughout the model space, or they can be
# finite, meaning displacement will be less towards the fault edges (which are defined by the extent of interface points used as input).
#
#
geo_model.set_is_fault(['Thrust_Mandach', 'Thrust_Jura', 'Fault_north_series',
'Fault_south_series', 'Vorwald_series', 'BIH_series'],
change_color=False)
geo_model.set_is_finite_fault(series_fault=['BIH_series', 'Vorwald_series'],
toggle=True)
# ## Bottom relation
# To set whether a surface is eroding or not, we can set a series' `bottom_relation`. Per default, it is set to `Erosion`, meaning the base of a younger surface (higher up in the stratigraphic pile)
# will cut through older surfaces. Setting the `bottom_relation` to `Onlap` will cause the opposite, i.e. younger surfaces stop on older ones.
# We set the _Graben_series_ to onlap, as most of it is only present in the graben, i.e. hanging wall of the normal faults, but not in the foot wall.
#
#
geo_model.set_bottom_relation(series=['Graben_series'], bottom_relation='Onlap')
# ## Define Fault relations
# With cross-cutting faults, we need to define fault relations, i.e. which fault stops at which. This is important, as some normal faults stop at others, e.g. :code:`BIH_Series` stops at
# :code:`Fault_south_series`. Fault relations are set in a matrix, where :code:`True` sets that one fault stops at the other. If set to :code:`False` (the default), faults cross-cut each other
# without any effects.
#
# Further, fault relations are used to define whether a fault displaces lithological series, or not. Per default, all faults displace the lithological series, but not other faults.
# This can be seen, if we plot the :code:`fault_relations` matrix:
#
#
geo_model.faults.faults_relations_df
# We know that faults do not affect all lithological series equally. For instance, thrusts will not affect the paleozoic sediments filling the graben.
# Just as the mesozoic units are not affected by the normal faults. Thus we set up a fault relation matrix, considering:
#
# - thrusts only affect Mesozoic units
# - normal faults only affect Basement, Graben_series
# - normal faults stop at thrusts
# We can update the fault relations by creating a boolean matrix of shape similar to :code:`faults_relations_df`, to assign which fault displaces which unit, etc. Then we use this
# boolean matrix to set fault relations using the :code:`set_fault_relation()` method.
#
#
fr = np.array([[False, False, False, False, False, False, True, False, False, False],
[False, False, False, True, False, False, True, False, False, False],
[False, False, False, False, True, False, False, True, True, True],
[False, False, False, False, False, True, False, False, True, True],
[False, False, False, False, False, False, False, True, True, True],
[False, False, False, False, False, False, False, False, True, True],
[False, False, False, False, False, False, False, False, False, False],
[False, False, False, False, False, False, False, False, False, False],
[False, False, False, False, False, False, False, False, False, False],
[False, False, False, False, False, False, False, False, False, False]])
geo_model.set_fault_relation(fr)
# Remember when we had a look at the input data and briefly mentioned the interpolator? We now set the interpolator function for the underlying co-kriging interpolation using theano:
#
#
gp.set_interpolator(geo_model,
compile_theano=True,
theano_optimizer='fast_compile',
verbose=[])
# ## Creating the model
# Now that we set the parameters and fault relations, it is time to start the modeling process. In Gempy, this is done using a single function :code:`gempy.comput_model` giving the prepared _geo_model_
# as input.
#
#
sol = gp.compute_model(geo_model, compute_mesh=True)
# For comparing model results with geological interpretations of the aforementioned seismic sections, we plot the model units on top of the seismic profiles.
# Profiles 4.3 and 4.4 (nomenclature is taken from [1]) strike across the graben axis, while profile 4.8 goes roughly along the graben.
#
# In the following plot, we model all profiles with the resulting geological grid, in the order from left to right: Profile 4.3, Profile 4.4, Profile 4.8.
#
#
gp.plot_2d(geo_model, section_names=list(section_dict), show_block=True, show_boundaries=False, show_data=False,
show_topography=True, show_results=True)
# ## References
# | [1]: <NAME>., and <NAME>.: Tektonische Karte des Nordschweizer Permokarbontrogs: Aktualisierung basierend auf 2D-Seismik und Schweredaten. Nagra Arbeitsbericht NAB 14-017, (2014).
# | [2]: <NAME>., <NAME>., <NAME>., & <NAME>.: Regional Hydrogeo-logical Model of Northern Switzerland. Nagra Arbeitsbericht NAB, 13-23, (2014).
# | [3]: <NAME>., <NAME>., <NAME>.: Hydrogeological model Jura Ost. Nagra Arbeitsbericht NAB, 13-26, (2014).
# | [4]: <NAME>., <NAME>., and <NAME>.: GemPy 1.0: Open-source stochastic geological modeling and inversion. Geoscientific Model Development, 12(1), (2019), 1. doi:http://dx.doi.org/10.5194/gmd-12-1-2019.
# | [5]: <NAME>., & <NAME>. (ed): Nagra technical report 14-02, geological basics-Dossier I-Introduction and summary; SGT Etappe 2: Vorschlag weiter zu untersuchender geologischer Standortgebiete mit zugehörigen Standortarealen für die Oberflächenanlage--Geologische Grundlagen--Dossier I--Einleitung und Zusammenfassung, (2014).
# | [6]: GeoMol Team (2015): GeoMol – Assessing subsurface potentials of the Alpine Foreland Basins for sustainable planning and use of natural resources – Project Report, 188 pp. (Augsburg, LfU).
#
| docs/source/Examples/Model_PCT_GemPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Scraping mit Python: Abschluss-Portfolio – Vergleich liberaler und konservativer Zeitungen/Medien in den USA
#
# In diesem Notebook soll eine deskriptiv-explorative, vergleichende Analyse zweier amerikanischer Zeitungen und der Themen, über die dort berichtet wird, durchgeführt werden.
#
# Folgendes Notebook gliedert sich wie folgt:
#
# **1. Hintergrund: welche Zeitungen wurden ausgewählt und auf was zielt die Analyse ab?**
#
# **2. Technische Durchführung, Datenakquise**
#
# **3. Explorative Analysen mittels erhaltener Daten**
#
# **4. Ein kurzes abschließendes Fazit**
# ## 1. Hintergrund: welche Zeitungen wurden ausgewählt und auf was zielt die Analyse ab?
#
# Zunächst wird geklärt, welches Erkenntnisinteresse diesem Projekt zugrunde liegt – woraus sich nachfolgend auch die Auswahl der beiden Medien ergibt.
#
# Es kann durchaus behauptet werden, dass die US-amerikanische politische Landschaft aktuell ein problematisches Bild abgibt. Gerade der Diskurs um "Fake News", der durch Donald Trumps Präsidentschaft sowie seine haltlosen Bemerkungen, dass ihm ein vermeintlicher Wahlsieg gegen Joe Biden bei der letzten Wahl durch gezielte Manipulation gestohlen worden wäre, trugen dazu bei. Die ohnehin starke Rivalität zwischen den konservativen Republikanern und den liberaleren Demokraten, die durchaus schon vor Trumps Präsidentschaft bestand, wurde jedoch durch eben diese verschärft: man bekommt als außenstehender Europäer immer mehr den Eindruck, es handele sich um regelrechte Grabenkämpfe – eine Metapher, die sich durch die traurigen Ereignisse des Sturms auf das US-Kapitol vom 6. Januar 2021 noch stärker aufdrängt.
#
# Und dieses Bild der amerikanischen Politik lässt sich zu gewissen Teilen sicher auch auf die Medienlandschaft der USA übertragen. So konnte man unter dem letzten Präsidenten wohl einen starken Anstieg in der Wichtigkeit von Blogs und anderen News-Quellen feststellen, mit denen seine Anhänger ihre "alternativen Fakten" einholten.
#
# Doch sind für vorliegendes Portfolio vor allem die traditionellen Printmedien von Interesse, genauer gesagt deren Online-Versionen. Es soll um die Frage gehen, ***welche Themen in diesen behandelt werden und wo zwischen traditionell liberalen und traditionell konservativen Zeitungen bzw. Nachrichtenagenturen Unterschiede bestehen.***
#
# Hier wäre beispielsweise anzunehmen, dass sozusagen aus der "Ideologie" in den konservativen Zeitungen das "America first"-Prinzip vorherrschend ist und daher viele bzw. hauptsächlich Artikel im Inland- oder gar im Sicherheits-Ressort publiziert werden. Dem gegenüber könnte angenommen werden, dass in den liberaleren Zeitungen wohl eher die Prämisse eines "offeneren Amerikas" vorherrscht und hier im Gegenteil zu konservativen Zeitungen auch Beiträge aus den Ressorts "Global" und "International" oder auch "Wirtschaft" veröffentlicht werden.
# Diese Vermutungen sollen mittels dieses Portfolios einer ersten explorativen Überprüfung unterzogen werden, damit sie durch Fakten untermauert werden können: welche Themen werden also wirklich von den Nachrichtenportalen aufgegriffen?
