code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Modified Unknown (Probabilistic) Constraint Function in Emukit
# Author: <NAME> (Massachusetts Institute of Technology)
# Date modified: Aug 17, 2020
#
# Version: 0.1: tested with two 2D functions - Branin and Six-hump
# - Partly modified unknown constraint function to accomodate different inputs for X and Xc
# - The optimization package is not supported; only able to re-run the model iteratively with appending new data
# - `ProbabilisticConstraintBayesianOptimizationLoop` is the modified unknow constraint loop
# +
# Copyright 2018 Amazon.com, Inc. or its affiliates. All Rights Reserved.
# SPDX-License-Identifier: Apache-2.0
import numpy as np
from typing import Union
from emukit.bayesian_optimization.acquisitions import ExpectedImprovement, ProbabilityOfFeasibility, ProbabilityOfImprovement
from emukit.core.acquisition import Acquisition
from emukit.core.interfaces import IModel, IDifferentiable
from emukit.core.loop import FixedIntervalUpdater, OuterLoop, SequentialPointCalculator
from emukit.core.loop.loop_state import create_loop_state
from emukit.core.optimization import AcquisitionOptimizerBase
from emukit.core.optimization import GradientAcquisitionOptimizer
from emukit.core.parameter_space import ParameterSpace
from emukit.bayesian_optimization.acquisitions.log_acquisition import LogAcquisition
from emukit.bayesian_optimization.local_penalization_calculator import LocalPenalizationPointCalculator
class ProbabilisticConstraintBayesianOptimizationLoop(OuterLoop):
def __init__(self, space: ParameterSpace, model_objective: Union[IModel, IDifferentiable],
model_constraint: Union[IModel, IDifferentiable], acquisition: Acquisition = None,
update_interval: int = 1, batch_size: int = 1):
"""
Emukit class that implements a loop for building Bayesian optimization with an unknown constraint.
For more information see:
<NAME>, <NAME>, and <NAME>,
Bayesian Optimization with Unknown Constraints,
https://arxiv.org/pdf/1403.5607.pdf
:param space: Input space where the optimization is carried out.
:param model_objective: The model that approximates the underlying objective function
:param model_constraint: The model that approximates the unknown constraints
:param acquisition: The acquisition function for the objective function (default, EI).
:param update_interval: Number of iterations between optimization of model hyper-parameters. Defaults to 1.
:param batch_size: How many points to evaluate in one iteration of the optimization loop. Defaults to 1.
"""
# if not np.all(np.isclose(model_objective.X, model_constraint.X)):
# raise ValueError('Emukit currently only supports identical '
# 'training inputs for the constrained and objective model')
if acquisition is None:
acquisition = ExpectedImprovement(model_objective)
acquisition_constraint = ProbabilityOfFeasibility(model_constraint)
acquisition_constrained = acquisition * acquisition_constraint
model_updater_objective = FixedIntervalUpdater(model_objective, update_interval)
model_updater_constraint = FixedIntervalUpdater(model_constraint, update_interval,
lambda state: state.Y_constraint)
acquisition_optimizer = GradientAcquisitionOptimizer(space)
if batch_size == 1:
candidate_point_calculator = SequentialPointCalculator(acquisition_constrained, acquisition_optimizer)
else:
log_acquisition = LogAcquisition(acquisition_constrained)
candidate_point_calculator = LocalPenalizationPointCalculator(log_acquisition, acquisition_optimizer,
model_objective, space, batch_size)
loop_state = create_loop_state(model_objective.X, model_objective.Y)
#loop_state = create_loop_state(model_objective.X, model_objective.Y, Y_constraint=model_constraint.Y)
super(ProbabilisticConstraintBayesianOptimizationLoop, self).__init__(candidate_point_calculator,
[model_updater_objective, model_updater_constraint],
loop_state)
def suggest_next_locations(self):
return self.candidate_point_calculator.compute_next_points(self.loop_state)
# def acquisition_evaluate(self, X: loop_sate.X):
# return acquisition.evaluate(X), acquisition_constraint.evaluate(X), acquisition_constrained.evaluate(X)
# -
# import seaborn as sns
# sns.set(font_scale=1.1)
# sns.set_style("ticks",{'xtick.direction': 'in','ytick.direction':'in','xtick.top': True,'ytick.right': True})
def plot_acquisitions(param_bound, f_cons_acq, f_raw_acq, f_cons_pr, X, Xc, Xnew):
xy = []
n_steps = 30
for x in np.linspace(param_bound[0][0], param_bound[0][1], n_steps):
for y in np.linspace(param_bound[1][0], param_bound[1][1], n_steps):
xy.append([x,y])
z1 = f_cons_acq(np.array(xy))
z2 = f_raw_acq(np.array(xy))
z3 = f_cons_pr(np.array(xy))
xc = np.array(xy)[:,0].reshape(n_steps, n_steps)
yc = np.array(xy)[:,1].reshape(n_steps, n_steps)
zc1 = np.array(z1).reshape(n_steps, n_steps)
zc2 = np.array(z2).reshape(n_steps, n_steps)
zc3 = np.array(z3).reshape(n_steps, n_steps)
fontsize = 12
title_pad = 15
fig,axes = plt.subplots(1, 3, figsize=(10, 3), sharey = False)
