code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 10.557037, "end_time": "2021-12-27T05:54:47.880926", "exception": false, "start_time": "2021-12-27T05:54:37.323889", "status": "completed"} tags=[]
# !pip install yfinance
# + papermill={"duration": 7.487879, "end_time": "2021-12-27T05:54:55.381980", "exception": false, "start_time": "2021-12-27T05:54:47.894101", "status": "completed"} tags=[]
# !pip install pycausalimpact
# + papermill={"duration": 0.029912, "end_time": "2021-12-27T05:54:55.425788", "exception": false, "start_time": "2021-12-27T05:54:55.395876", "status": "completed"} tags=[]
import yfinance as yf
import pandas as pd
# + papermill={"duration": 0.020321, "end_time": "2021-12-27T05:54:55.459898", "exception": false, "start_time": "2021-12-27T05:54:55.439577", "status": "completed"} tags=[]
training_start = "2018-01-02"
training_end = "2018-09-05"
treatment_start = "2018-09-06"
treatment_end = "2018-09-07"
#we need to check what happened during these dates hence they are called treatment dates
end_stock = "2018-09-08"
# + papermill={"duration": 1.01443, "end_time": "2021-12-27T05:54:56.488015", "exception": false, "start_time": "2021-12-27T05:54:55.473585", "status": "completed"} tags=[]
stocks = ["TSLA","VOLVF","GOOG","BMW.DE","GM","DAI.DE"]
dataset = yf.download(stocks, start = training_start, end = end_stock,interval ="1d")
# + papermill={"duration": 0.057926, "end_time": "2021-12-27T05:54:56.561854", "exception": false, "start_time": "2021-12-27T05:54:56.503928", "status": "completed"} tags=[]
dataset.head()
# + papermill={"duration": 0.034494, "end_time": "2021-12-27T05:54:56.612525", "exception": false, "start_time": "2021-12-27T05:54:56.578031", "status": "completed"} tags=[]
dataset = dataset.iloc[:,:6]
dataset.columns = dataset.columns.droplevel()
dataset = dataset.dropna()
dataset.head()
# + papermill={"duration": 0.033583, "end_time": "2021-12-27T05:54:56.662755", "exception": false, "start_time": "2021-12-27T05:54:56.629172", "status": "completed"} tags=[]
dataset.info()
# + papermill={"duration": 1.393649, "end_time": "2021-12-27T05:54:58.073557", "exception": false, "start_time": "2021-12-27T05:54:56.679908", "status": "completed"} tags=[]
data_corr = dataset[dataset.index <= training_end]
import seaborn as sns
import matplotlib.pyplot as plt
#get correlations of each features in dataset
corrmat = data_corr.corr()
top_corr_features = corrmat.index
plt.figure(figsize=(10,10))
#plot heat map
g=sns.heatmap(data_corr[top_corr_features].corr(),annot=True,cmap="RdYlGn")
# + papermill={"duration": 0.026648, "end_time": "2021-12-27T05:54:58.118001", "exception": false, "start_time": "2021-12-27T05:54:58.091353", "status": "completed"} tags=[]
final_stocks = dataset[["TSLA","VOLVF","GOOG","GM","DAI.DE"]]
pre_period = [training_start,training_end]
post_period = [treatment_start, treatment_end]
# + papermill={"duration": 2.260026, "end_time": "2021-12-27T05:55:00.396013", "exception": false, "start_time": "2021-12-27T05:54:58.135987", "status": "completed"} tags=[]
from causalimpact import CausalImpact
causal_effect = CausalImpact(data=final_stocks, pre_period=pre_period,post_period=post_period)
# + papermill={"duration": 0.029179, "end_time": "2021-12-27T05:55:00.443878", "exception": false, "start_time": "2021-12-27T05:55:00.414699", "status": "completed"} tags=[]
print(causal_effect.summary())
# + [markdown] papermill={"duration": 0.01818, "end_time": "2021-12-27T05:55:00.480525", "exception": false, "start_time": "2021-12-27T05:55:00.462345", "status": "completed"} tags=[]
# A p value of less than 0.05 shows that the results are statistically significant.
#
#
# + papermill={"duration": 0.018381, "end_time": "2021-12-27T05:55:00.517381", "exception": false, "start_time": "2021-12-27T05:55:00.499000", "status": "completed"} tags=[]
| tesla-stocks-causalimpact.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="OgYMbUm0VlP-"
# <p><img alt="Yahoo logo" height="60px" src="https://www.ulacit.ac.cr/wp-content/uploads/Logo-Ulacit-Blanco.png" align="left" hspace="10px" vspace="0px"></p>
# + [markdown] id="wO-2zsXoZAvc"
# # Operadores Booleanos
#
# La actual computación basa sus operaciones más fundamentales en la lógica Booleana, en donde también sale a relucir los conceptos relacionados de compuertas lógicas. En este documento podemos encontrar los operadores lógicos en Python (A través de `True` o `False`).
# + [markdown] id="FCIjk10eXXc5"
# MIT License
#
# Profesor <NAME>
# + [markdown] id="GhnQVTiNPytK"
# 67-0001
# criptografía
# + [markdown] id="YuYFIvMUINcl"
#
#
# ---
#
#
# + [markdown] id="uLBzhf1nIH9g"
# Consultamos con la funcion Type, cual es el tipado de los datos True y False.
# + id="gQchT0G1ZbTu" outputId="c778222f-7698-4bce-a5fb-ce6ccfe63ffa" colab={"base_uri": "https://localhost:8080/"}
type(True)
# + colab={"base_uri": "https://localhost:8080/"} id="nquxF2DaIFeE" outputId="a118f21b-d646-4bda-921b-ecbd2f13e3e1"
type(False)
# + [markdown] id="zDV9Nk6YrCDi"
# ## Operadores Booleanos
#
# ### Negación (`not`)
#
# La negación devuelve `False` cuando es aplicada sobre un `True` y viceversa. Es un operador unario. Es decir, sólo necesita un operando para devolver un resultado.
# + id="anz7MCkEG-GR" outputId="6410ba43-4799-435c-fcf1-4031f03e8b90" colab={"base_uri": "https://localhost:8080/"}
not False
# + id="gi1oL9ejHGKH" outputId="593c4d62-c648-4ee3-85a3-0dc1b8673faa" colab={"base_uri": "https://localhost:8080/"}
not True
# + [markdown] id="bU_1eDlScm2D"
# ### Suma lógica (`or`)
#
# La suma lógica devuelve `True` siempre que alguno de sus operandos sea `True`. O lo que es lo mismo, devuelve `False` cuando ambos operando son `False`
# + id="sMwW0Z1Fc8ol" outputId="b741262c-f605-401a-8193-e500e2b18861" colab={"base_uri": "https://localhost:8080/"}
True or False
# + id="dscFMYeUc_AB" outputId="6b960952-f4ea-431e-c034-4407d5fc3b41" colab={"base_uri": "https://localhost:8080/"}
True or True
# + id="VEk4Y2V4dBTo" outputId="b6cab143-171d-4bc2-f228-107f5397a56a" colab={"base_uri": "https://localhost:8080/"}
False or True
# + id="61GTdhsGdC69" outputId="7c54df71-3880-4866-b523-dfd5c9e18821" colab={"base_uri": "https://localhost:8080/"}
False or False
# + [markdown] id="N0Dcm0kidInP"
# ### Producto lógico (`and`)
#
# El producto lógico devuelve `True` sólo si ambos operando son `True`. O lo que es lo mismo, si alguno de sus operandos es `False` el producto lógico devuelve `False`
# + id="HYcoQiemdjiR" outputId="e98fa57e-8f2a-45ca-beb9-1d2255ece66f" colab={"base_uri": "https://localhost:8080/"}
True and True
# + id="CcyIVgFqdl1a" outputId="1fc29fb2-4202-4f09-ec7b-3dab3a4ede92" colab={"base_uri": "https://localhost:8080/"}
True and False
# + id="jif2AmCldqEM" outputId="4452d6d3-a1e7-440f-852b-90eb3f6e0837" colab={"base_uri": "https://localhost:8080/"}
False and True
# + [markdown] id="3QB27J7FJFyI"
# ### Producto lógico (`XOR`)
#
# Si una salida verdadera resulta si una, y solo una de las entradas a la puerta es verdadera. Si ambas entradas son falsas o ambas son verdaderas, resulta en una salida falsa.
# + colab={"base_uri": "https://localhost:8080/"} id="k-T7PB79I7cx" outputId="af2002f5-dda9-4115-97af-ba7d99b358fb"
a = bool(1)
b = bool(0)
print(a^b)
# + colab={"base_uri": "https://localhost:8080/"} id="9A6I00RPJKIs" outputId="1389853d-de11-4fde-f8a2-666beca70157"
a and b
# + colab={"base_uri": "https://localhost:8080/"} id="iK0kcNujJMfp" outputId="13dc7e61-8c4d-48b6-e927-d0399894ee66"
a or b
| operadores_booleanos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import os, sys
import json
aPath='/Users/liuchang/Documents/STUDY/AM207/Final Project/Data/data subset'
os.chdir(aPath)
os.path.dirname(os.path.abspath('biz.json'))
# -
# # Sample data for a Yelp business
# {"business_id": "SQ0j7bgSTazkVQlF5AnqyQ", "full_address": "214 E Main St\nCarnegie\nCarnegie, PA 15106", "hours": {}, "open": true, "categories": ["Chinese", "Restaurants"], "city": "Carnegie", "review_count": 8, "name": "<NAME> Restaurant", "neighborhoods": ["Carnegie"], "longitude": -80.084861000000004, "state": "PA", "stars": 2.5, "latitude": 40.408343000000002, "attributes": {"Take-out": true, "Alcohol": "none", "Noise Level": "quiet", "Parking": {"garage": false, "street": false, "validated": false, "lot": false, "valet": false}, "Delivery": true, "Has TV": true, "Outdoor Seating": false, "Attire": "casual", "Waiter Service": false, "Good For Groups": false, "Price Range": 1}, "type": "business"}
#
# # Sample data for a Yelp user
#
# {"yelping_since": "2004-10", "votes": {"funny": 166, "useful": 278, "cool": 245}, "review_count": 108, "name": "Russel", "user_id": "18kPq7GPye-YQ3LyKyAZPw", "friends": .... "fans": 69, "average_stars": 4.1399999999999997, "type": "user", "compliments": {"profile": 8, "cute": 15, "funny": 11, "plain": 25, "writer": 9, "note": 20, "photos": 15, "hot": 48, "cool": 78, "more": 3}, "elite": [2005, 2006]}
#
# # Sample data for a Yelp review
#
# {"votes": {"funny": 0, "useful": 1, "cool": 0}, "user_id": "uK8tzraOp4M5u3uYrqIBXg", "review_id": "KAkcn7oQP1xX8KsZ-XmktA", "stars": 4, "date": "2013-10-20", "text": "This place was very good. I found out about Emil's when watching a show called \"25 Things I Love About Pittsburgh\" on WQED hosted by <NAME>. This place ain't a luxurious restaurant...it's a beer & a shot bar / lounge. But the people are friendly & the food is good. I had the fish sandwich which was great. It ain't in a great part of town, Rankin, but I've been in worse places!! Try this place.", "type": "review", "business_id": "mVHrayjG3uZ_RLHkLj-AMg"}
#
# ## brainstorming questions:
#
# 1. What attributes of users (mood, review counts, etc) tend to give high/lower ratings to a certain type of restaurants (cuisine, locations, money, service)?
#
# For example, how does
#
#
# 2. What attributes of reviews (words, tones, etc) correlate with the ratings of the restaurant?
#
def open_json(json_file):
with open(json_file) as f:
for lines in f:
#line = f.next()
d = json.loads(lines)
df = pd.DataFrame(columns=d.keys())
# df[d.keys()[0]]= d[d.keys()[0]]
return df
# +
# list of files
biz_file ="/Users/liuchang/Documents/STUDY/AM207/Final Project/Data/data subset/biz.json"
review_file ="/Users/liuchang/Documents/STUDY/AM207/Final Project/Data/data subset/review.json"
checkin_file ="/Users/liuchang/Documents/STUDY/AM207/Final Project/Data/data subset/checkin.json"
tip_file ="/Users/liuchang/Documents/STUDY/AM207/Final Project/Data/data subset/tip.json"
user_file ="/Users/liuchang/Documents/STUDY/AM207/Final Project/Data/data subset/user.json"
# -
text = "biz_file,review_file,checkin_file,tip_file,user_file".split(',')
for idx, i in enumerate([biz_file,review_file,checkin_file,tip_file,user_file]):
headers = open_json(i)
print text[idx]
print headers
print
#
print type("biz_file,review_file,checkin_file,tip_file,user_file".split(',')[0])
json_file="/Users/liuchang/Documents/STUDY/AM207/Final Project/Data/data subset/biz.json"
biz =pd.io.json.read_json(json_file,orient='Dataframe')
biz
import pandas as pd
import os
os.path.dirname(os.path.realpath("yelp_academic_dataset_business.json"))
os.chdir("")
# +
import json
def load_json_file(file_path):
"""
Builds a list of dictionaries from a JSON file
:type file_path: string
:param file_path: the path for the file that contains the businesses
data
:return: a list of dictionaries with the data from the files
"""
records = [json.loads(line) for line in open(file_path)]
return records
def tf_idf_tips(file_path):
records = load_json_file(file_path)
print type(records),type(records[4])
data = [record['stars'] for record in records if 'Restaurants' in record['categories']]
return data
tip_matrix = tf_idf_tips("yelp_academic_dataset_business.json")
# -
scores = tip_matrix
plt.hist(scores,bins=30)
plt.show()
# +
import numpy as np
import numpy.random as npr
import pylab
def bootstrap(data, num_samples, statistic, alpha):
"""Returns bootstrap estimate of 100.0*(1-alpha) CI for statistic."""
n = len(data)
idx = npr.randint(0, n, (num_samples, n))
samples = data[idx]
stat = np.sort(statistic(samples, 1))
return (stat[int((alpha/2.0)*num_samples)],
stat[int((1-alpha/2.0)*num_samples)])
if __name__ == '__main__':
# data of interest is bimodal and obviously not normal
#x = np.concatenate([npr.normal(3, 1, 100), npr.normal(6, 2, 200)])
x = scores
# find mean 95% CI and 100,000 bootstrap samples
low, high = bootstrap(x, 100000, np.mean, 0.05)
# make plots
pylab.figure(figsize=(15,10))
pylab.subplot(121)
pylab.hist(x, 50)
pylab.title('Historgram of data')
pylab.subplot(122)
pylab.plot([-0.03,0.03], [np.mean(x), np.mean(x)], 'r', linewidth=2)
pylab.scatter(0.1*(npr.random(len(x))-0.5), x)
pylab.plot([0.19,0.21], [low, low], 'r', linewidth=2)
pylab.plot([0.19,0.21], [high, high], 'r', linewidth=2)
pylab.plot([0.2,0.2], [low, high], 'r', linewidth=2)
pylab.xlim([-0.2, 0.3])
pylab.title('Bootstrap 95% CI for mean')
#pylab.savefig('examples/boostrap.png')
# -
len(x)
| .ipynb_checkpoints/parsing data files _ read json-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/VxctxrTL/daa_2021_1/blob/master/Tarea7.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="v1U8w4HGzHB7" outputId="0ab664a1-50c8-43e8-daff-9fc82fecdbcf"
def sumalista(listaNumeros):
if len(listaNumeros) == 1:
return listaNumeros[0]
else:
return listaNumeros[0] + sumalista(listaNumeros[1:])
print(sumalista([6,24,104,9,15]))
# + colab={"base_uri": "https://localhost:8080/"} id="mApJLDP092PX" outputId="7375b352-2072-4996-f7be-0e6962861272"
def cuentaReg (x):
if x > 0:
print (x)
cuentaReg(x-1)
else:
print ("Fin de la cuenta regresiva")
cuentaReg(3)
# + id="4J-T0cx7aSi4"
class adtCola:
def __init__ (self):
self.items = []
def pop (self):
try:
return self.items.pop(0)
except:
raise ValueError ("La cola esta vacia")
def push (self, x):
self.items.append(x)
| Tarea7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.insert(0, "segmentation_models.pytorch/")
from train import Trainer
import torch
import torch.nn as nn
try:
from efficientnet_pytorch import EfficientNet
except:
os.system(f"""pip install efficientnet-pytorch""")
from efficientnet_pytorch import EfficientNet
# ## Train
model = EfficientNet.from_pretrained('efficientnet-b3')
num_ftrs = model._fc.in_features
model._fc = nn.Linear(num_ftrs, 1)
# training
model_trainer = Trainer(model = model, optim = 'Ranger', lr = 1e-3, bs = 8, name = "b3-ranger")
model_trainer.fit(20)
| notebooks/training_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Autoencoder for ESNs using sklearn
#
# ## Introduction
#
# In this notebook, we demonstrate how the ESN can deal with multipitch tracking, a challenging multilabel classification problem in music analysis.
#
# As this is a computationally expensive task, we have pre-trained models to serve as an entry point.
#
# At first, we import all packages required for this task. You can find the import statements below.
#
# To use another objective than `accuracy_score` for hyperparameter tuning, check out the documentation of [make_scorer](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.make_scorer.html) or ask me.
# +
import numpy as np
from sklearn.metrics import make_scorer
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
from sklearn.utils.fixes import loguniform
from scipy.stats import uniform
from joblib import dump, load
from pyrcn.echo_state_network import SeqToSeqESNClassifier # SeqToSeqESNRegressor or SeqToLabelESNClassifier
from pyrcn.metrics import accuracy_score # more available or create custom score
from pyrcn.model_selection import SequentialSearchCV
import seaborn as sns
from matplotlib import pyplot as plt
from matplotlib import ticker
from mpl_toolkits.mplot3d import Axes3D # noqa: F401 unused import
# %matplotlib inline
#Options
plt.rc('image', cmap='RdBu')
plt.rc('font', family='serif', serif='Times')
plt.rc('text', usetex=True)
plt.rc('xtick', labelsize=8)
plt.rc('ytick', labelsize=8)
plt.rc('axes', labelsize=8)
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('png', 'pdf')
from mpl_toolkits.axes_grid1 import make_axes_locatable
# -
# ## Load and preprocess the dataset
#
# This might require a large amount of and memory.
# +
# At first, please load all training and test sequences and targets.
# Each sequence should be a numpy.array with the shape (n_samples, n_features)
# Each target should be
# - either be a numpy.array with the shape (n_samples, n_targets)
# - or a 1D numpy.array with the shape (n_samples, 1)
train_sequences = ......................
train_targets = ......................
if len(train_sequences) != len(train_targets):
raise ValueError("Number of training sequences does not match number of training targets!")
n_train_sequences = len(train_sequences)
test_sequences = ......................
test_targets = ......................
if len(test_sequences) != len(test_targets):
raise ValueError("Number of test sequences does not match number of test targets!")
n_test_sequences = len(test_sequences)
# Initialize training and test sequences
X_train = np.empty(shape=(n_train_sequences, ), dtype=object)
y_train = np.empty(shape=(n_train_sequences, ), dtype=object)
X_test = np.empty(shape=(n_test_sequences, ), dtype=object)
y_test = np.empty(shape=(n_test_sequences, ), dtype=object)
for k, (train_sequence, train_target) in enumerate(zip(train_sequences, train_targets)):
X_train[k] = train_sequence
y_train[k] = train_target
for k, (test_sequence, test_target) in enumerate(zip(test_sequences, test_targets)):
X_test[k] = test_sequence
y_test[k] = test_target
# -
# Initial variables to be equal in the Autoencoder and in the ESN
hidden_layer_size = 500
input_activation = 'relu'
# ## Train a MLP autoencoder
#
# Currently very rudimentary. However, it can be flexibly made deeper or more complex. Check [MLPRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.neural_network.MLPRegressor.html) documentation for hyper-parameters.
# +
mlp_autoencoder = MLPRegressor(hidden_layer_sizes=(hidden_layer_size, ), activation=input_activation)
# X_train is a numpy array of sequences - the MLP does not handle sequences. Thus, concatenate all sequences
# Target of an autoencoder is the input of the autoencoder
mlp_autoencoder.fit(np.concatenate(X_train), np.concatenate(X_train))
w_in = np.divide(mlp_autoencoder.coefs_[0], np.linalg.norm(mlp_autoencoder.coefs_[0], axis=0)[None, :])
# w_in = mlp_autoencoder.coefs_[0] # uncomment in case that the vector norm does not make sense
# -
# ## Set up an ESN
#
# To develop an ESN model, we need to tune several hyper-parameters, e.g., input_scaling, spectral_radius, bias_scaling and leaky integration.
#
# We define the search spaces for each step in a sequential search together with the type of search (a grid or random search in this context).
#
# At last, we initialize a SeqToSeqESNClassifier with the desired output strategy and with the initially fixed parameters.
# +
initially_fixed_params = {'hidden_layer_size': hidden_layer_size,
'k_in': 10,
'input_scaling': 0.4,
'input_activation': input_activation,
'bias_scaling': 0.0,
'spectral_radius': 0.0,
'leakage': 1.0,
'k_rec': 10,
'reservoir_activation': 'tanh',
'bi_directional': False,
'wash_out': 0,
'continuation': False,
'alpha': 1e-3,
'random_state': 42}
step1_esn_params = {'input_scaling': uniform(loc=1e-2, scale=1),
'spectral_radius': uniform(loc=0, scale=2)}
step2_esn_params = {'leakage': loguniform(1e-5, 1e0)}
step3_esn_params = {'bias_scaling': np.linspace(0.0, 1.0, 11)}
step4_esn_params = {'alpha': loguniform(1e-5, 1e1)}
kwargs_step1 = {'n_iter': 200, 'random_state': 42, 'verbose': 1, 'n_jobs': -1, 'scoring': make_scorer(accuracy_score)}
kwargs_step2 = {'n_iter': 50, 'random_state': 42, 'verbose': 1, 'n_jobs': -1, 'scoring': make_scorer(accuracy_score)}
kwargs_step3 = {'verbose': 1, 'n_jobs': -1, 'scoring': make_scorer(accuracy_score)}
kwargs_step4 = {'n_iter': 50, 'random_state': 42, 'verbose': 1, 'n_jobs': -1, 'scoring': make_scorer(accuracy_score)}
# The searches are defined similarly to the steps of a sklearn.pipeline.Pipeline:
searches = [('step1', RandomizedSearchCV, step1_esn_params, kwargs_step1),
('step2', RandomizedSearchCV, step2_esn_params, kwargs_step2),
('step3', GridSearchCV, step3_esn_params, kwargs_step3),
('step4', RandomizedSearchCV, step4_esn_params, kwargs_step4)]
base_esn = SeqToSeqESNClassifier(input_to_node=PredefinedWeightsInputToNode(predefined_input_weights=w_in),
**initially_fixed_params)
# -
# ## Optimization
#
# We provide a SequentialSearchCV that basically iterates through the list of searches that we have defined before. It can be combined with any model selection tool from scikit-learn.
try:
sequential_search = load("sequential_search.joblib")
except FileNotFoundError:
print(FileNotFoundError)
sequential_search = SequentialSearchCV(base_esn, searches=searches).fit(X_train, y_train)
dump(sequential_search, "sequential_search.joblib")
# ## Visualize hyper-parameter optimization
#
# ### First optimization step: input scaling and spectral radius
#
# Either create a scatterplot - useful in case of a random search to optimize input scaling and spectral radius
# +
df = pd.DataFrame(sequential_search.all_cv_results_["step1"])
fig = plt.figure()
ax = sns.scatterplot(x="param_spectral_radius", y="param_input_scaling", hue="mean_test_score", palette='RdBu', data=df)
plt.xlabel("Spectral Radius")
plt.ylabel("Input Scaling")
norm = plt.Normalize(0, df['mean_test_score'].max())
sm = plt.cm.ScalarMappable(cmap="RdBu", norm=norm)
sm.set_array([])
plt.xlim((0, 2.05))
plt.ylim((0, 1.05))
# Remove the legend and add a colorbar
ax.get_legend().remove()
ax.figure.colorbar(sm)
fig.set_size_inches(4, 2.5)
tick_locator = ticker.MaxNLocator(5)
ax.yaxis.set_major_locator(tick_locator)
ax.xaxis.set_major_locator(tick_locator)
# -
# Or create a heatmap - useful in case of a grid search to optimize input scaling and spectral radius
# +
df = pd.DataFrame(sequential_search.all_cv_results_["step1"])
pvt = pd.pivot_table(df,
values='mean_test_score', index='param_input_scaling', columns='param_spectral_radius')
pvt.columns = pvt.columns.astype(float)
pvt2 = pd.DataFrame(pvt.loc[pd.IndexSlice[0:1], pd.IndexSlice[0.0:1.0]])
fig = plt.figure()
ax = sns.heatmap(pvt2, xticklabels=pvt2.columns.values.round(2), yticklabels=pvt2.index.values.round(2), cbar_kws={'label': 'Score'})
ax.invert_yaxis()
plt.xlabel("Spectral Radius")
plt.ylabel("Input Scaling")
fig.set_size_inches(4, 2.5)
tick_locator = ticker.MaxNLocator(10)
ax.yaxis.set_major_locator(tick_locator)
ax.xaxis.set_major_locator(tick_locator)
# -
# ### Second optimization step: leakage
df = pd.DataFrame(sequential_search.all_cv_results_["step2"])
fig = plt.figure()
fig.set_size_inches(2, 1.25)
ax = sns.lineplot(data=df, x="param_leakage", y="mean_test_score")
ax.set_xscale('log')
plt.xlabel("Leakage")
plt.ylabel("Score")
plt.xlim((1e-5, 1e0))
tick_locator = ticker.MaxNLocator(10)
ax.xaxis.set_major_locator(tick_locator)
ax.yaxis.set_major_formatter(ticker.FormatStrFormatter('%.4f'))
plt.grid()
# ### Third optimization step: bias_scaling
df = pd.DataFrame(sequential_search.all_cv_results_["step3"])
fig = plt.figure()
fig.set_size_inches(2, 1.25)
ax = sns.lineplot(data=df, x="param_bias_scaling", y="mean_test_score")
plt.xlabel("Bias Scaling")
plt.ylabel("Score")
plt.xlim((0, 1))
tick_locator = ticker.MaxNLocator(5)
ax.xaxis.set_major_locator(tick_locator)
ax.yaxis.set_major_formatter(ticker.FormatStrFormatter('%.5f'))
plt.grid()
# ### Fourth optimization step: alpha (regularization)
df = pd.DataFrame(sequential_search.all_cv_results_["step4"])
fig = plt.figure()
fig.set_size_inches(2, 1.25)
ax = sns.lineplot(data=df, x="param_alpha", y="mean_test_score")
ax.set_xscale('log')
plt.xlabel("Alpha")
plt.ylabel("Score")
plt.xlim((1e-5, 1e0))
tick_locator = ticker.MaxNLocator(5)
ax.xaxis.set_major_locator(tick_locator)
ax.yaxis.set_major_formatter(ticker.FormatStrFormatter('%.5f'))
plt.grid()
# ## Test the ESN
#
# Finally, we test the ESN on unseen data.
y_pred = esn.predict(X_train)
y_pred_proba = esn.predict_proba(X_train)
| examples/sklearn_autoencoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7v0Bo63NffSK"
# A API utilizada neste artigo é um pacote chamado yahooquery, criado pelo desenvolvedor <NAME>. Desde o cancelamento das APIs oficiais do Google e Yahoo, a comunidade de ciência de dados vem criando maneiras de extrair informações do mercado financeiro. Essa API não-oficial é uma maneira alternativa de se obter tais dados.
# + [markdown] id="KYxOMPdJdK9o"
# 1. Instalando biblioteca
# + id="q7-nlTuXYRkA"
# !pip install yahooquery
# + [markdown] id="DVX3hn-8dZ3Q"
# 2. Importando biblioteca
# + id="WXCj39JDbi1p"
from yahooquery import Ticker
# + [markdown] id="3aUaFbhzf3f4"
# 3. Extraindo dados diários
#
# Um ponto muito importante, os tickers (siglas que identificam as ações) usados pelo Yahoo Finance são um pouco diferentes dos oficiais da Bovespa. Antes de fazer a consulta, é preciso conferir no site qual é o ticker correto,confira alguns exemplos abaixo:
#
# | Oficial | Yahoo Finance |
# |-----------|-----------------|
# | PETR4 | PETR4.SA |
# | ABEV3 | ABEV3.SA |
# | MGLU3 | MGLU3.SA |
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="JAtzaF0Gb_uS" outputId="c7f9d5b6-5048-47b8-f0db-89f4bf1687fe"
CIEL3 = Ticker("CIEL3.SA")
CIEL3.history(period='max')
# + [markdown] id="l1KHSTbihWzl"
# Consultando um intervalo de tempo específico, adicionando datas de início e fim aos parâmetros:
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="wBVIOIIwcAUS" outputId="87a7109b-be95-4f5f-9fad-2bf2c2eda90b"
CIEL3.history(start='2001-01-01', end='2021-04-22')
# + [markdown] id="sEFGVeiIhgD7"
# 4. Extraindo dados intraday
#
# Para dados intraday, o período disponível varia de acordo com o intervalo escolhido. Os intervalos suportados são 30 min, 15 min, 5 min, 2 min e 1 min.
#
# | Intervalos intraday | |
# |--------------------------------| |
# | 30 min | últimos 60 dias úteis |
# | 15 min | últimos 60 dias úteis |
# | 5 min | últimos 60 dias úteis |
# | 2 min | últimos 31 dias úteis |
# | 1 min | últimos 7 dias úteis |
#
# Extraindo intervalos de 30 min:
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="3Lv2dRhicSyI" outputId="34f6aafa-7313-415b-b06b-ae6001d0d501"
CIEL3 = Ticker('CIEL3.SA')
CIEL3.history(period='60d', interval = "30m")
# + [markdown] id="egYMtcpWixCp"
# Extraindo intervalos de 1 minuto:
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="omleAovBcvho" outputId="addf4fcb-29c4-4b04-d611-7e4b18391f2c"
CIEL3 = CIEL3.history(period='7d', interval = "1m")
CIEL3
# + [markdown] id="FpnsjwTIdJKV"
# 4. Informações financeiras
#
# Extraindo dados financeiros para análise fundamentalista. Para isso, são retornados os principais indicadores presentes nos relatórios anuais das empresas, tais como receita, lucro, Ebit, gastos com pesquisa e inovação, etc.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="fECTT99XdI0N" outputId="629fc588-ff85-4e65-bd52-7876b8bc5959"
CIEL3 = Ticker("CIEL3.SA") # Coleta dados
CIEL3.income_statement() # Chama função de Demonstração de resultados
CIEL3 = CIEL3.income_statement().transpose() # Transpõe a matriz
CIEL3.columns = CIEL3.iloc[0,:] # Renomeia colunas
CIEL3 = CIEL3.iloc[2:,:-1] # Seleciona dados
CIEL3 = CIEL3.iloc[:, ::-1] # Inverte colunas
CIEL3
| Extrair_dados_ticker_bovespa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Start your first Spark Session 🎈
# Findspark helps us to **find** and **start** Spark
import findspark
# +
# Provide the path where you installed Spark
findspark.init("c:\spark")
# +
# Create Spark Sesion
from pyspark.sql import SparkSession
# +
# This piece of code will generate a Spark session, however i is not named or configured.
# In real-life, best practice is to create tailored and named sessions hat works best for your case.
spark = SparkSession.builder.getOrCreate()
# -
# Check Spark version
spark.version
# From this point, we have created our Spark Session. It is ready for usage. To make sure your session is created successfully, go to http://localhost:4040/jobs/
#
# From this page, you can see and manage all the jobs you have created!
| PySpark_CreateSession_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BPE vocabulary analysis
#
# (C) <NAME>, 2019, Mindscan
#
# Recently I build some bpe-vocabularies, which had way too much tokens because of arabic and asian languages. This notebook is intended to review these vocabularies...
#
# Current state: the vocabulary was cleaned up... So the initial reason for this notebook is no longer given.
#
import sys
sys.path.insert(0,'../src')
import functools
from de.mindscan.fluentgenesis.bpe.bpe_model import BPEModel
model = BPEModel("16K-full", "../src/de/mindscan/fluentgenesis/bpe/")
model.load_hparams()
model_vocabulary = model.load_tokens()
model_bpe_data = model.load_bpe_pairs()
# +
model_vocabulary_statistics_length = {}
for token, _ in model_vocabulary.items():
lengthOfToken = len(token)
if lengthOfToken not in model_vocabulary_statistics_length:
model_vocabulary_statistics_length[lengthOfToken] = 1
else:
model_vocabulary_statistics_length[lengthOfToken] += 1
print(model_vocabulary_statistics_length)
# -
# The result is something like this:
#
# {1: 24804, 2: 1462, 3: 2465, 4: 2213, 5: 1906, 6: 1639, 7: 1426, 8: 1255, 9: 971, 10: 738, 11: 505, 12: 349, 13: 277, 14: 202, 15: 156, 16: 107, 17: 84, 18: 64, 19: 44, 20: 35, 21: 22, 22: 21, 23: 8, 24: 9, 25: 13, 26: 7, 27: 3, 28: 3, 29: 2, 30: 4, 32: 2, 33: 2, 34: 1, 36: 1, 37: 1, 42: 1, 43: 1, 51: 1}
#
# The intended dictionary size was about 16K tokens...
#
functools.reduce( lambda sum,y: sum + model_vocabulary_statistics_length[y] ,model_vocabulary_statistics_length.keys() )
# The result for all tokenlength longer than one char is 16001 (oops, off by one in token calculation...) we can see, that we have another 24804 Tokens of length one. Since we might can not encode every words with pairs, and to takle the unknown word problem, we have to emit all unpaired tokens at the end of the process. There are nearly twice the number of unpaired tokens than paired tokens...
#
# These originate from strings containing other languages, so lets identify these
#
#
#
# +
import collections
unsupported_vocab_ranges = [
# Latin Extended
(0x0100, 0x017F), # Latin Extended-A
(0x0180, 0x024F), # Latin Extended-B
(0x1E00, 0x1EFF), # Latin Extended Additional
(0x2C60, 0x2C7F), # Latin Extended-C
(0xA720, 0xA7FF), # Latin Extended-D
(0xAB30, 0xAB6F), # Latin Extended-E
# Diacritical
(0x0300, 0x036F), # Combining Diacritical Marks
(0x1AB0, 0x1AFF), # Combining Diacritical Marks Extended
(0x1DC0, 0x1DFF), # Combining Diacritical Marks Supplement
(0x20D0, 0x20FF), # Combining Diacritical Marks for Symbols
# IPA & phonetic Extemsion
(0x0250, 0x02AF), # IPA Extensions
(0x1D00, 0x1D7F), # Phonetic Extensions
(0x1D80, 0x1DBF), # Phonetic Extensions Supplement
# Spacing Modifier Letters
(0x02B0, 0x02FF), # Spacing Modifier Letters
# Greek Coptic
(0x0370, 0x03FF), # Greek and coptic
(0x1F00, 0x1FFF), # Greek Extended
(0x2C80, 0x2CFF), # Coptic
# Cyrillic
(0x0400, 0x04FF), # Cyrillic)
(0x0500, 0x052F), # Cyrillic Supplement
(0x2DE0, 0x2DFF), # Cyrillic Extended-A
(0xA640, 0xA69F), # Cyrillic Extended-B
(0x1C80, 0x1C8F), # Cyrillic Extended-C
# Armenian
(0x0530, 0x058F), # Armenian
# Hebrew
(0x0590, 0x05FF), # Hebrew
# Arabic
(0x0600, 0x06FF), # Arabic
(0x0750, 0x077F), # Arabic Supplement
(0x08A0, 0x08FF), # Arabic Extended-A
(0xFB50, 0xFDFF), # Arabic Presentation Forms A
(0xFE70, 0xFEFF), # Arabic Presentation Forms B
# Syriac
(0x0700, 0x074F), # Syriac
(0x0860, 0x086F), # Syriac Supplement
# Thaana
(0x0780, 0x07BF), # Thaana
# NKo
(0x07C0, 0x07FF), # NKo
# Samritan
(0x0800, 0x083F), # Samaritan
# Mandaic
(0x0840, 0x085F), # Mandaic
# Invalid range
(0x0870, 0x089F), # Invalid range
# Indian Subkontinent Languages
(0x0900, 0x097F), # Devanagari
(0xA8E0, 0xA8FF), # Devanagari Extended
# (0x0980, 0x09FF), # Bengali
# ...
(0x0900, 0x0DFF), # India - covers multiple languages
(0xA830, 0xA83F), # Common Indic Number Forms
(0x0E00, 0x0E7F), # Thai
(0x0E80, 0x0EFF), # Lao
(0x0F00, 0x0FFF), # Tibetan
# Myanmar
(0x1000, 0x109F), # Myanmar
(0xAA60, 0xAA7F), # Myanmar Extended-A
(0xA9E0, 0xA9FF), # Myanmar Extended-B
# Georgian
(0x10A0, 0x10FF), # Georgian
(0x2D00, 0x2D2F), # Georgian Supplement
(0x1C90, 0x1CBF), # Georgian Extended
# Korean
(0x1100, 0x11FF), # Hangul Jamo
(0x3130, 0x318F), # Hangul Compatibility Jamo
(0xAC00, 0xD7AF), # Hangul Syllables
(0xA960, 0xA97F), # Hangul Jamo Extended-A
(0xD7B0, 0xD7FF), # Hangul Jamo Extended B
# Ethiopic
(0x1200, 0x139f), # Ethiopic, Ethopic Supplement
(0xAB00, 0xAB2F), # Ethiopic Extended-A
(0x2D80, 0x2DDF), # Ethiopic Extended
# Cherokee
(0x13A0, 0x13FF), # Cherokee
(0xAB70, 0xABBF), # Cherokee Supplement
# Canadian Aboriginal
(0x1400, 0x167F), # Unified Canadian Aboriginal Syllabics
(0x18B0, 0x18FF), # Unified Canadian Aboriginal Syllabics Extended
(0x1680, 0x169F), # Ogham
(0x16A0, 0x16FF), # Runic
(0x1700, 0x171F), # Tagalog
(0x1720, 0x173F), # Hanunoo
(0x1740, 0x175F), # Buhid
(0x1760, 0x177F), # Tagbanwa
# Khmer
(0x1780, 0x17FF), # Khmer
(0x19E0, 0x19FF), # Khmer Symbols
# Mongolian
(0x1800, 0x18AF), # Mongolian
(0x1900, 0x194F), # Limbu
(0x1950, 0x197F), # Tai Le
(0x1980, 0x19DF), # New Tai Lue
(0x1A00, 0x1A1F), # Buginese
(0x1A20, 0x1AAF), # Tai Tham
(0x1B00, 0x1B7F), # Balinese
(0x1B80, 0x1BBF), # Sundanese
(0x1CC0, 0x1CCF), # Sundanese Supplement
(0x1BC0, 0x1BFF), # Batak
(0x1C00, 0x1C4F), # Lepcha
(0x1C50, 0x1C7F), # Ol Chiki
(0x1CD0, 0x1CFF), # Vedic Extensions
# Punctuation
(0x2000, 0x206F), # General Punctuation
(0x2E00, 0x2E7F), # Supplemental Punctuation
(0x3000, 0x303F), # CJK Symbols and Punctuation
(0x2070, 0x209F), # Superscripts and Subscripts
(0x20A0, 0x20CF), # Currency Symbols
# Symbols
(0x2100, 0x26ff), # Letterlike Symbols, ... Miscelaneous Symbols
(0x2700, 0x27FF), # Dingbats & co
(0x2800, 0x28FF), # Braille
(0x2900, 0x2BFF), # Symbols Arrows math
(0x2D30, 0x2D7F), # Tifinagh
(0x2f00, 0x2FFF), # Kangxi radicals, Ideographic Description Characters
# CJK
(0x3000, 0xa4FF),
(0xFE30, 0xFE4F), # CJK Compatibility Forms
(0xF900, 0xFAFF), # CJK Compatibility Ideographs
(0x2E80, 0x2EFF), # CJK Radicals Supplement
#
(0xA500, 0xA63F), # Vai
(0xA6A0, 0xA6FF), # Bamum
(0xA700, 0xA71F), # Modifier Tone Letters
(0xA800, 0xA82F), # <NAME>
(0xA840, 0xA87F), # Phags-pa
(0xA880, 0xA8DF), # Saurashtra
(0xA900, 0xA92F), # Kayah Li
(0xA930, 0xA95F), # Rejang
(0xA980, 0xA9DF), # Javanese
(0xAA00, 0xAA5F), # Cham
(0xAA80, 0xAADF), # Tai Viet
(0xABC0, 0xABFF), # Meetei Mayek
(0xAAE0, 0xAAFF), # Meetei Mayek Extensions
# Private Area
(0xE000,0xF8FF), # Private Use Area
(0xD800, 0xDB7F), # High Surrogates
(0xDB80, 0xDBFF), # High Private Use Surrogates
(0xDC00, 0xDFFF), # Low Surrogates
# # ???
(0x2c00, 0x2c5f), # Glagolitic
(0xFB00, 0xFB4F), # Alphabetic Presentation Forms
(0xFE00, 0xFE0F), # Variation Selectors
(0xFE10, 0xFE1F), # Vertical Forms
(0xFE20, 0xFE2F), # Combining Half Marks
(0xFE50, 0xFE6F), # Small Form Variants
(0xFF00, 0xFFEF), # Halfwidth and Fullwidth Forms
# OLD and OLDER
(0x010000, 0x10FFFF) # Basically everything what is not as important to be in first ~65000 Codes
]
def is_unsupported_character(char):
char=ord(char)
for bottom, top in unsupported_vocab_ranges:
if char >= bottom and char <= top:
return True
return False
one_char_elements = filter(lambda x: len(x)==1,model_vocabulary )
chars = filter(lambda char: not(is_unsupported_character(char)), one_char_elements)
charsasArray = [x for x in chars]
orderedChars = sorted(charsasArray, key=lambda item: item)
print(len(orderedChars))
print(orderedChars)
print(["0x%x"%ord(item) for item in orderedChars])
# -
| ipynb/vocabulary_analyzer_0x01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import cPickle as pickle
import scipy.io
import time
import ssn
import ks_test3
from hyperopt import fmin, tpe, hp, STATUS_OK, Trials
# %matplotlib inline
# -
# Define Hyperopt search space:
space = [hp.uniform('sig_EE',7,9),
hp.uniform('sig_IE',10,16),
hp.uniform('sig_EI',3,5),
hp.uniform('sig_II',3,5),
hp.uniform('J_EI',0.089,0.105),
hp.uniform('J_II',0.08,0.105)]
# +
# load Blasdel orientation and ocular dominance maps (previously processed,
# see map_analysis.ipynb
st = time.time()
[OD_map_full, OP_map_full] = np.load('saved_vars/maps-Nov-7.p', 'rb')
print "Elapsed time to load maps: %d seconds" % (time.time() - st)
# plt.figure()
# plt.imshow(OD_map_full)
# plt.colorbar()
# plt.title('Full ocular dominance map, Obermayer and Blasdel')
# plt.figure()
# plt.imshow(OP_map_full)
# plt.colorbar()
# plt.title('Full orientation map, Obermayer and Blasdel')
OD_map = OD_map_full[-75:,-75:]
OP_map = np.floor(OP_map_full[-75:,-75:])
# +
n_units = 50
selected_units = np.floor( ss_net.N_pairs*np.random.rand(n_units, 2) )
OD_prefs = np.zeros(len(selected_units))
for i in range(len(selected_units)):
xi = selected_units[i,0]
yi = selected_units[i,1]
OD_prefs[i] = OD_map[yi,xi]
# -
# Define objective funtion for hyperopt:
def iot_ssn_ks2d(args):
sig_EE, sig_IE, sig_EI, sig_II, J_EI, J_II = args
# Generate SSN with specified hyperparams:
ss_net = ssn.SSNetwork(sig_EE, sig_IE, sig_EI, sig_II, J_EE=0.1, J_IE=0., J_EI, J_II, OP_map=OP_map, OD_map=OD_map)
# TODO: Check the stability of the network and abort if unstable (return high value)
c = 40
dt = 0.005
timesteps = 100
dx = ss_net.dx
N_pairs = ss_net.N_pairs
# first find the summation field size (optimal CRF stimulus) for each unit (both E and I)
stim_sizes = np.linspace(0.75, 2, 5)
crf_bank = np.zeros( (n_units, 2, len(stim_sizes), N_pairs, N_pairs) )
for i in range(n_units):
xi = selected_units[i,0]
yi = selected_units[i,1]
ocularity = np.round( OD_map[yi,xi] )
ori = OP_map[yi,xi]
for j in range(len(stim_sizes)):
crf_bank[i,0,j,:,:] = ssn.generate_mono_stimulus( ori, stim_sizes[j], [dx*xi, dx*yi], OP_map )
crf_bank[i,1,j,:,:] = ssn.generate_ext_stimulus( ori, stim_sizes[j], [dx*xi, dx*yi], OP_map, OD_map, ocularity)
# Store the summation field sizes (SFS) for both E and I units
sfs_E = np.zeros( n_units )
sfs_I = np.copy(sfs_E)
max_fr_E = np.copy(sfs_E)
max_fr_I = np.copy(sfs_E)
# run to find monocular SFS:
for i in range(n_units):
xi = selected_units[i,0]
yi = selected_units[i,1]
e_found = False
i_found = False
for j in range(len(stim_sizes)):
if e_found == True and i_found == True:
break
h = crf_bank[i,1,j,:,:]
[r_E, r_I, I_E, I_I] = ss_net.run_simulation(dt, timesteps, c, h )
if r_E[-1,yi,xi] >= max_fr_E[i]:
max_fr_E[i] = r_E[-1,yi,xi]
sfs_E[i] = stim_sizes[j]
else:
e_found = True
if r_I[-1,yi,xi] >= max_fr_I[i]:
max_fr_I[i] = r_I[-1,yi,xi]
sfs_I[i] = stim_sizes[j]
else:
i_found = True
# Generate non-dominant CRF stimuli
non_dom_stimuli = np.zeros((len(selected_units), 2, N_pairs, N_pairs))
for i in range(len(selected_units)):
xi = selected_units[i,0]
yi = selected_units[i,1]
ocularity = np.abs( np.round(OD_prefs[i]) - 1)
non_dom_stimuli[i,0,:,:] = ssn.generate_ext_stimulus( ori, sfs_E[i], [dx*xi, dx*yi], OP_map, OD_map, ocularity)
if sfs_E[i] != sfs_I[i]:
non_dom_stimuli[i,1,:,:] = ssn.generate_ext_stimulus( ori, sfs_I[i], [dx*xi, dx*yi], OP_map, OD_map, ocularity)
non_dom_results = np.zeros((len(selected_units), 2))
for i in range(len(selected_units)):
xi = selected_units[i,0]
yi = selected_units[i,1]
h = non_dom_stimuli[i,0,:,:]
[r_E, r_I, I_E, I_I] = ss_net.run_simulation(dt, timesteps, c, h )
non_dom_results[i,0] = r_E[-1,yi,xi]
non_dom_results[i,1] = r_I[-1,yi,xi]
if sfs_E[i] != sfs_I[i]:
h = non_dom_stimuli[i,1,:,:]
[r_E, r_I, I_E, I_I] = ss_net.run_simulation(dt, timesteps, c, h )
non_dom_results[i,1] = r_I[-1,yi,xi]
threshold = 1 # threshold for Webb's "reliable response" criterion
# Only carry on with units whose non-dom CRF response is above the threshold:
thresh_units_E = selected_units[np.where(non_dom_results[:,0]>=threshold),:][0]
thresh_units_I = selected_units[np.where(non_dom_results[:,1]>=threshold),:][0]
thresh_units_sfs_E = sfs_E[np.where(non_dom_results[:,0]>=threshold)]
thresh_units_sfs_I = sfs_I[np.where(non_dom_results[:,1]>=threshold)]
thresh_units_max_fr_E = max_fr_E[np.where(non_dom_results[:,0]>=threshold)]
thresh_units_max_fr_I = max_fr_I[np.where(non_dom_results[:,1]>=threshold)]
# Now find which units which are above threshold also suppress below 90% with non-dom surround:
non_dom_surround_stim_E = np.zeros((len(thresh_units_E), N_pairs, N_pairs))
dom_surround_stim_E = np.copy(non_dom_surround_stim_E)
dom_crf_stim_E = np.copy(non_dom_surround_stim_E)
for i in range(len(thresh_units_E)):
xi = thresh_units_E[i,0]
yi = thresh_units_E[i,1]
inner_d = thresh_units_sfs_E[i]
outer_d = inner_d + 3
centre = [dx*xi, dx*yi]
ocularity = np.abs( np.round(OD_map[yi,xi]) - 1)
non_dom_surround_stim_E[i] = ssn.generate_ring_stimulus(OP_map[yi,xi], inner_d, outer_d, centre, ocularity, OP_map, OD_map)
dom_surround_stim_E[i] = ssn.generate_ring_stimulus(OP_map[yi,xi], inner_d, outer_d, centre, np.round(OD_map[yi,xi]), OP_map, OD_map)
dom_crf_stim_E[i] = ssn.generate_ext_stimulus( ori, inner_d, [dx*xi, dx*yi], OP_map, OD_map, np.round(OD_map[yi,xi]) )
# Run simulations to analyze non dominant suppression:
non_dom_surround_results = np.zeros((len(thresh_units_E)))
dom_surround_results = np.copy(non_dom_surround_results)
for i in range(len(thresh_units_E)):
xi = thresh_units_E[i,0]
yi = thresh_units_E[i,1]
h = non_dom_surround_stim_E[i] + dom_crf_stim_E[i]
[r_E, r_I, I_E, I_I] = ss_net.run_simulation(dt, timesteps, c, h )
non_dom_surround_results[i] = r_E[-1,yi,xi]
h = dom_surround_stim_E[i] + dom_crf_stim_E[i]
[r_E, r_I, I_E, I_I] = ss_net.run_simulation(dt, timesteps, c, h )
dom_surround_results[i] = r_E[-1,yi,xi]
dominant_SI_E = (thresh_units_max_fr_E - dom_surround_results) / thresh_units_max_fr_E
non_dom_SI_E = (thresh_units_max_fr_E - non_dom_surround_results) / thresh_units_max_fr_E
# Now do all the same stuff for the I units:
non_dom_surround_stim_I = np.zeros((len(thresh_units_I), N_pairs, N_pairs))
dom_surround_stim_I = np.copy(non_dom_surround_stim_I)
dom_crf_stim_I = np.copy(non_dom_surround_stim_I)
for i in range(len(thresh_units_I)):
xi = thresh_units_I[i,0]
yi = thresh_units_I[i,1]
inner_d = thresh_units_sfs_I[i]
outer_d = inner_d + 3
centre = [dx*xi, dx*yi]
ocularity = np.abs( np.round(OD_map[yi,xi]) - 1)
non_dom_surround_stim_I[i] = ssn.generate_ring_stimulus(OP_map[yi,xi], inner_d, outer_d, centre, ocularity, OP_map, OD_map)
dom_surround_stim_I[i] = ssn.generate_ring_stimulus(OP_map[yi,xi], inner_d, outer_d, centre, np.round(OD_map[yi,xi]), OP_map, OD_map)
dom_crf_stim_I[i] = ssn.generate_ext_stimulus( ori, inner_d, [dx*xi, dx*yi], OP_map, OD_map, np.round(OD_map[yi,xi]))
# Run simulations to analyze non dominant suppression:
non_dom_surround_results_I = np.zeros((len(thresh_units_I)))
dom_surround_results_I = np.copy(non_dom_surround_results_I)
for i in range(len(thresh_units_I)):
xi = thresh_units_I[i,0]
yi = thresh_units_I[i,1]
h = non_dom_surround_stim_I[i] + dom_crf_stim_I[i]
[r_E, r_I, I_E, I_I] = ss_net.run_simulation(dt, timesteps, c, h )
non_dom_surround_results_I[i] = r_I[-1,yi,xi]
h = dom_surround_stim_I[i] + dom_crf_stim_I[i]
[r_E, r_I, I_E, I_I] = ss_net.run_simulation(dt, timesteps, c, h )
dom_surround_results_I[i] = r_I[-1,yi,xi]
dominant_SI_I = (thresh_units_max_fr_I - dom_surround_results_I) / thresh_units_max_fr_I
non_dom_SI_I = (thresh_units_max_fr_I - non_dom_surround_results_I) / thresh_units_max_fr_I
# Concatenate the E and I results
model_data_x = np.concatenate((dominant_SI_E, dominant_SI_I))
model_data_y = np.concatenate((non_dom_SI_E, non_dom_SI_I))
webb_data = np.array([[0.3538, 0.3214],
[0.5513, 0.2271],
[0.5154, 0.5064],
[0.5641, 0.5681],
[0.6077, 0.5605],
[0.7179, 0.6172],
[0.7487, 0.6865],
[0.8282, 0.6406],
[0.8923, 0.5459],
[0.9282, 0.5690],
[0.6308, 0.4093],
[0.7385, 0.4557],
[0.7923, 0.4866],
[0.7385, 0.5352],
[0.9974, 0.9846]])
d, prob = ks_test3.ks2d2s(webb_data[:,0], webb_data[:,1], model_data_x, model_data_y)
return {
'status': 'ok',
'loss':, 1-prob,
'attachments': {'units_probed':pickle.dumps([thresh_units_E, thresh_units_I, thresh_untits_max_fr_E, thresh_units_max_fr_I, dom_surround_results, dom_surround_results_I, sfs_E, sfs_I])}
}
# +
# create a Trials database to store experiment results:
trials = Trials()
st = time.time()
best = fmin(iot_ssn_ks2d, space, algo=tpe.suggest, max_evals=10, trials=trials)
print "Elapsed time for 10 hyperopt sims: %d seconds." % (time.time()-st)
print 'tpe:', best
| mechanistic/IOT_hyperopt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # K-Means Clustering
# ## <NAME>
# ### Importing the required libraries
import numpy as np
import matplotlib.pyplot as plt
import random
from sklearn.cluster import KMeans
from sklearn.datasets.samples_generator import make_blobs
# ### Creating the dataset
np.random.seed(0)
X, y = make_blobs(n_samples=5000, centers=[[4,4], [-2, -1], [2, -3], [1, 1]], cluster_std=0.9)
# The **make_blobs** class can take in many inputs, but we will be using these specific ones.
#
# **Input**
# - **n_samples**: The total number of points equally divided among clusters.
# - Value will be: 5000
# - **centers**: The number of centers to generate, or the fixed center locations.
# - Value will be: [[4, 4], [-2, -1], [2, -3],[1,1]]
# - **cluster_std**: The standard deviation of the clusters.
# - Value will be: 0.9
#
# **Output**
# - **X**: Array of shape [n_samples, n_features]. (Feature Matrix)
# - The generated samples.
#
# - **y**: Array of shape [n_samples]. (Response Vector)
# - The integer labels for cluster membership of each sample.
#
plt.figure(figsize=(12, 8))
plt.scatter(X[:, 0], X[:, 1], marker='.')
# ### Modeling
k_means = KMeans(init = "k-means++", n_clusters = 4, n_init = 12)
k_means.fit(X)
# ### Cluster labels of each point
k_means_labels = k_means.labels_
k_means_labels
# ### Center points of each clusters
k_means_cluster_centers = k_means.cluster_centers_
k_means_cluster_centers
# ### Create a Visual Plot
# +
# Initialize the plot with the specified dimensions.
fig = plt.figure(figsize=(12, 8))
# Colors uses a color map, which will produce an array of colors based on
# the number of labels there are. We use set(k_means_labels) to get the
# unique labels.
# colors = plt.cm.Spectral(np.linspace(0, 1, len(set(k_means_labels))))
colors = ['red', 'green', 'blue', 'black']
# Create a plot
ax = fig.add_subplot(1, 1, 1)
# For loop that plots the data points and centroids.
# k will range from 0-3, which will match the possible clusters that each
# data point is in.
for k, col in zip(range(len([[4,4], [-2, -1], [2, -3], [1, 1]])), colors):
# Create a list of all data points, where the data poitns that are
# in the cluster (ex. cluster 0) are labeled as true, else they are
# labeled as false.
my_members = (k_means_labels == k)
# Define the centroid, or cluster center.
cluster_center = k_means_cluster_centers[k]
# Plots the datapoints with color col.
ax.plot(X[my_members, 0], X[my_members, 1], 'w', markerfacecolor=col, marker='.')
# Plots the centroids with specified color, but with a darker outline
ax.plot(cluster_center[0], cluster_center[1], 'o', markerfacecolor=col, markeredgecolor='k', markersize=6)
# Title of the plot
ax.set_title('KMeans')
# Remove x-axis ticks
ax.set_xticks(())
# Remove y-axis ticks
ax.set_yticks(())
# Show the plot
plt.show()
# -
# ### With 3 Clusters
# +
k_means = KMeans(init = "k-means++", n_clusters = 3, n_init = 12)
k_means.fit(X)
# -
k_means_labels = k_means.labels_
k_means_labels
k_means_cluster_centers = k_means.cluster_centers_
k_means_cluster_centers
# +
# Initialize the plot with the specified dimensions.
fig = plt.figure(figsize=(12, 8))
# Colors uses a color map, which will produce an array of colors based on
# the number of labels there are. We use set(k_means_labels) to get the
# unique labels.
# colors = plt.cm.Spectral(np.linspace(0, 1, len(set(k_means_labels))))
colors = ['red', 'green', 'blue']
# Create a plot
ax = fig.add_subplot(1, 1, 1)
# For loop that plots the data points and centroids.
# k will range from 0-3, which will match the possible clusters that each
# data point is in.
for k, col in zip(range(len([[4,4], [-2, -1], [2, -3], [1, 1]])), colors):
# Create a list of all data points, where the data poitns that are
# in the cluster (ex. cluster 0) are labeled as true, else they are
# labeled as false.
my_members = (k_means_labels == k)
# Define the centroid, or cluster center.
cluster_center = k_means_cluster_centers[k]
# Plots the datapoints with color col.
ax.plot(X[my_members, 0], X[my_members, 1], 'w', markerfacecolor=col, marker='.')
# Plots the centroids with specified color, but with a darker outline
ax.plot(cluster_center[0], cluster_center[1], 'o', markerfacecolor=col, markeredgecolor='k', markersize=6)
# Title of the plot
ax.set_title('KMeans')
# Remove x-axis ticks
ax.set_xticks(())
# Remove y-axis ticks
ax.set_yticks(())
# Show the plot
plt.show()
| 03_Clustering/01_K-Means/KMeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import pandas as pd
import datetime as datetime
from statsmodels.tsa.ar_model import AR
from random import random
from statsmodels.tsa.seasonal import seasonal_decompose
import os as os
import hvplot.pandas
import matplotlib.pylab as plt
flx_hh = pd.read_csv('FLUXNET/AMF_US-ARM_BASE-BADM_8-5/AMF_US-ARM_BASE_HH_8-5.csv',index_col=0, parse_dates=True, header = 2, na_values = -9999.0)
flx_daily=flx_hh.resample('1d').mean()
flx_daily.describe()
# +
# Additive Decomposition
result_add = seasonal_decompose(flx_daily.FC_1_1_1.fillna(method='pad'), model='additive', freq=365, extrapolate_trend='freq')
# Plot
#result_mul.plot().suptitle('Multiplicative Decompose', fontsize=22)
result_add.plot();
# -
flx_daily.FC_1_1_1.fillna(method='pad').hvplot()
# +
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
fig,axes = plt.subplots(1, 2, figsize = (10,4))
plot_pacf(result_add.resid, lags = 60, ax = axes[0])
plot_acf(result_add.resid, lags = 60, ax = axes[1]);
# -
| nbs/XXX_test_timeseries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.12 ('img')
# language: python
# name: python3
# ---
# +
#open ft_1024_samples.npz
import numpy as np
samples = np.load('ft_1024_samples.npz')
samples = samples['arr_0']
import PIL
from PIL import Image
#save all samples as images
for i in range(samples.shape[0]):
img = Image.fromarray(samples[i].astype('uint8'))
img.save('./fake/sample_' + str(i) + '.png')
# +
#open ft_1024_samples.npz
import numpy as np
import PIL
from PIL import Image
samples = np.load('samples_128x64x64x3.npz')
samples = samples['arr_0']
#save all samples as images
for i in range(samples.shape[0]):
img = Image.fromarray(samples[i].astype('uint8'))
img.save('./fakeb/sampleb_' + str(i) + '.png')
# -
import PIL
from PIL import Image
from tqdm import tqdm
sample_path = '/Volumes/Elements/sample'
import os
#loop through all files in sample_path
for file in tqdm(os.listdir(sample_path)):
try:
#crop the image to be a square centered but keep the original aspect ratio
img = Image.open(sample_path + '/' + file)
#center crop
img = img.crop(((img.size[0]-img.size[1])/2, 0, (img.size[0]+img.size[1])/2, img.size[1]))
#resize the image to 64x64
img = img.resize((64, 64))
#save the image
img.save('./sample_resize/' + file)
except:
#write the file name to a file if there is an error
#remove the file from the ./sample_resize folder
os.remove('./sample_resize/' + file)
with open('./sample_resize/error.txt', 'a') as f:
f.write(file + '\n')
# #remove images that are in the error file
# import os
# import shutil
# with open('./sample_resize/error.txt') as f:
# lines = f.readlines()
# for line in lines:
# os.remove('./sample_resize/' + line.strip())
sample_path = './sample_resize'
# !python -m pytorch_fid ./fake $sample_path
print('FID for fine-tuned samples^')
sample_path = './sample_resize'
# !python -m pytorch_fid ./fakeb $sample_path
print('FID for base model samples^')
file = "/Users/isaiahwilliams/classes/cs236/sample_resize/1078811.jpg"
img = Image.open(file)
img.show()
import os
from PIL import Image
folder_path = '/Users/isaiahwilliams/classes/cs236/sample_resize/'
extensions = []
for filee in os.listdir(folder_path):
file_path = os.path.join(folder_path, filee)
print('** Path: {} **'.format(file_path), end="\r", flush=True)
im = Image.open(file_path)
rgb_im = im.convert('RGB')
if filee.split('.')[1] not in extensions:
extensions.append(filee.split('.')[1])
| pre&post-process+FID.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from os.path import join
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.cluster import AgglomerativeClustering
from scipy.cluster.hierarchy import dendrogram
sns.set()
# -
# ## Import preprocessed data
df = pd.read_csv(join('..', 'data', 'tugas_preprocessed.csv'))
df.head()
df.columns
# Splitting feature names into groups
non_metric_features = df.columns[df.columns.str.startswith('x')]
pc_features = df.columns[df.columns.str.startswith('PC')]
metric_features = df.columns[~df.columns.str.startswith('x') & ~df.columns.str.startswith('PC')]
# ## Hierarchical Clustering
#
# What is hierarchical clustering? How does it work? How does it relate to the distance matrix we discussed at the beggining of the course? ;)
#
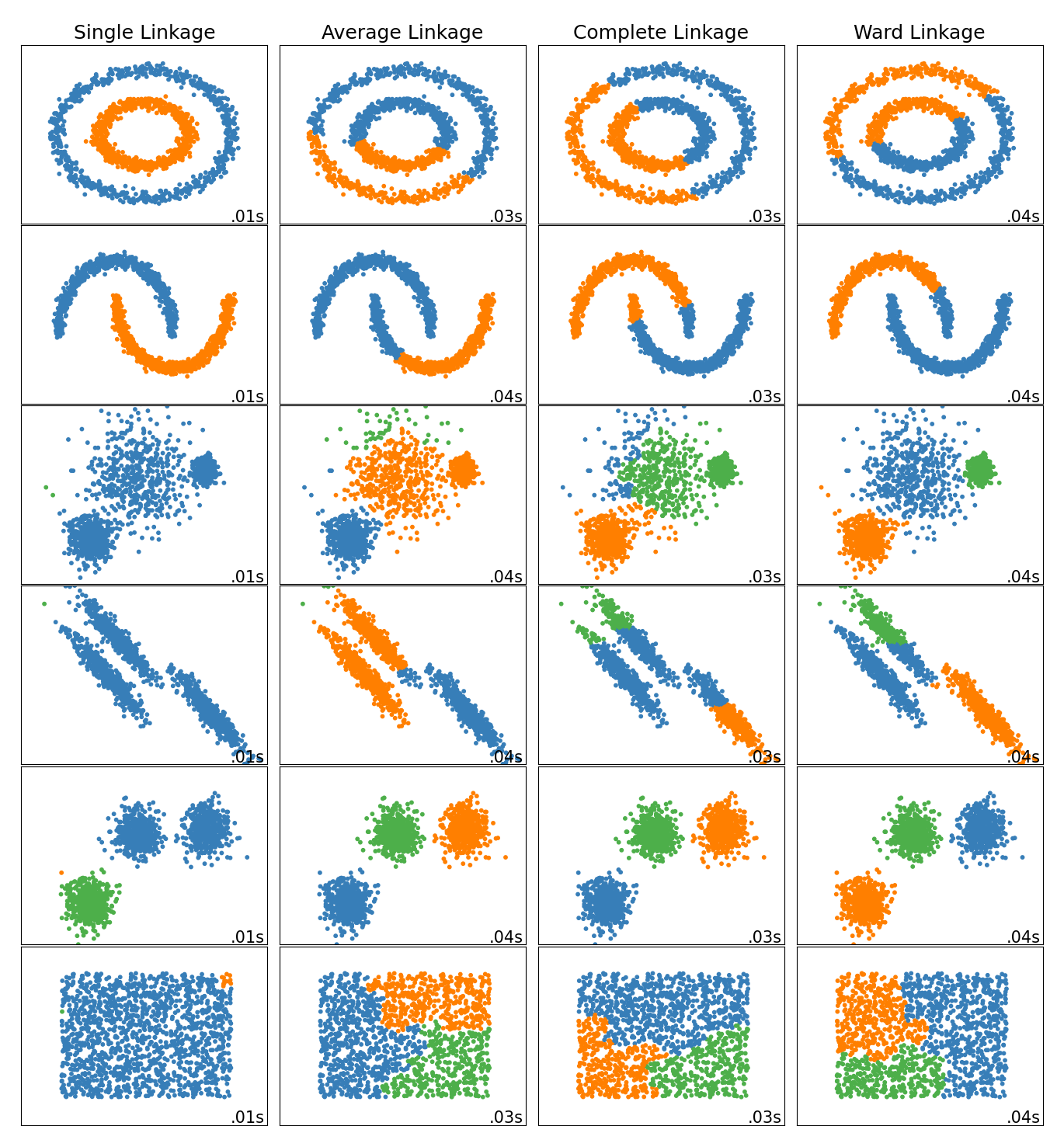
# ### Different types of linkage
# 
#
# ### How are they computed?
# 
#
# **Ward linkage**: minimizes the sum of squared differences within all clusters. It is a variance-minimizing approach and in this sense is similar to the k-means objective function but tackled with an agglomerative hierarchical approach.
#
# ### The distance matrix
# 
# ### Characteristics:
# - *bottom up approach*: each observation starts in its own cluster, and clusters are successively merged together
# - *greedy/local algorithm*: at each iteration tries to minimize the distance of cluster merging
# - *no realocation*: after an observation is assigned to a cluster, it can no longer change
# - *deterministic*: you always get the same answer when you run it
# - *scalability*: can become *very slow* for a large number of observations
# ### How to apply Hierarchical Clustering?
# **Note: Which types of variables should be used for clustering?**
# Performing HC
hclust = AgglomerativeClustering(linkage='ward', affinity='euclidean', n_clusters=5)
hc_labels = hclust.fit_predict(df[metric_features])
hc_labels
# Characterizing the clusters
df_concat = pd.concat((df, pd.Series(hc_labels, name='labels')), axis=1)
df_concat.groupby('labels').mean()
# ### Defining the linkage method to choose:
# **We need to understand that:**
# $$SS_{t} = SS_{w} + SS_{b}$$
#
# ---
#
# $$SS_{t} = \sum\limits_{i = 1}^n {{{({x_i} - \overline x )}^2}}$$
#
# $$SS_{w} = \sum\limits_{k = 1}^K {\sum\limits_{i = 1}^{{n_k}} {{{({x_i} - {{\overline x }_k})}^2}} }$$
#
# $$SS_{b} = \sum\limits_{k = 1}^K {{n_k}{{({{\overline x }_k} - \overline x )}^2}}$$
#
# , where $n$ is the total number of observations, $x_i$ is the vector of the $i^{th}$ observation, $\overline x$ is the centroid of the data, $K$ is the number of clusters, $n_k$ is the number of observations in the $k^{th}$ cluster and $\overline x_k$ is the centroid of the $k^{th}$ cluster.
# +
# Computing SST
X = df[metric_features].values
sst = np.sum(np.square(X - X.mean(axis=0)), axis=0)
# Computing SSW
ssw_iter = []
for i in np.unique(hc_labels):
X_k = X[hc_labels == i]
ssw_iter.append(np.sum(np.square(X_k - X_k.mean(axis=0)), axis=0))
ssw = np.sum(ssw_iter, axis=0)
# Computing SSB
ssb_iter = []
for i in np.unique(hc_labels):
X_k = X[hc_labels == i]
ssb_iter.append(X_k.shape[0] * np.square(X_k.mean(axis=0) - X.mean(axis=0)))
ssb = np.sum(ssb_iter, axis=0)
# Verifying the formula
np.round(sst) == np.round((ssw + ssb))
# -
def get_r2_hc(df, link_method, max_nclus, min_nclus=1, dist="euclidean"):
"""This function computes the R2 for a set of cluster solutions given by the application of a hierarchical method.
The R2 is a measure of the homogenity of a cluster solution. It is based on SSt = SSw + SSb and R2 = SSb/SSt.
Parameters:
df (DataFrame): Dataset to apply clustering
link_method (str): either "ward", "complete", "average", "single"
max_nclus (int): maximum number of clusters to compare the methods
min_nclus (int): minimum number of clusters to compare the methods. Defaults to 1.
dist (str): distance to use to compute the clustering solution. Must be a valid distance. Defaults to "euclidean".
Returns:
ndarray: R2 values for the range of cluster solutions
"""
def get_ss(df):
ss = np.sum(df.var() * (df.count() - 1))
return ss # return sum of sum of squares of each df variable
sst = get_ss(df) # get total sum of squares
r2 = [] # where we will store the R2 metrics for each cluster solution
for i in range(min_nclus, max_nclus+1): # iterate over desired ncluster range
cluster = AgglomerativeClustering(n_clusters=i, affinity=dist, linkage=link_method)
hclabels = cluster.fit_predict(df) #get cluster labels
df_concat = pd.concat((df, pd.Series(hclabels, name='labels')), axis=1) # concat df with labels
ssw_labels = df_concat.groupby(by='labels').apply(get_ss) # compute ssw for each cluster labels
ssb = sst - np.sum(ssw_labels) # remember: SST = SSW + SSB
r2.append(ssb / sst) # save the R2 of the given cluster solution
return np.array(r2)
# +
# Prepare input
hc_methods = ["ward", "complete", "average", "single"]
# Call function defined above to obtain the R2 statistic for each hc_method
max_nclus = 10
r2_hc_methods = np.vstack(
[
get_r2_hc(df=df[metric_features], link_method=link, max_nclus=max_nclus)
for link in hc_methods
]
).T
r2_hc_methods = pd.DataFrame(r2_hc_methods, index=range(1, max_nclus + 1), columns=hc_methods)
sns.set()
# Plot data
fig = plt.figure(figsize=(11,5))
sns.lineplot(data=r2_hc_methods, linewidth=2.5, markers=["o"]*4)
# Finalize the plot
fig.suptitle("R2 plot for various hierarchical methods", fontsize=21)
plt.gca().invert_xaxis() # invert x axis
plt.legend(title="HC methods", title_fontsize=11)
plt.xticks(range(1, max_nclus + 1))
plt.xlabel("Number of clusters", fontsize=13)
plt.ylabel("R2 metric", fontsize=13)
plt.show()
# -
# ### Defining the number of clusters:
# Where is the **first big jump** on the Dendrogram?
# setting distance_threshold=0 and n_clusters=None ensures we compute the full tree
linkage = 'ward'
distance = 'euclidean'
hclust = AgglomerativeClustering(linkage=linkage, affinity=distance, distance_threshold=0, n_clusters=None)
hclust.fit_predict(df[metric_features])
# +
# Adapted from:
# https://scikit-learn.org/stable/auto_examples/cluster/plot_agglomerative_dendrogram.html#sphx-glr-auto-examples-cluster-plot-agglomerative-dendrogram-py
# create the counts of samples under each node (number of points being merged)
counts = np.zeros(hclust.children_.shape[0])
n_samples = len(hclust.labels_)
# hclust.children_ contains the observation ids that are being merged together
# At the i-th iteration, children[i][0] and children[i][1] are merged to form node n_samples + i
for i, merge in enumerate(hclust.children_):
# track the number of observations in the current cluster being formed
current_count = 0
for child_idx in merge:
if child_idx < n_samples:
# If this is True, then we are merging an observation
current_count += 1 # leaf node
else:
# Otherwise, we are merging a previously formed cluster
current_count += counts[child_idx - n_samples]
counts[i] = current_count
# the hclust.children_ is used to indicate the two points/clusters being merged (dendrogram's u-joins)
# the hclust.distances_ indicates the distance between the two points/clusters (height of the u-joins)
# the counts indicate the number of points being merged (dendrogram's x-axis)
linkage_matrix = np.column_stack(
[hclust.children_, hclust.distances_, counts]
).astype(float)
# Plot the corresponding dendrogram
sns.set()
fig = plt.figure(figsize=(11,5))
# The Dendrogram parameters need to be tuned
y_threshold = 100
dendrogram(linkage_matrix, truncate_mode='level', p=5, color_threshold=y_threshold, above_threshold_color='k')
plt.hlines(y_threshold, 0, 1000, colors="r", linestyles="dashed")
plt.title(f'Hierarchical Clustering - {linkage.title()}\'s Dendrogram', fontsize=21)
plt.xlabel('Number of points in node (or index of point if no parenthesis)')
plt.ylabel(f'{distance.title()} Distance', fontsize=13)
plt.show()
# -
# ### Final Hierarchical clustering solution
# 4 cluster solution
linkage = 'ward'
distance = 'euclidean'
hc4lust = AgglomerativeClustering(linkage=linkage, affinity=distance, n_clusters=4)
hc4_labels = hc4lust.fit_predict(df[metric_features])
# Characterizing the 4 clusters
df_concat = pd.concat((df, pd.Series(hc4_labels, name='labels')), axis=1)
df_concat.groupby('labels').mean()
# 5 cluster solution
linkage = 'ward'
distance = 'euclidean'
hc5lust = AgglomerativeClustering(linkage=linkage, affinity=distance, n_clusters=5)
hc5_labels = hc5lust.fit_predict(df[metric_features])
# Characterizing the 5 clusters
df_concat = pd.concat((df, pd.Series(hc5_labels, name='labels')), axis=1)
df_concat.groupby('labels').mean()
| notebooks_solutions/lab09_hierarchical_clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.stats as stats
import pymc3 as pm
import arviz as az
import torch
import pyro
from pyro.optim import Adam
from pyro.infer import SVI, Trace_ELBO
import pyro.distributions as dist
from pyro.infer.autoguide import AutoDiagonalNormal, AutoMultivariateNormal, init_to_mean
# +
## Join data
features = pd.read_csv('data/dengue_features_train.csv')
labels = pd.read_csv('data/dengue_labels_train.csv')
df = features.copy()
df['total_cases'] = labels.total_cases
# Target is in column 'total_cases'
# -
df.head()
# +
cities = list(df.city.unique())
sj_df = df.loc[df.city=='sj']
data_df = sj_df.copy()
data_df = data_df.dropna()
ommit_cols = ('total_cases', 'city', 'year', 'weekofyear', 'week_start_date')
for c in data_df.columns:
if c not in ommit_cols:
m, std, mm = data_df[c].mean(), data_df[c].std(), data_df[c].max()
data_df[c] = (data_df[c]- m)/std
# +
plt.plot()
ax1 = data_df.total_cases.plot()
ax2 = ax1.twinx()
ax2.spines['right'].set_position(('axes', 1.0))
data_df.reanalysis_precip_amt_kg_per_m2.plot(ax=ax2, color='green')
plt.show()
# -
sub_df = data_df.loc[:]
with pm.Model() as simple_model:
b0 = pm.Normal("b0_intercept", mu=0, sigma=2)
b1 = pm.Normal("b1_variable", mu=0, sigma=2)
b2 = pm.Normal("b2_variable", mu=0, sigma=2)
b3 = pm.Normal("b3_variable", mu=0, sigma=2)
b4 = pm.Normal("b4_variable", mu=0, sigma=2)
θ = (
b0
+ b1 * sub_df.reanalysis_precip_amt_kg_per_m2
+ b2 * sub_df.station_diur_temp_rng_c
+ b3 * sub_df.reanalysis_max_air_temp_k
+ b4 * sub_df.station_precip_mm
)
y = pm.Poisson("y", mu=np.exp(θ), observed=sub_df.total_cases)
#start = {'b0_intercept': 5., 'b1_variable': 1., 'b2_variable': 1., 'b3_variable': 1., 'b4_variable': 1.}
with simple_model:
step = pm.Slice()
inf_model = pm.sample(10000, step=step, return_inferencedata=True,)
az.plot_trace(inf_model)
# ## Variational Inference approach
# +
torch.set_default_dtype(torch.float64)
cols = list(set(data_df.columns) - set(ommit_cols))
#cols = ['reanalysis_precip_amt_kg_per_m2', 'station_diur_temp_rng_c', 'reanalysis_max_air_temp_k', 'station_precip_mm']
x_data = torch.tensor(data_df[cols].values).float()
y_data = torch.tensor(data_df.total_cases.values).float()
# +
M = len(cols)
def model(x_data, y_data):
b = pyro.sample('b', dist.Normal(0.0, 2.0))
w = pyro.sample('w', dist.Normal(0.0, 2.0).expand([M]).to_event(1))
with pyro.plate('observe_data', size=len(y_data), subsample_size=100) as ind:
θ = (x_data.index_select(0, ind) * w).sum(axis=1) + b
pyro.sample('obs', dist.Poisson(θ.exp()), obs=y_data.index_select(0, ind))
guide = AutoMultivariateNormal(model, init_loc_fn=init_to_mean)
# +
pyro.clear_param_store()
adam = Adam({"lr": 0.001})
svi = SVI(model, guide, adam, loss=Trace_ELBO())
n_steps = 10000
losses = []
for step in range(n_steps):
losses.append(svi.step(x_data, y_data))
if step %1000 == 0:
print("Done with step {}".format(step))
# +
guide.requires_grad_(False)
for name, value in pyro.get_param_store().items():
print(name, pyro.param(name))
# -
plt.plot(losses)
plt.title('ELBO loss')
y_data.mean()
| Exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="KfJYMZwH2UfO"
# # Калькулятор краски
# -
# Введите последовательно требуемые параметры, следуя подсказкам программы (необходимо будет поочередно ввести значения - ширина поверхности, м; высота (длина) поверхности, м; расход краски кв.м / л.; объем банки в литрах, целое число; процент запаса, целое число) для получения результирующих показателей (площадь окрашивания, количество литров, количество банок, неиспользуемых литров краски).
# + colab={"base_uri": "https://localhost:8080/"} id="QE-FAQ2o2U9h" outputId="64e4c4ae-2f82-44fc-ffd8-4cc6c4bb854c"
print('Введите значение ширины поверхности в метрах: ')
surface_width = float(input()) # ширина поверхности, м
print('Введите значение высоты (длины) поверхности в метрах: ')
surface_height = float(input()) # высота (длина) поверхности, м
print('Введите значение расхода краски в кв.м / л.: ')
paints_in_meters_to_liter = float(input()) # расход краски кв.м / л.
print('Введите значение объема банки в литрах (целое число): ')
tin_of_paint_in_liters = int(input()) # объем банки в литрах, целое число
print('Введите значение процента запаса (целое число): ')
reserve_percent = int(input()) # процент запаса, целое число
surface_area = surface_width * surface_height # площадь окрашивания
number_of_liters = (surface_area / paints_in_meters_to_liter) * (1 + reserve_percent / 100) # количество литров
number_of_tins = int(number_of_liters // tin_of_paint_in_liters + 1) # количество банок
not_used_paints_in_liters = number_of_tins * tin_of_paint_in_liters - number_of_liters # неиспользуемых литров краски
print('Площадь окрашивания равна: ')
print(round(surface_area, 2))
print('Количество литров равно: ')
print(round(number_of_liters, 2))
print('Количество банок равно: ')
print(number_of_tins)
print('Количество неиспользуемых литров краски равно: ')
print(round(not_used_paints_in_liters, 2))
| Paints_Calculator.ipynb |
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ### 練習問題2.5
;; $a$ と $b$ のペアを積 $2^a \cdot 3^b$
;; の整数で表現することによって、
;; ⾮負整数のペアを数値と数値演算だけを使って表現できるということを⽰せ。
;; それに対応する cons, car, cdr ⼿続きを定義せよ。
(define (expr x n)
(define (iter m result)
(if (= n m) result
(iter (+ m 1) (* result x))
)
)
(iter 0 1)
)
;(display (expr 3 4))
;(newline)
(define (div x base)
(define (iter y result)
(if (not (= (% y base) 0)) result
(iter (/ y base) (+ result 1))
)
)
(iter x 0)
)
;(display (div 972 2))
;(newline)
(define (cons a b)
(* (expr 2 a) (expr 3 b))
)
(define (car z)
(div z 2))
(define (cdr z)
(div z 3))
(define x (cons 10 4))
(display (car x))
(newline)
(display (cdr x))
(newline)
| exercises/2.05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# 
# # Reinforcement Learning in Azure Machine Learning - Training a Minecraft agent using custom environments
#
# This tutorial will show how to set up a more complex reinforcement
# learning (RL) training scenario. It demonstrates how to train an agent to
# navigate through a lava maze in the Minecraft game using Azure Machine
# Learning.
#
# **Please note:** This notebook trains an agent on a randomly generated
# Minecraft level. As a result, on rare occasions, a training run may fail
# to produce a model that can solve the maze. If this happens, you can
# re-run the training step as indicated below.
#
# **Please note:** This notebook uses 1 NC6 type node and 8 D2 type nodes
# for up to 5 hours of training, which corresponds to approximately $9.06 (USD)
# as of May 2020.
#
# Minecraft is currently one of the most popular video
# games and as such has been a study object for RL. [Project
# Malmo](https://www.microsoft.com/en-us/research/project/project-malmo/) is
# a platform for artificial intelligence experimentation and research built on
# top of Minecraft. We will use Minecraft [gym](https://gym.openai.com) environments from Project
# Malmo's 2019 MineRL competition, which are part of the
# [MineRL](http://minerl.io/docs/index.html) Python package.
#
# Minecraft environments require a display to run, so we will demonstrate
# how to set up a virtual display within the docker container used for training.
# Learning will be based on the agent's visual observations. To
# generate the necessary amount of sample data, we will run several
# instances of the Minecraft game in parallel. Below, you can see a video of
# a trained agent navigating a lava maze. Starting from the green position,
# it moves to the blue position by moving forward, turning left or turning right:
#
# <table style="width:50%">
# <tr>
# <th style="text-align: center;">
# <img src="./images/lava_maze_minecraft.gif" alt="Minecraft lava maze" align="middle" margin-left="auto" margin-right="auto"/>
# </th>
# </tr>
# <tr style="text-align: center;">
# <th>Fig 1. Video of a trained Minecraft agent navigating a lava maze.</th>
# </tr>
# </table>
#
# The tutorial will cover the following steps:
# - Initializing Azure Machine Learning resources for training
# - Training the RL agent with Azure Machine Learning service
# - Monitoring training progress
# - Reviewing training results
#
#
# ## Prerequisites
#
# The user should have completed the Azure Machine Learning introductory tutorial.
# You will need to make sure that you have a valid subscription id, a resource group and a
# workspace. For detailed instructions see [Tutorial: Get started creating
# your first ML experiment.](https://docs.microsoft.com/en-us/azure/machine-learning/tutorial-1st-experiment-sdk-setup)
#
# In addition, please follow the instructions in the [Reinforcement Learning in
# Azure Machine Learning - Setting Up Development Environment](../setup/devenv_setup.ipynb)
# notebook to correctly set up a Virtual Network which is required for completing
# this tutorial.
#
# While this is a standalone notebook, we highly recommend going over the
# introductory notebooks for RL first.
# - Getting started:
# - [RL using a compute instance with Azure Machine Learning service](../cartpole-on-compute-instance/cartpole_ci.ipynb)
# - [Using Azure Machine Learning compute](../cartpole-on-single-compute/cartpole_sc.ipynb)
# - [Scaling RL training runs with Azure Machine Learning service](../atari-on-distributed-compute/pong_rllib.ipynb)
#
#
# ## Initialize resources
#
# All required Azure Machine Learning service resources for this tutorial can be set up from Jupyter.
# This includes:
# - Connecting to your existing Azure Machine Learning workspace.
# - Creating an experiment to track runs.
# - Creating remote compute targets for [Ray](https://docs.ray.io/en/latest/index.html).
#
# ### Azure Machine Learning SDK
#
# Display the Azure Machine Learning SDK version.
import azureml.core
print("Azure Machine Learning SDK Version: ", azureml.core.VERSION)
# ### Connect to workspace
#
# Get a reference to an existing Azure Machine Learning workspace.
# +
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.location, ws.resource_group, sep=' | ')
# -
# ### Create an experiment
#
# Create an experiment to track the runs in your workspace. A
# workspace can have multiple experiments and each experiment
# can be used to track multiple runs (see [documentation](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.experiment.experiment?view=azure-ml-py)
# for details).
# + nbpresent={"id": "bc70f780-c240-4779-96f3-bc5ef9a37d59"}
from azureml.core import Experiment
exp = Experiment(workspace=ws, name='minecraft-maze')
# -
# ### Create or attach an existing compute resource
#
# A compute target is a designated compute resource where you
# run your training script. For more information, see [What
# are compute targets in Azure Machine Learning service?](https://docs.microsoft.com/en-us/azure/machine-learning/concept-compute-target).
#
# #### GPU target for Ray head
#
# In the experiment setup for this tutorial, the Ray head node
# will run on a GPU-enabled node. A maximum cluster size
# of 1 node is therefore sufficient. If you wish to run
# multiple experiments in parallel using the same GPU
# cluster, you may elect to increase this number. The cluster
# will automatically scale down to 0 nodes when no training jobs
# are scheduled (see `min_nodes`).
#
# The code below creates a compute cluster of GPU-enabled NC6
# nodes. If the cluster with the specified name is already in
# your workspace the code will skip the creation process.
#
# Note that we must specify a Virtual Network during compute
# creation to allow communication between the cluster running
# the Ray head node and the additional Ray compute nodes. For
# details on how to setup the Virtual Network, please follow the
# instructions in the "Prerequisites" section above.
#
# **Note: Creation of a compute resource can take several minutes**
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# please enter the name of your Virtual Network (see Prerequisites -> Workspace setup)
vnet_name = 'your_vnet'
# name of the Virtual Network subnet ('default' the default name)
subnet_name = 'default'
gpu_cluster_name = 'gpu-cluster-nc6'
try:
gpu_cluster = ComputeTarget(workspace=ws, name=gpu_cluster_name)
print('Found existing compute target')
except ComputeTargetException:
print('Creating a new compute target...')
compute_config = AmlCompute.provisioning_configuration(
vm_size='Standard_NC6',
min_nodes=0,
max_nodes=1,
vnet_resourcegroup_name=ws.resource_group,
vnet_name=vnet_name,
subnet_name=subnet_name)
gpu_cluster = ComputeTarget.create(ws, gpu_cluster_name, compute_config)
gpu_cluster.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
print('Cluster created.')
# -
# #### CPU target for additional Ray nodes
#
# The code below creates a compute cluster of D2 nodes. If the cluster with the specified name is already in your workspace the code will skip the creation process.
#
# This cluster will be used to start additional Ray nodes
# increasing the clusters CPU resources.
#
# **Note: Creation of a compute resource can take several minutes**
# +
cpu_cluster_name = 'cpu-cluster-d2'
try:
cpu_cluster = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing compute target')
except ComputeTargetException:
print('Creating a new compute target...')
compute_config = AmlCompute.provisioning_configuration(
vm_size='STANDARD_D2',
min_nodes=0,
max_nodes=10,
vnet_resourcegroup_name=ws.resource_group,
vnet_name=vnet_name,
subnet_name=subnet_name)
cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
cpu_cluster.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
print('Cluster created.')
# -
# ## Training the agent
#
# ### Training environments
#
# This tutorial uses custom docker images (CPU and GPU respectively)
# with the necessary software installed. The
# [Environment](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-use-environments)
# class stores the configuration for the training environment. The docker
# image is set via `env.docker.base_image` which can point to any
# publicly available docker image. `user_managed_dependencies`
# is set so that the preinstalled Python packages in the image are preserved.
#
# Note that since Minecraft requires a display to start, we set the `interpreter_path`
# such that the Python process is started via **xvfb-run**.
# +
import os
from azureml.core import Environment
max_train_time = os.environ.get("AML_MAX_TRAIN_TIME_SECONDS", 5 * 60 * 60)
def create_env(env_type):
env = Environment(name='minecraft-{env_type}'.format(env_type=env_type))
env.docker.enabled = True
env.docker.base_image = 'akdmsft/minecraft-{env_type}'.format(env_type=env_type)
env.python.interpreter_path = "xvfb-run -s '-screen 0 640x480x16 -ac +extension GLX +render' python"
env.environment_variables["AML_MAX_TRAIN_TIME_SECONDS"] = str(max_train_time)
env.python.user_managed_dependencies = True
return env
cpu_minecraft_env = create_env('cpu')
gpu_minecraft_env = create_env('gpu')
# -
# ### Training script
#
# As described above, we use the MineRL Python package to launch
# Minecraft game instances. MineRL provides several OpenAI gym
# environments for different scenarios, such as chopping wood.
# Besides predefined environments, MineRL lets its users create
# custom Minecraft environments through
# [minerl.env](http://minerl.io/docs/api/env.html). In the helper
# file **minecraft_environment.py** provided with this tutorial, we use the
# latter option to customize a Minecraft level with a lava maze
# that the agent has to navigate. The agent receives a negative
# reward of -1 for falling into the lava, a negative reward of
# -0.02 for sending a command (i.e. navigating through the maze
# with fewer actions yields a higher total reward) and a positive reward
# of 1 for reaching the goal. To encourage the agent to explore
# the maze, it also receives a positive reward of 0.1 for visiting
# a tile for the first time.
#
# The agent learns purely from visual observations and the image
# is scaled to an 84x84 format, stacking four frames. For the
# purposes of this example, we use a small action space of size
# three: move forward, turn 90 degrees to the left, and turn 90
# degrees to the right.
#
# The training script itself registers the function to create training
# environments with the `tune.register_env` function and connects to
# the Ray cluster Azure Machine Learning service started on the GPU
# and CPU nodes. Lastly, it starts a RL training run with `tune.run()`.
#
# We recommend setting the `local_dir` parameter to `./logs` as this
# directory will automatically become available as part of the training
# run's files in the Azure Portal. The Tensorboard integration
# (see "View the Tensorboard" section below) also depends on the files'
# availability. For a list of common parameter options, please refer
# to the [Ray documentation](https://docs.ray.io/en/latest/rllib-training.html#common-parameters).
#
#
# ```python
# # Taken from minecraft_environment.py and minecraft_train.py
#
# # Define a function to create a MineRL environment
# def create_env(config):
# mission = config['mission']
# port = 1000 * config.worker_index + config.vector_index
# print('*********************************************')
# print(f'* Worker {config.worker_index} creating from mission: {mission}, port {port}')
# print('*********************************************')
#
# if config.worker_index == 0:
# # The first environment is only used for checking the action and observation space.
# # By using a dummy environment, there's no need to spin up a Minecraft instance behind it
# # saving some CPU resources on the head node.
# return DummyEnv()
#
# env = EnvWrapper(mission, port)
# env = TrackingEnv(env)
# env = FrameStack(env, 2)
#
# return env
#
#
# def stop(trial_id, result):
# return result["episode_reward_mean"] >= 1 \
# or result["time_total_s"] > 5 * 60 * 60
#
#
# if __name__ == '__main__':
# tune.register_env("Minecraft", create_env)
#
# ray.init(address='auto')
#
# tune.run(
# run_or_experiment="IMPALA",
# config={
# "env": "Minecraft",
# "env_config": {
# "mission": "minecraft_missions/lava_maze-v0.xml"
# },
# "num_workers": 10,
# "num_cpus_per_worker": 2,
# "rollout_fragment_length": 50,
# "train_batch_size": 1024,
# "replay_buffer_num_slots": 4000,
# "replay_proportion": 10,
# "learner_queue_timeout": 900,
# "num_sgd_iter": 2,
# "num_data_loader_buffers": 2,
# "exploration_config": {
# "type": "EpsilonGreedy",
# "initial_epsilon": 1.0,
# "final_epsilon": 0.02,
# "epsilon_timesteps": 500000
# },
# "callbacks": {"on_train_result": callbacks.on_train_result},
# },
# stop=stop,
# checkpoint_at_end=True,
# local_dir='./logs'
# )
# ```
#
# ### Submitting a training run
#
# Below, you create the training run using a `ReinforcementLearningEstimator`
# object, which contains all the configuration parameters for this experiment:
# - `source_directory`: Contains the training script and helper files to be
# copied onto the node running the Ray head.
# - `entry_script`: The training script, described in more detail above..
# - `compute_target`: The compute target for the Ray head and training
# script execution.
# - `environment`: The Azure machine learning environment definition for
# the node running the Ray head.
# - `worker_configuration`: The configuration object for the additional
# Ray nodes to be attached to the Ray cluster:
# - `compute_target`: The compute target for the additional Ray nodes.
# - `node_count`: The number of nodes to attach to the Ray cluster.
# - `environment`: The environment definition for the additional Ray nodes.
# - `max_run_duration_seconds`: The time after which to abort the run if it
# is still running.
# - `shm_size`: The size of docker container's shared memory block.
#
# For more details, please take a look at the [online documentation](https://docs.microsoft.com/en-us/python/api/azureml-contrib-reinforcementlearning/?view=azure-ml-py)
# for Azure Machine Learning service's reinforcement learning offering.
#
# We configure 8 extra D2 (worker) nodes for the Ray cluster, giving us a total of
# 22 CPUs and 1 GPU. The GPU and one CPU are used by the IMPALA learner,
# and each MineRL environment receives 2 CPUs allowing us to spawn a total
# of 10 rollout workers (see `num_workers` parameter in the training script).
#
#
# Lastly, the `RunDetails` widget displays information about the submitted
# RL experiment, including a link to the Azure portal with more details.
# +
from azureml.contrib.train.rl import ReinforcementLearningEstimator, WorkerConfiguration
from azureml.widgets import RunDetails
worker_config = WorkerConfiguration(
compute_target=cpu_cluster,
node_count=8,
environment=cpu_minecraft_env)
rl_est = ReinforcementLearningEstimator(
source_directory='files',
entry_script='minecraft_train.py',
compute_target=gpu_cluster,
environment=gpu_minecraft_env,
worker_configuration=worker_config,
max_run_duration_seconds=6 * 60 * 60,
shm_size=1024 * 1024 * 1024 * 30)
train_run = exp.submit(rl_est)
RunDetails(train_run).show()
# +
# If you wish to cancel the run before it completes, uncomment and execute:
#train_run.cancel()
# -
# ## Monitoring training progress
#
# ### View the Tensorboard
#
# The Tensorboard can be displayed via the Azure Machine Learning service's
# [Tensorboard API](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-monitor-tensorboard).
# When running locally, please make sure to follow the instructions in the
# link and install required packages. Running this cell will output a URL
# for the Tensorboard.
#
# Note that the training script sets the log directory when starting RLlib
# via the `local_dir` parameter. `./logs` will automatically appear in
# the downloadable files for a run. Since this script is executed on the
# Ray head node run, we need to get a reference to it as shown below.
#
# The Tensorboard API will continuously stream logs from the run.
#
# **Note: It may take a couple of minutes after the run is in "Running" state
# before Tensorboard files are available and the board will refresh automatically**
# +
import time
from azureml.tensorboard import Tensorboard
head_run = None
timeout = 60
while timeout > 0 and head_run is None:
timeout -= 1
try:
head_run = next(r for r in train_run.get_children() if r.id.endswith('head'))
except StopIteration:
time.sleep(1)
tb = Tensorboard([head_run])
tb.start()
# -
# ## Review results
#
# Please ensure that the training run has completed before continuing with this section.
# +
train_run.wait_for_completion()
print('Training run completed.')
# -
# **Please note:** If the final "episode_reward_mean" metric from the training run is negative,
# the produced model does not solve the problem of navigating the maze well. You can view
# the metric on the Tensorboard or in "Metrics" section of the head run in the Azure Machine Learning
# portal. We recommend training a new model by rerunning the notebook starting from "Submitting a training run".
#
#
# ### Export final model
#
# The key result from the training run is the final checkpoint
# containing the state of the IMPALA trainer (model) upon meeting the
# stopping criteria specified in `minecraft_train.py`.
#
# Azure Machine Learning service offers the [Model.register()](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.model.model?view=azure-ml-py)
# API which allows you to persist the model files from the
# training run. We identify the directory containing the
# final model written during the training run and register
# it with Azure Machine Learning service. We use a Dataset
# object to filter out the correct files.
# +
import re
import tempfile
from azureml.core import Dataset
path_prefix = os.path.join(tempfile.gettempdir(), 'tmp_training_artifacts')
run_artifacts_path = os.path.join('azureml', head_run.id)
datastore = ws.get_default_datastore()
run_artifacts_ds = Dataset.File.from_files(datastore.path(os.path.join(run_artifacts_path, '**')))
cp_pattern = re.compile('.*checkpoint-\\d+$')
checkpoint_files = [file for file in run_artifacts_ds.to_path() if cp_pattern.match(file)]
# There should only be one checkpoint with our training settings...
final_checkpoint = os.path.dirname(os.path.join(run_artifacts_path, os.path.normpath(checkpoint_files[-1][1:])))
datastore.download(target_path=path_prefix, prefix=final_checkpoint.replace('\\', '/'), show_progress=True)
print('Download complete.')
# +
from azureml.core.model import Model
model_name = 'final_model_minecraft_maze'
model = Model.register(
workspace=ws,
model_path=os.path.join(path_prefix, final_checkpoint),
model_name=model_name,
description='Model of an agent trained to navigate a lava maze in Minecraft.')
# -
# Models can be used through a varity of APIs. Please see the
# [documentation](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-deploy-and-where)
# for more details.
#
# ### Test agent performance in a rollout
#
# To observe the trained agent's behavior, it is a common practice to
# view its behavior in a rollout. The previous reinforcement learning
# tutorials explain rollouts in more detail.
#
# The provided `minecraft_rollout.py` script loads the final checkpoint
# of the trained agent from the model registered with Azure Machine Learning
# service. It then starts a rollout on 4 different lava maze layouts, that
# are all larger and thus more difficult than the maze the agent was trained
# on. The script further records videos by replaying the agent's decisions
# in [Malmo](https://github.com/microsoft/malmo). Malmo supports multiple
# agents in the same environment, thus allowing us to capture videos that
# depict the agent from another agent's perspective. The provided
# `malmo_video_recorder.py` file and the Malmo Github repository have more
# details on the video recording setup.
#
# You can view the rewards for each rollout episode in the logs for the 'head'
# run submitted below. In some episodes, the agent may fail to reach the goal
# due to the higher level of difficulty - in practice, we could continue
# training the agent on harder tasks starting with the final checkpoint.
# +
script_params = {
'--model_name': model_name
}
rollout_est = ReinforcementLearningEstimator(
source_directory='files',
entry_script='minecraft_rollout.py',
script_params=script_params,
compute_target=gpu_cluster,
environment=gpu_minecraft_env,
shm_size=1024 * 1024 * 1024 * 30)
rollout_run = exp.submit(rollout_est)
RunDetails(rollout_run).show()
# -
# ### View videos captured during rollout
#
# To inspect the agent's training progress you can view the videos captured
# during the rollout episodes. First, ensure that the training run has
# completed.
# +
rollout_run.wait_for_completion()
head_run_rollout = next(r for r in rollout_run.get_children() if r.id.endswith('head'))
print('Rollout completed.')
# -
# Next, you need to download the video files from the training run. We use a
# Dataset to filter out the video files which are in tgz archives.
# +
rollout_run_artifacts_path = os.path.join('azureml', head_run_rollout.id)
datastore = ws.get_default_datastore()
rollout_run_artifacts_ds = Dataset.File.from_files(datastore.path(os.path.join(rollout_run_artifacts_path, '**')))
video_archives = [file for file in rollout_run_artifacts_ds.to_path() if file.endswith('.tgz')]
video_archives = [os.path.join(rollout_run_artifacts_path, os.path.normpath(file[1:])) for file in video_archives]
datastore.download(
target_path=path_prefix,
prefix=os.path.dirname(video_archives[0]).replace('\\', '/'),
show_progress=True)
print('Download complete.')
# -
# Next, unzip the video files and rename them by the Minecraft mission seed used
# (see `minecraft_rollout.py` for more details on how the seed is used).
# +
import tarfile
import shutil
training_artifacts_dir = './training_artifacts'
video_dir = os.path.join(training_artifacts_dir, 'videos')
video_files = []
for tar_file_path in video_archives:
seed = tar_file_path[tar_file_path.index('rollout_') + len('rollout_'): tar_file_path.index('.tgz')]
tar = tarfile.open(os.path.join(path_prefix, tar_file_path).replace('\\', '/'), 'r')
tar_info = next(t_info for t_info in tar.getmembers() if t_info.name.endswith('mp4'))
tar.extract(tar_info, video_dir)
tar.close()
unzipped_folder = os.path.join(video_dir, next(f_ for f_ in os.listdir(video_dir) if not f_.endswith('mp4')))
video_file = os.path.join(unzipped_folder,'video.mp4')
final_video_path = os.path.join(video_dir, '{seed}.mp4'.format(seed=seed))
shutil.move(video_file, final_video_path)
video_files.append(final_video_path)
shutil.rmtree(unzipped_folder)
# Clean up any downloaded 'tmp' files
shutil.rmtree(path_prefix)
print('Local video files:\n', video_files)
# -
# Finally, run the cell below to display the videos in-line. In some cases,
# the agent may struggle to find the goal since the maze size was increased
# compared to training.
# +
from IPython.core.display import display, HTML
index = 0
while index < len(video_files) - 1:
display(
HTML('\
<video controls alt="cannot display video" autoplay loop width=49%> \
<source src="{f1}" type="video/mp4"> \
</video> \
<video controls alt="cannot display video" autoplay loop width=49%> \
<source src="{f2}" type="video/mp4"> \
</video>'.format(f1=video_files[index], f2=video_files[index + 1]))
)
index += 2
if index < len(video_files):
display(
HTML('\
<video controls alt="cannot display video" autoplay loop width=49%> \
<source src="{f1}" type="video/mp4"> \
</video>'.format(f1=video_files[index]))
)
# -
# ## Cleaning up
#
# Below, you can find code snippets for your convenience to clean up any resources created as part of this tutorial you don't wish to retain.
# +
# to stop the Tensorboard, uncomment and run
#tb.stop()
# +
# to delete the gpu compute target, uncomment and run
#gpu_cluster.delete()
# +
# to delete the cpu compute target, uncomment and run
#cpu_cluster.delete()
# +
# to delete the registered model, uncomment and run
#model.delete()
# +
# to delete the local video files, uncomment and run
#shutil.rmtree(training_artifacts_dir)
# -
# ## Next steps
#
# This is currently the last introductory tutorial for Azure Machine Learning
# service's Reinforcement
# Learning offering. We would love to hear your feedback to build the features
# you need!
#
#
| how-to-use-azureml/reinforcement-learning/minecraft-on-distributed-compute/minecraft.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Multi-State Model first example
# ## In this notebook
# This notebook provides a simple setting which illustrates basic usage of the model.
# ## Typical settings
# In a typical setting of modelling patient illness trajectories, there are multiple sources of complexity:
#
# 1. There could be many states (mild, severe, recovered, released from hospital, death etc.)
# 2. The probability of each transition and the duration of the stay in each state depend on patient covariates.
# 3. Patient covariates can change over time, possibly in a manner which depends on the states visited.
#
# In order to introduce the multi-state-model we shall use a much simpler setting where our data arrives from a simple 3 state model and covariates do not change over time or affect the probabilities of transitions between states.
# ## A Simple Multi-State Setting
# Patients start at state 1, state 3 shall be a terminal state and states 1,2 shall be identical in the sense that from both:
#
# 1. With probability 1/2 you transition to state 3 within 1 day.
# 2. With probability 1/2 you transition to state 2 or 1 (depending on the present state), within $t∼exp(λ)$
# +
from pymsm.plotting import state_diagram
state_diagram(
"""
s1 : 1
s2: 2
s3: 3
s1 --> s2: P=0.5, t~exp(lambda)
s1 --> s3: P=0.5, t=1
s2 --> s3: P=0.5, t=1
"""
)
# -
# A simple Multi-State Model
# For this setting, one can show that the expected time until reaching a terminal state is $1+\frac{1}{λ}$ (see proof at the end of this notebook.)
# ## The Dataset Structure
# Let’s load the dataset, which was constructed based on the graph above
from pymsm.examples.first_example_utils import create_toy_setting_dataset
dataset = create_toy_setting_dataset(lambda_param=2)
print('dataset type: {}'.format(type(dataset)))
print('elemnets type: {}'.format(type(dataset[0])))
# The dataset is a list of elements from class PathObject. Each PathObject in the list corresponds to a single sample’s (i.e “patient’s”) observed path. Let’s look at one such object in detail:
# +
first_path = dataset[0]
print(type(first_path))
print('\n------covariates------')
print(first_path.covariates)
print('\n-------states---------')
print(first_path.states)
print('\n--time at each state--')
print(first_path.time_at_each_state)
print('\n------sample id-------')
print(first_path.sample_id)
# -
# We see the following attributes:
#
# 1. *covariates* : These are the sample’s covariates. In this case they were randomally generated and do not affect the state transitions, but for a patient this could be a numerical vector with entries such as:
# * “age in years”
# * “is male”
# * “number of days that have passed since hospitalization”
# * etc..
# 2. *states* : These are the observed states the sample visited, encoded as positive integers. Here we can see the back and forth between states 1 and 2, ending with the only terminal state (state 3).
# 3. *time_at_each_state* : These are the observed times spent at each state.
# 4. *sample_id* : (optional) a unique identifier of the patient.
#
# Note: if the last state is a terminal state, then the vector of times should be shorter than the vector of states by 1. Conversely, if the last state is not a terminal state, then the length of the vector of times should be the same as that of the states. In such a case, the sample is inferred to be right censored.