#
# Zwecks der dafür genutzten Medien soll auf der "liberalen Seite" auf die [New York Times](https://www.nytimes.com) zurückgegriffen werden. Als konservatives Gegenstück wurde sich für die [Washington Times](https://www.washingtontimes.com) entschieden. Alternativ würde hier auch das News-Portal [Fox News](https://www.foxnews.com/) als eher konservatives Beispiel fungieren können – in Ermangelung von Informationen über eine möglicherweise verfügbare API wurden beide News-Portale kontaktiert, da diese den Prozess der Informationsgewinnung wesentlich vereinfachen würde. Zur Datenbeschaffung soll jedoch später unter Punkt 2 mehr Information gegeben werden. Diese Entscheidungen wurden beispielsweise aufgrund von [Rankings der University of Michigan in Zusammenarbeit mit AllSides](https://guides.lib.umich.edu/c.php?g=637508&p=4462444) getroffen.
#
# AllSides ist ein eigenes Nachrichten-Portal, welches wie der Name schon sagt versucht, "beide Seiten" des (nachrichten)politischen Spektrums zu erfassen und somit ein möglichst umfassendes Nachrichtenbild zu gewährleisten. Dort werden Seiten mit einem eigenen [Ranking](https://www.allsides.com/media-bias/media-bias-ratings) erfasst, auf welches von Seiten der University of Michigan und auch von Seiten dieses Portfolios zurückgegriffen wurde.
# Es wurde dabei bewusst nur auf die Kategorien "lean left" und "lean right" geblickt, da u. a. auch Nachrichtenagenturen wie Breitbart erfasst wurden, welche von vorliegender Arbeit nicht wirklich als seriöse "traditionelle Nachrichtenquelle" erachtet wird.
# # 2. Technische Durchführung, Datenakquise
# Im zweiten Schritt sollen dann die Daten von den ausgewählten Medien-APIs gezogen werden. Dieser Paragraph soll kurz als Vorabklärung zu den APIs dienen - etwaige geschriebene Funktionen werden dann jedoch an Ort und Stelle näher erläutert oder mit ausreichend erklärenden Kommentaren versehen.
#
# Zunächst zur **API der New York Times**. Hier konnte ganz einfach ein Zugang und ein Key angefragt werden, welcher dann sofort einsetzbar war. Hier können verschiedene Zugänge gewählt werden, besonders interessant für den hier vorliegenden Anwendungsfall sind jedoch *Article Archive* (hier kann zeitlich gesucht werden, nach Monaten) und *Article Search* (hier kann nach einem Begriff gesucht werden, zu dem dann Artikel gezeigt werden).
#
# Informationen zur Washington Times und zu Fox News folgen hier noch.
#
# Für die [Currents API](https://www.currentsapi.services/en) wurde ebenfalls ein API-Key beantragt, welcher auch nutzbar ist. Diese API versucht, über verschiedenste Nachrichtenquellen hinweg den Zugang über eine Schnittstelle zu gewährleisten – sie ist sozusagen als Meta-API zu sehen. Sollten für die einzelnen Portale keine Keys zu erhalten sein, dann wird auf Currents zurückgegriffen werden.
# +
import requests
import datetime
import pandas as pd
import seaborn as sea
from pprint import pprint
########################
myAPI = '<KEY>'
# +
# hier könnte IHRE funktion stehen!
#def req_currents(keywords, )
# + tags=["outputPrepend"]
# keyword search
latest = ('https://api.currentsapi.services/v1/latest-news?domain=washingtontimes.com&' +
'apiKey=<KEY>')
latest_resp = requests.get(latest)
pprint(latest_resp.json())
# + tags=["outputPrepend"]
# keyword search with domain specified
keywords = 'Biden'
search = (f"https://api.currentsapi.services/v1/search?domain=washingtontimes.com&keywords={keywords}&" +
"apiKey=<KEY>")
search_res = requests.get(search)
data = search_res.json()
pprint(data)
# +
def keys_currents(domain, keywords):
# definieren, dass die api mit such-funktion angesteuert wird
url = "https://api.currentsapi.services/v1/search?"
# welche domain soll durchsucht werden? (aufpassen: spezifizieren ohne web-protokoll und ohne "www.")
search_domain = f"domain={domain}&"
# mode == welches keyword mit meinem api-key gesucht werden soll
mode = f"keywords={keywords}&" + "apiKey=<KEY>"
# was passiert mit den resultaten davon:
gesamt = url + search_domain + mode
ergebnis = requests.get(gesamt)
# am ende ausgabe als json
daten = ergebnis.json()
return pprint(daten)
# + tags=["outputPrepend"]
nyt_news_keywords = keys_currents(domain='nytimes.com', keywords='Biden')
pd.DataFrame(nyt_news_keywords)
# + tags=["outputPrepend"]
keys_currents(domain='washingtontimes.com', keywords='Biden')
# -
def latest_currents(domain):
# definieren, dass die api mit such-funktion angesteuert wird
url2 = "https://api.currentsapi.services/v1/latest-news?"
# welche domain soll durchsucht werden? (aufpassen: spezifizieren ohne web-protokoll und ohne "www.")
search_domain2 = f"domain={domain}&"
# mode == diesmal nur der api-key eingegeben
api = "apiKey=<KEY>"
# was passiert mit den resultaten davon:
gesamt2 = url2 + search_domain2 + api
ergebnis2 = requests.get(gesamt2)
# am ende ausgabe als json
daten2 = ergebnis2.json()
daten_voll = pd.DataFrame(daten2["news"])
return(daten_voll)
# + tags=["outputPrepend"]
nyt = latest_currents(domain='nytimes.com')
# -
nyt["description"][0]
# + tags=["outputPrepend"]
wtimes = latest_currents(domain='washingtontimes.com')
# -
wtimes["description"][0]
foxnews = latest_currents(domain='foxnews.com')
foxnews["description"][1]
# # Versuche mit NLTK an NLP
import numpy
import re
from nltk.tokenize import word_tokenize
from nltk import pos_tag
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from sklearn.preprocessing import LabelEncoder
from collections import defaultdict
from nltk.corpus import wordnet as wn
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn import model_selection, naive_bayes, svm
from sklearn.metrics import accuracy_score
| abschlussnotebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content-dl/blob/main/tutorials/W2D1_ConvnetsAndRecurrentNeuralNetworks/student/W2D1_Tutorial3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Tutorial 3: Introduction to RNNs
# **Week 2, Day 1: Convnets And Recurrent Neural Networks**
#
# **By Neuromatch Academy**
#
# __Content creators:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# __Content editors:__ <NAME>
#
# __Production editors:__ <NAME>, <NAME>
#
# *Based on material from:* <NAME>, <NAME>, <NAME>, <NAME>
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# ---
# # Tutorial Objectives
# At the end of this tutorial, we will be able to:
# - Understand the structure of a Recurrent Neural Network (RNN)
# - Build a simple RNN model
#
#
# + cellView="form"
# @title Tutorial slides
# @markdown These are the slides for the videos in this tutorial
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/5asx2/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# -
# ---
# # Setup
# + cellView="form"
# @title Install dependencies
# !pip install livelossplot --quiet
# !pip install unidecode
# +
# Imports
import time
import math
import torch
import random
import numpy as np
import torch.nn as nn
import matplotlib.pyplot as plt
from torch.autograd import Variable
from tqdm.notebook import tqdm
# + cellView="form"
# @title Figure settings
import ipywidgets as widgets # interactive display
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/content-creation/main/nma.mplstyle")
plt.rcParams["mpl_toolkits.legacy_colorbar"] = False
import warnings
warnings.filterwarnings("ignore", category=UserWarning, module="matplotlib")
# + cellView="form"
# @title Helper functions
# https://github.com/spro/char-rnn.pytorch
import unidecode
import string
# Reading and un-unicode-encoding data
all_characters = string.printable
n_characters = len(all_characters)
def read_file(filename):
file = unidecode.unidecode(open(filename).read())
return file, len(file)
# Turning a string into a tensor
def char_tensor(string):
tensor = torch.zeros(len(string)).long()
for c in range(len(string)):
try:
tensor[c] = all_characters.index(string[c])
except:
continue
return tensor
# Readable time elapsed
def time_since(since):
s = time.time() - since
m = math.floor(s / 60)
s -= m * 60
return '%dm %ds' % (m, s)
def generate(decoder, prime_str='A', predict_len=100, temperature=0.8,
device='cpu'):
hidden = decoder.init_hidden(1)
prime_input = Variable(char_tensor(prime_str).unsqueeze(0))
hidden = hidden.to(device)
prime_input = prime_input.to(device)
predicted = prime_str
# Use priming string to "build up" hidden state
for p in range(len(prime_str) - 1):
_, hidden = decoder(prime_input[:,p], hidden)
inp = prime_input[:,-1]
for p in range(predict_len):
output, hidden = decoder(inp, hidden)
# Sample from the network as a multinomial distribution
output_dist = output.data.view(-1).div(temperature).exp()
top_i = torch.multinomial(output_dist, 1)[0]
# Add predicted character to string and use as next input
predicted_char = all_characters[top_i]
predicted += predicted_char
inp = Variable(char_tensor(predicted_char).unsqueeze(0))
inp = inp.to(device)
return predicted
# + cellView="form"
# @title Set random seed
# @markdown Executing `set_seed(seed=seed)` you are setting the seed
# for DL its critical to set the random seed so that students can have a
# baseline to compare their results to expected results.
# Read more here: https://pytorch.org/docs/stable/notes/randomness.html
# Call `set_seed` function in the exercises to ensure reproducibility.
import random
import torch
def set_seed(seed=None, seed_torch=True):
if seed is None:
seed = np.random.choice(2 ** 32)
random.seed(seed)
np.random.seed(seed)
if seed_torch:
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
print(f'Random seed {seed} has been set.')