c_plt1 = axes[0].contourf(xc, yc, zc1, cmap='plasma')
fig.colorbar(c_plt1, ax= axes[0])
axes[0].scatter(X[:,0],X[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'white')
axes[0].scatter(Xnew[:,0],Xnew[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'green')
axes[0].set_title('Constrained Acquisition', fontsize=fontsize,pad = title_pad)
c_plt2 = axes[1].contourf(xc, yc, zc2, cmap='plasma')
fig.colorbar(c_plt2, ax= axes[1])
axes[1].scatter(X[:,0],X[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'white')
axes[1].scatter(Xnew[:,0],Xnew[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'green')
axes[1].set_title('Raw Acquisition', fontsize=fontsize, pad = title_pad)
c_plt3 = axes[2].contourf(xc, yc, zc3, np.arange(0,1.01,0.1), cmap='plasma')
fig.colorbar(c_plt3, ax= axes[2])
axes[2].scatter(Xc[:,0],Xc[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'white')
axes[2].scatter(Xnew[:,0],Xnew[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'green')
axes[2].set_title('Probabilistic Constraint', fontsize=fontsize, pad = title_pad)
for i in range(len(axes)):
axes[i].set_xlabel(obj_fcn+'_X1',fontsize=fontsize)
axes[i].set_ylabel(obj_fcn+'_X2',fontsize=fontsize)
fig.tight_layout()
plt.show()
# import seaborn as sns
# sns.set(font_scale=1.5)
# sns.set_style("ticks",{'xtick.direction': 'in','ytick.direction':'in','xtick.top': True,'ytick.right': True})
def plot_gp_functions(param_bound, f_obj, f_cons, X, Xc, Xnew):
xy = []
n_steps = 50
for x in np.linspace(param_bound[0][0], param_bound[0][1], n_steps):
for y in np.linspace(param_bound[1][0], param_bound[1][1], n_steps):
xy.append([x,y])
z1_mean, z1_var = f_obj(np.array(xy))
z2_mean, z2_var = f_cons(np.array(xy))
xc = np.array(xy)[:,0].reshape(n_steps, n_steps)
yc = np.array(xy)[:,1].reshape(n_steps, n_steps)
zc1_mean = np.array(z1_mean).reshape(n_steps, n_steps)
zc2_mean = np.array(z2_mean).reshape(n_steps, n_steps)
zc1_var = np.array(z1_var).reshape(n_steps, n_steps)
zc2_var = np.array(z2_var).reshape(n_steps, n_steps)
fontsize = 12
title_pad = 15
for zc1, zc2, label in zip([zc1_mean, zc1_var],[zc2_mean, zc2_var],['Mean', 'Variance']):
fig,axes = plt.subplots(1, 2, figsize=(7, 3), sharey = False)
c_plt1 = axes[0].contourf(xc, yc, zc1, cmap='plasma')
fig.colorbar(c_plt1, ax= axes[0])
axes[0].scatter(X[:,0],X[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'white')
axes[0].scatter(Xnew[:,0],Xnew[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'green')
axes[0].set_title('Objective Function - '+label, fontsize=fontsize, pad=title_pad)
c_plt2 = axes[1].contourf(xc, yc, zc2, cmap='plasma')
fig.colorbar(c_plt2, ax= axes[1])
axes[1].scatter(Xc[:,0],Xc[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'white')
axes[1].scatter(Xnew[:,0],Xnew[:,1], s = 30, facecolors='none', alpha = 1, edgecolor = 'green')
axes[1].set_title('Constrained Function - '+label, fontsize=fontsize, pad=title_pad)
for i in range(len(axes)):
axes[i].set_xlabel(obj_fcn+' X1',fontsize=fontsize)
axes[i].set_ylabel(obj_fcn+' X2',fontsize=fontsize)
fig.tight_layout()
plt.show()
# import seaborn as sns
# sns.set(font_scale=1.5)
# sns.set_style("ticks",{'xtick.direction': 'in','ytick.direction':'in','xtick.top': True,'ytick.right': True})
def plot_true_functions(param_bound, f_obj, f_cons):
xy = []
n_steps = 200
for x in np.linspace(param_bound[0][0], param_bound[0][1], n_steps):
for y in np.linspace(param_bound[1][0], param_bound[1][1], n_steps):
xy.append([x,y])
z1_mean = f_obj(np.array(xy))
z2_mean = f_cons(np.array(xy))
xc = np.array(xy)[:,0].reshape(n_steps, n_steps)
yc = np.array(xy)[:,1].reshape(n_steps, n_steps)
zc1 = np.array(z1_mean).reshape(n_steps, n_steps)
zc2 = np.array(z2_mean).reshape(n_steps, n_steps)
fontsize = 12
title_pad = 15
fig,axes = plt.subplots(1, 2, figsize=(7, 3), sharey = False)
c_plt1 = axes[0].contourf(xc, yc, zc1, cmap='plasma')
fig.colorbar(c_plt1, ax= axes[0])
axes[0].set_title('Objective Function', fontsize=fontsize, pad=title_pad)
c_plt2 = axes[1].contourf(xc, yc, zc2, cmap='plasma')
fig.colorbar(c_plt2, ax= axes[1])
axes[1].set_title('Constrained Function', fontsize=fontsize, pad=title_pad)
for i in range(len(axes)):
axes[i].set_xlabel(obj_fcn+' X1',fontsize=fontsize)
axes[i].set_ylabel(obj_fcn+' X2',fontsize=fontsize)
fig.tight_layout()
plt.show()
# +
import GPy
from GPy.models import GPRegression
import matplotlib.pyplot as plt
import numpy as np
# Test Functions
from emukit.test_functions import sixhumpcamel_function, branin_function
from emukit.core import ParameterSpace, ContinuousParameter, DiscreteParameter
from emukit.core.initial_designs.random_design import RandomDesign
from emukit.core.initial_designs.latin_design import LatinDesign
# Model Wrappers
from emukit.model_wrappers import GPyModelWrapper
from emukit.model_wrappers.gpy_quadrature_wrappers import BaseGaussianProcessGPy, RBFGPy
import warnings
#warnings.filterwarnings('ignore')