# ## Updating Covariates Over Time
# In order to update the patient covariates over time, we need to define a state-transition function.
# In this simple case, the covariates do not change and the function is trivial
def default_update_covariates_function(covariates_entering_origin_state, origin_state=None, target_state=None,
time_at_origin=None, abs_time_entry_to_target_state=None):
return covariates_entering_origin_state
# You can define any function, as long as it recieves the following parameter types (in this order):
# 1. pandas Series (sample covariates when entering the origin state)
# 2. int (origin state number)
# 3. int (target state number)
# 4. float (time spent at origin state)
# 5. float (absolute time of entry to target state)
#
# If some of the parameters are not used in the function, use a default value of None, as in the example above.
# ## Defining terminal states
terminal_states = [3]
# ## Fitting the model
# Import and init the Model
from pymsm.multi_state_competing_risks_model import MultiStateModel
multi_state_model = MultiStateModel(dataset, terminal_states, default_update_covariates_function,
['covariate_1', 'covariate_2'])
# Fit the Model
multi_state_model.fit()
# ## Making predictions
#
# Predictions are done via monte carlo simulation. Initial patient covariates, along with the patient’s current state are supplied. The next states are sequentially sampled via the model parameters. The process concludes when the patient arrives at a terminal state or the number of transitions exceeds the specified maximum.
# +
import numpy as np
all_mcs = multi_state_model.run_monte_carlo_simulation(
# the current covariates of the patient.
# especially important to use updated covariates in case of
# time varying covariates along with a prediction from a point in time
# during hospitalization
sample_covariates = np.array([0.2,-0.3]),
# in this setting samples start at state 1, but
# in general this can be any non-terminal state which
# then serves as the simulation starting point
origin_state = 1,
# in this setting we start predictions from time 0, but
# predictions can be made from any point in time during the
# patient's trajectory
current_time = 0,
# If there is an observed upper limit on the number of transitions, we recommend
# setting this value to that limit in order to prevent generation of outlier paths
max_transitions = 100,
# the number of paths to simulate:
n_random_samples = 1000)
# -
# ## The Simulation Results Format:
#
# Each run is described by a list of states and times spent at each state (same format as the dataset the model is fit to).
# +
mc = all_mcs[0]
print(mc.states)
print(mc.time_at_each_state)
mc = all_mcs[1]
print(mc.states)
print(mc.time_at_each_state)
# -
# ## Analyzing The Results
#
# Recall we could compute the expected time for this simple setting? We will now see that the model provides an accurate estimate of this expected value of $1+\frac{1}{\lambda}$
# +
from pymsm.examples.first_example_utils import plot_total_time_until_terminal_state
plot_total_time_until_terminal_state(all_mcs, true_lambda=2)
# -
# ## Conclusions
# This notebook provides a simple example usage of the multi-state model, beginning with the structure of the dataset used to fit the model and up to a simple analysis of the model’s predictions.
#
# By following this process you can fit the model to any such dataset and make predictions
# ## Appendix 1 - Demonstrating that the expected time until reaching the terminal state is $1+\frac{1}{λ}$
# Let T be the random variable denoting the time until reaching the terminal state #3, and let $S2$ be the random variable denoting the second state visited by the sample (recall all patients start at state 1, that is: $S1=1$).
# From the law of total expectation:
# \begin{equation}
# \mathbf{E}[T] = \mathbf{E}[\mathbf{E}[T|S_2]] = \mathbf{P}(S_2 = 3)\cdot\mathbf{E}[T|S_2 = 3] + \mathbf{P}(S_2 = 2)\cdot\mathbf{E}[T|S_2 = 2]
# \end{equation}
# Denote $T=T_1+T_{2^+}$ (“The total time is the sum of the time of the first transition plus the time from arrival to the second state onwards”). Then:
# \begin{equation}
# =\frac{1}{2}\cdot1 + \frac{1}{2}\cdot\mathbf{E}[T_1 + T_{2^+}|S_2 = 2] = \frac{1}{2}+\frac{1}{2}\cdot(\mathbf{E}[T_1|S_2 = 2] + \mathbf{E}[T_{2^+}]|S_2 = 2) \\= \frac{1}{2}\cdot1 + \frac{1}{2}\cdot(\frac{1}{λ}+\mathbf{E}[T])
# \end{equation}
# We then have:
# \begin{equation}
# 2\cdot\mathbf{E}[T] = 1 + (\frac{1}{λ} + \mathbf{E}[T])
# \end{equation}
# and:
# \begin{equation}
# {E}[T] = 1 + \frac{1}{λ}
# \end{equation}
| src/pymsm/archive/first_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 180816 오후수업
#
# ## Kaggle Titanic
#
# ### Kaggle titanic 데이터로 시각화하기
#
# `conda install -c conda-forge missingno`
#
# ```python
# import missingno as msno
# msno.matrix(train)
# ```
# #### 1. pivot_table과 groupby 활용해 보기
#
# - [pandas.DataFrame.groupby — pandas 0.23.4 documentation] (https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.groupby.html)
#
# ```python
# data.groupby(['col1', 'col2']).mean()
# data.groupby('col1')[['col2']].mean()
#
# data.pivot_table(index=['col1'], values=['col2'], aggfunc=np.mean)
# ```
#
# #### 2. 타이타닉 데이터 시각화
#
# https://www.rstudio.com/wp-content/uploads/2015/03/ggplot2-cheatsheet.pdf
#
# ```python
# train['Survived'] = train['Survived'].astype('category')
#
# (ggplot(train)
# + aes(x='Sex', y='Survived')
# + geom_col() + ggtitle('성별 생존률')
# + theme(text=element_text(family='NanumBarunGothic'))
# )
#
# (ggplot(train)
# + aes(x='Age', y='Fare', fill='Pclass')
# + geom_point() + ggtitle('생존 여부별 성별 요금 등급 분포')
# + facet_wrap('~Survived')
# + theme(text=element_text(family='NanumBarunGothic'))
# )
#
# (ggplot(train)
# + aes(x='Age')
# + geom_histogram(binwidth=10)
# + ggtitle('연령대 분포')
# + theme(text=element_text(family='NanumBarunGothic'))
# )
# ```
# #### 3. 결측치 처리
#
# 결측치 처리 후 시각화 해보기
#
# `train['Age'] = train['Age'].dropna()`
# ## Kaggle Titanic 데이터 분석 실습
import pandas as pd
import numpy as np
from plotnine import *
train = pd.read_csv("/Users/jinyoungpark/Desktop/Projects/Titanic/train.csv")
test = pd.read_csv("/Users/jinyoungpark/Desktop/Projects/Titanic/test.csv")
train.head()
train.groupby('Sex').mean()
(ggplot(train)
+ aes(x='Sex', y='Survived')
+ geom_col()
+ ggtitle("성별생존률")
+ theme(text=element_text(family="NanumBarunGothic"))
)
# +
train=train.dropna()
#Embarked에 결측치가 2개 있어서 없애줌. (아니면 에러남)
(
ggplot(train)
+ aes (x='Sex', y='Survived', fill='Embarked')
+ geom_col()
)
# +
# 각 조가 생각한 조건을 적용시키고 Kaggle에 올려본다
# -
test['Survived'] = ((test['Sex'] == 'female')|(test['Age']<=9)|((test["Sex"]=='male')&(test['Pclass']!=3)))
submission = test[['PassengerId','Survived']].copy()
submission['Survived']=submission['Survived'].astype(int)
submission.to_csv('/Users/jinyoungpark/Desktop/Projects/Titanic/submission_fourth.csv', index=False)
# +
# 파일이 제대로 만들어졌는지 확인
# index가 있는 채로 업로드하면 Kaggle에서 패널티를 받으므로 index=False로 지정
# -
pd.read_csv("/Users/jinyoungpark/Desktop/Projects/Titanic/submission_fourth.csv")
| Tutorial_20180816_Kaggle_titanic_data_viasualization.md.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
from numba import jit
import numpy as np
@jit
def sum2d(arr):
M, N = arr.shape
result = 0.0
for i in range(M):
for j in range(N):
result += arr[i, j]
return result
a = np.arange(9).reshape(3, 3)
a
print(sum2d(a))
# +
from timeit import default_timer as timer
from matplotlib.pyplot import imshow, jet, show, ion
import numpy as np
from numba import jit
# +
# @jit
def mandel(x, y, max_iters):
'''
Given the real and imaginary parts of a complex number,
determine if it is a candidate for membership in the Mandelbrot
set given a fixed number of iterations.
'''
i = 0
c = complex(x, y)
z = 0.0j
for i in range(max_iters):
z = z*z + c
if (z.real * z.real + z.imag * z.imag) >= 4:
return i
return 25
# @jit
def create_fractal(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
for x in range(width):
real = min_x + x * pixel_size_x
for y in range(height):
imag = min_y + y * pixel_size_y
color = mandel(real, imag, iters)
image[y, x] = color
return image
# +
image = np.zeros((500 * 2, 750 * 2), dtype=np.uint8)
s = timer()
create_fractal(-2.0, 1.0, -1.0, 1.0, image, 20)
e = timer()
print(e - s)
imshow(image)
# jet()
# ion()
show()
# -
| numba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"} language="html"
# <style>
# .cell.selected~.unselected { display: none; }
# .cell.code_cell.unselected .input { display: none; }
# </style>
# -
# ## Bar chart
#
# + pycharm={"name": "#%%\n"}
# %load_ext autoreload
# %autoreload
# %run ./prepare_data.ipynb
# %run ./analysis.ipynb
import matplotlib.pyplot as plt
# + pycharm={"name": "#%%\n"}
votes_links.plot.bar(y='votes'); # plot histogram
# + pycharm={"name": "#%%\n"}
total_votes.plot.bar(y='votes'); # plot histogram
#plt.savefig('data/total_votes.pdf', bbox_inches='tight')
# + pycharm={"name": "#%%\n"}
if len(df_raw) < 30:
guess_stats.plot.bar(x='Current URL'); # plot histogram
# + pycharm={"name": "#%%\n"}
# bar chart per URL, the correct answer is colored in green, the rest of the guesses are blue
fig, axes = plt.subplots(len(urls),1,figsize=(20, 60))
plt.subplots_adjust(wspace=0.3, hspace=0.3, top=1)
for i,ax in enumerate(axes):
url = urls[i]
temp_df = df_guess.loc[df_guess['Current URL'] == url]
print(temp_df)
labels = temp_df['Guess']
cor_ans = df2_raw[df2_raw['URL'] == url]['Player'].values[0]
print(cor_ans)
ind=labels.isin([cor_ans])
color=list(map(lambda x: 'C2' if x else 'C0', ind))
#color=list(map(lambda x: 'C0' if x else 'C0', ind))
width = len(ind) * [0.2]
temp_df.loc[:,['Guess','votes']].plot.bar(ax=ax,subplots=True, x='Guess',y='votes',color=color, fontsize=12);
ax.get_legend().remove()
ax.set_title(url,fontdict={'fontsize':12})
ax.set(xlabel=None)
plt.savefig('../data/charts/bar-chart.pdf', bbox_inches='tight')
#plt.show()
# -
# ## Pie chart
# + pycharm={"name": "#%%\n"}
# combined pie chart, only for small dataframes
if len(df_raw) < 30:
guess_stats.plot.pie(subplots=True,y='votes',figsize=(6, 6),autopct='%1.1f%%',shadow = True);
# + pycharm={"name": "#%%\n"}
# pie chart per URL, the correct answers are "exploded"
fig, axes = plt.subplots(len(urls),1,figsize=(30, 20))
for i,ax in enumerate(axes):
url = urls[i]
temp_df = df_guess.loc[df_guess['Current URL'] == url]
labels = temp_df['Guess']
cor_ans = df2_raw[df2_raw['URL'] == url]['Player'].values[0]
ind=labels.isin([cor_ans])
#explode=list(map(int, ind))
explode=list(map(lambda x: 0.2 if x else 0, ind)) #only explode correct guess
temp_df.loc[:,['Guess','votes']].plot.pie(ax=ax,subplots=True,labels = labels, y='votes', autopct='%1.1f%%',shadow = True,explode=explode);
ax.get_legend().remove()
ax.set_title(url)
plt.show()
# + pycharm={"name": "#%%\n"}
corr_player_stats.sort_values('Guess')
fig = corr_player_stats.plot.bar(y='Guess',x='Player').get_figure()
fig.savefig('../data/charts/correct_guess_stats.png', bbox_inches='tight')
# + pycharm={"name": "#%%\n"}
corr_guess_stats.sort_values('Guess')
fig = corr_guess_stats.plot.bar(y='Guess',x='Player').get_figure()
fig.savefig('../data/charts/correct_player_stats.png', bbox_inches='tight')
# + pycharm={"name": "#%%\n"}
# total votes for player
#total_votes.sort_values
fig = total_votes.plot.bar(y='votes',x='Player').get_figure()
fig.savefig('../data/charts/guessed_player_stats.png', bbox_inches='tight')
# + pycharm={"name": "#%%\n"}
# wrongs votes for player
fig = wrong_votes.plot.bar(y='votes',x='Player').get_figure()
fig.savefig('../data/charts/wrong_guesses_stats.png', bbox_inches='tight')
| notebooks/guess-url/create_charts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''website_projects'': conda)'
# name: python388jvsc74a57bd0fbddea5140024843998ae64bf59a7579a9660d103062604797ea5984366c686c
# ---
# # Feature Engineering: `patsy` as `FormulaTransformer`
#
# In this notebook I want to describe how to create features inside [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html) using [patsy-like formulas](https://patsy.readthedocs.io/en/latest/formulas.html). I have used this approach to generate features in a previous post: [GLM in PyMC3: Out-Of-Sample Predictions](https://juanitorduz.github.io/glm_pymc3/), so I will consider the same data set here for the sake of comparison.
# ## Prepare Notebook
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import patsy
import seaborn as sns
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.compose import ColumnTransformer
from sklearn.inspection import plot_partial_dependence
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
from sklearn.model_selection import GridSearchCV, train_test_split
from sklearn.pipeline import Pipeline
from sklearn.metrics import plot_confusion_matrix, RocCurveDisplay, auc, roc_curve
from sklearn.preprocessing import StandardScaler
sns.set_style(
style='darkgrid',
rc={'axes.facecolor': '0.9', 'grid.color': '0.8'}
)
sns.set_palette(palette='deep')
sns_c = sns.color_palette(palette='deep')
plt.rcParams['figure.figsize'] = [7, 6]
plt.rcParams['figure.dpi'] = 100
# -
# ## Generate Sample Data
# We want to fit a logistic regression model where there is a multiplicative interaction between two numerical features.
# +
SEED = 42
np.random.seed(SEED)
# Number of data points.
n = 250
# Create features.
x1 = np.random.normal(loc=0.0, scale=2.0, size=n)
x2 = np.random.normal(loc=0.0, scale=2.0, size=n)
epsilon = np.random.normal(loc=0.0, scale=0.5, size=n)
# Define target variable.
intercept = -0.5
beta_x1 = 1
beta_x2 = -1
beta_interaction = 2
z = intercept + beta_x1 * x1 + beta_x2 * x2 + beta_interaction * x1 * x2
p = 1 / (1 + np.exp(-z))
y = np.random.binomial(n=1, p=p, size=n)
df = pd.DataFrame(dict(x1=x1, x2=x2, y=y))
df.head()
# -
# This is data set is relatively well balanced:
df['y'].value_counts() / df['y'].shape[0]
# Let us now split the data into the given features and the target variable
# +
numeric_features = ['x1', 'x2']
x = df[numeric_features]
y = df['y']
# -
# Next, we do a random train-test split.
x_train, x_test, y_train, y_test = train_test_split(
x, y, train_size=0.7, random_state=SEED
)
# ## Model Pipeline
# ### Implement `FormulaTransformer`
#
# We can follow [scikit-learn's guide](https://scikit-learn.org/stable/developers/develop.html) to implement custom transformations inside a pipeline. The following transformer creates features from a `formula` as described [here](https://patsy.readthedocs.io/en/latest/formulas.html). Please note that by default it will add an `Intercept` term (i.e. columns of ones). If we do not want this feature we can simply add a `- 1` inside the formula.
class FormulaTransformer(BaseEstimator, TransformerMixin):
def __init__(self, formula):
self.formula = formula
def fit(self, X, y=None):
return self
def transform(self, X):
X_formula = patsy.dmatrix(formula_like=self.formula, data=X)
columns = X_formula.design_info.column_names
return pd.DataFrame(X_formula, columns=columns)
# ### Define and Fit the Model Pipeline
#
# Now we define and fit our model (see [Column Transformer with Mixed Types guide](https://scikit-learn.org/stable/auto_examples/compose/plot_column_transformer_mixed_types.html)). We use a $l_2$-regularized logistic regression on a multiplicative interaction of the features. Note that we are not including the intercept in the `formula` but rather in the `estimator` itself.
# +
numeric_transformer = Pipeline(steps=[
('formula_transformer', FormulaTransformer(formula='x1 * x2 - 1')),
('scaler', StandardScaler())
])
preprocessor = ColumnTransformer(transformers=[
('numeric', numeric_transformer, numeric_features)
])
estimator = LogisticRegression(
fit_intercept=True,
penalty='l2',
class_weight='balanced'
)
pipeline = Pipeline(steps=[
('preprocessor', preprocessor),
('logistic_regression', estimator)
])
param_grid = {
'logistic_regression__C' : np.logspace(start=-2, stop=2, num=20)
}
grid_search = GridSearchCV(
estimator=pipeline,
param_grid=param_grid,
scoring='roc_auc',
cv=10
)
grid_search = grid_search.fit(X=x_train, y=y_train)
# -
# **Remark:** Note that one could have used the in-build [PolynomialFeatures](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html) transformer for this particular example. Nevertheless, this *formula* approach can give additional flexibility to explore feature transformations. For example, look how easy is to include *numpy-transformations* (which can actually be encoded using a [FunctionTransformer](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.FunctionTransformer.html#sklearn.preprocessing.FunctionTransformer) as well):
formula_transformer = FormulaTransformer(formula='x1 + x2 + np.sin(x1) + np.abs(x2) - 1')
x_train_features = formula_transformer.fit_transform(X=x_train)
x_train_features.head()
# Let us visualize the transformed features:
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(10, 6))
sns.lineplot(x='x1', y='np.sin(x1)', data=x_train_features, marker='o', color=sns_c[1], ax=ax[0])
sns.lineplot(x='x2', y='np.abs(x2)', data=x_train_features, marker='o', color=sns_c[2], ax=ax[1])
ax[0].set(title=r'$x_1 \longmapsto \sin(x_1)$')
ax[1].set(title=r'$x_2 \longmapsto |x_2|$')
fig.suptitle('Feature Transformations', y=0.93);
# ## Predictions and Evaluation Metrics
#
# Now we simply evaluate the model by looking into its performance in out-of-sample data. The results are similar to the ones in the [previous post](https://juanitorduz.github.io/glm_pymc3/). First let us get generate predictions (both probabilities and predictions) and compute the accuracy:
# +
p_test_pred = grid_search.predict_proba(X=x_test)[:, 1]
y_test_pred = grid_search.predict(X=x_test)
print(f"accuracy = {accuracy_score(y_true=y_test, y_pred=y_test_pred): 0.3f}")
# -
# Now let us plot the confusion matrix:
# +
fig, ax = plt.subplots()
plot_confusion_matrix(
estimator=grid_search,
X=x_test,
y_true=y_test,
normalize='all',
cmap='viridis_r',
ax=ax
)
ax.set(title='Confusion Matrix');
# -
# It seems the false positives and false negatives shares are balanced.
#
# Next we look into the [roc curve](https://en.wikipedia.org/wiki/Receiver_operating_characteristic):
# +
fpr, tpr, thresholds = roc_curve(
y_true=y_test, y_score=p_test_pred, pos_label=1, drop_intermediate=False
)
roc_auc = auc(fpr, tpr)
fig, ax = plt.subplots()
roc_display = RocCurveDisplay(fpr=fpr, tpr=tpr, roc_auc=roc_auc)
roc_display = roc_display.plot(ax=ax, marker="o", color=sns_c[4], markersize=4)
ax.set(title='ROC - Test Set');
# -
# ## Partial Dependency Plot
#
# Finally let us compute the [partial dependency plot](https://christophm.github.io/interpretable-ml-book/pdp.html) for both features `x1` and `x2`. First we need to make sure the features are not highly correlated:
print(f'Input features correlation: {x_test.corr().loc["x1", "x2"]: 0.3f}')
# Now let us visualize the plot:
# +
fig, ax = plt.subplots()
cmap = sns.color_palette('Spectral_r', as_cmap=True)
plot_partial_dependence(
estimator=grid_search,
X=x_test,
features=[('x1', 'x2')],
contour_kw={'cmap': cmap},
ax=ax
)
ax.set(title='Partial Dependency Plot');
# -
# Note how similar it looks as compared with the model decision boundary in the [previous post](https://juanitorduz.github.io/glm_pymc3/).
| Python/formula_transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Synthetic Model
# use gaussian model to simultate training and learning dataset. use compression and error correcting code to handle the single label classification problem.
#
# ## Data Generation
# Y -> label: L dimension one hot vector. each entry represent a class.
#
# X -> featues: $4\times L$ dimentsion. mean=Bernouli($\frac{1}{2}$), variance = var.
#
# ## Training Process
# We compress the label Y into binary representation of $\hat Y=\{0,1\}^{\hat L}$ where $\hat L = \log L$. Then train a bianry regressor on each bit of $\hat Y$.
#
# In the testing process, we view the classifier as a Binary Erasure Channel and threshold the score of each label.
#
BIT_LABEL = 8
DIM_LABEL = 2**8
DIM_FEATURE = DIM_LABEL*4
VARIANCE = 0.8
THRESHOLD = 0.25
NUM_DATA = 10000
SEED = 0
# +
# genetate synthetic data
import numpy as np
from numpy.random import binomial
from numpy.random import normal
from numpy.random import randint
np.random.seed(SEED)
X_mean = binomial(n=1, p=0.5, size=(DIM_LABEL, DIM_FEATURE))#genrate mean of each class
Y = randint(DIM_LABEL, size=(NUM_DATA))
X = np.array([normal(X_mean[y], scale=VARIANCE) for y in Y])
from util import num_to_bin
Y_bin = np.array([num_to_bin(y, BIT_LABEL) for y in Y]) # compress Y into binary format
#split training and test data set
from sklearn.model_selection import train_test_split
X_tr, X_te, Y_tr, Y_te = train_test_split(X, Y_bin, test_size=0.33)
# -
# train OvsA classifiers on raw data
from sklearn.linear_model import LogisticRegression
def train_bit(bit):
clf = LogisticRegression(solver='sag')
clf.fit(y=Y_tr[:, bit], X=X_tr)
return clf
from joblib import Parallel, delayed # Multitread
origin_clfs = Parallel(n_jobs=-1)(delayed(train_bit)( i) for i in range(Y_tr.shape[1]))
Y_prob = np.array([clf.predict_proba(X_te)[:,1] for clf in origin_clfs]).T
Y_pred = np.array([clf.predict(X_te) for clf in origin_clfs]).T
(Y_pred==Y_te).sum()/float(Y_te.shape[0]*Y_te.shape[1])
# test the accuracy of threshold data, 0.5 if it is erased
acc = []
erase = []
for clip in np.arange(0.01, 0.5, 0.01):
Y_clip = np.apply_along_axis(lambda row: \
np.array([0 if x < clip else 1 if x>1-clip else 0.5 for x in row]), \
0, Y_prob)
acc.append((Y_clip==Y_te).sum()/float((Y_clip!=0.5).sum()))
erase.append((Y_clip==0.5).sum()/float(Y_te.shape[0]*Y_te.shape[1]))
# %matplotlib inline
import matplotlib.pyplot as plt
plt.plot(np.arange(0.01, 0.5, 0.01), acc, label='acc')
plt.plot(np.arange(0.01, 0.5, 0.01), erase, label='erase')
plt.xlabel('threshold')
plt.legend()
from reedsolo import RSCodec, ReedSolomonError
rs = RSCodec()
list(rs.decode(rs.encode([0,1,1,0])))
| src/Synthetic Model1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ... ***CURRENTLY UNDER DEVELOPMENT*** ...
#
# ## Obtain wave families
# ### In this case, two swell systems, one local sea
#
# inputs required:
# * Historical wave conditions (GOW2 hindcast)
# * Wave families sectors; the split is based on wave direction
#
# in this notebook:
# * Split energy based on defined sectors
# * Remove TC associated waves to avoid double counting
#
# ### Workflow:
#
# <div>
# <img src="resources/nb01_09.png" width="300px">
# </div>
#
# +
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
# common
import os
import os.path as op
from datetime import datetime
# pip
import numpy as np
import xarray as xr
# DEV: override installed teslakit
import sys
sys.path.insert(0, op.join(os.path.abspath(''), '..', '..', '..'))
# teslakit
from teslakit.database import Database
from teslakit.waves import GetDistribution_gow, GetDistribution_ws
# wavespectra
from wavespectra.specdataset import SpecDataset
# -
#
# ## Database and Site parameters
# +
# --------------------------------------
# Teslakit database
p_data = r'/media/administrador/HD/Dropbox/Guam/teslakit/data'
#p_data=r'/Users/laurac/Dropbox/Guam/teslakit/data'
db = Database(p_data)
# set site
db.SetSite('GUAM')
# +
# --------------------------------------
# set waves families parameters
_, TCs_r1_params = db.Load_TCs_r1_hist() # TCs historical parameters inside big radius
_, TCs_r2_params = db.Load_TCs_r2_hist() # TCs historical parameters inside small radius
# wave families sectors
#fams_sectors = [(315, 45), (45, 202.5), (202.5, 315)]
fams_sectors = [(202.5, 60), (60, 202.5)]
# date limits for TCs removal from waves data, and TC time window (hours)
tc_rm_date1 = '1979-01-01'
tc_rm_date2 = '2019-12-31'
tc_time_window = 12
# -
#
# ## Calculate Waves Partitions from Waves Spectra (CSIRO + wavespectra)
# +
# aux.
def fix_dir(base_dirs):
'fix csiro direction for wavespectra (from -> to)'
new_dirs = base_dirs + 180
new_dirs[np.where(new_dirs>=360)] = new_dirs[np.where(new_dirs>=360)] - 360
return new_dirs
# +
# --------------------------------------
# load waves spectra point (CSIRO spec)
WVS_spec = db.Load_WAVES_spectra()
print(WVS_spec)
print()
# direction data fix
# WVS_spec['direction'] = fix_dir(WVS_spec['direction'])
# # rename variables
# WVS_spec = WVS_spec.rename(
# {
# 'frequency':'freq',
# 'direction':'dir',
# 'Efth':'efth',
# }
# ).set_coords({'freq','dir'})
# efth: rad to º
# WVS_spec['efth'] = WVS_spec['efth'] #* np.pi/180
# wavespectra parameters
wcut = 0.00000000001 # wcut = 0.3333
msw = 8
agef = 1.7
# bulk wavespectra
bulk_params = WVS_spec.spec.stats(['hs','tp','tm02','dpm','dspr'])
# partitions
ds_part = WVS_spec.spec.partition(
WVS_spec.Wspeed, WVS_spec.Wdir, WVS_spec.Depth,
wscut = wcut, max_swells = msw, agefac = agef,
)
WVS_parts = ds_part.spec.stats(['hs','tp','tm02','dpm','dspr'])
# Add bulk Hs, Tp, Dir variables
WVS_parts['Hs'] = bulk_params['hs']
WVS_parts['Tp'] = bulk_params['tp']
WVS_parts['Dir'] = bulk_params['dpm']
# drop station id
# WVS_parts = WVS_parts.drop('station')
# # Save partitions data
db.Save_WAVES_partitions(WVS_parts)
# -
print(WVS_parts)
#
# ## Calculate Historical Waves Families (CSIRO + wavespectra)
# +
# --------------------------------------
# Calculate wave families from waves partitions data and waves sectors
WVS_parts = db.Load_WAVES_partitions() # waves partitions data (from CSIRO spectra and wavespectra toolbox)
WVS = GetDistribution_ws(WVS_parts, fams_sectors, n_partitions=8)
# Add wavespectra bulk Hs, Tp, Dir variables
WVS['Hs'] = WVS_parts['Hs']
WVS['Tp'] = WVS_parts['Tp']
WVS['Dir'] = WVS_parts['Dir']
# ensure time dimension does not repeat values
_, index = np.unique(WVS['time'], return_index=True)
WVS = WVS.isel(time=index)
print(WVS)
# -
#
# ## Calculate Historical Waves Families (GOW)
# +
# --------------------------------------
# Calculate wave families from waves partitions data and waves sectors
#WVS_pts = db.Load_WAVES_partitions_GOW() # waves partitions data (GOW)
#WVS = GetDistribution_gow(WVS_pts, fams_sectors, n_partitions=5)
# Add GOW Hs, Tp, Dir variables
#WVS['Hs'] = WVS_pts['hs']
#WVS['Tp'] = WVS_pts['tp']
#WVS['Dir'] = WVS_pts['dir']
# -
#
# ## TCs: Waves Selection
# +
# --------------------------------------
# Locate TCs and set category alongside WAVES data
# remove TCs before 1979 and after 2020 (r1)
dds = TCs_r1_params.dmin_date.values[:]
ix = np.where((dds >= np.datetime64(tc_rm_date1)) & (dds <= np.datetime64(tc_rm_date2)))[0]
TCs_r1_params = TCs_r1_params.isel(storm=ix)
# select storms inside big circle
storms_sel = TCs_r1_params.storm.values[:]
# add TCs category alongside WAVES data
WVS['TC_category'] = (('time',), np.empty(len(WVS.time))*np.nan)
for s in storms_sel:
# waves at storm dates
ss = TCs_r1_params.sel(storm=s)
wvs_s = WVS.sel(time = slice(ss.dmin_date, ss.last_date))
# get hs_max date
t_hs_max = wvs_s.where(wvs_s.Hs == wvs_s.Hs.max(), drop=True).time.values[:][0]
# hs_max time window
w1 = t_hs_max - np.timedelta64(tc_time_window,'h')
w2 = t_hs_max + np.timedelta64(tc_time_window,'h')
# set category alongside WAVES data
ixs = np.where((WVS.time >= w1) & (WVS.time <= w2))[0]
WVS['TC_category'][ixs] = ss.category
print(WVS)
# Store historical WAVES data
db.Save_WAVES_hist(WVS)
# -
| notebooks/GUAM/GUAM/01_Offshore/09b_WAVES_CalculateFamilies_SelectTCs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Model Monitoring (beta)
# -
# ### Model Endpoints Overview Section
# 
#
# ### Model Endpoint Details Section
# 
#
# ### Model Endpoint Performance Section
# 
#
# ## Initial set up (and pre-requisites)
# 1. Make sure you have the `mlrun-api` as a Grafana data source configured in your Grafana instance. If not configured,
# otherwise add it by:
# 1. Open your grafana instance
# 2. Navigate to `Configuration -> Data Sources`
# 3. Press `Add data source` and configure the following parameters
# ```
# Name: mlrun-api
# URL: http://mlrun-api:8080/api/grafana-proxy/model-endpoints
# Access: Server (default)
#
# ## Add a custom header of:
# X-V3io-Session-Key: <YOUR ACCESS KEY>
# ```
# 4. Press `Save & Test` to make sure it works, a confirmation message should appear when this button is pressed
#
# 2. Import the available [dashboards](./dashboards) in your Grafana instance
# 3. To allow the system to utilize drift measurement, make sure you supply the train set when logging the model on the
# training step
#
# ```python
# # Log model
# context.log_model(
# "model",
# body=dumps(model),
# artifact_path=context.artifact_subpath("models"),
# extra_data=eval_metrics,
# model_file="model.pkl",
# metrics=context.results,
# training_set=<TRAIN_SET>, # <-
# label_cols=<LABEL_COLS>, # <-
# labels={"class": "sklearn.linear_model.LogisticRegression"}
# )
# ```
# 4. When serving the model, make sure that the Nuclio function is deployed with tracking enabled by applying
# `fn.set_tracking()` on the serving function
#
# ## Configuration
# The stream processing portion of the model monitoring, can be deployed under multiple configuration options. The
# available configurations can be found under `stream.Config`. Once configured it should be supplied as environment
# parameters to the Nuclio function by setting `fn.set_envs`
#
# ```python
# project: str # project name
# sample_window: int # The sampling window for the data that flows into the TSDB and the KV
# tsdb_batching_max_events: int # The max amount of event to batch before writing the batch to tsdb
# tsdb_batching_timeout_secs: int # The max amount of seconds a given batch can be gathered before being emitted
# parquet_batching_max_events: int # The max amount of event to batch before writing the batch to parquet
# parquet_batching_timeout_secs: int # The max amount of seconds, a given batch can be gathered before being written to parquet
# aggregate_count_windows: List[str] # List of window sizes for predictions count
# aggregate_count_period: str # Period of predictions count windows
# aggregate_avg_windows: List[str] # List of window sizes for average latency
# aggregate_avg_period: str # Period of average latency windows
# v3io_access_key: str # V3IO Access key, if not set will be taken from environment
# v3io_framesd: str # V3IO framesd URL, if not set will be taken from environment
# ```
# + pycharm={"name": "#%%\n"}
# Set project name
project = ""
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Deploy Model Servers
# + pycharm={"name": "#%%\n"}
import pandas as pd
from sklearn.datasets import load_iris
from mlrun import import_function, get_dataitem
from mlrun import projects
from mlrun.platforms import auto_mount
proj = projects.new_project(project)
get_dataitem("https://s3.wasabisys.com/iguazio/models/iris/model.pkl").download("model.pkl")
iris = load_iris()
train_set = pd.DataFrame(iris['data'], columns=['sepal_length_cm', 'sepal_width_cm', 'petal_length_cm', 'petal_width_cm'])
model_names = [
"sklearn_ensemble_RandomForestClassifier",
"sklearn_linear_model_LogisticRegression",
"sklearn_ensemble_AdaBoostClassifier"
]
serving_fn = import_function('hub://v2_model_server').apply(auto_mount())
for name in model_names:
proj.log_model(name, model_file="model.pkl", training_set=train_set)
serving_fn.add_model(name, model_path=f"store://models/{project}/{name}:latest")
serving_fn.metadata.project = project
serving_fn.set_tracking()
serving_fn.deploy()
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Deploy Stream Processing
# +
import os
from mlrun import import_function
from mlrun.platforms import mount_v3io
from mlrun.runtimes import RemoteRuntime
import json
fn: RemoteRuntime = import_function("hub://model_monitoring_stream")
fn.add_v3io_stream_trigger(
stream_path=f"projects/{project}/model-endpoints/stream",
name="monitoring_stream_trigger",
)
fn.set_env("MODEL_MONITORING_PARAMETERS", json.dumps({"project": project, "v3io_framesd": os.environ.get("V3IO_FRAMESD")}))
fn.metadata.project = project
fn.apply(mount_v3io())
fn.deploy()
# -
# ## Deploy Batch Processing
# + pycharm={"name": "#%%\n"}
from mlrun import import_function
from mlrun.platforms import mount_v3io
from mlrun.runtimes import KubejobRuntime
fn: KubejobRuntime = import_function("hub://model_monitoring_batch")
fn.metadata.project = project
fn.apply(mount_v3io())
fn.run(name='model-monitoring-batch', schedule="0 */1 * * *", params={"project": project})
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Simulating Requests
# + pycharm={"name": "#%%\n"}
import json
from time import sleep
from random import choice, uniform
from sklearn.datasets import load_iris
iris = load_iris()
iris_data = iris['data'].tolist()
while True:
for name in model_names:
data_point = choice(iris_data)
serving_fn.invoke(f'v2/models/{name}/infer', json.dumps({'inputs': [data_point]}))
sleep(uniform(0.1, 0.4))
sleep(uniform(0.2, 1.7))
| docs/model_monitoring/model-monitoring-deployment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyse Telegram Chat
#
# Anlayse and visualize the exported messages from `Telegram Desktop`
#
# -----
# ### Telegram Chat Export Tool
#
# For documentation how to export chats head over to the doc **with video** at [blog of Telegram](https://www.telegram.org/blog/export-and-more)
#
# -----
#
# ## Paramter
# +
# The directory containing the data export
# EDIT the date to match your export!!!
from pathlib import Path
HOME_PATH = str(Path.home()) # Python 3.5+
EXPORT_FOLDER = HOME_PATH + "/Downloads/Telegram Desktop/ChatExport_20_01_2020/"
# -
# Store the generated charts to this folder
OUTPUT_DIR = "CHARTS/" # Note the "/"" at the end!
# ## Parse Telegram Data
# +
"""
Author: <NAME> @jagoleni
Source: https://github.com/jagoleni/tele-data/blob/master/tele-data.py
"""
import os
from lxml import etree
import pandas as pd
def parse_file(html_string):
data = []
parser = etree.HTMLParser()
root = etree.HTML(html_string)
for element in root.iter():
if "id" in element.attrib:
message = {}
message["message_id"] = element.attrib["id"]
for child in element.getchildren():
if element.attrib["class"] == "message service" and \
child.attrib["class"] == "body details":
message["text"] = child.text.strip()
message['type'] = 'service_message'
if child.attrib["class"] == "body":
for grandchild in child.getchildren():
if grandchild.attrib["class"] == "from_name":
name = grandchild.text.strip()
message["name"] = name
if grandchild.attrib["class"] == "pull_right date details":
message['timestamp'] = grandchild.attrib["title"]
if grandchild.attrib["class"] == "text":
message['text'] = grandchild.text.strip()
message['type'] = 'text'
if grandchild.attrib["class"] == "forwarded body":
message['type'] = "forwarded_message"
if grandchild.attrib["class"] == "media_wrap clearfix":
message['type'] = \
grandchild.getchildren()[0].attrib["class"].split()[-1]
if element.attrib["class"] == "message default clearfix joined":
message["joined_message"] = True
message["name"] = name
if element.attrib["class"] == "message default clearfix":
message["joined_message"] = False
data.append(message)
return data
data = []
filecount = 0
for fname in os.listdir(EXPORT_FOLDER):
fpath = os.path.join(EXPORT_FOLDER, fname)
if os.path.isfile(fpath) and os.path.splitext(fpath)[-1] == ".html":
with open(fpath, encoding='utf8') as f:
# print("Reading", fname, "...")
data += parse_file(f.read())
filecount += 1
df = pd.DataFrame(data)
df["timestamp"] = pd.to_datetime(df["timestamp"], format="%d.%m.%Y %H:%M:%S")
#plot_message_count_by_name(df)
print("Queried ", len(df.index), "raw messages from ", filecount, "files.")
# -
# filter out all service_message
messages = df.loc[df['type'] == "text"]
# Display parsed messages
messages
# -----
# # Visualize
#
# To setup Plotly with Jupyter Lab follow the instructions at [plotly > Getting Started](https://plot.ly/python/getting-started/) - especially the [Jupyter Lab instructions](https://plot.ly/python/getting-started/#jupyterlab-support-python-35)
# +
# Install wordcloud
# #!conda install -c conda-forge wordcloud -y
# -
import plotly.graph_objects as go
# + active=""
# # Plotly-Test
# import plotly.graph_objects as go
# fig = go.Figure(data=go.Bar(y=[2, 3, 1]))
# fig.show()
# -
# # Total Messages
print("Total text messages: ", len(messages.index))
# # Messages by date
# +
# Date
dates = pd.DataFrame( messages['timestamp'].dt.date )
date_counts = pd.DataFrame( dates.stack().value_counts(sort=False) )
data = [go.Bar(x=date_counts.index, y=date_counts[0])]
# Plot chart
fig = go.Figure(data)
fig.show()
# Save to file
filename = "Telegram_by_date"
fig.write_image(OUTPUT_DIR + filename + ".png")
fig.write_image(OUTPUT_DIR + filename + ".svg")
# +
# Year
years = pd.DataFrame( messages['timestamp'].dt.year )
year_counts = pd.DataFrame( years.stack().value_counts(sort=False) )
data = [go.Bar(x=year_counts.index, y=year_counts[0])]
# Plot chart
fig = go.Figure(data)
fig.show()
# Save to file
filename = "Telegram_by_year"
fig.write_image(OUTPUT_DIR + filename + ".png")
fig.write_image(OUTPUT_DIR + filename + ".svg")
# -
# # Messages by Time
#
# +
# Time
hours = pd.DataFrame( pd.DatetimeIndex(messages['timestamp']).hour )
time_counts = pd.DataFrame( hours.stack().value_counts(sort=False) )
data = [go.Bar(x=time_counts.index, y=time_counts[0])]
# Plot chart
fig = go.Figure(data)
fig.show()
# Save to file
filename = "Telegram_by_time"
fig.write_image(OUTPUT_DIR + filename + ".png")
fig.write_image(OUTPUT_DIR + filename + ".svg")
# -
# lbls = list( time_counts.index.astype(str) )
# data = [go.Pie(labels=lbls, values=time_counts[0], hole=.3)] #, marker_colors=night_colors)]
#
# fig.update_traces(textposition='inside', textinfo='label')
# fig = go.Figure(data=data)
# fig.show()
# # Word count
#
message_texts = messages['text']
# +
words = []
for txt in message_texts:
word_tokens = txt.split()
words += word_tokens
message_text_flat = " ".join(words)
print("Total words: ", len(words))
print("Words per message: ", len(words) / len(messages))
# +
# Based on: https://amueller.github.io/word_cloud/auto_examples/single_word.html#sphx-glr-auto-examples-single-word-py
import numpy as np
import matplotlib.pyplot as plt
from wordcloud import WordCloud
#x, y = np.ogrid[:300, :300]
x, y = np.ogrid[:1000, :1000]
#mask = (x - 150) ** 2 + (y - 150) ** 2 > 130 ** 2
mask = (x - 500) ** 2 + (y - 500) ** 2 > 400 ** 2
mask = 255 * mask.astype(int)
wc = WordCloud(background_color="white", repeat=True, mask=mask)
wc.generate(message_text_flat)
# store to file
wc.to_file(OUTPUT_DIR + "Telegram_cloud.png") # PNG
print(wc.to_svg(embed_font=True), file=open(OUTPUT_DIR + "Telegram_cloud.svg", 'w')) # SVG
# show
plt.axis("off")
plt.imshow(wc, interpolation="bilinear")
plt.show()
# -
# ### Emoji Count
# +
# extended Count emojis
# Also counts "Emoji words" (multiple Emojis in a row)
import emoji
TEST_STR = "Get Emoji — All Emojis to ️ Copy and 📋 Paste 👌"
emoji_total_count = 0
emoji_word_total_count = 0
emoji_counts = {}
emoji_word_counts = {}
emoji_word = ""
#for idx, ch in message_text_flat:
for idx in range(len(message_text_flat)):
ch = message_text_flat[idx]
if ch in emoji.unicode_codes.UNICODE_EMOJI:
# Character is an Emoji
emoji_total_count += 1
emoji_word += ch # Append current Emoji to Emoji word
if ch in emoji_counts:
# Increment existing entry
emoji_counts[ch] += 1
else:
# Create new enty in dictionary
emoji_counts[ch] = 1
elif ch == " ":
# Ignore spaces between emojis
;
elif len(emoji_word) > 1:
# Characters (etc.) terminate Emoji word
emoji_word_total_count += 1
if emoji_word in emoji_counts:
# Increment existing entry
emoji_word_counts[emoji_word] += 1
else:
# Create new enty in dictionary
emoji_word_counts[emoji_word] = 1
emoji_word = "" # Start over Emoji word
print("Found", emoji_total_count, "emojis.")
print("Found", emoji_word_total_count, "emoji words.")
# emoji_counts
# -
# ### Emojis
# +
# Visualize Emojis
emoji_counts_df = pd.DataFrame( list(emoji_counts.items()) )
# Sort by frequency / count
emoji_counts_df.sort_values(1, ascending=False, inplace=True) # Sort by col
data = [go.Bar(x=emoji_counts_df[0], y=emoji_counts_df[1])]
# Plot chart
fig = go.Figure(data)
fig.show()
# Save to file
filename = "Telegram_emojis"
fig.write_image(OUTPUT_DIR + filename + ".png")
fig.write_image(OUTPUT_DIR + filename + ".svg")
# -
# ### Emoji Words
# +
# Visualize Emoji words
emoji_word_counts_df = pd.DataFrame( list(emoji_word_counts.items()) )
# Sort by length of Emoji word
emoji_word_counts_df['length'] = emoji_word_counts_df[0].str.len()
emoji_word_counts_df.sort_values('length', ascending=False, inplace=True)
data = [go.Bar(x=emoji_word_counts_df[0], y=emoji_word_counts_df[1])]
# Plot chart
fig = go.Figure(data)
fig.show()
# Save to file
filename = "Telegram_emoji_words"
fig.write_image(OUTPUT_DIR + filename + ".png")
fig.write_image(OUTPUT_DIR + filename + ".svg")
# -
| TelegramAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:python-tutorial]
# language: python
# name: conda-env-python-tutorial-py
# ---
# # Objects in Python
#
# Objects are _instances_ of classes. That is, objects are the constructs that are created in your computer's memory. All objects in Python _inherit_ from the same basic object, which means that every object has the same basic functionality "under the hood."
# ## What type of object?
#
# Python has a built-in function called `type` that will tell you what kind of object something is. In the next cell, we assign a number to a variable `v`, and then we output that variable's type.
i = 1
type(i)
x = 1.234
type(x)
# ## Types are objects, too!
#
# The type of an object is also an object. I know this gets confusing, but don't think too hard about it, yet. Just remember that any object can be assigned to a variable. For example:
i_type = type(i)
type(i_type)
# ## Objects hold data, but they also have functionality of their own!
# Conceptually, objects are things that group data together with functions that act on that data. For example, suppose you have an object `x` that represents an array of data. The object contains the data, but it also might have _methods_ attached to it that do things like compute the _sum_ of the data, or that computes the _mean_ of the data.
i_type
x
type(x)
i_type(x)
type(i_type(x))
# This just converted a `float` into an `int`. But we didn't use the `int` type explicitly. Instead, we used a variable (`i_type`) as a _pointer_ to the `int` type.
# ## Period
#
# _Methods_ of an object are additional functions that "come with" with object and the object's data. To search for _methods_ of an object, use the `.` operator and (in Jupyter) the _TAB_ key to search for methods of a given object.
x
x.
# Did you notice the options?
#
# Note that there is an `is_integer` method attached to this float. What does that do?
# ## Help
#
# Another really handy built-in function in Python is the `help` function. This will give you some simple documentation (if the developer's wrote it!) about the object you are asking about.
help(x)
# There's a _ton_ of information here. If you are curious about any of this, I recommend that you read up on the [Python Data Model](https://docs.python.org/3/reference/datamodel.html).
#
# You can use `help` to give you information about functions, too:
help(print)
# Jupyter notebooks give you a quick shortcut to the help function, too. Just use the `?` symbol.
# +
# print?
# -
# Notice, though, that the information displays is _essentially_ the same, but not exactly.
# Let's look at that `is_integer` method, now:
# +
# x.is_integer?
# -
x.is_integer()
# And that makes sense. (I hope!)
# # Functions are objects, too!
#
# Any function that you define is also an object, which (remember!) means you can assign it to a variable.
def f(g):
print(g)
f
p_f = f
type(p_f)
p_f(x)
# But if you can assign a function to a variable, then you can also pass that variable into a function!
f(p_f)
# ## Variables _point_ to objects (i.e., they are _pointers_)
# One of the biggest confusions in Python is the fact that variables are (essentially) pointer to objects. That is, the variable _symbol_ is a thing that Python uses to reference a particular object in memory.
id(p_f)
# Note: The `hex` function is a built-in function in Python that convert a number to hexadecimal.
#
# Note: The `id` of the `p_f` variable is what is displayed when I print out the variable itself.
hex(id(p_f))
# Now, let's create 2 objects (`list`s) that _look_ the same.
l1 = [1,2,3,4,5]
l2 = [1,2,3,4,5]
# They should have the same _values_, which we can check with the `==` comparison operator:
l1 == l2
# As we expected! But do these two variables _point_ to the same object in memory?
hex(id(l1))
hex(id(l2))
# These are _not_ the same! Python gives you a quick way to check this, using the `is` comparison operator:
# <div class="alert alert-block alert-success">
# <p>Previous: <a href="00_introduction.ipynb">Introduction</a></p>
# <p>Next: <a href="02_operators.ipynb">Operators</a></p>
# </div>
| notebooks/bytopic/python-basics/01_objects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
a=np.arange(1,11)
a
np.insert(a,1,50)
np.insert(a,1,50.5)
np.insert(a,(1,3,7),50)
np.delete(a,2)
a
| insert and delete functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://pythonista.mx)
# ## El paquete _urllib_.
#
# El módulo _urllib_ permite acceder a cualquier recurso publicado en la red (páginas web, archivos, directorios, imágenes, etc) mediante diversos protocolos (HTTP, FTP, SFTP, etc).
#
# ### El módulo _urllib.request_.
#
# El módulo _urllib.request_ permite acceder a un recurso publicado en internet mediante su URL y guardar la respuesta del servidor.
import urllib
help(urllib)
import urllib.request
pagina = urllib.request.urlopen('https://pythonista.io')
print(pagina.read())
# ## Escrutinio de HTML/XML con _BeautifulSoup_.
#
# El paquete *BeautifuSoup* contiene una biblioteca especialziada en análizar y buscar datos dentro de un archivo HTML mediante diversos tipos de criterios como:
#
# * Búsquedas de elementos HTML por medio de la estructura del DOM.
# * Búsquedas por medio de selectores.
# * Búsquedas de etiquetas.
#
# El nombre del paquete una vez instalado es *bs4*.
# !pip install beautifulsoup4
# **Ejemplo:**
#
# Se cargará la página [html/index.html](html/index.html).
#
# **Nota:** Asegúrese que la ruta absoluta al archivo sea correcta.
# %pwd
pagina = urllib.request.urlopen('file://C:\\Users\\josech\\Dropbox\\Codigo\\Pythonista\\Cursos\\py121\\html\\index.html')
html = pagina.read()
print(html)
# ### La clase *bs4.BeautifulSoup*.
#
# Para poder realizar operaciones con un documento HTML es necesario crear un objeto a partir de la clase *bs4.BeautifulSoup* ingresando un objeto tipo *str* que contiene el código HTML y seleccionando el tipo de analizador a utilizar.
#
# ```
# bs4.BeautifulSoup(<objeto tipo str>, <tipo de analizador>)
# ```
#
# Beautifulsoup soporta varios tipos de analizadores del código, pero por defecto se utiliza *'html_parser'*.
#
# Para conocer más sobre las opciones de analizadores puede consultar https://www.crummy.com/software/BeautifulSoup/bs4/doc/#installing-a-parser.
import bs4
sopa = bs4.BeautifulSoup(html, 'html.parser')
type(sopa)
dir(sopa)
# ### Despliegue del código HTML.
# El atributo _prettify()_ permite desplegar el código HTML de forma legible para los humanos.
print(sopa.prettify())
# ### Acceso a elementos mediante DOM.
#
# DOM es el acrónimo de Modelo de Objeto de Documento por sus siglas en inglés y es la forma en la que un navegador interpreta un documento HTML dentro de una ventana.
#
# El DOM presenta una estructura similar a la del tronco de un árbol a partir del cual se bifurcan ramas. Se dice que el elemento HTML que contiene a otros elementos es el padre de estos.
#
# ```
# padre(parent)
# | hijos(children)
# ├──elemento
# ├──hermano(sibling)
# ```
#
# Cuando se realizan búsquedas mediante el DOM, BeautifulSoup regresa el primer elemento con la etiqueta HTML que coincida.
# ### Los objetos de tipo *bs4.element.Tag*.
#
# El resultado es un objeto de tipo *bs4.element.Tag* que también contiene métodos y atributos de búsqueda e información del elemento. Los atributos principales que tienen estos objetos son:
#
# * *name*
# * *text*
# * *attribs*
# * *parent*
# * *children*
# **Ejemplos:**
sopa.title
type(sopa.title)
sopa.title.text
sopa.body.text
[elemento for elemento in sopa.body.children]
sopa.a
sopa.a.name
sopa.a.text
sopa.a.attrs
sopa.footer
sopa.footer.parent.name
[elemento for elemento in sopa.footer.children]
sopa.head
sopa.head.meta
sopa.body.a
sopa.head.meta.attrs
sopa.footer.parent.name
# ### Acceso a elementos mediante selectores.
#
# El método *select()* permite definir búsquedads mendiante selectores, trayendo por resultado un objeto tipo *list* con todos los elementos que coinciden con al búsqueda comop objetos tipo *bs4.element.Tag*.
#
# Para saber más de selectores puede consultar:
#
# [https://developer.mozilla.org/en-US/docs/Learn/CSS/Introduction_to_CSS/Selectors](https://developer.mozilla.org/en-US/docs/Learn/CSS/Introduction_to_CSS/Selectors)
sopa.body.select('a')
sopa.body.select('a')[0]
type(sopa.body.select('a')[0])
# ### Otros métodos para acceder a elementos y datos de HTML.
# Los siguientes métodos también permiten realizar búsquedas de elementos.
# * *find()*.
# * *findall()*.
# * *findNext()*.
# * *findPrevious()*.
# * *fetchParents()*.
# * *fetchNextSiblings()*.
# * *fetchPreviousSiblings()*.
# * *fetchChildren()*.
# * *get_text()*.
# * *get()*.
#
sopa.find_all("a")
sopa.find("a")
sopa.find("a").findNext("a")
sopa.find_all('img')
sopa.find_all('img')[0].parent
sopa.find("div", {"id" : 'contenedor'})
sopa.head.fetchNextSiblings()
sopa.footer.get_text()
sopa.a.get('href')
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2018.</p>
| 10_urllib_beautifulsoup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import onnxmltools
from keras.models import load_model
input_keras_path = 'model.h5' # Change this path to your keras model path
output_onnx_path= 'model.onnx' # Change this path to the output name and path for the onnx model
model = load_model(input_keras_path)
onnx_model = onnxmltools.convert_keras(model)
onnxmltools.utils.save_model(onnx_model, output_onnx_path)
| scripts/keras_onnx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
# default_exp datasets.mts
# -
# # MTS Dataset
# > data from the MTS Kion application on user interactions with content for a period of 6 months.
#
# The presented dataset contains data on users and objects (series / movies), as well as on their interactions (content viewing by the user) from the Kion online cinema. Content view data collected for ~6 months, from 2021-03-13 to 2021-08-22 inclusive, and diluted with random noise. User and content IDs are anonymised.
#
# ### users.csv
# This file contains information about users:
# - user_id - User ID
# - age - user's age group, string like "M_N".
# - 18_24 - from 18 to 24 years old inclusive
# - 25_34 - from 25 to 34 years old inclusive
# - 35_44 - from 35 to 44 years old inclusive
# - 45_54 - from 45 to 54 years old inclusive
# - 55_64 - from 55 to 64 years old inclusive
# - 65_inf - from 65 and older
# - sex - user gender
# - M - man
# - F - woman
# - income - user's income, string like "M_N
# - income_0_20
# - income_20_40
# - income_40_60
# - income_60_90
# - income_90_150
# - income_150_inf
# - kids_flg - flag "presence of a child
#
# ### items.csv
# This file contains information about objects (movies/series):
# - item_id - Content ID
# - content_type - Type of content (movie, series)
# - title - Title in Russian
# - title_orig - original name
# - genres - Genres from source (online movie theaters)
# - countries - country
# - for_kids - flag "content for children"
# - age_rating - age rating
# - studios - studios
# - directors - directors
# - actors - actors
# - keywords - keywords
# - description - description
#
# ### interactions.csv
# This file contains information about user interactions with content:
# - user_id - User ID
# - item_id - Content ID
# - last_watch_dt - Date last viewed
# - total_dur - The total duration of all views of this content in seconds
# - content_type - Type of content (movie, series)
#hide
from nbdev.showdoc import *
from fastcore.nb_imports import *
from fastcore.test import *
# +
#export
from typing import Any, Iterable, List, Optional, Tuple, Union, Callable
import os
import pandas as pd
import numpy as np
from recohut.utils.common_utils import *
from recohut.datasets.bases.interactions import InteractionsDataset
# -
#export
class MTSDataset(InteractionsDataset):
url_users = "https://github.com/RecoHut-Datasets/mts_kion/raw/v1/users.parquet.snappy"
url_items = "https://github.com/RecoHut-Datasets/mts_kion/raw/v1/items.parquet.snappy"
url_inter = "https://github.com/RecoHut-Datasets/mts_kion/raw/v1/interactions.parquet.snappy"
def __init__(self, sample_frac=1, **kwargs):
self.sample_frac = sample_frac
super().__init__(**kwargs)
@property
def raw_file_names(self):
return ['users.parquet.snappy',
'items.parquet.snappy',
'interactions.parquet.snappy']
@property
def processed_file_names(self):
return ['users_processed.csv',
'items_processed.csv',
'interactions_processed.csv',
'item_stats.csv']
def download(self):
_ = download_url(self.url_users, self.raw_dir)
_ = download_url(self.url_items, self.raw_dir)
_ = download_url(self.url_inter, self.raw_dir)
def load_users_df(self):
df = pd.read_parquet(self.raw_paths[0])
return df
def load_items_df(self):
df = pd.read_parquet(self.raw_paths[1])
return df
def load_ratings_df(self):
df = pd.read_parquet(self.raw_paths[2])
df = df.sample(frac=self.sample_frac)
return df
@staticmethod
def add_user_stats(interactions_df, users_df, split_name=''):
"""
Computes user watches stats for particular interactions date split
and adds them to users dataframe with specific name
"""
user_watch_count_all = interactions_df[
interactions_df['total_dur'] > 300].groupby(by='user_id')['item_id'].count()
max_date_df = interactions_df['last_watch_dt'].max()
user_watch_count_last_14 = interactions_df[
(interactions_df['total_dur'] > 300) &
(interactions_df['last_watch_dt'] >= (max_date_df - pd.Timedelta(days=14)))
].groupby(by='user_id')['item_id'].count()
user_watch_count_all.name = split_name + "user_watch_cnt_all"
user_watch_count_last_14.name = split_name + "user_watch_cnt_last_14"
user_watches = pd.DataFrame(user_watch_count_all).join(user_watch_count_last_14,
how='outer')
user_watches.fillna(0, inplace=True)
cols = user_watches.columns
user_watches[cols] = user_watches[cols].astype('int64')
users_df = users_df.join(user_watches, on='user_id', how='outer')
users_df[cols] = users_df[cols].fillna(0)
users_df['age'] = users_df['age'].fillna('age_unknown')
users_df['income'] = users_df['income'].fillna('income_unknown')
users_df['sex'] = users_df['sex'].fillna('sex_unknown')
users_df['kids_flg'] = users_df['kids_flg'].fillna(False)
return users_df
@staticmethod
def add_item_watches_stats(interactions_df, items_df, item_stats):
"""
Computes item watches stats for particular interactions date split
and adds them to item_stats dataframe
"""
def smooth(series, window_size, smoothing_func):
"""Computes smoothed interactions statistics for item"""
series = np.array(series)
ext = np.r_[2 * series[0] - series[window_size - 1::-1],
series,
2 * series[-1] - series[-1:-window_size:-1]]
weights = smoothing_func(window_size)
smoothed = np.convolve(weights / weights.sum(), ext, mode='same')
return smoothed[window_size:-window_size + 1]
def trend_slope(series, window_size=7, smoothing_func=np.hamming):
"""Computes trend slope for item interactions"""
smoothed = smooth(series, window_size, smoothing_func)
return smoothed[-1] - smoothed[-2]
keep = item_stats.columns
max_date = interactions_df['last_watch_dt'].max()
cols = list(range(7))
for col in cols:
watches = interactions_df[
interactions_df['last_watch_dt'] ==
max_date - pd.Timedelta(days=6 - col)]
item_stats = item_stats.join(
watches.groupby('item_id')['user_id'].count(), lsuffix=col)
item_stats.fillna(0, inplace=True)
new_colnames = ['user_id' + str(i) for i in range(1, 7)] + ['user_id']
trend_slope_to_row = lambda row: trend_slope(row[new_colnames],
window_size=7)
item_stats['trend_slope'] = item_stats.apply(trend_slope_to_row,
axis=1)
item_stats['watched_in_7_days'] = item_stats[new_colnames].apply(
sum, axis=1)
item_stats['watch_ts_quantile_95'] = 0
item_stats['watch_ts_median'] = 0
item_stats['watch_ts_std'] = 0
for item_id in item_stats.index:
watches = interactions_df[interactions_df['item_id'] == item_id]
day_of_year = watches['last_watch_dt'].apply(
lambda x: x.dayofyear).astype(np.int64)
item_stats.loc[item_id, 'watch_ts_quantile_95'] = \
day_of_year.quantile(q=0.95, interpolation='nearest')
item_stats.loc[item_id, 'watch_ts_median'] = \
day_of_year.quantile(q=0.5, interpolation='nearest')
item_stats.loc[item_id, 'watch_ts_std'] = day_of_year.std()
item_stats['watch_ts_quantile_95_diff'] = \
max_date.dayofyear - item_stats['watch_ts_quantile_95']
item_stats['watch_ts_median_diff'] = max_date.dayofyear - \
item_stats['watch_ts_median']
watched_all_time = interactions_df.groupby('item_id')['user_id'].count()
watched_all_time.name = 'watched_in_all_time'
item_stats = item_stats.join(watched_all_time, on='item_id', how='left')
item_stats.fillna(0, inplace=True)
added_cols = ['trend_slope',
'watched_in_7_days',
'watch_ts_quantile_95_diff',
'watch_ts_median_diff',
'watch_ts_std',
'watched_in_all_time']
return item_stats[list(keep) + added_cols]
@staticmethod
def add_age_stats(interactions, item_stats, users_df):
"""
Computes watchers age stats for items with particular interactions
date split and adds them to item_stats dataframe
"""
item_stats.reset_index(inplace=True)
interactions = interactions.set_index('user_id').join(
users_df[['user_id', 'sex', 'age', 'income']].set_index('user_id'))
interactions.reset_index(inplace=True)
interactions['age_overall'] = interactions['age'].replace(
to_replace={'age_18_24': 'less_35',
'age_25_34': 'less_35',
'age_35_44': 'over_35',
'age_45_54': 'over_35',
'age_65_inf': 'over_35',
'age_55_64': 'over_35'})
age_stats = interactions.groupby('item_id')['age_overall'] \
.value_counts(normalize=True)
age_stats = pd.DataFrame(age_stats)
age_stats.columns = ['value']
age_stats.reset_index(inplace=True)
age_stats.columns = ['item_id', 'age_overall', 'value']
age_stats = age_stats.pivot(
index='item_id', columns='age_overall', values='value').drop(
'age_unknown', axis=1)
age_stats.fillna(0, inplace=True)
item_stats = item_stats.set_index('item_id').join(age_stats)
item_stats[['less_35', 'over_35']] = item_stats[['less_35', 'over_35']] \
.fillna(0)
item_stats.rename(columns={'less_35': 'younger_35_fraction',
'over_35': 'older_35_fraction'},
inplace=True)
return item_stats
@staticmethod
def add_sex_stats(interactions, item_stats, users_df):
"""
Computes watchers sex stats for items with particular interactions date split
and adds them to item_stats dataframe
"""
item_stats.reset_index(inplace=True)
interactions = interactions.set_index('user_id') \
.join(users_df[['user_id', 'sex', 'age', 'income']]
.set_index('user_id'))
interactions.reset_index(inplace=True)
sex_stats = interactions.groupby('item_id')['sex'] \
.value_counts(normalize=True)
sex_stats = pd.DataFrame(sex_stats)
sex_stats.columns = ['value']
sex_stats.reset_index(inplace=True)
sex_stats.columns = ['item_id', 'sex', 'value']
sex_stats = sex_stats.pivot(index='item_id',
columns='sex',
values='value').drop('sex_unknown', axis=1)
sex_stats.fillna(0, inplace=True)
item_stats = item_stats.set_index('item_id').join(sex_stats)
item_stats[['F', 'M']] = item_stats[['F', 'M']].fillna(0)
item_stats.rename(columns={'F': 'female_watchers_fraction',
'M': 'male_watchers_fraction'},
inplace=True)
return item_stats
@staticmethod
def get_coo_matrix(df,
user_col='user_id',
item_col='item_id',
weight_col=None,
users_mapping={},
items_mapping={}):
if weight_col is None:
weights = np.ones(len(df), dtype=np.float32)
else:
weights = df[weight_col].astype(np.float32)
interaction_matrix = sp.coo_matrix((
weights,
(
df[user_col].map(users_mapping.get),
df[item_col].map(items_mapping.get)
)
))
return interaction_matrix
@staticmethod
def create_mapping():
# Creating items and users mapping
users_inv_mapping = dict(enumerate(df['user_id'].unique()))
users_mapping = {v: k for k, v in users_inv_mapping.items()}
items_inv_mapping = dict(enumerate(df['item_id'].unique()))
items_mapping = {v: k for k, v in items_inv_mapping.items()}
def process(self):
# load data
print('Loading data')
users_df = self.load_users_df()
items_df = self.load_items_df()
interactions_df = self.load_ratings_df()
# users info preprocessing
print('Processing users info')
users_df['age'] = users_df['age'].fillna('age_unknown')
users_df['age'] = users_df['age'].astype('category')
users_df['income'] = users_df['income'].fillna('income_unknown')
users_df['income'] = users_df['income'].astype('category')
users_df['sex'] = users_df['sex'].fillna('sex_unknown')
users_df.loc[users_df.sex == 'М', 'sex'] = 'M'
users_df.loc[users_df.sex == 'Ж', 'sex'] = 'F'
users_df['sex'] = users_df['sex'].astype('category')
users_df['kids_flg'] = users_df['kids_flg'].astype('bool')
# items info preprocessing
print('Processing items info')
items_df['content_type'] = items_df['content_type'].astype('category')
items_df['title'] = items_df['title'].str.lower()
items_df['title_orig'] = items_df['title_orig'].fillna('None')
items_df.loc[items_df['release_year'] < 1980, 'release_novelty'] = 1
items_df.loc[items_df['release_year'] >= 2020, 'release_novelty'] = 6
novelty = 1
for i in range(1980, 2020, 10):
novelty += 1
items_df.loc[(items_df['release_year'] >= i) &
(items_df['release_year'] < i + 10), 'release_novelty'] = novelty
items_df = items_df.drop(columns=['release_year'])
items_df['for_kids'] = items_df['for_kids'].fillna(0)
items_df['for_kids'] = items_df['for_kids'].astype('bool')
items_df.loc[items_df.age_rating.isna(), 'age_rating'] = 0
items_df['age_rating'] = items_df['age_rating'].astype('category')
items_df['genres_list'] = items_df['genres'].apply(lambda x: x.split(', '))
num_genres = pd.Series(np.hstack(items_df['genres_list'].values)).value_counts()
items_df['genres_min'] = items_df['genres_list'].apply(
lambda x: min([num_genres[el] for el in x]))
items_df['genres_max'] = items_df['genres_list'].apply(
lambda x: max([num_genres[el] for el in x]))
items_df['genres_med'] = items_df['genres_list'].apply(
lambda x: (np.median([num_genres[el] for el in x])))
items_df['countries'].fillna('None', inplace=True)
items_df['countries'] = items_df['countries'].str.lower()
items_df['countries_list'] = items_df['countries'].apply(
lambda x: x.split(', ') if ', ' in x else [x])
num_countries = pd.Series(np.hstack(items_df['countries_list'].values)).value_counts()
items_df['countries_max'] = items_df['countries_list'].apply(
lambda x: max([num_countries[el] for el in x]))
items_df['studios'].fillna('None', inplace=True)
items_df['studios'] = items_df['studios'].str.lower()
items_df['studios_list'] = items_df['studios'].apply(
lambda x: x.split(', ') if ', ' in x else [x])
num_studios = pd.Series(np.hstack(items_df['studios_list'].values)).value_counts()
items_df['studios_max'] = items_df['studios_list'].apply(
lambda x: max([num_studios[el] for el in x]))
items_df.drop(['countries_list', 'genres_list', 'studios_list'],
axis=1, inplace=True)
# interactions preprocessing
print('Processing interactions')
interactions_df['watched_pct'] = interactions_df['watched_pct'].astype(
pd.Int8Dtype())
interactions_df['watched_pct'] = interactions_df['watched_pct'].fillna(0)
interactions_df['last_watch_dt'] = pd.to_datetime(
interactions_df['last_watch_dt'])
interactions_df.sort_values(by='last_watch_dt', inplace=True)
# user stats feature engineering
print('Processing users stats')
max_date = interactions_df['last_watch_dt'].max()
boosting_split_date = max_date - pd.Timedelta(days=14)
interactions_boost = interactions_df[
interactions_df['last_watch_dt'] <= boosting_split_date]
users_df = self.add_user_stats(interactions_boost, users_df, split_name='boost_')
users_df = self.add_user_stats(interactions_df, users_df, split_name='')
# Item stats
print('Processing items stats')
item_stats = items_df[['item_id']]
item_stats = item_stats.set_index('item_id')
item_stats = self.add_item_watches_stats(interactions_boost, items_df, item_stats)
item_stats.fillna(0, inplace=True)
item_stats = self.add_sex_stats(interactions_boost, item_stats, users_df)
item_stats = self.add_age_stats(interactions_boost, item_stats, users_df)
# Saving preprocessed files
users_df.to_csv(self.processed_paths[0], index=False)
items_df.to_csv(self.processed_paths[1], index=False)
interactions_df.to_csv(self.processed_paths[2], index=False)
item_stats.to_csv(self.processed_paths[3], index=True)
# Example
#hide
# !apt-get -qq install tree
# !pip install -q watermark
# !pip install -q pytorch-lightning
ds = MTSDataset(data_dir='/content/data')
# !tree --du -h -C ./data
# > **References**
# > - https://ods.ai/tracks/recsys-course2021/competitions/competition-recsys-21
# > - https://github.com/blondered/ods_MTS_RecSys_Challenge_solution
#hide
# %reload_ext watermark
# %watermark -a "Sparsh A." -m -iv -u -t -d -p recohut
| nbs/datasets/datasets.mts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gap Framework - Natural Language Processing
# ## Splitter Module
#
# <b>[Github] (https://github.com/andrewferlitsch/gap)</b>
# # Automated PDF, Fax, Image Capture Text Extraction with Gap (Session 1)
#
# Let's start with the basics. We will be using the <b style='color: saddlebrown'>SPLITTER</b> module in the **Gap** Framework.
#
# Steps:
# 1. Import the <b style='color: saddlebrown'>Document</b> and <b style='color: saddlebrown'>Page</b> class from the <b style='color: saddlebrown'>splitter</b> module.
# 2. Create a <b style='color: saddlebrown'>Document</b> object.
# 3. Pass a PDF (text or scanned), Facsimile (TIFF) or image captured document to the <b style='color: saddlebrown'>Document</b> object.
# 4. Wait for the results :)
# let's go to the directory where Gap Framework is installed
import os
os.chdir("../")
# %ls
# import Document and Page from the document module
from gapnlp.splitter import Document, Page
# ## <span style='color: saddlebrown'>Document</span> Object
#
# The initializer (constructor) takes the following arguments:<br/>
#
# document - path to the document
# dir - directory where to store extracted pages and text
# ehandler - function to invoke when processing is completed in asynchronous mode
# config - configuration settings for SYNTAX module
#
# Let's start by preprocessing a 105 page PDF, which is a medical benefits plan. We should see:
#
# - Split into individual PDF pages
# - Text extracted from each page
# - Individual page PDF and text stored in specified directory.
#
# *Note, for brevity, we reduced the size of the PDF document to 10 pages for this code along.*
doc = Document("train/10nc.pdf", "train/nc")
# Ok, we are done! Let's look at the last page (page 105 ~ page 10 in the shorten version).
#
# Wow, that's the foreign language translation page - see how it handles other (non-latin) character sets.
# +
# Let's use the name property to see the name of the document
print( doc.name )
# Use the len() operator to find out how many pages are in the document
print( len(doc) )
# -
# ## <span style='color: saddlebrown'>Page</span> Object
#
# Let's now dive deeper. When the document was processed, each page was put into a <b style='color: saddlebrown'>Page</b> object. Here are some things we can do:
#
# 1. Walk thru each page sequentially as an array index (list).<br/>
# 2. See the original text from the page.<br/>
# 3. See the "default" NLP preprocessing of the text on the page (which can be modified with *config* settings).<br/>
#
# +
# Let's take a look at one of the pages
pages = doc.pages
# total number of pages
print(len(pages))
# Last page in the document
pages[9]
# -
# Let's look at the text for that page (page 10)
page = pages[9]
page.text
# Let's look at the default NLP preprocessing of the text (stemming, stopword removal, punct removal)
page.words
# We can see that some words appear a lot, like preventive, health and protection. Let's get information on the distribution of words in the page. There are two properties we can use for this purpose:
#
# freqDist - The count of the number of occurrences of each word.
# termFreq - The percentage the word appears on the page (TF -> Term Frequency).
# Let's see the frequency distribution (word counts) for the page
page.freqDist
# Let's see the term frequency (TF)
page.termFreq
# ## <span style='color: saddlebrown'>Document</span> Object (Advanced)
#
# Let's look at more advanced features of the <b style='color:saddlebrown'>Document</b> object.
#
# 1. Word Count and Term Frequency
# 2. Save and Restore
# 3. Asychronous Processing of Documents
# 4. Scanned PDF / OCR
# ### Frequency Distribution
#
# Let's look at a frequency distribution (word count) for the whole document. Note that if we look at just the top 10 word counts (after removing stopwords), it is very clear what the document is about: service, benefit, cover, health, medical, care, coverage, ...
#
# If we look at the top 25 word counts, we can see secondary classification indicators, like: plan, medication, treatment, deductible, eligible, dependent, hospital, claim, authorization, prescription and limit.
#
# HINT: It's a Healthcare Benefit Plan.
doc.freqDist
# ### (Re) Load
#
# When a <b style='color:saddlebrown'>Document</b> object is created, the individual PDF pages, text extraction and NLP analysis are stored.
#
# The document can then be subsequently reloaded from storage without reprocessing.
# Let's first delete the Document object from memory
doc = None
# Let's reload the document from storage.
doc = Document()
doc.load("train/10nc.pdf", "train/nc")
# Let's show some examples of how the document was reconstructed from memory.
# Document Name, Number of Pages
print(doc.document)
print(len(doc))
# Let's print text from the last page
page = doc[9]
page.text
# Let's print the word (count) frequency distribution
doc.freqDist
# ### Async Execution
#
# Let's say you have PDF files arriving for processing in real-time from various sources. The *ehandler* option provides asynchronous processing of documents. When this option is specified, the document is processed on an independent process thread, and when complete the specified event handler is called.
# +
def done(document):
print("EVENT HANDLER: done")
doc = Document("train/crash_2015.pdf", "train/crash", ehandler=done)
# -
# Let's get a frequency distribution for this document. BTW, it is a 2015 State of Oregon table of crash statistics (single page) from a multi-page report. Note how the top ten words (after stopword removal) indicate what the document is about: serious, injury, fatal, crash, highway, death.
doc.freqDist
# ### Scanned PDF / OCR
#
# Let's now process a scanned PDF. That's a PDF which is effective a scanned image of a text document, which is then wrapped inside a PDF.
#
# - Split into pages
# - Extract page image
# - OCR the image image into text
# - Extract the text
# +
Document.SCANCHECK = 0
# OCR the scanned PDF and extract text
doc = Document("train/4scan.pdf", "train/4scan")
# -
# Let's now look at a few properties of the preprocessed document.
# +
# The scanned property indicates the document was a scanned PDF (true)
print( doc.scanned )
# Let's print the number of pages
print( len(doc) )
# -
# Let's now look at a page.
# +
# Get the first page
page = doc[0]
page.text
# -
#clean dir
import shutil
shutil.rmtree('train/nc')
shutil.rmtree('train/4scan')
shutil.rmtree('train/crash')
# ## END OF SESSION 1
| train/session 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import subprocess
import matplotlib
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Preprocessing Functions
def load_dat(path):
print('Loading "%s" ...' % path)
with open(path, 'rb') as f:
n, k = np.fromfile(f, dtype=np.int32, count=2)
dat = np.fromfile(f, dtype=np.float32, count=n*k).reshape((n, k))
# Make sure that we've reached EOF
assert len(f.read(1)) == 0
print('=> shape: (%d, %d)' % dat.shape)
# Transpose `dat` to make summations over data index fast.
# Convert to double precision because PCA turned out a bit unstable with single precision.
return dat.T.astype(np.float64)
def fit_pca(dat, out_ks=[16, 32, 64]):
mean = dat.mean(axis=1, keepdims=True)
normalized = dat - mean
variance = (normalized ** 2).mean(axis=1, keepdims=True)
cov_unscaled = normalized.dot(normalized.T) / normalized.shape[1]
normalized /= np.sqrt(variance)
cov_scaled = normalized.dot(normalized.T) / normalized.shape[1]
del normalized
assert np.allclose(np.diag(cov_unscaled), variance[:, 0])
assert np.allclose(np.diag(cov_scaled), np.ones((variance.shape[0],)))
eigvals, eigvecs = np.linalg.eigh(cov_scaled)
assert np.allclose(eigvecs.dot(np.diag(eigvals)).dot(eigvecs.T), cov_scaled)
assert np.allclose(eigvecs.T.dot(cov_scaled).dot(eigvecs), np.diag(eigvals))
transform = eigvecs.T.copy() / np.sqrt(variance.reshape((1, -1)) * eigvals.reshape((-1, 1)))
assert np.allclose(transform.dot(cov_unscaled).dot(transform.T), np.eye(len(eigvals)))
fig, ax = plt.subplots()
ax.plot(range(len(eigvals), 0, -1), eigvals)
for k in out_ks:
ax.plot((k, k), (eigvals[0], eigvals[-1]), ':', label='k=%d' % k)
ax.set_yscale('log')
ax.legend()
return mean, transform
def reduce_dim(dat, mean, transform, base_out_path, out_ks=[16, 32, 64]):
for k in out_ks:
small = transform[-k:, :].dot(dat - mean).T.astype(np.float32)
out_path = '%s-k%d.np' % (base_out_path, k)
print('Saving (%d, %d) array to "%s" ...' % tuple(list(small.shape) + [out_path]))
with open(out_path, 'wb') as out_file:
np.array(small.shape, dtype=np.int32).tofile(out_file)
small.tofile(out_file)
print('Done.')
# ## Process Wikipedia-500K
wdat = load_dat('../../dat/Wikipedia-500K/train-features-full.np')
wmean, wtransform = fit_pca(wdat)
reduce_dim(wdat, wmean, wtransform, '../../dat/Wikipedia-500K/train-features')
del wdat
wdat_valid = load_dat('../../dat/Wikipedia-500K/valid-features-full.np')
reduce_dim(wdat_valid, wmean, wtransform, '../../dat/Wikipedia-500K/valid-features')
del wdat_valid
wdat_test = load_dat('../../dat/Wikipedia-500K/test-features-full.np')
reduce_dim(wdat_test, wmean, wtransform, '../../dat/Wikipedia-500K/test-features')
del wdat_test, wmean, wtransform
# ## Process Amazon-670K
adat = load_dat('../../dat/Amazon-670K/train-features-full.np')
amean, atransform = fit_pca(adat)
reduce_dim(adat, amean, atransform, '../../dat/Amazon-670K/train-features')
del adat
adat_valid = load_dat('../../dat/Amazon-670K/valid-features-full.np')
reduce_dim(adat_valid, amean, atransform, '../../dat/Amazon-670K/valid-features')
del adat_valid
adat_test = load_dat('../../dat/Amazon-670K/test-features-full.np')
reduce_dim(adat_test, amean, atransform, '../../dat/Amazon-670K/test-features')
del adat_test, amean, atransform
| preprocess-extreme-predicton/pca.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
from IPython import display
import matplotlib.pyplot as plt
import torch
from torch import nn
import torchvision
import torchvision.transforms as transforms
import time
import sys
sys.path.append("../")
import d2lzh1981 as d2l
from tqdm import tqdm
print(torch.__version__)
print(torchvision.__version__)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# -
mnist_train = torchvision.datasets.FashionMNIST(root='/Users/nick/Documents/dataset/FashionMNIST2065',
train=True, download=False)
mnist_test = torchvision.datasets.FashionMNIST(root='/Users/nick/Documents/dataset/FashionMNIST2065',
train=False, download=False)
num_id = 0
for x, y in mnist_train:
if num_id % 1000 == 0:
print(num_id)
x.save("/Users/nick/Documents/dataset/FashionMNIST_img/train/{}_{}.png".format(y, num_id))
num_id += 1
num_id = 0
for x, y in mnist_test:
if num_id % 1000 == 0:
print(num_id)
x.save("/Users/nick/Documents/dataset/FashionMNIST_img/test/{}_{}.png".format(y, num_id))
num_id += 1
mnist_train = torchvision.datasets.FashionMNIST(root='/Users/nick/Documents/dataset/FashionMNIST2065',
train=True, download=False, transform=transforms.ToTensor())
mnist_test = torchvision.datasets.FashionMNIST(root='/Users/nick/Documents/dataset/FashionMNIST2065',
train=False, download=False, transform=transforms.ToTensor())
# +
def vgg_block(num_convs, in_channels, out_channels): #卷积层个数,输入通道数,输出通道数
blk = []
for i in range(num_convs):
if i == 0:
blk.append(nn.Conv2d(in_channels, out_channels, kernel_size=3, padding=1))
else:
blk.append(nn.Conv2d(out_channels, out_channels, kernel_size=3, padding=1))
blk.append(nn.ReLU())
blk.append(nn.MaxPool2d(kernel_size=2, stride=2)) # 这里会使宽高减半
return nn.Sequential(*blk)
def vgg(conv_arch, fc_features, fc_hidden_units=4096):
net = nn.Sequential()
# 卷积层部分
for i, (num_convs, in_channels, out_channels) in enumerate(conv_arch):
# 每经过一个vgg_block都会使宽高减半
net.add_module("vgg_block_" + str(i+1), vgg_block(num_convs, in_channels, out_channels))
# 全连接层部分
net.add_module("fc", nn.Sequential(d2l.FlattenLayer(),
nn.Linear(fc_features, fc_hidden_units),
nn.ReLU(),
nn.Dropout(0.5),
nn.Linear(fc_hidden_units, fc_hidden_units),
nn.ReLU(),
nn.Dropout(0.5),
nn.Linear(fc_hidden_units, 10)
))
return net
def evaluate_accuracy(data_iter, net, device=None):
if device is None and isinstance(net, torch.nn.Module):
# 如果没指定device就使用net的device
device = list(net.parameters())[0].device
acc_sum, n = 0.0, 0
with torch.no_grad():
for X, y in data_iter:
if isinstance(net, torch.nn.Module):
net.eval() # 评估模式, 这会关闭dropout
acc_sum += (net(X.to(device)).argmax(dim=1) == y.to(device)).float().sum().cpu().item()
net.train() # 改回训练模式
else: # 自定义的模型, 3.13节之后不会用到, 不考虑GPU
if('is_training' in net.__code__.co_varnames): # 如果有is_training这个参数
# 将is_training设置成False
acc_sum += (net(X, is_training=False).argmax(dim=1) == y).float().sum().item()
else:
acc_sum += (net(X).argmax(dim=1) == y).float().sum().item()
n += y.shape[0]
return acc_sum / n
# +
batch_size = 100
if sys.platform.startswith('win'):
num_workers = 0
else:
num_workers = 4
train_iter = torch.utils.data.DataLoader(mnist_train, batch_size=batch_size,
shuffle=True, num_workers=num_workers)
test_iter = torch.utils.data.DataLoader(mnist_train, batch_size=batch_size,
shuffle=False, num_workers=num_workers)
# +
conv_arch = ((1, 1, 64), (1, 64, 128))
# 经过5个vgg_block, 宽高会减半5次, 变成 224/32 = 7
fc_features = 128 * 7 * 7 # c * w * h
fc_hidden_units = 4096 # 任意
# ratio = 8
# small_conv_arch = [(1, 1, 64//ratio), (1, 64//ratio, 128//ratio), (2, 128//ratio, 256//ratio),
# (2, 256//ratio, 512//ratio), (2, 512//ratio, 512//ratio)]
# net = vgg(small_conv_arch, fc_features // ratio, fc_hidden_units // ratio)
# +
net = vgg(conv_arch, fc_features, fc_hidden_units)
lr, num_epochs = 0.001, 5
optimizer = torch.optim.Adam(net.parameters(), lr=lr)
net = net.to(device)
print("training on ", device)
loss = torch.nn.CrossEntropyLoss()
# -
for epoch in range(num_epochs):
train_l_sum, train_acc_sum, n, batch_count, start = 0.0, 0.0, 0, 0, time.time()
for X, y in tqdm(train_iter):
X = X.to(device)
y = y.to(device)
y_hat = net(X)
l = loss(y_hat, y)
optimizer.zero_grad()
l.backward()
optimizer.step()
train_l_sum += l.cpu().item()
train_acc_sum += (y_hat.argmax(dim=1) == y).sum().cpu().item()
n += y.shape[0]
batch_count += 1
test_acc = evaluate_accuracy(test_iter, net)
print('epoch %d, loss %.4f, train acc %.3f, test acc %.3f, time %.1f sec'
% (epoch + 1, train_l_sum / batch_count, train_acc_sum / n, test_acc, time.time() - start))
test_acc = evaluate_accuracy(test_iter, net)
test_acc
# +
for X, y in train_iter:
X = X.to(device)
predict_y = net(X)
print(y)
print(predict_y.argmax(dim=1))
break
# predict_y.argmax(dim=1)
# -
| ReadBook/zh_gluon_ai/fashion-mnist/VGG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import shared
import json
ve_or_ju = "ju"
runs_df = pd.read_csv(f'data/runs_{ve_or_ju}.tsv', delimiter="\t")
# +
country_counts = runs_df["team_country"].value_counts()
top_country_counts = country_counts[country_counts > 50]
top_countries = top_country_counts.keys().tolist()
display(top_countries)
with open(f"data/top_countries_{ve_or_ju}.json", 'w') as outfile:
json.dump(top_countries, outfile)
# +
runs_df["first_name"] = runs_df.name.str.split(" ", expand=True).iloc[:, 0]
fn_counts = runs_df["first_name"].value_counts()
top_fn_counts = fn_counts[fn_counts > 20]
top_first_names = top_fn_counts.keys().tolist()
display(top_first_names)
with open(f"data/top_first_names_{ve_or_ju}.json", 'w') as outfile:
json.dump(top_first_names, outfile)
# -
# Temporarily remove 2018 in order to try predict it in other notebook
runs_df = runs_df[runs_df.year != 2018]
runs_df
features = shared.preprocess_features(runs_df, top_countries, top_first_names)
features.head(10)
features.info()
# +
x = features.values
#x = features[["team_id", "team_id_log10", "team_id_log100", "team_id_log2", "team_id_square", "leg_id_1", "leg_id_2", "leg_id_3", "leg_id_4", "leg_id_5", "leg_id_6", "leg_id_7"]].values # Poista tää.
y = np.log(runs_df.pace.values)
y = y.reshape(len(y), 1)
display(x.shape)
display(y.shape)
# -
import sklearn
from sklearn import linear_model
from sklearn import ensemble
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import train_test_split
import joblib
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.2, random_state=2019)
x_train.shape
import matplotlib.pyplot as plt
def fit_and_test_model(model, x_train, x_test, y_train, y_test, fit_params={}):
model.fit(x_train, y_train.ravel(), **fit_params)
y_pred = np.exp(model.predict(x_test))
print(f"Shapes: y_test={np.exp(y_test).shape} y_pred={y_pred.shape}")
print("Mean squared error: %.3f" % mean_squared_error(np.exp(y_test), y_pred))
print('Explained variance score: %.3f' % r2_score(np.exp(y_test), y_pred))
plt.scatter(x_test[:,0], np.exp(y_test), color='red', alpha=0.01)
plt.scatter(x_test[:,0], y_pred, color='blue', alpha=0.01)
plt.ylim(4, 20)
plt.show()
y_train.shape
# %%time
linear = linear_model.LinearRegression()
fit_and_test_model(linear, x_train, x_test, y_train, y_test)
pd.DataFrame({'name':features.keys(), 'coef':linear.coef_})
# +
# %%time
gbr_num_estimators=7100
gbr = sklearn.ensemble.GradientBoostingRegressor(n_estimators=gbr_num_estimators, random_state=0, verbose=1)
fit_and_test_model(gbr, x_train, x_test, y_train, y_test)
#print(f"feature_importances_: {gbr.feature_importances_}")
#gbr_features = pd.DataFrame({'feature':first_names.columns, 'importance': gbr.feature_importances_})
#gbr_features['feature'] = gbr_features['feature'].str.replace('top_first_name_','')
#display(gbr_features.sort_values(by="importance", ascending=False))
# -
gbr_num_estimators_quantile=int(gbr_num_estimators/2)
gbr_q_low = sklearn.ensemble.GradientBoostingRegressor(loss='quantile', alpha=0.159, n_estimators=gbr_num_estimators_quantile, random_state=0, verbose=1)
fit_and_test_model(gbr_q_low, x_train, x_test, y_train, y_test)
# +
gbr_q_high = sklearn.ensemble.GradientBoostingRegressor(loss='quantile', alpha=0.841, n_estimators=gbr_num_estimators_quantile, random_state=0, verbose=1)
fit_and_test_model(gbr_q_high, x_train, x_test, y_train, y_test)
# -
joblib.dump(gbr, 'gbr.sav')
joblib.dump(gbr_q_low, 'gbr_q_low.sav')
joblib.dump(gbr_q_high, 'gbr_q_high.sav')
gbr_preds = gbr.predict(pd.DataFrame(x_test))
gbr_q_low_preds = gbr_q_low.predict(pd.DataFrame(x_test))
gbr_q_high_preds = gbr_q_high.predict(pd.DataFrame(x_test))
# +
gbr_q_pred_errors = pd.DataFrame({
'q_low':np.exp(gbr_q_low_preds),
'true':np.exp(y_test).ravel(),
'predicted':np.exp(gbr_preds),
'q_high':np.exp(gbr_q_high_preds),
})
gbr_q_pred_errors["q_low_error"] = gbr_q_pred_errors.true < gbr_q_pred_errors.q_low
gbr_q_pred_errors["q_high_error"] = gbr_q_pred_errors.true > gbr_q_pred_errors.q_high
gbr_q_pred_errors["q_error"] = np.logical_or(gbr_q_pred_errors.q_low_error, gbr_q_pred_errors.q_high_error)
# Intentionally don't use log scale for calculation to get bigger std
gbr_q_pred_errors["std"] = (gbr_q_pred_errors.q_high - gbr_q_pred_errors.q_low) / 2
gbr_q_pred_errors["std_correct"] = np.exp((gbr_q_high_preds - gbr_q_low_preds) / 2)
gbr_q_pred_errors["abs_error"] = np.abs(gbr_q_pred_errors.predicted - gbr_q_pred_errors.true)
gbr_q_pred_errors["abs_error_in_stds"] = gbr_q_pred_errors.abs_error / np.exp(gbr_q_pred_errors["std_correct"])
display(gbr_q_pred_errors.tail(15).round(3))
display(gbr_q_pred_errors.q_low_error.mean())
display(gbr_q_pred_errors.q_high_error.mean())
display(gbr_q_pred_errors.q_error.mean())
display(gbr_q_pred_errors["std"].mean())
display(gbr_q_pred_errors["std_correct"].mean())
display(gbr_q_pred_errors["abs_error_in_stds"].mean())
# -
np.exp(1.138)
# %%date
STOP_HERE
# +
import os
#os.environ['MKL_THREADING_LAYER'] = 'GNU'
os.environ['THEANO_FLAGS'] = 'device=cpu'
#os.environ['THEANO_FLAGS'] = 'device=cuda,floatX=float32,force_device=True'
import pymc3 as pm
import pmlearn
from pmlearn.linear_model import LinearRegression
print('Running on pymc-learn v{}'.format(pmlearn.__version__))
# -
import multiprocessing
multiprocessing.cpu_count()
# +
pmlearn_linear = LinearRegression()
fit_params={
"inference_type": "nuts",
"inference_args": {
"cores": multiprocessing.cpu_count() -1,
#"chains":2,
"init": 'adapt_diag',
#"tune": 2000,
"target_accept": 0.9999
}
}
fit_and_test_model(pmlearn_linear, pd.DataFrame(data=x_train), pd.DataFrame(x_test), y_train, y_test,fit_params)
# -
pmlearn_preds = pmlearn_linear.predict(pd.DataFrame(x_test), return_std=True)
# +
joblib.dump(pmlearn_linear, 'pmlearn_linear.sav')
# -
pred_errors = pd.DataFrame({
'mean':np.exp(pmlearn_preds[0]),
'std':np.exp(pmlearn_preds[1]),
'true':np.exp(y_test).ravel(),
'error':np.abs(np.exp(y_test).ravel() -np.exp(pmlearn_preds[0])) / np.exp(pmlearn_preds[1])
})
display(pred_errors.head(15))
pred_errors.error.mean()
pmlearn_linear.plot_elbo()
pm.traceplot(pmlearn_linear.trace)
pm.forestplot(pmlearn_linear.trace, varnames=["betas", "alpha", "s"]);
summary_df = pm.summary(pmlearn_linear.trace, varnames=["betas", "alpha", "s"])
summary_df
pm.plot_posterior(pmlearn_linear.trace, varnames=["betas", "alpha", "s"],
figsize = [14, 8])
pm.gelman_rubin(pmlearn_linear.trace, varnames=["betas", "alpha", "s"])
| preprocess-priors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importing Libraries
import pandas as pd
import numpy as np
import seaborn as sns
# ## Reading Data
patient_info = pd.read_csv('../../data/PatientInfo.csv')
patient_info.head()
patient_info.describe()
patient_info.hist();
sns.heatmap(patient_info.corr(), annot=True, fmt=".2f");
sns.pairplot(patient_info.select_dtypes(include=['float']).dropna());
# ## Questions
# #### 1. How long does the infection endure in different types of patients? (Descriptive)
# #### 2. How did the disease spread through the country over time? (Descriptive)
# #### 3. How did the disease spread within the provinces over time? (Descriptive)
# #### 4. What characteristics are good predictors of the patient status? (Prediction / Inference)
# #### 5. What characteristics are good predictors of the patient time of recovery? (Prediction / Inference)
# #### 6. Do weather changes correlate to the spread of the disease? (Inference)
# #### 7. Do weather changes correlate to the patient status / time of recovery? (Inference)
# #### 8. Does the locomotion of the patients correlate to the spread of the disease? (Inference)
# #### 9. Does the distance covered by the patients (route) correlate to their status / time of recovery? (Descriptive / Exploratory)
# #### 10. What are the riskiest COVID-19 infection-prone places in Seoul (where most infected people have been)? (Descriptive / Exploratory)
# ## Selected Questions
# #### 3. How did the disease spread within the provinces over time? (Descriptive)
# #### 4. What characteristics are good predictors of the patient status? (Prediction / Inference)
# #### 10. What are the riskiest COVID-19 infection-prone places in Seoul (where most infected people have been)?
| code/draft-notebooks/data-exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
par_dir = os.path.abspath(os.path.join(os.getcwd(), os.pardir))
if par_dir not in sys.path:
sys.path.append(par_dir)
# -
import numpy as np
run = 7
softmaxes = np.load("softmax" + str(run) + ".npy")
labels = np.load("labels" + str(run) + ".npy")
predictions = np.load("predictions" + str(run) + ".npy")
energies = np.load("energies" + str(run) + ".npy")
# +
label_dict_1 = {"gamma":0, "e":1}
label_dict_2 = {"gamma":0, "mu":2}
label_dict_3 = {"e":1, "muon":2}
label_dict_all = {"gamma":0, "e":1, "mu":2}
# -
# %matplotlib inline
# # ROC Curve fix
#
# Fix the ROC curve such that it is one vs the other not one vs all
from plot_utils import plot_utils
plot_utils.plot_ROC_curve_one_vs_one(softmaxes, labels, label_dict_all, "gamma", "e")
plot_utils.plot_ROC_curve_one_vs_one(softmaxes, labels, label_dict_all, "e", "gamma")
plot_utils.plot_ROC_curve_one_vs_one(softmaxes, labels, label_dict_all, "gamma", "mu")
plot_utils.plot_ROC_curve_one_vs_one(softmaxes, labels, label_dict_all, "mu", "gamma")
| notebooks/notebooks_archive/scratch_nb_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize OK
from client.api.notebook import Notebook
ok = Notebook('lab04.ok')
# -
# # Lab 4: Principal Component Analysis
#
# In this lab assignment, we will walk through an example of using Principal Component Analysis (PCA) on a dataset involving [iris plants](https://en.wikipedia.org/wiki/Iris_(plant)).
from sklearn.datasets import load_iris
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# To begin, run the following cell to load the dataset into this notebook.
# * `iris_features` will contain a numpy array of 4 attributes for 150 different plants (shape 150 x 4).
# * `iris_target` will contain the class of each plant. There are 3 classes of plants in the dataset: Iris-Setosa, Iris-Versicolour, and Iris-Virginica. The class names will be stored in `iris_target_names`.
# * `iris_feature_names` will be a list of 4 names, one for each attribute in `iris_features`.
#
# Additional information on the dataset will be included in the description printed at the end of the following cell.
# +
iris_data = load_iris() # Loading the dataset
# Unpacking the data into arrays
iris_features = iris_data['data']
iris_target = iris_data['target']
iris_feature_names = iris_data['feature_names']
iris_target_names = iris_data['target_names']
# Convert iris_target to string labels instead of int labels currently (0, 1, 2) for the classes
iris_target = iris_target_names[iris_target]
#print(iris_data['DESCR'])
iris_target
# + [markdown] deletable=false editable=false
# ## Question 1
#
# Let's explore the data by creating a scatter matrix of our iris features. To do this, we'll create 2D scatter plots for every possible pair of our four features. This should result in six total scatter plots in our scatter matrix.
#
# Complete the code below using `sns.scatterplot` to create the scatter matrix.
#
# **Hint:** Use the `hue` argument of `sns.scatterplot` to color the points by class. A legend should then appear in each scatter plot automatically.
#
# <!--
# BEGIN QUESTION
# name: q1
# -->
# -
plt.figure(figsize=(14, 10))
plt.suptitle("Scatter Matrix of Iris Features")
plt.subplots_adjust(wspace=0.3, hspace=0.3)
for i in range(1, 4):
for j in range(i):
plt.subplot(3, 3, i+3*j)
...
plt.xlabel(iris_feature_names[i])
plt.ylabel(iris_feature_names[j])
# + [markdown] deletable=false editable=false
# ## Question 2a
#
# To apply PCA, we will first need to "center" the data so that the mean of each feature is 0. Additionally, we will need to scale the centered data by $\frac{1}{\sqrt n}$, where $n$ is the number of samples (rows) we have in our dataset.
#
# Compute the columnwise mean of `iris_features` in the cell below and store it in `iris_mean` (should be a numpy array of 4 means, 1 for each attribute). Then, subtract `iris_mean` from `iris_features`, divide the result by the $\sqrt n$, and save the result in `normalized_features`.
#
# **Hints:**
# * Use `np.mean` or `np.average` to compute `iris_mean`, and pay attention to the `axis` argument.
# * If you are confused about how numpy deals with arithmetic operations between arrays of different shapes, see this note about [broadcasting](https://docs.scipy.org/doc/numpy/user/basics.broadcasting.html) for explanations/examples.
#
# <!--
# BEGIN QUESTION
# name: q2a
# -->
# -
n = iris_features.shape[0] # should be 150
iris_mean = ...
normalized_features = ...
# + deletable=false editable=false
ok.grade("q2a");
# + [markdown] deletable=false editable=false
# ## Question 2b
#
# As you may recall from lecture, PCA is a specific application of the singular value decomposition (SVD) for matrices. In the following cell, let's use the [`np.linalg.svd`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.svd.html) function compute the SVD of our `normalized_features`. Store the left singular vectors, singular values, and right singular vectors in `u`, `s`, and `vt` respectively.
#
# **Hint:** Set the `full_matrices` argument of `np.linalg.svd` to `False`.
#
# <!--
# BEGIN QUESTION
# name: q2b
# -->
# -
...
u.shape, s, vt.shape
# + deletable=false editable=false
ok.grade("q2b");
# + [markdown] deletable=false editable=false
# ## Question 2c
#
# What can we learn from the singular values in `s`? First, we can compute the total variance of the data by summing the squared singular values. We will later be able to use this value to determine the variance captured by a subset of our principal components.
#
# Compute the total variance below by summing the square of `s` and store the result in the variable `total_variance`.
#
# <!--
# BEGIN QUESTION
# name: q2c
# -->
# -
total_variance = ...
print("total_variance: {:.3f} should approximately equal the sum of feature variances: {:.3f}"
.format(total_variance, np.sum(np.var(iris_features, axis=0))))
# + deletable=false editable=false
ok.grade("q2c");
# + [markdown] deletable=false editable=false
# ## Question 3a
#
# Let's now use only the first two principal components to see what a 2D version of our iris data looks like.
#
# First, construct the 2D version of the iris data by matrix-multiplying our `normalized_features` by the first two right singular vectors in `v`. This will project the iris data down from a 4D subspace to a 2D subspace, and the first two right singular vectors are directions for the first two principal components.
#
# **Hints:**
# * To matrix multiply two numpy arrays, use @ or np.dot.
# * The first two right singular vectors in `v` will be the first two columns of `v`, or the first two rows of `vt` (transposed to be column vectors instead of row vectors).
# * Since we want to obtain a 2D version of our iris dataset, the shape of `iris_2d` should be (150, 2).
#
# <!--
# BEGIN QUESTION
# name: q3a
# -->
# -
iris_2d = ...
# + deletable=false editable=false
ok.grade("q3a");
# -
# Now, run the cell below to create the scatter plot of our 2D version of the iris data, `iris_2d`.
plt.figure(figsize=(9, 6))
plt.title("PC2 vs. PC1 for Iris Data")
plt.xlabel("Iris PC1")
plt.ylabel("Iris PC2")
sns.scatterplot(iris_2d[:, 0], iris_2d[:, 1], hue=iris_target);
# + [markdown] deletable=false editable=false
# ## Question 3b
#
# What do you observe about the plot above? If you were given a point in the subspace defined by PC1 and PC2, how well would you be able to classify the point as one of the three iris types?
#
# <!--
# BEGIN QUESTION
# name: q3b
# -->
# -
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# ## Question 3c
#
# What proportion of the total variance is accounted for when we project the iris data down to two dimensions? Compute this quantity in the cell below by dividing the sum of the first two squared singular values (also known as component scores) in `s` by the `total_variance` you calculated previously. Store the result in `two_dim_variance`.
#
# <!--
# BEGIN QUESTION
# name: q3c
# -->
# -
two_dim_variance = ...
two_dim_variance
# + deletable=false editable=false
ok.grade("q3c");
# + [markdown] deletable=false editable=false
# ## Question 4
#
# As a last step, let's create a [scree plot](https://en.wikipedia.org/wiki/Scree_plot) to visualize the weight of each of each principal component. In the cell below, create a scree plot by plotting a line plot of the square of the singular values in `s` vs. the principal component number (1st, 2nd, 3rd, or 4th).
#
# <img src="scree.png" width="400px" />
#
# <!--
# BEGIN QUESTION
# name: q4
# -->
# -
...
# ### You have completed Lab 4!
# + [markdown] deletable=false editable=false
# # Submit
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output.
# **Please save before submitting!**
# + deletable=false editable=false
# Save your notebook first, then run this cell to submit.
ok.submit()
| labs/lab04/lab04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
# # Q1
# +
import xlrd
book = xlrd.open_workbook("house_price.xls")
sheet = book.sheet_by_name('price')
price_list = []
bath_list= []
htype_list= []
lot_size_list = []
year_list = []
area_list = []
for i in range (sheet.nrows):
price,bath,htype,lot_size,year,area = sheet.row_values(i)
if i !=0:
(price,bath,htype,lot_size,year,area)
price_list.append(price)
bath_list.append(bath)
htype_list.append(htype)
lot_size_list.append(lot_size)
year_list.append(year)
area_list.append(area)
print(price_list)
print(bath_list)
print(htype_list)
print(lot_size_list)
print(year_list)
print(area_list)
# -
# # Q2
plt.plot(year_list,price_list)
plt.xlabel('year')
plt.ylabel('price')
plt.show()
# # Q3
plt.hist(price_list)
# # Q4
dot_size = [i**3 for i in bath_list]
print(dot_size)
plt.scatter(area_list,price_list, s = dot_size)
plt.show()
# # Q5
# +
from collections import Counter
result = Counter(htype_list)
print(result)
print(result.keys())
print(result.values())
plt.pie(result.values(),labels = result.keys())
plt.show()
# -
# # Q6
| Lab10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Challenge Notebook
# ## Problem: Create a binary search tree with minimal height from a sorted array.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Is the array in increasing order?