# In case that `DataLoader` is used
def seed_worker(worker_id):
worker_seed = torch.initial_seed() % 2**32
np.random.seed(worker_seed)
random.seed(worker_seed)
# + cellView="form"
#@title Set device (GPU or CPU). Execute `set_device()`
# especially if torch modules used.
# inform the user if the notebook uses GPU or CPU.
def set_device():
device = "cuda" if torch.cuda.is_available() else "cpu"
if device != "cuda":
print("WARNING: For this notebook to perform best, "
"if possible, in the menu under `Runtime` -> "
"`Change runtime type.` select `GPU` ")
else:
print("GPU is enabled in this notebook.")
return device
# -
SEED = 2021
set_seed(seed=SEED)
DEVICE = set_device()
# ---
# # Section 1: Recurrent Neural Networks (RNNs)
# + cellView="form"
# @title Video 1: RNNs
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1L44y1m7PP", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"PsZjS125lLs", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# RNNs are compact models that operate over timeseries, and have the ability to remember past input. They also save parameters by using the same weights at every time step. If you've heard of Transformers, those models don't have this kind of temporal weight sharing, and so they are *much* larger.
#
# The code below is adapted from [this github repository](https://github.com/spro/char-rnn.pytorch).
# + cellView="form"
# @title Run Me to get the data
# !wget --output-document=/content/sample_data/twain.txt https://raw.githubusercontent.com/amfyshe/amfyshe.github.io/master/twain.txt
# -
# RNN
# https://github.com/spro/char-rnn.pytorch
class CharRNN(nn.Module):
def __init__(self, input_size, hidden_size, output_size, model="gru", n_layers=1):
super(CharRNN, self).__init__()
self.model = model.lower()
self.input_size = input_size
self.hidden_size = hidden_size
self.output_size = output_size
self.n_layers = n_layers
self.encoder = nn.Embedding(input_size, hidden_size)
if self.model == "gru":
self.rnn = nn.GRU(hidden_size, hidden_size, n_layers)
elif self.model == "lstm":
self.rnn = nn.LSTM(hidden_size, hidden_size, n_layers)
elif self.model == "rnn":
self.rnn = nn.RNN(hidden_size, hidden_size, n_layers)
self.decoder = nn.Linear(hidden_size, output_size)
def forward(self, input, hidden):
batch_size = input.size(0)
encoded = self.encoder(input)
output, hidden = self.rnn(encoded.view(1, batch_size, -1), hidden)
output = self.decoder(output.view(batch_size, -1))
return output, hidden
def init_hidden(self, batch_size):
if self.model == "lstm":
return (Variable(torch.zeros(self.n_layers, batch_size, self.hidden_size)),
Variable(torch.zeros(self.n_layers, batch_size, self.hidden_size)))
return Variable(torch.zeros(self.n_layers, batch_size, self.hidden_size))
# This next section of code takes care of training the RNN on several of <NAME>'s books. In this short section, we won't dive into the code, but you'll get to learn a lot more about RNNs in a few days! For now, we are just going to observe the training process.
#
# One cool thing about RNNs is that they can be used to _generate_ language based on what the network sees during training. As the network makes predictions, instead of confirming of those predictions are correct against some training text, we just feed them back into the model as the next observed token. Starting from a random vector for the hidden state, we can generate many original sentences! And what the network generates will reflect the text it was trained on.
#
# Let's try it! Run the code below. As the network trains, it will output samples of generated text every 25 epochs. Notice that as the training progresses, the model learns to spell short words, then learns to string some words together, and eventually can produce meaningful sentences (sometimes)! Keep in mind that this is a relatively small network, and doesn't employ some of the cool things you'll learn about later in the week (e.g. LSTMs, though you can change that in the code below by changing the value of the `model` variable if you wish!)
#
# After running the model, and observing the output, get together with your pod, and talk about what you noticed during training. Did your network produce anything interesting? Did it produce anything characteristic of Twain?
#
# **Note:** training for the full 2000 epochs is likely to take a while, so you may need to stop it before it finishes.
#
# **Important:** You can increase the number of epochs `n_epochs` to 2000.
# + cellView="code"
# https://github.com/spro/char-rnn.pytorch
batch_size = 50
chunk_len = 200
model = "rnn" # other options: `lstm`, `gru`
# hyperparams
n_layers = 2
hidden_size = 200
n_epochs = 1000 # initial was set to 2000
learning_rate = 0.01
print_every = 25
def train(inp, target, device='cpu'):
hidden = decoder.init_hidden(batch_size)
decoder.zero_grad()
loss = 0
for c in range(chunk_len):
output, hidden = decoder(inp[:,c].to(device), hidden.to(device))
loss += criterion(output.view(batch_size, -1), target[:,c])
loss.backward()
decoder_optimizer.step()
return loss.item() / chunk_len
file, file_len = read_file('/content/sample_data/twain.txt')
def random_training_set(chunk_len, batch_size, device='cpu'):
inp = torch.LongTensor(batch_size, chunk_len).to(device)
target = torch.LongTensor(batch_size, chunk_len).to(device)
for bi in range(batch_size):
start_index = random.randint(0, file_len - chunk_len - 1)
end_index = start_index + chunk_len + 1
chunk = file[start_index:end_index]
inp[bi] = char_tensor(chunk[:-1])
target[bi] = char_tensor(chunk[1:])
inp = Variable(inp)
target = Variable(target)
return inp, target
decoder = CharRNN(
n_characters,
hidden_size,
n_characters,
model=model,
n_layers=n_layers,
).to(DEVICE)
decoder_optimizer = torch.optim.Adagrad(decoder.parameters(), lr=learning_rate)
criterion = nn.CrossEntropyLoss()
start = time.time()
all_losses = []
loss_avg = 0
print("Training for %d epochs..." % n_epochs)
for epoch in tqdm(range(1, n_epochs + 1), position=0, leave=True):
loss = train(*random_training_set(chunk_len, batch_size, DEVICE), device=DEVICE)
loss_avg += loss
if epoch % print_every == 0:
print('[%s (%d %d%%) %.4f]' % (time_since(start), epoch,\
epoch / n_epochs * 100, loss))
print(f"{generate(decoder, prime_str='Wh', predict_len=100, device=DEVICE)}\n")
# -
# Now you can generate more examples using a trained model. Recall that `generate` takes the mentioned below arguments arguments to work:
#
# ```python
# generate(decoder, prime_str='A', predict_len=100, temperature=0.8, device='cpu')
# ```
#
# Try it by yourself
print(f"{generate(decoder, prime_str='Wh', predict_len=100, device=DEVICE)}\n")
# ---
# # Section 2: Power consumption in Deep Learning
#
# Training NN models can be incredibly costly, both in actual money but also in power consumption.
# + cellView="form"
# @title Video 2: Carbon Footprint of AI
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1My4y1j7HJ", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"as6C334LmRs", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# Take a few moments to chat with your pod about the following points:
# * Which societal costs of training do you find most compelling?
# * When is training an AI model worth the cost? Who should make that decision?
# * Should there be additional taxes on energy costs for compute centers?
# ## Exercise 2: Calculate the carbon footprint that your pod generated today.
# You can use this [online calculator](https://mlco2.github.io/impact/#compute)
# ---
# # Summary
#
# What a day! We've learned a lot! The basics of CNNs and RNNs, and how changes to architecture that allow models to parameter share can greatly reduce the size of the model. We learned about convolution and pooling, as well as the basic idea behind RNNs. To wrap up we thought about the impact of training large NN models.
| tutorials/W2D1_ConvnetsAndRecurrentNeuralNetworks/student/W2D1_Tutorial3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Proposed Course Schedule
#
# - Meetings 2 through 16 (7 weeks) are programming focused.
# - By end of meeting 9 would have enough material to build a prediction engine to compute roots of numbers using reduced instruction sets (multiply, divide, add, subtract, compare, iterate) as a computational thinking exercise.