# Decision loops
from emukit.bayesian_optimization.loops import BayesianOptimizationLoop
# Acquisition functions
from emukit.bayesian_optimization.acquisitions import ExpectedImprovement, \
NegativeLowerConfidenceBound, \
MaxValueEntropySearch, \
MultipointExpectedImprovement,\
ProbabilityOfFeasibility
from emukit.core.acquisition import IntegratedHyperParameterAcquisition
from emukit.experimental_design.acquisitions import ModelVariance
# Acquistions optimizers
from emukit.core.optimization import GradientAcquisitionOptimizer
# Stopping conditions
from emukit.core.loop import FixedIterationsStoppingCondition
from emukit.core.loop import ConvergenceStoppingCondition
### Define the objective function
### In this case we use the Branin function available in Emukit.
obj_fcn = 'branin'
if obj_fcn == 'six_hump':
f, parameter_space = sixhumpcamel_function()
def fc(x):#sixhumpcamel constraints
return np.transpose([((x[:,0]-0.1)**2+(x[:,1]+0.6)**2-0.16)])+(np.random.random(1)-0.5)/10
elif obj_fcn == 'branin':
f, parameter_space = branin_function()
def fc(x): #branin constraints
return np.transpose([((x[:,0]-3)**2+(x[:,1]-3)**2-9)])+(np.random.random(1)-0.5)*4
else:
raise ValueError('Need to eitherdefine an objective function and its constraint functions'
'Or, use external X and Y inputs iteratively')
plot_true_functions(parameter_space.get_bounds(), f, fc)
# +
num_init_points = 5
#pre-sampling of objective function
design = LatinDesign(parameter_space)
x_init = design.get_samples(num_init_points)
y_init = f(x_init)
#pre-sampling of constraint function
num_add_cons_points = 5
xc_init = np.concatenate([x_init, design.get_samples(num_add_cons_points)])
yc_init = fc(xc_init)
X, Y, Xc, Yc = [x_init, y_init, xc_init, yc_init]
acq_val, acq_fcn, acq_cons = [[],[],[]]
n_iterations = 5
batch = 5
for i in range(n_iterations+1):
### Fit and wrap a model to the collected data
input_dim = len(X[0])
ker = GPy.kern.Matern52(input_dim = input_dim, ARD =False)#
ker.lengthscale.constrain_bounded(1e-5, 100.1)
ker.variance.constrain_bounded(1e-5, 1000.1)
ker += GPy.kern.Bias(input_dim = input_dim)
model_gpy = GPRegression(X , Y, ker)
# model_gpy.kern.set_prior(GPy.priors.Uniform(0,5))
# model_gpy.likelihood.variance.constrain_fixed(0.1)
model_gpy.randomize()
model_gpy.optimize_restarts(num_restarts=20,verbose =False, messages=False)
objective_model = GPyModelWrapper(model_gpy)
# Make GPy constraint model
ker = GPy.kern.Matern52(input_dim = len(Xc[0]), ARD =False)
ker.lengthscale.constrain_bounded(1e-5, 1000.1)
ker.variance.constrain_bounded(1e-5, 1000.1)
ker += GPy.kern.Bias(input_dim = input_dim)
gpy_constraint_model = GPRegression(Xc, Yc, ker)
gpy_constraint_model.randomize()
gpy_constraint_model.optimize_restarts(num_restarts=20,verbose =False, messages=False)
constraint_model = GPyModelWrapper(gpy_constraint_model)
## Expeceted Improvement (EI)
acquisition = ExpectedImprovement(objective_model, jitter=.01)
## Lower Confidence Bound (LCB)
#acquisition = NegativeLowerConfidenceBound(objective_model, beta = 1)
## fully integreated acquisition
# acquisition_generator = lambda m: ExpectedImprovement(m, jitter=.01)
# expected_improvement_integrated = IntegratedHyperParameterAcquisition(objective_model, acquisition_generator)
# acquisition = expected_improvement_integrated
# Make loop and collect points
bayesopt_cons_pr = ProbabilisticConstraintBayesianOptimizationLoop(model_objective=objective_model,
space=parameter_space,
acquisition=acquisition,
model_constraint=constraint_model,
batch_size = batch)
X_new = bayesopt_cons_pr.candidate_point_calculator.compute_next_points(bayesopt_cons_pr.loop_state)
Y_new = f(X_new) # replace by new experiment results
Xc_new = X_new
Yc_new = fc(X_new) # replace by new experiment results
f_cons_acq = bayesopt_cons_pr.candidate_point_calculator.acquisition.acquisition.evaluate
f_raw_acq = bayesopt_cons_pr.candidate_point_calculator.acquisition.acquisition.acquisition_1.evaluate
f_cons_pr = bayesopt_cons_pr.candidate_point_calculator.acquisition.acquisition.acquisition_2.evaluate
acq_product_iter = f_cons_acq(X_new)
acq_fcn_iter = f_raw_acq(X_new)
acq_cons_iter = f_cons_pr(X_new)
acq_val.append(acq_product_iter)
acq_fcn.append(acq_fcn_iter)
acq_cons.append(acq_cons_iter)
#print(acq_val)
param_bound = parameter_space.get_bounds()
f_obj = objective_model.model.predict
f_cons = constraint_model.model.predict
if len(bayesopt_cons_pr.loop_state.X)%10 == 0:
print('total sample no.', len(bayesopt_cons_pr.loop_state.X))
# constraint_model.model.plot()
# objective_model.model.plot()
# plt.show()
plot_gp_functions(param_bound, f_obj, f_cons, X, Xc, X_new)
plot_acquisitions(param_bound, f_cons_acq, f_raw_acq, f_cons_pr, X, Xc, X_new)
X = np.concatenate([X, X_new])
Y = np.concatenate([Y, Y_new])
Xc = np.concatenate([Xc, Xc_new])
Yc = np.concatenate([Yc, Yc_new])
# +
fig, ax = plt.subplots(3, 1, figsize=(6,8))
font_size = 14
marker_size = 5
ax[0].plot(np.arange(len(Y)), np.minimum.accumulate(Y),
marker = 'o', ms = marker_size, c = 'black')
ax[0].scatter(np.arange(len(Y)), Y, c = 'cyan')
ax[0].set_ylabel('Current best', fontsize = font_size)
ax[0].set_xlim(0, len(Y)+5)
ax[0].set_ylim(min(np.minimum.accumulate(Y))-1,
np.max(np.minimum.accumulate(Y))+abs(np.max(np.minimum.accumulate(Y)))*0.5)
ax[1].plot(np.arange(len(Y)-num_init_points)+num_init_points, np.concatenate(acq_cons),
marker = 'o', ms = marker_size)
ax[1].set_ylabel('Probabilistic Constrants', fontsize = font_size)
ax[1].set_xlim(0, len(Y)+5)
ax[1].set_ylim(0, 1.1)
ax[2].plot(np.arange(len(Y)-num_init_points)+num_init_points, np.concatenate(acq_fcn),
marker = 'o', ms = marker_size, c = 'red', label = 'raw_acqui')
ax[2].plot(np.arange(len(Y)-num_init_points)+num_init_points, np.concatenate(acq_val),'--',
marker = 'o', ms = marker_size,c = 'green', label = 'cons_acqui')
ax[2].set_ylabel('Acquisition Values', fontsize = font_size)
ax[2].set_xlabel('Iteration', fontsize = font_size)
ax[2].set_xlim(0, len(Y)+5)
ax[2].set_ylim(0, np.max(acq_fcn)*1.1)
ax[2].legend()
ax[2].set_xlim(0, len(Y)+5)
fig.tight_layout()
for a in ax:
a.plot(np.ones(1000)*(num_init_points+batch*(n_iterations)), np.arange(1000)-500, '--', linewidth = 1, color = 'gray' )
plt.show()
# -
| Synthetic Example/Emukit_test_modified_unknown_constraint_function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/python11312/projekt-transformacja/blob/master/day4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="O35SyjWNgLRf" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error