# * Yes
# * Are the array elements unique?
# * Yes
# * Can we assume we already have a Node class with an insert method?
# * Yes
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * 0, 1, 2, 3, 4, 5, 6 -> height 3
# * 0, 1, 2, 3, 4, 5, 6, 7 -> height 4
# ## Algorithm
#
# Refer to the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/graphs_trees/bst_min/bst_min_solution.ipynb). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
# %run ../bst/bst.py
# %load ../bst/bst.py
# +
class Node(object):
def __init__(self, data):
self.data = data
self.left = None
self.right = None
def insert(self, data):
if data <= self.data:
if self.left:
self.left.insert(data)
else:
self.left = Node(data)
else:
if self.right:
self.right.insert(data)
else:
self.right = Node(data)
class MinBst(object):
def create_min_bst(self, array):
if not array:
return None
mid = len(array) // 2
node = Node(array[mid])
node.left = self.create_min_bst(array[:mid])
node.right = self.create_min_bst(array[mid+1:])
return node
# -
# ## Unit Test
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_bst_min.py
from nose.tools import assert_equal
def height(node):
if node is None:
return 0
return 1 + max(height(node.left),
height(node.right))
class TestBstMin(object):
def test_bst_min(self):
min_bst = MinBst()
array = [0, 1, 2, 3, 4, 5, 6]
root = min_bst.create_min_bst(array)
assert_equal(height(root), 3)
min_bst = MinBst()
array = [0, 1, 2, 3, 4, 5, 6, 7]
root = min_bst.create_min_bst(array)
assert_equal(height(root), 4)
print('Success: test_bst_min')
def main():
test = TestBstMin()
test.test_bst_min()
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/graphs_trees/bst_min/bst_min_solution.ipynb) for a discussion on algorithms and code solutions.
| graphs_trees/bst_min/bst_min_challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="fx4faMrck2ri" outputId="71fac306-fd34-4dc7-9109-ca173fc7cce8"
# !pip install kora
import kora.install.rdkit
# + id="vIP9SEWDmOWC"
#importing libraries
import numpy as np
from rdkit import DataStructs
import pandas as pd
# + id="8lCKkWBymwYr"
# auxillary function to create atom pairs
def to_numpyarray_to_list(desc):
arr = np.zeros((1,))
DataStructs.ConvertToNumpyArray(desc, arr)
return arr.tolist()
# + id="RLwbustum3zu"
# function for creating descriptions
#importing libraries
from rdkit.Chem import Descriptors, Lipinski
# + id="wV9hjATNnCQk"
def calc_descriptors(df_molecules, write=False):
# Making a copy of the molecule dataframe
df_mols_desc = df_molecules.copy()
# Create the descriptors (9)
df_mols_desc["molweight"] = df_mols_desc["mols"].apply(Descriptors.ExactMolWt)
df_mols_desc["hatommolwt"] = df_mols_desc["mols"].apply(Descriptors.HeavyAtomMolWt)
df_mols_desc["maxabspartcharge"] = df_mols_desc["mols"].apply(Descriptors.MaxAbsPartialCharge)
df_mols_desc["maxpartcharge"] = df_mols_desc["mols"].apply(Descriptors.MaxPartialCharge)
df_mols_desc["minabspc"] = df_mols_desc["mols"].apply(Descriptors.MinAbsPartialCharge)
df_mols_desc["minpartcharge"] = df_mols_desc["mols"].apply(Descriptors.MinPartialCharge)
df_mols_desc["molwt"] = df_mols_desc["mols"].apply(Descriptors.MolWt)
df_mols_desc["numrade"] = df_mols_desc["mols"].apply(Descriptors.NumRadicalElectrons)
df_mols_desc["numval"] = df_mols_desc["mols"].apply(Descriptors.NumValenceElectrons)
#Lipinski (18)
df_mols_desc["fracsp33"] = df_mols_desc["mols"].apply(Lipinski.FractionCSP3)
df_mols_desc["heavyatomcount"] = df_mols_desc["mols"].apply(Lipinski.HeavyAtomCount)
df_mols_desc["nhohcount"] = df_mols_desc["mols"].apply(Lipinski.NHOHCount)
df_mols_desc["nocount"] = df_mols_desc["mols"].apply(Lipinski.NOCount)
df_mols_desc["aliphcarbocycles"] = df_mols_desc["mols"].apply(Lipinski.NumAliphaticCarbocycles)
df_mols_desc["aliphhetcycles"] = df_mols_desc["mols"].apply(Lipinski.NumAliphaticHeterocycles)
df_mols_desc["aliphrings"] = df_mols_desc["mols"].apply(Lipinski.NumAliphaticRings)
df_mols_desc["arocarbocycles"] = df_mols_desc["mols"].apply(Lipinski.NumAromaticCarbocycles)
df_mols_desc["arohetcycles"] = df_mols_desc["mols"].apply(Lipinski.NumAromaticHeterocycles)
df_mols_desc["arorings"] = df_mols_desc["mols"].apply(Lipinski.NumAromaticRings)
df_mols_desc["numhacceptors"] = df_mols_desc["mols"].apply(Lipinski.NumHAcceptors)
df_mols_desc["numhdonors"] = df_mols_desc["mols"].apply(Lipinski.NumHDonors)
df_mols_desc["numhatoms"] = df_mols_desc["mols"].apply(Lipinski.NumHeteroatoms)
df_mols_desc["numrotbonds"] = df_mols_desc["mols"].apply(Lipinski.NumRotatableBonds)
df_mols_desc["numsatcarbcycles"] = df_mols_desc["mols"].apply(Lipinski.NumSaturatedCarbocycles)
df_mols_desc["numsathetcycles"] = df_mols_desc["mols"].apply(Lipinski.NumSaturatedHeterocycles)
df_mols_desc["numsatrings"] = df_mols_desc["mols"].apply(Lipinski.NumSaturatedRings)
df_mols_desc["ringcount"] = df_mols_desc["mols"].apply(Lipinski.RingCount)
#Drop SMILES and MOLS
df_mols_desc.drop("mols", inplace=True, axis=1)
#Fill NaN with 0
df_mols_desc = df_mols_desc.fillna(0)
df_mols_desc.to_csv("df_mols_desc.csv")
return df_mols_desc
# + id="9Uwc9WI2nQdR"
# creating fingerprints
#importing libraries
from rdkit.Chem import rdMolDescriptors
# + id="_QvLaBB-neWf"
def get_morgan(molecule, length=512):
try:
# radius=2 = ECFP4, radius=3 = ECFP6, etc.
desc = rdMolDescriptors.GetMorganFingerprintAsBitVect(molecule, 2, nBits=length)
except Exception as e:
print(e)
print('error ' + str(molecule))
desc = np.nan
return desc
# + id="XKqv6YAinlKl"
def get_maccs(molecule):
try:
maccs = rdMolDescriptors.GetMACCSKeysFingerprint(molecule)
# Does not have length
except Exception as e:
print(e)
print("error" + str(molecule))
maccs = np.nan
return maccs
# + id="G4obQzR_nn17"
def get_atompairs(molecule, length=512):
try:
atompairs = rdMolDescriptors.GetHashedAtomPairFingerprintAsBitVect(molecule, nBits=length)
except Exception as e:
print(e)
print("error" + str(molecule))
atompairs = np.nan
return atompairs
# + id="XAksRtionrJ7"
def get_topological_torsion(molecule, length=512):
try:
tt = rdMolDescriptors.GetHashedTopologicalTorsionFingerprintAsBitVect(molecule, nBits=length)
except Exception as e:
print(e)
print("error" + str(molecule))
tt = np.nan
return tt
# + id="8EDTO6Cuntqt"
def create_ecfp4_fingerprint(df_molecules, length=512, write=False):
# Morgan Fingerprint (ECFP4)
df_w = df_molecules.copy()
df_w["ECFP4"] = df_w["mols"].apply(lambda x: get_morgan(x, length)).apply(to_numpyarray_to_list)
# New DF with one column for each ECFP bit
ecfp_df = df_w['ECFP4'].apply(pd.Series)
ecfp_df = ecfp_df.rename(columns=lambda x: 'ECFP4_' + str(x + 1))
# Write to csv
ecfp_df.to_csv("ecfp4.csv")
return ecfp_df
# + id="eBtVpIgzn0XG"
def create_maccs_fingerprint(df_molecules, write=False):
# MACCS keys
df_w = df_molecules.copy()
df_w["MACCS"] = df_w["mols"].apply(get_maccs).apply(to_numpyarray_to_list)
# New DF with one column for each MACCS key
maccs_df = df_w['MACCS'].apply(pd.Series)
maccs_df = maccs_df.rename(columns=lambda x: 'MACCS_' + str(x + 1))
# Write to csv
maccs_df.to_csv("maccs.csv")
return maccs_df
# + id="UUS0iKoFn9G5"
def create_atompairs_fingerprint(df_molecules, length=512, write=False):
# ATOM PAIRS
df_w = df_molecules.copy()
df_w["ATOMPAIRS"] = df_w["mols"].apply(lambda x: get_atompairs(x, length)).apply(
to_numpyarray_to_list)
# New DF with one column for each ATOM PAIRS key
atom_pairs_df = df_w['ATOMPAIRS'].apply(pd.Series)
atom_pairs_df = atom_pairs_df.rename(columns=lambda x: 'ATOMPAIR_' + str(x + 1))
# Write to csv
atom_pairs_df.to_csv("atom_pairs.csv")
return atom_pairs_df
# + id="4HC5tBYpoELE"
def create_topological_torsion_fingerprint(df_molecules, length=512, write=False):
# Topological Torsion
df_w = df_molecules.copy()
df_w["TT"] = df_w["mols"].apply(lambda x: get_topological_torsion(x, length)).apply(to_numpyarray_to_list)
# New DF with one column for each Topological torsion key
tt_df = df_w['TT'].apply(pd.Series)
tt_df = tt_df.rename(columns=lambda x: 'TT' + str(x + 1))
# Write to csv
tt_df.to_csv("topological_torsion.csv")
return tt_df
# + id="1MoKRm9goL-L"
pip install matplotlib
# -
pip install tqdm
# plotting libraries
import matplotlib.pyplot as plt
from tqdm import tqdm
# %matplotlib inline
pip install sklearn
pip install xgboost
# + id="cG6RF7_ZoWLT"
# Models
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier, VotingClassifier
import xgboost as xgb
# -
pip install imblearn
pip install requests
# + colab={"base_uri": "https://localhost:8080/"} id="eijx81pjoYyz" outputId="0cf9179e-edd7-4048-b0d8-8e12cde749d0"
# Misc
from rdkit import Chem
from sklearn.model_selection import GridSearchCV, cross_validate, RandomizedSearchCV, StratifiedKFold
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.metrics import classification_report, confusion_matrix, precision_score, recall_score, f1_score, \
roc_auc_score, precision_recall_curve, average_precision_score
from imblearn.pipeline import make_pipeline
from imblearn.over_sampling import SMOTENC
from collections import Counter
import re, requests
# + id="GFwe4TEXofnM"
def create_original_df(usedf=False, file=None, write_s=False, write_off=False):
# Create dataframe from csv
if not usedf:
df = pd.read_csv("sider.csv", skipinitialspace=True)
else:
df = file.copy()
# Extract SMILES column
df_molecules = pd.DataFrame(df["smiles"])
# Converting to molecules
df_molecules["mols"] = df_molecules["smiles"].apply(Chem.MolFromSmiles)
# Droping mols and smiles
df_y = df.drop("smiles", axis=1)
# Write to csv
df_molecules.to_csv("df_molecules.csv")
df_y.to_csv("df_y.csv")
df_molecules.to_csv("df_off_mols.csv")
df_y.to_csv("df_off_y.csv")
return df_y, df_molecules
# + id="EBBR2ljgoz3Z"
def createfingerprints(df_mols, length): # using fingerprints functions
# Morgan Fingerprint (ECFP4)
ecfp_df = create_ecfp4_fingerprint(df_mols, length, False)
# MACCS keys (always 167)
maccs_df = create_maccs_fingerprint(df_mols, False)
# ATOM PAIRS
atom_pairs_df = create_atompairs_fingerprint(df_mols, length, False)
# Topological torsion
tt_df = create_topological_torsion_fingerprint(df_mols, length, False)
return ecfp_df, maccs_df, atom_pairs_df, tt_df
# + id="0egTUHgJpB1U"
def createdescriptors(df_molecules): # using descriptions function
# Descriptors
df_mols_desc = calc_descriptors(df_molecules, False)
return df_mols_desc
# + id="vexZNsX9pJkq"
def test_fingerprint_size(df_mols, df_y, model, colname="Hepatobiliary disorders", num_sizes_to_test=20, min_size=100,
max_size=2048, cv=10, makeplots=False, write=False):
# Fingerprint length type and selection
# Scoring metrics to use
scoring_metrics = ("f1_micro", "f1_macro", "f1", "roc_auc", "recall", "precision", "average_precision")
sizes = np.linspace(min_size, max_size, num_sizes_to_test, dtype=int)
# Create results dataframes for each metric
results_f1 = np.zeros([4, len(sizes)])
results_rocauc = np.zeros([4, len(sizes)])
results_precision = np.zeros([4, len(sizes)])
results_recall = np.zeros([4, len(sizes)])
results_average_precision = np.zeros([4, len(sizes)])
results_f1_micro = np.zeros([4, len(sizes)])
results_f1_macro = np.zeros([4, len(sizes)])
# Get test sizes
c = 0
# Size testing using SVC with scale gamma (1 / (n_features * X.var()))
for s in tqdm(sizes):
# Create fingerprint with size S
fingerprints = createfingerprints(df_mols, int(s))
r = 0
for fp in fingerprints:
X = fp.copy()
# Using "Hepatobiliary disorders" as an results example since its balanced
y = df_y[colname].copy()
# 10-fold cross validation
cv_scores = cross_validate(model, X, y, cv=cv, scoring=scoring_metrics, return_train_score=False, n_jobs=-1)
for k, v in cv_scores.items():
if k == "test_roc_auc":
results_rocauc[r, c] = v.mean()
if k == "test_precision":
results_precision[r, c] = v.mean()
if k == "test_recall":
results_recall[r, c] = v.mean()
if k == "test_average_precision":
results_average_precision[r, c] = v.mean()
if k == "test_f1":
results_f1[r, c] = v.mean()
if k == "test_f1_micro":
results_f1_micro[r, c] = v.mean()
if k == "test_f1_macro":
results_f1_macro[r, c] = v.mean()
r += 1
c += 1
all_results = (results_rocauc, results_precision, results_recall, results_average_precision, results_f1,
results_f1_micro, results_f1_macro)
# Create dataframe for results
df_results_rocauc_size_SVC = pd.DataFrame(results_rocauc, columns=sizes)
df_results_precision_size_SVC = pd.DataFrame(results_precision, columns=sizes)
df_results_recall_size_SVC = pd.DataFrame(results_recall, columns=sizes)
df_results_av_prec_size_SVC = pd.DataFrame(results_average_precision, columns=sizes)
df_results_f1_size_SVC = pd.DataFrame(results_f1, columns=sizes)
df_results_f1_micro_size_SVC = pd.DataFrame(results_f1_micro, columns=sizes)
df_results_f1_macro_size_SVC = pd.DataFrame(results_f1_macro, columns=sizes)
all_df_results = (
df_results_rocauc_size_SVC, df_results_precision_size_SVC, df_results_recall_size_SVC,
df_results_av_prec_size_SVC, df_results_f1_size_SVC, df_results_f1_micro_size_SVC, df_results_f1_macro_size_SVC)
# Save to file
df_results_rocauc_size_SVC.to_csv("df_results_rocauc_size_SVC.csv")
df_results_precision_size_SVC.to_csv("df_results_precision_size_SVC.csv")
df_results_recall_size_SVC.to_csv("df_results_recall_size_SVC.csv")
df_results_av_prec_size_SVC.to_csv("df_results_av_prec_size_SVC.csv")
df_results_f1_size_SVC.to_csv("df_results_f1_size_SVC.csv")
df_results_f1_micro_size_SVC.to_csv("df_results_f1_micro_size_SVC.csv")
df_results_f1_macro_size_SVC.to_csv("df_results_f1_macro_size_SVC.csv")
if makeplots:
fp_names = ["ECFP-4", "MACCS", "Atom Pairs", "Topological Torsion"]
m = 0
for d in all_results:
fig = plt.figure(figsize=(10, 10))
for i in range(len(fingerprints)):
plt.plot(sizes, d[i, :], "-")
plt.title(f"SVC, {scoring_metrics[m]} vs fingerprint length", fontsize=25)
plt.ylabel(f"{scoring_metrics[m]}", fontsize=20)
plt.xlabel("Fingerprint Length", fontsize=20)
plt.legend(fp_names, fontsize=15)
plt.ylim([0, 1])
plt.show()
m += 1
return all_df_results
# + id="48vj-0PGpq88"
def select_best_descriptors_multi(df_desc, y_all, out_names=[], score_func=f_classif, k=1):
# Select k highest scoring feature from X to every y and return new df with only the selected ones
if not out_names:
print("Column names necessary")
return None
selected = []
for n in tqdm(out_names):
skb = SelectKBest(score_func=score_func, k=k).fit(df_desc, y_all[n])
n_sel_bol = skb.get_support()
sel = df_desc.loc[:, n_sel_bol].columns.to_list()
for s in sel:
if s not in selected:
selected.append(s)
return selected
# + id="le7CyKJbpvMI"
def select_best_descriptors(X, y, score_func=f_classif, k=2):
# Select k highest scoring feature from X to y with a score function, f_classif by default
skb = SelectKBest(score_func=score_func, k=k).fit(X, y)
n_sel_bol = skb.get_support()
sel = X.loc[:, n_sel_bol].columns.to_list()
assert sel
return sel
# + id="y0RYrMvupyeF"
def create_dataframes_dic(df_desc_base_train, df_desc_base_test, X_train_fp, X_test_fp, y_train, out_names,
score_func=f_classif, k=3):
# Create 3 dictionaries, one with the train dataframes, one with the test dataframes and one with the selected
# features for each label
# Initialize dictonaries
train_series_dic = {name: None for name in out_names}
test_series_dic = {name: None for name in out_names}
selected_name = {name: None for name in out_names}
# For each of the tasks build the train and test dataframe with the selected descriptors
for name in tqdm(out_names):
# Select best descriptors for the task
sel_col = select_best_descriptors(df_desc_base_train, y_train[name], score_func=score_func, k=k)
selected_name[name] = sel_col # Keep track of selected columns
df_desc_train = df_desc_base_train.loc[:, sel_col].copy() # Get train dataframe with only selected columns
df_desc_test = df_desc_base_test.loc[:, sel_col].copy() # Get test dataframe with only selected columns
X_train = pd.concat([X_train_fp, df_desc_train], axis=1)
X_test = pd.concat([X_test_fp, df_desc_test], axis=1)
# Add to the dictionary
train_series_dic[name] = X_train
test_series_dic[name] = X_test
# Return the dictionaries
return train_series_dic, test_series_dic, selected_name
# + id="WEsue618p65Q"
def balance_dataset(X_train_dic, y_train_dic, out_names, random_state=0, n_jobs=-1, verbose=False):
# Initialize the dictionaries and boolean array for categorical features
train_series_dic_bal = {name: None for name in out_names}
y_dic_bal = {name: None for name in out_names}
cat_shape = np.full((1128,), True, dtype=bool)
cat_shape[-3:] = False
# For each classficiation label
for label in tqdm(out_names):
X_imb = X_train_dic[label]
y_imb = y_train_dic[label]
X_bal, y_bal = SMOTENC(categorical_features=cat_shape, random_state=random_state, n_jobs=n_jobs).fit_resample(
X_imb, y_imb)
train_series_dic_bal[label] = X_bal
y_dic_bal[label] = y_bal
# Print new counts
if verbose:
for label in out_names:
print(f"For {label}")
print(sorted(Counter(y_train_dic[label]).items()))
print(sorted(Counter(y_dic_bal[label]).items()))
# Return the new dictionaries
return train_series_dic_bal, y_dic_bal
# + id="8x63aoRip_6C"
def grid_search(X_train, y_train, model, params_to_test, X_test=None, y_test=None, balancing=False, n_splits=5,
scoring="f1", n_jobs=-1, verbose=False, random_state=None):
# Define grid search
if balancing:
# Save index of categorical features
cat_shape = np.full((1128,), True, dtype=bool)
cat_shape[-3:] = False
# Prepatre SMOTENC
smotenc = SMOTENC(categorical_features=cat_shape, random_state=random_state, n_jobs=n_jobs)
# Make a pipeline with the balancing and the estimator, balacing is only called when fitting
pipeline = make_pipeline(smotenc, model)
# Determine stratified k folds
kf = StratifiedKFold(n_splits=n_splits, random_state=random_state, shuffle=True)
# Call cross validate
grid_search = GridSearchCV(pipeline, params_to_test, cv=kf, n_jobs=n_jobs, verbose=verbose, scoring=scoring)
else:
kf = StratifiedKFold(n_splits=n_splits, random_state=random_state)
grid_search = GridSearchCV(model, params_to_test, cv=kf, n_jobs=n_jobs, verbose=verbose, scoring=scoring)
# Fit X and y to test parameters
grid_search.fit(X_train, y_train)
means = grid_search.cv_results_["mean_test_score"]
stds = grid_search.cv_results_["std_test_score"]
if verbose:
# Print scores
print()
print("Score for development set:")
for mean, std, params in zip(means, stds, grid_search.cv_results_["params"]):
print("%0.3f (+/-%0.03f) for %r" % (mean, std * 1.96, params))
print()
# Print best parameters
print()
print("Best parameters set found:")
print(grid_search.best_params_)
print()
if X_test and y_test:
# Detailed Classification report
print()
print("Detailed classification report:")
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, grid_search.predict(X_test)
print(classification_report(y_true, y_pred))
print()
print("Confusion Matrix as")
print("""
TN FP
FN TP
""")
print(confusion_matrix(y_true, y_pred))
# Save best estimator
best_estimator = grid_search.best_estimator_
best_params = grid_search.best_params_
# And return it
return best_params, best_estimator
# + id="sJuCjUo4qMuF"
def multi_label_grid_search(X_train_dic, y_train, out_names, model, params_to_test, balancing=False, X_test=None,
y_test=None, n_splits=5, scoring="f1", n_jobs=-1, verbose=False, random_state=None):
# Creates a dictionary with the best params in regards to chosen metric for each label
# Creates the dictionary
best_params_by_label = {label: None for label in out_names}
# If X_test and y_test is given so that generalization evalutation can happen
if X_test and y_test:
for label in tqdm(out_names):
print()
print(f"Scores for {label}")
best_params, _ = grid_search(X_train_dic[label], y_train[label], model, params_to_test[label],
X_test[label], y_test[label], n_splits=n_splits, scoring=scoring,
verbose=verbose, n_jobs=n_jobs, balancing=balancing, random_state=random_state)
best_params_by_label[label] = best_params
else:
for label in tqdm(out_names):
print()
print(f"Scores for {label}")
best_params, _ = grid_search(X_train_dic[label], y_train[label], model, params_to_test[label],
n_splits=n_splits, scoring=scoring, verbose=verbose, n_jobs=n_jobs,
balancing=balancing, random_state=random_state)
best_params_by_label[label] = best_params
return best_params_by_label
# + id="mFn8LwCZqRux"
def random_search(X_train, y_train, model, params_to_test, X_test=None, y_test=None, balancing=False,
n_iter=100, n_splits=5, scoring="f1", n_jobs=-1, verbose=False, random_state=None):
# Define random search
if balancing:
# Save index of categorical features
cat_shape = np.full((1128,), True, dtype=bool)
cat_shape[-3:] = False
# Prepatre SMOTENC
smotenc = SMOTENC(categorical_features=cat_shape, random_state=random_state, n_jobs=n_jobs)
# Make a pipeline with the balancing and the estimator, balacing is only called when fitting
pipeline = make_pipeline(smotenc, model)
# Determine stratified k folds
kf = StratifiedKFold(n_splits=n_splits, random_state=random_state, shuffle=True)
# Call cross validate
rs = RandomizedSearchCV(pipeline, params_to_test, n_iter=n_iter, cv=kf, n_jobs=n_jobs, verbose=verbose,
scoring=scoring)
else:
kf = StratifiedKFold(n_splits=n_splits, random_state=random_state, shuffle=True)
rs = RandomizedSearchCV(model, params_to_test, n_iter=n_iter, cv=kf, n_jobs=n_jobs, verbose=verbose,
scoring=scoring)
# Fit parameters
rs.fit(np.asarray(X_train), np.asarray(y_train))
means = rs.cv_results_["mean_test_score"]
stds = rs.cv_results_["std_test_score"]
# Print scores
if verbose:
print()
print("Score for development set:")
for mean, std, params in zip(means, stds, rs.cv_results_["params"]):
print("%0.3f (+/-%0.03f) for %r" % (mean, std * 1.96, params))
print()
# Print best parameters
print()
print("Best parameters set found:")
print(rs.best_params_)
print()
if X_test and y_test:
# Detailed Classification report
print()
print("Detailed classification report:")
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, rs.predict(X_test)
print(classification_report(y_true, y_pred))
print()
"""
print("Confusion matrix as:")
print(
TN FP
FN TP
)
print(confusion_matrix(y_true, y_pred))
print()
"""
# Save best estimator
best_estimator = rs.best_estimator_
best_params = rs.best_params_
# And return it
return best_params, best_estimator
# + id="HPAEHy5XqcvH"
def multi_label_random_search(X_train_dic, y_train, out_names, model, params_to_test, balancing=False, X_test=None,
y_test=None, n_iter=100, n_splits=5, scoring="f1", n_jobs=-1, verbose=False,
random_state=None):
# Creates a dictionary with the best params in regards to chosen metric for each label
# Creates the dictionary
best_params_by_label = {label: None for label in out_names}
# If X_test and y_test is given so that generalization evalutation can happen
if X_test and y_test:
for label in tqdm(out_names):
print()
print(f"Scores for {label}")
best_params, _ = random_search(X_train_dic[label], y_train[label], model, params_to_test[label],
X_test[label], y_test[label], n_iter=n_iter, n_splits=n_splits,
scoring=scoring, verbose=verbose, n_jobs=n_jobs, random_state=random_state,
balancing=balancing)
best_params_by_label[label] = best_params
else:
for label in tqdm(out_names):
print()
print(f"Scores for {label}")
best_params, _ = random_search(X_train_dic[label], y_train[label], model, params_to_test[label],
n_iter=n_iter, n_splits=n_splits, scoring=scoring, verbose=verbose,
n_jobs=n_jobs, random_state=random_state, balancing=balancing)
best_params_by_label[label] = best_params
return best_params_by_label
# + id="-pn_8ENvqgrC"
def score_report(estimator, X_test, y_test, verbose=False, plot=False, name=None):
# Predicting value
y_true, y_pred = y_test, estimator.predict(X_test)
y_score = estimator.predict_proba(X_test)
y_score = y_score[:, 1]
# Individual metrics
f1_micr_score = f1_score(y_true, y_pred, average="micro")
f1_macro_score = f1_score(y_true, y_pred, average="macro")
f1_s_score = f1_score(y_true, y_pred, average="binary")
auc = roc_auc_score(y_true, y_pred)
rec = recall_score(y_true, y_pred, average="binary")
prec = precision_score(y_true, y_pred, average="binary")
average_precision = average_precision_score(y_true, y_score)
# Detailed Classification report
if verbose:
print()
print("The scores are computed on the full evaluation set")
print("These are not used to train or optimize the model")
print()
print("Detailed classification report:")
print(classification_report(y_true, y_pred))
print()
print("Confusion matrix as:")
print("""
TN FP
FN TP
""")
print(confusion_matrix(y_true, y_pred))
print()
print("Individual metrics:")
print(f"F1 Micro score: {f1_micr_score:.3f}")
print(f"F1 Macro score: {f1_macro_score:.3f}")
print(f"F1 Binary score: {f1_s_score:.3f}")
print(f"AUROC score: {auc:.3f}")
print(f"Recall score: {rec:.3f}")
print(f"Precision score: {prec:.3f}")
print(f"Average precision-recall score: {average_precision:.3f}")
print()
if plot:
precision, recall, _ = precision_recall_curve(y_true, y_score)
# step_kwargs = ({'step': 'post'}
# if 'step' in signature(plt.fill_between).parameters
# else {})
plt.step(recall, precision, color="r", alpha=0.2, where="post")
plt.fill_between(recall, precision, step="post", alpha=0.2, color="#F59B00")
plt.xlabel("Recall")
plt.ylabel("Precision")
plt.ylim([0.0, 1.05])
plt.xlim([0.0, 1.0])
plt.title(f'{name} \n Precision-Recall curve: AP={average_precision:0.2f}')
plt.savefig(f"Precision-Recall curve.png")
plt.clf()
return {"f1_micr_score": f1_micr_score, "auc_score": auc, "rec_score": rec, "prec_score": prec,
"f1_macro_score": f1_macro_score, "f1_s_score": f1_s_score, "prec_rec_score": average_precision}
# + id="rjk_-s_Yqng7"
def cv_report(estimator, X_train, y_train, balancing=False, n_splits=5,
scoring_metrics=("f1_micro", "f1_macro", "f1", "roc_auc", "recall", "precision", "average_precision"),
random_state=None, n_jobs=-1, verbose=False):
if balancing:
# Save index of categorical features
cat_shape = np.full((1128,), True, dtype=bool)
cat_shape[-3:] = False
# Prepare SMOTENC
smotenc = SMOTENC(categorical_features=cat_shape, random_state=random_state, n_jobs=n_jobs)
# Make a pipeline with the balancing and the estimator, balacing is only called when fitting
pipeline = make_pipeline(smotenc, estimator)
# Determine stratified k folds
kf = StratifiedKFold(n_splits=n_splits, random_state=random_state, shuffle=True)
# Call cross validate
scores = cross_validate(pipeline, np.asarray(X_train), np.asarray(y_train), scoring=scoring_metrics, cv=kf,
n_jobs=n_jobs, verbose=verbose, return_train_score=False)
else:
# Normal cross validation
kf = StratifiedKFold(n_splits=n_splits, random_state=random_state,shuffle=True)
scores = cross_validate(estimator, np.asarray(X_train), np.asarray(y_train), scoring=scoring_metrics, cv=kf,
n_jobs=n_jobs, verbose=verbose, return_train_score=False)
# Means
f1_s = np.mean(scores["test_f1_micro"])
f1_ms = np.mean(scores["test_f1_macro"])
f1_bs = np.mean(scores["test_f1"])
auc_s = np.mean(scores["test_roc_auc"])
rec_s = np.mean(scores["test_recall"])
prec_s = np.mean(scores["test_precision"])
avp_s = np.mean(scores["test_average_precision"])
# STD
f1_std = np.std(scores["test_f1_micro"])
f1_mstd = np.std(scores["test_f1_macro"])
f1_bstd = np.std(scores["test_f1"])
auc_std = np.std(scores["test_roc_auc"])
rec_std = np.std(scores["test_recall"])
prec_std = np.std(scores["test_precision"])
avp_std = np.std(scores["test_average_precision"])
if verbose:
print()
print("Individual metrics")
print(f"F1 Micro Score: Mean: {f1_s:.3f} (Std: {f1_std:.3f})")
print(f"F1 Macro Score: Mean: {f1_ms:.3f} (Std: {f1_mstd:.3f})")
print(f"F1 Binary Score: Mean: {f1_bs:.3f} (Std: {f1_bstd:.3f})")
print(f"AUROC score: Mean: {auc_s:.3f} (Std: {auc_std:.3f})")
print(f"Recall score: Mean: {rec_s:.3f} (Std: {rec_std:.3f})")
print(f"Precision score: Mean: {prec_s:.3f} (Std: {prec_std:.3f})")
print(f"Average Precision score: Mean: {avp_s:.3f} (Std: {avp_std:.3f})")
print()
return {"f1_micr_score": f1_s, "f1_micr_std": f1_std, "auc_score": auc_s, "auc_std": auc_std, "rec_score": rec_s,
"rec_std": rec_std, "prec_score": prec_s, "prec_std": prec_std, "f1_macro_score": f1_ms,
"f1_macro_std": f1_mstd, "f1_score": f1_bs, "f1_std": f1_bstd, "avp_score": avp_s, "avp_std": avp_std}
# + id="G2f47LLUqxOb"
def cv_multi_report(X_train_dic, y_train, out_names, model=None, balancing=False, modelname=None, spec_params=None,
random_state=None, n_splits=5, n_jobs=-1, verbose=False):
# Creates a scores report dataframe for each classification label with cv
# Initizalize the dataframe
report = pd.DataFrame(
columns=["F1 Binary", "F1 Micro", "F1 Macro", "ROC_AUC", "Recall", "Precision", "Average Precision"],
index=out_names)
scoring_metrics = ("f1_micro", "f1_macro", "f1", "roc_auc", "recall", "precision", "average_precision")
# For each label
for name in tqdm(out_names):
if verbose:
print()
print(f"Scores for {name}")
# Calculate the score for the current label using the respective dataframe
if spec_params:
# Define the specific parameters for each model for each label
if modelname[name] == "SVC":
model_temp = SVC(random_state=random_state, probability=True)
model_temp.set_params(C=spec_params[name]["svc__C"],
gamma=spec_params[name]["svc__gamma"],
kernel=spec_params[name]["svc__kernel"])
elif modelname[name] == "RF":
model_temp = RandomForestClassifier(n_estimators=100, random_state=random_state)
model_temp.set_params(bootstrap=spec_params[name]["randomforestclassifier__bootstrap"],
max_depth=spec_params[name]["randomforestclassifier__max_depth"],
max_features=spec_params[name]["randomforestclassifier__max_features"],
min_samples_leaf=spec_params[name]["randomforestclassifier__min_samples_leaf"],
min_samples_split=spec_params[name]["randomforestclassifier__min_samples_split"],
n_estimators=spec_params[name]["randomforestclassifier__n_estimators"])
elif modelname[name] == "XGB":
model_temp = xgb.XGBClassifier(objective="binary:logistic", random_state=random_state)
model_temp.set_params(colsample_bytree=spec_params[name]["xgbclassifier__colsample_bytree"],
eta=spec_params[name]["xgbclassifier__eta"],
gamma=spec_params[name]["xgbclassifier__gamma"],
max_depth=spec_params[name]["xgbclassifier__max_depth"],
min_child_weight=spec_params[name]["xgbclassifier__min_child_weight"],
subsample=spec_params[name]["xgbclassifier__subsample"])
elif modelname[name] == "VotingClassifier":
# Spec params must be the list of the dictionaries with the params in order (SVC - RF - XGB)
model_svc = SVC(random_state=random_state, probability=True)
model_rf = RandomForestClassifier(n_estimators=100, random_state=random_state)
model_xgb = xgb.XGBClassifier(objective="binary:logistic", random_state=random_state)
model_svc.set_params(C=spec_params[0][name]["svc__C"],
gamma=spec_params[0][name]["svc__gamma"],
kernel=spec_params[0][name]["svc__kernel"])
model_rf.set_params(bootstrap=spec_params[1][name]["randomforestclassifier__bootstrap"],
max_depth=spec_params[1][name]["randomforestclassifier__max_depth"],
max_features=spec_params[1][name]["randomforestclassifier__max_features"],
min_samples_leaf=spec_params[1][name]["randomforestclassifier__min_samples_leaf"],
min_samples_split=spec_params[1][name]["randomforestclassifier__min_samples_split"],
n_estimators=spec_params[1][name]["randomforestclassifier__n_estimators"])
model_xgb.set_params(colsample_bytree=spec_params[2][name]["xgbclassifier__colsample_bytree"],
eta=spec_params[2][name]["xgbclassifier__eta"],
gamma=spec_params[2][name]["xgbclassifier__gamma"],
max_depth=spec_params[2][name]["xgbclassifier__max_depth"],
min_child_weight=spec_params[2][name]["xgbclassifier__min_child_weight"],
subsample=spec_params[2][name]["xgbclassifier__subsample"])
model_temp = VotingClassifier(estimators=[("svc", model_svc), ("rf", model_rf), ("xgb", model_xgb)],
voting="soft", n_jobs=n_jobs)
else:
print("Please specify used model (SVC, RF, XGB)")
return None
scores = cv_report(model_temp, X_train_dic[name], y_train[name], balancing=balancing, n_splits=n_splits,
scoring_metrics=scoring_metrics, n_jobs=n_jobs, verbose=verbose,
random_state=random_state)
else:
scores = cv_report(model, X_train_dic[name], y_train[name], balancing=balancing, n_splits=n_splits,
scoring_metrics=scoring_metrics, n_jobs=n_jobs, verbose=verbose,
random_state=random_state)
report.loc[name, "F1 Micro"] = round(float(scores["f1_micr_score"]), 3)
report.loc[name, "F1 Macro"] = round(float(scores["f1_macro_score"]), 3)
report.loc[name, "F1 Binary"] = round(float(scores["f1_score"]), 3)
report.loc[name, "ROC_AUC"] = round(float(scores["auc_score"]), 3)
report.loc[name, "Recall"] = round(float(scores["rec_score"]), 3)
report.loc[name, "Precision"] = round(float(scores["prec_score"]), 3)
report.loc[name, "Average Precision"] = round(float(scores["avp_score"]), 3)
report = report.apply(pd.to_numeric)
return report
# + id="w9dUsDs2rAr2"
def test_score_multi_report(X_train_dic, y_train, X_test, y_test, out_names, model=None, modelname=None,
spec_params=None, balancing=False, random_state=None, plot=False, verbose=False, n_jobs=-1):
# Creates a scores report dataframe for each classification label with cv
# Initizalize the dataframe
report = pd.DataFrame(columns=["F1 Binary", "F1 Micro", "F1 Macro", "ROC_AUC", "Recall", "Precision"],
index=out_names)
# For each label
for name in tqdm(out_names):
if verbose:
print()
print(f"Scores for {name}")
# Calculate the score for the current label using the respective dataframe
if spec_params:
# Define the specific parameters for each model for each label
if modelname[name] == "SVC":
model_temp = SVC(random_state=random_state, probability=True)
model_temp.set_params(C=spec_params[name]["svc__C"],
gamma=spec_params[name]["svc__gamma"],
kernel=spec_params[name]["svc__kernel"])
elif modelname[name] == "RF":
model_temp = RandomForestClassifier(n_estimators=100, random_state=random_state)
model_temp.set_params(bootstrap=spec_params[name]["randomforestclassifier__bootstrap"],
max_depth=spec_params[name]["randomforestclassifier__max_depth"],
max_features=spec_params[name]["randomforestclassifier__max_features"],
min_samples_leaf=spec_params[name]["randomforestclassifier__min_samples_leaf"],
min_samples_split=spec_params[name]["randomforestclassifier__min_samples_split"],
n_estimators=spec_params[name]["randomforestclassifier__n_estimators"])
elif modelname[name] == "XGB":
model_temp = xgb.XGBClassifier(objective="binary:logistic", random_state=random_state)
model_temp.set_params(colsample_bytree=spec_params[name]["xgbclassifier__colsample_bytree"],
eta=spec_params[name]["xgbclassifier__eta"],
gamma=spec_params[name]["xgbclassifier__gamma"],
max_depth=spec_params[name]["xgbclassifier__max_depth"],
min_child_weight=spec_params[name]["xgbclassifier__min_child_weight"],
subsample=spec_params[name]["xgbclassifier__subsample"])
else:
print("Please specify used model (SVC, RF, XGB)")
return None
if balancing:
# Save index of categorical features
cat_shape = np.full((1128,), True, dtype=bool)
cat_shape[-3:] = False
# Prepatre SMOTENC
smotenc = SMOTENC(categorical_features=cat_shape, random_state=random_state, n_jobs=n_jobs)
# Make a pipeline with the balancing and the estimator, balacing is only called when fitting
pipeline = make_pipeline(smotenc, model_temp)
# Fit and test
pipeline.fit(np.asarray(X_train_dic[name]), np.asarray(y_train[name]))
scores = score_report(pipeline, np.asarray(X_test[name]), np.asarray(y_test[name]), plot=plot,
verbose=verbose, name=name)
else:
model_temp.fit(np.asarray(X_train_dic[name]), np.asarray(y_train[name]))
scores = score_report(model_temp, np.asarray(X_test[name]), np.asarray(y_test[name]), plot=plot,
verbose=verbose, name=name)
else:
if balancing:
# Save index of categorical features
cat_shape = np.full((1128,), True, dtype=bool)
cat_shape[-3:] = False
# Prepatre SMOTENC
smotenc = SMOTENC(categorical_features=cat_shape, random_state=random_state, n_jobs=n_jobs)
# Make a pipeline with the balancing and the estimator, balacing is only called when fitting
pipeline = make_pipeline(smotenc, model)
# Fit and test
pipeline.fit(np.asarray(X_train_dic[name]), np.asarray(y_train[name]))
scores = score_report(pipeline, np.asarray(X_test[name]), np.asarray(y_test[name]), plot=plot,
verbose=verbose, name=name)
else:
model.fit(np.asarray(X_train_dic[name]), np.asarray(y_train[name]))
scores = score_report(model, np.asarray(X_test[name]), np.asarray(y_test[name]), plot=plot,
verbose=verbose, name=name)
report.loc[name, "F1 Micro"] = round(float(scores["f1_micr_score"]), 3)
report.loc[name, "F1 Macro"] = round(float(scores["f1_macro_score"]), 3)
report.loc[name, "F1 Binary"] = round(float(scores["f1_s_score"]), 3)
report.loc[name, "ROC_AUC"] = round(float(scores["auc_score"]), 3)
report.loc[name, "Recall"] = round(float(scores["rec_score"]), 3)
report.loc[name, "Precision"] = round(float(scores["prec_score"]), 3)
report.loc[name, "Average Prec-Rec"] = round(float(scores["prec_rec_score"]), 3)
# prec_rec_score
report = report.apply(pd.to_numeric)
return report
# + id="VWlFDqWMrNOx"
def get_smile_from_cid(cid):
# Trim CID
ct = re.sub("^CID[0]*", "", cid)
# Getting smile
res = requests.get(f"https://pubchem.ncbi.nlm.nih.gov/rest/pug/compound/cid/{ct}/property/CanonicalSMILES/txt")
# Checking for Error 400
try:
res.raise_for_status()
except Exception as e:
print(f"Problem retrieving smile for {cid}: {e}")
# If everything is ok, get smile text
res_t = res.text.strip("\n")
# Return smile
return res_t
# + id="Zk1i-UUkrR5O"
def create_offside_df(out_names, write=False):
oss = pd.read_csv("offsides_socs.csv")
oss_df = oss[["stitch_id", "SOC"]].copy()
stitchs = oss_df.stitch_id.unique()
sti_to_smil = {stitch: get_smile_from_cid(stitch) for stitch in tqdm(stitchs)}
d = {"stitch_id": stitchs}
mod_off = pd.DataFrame(data=d)
mod_off["smiles"] = mod_off.stitch_id.apply(lambda x: sti_to_smil[x])
for name in out_names:
mod_off[name] = 0
for index, row in tqdm(oss_df.iterrows()):
if row["SOC"] in out_names:
mod_off.loc[mod_off["stitch_id"] == row["stitch_id"], row["SOC"]] = 1
mod_off.drop("stitch_id", inplace=True, axis=1)
mod_off.to_csv("offside_socs_modified.csv", index=False)
return mod_off
# +
# Importing libraries for modelling
from sklearn.model_selection import train_test_split
from pprint import pprint
import matplotlib.pyplot as plt
from IPython.display import Image
# %matplotlib inline
# Functions
from params_by_label import *
# Fixing the seed
seed = 6
np.random.seed(seed)
# +
# model development
# Creating base df_molecules, df_y with the results vectors, and df_mols_descr with the descriptors
print("Creating Dataframes")
y_all, df_molecules = create_original_df(write_s=False)
df_molecules.drop("smiles", axis=1, inplace=True)
todrop = ["Product issues", "Investigations", "Social circumstances"]
y_all.drop(todrop, axis=1, inplace=True) # No real connection with the molecule, multiple problems
out_names = y_all.columns.tolist() # Get class labels
# Separating in a DF_mols_train and an Df_mols_test, in order to avoid data snooping and fitting the model to the test
df_mols_train, df_mols_test, y_train, y_test = train_test_split(df_molecules, y_all, test_size=0.2, random_state=seed)
# -
d = {"Positives": y_all.sum(axis=0), "Negatives": 1427 - y_all.sum(axis=0)}
countsm = pd.DataFrame(data=d)
countsm.plot(kind='bar', figsize=(16, 10), title="Adverse Drug Reactions Counts", ylim=(0, 1500), stacked=True)
# # FEATURE GENERATION AND SELECTION
# +
# fingerprint length
all_df_results_svc = test_fingerprint_size(df_mols_train, y_train, SVC(gamma="scale", random_state=seed), makeplots=True, write=False)
# Best result with ECFP-4 at 1125 - This will be used to all results
# +
# Create X datasets with fingerprint length
X_all, _, _, _ = createfingerprints(df_molecules, length=1125)
X_train_fp, _, _, _ = createfingerprints(df_mols_train, length=1125)
X_test_fp, _, _, _ = createfingerprints(df_mols_test, length=1125)
# Selects and create descriptors dataset
df_desc = createdescriptors(df_molecules) # Create all descriptors
# Splits in train and test
df_desc_base_train, df_desc_base_test = train_test_split(df_desc, test_size=0.2, random_state=seed)
# Creates a dictionary with key = class label and value = dataframe with fingerprint + best K descriptors for that label
X_train_dic, X_test_dic, selected_cols = create_dataframes_dic(df_desc_base_train, df_desc_base_test, X_train_fp,
X_test_fp, y_train, out_names, score_func=f_classif, k=3)
# Creates a y dictionary for all labels
y_train_dic = {name: y_train[name] for name in out_names}
modelnamesvc = {name: "SVC" for name in out_names}
modelnamerf = {name: "RF" for name in out_names}
modelnamexgb = {name: "XGB" for name in out_names}
modelnamevot = {name: "VotingClassifier" for name in out_names}
# -
print("Selected descriptors by label:")
pprint(selected_cols, width=-1)
# The base RF model performed not only better than the base SVC model for most of the classification tasks, it also out-performed the optimized SVC model
# # MODELING
# SVC MODEL DEVELOPMENT
print("SVC")
print("Base SVC without balancing:")
base_svc_report = cv_multi_report(X_train_dic, y_train, out_names, SVC(gamma="auto", random_state=seed), n_splits=5,
n_jobs=-2, verbose=False)
print("Scores for SVC without balancing:")
base_svc_report
| Descriptions_Fingerprints_Processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9v1fFy6VT1e9" colab_type="code" outputId="57e3099c-92d0-4534-ce50-65976588524c" colab={"base_uri": "https://localhost:8080/", "height": 132}
from google.colab import files, drive
drive.mount('/content/gdrive')
# + id="dJoQf-I2n1H5" colab_type="code" outputId="bcd2af38-7780-44a0-dca6-718f28c2f1df" colab={"base_uri": "https://localhost:8080/", "height": 112}
# !ln -s gdrive/My\ Drive/donkey donkey
# !ls -l donkey/
# + id="rw73EI5Tu4Ze" colab_type="code" cellView="both" colab={}
# Importy Pythonowe
import json
import os
import zipfile
import glob
import matplotlib.animation
import matplotlib.pyplot as plt
import numpy as np
from scipy import ndimage
from tqdm import tqdm
import seaborn as sns
sns.set_style('whitegrid')
import keras as K
import keras.layers as KL
import IPython.display
from IPython.display import HTML
# + id="Cr2PzakVZidX" colab_type="code" outputId="24d7b21d-533f-45f5-f112-681b434cfa94" colab={"base_uri": "https://localhost:8080/", "height": 340}
# Czytanie danych do pamięci
records = []
for archive_f in ['donkey/record2.zip', 'donkey/record3.zip']:
with zipfile.ZipFile(archive_f) as archive:
for fileinfo in tqdm(archive.filelist):
filename = fileinfo.filename
if not filename.endswith('.json') or filename.endswith('meta.json'):
continue
with archive.open(filename) as f:
data = json.load(f)
basename = os.path.basename(filename)
dirname = os.path.dirname(filename)
# ucinamy 'record_' z przodu i '.json' z tylu
step_number = int(basename[7:-5])
with archive.open(
os.path.join(dirname, data['image_array'])) as image_file:
image = ndimage.imread(image_file) / 255.0
records.append((step_number, image, data['user_angle'], data['user_throttle']))
records.sort(key=lambda x: x[0])
images = np.array([r[1] for r in records], dtype='float32')
angles = np.array([r[2] for r in records], dtype='float32')[:, None]
throttles = np.array([r[3] for r in records], dtype='float32')[:, None]
# + id="0ePbexK90mdB" colab_type="code" colab={}
images = images[:,72:,:,:]
# + id="Np4FzXl9xqNk" colab_type="code" outputId="d11f9925-c9a7-41a5-e9fa-f8c13ef6c328" colab={"base_uri": "https://localhost:8080/", "height": 93}
print('Images to tablica o wymiarach:', images.shape)
print('Angles to tablica o wymiarach:', angles.shape)
print('Throttles to tablica o wymiarach:', throttles.shape)
# Dane (images, angles) podzielimy na 3 podzbiory:
# - dane uczące (train_images, train_angles), na których będziemy trenować modele
# - dane walidacyjne (valid_images, valid_angles) również użyte do treningu modeli
# - dane testowe (test_images, test_angles) na których ocenimy jak dobrze działa
# nasz model
train_images, valid_images, test_images = np.split(images, [-1000, -500])
train_angles, valid_angles, test_angles = np.split(angles, [-1000, -500])
(train_throttles, valid_throttles, test_throttles
) = np.split(throttles, [-1000, -500])
print('Dane uczące mają %d klatek, walidacyjne %d i testowe %d.' %
(train_images.shape[0], valid_images.shape[0], test_images.shape[0]))
# + id="u7-uaJrpkXr8" colab_type="code" outputId="ec71c696-f489-484b-fab7-a6d3531fb31a" colab={"base_uri": "https://localhost:8080/", "height": 329}
# Wizualizacja danych
k = 5
plt.figure(figsize=(15, 5))
for i, frame in enumerate(range(k*9*9, (k+1)*9*9, 9)):
plt.subplot(3, 3, i + 1)
plt.imshow(images[frame])
plt.title("Klatka %d" % (frame,))
# + id="bV9Kj0uCl7wg" colab_type="code" outputId="87ebef19-4bd1-4502-c0a0-b9082286d694" colab={"base_uri": "https://localhost:8080/", "height": 376}
plt.plot(angles, color='g', label='recorded steering')
plt.plot(throttles, color='b', label='recorded steering')
plt.title('Nagrane sterowanie')
plt.xlabel('Nr klatki')
plt.ylabel('Skręt kierownicy')
None
# + id="3-HN9MEeA4j2" colab_type="code" colab={}
img_in = KL.Input(shape=(6, 48, 160, 3), name='img_in')
x = img_in
x = KL.BatchNormalization()(x)
x = KL.Convolution3D(filters=24, kernel_size=(1, 5, 5), padding='same', activation='relu')(x)
x = KL.Convolution3D(filters=32, kernel_size=(1, 5, 5), padding='same', activation='relu')(x)
x = KL.MaxPooling3D(pool_size=(1, 1, 2))(x)
x = KL.BatchNormalization()(x)
x = KL.Convolution3D(filters=64, kernel_size=(2, 5, 5), activation='relu')(x)
x = KL.Convolution3D(filters=64, kernel_size=(2, 3, 3), activation='relu')(x)
x = KL.MaxPooling3D(pool_size=(1, 2, 2))(x)
x = KL.BatchNormalization()(x)
x = KL.Convolution3D(filters=64, kernel_size=(2, 3, 3), activation='relu')(x)
x = KL.Convolution3D(filters=32, kernel_size=(2, 3, 3), activation='relu')(x)
x = KL.MaxPooling3D(pool_size=(1, 2, 2))(x)
x = KL.BatchNormalization()(x)
x = KL.Flatten(name='flattened')(x)
x = KL.Dense(units=64, activation='linear')(x)
x = KL.BatchNormalization()(x)
# categorical output of the angle
angle_out = KL.Dense(units=1, activation='linear', name='angle_out')(x)
# continous output of throttle
throttle_out = KL.Dense(units=1, activation='linear', name='throttle_out')(x)
model = K.Model(inputs=[img_in], outputs=[angle_out, throttle_out])
model.compile(optimizer='adam',
loss={'angle_out': 'mean_squared_error',
'throttle_out': 'mean_squared_error'},
loss_weights={'angle_out': 0.5, 'throttle_out': 0.5})
# + id="W26zQodlsP4I" colab_type="code" colab={}
def data_generator(X, Y, batch_size, shuffle=True):
A, T = Y
back_ind = np.array([1, 2, 4, 8, 16, 32]).reshape(1, -1)
while True:
n = X.shape[0]
indices = np.arange(n)
if shuffle:
np.random.shuffle(indices)
for i in range(0, n, batch_size):
batch_ind = indices[i:i+batch_size]
batch_back_ind = np.maximum(0, batch_ind.reshape(-1, 1) - back_ind)
yield (X[batch_back_ind], [A[batch_ind], T[batch_ind]])
# + id="lZINkPNnZI8E" colab_type="code" outputId="6ae9a8bf-4eaa-468f-f928-64d9d537914a" colab={"base_uri": "https://localhost:8080/", "height": 189}
callbacks = [
K.callbacks.ModelCheckpoint('model', save_best_only=True),
K.callbacks.EarlyStopping(monitor='val_loss',
min_delta=.0005,
patience=5,
verbose=True,
mode='auto')
]
batch_size=16
# Model uczymy na danych uczących.
# Po każdej epoce (ang. epoch) policzymy błąd na danych walidacyjnych i jeśli
# model jest lepszy (błąd jest mniejszy), zapisujemy go.
hist = model.fit_generator(data_generator(train_images, (train_angles, train_throttles), batch_size),
steps_per_epoch=len(train_images)/batch_size,
epochs=200,
validation_data=data_generator(valid_images, (valid_angles, valid_throttles), batch_size),
validation_steps=len(valid_images)/batch_size,
callbacks=callbacks)
# + [markdown] id="2d8CH-8x4X-i" colab_type="text"
# ## Analiza wyuczonej sieci
# Najpierw sprawdzimy błąd MSE osiągnięty przez sieć.
# + id="avL4PSMZ4amr" colab_type="code" outputId="c45ff06b-e382-465b-bd7b-8b9ee7957d37" colab={"base_uri": "https://localhost:8080/", "height": 55}
#@title Wczytywanie najlepszej sieci
best_model = K.models.load_model('model')
print('Obliczony przez Keras błąd walidacyjny:',
best_model.evaluate_generator(data_generator(valid_images,
(valid_angles, valid_throttles), batch_size),
steps=len(valid_images)/batch_size, verbose=0))
print('Obliczony przez Keras błąd testowy:',
best_model.evaluate_generator(data_generator(test_images,
(test_angles, test_throttles), batch_size),
steps=len(test_images)/batch_size, verbose=0))
# + [markdown] id="vqNq4usZ8anI" colab_type="text"
# Teraz zobaczymy jak można wykorzystać sieć do przewidywania sterowania na danych testowych:
# + id="jeF1bHEq4suo" colab_type="code" colab={}
#@title Użycie najlepszej sieci
# Obliczamy kąty przewidziane przez sieć na danych testowych
pred_test_angles, pred_test_throttles = best_model.predict_generator(data_generator(test_images,
(test_angles, test_throttles), 1, shuffle=False), steps=len(test_images))
# + id="MA6zDza4F-yN" colab_type="code" outputId="dccea693-5fa6-4d87-b773-109f271a34e2" colab={"base_uri": "https://localhost:8080/", "height": 362}
plt.plot(test_angles, label='recorded', color='g', alpha=0.5)
plt.plot(pred_test_angles, label='predicted', color='r')
plt.legend(loc='upper right')
plt.title('Nagrane i przewidywane sterowanie samochodem.')
None
# + id="ZmT_zIuyuh6k" colab_type="code" colab={}
files.download('model')
# + id="X0lbjl_7gJl7" colab_type="code" colab={}
| notebooks/train_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:analysis]
# language: python
# name: conda-env-analysis-py
# ---
# # ESMLab Scaling Study
# +
# %matplotlib inline
import math, time
from dask.distributed import Client, wait
import ncar_jobqueue
import dask
import xarray as xr
import intake
import numpy as np
import matplotlib.pyplot as plt
# -
cluster = ncar_jobqueue.NCARCluster(walltime="00:30:00", cores=36, memory='109GB', processes=2, project='NTDD0005')
client = dask.distributed.Client(cluster)
n_workers = 2 * 6
cluster.scale(n_workers)
client
col = intake.open_esm_metadatastore(collection_input_definition='cesm1-le-collection.yml',
overwrite_existing=False)
col.df.info()
arbitrary_pop_file = col.search(experiment='20C', stream='pop.h').query_results.file_fullpath.tolist()[0]
ds = xr.open_dataset(arbitrary_pop_file, decode_times=False, decode_coords=False)
grid_vars = ['KMT', 'z_t', 'TAREA', 'dz']
ds = ds.drop([v for v in ds.variables if v not in grid_vars]).compute()
ds
# +
nk = len(ds.z_t)
nj = ds.KMT.shape[0]
ni = ds.KMT.shape[1]
# make 3D array of 0:km
k_vector_one_to_km = xr.DataArray(np.arange(0, nk), dims=('z_t'), coords={'z_t': ds.z_t})
ONES_3d = xr.DataArray(np.ones((nk, nj, ni)), dims=('z_t', 'nlat', 'nlon'), coords={'z_t': ds.z_t})
MASK = (k_vector_one_to_km * ONES_3d)
# mask out cells where k is below KMT
MASK = MASK.where(MASK <= ds.KMT - 1)
MASK = xr.where(MASK.notnull(), 1., 0.)
plt.figure()
MASK.isel(z_t=0).plot()
plt.title('Surface mask')
plt.figure()
MASK.isel(nlon=200).plot(yincrease=False)
plt.title('Pacific transect')
# +
MASKED_VOL = ds.dz * ds.TAREA * MASK
MASKED_VOL.attrs['units'] = 'cm^3'
MASKED_VOL.attrs['long_name'] = 'masked volume'
plt.figure()
MASKED_VOL.isel(z_t=0).plot()
plt.title('Surface mask')
plt.figure()
MASKED_VOL.isel(nlon=200).plot(yincrease=False)
plt.title('Pacific transect')
# -
member_ids = col.search(experiment=['20C', 'RCP85'], has_ocean_bgc=True).query_results.ensemble.unique().tolist()
print(member_ids)
variable = ['O2']
query = dict(ensemble=member_ids[:2], experiment=['20C', 'RCP85'],
stream='pop.h', variable=variable, direct_access=True)
col_subset = col.search(**query)
col_subset.query_results.info()
# +
import socket
import re
def get_machine():
"""Function to determine which base class to use.
"""
filter1 = r'^cheyenne'
filter2 = r'r\d(i)\d(n)\d*'
cheyenne_filter = re.compile('|'.join([filter1, filter2]))
dav_filter = re.compile(r'^casper')
hostname = socket.gethostname()
host_on_cheyenne = cheyenne_filter.search(hostname)
host_on_dav = dav_filter.search(hostname)
try:
if host_on_cheyenne:
return 'cheyenne'
elif host_on_dav:
return 'dav'
else:
return 'local'
except:
return 'local'
# +
from distributed.client import default_client
from distributed.utils import format_bytes
import pandas as pd
def run(func, col_subset, chunks, weights, client=None):
client = client or default_client()
client.restart()
info = client.cluster.scheduler.identity()
workers = len(info['workers'])
cores = sum(w['ncores'] for w in info['workers'].values())
memory = sum(w['memory_limit'] for w in info['workers'].values())
memory = memory / 1e6
n = sum(client.ncores().values())
coroutine = func(col_subset, chunks, weights)
machine = get_machine()
name, unit, numerator, chunks_, chunk_size, dataset_size, dataset_shape = next(coroutine)
out = []
while True:
# time.sleep(1)
start = time.time()
try:
next_name, next_unit, next_numerator, next_chunks_, next_chunk_size, next_dataset_size, next_dataset_shape = next(coroutine)
except StopIteration:
break
finally:
end = time.time()
record = { 'machine': machine,
'workers': workers,
'memory': memory,
'name': name,
'duration': end - start,
'unit': unit,
'rate': numerator / (end - start),
'cores': n,
'chunks': chunks_ ,
'chunk_size': chunk_size,
'dataset_size': dataset_size,
'dataset_shape': dataset_shape}
out.append(record)
name = next_name
unit = next_unit
numerator = next_numerator
chunks_ = next_chunks_
chunk_size = next_chunk_size
dataset_size = next_dataset_size
dataset_shape = next_dataset_shape
return pd.DataFrame(out)
# -
def core(col_subset, chunks, weights):
import esmlab
from distributed.utils import format_bytes
from operator import mul
from functools import reduce
ds = col_subset.to_xarray(chunks=chunks, decode_times=False, decode_coords=False)
itemsize = ds.O2.data.dtype.itemsize
yield 'Reading data', 'MB', ds.nbytes / 1e6, str(ds.O2.data.chunksize), (itemsize * reduce(mul, ds.O2.data.chunksize)) / 1e6, ds.nbytes / 1e6, str(ds.O2.data.shape)
ds = ds.persist()
wait(ds)
yield 'Weighted Global mean', 'MB', ds.nbytes / 1e6, str(ds.O2.data.chunksize), (itemsize * reduce(mul, ds.O2.data.chunksize)) / 1e6 , ds.nbytes / 1e6, str(ds.O2.data.shape)
global_mean = esmlab.resample(ds, freq='ann', time_coord_name='time')
global_mean = esmlab.statistics.weighted_mean(global_mean,
weights=weights,
dim=['z_t', 'nlat', 'nlon']).persist()
wait(global_mean)
L = []
chunks=[{'time': 12, 'z_t': 30}, {'time': 24, 'z_t': 20}, {'time': 24, 'z_t': 30},
{'time': 48, 'z_t': 30}, {'time': 60, 'z_t': 10}, {'time': 72, 'z_t': 6}]
chunks = chunks[2:]
chunks
# %%time
for ch in chunks:
for func in [core]:
print(ch, func.__name__)
df = run(func, col_subset, ch, weights=MASKED_VOL, client=client)
L.append(df)
ddf = pd.concat(L)
ddf.head()
ddf.groupby(['dataset_size', 'dataset_shape', 'chunks', 'name']).median()
import datetime
filename =f"results/esmlab_scaling_{datetime.datetime.now().strftime('%Y-%m-%d_%H%M.%S')}_.csv"
filename
ddf.to_csv(filename)
# %load_ext watermark
# %watermark --iversion -g -m -v -u -d -h
| benchmarks/scaling-esmlab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: bluesky_2021_1
# language: python
# name: bluesky_2021_1
# ---
# # WIP: save reflection information in stream
# See: https://github.com/bluesky/hklpy/issues/158
# +
import gi
gi.require_version('Hkl', '5.0')
from bluesky import RunEngine
from bluesky.callbacks.best_effort import BestEffortCallback
import bluesky.plans as bp
import bluesky.plan_stubs as bps
import bluesky.preprocessors as bpp
import databroker
import hkl
from hkl import * # TODO: wildcard import, yikes!
import numpy as np
import pyRestTable
from ophyd import Component, Device, EpicsSignal, Signal
from ophyd.signal import AttributeSignal, ArrayAttributeSignal
from ophyd.sim import *
import pandas as pd
bec = BestEffortCallback()
bec.disable_plots()
cat = databroker.temp().v2
RE = RunEngine({})
RE.subscribe(bec)
RE.subscribe(cat.v1.insert)
RE.md["notebook"] = "tst_UB_in_stream"
RE.md["objective"] = "Demonstrate UB matrix save & restore in stream of bluesky run"
# +
from collections import namedtuple
Lattice = namedtuple("LatticeTuple", "a b c alpha beta gamma")
Position = namedtuple("PositionTuple", "omega chi phi tth")
Reflection = namedtuple("ReflectionTuple", "h k l position")
class Holder:
samples = {}
class Reflections:
reflections = []
class MyDevice(Device):
uptime = Component(EpicsSignal, "gp:UPTIME", kind="normal")
apple = Component(Signal, value="Fuji", kind="omitted")
orange = Component(Signal, value="Valencia", kind="omitted")
octopus = Component(Signal, value="spotted", kind="omitted")
stream_name = Component(AttributeSignal, attr="_stream_name", doc="stream name", kind="omitted")
stream_attrs = Component(AttributeSignal, attr="_stream_attrs", doc="attributes in stream", kind="omitted")
_samples = {}
_stream_name = "bozo"
# _stream_attrs = "orange octopus samples stream_name stream_attrs".split()
_stream_attrs = "orange octopus stream_name stream_attrs".split()
# cannot make AttributeSignal from these that can be written by bluesky
paddle = Holder()
spots = Reflections()
def other_streams(self, label=None):
label = label or self._stream_name
yield from bps.create(name=label)
for attr in self._stream_attrs:
yield from bps.read(getattr(self, attr))
yield from bps.save()
yield from bps.create("fruit")
yield from bps.read(self.apple)
yield from bps.read(self.orange)
yield from bps.save()
yield from bps.create("animal")
yield from bps.read(self.octopus)
yield from bps.save()
nitwit = MyDevice("", name="nitwit")
nitwit.paddle.samples["main"] = Lattice(1,1,1,30,60,90)
nitwit.paddle.samples["second"] = Lattice(2,2,2, 2,2,2)
def try_it():
yield from bps.open_run()
yield from bps.create()
yield from bps.read(nitwit)
yield from bps.save()
yield from nitwit.other_streams()
yield from bps.close_run()
# -
nitwit.wait_for_connection()
nitwit.read()
# RE(bp.count([nitwit]))
RE(try_it())
cat[-1]
run = cat[-1]
for stream in list(run):
print(f"{stream = }")
print(getattr(run, stream).read())
nitwit.paddle.samples["main"][:]
nitwit.octopus.read()
for k, v in nitwit.paddle.samples.items():
print(f"{k = }")
print(f"{v[:] = }")
sig = Signal(name="sig", value=dict(main=nitwit.paddle.samples["main"]._asdict()))
print(f"{sig.read() = }")
print(f"{nitwit.paddle.samples['main'] = }")
r = Reflection(4.0, 0., 0., Position(omega=-145.451, chi=0.0, phi=0.0, tth=69.0966))
r[-1][:]
# +
def read_soft_signal(key, value):
yield from bps.read(Signal(name=key, value=value))
def stream_dict(dictionary, label):
yield from bps.create(label)
for k, v in dictionary.items():
yield from read_soft_signal(k, v)
yield from bps.save()
def stream_samples(samples, label="samples"):
if len(samples):
yield from bps.create(label)
for sname, lattice in samples.items():
yield from read_soft_signal(sname, lattice[:])
yield from read_soft_signal("_keys", list(lattice._fields))
yield from bps.save()
else:
# because you have to yield _something_
yield from bps.null()
def stream_test(reflections, label="reflections"):
if len(reflections):
yield from bps.create(label)
keys = []
for i, refl in enumerate(reflections):
key = f"r{i+1}"
keys.append(key)
yield from read_soft_signal(key, (*refl[:3], refl[3][:]))
yield from read_soft_signal(key+"_hkl", refl[:3])
yield from read_soft_signal(key+"_axis_values", refl[3])
yield from read_soft_signal(key+"_wavelength", 2.101)
yield from read_soft_signal("_keys", keys)
yield from bps.save()
else:
# because you have to yield _something_
yield from bps.null()
def stream_reflections(self, label="reflections"):
reflections = self.calc.sample._sample.reflections_get()
if len(reflections):
yield from bps.create(label)
orient_refls = self.calc.sample._orientation_reflections
keys = []
for i, refl in enumerate(reflections):
key = f"r{i+1}"
keys.append(key)
hkl_tuple = refl.hkl_get()
geom = refl.geometry_get()
yield from read_soft_signal(key, (*hkl_tuple[:], geom.axis_values_get(1)))
yield from read_soft_signal(key+"_hkl", hkl_tuple[:])
yield from read_soft_signal(key+"_axis_names", geom.axis_names_get(1))
yield from read_soft_signal(key+"_axis_values", geom.axis_values_get(1))
yield from read_soft_signal(key+"_wavelength", geom.wavelength_get(1))
yield from read_soft_signal(key+"_for_calcUB", refl in orient_refls)
# ignore `flag`, no documentation for it, always 1 (?used by libhkl's GUI?)
yield from read_soft_signal("_keys", keys)
yield from bps.save()
else:
# because you have to yield _something_
yield from bps.null()
def stream_multi(label="multi"):
for i in range(3):
yield from bps.create(label)
yield from read_soft_signal("a", 1.2345 + i)
yield from read_soft_signal("b", f"4.{i}56")
yield from read_soft_signal("arr", [-1, i , 1.1])
yield from bps.save()
else:
# because you have to yield _something_
yield from bps.null()
def streams():
yield from bps.open_run()
yield from stream_samples(nitwit.paddle.samples)
# yield from stream_reflections(
yield from stream_test(
[
Reflection(4.0, 0., 0., Position(omega=-145.451, chi=0.0, phi=0.0, tth=69.0966)),
Reflection(0., 4.0, 0., Position(omega=-145.451, chi=0.0, phi=90.0, tth=69.0966))
]
)
# yield from stream_multi()
yield from bps.close_run()
RE(streams())
for nm, stream in cat[-1].items():
print(f"{nm = }")
print(stream.read())
print("-"*30)
# -
k = "r1"
r = cat[-1].reflections.read()[k+"_hkl"][0]
a = cat[-1].reflections.read()[k+"_axis_values"][0]
(*r.data, a.data)
| examples/tst_UB_in_stream.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="DSPCom-KmApV"
# # Convolutional Neural Network (CNN)
#
# **Learning Objectives**
# 1. We will learn how to configure our CNN to process inputs of CIFAR images
# 2. We will learn how to compile and train the CNN model
# 3. We will learn how to evaluate the CNN model
#
# + [markdown] id="qLGkt5qiyz4E"
# ## Introduction
# This notebook demonstrates training a simple [Convolutional Neural Network](https://developers.google.com/machine-learning/glossary/#convolutional_neural_network) (CNN) to classify [CIFAR images](https://www.cs.toronto.edu/~kriz/cifar.html). Because this notebook uses the [Keras Sequential API](https://www.tensorflow.org/guide/keras/overview), creating and training our model will take just a few lines of code.
#
# Each learning objective will correspond to a __#TODO__ in the notebook where you will complete the notebook cell's code before running. Refer to the [solution](https://github.com/GoogleCloudPlatform/training-data-analyst/blob/master/courses/machine_learning/deepdive2/image_understanding/solutions/cnn.ipynb) for reference.
#
# -
# Use the chown command to change the ownership of the repository.
# !sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# + [markdown] id="m7KBpffWzlxH"
# ## Import TensorFlow
# + id="iAve6DCL4JH4"
# Importing necessary TF version and modules
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
import matplotlib.pyplot as plt
# + [markdown] id="m7KBpffWzlxH"
# This notebook uses TF2.x. Please check your tensorflow version using the cell below.
# -
# Show the currently installed version of TensorFlow
print(tf.__version__)
# + [markdown] id="jRFxccghyMVo"
# ## Download and prepare the CIFAR10 dataset
#
#
# The CIFAR10 dataset contains 60,000 color images in 10 classes, with 6,000 images in each class. The dataset is divided into 50,000 training images and 10,000 testing images. The classes are mutually exclusive and there is no overlap between them.
# + id="JWoEqyMuXFF4"
# Download the CIFAR10 dataset.
(train_images, train_labels), (test_images, test_labels) = datasets.cifar10.load_data()
# Normalize pixel values to be between 0 and 1
train_images, test_images = train_images / 255.0, test_images / 255.0
# + [markdown] id="7wArwCTJJlUa"
# ### Verify the data
#
# To verify that the dataset looks correct, let's plot the first 25 images from the training set and display the class name below each image.
#
# + id="K3PAELE2eSU9"
# Plot the first 25 images and display the class name below each image.
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
# The CIFAR labels happen to be arrays,
# which is why you need the extra index
plt.xlabel(class_names[train_labels[i][0]])
plt.show()
# + [markdown] id="Oewp-wYg31t9"
# ## Lab Task 1: Create the convolutional base
# + [markdown] id="3hQvqXpNyN3x"
# The 6 lines of code below define the convolutional base using a common pattern: a stack of [Conv2D](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2D) and [MaxPooling2D](https://www.tensorflow.org/api_docs/python/tf/keras/layers/MaxPool2D) layers.
#
# As input, a CNN takes tensors of shape (image_height, image_width, color_channels), ignoring the batch size. If you are new to these dimensions, color_channels refers to (R,G,B). In this example, you will configure our CNN to process inputs of shape (32, 32, 3), which is the format of CIFAR images. You can do this by passing the argument `input_shape` to our first layer.
#
# + id="L9YmGQBQPrdn"
# TODO 1 - Write a code to configure our CNN to process inputs of CIFAR images.
# + [markdown] id="lvDVFkg-2DPm"
# Let's display the architecture of our model so far.
# + id="8-C4XBg4UTJy"
# Now, print a useful summary of the model.
model.summary()
# + [markdown] id="_j-AXYeZ2GO5"
# Above, you can see that the output of every Conv2D and MaxPooling2D layer is a 3D tensor of shape (height, width, channels). The width and height dimensions tend to shrink as you go deeper in the network. The number of output channels for each Conv2D layer is controlled by the first argument (e.g., 32 or 64). Typically, as the width and height shrink, you can afford (computationally) to add more output channels in each Conv2D layer.
# + [markdown] id="_v8sVOtG37bT"
# ### Add Dense layers on top
# To complete our model, you will feed the last output tensor from the convolutional base (of shape (4, 4, 64)) into one or more Dense layers to perform classification. Dense layers take vectors as input (which are 1D), while the current output is a 3D tensor. First, you will flatten (or unroll) the 3D output to 1D, then add one or more Dense layers on top. CIFAR has 10 output classes, so you use a final Dense layer with 10 outputs and a softmax activation.
# + id="mRs95d6LUVEi"
# Here, the model.add() method adds a layer instance incrementally for a sequential model.
model.add(layers.Flatten())
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(10))
# + [markdown] id="ipGiQMcR4Gtq"
# Here's the complete architecture of our model.
# + id="8Yu_m-TZUWGX"
# Print a useful summary of the model.
model.summary()
# + [markdown] id="xNKXi-Gy3RO-"
# As you can see, our (4, 4, 64) outputs were flattened into vectors of shape (1024) before going through two Dense layers.
# + [markdown] id="P3odqfHP4M67"
# ## Lab Task 2: Compile and train the model
# + id="MdDzI75PUXrG"
# TODO 2 - Write a code to compile and train a model
# + [markdown] id="jKgyC5K_4O0d"
# ## Lab Task 3: Evaluate the model
# + id="gtyDF0MKUcM7"
# TODO 3 - Write a code to evaluate a model.
# + id="0LvwaKhtUdOo"
# Print the test accuracy.
print(test_acc)
# + [markdown] id="8cfJ8AR03gT5"
# Our simple CNN has achieved a test accuracy of over 70%. Not bad for a few lines of code! For another CNN style, see an example using the Keras subclassing API and a `tf.GradientTape` [here](https://www.tensorflow.org/tutorials/quickstart/advanced).
| courses/machine_learning/deepdive2/image_understanding/labs/cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from ipyleaflet import Map, Heatmap
from random import uniform
import time
def create_random_data(length):
"Return a list of some random lat/lon/value triples."
return [[uniform(-80, 80),
uniform(-180, 180),
uniform(0, 1000)] for i in range(length)]
m = Map(center=[0, 0], zoom=2)
m
heat = Heatmap(locations=create_random_data(1000), radius=20, blur=10)
m.add_layer(heat)
for i in range(100):
heat.locations = create_random_data(1000)
time.sleep(0.1)
heat.radius = 30
heat.blur = 50
heat.max = 0.5
heat.gradient = {0.4: 'red', 0.6: 'yellow', 0.7: 'lime', 0.8: 'cyan', 1.0: 'blue'}
heat.locations = [[uniform(-80, 80), uniform(-180, 180), uniform(0, 1000)] for i in range(1000)]
| examples/Heatmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Proprietary content. © Great Learning. All Rights Reserved. Unauthorized use or distribution prohibited.
# + [markdown] id="hirWJRX7qKqI"
# ## Read PDF file with python
# + colab={"base_uri": "https://localhost:8080/"} id="pHNNH6gOqNb4" outputId="22f502bc-15b3-4ff2-ba25-dc2520238446"
# !pip install PyPDF2
# + colab={"base_uri": "https://localhost:8080/"} id="qWC1DRjKqS7s" outputId="74f3f0b4-2876-4559-fd1d-148612a617b8"
# importing required modules
import PyPDF2
# creating a pdf file object
data = open('/content/Introduction_to_python_week_1.pdf', 'rb')
# creating a pdf reader object
reader = PyPDF2.PdfFileReader(data)
# printing number of pages in pdf file
print(reader.numPages)
# + id="OjjcNckVq5GD"
# creating a page object
page= reader.getPage(0)
# + colab={"base_uri": "https://localhost:8080/"} id="xmrD80d1q-Nh" outputId="c93560a8-b84d-4e2e-8594-6da0d02edc0a"
# extracting text from page
print(page.extractText())
# closing the pdf file object
data.close()
# + [markdown] id="iFwak8iArHeQ"
# ## Working with text file in Python
# + colab={"base_uri": "https://localhost:8080/"} id="6GKywv3trLPM" outputId="3ec20171-032f-4d74-b65a-bca130cce6a6"
text_data= open("/content/demo_file.txt", "r")
print(text_data.read())
# + [markdown] id="BXe11qXBrrzx"
# ## Working with CSV file
# + id="bvJYp7g7tXhi"
import numpy as np
import pandas as pd
# + id="DQgpaxkFusT8"
data=pd.read_csv("/content/Reddit_Data.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="c1B1OG43x6Ia" outputId="ebc9507f-0c97-439b-8861-c5688df818ea"
data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="rCfL8CZPrvOP" outputId="23e1a232-3e10-425b-be82-5fe3a33af6fc"
data.columns
# + colab={"base_uri": "https://localhost:8080/"} id="bhYTxz-Trw0W" outputId="e10d0acd-83c7-44c4-bf64-6c957e1440ca"
data.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="PJx49Pk5rydB" outputId="b46d1977-1c63-4d5c-979d-c0128d07843e"
data.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="W6e9nrRIr0qw" outputId="683bf41a-0745-4158-f112-1276ef1ba481"
data.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="2tkeSfvbr5ys" outputId="28824955-015e-43c1-f918-32280898bdb9"
data.isnull().any()
# + id="h_9X5Pn_r-t1"
data_part=data.dropna()
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="o2veumzssUe-" outputId="fb60dbec-0128-44b9-e541-cd4dd728e1c6"
data_part
# + colab={"base_uri": "https://localhost:8080/"} id="MYrAhAtGsVzh" outputId="99d42d15-c5c3-4135-8cfc-97b7c593867b"
data_part.shape
# + id="Qdn__bQIxHnv"
# + [markdown] id="cxUgjr3Dx_O-"
# ## Line tokenization
# + colab={"base_uri": "https://localhost:8080/"} id="zhZLxffasX5K" outputId="15c5ba6e-5e76-494c-bc00-ce5837fd80c5"
import nltk
data = "Welcome to Great Learning!! "
tokens = nltk.sent_tokenize(data)
print (tokens)
# + [markdown] id="0hzGl1QjyClA"
# ## Word tokenization
# + colab={"base_uri": "https://localhost:8080/"} id="qxJ5sbbUxECn" outputId="6c796862-cf0a-42b6-bd0b-c95e791ae354"
tokens = nltk.word_tokenize(data)
print (tokens)
# + [markdown] id="DV-CwHe4ypiQ"
# ## Stemming
# + colab={"base_uri": "https://localhost:8080/"} id="1a8AwRDIyOav" outputId="ea2a7823-c7c6-4536-98ee-5ac5931b819b"
# import these modules
from nltk.stem import PorterStemmer
ps = PorterStemmer()
# choose some words to be stemmed
words = ["Like","Liking","Likes"]
for i in words:
print(i, " : ", ps.stem(w))
# + [markdown] id="3XU-mee5zK07"
# ## Lemmatization
# + colab={"base_uri": "https://localhost:8080/"} id="JIifxZskzPZS" outputId="0a9f1415-78e1-41dd-ba5d-6c7591bc76ee"
import nltk
nltk.download('wordnet')
# + colab={"base_uri": "https://localhost:8080/"} id="FGOuSuxNyzDY" outputId="7c10749c-4a60-4e1c-c998-dbc1d6635700"
# import these modules
from nltk.stem import WordNetLemmatizer
lemmati = WordNetLemmatizer()
print("socks:", lemmati.lemmatize("socks"))
print("sons:", lemmati.lemmatize("sons"))
# + [markdown] id="6rFlCwwi0GIk"
# ## Removing stop words
# + colab={"base_uri": "https://localhost:8080/"} id="o5-kXnwzzHD6" outputId="e432061f-6c9e-45ba-f773-a4663cb2db42"
from nltk.corpus import stopwords
import nltk
nltk.download('stopwords')
data = """Data science is one of the most trendind field to work with. It needs data to give prediction by using the past scenarios"""
stop_word = set(stopwords.words('english'))
print(stopwords.words() [620:680])
# + id="riWuQgpu0VmN"
data= nltk.word_tokenize(data)
# + id="gGuRxvTL1Jw2"
stops = set(stopwords.words('english'))
# + colab={"base_uri": "https://localhost:8080/"} id="-qPgOusb1Odd" outputId="b8ea4cf7-db94-42b3-9e74-e13e9e9518d7"
for word in data:
if word not in stops:
print(word)
# + id="xzi-9N8n1TkY"
| Movie Review Analysis/demo_of_nlp.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SITS
# language: R
# name: sits
# ---
# <img src="../../../img/logo-bdc.png" align="right" width="64"/>
#
# # <span style="color: #336699">Earth Observation Data Cubes for Brazil: Requirements, Methodology and Products</span>
# <hr style="border:2px solid #0077b9;">
#
# <br/>
#
# <div style="text-align: center;font-size: 90%;">
# <NAME> <sup><a href="mailto:<EMAIL>"><i class="far fa-lg fa-envelope"></i></a> <a href="https://orcid.org/0000-0003-0953-4132"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>, <NAME> <sup><a href="mailto:<EMAIL>"><i class="far fa-lg fa-envelope"></i></a> <a href="https://orcid.org/0000-0001-7966-2880"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>, <NAME> <sup><a href="mailto:<EMAIL>"><i class="far fa-lg fa-envelope"></i></a> <a href="https://orcid.org/0000-0002-3334-4315"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>, <NAME> <sup><a href="mailto:<EMAIL>"><i class="far fa-lg fa-envelope"></i></a> <a href="https://orcid.org/0000-0002-3397-6232"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>,<br/>
# <NAME> <sup><a href="mailto:<EMAIL>"><i class="far fa-lg fa-envelope"></i></a> <a href="https://orcid.org/0000-0003-2656-5504"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>, <NAME> <sup><a href="mailto:<EMAIL>"><i class="far fa-lg fa-envelope"></i></a> <a href="https://orcid.org/0000-0003-1104-3607"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>, <NAME><sup>* <a href="mailto:<EMAIL>"><i class="far fa-lg fa-envelope"></i></a> <a href="https://orcid.org/0000-0001-7534-0219"><i class="fab fa-lg fa-orcid" style="color: #a6ce39"></i></a></sup>
# <br/><br/>
# Earth Observation and Geoinformatics Division, National Institute for Space Research (INPE)
# <br/>
# Avenida dos Astronautas, 1758, Jardim da Granja, São José dos Campos, SP 12227-010, Brazil
# <br/><br/>
# <sup>*</sup> Author to whom correspondence should be addressed.
# <br/><br/>
# February 24, 2021
# </div>
#
# <br/>
#
# <div style="text-align: justify; margin-left: 10%; margin-right: 10%;">
# <b>Abstract.</b> This Jupyter Notebook compendium contains useful information for the creation of land use and land cover (LULC) maps using Earth observations data cubes and machine learning (ML) techniques. The code is based on the research pipeline described in the paper <em>Earth Observation Data Cubes for Brazil: Requirements, Methodology and Products</em>. All the datasets required to the reproducibility of the work is also available.
# </div>
#
# <br/>
# <div style="text-align: justify; margin-left: 15%; margin-right: 15%;font-size: 75%; border-style: solid; border-color: #0077b9; border-width: 1px; padding: 5px;">
# <b>This Jupyter Notebook is supplement to the <a href="https://www.mdpi.com/2072-4292/12/24/4033/htm#sec5-remotesensing-12-04033" target="_blank">Section 5</a> of the following paper:</b>
# <div style="margin-left: 10px; margin-right: 10px">
# <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>. 2020. Earth Observation Data Cubes for Brazil: Requirements, Methodology and Products. Remote Sens. 12, no. 24: 4033. DOI: <a href="https://doi.org/10.3390/rs12244033" target="_blank">10.3390/rs12244033</a>.
# </div>
# </div>
# # <span style="color: #336699">Land Use and Cover Mapping from Sentinel-2/MSI Data Cubes</span>
# <hr style="border:1px solid #0077b9;">
#
# This document will present the steps used to generate the Sentinel-2/MSI classification map presented in the paper. As presented in the article, the classification process was done using the [SITS R package](https://github.com/e-sensing/sits).
# ## <span style="color: #336699">Study Area and samples</span>
# <hr style="border:0.5px solid #0077b9;">
#
# The article associated with this example of reproduction uses a region of Bahia, Brazil, between the Cerrado and Caatinga biomes, as the study area. In this example, the classification will be done using a small region within the research paper study area to reduce computational complexity.
#
# On the other hand, the samples used will be the same ones presented in the article, with the difference that these will have the time series associated with each sample extracted again. The figure below shows the selected region for the classification and used samples.
#
# <div align="center">
# <img src="../../../img/bdc-article/study-area.png" width="600px">
# </div>
# <br/>
# <center><b>Figure 1</b> - Study area in relation to Brazil and its biomes.</center>
# ## <span style="color: #336699">Parameters</span>
# <hr style="border:0.5px solid #0077b9;">
#
# If you want to download and run this notebook in a workflow as a script, you can perform its parameterization through the [papermill library](https://github.com/nteract/papermill).
# + tags=["parameters"]
classification_memsize <- 20 # in GB
classification_multicores <- 20
start_date <- "2018-09-01"
end_date <- "2019-08-31"
MY_ACCESS_KEY <- "My-Token"
Sys.setenv(BDC_ACCESS_KEY = MY_ACCESS_KEY)
# -
# ## <span style="color: #336699">Samples and ROI definition</span>
# <hr style="border:0.5px solid #0077b9;">
#
# +
# fixed parameters
collection <- "S2_10_16D_STK-1"
# define the roi and load samples file
roi <- readRDS(url("https://brazildatacube.dpi.inpe.br/geo-knowledge-hub/bdc-article/roi/roi.rds"))
samples <- readRDS(url("https://brazildatacube.dpi.inpe.br/geo-knowledge-hub/bdc-article/training-samples/rds/training-samples/rds/S2_10_16D_STK_1.rds"))
# -
# > All the results generated in this document will be saved in your user's `/home` directory, inside `out` directory
# +
output_dir <- paste0(path.expand('~/work'), "/bdc-article", "/results", "/S2_10_16D_STK_1")
dir.create(
path = output_dir,
showWarnings = FALSE,
recursive = TRUE
)
# +
set.seed(777) # pseudo-randomic seed
library(sits)
library(rgdal)
# -
# ## <span style="color: #336699">Generating datacube using BDC-STAC</span>
# <hr style="border:0.5px solid #0077b9;">
#
# The classification process was done with the use of STAC. In this approach, the data cubes used for the classification are consumed directly through the STAC service. This process is useful for avoiding data movement.
#
# Following the definitions of the article, below is the definition of the data cube used. The spectral bands `Red`, `Green`, `Blue`, `Near-Infrared (NIR)` and the vegetation indices `EVI` and `NDVI` are applied in the created cube. The temporal extension used in the research paper covers the period of `2018-09` to `2019-08`.
cube <- sits_cube(
type = "BDC",
name = "cube_to_classify",
url = "https://brazildatacube.dpi.inpe.br/stac/",
collection = collection,
start_date = start_date,
end_date = end_date,
roi = roi$search_roi
)
# ## <span style="color: #336699">MultiLayer Perceptron model definition</span>
# <hr style="border:0.5px solid #0077b9;">
#
# For the classification of data cubes, the article presents the use of an MLP network with five hidden layers with 512 neurons, trained with the backpropagation algorithm, using the Adam optimizer. The model uses the ReLu activation function.
#
# Below is the definition of this model using the [SITS package](https://github.com/e-sensing/sits).
#
mlp_model <- sits_deeplearning(layers = c(512, 512, 512, 512, 512),
activation = "relu",
optimizer = keras::optimizer_adam(lr = 0.001),
epochs = 200)
# Below, the defined model is trained using the same samples used in the article.
dl_model <- sits_train(samples, mlp_model)
# ## <span style="color: #336699">Classify the datacube</span>
# <hr style="border:0.5px solid #0077b9;">
#
# > This is a time-consuming process
#
probs <- sits_classify(data = cube,
ml_model = dl_model,
memsize = classification_memsize,
multicores = classification_multicores,
output_dir = output_dir)
# ## <span style="color: #336699">Generate classification label map</span>
# <hr style="border:0.5px solid #0077b9;">
probs_smoothed <- sits_smooth(probs, type = "bayes", output_dir = output_dir)
labels <- sits_label_classification(probs_smoothed, output_dir = output_dir)
# ## <span style="color: #336699">Visualizing classification map</span>
# <hr style="border:0.5px solid #0077b9;">
#
# > The raster load in this step was generated automaticaly with `sits_label_classification` function
#
# +
gdalUtils::mosaic_rasters(c(
paste0(output_dir, "/cube_to_classify_088097_probs_class_2018_8_2019_7_v1.tif"),
paste0(output_dir, "/cube_to_classify_089097_probs_class_2018_8_2019_7_v1.tif"),
paste0(output_dir, "/cube_to_classify_089098_probs_class_2018_8_2019_7_v1.tif")
), paste0(output_dir, "/cube_to_classify_merged_probs_class_2018_8_2019_7_v1.tif"))
plot(
raster::raster(paste0(output_dir, "/cube_to_classify_merged_probs_class_2018_8_2019_7_v1.tif"))
)
# -
# ## <span style="color: #336699">Save the results</span>
# <hr style="border:0.5px solid #0077b9;">
# +
# labels
saveRDS(
labels, file = paste0(output_dir, "/labels.rds")
)
# probs
saveRDS(
probs, file = paste0(output_dir, "/probs_cube.rds")
)
# smoothed probs
saveRDS(
probs_smoothed, file = paste0(output_dir, "/probs_smoothed_cube.rds")
)
| jupyter/R/bdc-article/06_S2_10_16D_STK-1_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [Table of Contents](http://nbviewer.ipython.org/github/rlabbe/Kalman-and-Bayesian-Filters-in-Python/blob/master/table_of_contents.ipynb)
# # The Extended Kalman Filter
#format the book
# %matplotlib inline
from __future__ import division, print_function
from book_format import load_style
load_style()
# We have developed the theory for the linear Kalman filter. Then, in the last two chapters we broached the topic of using Kalman filters for nonlinear problems. In this chapter we will learn the Extended Kalman filter (EKF). The EKF handles nonlinearity by linearizing the system at the point of the current estimate, and then the linear Kalman filter is used to filter this linearized system. It was one of the very first techniques used for nonlinear problems, and it remains the most common technique.
#
# The EKF provides significant mathematical challenges to the designer of the filter; this is the most challenging chapter of the book. To be honest, I do everything I can to avoid the EKF in favor of other techniques that have been developed to filter nonlinear problems. However, the topic is unavoidable; all classic papers and a majority of current papers in the field use the EKF. Even if you do not use the EKF in your own work you will need to be familiar with the topic to be able to read the literature.
# ## Linearizing the Kalman Filter
#
# The Kalman filter uses linear equations, so it does not work with nonlinear problems. Problems can be nonlinear in two ways. First, the process model might be nonlinear. An object falling through the atmosphere encounters drag which reduces its acceleration. The drag coefficient varies based on the velocity the object. The resulting behavior is nonlinear - it cannot be modeled with linear equations. Second, the measurements could be nonlinear. For example, a radar gives a range and bearing to a target. We use trigonometry, which is nonlinear, to compute the position of the target.
#
# For the linear filter we have these equations for the process and measurement models:
#
# $$\begin{aligned}\overline{\mathbf x} &= \mathbf{Ax} + \mathbf{Bu} + w_x\\
# \mathbf z &= \mathbf{Hx} + w_z
# \end{aligned}$$
#
# For the nonlinear model these equations must be modified to read:
#
# $$\begin{aligned}\overline{\mathbf x} &= f(\mathbf x, \mathbf u) + w_x\\
# \mathbf z &= h(\mathbf x) + w_z
# \end{aligned}$$
#
# The linear expression $\mathbf{Ax} + \mathbf{Bu}$ is replaced by a nonlinear function $f(\mathbf x, \mathbf u)$, and the linear expression $\mathbf{Hx}$ is replaced by a nonlinear function $h(\mathbf x)$.
#
# You might imagine that we proceed by finding a new set of Kalman filter equations that optimally solve these equations. But if you remember the charts in the **Nonlinear Filtering** chapter you'll recall that passing a Gaussian through a nonlinear function results in a probability distribution that is no longer Gaussian. So this will not work.
#
# The EKF does not alter the Kalman filter's linear equations. Instead, it *linearizes* the nonlinear equations at the point of the current estimate, and uses this linearization in the linear Kalman filter.
#
# *Linearize* means what it sounds like. We find a line that most closely matches the curve at a defined point. The graph below linearizes the parabola $f(x)=x^2−2x$ at $x=1.5$.
import ekf_internal
ekf_internal.show_linearization()
# If the curve above is the process model, then the dotted lines shows the linearization of that curve for the estimate $x=1.5$.
#
# We linearize systems by taking the derivative, which finds an expression for the slope of the curve:
#
# $$\begin{aligned}
# f(x) &= x^2 -2x \\
# \frac{df}{dx} &= 2x - 2
# \end{aligned}$$
#
# and then evaluating it at $x$:
#
# $$\begin{aligned}m &= f'(x=1.5) \\&= 2(1.5) - 2 \\&= 1\end{aligned}$$
#
# Linearizing systems of differential equations is more complicated. We linearize $f(\mathbf x, \mathbf u)$, and $h(\mathbf x)$ by taking the partial derivatives of each to evaluate $\mathbf A$ and $\mathbf H$ at the point $\mathbf x_t$ and $\mathbf u_t$. We call the partial derivative of a matrix the *Jacobian*. This gives us the the system dynamics matrix and measurement model matrix:
#
# $$
# \begin{aligned}
# \mathbf A
# &= {\frac{\partial{f(\mathbf x_t, \mathbf u_t)}}{\partial{\mathbf x}}}\biggr|_{{\mathbf x_t},{\mathbf u_t}} \\
# \mathbf H &= \frac{\partial{h(\mathbf x_t)}}{\partial{\mathbf x}}\biggr|_{\mathbf x_t}
# \end{aligned}
# $$
#
# $h(\overline{\mathbf x})$ is computed with the prior, but I drop the bar on for notational convenience.
# Finally, we find the discrete state transition matrix $\mathbf F$ by using the Taylor series expansion of $e^{\mathbf A \Delta t}$:
#
# $$\mathbf F = e^{\mathbf A\Delta t} = \mathbf{I} + \mathbf A\Delta t + \frac{(\mathbf A\Delta t)^2}{2!} + \frac{(\mathbf A\Delta t)^3}{3!} + ... $$
#
# Alternatively, you can use one of the other techniques we learned in the **Kalman Math** chapter.
#
# This leads to the following equations for the EKF. I placed them beside the equations for the linear Kalman filter, and put boxes around the changes:
#
# $$\begin{array}{l|l}
# \text{linear Kalman filter} & \text{EKF} \\
# \hline
# & \boxed{\mathbf A = {\frac{\partial{f(\mathbf x_t, \mathbf u_t)}}{\partial{\mathbf x}}}\biggr|_{{\mathbf x_t},{\mathbf u_t}}} \\
# & \boxed{\mathbf F = e^{\mathbf A \Delta t}} \\
# \mathbf{\overline x} = \mathbf{Fx} + \mathbf{Bu} & \boxed{\mathbf{\overline x} = f(\mathbf x, \mathbf u)} \\
# \mathbf{\overline P} = \mathbf{FPF}^\mathsf{T}+\mathbf Q & \mathbf{\overline P} = \mathbf{FPF}^\mathsf{T}+\mathbf Q \\
# \hline
# & \boxed{\mathbf H = \frac{\partial{h(\mathbf x_t)}}{\partial{\mathbf x}}\biggr|_{\mathbf x_t}} \\
# \textbf{y} = \mathbf z - \mathbf{H \bar{x}} & \textbf{y} = \mathbf z - \boxed{h(\bar{x})}\\
# \mathbf{K} = \mathbf{\bar{P}H}^\mathsf{T} (\mathbf{H\bar{P}H}^\mathsf{T} + \mathbf R)^{-1} & \mathbf{K} = \mathbf{\bar{P}H}^\mathsf{T} (\mathbf{H\bar{P}H}^\mathsf{T} + \mathbf R)^{-1} \\
# \mathbf x=\mathbf{\bar{x}} +\mathbf{K\textbf{y}} & \mathbf x=\mathbf{\bar{x}} +\mathbf{K\textbf{y}} \\
# \mathbf P= (\mathbf{I}-\mathbf{KH})\mathbf{\bar{P}} & \mathbf P= (\mathbf{I}-\mathbf{KH})\mathbf{\bar{P}}
# \end{array}$$
#
# We don't normally use $\mathbf{Fx}$ to propagate the state for the EKF as the linearization causes inaccuracies. It is typical to compute $\overline{\mathbf x}$ using a suitable numerical integration technique such as Euler or Runge Kutta. Thus I wrote $\mathbf{\overline x} = f(\mathbf x, \mathbf u)$. For the same reasons we don't use $\mathbf{H\overline{x}}$ in the computation for the residual, opting for the more accurate $h(\overline{\mathbf x})$.
#
# I think the easiest way to understand the EKF is to start off with an example. Later you may want to come back and reread this section.
# ## Example: Tracking a Airplane
# This example tracks an airplane using ground based radar. We implemented a UKF for this problem in the last chapter. Now we will implement an EKF for the same problem so we can compare both the filter performance and the level of effort required to implement the filter.
#
# Radars work by emitting a beam of radio waves and scanning for a return bounce. Anything in the beam's path will reflects some of the signal back to the radar. By timing how long it takes for the reflected signal to get back to the radar the system can compute the *slant distance* - the straight line distance from the radar installation to the object.
#
# The relationship between the radar's slant range distance and bearing with the horizontal position $x$ and altitude $y$ of the aircraft is illustrated in the figure below:
import ekf_internal
ekf_internal.show_radar_chart()
# This gives us the equalities:
#
# $$\theta = \tan^{-1} \frac y x\\
# r^2 = x^2 + y^2$$
# ### Design the State Variables
#
# We want to track the position of an aircraft assuming a constant velocity and altitude, and measurements of the slant distance to the aircraft. That means we need 3 state variables - horizontal distance, horizonal velocity, and altitude:
#
# $$\mathbf x = \begin{bmatrix}\mathtt{distance} \\\mathtt{velocity}\\ \mathtt{altitude}\end{bmatrix}= \begin{bmatrix}x \\ \dot x\\ y\end{bmatrix}$$
# ### Design the Process Model
#
# We assume a Newtonian, kinematic system for the aircraft. We've used this model in previous chapters, so by inspection you may recognize that we want
#
# $$\mathbf F = \left[\begin{array}{cc|c} 1 & \Delta t & 0\\
# 0 & 1 & 0 \\ \hline
# 0 & 0 & 1\end{array}\right]$$
#
# I've partioned the matrix into blocks to show the upper left block is a constant velocity model for $x$, and the lower right block is a constant position model for $y$.
#
# However, let's practice finding these matrix for a nonlinear system. We model nonlinear systems with a set of differential equations. We need an equation in the form
#
# $$\dot{\mathbf x} = \mathbf{Ax} + \mathbf{w}$$
# where $\mathbf{w}$ is the system noise.
#
# The variables $x$ and $y$ are independent so we can compute them separately. The differential equations for motion in one dimension are:
#
# $$\begin{aligned}v &= \dot x \\
# a &= \ddot{x} = 0\end{aligned}$$
#
# Now we put the differential equations into state-space form. If this was a second or greater order differential system we would have to first reduce them to an equivalent set of first degree equations. The equations are first order, so we put them in state space matrix form as
#
# $$\begin{aligned}\begin{bmatrix}\dot x \\ \ddot{x}\end{bmatrix} &= \begin{bmatrix}0&1\\0&0\end{bmatrix} \begin{bmatrix}x \\
# \dot x\end{bmatrix} \\ \dot{\mathbf x} &= \mathbf{Ax}\end{aligned}$$
# where $\mathbf A=\begin{bmatrix}0&1\\0&0\end{bmatrix}$.
#
# Recall that $\mathbf A$ is the *system dynamics matrix*. It describes a set of linear differential equations. From it we must compute the state transition matrix $\mathbf F$. $\mathbf F$ describes a discrete set of linear equations which compute $\mathbf x$ for a discrete time step $\Delta t$.
#
# A common way to compute $\mathbf F$ is to use the power series expansion of the matrix exponential:
#
# $$\mathbf F(\Delta t) = e^{\mathbf A\Delta t} = \mathbf{I} + \mathbf A\Delta t + \frac{(\mathbf A\Delta t)^2}{2!} + \frac{(\mathbf A \Delta t)^3}{3!} + ... $$
#
#
# $\mathbf A^2 = \begin{bmatrix}0&0\\0&0\end{bmatrix}$, so all higher powers of $\mathbf A$ are also $\mathbf{0}$. Thus the power series expansion is:
#
# $$
# \begin{aligned}
# \mathbf F &=\mathbf{I} + \mathbf At + \mathbf{0} \\
# &= \begin{bmatrix}1&0\\0&1\end{bmatrix} + \begin{bmatrix}0&1\\0&0\end{bmatrix}\Delta t\\
# \mathbf F &= \begin{bmatrix}1&\Delta t\\0&1\end{bmatrix}
# \end{aligned}$$
#
# This is the same result used by the kinematic equations! This exercise was unnecessary other than to illustrate finding the state transition matrix from linear differential equations. We will conclude the chapter with an example that will require the use of this technique.
# ### Design the Measurement Model
#
# The measurement function takes the state estimate of the prior $\overline{\mathbf x}$ and turn it into a measurement of the slant range distance. For notational convenience I will use $\mathbf x$, not $\overline{\mathbf x}$. We use the Pythagorean theorem to derive:
#
# $$h(\mathbf x) = \sqrt{x^2 + y^2}$$
#
# The relationship between the slant distance and the position on the ground is nonlinear due to the square root. We linearize it by evaluating its partial derivative at $\mathbf x_t$:
#
# $$
# \mathbf H = \frac{\partial{h(\mathbf x)}}{\partial{\mathbf x}}\biggr|_{\mathbf x_t}
# $$
#
# The partial derivative of a matrix is called a Jacobian, and takes the form
#
# $$\frac{\partial \mathbf H}{\partial \mathbf x} =
# \begin{bmatrix}
# \frac{\partial h_1}{\partial x_1} & \frac{\partial h_1}{\partial x_2} &\dots \\
# \frac{\partial h_2}{\partial x_1} & \frac{\partial h_2}{\partial x_2} &\dots \\
# \vdots & \vdots
# \end{bmatrix}
# $$
#
# In other words, each element in the matrix is the partial derivative of the function $h$ with respect to the $x$ variables. For our problem we have
#
# $$\mathbf H = \begin{bmatrix}{\partial h}/{\partial x} & {\partial h}/{\partial \dot{x}} & {\partial h}/{\partial y}\end{bmatrix}$$
#
# Solving each in turn:
#
# $$\begin{aligned}
# \frac{\partial h}{\partial x} &= \frac{\partial}{\partial x} \sqrt{x^2 + y^2} \\
# &= \frac{x}{\sqrt{x^2 + y^2}}
# \end{aligned}$$
#
# and
#
# $$\begin{aligned}
# \frac{\partial h}{\partial \dot{x}} &=
# \frac{\partial}{\partial \dot{x}} \sqrt{x^2 + y^2} \\
# &= 0
# \end{aligned}$$
#
# and
#
# $$\begin{aligned}
# \frac{\partial h}{\partial y} &= \frac{\partial}{\partial y} \sqrt{x^2 + y^2} \\
# &= \frac{y}{\sqrt{x^2 + y^2}}
# \end{aligned}$$
#
# giving us
#
# $$\mathbf H =
# \begin{bmatrix}
# \frac{x}{\sqrt{x^2 + y^2}} &
# 0 &
# &
# \frac{y}{\sqrt{x^2 + y^2}}
# \end{bmatrix}$$
#
# This may seem daunting, so step back and recognize that all of this math is doing something very simple. We have an equation for the slant range to the airplane which is nonlinear. The Kalman filter only works with linear equations, so we need to find a linear equation that approximates $\mathbf H$. As we discussed above, finding the slope of a nonlinear equation at a given point is a good approximation. For the Kalman filter, the 'given point' is the state variable $\mathbf x$ so we need to take the derivative of the slant range with respect to $\mathbf x$. For the linear Kalman filter $\mathbf H$ was a constant that we computed prior to running the filter. For the EKF $\mathbf H$ is updated at each step as the evaluation point $\overline{\mathbf x}$ changes at each epoch.
#
# To make this more concrete, let's now write a Python function that computes the Jacobian of $h$ for this problem.
from math import sqrt
def HJacobian_at(x):
""" compute Jacobian of H matrix at x """
horiz_dist = x[0]
altitude = x[2]
denom = sqrt(horiz_dist**2 + altitude**2)
return array ([[horiz_dist/denom, 0., altitude/denom]])
# Finally, let's provide the code for $h(\mathbf x)$
def hx(x):
""" compute measurement for slant range that
would correspond to state x.
"""
return (x[0]**2 + x[2]**2) ** 0.5
# Now lets write a simulation for our radar.
# +
from numpy.random import randn
import math
class RadarSim(object):
""" Simulates the radar signal returns from an object
flying at a constant altityude and velocity in 1D.
"""
def __init__(self, dt, pos, vel, alt):
self.pos = pos
self.vel = vel
self.alt = alt
self.dt = dt
def get_range(self):
""" Returns slant range to the object. Call once
for each new measurement at dt time from last call.
"""
# add some process noise to the system
self.vel = self.vel + .1*randn()
self.alt = self.alt + .1*randn()
self.pos = self.pos + self.vel*self.dt
# add measurement noise
err = self.pos * 0.05*randn()
slant_dist = math.sqrt(self.pos**2 + self.alt**2)
return slant_dist + err
# -
# ### Design Process and Measurement Noise
#
# The radar measures the range to a target. We will use $\sigma_{range}= 5$ meters for the noise. This gives us
#
# $$\mathbf R = \begin{bmatrix}\sigma_{range}^2\end{bmatrix} = \begin{bmatrix}25\end{bmatrix}$$
#
#
# The design of $\mathbf Q$ requires some discussion. The state $\mathbf x= \begin{bmatrix}x & \dot x & y\end{bmatrix}^\mathtt{T}$. The first two elements are position (down range distance) and velocity, so we can use `Q_discrete_white_noise` noise to compute the values for the upper left hand side of $\mathbf Q$. The third element of $\mathbf x$ is altitude, which we are assuming is independent of the down range distance. That leads us to a block design of $\mathbf Q$ of:
#
# $$\mathbf Q = \begin{bmatrix}\mathbf Q_\mathtt{x} & 0 \\ 0 & \mathbf Q_\mathtt{y}\end{bmatrix}$$
# ### Implementation
#
# `FilterPy` provides the class `ExtendedKalmanFilter`. It works similarly to the `KalmanFilter` class we have been using, except that it allows you to provide a function that computes the Jacobian of $\mathbf H$ and the function $h(\mathbf x)$.
#
# We start by importing the filter and creating it. The dimension of `x` is 3 and `z` has dimension 1.
#
# ```python
# from filterpy.kalman import ExtendedKalmanFilter
#
# rk = ExtendedKalmanFilter(dim_x=3, dim_z=1)
# ```
# We create the radar simulator:
# ```python
# radar = RadarSim(dt, pos=0., vel=100., alt=1000.)
# ```
# We will initialize the filter near the airplane's actual position:
#
# ```python
# rk.x = array([radar.pos, radar.vel-10, radar.alt+100])
# ```
#
# We assign the system matrix using the first term of the Taylor series expansion we computed above:
#
# ```python
# dt = 0.05
# rk.F = eye(3) + array([[0, 1, 0],
# [0, 0, 0],
# [0, 0, 0]])*dt
# ```
#
# After assigning reasonable values to $\mathbf R$, $\mathbf Q$, and $\mathbf P$ we can run the filter with a simple loop. We pass the functions for computing the Jacobian of $\mathbf H$ and $h(x)$ into the `update` method.