# - By end of meeting 17 would have enough material to visually fit data models (curves) to observations (markers)
# <table>
# <tr>
# <th> Lecture </th>
# <th> Laboratory </th>
# </tr>
# <tr>
# <td>
#
# 0. introduction
# - syllabus
# - computational thinking
# - jupyterlab/computing environment
#
# 1. problem solving
# - decomposition
# - examples
#
# </td>
# <td>
#
# ```c++
# int foo() {
# int x = 4;
# return x;
# }
# ```
#
# </td>
# </tr>
# </table>
# 0. introduction
# - syllabus
# - computational thinking
# - jupyterlab/computing environment
#
# 1. problem solving
# - decomposition
# - examples
#
# 2. variables, data type
# - integer
# - float
# - string
#
# 3. expressions
# - assignment
# - evaluating arithmetic expressions
# - simple output: print()
#
# 4. simple input
# - replacing assignment with input()
# - reading from an existing file
#
# 5. mutable data structures
# - lists
# - tuples
# - name;position;value
#
# 6. immutable data structures
# - dictionaries
# - name;position;value
#
# 7. algorithm building blocks
# - sequence
# - selection
# - repetition
#
# 8. selection structures
# - Inline IF
# - IF ... ELSE block
# - nested if structures
#
# 9. count controlled repetition
# - FOR structures
# - selection based loop exit/skip
# - nested loop structures
#
# 10. sentinel controlled repetition
# - WHILE structures
# - selection based loop exit/skip
# - nested loop structures
#
# 11. file manipulation
# - write to an existing file
# - create new file
# - get a file from a URL
#
# 12. functions
# - intrinsic
# - external (core: e.g. math)
# - user-defined; concept of a prototype function, variable scope
#
# 13. external modules
# - saving a user-defined function to a file
# - importing the file
# - running a file as a script
#
# 14. array processing
# - arrays:a type of list
# - matrix arithmetic
# - scalars, vectors, matrix
# - linear algebra using array
# - linear algebra using using numpy
# - solve linear system in numpy
#
# 15. database processing
# - records,fields
# - unique identifiers/keys
# - create, read, update, delete
# - database processing with pandas
#
# 16. visual display of data
# - plot types
# - plot uses
# - plot conventions
# - building plots with matplotlib
#
# 17. data models and visualization
# - interpolation using lagrange polynomials
# - fitting an arbitrary function (trial-and-error)
# - fitting a polynomial (linear system)
#
# 18. title
# - subtopic
# - subtopic
# - subtopic
#
# 19. title
# - subtopic
# - subtopic
# - subtopic
#
# 20. title
# - subtopic
# - subtopic
# - subtopic
#
# 21. title
# - subtopic
# - subtopic
# - subtopic
#
# 22. title
# - subtopic
# - subtopic
# - subtopic
#
# 23. title
# - subtopic
# - subtopic
# - subtopic
#
# 24. title
# - subtopic
# - subtopic
# - subtopic
#
# 25. title (last lesson before exam 3)
# - subtopic
# - subtopic
# - subtopic
#
# 26. title (not assessed - use for project presentations)
# - subtopic
# - subtopic
# - subtopic
#
# 27. title (not assessed - use for project presentations)
# - subtopic
# - subtopic
# - subtopic
#
#
#
#
#
| course-outline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import uproot
import awkward
import numpy as np
import os
from scipy.sparse import csr_matrix, find
from scipy.spatial import cKDTree
from tqdm import tqdm_notebook as tqdm
from graph import SparseGraph, make_sparse_graph, \
save_graph, save_graphs, load_graph, \
load_graphs, make_sparse_graph, graph_from_sparse, \
draw_sample_validation, draw_sample3d, Graph
from preprocessing import make_graph_xy, make_graph_etaphi, make_graph_knn, make_graph_kdtree, make_graph_noedge
preprocessing_algo = make_graph_noedge
#preprocessing_algo = make_graph_etaphi
#grouping_algo = make_graph_knn
#preprocessing_args= dict(r = 0.07)
#preprocessing_args= dict(k=4)
#layer_norm = 150
#fname = '../../../data/D41/photon_flatE/FlatRandomEGunProducer_sitong_20190703/partGun_PDGid22_x1000_E2.0To100.0_NTUP_1.root'
#fname = './partGun_PDGid22_x1000_E2.0To100.0_NTUP_10.root'
fname = '../../data/ntup/partGun_PDGid15_x1000_Pt3.0To100.0_NTUP_1.root'
test = uproot.open(fname)['ana']['hgc']
# %load_ext autoreload
# %autoreload 2
# +
#example of generating a binary ground-truth adjacency matrix
#for both endcaps in all events for all clusters
#truth is now that hits in adjacent layers are connected
#and so are hits in the same layer within delta-R < 2
arrays = test.arrays([b'simcluster_hits_indices'])
rechit = test.arrays([b'rechit_x',b'rechit_y', b'rechit_z', b'rechit_eta', b'rechit_phi',
b'rechit_layer',b'rechit_time',b'rechit_energy'])
NEvents = rechit[b'rechit_z'].shape[0]
rechit[b'rechit_x'].content[rechit[b'rechit_z'].content < 0] *= -1
sim_indices = awkward.fromiter(arrays[b'simcluster_hits_indices'])
valid_sim_indices = sim_indices[sim_indices > -1]
def get_features(ievt,mask):
x = rechit_x[ievt][mask]
y = rechit_y[ievt][mask]
layer = rechit_layer[ievt][mask]
time = rechit_time[ievt][mask]
energy = rechit_energy[ievt][mask]
return np.stack((x,y,layer,time,energy)).T
for ievt in tqdm(range(NEvents),desc='events processed'):
#make input graphs
#pos_graph = preprocessing_algo(rechit, valid_sim_indices, ievt = ievt, mask = rechit[b'rechit_z'][ievt] > 0,
# layered_norm = layer_norm, algo=grouping_algo, preprocessing_args=preprocessing_args)
#neg_graph = preprocessing_algo(rechit, valid_sim_indices, ievt = ievt, mask = rechit[b'rechit_z'][ievt] < 0,
# layered_norm = layer_norm, algo=grouping_algo, preprocessing_args=preprocessing_args)
pos_graph = preprocessing_algo(rechit, valid_sim_indices, ievt = ievt, mask = rechit[b'rechit_z'][ievt] > 0)
neg_graph = preprocessing_algo(rechit, valid_sim_indices, ievt = ievt, mask = rechit[b'rechit_z'][ievt] < 0)
#write the graph and truth graph out
outbase = fname.split('/')[-1].replace('.root','')
outdir = "/".join(fname.split('/')[:-2]) + "/npz/" + outbase
if not os.path.exists( outdir):
os.makedirs(outdir)
#graph = make_sparse_graph(*pos_graph)
#save_graph(graph, '%s/%s_hgcal_graph_pos_evt%d.npz'%(outdir,outbase,ievt))
save_graph(pos_graph, '%s/%s_hgcal_graph_pos_evt%d.npz'%(outdir,outbase,ievt))
#graph = make_sparse_graph(*neg_graph)
#save_graph(graph, '%s/%s_hgcal_graph_neg_evt%d.npz'%(outdir,outbase,ievt))
save_graph(neg_graph, '%s/%s_hgcal_graph_neg_evt%d.npz'%(outdir,outbase,ievt))
# +
ievt = 0
#g1sparse = load_graph('%s/%s_hgcal_graph_pos_evt%d.npz'%(outdir,outbase,ievt))
#g1 = graph_from_sparse(g1sparse)
g1 = load_graph('%s/%s_hgcal_graph_pos_evt%d.npz'%(outdir,outbase,ievt), graph_type=Graph)
#g1 = load_graph('partGun_PDGid13_x1000_Pt3.0To100.0_NTUP_1_hgcal_graph_neg_evt0.npz')
draw_sample_validation(g1.X,g1.Ri,g1.Ro,g1.y,
sim_list=g1.simmatched,
skip_false_edges=False)
# +
ievt = 2
#g2sparse = load_graph('%s/%s_hgcal_graph_pos_evt%d.npz'%(outbase,outbase,ievt))
#g2 = graph_from_sparse(g2sparse)
g2 = load_graph('%s/%s_hgcal_graph_pos_evt%d.npz'%(outdir,outbase,ievt), graph_type=Graph)
draw_sample_validation(g2.X,g2.Ri,g2.Ro,g2.y,
sim_list=g2.simmatched)
# -
| notebooks/graph_generation/make_graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py39]
# language: python
# name: conda-env-py39-py
# ---
# # "Five Tips to Customize fastpages"
# > "Create a personalized blog hosted free on Github."
#
# - toc: false
# - branch: master
# - image: images/post_images/logo.png
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [fastpages, jupyter]
# The fastpages platform is an excellent option for creating your own blog, hosted free on Github. The platform supports Jupyter notebooks, Word documents and Markdown, which is right in the wheelhouse for many analysts, data scientists and programmers. My knowledge of HTML and website development is limited and I've just recently created my first blog on the fastpages platform. I discovered the learning curve to customize the platform can be steep, but I enjoy digging into new technology. I thought it would be beneficial to assemble a post to help others in their journey.
#
# The basic setup for [fastpages](https://github.com/fastai/fastpages) is straightforward, so I'm not going to spend any time on that here. If you are reading this post, you have probably already setup a blog on the fastpages platform and searching for ways to personalize it. In this post, I will review some simple code that you can easily implement to personalize your blog.
#
# On a side note, I will be using [Visual Studio Code](https://code.visualstudio.com/) to modify the code in the following examples. If your using another editor, the process may be slightly different.
# ---
# ## Add a Unique Title Font
# Adding a unique title font is one of the fastest ways to customize your blog. Personally, I prefer the informal charm of Rock Salt, but there are hundreds of other choices available from [Google Fonts](https://fonts.google.com/). If you desire something beyond the standard flavor, you can check out their offerings by following the link above.
#
# Once you have made your selection, there are only a few lines of code required to make the changes. I'll use the Rock Salt example from my blog, to demonstrate the implementation.
#
#
# **Select a title font from [Google Fonts](https://fonts.google.com/)**
#
# - Click on the icon in the top right hand corner to view your selected families
#
# - Select the @import option and copy the corresponding code such as:
#
# ```
# @import url('https://fonts.googleapis.com/css2?family=Roboto:wght@100&family=Rock+Salt&display=swap');
# ```
#
# **Open the repository and navigate to the `_sass\minima` directory**
#
# - Open the `custom-styles.scss` file
#
# **Paste the code into the `custom-styles.scss` file**
#
# ```
# /* Import Rock Salt Font for Title */
# @import url('https://fonts.googleapis.com/css2?family=Rock+Salt&display=swap');
# ```
#
# **Update the font-family specified for the `site-title` in the `custom-styles.scss` file**
#
# ```
# /*Change Site Title Font Color and Size*/
# .site-title, .site-title:visited {
# font-family: 'Rock Salt';
# color: #f00;
# font-size: 275%;
# }
# ```
#
# **Save your updates (Ctrl + S) and commit the changes**
#
#
# Be aware that sometime it takes few minutes for the updates to flow through to your website, even after all of the workflows have completed successfully. So if your changes are not implemented immediately, sometimes all we need is just a little patience.
