code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] jupyter={"outputs_hidden": true}
# # Maximal Coverage Location Problem
#
# *Authors:* [<NAME>](https://github.com/gegen07), [<NAME>](https://github.com/jGaboardi), [<NAME>](https://github.com/ljwolf), [<NAME>](https://github.com/qszhao)
#
# LSCP try to minimize the amount of facilities candidate sites in a maximum service standard but then arise another problem: the budget. Sometimes it requires many facilities sites to reach a complete coverage, and there are circumstances when the resources are not available and it's plausible to know how much coverage we can reach using a exact number of facilities. MCLP class try to solve this problem:
#
# _Maximize the amount of demand covered within a maximal service distance or time standard by locating a fixed number of facilities_
#
# **MCLP in math notation:**
#
# $\begin{array} \displaystyle \textbf{Maximize} & \sum_{i=1}^{n}{a_iy_i} && (1) \\
# \displaystyle \textbf{Subject to:} & \sum_{j\in N_i}{x_j \geq y_i} & \forall i & (2) \\
# & \sum_{j}{x_j = p} & \forall j & (3) \\
# & y_i \in \{0,1\} & \forall i & (4) \\
# & x_j \in \{0,1\} & \forall j & (5) \\ \end{array}$
#
# $\begin{array} \displaystyle \textbf{Where:}\\ & & \displaystyle i & \small = & \textrm{index referencing nodes of the network as demand} \\
# & & j & \small = & \textrm{index referencing nodes of the network as potential facility sites} \\
# & & S & \small = & \textrm{maximal acceptable service distance or time standard} \\
# & & d_{ij} & \small = & \textrm{shortest distance or travel time between nodes} i \textrm{and} j \\
# & & N_i & \small = & \{j | d_{ij} < S\} \\
# & & p & \small = & \textrm{number of facilities to be located} \\
# & & x_j & \small = & \begin{cases}
# 1, \text{if a facility is located at node } j \\
# 0, \text{otherwise} \\
# \end{cases} \\
# & & y_i & \small = & \begin{cases}
# 1, \textrm{if demand } i \textrm{ is covered within a service standard} \\
# 0, \textrm{otherwise} \\
# \end{cases}\end{array}$
#
# _This excerpt above was quoted from Church L., <NAME>. (2018)_
#
#
# This tutorial solves MCLP using `spopt.locate.coverage.MCLP` instance that depends on a array 2D representing the costs between facilities candidate sites and demand points. For that it uses a lattice 10x10 with simulated points to calculate the costs.
# +
from spopt.locate.coverage import MCLP
from spopt.locate.util import simulated_geo_points
import numpy
import geopandas
import pulp
import spaghetti
from shapely.geometry import Point
import matplotlib.pyplot as plt
# -
# Since the model needs a distance cost matrix we should define some variables. In the comments, it's defined what these variables are for but solver. The solver, assigned below as `pulp.PULP_CBC_CMD`, is an interface to optimization solver developed by [COIN-OR](https://github.com/coin-or/Cbc). If you want to use another optimization interface as Gurobi or CPLEX see this [guide](https://coin-or.github.io/pulp/guides/how_to_configure_solvers.html) that explains how to achieve this.
# +
CLIENT_COUNT = 100 # quantity demand points
FACILITY_COUNT = 5 # quantity supply points
MAX_COVERAGE = 7 # maximum service radius
P_FACILITIES = 4
# Random seeds for reproducibility
CLIENT_SEED = 5
FACILITY_SEED = 6
solver = pulp.PULP_CBC_CMD(msg=False) # see solvers available in pulp reference
# -
# ## Lattice 10x10
# Create lattice 10x10 with 9 vertical lines in interior.
lattice = spaghetti.regular_lattice((0, 0, 10, 10), 9, exterior=True)
ntw = spaghetti.Network(in_data=lattice)
# Transform spaghetti instance to geopandas geodataframe.
# +
street = spaghetti.element_as_gdf(ntw, arcs=True)
street_buffered = geopandas.GeoDataFrame(
geopandas.GeoSeries(street["geometry"].buffer(0.2).unary_union),
crs=street.crs,
columns=["geometry"],
)
# -
# Plotting the network created by spaghetti we can verify that it seems a district with quarters and streets.
street.plot()
# ## Simulate points in a network
# The function `simulated_geo_points` simulates points inside a network. In this case, it uses a lattice network 10x10 created by using spaghetti package.
# Below we use the function defined above and simulate the points inside lattice bounds.
client_points = simulated_geo_points(street_buffered, needed=CLIENT_COUNT, seed=CLIENT_SEED)
facility_points = simulated_geo_points(
street_buffered, needed=FACILITY_COUNT, seed=FACILITY_SEED
)
# Plotting the 100 client and 5 facility points we can see that the function generates dummy points to an area of 10x10 which is the area created by our lattice created on previous cells.
fig, ax = plt.subplots(figsize=(6, 6))
street.plot(ax=ax, alpha=0.8, zorder=1, label='streets')
facility_points.plot(ax=ax, color='red', zorder=2, label='facility candidate sites ($n$=5)')
client_points.plot(ax=ax, color='black', label='clients points ($n$=100)')
plt.legend(loc='upper left', bbox_to_anchor=(1.05, 1))
# Here, for each client point the model suppose that there is a weight. So, we use randint function from numpy to also simulate these weights.
ai = numpy.random.randint(1, 12, CLIENT_COUNT)
# The weight is simulate with a 1-12 range, the minimum is 1 and the maximum is 12.
ai
# ## Transform simulated points to real points
# To use cost matrix or geodataframes we have to pay attention in some details. The client and facility points simulated don't belong to network, so if we calculate the distances now we are supposed to receive a wrong result. Before calculating distances we snap points to the networok and then calculate the distances.
# Below we snap points that is not spatially belong to network and create new real points geodataframes.
# +
ntw.snapobservations(client_points, "clients", attribute=True)
clients_snapped = spaghetti.element_as_gdf(
ntw, pp_name="clients", snapped=True
)
ntw.snapobservations(facility_points, "facilities", attribute=True)
facilities_snapped = spaghetti.element_as_gdf(
ntw, pp_name="facilities", snapped=True
)
# -
# Now the plot seems more organized as the points belong to network.
# The network created is plotted below with facility points and clients points:
fig, ax = plt.subplots(figsize=(6, 6))
street.plot(ax=ax, alpha=0.8, zorder=1, label='streets')
facilities_snapped.plot(ax=ax, color='red', zorder=2, label='facility candidate sites ($n$=5)')
clients_snapped.plot(ax=ax, color='black', label='clients points ($n$=100)')
plt.legend(loc='upper left', bbox_to_anchor=(1.05, 1))
# ## Calculating the cost matrix
# Calculate distance between clients and facilities.
cost_matrix = ntw.allneighbordistances(
sourcepattern=ntw.pointpatterns["clients"],
destpattern=ntw.pointpatterns["facilities"],
)
# The expected result here is a Dijkstra distance between clients and facilities points, so we our case an array 2D 100x5.
cost_matrix
# With ``MCLP.from_cost_matrix`` we model the MCL problem to cover all demand points with $p$ facility points within a `max_coverage` meters as service radius using cost matrix calculated previously.
mclp_from_cost_matrix = MCLP.from_cost_matrix(cost_matrix, ai, MAX_COVERAGE, p_facilities=P_FACILITIES)
result = mclp_from_cost_matrix.solve(solver)
# Expected result is an instance of MCLP.
mclp_from_cost_matrix
# ## Using GeoDataFrame
# Assigning service load array to demand geodataframe
clients_snapped['weights'] = ai
clients_snapped
# With ``MCLP.from_geodataframe`` we model the MCL problem to cover all demand points with $p$ facility points within a `max_coverage` meters as service radius using geodataframes without calculating the cost matrix previously.
mclp_from_geodataframe = MCLP.from_geodataframe(
clients_snapped,
facilities_snapped,
"geometry",
"geometry",
"weights",
MAX_COVERAGE,
p_facilities=P_FACILITIES,
distance_metric="euclidean"
)
mclp_from_geodataframe = mclp_from_geodataframe.solve(solver)
# Expected result is an instance of MCLP.
mclp_from_geodataframe
# ## Plotting the results
# The cell below describe the plotting of the results. For each method from MCLP class (from_cost_matrix, from_geodataframe) there is a plot displaying the facility site that was selected with a star colored and the points covered with the same color. Sometimes the demand points will be colored with not expected colors, it represents the coverage overlapping.
# +
from matplotlib.patches import Patch
import matplotlib.lines as mlines
dv_colors = [
"darkcyan",
"mediumseagreen",
"cyan",
"darkslategray",
"lightskyblue",
"limegreen",
"darkgoldenrod",
"peachpuff",
"coral",
"mediumvioletred",
"blueviolet",
"fuchsia",
"thistle",
"lavender",
"saddlebrown",
]
def plot_results(model, facility_points):
arr_points = []
fac_sites = []
for i in range(FACILITY_COUNT):
if model.fac2cli[i]:
geom = client_points.iloc[model.fac2cli[i]]['geometry']
arr_points.append(geom)
fac_sites.append(i)
fig, ax = plt.subplots(figsize=(6, 6))
legend_elements = []
street.plot(ax=ax, alpha=1, color='black', zorder=1)
legend_elements.append(mlines.Line2D(
[],
[],
color='black',
label='streets',
))
facility_points.plot(ax=ax, color='brown', marker="*", markersize=80, zorder=2)
legend_elements.append(mlines.Line2D(
[],
[],
color='brown',
marker="*",
linewidth=0,
label=f'facility sites ($n$={FACILITY_COUNT})'
))
for i in range(len(arr_points)):
gdf = geopandas.GeoDataFrame(arr_points[i])
label = f"coverage_points by y{fac_sites[i]}"
legend_elements.append(Patch(facecolor=dv_colors[i], edgecolor="k", label=label))
gdf.plot(ax=ax, zorder=3, alpha=0.7, edgecolor="k", color=dv_colors[i], label=label)
facility_points.iloc[[fac_sites[i]]].plot(ax=ax,
marker="*",
markersize=200 * 3.0,
alpha=0.8,
zorder=4,
edgecolor="k",
facecolor=dv_colors[i])
legend_elements.append(mlines.Line2D(
[],
[],
color=dv_colors[i],
marker="*",
ms=20 / 2,
markeredgecolor="k",
linewidth=0,
alpha=0.8,
label=f"y{fac_sites[i]} facility selected",
))
plt.title("MCLP", fontweight="bold")
plt.legend(handles = legend_elements, loc='upper left', bbox_to_anchor=(1.05, 1))
# -
# ### MCLP built from cost matrix
mclp_from_cost_matrix.facility_client_array()
plot_results(mclp_from_cost_matrix, facility_points)
# ### MCLP built from geodataframes
mclp_from_geodataframe.facility_client_array()
plot_results(mclp_from_geodataframe, facility_points)
# You may notice that the models are different. This result is expected as the distance between facility and demand points is calculated with different metrics. The cost matrix is calculated with dijkstra distance while the distance using geodataframe is calculated with euclidean distance.
# ## References
#
# - [<NAME>., & <NAME>. (2018). Location covering models: History, applications and advancements (1st edition 2018). Springer](https://www.springer.com/gb/book/9783319998459)
| notebooks/mclp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.6 64-bit
# name: python36664bitea6884f10f474b21a2a2f022451e0d09
# ---
# +
from tqdm import tqdm
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.layers import Dense, Dropout, LSTM, Embedding, Bidirectional
from tensorflow.keras.models import Sequential
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.utils import to_categorical
from sklearn.model_selection import train_test_split
from sklearn.datasets import fetch_20newsgroups
import numpy as np
from glob import glob
import random
import os
# +
def get_embedding_vectors(word_index, dim=100):
embedding_matrix = np.zeros((len(word_index) + 1, dim))
with open(f"data/glove.6B.{dim}d.txt", encoding="utf8") as f:
for line in tqdm(f, "Reading GloVe"):
values = line.split()
# get the word as the first word in the line
word = values[0]
if word in word_index:
idx = word_index[word]
# get the vectors as the remaining values in the line
embedding_matrix[idx] = np.array(values[1:], dtype="float32")
return embedding_matrix
def create_model(word_index, units=128, n_layers=2, cell=LSTM, bidirectional=False,
embedding_size=100, sequence_length=100, dropout=0.3,
loss="categorical_crossentropy", optimizer="adam",
output_length=2):
"""
Constructs a RNN model given its parameters
"""
embedding_matrix = get_embedding_vectors(word_index, embedding_size)
model = Sequential()
# add the embedding layer
model.add(Embedding(len(word_index) + 1,
embedding_size,
weights=[embedding_matrix],
trainable=False,
input_length=sequence_length))
for i in range(n_layers):
if i == n_layers - 1:
# last layer
if bidirectional:
model.add(Bidirectional(cell(units, return_sequences=False)))
else:
model.add(cell(units, return_sequences=False))
else:
# first layer or hidden layers
if bidirectional:
model.add(Bidirectional(cell(units, return_sequences=True)))
else:
model.add(cell(units, return_sequences=True))
model.add(Dropout(dropout))
model.add(Dense(output_length, activation="softmax"))
# compile the model
model.compile(optimizer=optimizer, loss=loss, metrics=["accuracy"])
return model
def save_imdb_data():
pos_training_files = glob("data/aclImdb/train/pos/*.txt")
neg_training_files = glob("data/aclImdb/train/neg/*.txt")
pos_testing_files = glob("data/aclImdb/test/pos/*.txt")
neg_testing_files = glob("data/aclImdb/test/neg/*.txt")
print("total pos training files:", len(pos_training_files))
print("total neg training files:", len(neg_training_files))
print("total pos testing files:", len(pos_testing_files))
print("total neg testing files:", len(neg_testing_files))
# load the data, 0 for negative sentiment, 1 for positive sentiment
data = []
for file in tqdm(pos_training_files, "Loading positive training data"):
data.append((open(file).read().strip(), 1))
for file in tqdm(neg_training_files, "Loading negative training data"):
data.append((open(file).read().strip(), 0))
for file in tqdm(pos_testing_files, "Loading positive testing data"):
data.append((open(file).read().strip(), 1))
for file in tqdm(neg_testing_files, "Loading negative testing data"):
data.append((open(file).read().strip(), 0))
# shuffle the data
random.shuffle(data)
with open("data/reviews.txt", "w") as reviews_file:
with open("data/labels.txt", "w") as labels_file:
for review, label in tqdm(data, "Writing data to files"):
print(review, file=reviews_file)
print(label, file=labels_file)
def load_imdb_data(num_words, sequence_length, test_size=0.25, oov_token=None):
# read reviews
reviews = []
with open("data/reviews.txt") as f:
for review in f:
review = review.strip()
reviews.append(review)
labels = []
with open("data/labels.txt") as f:
for label in f:
label = label.strip()
labels.append(label)
tokenizer = Tokenizer(num_words=num_words, oov_token=oov_token)
tokenizer.fit_on_texts(reviews)
X = tokenizer.texts_to_sequences(reviews)
X, y = np.array(X), np.array(labels)
# pad sequences with 0's
X = pad_sequences(X, maxlen=sequence_length)
# convert labels to one-hot encoded
y = to_categorical(y)
# split data to training and testing sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size, random_state=1)
data = {}
data["X_train"] = X_train
data["X_test"]= X_test
data["y_train"] = y_train
data["y_test"] = y_test
data["tokenizer"] = tokenizer
data["int2label"] = {0: "negative", 1: "positive"}
data["label2int"] = {"negative": 0, "positive": 1}
return data
def load_20_newsgroup_data(num_words, sequence_length, test_size=0.25, oov_token=None):
# load the 20 news groups dataset
# shuffling the data & removing each document's header, signature blocks and quotation blocks
dataset = fetch_20newsgroups(subset="all", shuffle=True, remove=("headers", "footers", "quotes"))
documents = dataset.data
labels = dataset.target
tokenizer = Tokenizer(num_words=num_words, oov_token=oov_token)
tokenizer.fit_on_texts(documents)
X = tokenizer.texts_to_sequences(documents)
X, y = np.array(X), np.array(labels)
# pad sequences with 0's
X = pad_sequences(X, maxlen=sequence_length)
# convert labels to one-hot encoded
y = to_categorical(y)
# split data to training and testing sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size, random_state=1)
data = {}
data["X_train"] = X_train
data["X_test"]= X_test
data["y_train"] = y_train
data["y_test"] = y_test
data["tokenizer"] = tokenizer
data["int2label"] = { i: label for i, label in enumerate(dataset.target_names) }
data["label2int"] = { label: i for i, label in enumerate(dataset.target_names) }
return data
# +
# max number of words in each sentence
SEQUENCE_LENGTH = 300
# N-Dimensional GloVe embedding vectors
# using 100 here, feel free to use 200 or 300
EMBEDDING_SIZE = 300
# number of words to use, discarding the rest
N_WORDS = 10000
# out of vocabulary token
OOV_TOKEN = None
# 30% testing set, 70% training set
TEST_SIZE = 0.3
# number of CELL layers
N_LAYERS = 1
# the RNN cell to use, LSTM in this case
RNN_CELL = LSTM
# whether it's a bidirectional RNN
IS_BIDIRECTIONAL = False
# number of units (RNN_CELL ,nodes) in each layer
UNITS = 128
# dropout rate
DROPOUT = 0.4
### Training parameters
LOSS = "categorical_crossentropy"
OPTIMIZER = "adam"
BATCH_SIZE = 64
EPOCHS = 6
def get_model_name(dataset_name):
# construct the unique model name
model_name = f"{dataset_name}-{RNN_CELL.__name__}-seq-{SEQUENCE_LENGTH}-em-{EMBEDDING_SIZE}-w-{N_WORDS}-layers-{N_LAYERS}-units-{UNITS}-opt-{OPTIMIZER}-BS-{BATCH_SIZE}-d-{DROPOUT}"
if IS_BIDIRECTIONAL:
# add 'bid' str if bidirectional
model_name = "bid-" + model_name
if OOV_TOKEN:
# add 'oov' str if OOV token is specified
model_name += "-oov"
return model_name
# +
from tensorflow.keras.callbacks import TensorBoard, ModelCheckpoint
import os
import pickle
# create these folders if they does not exist
if not os.path.isdir("results"):
os.mkdir("results")
if not os.path.isdir("logs"):
os.mkdir("logs")
if not os.path.isdir("data"):
os.mkdir("data")
# load the data
data = load_imdb_data(N_WORDS, SEQUENCE_LENGTH, TEST_SIZE, oov_token=OOV_TOKEN)
# data = load_20_newsgroup_data(N_WORDS, SEQUENCE_LENGTH, TEST_SIZE, oov_token=OOV_TOKEN)
# save the tokenizer object to use later in testing
# pickle.dump(data["tokenizer"], open(f"results/{model_name}_tokenizer.pickle", "wb"))
model = create_model(data["tokenizer"].word_index, units=UNITS, n_layers=N_LAYERS,
cell=RNN_CELL, bidirectional=IS_BIDIRECTIONAL, embedding_size=EMBEDDING_SIZE,
sequence_length=SEQUENCE_LENGTH, dropout=DROPOUT,
loss=LOSS, optimizer=OPTIMIZER, output_length=data["y_train"][0].shape[0])
# checkpointer = ModelCheckpoint(os.path.join("results", model_name),
# save_weights_only=True, save_best_only=True,
# verbose=1)
model.summary()
tensorboard = TensorBoard(log_dir=os.path.join("logs", model_name))
history = model.fit(data["X_train"], data["y_train"],
batch_size=BATCH_SIZE,
epochs=EPOCHS,
validation_data=(data["X_test"], data["y_test"]),
# callbacks=[checkpointer, tensorboard],
callbacks=[tensorboard],
verbose=1)
model.save(os.path.join("results", model_name) + ".h5")
# -
def get_predictions(text):
sequence = data["tokenizer"].texts_to_sequences([text])
# pad the sequences
sequence = pad_sequences(sequence, maxlen=SEQUENCE_LENGTH)
# get the prediction
prediction = model.predict(sequence)[0]
return prediction, data["int2label"][np.argmax(prediction)]
text = "Not very good, but pretty good try."
output_vector, prediction = get_predictions(text)
print("="*50)
print("Output vector:", output_vector)
print("Prediction:", prediction)
| machine-learning/nlp/text-classification/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Decision Tree Regressor
# ## Python version (sklearn)
#
# http://scikit-learn.org/stable/auto_examples/tree/plot_iris.html#example-tree-plot-iris-py
# +
# %matplotlib inline
import pylab
pylab.rcParams['figure.figsize'] = (16.0, 8.0)
import numpy as np
from sklearn.tree import DecisionTreeRegressor
import matplotlib.pyplot as plt
# Create a random dataset
rng = np.random.RandomState(1234)
X = np.sort(5 * rng.rand(80, 1), axis=0)
y = np.sin(X).ravel()
y[::5] += 3 * (0.5 - rng.rand(16))
# Fit regression model
regr_1 = DecisionTreeRegressor(max_depth=2)
regr_2 = DecisionTreeRegressor(max_depth=5)
regr_1.fit(X, y)
regr_2.fit(X, y)
# Predict
X_test = np.arange(0.0, 5.0, 0.01)[:, np.newaxis]
y_1 = regr_1.predict(X_test)
y_2 = regr_2.predict(X_test)
# Plot the results
plt.figure()
plt.scatter(X, y, c="k", label="data")
plt.plot(X_test, y_1, c="g", label="max_depth=2", linewidth=2)
plt.plot(X_test, y_2, c="r", label="max_depth=5", linewidth=2)
plt.xlabel("data")
plt.ylabel("target")
plt.title("Decision Tree Regression")
plt.legend()
plt.show()
# -
# ## Crystal version (crystal-learn)
# ```ruby
# require "random"
# require "../math"
# require "../random"
# require "../array"
# require "../trees"
# require "csv"
#
# x = Random.sequence(80).map {|x| x * 5}
# x.sort!
# y = Math.sin(x)
#
# seq = Random.sequence(16).map{|x| 3 * (0.5 - x)}
# y.each_with_index do |e, i|
# y[i] += seq[i/5] if i%5 == 0
# end
#
# regr2 = ML::Classifiers::DecisionTreeRegressor.new(max_depth: 2)
# regr5 = ML::Classifiers::DecisionTreeRegressor.new(max_depth: 5)
#
# x = x.map {|xi| [xi]}
#
# regr2.fit(x, y)
# regr5.fit(x, y)
#
# # Predict
# x_test = ML.arange(0.0, 5.0, step: 0.01).map {|x| [x]}
# y_pred2 = regr2.predict(x_test).map {|x| x.round(2)}
# y_pred5 = regr5.predict(x_test).map {|x| x.round(2)}
#
# puts "regresor max_depth: 2"
# regr2.show_tree(column_names: ["x", "y"])
#
# # puts "regresor max_depth: 5"
# # regr5.show_tree(column_names: ["x", "y"])
#
#
# f = File.open("regressor.csv", mode: "w")
#
# result = CSV.build(f) do |csv|
# x.zip(y).each do |x_i, y_i|
# csv.row x_i[0], y_i, "true"
# end
# x_test.zip(y_pred2).each do |x_i, y_i|
# csv.row x_i[0], y_i, "pred_2"
# end
# x_test.zip(y_pred5).each do |x_i, y_i|
# csv.row x_i[0], y_i, "pred_5"
# end
# end
#
# f.close()
#
# ```
#
# +
import pandas as pd
data = pd.read_csv('regressor.csv', names = ["x", "y", "kind"])
data_true = data[data.kind == "true"]
data_pred2 = data[data.kind == "pred_2"]
data_pred5 = data[data.kind == "pred_5"]
X = np.array(data_true.x)
y = np.array(data_true.y)
X_test = np.array(data_pred.x)
y_pred2 = np.array(data_pred2.y)
y_pred5 = np.array(data_pred5.y)
plt.figure()
plt.scatter(X, y, c="k", label="data")
plt.plot(X_test, y_pred2, c="g", label="max_depth=2", linewidth=2)
plt.plot(X_test, y_pred5, c="r", label="max_depth=5", linewidth=2)
plt.xlabel("data")
plt.ylabel("target")
plt.title("Decision Tree Regression (Crystal)")
plt.legend()
plt.show()
# -
from sklearn import tree
tree.export_graphviz(regr_1, out_file='tree.dot')
| examples/replicate-sklearn/sklearn-dtregressor-comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lamps08/Tensor-flow/blob/master/Tensor_flow.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pVrb1Uh7sxOx" colab_type="code" colab={}
import numpy as np
import os
import PIL
import PIL.Image
import tensorflow as tf
import pandas as pd
# + id="Z0OO1yEbyFf0" colab_type="code" colab={}
import pathlib
data_dir = pathlib.Path("/content/drive/My Drive/chest_xray/train/")
# + id="tynbiig-2g_2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="14027a89-dd63-4ca9-86e4-8a78bfbbd06f"
image_count = len(list(data_dir.glob('*/*.j')))
print(image_count)
# + id="uz7PDLJJ3KmK" colab_type="code" colab={}
from IPython.display import Image, display
display(Image ("/content/drive/My Drive/chest_xray/train/NORMAL/IM-0115-0001.jpeg"))
# + id="lkulujHX5D_N" colab_type="code" colab={}
Normal = list(data_dir.glob('train/NORMAL/*'))
#PIL.Image.open(str(Normal[0]))
# + id="-aplzYf0874-" colab_type="code" colab={}
pneumonia = list(data_dir.glob('train/pneumonia/*'))
# + id="6nZQnudC-CH6" colab_type="code" colab={}
batch_size = 32
img_height = 180
img_width = 180
# + id="VtKhY1RdIQDy" colab_type="code" colab={}
# #!pip install tf-nightly
# + id="T8VWq0It_U-H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="b0e2a6e3-0341-4dc1-fea0-f57952350f47"
# (run this line if you get image_dataset error)
train_ds = tf.keras.preprocessing.image_dataset_from_directory(
data_dir,
validation_split=0.2,
subset="training",
seed=123,
image_size=(img_height, img_width),
batch_size=batch_size)
# + id="X4b0tODYGeF_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="6ca79d1a-8134-4f54-dcb3-ff83c25ed831"
val_ds = tf.keras.preprocessing.image_dataset_from_directory(
data_dir,
validation_split=0.2,
subset="validation",
seed=123,
image_size=(img_height, img_width),
batch_size=batch_size)
# + id="QKPin2Fm_i2G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9aa25af0-d8c3-4d85-a710-21a1a85b8fb2"
class_names = train_ds.class_names
print(class_names)
# + id="_54IG1F2BEP4" colab_type="code" colab={}
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 10))
for images, labels in train_ds.take(1):
for i in range(4):
ax = plt.subplot(2, 2, i+1)
plt.imshow(images[i].numpy().astype("uint8"))
plt.title(class_names[labels[i]])
plt.axis("off")
# + id="1txB7YmLBI3Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="b8ffa1fd-979a-424a-b2e7-79459adec767"
for image_batch, labels_batch in train_ds:
print(image_batch.shape)
print(labels_batch.shape)
break
# + id="9xm48wJEEii8" colab_type="code" colab={}
from tensorflow.keras import layers
normalization_layer = tf.keras.layers.experimental.preprocessing.Rescaling(1./255)
# + id="b_ZgBBWNEq9o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="23f5bfa9-cd0b-44e8-c590-47918e8fa479"
normalized_ds = train_ds.map(lambda x, y: (normalization_layer(x), y))
image_batch, labels_batch = next(iter(normalized_ds))
first_image = image_batch[0]
# Notice the pixels values are now in `[0,1]`.
print(np.min(first_image), np.max(first_image))
# + id="gYP3GPIBFRfz" colab_type="code" colab={}
AUTOTUNE = tf.data.experimental.AUTOTUNE
train_ds = train_ds.cache().prefetch(buffer_size=AUTOTUNE)
val_ds = val_ds.cache().prefetch(buffer_size=AUTOTUNE)
# + id="My7hKI7UFXye" colab_type="code" colab={}
num_classes = 2
model = tf.keras.Sequential([
layers.experimental.preprocessing.Rescaling(1./255),
layers.Conv2D(32, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(32, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(32, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dense(num_classes)
])
# + id="PXRfxlDpFtiI" colab_type="code" colab={}
model.compile(
optimizer='adam',
loss=tf.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# + id="JNH-o52wFwrh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="c08db27b-3232-411e-a2b3-b6647ae45ec2"
model.fit(
train_ds,
batch_size=batch_size,
validation_data=val_ds,
epochs=3
)
# + id="3fqOUiGzFy1l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="10309643-fe93-40f3-931d-b37824a917de"
#model = create_model()
#model.fit(train_images, train_labels, epochs=5)
# Save the entire model as a SavedModel.
# !mkdir -p saved_model
model.save('saved_model/my_model')
# + id="WWawFfzzWe5P" colab_type="code" colab={}
| Tensor_flow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The CLTK has a distributed infrastructure that lets you download official CLTK texts or other corpora shared by others. For full docs, see <http://docs.cltk.org/en/latest/importing_corpora.html>.
#
# To get started, from the Terminal, open a new Jupyter notebook from within your `~/cltk` directory (see notebook 1 "CLTK Setup" for instructions): `jupyter notebook`. Then go to <http://localhost:8888>.
# # See what corpora are available
#
# First we need to "import" the right part of the CLTK library. Think of this as pulling just the book you need off the shelf and having it ready to read.
# +
# This is the import of the right part of the CLTK library
from cltk.corpus.utils.importer import CorpusImporter
# +
# See https://github.com/cltk for all official corpora
my_latin_downloader = CorpusImporter('latin')
# Now 'my_latin_downloader' is the variable by which we call the CorpusImporter
# -
my_latin_downloader.list_corpora
# # Import several corpora
my_latin_downloader.import_corpus('latin_text_latin_library')
my_latin_downloader.import_corpus('latin_models_cltk')
# You can verify the files were downloaded in the Terminal with `$ ls -l ~/cltk_data/latin/text/latin_text_latin_library/`
# +
# Let's get some Greek corpora, too
my_greek_downloader = CorpusImporter('greek')
my_greek_downloader.import_corpus('greek_models_cltk')
my_greek_downloader.list_corpora
# -
my_greek_downloader.import_corpus('greek_text_lacus_curtius')
# Likewise, verify with `ls -l ~/cltk_data/greek/text/greek_text_lacus_curtius/plain/`
my_greek_downloader.import_corpus('greek_text_first1kgreek')
# !ls -l ~/cltk_data/greek/text/greek_text_first1kgreek/
# # Convert TEI XML texts
#
# Here we'll convert the First 1K Years' Greek corpus from TEI XML to plain text.
from cltk.corpus.greek.tei import onekgreek_tei_xml_to_text
# +
# #! If you get the following error: 'Install `bs4` and `lxml` to parse these TEI files.'
# then run: `pip install bs4 lxml`.
onekgreek_tei_xml_to_text()
# +
# Count the converted plaintext files
# !ls -l ~/cltk_data/greek/text/greek_text_first1kgreek_plaintext/ | wc -l
# -
# # Import local corpora
my_latin_downloader.import_corpus('phi5', '~/cltk/corpora/PHI5/')
my_latin_downloader.import_corpus('phi7', '~/cltk/corpora/PHI7/')
my_greek_downloader.import_corpus('tlg', '~/cltk/corpora/TLG_E/')
# !ls -l /home/kyle/cltk_data/originals/
| 2 Import corpora.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Install a pip package in the current Jupyter kernel
import sys
# !{sys.executable} -m pip install scipy
# !{sys.executable} -m pip install matplotlib
# +
from scipy.signal import butter, lfilter
import matplotlib.pyplot as plt
def butter_bandpass(lowcut, highcut, fs, order=5):
nyq = 0.5 * fs
low = lowcut / nyq
high = highcut / nyq
b, a = butter(order, [low, high], btype='band')
return b, a
def butter_bandpass_filter(data, lowcut, highcut, fs, order=5):
b, a = butter_bandpass(lowcut, highcut, fs, order=order)
y = lfilter(b, a, data)
return y
# -
def run():
import numpy as np
import matplotlib.pyplot as plt
from scipy.signal import freqz
# Sample rate and desired cutoff frequencies (in Hz).
fs = 5000.0
lowcut = 500.0
highcut = 1250.0
# Plot the frequency response for a few different orders.
plt.figure(1)
plt.clf()
for order in [3, 6, 9]:
b, a = butter_bandpass(lowcut, highcut, fs, order=order)
w, h = freqz(b, a, worN=2000)
plt.plot((fs * 0.5 / np.pi) * w, abs(h), label="order = %d" % order)
plt.plot([0, 0.5 * fs], [np.sqrt(0.5), np.sqrt(0.5)],
'--', label='sqrt(0.5)')
plt.xlabel('Frequency (Hz)')
plt.ylabel('Gain')
plt.grid(True)
plt.legend(loc='best')
# Filter a noisy signal.
T = 0.05
nsamples = int(T * fs)
t = np.linspace(0, T, nsamples, endpoint=False)
a = 0.02
f0 = 600.0
# The input of scipy filter should be a Numpy array
x = 0.1 * np.sin(2 * np.pi * 1.2 * np.sqrt(t))
print(type(x))
x += 0.01 * np.cos(2 * np.pi * 312 * t + 0.1)
x += a * np.cos(2 * np.pi * f0 * t + .11)
x += 0.03 * np.cos(2 * np.pi * 2000 * t)
plt.figure(2)
plt.clf()
plt.plot(t, x, label='Noisy signal')
y = butter_bandpass_filter(x, lowcut, highcut, fs, order=6)
plt.plot(t, y, label='Filtered signal (%g Hz)' % f0)
plt.xlabel('time (seconds)')
plt.hlines([-a, a], 0, T, linestyles='--')
plt.grid(True)
plt.axis('tight')
plt.legend(loc='upper left')
plt.show()
run()
| signal_processing/.ipynb_checkpoints/bandpass_filter-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# `use bit shift operator >> , << to define tasks order of execuation `
# `if the order execuation not definded workflow will execuate the tasks without order sequentails`
# `all upstream >> tasks must run be for downstream tasks <<`
# Task Defination
# </br>
# Define simple arguments
# ####
# Define the DAG
# ####
# Define Tasks and connected to dag
# ####
# Define task Order
#
from airflow.operators.bash import BashOperator
from datetime import datetime;
from airflow.models import DAG;
default_args={
"owner":"BK",
"start_date":datetime(2021,2,10)
}
dag=DAG("DAG_Example",default_args=default_args)
task_id_1 = BashOperator(task_id="task_1",bash_command="echo 'TASK 1 Execuation'",dag=dag)
task_id_2=BashOperator(task_id="task_id_2",bash_command="echo 'Task 2 Execuation'",dag=dag)
task_id_1 >> task_id_2
| Introduction to Airflow in Python/Implementing Airflow DAGS/practice/Workflow-DAG example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Introduction-(top-down-method)" data-toc-modified-id="Introduction-(top-down-method)-1"><span class="toc-item-num">1 </span>Introduction (top down method)</a></span></li><li><span><a href="#Script-setup" data-toc-modified-id="Script-setup-2"><span class="toc-item-num">2 </span>Script setup</a></span></li><li><span><a href="#Data-directory-preparation" data-toc-modified-id="Data-directory-preparation-3"><span class="toc-item-num">3 </span>Data directory preparation</a></span></li><li><span><a href="#Data-file-preperation" data-toc-modified-id="Data-file-preperation-4"><span class="toc-item-num">4 </span>Data file preperation</a></span></li><li><span><a href="#Load-data-functions" data-toc-modified-id="Load-data-functions-5"><span class="toc-item-num">5 </span>Load data functions</a></span></li><li><span><a href="#Load-data-sets" data-toc-modified-id="Load-data-sets-6"><span class="toc-item-num">6 </span>Load data sets</a></span></li><li><span><a href="#Methodology---MAP,-AP-and-pure-heat-(EEA-method)" data-toc-modified-id="Methodology---MAP,-AP-and-pure-heat-(EEA-method)-7"><span class="toc-item-num">7 </span>Methodology - MAP, AP and pure heat (EEA method)</a></span><ul class="toc-item"><li><span><a href="#Energy-related-calculations" data-toc-modified-id="Energy-related-calculations-7.1"><span class="toc-item-num">7.1 </span>Energy related calculations</a></span></li><li><span><a href="#Emissions-related-calculations" data-toc-modified-id="Emissions-related-calculations-7.2"><span class="toc-item-num">7.2 </span>Emissions related calculations</a></span></li><li><span><a href="#Carbon-intensity-calculation" data-toc-modified-id="Carbon-intensity-calculation-7.3"><span class="toc-item-num">7.3 </span>Carbon intensity calculation</a></span></li></ul></li><li><span><a href="#Methodology---MAP,-AP-and-only-heat-from-CHP-($\sigma=1$)" data-toc-modified-id="Methodology---MAP,-AP-and-only-heat-from-CHP-($\sigma=1$)-8"><span class="toc-item-num">8 </span>Methodology - MAP, AP and only heat from CHP ($\sigma=1$)</a></span><ul class="toc-item"><li><span><a href="#Energy-related-calculations" data-toc-modified-id="Energy-related-calculations-8.1"><span class="toc-item-num">8.1 </span>Energy related calculations</a></span></li><li><span><a href="#Emissions-related-calculations" data-toc-modified-id="Emissions-related-calculations-8.2"><span class="toc-item-num">8.2 </span>Emissions related calculations</a></span></li><li><span><a href="#Carbon-intensity-calculation" data-toc-modified-id="Carbon-intensity-calculation-8.3"><span class="toc-item-num">8.3 </span>Carbon intensity calculation</a></span></li></ul></li><li><span><a href="#Methodology---MAP,-AP-and-without-heat-from-CHP-($\sigma=0$)" data-toc-modified-id="Methodology---MAP,-AP-and-without-heat-from-CHP-($\sigma=0$)-9"><span class="toc-item-num">9 </span>Methodology - MAP, AP and without heat from CHP ($\sigma=0$)</a></span><ul class="toc-item"><li><span><a href="#Energy-related-calculations" data-toc-modified-id="Energy-related-calculations-9.1"><span class="toc-item-num">9.1 </span>Energy related calculations</a></span></li><li><span><a href="#Emissions-related-calculations" data-toc-modified-id="Emissions-related-calculations-9.2"><span class="toc-item-num">9.2 </span>Emissions related calculations</a></span></li><li><span><a href="#Carbon-intensity-calculation" data-toc-modified-id="Carbon-intensity-calculation-9.3"><span class="toc-item-num">9.3 </span>Carbon intensity calculation</a></span></li></ul></li><li><span><a href="#Methodology---MAP-and-only-heat-from-CHP--($\sigma=1$)" data-toc-modified-id="Methodology---MAP-and-only-heat-from-CHP--($\sigma=1$)-10"><span class="toc-item-num">10 </span>Methodology - MAP and only heat from CHP ($\sigma=1$)</a></span><ul class="toc-item"><li><span><a href="#Energy-related-calculations" data-toc-modified-id="Energy-related-calculations-10.1"><span class="toc-item-num">10.1 </span>Energy related calculations</a></span></li><li><span><a href="#Emissions-related-calculations" data-toc-modified-id="Emissions-related-calculations-10.2"><span class="toc-item-num">10.2 </span>Emissions related calculations</a></span></li><li><span><a href="#Carbon-intensity-calculation" data-toc-modified-id="Carbon-intensity-calculation-10.3"><span class="toc-item-num">10.3 </span>Carbon intensity calculation</a></span></li></ul></li><li><span><a href="#Results" data-toc-modified-id="Results-11"><span class="toc-item-num">11 </span>Results</a></span><ul class="toc-item"><li><span><a href="#Some-country-results-as-an-overview" data-toc-modified-id="Some-country-results-as-an-overview-11.1"><span class="toc-item-num">11.1 </span>Some country results as an overview</a></span></li><li><span><a href="#Plots" data-toc-modified-id="Plots-11.2"><span class="toc-item-num">11.2 </span>Plots</a></span><ul class="toc-item"><li><span><a href="#Comparing-sigma-1-and-sigma-0-methods-for-year-2018" data-toc-modified-id="Comparing-sigma-1-and-sigma-0-methods-for-year-2018-11.2.1"><span class="toc-item-num">11.2.1 </span>Comparing sigma 1 and sigma 0 methods for year 2018</a></span></li><li><span><a href="#CI_MAP_AP_without_heat-for-the-10-largest-energy-consumer-countries" data-toc-modified-id="CI_MAP_AP_without_heat-for-the-10-largest-energy-consumer-countries-11.2.2"><span class="toc-item-num">11.2.2 </span>CI_MAP_AP_without_heat for the 10 largest energy consumer countries</a></span></li><li><span><a href="#CI_1-for-the-three-base-years-for-all-countries" data-toc-modified-id="CI_1-for-the-three-base-years-for-all-countries-11.2.3"><span class="toc-item-num">11.2.3 </span>CI_1 for the three base years for all countries</a></span></li></ul></li></ul></li><li><span><a href="#Export-final-CI-to-csv" data-toc-modified-id="Export-final-CI-to-csv-12"><span class="toc-item-num">12 </span>Export final CI to csv</a></span></li></ul></div>
# -
# # Introduction (top down method)
# In this script, we determine country specific carbon intensity factors (CI) for European countries. The applied methods are based on a published procedures from the European Environment Agency (EEA).
#
# EEA method documentation: https://www.eea.europa.eu/data-and-maps/data/co2-intensity-of-electricity-generation/
# # Script setup
# +
import os
import logging
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.dates import DateFormatter
from IPython.display import Image
# %matplotlib inline
plt.style.use('seaborn')
plt.rcParams['figure.figsize'] = [15, 10]
#helpers
from helpers import get_country
from helpers import aligndata
# -
# # Data directory preparation
# Create input, processed and output folders if they don't exist
# If the paths are relative, the corresponding folders will be created inside the current working directory.
# - input -> all needed input data
# - processed -> save point and exchange with other scripts
# - output -> final emission factors
# +
input_directory_path = os.path.join('input')
top_down_method_input_directory_path = os.path.join('input', 'top_down_method')
processed_directory_path = 'processed'
output_directory_path = os.path.join('output')
os.makedirs(input_directory_path, exist_ok=True)
os.makedirs(top_down_method_input_directory_path, exist_ok=True)
os.makedirs(processed_directory_path, exist_ok=True)
os.makedirs(output_directory_path, exist_ok=True)
# -
# # Data file preperation
# The directory `input/top_down_method` should contain all necessary raw data files.
#
# - 1) Eurostat energy balance database https://ec.europa.eu/eurostat/web/energy/data/database
# - Complete energy balances as ZIP archive -> nrg_bal_c.tsv (tab separated file) https://ec.europa.eu/eurostat/estat-navtree-portlet-prod/BulkDownloadListing?file=data/nrg_bal_c.tsv.gz
#
# - 2) National emissions reported to the UNFCCC and to the EU Greenhouse Gas Monitoring Mechanism https://www.eea.europa.eu/data-and-maps/data/national-emissions-reported-to-the-unfccc-and-to-the-eu-greenhouse-gas-monitoring-mechanism-16
# - Reported emissions as ZIP archive -> UNFCCC_v23.csv (ASCII delimited) https://www.eea.europa.eu/data-and-maps/data/national-emissions-reported-to-the-unfccc-and-to-the-eu-greenhouse-gas-monitoring-mechanism-16/national-greenhouse-gas-inventories-ipcc-common-reporting-format-sector-classification/ascii-delimited-zip-2/at_download/file
#
#
# +
# Checks if the the input directories are empty or not
# Checks all filenames in the input directory
if not os.listdir(top_down_method_input_directory_path) :
print("The directory for the method is empty. Please provide the data to the directory as described in the instructions above.")
filenames = [os.path.join(top_down_method_input_directory_path, fn) for fn in os.listdir(top_down_method_input_directory_path)]
print(filenames)
# -
# # Load data functions
# +
def load_energy_balance_data(path, fn):
"""
Load the raw energy balances reported in the Eurostat database for all European countries from input directory.
Parameters
----------
path: str
path to data
fn : str
filename
"""
logging.info(f'Loading data from {fn}')
df = pd.read_csv(os.path.join(path, fn),sep = '\t', header=0)
# rename column (0) for identifier
df = df.rename(columns = {df.columns[0]:'use_substance_unit_country'})
return df
def load_UNFCC_data(path, fn):
"""
Load and standardize the raw UNFCC database for all European countries from input directory.
Filter data: only sector '1.A.1.a' (CO2 for all energy production from Public Electricity Generation, Public Combined Heat and Power and Public Heat Plants)
Filter data: only direct CO2 emissions
Filter data: only from year 1990 upwards
CO2 Emissions in million tones Tg
Parameters
----------
path: str
path to data
fn : str
filename
"""
logging.info(f'Loading data from {fn}')
df = pd.read_csv(os.path.join(path, fn), sep = ',', header =0, encoding = 'unicode_escape',low_memory=False)
#sector and pollutant selection
sector = '1.A.1.a' # CO2 for all energy production from Public Electricity Generation, Public Combined Heat and Power and Public Heat Plants
pollutant = 'CO2' # only direct CO2
df = df.query('Sector_code == @sector').query('Pollutant_name == @pollutant')
# data only from year 1990 upwards
df = df[~df['Year'].isin(['1985-1987'])] # skip an entry with several years
# convert all years to string
df['Year']= df['Year'].apply(lambda x: int(x)).apply(lambda x: str(x))
df = df[~df['Year'].isin(['1985','1986','1987','1988','1989'])] # filter for years
# Create table with countries as columns and years as rows
df = pd.pivot_table(df, values = 'emissions', index = 'Year', columns = ['Country_code'])
return df
# -
# # Load data sets
# The following image give information about the used data sets, abbreviations and calculation method
Image(filename= os.path.join(top_down_method_input_directory_path, 'top_down_method.png'))
# #### load CO2_emissions reported from UNFCC
# +
CO2_emissions_UNFCC = load_UNFCC_data(top_down_method_input_directory_path, 'UNFCCC_v23.csv')
# attention public electricity and heat generation only no autoproducers
# rename UK to GB
CO2_emissions_UNFCC.rename(columns={'UK':'GB'}, inplace=True)
# missing countries MD, ME, MK, RS,
# -
# #### load energy balance sheet reported from eurostat
nrg_bal_c = load_energy_balance_data(top_down_method_input_directory_path, 'nrg_bal_c.csv')
# +
# replace EU country code by ISO country code
nrg_bal_c.use_substance_unit_country.replace(to_replace='EL', value='GR' , regex=True, inplace=True)
nrg_bal_c.use_substance_unit_country.replace(to_replace='UK', value='GB' , regex=True, inplace=True)
#missing countries: CH, LI
# -
# # Methodology - MAP, AP and pure heat (EEA method)
# ## Energy related calculations
# Calculation for ei_MAP, ei_AP, dh_MAP, dh_AP, GEP
# see image for further information
# +
#using only substances provided in the example excel-sheet (Austria Example). Nevertheless, the methodology on the website
#suggests using two more substances namely primary biogases and primary solid biofuels
# Solid fossil fuel; C0000X0350-0370;
# Oil and petroleum products (excl. Biofuel); O4000XBIO;
# Natural Gas; G3000;
# Manufactured gases; C0350-0370;
# Peat and peat products; P1000;
# Oil shale and Oil sands; S2000;
#--- not in the example but in the documentation Primary solid biofuels; R5110-5150_W6000RI;
#--- not in the example but in the documentation Primary Biogases; R5300;
# Non-renewable waste; W6100_6220.
# set filter substance
substances = 'C0000X0350-0370|O4000XBIO|G3000|C0350-0370|P1000|S2000|W6100_6220'
# +
#energy input of main activity producers (ei_MAP)
#Transformation input Electricity & heat generation
#Main activity producer electricity only; TI_EHG_MAPE_E
#Main activity producer CHP; TI_EHG_MAPCHP_E
#Main activity producer heat only; TI_EHG_MAPH_E
ei_MAP_string_nrgbalc = 'TI_EHG_MAPE_E|TI_EHG_MAPCHP_E|TI_EHG_MAPH_E'
#filter with ei_MAP_string_nrgbalc and substances
ei_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(ei_MAP_string_nrgbalc, regex=True)]
ei_MAP = ei_MAP.loc[ei_MAP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
ei_MAP = ei_MAP.loc[ei_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
# split country from identifier
ei_MAP.use_substance_unit_country = ei_MAP.use_substance_unit_country.apply(lambda string: get_country(string))
# rename columns and set index
ei_MAP = ei_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_MAP.index.name,level = 0).sum()\
.T
# +
#calculating the energy input of Autoproducters (ei_AP)
#Transformation input Electricity & heat generation
#Autoproducer electricity only; TI_EHG_APE_E
#Autoproducer producer CHP; TI_EHG_APCHP_E
#Autoproducer producer heat only; TI_EHG_APH_E
AP_string = 'TI_EHG_APE_E|TI_EHG_APCHP_E|TI_EHG_APH_E'
ei_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(AP_string, regex=True)]
ei_AP = ei_AP.loc[ei_AP['use_substance_unit_country']\
.str.contains(substances,regex=True)]#
ei_AP = ei_AP.loc[ei_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
ei_AP.use_substance_unit_country = ei_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
ei_AP = ei_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_AP.index.name,level = 0).sum()\
.T
# +
#calculating the derived heat of main activity producers(dh_MAP)
#Gross heat production
#Main activity producer CHP; GHP_MAPCHP
#Main activity producer heat only; GHP_MAPH
#filter
derived_heat_string_nrgbalc = 'GHP_MAPCHP|GHP_MAPH'
dh_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(derived_heat_string_nrgbalc, regex=True)]
dh_MAP = dh_MAP.loc[dh_MAP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
dh_MAP = dh_MAP.loc[dh_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
dh_MAP.use_substance_unit_country = dh_MAP.use_substance_unit_country\
.apply(lambda string: get_country(string))
dh_MAP = dh_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=dh_MAP.index.name,level = 0).sum()\
.T\
#/ 0.9 # Estimating 90% efficiency heat production
# +
#calculating the derived heat of Autoproducters (dh_AP)
#Gross heat production
#Autoproducer CHP; GHP_APCHP
#Autoproducer heat only; GHP_APH
dh_AP_string = 'GHP_APCHP|GHP_APH'
dh_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(dh_AP_string, regex=True)]
dh_AP = dh_AP.loc[dh_AP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
dh_AP = dh_AP.loc[dh_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
dh_AP.use_substance_unit_country = dh_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
dh_AP = dh_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=dh_AP.index.name,level = 0).sum()\
.T\
#/ 0.9 # Estimating 90% efficiency heat production
# +
#Calculating gross electricity production of main activity producers (GEP)
#Gross electricity production
#Main activity producer electricity only; GEP_MAPE
#Main activity producer CHP; GEP_MAPCHP
MAP_string = 'GEP_MAPE|GEP_MAPCHP'
GEP_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(MAP_string, regex=True)]
GEP_MAP = GEP_MAP.loc[GEP_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)(?=.*TOTAL)')]
GEP_MAP.use_substance_unit_country = GEP_MAP.use_substance_unit_country\
.apply(lambda string: get_country(string))
GEP_MAP = GEP_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=GEP_MAP.index.name,level = 0).sum()\
.T
# +
#Calculating gross electricity production of Autoproductersof (GEP_AP)
#Gross electricity production
#Autoproducer electricity only; GEP_APE
#Autoproducer CHP; GEP_APCHP
AP_string = 'GEP_APE|GEP_APCHP'
GEP_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(AP_string, regex=True)]
GEP_AP = GEP_AP.loc[GEP_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)(?=.*TOTAL)')]
GEP_AP.use_substance_unit_country = GEP_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
GEP_AP = GEP_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=GEP_AP.index.name,level = 0).sum()\
.T
# -
# ## Emissions related calculations
#filter CO2_emissions_UNFCC using only columns that exist in both dataframes
CO2_cleaned = CO2_emissions_UNFCC[pd.Series(CO2_emissions_UNFCC.columns)[pd.Series(CO2_emissions_UNFCC.columns).apply(lambda x: x in ei_MAP.columns)]]
# +
# fit data to CO2_cleaned data columns and rows
#Transformation input Main activity producer electricity only, CHP and heat
ei_MAP = aligndata(ei_MAP,CO2_cleaned)
#Gross heat production Main activity producer CHP and heat
dh_MAP= aligndata(dh_MAP, CO2_cleaned)
#Transformation input Autoproducer electricity only, CHP and heat
ei_AP = aligndata(ei_AP, CO2_cleaned)
#Gross heat production Autoproducer CHP and heat
dh_AP = aligndata(dh_AP, CO2_cleaned)
#electricity production Main activity producer
GEP_MAP= aligndata(GEP_MAP, CO2_cleaned)
#electricity production Autoproducer
GEP_AP = aligndata(GEP_AP, CO2_cleaned)
#Gross electricity production
GEP_EEA = GEP_MAP + GEP_AP
# -
# ## Carbon intensity calculation
# +
# CO2 intensity of total electricity generation
# Is calculated by the ratio of all CO2 emissions from all electricity production (public main activity producers and autoproducers)
# against total electricity generation including all sources.
# Main activity producer
# First, the CO2 emissions of gross electricity production:
# total CO2 emissions multiplying by
# electricity production (plus energy losses) from public conventional thermal power stations
# versus all electrical energy production from public power stations and combined heat power station
# CO2 emissions * ((electrical energy + energy losses)/(electrical energy + derived heat + energy losses))
# 1) assumption
# (electrical energy + energy losses) = (electrical energy + derived heat + energy losses)–(derived heat)
# (electrical energy + derived heat + energy losses) = ei_MAP
# 2) assumption
# energy input calculation for derived heat with eff. of 0.9
# (derived heat) = (dh_MAP / 0.9)
# resulting in:
# energy input electrical energy production
# (electrical energy + derived heat + energy losses)–(derived heat) = ei_MAP - (dh_MAP / 0.9)
ei_elec_MAP = ei_MAP - (dh_MAP / 0.9)
# ((electrical energy + energy losses)/(electrical energy + derived heat + energy losses)) = ((ei_MAP - (dh_MAP / 0.9)) / ei_MAP)
# share of CO2 emissions of electricity production Main activity producer
CO2_from_MAP_elec = CO2_cleaned * (ei_elec_MAP / ei_MAP)
# Autoproducer
# The reported CO2 emissions in class 1A1a do not include CO2 emissions from autoproducers.
# Emissions from autoproducers were therefore estimated
# electricity output of autoproducers (same assumption as above)
ei_elec_AP = ei_AP - (dh_AP / 0.9)
# 1) assumption
# this was done by multiplying the electricity output of autoproducers
# by a calculated CO2 emission ratio for main activity producers
# share of CO2 emissions of electricity production Autoproducer
CO2_from_AP_elec = CO2_from_MAP_elec * (ei_elec_AP / ei_elec_MAP)
# CO2 intensity of total electricity generation
# sum of CO2 from MAP and CO2 from AP [CO2 in Gigagramm (Gg)] / Gross electricity production [GWh](85.98: Ktoe-->GWh)
CI_EEA = ((CO2_from_MAP_elec + CO2_from_AP_elec)/(GEP_EEA/85.98)) # CO2 intensity in [g CO2/kWh]
# -
# # Methodology - MAP, AP and only heat from CHP ($\sigma=1$)
# ## Energy related calculations
# Calculation for ei_MAP, ei_MAP_heat, ei_AP, ei_AP_heat, dh_MAP, dh_AP, GEP
# +
#using only substances provided in the example excel-sheet (Austria Example). Nevertheless, the methodology on the website
#suggests using two more substances namely primary biogases and primary solid biofuels
# Solid fossil fuel; C0000X0350-0370;
# Oil and petroleum products (excl. Biofuel); O4000XBIO;
# Natural Gas; G3000;
# Manufactured gases; C0350-0370;
# Peat and peat products; P1000;
# Oil shale and Oil sands; S2000;
#--- not in the example but in the documentation Primary solid biofuels; R5110-5150_W6000RI;
#--- not in the example but in the documentation Primary Biogases; R5300;
# Non-renewable waste; W6100_6220.
# set filter substance
substances = 'C0000X0350-0370|O4000XBIO|G3000|C0350-0370|P1000|S2000|W6100_6220'
# +
#energy input of main activity producers (ei_MAP)
#Transformation input Electricity
#Main activity producer electricity only; TI_EHG_MAPE_E
#Main activity producer CHP; TI_EHG_MAPCHP_E
ei_MAP_string_nrgbalc = 'TI_EHG_MAPE_E|TI_EHG_MAPCHP_E'
#filter with ei_MAP_string_nrgbalc and substances
ei_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(ei_MAP_string_nrgbalc, regex=True)]
ei_MAP = ei_MAP.loc[ei_MAP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
ei_MAP = ei_MAP.loc[ei_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
# split country from identifier
ei_MAP.use_substance_unit_country = ei_MAP.use_substance_unit_country.apply(lambda string: get_country(string))
# rename columns and set index
ei_MAP = ei_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_MAP.index.name,level = 0).sum()\
.T
# +
#energy input of main activity producers heat (ei_MAP_heat)
#Transformation input heat generation
#Main activity producer heat only; TI_EHG_MAPH_E
ei_MAP_heat_string_nrgbalc = 'TI_EHG_MAPH_E'
#filter with ei_MAP_string_nrgbalc and substances
ei_MAP_heat = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(ei_MAP_heat_string_nrgbalc, regex=True)]
ei_MAP_heat = ei_MAP_heat.loc[ei_MAP_heat['use_substance_unit_country']\
.str.contains(substances, regex=True)]
ei_MAP_heat = ei_MAP_heat.loc[ei_MAP_heat['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
# split country from identifier
ei_MAP_heat.use_substance_unit_country = ei_MAP_heat.use_substance_unit_country.apply(lambda string: get_country(string))
# rename columns and set index
ei_MAP_heat = ei_MAP_heat.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_MAP.index.name,level = 0).sum()\
.T
# +
#calculating the energy input of Autoproducters (ei_AP)
#Transformation input Electricity
#Autoproducer electricity only; TI_EHG_APE_E
#Autoproducer producer CHP; TI_EHG_APCHP_E
AP_string = 'TI_EHG_APE_E|TI_EHG_APCHP_E'
ei_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(AP_string, regex=True)]
ei_AP = ei_AP.loc[ei_AP['use_substance_unit_country']\
.str.contains(substances,regex=True)]#
ei_AP = ei_AP.loc[ei_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
ei_AP.use_substance_unit_country = ei_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
ei_AP = ei_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_AP.index.name,level = 0).sum()\
.T
# +
#calculating the energy input of Autoproducters (ei_AP_heat)
#Transformation input heat generation
#Autoproducer producer heat only; TI_EHG_APH_E
AP_heat_string = 'TI_EHG_APH_E'
ei_AP_heat = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(AP_heat_string, regex=True)]
ei_AP_heat = ei_AP_heat.loc[ei_AP_heat['use_substance_unit_country']\
.str.contains(substances,regex=True)]#
ei_AP_heat = ei_AP_heat.loc[ei_AP_heat['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
ei_AP_heat.use_substance_unit_country = ei_AP_heat.use_substance_unit_country\
.apply(lambda string: get_country(string))
ei_AP_heat = ei_AP_heat.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_AP.index.name,level = 0).sum()\
.T
# +
#calculating the derived heat of main activity producers(dh_MAP) CHP only
#Gross heat production
#Main activity producer CHP; GHP_MAPCHP
#filter
derived_heat_string_nrgbalc = 'GHP_MAPCHP'
dh_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(derived_heat_string_nrgbalc, regex=True)]
dh_MAP = dh_MAP.loc[dh_MAP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
dh_MAP = dh_MAP.loc[dh_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
dh_MAP.use_substance_unit_country = dh_MAP.use_substance_unit_country\
.apply(lambda string: get_country(string))
dh_MAP = dh_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=dh_MAP.index.name,level = 0).sum()\
.T\
#/ 0.9 # Estimating 90% efficiency heat production
# +
#calculating the derived heat of Autoproducters (dh_AP) CHP only
#Gross heat production
#Autoproducer CHP; GHP_APCHP
dh_AP_string = 'GHP_APCHP'
dh_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(dh_AP_string, regex=True)]
dh_AP = dh_AP.loc[dh_AP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
dh_AP = dh_AP.loc[dh_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
dh_AP.use_substance_unit_country = dh_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
dh_AP = dh_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=dh_AP.index.name,level = 0).sum()\
.T\
#/ 0.9 # Estimating 90% efficiency heat production
# +
#Calculating gross electricity production of main activity producers (GEP)
#Gross electricity production
#Main activity producer electricity only; GEP_MAPE
#Main activity producer CHP; GEP_MAPCHP
MAP_string = 'GEP_MAPE|GEP_MAPCHP'
GEP_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(MAP_string, regex=True)]
GEP_MAP = GEP_MAP.loc[GEP_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)(?=.*TOTAL)')]
GEP_MAP.use_substance_unit_country = GEP_MAP.use_substance_unit_country\
.apply(lambda string: get_country(string))
GEP_MAP = GEP_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=GEP_MAP.index.name,level = 0).sum()\
.T
# +
#Calculating gross electricity production of Autoproductersof (GEP_AP)
#Gross electricity production
#Autoproducer electricity only; GEP_APE
#Autoproducer CHP; GEP_APCHP
AP_string = 'GEP_APE|GEP_APCHP'
GEP_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(AP_string, regex=True)]
GEP_AP = GEP_AP.loc[GEP_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)(?=.*TOTAL)')]
GEP_AP.use_substance_unit_country = GEP_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
GEP_AP = GEP_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=GEP_AP.index.name,level = 0).sum()\
.T
# -
# ## Emissions related calculations
#filter CO2_emissions_UNFCC using only columns that exist in both dataframes
CO2_cleaned = CO2_emissions_UNFCC[pd.Series(CO2_emissions_UNFCC.columns)[pd.Series(CO2_emissions_UNFCC.columns).apply(lambda x: x in ei_MAP.columns)]]
# +
# fit data to CO2_cleaned data columns and rows
#Transformation input Main activity producer electricity only, CHP
ei_MAP = aligndata(ei_MAP,CO2_cleaned)
#Transformation input Main activity producer heat
ei_MAP_heat = aligndata(ei_MAP_heat, CO2_cleaned)
#Gross heat production Main activity producer CHP
dh_MAP= aligndata(dh_MAP, CO2_cleaned)
#Transformation input Autoproducer electricity only, CHP
ei_AP = aligndata(ei_AP, CO2_cleaned)
#Transformation input Autoproducer heat
ei_AP_heat = aligndata(ei_AP_heat, CO2_cleaned)
#Gross heat production Autoproducer CHP
dh_AP = aligndata(dh_AP, CO2_cleaned)
#electricity production Main activity producer
GEP_MAP= aligndata(GEP_MAP, CO2_cleaned)
#electricity production Autoproducer
GEP_AP = aligndata(GEP_AP, CO2_cleaned)
#Gross electricity production
GEP = (GEP_MAP + GEP_AP)
#converting gross elec to net elec assuming 6% self consumption
NEP_1 = GEP * 0.94
# set sigma for heat allocation
sigma = 1
# -
# ## Carbon intensity calculation
# +
# CO2 intensity of total electricity generation
# Is calculated by the ratio of all CO2 emissions from all electricity production (public main activity producers and autoproducers)
# against total electricity generation including all sources.
# Main activity producer
# energy input electrical energy production
ei_elec_MAP = ei_MAP - (sigma * (dh_MAP / 0.9))
# share of CO2 emissions of electricity production Main activity producer
CO2_from_MAP_elec = CO2_cleaned * (ei_elec_MAP / (ei_MAP + ei_MAP_heat))
# Autoproducer
# The reported CO2 emissions in class 1A1a do not include CO2 emissions from autoproducers.
# Emissions from autoproducers were therefore estimated
# electricity output of autoproducers (same assumption as above)
ei_elec_AP = ei_AP - (sigma * (dh_AP / 0.9))
# 1) assumption
# this was done by multiplying the electricity output of autoproducers
# by a calculated CO2 emission ratio for main activity producers
# share of CO2 emissions of electricity production Autoproducer
CO2_from_AP_elec = CO2_from_MAP_elec * (ei_elec_AP / ei_elec_MAP)
# CO2 intensity of total electricity generation
# sum of CO2 from MAP and CO2 from AP [CO2 in Gigagramm (Gg)] / Gross electricity production [GWh](85.98: Ktoe-->GWh)
CI_1 = ((CO2_from_MAP_elec + CO2_from_AP_elec)/(NEP_1/85.98)) # CO2 intensity in [g CO2/kWh]
# -
# # Methodology - MAP, AP and without heat from CHP ($\sigma=0$)
# ## Energy related calculations
# Calculation for ei_MAP, ei_MAP_heat, ei_AP, ei_AP_heat, dh_MAP, dh_AP, GEP
# +
#using only substances provided in the example excel-sheet (Austria Example). Nevertheless, the methodology on the website
#suggests using two more substances namely primary biogases and primary solid biofuels
# Solid fossil fuel; C0000X0350-0370;
# Oil and petroleum products (excl. Biofuel); O4000XBIO;
# Natural Gas; G3000;
# Manufactured gases; C0350-0370;
# Peat and peat products; P1000;
# Oil shale and Oil sands; S2000;
#--- not in the example but in the documentation Primary solid biofuels; R5110-5150_W6000RI;
#--- not in the example but in the documentation Primary Biogases; R5300;
# Non-renewable waste; W6100_6220.
# set filter substance
substances = 'C0000X0350-0370|O4000XBIO|G3000|C0350-0370|P1000|S2000|W6100_6220'
# +
#energy input of main activity producers (ei_MAP)
#Transformation input Electricity
#Main activity producer electricity only; TI_EHG_MAPE_E
#Main activity producer CHP; TI_EHG_MAPCHP_E
ei_MAP_string_nrgbalc = 'TI_EHG_MAPE_E|TI_EHG_MAPCHP_E'
#filter with ei_MAP_string_nrgbalc and substances
ei_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(ei_MAP_string_nrgbalc, regex=True)]
ei_MAP = ei_MAP.loc[ei_MAP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
ei_MAP = ei_MAP.loc[ei_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
# split country from identifier
ei_MAP.use_substance_unit_country = ei_MAP.use_substance_unit_country.apply(lambda string: get_country(string))
# rename columns and set index
ei_MAP = ei_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_MAP.index.name,level = 0).sum()\
.T
# +
#energy input of main activity producers heat (ei_MAP_heat)
#Transformation input heat generation
#Main activity producer heat only; TI_EHG_MAPH_E
ei_MAP_heat_string_nrgbalc = 'TI_EHG_MAPH_E'
#filter with ei_MAP_string_nrgbalc and substances
ei_MAP_heat = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(ei_MAP_heat_string_nrgbalc, regex=True)]
ei_MAP_heat = ei_MAP_heat.loc[ei_MAP_heat['use_substance_unit_country']\
.str.contains(substances, regex=True)]
ei_MAP_heat = ei_MAP_heat.loc[ei_MAP_heat['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
# split country from identifier
ei_MAP_heat.use_substance_unit_country = ei_MAP_heat.use_substance_unit_country.apply(lambda string: get_country(string))
# rename columns and set index
ei_MAP_heat = ei_MAP_heat.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_MAP.index.name,level = 0).sum()\
.T
# +
#calculating the energy input of Autoproducters (ei_AP)
#Transformation input Electricity
#Autoproducer electricity only; TI_EHG_APE_E
#Autoproducer producer CHP; TI_EHG_APCHP_E
AP_string = 'TI_EHG_APE_E|TI_EHG_APCHP_E'
ei_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(AP_string, regex=True)]
ei_AP = ei_AP.loc[ei_AP['use_substance_unit_country']\
.str.contains(substances,regex=True)]#
ei_AP = ei_AP.loc[ei_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
ei_AP.use_substance_unit_country = ei_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
ei_AP = ei_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_AP.index.name,level = 0).sum()\
.T
# +
#calculating the energy input of Autoproducters (ei_AP_heat)
#Transformation input heat generation
#Autoproducer producer heat only; TI_EHG_APH_E
AP_heat_string = 'TI_EHG_APH_E'
ei_AP_heat = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(AP_heat_string, regex=True)]
ei_AP_heat = ei_AP_heat.loc[ei_AP_heat['use_substance_unit_country']\
.str.contains(substances,regex=True)]#
ei_AP_heat = ei_AP_heat.loc[ei_AP_heat['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
ei_AP_heat.use_substance_unit_country = ei_AP_heat.use_substance_unit_country\
.apply(lambda string: get_country(string))
ei_AP_heat = ei_AP_heat.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_AP.index.name,level = 0).sum()\
.T
# +
#calculating the derived heat of main activity producers(dh_MAP) CHP only
#Gross heat production
#Main activity producer CHP; GHP_MAPCHP
#filter
derived_heat_string_nrgbalc = 'GHP_MAPCHP'
dh_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(derived_heat_string_nrgbalc, regex=True)]
dh_MAP = dh_MAP.loc[dh_MAP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
dh_MAP = dh_MAP.loc[dh_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
dh_MAP.use_substance_unit_country = dh_MAP.use_substance_unit_country\
.apply(lambda string: get_country(string))
dh_MAP = dh_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=dh_MAP.index.name,level = 0).sum()\
.T\
#/ 0.9 # Estimating 90% efficiency heat production
# +
#calculating the derived heat of Autoproducters (dh_AP) CHP only
#Gross heat production
#Autoproducer CHP; GHP_APCHP
dh_AP_string = 'GHP_APCHP'
dh_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(dh_AP_string, regex=True)]
dh_AP = dh_AP.loc[dh_AP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
dh_AP = dh_AP.loc[dh_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
dh_AP.use_substance_unit_country = dh_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
dh_AP = dh_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=dh_AP.index.name,level = 0).sum()\
.T\
#/ 0.9 # Estimating 90% efficiency heat production
# +
#Calculating gross electricity production of main activity producers (GEP)
#Gross electricity production
#Main activity producer electricity only; GEP_MAPE
#Main activity producer CHP; GEP_MAPCHP
MAP_string = 'GEP_MAPE|GEP_MAPCHP'
GEP_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(MAP_string, regex=True)]
GEP_MAP = GEP_MAP.loc[GEP_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)(?=.*TOTAL)')]
GEP_MAP.use_substance_unit_country = GEP_MAP.use_substance_unit_country\
.apply(lambda string: get_country(string))
GEP_MAP = GEP_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=GEP_MAP.index.name,level = 0).sum()\
.T
# +
#Calculating gross electricity production of Autoproductersof (GEP_AP)
#Gross electricity production
#Autoproducer electricity only; GEP_APE
#Autoproducer CHP; GEP_APCHP
AP_string = 'GEP_APE|GEP_APCHP'
GEP_AP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(AP_string, regex=True)]
GEP_AP = GEP_AP.loc[GEP_AP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)(?=.*TOTAL)')]
GEP_AP.use_substance_unit_country = GEP_AP.use_substance_unit_country\
.apply(lambda string: get_country(string))
GEP_AP = GEP_AP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=GEP_AP.index.name,level = 0).sum()\
.T
# -
# ## Emissions related calculations
#filter CO2_emissions_UNFCC using only columns that exist in both dataframes
CO2_cleaned = CO2_emissions_UNFCC[pd.Series(CO2_emissions_UNFCC.columns)[pd.Series(CO2_emissions_UNFCC.columns).apply(lambda x: x in ei_MAP.columns)]]
# +
# fit data to CO2_cleaned data columns and rows
#Transformation input Main activity producer electricity only, CHP
ei_MAP = aligndata(ei_MAP,CO2_cleaned)
#Transformation input Main activity producer heat
ei_MAP_heat = aligndata(ei_MAP_heat, CO2_cleaned)
#Gross heat production Main activity producer CHP
dh_MAP= aligndata(dh_MAP, CO2_cleaned)
#Transformation input Autoproducer electricity only, CHP
ei_AP = aligndata(ei_AP, CO2_cleaned)
#Transformation input Autoproducer heat
ei_AP_heat = aligndata(ei_AP_heat, CO2_cleaned)
#Gross heat production Autoproducer CHP
dh_AP = aligndata(dh_AP, CO2_cleaned)
#electricity production Main activity producer
GEP_MAP= aligndata(GEP_MAP, CO2_cleaned)
#electricity production Autoproducer
GEP_AP = aligndata(GEP_AP, CO2_cleaned)
#Gross electricity production
GEP = GEP_MAP + GEP_AP
#converting gross elec to net elec assuming 6% self consumption
NEP_0 = GEP * 0.94
# set sigma for heat allocation
sigma = 0
# -
# ## Carbon intensity calculation
# +
# CO2 intensity of total electricity generation
# Is calculated by the ratio of all CO2 emissions from all electricity production (public main activity producers and autoproducers)
# against total electricity generation including all sources.
# Main activity producer
# energy input electrical energy production
ei_elec_MAP = ei_MAP - (sigma * (dh_MAP / 0.9))
# share of CO2 emissions of electricity production Main activity producer
CO2_from_MAP_elec = CO2_cleaned * (ei_elec_MAP / (ei_MAP + ei_MAP_heat))
# Autoproducer
# The reported CO2 emissions in class 1A1a do not include CO2 emissions from autoproducers.
# Emissions from autoproducers were therefore estimated
# electricity output of autoproducers (same assumption as above)
ei_elec_AP = ei_AP - (sigma * (dh_AP / 0.9))
# 1) assumption
# this was done by multiplying the electricity output of autoproducers
# by a calculated CO2 emission ratio for main activity producers
# share of CO2 emissions of electricity production Autoproducer
CO2_from_AP_elec = CO2_from_MAP_elec * (ei_elec_AP / ei_elec_MAP)
# CO2 intensity of total electricity generation
# sum of CO2 from MAP and CO2 from AP [CO2 in Gigagramm (Gg)] / Gross electricity production [GWh](85.98: Ktoe-->GWh)
CI_0 = ((CO2_from_MAP_elec + CO2_from_AP_elec)/(NEP_0/85.98)) # CO2 intensity in [g CO2/kWh]
# -
# # Methodology - MAP and only heat from CHP ($\sigma=1$)
# CI for the MAP only, the AP are not considered
# ## Energy related calculations
# Calculation for ei_MAP, dh_MAP, GEP
# +
#using only substances provided in the example excel-sheet (Austria Example). Nevertheless, the methodology on the website
#suggests using two more substances namely primary biogases and primary solid biofuels
# Solid fossil fuel; C0000X0350-0370;
# Oil and petroleum products (excl. Biofuel); O4000XBIO;
# Natural Gas; G3000;
# Manufactured gases; C0350-0370;
# Peat and peat products; P1000;
# Oil shale and Oil sands; S2000;
#--- not in the example but in the documentation Primary solid biofuels; R5110-5150_W6000RI;
#--- not in the example but in the documentation Primary Biogases; R5300;
# Non-renewable waste; W6100_6220.
# set filter substance
substances = 'C0000X0350-0370|O4000XBIO|G3000|C0350-0370|P1000|S2000|W6100_6220'
# +
#energy input of main activity producers (ei_MAP)
#Transformation input Electricity
#Main activity producer electricity only; TI_EHG_MAPE_E
#Main activity producer CHP; TI_EHG_MAPCHP_E
ei_MAP_string_nrgbalc = 'TI_EHG_MAPE_E|TI_EHG_MAPCHP_E'
#filter with ei_MAP_string_nrgbalc and substances
ei_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(ei_MAP_string_nrgbalc, regex=True)]
ei_MAP = ei_MAP.loc[ei_MAP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
ei_MAP = ei_MAP.loc[ei_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
# split country from identifier
ei_MAP.use_substance_unit_country = ei_MAP.use_substance_unit_country.apply(lambda string: get_country(string))
# rename columns and set index
ei_MAP = ei_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_MAP.index.name,level = 0).sum()\
.T
# +
#energy input of main activity producers heat (ei_MAP_heat)
#Transformation input heat generation
#Main activity producer heat only; TI_EHG_MAPH_E
ei_MAP_heat_string_nrgbalc = 'TI_EHG_MAPH_E'
#filter with ei_MAP_string_nrgbalc and substances
ei_MAP_heat = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(ei_MAP_heat_string_nrgbalc, regex=True)]
ei_MAP_heat = ei_MAP_heat.loc[ei_MAP_heat['use_substance_unit_country']\
.str.contains(substances, regex=True)]
ei_MAP_heat = ei_MAP_heat.loc[ei_MAP_heat['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
# split country from identifier
ei_MAP_heat.use_substance_unit_country = ei_MAP_heat.use_substance_unit_country.apply(lambda string: get_country(string))
# rename columns and set index
ei_MAP_heat = ei_MAP_heat.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=ei_MAP.index.name,level = 0).sum()\
.T
# +
#calculating the derived heat of main activity producers(dh_MAP)
#Gross heat production
#Main activity producer CHP; GHP_MAPCHP
#filter
derived_heat_string_nrgbalc = 'GHP_MAPCHP'
dh_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country']\
.str.contains(derived_heat_string_nrgbalc, regex=True)]
dh_MAP = dh_MAP.loc[dh_MAP['use_substance_unit_country']\
.str.contains(substances, regex=True)]
dh_MAP = dh_MAP.loc[dh_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)')]
dh_MAP.use_substance_unit_country = dh_MAP.use_substance_unit_country\
.apply(lambda string: get_country(string))
dh_MAP = dh_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=dh_MAP.index.name,level = 0).sum()\
.T\
#/ 0.9 # Estimating 90% efficiency heat production
# +
#Calculating gross electricity production of main activity producers (GEP)
#Gross electricity production
#Main activity producer electricity only; GEP_MAPE
#Main activity producer CHP; GEP_MAPCHP
MAP_string = 'GEP_MAPE|GEP_MAPCHP'
GEP_MAP = nrg_bal_c.loc[nrg_bal_c['use_substance_unit_country'].str.contains(MAP_string, regex=True)]
GEP_MAP = GEP_MAP.loc[GEP_MAP['use_substance_unit_country']\
.str.contains(r'^(?=.*KTOE)(?=.*TOTAL)')]
GEP_MAP.use_substance_unit_country = GEP_MAP.use_substance_unit_country\
.apply(lambda string: get_country(string))
GEP_MAP = GEP_MAP.rename(columns = {'use_substance_unit_country':'Country_code'})\
.set_index('Country_code')\
.replace(': ',0.)\
.replace(': z',0.)\
.apply(lambda x: x.apply(lambda y:float(y)))\
.groupby(by=GEP_MAP.index.name,level = 0).sum()\
.T
# -
# ## Emissions related calculations
#filter CO2_emissions_UNFCC using only columns that exist in both dataframes
CO2_cleaned = CO2_emissions_UNFCC[pd.Series(CO2_emissions_UNFCC.columns)[pd.Series(CO2_emissions_UNFCC.columns).apply(lambda x: x in ei_MAP.columns)]]
# +
# fit data to CO2_cleaned data columns and rows
#Transformation input Main activity producer electricity only, CHP
ei_MAP = aligndata(ei_MAP,CO2_cleaned)
#Transformation input Main activity producer heat
ei_MAP_heat = aligndata(ei_MAP_heat, CO2_cleaned)
#Gross heat production Main activity producer CHP
dh_MAP= aligndata(dh_MAP, CO2_cleaned)
#electricity production Main activity producer
GEP_MAP= aligndata(GEP_MAP, CO2_cleaned)
#Gross electricity production
GEP = GEP_MAP
#converting gross elec to net elec assuming 6% self consumption
NEP_MAP_1 = GEP * 0.94
# set sigma for heat allocation
sigma = 1
# -
# ## Carbon intensity calculation
# +
# CO2 intensity of total electricity generation
# Is calculated by the ratio of all CO2 emissions from all electricity production (public main activity producers and autoproducers)
# against total electricity generation including all sources.
# Main activity producer
# energy input electrical energy production
# (electrical energy + derived heat + energy losses)–(derived heat) = ei_MAP - (dh_MAP / 0.9)
ei_elec_MAP = ei_MAP - (sigma * (dh_MAP / 0.9))
# share of CO2 emissions of electricity production Main activity producer
CO2_from_MAP_elec = CO2_cleaned * (ei_elec_MAP / (ei_MAP + ei_MAP_heat))
# CO2 intensity of total electricity generation
# sum of CO2 from MAP and CO2 from AP [CO2 in Gigagramm (Gg)] / Gross electricity production [GWh](85.98: Ktoe-->GWh)
CI_MAP_1 = ((CO2_from_MAP_elec)/(NEP_MAP_1/85.98)) # CO2 intensity in [g CO2/kWh]
# -
# # Results
# ## Some country results as an overview
# +
# DE
country = 'DE'
print('CI_EEA = ' + str(CI_EEA[country].loc['2018']))
print('CI_1 = ' + str(CI_1[country].loc['2018']))
print('CI_0 = ' + str(CI_0[country].loc['2018']))
print('CI_MAP_1 = ' + str(CI_MAP_1[country].loc['2018']))
print('GEP_EEA = ' + str(GEP_EEA[country].loc['2018']))
print('NEP_1 = ' + str(NEP_1[country].loc['2018']))
print('NEP_0 = ' + str(NEP_0[country].loc['2018']))
print('NEP_MAP_1 = ' + str(NEP_MAP_1[country].loc['2018']))
# +
# DK
country = 'SE'
print('CI_EEA = ' + str(CI_EEA[country].loc['2018']))
print('CI_1 = ' + str(CI_1[country].loc['2018']))
print('CI_0 = ' + str(CI_0[country].loc['2018']))
print('CI_MAP_1 = ' + str(CI_MAP_1[country].loc['2018']))
print('GEP_EEA = ' + str(GEP_EEA[country].loc['2018']))
print('NEP_1 = ' + str(NEP_1[country].loc['2018']))
print('NEP_0 = ' + str(NEP_0[country].loc['2018']))
print('NEP_MAP_1 = ' + str(NEP_MAP_1[country].loc['2018']))
# -
# ## Plots
# ### Comparing sigma 1 and sigma 0 methods for year 2018
# +
CI_compar = CI_1.loc[['2018']].transpose()
CI_compar.rename(columns={'2018':'CI_1'},inplace=True)
CI_compar['CI_0'] = CI_0.loc[['2018']].transpose()
#can be used to add data from other methods
#CI_compar['CI_MAP_CHP_heat'] = CI_MAP_CHP_heat.loc[['2018']].transpose()
#CI_compar['CI_MAP_AP_without_heat'] = CI_MAP_AP_without_heat.loc[['2018']].transpose()
# +
# Reorder the values
ordered_df = CI_compar.sort_values(by='CI_1', ascending=True)
my_range=range(1,len(CI_compar.index)+1)
fig, ax = plt.subplots(1,1)
# The vertical plot is made using the vline function
ax.vlines(x=my_range, ymin=ordered_df['CI_1'], ymax=ordered_df['CI_0'], color='grey', alpha=0.4, linewidths=2.5)
ax.scatter(my_range, ordered_df['CI_1'], color='green', alpha=0.6 , label='CI ($\sigma=1$)',s=90,edgecolor='black')
ax.scatter(my_range, ordered_df['CI_0'], color='red', alpha=0.6 , label='CI ($\sigma=0$)',s=90,edgecolor='black')
#ax.scatter(my_range , ordered_df['CI_MAP_CHP_heat'], color='yellow', alpha=0.6, label='CI_MAP_CHP_heat',s=90,edgecolor='black')
#ax.scatter(my_range , ordered_df['CI_MAP_AP_without_heat'], color='skyblue', alpha=0.6, label='CI_MAP_AP_without_heat',s=90,edgecolor='black')
handles, labels = ax.get_legend_handles_labels()
ax.legend(handles, labels, loc='upper left',
handletextpad=0., columnspacing=0.5, ncol=1,
fontsize=14, frameon=True)
# Add title and axis names
ax.set_xlabel('Country', fontsize=22)
ax.set_ylabel('Carbon intensity (g CO2 / kWh)', fontsize=22)
ax.set_xticks(my_range)
ax.set_xticklabels(ordered_df.index)
ax.tick_params(axis='x',labelsize=19, rotation=45)
ax.tick_params(axis='y',labelsize=19)
# using pass to not print the return from set_xticks and ...
pass
# -
fig.savefig(os.path.join(output_directory_path + '/_CO2_intensity_comparing_methods.png'))
# ### CI_MAP_AP_without_heat for the 10 largest energy consumer countries
GEP.loc['2018'].sort_values(ascending=False)[:10].index
# +
# CI_1 developent from 1990 upto 2018
sns.set(font_scale = 1.2)
fig, ax = plt.subplots()
ax = sns.lineplot(data=CI_1[GEP.loc['2018'].sort_values(ascending=False)[:10].index],
markers=False,
dashes=True,
linewidth=2,
markersize=9,
palette=sns.color_palette("muted"),
legend='full')
handles, labels = ax.get_legend_handles_labels()
ax.legend(handles, labels, loc='upper right',
handletextpad=0., columnspacing=0.5, ncol=1,
title='Country', fontsize=14, frameon=True)
ax.set_ylabel("CO2 Intensity (g CO2/kWh)", fontsize=15)
ax.set_xlabel("Year", fontsize=15)
ax.tick_params(axis='x', rotation=45)
# -
fig.savefig(os.path.join(output_directory_path + '/_CO2_intensity_by_large_consumer_countries_ver1.png'))
# +
# CI_1 developent from 1990 upto 2018
sns.set(font_scale = 1.2)
fig, ax = plt.subplots()
ax = sns.lineplot(data=CI_1[GEP.loc['2018'].sort_values(ascending=False)[:10].index],
markers=True,
dashes=False,
linewidth=0.7,
markersize=9,
palette=sns.color_palette("muted"),
legend='full')
handles, labels = ax.get_legend_handles_labels()
ax.legend(handles, labels, loc='upper right',
handletextpad=0., columnspacing=0.5, ncol=1,
title='Country', fontsize=14, frameon=True)
ax.set_ylabel("CO2 Intensity (g CO2/kWh)", fontsize=15)
ax.set_xlabel("Year", fontsize=15)
ax.tick_params(axis='x', rotation=45)
# -
fig.savefig(os.path.join(output_directory_path + '/_CO2_intensity_by_large_consumer_countries_ver2.png'))
# ### CI_1 for the three base years for all countries
# +
CI_compar = CI_1.loc[['2018','2010','2000','1990']].transpose()
# Reorder it following the values of the first value:
ordered_df = CI_compar.sort_values(by='2018', ascending=True)
my_range=range(1,len(CI_compar.index)+1)
fig, ax = plt.subplots(1,1)
# The vertical plot is made using the vline function
ax.vlines(x=my_range, ymin=ordered_df['2018'], ymax=ordered_df['2000'], color='grey', alpha=0.4, linewidths=2.5)
ax.vlines(x=my_range, ymin=ordered_df['2000'], ymax=ordered_df['1990'], color='grey', alpha=0.4, linewidths=2.5)
ax.scatter(my_range, ordered_df['1990'], color='green', alpha=0.6 , label='1990',s=90,edgecolor='black')
ax.scatter(my_range, ordered_df['2000'], color='red', alpha=0.6 , label='2000',s=90,edgecolor='black')
ax.scatter(my_range , ordered_df['2018'], color='skyblue', alpha=0.6, label='2018',s=90,edgecolor='black')
#plt.scatter(my_range, ordered_df['2010'], color='green', alpha=0.4 , label='2010',s=90,edgecolor='black')
handles, labels = ax.get_legend_handles_labels()
ax.legend(handles, labels, loc='upper left',
handletextpad=0., columnspacing=0.5, ncol=1,
title='Years', fontsize=14, frameon=True)
# Add title and axis names
ax.set_xlabel('Country', fontsize=22)
ax.set_ylabel('Carbon intensity (g CO2 / kWh)', fontsize=22)
ax.set_xticks(my_range)
ax.set_xticklabels(ordered_df.index)
ax.tick_params(axis='x',labelsize=19, rotation=45)
ax.tick_params(axis='y',labelsize=19)
# using pass to not print the return from set_xticks and ...
pass
# -
fig.savefig(os.path.join(output_directory_path + '/_CO2_intensity_develoment_by_countries.png'))
# # Export final CI to csv
CI_1.to_csv(processed_directory_path + '/CI_1_top_down.csv')
CI_1.to_csv(output_directory_path + '/CI_1_top_down.csv')
CI_0.to_csv(processed_directory_path + '/CI_0_top_down.csv')
CI_0.to_csv(output_directory_path + '/CI_0_top_down.csv')
CI_MAP_1.to_csv(processed_directory_path + '/CI_MAP_1_top_down.csv')
CI_MAP_1.to_csv(output_directory_path + '/CI_MAP_1_top_down.csv')
# +
ei_MAP.to_csv(output_directory_path + '/ei_MAP_top_down.csv')
ei_AP.to_csv(output_directory_path + '/ei_AP_top_down.csv')
ei_MAP.to_csv(processed_directory_path + '/ei_MAP_top_down.csv')
ei_AP.to_csv(processed_directory_path + '/ei_AP_top_down.csv')
# -
| CI calculation top down method.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# language: python
# name: python38564bit02a66c47ce504b05b2ef5646cfed96c2
# ---
# # Spark Low Level API
# ## Outline
#
# - Foundations
# - Functional programming (Spark RDDs)
# - Use of pandas (Spark DataFramej
# - Use of SQL (Spark SQL)
# - Catalyst optimizer
# - Analysis
# - DataFrame
# - SQL AST
# - Logical plan
# - Catalog resolution of names
# - Rule-based optimization
# - Boolean simplification
# - Predicate pushdown
# - Constant folding
# - Projection pruning
# - Physical plan
# - Convert to RDD
# - Cost-based optimization
# - Code generation
# - Project Tungsten
# - Performs role of compiler
# - Generates Java bytecode
#
# 
#
# Resilient distributed datasets (RDD)
# - In-memory distributed collections
# - Removes need to write to disk for fault tolerance
# - Resilience
# - Reconstruct from lineage
# - Immutable
# - Distributed
# - RDD data live in one or more partitions
# - Dataset
# - Consists of records
# - Each partition consists of distinct set of records that can be operated on independently
# - Shared nothing philosophy
#
# 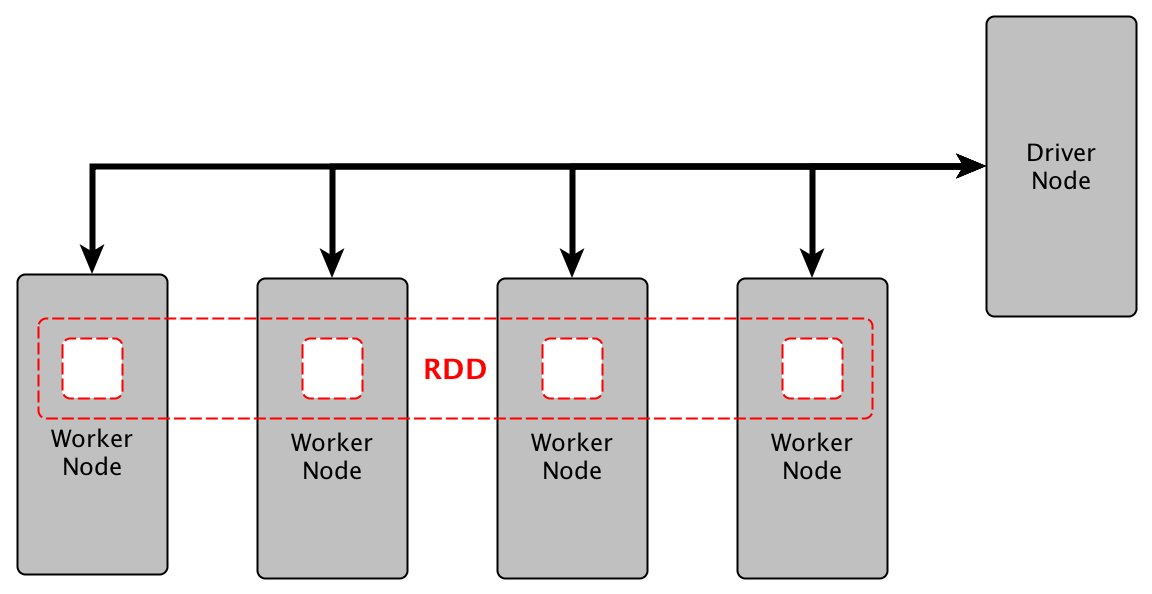
#
#
# - Loading data into RDDs
# - Programmatically
# - range()
# - parallelize()
# - From file
# - Compression
# - Splittable and non-splittable formats
# - Non-splittable files cannot be distributed
# - Splittable formats - LZO, Snappy
# - Non-splittable formats - gzip, zip
# - Data locality
# - Worker partitions from nearby DFS partitions
# - Default partition size is 128 MB
# - Local file system
# - Networked filesystem
# - Distributed filesystem
# - textFile()
# - WoleTextFiles()
# - From data resource
# - From stream
# - Persistence
# - persist)
# - cache()
# - Types of RDDs
# - Base RDD
# - Pair RDD
# - Double RDD
# - Many others
#
# Base RDD
#
# - Narrow transformations
# - map()
# - filter()
# - flatMap(j
# - distinct()
# - Broad transformations
# - reduce()
# - groupby()
# - sortBy()
# - join()
# - Actions
# - count()
# - take()
# - takeOrdered()
# - top()
# - collect()
# - saveAsTextFile()
# - first()
# - reduce()
# - fold()
# - aggregate()
# - foreach()
#
# PairedRDD
#
# - Dictionary functions
# - keys()
# - values()
# - keyBy()
# - Functional transformations
# - mapValues()
# - flatMapValues()
# - Grouping, sorting and aggregation
# - groupByKey()
# - reduceByKey()
# - foldByKey()
# - sortByKey()
# - Joins
# - Join large by small
# - join()
# - leftOuterJoin()
# - rightOuterJoin()
# - fullOuterJoin()
# - cogroup()
# - cartesian()
# - Set operations
# - union()
# - intersection(j
# - subtract()
# - subtractByKey()
#
# Numeric RDD
#
# - min()
# - max()
# - sum()
# - mean()
# - stdev()
# - variance()
# ## Architecture of a Spark Application
#
# ### Big picture
#
# You will type your commands iin a local Spark session, and the SparkContext will take care of running your instructions distributed across the workers (executors) on a cluster. Each executor can have 1 or more CPU cores, its own memory cahe, and is responsible for handling its own distributed tasks. Communicaiton between local and workers and between worker and worker is handled by a cluster manager.
#
# 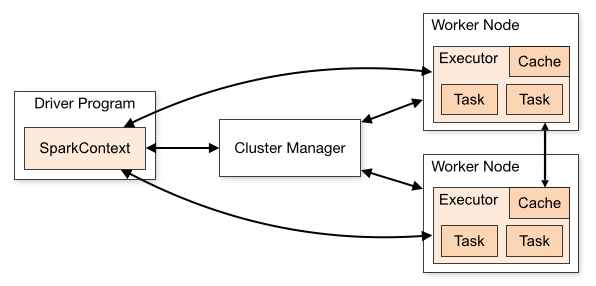
#
# Source: http://spark.apache.org/docs/latest/img/cluster-overview.png
#
# ### Organizaiton of Spark tasks
#
# Spark organizes tasks that can be performed without exchanging data across partitions into stages. The sequecne of tasks to be perfomed are laid out as a Directed Acyclic Graph (DAG). Tasks are differenitated into transforms (lazy evalutation - just add to DAG) and actions (eager evaluation - execute the specified path in the DAG). Note that calculations are not cached unless requested. Hence if you have triggered the action after RDD3 in the figure, then trigger the aciton after RDD6, RDD2 will be re-generated from RDD1 twice. We can avoid the re-calculation by persisting or cacheing RDD2.