import numpy as np
from sklearn.model_selection import cross_val_score
# + id="V3mh3xHShr0n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d9a724ab-54ca-48a1-ef7a-441f53875cb9"
# cd '/content/drive/My Drive/Colab Notebooks/projekt-transformacja'
# + id="bpt8Xn7Xh6Bm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5192f529-2b59-49cf-91ec-aafdd033bba6"
# ls data
# + id="IbXvzjHCh8oN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9db6b290-7a15-4c78-e2ab-7533c0e5cc48"
df = pd.read_csv('data/men_shoes.csv',low_memory=False)
df.shape
# + id="tDRNZ6EaiKEN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="2953d81e-4483-4152-b2bc-7e9e983aa914"
df.columns
# + id="ENL28Wd3iXBC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f757963b-6498-4a79-e5f1-928c7267c4c9"
mean_price = np.mean(df['prices_amountmin'])
mean_price
# + id="VI2yORtriyDT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fa1ccf22-a785-4db0-dcbb-f78090294aaf"
y_true = df['prices_amountmin']
y_pred = [mean_price] * y_true.shape[0]
mean_absolute_error( y_true , y_pred )
# + id="Sw5MBYwmjOLD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="54ae2e26-66f3-43a4-cbab-83e1bf0b2073"
np.log( df['prices_amountmin'] + 1) .hist(bins = 100)
# + id="pPc1kKBnkOLg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b60fa5e5-4f7c-45cb-e3b5-3dfb2d305448"
y_true = df['prices_amountmin']
y_pred = [np.median(y_true)] * y_true.shape[0]
mean_absolute_error( y_true , y_pred )
# + id="uERpdjunkurl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2911f365-9bbc-4c01-e2f8-71de007a2c76"
y_true = df['prices_amountmin']
price_log_mean = np.expm1( np.mean(np.log1p(y_true) ) )
y_pred = [price_log_mean] * y_true.shape[0]
mean_absolute_error( y_true , y_pred )
# + id="O83pGg1tl6mh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="f75c5164-5027-4097-8b39-89f873aa9e77"
df.columns
# + id="CdzlGCJYmRjT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="718ff2f4-5e88-4311-dad0-452d7fe6a22b"
df.brand.value_counts()
# + id="TXpXM4QimUvZ" colab_type="code" colab={}
df['brand_cat'] = df['brand'].factorize()[0]
# + id="AqlvSDRktBPq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="91495bba-0334-4651-a8d8-58e7cd713762"
feats = ['brand_cat']
x = df[ feats ].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
score = cross_val_score(model, x, y, scoring='neg_mean_absolute_error')
np.mean(score), np.std(score)
# + id="C3ZfcNfNQ8Oh" colab_type="code" colab={}
# + id="4Zi-dl3PSIz4" colab_type="code" colab={}
# + id="0dhfGCMuSOhr" colab_type="code" colab={}
# + id="-MPcmwlrSR_v" colab_type="code" colab={}
def run_model(feats):
x = df[ feats ].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
score = cross_val_score(model, x, y, scoring='neg_mean_absolute_error')
return np.mean(score), np.std(score)
run_model('brand_cat')
# + id="QOLDoeR5TP7P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="475e6a5e-abc3-4131-d05c-0a04af8b9720"
run_model(['brand_cat'])
# + id="31i7ba_-Tif6" colab_type="code" colab={}
# + id="B4SXmacLUOHe" colab_type="code" colab={}
# + id="DbgAfG8UUWR2" colab_type="code" colab={}
#run_model(['manufacturer'])
# + id="rZNhAyzNUdp9" colab_type="code" colab={}
#def run_model(feats):
# x = df[ feats ].values
#y = df['prices_amountmin'].values
#model = DecisionTreeRegressor(max_depth=5)
#score = cross_val_score(model, x, y, scoring='neg_mean_absolute_error')
#return np.mean(score), np.std(score)
# + id="BSAYm4KSUoKg" colab_type="code" colab={}
#run_model(['manufacturer'])
# + id="ILEENPZLUz4Y" colab_type="code" colab={}
#feats = ['manufacturer_cat']
#x = df[ feats ].values
#y = df['prices_amountmin'].values
#model = DecisionTreeRegressor(max_depth=5)
#score = cross_val_score(model, x, y, scoring='neg_mean_absolute_error')
#np.mean(score), np.std(score)
# + id="1lnTH4SnU74M" colab_type="code" colab={}
#run_model(['manufacturer'],['brand_cat'])
# + id="alYeAa9IVgDC" colab_type="code" colab={}
#run_model(['manufacturer','brand_cat'])
# + id="3hT2_qv5VucQ" colab_type="code" colab={}
#df.manufacturer.values_counts()
# + id="LkQ8qR0tV27_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="84da9a40-a891-4f5d-e488-7c78a7c4737f"
df.manufacturer.value_counts()
# + id="dCCdr6efWD-2" colab_type="code" colab={}
df['manufacturer_cat'] = df['manufacturer'].factorize()[0]
# + id="8O_Qj_SkWf64" colab_type="code" colab={}
feats = ['manufacturer_cat']
def run_code(feats):
x = df[ feats ].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
score = cross_val_score(model, x, y, scoring='neg_mean_absolute_error')
return np.mean(score), np.std(score)
# + id="hFu6qCVEWp53" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ae4775d9-821d-4e8b-8cb3-ab56a3f4faba"
run_model(['manufacturer_cat'])
# + id="vXR_mJ75W2bx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bcac42b2-28d3-4720-d93b-a009d646535b"
run_model(['manufacturer_cat','brand_cat'])
# + id="ZlUBVyiRW8sb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="185d1303-c0e7-46e5-947a-c085c50b05c3"
# !git push -u origin master
# + id="rN54MioUXsDs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5f4c3a54-221d-4518-a792-6179c59d497c"
# ls
# + id="M_jfSecSX3QV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e0e1d5b9-ac09-4d78-ed6f-08c01f7f97b3"
# !git add matrix_one/day4.ipynb
# + id="yexMuh6vYBzs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a6de207d-d8de-456c-b781-ba5eeaa199dc"
# ls
# + id="y-0Ua-PMYYDH" colab_type="code" colab={}
# !git add day4.ipynb
# + id="Yswy40MkYdQy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c2914073-29a3-45f7-8935-34d9991ec7bf"
# !git push-u origin master
# + id="ocWr5sTgYjKb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="bb5a9641-29fd-4ece-9246-8cf164cf3460"
# !git push -u origin master
# + id="P6ewmduhYngn" colab_type="code" colab={}
| day4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <center>
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/Logos/organization_logo/organization_logo.png" width="300" alt="cognitiveclass.ai logo" />
# </center>
#
# # Area Plots, Histograms, and Bar Plots
#
# Estimated time needed: **30** minutes
#
# ## Objectives
#
# After completing this lab you will be able to:
#
# - Create additional labs namely area plots, histogram and bar charts
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Table of Contents