#
# There is a screen shot below to display the new code that should reside in the `custom-styles.scss` file, if you have followed the directions correctly. Many of the customizations I will discuss require accessing the `custom-styles.scss` file, so you should become very familiar with the `_sass\minima` directory.
#
# As a sidenote, you can also see the code required to customize the site-header in the image below. The size and borders of the header can be easily adjusted to accommodate the new font.
# 
# ---
# ## Create Additional Pages
# The backbone of the fastpages platform is the index page. There is a section for summary content beneath the header, followed by another section with links to posts created in Jupyter notebook, Markdown, or Word. Within the header there are navigation links to three other pages; About Me, Search and Tags. If you would like to create additional pages with a navigation link in the header, the process is simple.
#
# **Open the repository and navigate to the `_pages` directory**
#
# **Create a copy of the `about.md` file**
# - Copy and paste into the `_pages directory`
# - Rename the file (e.g. Another Page.md)
#
#
# **Update the Front Matter**
# - Provide a new title and permalink
# - Add any new content you like with Markdown under the Front Matter (i.e. below the ---)
# - Here is a great resource if you are unfamiliar with [Markdown](https://www.markdownguide.org/cheat-sheet/)
#
# **Save your updates (Ctrl + S) and commit the changes**
#
# If you followed these steps correctly, a new page will be created for your blog. The navigation links in the header will update automatically to reflect the changes.
# 
# ---
# ## Create a Custom Favicon
# If your browsing behavior resembles mine, you probably have links to all your favorite websites bookmarked in your browser. A favicon is a shortcut icon you can save in your bookmarks toolbar, which is more compact and often preferable to text. The fastpages platform comes complete with a favicon, which is a nice touch. If you want to create a custom favicon for your blog, the process is simple.
#
# The favicon for the fastpages platform is saved as an ICO file in the `images` directory. Simply create a new favicon image and save it with the same name as the original file. The image dimensions for the original file is 25px x 35px, so aim for an image file with similar dimensions. If you already have an image, fantastic. Otherwise, I would recommend heading over to [favicon.io](https://favicon.io/) to create a custom favicon from a PNG image, text or even an emoji. 👍
#
# Here is a breakdown of the process:
#
# **Create an ICO file with the new favicon image**
#
#
# **Save the new file to the `images` directory**
# - Rename the file `favicon.ico`
# - Agree to replace the existing file
#
# **Save your updates (Ctrl + S) and commit the changes**
#
# That is all it takes to create a custom favicon for your blog.
# 
# ---
# ## Add Images to Jupyter Notebook
# The ability to quickly generate a blog post with Jupyter notebook is part of the allure of the fastpages platform. Initially, I struggled while attempting to post a notebook containing an image pasted in a cell. There are many ways to add an image to Jupyter notebook, but if the correct protocols are not observed the post conversion process for fastpages will fail. I've reviewed several methods for posting images and here is the easiest routine I have found to work consistently.
#
# **Create a new folder to organize image files**
# - Use the images directory (e.g. `blog\images\post_images`)
#
#
# **Navigate to the repository on Github and upload the image file to the new directory**
#
# - Click on Add file, then Upload files
#
# - Select your file and upload, then Commit changes
#
# - Open the new image file and right click to copy image link
#
#
# **Paste the link to the image within a markdown cell**
#
# - Use the following syntax to display the image
#
# ```
# 
# ```
#
# **Save your updates (Ctrl + S) and commit the changes**
#
#
# If you have found a better method that works consistently, please leave a comment below. I'm a big fan of consistency and efficiency. If something better comes along, I'll be sure to edit this post.
# 
# ---
# ## Eliminate the Footer
# <NAME> proclaimed "Simplicity is the ultimate sophistication". If you are seeking a simple and elegant solution to remove the footer content, there are only two lines of code required(the comment is optional). Simply copy and paste the following lines of code to the `custom-styles.scss` file in the `_sass\minima` directory.
#
#
#
# ```
# /* --- The following Code Removes the Footer --- */
# .site-footer {
# display: none;}
# ```
#
# Once this simple update is complete, save the update (Ctrl + S) and commit the changes.
#
# Voila... the footer is gone.
# ---
# The options for personalizing your blog are nearly infinite, but that is a wrap for this post. You can check out more of the customizations I have implemented by reviewing the [repository](https://github.com/awault/blog) for my blog on Github. If you have made it this far, thank you for taking the time, I hope you found this post helpful. If you noticed any errors or have any suggestions, please leave a comment.
#
# AA
| _notebooks/2022-01-24-Customizefastpages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib
import numpy as np
import pandas as pd
from collections import Counter
import warnings
warnings.filterwarnings('ignore')
# ### Read data
# All other countries have the same attributes as Canada. So I just analyse this single CSV file.
CAvideos_df = pd.read_csv("./dataset/youtube_trading/CAvideos.csv")
CAvideos_df.head(5)
# ### Data reviews
# Although attribute 'category_id' is an integer type, but literally, only attribute 'views', 'likes', 'dislikes', 'comment_count' belong to numeric attribute.
CAvideos_df.info()
CAvideos_df.describe()
# #### As usual, I will present the frequency of each category in each nominal attribute.
CAvideos_df['video_id'].value_counts()
CAvideos_df['title'].value_counts()
CAvideos_df['channel_title'].value_counts()
CAvideos_df['category_id'].value_counts()
CAvideos_df['tags'].value_counts()
# #### For numeric attributes, five-number summary and the count of null value will be presented.
# > views: Minimum:733 Q1:143902 Median:371204 Q3:963302 Maximum:1.378431e+08
#
# > likes: Minimum:0 Q1:2191 Median:8780 Q3:28717 Maximum:5053338
#
# > dislikes: Minimum:0 Q1:99 Median:303 Q3:950 Maximum:1602383
#
# > comment_count: Minimum:0 Q1:417 Median:1301 Q3:3713 Maximum:1114800
CAvideos_df.describe()
# ### Data visualization for numeric attributes
# As we can see, some attributes have extreme skewed character.
plt.figure(dpi=300)
plt.hist(x=CAvideos_df['category_id'], bins='auto')
plt.xlabel("category id")
plt.ylabel("Frequency")
plt.figure(dpi=300)
plt.hist(x=CAvideos_df["views"], bins="auto")
plt.xlabel("views for each video")
plt.ylabel("Frequency")
plt.figure(dpi=300)
plt.hist(x=CAvideos_df["likes"], bins="auto")
plt.xlabel("likes for each video")
plt.ylabel("Frequency")
plt.figure(dpi=300)
plt.hist(x=CAvideos_df["comment_count"], bins="auto")
plt.xlabel("comment_count for each video")
plt.ylabel("Frequency")
CAvideos_df_for_boxplot = CAvideos_df[['views', 'likes', 'dislikes', 'comment_count', 'category_id']]
# #### In order to make the drawing easier, we do logarithmic processing for each attribute.
CAvideos_df_for_boxplot['views'] = np.log(CAvideos_df_for_boxplot['views'] + 1)
CAvideos_df_for_boxplot['likes'] = np.log(CAvideos_df_for_boxplot['likes'] + 1)
CAvideos_df_for_boxplot['dislikes'] = np.log(CAvideos_df_for_boxplot['dislikes'] + 1)
CAvideos_df_for_boxplot['comment_count'] = np.log(CAvideos_df_for_boxplot['comment_count'] + 1)
# #### Next, I will draw a boxplot for each numerical attribute according to each category ID.
import seaborn as sns
fig, ax = plt.subplots(figsize=(20, 24))
ax = sns.boxplot(y="category_id", x='views', data=CAvideos_df_for_boxplot, orient="h")
plt.xlabel('views(log)', fontsize=20)
plt.ylabel('category id', fontsize=20)
plt.show()
import seaborn as sns
fig, ax = plt.subplots(figsize=(20, 24))
ax = sns.boxplot(y="category_id", x='likes', data=CAvideos_df_for_boxplot, orient="h")
plt.xlabel('likes(log)', fontsize=20)
plt.ylabel('category id', fontsize=20)
plt.show()
import seaborn as sns
fig, ax = plt.subplots(figsize=(20, 24))
ax = sns.boxplot(y="category_id", x='dislikes', data=CAvideos_df_for_boxplot, orient="h")
plt.xlabel('dislikes(log)', fontsize=20)
plt.ylabel('category id', fontsize=20)
plt.show()
import seaborn as sns
fig, ax = plt.subplots(figsize=(20, 24))
ax = sns.boxplot(y="category_id", x='comment_count', data=CAvideos_df_for_boxplot, orient="h")
plt.xlabel('comment count(log)', fontsize=20)
plt.ylabel('category id', fontsize=20)
plt.show()
# ### Handle null values
def nullplot(data):
'''
data: a dataframe on which we want to perform null plot
'''
df_null = data.isna()
nulls_percent_per_col = df_null.sum(axis=0) / len(data)
plt.figure(dpi=600)
fig, (ax_1, ax_2) = plt.subplots(nrows=2, ncols=1, sharex=True, figsize=(8, 10))
pla = matplotlib.cm.get_cmap("plasma")
colormap = matplotlib.colors.ListedColormap([pla(0), 'gold'])
sns.heatmap(data.isnull(), cmap=colormap, ax=ax_1, cbar=False, yticklabels=False)
nulls_percent_per_col.plot(kind="bar", color="gold", x=nulls_percent_per_col, y=nulls_percent_per_col.index,
ax=ax_2, label="Null value percent")
ax_2.set_ylim((0,1))
plt.show()