#
# 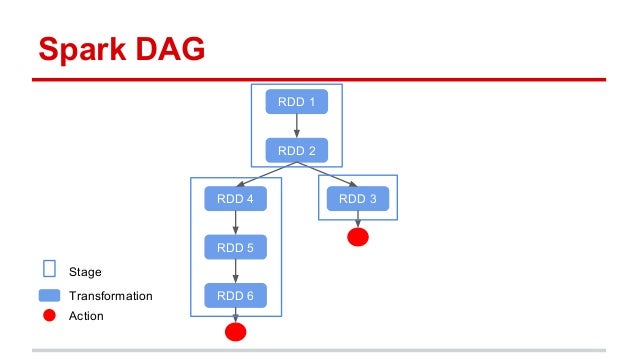
#
# Source: https://image.slidesharecdn.com/mapreducevsspark-150512052504-lva1-app6891/95/map-reduce-vs-spark-16-638.jpg?cb=1431408380
# - [PySpark API](https://spark.apache.org/docs/latest/api/python/pyspark.html)
# ## SparkContext
#
# A SparkContext represents the connection to a Spark cluster, and can be used to create RDDs, accumulators and broadcast variables on that cluster. Here we set it up to use local nodes - the argument `locals[*]` means to use the local machine as the cluster, using as many worker threads as there are cores. You can also explicitly set the number of cores with `locals[k]` where `k` is an integer.
#
# With Saprk 2.0 onwards, there is also a SparkSession that manages DataFrames, which is the preferred abstraction for working in Spark. However DataFrames are composed of RDDs, and it is still necesaary to understand how to use and mainpulate RDDs for low level operations.
# Depending on your setup, you many have to import SparkContext. This is not necessary in our Docker containers as we will be using `livy`.
# Start spark
from pyspark import SparkContext
sc = SparkContext.getOrCreate()
from pyspark.sql import SparkSession
spark = (
SparkSession.builder
.master("local")
.appName("BIOS-823")
.config("spark.executor.cores", 4)
.getOrCreate()
)
# Use this instead if using Spark on vm-manage containers.
#
# ```python
# # %%spark
# ```
import numpy as np
# Version
sc.version
# Number of workers
sc.defaultParallelism
# Data in an RDD is distributed across partitions. It is most efficient if data does not have to be transferred across partitions. We can see the default minimumn number of partitions, and the actual number in an RDD later.
sc.defaultMinPartitions
# ## Resilient Distributed Datasets (RDD)
#
#
# ### Creating an RDD
#
# The RDD (Resilient Distributed Dataset) is a data storage abstraction - you can work with it as though it were single unit, while it may actually be distributed over many nodes in the computing cluster.
# #### A first example
# Distribute the data set to the workers
xs = sc.parallelize(range(10))
xs
xs.getNumPartitions()
# Return all data within each partition as a list. Note that the glom() operation operates on the distributed workers without centralizing the data.
xs.glom().collect()
# Only keep even numbers
xs = xs.filter(lambda x: x % 2 == 0)
xs
# Square all elements
xs = xs.map(lambda x: x**2)
xs
# Execute the code and return the final dataset
xs.collect()
# Reduce also triggers a calculation
xs.reduce(lambda x, y: x+y)
# #### A common Spark idiom chains mutiple functions together
(
sc.parallelize(range(10))
.filter(lambda x: x % 2 == 0)
.map(lambda x: x**2)
.collect()
)
# Actions and transforms
# ----
# A **transform** maps an RDD to another RDD - it is a lazy operation. To actually perform any work, we need to apply an **action**.
# ### Actions
x = sc.parallelize(np.random.randint(1, 6, 10))
x.collect()
x.take(5)
x.first()
x.top(5)
x.takeSample(True, 15)
x.count()
x.distinct().collect()
x.countByValue()
x.sum()
x.max()
x.mean()
x.stats()
# #### Reduce, fold and aggregate actions
# **From the API**:
#
# - reduce(f)
#
# > Reduces the elements of this RDD using the specified commutative and associative binary operator. Currently reduces partitions locally.
#
# - fold(zeroValue, op)
#
# > Aggregate the elements of each partition, and then the results for all the partitions, using a given associative function and a neutral “zero value.”
#
# > The function op(t1, t2) is allowed to modify t1 and return it as its result value to avoid object allocation; however, it should not modify t2.
#
# > This behaves somewhat differently from fold operations implemented for non-distributed collections in functional languages like Scala. This fold operation may be applied to partitions individually, and then fold those results into the final result, rather than apply the fold to each element sequentially in some defined ordering. For functions that are not commutative, the result may differ from that of a fold applied to a non-distributed collection.
#
# - aggregate(zeroValue, seqOp, combOp)
#
# > Aggregate the elements of each partition, and then the results for all the partitions, using a given combine functions and a neutral “zero value.”
#
# > The functions op(t1, t2) is allowed to modify t1 and return it as its result value to avoid object allocation; however, it should not modify t2.
#
# > The first function (seqOp) can return a different result type, U, than the type of this RDD. Thus, we need one operation for merging a T into an U and one operation for merging two U
#
# Notes:
#
# - All 3 operations take a binary op with signature op(accumulator, operand)
x = sc.parallelize(np.random.randint(1, 10, 12))
x.collect()
# **max** using reduce
x.reduce(lambda x, y: x if x > y else y)
# **sum** using `reduce`
x.reduce(lambda x, y: x+y)
# **sum** using fold
x.fold(0, lambda x, y: x+y)
# **prod** using reduce
x.reduce(lambda x, y: x*y)
# **prod** using fold
x.fold(1, lambda x, y: x*y)
# **sum** using aggregate
x.aggregate(0, lambda x, y: x + y, lambda x, y: x + y)
# **count** using aggregate
x.aggregate(0, lambda acc, _: acc + 1, lambda x, y: x+y)
# **mean** using aggregate
sum_count = x.aggregate([0,0],
lambda acc, x: (acc[0]+x, acc[1]+1),
lambda acc1, acc2: (acc1[0] + acc2[0], acc1[1]+ acc2[1]))
sum_count[0]/sum_count[1]
# **Warning**: Be very careful with fold and aggregate - the `zero` value must be "neutral". The behavior can be different from Python's reduce with an initial value.
#
# This is because there are two levels of operations that use the `zero` value - first locally, then globally.
xs = x.collect()
xs = np.array(xs)
xs
sum(xs)
# **Exercise**: Explain the results shown below:
from functools import reduce
reduce(lambda x, y: x + y, xs, 1)
x.fold(1, lambda acc, val: acc + val)
x.aggregate(1, lambda x, y: x + y, lambda x, y: x + y)
# **Exercise**: Explain the results shown below:
reduce(lambda x, y: x + y**2, xs, 0)
np.sum(xs**2)
x.fold(0, lambda x, y: x + y**2)
x.aggregate(0, lambda x, y: x + y**2, lambda x, y: x + y)
# **Exercise**: Explain the results shown belwo:
x.fold([], lambda acc, val: acc + [val])
seqOp = lambda acc, val: acc + [val]
combOp = lambda acc, val: acc + val
x.aggregate([], seqOp, combOp)
# ### Transforms
x = sc.parallelize([1,2,3,4])
y = sc.parallelize([3,3,4,6])
x.map(lambda x: x + 1).collect()
x.filter(lambda x: x%3 == 0).collect()
# #### Think of flatMap as a map followed by a flatten operation
x.flatMap(lambda x: range(x-2, x)).collect()
x.sample(False, 0.5).collect()
# #### Set-like transformss
y.distinct().collect()
x.union(y).collect()
x.intersection(y).collect()
x.subtract(y).collect()
x.cartesian(y).collect()
# Note that flatmap gets rid of empty lists, and is a good way to ignore "missing" or "malformed" entires.
def conv(x):
try:
return [float(x)]
except:
return []
# +
s = "Thee square root of 3 is less than 3.14 unless you divide by 0".split()
x = sc.parallelize(s)
x.collect()
# -
x.map(conv).collect()
x.flatMap(conv).collect()
# Working with key-value pairs
# ----
#
# RDDs consissting of key-value pairs are required for many Spark operatinos. They can be created by using a function that returns an RDD composed of tuples.
data = [('ann', 'spring', 'math', 98),
('ann', 'fall', 'bio', 50),
('bob', 'spring', 'stats', 100),
('bob', 'fall', 'stats', 92),
('bob', 'summer', 'stats', 100),
('charles', 'spring', 'stats', 88),
('charles', 'fall', 'bio', 100)
]
rdd = sc.parallelize(data)
rdd.keys().collect()
rdd.collect()
# #### Functions `ByKey`
# Sum values by key
(
rdd.
map(lambda x: (x[0], x[3])).
reduceByKey(lambda x, y: x + y).
collect()
)
# Running list of values by key
(
rdd.
map(lambda x: ((x[0], x[3]))).
aggregateByKey([], lambda x, y: x + [y], lambda x, y: x + y).
collect()
)
# Average by key
(
rdd.
map(lambda x: ((x[0], x[3]))).
aggregateByKey([], lambda x, y: x + [y], lambda x, y: x + y).
map(lambda x: (x[0], sum(x[1])/len(x[1]))).
collect()
)
# Using a different key
(
rdd.
map(lambda x: ((x[2], x[3]))).
aggregateByKey([], lambda x, y: x + [y], lambda x, y: x + y).
map(lambda x: (x[0], sum(x[1])/len(x[1]))).
collect()
)
# ### Using key-value pairs to find most frequent words in Ulysses
#
# Note: This part assumes that there we are using HDFS.
#
# If you want to install locally, there are tutorials on the web. For Macbook I followed this [guide](https://www.quickprogrammingtips.com/big-data/how-to-install-hadoop-on-mac-os-x-el-capitan.html).
# +
hadoop = sc._jvm.org.apache.hadoop
fs = hadoop.fs.FileSystem
conf = hadoop.conf.Configuration()
path = hadoop.fs.Path('./data/texts')
for f in fs.get(conf).listStatus(path):
print(f.getPath())
# -
ulysses = sc.textFile('./data/texts/Portrait.txt')
# Note that we can also read in entire docs as the values.
docs = sc.wholeTextFiles('./data/texts/')
docs.keys().collect()
ulysses.take(10)
import string
def tokenize(line):
table = dict.fromkeys(map(ord, string.punctuation))
return line.translate(table).lower().split()
words = ulysses.flatMap(lambda line: tokenize(line))
words.take(10)
words = words.map(lambda x: (x, 1))
words.take(10)
counts = words.reduceByKey(lambda x, y: x+y)
counts.take(10)
counts.takeOrdered(10, key=lambda x: -x[1])
# ### Word count chained version
(
ulysses.flatMap(lambda line: tokenize(line))
.map(lambda word: (word, 1))
.reduceByKey(lambda x, y: x + y)
.takeOrdered(10, key=lambda x: -x[1])
)
# ### Avoiding slow Python UDF tokenize
#
# We will see how to to this in the DataFrames notebook.
# ### CountByValue Action
# If you are sure that the results will fit into memory, you can get a dacitonary of counts more easily.
wc = (
ulysses.
flatMap(lambda line: tokenize(line)).
countByValue()
)
wc['the']
# Persisting data
# ----
#
# The `top_word` program will repeat ALL the computations each time we take an action such as `takeOrdered`. We need to `persist` or `cache` the results - they are similar except that `persist` gives more control over how the data is retained.
counts.is_cached
counts.persist()
counts.is_cached
counts.takeOrdered(5, lambda x: -x[1])
counts.take(5)
counts.takeOrdered(5, lambda x: x[0])
counts.keys().take(5)
counts.values().take(5)
count_dict = counts.collectAsMap()
count_dict['circle']
# #### Using cache instead of persist
counts.unpersist()
counts.is_cached
counts.cache()
counts.is_cached
# ### Merging key, value datasets
#
# We will build a second counts key: value RDD from another of Joyce's works - Portrait of the Artist as a Young Man.
portrait = sc.textFile('./data/texts/Portrait.txt')
counts1 = (
portrait.flatMap(lambda line: tokenize(line))
.map(lambda x: (x, 1))
.reduceByKey(lambda x,y: x+y)
)
counts1.persist()
# #### Combine counts for words found in both books
joined = counts.join(counts1)
joined.take(5)
# #### sum counts over words
s = joined.mapValues(lambda x: x[0] + x[1])
s.take(5)
# #### average counts across books
avg = joined.mapValues(lambda x: np.mean(x))
avg.take(5)
| notebooks/C02_Sprak_Low_Level_API.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import cooler
import bioframe
from cooltools.sandbox import obs_over_exp_cooler
import cooltools
from scipy.sparse import coo_matrix
from matplotlib import colors
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# clr = cooler.Cooler("./ALV-repo/Hap1-WT-combined.mcool::/resolutions/500000")
# download test data
# this file is 145 Mb, and may take a few seconds to download
cool_file = cooltools.download_data("HFF_MicroC", cache=True, data_dir='./')
print(cool_file)
# Load a Hi-C map at a 1Mb resolution from a cooler file.
clr = cooler.Cooler('./test.mcool::/resolutions/1000000')
# Use bioframe to fetch the genomic features from the UCSC.
hg38_chromsizes = bioframe.fetch_chromsizes('hg38')
hg38_cens = bioframe.fetch_centromeres('hg38')
# create a view with chromosome arms using chromosome sizes and definition of centromeres
hg38_arms = bioframe.make_chromarms(hg38_chromsizes, hg38_cens)
# select only those chromosomes available in cooler
hg38_arms = hg38_arms[hg38_arms.chrom.isin(clr.chromnames)].reset_index(drop=True)
hg38_arms
# calculate full expected (cis + trans)
expected_df = obs_over_exp_cooler.expected_full(
clr,
view_df=hg38_arms,
smooth_cis=False,
aggregate_trans=True,
expected_column_name="expected",
nproc=4,
)
# collect obs/exp for chunks of pixel table (in memory for 1Mb cooler)
results = []
for oe_chunk in obs_over_exp_cooler.obs_over_exp_generator(
clr,
expected_df,
view_df=hg38_arms,
expected_column_name="expected",
oe_column_name='oe',
chunksize=1_000_000,
):
results.append(oe_chunk)
# concat chunks into single DataFrame - res_df - is a new pixel table - sparse matrix
res_df = pd.concat(results, ignore_index=True)
res_df.head()
# res_df: sparse matrix -> dense matrix for plotting
N = len(clr.bins())
oe = coo_matrix(
(res_df["oe"], (res_df["bin1_id"], res_df["bin2_id"])),
shape=(N,N),
).toarray()
# make it symmetric ...
oe = oe + oe.T
print(f"generated symmetrix obs/exp matrix of size {N} X {N}")
# +
# plot observed and stitched obs/exp side by side
istart, iend = 0, 327
obs = clr.matrix()[istart:iend, istart:iend]
obs_exp = oe[istart:iend, istart:iend]
f,axs = plt.subplots(1,2,figsize=(14,10))
img = axs[0].imshow(
obs,
interpolation="none",
cmap="YlOrRd",
norm=colors.LogNorm(vmin=0.00005,vmax=0.01)
)
plt.colorbar(img,ax=axs[0],orientation="horizontal")
img = axs[1].imshow(
obs_exp,
interpolation="none",
cmap="coolwarm",
norm=colors.LogNorm(vmin=0.4,vmax=2.5)
)
plt.colorbar(img,ax=axs[1],orientation="horizontal")
# -
# ### Try higher resolution data and write directly into cooler
# try 10kb ...
clr = cooler.Cooler('./test.mcool::/resolutions/10000')
# generate bins table with weights=1, and NaN for bad bins ...
bins_oe = clr.bins()[:]
_bad_mask = bins_oe["weight"].isna()
bins_oe["weight"] = 1.
bins_oe.loc[_bad_mask,"weight"] = np.nan
# re-calculate full expected (cis + trans) at higher resolution
expected_df = obs_over_exp_cooler.expected_full(
clr,
view_df=hg38_arms,
smooth_cis=False,
aggregate_trans=True,
expected_column_name="expected",
nproc=4,
)
# setup a generator (lazy) of obs/exp pixels
oe_pixels_stream = obs_over_exp_cooler.obs_over_exp_generator(
clr,
expected_df,
view_df=hg38_arms,
expected_column_name="expected",
oe_column_name='oe',
chunksize=10_000_000
)
# write oe_pixels_stream into cooler - with custom column "oe" (can do "count":float for higlass)
cooler.create_cooler(
cool_uri = "fun.cool",
bins = bins_oe,
pixels = oe_pixels_stream,
columns=["oe"],
dtypes={"oe":np.float64},
)
# +
# plot observed and stitched obs/exp side by side directly from the new cooler
istart, iend = 23_000, 25_000
obs = clr.matrix()[istart:iend, istart:iend]
obs_exp = cooler.Cooler("fun.cool").matrix(field="oe")[istart:iend, istart:iend]
f,axs = plt.subplots(1,2,figsize=(14,10))
img = axs[0].imshow(
obs,
interpolation="none",
cmap="YlOrRd",
norm=colors.LogNorm(vmin=0.00005,vmax=0.01)
)
plt.colorbar(img,ax=axs[0],orientation="horizontal")
# make sure zeros are displayed as "lowest" obs/exp according to the colormap
cm = plt.cm.get_cmap("coolwarm")
cm.set_under(cm(0))
img = axs[1].imshow(
obs_exp+10**-8,
interpolation="none",
cmap=cm,
norm=colors.LogNorm(vmin=0.4,vmax=2.5)
# add color to the "under" - to avoid adding "floor" to obs_exp
#
)
plt.colorbar(img,ax=axs[1],orientation="horizontal")
| cooltools/sandbox/observed_over_expected_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <div class="alert block alert-info alert">
#
# # <center> Scientific Programming in Python
#
# ## <center><NAME><br>Bonn-Rhein-Sieg University of Applied Sciences<br>Sankt Augustin, Germany
#
# # <center> Pandas
# #### <center> (Reading in, manipulating, analyzing and visualizing datasets)</center>
#
# **Source**: https://pandas.pydata.org/
# <br><br>
#
# "...providing fast, flexible, and expressive data structures designed to make working with “relational” or “labeled” data both easy and intuitive. It aims to be the fundamental high-level building block for doing practical, real world data analysis in Python." -- http://pandas.pydata.org/pandas-docs/stable/
#
# - Tabular data with heterogeneously-typed columns, (CSV, SQL, or Excel spreadsheet)
# - Ordered and unordered time series data.
# - Arbitrary matrix data with row and column labels
#
#
# **Significant things to note**:
# - Allows you to operate in any diretion on your data (i.e. by rows or by columns)
# - Database experts will find this interesting
# - SQL: maniplate data by rows (i.e. row-focused)
# - Columnar databases: manipulate data by columns (i.e. column-focused)
# - Operate data on data using 1-2 lines of code
#
#
# - Data structures
# - Series - 1 dimensional data
# - DataFrame - 2 dimensional data
#
#
# - Index data
# - can organize your data quickly and logically (e.g. based on calendar dates
# - can handle missing data
#
#
# - Missing data
# - NaN
# - mean
# - fill forward and backwards
#
# #### Basic Functionalities to Know
#
# https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html
# 1. Head and tail: (https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html#head-and-tail)
# 1. Attributes and underlying data (relevant for the numpy lecture): (https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html#attributes-and-underlying-data)
# 1. Descriptive statistics: (https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html#descriptive-statistics)
# 1. Reindexing and altering labels: (https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html#reindexing-and-altering-labels)
# 1. Iteration: (https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html#iteration)
# 1. Sorting: (https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html#sorting)
# 1. Copying: (https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html#copying)
# 1. dtypes: (https://pandas.pydata.org/pandas-docs/stable/user_guide/basics.html#dtypes)
#
# #### Underlying libraries (used but not seen)
# 1. Numpy
# 2. Matplotlib
#
# <br>
#
# #### Note about citations (i.e. referencing):
#
# **For citing Pandas**: (via https://pandas.pydata.org/about/citing.html - modify for your Pandas version)
#
# **Bibtex**
#
# @software{reback2020pandas,
# author = {The pandas development team},
# title = {pandas-dev/pandas: Pandas},
# month = feb,
# year = 2020,
# publisher = {Zenodo},
# version = {latest},
# doi = {10.5281/zenodo.3509134},
# url = {https://doi.org/10.5281/zenodo.3509134}
# }
#
# @InProceedings{mckinney-proc-scipy-2010,
# author = {{<NAME>}c{K}inney},
# title = {{D}ata {S}tructures for {S}tatistical {C}omputing in {P}ython},
# booktitle = {{P}roceedings of the 9th {P}ython in {S}cience {C}onference},
# pages = {56 - 61},
# year = {2010},
# editor = {{S}<NAME> {W}alt and {J}arrod {M}illman},
# doi = {10.25080/Majora-92bf1922-00a}
# }
#
# <br>
#
# #### Sources
# 1. The pandas development team, pandas-dev/pandas: Pandas, Zenodo, 2020, https://doi.org/10.5281/zenodo.3509134, visited on June 7, 2021
#
# 2. <NAME>., 2010, June. Data structures for statistical computing in python. In Proceedings of the 9th Python in Science Conference, van der Walt, S. & Millman, J. (Eds.), vol. 445 pp. 51-56).
#
#
# #### Additional sources
#
# 1. <NAME>, Python for Data Analysis; Data Wrangling with Pandas, Numpy and Ipython, O'Reilly, Second Edition, 2018.
#
# <hr style="border:2px solid gray"></hr>
import pandas as pd
# ## Pandas Series
#
# Series contain two components:
# 1. one-dimensional array-like object that contains a sequence of data values
# 2. an associated array of data labels (i.e. 'index')
#
# Note: indexes start at '0'
#
# #### Creating
#
# Create a series that contains 5 integers:
series_data = pd.Series([5, 10, 15, 20, 25])
series_data
# #### Indexes
# Now, let us add some indexes to help lables the integers:
series_data = pd.Series([5, 10, 15, 20, 25], index=['d', 'e', 'a', 'simulation', 'average'])
series_data
# We can alter these indexes at any time.
series_data.index = ['Norway', 'Italy', 'Germany', 'simulation', 'average']
series_data
# #### Accessing the series
#
# Access only the values:
series_data.values
# Access the data via an index label:
series_data['simulation']
# Or by a position:
series_data[3]
# #### Using operators
series_data**2
# What happens when one of the series has missing data?
#
# Let's create an alternate series that has the **Italian data missing**, and then **add them** to the original series:
# +
series_data_missing = pd.Series([5, 10, 20, 25], index=['Germany', 'Norway', 'simulation', 'average'])
series_data + series_data_missing
# -
# #### Filtering and Sorting
#
# Filter the data:
series_data[series_data >= 15]
# Sorting a series by its index:
series_data.sort_index()
# Notice the sorting goes by:
# 1. Capital case letters
# 1. Lower case letters
# Sorting a series by data values:
series_data.sort_values()
# ---
# ## Dataframes
# - dataframes represents a **rectangular, ordered** table of data (numbers, strings, etc.)
#
# - just like you are familiar with in a spreedsheet
#
# Let's create a simple user function that will allow us to reset our example dataframe as needed
# 1. First create a dictionary
# 2. Convert the dictionary to a dataframe
def dict2dataframe():
'''Create a dataframe 'by hand' using a dictionary that has equal lengths'''
data = {'group': ['Deichkind', 'Die Fantastischen Vier', 'Seeed', '<NAME>'],
'year': [2015, 2106, 2017, 2018],
'attendence (x1000)': [50, 60, 70, 90]}
dataframe = pd.DataFrame(data) # convert the dictionary to a pandas' dataframe
return dataframe
example_df = dict2dataframe()
example_df
# Alter these indexes in the same way we did for the series:
example_df.index = ['band 1', 'band 2', 'band 3', 'band 4']
example_df
# Note that index don't need to be unique for each row, but this can cause problems (for example, later we will delete based on the index label).
#
# Assign `band 1` to the first two index positions
example_df.index = ['band 1', 'band 1', 'band 3', 'band 4']
example_df
# #### Inserting columns
#
# Insert columns (simple):
example_df['quality'] = ['good', 'excellent', 'good', 'average']
example_df
# Inserting a new column, and fill it using 'NaN':
example_df['number of total concerts'] = pd.Series(data='NaN')
example_df
# **Inserting a new row**:
example_df = example_df.append({'group':'Scorpions', 'year':1965, 'attendence (x1000)':100},
ignore_index=True)
example_df
# Notice
# 1. how the index change to integers.
# 1. how `NaN` is added to the columns not specified (i.e. to `quality` and `number of total concerts`)
# ### Dropping data entries
# - pandas.drop will **drop columns** and **rows** using the **axis** keyword
# - `axis='rows'` ;`axis=0` ; `axis='index'`
# - `axis='columns'` ; `axis=1`
# #### Removing columns
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.drop.html
# - axis='columns' ; axis=1
example_df = example_df.drop(['year'], axis='columns')
example_df
# Interestingly, you don't need the list brackets:
example_df = example_df.drop('attendence (x1000)', axis='columns')
example_df
example_df = example_df.drop(['quality', 'number of total concerts'], axis='columns')
example_df
# #### Removing rows
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.drop.html
# - `axis='row'` ; `axis='rows'` ;`axis=0` ; `axis='index'`
example_df = example_df.drop([0], axis='index')
example_df
# As with columns, not specifing the list brackets also works:
example_df = example_df.drop(1, axis='index')
example_df
example_df = example_df.drop([3, 4], axis='rows')
example_df
# **Additional examples**
#
# Indexes that are strings
example_df = dict2dataframe()
example_df.index = ['band 1', 'band 2', 'band 3', 'band 4']
example_df
example_df.drop(['band 2', 'band 3'], axis='rows')
# What happens if you have rows with the same index?
#
# Let's reset, and set two rows as `band 3`:
example_df = dict2dataframe()
example_df.index = ['band 1', 'band 3', 'band 3', 'band 4']
example_df
example_df = example_df.drop(['band 3'])
example_df
# ---
# ## Accessing, selecting and filtering data
# - there are many ways to do this (df: dataframe)
# - `df[val]` and `df[[]]`
# - `df.loc[val]`: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.loc.html
# - `df.loc[row_val, col_val]`
# - `df.iloc[row_index, col_index]`: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.iloc.html#pandas.DataFrame.iloc
# - and more
#
# **Suggestion** - choose one method like `df.loc` and learn it first
# - Reset the example, and
# - Reindex the dataframe:
# +
example_df = dict2dataframe()
example_df.index = ['band 1', 'band 2', 'band 3', 'band 4']
example_df
# -
# #### Accessing/Selecting rows (by the index)
#
# <font color='dodgerblue'>**Single row:**</font>
# - Using slicing `:`
#
# via index names:
example_df['band 1':'band 1']
# via index numbers:
example_df[0:1]
# Alternative
# - `loc` with double `[[ ]]` (passing a list)
example_df.loc[['band 1']]
# <font color='dodgerblue'>**Multiple rows**</font>
#
# - Using slicing `:`
#
# via index names:
example_df['band 1':'band 3']
# via index numbers:
example_df[0:3]
# Alternative apporaches:
# - `loc` with double `[[ ]]`
#
# **Notice**: How we skip `band 2` in the following, so it is really is not a range.
example_df.loc[['band 1', 'band 3']]
# #### Access a specific cell (index, labels)
# +
example_df.loc['band 3', 'group']
# -
# Or by index number
# - `iloc`
example_df.iloc[2, 0]
# #### Substitute a value at a specific cell
example_df.loc['band 3', 'number of total concerts'] = 10000
example_df
# ### Accessing/Selecting columns
# #### Accessing columns (by label)
#
# <font color='dodgerblue'>Single column:</font>
example_df['group']
# <font color='dodgerblue'>Multiple column:</font>
#
# - the double `[[ ]]` (passing a list to the dataframe)
example_df[['group', 'year']]
# Alternative approaches
# - the `df.columns` command
example_df[example_df.columns[0:2]]
# - `loc`
#
# Notice that the rows designation is left as `:`, followed by a `,` and then the columns
example_df.loc[:, 'group':'attendence (x1000)']
example_df
# Now, let's putting everything together
# - slicing for rows (e.g. `'band 1':'band 3'`) and
# - slicing the columns (e.g. `'group':'attendence (x1000)'`)
example_df.loc['band 1':'band 3', 'group':'attendence (x1000)']
# ---
# ## Essential Functions
# ### Reminder about reordering the rows by their indexes
#
# - demonstrates what happens to a dataframe with multiple columns
#
# - `reindex`
# +
example_df = dict2dataframe()
example_df.index = ['band 1', 'band 2', 'band 3', 'band 4']
example_df
# -
example_df = example_df.reindex(['band 3', 'band 4', 'band 1', 'band 2'])
example_df
# ### Factorize catagorical data
# - This is something that is sometimes done when performing data analysis
# - e.g. Machine learning
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.factorize.html
# +
example_df = dict2dataframe()
example_df.index = ['band 1', 'band 2', 'band 3', 'band 4']
example_df['quality'] = ['good', 'excellent', 'good', 'average']
example_df
# -
codes, uniques = example_df['quality'].factorize()
codes
uniques
example_df['quality_numeric'] = codes
example_df
# ### Iterate over rows
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.iterrows.html#pandas-dataframe-iterrows
for index, row in example_df.iterrows():
print(f"Index: {index} ; Group: {row['group']}")
print()
# ### tolists
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.tolist.html
#
# First let's see what `dataframe.columns` does
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.columns.html
example_df.columns
# Convert column names to a list
example_df.columns.tolist()
# ---
#
# ## Combining dataframes
# - take the columns from different dataframes and put them together into a single collumn
# 1. concat: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.concat.html#pandas.concat
# 2. append
#
# **Example** - student grades on homeworks
homework_1_grades = pd.DataFrame({'student': [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13],
'homework 1': [63.0, 76.0, 76.0,
76.0, 0.0, 0.0,
88.0, 86.0, 76.0,
86.0, 70.0, 0.0, 80.0]})
homework_1_grades
homework_2_grades = pd.DataFrame({'student': [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13],
'homework 2': [70.0, 73.0, 91.0,
89.0, 58.0, 0.0,
77.0, 91.0, 86.0,
78.0, 100.0, 61.5, 71.0]})
homework_2_grades
new_df_concat = pd.concat([ homework_1_grades['homework 1'], homework_2_grades['homework 2'] ], axis='rows')
new_df_concat
type(new_df_concat)
# Alternative approach using 'append'
new_df_append = homework_1_grades['homework 1'].append(homework_2_grades['homework 2'])
new_df_append
type(new_df_append)
# - Combine two dataframe based on a common keys.
# 1. merge: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.merge.html#pandas.merge
#
# (This is just one example from many ways to do this, including when the keys might not be shared.)
pd.merge(homework_1_grades, homework_2_grades, on='student')
# ---
# ## Math operators
#
# Let's perform some math on a dataframe.
#
# Dataframe:
# - 5 rectangles that are defined by
# - length
# - height
# +
rectangles_dict = {'length': [0.1, 9.4, 6.2, 3.8, 9.4],
'height': [8.7, 6.2, 9.4, 5.6, 3.3]}
rectangles_data = pd.DataFrame(rectangles_dict)
rectangles_data
# -
# #### Operate on all columns (e.g. dividing by 10)
rectangles_data/10
# #### Operatation using two columns (e.g. for the area of a rectangle)
rectangles_data['length'] * rectangles_data['height']
# #### Create a new column based on math using other columns
rectangles_data['area'] = rectangles_data['length'] * rectangles_data['height']
rectangles_data
# ### Descriptive statistics
#
# Using **python built-in functions** (e.g. max, min) on a Pandas dataframe:
max(rectangles_data['area'])
min(rectangles_data['area'])
# Notice above: how the dataframe is given within the parentheses.
# Using **pandas functions**
#
# - count (https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.count.html)
# - sum (https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.sum.html)
# - median (https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.median.html)
# - std (https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.std.html)
# - var (https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.var.html)
# - max (https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.max.html)
# - min (https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.min.html)
# - correlation analysis (https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.corr.html)
# - and many more
#
#
#
# **Notice below** how **the dataframe is given first**, followed by the function (e.g. `df.max()`)
#
# On all dataframe columns:
rectangles_data.max()
# One a specific column:
rectangles_data['area'].max()
# `idxmin` and `idxmax`
#
# "Return **index** of first occurrence of maximum over requested axis."[1]
#
# 1. https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.idxmax.html
rectangles_data
maximum_index = rectangles_data['area'].idxmax()
maximum_index
rectangles_data.loc[maximum_index]['area']
rectangles_data.loc[maximum_index]['length']
# But **note** it is the **FIRST OCCURANCE**
# - Returns the row with a length=9.4, width=6.2 and an area=58.28 (i.e. index = 1)
# - It does NOT return values for the rows that contain
# - length=6.2, width=9.4 and an area=58.28 (i.e. index=2)
rectangles_data['area'].count()
rectangles_data['area'].mean()
rectangles_data['area'].std()
# #### Moving averages (data smoothing)
# - https://en.wikipedia.org/wiki/Moving_average
#
# - rolling mean of data via pandas
#
# https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.rolling.html?highlight=rolling#pandas.DataFrame.rolling
rectangles_data['area moving avg'] = rectangles_data['area'].rolling(window=2, win_type=None).mean()
rectangles_data
# ### Unique values
# - Unique values
rectangles_data['area'].unique()
# - Unique values and count their occurance
rectangles_data['area'].value_counts()
# #### How to using other libraries (e.g. statistics)
# - Make sure you have a good reason to do this (i.e. be consistent)
# - Notice that the format is similar to using a built-in function (shown above)
import statistics
statistics.mean(rectangles_data['area'])
# ### Sorting dataframes
# - similiar to how the series was done above, but with a twist
# - https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.sort_values.html
# - `df.sort_values()`
#
# Our original, unsorted dataframe:
rectangles_data
# - sort by a single column's values
rectangles_data.sort_values(by='area')
# - sort by multiple columns
# - consecutively done
## rows index 1 and 2 should switch due to length value
rectangles_data.sort_values(by=['area', 'length'])
# ### Filter by boolean operators
rectangles_data
rectangles_data['area'] > 7.0
# #### return a dataframe based on one boolean condition
rectangles_data[rectangles_data['area'] > 7.0]
# #### return a dataframe based on multiple boolean condition
rectangles_data[ (rectangles_data['area'] > 7.0) & (rectangles_data['area'] < 50.0) ]
# ---
# ## Data from a csv-formatted file
#
# - The example CSV data file used below can be found at https://github.com/karlkirschner/2020_Scientific_Programming/blob/master/data_3d.csv
# +
## For Colabs
## In order to upload data
#from google.colab import files
#uploaded = files.upload()
# -
# !head data_3d.csv --lines=10
# For files without a header you can:
# 1. have pandas assign an index value as the header (e.g. 1 2 3)
df = pd.read_csv('data_3d.csv', header=None, sep=',')
df
# 2. Read in a csv file, using the first row (i.e. 0) as the header, with a comma separator
#
df = pd.read_csv('data_3d.csv', header=0, sep=',')
df
# 3. Assign the headers yourself
# - use `skiprows` if the first row labels are present, as in this example
df = pd.read_csv('data_3d.csv', sep=',', skiprows=1, names=['header 1', 'header 2', 'average'])
df
# #### Save data to a new csv file, printing out to the first decimal place
df.to_csv('pandas_out.csv',
sep=',', float_format='%.1f',
index=False, encoding='utf-8')
# ## Visualizing the data via Pandas plotting
#
# https://pandas.pydata.org/pandas-docs/version/0.23.4/generated/pandas.DataFrame.plot.html
#
#
# #### Types of plots
#
# The type of plot is specified through the pandas.DataFrame.plot's `kind` keyword.
#
# 1. ‘line’ : line plot (default)
# 1. ‘bar’ : vertical bar plot
# 1. ‘barh’ : horizontal bar plot
# 1. ‘hist’ : histogram
# 1. ‘box’ : boxplot
# 1. ‘kde’ : Kernel Density Estimation plot
# 1. ‘density’ : same as ‘kde’
# 1. ‘area’ : area plot
# 1. ‘pie’ : pie plot
# 1. ‘scatter’ : scatter plot
# 1. ‘hexbin’ : hexbin plot
df = pd.read_csv('data_3d.csv', header=0, sep=',')
# In Pandas v. 1.1.0, xlabel and ylabel was introduced:
# +
## kind = line, box, hist, kde
df.plot(x='Time', y=['Exp', 'Theory'], kind='line',
xlabel='X-Label', ylabel='Y-Label',
title=['Example Plot: Exp', 'Example Plot: Theory'], fontsize=16, subplots=True)
# -
# An **alternative way** (also usuable with older Pandas version) that gives you a bit **more control** over, for example
# 1. the fontsize of different elements, for example
# - axis label
# - title
# 1. legend location
#
# This is similar to how matplotlib works.
# +
graphs = df.plot(x='Time', y=['Exp', 'Theory'], kind='line', fontsize=16, subplots=True)
graphs[0].set_title("Example Plot: Exp", fontsize=16)
graphs[0].set_ylabel("X-Label", fontsize=16)
graphs[0].legend(loc='upper left')
graphs[1].set_title("Example Plot: Theory", fontsize=16)
graphs[1].set_xlabel("X-Label", fontsize=16)
graphs[1].set_ylabel("Y-Label", fontsize=16)
graphs[1].legend(loc='upper left')
# -
# #### Combining multiple data lines onto one plot
# - demo by doing a rolling average on the above theory data and then plotting it.
df['Exp Rolling'] = df['Exp'].rolling(window=4, win_type=None).mean()
df['Theory Rolling'] = df['Theory'].rolling(window=4, win_type=None).mean()
df
df.plot(x='Time', y=['Theory', 'Theory Rolling'], kind='line',
title='Example Plot', fontsize=16)
# ---
# # Side Topics
#
# ## Pandas to Latex
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.to_latex.html
print(df.to_latex(index=False))
# ***
# ## Import Data from a European data csv file
# (e.g. decimal usage: 10.135,11)
# +
## CSV data file acan be found at
## https://github.com/karlkirschner/2020_Scientific_Programming/blob/master/data_eu.csv
## For Colabs
## In order to upload data
#from google.colab import files
#uploaded = files.upload()
# -
# !head data_eu.csv --lines=10
df = pd.read_csv('data_eu.csv', decimal=',', thousands='.', sep=';')
df.columns
df['Value']
| pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("weather.csv")
df
df.pivot(index="date",columns="city",values="humidity")
df.pivot(index="humidity",columns="city")
df = pd.read_csv("weather2.csv")
df
df.pivot_table(index="city",columns="date")
df.pivot_table(index="city",columns="date",aggfunc="count")
df.pivot_table(index="city",columns="date",margins=True)
df = pd.read_csv("weather3.csv")
df
df['date'] = pd.to_datetime(df['date'])
df
df.pivot_table(index=pd.Grouper(freq='M',key="date"),columns="city")
| path_of_ML/Pandas/pivot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import os
import sys
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# base = !pwd
base = os.path.dirname(os.path.dirname(base[0]))
p = os.path.join(base, 'neural-straight/nstraight/')
if p not in sys.path:
sys.path.append(p)
data_dir = os.path.join(base, 'neural-straight/data/')
if data_dir not in sys.path:
sys.path.append(data_dir)
from data.datasets import MovieSet
from curvature.curvature_schemas import CurvaturePixels, CurvatureResponse, TemporalFilter, SpatialRescale
from data import data_schemas as data
from utils.utils import get_trial_idx, type_object_movie
from visualization.visualize import scatter_brain_areas, brain_area_curvature, histogram_object_types, scatter_pix_responses
# -
# ### Pixel vs Responses in V1 of Naturalistic stimuli
# +
scan = data.MovieScan.proj() & 'animal_id > 20000'
rel_px_resp = (CurvaturePixels * (TemporalFilter.Butterworth() & dict(order =2) & scan) * SpatialRescale.No).proj('avg_pixel_curvature', 'median_pixel_curvature') * \
(CurvatureResponse * (TemporalFilter.Butterworth() & dict(order =2)) & scan).proj('avg_curvature', 'median_curvature', 'num_neurons') * data.ConditionClip.proj('movie_name')
df_px_resp = pd.DataFrame(rel_px_resp.fetch())
# +
sns.set_context('paper', font_scale=1.1)
sns.set_palette(sns.xkcd_palette(['grey', 'golden yellow']))
with sns.axes_style("ticks"):
g = sns.FacetGrid(df_px_resp, col = 'movie_name', aspect=1, col_wrap = 6)
g.map(sns.scatterplot, 'avg_pixel_curvature', 'avg_curvature')
for ax in g.axes.flatten():
ax.plot([12, 30], [12, 30], '--k')
ax.set_aspect('equal')
#ax.plot([15, 26], [15, 26], '--k')
sns.despine(trim=True)
g.fig.set_dpi(200)
# -
| notebooks/analysis_natural_movies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# # XGBoost による顧客離反分析 (Churn Analysis)
#
# ---
#
# ---
#
# ## コンテンツ
#
# 1. [背景](#1.背景)
# 1. [セットアップ](#2.セットアップ)
# 1. [データ](#3.データ)
# 1. [学習](#4.学習)
# 1. [ホスティング](#5.ホスティング)
# 1. [評価](#5-1.評価)
# 1. [推論エラーのコスト](#5-2.推論エラーのコスト)
# 1. [最適な閾値を探す](#5-3.最適な閾値を探す)
# 1. [エンドポイントの削除](#6.エンドポイントの削除)
# ---
#
# ## 1.背景
#
# _このノートブックで実施する内容は、[AWS blog post](https://aws.amazon.com/blogs/ai/predicting-customer-churn-with-amazon-machine-learning/)にも記載されています。_
#
# どのようなビジネスであっても、顧客を失うことは大きな損害です。もし、満足していない顧客を早期に見つけることができれば、そのような顧客をキープするためのインセンティブを提供できる可能性があるでしょう。このノートブックでは、満足していない顧客を自動で認識するために機械学習 (Machine Learning, ML) を利用する方法を説明します。このような顧客の離反分析は Customer Churn Prediction と呼ばれています。機械学習モデルは完璧な予測を行えないので、このノートブックでは予測のエラーが生じたときの相対的なコストを考慮して、機械学習を利用したときの成果を金額で評価します。
#
# ここでは、私達にとってなじみのある離反分析、携帯電話会社からの離反を取り上げます。携帯電話会社が、ある顧客が離反しそうと察知したら、その顧客にタイムリーにインセンティブを与えます。つまり、電話をアップグレードしたり、新しい機能を使えるようになったりして、引き続き携帯電話会社を使おうと思うかもしれません。インセンティブは、顧客が離反して再度獲得するまでにかかるコストよりもずっと小さいことが多いです。
#
#
#
# ---
#
# ## 2.セットアップ
#
# まず、このノートブックインスタンスに付与されている IAM role を `get_execution_role()` から取得しましょう。後ほど、SageMaker の学習やホスティングを行いますが、そこで IAM role が必要になります。そこで、ノートブックインスタンスの IAM role を、学習やホスティングでも利用します。
# 通常、role を取得するためにはAWS SDKを利用した数行のコードを書く必要があります。ここでは `get_execution_role()` のみで role を取得可能です。SageMaker Python SDK は、データサイエンティストが機械学習以外のコードを簡潔に済ませるために、このような関数を提供しています。
#
# + isConfigCell=true
# bucket = '<your_s3_bucket_name_here>'
# prefix = 'sagemaker/DEMO-xgboost-churn'
# Define IAM role
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
# -
# 以降で利用するライブラリをここで読み込んでおきます。
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import io
import os
import sys
import time
import json
from IPython.display import display
from time import strftime, gmtime
import sagemaker
from sagemaker.predictor import csv_serializer
# ---
# ## 3.データ
#
# 携帯電話会社は、どの顧客が最終的に離反したか、または、サービスを使い続けたかの履歴データをもっています。この履歴データに対して学習を行うことで、携帯電話会社の顧客離反を予想するモデルを構築します。モデルの学習が終わった後、任意の顧客のデータ (モデルの学習で利用したものと同じ情報を利用します)をモデルに入力すると、モデルはその顧客が離反しそうかどうかを予測します。もちろん、モデルは誤って予測することも考えられるので、将来を予測することはやはり難しいですが、そのような誤りに対応する方法も紹介します。
#
# ここで利用するデータセットは一般的に利用可能で、書籍 [Discovering Knowledge in Data](https://www.amazon.com/dp/0470908742/) の中で <NAME> が言及しているものです。そのデータセットは、著者によって University of California Irvine Repository of Machine Learning Datasets に提供されています。ここでは、そのデーセットをダウンロードして読み込んでみます。
#
# Jupyter notebook では、冒頭に `!` を入力することで、シェルコマンドを実行することができます。`wget` を利用してファイルをダウンロードし、`unzip` で展開します。
# !wget http://dataminingconsultant.com/DKD2e_data_sets.zip
# !unzip -o DKD2e_data_sets.zip
# 展開すると `./Data sets/` 以下にデータが展開されます。`churn.txt` を `pandas` を利用して読み込んでみます。 `pandas` は、表形式のデータを読み込んで、様々な加工ができるライブラリです。例えば、以下を実行すると表形式でのデータ表示が可能です。
churn = pd.read_csv('./Data sets/churn.txt')
pd.set_option('display.max_columns', 500)
churn
# データをみると 3,333 行のデータしかなく、現在の機械学習の状況から見ると、やや小さいデータセットです。各データのレコードは、ある米国の携帯電話会社の顧客のプロフィールを説明する21の属性からなります。その属性というのは、
#
# - `State`: 顧客が居住している米国州で、2文字の省略形で記載されます (OHとかNJのように)
# - `Account Length`: アカウントが利用可能になってからの経過日数
# - `Area Code`: 顧客の電話番号に対応する3桁のエリアコード
# - `Phone`: 残りの7桁の電話番号
# - `Int’l Plan`: 国際電話のプランに加入しているかどうか (yes/no)
# - `VMail Plan`: Voice mail の機能を利用しているかどうか (yes/no)
# - `VMail Message`: 1ヶ月の Voice mail のメッセージの平均長
# - `Day Mins`: 1日に通話した時間(分)の総和
# - `Day Calls`: 1日に通話した回数の総和
# - `Day Charge`: 日中の通話にかかった料金
# - `Eve Mins, Eve Calls, Eve Charge`: 夜間通話にかかった料金
# - `Night Mins`, `Night Calls`, `Night Charge`: 深夜通話にかかった料金
# - `Intl Mins`, `Intl Calls`, `Intl Charge`: 国際通話にかかった料金
# - `CustServ Calls`: カスタマーサービスに電話をかけた回数
# - `Churn?`: そのサービスから離反したかどうか (true/false)
#
# 最後の属性 `Churn?` は目的変数として知られ、MLのモデルで予測する属性になります。目的変数は2値 (binary) なので、ここで作成するモデルは2値の予測を行います。これは2値分類といわれます。
#
# それではデータを詳しく見てみます。
#
# まずはカテゴリデータごとにデータの頻度をみてみます。カテゴリデータは、`State`, `Area code`, `Phone`, `Int’l Plan`, `VMail Plan`, `Churn?`で、カテゴリを表す文字列や数値がデータとして与えられているものです。`pandas`ではある程度自動で、カテゴリデータを認識し、`object`というタイプでデータを保存します。以下では、`object` 形式のデータをとりだして、カテゴリごとの頻度を表示します。
#
# また `describe()`を利用すると各属性の統計量を一度に見ることができます。
# +
# Frequency tables for each categorical feature
for column in churn.select_dtypes(include=['object']).columns:
display(pd.crosstab(index=churn[column], columns='% observations', normalize='columns'))
# Histograms for each numeric features
display(churn.describe())
# %matplotlib inline
hist = churn.hist(bins=30, sharey=True, figsize=(10, 10))
# -
# データを見てみると以下のことに気づくと思います。
#
# - `State` の各頻度はだいたい一様に分布しています。
# - `Phone` はすべて同じ数値になっていて手がかりになりそうにありません。この電話番号の最初の3桁はなにか意味がありそうですが、その割当に意味がないのであれば、使うのは止めるべきでしょう
# - たった14%の顧客が離反しているので、インバランスなデータと言えるでしょうが、そこまで極端ではありません
# - 数値的な特徴量は都合の良い形で分布しており、多くは釣り鐘のようなガウス分布をしています。ただ、`VMail Message`は例外です。
# - `Area code` は数値データとみなされているようなので、非数値に変換しましょう
#
# さて、実際に`Phone`の列を削除して、`Area code`を非数値に変換します。
churn = churn.drop('Phone', axis=1)
churn['Area Code'] = churn['Area Code'].astype(object)
# それでは次に各属性の値を、目的変数の True か False か、にわけて見てみます。
# +
for column in churn.select_dtypes(include=['object']).columns:
if column != 'Churn?':
display(pd.crosstab(index=churn[column], columns=churn['Churn?'], normalize='columns'))
for column in churn.select_dtypes(exclude=['object']).columns:
print(column)
hist = churn[[column, 'Churn?']].hist(by='Churn?', bins=30)
plt.show()
# -
# データ分析の結果から、離反する顧客について、以下のような傾向が考えられます。
#
# - 地理的にもほぼ一様に分散している
# - 国際通話を利用している
# - VoiceMailを利用していない
# - 通話時間で見ると長い通話時間と短い通話時間の人に分かれる
# - カスタマーサービスへの通話が多い (多くの問題を経験した顧客ほど離反するというのは理解できる)
#
# 加えて、離反する顧客に関しては、`Day Mins` と `Day Charge` で似たような分布を示しています。しかし、話せば話すほど、通常課金されるので、驚くことではないです。もう少し深く調べてみましょう。`corr()` を利用すると相関係数を求めることができます。
display(churn.corr())
pd.plotting.scatter_matrix(churn, figsize=(12, 12))
plt.show()
# いくつかの特徴は互いに100%の相関をもっています。このような特徴があるとき、機械学習のアルゴリズムによっては全くうまくいかないことがあり、そうでなくても結果が偏ったりしてしまうことがあります。これらの相関の強いペアは削除しましょう。Day Mins に対する Day Charge、Night Mins に対する Night Charge、Intl Mins に対する Intl Charge を削除します。
churn = churn.drop(['Day Charge', 'Eve Charge', 'Night Charge', 'Intl Charge'], axis=1)
# ここまででデータセットの前処理は完了です。これから利用する機械学習のアルゴリズムを決めましょう。前述したように、数値の大小 (中間のような数値ではなく)で離反を予測するような変数を用意すると良さそうです。線形回帰のようなアルゴリズムでこれを行う場合は、複数の項(もしくはそれらをまとめた項)を属性として用意する必要があります。
#
# そのかわりに、これを勾配ブースティング木 (Gradient Boosted Tree)を利用しましょう。Amazon SageMaker は、マネージドで、分散学習が設定済みで、リアルタイム推論のためのホスティングも可能な XGBoost コンテナを用意しています。XGBoost は、特徴感の非線形な関係を考慮した勾配ブースティング木を利用しており、特徴感の複雑な関連性を扱うことができます。
#
# Amazon SageMaker の XGBoostは、csv または LibSVM 形式のデータを学習することができます。ここでは csv を利用します。csv は以下のようなデータである必要があります。
#
# - 1列目が予測対象のデータ
# - ヘッダ行はなし
#
# まずはじめに、カテゴリ変数を数値データに変換する必要があります。`get_dummies()` を利用すると数値データへの変換が可能です。
#
# そして、`Churn?_True`のデータを最初の列にもってきて、`Churn?_False.`, `Churn?_True.`のデータを削除した残りのデータをconcatenate (連結) します。
#
#
model_data = pd.get_dummies(churn)
model_data = pd.concat([model_data['Churn?_True.'], model_data.drop(['Churn?_False.', 'Churn?_True.'], axis=1)], axis=1)
# ここで学習用、バリデーション用、テスト用データにわけましょう。これによって overfitting (学習用データには精度が良いが、実際に利用すると制度が悪い、といった状況) を回避しやすくなり、未知のテストデータに対する精度を確認することができます。
train_data, validation_data, test_data = np.split(model_data.sample(frac=1, random_state=1729), [int(0.7 * len(model_data)), int(0.9 * len(model_data))])
train_data.to_csv('train.csv', header=False, index=False)
validation_data.to_csv('validation.csv', header=False, index=False)
# 学習には学習用データとバリデーション用データのみが必要です。上で csv に出力したデータをS3にアップロードして学習に利用できるようにします。
sagemaker_session = sagemaker.Session()
input_train = sagemaker_session.upload_data(path='train.csv', key_prefix='sagemaker/DEMO-xgboost-churn')
input_validation = sagemaker_session.upload_data(path='validation.csv', key_prefix='sagemaker/DEMO-xgboost-churn')
# `input_train` と `input_validation` にはアップロードしたファイルのS3パスが保存されています。これらは csv ファイルですが、Amazon SageMaker が用意している XGBoost のコンテナは、ファイルをデフォルトで libsvm 形式と認識してしまうため、このままだとエラーが発生します。
# `s3_input`という関数を利用して、`content_type='text/csv'`を明示的に指定することで、csv 形式と認識させることができます。
# +
from sagemaker.session import s3_input
s3_input_train = s3_input(s3_data=input_train, content_type='text/csv')
s3_input_validation = s3_input(s3_data=input_validation, content_type='text/csv')
# -
# ---
# ## 4.学習
#
# それでは学習を始めましょう。まず、XGBoost のコンテナの場所を取得します。コンテナ自体は SageMaker 側で用意されているので、場所を指定すれば利用可能です。
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, 'xgboost', '0.90-1')
# 学習のためにハイパーパラメータを指定したり、学習のインスタンスの数やタイプを指定することができます。XGBoost における主要なハイパーパラメータは以下のとおりです。
#
# - `max_depth` アルゴリズムが構築する木の深さをコントロールします。深い木はより学習データに適合しますが、計算も多く必要で、overfiting になる可能性があります。たくさんの浅い木を利用するか、少数の深い木を利用するか、モデルの性能という面ではトレードオフがあります。
# - `subsample` 学習データのサンプリングをコントロールします。これは overfitting のリスクを減らしますが、小さすぎるとモデルのデータが不足してしまいます。
# - `num_round` ブースティングを行う回数をコントロールします。以前のイテレーションで学習したときの残差を、以降のモデルにどこまで利用するかどうかを決定します。多くの回数を指定すると学習データに適合しますが、計算も多く必要で、overfiting になる可能性があります。
# - `eta` 各ブースティングの影響の大きさを表します。大きい値は保守的なブースティングを行います。
# - `gamma` ツリーの成長の度合いをコントロールします。大きい値はより保守的なモデルを生成します。
#
# XGBoostのhyperparameterに関する詳細は [github](https://github.com/dmlc/xgboost/blob/master/doc/parameter.rst) もチェックしてください。
# +
sess = sagemaker.Session()
xgb = sagemaker.estimator.Estimator(container,
role,
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
sagemaker_session=sess)
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
silent=0,
objective='binary:logistic',
num_round=100)
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
# -
# ---
# ## 5.ホスティング
#
# 学習が終われば、`deploy()`を実行することで、エンドポイントを作成してモデルをデプロイできます。
xgb_predictor = xgb.deploy(initial_instance_count = 1, instance_type = 'ml.m4.xlarge')
# ### 5-1.評価
#
# 現在、エンドポイントをホストしている状態で、これを利用して簡単に予測を行うことができます。予測は http の POST の request を送るだけです。
# ここではデータを `numpy` の `array` の形式で送って、予測を得られるようにしたいと思います。しかし、endpoint は `numpy` の `array` を受け取ることはできません。
#
# このために、`csv_serializer` を利用して、csv 形式に変換して送ることができます。
xgb_predictor.content_type = 'text/csv'
xgb_predictor.serializer = csv_serializer
xgb_predictor.deserializer = None
# 作成済みのテストデータを受け取ると、これをデフォルト500行ずつのデータにわけて、エンドポイントに送信する `predict` という関数を用意します。あとは `predict` を実行して予測結果を受け取ります。
# +
def predict(data, rows=500):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, xgb_predictor.predict(array).decode('utf-8')])
return np.fromstring(predictions[1:], sep=',')
dtest = test_data.values
predictions = []
predictions.append(predict(dtest[:, 1:]))
predictions = np.array(predictions).squeeze()
# -
# 機械学習の性能を比較評価する方法はいくつかありますが、単純に、予測値と実際の値を比較しましょう。今回は、顧客が離反する `1` と離反しない `0` を予測しますので、この混同行列を作成します。
pd.crosstab(index=test_data.iloc[:, 0], columns=np.round(predictions), rownames=['actual'], colnames=['predictions'])
# _注意点, アルゴリズムにはランダムな要素があるので結果は必ずしも一致しません._
#
# 48人の離反者がいて、それらの39名 (true positives) を正しく予測できました。そして、4名の顧客は離反すると予測しましたが、離反していません (false positives)。9名の顧客は離反しないと予測したにもかかわらず離反してしまいました (false negatives)。
#
# 重要な点として、離反するかどうかを `np.round()` という関数で、しきい値0.5で判断しています。`xgboost` が出力する値は0から1までの連続値で、それらを離反する `1` と 離反しない `0` に分類します。しかし、その連続値 (離反する確率) が示すよりも、顧客の離反というのは損害の大きい問題です。つまり離反する確率が低い顧客も、しきい値を0.5から下げて、離反するとみなす必要があるかもしれません。もちろんこては、false positives (離反すると予測したけど離反しなかった)を増やすと思いますが、 true positives (離反すると予測して離反した) を増やし、false negatives (離反しないと予測して離反した)を減らせます。
#
# 直感的な理解のため、予測結果の連続値をみてみましょう。
plt.hist(predictions)
plt.show()
# 連続値は0から1まで歪んでいますが、0.1から0.9までの間で、しきい値を調整するにはちょうど良さそうです。
pd.crosstab(index=test_data.iloc[:, 0], columns=np.where(predictions > 0.3, 1, 0))
# 例えば、しきい値を0.5から0.3まで減らしてみたとき、true positives は 1 つ、false positives は 3 つ増え、false negatives は 1 つ減りました。全体からみると小さな値ですが、全体の6-10%の顧客が、しきい値の変更で、予測結果が変わりました。ここで5名にインセンティブを与えることによって、インセンティブのコストが掛かりますが、3名の顧客を引き止めることができるかもしれません。
# つまり、最適な閾値を決めることは、実世界の問題を機械学習で解く上で重要なのです。これについてもう少し広く議論し、仮説的なソリューションを考えたいと思います。
#
# ### 5-2.推論エラーのコスト
#
# 2値分類の問題においては、しきい値に注意しなければならないという、似たような状況に直面することが多いです。それ自体は問題ではありません。もし、出力の連続値が2クラスで完全に別れていれば、MLを使うことなく単純なルールで解くことができると考えられます。
#
# 重要なこととして、MLモデルを正版環境に導入する際、モデルが false positives と false negatives に誤って入れたときのコストがあげられます。しきい値の選択は4つの指標に影響を与えます。4つの指標に対して、ビジネス上の相対的なコストを考える必要があるでしょう。
#
# #### コストの割当
#
# 携帯電話会社の離反の問題において、コストとはなんでしょうか?コストはビジネスでとるべきアクションに結びついています。いくつかの仮定をおいてみましょう。
#
# まず、true negatives のコストとして \$0 を割り当てます。満足しているお客様を正しく認識できていれば何も実施しません。
#
# false negatives が一番問題で、なぜなら、離反していく顧客を正しく予測できないからです。顧客を失えば、再獲得するまでに多くのコストを払う必要もあり、例えば逸失利益、広告コスト、管理コスト、販売管理コスト、電話の購入補助金などがあります。インターネットを簡単に検索してみると、そのようなコストは数百ドルとも言われ、ここでは \$500 としましょう。これが false negatives に対するコストです。
#
# 最後に、離反していくと予測された顧客に\$100のインセンティブを与えることを考えましょう。
# 携帯電話会社がそういったインセンティブを提供するなら、2回くらいは離反の前に考え直すかもしれません。これは true positive と false negative のコストになります。false positives の場合 (顧客は満足していて、モデルが誤って離反しそうと予測した場合)、\$100 のインセンティブは捨てることになります。その \$100 を効率よく消費してしまうかもしれませんが、優良顧客へのロイヤリティを増やすという意味では悪くないかもしれません。
#
# ### 5-3.最適な閾値を探す
#
# false negatives が false positives よりもコストが高いことは説明しました。そこで、顧客の数ではなく、コストを最小化するように、しきい値を最適化することを考えましょう。コストの関数は以下のようなものになります。
#
# ```txt
# $500 * FN(C) + $0 * TN(C) + $100 * FP(C) + $100 * TP(C)
# ```
#
# FN(C) は false negative の割合で、しきい値Cの関数です。同様にTN, FP, TP も用意します。この関数の値が最小となるようなしきい値Cを探します。
# 最も単純な方法は、候補となる閾値で何度もシミュレーションをすることです。以下では100個の値に対してループで計算を行います。
# +
cutoffs = np.arange(0.01, 1, 0.01)
costs = []
for c in cutoffs:
_predictions = pd.Categorical(np.where(predictions > c, 1, 0), categories=[0, 1])
matrix_a = np.array([[0, 100], [500, 100]])
matrix_b = pd.crosstab(index=test_data.iloc[:, 0], columns=_predictions, dropna=False)
costs.append(np.sum(np.sum(matrix_a * matrix_b)))
costs = np.array(costs)
plt.plot(cutoffs, costs)
plt.show()
print('Cost is minimized near a cutoff of:', cutoffs[np.argmin(costs)], 'for a cost of:', np.min(costs))
# -
# ## 6.エンドポイントの削除
#
# エンドポイントは起動したままだとコストがかかります。不要な場合は削除します。
sagemaker.Session().delete_endpoint(xgb_predictor.endpoint)
| xgboost_customer_churn/xgboost_customer_churn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Packaging
#
# Once we've made a working program, we'd like to be able to share it with others.
#
# A good cross-platform build tool is the most important thing: you can always
# have collaborators build from source.
#
# ### Distribution tools
# Distribution tools allow one to obtain a working copy of someone else's package.
#
# - Language-specific tools:
# - python: PyPI,
# - ruby: Ruby Gems,
# - perl: CPAN,
# - R: CRAN
#
# - Platform specific packagers e.g.:
# - `brew` for MacOS,
# - `apt`/`yum` for Linux or
# - [`choco`](https://chocolatey.org/) for Windows.
# ### Laying out a project
#
# When planning to package a project for distribution, defining a suitable
# project layout is essential.
#
#
#
# + language="bash"
# tree --charset ascii greetings -I "doc|build|Greetings.egg-info|dist|*.pyc"
# -
# We can start by making our directory structure. You can create many nested directories at once using the `-p` switch on `mkdir`.
# + language="bash"
# mkdir -p greetings/greetings/test/fixtures
# mkdir -p greetings/scripts
# -
# ### Using setuptools
#
# To make python code into a package, we have to write a `setup.py` file:
# ```python
# from setuptools import setup, find_packages
#
# setup(
# name="Greetings",
# version="0.1.0",
# packages=find_packages(exclude=['*test']),
# )
# ```
# We can now install this code with
# ```
# pip install .
# ```
#
# And the package will be then available to use everywhere on the system.
#
from greetings.greeter import greet
greet("Terry","Gilliam")
# ### Convert the script to a module
#
# Of course, there's more to do when taking code from a quick script and turning it into a proper module:
# We need to add docstrings to our functions, so people can know how to use them.
from IPython.display import Code
Code("greetings/greetings/greeter.py")
import greetings
help(greetings.greeter.greet)
# The documentation string explains how to use the function; don't worry about this for now, we'll consider
# this on [the next section](./04documentation.html) ([notebook version](./04documentation.ipynb)).
# ### Write an executable script
#
#
#
#
#
#
Code("greetings/greetings/command.py")
#
#
#
# ### Specify dependencies
# We use the setup.py file to specify the packages we depend on:
# ```python
# setup(
# name="Greetings",
# version="0.1.0",
# packages=find_packages(exclude=['*test']),
# install_requires=['numpy', 'pyyaml'] # NOTE: this is an example to ilustrate how to add dependencies.
# ) # Greetings doesn't have any external dependency.
# ```
# ### Specify entry point
# This allows us to create a command to execute part of our library. In this case when we execute `greet` on the terminal, we will be calling the `process` function under `greetings/command.py`.
#
Code("greetings/setup.py")
#
# And the scripts are now available as command line commands:
#
#
#
# + language="bash"
# greet --help
# + language="bash"
# greet <NAME>
# greet --polite <NAME>
# greet <NAME> --title Cartoonist
# -
# ### Installing from GitHub
#
# We could now submit "greeter" to PyPI for approval, so everyone could `pip install` it.
#
# However, when using git, we don't even need to do that: we can install directly from any git URL:
#
# ```
# pip install git+git://github.com/ucl-rits/greeter
# ```
# + language="bash"
# greet Lancelot the-Brave --title Sir
# -
#
#
# ### Write a readme file
# e.g.:
Code("greetings/README.md")
# ### Write a license file
# e.g.:
Code("greetings/LICENSE.md")
# ### Write a citation file
# e.g.:
Code("greetings/CITATION.md")
# You may well want to formalise this using the [codemeta.json](https://codemeta.github.io/) standard or the [citation file format](http://citation-file-format.github.io/) - these don't have wide adoption yet, but we recommend it.
# ### Define packages and executables
# + language="bash"
# touch greetings/greetings/test/__init__.py
# touch greetings/greetings/__init__.py
# -
# ### Write some unit tests
#
# Separating the script from the logical module made this possible:
#
#
#
#
#
#
Code("greetings/greetings/test/test_greeter.py")
#
#
#
# Add a fixtures file:
#
#
#
#
#
#
Code("greetings/greetings/test/fixtures/samples.yaml")
# + magic_args="--no-raise-error" language="bash"
# pytest
# -
# However, this hasn't told us that also the third test is wrong! A better aproach is to parametrize the test as follows:
# +
# %%writefile greetings/greetings/test/test_greeter.py
import yaml
import os
import pytest
from ..greeter import greet
def read_fixture():
with open(os.path.join(os.path.dirname(__file__),
'fixtures',
'samples.yaml')) as fixtures_file:
fixtures = yaml.safe_load(fixtures_file)
return fixtures
@pytest.mark.parametrize("fixture", read_fixture())
def test_greeter(fixture):
answer = fixture.pop('answer')
assert greet(**fixture) == answer
# -
# Now when we run `pytest`, we get a failure per element in our fixture and we know all that fails.
# + magic_args="--no-raise-error" language="bash"
# pytest
# -
# We can also make pytest to check whether the docstrings are correct by adding the `--doctest-modules` flag:
# + magic_args="--no-raise-error" language="bash"
# pytest --doctest-modules
# -
# ### Developer Install
#
# If you modify your source files, you would now find it appeared as if the program doesn't change.
#
# That's because pip install **copies** the files.
#
# If you want to install a package, but keep working on it, you can do:
# ```
# pip install --editable .
# ```
# ### Distributing compiled code
#
# If you're working in C++ or Fortran, there is no language specific repository.
# You'll need to write platform installers for as many platforms as you want to
# support.
#
# Typically:
#
# * `dpkg` for `apt-get` on Ubuntu and Debian
# * `rpm` for `yum`/`dnf` on Redhat and Fedora
# * `homebrew` on OSX (Possibly `macports` as well)
# * An executable `msi` installer for Windows.
#
# #### Homebrew
#
# Homebrew: A ruby DSL, you host off your own webpage
#
# See an [installer for the cppcourse example](http://github.com/jamespjh/homebrew-reactor)
#
# If you're on OSX, do:
#
# ```
# brew tap jamespjh/homebrew-reactor
# brew install reactor
# ```
| ch04packaging/03Packaging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import re
import pickle
from itertools import chain
from collections import namedtuple
from datetime import datetime
import pandas as pd
# -
Event = namedtuple('Event', 'id, card, abstract, authors, title, datetime, etype')
with open('nips2016-presenting.pk', 'rb') as fin:
events = pickle.load(fin)
events_frame = pd.DataFrame(list(events.values()), columns=Event._fields)
events_frame.head()
# ## Let's stalk authors!!
author_list = list(chain(*[map(lambda n: n.strip(), a.split('·'))
for a in events_frame['authors']]))
# Ooooo... so many papers with repeated authors...
# Or authors with repeated names...
len(author_list), len(set(author_list))
# ## Only 6 Machine Translation papers?!
len([i for i in events_frame['abstract'] if 'machine translation' in i.strip()])
# ## Date munging
def munge_time(s):
time, space = s.split(' @ ')
try: # Don't cross AM -- PM
day, month, date, start, _, end, ampm2 = time.split()
ampm1 = ampm2
except ValueError: # When crossing AM -- PM
day, month, date, start, ampm1, _, end, ampm2 = time.split()
date = re.findall(r'\d+',date)[0]
start_time = ' '.join([month, date, '2016', start+ampm1])
end_time = ' '.join([month, date, '2016', end+ampm2])
return day, start_time, end_time
s = 'Tue Dec 6th 03:00 -- 03:50 PM @ Area 1 + 2'
day, start, end = munge_time(s)
datetime.strptime(start, '%b %d %Y %I:%M%p')
print (datetime.strptime(start, '%b %d %Y %I:%M%p'))
for i, row in events_frame.iterrows():
##print (row.datetime)
day, start, end = munge_time(row.datetime)
events_frame.set_value(i,'day',day)
events_frame.set_value(i,'start_time',start)
events_frame.set_value(i,'end_time',end)
events_frame
| NIPS2016.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="3JAwvw7FFdMA" outputId="6aa5f633-f7ef-490a-d0fc-86a0390fa309" colab={"base_uri": "https://localhost:8080/", "height": 605}
# !pip install bilby
# + id="1sn3f821hc_2" outputId="b7f511b8-f73a-4173-8760-cf3974263842" colab={"base_uri": "https://localhost:8080/", "height": 655}
# !pip install lalsuite
# + id="lyMd_fF2hoT0" outputId="b329b157-8ce4-4548-c582-357c856e142d" colab={"base_uri": "https://localhost:8080/", "height": 588}
# !pip install gwpy
# + id="R8XvFnS95QLN" outputId="cfa8486e-e655-4c08-e7df-77039bdbf770" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#necessary modules are downloaded
"""
A script to sample a lensed signal by assuming that there is no lensing present
"""
from __future__ import division, print_function
import bilby
import numpy as np
import scipy
from scipy.special import hyp1f1
import mpmath as mp
import matplotlib.pyplot as plt
import lal
import lalsimulation
# First set up logging and some output directories and labels
outdir = 'outdir'
label = 'create_your_own_bbh_model'
fs = 128 #sampling_frequency
T_obs = 1 #duration
#lens model function - point mass model
def F(w,y):
if(y>0):
xm=0.5*(y+np.sqrt(y*y+4.0))
phim=0.5*((xm-y)**2)-np.log(xm)
HYP = [complex(mp.hyp1f1(((1j/2)*z),1.0,((1j/2)*z*y*y))) for z in w]
F = ((np.exp((np.pi*w)/4)) * (scipy.special.gamma(1-((1j/2)*w))) * HYP * (np.exp(0.5j*w*(np.log(0.5*w)-2.0*phim))))
else:
F=[1.0 for z in w]
return F
# Here we define out source model - this is the BBH merger signal in the frequency domain.
def gen_bbh(f, mass_1, mass_2, iota, phi, ra, dec, psi, d, geocent_time,y,M):
"""
generates a BBH frequency domain signal
"""
Lens_mass = M * scipy.constants.G * lal.MSUN_SI / scipy.constants.c**3 #lens mass - scaled to time
w = 8*np.pi*Lens_mass*f
N = T_obs * fs # the total number of time samples
dt = 1.0 / fs # the sampling time (sec)
df = 1.0/T_obs # the sampling frequency
f_low = 12.0 # lowest frequency of waveform (Hz)
f_max = 64.0
approximant = lalsimulation.IMRPhenomD
dist = d*1e6*lal.PC_SI # put it as 1 MPc
Mag=F(w,y)
Mag[0]=1.0
if(mass_1<mass_2):
print(mass_1,mass_2)
# make waveform
hp, hc = lalsimulation.SimInspiralChooseFDWaveform(mass_1 * lal.MSUN_SI, mass_2 * lal.MSUN_SI, 0, 0, 0, 0, 0, 0,
dist, iota, phi, 0, 0, 0,
df, f_low, f_max, f_low , lal.CreateDict(), approximant)
return {'plus': Mag*hp.data.data, 'cross': Mag*hc.data.data} #adding lens model while returning plus and cross polarisation
#injection parameters
injection_parameters = dict(mass_1=36.0,mass_2=29.0,iota=150*np.pi/180,phi=0,ra=0, dec=0, psi=0,d=500, geocent_time=0,y=0.1,M=4000)
# Now we pass our source function to the WaveformGenerator
waveform_generator = bilby.gw.waveform_generator.WaveformGenerator(
duration=T_obs, sampling_frequency=fs,
frequency_domain_source_model=gen_bbh)
# Set up interferometers.
ifos = bilby.gw.detector.InterferometerList(['H1'])
ifos.set_strain_data_from_power_spectral_densities(
sampling_frequency=fs, duration=T_obs,
start_time=injection_parameters['geocent_time'] - 3)
ifos.inject_signal(waveform_generator=waveform_generator,
parameters=injection_parameters)
# Here we define the priors for the search. We use the injection parameters
# except for the amplitude, f0, and geocent_time
from bilby.core.prior import PriorDict, Uniform, Constraint
prior = bilby.gw.prior.BBHPriorDict()
for key in ['iota', 'phi', 'psi', 'ra', 'dec', 'geocent_time','M']:
prior[key] = injection_parameters[key]
prior['y']=-1 #to recover as if there is no lensing effect
prior['theta_jn']=0
prior['phase']=0
prior['luminosity_distance']=0
prior['a_1']=0
prior['a_2']=0
prior['tilt_1']=0
prior['tilt_2']=0
prior['phi_12']=0
prior['phi_jl']=0
prior['chirp_mass'] = bilby.prior.Constraint(
name='chirp_mass', latex_label='$M$', minimum=20.0, maximum=40.0,
unit='$M_{\\odot}$')
prior['mass_ratio'] = bilby.prior.Constraint(
name='mass_ratio', latex_label='$q$', minimum=0.5, maximum=1.0)
prior['mass_1'] = Uniform(name='mass_1', minimum=0, maximum=50)
prior['mass_2'] = Uniform(name='mass_2', minimum=0, maximum=50)
prior['d'] = bilby.core.prior.PowerLaw(alpha=2, name='luminosity_distance', minimum=20, maximum=1000, unit='Mpc', latex_label='$d_L$')
likelihood = bilby.gw.likelihood.GravitationalWaveTransient(
interferometers=ifos, waveform_generator=waveform_generator)
#plot corner plots
result = bilby.core.sampler.run_sampler(
likelihood, prior, sampler='dynesty', outdir=outdir, label=label,
resume=False, sample='unif', injection_parameters=injection_parameters)
result.plot_corner()
| Sampling_without_considering_the_lens_effect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # SQL - Interfejs Pythona dla baz danych SQLite
#
# _<NAME>_
#
# 
#
# * Interfejs sqlite3
# * Ćwiczenia
# * Test
# * Materiały dodatkowe
# ## Interfejs sqlite3
#
# Interfejs sqlite3 to biblioteka C, która zapewnia lekką dyskową bazę danych, która nie wymaga oddzielnego procesu serwera i umożliwia dostęp do bazy danych przy użyciu niestandardowego wariantu języka zapytań SQL. Niektóre aplikacje mogą używać sqlite3 do wewnętrznego przechowywania danych. Możliwe jest również prototypowanie aplikacji przy użyciu sqlite3, a następnie przeniesienie kodu do większej bazy danych, takiej jak PostgreSQL lub Oracle.
#
# Moduł sqlite3 został napisany przez <NAME>. Zapewnia interfejs SQL zgodny ze specyfikacją DB-API 2.0 opisaną w [PEP 249](https://www.python.org/dev/peps/pep-0249).
#
# Aby użyć modułu, musisz najpierw utworzyć obiekt [`Connection`](https://docs.python.org/3/library/sqlite3.html#sqlite3.Connection), który reprezentuje bazę danych. Tutaj dane zostaną zapisane w pliku `example.db`:
import os
os.remove('example.db')
import sqlite3
conn = sqlite3.connect('example.db')
# Możesz również podać specjalną nazwę `:memory:` aby utworzyć bazę danych w pamięci RAM.
#
# Gdy masz już połączenie ([`Connection`](https://docs.python.org/3/library/sqlite3.html#sqlite3.Connection)), możesz utworzyć obiekt [Cursor](https://docs.python.org/3/library/sqlite3.html#sqlite3.Cursor) i wywołać jego metodę [execute()](https://docs.python.org/3/library/sqlite3.html#sqlite3.Cursor.execute) w celu wykonania poleceń SQL:
c = conn.cursor()
# Utwórz tabelę:
c.execute('''CREATE TABLE stocks
(date text, trans text, symbol text, qty real, price real)''')
# Wstaw wiersz danych:
c.execute("INSERT INTO stocks VALUES ('2006-01-05','BUY','RHAT',100,35.14)")
# Zapisz (zatwierdź, ang. _commit_) zmiany:
conn.commit()
# Możemy również zamknąć połączenie, jeśli skończymy.
#
# Tylko upewnij się, że wszelkie zmiany zostały wprowadzone, w przeciwnym razie zostaną utracone:
conn.close()
# Zapisane dane są trwałe i są dostępne w kolejnych sesjach:
import sqlite3
conn = sqlite3.connect('example.db')
c = conn.cursor()
# Nigdy tego nie rób (niepewne!):
symbol = 'RHAT'
c.execute("SELECT * FROM stocks WHERE symbol = '%s'" % symbol)
# Zrób to zamiast tego:
t = ('RHAT',)
c.execute('SELECT * FROM stocks WHERE symbol=?', t)
print(c.fetchone())
# Argument `execute`, który reprezentuje wartości, które chcesz wstawić do bazy danych, powinien być krotką (sekwencją)!
#
# Większy przykład, który wstawia wiele rekordów naraz:
purchases = [('2006-03-28', 'BUY', 'IBM', 1000, 45.00),
('2006-04-05', 'BUY', 'MSFT', 1000, 72.00),
('2006-04-06', 'SELL', 'IBM', 500, 53.00),
]
c.executemany('INSERT INTO stocks VALUES (?,?,?,?,?)', purchases)
# Aby pobrać dane po wykonaniu instrukcji SELECT, możesz traktować kursor jako [iterator](https://docs.python.org/3/glossary.html#term-iterator), wywołać metodę [`fetchone()`](https://docs.python.org/3/library/sqlite3.html#sqlite3.Cursor.fetchone) kursora w celu pobrania pojedynczego pasującego wiersza lub wywołać funkcję [`fetchall()`](https://docs.python.org/3/library/sqlite3.html#sqlite3.Cursor.fetchall) w celu uzyskania listy pasujących wierszy.
#
# W tym przykładzie zastosowano formę iteratora:
for row in c.execute('SELECT * FROM stocks ORDER BY price'):
print(row)
# Metoda **`iterdump()`** zwraca iterator do zrzutu bazy danych w formacie tekstowym SQL. Przydatne podczas zapisywania bazy danych w pamięci w celu późniejszego przywrócenia.
#
# Można więc napisać program w Pythonie, aby utworzyć kopię zapasową bazy danych SQLite:
import sqlite3
import io
conn = sqlite3.connect('CodeBrainers.db')
with io.open('CodeBrainers_dump.sql', 'w') as f:
for line in conn.iterdump():
f.write('%s\n' % line)
print('Kopia zapasowa została wykonana pomyślnie.')
print('Zapisano jako CodeBrainers_dump.sql')
conn.close()
# ## Ćwiczenia
# Python i baza danych SQLite - ćwiczenia i praktyka
# ### Utwórz połączenie bazy danych SQLite z bazą danych znajdującą się w pamięci
# #### Ćwiczenie
# Napisz program w języku Python, aby utworzyć połączenie bazy danych SQLite z bazą danych znajdującą się w pamięci.
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
conn = sqlite3.connect(':memory:')
print("Baza danych pamięci utworzona i połączona z SQLite.")
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Utwórz bazę danych SQLite i połącz się z bazą danych oraz wydrukuj wersje
# #### Ćwiczenie
# Napisz program w Pythonie, aby utworzyć bazę danych SQLite w pliku i połączyć się z bazą danych oraz wydrukować wersję bazy danych SQLite i numer wersji modułu sqlite3 w postaci ciągu.
#
# Podpowiedzi:
# * Atrybut SQLite **sqlite_version** zwraca ciąg znaków wersji dla uruchomionej biblioteki SQLite.
# * Funkcja Pythona `sqlite3.`**`version`** zwraca numer wersji modułu sqlite3 w postaci ciągu. To nie jest wersja biblioteki SQLite!
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
conn = sqlite3.connect('temp.db')
c = conn.cursor()
print("Baza danych stworzona i połączona z SQLite.")
query = "SELECT sqlite_version();"
c.execute(query)
record = c.fetchall()
print("Wersja bazy danych SQLite to: ", record)
print("Wersja modułu sqlite3 to: ", sqlite3.version)
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Utwórz bazę danych SQLite, połącz się z bazą danych oraz zabezpiecz się przed wyjątkami
# #### Ćwiczenie
# Napisz program w Pythonie, aby utworzyć bazę danych SQLite i połączyć się z bazą danych oraz zabezpieczyć się przed wyjątkami. Wywołaj sztucznie wyjątek, wykonując błędne zapytanie.
#
# Podpowiedź: _wyjątek_ `sqlite3.`**`Error`** jest to klasa bazowa pozostałych wyjątków w module sqlite3. Jest to zarazem podklasa [`Exception`](https://docs.python.org/3/library/exceptions.html#Exception).
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
try:
conn = sqlite3.connect(':memory:')
c = conn.cursor()
print("Baza danych stworzona i połączona z SQLite.")
c.execute("SELECT * FROM users;")
except sqlite3.Error as error:
print("Błąd podczas łączenia się z SQLite", error)
finally:
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Utwórz tabelę w bazie danych SQLite
# #### Ćwiczenie
# Napisz program w Pythonie, aby połączyć bazę danych SQLite, utworzyć tabelę w bazie danych i zweryfikować jej utworzenie.
#
# Podpowiedź: Każda baza danych SQLite zawiera pojedynczą „tabelę schematów”, która przechowuje schemat tej bazy danych. Schemat bazy danych to opis wszystkich innych tabel, indeksów, wyzwalaczy i widoków zawartych w bazie danych. Tabela schematów wygląda następująco:
#
# ```sqlite
# CREATE TABLE sqlite_schema(
# type text,
# name text,
# tbl_name text,
# rootpage integer,
# sql text
# );
# ```
#
# Tabela `sqlite_schema` zawiera po jednym wierszu dla każdej tabeli, indeksu, widoku i wyzwalacza (zbiorczo „obiekty”) w schemacie, z wyjątkiem tego, że nie ma wpisu dla samej tabeli `sqlite_schema`. Zapoznaj się z podsekcją dotyczącą [przechowywania schematów](https://www.sqlite.org/fileformat2.html#ffschema) w dokumentacji [formatu plików](https://www.sqlite.org/fileformat2.html), aby uzyskać dodatkowe informacje o tym, jak program SQLite używa wewnętrznie tabeli `sqlite_schema`.
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
#
# # **UWAGA PONIŻSZY KOD NIE DZIAŁA POD PYCHARM!!!!**
import sqlite3
conn = sqlite3.connect(':memory:')
c = conn.cursor()
c.execute("CREATE TABLE users(login VARCHAR(8) NOT NULL, name VARCHAR(40) NOT NULL, phone_no VARCHAR(15));")
print("Utworzono tabelę Users.")
c.execute("SELECT * FROM sqlite_schema;")
record = c.fetchall()
print(record)
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Wypisz zawartość tabel z pliku bazy danych SQLite CodeBrainers
# #### Ćwiczenie
# Napisz program w Pythonie, który wyświetli listy zawartości tabel pliku bazy danych SQLite CodeBrainers.
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
conn = sqlite3.connect('CodeBrainers.db')
c = conn.cursor()
print("Wykaz zawartości tabel:")
c.execute("SELECT * FROM product")
print(c.fetchall())
c.execute("SELECT * FROM customer")
print(c.fetchall())
c.execute("SELECT * FROM order_product")
print(c.fetchall())
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Utwórz i wypełnij tabelę w bazie danych SQLite
# #### Ćwiczenie
# Napisz program w Pythonie, aby połączyć bazę danych SQLite i utworzyć tabelę w bazie danych:
#
# Struktura tabeli `Users`:
#
# ```sqlite
# login VARCHAR(8) NOT NULL
# name VARCHAR(40) NOT NULL
# phone_no VARCHAR(15)
# ```
#
# Następnie spróbuj dodać jeden wiersz (np.: `'user', '<NAME>', '1234567890'`) i go wyświetlić z bazy danych.
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
conn = sqlite3.connect(':memory:')
c = conn.cursor()
c.execute("CREATE TABLE users(login VARCHAR(8) NOT NULL, name VARCHAR(40) NOT NULL, phone_no VARCHAR(15));")
print("Utworzono tabelę Users.")
c.execute("INSERT INTO users VALUES ('user', '<NAME>', '1234567890');")
c.execute("SELECT * FROM users;")
record = c.fetchall()
print(record)
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Wstaw listę rekordów do podanej tabeli SQLite i policz liczbę wierszy w danej tabeli SQLite
# #### Ćwiczenie
# Napisz program w Pythonie, który wstawi listę kilku rekordów do podanej tabeli SQLite o kilku kolumnach (jednym poleceniem). Policz liczbę wierszy w danej tabeli SQLite (przed i po wstawieniu wierszy).
#
# Przykładowe kolumny tabeli:
#
# ```sqlite
# id SMALLINT
# name VARCHAR(30)
# city VARCHAR(35)
# ```
#
# Przykładowe rekordy:
#
# ```sqlite
# 5001, '<NAME>', 'Warszawa'
# 5002, '<NAME>', 'Kraków'
# 5003, '<NAME>', 'Łódź'
# 5004, '<NAME>', 'Kraków'
# 5005, '<NAME>', 'Wrocław'
# ```
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
conn = sqlite3.connect(':memory:')
c = conn.cursor()
# Utwórz tabelę
c.execute("CREATE TABLE users(id SMALLINT, name VARCHAR(30), city VARCHAR(35));")
print("Liczba rekordów przed wstawieniem wierszy:")
cursor = c.execute('SELECT * FROM users;')
print(len(cursor.fetchall()))
query = "INSERT INTO users (id, name, city) VALUES (?, ?, ?);"
# Wstaw rekordy
rows = [(5001,'<NAME>', 'Warszawa'),
(5002,'<NAME>', 'Kraków' ),
(5003,'<NAME>', 'Łódź' ),
(5004,'<NAME>', 'Kraków' ),
(5005,'<NAME>', 'Wrocław' )]
c.executemany(query, rows)
conn.commit()
print("Liczba rekordów po wstawieniu wierszy:")
cursor = c.execute('SELECT * FROM users;')
print(len(cursor.fetchall()))
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Wstaw wartości do tabeli z danych wejściowych użytkownika
# #### Ćwiczenie
# Napisz program w Pythonie, który będzie wstawiał wartości do tabeli z danych wejściowych użytkownika.
#
# Przykładowe kolumny tabeli:
#
# ```sqlite
# id SMALLINT
# name VARCHAR(30)
# city VARCHAR(35)
# ```
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
conn = sqlite3.connect(':memory:')
c = conn.cursor()
# Utwórz tabelę użytkowników
c.execute("CREATE TABLE users(id SMALLINT, name VARCHAR(30), city VARCHAR(35));")
input_id = input('ID:')
input_name = input('Name:')
input_city = input('City:')
c.execute("INSERT INTO users(id, name, city) VALUES (?,?,?)", (input_id, input_name, input_city))
conn.commit()
print('Dane wprowadzone pomyślnie.')
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Zaktualizuj określoną wartość kolumny w danej tabeli
# #### Ćwiczenie
# Napisz program w Pythonie, aby zaktualizować określoną wartość kolumny w danej tabeli i wybrać/wyświetlić w pętli wszystkie wiersze przed i po aktualizacji tej tabeli.
#
# Przykładowe kolumny tabeli:
#
# ```sqlite
# id SMALLINT
# name VARCHAR(30)
# city VARCHAR(35)
# ```
#
# Przykładowe rekordy:
#
# ```sqlite
# 5001, '<NAME>', 'Warszawa'
# 5002, '<NAME>', 'Kraków'
# 5003, '<NAME>', 'Łódź'
# 5004, '<NAME>', 'Kraków'
# 5005, '<NAME>', 'Wrocław'
# ```
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
conn = sqlite3.connect(':memory:')
c = conn.cursor()
# Utwórz tabelę
c.execute("CREATE TABLE users(id SMALLINT, name VARCHAR(30), city VARCHAR(35));")
query = "INSERT INTO users (id, name, city) VALUES (?, ?, ?);"
# Wstaw rekordy
rows = [(5001,'<NAME>', 'Warszawa'),
(5002,'<NAME>', 'Kraków' ),
(5003,'<NAME>', 'Łódź' ),
(5004,'<NAME>', 'Kraków' ),
(5005,'<NAME>', 'Wrocław' )]
c.executemany(query, rows)
conn.commit()
rows = c.execute('SELECT * FROM users;')
rows = c.fetchall()
print("Dane użytkowników:")
for row in rows:
print(row)
print("Zaktualizuj miasto Łódź do Poznania, gdzie id to 5003:")
c.execute('UPDATE users SET city = "Poznań" WHERE id = 5003;')
print("Rekord zaktualizowany pomyślnie.")
rows = c.execute('SELECT * FROM users;')
rows = c.fetchall()
print("Po zaktualizowaniu danych użytkowników:")
for row in rows:
print(row)
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ### Usuń określony wiersz z podanej tabeli SQLite
# #### Ćwiczenie
# Napisz program w Pythonie, aby usunąć określony (danymi wejściowymi uzytkownika) wiersz z podanej tabeli SQLite i wybrać/wyświetlić w pętli wszystkie wiersze przed i po aktualizacji tej tabeli.
#
# Przykładowe kolumny tabeli:
#
# ```sqlite
# id SMALLINT
# name VARCHAR(30)
# city VARCHAR(35)
# ```
#
# Przykładowe rekordy:
#
# ```sqlite
# 5001, '<NAME>', 'Warszawa'
# 5002, '<NAME>', 'Kraków'
# 5003, '<NAME>', 'Łódź'
# 5004, '<NAME>', 'Kraków'
# 5005, '<NAME>', 'Wrocław'
# ```
# ---
# #### Rozwiązanie
# Kod Pythona (przykładowe rozwiązanie i przykładowe dane wyjściowe):
import sqlite3
conn = sqlite3.connect(':memory:')
c = conn.cursor()
# Utwórz tabelę
c.execute("CREATE TABLE users(id SMALLINT, name VARCHAR(30), city VARCHAR(35));")
query = "INSERT INTO users (id, name, city) VALUES (?, ?, ?);"
# Wstaw rekordy
rows = [(5001,'<NAME>', 'Warszawa'),
(5002,'<NAME>', 'Kraków' ),
(5003,'<NAME>', 'Łódź' ),
(5004,'<NAME>', 'Kraków' ),
(5005,'<NAME>', 'Wrocław' )]
c.executemany(query, rows)
conn.commit()
rows = c.execute('SELECT * FROM users;')
rows = c.fetchall()
print("Dane użytkowników:")
for row in rows:
print(row)
input_id = input('ID:')
print("Usuń użytkownika o ID", input_id, ":")
c.execute('DELETE FROM users WHERE id = ?;', (input_id,))
conn.commit()
print("Rekord zaktualizowany pomyślnie.")
rows = c.execute('SELECT * FROM users;')
rows = c.fetchall()
print("Po zaktualizowaniu danych użytkowników:")
for row in rows:
print(row)
conn.close()
print("Połączenie SQLite jest zamknięte.")
# ## [Test](https://www.w3schools.com/sql/sql_quiz.asp)
#
# Możesz sprawdzić swoje umiejętności SQL w Quizie.
#
# Test zawiera 25 pytań i nie ma ograniczenia czasowego.
#
# Test nie jest oficjalny, jest po prostu przyjemnym sposobem sprawdzenia, ile wiesz lub nie wiesz o SQL.
# ## Materiały dodatkowe
#
# * [SQL e-learning](http://zasoby.open.agh.edu.pl/~11smdrobniak/)
# * [Podstawy baz danych z encyklopedią SQL](http://zasoby.open.agh.edu.pl/~09seenglert/)
# * [MySQL - podstawy](http://www.galaxy.agh.edu.pl/~pamalino/programowanie/mysql/)
# * [SQL](https://github.com/pkociepka/sql)
| SQL/06 SQL - Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python395jvsc74a57bd07dea30b6c03ab7e134c0e74006a546b873dc5838ff7024ff5edf21dbf1fb34a3
# ---
# +
# Object-Oriented Programming Fundamentals
# +
# Using the simplest method
profile1_name = 'Mateus'
profile1_number = '000-000'
profile1_company = 'Undefined'
profile2_name = 'Naruchan'
profile2_number = '111-111'
profile2_company = 'Undefined'
# -
# Using dictonary {'key': 'value'}
profile = {'name': 'Raquel', 'number': '222-222'}
print(profile['name'])
print(profile['number'])
from models import Profile
| 4. Object_Oriented_Programming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# ##### array
# - ndim : 행렬의 차원
# - shape: 행렬의 모양
# 1차원 배열 (vector)
array = np.array([0,1,2,3,4,5])
print(type(array))
print(array)
# 2차원 배열
array = np.array([[0],[1],[2]])
print(array)
array = np.array([
[[0,1,2,3],
[4,5,6,7],
[8,9,10,11]],
[[0,1,2,3],
[4,5,6,7],
[8,9,10,11]]
])
print(array)
print(array.ndim, array.shape) # 바깥에서 안으로의 모양
# #### reshape
# - 행렬의 모양 바꿀 수 있다.
na = np.array([1,2,3,4,5,6,7,8])
na
na.reshape((2,4)) # 2 X 4 matrix change
rna = np.reshape(na, (4,2)) # 4 X 2 matrix change
rna
# 전치행렬
rna.T # 2 X 4 matrix change
# #### index
# - 행렬의 특정 위치 데이터 값 가져오기
# - 행렬 자르기
array
print(array[1][1][2])
print(array[1,1,2])
result = array[1:,1:,2:]
print(result)
print(result.ndim, result.shape)
result[0].ndim
# 데이터 수정
data = np.zeros(5, dtype=int)
data
data[1] = 1
data
data[3:] = 2
data
z = np.zeros((3,4))
z
z[1:,2:] = 2
z
# #### zeros
# - 행렬을 만들고 안에 0으로 채워 넣는다.
z = np.zeros((2,3,4))
z
# #### ones
# - 행렬을 만들고 안에 1을 채워 넣는다.
o = np.ones([2,3,4])
print(o)
print(o.ndim, o.shape)
# #### eye
# - 단위행렬 데이터를 만든다
np.eye(5)
# #### like
# - 행렬 안에 있는 모든 데이터를 0이나 1로 바꿀 수 있다.
z
o
ls_1 = np.zeros_like(o)
print(ls_1)
print(ls_1.ndim, ls_1.shape)
ls_2 = np.ones_like(z)
print(ls_2)
print((ls_2.ndim, ls_2.shape))
# #### empty
# - 빈 행렬을 만든다
# - 실제로는 빈데이터가 아니라 더미 데이터로 들어간다.
a = np.empty((5,5))
a
b = np.empty((6,4))
b
# #### arange
# - range와 같다.
# - 하지만 큰 데이터에서 range보다 속도가 빠르다.
np.arange(10)
np.arange(5,10)
np.arange(2,10,3)
np.arange(10,1,-2)
# #### linspace & logspace
np.linspace(0,100,3)
# 2~4까지 3개로 나누기
# logx = 2, 3, 4이 되도록 하는 x값을 출력하라
print(np.logspace(2,4,3))
print(np.logspace(1,4,4))
# ##### random
# - seed
# - rand (uniform)
# - randn (gaussian)
# - randint
# - shuffle
# - choice
np.random.seed(0)
rd = np.random.rand(10)
rdn = np.random.randn(10)
print(rd)
print(rdn)
r = np.random.rand(2,3,4)
r
# 지정된 범위의 수로 행렬 만들기
r = np.random.randint(5, 10, (3,3))
r
# 시작과 끝을 모두 안적어주는 경우 : range와 비슷
r = np.random.randint(10,size=(3,3)) # 10보다 작은 수로 행렬 만들어라
r
# shuffle로 데이터 순서 바꾸기
np.random.shuffle(r)
r
# p의 확률로 range(5)를 10개 추출
print(list(range(5)))
print([0.1,0,0.3,0.6,0])
np.random.choice(5,10, p=[0.1,0,0.3,0.6,0]) # 0~4중 10개를 각 확률을 바탕으로 추출
# unique로 데이터의 유일한 값에 대한 리스트와 값의 갯수를 구할 수 있다.
index, count = np.unique(r, return_counts=True)
print(index) # unique한 값(중복제거)
print(count) # unique한 값들의 갯수
# #### stack
# - 행렬을 쌓는 것과 같이 합치는 방법
na1 = np.arange(6)
na2 = np.arange(6,12)
na1 = np.random.randint(10, size=(2,3))
na2 = np.random.randint(10, size=(2,3))
print(na1, na1.shape)
print(na2, na2.shape)
na3 = np.stack((na1,na2))
print(na3, na3.shape)
na4 = np.stack((na1,na2), axis=1)
print(na4, na4.shape)
na5 = np.stack((na1,na2), axis=2)
print(na5, na5.shape)
ls = []
for i in range(5):
print(range(7), i, i+3)
print(range(7)[i:i+3])
print(list(range(7)[i:i+3]), end="\n\n")
ls.append(range(7)[i:i+3])
# ls
np.vstack(ls) # 위로 쌓아줌
np.hstack(ls) # 옆으로 쌓아줌(하나의 리스트형태)
data = [list(range(7))[i:i+3] for i in range(5)]
print(data)
np.stack(data, axis=1) # axis가 커질수록 차원이 안쪽으로 간다.