#
# ```python
# for i in range(int(20/dt)):
# z = radar.get_range()
# rk.update(array([z]), HJacobian_at, hx)
# rk.predict()
# ```
#
# Adding some boilerplate code to save and plot the results we get:
# +
from filterpy.common import Q_discrete_white_noise
from filterpy.kalman import ExtendedKalmanFilter
from numpy import eye, array, asarray
import numpy as np
dt = 0.05
rk = ExtendedKalmanFilter(dim_x=3, dim_z=1)
radar = RadarSim(dt, pos=0., vel=100., alt=1000.)
# make an imperfect starting guess
rk.x = array([radar.pos-100, radar.vel+100, radar.alt+1000])
rk.F = eye(3) + array([[0, 1, 0],
[0, 0, 0],
[0, 0, 0]]) * dt
range_std = 5. # meters
rk.R = np.diag([range_std**2])
rk.Q[0:2, 0:2] = Q_discrete_white_noise(2, dt=dt, var=0.1)
rk.Q[2,2] = 0.1
rk.P *= 50
xs, track = [], []
for i in range(int(20/dt)):
z = radar.get_range()
track.append((radar.pos, radar.vel, radar.alt))
rk.update(array([z]), HJacobian_at, hx)
xs.append(rk.x)
rk.predict()
xs = asarray(xs)
track = asarray(track)
time = np.arange(0, len(xs)*dt, dt)
ekf_internal.plot_radar(xs, track, time)
# -
# ## Using SymPy to compute Jacobians
#
# Depending on your experience with derivatives you may have found the computation of the Jacobian difficult. Even if you found it easy, a slightly more difficult problem easily leads to very difficult computations.
#
# As explained in Appendix A, we can use the SymPy package to compute the Jacobian for us.
# +
import sympy
sympy.init_printing(use_latex=True)
x, x_vel, y = sympy.symbols('x, x_vel y')
H = sympy.Matrix([sympy.sqrt(x**2 + y**2)])
state = sympy.Matrix([x, x_vel, y])
H.jacobian(state)
# -
# This result is the same as the result we computed above, and with much less effort on our part!
# ## Robot Localization
#
# It's time to try a real problem. I warn you that this section is difficult. However, most books choose simple, textbook problems with simple answers, and you are left wondering how to solve a real world problem.
#
# We will consider the problem of robot localization. We already implemented this in the **Unscented Kalman Filter** chapter, and I recommend you read it now if you haven't already. In this scenario we have a robot that is moving through a landscape using a sensor to detect landmarks. This could be a self driving car using computer vision to identify trees, buildings, and other landmarks. It might be one of those small robots that vacuum your house, or a robot in a warehouse.
#
# The robot has 4 wheels in the same configuration used by automobiles. It maneuvers by pivoting the front wheels. This causes the robot to pivot around the rear axle while moving forward. This is nonlinear behavior which we will have to model.
#
# The robot has a sensor that measures the range and bearing to known targets in the landscape. This is nonlinear because computing a position from a range and bearing requires square roots and trigonometry.
#
# Both the process model and measurement models are nonlinear. The EKF accommodates both, so we provisionally conclude that the EKF is a viable choice for this problem.
# ### Robot Motion Model
#
# At a first approximation an automobile steers by pivoting the front tires while moving forward. The front of the car moves in the direction that the wheels are pointing while pivoting around the rear tires. This simple description is complicated by issues such as slippage due to friction, the differing behavior of the rubber tires at different speeds, and the need for the outside tire to travel a different radius than the inner tire. Accurately modeling steering requires a complicated set of differential equations.
#
# For lower speed robotic applications a simpler *bicycle model* has been found to perform well. This is a depiction of the model:
ekf_internal.plot_bicycle()
# In the **Unscented Kalman Filter** chapter we derived these equations:
#
# $$\begin{aligned}
# \beta &= \frac d w \tan(\alpha) \\
# x &= x - R\sin(\theta) + R\sin(\theta + \beta) \\
# y &= y + R\cos(\theta) - R\cos(\theta + \beta) \\
# \theta &= \theta + \beta
# \end{aligned}
# $$
#
# where $\theta$ is the robot's heading.
#
# You do not need to understand this model in detail if you are not interested in steering models. The important thing to recognize is that our motion model is nonlinear, and we will need to deal with that with our Kalman filter.
# ### Design the State Variables
#
# For our filter we will maintain the position $x,y$ and orientation $\theta$ of the robot:
#
# $$\mathbf x = \begin{bmatrix}x \\ y \\ \theta\end{bmatrix}$$
#
# Our control input $\mathbf u$ is the velocity $v$ and steering angle $\alpha$:
#
# $$\mathbf u = \begin{bmatrix}v \\ \alpha\end{bmatrix}$$
# ### Design the System Model
#
# We model our system as a nonlinear motion model plus noise.
#
# $$\overline x = x + f(x, u) + \mathcal{N}(0, Q)$$
#
#
#
# Using the motion model for a robot that we created above, we can expand this to
#
# $$\overline{\begin{bmatrix}x\\y\\\theta\end{bmatrix}} = \begin{bmatrix}x\\y\\\theta\end{bmatrix} +
# \begin{bmatrix}- R\sin(\theta) + R\sin(\theta + \beta) \\
# R\cos(\theta) - R\cos(\theta + \beta) \\
# \beta\end{bmatrix}$$
# We find The $\mathbf F$ by taking the Jacobian of $f(x,u)$.
#
# $$\mathbf F = \frac{\partial f(x, u)}{\partial x} =\begin{bmatrix}
# \frac{\partial \dot x}{\partial x} &
# \frac{\partial \dot x}{\partial y} &
# \frac{\partial \dot x}{\partial \theta}\\
# \frac{\partial \dot y}{\partial x} &
# \frac{\partial \dot y}{\partial y} &
# \frac{\partial \dot y}{\partial \theta} \\
# \frac{\partial \dot{\theta}}{\partial x} &
# \frac{\partial \dot{\theta}}{\partial y} &
# \frac{\partial \dot{\theta}}{\partial \theta}
# \end{bmatrix}
# $$
#
# When we calculate these we get
#
# $$\mathbf F = \begin{bmatrix}
# 1 & 0 & -R\cos(\theta) + R\cos(\theta+\beta) \\
# 0 & 1 & -R\sin(\theta) + R\sin(\theta+\beta) \\
# 0 & 0 & 1
# \end{bmatrix}$$
#
# We can double check our work with SymPy.
# +
import sympy
from sympy.abc import alpha, x, y, v, w, R, theta
from sympy import symbols, Matrix
sympy.init_printing(use_latex="mathjax", fontsize='16pt')
time = symbols('t')
d = v*time
beta = (d/w)*sympy.tan(alpha)
r = w/sympy.tan(alpha)
fxu = Matrix([[x-r*sympy.sin(theta) + r*sympy.sin(theta+beta)],
[y+r*sympy.cos(theta)- r*sympy.cos(theta+beta)],
[theta+beta]])
F = fxu.jacobian(Matrix([x, y, theta]))
F
# -
# That looks a bit complicated. We can use SymPy to substitute terms:
# reduce common expressions
B, R = symbols('beta, R')
F = F.subs((d/w)*sympy.tan(alpha), B)
F.subs(w/sympy.tan(alpha), R)
# This form verifies that the computation of the Jacobian is correct.
#
# Now we can turn our attention to the noise. Here, the noise is in our control input, so it is in *control space*. In other words, we command a specific velocity and steering angle, but we need to convert that into errors in $x, y, \theta$. In a real system this might vary depending on velocity, so it will need to be recomputed for every prediction. I will choose this as the noise model; for a real robot you will need to choose a model that accurately depicts the error in your system.
#
# $$\mathbf{M} = \begin{bmatrix}\sigma_{vel}^2 & 0 \\ 0 & \sigma_\alpha^2\end{bmatrix}$$
#
# If this was a linear problem we would convert from control space to state space using the by now familiar $\mathbf{FMF}^\mathsf T$ form. Since our motion model is nonlinear we do not try to find a closed form solution to this, but instead linearize it with a Jacobian which we will name $\mathbf{V}$.
#
# $$\mathbf{V} = \frac{\partial f(x, u)}{\partial u} \begin{bmatrix}
# \frac{\partial \dot x}{\partial v} & \frac{\partial \dot x}{\partial \alpha} \\
# \frac{\partial \dot y}{\partial v} & \frac{\partial \dot y}{\partial \alpha} \\
# \frac{\partial \dot{\theta}}{\partial v} & \frac{\partial \dot{\theta}}{\partial \alpha}
# \end{bmatrix}$$
#
# These partial derivatives become very difficult to work with. Let's compute them with SymPy.
V = fxu.jacobian(Matrix([v, alpha]))
V = V.subs(sympy.tan(alpha)/w, 1/R)
V = V.subs(time*v/R, B)
V = V.subs(time*v, 'd')
V
# This should give you an appreciation of how quickly the EKF become mathematically intractable.
#
# This gives us the final form of our prediction equations:
#
# $$\begin{aligned}
# \mathbf{\overline x} &= \mathbf x +
# \begin{bmatrix}- R\sin(\theta) + R\sin(\theta + \beta) \\
# R\cos(\theta) - R\cos(\theta + \beta) \\
# \beta\end{bmatrix}\\
# \mathbf{\overline P} &=\mathbf{FPF}^{\mathsf T} + \mathbf{VMV}^{\mathsf T}
# \end{aligned}$$
#
# This form of linearization is not the only way to predict $\mathbf x$. For example, we could use a numerical integration technique such as *Runge Kutta* to compute the movement
# of the robot. This will be required if the time step is relatively large. Things are not as cut and dried with the EKF as for the Kalman filter. For a real problem you have to carefully model your system with differential equations and then determine the most appropriate way to solve that system. The correct approach depends on the accuracy you require, how nonlinear the equations are, your processor budget, and numerical stability concerns.
# ### Design the Measurement Model
#
# The robot's sensor provides a noisy bearing and range measurement to multiple known locations in the landscape. The measurement model must convert the state $\begin{bmatrix}x & y&\theta\end{bmatrix}^\mathsf T$ into a range and bearing to the landmark. If $\mathbf p$
# is the position of a landmark, the range $r$ is
#
# $$r = \sqrt{(p_x - x)^2 + (p_y - y)^2}$$
#
# The sensor provides bearing relative to the orientation of the robot, so we must subtract the robot's orientation from the bearing to get the sensor reading, like so:
#
# $$\phi = \arctan(\frac{p_y - y}{p_x - x}) - \theta$$
#
#
# Thus our measurement model $h$ is
#
#
# $$\begin{aligned}
# \mathbf z& = h(\bar{\mathbf x}, \mathbf p) &+ \mathcal{N}(0, R)\\
# &= \begin{bmatrix}
# \sqrt{(p_x - x)^2 + (p_y - y)^2} \\
# \arctan(\frac{p_y - y}{p_x - x}) - \theta
# \end{bmatrix} &+ \mathcal{N}(0, R)
# \end{aligned}$$
#
# This is clearly nonlinear, so we need linearize $h$ at $\mathbf x$ by taking its Jacobian. We compute that with SymPy below.
px, py = symbols('p_x, p_y')
z = Matrix([[sympy.sqrt((px-x)**2 + (py-y)**2)],
[sympy.atan2(py-y, px-x) - theta]])
z.jacobian(Matrix([x, y, theta]))
# Now we need to write that as a Python function. For example we might write:
# +
from math import sqrt
def H_of(x, landmark_pos):
""" compute Jacobian of H matrix where h(x) computes
the range and bearing to a landmark for state x """
px = landmark_pos[0]
py = landmark_pos[1]
hyp = (px - x[0, 0])**2 + (py - x[1, 0])**2
dist = sqrt(hyp)
H = array(
[[-(px - x[0, 0]) / dist, -(py - x[1, 0]) / dist, 0],
[ (py - x[1, 0]) / hyp, -(px - x[0, 0]) / hyp, -1]])
return H
# -
# We also need to define a function that converts the system state into a measurement.
# +
from math import atan2
def Hx(x, landmark_pos):
""" takes a state variable and returns the measurement
that would correspond to that state.
"""
px = landmark_pos[0]
py = landmark_pos[1]
dist = sqrt((px - x[0, 0])**2 + (py - x[1, 0])**2)
Hx = array([[dist],
[atan2(py - x[1, 0], px - x[0, 0]) - x[2, 0]]])
return Hx
# -
# ### Design Measurement Noise
#
# It is reasonable to assume that the noise of the range and bearing measurements are independent, hence
#
# $$\mathbf R=\begin{bmatrix}\sigma_{range}^2 & 0 \\ 0 & \sigma_{bearing}^2\end{bmatrix}$$
# ### Implementation
#
# We will use `FilterPy`'s `ExtendedKalmanFilter` class to implement the filter. Its `predict()` method uses the standard linear equations for the process model. Our's is nonlinear, so we will have to override `predict()` with our own implementation. I'll want to also use this class to simulate the robot, so I'll add a method `move()` that computes the position of the robot which both `predict()` and my simulation can call.
#
# The matrices for the prediction step are quite large. While writing this code I made several errors before I finally got it working. I only found my errors by using SymPy's `evalf` function. `evalf` evaluates a SymPy `Matrix` with specific values for the variables. I decided to demonstrate this technique to you, and used `evalf` in the Kalman filter code. You'll need to understand a couple of points.
#
# First, `evalf` uses a dictionary to specify the values. For example, if your matrix contains an `x` and `y`, you can write
#
# ```python
# M.evalf(subs={x:3, y:17})
# ```
#
# to evaluate the matrix for `x=3` and `y=17`.
#
# Second, `evalf` returns a `sympy.Matrix` object. Use `numpy.array(M).astype(float)` to convert it to a NumPy array. `numpy.array(M)` creates an array of type `object`, which is not what you want.
#
# Here is the code for the EKF:
from filterpy.kalman import ExtendedKalmanFilter as EKF
from numpy import dot, array, sqrt
class RobotEKF(EKF):
def __init__(self, dt, wheelbase, std_vel, std_steer):
EKF.__init__(self, 3, 2, 2)
self.dt = dt
self.wheelbase = wheelbase
self.std_vel = std_vel
self.std_steer = std_steer
a, x, y, v, w, theta, time = symbols(
'a, x, y, v, w, theta, t')
d = v*time
beta = (d/w)*sympy.tan(a)
r = w/sympy.tan(a)
self.fxu = Matrix(
[[x-r*sympy.sin(theta)+r*sympy.sin(theta+beta)],
[y+r*sympy.cos(theta)-r*sympy.cos(theta+beta)],
[theta+beta]])
self.F_j = self.fxu.jacobian(Matrix([x, y, theta]))
self.V_j = self.fxu.jacobian(Matrix([v, a]))
# save dictionary and it's variables for later use
self.subs = {x: 0, y: 0, v:0, a:0,
time:dt, w:wheelbase, theta:0}
self.x_x, self.x_y, = x, y
self.v, self.a, self.theta = v, a, theta
def predict(self, u=0):
self.x = self.move(self.x, u, self.dt)
self.subs[self.theta] = self.x[2, 0]
self.subs[self.v] = u[0]
self.subs[self.a] = u[1]
F = array(self.F_j.evalf(subs=self.subs)).astype(float)
V = array(self.V_j.evalf(subs=self.subs)).astype(float)
# covariance of motion noise in control space
M = array([[self.std_vel*u[0]**2, 0],
[0, self.std_steer**2]])
self.P = dot(F, self.P).dot(F.T) + dot(V, M).dot(V.T)
def move(self, x, u, dt):
hdg = x[2, 0]
vel = u[0]
steering_angle = u[1]
dist = vel * dt
if abs(steering_angle) > 0.001: # is robot turning?
beta = (dist / self.wheelbase) * tan(steering_angle)
r = self.wheelbase / tan(steering_angle) # radius
dx = np.array([[-r*sin(hdg) + r*sin(hdg + beta)],
[r*cos(hdg) - r*cos(hdg + beta)],
[beta]])
else: # moving in straight line
dx = np.array([[dist*cos(hdg)],
[dist*sin(hdg)],
[0]])
return x + dx
# Now we have another issue to handle. The residual is notionally computed as $y = z - h(x)$ but this will not work because our measurement contains an angle in it. Suppose z has a bearing of $1^\circ$ and $h(x)$ has a bearing of $359^\circ$. Naively subtracting them would yield a angular difference of $-358^\circ$, whereas the correct value is $2^\circ$. We have to write code to correctly compute the bearing residual.
def residual(a, b):
""" compute residual (a-b) between measurements containing
[range, bearing]. Bearing is normalized to [-pi, pi)"""
y = a - b
y[1] = y[1] % (2 * np.pi) # force in range [0, 2 pi)
if y[1] > np.pi: # move to [-pi, pi)
y[1] -= 2 * np.pi
return y
# The rest of the code runs the simulation and plots the results, and shouldn't need too much comment by now. I create a variable `landmarks` that contains the landmark coordinates. I update the simulated robot position 10 times a second, but run the EKF only once per second. This is for two reasons. First, we are not using Runge Kutta to integrate the differental equations of motion, so a narrow time step allows our simulation to be more accurate. Second, it is fairly normal in embedded systems to have limited processing speed. This forces you to run your Kalman filter only as frequently as absolutely needed.
# +
from filterpy.stats import plot_covariance_ellipse
from math import sqrt, tan, cos, sin, atan2
import matplotlib.pyplot as plt
dt = 1.0
def z_landmark(lmark, sim_pos, std_rng, std_brg):
x, y = sim_pos[0, 0], sim_pos[1, 0]
d = np.sqrt((lmark[0] - x)**2 + (lmark[1] - y)**2)
a = atan2(lmark[1] - y, lmark[0] - x) - sim_pos[2, 0]
z = np.array([[d + randn()*std_rng],
[a + randn()*std_brg]])
return z
def ekf_update(ekf, z, landmark):
ekf.update(z, HJacobian=H_of, Hx=Hx,
residual=residual,
args=(landmark), hx_args=(landmark))
def run_localization(landmarks, std_vel, std_steer,
std_range, std_bearing,
step=10, ellipse_step=20, ylim=None):
ekf = RobotEKF(dt, wheelbase=0.5, std_vel=std_vel,
std_steer=std_steer)
ekf.x = array([[2, 6, .3]]).T # x, y, steer angle
ekf.P = np.diag([.1, .1, .1])
ekf.R = np.diag([std_range**2, std_bearing**2])
sim_pos = ekf.x.copy() # simulated position
# steering command (vel, steering angle radians)
u = array([1.1, .01])
plt.scatter(landmarks[:, 0], landmarks[:, 1],
marker='s', s=60)
track = []
for i in range(200):
sim_pos = ekf.move(sim_pos, u, dt/10.) # simulate robot
track.append(sim_pos)
if i % step == 0:
ekf.predict(u=u)
if i % ellipse_step == 0:
plot_covariance_ellipse(
(ekf.x[0,0], ekf.x[1,0]), ekf.P[0:2, 0:2],
std=6, facecolor='k', alpha=0.3)
x, y = sim_pos[0, 0], sim_pos[1, 0]
for lmark in landmarks:
z = z_landmark(lmark, sim_pos,
std_range, std_bearing)
ekf_update(ekf, z, lmark)
if i % ellipse_step == 0:
plot_covariance_ellipse(
(ekf.x[0,0], ekf.x[1,0]), ekf.P[0:2, 0:2],
std=6, facecolor='g', alpha=0.8)
track = np.array(track)
plt.plot(track[:, 0], track[:,1], color='k', lw=2)
plt.axis('equal')
plt.title("EKF Robot localization")
if ylim is not None: plt.ylim(*ylim)
plt.show()
return ekf
# +
landmarks = array([[5, 10], [10, 5], [15, 15]])
ekf = run_localization(
landmarks, std_vel=0.1, std_steer=np.radians(1),
std_range=0.3, std_bearing=0.1)
print('Final P:', ekf.P.diagonal())
# -
# I have plotted the landmarks as solid squares. The path of the robot is drawn with a black line. The covariance ellipses for the predict step are light gray, and the covariances of the update are shown in green. To make them visible at this scale I have set the ellipse boundary at 6$\sigma$.
#
# We can see that there is a lot of uncertainty added by our motion model, and that most of the error in in the direction of motion. We determine that from the shape of the blue ellipses. After a few steps we can see that the filter incorporates the landmark measurements and the errors improve.
#
# I used the same initial conditions and landmark locations in the UKF chapter. The UKF achieves much better accuracy in terms of the error ellipse. Both perform roughly as well as far as their estimate for $\mathbf x$ is concerned.
#
# Now lets add another landmark.
# +
landmarks = array([[5, 10], [10, 5], [15, 15], [20, 5]])
ekf = run_localization(
landmarks, std_vel=0.1, std_steer=np.radians(1),
std_range=0.3, std_bearing=0.1)
plt.show()
print('Final P:', ekf.P.diagonal())
# -
# The uncertainly in the estimates near the end of the track are smaller. We can see the effect that multiple landmarks have on our uncertainty by only using the first two landmarks.
ekf = run_localization(
landmarks[0:2], std_vel=1.e-10, std_steer=1.e-10,
std_range=1.4, std_bearing=.05)
print('Final P:', ekf.P.diagonal())
# The estimate quickly diverges from the robot's path after passing the landmarks. The covariance also grows quickly. Let's see what happens with only one landmark:
ekf = run_localization(
landmarks[0:1], std_vel=1.e-10, std_steer=1.e-10,
std_range=1.4, std_bearing=.05)
print('Final P:', ekf.P.diagonal())
# As you probably suspected, one landmark produces a very bad result. Conversely, a large number of landmarks allows us to make very accurate estimates.
# +
landmarks = array([[5, 10], [10, 5], [15, 15], [20, 5], [15, 10],
[10,14], [23, 14], [25, 20], [10, 20]])
ekf = run_localization(
landmarks, std_vel=0.1, std_steer=np.radians(1),
std_range=0.3, std_bearing=0.1, ylim=(0, 21))
print('Final P:', ekf.P.diagonal())
# -
# ### Discussion
#
# I said that this was a real problem, and in some ways it is. I've seen alternative presentations that used robot motion models that led to simpler Jacobians. On the other hand, my model of the movement is also simplistic in several ways. First, it uses a bicycle model. A real car has two sets of tires, and each travels on a different radius. The wheels do not grip the surface perfectly. I also assumed that the robot responds instantaneously to the control input. <NAME> writes in *Probabilistic Robots* that simplified models are justified because the filters perform well when used to track real vehicles. The lesson here is that while you have to have a reasonably accurate nonlinear model, it does not need to be perfect to operate well. As a designer you will need to balance the fidelity of your model with the difficulty of the math and the CPU time required to perform the linear algebra.
#
# Another way in which this problem was simplistic is that we assumed that we knew the correspondance between the landmarks and measurements. But suppose we are using radar - how would we know that a specific signal return corresponded to a specific building in the local scene? This question hints at SLAM algorithms - simultaneous localization and mapping. SLAM is not the point of this book, so I will not elaborate on this topic.
# ## UKF vs EKF
#
#
# In the last chapter I used the UKF to solve this problem. The difference in implementation should be very clear. Computing the Jacobians for the state and measurement models was not trivial despite a rudimentary motion model. A different problem could result in a Jacobian which is difficult or impossible to derive analytically. In contrast, the UKF only requires you to provide a function that computes the system motion model and another for the measurement model.
#
# There are many cases where the Jacobian cannot be found analytically. The details are beyond the scope of this book, but you will have to use numerical methods to compute the Jacobian. That undertaking is not trivial, and you will spend a significant portion of a master's degree at a STEM school learning techniques to handle such situations. Even then you'll likely only be able to solve problems related to your field - an aeronautical engineer learns a lot about Navier Stokes equations, but not much about modelling chemical reaction rates.
#
# So, UKFs are easy. Are they accurate? In practice they often perform better than the EKF. You can find plenty of research papers that prove that the UKF outperforms the EKF in various problem domains. It's not hard to understand why this would be true. The EKF works by linearizing the system model and measurement model at a single point, and the UKF uses $2n+1$ points.
#
# Let's look at a specific example. Take $f(x) = x^3$ and pass a Gaussian distribution through it. I will compute an accurate answer using a monte carlo simulation. I generate 50,000 points randomly distributed according to the Gaussian, pass each through $f(x)$, then compute the mean and variance of the result.
#
# The EKF linearizes the function by taking the derivative to find the slope at the evaluation point $x$. This slope becomes the linear function that we use to transform the Gaussian. Here is a plot of that.
import nonlinear_plots
nonlinear_plots.plot_ekf_vs_mc()
# The EKF computation is rather inaccurate. In contrast, here is the performance of the UKF:
nonlinear_plots.plot_ukf_vs_mc(alpha=0.001, beta=3., kappa=1.)
# Here we can see that the computation of the UKF's mean is accurate to 2 decimal places. The standard deviation is slightly off, but you can also fine tune how the UKF computes the distribution by using the $\alpha$, $\beta$, and $\gamma$ parameters for generating the sigma points. Here I used $\alpha=0.001$, $\beta=3$, and $\gamma=1$. Feel free to modify them to see the result. You should be able to get better results than I did. However, avoid over-tuning the UKF for a specific test. It may perform better for your test case, but worse in general.
| 11-Extended-Kalman-Filters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ensegment: default program
from default import *
# ## Documentation
#
# Write some beautiful documentation of your program here.
Pw = Pdist(data=datafile("data/count_1w.txt"))
segmenter = Segment(Pw)
with open("data/input/dev.txt") as f:
for line in f:
print(" ".join(segmenter.segment(line.strip())))
# ## Analysis
#
# Do some analysis of the results. What ideas did you try? What worked and what did not?
| ensegment/default.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# The %... is an iPython thing, and is not part of the Python language.
# In this case we're just telling the plotting library to draw things on
# the notebook, instead of on a separate window.
# %matplotlib inline
# See all the "as ..." contructs? They're just aliasing the package names.
# That way we can call methods like plt.plot() instead of matplotlib.pyplot.plot().
import numpy as np
import scipy as sp
import matplotlib as mpl
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import pandas as pd
import time
pd.set_option('display.width', 500)
pd.set_option('display.max_columns', 100)
pd.set_option('display.max_rows', 40)
pd.set_option('display.notebook_repr_html', True)
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("notebook")
from bs4 import BeautifulSoup
from collections import OrderedDict # provides the ordered dictionary
import re # for regular expressions used below
import urllib # to read from URLs
import json
import networkx as nx # network analysis
from networkx.readwrite import json_graph
import itertools
import os.path
from datetime import datetime # for time measurement
import sys
import os
import pickle
import subprocess as subp
import gzip
import math
import codecs
from jellyfish import jaro_distance, jaro_winkler, hamming_distance, levenshtein_distance, metaphone, nysiis, soundex
import scipy.cluster.hierarchy as scipycluster
from sklearn.feature_extraction.text import TfidfVectorizer
from skimage import io, exposure
from scipy.spatial import distance
# import the k-means algorithm
from sklearn.cluster import KMeans, MiniBatchKMeans
from sklearn.metrics import pairwise_distances_argmin,pairwise_distances_argmin_min, pairwise_distances
from sklearn.metrics.pairwise import euclidean_distances
def printLog(text):
now=str(datetime.now())
print("["+now+"]\t"+text)
# forces to output the result of the print command immediately, see: http://stackoverflow.com/questions/230751/how-to-flush-output-of-python-print
sys.stdout.flush()
def pickleCompress(fileName,pickledObject):
printLog("Pickling to '%s'" %fileName)
f = gzip.open(fileName,'wb')
pickle.dump(pickledObject,f)
f.close()
printLog("Pickling done.")
def pickleDecompress(fileName):
#restore the object
printLog("Depickling from '%s'" %fileName)
f = gzip.open(fileName,'rb')
pickledObject = pickle.load(f)
f.close()
printLog("Depickling done.")
return pickledObject
# -
# !pip install jellyfish
# +
words = u'Berlin Balin Cölln Köln Cologne Zürich Zurich Bern'.split()
print("Number of words: %i" % len(words))
for i,val in enumerate(words):
print(str(i)+":\t "+str(val.encode('utf-8')))
r=np.triu_indices(n=len(words), k=1)
r
# +
def d_demo(coord):
i, j = coord
# the distance fix we have learnt about before...
return 1-jaro_distance(words[i], words[j])
# http://docs.scipy.org/doc/numpy-1.10.1/reference/generated/numpy.set_printoptions.html
np.set_printoptions(precision=4)
# axis (3rd parameter): 0= along y axis, 1= along x axis
r2=np.apply_along_axis(d_demo, 0, r)
r2
# +
Z=scipycluster.linkage(r2,method='single')
plt.figure(figsize=(25, 10))
#plt.title('Hierarchical Clustering Dendrogram')
plt.xlabel('Term')
plt.ylabel('Distance')
scipycluster.dendrogram(
Z,
leaf_rotation=90., # rotates the x axis labels
leaf_font_size=16., # font size for the x axis labels
labels=words
)
plt.show()
# +
Z=scipycluster.linkage(r2,method='average')
plt.figure(figsize=(25, 10))
#plt.title('Hierarchical Clustering Dendrogram')
plt.xlabel('Term')
plt.ylabel('Distance')
scipycluster.dendrogram(
Z,
leaf_rotation=90., # rotates the x axis labels
leaf_font_size=16., # font size for the x axis labels
labels=words
)
plt.show()
# +
Z=scipycluster.linkage(r2,method='ward')
plt.figure(figsize=(25, 10))
#plt.title('Hierarchical Clustering Dendrogram')
plt.xlabel('Term')
plt.ylabel('Distance')
scipycluster.dendrogram(
Z,
leaf_rotation=90., # rotates the x axis labels
leaf_font_size=16., # font size for the x axis labels
labels=words
)
plt.show()
# +
words = u'Berlin Balin Cölln Köln Cologne Zürich Zurich Bern'.split()
words2=list(map(metaphone,words))
term_labels=list(zip(words2,words))
words=words2
print("Number of words: %i" % len(words))
for i,val in enumerate(words):
print(str(i)+":\t "+str(val.encode('utf-8')))
r=np.triu_indices(n=len(words), k=1)
r
r2=np.apply_along_axis(d_demo, 0, r)
r2
Z=scipycluster.linkage(r2,method='single')
plt.figure(figsize=(25, 10))
#plt.title('Hierarchical Clustering Dendrogram')
plt.xlabel('Term (in Metaphone)')
plt.ylabel('Distance')
scipycluster.dendrogram(
Z,
leaf_rotation=90., # rotates the x axis labels
leaf_font_size=16., # font size for the x axis labels
labels=term_labels
)
plt.show()
# -
| clustering_samples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="MSnsTgZLKO72"
# # Practice Loading Datasets
#
# This assignment is purposely semi-open-ended you will be asked to load datasets both from github and also from CSV files from the [UC Irvine Machine Learning Repository](https://archive.ics.uci.edu/ml/index.php).
#
# Remember that the UCI datasets may not have a file type of `.csv` so it's important that you learn as much as you can about the dataset before you try and load it. See if you can look at the raw text of the file either locally, on github, using the `!curl` shell command, or in some other way before you try and read it in as a dataframe, this will help you catch what would otherwise be unforseen problems.
#
# + [markdown] colab_type="text" id="xn07qH1u-QFt"
#
# + [markdown] colab_type="text" id="156P6ndeKojO"
# ## 1) Load a dataset from Github (via its *RAW* URL)
#
# Pick a dataset from the following repository and load it into Google Colab. Make sure that the headers are what you would expect and check to see if missing values have been encoded as NaN values:
#
# <https://github.com/ryanleeallred/datasets>
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="NJdISe69ZT7E" outputId="51d772b5-f548-4479-f1d8-30a2fe30e479"
# Load the data
import pandas as pd
url = 'https://raw.githubusercontent.com/ryanleeallred/datasets/master/heart.csv'
df = pd.read_csv(url)
# Verify we have something going on
df.head()
# Looks like we got the headers correct.
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="Gxe3yIYSmw2l" outputId="d3069af4-f7e5-4934-db89-63e662157d10"
# we got 303 coloumns without the header.
df.count()
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="SvJDj2qhn7TF" outputId="b8d00c4c-b75b-4ab1-ba32-bc18526523d8"
# Does it have missing values?
df.isna().sum()
# there are no missing values.
# + [markdown] colab_type="text" id="-gFnZR6iLLPY"
# ## 2) Load a dataset from your local machine
# Download a dataset from the [UC Irvine Machine Learning Repository](https://archive.ics.uci.edu/ml/index.php) and then upload the file to Google Colab either using the files tab in the left-hand sidebar or by importing `files` from `google.colab` The following link will be a useful resource if you can't remember the syntax: <https://towardsdatascience.com/3-ways-to-load-csv-files-into-colab-7c14fcbdcb92>
#
# While you are free to try and load any dataset from the UCI repository, I strongly suggest starting with one of the most popular datasets like those that are featured on the right-hand side of the home page.
#
# Some datasets on UCI will have challenges associated with importing them far beyond what we have exposed you to in class today, so if you run into a dataset that you don't know how to deal with, struggle with it for a little bit, but ultimately feel free to simply choose a different one.
#
# - Make sure that your file has correct headers, and the same number of rows and columns as is specified on the UCI page. If your dataset doesn't have headers use the parameters of the `read_csv` function to add them. Likewise make sure that missing values are encoded as `NaN`.
# + colab={"base_uri": "https://localhost:8080/", "height": 75, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "headers": [["content-type", "application/javascript"]], "ok": true, "status": 200, "status_text": ""}}} colab_type="code" id="qUmwX-ZoM9cq" outputId="dcc90e66-6c81-4e7e-c5f1-f8f11b88d525"
# TODO your work here!
# And note you should write comments, descriptions, and add new
# code and text blocks as needed
from google.colab import files
uploaded = files.upload()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="d0nCMLSaxGUT" outputId="c3b4e329-7d06-49c3-c90e-71fd532275c8"
df = pd.read_csv('iris.data')
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="7MA_TUZ6Lpms" outputId="5a6c658b-7045-4bde-c75f-3dbed308b546"
df.count()
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="Q_Dum7EaLub9" outputId="4391dcfb-25ff-4024-e92c-dac2169a426c"
df.isna().sum()
# + [markdown] colab_type="text" id="mq_aQjxlM-u5"
# ## 3) Load a dataset from UCI using `!wget`
#
# "Shell Out" and try loading a file directly into your google colab's memory using the `!wget` command and then read it in with `read_csv`.
#
# With this file we'll do a bit more to it.
#
# - Read it in, fix any problems with the header as make sure missing values are encoded as `NaN`.
# - Use the `.fillna()` method to fill any missing values.
# - [.fillna() documentation](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.fillna.html)
# - Create one of each of the following plots using the Pandas plotting functionality:
# - Scatterplot
# - Histogram
# - Density Plot
#
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="-Xc-ZjA98O8F" outputId="8633065e-73c9-4042-b36a-87626da2f018"
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data
# + colab={} colab_type="code" id="bUZswS1kSAls"
column_headers = ['age', 'workclass', 'fnlwgt', 'education', 'education-num', 'marital-status',
'occupation', 'relationship', 'race', 'sex', 'capital-gain', 'capital-loss',
'hours-per-week', 'native-country', 'income']
df = pd.read_csv('adult.data', names=column_headers, na_values=' ?')
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="smHfCCkGUhPw" outputId="aa737cce-7d57-4ae4-9a06-6fa80391cde6"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" id="-1DL7RwsUrLB" outputId="fbb627ee-81fd-4fc6-9f2f-5b1571a8d60f"
df.count()
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" id="jDKknm2sUu3K" outputId="78353140-6c2d-4d95-d031-80a7d32c7c31"
df.isna().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="__T11itcVoFw" outputId="319f50b4-b189-4a45-9fed-be42d5016f43"
df.fillna('unknown')
# + colab={"base_uri": "https://localhost:8080/", "height": 297} colab_type="code" id="30zG3kYoZVaN" outputId="e01103f3-e79c-4f62-a86f-79f196f0280f"
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" id="rsthvlLMa39_" outputId="868a057b-120c-4c46-9f43-4c68f287a9ba"
# Histogram
df['age'].hist()
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="ww0REBcycCXP" outputId="004308b5-5a8a-4abb-f9a2-d59b64d416c0"
# Scatter Plot
df.plot.scatter('age','hours-per-week');
# + colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" id="_NdYmROncMrp" outputId="1e166d65-47ce-4b76-fe0b-c07a3e6c7cc4"
# Density Plot
df['age'].plot.density()
# + [markdown] colab_type="text" id="MZCxTwKuReV9"
# ## Stretch Goals - Other types and sources of data
#
# Not all data comes in a nice single file - for example, image classification involves handling lots of image files. You still will probably want labels for them, so you may have tabular data in addition to the image blobs - and the images may be reduced in resolution and even fit in a regular csv as a bunch of numbers.
#
# If you're interested in natural language processing and analyzing text, that is another example where, while it can be put in a csv, you may end up loading much larger raw data and generating features that can then be thought of in a more standard tabular fashion.
#
# Overall you will in the course of learning data science deal with loading data in a variety of ways. Another common way to get data is from a database - most modern applications are backed by one or more databases, which you can query to get data to analyze. We'll cover this more in our data engineering unit.
#
# How does data get in the database? Most applications generate logs - text files with lots and lots of records of each use of the application. Databases are often populated based on these files, but in some situations you may directly analyze log files. The usual way to do this is with command line (Unix) tools - command lines are intimidating, so don't expect to learn them all at once, but depending on your interests it can be useful to practice.
#
# One last major source of data is APIs: https://github.com/toddmotto/public-apis
#
# API stands for Application Programming Interface, and while originally meant e.g. the way an application interfaced with the GUI or other aspects of an operating system, now it largely refers to online services that let you query and retrieve data. You can essentially think of most of them as "somebody else's database" - you have (usually limited) access.
#
# *Stretch goal* - research one of the above extended forms of data/data loading. See if you can get a basic example working in a notebook. Image, text, or (public) APIs are probably more tractable - databases are interesting, but there aren't many publicly accessible and they require a great deal of setup.
# +
# https://www.dataquest.io/blog/python-api-tutorial/
import json
import request
response = requests.get("")
# + colab={} colab_type="code" id="f4QP6--JBXNK"
# HOW TOs:
# https://www.digitalocean.com/community/tutorials/how-to-use-web-apis-in-python-3
# https://calendarific.com/api-documentation
# https://calendarific.com
import json
import requests
response = requests.get("https://calendarific.com/api/v2/holidays?api_key=<KEY>")
print(response.status_code)
# +
# parameters = {"country": US, "year": 2019}
# had to append &country=US&year=2019 below
response = requests.get("https://calendarific.com/api/v2/holidays?api_key=fa5549d6ddc82939198f033b1cdddc1d7e3ce19c&country=US&year=2019")
print(response.content)
# +
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_json.html
import pandas as pd
calender = pd.read_json(response.content)
# -
calender.head()
calender.info
# This is what the response looks like
{
"meta": {
"code": 200
},
"response": {
"holidays": [
{
"name": "Name of holiday goes here",
"description": "Description of holiday goes here",
"date": {
"iso": "2018-12-31",
"datetime": {
"year": 2018,
"month": 12,
"day": 31
}
},
"type": [
"Type of Observance goes here"
]
}
]
}
}
| module2-loadingdata/JM_LS_DS_112_Loading_Data_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="FeuofmT9R5xX"
# Reference: https://www.kaggle.com/datasets/shivamb/netflix-shows/code
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" id="GL1MqKK-S7H9"
# import librairies
import networkx as nx
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import math as math
import time
plt.style.use('seaborn')
plt.rcParams['figure.figsize'] = [14,14]
# + colab={"base_uri": "https://localhost:8080/"} id="Nb50CG7NTGRE" outputId="a05fd2d2-b331-4a29-d9d0-3035e29753b9"
from google.colab import drive
drive.mount('/gdrive')
# %cd /gdrive
# + [markdown] id="yqlVS0p2S7ID"
# # Load the data
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="rpyT4OYCS7IE" outputId="a54ef4f8-f8ee-4cf3-b339-d2eae41a9b79"
# load the data
df = pd.read_csv('MyDrive/IDS_Datasets/netflix_titles.csv')
# convert to datetime
df["date_added"] = pd.to_datetime(df['date_added'])
df['year'] = df['date_added'].dt.year
df['month'] = df['date_added'].dt.month
df['day'] = df['date_added'].dt.day
# convert columns "director, listed_in, cast and country" in columns that contain a real list
# the strip function is applied on the elements
# if the value is NaN, the new column contains a empty list []
df['directors'] = df['director'].apply(lambda l: [] if pd.isna(l) else [i.strip() for i in l.split(",")])
df['categories'] = df['listed_in'].apply(lambda l: [] if pd.isna(l) else [i.strip() for i in l.split(",")])
df['actors'] = df['cast'].apply(lambda l: [] if pd.isna(l) else [i.strip() for i in l.split(",")])
df['countries'] = df['country'].apply(lambda l: [] if pd.isna(l) else [i.strip() for i in l.split(",")])
df.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="v_vnm_hhS7IG" outputId="2a7599aa-2af0-4535-8a24-031fd0509a2c"
print(df.shape)
# + [markdown] id="uhEr6-RpS7IH"
# # KMeans clustering with TF-IDF
# + colab={"base_uri": "https://localhost:8080/"} id="cuKbtr08S7IH" outputId="3ea95208-045c-43db-a45c-955490918610"
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import linear_kernel
from sklearn.cluster import MiniBatchKMeans
# Build the tfidf matrix with the descriptions
start_time = time.time()
text_content = df['description']
vector = TfidfVectorizer(max_df=0.4, # drop words that occur in more than X percent of documents
min_df=1, # only use words that appear at least X times
stop_words='english', # remove stop words
lowercase=True, # Convert everything to lower case
use_idf=True, # Use idf
norm=u'l2', # Normalization
smooth_idf=True # Prevents divide-by-zero errors
)
tfidf = vector.fit_transform(df['description'].apply(lambda x: np.str_(x)) )
# Clustering Kmeans
k = 200
kmeans = MiniBatchKMeans(n_clusters = k)
kmeans.fit(tfidf)
centers = kmeans.cluster_centers_.argsort()[:,::-1]
terms = vector.get_feature_names()
# print the centers of the clusters
# for i in range(0,k):
# word_list=[]
# print("cluster%d:"% i)
# for j in centers[i,:10]:
# word_list.append(terms[j])
# print(word_list)
request_transform = vector.transform(df['description'].apply(lambda x: np.str_(x)))
# new column cluster based on the description
df['cluster'] = kmeans.predict(request_transform)
df['cluster'].value_counts().head()
# + [markdown] id="MZ1ogQW0S7II"
# <div class="alert alert-block alert-warning"><span>«</span>column cluster are not going to be used because clusters are two unbalanced <br/> But tfidf will be used in order to find similar description<span>»</span></div>
# + id="Ir41sn3QS7II"
# Find similar : get the top_n movies with description similar to the target description
def find_similar(tfidf_matrix, index, top_n = 5):
cosine_similarities = linear_kernel(tfidf_matrix[index:index+1], tfidf_matrix).flatten()
related_docs_indices = [i for i in cosine_similarities.argsort()[::-1] if i != index]
return [index for index in related_docs_indices][0:top_n]
# + [markdown] id="EqKqIOq5S7II"
# # Load the graph (undirected graph)
# Nodes are :
# * Movies
# * Person ( actor or director)
# * Categorie
# * Countrie
# * Cluster (description)
# * Sim(title) top 5 similar movies in the sense of the description
#
# Edges are :
# * ACTED_IN : relation between an actor and a movie
# * CAT_IN : relation between a categrie and a movie
# * DIRECTED : relation between a director and a movie
# * COU_IN : relation between a country and a movie
# * DESCRIPTION : relation between a cluster and a movie
# * SIMILARITY in the sense of the description
#
# <span>«</span>so, two movies are not directly connected, but they share persons, categories,clusters and countries<span>»</span>
#
# + colab={"base_uri": "https://localhost:8080/"} id="nGxb9DygS7IJ" outputId="dd76958f-ff81-4a33-a499-bf794da9bbae"
G = nx.Graph(label="MOVIE")
start_time = time.time()
for i, rowi in df.iterrows():
if (i%1000==0):
print(" iter {} -- {} seconds --".format(i,time.time() - start_time))
G.add_node(rowi['title'],key=rowi['show_id'],label="MOVIE",mtype=rowi['type'],rating=rowi['rating'])
# G.add_node(rowi['cluster'],label="CLUSTER")
# G.add_edge(rowi['title'], rowi['cluster'], label="DESCRIPTION")
for element in rowi['actors']:
G.add_node(element,label="PERSON")
G.add_edge(rowi['title'], element, label="ACTED_IN")
for element in rowi['categories']:
G.add_node(element,label="CAT")
G.add_edge(rowi['title'], element, label="CAT_IN")
for element in rowi['directors']:
G.add_node(element,label="PERSON")
G.add_edge(rowi['title'], element, label="DIRECTED")
for element in rowi['countries']:
G.add_node(element,label="COU")
G.add_edge(rowi['title'], element, label="COU_IN")
indices = find_similar(tfidf, i, top_n = 5)
snode="Sim("+rowi['title'][:15].strip()+")"
G.add_node(snode,label="SIMILAR")
G.add_edge(rowi['title'], snode, label="SIMILARITY")
for element in indices:
G.add_edge(snode, df['title'].loc[element], label="SIMILARITY")
print(" finish -- {} seconds --".format(time.time() - start_time))
# + [markdown] id="TJkCscwsS7IJ"
# # To see what's going on,a sub-graph with only two movies ...
# + id="M87OYq44S7IJ"
def get_all_adj_nodes(list_in):
sub_graph=set()
for m in list_in:
sub_graph.add(m)
for e in G.neighbors(m):
sub_graph.add(e)
return list(sub_graph)
def draw_sub_graph(sub_graph):
subgraph = G.subgraph(sub_graph)
colors=[]
for e in subgraph.nodes():
if G.nodes[e]['label']=="MOVIE":
colors.append('blue')
elif G.nodes[e]['label']=="PERSON":
colors.append('red')
elif G.nodes[e]['label']=="CAT":
colors.append('green')
elif G.nodes[e]['label']=="COU":
colors.append('yellow')
elif G.nodes[e]['label']=="SIMILAR":
colors.append('orange')
elif G.nodes[e]['label']=="CLUSTER":
colors.append('orange')
nx.draw(subgraph, with_labels=True, font_weight='bold',node_color=colors)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="BTGnCywTS7IK" outputId="70d43b93-5708-4714-e8d6-cd253aa4b9fa"
list_in=["Ocean's Twelve","Ocean's Thirteen"]
sub_graph = get_all_adj_nodes(list_in)
draw_sub_graph(sub_graph)
# + [markdown] id="SbI0-pNfS7IK"
# # The recommendation function
# <div class="alert alert-block alert-info">
# <li> Explore the neighborhood of the target film <span>→</span> this is a list of actor, director, country, categorie</li>
# <li> Explore the neighborhood of each neighbor <span>→</span> discover the movies that share a node with the target field</li>
# <li> Calcul Adamic Adar measure <span>→</span> final results</li>
# </div>
# + id="FYzhXl9aS7IK"
def get_recommendation(root):
commons_dict = {}
for e in G.neighbors(root):
for e2 in G.neighbors(e):
if e2==root:
continue
if G.nodes[e2]['label']=="MOVIE":
commons = commons_dict.get(e2)
if commons==None:
commons_dict.update({e2 : [e]})
else:
commons.append(e)
commons_dict.update({e2 : commons})
movies=[]
weight=[]
for key, values in commons_dict.items():
w=0.0
for e in values:
w=w+1/math.log(G.degree(e))
movies.append(key)
weight.append(w)
result = pd.Series(data=np.array(weight),index=movies)
result.sort_values(inplace=True,ascending=False)
return result;
# + [markdown] id="5w6iq5QwS7IK"
# # Let's test it ...
# + colab={"base_uri": "https://localhost:8080/"} id="1HtySRo0S7IL" outputId="ece1b674-9476-44f0-aae9-ca64c0f8980c"
result = get_recommendation("Ocean's Twelve")
result2 = get_recommendation("Ocean's Thirteen")
result3 = get_recommendation("The Devil Inside")
result4 = get_recommendation("Stranger Things")
print("*"*40+"\n Recommendation for 'Ocean's Twelve'\n"+"*"*40)
print(result.head())
print("*"*40+"\n Recommendation for 'Ocean's Thirteen'\n"+"*"*40)
print(result2.head())
print("*"*40+"\n Recommendation for 'Belmonte'\n"+"*"*40)
print(result3.head())
print("*"*40+"\n Recommendation for 'Stranger Things'\n"+"*"*40)
print(result4.head())
# + [markdown] id="KACaPjXlS7IL"
# # Draw top recommendations, to see the common nodes
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="5Y2NmusBS7IL" outputId="58f0eb79-df7c-4a52-ee7d-14bb59c4403b"
reco=list(result.index[:4].values)
reco.extend(["Ocean's Twelve"])
sub_graph = get_all_adj_nodes(reco)
draw_sub_graph(sub_graph)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="xCpJu2GCS7IL" outputId="c72a1b74-12d4-4dd0-c1d2-52d33805044d"
reco=list(result4.index[:4].values)
reco.extend(["Stranger Things"])
sub_graph = get_all_adj_nodes(reco)
draw_sub_graph(sub_graph)
| recommendation_engine_with_networkx_Netflix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Missing Data Analysis of CrossFit Open 2019 Dataset
# The full Open 2019 dataset with it's 72 features and over 338k observations contains a lot missing data. Since most machine learning algorithms can not handle missing data this notebook analyses the amount and structure of missing values. It concludes with recommended actions for the data preparation.
# +
# import relevant libraries
import pandas as pd
import numpy as np
import missingno as msno
import warnings
# set options
warnings.filterwarnings('ignore')
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
# -
# read clean Open 2019 dataset and drop unnamed column
df = pd.read_csv('./data/19_clean.csv')
df = df.loc[:, ~df.columns.str.contains('^Unnamed')]
# # Data Overview
df.shape
df.head()
# # Missing Value Analysis
df.isna().sum()
# Missing values are present in:
# 1. Workout Results:
# * 19.1 - 13,219 (4%)
# * 19.2 - 35,850 (11%) (time 35,923 (11%); tiebreak 43,582 (13%))
# * 19.3 - 46,284 (14%) (time 46,287 (14%); tiebreak 49,047 (15%))
# * 19.4 - 58,225 (17%) (time 58,262 (17%); tiebreak 75,553 (22%))
# * 19.5 - 84,115 (25%)
# 2. Body Measurements:
# * age - 4 (0%)
# * height - 157,539 (47%)
# * weight - 146,363 (43%)
# 3. Benchmark Statistics:
# * bs_backsquat - 246,431 (73%)
# * bs_cleanandjerk - 253,771 (75%)
# * bs_snatch - 256,922 (76%)
# * bs_deadlift - 243,992 (72%)
# * bs_fightgonebad - 318,136 (94%)
# * bs_maxpull_ups - 304,877 (90%)
# * bs_fran - 295,350 (87%)
# * bs_grace - 303,716 (90%)
# * bs_helen - 313,636 (93%)
# * bs_filthy50 - 325,487 (96%)
# * bs_sprint400m - 322,311 (95%)
# * bs_run5k - 307,394 (91%)
#
# ### Workout Results
cols_19 = ['overallrank','w1_reps_total', \
'w2_reps_total','time_2','w2_tiebreak', \
'w3_reps_total','time_3','w3_tiebreak', \
'w4_reps_total','time_4','w4_tiebreak', \
'w5_reps_total']
msno.matrix(df[cols_19].sort_values(by='overallrank'));
cols_part = ['overallrank','w1_reps_total','w2_reps_total','w3_reps_total', \
'w4_reps_total','w5_reps_total']
df[cols_part].dropna()['overallrank'].count()
# A missing of workout results means that athletes did not participate. Just 65% competitors participated in all five workouts and would be left when dropping observations. The missing of workout results (wi_reps_total) has an influence on the performance in other workouts, including especially the overall ranking position. The better the overall rank the less missing values Thus, there is MNAR (Missing Not At Random) of missing participation.
# Recommended action:
# 1. Since most athletes do not finish workouts within timecap, time features can be dropped.
# 2. missing participation:
# * complete case analysis (drop missing observations)
# * imputation to 0
# * labeling new participation feature
# ### Body Measurements
cols_hw = ['overallrank','age','height','weight']
msno.matrix(df[cols_hw].sort_values(by='overallrank'));
# There are just 4 missing age values, these observation can be dropped. Very top athletes do reveal their height and weight data. Most missing values are randomly distributed, the is MAR (Missing At Random). Thus, the imputation of missing height and weight values is recommended, as well as dropping heights and weights.
# ### Benchmark Statistics
cols_bs = ['overallrank','bs_backsquat','bs_cleanandjerk','bs_snatch','bs_deadlift', \
'bs_fightgonebad','bs_maxpull_ups','bs_fran','bs_grace', \
'bs_helen','bs_filthy50','bs_sprint400m','bs_run5k']
msno.matrix(df[cols_bs].sort_values(by='overallrank'));
df_bs = df[(df['bs_backsquat'].notna()) |
(df['bs_cleanandjerk'].notna()) |
(df['bs_snatch'].notna()) |
(df['bs_deadlift'].notna()) |
(df['bs_fightgonebad'].notna()) |
(df['bs_maxpull_ups'].notna()) |
(df['bs_fran'].notna()) |
(df['bs_grace'].notna()) |
(df['bs_helen'].notna()) |
(df['bs_filthy50'].notna()) |
(df['bs_sprint400m'].notna()) |
(df['bs_run5k'].notna())
]
msno.matrix(df_bs[cols_bs].sort_values(by='overallrank'));
df_lift = df[(df['bs_backsquat'].notna()) |
(df['bs_cleanandjerk'].notna()) |
(df['bs_snatch'].notna()) |
(df['bs_deadlift'].notna())]
cols_lift = ['overallrank','bs_backsquat','bs_cleanandjerk','bs_snatch','bs_deadlift']
msno.matrix(df_lift[cols_lift].sort_values(by='overallrank'));
df_lift.shape
# The very most part of benchmark statistics is missing. Amongst these features the olympic lifts are most commonly present. Considering observations with minimum one lift entry, there is still 30% of all competitors left. Since the workout results and the ranking position are influenced by missing of benchmark data, the missing structure is MNAR.
# Recommendations:
# * drop benchmark features besides lifts
# * lifting features:
# - complete case analysis (drop missing observations)
# - multiple imputation of missing lifts
| 5_Missing_Data_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 9.11 (Optional) Stack Unwinding and Tracebacks
def function1():
function2()
def function2():
raise Exception('An exception occurred')
function1()
# ### Traceback Details
# ### Stack Unwinding
# ### Tip for Reading Tracebacks
# ### Exceptions in finally Suites
##########################################################################
# (C) Copyright 2019 by Deitel & Associates, Inc. and #
# Pearson Education, Inc. All Rights Reserved. #
# #
# DISCLAIMER: The authors and publisher of this book have used their #
# best efforts in preparing the book. These efforts include the #
# development, research, and testing of the theories and programs #
# to determine their effectiveness. The authors and publisher make #
# no warranty of any kind, expressed or implied, with regard to these #
# programs or to the documentation contained in these books. The authors #
# and publisher shall not be liable in any event for incidental or #
# consequential damages in connection with, or arising out of, the #
# furnishing, performance, or use of these programs. #
##########################################################################
| examples/ch09/snippets_ipynb/09_11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp tsne_visualisation
# -
# # tSNE visualisation
#
# > tsne visualisation
#hide
from nbdev.showdoc import *
>>> import numpy as np
>>> from sklearn.manifold import TSNE
>>> X = np.array([[0, 0, 0], [0, 1, 1], [1, 0, 1], [1, 1, 1]])
>>> X_embedded = TSNE(n_components=2).fit_transform(X)
>>> X_embedded.shape
X_embedded = X_embedded/100
# +
# tsne2 = TSNE(n_components=2, random_state=0)
# do_plot(tsne2.fit_transform(X), 't-SNE')
# +
import matplotlib as mpl
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE, Isomap, LocallyLinearEmbedding, MDS, SpectralEmbedding
from sklearn.preprocessing import StandardScaler
np.set_printoptions(suppress=True)
np.set_printoptions(precision=4)
plt_style = 'seaborn-talk'
# -
labels = [0, 1, 2, 3, 4, 5, 6]
def do_plot(X_fit, title, labels):
dimension = X_fit.shape[1]
label_types = sorted(list(set(labels)))
num_labels = len(label_types)
colors = cm.Accent(np.linspace(0, 1, num_labels))
with plt.style.context(plt_style):
fig = plt.figure()
if dimension == 2:
ax = fig.add_subplot(111)
for lab, col in zip(label_types, colors):
ax.scatter(X_fit[labels==lab, 0],
X_fit[labels==lab, 1],
c=col)
elif dimension == 3:
ax = fig.add_subplot(111, projection='3d')
for lab, col in zip(label_types, colors):
ax.scatter(X_fit[labels==lab, 0],
X_fit[labels==lab, 1],
X_fit[labels==lab, 2],
c=col)
else:
raise Exception('Unknown dimension: %d' % dimension)
plt.title(title)
plt.show()
# +
# do_plot(X_embedded,"tSME",[0,1])
# -
def tsne_plot(encoded, labels):
"Creates and TSNE model and plots it"
new_values = TSNE(n_components=2).fit_transform(encoded)
x, y = [], []
for value in new_values:
x.append(value[0])
y.append(value[1])
plt.figure(figsize=(16, 16))
for i in range(len(x)):
plt.scatter(x[i],y[i])
plt.annotate(labels[i], xy=(x[i], y[i]), xytext=(5, 2), textcoords='offset points', ha='right', va='bottom')
plt.show()
tsne_plot(X_embedded,[0,1,2,3])
| 16_tsne_visualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import the necessary packages
from imutils import paths
import numpy as np
import argparse
import cv2
import os
from matplotlib import pyplot as plt
# +
def image_resize(image, width = None, height = None, inter = cv2.INTER_AREA):
# initialize the dimensions of the image to be resized and
# grab the image size
dim = None
(h, w) = image.shape[:2]
# if both the width and height are None, then return the
# original image
if width is None and height is None:
return image
# check to see if the width is None
if width is None:
# calculate the ratio of the height and construct the
# dimensions
r = height / float(h)
dim = (int(w * r), height)
# otherwise, the height is None
else:
# calculate the ratio of the width and construct the
# dimensions
r = width / float(w)
dim = (width, int(h * r))
# resize the image
resized = cv2.resize(image, dim, interpolation = inter)
# return the resized image
return resized
dataset_path = "F:\\Datasets\\201912 - dogs vs cats\\training\\cat"
img_path = os.path.join(dataset_path, "cat.1801.jpg")
img = cv2.imread(img_path)
print(img.shape)
img_resized = image_resize(img, height=200)
print(img_resized.shape)
img_resized = img_resized[:,:250,:]
print(img_resized.shape)
plt.imshow(img_resized[:,:,::-1])
| WatermarkRemoval/Image Resizing - Shoaib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
a = np.array([0,1,2,3,4])
print("datatype:", type(a), a.dtype)
print(a.itemsize)
print(a.shape)
print(np.shape(a))
print(a.size)
print(np.size(a))
print(a.nbytes)
a.fill(123)
print(a)
a[:] = 321
print(a)
# #### Slicing
a = np.array([10,11,12,13,14,15,16])
print(a[1:3])
print("negative index")
print(a[3:-3])
print(a[:3])
print(a[3:])
print("STEP")
print("every other element", a[::2])
print("every third element", a[::3])
print("starting from the first", a[1::2])
# #### Multi-dimensional
a = np.array([[0,1, 2, 3, 4],
[8,9,10,11,12]])
print(a.shape)
print(a.size)
print(a.ndim)
print(a[1,2])
print(a[-1,3])
print(a[1])
# #### Indexing
| scripts/scripts_ipynb/numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model with Binary Data
#
# Example testing a model which uses binary data
#
# !cp ../../../proto/prediction.proto ./proto
# !cp -r ../../../proto/tensorflow/tensorflow .
# !python -m grpc.tools.protoc -I. --python_out=. --grpc_python_out=. ./proto/prediction.proto
# +
import json
import requests
import base64
from proto import prediction_pb2
from proto import prediction_pb2_grpc
import grpc
import numpy as np
import pickle
def get_payload():
array_2d = np.arange(9).reshape(3, 3)
print(array_2d)
bdata = pickle.dumps(array_2d)
return bdata
# +
def rest_request_ambassador(deploymentName,namespace,request,endpoint="localhost:8003"):
response = requests.post(
"http://"+endpoint+"/seldon/"+namespace+"/"+deploymentName+"/api/v0.1/predictions",
json=request)
print(response.status_code)
print(response.text)
return response.json()
def grpc_request_ambassador(deploymentName,namespace,data,endpoint="localhost:8004"):
request = prediction_pb2.SeldonMessage(binData = data)
channel = grpc.insecure_channel(endpoint)
stub = prediction_pb2_grpc.SeldonStub(channel)
metadata = [('seldon',deploymentName),('namespace',namespace)]
response = stub.Predict(request=request,metadata=metadata)
return response
def rest_request_docker(request,endpoint="localhost:5000"):
response = requests.post(
"http://"+endpoint+"/predict",
data={"json":json.dumps(request),"isDefault":True})
print(response.text)
return response.json()
def grpc_request_docker(data,endpoint="localhost:5000"):
request = prediction_pb2.SeldonMessage(binData = data)
channel = grpc.insecure_channel(endpoint)
stub = prediction_pb2_grpc.ModelStub(channel)
response = stub.Predict(request=request)
return response
# -
# ## REST
# !s2i build -E environment_rest . seldonio/seldon-core-s2i-python36:0.12 model-with-bindata-rest:0.1
# !docker run --name "model-with-bindata" -d --rm -p 5000:5000 model-with-bindata-rest:0.1
bdata = get_payload()
bdata_base64 = base64.b64encode(bdata).decode('utf-8')
payload = {"meta":{},"binData":bdata_base64}
response = rest_request_docker(payload)
bdata2 = base64.b64decode(response["binData"])
arr_resp = pickle.loads(bdata2)
print(arr_resp)
print(arr_resp.shape)
# !docker rm model-with-bindata --force
# ## gRPC
# !s2i build -E environment_grpc . seldonio/seldon-core-s2i-python36:0.12 model-with-bindata-grpc:0.1
# !docker run --name "model-with-bindata" -d --rm -p 5000:5000 model-with-bindata-grpc:0.1
payload = get_payload()
bdata = get_payload()
resp = grpc_request_docker(bdata)
bdata2 = resp.binData
arr_resp = pickle.loads(bdata2)
print(arr_resp)
print(arr_resp.shape)
# !docker rm model-with-bindata --force
# ## Test using Minikube
#
# **Due to a [minikube/s2i issue](https://github.com/SeldonIO/seldon-core/issues/253) you will need [s2i >= 1.1.13](https://github.com/openshift/source-to-image/releases/tag/v1.1.13)**
# !minikube start --vm-driver kvm2 --memory 4096
# !kubectl create clusterrolebinding kube-system-cluster-admin --clusterrole=cluster-admin --serviceaccount=kube-system:default
# !helm init
# !kubectl rollout status deploy/tiller-deploy -n kube-system
# !helm install ../../../helm-charts/seldon-core-operator --name seldon-core --set usageMetrics.enabled=true --namespace seldon-system
# !kubectl rollout status deploy/seldon-controller-manager -n seldon-system
# ## Setup Ingress
# Please note: There are reported gRPC issues with ambassador (see https://github.com/SeldonIO/seldon-core/issues/473).
# !helm install stable/ambassador --name ambassador --set crds.keep=false
# !kubectl rollout status deployment.apps/ambassador
# # REST
# !eval $(minikube docker-env) && s2i build -E environment_rest . seldonio/seldon-core-s2i-python36:0.12 model-with-bindata-rest:0.1
# !kubectl create -f deployment-rest.json
# !kubectl rollout status deploy/mymodel-mymodel-b0e3779
# ## Test predict
# minikube_ip = !minikube ip
# minikube_port = !kubectl get svc -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].spec.ports[0].nodePort}'
bdata = get_payload()
bdata_base64 = base64.b64encode(bdata).decode('utf-8')
payload = {"meta":{},"binData":bdata_base64}
response = rest_request_ambassador("mymodel","default",payload,minikube_ip[0]+":"+minikube_port[0])
bdata2 = base64.b64decode(response["binData"])
arr_resp = pickle.loads(bdata2)
print(arr_resp)
print(arr_resp.shape)
# !kubectl delete -f deployment-rest.json
# # gRPC
# !eval $(minikube docker-env) && s2i build -E environment_grpc . seldonio/seldon-core-s2i-python36:0.12 model-with-bindata-grpc:0.1
# !kubectl create -f deployment-grpc.json
# Wait until ready (replicas == replicasAvailable)
# !kubectl rollout status deploy/mymodel-mymodel-a9ecaa4
bdata = get_payload()
response = grpc_request_ambassador("mymodel","default",bdata,minikube_ip[0]+":"+minikube_port[0])
bdata2 = response.binData
arr_resp = pickle.loads(bdata2)
print(arr_resp)
print(arr_resp.shape)
# !kubectl delete -f deployment-grpc.json
# !minikube delete
| examples/models/template_model_bindata/modelWithBindata.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Answers: Collections & Loops
#
# Provided here are answers to the practice questions at the end of "Collections & Loops".
# ## Collections
# **Collections Q1**.
# actual approaches could differ
trees_a = trees[5]
trees_b = trees[-3:]
trees_c = trees[2:9:2]
trees_d = trees[::3]
# **Collections Q2**.
#
# Part I.
# actual values will differ
practice_list = ['a', 'b', 2.9, True,
{'A':1, 'B':2}, (2, 3)]
# Part II.
slice_1 = practice_list[1:4]
slice_2 = practice_list[1::2]
slice_3 = practice_list[-3]
# **Collections Q3**.
practice_dict = {'name': 'Shannon',
'favorite_game' : 'Coup',
'height': 65}
# **Collections Q4**.
#
# Part I.
grading = {'A': (90, 100),
'B': (80, 90),
'C': (70, 80),
'D': (60, 70),
'F': (0, 60)}
# Part II.
A_lower = grading['A'][0]
A_upper = grading['A'][1]
# **Collections Q5**.
dict_a = cogs18_dict['Ellis']
dict_b = type(cogs18_dict)
dict_c = len(cogs18_dict)
# ## Loops
# **Loops Q1**.
# +
my_name = 'Shannon'
counter = 0
for char in my_name:
counter += 1
# -
# **Loops Q2**.
# +
# there are other approaches that use string methods
sentence = ''
for word in vaccination_list:
sentence += word
# -
# ## Topic Synthesis
# **Synthesis Q1**.
# +
output = []
val = 0
while val <= 100:
if val % 10 == 0:
output.append(val)
val += 1
# -
# **Synthesis Q2**.
# +
# output will differ based on my_name
my_name = 'Shannon'
counter = 0
for char in my_name:
char = char.lower()
if char not in ['a', 'e', 'i', 'o', 'u']:
counter += 1
# -
# **Synthesis Q3**.
#
# Part I.
# +
# output will differ based on my_name
my_name = 'Shannon'
name_consonants = {}
vowels = ['a', 'e', 'i', 'o', 'u']
for char in my_name:
char = char.lower()
if char not in vowels:
if char not in name_consonants:
name_consonants[char] = 1
else:
name_consonants[char] += 1
# -
# Part II.
# +
# output will differ based on who is taking the exam
consonant_count = 0
for key in name_consonants:
consonant_count += name_consonants[key]
# -
# **Synthesis Q4**.
# +
to_contact = []
for person in staff:
if '_IA' in person:
to_contact.append(person)
# -
# **Synthesis Q5**.
# +
# there are multiple possible solutions/approaches
cogs18_students = 0
for key in ellis_courses:
if 'cogs18' in key:
cogs18_students = cogs18_students + ellis_courses[key]
cogs18_students
# -
# **Synthesis Q6**.
#
# Part I.
# +
# there are multiple possible solutions/approaches
steps = []
to_do = []
grocery = []
for value in to_do_list:
if isinstance(value, int):
steps.append(value)
else:
if 'cogs' in value:
to_do.append(value)
else:
grocery.append(value)
# -
# Part II.
# +
days_of_week = ['Monday', 'Tuesday', 'Wednesday', 'Thursday',
'Friday', 'Saturday', 'Sunday']
steps_dict = {}
for i in range(0,7):
key = days_of_week[i]
val = steps[i]
steps_dict[key] = val
| _build/jupyter_execute/content/0X-answers/03-answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Intro to Pandas, Data Manipulation, and Visualization in Python
# In this section, we will learn and practice how to read in data, conduct data manipulation and visualization in `Python`. In particular, we will be learning the `Pandas` package, which provides a fast and powerful interface to dataframes.
#
# ## Pandas
# <img src="https://img.youtube.com/vi/lsJLLEwUYZM/0.jpg" align="right">
# `Pandas` is a library that provides high-performance, easy-to-use data structures and data analysis tools for `Python`.
#
#
#
# Let's load the package `pandas` as well as `numpy`, and `matplotlib` for visualization later. The next few parameters set up the inline plotting to look nicely for the notebook. This is standard preamble for data processing in `ipython` notebooks that you can use in the future. There are some other variations such as giving `matplotlib` the `ggplot` theme from `R` if you wish (add `plt.style.use('ggplot')`).
#
#
#
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
# %matplotlib inline
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 15, 6
# ## Case Study
#
# ### Instacart orders
#
#
# In this problem, we'll use the dataset from Instacart.com (https://www.instacart.com/datasets/grocery-shopping-2017), a "sometimes-same-day" grocery delivery service that connects customers with Personal Shoppers who pick up and deliver the groceries from local stores. The open data contains order, product, and aisles detailed information. We took a 5% sample of orders in this tutorial.
#
#
#
# ### Read in Data
# Now let's read in a csv file for the dataset `orders.csv` and `orders_products.csv` using the `read_csv` function in pd. Index is very important in Pandas for reasons we will talk about later (subset, merge, ...). Let's specify the index when we read in the data with `index_col = ` parameter.
#
# To get a glimpse of the data, you can do:
# * `.shape` to look at the dimension / size / shape of the dataframe,
# * `.describe()` to see a summary of the data,
# * `.head()` to view first 5 rows, or you can do it with `[:5]`.
orders = pd.read_csv('../data/orders.csv', index_col='order_id');
print(orders.shape)
print(orders.describe())
orders.head()
orders_products = pd.read_csv('../data/orders_products.csv', index_col='order_id');
orders_products.head()
# ### Data Indexing
#
# After reading in the datasets and taking a look at the description or the first few rows, we are interested in some basic dataframe manipulations.
# * **Subset columns:** To select a column, we can:
# 1. index with the name of the column as a string,
# 2. use the attribute operator . on the column name,
# 3. use the `loc[:, ]` function on the column name,
# 4. use the `iloc[:, ]` function on the column index (remember zero indexing in Python!)
#
# In this example we take the `order_hour_of_day` column using each of the method.
print(orders['order_hour_of_day'].head())
print(orders.order_hour_of_day.head())
print(orders.loc[:, 'order_hour_of_day'].head())
print(orders.iloc[:, 4].head())
# You can also select multiple columns by indexing the list of columns you would like to select:
orders[['order_dow', 'order_hour_of_day']].head()
# * **Subset rows**: You can subset the rows of a dataframe by
# 1. `iloc[]`: based on the row numbers
# 2. `loc[]`: based on index value
# 3. `[]` with a logical condition
#
# Let's look at the following examples:
# To get the first 5 rows, use `iloc`:
orders.iloc[range(5)]
# To subset based on the index value, use the `loc` command:
orders.loc[[1076138,1609528]]
# If we only want to look at the order hour of day being 6pm:
orders_18pm = orders[orders['order_hour_of_day'] == 18]
print(orders_18pm.shape)
orders_18pm.iloc[:5,:]
# Another example use case could be that if we want to only keep observations with non-NA values for `days_since_prior_order`, we can use the `isnull()` function which returns a boolean array for indexing:
orders_noNA = orders[~orders['days_since_prior_order'].isnull()];
orders_noNA.head()
# ## Exercise 1: Explore `aisles` and `products` data
# Please read in the `aisles.csv` and `products.csv` files. Answer the following questions:
# * How many distinct aisles are there?
# * What's the name for aisle_id = 61?
# * How many products are there in aisle_id = 61?
#
aisles = pd.read_csv('../data/aisles.csv', index_col='aisle_id');
print(aisles.shape)
print(aisles[['aisle']].drop_duplicates().shape)
print(aisles.loc[61])
aisles.head()
products = pd.read_csv('../data/products.csv', index_col='product_id');
print(products[products['aisle_id'] == 61].shape)
# ### Visualization
#
# Let's try to look at the relationship between the day-of-the-week and the hour-of-day for all orders. We can tabulate them by using the `crosstab()` function:
orders_counts = pd.crosstab(orders['order_hour_of_day'], orders['order_dow'])
orders_counts
# One interesting way to look at this data is to plot the distribution of hour-of-day by different day-of-the-week. A `Pandas` dataframe has some plot functions that can be called directly on it. For example, to do a line plot of the counts by each,
orders_counts.plot()
# That's pretty good already, without us needing to supply any arguments to the plot function. Monday and Sundays seem to be the days with more orders placed than other days. It also seems like there is a little bump on Monday, at around 9 to 10am.
# ## Group, Summarize, and Sort
#
# Suppose we are intereted in knowing something on the individual user level. For example, what's the total number of orders each user had? We can use the `groupby` and `size`. For a single variable this achieves similar effect as `value_counts()` function on that column:
orders.groupby('user_id').size().iloc[0:5]
orders['user_id'].value_counts().head()
# We can sort it further by the `sort_values()` function (and specifying `ascending = False` for decreasing order):
orders.groupby('user_id').size().sort_values(ascending=False).head()
# More generally, you can use the `agg` for aggregating specific summary statistic.
# * You can supply a single type and it will be performed on all variables: for example, getting the mean of each variable on each user:
orders.groupby('user_id').agg('mean')[:5]
# * or supply a dictionary that the specfic variable as key: here only summarizses the mean of the `order_hour_of_day`, and the maximum of the `days_since_prior_order`:
orders.groupby('user_id').agg({'order_hour_of_day': 'mean', 'days_since_prior_order': 'max'})[:5]
# ## Exercise 2: Describing User Patterns
#
# From the `orders_products` dataframe, answer the following questions:
# * Are there more products that are reordered, or never ordered again?
# * Which `product_id` is the most frequently ordered?
# * What is that product called from the `products` dataframe?
# * Is there a relationship between the order when a product is added to cart (`add_to_cart_order`), and whether a product is reordered?
# * What about the relationship between whether the product is reordered and the total number of items in the order?
orders_products['reordered'].value_counts()
orders_products.groupby('product_id').size().sort_values(ascending=False).head()
products.loc[24852]
orders_products.groupby('reordered').agg({'add_to_cart_order': 'mean'})
# ## Merge and Join
#
# With many separate dataframes, it is often useful to join them to understand the relationship between variables and also to create additional features in predictions. Pandas provides high-performance, in-memory join operations that are similar to relational databases such as SQL.
#
# When the two dataframes are both indexed by the same variable that you want to join on, it is easy: use the `join` function on the left dataframe, and the right data frame is supplied as second argument.
#
# If the current index is not the right variable, we can first reindex the data using the `set_index` command:
# ```python
# orders_products=orders_products.set_index('order_id')
# orders=orders.set_index('order_id')
# ```
orders_products_joined = orders_products.join(orders);
orders_products_joined.head()
# When the two columns do not have the same index, we can use the more flexible `merge` function:
# * the `left_on` and `right_on` options specify the column names to be joined on
# * if the variable to be joined on is an index, set `left_index` / `right_index` to `True`.
# * Finally, the `how` option allows you to specify the type of joins (left, right, inner, outer).
order_products_desc = orders_products.merge(products, left_on='product_id', right_index=True, how='left')
order_products_desc.head()
# This way, we can look at the most popular products.
order_products_desc['product_name'].value_counts()[:10].plot(kind='bar')
# ## Exercise 3: Summarizing orders by aisle information
#
# We are interested in knowing which aisles are *LEAST* popular and can make management decisions based on that. Try to answer the following questions:
# * Which aisle has the least number of products ordered from?
# * What about only among the reordered products?
#
#
# **Challenge:** Suppose there was a software glitch that all products with the `add_to_cart_order` more than 30 was not correctly charged. What percent of all orders are affected by this glitch?
orders_products_aisles = order_products_desc.set_index('aisle_id').join(aisles)
orders_products_aisles.head()
orders_products_aisles.groupby('aisle').size().sort_values()
orders_products_aisles[orders_products_aisles['reordered'] ==1].groupby('aisle').size().sort_values()
# ## Reading data from SQL databases
#
# (Note: The content for this section is adapted from the Pandas Cookbook Chapter 9.)
#
# Pandas can read from HTML, JSON, SQL, Excel, HDF5, Stata, and a few other things. We'll talk about reading data from SQL databases now.
#
# You can read data from a SQL database using the `pd.read_sql` function. `read_sql` will automatically convert SQL column names to DataFrame column names.
#
# `read_sql` takes 2 arguments: a `SELECT` statement, and a database connection object. It means you can read from *any* kind of SQL database -- it doesn't matter if it's MySQL, SQLite, PostgreSQL, or something else.
import pandas as pd
import sqlite3
con = sqlite3.connect("../data/weather_2012.sqlite")
df = pd.read_sql("SELECT * from weather_2012 LIMIT 5", con)
df
# And that's it! If you are familiar with SQL type statements, you can try some advanced `SELECT` statements; otherwise, just select everything you need and do the data cleaning in Pandas.
# ## Getting Data Ready for SciKitLearn
#
# Having a cleaned `Pandas` dataframe does not allow you to run through machine learning packages directly yet. The dataframe may need to be appropriately transformed (onehot encoded for categorical variables, scaled, etc.). Both Pandas and scikit-learn offer some useful preprocessing functions.
#
# `pd.get_dummies` takes the dataframe, and a list of categorical columns to be converted into a dummified dataframe. See the following example, where we take the `orders_products_aisles` dataframe that we merged earlier, and convert the string categorical variable `aisle` to be onehot encoded:
X_dum = pd.get_dummies(orders_products_aisles.drop(['reordered','product_name'], axis=1) , columns=['aisle'])
X_dum.head()
# Now we can run it through scikit learn, a wonderful machine learning package, which will be the topic of the SIP lessons.
# +
#If you have the error ModuleNotFoundError: No module named 'sklearn',
#It means you need to install scikit-learn. Depending on how you installed Python,
#You may execute in your terminal the following command:
#pip install scikit-learn
from sklearn import tree
Y = order_products_desc['reordered']
clf = tree.DecisionTreeClassifier()
clf = clf.fit(X_dum, Y)
clf.predict_proba(X_dum)[:,1]
# -
# ## Other useful references
#
# * A useful cheatsheet: https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf
# * The mapping between `R` commands and `Pandas` can be found here, if you are coming from a more `R`-type background:
# https://pandas.pydata.org/pandas-docs/version/0.18.1/comparison_with_r.html
| intro_to_python/7_Dataframes_with_Pandas_Complete.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Excercises Electric Machinery Fundamentals
# ## Chapter 6
# ## Problem 6-26
# + slideshow={"slide_type": "skip"}
# %pylab notebook
# -
# ### Description
# A 460-V 50-hp six-pole $\Delta$ -connected 60-Hz three-phase induction motor has a full-load slip of 4 percent, an efficiency of 91 percent, and a power factor of 0.87 lagging. At start-up, the motor develops 1.75 times the full-load torque but draws 7 times the rated current at the rated voltage. This motor is to be started with an autotransformer reduced voltage starter.
#
# #### (a)
#
# * What should the output voltage of the starter circuit be to reduce the starting torque until it equals the rated torque of the motor?
#
# #### (b)
#
# * What will the motor starting current and the current drawn from the supply be at this voltage?
Vt = 460 # [V]
Wperhp = 746 # official conversion rate of "electrical horsepowers"
Pout = 50 * Wperhp # [W]
PF = 0.87
eta = 0.91
times_tor = 1.75
times_cur = 7
# ### SOLUTION
# #### (a)
# The starting torque of an induction motor is proportional to the square of $V_{TH}$ ,
#
# $$\frac{\tau_\text{start2}}{\tau_\text{start1}} = \left(\frac{V_\text{TH2}}{V_\text{TH1}}\right)^2 = \left(\frac{V_\text{T2}}{V_\text{T2}}\right)^2$$
# If a torque of 1.75 $\tau_{rated}$ is produced by a voltage of 460 V, then a torque of 1.00 $\tau_\text{rated}$ would be produced by a voltage of:
#
# $$\frac{1.00\tau_\text{rated}}{1.75\tau_\text{rated}} = \left(\frac{V_{T2}}{460V}\right)^2$$
Vt2 = sqrt(1.00/times_tor * Vt**2)
print('''
Vt2 = {:.0f} V
==========='''.format(Vt2))
# #### (b)
# The motor starting current is directly proportional to the starting voltage, so
#
# $$I_{L2} = \left(\frac{V_{T2}}{V_T}\right)I_{L1}$$
Il2_Il1 = Vt2/Vt
Il1_Irated = times_cur
Il2_Irated = Il2_Il1 * Il1_Irated
print('''
Il2 = {:.2f} Irated
================='''.format(Il2_Irated))
# The input power to this motor is:
# $$P_\text{in} = \frac{P_\text{out}}{\eta}$$
Pin = Pout / eta
print('Pin = {:.1f} kW'.format(Pin/1000))
# The rated current is equal to:
# $$I_\text{rated} = \frac{P_\text{in}}{\sqrt{3}V_TPF}$$
Irated = Pin / (sqrt(3)*Vt*PF)
print('Irated = {:.2f} A'.format(Irated))
# Therefore, the motor starting current is
Il2 = Il2_Irated * Irated
print('''
Il2 = {:.1f} A
============='''.format(Il2))
# The turns ratio of the autotransformer that produces this starting voltage is:
#
# $$\frac{N_{SE}+N_C}{N_C} = \frac{V_T}{V_{T2}} = a$$
a = Vt/Vt2
print('a = {:.3f}'.format(a))
# so the current drawn from the supply will be:
# $$I_\text{line} = \frac{I_\text{start}}{a}$$
Iline = Il2 / a
print('''
Iline = {:.0f} A
============='''.format(Iline))
| Chapman/Ch6-Problem_6-26.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test 2D CAE
#
# 2D CNN-based AE (CAE) have shown remarkable results for image data, e.g., MNIST. In this notebook I am testing if a 2D CAE with a similar architecture to the CAE that shows favorable results for MNIST (https://blog.keras.io/building-autoencoders-in-keras.html).
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
import os
import sys
import warnings
import matplotlib.pyplot as plt
# Ignore warnings. They just pollute the output
warnings.filterwarnings('ignore')
# Allow importing from parent directories
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
assembly = 'hg19'
window_size = 2400
step_size = window_size / 2
aggregation = 100
chroms = ['chr1', 'chr22']
# -
# ## Load the data
#
# #### Download dataset
# +
import wget
from pathlib import Path
Path('data').mkdir(parents=True, exist_ok=True)
print('Download data...')
# GM12878 ChIP-seq H3K27ac log10 p-val
bw = 'data/ENCFF258KTL.bigWig'
if not Path(bw).is_file():
wget.download(
'https://www.encodeproject.org/files/ENCFF258KTL/@@download/ENCFF258KTL.bigWig',
'data/ENCFF258KTL.bigWig',
)
print('Done!')
# -
# #### Retrieved binned genomic windows
# +
"""Chunk up the bigWig file"""
from ae import bigwig
data_train, data_test = bigwig.chunk(
bw,
window_size,
step_size,
aggregation,
chroms,
verbose=True,
)
# -
# #### Normalize the data
#
# The data is capped at the 99.9th percentile and normalized to `[0, 1]`.
# +
import numpy as np
from sklearn.preprocessing import MinMaxScaler
cutoff = np.percentile(data_train, (0, 99.9))
data_train_norm = np.copy(data_train)
data_train_norm[np.where(data_train_norm < cutoff[0])] = cutoff[0]
data_train_norm[np.where(data_train_norm > cutoff[1])] = cutoff[1]
cutoff = np.percentile(data_test, (0, 99.9))
data_test_norm = np.copy(data_test)
data_test_norm[np.where(data_test_norm < cutoff[0])] = cutoff[0]
data_test_norm[np.where(data_test_norm > cutoff[1])] = cutoff[1]
print('Train max: {} | Train norm max: {}'.format(np.max(data_train), np.max(data_train_norm)))
print('Test max: {} | Test norm max: {}'.format(np.max(data_test), np.max(data_test_norm)))
data_train_norm = MinMaxScaler().fit_transform(data_train_norm)
data_test_norm = MinMaxScaler().fit_transform(data_test_norm)
# -
# #### Convert the 1D data into 2D bar chart images with 1 channel
# +
from ae.utils import to_2d
ydim = 12
val_max = 1.0
dtype = np.float32
data_2d_train_norm = to_2d(data_train_norm, ydim, val_max, dtype)
data_2d_test_norm = to_2d(data_test_norm, ydim, val_max, dtype)
# +
import seaborn as sns
signal = np.sum(
data_2d_train_norm.reshape(data_2d_train_norm.shape[0], data_2d_train_norm.shape[1] * data_2d_train_norm.shape[2]),
axis=1
)
def logistic(x):
return 1 / (1 + 1000 * np.exp(-x / 30))
plt.figure(figsize=(20, 2))
sns.distplot(signal)
plt.figure(figsize=(20, 2))
plt.plot(logistic(np.arange(300)))
sample_weights = logistic(signal)
plt.figure(figsize=(20, 2))
# -
# #### Plot the 10 windows with the highest amount of signal
# +
import matplotlib.pyplot as plt
offset = np.argsort(signal[np.where(signal > threshold)])[-10:]
n = 10
plt.figure(figsize=(20, n))
for i, k in enumerate(offset):
# display original
ax = plt.subplot(n, 5, i + 1)
plt.imshow(data_2d_train_norm_sub[k], cmap="gray")
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# -
# ## Build the 2D CAE
#
# The architecture is similar to the CAE used for MNIST (https://blog.keras.io/building-autoencoders-in-keras.html).
# +
from ae.cnn import cae2d
encoder, decoder, autoencoder = cae2d(
(ydim, window_size // aggregation, 1),
optimizer='adadelta',
loss='binary_crossentropy',
summary=True
)
# -
# ## Train the model
# +
from ae.utils import train
train(
autoencoder,
data_2d_train_norm_sub.reshape((
data_2d_train_norm_sub.shape[0],
data_2d_train_norm_sub.shape[1],
data_2d_train_norm_sub.shape[2],
1
)),
data_2d_test_norm.reshape(
data_2d_test_norm.shape[0],
data_2d_test_norm.shape[1],
data_2d_test_norm.shape[2],
1
),
epochs=15,
batch_size=256,
)
# -
# ## Predict genomic windows and get the prediction loss
# +
from keras.metrics import binary_crossentropy
from ae.utils import predict_2d
predicted, loss, encoded = predict_2d(
encoder,
decoder,
data_2d_test_norm.reshape(
data_2d_test_norm.shape[0],
data_2d_test_norm.shape[1],
data_2d_test_norm.shape[2],
1
),
validator=binary_crossentropy
)
# -
# ## Compare predictions against the ground truth
#
# The groundtruth is plotted in black and white. The predictions are plotted in a red colormap.
# +
import matplotlib.pyplot as plt
ymax = 1.0
offset = np.arange(3809, 3827) # 2011: good to bad
offset = np.arange(32370, 32380)
n = offset.size
plt.figure(figsize=(20, n))
for i, k in enumerate(offset):
# display original
ax = plt.subplot(n, 10, i * 2 + 1)
plt.imshow(data_2d_test_norm[k], cmap="gray", interpolation='nearest')
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(n, 10, i * 2 + 2)
plt.imshow(predicted[k], cmap="Reds", interpolation='nearest')
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# -
# ---
#
# ## As a sanity check: run a very similar CAE against MNIST
# +
from keras.datasets import mnist
(x_train, _), (x_test, _) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
x_train = np.reshape(x_train, (len(x_train), 28, 28, 1)) # adapt this if using `channels_first` image data format
x_test = np.reshape(x_test, (len(x_test), 28, 28, 1)) # adapt this if using `channels_first` image data format
from ptfind.nn import cae2d
encoder, decoder, autoencoder = cae2d(
(28, 28, 1),
filters=[32, 64, 128, 10],
kernel_sizes=[5, 5, 3],
optimizer='adadelta',
loss='mse',
summary=True
)
autoencoder.fit(
x_train, x_train,
epochs=25,
batch_size=128,
shuffle=True,
validation_data=(x_test, x_test),
)
# -
# #### Visualize the predictions on MNIST
# +
decoded_imgs = autoencoder.predict(x_test)
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(2, n, i + n + 1)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
| experiments/notebooks/Test CAE 2.4Kb 2D (old).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="t3_UKdma4cEB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 493} outputId="8810aa07-93cf-4c41-af69-b4bfe2392810"
# !pip install --upgrade tables
# !pip install eli5
# !pip install xgboost
# + id="pMhQ9x824lc-" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
import eli5
from eli5.sklearn import PermutationImportance
# + id="V6mPX0sW5Uiu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="81716e44-b41c-491d-98ff-c2f77395d01e"
# cd '/content/drive/My Drive/Colab Notebooks/matrix/matrix_two/dw_matrix_car'
# + id="iiPjRoHY5gqm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="28da45fc-4360-46fa-b5d3-320af1bbb7d6"
df = pd.read_hdf('data/car.h5')
df.shape
# + id="lh7iJ7LI5njm" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0],list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat+ SUFFIX_CAT] = factorized_values
# + id="U8uN6E8459En" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a3d22803-838c-4d0f-ce66-e764081f3e3e"
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="7uNMipwf5_ue" colab_type="code" colab={}
def run_model(model, feats):
X = df[feats].values
y = df['price_value'].values
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + [markdown] id="NsZnuGga7cE2" colab_type="text"
# ## DecisionTree
# + id="8HwPX5Rg6Gou" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="32ddc8ea-084a-447e-da05-93f7eeef8577"
run_model( DecisionTreeRegressor(max_depth=5), cat_feats)
# + [markdown] id="DGsa7Fy37gA2" colab_type="text"
# ## Random Forest
# + id="p5vQ0tzH7hXe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7d725d1a-985b-46ad-b31f-5400ed08f1d0"
model = RandomForestRegressor(max_depth=5, n_estimators=50, random_state=0)
run_model(model, cat_feats)
# + [markdown] id="Jt1vNCHM87zs" colab_type="text"
# ## XGBoost
# + id="E7bcSe-g-hy_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="b3073e0c-c7fe-45b3-cb4a-e75bbc3ec964"
xgb_params = {
'max_depth':5,
'n_estimators':50,
'learning_rate':0.1,
'seed': 0
}
run_model(xgb.XGBRegressor(**xgb_params), cat_feats)
# + id="P2cQIPPP7SHP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="1aeb3802-cee8-4d0a-f847-b654ba4c67eb"
m = xgb.XGBRegressor(max_depth=5, n_estimators=50, learning_rate=0.1, seed=0)
m.fit(X,y)
imp=PermutationImportance(m, random_state=0).fit(X,y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="7ewTc23t_-OS" colab_type="code" colab={}
feats= ['param_napęd__cat',
'param_rok-produkcji', #usunieto __cat
'param_stan__cat',
'param_skrzynia-biegów__cat',
'param_faktura-vat__cat',
'param_moc', #usunieto __cat
'param_marka-pojazdu__cat',
'feature_kamera-cofania__cat',
'param_typ__cat',
'param_pojemność-skokowa', #usunieto __cat
'seller_name__cat',
'feature_wspomaganie-kierownicy__cat',
'param_model-pojazdu__cat',
'param_wersja__cat',
'param_kod-silnika__cat',
'feature_system-start-stop__cat',
'feature_asystent-pasa-ruchu__cat',
'feature_czujniki-parkowania-przednie__cat',
'feature_łopatki-zmiany-biegów__cat',
'feature_regulowane-zawieszenie__cat']
# + id="YqJ6LvcCCGr-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="91a1706e-7910-4cee-8d54-9d7c14ee6bb5"
len(feats)
# + id="R02hmTQ_Cir3" colab_type="code" colab={}
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x: -1 if str(x) == 'None' else int(x)) # edytujemy poszczegolne zmienne tak, zeby miały większą wartość objaśniającą (zmienne ciągłe)
df['param_moc'] = df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
# + id="gdaVljf9CJev" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="ac23877e-2b6f-43ab-8b0e-2bd695de7040"
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="8jNJHxdRCTma" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="cad45d61-ad23-4ae1-b57a-ed156237a261"
df['param_pojemność-skokowa'].unique()
# + id="yZH9zouSD0Wy" colab_type="code" colab={}
df['param_pojemność-skokowa'] = df['param_pojemność-skokowa'].map(lambda x: -1 if str(x) == 'None' else int(str(x).split('cm')[0].replace(' ','')))
# + id="Gt_lCE9tEj_w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="f17de2dd-d6a2-43e4-fb64-658c716c08e5"
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="80txSEbwExKX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7eba66c8-c0cf-4364-8a70-a1b7d2735256"
# !git init
# !git add .
# + id="25SudIBRJ21o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="2f3b5a17-0cea-4008-b2b3-e5e04b10d45f"
# !git config --global user.email '<EMAIL>'
# !git config --global user.name 'KonradWasikiewicz'
# !git commit
# + id="b-1mzRduJ4mh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fc525018-a887-495f-eb58-eed064ecca5c"
# !git remote add origin https://github.com/KonradWasikiewicz/dw_matrix_car.git
# + id="JI_ZRR0CKEuw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="32ab3b7f-60bd-4c06-a2ea-5e20b3050ac7"
# !git push --all origin
# + id="bwgNUnNKKqmR" colab_type="code" colab={}
| day4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
# +
from spectralCollocationSolver import SpectralCollocationSolver
import numpy as np
import matplotlib.pyplot as plt
# -
# If the barycentricInterpolation module can not be found ("ModuleNotFoundError: No module named 'barycentricInterpolation'"), probably you just forgot to install it. The package is provided in my github repository with the same name (https://github.com/lubo92/barycentricLagrangeInterpolation/). Download or clone the repository and then run the setup.py installation utility. For details see the readme in the repository.
# # Define the Function to Be Solved
# To some people the following coding approach may appear a little bit strange. I'm not sure if this is a good concept or not - but it works!
# In the provided python package (`spectralCollocationSolver`) a class `SepctralCollocationSolver` exists. This class contains all functionality to solve differential equations - with one exception: The traget function is not contained.
# In order to define the function to be solved, follow the provided exmaple:
# * First you need to create a child class of the `SpectralCollocationSolver`. In the example below it's the `HarmonicOscillator` class.
# * Copy the `__init__` method from the exmaple to your function.
# * Create the `residuals` function. In this function you define your differential equation to be solved. You have to return the residual of the interpolating function solving your differential equation at every node as well as the residuals for all boundary conditions.
# ---
# **CAVEAT:**
#
# Since we are using a barycentric Langrange Interpolation to solve the differential equation, the target function is only defined on the intervall [-1,1]. If your differential equation is defined on an other domain you have to transform your coordinates, e.g. with a tangent.
#
# Be cautious: If you transform your coordinates, you need also transform your derivatives/ differential equations. For details see my repository on the Blasius equation, where I apply this Python package.
#
# ---
# **How to define the differential equation:**
# * `values` is the array containing the values of the interpolating function at all nodes
# * `self.f.derivative(values,self.D1)` returns the (values of the) first derivative of the function defined by '`values`' at the nodes
# * `self.f.derivative(values,self.D2)` returns the (values of the) second derivative of the function defined by '`values`' at the nodes
# * `self.f.evaluateFunction(-1,values,self.weights,self.nodes)` gives you the function at the point -1
class HarmonicOscillator(SpectralCollocationSolver):
def __init__(self):
SpectralCollocationSolver.__init__(self)
def residuals(self,values,args):
"""This function calculates some residuals:
- the residuals of the harmonic oscillator equation at every node
- the residuals of every boundary condition
Parameters:
values(ndarray[nNodes]): values of the function to calculate residuals for - given at every sampling point
args(tuple): (omega^2,f(0),f'(0))
Returns:
ndarray[nNodes+2] : residuals at every node as well as residuals of the two boundary conditions
"""
omega2 = args[0]
boundary1 = args[1]
boundary2 = args[2]
#residuals at the sampling points
residuals1 = omega2*values + self.f.derivative(values,self.D2)
#residuals for boundary conditions
residuals2 = self.f.evaluateFunction(-1,values,self.weights,self.nodes)-boundary1
residuals3 = self.f.evaluateFunction(-1,self.f.derivative(values,self.D1),self.weights,self.nodes)-boundary2
temp2 = np.reshape(residuals2,1)
temp3 = np.reshape(residuals3,1)
return np.concatenate((residuals1, temp2, temp3))
# # Initialize Class
solver = HarmonicOscillator()
# # Set up Solver
# 'nNodes' is the number of nodes, at which the differential equation is solved. A low number of nodes gives very quick results and is in most cases nummerically stable, but it might not be possible approximate the target function well with only very few nodes. A high number of nodes may take longer to calculate and get's at some point nummerically unstable.
solver.setupUtilities(nNodes = 70)
# # Solve Equation
omegaSquare = 30
x0 = 1
v0 = 0
init = np.random.normal(size=solver.nNodes)
solution = solver.solve(init,(omegaSquare,x0,v0))
# # Check Result
# In order to check the result, we evalute the function at the point `t = 2*pi/omega` and subtract the initial vaule `x0=1`. We expect an result close to zero.
solver.f.evaluateFunction(-1+2*np.pi/np.sqrt(omegaSquare),solution.x,solver.weights,solver.nodes)-1
| exmaple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Examples for general profile shapes
from shape_generator import CrossSectionHolding as CrossSection, Circle, csv
# ## Example for standard egg cross section
profile_dimensions = csv("""
label,name,r
P0,0,30
P1,1,35
P2,2,40
P3,3,45
P4,4,50
P5,5,55
P6,6,60
P7,6a,65
""")
unit = 'cm'
profile_dimensions
# +
label = 'P0'
name, r = profile_dimensions.loc[label].values
R = 3 * r
roh = r / 2
height = r * 3
width = r * 2
# h1 = roh - (r + roh) / (R - roh) * roh
h1 = r/ 5
cross_section = CrossSection(label=label, description=name, width=width, height=height, unit=unit)
cross_section.add(Circle(roh, x_m=roh))
cross_section.add(h1)
cross_section.add(Circle(R, x_m=2 * r, y_m=-(R - r)))
cross_section.add(2 * r)
cross_section.add(Circle(r, x_m=2 * r))
# -
import pandas as pd
pd.DataFrame(cross_section.get_points()).T.rename(columns={0:'$h_i$', 1:'$A_i$'})
fig = cross_section.profile_figure()
# ## Example for custom cross section
# + pycharm={"name": "#%%\n"}
def add_and_show(cs, *args, **kwargs):
cs.add(*args, **kwargs)
print('-' * 5, *cs.shape_description, '-' * 5, sep='\n')
cs.profile_figure()
# +
no = 'test'
name = 'Kreis'
r = 20 # cm
unit = 'cm'
kreis = CrossSection(label=no, description=name, height=2*r, unit=unit)
add_and_show(kreis, Circle(r, x_m=r))
# -
add_and_show(kreis, r)
add_and_show(kreis, 30, '°slope')
add_and_show(kreis, None,1.5*r)
| docs/show_case.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''venv'': venv)'
# name: python3
# ---
# + [markdown] id="O-W2ZQ6CN-gZ" pycharm={"name": "#%% md\n"}
# # Query Classifier Tutorial
# [](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial14_Query_Classifier.ipynb)
#
# In this tutorial we introduce the query classifier the goal of introducing this feature was to optimize the overall flow of Haystack pipeline by detecting the nature of user queries. Now, the Haystack can detect primarily three types of queries using both light-weight SKLearn Gradient Boosted classifier or Transformer based more robust classifier. The three categories of queries are as follows:
#
#
# ### 1. Keyword Queries:
# Such queries don't have semantic meaning and merely consist of keywords. For instance these three are the examples of keyword queries.
#
# * arya stark father
# * jon snow country
# * arya stark younger brothers
#
# ### 2. Interrogative Queries:
# In such queries users usually ask a question, regardless of presence of "?" in the query the goal here is to detect the intent of the user whether any question is asked or not in the query. For example:
#
# * who is the father of arya stark ?
# * which country was jon snow filmed ?
# * who are the younger brothers of arya stark ?
#
# ### 3. Declarative Queries:
# Such queries are variation of keyword queries, however, there is semantic relationship between words. Fo example:
#
# * Arya stark was a daughter of a lord.
# * <NAME> was filmed in a country in UK.
# * Bran was brother of a princess.
#
# In this tutorial, you will learn how the `TransformersQueryClassifier` and `SklearnQueryClassifier` classes can be used to intelligently route your queries, based on the nature of the user query. Also, you can choose between a lightweight Gradients boosted classifier or a transformer based classifier.
#
# Furthermore, there are two types of classifiers you can use out of the box from Haystack.
# 1. Keyword vs Statement/Question Query Classifier
# 2. Statement vs Question Query Classifier
#
# As evident from the name the first classifier detects the keywords search queries and semantic statements like sentences/questions. The second classifier differentiates between question based queries and declarative sentences.