# Only attribute 'description' has a few of null values.
CAvideos_df_2 = CAvideos_df.copy()
nullplot(CAvideos_df_2)
CAvideos_df_2.isnull().sum()
# #### Eliminate null values --- drop rows
CAvideos_df_3 = CAvideos_df.copy()
CAvideos_df_3 = CAvideos_df_3.dropna(axis=0)
nullplot(CAvideos_df_3)
| youtube trading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# make Notebook wider
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:90% !important; }</style>"))
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('./'):
for filename in filenames:
print(os.path.join(dirname, filename))
df = pd.read_csv("../immo_data.csv")
print("df shape:", df.shape)
display(df.info())
display(df.head())
# +
import matplotlib.pyplot as plt
# pd.set_option('max_columns', None)
def plot_barh(dict, title=""):
full_height = (len(dict.keys())) / 3
plt.figure(figsize=(10, full_height))
plt.gca().spines['right'].set_color('none')
plt.gca().spines['top'].set_color('none')
plt.title(title)
pp = plt.barh(list(dict.keys()), list(dict.values()), align='center', alpha=0.5)
for p in pp:
width = p.get_width()
plt.annotate('{}'.format(width),
xy=(width, p.get_y()),
xytext=(3, 6),
textcoords="offset points",
ha='left', va='center')
plt.show()
# -
record_count_by_region = df.groupby(['regio1']).size().sort_values(ascending=True).to_dict()
plot_barh(record_count_by_region, "Records count by region")
# +
berlinDf = df[df['regio1']=='Berlin']
records_by_neighbourhood_in_berlin = berlinDf.groupby(['regio3']).size().sort_values(ascending=True).to_dict()
plot_barh(records_by_neighbourhood_in_berlin, "Records by neighbourhood in Berlin")
# +
import seaborn as sns
useful_columns = [
'regio1', 'regio3', 'heatingType', 'newlyConst', 'balcony', 'picturecount', 'totalRent', 'yearConstructed', 'noParkSpaces', 'firingTypes', 'hasKitchen', 'geo_bln', 'cellar', 'baseRent', 'livingSpace', 'condition', 'interiorQual', 'petsAllowed', 'lift', 'typeOfFlat', 'noRooms', 'floor', 'numberOfFloors', 'garden', 'heatingCosts', 'energyEfficiencyClass', 'lastRefurbish',
]
berlinDf_select = berlinDf[useful_columns]
print("NA records by feature:")
display(sns.heatmap(berlinDf_select.isna(), yticklabels=False, cbar=False, ))
berlinDf_select.isna().sum()
# +
# We will be removing: ['yearConstructed', 'noParkSpaces', 'floor', 'numberOfFloors', 'typeOfFlat', 'heatingCosts', 'lastRefurbish', 'interiorQual']
# because there are too many NA values, and feature are not that important.
berlinDf_select = berlinDf_select.drop(['yearConstructed', 'noParkSpaces', 'floor', 'numberOfFloors', 'typeOfFlat', 'heatingCosts', 'lastRefurbish', 'interiorQual', 'petsAllowed', 'energyEfficiencyClass', 'firingTypes'], axis='columns')
# Also ['regio1', 'totalRent', 'picturecount', 'condition', 'geo_bln'] is not a value that we want to predict on.
berlinDf_select = berlinDf_select.drop(['regio1', 'totalRent', 'picturecount', 'condition', 'geo_bln'], axis='columns')
berlinDf_select.isna().sum()
# We will be keeping ['heatingType', 'firingTypes'] because I still think they are relevant. We will fill NA with NO_INFORMATION and experiemtn with the model.
# +
# Show correlation between numerical types:
correlation = berlinDf_select.corr()
import seaborn as sn
import matplotlib.pyplot as plt
plt.figure(figsize=(18, 18))
sn.heatmap(correlation, annot=True, cbar=False, linewidths=.5, cmap="YlGnBu", fmt='.2f', annot_kws={'size': 15})
plt.tick_params(axis='both', which='major', labelsize=23, labelbottom = True, bottom=True, top = True, labeltop=True, right=True, labelright=True)
plt.xticks(rotation=90)
plt.yticks(rotation=0)
plt.show()
# Checking the baseRent corelation with possible features:
# Data shows that livingSpace and noRooms has the highest corelation ~0.8.
# But constucton year is important too ~0.5.
# Surprisingly seems like heatingCost is also directly corelated with baseRent too. I was expecting this to be inverse corelated.
# Result: We will experiement with following columns for training: livingSpace, noRooms, heatingCosts, hasKitchen, cellar, garden, balcony
# Plus categorical columns for: heatingType, firingTypes, condition', 'interiorQual', 'petsAllowed, energyEfficiencyClass
# +
import matplotlib.pyplot as plt
def plot_ticks(x, y, x_label, y_label):
plt.figure(figsize=(10, 5))
plt.plot(x, y, 'o')
plt.xlabel(x_label)
plt.ylabel(y_label)
plt.show()
for c in ["livingSpace", "noRooms", "hasKitchen", "cellar", "garden", "balcony"]:
plot_ticks(berlinDf_select["baseRent"], berlinDf_select[c], 'baseRent', c)
# -
for c in ["heatingType"]:
berlinDf_select[c] = berlinDf_select[c].fillna("NO_INFORMATION")
classes = berlinDf_select[c].unique()
data = {}
for cl in classes:
mean = berlinDf_select[berlinDf_select[c]==cl]["baseRent"].mean()
data[str(cl)] = round(mean,0)
plot_barh(data, c)
# +
# we need to reduce the heatingType to 2..3 categorical values to not be hard to predict for users
# new 'heating' column: ['low', 'normal', 'high']
# 'NO_INFORMATION' will result in 'heating'='normal'
berlinDf_select['heatingType'].unique()
conditions = [
(berlinDf_select['heatingType']== 'night_storage_heater'),
(berlinDf_select['heatingType']== 'solar_heating'),
(berlinDf_select['heatingType']== 'self_contained_central_heating'),
(berlinDf_select['heatingType']== 'heat_pump'),
(berlinDf_select['heatingType']== 'gas_heating'),
(berlinDf_select['heatingType']== 'combined_heat_and_power_plant'),
(berlinDf_select['heatingType']== 'district_heating'),
(berlinDf_select['heatingType']== 'floor_heating'),
(berlinDf_select['heatingType']== 'central_heating'),
(berlinDf_select['heatingType']== 'stove_heating'),
(berlinDf_select['heatingType']== 'electric_heating'),
(berlinDf_select['heatingType']== 'wood_pellet_heating'),
(berlinDf_select['heatingType']== 'oil_heating'),
(berlinDf_select['heatingType']== 'NO_INFORMATION'),
]
result = ['low', 'low',
'normal', 'normal', 'normal', 'normal', 'normal', 'normal', 'normal',
'high', 'high', 'high', 'high',
'normal']
berlinDf_select['heating'] = np.select(conditions, result)
berlinDf_select.drop('heatingType', axis='columns', inplace=True)
berlinDf_select
# +
# 'regio3' contains the neighbourhoods and also the subneighbourhoods in following format: "subneighbourhood_neighbourhoods"
# we will only keep neighbourhood to be easier to predit
berlinDf_select['regio3'].unique()
berlinDf_select[['subneighbourhoods','neighbourhoods']] = pd.DataFrame(berlinDf_select.regio3.str.split('_', 1, expand=True))
berlinDf_select['neighbourhoods'] = ['Prenzlauer_Berg' if n == "Berg_Prenzlauer_Berg" else n for n in berlinDf_select['neighbourhoods']]
berlinDf_select['neighbourhoods'] = ['Hohenschönhausen' if n == "Hohenschönhausen_Hohenschönhausen" else n for n in berlinDf_select['neighbourhoods']]
berlinDf_select['neighbourhoods'] = ['Pankow' if n == "Buchholz_Pankow" else n for n in berlinDf_select['neighbourhoods']]
# neighbourhoods to lowercase
berlinDf_select['neighbourhoods'] = [n.lower() for n in berlinDf_select['neighbourhoods']]
#rename neighbourhoodscolumn name
berlinDf_select.rename(columns = {'neighbourhoods': 'neighbourhood'}, inplace = True)
berlinDf_select.rename(columns = {'subneighbourhoods': 'subneighbourhood'}, inplace = True)
print("subneighbourhoods count:", len(berlinDf_select['subneighbourhood'].unique()))
neighbourhoods = berlinDf_select['neighbourhood'].unique()
print("neighbourhoods count:", len(neighbourhoods))
print("neighbourhoods:", neighbourhoods)
berlinDf_select.drop('regio3', axis='columns', inplace=True)
# berlinDf_select.drop('subneighbourhood', axis='columns', inplace=True)
berlinDf_select
# +
# Calculating target encoding for subreagions:
subneighbourhood_names = berlinDf_select['subneighbourhood'].unique()
subneighbourhood_mean_baseRent = {}