# #### concatenate 결합
# - 가로결합 : 행의 갯수가 같아야함.
# - 세로결합 : 열의 갯수가 같아야함.
r1 = np.random.randint(10, size=(2,3)) #10이하의 숫자로 2X3행렬 만들기
r1
r2 = np.random.randint(10, size=(3,2)) #10이하의 숫자로 3X2행렬 만들기
r2
r3 = np.random.randint(10, size=(3,3)) #10이하의 숫자로 3X3행렬 만들기
r3
np.concatenate([r1,r3])
np.concatenate([r2,r3], axis=1)
# #### split
r = np.arange(10)
r # 0앞에서부터 0으로 셈
r1 = np.split(r, [5]) # 0앞에서부터 0으로 세서 5번째에서 자름.
r1
r2 = np.split(r, [2,4,6,8])
r2
# +
# vsplit
# -
r = np.random.randint(10, size=(4,6))
r
np.vsplit(r, [2,3])
# +
# hsplit
# -
np.hsplit(r, [2])
| python/02_numpy_&_pandas/10_Numpy_make_review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy.stats import poisson, chi2
import pandas as pd
from collections import defaultdict
import matplotlib.pyplot as plt
import seaborn as sns
% matplotlib inline
from sklearn.ensemble import GradientBoostingRegressor
from skgarden import RandomForestQuantileRegressor
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from xgboost import XGBClassifier
from tqdm.auto import tqdm
from sklearn.model_selection import GridSearchCV
import os
import statsmodels.api as sm
import sys, traceback
class Suppressor(object):
def __enter__(self):
self.stdout = sys.stdout
sys.stdout = self
def __exit__(self, type, value, traceback):
sys.stdout = self.stdout
if type is not None:
pass
def write(self, x): pass
# -
# ## Tables 2 (section 4) and Table 3 (Supp. Mat)
# +
directory = 'sims/table2/'
files = [x for x in os.listdir(directory) if 'csv' in x and 'truth' not in x]
for line in files:
print(line)
final_df = None
for flnm in files:
temp_df = pd.read_csv(directory + flnm, index_col=0)
temp_df['classifier'] = temp_df['classifier'].apply(lambda x: x.replace('\n ', ''))
temp_df['classifier_cde'] = temp_df['classifier_cde'].apply(lambda x: x.replace('\n ', ''))
if 'in_true_interval' in temp_df.columns:
temp_df['in_true_interval'] = temp_df['in_true_interval'].values
if final_df is None:
final_df = temp_df.copy()
else:
final_df = final_df.append(temp_df.copy())
# -
print(final_df['b_prime'].unique())
print(final_df['b'].unique())
print(final_df['classifier'].unique())
print(final_df['classifier_cde'].unique())
print(final_df['run'].unique())
print(final_df['sample_size_obs'].unique())
print(final_df['rep'].unique())
# +
def print_table_to_latex(df, hue='classifier'):
final_row = '\%s \hline' % ('\\')
hue_vals = df[hue].unique()
b_vals = df['b'].unique()
out = []
for b_val in b_vals:
for jj, hue_val in enumerate(hue_vals):
temp_df = df[(df[hue] == hue_val) & (df['b'] == b_val)]
temp_line = '%s & %s & %.2f $\pm$ %.2f & %.2f & %.1f $\pm$ %.1f' % (
'\multirow{3}{*}{%s}' % ('{:,}'.format(b_val)) if jj == 0 else '',
hue_val,
temp_df['cross_entropy_loss average'].values[0],
temp_df['cross_entropy_loss std'].values[0],
temp_df['out_confint notrue'].values[0],
temp_df['size_CI average'].values[0],
temp_df['size_CI std'].values[0]
)
if jj == 2:
out.append(temp_line + final_row)
else:
out.append(temp_line + '\%s' % ('\\'))
for line in out:
print(line)
def print_coverage_table_to_latex(df, hue='classifier'):
final_row = '\%s \hline' % ('\\')
hue_vals = df[hue].unique()
b_vals = df['b'].unique()
out = []
for b_val in b_vals:
for jj, hue_val in enumerate(hue_vals):
temp_df = df[(df[hue] == hue_val) & (df['b'] == b_val)]
temp_line = '%s & %s & %.2f' % (
'\multirow{3}{*}{%s}' % ('{:,}'.format(b_val)) if jj == 0 else '',
hue_val,
temp_df['in_confint average'].values[0])
if jj == 2:
out.append(temp_line + final_row)
else:
out.append(temp_line + '\%s' % ('\\'))
for line in out:
print(line)
# +
sample_size_val = 10
color_palette = sns.color_palette("cubehelix", 3)
for run in ['poisson', 'gmm']:
b_prime_val = 2500 if run == 'poisson' else 5000
class_cde = 'lgb' if run=='poisson' else 'xgb_d3_n100'
plot_df = final_df[(final_df['run'] == run) &
(final_df['classifier_cde'] == class_cde) &
(final_df['sample_size_obs'] == sample_size_val) &
(final_df['b_prime'] == b_prime_val)]
true_t0 = plot_df[plot_df['on_true_t0'] == 1]['theta_0_current'].values[0]
size_CI_df = plot_df[['b', 'classifier', 'cross_entropy_loss', 'size_CI', 'out_confint', 'on_true_t0']]
out_confint_outint = size_CI_df[size_CI_df['on_true_t0']==0].groupby(['b', 'classifier']).agg({
'out_confint': [np.average]}).round(2)['out_confint']
size_CI_df['size_CI'] = size_CI_df['size_CI'].values * 100
size_CI_df = size_CI_df.groupby(['b', 'classifier']).agg({'size_CI': [np.average, np.std],
'cross_entropy_loss': [np.average, np.std],
'out_confint': [np.average]}).round(2).reset_index()
size_CI_df.columns = [' '.join(col).strip() for col in size_CI_df.columns.values]
size_CI_df['out_confint notrue'] = out_confint_outint.values
print(size_CI_df)
print_table_to_latex(size_CI_df)
print('\n')
coverage_df = plot_df[plot_df['on_true_t0'] == 1.0][
['b', 'classifier', 'cross_entropy_loss', 'in_confint']]
coverage_df = coverage_df.groupby(['b', 'classifier']).agg({'in_confint': [np.average, np.std],
'cross_entropy_loss': [np.average, np.std]}).round(2).reset_index()
coverage_df.columns = [' '.join(col).strip() for col in coverage_df.columns.values]
print_coverage_table_to_latex(coverage_df)
truth_flnm = [x for x in os.listdir(directory) if 'truth' in x and run in x][0]
truth_df = pd.read_csv(directory + truth_flnm).set_index('Unnamed: 0')
power_vec = 1.0 - truth_df[truth_df['on_true_t0']==0].groupby(['classifier']).agg(
{'in_true_interval': np.average}).reset_index()['in_true_interval'].values
summary_truth_df = truth_df.groupby(['classifier']).agg({'size_true_int': [np.average, np.std],
'true_entropy': [np.average, np.std]})
summary_truth_df['power'] = power_vec
print(summary_truth_df.round(4))
classifier_column_name = 'OR Classifier'
b_col_name = "Sample Size B"
plot_df[b_col_name] = np.array(plot_df['b'].values)
plot_df[classifier_column_name] = plot_df['classifier']
plot_df = plot_df[[classifier_column_name, 'theta_0_current', 'out_confint', b_col_name]].groupby(
[classifier_column_name, 'theta_0_current', b_col_name]).mean().reset_index()
fig = plt.figure(figsize=(21,6))
for jj, clf_odds in enumerate(plot_df[classifier_column_name].unique()):
temp_df = plot_df[plot_df[classifier_column_name] == clf_odds]
ax = fig.add_subplot(1, 3, jj + 1)
sns.lineplot(x='theta_0_current', y='out_confint', color=color_palette[jj],
style=b_col_name, linewidth=3, data=temp_df)
plt.xlabel(r'$\theta$' if run == 'poisson' else r'$\mu$', fontsize=24)
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
if jj == 0:
plt.ylabel('Power', fontsize=24)
else:
plt.ylabel('')
plt.ylim([0, 1.01])
plt.title("%s Classifier" % (clf_odds), fontsize=22)
plt.axvline(x=true_t0, color='red', linestyle='--')
if jj == 0 and run=='poisson':
plt.legend(loc='upper left', fontsize=19)
elif jj == 2 and run =='gmm':
plt.legend(loc='upper right', fontsize=19)
else:
ax.get_legend().remove()
title_run = 'Poisson' if run == 'poisson' else 'GMM'
plt.suptitle("Power as Function of B, %s (B'=%s, %s, n=%s)" % (
title_run, '5,000', r'$\alpha=0.9$', sample_size_val), y=1.025, fontsize=28)
image_name = 'power_plot_function_bprime%s_n%s_%s.pdf' % (5000, sample_size_val, run)
plt.savefig('images/toy_examples/' + image_name,
bbox_inches='tight')
plt.show()
print('\n')
# -
# ## Figure 6 (<NAME>.)
# +
directory = 'sims/figure6/'
files = [x for x in os.listdir(directory) if 'csv' in x]
final_df_cov = None
for flnm in files:
temp_df = pd.read_csv(directory + flnm, index_col=0)
temp_df['classifier'] = temp_df['classifier'].apply(lambda x: x.replace('\n', ''))
temp_df['classifier_cde'] = temp_df['classifier_cde'].apply(lambda x: x.replace('\n', ''))
temp_df['B'] = temp_df['b_prime']
temp_df['B_PRIME'] = temp_df['b']
if final_df_cov is None:
final_df_cov = temp_df.copy()
else:
final_df_cov = final_df_cov.append(temp_df.copy())
final_df_cov['b'] = final_df_cov['B']
final_df_cov['b_prime'] = final_df_cov['B_PRIME']
# -
print(final_df_cov['b_prime'].unique())
print(final_df_cov['b'].unique())
print(final_df_cov['classifier'].unique())
print(final_df_cov['classifier_cde'].unique())
print(final_df_cov['run'].unique())
print(final_df_cov['sample_size_obs'].unique())
print(final_df_cov['rep'].unique())
print(final_df_cov.columns)
# +
## Coverage of Plot -- Varying as a function of B
b_val = 1000
for run in ['poisson', 'gmm']:
for class_cde in ['XGBoost (d3, n500)']:
plt.figure(figsize=(12,6))
plot_df = final_df_cov[(final_df_cov['run'] == run) &
(final_df_cov['classifier_cde'] == 'XGBoost (d3, n500)') &
(final_df_cov['sample_size_obs'] == 10)]
plot_df = plot_df[plot_df['B'] == b_val]
plot_df['b_prime'] = plot_df['b_prime'].apply(lambda x: "B' = %s" % str(x))
coverage_df = plot_df[['B_PRIME', 'classifier', 'pinball_loss', 'in_confint']]
print(coverage_df.groupby(['B_PRIME', 'classifier']).agg({'in_confint': [np.average, np.std, np.min, np.max],
'pinball_loss': [np.average, np.std]}).round(2))
class_combo_name = 'Odds Class./Critical Value Class.'
b_col_name = "Sample size"
plot_df[class_combo_name] = plot_df[['classifier', 'classifier_cde']].apply(lambda x: x[0] + '/' + x[1], axis = 1)
plot_df[b_col_name] = plot_df['b_prime']
plot_df = plot_df[[class_combo_name, 'theta_0_current', 'in_confint', b_col_name]].groupby(
[class_combo_name, 'theta_0_current', b_col_name]).mean().reset_index()
sns.lineplot(x='theta_0_current', y='in_confint', style=b_col_name,
style_order=sorted(plot_df[b_col_name].unique(), key=lambda x: int(x.split('=')[1])),
data=plot_df)
plt.xlabel(r'$\theta$', fontsize=25)
plt.ylabel('Observed Coverage', fontsize=25)
plt.xticks(fontsize=20)
plt.yticks(fontsize=20)
plt.title("Observed MC Coverage as Function of %s (%s)" % (
r'$\theta$','Poisson Model' if run == 'poisson' else 'GMM'),
fontsize=28, y=1.01)
plt.axhline(y=0.9, color='red', linestyle='--')
plt.legend(loc='lower left', fontsize=20)
plt.ylim([0,1])
plt.tight_layout()
image_name = 'coverage_MC_plot_function_bbprime_b%s_classcde%s_n100_%s.pdf' % (
run, b_val, class_cde.replace(' ', '_'))
plt.savefig('images/toy_examples/' + image_name)
plt.show()
# -
# ## Figures 3 (Section 4) and 5 (Supp. Mat.)
# +
directory = 'sims/figures3-5/'
files = [x for x in os.listdir(directory) if 'csv' in x]
final_df_cov = None
for flnm in files:
temp_df = pd.read_csv(directory + flnm, index_col=0)
temp_df['classifier'] = temp_df['classifier'].apply(lambda x: x.replace('\n', ''))
temp_df['classifier_cde'] = temp_df['classifier_cde'].apply(lambda x: x.replace('\n', ''))
temp_df['B'] = temp_df['b_prime']
temp_df['B_PRIME'] = temp_df['b']
if final_df_cov is None:
final_df_cov = temp_df.copy()
else:
final_df_cov = final_df_cov.append(temp_df.copy())
final_df_cov['b'] = final_df_cov['B']
final_df_cov['b_prime'] = final_df_cov['B_PRIME']
# -
print(final_df_cov['b_prime'].unique())
print(final_df_cov['b'].unique())
print(final_df_cov['classifier'].unique())
print(final_df_cov['classifier_cde'].unique())
print(final_df_cov['run'].unique())
print(final_df_cov['sample_size_obs'].unique())
print(final_df_cov['rep'].unique())
print(final_df_cov.columns)
# +
b_val = 1000
color_vec = ['red', 'blue', 'green']
n_grid = len(final_df_cov['theta_0_current'].unique())
for run in ['poisson', 'gmm']:
plot_df = final_df_cov[(final_df_cov['run'] == run) &
(final_df_cov['classifier_cde'] == 'XGBoost (d3, n500)') &
(final_df_cov['sample_size_obs'] == 10)]
plot_df = plot_df[plot_df['b'] == b_val]
class_combo_name = 'Odds Class./Critical Value Class.'
b_col_name = "Number of Available B'"
plot_df[class_combo_name] = plot_df['classifier']
plot_df[b_col_name] = plot_df['b_prime']
plot_df = plot_df[[class_combo_name, 'theta_0_current', 'in_confint', b_col_name]].groupby(
[class_combo_name, 'theta_0_current', b_col_name]).mean().reset_index()
b_vec = np.sort(plot_df[b_col_name].unique())
fig = plt.figure(figsize=(12,6))
for ii, b_prime_val in enumerate(b_vec):
temp_df = plot_df[plot_df[b_col_name] == b_prime_val]
x = temp_df['theta_0_current'].values
y = temp_df['in_confint'].values
# estimate the model
X = sm.add_constant(x)
with Suppressor():
model = sm.Logit(y, X).fit(full_output=False)
proba = model.predict(X)
# estimate confidence interval for predicted probabilities
cov = model.cov_params()
gradient = (proba * (1 - proba) * X.T).T # matrix of gradients for each observation
std_errors = np.array([np.sqrt(np.dot(np.dot(g, cov), g)) for g in gradient])
c = 1 # multiplier for confidence interval
upper = np.maximum(0, np.minimum(1, proba + std_errors * c))
lower = np.maximum(0, np.minimum(1, proba - std_errors * c))
x_plot = x[:n_grid]
proba_plot = proba[:n_grid]
lower_plot = lower[:n_grid]
upper_plot = np.clip(upper[:n_grid], a_min=0, a_max=1.0)
ax = fig.add_subplot(3, 1, ii + 1)
sns.lineplot(x=x_plot, y=proba_plot, color=color_vec[ii], label="B'=%s" % b_prime_val)
sns.lineplot(x=x_plot, y=lower_plot, color=color_vec[ii])
sns.lineplot(x=x_plot, y=upper_plot, color=color_vec[ii])
plt.fill_between(x=x_plot, y1=lower_plot, y2=upper_plot, alpha=0.1, color=color_vec[ii])
plt.axhline(y=0.9, color='black', linestyle='--', linewidth=3)
plt.legend(loc='lower left', fontsize=20)
plt.ylim([0.5,1])
plt.xlim([plot_df['theta_0_current'].min(), plot_df['theta_0_current'].max()])
if ii == 0:
plt.title("Coverage as Function of %s (%s)" % (
r'$\theta$','Poisson Model' if run == 'poisson' else 'GMM'),
fontsize=28, y=1.01)
if ii == 2:
plt.xticks(fontsize=18)
plt.xlabel(r'$\theta$', fontsize=24)
else:
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
labelbottom=False) # labels along the bottom edge are off
plt.yticks(fontsize=18)
if ii == 1:
plt.ylabel('Estimated Coverage', fontsize=24)
image_name = 'coverage_plot_function_bprime_b%s_n100_%s.pdf' % (
b_val, run)
plt.savefig('images/toy_examples/' + image_name,
bbox_inches='tight')
plt.show()
| paper_figures/toy_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Finger 1: Análisis de datos de avisos laborales y postulaciones
# +
# importacion general de librerias y de visualizacion (matplotlib y seaborn)
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
plt.style.use('default') # haciendo los graficos un poco mas bonitos en matplotlib
plt.rcParams['figure.figsize'] = (20, 10)
sns.set(style="whitegrid") # seteando tipo de grid en seaborn
# -
applications = pd.read_csv('/Users/ignacio.iglesias/Dev/datos/fingers/finger1/datos_navent_fiuba/fiuba_4_postulaciones.csv')
applications.head()
online_offers = pd.read_csv('/Users/ignacio.iglesias/Dev/datos/fingers/finger1/datos_navent_fiuba/fiuba_5_avisos_online.csv')
online_offers.head()
offers = pd.read_csv('/Users/ignacio.iglesias/Dev/datos/fingers/finger1/datos_navent_fiuba/fiuba_6_avisos_detalle.csv')
offers.head()
applications['fechapostulacion'].value_counts()
applications['fechapostulacion'].head()
applications['fechapostulacion'].isnull().head()
applications['fechapostulacion_dt'] = pd.to_datetime(applications['fechapostulacion'])
applications.head()
applications['fechapostulacion_dt'].head()
import datetime
applications['fechapostulacion_dt'][2].weekday()
from datetime import date
import calendar
calendar.day_name[applications['fechapostulacion_dt'][0].weekday()]
type(applications[['fechapostulacion_dt']])
applications['weekday'] = applications['fechapostulacion_dt'].apply(lambda x: x.weekday())
applications.head()
applications['weekday'] = applications['weekday'].apply(lambda x: calendar.day_name[x])
applications.head()
applications_by_weekday = applications['weekday'].value_counts()
applications_by_weekday
g = sns.barplot(x=applications_by_weekday.values, y=applications_by_weekday.index, orient='h')
g.set_title("Postulaciones según el día de la semana", fontsize=15)
g.set_xlabel("Número de postulaciones", fontsize=12)
g.set_ylabel("Día de la semana", fontsize=12)
type(applications['fechapostulacion_dt'][0])
applications_by_year = applications['fechapostulacion_dt'].apply(lambda x: x.year)
applications_by_year.value_counts()
applications_by_month = applications['fechapostulacion_dt'].apply(lambda x: x.month).value_counts()
applications_by_month
applications_by_date = applications['fechapostulacion_dt'].apply(lambda x: x.date()).value_counts()
applications_by_date
g = applications_by_date.plot(color='lightblue')
g.set_title("Postulaciones en el tiempo", fontsize=18)
g.set_xlabel("Fecha",fontsize=18)
g.set_ylabel("Cantidad de postulaciones", fontsize=18)
# g.set(xticks=applications_by_date)
applications.groupby(['weekday']).agg({'weekday':'count'}).sort_values('weekday', ascending=False)
applications.head()
offers.head()
offers.groupby(['nombre_area']).agg({'nombre_area':'count'}).sort_values('nombre_area', ascending=False).head(20)
top20 = offers.groupby(['nombre_area']).agg({'nombre_area':'count'}).sort_values('nombre_area', ascending=False).head(20)
top20['nombre_area'].values
g = sns.barplot(x=top20['nombre_area'].values, y=top20['nombre_area'].index, orient='h')
g.set_title("Top 20 de avisos por área", fontsize=15)
g.set_xlabel("Cantidad de avisos", fontsize=12)
g.set_ylabel("Área de trabajo", fontsize=12)
grupped_by_area = offers.groupby(['nombre_area']).agg({'nombre_area':'count'}).sort_values('nombre_area', ascending=False)
grupped_by_area = offers.groupby(['nombre_area'])
dfVentas = grupped_by_area.get_group('Ventas')
type(dfVentas)
dfZonasVentas = dfVentas.groupby(['nombre_zona']).agg({'nombre_zona':'count'}).sort_values('nombre_zona', ascending=False)
dfZonasVentas
g = sns.barplot(x=dfZonasVentas['nombre_zona'].values, y=dfZonasVentas['nombre_zona'].index, orient='h')
g.set_title("Avisos de ventas distribuidas por zonas", fontsize=15)
g.set_xlabel("Cantidad de avisos", fontsize=12)
g.set_ylabel("Zonas", fontsize=12)
g = sns.barplot(x=dfZonasVentas['nombre_zona'].index, y=dfZonasVentas['nombre_zona'].values, orient='v')
g.set_title("Avisos de ventas distribuidas por zonas", fontsize=15)
g.set_ylabel("Cantidad de avisos", fontsize=12)
g.set_xlabel("Zonas", fontsize=12)
| fingers/01/Finger 01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 3
import spacy
from spacy.tokens import Span, Doc
from spacy.matcher import Matcher, PhraseMatcher
nlp = spacy.load('en_core_web_sm')
print(nlp.pipe_names)
# +
# @Language.component('my_pipeline')
def my_pipeline(doc):
print('Doc length:', len(doc))
return doc
nlp.add_pipe(my_pipeline)
print(nlp.pipe_names)
# -
| Courses/spaCy/.ipynb_checkpoints/Chapter 3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="n5oInnkZwjwm" colab_type="code" colab={}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# + id="hLVN_Ave_c2-" colab_type="code" colab={}
# + id="1wzEF7p8wlMT" colab_type="code" colab={}
x=pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_features_train.csv')
y=pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_labels_train.csv')
x_test=pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_features_test.csv')
submission_file=pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/submission_format.csv')
# + id="qRivwzaAwwmc" colab_type="code" outputId="f5d25510-0885-490e-c783-af126dc9a53e" colab={"base_uri": "https://localhost:8080/", "height": 521}
x.fillna(method='ffill', inplace=True)
x_test.fillna(method='ffill', inplace=True)
x.info()
# + id="MICvQ6zF7BXH" colab_type="code" colab={}
features=['city','year','weekofyear','ndvi_se','station_avg_temp_c','ndvi_sw','reanalysis_dew_point_temp_k','reanalysis_air_temp_k','ndvi_ne','reanalysis_max_air_temp_k','reanalysis_min_air_temp_k','ndvi_nw','reanalysis_tdtr_k','precipitation_amt_mm','reanalysis_precip_amt_kg_per_m2','station_precip_mm']
# + id="QQPZ_0r_7MAD" colab_type="code" outputId="f3a3f26c-446c-4b2e-dd45-c40055a2fb29" colab={"base_uri": "https://localhost:8080/", "height": 382}
X=x[features]
X.info()
# + id="-ddUNjmL7aIp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 208} outputId="92b4baf5-9e16-41ac-e3fc-788f54b4eac3"
X['city1']=np.where(X['city']=='sj',1,-1)
X.drop(columns='city',inplace=True)
# + id="CDmfICBc_YXf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 382} outputId="f8644b9e-fd70-4e32-b227-154853896e30"
X.info()
# + id="Pm6k_0_G_lU_" colab_type="code" colab={}
y.drop(columns='city',inplace=True)
y.drop(columns='year',inplace=True)
y.drop(columns='weekofyear',inplace=True)
# + id="fyn3sgivALuu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="1fbaa47c-6827-4064-8e7b-62b08fd298f3"
y.info()
# + id="rhx6xb3yANVD" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
seed = 5
test_size = 0.3
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size, random_state=seed)
# + id="cOEWrSuKARso" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="ae4edabd-6708-4a05-f458-9f42aa679aa7"
from xgboost import XGBRegressor
my_model = XGBRegressor()
my_model.fit(X_train, y_train)
# + id="sUwfSYkwAWzC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="37326bf8-83b4-4f0e-be60-18ba0013dba2"
from sklearn.metrics import mean_absolute_error
predictions = my_model.predict(X_test)
print("Mean Absolute Error: " + str(mean_absolute_error(predictions, y_test)))
# + id="1LkkBQUIAc5n" colab_type="code" colab={}
| relevent_features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
#
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# # Plotly - Create Mapchart world
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Plotly/Plotly_Create_Mapchart_world.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# **Tags:** #plotly #chart #worldmap #dataviz #snippet #operations #image #html
# + [markdown] papermill={} tags=["naas", "awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# **Author:** [<NAME>](https://www.linkedin.com/in/ACoAAAJHE7sB5OxuKHuzguZ9L6lfDHqw--cdnJg/)
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ## Input
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ### Import libraries
# + papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
import naas
import plotly.graph_objects as go
import pandas as pd
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ### Variables
# + papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
title = "Worldmap"
# Output paths
output_image = f"{title}.png"
output_html = f"{title}.html"
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ### Get data
# Columns :
# 1. ISO code of country
# 2. Value
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# To use the built-in countries geometry, provide locations as [three-letter ISO country codes](https://en.wikipedia.org/wiki/ISO_3166-1_alpha-3).
# + papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/2014_world_gdp_with_codes.csv')
df
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ## Model
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ### Create the plot
# + papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
fig = go.Figure()
fig = go.Figure(data=go.Choropleth(
locations = df['CODE'],
z = df['GDP (BILLIONS)'],
text = df['COUNTRY'],
colorscale = 'Blues',
autocolorscale=False,
reversescale=True,
marker_line_color='darkgray',
marker_line_width=0.5,
colorbar_tickprefix = '$',
colorbar_title = 'GDP<br>Billions US$',
))
fig.update_layout(
title=title ,
plot_bgcolor="#ffffff",
legend_x=1,
geo=dict(
showframe=False,
showcoastlines=False,
#projection_type='equirectangular'
),
dragmode= False,
width=1200,
height=800,
)
config = {'displayModeBar': False}
fig.show(config=config)
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ## Output
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ### Export in PNG and HTML
# + papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
fig.write_image(output_image, width=1200)
fig.write_html(output_html)
# + [markdown] papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
# ### Generate shareable assets
# + papermill={} tags=["awesome-notebooks/Plotly/Plotly_Create_Mapchart_world.ipynb"]
link_image = naas.asset.add(output_image)
link_html = naas.asset.add(output_html, {"inline":True})
#-> Uncomment the line below to remove your assets
# naas.asset.delete(output_image)
# naas.asset.delete(output_html)
| Plotly/Plotly_Create_Mapchart_world.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Problem 21
# ## Amicable numbers
#
# Let $d(n)$ be defined as the sum of proper divisors of $n$ (numbers less than $n$ which divide evenly into $n$).
#
# If $d(a) = b$ and $d(b) = a$, where $a \ne b$, then $a$ and $b$ are an amicable pair and each of $a$ and $b$ are called amicable numbers.
#
# For example, the proper divisors of $220$ are $1, 2, 4, 5, 10, 11, 20, 22, 44, 55$ and $110$; therefore $d(220) = 284$. The proper divisors of $284$ are $1, 2, 4, 71$ and $142$; so $d(284) = 220$.
#
# Evaluate the sum of all the amicable numbers under $10000$.
#
# OEIS Sequence: [A063990](https://oeis.org/A063990)
#
# ## Solution
# + pycharm={"name": "#%%\n"}
from math import isqrt
from euler.primes import prime_numbers
from euler.numbers import sum_proper_divisors
# + pycharm={"name": "#%%\n"}
def compute(n: int) -> int:
primes = list(prime_numbers(isqrt(n)))
sum_factors = [0] * (n + 1)
result = 0
for i in range(1, n + 1):
sum_factors[i] = sum_proper_divisors(i)
for i in range(2, n + 1):
j = sum_factors[i]
if j != i and j <= n and sum_factors[j] == i:
result += i
return result
# + pycharm={"name": "#%%\n"}
compute(1_000)
# + pycharm={"name": "#%%\n"}
compute(10_000)
# + pycharm={"name": "#%%\n"}
# %timeit -n 100 -r 1 -p 6 compute(10_000)
| problems/0021/solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classify different data sets
# ### Basic includes
# +
# Using pandas to load the csv file
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from keras import models
from keras import layers
from keras import callbacks
from keras.utils import to_categorical
# reuters and fashin mnist data set from keras
from keras.datasets import reuters
from keras.datasets import fashion_mnist
# needed to preprocess text
from keras.preprocessing.text import Tokenizer
# -
# ### Classify the Fashion Mnist
#
# ---
# +
(fashion_train_data, fashion_train_labels), (fashion_test_data, fashion_test_labels) = fashion_mnist.load_data()
print(fashion_train_data.shape)
test_index = 10
plt.title("Label: " + str(fashion_train_labels[test_index]))
plt.imshow(fashion_train_data[test_index], cmap="gray")
# -
# #### TO DO: Preprocess the data
#
# 1. Normalize the input data set
# 2. Perform one hot encoding
# 3. Create a train, test, and validation set
# #### TO DO: Define and train a network, then plot the accuracy of the training, validation, and testing
#
# 1. Use a validation set
# 2. Propose and train a network
# 3. Print the history of the training
# 4. Evaluate with a test set
fashion_train_data = fashion_train_data.reshape((60000, 28 * 28))# Normalizar datos
fashion_train_data = fashion_train_data.astype('float32') / 255# Normalizar datos
fashion_test_data = fashion_test_data.reshape((10000, 28 * 28))
fashion_test_data = fashion_test_data.astype('float32') / 255
fashion_train_labels = to_categorical(fashion_train_labels) #hot encoding
fashion_test_labels = to_categorical(fashion_test_labels)#hot encoding
validation_data = fashion_train_data[:30000] #Validación
validation_labels = fashion_train_labels[:30000] #Validación
x_data = fashion_train_data[30000:]
y_data = fashion_train_labels[30000:]
network = models.Sequential()
network.add(layers.Dense(128, activation='relu', input_shape= (28 * 28,)))
network.add(layers.Dropout(0.3))
network.add(layers.Dense(64, activation='relu'))
network.add(layers.Dropout(0.3))
network.add(layers.Dense(10, activation='softmax'))
network.summary()
early_stop = callbacks.EarlyStopping(monitor='val_loss', patience=2)
network.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
history = network.fit(x_data, y_data, epochs=25, validation_data = (validation_data, validation_labels), callbacks=[early_stop], verbose=2)
# +
test_loss, test_acc = network.evaluate(fashion_test_data, fashion_test_labels)
print("test loss: ", test_loss, "test accuracy: ", test_acc) #imprimir evaluación
history_dict = history.history
acc = history_dict['acc']
val_acc = history_dict['val_acc']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, loss, 'bo', label='Training loss') #graficas
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# +
plt.clf()
plt.plot(epochs, acc, 'bo', label='Training acc') #Grñafica de entrenamiento
plt.plot(epochs, val_acc, 'b', label='Validation acc') #Gráfica de validación
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# -
# ## Classifying newswires
#
# ---
#
# Build a network to classify Reuters newswires into 46 different mutually-exclusive topics.
# ### Load and review the data
# +
(reuters_train_data, reuters_train_labels), (reuters_test_data, reuters_test_labels) = reuters.load_data(num_words=10000)
print(reuters_train_data.shape)
print(reuters_train_labels.shape)
print(reuters_train_data[0])
print(reuters_train_labels[0])
print(set(reuters_train_labels))
# -
# Load the word index to decode the train data.
# +
word_index = reuters.get_word_index()
reverse_index = dict([(value+3, key) for (key, value) in word_index.items()])
reverse_index[0] = "<PAD>"
reverse_index[1] = "<START>"
reverse_index[2] = "<UNKNOWN>" # unknown
reverse_index[3] = "<UNUSED>"
decoded_review = ' '.join([reverse_index.get(i,'?') for i in reuters_train_data[0]])
print(decoded_review)
# -
# #### TO DO: Preprocess the data
#
# 1. Normalize the input data set
# 2. Perform one hot encoding
# 3. Create a train, test, and validation set
# #### TO DO: Define and train a network, then plot the accuracy of the training, validation, and testing
#
# 1. Use a validation set
# 2. Propose and train a network
# 3. Print the history of the training
# 4. Evaluate with a test set
# +
tokenizer = Tokenizer(num_words=8982)
train_data_token = tokenizer.sequences_to_matrix(reuters_train_data, mode='binary') #Normalizando
test_data_token = tokenizer.sequences_to_matrix(reuters_test_data, mode='binary')
one_hot_train_labels = to_categorical(reuters_train_labels) #hot encoding
one_hot_test_labels = to_categorical(reuters_test_labels)
# -
validation_data = train_data_token[:3000] #validación
validation_labels = one_hot_train_labels[:3000]
x_data = train_data_token[3000:]
y_data = one_hot_train_labels[3000:]
# +
network = models.Sequential()
network.add(layers.Dense(92, activation='relu', input_shape= (8982,)))
network.add(layers.Dropout(0.3))
network.add(layers.Dense(46, activation='softmax'))
network.add(layers.Dropout(0.2))
network.summary()
# -
early_stop = callbacks.EarlyStopping(monitor='val_loss', patience=6)
network.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
history = network.fit(x_data, y_data, epochs=15, validation_data = (validation_data, validation_labels), callbacks=[early_stop], verbose=2) #entrenamiento
test_loss, test_acc = network.evaluate(test_data_token, one_hot_test_labels) #evaluando
print("test loss: ", test_loss, "test accuracy: ", test_acc)
# +
history_dict = history.history
acc = history_dict['acc']
val_acc = history_dict['val_acc']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
# +
plt.plot(epochs, loss, 'bo', label='Training loss') #gráfica
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# +
plt.clf()
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy') #gráfica de validación
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# -
# ## Predicting Student Admissions
#
# ---
#
# Predict student admissions based on three pieces of data:
#
# - GRE Scores
# - GPA Scores
# - Class rank
# ### Load and visualize the data
student_data = pd.read_csv("data/student_data.csv") #se creo carpeta data
print(student_data)
# Plot of the GRE and the GPA from the data.
# +
X = np.array(student_data[["gre","gpa"]])
y = np.array(student_data["admit"])
admitted = X[np.argwhere(y==1)]
rejected = X[np.argwhere(y==0)]
plt.scatter([s[0][0] for s in rejected], [s[0][1] for s in rejected], s = 25, color = 'red', edgecolor = 'k')
plt.scatter([s[0][0] for s in admitted], [s[0][1] for s in admitted], s = 25, color = 'cyan', edgecolor = 'k')
plt.xlabel('Test (GRE)')
plt.ylabel('Grades (GPA)')
plt.show()
# -
# Plot of the data by class rank.
# +
f, plots = plt.subplots(2, 2, figsize=(20,10))
plots = [plot for sublist in plots for plot in sublist]
for idx, plot in enumerate(plots):
data_rank = student_data[student_data["rank"]==idx+1]
plot.set_title("Rank " + str(idx+1))
X = np.array(data_rank[["gre","gpa"]])
y = np.array(data_rank["admit"])
admitted = X[np.argwhere(y==1)]
rejected = X[np.argwhere(y==0)]
plot.scatter([s[0][0] for s in rejected], [s[0][1] for s in rejected], s = 25, color = 'red', edgecolor = 'k')
plot.scatter([s[0][0] for s in admitted], [s[0][1] for s in admitted], s = 25, color = 'cyan', edgecolor = 'k')
plot.set_xlabel('Test (GRE)')
plot.set_ylabel('Grades (GPA)')
# -
# #### TO DO: Preprocess the data
#
# 1. Normalize the input data set
# 2. Perform one hot encoding
# 3. Create a train, test, and validation set
# #### TO DO: Define and train a network, then plot the accuracy of the training, validation, and testing
#
# 1. Use a validation set
# 2. Propose and train a network
# 3. Print the history of the training
# 4. Evaluate with a test set
# +
student_data = student_data.fillna(0)
normalized_student_data = pd.get_dummies(student_data, columns=['rank']) #normalizando
normalized_student_data["gre"] = normalized_student_data["gre"] / 800
normalized_student_data["gpa"] = normalized_student_data["gpa"] / 4
np.random.shuffle(normalized_student_data.values)
student_x = np.array(normalized_student_data)[:,1:]
student_x = student_x.astype('float32')
student_y = to_categorical(student_data["admit"])
# +
student_validation_data = student_x[:100] #validando
student_validation_labels = student_y[:100]
student_x_data = student_x[100:]
student_y_data = student_y[100:]
# +
student_NN_model = models.Sequential([
layers.Dense(128, activation='relu', kernel_initializer='random_uniform', input_shape=(7,)),
layers.Dropout(0.3),
layers.Dense(64, activation='relu'),
layers.Dropout(0.3),
layers.Dense(32, activation='relu'),
layers.Dense(2, activation='softmax')
])
student_NN_model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
student_NN_model.summary()
student_history = student_NN_model.fit(student_x_data, student_y_data, epochs=15, batch_size=100, validation_data=(student_validation_data,student_validation_labels))
student_results = student_NN_model.evaluate(student_x[80:200], student_y[80:200])
print('student Test accuracy:', student_results)
# +
student_history_dict = student_history.history
print(student_history_dict.keys())
student_acc = student_history_dict['acc']
student_val_acc = student_history_dict['val_acc']
student_loss = student_history_dict['loss']
student_val_loss = student_history_dict['val_loss']
student_epochs = range(1, len(student_acc) + 1)
# +
plt.plot(student_epochs, student_loss, 'bo', label='Training loss')
plt.plot(student_epochs, student_val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# +
plt.clf()
plt.plot(student_epochs, student_acc, 'bo', label='Training acc')
plt.plot(student_epochs, student_val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# -
| Keras_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import division
from collections import OrderedDict
import time
import copy
import pickle
import os
import random
import pandas as pd
import numpy as np
import sched
# -
from matplotlib import pyplot as plt
import seaborn as sns
# %matplotlib inline
import matplotlib as mpl
mpl.rc('savefig', dpi=300)
mpl.rc('text', usetex=True)
def sample_arrival_times(all_items, arrival_rate, start_time):
"""
Sample item arrival times for init_data['arrival_time_of_item'],
which gets passed to the StatelessLQNScheduler constructor
:param set[str] all_items: A set of item ids
:param float arrival_rate: The arrival rate for the Poisson process
:param int start_time: Start time (unix epoch) for the arrival process
"""
all_items = list(all_items)
random.shuffle(all_items)
inter_arrival_times = np.random.exponential(1 / arrival_rate, len(all_items))
arrival_times = start_time + np.cumsum(inter_arrival_times, axis=0).astype(int)
return OrderedDict(zip(all_items, arrival_times))
# Sanity check
# +
init_data = {
'arrival_time_of_item' : {0: int(time.time())},
'review_rates' : np.array([0.25, 0.25, 0.25, 0.25])[np.newaxis, :],
'difficulty_of_item' : {0: 0.01},
'difficulty_rate' : 100,
'max_num_items_in_deck' : None
}
scheduler = sched.ExtLQNScheduler(init_data)
history = []
assert scheduler.next_item() == 0
# -
# Simulations
global_item_difficulty = 0.0076899999999998905
difficulty_rate = 1 / global_item_difficulty
using_global_difficulty = True
num_items = 50
difficulty_of_item = np.ones(num_items) * global_item_difficulty if using_global_difficulty else np.random.exponential(1 / difficulty_rate, num_items)
arrival_rate = 0.05
num_timesteps_in_sim = 1000
all_items = range(num_items)
start_time = int(time.time())
init_data = {
'arrival_time_of_item' : sample_arrival_times(all_items, arrival_rate, start_time),
'review_rates' : np.array([[0.125, 0.125, 0.125, 0.125], [0.125, 0.125, 0.125, 0.125]]),
'difficulty_of_item' : {i: x for i, x in enumerate(difficulty_of_item)},
'difficulty_rate' : difficulty_rate,
'max_num_items_in_deck' : None
}
scheduler = sched.ExtLQNScheduler(init_data)
num_systems = len(init_data['review_rates'])
num_decks = len(init_data['review_rates'][0])
work_rate = 0.19020740740740741
inter_arrival_times = np.random.exponential(1 / work_rate, num_timesteps_in_sim)
timesteps = int(time.time()) + np.cumsum(inter_arrival_times, axis=0).astype(int)
# +
history = []
deck_of_item = {item: 1 for item in all_items}
latest_timestamp_of_item = {item: 0 for item in all_items}
for current_time in timesteps:
try:
next_item = scheduler.next_item(current_time=current_time)
except sched.ExhaustedError:
continue
delay = current_time - latest_timestamp_of_item[next_item]
latest_timestamp_of_item[next_item] = current_time
deck = deck_of_item[next_item]
outcome = 1 if np.random.random() < np.exp(-difficulty_of_item[next_item] * delay / deck) else 0
deck_of_item[next_item] = max(1, deck + 2 * outcome - 1)
history.append({'item_id' : next_item, 'outcome' : outcome, 'timestamp' : current_time})
scheduler.update(next_item, outcome, current_time)
# -
df = pd.DataFrame(history)
np.mean(df['outcome'])
def deck_promotion_rates(init_data, history):
"""
Compute the observed rates at which items move from deck i to deck i+1
:param pd.DataFrame history: The logs for a single user
:rtype: list[float]
:return: The average promotion rate (items per second) for each deck
"""
deck_of_item = {item: 1 for item in init_data['arrival_time_of_item']}
num_decks = len(init_data['review_rates'][0])
num_promotions_of_deck = {deck: 0 for deck in xrange(1, num_decks + 1)}
for ixn in history:
item = ixn['item_id']
outcome = ixn['outcome']
current_deck = deck_of_item[item]
if outcome == 1:
if current_deck >= 1 and current_deck <= num_decks:
num_promotions_of_deck[current_deck] += 1
deck_of_item[item] += 1
elif outcome == 0 and current_deck > 1:
deck_of_item[item] -= 1
duration = max(ixn['timestamp'] for ixn in history) - min(ixn['timestamp'] for ixn in history)
promotion_rate_of_deck = {deck: (num_promotions / (1 + duration)) for deck, num_promotions in num_promotions_of_deck.iteritems()}
return promotion_rate_of_deck
deck_promotion_rates(init_data, history)
max_num_items_in_deck = None
def run_sim(arrival_rate, num_items, difficulty_rate, difficulty_of_item, review_rates, work_rate, num_timesteps_in_sim, expected_recall_likelihoods=None):
all_items = range(num_items)
start_time = int(time.time())
init_data = {
'arrival_time_of_item' : sample_arrival_times(all_items, arrival_rate, start_time),
'review_rates' : review_rates,
'difficulty_of_item' : {i: x for i, x in enumerate(difficulty_of_item)},
'difficulty_rate' : difficulty_rate,
'max_num_items_in_deck' : max_num_items_in_deck
}
num_decks = len(init_data['review_rates'][0])
scheduler = sched.ExtLQNScheduler(init_data)
history = []
deck_of_item = {item: 1 for item in all_items}
latest_timestamp_of_item = {item: 0 for item in all_items}
inter_arrival_times = np.random.exponential(1 / work_rate, num_timesteps_in_sim)
timesteps = int(time.time()) + np.cumsum(inter_arrival_times, axis=0).astype(int)
for current_time in timesteps:
try:
next_item = scheduler.next_item(current_time=current_time)
except sched.ExhaustedError:
continue
deck = deck_of_item[next_item]
if expected_recall_likelihoods is None:
delay = current_time - latest_timestamp_of_item[next_item]
recall_likelihood = np.exp(-difficulty_of_item[next_item] * delay / deck)
else:
recall_likelihood = expected_recall_likelihoods[deck-1]
outcome = 1 if np.random.random() < recall_likelihood else 0
latest_timestamp_of_item[next_item] = current_time
deck_of_item[next_item] = max(1, deck + 2 * outcome - 1)
history.append({'item_id' : next_item, 'outcome' : outcome, 'timestamp' : current_time})
scheduler.update(next_item, outcome, current_time)
if history == []:
return 0
promotion_rate_of_deck = deck_promotion_rates(init_data, history)
return promotion_rate_of_deck[num_decks]
num_sim_repeats = 10
num_systems = 1
num_decks = 5
work_rate = 0.19020740740740741
num_timesteps_in_sim = 500
review_rates = 1 / np.sqrt(np.arange(1, num_decks + 1, 1))
review_rates /= review_rates.sum()
review_rates = review_rates[np.newaxis, :]
run_sim(1., num_items, difficulty_rate, difficulty_of_item, review_rates, work_rate, num_timesteps_in_sim)
std_err = lambda x: np.nanstd(x) / np.sqrt(len(x))
# Compared simulations with clocked delay to simulations with the mean-recall approximation
arrival_rates = np.arange(0.001, 0.01+1e-6, 0.0005)
# from lqn_properties.ipynb
expected_recall_likelihoods = [[0.8816669326889862,0.912726114951097,0.92719714973377,0.9360503409439428,0.9423109652525123],
[0.8802159353708854,0.9112980491805251,0.9257778382415257,0.9346388744415259,0.94097809417084],
[0.8787197882984757,0.9098132332840261,0.9242928082947981,0.9331551225315882,0.9395782276183158],
[0.8771758043736176,0.9082675509248022,0.9227366399542867,0.9315926867387265,0.9381058586147527],
[0.8755810321617427,0.9066564105056785,0.9211032176252993,0.9299443133327762,0.9365548225241448],
[0.8739322170539264,0.9049746654250952,0.9193856028066731,0.9282017344908691,0.9349181903609529],
[0.8722257544837021,0.9032165160944434,0.9175758756485984,0.9263554706857485,0.9331881396027241],
[0.8704576330412577,0.9013753882808239,0.915664935506246,0.9243945823202996,0.9313557965386431],
[0.8686233645757845,0.8994437802941111,0.9136422468301503,0.9223063540631219,0.9294110422347164],
[0.8667178972966402,0.8974130685424716,0.9114955110493042,0.9200758886676667,0.9273422714786288],
[0.864735506299407,0.8952732565009774,0.909210236521236,0.9176855771145564,0.9251360902192634],
[0.8626696535627787,0.8930126452763714,0.9067691653630414,0.91511439677798,0.9227769314748718],
[0.8605128057906617,0.8906173931562718,0.9041514949450417,0.9123369657072056,0.9202465615555917],
[0.858256192636374,0.8880709140279719,0.9013317974714279,0.9093222432855949,0.9175234362738623],
[0.8558894786472526,0.8853530375953804,0.8982784973778067,0.906031726907586,0.9145818567364079],
[0.8534003061323808,0.8824387858876376,0.8949515522602463,0.9024167152103361,0.9113907773426282],
[0.8507736192636972,0.8792965769000917,0.8912991768515758,0.8984146080316402,0.9079123035650984],
[0.8479907015893672,0.8758854112910051,0.8872523574499168,0.8939427107145428,0.9040993839676039],
[0.8450275725143461,0.8721502713471428,0.8827160178057104,0.8888886821757709,0.8998926094017514]]
assert len(expected_recall_likelihoods) == len(arrival_rates)
ys = [[run_sim(x, num_items, difficulty_rate, difficulty_of_item, review_rates, work_rate-x, num_timesteps_in_sim) for _ in xrange(num_sim_repeats)] for x in arrival_rates]
exp_ys = [[run_sim(x, num_items, difficulty_rate, difficulty_of_item, review_rates, work_rate-x, num_timesteps_in_sim, expected_recall_likelihoods=y) for _ in xrange(num_sim_repeats)] for x, y in zip(arrival_rates, expected_recall_likelihoods)]
mean_ys = [np.mean(y) for y in ys]
std_err_ys = [std_err(y) for y in ys]
mean_exp_ys = [np.mean(y) for y in exp_ys]
std_err_exp_ys = [std_err(y) for y in exp_ys]
plt.xlabel(r'Arrival Rate $\lambda_{ext}$ (Items Per Second)')
plt.ylabel(r'Throughput $\lambda_{out}$ (Items Per Second)')
plt.errorbar(arrival_rates, mean_exp_ys, yerr=std_err_exp_ys, label='Simulated (Mean-Recall Approximation)')
plt.errorbar(arrival_rates, mean_ys, yerr=std_err_ys, label='Simulated (Clocked Delay)')
plt.plot(np.arange(arrival_rates[0], arrival_rates[-1], 0.0001), np.arange(arrival_rates[0], arrival_rates[-1], 0.0001), '--', label='Theoretical Steady-State Behavior')
plt.legend(loc='best')
plt.savefig(os.path.join('figures', 'lqn', 'clocked-vs-expected-delays.pdf'))
plt.show()
with open(os.path.join('results', 'clocked-vs-expected-delays.pkl'), 'wb') as f:
pickle.dump((arrival_rates, ys, exp_ys), f, pickle.HIGHEST_PROTOCOL)
# Compare theoretical phase transition threshold to simulations
arrival_rates = np.arange(0.001, 0.15, 0.005)
theoretical_phase_transition_threshold = 0.013526062011718753 # from lqn_properties.ipynb
ys = [[run_sim(x, num_items, difficulty_rate, difficulty_of_item, review_rates, work_rate-x, num_timesteps_in_sim) for _ in xrange(num_sim_repeats)] for x in arrival_rates]
plt.xlabel(r'Arrival Rate $\lambda_{ext}$ (Items Per Second)')
plt.ylabel(r'Throughput $\lambda_{out}$ (Items Per Second)')
plt.errorbar(arrival_rates, [np.mean(y) for y in ys], yerr=[std_err(y) for y in ys], label='Simulated (Clocked Delay)')
plt.axvline(x=theoretical_phase_transition_threshold, label=r'Phase Transition Threshold (Theoretical)', linestyle='--')
plt.legend(loc='best')
plt.savefig(os.path.join('figures', 'lqn', 'theoretical-vs-simulated-phase-transition.pdf'))
plt.show()
with open(os.path.join('results', 'theoretical-vs-simulated-phase-transition.pkl'), 'wb') as f:
pickle.dump((arrival_rates, ys, theoretical_phase_transition_threshold), f, pickle.HIGHEST_PROTOCOL)
# Compare simulations of different lengths (i.e., transient vs. steady-state behavior)
arrival_rates = np.arange(0.001, 0.15, 0.0001)
sim_lengths = [500, 1000, 5000, 10000]
num_items = 500
difficulty_of_item = np.ones(num_items) * global_item_difficulty if using_global_difficulty else np.random.exponential(global_item_difficulty, num_items)
ys = [[[run_sim(x, num_items, difficulty_rate, difficulty_of_item, review_rates, work_rate-x, y) for _ in xrange(num_sim_repeats)] for x in arrival_rates] for y in sim_lengths]
plt.xlabel(r'Arrival Rate $\lambda_{ext}$ (Items Per Second)')
plt.ylabel(r'Throughput $\lambda_{out}$ (Items Per Second)')
for nts, ds in zip(sim_lengths, ys):
plt.errorbar(
arrival_rates, [np.mean(y) for y in ds], yerr=[std_err(y) for y in ds],
label='Simulated Session Length = %d Reviews' % nts)
plt.axvline(x=theoretical_phase_transition_threshold, label=r'Phase Transition Threshold (Theoretical)', linestyle='--')
plt.legend(loc='best')
#plt.savefig(os.path.join('figures', 'lqn', 'throughput-vs-arrival-rate-vs-simulated-session-length.pdf'))
plt.show()
with open(os.path.join('results', 'throughput-vs-arrival-rate-vs-simulated-session-length.pkl'), 'wb') as f:
pickle.dump((arrival_rates, ys, sim_lengths), f, pickle.HIGHEST_PROTOCOL)
# If difficulties are item-specific, does creating parallel queueing systems help?
arrival_rates = np.arange(0.001, 0.15, 0.001)
sim_length = 1000
num_systems_set = range(1, 11)
# from ext_lqn_properties.ipynb
optimal_review_rates = [[0.045399259994105136,0.03847275719791948,0.03431551304800672,0.031248391825640848,0.02572273483748578],
[0.01607388427821009,0.028716279906786,0.01418113985931187,0.02399628349725895,0.013071220952213317,
0.021141555920711676, 0.012316601866189176,0.018942481766948866,0.010864112665063222,0.015106339580169872],
[0.009574995362327697,0.013960067171853526,0.02109463480774494,0.00858820264078027,0.011980694695535053,
0.017530535748704914,0.00801450278574112,0.01080151621076895,0.015369501281372995,0.0076332471983635635,
0.009958108467235074,0.013670630601002836,0.006884757991376857,0.008401291880749036,0.010749861464653288],
[0.006714438008920156,0.009237688453210182,0.011835423783583859,0.016773213565409055,0.006086273015367501,
0.00803496737482064,0.010050219385669908,0.013894393221450899,0.005722942578342616,0.007324062335706905,
0.008980026869794863,0.012146727442443267,0.00548448429102778,0.006829139163046215,0.008194754766998413,
0.010754894496614424,0.005011122270095049,0.005895292937864199,0.006774075922036427,0.008383621517784401],
[0.005127001550263364,0.006820580820242886,0.008371531015656583,0.010225672065252102,0.013979049421944282,
0.004682866183510203,0.005989275620204075,0.007190541306872763,0.00863085347246327,0.011554290753403687,
0.00442688588752093,0.005500536428474011,0.006487332446142775,0.0076714396465655905,0.010081243070342938,
0.004260252953890548,0.005165797901650487,0.005985285503196308,0.00695558005354823,0.00889706475211073,
0.003927023179050156,0.004525506061992437,0.005057213183584693,0.005676940018220873,0.00689257709346182],
[0.00412572602148644,0.005362992945624901,0.006429352327291899,0.007564488283251954,0.009001759456367844,
0.012017043371052446,0.003790559014989381,0.004744148390001485,0.005569119841589949,0.006449655310347438,
0.0075670530359376416,0.009916297749941835,0.003597894068777136,0.004381775745773768,0.005059433818363058,
0.005782936024405004,0.006702006503247498,0.008639533531856102,0.0034732157615651534,0.004136386902729236,
0.004701815726974527,0.005298739572206223,0.006048611381527559,0.00760565575491652,0.003222540185218408,
0.003662443460112906,0.00403136487467274,0.004415692124334982,0.004892211612597641,0.005864239644389311],
[0.0034401633349379116,0.004394704157125355,0.005186881477660381,0.0059814445727146195,0.006876825737061398,
0.008046795771934807,0.010559628117936837,0.0031757313190138073,0.003910920837911669,0.004523219770707268,
0.005138901320564553,0.005834131631513151,0.006744225452490461,0.008702401851729078,0.0030240401605374175,
0.0036285269860739803,0.004131519129134704,0.004637289911571074,0.005208717488592617,0.00595760008874033,
0.0075733194721484855,0.002926320252875975,0.003438916136205053,0.0038600836076054467,0.00427936448089452,
0.004748680369203616,0.00535773907644157,0.006653618536501655,0.0027290330866415486,0.003070008926448823,
0.003345952442723963,0.003617426242653257,0.003917980127278529,0.004303562296616209,0.005110681849743032],
[0.0029431275158235135,0.0037082224895140787,0.0043268822702673135,0.004925013161090805,0.005560113570175186,
0.006298239392313956,0.007282382651210947,0.009431981371507974,0.002727649005498385,0.0033166049799372727,
0.0037944349700944654,0.004257502451900565,0.0047501414270776,0.005323661589448345,0.006089516377738369,
0.007764881354195466,0.002604247725947978,0.0030885985788751606,0.0034811663437407433,0.003861538854976871,
0.004266312422755438,0.004737857206584536,0.005368292619809832,0.006751192742917353,0.002525038731676661,
0.0029365253223470676,0.0032661403715068065,0.0035826151872423457,0.003916626337287045,0.0043025787575814815,
0.00481401565348028,0.005921396686000432,0.002364590827360334,0.0026389232162923595,0.002855592702119905,
0.0030613834668636623,0.003276474466994089,0.003522644695641686,0.0038454685865182605,0.004533893590042749],
[0.0025672266787736156,0.0031980909444892804,0.003698576382497309,0.0041704505568082055,0.004653476113431633,
0.0051823447605420245,0.0058091667021883935,0.006656823034227595,0.008532099465993204,0.0023872752517825166,
0.002872682111864994,0.0032590014448035673,0.0036240565467501274,0.003998423747529171,0.004408973098197347,
0.004896260255356827,0.005556106208709167,0.007017828670240587,0.002284364303991858,0.002683632284553226,
0.003001055220896595,0.003300917551748293,0.003608460289737564,0.003945865808164509,0.004346635047784988,
0.004889992851317649,0.0060969521090462955,0.002218503429820102,0.002558225532599843,0.0028253460230545173,
0.003075561925905218,0.0033302671312246787,0.003607704544024131,0.003934832932856721,0.004374723998180398,
0.00533993213383286,0.0020847271648438016,0.002311633767453025,0.0024876942580310995,0.0026509603375305824,
0.0028156871399817435,0.0029936045076368475,0.0032015865400330874,0.0034785813597489243,0.004077684309311139],
[0.00227355667675018,0.002805236836620471,0.0032209116758492025,0.0036057083730374954,0.003989892160231512,
0.0043953996973899,0.004848079897839314,0.005391966947519379,0.00613527079964084,0.007796363594529018,
0.002120348223628812,0.0025292816269610312,0.002849968885186416,0.003147474253595381,0.0034450261101616874,
0.0037595685880610036,0.004111178627856744,0.004534161398886372,0.005112916878636409,0.006407798093046678,
0.002032835685961884,0.0023692581475593714,0.0026327859726142794,0.0028771706280849574,0.00312158927159844,
0.003380028387972353,0.003669066348227619,0.00401704963475816,0.004493780090372081,0.005563296418237351,
0.0019769691003886968,0.002263586712766765,0.002485770365355199,0.002690184421709366,0.0028932097241258156,
0.003106484759524572,0.003343476885060724,0.0036268807402428196,0.0040121676307676175,0.00486649155814749,
0.0018632300009208923,0.002054965563744719,0.0022017373363445885,0.0023354982935077867,0.002467260542173357,
0.0026046137205851167,0.0027560872050753347,0.0029357950123730468,0.0031779163834028913,0.003707486585455173]]
num_decks = 5
normalize = lambda x: x / x.sum()
optimal_review_rates = np.array([normalize(np.reshape(x, (y, num_decks))) for x, y in zip(optimal_review_rates, num_systems_set)])
# from ext_lqn_properties.ipynb
transition_thresholds = [0.015048737992973047,0.015797495306389367,0.01599484746534838,0.016079634446755746,0.016125055496048188,
0.016152717166862837,0.01617103990361111,0.016183926264181508,0.01619340146712257,0.016200613478280345]
num_items = 100
difficulty_of_item = np.random.exponential(1 / difficulty_rate, num_items)
ys = [[[run_sim(x, num_items, difficulty_rate, difficulty_of_item, review_rates, work_rate-x, sim_length) for _ in xrange(num_sim_repeats)] for x in arrival_rates] for y, z in zip(num_systems_set, optimal_review_rates)]
plt.xlabel(r'Arrival Rate $\lambda_{ext}$ (Items Per Second)')
plt.ylabel(r'Throughput $\lambda_{out}$ (Items Per Second)')
for ns, ds, th in zip(num_systems_set, ys, transition_thresholds):
plt.errorbar(
arrival_rates, [np.mean(y) for y in ds], yerr=[std_err(y) for y in ds],
label=r'Simulated ($n = %d$)' % ns)
plt.axvline(x=th, label=r'Predicted ($n = %d$)' % ns, linestyle='--')
plt.legend(loc='best')
plt.savefig(os.path.join('figures', 'lqn', 'throughput-vs-arrival-rate-vs-num-systems.pdf'))
plt.show()
with open(os.path.join('results', 'throughput-vs-arrival-rate-vs-num-systems.pkl'), 'wb') as f:
pickle.dump((arrival_rates, ys, num_systems_set, transition_thresholds), f, pickle.HIGHEST_PROTOCOL)
| nb/lqn_simulations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sumanEnv
# language: python
# name: sumanenv
# ---
# # Importing Packges
# + colab={} colab_type="code" executionInfo={"elapsed": 3586, "status": "ok", "timestamp": 1598629644701, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="mcR2OGs_kRno"
# %matplotlib inline
import numpy as np
import joblib
import pandas as pd
import matplotlib.pyplot as plt
import math
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
from sklearn.metrics import accuracy_score, f1_score, recall_score,precision_score, classification_report, confusion_matrix
import collections
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import precision_recall_curve, roc_curve
from sklearn.multiclass import OneVsRestClassifier
from sklearn.preprocessing import label_binarize
np.random.seed(1337) # for reproducibility
from sklearn.ensemble import VotingClassifier
# -
# # Loading data (CTTD features and Labels)
# + colab={} colab_type="code" executionInfo={"elapsed": 10269, "status": "ok", "timestamp": 1598629651412, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="dY-T9ZviojBz"
X_train = np.load('../data/train/X_train.npy')
Y_train = np.load('../data/train/Y_train.npy')
X_test = np.load('../data/test/set1/X_test.npy')
Y_test = np.load('../data/test/set1/Y_test.npy')
X_test2 = np.load('../data/test/set2/X_test2.npy')
Y_test2 = np.load('../data/test/set2/Y_test2.npy')
# -
# # Scaling features
# + colab={} colab_type="code" executionInfo={"elapsed": 775, "status": "ok", "timestamp": 1598629657354, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="4cJXuCsmpz8D"
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.fit_transform(X_test)
X_test2 = scaler.fit_transform(X_test2)
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" executionInfo={"elapsed": 1402, "status": "ok", "timestamp": 1598629663256, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="Ph-dUh8ZqEa9" outputId="7f878c8c-946d-4c82-8d1e-ac9a26edb5a1"
print(X_train.shape)
print(Y_train.shape)
print(X_test.shape)
print(Y_test.shape)
print(X_test2.shape)
print(Y_test2.shape)
# -
# # Loading models
# + colab={"base_uri": "https://localhost:8080/", "height": 105} colab_type="code" executionInfo={"elapsed": 1314, "status": "ok", "timestamp": 1598629960623, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="RW_ws22wqLi0" outputId="2b428f9b-31fb-463e-e4c0-5b896489a255"
SVM_clf = joblib.load('../models/svm_emg_clf.pkl')
LR_clf = joblib.load('../models/LR_emg_clf.pkl')
RF_clf = joblib.load('../models/RF_emg_clf.pkl')
ET_clf = joblib.load('../models/ET_emg_clf.pkl')
# + colab={} colab_type="code" executionInfo={"elapsed": 1378, "status": "ok", "timestamp": 1598630015761, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="FUpO8EImsfiX"
estimators=[('LR',LR_clf),('SVM',SVM_clf),('RF',RF_clf),('ET',ET_clf)]
# + colab={} colab_type="code" executionInfo={"elapsed": 1674, "status": "ok", "timestamp": 1598630041632, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="IzeYCKYyvSGC"
voting_clf=VotingClassifier(estimators,voting='soft')
# + colab={"base_uri": "https://localhost:8080/", "height": 510} colab_type="code" executionInfo={"elapsed": 204096, "status": "ok", "timestamp": 1598630270403, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="1ZGKeLj0viwy" outputId="016f5444-f0e0-4983-b8c2-33297f1c7beb"
voting_clf.fit(X_train,Y_train)
# + colab={} colab_type="code" executionInfo={"elapsed": 3309, "status": "ok", "timestamp": 1598630288279, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="_XQ3s4NQvluy"
Y_predict = voting_clf.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1223, "status": "ok", "timestamp": 1598630290667, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="qit8WlR1vtxI" outputId="ebd3c93e-00ab-43ac-987d-65270198b798"
accuracy_score(Y_test, Y_predict)
# -
# # SVC class don't compute probability by default, so we have to train the model by setting the parameter probability=True.
# + colab={} colab_type="code" executionInfo={"elapsed": 1876, "status": "ok", "timestamp": 1598630477437, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="aOIBSVovzIdT"
SVM_clf = SVC(kernel = 'rbf', C = 35.4, gamma= 'scale', class_weight = 'balanced',probability=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 15927, "status": "ok", "timestamp": 1598630494539, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="Pe9WoA-NzTVq" outputId="0f2e08c8-e731-4142-9cd7-5005c276e92b"
SVM_clf.fit(X_train, Y_train)
# -
# # Evaluation on Test set 1
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 5075, "status": "ok", "timestamp": 1598630574162, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="-_gcq3uXvu80" outputId="3c27be6c-f3bf-4738-fd17-d7eea3435a5c"
for clf in (LR_clf,SVM_clf,RF_clf,ET_clf,voting_clf):
y_predict=clf.predict(X_test)
print(clf.__class__.__name__,accuracy_score(Y_test, y_predict))
# -
# # Evaluation on Test set 2
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 3546, "status": "ok", "timestamp": 1598630634131, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11638296981034036456"}, "user_tz": -330} id="qZR_2eNxzzxJ" outputId="24b0445f-bd6f-4742-e5e1-345ae5f29442"
for clf in (LR_clf,SVM_clf,RF_clf,ET_clf,voting_clf):
y_predict=clf.predict(X_test2)
print(clf.__class__.__name__,accuracy_score(Y_test2, y_predict))
| notebooks/.ipynb_checkpoints/demo_voting_classifier-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import cross_validate
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.metrics.scorer import make_scorer
from sklearn.decomposition import PCA
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LassoCV
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import f_regression
from sklearn.linear_model import ElasticNetCV
from sklearn.linear_model import RidgeCV
from sklearn.model_selection import TimeSeriesSplit
from sklearn.model_selection import RandomizedSearchCV
from sklearn.linear_model import Lasso
from matplotlib import pyplot as plt
import seaborn as sns
import missingno as msno
import sys
sys.path.append('..')
from utils import preprocess, missing, evaluate
# -
# ### Global Variables and Load Data
target = 'SI.POV.DDAY'
predict_year=2010
#percent of input Indicators to use (set to 100 for full set of input features)
percent = 50
# +
#Load the data from disk
input_dir = '.\\..\\data\\'
data_input = "cleaned_data.pkl"
data = pd.read_pickle(input_dir + data_input)
#Possible subset of data choosen to reduce calulation time
#For percetages less than 100% we try to choose a subset that represents the spread of variables
if percent == 100:
pass
else:
num_indicators_original = data.shape[1]
step = int(100/percent)
data_new = data.iloc[:,::step].copy()
#Add the target column if not already included
if target not in data_new.columns:
data_new[target] = data[target]
data = data_new
print(data.shape[1], "indicators included")
# -
# ### Multicountry plot across the first 20 independent variables and the dependent variable (SI.POV.DDAY)
#
# The following plot is a quick illustration of the state of data in across multiple coutries.
#
# One thing that it clearly shows is that there is a strong correlation of location of missing values for many of the variables. However for some others, such as SI.POV.DDAY and SI.POV.GINI this is less the case.
#
# Secondly, it is clear that we will need a model that is robust to missing values.
#
# A key question here is if these patterns help us know if we have to model each country seperately or if one model fits all. Is there a method that can help us to work this out? Maybe there is a middle ground of number of models (for example a different model for every continent.
#
# There are a number of outlier countries that could be investigated as well. For example, in SE.PRM.UNER. What should be done with these? How should our respond to these outliers?
# +
#Create a list of coutries to be plotted in the summary plot below
countries = list(data.index.levels[0])
#pyplot.figure(figsize=(16, 20))
inputs = list(data.iloc[:,:20].columns)
fig, axes = plt.subplots(len(inputs), 1, sharex=True ,figsize=(16, 32))
for i in range(len(inputs)):
axes[i].set_xticks(list(range(1972,2018)), minor=True)
axes[i].set_title(inputs[i])
#ax.set_yticklabels([])
column = inputs[i]
for country in countries:
values = data.loc[country][column].values
axes[i].plot( list(range(1972,2019)), values)
plt.subplots_adjust(wspace=0, hspace=0.4)
#fig.invert_xaxis()
#pyplot.show()
# -
# ### Break data into windows
# Break up input dataframe (countries/year as 2-level index and economic indicators as columns) into windowed dataframes for training and testing.
#
# A windowed dataframe can be seen below. Each window row is a sliding window (of size 'lag') of the indictor data.
#
# %time data_regressors, data_targets = \
# preprocess.window_data(data, lag=3,num_windows=10, step=1, predict_year=2010, \
# target=target, impute_type='interpolation')
# +
#Break up into training and testing data.
idx = pd.IndexSlice
data_train_regressors = data_regressors.loc[idx[:,2:10],:]
data_train_targets = data_targets.loc[idx[:,2:10],:]
data_test_regressors = data_regressors.loc[idx[:,1],:]
data_test_targets= data_targets.loc[idx[:,1],:]
# -
# Widowed dataframe:
data_train_regressors.head(13)
# ### Postprocess windows
# Here we deal with windows that do not have any target value. Note that in the windowed dataframe, every row is a window. Each window will act as an observation in the input to our machine learning algorithm.
#
# In the case of the training data, we get rid of these windows (both in the regressor training dataframe and the target training dataframe).
#
# *TODO: Investigate possibility of imputing missing target data. One lead is [here](https://www.analyticbridge.datasciencecentral.com/forum/topics/missing-values-in-target)*
#
# In the case of the test data we also do the same, as without a target, it is impossible to evaluate the error in prediction.
#
# +
#For Training, only consider windows that don't have a missing target as they offer nothing to training
#Therefore, remove those observations from both the training regressors and targets datasets.
data_train_regressors_subset = data_train_regressors[~np.isnan(list(data_train_targets.values.flatten()))]
data_train_targets_subset = data_train_targets[~np.isnan(list(data_train_targets.values.flatten()))]
#For testing, also remove windows with no target variable as it is impossible to measure preformance.
data_test_regressors_subset = data_test_regressors[~np.isnan(list(data_test_targets.values.flatten()))]
data_test_targets_subset = data_test_targets[~np.isnan(list(data_test_targets.values.flatten()))]
# -
# ### Standardize
# +
scaler = StandardScaler()
#Fit the scaler on the training data only
scaler.fit(data_train_regressors_subset)
data_train_regressors_std_array = scaler.transform(data_train_regressors_subset)
data_test_regressors_std_array = scaler.transform(data_test_regressors_subset)
# -
data_train_regressors_std = pd.DataFrame(data_train_regressors_std_array, index=data_train_regressors_subset.index, columns=data_train_regressors_subset.columns)
data_test_regressors_std = pd.DataFrame(data_test_regressors_std_array, index=data_test_regressors_subset.index, columns=data_test_regressors_subset.columns)
# ### Linear Regression Prediction
# #### Historical values of the target only
#
# This is very much an naive predictor. If we are limited to only historical values of the target variable then there are well established time series forecasting methods that can be used such as ARIMA.
# +
model_target_only = LinearRegression()
model_target_only.fit(data_train_regressors_std.loc[:,target],data_train_targets_subset)
#Make predictions
predictions_target_only = model_target_only.predict(data_test_regressors_std.loc[:,target])
mse_target_only = mean_squared_error(data_test_targets_subset, predictions_target_only)
print("RMSE of linear regression using historical target values only is:", np.sqrt(mse_target_only))
# -
# #### OLS Linear Regression
#
# Here we use the linear regression function in scikit-learn. From an implementation point of view, this is just plain ordinary least squares (scipy.linalg.lstsq) wrapped as a scikit-learn predictor object.
# +
model_linear = LinearRegression()
model_linear.fit(data_train_regressors_std,data_train_targets_subset)
#Make predictions
predictions = model_linear.predict(data_test_regressors_std)
mse= mean_squared_error(data_test_targets, predictions)
print("RMSE of OLS linear regression using all target values only is:", np.sqrt(mse))
# -
# #### Ridge Regression
# +
scorer = make_scorer(mean_squared_error)
model_ridge = RidgeCV(scoring=scorer, cv=5)
model_ridge.fit(data_train_regressors_std,data_train_targets_subset)
#Make predictions
predictions = model_ridge.predict(data_test_regressors_std)
mse= mean_squared_error(data_test_targets, predictions)
print("RMSE of ridge regression using all target values only is:", np.sqrt(mse))
# -
# #### Lasso Regression
# The choice of Lasso for this problem is discussed in the associated blog. (see README.md for a link)
#
# It is evaluated here using the cross-validation Scikit-learn estimator LassoCV. As well as using cross-validation testing to evaluate the fit, it also picks the best value of alpha that gives best predictive performance.
#
# The default values for the range of alpha and the number of values to test set by eps,n_alphas were tested to be ideal.
#
# The cross validation technique here according to the textbooks is not appropiate for time-series in that there is data leakage as some of the training sets are from timestamps later in time that that the timestamps of the test sets. As noted below in the time-series cross validation section, this actually generalised beter accroding to our final test set.
#
# +
model_lasso = LassoCV(cv=5, eps=1e-3, selection='random', random_state=0)
model_lasso.fit(data_train_regressors_std.values,data_train_targets_subset.values.flatten())
#Make predictions
predictions = model_lasso.predict(data_test_regressors_std)
mse= mean_squared_error(data_test_targets.values.flatten(), predictions)
print("RMSE of lasso regression using all target values only is:", np.sqrt(mse))
# +
num_of_values_to_skip = 23 #We want to focus on the lower values of alpha for our graphs
mses = np.sqrt(np.mean(model_lasso.mse_path_[num_of_values_to_skip:], axis=1))
alphas = model_lasso.alphas_[num_of_values_to_skip:]
sns.set()
ax = sns.lineplot(alphas, mses)
ax.set_title("RMSE for different values of Alpha in Lasso Cross-Validation")
ax.axvline(alphas[np.argmin(mses)], color='green', linewidth=1)
ax.set(xlabel='alpha value', ylabel='RMSE')
# -
#Convert to dataframe with country names as index
predictions_df = pd.DataFrame(predictions.flatten(), index=data_test_regressors_subset.index, columns=[target])
#Reindex in order to put back in the coutries that were removed due to lack of target.
predictions_df = predictions_df.reindex(data_test_targets.index)
(abs(predictions_df - data_test_targets)).sort_values(by='SI.POV.DDAY',ascending=False).head(10)
# The biggest errors were on countries that did not have a value measured for SI.POV.DDAY for a number of years. On the surface these could be considered outliers but instead, they are infact probably our most important coutries to predict poverty as accuractely as we can for.
#
# IT could be a sensible approch to focus on these countries and relook at the problem at hand for them specifically. What extra features could we add to improve their performance? Is our interpolation method serving these countries well?
# #### Lars
from sklearn.linear_model import LassoLarsCV
# +
model_lars = LassoLarsCV(cv=5, max_iter=100)
model_lars.fit(data_train_regressors_std.values,data_train_targets_subset.values.flatten())
#Make predictions
predictions = model_lars.predict(data_test_regressors_std)
mse= mean_squared_error(data_test_targets.values.flatten(), predictions)
print("RMSE of lars regression using all target values only is:", np.sqrt(mse))
# -
# #### ElasticNet
# ElasticNet is a method that has a tuning parameter than on one extreme is identical to Lasso and at the other it's identical to Ridge.
# This brought nothing to the table as our optimal solution is 100% Lasso.
# +
model_elastic = ElasticNetCV(l1_ratio=.1,cv=5, random_state=0)
model_elastic.fit(data_train_regressors_std.values,data_train_targets_subset.values.flatten())
#Make predictions
predictions = model_elastic.predict(data_test_regressors_std)
mse= mean_squared_error(data_test_targets.values.flatten(), predictions)
print("RMSE of ElasticNet regression using all target values only is:", np.sqrt(mse))
# -
# ### Time Series Split
X = data_train_regressors_std.copy()
y = data_train_targets_subset.copy()
# +
#We first need to sort the the rows in our test set as to have the rows in order of..
#data rather than country
#Before we do the sorting lets combine the two datasets so that we can sort both together
Xy = X
Xy.loc[:,('temp', '1')] = y.values
#Sort according to the window number
Xy = Xy.sort_index(level=1, ascending = False)
#This will be used to obtain CV splits for the following algos
tscv = TimeSeriesSplit(n_splits=5)
#Reconstruct the X and y datasets from the combined one
y = Xy.loc[:,('temp','1')].values
X = Xy.drop('temp', level=0, axis=1).values
# -
# #### Lasso with Time-series splits
# Surpisingly, the time-series cross validation does not perofrm as well on the test set as the standard cross-validation used above.
# This probably warrants some more time to investigate if there is a bug somewhere in the preparation of the data.
# +
model_lasso = LassoCV(cv=tscv.split(Xy), selection='random', random_state=0)
model_lasso.fit(X,y)
#Make predictions
predictions = model_lasso.predict(data_test_regressors_std)
mse= mean_squared_error(data_test_targets.values.flatten(), predictions)
print("RMSE of time-series CV lasso regression using all target values only is:", np.sqrt(mse))
| notebooks/linear_regression_predictor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib as mtl
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.express as px
import plotly.graph_objects as go
telco_df = pd.read_csv("WA_Fn-UseC_-Telco-Customer-Churn.csv")
telco_df.tail(10)
telco_df.info()
# +
#since Total Chrges is of type Object, it should be of type float to allow calculations as it's numeric
telco_df['TotalCharges'] = pd.to_numeric(telco_df['TotalCharges'], errors='coerce')
telco_df.info()
# -
telco_df.isnull().sum()
telco_df.describe(include = "all")
#removing the customerID from the dataset as it has no contribution in Analysis
telco_df = telco_df.iloc[:, 1:]
telco_df
# ### Demographics Analysis
# +
#Overall number of Churn between the gender
pd.crosstab(telco_df['gender'],telco_df['Churn'])
# +
#Use the demographics of genders with those with partners and dependednts
telco_df.groupby(['gender','Partner','Dependents','Churn']).size().reset_index(name='Count')
# -
# 1a) The above analysis shows that there isn't much of a difference between the Gender however those who have no partners and Dependents are more likely to Churn than those who have Partners and or dependents.
# 2b) if the market team wants to increase their customer retention then they should targets those clients who have no partners and or dependents
# ### Services Analysis
#convert the label column in to numeric from categorical
telco_df['Churn'].replace(to_replace='Yes', value=1, inplace=True)
telco_df['Churn'].replace(to_replace='No', value=0, inplace=True)
#create_dummies to converting all of the categorical features columns data into numerical data
telco_dummies = pd.get_dummies(telco_df)
telco_dummies.head(10)
telco_dummies.info()
# +
#plot the graph to show the crrelation between the features and the label variables;
monthly_correlation = telco_dummies.corr()['MonthlyCharges'].sort_values(ascending=False)
fig = go.Figure(go.Bar(x=monthly_correlation.keys(),
y=monthly_correlation,
name='Churn Correlation',
marker={'color': monthly_correlation, 'colorscale': 'Viridis'}))
fig.update_layout(yaxis={'categoryorder':'category ascending'})
fig.update_layout(
title='Correlation to Monthly Charges',
xaxis_tickfont_size=10,
xaxis_tickangle=-45,
yaxis=dict(
title='Correlation',
titlefont_size=16,
tickfont_size=14,
),
bargap=0.15, # gap between bars of adjacent location coordinates.
)
fig.show()
# -
# 2a) From the above figure, we can see that the 3 services that contributes greatly to monthly charges are;
# Fiber Optic internet, Tv and Movies streaming services
# +
#finding the type of contract that would encourage customer retention if the company offered only Phone services
fig = px.bar(telco_df, x="Contract", color ="PhoneService", barmode="group", facet_col="Churn")
fig.show()
# -
# 2b) Based on the above visual, it shows that if the company was offering only Phone Services then the contract type that would encourage retention would be the Month to Month contract
# ### Payments Analysis:
# +
#plot the graph to show the correlation between the features and the label variables;
churn_correlation = telco_dummies.corr()['Churn'].sort_values(ascending=False)
fig = go.Figure(go.Bar(x=churn_correlation.keys(),
y=churn_correlation,
name='Churn Correlation',
marker={'color': churn_correlation, 'colorscale': 'Viridis'}))
fig.update_layout(yaxis={'categoryorder':'category ascending'})
fig.update_layout(
title='Correlation to Churn',
xaxis_tickfont_size=10,
xaxis_tickangle=-45,
yaxis=dict(
title='Correlation',
titlefont_size=16,
tickfont_size=14,
),
bargap=0.15, # gap between bars of adjacent location coordinates.
)
fig.show()
# -
# 3b) Based on the above graph, the compant should consider other payments options such as automatic Bank transfer, automatic credit cards and even Mailed check as those payments methods have higher retention than the paperless method
| ADS-Assignment-4-main/.ipynb_checkpoints/Jay_Assignment4-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prepare CSJ (japanese dataset)
# 1. grab linguistic units from dataset
# 2. grab dataset statistics
import numpy as np
from glob import glob
from tqdm.autonotebook import tqdm
from parallelspaper.speech_datasets import prep_CSJ
import pandas as pd
from parallelspaper.config.paths import DATA_DIR
CSJ_DIR = '/mnt/cube/Datasets/Japanese/XML/BaseXML/core/'
xml_locs = glob(CSJ_DIR+'*.xml')
# ### Load data
(words, pos, mora, phonemes, phones, phone_class, session_lens,
IPU_lens, phone_lens, word_lens, session_lens, IPU_phonemes) = prep_CSJ(xml_locs)
# ### Get dataset statistics
num_phonemes = len(np.concatenate(phonemes))
num_words = len(np.concatenate(words))
word_durations_s = np.nan
word_length_phones = word_lens
phone_duration_s = phone_lens
unique_phones = len(np.unique(np.concatenate(phonemes)))
unique_words = len(np.unique(np.concatenate(words)))
utterance_length_phones = [len(i) for i in np.concatenate(IPU_phonemes)]
n_sessions = len(phones)
session_durations = [np.sum(i) for i in session_lens]
total_duration = np.sum(IPU_lens)
stats_df = pd.DataFrame([[
num_phonemes,
num_words,
word_durations_s,
word_length_phones,
phone_duration_s,
unique_phones,
unique_words,
utterance_length_phones,
n_sessions,
session_durations,
total_duration
]],
columns=[
'num_phonemes',
'num_words',
'word_durations_s',
'word_length_phones',
'phone_duration_s',
'unique_phones',
'unique_words',
'utterance_length_phones',
'n_sessions',
'session_durations',
'total_duration'
])
stats_df
# statistics for this language
stats_df.to_pickle((DATA_DIR / 'stats_df/CSJ_stats_df.pickle'))
# ### make sequence dataframes
# +
# words, pos, mora, phonemes, phones, phone_class
# -
seq_df = pd.DataFrame(columns = ['language', 'levels', 'data'])
seq_df.loc[len(seq_df)] = ['japanese', 'speaker/IPU/phonemes', IPU_phonemes]
seq_df.loc[len(seq_df)] = ['japanese', 'speaker/word', words]
seq_df.loc[len(seq_df)] = ['japanese', 'speaker/pos', pos]
seq_df.loc[len(seq_df)] = ['japanese', 'speaker/word/mora', mora]
seq_df.loc[len(seq_df)] = ['japanese', 'speaker/word/phonemes', phonemes]
seq_df.loc[len(seq_df)] = ['japanese', 'speaker/word/phones', phones]
seq_df.loc[len(seq_df)] = ['japanese', 'speaker/word/phone_class', phone_class]
seq_df.to_pickle((DATA_DIR / 'speech_seq_df/CSJ_seq_df.pickle'))
| notebooks/language/02.0-CSJ-prep-dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ##ThinkDSP
#
# This notebook contains solutions to exercises in Chapter 5: Autocorrelation
#
# Copyright 2015 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](http://creativecommons.org/licenses/by/4.0/)
# +
from __future__ import print_function, division
import thinkdsp
import thinkplot
import thinkstats2
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# -
# **Exercise:** If you did the exercises in the previous chapter, you downloaded
# the historical price of BitCoins and estimated the power spectrum
# of the price changes. Using the same data, compute the autocorrelation
# of BitCoin prices. Does the autocorrelation function drop off quickly? Is there evidence of periodic behavior?
df = pd.read_csv('coindesk-bpi-USD-close.csv', nrows=1625, parse_dates=[0])
ys = df.Close.values
wave = thinkdsp.Wave(ys, framerate=1)
wave.plot()
thinkplot.config(xlabel='Time (days)',
ylabel='Price of BitCoin ($)')
# Here's the autocorrelation function using the statistical definition, which unbiases, normalizes, and standardizes; that is, it shifts the mean to zero, divides through by standard deviation, and divides the sum by N.
# +
from autocorr import autocorr
lags, corrs = autocorr(wave)
thinkplot.plot(lags, corrs)
thinkplot.config(xlabel='Lag',
ylabel='Correlation')
# -
# The ACF drops off slowly as lag increases, suggesting some kind of pink noise. And it looks like there are moderate correlations with lags near 200, 425 and 700 days.
# We can compare my implementation of `autocorr` with `np.correlate`, which uses the definition of correlation used in signal processing. It doesn't unbias, normalize, or standardize the wave.
N = len(wave)
corrs2 = np.correlate(wave.ys, wave.ys, mode='same')
lags = np.arange(-N//2, N//2)
thinkplot.plot(lags, corrs2)
thinkplot.config(xlabel='Lag',
ylabel='Dot product')
# The second half of the result corresponds to positive lags:
N = len(corrs2)
half = corrs2[N//2:]
thinkplot.plot(half)
thinkplot.config(xlabel='Lag',
ylabel='Dot product')
# We can standardize the results after the fact by dividing through by `lengths`:
lengths = range(N, N//2, -1)
half /= lengths
half /= half[0]
thinkplot.plot(half)
thinkplot.config(xlabel='Lag',
ylabel='Dot product')
# But even after standardizing, the results look very different. In the results from `correlate`, the peak at lag 200 is less apparent, and the other two peaks are obliterated.
thinkplot.preplot(2)
thinkplot.plot(corrs, label='autocorr')
thinkplot.plot(half, label='correlate')
thinkplot.config(xlabel='Lag', ylabel='Correlation')
# I think the reason the results are so different the data look very different in different parts of the range; in particular, the variance changes a lot over time.