#
# <div class="alert alert-block alert-info" style="margin-top: 20px">
#
# 1. [Exploring Datasets with _pandas_](#0)<br>
# 2. [Downloading and Prepping Data](#2)<br>
# 3. [Visualizing Data using Matplotlib](#4) <br>
# 4. [Area Plots](#6) <br>
# 5. [Histograms](#8) <br>
# 6. [Bar Charts](#10) <br>
# </div>
# <hr>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Exploring Datasets with _pandas_ and Matplotlib<a id="0"></a>
#
# Toolkits: The course heavily relies on [_pandas_](http://pandas.pydata.org?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ) and [**Numpy**](http://www.numpy.org?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ) for data wrangling, analysis, and visualization. The primary plotting library that we are exploring in the course is [Matplotlib](http://matplotlib.org?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
#
# Dataset: Immigration to Canada from 1980 to 2013 - [International migration flows to and from selected countries - The 2015 revision](http://www.un.org/en/development/desa/population/migration/data/empirical2/migrationflows.shtml?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ) from United Nation's website.
#
# The dataset contains annual data on the flows of international migrants as recorded by the countries of destination. The data presents both inflows and outflows according to the place of birth, citizenship or place of previous / next residence both for foreigners and nationals. For this lesson, we will focus on the Canadian Immigration data.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Downloading and Prepping Data <a id="2"></a>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Import Primary Modules. The first thing we'll do is import two key data analysis modules: _pandas_ and **Numpy**.
#
# + button=false new_sheet=false run_control={"read_only": false}
import numpy as np # useful for many scientific computing in Python
import pandas as pd # primary data structure library
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Let's download and import our primary Canadian Immigration dataset using _pandas_ `read_excel()` method. Normally, before we can do that, we would need to download a module which _pandas_ requires to read in excel files. This module is **xlrd**. For your convenience, we have pre-installed this module, so you would not have to worry about that. Otherwise, you would need to run the following line of code to install the **xlrd** module:
#
# ```
# # # !conda install -c anaconda xlrd --yes
# ```
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Download the dataset and read it into a _pandas_ dataframe.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can = pd.read_excel('https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork/Data%20Files/Canada.xlsx',
sheet_name='Canada by Citizenship',
skiprows=range(20),
skipfooter=2
)
print('Data downloaded and read into a dataframe!')
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Let's take a look at the first five items in our dataset.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Let's find out how many entries there are in our dataset.
#
# + button=false new_sheet=false run_control={"read_only": false}
# print the dimensions of the dataframe
print(df_can.shape)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Clean up data. We will make some modifications to the original dataset to make it easier to create our visualizations. Refer to `Introduction to Matplotlib and Line Plots` lab for the rational and detailed description of the changes.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### 1. Clean up the dataset to remove columns that are not informative to us for visualization (eg. Type, AREA, REG).
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can.drop(['AREA', 'REG', 'DEV', 'Type', 'Coverage'], axis=1, inplace=True)
# let's view the first five elements and see how the dataframe was changed
df_can.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Notice how the columns Type, Coverage, AREA, REG, and DEV got removed from the dataframe.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### 2. Rename some of the columns so that they make sense.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can.rename(columns={'OdName':'Country', 'AreaName':'Continent','RegName':'Region'}, inplace=True)
# let's view the first five elements and see how the dataframe was changed
df_can.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Notice how the column names now make much more sense, even to an outsider.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### 3. For consistency, ensure that all column labels of type string.
#
# + button=false new_sheet=false run_control={"read_only": false}
# let's examine the types of the column labels
all(isinstance(column, str) for column in df_can.columns)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Notice how the above line of code returned _False_ when we tested if all the column labels are of type **string**. So let's change them all to **string** type.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can.columns = list(map(str, df_can.columns))
# let's check the column labels types now
all(isinstance(column, str) for column in df_can.columns)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### 4. Set the country name as index - useful for quickly looking up countries using .loc method.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can.set_index('Country', inplace=True)
# let's view the first five elements and see how the dataframe was changed
df_can.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Notice how the country names now serve as indices.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### 5. Add total column.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can['Total'] = df_can.sum(axis=1)
# let's view the first five elements and see how the dataframe was changed
df_can.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Now the dataframe has an extra column that presents the total number of immigrants from each country in the dataset from 1980 - 2013. So if we print the dimension of the data, we get:
#
# + button=false new_sheet=false run_control={"read_only": false}
print ('data dimensions:', df_can.shape)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# So now our dataframe has 38 columns instead of 37 columns that we had before.