# + [markdown] id="yaaKv3_ZN-gb" pycharm={"name": "#%% md\n"}
# ### Prepare environment
#
# #### Colab: Enable the GPU runtime
# Make sure you enable the GPU runtime to experience decent speed in this tutorial.
# **Runtime -> Change Runtime type -> Hardware accelerator -> GPU**
#
# <img src="https://raw.githubusercontent.com/deepset-ai/haystack/master/docs/img/colab_gpu_runtime.jpg">
# + [markdown] id="TNlqD5HeN-gc" pycharm={"name": "#%% md\n"}
# These lines are to install Haystack through pip
# + colab={"base_uri": "https://localhost:8080/"} id="CjA5n5lMN-gd" outputId="da688e25-ad0e-41d3-94cf-581858fc05a4" pycharm={"name": "#%%\n"}
# Install the latest release of Haystack in your own environment
# #! pip install farm-haystack
# Install the latest master of Haystack
# !pip install --upgrade pip
# !pip install git+https://github.com/deepset-ai/haystack.git#egg=farm-haystack[colab]
# Install pygraphviz
# !apt install libgraphviz-dev
# !pip install pygraphviz
# In Colab / No Docker environments: Start Elasticsearch from source
# ! wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-7.9.2-linux-x86_64.tar.gz -q
# ! tar -xzf elasticsearch-7.9.2-linux-x86_64.tar.gz
# ! chown -R daemon:daemon elasticsearch-7.9.2
import os
from subprocess import Popen, PIPE, STDOUT
es_server = Popen(
["elasticsearch-7.9.2/bin/elasticsearch"], stdout=PIPE, stderr=STDOUT, preexec_fn=lambda: os.setuid(1) # as daemon
)
# wait until ES has started
# ! sleep 30
# + [markdown] id="fAfd2cOQN-gd" pycharm={"name": "#%% md\n"}
# If running from Colab or a no Docker environment, you will want to start Elasticsearch from source
# + [markdown] id="Z7Tu5OQnN-ge" pycharm={"name": "#%% md\n"}
# ## Initialization
#
# Here are some core imports
# + [markdown] id="Vm9gqTioN-gf" pycharm={"name": "#%% md\n"}
# Then let's fetch some data (in this case, pages from the Game of Thrones wiki) and prepare it so that it can
# be used indexed into our `DocumentStore`
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["e827efc5cd744b55b7d7d702663b8250", "d7f2e3e918514031acf261116b690c5b", "e7c2e561bfdf49aa88814126e0471a58", "3838eb97db0f46a8828473c0e686f5df", "769ee7fdb9c549e2a8d7f0e0719477eb", "<KEY>", "f069997c1a114e46947c8a07cebc709b", "<KEY>", "02b9c7b799a846b6974eac600c0178ba", "667c145e9bef4f5b99fffda361a2ea47", "<KEY>", "<KEY>", "216fd635f06847c5a7daf25d543e321b", "fe18331098014a22aeb390730d35fedb", "b8dddaca70054b5cab955e0dab9d6d0a", "490edef475d646a2ba12b429c1244de4", "<KEY>", "cbe8cc2b1394456bac529507081a2263", "<KEY>", "6bf5fe6ae4bf494e9c33e0940da62951", "<KEY>", "<KEY>", "ef99a76fdfbc48669ed4d18e118bfa42", "4de441f28ec040e995c285b1e3fcadb9", "<KEY>", "9a4e9075d67b4bbaab47ff97e8ea539c", "<KEY>", "<KEY>", "b2249a087bd343999a5319152d7e0548", "1618f5a1981c4d18a67be3965745aa49", "<KEY>", "a226c1daa4454c55a03b1edf595a33ae", "6de6dbe386d447f588c982d3501bd1c1", "dc143ea6a5594b769b3a4880d83f0e1b", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "e80fc1c849354ee89fa0d51f7de4c78e", "b4eb040d5e224e619414a4d4740badc5", "30413c25f3774ef6860be4230804e31c", "<KEY>", "<KEY>", "<KEY>", "4d37fd7e90a64ff68870b86334cfad7e", "394b26eab89b410db98a07cdf6609ead", "<KEY>", "027441d4e1f14648919f54ba9d9a5204", "b27ce7d9f5e74ef895b809e519673c60", "<KEY>", "<KEY>", "de371bd60c64464284f47eeef0a07bed", "<KEY>", "<KEY>", "7ea5f945a5aa4eb4adc96e9f986a0d71", "<KEY>", "<KEY>", "6b97bc4ee3924dfdbe72738d137c967e", "c05f2ee787944a9e931f74de9831599c", "14cd6229f5e043f8862c6fe1773df4e0", "7541bd7ec86f4c028405e6e9ca6f4d06", "853a80d353144e8d9cd6ce8788c7a9bb", "<KEY>", "d4e2dd1fa16b40c3bb115ecccba69a03", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "24a7839d950740c9958e2d88b7ad829a", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "56660d608cbe451e9ec0701c1941a7ab", "<KEY>", "<KEY>", "<KEY>", "9d36e8f0ab3e4a18b519e5dc04a51061", "5b2cc531f1e44c318488c993024a94b7", "463ff451eab94f509f4ba6ce4e1ae1ef", "7e59ccb05ef3494eb8ad400dd8f606e0", "<KEY>", "42819b5d13df44b392be19add1bdd122", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "41635bd3da9c4c1da50fe86a01da25bf", "3fa3082333c344d1be66344a7d0ff542", "a2905ce60084476da1c96d46a63f138d", "4de5d23d236646629ee2cabf08f4621b", "2d5ce6e282b14019ab5aece4ceb4e643", "f028104b8e9548f0a7e53f1edcd63c4e", "4cea79778d26402dad42393c2bebfe10", "<KEY>", "<KEY>", "6445465e075f4254b18874e0f7353885", "<KEY>", "<KEY>", "e0599025f4534a89932b42945c3c1e21", "<KEY>", "09dd2dd5a35c4f418ea1ef010e6831d2", "<KEY>", "b780dea3ed6e4057a09e432a067596fb", "1d14929d95684c05a66c958fd8a5b89d", "11d32ef999d94db187dab41a1bbf7723", "3d71e9d26ebb41d980819d608ddb115a", "<KEY>", "96b976f448be47d892dc1923c51c470e", "<KEY>", "5fa38582019043d084a6ecb2f2fea016", "1c10236190604ad49fa72d4d851ac316", "<KEY>", "66f26219cc6248288423c2e7ba5304c4", "33143aed23264e82a7750d5fbc8e22a7", "<KEY>", "<KEY>", "0cab5420cdb144e9aa3dfa907e1ed729", "28b880a5c9864a63b2d557a1ec9458a5", "6e5b975419994befb261100d7719e991", "0f7ab0ce7ee24838a6cbffac4cdb3040", "<KEY>", "ba67c6d2324d440abbd25de05161b381", "84cea882fd1e431b9c93631ee3f4ed17", "cea38043b06d419fa63695a316233088", "<KEY>", "f53911fd1c9c47c69e69206c3169158a", "33b89e0dfe124edaa5c90ada36a4c7a8", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "814e632c690e4745a93fdef461fd48b6", "56b67d80ef05478da31ef471eb7de385", "1bc6ff28903f4b2e9aefb6275aa047be", "<KEY>", "<KEY>", "409718ada73943e69bee6f73e0db5b94", "e1e04bcfd0754a3cb1ea6ac0d8ba0efa", "fc67d27eeac0448ba310e3fb991b9b5a", "<KEY>", "e7227229d21d4d7292733f373fa7f9db", "adbe499b09574ba6853581f3f6cc21d5", "<KEY>", "<KEY>", "209eb3ac83504ba3920b13e5aec28da6", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "399fa7054fc74dfea45d22ee20765b30", "26be6fe237fd4b8c8d8fac787d72e823", "<KEY>", "d03e8b5ed27c4c97a8a25bdfa82a752e", "f27ffc683e714a28a0842a730e736716", "b5efbf17543a4db7a3b01ee0e28743c9", "<KEY>", "892b9273604c4440ad179ed4a8dfc5e1", "96f73362e4694c24acb058e1b7d69a14", "<KEY>", "<KEY>", "76e76e39d6d64438a24919e973e0053c", "<KEY>", "d377b1dec2a84337bc1644ff51464e4d", "038267d5661343deb4be8a3c6d16a221", "443739aca635464ea01e08e800dd40aa", "484eafd3786b43219fd4103aba12c2e9", "<KEY>", "<KEY>", "580a190f9eee4b0e8d3008e529535aa7", "3fc82442548e42e887ba9ad11dfd5d77", "c7ca455474c04af18ade2d3fd906727e", "d31dc63868a4475fbcfc7406723ad359", "2ed78454dfa347aaa0ab9be6be341264", "0ace7e3fad4b430c9fb09d6736ea9ac3", "b6c733e5327a40259700554efe576527", "1b81083a5e9644c08eb711693d480ed2", "ac67a20654414b0ea3b36acdc6e86171", "<KEY>", "518539c52ba54f1f9d59ec0dfdca132d", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "f1238867e4e249c1a0daa553d459e98b", "<KEY>", "4bbce51f2459479f8a95064c17b73e2c", "<KEY>", "<KEY>", "01781b50453c401785af24b5ea608440", "6476b24e759f4ae4ba594ec63190506c", "<KEY>", "6d34f638e428428f8463026b19172847", "<KEY>", "<KEY>", "<KEY>", "3ff47ea8d9a641cd9cfece170597dd51", "<KEY>"]} id="Ig7dgfdHN-gg" outputId="62383a2b-d653-4f9f-e3a9-4b2cb90ee007" pycharm={"name": "#%%\n"}
from haystack.utils import (
print_answers,
print_documents,
fetch_archive_from_http,
convert_files_to_docs,
clean_wiki_text,
launch_es,
)
from haystack.pipelines import Pipeline
from haystack.document_stores import ElasticsearchDocumentStore
from haystack.nodes import (
ElasticsearchRetriever,
EmbeddingRetriever,
FARMReader,
TransformersQueryClassifier,
SklearnQueryClassifier,
)
# Download and prepare data - 517 Wikipedia articles for Game of Thrones
doc_dir = "data/tutorial14"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/documents/wiki_gameofthrones_txt14.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# convert files to dicts containing documents that can be indexed to our datastore
got_docs = convert_files_to_docs(dir_path=doc_dir, clean_func=clean_wiki_text, split_paragraphs=True)
# Initialize DocumentStore and index documents
launch_es()
document_store = ElasticsearchDocumentStore()
document_store.delete_documents()
document_store.write_documents(got_docs)
# Initialize Sparse retriever
es_retriever = ElasticsearchRetriever(document_store=document_store)
# Initialize dense retriever
embedding_retriever = EmbeddingRetriever(
document_store=document_store,
model_format="sentence_transformers",
embedding_model="sentence-transformers/multi-qa-mpnet-base-dot-v1",
)
document_store.update_embeddings(embedding_retriever, update_existing_embeddings=False)
reader = FARMReader(model_name_or_path="deepset/roberta-base-squad2")
# + [markdown] id="5kPAbP4EN-gk" pycharm={"name": "#%% md\n"}
# ## Keyword vs Question/Statement Classifier
#
# The keyword vs question/statement query classifier essentially distinguishes between the keyword queries and statements/questions. So you can intelligently route to different retrieval nodes based on the nature of the query. Using this classifier can potentially yield the following benefits:
#
# * Getting better search results (e.g. by routing only proper questions to DPR / QA branches and not keyword queries)
# * Less GPU costs (e.g. if 50% of your traffic is only keyword queries you could just use elastic here and save the GPU resources for the other 50% of traffic with semantic queries)
#
# ![image]()
#
# + [markdown] id="K4wZ3xkQCHjY"
# Below, we define a `SklearnQueryClassifier` and show how to use it:
#
# Read more about the trained model and dataset used [here](https://ext-models-haystack.s3.eu-central-1.amazonaws.com/gradboost_query_classifier/readme.txt)
# + id="Sz-oZ5eJN-gl" pycharm={"name": "#%%\n"}
# Here we build the pipeline
sklearn_keyword_classifier = Pipeline()
sklearn_keyword_classifier.add_node(component=SklearnQueryClassifier(), name="QueryClassifier", inputs=["Query"])
sklearn_keyword_classifier.add_node(
component=embedding_retriever, name="EmbeddingRetriever", inputs=["QueryClassifier.output_1"]
)
sklearn_keyword_classifier.add_node(component=es_retriever, name="ESRetriever", inputs=["QueryClassifier.output_2"])
sklearn_keyword_classifier.add_node(component=reader, name="QAReader", inputs=["ESRetriever", "EmbeddingRetriever"])
sklearn_keyword_classifier.draw("pipeline_classifier.png")
# + id="fP6Cpcb-o0HK"
# Run only the dense retriever on the full sentence query
res_1 = sklearn_keyword_classifier.run(query="Who is the father of Arya Stark?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_1, details="minimum")
# Run only the sparse retriever on a keyword based query
res_2 = sklearn_keyword_classifier.run(query="arya stark father")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_2, details="minimum")
# + id="EZru--pao1UG"
# Run only the dense retriever on the full sentence query
res_3 = sklearn_keyword_classifier.run(query="which country was jon snow filmed ?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_3, details="minimum")
# Run only the sparse retriever on a keyword based query
res_4 = sklearn_keyword_classifier.run(query="jon snow country")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_4, details="minimum")
# + id="MWCMG8MJo1tB"
# Run only the dense retriever on the full sentence query
res_5 = sklearn_keyword_classifier.run(query="who are the younger brothers of arya stark ?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_5, details="minimum")
# Run only the sparse retriever on a keyword based query
res_6 = sklearn_keyword_classifier.run(query="arya stark younger brothers")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_6, details="minimum")
# + [markdown] id="dQ5YMyd4CQPC"
# ## Transformer Keyword vs Question/Statement Classifier
#
# Firstly, it's essential to understand the trade-offs between SkLearn and Transformer query classifiers. The transformer classifier is more accurate than SkLearn classifier however, it requires more memory and most probably GPU for faster inference however the transformer size is roughly `50 MBs`. Whereas, SkLearn is less accurate however is much more faster and doesn't require GPU for inference.
#
# Below, we define a `TransformersQueryClassifier` and show how to use it:
#
# Read more about the trained model and dataset used [here](https://huggingface.co/shahrukhx01/bert-mini-finetune-question-detection)
# + id="yuddZL3FCPeq"
# Here we build the pipeline
transformer_keyword_classifier = Pipeline()
transformer_keyword_classifier.add_node(
component=TransformersQueryClassifier(), name="QueryClassifier", inputs=["Query"]
)
transformer_keyword_classifier.add_node(
component=embedding_retriever, name="EmbeddingRetriever", inputs=["QueryClassifier.output_1"]
)
transformer_keyword_classifier.add_node(component=es_retriever, name="ESRetriever", inputs=["QueryClassifier.output_2"])
transformer_keyword_classifier.add_node(component=reader, name="QAReader", inputs=["ESRetriever", "EmbeddingRetriever"])
transformer_keyword_classifier.draw("pipeline_classifier.png")
# + id="uFmJJIb_q-X7"
# Run only the dense retriever on the full sentence query
res_1 = transformer_keyword_classifier.run(query="Who is the father of <NAME>?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_1, details="minimum")
# Run only the sparse retriever on a keyword based query
res_2 = transformer_keyword_classifier.run(query="arya stark father")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_2, details="minimum")
# + id="GMPNcTz8rdix"
# Run only the dense retriever on the full sentence query
res_3 = transformer_keyword_classifier.run(query="which country was jon snow filmed ?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_3, details="minimum")
# Run only the sparse retriever on a keyword based query
res_4 = transformer_keyword_classifier.run(query="jon snow country")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_4, details="minimum")
# + id="jN5zdLJbrzOh"
# Run only the dense retriever on the full sentence query
res_5 = transformer_keyword_classifier.run(query="who are the younger brothers of arya stark ?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_5, details="minimum")
# Run only the sparse retriever on a keyword based query
res_6 = transformer_keyword_classifier.run(query="arya stark younger brothers")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_6, details="minimum")
# + [markdown] id="zLwdVwMXDcoS"
# ## Question vs Statement Classifier
#
# One possible use case of this classifier could be to route queries after the document retrieval to only send questions to QA reader and in case of declarative sentence, just return the DPR/ES results back to user to enhance user experience and only show answers when user explicitly asks it.
#
# ![image]()
#
# + [markdown] id="SMVFFRtMPVIt"
# Below, we define a `TransformersQueryClassifier` and show how to use it:
#
# Read more about the trained model and dataset used [here](https://huggingface.co/shahrukhx01/question-vs-statement-classifier)
# + id="BIisEJrzDr-9"
# Here we build the pipeline
transformer_question_classifier = Pipeline()
transformer_question_classifier.add_node(component=embedding_retriever, name="EmbeddingRetriever", inputs=["Query"])
transformer_question_classifier.add_node(
component=TransformersQueryClassifier(model_name_or_path="shahrukhx01/question-vs-statement-classifier"),
name="QueryClassifier",
inputs=["EmbeddingRetriever"],
)
transformer_question_classifier.add_node(component=reader, name="QAReader", inputs=["QueryClassifier.output_1"])
transformer_question_classifier.draw("question_classifier.png")
# Run only the QA reader on the question query
res_1 = transformer_question_classifier.run(query="Who is the father of <NAME>?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_1, details="minimum")
res_2 = transformer_question_classifier.run(query="<NAME> was the daughter of a Lord.")
print("ES Results" + "\n" + "=" * 15)
print_documents(res_2)
# + [markdown] id="sJcWRK4Hwyx2"
# ## Standalone Query Classifier
# Below we run queries classifiers standalone to better understand their outputs on each of the three types of queries
# + colab={"base_uri": "https://localhost:8080/"} id="XhPMEqBzxA8V" outputId="be3ba2ac-b557-4cb3-9eed-41928f644b6e"
# Here we create the keyword vs question/statement query classifier
from haystack.nodes import TransformersQueryClassifier
queries = [
"arya stark father",
"jon snow country",
"who is the father of <NAME>",
"which country was jon snow filmed?",
]
keyword_classifier = TransformersQueryClassifier()
for query in queries:
result = keyword_classifier.run(query=query)
if result[1] == "output_1":
category = "question/statement"
else:
category = "keyword"
print(f"Query: {query}, raw_output: {result}, class: {category}")
# + colab={"base_uri": "https://localhost:8080/"} id="l4eH3SSaxZ0O" outputId="53384108-3d4c-4547-d32a-4a63aa1b74a0"
# Here we create the question vs statement query classifier
from haystack.nodes import TransformersQueryClassifier
queries = [
"<NAME> was the father of A<NAME>.",
"<NAME> was filmed in United Kingdom.",
"who is the father of <NAME>?",
"Which country was jon snow filmed in?",
]
question_classifier = TransformersQueryClassifier(model_name_or_path="shahrukhx01/question-vs-statement-classifier")
for query in queries:
result = question_classifier.run(query=query)
if result[1] == "output_1":
category = "question"
else:
category = "statement"
print(f"Query: {query}, raw_output: {result}, class: {category}")
# + [markdown] id="9VMUHR-BN-gl" pycharm={"name": "#%% md\n"}
# ## Conclusion
#
# The query classifier gives you more possibility to be more creative with the pipelines and use different retrieval nodes in a flexible fashion. Moreover, as in the case of Question vs Statement classifier you can also choose the queries which you want to send to the reader.
#
# Finally, you also have the possible of bringing your own classifier and plugging it into either `TransformersQueryClassifier(model_name_or_path="<huggingface_model_name_or_file_path>")` or using the `SklearnQueryClassifier(model_name_or_path="url_to_classifier_or_file_path_as_pickle", vectorizer_name_or_path="url_to_vectorizer_or_file_path_as_pickle")`
# -
# ## About us
#
# This [Haystack](https://github.com/deepset-ai/haystack/) notebook was made with love by [deepset](https://deepset.ai/) in Berlin, Germany
#
# We bring NLP to the industry via open source!
# Our focus: Industry specific language models & large scale QA systems.
#
# Some of our other work:
# - [German BERT](https://deepset.ai/german-bert)
# - [GermanQuAD and GermanDPR](https://deepset.ai/germanquad)
# - [FARM](https://github.com/deepset-ai/FARM)
#
# Get in touch:
# [Twitter](https://twitter.com/deepset_ai) | [LinkedIn](https://www.linkedin.com/company/deepset-ai/) | [Slack](https://haystack.deepset.ai/community/join) | [GitHub Discussions](https://github.com/deepset-ai/haystack/discussions) | [Website](https://deepset.ai)
#
# By the way: [we're hiring!](https://www.deepset.ai/jobs)
| tutorials/Tutorial14_Query_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Working with FB Prophet
# ## begin with [Forecasting Growth](https://facebookincubator.github.io/prophet/docs/forecasting_growth.html) example from FB page
#
# Lok at time series of daily page views for the Wikipedia page of the R programming language. The csv is available [here](https://github.com/facebookincubator/prophet/blob/master/examples/example_wp_R.csv)
# + deletable=true editable=true
wp_R_dataset_url = 'https://github.com/facebookincubator/prophet/blob/master/examples/example_wp_R.csv'
wp_R_filename = '../datasets/example_wp_R.csv'
# + deletable=true editable=true
import pandas as pd
import numpy as np
from fbprophet import Prophet
# + deletable=true editable=true
# NB: this didn't work as of 8/22/17
#import io
#import requests
#s=requests.get(peyton_dataset_url).content
#df=pd.read_csv(io.StringIO(s.decode('utf-8')))#df = pd.read_csv(peyton_dataset_url)
# -
# ### import the data and transform to log-scale
# + deletable=true editable=true
df = pd.read_csv(wp_R_filename)
# transform to log scale
df['y']=np.log(df['y'])
df.head()
# + [markdown] deletable=true editable=true
# By default, Prophet uses a linear model for its forecast. When forecasting growth, there is usually some maximum achievable point: total market size, total population size, etc. This is called the carrying capacity, and the forecast should saturate at this point.
#
# Prophet allows you to make forecasts using a [logistic growth](https://en.wikipedia.org/wiki/Logistic_function) trend model, with a specified carrying capacity. We illustrate this with the log number of page visits to the R (programming language) page on Wikipedia.
# + [markdown] deletable=true editable=true
# We must specify the carrying capacity in a column `cap`. Here we will assume a particular value, but this would usually be set using data or expertise about the market size.
# + deletable=true editable=true
df['cap']=8.5
# -
df.tail()
# The important things to note are that `cap` must be specified for every row in the dataframe, and that it does not have to be constant. If the market size is growing, then `cap` can be an increasing sequence.
#
#
# We then fit the model as before, except pass in an additional argument to specify logistic growth:
# + deletable=true editable=true
m = Prophet(growth='logistic')
m.fit(df)
# -
# We make a dataframe for future predictions as before, except we must also specify the capacity in the future. Here we keep capacity constant at the same value as in the history, and forecast 3 years into the future.
future = m.make_future_dataframe(periods=3*365+1) # covers leap year of 2016 this way
# %matplotlib inline
future['cap'] = 8.5
fcst = m.predict(future)
m.plot(fcst);
# +
# m.plot?
# -
# ### go back and make a prediction using simple linear growth model
m_linear = Prophet()
m_linear.fit(df);
future_lin = m_linear.make_future_dataframe(periods=3*365+1) # covers leap year of 2016 this way
fcst_lin = m_linear.predict(future_lin)
m_linear.plot(fcst_lin);
# ## poke around a little more at what is under the hood of FB prophet
| notebooks/Prophet_ForecastingGrowth_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Welcome to the Hour of CI!
#
# The Hour of Cyberinfrastructure (Hour of CI) project will introduce you to the world of cyberinfrastructure (CI). If this is your first lesson, then we recommend starting with the **[Gateway Lesson](https://www.hourofci.org/gateway-lesson)**, which will introduce you to the Hour of CI project and the eight knowledge areas that make up Cyber Literacy for Geographic Information Science. This is the **Beginner Geospatial Data** lesson.
#
# <div style="float:left;" >
# To start, click on the "Run this cell" button below to setup your Hour of CI environment. It looks like this: <img style="float: right; padding:0px; margin-top:4px; margin-left:5px;" src="../../gateway-lesson/gateway/supplementary/play-button.png" alt="Play button image">
# </div>
# -
# !cd ../..; sh setupHourofCI # Run this cell (button on left) to setup your Hour of CI environment
# + [markdown] slideshow={"slide_type": "-"}
# After your Hour of CI setup is complete, then
#
# <font size="+1"><a style="background-color:blue;color:white;padding:12px;margin:10px;font-weight:bold;" href="gd-1.ipynb">Click here to launch the Beginner Geospatial Data Lesson</a></font>
| beginner-lessons/geospatial-data/Welcome.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:illustrip3d]
# language: python
# name: python3
# ---
# + [markdown] id="toWe1IoH7X35"
# # IllusTrip: Text to Video 3D
#
# Part of [Aphantasia](https://github.com/eps696/aphantasia) suite, made by <NAME> [[eps696](https://github.com/eps696)]
# Based on [CLIP](https://github.com/openai/CLIP) + FFT/pixel ops from [Lucent](https://github.com/greentfrapp/lucent).
# 3D part by [deKxi](https://twitter.com/deKxi), based on [AdaBins](https://github.com/shariqfarooq123/AdaBins) depth.
# thanks to [<NAME>](https://twitter.com/advadnoun), [<NAME>](https://twitter.com/jonathanfly), [@eduwatch2](https://twitter.com/eduwatch2) for ideas.
#
# ## Features
# * continuously processes **multiple sentences** (e.g. illustrating lyrics or poems)
# * makes **videos**, evolving with pan/zoom/rotate motion
# * works with [inverse FFT](https://github.com/greentfrapp/lucent/blob/master/lucent/optvis/param/spatial.py) representation of the image or **directly with RGB** pixels (no GANs involved)
# * generates massive detailed textures (a la deepdream), **unlimited resolution**
# * optional **depth** processing for 3D look
# * various CLIP models
# * can start/resume from an image
#
# + [markdown] id="QytcEMSKBtN-"
# **Run the cell below after each session restart**
#
# Ensure that you're given Tesla T4/P4/P100 GPU, not K80!
# +
# @title General setup
import GPUtil as GPU
from progress_bar import ProgressIPy as ProgressBar
import depth
import transforms
from utils import slice_imgs, derivat, pad_up_to, slerp, checkout, sim_func, latent_anima
from utils import basename, file_list, img_list, img_read, txt_clean, plot_text, old_torch
from clip_fft import to_valid_rgb, fft_image, rfft2d_freqs, img2fft, pixel_image, un_rgb
import lpips
import kornia
from sentence_transformers import SentenceTransformer
import clip
import warnings
import ipywidgets as ipy
from IPython.core.interactiveshell import InteractiveShell
from IPython.display import HTML, Image, display, clear_output
from torch.autograd import Variable
from torchvision import transforms as T
import torchvision
import torch.nn.functional as F
import torch.nn as nn
import torch
from easydict import EasyDict as edict
import shutil
from base64 import b64encode
import PIL
import numpy as np
import imageio
import random
import math
import time
import io
from pathlib import Path
import os
# !pip install ftfy == 5.8 transformers
# !pip install gputil ffpb
# !pip install imageio
# !pip install easydict
# !pip3 install torch == 1.10.0+cu113 torchvision == 0.11.1+cu113 torchaudio == =0.10.0+cu113 - f https: // download.pytorch.org/whl/cu113/torch_stable.html
# !pip install ipywidgets
try:
# !pip3 install googletrans == 3.1.0a0
from googletrans import Translator, constants
translator = Translator()
except:
pass
# # !apt-get -qq install ffmpeg
work_dir = Path.cwd()
illustrip_dir = work_dir / 'illustrip'
Path.mkdir(illustrip_dir, exist_ok=True)
a = edict()
InteractiveShell.ast_node_interactivity = "all"
if os.name != "nt":
from google.colab import output, files
warnings.filterwarnings("ignore")
# !pip install git+https: // github.com/openai/CLIP.git - -no-deps
# !pip install sentence_transformers
# !pip install kornia
# !pip install lpips
# !pip install PyWavelets == 1.1.1
# !pip install git+https: // github.com/fbcotter/pytorch_wavelets
shutil.copy('mask.jpg', illustrip_dir)
depth_mask_file = illustrip_dir / 'mask.jpg'
clear_output()
def save_img(img, fname=None):
img = np.array(img)[:, :, :]
img = np.transpose(img, (1, 2, 0))
img = np.clip(img*255, 0, 255).astype(np.uint8)
if fname is not None:
imageio.imsave(fname, np.array(img))
imageio.imsave('result.jpg', np.array(img))
def makevid(seq_dir, size=None):
seq_dir = Path(seq_dir).as_posix()
char_len = len(basename(img_list(seq_dir)[0]))
out_sequence = seq_dir + '/%0{}d.jpg'.format(char_len)
out_video = seq_dir + '.mp4'
print('.. generating video ..')
# !ffmpeg - y - v warning - i $out_sequence - crf 18 $out_video
data_url = "data:video/mp4;base64," + \
b64encode(open(out_video, 'rb').read()).decode()
wh = '' if size is None else 'width=%d height=%d' % (size, size)
# return """<video %s controls><source src="%s" type="video/mp4"></video>""" % (wh, data_url)
def makevidloop1(seq_dir, size=None):
seq_dir = Path(seq_dir).as_posix()
char_len = len(basename(img_list(seq_dir)[0]))
out_sequence = seq_dir + '/%0{}d.jpg'.format(char_len)
out_video = seq_dir + '.mp4'
print('.. generating video ..')
# !ffmpeg - r 24 - i $out_sequence - filter_complex "[0]reverse[r];[0][r]concat,setpts=N/24/TB" - crf 18 - pix_fmt yuv420p $out_video
data_url = "data:video/mp4;base64," + \
b64encode(open(out_video, 'rb').read()).decode()
wh = '' if size is None else 'width=%d height=%d' % (size, size)
# return """<video %s controls><source src="%s" type="video/mp4"></video>""" % (wh, data_url)
def makevidloop2(seq_dir, size=None):
seq_dir = Path(seq_dir).as_posix()
char_len = len(basename(img_list(seq_dir)[0]))
out_sequence = seq_dir + '/%0{}d.jpg'.format(char_len)
out_video = seq_dir + '.mp4'
print('.. generating video ..')
# !ffmpeg - r 24 - i $out_sequence - c: v h264_nvenc - filter_complex "[0]reverse[r];[0][r]concat,setpts=N/24/TB" - preset slow - rc constqp - qp 30 - pix_fmt yuv420p $out_video
wh = '' if size is None else 'width=%d height=%d' % (size, size)
# return """<video %s controls><source src="%s" type="video/mp4"></video>""" % (wh, data_url)
# Hardware check
gpu = GPU.getGPUs()[0] # XXX: only one GPU on Colab and isn’t guaranteed
print("(GPU RAM {0:.0f}MB | Free {1:.0f}MB)".format(
gpu.memoryTotal, gpu.memoryFree))
# + cellView="form" id="JUvpdy8BWGuM"
# @title Load inputs
# @markdown **Content** (either type a text string, or upload a text file):
# @param {type:"string"}
content = "pink brains connected to computers by a jumble of colorful wires."
upload_texts = False # @param {type:"boolean"}
# @markdown **Style** (either type a text string, or upload a text file):
style = "" # @param {type:"string"}
upload_styles = False # @param {type:"boolean"}
# @markdown For non-English languages use Google translation:
translate = False # @param {type:"boolean"}
# @markdown Resume from the saved `.pt` snapshot, or from an image
# @markdown (resolution settings below will be ignored in this case):
resume = False # @param {type:"boolean"}
if upload_texts:
print('Upload main text file')
uploaded = files.upload()
text_file = list(uploaded)[0]
texts = list(uploaded.values())[0].decode().split('\n')
texts = [tt.strip()
for tt in texts if len(tt.strip()) > 0 and tt[0] != '#']
print(' main text:', text_file, len(texts), 'lines')
workname = txt_clean(basename(text_file))
else:
texts = [content]
workname = txt_clean(content)[:44]
if upload_styles:
print('Upload styles text file')
uploaded = files.upload()
text_file = list(uploaded)[0]
styles = list(uploaded.values())[0].decode().split('\n')
styles = [tt.strip()
for tt in styles if len(tt.strip()) > 0 and tt[0] != '#']
print(' styles:', text_file, len(styles), 'lines')
else:
styles = [style]
if resume:
print('Upload file to resume from')
resumed = files.upload()
resumed_filename = list(resumed)[0]
resumed_bytes = list(resumed.values())[0]
assert len(texts) > 0 and len(texts[0]) > 0, 'No input text[s] found!'
tempdir = illustrip_dir / workname
counter = 1
workname_num = workname
while tempdir.exists():
counter += 1
workname_num = workname + "_" + str(counter)
tempdir = illustrip_dir / workname_num
Path.mkdir(tempdir)
print('main dir', tempdir)
print(workname_num)
# + [markdown] id="PQFGziYKtHSa"
# **`content`** (what to draw) is your primary input; **`style`** (how to draw) is optional, if you want to separate such descriptions.
# If you load text file[s], the imagery will interpolate from line to line (ensure equal line counts for content and style lists, for their accordance).
# + cellView="form" id="64mlBCAYeOrB"
# @title Main settings
sideX = 1920 # @param {type:"integer"} 1920
sideY = 1080 # @param {type:"integer"} 1080
steps = 100 # @param {type:"integer"}
frame_step = 100 # @param {type:"integer"}
# @markdown > Config
method = 'RGB' # @param ['FFT', 'RGB']
# @param ['ViT-B/16', 'ViT-B/32', 'RN101', 'RN50x16', 'RN50x4', 'RN50']
model = 'ViT-B/32'
# Default settings
if method == 'RGB':
align = 'overscan'
colors = 2
contrast = 1.2
sharpness = -1.
aug_noise = 0.
smooth = False
else:
align = 'uniform'
colors = 1.8
contrast = 1.1
sharpness = 1.
aug_noise = 2.
smooth = True
interpolate_topics = True
style_power = 1.
samples = 200
save_step = 1
learning_rate = 1.
aug_transform = 'custom'
similarity_function = 'cossim'
macro = 0.4
enforce = 0.
expand = 0.
zoom = 0.012
shift = 10
rotate = 0.8
distort = 0.3
animate_them = True
sample_decrease = 1.
DepthStrength = 0.
print(' loading CLIP model..')
model_clip, _ = clip.load(model, jit=old_torch())
modsize = model_clip.visual.input_resolution
xmem = {'ViT-B/16': 0.25, 'RN50': 0.5,
'RN50x4': 0.16, 'RN50x16': 0.06, 'RN101': 0.33}
if model in xmem.keys():
sample_decrease *= xmem[model]
clear_output()
print(' using CLIP model', model)
# + [markdown] id="JIWNmmd5uuSn"
# **`FFT`** method uses inverse FFT representation of the image. It allows flexible motion, but is either blurry (if smoothed) or noisy (if not).
# **`RGB`** method directly optimizes image pixels (without FFT parameterization). It's more clean and stable, when zooming in.
# There are few choices for CLIP `model` (results do vary!). I prefer ViT-B/32 for consistency, next best bet is ViT-B/16.
#
# **`steps`** defines the length of animation per text line (multiply it to the inputs line count to get total video duration in frames).
# `frame_step` sets frequency of the changes in animation (how many frames between motion keypoints).
#
#
# + [markdown] id="f3Sj0fxmtw6K"
# ## Other settings [optional]
# + cellView="form" id="P88_xcpAIXlq"
# @title Run this cell to override settings, if needed
# @markdown [to roll back defaults, run "Main settings" cell again]
style_power = 1. # @param {type:"number"}
overscan = True # @param {type:"boolean"}
align = 'overscan' if overscan else 'uniform'
interpolate_topics = True # @param {type:"boolean"}
# @markdown > Look
colors = 2 # @param {type:"number"}
contrast = 1.2 # @param {type:"number"}
sharpness = 0. # @param {type:"number"}
# @markdown > Training
samples = 500 # @param {type:"integer"}
save_step = 12 # @param {type:"integer"}
learning_rate = 2 # @param {type:"number"}
# @markdown > Tricks
aug_transform = 'custom' # @param ['elastic', 'custom', 'none']
aug_noise = 0. # @param {type:"number"}
macro = 0.4 # @param {type:"number"}
enforce = 0.5 # @param {type:"number"}
expand = 0.5 # @param {type:"number"}
# @param ['cossim', 'spherical', 'mixed', 'angular', 'dot']
similarity_function = 'cossim'
# @markdown > Motion
zoom = 0.012 # @param {type:"number"}
shift = 10 # @param {type:"number"}
rotate = 0.8 # @param {type:"number"}
distort = 0.3 # @param {type:"number"}
animate_them = True # @param {type:"boolean"}
smooth = True # @param {type:"boolean"}
if method == 'RGB':
smooth = False
# + [markdown] id="QYrJTb8xDm9C"
# `style_power` controls the strength of the style descriptions, comparing to the main input.
# `overscan` provides better frame coverage (needed for RGB method).
# `interpolate_topics` changes the subjects smoothly, otherwise they're switched by cut, making sharper transitions.
#
# Decrease **`samples`** if you face OOM (it's the main RAM eater), or just to speed up the process (with the cost of quality).
# `save_step` defines, how many optimization steps are taken between saved frames. Set it >1 for stronger image processing.
#
# Experimental tricks:
# `aug_transform` applies some augmentations, which quite radically change the output of this method (and slow down the process). Try yourself to see which is good for your case. `aug_noise` augmentation [FFT only!] seems to enhance optimization with transforms.
# `macro` boosts bigger forms.
# `enforce` adds more details by enforcing similarity between two parallel samples.
# `expand` boosts diversity (up to irrelevant) by enforcing difference between prev/next samples.
#
# Motion section:
# `shift` is in pixels, `rotate` in degrees. The values will be used as limits, if you mark `animate_them`.
#
# `smooth` reduces blinking, but induces motion blur with subtle screen-fixed patterns (valid only for FFT method, disabled for RGB).
# + [markdown] id="YdVubN0vb3TU"
# ## Add 3D depth [optional]
# + cellView="form" id="vl-rm1Nm03lK"
import gdown
# !pip install gdown
# deKxi:: This whole cell contains most of whats needed,
# with just a few changes to hook it up via frame_transform
# (also glob_step now as global var)
# I highly recommend performing the frame transformations and depth *after* saving,
# (or just the depth warp if you prefer to keep the other affines as they are)
# from my testing it reduces any noticeable stretching and allows the new areas
# revealed from the changed perspective to be filled/detailed
# pretrained models: Nyu is much better but Kitti is an option too
depth_model = 'nyu' # @ param ["nyu","kitti"]
DepthStrength = 0.01 # @param{type:"number"}
MaskBlurAmt = 33 # @param{type:"integer"}
save_depth = False # @param{type:"boolean"}
size = (sideY, sideX)
# @markdown NB: depth computing may take up to ~3x more time. Read the comments inside for more info.
# @markdown Courtesy of [deKxi](https://twitter.com/deKxi)
if DepthStrength > 0:
if not os.path.exists("AdaBins_nyu.pt"):
# !gdown https: // drive.google.com/uc?id = 1lvyZZbC9NLcS8a__YPcUP7rDiIpbRpoF
if not os.path.exists('AdaBins_nyu.pt'):
# !wget https: // www.dropbox.com/s/tayczpcydoco12s/AdaBins_nyu.pt
# if depth_model=='kitti' and not os.path.exists(os.path.join(workdir_depth, "pretrained/AdaBins_kitti.pt")):
# # !gdown https://drive.google.com/uc?id=1HMgff-FV6qw1L0ywQZJ7ECa9VPq1bIoj
if save_depth:
depthdir = os.path.join(tempdir, 'depth')
os.makedirs(depthdir, exist_ok=True)
print('depth dir', depthdir)
else:
depthdir = None
depth_infer, depth_mask = depth.init_adabins(
model_path='AdaBins_nyu.pt', mask_path='mask.jpg', size=size)
def depth_transform(img_t, img_np, depth_infer, depth_mask, size, depthX=0, scale=1., shift=[0, 0], colors=1, depth_dir=None, save_num=0):
# d X/Y define the origin point of the depth warp, effectively a "3D pan zoom", [-1..1]
# plus = look ahead, minus = look aside
dX = 100. * shift[0] / size[1]
dY = 100. * shift[1] / size[0]
# dZ = movement direction: 1 away (zoom out), 0 towards (zoom in), 0.5 stay
dZ = 0.5 + 23. * (scale[0]-1)
# dZ += 0.5 * float(math.sin(((save_num % 70)/70) * math.pi * 2))
if img_np is None:
img2 = img_t.clone().detach()
par, imag, _ = pixel_image(img2.shape, resume=img2)
img2 = to_valid_rgb(imag, colors=colors)()
img2 = img2.detach().cpu().numpy()[0]
img2 = (np.transpose(img2, (1, 2, 0))) # [h,w,c]
img2 = np.clip(img2*255, 0, 255).astype(np.uint8)
image_pil = T.ToPILImage()(img2)
del img2
else:
image_pil = T.ToPILImage()(img_np)
size2 = [s//2 for s in size]
img = depth.depthwarp(img_t, image_pil, depth_infer, depth_mask, size2, depthX, [
dX, dY], dZ, rescale=0.5, clip_range=2, save_path=depth_dir, save_num=save_num)
return img
# + [markdown] id="YZFuwNux8oEg"
# ## Generate
# + cellView="form" id="Nq0wA-wc-P-s"
# @title Generate
# Delete memory from previous runs
# !nvidia-smi - caa
for var in ['model_clip', 'perceptor', 'z']:
try:
del globals()[var]
except:
pass
try:
import gc
gc.collect()
except:
pass
try:
torch.cuda.empty_cache()
except:
pass
# End Clear Memory
if aug_transform == 'elastic':
trform_f = transforms.transforms_elastic
sample_decrease *= 0.95
elif aug_transform == 'custom':
trform_f = transforms.transforms_custom
sample_decrease *= 0.95
else:
trform_f = transforms.normalize()
if enforce != 0:
sample_decrease *= 0.5
samples = int(samples * sample_decrease)
print(' using %s method, %d samples' % (method, samples))
if translate:
translator = Translator()
def enc_text(txt):
if translate:
txt = translator.translate(txt, dest='en').text
emb = model_clip.encode_text(clip.tokenize(txt).cuda()[:77])
return emb.detach().clone()
# Encode inputs
count = 0 # max count of texts and styles
key_txt_encs = [enc_text(txt) for txt in texts]
count = max(count, len(key_txt_encs))
key_styl_encs = [enc_text(style) for style in styles]
count = max(count, len(key_styl_encs))
assert count > 0, "No inputs found!"
# # !rm -rf $tempdir
# os.makedirs(tempdir, exist_ok=True)
# opt_steps = steps * save_step # for optimization
glob_steps = count * steps # saving
if glob_steps == frame_step: frame_step = glob_steps // 2 # otherwise no motion
outpic = ipy.Output()
outpic
if method == 'RGB':
if resume:
img_in = imageio.imread(resumed_bytes) / 255.
params_tmp = torch.Tensor(img_in).permute(
2, 0, 1).unsqueeze(0).float().cuda()
params_tmp = un_rgb(params_tmp, colors=1.)
sideY, sideX = img_in.shape[0], img_in.shape[1]
else:
params_tmp = torch.randn(1, 3, sideY, sideX).cuda() # * 0.01
else: # FFT
if resume:
if os.path.splitext(resumed_filename)[1].lower()[1:] in ['jpg', 'png', 'tif', 'bmp']:
img_in = imageio.imread(resumed_bytes)
params_tmp = img2fft(img_in, 1.5, 1.) * 2.
else:
params_tmp = torch.load(io.BytesIO(resumed_bytes))
if isinstance(params_tmp, list): params_tmp = params_tmp[0]
params_tmp = params_tmp.cuda()
sideY, sideX = params_tmp.shape[2], (params_tmp.shape[3]-1)*2
else:
params_shape = [1, 3, sideY, sideX//2+1, 2]
params_tmp = torch.randn(*params_shape).cuda() * 0.01
params_tmp = params_tmp.detach()
# function() = torch.transformation(linear)
# animation controls
if animate_them:
if method == 'RGB':
m_scale = latent_anima([1], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[-0.3])
m_scale = 1 + (m_scale + 0.3) * zoom # only zoom in
else:
m_scale = latent_anima([1], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[0.6])
m_scale = 1 - (m_scale-0.6) * zoom # ping pong
m_shift = latent_anima([2], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[0.5,0.5])
m_angle = latent_anima([1], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[0.5])
m_shear = latent_anima([1], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[0.5])
m_shift = (m_shift-0.5) * shift * abs(m_scale-1.) / zoom
m_angle = (m_angle-0.5) * rotate * abs(m_scale-1.) / zoom
m_shear = (m_shear-0.5) * distort * abs(m_scale-1.) / zoom
def get_encs(encs, num):
cnt = len(encs)
if cnt == 0: return []
enc_1 = encs[min(num, cnt-1)]
enc_2 = encs[min(num+1, cnt-1)]
return slerp(enc_1, enc_2, steps)
def frame_transform(img, size, angle, shift, scale, shear):
if old_torch(): # 1.7.1
img = T.functional.affine(img, angle, shift, scale, shear, fillcolor=0, resample=PIL.Image.BILINEAR)
img = T.functional.center_crop(img, size)
img = pad_up_to(img, size)
else: # 1.8+
img = T.functional.affine(img, angle, shift, scale, shear, fill=0, interpolation=T.InterpolationMode.BILINEAR)
img = T.functional.center_crop(img, size) # on 1.8+ also pads
return img
global img_np
img_np = None
prev_enc = 0
stop_on_next_loop = False # Make sure GPU memory doesn't get corrupted from cancelling the run mid-way through, allow a full frame to complete
def process(num):
if stop_on_next_loop:
break
global params_tmp, img_np, opt_state, params, image_f, optimizer, pbar
if interpolate_topics:
txt_encs = get_encs(key_txt_encs, num)
styl_encs = get_encs(key_styl_encs, num)
else:
txt_encs = [key_txt_encs[min(num, len(key_txt_encs)-1)][0]] * steps if len(key_txt_encs) > 0 else []
styl_encs = [key_styl_encs[min(num, len(key_styl_encs)-1)][0]] * steps if len(key_styl_encs) > 0 else []
if len(texts) > 0: print(' ref text: ', texts[min(num, len(texts)-1)][:80])
if len(styles) > 0: print(' ref style: ', styles[min(num, len(styles)-1)][:80])
for ii in range(steps):
glob_step = num * steps + ii # saving/transforming
# animation: transform frame, reload params
h, w = sideY, sideX
# transform frame for motion
scale = m_scale[glob_step] if animate_them else 1-zoom
trans = tuple(m_shift[glob_step]) if animate_them else [0, shift]
angle = m_angle[glob_step][0] if animate_them else rotate
shear = m_shear[glob_step][0] if animate_them else distort
if method == 'RGB':
if DepthStrength > 0:
params_tmp = depth_transform(params_tmp, img_np, depth_infer, depth_mask, size, DepthStrength, scale, trans, colors, depthdir, glob_step)
params_tmp = frame_transform(params_tmp, (h,w), angle, trans, scale, shear)
params, image_f, _ = pixel_image([1,3,h,w], resume=params_tmp)
img_tmp = None
else: # FFT
if old_torch(): # 1.7.1
img_tmp = torch.irfft(params_tmp, 2, normalized=True, signal_sizes=(h,w))
if DepthStrength > 0:
img_tmp = depth_transform(img_tmp, img_np, depth_infer, depth_mask, size, DepthStrength, scale, trans, colors, depthdir, glob_step)
img_tmp = frame_transform(img_tmp, (h,w), angle, trans, scale, shear)
params_tmp = torch.rfft(img_tmp, 2, normalized=True)
else: # 1.8+
if type(params_tmp) is not torch.complex64:
params_tmp = torch.view_as_complex(params_tmp)
img_tmp = torch.fft.irfftn(params_tmp, s=(h,w), norm='ortho')
if DepthStrength > 0:
img_tmp = depth_transform(img_tmp, img_np, depth_infer, depth_mask, size, DepthStrength, scale, trans, colors, depthdir, glob_step)
img_tmp = frame_transform(img_tmp, (h,w), angle, trans, scale, shear)
params_tmp = torch.fft.rfftn(img_tmp, s=[h,w], dim=[2,3], norm='ortho')
params_tmp = torch.view_as_real(params_tmp)
params, image_f, _ = fft_image([1,3,h,w], resume=params_tmp, sd=1.)
image_f = to_valid_rgb(image_f, colors=colors)
del img_tmp
optimizer = torch.optim.Adam(params, learning_rate)
# optimizer = torch.optim.AdamW(params, learning_rate, weight_decay=0.01, amsgrad=True)
if smooth is True and num + ii > 0:
optimizer.load_state_dict(opt_state)
# get encoded inputs
txt_enc = txt_encs[ii % len(txt_encs)].unsqueeze(0) if len(txt_encs) > 0 else None
styl_enc = styl_encs[ii % len(styl_encs)].unsqueeze(0) if len(styl_encs) > 0 else None
# optimization
for ss in range(save_step):
loss = 0
noise = aug_noise * (torch.rand(1, 1, *params[0].shape[2:4], 1)-0.5).cuda() if aug_noise > 0 else 0.
img_out = image_f(noise)
img_sliced = slice_imgs([img_out], samples, modsize, trform_f, align, macro)[0]
out_enc = model_clip.encode_image(img_sliced)
if method == 'RGB': # empirical hack
loss += 1.5 * abs(img_out.mean((2,3)) - 0.45).mean() # fix brightness
loss += 1.5 * abs(img_out.std((2,3)) - 0.17).sum() # fix contrast
if txt_enc is not None:
loss -= sim_func(txt_enc, out_enc, similarity_function)
if styl_enc is not None:
loss -= style_power * sim_func(styl_enc, out_enc, similarity_function)
if sharpness != 0: # mode = scharr|sobel|naive
loss -= sharpness * derivat(img_out, mode='naive')
# loss -= sharpness * derivat(img_sliced, mode='scharr')
if enforce != 0:
img_sliced = slice_imgs([image_f(noise)], samples, modsize, trform_f, align, macro)[0]
out_enc2 = model_clip.encode_image(img_sliced)
loss -= enforce * sim_func(out_enc, out_enc2, similarity_function)
del out_enc2; torch.cuda.empty_cache()
if expand > 0:
global prev_enc
if ii > 0:
loss += expand * sim_func(prev_enc, out_enc, similarity_function)
prev_enc = out_enc.detach().clone()
del img_out, img_sliced, out_enc; torch.cuda.empty_cache()
optimizer.zero_grad()
loss.backward()
optimizer.step()
# save params & frame
params_tmp = params[0].detach().clone()
if smooth is True:
opt_state = optimizer.state_dict()
with torch.no_grad():
img_t = image_f(contrast=contrast)[0].permute(1,2,0)
img_np = torch.clip(img_t*255, 0, 255).cpu().numpy().astype(np.uint8)
imageio.imsave(os.path.join(tempdir, '%05d.jpg' % glob_step), img_np, quality=95)
shutil.copy(os.path.join(tempdir, '%05d.jpg' % glob_step), 'result.jpg')
outpic.clear_output()
with outpic:
display(Image('result.jpg'))
del img_t
pbar.upd()
params_tmp = params[0].detach().clone()
outpic = ipy.Output()
outpic
pbar = ProgressBar(glob_steps)
for i in range(count):
process(i)
# -
# makevid(tempdir)
# makevidloop1(tempdir)
# makevidloop2(tempdir)
# import torch
# print(torch.__version__)
# torch.cuda.is_available()
# torch.cuda.empty_cache()
# # !nvidia-smi -L
# # !nvidia-smi
# !nvidia-smi
# +
# !nvidia-smi - caa
for var in ['model_clip']:
try:
del globals()[var]
except:
pass
try:
import gc
gc.collect()
except:
pass
try:
torch.cuda.empty_cache()
except:
pass
# -
| IllusTrip3D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="xyxgU7hwBN4K"
# ## What is Quickvision ?
# + [markdown] id="_GKAHq41DO4p"
# - Quickvision is a Computer Vision Library built on Top of Torchvision, PyTorch and Lightning
#
# A brief description on how it works.
#
# - It provides Easy to use torch native API, for fit(), train_step(), val_step() of models.
#
# - It is Easily customizable and configurable models with various backbones.
#
# - A complete torch native interface. All models are nn.Module all the training APIs are optional and not binded to models.
#
# - Tensor First library. No abstraction and classes over it !
#
# - A lightning API which helps to accelerate training over multiple GPUs, TPUs.
#
# - A datasets API to common data format very easily and quickly to torch formats.
#
# - A minimal package, with very low dependencies.
#
#
# + [markdown] id="wawtpa3CDq4-"
# ### Let's explore these with a simple Image Classification task.
# + [markdown] id="2BHtveDiD2b9"
# ## Install
# + [markdown] id="memMtSeqD3d9"
# Install directly from GitHub. Very soon it will be available over PyPi.
#
#
# + id="3Kb2KnJiBBQF"
# !pip install -q git+https://github.com/Quick-AI/quickvision.git
# + [markdown] id="x8PxnXi0E7tF"
# ## Some imports
# + id="eKK7lmkOE80m"
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision
import torchvision.transforms as T
from tqdm import tqdm
import pytorch_lightning as pl
# And finally our Hero !
import quickvision
# + [markdown] id="w2X9FYlBEodG"
# ## Create a dataset and dataloader
# + [markdown] id="IeLHW85wEr2l"
# - This process is completely as in PyTorch.
#
# - For example let's take the CIFAR10 dataset available in torchvision.
#
# + [markdown] id="Jnhsjy-wFGfH"
# ### Training and testing transfomrs
#
# Quickivision is not binded to any library for transforms.
#
# Let's keep it simple and use torchvision transforms for now.
# + id="s1q51DuPFFpF"
# Train and validation Transforms which you would like
train_transforms = T.Compose([T.ToTensor(), T.Normalize((0.5,), (0.5,))])
valid_transforms = T.Compose([T.ToTensor(), T.Normalize((0.5,), (0.5,))])
# + id="SBr0O1r7ES8W"
train_set = torchvision.datasets.CIFAR10("./data", download=True, train=True, transform=train_transforms)
valid_set = torchvision.datasets.CIFAR10("./data", download=True, train=False, transform=valid_transforms)
# + id="FKNDI7c2FYU3"
train_loader = torch.utils.data.DataLoader(train_set, batch_size=32, shuffle=True)
valid_loader = torch.utils.data.DataLoader(valid_set, batch_size=32, shuffle=False)
# + [markdown] id="C4pnUs0KFqU6"
# ## Creating model !!
# + [markdown] id="s0mdPX8VFzB4"
# - Quickvision Provides simple functions to create models with pretrained weights.
# + id="Y_ljlgF2FyE4"
from quickvision.models.classification import cnn
# + id="OURry2JrGAJl"
# To create model with imagenet pretrained weights
model = cnn.create_vision_cnn("resnet50", num_classes=10, pretrained="imagenet")
# Alternatively if you don't need pretrained weights
model_bare = cnn.create_vision_cnn("resnet50", num_classes=10, pretrained=None)
# It also supports other weights, do check a list which are supported !
model_ssl = cnn.create_vision_cnn("resnet50", num_classes=10, pretrained="ssl")
# + [markdown] id="q8RZJ_4SG2_o"
# ### Loss Function and Optimizer
# + [markdown] id="B5HX4EtBG4uO"
# - Again this is just like in torch
# + id="mwVXfhsjG20Q"
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=1e-3)
# + [markdown] id="fJa724TuH2a-"
# ## Training
# + [markdown] id="Nq6xJJgWGoDu"
# ### The usual boring PyTorch procedure.
# + [markdown] id="BE3RAwLaLC-j"
# - This too works with Quickvision, we just have models as `nn.Module` .
#
# It is same way as you would train a CNN.
#
# + id="LsrahKZrFmoz"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# + id="LFTyhOYIF-J4" outputId="99bbd549-370e-4bba-d93a-b0fa33907b87" colab={"base_uri": "https://localhost:8080/"}
model = model.to(device)
for epoch in range(2):
for batch_idx, (inputs, target) in enumerate(train_loader):
optimizer.zero_grad()
inputs = inputs.to(device)
target = target.to(device)
out = model(inputs)
loss = criterion(out, target)
loss.backward()
optimizer.step()
print("Done !!")
# And the boring boiler plate continues
# + [markdown] id="NNsunzCzHupY"
# ### The quickvision way of doing it !
# + [markdown] id="1O5XVd9tH5Dh"
# - We have already implemented these boring procedures for you to speed up training !
# + id="Pp8gRByvHmLh"
model = cnn.create_vision_cnn("resnet50", num_classes=10, pretrained="imagenet")
# + id="xwoHZL9MIpwH"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# + id="dLdfd3YuILaJ"
history = cnn.fit(model=model, epochs=2, train_loader=train_loader,
val_loader=valid_loader, criterion=criterion, device=device, optimizer=optimizer)
# + [markdown] id="EDxBb1--I2QE"
# - If you need a granular control over the training.
#
# You can use `train_step` and `val_step` methods.
#
# We calculate commonly used metrics such as `accuracy` here for you !
#
#
# + id="rKpYJvzZItS0"
for epoch in tqdm(range(2)):
print()
print(f"Training Epoch = {epoch}")
train_metrics = cnn.train_step(model, train_loader, criterion, device, optimizer)
print()
print(f"Validating Epoch = {epoch}")
valid_metrics = cnn.val_step(model, valid_loader, criterion, device)
# + [markdown] id="8jz9neAdKMoG"
# ### Train with Lightning !
# + [markdown] id="ca6on8sMKQKv"
# - We have the same logics implemented for PyTorch Lightning as well.
# - This directly allows you to use all Lighning features such as Multi-GPU training, TPU Training, logging etc.
#
# Quickly Prototype with Torch, transfer it to Lightning !
#
#
# + id="QPLTOD4rKO69"
model_imagenet = cnn.lit_cnn("resnet18", num_classes=10, pretrained="imagenet")
gpus = 1 if torch.cuda.is_available() else 0
# Again use all possible Trainer Params from Lightning here !!
trainer = pl.Trainer(gpus=gpus, max_epochs=2)
trainer.fit(model_imagenet, train_loader, valid_loader)
# + [markdown] id="IRAsNz4TLxUY"
# ## Some more Features of Quickvision
# + [markdown] id="Mv36qVgNL0XZ"
# - Support for mixed precision training in PyTorch API over GPUs.
#
# - Supports Detection models such as `Detr`, `FRCNN`, `RetinaNet`.
#
#
#
# + [markdown] id="Riy8NgHqJiAY"
# # Conclusion
# + [markdown] id="d3Cr6NrqJmTN"
# 1. Quickvision allows you to bring your own `Dataset`, `Model` or `Code` Recipe
#
# 2. You may use models, training functions or Dataset loading utilites from quickvision.
#
# 3. Seamless API to connect with Lightning as well.
#
# 4. Faster Experimentation with same control with PyTorch or Lightning.
#
# Visit us [here](https://github.com/Quick-AI/quickvision) over GitHub !
#
# We are happy for new contributions / improvements to our package.
#
# Quickivison is a library built for faster but no compromise PyTorch Training !
#
#
| examples/notebooks/CNNs_with_Quickvision.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tum]
# language: python
# name: conda-env-tum-py
# ---
# #### Variables coded in this notebook: 6, 11, 16, 19
import os, pandas as pd, re
exportdate = 20180327
projectname = 'repract'
df = pd.read_csv(f'../../data/{exportdate}{projectname}.csv')
df.head(2)
basedir = '../../data/freetext'
freetextfiles = os.listdir(basedir)
dfs = {file[:-4]:pd.read_csv(f'{basedir}/{file}') for file in freetextfiles}
dfs.keys()
def add_unique_code(df, func, varname):
df[f'{varname}_coded'] = func(df[varname])
return df
codedir = '../../analysis/freetext'
def write_coded(df, varname, sep=','):
filepath = f'{codedir}/{varname}_coded.csv'
df.to_csv(f'{codedir}/{varname}_coded.csv', index=False, sep=sep)
print(f'File stored at {filepath}.')
# #### Variable 6 (_Other_ for variable 5) - primary working area
# Options shown were:
# * Requirements Engineer
# * Business Analyst
# * Architect
# * Tester / Test Manager
# * Project Manager
# * Developer
# * Product Owner
# * Designer
# * Other (please specify)
def code_var_6(series):
coded_series = []
for value in series:
value = value.lower()
val = None
if re.search('lecturer|phd\scandidate|researcher|r&d', value):
val = 'Researcher'
elif re.search('consultant', value):
val = 'Consultant'
elif re.search('systems?\sengineer', value):
val = 'Architect'
elif re.search('processes', value):
val = 'Process Designer'
elif re.search('design', value):
val = 'Designer'
elif re.search('marketing|iso\s\d+', value):
val = 'Context Roles' # this was: 1 Marketing, 1 Regulator
elif re.search('manag|cto', value):
val = 'Manager'
elif (re.search('different|changing|both|depend(?:s|ing)|combin',
value) or (len(re.findall(',', value)) > 1)):
val = 'Multiple Roles'
else:
raise Exception(f'Difficulty Coding Entry: {value}')
coded_series.append(val)
return coded_series
coded_v6 = add_unique_code(dfs['v_6'], code_var_6, 'v_6')
coded_v6.head()
coded_v6.groupby('v_6_coded').count()[['lfdn']]
assert coded_v6.groupby('v_6_coded').count().v_6.sum() == len(dfs['v_6']['v_6'])
write_coded(coded_v6, 'v_6', sep=';')
# #### Variable 11 - years of experience
def code_var_11(series):
coded_series = []
replace_dict = {'years?\.?|y(?!\w)|about': '',
'six':'6',
'one':'1',
',':'.',
'\+|>':''}
this_year = 2018
for value in series:
value = value.lower()
val = None
try:
val = float(value)
except:
val = value
for k, v in replace_dict.items():
val = re.sub(k, v, val)
try:
val = float(val)
except:
if re.search('since\s(\d{4})', val):
val = 2018 - float(re.search('since\s(\d{4})', val).group(1))
elif re.search('\d+', val):
val = sum([float(x) for x in re.findall('\d+', val)])
else:
raise Exception(f'Difficulty Coding Entry: {value}')
coded_series.append(val)
return coded_series
coded_v11 = add_unique_code(dfs['v_11'], code_var_11, 'v_11')
coded_v11.head()
# %matplotlib notebook
import seaborn as sns
sns.set_style('darkgrid')
import matplotlib.pyplot as plt
coded_v11.v_11_coded.plot.hist(bins=50, color='k', alpha=0.5)
plt.xlabel('Years of Experience');
plt.xlim(0,1)
plt.xticks(range(0,51,5))
plt.yticks(range(0,21,2));
# +
#write_coded(coded_v11, 'v_11', sep=';')
# -
# #### Variable 16 (_Other_ for Variable 15) - class of system
# Options shown were:
# - Software-intensive embedded systems
# - (Business) information systems
# - Hybrid / mix of embedded systems and information systems
# - Other (please specify)
# NB: I feel the coding is somewhat arbitrary - but at least it's transparently arbitrary...
def code_var_16(series):
coded_series = []
for value in series:
value = value.lower()
val = None
if re.search('all.*?above', value):
val = 'Hybrid / mix of embedded systems and information systems'
elif (re.search('c(?:ustomer|onsumer)|online|information', value) # infosys
or re.search('(?<!\w)erp(?!\w)', value)): # infosys, special (and doubtful ;-))
val = '(Business) information systems'
elif re.search('machine|infrastructure|processor', value):
# or would you want to class these as hybrid?
val = 'Hardware'
elif re.search('aeronautics|railway', value): # guessing this one
val = 'Hybrid / mix of embedded systems and information systems'
else:
raise Exception(f'Difficulty Coding Entry: {value}')
coded_series.append(val)
return coded_series
len(dfs['v_16'])
coded_v16 = add_unique_code(dfs['v_16'], code_var_16, 'v_16')
coded_v16.head()
coded_v16.groupby('v_16_coded').count()[['lfdn']]
# +
#write_coded(coded_v16, 'v_16', sep=';')
# -
# #### Variable 19 - Industry Sector (not standardized as in NaPiRE!)
# NB: This is - of course - drastically overfitting, but there's hardly another option (the NaPiRE categories are very problematic and thus should not be reused).
def code_var_19(series):
coded_series = []
for value in series:
value = value.lower()
val = None
if (re.search('mixed|varies(?!\w)|several|ecosystem|(?<!\s)services(?!\s)',
value)
or (len(re.findall(',', value)) > 1) or re.search('and', value)
and not re.search('oil.*?gas|bank.*?fin|ins.*?bank|aero.*?defen|well.*heal', value)):
val = 'Multiple Sectors'
elif re.search('university|research|academi', value):
val = 'Academia'
elif re.search('aero|avi(?:on|at)', value):
val = 'Aeronautics'
elif re.search('automation', value):
val = 'Automation'
elif re.search('automotive', value):
val = 'Automotive'
elif re.search('consult', value):
val = 'Consulting'
elif re.search('e\-?commerc|online', value):
val = 'E-Commerce'
elif re.search('educati', value):
val = 'Education'
elif re.search('energy|(?:oil|gas)(?!\w)', value):
val = 'Energy'
elif re.search('financ|banki|insuran', value):
val = 'Financial Services'
elif re.search('semiconductor|robotics|computer\sengin|industrial\ssys', value):
val = 'Hardware'
elif re.search('medic(?:al|ine)|heal?th|wellness', value):
val = 'Healthcare'
elif re.search('railway|building|pipelines', value):
val = 'Infrastructure'
elif re.search('government|public\s(?!transport)|defen[cs]e', value):
val = 'Public Sector'
elif re.search('software|saas', value):
val = 'Software'
elif re.search('transport|logis\w|marine', value):
val = 'Transportation'
elif re.search('tourism', value):
val = 'Tourism'
elif re.search(('commun|telecom|(?<!\w)ict(?!\w)|(?<!\w)it(?!\w)|(?<!\w)iot(?!\w)|'
+'intranet|electron|network|information'), value):
val = 'ICT'
else:
raise Exception(f'Difficulty Coding Entry: {value}')
coded_series.append(val)
return coded_series
coded_v19 = add_unique_code(dfs['v_19'], code_var_19, 'v_19')
coded_v19.head()
# +
#write_coded(coded_v19, 'v_19', sep=';')
# -
coded_v19.groupby('v_19_coded').count()[['lfdn']]
# The End.
| notebooks/freetextcoding/01a_assigning_codes_short_answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Implement strStr().
#
# Return the index of the first occurrence of needle in haystack, or -1 if needle is not part of haystack.
#
# Example 1:
#
# Input: haystack = "hello", needle = "ll"
# Output: 2
# Example 2:
#
# Input: haystack = "aaaaa", needle = "bba"
# Output: -1
# Clarification:
#
# What should we return when needle is an empty string? This is a great question to ask during an interview.
#
# For the purpose of this problem, we will return 0 when needle is an empty string. This is consistent to C's strstr() and Java's indexOf().
# +
class Solution:
def strStr(self, strings: str, pattern: str) -> int:
if not pattern:
return 0
m,n = len(strings),len(pattern)
i,j = 0,0
while i < m and j < n:
if strings[i] == pattern[j]:
i += 1
j += 1
else:
i = i-j+1
j = 0
if j == n:
return i-j
return -1
# test
s = "mississippi"
p = "issip"
Solution().strStr(s,p)
# -
class Solution:
def strStr(self, s: str, p: str) -> int:
# when pattern is empty, we should return 0
if not p:
return 0
m, n = len(s), len(p)
# lps means the longest proper prefix that same as suffix
# initialize lps array generated from pattern
lps = [0] * n
# compute lps
self.findlps(p, lps, n)
i, j = 0, 0
while i < m and j < n:
# if string matched, move i and j one step ahead
if s[i] == p[j]:
i,j = i+1, j+1
# if string not matched,
else:
# if j backtrack to 0, which indicate that at i-th index
# there is no match, so i move one step ahead
if j == 0:
i += 1
# if j not backtrack to 0, now j backtracks to previous loc in lps
else:
j = lps[j-1]
# finally, if j equals to size of pattern
# which indicates that pattern is fully matched
# return the index (i locates last char in pattern in s to be found)
# thus i should minus the index j (which index the length of pattern)
if j == n:
return i - j
# if no match, return error code -1
return -1
def findlps(self, p, lps, n):
# pre indicates the longest prefix
pre, i = 0,1
# lps[0] is always zero!
while i < n:
if p[i] == p[pre]:
pre += 1
lps[i] = pre
i += 1
else:
# at the very beginnning, lps[0] = 0, and i = i+1
if pre == 0:
lps[i] = 0
i += 1
else:
pre = lps[pre-1]
# test
s = "mississippi"
p = "issip"
Solution().strStr(s,p)
| DSA/string/strStr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parallel programming in Python
#
# It is sometimes stated that parallel code is difficult in Python. However, for most scientific applications, we can achieve parallel code without much effort. In this notebook I will show some simple ways to get parallel code execution in Python.
#
# 1. With NumPy
# 2. With Joblib and multiprocessing
# 3. With Numba
# 4. With Cython
#
# These methods range in complexity from easiest to most difficult.
#
# After discussing Cython, there is a short example with Numba vectorize, which can be used for functions that should be applied element-wise on numerical NumPy arrays.
# ## Parallel code with NumPy
#
# By default, NumPy will dispatch the computations to an efficient BLAS (basic linear algebra subproblem) and LAPACK (Linear Algebra PACKage) implementation. BLAS and LAPACK routines are very efficient linear algebra functions, which are implemented by experts who can get as much speed as possible out of your CPU. There is absolutely no way we compete with these functions if we are doing linear algebra.
#
# A benefit and downside with NumPy is that it will likely parallelise the code for you without your knowledge. Try to compute the matrix product between two large matrices and look at your CPU load. It will likely use all of your hardware threads (hardware threads are essentially cores).
# ## Moving the parallelism to the outer loop
#
# Often, when we program, we have nested loops, like we see below
# +
x = [[] for _ in range(10)]
for i in range(10):
for j in range(10):
x[i].append(i*j)
for row in x:
print(row)
# -
# Here, we have two nested loops, so there are three ways to make this parallel: performing multiple iterations of the outer loop (`for i in range(10)`) simultaneously, performing multiple iterations of the inner loop (`for j in range(10)`) simultaneously, or both.
#
# Generally, we prefer to have the parallel code on the outer loop. That way, each worker has a lot of work to do, and is therefore less likely to stay idle. If there are more hardware threads available than there are iterations on the outer loop, we can split it up and have some of the parallelism on the inner loop as well. However, it is important to make sure that we don't try to do more things in parallel than we have hardware threads available, as otherwise, much time will be spent switching between tasks rather than actually performing the computations.
#
# ## Disabling parallel code execution in NumPy routines
# Unfortunately, if we use a NumPy function in our loop, then that function likely runs in parallel and is using all available hardware threads. To avoid this, we have to set some envionment variables *before* importing NumPy. Specifically, we should set
#
# OMP_NUM_THREADS=NUM_THREADS
# OPENBLAS_NUM_THREADS=NUM_THREADS
# MKL_NUM_THREADS=NUM_THREADS
# VECLIB_MAXIMUM_THREADS=NUM_THREADS
# NUMEXPR_NUM_THREADS=NUM_THREADS
#
# The first variable sets the number of OpenMP threads to `NUM_THREADS`. OpenMP is used by many software packages to obtain parallel code. The next three variables sets the number of threads for NumPy with various BLAS backends. Finally, the last line sets the number of threads for a useful package called NumExpr, which can be used to optimise operations on the form `a*x + b*x - c`. With pure NumPy, such an expression would entail four separate loops, but with NumExpr, it is compiled to a single parallel loop.
#
# We can either set these variables directly from Python, or we can set it as a global environment variable. If we set these variables directly from Python, then we must do it before we (or any other library) import NumPy. If you instead want to set the variables globally on your computer, and you are using Linux, then you can add these lines to your [`~/.profile` file](https://www.quora.com/What-is-profile-file-in-Linux):
#
# ```
# OPENBLAS_NUM_THREADS=1
# MKL_NUM_THREADS=1
# VECLIB_MAXIMUM_THREADS=1
# NUMEXPR_NUM_THREADS=1
# ```
#
# Notice how we did not set the number of OpenMP threads to 1 in the `~/.profile` file! If we did that, then we would likely disable parallelism for most programs that use OpenMP.
#
# **Note that if we set `OMP_NUM_THREADS` to 1, then parallelism with Numba and Cython will not work.**
# +
import os
def set_threads(
num_threads,
set_blas_threads=True,
set_numexpr_threads=True,
set_openmp_threads=False
):
num_threads = str(num_threads)
if not num_threads.isdigit():
raise ValueError("Number of threads must be an integer.")
if set_blas_threads:
os.environ["OPENBLAS_NUM_THREADS"] = num_threads
os.environ["MKL_NUM_THREADS"] = num_threads
os.environ["VECLIB_MAXIMUM_THREADS"] = num_threads
if set_numexpr_threads:
os.environ["NUMEXPR_NUM_THREADS"] = num_threads
if set_openmp_threads:
os.environ["OMP_NUM_THREADS"] = num_threads
set_threads(1)
# -
# Now, we can import numpy to our code and it will only run on only one core.
import numpy as np
# ## Parallel code with Joblib and multiprocessing
#
# Python does not support parallel threading. This means that each Python process can only do one thing at a time. The reason for this lies with the way Python code is run on your computer. Countless hours has been spent trying to remove this limitation, but all sucessfull attempts severly impaired the speed of the language (the most well known attempt is Larry Hasting's [gilectomy](https://github.com/larryhastings/gilectomy)).
#
# Since we cannot run code in parallel within a single Python process, we need to start new processes for each task we wish to compute in parallel and send the relevant information to these processes. This leads to a lot of overhead, and if we plan to have any performance gain using multiple processes, then we should parallelise substantial tasks.
# ### The best approach: Joblib
#
# The best approach to multiprocessing in Python is through the Joblib library. It overcomes some of the shortcomings of multiprocessing (that you may not realise is a problem until you encounter them) at the cost of an extra dependency in your code. Below, we see an example of parallel code with Joblib
# +
from joblib import Parallel, delayed
def f(x):
return x + 2
numbers1 = Parallel(n_jobs=2)(delayed(f)(x) for x in range(10))
print(numbers1)
# -
# Here we see how Joblib can help us parallelise simple for loops. We wrap what we wish to compute in a function and use it in a list comprehension. The `n_jobs` argument specifies how many processes to spawn. If it is a positive number (1, 2, 4, etc.) then it is the number of processes to spawn and if it is a negative number then joblib will spawn (n_cpu_threads + 1 + n_jobs). Thus `n_jobs=-1` will spawn as many processes as there are CPU threads available, `n_jobs=-2` will spawn n-1 CPU threads, etc.
#
# I recommend setting `n_jobs=-2` so you have one CPU thread free to surf the web while you run hard-core experiments on your computer.
# +
numbers1 = Parallel(n_jobs=-2)(delayed(f)(x) for x in range(10))
print(numbers1)
# -
# If we cannot wrap all the logic within a single function, but need to have two separate parallel loops, then we should use the `Parallel` object in a slightly different fashion. If we do the following:
# +
from joblib import Parallel, delayed
def f(x):
return x + 2
numbers1 = Parallel(n_jobs=2)(delayed(f)(x) for x in range(10))
numbers2 = Parallel(n_jobs=2)(delayed(f)(x) for x in range(20))
print(numbers1)
print(numbers2)
# -
# Then we will first create two new Python processes, compute the parallel list comprehension, close these two processess before spawning two new Python processes and computing the second parallel list comprehension. This is obviously not ideal, and we can reuse the pool of processes with a context manager:
# +
with Parallel(n_jobs=2) as parallel:
numbers1 = parallel(delayed(f)(x) for x in range(10))
numbers2 = parallel(delayed(f)(x) for x in range(20))
print(numbers1)
print(numbers2)
# -
# Here, the same processes are used for both list comprehensions!
# ## Async operations with multiprocessing
# An alternative to using Joblib for multiprocessing in Python is to use the builtin multiprocessing module.
# This module is not as user friendly as joblib, and may break with weird error messages.
# +
import multiprocessing
def add_2(x):
return x + 2
with multiprocessing.Pool(4) as p:
print(p.map(add_2, range(10)))
# -
# Here, we see that multiprocessing also requires us to wrap the code we wish to run in parallel in a function.
#
# However, one particular of multiprocessing is that it requires all inputs to be picklable. That means that we cannot use output a factory function and you may also have problems with using multiprocessing with instance methods. Below is an example that fails.
# +
import multiprocessing
def add(x):
def add_x(y):
return x + y
return add_x
add_2 = add(2)
print(add_2(2))
with multiprocessing.Pool(4) as p:
p.map(add_2, range(10))
# -
# We see that local functions aren't picklable, however, the same code runs with joblib:
print(Parallel(n_jobs=4)(delayed(add_2)(x) for x in range(10)))
# ### So why use multiprocessing?
# Unfortunately, Joblib blocks the python interpreter, so that while the other processess run, no work can be done on the mother process. See the example below:
# +
from time import sleep, time
def slow_function(x):
sleep(3)
return x
start_time = time()
Parallel(n_jobs=-2)(delayed(slow_function)(i) for i in range(10))
print(time() - start_time)
# -
# Meanwhile, with multiprocessing, we can start the processes, let those run in the background, and do other tasks while waiting. Here is an example
with multiprocessing.Pool(6) as p:
# Start ten processes.
# The signature for the apply_async method is as follows
# apply_async(function, args, kwargs)
# the args iterable is fed into the function using tuple unpacking
# the kwargs iterable is fed into the function using dictionary unpacking
tasks = [p.apply_async(slow_function, [i]) for i in range(10)]
prev_ready = 0
num_ready = sum(task.ready() for task in tasks)
while num_ready != len(tasks):
if num_ready != prev_ready:
print(f"{num_ready} out of {len(tasks)} completed tasks")
prev_ready = num_ready
num_ready = sum(task.ready() for task in tasks)
results = [task.get() for task in tasks]
print(results)
# This means that if you have to do some post processing of the output of the parallel loop, then you can start doing that with the elements that are done. Here is a very simple example
# +
with multiprocessing.Pool(6) as p:
# Start ten processes.
# The signature for the apply_async method is as follows
# apply_async(function, args, kwargs)
# the args iterable is fed into the function using tuple unpacking
# the kwargs iterable is fed into the function using dictionary unpacking
tasks = [p.apply_async(slow_function, [i]) for i in range(10)]
finished = {}
while len(finished) != len(tasks):
for i, task in enumerate(tasks):
if task.ready():
if i not in finished:
print(f"Task {i} just finished, its result was {task.get()}")
finished[i] = task.get()
print([finished[i] for i in range(10)])
# -
# ## Parallelising with Numba
#
# Numba is an almost magical tool that lets us write Python code which is just in time compiled (JIT) to machine code using LLVM. Consequently, we can get C-speed with our Python code!
#
# Unfortunately, the price of this black magic is that Numba doesn't support the whole Python language. It supports only a subset of it. Especially if you enable `nopython` mode to get the largest speedups.
#
# Let us start by looking at some simple non-parallel Numba tricks
# +
import numba
def python_sum(A):
s = 0
for i in range(A.shape[0]):
s += A[i]
return s
@numba.jit
def numba_normal_sum(A):
s = 0
for i in range(A.shape[0]):
s += A[i]
return s
# -
x = np.random.randn(10000)
print("Pure python")
# %timeit python_sum(x)
print("Numba")
# %timeit numba_normal_sum(x)
# We see that the numba compiled code is much faster than the plain python code (more than x100). There are two downsides with writing code this way.
#
# 1. Error messages are cryptic
# 1. JIT compiled functions can only call other JIT compiled functions and a subset of all Python, NumPy and SciPy.
# * See the documentation for more info on this.
#
# However, we can also parallelise code with Numba, using the `numba.prange` (parallel range) function. This code is cheekily stolen from the numba [documentation](https://numba.pydata.org/numba-doc/0.11/prange.html).
@numba.jit(parallel=True, nogil=True)
def numba_parallel_sum(A):
s = 0
for i in numba.prange(A.shape[0]):
s += A[i]
return s
x = np.random.randn(10000)
numba_normal_sum(x) # compile it once
numba_parallel_sum(x) # compile it once
print("Pure python")
# %timeit python_sum(x)
print("\nNumba")
# %timeit numba_normal_sum(x)
print("\nParallel Numba")
# %timeit numba_parallel_sum(x)
# Here, we see that the performance actually deteriorates by parallising the code! This is because of the extra overhead needed to organise multiple workers. However, sometimes parallelising code this way can lead to significant speedups (especially if each iteration is costly).
#
# We can use Cython to reduce the overhead that we experience with the parallel sum.
#
# **Note:** It is difficult to use Numba on the outer loop, so if you have costly outer loops, then you should use Joblib to have the parallelism there instead.
# ## Parallel code with Cython
# Finally, we look at Cython to optimise and paralellise code. Cython is a language that will let us write Python-like code that is transpiled into a Python C extension. This means several things:
#
# 1. We can get C speed without much effort
# 2. Cython is a superset of Python, so any Python code can be compiled
# 3. It is easier to write Cython, but it requires manual compilation.
#
# The first two points here make Cython a very attractive alternative. However, the final point can be very problematic. Whenever you make a change to a Cython file, you need to compile it again. This is generally done via a `setup.py` file that contains the build instructions for your Cython files.
#
# Luckily, we can prototype some Cython code in a notebook, using the `%%cython` magic command.
# %load_ext Cython
# The code below is just copy pasted from above, but the inclusion of the `%%cython` cell magic means that the code is now compiled and can run faster its pure Python counterpart. Just copy pasting code this way will not massively improve runtime.
# + language="cython"
# import numpy as np
#
# def cython_sum(A):
# s = 0
# for i in range(A.shape[0]):
# s += A[i]
#
# return s
# -
# Unfortunately, the code above is sill running in the CPython virtual machine which are a lot slower than a pure C function. Let us fix this, to do that, we avoid any Python data types, and only use the C counterparts.
# + language="cython"
# cimport cython
# cimport numpy as np
# import numpy as np
#
# @cython.boundscheck(False) # Do not check if Numpy indexing is valid
# @cython.wraparound(False) # Deactivate negative Numpy indexing.
# cpdef smart_cython_sum(np.ndarray[np.float_t] A):
# # ^ Notice cpdef instead of def. Define it as a C function and Python function simultaneously.
# cdef float s = 0
# cdef int i
# for i in range(A.shape[0]):
# s += A[i]
#
# return s
# -
# Now, we can look at how to make this run in parallel. To do this, we need OpenMP (which runs a bit differently on Linux and Windows, so your first line may be different if you are using a Windows machine).
# + magic_args="--compile-args=-fopenmp --link-args=-fopenmp --force" language="cython"
# from cython.parallel import prange
# import numpy as np
#
# cimport cython
# cimport numpy as np
# cimport openmp
#
#
# @cython.boundscheck(False) # Do not check if Numpy indexing is valid
# @cython.wraparound(False) # Deactivate negative Numpy indexing.
# cpdef parallel_cython_sum(np.ndarray[np.float_t] A):
# # ^ Notice cpdef instead of def. Define it as a C function and Python function simultaneously.
# cdef float s = 0
# cdef int i
# for i in prange(A.shape[0], nogil=True, num_threads=8):
# s += A[i]
#
#
# return s
#
# -
x = np.random.randn(10000)
print("Pure python")
# %timeit python_sum(x)
print("\nNumba")
# %timeit numba_normal_sum(x)
print("\nParallel Numba")
# %timeit numba_parallel_sum(x)
print("\nNaive Cython")
# %timeit cython_sum(x)
print("\nSmart Cython")
# %timeit smart_cython_sum(x)
print("\nParallel Cython")
# %timeit parallel_cython_sum(x)
# From this, we see a couple of things
#
# 1. There is little difference between pure Python and naive Cython
# 1. Pure Numba is faster than sophisticated Cython for this example
# 1. The parallel code in Cython has much less overhead than the parallel code in Numba
#
# In general, it is difficult to say if Numba or Cython will be fastest. The reason for this is that Numba may or may not be able to lift the CPython virtual machine. If it isn't able to do so, it will often be much slower than Cython.
#
# Thus, if Numba works, it is often as good as, if not better than Cython. You should therefore start with Numba, and if that doesn't provide a good enough speed up, then you can try Cython.
#
# Long functions are often easier to implement in Cython and small ones are often best implemented in Numba. A big downside with Cython (that cannot be stressed enough) is that it adds huge overhead for building and distributing the code. I therefore discourage the use of Cython for code that will be made public unless all other options are tested first.
# ## Vectorisation with Numba
# Finally, we will look at vectorisation of functions with Numba. Vectorisation is when we wish to apply the same function to all elements of an array. For example, the `exp` function in NumPy is a vectorised function.
#
# Let us create a vectorised function to compute the Mandelbrot set.
# +
@np.vectorize
def compute_mandelbrot_np(x):
C = x
for i in range(20):
if abs(x) >= 4:
return i
x = x**2 + C
return -1
@numba.vectorize([numba.int32(numba.complex128)], target="cpu")
def compute_mandelbrot_cpu(x):
C = x
for i in range(20):
if abs(x) >= 4:
return i
x = x**2 + C
return -1
@numba.vectorize([numba.int32(numba.complex128)], target="parallel")
def compute_mandelbrot_parallel(x):
C = x
for i in range(20):
if abs(x) >= 4:
return i
x = x**2 + C
return -1
# +
X = -0.235
Y = 0.827
R = 4.0e-1
x = np.linspace(X - R, X + R, 100)
y = np.linspace(Y - R, Y + R, 100)
xx, yy = np.meshgrid(x, y)
zz = xx + 1j*yy
compute_mandelbrot_cpu(zz) # Compile once
compute_mandelbrot_parallel(zz) # Compile once
print("Single core NumPy")
# %timeit compute_mandelbrot_np(zz)
print("Single core Numba")
# %timeit compute_mandelbrot_cpu(zz)
print("Multi core Numba")
# %timeit compute_mandelbrot_parallel(zz)
# -
# Here, we see not only the effect of just in time compiling our function but also that all our CPU cores are fully utilised when vectorizing functions! Let us plot a section of the mandelbrot set!
import matplotlib.pyplot as plt
# %config InlineBackend.figure_format = 'retina'
# +
X = -0.235
Y = 0.827
R = 4.0e-1
x = np.linspace(X - R, X + R, 1000)
y = np.linspace(Y - R, Y + R, 1000)
xx, yy = np.meshgrid(x, y)
zz = xx + 1j*yy
mandelbrot = compute_mandelbrot_parallel(zz)
plt.figure(figsize=(10, 10), dpi=300)
plt.imshow(mandelbrot)
plt.axis('off')
plt.show()
| assets/notebooks/2020-06-13-parallel-python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/seldoncode/Python_CoderDojo/blob/main/Python_CoderDojo15.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="M6LYA7QGNu4G"
# # Trabajando con archivos ```.py``` en Visual Studio Code
# ## Librería ```os```
# * Importamos la librería que nos permite acceder a los comandos del sistema operativo.
# * Vamos a limpiar la pantalla con:
# - clear en Linux
# - cls en Windows
# * Si no se iguala a una variable aparece un 0
# * En Jupyter Notebook no funciona bien en una celda lo de limpiar la pantalla pero si se ejecuta el fichero .py en una terminal si limpia la terminal.
# * El último comando se asigna a una variable para evitar que se vea un cero al final, si se ejecuta en la terminal.
# + id="vDCMvj7hNKn3"
import os
x = os.system("clear") # en Windows es cls
# + [markdown] id="fRn7fMo6F1al"
# ### Creamos el fichero saludo.py
# * El fichero se crea usando un editor de código como por ejemplo **[Visual Studio Code](https://code.visualstudio.com/)**.
# * Se graba en una carpeta que hemos llamado *mypy*
# * La carpeta está en el repositorio de GitHub llamado:
# - https://github.com/seldoncode/Python_CoderDojo
# * El fichero es este:
# - https://github.com/seldoncode/Python_CoderDojo/mypy/saludo.py
# * El fichero se ejecuta desde la terminal o desde la terminal integrada en Visual Studio Code.
# * Al ejecutarlo desde Jupyter Notebook no veremos lo mismo, ya que en la terminal se va limpiando la pantalla después de escribir cada print y en Jupyter no se limpia.
# + colab={"base_uri": "https://localhost:8080/"} id="reni-o9PO2u8" outputId="2aae6d24-1ff9-4ac8-e358-8a7affdf3f1e"
import os
import time
os.system("clear") # en Windows es cls
print("¡Hola!")
time.sleep(2)
os.system("clear")
print("¿Qué tal?")
time.sleep(2)
os.system("clear")
print("Bye, bye.")
time.sleep(2)
# + [markdown] id="fdYdnO5gVfMU"
# ### Simular movimiento
# * Crear el archivo movimiento1.py para la versión 1
# * Crear el archivo movimiento2.py para la versión 2
# * Al ejecutarlo en la terminal simula el movimiento de un personaje.
# * Son cuatro figuras de un personaje que se repiten
# * En cada figura vamos introduciendo un espacio más para ir simulando el movimiento
# + [markdown] id="OE4UJC4LcSGU"
# #### Versión 1. Sin bucle
# + id="2jzzYIIzVr8z" colab={"base_uri": "https://localhost:8080/"} outputId="54d5a3bb-20fa-4019-ea36-aaccb00be529"
# #!/usr/bin/env python
import os
import time
os.system("clear") # en Windows es cls
print()
print(" o ")
print(" /|\ ")
print(" / | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" (|\ ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" | \ ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|) ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" / | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" (|\ ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" | \ ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|) ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" / | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" (|\ ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" | \ ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|) ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" / | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" (|\ ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" | \ ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|) ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" / | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" (|\ ")
print(" | | ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|\ ")
print(" | \ ")
time.sleep(.3)
os.system("clear")
print()
print(" o ")
print(" /|) ")
print(" | | ")
time.sleep(.3)
os.system("clear")
# + [markdown] id="NJU3daD5cVW-"
# #### Versión 2. Con bucle
# * Introducimos un bucle ```for```
# * El archivo se llamará movimiento2.py
# + id="0iiqDfAFciot"
# #!/usr/bin/env python
import os
import time
espacios = ""
for i in range(20):
time.sleep(.2)
os.system("clear")
print()
print(f"{espacios} o ")
print(f"{espacios} /|\ ")
print(f"{espacios} / | ")
espacios += " "
time.sleep(.2)
os.system("clear")
print()
print(f"{espacios} o ")
print(f"{espacios} (|\ ")
print(f"{espacios} | | ")
espacios += " "
time.sleep(.2)
os.system("clear")
print()
print(f"{espacios} o ")
print(f"{espacios} /|\ ")
print(f"{espacios} | \ ")
espacios += " "
time.sleep(.2)
os.system("clear")
print()
print(f"{espacios} o ")
print(f"{espacios} /|) ")
print(f"{espacios} | | ")
espacios += " "
# + [markdown] id="U-5GkG7QT0-t"
# ### Juego de Muertos y Heridos
# * Comercialmente existe el Mastermind que se juega con colores
# * Nosotros emplearemos los números del 0 al 9.
# * Jugamos contra la máquina
# * La máquina piensa un número de **cuatro** cifras **sin repetición** empleando los diez dígitos entre 0 y 9.
# * Por ejemplo, piensa el número secreto 6293
# * El jugador intenta adivinar el número secreto
# * En cada tirada se van diciendo números de cuatro cifras **sin repetición**
# * Al decir un número se trata de ver cuantos muertos y heridos hay:
# - Un Muerto se produce cuando se ha adivinado un dígito y justo en su sitio
# - Un Herido se produce cuando se ha adividando un dígito pero no está en su sitio.
# - Por ejemplo, si el número secreto es 6293 y el jugador para comprobar dice: 1234, la máquina responde con el resultado: 1M1H, el muerto (M) es el 2 y el herido (H) es el 3.
# - Si el jugador para comprobar dice: 6230, la máquina responde con el resultado: 2M1H, los muertos (M) son el 6 y el 2, el herido (H) es el 3.
# * Si después de 15 intentos no se adivina el número secreto el juego termina
# * Se podrá salir en cualquier momento si tecleamos un valor que contenga algún caracter que no sea numérico.
# + id="VNGknOnyWmn1" colab={"base_uri": "https://localhost:8080/"} outputId="80bb455e-8e73-498b-cd95-d1849c203879"
import random
random.seed()
print("============= Juego de los Muertos y Heridos =============")
print("Dispone de 15 intentos para adivinar el número secreto de cuatro dígitos sin repetición.")
print("Puede salir en cualquier momento tecleando cualquier caracter no numérico.")
secreto = [str(x) for x in random.sample(range(10), 4)]
#print(secreto) # descomentar esta línea para hacer trampas
tirada = 0
jugando = True
while jugando:
tirada += 1
incorrectos = True # inicialmente pensamos que los número que dice el jugador están incorrectos
while incorrectos: # mientras pensemos que están incorrectos pediremos que introduzca un número sin repetición y de 4 dígitos
n = input(f"Tirada {tirada}: Diga un número de cuatro dígitos entre 0 y 9, sin repetir digitos: ")
if not(n.isnumeric()) or len(n) != 4:
print("Fin del juego.")
incorrectos=False # para salirnos del while interno
jugando=False # ya nos queremos salir
intento = list(n) # ['1', '2', '3', '4']
if sorted(intento) == sorted(list(set(intento))): # para evitar que el número que diga el jugador tenga repetidos
incorrectos = False
if not(jugando): break # si queremos salir, ya hemos dicho que jugando=False y ya nos saliemos del while principal
resultado = ""
m = h = 0 # inicializamos los muertos y heridos
for i in range(4):
if intento[i] in secreto: h += 1
if intento[i] == secreto[i]: m += 1; h -= 1
resultado = f"{m}M{h}H"
print(f"{intento[0]+intento[1]+intento[2]+intento[3]} : {resultado}")
if resultado == "4M0H":
print(f"Felicidades: ha adivinado el número secreto {secreto[0]+secreto[1]+secreto[2]+secreto[3]} en {tirada} tiradas.")
jugando = False
elif tirada==15:
print("Se han agotado los 15 intetos. Fin del juego.")
jugando = False
| Python_CoderDojo15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def get_descriptor_model(shape):
model = VGG16(include_top=False, input_shape=(224,224,3), weights='imagenet')
# Freeze all the layers
for layer in model.layers[:]:
layer.trainable = False
init_weights = keras.initializers.he_normal()
output = model.get_layer('block5_conv3').output
output = Conv2D(64, 3, padding = 'valid', use_bias = True, kernel_initializer=init_weights)(output)
output = BatchNormalization(axis = -1)(output)
output = Activation('relu')(output)
output = Conv2D(128, 3, padding = 'valid', use_bias = True, kernel_initializer=init_weights)(output)
output = BatchNormalization(axis = -1)(output)
output = Activation('relu')(output)
output = Conv2D(128,3, padding = 'valid', use_bias = True, kernel_initializer=init_weights)(output)
output = BatchNormalization(axis = -1)(output)
output = Activation('relu')(output)
output = Dropout(0.3)(output)
output = Conv2D(128, 8, padding = 'valid', use_bias = True, kernel_initializer=init_weights)(output)
output = Reshape((128,))(output)
model = Model(model.input, output)
newInput = Input(shape)
i_concat = Concatenate()([newInput, newInput, newInput])
i_resize = Lambda(lambda image: ktf.image.resize_images(image, (224,224)))(i_concat)
newOutput = model(i_resize)
model = Model(newInput, newOutput)
return model
| Improved Models/Modified VGG-16 Model Architecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing what happens in Union master
# One disadvantage to collecting all the simulation in the Union_master component, is that it is not possible to insert monitors between the parts to check on the beam. This issue is addressed by adding logger components that can record scattering and absorption events that occurs during the simulation. This notebook will show examples on the usage of loggers and their features.
# ### Set up materials and geometry to investigate
# First we set up the same mock cryostat we created in the advanced geometry tutorial to have an interesting system to investigate using the loggers.
from mcstasscript.interface import instr, functions, plotter
instrument = instr.McStas_instr("python_tutorial", input_path="run_folder")
# +
Al_inc = instrument.add_component("Al_inc", "Incoherent_process")
Al_inc.sigma = 0.0082
Al_inc.unit_cell_volume = 66.4
Al_pow = instrument.add_component("Al_pow", "Powder_process")
Al_pow.reflections = '"Al.laz"'
Al = instrument.add_component("Al", "Union_make_material")
Al.process_string = '"Al_inc,Al_pow"'
Al.my_absorption = 100*0.231/66.4 # barns [m^2 E-28] * Å^3 [m^3 E-30] = [m E-2], correct with factor 100.
Sample_inc = instrument.add_component("Sample_inc", "Incoherent_process")
Sample_inc.sigma = 3.4176
Sample_inc.unit_cell_volume = 1079.1
Sample_pow = instrument.add_component("Sample_pow", "Powder_process")
Sample_pow.reflections = '"Na2Ca3Al2F14.laz"'
Sample = instrument.add_component("Sample", "Union_make_material")
Sample.process_string = '"Sample_inc,Sample_pow"'
Sample.my_absorption = 100*2.9464/1079.1
src = instrument.add_component("source", "Source_div")
src.xwidth = 0.01
src.yheight = 0.035
src.focus_aw = 0.01
src.focus_ah = 0.01
instrument.add_parameter("wavelength", value=5.0, comment="Wavelength in [Ang]")
src.lambda0="wavelength"
src.dlambda="0.01*wavelength"
src.flux = 1E13
sample_geometry = instrument.add_component("sample_geometry", "Union_cylinder")
sample_geometry.yheight = 0.03
sample_geometry.radius = 0.0075
sample_geometry.material_string='"Sample"'
sample_geometry.priority = 100
sample_geometry.set_AT([0,0,1], RELATIVE=src)
container = instrument.add_component("sample_container", "Union_cylinder", RELATIVE=sample_geometry)
container.yheight = 0.03+0.003 # 1.5 mm top and button
container.radius = 0.0075 + 0.0015 # 1.5 mm sides of container
container.material_string='"Al"'
container.priority = 99
container_lid = instrument.add_component("sample_container_lid", "Union_cylinder")
container_lid.set_AT([0, 0.0155, 0], RELATIVE=container)
container_lid.yheight = 0.004
container_lid.radius = 0.013
container_lid.material_string='"Al"'
container_lid.priority = 98
inner_wall = instrument.add_component("cryostat_wall", "Union_cylinder")
inner_wall.set_AT([0,0,0], RELATIVE=sample_geometry)
inner_wall.yheight = 0.12
inner_wall.radius = 0.03
inner_wall.material_string='"Al"'
inner_wall.priority = 80
inner_wall_vac = instrument.add_component("cryostat_wall_vacuum", "Union_cylinder")
inner_wall_vac.set_AT([0,0,0], RELATIVE=sample_geometry)
inner_wall_vac.yheight = 0.12 - 0.008
inner_wall_vac.radius = 0.03 - 0.002
inner_wall_vac.material_string='"Vacuum"'
inner_wall_vac.priority = 81
outer_wall = instrument.add_component("outer_cryostat_wall", "Union_cylinder")
outer_wall.set_AT([0,0,0], RELATIVE=sample_geometry)
outer_wall.yheight = 0.15
outer_wall.radius = 0.1
outer_wall.material_string='"Al"'
outer_wall.priority = 60
outer_wall_vac = instrument.add_component("outer_cryostat_wall_vacuum", "Union_cylinder")
outer_wall_vac.set_AT([0,0,0], RELATIVE=sample_geometry)
outer_wall_vac.yheight = 0.15 - 0.01
outer_wall_vac.radius = 0.1 - 0.003
outer_wall_vac.material_string='"Vacuum"'
outer_wall_vac.priority = 61
instrument.print_components()
# -
instrument.show_components("Work directory")
# ## Adding Union logger components
# Union logger components need to be added before the *Union_master* component, as the master need to record the necessary information when the simulation is being performed. There are two different kind of Union logger components, the *loggers* that record scattering and the *abs_loggers* that record absorption. They have similar parameters and user interface. Here is a list of the currently available loggers:
#
# - Union_logger_1D
# - Union_logger_2D_space
# - Union_logger_2D_space_time
# - Union_logger_3D_space
# - Union_logger_2D_kf
# - Union_logger_2D_kf_time
# - Union_logger_2DQ
#
# - Union_abs_logger_1D_space
# - Union_abs_logger_1D_space_tof
# - Union_abs_logger_2D_space
#
# The most commonly used logger is probably the *Union_logger_2D_space*, this component records spatial distribution of scattering, here are the available parameters.
instrument.component_help("Union_logger_2D_space")
# ### Setting up a 2D_space logger
# One can select which two axis to record using *D_direction_1* and *D_direction_2*, and the range with for example *D1_min* and *D1_max*. When spatial information is recorded it is also important to place the logger at an appropriate position, here we center it on the sample position.
# +
logger_zx = instrument.add_component("logger_space_zx", "Union_logger_2D_space", RELATIVE=sample_geometry)
logger_zx.D_direction_1 = '"z"'
logger_zx.D1_min = -0.12
logger_zx.D1_max = 0.12
logger_zx.n1 = 300
logger_zx.D_direction_2 = '"x"'
logger_zx.D2_min = -0.12
logger_zx.D2_max = 0.12
logger_zx.n2 = 300
logger_zx.filename = '"logger_zx.dat"'
logger_zy = instrument.add_component("logger_space_zy", "Union_logger_2D_space", RELATIVE=sample_geometry)
logger_zy.D_direction_1 = '"z"'
logger_zy.D1_min = -0.12
logger_zy.D1_max = 0.12
logger_zy.n1 = 300
logger_zy.D_direction_2 = '"y"'
logger_zy.D2_min = -0.12
logger_zy.D2_max = 0.12
logger_zy.n2 = 300
logger_zy.filename = '"logger_zy.dat"'
master = instrument.add_component("master", "Union_master")
# -
# ### Running the simulation
# If mpi is installed, one can add mpi=N where N is the number of cores available to speed up the simulation.
data = instrument.run_full_instrument(ncount=1E7, foldername="data_folder/union_loggers",
increment_folder_name=True,
parameters={"wavelength" : 3.0})
functions.name_plot_options("logger_space_zx", data, log=True, orders_of_mag=4)
functions.name_plot_options("logger_space_zy", data, log=True, orders_of_mag=4)
plotter.make_sub_plot(data)
# ### Interpreting the data
# The zx logger views the cryostat from the top, while the zy loggers shows it from the side. These are histograms of scattered intensity, and it is clear the majority of the scattering happens in the direct beam. There are however scattering events in all parts of our mock cryostat, as neutrons that scattered in either the sample or cryostat walls could go in any direction due to the incoherent scattering. The aluminium and sample also have powder scattering, so some patterns can be seen from the debye scherrer cones.
# ## Logger targets
# It is possible to attach a logger to a certain geometry, or even a list of geometries using the *target_geometry* parameter. In that way one can for example view the scattering in the sample environment, while ignoring the sample. It is also possible to select a number of specific scattering processes to investigate with the *target_process* parameter. This is especially useful when working with a single crystal process, that only scatters when the Bragg condition is met.
#
# Let us modify our existing loggers to view certain parts of the simulated system, and then rerun the simulation. If mpi is installed, one can add mpi=N where N is the number of cores available to speed up the simulation.
# +
logger_zx.target_geometry = '"outer_cryostat_wall,cryostat_wall"'
logger_zy.target_geometry = '"sample_geometry"'
data = instrument.run_full_instrument(ncount=1E7, foldername="data_folder/union_loggers",
increment_folder_name=True,
parameters={"wavelength" : 3.0})
# -
functions.name_plot_options("logger_space_zx", data, log=True, orders_of_mag=4)
functions.name_plot_options("logger_space_zy", data, log=False)
plotter.make_sub_plot(data)
# ## Scattering order
# All loggers also have the option to only record given scattering orders. For example only record the second scattering.
# - order_total : Match given number of scattering events, counting all scattering events in the system
# - order_volume : Match given number of scattering events, only counting events in the current volume
# - order_volume_process : Match given number of scattering events, only counting events in current volume with current process
#
# We can modify our previous loggers to test out these features. The zx logger viewing from above will keep the target, but we remove the sample target on the zy logger, which is done by setting the *taget_geometry* to NULL. We choose to look at the second scattering event.
# +
logger_zx.order_total = 2
logger_zy.target_geometry = '"NULL"'
logger_zy.order_total = 2
data = instrument.run_full_instrument(ncount=1E7, foldername="data_folder/union_loggers",
increment_folder_name=True, mpi=4,
parameters={"wavelength" : 3.0})
# -
functions.name_plot_options("logger_space_zx", data, log=True, orders_of_mag=3)
functions.name_plot_options("logger_space_zy", data, log=True, orders_of_mag=3)
plotter.make_sub_plot(data)
# ## Demonstration of additional logger components
# Here we add a few more loggers to showcase what kind of information that can be displayed.
# - 1D logger that logs scattered intensity as function of time
# - 2D abs_logger that logs absorption projected onto the scattering plane
# - 2DQ logger that logs scattering vector projected onto the scattering plane
# - 2D kf logger that logs final wavevector projected onto the scattering plane
# +
logger_1D = instrument.add_component("logger_1D", "Union_logger_1D", before="master")
logger_1D.variable = '"time"'
logger_1D.min_value = 0.0006
logger_1D.max_value = 0.0012
logger_1D.n1 = 300
logger_1D.filename = '"logger_1D_time.dat"'
abs_logger_zx = instrument.add_component("abs_logger_space_zx", "Union_abs_logger_2D_space",before="master")
abs_logger_zx.set_AT([0,0,0], RELATIVE=sample_geometry)
abs_logger_zx.D_direction_1 = '"z"'
abs_logger_zx.D1_min = -0.12
abs_logger_zx.D1_max = 0.12
abs_logger_zx.n1 = 300
abs_logger_zx.D_direction_2 = '"x"'
abs_logger_zx.D2_min = -0.12
abs_logger_zx.D2_max = 0.12
abs_logger_zx.n2 = 300
abs_logger_zx.filename = '"abs_logger_zx.dat"'
logger_2DQ = instrument.add_component("logger_2DQ", "Union_logger_2DQ", before="master")
logger_2DQ.Q_direction_1 = '"z"'
logger_2DQ.Q1_min = -5.0
logger_2DQ.Q1_max = 5.0
logger_2DQ.n1 = 200
logger_2DQ.Q_direction_2 = '"x"'
logger_2DQ.Q2_min = -5.0
logger_2DQ.Q2_max = 5.0
logger_2DQ.n2 = 200
logger_2DQ.filename = '"logger_2DQ.dat"'
logger_2D_kf = instrument.add_component("logger_2D_kf", "Union_logger_2D_kf", before="master")
logger_2D_kf.Q_direction_1 = '"z"'
logger_2D_kf.Q1_min = -2.5
logger_2D_kf.Q1_max = 2.5
logger_2D_kf.n1 = 200
logger_2D_kf.Q_direction_2 = '"x"'
logger_2D_kf.Q2_min = -2.5
logger_2D_kf.Q2_max = 2.5
logger_2D_kf.n2 = 200
logger_2D_kf.filename = '"logger_2D_kf.dat"'
# -
# ### Runnig the simulation
# We now rerun the simulation with the new loggers. If mpi is installed, one can add mpi=N where N is the number of cores available to speed up the simulation.
data = instrument.run_full_instrument(ncount=1E7, foldername="data_folder/union_loggers",
increment_folder_name=True, mpi=4,
parameters={"wavelength" : 3.0})
# +
functions.name_plot_options("logger_space_zx", data, log=True, orders_of_mag=3)
functions.name_plot_options("logger_space_zy", data, log=True, orders_of_mag=3)
functions.name_plot_options("abs_logger_space_zx", data, log=True, orders_of_mag=3)
functions.name_plot_options("logger_1D", data, log=True, orders_of_mag=3)
functions.name_plot_options("logger_2DQ", data, log=True, orders_of_mag=3)
functions.name_plot_options("logger_2D_kf", data, log=True, orders_of_mag=3)
plotter.make_sub_plot(data[0:2])
plotter.make_sub_plot(data[2:4])
plotter.make_sub_plot(data[4:6])
# -
# ## Interpreting the data
# We see the scattered intensity as a function of time, here the peaks correspond to the direct beam intersecting the sides of the cryostat and sample. The source used release all neutrons at time 0, so it is a perfect pulse.
#
# The absorption monitor shows an image very similar to the scattered intensity, but this could be very different, for example when using materials meant as shielding.
#
# The 2D scattering vector is interesting, it shows a small sphere made of vertical lines, these are powder Bragg peaks. Since the wavevector is almost identical for all incoming neutrons, the first scattering can only access this smaller region of the space. The larger circle is incoherent scattering from second and later scattering events, where the incoming wavevector could be any direction since a scattering already happened.
#
# The 2D final wavevector plot shows mainly the powder Bragg peaks.
| notebooks/5_McStas/Union_tutorial_3_loggers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SkinAnaliticAI, Skin Cancer Detection with AI Deep Learning
#
# ## __Evaluation of Harvard Dataset with different AI classiffication techniques using FastClassAI papeline__
# Author: __<NAME>__
# <EMAIL>
# License: __MIT__
# ttps://opensource.org/licenses/MIT
# Copyright (C) 2021.01.30 <NAME>
# # PART 1. Setting Up Project Enviroment
# ---
# * __Alternative 1.__ CLONE SkinAnaliticAI projects or FastClassAI project from github
# https://github.com/PawelRosikiewicz/SkinAnaliticAI
#
# * __Alternative 2.__ Create file structure and copy/past content of src and notebook folders to corresponding directories,
# * For that solution, you need to create the file structure for storing scripts, notebooks, input data, etc... for FastClasAI pipeline, you may modify, basedir manually in each notebook, if necessary.
# * follow the instruction below
#
# ## Step 1. Create basedir file for your project, eg myproject/
# * then navigate to that file, and follow the instructions below,
# ## Step 2. Setup FastClassAI directory structure in basedir
# +
# imports,
import os # allow changing, and navigating files and folders,
import sys
import re # module to use regular expressions,
import glob # lists names in folders that match Unix shell patterns
# basedir
basedir = os.path.dirname(os.getcwd())
os.chdir(basedir)
sys.path.append(basedir)
print(basedir) # shoudl be ../myproject/
# create folders holing different types of data por notebooks,
files_to_create = {
"for whatever I dont use but wish to keep": os.path.join(basedir, "bin"),
"for random notes and materials created on project development": os.path.join(basedir, "notes"),
# ....
"for jupyter notebooks": os.path.join(basedir, "notebooks"),
"for tfhub model": os.path.join(basedir, "models"),
# ...
"for tools in .py format": os.path.join(basedir, "src"),
"IMPORTANT : HERE YOU MUST COPY ALL .py FILES with my functions": os.path.join(basedir, "src/utils"),
"for config files": os.path.join(basedir, "src/configs"),
# ....
"to store data and resuls": os.path.join(basedir, "data"),
"here you will donwload raw images and other files form the source": os.path.join(basedir, "data/raw"),
"to store matrices with extracted features": os.path.join(basedir, "data/interim"),
"for final results": os.path.join(basedir, "data/results")
}
# ....
for file_function in list(files_to_create.keys()):
try:
os.mkdir(files_to_create[file_function])
except:
print("file", file_function, " - - - was already created")
# -
# ## Step 3. Copy/past src
# * copy/past current notbook into basedir/notebooks;
# * copy/past config files to basedir/src/configs
# * copy/past .py toolts to basedir/src/
# * if avaibale, copy/past results into basedir/data/results
# * if avaibale, copy/past downloaded tf hub models into basedir/models (each model is one folder)
# ## Step 4. Test whether you can import one of my functions
# to test it, just type:
from src.utils.feature_extraction_tools import encode_images
# ---
# # Part 2. DOWNLOAD THE DATA AND TF-HUB MODELS FOR TRANFER LEARING
# ---
# ## Step 1. __Download Input data__
# ### __Dataset Description__
#
# * __Dataset Name__
# * The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions
# * __Source__
# * https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/DBW86T
# * __Size__
# * Apoprox 3GB
# * __Dataset Version__
# * HAM10000 dataset, has only one version, at the time of this project development, that was published in 2018.
# * __Related Publications__
# * <NAME>., <NAME>. & <NAME>. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 5, 180161 (2018). doi: 10.1038/sdata.2018.161 https://www.nature.com/articles/sdata2018161
# * __License__
# * Non-Commercial purposes only,
# * for more details;
# https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/DBW86T
#
#
# ### __Donwload Instructions__
# * data (images and metadata) can be found at the botton of the source site, in section download
# * Harvards site, contains 6 files that can be donwloaded. The followinbg three are required for that project:
# * HAM10000_images_part_1.zip
# * HAM10000_images_part_2.zip
# * HAM10000_metadata.tab
# * Unpack the files, and store all in basedir/data/raw
# ## Step 2. __Download Tf-hub Models used for feature extraction__
#
# ### NOTES
# * In order to work more reliably, I donwloaded several pretrained models for feature extraction from images, from tf-hub
# * My function, in section __Data Preparation__, can also use urls, however, it may be problematic in case of slow internet connection or reteated feature extractions perfomed on different data subsets (timeout occures frequently in these cases)
# * Important: the funciton that I implemented in section __Data Preparation__ for feature extraction, accepts models constructed with TF1 and TF2.