#
# For this dataset, the statistical definition of ACF, is probably more appropriate.
# **Exercise:** The example code in `chap05.ipynb` shows how to use autocorrelation
# to estimate the fundamental frequency of a periodic signal.
# Encapsulate this code in a function called `estimate_fundamental`,
# and use it to track the pitch of a recorded sound.
#
# To see how well it works, try superimposing your pitch estimates on a
# spectrogram of the recording.
wave = thinkdsp.read_wave('28042__bcjordan__voicedownbew.wav')
wave.normalize()
wave.make_audio()
# I'll use the same example from `chap05.ipynb`. Here's the spectrogram:
wave.make_spectrogram(2048).plot(high=4200)
thinkplot.config(xlabel='Time (s)',
ylabel='Frequency (Hz)',
xlim=[0, 1.4],
ylim=[0, 4200])
# And here's a function that encapsulates the code from Chapter 5. In general, finding the first, highest peak in the autocorrelation function is tricky. I kept it simple by specifying the range of lags to search.
def estimate_fundamental(segment, low=70, high=150):
lags, corrs = autocorr(segment)
lag = np.array(corrs[low:high]).argmax() + low
period = lag / segment.framerate
frequency = 1 / period
return frequency
# Here's an example of how it works.
duration = 0.01
segment = wave.segment(start=0.2, duration=duration)
freq = estimate_fundamental(segment)
freq
# And here's a loop that tracks pitch over the sample.
#
# The `ts` are the mid-points of each segment.
# +
step = 0.05
starts = np.arange(0.0, 1.4, step)
ts = []
freqs = []
for start in starts:
ts.append(start + step/2)
segment = wave.segment(start=start, duration=duration)
freq = estimate_fundamental(segment)
freqs.append(freq)
# -
# Here's the pitch-tracking curve superimposed on the spectrogram:
wave.make_spectrogram(2048).plot(high=900)
thinkplot.plot(ts, freqs, color='green')
thinkplot.config(xlabel='Time (s)',
ylabel='Frequency (Hz)',
xlim=[0, 1.4],
ylim=[0, 900])
# Looks pretty good!
| ThinkDSP-master/code/chap05soln.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Murtaza-Husain1/Super-Saiyan-Classifier/blob/master/Super_Saiyan_Classifier.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="fxPWoZ8JgfzO" colab_type="code" outputId="770ae7e5-a575-4351-da5c-c454cd8a4080" colab={"base_uri": "https://localhost:8080/", "height": 158}
# !git clone https://github.com/Murtaza-Husain1/Super-Saiyan-Classifier.git
# + id="QQvHiydP3kyi" colab_type="code" colab={}
# !git pull
# + id="7G7EQbCHvIGg" colab_type="code" outputId="fbc3207d-9bbd-4e8a-dd6b-8df8456ebf1a" colab={"base_uri": "https://localhost:8080/", "height": 34}
# # !git add *
# # !git commit -m ""
# # !git push
# + id="gTwjfcWo4Haq" colab_type="code" outputId="08aa9c0b-013f-48f4-8d7f-37ef832f7c44" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd /content/Super-Saiyan-Classifier
# # cd ..
# + id="1m94GUajRHQY" colab_type="code" colab={}
# + id="a2tvSO1NqWcQ" colab_type="code" outputId="0eff575e-cf6b-423b-faf0-54f57e3724d1" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls
# + [markdown] id="Nh5k3PVDWNs-" colab_type="text"
# import * for ease of experimentation
# + id="EsBiq5ieWEWW" colab_type="code" colab={}
from fastai.vision import *
# + [markdown] id="IuzizefaaWlo" colab_type="text"
# # File Handling
# + [markdown] id="Rq5tf0ylW3Ns" colab_type="text"
# Concatenate my text in all my .csv's (faster to do manually though)
# + id="EtRZ8Cc7WfJc" colab_type="code" colab={}
import os
import glob
import pandas as pd
# + id="Xej8_2zFXUtP" colab_type="code" colab={}
os.chdir(path)
# + id="Im4KF8jQXNly" colab_type="code" colab={}
extension = 'csv'
all_filenames = [i for i in glob.glob('*.{}'.format(extension))]
# + id="kY6TCpYrsoxN" colab_type="code" colab={}
folder = 'ss4'
f = 'ss4.csv'
# + id="dQzsfrCisvy3" colab_type="code" colab={}
path = Path('')
dest = path/folder
# + id="fHC6k3ggup6J" colab_type="code" colab={}
path = Path('')
# + id="wAnXR1zstRrR" colab_type="code" outputId="4b0a66da-f3b2-4ea2-e0c9-f22cdbff5e54" colab={"base_uri": "https://localhost:8080/", "height": 263}
path.ls()
# + id="_5TZK62QuJnG" colab_type="code" colab={}
download_images(path/f, dest, max_pics=200)
# + id="R_pdEwLXM0P6" colab_type="code" colab={}
# !zip -r file.zip ss4
# + id="NiF-uQNVwyQK" colab_type="code" colab={}
classes = ['ss1', 'ss2', 'ss3', 'ss4']
# + id="j-hdw4_cTG9O" colab_type="code" colab={}
for c in classes:
print(c)
verify_images(path/c, delete=True, max_size=800)
# + [markdown] id="ckV6p28vavUI" colab_type="text"
# # View Data
# + id="qhpMT84ya0ge" colab_type="code" colab={}
np.random.seed(1440)
data = ImageDataBunch.from_folder(path, train=".", valid_pct=.3,
ds_tfms=get_transforms(), size=224, num_workers=4).normalize(imagenet_stats)
# + id="WzxAVTdQbxd7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="035e066e-03d0-4b23-d754-841f71d8084e"
data.classes
# + id="yVhTxc5Ab1m3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 568} outputId="8d2163ff-cb42-4e73-97e5-1d06d43b5607"
data.show_batch(rows=3, figsize=(7,8))
# + id="J8Aeo9UocEZ-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0620bd71-31af-479c-b96d-4b530692d857"
data.classes, data.c, len(data.train_ds), len(data.valid_ds)
# + [markdown] id="UN4PtHjVd8cL" colab_type="text"
# # Train Model
# + colab_type="code" id="QkQtjkS7gP-L" colab={}
np.random.seed(1440)
data = ImageDataBunch.from_folder(path, train=".", valid_pct=.3,
ds_tfms=get_transforms(), size=224, num_workers=4).normalize(imagenet_stats)
# + id="sVQ8kszWcVkg" colab_type="code" colab={}
learn = cnn_learner(data, models.resnet34, metrics=error_rate)
# + id="biAkqwemc66O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 639} outputId="d32d5fd3-e680-424b-b4fe-b7609b300129"
learn.fit_one_cycle(20) # valid_pct=.5, size=224, resnet50
# + id="ueSg2fz7eX5O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="0e5ad9ef-7d76-461f-8353-c05f66e9b2b3"
learn.fit_one_cycle(5) # valid_pct=.5, size=276, resnet50
# + id="UQQ9IFUYftVi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="afd6b0f0-4f59-45eb-b339-a774ce067047"
learn.fit_one_cycle(5) # valid_pct=.3, size=224, resnet50
# + id="MX-gH9q8ilcz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 77} outputId="b1a807d0-825c-4695-a3e3-d6bc9f9cde08"
learn.fit_one_cycle(1) # valid_pct=.3, size=224, resnet34
# + id="T_b4pJwugvuC" colab_type="code" colab={}
learn.unfreeze()
# + id="GzvVFhNQgyIS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="11c105cb-020a-40c2-bb69-f06b3b445092"
learn.lr_find(start_lr=1e-5, end_lr=1e-1)
# + id="qHZsjcJlg-hw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="5907131f-b41e-42f1-abcb-21a17f68bb71"
learn.recorder.plot()
# + id="KOFVF-_EiCiP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 106} outputId="17a97552-5fa7-47ec-c4f1-5145c65f856d"
learn.fit_one_cycle(2, max_lr=slice(3e-4,3e-3))
# + id="oQdwVtmalBY5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 432} outputId="c2ce5a9a-93dc-40a8-fd24-6deda1685217"
learn.fit_one_cycle(13)
# + id="KYdadFZWmeRk" colab_type="code" colab={}
learn.save('stage-1')
# + id="Lz6r_8adnoXO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 935} outputId="e6501dfd-c144-4ee0-fb3a-1b24302f7242"
learn.fit_one_cycle(30)
# + [markdown] id="FYMbSHY3pyhv" colab_type="text"
# # Analysis
# + id="y14s2bZwp5OK" colab_type="code" colab={}
learn.load('stage-1')
# + id="xtaAJBH7qFia" colab_type="code" colab={}
interp = ClassificationInterpretation.from_learner(learn)
# + id="DZR6TsRiqJm4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 311} outputId="b2f20a10-e875-4217-c6e9-9d4f96324e21"
interp.plot_confusion_matrix()
# + [markdown] id="UlOwJvIervQq" colab_type="text"
# The model is fairly accurate for ss2, ss3, and ss4, but ss1 is a bit all over the place. It's possible that the training data looks too similar to ss2. I might try to get more data for ss1. 80% is okay but we can do better.
| Super_Saiyan_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bilinear Interpolation
import cv2
from matplotlib import pyplot as plot
import numpy as nm
img =cv2.imread("apple.jpg")[:,:,::-1]
plot.imshow(img)
plot.title("Original Image")
# #### Dimension Of Orignal Image
print(img.shape)
# ### Applying Bilinear Interpolation
bilinear_img = cv2.resize(img,None, fx = 10, fy = 10, interpolation = cv2.INTER_LINEAR)
plot.imshow(bilinear_img)
plot.title("Bilinear Interpolation Image")
# # Dimension after applying Bilinear- interpolation
print(bilinear_img.shape)
| ImageResize/Bilinear_Interpolation/.ipynb_checkpoints/bilinear_interpolation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="5lx-7K2fzK8G" outputId="7808a03a-648c-40b6-cf7d-f8d505b12d46"
# !pip install torchtext==0.6.0
# + id="Qx54Pe-X-Fcd"
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchtext
from torchtext import data
import time
import torch.optim as optim
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import classification_report,confusion_matrix
# + id="kdu-8gPjWQzv"
SEED = 1234
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
# + id="uONq22WixBVd"
def generate_bigrams(x):
n_grams = set(zip(*[x[i:] for i in range(2)]))
for n_gram in n_grams:
x.append(" ".join(n_gram))
return x
# + id="2VHQUKCZA9it"
TEXT = data.Field(tokenize="spacy",preprocessing=generate_bigrams,lower=True)
LABEL = data.LabelField()
fields = [(None,None),('text', TEXT),('label',LABEL), (None,None)]
# + id="XE2EK7hpTMcb"
train_data, test_data = data.TabularDataset.splits(
path = '/content/drive/MyDrive/data/benchmarking_data',
train = 'train.csv',
test = 'valid.csv',
format = 'csv',
fields = fields,
skip_header = True
)
# + colab={"base_uri": "https://localhost:8080/"} id="ztPDjhrlTfwx" outputId="7cc0506c-0f5d-43e8-b267-677cb7efc3a5"
print(vars(train_data[2]))
# + colab={"base_uri": "https://localhost:8080/"} id="re2to_BKVIxy" outputId="e1d3f176-ba5a-4682-d1d6-67dbd6136df9"
print(f"Number of training examples: {len(train_data)}")
print(f"Number of validation examples: {len(test_data)}")
# + [markdown] id="CgY9E--DWCsa"
# #### Create our own validation set
#
# + id="Z9FUieVgV-Sw"
import random
train_data,valid_data = train_data.split(random_state=random.seed(SEED))
# + colab={"base_uri": "https://localhost:8080/"} id="JcqbvM5jWdbQ" outputId="543ab8ba-316b-4383-9ea8-070f8b6d18a9"
print(f"Number of training examples: {len(train_data)}")
print(f"Number of validation examples: {len(valid_data)}")
print(f"Number of test examples: {len(test_data)}")
# + colab={"base_uri": "https://localhost:8080/"} id="s8ZJ_HD66Pyt" outputId="e14a3523-63c2-477a-8c04-f3cbfa1f4096"
MAX_VOCAB_SIZE = 25_000
TEXT.build_vocab(train_data,max_size=MAX_VOCAB_SIZE,
vectors="glove.6B.100d",
unk_init = torch.Tensor.normal_)
LABEL.build_vocab(train_data)
# + colab={"base_uri": "https://localhost:8080/"} id="hR7Le3b-eG1y" outputId="ae57d751-af14-43d7-ea72-0b71d3b65a58"
print(f"Unique tokens in text vocab:{len(TEXT.vocab)}")
print(f"Unique tokens in label vocab:{len(LABEL.vocab)}")
# + colab={"base_uri": "https://localhost:8080/"} id="WU2jTVCCj1vH" outputId="5640e7e7-93dd-4fbe-ad14-e13e23ab89a3"
print(TEXT.vocab.freqs.most_common(20))
# + colab={"base_uri": "https://localhost:8080/"} id="ZOj8KkfHk2pJ" outputId="00cdcf1c-b460-4a28-9b33-6b0d1ad5ebf9"
print(TEXT.vocab.itos[:10])
# + colab={"base_uri": "https://localhost:8080/"} id="I_1g38BdlAvI" outputId="70cc33d9-0e11-424f-9437-d0dcf242e95a"
print(LABEL.vocab.stoi)
# + id="Nxr1RSdalGgQ"
BATCH_SIZE = 64
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
train_iterator,valid_iterator,test_iterator = data.BucketIterator.splits(
(train_data,valid_data,test_data),
batch_size=BATCH_SIZE,
device=device,
sort_within_batch=True,
sort_key=lambda x:len(x.text))
# + id="_HoO2nQezb-Q"
class FastText(nn.Module):
def __init__(self,vocab_size,embedding_dim,output_dim,pad_idx):
super().__init__()
self.embedding = nn.Embedding(vocab_size,embedding_dim)
self.fc = nn.Linear(embedding_dim,output_dim)
def forward(self,text):
#text = [sent_len,batch_size]
embedded = self.embedding(text)
#embedded = [sent_len,batch_size,emb_dim]
embedded = embedded.permute(1,0,2)
#embedded = [batch_size,sent_len,emb_dim]
pooled = F.avg_pool2d(embedded,(embedded.shape[1],1)).squeeze(1)
#pooled = [batch_size,sent_len]
return self.fc(pooled)
# + id="IXX4NCulndAw"
INPUT_DIM = len(TEXT.vocab)
EMBEDDING_DIM = 100
HIDDEN_DIM = 256
OUTPUT_DIM = len(LABEL.vocab)
model = FastText(INPUT_DIM,EMBEDDING_DIM,HIDDEN_DIM,OUTPUT_DIM)
# + colab={"base_uri": "https://localhost:8080/"} id="O35bR5pin2aG" outputId="aa66a919-2a8d-4df3-a1bb-a6a8799ff10f"
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
# + id="ek4TDiegoRgX"
import torch.optim as optim
lr=1e-3
optimizer = optim.Adam(model.parameters(),lr=lr)
#sched = optim.lr_scheduler
# + id="YV4zn_OKo92l"
criterion = nn.CrossEntropyLoss()
# + id="lt0iqS9CpBwL"
model = model.to(device)
criterion = criterion.to(device)
# + id="xSVvk8h6pJaR"
def categorical_accuracy(preds, y):
"""
Returns accuracy per batch, i.e. if you get 8/10 right, this returns 0.8, NOT 8
"""
max_preds = preds.argmax(dim = 1, keepdim = True) # get the index of the max probability
correct = max_preds.squeeze(1).eq(y)
return correct.sum() / torch.FloatTensor([y.shape[0]])
# + id="7ylzhmb6pUCT"
def train(model, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
optimizer.zero_grad()
predictions = model(batch.text)
loss = criterion(predictions,batch.label)
acc = categorical_accuracy(predictions, batch.label)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + id="WfcWAhG4pbvj"
def evaluate(model, iterator, criterion):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
predictions = model(batch.text)
loss = criterion(predictions,batch.label)
acc = categorical_accuracy(predictions,batch.label)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + id="IE0RB-BSphO0"
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# + colab={"base_uri": "https://localhost:8080/"} id="lsoc4vNApkgB" outputId="366de94d-9749-4f52-cc1e-5fffe10890f2"
N_EPOCHS = 5
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), '/content/drive/MyDrive/Models/INTENT/rnn-model.pt')
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
# + colab={"base_uri": "https://localhost:8080/"} id="EtiehDXT4pWV" outputId="4302516a-8edf-44e9-8cf5-c7c308eb94fb"
model.load_state_dict(torch.load("/content/drive/MyDrive/Models/INTENT/bag-of-tricks-model.pt"))
test_loss, test_acc = evaluate(model,test_iterator,criterion)
print(f"Test Loss: {test_loss:.3f} | Test Accuracy: {test_acc:.2f}")
# + id="RVpri4_zrBpk"
def plot_confusion_matrix(test_y, predict_y):
C = confusion_matrix(test_y, predict_y)
# C = 9,9 matrix, each cell (i,j) represents number of points of class i are predicted class j
A =(((C.T)/(C.sum(axis=1))).T)
#divid each element of the confusion matrix with the sum of elements in that column
# C = [[1, 2],
# [3, 4]]
# C.T = [[1, 3],
# [2, 4]]
# C.sum(axis = 1) axis=0 corresonds to columns and axis=1 corresponds to rows in two diamensional array
# C.sum(axix =1) = [[3, 7]]
# ((C.T)/(C.sum(axis=1))) = [[1/3, 3/7]
# [2/3, 4/7]]
# ((C.T)/(C.sum(axis=1))).T = [[1/3, 2/3]
# [3/7, 4/7]]
# sum of row elements = 1
B =(C/C.sum(axis=0))
#divid each element of the confusion matrix with the sum of elements in that row
# C = [[1, 2],
# [3, 4]]
# C.sum(axis = 0) axis=0 corresonds to columns and axis=1 corresponds to rows in two diamensional array
# C.sum(axix =0) = [[4, 6]]
# (C/C.sum(axis=0)) = [[1/4, 2/6],
# [3/4, 4/6]]
labels = [1,2,3,4,5,6,7,8,9]
# representing A in heatmap format
print("-"*20, "Confusion matrix", "-"*20)
plt.figure(figsize=(20,7))
sns.heatmap(C, annot=True, cmap="YlGnBu", fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.show()
print("-"*20, "Precision matrix (Columm Sum=1)", "-"*20)
plt.figure(figsize=(20,7))
sns.heatmap(B, annot=True, cmap="YlGnBu", fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.show()
# representing B in heatmap format
print("-"*20, "Recall matrix (Row sum=1)", "-"*20)
plt.figure(figsize=(20,7))
sns.heatmap(A, annot=True, cmap="YlGnBu", fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.show()
B = (C/C.sum(axis=0))
labels = [1,2,3,4,5,6,7,8,9]
# + id="bHKqlcp7dZbG"
def get_predictions(model,iterator):
y_pred = []
y_true = []
model.eval()
with torch.no_grad():
for batch in iterator:
text = batch.text
predictions = model(text)
y_pred.extend(torch.argmax(predictions,axis=-1).tolist())
y_true.extend(batch.label.tolist())
return y_pred,y_true
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="PDlxQa3zCfoJ" outputId="f9dc85bb-10ac-41c0-c009-0cdc8b94df96"
y_pred,y_true = get_predictions(model,test_iterator)
plot_confusion_matrix(y_true,y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="uAZxWrFFCrGa" outputId="db6fd7be-5fb1-41d7-e00f-9df4afb87d7c"
print('Classification Report:')
print(classification_report(y_true, y_pred))
# + id="5qJBeOrMD391"
| intent-recogntion/nbs/Bag of Tricks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME>:
# We are in a competition to win the archery contest in Sherwood. With our bow and arrows we shoot on a target and try to hit as close as possible to the center.
#
# The center of the target is represented by the values (0, 0) on the coordinate axes.
#
# 
#
# ## Goals:
# * data structures: lists, sets, tuples
# * logical operators: if-elif-else
# * loop: while/for
# * minimum (optional sorting)
#
# ## Description:
# In the 2-dimensional space, a point can be defined by a pair of values that correspond to the horizontal coordinate (x) and the vertical coordinate (y). The space can be divided into 4 zones (quadrants): Q1, Q2, Q3, Q4. Whose single point of union is the point (0, 0).
#
# If a point is in Q1 both its x coordinate and the y are positive. I leave a link to wikipedia to familiarize yourself with these quadrants.
#
# https://en.wikipedia.org/wiki/Cartesian_coordinate_system
#
# https://en.wikipedia.org/wiki/Euclidean_distance
#
# ## Shots
# ```
# points = [(4, 5), (-0, 2), (4, 7), (1, -3), (3, -2), (4, 5),
# (3, 2), (5, 7), (-5, 7), (2, 2), (-4, 5), (0, -2),
# (-4, 7), (-1, 3), (-3, 2), (-4, -5), (-3, 2),
# (5, 7), (5, 7), (2, 2), (9, 9), (-8, -9)]
# ```
#
# ## Tasks
# 1. <NAME> is famous for hitting an arrow with another arrow. Did you get it?
# 2. Calculate how many arrows have fallen in each quadrant.
# 3. Find the point closest to the center. Calculate its distance to the center.
# 4. If the target has a radius of 9, calculate the number of arrows that must be picked up in the forest.
# +
# Variables
points = [(4, 5), (-0, 2), (4, 7), (1, -3), (3, -2), (4, 5),
(3, 2), (5, 7), (-5, 7), (2, 2), (-4, 5), (0, -2),
(-4, 7), (-1, 3), (-3, 2), (-4, -5), (-3, 2),
(5, 7), (5, 7), (2, 2), (9, 9), (-8, -9)]
# -
# 1. <NAME> is famous for hitting an arrow with another arrow. Did you get it?
if len(points) > len(set(points)):
print("We did hit an arrow with another arrow!")
else:
print("We did not hit an arrow with another arrow.")
# +
# 2. Calculate how many arrows have fallen in each quadrant.
"""
Q1 positive x, positive y
Q2 positive x, negative y
Q3 negative x, negative y
Q4 negative x, positive y
"""
q1 = 0
q2 = 0
q3 = 0
q4 = 0
x_values = []
y_values = []
list_points = []
for i in list(points):
list_points.append(list(i))
#print(list_points)
for i in list_points:
x_values.append(i[0])
y_values.append(i[1])
#print(x_values)
#print(y_values)
for i in range(len(points)):
#print(x_values[i])
#print(y_values[i])
if x_values[i] >= 0 and y_values[i] >= 0:
q1 += 1
elif x_values[i] >= 0 and y_values[i] < 0:
q2 += 1
elif x_values[i] < 0 and y_values[i] >= 0:
q4 += 1
else:
q3 += 1
print("The number of arrows with an x-value of zero or greater, and a y-value of zero or greater is ", q1)
print("The number of arrows with an x-value of zero or greater, and a y-value of below zero is ", q2)
print("The number of arrows with an x-value of below zero, and a y-value of below zero is ", q3)
print("The number of arrows with an x-value of below zero, and a y-value of zero or greater is ", q4)
# +
# 3. Find the point closest to the center. Calculate its distance to the center
# Defining a function that calculates the distance to the center can help.
# I should instead define a function that calculates the distance to the center, and returns the coordinate pair of the minimum-distance-value
distance_from_center = []
summation = 0
for i in range(len(points)):
summation = ((x_values[i] ** 2) + (y_values[i] ** 2))
distance_from_center.append(summation ** 0.5)
print("The shot closest to the center is index number", distance_from_center.index(min(distance_from_center)))
print("The distance of that shot to the center is", min(distance_from_center))
# +
# 4. If the target has a radius of 9, calculate the number of arrows that
# must be picked up in the forest.
forest_pickup = 0
for i in distance_from_center:
if i > 9:
forest_pickup += 1
#distance_from_center.sort(reverse=True)
#print(distance_from_center)
print("The number of arrows to retrive from the forest is", forest_pickup)
# -
| robin-hood/poad_solution_robinhood.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # python 多线程专题
# 1、线程创建
'''
threading.current_thread()表示当前线程,可以用name或getName()获取线程名称
'''
import threading
import time
def fun():
print('The {0} starting\n'.format(threading.current_thread().name))
time.sleep(1)
print('The {0} is end\n'.format(threading.current_thread().getName()))
if __name__=='__main__':
print('The {0} is starting\n'.format(threading.current_thread().name))
for _ in range(5):
th = threading.Thread(target = fun)
th.start()
print('The {0} is ended\n'.format(threading.current_thread().name))
# +
'''
线程创建,通过类的方式
__author__ = jyb
'''
import threading
class threadTest(threading.Thread):
def __init__(self):
threading.Thread.__init__(self)
def run(self):
print('%s is running'%threading.current_thread().name)
th1 = threadTest()
print('%s is running'%threading.current_thread().name)
th1.start()
# +
"""
join()函数的用法
"""
from threading import Thread
import time
t = time.time()
def myfun():
time.sleep(1)
print(1)
if __name__ =='__main__':
th_list = []
for _ in range(5):
th = threading.Thread(target = myfun)
th.start()
th_list.append(th)
for threadInstance in th_list:
threadInstance.join()
print(time.time()-t)
# -
# # 2、 线程锁机制
# ## LOCK锁,RLOCK锁
# +
"""
Locak
"""
import time
import threading
threadlock = threading.Lock()
a =100
def consumer():
threadlock.acquire()
global a
a = a-1
threadlock.release()
def producer():
threadlock.acquire()
global a
a = a+1
threadlock.release
th1 = threading.Thread(target=consumer)
th1.start()
th2 = threading.Thread(target=producer)
th2.start()
print(a)
# +
'''
RLOCK 锁,可重入锁,同一线程可以锁多次,锁的次数和释放次数要对应
'''
import threading
rlock = threading.RLock()
def myfun():
print('%s is apply for a lock firstly\n'%threading.current_thread().name)
if(rlock.acquire()):
print('%s get a lock successfully \n'%threading.current_thread().name)
time.sleep(1)
print('%s is apply for a lock again\n'%threading.current_thread().name)
if(rlock.acquire()):
print('%s get a lock again successfully\n'%threading.current_thread().name)
time.sleep(1)
print('%s release a lock firstly\n'%threading.current_thread().name)
rlock.release()
time.sleep(1)
print('%s release a lock again\n'%threading.current_thread().name)
rlock.release()
th1 = threading.Thread(target =myfun,name='threading-1')
th2 = threading.Thread(target = myfun,name='threading-2')
th3 = threading.Thread(target=myfun,name='threading-3')
th1.start()
th2.start()
th3.start()
# -
# 即使存在多核的时候,python多进程是真正的并行,而多线程是伪并行,实际上他只是交替执行。
#
# 是什么导致多线程,只能交替执行呢?是一个叫GIL(Global Interpreter Lock,全局解释器锁)的东西。
#
# 什么是GIL呢?
#
# 每个python进程有一个GIL锁,任何Python线程执行前,必须先获得GIL锁,然后,每执行100条字节码,解释器就自动释放GIL锁,让别的线程有机会执行。这个GIL全局锁实际上把所有线程的执行代码都给上了锁,所以,多线程在Python中只能交替执行,即使100个线程跑在100核CPU上,也只能用到1个核。
# # 3、线程通信机制:
# ## condition
# ## event
# ## queue
a = []
dir(iter(a))
b =(x for x in range(10))
dir(b)
| Threading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import package
from autodp import rdp_bank, rdp_acct, dp_acct,privacy_calibrator
import numpy as np
# declare the moment accountants
acct = rdp_acct.anaRDPacct()
# ## Some experiments to test how close RDP(inf) can be used to approximate RDP(2).
# +
import matplotlib.pyplot as plt
# %matplotlib inline
blist = [1,2,3]
rlist = []
ra_list = []
k=50
for b in blist:
func_laplace = lambda x: rdp_bank.RDP_laplace({'b': b}, x)
func_puredp = lambda x: rdp_bank.RDP_pureDP({'eps':1/b},x)
results = [func_laplace(i+1) for i in range(k)]
results1 = [func_puredp(i+1) for i in range(k)]
rlist.append(results)
ra_list.append(results1)
colorlist = ['C0', 'C1', 'C2']
plt.figure(num=1, figsize=(12, 8), dpi=80, facecolor='w', edgecolor='k')
for (item,item1,color) in zip(rlist,ra_list,colorlist):
plt.plot(range(k), item,'-o',color=color)
plt.plot(range(k),item1,':',color=color)
plt.legend(['b = 1','b=1 CDP','b=2','b=2 CDP', 'b=3','b=3 CDP'], loc='best')
plt.title('RDP of Laplace')
plt.xlabel('alpha')
plt.show()
# +
blist = np.exp(np.linspace(-5,5,100))
alphalist = [2,5, 10]
resultslist = []
for alpha in alphalist:
results =[]
for b in blist:
func_laplace = lambda x: rdp_bank.RDP_laplace({'b': 1.0*b}, x)
results.append(func_laplace(np.inf)/func_laplace(alpha))
resultslist.append(results)
plt.figure(num=1, figsize=(12, 8), dpi=80, facecolor='w', edgecolor='k')
for results in resultslist:
plt.loglog(blist, results,'-o')
plt.legend(['alpha = 2', 'alpha =5', 'alpha=10'], loc='best')
plt.xlabel('b')
plt.ylabel('PureDP divided by RDP(alpha)')
plt.show()
# -
# ## The better approximation
# +
blist = np.exp(np.linspace(-5,5,100))
alphalist = [2]
resultslist = []
for alpha in alphalist:
results =[]
for b in blist:
func_laplace = lambda x: rdp_bank.RDP_laplace({'b': 1.0*b}, x)
results.append(np.minimum(np.exp(func_laplace(np.inf))-1,(np.exp(func_laplace(np.inf))-1)**2)/(np.exp(func_laplace(alpha))-1))
resultslist.append(results)
plt.figure(num=1, figsize=(12, 8), dpi=80, facecolor='w', edgecolor='k')
for results in resultslist:
plt.semilogx(blist, results,'-o')
plt.legend([r'$\frac{\min \{e^{\epsilon}-1,(e^{\epsilon}-1)^2)}{e^{RDP(2)}-1}$'], loc='best',fontsize=20)
plt.xlabel('b')
plt.ylabel('ratio')
plt.show()
# +
acct = rdp_acct.anaRDPacct(m_lin_max=1300) # approx using eps(2) and eps
acct_approx = rdp_acct.anaRDPacct(m_lin_max=1300) # approx using only eps
acct_approx2 = rdp_acct.anaRDPacct(m_lin_max=1300) # approx using only subsampled cdp bound
acct_exact = rdp_acct.anaRDPacct(m_lin_max=1300) # direct calculation
b=5
prob = 0.01
params={}
params1 ={}
params2={}
params['b'] = b
laplace = lambda x: rdp_bank.RDP_laplace(params,x)
epsdp = lambda x:rdp_bank.RDP_pureDP({'eps':1/b},x)
params1['eps'] = laplace(np.inf)
params1['prob'] = prob
params2['eps'] = laplace(np.inf)
params2['eps2'] = laplace(2)
params2['prob'] = prob
func1 = lambda x: rdp_bank.RDP_subsampled_pureDP(params1, x)
func2 = lambda x: rdp_bank.RDP_subsampled_pureDP(params2, x)
acct.compose_mechanism(func2,coeff=1000)
acct_approx.compose_mechanism(func1,coeff=1000)
acct_exact.compose_poisson_subsampled_mechanisms(laplace, prob, coeff=1000)
acct_approx2.compose_poisson_subsampled_mechanisms(epsdp,prob,coeff=1000)
accts = [acct_exact,acct,acct_approx,acct_approx2]
resultslist = []
for item in accts:
item.build_zeroth_oracle()
rdps = [item.evalRDP(1.0*(i+1)) for i in range(2000)]
resultslist.append(rdps)
# +
plt.figure(num=1, figsize=(12, 8), dpi=80, facecolor='w', edgecolor='k')
for results in resultslist:
plt.loglog(np.linspace(1,2001,2000), results,'-o')
plt.legend(['exact','approx with eps(2) and eps(inf)',
'approx with eps(inf) only','approx with subsampled CDP'], loc='best',fontsize=20)
plt.xlabel('alpha')
plt.ylabel('RDP')
plt.show()
# -
| tutorials/legacy/pure-dp-approximation-scheme.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Visualization and Analysis
#
# This notebook is to go through and experiment ways to visualize and analyze the data.
#
#
import pandas as pd
import seaborn as sns
import numpy as np
from ggplot import *
import matplotlib.pyplot as plt
# +
###################
## Read in Data
####################
all_df = pd.read_table("../data/outputs/TFBS_map_DF_all_bicoid_test.csv", na_values = 'NA',sep= "\t", index_col = 0)
# +
# remove all rows with NAs
all_df = all_df.dropna()
# Check
print all_df
# -
ggplot(aes(x='species', y = 'strand'), data = all_df) +\
geom_bar()ga
| py/visualization_bicoid_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#In RL, the two major entities are agent and the environment
import random
class Environment:
def __init__(self):
self.steps = 10
def get_observations(self):
"""returns current environment's observation to the agent"""
return [0.0, 0.0, 0.0]
def get_actions(self):
return [0, 1]
def is_done(self):
return self.steps==0
def action(self, action):
if self.is_done():
raise Exception("Game Over")
self.steps -= 1
return random.random()
class Agent:
def __init__(self):
self.total_reward = 0.0
def step(self, env):
current_obs = env.get_observations()
actions = env.get_actions()
reward = env.action(random.choice(actions))
self.total_reward += reward
print(self.total_reward)
env = Environment()
agent = Agent()
while not env.is_done():
agent.step(env)
# -
| Basics of Reinforcement Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import keras
keras.__version__
# # Text generation with LSTM
#
# This notebook contains the code samples found in Chapter 8, Section 1 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
#
# ----
#
# [...]
#
# ## Implementing character-level LSTM text generation
#
#
# Let's put these ideas in practice in a Keras implementation. The first thing we need is a lot of text data that we can use to learn a
# language model. You could use any sufficiently large text file or set of text files -- Wikipedia, the Lord of the Rings, etc. In this
# example we will use some of the writings of Nietzsche, the late-19th century German philosopher (translated to English). The language model
# we will learn will thus be specifically a model of Nietzsche's writing style and topics of choice, rather than a more generic model of the
# English language.
# ## Preparing the data
#
# Let's start by downloading the corpus and converting it to lowercase:
# +
import keras
import numpy as np
path = keras.utils.get_file(
'nietzsche.txt',
origin='https://s3.amazonaws.com/text-datasets/nietzsche.txt')
text = open(path).read().lower()
print('Corpus length:', len(text))
# -
#
# Next, we will extract partially-overlapping sequences of length `maxlen`, one-hot encode them and pack them in a 3D Numpy array `x` of
# shape `(sequences, maxlen, unique_characters)`. Simultaneously, we prepare a array `y` containing the corresponding targets: the one-hot
# encoded characters that come right after each extracted sequence.
# +
# Length of extracted character sequences
maxlen = 60
# We sample a new sequence every `step` characters
step = 3
# This holds our extracted sequences
sentences = []
# This holds the targets (the follow-up characters)
next_chars = []
for i in range(0, len(text) - maxlen, step):
sentences.append(text[i: i + maxlen])
next_chars.append(text[i + maxlen])
print('Number of sequences:', len(sentences))
# List of unique characters in the corpus
chars = sorted(list(set(text)))
print('Unique characters:', len(chars))
# Dictionary mapping unique characters to their index in `chars`
char_indices = dict((char, chars.index(char)) for char in chars)
# Next, one-hot encode the characters into binary arrays.
print('Vectorization...')
x = np.zeros((len(sentences), maxlen, len(chars)), dtype=np.bool)
y = np.zeros((len(sentences), len(chars)), dtype=np.bool)
for i, sentence in enumerate(sentences):
for t, char in enumerate(sentence):
x[i, t, char_indices[char]] = 1
y[i, char_indices[next_chars[i]]] = 1
# -
# ## Building the network
#
# Our network is a single `LSTM` layer followed by a `Dense` classifier and softmax over all possible characters. But let us note that
# recurrent neural networks are not the only way to do sequence data generation; 1D convnets also have proven extremely successful at it in
# recent times.
# +
from keras import layers
model = keras.models.Sequential()
model.add(layers.LSTM(128, input_shape=(maxlen, len(chars))))
model.add(layers.Dense(len(chars), activation='softmax'))
# -
# Since our targets are one-hot encoded, we will use `categorical_crossentropy` as the loss to train the model:
optimizer = keras.optimizers.RMSprop(lr=0.01)
model.compile(loss='categorical_crossentropy', optimizer=optimizer)
# ## Training the language model and sampling from it
#
#
# Given a trained model and a seed text snippet, we generate new text by repeatedly:
#
# * 1) Drawing from the model a probability distribution over the next character given the text available so far
# * 2) Reweighting the distribution to a certain "temperature"
# * 3) Sampling the next character at random according to the reweighted distribution
# * 4) Adding the new character at the end of the available text
#
# This is the code we use to reweight the original probability distribution coming out of the model,
# and draw a character index from it (the "sampling function"):
def sample(preds, temperature=1.0):
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
#
# Finally, this is the loop where we repeatedly train and generated text. We start generating text using a range of different temperatures
# after every epoch. This allows us to see how the generated text evolves as the model starts converging, as well as the impact of
# temperature in the sampling strategy.
# +
import random
import sys
for epoch in range(1, 60):
print('epoch', epoch)
# Fit the model for 1 epoch on the available training data
model.fit(x, y,
batch_size=128,
epochs=1)
# Select a text seed at random
start_index = random.randint(0, len(text) - maxlen - 1)
generated_text = text[start_index: start_index + maxlen]
print('--- Generating with seed: "' + generated_text + '"')
for temperature in [0.2, 0.5, 1.0, 1.2]:
print('------ temperature:', temperature)
sys.stdout.write(generated_text)
# We generate 400 characters
for i in range(400):
sampled = np.zeros((1, maxlen, len(chars)))
for t, char in enumerate(generated_text):
sampled[0, t, char_indices[char]] = 1.
preds = model.predict(sampled, verbose=0)[0]
next_index = sample(preds, temperature)
next_char = chars[next_index]
generated_text += next_char
generated_text = generated_text[1:]
sys.stdout.write(next_char)
sys.stdout.flush()
print()
# -
#
# As you can see, a low temperature results in extremely repetitive and predictable text, but where local structure is highly realistic: in
# particular, all words (a word being a local pattern of characters) are real English words. With higher temperatures, the generated text
# becomes more interesting, surprising, even creative; it may sometimes invent completely new words that sound somewhat plausible (such as
# "eterned" or "troveration"). With a high temperature, the local structure starts breaking down and most words look like semi-random strings
# of characters. Without a doubt, here 0.5 is the most interesting temperature for text generation in this specific setup. Always experiment
# with multiple sampling strategies! A clever balance between learned structure and randomness is what makes generation interesting.
#
# Note that by training a bigger model, longer, on more data, you can achieve generated samples that will look much more coherent and
# realistic than ours. But of course, don't expect to ever generate any meaningful text, other than by random chance: all we are doing is
# sampling data from a statistical model of which characters come after which characters. Language is a communication channel, and there is
# a distinction between what communications are about, and the statistical structure of the messages in which communications are encoded. To
# evidence this distinction, here is a thought experiment: what if human language did a better job at compressing communications, much like
# our computers do with most of our digital communications? Then language would be no less meaningful, yet it would lack any intrinsic
# statistical structure, thus making it impossible to learn a language model like we just did.
#
#
# ## Take aways
#
# * We can generate discrete sequence data by training a model to predict the next tokens(s) given previous tokens.
# * In the case of text, such a model is called a "language model" and could be based on either words or characters.
# * Sampling the next token requires balance between adhering to what the model judges likely, and introducing randomness.
# * One way to handle this is the notion of _softmax temperature_. Always experiment with different temperatures to find the "right" one.
| 8.1-text-generation-with-lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !python ~/MasterProject/Code/ClinicaTools/AD-DL/clinicaaddl/clinicaaddl/main.py train /u/horlavanasta/MasterProject//DataAndExperiments/Experiments/Experiments-1.5T-3T/NNs_Bayesian/ResNet18/subject_model-ResNet18_preprocessing-linear_task-AD_CN_norm-1_loss-WeightedCrossEntropy_augmTrue --n_splits 1 --split 0 --batch_size 5
# +
def check_history(model_path, num_folds):
from visualize.data_utils import get_data_generic
return os.path.exists(os.path.join(model_path, "status.txt"))
def check_results(model_path, MS_list, num_folds):
import os
import pathlib
import numpy as np
currentDirectory = pathlib.Path(model_path)
currentPattern = "fold-*"
flag=True
for fold_dir in currentDirectory.glob(currentPattern):
fold = int(str(fold_dir).split("-")[-1])
selection_metrics = ["best_loss", "best_balanced_accuracy", "last_checkpoint"]
cnn_classification_dir = os.path.join(model_path, 'fold-%i' % fold, 'cnn_classification')
for selection_metric in selection_metrics:
modes = ['train', 'validation']
for ms_el in MS_list:
modes.append('test_' + ms_el)
for mode in modes:
if not os.path.exists(os.path.join(cnn_classification_dir, selection_metric,
'%s_image_level_metrics.tsv' % (mode))):
flag=False
return flag
def check_complete_test(model_path, num_folds, MS_list):
import json
path_params = os.path.join(model_path, "commandline_train.json")
return (check_history(model_path, num_folds) and check_results(model_path, MS_list, num_folds))
# -
def check_baesian_stat(model_path, MS_list, num_folds):
import os
import pathlib
import numpy as np
currentDirectory = pathlib.Path(model_path)
currentPattern = "fold-*"
flag=True
for fold_dir in currentDirectory.glob(currentPattern):
fold = int(str(fold_dir).split("-")[-1])
selection_metrics = ["best_loss", "best_balanced_accuracy", "last_checkpoint"]
cnn_classification_dir = os.path.join(model_path, 'fold-%i' % fold, 'cnn_classification')
for selection_metric in selection_metrics:
modes = ['test_' + ms_el for ms_el in MS_list]
for mode in modes:
if not os.path.exists(os.path.join(cnn_classification_dir, selection_metric,
'%s_image_level_stats.tsv' % (mode))):
flag=False
return flag
# +
import matplotlib.pyplot as plt
def get_rows_and_cols(data):
rows_matrix = {}
cols_matrix = {}
for data_type in data.keys():
cols_matrix[data_type] = [selection_metric.replace("_", " ") for selection_metric in data[data_type].keys()]
if data_type == "history":
cols_matrix[data_type] = ["loss", "balanced_accuracy"]
else:
cols_matrix[data_type] = [selection_metric.replace("_", " ") for selection_metric in data[data_type].keys()]
if data_type == "uncertainty_distribution":
rows_matrix[data_type] = [test_MS.replace("_", " ") for test_MS in
list(data[data_type][list(data[data_type].keys())[0]].groupby("mode", as_index=False, sort=False).groups.keys())]
else:
rows_matrix[data_type] = [None]
num_rows = sum([len(rows_matrix[row]) for row in rows_matrix.keys()])
num_cols = max([len(cols_matrix[col]) for col in cols_matrix.keys()])
return rows_matrix, cols_matrix, num_rows, num_cols
def plot_history(args, data, fig, row, figshape):
from .plot_utils import plot_history_ax
for col, history_mode in enumerate(args.history_modes):
ax = plt.subplot2grid(shape=figshape, loc=(row, col), fig=fig)
plot_history_ax(ax, data, mode=history_mode, aggregation_type=args.aggregation_type)
def plot_results(args, data, fig, row, figshape):
from .plot_utils import plot_results_ax, plot_results_agg_ax
for col, selection_mode in enumerate(list(data.keys())):
ax = plt.subplot2grid(shape=figshape, loc=(row, col), fig=fig)
if args.aggregation_type is not "all":
plot_results_ax(ax, data[selection_mode], args.result_metrics)
else:
plot_results_agg_ax(ax, data[selection_mode], args.result_metrics)
ax.set_title(selection_mode)
def plot_uncertainty_distribution(args, data, fig, row, figshape):
from .plot_utils import plot_catplot_ax, set_ylims_axes
axes = []
for col, selection_mode in enumerate(list(data.keys())):
for j, (mode, mode_group) in enumerate(data[selection_mode].groupby("mode", as_index=False, sort=False)):
ax = plt.subplot2grid(shape=figshape, loc=(row+j, col), fig=fig)
plot_catplot_ax(ax,mode_group, args.uncertainty_metric, args.ba_inference_mode, args.catplot_type )
ax.set_title(selection_mode+"; "+mode)
axes.append(ax)
set_ylims_axes(axes)
def plot_combined_plots(args, model_params, saved_file_path, data=None):
import matplotlib.pyplot as plt
readable_params = ['model', 'data_augmentation', 'batch_size', 'learning_rate', "loss", 'training MS']
rows_matrix, cols_matrix, num_rows, num_cols = get_rows_and_cols(data)
fig = plt.figure(figsize=((int(8 * num_cols), int(6 * num_rows))))
row = 0
for data_key in sorted(list(data.keys()), reverse=True):
eval("plot_%s" % (data_key))(args, data=data[data_key], fig=fig, figshape=(num_rows, num_cols), row=row)
row+=len(rows_matrix[data_key])
str_suptitle = "\n Params: "
for i, line in enumerate(readable_params):
str_suptitle += line + ': ' + str(model_params[line]) + "; "
str_suptitle +="\n"
plt.suptitle(str_suptitle)
# plt.subplots_adjust(left=0.05, right=0.95, top=0.95, bottom=0.05, wspace=0.1, hspace=0.1)
plt.subplots_adjust( left=0.05, right=0.95, top=0.95, bottom=0.05,hspace=0.3)
if saved_file_path is not None:
plt.savefig(saved_file_path)
else:
plt.show()
plt.close()
def plot_generic(
args,
training_MS,
):
import pathlib
import os
import json
import pandas as pd
from .data_utils import get_data_generic
currentDirectory = pathlib.Path(args.model_path)
path_params = os.path.join(currentDirectory, "commandline_train.json")
with open(path_params, "r") as f:
params = json.load(f)
params['training MS'] = training_MS
args.bayesian=params["bayesian"]
model_name = os.path.basename(os.path.normpath(currentDirectory))
folder_name = ''
for data_type in sorted(args.data_types):
if data_type=="uncertainty_distribution":
folder_name += '%s_uncertainty_%s' % (args.uncertainty_metric, args.catplot_type)
else:
folder_name += data_type
folder_name += "_"
data=[]
for f in args.models:
data = get_data_generic(args)
for fold_key in data.keys():
if args.aggregation_type=="separate":
folder_fold_name = os.path.join("separate_folds", "fold-%s"%fold_key)
else:
folder_fold_name = fold_key
if args.output_path:
saved_file_path = os.path.join(args.output_path, folder_fold_name, params["model"], folder_name)
os.makedirs(saved_file_path, exist_ok=True)
saved_file_path=os.path.join(saved_file_path, model_name + '.png')
else:
saved_file_path=None
plot_combined_plots(args, model_params=params, data=data[fold_key], saved_file_path=saved_file_path)
# -
# +
import pathlib
import pandas as pd
import os
import json
folders = []
MS_main_list = ['1.5T', "3T"]
MS_list_dict = {'1.5T':['1.5T', '3T'], "3T": ['3T', '1.5T'], "1.5T-3T": ["1.5T-3T"]}
home_folder='/u/horlavanasta/MasterProject/'
num_folds_arr=[5]
isBayesian_arr=[True]
num_folds=5
merged_file=os.path.join(home_folder,"DataAndExperiments/Data/DataStat", "merge.tsv")
for isBayesian in isBayesian_arr:
for MS in MS_main_list[:]:
print("MS %s \n ____________________________________________________________________________________________"%MS)
model_types = [ "ResNet18", "SEResNet18", "ResNet18Expanded", "SEResNet18Expanded", "Conv5_FC3", "ResNet50", "SEResNet50",]
MS_list = MS_list_dict[MS]
inference_modes=["mode", "mean"]
results_folder_general =os.path.join(home_folder, 'Code/ClinicaTools/AD-DL/results/', "Experiments_%d-fold"%num_folds, "Experiments_Bayesian" if isBayesian else "Experiments", 'Experiments-' + MS)
model_dir_general = os.path.join(home_folder,"DataAndExperiments/Experiments_%d-fold/Experiments-%s"%(num_folds, MS), "NNs_Bayesian" if isBayesian else "NNs")
for network in model_types[:]:
model_dir = os.path.join(model_dir_general, network)
# output_dir = pathlib.Path(output_dir)
modelPatter = "subject_model*"
folders = [f for f in pathlib.Path(model_dir).glob(modelPatter)]
for f in folders[:]:
if check_complete_test(f, num_folds=num_folds, MS_list=MS_list):
# pass
print(f)
for inference_mode in inference_modes:
results_dir=os.path.join(results_folder_general, "%s_inference"%inference_mode)
if not check_baesian_stat(f, num_folds=num_folds, MS_list=MS_list):
# !python ~/MasterProject/Code/ClinicaTools/AD-DL/clinicaaddl/clinicaaddl/main.py bayesian $f stat
# !python ~/MasterProject/Code/ClinicaTools/AD-DL/clinicaaddl/clinicaaddl/main.py visualize $f $results_dir uncertainty_distribution results history --merged_file $merged_file --ba_inference_mode $inference_mode --catplot_type violinplot --uncertainty_metric "total_variance"
# # !python ~/MasterProject/Code/ClinicaTools/AD-DL/clinicaaddl/clinicaaddl/main.py visualize $f $results_dir uncertainty_distribution results history --aggregation_type "separate" --ba_inference_mode $inference_mode --catplot_type violinplot --uncertainty_metric "total_variance"
else:
pass
# print(f)
# +
import pathlib
import pandas as pd
import os
import json
folders = []
MS_main_list = ["1.5T-3T", '1.5T', "3T"]
MS_list_dict = {'1.5T':['1.5T', '3T'], "3T": ['3T', '1.5T'], "1.5T-3T": ["1.5T-3T"]}
home_folder='/u/horlavanasta/MasterProject/'
num_folds_arr=[5]
isBayesian_arr=[False]
num_folds=5
merged_file=os.path.join(home_folder,"DataAndExperiments/Data/DataStat", "merge.tsv")
for isBayesian in isBayesian_arr:
for MS in MS_main_list[:]:
print("MS %s \n ____________________________________________________________________________________________"%MS)
model_types = [ "ResNet18", "SEResNet18", "ResNet18Expanded", "SEResNet18Expanded", "Conv5_FC3", "ResNet50", "SEResNet50" ]
MS_list = MS_list_dict[MS]
inference_modes=["mode", "mean"]
results_folder_general =os.path.join(home_folder, 'Code/ClinicaTools/AD-DL/results/', "Experiments_%d-fold"%num_folds, "Experiments_Bayesian" if isBayesian else "Experiments", 'Experiments-' + MS)
model_dir_general = os.path.join(home_folder,"DataAndExperiments/Experiments_%d-fold/Experiments-%s"%(num_folds, MS), "NNs_Bayesian" if isBayesian else "NNs")
for network in model_types[:]:
model_dir = os.path.join(model_dir_general, network)
# output_dir = pathlib.Path(output_dir)
modelPatter = "subject_model*"
folders = [f for f in pathlib.Path(model_dir).glob(modelPatter)]
for f in folders[:]:
if check_complete_test(f, num_folds=num_folds,MS_list=MS_list):
# pass
print(f)
results_dir=results_folder_general
# !python ~/MasterProject/Code/ClinicaTools/AD-DL/clinicaaddl/clinicaaddl/main.py visualize $f $results_dir results history --merged_file $merged_file --aggregation_type "all"
else:
pass
# print(f)
# +
def translate_parameters(args):
"""
Translate the names of the parameters between command line and source code.
"""
args.gpu = False
args.num_workers = args.nproc
args.optimizer = "Adam"
args.batch_size=9
# args.loss = "default"
if hasattr(args, "caps_dir"):
args.input_dir = args.caps_dir
if hasattr(args, "unnormalize"):
args.minmaxnormalization = not args.unnormalize
if hasattr(args, "slice_direction"):
args.mri_plane = args.slice_direction
if hasattr(args, "network_type"):
args.mode_task = args.network_type
if not hasattr(args, "selection_threshold"):
args.selection_threshold = None
if not hasattr(args, "verbose"):
args.verbose = 0
if not hasattr(args, "bayesian"):
args.bayesian = False
if not hasattr(args, "prepare_dl"):
if hasattr(args, "use_extracted_features"):
args.prepare_dl = args.use_extracted_features
elif hasattr(args, "use_extracted_patches") and args.mode == "patch":
args.prepare_dl = args.use_extracted_patches
elif hasattr(args, "use_extracted_slices") and args.mode == "slice":
args.prepare_dl = args.use_extracted_slices
elif hasattr(args, "use_extracted_roi") and args.mode == "roi":
args.prepare_dl = args.use_extracted_roi
return args
def show_fpg(data_batch, indices=None,plane="sag", num_rows=2,
num_cols=2, name=None, folder="/current_augmentations_examples/"):
import matplotlib.pyplot as plt
import numpy as np
fig, axes = plt.subplots(num_rows, num_cols, figsize=((int(8 * num_rows), int(6 * num_cols))))
data_batch=data_batch.numpy()
print(data_batch.shape)
data_batch=data_batch[:num_rows*num_cols].reshape(num_rows, num_cols, data_batch.shape[1], data_batch.shape[2], data_batch.shape[3],
data_batch.shape[4])
print(data_batch.shape)
for row in range(num_rows):
for col in range(num_cols):
i, j, k = indices
data=data_batch[row][col]
kwargs = dict(cmap='gray', interpolation='none')
slices=dict()
slices["sag"], slices["cor"], slices["axi"] = np.rot90(data[0, i]), np.rot90(data[0, :, j]), np.rot90(data[0, ..., k])
axes[row][col].imshow(slices[plane],**kwargs)
axes[row][col].axis('off')
# path = '../../outputs/'+folder
# if not os.path.exists(path):
# os.makedirs(path)
if name is not None:
fig.suptitle(name)
plt.subplots_adjust( left=0.05, right=0.95, top=0.95, bottom=0.05, wspace=0.05, hspace=0.05)
# plt.savefig(path + str(name) + '.png')
plt.show()
plt.close()
def show_data(model_folder, name=None, plane="sag"):
from tools.deep_learning.models import init_model
from tools.deep_learning.data import (get_transforms,
load_data,
return_dataset,
generate_sampler)
from tools.deep_learning.iotools import return_logger
from argparse import Namespace
from torch.utils.data import DataLoader
import torchvision.models
import hiddenlayer as hl
import torch
path_params = os.path.join(model_folder, "commandline_train.json")
with open(path_params, "r") as f:
params = json.load(f)
params = translate_parameters(Namespace(**params))
main_logger = return_logger(params.verbose, "main process")
train_transforms, all_transforms = get_transforms(params.mode,
minmaxnormalization=params.minmaxnormalization,
data_augmentation=None,
output_dir=None)
training_df, valid_df = load_data(
params.tsv_path,
params.diagnoses,
0,
n_splits=params.n_splits,
baseline=params.baseline,
logger=main_logger
)
data_valid = return_dataset(params.mode, params.input_dir, valid_df, params.preprocessing,
train_transformations=train_transforms, all_transformations=all_transforms,
params=params)
valid_loader = DataLoader(
data_valid,
batch_size=params.batch_size,
shuffle=False,
num_workers=params.num_workers,
pin_memory=True
)
sample = next(iter(valid_loader))
show_fpg(sample["image"], indices=(169//2, 208//2, 179//2), name=name, plane=plane)
# +
import pathlib
import pandas as pd
import os
import json
folders = []
MS_main_list = ['1.5T', "3T","1.5T-3T" ]
MS_list_dict = {'1.5T':['1.5T', '3T'], "3T": ['3T', '1.5T'], "1.5T-3T": ["1.5T-3T"]}
home_folder='/u/horlavanasta/MasterProject/'
isBayesian=True
for MS in MS_main_list[:1]:
print("MS %s \n ____________________________________________________________________________________________"%MS)
model_types = [ "ResNet18", "SEResNet18", "ResNet18Expanded", "SEResNet18Expanded", "Conv5_FC3" ]
MS_list = MS_list_dict[MS]
model_dir_general = os.path.join(home_folder,"DataAndExperiments/Experiments/Experiments-" + MS, "NNs" if isBayesian else "NNs")
for network in model_types[:]:
model_dir = os.path.join(model_dir_general, network)
# output_dir = pathlib.Path(output_dir)
modelPatter = "subject_model*"
folders = [f for f in pathlib.Path(model_dir).glob(modelPatter)]
for f in folders[:1]:
print(f)
show_model(f)
# show_data(f, plane="sag")
# show_data(f, plane="cor")
# show_data(f, plane="axi")
# -
hl_graph
| clinicaaddl/clinicaaddl/ShowResultsAllNetworks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # First steps through pyiron
# This section gives a brief introduction about fundamental concepts of pyiron and how they can be used to setup, run and analyze atomic simulations. As a first step we import the libraries [numpy](http://www.numpy.org/) for data analysis and [matplotlib](https://matplotlib.org/) for visualization.
import numpy as np
# %matplotlib inline
import matplotlib.pylab as plt
# To import pyiron simply use:
from pyiron import Project
# The Project object introduced below is central in pyiron. It allows to name the project as well as to derive all other objects such as structures, jobs etc. without having to import them. Thus, by code completion *Tab* the respective commands can be found easily.
# We now create a pyiron Project named 'first_steps'.
pr = Project(path='first_steps')
# The project name also applies for the directory that is created for the project.
# ## Perform a LAMMPS MD simulation
# Having created an instance of the pyiron Project we now perform a [LAMMPS](http://lammps.sandia.gov/) molecular dynamics simulation.
# For this basic simulation example we construct an fcc Al crystal in a cubic supercell (`cubic=True`). For more details on generating structures, please have a look at our [structures example](./structures.ipynb)
basis = pr.create_ase_bulk('Al', cubic=True)
supercell_3x3x3 = basis.repeat([3, 3, 3])
supercell_3x3x3.plot3d()
# Here `create_ase_bulk` uses the [ASE bulk module](https://wiki.fysik.dtu.dk/ase/ase/build/build.html). The structure can be modified - here we extend the original cell to a 3x3x3 supercell (`repeat([3, 3, 3]`). Finally, we plot the structure using [NGlview](http://nglviewer.org/nglview/latest/api.html).
# The project object allows to create various simulation job types. Here, we create a LAMMPS job.
job = pr.create_job(job_type=pr.job_type.Lammps, job_name='Al_T800K')
# Further, we specify a Molecular Dynamics simulation at $T=800$ K using the supercell structure created above.
job.structure = supercell_3x3x3
job.calc_md(temperature=800, pressure=0, n_ionic_steps=10000)
# To see all available interatomic potentials which are compatible with the structure (for our example they must contain Al) and the job type (here LAMMPS) we call `job.list_potentials()`.
job.list_potentials()
# From the above let us select the first potential in the list.
pot = job.list_potentials()[0]
print ('Selected potential: ', pot)
job.potential = pot
# To run the LAMMPS simulation (locally) we now simply use:
job.run()
# ## Analyze the calculation
# After the simulation has finished the information about the job can be accessed through the Project object.
job = pr['Al_T800K']
job
# Printing the job object (note that in Jupyter we don't have to call a print statement if the variable/object is in the last line). The output lists the variables (nodes) and the directories (groups). To get a list of all variables stored in the generic output we type:
job['output/generic']
# An animated 3d plot of the MD trajectories is created by:
job.animate_structure()
# To analyze the temperature evolution we plot it as function of the MD step.
temperatures = job['output/generic/temperature']
steps = job['output/generic/steps']
plt.plot(steps, temperatures)
plt.xlabel('MD step')
plt.ylabel('Temperature [K]');
# In the same way we can plot the trajectories.
pos = job['output/generic/positions']
x, y, z = [pos[:, :, i] for i in range(3)]
sel = np.abs(z) < 0.1
fig, axs = plt.subplots(1,1)
axs.scatter(x[sel], y[sel])
axs.set_xlabel('x [$\AA$]')
axs.set_ylabel('y [$\AA$]')
axs.set_aspect('equal', 'box');
# ## Perform a series of jobs
# To run the MD simulation for various temperatures we can simply loop over the desired temperature values.
for temperature in np.arange(200, 1200, 200):
job = pr.create_job(pr.job_type.Lammps,
'Al_T{}K'.format(int(temperature)))
job.structure = supercell_3x3x3
job.potential = pot
job.calc_md(temperature=temperature,
pressure=0,
n_ionic_steps=10000)
job.run()
# To inspect the list of jobs in our current project we type (note that the existing job from the previous excercise at $T=800$ K has been recognized and not run again):
pr
# We can now iterate over the jobs and extract volume and mean temperature.
vol_lst, temp_lst = [], []
for job in pr.iter_jobs(convert_to_object=False):
volumes = job['output/generic/volume']
temperatures = job['output/generic/temperature']
temp_lst.append(np.mean(temperatures[:-20]))
vol_lst.append(np.mean(volumes[:-20]))
# Then we can use the extracted information to plot the thermal expansion, calculated within the $NPT$ ensemble. For plotting the temperature values in ascending order the volume list is mapped to the sorted temperature list.
plt.figure()
vol_lst[:] = [vol_lst[np.argsort(temp_lst)[k]]
for k in range(len(vol_lst))]
plt.plot(sorted(temp_lst), vol_lst,
linestyle='-',marker='o',)
plt.title('Thermal expansion')
plt.xlabel('Temperature [K]')
plt.ylabel('Volume [$\AA^3$]');
# ## Create a series of projects
# We extend the previous example and compute the thermal expansion for three of the available aluminum potentials. First, let us create a new pyiron project named 'Al_potentials'. We can use the information of the previously run job 'Al_T200K' of the 'first_steps' project to find all the compatible potentials.
pr = Project('Al_potentials')
pot_lst = pr['../first_steps/Al_T200K'].load_object().list_potentials()[:3]
pot_lst
# Note again that `list_potentials()` automatically only returns the potentials that are compatible with the structure (chemical species) and the job type.
# We can now loop over the selected potentials and run the MD simulation for the desired temperature values for any of the potentials.
for pot in pot_lst:
print ('Interatomic potential used: ',pot)
pr_pot = pr.create_group(pot)
for temperature in np.arange(200, 1200, 200):
job = pr_pot.create_job(pr.job_type.Lammps,
'Al_T{}K'.format(int(temperature)))
job.structure = supercell_3x3x3
job.potential = pot
job.calc_md(temperature=temperature,
pressure=0,
n_ionic_steps=10000)
job.run()
# With the `pr.create_group()` command a new subproject (directory) is created named here by the name of the potential.
# For any particular potential the thermal expansion data can be obtained again by looping over the jobs performed using that potential. To obtain the thermal expansion curves for all the potentials used we can simply iterate over the subprojects (directories) created above by using the `pr.iter_groups()` command.
for p in pr.iter_groups():
vol_lst, temp_lst = [], []
for out in p.iter_jobs(path='output/generic'):
volumes = out['volume']
temperatures = out['temperature']
temp_lst.append(np.mean(temperatures[:-20]))
vol_lst.append(np.mean(volumes[:-20]))
# Plot only if there is a job in that group
if len(p.get_job_ids()) > 0:
plt.plot(temp_lst, vol_lst,
linestyle='-',marker='o',
label=p.name)
plt.legend(loc='best')
plt.title('Thermal expansion for different interatomic potentials')
plt.xlabel('Temperature [K]')
plt.ylabel('Volume [$\AA^3$]');
| notebooks/first_steps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## LSTM Keras method for stock prediction
#
# https://www.kdnuggets.com/2018/11/keras-long-short-term-memory-lstm-model-predict-stock-prices.html
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import pandas_datareader.data as web
# ### Load the dataset
start = pd.to_datetime('2016-01-01')
df = web.DataReader("^FCHI", data_source = 'yahoo', start = start )
df.head()
df.isnull().values.any()
# +
data = df [['Close']]
data = data.reset_index()
training_data = data[data['Date'] < pd.to_datetime('2020-01-01')].copy()
test_data = data[data['Date'] >= pd.to_datetime("2020-01-01")].copy()
training_data = training_data.set_index('Date')
test_data = test_data.set_index('Date')
# -
plt.figure(figsize=(14,4))
plt.plot(training_data.Close)
plt.plot(test_data.Close)
plt.ylabel("Price")
plt.xlabel("Date")
plt.legend(["Training Set", "Test Set"])
plt.title("CAC40 Close Price")
plt.show()
# ### Feature scaling
from sklearn.preprocessing import MinMaxScaler
sc = MinMaxScaler(feature_range = (0, 1))
training_set_scaled = sc.fit_transform(training_data)
len(training_set_scaled)
max(training_set_scaled)
# ### Creating Data with Timesteps
# +
X_train = []
y_train = []
for i in range(60, len(training_set_scaled)):
X_train.append(training_set_scaled[i-60:i, 0])
y_train.append(training_set_scaled[i, 0])
X_train, y_train = np.array(X_train), np.array(y_train)
X_train = np.reshape(X_train, (X_train.shape[0], X_train.shape[1], 1))
# -
# ### Building the LSTM
#
# In order to build the LSTM, we need to import a couple of modules from Keras:
#
# Sequential for initializing the neural network
# Dense for adding a densely connected neural network layer
# LSTM for adding the Long Short-Term Memory layer
# Dropout for adding dropout layers that prevent overfitting
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers import Dropout
from keras import optimizers
# +
regressor = Sequential()
regressor.add(LSTM(units = 50, return_sequences = True, input_shape = (X_train.shape[1], 1)))
regressor.add(Dropout(0.2))
regressor.add(LSTM(units = 50, return_sequences = True))
regressor.add(Dropout(0.2))
regressor.add(LSTM(units = 50, return_sequences = True))
regressor.add(Dropout(0.2))
regressor.add(LSTM(units = 50))
regressor.add(Dropout(0.2))
regressor.add(Dense(units = 1))
optimizer = optimizers.Adam(clipvalue=0.5)
regressor.compile(optimizer=optimizer, loss='mean_squared_error')
regressor.fit(X_train, y_train, epochs = 10, batch_size = 128)
# -
# ### Predicting future stock
dataset_test = pd.read_csv('^FCHI_test.csv')
real_stock_price = dataset_test.iloc[:, 3:4].values
len(real_stock_price)
# In order to predict future stock prices we need to do a couple of things after loading in the test set:
#
# Merge the training set and the test set on the 0 axis.
# Set the time step as 60 (as seen previously)
# Use MinMaxScaler to transform the new dataset
# Reshape the dataset as done previously
# After making the predictions we use inverse_transform to get back the stock prices in normal readable format.
# +
total_data = pd.concat((training_data, test_data), axis = 0)
inputs = total_data[len(total_data) - len(test_data) - 60:].values
inputs = inputs.reshape(-1,1)
inputs = sc.transform(inputs)
# shaping data from neural network
X_test = []
y_test = []
for i in range(60, len(inputs)):
X_test.append(inputs[i-60:i,0])
y_test.append(inputs[i,0])
X_test, y_test = np.array(X_test), np.array(y_test)
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
# -
predicted_price = regressor.predict(X_test)
predicted_price = sc.inverse_transform(predicted_price)
predicted_price = pd.DataFrame(predicted_price)
predicted_price.rename(columns = {0: 'CAC40_predicted'}, inplace=True)
predicted_price = predicted_price.round(decimals=0)
predicted_price.index = test_data.index
# ### Plotting the results
# +
from sklearn.metrics import mean_squared_error
plt.figure(figsize = (14,5))
mse = mean_squared_error(y_test, predicted_price)
plt.plot(predicted_price['CAC40_predicted'], color = 'red', label = 'Predicted CAC40 closing price')
plt.plot(test_data.Close, color = 'green', label = 'Actual CAC40 closing price')
plt.title ("CAC40 closing Price prediction- with MSE {:10.4f}".format(mse))
plt.xlabel('Time')
plt.ylabel('Price (EUR)')
plt.legend()
# -
| LSTM/LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plot Kmeans clusters stored in a GeoTiff
#
# This is a notebook plots the GeoTiffs created out of [kmeans](../stable/kmeans.ipynb). Such GeoTiffs contains the Kmeans cluster IDs.
# ## Dependencies
# +
import sys
sys.path.append("/usr/lib/spark/python")
sys.path.append("/usr/lib/spark/python/lib/py4j-0.10.4-src.zip")
sys.path.append("/usr/lib/python3/dist-packages")
import os
os.environ["HADOOP_CONF_DIR"] = "/etc/hadoop/conf"
import os
os.environ["PYSPARK_PYTHON"] = "python3"
os.environ["PYSPARK_DRIVER_PYTHON"] = "ipython"
from pyspark.mllib.clustering import KMeans, KMeansModel
from pyspark import SparkConf, SparkContext
from osgeo import gdal
from io import BytesIO
import matplotlib.pyplot as plt
import rasterio
from rasterio import plot
from rasterio.io import MemoryFile
# -
# ## Spark Context
# +
appName = "plot_kmeans_clusters"
masterURL="spark://pheno0.phenovari-utwente.surf-hosted.nl:7077"
try:
sc.stop()
except NameError:
print("A new Spark Context will be created.")
sc = SparkContext(conf = SparkConf().setAppName(appName).setMaster(masterURL))
# -
# ## Mode of Operation setup
#
# The user should modify the following variables to define which GeoTiffs should be loaded. In case it (s)he wants to visualize results that just came out of [kmeans](kmeans.ipnyb) laste execution, just copy the values set at its [**Mode of Operation Setup**](../stable/kmeans.ipynb#mode_of_operation_setup).
# +
#GeoTiffs to be read from "hdfs:///user/hadoop/spring-index/"
dir_path = "hdfs:///user/hadoop/spring-index/"
offline_dir_path = "hdfs:///user/pheno/spring-index/"
geoTiff_dir = "BloomFinal"
#Kmeans number of iterations and clusters
numIterations = 35
minClusters = 2
maxClusters = 15
stepClusters = 1
# -
# ## Mode of Operation verification
# +
geotiff_hdfs_paths = []
if minClusters > maxClusters:
maxClusters = minClusters
stepClusters = 1
if stepClusters < 1:
stepClusters = 1
numClusters_id = 0
numClusters = minClusters
while numClusters <= maxClusters :
path = offline_dir_path + geoTiff_dir + '/clusters_' + str(numClusters) + '_' + str(numIterations) + '.tif'
geotiff_hdfs_paths.append(path)
numClusters_id += 1
numClusters += stepClusters
# -
# ## Load GeoTiffs
#
# Load the GeoTiffs into MemoryFiles.
# +
clusters_dataByteArrays = []
numClusters_id = 0
numClusters = minClusters
while numClusters <= maxClusters :
clusters_data = sc.binaryFiles(geotiff_hdfs_paths[numClusters_id]).take(1)
clusters_dataByteArrays.append(bytearray(clusters_data[0][1]))
numClusters_id += 1
numClusters += stepClusters
# -
# ## Check GeoTiffs metadata
for val in clusters_dataByteArrays:
#Create a Memory File
memFile = MemoryFile(val).open()
print(memFile.profile)
memFile.close()
# ## Plot GeoTiffs
# +
# %matplotlib inline
numClusters_id = 0
numClusters = minClusters
while numClusters <= maxClusters :
print ("Plot for " + str(numClusters) + " clusters!!!")
memFile = MemoryFile(clusters_dataByteArrays[numClusters_id]).open()
plot.show((memFile,1))
if (numClusters < maxClusters) :
_ = input("Press [enter] to continue.")
memFile.close()
numClusters_id += 1
numClusters += stepClusters
# -
| applications/notebooks/stable/plot_kmeans_clusters-Light.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Jupyter notebooks
#
# This is a [Jupyter](http://jupyter.org/) notebook using Python. You can install Jupyter locally to edit and interact with this notebook.
#
# # Higher order finite difference methods
#
# ## Lagrange Interpolating Polynomials
#
# Suppose we are given function values $u_0, \dotsc, u_n$ at the distinct points $x_0, \dotsc, x_n$ and we would like to build a polynomial of degree $n$ that goes through all these points. This explicit construction is attributed to Lagrange (though he was not first):
#
# $$ p(x) = \sum_{i=0}^n u_i \prod_{j \ne i} \frac{x - x_j}{x_i - x_j} $$
#
# * What is the degree of this polynomial?
# * Why is $p(x_i) = u_i$?
# * How expensive (in terms of $n$) is it to evaluate $p(x)$?
# * How expensive (in terms of $n$) is it to convert to standard form $p(x) = \sum_{i=0}^n a_i x^i$?
# * Can we easily evaluate the derivative $p'(x)$?
# * What can go wrong? Is this formulation numerically stable?
#
# A general derivation of finite difference methods for approximating $p^{(k)}(x)$ using function values $u(x_i)$ is to construct the Lagrange interpolating polynomial $p(x)$ from the function values $u_i = u(x_i)$ and evaluate it or its derivatives at the target point $x$. We can do this directly from the formula above, but a more linear algebraic approach will turn out to be more reusable.
# #### Uniqueness
#
# Is the polynomial $p(x)$ of degree $m$ that interpolates $m+1$ points unique? Why?
#
# ### Vandermonde matrices
#
# We can compute a polynomial
#
# $$ p(x) = c_0 + c_1 x + c_2 x^2 + \dotsb $$
#
# that assumes function values $p(x_i) = u_i$ by solving a linear system with the Vandermonde matrix.
#
# $$ \underbrace{\begin{bmatrix} 1 & x_0 & x_0^2 & \dotsb \\
# 1 & x_1 & x_1^2 & \dotsb \\
# 1 & x_2 & x_2^2 & \dotsb \\
# \vdots & & & \ddots \end{bmatrix}}_V \begin{bmatrix} c_0 \\ c_1 \\ c_2 \\ \vdots \end{bmatrix} = \begin{bmatrix} u_0 \\ u_1 \\ u_2 \\ \vdots \end{bmatrix} .$$
# +
# %matplotlib inline
import numpy
from matplotlib import pyplot
pyplot.style.use('ggplot')
x = numpy.linspace(-2,2,4)
u = numpy.sin(x)
xx = numpy.linspace(-3,3,40)
c = numpy.linalg.solve(numpy.vander(x), u)
pyplot.plot(x, u, '*')
pyplot.plot(xx, numpy.vander(xx, 4).dot(c), label='p(x)')
pyplot.plot(xx, numpy.sin(xx), label='sin(x)')
pyplot.legend(loc='upper left');
# -
# Given the coefficients $c = V^{-1} u$, we find
#
# $$ \begin{align} p(0) &= c_0 \\ p'(0) &= c_1 \\ p''(0) &= c_2 \cdot 2! \\ p^{(k)}(0) &= c_k \cdot k! . \end{align} $$
#
# To compute the stencil coefficients $s_i^0$ for interpolation to $x=0$,
# $$ p(0) = s_0^0 u_0 + s_1^0 u_1 + \dotsb = \sum_i s_i^0 u_i $$
# we can write
# $$ p(0) = e_0^T \underbrace{V^{-1} u}_c = \underbrace{e_0^T V^{-1}}_{(s^0)^T} u $$
# where $e_0$ is the first column of the identity. Evidently $s^0$ can also be expressed as
# $$ s^0 = V^{-T} e_0 . $$
# We can compute stencil coefficients for any order derivative $p^{(k)}(0) = (s^k)^T u$ by solving the linear system
# $$ s^k = V^{-T} e_k \cdot k! . $$
# Alternatively, invert the Vandermonde matrix $V$ and scale row $k$ of $V^{-1}$ by $k!$.
# +
def fdstencil(z, x):
x = numpy.array(x)
V = numpy.vander(x - z, increasing=True)
scaling = numpy.array([numpy.math.factorial(i) for i in range(len(x))])
return (numpy.linalg.inv(V).T * scaling).T
x = numpy.linspace(0,3,4)
S = fdstencil(0, x)
print(S)
hs = 2.**(-numpy.arange(6))
errors = numpy.zeros((3,len(hs)))
for i,h in enumerate(hs):
z = 1 + .3*h
S = fdstencil(z, 1+x*h)
u = numpy.sin(1+x*h)
errors[:,i] = S[:3].dot(u) - numpy.array([numpy.sin(z), numpy.cos(z), -numpy.sin(z)])
pyplot.loglog(hs, numpy.abs(errors[0]), 'o', label="$p(0)$")
pyplot.loglog(hs, numpy.abs(errors[1]), '<', label="$p'(0)$")
pyplot.loglog(hs, numpy.abs(errors[2]), 's', label="$p''(0)$")
for k in (1,2,3):
pyplot.loglog(hs, hs**k, label='$h^{%d}$' % k)
pyplot.legend(loc='upper left');
# -
# ### Notes on accuracy
#
# * When using three points, we fit a polynomial of degree 2. The leading error term for interpolation $p(0)$ is thus $O(h^3)$.
# * Each derivative gives up one order of accuracy, therefore differencing to a general (non-centered or non-uniform grid) point is $O(h^2)$ for the first derivative and $O(h)$ for the second derivative.
# * Centered differences on uniform grids can provide cancelation, raising the order of accuracy by one. So our standard 3-point centered second derivative is $O(h^2)$ as we have seen in the Taylor analysis and numerically.
# * The Vandermonde matrix is notoriously ill-conditioned when using many points. For such cases, we recommend using a more numerically stable method from [Fornberg](https://doi.org/10.1137/S0036144596322507).
-fdstencil(0, numpy.linspace(-1,4,6))[2]
# ### Solving BVPs
#
# This `fdstencil` gives us a way to compute derivatives of arbitrary accuracy on arbitrary grids. We will need to use uncentered rules near boundaries, usually with more points to maintain order of accuracy. This will usually cost us symmetry. Implementation of boundary conditions is the bane of high order finite difference methods.
# ### Discretization stability measures: $h$-ellipticity
#
# Consider the test function $\phi(\theta, x) = e^{i\theta x}$ and apply the difference stencil centered at an arbitrary point $x$ with element size $h=1$:
#
# $$ \begin{bmatrix} -1 & 2 & -1 \end{bmatrix} \begin{bmatrix} e^{i \theta (x - 1)} \\ e^{i \theta x} \\ e^{i \theta (x+1)} \end{bmatrix}
# = \big( 2 - (e^{i\theta} + e^{-i\theta}) \big) e^{i\theta x}= 2 (1 - \cos \theta) e^{i \theta x} . $$
#
# Evidently $\phi(\theta,x) = e^{i \theta x}$ is an eigenfunction of the discrete differencing operator on an infinite grid and the corresponding eigenvalue is
# $$ L(\theta) = 2 (1 - \cos \theta), $$
# also known as the "symbol" of the operator. That $\phi(\theta,x)$ is an eigenfunction of the discrete differencing formula will generally be true for uniform grids.
#
# The highest frequency that is distinguishable using this stencil is $\theta_{\max} = \pi$ which results in a wave at the Nyquist frequency. If a higher frequency wave is sampled onto this grid, it will be aliased back into the interval $[-\pi, \pi)$.
x = numpy.linspace(-1, 1, 3)
s2 = -fdstencil(0, x)[2]
print(s2)
theta = numpy.linspace(-numpy.pi, numpy.pi)
phi = numpy.exp(1j*numpy.outer(x, theta))
pyplot.plot(theta, numpy.abs(s2.dot(phi)), '.')
pyplot.plot(theta, 2*(1-numpy.cos(theta)))
pyplot.plot(theta, theta**2);
# A measure of internal stability known as $h$-ellipticity is defined by
#
# $$ E^h(L) = \frac{\min_{\pi/2 \le |\theta| \le \pi} L(\theta)}{\max_{|\theta| \le \pi} L(\theta)} . $$
#
# * What is $E^h(L)$ for the second order "version 2" stencil?
# * How about for uncentered formulas and higher order?
# # Spectral collocation
#
# Suppose that instead of using only a fixed number of neighbors in our differencing stencil, we use all points in the domain?
# +
n = 10
x = numpy.linspace(-1, 1, n)
L = numpy.zeros((n,n))
for i in range(n):
L[i] = -fdstencil(x[i], x)[2]
u = numpy.cos(3*x)
pyplot.plot(x, L.dot(u), 'o')
pyplot.plot(x, 9*u);
# -
# We are suffering from two problems here. The first is that the monomial basis is very ill-conditioned when using many terms. This is true as continuous functions, not just when sampled onto a particular grid.
x = numpy.linspace(-1, 1, 50)
V = numpy.vander(x, 15)
pyplot.plot(x, V)
numpy.linalg.cond(V)
# ## Chebyshev polynomials
#
# Define $$ T_n(x) = \cos (n \arccos(x)) .$$
# This turns out to be a polynomial, though it may not be obvious why.
# Recall $$ \cos(a + b) = \cos a \cos b - \sin a \sin b .$$
# Let $y = \arccos x$ and check
# $$ \begin{split}
# T_{n+1}(x) &= \cos (n+1) y = \cos ny \cos y - \sin ny \sin y \\
# T_{n-1}(x) &= \cos (n-1) y = \cos ny \cos y + \sin ny \sin y
# \end{split}$$
# Adding these together produces a similar recurrence:
# $$\begin{split}
# T_0(x) &= 1 \\
# T_1(x) &= x \\
# T_{n+1}(x) &= 2 x T_n(x) - T_{n-1}(x)
# \end{split}$$
# which we can also implement in code
# +
def vander_chebyshev(x, n=None):
if n is None:
n = len(x)
T = numpy.ones((len(x), n))
if n > 1:
T[:,1] = x
for k in range(2,n):
T[:,k] = 2 * x * T[:,k-1] - T[:,k-2]
return T
x = numpy.linspace(-1, 1)
V = vander_chebyshev(x, 5)
pyplot.plot(x, V)
numpy.linalg.cond(V)
# -
# We can use the Chebyshev basis for interpolation
x = numpy.linspace(-2, 2, 4)
u = numpy.sin(x)
c = numpy.linalg.solve(vander_chebyshev(x), u)
pyplot.plot(x, u, '*')
pyplot.plot(xx, vander_chebyshev(xx, 4).dot(c), label='p(x)')
pyplot.plot(xx, numpy.sin(xx), label='sin(x)')
pyplot.legend(loc='upper left');
# ### Differentiation
#
# We can differentiate Chebyshev polynomials using the recurrence
#
# $$ \frac{T_n'(x)}{n} = 2 T_{n-1}(x) + \frac{T_{n-2}'(x)}{n-2} $$
#
# which we can differentiate to evaluate higher derivatives.
# +
def chebeval(z, n=None):
"""Build matrices to evaluate the n-term Chebyshev expansion and its derivatives at point(s) z"""
z = numpy.array(z, ndmin=1)
if n is None:
n = len(z)
Tz = vander_chebyshev(z, n)
dTz = numpy.zeros_like(Tz)
dTz[:,1] = 1
dTz[:,2] = 4*z
ddTz = numpy.zeros_like(Tz)
ddTz[:,2] = 4
for n in range(3,n):
dTz[:,n] = n * (2*Tz[:,n-1] + dTz[:,n-2]/(n-2))
ddTz[:,n] = n * (2*dTz[:,n-1] + ddTz[:,n-2]/(n-2))
return [Tz, dTz, ddTz]
n = 44
x = numpy.linspace(-1, 1, n)
T = vander_chebyshev(x)
print('cond = {:e}'.format(numpy.linalg.cond(T)))
Tinv = numpy.linalg.inv(T)
L = numpy.zeros((n,n))
for i in range(n):
L[i] = chebeval(x[i], n)[2].dot(Tinv)
u = numpy.cos(3*x)
pyplot.plot(x, L.dot(u), 'o')
xx = numpy.linspace(-1, 1, 100)
pyplot.plot(xx, -9*numpy.cos(3*xx));
# -
# ### Runge Effect
#
# Polynomial interpolation on equally spaced points is very ill-conditioned as the number of points grows. We've seen that in the growth of the condition number of the Vandermonde matrix, both for monomials and Chebyshev polynomials, but it's also true if the polynomials are measured in a different norm, such as pointwise values or merely the eyeball norm.
# +
def chebyshev_interp_and_eval(x, xx):
"""Matrix mapping from values at points x to values
of Chebyshev interpolating polynomial at points xx"""
A = vander_chebyshev(x)
B = vander_chebyshev(xx, len(x))
return B.dot(numpy.linalg.inv(A))
def runge1(x):
return 1 / (1 + 10*x**2)
x = numpy.linspace(-1,1,20)
xx = numpy.linspace(-1,1,100)
pyplot.plot(x, runge1(x), 'o')
pyplot.plot(xx, chebyshev_interp_and_eval(x, xx).dot(runge1(x)));
# -
x = cosspace(-1,1,8)
pyplot.plot(xx, chebyshev_interp_and_eval(x,xx))
numpy.outer(numpy.arange(4), [10,20])
ns = numpy.arange(5,20)
conds = [numpy.linalg.cond(chebyshev_interp_and_eval(numpy.linspace(-1,1,n),
numpy.linspace(-1,1,100)))
for n in ns]
pyplot.semilogy(ns, conds);
# This ill-conditioning cannot be fixed when using polynomial *interpolation* on equally spaced grids.
#
# ### Chebyshev nodes
#
# The Chebyshev polynomials assume their maximum value of 1 at points where their derivatives are zero (plus the endpoints). Choosing the roots of $T_n'(x)$ (plus endpoints) will control the polynomials and should lead to a well-conditioned formulation.
# +
def cosspace(a, b, n=50):
return (a + b)/2 + (b - a)/2 * (numpy.cos(numpy.linspace(-numpy.pi, 0, n)))
conds = [numpy.linalg.cond(chebyshev_interp_and_eval(cosspace(-1,1,n),
numpy.linspace(-1,1,100)))
for n in ns]
pyplot.figure()
pyplot.plot(ns, conds);
# -
x = cosspace(-1, 1, 7)
pyplot.plot(x, 0*x, 'o')
pyplot.plot(xx, chebeval(xx, 7)[1]);
x = cosspace(-1,1,20)
xx = numpy.linspace(-1,1,100)
pyplot.figure()
pyplot.plot(x, runge1(x), 'o')
pyplot.plot(xx, chebyshev_interp_and_eval(x, xx).dot(runge1(x)));
# ## Chebyshev solution of Boundary Value Problems
#
# If instead of an equally (or arbitrarily) spaced grid, we choose the Chebyshev nodes and compute derivatives in a stable way (e.g., via interpolating into the Chebyshev basis), we should have a very accurate method. Let's return to our test equation
#
# $$ -u''(x) = f(x) $$
#
# subject to some combination of Neumann and Dirichlet boundary conditions.
# +
def laplacian_cheb(n, rhsfunc, left, right):
"""Solve the Laplacian boundary value problem on (-1,1) using n elements with rhsfunc(x) forcing.
The left and right boundary conditions are specified as a pair (deriv, func) where
* deriv=0 for Dirichlet u(x_endpoint) = func(x_endpoint)
* deriv=1 for Neumann u'(x_endpoint) = func(x_endpoint)"""
x = cosspace(-1, 1, n+1) # n+1 points is n "elements"
T = chebeval(x)
L = -T[2]
rhs = rhsfunc(x)
for i,deriv,func in [(0, *left), (-1, *right)]:
L[i] = T[deriv][i]
rhs[i] = func(x[i])
return x, L.dot(numpy.linalg.inv(T[0])), rhs
class exact_tanh:
def __init__(self, k=1, x0=0):
self.k = k
self.x0 = x0
def u(self, x):
return numpy.tanh(self.k*(x - self.x0))
def du(self, x):
return self.k * numpy.cosh(self.k*(x - self.x0))**(-2)
def ddu(self, x):
return -2 * self.k**2 * numpy.tanh(self.k*(x - self.x0)) * numpy.cosh(self.k*(x - self.x0))**(-2)
ex = exact_tanh(5, 0.3)
x, L, rhs = laplacian_cheb(50, lambda x: -ex.ddu(x),
left=(0,ex.u), right=(0,ex.u))
uu = numpy.linalg.solve(L, rhs)
pyplot.plot(x, uu, 'o')
pyplot.plot(xx, ex.u(xx))
pyplot.plot(xx, chebeval(xx)[0][:,:51].dot(numpy.linalg.solve(chebeval(x)[0], uu)))
print(numpy.linalg.norm(numpy.linalg.solve(L, rhs) - ex.u(x), numpy.inf))
# +
def mms_error(n, discretize, sol):
x, L, f = discretize(n, lambda x: -sol.ddu(x), left=(0,sol.u), right=(1,sol.du))
u = numpy.linalg.solve(L, f)
return numpy.linalg.norm(u - sol.u(x), numpy.inf)
ns = numpy.arange(10,60,2)
errors = [mms_error(n, laplacian_cheb, ex) for n in ns]
pyplot.figure()
pyplot.semilogy(ns, errors, 'o', label='numerical')
for p in range(1,5):
pyplot.semilogy(ns, 1/ns**(p), label='$n^{-%d}$'%p)
pyplot.xlabel('n')
pyplot.ylabel('error')
pyplot.legend(loc='lower left');
# -
# # Homework 1: Due 2017-09-25
#
# Use a Chebyshev method to solve the second order ordinary differential equation
#
# $$ u''(t) + a u'(t) + b u(t) = f(t) $$
#
# from $t=0$ to $t=1$ with initial conditions $u(0) = 1$ and $u'(0) = 0$.
#
# 1. Do a grid convergence study to test the accuracy of your method.
# * Setting $f(t)=0$, experiment with the values $a$ and $b$ to identify two regimes with qualitatively different dynamics.
| FDHighOrder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
# +
import numpy as np
import matplotlib.pyplot as plt
import pycomlink as pycml
# -
# # Read in example data from one CML
cml = pycml.io.examples.read_one_cml()
# +
# Remove artifacts and plot data
cml.process.quality_control.set_to_nan_if('tx', '>=', 100)
cml.process.quality_control.set_to_nan_if('rx', '==', -99.9)
cml.plot_data(['tx', 'rx', 'txrx']);
# -
# # Do a simple wet/dry classification
cml.process.wet_dry.std_dev(window_length=30, threshold=0.8)
cml.plot_data(['txrx', 'wet']);
# # Derive a constant baseline
# Let's just focus on the rain events on 2016-10-25
cml.process.baseline.constant()
cml.process.baseline.calc_A()
ax = cml.plot_data(['txrx', 'wet', 'baseline', 'A']);
ax[0].set_xlim('2016-10-25 00:00', '2016-10-25 10:00');
# Save a copy of these results for comparing them to the linear baseline later
baseline_constant = cml.channel_1.data.baseline.copy()
A_constant = cml.channel_1.data.A.copy()
# # Or derive a linear baseline
cml.process.baseline.linear()
cml.process.baseline.calc_A()
ax = cml.plot_data(['txrx', 'wet', 'baseline', 'A']);
ax[0].set_xlim('2016-10-25 00:00', '2016-10-25 10:00');
# Save a copy of these results for comparing them to the constant baseline
baseline_linear = cml.channel_1.data.baseline.copy()
A_linear = cml.channel_1.data.A.copy()
# # Compare the results from constant and linear baseline
# +
fig, ax = plt.subplots(2, 1, figsize=(10, 4), sharex=True)
ax[0].plot(baseline_constant, color='C3', label='constant baseline')
ax[0].plot(baseline_linear, color='C4', label='linear baseline')
ax[1].plot(A_constant, color='C3', label='constant baseline')
ax[1].plot(A_linear, color='C4', label='linear baseline')
ax[0].set_xlim('2016-10-25 00:00', '2016-10-25 10:00');
ax[0].set_ylabel('baseline')
ax[1].set_ylabel('A')
ax[0].legend();
# -
# # NaN handling
# The algorithms for constant and linear baseline handle `NaN`s differently:
# * constant baseline:
# * For `NaN` values in `wet` the `baseline` is also set to `NaN`.
# * All `baseline` values following a `NaN` during a wet event are also set to `NaN` till the next dry event starts. This has to be done, since we do not know if a new wet event started during a `NaN` period and hence we do not know at which level the constant baseline should be.
# * linear baseline:
# * Default:
# The `baseline` for a whole wet event is set to `NaN` if there is at least one `wet` `NaN` within this period. This makes sense, since for the interpolation of the linear baseline the correct end of the wet period has to be known to its `txrx` value. Since the wet event could have ended during the `NaN` period, we do not know the end of the wet period and hence cannot savely assume a `txrx` endpoint for the interpolation.
# * Option to `ignore_nan`:
# If you know what you are doing, e.g. because you know that your only have very few consecutive `wet` `NaN`s and hecne can assume that a wet event will not stop during your `wet` `NaN`, then you can ignore all `NaN`s. This will take the next switch from wet to dry as endpoint of the wet event and do the interpolation accordingly.
# Exchange the current `wet` pd.Series in `channel_1` with a different series of floats with some `NaN`s
wet_temp = cml.channel_1.data.wet.astype(float)
wet_temp['2016-10-25 04:45': '2016-10-25 05:00'] = np.NaN
cml.channel_1.data.wet = wet_temp
# ## Constant baseline
cml.process.baseline.constant()
cml.process.baseline.calc_A()
ax = cml.plot_data(['txrx', 'wet', 'baseline', 'A']);
ax[0].set_xlim('2016-10-25 00:00', '2016-10-25 10:00');
# # Linear baseline (default)
# default = set `baseline` for whole wet event to `NaN` if it contains at least one `wet` `NaN`
cml.process.baseline.linear()
cml.process.baseline.calc_A()
ax = cml.plot_data(['txrx', 'wet', 'baseline', 'A']);
ax[0].set_xlim('2016-10-25 00:00', '2016-10-25 10:00');
# ## Linear baseline (ignoring `NaN`s)
cml.process.baseline.linear(ignore_nan=True)
cml.process.baseline.calc_A()
ax = cml.plot_data(['txrx', 'wet', 'baseline', 'A']);
ax[0].set_xlim('2016-10-25 00:00', '2016-10-25 10:00');
| notebooks/Baseline determination.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_pytorch_p36
# language: python
# name: conda_pytorch_p36
# ---
# ### Importing the required libraries
# +
import torch
import torchvision
import torch.nn as nn
import PIL
from PIL import Image
import numpy as np
import pandas as pd
import torchvision.models as models
import importlib
#Custom modules
import helping_material
importlib.reload(helping_material)
from helping_material.helper_fns import *
# -
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
from sklearn.neighbors import NearestNeighbors
import torchvision.transforms as transforms
# ### Making the dataset, and dataloader
# +
mean_nums = [0.485, 0.456, 0.406]
std_nums = [0.229, 0.224, 0.225]
trans=transforms.Compose([transforms.Resize((500,500)),
transforms.ToTensor(),
transforms.Normalize(mean_nums,std_nums)])
# -
images_path='data/5000-data/'
dataset=torchvision.datasets.ImageFolder(images_path,transform=trans)
dataset.classes
print(len(dataset))
train_dataloader=torch.utils.data.DataLoader(dataset,batch_size=4,shuffle=True)
#filenames = [s for s in train_dataloader.dataset.samples[0]]
filenames=[s for (_,_,s) in data_set_data_dir]
len(filenames)
# +
class ImageFolderWithPaths(torchvision.datasets.ImageFolder):
def __getitem__(self, index):
original_tuple = super(ImageFolderWithPaths, self).__getitem__(index)
path = self.imgs[index][0]
tuple_with_path = (original_tuple + (path,))
return tuple_with_path
dataset = ImageFolderWithPaths(root=images_path, transform=trans)
data_set_data_dir = torch.utils.data.DataLoader(dataset=dataset,batch_size=1)
for i, data in enumerate(data_set_data_dir):
images,labels,paths = data
print(paths[0])
if i==4:
break
# -
for i,(images,_) in enumerate(train_dataloader,0):
# show_tensor_images(images)
# sample_fname, _ = test_loader.dataset.samples[i]
print(i,train_dataloader.dataset.samples)
checking_loader=next(iter(train_dataloader))
#checking_loader[0].shape
show_tensor_images(checking_loader[0])
# * So, that we cna easily shift between cpu, and gpu
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# * Loading the pretrained model
resnet_model=models.resnet50(pretrained=True)
# ### Changing the model structure, for our use case
# * We can also use this technique
#
#
# model.classifier=nn.Sequential(*[model.classifier[i] for i in range(4)])
resnet_model.fc=nn.Identity()
resnet_model.to(device)
# +
#resnet_model
# -
# * Now using this model for getting the features of the images
# +
features_list=[]
with torch.no_grad():
for i,(images,_,_) in enumerate(data_set_data_dir):
images=images.to(device)
features=resnet_model(images)
if i%200==0:
print(features.shape)
for i in range(features.shape[0]):
features_list.append(features[i])
# -
print(len(features_list))
print(features_list[0].shape)
# * We need to have 2D array, for fitting the nearest neighbor algorith
features_list_array=np.array([features_list[i].cpu().numpy() for i in range(5000)])
features_list_array.shape
# ### Fitting Nearest-Neighbor algorithm to extracted features
neighbors=NearestNeighbors(n_neighbors=5,algorithm='ball_tree',metric='euclidean')
neighbors.fit(features_list_array)
# ### We need to have mapped filenames of all images, so that we can access them by indices, that are returned by Nearest-Neighbor Algorithm
# For this I have made Custom Dataloader, and got the names in filenames
filenames=filenames
# ### Now,we will extract the features of an image, not seen by model
test_img=Image.open('test_img3.jpeg')
test_img_t=trans(test_img)
show_tensor_images(test_img_t)
# * Getting the features
test_img_features=resnet_model(test_img_t.cuda().unsqueeze(0))
test_img_features.shape
# * Converting the features to numpy array
test_img_features_array=test_img_features.detach().cpu().numpy()
# * Getting the indices of similar images
_, indices = neighbors.kneighbors(test_img_features_array)
indices[0]
for i in indices[0]:
path=filenames[i]
print(path)
# * Showing the similar images
def similar_images(indices):
plt.figure(figsize=(15,10), facecolor='white')
plotnumber = 1
for index in indices:
if plotnumber<=len(indices) :
ax = plt.subplot(3,4,plotnumber)
plt.imshow(mpimg.imread(filenames[index][0]), interpolation='lanczos')
plotnumber+=1
plt.tight_layout()
similar_images(indices[0])
test_img_2=Image.open('TEST_TABLE.jpg')
test_img_t=trans(test_img_2)
show_tensor_images(test_img_t)
test_img_features=resnet_model(test_img_t.cuda().unsqueeze(0))
test_img_features_array=test_img_features.detach().cpu().numpy()
_, indices = neighbors.kneighbors(test_img_features_array)
similar_images(indices[0])
test_img_2=Image.open('TEST_BED.jpg')
test_img_t=trans(test_img_2)
show_tensor_images(test_img_t)
test_img_features=resnet_model(test_img_t.cuda().unsqueeze(0))
test_img_features_array=test_img_features.detach().cpu().numpy()
_, indices = neighbors.kneighbors(test_img_features_array)
similar_images(indices[0])
# ## So, it's pretty good that we are able to catch the complex features:
#
# * As for two seater, black sofa we are getting almost all two seaters
# * For table, we are getting 60% tables
# * And for bed, we are having 2 beds, that's bad
| Muhammad-Work/image_similarity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow 2.3 on Python 3.6 (CUDA 10.1)
# language: python
# name: python3
# ---
# + [markdown] id="UV3_mVndP6Hi"
# # 케라스로 심층 합성곱 신경망 만들기
# + [markdown] id="H2icE8zkP6Hm"
# 이 노트북에서 [LeNet-5](http://yann.lecun.com/exdb/publis/pdf/lecun-01a.pdf)과 비슷한 MNIST 손글씨 숫자를 분류하는 심층 합성곱 신경망을 만듭니다.
# + [markdown] id="G74ktsMRP6Hn"
# [](https://colab.research.google.com/github/rickiepark/dl-illustrated/blob/master/notebooks/10-1.lenet_in_keras.ipynb)
# + [markdown] id="ivEeMRHrP6Hn"
# #### 라이브러를 적재합니다.
# + id="ma9EMkCAP6Hn"
from tensorflow import keras
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout
from tensorflow.keras.layers import Flatten, Conv2D, MaxPooling2D # new!