#
# + button=false new_sheet=false run_control={"read_only": false}
# finally, let's create a list of years from 1980 - 2013
# this will come in handy when we start plotting the data
years = list(map(str, range(1980, 2014)))
years
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Visualizing Data using Matplotlib<a id="4"></a>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Import `Matplotlib` and **Numpy**.
#
# + button=false new_sheet=false run_control={"read_only": false}
# use the inline backend to generate the plots within the browser
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.style.use('ggplot') # optional: for ggplot-like style
# check for latest version of Matplotlib
print ('Matplotlib version: ', mpl.__version__) # >= 2.0.0
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Area Plots<a id="6"></a>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# In the last module, we created a line plot that visualized the top 5 countries that contribued the most immigrants to Canada from 1980 to 2013. With a little modification to the code, we can visualize this plot as a cumulative plot, also knows as a **Stacked Line Plot** or **Area plot**.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can.sort_values(['Total'], ascending=False, axis=0, inplace=True)
# get the top 5 entries
df_top5 = df_can.head()
# transpose the dataframe
df_top5 = df_top5[years].transpose()
df_top5.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Area plots are stacked by default. And to produce a stacked area plot, each column must be either all positive or all negative values (any NaN values will defaulted to 0). To produce an unstacked plot, pass `stacked=False`.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_top5.index = df_top5.index.map(int) # let's change the index values of df_top5 to type integer for plotting
df_top5.plot(kind='area',
stacked=False,
figsize=(20, 10), # pass a tuple (x, y) size
)
plt.title('Immigration Trend of Top 5 Countries')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# The unstacked plot has a default transparency (alpha value) at 0.5. We can modify this value by passing in the `alpha` parameter.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_top5.plot(kind='area',
alpha=0.25, # 0-1, default value a= 0.5
stacked=False,
figsize=(20, 10),
)
plt.title('Immigration Trend of Top 5 Countries')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Two types of plotting
#
# As we discussed in the video lectures, there are two styles/options of ploting with `matplotlib`. Plotting using the Artist layer and plotting using the scripting layer.
#
# **Option 1: Scripting layer (procedural method) - using matplotlib.pyplot as 'plt' **
#
# You can use `plt` i.e. `matplotlib.pyplot` and add more elements by calling different methods procedurally; for example, `plt.title(...)` to add title or `plt.xlabel(...)` to add label to the x-axis.
#
# ```python
# # Option 1: This is what we have been using so far
# df_top5.plot(kind='area', alpha=0.35, figsize=(20, 10))
# plt.title('Immigration trend of top 5 countries')
# plt.ylabel('Number of immigrants')
# plt.xlabel('Years')
# ```
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Option 2: Artist layer (Object oriented method) - using an `Axes` instance from Matplotlib (preferred) **
#
# You can use an `Axes` instance of your current plot and store it in a variable (eg. `ax`). You can add more elements by calling methods with a little change in syntax (by adding "_set__" to the previous methods). For example, use `ax.set_title()` instead of `plt.title()` to add title, or `ax.set_xlabel()` instead of `plt.xlabel()` to add label to the x-axis.
#
# This option sometimes is more transparent and flexible to use for advanced plots (in particular when having multiple plots, as you will see later).
#
# In this course, we will stick to the **scripting layer**, except for some advanced visualizations where we will need to use the **artist layer** to manipulate advanced aspects of the plots.
#
# + button=false new_sheet=false run_control={"read_only": false}
# option 2: preferred option with more flexibility
ax = df_top5.plot(kind='area', alpha=0.35, figsize=(20, 10))
ax.set_title('Immigration Trend of Top 5 Countries')
ax.set_ylabel('Number of Immigrants')
ax.set_xlabel('Years')
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Question**: Use the scripting layer to create a stacked area plot of the 5 countries that contributed the least to immigration to Canada **from** 1980 to 2013. Use a transparency value of 0.45.
#
# + button=false new_sheet=false run_control={"read_only": false}
### type your answer here
df_least5 = df_can.tail(5)
# transpose the dataframe
df_least5 = df_least5[years].transpose()
df_least5.head()
df_least5.index = df_least5.index.map(int) # let's change the index values of df_least5 to type integer for plotting
df_least5.plot(kind='area', alpha=0.45, figsize=(20, 10))
plt.title('Immigration Trend of 5 Countries with Least Contribution to Immigration')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <details><summary>Click here for a sample python solution</summary>
#
# ```python
# #The correct answer is:
# # get the 5 countries with the least contribution
# df_least5 = df_can.tail(5)
#
# # transpose the dataframe
# df_least5 = df_least5[years].transpose()
# df_least5.head()
#
# df_least5.index = df_least5.index.map(int) # let's change the index values of df_least5 to type integer for plotting
# df_least5.plot(kind='area', alpha=0.45, figsize=(20, 10))
#
# plt.title('Immigration Trend of 5 Countries with Least Contribution to Immigration')
# plt.ylabel('Number of Immigrants')
# plt.xlabel('Years')
#
# plt.show()
#
# ```
#
# </details>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Question**: Use the artist layer to create an unstacked area plot of the 5 countries that contributed the least to immigration to Canada **from** 1980 to 2013. Use a transparency value of 0.55.