for s in subneighbourhood_names :
selected = berlinDf_select[berlinDf_select['subneighbourhood']==s]
mean_baseRent = selected.baseRent.mean()
subneighbourhood_mean_baseRent[s] = int(mean_baseRent)
subneighbourhood_mean_baseRent_sorted = dict(sorted(subneighbourhood_mean_baseRent.items(), key=lambda x: x[1], reverse=False))
plot_barh(subneighbourhood_mean_baseRent_sorted, "Records by sub-neighbourhood in Berlin")
# +
# Replacing sub-neighbourhood with mean base rent as target encoded:
berlinDf_select['subneighbourhood_meanBaseRent'] = berlinDf_select['subneighbourhood'].apply(lambda x: subneighbourhood_mean_baseRent_sorted[x])
berlinDf_select.drop('subneighbourhood', axis='columns', inplace=True)
berlinDf_select
# -
# make baseRent, livingSpace and noRooms ab Int
berlinDf_select['baseRent'] = berlinDf_select['baseRent'].astype('int').values
berlinDf_select['livingSpace'] = berlinDf_select['livingSpace'].astype('int').values
berlinDf_select['noRooms'] = berlinDf_select['noRooms'].astype('int').values
berlinDf_select
# +
# We will:
# - later experiment training by exluding the ["hasKitchen", "cellar", "garden", "balcony"]
# - remove records with baseRent bigger than 5000 and 7500
# - remove record with noRooms > 10
berlinDf_final = berlinDf_select
berlinDf_final = berlinDf_final.drop(berlinDf_final[berlinDf_final['baseRent']>5000].index)
berlinDf_final = berlinDf_final.drop(berlinDf_final[berlinDf_final['noRooms']>10].index)
berlinDf_final
# +
print("Final data:")
display(berlinDf_final.isna().sum())
berlinDf_final.reset_index(drop=True, inplace=True)
pd.set_option('display.max_columns', None)
berlinDf_final
# +
# Split data
from sklearn.model_selection import train_test_split
def split_dataFrame(df_to_split):
df_full_train, df_test = train_test_split(df_to_split, test_size=0.2, random_state=11)
df_train, df_val = train_test_split(df_full_train, test_size=0.25, random_state=11)
df_full_train = df_full_train.reset_index(drop=True)
df_train = df_train.reset_index(drop=True)
df_val = df_val.reset_index(drop=True)
df_test = df_test.reset_index(drop=True)
y_full_train = df_full_train.baseRent
y_train = df_train.baseRent
y_val = df_val.baseRent
y_test = df_test.baseRent
del df_full_train["baseRent"]
del df_train["baseRent"]
del df_val["baseRent"]
del df_test["baseRent"]
# with pd.option_context('display.max_rows', 2, 'display.max_columns', None):
# display(df_test)
print("df_to_split length: ", len(df_to_split))
print()
print("df_full_train length: ", len(df_full_train))
print("df_train length: ", len(df_train))
print("df_val length: ", len(df_val))
print("df_test length: ", len(df_test))
print()
print("y_full_train length: ", len(y_full_train))
print("y_train length: ", len(y_train))
print("y_val length: ", len(y_val))
print("y_test length: ", len(y_test))
return df_full_train, df_train, df_val, df_test, y_full_train, y_train, y_val, y_test
split_dataFrame(berlinDf_select)
""
| capstone-project/target_encoding/data_analysis_target_encoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This jupyter notebooks provides the code to give an introduction to the PyWavelets library.
# ### To get some more background information, please have a look at the accompanying blog-post:
# ### http://ataspinar.com/2018/12/21/a-guide-for-using-the-wavelet-transform-in-machine-learning/
import pywt
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
# # 1. Which Wavelets are present in PyWavelets?
# +
wavelet_families = pywt.families(short=False)
discrete_mother_wavelets = pywt.wavelist(kind='discrete')
continuous_mother_wavelets = pywt.wavelist(kind='continuous')
print("PyWavelets contains the following families: ")
print(wavelet_families)
print()
print("PyWavelets contains the following Continuous families: ")
print(continuous_mother_wavelets)
print()
print("PyWavelets contains the following Discrete families: ")
print(discrete_mother_wavelets)
print()
for family in pywt.families():
print(" * The {} family contains: {}".format(family, pywt.wavelist(family)))
# -
# # 2. Visualizing several Discrete and Continuous wavelets
# +
discrete_wavelets = ['db5', 'sym5', 'coif5', 'bior2.4']
continuous_wavelets = ['mexh', 'morl', 'cgau5', 'gaus5']
list_list_wavelets = [discrete_wavelets, continuous_wavelets]
list_funcs = [pywt.Wavelet, pywt.ContinuousWavelet]
fig, axarr = plt.subplots(nrows=2, ncols=4, figsize=(16,8))
for ii, list_wavelets in enumerate(list_list_wavelets):
func = list_funcs[ii]
row_no = ii
for col_no, waveletname in enumerate(list_wavelets):
wavelet = func(waveletname)
family_name = wavelet.family_name
biorthogonal = wavelet.biorthogonal
orthogonal = wavelet.orthogonal
symmetry = wavelet.symmetry
if ii == 0:
_ = wavelet.wavefun()
wavelet_function = _[0]
x_values = _[-1]
else:
wavelet_function, x_values = wavelet.wavefun()
if col_no == 0 and ii == 0:
axarr[row_no, col_no].set_ylabel("Discrete Wavelets", fontsize=16)
if col_no == 0 and ii == 1:
axarr[row_no, col_no].set_ylabel("Continuous Wavelets", fontsize=16)
axarr[row_no, col_no].set_title("{}".format(family_name), fontsize=16)
axarr[row_no, col_no].plot(x_values, wavelet_function)
axarr[row_no, col_no].set_yticks([])
axarr[row_no, col_no].set_yticklabels([])
plt.tight_layout()
plt.show()
# -
# ## 3. Visualizing how the wavelet form depends on the order and decomposition level
# +
fig, axarr = plt.subplots(ncols=5, nrows=5, figsize=(20,16))
fig.suptitle('Daubechies family of wavelets', fontsize=16)
db_wavelets = pywt.wavelist('db')[:5]
for col_no, waveletname in enumerate(db_wavelets):
wavelet = pywt.Wavelet(waveletname)
no_moments = wavelet.vanishing_moments_psi
family_name = wavelet.family_name
for row_no, level in enumerate(range(1,6)):
wavelet_function, scaling_function, x_values = wavelet.wavefun(level = level)
axarr[row_no, col_no].set_title("{} - level {}\n{} vanishing moments\n{} samples".format(
waveletname, level, no_moments, len(x_values)), loc='left')
axarr[row_no, col_no].plot(x_values, wavelet_function, 'bD--')
axarr[row_no, col_no].set_yticks([])
axarr[row_no, col_no].set_yticklabels([])
plt.tight_layout()
plt.subplots_adjust(top=0.9)
plt.show()
# -
# ## 4.A Using the pywt.dwt() for the decomposition of a signal into the frequency sub-bands
# ### (and reconstrucing it again)
# +
time = np.linspace(0, 1, num=2048)
chirp_signal = np.sin(250 * np.pi * time**2)
(cA1, cD1) = pywt.dwt(chirp_signal, 'db2', 'smooth')
(cA2, cD2) = pywt.dwt(cA1, 'db2', 'smooth')
(cA3, cD3) = pywt.dwt(cA2, 'db2', 'smooth')
(cA4, cD4) = pywt.dwt(cA3, 'db2', 'smooth')
(cA5, cD5) = pywt.dwt(cA4, 'db2', 'smooth')
coefficients_level1 = [cA1, cD1]
coefficients_level2 = [cA2, cD2, cD1]
coefficients_level3 = [cA3, cD3, cD2, cD1]
coefficients_level4 = [cA4, cD4, cD3, cD2, cD1]
coefficients_level5 = [cA5, cD5, cD4, cD3, cD2, cD1]
reconstructed_signal_level1 = pywt.waverec(coefficients_level1, 'db2', 'smooth')
reconstructed_signal_level2 = pywt.waverec(coefficients_level2, 'db2', 'smooth')
reconstructed_signal_level3 = pywt.waverec(coefficients_level3, 'db2', 'smooth')
reconstructed_signal_level4 = pywt.waverec(coefficients_level4, 'db2', 'smooth')
reconstructed_signal_level5 = pywt.waverec(coefficients_level5, 'db2', 'smooth')
fig, ax = plt.subplots(figsize=(12,4))
ax.plot(chirp_signal, label='signal')
ax.plot(reconstructed_signal_level1, label='reconstructed level 1', linestyle='--')
ax.plot(reconstructed_signal_level2, label='reconstructed level 2', linestyle='--')
ax.plot(reconstructed_signal_level3, label='reconstructed level 3', linestyle='--')
ax.plot(reconstructed_signal_level4, label='reconstructed level 4', linestyle='--')
ax.plot(reconstructed_signal_level5, label='reconstructed level 5', linestyle='--')
ax.legend(loc='upper right')