# + [markdown] id="SsTnIg6aP6Ho"
# #### 데이터를 적재합니다.
# + id="NRkMN0KJP6Ho" outputId="b2e94361-70f7-4e76-8d30-6c196f7d04f1" colab={"base_uri": "https://localhost:8080/"}
(X_train, y_train), (X_valid, y_valid) = mnist.load_data()
# + [markdown] id="Hkf9icAJP6Hp"
# #### 데이터를 전처리합니다.
# + id="S_aWRMslP6Hp"
X_train = X_train.reshape(60000, 28, 28, 1).astype('float32')
X_valid = X_valid.reshape(10000, 28, 28, 1).astype('float32')
# + id="se51PfthP6Hp"
X_train /= 255
X_valid /= 255
# + id="KUE_VeFiP6Hp"
n_classes = 10
y_train = keras.utils.to_categorical(y_train, n_classes)
y_valid = keras.utils.to_categorical(y_valid, n_classes)
# + [markdown] id="5aE2huiNP6Hp"
# #### 신경망 모델을 만듭니다.
# + id="ozZ3asHxP6Hq"
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', input_shape=(28, 28, 1)))
model.add(Conv2D(64, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(n_classes, activation='softmax'))
# + id="A3JA2ahzP6Hq" outputId="cf0fc04e-d3e3-41e0-89b0-5c8a37e0ec6b" colab={"base_uri": "https://localhost:8080/"}
model.summary()
# + [markdown] id="RqMEQz_rP6Hq"
# #### 모델을 설정합니다.
# + id="xevWEuNDP6Hq"
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# + [markdown] id="Sa8I9xhCP6Hq"
# #### 훈련!
# + id="bsLCVNMPP6Hq" outputId="a9d85d71-687e-457d-c455-4e24611bcc07" colab={"base_uri": "https://localhost:8080/"}
model.fit(X_train, y_train, batch_size=128, epochs=10, verbose=1, validation_data=(X_valid, y_valid))
| notebooks/10-1.lenet_in_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
with open('Donline_lastepisode_heat.pickle', 'rb') as f:
last_heat = pickle.load(f)
with open('Donline_heat_unique0.pickle', 'rb') as f:
heat_uniq0 = pickle.load(f)
with open('Donline_heat_freq0.pickle', 'rb') as f:
heat_freq0 = pickle.load(f)
with open('Donline_heat_unique1.pickle', 'rb') as f:
heat_uniq1 = pickle.load(f)
with open('Donline_heat_freq1.pickle', 'rb') as f:
heat_freq1 = pickle.load(f)
# +
# with open('deep_random_corr0.pickle', 'rb') as f:
# rand_corr0 = pickle.load(f)
# with open('deep_random_corr1.pickle', 'rb') as f:
# rand_corr1 = pickle.load(f)
# with open('deep_recent_corr0.pickle', 'rb') as f:
# recen_corr0 = pickle.load(f)
# with open('deep_recent_corr1.pickle', 'rb') as f:
# recen_corr1 = pickle.load(f)
# -
num_episodes = len(heat_freq0)
num_actions = 15
num_sub = 500
np.unique(last_heat[:, 0, :, :], return_counts=True)
np.unique(last_heat[:, 0, :, :], return_counts=True)[1]/num_sub/num_actions**2
np.unique(last_heat[:, 1, :, :], return_counts=True)
np.unique(last_heat[:, 1, :, :], return_counts=True)[1]/num_sub/num_actions**2
np.sum(last_heat[:, 0, 0, 0] - last_heat[:, 1, 0, 0] < 0)/num_sub
np.sum(last_heat[:, 0, 0, 0] - last_heat[:, 1, 0, 0] == 0)/num_sub
np.sum(last_heat[:, 0, 0, 0] - last_heat[:, 1, 0, 0] > 0)/num_sub
plt.figure(figsize=(8, 6))
ax = sns.heatmap(last_heat[-1, 0, :, :], cbar=False, annot=True)
plt.xlabel('Player 1')
plt.ylabel('Player 0')
cbar = ax.figure.colorbar(ax.collections[0])
cbar.set_ticks([0, 2, 4, 6, 8, 10, 12, 14])
fig = ax.get_figure()
# fig.savefig('hybrid_heat0.eps', format='eps', dpi=500, bbox_inches='tight', pad_inches=0.1)
full_freq0 = np.zeros((num_episodes, num_actions))
for i in range(num_episodes):
full_freq0[i, heat_uniq0[i].astype(int)] = heat_freq0[i]
len(heat_uniq0[1000])
var = np.zeros(num_episodes)
for i in range(num_episodes):
var[i] = len(heat_uniq0[i])
plt.plot(var)
np.argmax(np.sum(full_freq0, axis=0))
max_price = np.zeros(num_episodes)
max_freq = np.zeros(num_episodes)
bottom8_freq = np.zeros(num_episodes)
bottom3_freq = np.zeros(num_episodes)
for i in range(num_episodes):
max_price[i] = np.max(heat_uniq0[i])
max_freq[i] = np.argmax(full_freq0[i, :])
bottom8_freq[i] = np.sum(full_freq0[i, :8])
bottom3_freq[i] = np.sum(full_freq0[i, :3])
fig, ax = plt.subplots(figsize=(14, 5), dpi=120)
ax.plot(bottom8_freq/112500, color='tab:blue', label=r'Price $\leq$ 1.71')
ax.plot(bottom3_freq/112500, color='tab:orange', label =r'Price $\leq$ 1.51')
ax.set_ylabel('Percent')
ax.set_xlabel('Episodes')
ax.legend(loc='best')
ax.grid(True)
# plt.savefig('bottom.eps', format='eps', dpi=1000, bbox_inches='tight', pad_inches=0.1)
plt.show()
np.unique(max_price, return_counts=True)
fig, ax = plt.subplots(figsize=(8, 6), dpi=120)
ax.plot(1.43 + 0.04*max_price, color='tab:blue', label='Highest')
ax.plot(1.43 + 0.04*max_freq, color='tab:orange', label ='Most frequent')
ax.set_ylabel('Price')
ax.yaxis.set_ticks(np.arange(1.43, 2.0, 0.04))
ax.set_xlabel('Episodes')
ax.legend(loc='best')
ax.grid(True)
plt.savefig('recent_max.eps', format='eps', dpi=1000, bbox_inches='tight', pad_inches=0.1)
plt.show()
full_freq1 = np.zeros((num_episodes, num_actions))
for i in range(num_episodes):
full_freq1[i, heat_uniq1[i].astype(int)] = heat_freq1[i]
max_price1 = np.zeros(num_episodes)
max_freq1 = np.zeros(num_episodes)
bottom8_freq1 = np.zeros(num_episodes)
bottom3_freq1 = np.zeros(num_episodes)
for i in range(num_episodes):
max_price1[i] = np.max(heat_uniq1[i])
max_freq1[i] = np.argmax(full_freq1[i, :])
bottom8_freq1[i] = np.sum(full_freq1[i, :8])
bottom3_freq1[i] = np.sum(full_freq1[i, :3])
fig, ax = plt.subplots(figsize=(18, 6), dpi=120)
ax.plot(bottom8_freq1/112500, color='tab:blue', label=r'Price $\leq$ 1.71')
ax.plot(bottom3_freq1/112500, color='tab:orange', label =r'Price $\leq$ 1.51')
ax.set_ylabel('Percent')
ax.set_xlabel('Episodes')
ax.legend(loc='best')
ax.grid(True)
# plt.savefig('.eps', format='eps', dpi=500, bbox_inches='tight', pad_inches=0.1)
plt.show()
fig, ax = plt.subplots(figsize=(8, 6), dpi=120)
ax.plot(1.43 + 0.04*max_price1, color='tab:blue', label='Highest')
ax.plot(1.43 + 0.04*max_freq1, color='tab:orange', label ='Most frequent')
ax.set_ylabel('Price')
ax.yaxis.set_ticks(np.arange(1.43, 2.0, 0.04))
ax.set_xlabel('Episodes')
ax.legend(loc='best')
ax.grid(True)
# plt.savefig('.eps', format='eps', dpi=500, bbox_inches='tight', pad_inches=0.1)
plt.show()
-3%8
ind
# +
N = 2000 - 30
ind = np.arange(N+1, N+31)
width = 0.5
# plt.style.use('default')
cm = plt.get_cmap('tab20')
plt.rcParams["axes.prop_cycle"] = plt.cycler('color', [cm(1.*i/num_actions) for i in range(num_actions)])
p = []
fig, ax = plt.subplots(figsize=(8,6), dpi=120)
for k in range(num_actions):
p.append(plt.bar(ind, full_freq0[N:N+30, k]/112500, width, bottom = np.sum(full_freq0[N:N+30, :k], axis=1)/112500))
plt.legend((p[0][0], p[1][0], p[2][0], p[3][0], p[4][0], p[5][0], p[6][0], p[7][0],
p[8][0], p[9][0], p[10][0], p[11][0], p[12][0], p[13][0], p[14][0]),
('1.43', '1.47', '1.51', '1.55', '1.59', '1.63', '1.67', '1.71',
'1.75', '1.79', '1.83', '1.87', '1.91', '1.95', '1.99'), bbox_to_anchor=(1.0, 1.0))
plt.xticks(ind)
plt.xticks(rotation=70)
ax.set_xlabel('Episodes')
ax.set_ylabel('Percent')
# plt.savefig('begin_bar.eps', format='eps', dpi=1000, bbox_inches='tight', pad_inches=0.1)
plt.show()
# -
fig, ax = plt.subplots(figsize=(8, 6), dpi=120)
ax.hist(rand_corr0, bins=200)
# ax.plot(bottom3_freq/112500, color='tab:orange', label =r'Price $\leq$ 1.51')
ax.set_ylabel('Frequency')
ax.set_xlabel('Correlation')
# ax.legend(loc='best')
# ax.grid(True)
# plt.savefig('rand_corr.eps', format='eps', dpi=1000, bbox_inches='tight', pad_inches=0.1)
plt.show()
fig, ax = plt.subplots(figsize=(8, 6), dpi=120)
ax.hist(recen_corr0, bins=200)
ax.set_ylabel('Frequency')
ax.set_xlabel('Correlation')
# ax.legend(loc='best')
# ax.grid(True)
# plt.savefig('recen_corr.eps', format='eps', dpi=1000, bbox_inches='tight', pad_inches=0.1)
plt.show()
fig, ax = plt.subplots(figsize=(14, 5), dpi=120)
ax.plot(recen_corr0, color='tab:blue')
# ax.plot(rand_corr0, 'o', markersize=2, color='tab:orange')
ax.set_ylabel('Correlation')
ax.set_xlabel('Iterations')
# ax.grid(True)
# plt.savefig('rand_corr_series.eps', format='eps', dpi=1000, bbox_inches='tight', pad_inches=0.1)
plt.show()
fig, ax = plt.subplots(figsize=(8, 6), dpi=120)
ax.plot(rand_corr0[:10000], 'o', markersize=1, color='tab:orange', label='Random')
ax.plot(recen_corr0[:10000], 'o', markersize=1, color='tab:blue', label='Recent')
ax.legend(loc='best')
ax.set_ylabel('Correlation')
ax.set_xlabel('Iterations')
# ax.grid(True)
# plt.savefig('corr_series_begin.eps', format='eps', dpi=200, bbox_inches='tight', pad_inches=0.1)
plt.show()
fig, ax = plt.subplots(figsize=(8, 6), dpi=120)
x_ticks = np.arange(len(recen_corr0) - 10000, len(recen_corr0))
ax.plot(x_ticks, rand_corr0[-10000:], 'o', markersize=1, color='tab:orange', label='Random')
ax.plot(x_ticks, recen_corr0[-10000:], 'o', markersize=1, color='tab:blue', label='Recent')
ax.legend(loc='best')
ax.set_ylabel('Correlation')
ax.set_xlabel('Iterations')
# ax.grid(True)
# plt.savefig('corr_series_end.eps', format='eps', dpi=1000, bbox_inches='tight', pad_inches=0.1)
plt.show()
| figs/Donline_figs_not_in_paper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Determine global irrigation topology
# ## which HRUs are served by a reservoir
#
# <NAME> - March 2021
#
# +
# import modules
# add path where utils modules are located to python path
import numpy as np
import geopandas as gpd
import pandas as pd
import pickle
import warnings
warnings.filterwarnings('ignore')
# import own functions
import pfaf.pfafstetter as pfaf # decode package from Naoki, see https://github.com/nmizukami/pfaf_decode
from utils_plotting import *
from utils_irrigtopo import *
# define data directory
data_dir = 'C:\\Users\\ivand\\OneDrive - Vrije Universiteit Brussel/PhD/3_reservoir_release/data_for_mizuroute/'
# Settings:
outlet_threshold = 700e3 # in m
tributary_threshold = 100e3 # m
# calculate or load reservoir dependency
calc_res_dependency = True # if True: Calculate, if False: load
# -
# ## Calculate or load reservoir dependencies
# +
# %%time
if calc_res_dependency:
# load global river segments
river_grand_attrs = gpd.read_file(data_dir+'river_with_lake_flag4_reorder/river_with_grand.shp')
pfaf_reservoirs = list(river_grand_attrs.loc[river_grand_attrs['H06_purpos']==1,'PFAF'].values)
river_shp = river_grand_attrs
# get their corresponding outlets
pfaf_outlets = get_outlets(river_shp,pfaf_reservoirs,outlet_threshold)
# get dependent reservoirs per segment
seg_dependency_dict = get_seg_dependency(pfaf_reservoirs, pfaf_outlets, river_shp, tributary_threshold)
# get weights of dependend reservoirs per segment
weights_dict = get_weights_per_seg(seg_dependency_dict, river_shp, pfaf_reservoirs, weigh_smax_with_nseg=True)
# get depenent segments per reservoir and corresponding weights
res_dependency_dict = get_res_dependency_and_weights(pfaf_reservoirs, seg_dependency_dict, weights_dict)
# save dependency dict as a pickle
f = open(data_dir+"irrigation_topology/res_dependency_HDMA_outletthres_"+str(int(outlet_threshold*10**-3))+"km_tribthres_"+str(int(tributary_threshold*10**-3))+"km.pkl","wb")
pickle.dump(res_dependency_dict,f)
f.close()
f = open(data_dir+"irrigation_topology/seg_dependency_HDMA_outletthres_"+str(int(outlet_threshold*10**-3))+"km_tribthres_"+str(int(tributary_threshold*10**-3))+"km.pkl","wb")
pickle.dump(seg_dependency_dict,f)
f.close()
else:
print('load reservoir and segment depenency')
print('with outlet threshold of '+str(int(outlet_threshold*10**-3))+' km and tributary threshold of '+str(int(tributary_threshold*10**-3))+' km')
# open res_dependency pickle
with open(data_dir+"/irrigation_topology/res_dependency_HDMA_outletthres_"+str(int(outlet_threshold*10**-3))+"km_tribthres_"+str(int(tributary_threshold*10**-3))+"km.pkl", 'rb') as handle:
res_dependency_dict = pickle.load(handle)
with open(data_dir+"/irrigation_topology/seg_dependency_HDMA_outletthres_"+str(int(outlet_threshold*10**-3))+"km_tribthres_"+str(int(tributary_threshold*10**-3))+"km.pkl", 'rb') as handle:
seg_dependency_dict = pickle.load(handle)
# -
# ## Apply topology on HRUs (for now only simplified ones)
# +
# laod HRUS
hru_simplify = gpd.read_file("C:\\Users\\ivand\\OneDrive - Vrije Universiteit Brussel/PhD/3_reservoir_release/water_demand/mizuRoute_data/HDMA_catchment/hdma_global_catch_simplify.gpkg")
hrus_shp = hru_simplify
# calculate number of dependend reservoirs per hru.
def count_res(row):
pfaf = row.PFAF
if pfaf in seg_dependency_dict.keys():
return len(seg_dependency_dict[pfaf])
else:
return 0
hrus_shp['n_res'] = hrus_shp.apply(lambda row: count_res(row),axis=1)
hrus_shp.to_file("C:\\Users\\ivand\\OneDrive - Vrije Universiteit Brussel/PhD/3_reservoir_release/water_demand/mizuRoute_data/HDMA_catchment/hru_simplify_nres.gpkg", driver="GPKG")
# -
| preprocessing/determine_irrigtopo_global.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://github.com/giswqs/gee-tutorials/blob/master/Image/image_overview.ipynb)
# **Image Overview**
#
# Raster data are represented as `Image` objects in Earth Engine. Images are composed of one or more bands and each band has its own name, data type, scale, mask and projection. Each image has metadata stored as a set of properties.
#
# In addition to loading images from the archive by an image ID, you can also create images from constants, lists or other suitable Earth Engine objects. The following illustrates methods for creating images, getting band subsets, and manipulating bands.
#
# More information about ee.Image can be found in the [Earth Engine Documentation](https://developers.google.com/earth-engine/guides/image_overview).
# +
# # !pip isntall geemap
# -
import ee
import geemap
# ## Loading a single-band image
#
# Images can be loaded by pasting an Earth Engine asset ID into the ee.Image constructor. You can find image IDs in the [data catalog](https://developers.google.com/earth-engine/datasets). For example, to load SRTM Digital Elevation Data:
Map = geemap.Map(center=(40, -100), zoom=4)
Map
dem = ee.Image('CGIAR/SRTM90_V4')
Map.addLayer(dem, {}, "DEM")
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
Map.addLayer(dem, vis_params, "DEM Vis")
# ## Loading a multi-band image
# +
Map = geemap.Map()
# Load an image.
image = ee.Image('LANDSAT/LC08/C01/T1_SR/LC08_044034_20140318')
# Center the map and display the image.
Map.centerObject(image, zoom=8)
Map.addLayer(image, {}, 'Landsat')
Map
# -
vis_params = {'bands': ['B5', 'B4', 'B3'], 'min': 0.0, 'max': 3000, 'opacity': 1.0, 'gamma': 1.2}
Map.addLayer(image, vis_params, "Landsat Vis")
# ## Getting image properties
image = ee.Image('LANDSAT/LC08/C01/T1_SR/LC08_044034_20140318')
props = geemap.image_props(image)
props.getInfo()
# ## Selecting image bands
image = ee.Image('LANDSAT/LC08/C01/T1_SR/LC08_044034_20140318')
bands = image.select(['B5', 'B4', 'B3'])
Map.addLayer(bands, vis_params, 'Landsat B543')
Map
# ## Renaming band names
image = ee.Image('LANDSAT/LC08/C01/T1_SR/LC08_044034_20140318')
new_image = image.select(['B5', 'B4', 'B3'], ['NIR', 'Red', 'Green'])
band_names = new_image.bandNames()
band_names.getInfo()
# ## Adding a legend
Map = geemap.Map()
Map.add_basemap('HYBRID')
Map
landcover = ee.Image('USGS/NLCD/NLCD2016').select('landcover')
Map.addLayer(landcover, {}, 'Land Cover')
Map.add_legend(builtin_legend='NLCD', layer_name='Land Cover')
| docs/Image/image_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
from datascience import *
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
plt.style.use('ggplot')
# # Intro to Regression
#
# A popular classification model is [logistic regression](https://en.wikipedia.org/wiki/Logistic_regression). This is what Underwood and Sellers use in their article to classify whether a text was reviewed or randomly selected from HathiTrust. Today we'll look at the difference between regression and classification tasks, and how we can use a logistic regression model to classify text like Underwood and Sellers. We won't have time to go through their full code, but if you're interested I've provided a walk-through in the second notebook.
#
# To explore the regression model let's first create some dummy data:
demo_tb = Table()
demo_tb['Study_Hours'] = [2.0, 6.9, 1.6, 7.8, 3.1, 5.8, 3.4, 8.5, 6.7, 1.6, 8.6, 3.4, 9.4, 5.6, 9.6, 3.2, 3.5, 5.9, 9.7, 6.5]
demo_tb['Grade'] = [67.0, 83.6, 35.4, 79.2, 42.4, 98.2, 67.6, 84.0, 93.8, 64.4, 100.0, 61.6, 100.0, 98.4, 98.4, 41.8, 72.0, 48.6, 90.8, 100.0]
demo_tb['Pass'] = [0, 1, 0, 1, 0, 1, 0, 1, 1, 0, 1, 0, 1, 1, 1, 0, 1, 0, 1, 1]
demo_tb.show()
# ## Intuiting the Linear Regression Model
#
# You may have encountered linear regression in previous coursework of yours. Linear regression, in its simple form, tries to model the relationship between two continous variables as a straight line. It interprets one variable as the input, and the other as the output.:
demo_tb.scatter('Study_Hours','Grade')
# In the example above, we're interested in `Study_Hours` and `Grade`. This is a natural "input" "output" situation. To plot the regression line, or ***best-fit***, we can feed in `fit_line=True` to the `scatter` method:
demo_tb.scatter('Study_Hours','Grade', fit_line=True)
# The better this line fits the points, the better we can predict one's `Grade` based on their `Study_Hours`, even if we've never seen anyone put in that number of study hours before.
#
# The regression model above can be expressed as:
#
# $GRADE_i= \alpha + \beta STUDYHOURS + \epsilon_i$
#
# The variable we want to predict (or model) is the left side `y` variable, here `GRADE`. The variable which we think has an influence on our left side variable is on the right side, the independent variable `STUDYHOURS`. The $\alpha$ term is the y-intercept and the $\epsilon_i$ describes the randomness.
#
# The $\beta$ coefficient on `STUDYHOURS` gives us the slope, in a univariate regression. That's the factor on `STUDYHOURS` to get `GRADE`.
#
# If we want to build a model for the regression, we can use the `sklearn` library. `sklearn` is by far the most popular machine learning library for Python, and its syntax is really important to learn. In the next cell we'll import the [`Linear Regression`](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html) model and assign it to a `linreg` variable:
from sklearn.linear_model import LinearRegression
linreg = LinearRegression()
# Before we go any further, `sklearn` likes our data in a very specific format. The `X` must be in an array of arrays, each sub array is an observation. Because we only have one independent variable, we'll have sub arrays of `len` 1. We can do that with the `reshape` method:
X = demo_tb['Study_Hours'].reshape(-1,1)
X
# Your output, or dependent variable, is just one array with no sub arrays.
y = demo_tb['Grade'].reshape(len(demo_tb['Grade']),)
y
# We then use the `fit` method to fit the model. This happens in-place, so we don't have to reassign the variable:
linreg.fit(X, y)
# We can get back the `intercept_` and $\beta$ `coef_` with attributes of the `linreg` object:
B0, B1 = linreg.intercept_, linreg.coef_[0]
B0, B1
# So this means:
#
# $GRADE_i= 42.897229302892598 + 5.9331153718275509 * STUDYHOURS + \epsilon_i$
#
# As a linear regression this is simple to interpret. To get our grade score, we take the number of study hours and multipy it by 5.9331153718275509 then we add 42.897229302892598 and that's our prediction.
#
# If we look at our chart again but using the model we just made, that looks about right:
y_pred = linreg.predict(X)
print(X)
print(y_pred)
plt.scatter(X, y)
plt.plot(X, y_pred)
# We can evaluate how great our model is with the `score` method. We need to give it the `X` and observed `y` values, and it will predict its own `y` values and compare:
linreg.score(X, y)
# For the Linear Regression, `sklearn` returns an **R-squared** from the `score` method. The R-squared tells us how much of the variation in the data can be explained by our model, .559 isn't that bad, but obviously more goes into your `Grade` than *just* `Study_Hours`.
#
# Nevertheless we can still predict a grade just like we did above to create that line, let's say I studied for 5 hours:
linreg.predict([[5]])
# Maybe I should study more?
linreg.predict([[20]])
# Wow! I rocked it.
# ## Intuiting the Logistic Regression Model
#
# But what happens if one of your variables is categorical, and not continuous? Suppose we don't care about the `Grade` score, but we just care if you `Pass` or not:
demo_tb.scatter('Study_Hours','Pass')
# How would we fit a line to that? That's where the [logistic function](https://en.wikipedia.org/wiki/Logistic_function) can be handy. The general logistic function is:
#
# $ f(x) = \frac{1}{1 + e^{-x}} $
#
# We can translate that to Python:
def logistic(p):
return 1 / (1 + np.exp(-p))
# We'll also need to assign a couple $\beta$ coefficients for the intercept and variable just like we saw in linear regression:
B0, B1 = 0, 1
# Let's plot the logistic curve:
# +
xmin, xmax = -10,10
xlist = [float(x)/int(1e4) for x in range(xmin*int(1e4), xmax*int(1e4))] # just a lot of points on the x-axis
ylist = [logistic(B0 + B1*x) for x in xlist]
plt.axis([-10, 10, -0.1,1.1])
plt.plot(xlist,ylist)
# -
# When things get complicated, however, with several independent variables, we don't want to write our own code. Someone has done that for us. We'll go back to `sklearn`.
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
# We'll `reshape` our arrays again too, since we know how `sklearn` likes them:
X = demo_tb['Study_Hours'].reshape(-1,1)
y = demo_tb['Pass'].reshape(len(demo_tb['Pass']),)
X, y
# We can use the `fit` function again on our `X` and `y`:
lr.fit(X, y)
# We can get those $\beta$ coefficients back out from `sklearn` for our grade data:
B0, B1 = lr.intercept_[0], lr.coef_[0][0]
B0, B1
# Then we can plot the curve just like we did earlier, and we'll add our points:
# +
xmin, xmax = 0,10
xlist = [float(x)/int(1e4) for x in range(xmin*int(1e4), xmax*int(1e4))]
ylist = [logistic(B0 + B1*x) for x in xlist]
plt.plot(xlist,ylist)
# add our "observed" data points
plt.scatter(demo_tb['Study_Hours'],demo_tb['Pass'])
# -
# How might this curve be used for a binary classification task?
# + active=""
#
# -
# ## Logistic Classification
#
# That's great, so we can begin to see how we might use such a model to conduct binary classification. In this task, we want to get a number of study hours as an observation, and place it in one of two bins: pass or fail.
#
# To create the model though, we have to train it on the data we have. [In machine learning, we also need to put some data aside as "testing data" so that we don't bias our model by using it in the training process.](https://en.wikipedia.org/wiki/Training,_test,_and_validation_sets) In Python we often see `X_train`, `y_train` and `X_test`, `y_test`:
# +
X_train = demo_tb.column('Study_Hours')[:-2]
y_train = demo_tb.column('Pass')[:-2]
X_test = demo_tb.column('Study_Hours')[-2:]
y_test = demo_tb.column('Pass')[-2:]
# -
# Let's see the observations we're setting aside for later:
print(X_test, y_test)
# Now we'll fit our model again but only on the `_train` data, and get out the $\beta$ coefficients:
lr.fit(X_train.reshape(-1,1),y_train.reshape(len(y_train),))
B0, B1 = lr.intercept_[0], lr.coef_[0]
# We can send these coefficients back into the `logistic` function we wrote earlier to get the probability that a student would pass given our `X_test` values:
fitted = [logistic(B1*th + B0) for th in X_test]
fitted
# We can take the probability and change this to a binary outcome based on probability `>` or `<` .5:
prediction = [pred >.5 for pred in fitted]
prediction
# The `sklearn` built-in methods can make this `predict` process faster:
lr.predict(X_test.reshape(-1, 1))
# To see how accurate our model is, we'd predict on the "unseen" `_test`ing data and see how many we got correct. In this case there's only two, so not a whole lot to test with:
prediction_eval = [prediction[i]==y_test[i] for i in range(len(prediction))]
float(sum(prediction_eval)/len(prediction_eval))
# We can do this quickly in `sklearn` too with the `score` method like in the linear regression example:
lr.score(X_test.reshape(-1, 1), y_test.reshape(len(y_test),))
# ---
#
# # Classification of Textual Data
#
# How can we translate this simple model of binary classification to text? I'm going to leave the more complicated model that Underwood and Sellers use for the next notebook if you're interested, today we're just going to work through the basic classfication pipeline. We'll download a pre-made corpus from `nltk`:
import nltk
nltk.download("movie_reviews")
# Now we import the `movie_reviews` object:
from nltk.corpus import movie_reviews
# As you might expect, this is a corpus of IMDB movie reviews. Someone went through and read each review, labeling it as either "positive" or "negative". The task we have before us is to create a model that can accurately predict whether a never-before-seen review is positive or negative. This is analogous to Underwood and Sellers looking at whether a poem volume was reviewed or randomly selected.
#
# From the `movie_reviews` object let's take out the reviews and the judgement:
reviews = [movie_reviews.raw(fileid) for fileid in movie_reviews.fileids()]
judgements = [movie_reviews.categories(fileid)[0] for fileid in movie_reviews.fileids()]
# Let's read the first review:
print(reviews[0])
# Do you consider this a positive or negative review? Let's see what the human annotator said:
print(judgements[0])
# So right now we have a list of movie reviews in the `reviews` variable and a list of their corresponding judgements in the `judgements` variable. Awesome. What does this sound like to you? Independent and dependent variables? You'd be right!
#
# `reviews` is our `X` array from above. `judgements` is our `y` array from above. Let's first reassign our `X` and `y` so we're explicit about what's going on. While we're at it, we're going to set the random `seed` for our computer. This just makes our result reproducible. We'll also `shuffle` so that we randomize the order of our observations, and when we split the testing and training data it won't be in a biased order:
# +
from sklearn.utils import shuffle
np.random.seed(1)
X, y = shuffle(reviews, judgements, random_state=0)
# -
# If you don't believe me that all we did is reassign and shuffle:
X[0], y[0]
# To get meaningful independent variables (words) we have to do some processing too (think DTM!). With `sklearn`'s text pipelines, we can quickly build a text a classifier in only a few lines of Python:
# +
# LogisticRegression?
# +
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer, TfidfTransformer
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score, train_test_split
text_clf = Pipeline([('vect', CountVectorizer(ngram_range=(1, 2))),
('tfidf', TfidfTransformer()),
('clf', LogisticRegression(random_state=0, penalty='l2', C=1000))
])
scores = cross_val_score(text_clf, X, y, cv=5)
print(scores, np.mean(scores))
# -
# ***Whoa! What just happened?!?*** The pipeline tells us three things happened:
#
# 1. `CountVectorizer`
#
# 2. `TfidfTransformer`
#
# 3. `LogisticRegression`
#
# Let's walk through this step by step.
#
# 1. A count vectorizer does exactly what we did last week with tokenization. It changes all the texts to words, and then simply counts the frequency of each word occuring in the corpus for each document. The feature array for each document at this point is simply the length of all unique words in a corpus, with the count for the frequency of each. This is the most basic way to provide features for a classifier---a document term matrix.
#
# 2. [tfidf](https://en.wikipedia.org/wiki/Tf%E2%80%93idf) (term frequency inverse document frequency) is an algorithm that aims to find words that are important to specific documents. It does this by taking the term frequency (tf) for a specific term in a specific document, and multiplying it by the term's inverse document frequency (idf), which is the total number of documents divided by the number of documents that contain the term at least once. Thus, idf is defined as:
#
# $$idf(t, d, D)= log\left(\frac{\mid D \mid}{\mid \{d \subset D : t \subset d \} \mid}\right )$$
#
# So tfidf is simply:
#
# $$tfidf(t, d, D)= f_{t,d}*log\left(\frac{\mid D \mid}{\mid \{d \subset D : t \subset d \} \mid}\right )$$
#
# A tfidf value is calculated for each term for each document. The feature arrays for a document is now the tfidf values. ***The tfidf matrix is the exact same as our document term matrix, only now the values have been weighted according to their distribution across documents.***
#
# The pipeline now sends these tfidf feature arrays to a 3. **Logistic Regression**, what we learned above. We add in an l2 penalization parameter because we have many more independent variables from our `dtm` than observations. The independent variables are the tfidf values of each word. In a simple linear model, that would look like:
#
# $$log(CLASSIFICATION_i)= \alpha + \beta DOG + \beta RABBIT + \beta JUMP + ... + \epsilon_i$$
#
# where $\beta DOG$ is the model's $\beta$ coefficient multiplied by the ***tfidf*** value for "dog".
#
# The code below breaks this down by each step, but combines the `CountVectorizer` and `TfidfTransformer` in the `TfidfVectorizer`.
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=50)
# get tfidf values
tfidf = TfidfVectorizer()
tfidf.fit(X)
X_train = tfidf.transform(X_train)
X_test = tfidf.transform(X_test)
# build and test logit
logit_class = LogisticRegression(random_state=0, penalty='l2', C=1000)
model = logit_class.fit(X_train, y_train)
model.score(X_test, y_test)
# -
# The concise code we first ran actually uses "cross validation", where we split up testing and training data `k` number of times and average our score on all of them. This is a more reliable metric than just testing the accuracy once. It's possible that you're random train/test split just didn't provide a good split, so averaging it over multiple splits is preferred.
#
# You'll also notice the `ngram_range` parameter in the `CountVectorizer` in the first cell. This expands our vocabulary document term matrix by including groups of words together. It's easier to understand an [ngram](https://en.wikipedia.org/wiki/N-gram) by just seeing one. We'll look at a bigram (bi is for 2):
# +
from nltk.util import ngrams
ngs = ngrams("Text analysis is so cool. I can really see why classification can be a valuable tool.".split(), 2)
list(ngs)
# -
# Trigram:
ngs = ngrams("Text analysis is so cool. I can really see why classification can be a valuable tool.".split(), 3)
list(ngs)
# You get the point. This helps us combat this "bag of words" idea, but doesn't completely save us. For our purposes here, just as we counted the frequency of individual words, we've added counting the frequency of groups of 2s and 3s.
#
# ---
#
# ### Important Features
#
# After we train the model we can then index the tfidf matrix for the words with the most significant coefficients (remember independent variables!) to get the most helpful features:
feature_names = tfidf.get_feature_names()
top10pos = np.argsort(model.coef_[0])[-10:]
print("Top features for positive reviews:")
print(list(feature_names[j] for j in top10pos))
print()
print("Top features for negative reviews:")
top10neg = np.argsort(model.coef_[0])[:10]
print(list(feature_names[j] for j in top10neg))
# ### Prediction
#
# We can also use our model to classify new reviews, all we have to do is extract the tfidf features from the raw text and send them to the model as our features (independent variables):
# +
new_bad_review = "This movie really sucked. I can't believe how long it dragged on. The actors are absolutely terrible. They should rethink their career paths"
features = tfidf.transform([new_bad_review])
model.predict(features)
# +
new_good_review = "I loved this film! The cinematography was incredible, and <NAME> is flawless. Super cute BTW."
features = tfidf.transform([new_good_review])
model.predict(features)
# -
# # Homework
#
# Let's examine more the three objects in the pipeline:
# +
# CountVectorizer?
# +
# TfidfTransformer?
# +
# LogisticRegression?
# -
# I've copied the cell from above below. Try playing with the parameters to these objects and see if you can improve the `cross_val_score` for the model.
# +
text_clf = Pipeline([('vect', CountVectorizer(ngram_range=(1, 2))),
('tfidf', TfidfTransformer()),
('clf', LogisticRegression(random_state=0))
])
scores = cross_val_score(text_clf, X, y, cv=5)
print(scores, np.mean(scores))
# -
# Why do you think your score improved (or didn't)?
# + active=""
#
# -
# ---
#
# # BONUS (not assigned)
#
# We're going to download the [20 Newsgroups](http://qwone.com/~jason/20Newsgroups/), a widely used corpus for demos of general texts:
#
# > The 20 Newsgroups data set is a collection of approximately 20,000 newsgroup documents, partitioned (nearly) evenly across 20 different newsgroups. To the best of my knowledge, it was originally collected by <NAME>, probably for his Newsweeder: Learning to filter netnews paper, though he does not explicitly mention this collection. The 20 newsgroups collection has become a popular data set for experiments in text applications of machine learning techniques, such as text classification and text clustering.
# First we'll import the data from `sklearn`:
from sklearn.datasets import fetch_20newsgroups
# Let's see what categories they have:
fetch_20newsgroups(subset="train").target_names
# The `subset` parameter will give you training and testing data. You can also use the `categories` parameter to choose only certain categories.
#
# If we wanted to get the training data for `sci.electronics` and `rec.autos` we would write this:
train = fetch_20newsgroups(subset="train", categories=['sci.electronics', 'rec.autos'])
# The list of documents (strings) is in the `.data` property, we can access the first one like so:
train.data[0]
# And here is the assigment category:
train.target[0]
# How many training documents are there?
len(train.data)
# We can do the same for the testing data:
test = fetch_20newsgroups(subset="test", categories=['sci.electronics', 'rec.autos'])
test.data[0]
test.target[0]
len(test.data)
# You now have your four lists:
#
# - `train.data` = `X_train` list of strings
# - `train.target` = `y_train` list of category assignments
# - `test.data` = `X_test` list of strings
# - `test.target` = `y_test` list of category assignments
#
# Build a classifier below. And then choose different categories and rebuild it. Which categories are easier to classify from each other? Why?
#
# ***TIP***: Don't forget to `shuffle` your data!
| 08-Classification/01-Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Hank-Cui/otis2019/blob/crawler/%E4%B8%80%E4%BA%9B%E7%88%AC%E8%99%AB.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="iU9ubOm7UKfT" colab_type="code" colab={}
#Install yfinance
# %pip install yfinance
# %pip install selenium
# + id="twfcBpDoxmn0" colab_type="code" colab={}
import time
import requests
from bs4 import BeautifulSoup
from selenium.webdriver.common.keys import Keys
import pandas as pd
import numpy as np
from pandas_datareader import data
import matplotlib.pyplot as plt
import yfinance as yf
# + id="hRvscfxD3kLG" colab_type="code" colab={}
from google.colab import drive
drive.mount('/gdrive')
# + id="yN25Jvc2b9Gv" colab_type="code" colab={}
# Selenium Colab
# 暂时不用了
install chromium, its driver, and selenium:
# !pip install selenium
# !apt-get update # to update ubuntu to correctly run apt install
# !apt install chromium-chromedriver
# !cp /usr/lib/chromium-browser/chromedriver /usr/bin
import sys
sys.path.insert(0,'/usr/lib/chromium-browser/chromedriver')
from selenium import webdriver
chrome_options = webdriver.ChromeOptions()
preferences = {"download.default_directory":"/content",
"safebrowsing.enabled":"false",
"profile.default_content_settings.popups": 0}
chrome_options.add_argument('--headless')
chrome_options.add_argument('--no-sandbox')
chrome_options.add_argument('--disable-dev-shm-usage')
chrome_options.add_experimental_option("prefs", preferences)
wd = webdriver.Chrome('chromedriver',chrome_options=chrome_options)
wd.get("https://www.google.com")
# + id="kowJFRJ-NtFm" colab_type="code" colab={}
# 获取页面HTML
# 调试用
def get_html(url):
wp = requests.get(url)
web_html = BeautifulSoup(wp.text, 'html.parser')
file = open('test.html', 'w') # 创建并打开html文件
file.write(web_html.prettify()) # 写入美丽的汤prettify后的文字
file.close()
# + id="GRPb2MG3Ubvx" colab_type="code" colab={}
# 用yfinace获取csv历史数据
def get_hist_data(tickets_list):
for ticket in tickets_list:
company = yf.Ticker(ticket)
company.info # get stock info
hist = company.history(period="max") # get historical market data
hist.to_csv(r'/content//' + ticket + ".csv")
# + id="zeGqGaY4Uf0R" colab_type="code" colab={}
# 在这个里面填股票代码
TICKETS = ['AAPL', 'GOOG']
get_hist_data(TICKETS)
# + id="WvRBzfJ0ZA0_" colab_type="code" colab={}
# 一堆不太好用的东西
# wd.get("https://finance.yahoo.com/quote/AAPL/history?period1=0&period2=1569297600&interval=1d&filter=history&frequency=1d")
# xpath = '''//*[@id="Col1-1-HistoricalDataTable-Proxy"]/section/div[1]/div[2]/span[2]/a'''
# wd.find_element_by_xpath(xpath).click()
# elem = wd.find_element_by_tag_name("body")
# no_of_scrowdowns = 100
# while no_of_scrowdowns:
# elem.send_keys(Keys.PAGE_DOWN)
# #time.sleep(0.1)
# no_of_scrowdowns-=1
# df = pd.read_html(wd.page_source)
# + id="79ozC_NaaQfd" colab_type="code" colab={}
wp = requests.get("http://vip.stock.finance.sina.com.cn/usstock/ustotal.php")
web_html = BeautifulSoup(wp.text, 'html.parser')
soup = BeautifulSoup(str(web_html))
urls = []
for a in soup.find_all('a', href=True):
if
urls.append(a['href'])
for i in range(len(urls)):
if len(urls[i]) == 58: urls[i] = urls[i][48:53]
elif len(urls[i]) == 57: urls[i] = urls[i][48:52]
elif len(urls[i]) == 56: urls[i] = urls[i][48:51]
elif len(urls[i]) == 55: urls[i] = urls[i][48:50]
elif len(urls[i]) == 54: urls[i] = urls[i][48:49]
get_hist_data(urls[21:-13])
| Web_crawlers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# * [Pandas](http://pandas.pydata.org/)
# * [Seaborn](http://stanford.edu/~mwaskom/software/seaborn/)
# * [Bokeh](http://bokeh.pydata.org/en/latest/)
# * [Pygal](http://www.pygal.org/en/latest/)
# * [ggplot](http://ggplot.yhathq.com/)
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas as pd
import seaborn as sns
from ggplot import *
import pygal
import numpy as np
from sklearn import datasets
iris = sns.load_dataset("iris")
titanic = sns.load_dataset("titanic")
titanic.head()
sns.pairplot(titanic[["alive", "pclass", "fare", "age"]], hue="alive")
titanic[["survived", "pclass", "age", "fare"]]
| _source/2016-2-20-python-visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow]
# language: python
# name: conda-env-tensorflow-py
# ---
# ### Sample notebook
#
# Author: (arl)
# +
import os
import numpy as np
import matplotlib.pyplot as plt
import tensorflow.keras as K
from skimage import io
# -
import cellx
print(cellx.example_function())
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 1c: Building the SPN Graph Using A Generator
# Randomly stuctured SPNs can be generated using only the leaf layer and some parameters to specify the number of sums per scope and the number of decompositions at every product layer.
import libspn as spn
import tensorflow as tf
# ## Build the SPN
# +
indicator_leaves = spn.IndicatorLeaf(
num_vars=2, num_vals=2, name="indicator_x")
# Generate random structure with 1 decomposition per product layer
# 2 subsets of variables per product (so 2 children) and 2 sums/mixtures per scope
dense_spn_generator = spn.DenseSPNGenerator(num_decomps=1, num_subsets=2, num_mixtures=2)
root = dense_spn_generator.generate(indicator_leaves)
# Connect a latent indicator
indicator_y = root.generate_latent_indicators(name="indicator_y") # Can be added manually
# Generate weights
spn.generate_weights(root, initializer=tf.initializers.random_uniform()) # Can be added manually
# -
# ## Inspect
# Inspect
print(root.get_num_nodes())
print(root.get_scope())
print(root.is_valid())
# ## Visualize the SPN Graph
# The visualization below uses `graphviz`. Depending on your setup (e.g. `jupyter lab` vs. `jupyter notebook`) this might fail to show. At least `Chrome` + `jupyter notebook` seems to work.
# Visualize SPN graph
spn.display_spn_graph(root)
| ipynb/Tutorial 1c.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import glob, re
import numpy as np
import pandas as pd
from sklearn import *
from datetime import datetime
from xgboost import XGBRegressor
data = {
'tra': pd.read_csv('../input/air_visit_data.csv'),
'as': pd.read_csv('../input/air_store_info.csv'),
'hs': pd.read_csv('../input/hpg_store_info.csv'),
'ar': pd.read_csv('../input/air_reserve.csv'),
'hr': pd.read_csv('../input/hpg_reserve.csv'),
'id': pd.read_csv('../input/store_id_relation.csv'),
'tes': pd.read_csv('../input/sample_submission.csv'),
'hol': pd.read_csv('../input/date_info.csv').rename(columns={'calendar_date':'visit_date'})
}
data['hr'] = pd.merge(data['hr'], data['id'], how='inner', on=['hpg_store_id'])
for df in ['ar','hr']:
data[df]['visit_datetime'] = pd.to_datetime(data[df]['visit_datetime'])
data[df]['visit_datetime'] = data[df]['visit_datetime'].dt.date
data[df]['reserve_datetime'] = pd.to_datetime(data[df]['reserve_datetime'])
data[df]['reserve_datetime'] = data[df]['reserve_datetime'].dt.date
data[df]['reserve_datetime_diff'] = data[df].apply(lambda r: (r['visit_datetime'] - r['reserve_datetime']).days, axis=1)
tmp1 = data[df].groupby(['air_store_id','visit_datetime'], as_index=False)[['reserve_datetime_diff', 'reserve_visitors']].sum().rename(columns={'visit_datetime':'visit_date', 'reserve_datetime_diff': 'rs1', 'reserve_visitors':'rv1'})
tmp2 = data[df].groupby(['air_store_id','visit_datetime'], as_index=False)[['reserve_datetime_diff', 'reserve_visitors']].mean().rename(columns={'visit_datetime':'visit_date', 'reserve_datetime_diff': 'rs2', 'reserve_visitors':'rv2'})
data[df] = pd.merge(tmp1, tmp2, how='inner', on=['air_store_id','visit_date'])
data['tra']['visit_date'] = pd.to_datetime(data['tra']['visit_date'])
data['tra']['dow'] = data['tra']['visit_date'].dt.dayofweek
data['tra']['year'] = data['tra']['visit_date'].dt.year
data['tra']['month'] = data['tra']['visit_date'].dt.month
data['tra']['visit_date'] = data['tra']['visit_date'].dt.date
data['tes']['visit_date'] = data['tes']['id'].map(lambda x: str(x).split('_')[2])
data['tes']['air_store_id'] = data['tes']['id'].map(lambda x: '_'.join(x.split('_')[:2]))
data['tes']['visit_date'] = pd.to_datetime(data['tes']['visit_date'])
data['tes']['dow'] = data['tes']['visit_date'].dt.dayofweek
data['tes']['year'] = data['tes']['visit_date'].dt.year
data['tes']['month'] = data['tes']['visit_date'].dt.month
data['tes']['visit_date'] = data['tes']['visit_date'].dt.date
unique_stores = data['tes']['air_store_id'].unique()
stores = pd.concat([pd.DataFrame({'air_store_id': unique_stores, 'dow': [i]*len(unique_stores)}) for i in range(7)], axis=0, ignore_index=True).reset_index(drop=True)
# +
#sure it can be compressed...
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].min().rename(columns={'visitors':'min_visitors'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].mean().rename(columns={'visitors':'mean_visitors'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].median().rename(columns={'visitors':'median_visitors'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].max().rename(columns={'visitors':'max_visitors'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].count().rename(columns={'visitors':'count_observations'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
stores = pd.merge(stores, data['as'], how='left', on=['air_store_id'])
# NEW FEATURES FROM <NAME>
stores['air_genre_name'] = stores['air_genre_name'].map(lambda x: str(str(x).replace('/',' ')))
stores['air_area_name'] = stores['air_area_name'].map(lambda x: str(str(x).replace('-',' ')))
lbl = preprocessing.LabelEncoder()
for i in range(10):
stores['air_genre_name'+str(i)] = lbl.fit_transform(stores['air_genre_name'].map(lambda x: str(str(x).split(' ')[i]) if len(str(x).split(' '))>i else ''))
stores['air_area_name'+str(i)] = lbl.fit_transform(stores['air_area_name'].map(lambda x: str(str(x).split(' ')[i]) if len(str(x).split(' '))>i else ''))
stores['air_genre_name'] = lbl.fit_transform(stores['air_genre_name'])
stores['air_area_name'] = lbl.fit_transform(stores['air_area_name'])
data['hol']['visit_date'] = pd.to_datetime(data['hol']['visit_date'])
data['hol']['day_of_week'] = lbl.fit_transform(data['hol']['day_of_week'])
data['hol']['visit_date'] = data['hol']['visit_date'].dt.date
train = pd.merge(data['tra'], data['hol'], how='left', on=['visit_date'])
test = pd.merge(data['tes'], data['hol'], how='left', on=['visit_date'])
train = pd.merge(train, stores, how='left', on=['air_store_id','dow'])
test = pd.merge(test, stores, how='left', on=['air_store_id','dow'])
for df in ['ar','hr']:
train = pd.merge(train, data[df], how='left', on=['air_store_id','visit_date'])
test = pd.merge(test, data[df], how='left', on=['air_store_id','visit_date'])
train['id'] = train.apply(lambda r: '_'.join([str(r['air_store_id']), str(r['visit_date'])]), axis=1)
train['total_reserv_sum'] = train['rv1_x'] + train['rv1_y']
train['total_reserv_mean'] = (train['rv2_x'] + train['rv2_y']) / 2
train['total_reserv_dt_diff_mean'] = (train['rs2_x'] + train['rs2_y']) / 2
test['total_reserv_sum'] = test['rv1_x'] + test['rv1_y']
test['total_reserv_mean'] = (test['rv2_x'] + test['rv2_y']) / 2
test['total_reserv_dt_diff_mean'] = (test['rs2_x'] + test['rs2_y']) / 2
# NEW FEATURES FROM JMBULL
train['date_int'] = train['visit_date'].apply(lambda x: x.strftime('%Y%m%d')).astype(int)
test['date_int'] = test['visit_date'].apply(lambda x: x.strftime('%Y%m%d')).astype(int)
train['var_max_lat'] = train['latitude'].max() - train['latitude']
train['var_max_long'] = train['longitude'].max() - train['longitude']
test['var_max_lat'] = test['latitude'].max() - test['latitude']
test['var_max_long'] = test['longitude'].max() - test['longitude']
# NEW FEATURES FROM Georgii Vyshnia
train['lon_plus_lat'] = train['longitude'] + train['latitude']
test['lon_plus_lat'] = test['longitude'] + test['latitude']
lbl = preprocessing.LabelEncoder()
train['air_store_id2'] = lbl.fit_transform(train['air_store_id'])
test['air_store_id2'] = lbl.transform(test['air_store_id'])
col = [c for c in train if c not in ['id', 'air_store_id', 'visit_date','visitors']]
train = train.fillna(-1)
test = test.fillna(-1)
# -
# +
def RMSLE(y, pred):
return metrics.mean_squared_error(y, pred)**0.5
#model1 = ensemble.GradientBoostingRegressor(learning_rate=0.2, random_state=3, n_estimators=200, subsample=0.8,
# max_depth =10))
model2 = neighbors.KNeighborsRegressor(n_jobs=-1, n_neighbors=4)
model3 = XGBRegressor(learning_rate=0.2, seed=3, n_estimators=200, subsample=0.8,
colsample_bytree=0.8, max_depth =10)
# +
#model1.fit(train[col], np.log1p(train['visitors'].values))
model2.fit(train[col], np.log1p(train['visitors'].values))
model3.fit(train[col], np.log1p(train['visitors'].values))
#preds1 = model1.predict(train[col])
preds2 = model2.predict(train[col])
preds3 = model3.predict(train[col])
#print('RMSE GradientBoostingRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds1))
print('RMSE KNeighborsRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds2))
print('RMSE XGBRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds3))
# -
from lightgbm import LGBMRegressor
model1 = LGBMRegressor(learning_rate=0.3, num_leaves=1400, max_depth=15, max_bin=300, min_child_weight=5)
model1.fit(train[col], np.log1p(train['visitors'].values))
preds1 = model1.predict(train[col])
print('RMSE LightGBM: ', RMSLE(np.log1p(train['visitors'].values), preds1))
preds1 = model1.predict(test[col])
preds2 = model2.predict(test[col])
preds3 = model3.predict(test[col])
pred = (preds1+preds2+preds3) / 3.0
test['visitors'] = pred
test['visitors'] = np.expm1(test['visitors']).clip(lower=0.)
sub1 = test[['id','visitors']].copy()
print "Hi"
# +
from __future__ import division
# from hklee
# https://www.kaggle.com/zeemeen/weighted-mean-comparisons-lb-0-497-1st/code
dfs = { re.search('/([^/\.]*)\.csv', fn).group(1):
pd.read_csv(fn)for fn in glob.glob('../input/*.csv')}
for k, v in dfs.items(): locals()[k] = v
wkend_holidays = date_info.apply(
(lambda x:(x.day_of_week=='Sunday' or x.day_of_week=='Saturday') and x.holiday_flg==1), axis=1)
date_info.loc[wkend_holidays, 'holiday_flg'] = 0
date_info['weight'] = ((date_info.index + 1) / len(date_info)) ** 5
visit_data = air_visit_data.merge(date_info, left_on='visit_date', right_on='calendar_date', how='left')
visit_data.drop('calendar_date', axis=1, inplace=True)
visit_data['visitors'] = visit_data.visitors.map(pd.np.log1p)
wmean = lambda x:( (x.weight * x.visitors).sum() / x.weight.sum() )
visitors = visit_data.groupby(['air_store_id', 'day_of_week', 'holiday_flg']).apply(wmean).reset_index()
visitors.rename(columns={0:'visitors'}, inplace=True) # cumbersome, should be better ways.
sample_submission['air_store_id'] = sample_submission.id.map(lambda x: '_'.join(x.split('_')[:-1]))
sample_submission['calendar_date'] = sample_submission.id.map(lambda x: x.split('_')[2])
sample_submission.drop('visitors', axis=1, inplace=True)
sample_submission = sample_submission.merge(date_info, on='calendar_date', how='left')
sample_submission = sample_submission.merge(visitors, on=[
'air_store_id', 'day_of_week', 'holiday_flg'], how='left')
missings = sample_submission.visitors.isnull()
sample_submission.loc[missings, 'visitors'] = sample_submission[missings].merge(
visitors[visitors.holiday_flg==0], on=('air_store_id', 'day_of_week'),
how='left')['visitors_y'].values
missings = sample_submission.visitors.isnull()
sample_submission.loc[missings, 'visitors'] = sample_submission[missings].merge(
visitors[['air_store_id', 'visitors']].groupby('air_store_id').mean().reset_index(),
on='air_store_id', how='left')['visitors_y'].values
sample_submission['visitors'] = sample_submission.visitors.map(pd.np.expm1)
sub2 = sample_submission[['id', 'visitors']].copy()
sub_merge = pd.merge(sub1, sub2, on='id', how='inner')
sub_merge['visitors'] = (sub_merge['visitors_x'] + sub_merge['visitors_y']* 1.1)/2
# -
sub_merge['visitors'] = sub_merge['visitors'].astype(int)
sub_merge[['id', 'visitors']].to_csv('submissionINT2.csv', index=False)
| Recruit Restaurant Visitor Forecasting/Notebooks/.ipynb_checkpoints/Rework-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.insert(0, '..')
# %load_ext autoreload
# %autoreload 2
from mail_parser.mail_parser import MailParser
test_0 = """
Dear Passenger,
Due to an extraordinarily high number of customer contacts in our service centers and at our stations, we are facing delays in replying to your emails. We apologize for any inconveniences and appeal to your understanding to allow us to prioritize the most urgent inquires.
Please use the following links which answer to most of your inquiries:
https://www.example.com/en/example/example
Thank you for your understanding!
SECRET COMPANY
logo1
"""
print(MailParser().parse_mail(test_0))
test_1 = """Hello,
Please send your email to <EMAIL>
Thank you
logo1
JACQUES LAMA
Example - Example Air Transport
Example, Jacques Lama Airport
E-mail: <EMAIL>
Website: www.example.com
From: "Support-Example" <<EMAIL>>
Sent: Friday, April 24, 2020 11:07
To: "<EMAIL>" <<EMAIL>>
Subject: Reply to your last feedback - XXXXXX000000 - [ ref:XXXXX0000000:ref ]
19, rue example tour eiffel
000000 Tour eiffel
France
<EMAIL>
Subject: Reply to your last feedback
Ref. Secret Company: XXXXX-0000000
Réf. Secret:
Date: XX/XX/XXXX
EXAMPLE
Madam/Sir,
We refer to your latest feedback, under reference XXX-0000, concerning the claim of our client(s) Jacques Example, Jacques Example.
Our clients indeed contacted you before but as you never replied to their claim, they decided to entrust us with with their claim.
Please proceed with our initial demand.
We are looking forward to hear from you.
For a bank transfer: For a cheque:
Owner of the account: Secret Company
Bank: Secret Bank – Paris, France (00000)
IBAN: 0000 0000 0000 0000 0000 0000
BIC: XXXXX To the order of " SECRET COMPANY "
Send it to the following address: 19, rue example tour eiffel
Obligation to indicate the reference number of the claim on the order of payment
So that we can make the payment to our client in accordance with the terms of our agreement, it is necessary that your company specifies the claim reference on the order of payment. Otherwise, your company will not be discharge of its obligation of indemnification.
Reference to mention in the payment: XXXX-000000000
Sincerely,
Secret Department
Secret Company
ref:aeuizriuzdsksdk_sdhjdsjhsdhjds:ref
…
[Message tronqué] Afficher l'intégralité du message"""
print(MailParser().parse_mail(test_1))
test_2 = """
To: XYZ
CC/BCC:
Subject: Invitation to a birthday party
Hi XYZ!
Hope this mail finds you in the best of your time. I am very happy to invite you to my birthday party on Nov 03 at ABC Hotel from 7:00 pm to 10:00 pm. The theme of the birthday party is ‘Pirate of the Caribbean”.
It would be great if you come and join us at the party. We will have a great time and fun together.
See You Soon
Sincerely
LMN
"""
print(MailParser().parse_mail(test_2))
test_3 = """
BLABLABLABLABLA
BLOBLOBLOBLOBLOBLO
Will the recipient of <EMAIL> is the same as <EMAIL>?
I want to prevent sending email to some email address with a special character (like é) to go directly to the spam folder by replacing special character in the email address. But I am afraid if I am doing so, the recipient will be different.
Would that be a problem if I mean to send an email to <EMAIL> but write it as <EMAIL>? Will it have the same recipient?
Thank you GMail team.
Warm Regards,
Kris
"""
print(MailParser().parse_mail(test_3))
test_4 = """
Image
Image
Our reference: XX-00000
Your reference: XXX-0000000-00000
----------Forwarding process-----------
Dear Sir/Madam,
We received your return regarding the compensation request on behalf of our common clients Mr. <NAME> and Mrs. Example Jacques for their XX-0000 flight.
As mentioned, the flight was delayed less than 3 hours (30 minutes), passengers are therefore not entitled to compensation. So, we cannot grant your request.
Thank you for your understanding.
Kind regards,
Example
Example Customer Service
"""
print(MailParser().parse_mail(test_4))
| mail_parser/examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fastai2
# language: python
# name: conda-env-fastai2-py
# ---
# # Chapter 1
#
# My notes on Chapter 1 of the [fast.ai](https://www.fast.ai) course [Deep Learning for Coders](https://course.fast.ai).
#
# ## Chapter notes
# ## End of Chapter Questions
#
# 1. Do you need these for deep learning? (true/false)
#
# 1. Lots of maths: no - high school maths is sufficient
# 1. Lots of data: no - record-breaking results with <50 data items
# 1. Lots of expensive computers: no - get what you need from free
# 1. A PhD: no. Accessible to those with basic programming skills
#
# 1. Name 5 areas where deep learning is now the best tool in the world
#
# 1. Do you need these for deep learning?
#
# Book page 3:
# - Lots of math False
# - Lots of data False
# - Lots of expensive computers False
# - A PhD False
#
# 1. Name five areas where deep learning is now the best in the world.
#
# Book page 4:
# - Natural Language Processing
# - Computer Vision
# - Image generation
# - Financial forecasting
# - Protein folding
#
#
# 1. What was the name of the first device that was based on the principle of the artificial neuron?
#
# Book page 5: the artificial perceptron
#
#
# 1. Based on the book of the same name, what are the requirements for parallel distributed processing (PDP)?
#
# Book page 6:
# - A set of processing units
# - A state of activation
# - An output function for each unit
# - A pattern of connectivity among units
# - A propagation rule for propagating patterns of activities through the network of connectivities
# - A learning rule whereby patterns of connectivity are modified by experience
# - An environment within which the system must operate
#
#
# 1. What were the two theoretical misunderstandings that held back the field of neural networks?
#
# Page 6/7:
# - single layer of neurons not sufficient for even simple functions like XOR
# - lack of computing power (is this really a *theoretical* limitation?
#
#
# 1. What is a GPU?
# 1. Open a notebook and execute a cell containing: `1+1`. What happens?
# 1. Follow through each cell of the stripped version of the notebook for this chapter. Before executing each cell, guess what will happen.
# 1. Complete the Jupyter Notebook online appendix.
# 1. Why is it hard to use a traditional computer program to recognize images in a photo?
# 1. What did Samuel mean by "weight assignment"?
# 1. What term do we normally use in deep learning for what Samuel called "weights"?
# 1. Draw a picture that summarizes Samuel's view of a machine learning model.
# 1. Why is it hard to understand why a deep learning model makes a particular prediction?
# 1. What is the name of the theorem that shows that a neural network can solve any mathematical problem to any level of accuracy?
# 1. What do you need in order to train a model?
# 1. How could a feedback loop impact the rollout of a predictive policing model?
# 1. Do we always have to use 224×224-pixel images with the cat recognition model?
# 1. What is the difference between classification and regression?
# 1. What is a validation set? What is a test set? Why do we need them?
# 1. What will fastai do if you don't provide a validation set?
# 1. Can we always use a random sample for a validation set? Why or why not?
# 1. What is overfitting? Provide an example.
# 1. What is a metric? How does it differ from "loss"?
# 1. How can pretrained models help?
# 1. What is the "head" of a model?
# 1. What kinds of features do the early layers of a CNN find? How about the later layers?
# 1. Are image models only useful for photos?
# 1. What is an "architecture"?
# 1. What is segmentation?
# 1. What is `y_range` used for? When do we need it?
# 1. What are "hyperparameters"?
# 1. What's the best way to avoid failures when using AI in an organization?
| .ipynb_checkpoints/chapter1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Neural Networks
# ## Machine learning on images
import pandas as pd
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# ### MNIST
from keras.datasets import mnist
(X_train, y_train), (X_test, y_test) = mnist.load_data('/tmp/mnist.npz')
X_train.shape
X_test.shape
# +
#X_train[0]
# -
plt.imshow(X_train[0], cmap='gray')
X_train = X_train.reshape(-1, 28*28)
X_test = X_test.reshape(-1, 28*28)
X_train.shape
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255.0
X_test /= 255.0
# +
#X_train[0]
# -
from keras.utils.np_utils import to_categorical
y_train_cat = to_categorical(y_train)
y_test_cat = to_categorical(y_test)
y_train[0]
y_train_cat[0]
y_train_cat.shape
y_test_cat.shape
# ### Fully connected on images
# +
from keras.models import Sequential
from keras.layers import Dense
import keras.backend as K
K.clear_session()
model = Sequential()
model.add(Dense(512, input_dim=28*28, activation='relu'))
model.add(Dense(256, activation='relu'))
model.add(Dense(128, activation='relu'))
model.add(Dense(32, activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
# -
h = model.fit(X_train, y_train_cat, batch_size=128, epochs=10, verbose=1, validation_split=0.3)
plt.plot(h.history['acc'])
plt.plot(h.history['val_acc'])
plt.legend(['Training', 'Validation'])
plt.title('Accuracy')
plt.xlabel('Epochs')
test_accuracy = model.evaluate(X_test, y_test_cat)[1]
test_accuracy
# ### Tensor Math
A = np.random.randint(10, size=(2, 3, 4, 5))
B = np.random.randint(10, size=(2, 3))
A
A[0, 1, 0, 3]
B
# #### A random colored image
img = np.random.randint(255, size=(4, 4, 3), dtype='uint8')
img
# +
plt.figure(figsize=(5, 5))
plt.subplot(221)
plt.imshow(img)
plt.title("All Channels combined")
plt.subplot(222)
plt.imshow(img[:, : , 0], cmap='Reds')
plt.title("Red channel")
plt.subplot(223)
plt.imshow(img[:, : , 1], cmap='Greens')
plt.title("Green channel")
plt.subplot(224)
plt.imshow(img[:, : , 2], cmap='Blues')
plt.title("Blue channel")
# -
# ### Tensor operations
2 * A
A + A
A.shape
B.shape
np.tensordot(A, B, axes=([0, 1], [0, 1]))
np.tensordot(A, B, axes=([0], [0])).shape
np.tensordot(A, B, axes=([0], [0]))
# ### 1D convolution
a = np.array([0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0], dtype='float32')
b = np.array([1, -1], dtype='float32')
c = np.convolve(a, b)
a
b
c
# +
plt.subplot(211)
plt.plot(a, 'o-')
plt.subplot(212)
plt.plot(c, 'o-')
# -
# ### Image filters with convolutions
from scipy.ndimage.filters import convolve
from scipy.signal import convolve2d
from scipy import misc
img = misc.ascent()
img.shape
plt.imshow(img, cmap='gray')
h_kernel = np.array([[ 1, 2, 1],
[ 0, 0, 0],
[-1, -2, -1]])
plt.imshow(h_kernel, cmap='gray')
# +
res = convolve2d(img, h_kernel)
plt.imshow(res, cmap='gray')
# -
# ## Convolutional neural networks
from keras.layers import Conv2D
img.shape
plt.figure(figsize=(5, 5))
plt.imshow(img, cmap='gray')
img_tensor = img.reshape((1, 512, 512, 1))
model = Sequential()
model.add(Conv2D(1, (3, 3), strides=(2,1), input_shape=(512, 512, 1)))
model.compile('adam', 'mse')
model.summary()
img_pred_tensor = model.predict(img_tensor)
img_pred_tensor.shape
img_pred = img_pred_tensor[0, :, :, 0]
plt.imshow(img_pred, cmap='gray')
weights = model.get_weights()
weights[0].shape
plt.imshow(weights[0][:, :, 0, 0], cmap='gray')
weights[0] = np.ones(weights[0].shape)
model.set_weights(weights)
img_pred_tensor = model.predict(img_tensor)
img_pred_tensor.shape
img_pred = img_pred_tensor[0, :, :, 0]
img_pred.shape
plt.imshow(img_pred, cmap='gray')
# +
model = Sequential()
model.add(Conv2D(1, (3, 3), input_shape=(512, 512, 1), padding='same'))
model.compile('adam', 'mse')
img_pred_tensor = model.predict(img_tensor)
img_pred_tensor.shape
# -
# ## Pooling layers
from keras.layers import MaxPool2D, AvgPool2D
model = Sequential()
model.add(MaxPool2D((5, 5), input_shape=(512, 512, 1)))
model.compile('adam', 'mse')
img_pred = model.predict(img_tensor)[0, :, :, 0]
plt.imshow(img_pred, cmap='gray')
model = Sequential()
model.add(AvgPool2D((5, 5), input_shape=(512, 512, 1)))
model.compile('adam', 'mse')
img_pred = model.predict(img_tensor)[0, :, :, 0]
plt.imshow(img_pred, cmap='gray')
# ## Final architecture
X_train = X_train.reshape(-1, 28, 28, 1)
X_test = X_test.reshape(-1, 28, 28, 1)
X_train.shape
X_test.shape
from keras.layers import Flatten, Activation
# +
K.clear_session()
model = Sequential()
model.add(Conv2D(32, (3, 3), input_shape=(28, 28, 1)))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
# -
model.summary()
model.fit(X_train, y_train_cat, batch_size=128,
epochs=10, verbose=1, validation_split=0.3)
model.evaluate(X_test, y_test_cat)
# ### Exercise 1
# You've been hired by a shipping company to overhaul the way they route mail, parcels and packages. They want to build an image recognition system capable of recognizing the digits in the zipcode on a package, so that it can be automatically routed to the correct location.
# You are tasked to build the digit recognition system. Luckily, you can rely on the MNIST dataset for the intial training of your model!
#
# Build a deep convolutional neural network with at least two convolutional and two pooling layers before the fully connected layer.
#
# - Start from the network we have just built
# - Insert a `Conv2D` layer after the first `MaxPool2D`, give it 64 filters.
# - Insert a `MaxPool2D` after that one
# - Insert an `Activation` layer
# - retrain the model
# - does performance improve?
# - how many parameters does this new model have? More or less than the previous model? Why?
# - how long did this second model take to train? Longer or shorter than the previous model? Why?
# - did it perform better or worse than the previous model?
# +
K.clear_session()
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(28, 28, 1)))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
# -
model.summary()
model.fit(X_train, y_train_cat, batch_size=128,
epochs=10, verbose=1, validation_split=0.3)
model.evaluate(X_test, y_test_cat)
# ### Exercise 2
#
# Pleased with your performance with the digits recognition task, your boss decides to challenge you with a harder task. Their online branch allows people to upload images to a website that generates and prints a postcard that is shipped to destination. Your boss would like to know what images people are loading on the site in order to provide targeted advertising on the same page, so he asks you to build an image recognition system capable of recognizing a few objects. Luckily for you, there's a dataset ready made with a collection of labeled images. This is the [Cifar 10 Dataset](http://www.cs.toronto.edu/~kriz/cifar.html), a very famous dataset that contains images for 10 different categories:
#
# - airplane
# - automobile
# - bird
# - cat
# - deer
# - dog
# - frog
# - horse
# - ship
# - truck
#
# In this exercise we will reach the limit of what you can achieve on your laptop and get ready for the next session on cloud GPUs.
#
# Here's what you have to do:
# - load the cifar10 dataset using `keras.datasets.cifar10.load_data()`
# - display a few images, see how hard/easy it is for you to recognize an object with such low resolution
# - check the shape of X_train, does it need reshape?
# - check the scale of X_train, does it need rescaling?
# - check the shape of y_train, does it need reshape?
# - build a model with the following architecture, and choose the parameters and activation functions for each of the layers:
# - conv2d
# - conv2d
# - maxpool
# - conv2d
# - conv2d
# - maxpool
# - flatten
# - dense
# - output
# - compile the model and check the number of parameters
# - attempt to train the model with the optimizer of your choice. How fast does training proceed?
# - If training is too slow (as expected) stop the execution and move to the next session!
from keras.datasets import cifar10
(X_train, y_train), (X_test, y_test) = cifar10.load_data()
X_train.shape
plt.imshow(X_train[1])
X_train = X_train.astype('float32') / 255.0
X_test = X_test.astype('float32') / 255.0
y_train.shape
y_train_cat = to_categorical(y_train, 10)
y_test_cat = to_categorical(y_test, 10)
y_train_cat.shape
# +
model = Sequential()
model.add(Conv2D(32, (3, 3),
padding='same',
activation='relu',
input_shape=(32, 32, 3)))
model.add(Conv2D(32, (3, 3), activation='relu'))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3), padding='same', activation='relu'))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(512, activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
# -
model.summary()
model.fit(X_train, y_train_cat,
batch_size=32,
epochs=2,
validation_data=(X_test, y_test_cat),
shuffle=True)
model.evaluate(X_test, y_test_cat)
| course/6 Convolutional Neural Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Amit2016-17/autokeras/blob/master/cnnpytorch_crowd.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="L0lAtR1r-l-t" colab_type="code" colab={}
# Clone the entire repo.
# !git clone -l -s https://github.com/vivek-bombatkar/CSRNet-pytorch.git
# + [markdown] id="OpZjbgAjcsW9" colab_type="text"
#
# + id="-B-KghnYEUxw" colab_type="code" colab={}
# # cd /content/CSRNet-pytorch/
# !ls
# + id="dlUgzjaySa3G" colab_type="code" colab={}
# !rm -rf CSRNet-pytorch/
# + id="Xwrkv2sBSjX5" colab_type="code" colab={}
# !ls
# + id="kzRc5KVxFXFn" colab_type="code" colab={}
# !python make_dataset.ipynb
| cnnpytorch_crowd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:climpred-dev] *
# language: python
# name: conda-env-climpred-dev-py
# ---
# # Setting up your own output
#
# This demo demonstrates how you can setup your raw model output with ``climpred.preprocessing`` to match `climpred`'s expectations.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
import climpred
# -
from climpred.preprocessing.shared import load_hindcast, set_integer_time_axis
from climpred.preprocessing.mpi import get_path
# Assuming your raw model output is stored in multiple files per member and initialization, `load_hindcast` is a nice wrapper function based on `get_path` designed for the output format of `MPI-ESM` to aggregated all hindcast output into one file as expected by `climpred`.
#
# The basic idea is to look over the output of all members and concatinate, then loop over all initializations and concatinate. Before concatination, it is important to make the `time` dimension identical in all input datasets for concatination.
#
# To reduce the data size, use the `preprocess` function provided to `xr.open_mfdataset` wisely in combination with `set_integer_axis`, e.g. additionally extracting only a certain region, time-step, time-aggregation or only few variables for a multi-variable input file as in MPI-ESM standard output.
# +
# check the code of load_hindcast
# # load_hindcast??
# +
v = "global_primary_production"
def preprocess_1var(ds, v=v):
"""Only leave one variable `v` in dataset """
return ds[v].to_dataset(name=v).squeeze()
# -
# lead_offset because yearmean output
# %time ds = load_hindcast(inits=range(1961, 1965), members=range(1, 3), preprocess=preprocess_1var, get_path=get_path)
# what we need for climpred
ds.coords
ds[v].data
# loading the data into memory
# if not rechunk
# %time ds = ds.load()
# +
# go on with creation of PredictionEnsemble
# climpred.HindcastEnsemble(ds).add_observations(obs).verify(metric='acc', comparison='e2o', dim='init', alignment='maximize')
# climpred.PerfectModelEnsemble(ds).add_control(control).verify(metric='acc', comparison='m2e', dim=['init','member'])
# -
# # `intake-esm` for cmorized output
# In case you have access to cmorized output of CMIP experiments, consider using `intake-esm`. With the `preprocess` function you can align the `time` dimension of all input files. Finally, `rename_to_climpred_dims` only renames.
from climpred.preprocessing.shared import rename_to_climpred_dims, set_integer_time_axis
# make to have to install intake-esm installed, which is not included in climpred-dev
import intake # this is enough for intake-esm to work
col_url = "/home/mpim/m300524/intake-esm-datastore/catalogs/mistral-cmip6.json"
col_url = "https://raw.githubusercontent.com/NCAR/intake-esm-datastore/master/catalogs/pangeo-cmip6.json"
col = intake.open_esm_datastore(col_url)
col.df.columns
# load 2 members for 2 inits for one variable from one model
query = dict(experiment_id=[
'dcppA-hindcast'], table_id='Amon', member_id=['r1i1p1f1', 'r2i1p1f1'], dcpp_init_year=[1970, 1971],
variable_id='tas', source_id='MPI-ESM1-2-HR')
cat = col.search(**query)
cdf_kwargs = {'chunks': {'time': 12}, 'decode_times': False}
cat.df.head()
def preprocess(ds):
# extract tiny spatial and temporal subset to make this fast
ds = ds.isel(lon=[50, 51, 52], lat=[50, 51, 52],
time=np.arange(12 * 2))
# make time dim identical
ds = set_integer_time_axis(ds,time_dim='time')
return ds
dset_dict = cat.to_dataset_dict(
cdf_kwargs=cdf_kwargs, preprocess=preprocess)
# get first dict value
_, ds = dset_dict.popitem()
ds.coords
# rename to comply with climpred's required dimension names
ds = rename_to_climpred_dims(ds)
# what we need for climpred
ds.coords
ds['tas'].data
# loading the data into memory
# if not rechunk
# this is here quite fast before we only select 9 grid cells
# %time ds = ds.load()
# +
# go on with creation of PredictionEnsemble
# climred.HindcastEnsemble(ds).add_observations(obs).verify(metric='acc', comparison='e2o', dim='init', alignment='maximize')
# climred.PerfectModelEnsemble(ds).add_control(control).verify(metric='acc', comparison='m2e', dim=['init','member'])
| docs/source/examples/preprocessing/setup_your_own_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import scipp as sc
import scippneutron as scn
import urllib.request
url = 'https://github.com/ess-dmsc-dram/loki_tube_scripts/raw/master/test/test_data/LARMOR00049338.nxs'
filename, _ = urllib.request.urlretrieve(url)
data = scn.load(filename=filename)
edges = sc.array(dims=['tof'], unit='us', values=np.linspace(5.0, 100000.0, num=201))
data = sc.rebin(data, 'tof', edges)
for i in [1,2,3,4,5]:
mon = data.attrs[f'monitor{i}']
mon.value = sc.rebin(mon.value, 'tof', edges)
data.to_hdf5(filename='loki-at-larmor.hdf5')
import urllib
url = 'http://172.16.17.32/ftp/external-data/MD5/d5ae38871d0a09a28ae01f85d969de1e'
filename, _ = urllib.request.urlretrieve(url, filename='PG3_4844_event.nxs')
# +
import scipp as sc
import scippneutron as scn
da = scn.load(filename=f'PG3_4844_event.nxs', load_pulse_times=True)
# Fake d-spacing shift
da = scn.convert(da, 'tof', 'dspacing', scatter=True)
proton_charge = da.attrs['proton_charge'].value
tmin = proton_charge.coords['time'].min()
tmax = proton_charge.coords['time'].max()
delta = sc.to_unit(tmax-tmin, 'us')
delta.unit = ''
scale = sc.to_unit(da.bins.coords['pulse_time'] - tmin, 'us')* sc.scalar(1, unit='Angstrom/us')
da.bins.coords['dspacing'] += 0.02*scale/delta
da = scn.convert(da, 'dspacing', 'tof', scatter=True)
da.coords['tof'] = da.coords['tof']['spectrum', 0]
# Fake prompt pulse
prompt_start = 4000.0 * sc.Unit('us')
prompt_stop = 5000.0 * sc.Unit('us')
tof = da.bins.coords['tof']
da.bins.data *= sc.where((prompt_start <= tof) & (tof < prompt_stop), 1.0+3.0*sc.exp(-(tof-prompt_start)/sc.scalar(200.0, unit='us')), sc.scalar(1.0))
# Reduce data size to 1/3
da['spectrum', 14100:].to_hdf5(filename='powder-event.h5')
| tools/make_tutorial_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import pickle as pkl
import matplotlib.pyplot as plt
import os, sys
datapath = '../../data/IEC_2/'
fPaths = os.listdir(datapath)
fPaths.sort()
# demo load data
for ifile, file in enumerate(fPaths):
loadfile = os.path.join(datapath,file)
with open(loadfile, 'rb') as f:
test= pkl.load(f)
if ifile == 0:
events = test.copy()
for key in events:
events[key] = pd.concat([events[key],test[key]])
# +
fig,ax = plt.subplots(2,1)
events['ETM']['WS'].hist(ax=ax[0],color='C1',edgecolor='k')
ax[0].set_xlabel('Wind Speed [m/s]')
ax[0].set_ylabel('Counts [-]')
events['ETM']['WD'].hist(ax=ax[1],color='C1',edgecolor='k')
ax[1].set_xlabel('Wind Direction [m/s]')
ax[1].set_ylabel('Counts [-]')
fig.tight_layout()
# -
| scripts/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="cwBY2-4nuGh3"
import torch
import numpy as np
# + id="-EnLIu97x4Cu"
# ! pip install drive/My\ Drive/Colab\ Notebooks/b3_proj_2020/MyModules
# + id="7r3bSp9LuGiA"
from my_quantizers.ops import quant
# Load my precious library
# + id="leU2D9YCuGiC"
a = torch.Tensor(np.arange(-140.0, 140.0, 3.33, dtype=np.float))
# + id="1W8eaQBwuGiF"
a
# + id="GS4LoOGCuGiI"
b = quant.quantize_forward(a)
# + id="xNcAStXBuGiK"
b
# + id="qzooChsNuGiM"
for i in zip(a, b):
print("%7.2f %7.2f" % i)
# + id="aB0vFiD_uGiO"
| Exercise05/05_01_direct_import.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# +
data_path_5 = "./Data/mean_std_ceytometry_SMAF.csv"
SMAF = np.loadtxt(open(data_path_5),delimiter=",",skiprows=0)
#print(SMAF.shape)
data_path_3 = "./Data/DeepAE distictc measurements_ceytometry.csv"
data_measurement = np.loadtxt(open(data_path_3),delimiter=",",skiprows=0)
PCC_3 = data_measurement[:,(9,6,3,0)]
EM_3 = data_measurement[:,(10,7,4,1)]
MAE_3 = data_measurement[:,(11,8,5,2)]
# -
SMAF
# +
ind = np.arange(4) # the x locations for the groups
width = 0.15 # the width of the bars
color_list = plt.cm.Set3(np.linspace(0, 1, 12))
from matplotlib import rcParams
rcParams.update({'font.size': 14, 'font.family': 'STIXGeneral'})
from matplotlib import ticker
formatter = ticker.ScalarFormatter(useMathText=True)
formatter.set_scientific(True)
formatter.set_powerlimits((0,1))
fig, axes = plt.subplots(nrows=1,ncols=3, figsize=(15,4))
rects1 = axes[0].bar(ind - width/2 - 2*width, SMAF[(2,10,18,26),0],
width, yerr=SMAF[(3,11,19,27),0], color=color_list[0], label='SVD')
rects2 = axes[0].bar(ind - width/2 - width, SMAF[(4,12,20,28),0],
width, yerr=SMAF[(5,13,21,29),0], color=color_list[7], label='k-SVD')
rects3 = axes[0].bar(ind - width/2, SMAF[(6,14,22,30),0],
width, yerr=SMAF[(7,15,23,31),0], color=color_list[2], label='sNMF')
rects4 = axes[0].bar(ind + width/2, SMAF[(0,8,16,24),0],
width, yerr=SMAF[(1,9,17,25),0], color=color_list[3], label='CS-SMAF')
rects5 = axes[0].bar(ind + width/2 + width, np.mean(PCC_3[0:5, :], axis=0),
width, yerr=np.std(PCC_3[0:5, :], axis=0), color=color_list[4], label='DeepAE')
axes[0].set_ylabel('PCC')
axes[0].set_xticks(ind)
axes[0].set_ylim(0.4,1)
axes[0].set_xticklabels(('10', '25', '50', '100'))
#axes[0,0].set_title('GSE45234')
rects1 = axes[1].bar(ind - width/2 - 2*width, SMAF[(2,10,18,26),1],
width, yerr=SMAF[(3,11,19,27),0], color=color_list[0], label='SVD')
rects2 = axes[1].bar(ind - width/2 - width, SMAF[(4,12,20,28),1],
width, yerr=SMAF[(5,13,21,29),0], color=color_list[7], label='k-SVD')
rects3 = axes[1].bar(ind - width/2, SMAF[(6,14,22,30),1],
width, yerr=SMAF[(7,15,23,31),0], color=color_list[2], label='sNMF')
rects4 = axes[1].bar(ind + width/2, SMAF[(0,8,16,24),1],
width, yerr=SMAF[(1,9,17,25),1], color=color_list[3], label='CS-SMAF')
rects5 = axes[1].bar(ind + width/2 + width, np.mean(EM_3[0:5, :], axis=0),
width, yerr=np.std(EM_3[0:5, :], axis=0), color=color_list[4], label='DeepAE')
axes[1].set_ylabel('EM')
axes[1].set_xticks(ind)
axes[1].set_xticklabels(('10', '25', '50', '100'))
axes[1].set_title('Mass cytometry data')
axes[1].yaxis.set_major_formatter(formatter)
axes[1].set_xlabel('Measurements')
rects1 = axes[2].bar(ind - width/2 - 2*width, SMAF[(2,10,18,26),2],
width, yerr=SMAF[(3,11,19,27),0], color=color_list[0], label='SVD')
rects2 = axes[2].bar(ind - width/2 - width, SMAF[(4,12,20,28),2],
width, yerr=SMAF[(5,13,21,29),2], color=color_list[7], label='k-SVD')
rects3 = axes[2].bar(ind - width/2, SMAF[(6,14,22,30),2],
width, yerr=SMAF[(7,15,23,31),0], color=color_list[2], label='sNMF')
rects4 = axes[2].bar(ind + width/2, SMAF[(0,8,16,24),2],
width, yerr=SMAF[(1,9,17,25),2], color=color_list[3], label='CS-SMAF')
rects5 = axes[2].bar(ind + width/2 + width, np.mean(MAE_3[0:5, :], axis=0),
width, yerr=np.std(MAE_3[0:5, :], axis=0), color=color_list[4], label='DeepAE')
axes[2].set_ylabel('MAE')
axes[2].set_xticks(ind)
axes[2].set_xticklabels(('10', '25', '50', '100'))
#axes[0,2].set_title('GSE45234')
axes[2].yaxis.set_major_formatter(formatter)
axes[2].legend(loc=1)
plt.show()
fig.savefig('./Data/ceytomtry.pdf', dpi=200, bbox_inches='tight')
# -
SMAF[(2,10,18,26),0]
| Plot_Bars_ceytometry_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="4f3CKqFUqL2-" slideshow={"slide_type": "slide"}
# # Hand tuning hyperparameters
# -
# **Learning Objectives:**
# * Use the `LinearRegressor` class in TensorFlow to predict median housing price, at the granularity of city blocks, based on one input feature
# * Evaluate the accuracy of a model's predictions using Root Mean Squared Error (RMSE)
# * Improve the accuracy of a model by hand-tuning its hyperparameters
# The data is based on 1990 census data from California. This data is at the city block level, so these features reflect the total number of rooms in that block, or the total number of people who live on that block, respectively. Using only one input feature -- the number of rooms -- predict house value.
# + [markdown] colab_type="text" id="6TjLjL9IU80G"
# ## Set Up
# In this first cell, we'll load the necessary libraries.
# +
import math
import shutil
import numpy as np
import pandas as pd
import tensorflow as tf
print(tf.__version__)
tf.logging.set_verbosity(tf.logging.INFO)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
# + [markdown] colab_type="text" id="ipRyUHjhU80Q"
# Next, we'll load our data set.
# -
df = pd.read_csv("https://storage.googleapis.com/ml_universities/california_housing_train.csv", sep=",")
# + [markdown] colab_type="text" id="HzzlSs3PtTmt" slideshow={"slide_type": "-"}
# ## Examine the data
#
# It's a good idea to get to know your data a little bit before you work with it.
#
# We'll print out a quick summary of a few useful statistics on each column.
#
# This will include things like mean, standard deviation, max, min, and various quantiles.
# -
df.head()
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "test": {"output": "ignore", "timeout": 600}} colab_type="code" id="gzb10yoVrydW" slideshow={"slide_type": "slide"}
df.describe()
# -
# In this exercise, we'll be trying to predict median_house_value. It will be our label (sometimes also called a target). Can we use total_rooms as our input feature? What's going on with the values for that feature?
#
# This data is at the city block level, so these features reflect the total number of rooms in that block, or the total number of people who live on that block, respectively. Let's create a different, more appropriate feature. Because we are predicing the price of a single house, we should try to make all our features correspond to a single house as well
df['num_rooms'] = df['total_rooms'] / df['households']
df.describe()
# Split into train and eval
np.random.seed(seed=1) #makes split reproducible
msk = np.random.rand(len(df)) < 0.8
traindf = df[msk]
evaldf = df[~msk]
# + [markdown] colab_type="text" id="Lr6wYl2bt2Ep" slideshow={"slide_type": "-"}
# ## Build the first model
#
# In this exercise, we'll be trying to predict `median_house_value`. It will be our label (sometimes also called a target). We'll use `num_rooms` as our input feature.
#
# To train our model, we'll use the [LinearRegressor](https://www.tensorflow.org/api_docs/python/tf/estimator/LinearRegressor) estimator. The Estimator takes care of a lot of the plumbing, and exposes a convenient way to interact with data, training, and evaluation.
# +
OUTDIR = './housing_trained'
def train_and_evaluate(output_dir, num_train_steps):
estimator = tf.estimator.LinearRegressor(
model_dir = output_dir,
feature_columns = [tf.feature_column.numeric_column('num_rooms')])
#Add rmse evaluation metric
def rmse(labels, predictions):
pred_values = tf.cast(predictions['predictions'],tf.float64)
return {'rmse': tf.metrics.root_mean_squared_error(labels, pred_values)}
estimator = tf.contrib.estimator.add_metrics(estimator,rmse)
train_spec=tf.estimator.TrainSpec(
input_fn = tf.estimator.inputs.pandas_input_fn(x = traindf[["num_rooms"]],
y = traindf["median_house_value"], # note the scaling
num_epochs = None,
shuffle = True),
max_steps = num_train_steps)
eval_spec=tf.estimator.EvalSpec(
input_fn = tf.estimator.inputs.pandas_input_fn(x = evaldf[["num_rooms"]],
y = evaldf["median_house_value"], # note the scaling
num_epochs = 1,
shuffle = False),
steps = None,
start_delay_secs = 1, # start evaluating after N seconds
throttle_secs = 10, # evaluate every N seconds
)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
# Run training
shutil.rmtree(OUTDIR, ignore_errors = True) # start fresh each time
train_and_evaluate(OUTDIR, num_train_steps = 100)
# -
# ## 1. Scale the output
# Let's scale the target values so that the default parameters are more appropriate.
# +
SCALE = 100000
OUTDIR = './housing_trained'
def train_and_evaluate(output_dir, num_train_steps):
estimator = tf.estimator.LinearRegressor(
model_dir = output_dir,
feature_columns = [tf.feature_column.numeric_column('num_rooms')])
#Add rmse evaluation metric
def rmse(labels, predictions):
pred_values = tf.cast(predictions['predictions'],tf.float64)
return {'rmse': tf.metrics.root_mean_squared_error(labels*SCALE, pred_values*SCALE)}
estimator = tf.contrib.estimator.add_metrics(estimator,rmse)
train_spec=tf.estimator.TrainSpec(
input_fn = tf.estimator.inputs.pandas_input_fn(x = traindf[["num_rooms"]],
y = traindf["median_house_value"] / SCALE, # note the scaling
num_epochs = None,
shuffle = True),
max_steps = num_train_steps)
eval_spec=tf.estimator.EvalSpec(
input_fn = tf.estimator.inputs.pandas_input_fn(x = evaldf[["num_rooms"]],
y = evaldf["median_house_value"] / SCALE, # note the scaling
num_epochs = 1,
shuffle = False),
steps = None,
start_delay_secs = 1, # start evaluating after N seconds
throttle_secs = 10, # evaluate every N seconds
)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
# Run training
shutil.rmtree(OUTDIR, ignore_errors = True) # start fresh each time
train_and_evaluate(OUTDIR, num_train_steps = 100)
# -
# ## 2. Change learning rate and batch size
# Can you come up with better parameters?
# +
SCALE = 100000
OUTDIR = './housing_trained'
def train_and_evaluate(output_dir, num_train_steps):
myopt = tf.train.FtrlOptimizer(learning_rate = 0.2) # note the learning rate
estimator = tf.estimator.LinearRegressor(
model_dir = output_dir,
feature_columns = [tf.feature_column.numeric_column('num_rooms')],
optimizer = myopt)
#Add rmse evaluation metric
def rmse(labels, predictions):
pred_values = tf.cast(predictions['predictions'],tf.float64)
return {'rmse': tf.metrics.root_mean_squared_error(labels*SCALE, pred_values*SCALE)}
estimator = tf.contrib.estimator.add_metrics(estimator,rmse)
train_spec=tf.estimator.TrainSpec(
input_fn = tf.estimator.inputs.pandas_input_fn(x = traindf[["num_rooms"]],
y = traindf["median_house_value"] / SCALE, # note the scaling
num_epochs = None,
batch_size = 512, # note the batch size
shuffle = True),
max_steps = num_train_steps)
eval_spec=tf.estimator.EvalSpec(
input_fn = tf.estimator.inputs.pandas_input_fn(x = evaldf[["num_rooms"]],
y = evaldf["median_house_value"] / SCALE, # note the scaling
num_epochs = 1,
shuffle = False),
steps = None,
start_delay_secs = 1, # start evaluating after N seconds
throttle_secs = 10, # evaluate every N seconds
)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
# Run training
shutil.rmtree(OUTDIR, ignore_errors = True) # start fresh each time
train_and_evaluate(OUTDIR, num_train_steps = 100)
# + [markdown] colab_type="text" id="QU5sLyYTqzqL" slideshow={"slide_type": "slide"}
# ### Is there a standard method for tuning the model?
#
# This is a commonly asked question. The short answer is that the effects of different hyperparameters is data dependent. So there are no hard and fast rules; you'll need to run tests on your data.
#
# Here are a few rules of thumb that may help guide you:
#
# * Training error should steadily decrease, steeply at first, and should eventually plateau as training converges.
# * If the training has not converged, try running it for longer.
# * If the training error decreases too slowly, increasing the learning rate may help it decrease faster.
# * But sometimes the exact opposite may happen if the learning rate is too high.
# * If the training error varies wildly, try decreasing the learning rate.
# * Lower learning rate plus larger number of steps or larger batch size is often a good combination.
# * Very small batch sizes can also cause instability. First try larger values like 100 or 1000, and decrease until you see degradation.
#
# Again, never go strictly by these rules of thumb, because the effects are data dependent. Always experiment and verify.
# + [markdown] colab_type="text" id="GpV-uF_cBCBU" slideshow={"slide_type": "slide"}
# ### 3: Try adding more features
#
# See if you can do any better by adding more features.
#
# Don't take more than 5 minutes on this portion.
# -
| 5-HandTuning models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:c10]
# language: python
# name: conda-env-c10-py
# ---
# +
# # 从leetcode上爬图片并生成gif
# import imageio
# from skimage.io import imread
# +
# url_sample = "https://pic.leetcode-cn.com/Figures/binary_tree/level_traversal/Slide11.png"
# gif_name = "level_order.gif"
# total_img_nums = 17
# url_sample = "https://pic.leetcode-cn.com/Figures/binary_tree/preorder_traversal/Slide13.png"
# gif_name = "preorder_traversal.gif"
# total_img_nums = 19
# url_sample = "https://pic.leetcode-cn.com/Figures/binary_tree/inorder_traversal/Slide22.png"
# gif_name = "inorder_traversal.gif"
# total_img_nums = 22
# url_sample = "https://pic.leetcode-cn.com/Figures/binary_tree/postorder_traversal/Slide19.png"
# gif_name = "inorder_traversal.gif"
# # total_img_nums = 19
# ext_suffix = "." + url_sample.split('/')[-1].split('.')[-1]
# url_template = url_sample[:-6]
# gif_duration = 1.0
# +
def getLeetcodeGif(url_sample: str, gif_name: str, gif_duration=1.0, src_img_ext: str = ".png"):
# 从leetcode上爬图片并生成gif
from imageio import mimsave
from skimage.io import imread
# url_sample = "https://pic.leetcode-cn.com/Figures/binary_tree/postorder_traversal/Slide19.png"
gif_name = gif_name + ".gif"
# total_img_nums = 19
ext_suffix = "." + url_sample.split('/')[-1].split('.')[-1]
url_template = url_sample[:-6]
gif_duration = 1.0
frames = []
ind = 1
while True:
url = url_template + str(ind).zfill(2) + ext_suffix
try:
img = imread(url)
print("GET >>> " + url)
ind += 1
frames.append(img)
except Exception as err:
if ind > 3:
print(f"Failed @ {ind}th image >>> {err}")
else:
if ind == 3:
print(f"Failed @ third image >>> {err}")
if ind == 2:
print(f"Failed @ sencod image >>> {err}")
if ind == 1:
print(f"Failed @ first image >>> {err}")
break
print(f"Got {len(frames)} image(s)")
mimsave(gif_name, frames, "GIF", duration=gif_duration)
print("Done!")
# -
getLeetcodeGif(
"https://pic.leetcode-cn.com/Figures/binary_tree/postorder_traversal/Slide19.png",
"postorder_traversal"
)
getLeetcodeGif(
"https://pic.leetcode-cn.com/Figures/binary_tree/top_down/Slide01.png",
"maxdepth_topdown"
)
getLeetcodeGif(
"https://pic.leetcode-cn.com/Figures/binary_tree/bottom_up/Slide01.png",
"maxdepth_bottomup"
)
| code/btree/images/gif.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/RachitBansal/RedditFlairDetector/blob/master/2_EDA%26PreProcessing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_3USh6cVScAz" colab_type="text"
# # EDA and Pre-Processing
# + [markdown] id="p5I-i-gs6zX8" colab_type="text"
# In this notebook, we will analyse the data which we collected in the previous step by performing appropriate EDA Techniques and will proprocess the data accordingly. Lastly, the final processed data would be saved as a csv file to be used for modelling.
#
# .
# + id="dVdFmu_JEqFo" colab_type="code" outputId="f58a519f-215c-48c2-e8c1-b8be250be01a" colab={"base_uri": "https://localhost:8080/", "height": 72}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# + id="UkFq9A8tE_T7" colab_type="code" outputId="4be6a62a-ae39-452d-e112-670732aea0e1" colab={"base_uri": "https://localhost:8080/", "height": 54}
from google.colab import drive
drive.mount('drive')
# + [markdown] id="CxHH2XBB7f5t" colab_type="text"
# .
#
# Reading the pre-final data file which is to be analysed and processed.
# + id="WGbs4KjTFMcO" colab_type="code" outputId="06fc160b-6fa3-413e-b455-e453c44935e9" colab={"base_uri": "https://localhost:8080/", "height": 72}
df = pd.read_csv('./drive/My Drive/rMIDAS_final.csv')
# + id="kK2G1_LzFfc5" colab_type="code" outputId="69d0fe57-366f-48be-d2f5-2847e43cb221" colab={"base_uri": "https://localhost:8080/", "height": 887}
df.head(10)
# + id="7JMxZMJ98MX4" colab_type="code" outputId="6137bd64-8b37-4131-aa4a-cd32ed99d6d1" colab={"base_uri": "https://localhost:8080/", "height": 34}
df.shape
# + [markdown] id="wwU7lutm8HNy" colab_type="text"
# We can see that we have close to .75M rows in our data
#
# .
#
# Analysing the number of Flairs in the entire data as well as the number of Data Points for each of them:
# + id="eYc2Snp9FlW-" colab_type="code" outputId="36da77bb-b63f-4734-e099-50118d06b306" colab={"base_uri": "https://localhost:8080/", "height": 538}
flairs_in_df = list( dict.fromkeys(list(df.loc[:, 'link_flair_text'].values)))
print(len(flairs_in_df))
for flair in flairs_in_df:
count = np.sum(df.loc[:, 'link_flair_text'].values == flair)
if(count>500):
print(flair, "\t", count)
# + [markdown] id="DaSwxzBnSwYk" colab_type="text"
# ## Handling redundant data
# + [markdown] id="iJvRXjUD8vJb" colab_type="text"
# From the above cell, it can be seen that there is a presence of some redundant Flairs in the data like 'Science & Technology' <-> 'Science/Technology' and 'Business & Finance' <-> 'Business/Finance'. These can be concatendated as done below:
# + id="kOfTTTfHv1HM" colab_type="code" colab={}
df['selftext'].replace('[deleted]', 'None', inplace=True)
df['link_flair_text'].replace('Science & Technology', 'Science/Technology', inplace=True)
df['link_flair_text'].replace('Business & Finance', 'Business/Finance', inplace=True)
df['link_flair_text'].replace('CAA-NRC', 'Politics', inplace=True)
df['link_flair_text'].replace('Demonetization', 'Policy/Economy', inplace=True)
df['link_flair_text'].replace('Policy & Economy', 'Policy/Economy', inplace=True)
# + [markdown] id="LyKtN23x9pYR" colab_type="text"
# .
# + id="jptCgYK0Fg8M" colab_type="code" outputId="7a517be3-c64a-46c7-c3f6-79592fdc70ec" colab={"base_uri": "https://localhost:8080/", "height": 260}
df.info()
# + [markdown] id="5jt15HtLS3c0" colab_type="text"
# ## Handling 'NaN's
# + [markdown] id="HND9uqoz9uxr" colab_type="text"
# It can be observed that close to .24M rows have our label (i.e. link_flair_text) as 'NaN' and around 0.05M rows have our main input (i.e. title) as 'NaN'. These rows can't be included in our data and are removed in the step below:
# + id="SxJQTw4mblEX" colab_type="code" outputId="28236124-759b-4fcf-c3c4-064be8d62f09" colab={"base_uri": "https://localhost:8080/", "height": 34}
df = df.dropna('index', subset = ['title', 'link_flair_text'])
df['num_comments'] = df['num_comments'].astype(int)
df = df.reset_index()
df = df.drop('index', 'columns')
print(df.shape)
# + id="ClK2gR51w1HM" colab_type="code" outputId="dd03bace-426a-4dcd-b33b-7e246377588a" colab={"base_uri": "https://localhost:8080/", "height": 451}
flairs_to_keep = []
flairs_in_df = list( dict.fromkeys(list(df.loc[:, 'link_flair_text'].values)))
print(len(flairs_in_df))
for flair in flairs_in_df:
count = np.sum(df.loc[:, 'link_flair_text'].values == flair)
if(count>500):
print(flair, "\t", count)
flairs_to_keep.append(flair)
# + [markdown] id="ehR07ZY3-sLS" colab_type="text"
# After performing the preprocessing steps so far, we are left with .45M rows and 700 Flairs in the data.
#
# .
#
# Out of these 700, only those Flairs are to be kept in the final data which have at lest 500 values and that is done by removing the others:
# + id="RZoGxrrwVQD7" colab_type="code" outputId="156d5207-af4b-47de-8465-48f403a46cc0" colab={"base_uri": "https://localhost:8080/", "height": 312}
for i in range(df.shape[0]):
if(i%10000 == 0):
print(i)
if(df.loc[i, 'link_flair_text'] not in flairs_to_keep):
df = df.drop(i, 'index')
continue
# + id="027mwhsTWoF5" colab_type="code" outputId="90060670-5863-4e7f-afd5-669d35a4802d" colab={"base_uri": "https://localhost:8080/", "height": 34}
df.reset_index(inplace=True)
df.drop('index', 'columns', inplace = True)
print(df.shape)
# + id="O_6Nt75C5wAr" colab_type="code" outputId="420eef56-afef-4123-c2fb-f69da591b895" colab={"base_uri": "https://localhost:8080/", "height": 206}
df.head()
# + [markdown] id="_ViwU-JIAJd1" colab_type="text"
# .
#
# The figure belolw shows the 'NaN' Values in the entire data in the form of a heatmap, it can be observed that only the 'selftext' column (which represents the description of a reddit post) has NaN values. These are replaced by a placeholder 'None'.