#
# ### __Module Description__
#
# * __Module name used in the project__
# * BiT_M # working name resnet,
# * __Full Module Name__
# * bit_m-r101x1_1
# * __url__
# * https://tfhub.dev/google/bit/m-r101x1/1
# * __Info__
# * __Input Image size__
# * (?, 224, 224, 3)
# * __Output Feature Number__
# * (?, 2048)
# * __Short Description__
# * Big Transfer (BiT) is a recipe for pre-training image classification models on large supervised datasets and efficiently fine-tuning them on any given target task. The recipe achieves excellent performance on a wide variety of tasks, even when using very few labeled examples from the target dataset.
# * This module implements the R101x1 architecture (ResNet-101), trained to perform multi-label classification on ImageNet-21k, a dataset with 14 milion images labeled with 21,843 classes. Its outputs are the 2048-dimensional feature vectors, before the multi-label classification head. This model can be used as a feature extractor or for fine-tuning on a new target task.
# * Instructions:
# module = hub.KerasLayer("https://tfhub.dev/google/bit/m-r101x1/1")
# images = ... # A batch of images with shape [batch_size, height, width, 3].
# features = module(images) # Features with shape [batch_size, 2048].
# ---
# # PART 3. Prepare Config Files - examples below
# ---
# The goal of that part is to define dataset names, dataset varinat names, what tf hub models you use, colors you asign to each class in a project etc...
#
# ## CONFIG FILES
# * location: __basedir/src/configs/project_configs__
# * there are 4 basic configs files that must be prepared
# * __tfhub_configs.py__
# * file that contains info on tf hub modules used for feature extraction
# * __project_configs.py__
# * basic description of the dataset
# * __dataset_configs.py__
# * contains dictionaries used to label images in each class, provide colors etc...
# * and select classes for statistics
# * __config_functions.py__
# * .py file with special functions used to select files for data processing and module training,
# * additionally there is a config file that contains model parameters used when training various ai models
# * this will be descibed later on,
#
# ## Notes
# * config files with CLASS_COLORS, and CLASS_DESCRIPTION, were prepared based on,
# * Links from: https://dermoscopedia.org
# * <NAME>., <NAME>. & <NAME>. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 5, 180161 (2018). doi: 10.1038/sdata.2018.161 https://www.nature.com/articles/sdata2018161
#
#
# ### Step 1. prepare tfhub_configs.py
#
# * this config file contains one dictiory TFHUB_MODELS
# * it is used for extracting features from images using dowlonaded tf hub modules,
# * each module has unique name and working name that may be more descriptive and used on plots,
# * the modules can be donwloaded from tf-hub and stored in basedir/models, or you may add "module_url" to each distionary that is also accepted by FastClassAI function,
#
# +
# config, ...........................................................................................
# Purpose: create config file for tf-hub module used,
# Localization: tfhub_configs.py
# values:
# "module_name" : str, name used on plots, and for file saving
# "working_name" : str, alternative to module_name (eg shorter), not used in my projects
# "file_name" : str, the name of the file donwloaded from tfhub, wiht a given module, (can be custom)
# "module_url" : str, url, to the module on tfhub
# "input_size" : tuple, (height, width) in pixes
# "output_size" : int, lenght of vector with extracted features,
# "note" : str, notes, whatether you this is important, for other users
#
# IMPORTANT, KEY NAME MUST BE THE SAME AS module name
# here is an example for BiT_M_Resnet101 module
TFHUB_MODELS = {
"BiT_M_Resnet101":{
"module_name": "BiT_M_Resnet101",
"working_name": "resnet",
"file_name": "bit_m-r101x1_1",
"module_url":"https://tfhub.dev/google/bit/m-r101x1/1",
"input_size": (224, 224),
"output_size": 2048,
"note":"tested on swissroads dataset, where it worked very well"
}
}# end
# -
# ### create project_configs.py
# * two variables are the most important:
# * PROJECT_NAME : just a string with a solid project name that will be usxed in the project
# * CLASS_DESCRIPTION : that contains description of each class in the original data, plus extra information such as links to external datasources, and class_description (created manually) that may be very usefull later on in the project, while evaliating the results or in EDA
#
# +
# config, ...........................................................................................
PROJECT_NAME = "SkinAnaliticAI_Harvard_dataset_evaluation"
# config, ...........................................................................................
# CLASS_DESCRIPTION
# Purpose: information on each class, used for creating new class arrangment and for providing info on each class,
# Localization: project_configs.py
#
#. "key" : str, class name used in original dataset downloaded form databse
# "original_name" : str, same as the key, but you can introduce other values in case its necessarly
# "class_full_name" : str, class name used on images, saved data etc, (more descriptive then class names, or sometimes the same according to situation)
# "class_group" : str, group of classes, if the classes are hierarchical,
# "class_description" : str, used as notes, or for class description available for the user/client
# "links" : list, with link to more data, on each class
CLASS_DESCRIPTION = {
'akiec':{
"original_name":'akiec',
"class_full_name": "squamous_cell_carcinoma", # prevoisly called "Actinic_keratoses" in my dataset, but ths name is easier to find in online resourses, noth names are correct,
"class_group": "Tumour_Benign",
"class_description": "Class that contains two subclasses:(A) Actinic_Keratoses or (B) Bowen’s disease. Actinic Keratoses (Solar Keratoses) and Intraepithelial Carcinoma (Bowen’s disease) are common non-invasive, variants of squamous cell carcinoma that can be treated locally without surgery. These lesions may progress to invasive squamous cell carcinoma – which is usually not pigmented. Both neoplasms commonly show surface scaling and commonly are devoid of pigment, Actinic keratoses are more common on the face and Bowen’s disease is more common on other body sites. Because both types are induced by UV-light the surrounding skin is usually typified by severe sun damaged except in cases of Bowen’s disease that are caused by human papilloma virus infection and not by UV. Pigmented variants exist for Bowen’s disease and for actinic keratoses",
"links":["https://dermoscopedia.org/Actinic_keratosis_/_Bowen%27s_disease_/_keratoacanthoma_/_squamous_cell_carcinoma"]
}
# 6 more classes follow ...
}
# -
# ### create dataset_configs.py
# * this is the config file with the largest number of variables,
# * it contains information on
# * DROPOUT_VALUE : a keword/value that can be introduced to batch labels and will be recognised by FastClassAI function to not use images labelled like that for model training, eg to undersample one or more classes, or to exlude images from some classes in model training,
#
# * CLASS_COLORS
# * a dictiionary with colors assigned to original class labels,
# * key: original class label, value: color (any name accepted nby Matlotlib)
#
#
# * CLASS_COLORS_zorder
# * because some classes can be merged to build larger classes in different dataset variants,
# I created that variale to assign proper colors to a class that emerges from joingin these towo or more classes,
# * eg if we join class 1: yellow (zorder=1), and class 2: blue (zorder=100), new class will have blue color,
#
# * CLASS_LABELS_CONFIGS
#
# +
# configs .......................................................
DROPOUT_VALUE = "to_dropout"
# configs .......................................................
'''
colors assigned to original class labels,
'''
CLASS_COLORS ={
'bkl': 'orange',
'nv': 'forestgreen',
'df': 'purple',
'mel': 'black',
'Vasc': 'red',
'bcc': 'dimgrey',
'akiec': 'steelblue'}
# configs .......................................................
CLASS_COLORS_zorder ={
'bkl': 300,
'nv': 500,
'df': 1,
'mel': 200,
'Vasc': 1,
'bcc': 1,
'akiec': 1}
# configs .......................................................
DATASET_CONFIGS = {
"HAM10000": {
"info": "raw data grouped with original classes, no augmentation, duplicates were removed",
"labels": ["Cancer_Detection_And_Classification",
"Cancer_Risk_Groups",
"Melanoma_Detection",
"Skin_Cancer_Detection",
"Cancer_Classification"
]
# configs .......................................................
# CLASS_LABELS_CONFIGS
# key : str, name of the classyficaiton system used
# "info" : str, notes for the user
# "class_labels_dict" : dict, key: original class label, value: labels used in that classyficaiton system
#. "melanoma_stat_labels_dict" : dict, custom dict, added to allow caulating accuracy statistucs, with one class containigni melanoma (POSITIVE),
# vs all other classes designated as NEGATIVE
CLASS_LABELS_CONFIGS = {
"Cancer_Detection_And_Classification":{
"info":"more informative class names for raw data",
"class_labels_dict":{
'akiec': 'Squamous_cell_carcinoma',
'bcc': 'Basal_cell_carcinoma',
'bkl': 'Benign_keratosis',
'df': 'Dermatofibroma',
'nv': 'Melanocytic_nevus',
'mel': 'Melanoma',
'Vasc': 'Vascular_skin_lesions'},
"melanoma_stat_labels_dict":{
'Squamous_cell_carcinoma': 'NEGATIVE',
'Basal_cell_carcinoma': 'NEGATIVE',
'Benign_keratosis': 'NEGATIVE',
'Dermatofibroma': 'NEGATIVE',
'Melanocytic_nevus': 'NEGATIVE',
'Vascular_skin_lesions':'NEGATIVE',
'Melanoma': 'POSITIVE'}
},
"Cancer_Risk_Groups":{
"info":"""
7 original classes were grouped into three oncological risk groups
with vasc&nv assigned into low lever skin lessions, all other cancer types into cancer benign,
and melanoma as separate category
""",
"class_labels_dict":{
'akiec': 'Medium-benign_cancer',
'bcc': 'Medium-benign_cancer',
'bkl': 'Medium-benign_cancer',
'df': 'Medium-benign_cancer',
'nv': 'Low-skin_lession',
'mel': 'High-melanoma',
'Vasc': 'Low-skin_lession'},
"melanoma_stat_labels_dict":{
'Low-skin_lession': 'NEGATIVE',
'Medium-benign_cancer': 'NEGATIVE',
'High-melanoma': 'POSITIVE'}
}
}
| notebooks/project_setup/01_Setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (system-wide)
# language: python
# metadata:
# cocalc:
# description: Python 3 programming language
# priority: 100
# url: https://www.python.org/
# name: python3
# resource_dir: /ext/jupyter/kernels/python3
# ---
# implements specialize containers data types - Tuple and Dictionaries
# +
from collections import Counter # use to count
mylist = [1,1,1,1,1,1,2,2,2,2,2,3,3,3,3]
mylist
# -
# count how many 1's 2's and 3's
Counter(mylist)
mylist = ['a','a','a',10,10,10]
Counter(mylist)
sentence = "How many times does each word show up in this sentence"
count_words = sentence.lower().split()
count_words
Counter(count_words)
letters = 'aaaaaabbbbbbccccccddddd'
c = Counter(letters)
from collections import defaultdict
# will assign a default value if there is an instance when a keyerror occurs
d = {'a':10}
d['a']
d['key']
d = defaultdict(lambda: 0) # default value
d['correct'] = 100
d['Wrong Key']
d['a']
mytuple = (10,20,30)
mytuple[0]
# +
# may have a very large tuple
from collections import namedtuple
Dog = namedtuple('Dog',['age','breed','name']) #2 params name reported as , field names as a list
# -
sammy = Dog(age=5,breed='Husky',name='Sam')
sammy
sammy.age
type(sammy)
| PythonCollections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Retrieve
#
# You have to download manually the EEG data from https://physionet.org/content/eegmmidb
# # Parse
#
# Go through all subjects from the dataset, read the EDF files and store them into NumPy arrays.
#
# **Notes**
#
# * You have to download the dataset yourself, and modify the `edf_dir` variable.
#
# * In some subjects, we drop the last 170 samples, to make sure equal number of samples across subjects.
import numpy as np
import pyedflib
import os
eyes_open = np.zeros((109, 64, 9600))
eyes_closed = np.zeros((109, 64, 9600))
# Define the directory where dataset is located
edf_dir = '/opt/Temp/physionet.nlm.nih.gov/pn4/eegmmidb/'
# ### Process the baseline file for "eyes open"
for sub_id in range(0, 109):
subj_prefix = "S{0:03}".format(sub_id + 1)
subj_dir = "{0}/{1}".format(edf_dir, subj_prefix)
baseline_eyes_open = "{0}/{1}R01".format(subj_dir, subj_prefix)
f = pyedflib.EdfReader(baseline_eyes_open + ".edf")
a = f.read_annotation()
n = f.signals_in_file
signal_labels = f.getSignalLabels()
for chan in np.arange(n):
eyes_open[sub_id, chan, :] = f.readSignal(chan)[0:9600]
# ### Process the baseline file for "eyes closed"
for sub_id in range(0, 109):
subj_prefix = "S{0:03}".format(sub_id + 1)
subj_dir = "{0}/{1}".format(edf_dir, subj_prefix)
baseline_eyes_closed = "{0}/{1}R02".format(subj_dir, subj_prefix)
f = pyedflib.EdfReader(baseline_eyes_closed + ".edf")
a = f.read_annotation() #baseline_eyes_open + ".edf.event")
n = f.signals_in_file
signal_labels = f.getSignalLabels()
for chan in np.arange(n):
eyes_closed[sub_id, chan, :] = f.readSignal(chan)[0:9600]
# ### Store files
# +
if not os.path.exists("data/"):
os.makedirs("data/")
np.save('data/eyes_opened.npy', eyes_open)
np.save('data/eyes_closed.npy', eyes_closed)
| tutorials/0 - Retrieve and parse EEG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="24gYiJcWNlpA"
# ##### Copyright 2020 Google LLC
# + cellView="form" colab={} colab_type="code" id="ioaprt5q5US7"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="ItXfxkxvosLH"
# # Graph-based Neural Structured Learning in TFX
#
# This tutorial describes graph regularization from the
# [Neural Structured Learning](https://www.tensorflow.org/neural_structured_learning/)
# framework and demonstrates an end-to-end workflow for sentiment classification
# in a TFX pipeline.
# + [markdown] colab_type="text" id="vyAF26z9IDoq"
# Note: We recommend running this tutorial in a Colab notebook, with no setup required! Just click "Run in Google Colab".
#
# <div class="devsite-table-wrapper"><table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/tfx/tutorials/tfx/neural_structured_learning">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/tfx/blob/master/docs/tutorials/tfx/neural_structured_learning.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png">Run in Google Colab</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/tfx/tree/master/docs/tutorials/tfx/neural_structured_learning.ipynb">
# <img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">View source on GitHub</a></td>
# </table></div>
# + [markdown] colab_type="text" id="z3otbdCMmJiJ"
# ## Overview
# + [markdown] colab_type="text" id="ApxPtg2DiTtd"
# This notebook classifies movie reviews as *positive* or *negative* using the
# text of the review. This is an example of *binary* classification, an important
# and widely applicable kind of machine learning problem.
#
# We will demonstrate the use of graph regularization in this notebook by building
# a graph from the given input. The general recipe for building a
# graph-regularized model using the Neural Structured Learning (NSL) framework
# when the input does not contain an explicit graph is as follows:
#
# 1. Create embeddings for each text sample in the input. This can be done using
# pre-trained models such as [word2vec](https://arxiv.org/pdf/1310.4546.pdf),
# [Swivel](https://arxiv.org/abs/1602.02215),
# [BERT](https://arxiv.org/abs/1810.04805) etc.
# 2. Build a graph based on these embeddings by using a similarity metric such as
# the 'L2' distance, 'cosine' distance, etc. Nodes in the graph correspond to
# samples and edges in the graph correspond to similarity between pairs of
# samples.
# 3. Generate training data from the above synthesized graph and sample features.
# The resulting training data will contain neighbor features in addition to
# the original node features.
# 4. Create a neural network as a base model using Estimators.
# 5. Wrap the base model with the `add_graph_regularization` wrapper function,
# which is provided by the NSL framework, to create a new graph Estimator
# model. This new model will include a graph regularization loss as the
# regularization term in its training objective.
# 6. Train and evaluate the graph Estimator model.
#
# In this tutorial, we integrate the above workflow in a TFX pipeline using
# several custom TFX components as well as a custom graph-regularized trainer
# component.
#
# Below is the schematic for our TFX pipeline. Orange boxes represent
# off-the-shelf TFX components and pink boxes represent custom TFX components.
#
# 
# + [markdown] colab_type="text" id="EIx0r9-TeVQQ"
# ## Upgrade Pip
#
# To avoid upgrading Pip in a system when running locally, check to make sure that we're running in Colab. Local systems can of course be upgraded separately.
# + colab={} colab_type="code" id="-UmVrHUfkUA2"
try:
import colab
# !pip install --upgrade pip
except:
pass
# + [markdown] colab_type="text" id="nDOFbB34KY1R"
# ## Install Required Packages
# + colab={} colab_type="code" id="yDUe7gk_ztZ-"
# !pip install -q -U \
# tensorflow \
# tfx \
# neural-structured-learning \
# tensorflow-hub \
# tensorflow-datasets
# + [markdown] colab_type="text" id="1CeGS8G_eueJ"
# ## Did you restart the runtime?
#
# If you are using Google Colab, the first time that you run the cell above, you must restart the runtime (Runtime > Restart runtime ...). This is because of the way that Colab loads packages.
# + [markdown] colab_type="text" id="x6FJ64qMNLez"
# ## Dependencies and imports
# + colab={} colab_type="code" id="2ew7HTbPpCJH"
import gzip as gzip_lib
import numpy as np
import os
import pprint
import shutil
import tempfile
import urllib
pp = pprint.PrettyPrinter()
import tensorflow as tf
import tfx
from tfx.components.base import base_component
from tfx.components.base import base_executor
from tfx.components.base import executor_spec
from tfx.components.evaluator.component import Evaluator
from tfx.components.example_gen.import_example_gen.component import ImportExampleGen
from tfx.components.example_validator.component import ExampleValidator
from tfx.components.model_validator.component import ModelValidator
from tfx.components.pusher.component import Pusher
from tfx.components.schema_gen.component import SchemaGen
from tfx.components.statistics_gen.component import StatisticsGen
from tfx.components.trainer.component import Trainer
from tfx.components.transform.component import Transform
from tfx.orchestration.experimental.interactive.interactive_context import InteractiveContext
from tfx.proto import evaluator_pb2
from tfx.proto import example_gen_pb2
from tfx.proto import pusher_pb2
from tfx.proto import trainer_pb2
from tfx.utils.dsl_utils import external_input
from tfx.types import artifact
from tfx.types import artifact_utils
from tfx.types import standard_artifacts
from tfx.types.component_spec import ChannelParameter
from tfx.types.component_spec import ExecutionParameter
from tfx.dsl.component.experimental.annotations import InputArtifact
from tfx.dsl.component.experimental.annotations import OutputArtifact
from tfx.dsl.component.experimental.annotations import Parameter
from tfx.dsl.component.experimental.decorators import component
from tfx.types import artifact_utils
from tfx.types.standard_artifacts import Examples
from tensorflow_metadata.proto.v0 import anomalies_pb2
from tensorflow_metadata.proto.v0 import schema_pb2
from tensorflow_metadata.proto.v0 import statistics_pb2
import tensorflow_transform as tft
import tensorflow_model_analysis as tfma
import tensorflow_data_validation as tfdv
import neural_structured_learning as nsl
import tensorflow_hub as hub
import tensorflow_datasets as tfds
print("TF Version: ", tf.__version__)
print("Eager mode: ", tf.executing_eagerly())
print(
"GPU is",
"available" if tf.config.list_physical_devices("GPU") else "NOT AVAILABLE")
print("NSL Version: ", nsl.__version__)
print("TFX Version: ", tfx.__version__)
print("TFDV version: ", tfdv.__version__)
print("TFT version: ", tft.__version__)
print("TFMA version: ", tfma.__version__)
print("Hub version: ", hub.__version__)
# + [markdown] colab_type="text" id="nGwwFd99n42P"
# ## IMDB dataset
#
# The
# [IMDB dataset](https://www.tensorflow.org/datasets/catalog/imdb_reviews)
# contains the text of 50,000 movie reviews from the
# [Internet Movie Database](https://www.imdb.com/). These are split into 25,000
# reviews for training and 25,000 reviews for testing. The training and testing
# sets are *balanced*, meaning they contain an equal number of positive and
# negative reviews.
# Moreover, there are 50,000 additional unlabeled movie reviews.
# + [markdown] colab_type="text" id="iAsKG535pHep"
# ### Download preprocessed IMDB dataset
#
# The following code downloads the IMDB dataset (or uses a cached copy if it has already been downloaded) using TFDS. To speed up this notebook we will use only 10,000 labeled reviews and 10,000 unlabeled reviews for training, and 10,000 test reviews for evaluation.
# + colab={} colab_type="code" id="__cZi2Ic48KL"
train_set, eval_set = tfds.load(
"imdb_reviews:1.0.0",
split=["train[:10000]+unsupervised[:10000]", "test[:10000]"],
shuffle_files=False)
# + [markdown] colab_type="text" id="nE9tNh-67Y3W"
# Let's look at a few reviews from the training set:
# + colab={} colab_type="code" id="LsnHde8T67Jz"
for tfrecord in train_set.take(4):
print("Review: {}".format(tfrecord["text"].numpy().decode("utf-8")[:300]))
print("Label: {}\n".format(tfrecord["label"].numpy()))
# + colab={} colab_type="code" id="0wG7v3rk-Cwo"
def _dict_to_example(instance):
"""Decoded CSV to tf example."""
feature = {}
for key, value in instance.items():
if value is None:
feature[key] = tf.train.Feature()
elif value.dtype == np.integer:
feature[key] = tf.train.Feature(
int64_list=tf.train.Int64List(value=value.tolist()))
elif value.dtype == np.float32:
feature[key] = tf.train.Feature(
float_list=tf.train.FloatList(value=value.tolist()))
else:
feature[key] = tf.train.Feature(
bytes_list=tf.train.BytesList(value=value.tolist()))
return tf.train.Example(features=tf.train.Features(feature=feature))
examples_path = tempfile.mkdtemp(prefix="tfx-data")
train_path = os.path.join(examples_path, "train.tfrecord")
eval_path = os.path.join(examples_path, "eval.tfrecord")
for path, dataset in [(train_path, train_set), (eval_path, eval_set)]:
with tf.io.TFRecordWriter(path) as writer:
for example in dataset:
writer.write(
_dict_to_example({
"label": np.array([example["label"].numpy()]),
"text": np.array([example["text"].numpy()]),
}).SerializeToString())
# + [markdown] colab_type="text" id="HdQWxfsVkzdJ"
# ## Run TFX Components Interactively
#
# In the cells that follow you will construct TFX components and run each one interactively within the InteractiveContext to obtain `ExecutionResult` objects. This mirrors the process of an orchestrator running components in a TFX DAG based on when the dependencies for each component are met.
# + colab={} colab_type="code" id="4aVuXUil7hil"
context = InteractiveContext()
# + [markdown] colab_type="text" id="L9fwt9gQk3BR"
# ### The ExampleGen Component
# In any ML development process the first step when starting code development is to ingest the training and test datasets. The `ExampleGen` component brings data into the TFX pipeline.
#
# Create an ExampleGen component and run it.
# + colab={} colab_type="code" id="WdH4ql3Y7pT4"
input_data = external_input(examples_path)
input_config = example_gen_pb2.Input(splits=[
example_gen_pb2.Input.Split(name='train', pattern='train.tfrecord'),
example_gen_pb2.Input.Split(name='eval', pattern='eval.tfrecord')
])
example_gen = ImportExampleGen(input=input_data, input_config=input_config)
context.run(example_gen, enable_cache=True)
# + colab={} colab_type="code" id="IeUp6xCCrxsS"
for artifact in example_gen.outputs['examples'].get():
print(artifact)
print('\nexample_gen.outputs is a {}'.format(type(example_gen.outputs)))
print(example_gen.outputs)
print(example_gen.outputs['examples'].get()[0].split_names)
# + [markdown] colab_type="text" id="0SXc2OGnDWz5"
# The component's outputs include 2 artifacts:
# * the training examples (10,000 labeled reviews + 10,000 unlabeled reviews)
# * the eval examples (10,000 labeled reviews)
#
# + [markdown] colab_type="text" id="pcPppPASQzFa"
# ### The IdentifyExamples Custom Component
# To use NSL, we will need each instance to have a unique ID. We create a custom component that adds such a unique ID to all instances across all splits.
# + colab={} colab_type="code" id="XHCUzXA5qeWe"
@component
def IdentifyExamples(orig_examples: InputArtifact[Examples],
identified_examples: OutputArtifact[Examples],
id_feature_name: Parameter[str],
component_name: Parameter[str]) -> None:
# Get a list of the splits in input_data
splits_list = artifact_utils.decode_split_names(
split_names=orig_examples.split_names)
next_id = 0
for split in splits_list:
input_dir = os.path.join(orig_examples.uri, split)
output_dir = os.path.join(identified_examples.uri, split)
os.mkdir(output_dir)
for tfrecord_filename in os.listdir(input_dir):
input_path = os.path.join(input_dir, tfrecord_filename)
output_path = os.path.join(output_dir, tfrecord_filename)
with tf.io.TFRecordWriter(output_path, options="GZIP") as writer:
# Read each tfrecord file in the input split
for tfrecord in tf.data.TFRecordDataset(
input_path, compression_type="GZIP"):
example = tf.train.Example()
example.ParseFromString(tfrecord.numpy())
example.features.feature.get_or_create(
id_feature_name).bytes_list.MergeFrom(
tf.train.BytesList(value=[str(next_id).encode("utf-8")]))
next_id += 1
writer.write(example.SerializeToString())
# For completeness, encode the splits names.
# We could also just use input_data.split_names.
identified_examples.split_names = artifact_utils.encode_split_names(
splits=splits_list)
return
# + colab={} colab_type="code" id="ZtLxNWHPO0je"
identify_examples = IdentifyExamples(
orig_examples=example_gen.outputs['examples'],
component_name=u'IdentifyExamples',
id_feature_name=u'id')
context.run(identify_examples, enable_cache=False)
# + [markdown] colab_type="text" id="csM6BFhtk5Aa"
# ### The StatisticsGen Component
#
# The `StatisticsGen` component computes descriptive statistics for your dataset. The statistics that it generates can be visualized for review, and are used for example validation and to infer a schema.
#
# Create a StatisticsGen component and run it.
# + colab={} colab_type="code" id="MAscCCYWgA-9"
# Computes statistics over data for visualization and example validation.
statistics_gen = StatisticsGen(
examples=identify_examples.outputs["identified_examples"])
context.run(statistics_gen, enable_cache=True)
# + [markdown] colab_type="text" id="HLKLTO9Nk60p"
# ### The SchemaGen Component
#
# The `SchemaGen` component generates a schema for your data based on the statistics from StatisticsGen. It tries to infer the data types of each of your features, and the ranges of legal values for categorical features.
#
# Create a SchemaGen component and run it.
# + colab={} colab_type="code" id="ygQvZ6hsiQ_J"
# Generates schema based on statistics files.
schema_gen = SchemaGen(statistics=statistics_gen.outputs['statistics'])
context.run(schema_gen, enable_cache=True)
# + [markdown] colab_type="text" id="kdtU3u01FR-2"
# The generated artifact is just a `schema.pbtxt` containing a text representation of a `schema_pb2.Schema` protobuf:
# + colab={} colab_type="code" id="L6-tgKi6A_gK"
train_uri = schema_gen.outputs['schema'].get()[0].uri
schema_filename = os.path.join(train_uri, 'schema.pbtxt')
schema = tfx.utils.io_utils.parse_pbtxt_file(
file_name=schema_filename, message=schema_pb2.Schema())
# + [markdown] colab_type="text" id="FaSgx5qIFelw"
# It can be visualized using `tfdv.display_schema()` (we will look at this in more detail in a subsequent lab):
# + colab={} colab_type="code" id="gycOsJIQFhi3"
tfdv.display_schema(schema)
# + [markdown] colab_type="text" id="V1qcUuO9k9f8"
# ### The ExampleValidator Component
#
# The `ExampleValidator` performs anomaly detection, based on the statistics from StatisticsGen and the schema from SchemaGen. It looks for problems such as missing values, values of the wrong type, or categorical values outside of the domain of acceptable values.
#
# Create an ExampleValidator component and run it.
# + colab={} colab_type="code" id="XRlRUuGgiXks"
# Performs anomaly detection based on statistics and data schema.
validate_stats = ExampleValidator(
statistics=statistics_gen.outputs['statistics'],
schema=schema_gen.outputs['schema'])
context.run(validate_stats, enable_cache=False)
# + [markdown] colab_type="text" id="g3f2vmrF_e9b"
# ### The SynthesizeGraph Component
# + [markdown] colab_type="text" id="3oCuXo4BPfGr"
# Graph construction involves creating embeddings for text samples and then using
# a similarity function to compare the embeddings.
# + [markdown] colab_type="text" id="Gf8B3KxcinZ0"
# We will use pretrained Swivel embeddings to create embeddings in the
# `tf.train.Example` format for each sample in the input. We will store the
# resulting embeddings in the `TFRecord` format along with the sample's ID.
# This is important and will allow us match sample embeddings with corresponding
# nodes in the graph later.
# + [markdown] colab_type="text" id="_hSzZNdbPa4X"
# Once we have the sample embeddings, we will use them to build a similarity
# graph, i.e, nodes in this graph will correspond to samples and edges in this
# graph will correspond to similarity between pairs of nodes.
#
# Neural Structured Learning provides a graph building library to build a graph
# based on sample embeddings. It uses **cosine similarity** as the similarity
# measure to compare embeddings and build edges between them. It also allows us to specify a similarity threshold, which can be used to discard dissimilar edges from the final graph. In the following example, using 0.99 as the similarity threshold, we end up with a graph that has 115,368 bi-directional edges.
# + [markdown] colab_type="text" id="nERXNfSWPa4Z"
# **Note:** Graph quality and by extension, embedding quality, are very important
# for graph regularization. While we use Swivel embeddings in this notebook, using BERT embeddings for instance, will likely capture review semantics more
# accurately. We encourage users to use embeddings of their choice and as appropriate to their needs.
# + colab={} colab_type="code" id="2bAttbhgPa4V"
swivel_url = 'https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim/1'
hub_layer = hub.KerasLayer(swivel_url, input_shape=[], dtype=tf.string)
def _bytes_feature(value):
"""Returns a bytes_list from a string / byte."""
return tf.train.Feature(bytes_list=tf.train.BytesList(value=value))
def _float_feature(value):
"""Returns a float_list from a float / double."""
return tf.train.Feature(float_list=tf.train.FloatList(value=value))
def create_embedding_example(example):
"""Create tf.Example containing the sample's embedding and its ID."""
sentence_embedding = hub_layer(tf.sparse.to_dense(example['text']))
# Flatten the sentence embedding back to 1-D.
sentence_embedding = tf.reshape(sentence_embedding, shape=[-1])
feature_dict = {
'id': _bytes_feature(tf.sparse.to_dense(example['id']).numpy()),
'embedding': _float_feature(sentence_embedding.numpy().tolist())
}
return tf.train.Example(features=tf.train.Features(feature=feature_dict))
def create_dataset(uri):
tfrecord_filenames = [os.path.join(uri, name) for name in os.listdir(uri)]
return tf.data.TFRecordDataset(tfrecord_filenames, compression_type='GZIP')
def create_embeddings(train_path, output_path):
dataset = create_dataset(train_path)
embeddings_path = os.path.join(output_path, 'embeddings.tfr')
feature_map = {
'label': tf.io.FixedLenFeature([], tf.int64),
'id': tf.io.VarLenFeature(tf.string),
'text': tf.io.VarLenFeature(tf.string)
}
with tf.io.TFRecordWriter(embeddings_path) as writer:
for tfrecord in dataset:
tensor_dict = tf.io.parse_single_example(tfrecord, feature_map)
embedding_example = create_embedding_example(tensor_dict)
writer.write(embedding_example.SerializeToString())
def build_graph(output_path, similarity_threshold):
embeddings_path = os.path.join(output_path, 'embeddings.tfr')
graph_path = os.path.join(output_path, 'graph.tfv')
nsl.tools.build_graph([embeddings_path], graph_path, similarity_threshold)
# + colab={} colab_type="code" id="ITkf2SLg1TG7"
"""Custom Artifact type"""
class SynthesizedGraph(tfx.types.artifact.Artifact):
"""Output artifact of the SynthesizeGraph component"""
TYPE_NAME = 'SynthesizedGraphPath'
PROPERTIES = {
'span': standard_artifacts.SPAN_PROPERTY,
'split_names': standard_artifacts.SPLIT_NAMES_PROPERTY,
}
@component
def SynthesizeGraph(identified_examples: InputArtifact[Examples],
synthesized_graph: OutputArtifact[SynthesizedGraph],
similarity_threshold: Parameter[float],
component_name: Parameter[str]) -> None:
# Get a list of the splits in input_data
splits_list = artifact_utils.decode_split_names(
split_names=identified_examples.split_names)
# We build a graph only based on the 'train' split which includes both
# labeled and unlabeled examples.
train_input_examples_uri = os.path.join(identified_examples.uri, 'train')
output_graph_uri = os.path.join(synthesized_graph.uri, 'train')
os.mkdir(output_graph_uri)
print('Creating embeddings...')
create_embeddings(train_input_examples_uri, output_graph_uri)
print('Synthesizing graph...')
build_graph(output_graph_uri, similarity_threshold)
synthesized_graph.split_names = artifact_utils.encode_split_names(
splits=['train'])
return
# + colab={} colab_type="code" id="H0ZkHvJMA-0G"
synthesize_graph = SynthesizeGraph(
identified_examples=identify_examples.outputs['identified_examples'],
component_name=u'SynthesizeGraph',
similarity_threshold=0.99)
context.run(synthesize_graph, enable_cache=False)
# + colab={} colab_type="code" id="o54M-0Q11FcS"
train_uri = synthesize_graph.outputs["synthesized_graph"].get()[0].uri
os.listdir(train_uri)
# + colab={} colab_type="code" id="IRK_rS_q1UcZ"
graph_path = os.path.join(train_uri, "train", "graph.tfv")
print("node 1\tnode 2\tsimilarity")
# !head {graph_path}
print("...")
# !tail {graph_path}
# + colab={} colab_type="code" id="uybqyWztvCGm"
# !wc -l {graph_path}
# + [markdown] colab_type="text" id="JPViEz5RlA36"
# ### The Transform Component
#
# The `Transform` component performs data transformations and feature engineering. The results include an input TensorFlow graph which is used during both training and serving to preprocess the data before training or inference. This graph becomes part of the SavedModel that is the result of model training. Since the same input graph is used for both training and serving, the preprocessing will always be the same, and only needs to be written once.
#
# The Transform component requires more code than many other components because of the arbitrary complexity of the feature engineering that you may need for the data and/or model that you're working with. It requires code files to be available which define the processing needed.
# + [markdown] colab_type="text" id="_USkfut69gNW"
# Each sample will include the following three features:
#
# 1. **id**: The node ID of the sample.
# 2. **text_xf**: An int64 list containing word IDs.
# 3. **label_xf**: A singleton int64 identifying the target class of the review: 0=negative, 1=positive.
# + [markdown] colab_type="text" id="XUYeCayFG7kH"
# Let's define a module containing the `preprocessing_fn()` function that we will pass to the `Transform` component:
# + colab={} colab_type="code" id="7uuWiQbOG9ki"
_transform_module_file = 'imdb_transform.py'
# + colab={} colab_type="code" id="v3EIuVQnBfH7"
# %%writefile {_transform_module_file}
import tensorflow as tf
import tensorflow_transform as tft
SEQUENCE_LENGTH = 100
VOCAB_SIZE = 10000
OOV_SIZE = 100
def tokenize_reviews(reviews, sequence_length=SEQUENCE_LENGTH):
reviews = tf.strings.lower(reviews)
reviews = tf.strings.regex_replace(reviews, r" '| '|^'|'$", " ")
reviews = tf.strings.regex_replace(reviews, "[^a-z' ]", " ")
tokens = tf.strings.split(reviews)[:, :sequence_length]
start_tokens = tf.fill([tf.shape(reviews)[0], 1], "<START>")
end_tokens = tf.fill([tf.shape(reviews)[0], 1], "<END>")
tokens = tf.concat([start_tokens, tokens, end_tokens], axis=1)
tokens = tokens[:, :sequence_length]
tokens = tokens.to_tensor(default_value="<PAD>")
pad = sequence_length - tf.shape(tokens)[1]
tokens = tf.pad(tokens, [[0, 0], [0, pad]], constant_values="<PAD>")
return tf.reshape(tokens, [-1, sequence_length])
def preprocessing_fn(inputs):
"""tf.transform's callback function for preprocessing inputs.
Args:
inputs: map from feature keys to raw not-yet-transformed features.
Returns:
Map from string feature key to transformed feature operations.
"""
outputs = {}
outputs["id"] = inputs["id"]
tokens = tokenize_reviews(_fill_in_missing(inputs["text"], ''))
outputs["text_xf"] = tft.compute_and_apply_vocabulary(
tokens,
top_k=VOCAB_SIZE,
num_oov_buckets=OOV_SIZE)
outputs["label_xf"] = _fill_in_missing(inputs["label"], -1)
return outputs
def _fill_in_missing(x, default_value):
"""Replace missing values in a SparseTensor.
Fills in missing values of `x` with the default_value.
Args:
x: A `SparseTensor` of rank 2. Its dense shape should have size at most 1
in the second dimension.
default_value: the value with which to replace the missing values.
Returns:
A rank 1 tensor where missing values of `x` have been filled in.
"""
return tf.squeeze(
tf.sparse.to_dense(
tf.SparseTensor(x.indices, x.values, [x.dense_shape[0], 1]),
default_value),
axis=1)
# + [markdown] colab_type="text" id="eeMVMafpHHX1"
# Create and run the `Transform` component, referring to the files that were created above.
# + colab={} colab_type="code" id="jHfhth_GiZI9"
# Performs transformations and feature engineering in training and serving.
transform = Transform(
examples=identify_examples.outputs['identified_examples'],
schema=schema_gen.outputs['schema'],
module_file=_transform_module_file)
context.run(transform)
# + [markdown] colab_type="text" id="_jbZO1ykHOeG"
# The `Transform` component has 2 types of outputs:
# * `transform_graph` is the graph that can perform the preprocessing operations (this graph will be included in the serving and evaluation models).
# * `transformed_examples` represents the preprocessed training and evaluation data.
# + colab={} colab_type="code" id="j4UjersvAC7p"
transform.outputs
# + [markdown] colab_type="text" id="wRFMlRcdHlQy"
# Take a peek at the `transform_graph` artifact: it points to a directory containing 3 subdirectories:
# + colab={} colab_type="code" id="E4I-cqfQQvaW"
train_uri = transform.outputs['transform_graph'].get()[0].uri
os.listdir(train_uri)
# + [markdown] colab_type="text" id="9374B4RpHzor"
# The `transform_fn` subdirectory contains the actual preprocessing graph. The `metadata` subdirectory contains the schema of the original data. The `transformed_metadata` subdirectory contains the schema of the preprocessed data.
#
# Take a look at some of the transformed examples and check that they are indeed processed as intended.
# + colab={} colab_type="code" id="-QPONyzDTswf"
def pprint_examples(artifact, n_examples=3):
print("artifact:", artifact)
uri = os.path.join(artifact.uri, "train")
print("uri:", uri)
tfrecord_filenames = [os.path.join(uri, name) for name in os.listdir(uri)]
print("tfrecord_filenames:", tfrecord_filenames)
dataset = tf.data.TFRecordDataset(tfrecord_filenames, compression_type="GZIP")
for tfrecord in dataset.take(n_examples):
serialized_example = tfrecord.numpy()
example = tf.train.Example.FromString(serialized_example)
pp.pprint(example)
# + colab={} colab_type="code" id="2zIepQhSQoPa"
pprint_examples(transform.outputs['transformed_examples'].get()[0])
# + [markdown] colab_type="text" id="vpGvPKielIvI"
# ### The GraphAugmentation Component
#
# Since we have the sample features and the synthesized graph, we can generate the
# augmented training data for Neural Structured Learning. The NSL framework
# provides a library to combine the graph and the sample features to produce
# the final training data for graph regularization. The resulting training data
# will include original sample features as well as features of their corresponding
# neighbors.
#
# In this tutorial, we consider undirected edges and use a maximum of 3 neighbors
# per sample to augment training data with graph neighbors.
# + colab={} colab_type="code" id="gI6P_-AXGm04"
def split_train_and_unsup(input_uri):
'Separate the labeled and unlabeled instances.'
tmp_dir = tempfile.mkdtemp(prefix='tfx-data')
tfrecord_filenames = [
os.path.join(input_uri, filename) for filename in os.listdir(input_uri)
]
train_path = os.path.join(tmp_dir, 'train.tfrecord')
unsup_path = os.path.join(tmp_dir, 'unsup.tfrecord')
with tf.io.TFRecordWriter(train_path) as train_writer, \
tf.io.TFRecordWriter(unsup_path) as unsup_writer:
for tfrecord in tf.data.TFRecordDataset(
tfrecord_filenames, compression_type='GZIP'):
example = tf.train.Example()
example.ParseFromString(tfrecord.numpy())
if ('label_xf' not in example.features.feature or
example.features.feature['label_xf'].int64_list.value[0] == -1):
writer = unsup_writer
else:
writer = train_writer
writer.write(tfrecord.numpy())
return train_path, unsup_path
def gzip(filepath):
with open(filepath, 'rb') as f_in:
with gzip_lib.open(filepath + '.gz', 'wb') as f_out:
shutil.copyfileobj(f_in, f_out)
os.remove(filepath)
def copy_tfrecords(input_uri, output_uri):
for filename in os.listdir(input_uri):
input_filename = os.path.join(input_uri, filename)
output_filename = os.path.join(output_uri, filename)
shutil.copyfile(input_filename, output_filename)
@component
def GraphAugmentation(identified_examples: InputArtifact[Examples],
synthesized_graph: InputArtifact[SynthesizedGraph],
augmented_examples: OutputArtifact[Examples],
num_neighbors: Parameter[int],
component_name: Parameter[str]) -> None:
# Get a list of the splits in input_data
splits_list = artifact_utils.decode_split_names(
split_names=identified_examples.split_names)
train_input_uri = os.path.join(identified_examples.uri, 'train')
eval_input_uri = os.path.join(identified_examples.uri, 'eval')
train_graph_uri = os.path.join(synthesized_graph.uri, 'train')
train_output_uri = os.path.join(augmented_examples.uri, 'train')
eval_output_uri = os.path.join(augmented_examples.uri, 'eval')
os.mkdir(train_output_uri)
os.mkdir(eval_output_uri)
# Separate out the labeled and unlabeled examples from the 'train' split.
train_path, unsup_path = split_train_and_unsup(train_input_uri)
output_path = os.path.join(train_output_uri, 'nsl_train_data.tfr')
pack_nbrs_args = dict(
labeled_examples_path=train_path,
unlabeled_examples_path=unsup_path,
graph_path=os.path.join(train_graph_uri, 'graph.tfv'),
output_training_data_path=output_path,
add_undirected_edges=True,
max_nbrs=num_neighbors)
print('nsl.tools.pack_nbrs arguments:', pack_nbrs_args)
nsl.tools.pack_nbrs(**pack_nbrs_args)
# Downstream components expect gzip'ed TFRecords.
gzip(output_path)
# The test examples are left untouched and are simply copied over.
copy_tfrecords(eval_input_uri, eval_output_uri)
augmented_examples.split_names = identified_examples.split_names
return
# + colab={} colab_type="code" id="r9MIEVDiOANe"
# Augments training data with graph neighbors.
graph_augmentation = GraphAugmentation(
identified_examples=transform.outputs['transformed_examples'],
synthesized_graph=synthesize_graph.outputs['synthesized_graph'],
component_name=u'GraphAugmentation',
num_neighbors=3)
context.run(graph_augmentation, enable_cache=False)
# + colab={} colab_type="code" id="gpSLs3Hx8viI"
pprint_examples(graph_augmentation.outputs['augmented_examples'].get()[0], 6)
# + [markdown] colab_type="text" id="OBJFtnl6lCg9"
# ### The Trainer Component
#
# The `Trainer` component trains models using TensorFlow.
#
# Create a Python module containing a `trainer_fn` function, which must return an estimator. If you prefer creating a Keras model, you can do so and then convert it to an estimator using `keras.model_to_estimator()`.
# + colab={} colab_type="code" id="5ajvClE6b2pd"
# Setup paths.
_trainer_module_file = 'imdb_trainer.py'
# + colab={} colab_type="code" id="_dh6AejVk2Oq"
# %%writefile {_trainer_module_file}
import neural_structured_learning as nsl
import tensorflow as tf
import tensorflow_model_analysis as tfma
import tensorflow_transform as tft
from tensorflow_transform.tf_metadata import schema_utils
NBR_FEATURE_PREFIX = 'NL_nbr_'
NBR_WEIGHT_SUFFIX = '_weight'
LABEL_KEY = 'label'
ID_FEATURE_KEY = 'id'
def _transformed_name(key):
return key + '_xf'
def _transformed_names(keys):
return [_transformed_name(key) for key in keys]
# Hyperparameters:
#
# We will use an instance of `HParams` to inclue various hyperparameters and
# constants used for training and evaluation. We briefly describe each of them
# below:
#
# - max_seq_length: This is the maximum number of words considered from each
# movie review in this example.
# - vocab_size: This is the size of the vocabulary considered for this
# example.
# - oov_size: This is the out-of-vocabulary size considered for this example.
# - distance_type: This is the distance metric used to regularize the sample
# with its neighbors.
# - graph_regularization_multiplier: This controls the relative weight of the
# graph regularization term in the overall
# loss function.
# - num_neighbors: The number of neighbors used for graph regularization. This
# value has to be less than or equal to the `num_neighbors`
# argument used above in the GraphAugmentation component when
# invoking `nsl.tools.pack_nbrs`.
# - num_fc_units: The number of units in the fully connected layer of the
# neural network.
class HParams(object):
"""Hyperparameters used for training."""
def __init__(self):
### dataset parameters
# The following 3 values should match those defined in the Transform
# Component.
self.max_seq_length = 100
self.vocab_size = 10000
self.oov_size = 100
### Neural Graph Learning parameters
self.distance_type = nsl.configs.DistanceType.L2
self.graph_regularization_multiplier = 0.1
# The following value has to be at most the value of 'num_neighbors' used
# in the GraphAugmentation component.
self.num_neighbors = 1
### Model Architecture
self.num_embedding_dims = 16
self.num_fc_units = 64
HPARAMS = HParams()
def optimizer_fn():
"""Returns an instance of `tf.Optimizer`."""
return tf.compat.v1.train.RMSPropOptimizer(
learning_rate=0.0001, decay=1e-6)
def build_train_op(loss, global_step):
"""Builds a train op to optimize the given loss using gradient descent."""
with tf.name_scope('train'):
optimizer = optimizer_fn()
train_op = optimizer.minimize(loss=loss, global_step=global_step)
return train_op
# Building the model:
#
# A neural network is created by stacking layers—this requires two main
# architectural decisions:
# * How many layers to use in the model?
# * How many *hidden units* to use for each layer?
#
# In this example, the input data consists of an array of word-indices. The
# labels to predict are either 0 or 1. We will use a feed-forward neural network
# as our base model in this tutorial.
def feed_forward_model(features, is_training, reuse=tf.compat.v1.AUTO_REUSE):
"""Builds a simple 2 layer feed forward neural network.
The layers are effectively stacked sequentially to build the classifier. The
first layer is an Embedding layer, which takes the integer-encoded vocabulary
and looks up the embedding vector for each word-index. These vectors are
learned as the model trains. The vectors add a dimension to the output array.
The resulting dimensions are: (batch, sequence, embedding). Next is a global
average pooling 1D layer, which reduces the dimensionality of its inputs from
3D to 2D. This fixed-length output vector is piped through a fully-connected
(Dense) layer with 16 hidden units. The last layer is densely connected with a
single output node. Using the sigmoid activation function, this value is a
float between 0 and 1, representing a probability, or confidence level.
Args:
features: A dictionary containing batch features returned from the
`input_fn`, that include sample features, corresponding neighbor features,
and neighbor weights.
is_training: a Python Boolean value or a Boolean scalar Tensor, indicating
whether to apply dropout.
reuse: a Python Boolean value for reusing variable scope.
Returns:
logits: Tensor of shape [batch_size, 1].
representations: Tensor of shape [batch_size, _] for graph regularization.
This is the representation of each example at the graph regularization
layer.
"""
with tf.compat.v1.variable_scope('ff', reuse=reuse):
inputs = features[_transformed_name('text')]
embeddings = tf.compat.v1.get_variable(
'embeddings',
shape=[
HPARAMS.vocab_size + HPARAMS.oov_size, HPARAMS.num_embedding_dims
])
embedding_layer = tf.nn.embedding_lookup(embeddings, inputs)
pooling_layer = tf.compat.v1.layers.AveragePooling1D(
pool_size=HPARAMS.max_seq_length, strides=HPARAMS.max_seq_length)(
embedding_layer)
# Shape of pooling_layer is now [batch_size, 1, HPARAMS.num_embedding_dims]
pooling_layer = tf.reshape(pooling_layer, [-1, HPARAMS.num_embedding_dims])
dense_layer = tf.compat.v1.layers.Dense(
16, activation='relu')(
pooling_layer)
output_layer = tf.compat.v1.layers.Dense(
1, activation='sigmoid')(
dense_layer)
# Graph regularization will be done on the penultimate (dense) layer
# because the output layer is a single floating point number.
return output_layer, dense_layer
# A note on hidden units:
#
# The above model has two intermediate or "hidden" layers, between the input and
# output, and excluding the Embedding layer. The number of outputs (units,
# nodes, or neurons) is the dimension of the representational space for the
# layer. In other words, the amount of freedom the network is allowed when
# learning an internal representation. If a model has more hidden units
# (a higher-dimensional representation space), and/or more layers, then the
# network can learn more complex representations. However, it makes the network
# more computationally expensive and may lead to learning unwanted
# patterns—patterns that improve performance on training data but not on the
# test data. This is called overfitting.
# This function will be used to generate the embeddings for samples and their
# corresponding neighbors, which will then be used for graph regularization.
def embedding_fn(features, mode):
"""Returns the embedding corresponding to the given features.
Args:
features: A dictionary containing batch features returned from the
`input_fn`, that include sample features, corresponding neighbor features,
and neighbor weights.
mode: Specifies if this is training, evaluation, or prediction. See
tf.estimator.ModeKeys.
Returns:
The embedding that will be used for graph regularization.
"""
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
_, embedding = feed_forward_model(features, is_training)
return embedding
def feed_forward_model_fn(features, labels, mode, params, config):
"""Implementation of the model_fn for the base feed-forward model.
Args:
features: This is the first item returned from the `input_fn` passed to
`train`, `evaluate`, and `predict`. This should be a single `Tensor` or
`dict` of same.
labels: This is the second item returned from the `input_fn` passed to
`train`, `evaluate`, and `predict`. This should be a single `Tensor` or
`dict` of same (for multi-head models). If mode is `ModeKeys.PREDICT`,
`labels=None` will be passed. If the `model_fn`'s signature does not
accept `mode`, the `model_fn` must still be able to handle `labels=None`.
mode: Optional. Specifies if this training, evaluation or prediction. See
`ModeKeys`.
params: An HParams instance as returned by get_hyper_parameters().
config: Optional configuration object. Will receive what is passed to
Estimator in `config` parameter, or the default `config`. Allows updating
things in your model_fn based on configuration such as `num_ps_replicas`,
or `model_dir`. Unused currently.
Returns:
A `tf.estimator.EstimatorSpec` for the base feed-forward model. This does
not include graph-based regularization.
"""
is_training = mode == tf.estimator.ModeKeys.TRAIN
# Build the computation graph.
probabilities, _ = feed_forward_model(features, is_training)
predictions = tf.round(probabilities)
if mode == tf.estimator.ModeKeys.PREDICT:
# labels will be None, and no loss to compute.
cross_entropy_loss = None
eval_metric_ops = None
else:
# Loss is required in train and eval modes.
# Flatten 'probabilities' to 1-D.
probabilities = tf.reshape(probabilities, shape=[-1])
cross_entropy_loss = tf.compat.v1.keras.losses.binary_crossentropy(
labels, probabilities)
eval_metric_ops = {
'accuracy': tf.compat.v1.metrics.accuracy(labels, predictions)
}
if is_training:
global_step = tf.compat.v1.train.get_or_create_global_step()
train_op = build_train_op(cross_entropy_loss, global_step)
else:
train_op = None
return tf.estimator.EstimatorSpec(
mode=mode,
predictions={
'probabilities': probabilities,
'predictions': predictions
},
loss=cross_entropy_loss,
train_op=train_op,
eval_metric_ops=eval_metric_ops)
# Tf.Transform considers these features as "raw"
def _get_raw_feature_spec(schema):
return schema_utils.schema_as_feature_spec(schema).feature_spec
def _gzip_reader_fn(filenames):
"""Small utility returning a record reader that can read gzip'ed files."""
return tf.data.TFRecordDataset(
filenames,
compression_type='GZIP')
def _example_serving_receiver_fn(tf_transform_output, schema):
"""Build the serving in inputs.
Args:
tf_transform_output: A TFTransformOutput.
schema: the schema of the input data.
Returns:
Tensorflow graph which parses examples, applying tf-transform to them.
"""
raw_feature_spec = _get_raw_feature_spec(schema)
raw_feature_spec.pop(LABEL_KEY)
# We don't need the ID feature for serving.
raw_feature_spec.pop(ID_FEATURE_KEY)
raw_input_fn = tf.estimator.export.build_parsing_serving_input_receiver_fn(
raw_feature_spec, default_batch_size=None)
serving_input_receiver = raw_input_fn()
transformed_features = tf_transform_output.transform_raw_features(
serving_input_receiver.features)
# Even though, LABEL_KEY was removed from 'raw_feature_spec', the transform
# operation would have injected the transformed LABEL_KEY feature with a
# default value.
transformed_features.pop(_transformed_name(LABEL_KEY))
return tf.estimator.export.ServingInputReceiver(
transformed_features, serving_input_receiver.receiver_tensors)
def _eval_input_receiver_fn(tf_transform_output, schema):
"""Build everything needed for the tf-model-analysis to run the model.
Args:
tf_transform_output: A TFTransformOutput.
schema: the schema of the input data.
Returns:
EvalInputReceiver function, which contains:
- Tensorflow graph which parses raw untransformed features, applies the
tf-transform preprocessing operators.
- Set of raw, untransformed features.
- Label against which predictions will be compared.
"""
# Notice that the inputs are raw features, not transformed features here.
raw_feature_spec = _get_raw_feature_spec(schema)
# We don't need the ID feature for TFMA.
raw_feature_spec.pop(ID_FEATURE_KEY)
raw_input_fn = tf.estimator.export.build_parsing_serving_input_receiver_fn(
raw_feature_spec, default_batch_size=None)
serving_input_receiver = raw_input_fn()
transformed_features = tf_transform_output.transform_raw_features(
serving_input_receiver.features)
labels = transformed_features.pop(_transformed_name(LABEL_KEY))
return tfma.export.EvalInputReceiver(
features=transformed_features,
receiver_tensors=serving_input_receiver.receiver_tensors,
labels=labels)
def _augment_feature_spec(feature_spec, num_neighbors):
"""Augments `feature_spec` to include neighbor features.
Args:
feature_spec: Dictionary of feature keys mapping to TF feature types.
num_neighbors: Number of neighbors to use for feature key augmentation.
Returns:
An augmented `feature_spec` that includes neighbor feature keys.
"""
for i in range(num_neighbors):
feature_spec['{}{}_{}'.format(NBR_FEATURE_PREFIX, i, 'id')] = \
tf.io.VarLenFeature(dtype=tf.string)
# We don't care about the neighbor features corresponding to
# _transformed_name(LABEL_KEY) because the LABEL_KEY feature will be
# removed from the feature spec during training/evaluation.
feature_spec['{}{}_{}'.format(NBR_FEATURE_PREFIX, i, 'text_xf')] = \
tf.io.FixedLenFeature(shape=[HPARAMS.max_seq_length], dtype=tf.int64,
default_value=tf.constant(0, dtype=tf.int64,
shape=[HPARAMS.max_seq_length]))
# The 'NL_num_nbrs' features is currently not used.
# Set the neighbor weight feature keys.
for i in range(num_neighbors):
feature_spec['{}{}{}'.format(NBR_FEATURE_PREFIX, i, NBR_WEIGHT_SUFFIX)] = \
tf.io.FixedLenFeature(shape=[1], dtype=tf.float32, default_value=[0.0])
return feature_spec
def _input_fn(filenames, tf_transform_output, is_training, batch_size=200):
"""Generates features and labels for training or evaluation.
Args:
filenames: [str] list of CSV files to read data from.
tf_transform_output: A TFTransformOutput.
is_training: Boolean indicating if we are in training mode.
batch_size: int First dimension size of the Tensors returned by input_fn
Returns:
A (features, indices) tuple where features is a dictionary of
Tensors, and indices is a single Tensor of label indices.
"""
transformed_feature_spec = (
tf_transform_output.transformed_feature_spec().copy())
# During training, NSL uses augmented training data (which includes features
# from graph neighbors). So, update the feature spec accordingly. This needs
# to be done because we are using different schemas for NSL training and eval,
# but the Trainer Component only accepts a single schema.
if is_training:
transformed_feature_spec =_augment_feature_spec(transformed_feature_spec,
HPARAMS.num_neighbors)
dataset = tf.data.experimental.make_batched_features_dataset(
filenames, batch_size, transformed_feature_spec, reader=_gzip_reader_fn)
transformed_features = tf.compat.v1.data.make_one_shot_iterator(
dataset).get_next()
# We pop the label because we do not want to use it as a feature while we're
# training.
return transformed_features, transformed_features.pop(
_transformed_name(LABEL_KEY))
# TFX will call this function
def trainer_fn(hparams, schema):
"""Build the estimator using the high level API.
Args:
hparams: Holds hyperparameters used to train the model as name/value pairs.
schema: Holds the schema of the training examples.
Returns:
A dict of the following:
- estimator: The estimator that will be used for training and eval.
- train_spec: Spec for training.
- eval_spec: Spec for eval.
- eval_input_receiver_fn: Input function for eval.
"""
train_batch_size = 40
eval_batch_size = 40
tf_transform_output = tft.TFTransformOutput(hparams.transform_output)
train_input_fn = lambda: _input_fn(
hparams.train_files,
tf_transform_output,
is_training=True,
batch_size=train_batch_size)
eval_input_fn = lambda: _input_fn(
hparams.eval_files,
tf_transform_output,
is_training=False,
batch_size=eval_batch_size)
train_spec = tf.estimator.TrainSpec(
train_input_fn,
max_steps=hparams.train_steps)
serving_receiver_fn = lambda: _example_serving_receiver_fn(
tf_transform_output, schema)
exporter = tf.estimator.FinalExporter('imdb', serving_receiver_fn)
eval_spec = tf.estimator.EvalSpec(
eval_input_fn,
steps=hparams.eval_steps,
exporters=[exporter],
name='imdb-eval')
run_config = tf.estimator.RunConfig(
save_checkpoints_steps=999, keep_checkpoint_max=1)
run_config = run_config.replace(model_dir=hparams.serving_model_dir)
estimator = tf.estimator.Estimator(
model_fn=feed_forward_model_fn, config=run_config, params=HPARAMS)
# Create a graph regularization config.
graph_reg_config = nsl.configs.make_graph_reg_config(
max_neighbors=HPARAMS.num_neighbors,
multiplier=HPARAMS.graph_regularization_multiplier,
distance_type=HPARAMS.distance_type,
sum_over_axis=-1)
# Invoke the Graph Regularization Estimator wrapper to incorporate
# graph-based regularization for training.
graph_nsl_estimator = nsl.estimator.add_graph_regularization(
estimator,
embedding_fn,
optimizer_fn=optimizer_fn,
graph_reg_config=graph_reg_config)
# Create an input receiver for TFMA processing
receiver_fn = lambda: _eval_input_receiver_fn(
tf_transform_output, schema)
return {
'estimator': graph_nsl_estimator,
'train_spec': train_spec,
'eval_spec': eval_spec,
'eval_input_receiver_fn': receiver_fn
}
# + [markdown] colab_type="text" id="GnLjStUJIoos"
# Create and run the `Trainer` component, passing it the file that we created above.
# + colab={} colab_type="code" id="MWLQI6t0b2pg"
# Uses user-provided Python function that implements a model using TensorFlow's
# Estimators API.
trainer = Trainer(
module_file=_trainer_module_file,
transformed_examples=graph_augmentation.outputs['augmented_examples'],
schema=schema_gen.outputs['schema'],
transform_graph=transform.outputs['transform_graph'],
train_args=trainer_pb2.TrainArgs(num_steps=10000),
eval_args=trainer_pb2.EvalArgs(num_steps=5000))
context.run(trainer)
# + [markdown] colab_type="text" id="pDiZvYbFb2ph"
# Take a peek at the trained model which was exported from `Trainer`.
# + colab={} colab_type="code" id="qDBZG9Oso-BD"
train_uri = trainer.outputs['model'].get()[0].uri
serving_model_path = os.path.join(train_uri, 'serving_model_dir', 'export',
'imdb')
latest_serving_model_path = os.path.join(serving_model_path,
max(os.listdir(serving_model_path)))
exported_model = tf.saved_model.load(latest_serving_model_path)
# + colab={} colab_type="code" id="KyT3ZVGCZWsj"
exported_model.graph.get_operations()[:10] + ["..."]
# + [markdown] colab_type="text" id="zIsspBf5GjKm"
# Let's visualize the model's metrics using Tensorboard.
# + colab={} colab_type="code" id="rnKeqLmcGqHH"
#docs_infra: no_execute
# Get the URI of the output artifact representing the training logs,
# which is a directory
model_dir = train_uri
# %load_ext tensorboard
# %tensorboard --logdir {model_dir}
# + [markdown] colab_type="text" id="LgZXZJBsGzHm"
# ## Model Serving
#
# Graph regularization only affects the training workflow by adding a regularization term to the loss function. As a result, the model evaluation and serving workflows remain unchanged. It is for the same reason that we've also omitted downstream TFX components that typically come after the *Trainer* component like the *Evaluator*, *Pusher*, etc.
# + [markdown] colab_type="text" id="qOh5FjbWiP-b"
# ## Conclusion
#
# We have demonstrated the use of graph regularization using the Neural Structured
# Learning (NSL) framework in a TFX pipeline even when the input does not contain
# an explicit graph. We considered the task of sentiment classification of IMDB
# movie reviews for which we synthesized a similarity graph based on review
# embeddings. We encourage users to experiment further by using different
# embeddings for graph construction, varying hyperparameters, changing the amount
# of supervision, and by defining different model architectures.
| site/en-snapshot/tfx/tutorials/tfx/neural_structured_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to classify Lego figures
# ## What better way to impress your significant other?
# ### Build a Lego Classifier with fastai
# !pip install kaggle
# We want to save the dataset into the folder /notebooks/storage/data/Lego-Classification:
# !kaggle datasets download -d ihelon/lego-minifigures-classification -p /notebooks/storage/data/Lego-Classification
# ### Unzip data using Pythons pathlib library
from pathlib import Path
p = Path('/notebooks/storage/data/Lego-Classification')
filename = Path('/notebooks/storage/data/Lego-Classification/lego-minifigures-classification.zip')
# Just as before, we can use bash commands from within Jupyter Notebook. So let's do that to unzip our data. -q is quiet mode, -d points to the direction where to unzip the data. Just see how well Pythons pathlib and bash work together!
# !unzip -q {str(filename)} -d {str(p/"train")}
# ### Imports
from fastbook import *
from fastai.vision.widgets import *
import pandas as pd
# Let's now use fastai's "get_image_files()" function to see how the unzipped data looks like in our destination path:
fns = get_image_files(p/"train")
fns
# Remember, we put the data into our directory '/notebooks/storage/data/Lego-Classification'. After having a quick look at our data it looks like the data is stored as follows: first the genre of our image (marvel/jurassic-world), then the classification of the figure (0001/0002 etc.). Within these folders we find many different pictures of that figure (001.jpg/002.jpg and so on).
# Let's confirm this by looking at the metadata.
df = pd.read_csv(f'{p}/index.csv', index_col=0)
df.tail(5)
df_metadata = pd.read_csv(f'{p}/metadata.csv', usecols=['class_id', 'lego_names', 'minifigure_name'])
df_metadata.head()
# Indeed, that's how this dataset is structured. What we want is a data structure with which fastai's data block can easily work with. So what we need is something that gives us the filename, the label and a label which data is for training and which one is for validation. Luckily we can get exactly this by combining the meta-data:
datablock_df = pd.merge(df, df_metadata, left_on='class_id', right_on='class_id').loc[:,['path', 'class_id', 'minifigure_name', 'train-valid']]
datablock_df['is_valid'] = datablock_df['train-valid']=='valid'
datablock_df.head()
# fastai gives us a brief overview of what to check before we can make optimal use of the datablock:
#
# what are the types of our inputs and targets? Images and labels.
# where is the data? In a dataframe.
# how do we know if a sample is in the training or the validation set? A column of our dataframe.
# how do we get an image? By looking at the column path.
# how do we know the label of an image? By looking at the column minifigure_name.
# do we want to apply a function to a given sample? Yes, we need to resize everything to a given size.
# do we want to apply a function to a batch after it's created? Yes, we want data augmentation.
#
lego_block = DataBlock(blocks=(ImageBlock, CategoryBlock),
splitter=ColSplitter(),
get_x=lambda x:p/"train"/f'{x[0]}',
get_y=lambda x:x[2],
item_tfms=Resize(224),
batch_tfms=aug_transforms())
# Now our datablock is called lego_block. See how it perfectly matches together?
#
# Let me briefly explain what the different steps within our lego_block are doing: first we tell the lego_block on what we want to split our data on (the default here is col='is_valid'), then we simply put our path column (x[0]) and combine it with our path p and the folder 'train' in which is is located in. get_y tells the lego_block where to find the labels in our dataset (x[2]), we then make all of our images the same size and apply transformation on them (checkout fastai for more information).
dls = lego_block.dataloaders(datablock_df)
dls.show_batch()
# Glorious!
# fastai tries to make our life easier. This blog is intended to show you guys how to easily and quickly manage to get a great classifier with it. In the upcoming blogs I will try to better explain what is going on behind the scenes. But for now, let's enjoy how fast we can build our classifier with fastai!
learn = cnn_learner(dls, resnet34, metrics=error_rate)
learn.fine_tune(20)
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix(figsize=(12,12), dpi=60)
interp.most_confused(min_val=2)
# Not to bad I would say. However, seeing an image of <NAME> and predicting it to be <NAME> - I'm not sure how much this will impress your significant other. On the other hand, Captain America is correctly predicted 100%.
#
# But we can still try to improve our model by unfreezing the weights, to make the model even better. Let's check this out:
learn.fit_one_cycle(3, 3e-3)
# Then we will unfreeze the parameters and learn at a slightly lower learning rate:
learn.unfreeze()
learn.fit_one_cycle(2, lr_max=1e-5)
# Wow! Down to only 10% error rate. I think that's quite impressive! Let's see the confusion matrix:
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix(figsize=(12,12), dpi=60)
# With these results I am sure you will impress your significant other!
# In one of the next posts I will show you how to use Jupyter to easily set up a small Web App with the help of Binder. So stay tuned!
#
# Lasse
| _notebooks/2020-09-16-Lego-Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 64-bit
# name: python38264bite7cea40d344342a09e3a12f2e4b8d6f2
# ---
# + tags=[]
import torch
from PIL import Image
import torchvision
torch.__version__
print(torch.cuda.current_device())
# + tags=[]
cuda = torch.device('cuda')
'''
image_path = '/home/medathati/Work/SpectralSelfSupervision/Data/ILSVRC/Data/CLS-LOC/train/n03944341/n03944341_7353.JPEG'
img = Image.open(image_path).convert('RGB')
img1 = torchvision.transform(img)
'''
I = torch.rand(5,3,5,5,device=cuda)
I_fft = torch.rfft(I, signal_ndim=2, onesided = False, normalized=False)
I_mag = ((I_fft[:,:,:,:,0]**2+I_fft[:,:,:,:,1]**2)**0.5)
I_mag_nth = I_mag**(1-0.1)
I_fft[:,:,:,:,0] = I_fft[:,:,:,:,0]/I_mag_nth
I_fft[:,:,:,:,1] = I_fft[:,:,:,:,1]/I_mag_nth
# batch x channels x height x width
I_fft[I_fft!=I_fft]=0
I_hat = torch.irfft(I_fft, signal_ndim=2, onesided = False, normalized=False)
print("Shape of input tensor: ",I.shape)
print("Shape of the FFT output: ",I_fft.shape)
print("Shape of the root filter output: ",I_hat.shape)
print(I)
print("---I_ht---")
print(I_hat)
# + tags=[]
a = torch.tensor([0.0,1.5,2.0,3.0],device=cuda)
b = torch.tensor([0.0,.75,1.0,1.5],device=cuda)
c = b/a
c[c!=c]=0
print(a**2)
print(a**1)
print(b/a)
print(c)
# -
a.type()
# + tags=[]
I = torch.rand(3,3,device=cuda)
I1 = torch.min(I, dim=0,keepdim=True)
print(I)
print(I1)
# +
class MinMaxScalerVectorized(object):
"""MinMax Scaler
Transforms each channel to the range [a, b].
Parameters
----------
feature_range : tuple
Desired range of transformed data.
"""
def __init__(self, **kwargs):
self.__dict__.update(kwargs)
def __call__(self, tensor):
"""Fit features
Parameters
----------
stacked_features : tuple, list
List of stacked features.
Returns
-------
tensor
A tensor with scaled features using requested preprocessor.
"""
tensor = torch.stack(tensor)
# Feature range
a, b = self.feature_range
dist = tensor.max(dim=0, keepdim=True)[0] - tensor.min(dim=0, keepdim=True)[0]
dist[dist == 0.0] = 1.0
scale = 1.0 / dist
tensor.mul_(scale).sub_(tensor.min(dim=0, keepdim=True)[0])
tensor.mul_(b - a).add_(a)
return tensor
scaler = MinMaxScalerVectorized(feature_range=(-1, 1))
I2 = scaler(I)
print(I2)
# + tags=[]
#a = torch.randn(3, 4, 16, 16, 16, 16)
a = torch.randn(3, 3,3)
def MinMaxNormalize(X):
X_channel_flat = X.view(*(X.size()[:-2]),1,-1)
X_channel_min,_ = torch.min(X_channel_flat,len(X.size())-1, keepdim=True, out=None)
X_channel_max,_ = torch.max(X_channel_flat,len(X.size())-1, keepdim=True, out=None)
X_channel_den = X_channel_max - X_channel_min
X_channel_den[X_channel_den==0] = 1.0 # To avoid division by zero
X_normalized_flat = (X_channel_flat - X_channel_min)/X_channel_den
X_normalized = X_normalized_flat.view(X.size())
return X_normalized
print(a)
print(MinMaxNormalize(a))
# + tags=[]
n = 5
s = 0
d = 1.0/n
for i in range(n):
s = s+d
print(i,d, s)
# + tags=[]
def root_filter(img,num_filters=2):
assert(num_filters>1)
img = MinMaxNormalize(img)
imgs = [img]
#I = torch.from_numpy(img.transpose([2,0,1])).float().to('cuda')
I_fft = torch.rfft(img, signal_ndim=2, onesided = False, normalized=False)
I_mag = ((I_fft[:,:,:,:,0]**2+I_fft[:,:,:,:,1]**2)**0.5)
#I_mag_nth = I_mag**(1-0.1)
pf = 1.0/num_filters
I_mag_nth = I_mag**(pf)
for i in range(num_filters):
I_fft[:,:,:,:,0] = I_fft[:,:,:,:,0]/I_mag_nth
I_fft[:,:,:,:,1] = I_fft[:,:,:,:,1]/I_mag_nth
I_fft[I_fft!=I_fft]=0
I_hat = torch.irfft(I_fft, signal_ndim=2, onesided = False, normalized=False)
I_hat_normalized = MinMaxNormalize(I_hat)
imgs.append(I_hat_normalized)
return imgs
cuda = torch.device('cuda')
img = torch.rand(1,1,5,5,device=cuda)
imgs = root_filter(img,num_filters=2)
print(img)
print(imgs)
print(img.size())
print(len(imgs))
# + tags=[]
import numpy as np
import matplotlib.pylab as plt
image_path = '/home/medathati/Work/SpectralSelfSupervision/Data/ILSVRC/Data/CLS-LOC/train/n03944341/n03944341_7355.JPEG'
img = Image.open(image_path).convert('HSV')
na = np.array(img).astype(np.float)
print(np.max(na))
nb = np.array(na)
na[:,:,0] = 1
im = Image.fromarray(na.astype('uint8'), mode='HSV')
plt.figure()
plt.imshow(im)
plt.show()
# -
from matplotlib import pylab as plt
plt.figure()
plt.imshow(img)
plt.show()
a = np.array([365,379],dtype=np.uint8)
# + tags=[]
print(a)
# -
365-109
# +
#import Image
import numpy as np
from PIL import Image
# Source: https://stackoverflow.com/questions/7274221/changing-image-hue-with-python-pil
def rgb_to_hsv(rgb):
# Translated from source of colorsys.rgb_to_hsv
# r,g,b should be a numpy arrays with values between 0 and 255
# rgb_to_hsv returns an array of floats between 0.0 and 1.0.
rgb = rgb.astype('float')
hsv = np.zeros_like(rgb)
# in case an RGBA array was passed, just copy the A channel
hsv[..., 3:] = rgb[..., 3:]
r, g, b = rgb[..., 0], rgb[..., 1], rgb[..., 2]
maxc = np.max(rgb[..., :3], axis=-1)
minc = np.min(rgb[..., :3], axis=-1)
hsv[..., 2] = maxc
mask = maxc != minc
hsv[mask, 1] = (maxc - minc)[mask] / maxc[mask]
rc = np.zeros_like(r)
gc = np.zeros_like(g)
bc = np.zeros_like(b)
rc[mask] = (maxc - r)[mask] / (maxc - minc)[mask]
gc[mask] = (maxc - g)[mask] / (maxc - minc)[mask]
bc[mask] = (maxc - b)[mask] / (maxc - minc)[mask]
hsv[..., 0] = np.select(
[r == maxc, g == maxc], [bc - gc, 2.0 + rc - bc], default=4.0 + gc - rc)
hsv[..., 0] = (hsv[..., 0] / 6.0) % 1.0
return hsv
def hsv_to_rgb(hsv):
# Translated from source of colorsys.hsv_to_rgb
# h,s should be a numpy arrays with values between 0.0 and 1.0
# v should be a numpy array with values between 0.0 and 255.0
# hsv_to_rgb returns an array of uints between 0 and 255.
rgb = np.empty_like(hsv)
rgb[..., 3:] = hsv[..., 3:]
h, s, v = hsv[..., 0], hsv[..., 1], hsv[..., 2]
i = (h * 6.0).astype('uint8')
f = (h * 6.0) - i
p = v * (1.0 - s)
q = v * (1.0 - s * f)
t = v * (1.0 - s * (1.0 - f))
i = i % 6
conditions = [s == 0.0, i == 1, i == 2, i == 3, i == 4, i == 5]
rgb[..., 0] = np.select(conditions, [v, q, p, p, t, v], default=v)
rgb[..., 1] = np.select(conditions, [v, v, v, q, p, p], default=t)
rgb[..., 2] = np.select(conditions, [v, p, t, v, v, q], default=p)
return rgb.astype('uint8')
def shift_hue(arr,hshift):
hsv=rgb_to_hsv(arr)
#hsv[...,0]=hshift #To set the hue
hsv[...,0]= (hsv[...,0] + hshift)%1.0
rgb=hsv_to_rgb(hsv)
return rgb
#img = Image.open('tweeter.png').convert('RGBA')
#arr = np.array(img)
image_path = '/home/medathati/Work/SpectralSelfSupervision/Data/ILSVRC/Data/CLS-LOC/train/n03944341/n03944341_7355.JPEG'
img = Image.open(image_path).convert('RGB')
arr = np.array(img)
if __name__=='__main__':
green_hue = (180-78)/360.0
red_hue = (180-180)/360.0
new_img = Image.fromarray(shift_hue(arr,270/360.0), 'RGB')
new_img.save('tweeter_red.png')
plt.figure()
plt.imshow(new_img)
plt.show()
new_img = Image.fromarray(shift_hue(arr,0/360), 'RGB')
new_img.save('tweeter_green.png')
plt.figure()
plt.imshow(new_img)
plt.show()
# + tags=[]
a = np.array([1.5,0.5,0.25,0.5])
a = a+0.5
print(a)
print(a%1.0)
| try_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What percentage of samples from a dataset fall into a range of another dataset?
# This notebook demonstrates using Python to compare two datasets with one variable. More specifically, it answers "What percentage of samples from dataset A fall into a particular range (e.g., the interquartile) of dataset B?" We use Python 3, numpy, and scipy.
# !python --version
import numpy as np
import scipy.stats
# Consider the following two datasets. I have chosen easy ones for clarity, but this methodology should work on any datasets of a single variable.
a = np.arange(10, 40)
b = np.arange(0, 100)
a, b
# How many data points from dataset A fall into the interquartile range of dataset B? For this, we can create a numpy histogram from dataset B, translate that into a random variable histogram distribution using scipy, and then call the `ppf` function for the range we want. Here, I round for convenience.
hist_b = np.histogram(b, bins=100)
dist_b = scipy.stats.rv_histogram(hist_b)
start = dist_b.ppf(0.25).round(2)
finish = dist_b.ppf(0.75).round(2)
start, finish
# Next, I create a similar distribution for dataset `a`, but now I take the ppf values from dataset `b` and call the `cdf` function. Taking the difference tells me the percentage of values from dataset `a` that are in the specified range in `b`.
hist_a = np.histogram(a, bins=100)
dist_a = scipy.stats.rv_histogram(hist_a)
(dist_a.cdf(finish) - dist_a.cdf(start)).round(2)
# In our sample, the answer is 50%.
| _notebooks/2020-11-07-what-percentage-of-samples-from-a-dataset-fall-into-a-range-of-another-dataset-using-python-numpy-scipy-stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MLPerf Inference v1.0 - Reproducing Xavier results with JetPack 4.5.1
# # Table of Contents
#
# 1. [System](#system)
# 1. [Installation](#installation)
# 1. [Clone the repo](#installation_repo)
# 1. [Install the dependencies](#installation_deps)
# 1. [Link the datasets](#installation_datasets)
# 1. [Download the models](#installation_models)
# 1. [Build the harness](#installation_harness)
# 1. [ResNet50](#resnet50)
# 1. [Offline](#resnet50_offline)
# 1. [Build](#resnet50_offline_build)
# 1. [Performance](#resnet50_offline_performance)
# 1. [Accuracy](#resnet50_offline_accuracy)
# 1. [SingleStream](#resnet50_singlestream)
# 1. [Build](#resnet50_singlestream_build)
# 1. [Performance](#resnet50_singlestream_performance)
# 1. [Accuracy](#resnet50_singlestream_accuracy)
# 1. [MultiStream](#resnet50_multistream)
# 1. [Build](#resnet50_multistream_build)
# 1. [Performance](#resnet50_multistream_performance)
# 1. [Accuracy](#resnet50_multistream_accuracy)
# 1. [SSD-ResNet34](#ssd-resnet34)
# 1. [Offline](#ssd-resnet34_offline)
# 1. [Build](#ssd-resnet34_offline_build)
# 1. [Performance](#ssd-resnet34_offline_performance)
# 1. [Accuracy](#ssd-resnet34_offline_accuracy)
# 1. [SingleStream](#ssd-resnet34_singlestream)
# 1. [Build](#ssd-resnet34_singlestream_build)
# 1. [Performance](#ssd-resnet34_singlestream_performance)
# 1. [Accuracy](#ssd-resnet34_singlestream_accuracy)
# 1. [MultiStream](#ssd-resnet34_multistream)
# 1. [Build](#ssd-resnet34_multistream_build)
# 1. [Performance](#ssd-resnet34_multistream_performance)
# 1. [Accuracy](#ssd-resnet34_multistream_accuracy)
# <a name="system"></a>
# # System: [NVIDIA AGX Xavier](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/systems/AGX_Xavier_TRT.json)
# ## Stats: JetPack 4.5.1
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ sudo -H python3 -m pip install jetson-stats -U
# ...
# Successfully installed jetson-stats-3.1.0
# </pre>
#
# <pre>
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ jetson_release
# Current serial number in output stream: 19
# <b> - NVIDIA Jetson AGX Xavier [16GB]</b>
# * Jetpack 4.5.1 [L4T 32.5.1]
# * NV Power Mode: <font color="#859900">MAXN</font> - Type: <font color="#859900">0</font>
# * jetson_stats.service: <font color="#859900">active</font>
# <b> - Libraries:</b>
# * CUDA: 10.2.89
# * cuDNN: 8.0.0.180
# * TensorRT: 7.1.3.0
# * Visionworks: 1.6.0.501
# * OpenCV: 4.1.1 compiled CUDA: <font color="#DC322F">NO</font>
# * VPI: ii libnvvpi1 1.0.15 arm64 NVIDIA Vision Programming Interface library
# * Vulkan: 1.2.70
# </pre>
#
# ## OS: Ubuntu 18.04.5 LTS
#
# <pre>
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ cat /etc/lsb-release
# DISTRIB_ID=Ubuntu
# DISTRIB_RELEASE=18.04
# DISTRIB_CODENAME=bionic
# DISTRIB_DESCRIPTION="Ubuntu 18.04.5 LTS"
# </pre>
#
# ## CPU: 8-core ARMv8 @ 2265 MHz
#
# <pre>
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ lscpu
# Architecture: aarch64
# Byte Order: Little Endian
# CPU(s): 8
# On-line CPU(s) list: 0-7
# Thread(s) per core: 1
# Core(s) per socket: 2
# Socket(s): 4
# Vendor ID: Nvidia
# Model: 0
# Model name: ARMv8 Processor rev 0 (v8l)
# Stepping: 0x0
# CPU max MHz: 2265.6001
# CPU min MHz: 115.2000
# BogoMIPS: 62.50
# L1d cache: 64K
# L1i cache: 128K
# L2 cache: 2048K
# L3 cache: 4096K
# Flags: fp asimd evtstrm aes pmull sha1 sha2 crc32 atomics fphp asimdhp
# </pre>
#
# ## GPU
#
# <pre>
# <font color="#859900"><b><EMAIL>on@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ sudo jetson_clocks
# <font color="#859900"><b><EMAIL>on@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ sudo jetson_clocks --show
# SOC family:tegra194 Machine:Jetson-AGX
# Online CPUs: 0-7
# cpu0: Online=1 Governor=schedutil MinFreq=2265600 MaxFreq=2265600 CurrentFreq=2265600 IdleStates: C1=0 c6=0
# cpu1: Online=1 Governor=schedutil MinFreq=2265600 MaxFreq=2265600 CurrentFreq=2265600 IdleStates: C1=0 c6=0
# cpu2: Online=1 Governor=schedutil MinFreq=2265600 MaxFreq=2265600 CurrentFreq=2265600 IdleStates: C1=0 c6=0
# cpu3: Online=1 Governor=schedutil MinFreq=2265600 MaxFreq=2265600 CurrentFreq=2265600 IdleStates: C1=0 c6=0
# cpu4: Online=1 Governor=schedutil MinFreq=2265600 MaxFreq=2265600 CurrentFreq=2265600 IdleStates: C1=0 c6=0
# cpu5: Online=1 Governor=schedutil MinFreq=2265600 MaxFreq=2265600 CurrentFreq=2265600 IdleStates: C1=0 c6=0
# cpu6: Online=1 Governor=schedutil MinFreq=2265600 MaxFreq=2265600 CurrentFreq=2265600 IdleStates: C1=0 c6=0
# cpu7: Online=1 Governor=schedutil MinFreq=2265600 MaxFreq=2265600 CurrentFreq=2265600 IdleStates: C1=0 c6=0
# GPU MinFreq=1377000000 MaxFreq=1377000000 CurrentFreq=1377000000
# EMC MinFreq=204000000 MaxFreq=2133000000 CurrentFreq=2133000000 FreqOverride=1
# Fan: PWM=0
# NV Power Mode: MAXN
# </pre>
#
#
# <pre>
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ ck compile program:tool-print-cuda-devices
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ ck run program:tool-print-cuda-devices
# ...
# GPU Device ID: 0
# GPU Name: Xavier
# GPU compute capability: 7.2
# CUDA driver version: 10.2
# CUDA runtime version: 10.2
# Global memory: 33479647232
# Max clock rate: 1377.000000 MHz
# Total amount of shared memory per block: 49152
# Total number of registers available per block: 65536
# Warp size: 32
# Maximum number of threads per multiprocessor: 2048
# Maximum number of threads per block: 1024
# Max dimension size of a thread block X: 1024
# Max dimension size of a thread block Y: 1024
# Max dimension size of a thread block Z: 64
# Max dimension size of a grid size X: 2147483647
# Max dimension size of a grid size Y: 65535
# Max dimension size of a grid size Z: 65535
# </pre>
#
#
# ## Disks
#
# <pre><font color="#859900"><b><EMAIL>on@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ df -h
# Filesystem Size Used Avail Use% Mounted on
# /dev/mmcblk0p1 28G 17G 9.4G 64% /
# none 16G 0 16G 0% /dev
# tmpfs 16G 56K 16G 1% /dev/shm
# tmpfs 16G 42M 16G 1% /run
# tmpfs 5.0M 4.0K 5.0M 1% /run/lock
# tmpfs 16G 0 16G 0% /sys/fs/cgroup
# /dev/mmcblk1p1 361G 314G 28G 92% /sd
# tmpfs 3.2G 12K 3.2G 1% /run/user/120
# tmpfs 3.2G 0 3.2G 0% /run/user/1000
# </pre>
# <a name="installation"></a>
# # Installation
# <a name="installation_repo"></a>
# ## Clone the v1.0 results repo
#
# <pre><font color="#859900"><b><EMAIL>on@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ git clone https://github.com/mlcommons/inference_results_v1.0.git</pre>
# <a name="installation_deps"></a>
# ## Install the dependencies
#
# To [quote](https://github.com/mlcommons/inference_results_v1.0/tree/master/closed/NVIDIA#prerequisites) NVIDIA:
# > Note that this might take a while, on the order of several hours.
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ chmod +x inference_results_v1.0/closed/NVIDIA/scripts/install_xavier_dependencies.sh
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ time sudo inference_results_v1.0/closed/NVIDIA/scripts/install_xavier_dependencies.sh
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ python3 -m pip install onnx
# </pre>
# <a name="installation_datasets"></a>
# ## Link the datasets
#
# We reused the datasets we generated while reproducing the v0.5 results.
#
# <pre>
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ export MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ mkdir $MLPERF_SCRATCH_PATH
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ cd $MLPERF_SCRATCH_PATH
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/mlperf_scratch_path</b></font>$ ln -s /datasets/inference_results_v0.5-nvidia/closed/NVIDIA/build/preprocessed_data preprocessed_data
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/mlperf_scratch_path</b></font>$ ln -s /datasets/inference_results_v0.5-nvidia/closed/NVIDIA/build/data data
# <font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/mlperf_scratch_path</b></font>$ ls -la /datasets/mlperf_scratch_path/
# total 20
# drwxrwsr-x 3 anton dvdt 4096 Jan 28 11:10 <font color="#268BD2"><b>.</b></font>
# drwxrwsr-x 17 root dvdt 4096 Jan 28 09:53 <font color="#268BD2"><b>..</b></font>
# lrwxrwxrwx 1 anton dvdt 64 Jan 28 11:10 <font color="#2AA198"><b>data</b></font> -> <font color="#268BD2"><b>/datasets/inference_results_v0.5-nvidia/closed/NVIDIA/build/data</b></font>
# drwxrwsr-x 5 anton dvdt 4096 Jan 28 09:54 <font color="#268BD2"><b>models</b></font>
# lrwxrwxrwx 1 an<NAME>dt 77 Jan 28 11:10 <font color="#2AA198"><b>preprocessed_data</b></font> -> <font color="#268BD2"><b>/datasets/inference_results_v0.5-nvidia/closed/NVIDIA/build/preprocessed_data</b></font>
# </pre>
# <a name="installation_models"></a>
# ## Download the models
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ git diff Makefile
# <b>diff --git a/closed/NVIDIA/Makefile b/closed/NVIDIA/Makefile</b>
# <b>index e7647663..2b9356f6 100644</b>
# <b>--- a/closed/NVIDIA/Makefile</b>
# <b>+++ b/closed/NVIDIA/Makefile</b>
# <font color="#2AA198">@@ -360,7 +360,7 @@</font> endif
#
# ############################## DOWNLOAD_MODEL ##############################
#
# <font color="#DC322F">-BENCHMARKS = resnet50 ssd-resnet34 ssd-mobilenet bert dlrm rnnt 3d-unet</font>
# <font color="#859900">+BENCHMARKS = resnet50 ssd-resnet34 ssd-mobilenet # bert dlrm rnnt 3d-unet</font>
#
# .PHONY: download_model
# download_model: link_dirs
#
# <font color="#2AA198">@@ -474,7 +474,6 @@</font> ifeq ($(ARCH), x86_64)
# cd build/plugins/DLRMBottomMLPPlugin \
# && cmake -DCMAKE_BUILD_TYPE=$(BUILD_TYPE) $(PROJECT_ROOT)/code/plugin/DLRMBottomMLPPlugin \
# && make -j
# <font color="#DC322F">-endif</font>
# mkdir -p build/plugins/RNNTOptPlugin
# cd build/plugins/RNNTOptPlugin \
# && cmake -DCMAKE_BUILD_TYPE=$(BUILD_TYPE) $(PROJECT_ROOT)/code/plugin/RNNTOptPlugin \
# <font color="#2AA198">@@ -491,6 +490,7 @@</font> endif
# cd build/plugins/conv3D1X1X1K4Plugin \
# && cmake -DCMAKE_BUILD_TYPE=$(BUILD_TYPE) $(PROJECT_ROOT)/code/plugin/conv3D1X1X1K4Plugin \
# && make -j
# <font color="#859900">+endif</font>
#
# # Build LoadGen.
# .PHONY: build_loadgen
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path make download_model
# ...
# Finished downloading all the models!
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ make build
# ...
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ ls -la build/
# total 28
# drwxrwsr-x 7 anton dvdt 4096 Apr 27 11:48 <font color="#268BD2"><b>.</b></font>
# drwxrwsr-x 14 anton dvdt 4096 Apr 27 11:29 <font color="#268BD2"><b>..</b></font>
# drwxrwsr-x 2 anton dvdt 4096 Apr 27 11:49 <font color="#268BD2"><b>bin</b></font>
# lrwxrwxrwx 1 anton dvdt 34 Apr 27 11:48 <font color="#2AA198"><b>data</b></font> -> <font color="#268BD2"><b>/datasets/mlperf_scratch_path/data</b></font>
# drwxrwsr-x 5 anton dvdt 4096 Apr 27 11:49 <font color="#268BD2"><b>harness</b></font>
# drwxrwsr-x 16 anton dvdt 4096 Apr 27 11:24 <font color="#268BD2"><b>inference</b></font>
# lrwxrwxrwx 1 anton dvdt 36 Apr 27 11:48 <font color="#2AA198"><b>models</b></font> -> <font color="#268BD2"><b>/datasets/mlperf_scratch_path/models</b></font>
# drwxrwsr-x 5 anton dvdt 4096 Apr 27 11:27 <font color="#268BD2"><b>plugins</b></font>
# drwxrwsr-x 7 anton dvdt 4096 Apr 27 11:24 <font color="#268BD2"><b>power-dev</b></font>
# lrwxrwxrwx 1 anton dvdt 47 Apr 27 11:48 <font color="#2AA198"><b>preprocessed_data</b></font> -> <font color="#268BD2"><b>/datasets/mlperf_scratch_path/preprocessed_data</b></font>
# </pre>
# <a name="resnet50"></a>
# # ResNet50
# <a name="resnet50_offline"></a>
# ## Offline
# ### [Config](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/configs/resnet50/Offline/config.json)
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>~</b></font>$ grep AGX_Xavier /datasets/inference_results_v1.0/closed/NVIDIA/configs/resnet50/Offline/config.json -A 15
# "<font color="#DC322F"><b>AGX_Xavier</b></font>": {
# "concurrent_offline_expected_qps": 2181,
# "config_ver": {
# "maxq": {
# "concurrent_offline_expected_qps": 1530
# }
# },
# "dla_batch_size": 32,
# "dla_copy_streams": 1,
# "dla_core": 0,
# "dla_inference_streams": 1,
# "dla_offline_expected_qps": 396,
# "gpu_batch_size": 64,
# "gpu_copy_streams": 1,
# "gpu_offline_expected_qps": 1478.33
# },
# </pre>
# <a name="resnet50_offline_build"></a>
# ### Build
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make generate_engines RUN_ARGS="--benchmarks=resnet50 --scenarios=offline"
# ...
# [2021-04-27 17:07:37,429 main.py:112 INFO] Finished building engines for resnet50 benchmark in Offline scenario.
# Time taken to generate engines: 65.42682266235352 seconds
#
# real 1m6.766s
# user 0m26.980s
# sys 0m6.420s
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ find . \
# -name resnet50-Offline*.plan -exec du -hs {} \;
# 48M ./build/engines/AGX_Xavier/resnet50/Offline/resnet50-Offline-gpu-b64-int8.default.plan
# 33M ./build/engines/AGX_Xavier/resnet50/Offline/resnet50-Offline-dla-b32-int8.default.plan
# </pre>
# <a name="resnet50_offline_performance"></a>
# ### Performance
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/resnet50/Offline/performance/run_1/mlperf_log_summary.txt#L7)
#
# <pre>
# Samples per second: 2071.51
# Result is : VALID
# </pre>
#
# #### Reproduced experiment (~53 W)
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=resnet50 --scenarios=offline --test_mode=PerformanceOnly"
# ...
# ================================================
# MLPerf Results Summary
# ================================================
# SUT name : LWIS_Server
# Scenario : Offline
# Mode : PerformanceOnly
# Samples per second: 2086.92
# Result is : VALID
# Min duration satisfied : Yes
# Min queries satisfied : Yes
#
# ================================================
# Additional Stats
# ================================================
# Min latency (ns) : 155755993
# Max latency (ns) : 689754096859
# Mean latency (ns) : 345039571231
# 50.00 percentile latency (ns) : 345089223369
# 90.00 percentile latency (ns) : 620810076791
# 95.00 percentile latency (ns) : 655270079960
# 97.00 percentile latency (ns) : 669065261726
# 99.00 percentile latency (ns) : 682850845028
# 99.90 percentile latency (ns) : 689068993553
#
# ================================================
# Test Parameters Used
# ================================================
# samples_per_query : 1439460
# target_qps : 2181
# target_latency (ns): 0
# max_async_queries : 1
# min_duration (ms): 600000
# max_duration (ms): 0
# min_query_count : 1
# max_query_count : 0
# qsl_rng_seed : 7322528924094909334
# sample_index_rng_seed : 1570999273408051088
# schedule_rng_seed : 3507442325620259414
# accuracy_log_rng_seed : 0
# accuracy_log_probability : 0
# accuracy_log_sampling_target : 0
# print_timestamps : 0
# performance_issue_unique : 0
# performance_issue_same : 0
# performance_issue_same_index : 0
# performance_sample_count : 2048
#
# No warnings encountered during test.
#
# No errors encountered during test.
#
# Finished running actual test.
# Device Device:0 processed:
# 1 batches of size 36
# 15545 batches of size 64
# Memcpy Calls: 0
# PerSampleCudaMemcpy Calls: 10368
# BatchedCudaMemcpy Calls: 15384
# Device Device:0.DLA-0 processed:
# 6938 batches of size 32
# Memcpy Calls: 0
# PerSampleCudaMemcpy Calls: 0
# BatchedCudaMemcpy Calls: 0
# Device Device:0.DLA-1 processed:
# 6954 batches of size 32
# Memcpy Calls: 0
# PerSampleCudaMemcpy Calls: 0
# BatchedCudaMemcpy Calls: 0
# &&&& PASSED Default_Harness # ./build/bin/harness_default
# [2021-04-27 17:22:17,724 main.py:280 INFO] Result: result_samples_per_second: 2086.92, Result is VALID
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-Offline:
# resnet50: result_samples_per_second: 2086.92, Result is VALID
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-Offline:
# resnet50: No accuracy results in PerformanceOnly mode.
#
#
# real 11m52.559s
# user 11m30.528s
# sys 0m12.236s
# </pre>
# <a name="resnet50_offline_accuracy"></a>
# ### Accuracy
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/resnet50/Offline/accuracy/accuracy.txt)
#
# <pre>
# accuracy=76.074%, good=38037, total=50000
# hash=3dc4add63a23f3f1f23e44abbf6b5e8f9c8a12e44412e4aa6a77f09cac22eee2
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=resnet50 --scenarios=offline --test_mode=AccuracyOnly"
# ...
# accuracy=76.040%, good=38020, total=50000
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-Offline:
# resnet50: Cannot find performance result. Maybe you are running in AccuracyOnly mode.
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-Offline:
# resnet50: Accuracy = 76.040, Threshold = 75.695. Accuracy test PASSED.
#
#
# real 3m29.147s
# user 0m58.836s
# sys 0m17.444s
# </pre>
# <a name="resnet50_singlestream"></a>
# ## SingleStream
# ### [Config](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/configs/resnet50/SingleStream/config.json)
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ grep AGX_Xavier /datasets/inference_results_v1.0/closed/NVIDIA/configs/resnet50/SingleStream/config.json -A 6
# "<font color="#DC322F"><b>AGX_Xavier</b></font>": {
# "config_ver": {
# "maxq": {}
# },
# "gpu_single_stream_expected_latency_ns": 2273000,
# "use_graphs": false
# },
# </pre>
# <a name="resnet50_singlestream_build"></a>
# ### Build
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make generate_engines RUN_ARGS="--benchmarks=resnet50 --scenarios=singlestream"
# ...
# [2021-04-27 17:35:34,949 main.py:112 INFO] Finished building engines for resnet50 benchmark in SingleStream scenario.
# Time taken to generate engines: 42.95994567871094 seconds
#
# real 0m44.209s
# user 0m23.192s
# sys 0m4.620s
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ find . \
# -name resnet50-SingleStream*.plan -exec du -hs {} \;
# 44M ./build/engines/AGX_Xavier/resnet50/SingleStream/resnet50-SingleStream-gpu-b1-int8.default.plan
# </pre>
# <a name="resnet50_singlestream_performance"></a>
# ### Performance
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/resnet50/SingleStream/performance/run_1/mlperf_log_summary.txt#L7)
#
# <pre>
# 90th percentile latency (ns) : 1980582
# Result is : VALID
# Min duration satisfied : Yes
# Min queries satisfied : Yes
# ...
# QPS w/ loadgen overhead : 509.00
# QPS w/o loadgen overhead : 512.43
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=resnet50 --scenarios=SingleStream --test_mode=PerformanceOnly"
# ...
# ================================================
# MLPerf Results Summary
# ================================================
# SUT name : LWIS_Server
# Scenario : Single Stream
# Mode : Performance
# 90th percentile latency (ns) : 2078719
# Result is : VALID
# Min duration satisfied : Yes
# Min queries satisfied : Yes
#
# ================================================
# Additional Stats
# ================================================
# QPS w/ loadgen overhead : 485.07
# QPS w/o loadgen overhead : 488.48
#
# Min latency (ns) : 1980890
# Max latency (ns) : 15165301
# Mean latency (ns) : 2047146
# 50.00 percentile latency (ns) : 2038621
# 90.00 percentile latency (ns) : 2078719
# 95.00 percentile latency (ns) : 2098912
# 97.00 percentile latency (ns) : 2118465
# 99.00 percentile latency (ns) : 2187844
# 99.90 percentile latency (ns) : 2564821
#
# ================================================
# Test Parameters Used
# ================================================
# samples_per_query : 1
# target_qps : 439.947
# target_latency (ns): 0
# max_async_queries : 1
# min_duration (ms): 60000
# max_duration (ms): 0
# min_query_count : 1024
# max_query_count : 0
# qsl_rng_seed : 12786827339337101903
# sample_index_rng_seed : 12640797754436136668
# schedule_rng_seed : 3135815929913719677
# accuracy_log_rng_seed : 0
# accuracy_log_probability : 0
# accuracy_log_sampling_target : 0
# print_timestamps : false
# performance_issue_unique : false
# performance_issue_same : false
# performance_issue_same_index : 0
# performance_sample_count : 2048
#
# No warnings encountered during test.
#
# No errors encountered during test.
# Finished running actual test.
# Device Device:0 processed:
# 29106 batches of size 1
# Memcpy Calls: 0
# PerSampleCudaMemcpy Calls: 0
# BatchedCudaMemcpy Calls: 29106
# &&&& PASSED Default_Harness # ./build/bin/harness_default
# [2021-01-29 23:25:15,883 main.py:341 INFO] Result: 90th percentile latency (ns) : 2078719 and Result is : VALID
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-SingleStream:
# resnet50: 90th percentile latency (ns) : 2078719 and Result is : VALID
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-SingleStream:
# resnet50: No accuracy results in PerformanceOnly mode.
#
#
# real 1m16.374s
# user 1m6.484s
# sys 0m5.004s
#
# </pre>
# <a name="resnet50_singlestream_accuracy"></a>
# ### Accuracy
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/resnet50/SingleStream/accuracy/accuracy.txt)
#
# <pre>
# accuracy=76.064%, good=38032, total=50000
# hash=7458cd3f1154670a0d063c87b38d2eba7aa8c1921f2558a46333cfef8d9b4036
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=resnet50 --scenarios=SingleStream --test_mode=AccuracyOnly"
# ...
# accuracy=76.078%, good=38039, total=50000
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-SingleStream:
# resnet50: Cannot find performance result. Maybe you are running in AccuracyOnly mode.
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-SingleStream:
# resnet50: Accuracy = 76.078, Threshold = 75.695. Accuracy test PASSED.
#
#
# real 4m27.773s
# user 2m9.392s
# sys 0m11.332s
# </pre>
# <a name="resnet50_multistream"></a>
# ## MultiStream
# ### [Config](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/configs/resnet50/MultiStream/config.json)
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ grep AGX_Xavier /datasets/inference_results_v1.0/closed/NVIDIA/configs/resnet50/MultiStream/config.json -A 14
# "<font color="#DC322F"><b>AGX_Xavier</b></font>": {
# "concurrent_multi_stream_samples_per_query": 96,
# "config_ver": {
# "maxq": {}
# },
# "dla_batch_size": 15,
# "dla_copy_streams": 2,
# "dla_core": 0,
# "dla_inference_streams": 4,
# "dla_multi_stream_samples_per_query": 15,
# "gpu_batch_size": 66,
# "gpu_copy_streams": 2,
# "gpu_inference_streams": 4,
# "gpu_multi_stream_samples_per_query": 66
# },
# </pre>
# <a name="resnet50_multistream_build"></a>
# ### Build
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make generate_engines RUN_ARGS="--benchmarks=resnet50 --scenarios=multistream"
# ...
# [2021-04-27 20:34:39,925 main.py:112 INFO] Finished building engines for resnet50 benchmark in MultiStream scenario.
# Time taken to generate engines: 118.26688885688782 seconds
#
# real 1m59.516s
# user 0m35.180s
# sys 0m15.528s
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ find . \
# -name resnet50-MultiStream*.plan -exec du -hs {} \;
# 96M ./build/engines/AGX_Xavier/resnet50/MultiStream/resnet50-MultiStream-gpu-b66-int8.default.plan
# 29M ./build/engines/AGX_Xavier/resnet50/MultiStream/resnet50-MultiStream-dla-b15-int8.default.plan
# </pre>
# <a name="resnet50_multistream_performance"></a>
# ### Performance
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/resnet50/MultiStream/performance/run_1/mlperf_log_summary.txt#L7)
#
# <pre>
# Samples per query : 96
# Result is : VALID
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=resnet50 --scenarios=MultiStream --test_mode=PerformanceOnly"
# [2021-04-27 20:38:49,061 main.py:701 INFO] Detected System ID: AGX_Xavier
# [2021-04-27 20:38:49,062 main.py:529 INFO] Using config files: configs/resnet50/MultiStream/config.json
# [2021-04-27 20:38:49,063 __init__.py:341 INFO] Parsing config file configs/resnet50/MultiStream/config.json ...
# [2021-04-27 20:38:49,063 main.py:542 INFO] Processing config "AGX_Xavier_resnet50_MultiStream"
# [2021-04-27 20:38:49,064 main.py:224 INFO] Running harness for resnet50 benchmark in MultiStream scenario...
# ...
# ================================================
# MLPerf Results Summary
# ================================================
# SUT name : LWIS_Server
# Scenario : MultiStream
# Mode : PerformanceOnly
# Samples per query : 96
# Result is : INVALID
# Performance constraints satisfied : NO
# Min duration satisfied : Yes
# Min queries satisfied : Yes
# Recommendations:
# * Reduce samples per query to improve latency.
#
# ================================================
# Additional Stats
# ================================================
# Intervals between each IssueQuery: "qps" : 20, "ms" : 50
# 50.00 percentile : 1
# 90.00 percentile : 1
# 95.00 percentile : 1
# 97.00 percentile : 1
# 99.00 percentile : 2
# 99.90 percentile : 2
#
# Per-query latency: "target_ns" : 50000000, "target_ms" : 50
# 50.00 percentile latency (ns) : 48702492
# 90.00 percentile latency (ns) : 49325203
# 95.00 percentile latency (ns) : 49511578
# 97.00 percentile latency (ns) : 49645776
# 99.00 percentile latency (ns) : 49978295
# 99.90 percentile latency (ns) : 50705066
#
# Per-sample latency:
# Min latency (ns) : 45861628
# Max latency (ns) : 182167011
# Mean latency (ns) : 48390461
# 50.00 percentile latency (ns) : 48353859
# 90.00 percentile latency (ns) : 49233149
# 95.00 percentile latency (ns) : 49415884
# 97.00 percentile latency (ns) : 49546249
# 99.00 percentile latency (ns) : 49851112
# 99.90 percentile latency (ns) : 50605105
#
# ================================================
# Test Parameters Used
# ================================================
# samples_per_query : 96
# target_qps : 20
# target_latency (ns): 50000000
# max_async_queries : 1
# min_duration (ms): 600000
# max_duration (ms): 0
# min_query_count : 270336
# max_query_count : 0
# qsl_rng_seed : 7322528924094909334
# sample_index_rng_seed : 1570999273408051088
# schedule_rng_seed : 3507442325620259414
# accuracy_log_rng_seed : 0
# accuracy_log_probability : 0
# accuracy_log_sampling_target : 0
# print_timestamps : 0
# performance_issue_unique : 0
# performance_issue_same : 0
# performance_issue_same_index : 0
# performance_sample_count : 2048
#
# No warnings encountered during test.
#
# No errors encountered during test.
# ...
# [2021-04-28 10:17:24,243 main.py:280 INFO] Result: requested_multi_stream_samples_per_query: 96, Result is INVALID
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-MultiStream:
# resnet50: requested_multi_stream_samples_per_query: 96, Result is INVALID
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-MultiStream:
# resnet50: No accuracy results in PerformanceOnly mode.
#
#
# real 228m2.179s
# user 12m59.700s
# sys 3m27.784s
# </pre>
# <a name="resnet50_multistream_accuracy"></a>
# ### Accuracy
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/resnet50/MultiStream/accuracy/accuracy.txt)
#
# <pre>
# accuracy=76.008%, good=38004, total=50000
# hash=12988c6c56d7af58b5e816d9b9d64b20f5ae80497e355dc51653ed2433b583bb
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=resnet50 --scenarios=MultiStream --test_mode=AccuracyOnly"
# ...
# accuracy=76.020%, good=38010, total=50000
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-MultiStream:
# resnet50: requested_multi_stream_samples_per_query: 96, Result validity unknown
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-MultiStream:
# resnet50: Accuracy = 76.020, Threshold = 75.695. Accuracy test PASSED.
#
#
# real 1m2.343s
# user 0m24.100s
# sys 0m7.468s
# </pre>
# <a name="ssd-resnet34"></a>
# # SSD-ResNet34
# <a name="ssd-resnet34_offline"></a>
# ## Offline
# ### [Config](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/configs/ssd-resnet34/Offline/config.json)
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ grep AGX_Xavier /datasets/inference_results_v1.0/closed/NVIDIA/configs/ssd-resnet34/Offline/config.json -A 16
# "<font color="#DC322F"><b>AGX_Xavier</b></font>": {
# "concurrent_offline_expected_qps": 56,
# "config_ver": {
# "maxq": {
# "concurrent_offline_expected_qps": 42
# }
# },
# "dla_batch_size": 1,
# "dla_copy_streams": 1,
# "dla_core": 0,
# "dla_inference_streams": 1,
# "dla_offline_expected_qps": 10,
# "gpu_batch_size": 2,
# "gpu_copy_streams": 4,
# "gpu_inference_streams": 1,
# "gpu_offline_expected_qps": 35.1243
# },
# </pre>
# <a name="ssd-resnet34_offline_build"></a>
# ### Build
#
# <pre><font color="#859900"><b><EMAIL>@<EMAIL></b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make generate_engines RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=offline"
# ...
# [2021-04-28 21:32:57,015 main.py:112 INFO] Finished building engines for ssd-resnet34 benchmark in Offline scenario.
# Time taken to generate engines: 381.31431436538696 seconds
#
# real 6m23.144s
# user 2m16.348s
# sys 0m46.172s
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ find . \
# -name ssd-resnet34-Offline*.plan -exec du -hs {} \;
# 147M ./build/engines/AGX_Xavier/ssd-resnet34/Offline/ssd-resnet34-Offline-gpu-b2-int8.default.plan
# 23M ./build/engines/AGX_Xavier/ssd-resnet34/Offline/ssd-resnet34-Offline-dla-b1-int8.default.plan
# </pre>
# <a name="ssd-resnet34_offline_performance"></a>
# ### Performance
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/ssd-resnet34/Offline/performance/run_1/mlperf_log_summary.txt#L7)
#
# <pre>
# Samples per second: 56.6721
# Result is : VALID
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=Offline --test_mode=PerformanceOnly"
# ...
# ================================================
# MLPerf Results Summary
# ================================================
# SUT name : LWIS_Server
# Scenario : Offline
# Mode : PerformanceOnly
# Samples per second: 51.1607
# Result is : VALID
# Min duration satisfied : Yes
# Min queries satisfied : Yes
#
# ================================================
# Additional Stats
# ================================================
# Min latency (ns) : 195940306
# Max latency (ns) : 722430107208
# Mean latency (ns) : 361257062980
# 50.00 percentile latency (ns) : 361271790879
# 90.00 percentile latency (ns) : 650292988467
# 95.00 percentile latency (ns) : 686348820151
# 97.00 percentile latency (ns) : 700764011699
# 99.00 percentile latency (ns) : 715213570646
# 99.90 percentile latency (ns) : 721724962037
#
# ================================================
# Test Parameters Used
# ================================================
# samples_per_query : 36960
# target_qps : 56
# target_latency (ns): 0
# max_async_queries : 1
# min_duration (ms): 600000
# max_duration (ms): 0
# min_query_count : 1
# max_query_count : 0
# qsl_rng_seed : 7322528924094909334
# sample_index_rng_seed : 1570999273408051088
# schedule_rng_seed : 3507442325620259414
# accuracy_log_rng_seed : 0
# accuracy_log_probability : 0
# accuracy_log_sampling_target : 0
# print_timestamps : 0
# performance_issue_unique : 0
# performance_issue_same : 0
# performance_issue_same_index : 0
# performance_sample_count : 64
#
# No warnings encountered during test.
#
# No errors encountered during test.
# ...
# [2021-04-28 21:49:22,497 main.py:280 INFO] Result: result_samples_per_second: 51.1607, Result is VALID
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-Offline:
# ssd-resnet34: result_samples_per_second: 51.1607, Result is VALID
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-Offline:
# ssd-resnet34: No accuracy results in PerformanceOnly mode.
#
#
# real 12m18.608s
# user 12m4.692s
# sys 0m8.680s
# </pre>
# <a name="ssd-resnet34_offline_accuracy"></a>
# ### Accuracy
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/ssd-resnet34/Offline/accuracy/accuracy.txt)
#
# <pre>
# Average Precision (AP) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.201
# Average Precision (AP) @[ IoU=0.50 | area= all | maxDets=100 ] = 0.379
# Average Precision (AP) @[ IoU=0.75 | area= all | maxDets=100 ] = 0.188
# Average Precision (AP) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.121
# Average Precision (AP) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.257
# Average Precision (AP) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.237
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 1 ] = 0.203
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 10 ] = 0.332
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.353
# Average Recall (AR) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.180
# Average Recall (AR) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.412
# Average Recall (AR) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.429
# mAP=20.052%
# hash=74f272c16708cc31a2c487a16086c6d6a03b121cf34156f94b036c4fad0ad7ac
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=Offline --test_mode=AccuracyOnly"
# ...
# Average Precision (AP) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.200
# Average Precision (AP) @[ IoU=0.50 | area= all | maxDets=100 ] = 0.380
# Average Precision (AP) @[ IoU=0.75 | area= all | maxDets=100 ] = 0.186
# Average Precision (AP) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.121
# Average Precision (AP) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.257
# Average Precision (AP) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.238
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 1 ] = 0.203
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 10 ] = 0.331
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.352
# Average Recall (AR) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.179
# Average Recall (AR) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.412
# Average Recall (AR) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.427
# mAP=20.031%
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-Offline:
# ssd-resnet34: Cannot find performance result. Maybe you are running in AccuracyOnly mode.
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-Offline:
# ssd-resnet34: Accuracy = 20.031, Threshold = 19.800. Accuracy test PASSED.
#
#
# real 10m50.156s
# user 6m3.140s
# sys 0m23.652s
# </pre>
# <a name="ssd-resnet34_singlestream"></a>
# ## SingleStream
# ### [Config](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/configs/ssd-resnet34/SingleStream/config.json)
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ grep AGX_Xavier /datasets/inference_results_v1.0/closed/NVIDIA/configs/ssd-resnet34/SingleStream/config.json -A 11
# "<font color="#DC322F"><b>AGX_Xavier</b></font>": {
# "gpu_batch_size": 1,
# "gpu_copy_streams": 1,
# "gpu_inference_streams": 1,
# "gpu_single_stream_expected_latency_ns": 29478000,
# "input_dtype": "int8",
# "input_format": "linear",
# "map_path": "data_maps/coco/val_map.txt",
# "precision": "int8",
# "tensor_path": "${PREPROCESSED_DATA_DIR}/coco/val2017/SSDResNet34/int8_linear",
# "use_graphs": false
# },
# </pre>
# <a name="ssd-resnet34_singlestream_build"></a>
# ### Build
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make generate_engines RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=singlestream"
# ...
# [2021-01-29 17:45:05,185 main.py:153 INFO] Finished building engines for ssd-resnet34 benchmark in SingleStream scenario.