#
# + button=false new_sheet=false run_control={"read_only": false}
### type your answer here
df_least5 = df_can.tail(5)
# transpose the dataframe
df_least5 = df_least5[years].transpose()
df_least5.head()
df_least5.index = df_least5.index.map(int) # let's change the index values of df_least5 to type integer for plotting
ax = df_least5.plot(kind='area', alpha=0.55, stacked=False, figsize=(20, 10))
ax.set_title('Immigration Trend of 5 Countries with Least Contribution to Immigration')
ax.set_ylabel('Number of Immigrants')
ax.set_xlabel('Years')
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <details><summary>Click here for a sample python solution</summary>
#
# ```python
# #The correct answer is:
#
# # get the 5 countries with the least contribution
# df_least5 = df_can.tail(5)
#
# # transpose the dataframe
# df_least5 = df_least5[years].transpose()
#
# df_least5.head()
#
# df_least5.index = df_least5.index.map(int) # let's change the index values of df_least5 to type integer for plotting
#
# ax = df_least5.plot(kind='area', alpha=0.55, stacked=False, figsize=(20, 10))
#
# ax.set_title('Immigration Trend of 5 Countries with Least Contribution to Immigration')
# ax.set_ylabel('Number of Immigrants')
# ax.set_xlabel('Years')
#
#
# ```
#
# </details>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Histograms<a id="8"></a>
#
# A histogram is a way of representing the _frequency_ distribution of numeric dataset. The way it works is it partitions the x-axis into _bins_, assigns each data point in our dataset to a bin, and then counts the number of data points that have been assigned to each bin. So the y-axis is the frequency or the number of data points in each bin. Note that we can change the bin size and usually one needs to tweak it so that the distribution is displayed nicely.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Question:** What is the frequency distribution of the number (population) of new immigrants from the various countries to Canada in 2013?
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Before we proceed with creating the histogram plot, let's first examine the data split into intervals. To do this, we will us **Numpy**'s `histrogram` method to get the bin ranges and frequency counts as follows:
#
# + button=false new_sheet=false run_control={"read_only": false}
# let's quickly view the 2013 data
df_can['2013'].head()
# + button=false new_sheet=false run_control={"read_only": false}
# np.histogram returns 2 values
count, bin_edges = np.histogram(df_can['2013'])
print(count) # frequency count
print(bin_edges) # bin ranges, default = 10 bins
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# By default, the `histrogram` method breaks up the dataset into 10 bins. The figure below summarizes the bin ranges and the frequency distribution of immigration in 2013. We can see that in 2013:
#
# - 178 countries contributed between 0 to 3412.9 immigrants
# - 11 countries contributed between 3412.9 to 6825.8 immigrants
# - 1 country contributed between 6285.8 to 10238.7 immigrants, and so on..
#
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DV0101EN/labs/Images/Mod2Fig1-Histogram.JPG" align="center" width=800>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# We can easily graph this distribution by passing `kind=hist` to `plot()`.
#
# + button=false new_sheet=false run_control={"read_only": false}
df_can['2013'].plot(kind='hist', figsize=(8, 5))
plt.title('Histogram of Immigration from 195 Countries in 2013') # add a title to the histogram
plt.ylabel('Number of Countries') # add y-label
plt.xlabel('Number of Immigrants') # add x-label
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# In the above plot, the x-axis represents the population range of immigrants in intervals of 3412.9. The y-axis represents the number of countries that contributed to the aforementioned population.
#
# Notice that the x-axis labels do not match with the bin size. This can be fixed by passing in a `xticks` keyword that contains the list of the bin sizes, as follows:
#
# + button=false new_sheet=false run_control={"read_only": false}
# 'bin_edges' is a list of bin intervals
count, bin_edges = np.histogram(df_can['2013'])
df_can['2013'].plot(kind='hist', figsize=(8, 5), xticks=bin_edges)
plt.title('Histogram of Immigration from 195 countries in 2013') # add a title to the histogram
plt.ylabel('Number of Countries') # add y-label
plt.xlabel('Number of Immigrants') # add x-label
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# _Side Note:_ We could use `df_can['2013'].plot.hist()`, instead. In fact, throughout this lesson, using `some_data.plot(kind='type_plot', ...)` is equivalent to `some_data.plot.type_plot(...)`. That is, passing the type of the plot as argument or method behaves the same.
#
# See the _pandas_ documentation for more info [http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.plot.html](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.plot.html?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# We can also plot multiple histograms on the same plot. For example, let's try to answer the following questions using a histogram.
#
# **Question**: What is the immigration distribution for Denmark, Norway, and Sweden for years 1980 - 2013?
#
# + button=false new_sheet=false run_control={"read_only": false}
# let's quickly view the dataset
df_can.loc[['Denmark', 'Norway', 'Sweden'], years]
# + button=false new_sheet=false run_control={"read_only": false}
# generate histogram
df_can.loc[['Denmark', 'Norway', 'Sweden'], years].plot.hist()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# That does not look right!
#
# Don't worry, you'll often come across situations like this when creating plots. The solution often lies in how the underlying dataset is structured.
#
# Instead of plotting the population frequency distribution of the population for the 3 countries, _pandas_ instead plotted the population frequency distribution for the `years`.
#
# This can be easily fixed by first transposing the dataset, and then plotting as shown below.
#
# + button=false new_sheet=false run_control={"read_only": false}
# transpose dataframe
df_t = df_can.loc[['Denmark', 'Norway', 'Sweden'], years].transpose()
df_t.head()
# + button=false new_sheet=false run_control={"read_only": false}
# generate histogram
df_t.plot(kind='hist', figsize=(10, 6))
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Let's make a few modifications to improve the impact and aesthetics of the previous plot:
#
# - increase the bin size to 15 by passing in `bins` parameter
# - set transparency to 60% by passing in `alpha` paramemter
# - label the x-axis by passing in `x-label` paramater
# - change the colors of the plots by passing in `color` parameter
#
# + button=false new_sheet=false run_control={"read_only": false}
# let's get the x-tick values
count, bin_edges = np.histogram(df_t, 15)
# un-stacked histogram
df_t.plot(kind ='hist',
figsize=(10, 6),
bins=15,
alpha=0.6,
xticks=bin_edges,
color=['coral', 'darkslateblue', 'mediumseagreen']
)
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Tip:
# For a full listing of colors available in Matplotlib, run the following code in your python shell:
#
# ```python
# import matplotlib
# for name, hex in matplotlib.colors.cnames.items():
# print(name, hex)
# ```
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# If we do no want the plots to overlap each other, we can stack them using the `stacked` paramemter. Let's also adjust the min and max x-axis labels to remove the extra gap on the edges of the plot. We can pass a tuple (min,max) using the `xlim` paramater, as show below.