# + id="78JCRjGAHeiN" colab_type="code" outputId="ec399a48-3d58-4917-e6fe-8a02deebcdd4" colab={"base_uri": "https://localhost:8080/", "height": 352}
sns.heatmap(df.isnull(),cbar=False,yticklabels=False,cmap = 'viridis')
# + id="rWDb-g1Y5-m6" colab_type="code" colab={}
df.fillna('None', inplace = True)
# + [markdown] id="TQcOSfUt_nLu" colab_type="text"
# The figure below shows the amount of data available corresponding to each Flair
# + id="FzGFwgW5ztY9" colab_type="code" outputId="46476a29-f4f6-41ce-e798-fbf57763a118" colab={"base_uri": "https://localhost:8080/", "height": 553}
flairs_count = {
'flair': [],
'count': []
}
for flair in flairs_to_keep:
count = np.sum(df.loc[:, 'link_flair_text'].values == flair)
flairs_count['flair'].append(flair)
flairs_count['count'].append(count)
pd.DataFrame(flairs_count) \
.round(decimals=2) \
.sort_values('count', ascending=False) \
.style.bar(color=['grey', 'red'], align='zero')
# + [markdown] id="mjdTqKX6jThx" colab_type="text"
# ## Balancing out the dataset
#
# As can be seen from the above figure, the data is highly unbalanced across the classes, thus data balancing is carried out by resampling and downsampling from this data. The most data-rich classes 'Politics' and 'Non-Politics' have been reduced to a comparable size as the other classes.
# + id="rrZD-wDGe8uD" colab_type="code" colab={}
politics_df = df.loc[df['link_flair_text'] == 'Politics']
non_pol_df = df.loc[df['link_flair_text'] == 'Non-Political']
# + colab_type="code" id="L2aS7gJ1g0b7" colab={}
n_p_df = df.loc[df['link_flair_text'] != 'Politics']
n_p_df = n_p_df.loc[n_p_df['link_flair_text'] != 'Non-Political']
# + id="MD1wp0Lrg1kf" colab_type="code" outputId="74ca82dd-9e60-40f9-d37b-068d374766d2" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(politics_df.shape, non_pol_df.shape, n_p_df.shape)
# + id="nGdfP65Gf_9l" colab_type="code" colab={}
from sklearn.utils import resample
pol_downs = resample(politics_df,
replace = False, # sample without replacement
n_samples = 75000, # match minority n
random_state = 27)
non_pol_downs = resample(non_pol_df,
replace = False, # sample without replacement
n_samples = 75000, # match minority n
random_state = 27)
# + id="jRbhywa0hX8j" colab_type="code" outputId="091809eb-fc0a-48d9-8cd1-7ba9afdd7253" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(pol_downs.shape, non_pol_downs.shape)
# + id="UQeAsc1ThfnI" colab_type="code" colab={}
df_bal = pd.concat([pol_downs, non_pol_downs, n_p_df])
# + id="udKI8asNhtni" colab_type="code" outputId="a95da12f-7204-4075-866d-e249d848c498" colab={"base_uri": "https://localhost:8080/", "height": 34}
df_bal.shape
# + id="6MCrhGW-iL0j" colab_type="code" colab={}
flairs_to_keep = list(set(df['link_flair_text']))
# + id="gc4DgcqWh4QB" colab_type="code" outputId="aa972bfd-51a9-4804-d39c-6c8a3c50e61d" colab={"base_uri": "https://localhost:8080/", "height": 553}
flairs_count = {
'flair': [],
'count': []
}
for flair in flairs_to_keep:
count = np.sum(df_bal.loc[:, 'link_flair_text'].values == flair)
flairs_count['flair'].append(flair)
flairs_count['count'].append(count)
pd.DataFrame(flairs_count) \
.round(decimals=2) \
.sort_values('count', ascending=False) \
.style.bar(color=['grey', 'red'], align='zero')
# + id="RtakylgUX8Pg" colab_type="code" colab={}
df_to_keep = []
for flair in flairs_to_keep:
df_ = df_bal.loc[df['link_flair_text'] == flair]
if(df_.shape[0] > 4000):
df_to_keep.append(df_)
# + id="DTGarI0TY0eU" colab_type="code" colab={}
df_bal_2 = pd.concat(df_to_keep)
# + id="F03ZZJG-ZAOc" colab_type="code" outputId="1cd06ffa-e6be-4abc-e95e-428cc9a1183c" colab={"base_uri": "https://localhost:8080/", "height": 297}
flairs_count = {
'flair': [],
'count': []
}
for flair in list(set(df_bal_2['link_flair_text'])):
count = np.sum(df_bal_2.loc[:, 'link_flair_text'].values == flair)
flairs_count['flair'].append(flair)
flairs_count['count'].append(count)
pd.DataFrame(flairs_count) \
.round(decimals=2) \
.sort_values('count', ascending=False) \
.style.bar(color=['grey', 'red'], align='zero')
# + id="aCz2kVQ6jeHI" colab_type="code" outputId="2d6ae7ad-9d87-4cb7-b937-83912118ffd6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df_bal_2 = df_bal_2.sample(frac=1).reset_index(drop=True)
df_bal_2.head(20)
# + [markdown] id="h982kKB__19U" colab_type="text"
# .
#
# Finally, after preprocessing, we are left with **446283 (.446M) Data Points** across **23 Classes (Flairs)** in the Unbalanced Data
# While, **315099 (.315M) Data Points** across **12 Classes (Flairs)** in the Balanced Data.
#
# .
# + [markdown] id="u4EhmL01TmbY" colab_type="text"
# ## Further Exploration of the Data
# + [markdown] id="ir4xzBTbBJxk" colab_type="text"
# The discussion from here on involves numerical EDA of the data by the following ways:
#
# - Studying the relation and correlation between [Number of Comments, Length of {Title, Self Text and URL}] wrt the Label (Flair) in the data.
#
# - Plotting the Correlation Matrix of the above quantities in various ways.
# + id="Uh9EYwyYfRBw" colab_type="code" outputId="ddaec744-a148-4e76-ef30-48fccdbe2f23" colab={"base_uri": "https://localhost:8080/", "height": 139}
# !pip install pytorch-nlp
# + id="VqSg75gbex--" colab_type="code" colab={}
import torchnlp
from torchnlp.encoders import LabelEncoder
# + [markdown] id="F6yeAJq0CYQh" colab_type="text"
# .
#
# Creating a corresponding dataset containing the numerical data as was explained above
#
# - LabelEncoding the flairs
# - {Title, Selftext, URL} --> Length of each
# + id="s14IjYZ3IXS9" colab_type="code" colab={}
df_cols = {'link_flair_text':[], 'num_comments':[], 'selftext':[], 'title':[], 'url':[]}
# + id="keEzo8hiJmo6" colab_type="code" colab={}
for i in range(df.shape[0]):
df_cols['link_flair_text'].append(df.loc[i, 'link_flair_text'])
df_cols['title'].append(len(df.loc[i, 'title']))
df_cols['url'].append(len(df.loc[i, 'url']))
df_cols['num_comments'].append(int(df.loc[i, 'num_comments']))
if type(df.loc[i, 'selftext']) != float:
df_cols['selftext'].append(len(df.loc[i, 'selftext']))
else:
df_cols['selftext'].append(0)
# + id="DS5ed6uqNell" colab_type="code" colab={}
encoder = LabelEncoder(df_cols['link_flair_text'])
# + id="FSRVecu4PD_n" colab_type="code" colab={}
df_cols['link_flair_text'] = encoder.batch_encode(df_cols['link_flair_text'])
# + id="05MOsAvSK_Qx" colab_type="code" outputId="b502ba91-a010-41a8-c854-573dcef004cd" colab={"base_uri": "https://localhost:8080/", "height": 363}
df_supp = pd.DataFrame(df_cols)
df_supp.head(10)
# + id="pkk4ewQ3j19h" colab_type="code" outputId="46b4347d-338c-4ddc-9eb9-4131ed26a8b8" colab={"base_uri": "https://localhost:8080/", "height": 300}
df_supp.describe()
# + id="SUpfk0S0LfDx" colab_type="code" outputId="96565081-93f3-4ab7-9a0a-7a17693a9bc2" colab={"base_uri": "https://localhost:8080/", "height": 356}
plt.figure(figsize=(6,4))
sns.heatmap(df_supp.corr(),cmap='Blues',annot=False)
# + [markdown] id="-bSM0i-rDK7g" colab_type="text"
# In the above figure, dark shades represents positive correlation while lighter shades represents negative correlation.
# + id="XIGTu0FePxuC" colab_type="code" outputId="b07f1620-e8a0-4e80-d411-b1a8e37450b6" colab={"base_uri": "https://localhost:8080/", "height": 396}
k = 5 #number of variables for heatmap
cols = df_supp.corr().nlargest(k, 'link_flair_text')['link_flair_text'].index
cm = df_supp[cols].corr()
plt.figure(figsize=(10,6))
sns.heatmap(cm, annot=True, cmap = 'viridis')
# + id="YfBGg-dgQud1" colab_type="code" outputId="31000bbe-4484-4bc5-90a8-8daa6f4ab661" colab={"base_uri": "https://localhost:8080/", "height": 291}
l = df_supp.columns.values
n_cols = 5
n_rows = len(l)-1/n_cols
plt.figure(figsize=(2*n_cols+15,5*n_rows))
for i in range(0,len(l)):
plt.subplot(n_rows+1, n_cols, i+1)
sns.distplot(df_supp[l[i]],kde=True)
# + [markdown] id="R2AnFKQsDhBS" colab_type="text"
# The above figure shows that the 'Title' length spans across a considerable range and can be approximated via a Gaussian Distribution with mean at 100, i.e., most of the Title Lengths have a length around 100.
#
# .
# + [markdown] id="Sh8EU6bREDVY" colab_type="text"
# ## Exporting the final preprocessed data; to be used in the next steps
# + id="84m10dGjlAOZ" colab_type="code" colab={}
df.to_csv('./drive/My Drive/rMIDAS.csv', index = False)
# + id="ZBRt9k6Q6Ye8" colab_type="code" outputId="21a3bafb-8d5c-4849-b989-78c8240ccdb4" colab={"base_uri": "https://localhost:8080/", "height": 380}
df = pd.read_csv('./drive/My Drive/rMIDAS.csv')
print(df.shape)
df.head(10)
# + id="llYM1IPLkCo5" colab_type="code" colab={}
df_bal_2.to_csv('./drive/My Drive/rMIDAS_bal_2.csv', index = False)
| 2_EDA&PreProcessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variability analysis for HBEC IFN experiment
import scanpy as sc
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
import sys
sys.path.append('/data/home/Github/scrna-parameter-estimation/dist/memento-0.0.2-py3.7.egg')
import memento
data_path = '/data_volume/ifn_hbec/'
# ### Read the processed RNA data
#
# Focus on the club and bc/club cells and type I interferons for now.
#
# Encode the timestamps to integers.
adata_processed = sc.read(data_path + 'HBEC_type_I_processed.h5ad')
adata = sc.read(data_path + 'HBEC_type_I_filtered_counts.h5ad')
adata.obs.donor.value_counts()
adata = adata[
adata.obs.cell_type.isin(['club', 'basal/club']) & \
adata.obs.stim.isin(['control','beta'])].copy()
adata.obs.donor.value_counts()
adata.shape
# ### Perform 1D test to find genes that are generally upregulated
#
# Use all time steps
time_converter={0:0, 3:1, 6:1, 9:1, 24:1, 48:1}
adata.obs['time_step'] = adata.obs['time'].astype(int).apply(lambda x: time_converter[x])
memento.create_groups(adata, label_columns=['time_step'], inplace=True, q=0.25*0.2)
memento.compute_size_factors(adata, trim_percent=0.05)
memento.compute_1d_moments(adata, inplace=True, filter_mean_thresh=0.2, min_perc_group=.9)
memento.ht_1d_moments(
adata,
formula_like='1 + time_step',
cov_column='time_step',
num_boot=10000,
verbose=1,
num_cpus=13)
result_1d_overall = memento.get_1d_ht_result(adata)
result_1d_overall['de_fdr'] = memento.util._fdrcorrect(result_1d_overall['de_pval'])
de_genes = result_1d_overall.query('de_fdr < 0.05 & de_coef > 0 & ~gene.str.contains("MT-").values').gene.tolist()
result_1d_overall.query('gene == "HES4"')
# ### Plot the progression for these DE genes
adata = sc.read(data_path + 'HBEC_type_I_filtered_counts.h5ad')
adata = adata[adata.obs.stim.isin(['control','alpha'])].copy()
time_converter={0:0, 3:1, 6:2, 9:3, 24:4, 48:5}
adata.obs['time_step'] = adata.obs['time'].astype(int).apply(lambda x: time_converter[x])
memento.create_groups(adata, label_columns=['time_step'], inplace=True, q=0.25*0.2)
memento.compute_size_factors(adata, trim_percent=0.05)
memento.compute_1d_moments(adata, inplace=True, filter_mean_thresh=0.2, min_perc_group=.9, filter_genes=False)
moments_mean, moments_var, _ = memento.get_1d_moments(adata)
def plot_time_expr(gene):
plt.figure(figsize=(6,3));
plt.subplot(1, 2, 1);
plt.title('{} mean'.format(gene))
plt.scatter(np.arange(6), moments_mean[moments_mean.columns.sort_values()].query('gene == "{}"'.format(gene)).iloc[0, 1:].values)
plt.subplot(1, 2, 2);
plt.title('{} variability'.format(gene))
plt.scatter(np.arange(6), moments_var[moments_var.columns.sort_values()].query('gene == "{}"'.format(gene)).iloc[0, 1:].values)
# + active=""
# for gene in de_genes:
#
# plot_time_expr(gene)
# plt.savefig('figures/{}.pdf'.format(gene))
# plt.close()
# -
plt.figure(figsize=(6, 3));
plot_time_expr('IRF9');
plt.figure(figsize=(6, 3));
plot_time_expr('ISG15');
plt.figure(figsize=(6, 3));
plot_time_expr('IRF1');
moments_mean[moments_mean.columns.sort_values()].query('gene == "ISG15"')
moments_var[moments_var.columns.sort_values()].query('gene == "ISG15"')
moments_mean[moments_mean.columns.sort_values()].query('gene == "SAT1"')
moments_var[moments_var.columns.sort_values()].query('gene == "SAT1"')
adata.shape
memento.ht_1d_moments(
adata,
formula_like='1 + time_step',
cov_column='time_step',
num_boot=10000,
verbose=1,
num_cpus=13)
moments_1d = memento.get_1d_moments(adata)
moments_1d[1][moments_1d[1].columns.sort_values()].query('gene == "TXN"')
result_1d = memento.get_1d_ht_result(adata)
moments_1d[1].query('gene == "SAT1"')
result_1d.query('gene == "SAT1"')
| analysis/ifn_hbec/chipseq/variability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3
# language: python
# name: py3
# ---
import matplotlib.pyplot as plt
from ipywidgets import interact, IntSlider
# %matplotlib inline
x = IntSlider()
@interact(n=x)
def f(n):
plt.plot([0,n])
plt.show()
@interact(n=x)
def f(n):
plt.plot([0,n,0,n])
plt.show()
| notebooks/delete-me.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''nerf_pl'': conda)'
# language: python
# name: python3
# ---
# +
from pathlib import Path
import shutil
from tqdm import tqdm
import cv2
import matplotlib.pyplot as plt
import numpy as np
# -
# It is a good idea to experiment with these values
THRESHOLD = 75
# Get the images
PATH = Path("../../data/llff/deer_v6_debug_script_picked_blurry_v2")
image_filenames = list((PATH / "images").rglob("**/*.JPG"))
len(image_filenames)
def variance_of_laplacian(image, quiet=True):
# compute the Laplacian of the image and then return the focus
# measure, which is simply the variance of the Laplacian
laplacian_img = cv2.Laplacian(image, cv2.CV_64F) # has same shape as the original image
if not quiet:
_, axes = plt.subplots(1, 2, figsize=(8, 16))
axes = axes.flatten()
for img, ax in zip([image, laplacian_img*255], axes):
ax.imshow(img, cmap="gray", vmin=0, vmax=255)
plt.show()
return laplacian_img.var()
# +
blurry_image_filenames = []
focus_measures = []
# loop over the input images
for i, img_fn in enumerate(tqdm(image_filenames, total=len(image_filenames), leave=False)):
# load the image, convert it to grayscale, and compute the
# focus measure of the image using the Variance of Laplacian method
image = cv2.imread(str(img_fn))
grayscale_image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
focus_measure = variance_of_laplacian(grayscale_image)
focus_measures.append(focus_measure)
# Check if blurry
if focus_measure < THRESHOLD:
blurry_image_filenames.append(img_fn)
# -
len(blurry_image_filenames), len(blurry_image_filenames) / len(image_filenames)
plt.hist(focus_measures, bins=50)
# +
# Uncomment below when you are ready to remove the images
# for img_fn in blurry_image_filenames:
# shutil.move(img_fn, str(PATH / "bad_images" / "blurry" / img_fn.name))
# -
| notebooks/detect_blurry_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES'] = "-1"
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import pandas as pd
from tqdm.auto import tqdm
import torch
from torch import nn
import gin
import pickle
import io
from sparse_causal_model_learner_rl.trainable.gumbel_switch import WithInputSwitch, sample_from_logits_simple
gin.enter_interactive_mode()
from sparse_causal_model_learner_rl.loss.losses import fit_loss
from sparse_causal_model_learner_rl.metrics.context_rewrite import context_rewriter
from sparse_causal_model_learner_rl.sacred_gin_tune.sacred_wrapper import load_config_files
from sparse_causal_model_learner_rl.learners.rl_learner import CausalModelLearnerRL
from sparse_causal_model_learner_rl.config import Config
from keychest.features_xy import dict_to_arr, arr_to_dict, obs_features_handcoded
from causal_util import load_env
load_config_files(['../keychest/config/5x5_1f1c1k.gin', '../sparse_causal_model_learner_rl/configs/rec_nonlin_gnn_gumbel_siamese_l2.gin'])
import ray
ray.init(address='10.90.38.7:6379', ignore_reinit_error=True)
learner = CausalModelLearnerRL(Config())
learner.collect_steps()
ctx = learner._context
# +
from keychest.features_xy import dict_to_arr, arr_to_dict, obs_features_handcoded
keys = sorted(obs_features_handcoded(learner.env.engine).keys())
keys_add = learner.additional_feature_keys
# +
ox_norm = ctx['obs_x']
ox = learner.normalizers['obs'].unnormalize(ox_norm)
ax = ctx['action_x']
add_features_y = torch.cat([ctx[k] for k in keys_add], dim=1)
# -
keys, keys_add
pd.DataFrame(ax.numpy()).hist()
pd.DataFrame(ox.numpy(), columns=keys).hist()
pd.DataFrame(add_features_y.numpy(), columns=learner.additional_feature_keys).hist()
def predict_done(f):
d = arr_to_dict(arr=f, keys=keys)
d = {x: int(round(y)) for x, y in d.items()}
done = d['health'] <= 1
if (d['player__x'], d['player__y']) == (d['food__00__x'], d['food__00__y']):
done = False
return done
dones_pred = np.array([predict_done(x.numpy()) for x in ox])
dones_true = add_features_y[:, keys_add.index('done_y')].numpy()
np.mean(dones_pred - dones_true)
def predict_rew(f):
d = arr_to_dict(arr=f, keys=keys)
d = {x: int(round(y)) for x, y in d.items()}
r = 0.0
r += learner.env.reward_dict['step']
if d['player__x'] == d['food__00__x'] and d['player__y'] == d['food__00__y']:
r += learner.env.reward_dict['food_collected']
if d['player__x'] == d['key__00__x'] and d['player__y'] == d['key__00__y']:
r += learner.env.reward_dict['key_collected']
if d['player__x'] == d['chest__00__x'] and d['player__y'] == d['chest__00__y'] and d['keys'] > 0:
r += learner.env.reward_dict['chest_opened']
return r
rew_pred = np.array([predict_rew(x.numpy()) for x in ox])
rew_true = add_features_y[:, keys_add.index('rew_y')].numpy()
np.max(np.abs(rew_pred - rew_true))
| debug/keychest_gofa_reward_done.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: gan-ensembling
# language: python
# name: gan-ensembling
# ---
# # Plot paper graphs using precomputed evaluation results
# +
import sys
import numpy as np
from matplotlib import rc
import matplotlib.pyplot as plt
import sklearn.metrics
from collections import defaultdict, OrderedDict
import os
from tqdm import tqdm
import pandas as pd
import seaborn as sns
pd.options.display.float_format = '{:0.2f}'.format
rc('font', **{'family': 'serif'})
from data import data_celebahq
# %matplotlib inline
# -
# ! mkdir -p pdfs
# # utility functions
# +
### plot format utilities ###
sns.set(style='whitegrid')
sns.set_style({'font.family': 'serif'})
def save(f, filename, extra_artists=None):
f.savefig(os.path.join('pdfs', filename), bbox_inches='tight', dpi=300, bbox_extra_artists=extra_artists)
def adjust_saturation(palette, s):
new_palette = [sns.set_hls_values(color=p, h=None, l=None, s=s)
for p in palette]
return new_palette
def bar_offset(group_size, n_groups, barwidth):
# utility function to get x-axis values for grouped bar plots
xvals = np.arange(1, n_groups+1)
halfwidth = barwidth / 2
offsets = [i * barwidth for i in range(group_size)]
if group_size % 2 == 1:
middle = offsets[int(len(offsets) / 2)]
if group_size % 2 == 0:
middle = np.mean(offsets[int(len(offsets) / 2)-1:int(len(offsets) / 2)+1])
offsets = [off - middle for off in offsets]
return [xvals + off for off in offsets]
def get_list_stats(l):
mean = np.mean(l)
stderr = np.std(l) / np.sqrt(len(l))
n = len(l)
return {'mean': mean, 'stderr': stderr, 'n': n}
def make_green_palette(n):
return sns.light_palette([0.39215686, 0.61960784, 0.45098039], n_colors=n)
def make_blue_palette(n):
return sns.light_palette([0.29803922, 0.44705882, 0.69019608], n_colors=n)
def make_purple_palette(n):
return sns.light_palette([0.5058823529411764, 0.4470588235294118, 0.7019607843137254], n_colors=n)
def make_yellow_palette(n):
return sns.light_palette([0.8666666666666667, 0.5176470588235295, 0.3215686274509804], n_colors=n)
def make_diverging_palette(n):
return sns.color_palette("vlag", n_colors=n)
# +
### data evaluation utilities ###
def softmax_to_prediction(softmax_prediction):
# converts softmax prediction to discrete class label
if np.ndim(softmax_prediction) == 2:
# N x ensembles binary prediction
return (softmax_prediction > 0.5).astype(int)
elif np.ndim(softmax_prediction) == 3:
# N x ensembles x classes
return np.argmax(softmax_prediction, axis=-1).squeeze()
else:
assert(False)
def get_accuracy_from_image_ensembles(data_file, key, resample=False, seed=0,
n_resamples=20, ens_size=32, verbose=True):
# helper function to extract ensembled accuracy from image augmentations
# e.g. image_ensemble_imcolor.npz or image_ensemble_imcrop.npz
encoded_data = np.load(data_file)
preds_original = softmax_to_prediction(encoded_data['original'])
acc_original = sklearn.metrics.accuracy_score(encoded_data['label'], preds_original) * 100
jitters = np.concatenate([encoded_data['original'], encoded_data[key]], axis=1)
jitters = np.mean(jitters, axis=1, keepdims=True)
preds_ensembled = softmax_to_prediction(jitters)
acc_ensembled = sklearn.metrics.accuracy_score(encoded_data['label'], preds_ensembled) * 100
resamples = None
if resample:
# sample num_samples batches with replacement, compute accuracy
resamples = []
rng = np.random.RandomState(seed)
jitters = np.concatenate([encoded_data['original'], encoded_data[key]], axis=1)
assert(jitters.shape[1] == ens_size) # sanity check
for i in range(n_resamples):
if verbose:
print('*', end='')
indices = rng.choice(jitters.shape[1], ens_size, replace=True)
jitters_resampled = jitters[:, indices]
jitters_resampled = np.mean(jitters_resampled, axis=1, keepdims=True)
preds_ensembled = softmax_to_prediction(jitters_resampled)
resamples.append(sklearn.metrics.accuracy_score(encoded_data['label'], preds_ensembled) * 100)
if verbose:
print("done")
return {'acc_original': acc_original, 'acc_ensembled': acc_ensembled, 'resamples': resamples}
def sample_ensemble(raw_preds, ens_size=None, seed=None):
# helper function to resample raw ensemble predictions
# raw_preds = N x ens_size for binary classification, or N x ens_size x classes
# ens_size = number of samples to take preds for ensembling, None takes all all samples
# seed = random seed to use when sampling with replacement, None takes samples in order
if ens_size is None:
ens_size = raw_preds.shape[1] # take all samples
if seed is None:
ensemble_preds = raw_preds[:, range(ens_size)] # take the samples in order
else: # sample the given preds with replacement
rng = np.random.RandomState(seed)
indices = rng.choice(raw_preds.shape[1], ens_size, replace=True)
ensemble_preds = raw_preds[:, indices]
return ensemble_preds
def get_accuracy_from_npz(data_file, expt_name, weight=None, ens_size=None, seed=None, return_preds=False,
add_aug=False, aug_name='image_ensemble_imcrop', aug_key='imcrop'):
# compute weighted accuracies combining original image and GAN reconstructions from an npz_file
# option to use either single original image, or multiple image augmentations for the image views
# setup
encoded_data = np.load(data_file)
df = defaultdict(list)
expt_settings = os.path.basename(data_file).split('.')[0]
if weight is not None:
weights = [weight]
else:
weights = np.linspace(0, 1, 21)
# determine image classification accuracy
if not add_aug:
# basic case: just load the image predictions from the data file
preds_original = softmax_to_prediction(encoded_data['original'])
original = encoded_data['original'] # full softmax distribution
else:
# ensemble also with the image augmentations data
print('.', end='')
im_aug_data = np.load(os.path.join(data_file.rsplit('/', 1)[0], '%s.npz' % aug_name))
im_aug_ens = np.concatenate([im_aug_data['original'], im_aug_data[aug_key]], axis=1)
im_aug_ens = sample_ensemble(im_aug_ens, ens_size, seed)
im_aug_ens = np.mean(im_aug_ens, axis=1, keepdims=True)
preds_original = softmax_to_prediction(im_aug_ens)
original = im_aug_ens # full softmax distribution
acc_original = sklearn.metrics.accuracy_score(encoded_data['label'], preds_original) * 100
# determine GAN reconstruction accuracy
preds_reconstructed = softmax_to_prediction(encoded_data['reconstructed'])
acc_reconstructed = sklearn.metrics.accuracy_score(encoded_data['label'], preds_reconstructed) * 100
# determine GAN ensemble accuracy
perturbed = encoded_data[expt_name] # N x ens_size x softmax distribution
gan_ens = np.concatenate((encoded_data['reconstructed'], perturbed), axis=1)
if ens_size == 0:
gan_ens = original # dummy case: don't use gan reconstructed images
else:
gan_ens = sample_ensemble(gan_ens, ens_size, seed)
for weight in weights: # alpha weighting hyperparameter
# for binary classification: original.shape = N x 1, gan_ens.shape = N x ens_size
# for multi-class classification: original.shape = N x 1 x classes; gan_ens.shape = N x ens_size x classes
ensembled = (1-weight) * original + weight * np.mean(gan_ens, axis=1, keepdims=True)
preds_ensembled = softmax_to_prediction(ensembled)
acc_ensembled = sklearn.metrics.accuracy_score(encoded_data['label'], preds_ensembled) * 100
df['acc'].append(acc_ensembled)
df['weight'].append(weight)
df['expt_name'].append(expt_name)
# table of expt_name x weight
df = pd.DataFrame.from_dict(df)
return_data = {'expt_settings': expt_settings,
'acc_original': acc_original,
'acc_reconstructed': acc_reconstructed,
'ensemble_table': df}
if return_preds:
assert(len(weights) == 1)
return_preds = {
'original': original, # original softmax
'reconstruction': gan_ens, # softmax of all gan views
'ensembled': ensembled, # softmax of the weighted ensemble
'pred_original': preds_original,
'pred_reconstruction': preds_reconstructed,
'pred_ensemble': preds_ensembled,
'label': encoded_data['label'],
}
return return_data, return_preds
return return_data
def compute_best_weight(val_data_file, test_data_file, expt_name,
verbose=True, ens_size=None, seed=None,
add_aug=False, aug_name='image_ensemble_imcrop', aug_key='imcrop'):
# given a val data file and a test data file, find the best weighting between
# image view and GAN-generated views on the val split, and use that weighting on the test split
# sanity checks
assert('val' in val_data_file)
assert('test' in test_data_file)
val_accuracy_info = get_accuracy_from_npz(val_data_file, expt_name,
weight=None, ens_size=ens_size, seed=seed,
add_aug=add_aug, aug_name=aug_name, aug_key=aug_key)
val_ensemble_table = val_accuracy_info['ensemble_table']
# find the optimal ensemble weight from validation
best_val_setting = val_ensemble_table.iloc[val_ensemble_table['acc'].argsort().iloc[-1], :]
if verbose:
print("Val original %0.4f Val reconstructed %0.4f" %
(val_accuracy_info['acc_original'], val_accuracy_info['acc_reconstructed']))
print("%0.4f @ %0.4f %s" % (best_val_setting['acc'], best_val_setting['weight'], best_val_setting['expt_name']))
test_accuracy_info = get_accuracy_from_npz(test_data_file, expt_name,
weight=best_val_setting['weight'],
ens_size=ens_size, seed=seed,
add_aug=add_aug, aug_name=aug_name, aug_key=aug_key)
test_ensemble_table = test_accuracy_info['ensemble_table']
assert(test_ensemble_table.shape[0] == 1) # it should only evaluate at the specified weight
test_setting_from_val = test_ensemble_table.iloc[0, :] # gets the single element from the table
if verbose:
print("Test original %0.4f Test reconstructed %0.4f" %
(test_accuracy_info['acc_original'], test_accuracy_info['acc_reconstructed']))
print("%0.4f @ %0.4f %s" % (test_setting_from_val['acc'], test_setting_from_val['weight'],
test_setting_from_val['expt_name']))
return {'val_info': val_accuracy_info, 'test_info': test_accuracy_info,
'val_setting': best_val_setting, 'test_setting': test_setting_from_val}
def resample_wrapper(val_file, test_file, expt_name, ens_size, add_aug, n_resamples=20, verbose=False,
aug_name='image_ensemble_imcrop', aug_key='imcrop'):
# due to randomness in sampling, it helps to sample multiple times and average the results for stability
# this function wraps compute_best_weight(), using the specified ensemble size and resampling multiple times
val_samples = []
test_samples = []
weights = []
assert(ens_size==31 or (ens_size==16 and add_aug==True))
# using ens_size=31 so that with the original image, total size=32; or 16 image views and 16 GAN views
for s in range(n_resamples):
res = compute_best_weight(val_file, test_file, expt_name, verbose=verbose, add_aug=add_aug,
ens_size=ens_size, seed=s, aug_name=aug_name, aug_key=aug_key)
val_samples.append(res['val_setting']['acc'])
test_samples.append(res['test_setting']['acc'])
weights.append(res['test_setting']['weight'])
return {'val_avg': np.mean(val_samples),
'test_avg': np.mean(test_samples),
'val_stderr': np.std(val_samples) / np.sqrt(n_resamples),
'test_stderr': np.std(test_samples) / np.sqrt(n_resamples),
'weights': weights,
'val_acc_original': res['val_info']['acc_original'],
'test_acc_original': res['test_info']['acc_original'],
'val_acc_rec': res['val_info']['acc_reconstructed'],
'test_acc_rec': res['test_info']['acc_reconstructed'],
}
# -
# # cars domain
# +
# sample 32 crops of images, compare to combination of 16 crops of images and 16 crops of gan
df = defaultdict(list)
for i, classifier in enumerate(['imageclassifier', 'latentclassifier',
'latentclassifier_stylemix_fine']):
print(classifier)
val_expts = [
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_isotropic_coarse_tensortransform.npz',
('isotropic_coarse_1.00', 'isotropic_coarse_1.50', 'isotropic_coarse_2.00'), 'Isotropic Coarse'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_isotropic_fine_tensortransform.npz',
('isotropic_fine_0.30', 'isotropic_fine_0.50', 'isotropic_fine_0.70'), 'Isotropic Fine'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_pca_coarse_tensortransform.npz',
('pca_coarse_1.00', 'pca_coarse_2.00', 'pca_coarse_3.00'), 'PCA Coarse'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_pca_fine_tensortransform.npz',
('pca_fine_1.00', 'pca_fine_2.00', 'pca_fine_3.00'), 'PCA Fine'),
# (f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_stylemix_coarse_tensortransform.npz',
# ('stylemix_coarse',), 'Style-mix Coarse'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_stylemix_fine_tensortransform.npz',
('stylemix_fine',), 'Style-mix Fine'),
]
test_expts = [(x.replace('_val/', '_test/'), y, z) for x, y, z in val_expts]
for val, test in zip(val_expts, test_expts):
expt_settings = []
print(val[-1])
for expt_name in val[1]:
resampled_accs = resample_wrapper(val[0], test[0], expt_name, ens_size=16,
add_aug=True, aug_name='image_ensemble_imcrop', verbose=False)
resampled_accs['expt_name'] = expt_name
expt_settings.append(resampled_accs)
print("done")
best_expt = max(expt_settings, key=lambda x: x['val_avg']) # take the val accuracy, avged over samples
df['classifier'].append(classifier+'_crop')
df['acc'].append(best_expt['test_avg'])
df['stderr'].append(best_expt['test_stderr'])
df['expt'].append(best_expt['expt_name'])
df['expt_group'].append(test[2])
df = pd.DataFrame.from_dict(df)
# -
df
# +
# plot it
f, ax = plt.subplots(1, 1, figsize=(7, 5))
data_file = f'results/precomputed_evaluations/car/output/imageclassifier_test/image_ensemble_imcrop.npz'
im_crops = get_accuracy_from_image_ensembles(data_file, 'imcrop', resample=True)
group_size = 5
bar_width=0.15
n_groups = 3
bar_offsets = bar_offset(group_size, n_groups, bar_width)
palette = make_blue_palette(3)[1:] + make_green_palette(3)[1:] + make_purple_palette(3)[1:]
resample_stats = get_list_stats(im_crops['resamples'])
ind = 0.2
ax.axhline(im_crops['acc_ensembled'], color='k', linestyle=':', label='Original Images')
xticklabels = []
for i in range(group_size):
indices = np.arange(i, n_groups*group_size, group_size)
bar_height = df.iloc[indices]['acc']
bar_err = df.iloc[indices]['stderr']
assert(all([x == df.iloc[indices[0]]['expt_group'] for x in df.iloc[indices]['expt_group']]))
ax.bar(bar_offsets[i], bar_height, width=bar_width, color=palette[i], yerr=bar_err,
label=df.iloc[indices[0]]['expt_group'], edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels.append(df.iloc[indices[0]]['classifier'].replace('_', '\n'))
ax.set_ylim([94, 99])
ax.set_xticks(list(range(1, n_groups+1)))
handles,labels = ax.get_legend_handles_labels()
# reorder it so it looks nicer
order = [0, 3, 1, 4, 2, 5]
handles = [handles[i] for i in order]
labels = [labels[i] for i in order]
ax.legend(handles, labels, loc='upper center', ncol=3, prop={'size': 11})
# ax.legend(handles, labels, loc='upper center', bbox_to_anchor=(0.5, -0.3), ncol=3, prop={'size': 11})
# ax.legend(loc='center left', bbox_to_anchor=(1, 0.5), prop={'size': 14})
ax.set_xticklabels(['Original\nImages', 'GAN\nRecontructions', 'Style-mix Fine\nAugmentations'], fontsize=12)
ax.set_xlabel('Classifier training distribution', fontsize=16)
ax.set_ylabel('Classification Accuracy', fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(14)
ax.set_title('Cars', fontsize=16)
f.tight_layout()
save(f, 'graph_cars_v2.pdf')
# +
# sample 32 crops of images, compare to combination of 16 crops of images and 16 crops of gan
# using all experiment settings for supplemental
df = defaultdict(list)
im_crop_data = []
for i, classifier in enumerate(['imageclassifier', 'latentclassifier',
'latentclassifier_isotropic_fine', 'latentclassifier_isotropic_coarse',
'latentclassifier_pca_fine', 'latentclassifier_pca_coarse',
'latentclassifier_stylemix_fine', 'latentclassifier_stylemix_coarse']):
print(classifier)
val_expts = [
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_isotropic_coarse_tensortransform.npz',
('isotropic_coarse_1.00', 'isotropic_coarse_1.50', 'isotropic_coarse_2.00'), 'Isotropic Coarse'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_isotropic_fine_tensortransform.npz',
('isotropic_fine_0.30', 'isotropic_fine_0.50', 'isotropic_fine_0.70'), 'Isotropic Fine'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_pca_coarse_tensortransform.npz',
('pca_coarse_1.00', 'pca_coarse_2.00', 'pca_coarse_3.00'), 'PCA Coarse'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_pca_fine_tensortransform.npz',
('pca_fine_1.00', 'pca_fine_2.00', 'pca_fine_3.00'), 'PCA Fine'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_stylemix_coarse_tensortransform.npz',
('stylemix_coarse',), 'Style-mix Coarse'),
(f'results/precomputed_evaluations/car/output/{classifier}_val/gan_ensemble_stylemix_fine_tensortransform.npz',
('stylemix_fine',), 'Style-mix Fine'),
]
test_expts = [(x.replace('_val/', '_test/'), y, z) for x, y, z in val_expts]
data_file = f'results/precomputed_evaluations/car/output/{classifier}_test/image_ensemble_imcrop.npz'
im_crop_data.append(get_accuracy_from_image_ensembles(data_file, 'imcrop', resample=True))
for val, test in zip(val_expts, test_expts):
expt_settings = []
print(val[-1])
for expt_name in val[1]:
resampled_accs = resample_wrapper(val[0], test[0], expt_name, ens_size=16,
add_aug=True, aug_name='image_ensemble_imcrop', verbose=False)
resampled_accs['expt_name'] = expt_name
expt_settings.append(resampled_accs)
print("done")
best_expt = max(expt_settings, key=lambda x: x['val_avg']) # take the val accuracy, avged over samples
df['classifier'].append(classifier+'_crop')
df['acc'].append(best_expt['test_avg'])
df['stderr'].append(best_expt['test_stderr'])
df['expt'].append(best_expt['expt_name'])
df['expt_group'].append(test[2])
df = pd.DataFrame.from_dict(df)
# -
df
# +
# plot it
f, ax = plt.subplots(1, 1, figsize=(14, 6))
group_size = 8
bar_width=0.1
n_groups = 8
bar_offsets = bar_offset(group_size, n_groups, bar_width)
palette = make_yellow_palette(3)[1:] + make_blue_palette(3)[1:] + make_green_palette(3)[1:] + make_purple_palette(3)[1:]
# resample_stats = get_list_stats(im_crops['resamples'])
ind = 0.2
# ax.axhline(im_crops['acc_ensembled'], color='k', linestyle=':', label='Original Images')
ax.bar(bar_offsets[0], [x['acc_original'] for x in im_crop_data], width=bar_width, color=palette[0],
label='Image Single Crop', edgecolor=(0.5, 0.5, 0.5), capsize=5)
ax.bar(bar_offsets[1], [get_list_stats(x['resamples'])['mean'] for x in im_crop_data],
width=bar_width, color=palette[1], yerr=[get_list_stats(x['resamples'])['stderr'] for x in im_crop_data],
label='Image Multi Crop', edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels = []
for i in range(6):
indices = np.arange(i, n_groups*6, 6)
bar_height = df.iloc[indices]['acc']
bar_err = df.iloc[indices]['stderr']
assert(all([x == df.iloc[indices[0]]['expt_group'] for x in df.iloc[indices]['expt_group']]))
ax.bar(bar_offsets[i+2], bar_height, width=bar_width, color=palette[i+2], yerr=bar_err,
label=df.iloc[indices[0]]['expt_group'], edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels.append(df.iloc[indices[0]]['classifier'].replace('_', '\n'))
ax.set_ylim([94, 100])
ax.set_xticks(list(range(1, n_groups+1)))
handles,labels = ax.get_legend_handles_labels()
ax.legend(handles, labels, loc='upper center', ncol=4, prop={'size': 11})
# ax.legend(handles, labels, loc='upper center', bbox_to_anchor=(0.5, -0.2), ncol=4, prop={'size': 11})
ax.set_xticklabels(['Original\nImages', 'GAN\nRecontructions',
'Isotropic Fine\nAugmentations', 'Isotropic Coarse\nAugmentations',
'PCA Fine\nAugmentations', 'PCA Coarse\nAugmentations',
'Style-mix Fine\nAugmentations', 'Style-mix Coarse\nAugmentations'], fontsize=12)
ax.set_xlabel('Classifier training distribution', fontsize=16)
ax.set_ylabel('Classification Accuracy', fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(14)
ax.set_title('Cars', fontsize=16)
f.tight_layout()
save(f, 'sm_graph_cars_all_settings.pdf')
# -
# # # cat face classifier
# +
# figure for main: cat face augmentations (does not use crop)
df = defaultdict(list)
for i, classifier in enumerate(['imageclassifier', 'latentclassifier',
'latentclassifier_stylemix_coarse']):
print(classifier)
val_expts = [
# also tried without _tensortransform, it's similar
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_isotropic_coarse_tensortransform.npz',
('isotropic_coarse_0.50', 'isotropic_coarse_0.70', 'isotropic_coarse_1.00'), 'Isotropic Coarse'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_isotropic_fine_tensortransform.npz',
('isotropic_fine_0.10', 'isotropic_fine_0.20', 'isotropic_fine_0.30'), 'Isotropic Fine'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_pca_coarse_tensortransform.npz',
('pca_coarse_0.50', 'pca_coarse_0.70', 'pca_coarse_1.00'), 'PCA Coarse'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_pca_fine_tensortransform.npz',
('pca_fine_0.50', 'pca_fine_0.70', 'pca_fine_1.00'), 'PCA Fine'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_stylemix_coarse_tensortransform.npz',
('stylemix_coarse',), 'Style-mix Coarse'),
# (f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_stylemix_fine_tensortransform.npz',
# ('stylemix_fine',), 'Style-mix Fine'),
]
test_expts = [(x.replace('_val/', '_test/'), y, z) for x, y, z in val_expts]
for val, test in zip(val_expts, test_expts):
expt_settings = []
print(val[-1])
for expt_name in val[1]:
resampled_accs = resample_wrapper(val[0], test[0], expt_name, ens_size=31,
add_aug=False, verbose=False)
resampled_accs['expt_name'] = expt_name
expt_settings.append(resampled_accs)
print("done")
best_expt = max(expt_settings, key=lambda x: x['val_avg']) # take the val accuracy, avged over samples
df['classifier'].append(classifier+'_crop')
df['acc'].append(best_expt['test_avg'])
df['stderr'].append(best_expt['test_stderr'])
df['expt'].append(best_expt['expt_name'])
df['expt_group'].append(test[2])
df = pd.DataFrame.from_dict(df)
# -
df
# +
# plot it
f, ax = plt.subplots(1, 1, figsize=(7, 5))
data_file = f'results/precomputed_evaluations/cat/output/imageclassifier_test/image_ensemble_imcrop.npz'
im_s = get_accuracy_from_image_ensembles(data_file, 'imcrop', resample=True)
group_size = 5
bar_width=0.15
n_groups = 3
bar_offsets = bar_offset(group_size, n_groups, bar_width)
palette = make_blue_palette(3)[1:] + make_green_palette(3)[1:] + make_purple_palette(3)[1:]
resample_stats = get_list_stats(im_s['resamples'])
ind = 0.2
# note: using acc_original here, as it's better
ax.axhline(im_s['acc_original'], color='k', linestyle=':', label='Original Images')
xticklabels = []
for i in range(group_size):
indices = np.arange(i, n_groups*group_size, group_size)
bar_height = df.iloc[indices]['acc']
bar_err = df.iloc[indices]['stderr']
assert(all([x == df.iloc[indices[0]]['expt_group'] for x in df.iloc[indices]['expt_group']]))
ax.bar(bar_offsets[i], bar_height, width=bar_width, color=palette[i], yerr=bar_err,
label=df.iloc[indices[0]]['expt_group'], edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels.append(df.iloc[indices[0]]['classifier'].replace('_', '\n'))
ax.set_ylim([90, 95])
ax.set_xticks(list(range(1, n_groups+1)))
handles,labels = ax.get_legend_handles_labels()
# reorder it so it looks nicer
order = [0, 3, 1, 4, 2, 5]
handles = [handles[i] for i in order]
labels = [labels[i] for i in order]
ax.legend(handles, labels, loc='upper center', ncol=3, prop={'size': 10.8})
# ax.legend(handles, labels, loc='upper center', bbox_to_anchor=(0.5, -0.3), ncol=3, prop={'size': 11})
# ax.legend(loc='center left', bbox_to_anchor=(1, 0.5), prop={'size': 14})
ax.set_xticklabels(['Original\nImages', 'GAN\nRecontructions', 'Style-mix Coarse\nAugmentations'], fontsize=12)
ax.set_xlabel('Classifier training distribution', fontsize=16)
ax.set_ylabel('Classification Accuracy', fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(14)
ax.set_title('Cats', fontsize=16)
f.tight_layout()
save(f, 'graph_cats_v2.pdf')
# +
# all settings for the supplemental
df = defaultdict(list)
im_crop_data = []
for i, classifier in enumerate(['imageclassifier', 'latentclassifier',
'latentclassifier_isotropic_fine', 'latentclassifier_isotropic_coarse',
'latentclassifier_pca_fine', 'latentclassifier_pca_coarse',
'latentclassifier_stylemix_fine', 'latentclassifier_stylemix_coarse']):
print(classifier)
val_expts = [
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_isotropic_coarse_tensortransform.npz',
('isotropic_coarse_0.50', 'isotropic_coarse_0.70', 'isotropic_coarse_1.00'), 'Isotropic Coarse'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_isotropic_fine_tensortransform.npz',
('isotropic_fine_0.10', 'isotropic_fine_0.20', 'isotropic_fine_0.30'), 'Isotropic Fine'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_pca_coarse_tensortransform.npz',
('pca_coarse_0.50', 'pca_coarse_0.70', 'pca_coarse_1.00'), 'PCA Coarse'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_pca_fine_tensortransform.npz',
('pca_fine_0.50', 'pca_fine_0.70', 'pca_fine_1.00'), 'PCA Fine'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_stylemix_coarse_tensortransform.npz',
('stylemix_coarse',), 'Style-mix Coarse'),
(f'results/precomputed_evaluations/cat/output/{classifier}_val/gan_ensemble_stylemix_fine_tensortransform.npz',
('stylemix_fine',), 'Style-mix Fine'),
]
test_expts = [(x.replace('_val/', '_test/'), y, z) for x, y, z in val_expts]
data_file = f'results/precomputed_evaluations/cat/output/{classifier}_test/image_ensemble_imcrop.npz'
im_crop_data.append(get_accuracy_from_image_ensembles(data_file, 'imcrop', resample=True))
for val, test in zip(val_expts, test_expts):
expt_settings = []
print(val[-1])
for expt_name in val[1]:
resampled_accs = resample_wrapper(val[0], test[0], expt_name, ens_size=31,
add_aug=False, verbose=False)
resampled_accs['expt_name'] = expt_name
expt_settings.append(resampled_accs)
print("done")
best_expt = max(expt_settings, key=lambda x: x['val_avg']) # take the val accuracy, avged over samples
df['classifier'].append(classifier)
df['acc'].append(best_expt['test_avg'])
df['stderr'].append(best_expt['test_stderr'])
df['expt'].append(best_expt['expt_name'])
df['expt_group'].append(test[2])
df = pd.DataFrame.from_dict(df)
# -
df
# +
# plot it
f, ax = plt.subplots(1, 1, figsize=(14, 6))
group_size = 8
bar_width=0.1
n_groups = 8
bar_offsets = bar_offset(group_size, n_groups, bar_width)
palette = make_yellow_palette(3)[1:] + make_blue_palette(3)[1:] + make_green_palette(3)[1:] + make_purple_palette(3)[1:]
ind = 0.2
# ax.axhline(im_crops['acc_ensembled'], color='k', linestyle=':', label='Original Images')
ax.bar(bar_offsets[0], [x['acc_original'] for x in im_crop_data], width=bar_width, color=palette[0],
label='Image Single Crop', edgecolor=(0.5, 0.5, 0.5), capsize=5)
ax.bar(bar_offsets[1], [get_list_stats(x['resamples'])['mean'] for x in im_crop_data],
width=bar_width, color=palette[1], yerr=[get_list_stats(x['resamples'])['stderr'] for x in im_crop_data],
label='Image Multi Crop', edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels = []
for i in range(6):
indices = np.arange(i, n_groups*6, 6)
bar_height = df.iloc[indices]['acc']
bar_err = df.iloc[indices]['stderr']
assert(all([x == df.iloc[indices[0]]['expt_group'] for x in df.iloc[indices]['expt_group']]))
ax.bar(bar_offsets[i+2], bar_height, width=bar_width, color=palette[i+2], yerr=bar_err,
label=df.iloc[indices[0]]['expt_group'], edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels.append(df.iloc[indices[0]]['classifier'].replace('_', '\n'))
ax.set_ylim([90, 94])
ax.set_xticks(list(range(1, n_groups+1)))
handles,labels = ax.get_legend_handles_labels()
ax.legend(handles, labels, loc='upper center', ncol=4, prop={'size': 11})
# ax.legend(handles, labels, loc='upper center', bbox_to_anchor=(0.5, -0.2), ncol=4, prop={'size': 11})
ax.set_xticklabels(['Original\nImages', 'GAN\nRecontructions',
'Isotropic Fine\nAugmentations', 'Isotropic Coarse\nAugmentations',
'PCA Fine\nAugmentations', 'PCA Coarse\nAugmentations',
'Style-mix Fine\nAugmentations', 'Style-mix Coarse\nAugmentations'], fontsize=12)
ax.set_xlabel('Classifier training distribution', fontsize=16)
ax.set_ylabel('Classification Accuracy', fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(14)
ax.set_title('Cats', fontsize=16)
f.tight_layout()
save(f, 'sm_graph_cats_all_settings.pdf')
# -
# # stylegan faces 40 attributes
# +
attr_mean = data_celebahq.attr_celebahq.mean(axis=0)[:-1]
attr_order = sorted([(abs(v-0.5), v, k) for k, v in attr_mean.to_dict().items()])
table_dict = OrderedDict([])
table_accs = OrderedDict([])
for i, (_, _, attr) in enumerate(tqdm(attr_order[:40])):
# print('========== %s ==========' % attr)
# gan jitter
val_file = f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_fine.npz'
test_file = f'results/precomputed_evaluations/celebahq/output/{attr}_test/gan_ensemble_stylemix_fine.npz'
expt_name = 'stylemix_fine'
# resample
resampled_accs = resample_wrapper(val_file, test_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
val_orig = resampled_accs['val_acc_original']
val_top1 = resampled_accs['val_avg']
test_orig = resampled_accs['test_acc_original']
test_top1_from_val = resampled_accs['test_avg']
# gan jitter with color/crop jitter
val_file = f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_fine_tensortransform.npz'
test_file = f'results/precomputed_evaluations/celebahq/output/{attr}_test/gan_ensemble_stylemix_fine_tensortransform.npz'
expt_name = 'stylemix_fine'
# resample
resampled_accs = resample_wrapper(val_file, test_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
val_orig_mix = resampled_accs['val_acc_original']
val_top1_mix = resampled_accs['val_avg']
test_orig_mix = resampled_accs['test_acc_original']
test_top1_from_val_mix = resampled_accs['test_avg']
# color jitter
val_file = f'results/precomputed_evaluations/celebahq/output/{attr}_val/image_ensemble_imcolor.npz'
im_ensemble = get_accuracy_from_image_ensembles(val_file, 'imcolor', resample=True, verbose=False)
val_color_orig = im_ensemble['acc_original']
val_color_ens = np.mean(im_ensemble['resamples']) # im_ensemble['acc_ensembled']
test_file = f'results/precomputed_evaluations/celebahq/output/{attr}_test/image_ensemble_imcolor.npz'
im_ensemble = get_accuracy_from_image_ensembles(test_file, 'imcolor', resample=True, verbose=False)
test_color_orig = im_ensemble['acc_original']
test_color_ens = np.mean(im_ensemble['resamples']) # im_ensemble['acc_ensembled']
# crop jitter
val_file = f'results/precomputed_evaluations/celebahq/output/{attr}_val/image_ensemble_imcrop.npz'
im_ensemble = get_accuracy_from_image_ensembles(val_file, 'imcrop', resample=True, verbose=False)
val_crop_orig = im_ensemble['acc_original']
val_crop_ens = np.mean(im_ensemble['resamples']) # im_ensemble['acc_ensembled']
test_file = f'results/precomputed_evaluations/celebahq/output/{attr}_test/image_ensemble_imcrop.npz'
im_ensemble = get_accuracy_from_image_ensembles(test_file, 'imcrop', resample=True, verbose=False)
test_crop_orig = im_ensemble['acc_original']
test_crop_ens = np.mean(im_ensemble['resamples']) # im_ensemble['acc_ensembled']
# sanity check
assert(test_color_orig == test_orig)
assert(test_crop_orig == test_orig)
assert(test_orig_mix == test_orig)
assert(val_color_orig == val_orig)
assert(val_crop_orig == val_orig)
assert(val_orig_mix == val_orig)
val_labels = ['Val Orig', 'Val Color', 'Val Crop', 'Val GAN', 'Val Combined']
val_values = [val_orig, val_color_ens, val_crop_ens, val_top1, val_top1_mix]
val_diffs = [x - val_values[0] for x in val_values]
test_labels = ['Test Orig', 'Test Color', 'Test Crop', 'Test GAN', 'Test Combined']
test_values = [test_orig, test_color_ens, test_crop_ens, test_top1_from_val, test_top1_from_val_mix]
test_diffs = [x - test_values[0] for x in test_values]
table_dict[attr] = val_diffs + test_diffs
table_accs[attr] = val_values + test_values
# -
table = pd.DataFrame.from_dict(table_dict, orient='index', columns=val_labels+test_labels)
table = table.append(table.mean(axis=0).rename('Avg'))
std = table.iloc[:-1, :].std(axis=0).rename('Std')
print(std / np.sqrt(40))
display(table.iloc[-1:, :])
table_acc = pd.DataFrame.from_dict(table_accs, orient='index', columns=val_labels+test_labels)
table_acc = table_acc.append(table_acc.mean(axis=0).rename('Avg'))
std_acc = table_acc.iloc[:-1, :].std(axis=0).rename('Std')
print(std_acc / np.sqrt(40))
display(table_acc.iloc[-1:, :])
df = table_acc.iloc[[-1], 5:].T
df = df.reset_index()
display(df)
f, ax = plt.subplots(1, 1, figsize=(6, 3))
palette = adjust_saturation(make_blue_palette(3), 0.3)
ax.bar(np.arange(len(df)), df.loc[:, 'Avg'], color=palette[-1], edgecolor=(0.5, 0.5, 0.5))
ax.set_ylim([88.5, 89.5])
ax.set_xticks(range(5))
ax.set_xticklabels(['Single\nImage', 'Color\nJitter', 'Crop\nJitter', 'Style-mix\nJitter', 'Combined\nJitter'],
fontsize=12)
ax.set_ylabel('Classification Accuracy', fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(12)
ax.set_xlabel('')
ax.set_xlim([-0.7, 4.7])
save(f, 'graph_face_testaug.pdf')
# +
f, ax = plt.subplots(1, 1, figsize=(6, 3))
diffs = table.iloc[:-1, 5:]
bar_height = diffs.mean(axis=0)
bar_err = diffs.std(axis=0) / np.sqrt(diffs.shape[0])
palette = adjust_saturation(make_blue_palette(3), 0.3)
ax.bar(range(5), bar_height, edgecolor=(0.5, 0.5, 0.5), yerr=bar_err, color=palette[-1], capsize=5)
ax.set_xticks(range(5))
ax.set_xticklabels(['Single\nImage', 'Color\nJitter', 'Crop\nJitter', 'Style-mix\nJitter', 'Combined\nJitter'],
fontsize=12)
ax.set_ylabel('Accuracy Difference', fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(12)
ax.set_xlabel('')
ax.set_xlim([-0.7, 4.7])
ax.set_ylim([-0.1, 0.2])
save(f, 'graph_face_testaug_diffs.pdf')
# -
# # stylegan idinvert
# +
attr_mean = data_celebahq.attr_celebahq.mean(axis=0)[:-1]
attr_order = sorted([(abs(v-0.5), v, k) for k, v in attr_mean.to_dict().items()])
table_dict = OrderedDict([])
table_accs = OrderedDict([])
for i, (_, _, attr) in enumerate(tqdm(attr_order[:40])):
# print('========== %s ==========' % attr)
# gan jitter
val_file = f'results/precomputed_evaluations/celebahq-idinvert/output/{attr}_val/gan_ensemble_stylemix_fine.npz'
test_file = f'results/precomputed_evaluations/celebahq-idinvert/output/{attr}_test/gan_ensemble_stylemix_fine.npz'
expt_name = 'stylemix_fine'
# resample
resampled_accs = resample_wrapper(val_file, test_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
val_orig = resampled_accs['val_acc_original']
val_top1 = resampled_accs['val_avg']
test_orig = resampled_accs['test_acc_original']
test_top1_from_val = resampled_accs['test_avg']
# gan jitter with color/crop jitter
val_file = f'results/precomputed_evaluations/celebahq-idinvert/output/{attr}_val/gan_ensemble_stylemix_fine_tensortransform.npz'
test_file = f'results/precomputed_evaluations/celebahq-idinvert/output/{attr}_test/gan_ensemble_stylemix_fine_tensortransform.npz'
expt_name = 'stylemix_fine'
# resample
resampled_accs = resample_wrapper(val_file, test_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
val_orig_mix = resampled_accs['val_acc_original']
val_top1_mix = resampled_accs['val_avg']
test_orig_mix = resampled_accs['test_acc_original']
test_top1_from_val_mix = resampled_accs['test_avg']
# sanity check
assert(test_orig_mix == test_orig)
assert(val_orig_mix == val_orig)
val_labels = ['Val Orig', 'Val GAN', 'Val Combined']
val_values = [val_orig, val_top1, val_top1_mix]
val_diffs = [x - val_values[0] for x in val_values]
test_labels = ['Test Orig', 'Test GAN', 'Test Combined']
test_values = [test_orig,test_top1_from_val, test_top1_from_val_mix]
test_diffs = [x - test_values[0] for x in test_values]
table_dict[attr] = val_diffs + test_diffs
table_accs[attr] = val_values + test_values
# +
table_idinvert = pd.DataFrame.from_dict(table_dict, orient='index', columns=val_labels+test_labels)
table_idinvert = table_idinvert.append(table_idinvert.mean(axis=0).rename('Avg'))
std = table_idinvert.iloc[:-1, :].std(axis=0).rename('Std')
print(std / np.sqrt(40))
display(table_idinvert.iloc[-1:, :])
# -
table_idinvert_acc = pd.DataFrame.from_dict(table_accs, orient='index', columns=val_labels+test_labels)
table_idinvert_acc = table_idinvert_acc.append(table_idinvert_acc.mean(axis=0).rename('Avg'))
std_acc = table_idinvert_acc.iloc[:-1, :].std(axis=0).rename('Std')
print(std_acc / np.sqrt(40))
display(table_idinvert_acc.iloc[-1:, :])
# +
f, ax = plt.subplots(1, 1, figsize=(6, 4))
ax.plot(table['Test GAN'], table_idinvert['Test GAN'], '*', label='GAN Aug')
ax.plot(table['Test Combined'], table_idinvert['Test Combined'], '*', label='Combined Aug')
ax.set_xlabel('Pre-trained FFHQ + Encoder\nAccuracy Difference', fontsize=14)
ax.set_ylabel('ID-Invert\nAccuracy Difference', fontsize=14)
ax.legend(loc='lower right')
from scipy.stats import pearsonr
corr, pval = pearsonr(table['Test GAN'].to_list() + table['Test Combined'].to_list(),
table_idinvert['Test GAN'].to_list() + table_idinvert['Test Combined'].to_list())
print('Pearsons correlation: %.3f pval %f' % (corr, pval))
save(f, 'sm_graph_face_idinvert.pdf')
# -
# # different training approaches
# +
# different training approaches
attr_mean = data_celebahq.attr_celebahq.mean(axis=0)[:-1]
attr_order = sorted([(abs(v-0.5), v, k) for k, v in attr_mean.to_dict().items()])
table_dict = OrderedDict([])
table_accs = OrderedDict([])
for i, (_, _, attribute) in enumerate(tqdm(attr_order)):
val_values = []
val_diffs = []
test_values = []
test_diffs = []
val_labels = ['Val ' + train_method + ' ' + eval_method for train_method in
['Im', 'latent', 'latent_stylemix', 'latent_stylemix_crop'] for eval_method in ['Single', 'GAN Ens', 'Combined Ens']]
test_labels = ['Test ' + train_method + ' ' + eval_method for train_method in
['Im', 'latent', 'latent_stylemix', 'latent_stylemix_crop'] for eval_method in ['Single', 'GAN Ens', 'Combined Ens']]
for suffix in ['', '__latent', '__latent_stylemix_fine', '__latent_stylemix_fine_crop']:
attr = attribute + suffix
# print('========== %s ==========' % attr)
# gan jitter
val_file = f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_fine.npz'
test_file = f'results/precomputed_evaluations/celebahq/output/{attr}_test/gan_ensemble_stylemix_fine.npz'
expt_name = 'stylemix_fine'
# resample
resampled_accs = resample_wrapper(val_file, test_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
val_orig = resampled_accs['val_acc_original']
val_top1 = resampled_accs['val_avg']
test_orig = resampled_accs['test_acc_original']
test_top1_from_val = resampled_accs['test_avg']
# gan jitter with color/crop jitter
val_file = f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_fine_tensortransform.npz'
test_file = f'results/precomputed_evaluations/celebahq/output/{attr}_test/gan_ensemble_stylemix_fine_tensortransform.npz'
expt_name = 'stylemix_fine'
# resample
resampled_accs = resample_wrapper(val_file, test_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
val_orig_mix = resampled_accs['val_acc_original']
val_top1_mix = resampled_accs['val_avg']
test_orig_mix = resampled_accs['test_acc_original']
test_top1_from_val_mix = resampled_accs['test_avg']
# sanity check
assert(test_orig_mix == test_orig)
assert(val_orig_mix == val_orig)
new_val_values = [val_orig, val_top1, val_top1_mix]
new_test_values = [test_orig, test_top1_from_val, test_top1_from_val_mix]
val_values.extend(new_val_values)
test_values.extend(new_test_values)
val_diffs.extend([x - val_values[0] for x in new_val_values])
test_diffs.extend([x - test_values[0] for x in new_test_values])
table_dict[attribute] = val_diffs + test_diffs
table_accs[attribute] = val_values + test_values
# +
table = pd.DataFrame.from_dict(table_dict, orient='index', columns=val_labels+test_labels)
table = table.append(table.mean(axis=0).rename('Avg'))
std = table.iloc[:-1, :].std(axis=0).rename('Std')
print(std / np.sqrt(40))
# table = table.append(table.iloc[:-1, :].std(axis=0).rename('Std'))
# display(table.iloc[-2:, 12:])
display(table.iloc[-1:, 12:])
# -
table_acc = pd.DataFrame.from_dict(table_accs, orient='index', columns=val_labels+test_labels)
table_acc = table_acc.append(table_acc.mean(axis=0).rename('Avg'))
# table_acc.iloc[:, 12:]
display(table_acc.iloc[-1:, 12:])
# show the IM and W columns
# +
assert(table_acc.iloc[:-1, 12:].shape[0] == 40)
df = {'train_method': ['Im', 'Im', 'Im', 'latent', 'latent', 'latent'] + ['latent_stylemix'] * 3 + ['latent_stylemix_crop'] * 3,
'ens_method': ['Single Image', 'Style-mix Ensemble', 'Combined Ensemble'] * 4,
'acc': table_acc.iloc[:-1, 12:].mean(axis=0),
'stderr': table_acc.iloc[:-1, 12:].std(axis=0) / np.sqrt(table_acc.iloc[:-1, 12:].shape[0])
}
df = pd.DataFrame.from_dict(df)
display(df)
f, ax = plt.subplots(1, 1, figsize=(6, 4))
group_size = 3
bar_width=0.2
n_groups = 4
bar_offsets = bar_offset(group_size, n_groups, bar_width)
palette = make_blue_palette(group_size)
xticklabels = []
for i in range(group_size):
indices = np.arange(i, n_groups*group_size, group_size)
bar_height = df.iloc[indices]['acc']
bar_err = df.iloc[indices]['stderr']
assert(all([x == df.iloc[indices[0]]['ens_method'] for x in df.iloc[indices]['ens_method']]))
ax.bar(bar_offsets[i], bar_height, width=bar_width, color=palette[i],
label=df.iloc[indices[0]]['ens_method'], edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels.append(df.iloc[indices[0]]['train_method'].replace('_', '\n'))
ax.set_ylim([88.5, 89.8])
ax.legend(prop={'size': 12}) # , loc='upper left')
# ax.legend(loc='upper center', bbox_to_anchor=(0.5, -0.1), ncol=2, prop={'size': 11})
ax.set_xticks(np.arange(1,n_groups+1))
ax.set_xticklabels(['Train\nImage', 'Train\nLatent', 'Train\nStyle-mix', 'Train\nCombined'], fontsize=14)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(12)
ax.set_xlabel('')
ax.set_ylabel('Accuracy', fontsize=16)
f.tight_layout()
save(f, 'graph_face_train_latent.pdf')
# +
assert(table.iloc[:-1, 12:].shape[0] == 40)
df = {'train_method': ['Im', 'Im', 'Im', 'latent', 'latent', 'latent'] + ['latent_stylemix'] * 3 + ['latent_stylemix_crop'] * 3,
'ens_method': ['Single Image', 'Style-mix Ensemble', 'Combined Ensemble'] * 4,
'acc': table.iloc[:-1, 12:].mean(axis=0),
'stderr': table.iloc[:-1, 12:].std(axis=0) / np.sqrt(table.iloc[:-1, 12:].shape[0])
}
df = pd.DataFrame.from_dict(df)
display(df)
f, ax = plt.subplots(1, 1, figsize=(6, 4))
group_size = 3
bar_width=0.2
n_groups = 4
bar_offsets = bar_offset(group_size, n_groups, bar_width)
palette = make_blue_palette(group_size)
xticklabels = []
for i in range(group_size):
indices = np.arange(i, n_groups*group_size, group_size)
bar_height = df.iloc[indices]['acc']
bar_err = df.iloc[indices]['stderr']
assert(all([x == df.iloc[indices[0]]['ens_method'] for x in df.iloc[indices]['ens_method']]))
ax.bar(bar_offsets[i], bar_height, width=bar_width, color=palette[i], yerr=bar_err,
label=df.iloc[indices[0]]['ens_method'], edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels.append(df.iloc[indices[0]]['train_method'].replace('_', '\n'))
# ax.set_ylim([88.5, 89.6])
ax.set_ylim([-0.3, 0.8])
ax.legend(prop={'size': 12}, loc='upper left')
# ax.legend(loc='upper center', bbox_to_anchor=(0.5, -0.1), ncol=2, prop={'size': 11})
ax.set_xticks(np.arange(1,n_groups+1))
ax.set_xticklabels(['Train\nImage', 'Train\nLatent', 'Train\nStyle-mix', 'Train\nCombined'], fontsize=14)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(12)
ax.set_xlabel('')
ax.set_ylabel('Accuracy Difference', fontsize=16)
f.tight_layout()
save(f, 'graph_face_train_latent_diff.pdf')
# -
# # distribution of classification accuracies
f, ax = plt.subplots(1, 1, figsize=(6, 4))
ax.hist(table_acc['Test Im Single'])
ax.set_xlim([50, 100])
ax.set_ylabel('Count', fontsize=14)
ax.set_xlabel('Test Accuracy', fontsize=14)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(12)
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(12)
save(f, 'sm_graph_face_acc_distribution.pdf')
# # over 12 attributes, plot stylemix, isotropic, and PCA fine and coarse
# +
attr_mean = data_celebahq.attr_celebahq.mean(axis=0)[:-1]
attr_order = sorted([(abs(v-0.5), v, k) for k, v in attr_mean.to_dict().items()])
df_val = defaultdict(list)
df_test = defaultdict(list)
for i, (_, _, attr) in enumerate(tqdm(attr_order[:12])):
# print('========== %s ==========' % attr)
val_expts = [
(f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_isotropic_coarse.npz',
('isotropic_coarse_0.10', 'isotropic_coarse_0.30'), 'Isotropic Coarse'),
(f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_isotropic_fine.npz',
('isotropic_fine_0.10', 'isotropic_fine_0.30'), 'Isotropic Fine'),
(f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_pca_coarse.npz',
('pca_coarse_1.00', 'pca_coarse_2.00', 'pca_coarse_3.00'), 'PCA Coarse'),
(f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_pca_fine.npz',
('pca_fine_1.00', 'pca_fine_2.00', 'pca_fine_3.00'), 'PCA Fine'),
(f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_coarse.npz',
('stylemix_coarse',), 'Style-mix Coarse'),
(f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_fine.npz',
('stylemix_fine',), 'Style-mix Fine'),
]
test_expts = [(x.replace('_val/', '_test/'), y, z) for x, y, z in val_expts]
for i, (val, test) in enumerate(zip(val_expts, test_expts)):
expt_settings = []
for expt_name in val[1]:
resampled_accs = resample_wrapper(val[0], test[0], expt_name, ens_size=31,
add_aug=False, verbose=False)
resampled_accs['expt_name'] = expt_name
expt_settings.append(resampled_accs)
# these should all be the same -- just standard test info
assert(all([x['val_acc_original'] == expt_settings[0]['val_acc_original'] for x in expt_settings]))
assert(all([x['test_acc_original'] == expt_settings[0]['test_acc_original'] for x in expt_settings]))
if i == 0:
df_val['attribute'].append(attr)
df_val['acc'].append(expt_settings[0]['val_acc_original'])
df_val['stderr'].append(0.)
df_val['expt_group'].append('Original Image')
df_val['expt'].append('original')
df_test['attribute'].append(attr)
df_test['acc'].append(expt_settings[0]['test_acc_original'])
df_test['stderr'].append(0.)
df_test['expt_group'].append('Original Image')
df_test['expt'].append('original')
# import pdb; pdb.set_trace()
best_expt = max(expt_settings, key=lambda x: x['val_avg']) # take the val accuracy
# val result
df_val['attribute'].append(attr)
df_val['acc'].append(best_expt['val_avg'])
df_val['stderr'].append(best_expt['val_stderr'])
df_val['expt'].append(best_expt['expt_name'])
df_val['expt_group'].append(val[2])
# test result
df_test['attribute'].append(attr)
df_test['acc'].append(best_expt['test_avg'])
df_test['stderr'].append(best_expt['test_stderr'])
df_test['expt'].append(best_expt['expt_name'])
df_test['expt_group'].append(test[2])
df_val = pd.DataFrame.from_dict(df_val)
df_test = pd.DataFrame.from_dict(df_test)
# +
df_per_attr_val = OrderedDict([])
group_size=7
num_attr=12
for i in range(0, num_attr*group_size, group_size):
attribute_names = list(df_val.iloc[i:i+group_size]['attribute'])
assert(all([x == attribute_names[0] for x in attribute_names]))
df_per_attr_val[attribute_names[0]] = list(df_val.iloc[i:i+group_size]['acc'])
df_per_attr_val = pd.DataFrame.from_dict(df_per_attr_val, orient='index', columns=['Original'] + [x for _,_, x in val_expts])
df_per_attr_test = OrderedDict([])
group_size=7
num_attr=12
for i in range(0, num_attr*group_size, group_size):
attribute_names = list(df_test.iloc[i:i+group_size]['attribute'])
assert(all([x == attribute_names[0] for x in attribute_names]))
df_per_attr_test[attribute_names[0]] = list(df_test.iloc[i:i+group_size]['acc'])
df_per_attr_test = pd.DataFrame.from_dict(df_per_attr_test, orient='index', columns=['Original'] + [x for _,_, x in test_expts])
# -
df_per_attr_test
# +
df_per_attr_val_diff = (df_per_attr_val.sub(df_per_attr_val['Original'], axis=0)).iloc[:, 1:]
df_per_attr_test_diff = (df_per_attr_test.sub(df_per_attr_test['Original'], axis=0)).iloc[:, 1:]
f, ax = plt.subplots(1, 1, figsize=(6, 3))
group_size = 2
bar_width=0.25
n_groups = 6
bar_offsets = bar_offset(group_size, n_groups, bar_width)
palette = sns.color_palette()
#### combined plot ####
for i, label in enumerate(df_per_attr_val_diff.columns):
# val
height = df_per_attr_val_diff[label].mean()
yerr = df_per_attr_val_diff[label].std() / np.sqrt(df_per_attr_val_diff.shape[0])
ax.bar(bar_offsets[0][i], height, yerr=yerr, width=bar_width, color=palette[0],
edgecolor=(0.5, 0.5, 0.5), capsize=5, label='Validation' if i == 0 else None)
# test
height = df_per_attr_test_diff[label].mean()
yerr = df_per_attr_test_diff[label].std() / np.sqrt(df_per_attr_test_diff.shape[0])
ax.bar(bar_offsets[1][i], height, yerr=yerr, width=bar_width, color=palette[1],
edgecolor=(0.5, 0.5, 0.5), capsize=5, label='Test' if i == 0 else None)
ax.legend()
ax.set_ylabel('Accuracy Difference', fontsize=14)
ax.set_xticks(np.arange(1,n_groups+1))
ax.set_xticklabels([x.replace(' ', '\n') for x in df_per_attr_val_diff.columns], fontsize=11)
save(f, 'graph_face_gan_aug_types.pdf')
# -
# # plot the accuracy vs alpha graph
for attr in ['Smiling',]:
val_expt = (f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_fine_tensortransform.npz',
('stylemix_fine',), 'Style-Mix Fine')
x, y, z = val_expt
test_expt = (x.replace('_val', '_test'), y, z)
val_res = get_accuracy_from_npz(val_expt[0], val_expt[1][0], add_aug=False, ens_size=31)
test_res = get_accuracy_from_npz(test_expt[0], test_expt[1][0], add_aug=False, ens_size=31)
f, ax = plt.subplots(1, 1, figsize=(6, 3)) # , sharey=True)
ax.plot(val_res['ensemble_table']['weight'], val_res['ensemble_table']['acc'], label='Validation')
ax.plot(test_res['ensemble_table']['weight'], test_res['ensemble_table']['acc'], label='Test')
# plot the ensemble weight
val_ensemble_table = val_res['ensemble_table']
best_val_setting = val_ensemble_table.iloc[val_ensemble_table['acc'].argsort().iloc[-1], :]
ax.axvline(best_val_setting.weight, color='k', linestyle=':', label='Selected Weight')
ax.set_ylabel('Accuracy')
ax.set_xlabel('Ensemble Weight')
#for tick in ax.yaxis.get_major_ticks():
# tick.label.set_fontsize(12)
#for tick in ax.xaxis.get_major_ticks():
# tick.label.set_fontsize(12)
if attr == 'Smiling':
ax.legend()
# ax.set_title('Attribute: ' + attr.replace('_', ' '), fontsize=16)
# ax[1].set_title('Test', fontsize=16)
# f.suptitle('Attribute: ' + attr.replace('_', ' '), fontsize=16, y=1.0)
f.tight_layout()
save(f, 'sm_ensemble_alpha_%s_v2.pdf' % attr)
# # stylegan corruptions
# +
# sample each 20 times
table_dict = OrderedDict([])
table_accs = OrderedDict([])
table_stderrs = OrderedDict([])
# axes = [col for row in axes for col in row]
n_samples = 20
for i, attribute in enumerate(['Smiling', 'Arched_Eyebrows', 'Young', 'Wavy_Hair']):
val_values = []
test_values = []
val_stderrs = []
test_stderrs = []
val_diffs = []
test_diffs = []
val_labels = ['Val ' + corruption + ' ' + eval_method for corruption in
['Im', 'Jpeg', 'Blur', 'Noise', 'FGSM', 'PGD', 'CW'] for eval_method in ['S', 'R', 'G', 'C']]
test_labels = ['Test ' + corruption + ' ' + eval_method for corruption in
['Im', 'Jpeg', 'Blur', 'Noise', 'FGSM', 'PGD', 'CW'] for eval_method in ['S', 'R', 'G', 'C']]
for prefix in ['', 'corruption_jpeg_', 'corruption_gaussian_blur_', 'corruption_gaussian_noise_', 'fgsm_', 'pgd_', 'cw_']:
attr = prefix + attribute
print(attr)
# gan jitter fine
val_file = f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_fine.npz'
test_file = f'results/precomputed_evaluations/celebahq/output/{attr}_test/gan_ensemble_stylemix_fine.npz'
expt_name = 'stylemix_fine'
# resample
resampled_accs = resample_wrapper(val_file, test_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
val_orig = resampled_accs['val_acc_original']
val_top1 = resampled_accs['val_avg']
val_stderr = resampled_accs['val_stderr']
val_rec = resampled_accs['val_acc_rec']
test_orig = resampled_accs['test_acc_original']
test_top1_from_val = resampled_accs['test_avg']
test_stderr = resampled_accs['test_stderr']
test_rec = resampled_accs['test_acc_rec']
# gan jitter with color/crop jitter
val_file = f'results/precomputed_evaluations/celebahq/output/{attr}_val/gan_ensemble_stylemix_fine_tensortransform.npz'
test_file = f'results/precomputed_evaluations/celebahq/output/{attr}_test/gan_ensemble_stylemix_fine_tensortransform.npz'
expt_name = 'stylemix_fine'
resampled_accs = resample_wrapper(val_file, test_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
val_orig_mix = resampled_accs['val_acc_original']
val_top1_mix = resampled_accs['val_avg']
val_stderr_mix = resampled_accs['val_stderr']
val_rec_mix = resampled_accs['val_acc_rec']
test_orig_mix = resampled_accs['test_acc_original']
test_top1_from_val_mix = resampled_accs['test_avg']
test_stderr_mix = resampled_accs['test_stderr']
test_rec_mix = resampled_accs['test_acc_rec']
# sanity check
assert(test_orig_mix == test_orig)
assert(test_rec_mix == test_rec)
assert(val_orig_mix == val_orig)
assert(val_rec_mix == val_rec)
new_val_values = [val_orig, val_rec, val_top1, val_top1_mix]
new_val_stderrs = [0., 0., val_stderr, val_stderr_mix]
new_test_values = [test_orig, test_rec, test_top1_from_val, test_top1_from_val_mix]
new_test_stderrs = [0., 0., test_stderr, test_stderr_mix]
val_values.extend(new_val_values)
test_values.extend(new_test_values)
val_stderrs.extend(new_val_stderrs)
test_stderrs.extend(new_test_stderrs)
val_diffs.extend([x - val_values[0] for x in new_val_values])
test_diffs.extend([x - test_values[0] for x in new_test_values])
table_dict[attribute] = val_diffs + test_diffs
table_accs[attribute] = val_values + test_values
table_stderrs[attribute] = val_stderrs + test_stderrs
# -
table = pd.DataFrame.from_dict(table_dict, orient='index', columns=val_labels+test_labels)
table.shape
# +
display(table.iloc[:, 28:])
table_acc = pd.DataFrame.from_dict(table_accs, orient='index', columns=val_labels+test_labels)
display(table_acc.iloc[:, 28:])
table_stderr = pd.DataFrame.from_dict(table_stderrs, orient='index', columns=val_labels+test_labels)
display(table_stderr.iloc[:, 28:])
# +
f, axes = plt.subplots(1, 4, figsize=(16, 3.5))
for row, attr in enumerate(table_acc.index):
ax = axes[row]
df = {'train_method': ['Uncorrupted'] * 4 + ['Jpeg'] * 4 + ['Blur'] * 4 + ['Noise'] * 4,
'ens_method': ['Image', 'Reconstruction', 'Style-mix Ensemble', 'Combined Ensemble'] * 4,
'acc': table_acc.iloc[row, 28:-12],
'stderr': table_stderr.iloc[row, 28:-12]
}
df = pd.DataFrame.from_dict(df)
# display(df)
palette = make_blue_palette(4)
group_size=4
n_groups=4
bar_width=0.2
bar_offsets = bar_offset(group_size, n_groups, bar_width)
xticklabels = []
for i in range(group_size):
indices = np.arange(i, n_groups*group_size, group_size)
bar_height = df.iloc[indices]['acc']
bar_err = df.iloc[indices]['stderr']
assert(all([x == df.iloc[indices[0]]['ens_method'] for x in df.iloc[indices]['ens_method']]))
ax.bar(bar_offsets[i], bar_height, width=bar_width, color=palette[i], # yerr=bar_err,
label=df.iloc[indices[0]]['ens_method'], edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels.append(df.iloc[indices[0]]['train_method'].replace('_', '\n'))
ax.set_ylim([np.min(df['acc'])-1.0, np.max(df['acc'])+1.0])
# ax.legend(loc='upper left', prop={'size': 12})
# ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
ax.set_xticks(np.arange(1, n_groups+1))
ax.set_xticklabels(['Clean', 'Jpeg', 'Blur', 'Noise'], fontsize=14)
ax.set_xlabel('')
ax.set_ylabel('Accuracy', fontsize=16)
ax.set_title(attr.replace('_', ' '), fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(12)
handles, labels = ax.get_legend_handles_labels() # on the last axis
lgd = f.legend(handles, labels, loc='lower center', ncol=4, prop={'size': 12},
bbox_to_anchor=(0.5, -0.08), edgecolor='1.0')
f.tight_layout()
save(f, 'graph_face_untargeted_corruption.pdf')
# +
f, axes = plt.subplots(1, 4, figsize=(16, 3.5))
for row, attr in enumerate(table_acc.index):
ax = axes[row]
df = {'train_method': ['Uncorrupted'] * 4 + ['FGSM'] * 4 + ['PGD'] * 4 + ['CW'] * 4,
'ens_method': ['Image', 'Reconstruction', 'Style-mix Ensemble', 'Combined Ensemble'] * 4,
'acc': table_acc.iloc[row, list(range(28, 32)) + list(range(44,56))],
'stderr': table_stderr.iloc[row, list(range(28, 32)) + list(range(44,56))]
}
df = pd.DataFrame.from_dict(df)
# display(df)
palette = make_blue_palette(4)
group_size=4
n_groups=4
bar_width=0.2
bar_offsets = bar_offset(group_size, n_groups, bar_width)
xticklabels = []
for i in range(group_size):
indices = np.arange(i, n_groups*group_size, group_size)
bar_height = df.iloc[indices]['acc']
bar_err = df.iloc[indices]['stderr']
assert(all([x == df.iloc[indices[0]]['ens_method'] for x in df.iloc[indices]['ens_method']]))
b = ax.bar(bar_offsets[i], bar_height, width=bar_width, color=palette[i], # yerr=bar_err,
label=df.iloc[indices[0]]['ens_method'], edgecolor=(0.5, 0.5, 0.5), capsize=5)
xticklabels.append(df.iloc[indices[0]]['train_method'].replace('_', '\n'))
# ax.set_ylim([np.min(df['acc'])-1.0, np.max(df['acc'])+1.0])
# ax.legend(loc='upper center', prop={'size': 12})
ax.set_ylim([0, 100])
#ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
ax.set_xticks(np.arange(1, n_groups+1))
ax.set_xticklabels(['Clean', 'FGSM', 'PGD', 'CW'], fontsize=14)
ax.set_xlabel('')
ax.set_ylabel('Accuracy', fontsize=16)
ax.set_title(attr.replace('_', ' '), fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(12)
# axes[0].legend([],[], frameon=False)
# axes[1].legend([],[], frameon=False)
# axes[2].legend([],[], frameon=False)
handles, labels = ax.get_legend_handles_labels() # on the last axis
lgd = f.legend(handles, labels, loc='lower center', ncol=4, prop={'size': 12}, bbox_to_anchor=(0.5, -0.08), edgecolor='1.0')
f.tight_layout()
save(f, 'graph_face_targeted_corruption.pdf')
# -
# # stylegan ensemble size
def compute_best_weight_ensemble_size(val_data_file, test_data_file, expt_name, verbose=True, add_aug=False, seed=None):
ens_sizes = [0, 2, 4, 8, 12, 16, 20, 24, 28, 30, 31]
num_samples = 16
assert('val' in val_data_file)
assert('test' in test_data_file)
# compute best val setting using full ensemble
val_accuracy_info = get_accuracy_from_npz(val_data_file, expt_name, add_aug=add_aug, ens_size=31, seed=seed)
val_ensemble_table = val_accuracy_info['ensemble_table']
# best_val_setting = val_ensemble_table.iloc[val_ensemble_table['acc'].idxmax(), :]
best_val_setting = val_ensemble_table.iloc[val_ensemble_table['acc'].argsort().iloc[-1], :]
if verbose:
print("Val original %0.4f Val reconstructed %0.4f" %
(val_accuracy_info['acc_original'], val_accuracy_info['acc_reconstructed']))
print("%0.4f @ %0.4f %s" % (best_val_setting['acc'], best_val_setting['weight'], best_val_setting['expt_name']))
# test: iterate through ensemble sizes, taking samples from each
accs_reconstructed = []
accs_original = []
test_table = OrderedDict([(ens_size, []) for ens_size in ens_sizes])
for ens_size in ens_sizes:
for sample in range(num_samples):
test_accuracy_info = get_accuracy_from_npz(test_data_file, expt_name, weight=best_val_setting['weight'],
add_aug=add_aug, ens_size=ens_size, seed=sample)
accs_reconstructed.append(test_accuracy_info['acc_reconstructed'])
accs_original.append(test_accuracy_info['acc_original'])
test_ensemble_table = test_accuracy_info['ensemble_table']
assert(test_ensemble_table.shape[0] == 1) # it should only evaluate at the specified weight
test_setting_from_val = test_ensemble_table.iloc[0, :]
test_table[ens_size].append(test_setting_from_val['acc'])
# sanity check
assert(all([x == accs_reconstructed[0] for x in accs_reconstructed]))
assert(all([x == accs_original[0] for x in accs_original]))
test_df = pd.DataFrame.from_dict(test_table, orient='index', columns=range(num_samples))
return {'val_info': val_accuracy_info, 'test_info': test_accuracy_info,
'val_setting': best_val_setting, 'test_df': test_df}
# +
expt_name = 'stylemix_fine'
expt_data = [
('Smiling', f'results/precomputed_evaluations/celebahq/output/%s_%s/gan_ensemble_stylemix_fine_tensortransform.npz'),
('Arched_Eyebrows', f'results/precomputed_evaluations/celebahq/output/%s_%s/gan_ensemble_stylemix_fine_tensortransform.npz'),
('Wavy_Hair', f'results/precomputed_evaluations/celebahq/output/%s_%s/gan_ensemble_stylemix_fine_tensortransform.npz'),
('Young', f'results/precomputed_evaluations/celebahq/output/%s_%s/gan_ensemble_stylemix_fine_tensortransform.npz')
]
f, axes = plt.subplots(1, 4, figsize=(16, 4))
# axes = [ax for row in axes for ax in row]
for i, (attr, data_file_base) in enumerate(expt_data):
ax = axes[i]
output = compute_best_weight_ensemble_size(data_file_base % (attr, 'val'),
data_file_base % (attr, 'test'),
expt_name)
plot_vals = output['test_df'].to_numpy()
m = np.mean(plot_vals, axis=1)
s = np.std(plot_vals, axis=1) / np.sqrt(plot_vals.shape[1])
ax.plot(output['test_df'].index, m)
ax.fill_between(output['test_df'].index, m-s, m+s, alpha=0.3)
ax.set_title(attr.replace('_', ' '), fontsize=16)
ax.set_xlabel('Number of\nGAN samples', fontsize=14)
ax.set_ylabel('Accuracy', fontsize=16)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(12)
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(12)
# ax.axhline(test_output[0][0])
# ax.axhline(test_output[2])
f.tight_layout()
save(f, 'graph_face_ensemble_size.pdf')
# -
# # cifar10
# +
table_dict = {}
for classifier in ['imageclassifier', 'latentclassifier', 'latentclassifier_layer6', 'latentclassifier_layer7']:
print("==================")
for expt_name in ['stylemix_layer6', 'stylemix_layer7']:
print("---> %s %s" % (classifier, expt_name))
val_data_file = f'results/precomputed_evaluations/cifar10/output/{classifier}_val/gan_ensemble_{expt_name}.npz'
test_data_file = val_data_file.replace('_val', '_test')
resampled_accs = resample_wrapper(val_data_file, test_data_file, expt_name, ens_size=31,
add_aug=False, verbose=False)
print("val improvement: %0.3f" % (resampled_accs['val_avg'] - resampled_accs['val_acc_original']))
print("test improvement: %0.3f" % (resampled_accs['test_avg'] - resampled_accs['test_acc_original']))
oracle = get_accuracy_from_npz(test_data_file, expt_name)
oracle_table = oracle['ensemble_table']
oracle_setting = oracle_table.iloc[oracle_table['acc'].argsort().iloc[-1], :]
print("oracle imrovement: %0.3f" % (oracle_setting['acc'] - resampled_accs['test_acc_original']))
if expt_name == 'stylemix_layer6':
# also extract the classifier acc on images
table_dict['%s %s' % (classifier, 'images')] = [np.nan, resampled_accs['val_acc_original'],
resampled_accs['test_acc_original'], np.nan, np.nan]
table_dict['%s %s' % (classifier, expt_name)] = [np.mean(resampled_accs['weights']), resampled_accs['val_avg'],
resampled_accs['test_avg'], oracle_setting['weight'],
oracle_setting['acc']]
# -
table = pd.DataFrame.from_dict(table_dict, orient='index',
columns=['val weight', 'val acc', 'test acc', 'oracle weight', 'oracle acc'])
table
# +
# plot it
f, ax = plt.subplots(1, 1, figsize=(6, 4))
group_size = 3
bar_width=0.2
n_groups = 4 # training configurations
bar_offsets = bar_offset(group_size, n_groups, bar_width)
palette = make_yellow_palette(2)[1:] + make_blue_palette(2)[1:] + make_green_palette(2)[1:]
ind = 0.2
# ax.axhline(im_crops['acc_ensembled'], color='k', linestyle=':', label='Original Images')
ax.bar(bar_offsets[0], table.loc[[x for x in table.index if x.endswith('images')]]['test acc'],
width=bar_width, color=palette[0], label='Image', edgecolor=(0.5, 0.5, 0.5), capsize=5)
for i, layer in enumerate([6, 7]):
ax.bar(bar_offsets[i+1], table.loc[[x for x in table.index if x.endswith('layer%d' % layer)]]['test acc'],
width=bar_width, color=palette[i+1], label='Style-mix Layer%d' % layer, edgecolor=(0.5, 0.5, 0.5), capsize=5)
ax.set_ylim([92, 96])
ax.set_ylabel('Classification Accuracy', fontsize=14)
ax.set_xticks(np.arange(1, n_groups+1))
ax.legend()
ax.set_xticklabels(['Original\nImages', 'GAN\nReconstructions',
'Style-mix\nLayer 6', 'Style-mix\nLayer 8'], fontsize=12)
ax.set_xlabel('Classifier training distribution', fontsize=16)
save(f, 'graph_cifar10.pdf')
| notebooks/plot_precomputed_evaluations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#download dataset from here (https://s3.amazonaws.com/content.udacity-data.com/nd089/Cat_Dog_data.zip)
import numpy as np
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data.sampler import SubsetRandomSampler
from torchvision import datasets, transforms
from torchsummary import summary
train_on_gpu = torch.cuda.is_available()
if not train_on_gpu:
print('CUDA is not available. Training on CPU ...')
else:
print('CUDA is available! Training on GPU ...')