ax.set_title('single reconstruction', fontsize=20)
ax.set_xlabel('time axis', fontsize=16)
ax.set_ylabel('Amplitude', fontsize=16)
plt.show()
# -
# ## 4.B Using the pywt.wavedec() for the decomposition of a signal into the frequency sub-bands
# ### (and reconstrucing it again)
# +
time = np.linspace(0, 1, num=2048)
chirp_signal = np.sin(250 * np.pi * time**2)
coefficients_level1 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=1)
coefficients_level2 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=2)
coefficients_level3 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=3)
coefficients_level4 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=4)
coefficients_level5 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=5)
reconstructed_signal_level1 = pywt.waverec(coefficients_level1, 'db2', 'smooth')
reconstructed_signal_level2 = pywt.waverec(coefficients_level2, 'db2', 'smooth')
reconstructed_signal_level3 = pywt.waverec(coefficients_level3, 'db2', 'smooth')
reconstructed_signal_level4 = pywt.waverec(coefficients_level4, 'db2', 'smooth')
reconstructed_signal_level5 = pywt.waverec(coefficients_level5, 'db2', 'smooth')
fig, ax = plt.subplots(figsize=(12,4))
ax.plot(chirp_signal, label='signal')
ax.plot(reconstructed_signal_level1, label='reconstructed level 1', linestyle='--')
ax.plot(reconstructed_signal_level2, label='reconstructed level 2', linestyle='--')
ax.plot(reconstructed_signal_level3, label='reconstructed level 3', linestyle='--')
ax.plot(reconstructed_signal_level4, label='reconstructed level 4', linestyle='--')
ax.plot(reconstructed_signal_level5, label='reconstructed level 5', linestyle='--')
ax.legend(loc='upper right')
ax.set_title('single reconstruction', fontsize=20)
ax.set_xlabel('time axis', fontsize=16)
ax.set_ylabel('Amplitude', fontsize=16)
plt.show()
# -
# ## 5. Reconstrucing a signal with only one level of coefficients
# +
fig = plt.figure(figsize=(6,8))
spec = gridspec.GridSpec(ncols=2, nrows=6)
ax0 = fig.add_subplot(spec[0, 0:2])
ax1a = fig.add_subplot(spec[1, 0])
ax1b = fig.add_subplot(spec[1, 1])
ax2a = fig.add_subplot(spec[2, 0])
ax2b = fig.add_subplot(spec[2, 1])
ax3a = fig.add_subplot(spec[3, 0])
ax3b = fig.add_subplot(spec[3, 1])
ax4a = fig.add_subplot(spec[4, 0])
ax4b = fig.add_subplot(spec[4, 1])
ax5a = fig.add_subplot(spec[5, 0])
ax5b = fig.add_subplot(spec[5, 1])
axarr = np.array([[ax1a, ax1b],[ax2a, ax2b],[ax3a, ax3b],[ax4a, ax4b],[ax5a, ax5b]])
time = np.linspace(0, 1, num=2048)
chirp_signal = np.sin(250 * np.pi * time**2)
# First we reconstruct a signal using pywt.wavedec() as we have also done at #4.2
coefficients_level1 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=1)
coefficients_level2 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=2)
coefficients_level3 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=3)
coefficients_level4 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=4)
coefficients_level5 = pywt.wavedec(chirp_signal, 'db2', 'smooth', level=5)
# pywt.wavedec() returns a list of coefficients. Below we assign these list of coefficients to variables explicitely.
[cA1_l1, cD1_l1] = coefficients_level1
[cA2_l2, cD2_l2, cD1_l2] = coefficients_level2
[cA3_l3, cD3_l3, cD2_l3, cD1_l3] = coefficients_level3
[cA4_l4, cD4_l4, cD3_l4, cD2_l4, cD1_l4] = coefficients_level4
[cA5_l5, cD5_l5, cD4_l5, cD3_l5, cD2_l5, cD1_l5] = coefficients_level5
# Since the the list of coefficients have been assigned explicitely to variables, we can set a few of them to zero.
approx_coeff_level1_only = [cA1_l1, None]
detail_coeff_level1_only = [None, cD1_l1]
approx_coeff_level2_only = [cA2_l2, None, None]
detail_coeff_level2_only = [None, cD2_l2, None]
approx_coeff_level3_only = [cA3_l3, None, None, None]
detail_coeff_level3_only = [None, cD3_l3, None, None]
approx_coeff_level4_only = [cA4_l4, None, None, None, None]
detail_coeff_level4_only = [None, cD4_l4, None, None, None]
approx_coeff_level5_only = [cA5_l5, None, None, None, None, None]
detail_coeff_level5_only = [None, cD5_l5, None, None, None, None]
# By reconstrucing the signal back from only one set of coefficients, we can see how
# the frequency-sub band for that specific set of coefficient looks like
rec_signal_cA_level1 = pywt.waverec(approx_coeff_level1_only, 'db2', 'smooth')
rec_signal_cD_level1 = pywt.waverec(detail_coeff_level1_only, 'db2', 'smooth')
rec_signal_cA_level2 = pywt.waverec(approx_coeff_level2_only, 'db2', 'smooth')
rec_signal_cD_level2 = pywt.waverec(detail_coeff_level2_only, 'db2', 'smooth')
rec_signal_cA_level3 = pywt.waverec(approx_coeff_level3_only, 'db2', 'smooth')
rec_signal_cD_level3 = pywt.waverec(detail_coeff_level3_only, 'db2', 'smooth')
rec_signal_cA_level4 = pywt.waverec(approx_coeff_level4_only, 'db2', 'smooth')
rec_signal_cD_level4 = pywt.waverec(detail_coeff_level4_only, 'db2', 'smooth')
rec_signal_cA_level5 = pywt.waverec(approx_coeff_level5_only, 'db2', 'smooth')
rec_signal_cD_level5 = pywt.waverec(detail_coeff_level5_only, 'db2', 'smooth')
ax0.set_title("Chirp Signal", fontsize=16)
ax0.plot(time, chirp_signal)
ax0.set_xticks([])
ax0.set_yticks([])
ax1a.plot(rec_signal_cA_level1, color='red')
ax1b.plot(rec_signal_cD_level1, color='green')
ax2a.plot(rec_signal_cA_level2, color='red')
ax2b.plot(rec_signal_cD_level2, color='green')
ax3a.plot(rec_signal_cA_level3, color='red')
ax3b.plot(rec_signal_cD_level3, color='green')
ax4a.plot(rec_signal_cA_level4, color='red')
ax4b.plot(rec_signal_cD_level4, color='green')
ax5a.plot(rec_signal_cA_level5, color='red')
ax5b.plot(rec_signal_cD_level5, color='green')
for ii in range(0,5):
axarr[ii,0].set_xticks([])
axarr[ii,0].set_yticks([])
axarr[ii,1].set_xticks([])
axarr[ii,1].set_yticks([])
axarr[ii,0].set_title("Approximation Coeff", fontsize=16)
axarr[ii,1].set_title("Detail Coeff", fontsize=16)
axarr[ii,0].set_ylabel("Level {}".format(ii+1), fontsize=16)
plt.tight_layout()
plt.show()
# -
| notebooks/WV1 - Using PyWavelets for Wavelet Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dask Array
# 
#
# Dask 数组协调许多 Numpy 数组,在网格内排列成块。它们支持 Numpy API 的很大一部分。
#
# Dask 数组使用阻塞算法提供了一个并行的、大于内存的 n 维数组。简单地说:分布式 Numpy
#
# - **并行**:使用计算机上的所有内核
#
# - **大于内存**:通过将数组分解成许多小块,按顺序操作这些块以最大限度地减少计算的内存占用,并有效地从磁盘流式传输数据,从而让您可以处理大于可用内存的数据集.
#
# - **阻塞算法**:通过执行许多较小的计算来执行大型计算
# +
# 连接/创建 Dask集群
from dask.distributed import Client
client = Client(n_workers=4, threads_per_worker=1)
client
# -
# ## 示例一
#
# **针对一个有10亿个随机数的大数组做加法,分而治之**
# dask使用chunks参数将大数组切分成小块,先针对小块数组进行计算,然后将小块计算的结果再计算,最后得到结果
import dask.array as da
x = da.random.random((100_000_000_000,), chunks=(10_000_000,))
x
result = x.sum()
result
# 8.56G of 16G default
# 9.73G chunk x10
result.compute()
# +
# numpy运行时间
# 15 G out of 16
import numpy as np
x = np.random.random((100_000_000_000,))
x.sum()
# -
# ## 示例二
#
# 1. **构建一个 20000x20000 的正态分布随机值数组,该数组被分成 1000x1000 大小的块**
# 2. **沿一个轴取平均值**
# 3. **取每 100 个元素**
# +
import numpy as np
import dask.array as da
x = da.random.normal(10, 0.1, size=(20000, 20000),
chunks=(1000, 1000))
x
# -
y = x.mean(axis=0)[::100]
y
# %%time
y.compute()
# - 更改chunks大小对执行有什么影响?
# - chunks = (10000, 10000)
# - chunks = (25, 25)
# +
import numpy as np
import dask.array as da
x = da.random.normal(10, 0.1, size=(20000, 20000),
chunks=(20000, 20000))
x
# -
y = x.mean(axis=0)[::100]
# %time y.compute()
# +
# %%time
# numpy运算时间
import numpy as np
x = np.random.normal(10, 0.1, size=(20000, 20000))
x.mean(axis=0)[::100]
# -
# # 示例三
#
# **持久化计算到内存中,加速后续计算**
import dask.array as da
x = da.random.random((10000, 10000), chunks=(1000, 1000))
x
y = x + x.T
z = y[::2, 5000:].mean(axis=1)
z
# %time z.compute()
# 持久化
y = y.persist()
# %time y[0, 0].compute()
# %time y[0, 0].compute()
# 清楚内存占用
client.cancel(y)
# or del y
client.shutdown()
| dask/dask-array.ipynb |