# Time taken to generate engines: 63.35827445983887 seconds
#
# real 1m5.118s
# user 0m15.400s
# sys 0m5.076s
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ find . \
# -name ssd-resnet34-SingleStream*.plan -exec du -hs {} \;
# 37M ./build/engines/AGX_Xavier/ssd-resnet34/SingleStream/ssd-resnet34-SingleStream-gpu-b1-int8.default.plan
# </pre>
# <a name="ssd-resnet34_singlestream_performance"></a>
# ### Performance
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/ssd-resnet34/SingleStream/performance/run_1/mlperf_log_summary.txt#L7)
#
# <pre>
# 90th percentile latency (ns) : 28531845
# Result is : VALID
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=SingleStream --test_mode=PerformanceOnly"
# ...
# ================================================
# MLPerf Results Summary
# ================================================
# SUT name : LWIS_Server
# Scenario : Single Stream
# Mode : Performance
# 90th percentile latency (ns) : 28554901
# Result is : VALID
# Min duration satisfied : Yes
# Min queries satisfied : Yes
#
# ================================================
# Additional Stats
# ================================================
# QPS w/ loadgen overhead : 35.17
# QPS w/o loadgen overhead : 35.23
#
# Min latency (ns) : 28109905
# Max latency (ns) : 32556991
# Mean latency (ns) : 28385289
# 50.00 percentile latency (ns) : 28361924
# 90.00 percentile latency (ns) : 28554901
# 95.00 percentile latency (ns) : 28625392
# 97.00 percentile latency (ns) : 28688816
# 99.00 percentile latency (ns) : 28884610
# 99.90 percentile latency (ns) : 29821107
#
# ================================================
# Test Parameters Used
# ================================================
# samples_per_query : 1
# target_qps : 33.9236
# target_latency (ns): 0
# max_async_queries : 1
# min_duration (ms): 60000
# max_duration (ms): 0
# min_query_count : 1024
# max_query_count : 0
# qsl_rng_seed : 12786827339337101903
# sample_index_rng_seed : 12640797754436136668
# schedule_rng_seed : 3135815929913719677
# accuracy_log_rng_seed : 0
# accuracy_log_probability : 0
# accuracy_log_sampling_target : 0
# print_timestamps : false
# performance_issue_unique : false
# performance_issue_same : false
# performance_issue_same_index : 0
# performance_sample_count : 64
#
# No warnings encountered during test.
#
# No errors encountered during test.
# Finished running actual test.
# Device Device:0 processed:
# 2111 batches of size 1
# Memcpy Calls: 0
# PerSampleCudaMemcpy Calls: 0
# BatchedCudaMemcpy Calls: 2111
# &&&& PASSED Default_Harness # ./build/bin/harness_default
# [2021-01-29 22:39:22,362 main.py:341 INFO] Result: 90th percentile latency (ns) : 28554901 and Result is : VALID
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-SingleStream:
# ssd-resnet34: 90th percentile latency (ns) : 28554901 and Result is : VALID
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-SingleStream:
# ssd-resnet34: No accuracy results in PerformanceOnly mode.
#
#
# real 1m9.437s
# user 1m4.720s
# sys 0m3.964s
# </pre>
# <a name="ssd-resnet34_singlestream_accuracy"></a>
# ### Accuracy
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/ssd-resnet34/SingleStream/accuracy/accuracy.txt)
#
# <pre>
# Average Precision (AP) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.201
# Average Precision (AP) @[ IoU=0.50 | area= all | maxDets=100 ] = 0.381
# Average Precision (AP) @[ IoU=0.75 | area= all | maxDets=100 ] = 0.188
# Average Precision (AP) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.121
# Average Precision (AP) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.258
# Average Precision (AP) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.238
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 1 ] = 0.203
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 10 ] = 0.332
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.353
# Average Recall (AR) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.179
# Average Recall (AR) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.411
# Average Recall (AR) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.430
# mAP=20.111%
# hash=526aac286ebb67218a3528397b4aecbff9269cbe01307069569345d9c3fbb445
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=SingleStream --test_mode=AccuracyOnly"
# ...
# Average Precision (AP) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.201
# Average Precision (AP) @[ IoU=0.50 | area= all | maxDets=100 ] = 0.381
# Average Precision (AP) @[ IoU=0.75 | area= all | maxDets=100 ] = 0.188
# Average Precision (AP) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.121
# Average Precision (AP) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.258
# Average Precision (AP) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.238
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 1 ] = 0.203
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 10 ] = 0.332
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.353
# Average Recall (AR) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.179
# Average Recall (AR) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.411
# Average Recall (AR) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.430
# mAP=20.111%
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-SingleStream:
# ssd-resnet34: Cannot find performance result. Maybe you are running in AccuracyOnly mode.
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-SingleStream:
# ssd-resnet34: Accuracy = 20.111, Threshold = 19.800. Accuracy test PASSED.
#
#
# real 12m55.238s
# user 8m38.988s
# sys 0m16.408s
# </pre>
# <a name="ssd-resnet34_multistream"></a>
# ## MultiStream
# ### [Config](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/configs/ssd-resnet34/MultiStream/config.json)
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ grep AGX_Xavier /datasets/inference_results_v1.0/closed/NVIDIA/configs/ssd-resnet34/MultiStream/config.json -A 9
# "<font color="#DC322F"><b>AGX_Xavier</b></font>": {
# "gpu_batch_size": 2,
# "gpu_multi_stream_samples_per_query": 2,
# "input_dtype": "int8",
# "input_format": "linear",
# "map_path": "data_maps/coco/val_map.txt",
# "precision": "int8",
# "tensor_path": "${PREPROCESSED_DATA_DIR}/coco/val2017/SSDResNet34/int8_linear",
# "use_graphs": false
# },
# </pre>
# <a name="ssd-resnet34_multistream_build"></a>
# ### Build
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make generate_engines RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=multistream"
# ...
# [2021-01-29 07:33:48,592 main.py:153 INFO] Finished building engines for ssd-resnet34 benchmark in MultiStream scenario.
# Time taken to generate engines: 263.8264467716217 seconds
#
# real 4m25.496s
# user 0m32.000s
# sys 0m35.784s
# </pre>
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ find . \
# -name ssd-resnet34-MultiStream*.plan -exec du -hs {} \;
# 148M ./build/engines/AGX_Xavier/ssd-resnet34/MultiStream/ssd-resnet34-MultiStream-gpu-b2-int8.default.plan
# </pre>
# <a name="ssd-resnet34_multistream_performance"></a>
# ### Performance
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/ssd-resnet34/MultiStream/performance/run_1/mlperf_log_summary.txt#L7)
#
# <pre>
# Samples per query : 2
# Result is : VALID
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=MultiStream --test_mode=PerformanceOnly"
# ...
# ================================================
# MLPerf Results Summary
# ================================================
# SUT name : LWIS_Server
# Scenario : Multi Stream
# Mode : Performance
# Samples per query : 2
# Result is : VALID
# Performance constraints satisfied : Yes
# Min duration satisfied : Yes
# Min queries satisfied : Yes
#
# ================================================
# Additional Stats
# ================================================
# Intervals between each IssueQuery: "qps" : 15, "ms" : 66.6667
# 50.00 percentile : 1
# 90.00 percentile : 1
# 95.00 percentile : 1
# 97.00 percentile : 1
# 99.00 percentile : 1
# 99.90 percentile : 1
#
# Per-query latency: "target_ns" : 66666666, "target_ms" : 66.6667
# 50.00 percentile latency (ns) : 55624812
# 90.00 percentile latency (ns) : 55913099
# 95.00 percentile latency (ns) : 56006924
# 97.00 percentile latency (ns) : 56067621
# 99.00 percentile latency (ns) : 56189283
# 99.90 percentile latency (ns) : 56401684
#
# Per-sample latency:
# Min latency (ns) : 55180175
# Max latency (ns) : 61310689
# Mean latency (ns) : 55646311
# 50.00 percentile latency (ns) : 55624812
# 90.00 percentile latency (ns) : 55913099
# 95.00 percentile latency (ns) : 56006924
# 97.00 percentile latency (ns) : 56067621
# 99.00 percentile latency (ns) : 56189283
# 99.90 percentile latency (ns) : 56401684
#
# ================================================
# Test Parameters Used
# ================================================
# samples_per_query : 2
# target_qps : 15
# target_latency (ns): 66666666
# max_async_queries : 1
# min_duration (ms): 60000
# max_duration (ms): 0
# min_query_count : 270336
# max_query_count : 0
# qsl_rng_seed : 12786827339337101903
# sample_index_rng_seed : 12640797754436136668
# schedule_rng_seed : 3135815929913719677
# accuracy_log_rng_seed : 0
# accuracy_log_probability : 0
# accuracy_log_sampling_target : 0
# print_timestamps : false
# performance_issue_unique : false
# performance_issue_same : false
# performance_issue_same_index : 0
# performance_sample_count : 64
#
# No warnings encountered during test.
#
# No errors encountered during test.
# Finished running actual test.
# Equivalent QPS computed by samples_per_query*target_qps : 30
# Device Device:0 processed:
# 270336 batches of size 2
# Memcpy Calls: 0
# PerSampleCudaMemcpy Calls: 0
# BatchedCudaMemcpy Calls: 270336
# &&&& PASSED Default_Harness # ./build/bin/harness_default
# [2021-01-29 13:12:12,503 main.py:341 INFO] Result: Samples per query : 2 and Result is : VALID
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-MultiStream:
# ssd-resnet34: Samples per query : 2 and Result is : VALID
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-MultiStream:
# ssd-resnet34: No accuracy results in PerformanceOnly mode.
#
#
# real 300m37.676s
# user 5m11.760s
# sys 0m27.856s
# </pre>
# <a name="ssd-resnet34_multistream_accuracy"></a>
# ### Accuracy
#
# #### [Submitted experiment](https://github.com/mlcommons/inference_results_v1.0/blob/master/closed/NVIDIA/results/AGX_Xavier_TRT/ssd-resnet34/MultiStream/accuracy/accuracy.txt)
#
# <pre>
# Average Precision (AP) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.201
# Average Precision (AP) @[ IoU=0.50 | area= all | maxDets=100 ] = 0.381
# Average Precision (AP) @[ IoU=0.75 | area= all | maxDets=100 ] = 0.188
# Average Precision (AP) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.121
# Average Precision (AP) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.258
# Average Precision (AP) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.238
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 1 ] = 0.203
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 10 ] = 0.332
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.353
# Average Recall (AR) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.179
# Average Recall (AR) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.411
# Average Recall (AR) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.430
# mAP=20.111%
# hash=859242388c9a94513b189eb58a55bd11ad1d2f7d094880dfb72157a7ac5e45fd
# </pre>
#
# #### Reproduced experiment
#
# <pre><font color="#859900"><b>anton@xavier</b></font>:<font color="#268BD2"><b>/datasets/inference_results_v1.0/closed/NVIDIA</b></font>$ time \
# MLPERF_SCRATCH_PATH=/datasets/mlperf_scratch_path \
# make run_harness RUN_ARGS="--benchmarks=ssd-resnet34 --scenarios=MultiStream --test_mode=AccuracyOnly"
# ...
# Average Precision (AP) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.201
# Average Precision (AP) @[ IoU=0.50 | area= all | maxDets=100 ] = 0.381
# Average Precision (AP) @[ IoU=0.75 | area= all | maxDets=100 ] = 0.188
# Average Precision (AP) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.121
# Average Precision (AP) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.258
# Average Precision (AP) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.238
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 1 ] = 0.203
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 10 ] = 0.332
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.353
# Average Recall (AR) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.179
# Average Recall (AR) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = 0.411
# Average Recall (AR) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = 0.430
# mAP=20.111%
#
# ======================= Perf harness results: =======================
#
# AGX_Xavier_TRT-default-MultiStream:
# ssd-resnet34: Cannot find performance result. Maybe you are running in AccuracyOnly mode.
#
#
# ======================= Accuracy results: =======================
#
# AGX_Xavier_TRT-default-MultiStream:
# ssd-resnet34: Accuracy = 20.111, Threshold = 19.800. Accuracy test PASSED.
#
#
# real 11m44.285s
# user 4m22.432s
# sys 0m17.144s
# </pre>
| jnotebook/mlperf-inference-v1.0-reproduce-xavier/reproduce-xavier-with-jetpack-4.5.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>Multiple Control Constructions</h2>
#
# [Watch Lecture](https://youtu.be/eoFJdS5BwkA)
# Remember that when appying CNOT gate, NOT operator is applied to the target qubit if the control qubit is in state $\ket{1}$:
#
# $$ CNOT= \mymatrix{cc|cc}{\blackbit{1} & 0 & 0 & 0 \\ 0 & \blackbit{1} & 0 & 0 \\ \hline 0 & 0 & 0 & \bluebit{1} \\ 0 & 0 & \bluebit{1} & 0} . $$
#
# How can we obtain the following operator, in which the NOT operator is applied to the target qubit if the control qubit is in state $ \ket{0} $?
#
# $$ C_0NOT = \mymatrix{cc|cc}{0 & \bluebit{1} & 0 & 0 \\ \bluebit{1} & 0 & 0 & 0 \\ \hline 0 & 0 & \blackbit{1} & 0 \\ 0 & 0 & 0 & \blackbit{1}} . $$
#
# As also mentioned in the notebook [Operators on Multiple Bits](B19_Operators_on_Multiple_Bits.ipynb), we can apply a $ NOT $ operator on the control bit before applying $ CNOT $ operator so that the $ NOT $ operator is applied to the target qubit when the control qubit has been in state $ \ket{0} $. To recover the previous value of the control qubit, we apply the $ NOT $ operator once more after the $ CNOT $ operator. In short:
# <ul>
# <li>apply $ NOT $ operator to the control qubit,</li>
# <li>apply $ CNOT $ operator, and,</li>
# <li>apply $ NOT $ operator to the control qubit.</li>
# </ul>
#
# We can implement this idea in Qiskit as follows.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
q = QuantumRegister(2, "q")
c = ClassicalRegister(2, "c")
qc = QuantumCircuit(q,c)
qc.x(q[1])
qc.cx(q[1],q[0])
# Returning control qubit to the initial state
qc.x(q[1])
job = execute(qc,Aer.get_backend('unitary_simulator'), shots = 1)
U=job.result().get_unitary(qc,decimals=3)
print("CNOT(0) = ")
for row in U:
s = ""
for value in row:
s = s + str(round(value.real,2)) + " "
print(s)
qc.draw(output="mpl", reverse_bits=True)
# -
# By using this trick, more complex conditional operators can be implemented.
# <h3>CCNOT</h3>
#
# Now we introduce $ CCNOT $ gate: **controlled-controlled-not operator** ([Toffoli gate](https://en.wikipedia.org/wiki/Toffoli_gate)), which is controlled by two qubits. The implementation of $CCNOT$ gate in Qiskit is as follows:
#
# circuit.ccx(control-qubit1,control-qubit2,target-qubit)
#
# That is, $ NOT $ operator is applied to the target qubit when both control qubits are in state $\ket{1}$. Its matrix representation is as follows:
#
# $$ CCNOT = \mymatrix{cc|cc|cc|cc}{\blackbit{1} & 0 & 0 & 0 & 0 & 0 & 0 & 0 \\ 0 & \blackbit{1} & 0 & 0 & 0 & 0 & 0 & 0 \\ \hline 0 & 0 & \blackbit{1} & 0 & 0 & 0 & 0 & 0 \\ 0 & 0 & 0 & \blackbit{1} & 0 & 0 & 0 & 0 \\ \hline 0 & 0 & 0 & 0 & \blackbit{1} & 0 & 0 & 0 \\ 0 & 0 & 0 & 0 & 0 & \blackbit{1} & 0 & 0 \\ \hline 0 & 0 & 0 & 0 & 0 & 0 & 0 & \bluebit{1} \\ 0 & 0 & 0 & 0 & 0 & 0 & \bluebit{1} & 0}. $$
# <a id="task1"></a>
# <h3> Task 1 </h3>
#
# Implement each of the following operators in Qiskit by using three qubits. Verify your implementation by using "unitary_simulator" backend. <br><br>
#
# <font size="-2">
# $$
# C_0C_0NOT = \mymatrix{cc|cc|cc|cc}{0 & \bluebit{1} & 0 & 0 & 0 & 0 & 0 & 0 \\ \bluebit{1} & 0 & 0 & 0 & 0 & 0 & 0 & 0 \\ \hline 0 & 0 & \blackbit{1} & 0 & 0 & 0 & 0 & 0 \\ 0 & 0 & 0 & \blackbit{1} & 0 & 0 & 0 & 0 \\ \hline 0 & 0 & 0 & 0 & \blackbit{1} & 0 & 0 & 0 \\ 0 & 0 & 0 & 0 & 0 & \blackbit{1} & 0 & 0 \\ \hline 0 & 0 & 0 & 0 & 0 & 0 & \blackbit{1} & 0 \\ 0 & 0 & 0 & 0 & 0 & 0 & 0 & \blackbit{1}}, ~~
# C_0C_1NOT = \mymatrix{cc|cc|cc|cc}{ \blackbit{1} & 0 & 0 & 0 & 0 & 0 & 0 & 0 \\ 0 & \blackbit{1} & 0 & 0 & 0 & 0 & 0 & 0 \\ \hline 0 & 0 & 0 & \bluebit{1} & 0 & 0 & 0 & 0 \\ 0 & 0 & \bluebit{1} & 0 & 0 & 0 & 0 & 0 \\ \hline 0 & 0 & 0 & 0 & \blackbit{1} & 0 & 0 & 0 \\ 0 & 0 & 0 & 0 & 0 & \blackbit{1} & 0 & 0 \\ \hline 0 & 0 & 0 & 0 & 0 & 0 & \blackbit{1} & 0 \\ 0 & 0 & 0 & 0 & 0 & 0 & 0 & \blackbit{1}}, ~~ \mbox{and} ~~
# C_1C_0NOT = \mymatrix{cc|cc|cc|cc}{\blackbit{1} & 0 & 0 & 0 & 0 & 0 & 0 & 0 \\ 0 & \blackbit{1} & 0 & 0 & 0 & 0 & 0 & 0 \\ \hline 0 & 0 & \blackbit{1} & 0 & 0 & 0 & 0 & 0 \\ 0 & 0 & 0 & \blackbit{1} & 0 & 0 & 0 & 0 \\ \hline 0 & 0 & 0 & 0 & 0 & \bluebit{1} & 0 & 0 \\ 0 & 0 & 0 & 0 & \bluebit{1} & 0 & 0 & 0 \\ \hline 0 & 0 & 0 & 0 & 0 & 0 & \blackbit{1} & 0 \\ 0 & 0 & 0 & 0 & 0 & 0 & 0 & \blackbit{1}}.
# $$
# </font>
# #### Implementation of $ C_0C_0NOT $ operator
# +
#
# your solution is here
#
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
q = QuantumRegister(3,"q")
c = ClassicalRegister(3,"c")
qc = QuantumCircuit(q,c)
qc.x(q[2])
qc.x(q[1])
qc.ccx(q[2],q[1],q[0])
qc.x(q[2])
qc.x(q[1])
job = execute(qc,Aer.get_backend('unitary_simulator'), shots = 1)
U=job.result().get_unitary(qc,decimals=3)
print("CCNOT(00) = ")
for row in U:
s = ""
for value in row:
s = s + str(round(value.real,2)) + " "
print(s)
qc.draw(output="mpl",reverse_bits=True)
# -
# #### Implementation of $ C_0C_1NOT $ operator
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
q = QuantumRegister(3,"q")
c = ClassicalRegister(3,"c")
qc = QuantumCircuit(q,c)
qc.x(q[2])
qc.ccx(q[2],q[1],q[0])
qc.x(q[2])
job = execute(qc,Aer.get_backend('unitary_simulator'), shots = 1)
U=job.result().get_unitary(qc,decimals=3)
print("CCNOT(01) = ")
for row in U:
s = ""
for value in row:
s = s + str(round(value.real,2)) + " "
print(s)
qc.draw(output="mpl",reverse_bits=True)
# -
# #### Implementation of $ C_1C_0NOT $ operator
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
q = QuantumRegister(3,"q")
c = ClassicalRegister(3,"c")
qc = QuantumCircuit(q,c)
qc.x(q[1])
qc.ccx(q[2],q[1],q[0])
qc.x(q[1])
job = execute(qc,Aer.get_backend('unitary_simulator'), shots = 1)
U=job.result().get_unitary(qc,decimals=3)
print("CCNOT(10) = ")
for row in U:
s = ""
for value in row:
s = s + str(round(value.real,2)) + " "
print(s)
qc.draw(output="mpl",reverse_bits=True)
# -
# <h3>More controls</h3>
#
# Here we present basic methods on how to implement $ NOT $ gates controlled by more than two qubits by using $CNOT$, $ CCNOT $, and some ancilla (auxiliary) qubits.
#
# *(Note that Qiskit has a method called "mct" to implement such gates. Another multiple-controlled operator in Qiskit is "mcrz".)*
# #### Implementation of CCCNOT gate
#
# We give the implementation of $ CCCNOT $ gate: $NOT$ operator is applied to target qubit when the control qubits are in state $ \ket{111} $. This gate requires 4 qubits. We also use an auxiliary qubit.
#
# Our qubits are $ q_{aux}, q_3, q_2, q_1, q_0 $, and the auxiliary qubit $q_{aux}$ should be in state $\ket{0}$ after each use. The implementation of the $ CCCNOT $ gate in Qiskit is given below. The short explanations are given as comments.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qaux = QuantumRegister(1,"qaux")
q = QuantumRegister(4,"q")
c = ClassicalRegister(4,"c")
qc = QuantumCircuit(q,qaux,c)
# step 1: set qaux to |1> if both q3 and q2 are in |1>
qc.ccx(q[3],q[2],qaux[0])
# step 2: apply NOT gate to q0 if both qaux and q1 are in |1>
qc.ccx(qaux[0],q[1],q[0])
# step 3: set qaux to |0> if both q3 and q2 are in |1> by reversing the affect of step 1
qc.ccx(q[3],q[2],qaux[0])
qc.draw(output="mpl",reverse_bits=True)
# -
# Now, we execute this circuit on every possible inputs and verify the correctness of the implementation experimentally.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
all_inputs=[]
for q3 in ['0','1']:
for q2 in ['0','1']:
for q1 in ['0','1']:
for q0 in ['0','1']:
all_inputs.append(q3+q2+q1+q0)
# print(all_inputs)
print("input --> output")
for the_input in all_inputs:
# create the circuit
qaux = QuantumRegister(1,"qaux")
q = QuantumRegister(4,"q")
c = ClassicalRegister(4,"c")
qc = QuantumCircuit(q,qaux,c)
# set the initial value of the circuit w.r.t. the input
if the_input[0] =='1': qc.x(q[3])
if the_input[1] =='1': qc.x(q[2])
if the_input[2] =='1': qc.x(q[1])
if the_input[3] =='1': qc.x(q[0])
# implement the CCNOT gates
qc.ccx(q[3],q[2],qaux[0])
qc.ccx(qaux[0],q[1],q[0])
qc.ccx(q[3],q[2],qaux[0])
# measure the main quantum register
qc.measure(q,c)
# execute the circuit
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=1)
counts = job.result().get_counts(qc)
for key in counts: the_output = key
printed_str = the_input[0:3]+" "+the_input[3]+" --> "+the_output[0:3]+" "+the_output[3]
if (the_input!=the_output): printed_str = printed_str + " the output is different than the input"
print(printed_str)
# -
# <h3>Task 2</h3>
#
# Provide an implementation of the NOT operator controlled by 4 qubits ($CCCCNOT$) in Qiskit. Verify its correctness by executing your solution on all possible inputs. (See the above example)
#
# *You may use two auxiliary qubits.*
# #### Desinging the operator
# +
#
# your solution is here
#
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qaux = QuantumRegister(2,"qaux")
q = QuantumRegister(5,"q")
c = ClassicalRegister(5,"c")
qc = QuantumCircuit(q,qaux,c)
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.ccx(qaux[1],qaux[0],q[0])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.draw(output="mpl",reverse_bits=True)
# -
# #### Executing this circuit on every possible inputs and verify the correctness of the implementation experimentially.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
all_inputs=[]
for q4 in ['0','1']:
for q3 in ['0','1']:
for q2 in ['0','1']:
for q1 in ['0','1']:
for q0 in ['0','1']:
all_inputs.append(q4+q3+q2+q1+q0)
#print(all_inputs)
print("input --> output")
for the_input in all_inputs:
# create the circuit
qaux = QuantumRegister(2,"qaux")
q = QuantumRegister(5,"q")
c = ClassicalRegister(5,"c")
qc = QuantumCircuit(q,qaux,c)
# set the initial value of the circuit w.r.t. the input
if the_input[0] =='1': qc.x(q[4])
if the_input[1] =='1': qc.x(q[3])
if the_input[2] =='1': qc.x(q[2])
if the_input[3] =='1': qc.x(q[1])
if the_input[4] =='1': qc.x(q[0])
# implement the CCNOT gates
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.ccx(qaux[1],qaux[0],q[0])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
# measure the main quantum register
qc.measure(q,c)
# execute the circuit
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=1)
counts = job.result().get_counts(qc)
for key in counts: the_output = key
printed_str = the_input[0:4]+" "+the_input[4]+" --> "+the_output[0:4]+" "+the_output[4]
if (the_input!=the_output): printed_str = printed_str + " the output is different than the input"
print(printed_str)
# -
# <h3>Task 3</h3>
#
# Repeat Task 2 for the operator $C_1C_0C_1C_0NOT$: $NOT$ operator is applied to the target qubit if the four control qubits are in state $ \ket{1010} $.
# #### Desinging the operator
# +
#
# your solution is here
#
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qaux = QuantumRegister(2,"qaux")
q = QuantumRegister(5,"q")
c = ClassicalRegister(5,"c")
qc = QuantumCircuit(q,qaux,c)
qc.x(q[3])
qc.x(q[1])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.ccx(qaux[1],qaux[0],q[0])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.x(q[3])
qc.x(q[1])
qc.draw(output="mpl",reverse_bits=True)
# -
# #### Executing this circuit on every possible inputs and verify the correctness of the implementation experimentially.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
all_inputs=[]
for q4 in ['0','1']:
for q3 in ['0','1']:
for q2 in ['0','1']:
for q1 in ['0','1']:
for q0 in ['0','1']:
all_inputs.append(q4+q3+q2+q1+q0)
#print(all_inputs)
print("input --> output")
for the_input in all_inputs:
# create the circuit
qaux = QuantumRegister(2,"qaux")
q = QuantumRegister(5,"q")
c = ClassicalRegister(5,"c")
qc = QuantumCircuit(q,qaux,c)
# set the initial value of the circuit w.r.t. the input
if the_input[0] =='1': qc.x(q[4])
if the_input[1] =='1': qc.x(q[3])
if the_input[2] =='1': qc.x(q[2])
if the_input[3] =='1': qc.x(q[1])
if the_input[4] =='1': qc.x(q[0])
# implement the CCNOT gates
qc.x(q[3])
qc.x(q[1])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.ccx(qaux[1],qaux[0],q[0])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.x(q[3])
qc.x(q[1])
# measure the main quantum register
qc.measure(q,c)
# execute the circuit
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=1)
counts = job.result().get_counts(qc)
for key in counts: the_output = key
printed_str = the_input[0:4]+" "+the_input[4]+" --> "+the_output[0:4]+" "+the_output[4]
if (the_input!=the_output): printed_str = printed_str + " the output is different than the input"
print(printed_str)
# -
# <h3>Task 4 (extra)</h3>
#
# Write a function taking a binary string "$ b_1 b_2 b_3 b_4$ that repeats Task 2 for the operator $ C_{b_1}C_{b_2}C_{b_3}C_{b_4}NOT $ gate, where $ b_1,\ldots,b_4$ are bits and $ NOT $ operator is applied to target qubit if the control qubits are in state $ \ket{b_1b_2b_3b_4} $.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
all_inputs=[]
for q4 in ['0','1']:
for q3 in ['0','1']:
for q2 in ['0','1']:
for q1 in ['0','1']:
for q0 in ['0','1']:
all_inputs.append(q4+q3+q2+q1+q0)
#print(all_inputs)
def c4not(control_state='1111'):
#
# drawing the circuit
#
print("Control state is",control_state)
print("Drawing the circuit:")
qaux = QuantumRegister(2,"qaux")
q = QuantumRegister(5,"q")
c = ClassicalRegister(5,"c")
qc = QuantumCircuit(q,qaux,c)
for b in range(4):
if control_state[b] == '0':
qc.x(q[4-b])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.ccx(qaux[1],qaux[0],q[0])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
for b in range(4):
if control_state[b] == '0':
qc.x(q[4-b])
display(qc.draw(output="mpl",reverse_bits=True))
#
# executing the operator on all possible inputs
#
print("Control state is",control_state)
print("input --> output")
for the_input in all_inputs:
# create the circuit
qaux = QuantumRegister(2,"qaux")
q = QuantumRegister(5,"q")
c = ClassicalRegister(5,"c")
qc = QuantumCircuit(q,qaux,c)
# set the initial value of the circuit w.r.t. the input
if the_input[0] =='1': qc.x(q[4])
if the_input[1] =='1': qc.x(q[3])
if the_input[2] =='1': qc.x(q[2])
if the_input[3] =='1': qc.x(q[1])
if the_input[4] =='1': qc.x(q[0])
# implement the CCNOT gates
for b in range(4):
if control_state[b] == '0':
qc.x(q[4-b])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
qc.ccx(qaux[1],qaux[0],q[0])
qc.ccx(q[4],q[3],qaux[1])
qc.ccx(q[2],q[1],qaux[0])
for b in range(4):
if control_state[b] == '0':
qc.x(q[4-b])
# measure the main quantum register
qc.measure(q,c)
# execute the circuit
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=1)
counts = job.result().get_counts(qc)
for key in counts: the_output = key
printed_str = the_input[0:4]+" "+the_input[4]+" --> "+the_output[0:4]+" "+the_output[4]
if (the_input!=the_output): printed_str = printed_str + " the output is different than the input"
print(printed_str)
# +
# try different values
#c4not()
#c4not('1001')
c4not('0011')
#c4not('1101')
#c4not('0000')
| QWorld's Global Quantum Programming Workshop/DAY 5 + DAY 6/27.Multiple_Control_Constructions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/NicolasPinell/Backtesting/blob/master/convolutional_networks.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="DDJwQPZcupab"
# # EECS 498-007/598-005 Assignment 3-2: Convolutional Neural Networks and Batch Normalization
#
# Before we start, please put your name and UMID in following format
#
# : Firstname LASTNAME, #00000000 // e.g.) <NAME>, #12345678
# + [markdown] colab_type="text" id="2KMxqLt1h2kx"
# **Your Answer:**
# Hello WORLD, #XXXXXXXX
# + [markdown] colab_type="text" id="aQW_w1Wzw72f" tags=["pdf-title"]
# # Convolutional networks
# So far we have worked with deep fully-connected networks, using them to explore different optimization strategies and network architectures. Fully-connected networks are a good testbed for experimentation because they are very computationally efficient, but in practice all state-of-the-art results use convolutional networks instead.
#
# First you will implement several layer types that are used in convolutional networks. You will then use these layers to train a convolutional network on the CIFAR-10 dataset.
# + [markdown] colab_type="text" id="ubB_0e-UAOVK"
# ## Install starter code
# We will continue using the utility functions that we've used for Assignment 1 and 2: [`coutils` package](https://github.com/deepvision-class/starter-code). Run this cell to download and install it.
#
# + colab_type="code" id="ASkY27ZtA7Is" colab={}
# !pip install git+https://github.com/deepvision-class/starter-code
# + [markdown] colab_type="text" id="MzqbYcKdz6ew"
# ## Setup code
# Run some setup code for this notebook: Import some useful packages and increase the default figure size.
# + colab_type="code" id="vixlKb-ew72h" tags=["pdf-ignore"] colab={}
import math
import torch
import coutils
from coutils import extract_drive_file_id, register_colab_notebooks, \
fix_random_seed, rel_error, compute_numeric_gradient, Solver
import matplotlib.pyplot as plt
import time
# for plotting
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# data type and device for torch.tensor
to_float = {'dtype': torch.float, 'device': 'cpu'}
to_float_cuda = {'dtype': torch.float, 'device': 'cuda'}
to_double = {'dtype': torch.double, 'device': 'cpu'}
to_double_cuda = {'dtype': torch.double, 'device': 'cuda'}
to_long = {'dtype': torch.long, 'device': 'cpu'}
to_long_cuda = {'dtype': torch.long, 'device': 'cuda'}
# + [markdown] colab_type="text" id="0Saw9jGNm-9-"
# ## Import functions from previous notebook
# This notebook will re-use some pieces of code that you implemented in the previous notebook.
#
# In order to do this, you will need the **Google Drive file ID** of your completed notebook `fully_connected_networks.ipynb`. You can find the this file ID by doing the following:
# 1. Make sure you have saved your completed `fully_connected_networks.ipynb` notebook to your own Google Drive
# 2. Open you finished `fully_connected_networks.ipynb` notebook in Colab.
# 3. Click the "Share" button at the top of the screen
# 4. Copy the "Notebook link" and paste it in the following cell, assigning it to the `FULLY_CONNECTED_NOTEBOOK_LINK` variable
#
# **Important:** If you modify the implementations of any functions in your `fully_connected_networks.ipynb` notebook, **they will not automatically be propagated to this notebook**. For changes to `fully_connected_networks.ipynb` to be propagated to this notebook, you will need to:
# 1. Make sure that you save your modified `fully_connected_networks.ipynb` notebook (File > Save)
# 2. Restart the runtime of this notebook (Runtime > Restart Runtime)
# 3. Rerun all cells in this notebook (in particular the import cell below)
# + colab_type="code" id="Xq5yHDeuklId" colab={}
FULLY_CONNECTED_NOTEBOOK_LINK = ""
fcn_id = extract_drive_file_id(FULLY_CONNECTED_NOTEBOOK_LINK)
print('Google Drive file id: "%s"' % fcn_id)
register_colab_notebooks({'fully_connected_networks': fcn_id})
from fully_connected_networks import get_CIFAR10_data
from fully_connected_networks import Linear, ReLU, Linear_ReLU, Dropout
from fully_connected_networks import svm_loss, softmax_loss
from fully_connected_networks import sgd_momentum, rmsprop, adam
print('Import successful!')
# + [markdown] colab_type="text" id="HpqGVD5mnjYF"
# ## Load CIFAR-10 data
#
# Here we load up our favorite CIFAR-10 data so we can use it to train a classifier on a real dataset.
# + colab_type="code" id="rEjt-AJRG5dp" colab={}
# Invoke the above function to get our data.
data_dict = get_CIFAR10_data(visualize=True)
print('Train data shape: ', data_dict['X_train'].shape)
print('Train labels shape: ', data_dict['y_train'].shape)
print('Validation data shape: ', data_dict['X_val'].shape)
print('Validation labels shape: ', data_dict['y_val'].shape)
print('Test data shape: ', data_dict['X_test'].shape)
print('Test labels shape: ', data_dict['y_test'].shape)
# + [markdown] colab_type="text" id="CJInAlccoI5e"
# # Convolutional layer
# As in the previous notebook, we will package each new neural network operator in a class that defines a `forward` and `backward` function.
# + colab_type="code" id="N8ZohWR-o_uc" colab={}
class Conv(object):
@staticmethod
def forward(x, w, b, conv_param):
raise NotImplementedError
@staticmethod
def backward(dout, cache):
raise NotImplementedError
# + [markdown] colab_type="text" id="x07DS91iw72o"
# ## Convolutional layer: forward
# The core of a convolutional network is the convolution operation. Implement the forward pass for the convolution layer in the function `Conv.forward`.
#
# You don't have to worry too much about efficiency at this point; just write the code in whatever way you find most clear.
#
# You can test your implementation by running the following:
# + colab_type="code" id="48TBbwG0n7WK" colab={}
def conv_forward(x, w, b, conv_param):
"""
A naive implementation of the forward pass for a convolutional layer.
The input consists of N data points, each with C channels, height H and
width W. We convolve each input with F different filters, where each filter
spans all C channels and has height HH and width WW.
Input:
- x: Input data of shape (N, C, H, W)
- w: Filter weights of shape (F, C, HH, WW)
- b: Biases, of shape (F,)
- conv_param: A dictionary with the following keys:
- 'stride': The number of pixels between adjacent receptive fields in the
horizontal and vertical directions.
- 'pad': The number of pixels that will be used to zero-pad the input.
During padding, 'pad' zeros should be placed symmetrically (i.e equally on both sides)
along the height and width axes of the input. Be careful not to modfiy the original
input x directly.
Returns a tuple of:
- out: Output data, of shape (N, F, H', W') where H' and W' are given by
H' = 1 + (H + 2 * pad - HH) / stride
W' = 1 + (W + 2 * pad - WW) / stride
- cache: (x, w, b, conv_param)
"""
out = None
##############################################################################
# TODO: Implement the convolutional forward pass. #
# Hint: you can use the function torch.nn.functional.pad for padding. #
# Note that you are NOT allowed to use anything in torch.nn in other places. #
##############################################################################
# Replace "pass" statement with your code
pass
#############################################################################
# END OF YOUR CODE #
#############################################################################
cache = (x, w, b, conv_param)
return out, cache
Conv.forward = conv_forward
# + [markdown] colab_type="text" id="VeK1fneVy0N3"
# After implementing the forward pass of the convolution operation, run the following to check your implementation. You should get a relative error less than `1e-7`.
# + colab_type="code" id="F5R_WY1Iw72p" colab={}
x_shape = torch.tensor((2, 3, 4, 4))
w_shape = torch.tensor((3, 3, 4, 4))
x = torch.linspace(-0.1, 0.5, steps=torch.prod(x_shape), **to_double_cuda).reshape(*x_shape)
w = torch.linspace(-0.2, 0.3, steps=torch.prod(w_shape), **to_double_cuda).reshape(*w_shape)
b = torch.linspace(-0.1, 0.2, steps=3, **to_double_cuda)
conv_param = {'stride': 2, 'pad': 1}
out, _ = Conv.forward(x, w, b, conv_param)
correct_out = torch.tensor([[[[-0.08759809, -0.10987781],
[-0.18387192, -0.2109216 ]],
[[ 0.21027089, 0.21661097],
[ 0.22847626, 0.23004637]],
[[ 0.50813986, 0.54309974],
[ 0.64082444, 0.67101435]]],
[[[-0.98053589, -1.03143541],
[-1.19128892, -1.24695841]],
[[ 0.69108355, 0.66880383],
[ 0.59480972, 0.56776003]],
[[ 2.36270298, 2.36904306],
[ 2.38090835, 2.38247847]]]], **to_double_cuda)
# Compare your output to ours; difference should be around e-8
print('Testing Conv.forward')
print('difference: ', rel_error(out, correct_out))
# + [markdown] colab_type="text" id="N5bKrl7Uw72t"
# ## Aside: Image processing via convolutions
#
# As fun way to both check your implementation and gain a better understanding of the type of operation that convolutional layers can perform, we will set up an input containing two images and manually set up filters that perform common image processing operations (grayscale conversion and edge detection). The convolution forward pass will apply these operations to each of the input images. We can then visualize the results as a sanity check.
# + colab_type="code" id="k8BffZxdw72u" tags=["pdf-ignore-input"] colab={}
from imageio import imread
from PIL import Image
from torchvision.transforms import ToTensor
kitten_url = 'https://web.eecs.umich.edu/~justincj/teaching/eecs498/assets/a3/kitten.jpg'
puppy_url = 'https://web.eecs.umich.edu/~justincj/teaching/eecs498/assets/a3/puppy.jpg'
kitten = imread(kitten_url)
puppy = imread(puppy_url)
# kitten is wide, and puppy is already square
d = kitten.shape[1] - kitten.shape[0]
kitten_cropped = kitten[:, d//2:-d//2, :]
img_size = 200 # Make this smaller if it runs too slow
resized_puppy = ToTensor()(Image.fromarray(puppy).resize((img_size, img_size)))
resized_kitten = ToTensor()(Image.fromarray(kitten_cropped).resize((img_size, img_size)))
x = torch.stack([resized_puppy, resized_kitten])
# Set up a convolutional weights holding 2 filters, each 3x3
w = torch.zeros(2, 3, 3, 3, dtype=x.dtype)
# The first filter converts the image to grayscale.
# Set up the red, green, and blue channels of the filter.
w[0, 0, :, :] = torch.tensor([[0, 0, 0], [0, 0.3, 0], [0, 0, 0]])
w[0, 1, :, :] = torch.tensor([[0, 0, 0], [0, 0.6, 0], [0, 0, 0]])
w[0, 2, :, :] = torch.tensor([[0, 0, 0], [0, 0.1, 0], [0, 0, 0]])
# Second filter detects horizontal edges in the blue channel.
w[1, 2, :, :] = torch.tensor([[1, 2, 1], [0, 0, 0], [-1, -2, -1]])
# Vector of biases. We don't need any bias for the grayscale
# filter, but for the edge detection filter we want to add 128
# to each output so that nothing is negative.
b = torch.tensor([0, 128], dtype=x.dtype)
# Compute the result of convolving each input in x with each filter in w,
# offsetting by b, and storing the results in out.
out, _ = Conv.forward(x, w, b, {'stride': 1, 'pad': 1})
def imshow_no_ax(img, normalize=True):
""" Tiny helper to show images as uint8 and remove axis labels """
if normalize:
img_max, img_min = img.max(), img.min()
img = 255.0 * (img - img_min) / (img_max - img_min)
plt.imshow(img)
plt.gca().axis('off')
# Show the original images and the results of the conv operation
plt.subplot(2, 3, 1)
imshow_no_ax(puppy, normalize=False)
plt.title('Original image')
plt.subplot(2, 3, 2)
imshow_no_ax(out[0, 0])
plt.title('Grayscale')
plt.subplot(2, 3, 3)
imshow_no_ax(out[0, 1])
plt.title('Edges')
plt.subplot(2, 3, 4)
imshow_no_ax(kitten_cropped, normalize=False)
plt.subplot(2, 3, 5)
imshow_no_ax(out[1, 0])
plt.subplot(2, 3, 6)
imshow_no_ax(out[1, 1])
plt.show()
# + [markdown] colab_type="text" id="Lqsg-NxHw72y"
# ## Convolutional layer: backward
# Implement the backward pass for the convolution operation in the function `Conv.backward`. Again, you don't need to worry too much about computational efficiency.
# + colab_type="code" id="sPE64kjCqtEz" colab={}
def conv_backward(dout, cache):
"""
A naive implementation of the backward pass for a convolutional layer.
Inputs:
- dout: Upstream derivatives.
- cache: A tuple of (x, w, b, conv_param) as in conv_forward_naive
Returns a tuple of:
- dx: Gradient with respect to x
- dw: Gradient with respect to w
- db: Gradient with respect to b
"""
dx, dw, db = None, None, None
#############################################################################
# TODO: Implement the convolutional backward pass. #
#############################################################################
# Replace "pass" statement with your code
pass
#############################################################################
# END OF YOUR CODE #
#############################################################################
return dx, dw, db
Conv.backward = conv_backward
# + [markdown] colab_type="text" id="fgj8WovMzYtJ"
# After implementing the convolution backward pass, run the following to test your implementation. You should get errors less than `1e-8`.
# + colab_type="code" id="77NxvIGpw720" colab={}
fix_random_seed(0)
x = torch.randn(4, 3, 5, 5, **to_double_cuda)
w = torch.randn(2, 3, 3, 3, **to_double_cuda)
b = torch.randn(2, **to_double_cuda)
dout = torch.randn(4, 2, 5, 5, **to_double_cuda)
conv_param = {'stride': 1, 'pad': 1}
dx_num = compute_numeric_gradient(lambda x: Conv.forward(x, w, b, conv_param)[0], x, dout)
dw_num = compute_numeric_gradient(lambda w: Conv.forward(x, w, b, conv_param)[0], w, dout)
db_num = compute_numeric_gradient(lambda b: Conv.forward(x, w, b, conv_param)[0], b, dout)
out, cache = Conv.forward(x, w, b, conv_param)
dx, dw, db = Conv.backward(dout, cache)
print('Testing Conv.backward function')
print('dx error: ', rel_error(dx, dx_num))
print('dw error: ', rel_error(dw, dw_num))
print('db error: ', rel_error(db, db_num))
# + [markdown] colab_type="text" id="CS8EsPacrpG8"
# # Max-pooling
# + colab_type="code" id="anBWiAVVr24F" colab={}
class MaxPool(object):
@staticmethod
def forward(x, pool_param):
raise NotImplementedError
@staticmethod
def backward(dout, cache):
raise NotImplementedError
# + [markdown] colab_type="text" id="elO_ys-8w723"
# ## Max-pooling: forward
# Implement the forward pass for the max-pooling operation. Again, don't worry too much about computational efficiency.
# + colab_type="code" id="0z5_oQ2PrWCZ" colab={}
def max_pool_forward(x, pool_param):
"""
A naive implementation of the forward pass for a max-pooling layer.
Inputs:
- x: Input data, of shape (N, C, H, W)
- pool_param: dictionary with the following keys:
- 'pool_height': The height of each pooling region
- 'pool_width': The width of each pooling region
- 'stride': The distance between adjacent pooling regions
No padding is necessary here.
Returns a tuple of:
- out: Output data, of shape (N, C, H', W') where H' and W' are given by
H' = 1 + (H - pool_height) / stride
W' = 1 + (W - pool_width) / stride
- cache: (x, pool_param)
"""
out = None
#############################################################################
# TODO: Implement the max-pooling forward pass #
#############################################################################
# Replace "pass" statement with your code
pass
#############################################################################
# END OF YOUR CODE #
#############################################################################
cache = (x, pool_param)
return out, cache
MaxPool.forward = max_pool_forward
# + [markdown] colab_type="text" id="PeKxeFUN0L8a"
# After implementing the forward pass for max-pooling, run the following to check your implementation. You should get errors less than `1e-7`.
# + colab_type="code" id="qmNJY6E7w724" colab={}
x_shape = torch.tensor((2, 3, 4, 4))
x = torch.linspace(-0.3, 0.4, steps=torch.prod(x_shape), **to_double_cuda).reshape(*x_shape)
pool_param = {'pool_width': 2, 'pool_height': 2, 'stride': 2}
out, _ = MaxPool.forward(x, pool_param)
correct_out = torch.tensor([[[[-0.26315789, -0.24842105],
[-0.20421053, -0.18947368]],
[[-0.14526316, -0.13052632],
[-0.08631579, -0.07157895]],
[[-0.02736842, -0.01263158],
[ 0.03157895, 0.04631579]]],
[[[ 0.09052632, 0.10526316],
[ 0.14947368, 0.16421053]],
[[ 0.20842105, 0.22315789],
[ 0.26736842, 0.28210526]],
[[ 0.32631579, 0.34105263],
[ 0.38526316, 0.4 ]]]], **to_double_cuda)
# Compare your output with ours. Difference should be on the order of e-8.
print('Testing MaxPool.forward function:')
print('difference: ', rel_error(out, correct_out))
# + [markdown] colab_type="text" id="lLjMhG4nw728"
# ## Max-pooling: backward
# Implement the backward pass for the max-pooling operation. You don't need to worry about computational efficiency.
# + colab_type="code" id="retdcruisTj7" colab={}
def max_pool_backward(dout, cache):
"""
A naive implementation of the backward pass for a max-pooling layer.
Inputs:
- dout: Upstream derivatives
- cache: A tuple of (x, pool_param) as in the forward pass.
Returns:
- dx: Gradient with respect to x
"""
dx = None
#############################################################################
# TODO: Implement the max-pooling backward pass #
#############################################################################
# Replace "pass" statement with your code
pass
#############################################################################
# END OF YOUR CODE #
#############################################################################
return dx
MaxPool.backward = max_pool_backward
# + [markdown] colab_type="text" id="48x6OP5J0WjS"
# Check your implementation of the max pooling backward pass with numeric gradient checking by running the following. You should get errors less than `1e-10`.
# + colab_type="code" id="9FTLuSsIw729" colab={}
fix_random_seed(0)
x = torch.randn(3, 2, 8, 8, **to_double_cuda)
dout = torch.randn(3, 2, 4, 4, **to_double_cuda)
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
dx_num = compute_numeric_gradient(lambda x: MaxPool.forward(x, pool_param)[0], x, dout)
out, cache = MaxPool.forward(x, pool_param)
dx = MaxPool.backward(dout, cache)
print('Testing MaxPool.backward function:')
print('dx error: ', rel_error(dx, dx_num))
# + [markdown] colab_type="text" id="0-GA5mHcw73A"
# # Fast layers
# Making convolution and pooling layers fast can be challenging. To spare you the pain, we've provided fast implementations of the forward and backward passes for convolution and pooling layers.
#
# The fast convolution implementation depends on `torch.nn`
#
# The API for the fast versions of the convolution and pooling layers is exactly the same as the naive versions that you implemented above: the forward pass receives data, weights, and parameters and produces outputs and a cache object; the backward pass recieves upstream derivatives and the cache object and produces gradients with respect to the data and weights.
#
# You can compare the performance of the naive and fast versions of these layers by running the following:
# + colab_type="code" id="KONf6gRfs7sl" colab={}
class FastConv(object):
@staticmethod
def forward(x, w, b, conv_param):
N, C, H, W = x.shape
F, _, HH, WW = w.shape
stride, pad = conv_param['stride'], conv_param['pad']
layer = torch.nn.Conv2d(C, F, (HH, WW), stride=stride, padding=pad)
layer.weight = torch.nn.Parameter(w)
layer.bias = torch.nn.Parameter(b)
tx = x.detach()
tx.requires_grad = True
out = layer(tx)
cache = (x, w, b, conv_param, tx, out, layer)
return out, cache
@staticmethod
def backward(dout, cache):
try:
x, _, _, _, tx, out, layer = cache
out.backward(dout)
dx = tx.grad.detach()
dw = layer.weight.grad.detach()
db = layer.bias.grad.detach()
layer.weight.grad = layer.bias.grad = None
except RuntimeError:
dx, dw, db = torch.zeros_like(tx), torch.zeros_like(layer.weight), torch.zeros_like(layer.bias)
return dx, dw, db
class FastMaxPool(object):
@staticmethod
def forward(x, pool_param):
N, C, H, W = x.shape
pool_height, pool_width = pool_param['pool_height'], pool_param['pool_width']
stride = pool_param['stride']
layer = torch.nn.MaxPool2d(kernel_size=(pool_height, pool_width), stride=stride)
tx = x.detach()
tx.requires_grad = True
out = layer(tx)
cache = (x, pool_param, tx, out, layer)
return out, cache
@staticmethod
def backward(dout, cache):
try:
x, _, tx, out, layer = cache
out.backward(dout)
dx = tx.grad.detach()
except RuntimeError:
dx = torch.zeros_like(tx)
return dx
# + [markdown] colab_type="text" id="Z7B_QYt206g4"
# We will now compare three different implementations of convolution (both forward and backward):
#
# 1. Your naive, non-vectorized implementation on CPU
# 2. The fast, vectorized implementation on CPU
# 3. The fast, vectorized implementation on GPU
#
# The differences between your implementation and FastConv should be less than `1e-10`. When moving from your implementation to FastConv CPU, you will likely see speedups of at least 100x. When comparing your implementation to FastConv CUDA, you will likely see speedups of more than 500x. (These speedups are not hard requirements for this assignment since we are not asking you to write any vectorized implementations)
# + colab_type="code" id="mSbc5Ttvw73C" colab={}
# Rel errors should be around e-11 or less
fix_random_seed(0)
x = torch.randn(10, 3, 31, 31, **to_double)
w = torch.randn(25, 3, 3, 3, **to_double)
b = torch.randn(25, **to_double)
dout = torch.randn(10, 25, 16, 16, **to_double)
x_cuda, w_cuda, b_cuda, dout_cuda = x.to('cuda'), w.to('cuda'), b.to('cuda'), dout.to('cuda')
conv_param = {'stride': 2, 'pad': 1}
t0 = time.time()
out_naive, cache_naive = Conv.forward(x, w, b, conv_param)
t1 = time.time()
out_fast, cache_fast = FastConv.forward(x, w, b, conv_param)
t2 = time.time()
out_fast_cuda, cache_fast_cuda = FastConv.forward(x_cuda, w_cuda, b_cuda, conv_param)
t3 = time.time()
print('Testing FastConv.forward:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Fast CUDA: %fs' % (t3 - t2))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('Speedup CUDA: %fx' % ((t1 - t0) / (t3 - t2)))
print('Difference: ', rel_error(out_naive, out_fast))
print('Difference CUDA: ', rel_error(out_naive, out_fast_cuda.to(out_naive.device)))
t0 = time.time()
dx_naive, dw_naive, db_naive = Conv.backward(dout, cache_naive)
t1 = time.time()
dx_fast, dw_fast, db_fast = FastConv.backward(dout, cache_fast)
t2 = time.time()
dx_fast_cuda, dw_fast_cuda, db_fast_cuda = FastConv.backward(dout_cuda, cache_fast_cuda)
t3 = time.time()
print('\nTesting FastConv.backward:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Fast CUDA: %fs' % (t3 - t2))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('Speedup CUDA: %fx' % ((t1 - t0) / (t3 - t2)))
print('dx difference: ', rel_error(dx_naive, dx_fast))
print('dw difference: ', rel_error(dw_naive, dw_fast))
print('db difference: ', rel_error(db_naive, db_fast))
print('dx difference CUDA: ', rel_error(dx_naive, dx_fast_cuda.to(dx_naive.device)))
print('dw difference CUDA: ', rel_error(dw_naive, dw_fast_cuda.to(dw_naive.device)))
print('db difference CUDA: ', rel_error(db_naive, db_fast_cuda.to(db_naive.device)))
# + [markdown] colab_type="text" id="HdvR1krO2P_q"
# We will now similarly compare your naive implementation of max pooling against the fast implementation. You should see differences of 0 between your implementation and the fast implementation.
#
# When comparing your implementation against FastMaxPool on CPU, you will likely see speedups of more than 100x. When comparing your implementation against FastMaxPool on GPU, you will likely see speedups of more than 500x.
# + colab_type="code" id="R0fykwCiw73F" colab={}
# Relative errors should be close to 0.0
fix_random_seed(0)
x = torch.randn(40, 3, 32, 32, **to_double)
dout = torch.randn(40, 3, 16, 16, **to_double)
x_cuda, dout_cuda = x.to('cuda'), dout.to('cuda')
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
t0 = time.time()
out_naive, cache_naive = MaxPool.forward(x, pool_param)
t1 = time.time()
out_fast, cache_fast = FastMaxPool.forward(x, pool_param)
t2 = time.time()
out_fast_cuda, cache_fast_cuda = FastMaxPool.forward(x_cuda, pool_param)
t3 = time.time()
print('Testing FastMaxPool.forward:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Fast CUDA: %fs' % (t3 - t2))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('Speedup CUDA: %fx' % ((t1 - t0) / (t3 - t2)))
print('Difference: ', rel_error(out_naive, out_fast))
print('Difference CUDA: ', rel_error(out_naive, out_fast_cuda.to(out_naive.device)))
t0 = time.time()
dx_naive = MaxPool.backward(dout, cache_naive)
t1 = time.time()
dx_fast = FastMaxPool.backward(dout, cache_fast)
t2 = time.time()
dx_fast_cuda = FastMaxPool.backward(dout_cuda, cache_fast_cuda)
t3 = time.time()
print('\nTesting FastMaxPool.backward:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Fast CUDA: %fs' % (t3 - t2))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('Speedup CUDA: %fx' % ((t1 - t0) / (t3 - t2)))
print('dx difference: ', rel_error(dx_naive, dx_fast))
print('dx difference CUDA: ', rel_error(dx_naive, dx_fast_cuda.to(dx_naive.device)))
# + [markdown] colab_type="text" id="Ix9ih6sKw73I"
# # Convolutional "sandwich" layers
# Previously we introduced the concept of "sandwich" layers that combine multiple operations into commonly used patterns. Below you will find sandwich layers that implement a few commonly used patterns for convolutional networks. Run the cells below to sanity check they're working.
# + colab_type="code" id="bx5o2y4Jxx2I" colab={}
class Conv_ReLU(object):
@staticmethod
def forward(x, w, b, conv_param):
"""
A convenience layer that performs a convolution followed by a ReLU.
Inputs:
- x: Input to the convolutional layer
- w, b, conv_param: Weights and parameters for the convolutional layer
Returns a tuple of:
- out: Output from the ReLU
- cache: Object to give to the backward pass
"""
a, conv_cache = FastConv.forward(x, w, b, conv_param)
out, relu_cache = ReLU.forward(a)
cache = (conv_cache, relu_cache)
return out, cache
@staticmethod
def backward(dout, cache):
"""
Backward pass for the conv-relu convenience layer.
"""
conv_cache, relu_cache = cache
da = ReLU.backward(dout, relu_cache)
dx, dw, db = FastConv.backward(da, conv_cache)
return dx, dw, db
class Conv_ReLU_Pool(object):
@staticmethod
def forward(x, w, b, conv_param, pool_param):
"""
A convenience layer that performs a convolution, a ReLU, and a pool.
Inputs:
- x: Input to the convolutional layer
- w, b, conv_param: Weights and parameters for the convolutional layer
- pool_param: Parameters for the pooling layer
Returns a tuple of:
- out: Output from the pooling layer
- cache: Object to give to the backward pass
"""
a, conv_cache = FastConv.forward(x, w, b, conv_param)
s, relu_cache = ReLU.forward(a)
out, pool_cache = FastMaxPool.forward(s, pool_param)
cache = (conv_cache, relu_cache, pool_cache)
return out, cache
@staticmethod
def backward(dout, cache):
"""
Backward pass for the conv-relu-pool convenience layer
"""
conv_cache, relu_cache, pool_cache = cache
ds = FastMaxPool.backward(dout, pool_cache)
da = ReLU.backward(ds, relu_cache)
dx, dw, db = FastConv.backward(da, conv_cache)
return dx, dw, db
# + [markdown] colab_type="text" id="JsWygDZi2tFy"
# Test the implementations of the sandwich layers by running the following. You should see errors less than `1e-7`.
# + colab_type="code" id="B-0DpEY8w73J" colab={}
fix_random_seed(0)
x = torch.randn(2, 3, 16, 16, **to_double_cuda)
w = torch.randn(3, 3, 3, 3, **to_double_cuda)
b = torch.randn(3, **to_double_cuda)
dout = torch.randn(2, 3, 8, 8, **to_double_cuda)
conv_param = {'stride': 1, 'pad': 1}
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
out, cache = Conv_ReLU_Pool.forward(x, w, b, conv_param, pool_param)
dx, dw, db = Conv_ReLU_Pool.backward(dout, cache)
dx_num = compute_numeric_gradient(lambda x: Conv_ReLU_Pool.forward(x, w, b, conv_param, pool_param)[0], x, dout)
dw_num = compute_numeric_gradient(lambda w: Conv_ReLU_Pool.forward(x, w, b, conv_param, pool_param)[0], w, dout)
db_num = compute_numeric_gradient(lambda b: Conv_ReLU_Pool.forward(x, w, b, conv_param, pool_param)[0], b, dout)
# Relative errors should be around e-8 or less
print('Testing Conv_ReLU_Pool')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
# + colab_type="code" id="WjtyDWxPw73M" colab={}
fix_random_seed(0)
x = torch.randn(2, 3, 8, 8, **to_double_cuda)
w = torch.randn(3, 3, 3, 3, **to_double_cuda)
b = torch.randn(3, **to_double_cuda)
dout = torch.randn(2, 3, 8, 8, **to_double_cuda)
conv_param = {'stride': 1, 'pad': 1}
out, cache = Conv_ReLU.forward(x, w, b, conv_param)
dx, dw, db = Conv_ReLU.backward(dout, cache)
dx_num = compute_numeric_gradient(lambda x: Conv_ReLU.forward(x, w, b, conv_param)[0], x, dout)
dw_num = compute_numeric_gradient(lambda w: Conv_ReLU.forward(x, w, b, conv_param)[0], w, dout)
db_num = compute_numeric_gradient(lambda b: Conv_ReLU.forward(x, w, b, conv_param)[0], b, dout)
# Relative errors should be around e-8 or less
print('Testing Conv_ReLU:')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
# + [markdown] colab_type="text" id="Kgp7ymihw73P"
# # Three-layer convolutional network
# Now that you have implemented all the necessary layers, we can put them together into a simple convolutional network.
#
# Complete the implementation of the `ThreeLayerConvNet` class. We STRONGLY recommend you to use the fast/sandwich layers (already imported for you) in your implementation. Run the following cells to help you debug:
# + colab_type="code" id="DIN4bw-R0v0D" colab={}
class ThreeLayerConvNet(object):
"""
A three-layer convolutional network with the following architecture:
conv - relu - 2x2 max pool - linear - relu - linear - softmax
The network operates on minibatches of data that have shape (N, C, H, W)
consisting of N images, each with height H and width W and with C input
channels.
"""
def __init__(self, input_dims=(3, 32, 32), num_filters=32, filter_size=7,
hidden_dim=100, num_classes=10, weight_scale=1e-3, reg=0.0,
dtype=torch.float, device='cpu'):
"""
Initialize a new network.
Inputs:
- input_dims: Tuple (C, H, W) giving size of input data
- num_filters: Number of filters to use in the convolutional layer
- filter_size: Width/height of filters to use in the convolutional layer
- hidden_dim: Number of units to use in the fully-connected hidden layer
- num_classes: Number of scores to produce from the final linear layer.
- weight_scale: Scalar giving standard deviation for random initialization
of weights.
- reg: Scalar giving L2 regularization strength
- dtype: A torch data type object; all computations will be performed using
this datatype. float is faster but less accurate, so you should use
double for numeric gradient checking.
- device: device to use for computation. 'cpu' or 'cuda'
"""
self.params = {}
self.reg = reg
self.dtype = dtype
############################################################################
# TODO: Initialize weights and biases for the three-layer convolutional #
# network. Weights should be initialized from a Gaussian centered at 0.0 #
# with standard deviation equal to weight_scale; biases should be #
# initialized to zero. All weights and biases should be stored in the #
# dictionary self.params. Store weights and biases for the convolutional #
# layer using the keys 'W1' and 'b1'; use keys 'W2' and 'b2' for the #
# weights and biases of the hidden linear layer, and keys 'W3' and 'b3' #
# for the weights and biases of the output linear layer. #
# #
# IMPORTANT: For this assignment, you can assume that the padding #
# and stride of the first convolutional layer are chosen so that #
# **the width and height of the input are preserved**. Take a look at #
# the start of the loss() function to see how that happens. #
############################################################################
# Replace "pass" statement with your code
pass
############################################################################
# END OF YOUR CODE #
############################################################################
def loss(self, X, y=None):
"""
Evaluate loss and gradient for the three-layer convolutional network.
Input / output: Same API as TwoLayerNet.
"""
X = X.to(self.dtype)
W1, b1 = self.params['W1'], self.params['b1']
W2, b2 = self.params['W2'], self.params['b2']
W3, b3 = self.params['W3'], self.params['b3']
# pass conv_param to the forward pass for the convolutional layer
# Padding and stride chosen to preserve the input spatial size
filter_size = W1.shape[2]
conv_param = {'stride': 1, 'pad': (filter_size - 1) // 2}
# pass pool_param to the forward pass for the max-pooling layer
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
scores = None
############################################################################
# TODO: Implement the forward pass for the three-layer convolutional net, #
# computing the class scores for X and storing them in the scores #
# variable. #
# #
# Remember you can use the functions defined in your implementation above. #
############################################################################
# Replace "pass" statement with your code
pass
############################################################################
# END OF YOUR CODE #
############################################################################
if y is None:
return scores
loss, grads = 0.0, {}
############################################################################
# TODO: Implement the backward pass for the three-layer convolutional net, #
# storing the loss and gradients in the loss and grads variables. Compute #
# data loss using softmax, and make sure that grads[k] holds the gradients #
# for self.params[k]. Don't forget to add L2 regularization! #
# #
# NOTE: To ensure that your implementation matches ours and you pass the #
# automated tests, make sure that your L2 regularization does not include #
# a factor of 0.5 #
############################################################################
# Replace "pass" statement with your code
pass
############################################################################
# END OF YOUR CODE #
############################################################################
return loss, grads
# + [markdown] colab_type="text" id="e_MfqAQXw73Q"
# ## Sanity check loss
# After you build a new network, one of the first things you should do is sanity check the loss. When we use the softmax loss, we expect the loss for random weights (and no regularization) to be about `log(C)` for `C` classes. When we add regularization the loss should go up slightly.
# + colab_type="code" id="h-XEWaw2w73R" colab={}
fix_random_seed(0)
model = ThreeLayerConvNet(**to_double_cuda)
N = 50
X = torch.randn(N, 3, 32, 32, **to_double_cuda)
y = torch.randint(10, size=(N,), **to_long_cuda)
loss, grads = model.loss(X, y)
print('Initial loss (no regularization): ', loss.item())
model.reg = 0.5
loss, grads = model.loss(X, y)
print('Initial loss (with regularization): ', loss.item())
# + [markdown] colab_type="text" id="QEIViSCjw73U"
# ## Gradient check
# After the loss looks reasonable, use numeric gradient checking to make sure that your backward pass is correct. When you use numeric gradient checking you should use a small amount of artificial data and a small number of neurons at each layer.
#
# You should see errors less than `1e-5`.
# + colab_type="code" id="xPrOgIsJw73V" colab={}
num_inputs = 2
input_dims = (3, 16, 16)
reg = 0.0
num_classes = 10
fix_random_seed(0)
X = torch.randn(num_inputs, *input_dims, **to_double_cuda)
y = torch.randint(num_classes, size=(num_inputs,), **to_long_cuda)
model = ThreeLayerConvNet(num_filters=3, filter_size=3,
input_dims=input_dims, hidden_dim=7,
weight_scale=5e-2, **to_double_cuda)
loss, grads = model.loss(X, y)
for param_name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
param_grad_num = compute_numeric_gradient(f, model.params[param_name])
print('%s max relative error: %e' % (param_name, rel_error(param_grad_num, grads[param_name])))
# + [markdown] colab_type="text" id="dUPRjnzww73Y"
# ## Overfit small data
# A nice trick is to train your model with just a few training samples. You should be able to overfit small datasets, which will result in very high training accuracy and comparatively low validation accuracy.
# + colab_type="code" id="pwwQ0XB7w73Z" colab={}
fix_random_seed(0)
num_train = 100
small_data = {
'X_train': data_dict['X_train'][:num_train],
'y_train': data_dict['y_train'][:num_train],
'X_val': data_dict['X_val'],
'y_val': data_dict['y_val'],
}
model = ThreeLayerConvNet(weight_scale=1e-3, **to_float_cuda)
solver = Solver(model, small_data,
num_epochs=30, batch_size=50,
update_rule=adam,
optim_config={
'learning_rate': 2e-3,
},
verbose=True, print_every=1,
device='cuda')
solver.train()
# + [markdown] colab_type="text" id="KTDOqqdLw73d"
# Plotting the loss, training accuracy, and validation accuracy should show clear overfitting:
# + colab_type="code" id="fypbffqsw73f" colab={}
plt.title('Training losses')
plt.plot(solver.loss_history, 'o')
plt.xlabel('iteration')
plt.ylabel('loss')
plt.gcf().set_size_inches(9, 4)
plt.show()
plt.title('Train and Val accuracies')
plt.plot(solver.train_acc_history, '-o')
plt.plot(solver.val_acc_history, '-o')
plt.legend(['train', 'val'], loc='upper left')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.gcf().set_size_inches(9, 4)
plt.show()
# + [markdown] colab_type="text" id="W2vnSbfjw73i"
# ## Train the net
# By training the three-layer convolutional network for one epoch, you should achieve greater than 50% accuracy on the training set:
# + colab_type="code" id="nfArKG-Gw73j" colab={}
fix_random_seed(0)
model = ThreeLayerConvNet(weight_scale=0.001, hidden_dim=500, reg=0.001, dtype=torch.float, device='cuda')
solver = Solver(model, data_dict,
num_epochs=1, batch_size=64,
update_rule=adam,
optim_config={
'learning_rate': 2e-3,
},
verbose=True, print_every=50, device='cuda')
solver.train()
# + [markdown] colab_type="text" id="vIYQ0nm2w73n"
# ## Visualize Filters
# You can visualize the first-layer convolutional filters from the trained network by running the following:
# + colab_type="code" id="n3FLipRY4NUv" colab={}
from torchvision.utils import make_grid
nrow = math.ceil(math.sqrt(model.params['W1'].shape[0]))
grid = make_grid(model.params['W1'], nrow=nrow, padding=1, normalize=True, scale_each=True)
plt.imshow(grid.to(device='cpu').permute(1, 2, 0))
plt.axis('off')
plt.gcf().set_size_inches(5, 5)
plt.show()
# + [markdown] colab_type="text" id="JleotK9yDcyv"
# # Deep convolutional network
# Next you will implement a deep convolutional network with an arbitrary number of conv layers in VGGNet style.
#
# Read through the `DeepConvNet` class.
#
# Implement the initialization, the forward pass, and the backward pass. For the moment don't worry about implementing batch normalization; we will add those features soon. Again, we STRONGLY recommend you to use the fast/sandwich layers (already imported for you) in your implementation.
# + colab_type="code" id="Ah-_nwx2BSxl" colab={}
class DeepConvNet(object):
"""
A convolutional neural network with an arbitrary number of convolutional
layers in VGG-Net style. All convolution layers will use kernel size 3 and
padding 1 to preserve the feature map size, and all pooling layers will be
max pooling layers with 2x2 receptive fields and a stride of 2 to halve the
size of the feature map.
The network will have the following architecture:
{conv - [batchnorm?] - relu - [pool?]} x (L - 1) - linear
Each {...} structure is a "macro layer" consisting of a convolution layer,
an optional batch normalization layer, a ReLU nonlinearity, and an optional
pooling layer. After L-1 such macro layers, a single fully-connected layer
is used to predict the class scores.
The network operates on minibatches of data that have shape (N, C, H, W)
consisting of N images, each with height H and width W and with C input
channels.
"""
def __init__(self, input_dims=(3, 32, 32),
num_filters=[8, 8, 8, 8, 8],
max_pools=[0, 1, 2, 3, 4],
batchnorm=False,
num_classes=10, weight_scale=1e-3, reg=0.0,
weight_initializer=None,
dtype=torch.float, device='cpu'):
"""
Initialize a new network.
Inputs:
- input_dims: Tuple (C, H, W) giving size of input data
- num_filters: List of length (L - 1) giving the number of convolutional
filters to use in each macro layer.
- max_pools: List of integers giving the indices of the macro layers that
should have max pooling (zero-indexed).
- batchnorm: Whether to include batch normalization in each macro layer
- num_classes: Number of scores to produce from the final linear layer.
- weight_scale: Scalar giving standard deviation for random initialization
of weights, or the string "kaiming" to use Kaiming initialization instead
- reg: Scalar giving L2 regularization strength. L2 regularization should
only be applied to convolutional and fully-connected weight matrices;
it should not be applied to biases or to batchnorm scale and shifts.
- dtype: A torch data type object; all computations will be performed using
this datatype. float is faster but less accurate, so you should use
double for numeric gradient checking.
- device: device to use for computation. 'cpu' or 'cuda'
"""
self.params = {}
self.num_layers = len(num_filters)+1
self.max_pools = max_pools
self.batchnorm = batchnorm
self.reg = reg
self.dtype = dtype
if device == 'cuda':
device = 'cuda:0'
############################################################################
# TODO: Initialize the parameters for the DeepConvNet. All weights, #
# biases, and batchnorm scale and shift parameters should be stored in the #
# dictionary self.params. #
# #
# Weights for conv and fully-connected layers should be initialized #
# according to weight_scale. Biases should be initialized to zero. #
# Batchnorm scale (gamma) and shift (beta) parameters should be initilized #
# to ones and zeros respectively. #
############################################################################
# Replace "pass" statement with your code
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# With batch normalization we need to keep track of running means and
# variances, so we need to pass a special bn_param object to each batch
# normalization layer. You should pass self.bn_params[0] to the forward pass
# of the first batch normalization layer, self.bn_params[1] to the forward
# pass of the second batch normalization layer, etc.
self.bn_params = []
if self.batchnorm:
self.bn_params = [{'mode': 'train'} for _ in range(len(num_filters))]
# Check that we got the right number of parameters
if not self.batchnorm:
params_per_macro_layer = 2 # weight and bias
else:
params_per_macro_layer = 4 # weight, bias, scale, shift
num_params = params_per_macro_layer * len(num_filters) + 2
msg = 'self.params has the wrong number of elements. Got %d; expected %d'
msg = msg % (len(self.params), num_params)
assert len(self.params) == num_params, msg
# Check that all parameters have the correct device and dtype:
for k, param in self.params.items():
msg = 'param "%s" has device %r; should be %r' % (k, param.device, device)
assert param.device == torch.device(device), msg
msg = 'param "%s" has dtype %r; should be %r' % (k, param.dtype, dtype)
assert param.dtype == dtype, msg
def loss(self, X, y=None):
"""
Evaluate loss and gradient for the deep convolutional network.
Input / output: Same API as ThreeLayerConvNet.
"""
X = X.to(self.dtype)
mode = 'test' if y is None else 'train'
# Set train/test mode for batchnorm params since they
# behave differently during training and testing.
if self.batchnorm:
for bn_param in self.bn_params:
bn_param['mode'] = mode
scores = None
# pass conv_param to the forward pass for the convolutional layer
# Padding and stride chosen to preserve the input spatial size
filter_size = 3
conv_param = {'stride': 1, 'pad': (filter_size - 1) // 2}
# pass pool_param to the forward pass for the max-pooling layer
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
scores = None
############################################################################
# TODO: Implement the forward pass for the DeepConvNet, computing the #
# class scores for X and storing them in the scores variable. #
# #
# You should use the fast versions of convolution and max pooling layers, #
# or the convolutional sandwich layers, to simplify your implementation. #
############################################################################
# Replace "pass" statement with your code
pass
############################################################################
# END OF YOUR CODE #
############################################################################
if y is None:
return scores
loss, grads = 0, {}
############################################################################
# TODO: Implement the backward pass for the DeepConvNet, storing the loss #
# and gradients in the loss and grads variables. Compute data loss using #
# softmax, and make sure that grads[k] holds the gradients for #
# self.params[k]. Don't forget to add L2 regularization! #
# #
# NOTE: To ensure that your implementation matches ours and you pass the #
# automated tests, make sure that your L2 regularization does not include #
# a factor of 0.5 #
############################################################################
# Replace "pass" statement with your code
pass
############################################################################
# END OF YOUR CODE #
############################################################################
return loss, grads
# + [markdown] colab_type="text" id="0AC0R6dv059E"
# ## Sanity check loss
# After you build a new network, one of the first things you should do is sanity check the loss. When we use the softmax loss, we expect the loss for random weights (and no regularization) to be about `log(C)` for `C` classes. When we add regularization the loss should go up slightly.
# + colab_type="code" id="MPfK4L5P059L" colab={}
fix_random_seed(0)
input_dims = (3, 32, 32)
model = DeepConvNet(num_filters=[8, 64], max_pools=[0, 1], **to_double_cuda)
N = 50
X = torch.randn(N, *input_dims, **to_double_cuda)
y = torch.randint(10, size=(N,), **to_long_cuda)
loss, grads = model.loss(X, y)
print('Initial loss (no regularization): ', loss.item())
model.reg = 1.
loss, grads = model.loss(X, y)
print('Initial loss (with regularization): ', loss.item())
# + [markdown] colab_type="text" id="8BfRDkEj1TX8"
# ## Gradient check
# After the loss looks reasonable, use numeric gradient checking to make sure that your backward pass is correct. When you use numeric gradient checking you should use a small amount of artifical data and a small number of neurons at each layer.
#
# For the check with reg=0, you should see relative errors less than `1e-5`. For the check with reg=3.14, you should see relative errors less than `1e-4`
# + colab_type="code" id="qiPiP1X11TYA" colab={}
num_inputs = 2
input_dims = (3, 8, 8)
num_classes = 10
fix_random_seed(0)
X = torch.randn(num_inputs, *input_dims, **to_double_cuda)
y = torch.randint(num_classes, size=(num_inputs,), **to_long_cuda)
for reg in [0, 3.14]:
print('Running check with reg = ', reg)
model = DeepConvNet(input_dims=input_dims, num_classes=num_classes,
num_filters=[8, 8, 8],
max_pools=[0, 2],
reg=reg,
weight_scale=5e-2, **to_double_cuda)
loss, grads = model.loss(X, y)
# The relative errors should be up to the order of e-6
for name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
grad_num = compute_numeric_gradient(f, model.params[name])
print('%s max relative error: %e' % (name, rel_error(grad_num, grads[name])))
if reg == 0: print()
# + [markdown] colab_type="text" id="1_njNfEh3cxs"
# ## Overfit small data
# As another sanity check, make sure you can overfit a small dataset of 50 images. In the following cell, tweak the **learning rate** and **weight initialization scale** to overfit and achieve 100% training accuracy within 30 epochs.
# + colab_type="code" id="2NccCDJ3e1DR" colab={}
# TODO: Use a DeepConvNet to overfit 50 training examples by
# tweaking just the learning rate and initialization scale.
fix_random_seed(0)
num_train = 50
small_data = {
'X_train': data_dict['X_train'][:num_train],
'y_train': data_dict['y_train'][:num_train],
'X_val': data_dict['X_val'],
'y_val': data_dict['y_val'],
}
input_dims = small_data['X_train'].shape[1:]
weight_scale = 2e-3 # Experiment with this!
learning_rate = 1e-5 # Experiment with this!
############################################################################
# TODO: Change weight_scale and learning_rate so your model achieves 100% #
# training accuracy within 30 epochs. #
############################################################################
# Replace "pass" statement with your code
pass
############################################################################
# END OF YOUR CODE #
############################################################################
model = DeepConvNet(input_dims=input_dims, num_classes=10,
num_filters=[8, 16, 32, 64],
max_pools=[0, 1, 2, 3],
reg=1e-5, weight_scale=weight_scale, **to_float_cuda)
solver = Solver(model, small_data,
print_every=10, num_epochs=30, batch_size=10,
update_rule=adam,
optim_config={
'learning_rate': learning_rate,
},
device='cuda',
)
solver.train()
plt.plot(solver.loss_history, 'o')
plt.title('Training loss history')
plt.xlabel('Iteration')
plt.ylabel('Training loss')
plt.show()
# + [markdown] colab_type="text" id="FI3LznMXRnad"
# # Kaiming initialization
# So far, you manually tuned the weight scale and for weight initialization.
# However, this is inefficient when it comes to training deep neural networks; practically, as your weight matrix is larger, the weight scale should be small.
# Below you will implement [Kaiming initialization](http://arxiv-web3.library.cornell.edu/abs/1502.01852). For more details, refer to [cs231n note](http://cs231n.github.io/neural-networks-2/#init) and [PyTorch documentation](https://pytorch.org/docs/stable/nn.init.html#torch.nn.init.kaiming_normal_).
# + colab_type="code" id="y90Z6I18AdjW" colab={}
def kaiming_initializer(Din, Dout, K=None, relu=True, device='cpu',
dtype=torch.float32):
"""
Implement Kaiming initialization for linear and convolution layers.
Inputs:
- Din, Dout: Integers giving the number of input and output dimensions for
this layer
- K: If K is None, then initialize weights for a linear layer with Din input
dimensions and Dout output dimensions. Otherwise if K is a nonnegative
integer then initialize the weights for a convolution layer with Din input
channels, Dout output channels, and a kernel size of KxK.
- relu: If ReLU=True, then initialize weights with a gain of 2 to account for
a ReLU nonlinearity (Kaiming initializaiton); otherwise initialize weights
with a gain of 1 (Xavier initialization).
- device, dtype: The device and datatype for the output tensor.
Returns:
- weight: A torch Tensor giving initialized weights for this layer. For a
linear layer it should have shape (Din, Dout); for a convolution layer it
should have shape (Dout, Din, K, K).
"""
gain = 2. if relu else 1.
weight = None
if K is None:
###########################################################################
# TODO: Implement Kaiming initialization for linear layer. #
# The weight scale is sqrt(gain / fan_in), #
# where gain is 2 if ReLU is followed by the layer, or 1 if not, #
# and fan_in = num_in_channels (= Din). #
# The output should be a tensor in the designated size, dtype, and device.#
###########################################################################
# Replace "pass" statement with your code
pass
###########################################################################
# END OF YOUR CODE #
###########################################################################
else:
###########################################################################
# TODO: Implement Kaiming initialization for convolutional layer. #
# The weight scale is sqrt(gain / fan_in), #
# where gain is 2 if ReLU is followed by the layer, or 1 if not, #
# and fan_in = num_in_channels (= Din) * K * K #
# The output should be a tensor in the designated size, dtype, and device.#
###########################################################################
# Replace "pass" statement with your code
pass
###########################################################################
# END OF YOUR CODE #
###########################################################################
return weight
# + [markdown] colab_type="text" id="XkZL_lsufSVZ"
# # Convolutional nets with Kaiming initialization
# Now that you have a working implementation for Kaiming initialization, go back to your [`DeepConvnet`](#scrollTo=Ah-_nwx2BSxl). Modify your implementation to add Kaiming initialization.
#
# Concretely, when the `weight_scale` is set to `'kaiming'` in the constructor, you should initialize weights of convolutional and linear layers using `kaiming_initializer`. Once you are done, run the following to see the effect of kaiming initialization in deep CNNs.
#
# In this experiment, we train a 31-layer network with four different weight initialization schemes. Among them, only the Kaiming initialization method should achieve a non-random accuracy after one epoch of training.
#
# You may see `nan` loss when `weight_scale` is large, this shows a catastrophe of inappropriate weight initialization.
# + colab_type="code" id="sQJKcH60jfTm" colab={}
fix_random_seed(0)
# Try training a deep convolutional net with different weight initialization methods
num_train = 10000
small_data = {
'X_train': data_dict['X_train'][:num_train],
'y_train': data_dict['y_train'][:num_train],
'X_val': data_dict['X_val'],
'y_val': data_dict['y_val'],
}
input_dims = data_dict['X_train'].shape[1:]
weight_scales = ['kaiming', 1e-1, 1e-2, 1e-3]
solvers = []
for weight_scale in weight_scales:
print('Solver with weight scale: ', weight_scale)
model = DeepConvNet(input_dims=input_dims, num_classes=10,
num_filters=([8] * 10) + ([32] * 10) + ([128] * 10),
max_pools=[9, 19],
weight_scale=weight_scale,
reg=1e-5, **to_float_cuda)
solver = Solver(model, small_data,
num_epochs=1, batch_size=128,
update_rule=sgd_momentum,
optim_config={
'learning_rate': 2e-3,
},
print_every=20, device='cuda')
solver.train()
solvers.append(solver)
# + colab_type="code" id="6WDM0E5lHBmD" colab={}
def plot_training_history_init(title, xlabel, solvers, labels, plot_fn, marker='-o'):
plt.title(title)
plt.xlabel(xlabel)
for solver, label in zip(solvers, labels):
data = plot_fn(solver)
label = 'weight_scale=' + str(label)
plt.plot(data, marker, label=label)
plt.legend(loc='lower center', ncol=len(solvers))
# + colab_type="code" id="uG-dQDRCH_XD" colab={}
plt.subplot(3, 1, 1)
plot_training_history_init('Training loss','Iteration', solvers, weight_scales,
lambda x: x.loss_history, marker='o')
plt.subplot(3, 1, 2)
plot_training_history_init('Training accuracy','Epoch', solvers, weight_scales,
lambda x: x.train_acc_history)
plt.subplot(3, 1, 3)
plot_training_history_init('Validation accuracy','Epoch', solvers, weight_scales,
lambda x: x.val_acc_history)
plt.gcf().set_size_inches(15, 15)
plt.show()
# + [markdown] colab_type="text" id="ogx_-gJ1e1EO"
# # Train a good model!
# Train the best convolutional model that you can on CIFAR-10, storing your best model in the `best_model` variable. We require you to get at least 72% accuracy on the validation set using a convolutional net, within 60 seconds of training.
#
# You might find it useful to use batch normalization in your model. However, since we do not ask you to implement it CUDA-friendly, it might slow down training.
#
# Hint: Your model does not have to be too deep.
#
# Hint 2: We used `batch_size = 128` for training a model with 76% validation accuracy. You don't have to follow this, but it would save your time for hyperparameter search.
# + colab_type="code" id="UPRQvUDJe1EQ" colab={}
input_dims = data_dict['X_train'].shape[1:]
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = True
model = None
solver = None
################################################################################
# TODO: Train the best DeepConvNet that you can on CIFAR-10 within 60 seconds. #
################################################################################
# Replace "pass" statement with your code
pass
################################################################################
# END OF YOUR CODE #
################################################################################
solver.train(time_limit=60)
torch.backends.cudnn.benchmark = False
# + [markdown] colab_type="text" id="uZWeaN-6e1ET"
# # Test your model!
# Run your best model on the validation and test sets. You should achieve above 72% accuracy on the validation set and 70% accuracy on the test set.
#
# (Our best model gets 76% validation accuracy and 75% test accuracy -- can you beat ours?)
#
# + colab_type="code" id="xdVs_GEse1EU" colab={}
print('Validation set accuracy: ', solver.check_accuracy(data_dict['X_val'], data_dict['y_val']))
print('Test set accuracy: ', solver.check_accuracy(data_dict['X_test'], data_dict['y_test']))
# + [markdown] colab_type="text" id="KqsqNYOVwypM" tags=["pdf-title"]
# # Batch Normalization
# One way to make deep networks easier to train is to use more sophisticated optimization procedures such as SGD+momentum, RMSProp, or Adam. Another strategy is to change the architecture of the network to make it easier to train.
# One idea along these lines is batch normalization which was proposed by [1] in 2015.
#
# The idea is relatively straightforward. Machine learning methods tend to work better when their input data consists of uncorrelated features with zero mean and unit variance. When training a neural network, we can preprocess the data before feeding it to the network to explicitly decorrelate its features; this will ensure that the first layer of the network sees data that follows a nice distribution. However, even if we preprocess the input data, the activations at deeper layers of the network will likely no longer be decorrelated and will no longer have zero mean or unit variance since they are output from earlier layers in the network. Even worse, during the training process the distribution of features at each layer of the network will shift as the weights of each layer are updated.
#
# The authors of [1] hypothesize that the shifting distribution of features inside deep neural networks may make training deep networks more difficult. To overcome this problem, [1] proposes to insert batch normalization layers into the network. At training time, a batch normalization layer uses a minibatch of data to estimate the mean and standard deviation of each feature. These estimated means and standard deviations are then used to center and normalize the features of the minibatch. A running average of these means and standard deviations is kept during training, and at test time these running averages are used to center and normalize features.
#
# It is possible that this normalization strategy could reduce the representational power of the network, since it may sometimes be optimal for certain layers to have features that are not zero-mean or unit variance. To this end, the batch normalization layer includes learnable shift and scale parameters for each feature dimension.
#
# [1] [<NAME> and <NAME>, "Batch Normalization: Accelerating Deep Network Training by Reducing
# Internal Covariate Shift", ICML 2015.](https://arxiv.org/abs/1502.03167)
# + colab_type="code" id="EbSBc8mt4so0" colab={}
class BatchNorm(object):
@staticmethod
def forward(x, gamma, beta, bn_param):
raise NotImplementedError
@staticmethod
def backward(dout, cache):
raise NotImplementedError
@staticmethod
def backward_alt(dout, cache):
raise NotImplementedError
# + [markdown] colab_type="text" id="s0ELNJIlwypX"
# ## Batch normalization: forward
# Implement the batch normalization forward pass in the function `BatchNorm.forward`. Once you have done so, run the following to test your implementation.
#
# Referencing the paper linked to above in [1] may be helpful!
# + colab_type="code" id="LhULDxfU43f1" colab={}
def batchnorm_forward(x, gamma, beta, bn_param):
"""
Forward pass for batch normalization.
During training the sample mean and (uncorrected) sample variance are
computed from minibatch statistics and used to normalize the incoming data.
During training we also keep an exponentially decaying running mean of the
mean and variance of each feature, and these averages are used to normalize
data at test-time.
At each timestep we update the running averages for mean and variance using
an exponential decay based on the momentum parameter:
running_mean = momentum * running_mean + (1 - momentum) * sample_mean
running_var = momentum * running_var + (1 - momentum) * sample_var
Note that the batch normalization paper suggests a different test-time
behavior: they compute sample mean and variance for each feature using a
large number of training images rather than using a running average. For
this implementation we have chosen to use running averages instead since
they do not require an additional estimation step; the PyTorch
implementation of batch normalization also uses running averages.
Input:
- x: Data of shape (N, D)
- gamma: Scale parameter of shape (D,)
- beta: Shift paremeter of shape (D,)
- bn_param: Dictionary with the following keys:
- mode: 'train' or 'test'; required
- eps: Constant for numeric stability
- momentum: Constant for running mean / variance.
- running_mean: Array of shape (D,) giving running mean of features
- running_var Array of shape (D,) giving running variance of features
Returns a tuple of:
- out: of shape (N, D)
- cache: A tuple of values needed in the backward pass
"""
mode = bn_param['mode']
eps = bn_param.get('eps', 1e-5)
momentum = bn_param.get('momentum', 0.9)
N, D = x.shape
running_mean = bn_param.get('running_mean', torch.zeros(D, dtype=x.dtype, device=x.device))
running_var = bn_param.get('running_var', torch.zeros(D, dtype=x.dtype, device=x.device))
out, cache = None, None
if mode == 'train':
#######################################################################
# TODO: Implement the training-time forward pass for batch norm. #
# Use minibatch statistics to compute the mean and variance, use #
# these statistics to normalize the incoming data, and scale and #
# shift the normalized data using gamma and beta. #
# #
# You should store the output in the variable out. Any intermediates #
# that you need for the backward pass should be stored in the cache #
# variable. #
# #
# You should also use your computed sample mean and variance together #
# with the momentum variable to update the running mean and running #
# variance, storing your result in the running_mean and running_var #
# variables. #
# #
# Note that though you should be keeping track of the running #
# variance, you should normalize the data based on the standard #
# deviation (square root of variance) instead! #
# Referencing the original paper (https://arxiv.org/abs/1502.03167) #
# might prove to be helpful. #
#######################################################################
# Replace "pass" statement with your code
pass
#######################################################################
# END OF YOUR CODE #
#######################################################################
elif mode == 'test':
#######################################################################
# TODO: Implement the test-time forward pass for batch normalization. #
# Use the running mean and variance to normalize the incoming data, #
# then scale and shift the normalized data using gamma and beta. #
# Store the result in the out variable. #
#######################################################################
# Replace "pass" statement with your code
pass
#######################################################################
# END OF YOUR CODE #
#######################################################################
else:
raise ValueError('Invalid forward batchnorm mode "%s"' % mode)
# Store the updated running means back into bn_param
bn_param['running_mean'] = running_mean.detach()
bn_param['running_var'] = running_var.detach()
return out, cache
BatchNorm.forward = batchnorm_forward
# + [markdown] colab_type="text" id="q9cFW4heSrpt"
# After implementing the forward pass for batch normalization, you can run the following to sanity check your implementation. After running batch normalization with beta=0 and gamma=1, the data should have zero mean and unit variance.
#
# After running batch normalization with nontrivial beta and gamma, the output data should have mean approximately equal to beta, and std approximatly equal to gamma.
# + colab_type="code" id="7iB1mAkLwypZ" colab={}
# Check the training-time forward pass by checking means and variances
# of features both before and after batch normalization
def print_mean_std(x,dim=0):
means = ['%.3f' % xx for xx in x.mean(dim=dim).tolist()]
stds = ['%.3f' % xx for xx in x.std(dim=dim).tolist()]
print(' means: ', means)
print(' stds: ', stds)
print()
# Simulate the forward pass for a two-layer network
fix_random_seed(0)
N, D1, D2, D3 = 200, 50, 60, 3
X = torch.randn(N, D1, **to_double_cuda)
W1 = torch.randn(D1, D2, **to_double_cuda)
W2 = torch.randn(D2, D3, **to_double_cuda)
a = X.matmul(W1).clamp(min=0.).matmul(W2)
print('Before batch normalization:')
print_mean_std(a,dim=0)
# Run with gamma=1, beta=0. Means should be close to zero and stds close to one
gamma = torch.ones(D3, **to_double_cuda)
beta = torch.zeros(D3, **to_double_cuda)
print('After batch normalization (gamma=1, beta=0)')
a_norm, _ = BatchNorm.forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,dim=0)
# Run again with nontrivial gamma and beta. Now means should be close to beta
# and std should be close to gamma.
gamma = torch.tensor([1.0, 2.0, 3.0], **to_double_cuda)
beta = torch.tensor([11.0, 12.0, 13.0], **to_double_cuda)
print('After batch normalization (gamma=', gamma.tolist(), ', beta=', beta.tolist(), ')')
a_norm, _ = BatchNorm.forward(a, gamma, beta, {'mode': 'train'})
print_mean_std(a_norm,dim=0)
# + [markdown] colab_type="text" id="qGUPz4t_Tam4"
# We can sanity-check the test-time forward pass of batch normalization by running the following. First we run the training-time forward pass many times to "warm up" the running averages. If we then run a test-time forward pass, the output should have approximately zero mean and unit variance.
# + colab_type="code" id="w8lPQyI9wype" colab={}
fix_random_seed(0)
N, D1, D2, D3 = 200, 50, 60, 3
W1 = torch.randn(D1, D2, **to_double_cuda)
W2 = torch.randn(D2, D3, **to_double_cuda)
bn_param = {'mode': 'train'}
gamma = torch.ones(D3, **to_double_cuda)
beta = torch.zeros(D3, **to_double_cuda)
for t in range(500):
X = torch.randn(N, D1, **to_double_cuda)
a = X.matmul(W1).clamp(min=0.).matmul(W2)
BatchNorm.forward(a, gamma, beta, bn_param)
bn_param['mode'] = 'test'
X = torch.randn(N, D1, **to_double_cuda)
a = X.matmul(W1).clamp(min=0.).matmul(W2)
a_norm, _ = BatchNorm.forward(a, gamma, beta, bn_param)
# Means should be close to zero and stds close to one, but will be
# noisier than training-time forward passes.
print('After batch normalization (test-time):')
print_mean_std(a_norm,dim=0)
# + [markdown] colab_type="text" id="jt0hsHxIwypj"
# ## Batch normalization: backward
# Now implement the backward pass for batch normalization in the function `BatchNorm.backward`.
#
# To derive the backward pass you should write out the computation graph for batch normalization and backprop through each of the intermediate nodes. Some intermediates may have multiple outgoing branches; make sure to sum gradients across these branches in the backward pass.
#
# Please don't forget to implement the train and test mode separately.
#
# Once you have finished, run the following to numerically check your backward pass.
# + colab_type="code" id="HpjnmV8D6e0Y" colab={}
def batchnorm_backward(dout, cache):
"""
Backward pass for batch normalization.
For this implementation, you should write out a computation graph for
batch normalization on paper and propagate gradients backward through
intermediate nodes.
Inputs:
- dout: Upstream derivatives, of shape (N, D)
- cache: Variable of intermediates from batchnorm_forward.
Returns a tuple of:
- dx: Gradient with respect to inputs x, of shape (N, D)
- dgamma: Gradient with respect to scale parameter gamma, of shape (D,)
- dbeta: Gradient with respect to shift parameter beta, of shape (D,)
"""
dx, dgamma, dbeta = None, None, None
###########################################################################
# TODO: Implement the backward pass for batch normalization. Store the #
# results in the dx, dgamma, and dbeta variables. #
# Referencing the original paper (https://arxiv.org/abs/1502.03167) #
# might prove to be helpful. #
# Don't forget to implement train and test mode separately. #
###########################################################################
# Replace "pass" statement with your code
pass
###########################################################################
# END OF YOUR CODE #
###########################################################################
return dx, dgamma, dbeta
BatchNorm.backward = batchnorm_backward
# + colab_type="code" id="n2pMzvTqwypk" colab={}
# Gradient check batchnorm backward pass
fix_random_seed(0)
N, D = 4, 5
x = 5 * torch.randn(N, D, **to_double_cuda) + 12
gamma = torch.randn(D, **to_double_cuda)
beta = torch.randn(D, **to_double_cuda)
dout = torch.randn(N, D, **to_double_cuda)
bn_param = {'mode': 'train'}
fx = lambda x: BatchNorm.forward(x, gamma, beta, bn_param)[0]
fg = lambda a: BatchNorm.forward(x, a, beta, bn_param)[0]
fb = lambda b: BatchNorm.forward(x, gamma, b, bn_param)[0]
dx_num = compute_numeric_gradient(fx, x, dout)
da_num = compute_numeric_gradient(fg, gamma.clone(), dout)
db_num = compute_numeric_gradient(fb, beta.clone(), dout)
_, cache = BatchNorm.forward(x, gamma, beta, bn_param)
dx, dgamma, dbeta = BatchNorm.backward(dout, cache)
# You should expect to see relative errors between 1e-12 and 1e-9
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
# + [markdown] colab_type="text" id="hFxuZMUAwypp"
# ## Batch normalization: alternative backward
# In class we talked about two different implementations for the sigmoid backward pass. One strategy is to write out a computation graph composed of simple operations and backprop through all intermediate values. Another strategy is to work out the derivatives on paper. For example, you can derive a very simple formula for the sigmoid function's backward pass by simplifying gradients on paper.