# +
data_dir = 'Cat_Dog_data'
valid_size = 0.2
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
test_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
# define samplers for obtaining training and validation batches
# Pass transforms in here, then run the next cell to see how the transforms look
train_data = datasets.ImageFolder(data_dir + '/train', transform=train_transforms)
test_data = datasets.ImageFolder(data_dir + '/test', transform=test_transforms)
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
trainloader = torch.utils.data.DataLoader(train_data, batch_size=64,sampler=train_sampler)
testloader = torch.utils.data.DataLoader(test_data, batch_size=64)
validloader = torch.utils.data.DataLoader(train_data, batch_size=64,
sampler=valid_sampler)
# +
class Net(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(in_channels = 3, out_channels = 16, kernel_size=(3, 3), stride=2, padding=1)
self.conv2 = nn.Conv2d(in_channels = 16, out_channels = 32, kernel_size=(3, 3), stride=2, padding=1)
self.conv3 = nn.Conv2d(in_channels = 32, out_channels = 64, kernel_size=(3, 3),padding=1)
self.conv4 = nn.Conv2d(in_channels = 64, out_channels = 128, kernel_size=(3, 3),padding=1)
self.conv5 = nn.Conv2d(in_channels = 128, out_channels = 256, kernel_size=(3, 3), padding=1)
self.fc1 = nn.Linear(in_features= 256, out_features=50)
self.fc2 = nn.Linear(in_features=50, out_features=10)
self.fc3 = nn.Linear(in_features=10, out_features=2)
def forward(self, X):
X = F.relu(self.conv1(X))
X = F.avg_pool2d(X, 2)
X = F.relu(self.conv2(X))
X = F.avg_pool2d(X, 2)
X = F.relu(self.conv3(X))
X = F.avg_pool2d(X, 2)
X = F.relu(self.conv4(X))
X = F.avg_pool2d(X, 2)
X = F.relu(self.conv5(X))
X = F.avg_pool2d(X, 2)
# print(X.shape)
X = X.view(X.shape[0], -1)
X = F.relu(self.fc1(X))
X = F.relu(self.fc2(X))
X = F.log_softmax(self.fc3(X), dim=1)
# X = torch.sigmoid(X)
return X
# +
model=Net()
model.cuda()
device="cuda"
model
#summary(model,(3,255,255))
# -
optimizer = optim.Adam(model.parameters(), lr=0.003)
criterion = nn.NLLLoss()
# +
n_epochs = 50
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# keep track of training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train()
for data, target in trainloader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model.eval()
for data, target in validloader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss += loss.item()*data.size(0)
# calculate average losses
train_loss = train_loss/len(trainloader.sampler)
valid_loss = valid_loss/len(validloader.sampler)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model.state_dict(), 'dogs2.pt')
valid_loss_min = valid_loss
# -
model.load_state_dict(torch.load('dogs2.pt'))
# +
# track test loss
classes=["cats","dogs"]
test_loss = 0.0
class_correct = list(0. for i in range(3))
class_total = list(0. for i in range(3))
model.eval()
# iterate over test data
for data, target in testloader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct_tensor = pred.eq(target.data.view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
# calculate test accuracy for each object class
for i in range(4):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# average test loss
test_loss = test_loss/len(testloader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(2):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# helper function to un-normalize and display an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
plt.imshow(np.transpose(img, (1, 2, 0)))
dataiter = iter(testloader)
images, labels = dataiter.next()
images.numpy()
# move model inputs to cuda, if GPU available
if train_on_gpu:
images = images.cuda()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds_tensor = torch.max(output, 1)
preds = np.squeeze(preds_tensor.numpy()) if not train_on_gpu else np.squeeze(preds_tensor.cpu().numpy())
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
imshow(images.cpu()[idx])
ax.set_title("{} ({})".format(classes[preds[idx]], classes[labels[idx]]),
color=("green" if preds[idx]==labels[idx].item() else "red"))
# -
torch.cuda.empty_cache()
| dog-cats-classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # %load first_cell.py
# %reload_ext autoreload
# %autoreload 2
from pathlib import Path
home = str(Path.home())
import sys
sys.path = sys.path + [f'{home}/.conda/envs/norm_env/lib/python37.zip',
f'{home}/.conda/envs/norm_env/lib/python3.7',
f'{home}/.conda/envs/norm_env/lib/python3.7/lib-dynload',
f'{home}/.conda/envs/norm_env/lib/python3.7/site-packages',
'../src']
sys.prefix = '/home/joaom/.conda/envs/norm_env'
from paths import RAW_PATH, TREAT_PATH, OUTPUT_PATH, FIGURES_PATH
from copy import deepcopy
import numpy as np
import pandas as pd
pd.options.display.max_columns = 999
import yaml
import matplotlib.pyplot as plt
import datetime
import warnings
warnings.filterwarnings('ignore')
# Plotting
import plotly
import plotly.graph_objs as go
import cufflinks as cf
plotly.offline.init_notebook_mode(connected=True)
def iplottitle(title, width=40):
return '<br>'.join(textwrap.wrap(title, width))
# Setting cufflinks
import textwrap
import cufflinks as cf
cf.go_offline()
cf.set_config_file(offline=False, world_readable=True)
import yaml
from jinja2 import Template
cf.themes.THEMES['custom'] = yaml.load(open('cufflinks_template.yaml', 'r'))
# -
import pickle
import networkx as nx
import pylab as plt
from networkx.drawing.nx_agraph import graphviz_layout, to_agraph
def replace_attribute_key(G, tnode, old_key, new_key):
value = G.node[tnode][old_key]
G.node[tnode].pop(old_key, None)
G.node[tnode][new_key] = value
G = pickle.load(open(OUTPUT_PATH / 'dependency_graph.p', 'rb'))
attr = G.node.data()._nodes
for k in attr.keys():
replace_attribute_key(G, k, 'name', '_name')
nx.nx_pydot.write_dot(G, open(OUTPUT_PATH / 'dependency.dot', 'w'))
from rdp import rdp
# +
import geojson
from shapely.geometry import shape
import pandas as pd
from pathlib import Path
def convert(path):
converted = []
for p in Path(path).glob('*.GeoJson'):
d = geojson.load(open(p, 'r'))
converted.append(dict(
country_name=d['properties']['name'],
country_iso=d['properties']['alltags']['ISO3166-1'],
region_slug='_'.join(['country'] + d['properties']['name'].lower().split(' ')),
region_name=d['properties']['name'],
region_type='country',
dashboard='TRUE',
population=d['properties']['alltags'].get('population'),
timezone=d['properties']['alltags'].get('timezone'),
region_shapefile_wkt_1=None,
region_shapefile_wkt=shape(d['geometry']).simplify(0.05, preserve_topology=False).wkt
))
pd.DataFrame(converted)[['country_name',
'country_iso',
'region_slug',
'region_name',
'region_type',
'dashboard',
'population',
'timezone',
'region_shapefile_wkt_1',
'region_shapefile_wkt']].to_csv(path / 'converted.csv', index=False)
# -
a = convert(RAW_PATH / 'countries-boundaries')
converted
import pandas as pd
url = 'https://docs.google.com/spreadsheets/d/1xBDV5Zm7zSRPK2TGBoVSHhkvPeqRT9hZFiD4AueoVeI/export?format=csv&id'
df = pd.read_csv(url)
ar = df.query('country_name == "Argentina"').query('region_type == "city"')
ar['day'].unique()
ar.pivot_table(columns='region_slug', index='dow', values='expected_2020').iplot()
i = 8
ar.query('region_slug == "cordoba"')[['region_shapefile_wkt']].iloc[0].values
ar.query('region_slug == "cordoba"')[['observed', 'expected_2020', 'day', 'ratio_20']].sort_values(by='observed')
ar.query(f'day in ({list(range(i, i+7))})')\
.pivot_table(columns='region_slug', index='dow', values='observed').iplot()
i = 8+7
ar.query(f'day in ({list(range(i, i+7))})')\
.pivot_table(columns='region_slug', index='dow', values='observed').iplot()
ar.pivot_table(columns='region_slug', index='dow', values='observed').iplot(kind='bar')
from src import utils
from datetime import datetime
conn = utils.connect_athena(path='../configs/athena.yaml')
df = pd.read_sql_query("""
select
*
from spd_sdv_waze_corona.prod_daily_daily_raw
where st_intersects(
st_polygon('Polygon ((-64.30875060953628974 -31.27541800853970955, -64.292083943961984 -31.27541800853970955, -64.292083943961984 -31.28375134244235056, -64.27541727838767827 -31.28375134244235056, -64.27541727838767827 -31.29208467634500579, -64.26708394560051829 -31.29208467634500579, -64.26708394560051829 -31.30041801024764681, -64.25875061281337253 -31.30041801024764681, -64.25875061281337253 -31.30875134415030203, -64.24208394723905258 -31.30875134415030203, -64.24208394723905258 -31.31708467805294305, -64.23375061445190681 -31.31708467805294305, -64.23375061445190681 -31.30875134415030203, -64.22541728166476105 -31.30875134415030203, -64.22541728166476105 -31.30041801024764681, -64.21708394887760107 -31.30041801024764681, -64.21708394887760107 -31.30875134415030203, -64.2087506160904411 -31.30875134415030203, -64.2087506160904411 -31.31708467805294305, -64.20041728330329533 -31.31708467805294305, -64.20041728330329533 -31.32541801195559827, -64.19208395051613536 -31.32541801195559827, -64.19208395051613536 -31.30875134415030203, -64.18375061772897539 -31.30875134415030203, -64.18375061772897539 -31.30041801024764681, -64.16708395215468386 -31.30041801024764681, -64.16708395215468386 -31.30875134415030203, -64.15875061936752388 -31.30875134415030203, -64.15875061936752388 -31.34208467976088031, -64.14208395379321814 -31.34208467976088031, -64.14208395379321814 -31.35041801366353553, -64.12541728821889819 -31.35041801366353553, -64.12541728821889819 -31.35875134756617655, -64.11708395543175243 -31.35875134756617655, -64.11708395543175243 -31.36708468146883177, -64.10875062264460666 -31.36708468146883177, -64.10875062264460666 -31.37541801537147279, -64.10041728985744669 -31.37541801537147279, -64.10041728985744669 -31.38375134927412802, -64.09208395707028671 -31.38375134927412802, -64.09208395707028671 -31.39208468317676903, -64.10041728985744669 -31.39208468317676903, -64.10041728985744669 -31.40041801707942426, -64.10875062264460666 -31.40041801707942426, -64.10875062264460666 -31.41708468488470629, -64.10041728985744669 -31.41708468488470629, -64.10041728985744669 -31.44208468659265776, -64.09208395707028671 -31.44208468659265776, -64.09208395707028671 -31.45041802049529878, -64.08375062428314095 -31.45041802049529878, -64.08375062428314095 -31.458751354397954, -64.07541729149599519 -31.458751354397954, -64.07541729149599519 -31.46708468830059502, -64.08375062428314095 -31.46708468830059502, -64.08375062428314095 -31.47541802220323603, -64.12541728821889819 -31.47541802220323603, -64.12541728821889819 -31.48375135610589126, -64.13375062100605817 -31.48375135610589126, -64.13375062100605817 -31.49208469000853228, -64.15875061936752388 -31.49208469000853228, -64.15875061936752388 -31.48375135610589126, -64.19208395051613536 -31.48375135610589126, -64.19208395051613536 -31.49208469000853228, -64.20041728330329533 -31.49208469000853228, -64.20041728330329533 -31.5004180239111875, -64.21708394887760107 -31.5004180239111875, -64.21708394887760107 -31.49208469000853228, -64.22541728166476105 -31.49208469000853228, -64.22541728166476105 -31.5004180239111875, -64.25041728002621255 -31.5004180239111875, -64.25041728002621255 -31.49208469000853228, -64.25875061281337253 -31.49208469000853228, -64.25875061281337253 -31.48375135610589126, -64.26708394560051829 -31.48375135610589126, -64.26708394560051829 -31.458751354397954, -64.27541727838767827 -31.458751354397954, -64.27541727838767827 -31.45041802049529878, -64.28375061117483824 -31.45041802049529878, -64.28375061117483824 -31.42541801878736152, -64.27541727838767827 -31.42541801878736152, -64.27541727838767827 -31.41708468488470629, -64.28375061117483824 -31.41708468488470629, -64.28375061117483824 -31.39208468317676903, -64.292083943961984 -31.39208468317676903, -64.292083943961984 -31.37541801537147279, -64.30041727674912977 -31.37541801537147279, -64.30041727674912977 -31.36708468146883177, -64.292083943961984 -31.36708468146883177, -64.292083943961984 -31.35875134756617655, -64.30041727674912977 -31.35875134756617655, -64.30041727674912977 -31.35041801366353553, -64.30875060953628974 -31.35041801366353553, -64.30875060953628974 -31.32541801195559827, -64.31708394232344972 -31.32541801195559827, -64.31708394232344972 -31.31708467805294305, -64.32541727511059548 -31.31708467805294305, -64.32541727511059548 -31.30875134415030203, -64.31708394232344972 -31.30875134415030203, -64.31708394232344972 -31.28375134244235056, -64.30875060953628974 -31.28375134244235056, -64.30875060953628974 -31.27541800853970955),(-64.23375061445190681 -31.31708467805294305, -64.23375061445190681 -31.32541801195559827, -64.22541728166476105 -31.32541801195559827, -64.22541728166476105 -31.31708467805294305, -64.23375061445190681 -31.31708467805294305))'),
st_line(line))""", conn)
day = df[ df['retrievaltime'].apply(lambda x: (x.day == 11) or (x.day == 12))]
df['day'] = df['retrievaltime'].apply(lambda x: x.day)
df.groupby('day').sum()['length'].iplot()
df.groupby('day').count()['length'].iplot(
title='Number of Records for Cordoba per day in March 2020',
theme='custom'
)
counta = pd.read_sql_query(
"""
with t as (
select
year(retrievaltime) as year,
month(retrievaltime) as month,
day(retrievaltime) as day,
day_of_week(retrievaltime) as dow,
date_parse(format_datetime(date_add('minute',
cast(date_diff('minute',
timestamp '2015-01-01 00:00:00', retrievaltime) / 5 as bigint) * 5,
timestamp '2015-01-01 00:00:00'), 'H:m'), '%H:%i') as time,
row_number() over (partition by uuid,
date_parse(format_datetime(date_add('minute',
cast(date_diff('minute',
timestamp '2015-01-01 00:00:00', retrievaltime) / 5 as bigint) * 5,
timestamp '2015-01-01 00:00:00'), 'H:m'), '%H:%i') order by retrievaltime) n_row
from (
select
uuid, arbitrary(from_unixtime(retrievaltime/1000)) retrievaltime
from "p-waze-parquet-waze"."jams"
where regexp_like(datetime, '202002')
and
speed >= 0
and length > 0
group by
uuid, pubmillis, country,
city, street, roadtype, level, length, speed,
speedkmh, delay, line, type, turntype,
blockingalertuuid, startnode, endnode))
select
month,
day,
count(*)
from t
where n_row = 1
group by month, day
""", conn)
counta.set_index(['month', 'day']).sort_index().iplot(
title='Deduplicated every 5 minutes Number of Records March 2020',
theme='custom'
)
counta = pd.read_sql_query(
"""
select
month(from_unixtime(retrievaltime/1000)) month,
day(from_unixtime(retrievaltime/1000)) day,
count(*) as counta
from "p-waze-parquet-waze"."jams"
where regexp_like(datetime, '202002|202003')
and
speed >= 0
and length > 0
group by
month(from_unixtime(retrievaltime/1000)),
day(from_unixtime(retrievaltime/1000))
""", conn)
counta.set_index(['month', 'day']).sort_index().iplot(
title='Raw Data Number of Records in Feb and March 2020',
theme='custom'
)
dupls = pd.read_sql_query(
"""
select
month,
day,
count(*) as counta
from (
select *
from "p-waze-parquet-waze"."jams"
where regexp_like(datetime, '202002|202003')
and speed >= 0
and length > 0
group by retrievaltime, uuid, pubmillis, country,
city, street, roadtype, level, length, speed,
speedkmh, delay, line, type, turntype,
blockingalertuuid, startnode, endnode, day, month, year, hour, datetime)
group by
month,
day
""", conn)
dupls.set_index(['month', 'day']).sort_index().iplot(
title='Full deduplication Raw Data Number of Records in Feb and March 2020',
theme='custom'
)
a = dupls.merge(counta, on=['month', 'day'])
a['diff'] = a['counta_x'] - a['counta_y']
a.set_index(['month', 'day']).sort_index()['diff'].iplot(
title='Difference of Raw Data and Deduplicated Number of Records',
theme='custom'
)
json = pd.read_sql_query(
"""
select
month,
day,
count(*) as counta
from "p-waze-json-waze"."jams"
where (month = '3') and year = '2020'
and day in ('1', '2', '3', '4', '5')
group by
month,
day
""", conn)
json.set_index(['month', 'day']).sort_index().iplot(
title='JSON number of files',
theme='custom',
xTitle='(month, day)'
)
cordoba_count = pd.read_sql_query(
"""
select
month(from_unixtime(retrievaltime/1000)) month,
day(from_unixtime(retrievaltime/1000)) day,
count(*) as counta
from "p-waze-parquet-waze"."jams"
where regexp_like(datetime, '202002|202003')
and
speed >= 0
and length > 0
and st_intersects(
st_polygon('Polygon ((-64.30875060953628974 -31.27541800853970955, -64.292083943961984 -31.27541800853970955, -64.292083943961984 -31.28375134244235056, -64.27541727838767827 -31.28375134244235056, -64.27541727838767827 -31.29208467634500579, -64.26708394560051829 -31.29208467634500579, -64.26708394560051829 -31.30041801024764681, -64.25875061281337253 -31.30041801024764681, -64.25875061281337253 -31.30875134415030203, -64.24208394723905258 -31.30875134415030203, -64.24208394723905258 -31.31708467805294305, -64.23375061445190681 -31.31708467805294305, -64.23375061445190681 -31.30875134415030203, -64.22541728166476105 -31.30875134415030203, -64.22541728166476105 -31.30041801024764681, -64.21708394887760107 -31.30041801024764681, -64.21708394887760107 -31.30875134415030203, -64.2087506160904411 -31.30875134415030203, -64.2087506160904411 -31.31708467805294305, -64.20041728330329533 -31.31708467805294305, -64.20041728330329533 -31.32541801195559827, -64.19208395051613536 -31.32541801195559827, -64.19208395051613536 -31.30875134415030203, -64.18375061772897539 -31.30875134415030203, -64.18375061772897539 -31.30041801024764681, -64.16708395215468386 -31.30041801024764681, -64.16708395215468386 -31.30875134415030203, -64.15875061936752388 -31.30875134415030203, -64.15875061936752388 -31.34208467976088031, -64.14208395379321814 -31.34208467976088031, -64.14208395379321814 -31.35041801366353553, -64.12541728821889819 -31.35041801366353553, -64.12541728821889819 -31.35875134756617655, -64.11708395543175243 -31.35875134756617655, -64.11708395543175243 -31.36708468146883177, -64.10875062264460666 -31.36708468146883177, -64.10875062264460666 -31.37541801537147279, -64.10041728985744669 -31.37541801537147279, -64.10041728985744669 -31.38375134927412802, -64.09208395707028671 -31.38375134927412802, -64.09208395707028671 -31.39208468317676903, -64.10041728985744669 -31.39208468317676903, -64.10041728985744669 -31.40041801707942426, -64.10875062264460666 -31.40041801707942426, -64.10875062264460666 -31.41708468488470629, -64.10041728985744669 -31.41708468488470629, -64.10041728985744669 -31.44208468659265776, -64.09208395707028671 -31.44208468659265776, -64.09208395707028671 -31.45041802049529878, -64.08375062428314095 -31.45041802049529878, -64.08375062428314095 -31.458751354397954, -64.07541729149599519 -31.458751354397954, -64.07541729149599519 -31.46708468830059502, -64.08375062428314095 -31.46708468830059502, -64.08375062428314095 -31.47541802220323603, -64.12541728821889819 -31.47541802220323603, -64.12541728821889819 -31.48375135610589126, -64.13375062100605817 -31.48375135610589126, -64.13375062100605817 -31.49208469000853228, -64.15875061936752388 -31.49208469000853228, -64.15875061936752388 -31.48375135610589126, -64.19208395051613536 -31.48375135610589126, -64.19208395051613536 -31.49208469000853228, -64.20041728330329533 -31.49208469000853228, -64.20041728330329533 -31.5004180239111875, -64.21708394887760107 -31.5004180239111875, -64.21708394887760107 -31.49208469000853228, -64.22541728166476105 -31.49208469000853228, -64.22541728166476105 -31.5004180239111875, -64.25041728002621255 -31.5004180239111875, -64.25041728002621255 -31.49208469000853228, -64.25875061281337253 -31.49208469000853228, -64.25875061281337253 -31.48375135610589126, -64.26708394560051829 -31.48375135610589126, -64.26708394560051829 -31.458751354397954, -64.27541727838767827 -31.458751354397954, -64.27541727838767827 -31.45041802049529878, -64.28375061117483824 -31.45041802049529878, -64.28375061117483824 -31.42541801878736152, -64.27541727838767827 -31.42541801878736152, -64.27541727838767827 -31.41708468488470629, -64.28375061117483824 -31.41708468488470629, -64.28375061117483824 -31.39208468317676903, -64.292083943961984 -31.39208468317676903, -64.292083943961984 -31.37541801537147279, -64.30041727674912977 -31.37541801537147279, -64.30041727674912977 -31.36708468146883177, -64.292083943961984 -31.36708468146883177, -64.292083943961984 -31.35875134756617655, -64.30041727674912977 -31.35875134756617655, -64.30041727674912977 -31.35041801366353553, -64.30875060953628974 -31.35041801366353553, -64.30875060953628974 -31.32541801195559827, -64.31708394232344972 -31.32541801195559827, -64.31708394232344972 -31.31708467805294305, -64.32541727511059548 -31.31708467805294305, -64.32541727511059548 -31.30875134415030203, -64.31708394232344972 -31.30875134415030203, -64.31708394232344972 -31.28375134244235056, -64.30875060953628974 -31.28375134244235056, -64.30875060953628974 -31.27541800853970955),(-64.23375061445190681 -31.31708467805294305, -64.23375061445190681 -31.32541801195559827, -64.22541728166476105 -31.32541801195559827, -64.22541728166476105 -31.31708467805294305, -64.23375061445190681 -31.31708467805294305))'),
st_line(line))
group by
month(from_unixtime(retrievaltime/1000)),
day(from_unixtime(retrievaltime/1000))
""", conn)
duplsdupls = pd.read_sql_query(
"""
select
month,
day,
count(*) as counta
from (
select uuid, pubmillis, country,
city, street, roadtype, level, length, speed,
speedkmh, delay, line, type, turntype,
blockingalertuuid, startnode, endnode, day, month, year, hour, datetime
from "p-waze-parquet-waze"."jams"
where regexp_like(datetime, '202002|202003')
and speed >= 0
and length > 0
group by uuid, pubmillis, country,
city, street, roadtype, level, length, speed,
speedkmh, delay, line, type, turntype,
blockingalertuuid, startnode, endnode, day, month, year, hour, datetime)
group by
month,
day
""", conn)
duplsdupls.set_index(['month', 'day']).sort_index().iplot(
title='JSON number of files',
theme='custom',
xTitle='(month, day)'
)
json = pd.read_sql_query(
"""
select
month(from_unixtime(retrievaltime/1000)) month,
day(from_unixtime(retrievaltime/1000)) day,
count(*) as counta
from spd_sdv_waze_reprocessed.jams_ready
where regexp_like(datetime, '202002|202003')
group by
month(from_unixtime(retrievaltime/1000)),
day(from_unixtime(retrievaltime/1000))
""", conn)
json.set_index(['month', 'day']).sort_index().iplot(
title='JSON number of files',
theme='custom',
xTitle='(month, day)'
)
day.to_csv(OUTPUT_PATH / 'cordoba_test.csv')
| notebooks/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Projectile Motion
#
# Recall that the equation for projectile motion when acceleration is constant can be written as $$y=y_{0}+v_{0}t+\frac{a}{2}t^{2}.$$
#
# 1. Create a Matlab or Python function that generates a plot of $y(t)$. You should be able to easily change the initial velocity, the initial position, and the acceleration. Be sure to include axis labels as appropriate. The function should take $y_{0}$, $v_{0}$, $a$ and a vector of $t$ as arguments. The function should:
#
# (a) Plot $y(t)$ with labeled axes and a grid on the plot.
#
# (b) Return the vector $y(t)$.
#
# 2. Determine $t_{\mathrm{end}}$ such that $y(t_{\mathrm{end}})=0$. In the event that there are two times that produce $y=0$ you should use the larger of the two. HINT: Solve the quadratic equation for the projectile.
#
# You do not need to submit a report for this problem - only your Matlab or Python files.
# # Solution
# ## Question 1
# Create a Matlab or Python function that generates a plot of $y(t)$. You should be able to easily change the initial velocity, the initial position, and the acceleration. Be sure to include axis labels as appropriate. The function should take $y_{0}$, $v_{0}$, $a$ and a vector of $t$ as arguments. The function should:
#
# (a) Plot $y(t)$ with labeled axes and a grid on the plot.
#
# (b) Return the vector $y(t)$.
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
def projectile(y0,v0,a,t):
"""The method's docstring
params:
y0: value
v0: value 2
"""
result = y0 + v0 * t + a/2.0 * t * t
plt.title('Projectile')
plt.xlabel('time (s)', fontsize=18)
plt.ylabel('Height (m)', fontsize=16)
plt.plot(t,result)
return result
y0=0.1
v0=10
a=-9.81
t = np.linspace(0,2,200)
y = projectile(y0,v0,a,t)
# ## Question 2
# Determine $t_{\mathrm{end}}$ such that $y(t_{\mathrm{end}})=0$. In the event that there are two times that produce $y=0$ you should use the larger of the two. HINT: Solve the quadratic equation for the projectile.
#
# What we can do here is use numpy's roots function
tend = np.max(np.roots([a/2,v0,y0]))
print('The Projectile will reach the ground in:', tend)
t = np.linspace(0,tend,200)
y = projectile(y0,v0,a,t)
| misc/Ex01. Projectile Motion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/carsonashby/DS-Unit-2-Linear-Models/blob/master/Copy_of_LS_DS_224_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="sz3wU5NUuMA3"
# Lambda School Data Science
#
# *Unit 2, Sprint 2, Module 4*
#
# ---
# + [markdown] id="nCc3XZEyG3XV"
# # Classification Metrics
#
# ## Assignment
# - [ ] If you haven't yet, [review requirements for your portfolio project](https://lambdaschool.github.io/ds/unit2), then submit your dataset.
# - [ ] Plot a confusion matrix for your Tanzania Waterpumps model.
# - [ ] Continue to participate in our Kaggle challenge. Every student should have made at least one submission that scores at least 70% accuracy (well above the majority class baseline).
# - [ ] Submit your final predictions to our Kaggle competition. Optionally, go to **My Submissions**, and _"you may select up to 1 submission to be used to count towards your final leaderboard score."_
# - [ ] Commit your notebook to your fork of the GitHub repo.
# - [ ] Read [Maximizing Scarce Maintenance Resources with Data: Applying predictive modeling, precision at k, and clustering to optimize impact](http://archive.is/DelgE), by Lambda DS3 student <NAME>. His blog post extends the Tanzania Waterpumps scenario, far beyond what's in the lecture notebook.
#
#
# ## Stretch Goals
#
# ### Reading
#
# - [Attacking discrimination with smarter machine learning](https://research.google.com/bigpicture/attacking-discrimination-in-ml/), by Google Research, with interactive visualizations. _"A threshold classifier essentially makes a yes/no decision, putting things in one category or another. We look at how these classifiers work, ways they can potentially be unfair, and how you might turn an unfair classifier into a fairer one. As an illustrative example, we focus on loan granting scenarios where a bank may grant or deny a loan based on a single, automatically computed number such as a credit score."_
# - [Notebook about how to calculate expected value from a confusion matrix by treating it as a cost-benefit matrix](https://github.com/podopie/DAT18NYC/blob/master/classes/13-expected_value_cost_benefit_analysis.ipynb)
# - [Visualizing Machine Learning Thresholds to Make Better Business Decisions](https://blog.insightdatascience.com/visualizing-machine-learning-thresholds-to-make-better-business-decisions-4ab07f823415)
#
#
# ### Doing
# - [ ] Share visualizations in our Slack channel!
# - [ ] RandomizedSearchCV / GridSearchCV, for model selection. (See module 3 assignment notebook)
# - [ ] Stacking Ensemble. (See module 3 assignment notebook)
# - [ ] More Categorical Encoding. (See module 2 assignment notebook)
# + id="lsbRiKBoB5RE"
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="BVA1lph8CcNX"
import pandas as pd
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="e4QOHD8BuMA_" outputId="ca57730d-3965-43f4-e70e-2dd3efd13847"
# %matplotlib inline
import category_encoders as ce
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.impute import SimpleImputer
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestClassifier
# + colab={"base_uri": "https://localhost:8080/"} id="FRgJiTZJ6PDg" outputId="46c3660d-f105-48b6-bc9d-c1b9f60061a7"
train['status_group'].value_counts()
# + id="dA6NWf6P6YNo"
train['needs_to_repair'] = train['status_group'].apply(lambda x : 0 if x == 'functional' else 1)
# + id="c4uxuofp6YDP"
X = train.drop(['needs_to_repair', 'status_group'], axis=1)
y = train['needs_to_repair']
# + id="bYR7c8nB6gkP"
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=42)
# + id="Ra61rQE26h7P"
model = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
RandomForestClassifier(n_estimators=50, n_jobs=-1, random_state=42)
)
# + colab={"base_uri": "https://localhost:8080/"} id="FyIDxCLY6jEf" outputId="8af151c6-a8d6-447e-f0a9-03315b2a0e6c"
model.fit(X_train, y_train)
# + id="ngDtfg006qJ8"
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="VI32w5Cx6qEK" outputId="2a8a7333-365e-410a-a308-341915e48226"
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(model, X_val, y_val, values_format='.0f')
| Copy_of_LS_DS_224_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import re
import spacy
import numpy as np
import gzip
import gensim.downloader
import torch
from torch.utils.data import Dataset, DataLoader
import pickle
from tqdm import tqdm
# +
# train = pd.read_csv("./nlp-getting-started/train.csv")
# test = pd.read_csv("./nlp-getting-started/test.csv")
# train["isTrain"] = True
# test["isTrain"] = False
# full = pd.concat([train, test])
# # print(full)
# def clean_text(row):
# clean = row["text"]
# if len(row["tags"]) != 0:
# for word in row["tags"]:
# clean = clean.replace(word, "")
# if len(row["links"]) != 0:
# for word in row["links"]:
# clean = clean.replace(word, "")
# #only remove the # symbol
# clean = clean.replace("#", "").replace("/", "").replace("(", "").replace(")", "")
# return clean.strip()
# def get_at(row):
# return re.findall("@[\w]+", row["text"])
# def get_http(row):
# return re.findall("http[\:\/\.\w]+", row["text"])
# def get_hashtags(row):
# return re.findall("#[\w]+", row["text"])
# def number_of_tags(row):
# return len(row["tags"])
# def number_of_links(row):
# return len(row["links"])
# def number_of_hashs(row):
# return len(row["hashtags"])
# full["tags"] = full.apply(lambda row: get_at(row), axis = 1)
# full["links"] = full.apply(lambda row: get_http(row), axis = 1)
# full["hashtags"] = full.apply(lambda row: get_hashtags(row), axis = 1)
# full["number_of_tags"] = full.apply(lambda row: number_of_tags(row), axis = 1)
# full["number_of_links"] = full.apply(lambda row: number_of_links(row), axis = 1)
# full["number_of_hashs"] = full.apply(lambda row: number_of_hashs(row), axis = 1)
# full["clean_text"] = full.apply(lambda row: clean_text(row), axis = 1)
# full.sample(5)
# +
with gzip.open('./test.txt.gz', 'rt') as f:
lines = [x.rstrip('\n') for x in f.readlines()]
sentences = [[]]
idx = 0
for line in lines:
if line != '':
sentences[idx].append(line.split()[0])
else:
sentences.append([])
idx += 1
# print(sentences[:100])
for sentence in sentences:
for i, word in enumerate(sentence):
if word == "'s":
sentence[i-1] = sentence[i-1] + "'s"
del sentence[i]
print(sentences[:100])
print(len(sentences))
# -
jpdy_data = pd.read_json("JEOPARDY_QUESTIONS1.json")
sentences = [sent.strip().split(' ') for sent in jpdy_data['question'] if sent.strip()]
# +
nlp = spacy.load('en_core_web_sm')
sub_toks = []
sub_indices = []
new_sentences = []
for i, sentence in tqdm(enumerate(sentences)):
sent = ' '.join(sentence)
# print(sent)
doc = nlp(sent)
temp = [tok for tok in doc if (tok.dep_ == "nsubj")]
if len(temp) == 1:
if str(temp[0]) in sentence:
new_sentences.append(sentence)
sub_toks.append(temp[0])
sub_indices.append(sentence.index(str(temp[0])))
if i + 1 % 200 == 0:
print('{} out of {} done'.format(i+1, len(sentences)))
print(sub_toks[:10])
print(sub_indices[:10])
print(new_sentences[:10])
# +
print(len(new_sentences))
print(len(sub_toks))
print(len(sub_indices))
split = len(new_sentences)//10*9
print(f'Split:{split}')
print(new_sentences[:10])
print(sub_toks[:10])
print(sub_indices[:10])
pickle.dump({'sentences': new_sentences[:split], 'sub_indices': sub_indices[:split]}, open('subdata_jpdy_train.pkl', 'wb'))
pickle.dump({'sentences': new_sentences[split:], 'sub_indices': sub_indices[split:]}, open('subdata_jpdy_test.pkl', 'wb'))
# +
print(list(gensim.downloader.info()['models'].keys()))
glove = gensim.downloader.load('glove-wiki-gigaword-200')
# -
a = glove[['augusts', 'bye']]
print(a.shape)
print('Hi' in glove)
class WordDataset(Dataset):
def __init__(self, sfile):
d = pickle.load(open(sfile, 'rb'))
self.sentences = d['sentences']
self.indices = d['sub_indices']
for sent in self.sentences:
for i, word in enumerate(sent):
sent[i] = word.lower().replace("'", "")
if sent[i] == '':
del sent[i]
self.glove = gensim.downloader.load('glove-wiki-gigaword-200')
self.sentence_embeddings = []
for sent in self.sentences:
temp = []
for word in sent:
if word in self.glove:
temp.append(self.glove[word])
self.sentence_embeddings.append(np.array(temp))
def __len__(self):
return len(self.sentence_embeddings)
def __getitem__(self, idx):
return (self.sentences[idx], self.sentence_embeddings[idx], self.indices[idx])
train_dataset = WordDataset('subdata.pkl')
test_dataset = WordDataset('subdata_test.pkl')
print(train_dataset[1][1].shape)
| preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "skip"}
# Required to load webpages
from IPython.display import IFrame
# + [markdown] slideshow={"slide_type": "slide"}
# [Table of contents](../toc.ipynb)
#
# <img src="https://github.com/sphinx-doc/sphinx/raw/master/doc/_static/sphinx.png" alt="Sphinx" width="400" align="right">
#
# # Sphinx
#
# Sphinx is a python library to create beautiful documentation pages for Python software projects. It was written for the Python doc itself and became number one tool for many other packages. Some examples of the Sphinx's output are:
# * [the Python documentation](https://docs.python.org/),
# * [scikit-learn](https://scikit-learn.org/stable/index.html),
# * [numpy](https://numpy.org/).
#
# Please find here an extensive list of projects using Sphinx: [https://www.sphinx-doc.org/en/master/examples.html](https://www.sphinx-doc.org/en/master/examples.html).
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Why care about documentation?
#
# There are good reasons to work continuously on documentation. New developers need to know about the "mechanics" of the software, users want to know how to apply the software and also want to know some background information,...
#
# The list of reasons for documentation is actually very long and you see that different people are addressed.
#
# This is the reason why it is recommended to split the documentation into four parts:
# 1. Tutorials (learning-oriented),
# 2. How-to guides (goal-oriented),
# 3. Explanation (understanding-oriented),
# 4. Reference (information-oriented),
#
# please read this great blog post: ["What nobody tells you about documentation"](https://www.divio.com/blog/documentation/).
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Sphinx getting started
#
# Sphinx is available as conda and pip package and there is a quickstart command which sets up the required folder structure, options and files.
#
# Sphinx docu is written with [reStructuredText](https://docutils.sourceforge.io/rst.html) markup language in `.rst` files. The syntax is explained in the previous link and easy to learn.
#
# Let us start with a small Sphinx documentation.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Exercise: `sphinx-quickstart` (10 minutes)
#
# <img src="../_static/exercise.png" alt="Exercise" width="75" align="left">
#
# * Install Sphinx.
# * Create a folder `doc` and change your terminal path to it.
# * Run `sphinx-quickstart` from a terminal and answer the questions.
# * The files `make.bat`, `Makefile`, `conf.py`, `index.rst` and folders `build`, `static` and `templates` should show up.
# * Now build this documentation with `make html`.
# * To see the result open the file `build/html/index.html` in your browser.
# * Take a look at the `conf.py` file, which contains the configuration and the `index.rst` file which contains the text of the main docu page.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Short intro to reStructuredText
#
# Here some short comments on reStructuredText syntax.
#
# ### Headers
#
# Can be generated with
# ```
# Chapter 1 Title
# ===============
#
# Section 1.1 Title
# -----------------
#
# Subsection 1.1.1 Title
# ~~~~~~~~~~~~~~~~~~~~~~
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Lists
# Bullets use stars.
# ```
# * bullet one
# * bullet two
# - sub bullet
# + subsub bullet
# ```
# And enumerated lists use numbers or letters.
# ```
# A. bullet one
# B. bullet two
# 1. sub bullet
# a. subsub bullet
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Text styles
#
# `*italics*`, `**bold**`, and back ticks are used for `fixed spaced text`
#
# ### Images
#
# `.. image:: some_image.png`
#
# ### Hyperlinks
#
# `google.com <http://google.com>`_
#
# ### Try reStructuredText online
#
# There is a website where you can paste and try reStructured text interactively [http://rst.ninjs.org/](http://rst.ninjs.org/).
#
# Please find more details in the [reStruckturedText docu](https://docutils.sourceforge.io/rst.html), [cheat sheet](https://docutils.sourceforge.io/docs/user/rst/cheatsheet.txt), and [Sphinx page about reStructuredText](https://www.sphinx-doc.org/en/master/usage/restructuredtext/index.html).
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Sphinx extension autodoc
#
# [Sphinx autodoc](https://www.sphinx-doc.org/en/master/usage/quickstart.html#autodoc) is an extension to generate documentation out of doc strings in Python code. Hence, you can document your code directly in the doc string with this extension.
#
# You can configure this extension with
#
# `extensions = ['sphinx.ext.autodoc']` in the `conf.py` file.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### An autodoc example
#
# Assume this file and folder structure of a project,
# ```
# |___ doc
# |___ conv.py
# |___ index.rst
# |___ make.bat
# |___ Makefile
# |___src
# |___ __init__.py
# |___ matrix_comp.py
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# where the `matrix_comp.py` file contains
#
# ```python
# """
# .. module:: matrix_comp
#
# The :py:mod:`matrix_comp` module provides ...
# """
#
# def matrix_multiply(a, b):
# """ The :py:func:`matrix_algebra.matrix_multiply` function computes
# the product of two matrices.
#
# Args:
# a (numpy.ndarray): first matrix
# b (numpy.ndarray): second matrix
#
# Returns:
# (numpy.ndarray): product of a and b
#
# """
# return a @ b
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# The autodoc extension needs these settings in `conf.py`.
#
# ```python
# import os
# import sys
# sys.path.insert(0, os.path.abspath('.'))
#
# extensions = ['sphinx.ext.autodoc']
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# To include the module in your documentation, call `automodule` in `index.rst` file like this.
#
# ```
# Matrix computation module
# -------------------------
# .. automodule:: src.matrix_comp
# :members:
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# If you run `make html`, documentation start page will result in.
#
# <img src="sphinx_result.png" alt="Sphinx_result" width="800">
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Sphinx-Gallery extension
#
# [Sphinx-Gallery](https://sphinx-gallery.github.io/stable/index.html) renders Python files as notebooks and is ideal if you want to write tutorials for your software.
#
# Sphinx gallery must be installed as conda or pip package and must be called in `conf.py` with
# ```python
# extensions = [...
# 'sphinx_gallery.gen_gallery',
# ]
# ```
#
# Many popular packages like scikit-learn use Sphinx gallery to present tutorials, see also [who uses Sphinx-Gallery](https://sphinx-gallery.github.io/stable/projects_list.html).
#
# In the next cell some gallery examples from Sphinx-Gallery page are linked.
# + slideshow={"slide_type": "subslide"}
IFrame(src='https://sphinx-gallery.github.io/stable/auto_examples/index.html',
width=1000, height=600)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## References
#
# * There are many more extensions for Sphinx available. This was just a short introduction.
#
# * There is one German book called [Software-Dokumentation mit Sphinx](http://www.worldcat.org/oclc/889425279), but best is to look at github projects which have good documentation.
#
# * It is very common to store the Sphinx files in a `doc` folder on root level. You can learn from open source best how to create more advanced documents with Sphinx.
| 03_software-development/02_sphinx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <center> <img src ="https://i.postimg.cc/1X8H7YYt/BITS-Logo.png" width = "400" alt="BITS Pilani Logo" /> </center>
# <font color='green'> <h1> <center> Python - Output Statements </center> </h1> </font>
#
# <b>First Python Program<b>
# Communicate to the outside world with a greeting <br>
print("Hello World from Python!")
# Whatever is specified in the print statement gets outputed on the console
print('Single quotes also works!')
print("Multiline output \n can also be printed.")
print('String concatenation', 'also', 'works.')
# <b>Python Outputs </b>
# print function requires paranthesis around its arguments. Anything within the quotes will be printed as it is.
print('3+4') #treats '3+4' as string and outputs as is
print(3+4) #evaluates the expression and then prints the outcome
print('3 + 4 = ', (3+4) ) # strings and expressions can be concatenanted in the single print function call
print('age ', 35, 'name', 'ABC') # strings and numbers can also be concatenated in single statement
# <b>Using Separator<b>
# The detault separator between two values in print statements is space.
print("This", "is", "first", "line")
# The default separator can be changed with optional parameter 'sep' as follows
print("This", "is", "first", "line", sep=", ")
# <b>Using end<b>
# The default endline character is newline i.e. \n
print("this is first line")
print("this is second line")
# This can be changed by optional parameter 'end' as shown below
print("this is first line", end='.')
print("this is second line")
# <b>Exercise <b>
# Q.1 Print a traingle like one below
# + active=""
# *
# **
# ***
# ****
# *****
# -
#Try it here
star_var = ""
for i in range(1, 6):
star_var += "*"
print(star_var)
# Q2. Write a program that prints result of following expression.
#
# + active=""
# 123 - 282
# ----------
# 47.34 + 23
# -
#Try it here
print((123-282)/(47.34+23))
| Python Fundamentals for Data Science/S1/Additional Notebooks S1/0_Python - Output Statements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.10.4 (''fantasysidelines.venv'': venv)'
# language: python
# name: python3
# ---
# + tags=[]
# -*- coding: utf-8 -*-
"""
{Description}
MIT License
Copyright (c) 2021, Fantasy-Sidelines
Permission is hereby granted, free of charge, to any person obtaining a copy
of this software and associated documentation files (the "Software"), to deal
in the Software without restriction, including without limitation the rights
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
copies of the Software, and to permit persons to whom the Software is
furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY +OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
Sportradar API details and documentation: https://developer.sportradar.com/docs
MySportsFeeds API details and documentation: https://www.mysportsfeeds.com/data-feeds/api-docs/
www.pro-football-reference.come and documentation: https://www.sports-reference.com/termsofuse.html
"""
import time
import pyodbc
import os
# import pandas as pd
import pandas as pd
import numpy as np
from sqlalchemy import create_engine
from sqlalchemy import inspect
from sqlalchemy.engine import URL
from dotenv import load_dotenv
__author__ = "<NAME>"
__copyright__ = "Copyright (c) 2021, Fantasy-Sidelines"
__credits__ = ["<NAME>", "Sportradar API", "Fantasy Sharks", "MySportsFeeds API"]
__license__ = "MIT License"
__version__ = "1.1.0"
__maintainer__ = "<NAME>"
__email__ = "<EMAIL>"
__status__ = "Dev"
from sql_upload import *
# + tags=[]
load_dotenv()
sql_driver = os.getenv("sql_driver")
sql_server = os.getenv("sql_server")
sql_database = os.getenv("sql_database")
sql_username = os.getenv("sql_username")
sql_password = os.getenv("sql_password")
# api_key = os.getenv("sportradar_apikey")
# year = [2020, 2019, 2018, 2017, 2016, 2015]
connection_string = (
"DRIVER={"
+ sql_driver
+ "};SERVER="
+ sql_server
+ ";DATABASE="
+ sql_database
+ ";UID="
+ sql_username
+ ';PWD='
+ sql_password
+ ";Trusted_Connection=yes;"
)
# cxn = pyodbc.connect(connection_string)
connection_url = URL.create("mssql+pyodbc", query={"odbc_connect": connection_string})
engine = create_engine(connection_url)
conn = engine.connect()
inspector = inspect(engine)
print(inspector.get_table_names())
# + tags=[]
pd.set_option("display.max_columns", None)
pd.set_option("display.max_colwidth", 15)
pd.set_option("display.max_rows", None)
# schedule_stats_api_sql(api_key, year, engine)
# snaps(2016, 2020, engine)
# injuries(y1, y2, engine)
# practice_participation(season_start, season_end, engine)
# player_table(engine)
# game_table(engine)
# season_table(engine)
# week_table(engine)
# team_table(engine)
# venue_table(engine)
# calendar_table("8/1/2016", "2/1/2021", engine)
# weekly_stats_offense(conn, engine)
# calendar_ID = pd.read_sql_table("IDCalendarTable", con=conn)
# game_ID = pd.read_sql_table("IDGameTable", con=conn)
# team_ID = pd.read_sql_table("IDTeamTable", con=conn)
# venue_ID = pd.read_sql_table("IDVenueTable", con=conn)
# week_ID = pd.read_sql_table("IDWeekTable", con=conn)
# player_practice = pd.read_sql_table("playerPractice", con=conn)
# player_snaps = pd.read_sql_table("playerSnaps", con=conn)
# player_stats = pd.read_sql_table("playerStats", con=conn)
# schedule = pd.read_sql_table("schedule", con=conn)
# team_stats = pd.read_sql_table("teamStats", con=conn)
weekly_stats = pd.read_sql_table("weeklyStats", con=conn)
# +
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
# %matplotlib inline
weekly_stats.sort_values(
["season.year", "week.sequence", "team.id", "player.id"], inplace=True
)
# +
variables = ["off.snaps", "total_fan_pts.half.kick_yrds"]
position = ["QB", "RB", "WR", "TE"]
status_cats = ["date1.status", "date2.status", "date3.status", "game.status"]
injury_cats = [
"head",
"face",
"neck",
"shoulder",
"upper_arm",
"elbow",
"forearm",
"wrist",
"hand_finger",
"thumb",
"back",
"chest",
"abdomen",
"hip",
"groin",
"quadricep",
"hamstring",
"thigh",
"knee",
"lower_leg",
"achilles",
"ankle",
"foot",
"toe",
"illness",
]
alpha = 0.05
hypothesis_results = {
2: "Reject the null: There is significance between injury status/injury and fantasy production.".upper(),
1: "Reject the null: There is significance between injury status/injury and offense snap counts.".upper(),
}
# -
hypothesis_1 = weekly_stats[
(weekly_stats["played"] >= 1) & (weekly_stats["off.snaps"] >= 15)
]
# + tags=[]
for pos in position:
for status_day in status_cats:
for injury in injury_cats:
hypothesis_1_full = hypothesis_1[
(
(hypothesis_1[status_day].isnull())
| (hypothesis_1[status_day] == "Full")
)
& (hypothesis_1["player.position"] == pos)
]
hypothesis_1_status = hypothesis_1[
~(
(hypothesis_1[status_day].isnull())
| (hypothesis_1[status_day] == "Full")
)
& (hypothesis_1["player.position"] == pos)
& (hypothesis_1[injury] > 0)
]
try:
p_mannwhiteyu_snaps = stats.mannwhitneyu(
hypothesis_1_full[variables[0]], hypothesis_1_status[variables[0]]
)[1]
if p_mannwhiteyu_snaps > alpha:
pass
elif p_mannwhiteyu_snaps <= alpha:
print(
pos,
injury.upper(),
status_day.upper(),
"Snaps",
hypothesis_results[1],
"p_value = " + str(p_mannwhiteyu_snaps),
"No designation Snaps:",
len(hypothesis_1_full[variables[0]]),
"Designation Snaps:",
len(hypothesis_1_status[variables[0]]),
"\n",
sep="\n",
)
p_mannwhiteyu_fp = stats.mannwhitneyu(
hypothesis_1_full[variables[1]], hypothesis_1_status[variables[1]]
)[1]
if p_mannwhiteyu_fp > alpha:
pass
elif p_mannwhiteyu_fp <= alpha:
print(
pos,
injury.upper(),
status_day.upper(),
"FP",
hypothesis_results[2],
"p_value = " + str(p_mannwhiteyu_fp),
"No designation FP:",
len(hypothesis_1_full[variables[1]]),
"Designation FP:",
len(hypothesis_1_status[variables[1]]),
"\n",
sep="\n",
)
except:
continue
# -
for status_day in status_cats:
hypothesis_1_full = hypothesis_1[
((hypothesis_1[status_day].isnull()) | (hypothesis_1[status_day] == "Full"))
]
hypothesis_1_status = hypothesis_1[
# (
# (hypothesis_1[status_day] == "Questionable")
# | (hypothesis_1[status_day] == "Doubtful")
# | (hypothesis_1[status_day] == "Limited")
# | (hypothesis_1[status_day] == "DNP")
# )
~((hypothesis_1[status_day].isnull()) | (hypothesis_1[status_day] == "Full"))
]
try:
p_mannwhiteyu_snaps = stats.mannwhitneyu(
hypothesis_1_full[variables[0]], hypothesis_1_status[variables[0]]
)[1]
if p_mannwhiteyu_snaps > alpha:
pass
elif p_mannwhiteyu_snaps <= alpha:
print(
status_day.upper(),
"Snaps",
hypothesis_results[1],
"p_value = " + str(p_mannwhiteyu_snaps),
"No designation Snaps:",
len(hypothesis_1_full[variables[0]]),
"Designation Snaps:",
len(hypothesis_1_status[variables[0]]),
"\n",
sep="\n",
)
p_mannwhiteyu_fp = stats.mannwhitneyu(
hypothesis_1_full[variables[1]], hypothesis_1_status[variables[1]]
)[1]
if p_mannwhiteyu_fp > alpha:
pass
elif p_mannwhiteyu_fp <= alpha:
print(
status_day.upper(),
"FP",
hypothesis_results[2],
"p_value = " + str(p_mannwhiteyu_fp),
"No designation FP:",
len(hypothesis_1_full[variables[1]]),
"Designation FP:",
len(hypothesis_1_status[variables[1]]),
"\n",
sep="\n",
)
except:
continue
"""
paired t-test
assume normal distribution
assume no outliers
Wilcoxon Signed-Rank test:
not normal distribution
"""
conn.close()
# cxn.close()
# +
# import seaborn as sns
# import matplotlib.pyplot as plt
# from scipy import stats
# # %matplotlib inline
# weekly_stats = pd.read_sql_table("weeklyStats", con=conn)
# mask = (weekly_stats["player.position"] == "RB") & (weekly_stats["total.snaps"] >= 5) & (weekly_stats["total_fan_pts.half.kick_yrds"] >= 0)
# sns.displot(weekly_stats[mask], x="total_fan_pts.half.kick_yrds", binwidth=1, kde=True)
# corr = weekly_stats.corr()
# def background_pandas(x):
# try:
# if x >= 0.5:
# bg_color = "green"
# elif x <= -0.5:
# bg_color = "green"
# else:
# bg_color = ""
# return f"background-color : {bg_color}"
# except:
# return "background-color: ''"
# corr.style.format(precision=3).applymap(background_pandas).to_excel("correlations.xlsx", engine="openpyxl")
| scripts/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Import all the necessary libraries here
import numpy as np
import pandas as pd
import seaborn as sns
# # Footwear
# The given dataset contains the profits generated(in %) by all the suppliers of a footwear company in 4 major cities of India - Delhi, Mumbai, Jaipur and Hyderabad. The company wants to invest more money in the city that is showing the most promise. Analyse the dataset and answer the following questions.
#loading data
df=pd.read_csv("Footwear_v2.csv")
df.head()
# +
#we note that there are no null values, and the values are treated as objects and not floats, we will have to clean the
# '%' sign at the end of all and change it to float
#we will write a function to do this like the last session
def clean(string):
clean="".join(filter(lambda x: x!='%', string))
return float(clean)
# you can also use replace
# def clean(val):
# return float(val.replace("%",""))
#
# -
# we also see the supplier column has few 'S' as upper case and few lowercase
#lets clean that too
def supply_cleaner(string):
return string.lower()
#clean the df
df['Supplier']=df['Supplier'].apply(supply_cleaner)
df['Mumbai']=df['Mumbai'].apply(clean)
df['Delhi']=df['Delhi'].apply(clean)
df['Jaipur']=df['Jaipur'].apply(clean)
df['Hyderabad']=df['Hyderabad'].apply(clean)
# ## 1. Average
# Q1)The absolute difference in the average profit percentages of Delhi and Mumbai comes out to be approximately ____
#
# a) 1.67
#
# b) 1.57
#
# c) 1.77
#
# d) 1.47
#
#Your code here
df.describe()
# ## 2. Invest More
# Q2) Which city amongst the four should the company invest more money in?
#
# Hint: You need to see which city is showing most consistency in profits
#
# a) Delhi
#
# b) Mumbai
#
# c) Jaipur
#
# d) Hyderabad
#
#
#Your code here
sub_df=df[['Delhi', 'Mumbai', 'Jaipur', 'Hyderabad']]
sub_df.boxplot()
# # Crypto Currencies
# The following datasets contain the prices of some popular cryptocurrencies such as bitcoin, litecoin, ethereum, monero, neo, quantum and ripple.Now, you would like to know how the prices of these currencies vary with each other.
#
# The datasets containing their price values over several days is mentioned. The attributes are as follows:
#
# - Date - The date of trading
# - Open - Opening Price
# - High - Highest Price
# - Low - Lowest Price
# - Close - Closing Price
# - Volume - Total Volume
# - Market Cap- Market Capitalisation
#
#
#
bitcoin = pd.read_csv("crypto_data/bitcoin_price.csv")
ethereum = pd.read_csv("crypto_data/ethereum_price.csv")
litecoin = pd.read_csv("crypto_data/litecoin_price.csv")
monero = pd.read_csv("crypto_data/monero_price.csv")
neo = pd.read_csv("crypto_data/neo_price.csv")
qtum = pd.read_csv("crypto_data/qtum_price.csv")
ripple = pd.read_csv("crypto_data/ripple_price.csv")
print("Bitcoin: ", bitcoin.shape)
print("Ethereum: ", ethereum.shape)
print("Litecoin: ", litecoin.shape)
print("Monero: ", monero.shape)
print("Neo: ", neo.shape)
print("Qtum: ", qtum.shape)
print("Ripple: ", ripple.shape)
bitcoin = pd.concat([bitcoin.Date, bitcoin.Close], axis=1)
ethereum = pd.concat([ethereum.Date, ethereum.Close], axis=1)
litecoin = pd.concat([litecoin.Date, litecoin.Close], axis=1)
monero = pd.concat([monero.Date, monero.Close], axis=1)
neo = pd.concat([neo.Date, neo.Close], axis=1)
qtum = pd.concat([qtum.Date, qtum.Close], axis=1)
ripple = pd.concat([ripple.Date, ripple.Close], axis=1)
data_frames = [ethereum, litecoin, monero, neo, qtum, ripple]
crypto_coins = bitcoin
for df in data_frames:
crypto_coins = pd.merge(crypto_coins, df, on="Date", how="inner")
crypto_coins.columns = ["Date", "bitcoin", "ethereum", "litecoin", "monero", "neo", "qtum", "ripple"]
crypto_coins.head()
# ## 1. Correct Statements
# Q1) Combine all the datasets by merging on the date column and create a dataframe with only the closing prices for all the currencies. Next, create a pair plot with all these columns and choose the correct statements from the given ones:
#
# I)There is a good trend between litecoin and monero, one increases as the other
#
# II)There is a weak trend between bitcoin and neo.
#
# a)I
#
# b)II
#
# c)Both I and II
#
# d)None of the above.
#
#Your code here
sns.pairplot(crypto_coins)
# ## Heatmap
# Q2)As mentioned earlier, Heat Maps are predominantly utilised for analysing Correlation Matrix. A high positive correlation (values near 1) means a good positive trend - if one increases, then the other also increases. A negative correlation on the other hand(values near -1) indicate good negative trend - if one increases, then the other decreases. A value near 0 indicates no correlation, as in one variable doesn’t affect the other.
#
#
# a)Ethereum and Quantum have high correlation
#
# b)Neo and Bitcoin have pretty low correlation
#
# c)Ethereum has similar correlation with litecoin and neo
#
#
#Your code here
crypto_corr = crypto_coins.corr()
sns.heatmap(crypto_corr,cmap="Greens", annot=True)
| Course_1-PreLaunch_Preparatory_Content/Module_3-Data_Visualisation_in_Python/2-Introduction_to_Data_Visualization/Practice_Exercise_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Carrinheiros
#
# "Carrinheiros" are collectors of recyclable materials that use human propulsion vehicles in the selective collection. The problem is that route can be very tiring for waste pickers according to the increase in vehicle weight and the roads' slope. Therefore, this work proposes a route suggestion service to minimize the collection time or the pickers' physical effort on the route. The characteristics of the scenario addressed in this proposal are the distance between collect points, the depot's geographical position, the inclination of the roads, and the use of vehicles with human propulsion. Besides, this work should consider the variation in vehicle weight along the route.
# <img src="https://raw.githubusercontent.com/vivirodrigues/Carrinheiros/main/documentation/carrinheiro.png">
# Tools employed in this work:
#
# * This work used the osmnx package to construct geographic graphs in Python. The osmnx documentation is available on https://osmnx.readthedocs.io/en/stable/.
#
# * The tool Networkx is used to manipulate and construct graphs in Python. The Networkx documentation is available on https://networkx.org/documentation/stable/.
#
# * The geographic data was obtained from Open Street Map (https://www.openstreetmap.org/), and Brazil's elevation data was obtained from Topodata (http://www.dsr.inpe.br/topodata/index.php).
#
# * The proposal's validation was performed through computer simulations using Simulation of Urban MObility (SUMO). The SUMO documentation is available on https://sumo.dlr.de/docs/.
#
from IPython.display import Image
# To execute the simulations, access 'Main.py' file. The Main.py has the "get_seed" function, which provides the random seed to guarantee the reproducibility of the simulation. The seed_id input is the index of 'seeds' vector.
def get_seed(seed_id):
seeds = [960703545, 1277478588, 1936856304,
186872697, 1859168769, 1598189534,
1822174485, 1871883252, 694388766,
188312339, 773370613, 2125204119,
2041095833, 1384311643, 1000004583,
358485174, 1695858027, 762772169,
437720306, 939612284, 425414105,
1998078925, 981631283, 1024155645,
558746720, 1349341884, 678622600,
1319566104, 538474442, 722594620,
1700738670, 1995749838, 1147024708,
346983590, 565528207, 513791680,
1996632795, 2081634991, 1769370802,
349544396, 1996610406, 1973272912,
1972392646, 605846893, 934100682,
222735214, 2101442385, 2009044369,
1895218768, 701857417, 89865291,
144443207, 720236707, 822780843,
898723423, 1644999263, 985046914,
1859531344, 1024155645, 764283187,
778794064, 683102175, 1334983095,
1072664641, 999157082]
return seeds[seed_id]
# The 'main' function executes the experiments on the SUMO simulator. First of all, it sets the characteristics of the scenario: number of collect points, values of vehicle weight increment, city of the scenario, number of repetitions of the simulations, etc. Then, it creates pseudo-random collect points/stop points of the scenario. Finally, it calls the 'create_route' function.
# +
def main():
# number of collect points
n_points = 10
# maximum increment of vehicle weight at the collect point (material mass)
max_mass_material = 50
# random seed of mass increment
random.seed(get_seed(0))
# scenarios: 'Belo Horizonte' and 'Belem'
city = 'Belo Horizonte'
# mean of the gaussian function that creates the collect points
if city == 'Belo Horizonte':
mean_lon = [-43.9438]
mean_lat = [-19.9202]
elif city == 'Belem':
mean_lon = [-48.47000]
mean_lat = [-1.46000]
# standard deviation of the gaussian function that creates the collect points
sigma = 0.005
# vector with vehicle weight increment in the collect points
mass_increments = [random.randint(0, max_mass_material) for i in range(n_points-2)]
# add unit of measurement of vehicle weight increment in the collect points
material_weights = [(mass_increments[i], 'Kg') for i in range(n_points-2)]
# the arrival point must not increment the vehicle weight
material_weights.append((0, 'Kg'))
# the starting point must not increment the vehicle weight
material_weights.insert(0, (0, 'Kg'))
# number of repetitions of the simulations
n_seeds = 30
json_files = []
materials = {}
for n in range(0, n_seeds):
# gets the current random seed
random.seed(get_seed(n))
# creates a vector with pseudo-random longitude and latitude values
longitudes = [random.gauss(mean_lon[0], sigma) for i in range(n_points)]
latitudes = [random.gauss(mean_lat[0], sigma) for i in range(n_points)]
# creates the collect points with longitude and latitudes values
stop_points = [(float(latitudes[i]), float(longitudes[i])) for i in range(len(latitudes))]
# creates a dict with vehicle mass increment in each collect point
[materials.update([((latitudes[i], longitudes[i]), material_weights[i])]) for i in range(len(latitudes))]
# creates the routes and writes the simulation results on json file. It returns the name of the file
json_files = create_route(stop_points, materials, json_files, n)
# -
# The "create route" function generates two graphs: the geographic scenario graph and the ordering graph. It uses the osmnx to creates the geographic graph based on Open Street Map data. Besides, the networkx is used to generates the ordering graph, which is complete, and each vertex corresponds to a collection point. The Nearest Neighbor search is the heuristic used to order the vertexes. Also, the Shortest Path Faster Algorithm is employed to create a route between two collection points in the geographic graph.
#
# Figure 1 shows the ordering graph with a red connection between 8 and 9 vertexes. This connection corresponds to the path in the geographic graph exhibited in Figure 2.
# <table>
# <tr>
# <td>
# <img src="https://raw.githubusercontent.com/vivirodrigues/Carrinheiros/main/documentation/grafo1.png">
# </td>
# <td>
# <img src="https://raw.githubusercontent.com/vivirodrigues/Carrinheiros/main/documentation/grafo_a.png">
# </td>
# </tr>
# <tr>
# <td>
# <p>Figure 1: Ordering graph</p>
# </td>
# <td>
# <p>Figure 2: Geogrpahic graph</p>
# </td>
# </tr>
# </table>
# The combination of SPFA and nearest neighbor provides routes based on three edge costing policies: Less Work Policy (LWP), Less Impedance Policy (LIP), and Short Distance Policy (SDP). The LWP minimizes the work required to push the vehicle, LIP avoids steep slopes and SDP minimizes the total distance. Using the first random seed (960703545), the algorithm generates three routes based on the policies:
# <table>
# <tr>
# <td>
# <img src="https://raw.githubusercontent.com/vivirodrigues/Carrinheiros/main/documentation/weight.png">
# </td>
# <td>
# <img src="https://raw.githubusercontent.com/vivirodrigues/Carrinheiros/main/documentation/impedance.png">
# </td>
# <td>
# <img src="https://raw.githubusercontent.com/vivirodrigues/Carrinheiros/main/documentation/distance.png">
# </td>
# </tr>
# <tr>
# <td>
# <p>Figure 3: Route generated using LWP</p>
# </td>
# <td>
# <p>Figure 4: Route generated using LIP</p>
# </td>
# <td>
# <p>Figure 5: Route generated using SDP</p>
# </td>
# </tr>
# </table>
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multiple Kernel Learning
# #### By <NAME> - <a href="https://github.com/Saurabh7">github.com/Saurabh7</a>
# This notebook is about multiple kernel learning in shogun. We will see how to construct a combined kernel, determine optimal kernel weights using MKL and use it for different types of [classification](http://en.wikipedia.org/wiki/Statistical_classification) and [novelty detection](http://en.wikipedia.org/wiki/Novelty_detection).
# 1. [Introduction](#Introduction)
# 2. [Mathematical formulation](#Mathematical-formulation-(skip-if-you-just-want-code-examples))
# 3. [Using a Combined kernel](#Using-a-Combined-kernel)
# 4. [Example: Toy Data](#Prediction-on-toy-data)
# 1. [Generating Kernel weights](#Generating-Kernel-weights)
# 5. [Binary classification using MKL](#Binary-classification-using-MKL)
# 6. [MKL for knowledge discovery](#MKL-for-knowledge-discovery)
# 7. [Multiclass classification using MKL](#Multiclass-classification-using-MKL)
# 8. [One-class classification using MKL](#One-class-classification-using-MKL)
# +
# %matplotlib inline
import os
import numpy as np
import matplotlib.pyplot as plt
import shogun as sg
SHOGUN_DATA_DIR=os.getenv('SHOGUN_DATA_DIR', '../../../data')
# -
# ### Introduction
# <em>Multiple kernel learning</em> (MKL) is about using a combined kernel i.e. a kernel consisting of a linear combination of arbitrary kernels over different domains. The coefficients or weights of the linear combination can be learned as well.
#
# [Kernel based methods](http://en.wikipedia.org/wiki/Kernel_methods) such as support vector machines (SVMs) employ a so-called kernel function $k(x_{i},x_{j})$ which intuitively computes the similarity between two examples $x_{i}$ and $x_{j}$. </br>
# Selecting the kernel function
# $k()$ and it's parameters is an important issue in training. Kernels designed by humans usually capture one aspect of data. Choosing one kernel means to select exactly one such aspect. Which means combining such aspects is often better than selecting.
#
# In shogun the [MKL](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MKL.html) is the base class for MKL. We can do classifications: [binary](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MKLClassification.html), [one-class](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MKLOneClass.html), [multiclass](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MKLMulticlass.html) and regression too: [regression](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MKLRegression.html).
# ### Mathematical formulation (skip if you just want code examples)
# </br>In a SVM, defined as:
# $$f({\bf x})=\text{sign} \left(\sum_{i=0}^{N-1} \alpha_i k({\bf x}, {\bf x_i})+b\right)$$</br>
# where ${\bf x_i},{i = 1,...,N}$ are labeled training examples ($y_i \in {±1}$).
#
# One could make a combination of kernels like:
# $${\bf k}(x_i,x_j)=\sum_{k=0}^{K} \beta_k {\bf k_k}(x_i, x_j)$$
# where $\beta_k > 0$ and $\sum_{k=0}^{K} \beta_k = 1$
#
# In the multiple kernel learning problem for binary classification one is given $N$ data points ($x_i, y_i$ )
# ($y_i \in {±1}$), where $x_i$ is translated via $K$ mappings $\phi_k(x) \rightarrow R^{D_k} $, $k=1,...,K$ , from the input into $K$ feature spaces $(\phi_1(x_i),...,\phi_K(x_i))$ where $D_k$ denotes dimensionality of the $k$-th feature space.
#
# In MKL $\alpha_i$,$\beta$ and bias are determined by solving the following optimization program. For details see [1].
#
# $$\mbox{min} \hspace{4mm} \gamma-\sum_{i=1}^N\alpha_i$$
# $$ \mbox{w.r.t.} \hspace{4mm} \gamma\in R, \alpha\in R^N \nonumber$$
# $$\mbox {s.t.} \hspace{4mm} {\bf 0}\leq\alpha\leq{\bf 1}C,\;\;\sum_{i=1}^N \alpha_i y_i=0 \nonumber$$
# $$ {\frac{1}{2}\sum_{i,j=1}^N \alpha_i \alpha_j y_i y_j \leq \gamma}, \forall k=1,\ldots,K\nonumber\\
# $$
#
#
# Here C is a pre-specified regularization parameter.
# Within shogun this optimization problem is solved using [semi-infinite programming](http://en.wikipedia.org/wiki/Semi-infinite_programming). For 1-norm MKL one of the two approaches described in [1] is used.
# The first approach (also called the wrapper algorithm) wraps around a single kernel SVMs, alternatingly solving for $\alpha$ and $\beta$. It is using a traditional SVM to generate new violated constraints and thus requires a single kernel SVM and any of the SVMs contained in shogun can be used. In the MKL step either a linear program is solved via [glpk](http://en.wikipedia.org/wiki/GNU_Linear_Programming_Kit) or cplex or analytically or a newton (for norms>1) step is performed.
#
# The second much faster but also more memory demanding approach performing interleaved optimization, is integrated into the chunking-based [SVMlight](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1SVMLight.html).
#
#
# ### Using a Combined kernel
# Shogun provides an easy way to make a combination of kernels using the [CombinedKernel](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CombinedKernel.html) class, to which we can append any [kernel](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1Kernel.html) from the many options shogun provides. It is especially useful to combine kernels working on different domains and to combine kernels looking at independent features and requires [CombinedFeatures](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CombinedFeatures.html) to be used. Similarly the CombinedFeatures is used to combine a number of feature objects into a single CombinedFeatures object
kernel = sg.CombinedKernel()
# ### Prediction on toy data
# In order to see the prediction capabilities, let us generate some data using the [GMM](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CGMM.html) class. The data is sampled by setting means ([GMM notebook](http://www.shogun-toolbox.org/static/notebook/current/GMM.html)) such that it sufficiently covers X-Y grid and is not too easy to classify.
# +
num=30;
num_components=4
means=np.zeros((num_components, 2))
means[0]=[-1,1]
means[1]=[2,-1.5]
means[2]=[-1,-3]
means[3]=[2,1]
covs=np.array([[1.0,0.0],[0.0,1.0]])
# gmm=sg.distribution("GMM")
# gmm.set_pseudo_count(num_components)
gmm=sg.GMM(num_components)
[gmm.set_nth_mean(means[i], i) for i in range(num_components)]
[gmm.set_nth_cov(covs,i) for i in range(num_components)]
gmm.set_coef(np.array([1.0,0.0,0.0,0.0]))
xntr=np.array([gmm.sample() for i in range(num)]).T
xnte=np.array([gmm.sample() for i in range(5000)]).T
gmm.set_coef(np.array([0.0,1.0,0.0,0.0]))
xntr1=np.array([gmm.sample() for i in range(num)]).T
xnte1=np.array([gmm.sample() for i in range(5000)]).T
gmm.set_coef(np.array([0.0,0.0,1.0,0.0]))
xptr=np.array([gmm.sample() for i in range(num)]).T
xpte=np.array([gmm.sample() for i in range(5000)]).T
gmm.set_coef(np.array([0.0,0.0,0.0,1.0]))
xptr1=np.array([gmm.sample() for i in range(num)]).T
xpte1=np.array([gmm.sample() for i in range(5000)]).T
traindata=np.concatenate((xntr,xntr1,xptr,xptr1), axis=1)
trainlab=np.concatenate((-np.ones(2*num), np.ones(2*num)))
testdata=np.concatenate((xnte,xnte1,xpte,xpte1), axis=1)
testlab=np.concatenate((-np.ones(10000), np.ones(10000)))
#convert to shogun features and generate labels for data
feats_train=sg.features(traindata)
labels=sg.BinaryLabels(trainlab)
# -
_=plt.jet()
plt.figure(figsize=(18,5))
plt.subplot(121)
# plot train data
_=plt.scatter(traindata[0,:], traindata[1,:], c=trainlab, s=100)
plt.title('Toy data for classification')
plt.axis('equal')
colors=["blue","blue","red","red"]
# a tool for visualisation
from matplotlib.patches import Ellipse
def get_gaussian_ellipse_artist(mean, cov, nstd=1.96, color="red", linewidth=3):
vals, vecs = np.linalg.eigh(cov)
order = vals.argsort()[::-1]
vals, vecs = vals[order], vecs[:, order]
theta = np.degrees(np.arctan2(*vecs[:, 0][::-1]))
width, height = 2 * nstd * np.sqrt(vals)
e = Ellipse(xy=mean, width=width, height=height, angle=theta, \
edgecolor=color, fill=False, linewidth=linewidth)
return e
for i in range(num_components):
plt.gca().add_artist(get_gaussian_ellipse_artist(means[i], covs, color=colors[i]))
# ### Generating Kernel weights
# Just to help us visualize let's use two gaussian kernels ([GaussianKernel](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1GaussianKernel.html)) with considerably different widths. As required in MKL, we need to append them to the Combined kernel. To generate the optimal weights (i.e $\beta$s in the above equation), training of [MKL](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MKLClassification.html) is required. This generates the weights as seen in this example.
# +
width0=0.5
kernel0=sg.kernel("GaussianKernel", log_width=np.log(width0))
width1=25
kernel1=sg.kernel("GaussianKernel", log_width=np.log(width1))
#combine kernels
kernel.append_kernel(kernel0)
kernel.append_kernel(kernel1)
kernel.init(feats_train, feats_train)
mkl = sg.MKLClassification()
#set the norm, weights sum to 1.
mkl.set_mkl_norm(1)
mkl.set_C(1, 1)
mkl.set_kernel(kernel)
mkl.set_labels(labels)
#train to get weights
mkl.train()
w=kernel.get_subkernel_weights()
print(w)
# -
# ### Binary classification using MKL
# Now with the data ready and training done, we can do the binary classification. The weights generated can be intuitively understood. We will see that on plotting individual subkernels outputs and outputs of the MKL classification. To apply on test features, we need to reinitialize the kernel with `kernel.init` and pass the test features. After that it's just a matter of doing `mkl.apply` to generate outputs.
# +
size=100
x1=np.linspace(-5, 5, size)
x2=np.linspace(-5, 5, size)
x, y=np.meshgrid(x1, x2)
#Generate X-Y grid test data
grid=sg.features(np.array((np.ravel(x), np.ravel(y))))
kernel0t=sg.kernel("GaussianKernel", log_width=np.log(width0))
kernel1t=sg.kernel("GaussianKernel", log_width=np.log(width1))
kernelt=sg.CombinedKernel()
kernelt.append_kernel(kernel0t)
kernelt.append_kernel(kernel1t)
#initailize with test grid
kernelt.init(feats_train, grid)
mkl.set_kernel(kernelt)
#prediction
grid_out=mkl.apply()
z=grid_out.get_values().reshape((size, size))
plt.figure(figsize=(10,5))
plt.title("Classification using MKL")
c=plt.pcolor(x, y, z)
_=plt.contour(x, y, z, linewidths=1, colors='black')
_=plt.colorbar(c)
# -
# To justify the weights, let's train and compare two subkernels with the MKL classification output. Training MKL classifier with a single kernel appended to a combined kernel makes no sense and is just like normal single kernel based classification, but let's do it for comparison.
# +
z=grid_out.get_labels().reshape((size, size))
# MKL
plt.figure(figsize=(20,5))
plt.subplot(131, title="Multiple Kernels combined")
c=plt.pcolor(x, y, z)
_=plt.contour(x, y, z, linewidths=1, colors='black')
_=plt.colorbar(c)
comb_ker0=sg.CombinedKernel()
comb_ker0.append_kernel(kernel0)
comb_ker0.init(feats_train, feats_train)
mkl.set_kernel(comb_ker0)
mkl.train()
comb_ker0t=sg.CombinedKernel()
comb_ker0t.append_kernel(kernel0)
comb_ker0t.init(feats_train, grid)
mkl.set_kernel(comb_ker0t)
out0=mkl.apply()
# subkernel 1
z=out0.get_labels().reshape((size, size))
plt.subplot(132, title="Kernel 1")
c=plt.pcolor(x, y, z)
_=plt.contour(x, y, z, linewidths=1, colors='black')
_=plt.colorbar(c)
comb_ker1=sg.CombinedKernel()
comb_ker1.append_kernel(kernel1)
comb_ker1.init(feats_train, feats_train)
mkl.set_kernel(comb_ker1)
mkl.train()
comb_ker1t=sg.CombinedKernel()
comb_ker1t.append_kernel(kernel1)
comb_ker1t.init(feats_train, grid)
mkl.set_kernel(comb_ker1t)
out1=mkl.apply()
# subkernel 2
z=out1.get_labels().reshape((size, size))
plt.subplot(133, title="kernel 2")
c=plt.pcolor(x, y, z)
_=plt.contour(x, y, z, linewidths=1, colors='black')
_=plt.colorbar(c)
# -
# As we can see the multiple kernel output seems just about right. Kernel 1 gives a sort of overfitting output while the kernel 2 seems not so accurate. The kernel weights are hence so adjusted to get a refined output. We can have a look at the errors by these subkernels to have more food for thought. Most of the time, the MKL error is lesser as it incorporates aspects of both kernels. One of them is strict while other is lenient, MKL finds a balance between those.
# +
kernelt.init(feats_train, sg.features(testdata))
mkl.set_kernel(kernelt)
out = mkl.apply()
evaluator = sg.evaluation("ErrorRateMeasure")
print("Test error is %2.2f%% :MKL" % (100*evaluator.evaluate(out,sg.BinaryLabels(testlab))))
comb_ker0t.init(feats_train, sg.features(testdata))
mkl.set_kernel(comb_ker0t)
out = mkl.apply()
evaluator = sg.evaluation("ErrorRateMeasure")
print("Test error is %2.2f%% :Subkernel1"% (100*evaluator.evaluate(out,sg.BinaryLabels(testlab))))
comb_ker1t.init(feats_train, sg.features(testdata))
mkl.set_kernel(comb_ker1t)
out = mkl.apply()
evaluator = sg.evaluation("ErrorRateMeasure")
print("Test error is %2.2f%% :subkernel2" % (100*evaluator.evaluate(out,sg.BinaryLabels(testlab))))
# -
# ### MKL for knowledge discovery
# MKL can recover information about the problem at hand. Let us see this with a binary classification problem. The task is to separate two concentric classes shaped like circles. By varying the distance between the boundary of the circles we can control the separability of the problem. Starting with an almost non-separable scenario, the data quickly becomes separable as the distance between the circles increases.
# +
def circle(x, radius, neg):
y=np.sqrt(np.square(radius)-np.square(x))
if neg:
return[x, -y]
else:
return [x,y]
def get_circle(radius):
neg=False
range0=np.linspace(-radius,radius,100)
pos_a=np.array([circle(i, radius, neg) for i in range0]).T
neg=True
neg_a=np.array([circle(i, radius, neg) for i in range0]).T
c=np.concatenate((neg_a,pos_a), axis=1)
return c
def get_data(r1, r2):
c1=get_circle(r1)
c2=get_circle(r2)
c=np.concatenate((c1, c2), axis=1)
feats_tr=sg.features(c)
return c, feats_tr
l=np.concatenate((-np.ones(200),np.ones(200)))
lab=sg.BinaryLabels(l)
#get two circles with radius 2 and 4
c, feats_tr=get_data(2,4)
c1, feats_tr1=get_data(2,3)
_=plt.gray()
plt.figure(figsize=(10,5))
plt.subplot(121)
plt.title("Circles with different separation")
p=plt.scatter(c[0,:], c[1,:], c=lab.get_labels())
plt.subplot(122)
q=plt.scatter(c1[0,:], c1[1,:], c=lab.get_labels())
# -
# These are the type of circles we want to distinguish between. We can try classification with a constant separation between the circles first.
# +
def train_mkl(circles, feats_tr):
#Four kernels with different widths
kernel0=sg.kernel("GaussianKernel", log_width=np.log(1))
kernel1=sg.kernel("GaussianKernel", log_width=np.log(5))
kernel2=sg.kernel("GaussianKernel", log_width=np.log(7))
kernel3=sg.kernel("GaussianKernel", log_width=np.log(10))
kernel = sg.CombinedKernel()
kernel.append_kernel(kernel0)
kernel.append_kernel(kernel1)
kernel.append_kernel(kernel2)
kernel.append_kernel(kernel3)
kernel.init(feats_tr, feats_tr)
mkl = sg.MKLClassification()
mkl.set_mkl_norm(1)
mkl.set_C(1, 1)
mkl.set_kernel(kernel)
mkl.set_labels(lab)
mkl.train()
w=kernel.get_subkernel_weights()
return w, mkl
def test_mkl(mkl, grid):
kernel0t=sg.kernel("GaussianKernel", log_width=np.log(1))
kernel1t=sg.kernel("GaussianKernel", log_width=np.log(5))
kernel2t=sg.kernel("GaussianKernel", log_width=np.log(7))
kernel3t=sg.kernel("GaussianKernel", log_width=np.log(10))
kernelt = sg.CombinedKernel()
kernelt.append_kernel(kernel0t)
kernelt.append_kernel(kernel1t)
kernelt.append_kernel(kernel2t)
kernelt.append_kernel(kernel3t)
kernelt.init(feats_tr, grid)
mkl.set_kernel(kernelt)
out=mkl.apply()
return out
size=50
x1=np.linspace(-10, 10, size)
x2=np.linspace(-10, 10, size)
x, y=np.meshgrid(x1, x2)
grid=sg.features(np.array((np.ravel(x), np.ravel(y))))
w, mkl=train_mkl(c, feats_tr)
print(w)
out=test_mkl(mkl,grid)
z=out.get_values().reshape((size, size))
plt.figure(figsize=(5,5))
c=plt.pcolor(x, y, z)
_=plt.contour(x, y, z, linewidths=1, colors='black')
plt.title('classification with constant separation')
_=plt.colorbar(c)
# -
# As we can see the MKL classifier classifies them as expected. Now let's vary the separation and see how it affects the weights.The choice of the kernel width of the Gaussian kernel used for classification is expected to depend on the separation distance of the learning problem. An increased distance between the circles will correspond to a larger optimal kernel width. This effect should be visible in the results of the MKL, where we used MKL-SVMs with four kernels with different widths (1,5,7,10).
# +
range1=np.linspace(5.5,7.5,50)
x=np.linspace(1.5,3.5,50)
temp=[]
for i in range1:
#vary separation between circles
c, feats=get_data(4,i)
w, mkl=train_mkl(c, feats)
temp.append(w)
y=np.array([temp[i] for i in range(0,50)]).T
# -
plt.figure(figsize=(20,5))
_=plt.plot(x, y[0,:], color='k', linewidth=2)
_=plt.plot(x, y[1,:], color='r', linewidth=2)
_=plt.plot(x, y[2,:], color='g', linewidth=2)
_=plt.plot(x, y[3,:], color='y', linewidth=2)
plt.title("Comparison between kernel widths and weights")
plt.ylabel("Weight")
plt.xlabel("Distance between circles")
_=plt.legend(["1","5","7","10"])
# In the above plot we see the kernel weightings obtained for the four kernels. Every line shows one weighting. The courses of the kernel weightings reflect the development of the learning problem: as long as the problem is difficult the best separation can be obtained when using the kernel with smallest width. The low width kernel looses importance when the distance between the circle increases and larger kernel widths obtain a larger weight in MKL. Increasing the distance between the circles, kernels with greater widths are used.
# ### Multiclass classification using MKL
# MKL can be used for multiclass classification using the [MKLMulticlass](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MKLMulticlass.html) class. It is based on the GMNPSVM Multiclass SVM. Its termination criterion is set by `set_mkl_epsilon(float64_t eps )` and the maximal number of MKL iterations is set by `set_max_num_mkliters(int32_t maxnum)`. The epsilon termination criterion is the L2 norm between the current MKL weights and their counterpart from the previous iteration. We set it to 0.001 as we want pretty accurate weights.
#
# To see this in action let us compare it to the normal [GMNPSVM](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CGMNPSVM.html) example as in the [KNN notebook](http://www.shogun-toolbox.org/static/notebook/current/KNN.html#Comparison-to-Multiclass-Support-Vector-Machines), just to see how MKL fares in object recognition. We use the [USPS digit recognition dataset](http://www.gaussianprocess.org/gpml/data/).
# +
from scipy.io import loadmat, savemat
from os import path, sep
mat = loadmat(sep.join(['..','..','..','data','multiclass', 'usps.mat']))
Xall = mat['data']
Yall = np.array(mat['label'].squeeze(), dtype=np.double)
# map from 1..10 to 0..9, since shogun
# requires multiclass labels to be
# 0, 1, ..., K-1
Yall = Yall - 1
np.random.seed(0)
subset = np.random.permutation(len(Yall))
#get first 1000 examples
Xtrain = Xall[:, subset[:1000]]
Ytrain = Yall[subset[:1000]]
Nsplit = 2
all_ks = range(1, 21)
print(Xall.shape)
print(Xtrain.shape)
# -
# Let's plot five of the examples to get a feel of the dataset.
# +
def plot_example(dat, lab):
for i in range(5):
ax=plt.subplot(1,5,i+1)
plt.title(int(lab[i]))
ax.imshow(dat[:,i].reshape((16,16)), interpolation='nearest')
ax.set_xticks([])
ax.set_yticks([])
_=plt.figure(figsize=(17,6))
plt.gray()
plot_example(Xtrain, Ytrain)
# -
# We combine a [Gaussian kernel](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1GaussianKernel.html) and a [PolyKernel](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CPolyKernel.html). To test, examples not included in training data are used.
#
# This is just a demonstration but we can see here how MKL is working behind the scene. What we have is two kernels with significantly different properties. The gaussian kernel defines a function space that is a lot larger than that of the linear kernel or the polynomial kernel. The gaussian kernel has a low width, so it will be able to represent more and more complex relationships between the training data. But it requires enough data to train on. The number of training examples here is 1000, which seems a bit less as total examples are 10000. We hope the polynomial kernel can counter this problem, since it will fit the polynomial for you using a lot less data than the squared exponential. The kernel weights are printed below to add some insight.
# +
# MKL training and output
labels = sg.MulticlassLabels(Ytrain)
feats = sg.features(Xtrain)
#get test data from 5500 onwards
Xrem=Xall[:,subset[5500:]]
Yrem=Yall[subset[5500:]]
#test features not used in training
feats_rem = sg.features(Xrem)
labels_rem = sg.MulticlassLabels(Yrem)
kernel = sg.CombinedKernel()
feats_train = sg.CombinedFeatures()
feats_test = sg.CombinedFeatures()
#append gaussian kernel
subkernel = sg.kernel("GaussianKernel", log_width=np.log(15))
feats_train.append_feature_obj(feats)
feats_test.append_feature_obj(feats_rem)
kernel.append_kernel(subkernel)
#append PolyKernel
feats = sg.features(Xtrain)
subkernel = sg.kernel('PolyKernel', degree=10, c=2)
feats_train.append_feature_obj(feats)
feats_test.append_feature_obj(feats_rem)
kernel.append_kernel(subkernel)
kernel.init(feats_train, feats_train)
mkl = sg.MKLMulticlass(1.2, kernel, labels)
mkl.set_epsilon(1e-2)
mkl.set_mkl_epsilon(0.001)
mkl.set_mkl_norm(1)
mkl.train()
#initialize with test features
kernel.init(feats_train, feats_test)
out = mkl.apply()
evaluator = sg.evaluation("MulticlassAccuracy")
accuracy = evaluator.evaluate(out, labels_rem)
print("Accuracy = %2.2f%%" % (100*accuracy))
idx=np.where(out.get_labels() != Yrem)[0]
Xbad=Xrem[:,idx]
Ybad=Yrem[idx]
_=plt.figure(figsize=(17,6))
plt.gray()
plot_example(Xbad, Ybad)
# -
w=kernel.get_subkernel_weights()
print(w)
# +
# Single kernel:PolyKernel
C=1
pk = sg.kernel('PolyKernel', degree=10, c=2)
svm = sg.GMNPSVM(C, pk, labels)
_=svm.train(feats)
out=svm.apply(feats_rem)
evaluator = sg.evaluation("MulticlassAccuracy")
accuracy = evaluator.evaluate(out, labels_rem)
print("Accuracy = %2.2f%%" % (100*accuracy))
idx=np.where(out.get_labels() != Yrem)[0]
Xbad=Xrem[:,idx]
Ybad=Yrem[idx]
_=plt.figure(figsize=(17,6))
plt.gray()
plot_example(Xbad, Ybad)
# +
#Single Kernel:Gaussian kernel
width=15
C=1
gk=sg.kernel("GaussianKernel", log_width=np.log(width))
svm=sg.GMNPSVM(C, gk, labels)
_=svm.train(feats)
out=svm.apply(feats_rem)
evaluator = sg.evaluation("MulticlassAccuracy")
accuracy = evaluator.evaluate(out, labels_rem)
print("Accuracy = %2.2f%%" % (100*accuracy))
idx=np.where(out.get_labels() != Yrem)[0]
Xbad=Xrem[:,idx]
Ybad=Yrem[idx]
_=plt.figure(figsize=(17,6))
plt.gray()
plot_example(Xbad, Ybad)
# -
# The misclassified examples are surely pretty tough to predict. As seen from the accuracy MKL seems to work a shade better in the case. One could try this out with more and different types of kernels too.
# ### One-class classification using MKL
# [One-class classification](http://en.wikipedia.org/wiki/One-class_classification) can be done using MKL in shogun. This is demonstrated in the following simple example using [MKLOneClass](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MKLOneClass.html). We will see how abnormal data is detected. This is also known as novelty detection. Below we generate some toy data and initialize combined kernels and features.
# +
X = -0.3 * np.random.randn(100,2)
traindata = np.r_[X + 2, X - 2].T
X = -0.3 * np.random.randn(20, 2)
testdata = np.r_[X + 2, X - 2].T
trainlab=np.concatenate((np.ones(99),-np.ones(1)))
#convert to shogun features and generate labels for data
feats=sg.features(traindata)
labels=sg.BinaryLabels(trainlab)
# +
xx, yy = np.meshgrid(np.linspace(-5, 5, 500), np.linspace(-5, 5, 500))
grid=sg.features(np.array((np.ravel(xx), np.ravel(yy))))
#test features
feats_t=sg.features(testdata)
x_out=(np.random.uniform(low=-4, high=4, size=(20, 2))).T
feats_out=sg.features(x_out)
kernel=sg.CombinedKernel()
feats_train=sg.CombinedFeatures()
feats_test=sg.CombinedFeatures()
feats_test_out=sg.CombinedFeatures()
feats_grid=sg.CombinedFeatures()
#append gaussian kernel
subkernel=sg.kernel("GaussianKernel", log_width=np.log(8))
feats_train.append_feature_obj(feats)
feats_test.append_feature_obj(feats_t)
feats_test_out.append_feature_obj(feats_out)
feats_grid.append_feature_obj(grid)
kernel.append_kernel(subkernel)
#append PolyKernel
feats = sg.features(traindata)
subkernel = sg.kernel('PolyKernel', degree=10, c=3)
feats_train.append_feature_obj(feats)
feats_test.append_feature_obj(feats_t)
feats_test_out.append_feature_obj(feats_out)
feats_grid.append_feature_obj(grid)
kernel.append_kernel(subkernel)
kernel.init(feats_train, feats_train)
mkl = sg.MKLOneClass()
mkl.set_kernel(kernel)
mkl.set_labels(labels)
mkl.set_interleaved_optimization_enabled(False)
mkl.set_epsilon(1e-2)
mkl.put('mkl_epsilon', 0.1)
mkl.set_mkl_norm(1)
# -
# Now that everything is initialized, let's see MKLOneclass in action by applying it on the test data and on the X-Y grid.
# +
mkl.train()
print("Weights:")
w=kernel.get_subkernel_weights()
print(w)
#initialize with test features
kernel.init(feats_train, feats_test)
normal_out = mkl.apply()
#test on abnormally generated data
kernel.init(feats_train, feats_test_out)
abnormal_out = mkl.apply()
#test on X-Y grid
kernel.init(feats_train, feats_grid)
grid_out=mkl.apply()
z=grid_out.get_values().reshape((500,500))
z_lab=grid_out.get_labels().reshape((500,500))
a=abnormal_out.get_labels()
n=normal_out.get_labels()
#check for normal and abnormal classified data
idx=np.where(normal_out.get_labels() != 1)[0]
abnormal=testdata[:,idx]
idx=np.where(normal_out.get_labels() == 1)[0]
normal=testdata[:,idx]
plt.figure(figsize=(15,6))
pl =plt.subplot(121)
plt.title("One-class classification using MKL")
_=plt.pink()
c=plt.pcolor(xx, yy, z)
_=plt.contour(xx, yy, z_lab, linewidths=1, colors='black')
_=plt.colorbar(c)
p1=pl.scatter(traindata[0, :], traindata[1,:], cmap=plt.gray(), s=100)
p2=pl.scatter(normal[0,:], normal[1,:], c="red", s=100)
p3=pl.scatter(abnormal[0,:], abnormal[1,:], c="blue", s=100)
p4=pl.scatter(x_out[0,:], x_out[1,:], c=a, cmap=plt.jet(), s=100)
_=pl.legend((p1, p2, p3), ["Training samples", "normal samples", "abnormal samples"], loc=2)
plt.subplot(122)
c=plt.pcolor(xx, yy, z)
plt.title("One-class classification output")
_=plt.gray()
_=plt.contour(xx, yy, z, linewidths=1, colors='black')
_=plt.colorbar(c)
# -
# MKL one-class classification will give you a bit more flexibility compared to normal classifications. The kernel weights are expected to be more or less similar here since the training data is not overly complicated or too easy, which means both the gaussian and polynomial kernel will be involved. If you don't know the nature of the training data and lot of features are invoved, you could easily use kernels with much different properties and benefit from their combination.
# ### References:
# [1] <NAME>, <NAME>, <NAME>, and <NAME>. Large Scale Multiple Kernel Learning. Journal of Machine Learning Research, 7:1531-1565, July 2006.
#
# [2]<NAME>, <NAME>, and <NAME>. Multiple kernel learning, conic duality, and
# the SMO algorithm. In <NAME>, editor, Twenty-first international conference on Machine
# learning. ACM, 2004
#
# [3] Kernel Methods for Object Recognition , <NAME>
| doc/ipython-notebooks/classification/MKL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# ## Dataset: Top 100,000 YouTube channels by followers
# +
import pandas as pd
df = pd.read_csv("channels.csv")
df
# -
set(df['category_name'])
Edu=df[df["category_id"]==27]
Edu=Edu.sort_values(["followers"], ascending = False)
Edu=Edu[['category_name','title','description','followers','videos','join_date']]
Edu
# ## Data Cleaning
duplicate = Edu.duplicated()
print('There are', duplicate.sum(), 'duplicated values.')
Edu.drop_duplicates(inplace=True)
Edu=Edu[Edu.title.map(lambda x: x.isascii())] #remove non-english title
# +
#Edu=Edu[Edu.description.map(lambda x: x.isascii())]
#remove non-english description will result in essential lost in dataset
# -
Edu=Edu.dropna(subset=["description"]) #remove those with empty description
len(Edu)
Edu["description"]=Edu["description"].str.lower()
Edu["title"]=Edu["title"].str.lower()
# ## Remove channels for kids education
Edu_kids=Edu[Edu["title"].str.contains("kids")
|Edu["description"].str.contains("kids")
|Edu["description"].str.contains("nursery")
|Edu["description"].str.contains("baby")
|Edu["description"].str.contains("children")
|Edu["description"].str.contains("kindergarten")
|Edu["title"].str.contains("baby")]
len(Edu_kids)
Edu_nokids=pd.concat([Edu_kids, Edu]).drop_duplicates(keep=False)
len(Edu_nokids)
# ## 1. Top 5 Education Channel(remove those for kids)
Edu_nokids1=Edu_nokids.head(5)
Edu_nokids1
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1,2,figsize=(20,3))
ax[0].bar(Edu_nokids1["title"], Edu_nokids1["followers"])
ax[1].bar(Edu_nokids1["title"], Edu_nokids1["videos"])
ax[0].set(ylabel = "Number of followers", title = "Number of followers of Top 5 Education Channel")
ax[1].set(ylabel = "Number of videos",title = "Number of videos of Top 5 Education Channel")
plt.show()
# -
# #### The King of Random (aka TKOR)
# a place where curiosity, creativity, and experimentation meet. We're all about learning how things work, doing cool projects, and sharing our discoveries with you.
#
# #### Crash Course
# produced more than 32 courses on a wide variety of subjects, including organic chemistry, literature, world history, biology, philosophy, theater, ecology, and many more! **Including many courses for investing/taxes/insurance, each about 10 min long. Might be Onomy's competitor.**
#
# #### TED-Ed
# creating lessons worth sharing is an extension of TED’s mission of spreading great ideas. **Cover topics about investing, not the rest. Each video is about 5 min with carton-like pictures.**
#
# #### <NAME>
# a Canadian YouTuber and comedian. He produces top ten lists and "50 Amazing Facts" videos on his main channel.
#
# #### <NAME>
# answers fan-submitted questions in a way that is completely inaccurate (most of the time), but is meant to convince the audience of its validity, or simply for entertainment purpose.
# ## 2. Top 5 Courses Related Channels
courses=Edu_nokids[Edu_nokids['title'].str.contains("course")
|Edu_nokids['description'].str.contains("courses")
|Edu_nokids['description'].str.contains("course")
|Edu_nokids['description'].str.contains("lessons")
|Edu_nokids['description'].str.contains("instruction")]
len(courses)
courses1=courses.head(5)
courses1['title']
# #### Unacademy
# india’s largest free education initiative focuses on Academic teaching. Almost no overlap with Onomy.
#
# #### Learn English with Let's Talk - Free English Lessons
# India platform of learning English for Business, IELTS, and TOFEL. Uploading 10 videos per month, each about 10 min. Their topics have almost no overlap with Onomy.
#
# #### Khan Academy
# Our interactive practice problems, articles, and videos help students succeed in math, biology, chemistry, physics, history, economics, finance, grammar, and many other topics.**Including many courses for investing/taxes/insurance, each about 10 min long. Might be Onomy's competitor.**
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1,figsize=(18,3))
plt.bar(courses1["title"], courses1["followers"])
plt.suptitle("Number of followers of Top 5 Course-related Channel")
plt.show()
# +
fig, ax = plt.subplots(1,figsize=(18,3))
plt.bar(courses1["title"], courses1["videos"])
plt.suptitle("Number of videos of Top 5 Course-related Channel")
plt.show()
# -
# ### Attempts to find relationship between number of followers vs videos
fig, axes = plt.subplots(1,figsize=(8,3))
plt.scatter(courses1["videos"], courses1["followers"])
plt.xlabel('Number of videos')
plt.ylabel('Number of followers')
plt.title('Number of videos vs Number of followers')
Edu_nokids1=Edu_nokids.head(100)
fig, axes = plt.subplots(1,figsize=(8,3))
plt.scatter(Edu_nokids["videos"], Edu_nokids["followers"])
plt.xlabel('Number of videos')
plt.ylabel('Number of followers')
plt.title('Number of videos vs Number of followers')
Edu_nokids.sort_values("videos",ascending=False).head(9)
fig, axes = plt.subplots(1,figsize=(8,3))
plt.scatter(courses["videos"], courses["followers"])
plt.xlabel('Number of videos')
plt.ylabel('Number of followers')
plt.title('Number of videos vs Number of followers')
courses.sort_values("videos",ascending=False).head(9)
# ## 3. Potential Competitors Identified based on "similar content"
#get these 4 channels by previous research about their content stated in description(overlap with Onomy)
BigFish=courses.loc[[216,332,802,2669],:]
BigFish
# +
fig, ax = plt.subplots(1,2,figsize=(20,3))
ax[0].bar(BigFish["title"], BigFish["followers"])
ax[1].bar(BigFish["title"], BigFish["videos"])
ax[0].set(ylabel = "Number of followers", title = "Number of followers of Potential Competitors")
ax[1].set(ylabel = "Number of videos",title = "Number of videos of Potential Competitors")
plt.show()
# -
# Number of videos has no relationship with number of followers. For Khan Academy and MIT Opencourseware, they both provide wide range of courses including academic one, so their video numbers are large. For Crashcourse, it has the least number of videos but with the most followers. Considered that Crashcourse covers most of topics taught in Onomy, it might be insightful to study Crashcourse's strategies in attracting viewers.
# ### Thoughts
# Both CrashCourse and Khan Academy offer similar courses as Onomy. Since they offer all kinds of courses including the academic one, college students(people struggling with adulting concern) may be more familiar with these learning platforms and more likely to click in their videos instead of Onomy.
#
# Questions:
# 1. How does Onomy define their target customers?(Age, educational level, US/Inl) How does it differ from target customers of CrashCourse and Khan Academy?
# 2. What are the comparative advantage of Onomy to attract users to take Onomy courses instead of similar courses above?
# 3. What are current marketing strategies/channels that make people get to know about Onomy?
# ## 4. Filter channel description by keywords
# ### For all Education channels that mention Onomy's topic
Invest=Edu_nokids[Edu_nokids['description'].str.contains("investing")
|Edu_nokids['description'].str.contains("invest")
|Edu_nokids['description'].str.contains("finance")]
len(Invest)
Invest.head(6)
# #### <NAME>
# <NAME> is an investor, partner, consultant, or advisor to over 20 multi-million dollar businesses.
#
# #### <NAME>
# <NAME> is the author of eight business books, thirteen business programs, and is the CEO of seven privately held companies. Forbes calls him one of the top social media business influencers in the world.
#
# #### Finnovationz.com
# a platform designed for learners who are passionate about learning stock market investments, mutual funds, TA, etc. The company's goal is to answer people's questions about the stock market. It is the beginners’ guide to the world of finance, and it brings exclusive videos, news updates, courses, blogs and many other products, aimed at increasing financial awareness among people.
#
# #### <NAME>
# We make simple videos on HOW TO START TRADING IN the stock market, ETF, Penny Stocks, Swing & Day Trading, Cryptocurrency, Bitcoin, Startups, Real Estate, Forex, Binary Option, Affiliate marketing, Digital Marketing, Online Sales, and creative methods to make money online.
healthcare=Edu_nokids[Edu_nokids['description'].str.contains("healthcare")]
len(healthcare)
healthcare.head(3)
# #### Nucleus Medical Media
# Nucleus Medical Media creates visual content for healthcare, education, media, pharma and medical device companies. The animations on this channel are predominantly created for hospital patient education and content marketing.
#
# #### Global Health Media Project
# Global Health Media Project produces and distributes teaching videos for frontline healthcare workers and communities in low-resource settings. Based on international standards of care, the videos provide high-quality, step-by-step visual instructions that are easy to understand and put into action. The videos are professionally filmed on-location in developing world health clinics and voiced over to enable narration in local languages.
#
# #### Be Natural
# BE NATURAL is all about hair care, skin care, DIYs, makeup, beauty, style, fitness, fashion, women health & hygiene, women empowerment, girl power, motivational video, vlogs, lifestyle and videos platform for everyone.
tax=Edu_nokids[Edu_nokids['description'].str.contains("tax")
|Edu_nokids['description'].str.contains("taxes")]
len(tax)
tax.head(3)
# #### <NAME>
# We "Badlani Classes" provides CA / CS / CMA online classes not only for students but also for Tax Professionals.
#
# #### <NAME>
# Welcome to City Commerce Academy Online Classes. Learn Accounts and Taxation in just 45 days even if you don't have any accounts background
Credit_Loan=Edu_nokids[Edu_nokids['description'].str.contains("credit card")
|Edu_nokids['description'].str.contains("loans")
|Edu_nokids['description'].str.contains("loan")
|Edu_nokids['description'].str.contains("mortgages")
|Edu_nokids['description'].str.contains("mortgage")]
len(Credit_Loan)
Credit_Loan.head(3)
# #### VIP Financial Education
# VIP Financial Education is the trusted pilot for people who want to dominate the banks.
# Companies from NASA, to RE/MAX have relied on this curriculum to empower borrowers with strategies that quickly grow credit, unlock massive capital, and rapidly wipe out mortgage and non-mortgage debts.
#
# #### 100 Percent Financed
# My name is <NAME> and I help investors get multi-unit rentals using my predictable and automated cashflow cycle system. Teach people how to increase credit score and several Real Estate strategies.
len(Edu_nokids) #4448
Percentage={'Invest':len(Invest)/len(Edu_nokids)*100,
'Healthcare':len(healthcare)/len(Edu_nokids)*100,
'Taxes':len(tax)/len(Edu_nokids)*100,
'Credit/Loan':len(Credit_Loan)/len(Edu_nokids)*100}
Percentage
fig, ax = plt.subplots(1,figsize=(8,5))
plt.bar(range(len(Percentage)), list(Percentage.values()), align='center')
plt.xticks(range(len(Percentage)), list(Percentage.keys()))
plt.suptitle("Percentage of Onomy's topics among all Education Channels",fontsize=15)
plt.ylabel("Percentage")
plt.show()
# ### Filter keywords in all courses-related channels
Invest1=courses[courses['description'].str.contains("investing")
|courses['description'].str.contains("invest")
|courses['description'].str.contains("finance")]
len(Invest1)
healthcare1=courses[courses['description'].str.contains("healthcare")]
len(healthcare1)
healthcare1.head(2)
# #### Healthcare Triage
# Healthcare Triage is a series about healthcare hosted by Dr. <NAME> who explains healthcare policy, medical research, and answers a lot of other questions you may have about medicine, health, and healthcare.
tax1=courses[courses['description'].str.contains("tax")
|courses['description'].str.contains("taxes")]
len(tax1)
tax
# #### Farhat's Accounting Lectures
# offers a growing number of free accounting lectures and accounting courses that cover college level Accounting courses including Financial Accounting, Managerial Accounting, Intermediate Accounting, Advanced Accounting, Taxation, Auditing, Cost Accounting and CPA prep material.
Credit_Loan1=courses[courses['description'].str.contains("credit card")
|courses['description'].str.contains("loans")
|courses['description'].str.contains("loan")
|courses['description'].str.contains("mortgages")
|courses['description'].str.contains("mortgage")]
len(Credit_Loan1)
len(courses) #451
Percentage1={'Invest':len(Invest1)/len(courses)*100,
'Healthcare':len(healthcare1)/len(courses)*100,
'Taxes':len(tax1)/len(courses)*100,
'Credit/Loan':len(Credit_Loan1)/len(courses)*100}
Percentage1
#bar plot
fig, ax = plt.subplots(1,figsize=(7,5))
plt.bar(range(len(Percentage1)), list(Percentage1.values()), align='center')
plt.xticks(range(len(Percentage1)), list(Percentage1.keys()))
plt.title("Percentage of Onomy's topics among all courses Channels")
plt.ylabel("Percentage")
plt.show()
# ### Try Topic Modeling for all Education Channel
X=pd.DataFrame({'text':Edu_nokids['description']})
X
from sklearn.feature_extraction.text import CountVectorizer
vec=CountVectorizer(max_df=0.15,min_df=0.03, stop_words='english')
counts=vec.fit_transform(X['text'])
counts=counts.toarray()
count_df1=pd.DataFrame(counts,columns=vec.get_feature_names())
count_df1=count_df1.drop(['want','website','twitter',
'facebook','online','things','http',
'https','people'],axis=1)
count_df1
# +
from sklearn.decomposition import NMF
model1=NMF(n_components=6,init="random",random_state=0)
model1.fit(count_df1)
#model1.components_
# -
import numpy as np
def top_words(X, model, component, num_words):
"""
Extract the top words from the specified component
for a topic model trained on data.
X: a term-document matrix, assumed to be a pd.DataFrame
model: a sklearn model with a components_ attribute, e.g. NMF
component: the desired component, specified as an integer.
Must be less than than the total number of components in model
num_words: the number of words to return.
"""
orders = np.argsort(model.components_, axis = 1)
important_words = np.array(X.columns)[orders]
return important_words[component][-num_words:]
topic1=pd.DataFrame({'Topic 1':top_words(count_df1, model1, 0, 6),
'Topic 2':top_words(count_df1, model1, 1, 6),
'Topic 3':top_words(count_df1, model1, 2, 6),
'Topic 4':top_words(count_df1, model1, 3, 6),
'Topic 5':top_words(count_df1, model1, 4, 6),
'Topic 6':top_words(count_df1, model1, 5, 6)})
topic1
Y=pd.DataFrame({'text':courses['description']})
Y
from sklearn.feature_extraction.text import CountVectorizer
vec=CountVectorizer(max_df=0.3,min_df=0.03, stop_words='english')
counts=vec.fit_transform(Y['text'])
counts=counts.toarray()
count_df2=pd.DataFrame(counts,columns=vec.get_feature_names())
count_df2=count_df2.drop(['http','https','video'],axis=1)
count_df2
# +
from sklearn.decomposition import NMF
model2=NMF(n_components=4,init="random",random_state=0)
model2.fit(count_df2)
#model2.components_
# -
topic2=pd.DataFrame({'Topic 1':top_words(count_df2, model2, 0, 4),
'Topic 2':top_words(count_df2, model2, 1, 4),
'Topic 3':top_words(count_df2, model2, 2, 4),
'Topic 4':top_words(count_df2, model2, 3, 4)})
topic2
# ## Try to find category composition of Education channels
Language=Edu_nokids[Edu_nokids['description'].str.contains("language")
|Edu_nokids['description'].str.contains("english")
|Edu_nokids['title'].str.contains("english")]
len(Language)
History=Edu_nokids[Edu_nokids['description'].str.contains("history")
|Edu_nokids['title'].str.contains("history")]
len(History)
Music=Edu_nokids[Edu_nokids['description'].str.contains("music")
|Edu_nokids['title'].str.contains("music")]
len(Music)
Computer=Edu_nokids[Edu_nokids['description'].str.contains("computer")
|Edu_nokids['title'].str.contains("computer")]
len(Computer)
Sci=Edu_nokids[Edu_nokids['description'].str.contains("science")
|Edu_nokids['description'].str.contains("technology")
|Edu_nokids['title'].str.contains("technology")
|Edu_nokids['title'].str.contains("science")]
len(Sci)
Health=Edu_nokids[Edu_nokids['description'].str.contains("health")
|Edu_nokids['title'].str.contains("health")]
len(Health)
game=Edu_nokids[Edu_nokids['description'].str.contains("game")
|Edu_nokids['title'].str.contains("game")]
len(game)
exam=Edu_nokids[Edu_nokids['description'].str.contains("exam")
|Edu_nokids['title'].str.contains("exam")]
len(exam)
uni=Edu_nokids[Edu_nokids['description'].str.contains("university")
|Edu_nokids['title'].str.contains("university")]
len(uni)
len(Edu_nokids) #4441
Percentage2={'Language':len(Language)/len(Edu_nokids)*100,
'Science':len(Sci)/len(Edu_nokids)*100,
'Health':len(Health)/len(Edu_nokids)*100,
'History':len(History)/len(Edu_nokids)*100,
'Music':len(Music)/len(Edu_nokids)*100,
'University':len(uni)/len(Edu_nokids)*100,
'Exam Prep':len(exam)/len(Edu_nokids)*100,
'Game':len(game)/len(Edu_nokids)*100,
'Computer':len(Computer)/len(Edu_nokids)*100,
'Invest':len(Invest)/len(Edu_nokids)*100,
'Healthcare':len(healthcare)/len(Edu_nokids)*100,
'Taxes':len(tax)/len(Edu_nokids)*100,
'Credit/Loan':len(Credit_Loan)/len(Edu_nokids)*100}
Percentage2
#bar plot
color=['black']*9+['orange']*4
fig, ax = plt.subplots(1,figsize=(20,5))
plt.bar(range(len(Percentage2)), list(Percentage2.values()), align='center',color=color)
plt.xticks(range(len(Percentage2)), list(Percentage2.keys()),fontsize=15)
plt.title("Category Composition of All Education Channels",fontsize=20)
plt.ylabel("Percentage",fontsize=15)
plt.show()
total=len(healthcare)+len(Invest)+len(Language)+len(Music)+len(History)+len(Sci)+len(tax)+len(Credit_Loan)+len(Computer)+len(Health)+len(game)+len(exam)+len(uni)
known=pd.concat([Music, History,Sci,Language,Invest,healthcare,tax,Credit_Loan,Computer,Health,game,exam,uni]).drop_duplicates(keep='first')
total-len(known) #number of overlap channels
unknown=pd.concat([known,Edu_nokids]).drop_duplicates(keep=False)
len(unknown) #still have large amount of channels that could not be catigorized by description/title
# ## Draft of previous versions(DON'T READ)
# +
# Pie Chart will not be used
import matplotlib.pyplot as plt
# Pie chart, where the slices will be ordered and plotted counter-clockwise:
labels = 'Loan/Mortgages','insurance', 'tax', 'credit', 'Invest','Other'
sizes = [9,9, 13, 33, 41, 346]
fig1, ax1= plt.subplots(figsize=(9,12))
ax1.pie(sizes, labels=labels, autopct='%1.1f%%',startangle=90,rotatelabels=True)
ax1.axis('equal') # Equal aspect ratio ensures that pie is drawn as a circle.
ax1.set(title="Onomy's topic as percentage of all Youtube courses")
plt.show()
# -
s1 = pd.merge(Invest, insurance, how='inner', on=['title'])
s1['title']
s2 = pd.merge(Invest, tax, how='inner', on=['title'])
s2['title']
s3 = pd.merge(Invest, credit, how='inner', on=['title'])
s3['title']
s4 = pd.merge(Invest, loan_mortgage, how='inner', on=['title'])
s4['title']
s5 = pd.merge(credit, loan_mortgage, how='inner', on=['title'])
s5['title']
s6 = pd.merge(credit, tax, how='inner', on=['title'])
s6['title']
s7 = pd.merge(credit, insurance, how='inner', on=['title'])
s7['title']
s8 = pd.merge(credit, insurance, how='inner', on=['title'])
s8['title']
from matplotlib_venn import venn3
venn3(subsets = (10, 8, 22, 6,9,4,2))
plt.show()
| EDA of Online Edu Market/Onomy x Youtube EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import matplotlib.pyplot as plt
import csv
import pickle
import math
# Don't edit
done_load=0
load_dest=""
# +
import time
def deleteDB(db='ycsb', host='vmtest3.westus.cloudapp.azure.com:27017', mongo_dir=r"C:\Program Files\MongoDB\Server\3.6\bin"):
curr_dir=os.getcwd()
os.chdir(mongo_dir)
status = os.system(r'mongo ycsb --host "' + host + '" --eval "db.usertable.drop()"')
os.chdir(curr_dir)
return status
def deleteDBMongo():
deleteDB(host='mongotcoa.westus.cloudapp.azure.com:27017')
def deleteDBAtlas(mongo_dir=r"C:\Program Files\MongoDB\Server\3.6\bin"):
curr_dir=os.getcwd()
os.chdir(mongo_dir)
u=r"anfeldma"
p=r"O!curmt0"
host=r"mongodb+srv://atlas36shard1ncaluswest.fr0to.mongodb.net/ycsb"
run_str=r'mongo "' + host + r'" --username anfeldma --password O!<PASSWORD>' + r' --eval "db.usertable.drop()"'
print(run_str)
status = os.system(run_str)
# create_cmd=r'mongo ycsb --host ' + host + r' -u ' + u + r' -p ' + p + r' --ssl < inp.txt'
# status = os.system(create_cmd)
os.chdir(curr_dir)
time.sleep(2)
def deleteDBCosmos(mongo_dir=r"C:\Program Files\MongoDB\Server\3.6\bin"):
curr_dir=os.getcwd()
os.chdir(mongo_dir)
u=r"mongo-api-benchmark"
p=r"KiYRdcJp41NN268oTcyeM2ilpLwYUAo8tsX9sYoBNTd6DzjXuJHtcaSylh5VJNGs2wg1FVGExRC0m5Z6pEk7ow=="
host=r"mongo-api-benchmark.mongo.cosmos.azure.com:10255"
run_str=r'mongo ycsb --host ' + host + r' -u ' + u + r' -p ' + p + r' --ssl --eval "db.usertable.drop()"'
status = os.system(run_str)
create_cmd=r'mongo ycsb --host ' + host + r' -u ' + u + r' -p ' + p + r' --ssl < inp.txt'
print(create_cmd)
status = os.system(create_cmd)
os.chdir(curr_dir)
time.sleep(2)
return status
# deleteDB(host=r'mongo-api-benchmark:KiYRdcJp41NN268oTcyeM2ilpLwYUAo8tsX9sYoBNTd6DzjXuJHtcaSylh5VJNGs2wg1FVGExRC0m5Z6pEk7ow^=^=<EMAIL>:10255/?ssl^=true^&replicaSet^=globaldb^&retrywrites^=false^&maxIdleTimeMS^=120000^&appName^=@mongo-<EMAIL>-benchmark@')
# deleteDB(host=r'mongo-api-benchmark:KiYRdcJp41NN268oTcyeM2ilpLwYUAo8tsX9sYoBNTd6DzjXuJHtcaSylh5VJNGs2wg1FVGExRC0<EMAIL>==<EMAIL>:10255/?ssl=true&replicaSet=globaldb&retrywrites=false&maxIdleTimeMS=120000&appName=@mongo-api-benchmark@')
# +
def runYCSB(cmd="run", ycsb_dir=r'C:\Users\anfeldma\codeHome\YCSB\bin',workload_dir=r'C:\Users\anfeldma\codeHome\YCSB\workloads',workload='workloadw', \
mongo_endpoint=r'mongodb://vmtest3.westus.cloudapp.azure.com:27017/',operation_count=1000,record_count=100, \
nthreads=1,logdir=".\\",logfn="log.csv"):
curr_dir=os.getcwd()
os.chdir(ycsb_dir)
ycsb_str=r'ycsb ' + cmd + ' mongodb -s -P "' + workload_dir + "\\" + workload + r'" -p mongodb.url="' + mongo_endpoint + \
r'" -p operationcount=' + str(operation_count) + r' -p recordcount=' + str(record_count) + r' -threads ' + str(nthreads) + \
r" " + \
' > ' + logdir + logfn
# r"^&maxPoolSize^=" + str(10*nthreads) + \
print(ycsb_str)
#status=0
os.system(ycsb_str)
os.chdir(curr_dir)
return ycsb_str
def runYCSBMongo36(execmd="run", op_count=10000, rec_count=10000, nthr=1, wkld="workloadw"):
return runYCSB(cmd=execmd, operation_count=op_count, record_count=rec_count, nthreads=nthr, workload=wkld, mongo_endpoint=r"mongodb://mongotcoa.westus.cloudapp.azure.com:27017/")
def runYCSBCosmos36(execmd="run", op_count=10000, rec_count=10000, nthr=1, wkld="workloadw"):
return runYCSB(cmd=execmd, mongo_endpoint=r'mongodb://mongo-api-benchmark:KiYRdcJp41NN268oTcyeM2ilpLwYUAo8tsX9sYoBNTd6DzjXuJHtcaSylh5VJNGs2wg1FVGExRC0m5Z6pEk7ow^=^=<EMAIL>:10255/?ssl^=true^&replicaSet^=globaldb^&retrywrites^=false^&maxIdleTimeMS^=120000^&appName^=@mongo-api-benchmark@', \
operation_count=op_count, record_count=rec_count, nthreads=nthr, workload=wkld)
def runYCSBAtlas36(execmd="run", op_count=10000, rec_count=10000, nthr=1, wkld="workloadw"):
return runYCSB(cmd=execmd, mongo_endpoint=r'mongodb+srv://anfeldma:O%21curmt0@atlas36shard1ncaluswest.fr0to.mongodb.net/ycsb?authSource^=admin^&retryWrites^=true^&w^=majority', \
operation_count=op_count, record_count=rec_count, nthreads=nthr, workload=wkld)
# -
def parseLog(logdir=r'C:\Users\anfeldma\codeHome\YCSB\bin', logfn='log.csv'):
metrics_dict={}
with open(logdir + '\\' + logfn, newline='') as csvfile:
csvrdr = csv.reader(csvfile)#csv.reader(csvfile, delimiter='', quotechar='|')
for row in csvrdr:
if len(row) > 0 and row[0][0] == "[":
arg0 = row[0].lstrip().rstrip()
arg1 = row[1].lstrip().rstrip()
met_val = row[2].lstrip().rstrip()
if not(arg0 in metrics_dict):
metrics_dict[arg0] = {}
metrics_dict[arg0][arg1] = float(met_val)
return metrics_dict
def getIndividualMetrics(met_thrpt_dict_array):
# Plot response curve
thrpt_list=[]
metric_list=[]
max_thrpt=0
for idx in range(len(met_thrpt_dict_array)):
thrpt_list.append(met_thrpt_dict_array[idx][rt_thrpt_field][thrpt_field])
metric_list.append(met_thrpt_dict_array[idx][optype_field][metric_field])
return thrpt_list, metric_list, max_thrpt
def plotResponseCurve(thrpt_list, metric_list, max_thrpt, optype_field):
plt.plot(thrpt_list, metric_list, marker="x")
ax = plt.gca()
for idx in range(len(met_thrpt_dict_array)):
ax.annotate(str(thrpt_list[idx]),
xy=(thrpt_list[idx], metric_list[idx]))
plt.grid(True)
plt.title(optype_field)
plt.xlabel(thrpt_field)
plt.ylabel(metric_field)
fig=plt.gcf()
plt.show()
return fig
def saveResult(met_thrpt_dict_array,thrpt_list,metric_list,nthread_list,max_thrpt,optype_field,ycsb_str,fig):
print("Making " + optype_field + " dir.")
os.makedirs(optype_field, exist_ok=True)
print("Saving result data...")
dumpObj={}
with open(optype_field + "\\pickle.obj", "wb") as fileObj:
dumpObj["met_thrpt_dict_array"]=met_thrpt_dict_array
dumpObj["thrpt_list"]=thrpt_list
dumpObj["metric_list"]=metric_list
dumpObj["nthread_list"]=nthread_list
dumpObj["max_thrpt"]=max_thrpt
dumpObj["optype_field"]=optype_field
dumpObj["ycsb_str"]=max_thrpt
pickle.dump(dumpObj,fileObj)
print("Saving plot...")
fig.savefig(optype_field + "\\" + optype_field + ".png")
def saveComparison(op_max_rate):
print("Making " + "ycsb_op_comparison" + " dir.")
os.makedirs("ycsb_op_comparison", exist_ok=True)
print("Saving comparison data...")
dumpObj={}
with open(optype_field + "\\pickle.obj", "wb") as fileObj:
dumpObj["op_max_rate"]=op_max_rate
pickle.dump(dumpObj,fileObj)
# +
op_mapping={"insert":{"optype_field":"[INSERT]","workload_name":"workloadw"}, \
"read":{"optype_field":"[READ]","workload_name":"workloadr"}, \
"update":{"optype_field":"[UPDATE]","workload_name":"workloadu"} \
}
db_type="atlas" #"cosmos", "mongo", "atlas"
rt_thrpt_field="[OVERALL]"
rt_field="RunTime(ms)"
thrpt_field="Throughput(ops/sec)"
ops_list=["read","update"] #["insert","read","update"]
opname=""
optype_field=""
workload_name=""
metric_field="99thPercentileLatency(us)"
doc_count=10000000#4000000
nthread_list=[50]#range(65,73,1)#[20,50,64,100] #[10,12,14,16,18,20] # [1,2,5,10,20,50,64,100]
# -
print(str(range(65,73,1)[-1]))
# +
met_thrpt_dict_array = []
os.chdir(r"C:\Users\anfeldma\codeHome\YCSB")
op_max_rate={}
for jdx in range(len(ops_list)):
opname = ops_list[jdx]
optype_field=op_mapping[opname]["optype_field"]
workload_name=op_mapping[opname]["workload_name"]
if opname != "insert":
if (done_load>=doc_count and load_dest==db_type):
print("Already loaded data.")
else:
print("Deleting existing data.")
if db_type=="mongo":
deleteDBMongo()
print("Starting YCSB load using max thread count...")
runYCSBMongo36(execmd="load",op_count=doc_count, rec_count=doc_count, nthr=max(nthread_list), wkld=workload_name)
elif db_type=="atlas":
deleteDBAtlas()
print("Starting YCSB load using max thread count...")
runYCSBAtlas36(execmd="load",op_count=doc_count, rec_count=doc_count, nthr=max(nthread_list), wkld=workload_name)
elif db_type=="cosmos":
deleteDBCosmos()
print("Starting YCSB load using max thread count...")
runYCSBCosmos36(execmd="load",op_count=doc_count, rec_count=doc_count, nthr=max(nthread_list), wkld=workload_name)
done_load=doc_count
load_dest=db_type
print("Finished YCSB load.")
for idx in range(len(nthread_list)):
print("Starting YCSB " + db_type + " run, opname " + opname + ", workload " + workload_name + ", thread count " + str(nthread_list[idx]))
if opname=="insert":
if db_type=="mongo":
deleteDBMongo()
elif db_type=="atlas":
deleteDBAtlas()
elif db_type=="cosmos":
deleteDBCosmos()
print("Done deleting existing YCSB dataset.")
done_load=0
operation_count=doc_count
if opname=="read" or opname=="update":
print(opname)
operation_count=int(doc_count/3)
elif opname=="insert":
print(opname)
operation_count=int(doc_count/3)
if db_type=="mongo":
ycsb_str=runYCSBMongo36(op_count=operation_count, rec_count=doc_count, nthr=nthread_list[idx], wkld=workload_name)
elif db_type=="atlas":
ycsb_str=runYCSBAtlas36(op_count=operation_count, rec_count=doc_count, nthr=nthread_list[idx], wkld=workload_name)
elif db_type=="cosmos":
ycsb_str=runYCSBCosmos36(op_count=operation_count, rec_count=doc_count, nthr=nthread_list[idx], wkld=workload_name)
met_thrpt_dict_array.append(parseLog())
print("Finished YCSB run, thread count " + str(nthread_list[idx]))
thrpt_list, metric_list, max_thrpt = getIndividualMetrics(met_thrpt_dict_array)
max_thrpt=max(thrpt_list)
met_thrpt_dict_array=[]
fig=plotResponseCurve(thrpt_list, metric_list, max_thrpt, opname)
saveResult(met_thrpt_dict_array,thrpt_list,metric_list,nthread_list,max_thrpt,optype_field,ycsb_str,fig)
print("Max throughput: " + str(max_thrpt))
op_max_rate[opname]=max_thrpt
saveComparison(op_max_rate)
print(op_max_rate)
# -
met_thrpt_dict_array
os.getcwd()
| ResponseCurveAutomation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/helenpiepie/hl.github.io/blob/master/Food_Recommendation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="wwHjj7Br-sUk" colab_type="text"
# # Food Recommendation: Clustering Models
# + [markdown] id="yY9gKLFD-03J" colab_type="text"
# Image we are a company who designs food menu for **Hyperglycemic patients**. One of our services is to help patients prevent or **reduce the intake of high-carbohydrate and high-fat food**. In this notebook, 45027 kinds of food are listed and clustered as either 'healthy' or 'unhealthy' food.
# + [markdown] id="Y8a6auMZTzoa" colab_type="text"
# ## Ingest
# + id="co7HfeLmvvZV" colab_type="code" colab={}
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
import matplotlib.cbook
warnings.filterwarnings("ignore",category=matplotlib.cbook.mplDeprecation)
sns.set(style="white", palette="muted", color_codes=True)
# + id="77Ih-vxcv-BR" colab_type="code" outputId="fc0d0b2d-8cca-4f75-a438-b7713e97f7c2" colab={"base_uri": "https://localhost:8080/", "height": 377}
df = pd.read_csv(
"https://raw.githubusercontent.com/noahgift/food/master/data/features.en.openfoodfacts.org.products.csv")
df.drop(["Unnamed: 0", "exceeded", "g_sum", "energy_100g"], axis=1, inplace=True) #drop two rows we don't need
df = df.drop(df.index[[1,11877]]) #drop outlier
df.rename(index=str, columns={"reconstructed_energy": "energy_100g"}, inplace=True)
df.head()
# + [markdown] id="P_AMOqmFY7nm" colab_type="text"
# ## EDA
# + id="Bga4PDs8PJ63" colab_type="code" outputId="b5c8fbe0-eb1c-4dc1-b35a-1d70f1127a20" colab={"base_uri": "https://localhost:8080/", "height": 68}
df.columns
# + id="pCpP5sT0wK4u" colab_type="code" outputId="cd9381ac-2cbf-4753-ebea-a8aa84170e3d" colab={"base_uri": "https://localhost:8080/", "height": 34}
df.shape
# + id="in5EsQyLMfvk" colab_type="code" outputId="6f5ebe09-d709-4c6f-f328-2fa9875db9aa" colab={"base_uri": "https://localhost:8080/", "height": 221}
df.info()
# + [markdown] id="bq_K-nnPWQcA" colab_type="text"
# ### Sort by Sugar, Carbohydrate, Fat
# + id="iI2ogrIvKYIm" colab_type="code" outputId="d7b185de-430c-465f-bf74-41605b326d5a" colab={"base_uri": "https://localhost:8080/", "height": 889}
df.sort_values(by=["sugars_100g","carbohydrates_100g","fat_100g"], ascending=[True, True, True]).head(10)
# + [markdown] id="XWnJKyYXbHBU" colab_type="text"
# ### Histogram
# + [markdown] id="V8Pi_OEGbJv1" colab_type="text"
# Generate distributions based on fat, sugar, and carbohydrates
# + id="8LykCes1bMwP" colab_type="code" outputId="92d0a981-6f05-4c96-8baf-45071f5ce29d" colab={"base_uri": "https://localhost:8080/", "height": 623}
# Set up the matplotlib figure
f, axes = plt.subplots(2, 2, figsize=(10, 10), sharex=True)
sns.despine(left=True)
# Plot a simple histogram with binsize determined automatically
sns.distplot(df.fat_100g, color="b", ax=axes[0, 0])
sns.distplot(df.sugars_100g, color="g", ax=axes[0, 1])
sns.distplot(df.carbohydrates_100g, color="m", ax=axes[1, 0])
# + [markdown] id="pxh-Xls2a_kc" colab_type="text"
# ### Word Cloud
# + id="TS_vESIbaTMH" colab_type="code" colab={}
from wordcloud import WordCloud, STOPWORDS
import matplotlib.pyplot as plt
# + [markdown] colab_type="text" id="LKKQX0f0iW5a"
# #### High fat and sugar foods
# + [markdown] colab_type="text" id="R_9G_7JziW5c"
# Find fatty and sweet foods in the 99th percentile
# + id="YuN7ragAOATd" colab_type="code" colab={}
high_fat_df = df[df.fat_100g > df.fat_100g.quantile(.99)]
high_sugar_df = df[df.sugars_100g > df.sugars_100g.quantile(.99)]
high_carbohydrates_df = df[df.carbohydrates_100g > df.carbohydrates_100g.quantile(.99)]
high_fat_and_sugar_df = high_fat_df.append(pd.DataFrame(data = high_sugar_df), ignore_index=True)
high_fat_and_sugar_df = high_fat_df.append(pd.DataFrame(data = high_carbohydrates_df), ignore_index=True)
# + id="Ie786XHuOqXq" colab_type="code" outputId="c2be1dfc-05aa-4057-c300-361606393350" colab={"base_uri": "https://localhost:8080/", "height": 34}
high_fat_and_sugar_text = high_fat_and_sugar_df['product'].values
len(high_fat_and_sugar_text)
# + [markdown] colab_type="text" id="Y_0zg1itiW5f"
# Word Cloud High Fat and sugar
# + colab_type="code" outputId="7f5e0140-0ffd-480f-907f-ff36725fdc86" id="6B6Ql3bXiW5g" colab={"base_uri": "https://localhost:8080/", "height": 627}
wordcloud = WordCloud(
width = 3000,
height = 2000,
background_color = 'black',
stopwords = STOPWORDS).generate(str(high_fat_and_sugar_text))
fig = plt.figure(
figsize = (12, 8),
facecolor = 'k',
edgecolor = 'k')
plt.imshow(wordcloud, interpolation = 'bilinear')
plt.axis('off')
plt.tight_layout(pad=0)
plt.show()
# + [markdown] id="9cvboB89IzkW" colab_type="text"
# ## Modeling
# + [markdown] id="qXTQIDrhP4Aw" colab_type="text"
# ### Create Features to Cluster
# + id="zxGmh-6sI2oE" colab_type="code" outputId="2648188b-149f-4741-f5c2-d9602a8f63b6" colab={"base_uri": "https://localhost:8080/", "height": 68}
df.columns
# + [markdown] id="NnZGyDKvPztU" colab_type="text"
# We will cluster based on fat, carbohydrates, and sugars.
# + id="4P18IBIPI-JD" colab_type="code" colab={}
df_cluster_features = df.drop(['proteins_100g',
'salt_100g', 'energy_100g','product'], axis=1)
# + id="nPXkU5o1Q1TU" colab_type="code" outputId="e7e1ed9b-5c53-4e24-9f9b-d8b0be8f01e7" colab={"base_uri": "https://localhost:8080/", "height": 297}
df_cluster_features.describe()
# + [markdown] id="0ikpFFC3K8eS" colab_type="text"
# ### Scale the data
# + id="xthd6KraJHkP" colab_type="code" outputId="d450eb21-ad75-4583-8c64-d1282e089878" colab={"base_uri": "https://localhost:8080/", "height": 153}
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
print(scaler.fit(df_cluster_features))
print(scaler.transform(df_cluster_features))
# + [markdown] id="IS9657wxQdKV" colab_type="text"
# ### Determine Cluster Number
# + [markdown] id="5d2si5l2SexW" colab_type="text"
# #### Yellowbrick Visualizer Elbow Method
# + id="JnHNAvG_SiUY" colab_type="code" outputId="ea3b82c4-6973-4caf-9f60-04293ef711b6" colab={"base_uri": "https://localhost:8080/", "height": 376}
from sklearn.cluster import KMeans
from yellowbrick.cluster import KElbowVisualizer
# Instantiate the clustering model and visualizer
model = KMeans()
visualizer = KElbowVisualizer(model, k=(1,11))
visualizer.fit(df_cluster_features) # Fit the data to the visualizer
visualizer.poof() # Draw/show/poof the data
# + [markdown] id="vURiLsllHYK9" colab_type="text"
# **Two clusters seem ideal (healthy and unhealthy food).**
# + [markdown] id="YvcpnRK2TQQ5" colab_type="text"
# #### Yellowbrick Silhouette Visualizer
#
# + id="K-DcBLiOTZrr" colab_type="code" outputId="40138e2e-94b9-4783-f1ac-ae003fb79114" colab={"base_uri": "https://localhost:8080/", "height": 376}
from sklearn.cluster import MiniBatchKMeans
from yellowbrick.cluster import SilhouetteVisualizer
# Instantiate the clustering model and visualizer
model = MiniBatchKMeans(2)
visualizer = SilhouetteVisualizer(model)
visualizer.fit(df_cluster_features) # Fit the training data to the visualizer
visualizer.poof() # Draw/show/poof the data
# + [markdown] id="b7zfwUzjGx1t" colab_type="text"
# ### Clustering
# + id="mOOGRYjKQxqW" colab_type="code" colab={}
new_df = df.drop(['proteins_100g','salt_100g', 'energy_100g'], axis=1)
# + id="I7yEPaPQaT-w" colab_type="code" outputId="c05c12a0-805c-46ea-e1b2-0bb9d6166dca" colab={"base_uri": "https://localhost:8080/", "height": 364}
X = new_df.iloc[:, [0, 1, 2]].values
y_kmeans = kmeans.fit_predict(X)
plt.scatter(X[y_kmeans == 0, 0], X[y_kmeans == 0, 1], s = 100, c = 'lightgreen', label = 'cluster_A')
plt.scatter(X[y_kmeans == 1, 0], X[y_kmeans == 1, 1], s = 100, c = 'lightblue', label = 'cluster_B')
#Plotting the centroids of the clusters
plt.scatter(kmeans.cluster_centers_[:, 0], kmeans.cluster_centers_[:,1], s = 100, c = 'yellow', label = 'Centroide')
plt.legend()
# + id="pA7irconcxst" colab_type="code" colab={}
from sklearn.cluster import KMeans
k_means = KMeans(n_clusters=2)
kmeans = k_means.fit(scaler.transform(df_cluster_features))
# + id="SK6JK4qZG0ol" colab_type="code" outputId="338fd78a-0220-4031-b977-31cbb64072bc" colab={"base_uri": "https://localhost:8080/", "height": 410}
new_df['cluster'] = kmeans.labels_
new_df.head(10)
# + [markdown] id="ayBs1YuESRkr" colab_type="text"
# **Examine what does cluster 1 and clsuter 0 represent**
# + id="CvyXzFCySGzT" colab_type="code" outputId="0c576bbb-d67e-4ae0-aa89-00289bb5aaf9" colab={"base_uri": "https://localhost:8080/", "height": 204}
new_df.sort_values(by=["sugars_100g","carbohydrates_100g","fat_100g"], ascending=[True, True, True]).head()
# + id="loMY7vq-SrKP" colab_type="code" outputId="c117291b-6a51-4d86-a480-d768f2fa6365" colab={"base_uri": "https://localhost:8080/", "height": 204}
new_df.sort_values(by=["sugars_100g","carbohydrates_100g","fat_100g"], ascending=[False, False, False]).head()
# + [markdown] id="1WD4NGdEVlTk" colab_type="text"
# ## Conclusion
# + [markdown] id="MKPj5s5xVoIt" colab_type="text"
#
#
# * What we want to achieve from the dataset determines how to choose appropriate features. This notebook is to target Hyperglycemic patients and to help them prevent or reduce the intake of high-carbohydrate and high-fat food. Therefore, choosing fat, carbohydrates, and sugars as the clustering features is more reasonable.
# * Using the clustering results, Hyperglycemic patients can look in the clustered data frame and have a basic idea of whether the chosen food is good for them or not.
#
#
| Food_Recommendation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We say an operator $S$
# $$
# A: \mathcal P^+ \to \mathcal P^+
# $$
#
# - is $\alpha$-scalable for some $\alpha\in \mathbb R$ if
# $$
# A(\theta f) = \theta^\alpha T f, \quad \theta \geq 0, f\in \mathcal P^+.
# $$
#
# - is monotonic if for each $f, g\in \mathcal P^+$ with $f\leq g$, we have
# $$
# Af \leq Ag
# $$
# We say an operator $A: D\subset \mathcal P^* \to \mathcal P^*$ is $\alpha$-controlled for some $\alpha \in \mathbb R$ if there is $M$ a monotonic operator and #S# an $\alpha$-scalable operator such that
# $$
# |Af|\leq M|f| \leq S|f|,\quad f\in D
# $$
# Let $A$ and $A_0$ be an $\alpha$-controllable and an $\alpha_0$-controllable operator, repectively.
# Denote by $M$ and $R$ the monotonic operator and the $\alpha$-scalable operator corresponding to $A$.
# Denote by $M_0$ and $R_0$ similar for $A_0$.
# Then
# $$
# |AA_0 f|\leq M |A_0f| \leq M M_0 |f|
# \leq MR_0 f = M|R_0 f|\leq RR_0f
# $$
# $$
# T_k^\alpha
# $$
# is 1-scalable with
# $$
# |T_t^\alpha g| \leq e^{-\kappa(g)bt} R_g
# $$
# Then
# $$
# |AT_t^\alpha g| \leq M|T_t^\alpha g| \leq M(e^{-\kappa(g)bt}Qg) \leq R e^{-\kappa(g)bt}Qg
# $$
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="r3cas2_1T98w"
# # Decision Tree Regression
# + [markdown] colab_type="text" id="IODliia6U1xO"
# ## Importing the libraries
# + colab={} colab_type="code" id="y98nA5UdU6Hf"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] colab_type="text" id="jpjZ43YlU8eI"
# ## Importing the dataset
# + colab={} colab_type="code" id="pLVaXoYVU_Uy"
dataset = pd.read_csv('Position_Salaries.csv')
X = dataset.iloc[:, 1:-1].values
y = dataset.iloc[:, -1].values
# + [markdown] colab_type="text" id="g16qFkFQVC35"
# ## Training the Decision Tree Regression model on the whole dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 121} colab_type="code" id="SLDKyv1SVUqS" outputId="a633ebbf-6fea-4b97-ccd8-1f8851e9d363"
from sklearn.tree import DecisionTreeRegressor
regressor = DecisionTreeRegressor(random_state = 0)
regressor.fit(X, y)
# + [markdown] colab_type="text" id="MQRGPTH3VcOn"
# ## Predicting a new result
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="_FpGZf7vVgrK" outputId="54f36048-d4a1-4143-8b2b-b5aa32233b68"
regressor.predict([[6.5]])
# + [markdown] colab_type="text" id="ph8ExBj0VkIT"
# ## Visualising the Decision Tree Regression results (higher resolution)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="zzH1Vv1oVrqe" outputId="84111519-5c51-498c-c330-0d53825849e3"
X_grid = np.arange(min(X), max(X), 0.01)
X_grid = X_grid.reshape((len(X_grid), 1))
plt.scatter(X, y, color = 'red')
plt.plot(X_grid, regressor.predict(X_grid), color = 'blue')
plt.title('Truth or Bluff (Decision Tree Regression)')
plt.xlabel('Position level')
plt.ylabel('Salary')
plt.show()
| Regression Models/Decision Trees/decision_tree_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="jfWMKrNz13IB"
# # Qiita記事のタイトルを自動生成するモデルの学習
#
# 事前学習済み日本語T5モデルを、今回の自動生成タスク用に転移学習(ファインチューニング)します。
#
# - **T5(Text-to-Text Transfer Transformer)**: テキストを入力されるとテキストを出力するという統一的枠組みで様々な自然言語処理タスクを解く深層学習モデル([解説](https://www.ogis-ri.co.jp/otc/hiroba/technical/similar-document-search/part7.html))
# - **事前学習**: 個別のタスク用に学習をする前に文法や一般的な言葉の意味を学習させること(教師なし学習や自己教師あり学習とWikipedia等の大規模データ(コーパス)を用いることで広く一般的な知識を持ったモデルを作れる)
# - **転移学習、ファインチューニング**: 事前学習済みモデルを初期値にして、特定のタスク用に追加で学習を行うこと(主に教師あり学習)
#
#
# 今回は入出力が次の形式を持ったタスク用に転移学習します。
# - **入力**: "body: {body}"をトークナイズしたトークンID列
# - **出力**: "title: {title}"をトークナイズしたトークンID列
#
# ここで、{body}は記事本文(を前処理したもの)、{title}はタイトルです。
#
# ### 補足
#
# 今回はタイトル自動生成タスク用のモデルを作りますが、データセット(TSVファイル)を他のタスク用のものに入れ替えれば、ほぼコードを変えることなく(最大入出力トークン数を指定するmax_input_lengthとmax_target_lengthは適切な値に変える必要があるでしょう)、他のタスク用のモデルを作ることもできます。ぜひ色々試してみてください。
# + [markdown] id="-_0AfdGtSqup"
# ## ライブラリの準備
# + [markdown] id="lf9IXZuZ2J-3"
# ### パスワード設定
#
# **※学習に用いるデータセットとモデルの解凍にはパスワードが必要です(一般には公開しません)。**
# **※次の変数 RESOURCES_PASSWORD に解凍用の秘密のパスワードを設定してから、以降の処理を実行してください。**
# + id="UxrYLtEz2ISx"
# データセットとモデルの解凍用パスワード
RESOURCES_PASSWORD = ""
# + [markdown] id="VXwcsE_8xtFL"
# ### 依存ライブラリのインストール
# + id="5JQI6sIAAR2T" colab={"base_uri": "https://localhost:8080/"} outputId="788f3634-c5ed-41b6-f4b9-ac018e6f1275"
# !pip install -qU torch==1.7.1 torchtext==0.8.0 torchvision==0.8.2
# !pip install -q transformers==4.2.2 pytorch_lightning==1.2.1 sentencepiece
# + [markdown] id="ZkIGrSKTxwa6"
# ### 学習に利用するデータセットと事前学習済みモデルをダウンロード
# + colab={"base_uri": "https://localhost:8080/"} id="oFD6mIaPs1h8" outputId="94d4cc80-1939-448f-8711-5a9240ebcdfc"
# !wget -O resources.tar "https://www.floydhub.com/api/v1/resources/idKgfo9obBzdvzEXcXFJxj?content=true&download=true"
# !tar xvf resources.tar
# + [markdown] id="UjsS8yqdx3va"
# ### データと事前学習済みモデルの解凍
# + colab={"base_uri": "https://localhost:8080/"} id="XJseFTsGxD60" outputId="704ff791-3086-46e4-ab33-b28ea8c4bcb9"
# !unzip -P {RESOURCES_PASSWORD} qiita_title_generation.zip
# + [markdown] id="6JbdTnSqzvpy"
# ### 各種ディレクトリの作成
#
# - data: 学習データの置き場所
# - model: 学習済みモデルの出力先
# + id="XKBBoT1OBo3G"
# !mkdir -p /content/data /content/model
# + id="EanRC6WDOTu5"
OUTPUT_MODEL_DIR = "/content/model"
# + [markdown] id="7_Z2A5YQ-TaY"
# ※/content/modelディレクトリは、このColabインスタンスがリセットされたら削除されます。
# 削除されない場所に保存したいのであれば、Google Driveに保存するのが簡単です。
#
# ColabからGoogle Driveを利用する方法: https://qiita.com/kado_u/items/45b76f9a6f920bf0f786
# + id="mXTIcz3-DDHB"
# もし、Google Driveに学習済みモデルを保存する場合は次の処理を実行
# ※Googleアカウントの認証が必要です。
# from google.colab import drive
# drive.mount('/content/drive')
# OUTPUT_MODEL_DIR = "/content/drive/MyDrive/qiita_title_generation_model"
# # !mkdir -p {OUTPUT_MODEL_DIR}
# + [markdown] id="kaI8M-lsDvI0"
# ## 学習可能な形式にデータを変換
# + [markdown] id="Bn7zUi7BEHaB"
# ### 前処理の定義
# + [markdown] id="Ef1vVbNJFW7x"
# #### 文字列の正規化
#
# 表記揺れを減らします。今回は[neologdの正規化処理](https://github.com/neologd/mecab-ipadic-neologd/wiki/Regexp.ja)を一部改変したものを利用します。
# 処理の詳細はリンク先を参照してください。
# + id="7TU3L01WgKmE"
# https://github.com/neologd/mecab-ipadic-neologd/wiki/Regexp.ja から引用・一部改変
from __future__ import unicode_literals
import re
import unicodedata
def unicode_normalize(cls, s):
pt = re.compile('([{}]+)'.format(cls))
def norm(c):
return unicodedata.normalize('NFKC', c) if pt.match(c) else c
s = ''.join(norm(x) for x in re.split(pt, s))
s = re.sub('-', '-', s)
return s
def remove_extra_spaces(s):
s = re.sub('[ ]+', ' ', s)
blocks = ''.join(('\u4E00-\u9FFF', # CJK UNIFIED IDEOGRAPHS
'\u3040-\u309F', # HIRAGANA
'\u30A0-\u30FF', # KATAKANA
'\u3000-\u303F', # CJK SYMBOLS AND PUNCTUATION
'\uFF00-\uFFEF' # HALFWIDTH AND FULLWIDTH FORMS
))
basic_latin = '\u0000-\u007F'
def remove_space_between(cls1, cls2, s):
p = re.compile('([{}]) ([{}])'.format(cls1, cls2))
while p.search(s):
s = p.sub(r'\1\2', s)
return s
s = remove_space_between(blocks, blocks, s)
s = remove_space_between(blocks, basic_latin, s)
s = remove_space_between(basic_latin, blocks, s)
return s
def normalize_neologd(s):
s = s.strip()
s = unicode_normalize('0-9A-Za-z。-゚', s)
def maketrans(f, t):
return {ord(x): ord(y) for x, y in zip(f, t)}
s = re.sub('[˗֊‐‑‒–⁃⁻₋−]+', '-', s) # normalize hyphens
s = re.sub('[﹣-ー—―─━ー]+', 'ー', s) # normalize choonpus
s = re.sub('[~∼∾〜〰~]+', '〜', s) # normalize tildes (modified by <NAME>)
s = s.translate(
maketrans('!"#$%&\'()*+,-./:;<=>?@[¥]^_`{|}~。、・「」',
'!”#$%&’()*+,-./:;<=>?@[¥]^_`{|}〜。、・「」'))
s = remove_extra_spaces(s)
s = unicode_normalize('!”#$%&’()*+,-./:;<>?@[¥]^_`{|}〜', s) # keep =,・,「,」
s = re.sub('[’]', '\'', s)
s = re.sub('[”]', '"', s)
return s
# + [markdown] id="QT_O4Pf6FZu_"
# #### Markdownのクリーニング
#
# タイトルを考える上で関係のなさそうな文章を削る処理を行います。
# 以下のノイズとなるデータを削除し、タブや改行を空白文字にしたり、文字を小文字に揃える等の処理を行います。
#
# - ソースコード
# - URLやリンク
# - 画像
#
# 現状、img以外のHTML要素を残していますが、タイトルに関係なさそうな要素を削ると精度が上がるかもしれません。
# + id="kYaC9tcbGwJD"
import re
CODE_PATTERN = re.compile(r"```.*?```", re.MULTILINE | re.DOTALL)
LINK_PATTERN = re.compile(r"!?\[([^\]\)]+)\]\([^\)]+\)")
IMG_PATTERN = re.compile(r"<img[^>]*>")
URL_PATTERN = re.compile(r"(http|ftp)s?://[^\s]+")
NEWLINES_PATTERN = re.compile(r"(\s*\n\s*)+")
def clean_markdown(markdown_text):
markdown_text = CODE_PATTERN.sub(r"", markdown_text)
markdown_text = LINK_PATTERN.sub(r"\1", markdown_text)
markdown_text = IMG_PATTERN.sub(r"", markdown_text)
markdown_text = URL_PATTERN.sub(r"", markdown_text)
markdown_text = NEWLINES_PATTERN.sub(r"\n", markdown_text)
markdown_text = markdown_text.replace("`", "")
return markdown_text
def normalize_text(markdown_text):
markdown_text = clean_markdown(markdown_text)
markdown_text = markdown_text.replace("\t", " ")
markdown_text = normalize_neologd(markdown_text).lower()
markdown_text = markdown_text.replace("\n", " ")
return markdown_text
def preprocess_qiita_body(markdown_text):
return "body: " + normalize_text(markdown_text)[:4000]
def preprocess_qiita_title(markdown_text):
# return normalize_text(markdown_text)
return "title: " + normalize_text(markdown_text)
# + [markdown] id="GeuZcreUFhWe"
# ### Qiita記事データ(JSON形式)を学習データ(TSV形式)に変換
# + colab={"base_uri": "https://localhost:8080/"} id="M1WCYH7GFhCH" outputId="80d6fc08-3735-4927-f44c-59516c0c47e6"
import json
import gzip
import random
import math
# Qiita記事データ(JSON形式)を読み込む
with gzip.open("/content/qiita_title_generation/qiita/qiita_articles.json.gz",
"rt", encoding="utf8") as f_in:
dataset = json.load(f_in)
# 前処理を施す
for data in dataset:
body = data["body"]
title = data["title"]
data["preprocessed_body"] = preprocess_qiita_body(body)
data["preprocessed_title"] = preprocess_qiita_title(title)
# 再現性のため、ランダムシードを固定してデータをシャッフルする
random.seed(42)
random.shuffle(dataset)
# 学習データを訓練データ/開発(バリデーション)データ/テスト(評価)データに分割する。
total_count = len(dataset)
train_count = math.ceil(total_count * 0.92)
dev_count = math.ceil(total_count * 0.04)
test_count = total_count - train_count - dev_count
with open("/content/data/train.tsv", "w", encoding="utf8") as f_train, \
open("/content/data/dev.tsv", "w", encoding="utf8") as f_dev, \
open("/content/data/test.tsv", "w", encoding="utf8") as f_test:
for i, data in enumerate(dataset):
preprocessed_body = data["preprocessed_body"]
preprocessed_title = data["preprocessed_title"]
if i < train_count:
f_out = f_train
elif i < train_count + dev_count:
f_out = f_dev
else:
f_out = f_test
f_out.write(f"{preprocessed_body}\t{preprocessed_title}\n")
print(train_count, dev_count, test_count)
# + [markdown] id="ePouLoxLzKgG"
# ## 学習に必要なクラス等の定義
#
# 学習にはPyTorch/PyTorch-lightning/Transformersを利用します。
# + id="15ZzooLKA-j5"
import argparse
import os
import random
import numpy as np
import torch
from torch.utils.data import Dataset, DataLoader
import pytorch_lightning as pl
from transformers import T5ForConditionalGeneration, T5Tokenizer
from transformers import AdamW,get_linear_schedule_with_warmup
# 乱数シードの設定
def set_seed(seed):
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
if torch.cuda.is_available():
torch.cuda.manual_seed_all(seed)
set_seed(12345)
# + id="v1EyFckzBV_Y"
# 事前学習済みモデルの置き場所
PRETRAINED_MODEL_PATH = "/content/qiita_title_generation/pretrained_model"
# GPU利用有無
USE_GPU = torch.cuda.is_available()
# 各種ハイパーパラメータ
args_dict = dict(
data_dir="/content/data", # データセットのディレクトリ
output_dir=OUTPUT_MODEL_DIR, # 学習済みモデルの出力先
model_name_or_path=PRETRAINED_MODEL_PATH,
tokenizer_name_or_path=PRETRAINED_MODEL_PATH,
learning_rate=1e-3, # 学習率
adam_epsilon=1e-8,
weight_decay=0.0,
warmup_steps=0,
gradient_accumulation_steps=1,
# max_input_length=512,
# max_target_length=64,
# train_batch_size=8,
# eval_batch_size=8,
# num_train_epochs=4,
n_gpu=1 if USE_GPU else 0,
early_stop_callback=False,
fp_16=False,
opt_level='O1',
max_grad_norm=1.0,
seed=12345,
)
# + [markdown] id="Hv6jWWCe1Iaf"
# ### TSVデータセットクラス
#
# TSV形式のファイルをデータセットとして読み込みます。
# 形式は"{入力される文字列}\t{出力される文字列}"です。
# + id="uEgMB3pjD6q_"
class TsvDataset(Dataset):
def __init__(self, tokenizer, data_dir, type_path, input_max_len=512, target_max_len=512):
self.file_path = os.path.join(data_dir, type_path)
self.input_max_len = input_max_len
self.target_max_len = target_max_len
self.tokenizer = tokenizer
self.inputs = []
self.targets = []
self._build()
def __len__(self):
return len(self.inputs)
def __getitem__(self, index):
source_ids = self.inputs[index]["input_ids"].squeeze()
target_ids = self.targets[index]["input_ids"].squeeze()
source_mask = self.inputs[index]["attention_mask"].squeeze()
target_mask = self.targets[index]["attention_mask"].squeeze()
return {"source_ids": source_ids, "source_mask": source_mask,
"target_ids": target_ids, "target_mask": target_mask}
def _build(self):
# ファイルからデータを読み込む
with open(self.file_path, "r", encoding="utf-8") as f:
for line in f:
line = line.strip().split("\t")
assert len(line) == 2
assert len(line[0]) > 0
assert len(line[1]) > 0
inputs = [line[0]]
targets = [line[1]]
tokenized_inputs = self.tokenizer.batch_encode_plus(
inputs, max_length=self.input_max_len, truncation=True,
padding="max_length", return_tensors="pt"
)
tokenized_targets = self.tokenizer.batch_encode_plus(
targets, max_length=self.target_max_len, truncation=True,
padding="max_length", return_tensors="pt"
)
self.inputs.append(tokenized_inputs)
self.targets.append(tokenized_targets)
# + [markdown] id="yOtEZl6t3z-a"
# 試しにテストデータ(test.tsv)を読み込み、トークナイズ結果をみてみます。
# + id="LWrvaBiIHtza"
# トークナイザー(SentencePiece)モデルの読み込み
tokenizer = T5Tokenizer.from_pretrained(PRETRAINED_MODEL_PATH, is_fast=True)
# テストデータセットの読み込み
train_dataset = TsvDataset(tokenizer, args_dict["data_dir"],
"test.tsv",
input_max_len=512, target_max_len=64)
# + [markdown] id="g9Ds_y4K4TdP"
# テストデータの1レコード目をみてみます。
# + id="iKKjKUi4IUdI" colab={"base_uri": "https://localhost:8080/"} outputId="dc5bb6c2-9f6c-45c1-c2e3-c6830a79a4b6"
for data in train_dataset:
print("A. 入力データの元になる文字列")
print(tokenizer.decode(data["source_ids"]))
print()
print("B. 入力データ(Aの文字列がトークナイズされたトークンID列)")
print(data["source_ids"])
print()
print("C. 出力データの元になる文字列")
print(tokenizer.decode(data["target_ids"]))
print()
print("D. 出力データ(Cの文字列がトークナイズされたトークンID列)")
print(data["target_ids"])
break
# + [markdown] id="cMy5OZ7q5r1J"
# ### 学習処理クラス
#
# [PyTorch-Lightning](https://github.com/PyTorchLightning/pytorch-lightning)を使って学習します。
#
# PyTorch-Lightningとは、機械学習の典型的な処理を簡潔に書くことができるフレームワークです。
# + id="9VTRkMzrw_Ec"
import os
class T5FineTuner(pl.LightningModule):
def __init__(self, hparams):
super().__init__()
self.hparams = hparams
# 事前学習済みモデルの読み込み
self.model = T5ForConditionalGeneration.from_pretrained(hparams.model_name_or_path)
# トークナイザーの読み込み
self.tokenizer = T5Tokenizer.from_pretrained(hparams.tokenizer_name_or_path, is_fast=True)
def forward(self, input_ids, attention_mask=None, decoder_input_ids=None,
decoder_attention_mask=None, labels=None):
"""順伝搬"""
return self.model(
input_ids,
attention_mask=attention_mask,
decoder_input_ids=decoder_input_ids,
decoder_attention_mask=decoder_attention_mask,
labels=labels
)
def _step(self, batch):
"""ロス計算"""
labels = batch["target_ids"]
labels[labels[:, :] == self.tokenizer.pad_token_id] = -100
outputs = self(
input_ids=batch["source_ids"],
attention_mask=batch["source_mask"],
decoder_attention_mask=batch["target_mask"],
labels=labels
)
loss = outputs[0]
return loss
def training_step(self, batch, batch_idx):
"""訓練ステップ処理"""
loss = self._step(batch)
self.log("train_loss", loss)
return {"loss": loss}
def _precision_recall_f1_step(self, batch):
"""トークンレベルの一致率(precision/recall/f1)を求める"""
# ※文章の生成の場合、BLEUやROUGEの方がより良い場合もある。
input_ids = batch["source_ids"]
input_mask = batch["source_mask"]
targets = batch["target_ids"]
outputs = self.model.generate(
input_ids=input_ids,
attention_mask=input_mask,
max_length=args.max_target_length,
repetition_penalty=1.5,
)
outputs = [set(output.cpu().tolist()) for output in outputs]
targets = [set(target.cpu().tolist()) for target in targets]
precisions = [len(o & t) / len(o) for o, t in zip(outputs, targets)]
recalls = [len(o & t) / len(t) for o, t in zip(outputs, targets)]
f1s = [2 * p * r / (p + r) if p + r > 0 else 0 for p, r in zip(precisions, recalls)]
p = precisions
r = recalls
f1 = f1s
return p, r, f1
def validation_step(self, batch, batch_idx):
"""バリデーションステップ処理"""
loss = self._step(batch)
p, r, f1 = self._precision_recall_f1_step(batch)
self.log("val_loss", loss)
return {"val_loss": loss, "val_p": p, "val_r": r, "val_f1": f1}
def validation_epoch_end(self, outputs):
"""バリデーション完了処理"""
avg_loss = torch.stack([x["val_loss"] for x in outputs]).mean()
avg_p = np.mean([v for x in outputs for v in x["val_p"]])
avg_r = np.mean([v for x in outputs for v in x["val_r"]])
avg_f1 = np.mean([v for x in outputs for v in x["val_f1"]])
self.log("val_loss", avg_loss, prog_bar=True)
self.log("val_p", avg_p, prog_bar=True)
self.log("val_r", avg_r, prog_bar=True)
self.log("val_f1", avg_f1, prog_bar=True)
return {"val_loss": avg_loss, "val_p": avg_p, "val_r": avg_r, "val_f1": avg_f1}
def test_step(self, batch, batch_idx):
"""テストステップ処理"""
loss = self._step(batch)
p, r, f1 = self._precision_recall_f1_step(batch)
self.log("test_loss")
return {"test_loss": loss, "test_p": p, "test_r": r, "test_f1": f1}
def test_epoch_end(self, outputs):
"""テスト完了処理"""
avg_loss = torch.stack([x["test_loss"] for x in outputs]).mean()
avg_p = torch.stack([x["test_p"] for x in outputs]).mean()
avg_r = torch.stack([x["test_r"] for x in outputs]).mean()
avg_f1 = torch.stack([x["test_f1"] for x in outputs]).mean()
self.log("test_loss", avg_loss, prog_bar=True)
self.log("test_p", avg_p, prog_bar=True)
self.log("test_r", avg_r, prog_bar=True)
self.log("test_f1", avg_f1, prog_bar=True)
def configure_optimizers(self):
"""オプティマイザーとスケジューラーを作成する"""
model = self.model
no_decay = ["bias", "LayerNorm.weight"]
optimizer_grouped_parameters = [
{
"params": [p for n, p in model.named_parameters()
if not any(nd in n for nd in no_decay)],
"weight_decay": self.hparams.weight_decay,
},
{
"params": [p for n, p in model.named_parameters()
if any(nd in n for nd in no_decay)],
"weight_decay": 0.0,
},
]
optimizer = AdamW(optimizer_grouped_parameters,
lr=self.hparams.learning_rate,
eps=self.hparams.adam_epsilon)
# scheduler = get_linear_schedule_with_warmup(
# optimizer, num_warmup_steps=self.hparams.warmup_steps, num_training_steps=self.t_total
# )
# return [optimizer], [{"scheduler": scheduler, "interval": "step", "frequency": 1}]
return [optimizer]
def get_dataset(self, tokenizer, type_path, args):
"""データセットを作成する"""
return TsvDataset(
tokenizer=tokenizer,
data_dir=args.data_dir,
type_path=type_path,
input_max_len=args.max_input_length,
target_max_len=args.max_target_length)
def setup(self, stage=None):
"""初期設定(データセットの読み込み)"""
if stage == "fit" or stage is None:
train_dataset = self.get_dataset(tokenizer=self.tokenizer,
type_path="train.tsv", args=self.hparams)
self.train_dataset = train_dataset
val_dataset = self.get_dataset(tokenizer=self.tokenizer,
type_path="dev.tsv", args=self.hparams)
self.val_dataset = val_dataset
# self.t_total = (
# (len(train_dataset) // (self.hparams.train_batch_size * max(1, self.hparams.n_gpu)))
# // self.hparams.gradient_accumulation_steps
# * float(self.hparams.num_train_epochs)
# )
def train_dataloader(self):
"""訓練データローダーを作成する"""
return DataLoader(self.train_dataset, batch_size=self.hparams.train_batch_size, drop_last=True, shuffle=True, num_workers=4)
def val_dataloader(self):
"""バリデーションデータローダーを作成する"""
return DataLoader(self.val_dataset, batch_size=self.hparams.eval_batch_size, num_workers=4)
# + [markdown] id="abeNz6_TjtnI"
# Transformersのモデルのみを保存するようにする
#
# + id="OBBuGb_DjrTl"
import os
class TransformerTrainer(pl.Trainer):
def save_checkpoint(self, filepath, weights_only=False):
"""モデルの重みを保存する"""
# 学習済みモデル(とトークナイザー)を保存する
if self.is_global_zero:
print(f"save model: {filepath}")
model_dir_name = os.path.dirname(filepath)
lightningmodel = self.get_model()
lightningmodel.model.save_pretrained(model_dir_name)
lightningmodel.tokenizer.save_pretrained(model_dir_name)
# + [markdown] id="MGh2UwbL95fQ"
# ## 転移学習を実行
# + id="e80VBR4bIhnM"
# 学習に用いるハイパーパラメータを設定する
args_dict.update({
"max_input_length": 512, # 入力文の最大トークン数 (<512)
"max_target_length": 64, # 出力文の最大トークン数 (<512)
"train_batch_size": 8,
"eval_batch_size": 8,
"num_train_epochs": 4, # 適切なエポック数にしてください。2~10くらいのはず。
})
args = argparse.Namespace(**args_dict)
checkpoint_callback = pl.callbacks.ModelCheckpoint(
args.output_dir,
# バリデーションセット(dev.tsv)におけるF1値が最大のモデルを1個だけ保存します。
monitor="val_f1", mode="max", save_top_k=1
)
train_params = dict(
gpus=args.n_gpu,
max_epochs=args.num_train_epochs,
precision= 16 if args.fp_16 else 32,
amp_level=args.opt_level,
gradient_clip_val=args.max_grad_norm,
accumulate_grad_batches=args.gradient_accumulation_steps,
checkpoint_callback=checkpoint_callback,
)
# + id="0Ps_lWvCL0tW" colab={"base_uri": "https://localhost:8080/", "height": 390, "referenced_widgets": ["83eb97e631b84468b306dcadddddc936", "e117b46b64264f63890710d66e5b8d0c", "b60eef2d46e04e4a9050cb525aaf4ce2", "4d017b3e6ad74436b37b48e0baf32553", "ff7736971bb14508a0af8fa00d8235d1", "<KEY>", "<KEY>", "<KEY>", "51268dd4ed6641dabf0c13391e4bbe0f", "<KEY>", "f683fff6a7484442aebabdc7872e8504", "<KEY>", "07035b5f1f3c4c549407eaf06ee0fd5e", "8286d9c14e7c4d2986adae1b73e52030", "<KEY>", "6875c1755f334a2ea935096e33ab1e2a", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "846cda8406b84d4d9f8e70003559a68d", "<KEY>", "ddf5fb4d4dd54cc58669ade874f875d4", "08a510c4269b4be19e9763a47d54660c", "<KEY>", "22bec67ef23143aba3e234e54e2e59f6", "7098c4aaaad24554a26f563f2020e319", "f6b72c4753084e8d9db65323edaf1b02", "<KEY>", "40f5f071275641ada3a5a46c6e38e439", "<KEY>", "f5ad9cca566b42dfb04ea3dbd1cb1ab6", "<KEY>", "672b7d77fd2e4fe4993a1ca5cb59c588", "885b28a092c348efb774a070cec7a972", "<KEY>", "27ae7acb9212441ba1be8adefaf64bba", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "d2336b5e4d5c4a9187fa63912c305964", "<KEY>", "<KEY>", "<KEY>"]} outputId="f2f32a46-6c83-4327-c32f-c81a3f705e79"
# 転移学習の実行(GPUを利用すれば1エポック10分程度)
model = T5FineTuner(args)
trainer = TransformerTrainer(**train_params)
trainer.fit(model)
# + [markdown] id="g76MuSLIFdaB"
# ## 学習済みモデルの読み込み
# + id="YHvq7JNCQtvD"
import torch
from torch.utils.data import Dataset, DataLoader
from transformers import T5ForConditionalGeneration, T5Tokenizer
# トークナイザー(SentencePiece)
tokenizer = T5Tokenizer.from_pretrained(OUTPUT_MODEL_DIR, is_fast=True)
# 学習済みモデル
trained_model = T5ForConditionalGeneration.from_pretrained(OUTPUT_MODEL_DIR)
# GPUの利用有無
USE_GPU = torch.cuda.is_available()
if USE_GPU:
trained_model.cuda()
# + [markdown] id="Bv8If9WYF4I7"
# ## 全テストデータに対してタイトル生成を実行
# + id="H7qyYdvrEm-9" colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["7f22fe83f13641ea9df55e1e8d2059ee", "d4a55dc8780f4d2580d170b73744a881", "<KEY>", "<KEY>", "5d9202de1ba1437f97e02ae808a3fd75", "788860e8d5d0447cac5abeeedc264b7d", "9de6bbf345364a1e8d6ab462ba183f0f", "6bdb5529886f4d828e829517ca69e845"]} outputId="19799dd1-376a-47d4-8012-34bc67a2c18f"
import textwrap
from tqdm.auto import tqdm
from sklearn import metrics
# テストデータの読み込み
test_dataset = TsvDataset(tokenizer, args_dict["data_dir"],
"test.tsv",
input_max_len=args.max_input_length,
target_max_len=args.max_target_length)
test_loader = DataLoader(test_dataset, batch_size=4, num_workers=4)
# 推論モード
trained_model.eval()
outputs = []
targets = []
for batch in tqdm(test_loader):
input_ids = batch['source_ids']
input_mask = batch['source_mask']
if USE_GPU:
input_ids = input_ids.cuda()
input_mask = input_mask.cuda()
outs = trained_model.generate(input_ids=input_ids,
attention_mask=input_mask,
max_length=args.max_target_length,
repetition_penalty=1.5,
)
dec = [tokenizer.decode(ids, skip_special_tokens=True,
clean_up_tokenization_spaces=False)
for ids in outs]
target = [tokenizer.decode(ids, skip_special_tokens=True,
clean_up_tokenization_spaces=False)
for ids in batch["target_ids"]]
output_idset = [set(ids.cpu()) for ids in outs]
output_idset = [set(ids) for ids in outs]
outputs.extend(dec)
targets.extend(target)
# + id="XQlUn1MEgnv7"
def postprocess_title(title):
return re.sub(r"^title: ", "", title)
# + [markdown] id="bIqmdKiwTM4W"
# generatedが自動生成されたタイトル、actualが実際に人が付けたタイトルです。
# + id="ammyb4W9KoqD" colab={"base_uri": "https://localhost:8080/"} outputId="7cc1135d-0b0e-463f-e514-60d9e6597ca3"
import re
for output, target in list(zip(outputs, targets)):
print(f"generated: {postprocess_title(output)}")
print(f"actual: {postprocess_title(target)}")
print()
# + [markdown] id="GY-FprEDGuIq"
# ## 以降は次のNotebookと同じ内容のため説明は省略
#
# https://github.com/sonoisa/qiita-title-generation/blob/main/T5_ja_qiita_title_generation.ipynb
# + id="jr-SYFb9G2u7"
qiita_body = """
AIの進歩はすごいですね。
今回は深層学習を用いて、記事(Qiita)のタイトルを自動生成してくれるAIさんを試作してみました。
この実験は自然言語処理について新人さんに教えるためのハンズオンネタを探索する一環で行ったものになります。
作ったAIは、Qiitaの記事本文(少し前処理したテキスト)を与えると、適したタイトル文字列を作文して返してくれるというものです。
なお、学習データは(2019年頃に)Qiitaの殿堂を入り口にして、評価の高い記事(いいねが50個以上)をスクレイピングしたものを使いました。
つまりヒットした記事のタイトルの付け方を学んだAIであるといえます。
* もう少し詳細:
* 学習データの例:
* 入力: "body: hiveqlではスピードに難を感じていたため、私もprestoを使い始めました。 mysqlやhiveで使っていたクエリ..."
* 出力: "title: hadoop利用者ならきっと知ってる、hive/prestoクエリ関数の挙動の違い"
* 学習方法: 独自に作った日本語T5の事前学習モデルをこの学習データを用いて転移学習
以下、結果(抜粋)です。generatedが生成されたもの、actualが人が付けたタイトルです。
"""
# + colab={"base_uri": "https://localhost:8080/", "height": 139} id="op1BR0S9G3Uz" outputId="582c4b06-fdf2-4364-ff43-6e79235bd1d4"
preprocess_qiita_body(qiita_body)
# + colab={"base_uri": "https://localhost:8080/"} id="av0NevpMG6fc" outputId="f6f7a1fd-098e-4144-d206-138c292bac76"
MAX_SOURCE_LENGTH = 512 # 入力される記事本文の最大トークン数
MAX_TARGET_LENGTH = 64 # 生成されるタイトルの最大トークン数
# 推論モード設定
trained_model.eval()
# 前処理とトークナイズを行う
inputs = [preprocess_qiita_body(qiita_body)]
batch = tokenizer.batch_encode_plus(
inputs, max_length=MAX_SOURCE_LENGTH, truncation=True,
padding="longest", return_tensors="pt")
input_ids = batch['input_ids']
input_mask = batch['attention_mask']
if USE_GPU:
input_ids = input_ids.cuda()
input_mask = input_mask.cuda()
# 生成処理を行う
outputs = trained_model.generate(
input_ids=input_ids, attention_mask=input_mask,
max_length=MAX_TARGET_LENGTH,
temperature=1.0, # 生成にランダム性を入れる温度パラメータ
num_beams=10, # ビームサーチの探索幅
diversity_penalty=1.0, # 生成結果の多様性を生み出すためのペナルティ
num_beam_groups=10, # ビームサーチのグループ数
num_return_sequences=10, # 生成する文の数
repetition_penalty=1.5, # 同じ文の繰り返し(モード崩壊)へのペナルティ
)
# 生成されたトークン列を文字列に変換する
generated_titles = [tokenizer.decode(ids, skip_special_tokens=True,
clean_up_tokenization_spaces=False)
for ids in outputs]
# 生成されたタイトルを表示する
for i, title in enumerate(generated_titles):
print(f"{i+1:2}. {postprocess_title(title)}")
# + id="flxsJkTZI8NG"
| T5_ja_training_qiita_title_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ¿Cómo medir rendimiento y riesgo en un portafolio? II
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="http://www.picpedia.org/clipboard/images/stock-portfolio.jpg" width="600px" height="400px" />
#
# > La clase pasada y la presente, están dedicadas a obtener medidas de rendimiento y riesgo en un portafolio.
#
# > Vimos que podemos obtener los rendimientos de un portafolio mediante la relación $r_p=\sum_{i=1}^{n}w_ir_i$, y una vez teniendo los rendimientos del portafolio, lo podemos tratar como un activo individual.
#
# > Por otra parte, vimos que si conocemos los rendimientos esperados de cada activo que conforma el portafolio $E[r_i]$, podemos calcular el rendimiento esperado del portafolio como el promedio ponderado de los rendimientos esperados de los activos $E[r_p]=\sum_{i=1}^{n}w_iE[r_i]$.
#
# > Sin embargo, vimos que esto no es válido para la medida de riesgo (desviación estándar). Es decir, la varianza (o volatilidad, o desviación estándar) no es el promedio ponderado de las varianzas individuales. Anticipamos que esto es clave en el concepto de **diversificación**.
# **Objetivos:**
# - Medir el riesgo en un portafolio a partir del riesgo de cada uno de los activos que lo conforman.
#
# *Referencias*
# - Notas del curso "Portfolio Selection and Risk Management", Rice University, disponible en Coursera.
# ___
# ## 1. Midiendo el riesgo en un portafolio
#
# ### 1.1. Volatilidad de un portafolio
#
# Retomamos el ejemplo qur veníamos trabajando la clase pasada...
# **Ejemplo.** Supongamos que tenemos inversión en activos de Toyota, Walmart y Pfizer. Tenemos cuatro posibles estados económicos:
import numpy as np
import pandas as pd
# +
# Creamos tabla
tabla = pd.DataFrame(columns=['Prob', 'Toyota', 'Walmart', 'Pfizer'],
index=['Expansion', 'Normal', 'Recesion', 'Depresion'])
tabla.index.name = 'Estado'
tabla['Prob']=np.array([0.1, 0.4, 0.3, 0.2])
tabla['Toyota']=np.array([0.06, 0.075, 0.02, -0.03])
tabla['Walmart']=np.array([0.045, 0.055, 0.04, -0.01])
tabla['Pfizer']=np.array([0.025, -0.005, 0.01, 0.13])
tabla.round(4)
# -
## Rendimientos esperados
# Toyota
ErT = (tabla['Prob'] * tabla['Toyota']).sum()
# Walmart
ErW = (tabla['Prob'] * tabla['Walmart']).sum()
# Pfizer
ErP = (tabla['Prob'] * tabla['Pfizer']).sum()
# Mostrar
ErT, ErW, ErP
## Volatilidad
# Toyota
sT = (tabla['Prob'] * (tabla['Toyota'] - ErT)**2).sum()**0.5
# Walmart
sW = (tabla['Prob'] * (tabla['Walmart'] - ErW)**2).sum()**0.5
# Pfizer
sP = (tabla['Prob'] * (tabla['Pfizer'] - ErP)**2).sum()**0.5
# Mostrar
sT, sW, sP
# Portafolio 0.5Toyota+0.5Pfizer
tabla['PortTP'] = 0.5 * tabla['Toyota'] + 0.5 * tabla['Pfizer']
tabla
# Rendimiento portafolio (como activo individual)
ErTP = (tabla['Prob'] * tabla['PortTP']).sum()
# Rendimiento portafolio (como suma ponderada de rendimientos individuales)
ErTP1 = 0.5 * ErT + 0.5 * ErP
ErTP, ErTP1
# Volatilidad del portafolio
sTP = (tabla['Prob'] * (tabla['PortTP'] - ErTP)**2).sum()**0.5
sTP
# Notar que sTP < 0.5 * sT + 0.5 * sP
# la volatilidad del portafolio siempre es menor
# a la suma ponderada de las volatilidades individuales
0.5 * sT + 0.5 * sP
# **Actividad.** Encontrar la volatilidad del portafolio formado $0.5$ Toyota y $0.5$ Walmart.
# Encontrar los rendimientos del portafolio en cada estado de la economía
tabla['PortTW'] = 0.5 * tabla['Toyota'] + 0.5 * tabla['Walmart']
tabla
# Encontrar el rendimiento esperado del portafolio
ErTW = (tabla['Prob'] * tabla['PortTW']).sum()
ErTW1 = 0.5 * ErT + 0.5 * ErW
ErTW, ErTW1
# Encontrar la volatilidad de Toyota, Walmart y el portafolio
sTW = (tabla['Prob'] * (tabla['PortTW'] - ErTW)**2).sum()**0.5
sTW
# Notar que sTW < 0.5 * sT + 0.5 * sW
# la volatilidad del portafolio siempre es menor
# a la suma ponderada de las volatilidades individuales
0.5 * sT + 0.5 * sW
# ### 1.2. Midiendo el co-movimiento entre instrumentos
#
# - Una vez más, concluimos que la volatilidad (varianza) **NO** es el promedio ponderado de las varianzas individales.
#
# - Por el contrario, la varianza de los rendimientos de un portafolio está afectada por el movimiento relativo de un activo individual respecto a otro.
#
# - Por tanto, necesitamos definir las medidas de **covarianza** y **correlación**, que nos permiten evaluar las fluctuaciones relativas entre los activos.
# #### Covarianza:
#
# Es una medida el movimiento relativo entre dos instrumentos.
#
# Matemáticamente, si tenemos dos activos $A_1$ y $A_2$ cuyos rendimientos son $r_1$ y $r_2$, respectivamente, entonces la covarianza de los rendimientos de los activos es
#
# $$\text{cov}(r_1,r_2)=\sigma_{12}=\sum_{j=1}^{m}p_j(r_{1j}-E[r_1])(r_{2j}-E[r_2]).$$
#
# $$\text{cov}(r_2,r_1)=\sigma_{21}=\sum_{j=1}^{m}p_j(r_{2j}-E[r_2])(r_{1j}-E[r_1]) = \sigma_{12}.$$
# Podemos notar fácilmente que la covarianza de los rendimientos de un activo con los rendimientos del mismo activo corresponde a la varianza
#
# $$\text{cov}(r_1,r_1)=\sigma_{11}=\sum_{j=1}^{m} p_j(r_{1j}-E[r_1])(r_{1j}-E[r_1])=\sigma_1^2=\text{var}(r_1).$$
# **Ejemplo.** Calcular la covarianza entre los rendimientos de Toyota y Pfizer.
# Mostrar tabla
tabla
tabla[['Toyota', 'Pfizer']]
# Calcular la covarianza
covTP = (tabla['Prob'] * (tabla['Toyota'] - ErT) * (tabla['Pfizer'] - ErP)).sum()
covTP
# **Actividad.** Calcular la covarianza entre los rendimientos de Toyota y Walmart.
# Calcular la covarianza
covTW = (tabla['Prob'] * (tabla['Toyota'] - ErT) * (tabla['Walmart'] - ErW)).sum()
covTW
tabla[['Toyota', 'Walmart']]
# ¿Qué nos dice este número?
# - El signo nos dice las direcciones relativas entre los rendimientos de cada activo. Por ejemplo, la covarianza entre los rendimientos de Toyota y Pfizer es negativa... ver los rendimientos.
# - La magnitud de la covarianza no nos dice mucho acerca de la fuerza con que se relacionan o no estos rendimientos.
# **Correlación:**
#
# Un posible problema de la covarianza es que la magnitud de esta medida no nos dice mucho acerca de la fuerza de los movimientos relativos. La *correlación* es una medida normalizada del movimiento relativo entre los rendimientos de dos activos.
#
# Matemáticamente,
#
# $$\text{corr}(r_1,r_2)=\rho_{12}=\rho_{21}=\frac{\sigma_{12}}{\sigma_1\sigma_{2}}.$$
# Propiedades:
#
# - Podemos notar fácilmente que la correlación de los rendimientos de un activo con los rendimientos del mismo activo es $1$: $$\text{corr}(r_1,r_1)=\rho_{11}=\frac{\sigma_{11}}{\sigma_1\sigma_1}=\frac{\sigma_{1}^2}{\sigma_1\sigma_1}=1.$$
# - El signo de la correlación y la covarianza es el mismo.
# - La correlación satisface: $$-1\leq\rho_{12}\leq 1.$$
# **Ejemplo.** Calcular la correlación entre los rendimientos de Toyota y Pfizer.
corrTP = covTP / (sT * sP)
corrTP
# **Actividad.** Calcular la correlación entre los rendimientos de Toyota y Walmart.
corrTW = covTW / (sT * sW)
corrTW
# **Conclusión.**
# - Es una medida normalizada de la fluctuación relativa de los rendimientos de dos activos.
# - En los ejemplos que vimos, sería conveniente invertir en el portafolio de Toyota y Pfizer puesto que su correlación es negativa, y esto impactaría positivamente en la diversificación del riesgo.
#
# ___
# ## 2. Uniendo todo...
# - Entonces, vimos mediante ejemplos que el riesgo en un portafolio se ve afectado significativamente por como los rendimientos de los activos se mueven relativamente.
# - Este movimiento relativo lo medimos mediante la covarianza o la correlación.
# - Si se mueven de una manera que no están perfectamente correlacionados ($\rho<1$), entonces el riesgo del portafolio siempre será menor que el promedio ponderado de los riesgos individuales.
# <img style="float: left; margin: 0px 0px 15px 15px;" src="https://www.publicdomainpictures.net/pictures/20000/velka/happy-child.jpg" width="300px" height="200px" />
#
# ## Ésta es la razón por la que combinar activos en un portafolio permite diversificar el riesgo...
# Entonces, ¿cómo podemos incorporar esta medida en el cálculo de la varianza del portafolio?
# - <font color=blue> Ver en el tablero...</font>
#
# $$
# \sigma_p^2 = \sum_{i=1}^{n} \sum_{k=1}^{n} w_i w_k \sigma_{ik} = w^T \Sigma w
# $$
#
# - ¿Cómo sería para dos activos?
#
# \begin{align}
# \sigma_p^2 & = w_1^2 \sigma_1^2 + w_2^2 \sigma_2^2 + 2 w_1 w_2 \sigma_{12} \\
# & = w_1^2 \sigma_1^2 + w_2^2 \sigma_2^2 + 2 w_1 w_2 \rho_{12} \sigma_1 \sigma_2
# \end{align}
# **Ejemplo.** Calcular por fórmula para el portafolio de Toyota y Pfizer. Comparar.
sTP2 = 0.5**2 * sT**2 + 0.5**2 * sP**2 + 2 * 0.5**2 * covTP
sTP1 = sTP2**0.5
w = np.array([0.5, 0.5])
Sigma = np.array([[sT**2, covTP],
[covTP, sP**2]])
sTP_ = (w.T.dot(Sigma).dot(w))**0.5
sTP, sTP1, sTP_
# **Actividad.** Calcular por fórmula para el portafolio de Toyota y Walmart. Comparar.
sTW2 = 0.5**2 * sT**2 + 0.5**2 * sW**2 + 2 * 0.5**2 * covTW
sTW1 = sTW2**0.5
sTW, sTW1
# ## 2.1. <font color=blue> Ver en el tablero...</font>
# ### Matriz de varianza covarianza.
# ### Matriz de correlación.
# # Anuncios parroquiales
#
# ## 1. Recordar quiz la siguiente clase. Temas: Clases 6 y 7.
# ## 2. Tarea: revisar archivo "Tarea4_MidiendoRendimientoRiesgo" en clase.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Modulo2/Clase7_RendimientoRiesgoPortafoliosII.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from transformers import AutoModelWithLMHead, AutoTokenizer
tokenizer = AutoTokenizer.from_pretrained("microsoft/DialoGPT-medium")
new_special_tokens_dict = {'additional_special_tokens':['🙂', '☹️', '😮', '😡', 'firewood']}
num_added = tokenizer.add_special_tokens(new_special_tokens_dict)
print(num_added)
tokenizer.tokenize('🙂 firewood ☹️ 😮 😡')
| kc_work/test_tokenizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (jupyter env)
# language: python
# name: jupyter_venv
# ---
# # Notes on Chapter 1 of *Hands-On Machine Learning with Scikit-Learn, Keras, & TensorFlow* by <NAME>
#
# ## Exercises:
# ### 1
# A set of algorithms whose performance improves with exposure to data
#
# ### 2
# Image recognition, speech recognition, turn-based games, natural language processing
#
# ### 3
# A set of sample inputs and matching desired outputs for a prediction task
#
# ### 4
# Regression and classification
#
# ### 5
# Clustering, anomaly detection, dimensional reduction, and association rule learning
#
# ### 6
# Reinforcement learning
#
# ### 7
# Clustering
#
# ### 8
# Supervised learning
#
# ### 9
# One whose performance improves from instance to instance during training.
#
# ### 10
# Algorithms that can learn from datasets that are larger than available memory
#
# ### 11
# Instance-based learning
#
# ### 12
# A model parameter is optimized as part of the learning process, whereas a hyperparameter is set externally to the learning process
#
# ### 13
# Model-based learning searches for optimal values of set of prediction parameters (generally much smaller than the training data set). Usually an approach such as maximizing log-likelyhood is used to tune the parameters, and the model outputs are then used at the predictions.
#
# ### 14
# Insufficient data, non-representative sample data, poor quality data, irrelevant features
#
# ### 15
# Overfitting, consider reducing the free parameters or adding a regularization term.
#
# ### 16
# A set of data that is independent of the data used to develop the model, which can provide a better estimate of the ability of the model to generalize to new data.
#
# ### 17
# A validation set can be used to estimate the performance of the model on new data, and is used during model selection and hyperparameter tuning
#
# ### 18
# At times potentially non-representitive data is used to train a model because it is more readily available than representative data. The train-dev dataset is composed of a sample of this non-representative data that was not used for training, and allows you to distingish between failures to generalize due to overfitting and failures to generalize secondary to this non-representative data.
#
# ### 19
# You are prone to falsely overestimate the accuracy of your model on new data.
| handsonml2e/handson-ml2-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [pandas - restructuring dataframe using column names](https://stackoverflow.com/questions/52336218/pandas-restructuring-dataframe-using-column-names/52339373#52339373)
import pandas as pd
from pprint import pprint as pp
df = pd.read_csv('data/2018-09-14_stock_data.csv')
df.columns
# +
headers_new = {}
for x in list(df.columns):
headers_new[x] = x[-4:] + x[:-4]
headers_new
# -
df = df.rename(index=str, columns=headers_new)
df
df_long = pd.wide_to_long(df, ['(PO)', '(PI)'], i='Date', j='stock', suffix=r'(?<=\))(.*)')
df_long
df2 = df_long.reset_index()
df2.head()
# ## How to index
df_long.columns
df_long.index
df2.columns
df2.index
df_long.loc[('1978-09-25 0:00', 'S&PCOMP')]
df_long.loc[('1978-09-25 0:00')]
df2.iloc[:3]
# ## additional resources
#
# * [Data Reshaping with Pandas Explained](https://medium.com/@wangyuw/data-reshaping-with-pandas-explained-80b2f51f88d2)
# * [pandas iloc vs ix vs loc explanation, how are they different?](https://stackoverflow.com/questions/31593201/pandas-iloc-vs-ix-vs-loc-explanation-how-are-they-different)
# * [MultiIndex / Advanced Indexing](http://pandas.pydata.org/pandas-docs/stable/advanced.html)
| complete_solutions/2018-09-14_wide_to_long.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: twtenv
# language: python
# name: twtenv
# ---
# # Setup
# + id="nF-7m57sBgdp"
import json
# -
users = []
with open('../data/users.json') as json_file:
data = json.load(json_file)
users = data["all"]
len(users)
print(users[:5])
print(users[-5:])
| codes/read_accounts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.12 64-bit (''work'': conda)'
# name: python3
# ---
import xml.etree.ElementTree as et
import numpy as np
import matplotlib.pyplot as plt
hsa = et.parse("hsa00140.xml")
root = hsa.getroot()
for child in root[1]:
pathway = child.attrib['name'].replace('TITLE:', '')
for i in range(len(root)):
for child in root[i]:
try:
print(child.attrib)
except:
pass
break
for child in root[0]:
print(child.attrib)
genes = [gene for gene in root[0].attrib['name'].split(' ')]
print(genes)
expr = [0, 1, 0, 1, 1, 1]
# +
label_loc = np.linspace(start=0, stop=2 * np.pi, num=len(x))
plt.figure(figsize=(8, 8))
plt.subplot(polar=True)
plt.plot(label_loc, x, label=pathway)
plt.title('Pathway Analysis', size=20)
lines, labels = plt.thetagrids(np.degrees(label_loc), labels=genes)
plt.legend()
plt.show()
# -
| radar_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Neural Network
# Modules
import torch
import torch.nn as nn
import torchvision
import torchvision.transforms as transforms
# Device configuration
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# Hyper parameters
num_epochs = 5
num_classes = 10
batch_size = 100
learning_rate = 0.001
# MNIST dataset
# +
train_dataset = torchvision.datasets.MNIST(root='../../data/',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = torchvision.datasets.MNIST(root='../../data/',
train=False,
transform=transforms.ToTensor())
# -
# Data loader
# +
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# -
# Convolutional neural network (two convolutional layers)
class ConvNet(nn.Module):
def __init__(self, num_classes=10):
super(ConvNet, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(1, 16, kernel_size=5, stride=1, padding=2),
nn.BatchNorm2d(16),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.layer2 = nn.Sequential(
nn.Conv2d(16, 32, kernel_size=5, stride=1, padding=2),
nn.BatchNorm2d(32),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.fc = nn.Linear(7*7*32, num_classes)
def forward(self, x):
out = self.layer1(x)
out = self.layer2(out)
out = out.reshape(out.size(0), -1)
out = self.fc(out)
return out
model = ConvNet(num_classes).to(device)
# Loss and optimizer
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# Train the model
total_step = len(train_loader)
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(train_loader):
images = images.to(device)
labels = labels.to(device)
# Forward pass
outputs = model(images)
loss = criterion(outputs, labels)
# Backward and optimize
optimizer.zero_grad()
loss.backward()
optimizer.step()
if (i+1) % 100 == 0:
print ('Epoch [{}/{}], Step [{}/{}], Loss: {:.4f}'
.format(epoch+1, num_epochs, i+1, total_step, loss.item()))
# Test the model
model.eval() # eval mode (batchnorm uses moving mean/variance instead of mini-batch mean/variance)
with torch.no_grad():
correct = 0
total = 0
for images, labels in test_loader:
images = images.to(device)
labels = labels.to(device)
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Test Accuracy of the model on the 10000 test images: {} %'.format(100 * correct / total))
# Save the model checkpoint
torch.save(model.state_dict(), 'model.ckpt')
| tutorials_jupyter/02-intermediate/convolutional_neural_network/main.ipynb |