#
# Surprisingly, it turns out that you can do a similar simplification for the batch normalization backward pass too!
#
# In the forward pass, given a set of inputs $X=\begin{bmatrix}x_1\\x_2\\...\\x_N\end{bmatrix}$,
#
# we first calculate the mean $\mu$ and variance $v$.
# With $\mu$ and $v$ calculated, we can calculate the standard deviation $\sigma$ and normalized data $Y$.
# The equations and graph illustration below describe the computation ($y_i$ is the i-th element of the vector $Y$).
#
# \begin{align}
# & \mu=\frac{1}{N}\sum_{k=1}^N x_k & v=\frac{1}{N}\sum_{k=1}^N (x_k-\mu)^2 \\
# & \sigma=\sqrt{v+\epsilon} & y_i=\frac{x_i-\mu}{\sigma}
# \end{align}
# + [markdown] colab_type="text" id="gVoopiQ7wypr"
# <img src="https://web.eecs.umich.edu/~justincj/teaching/eecs498/assets/a3/batchnorm_graph.png" width=691 height=202>
# + [markdown] colab_type="text" id="EDTJ6AXawypt" tags=["pdf-ignore"]
# The meat of our problem during backpropagation is to compute $\frac{\partial L}{\partial X}$, given the upstream gradient we receive, $\frac{\partial L}{\partial Y}.$ To do this, recall the chain rule in calculus gives us $\frac{\partial L}{\partial X} = \frac{\partial L}{\partial Y} \cdot \frac{\partial Y}{\partial X}$.
#
# The unknown/hart part is $\frac{\partial Y}{\partial X}$. We can find this by first deriving step-by-step our local gradients at
# $\frac{\partial v}{\partial X}$, $\frac{\partial \mu}{\partial X}$,
# $\frac{\partial \sigma}{\partial v}$,
# $\frac{\partial Y}{\partial \sigma}$, and $\frac{\partial Y}{\partial \mu}$,
# and then use the chain rule to compose these gradients (which appear in the form of vectors!) appropriately to compute $\frac{\partial Y}{\partial X}$.
#
# If it's challenging to directly reason about the gradients over $X$ and $Y$ which require matrix multiplication, try reasoning about the gradients in terms of individual elements $x_i$ and $y_i$ first: in that case, you will need to come up with the derivations for $\frac{\partial L}{\partial x_i}$, by relying on the Chain Rule to first calculate the intermediate $\frac{\partial \mu}{\partial x_i}, \frac{\partial v}{\partial x_i}, \frac{\partial \sigma}{\partial x_i},$ then assemble these pieces to calculate $\frac{\partial y_i}{\partial x_i}$.
#
# You should make sure each of the intermediary gradient derivations are all as simplified as possible, for ease of implementation.
#
# After doing so, implement the simplified batch normalization backward pass in the function `BatchNorm.backward_alt` and compare the two implementations by running the following. Your two implementations should compute nearly identical results, but the alternative implementation should be a bit faster.
# + colab_type="code" id="zTh1EemM7MD3" colab={}
def batchnorm_backward_alt(dout, cache):
"""
Alternative backward pass for batch normalization.
For this implementation you should work out the derivatives for the batch
normalizaton backward pass on paper and simplify as much as possible. You
should be able to derive a simple expression for the backward pass.
See the jupyter notebook for more hints.
Note: This implementation should expect to receive the same cache variable
as batchnorm_backward, but might not use all of the values in the cache.
Inputs / outputs: Same as batchnorm_backward
"""
dx, dgamma, dbeta = None, None, None
###########################################################################
# TODO: Implement the backward pass for batch normalization. Store the #
# results in the dx, dgamma, and dbeta variables. #
# #
# After computing the gradient with respect to the centered inputs, you #
# should be able to compute gradients with respect to the inputs in a #
# single statement; our implementation fits on a single 80-character line.#
###########################################################################
# Replace "pass" statement with your code
pass
###########################################################################
# END OF YOUR CODE #
###########################################################################
return dx, dgamma, dbeta
BatchNorm.backward_alt = batchnorm_backward_alt
# + colab_type="code" id="cJZp2i7ywypv" colab={}
fix_random_seed(0)
N, D = 128, 2048
x = 5 * torch.randn(N, D, **to_double_cuda) + 12
gamma = torch.randn(D, **to_double_cuda)
beta = torch.randn(D, **to_double_cuda)
dout = torch.randn(N, D, **to_double_cuda)
bn_param = {'mode': 'train'}
out, cache = BatchNorm.forward(x, gamma, beta, bn_param)
t1 = time.time()
dx1, dgamma1, dbeta1 = BatchNorm.backward(dout, cache)
t2 = time.time()
dx2, dgamma2, dbeta2 = BatchNorm.backward_alt(dout, cache)
t3 = time.time()
print('dx difference: ', rel_error(dx1, dx2))
print('dgamma difference: ', rel_error(dgamma1, dgamma2))
print('dbeta difference: ', rel_error(dbeta1, dbeta2))
print('speedup: %.2fx' % ((t2 - t1) / (t3 - t2)))
# + [markdown] colab_type="text" id="uIJWjzFZw73z"
# # Spatial Batch Normalization
# As proposed in the original paper, batch normalization can also be used for convolutional networks, but we need to tweak it a bit; the modification will be called "spatial batch normalization."
#
# Normally batch-normalization accepts inputs of shape `(N, D)` and produces outputs of shape `(N, D)`, where we normalize across the minibatch dimension `N`. For data coming from convolutional layers, batch normalization needs to accept inputs of shape `(N, C, H, W)` and produce outputs of shape `(N, C, H, W)` where the `N` dimension gives the minibatch size and the `(H, W)` dimensions give the spatial size of the feature map.
#
# If the feature map was produced using convolutions, then we expect every feature channel's statistics e.g. mean, variance to be relatively consistent both between different images, and different locations within the same image -- after all, every feature channel is produced by the same convolutional filter! Therefore spatial batch normalization computes a mean and variance for each of the `C` feature channels by computing statistics over the minibatch dimension `N` as well the spatial dimensions `H` and `W`.
#
#
# [1] [<NAME> and <NAME>, "Batch Normalization: Accelerating Deep Network Training by Reducing
# Internal Covariate Shift", ICML 2015.](https://arxiv.org/abs/1502.03167)
# + colab_type="code" id="8Czv0Qu0BsDh" colab={}
class SpatialBatchNorm(object):
@staticmethod
def forward(x, gamma, beta, bn_param):
raise NotImplementedError
@staticmethod
def backward(dout, cache):
raise NotImplementedError
# + [markdown] colab_type="text" id="oqCmKCw7w730"
# ## Spatial batch normalization: forward
#
# Implement the forward pass for spatial batch normalization in the function `SpatialBatchNorm.forward`. Check your implementation by running the following:
# + colab_type="code" id="oBjsUeK_Bq7-" colab={}
def spatial_batchnorm_forward(x, gamma, beta, bn_param):
"""
Computes the forward pass for spatial batch normalization.
Inputs:
- x: Input data of shape (N, C, H, W)
- gamma: Scale parameter, of shape (C,)
- beta: Shift parameter, of shape (C,)
- bn_param: Dictionary with the following keys:
- mode: 'train' or 'test'; required
- eps: Constant for numeric stability
- momentum: Constant for running mean / variance. momentum=0 means that
old information is discarded completely at every time step, while
momentum=1 means that new information is never incorporated. The
default of momentum=0.9 should work well in most situations.
- running_mean: Array of shape (D,) giving running mean of features
- running_var Array of shape (D,) giving running variance of features
Returns a tuple of:
- out: Output data, of shape (N, C, H, W)
- cache: Values needed for the backward pass
"""
out, cache = None, None
###########################################################################
# TODO: Implement the forward pass for spatial batch normalization. #
# #
# HINT: You can implement spatial batch normalization by calling the #
# vanilla version of batch normalization you implemented above. #
# Your implementation should be very short; ours is less than five lines. #
###########################################################################
# Replace "pass" statement with your code
pass
###########################################################################
# END OF YOUR CODE #
###########################################################################
return out, cache
SpatialBatchNorm.forward = spatial_batchnorm_forward
# + [markdown] colab_type="text" id="5ZHLPa6-UkY1"
# After implementing the forward pass for spatial batch normalization, you can run the following to sanity check your code.
# + colab_type="code" id="3x-vfMnIw732" colab={}
fix_random_seed(0)
# Check the training-time forward pass by checking means and variances
# of features both before and after spatial batch normalization
N, C, H, W = 2, 3, 4, 5
x = 4 * torch.randn(N, C, H, W, **to_double_cuda) + 10
print('Before spatial batch normalization:')
print(' Shape: ', x.shape)
print(' Means: ', x.mean(dim=(0, 2, 3)))
print(' Stds: ', x.std(dim=(0, 2, 3)))
# Means should be close to zero and stds close to one
gamma, beta = torch.ones(C, **to_double_cuda), torch.zeros(C, **to_double_cuda)
bn_param = {'mode': 'train'}
out, _ = SpatialBatchNorm.forward(x, gamma, beta, bn_param)
print('After spatial batch normalization:')
print(' Shape: ', out.shape)
print(' Means: ', out.mean(dim=(0, 2, 3)))
print(' Stds: ', out.std(dim=(0, 2, 3)))
# Means should be close to beta and stds close to gamma
gamma, beta = torch.tensor([3, 4, 5], **to_double_cuda), torch.tensor([6, 7, 8], **to_double_cuda)
out, _ = SpatialBatchNorm.forward(x, gamma, beta, bn_param)
print('After spatial batch normalization (nontrivial gamma, beta):')
print(' Shape: ', out.shape)
print(' Means: ', out.mean(dim=(0, 2, 3)))
print(' Stds: ', out.std(dim=(0, 2, 3)))
# + [markdown] colab_type="text" id="RxzqSGqZUr2J"
# Similar to the vanilla batch normalization implementation, run the following to sanity-check the test-time forward pass of spatial batch normalization.
# + colab_type="code" id="7z2Eu2Xlw736" colab={}
fix_random_seed(0)
# Check the test-time forward pass by running the training-time
# forward pass many times to warm up the running averages, and then
# checking the means and variances of activations after a test-time
# forward pass.
N, C, H, W = 10, 4, 11, 12
bn_param = {'mode': 'train'}
gamma = torch.ones(C, **to_double_cuda)
beta = torch.zeros(C, **to_double_cuda)
for t in range(50):
x = 2.3 * torch.randn(N, C, H, W, **to_double_cuda) + 13
SpatialBatchNorm.forward(x, gamma, beta, bn_param)
bn_param['mode'] = 'test'
x = 2.3 * torch.randn(N, C, H, W, **to_double_cuda) + 13
a_norm, _ = SpatialBatchNorm.forward(x, gamma, beta, bn_param)
# Means should be close to zero and stds close to one, but will be
# noisier than training-time forward passes.
print('After spatial batch normalization (test-time):')
print(' means: ', a_norm.mean(dim=(0, 2, 3)))
print(' stds: ', a_norm.std(dim=(0, 2, 3)))
# + [markdown] colab_type="text" id="dVdE3j3iw739"
# ## Spatial batch normalization: backward
# Implement the backward pass for spatial batch normalization in the function `SpatialBatchNorm.backward`.
# + colab_type="code" id="v9TqgGJ0DlCV" colab={}
def spatial_batchnorm_backward(dout, cache):
"""
Computes the backward pass for spatial batch normalization.
Inputs:
- dout: Upstream derivatives, of shape (N, C, H, W)
- cache: Values from the forward pass
Returns a tuple of:
- dx: Gradient with respect to inputs, of shape (N, C, H, W)
- dgamma: Gradient with respect to scale parameter, of shape (C,)
- dbeta: Gradient with respect to shift parameter, of shape (C,)
"""
dx, dgamma, dbeta = None, None, None
###########################################################################
# TODO: Implement the backward pass for spatial batch normalization. #
# #
# HINT: You can implement spatial batch normalization by calling the #
# vanilla version of batch normalization you implemented above. #
# Your implementation should be very short; ours is less than five lines. #
###########################################################################
# Replace "pass" statement with your code
pass
###########################################################################
# END OF YOUR CODE #
###########################################################################
return dx, dgamma, dbeta
SpatialBatchNorm.backward = spatial_batchnorm_backward
# + [markdown] colab_type="text" id="B5efGyfwU3rK"
# After implementing the backward pass for spatial batch normalization, run the following to perform numeric gradient checking on your implementation. You should see errors less than `1e-6`.
# + colab_type="code" id="5Z2j4mQlw73_" colab={}
fix_random_seed(0)
N, C, H, W = 2, 3, 4, 5
x = 5 * torch.randn(N, C, H, W, **to_double_cuda) + 12
gamma = torch.randn(C, **to_double_cuda)
beta = torch.randn(C, **to_double_cuda)
dout = torch.randn(N, C, H, W, **to_double_cuda)
bn_param = {'mode': 'train'}
fx = lambda x: SpatialBatchNorm.forward(x, gamma, beta, bn_param)[0]
fg = lambda a: SpatialBatchNorm.forward(x, gamma, beta, bn_param)[0]
fb = lambda b: SpatialBatchNorm.forward(x, gamma, beta, bn_param)[0]
dx_num = compute_numeric_gradient(fx, x, dout)
da_num = compute_numeric_gradient(fg, gamma, dout)
db_num = compute_numeric_gradient(fb, beta, dout)
_, cache = SpatialBatchNorm.forward(x, gamma, beta, bn_param)
dx, dgamma, dbeta = SpatialBatchNorm.backward(dout, cache)
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
# + [markdown] colab_type="text" id="9qS4seVFEkAN"
# # "Sandwich" layers with batch normalization
# Again, below you will find sandwich layers that implement a few commonly used patterns for convolutional networks.
# + colab_type="code" id="FUY0tkJTEkAU" colab={}
class Linear_BatchNorm_ReLU(object):
@staticmethod
def forward(x, w, b, gamma, beta, bn_param):
"""
Convenience layer that performs an linear transform, batch normalization,
and ReLU.
Inputs:
- x: Array of shape (N, D1); input to the linear layer
- w, b: Arrays of shape (D2, D2) and (D2,) giving the weight and bias for
the linear transform.
- gamma, beta: Arrays of shape (D2,) and (D2,) giving scale and shift
parameters for batch normalization.
- bn_param: Dictionary of parameters for batch normalization.
Returns:
- out: Output from ReLU, of shape (N, D2)
- cache: Object to give to the backward pass.
"""
a, fc_cache = Linear.forward(x, w, b)
a_bn, bn_cache = BatchNorm.forward(a, gamma, beta, bn_param)
out, relu_cache = ReLU.forward(a_bn)
cache = (fc_cache, bn_cache, relu_cache)
return out, cache
@staticmethod
def backward(dout, cache):
"""
Backward pass for the linear-batchnorm-relu convenience layer.
"""
fc_cache, bn_cache, relu_cache = cache
da_bn = ReLU.backward(dout, relu_cache)
da, dgamma, dbeta = BatchNorm.backward(da_bn, bn_cache)
dx, dw, db = Linear.backward(da, fc_cache)
return dx, dw, db, dgamma, dbeta
class Conv_BatchNorm_ReLU(object):
@staticmethod
def forward(x, w, b, gamma, beta, conv_param, bn_param):
a, conv_cache = FastConv.forward(x, w, b, conv_param)
an, bn_cache = SpatialBatchNorm.forward(a, gamma, beta, bn_param)
out, relu_cache = ReLU.forward(an)
cache = (conv_cache, bn_cache, relu_cache)
return out, cache
@staticmethod
def backward(dout, cache):
conv_cache, bn_cache, relu_cache = cache
dan = ReLU.backward(dout, relu_cache)
da, dgamma, dbeta = SpatialBatchNorm.backward(dan, bn_cache)
dx, dw, db = FastConv.backward(da, conv_cache)
return dx, dw, db, dgamma, dbeta
class Conv_BatchNorm_ReLU_Pool(object):
@staticmethod
def forward(x, w, b, gamma, beta, conv_param, bn_param, pool_param):
a, conv_cache = FastConv.forward(x, w, b, conv_param)
an, bn_cache = SpatialBatchNorm.forward(a, gamma, beta, bn_param)
s, relu_cache = ReLU.forward(an)
out, pool_cache = FastMaxPool.forward(s, pool_param)
cache = (conv_cache, bn_cache, relu_cache, pool_cache)
return out, cache
@staticmethod
def backward(dout, cache):
conv_cache, bn_cache, relu_cache, pool_cache = cache
ds = FastMaxPool.backward(dout, pool_cache)
dan = ReLU.backward(ds, relu_cache)
da, dgamma, dbeta = SpatialBatchNorm.backward(dan, bn_cache)
dx, dw, db = FastConv.backward(da, conv_cache)
return dx, dw, db, dgamma, dbeta
# + [markdown] colab_type="text" id="1C8k4fRgwyp1"
# # Convolutional nets with batch normalization
# Now that you have a working implementation for batch normalization, go back to your [`DeepConvnet`](#scrollTo=Ah-_nwx2BSxl). Modify your implementation to add batch normalization.
#
# Concretely, when the `batchnorm` flag is set to `True` in the constructor, you should insert a batch normalization layer before each ReLU nonlinearity. The outputs from the last linear layer of the network should not be normalized. Once you are done, run the following to gradient-check your implementation.
#
# In the reg=0 case, you should see errors less than `1e-6` for all weights and batchnorm parameters (beta and gamma); for biases you should see errors less than `1e-2`.
#
# In the reg=3.14 case, you should see errors less than `1e-6` for all parameters.
# + colab_type="code" id="E5sLZ6J1wyp6" colab={}
num_inputs = 2
input_dims = (3, 8, 8)
num_classes = 10
fix_random_seed(0)
X = torch.randn(num_inputs, *input_dims, **to_double_cuda)
y = torch.randint(num_classes, size=(num_inputs,), **to_long_cuda)
for reg in [0, 3.14]:
print('Running check with reg = ', reg)
model = DeepConvNet(input_dims=input_dims, num_classes=num_classes,
num_filters=[8, 8, 8],
max_pools=[0, 2],
reg=reg, batchnorm=True,
weight_scale='kaiming',
**to_double_cuda)
loss, grads = model.loss(X, y)
# The relative errors should be up to the order of e-3
for name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
grad_num = compute_numeric_gradient(f, model.params[name])
print('%s max relative error: %e' % (name, rel_error(grad_num, grads[name])))
if reg == 0: print()
# + [markdown] colab_type="text" id="T-D9TV8qwyp9"
# # Batchnorm for deep convolutional networks
# Run the following to train a deep convolutional network on a subset of 500 training examples both with and without batch normalization.
# + colab_type="code" id="IrrKxsVGwyp_" colab={}
fix_random_seed(0)
# Try training a deep convolutional net with batchnorm
num_train = 500
small_data = {
'X_train': data_dict['X_train'][:num_train],
'y_train': data_dict['y_train'][:num_train],
'X_val': data_dict['X_val'],
'y_val': data_dict['y_val'],
}
input_dims = data_dict['X_train'].shape[1:]
bn_model = DeepConvNet(input_dims=input_dims, num_classes=10,
num_filters=[16, 32, 32, 64, 64],
max_pools=[0, 1, 2, 3, 4],
weight_scale='kaiming',
batchnorm=True,
reg=1e-5, **to_float_cuda)
model = DeepConvNet(input_dims=input_dims, num_classes=10,
num_filters=[16, 32, 32, 64, 64],
max_pools=[0, 1, 2, 3, 4],
weight_scale='kaiming',
batchnorm=False,
reg=1e-5, **to_float_cuda)
print('Solver with batch norm:')
bn_solver = Solver(bn_model, small_data,
num_epochs=10, batch_size=100,
update_rule=adam,
optim_config={
'learning_rate': 1e-3,
},
print_every=20, device='cuda')
bn_solver.train()
print('\nSolver without batch norm:')
solver = Solver(model, small_data,
num_epochs=10, batch_size=100,
update_rule=adam,
optim_config={
'learning_rate': 1e-3,
},
print_every=20, device='cuda')
solver.train()
# + [markdown] colab_type="text" id="4XZ7jZE9wyqE"
# Run the following to visualize the results from two networks trained above. You should find that using batch normalization helps the network to converge much faster.
# + colab_type="code" id="1oBBnMuGehfi" colab={}
def plot_training_history_bn(title, label, solvers, bn_solvers, plot_fn, bl_marker='.', bn_marker='.', labels=None):
"""utility function for plotting training history"""
plt.title(title)
plt.xlabel(label)
bn_plots = [plot_fn(bn_solver) for bn_solver in bn_solvers]
bl_plots = [plot_fn(solver) for solver in solvers]
num_bn = len(bn_plots)
num_bl = len(bl_plots)
for i in range(num_bn):
label='w/ BN'
if labels is not None:
label += str(labels[i])
plt.plot(bn_plots[i], bn_marker, label=label)
for i in range(num_bl):
label='w/o BN'
if labels is not None:
label += str(labels[i])
plt.plot(bl_plots[i], bl_marker, label=label)
plt.legend(loc='lower center', ncol=num_bn+num_bl)
# + colab_type="code" id="PVYMhWo9wyqG" tags=["pdf-ignore-input"] colab={}
plt.subplot(3, 1, 1)
plot_training_history_bn('Training loss','Iteration', [solver], [bn_solver], \
lambda x: x.loss_history, bl_marker='-o', bn_marker='-o')
plt.subplot(3, 1, 2)
plot_training_history_bn('Training accuracy','Epoch', [solver], [bn_solver], \
lambda x: x.train_acc_history, bl_marker='-o', bn_marker='-o')
plt.subplot(3, 1, 3)
plot_training_history_bn('Validation accuracy','Epoch', [solver], [bn_solver], \
lambda x: x.val_acc_history, bl_marker='-o', bn_marker='-o')
plt.gcf().set_size_inches(15, 15)
plt.show()
# + [markdown] colab_type="text" id="YV6-pJEdSXMh"
# # Batch normalization and learning rate
# We will now run a small experiment to study the interaction of batch normalization and learning rate.
#
# The first cell will train convolutional networks with different learning rates. The second layer will plot training accuracy and validation set accuracy over time. You should find that using batch normalization helps the network to be less dependent to the learning rate.
# + colab_type="code" id="yVSQ5SesSXMq" tags=["pdf-ignore-input"] colab={}
fix_random_seed(0)
# Try training a very deep net with batchnorm
num_train = 10000
small_data = {
'X_train': data_dict['X_train'][:num_train],
'y_train': data_dict['y_train'][:num_train],
'X_val': data_dict['X_val'],
'y_val': data_dict['y_val'],
}
input_dims = data_dict['X_train'].shape[1:]
num_epochs = 5
lrs = [2e-1, 1e-1, 5e-2]
lrs = [5e-3, 1e-2, 2e-2]
solvers = []
for lr in lrs:
print('No normalization: learning rate = ', lr)
model = DeepConvNet(input_dims=input_dims, num_classes=10,
num_filters=[8, 8, 8],
max_pools=[0, 1, 2],
weight_scale='kaiming',
batchnorm=False,
reg=1e-5, **to_float_cuda)
solver = Solver(model, small_data,
num_epochs=num_epochs, batch_size=100,
update_rule=sgd_momentum,
optim_config={
'learning_rate': lr,
},
verbose=False, device='cuda')
solver.train()
solvers.append(solver)
bn_solvers = []
for lr in lrs:
print('Normalization: learning rate = ', lr)
bn_model = DeepConvNet(input_dims=input_dims, num_classes=10,
num_filters=[8, 8, 16, 16, 32, 32],
max_pools=[1, 3, 5],
weight_scale='kaiming',
batchnorm=True,
reg=1e-5, **to_float_cuda)
bn_solver = Solver(bn_model, small_data,
num_epochs=num_epochs, batch_size=128,
update_rule=sgd_momentum,
optim_config={
'learning_rate': lr,
},
verbose=False, device='cuda')
bn_solver.train()
bn_solvers.append(bn_solver)
# + colab_type="code" id="kBE8AT5SSXMv" colab={}
plt.subplot(2, 1, 1)
plot_training_history_bn('Training accuracy (Batch Normalization)','Epoch', solvers, bn_solvers, \
lambda x: x.train_acc_history, bl_marker='-^', bn_marker='-o', labels=[' lr={:.0e}'.format(lr) for lr in lrs])
plt.subplot(2, 1, 2)
plot_training_history_bn('Validation accuracy (Batch Normalization)','Epoch', solvers, bn_solvers, \
lambda x: x.val_acc_history, bl_marker='-^', bn_marker='-o', labels=[' lr={:.0e}'.format(lr) for lr in lrs])
plt.gcf().set_size_inches(15, 10)
plt.show()
| convolutional_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Train a VAE on L1000 Data
# +
import sys
import pathlib
import numpy as np
import pandas as pd
sys.path.insert(0, "../../scripts")
from utils import load_data, infer_L1000_features
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
from sklearn.decomposition import PCA
from tensorflow import keras
from vae import VAE
from tensorflow.keras.models import Model, Sequential
import seaborn
import tensorflow as tf
# -
data_splits = ["train", "valid", "test", "complete"]
data_dict = load_data(data_splits, dataset="L1000")
# +
# Prepare data for training
meta_features = infer_L1000_features(data_dict["train"], metadata=True)
profile_features = infer_L1000_features(data_dict["train"])
train_features_df = data_dict["train"].reindex(profile_features, axis="columns")
train_meta_df = data_dict["train"].reindex(meta_features, axis="columns")
valid_features_df = data_dict["valid"].reindex(profile_features, axis="columns")
valid_meta_df = data_dict["valid"].reindex(meta_features, axis="columns")
test_features_df = data_dict["test"].reindex(profile_features, axis="columns")
test_meta_df = data_dict["test"].reindex(meta_features, axis="columns")
complete_features_df = data_dict["complete"].reindex(profile_features, axis="columns")
complete_meta_df = data_dict["complete"].reindex(meta_features, axis="columns")
# -
print(train_features_df.shape)
train_features_df.head(3)
print(valid_features_df.shape)
valid_features_df.head(3)
print(test_features_df.shape)
test_features_df.head(3)
print(complete_features_df.shape)
complete_features_df.head(3)
# +
encoder_architecture = [500]
decoder_architecture = [500]
# +
L1000_vae = VAE(
input_dim=train_features_df.shape[1],
latent_dim=65,
batch_size=512,
encoder_batch_norm=True,
epochs=180,
learning_rate=0.001,
encoder_architecture=encoder_architecture,
decoder_architecture=decoder_architecture,
beta=1,
verbose=True,
)
L1000_vae.compile_vae()
#1495
# -
L1000_vae.train(x_train=train_features_df, x_test=valid_features_df)
L1000_vae.vae
# Save training performance
history_df = pd.DataFrame(L1000_vae.vae.history.history)
history_df
#1067 for level 1
history_df.to_csv('training_data/twolayer_training_vanilla.csv')
plt.figure(figsize=(10, 5))
plt.plot(history_df["loss"], label="Training data")
plt.plot(history_df["val_loss"], label="Validation data")
plt.title("Loss for VAE training on L1000 data")
plt.ylabel("MSE + KL Divergence")
plt.ylabel("Loss")
plt.xlabel("No. Epoch")
plt.legend()
plt.show()
# evaluating performance using test set
L1000_vae.vae.evaluate(test_features_df)
reconstruction = pd.DataFrame(
L1000_vae.vae.predict(test_features_df), columns=profile_features
)
(sum(sum((np.array(test_features_df) - np.array(reconstruction)) ** 2))) ** 0.5
# latent space heatmap
fig, ax = plt.subplots(figsize=(10, 10))
encoder = L1000_vae.encoder_block["encoder"]
latent = np.array(encoder.predict(test_features_df)[2])
seaborn.heatmap(latent, ax=ax)
# +
reconstruction = pd.DataFrame(
L1000_vae.vae.predict(test_features_df), columns=profile_features
)
pca = PCA(n_components=2).fit(test_features_df)
pca_reconstructed_latent_df = pd.DataFrame(pca.transform(reconstruction))
pca_test_latent_df = pd.DataFrame(pca.transform(test_features_df))
# -
figure(figsize=(10, 10), dpi=80)
plt.scatter(pca_test_latent_df[0],pca_test_latent_df[1], marker = ".", alpha = 0.5)
plt.scatter(pca_reconstructed_latent_df[0],pca_reconstructed_latent_df[1], marker = ".", alpha = 0.5)
import umap
reducer = umap.UMAP().fit(test_features_df)
original_test_embedding = reducer.transform(test_features_df)
reconstructed_test_embedding = reducer.transform(reconstruction)
figure(figsize=(10, 10), dpi=80)
plt.scatter(original_test_embedding[:,0],original_test_embedding[:,1], marker = ".", alpha = 0.5)
plt.scatter(reconstructed_test_embedding[:,0],reconstructed_test_embedding[:,1], marker = ".", alpha = 0.5)
decoder = L1000_vae.decoder_block["decoder"]
# +
pca_training = PCA(n_components=2).fit(train_features_df)
simulated_df = pd.DataFrame(np.random.normal(size=(94440, 65)), columns=np.arange(0,65))
reconstruction_of_simulated = decoder.predict(simulated_df)
pca_reconstruction_of_simulated = pd.DataFrame(pca_training.transform(reconstruction_of_simulated))
pca_train_latent_df = pd.DataFrame(pca_training.transform(train_features_df))
fig, (ax1,ax2) = plt.subplots(1, 2, figsize=(16,8), sharey = True, sharex = True)
ax1.scatter(pca_train_latent_df[0],pca_train_latent_df[1], marker = ".", alpha = 0.5)
ax2.scatter(pca_reconstruction_of_simulated[0],pca_reconstruction_of_simulated[1], marker = ".", alpha = 0.5)
# -
from scipy.spatial.distance import directed_hausdorff
max(directed_hausdorff(reconstruction_of_simulated, train_features_df)[0],directed_hausdorff(train_features_df,reconstruction_of_simulated)[0])
#NOTE: IF YOU RUN THIS, YOU WILL NOT BE ABLE TO REPRODUCE THE EXACT RESULTS IN THE EXPERIMENT
latent_complete = np.array(encoder.predict(complete_features_df)[2])
latent_df = pd.DataFrame(latent_complete)
latent_df.to_csv("../3.application/latentTwoLayer_vanilla.csv")
#NOTE: IF YOU RUN THIS, YOU WILL NOT BE ABLE TO REPRODUCE THE EXACT RESULTS IN THE EXPERIMENT
decoder.save('./models/L1000twolayerDecoder_vanilla')
#NOTE: IF YOU RUN THIS, YOU WILL NOT BE ABLE TO REPRODUCE THE EXACT RESULTS IN THE EXPERIMENT
encoder.save('./models/L1000twolayerEncoder_vanilla')
| L1000/2.train/0.L1000-vae-TwoLayer-vanilla.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Loan Application Environment
# language: python
# name: loan_env
# ---
# +
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import plotly.express as px
from sklearn.model_selection import train_test_split
import statsmodels.api as sm
from sklearn.base import BaseEstimator,TransformerMixin
from sklearn.impute import KNNImputer
from sklearn.pipeline import Pipeline,FeatureUnion
from sklearn.preprocessing import OrdinalEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import f1_score,auc,classification_report,confusion_matrix,roc_curve,roc_auc_score,recall_score
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import FeatureUnion
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import FunctionTransformer
from sklearn.preprocessing import LabelEncoder
from sklearn.ensemble import RandomForestClassifier
from imblearn.over_sampling import SMOTE
# from imblearn.pipeline import Pipeline
#
# from imblearn.over_sampling import SMOTE
import joblib
# -
df = pd.read_csv(r"../data/raw/data.csv")
df.head(15)
df[["Loan_Amount_Term","Loan_ID"]].groupby(by=["Loan_Amount_Term"]).count().sort_values(by=["Loan_ID"])
# +
cat_columns = ['Self_Employed',
'Dependents',
'Gender',
'Married',
'Education',
'Property_Area',
'Credit_History']
num_columns = ['LoanAmount','ApplicantIncome','TotalApplicantIncome']
# -
class PipeCustomOrdinalEncoder(BaseEstimator,TransformerMixin):
'''
Converts categories into numbers for KNNImputing
'''
def __init__(self):
self.ord_encoder = OrdinalEncoder()
# self.feature = feature_name
def fit(self,X,y=None):
na_indices = X[X.isna()].index.values
not_na_rows = X[X.notna()]
not_na_np = not_na_rows.to_numpy().reshape(-1,1)
self.ord_encoder = self.ord_encoder.fit(not_na_np)
return self
def transform(self,X,y=None):
na_indices = X[X.isna()].index.values
not_na_rows = X[X.notna()]
not_na_np = not_na_rows.to_numpy().reshape(-1,1)
transformed_data = self.ord_encoder.transform(not_na_np)
not_na_encode = pd.Series(transformed_data.flatten(),index = not_na_rows.index.values)
return pd.DataFrame(pd.concat([not_na_encode,X[X.isna()]]).sort_index())
def inverse_transform(self,X,y=None):
col = X.to_numpy().reshape(-1,1)
return self.ord_encoder.inverse_transform(col)
class CustomKNNImputer(BaseEstimator,TransformerMixin):
def __init__(self,n_neighbors):
self.n_neighbors = n_neighbors
self.imputer = KNNImputer(n_neighbors=self.n_neighbors)
def fit(self,X,y=None):
self.imputer = self.imputer.fit(X)
return self.imputer
def transform(self,X,y=None):
return pd.DataFrame(self.imputer.transform(X),columns=X.columns)
def fit_transform(self,X,y=None):
self.imputer = self.imputer.fit(X)
return self.transform(X,y)
def return_df(x,cols=None):
return pd.DataFrame(x,columns=cols)
def transform_dataset(data):
data.loc[:,"LoanAmount"] = np.log(data.LoanAmount)
data.loc[:,"TotalApplicantIncome"] = np.log(data.ApplicantIncome+data.CoapplicantIncome)
data.loc[:,"ApplicantIncome"] = np.log(data.ApplicantIncome)
data = data.drop(columns=['CoapplicantIncome','Loan_ID','Loan_Amount_Term'])
return data
# +
# class custom_smote(BaseEstimator, TransformerMixin):
# def __init__(self):
# self.smt = SMOTE(k_neighbors=5)
# def fit(self,X,y=None):
# self.smt = self.smt.fit(X,y)
# return self
# def transform(self,X,y=None):
# x,_ = self.smt.fit_resample(X,y)
# return x
# + [markdown] tags=[]
# ## Pipelines
# -
transformations = FunctionTransformer(transform_dataset)
# +
cat_preprocess = ColumnTransformer(transformers=[
('employed',PipeCustomOrdinalEncoder(),'Self_Employed'),
('dependents',PipeCustomOrdinalEncoder(),'Dependents'),
('married',PipeCustomOrdinalEncoder(),'Married'),
('gender',PipeCustomOrdinalEncoder(),'Gender'),
('education',PipeCustomOrdinalEncoder(),'Education'),
('property',PipeCustomOrdinalEncoder(),'Property_Area'),
('credit-history',PipeCustomOrdinalEncoder(),'Credit_History')
])
categories = Pipeline(steps=[
('cat',cat_preprocess),
('cat_names',FunctionTransformer(return_df,kw_args={"cols":cat_columns}))
])
# +
num_preprocess = ColumnTransformer(
transformers=[
('num_scaling',MinMaxScaler(),num_columns)
])
numerical = Pipeline(steps=[
('num',num_preprocess),
('num_names',FunctionTransformer(return_df,kw_args={"cols":num_columns}))
])
# -
impute_preprocess = Pipeline(steps=[
('transformations',transformations),
('feature_union',FeatureUnion(transformer_list=[
('cat_pipe',categories),
('num_pipe',numerical)
])),
('test',FunctionTransformer(return_df,kw_args={"cols":cat_columns+num_columns})),
('imputer',CustomKNNImputer(n_neighbors=1,))
])
one_hot_encoding_preprocess = FeatureUnion(transformer_list=[
('cat_features',ColumnTransformer(
transformers=[('categorical',OneHotEncoder(),cat_columns),]
)),
('num_features',ColumnTransformer(
transformers=[('numeric','passthrough',num_columns),]
))
])
preprocessing = Pipeline(steps=[
('impute',impute_preprocess),
('one_hot_encoded',one_hot_encoding_preprocess)
])
pipe = Pipeline(steps=[
('preprocess',preprocessing),
# ('smote',custom_smote()),
('model',LogisticRegression(random_state=123,fit_intercept=True,max_iter=1000))
],verbose=True)
# ## Modelling
X = df.drop(columns=['Loan_Status'])
y = df['Loan_Status']
X_train,X_test,y_train,y_test = train_test_split(X,y,stratify=df['Loan_Status'],random_state=60,train_size=0.6)
X_train = X_train.sort_index()
y_train = y_train.sort_index()
X_test = X_test.sort_index()
y_test = y_test.sort_index()
# + tags=[]
label_encoder = LabelEncoder()
label_encoder = label_encoder.fit(y_train)
y_train = label_encoder.transform(y_train)
y_test = label_encoder.transform(y_test)
# + tags=[]
# X_train_1 = preprocessing.fit_transform(X_train)
# X_test_1 = preprocessing.transform(X_test)
# +
# clf = LogisticRegression(random_state=123,fit_intercept=True,max_iter=1000)
# -
params = {}
# + jupyter={"outputs_hidden": true} tags=[]
model = GridSearchCV(pipe,param_grid=params)
model = model.fit(X_train,y_train)
# + tags=[]
f1_score(y_train,model.predict(X_train))
# + tags=[]
f1_score(y_test,model.predict(X_test))
# -
print(classification_report(y_test,model.predict(X_test)))
print(confusion_matrix(y_test,model.predict(X_test)))
X_test
X_test[~np.where(model.predict(X_test)==1,True,False)]
raw_row = X_train.sample(1)
raw_row
new_row = pd.Series(raw_row.to_dict('records')[0])
new_row = pd.DataFrame(new_row).T
new_row
data_types = raw_row.dtypes.to_dict()
type(data_types['Loan_ID'])
new_row = new_row.astype(data_types)
model.predict(raw_row)
model.predict(new_row)
recall_score(y_test,model.predict(X_test))
recall_score(y_train,model.predict(X_train))
y_pred_proba = model.predict_proba(X_test)[:,1]
fpr, tpr, _ = roc_curve(y_test, y_pred_proba)
auc = roc_auc_score(y_test, y_pred_proba)
plt.plot(fpr,tpr,label="data 1, auc="+str(auc))
plt.legend(loc=4)
plt.show()
# +
from sklearn import metrics
preds = model.predict_proba(X_test)
tpr, tpr, thresholds = roc_curve(y_test,preds[:,1])
print (thresholds)
accuracy_ls = []
for thres in thresholds:
y_pred = np.where(preds[:,1]>thres,1,0)
# Apply desired utility function to y_preds, for example accuracy.
accuracy_ls.append(f1_score(y_test, y_pred))
# -
joblib.dump(model,r"../models/1_base_model.sav")
# +
random_forest_pipe = Pipeline(steps=[
('preprocess',preprocessing),
('model',RandomForestClassifier())
],verbose=True)
# -
params= {
"model__n_estimators" : [20,50,100,150],
"model__max_depth" : [2,3,5,6],
}
model_forest = GridSearchCV(random_forest_pipe,param_grid=params,cv=4)
X_train,X_test,y_train,y_test = train_test_split(X,y,stratify=df['Loan_Status'],random_state=60,train_size=0.6)
X_train = X_train.sort_index()
y_train = y_train.sort_index()
X_test = X_test.sort_index()
y_test = y_test.sort_index()
# + tags=[]
label_encoder = LabelEncoder()
label_encoder = label_encoder.fit(y_train)
y_train = label_encoder.transform(y_train)
y_test = label_encoder.transform(y_test)
# -
params = {}
# + jupyter={"outputs_hidden": true} tags=[]
model_forest.fit(X_train,y_train)
# -
f1_score(y_train,model_forest.predict(X_train))
# + jupyter={"outputs_hidden": true} tags=[]
model_forest.best_estimator_
# -
f1_score(y_test,model_forest.predict(X_test))
recall_score(y_test,model_forest.predict(X_test))
recall_score(y_train,model_forest.predict(X_train))
print(confusion_matrix(y_test,model_forest.predict(X_test)))
import xgboost as xgb
X_train,X_test,y_train,y_test = train_test_split(X,y,stratify=df['Loan_Status'],random_state=60,train_size=0.6)
X_train = X_train.sort_index()
y_train = y_train.sort_index()
X_test = X_test.sort_index()
y_test = y_test.sort_index()
label_encoder = LabelEncoder()
label_encoder = label_encoder.fit(y_train)
y_train = label_encoder.transform(y_train)
y_test = label_encoder.transform(y_test)
params= {
"model__n_estimators" : [20,50,100,150],
"model__max_depth" : [2,3,5,6],
}
# +
xgb_pipe = Pipeline(steps=[
('preprocess',preprocessing),
('model',xgb.XGBClassifier())
],verbose=True)
model_xgb = GridSearchCV(xgb_pipe,param_grid=params,cv=4)
# + jupyter={"outputs_hidden": true} tags=[]
model_xgb.fit(X_train,y_train)
# -
f1_score(y_test,model_xgb.predict(X_test))
f1_score(y_train,model_xgb.predict(X_train))
recall_score(y_test,model_xgb.predict(X_test))
recall_score(y_train,model_xgb.predict(X_train))
print(confusion_matrix(y_test,model_xgb.predict(X_test)))
| notebooks/4.1_Pipelines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 7.2
# language: ''
# name: sagemath
# ---
runfile SuperTriangle.py
runfile ComparisonCode.py
G = SuperTriangle(2)
G.show()
#L = G.laplacian_matrix()
#sp = L.eigenvalues()
#sp.sort()
#sp
runfile ResistorNetworkTransformations.py
# +
G = SuperTriangle(2)
G = GraphtoWeighted(G)
elist = G.edges()
G = DeltaY(G,0,1,2)
G = DeltaY(G,2,4,5)
G = DeltaY(G,1,3,4)
G.plot(edge_labels=True)
# -
G = series(G, 1)
G = series(G, 2)
G = series(G, 4)
G.plot(edge_labels = True)
# +
edgelist = G.edges()
print edgelist
G = DeltaY(G,6,7,8)
G.plot(edge_labels=True)
# -
G = SuperTriangle(3)
G.show()
G = GraphtoWeighted(G)
el = G.edges()
G = DeltaY(G,0,1,2 )
G = DeltaY(G,1,3,4 )
G = DeltaY(G,2,4,5 )
G = DeltaY(G,3,6,7 )
G = DeltaY(G,4,7,8 )
G = DeltaY(G,5,8,9 )
G.plot(edge_labels = True)
G = series(G, 1)
G = series(G, 2)
G = series(G, 3)
G = series(G, 5)
G = series(G, 7)
G = series(G, 8)
G = YDelta(G, 4)
G.plot(edge_labels = True)
G = DeltaY(G, 10,11,12)
G = DeltaY(G, 11,13,14)
G = DeltaY(G, 12,14,15)
G.plot(edge_labels = True)
G = series(G,11)
G = series(G,12)
G = series(G,14)
G.plot(edge_labels = True)
G = DeltaY(G, (16,18,4/7), (16,17,4/7), (17,18,4/7))
G.plot(edge_labels = True)
s = 4/21 + 4/21 + 1/3
s*2
G = SuperTriangle(4)
G.show()
G = GraphtoWeighted(G)
G = DeltaY(G, 0,1,2)
G = DeltaY(G, 1,3,4)
G = DeltaY(G, 2,4,5)
G = DeltaY(G, 3,6,7)
G = DeltaY(G, 4,7,8)
G = DeltaY(G, 5,8,9)
G = DeltaY(G, 6,10,11)
G = DeltaY(G, 7,11,12)
G = DeltaY(G, 8,12,13)
G = DeltaY(G, 9,13,14)
G.plot(edge_labels = True)
# +
G = series(G,1)
G = series(G,3)
G = series(G,6)
G = series(G,11)
G = series(G,12)
G = series(G,13)
G = series(G,2)
G = series(G,5)
G = series(G,9)
G=YDelta(G, 4)
G=YDelta(G, 7)
G=YDelta(G, 8)
G.plot(edge_labels = True)
# +
G = DeltaY(G, 15,16,17)
G = DeltaY(G, 16,18,19)
G = DeltaY(G, 17,19,20)
G = DeltaY(G, 18,21,22)
G = DeltaY(G, 19,22,23)
G = DeltaY(G, 20,23,24)
G.plot(edge_labels = True)
# +
G = series(G,16)
G = series(G,17)
G = series(G,18)
G = series(G,20)
G = series(G,22)
G = series(G,23)
G.plot(edge_labels = True)
# -
G = YDelta(G, 19)
G.plot(edge_labels = True)
G = DeltaY(G, 26,28,29)
G = DeltaY(G, 25,26,27)
G = DeltaY(G, 27,29,30)
G.plot(edge_labels = True)
G = series(G, 27)
G = series(G, 26)
G = series(G, 29)
G.plot(edge_labels = True)
G = DeltaY(G, 31,32,33)
G.plot(edge_labels = True)
1/3 + 4/21 + 75/574 + 15/82
2*103/123
# +
# -
| Resistance for SuperTriangles3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/krakowiakpawel9/data-science-bootcamp/blob/master/06_uczenie_maszynowe/01_sklearn_intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="VLSdropBJv7f" colab_type="text"
# * @author: <EMAIL>
# * @site: e-smartdata.org
# + [markdown] id="mj9rJ7JCQAFO" colab_type="text"
# ### scikit-learn
# >Strona biblioteki: [https://scikit-learn.org](https://scikit-learn.org)
# >
# >Dokumentacja/User Guide: [https://scikit-learn.org/stable/user_guide.html](https://scikit-learn.org/stable/user_guide.html)
# >
# >Podstawowa biblioteka do uczenia maszynowego w języku Python.
# >
# >Aby zainstalować bibliotekę scikit-learn, użyj polecenia poniżej:
# ```
# pip install scikit-learn
# ```
# >Aby zaktualizować bibliotekę scikit-learn, użyj polecenia poniżej:
# ```
# pip install -q --upgrade scikit-learn
# ```
# Główne kategorie uczenia maszynowego:
# * Uczenie Nadzorowane:
# - klasyfikacja
# - regresja
# * Uczenie Nienadzorowane:
# - klasteryzacja
# - redukcja wymiarowości
# + id="AEE5ZeP_mtQo" colab_type="code" outputId="7b8e0a1e-ee50-48ae-9626-24f175828662" colab={"base_uri": "https://localhost:8080/", "height": 34}
import sklearn
sklearn.__version__
# + [markdown] id="8KHlhP68bKId" colab_type="text"
# ### Klasyfikacja
# + id="jU_GseY8cE0Y" colab_type="code" outputId="13fd7205-6f90-4ef0-cbb5-cab28f17a065" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import numpy as np
from sklearn import datasets
np.random.seed(10)
raw_data = datasets.load_iris()
raw_data
# + id="lOpB2oWLcTTj" colab_type="code" outputId="f9c5eed1-5120-4d97-ef0a-d7513e11a5a5" colab={"base_uri": "https://localhost:8080/", "height": 34}
raw_data.keys()
# + id="mQQRmWpNcYyk" colab_type="code" outputId="cbe76e88-ddce-4178-e644-5dce7b66f9cd" colab={"base_uri": "https://localhost:8080/", "height": 1000}
print(raw_data.DESCR)
# + id="0MblEs6QckdJ" colab_type="code" colab={}
data = raw_data.data
target = raw_data.target
# + id="iFv2UgJ_c2Bg" colab_type="code" outputId="8342dc1b-56c5-4f0c-c31f-0db423537159" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(data.shape)
print(target.shape)
# + id="U15NZiG0dGp0" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
data_train, data_test, target_train, target_test = train_test_split(data, target, test_size=0.3)
# + id="Sam7WYGpdtId" colab_type="code" outputId="933ce065-bda1-48ef-9a38-720d74630f8b" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(data_train.shape)
print(target_train.shape)
# + id="fBgxSYbmdzY9" colab_type="code" outputId="85f9a6ae-7999-40a4-fa63-55de47887a66" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(data_test.shape)
print(target_test.shape)
# + id="UDrpnhvPd73o" colab_type="code" outputId="f1557f25-e23c-4749-8b02-68bff98f2568" colab={"base_uri": "https://localhost:8080/", "height": 191}
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(data_train, target_train)
# + id="0Rle8DPbeT4O" colab_type="code" outputId="cad4c595-58b0-4b00-bb66-e7edc167c176" colab={"base_uri": "https://localhost:8080/", "height": 68}
target_pred = model.predict(data_test)
target_pred
# + id="fhiOLiMfegaI" colab_type="code" outputId="5cb751cc-e96e-41cd-ade6-bbe5d5eeba0e" colab={"base_uri": "https://localhost:8080/", "height": 68}
from sklearn.metrics import confusion_matrix
confusion_matrix(target_test, target_pred)
# + id="8ORNQp4Fe5tQ" colab_type="code" outputId="11472695-44e1-4dd1-884a-0a51e841d245" colab={"base_uri": "https://localhost:8080/", "height": 188}
from sklearn.metrics import classification_report
print(classification_report(target_test, target_pred))
# + id="4RKoAVMMfNwL" colab_type="code" colab={}
| 06_uczenie_maszynowe/01_sklearn_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Python code for generating figures used in the paper "What Determines the Sizes of Bars in Spiral Galaxies?" (Erwin 2019, submitted)
# ## Setup
# ### General Setup
# +
# %pylab inline
matplotlib.rcParams['figure.figsize'] = (8,6)
matplotlib.rcParams['xtick.labelsize'] = 16
matplotlib.rcParams['ytick.labelsize'] = 16
matplotlib.rcParams['axes.labelsize'] = 20
# kludge to fix matplotlib's font_manager bug which mistakenly ID's "Times New Roman Bold.ttf" as
# indicating a "roman" (i.e., "normal") weight
try:
del matplotlib.font_manager.weight_dict['roman']
except KeyError:
pass
matplotlib.font_manager._rebuild()
# +
import os
import scipy.stats
import plotutils as pu
from loess import loess_1d as cap_loess_1d
# CHANGE THIS TO POINT TO APPROPRIATE LOCAL DIRECTORY (DEFAULT = SAME DIRECTORY AS THIS NOTEBOOK)
#projDir = "/Users/erwin/Documents/Working/Projects/Project_BarSizes/"
projDir = os.getcwd() + "/"
sys.path.append(projDir)
import datautils as du
import fitting_barsizes
# CHANGE THIS IF BAR-SIZE DATA TABLE IS NOT IN SAME DIRECTORY AS THIS NOTEBOOK
dataDir = projDir
s4gdata = du.ReadCompositeTable(dataDir + "s4gbars_table.dat", columnRow=25, dataFrame=True)
# This is where the best-fit parameter coefficients are saved
bestfitParamsFile = projDir + "bestfit_parameters.txt"
# CHANGE THIS TO POINT TO APPROPRIATE LOCAL DIRECTORY -- this is where PDF figures will be saved
baseDir = "/Users/erwin/Documents/Working/Papers/Papers-s4gbars/"
plotDir = baseDir + "plots/"
nDisksTotal = len(s4gdata.name)
mm = np.arange(8,12,0.01)
# set the following to True in order to save the figures as PDF files
savePlots = False
# -
# ### Useful Functions
# +
# code to make use of Cappellari LOESS function more useful
def GetSortedLoess1D( x, y, frac=0.5, degree=1 ):
ii_sorted = np.argsort(x)
x_sorted = x[ii_sorted]
y_sorted = y[ii_sorted]
x_sorted, y_predicted, w = cap_loess_1d.loess_1d(x_sorted, y_sorted, frac=frac, degree=degree)
return x_sorted, y_predicted
def GetSortedLoess1DResiduals( x, y, frac=0.5, degree=1 ):
ii_sorted = np.argsort(x)
x_sorted = x[ii_sorted]
y_sorted = y[ii_sorted]
x_sorted, y_predicted, w = cap_loess_1d.loess_1d(x_sorted, y_sorted, frac=frac, degree=degree)
return x_sorted, y_sorted - y_predicted, ii_sorted
# code for fits
def simplefunc_brokenlinear( params, x ):
"""Simple function to compute broken-linear fit:
y = alpha_1 + beta_1*x, x < x_break
= alpha_2 + beta_2*x, x >= x_break
Parameters
----------
params : sequence of float
[alpha_1, beta_1, x_break, beta_2]
note that alpha_2 can be computed from the other parameters
x : float or numpy ndarray of float
x values
Returns
-------
y : float or numpy ndarray of float
see definition above
"""
alpha_1 = params[0]
beta_1 = params[1]
x_break = params[2]
beta_2 = params[3]
alpha_2 = alpha_1 + (beta_1 - beta_2)*x_break
if (np.iterable(x) == 0):
x = [x]
npts = len(x)
vals = []
for i in range(npts):
if x[i] < x_break:
value = alpha_1 + beta_1*x[i]
else:
value = alpha_2 + beta_2*x[i]
vals.append(value)
return np.array(vals)
def simplefunc_linear( params, x ):
"""Simple function to compute linear fit:
y = alpha + beta*x
Parameters
----------
params : sequence of float
[alpha, beta]
x : float or numpy ndarray of float
x values
Returns
-------
y : float or numpy ndarray of float
y = alpha + beta*x
"""
alpha = params[0]
beta = params[1]
npts = len(x)
vals = []
for i in range(npts):
value = alpha + beta*x[i]
vals.append(value)
return np.array(vals)
def GetBinMeansAndErrors( xVals, indices, binStart, binStop, binWidth ):
"""Generate binned mean values and standard deviations for bar size,
binned by xVals (e.g., logMstar).
We assume that xVals are logarithmic values.
Parameters
----------
xVals : sequence of float
x values in logarithmic form (e.g., log10(M_star))
indices : list of int
indices into xVals specifying a particular subsample
binStart : float
binStop : float
binWidth : float
specifications for bin boundaries
Returns
-------
(x_bin_centers, bin_means, bin_errs) : tuple of float
"""
xx = xVals[indices]
yy = np.log10(s4gdata.sma_dp_kpc2[indices])
x_bins = np.arange(binStart, binStop, binWidth)
halfWidth = binWidth / 2
x_bin_centers = np.arange(binStart + halfWidth, binStop - halfWidth, binWidth)
digitized = np.digitize(xx, x_bins)
bin_means = np.array([yy[digitized == i].mean() for i in range(1, len(x_bins))])
bin_stddev = np.array([yy[digitized == i].std() for i in range(1, len(x_bins))])
# convert means and errors to linear values
bin_means_lin = 10**bin_means
bin_err_low = bin_means_lin - 10**(bin_means - bin_stddev)
bin_err_high = 10**(bin_means + bin_stddev) - bin_means_lin
bin_errs = np.array([bin_err_low,bin_err_high])
return (x_bin_centers, bin_means_lin, bin_errs)
# -
# ### Plot labels
# +
# useful definitions for printing and plotting labels
xtmstar = r"$\log \: (M_{\star} / M_{\odot})$"
xth_kpc = r"Disc scale length $h$ [kpc]"
xtRe_kpc = r"Galaxy half-light radius $R_{e}$ [kpc]"
xtfgas = r"$\log \: (M_{\rm HI} / M_{\star})$"
xtHtype = r"Hubble Type $T$"
ytbarsize_kpc = r"Bar size $a_{\rm vis}$ [kpc]"
ytresid = r"Residuals: $\log \, (a_{\rm vis}) - \log \, ({\rm predicted})$"
ytresid_Re = r"Residuals: $\log \, R_{e, {\rm obs}} - \log \, ({\rm predicted})$"
ytresid_h = r"Residuals: $\log \, h - \log \, ({\rm predicted})$"
s4g_txt = r"S$^{4}$G"
s4g_fwhm = r"S$^{4}$G $\langle$FWHM$\rangle$"
s4g_2fwhm = r"$2 \times$ S$^{4}$G $\langle$FWHM$\rangle$"
# -
# ### Subsamples
# Importing the variables defined in `sample_defs.py` provides lists of int values, which can be used to index numpy 1D arrays, thereby defining different subsamples of S4G.
#
# E.g., the R_e values of all the barred galaxies can be defined as
#
# s4gdata.Re_kpc[ii_barred]
# **Location of subsample definitions:**
#
# projDir + sample_defs.py
from sample_defs import *
# For reference, here are the definitions in that file:
# +
# "limited2" or "lim2" = D <= 30 Mpc
# "m9to11 = logmstar = 9--11
# general subsamples: all barred, all unbarred, all spirals
ii_barred = [i for i in range(nDisksTotal) if s4gdata.sma[i] > 0]
ii_unbarred = [i for i in range(nDisksTotal) if s4gdata.sma[i] <= 0]
ii_spirals = [i for i in range(nDisksTotal) if s4gdata.t_s4g[i] > -0.5]
# Limited subsample 2: spirals with D <= 30 Mpc -- 856 galaxies: 483 barred, 373 unbarred
ii_all_limited2 = [i for i in ii_spirals if s4gdata.dist[i] <= 30]
ii_barred_limited2 = [i for i in ii_all_limited2 if i in ii_barred]
ii_unbarred_limited2 = [i for i in ii_all_limited2 if i not in ii_barred]
# Parent Spiral Sample: spirals with D <= 30 Mpc + logMstar = 9--11:
ii_all_limited2_m9to11 = [i for i in ii_all_limited2 if s4gdata.logmstar[i] >= 9 and s4gdata.logmstar[i] <= 11]
ii_barred_limited2_m9to11 = [i for i in ii_all_limited2_m9to11 if i in ii_barred]
# Spirals with R_e *and* h
ii_all_Reh = [i for i in ii_spirals if s4gdata.Re_kpc[i] > 0 and s4gdata.h_kpc[i] > 0]
ii_barred_Reh = [i for i in ii_all_Reh if i in ii_barred]
ii_unbarred_Reh = [i for i in ii_all_Reh if i not in ii_barred]
ii_all_Reh_m9to11 = [i for i in ii_spirals if s4gdata.logmstar[i] >= 9 and s4gdata.logmstar[i] <= 11 and s4gdata.Re_kpc[i] > 0 and s4gdata.h_kpc[i] > 0]
ii_barred_Reh_m9to11 = [i for i in ii_all_Reh_m9to11 if i in ii_barred]
ii_unbarred_Reh_m9to11 = [i for i in ii_all_Reh_m9to11 if i not in ii_barred]
ii_all_limited2_Reh = [i for i in ii_all_Reh if s4gdata.dist[i] <= 30]
ii_barred_limited2_Reh = [i for i in ii_all_limited2_Reh if i in ii_barred]
ii_unbarred_limited2_Reh = [i for i in ii_all_limited2_Reh if i not in ii_barred]
# Main Spiral Sample: Spirals with D < 30 Mpc, valid R_e *and* h, *and* logMstar = 9--11:
ii_all_lim2m9to11_Reh = [i for i in ii_all_limited2_Reh if s4gdata.logmstar[i] >= 9 and s4gdata.logmstar[i] <= 11]
ii_barred_lim2m9to11_Reh = [i for i in ii_all_lim2m9to11_Reh if i in ii_barred]
ii_unbarred_lim2m9to11_Reh = [i for i in ii_all_lim2m9to11_Reh if i not in ii_barred]
# useful aliases which are more directly descriptive
ii_all_D30 = ii_all_limited2
ii_all_D30_m9to11 = ii_all_limited2_m9to11
ii_barred_D30_m9to11 = ii_barred_limited2_m9to11
ii_barred_D30_m9to11_Reh = ii_barred_lim2m9to11_Reh
# +
print("Parent Disc Sample:")
N_parent_disc = len(ii_barred) + len(ii_unbarred)
print("\tN(all): %d" % N_parent_disc)
print("\tN(ii_barred): %d" % len(ii_barred))
print("Parent Spiral Sample: Spirals with D < 30 Mpc and logMstar = 9--11")
print("\tN(ii_all_limited2_m9to11): %d" % len(ii_all_limited2_m9to11))
print("\tN(ii_barred_limited2_m9to11 = ii_barred_D30_m9to11): %d" % len(ii_barred_limited2_m9to11))
print("Main Spiral Sample: Spirals with D < 30 Mpc, valid R_e *and* h, *and* logMstar = 9--11")
print("\tN(ii_all_lim2m9to11_Reh): %d" % len(ii_all_lim2m9to11_Reh))
print("\tN(ii_barred_lim2m9to11_Reh = ii_barred_D30_m9to11_Reh): %d" % len(ii_barred_lim2m9to11_Reh))
# -
# global data vectors in log10 format
logbarsize_all = np.log10(s4gdata.sma_dp_kpc2)
logRe_all = np.log10(s4gdata.Re_kpc)
logh_all = np.log10(s4gdata.h_kpc)
# ### Get best-fit parameters from file
# **Note:** Best-fit parameters are generated (and stored in output file) by Jupyter notebook barsize_fits.ipynb
dlines = [line for line in open(bestfitParamsFile) if line[0] != '#']
fitDict = {}
for line in dlines:
pp = line.split(":")
fitName = pp[0]
ppp = pp[1].split()
params = [float(p) for p in ppp]
fitDict[fitName] = params
# # Figures
# ## Figure 1: Barsize vs logMstar w/ LOESS and linear + broken-linear fit
# plots/barsize-logMstar_loess+fits -- **assembled in Adobe Illustrator**:
# 1. plots/barsize-vs-mass-with-loess.pdf
# 2. plots/barsize-vs-mass-with-fits_final.pdf
#
# Generate binned mean values and standard deviations
logmstar_bin_centers, bin_means_lin, bin_errs = GetBinMeansAndErrors(s4gdata.logmstar, ii_barred_limited2, 8.5,11.5,0.25)
# +
# LOESS fit
s4gxx_05, s4gyy_05 = GetSortedLoess1D(s4gdata.logmstar[ii_barred_limited2], np.log10(s4gdata.sma_dp_kpc2[ii_barred_limited2]), frac=0.5)
s4gyy_05 = 10**s4gyy_05
semilogy(s4gdata.logmstar[ii_barred_limited2], s4gdata.sma_dp_kpc2[ii_barred_limited2], 'ko', mfc='0.8',ms=5, alpha=0.7, zorder=1)
errorbar(logmstar_bin_centers, bin_means_lin, yerr=bin_errs, fmt='s', mfc='None', mec='k', mew=1.2, ms=15, ecolor='k', capsize=5, elinewidth=1.2, zorder=2)
xlim(8.5,11.5);ylim(0.1,30)
plot(s4gxx_05, s4gyy_05, 'k', zorder=3)
semilogy(10.7, 6.0, '*', mec='g', mew=2.5, mfc='None', ms=20, alpha=0.75, zorder=4)
xlabel(xtmstar);ylabel(ytbarsize_kpc)
pu.MakeNiceLogAxes(whichAxis="y")
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
savePlots = True
if savePlots:
plt.savefig(plotDir+"barsize-vs-mass-with-loess.pdf")
# +
semilogy(s4gdata.logmstar[ii_barred_limited2], s4gdata.sma_dp_kpc2[ii_barred_limited2], 'ko', mfc='0.8',ms=5, alpha=0.7, zorder=1)
semilogy(s4gdata.logmstar[ii_barred_D30_m9to11], s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11], 'ko', mfc='0.5',ms=5, alpha=0.7, zorder=2)
xlim(8.5,11.5);plt.ylim(0.1,30)
plot(mm, 10**np.array(simplefunc_linear(fitDict['barsize-vs-Mstar_lin'], mm)), 'b--', lw=1.5, zorder=3)
plot(mm, 10**np.array(simplefunc_brokenlinear(fitDict['barsize-vs-Mstar_parent_brokenlin'], mm)), 'r--', lw=2.5, zorder=4)
# plot Milky Way
semilogy(10.7, 6.0, '*', mec='g', mew=2.5, mfc='None', ms=20, alpha=0.9, zorder=5)
xlabel(xtmstar);ylabel(ytbarsize_kpc)
pu.MakeNiceLogAxes(whichAxis="y")
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
plt.savefig(plotDir+"barsize-vs-mass-with-fits_final.pdf")
# -
# ## Figure 2 -- Barsize vs logMstar using linear barsize
# plots/barsize-vs-mass-with-fit_linear.pdf
plot(s4gdata.logmstar[ii_barred_limited2], s4gdata.sma_dp_kpc2[ii_barred_limited2], 'ko', mfc='0.8',ms=5, alpha=0.7, zorder=1)
plot(s4gdata.logmstar[ii_barred_D30_m9to11], s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11], 'ko', mfc='0.5',ms=5, alpha=0.7, zorder=2)
xlim(8.5,11.5);ylim(0,14)
plot(mm, 10**np.array(simplefunc_brokenlinear(fitDict['barsize-vs-Mstar_brokenlin'], mm)), 'r--', lw=2.5, zorder=3)
# plot Milky Way
plot(10.7, 5.0, '*', mec='g', mew=2.5, mfc='None', ms=20, alpha=0.75, zorder=4)
xlabel(xtmstar);ylabel(ytbarsize_kpc)
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
plt.savefig(plotDir+"barsize-vs-mass-with-fit_linear.pdf")
# ## Figure 3 -- Residuals of barsize vs logR_e and logh, including residuals vs logMstar (4-panel)
# plots/barsize-h-Re-fits-and-residuals_4-panel -- **assembled in Adobe Illustrator**:
# 1. barsize-Re-fit.pdf
# 2. barsize-h-fit.pdf
# 3. barsize-Re-residuals-vs_logMstar.pdf
# 4. barsize-h-residuals-vs_logMstar.pdf
#
# ### Upper-left panel
# Generate binned mean values and standard deviations
logRe_bin_centers, bin_means_lin, bin_errs = GetBinMeansAndErrors(logRe_all, ii_barred_D30_m9to11_Reh, 0,1.5,0.25)
Re_bin_centers_lin = 10**logRe_bin_centers
# +
re_vect = np.arange(-0.5,2.6,0.1)
a,b = fitDict['barsize-vs-Re_lin_Reh']
barsize_predicted_lin = 10**simplefunc_linear(fitDict['barsize-vs-Re_lin_Reh'], re_vect)
re_vect_lin = 10**re_vect
loglog(s4gdata.Re_kpc[ii_barred_limited2_Reh], s4gdata.sma_dp_kpc2[ii_barred_limited2_Reh], 'ko', mfc='0.8', alpha=0.7, ms=5, zorder=1)
loglog(s4gdata.Re_kpc[ii_barred_D30_m9to11_Reh], s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11_Reh], 'ko', mfc='0.5', ms=5, alpha=0.7, zorder=2)
errorbar(Re_bin_centers_lin, bin_means_lin, yerr=bin_errs, fmt='s', mfc='None', mec='0.8', mew=1.5, ms=17, ecolor='0.7', capsize=5, elinewidth=2.5, zorder=3)
errorbar(Re_bin_centers_lin, bin_means_lin, yerr=bin_errs, fmt='s', mfc='None', mec='b', mew=2, ms=13, ecolor='b', capsize=5, elinewidth=1.5, zorder=4)
#errorbar(Re_bin_centers_lin, bin_means_lin, yerr=bin_errs, fmt='s', mfc='None', mec='0.75', mew=1.5, ms=17, ecolor='0.5', capsize=5, elinewidth=0.5)
loglog(re_vect_lin, barsize_predicted_lin, 'r--', lw=2, zorder=5)
xlim(0.3,300);ylim(0.05,50)
xlabel(r"$R_e$ [kpc]")
ylabel(ytbarsize_kpc)
pu.MakeNiceLogAxes(whichAxis="xy")
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-Re-fit.pdf")
# -
# ### Upper-right panel
# Generate binned mean values and standard deviations
logh_bin_centers, bin_means_lin, bin_errs = GetBinMeansAndErrors(np.log10(s4gdata.h_kpc), ii_barred_D30_m9to11_Reh, -0.25,1.25,0.25)
h_bin_centers_lin = 10**logh_bin_centers
# +
h_vect = np.arange(-0.5,1.31,0.1)
barsize_predicted_lin = 10**simplefunc_linear(fitDict['barsize-vs-h_lin_Reh'], h_vect)
h_vect_lin = 10**h_vect
loglog(s4gdata.h_kpc[ii_barred_limited2_Reh], s4gdata.sma_dp_kpc2[ii_barred_limited2_Reh], 'ko', mfc='0.8', alpha=0.7, ms=5, zorder=1)
loglog(s4gdata.h_kpc[ii_barred_D30_m9to11_Reh], s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11_Reh], 'ko', mfc='0.5', alpha=0.7, ms=5, zorder=2)
errorbar(h_bin_centers_lin, bin_means_lin, yerr=bin_errs, fmt='s', mfc='None', mec='0.8', mew=1.5, ms=17, ecolor='0.5', capsize=5, elinewidth=2.5, zorder=3)
errorbar(h_bin_centers_lin, bin_means_lin, yerr=bin_errs, fmt='s', mfc='None', mec='b', mew=2, ms=13, ecolor='b', capsize=5, elinewidth=1.5, zorder=4)
loglog(h_vect_lin, barsize_predicted_lin, 'r--', lw=2, zorder=5)
# 1:1 relation (a_vis = h)
#loglog([0.3,10],[0.3,10], 'g')
xlim(0.3,10);ylim(0.05,50)
xlabel(r"$h$ [kpc]")
ylabel(ytbarsize_kpc)
pu.MakeNiceLogAxes(whichAxis="xy")
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-h-fit.pdf")
# -
# ### Compute residuals to fits
# +
barsize_Re_predicted_log = simplefunc_linear(fitDict['barsize-vs-Re_lin_Reh'], logRe_all)
residuals_Re_log = logbarsize_all - barsize_Re_predicted_log
barsize_h_predicted_log = simplefunc_linear(fitDict['barsize-vs-h_lin_Reh'], logh_all)
residuals_h_log = logbarsize_all - barsize_h_predicted_log
# -
# ### Lower-left panel
# +
s4gxx_03, s4gyy_03 = GetSortedLoess1D(s4gdata.logmstar[ii_barred_limited2_Reh], residuals_Re_log[ii_barred_limited2_Reh], frac=0.3)
plot(s4gdata.logmstar[ii_barred_D30_m9to11_Reh], residuals_Re_log[ii_barred_D30_m9to11_Reh], 'ko', ms=5)
plot(s4gxx_03, s4gyy_03, 'r-', lw=2)
xlim(9,11.0);ylim(-0.8,0.8)
xlabel(xtmstar);ylabel(ytresid)
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-Re-residuals_vs_logMstar.pdf")
# -
# ### Lower-right panel
# +
s4gxx_03, s4gyy_03 = GetSortedLoess1D(s4gdata.logmstar[ii_barred_limited2_Reh], residuals_h_log[ii_barred_limited2_Reh], frac=0.3)
plot(s4gdata.logmstar[ii_barred_D30_m9to11_Reh], residuals_h_log[ii_barred_D30_m9to11_Reh], 'ko', ms=5)
plot(s4gxx_03, s4gyy_03, 'r', lw=2)
xlim(9,11.);ylim(-0.8,0.8)
xlabel(xtmstar);ylabel(ytresid)
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-h-residuals_vs_logMstar.pdf")
# -
# ## Figure 4 -- Barsize vs gas mass fraction
# plots/barsize-vs-logfgas.pdf
# +
s4gxx, s4gyy = GetSortedLoess1D(s4gdata.logfgas[ii_barred_D30_m9to11], np.log10(s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11]), frac=0.3)
s4gyy = 10**s4gyy
semilogy(s4gdata.logfgas[ii_barred_D30_m9to11], s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11], 'o', color='0.5',mew=0.6, ms=5, alpha=0.7)
semilogy(s4gdata.logfgas[ii_barred_D30_m9to11_Reh], s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11_Reh], 'o', color='k',mew=0.6, ms=5, alpha=0.7)
semilogy(s4gxx,s4gyy, 'r')
xlim(-3.5,0.5);ylim(0.1,30)
xlabel(xtfgas)
ylabel(ytbarsize_kpc)
pu.MakeNiceLogAxes(whichAxis="y")
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-vs-logfgas.pdf")
# -
# ## Figure 5 -- Residuals of barsize vs gas mass fraction (2-panel)
# plots/barsize-residuals-vs-logfgas -- **assembled in Adobe Illustrator**:
# 1. barsize-ReMstar-residuals_vs_logfgas.pdf
# 2. barsize-hMstar-residuals_vs_logfgas.pdf
#
# ### Compute residuals to fits
# +
# data vectors for Main Barred Spiral Sample
xxReh_Re = np.log10(s4gdata.Re_kpc[ii_barred_D30_m9to11_Reh])
xxReh_h = np.log10(s4gdata.h_kpc[ii_barred_D30_m9to11_Reh])
xxReh_mstar = s4gdata.logmstar[ii_barred_D30_m9to11_Reh]
yyReh = np.log10(s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11_Reh])
# compute predicted bar sizes, then residuals, for fit of barsize-vs-Re+Mstar
barsize_predicted_log = fitting_barsizes.fmulti_lin_brokenlin((xxReh_Re,xxReh_mstar), *fitDict['barsize-vs-Re+Mstar_Reh'])
resid_logbarsize_ReMstar = yyReh - barsize_predicted_log
# compute predicted bar sizes, then residuals, for fit of barsize-vs-h+Mstar
barsize_predicted_log = fitting_barsizes.fmulti_lin_brokenlin((xxReh_h,xxReh_mstar), *fitDict['barsize-vs-h+Mstar_Reh'])
resid_logbarsize_hMstar = yyReh - barsize_predicted_log
# -
# ### Plot: Left panel
# +
s4gxx_03, s4gyy_03 = GetSortedLoess1D(s4gdata.logfgas[ii_barred_D30_m9to11_Reh], resid_logbarsize_ReMstar, frac=0.3)
plot(s4gdata.logfgas[ii_barred_D30_m9to11_Reh], resid_logbarsize_ReMstar, 'ko', ms=5)
plot(s4gxx_03, s4gyy_03, 'r', lw=1.5)
axhline(0, color='0.5')
xlim(-2.5,0.5);ylim(-0.8,0.8)
xlabel(xtfgas);ylabel(ytresid)
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-ReMstar-residuals_vs_logfgas.pdf")
# -
# ### Plot: Right panel
# +
s4gxx_03, s4gyy_03 = GetSortedLoess1D(s4gdata.logfgas[ii_barred_D30_m9to11_Reh], resid_logbarsize_hMstar, frac=0.3)
plot(s4gdata.logfgas[ii_barred_D30_m9to11_Reh], resid_logbarsize_hMstar, 'ko', ms=5)
plot(s4gxx_03, s4gyy_03, 'r', lw=1.5)
axhline(0, color='0.5')
xlim(-2.5,0.5);ylim(-0.8,0.8)
xlabel(xtfgas);ylabel(ytresid)
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-hMstar-residuals_vs_logfgas.pdf")
# -
# ## Figure 6 -- Barsize vs Hubble type
# plots/barsize-vs-T.pdf
# +
s4gxx, s4gyy = GetSortedLoess1D(s4gdata.t_leda[ii_barred_D30_m9to11], np.log10(s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11]), frac=0.3)
s4gyy = 10**s4gyy
semilogy(s4gdata.t_leda[ii_barred_limited2], s4gdata.sma_dp_kpc2[ii_barred_limited2], 'ko', mfc='0.8', ms=5, alpha=0.7)
semilogy(s4gdata.t_leda[ii_barred_D30_m9to11], s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11], 'ko', mfc='0.5', ms=5, alpha=0.7)
semilogy(s4gxx,s4gyy, 'r', lw=1.5)
xlim(-1,11);ylim(0.1,30)
xlabel(r'Hubble Type $T$')
ylabel(ytbarsize_kpc)
pu.MakeNiceLogAxes(whichAxis="y")
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-vs-T.pdf")
# -
# ## Figure 7 -- Residuals of barsize vs Hubble type (2-panel)
# plots/barsize-residuals-vs-T_2panel -- **assembled in Adobe Illustrator**:
# 1. barsize-ReMstar-residuals_vs_T.pdf
# 2. barsize-hMstar-residuals_vs_T.pdf
# Note that we can re-use the residuals to the barsize-vs-logMstar+Re/h fits from Section 5 above.
# ### Left panel
# +
htype_Reh = s4gdata.t_leda[ii_barred_D30_m9to11_Reh]
s4gxx, s4gyy = GetSortedLoess1D(htype_Reh, resid_logbarsize_ReMstar, frac=0.3)
plot(htype_Reh, resid_logbarsize_ReMstar, 'ko', ms=5)
plot(s4gxx, s4gyy, 'r-', lw=1.5)
axhline(0,color='0.5')
xlim(-1,11);ylim(-0.8,0.8)
xlabel(r"Hubble type $T$");ylabel(ytresid)
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-ReMstar-residuals_vs_T.pdf")
# -
# ### Right panel
# +
htype_Reh = s4gdata.t_leda[ii_barred_D30_m9to11_Reh]
s4gxx, s4gyy = GetSortedLoess1D(htype_Reh, resid_logbarsize_hMstar, frac=0.3)
plot(htype_Reh, resid_logbarsize_hMstar, 'ko', ms=5)
plot(s4gxx, s4gyy, 'r-', lw=1.5)
axhline(0,color='0.5')
xlim(-1,11);ylim(-0.8,0.8)
xlabel(r"Hubble type $T$");ylabel(ytresid)
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"barsize-hMstar-residuals_vs_T.pdf")
# -
r1,p1 = scipy.stats.spearmanr(htype_Reh, resid_logbarsize_ReMstar)
print("Correlation of barsize-Mstar+Re fit residuals vs T for logMstar = 9--11 barred galaxies: r = %g, P = %g" % (r1,p1))
r2,p2 = scipy.stats.spearmanr(htype_Reh, resid_logbarsize_hMstar)
print("Correlation of barsize-Mstar+h fit residuals vs T for logMstar = 9--11 barred galaxies: r = %g, P = %g" % (r2,p2))
# ## Figure 8 -- Residuals of Re-Mstar or h-Mstar vs bar sizes
# [~/Documents/Working/Projects/Project_BarSizes/notebooks/barsizes_galaxy_scaling_relations.ipynb]
#
# plots/size-vs-mstar_residuals-vs-barsize.pdf
#
# ### Compute residuals of fits to log(Mstar) vs R_e or vs h
# +
# broken-linear fit to Main Spiral Sample: log R_e vs logMstar
predRe = simplefunc_brokenlinear(fitDict['Re-vs-Mstar_Reh'], s4gdata.logmstar)
residRe = logRe_all - predRe
# broken-linear fit to Main Spiral Sample: log h vs logMstar
predh = simplefunc_brokenlinear(fitDict['h-vs-Mstar_Reh'], s4gdata.logmstar)
residh = logh_all - predh
# compute simple linear fits to residuals as a function of barsize
def flin( x, a, b ):
return a + b*x
p0_lin = [-2, 0.3]
pp_lin_reresid, pcov = scipy.optimize.curve_fit(flin, logbarsize_all[ii_barred_D30_m9to11_Reh], residRe[ii_barred_D30_m9to11_Reh], p0=p0_lin)
pp_lin_hresid, pcov = scipy.optimize.curve_fit(flin, logbarsize_all[ii_barred_D30_m9to11_Reh], residh[ii_barred_D30_m9to11_Reh], p0=p0_lin)
# -
# ### Make 2-panel plot
# +
xx = np.arange(-1.5,1.6,0.1)
xx_lin = 10**xx
fitline_Re = pp_lin_reresid[0] + pp_lin_reresid[1]*xx
fitline_h = pp_lin_hresid[0] + pp_lin_hresid[1]*xx
f,(ax1,ax2) = subplots(1,2, figsize=(15,5))
ax1.semilogx(s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11_Reh], residRe[ii_barred_D30_m9to11_Reh], 'o', color='0.5', ms=3.5)
ax1.semilogx(xx_lin, fitline_Re, 'k--')
ax1.axhline(0, lw=0.5)
ax1.set_ylim(-1,2.0)
ax1.set_xlim(0.2,20)
ax1.set_xlabel(ytbarsize_kpc)
ax1.set_ylabel(ytresid_Re, fontsize=17)
pu.MakeNiceLogAxes(whichAxis="x", axisObj=ax1)
ax2.semilogx(s4gdata.sma_dp_kpc2[ii_barred_D30_m9to11_Reh], residh[ii_barred_D30_m9to11_Reh], 'o', color='0.5', ms=3.5)
ax2.semilogx(xx_lin, fitline_h, 'k--')
ax2.axhline(0, lw=0.5)
ax2.set_ylim(-1,2.0)
ax2.set_xlim(0.2,20)
ax2.set_xlabel(ytbarsize_kpc)
ax2.set_ylabel(ytresid_h, fontsize=17)
pu.MakeNiceLogAxes(whichAxis="x", axisObj=ax2)
if savePlots:
plt.savefig(plotDir+"size-vs-mstar_residuals-vs-barsize.pdf")
# -
# ## Figure 9 -- Scaling relations: R_e or h vs logMstar: barred, unbarred, all (2-panel)
# plots/size-vs-mstar_barred-v-unbarred-loess_2panel.pdf --> size-vs-mstar_barred-v-unbarred-loess_2panel_tweaked.pdf
#
# This is generated in this notebook as a 2-panel figure; the bounding-box/canvas is then
# tweaked in Adobe Illustrator.
# +
s4gxx, loessFit_Re = GetSortedLoess1D(s4gdata.logmstar[ii_all_Reh], np.log10(s4gdata.Re_kpc[ii_all_Reh]), frac=0.3)
s4gxx_barred, loessFit_Re_barred = GetSortedLoess1D(s4gdata.logmstar[ii_barred_Reh], np.log10(s4gdata.Re_kpc[ii_barred_Reh]), frac=0.3)
s4gxx_unbarred, loessFit_Re_unbarred = GetSortedLoess1D(s4gdata.logmstar[ii_unbarred_Reh], np.log10(s4gdata.Re_kpc[ii_unbarred_Reh]), frac=0.3)
loessFit_Re = 10**loessFit_Re
loessFit_Re_barred = 10**loessFit_Re_barred
loessFit_Re_unbarred = 10**loessFit_Re_unbarred
s4gxx, s4gyy = GetSortedLoess1D(s4gdata.logmstar[ii_all_Reh], logh_all[ii_all_Reh], frac=0.3)
loessFit_h = s4gyy
s4gxx_barred, s4gyy_barred = GetSortedLoess1D(s4gdata.logmstar[ii_barred_Reh], logh_all[ii_barred_Reh], frac=0.3)
loessFit_h_barred = s4gyy_barred
s4gxx_unbarred, s4gyy_unbarred = GetSortedLoess1D(s4gdata.logmstar[ii_unbarred_Reh], logh_all[ii_unbarred_Reh], frac=0.3)
loessFit_h_unbarred = s4gyy_unbarred
loessFit_h = 10**loessFit_h
loessFit_h_barred = 10**loessFit_h_barred
loessFit_h_unbarred = 10**loessFit_h_unbarred
# +
f,(ax1,ax2) = subplots(1,2, figsize=(15,5))
ax1.semilogy(s4gdata.logmstar, s4gdata.Re_kpc, 'o', mec='k', mfc="None", ms=3.5)
ax1.semilogy(s4gdata.logmstar[ii_barred], s4gdata.Re_kpc[ii_barred], 'o', color="0.5", ms=3.5)
ax1.plot(s4gxx_barred, loessFit_Re_barred, 'b', lw=2, label='barred')
ax1.plot(s4gxx_unbarred, loessFit_Re_unbarred, 'g', lw=2, label='unbarred')
ax1.plot(s4gxx, loessFit_Re, 'r-', lw=3, label='all', alpha=0.3)
ax1.set_xlim(7,11.5);ax1.set_ylim(0.1,50)
ax1.legend()
pu.MakeNiceLogAxes('y', axisObj=ax1)
ax1.set_xlabel(xtmstar, fontsize=16); ax1.set_ylabel(xtRe_kpc, fontsize=16)
ax2.semilogy(s4gdata.logmstar, s4gdata.h_kpc, 'o', mfc="None", mec="k", ms=3.5)
ax2.semilogy(s4gdata.logmstar[ii_barred], s4gdata.h_kpc[ii_barred], 'o', color="0.5", ms=3.5)
ax2.plot(s4gxx_barred, loessFit_h_barred, 'b', lw=2, label='barred')
ax2.plot(s4gxx_unbarred, loessFit_h_unbarred, 'g', lw=2, label='unbarred')
ax2.plot(s4gxx, loessFit_h, 'r-', lw=3, label='all', alpha=0.3)
ax2.set_xlim(7,11.5);ax2.set_ylim(0.1,20)
ax2.legend()
pu.MakeNiceLogAxes(whichAxis="y", axisObj=ax2)
ax2.set_xlabel(xtmstar, fontsize=16); ax2.set_ylabel(xth_kpc, fontsize=16)
if savePlots:
savefig(plotDir+"size-vs-mstar_barred-v-unbarred-loess_2panel.pdf")
# -
# ## Figure A1 -- Scaling relations: R_e or h vs logMstar: all (2-panel) [Appendix]
# plots/size-vs-mstar-with-fits_2panel.pdf
# +
# broken-linear fit to Main Spiral Sample: log R_e vs logMstar
#a1, b1, x_brk, b2 = pp_brokenlin_revsmstar
xx = np.arange(7,12,0.05)
predRe = 10**simplefunc_brokenlinear(fitDict['Re-vs-Mstar_Reh'], xx)
# broken-linear fit to Main Spiral Sample: log h vs logMstar
#a1, b1, x_brk, b2 = pp_brokenlin_hvsmstar
xx = np.arange(7,12,0.05)
predh = 10**simplefunc_brokenlinear(fitDict['h-vs-Mstar_Reh'], xx)
# make the plots
f,(ax1,ax2) = subplots(1,2, figsize=(15,5))
ax1.semilogy(s4gdata.logmstar, s4gdata.Re_kpc, 'o', mfc='None', mec='0.1', ms=3.5)
ax1.semilogy(s4gdata.logmstar[ii_all_Reh_m9to11], s4gdata.Re_kpc[ii_all_Reh_m9to11], 'o', color="k", ms=3.5)
ax1.plot(s4gxx, loessFit_Re, 'r-', lw=3, label='all', alpha=0.3)
ax1.semilogy(xx, predRe, 'c--', lw=2.5)
ax1.set_xlim(7,11.5);ax1.set_ylim(0.1,50)
pu.MakeNiceLogAxes('y', axisObj=ax1)
ax1.set_xlabel(xtmstar, fontsize=16); ax1.set_ylabel(xtRe_kpc, fontsize=16)
ax2.semilogy(s4gdata.logmstar, s4gdata.h_kpc, 'o', color='0.5', ms=3.5)
ax2.semilogy(s4gdata.logmstar[ii_all_Reh_m9to11], s4gdata.h_kpc[ii_all_Reh_m9to11], 'o', color="k", ms=3.5)
ax2.plot(s4gxx, loessFit_h, 'r-', lw=3, label='all', alpha=0.3)
ax2.semilogy(xx, predh, 'c--', lw=2.5)
ax2.set_xlim(7,11.5);ax2.set_ylim(0.1,20)
pu.MakeNiceLogAxes(whichAxis="y", axisObj=ax2)
ax2.set_xlabel(xtmstar, fontsize=16); ax2.set_ylabel(xth_kpc, fontsize=16)
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
savefig(plotDir+"size-vs-mstar-with-fits_2panel.pdf")
# -
# ## Figure A2 -- Gas mass ratio vs logMstar
# +
plot(s4gdata.logmstar[ii_spirals], s4gdata.logfgas[ii_spirals], 'o', mfc='0.6', mec='None',ms=5, alpha=0.8, zorder=-1)
plot(s4gdata.logmstar[ii_all_D30_m9to11], s4gdata.logfgas[ii_all_D30_m9to11], 'ko', mec='None',ms=5, alpha=0.8, zorder=-1)
xlim(8,11.5);ylim(-3.5,1.5)
xlabel(xtmstar)
ylabel(r'Gas mass ratio [$\log \, (M_{\rm HI} / M_{\odot})]$')
# push bottom of plot upwards so that x-axis label isn't clipped in PDF output
plt.subplots_adjust(bottom=0.14)
if savePlots:
savefig(plotDir+"logfgas-vs-mstar.pdf")
| barsizes_figures_for_paper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import sklearn
from scipy.stats.stats import pearsonr
'''Notes predict target. 1 means borrower had issues making payment 0 all other predict if borrower
will have problems paying back the loan. Visual data scan:
def_60_cnt_social bureau (loan payment problems in area)
Step 1 Find correcation between target and other variables
'''
df = pd.read_csv('../data/application_train.csv')
# df['DEF_30_CNT_SOCIAL_CIRCLE']
df.head()
# +
# TODO recursive way to clean up data
def normalize_boolean(value):
if value == type(bool):
return value * 1
def replace_yes_no(value):
if value == 'Y':
return 1
elif value == 'N':
return 0
# -
# Visual check for seemingly useless column lets drop
drop = [col for col in df if col.startswith('FLAG_DOCUMENT_')]
df = df.drop(drop, axis=1)
df.head()
# Manual clean up bad! Keep for reference
df['FLAG_OWN_REALTY'] = df['FLAG_OWN_REALTY'].map({'Y':1, 'N':0})
df['FLAG_OWN_CAR'] = df['FLAG_OWN_CAR'].map({'Y':1, 'N':0})
df['CODE_GENDER'] = df['CODE_GENDER'].map({'M':1, 'F':0})
df.head()
# +
# check for non numerical values
#df.select_dtypes(exclude=[np.number])
#check for dtypes in table
#g = df.columns.to_series().groupby(df.dtypes).groups
df_build_in_cat = pd.get_dummies(df)
df_build_in_cat.head()
#df_test = df_build_in_cat.fillna(value=-999999, inplace=True)
# try build in catergorizing of data for 1 column
# df['NAME_CONTRACT_TYPE'].astype('category')
# dtype(df['NAME_CONTRACT_TYPE'])
# +
# Check for machine unfriendly values
df_build_in_cat.select_dtypes(exclude=[np.number])
print(df_build_in_cat.isnull().values.any())
df_build_in_cat.fillna(value=-999999, inplace=True)
if df_build_in_cat.select_dtypes(include=[object]).empty:
print('No NaNs')
df_build_in_cat.head()
# -
# X features, y as label
# y = df_build_in_cat.target
# X = np.array(df_build_in_cat.drop(['TARGET', 'SK_ID_CURR'], axis=1), dtype='f')
# y what are we predicting
y = np.array(df_build_in_cat['TARGET'])
# X what are we using to predict
X = np.array(df_build_in_cat.drop(['TARGET', 'SK_ID_CURR'], axis=1), dtype='f')
# Check np array for evil value
print('Do we have any NaN\'s: {}'.format(np.any(np.isnan(X))))
print('Make sure we have no infinity? {}'.format(np.all(np.isfinite(X))))
# +
from sklearn.tree import DecisionTreeRegressor
tree_regressor_model = DecisionTreeRegressor()
tree_regressor_model.fit(X, y)
# -
tree_regressor_model.predict(X)
from sklearn.metrics import mean_absolute_error
predictions = tree_regressor_model.predict(X)
mean_absolute_error(y, predictions)
'''
Merge them tables
'''
df_b_app =pd.merge(df_bureau, df, on='SK_ID_CURR')
'''
Normalize Bureau datas TODO
'''
'''
Check for correlation
'''
df[(df.TARGET == 1)][['TARGET', 'FLAG_OWN_REALTY']]
df[(df.TARGET == 1)][['TARGET','DEF_30_CNT_SOCIAL_CIRCLE']]
drop_na_df = df.dropna()
print(pearsonr(drop_na_df.TARGET, drop_na_df.DEF_30_CNT_SOCIAL_CIRCLE))
| sz/ZabaPredicts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
import urllib.request
import time
from bs4 import BeautifulSoup
import pandas as pd
import numpy as np
businessName=[]
url = 'https://members.hamptonroadschamber.com/sbaweb/members/members.asp?&id=1&wpid=-101&kwaction=1&kwwhere=[NEW]&newsession=FALSE&sid=119858852'
response = requests.get(url)
soup = BeautifulSoup(response.text, "html.parser")
soup.findAll('a')
for a in soup.findAll('a',href=True, attrs={'class': 'sbaLinkMenu'}):
name = a.find('div', attrs = {'class': 'body'})
businessName.append(name.text)
df = pd.DataFrame({'BusinessName': businessName})
df.to_csv('businessName.csv', index = False, encoding = 'utf-8')
# +
import requests
import urllib.request
from bs4 import BeautifulSoup
from selenium import webdriver
import pandas as pd
import numpy as np
driver = webdriver.chrome('C:\Program Files (x86)\Google\Chrome\Application')
businessName=[]
url = 'https://members.hamptonroadschamber.com/sbaweb/members/members.asp?&id=1&wpid=-101&kwaction=1&kwwhere=[NEW]&newsession=FALSE&sid=119858852'
response = requests.get(url)
soup = BeautifulSoup(response.text, "html.parser")
for a in soup.findAll('a',href=True, attrs={'class': 'sbaLinkMenu'}):
name = a.find('div', attrs = {'class': 'body'})
businessName.append(name.text)
df = pd.DataFrame({'BusinessName': businessName})
df.to_csv('businessName.csv', index = False, encoding = 'utf-8')
| Project-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + Collapsed="false"
import numpy as np
import math
# + Collapsed="false"
output = np.arange(0, 128, 1)
target = np.arange(0, 128, 1)
# + Collapsed="false"
output.max()
# + Collapsed="false"
bce = np.empty((128,128), dtype=np.float)
eps = 1e-6
for x in output:
for y in target:
out = float(x)/127
tar = float(y)/127
temp = tar * math.log(out+eps)
temp += (1 - tar) * math.log(1 - out+eps)
bce[y,x] = -temp
bce.shape
# + Collapsed="false"
bce
# + Collapsed="false"
np.savetxt('bce', bce, delimiter=',')
# + Collapsed="false"
# Compute cross entropy from probabilities.
epsilon = 1e-13 # to avoid log0
target = 1
output = 0.3
bce = target * math.log(output+epsilon)
bce += (1 - target) * math.log(1 - output+epsilon)
-bce
# + Collapsed="false"
| examples/2D/BinaryCrossEntropy on continuous probability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Praxis-bs/DEMD/blob/main/Linear_Regression_Wine_Analysis_Predict.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ikOWiOLJGEfQ"
# <br>
#
#
# <hr>
#
# [<NAME>](http://www.linkedin.com/in/prithwis)<br>
# + [markdown] id="ndjcQmR_GK52"
# #Linear Regression - Wine Analysis - Predict
# based on https://medium.freecodecamp.org/a-beginners-guide-to-training-and-deploying-machine-learning-models-using-python-48a313502e5a <br>
# This notebook needs the [Training Notebook](https://github.com/Praxis-bs/DEMD/blob/64989c5b5de8f59637c123bd8867a02be873cf7d/Linear_Regression_Wine_Analysis_Train.ipynb)
# + id="ejVRNvtW068x"
import pickle
from sklearn import linear_model
# + colab={"base_uri": "https://localhost:8080/"} id="8iyzk6t91rfa" outputId="f5f440e9-6e69-416b-f478-4d696fae62ec"
#get the model
# https://drive.google.com/file/d/1IsVRzeoP9E-C9O0-e2zESbwNHjbOcZId/view?usp=sharing
# !gdown https://drive.google.com/uc?id=1IsVRzeoP9E-C9O0-e2zESbwNHjbOcZId
# + id="urho2_Ql0683"
model = pickle.load(open("lrw-model.pkl","rb"))
# + id="MUf3_b7U0684"
feature_array = [7.4,0.66,0,1.8,0.075,13,40,0.9978,3.51,0.56,9.4]
# + id="3iHVItln0684" colab={"base_uri": "https://localhost:8080/"} outputId="bc24f143-bc46-4ea9-9f26-e4bed9acb686"
#our model rates the wine based on the input array
prediction = model.predict([feature_array]).tolist()
print(prediction)
# + [markdown] id="_00MgHT9G6RG"
# #Chronobooks <br>
# <hr>
# Chronotantra and Chronoyantra are two science fiction novels that explore the collapse of human civilisation on Earth and then its rebirth and reincarnation both on Earth as well as on the distant worlds of Mars, Titan and Enceladus. But is it the human civilisation that is being reborn? Or is it some other sentience that is revealing itself.
# If you have an interest in AI and found this material useful, you may consider buying these novels, in paperback or kindle, from [http://bit.ly/chronobooks](http://bit.ly/chronobooks)
| Linear_Regression_Wine_Analysis_Predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Introduction
#
# This notebook demonstrates how to plot time series from the UKESM1 Geoengineering simulations
# #### Firstly, import packages and define functions for calculations
'''Import packages for loading data, analysing, and plotting'''
import xarray
import matplotlib
import numpy
import cftime
# %matplotlib inline
# # Plotting time series computed by the CVDP package
#
# The Climate Variability Diagnostics Package computes a whole bunch of time series. Some of then, such as the Nino3.4 index, need to be treated with caution. But some are really useful.
# *Use this cells if you are using the UCL JupyterHub. If you are running this on your own laptop use the cell at the very end instead.*
# Specify the filenames for the 3 scenarios
filenameG6='~/geog0121_shared_space/PMIP_GeoMIP_summary_files/UKESM1_G6sulfur_1850-1900.cvdp_data.1850-2100.nc'
filename245='~/geog0121_shared_space/UKESM1_summary_files/UKESM1_ssp245_1851-1900.cvdp_data.1850-2100.nc'
filename585='~/geog0121_shared_space/UKESM1_summary_files/UKESM1_ssp585_1851-1900.cvdp_data.1850-2100.nc'
# First, let us remind ourselves about the difference between the two standard scenarios - SSP585 and SSP245.
# Here we will plot the global average temperature over land.
# +
#Start with SSP245
# open the file to get the data
ssp245=xarray.open_dataset(filename245)
# Let smooth this to create a 5-year mean
glt_245=ssp245.ipcc_GLOBAL_lnd_tas
glt_245_smoothed=glt_245.rolling(time=60,center=True).mean() #take the 5-year running mean (i.e. rolling over 60 months)
# create a plot of the variable called ipcc_GLOBAL_lnd_tas
glt_245_smoothed.plot() #by default the first line is blue
#Then move onto SSP585
# open the file to get the data
ssp585=xarray.open_dataset(filename585)
glt_585=ssp585.ipcc_GLOBAL_lnd_tas
glt_585_smoothed=glt_585.rolling(time=60,center=True).mean() #take the 5-year running mean (i.e. rolling over 60 months)
# create a plot of the variable called ipcc_GLOBAL_all_tas
glt_585_smoothed.plot(color='firebrick') #by default the second line is orange
# -
# #### Introducing the G6sulfur scenario
# So the SSP585 scenario results in the land surface warming by approaching 10oC above preindustrial, which is not pleasant. Perhaps we might want to intervene and deploy some technology to mask out the effect of the increasing carbon dioxide levels.
# +
#Now lets look at the G6sulfur experiment...
# open the file to get the data
G6=xarray.open_dataset(filenameG6)
# Let's smooth this to create a 5-year mean
glt_G6=G6.ipcc_GLOBAL_lnd_tas
glt_G6_smoothed=glt_G6.rolling(time=60,center=True).mean() #take the 5-year running mean (i.e. rolling over 60 months)
# create a plot of the variable called nino34.
glt_245_smoothed.plot() #by default the first line is blue
glt_585_smoothed.plot(color='firebrick')
glt_G6_smoothed.plot(color='green') #by default the second line is orangeglt_G6_smoothed.plot(color='green')
# -
# So injecting sulphur into the atmosphere (a bit like artificial volcanoes) can successfully change the temperature trajectory.
#
# #### What about precipitation though?
# We can also look at other properties of the climate system, to see if it masks all the impacts of climate change though. Let us look at regional rainfall. Here will pick the South East Asian summer rainfall, which is roughly akin to the summer monsoon.
# +
# Let smooth this to create a 5-year mean
sea_245=ssp245.ipcc_SEA_lnd_pr #load the monthly time series
sea_245_seasonal=sea_245.resample(time="QS-DEC").mean()
sea_sm245=sea_245_seasonal[2::4]
sea_sm245_smoothed=sea_sm245.rolling(time=10,center=True).mean() #take the 5-year running mean (i.e. rolling over 60 months)
# Let smooth this to create a 5-year mean
sea_585=ssp585.ipcc_SEA_lnd_pr #load the monthly time series
sea_585_seasonal=sea_585.resample(time="QS-DEC").mean()
sea_sm585=sea_585_seasonal[2::4]
sea_sm585_smoothed=sea_sm585.rolling(time=10,center=True).mean() #take the 5-year running mean (i.e. rolling over 60 months)
# Let smooth this to create a 5-year mean
sea_G6=G6.ipcc_SEA_lnd_pr #load the monthly time series
sea_G6_seasonal=sea_G6.resample(time="QS-DEC").mean()
sea_smG6=sea_G6_seasonal[2::4]
sea_smG6_smoothed=sea_smG6.rolling(time=10,center=True).mean() #take the 5-year running mean (i.e. rolling over 60 months)
sea_sm245_smoothed.plot() #by default the first line is blue
sea_sm585_smoothed.plot(color='firebrick')
sea_smG6_smoothed.plot(color='green') #by default the second line is orangeglt_G6_smoothed.plot(color='green')
start_date=cftime.Datetime360Day(2010,1,1,0,0,0)
end_date=cftime.Datetime360Day(2100,12,30,0,0,0)
# alter the limits of the axes
matplotlib.pyplot.axis([start_date,end_date, -1.5, 0.5]) #[xmin, xmax, ymin, ymax]; Note that the years are strings
# add a grid in the background
matplotlib.pyplot.grid(True)
#Finally save the figure to your current directory...
#plt.savefig("nino34_timeseries_plot.pdf")
# -
# Here we can see that the geoengineering does not ameliorate the reduction in rainfall over Indonesia. In fact, it makes it even worse.
# ## Alternative location code.
# **If you are not using the UCL JupyterHub, you will not be able to access the shared data files. You will need to download versions yourselves. This can be done using the following cell instead.**
# +
'''Select file and variable names'''
#first of all specify some names
reference_period='1851-1900'
ssp_names=["ssp126", "ssp245", "ssp370","ssp585"]
directory='Data'
# determine the filenames for the 3 scenarios
filename245='%s/UKESM1_%s_%s.cvdp_data.1850-2100.nc'%(directory,ssp_names[1],reference_period)
filename585='%s/UKESM1_%s_%s.cvdp_data.1850-2100.nc'%(directory,ssp_names[3],reference_period)
filenameG6='%s/UKESM1_%s_%s.cvdp_data.1850-2100.nc'%(directory,"G6sulfur",reference_period)
#Download the files if they are not already here
import os
import urllib
if not os.path.isfile(filename245):
# !mkdir Data
urllib.request.urlretrieve("http://www2.geog.ucl.ac.uk/~ucfaccb/geog0121/downloads/UKESM1_ssp245_1851-1900.cvdp_data.1850-2100.nc", filename245)
if not os.path.isfile(filename585):
urllib.request.urlretrieve("http://www2.geog.ucl.ac.uk/~ucfaccb/geog0121/downloads/UKESM1_ssp585_1851-1900.cvdp_data.1850-2100.nc", filename585)
if not os.path.isfile(filenameG6):
urllib.request.urlretrieve("http://www2.geog.ucl.ac.uk/~ucfaccb/geog0121/downloads/UKESM1_G6sulfur_1850-1900.cvdp_data.1850-2100.nc", filenameG6)
| practicals/UKESM1_in_Python/GeoMIP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="IOp0AJeskbEl"
# # Binary Representation_answers
# + [markdown] colab_type="text" id="FtRxid4BkbEt"
# Question 1:
#
# What is the binary representation of 12?
# + colab={} colab_type="code" id="lAP7izWJkbEv" outputId="325df176-cdb0-48b8-f6c4-1f7843037bf2"
bin(12)
# + [markdown] colab_type="text" id="jVp6xCxgkbE7"
# Question 2:
#
# What is the binary representation of 4?
# + colab={} colab_type="code" id="xePUwPr6kbE8" outputId="93c2788d-faa3-41ef-bb7c-15f9513b462f"
bin(4)
# + [markdown] colab_type="text" id="TYTT7AYpkbFA"
# Question 3:
#
# Using bitwise OR, find the number which combines the bits of 12 and 4 and its binary representation.
# + colab={} colab_type="code" id="ybfdO5fkkbFB" outputId="ee3c51ae-d18a-406e-d212-5a92324e2715"
12 | 4
# + colab={} colab_type="code" id="zigVWjHtkbFI" outputId="60863137-18a6-4629-b993-2afd3958df0e"
bin(12 | 4)
| Variable_Operations/Binary_representation_answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Snapshot Explain
#
# This notebook shows how to run a Snapshot Explain operation with the minimal steps and a simple query.
#
# In this notebook...
# * [Dependencies and Initialisation](#dependencies-and-initialisation)
# * [The Where Statement](#the-where-statement)
# * [Running the Explain Operation](#running-the-explain-operation)
# * [Next Steps](#next-steps)
# ## Dependencies and Initialisation
# Import statements and environment initialisation using the package `dotenv`. More details in the [Configuration notebook](0.2_configuration.ipynb).
from factiva.news import Snapshot
from dotenv import load_dotenv
load_dotenv()
print('Done!')
# ## The Where Statement
#
# This notebook uses a simple query for illustration purposes. For more tips about queries, or guidance on how to build complex or large queries, checkout the [query reference](2.1_complex_large_queries.ipynb) notebook.
# +
# Industry i3432 is for Batteries
where_statement = (
r" publication_datetime >= '2016-01-01 00:00:00' "
r" AND LOWER(language_code) IN ('en', 'de', 'fr') "
r" AND REGEXP_CONTAINS(industry_codes, r'(?i)(^|,)(i3432)($|,)') "
)
s = Snapshot(query=where_statement)
# -
# ## Running the Explain Operation
#
# This operation returns the number of documents matching the provided query in the Factiva Analytics archive.
#
# The goal of this operation is to have a rough idea of the document volume. When used iteratively, helps deciding on the used criteria to add/delete/modify the criteria to verify the impact on the matching items.
#
#
# The `<Snapshot>.process_explain()` function directly returns this value. If a more manual process is required (send job, monitor job, get results), please see the [detailed package documentation](https://factiva-news-python.readthedocs.io/).
# %%time
s.process_explain()
print(f'Explain operation ID: {s.last_explain_job.job_id}')
print(f'Document volume estimate: {s.last_explain_job.document_volume}')
# ## Getting Explain Samples
# As an extension of the Explain operation, it is possible to request a set of random article metadata samples matching the Explain criteria. The main requirement in this case is just using the previously obtained Explain Job ID. It uses the ID from the `last_explain_job` instance within the `Snapshot` instance.
#
# The operation `get_explain_job_samples(num_samples=10)` returns a Pandas DataFrame with the metadata content from a random selection of items. It accepts the parameter `num_samples` that is an integer between `1` and `100`.
samples = s.get_explain_job_samples(num_samples=10)
# The following code shows the list of columns that can be used for a quick analysis.
samples.columns
# Displaying a subset of the content.
samples[['an', 'source_name', 'title', 'word_count']]
# # Next Steps
#
# * Run an [analytics](1.5_snapshot_analytics.ipynb) to get a detailed time-series dataset of the estimates.
# * Run an [extraction](1.6_snapshot_extraction.ipynb) and download the matching content.
# * Fine-tune the query by adding/modifying the query criteria (where_statement) according to the [query reference](2.1_complex_large_queries.ipynb).
| 1.4_snapshot_explain.ipynb |