#
# + button=false new_sheet=false run_control={"read_only": false}
count, bin_edges = np.histogram(df_t, 15)
xmin = bin_edges[0] - 10 # first bin value is 31.0, adding buffer of 10 for aesthetic purposes
xmax = bin_edges[-1] + 10 # last bin value is 308.0, adding buffer of 10 for aesthetic purposes
# stacked Histogram
df_t.plot(kind='hist',
figsize=(10, 6),
bins=15,
xticks=bin_edges,
color=['coral', 'darkslateblue', 'mediumseagreen'],
stacked=True,
xlim=(xmin, xmax)
)
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Question**: Use the scripting layer to display the immigration distribution for Greece, Albania, and Bulgaria for years 1980 - 2013? Use an overlapping plot with 15 bins and a transparency value of 0.35.
#
# + button=false new_sheet=false run_control={"read_only": false}
### type your answer here
df_cof = df_can.loc[['Greece', 'Albania', 'Bulgaria'], years]
# transpose the dataframe
df_cof = df_cof.transpose()
# let's get the x-tick values
count, bin_edges = np.histogram(df_cof, 15)
# Un-stacked Histogram
df_cof.plot(kind ='hist',
figsize=(10, 6),
bins=15,
alpha=0.35,
xticks=bin_edges,
color=['coral', 'darkslateblue', 'mediumseagreen']
)
plt.title('Histogram of Immigration from Greece, Albania, and Bulgaria from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <details><summary>Click here for a sample python solution</summary>
#
# ```python
# #The correct answer is:
#
# # create a dataframe of the countries of interest (cof)
# df_cof = df_can.loc[['Greece', 'Albania', 'Bulgaria'], years]
#
# # transpose the dataframe
# df_cof = df_cof.transpose()
#
# # let's get the x-tick values
# count, bin_edges = np.histogram(df_cof, 15)
#
# # Un-stacked Histogram
# df_cof.plot(kind ='hist',
# figsize=(10, 6),
# bins=15,
# alpha=0.35,
# xticks=bin_edges,
# color=['coral', 'darkslateblue', 'mediumseagreen']
# )
#
# plt.title('Histogram of Immigration from Greece, Albania, and Bulgaria from 1980 - 2013')
# plt.ylabel('Number of Years')
# plt.xlabel('Number of Immigrants')
#
# plt.show()
#
#
# ```
#
# </details>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Bar Charts (Dataframe) <a id="10"></a>
#
# A bar plot is a way of representing data where the _length_ of the bars represents the magnitude/size of the feature/variable. Bar graphs usually represent numerical and categorical variables grouped in intervals.
#
# To create a bar plot, we can pass one of two arguments via `kind` parameter in `plot()`:
#
# - `kind=bar` creates a _vertical_ bar plot
# - `kind=barh` creates a _horizontal_ bar plot
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Vertical bar plot**
#
# In vertical bar graphs, the x-axis is used for labelling, and the length of bars on the y-axis corresponds to the magnitude of the variable being measured. Vertical bar graphs are particuarly useful in analyzing time series data. One disadvantage is that they lack space for text labelling at the foot of each bar.
#
# **Let's start off by analyzing the effect of Iceland's Financial Crisis:**
#
# The 2008 - 2011 Icelandic Financial Crisis was a major economic and political event in Iceland. Relative to the size of its economy, Iceland's systemic banking collapse was the largest experienced by any country in economic history. The crisis led to a severe economic depression in 2008 - 2011 and significant political unrest.
#
# **Question:** Let's compare the number of Icelandic immigrants (country = 'Iceland') to Canada from year 1980 to 2013.
#
# + button=false new_sheet=false run_control={"read_only": false}
# step 1: get the data
df_iceland = df_can.loc['Iceland', years]
df_iceland.head()
# + button=false new_sheet=false run_control={"read_only": false}
# step 2: plot data
df_iceland.plot(kind='bar', figsize=(10, 6))
plt.xlabel('Year') # add to x-label to the plot
plt.ylabel('Number of immigrants') # add y-label to the plot
plt.title('Icelandic immigrants to Canada from 1980 to 2013') # add title to the plot
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# The bar plot above shows the total number of immigrants broken down by each year. We can clearly see the impact of the financial crisis; the number of immigrants to Canada started increasing rapidly after 2008.
#
# Let's annotate this on the plot using the `annotate` method of the **scripting layer** or the **pyplot interface**. We will pass in the following parameters:
#
# - `s`: str, the text of annotation.
# - `xy`: Tuple specifying the (x,y) point to annotate (in this case, end point of arrow).
# - `xytext`: Tuple specifying the (x,y) point to place the text (in this case, start point of arrow).
# - `xycoords`: The coordinate system that xy is given in - 'data' uses the coordinate system of the object being annotated (default).
# - `arrowprops`: Takes a dictionary of properties to draw the arrow:
# - `arrowstyle`: Specifies the arrow style, `'->'` is standard arrow.
# - `connectionstyle`: Specifies the connection type. `arc3` is a straight line.
# - `color`: Specifes color of arror.
# - `lw`: Specifies the line width.
#
# I encourage you to read the Matplotlib documentation for more details on annotations:
# [http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.annotate](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.annotate?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
#
# + button=false new_sheet=false run_control={"read_only": false}
df_iceland.plot(kind='bar', figsize=(10, 6), rot=90) # rotate the xticks(labelled points on x-axis) by 90 degrees
plt.xlabel('Year')
plt.ylabel('Number of Immigrants')
plt.title('Icelandic Immigrants to Canada from 1980 to 2013')
# Annotate arrow
plt.annotate('', # s: str. Will leave it blank for no text
xy=(32, 70), # place head of the arrow at point (year 2012 , pop 70)
xytext=(28, 20), # place base of the arrow at point (year 2008 , pop 20)
xycoords='data', # will use the coordinate system of the object being annotated
arrowprops=dict(arrowstyle='->', connectionstyle='arc3', color='blue', lw=2)
)
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Let's also annotate a text to go over the arrow. We will pass in the following additional parameters:
#
# - `rotation`: rotation angle of text in degrees (counter clockwise)
# - `va`: vertical alignment of text [‘center’ | ‘top’ | ‘bottom’ | ‘baseline’]
# - `ha`: horizontal alignment of text [‘center’ | ‘right’ | ‘left’]
#
# + button=false new_sheet=false run_control={"read_only": false}
df_iceland.plot(kind='bar', figsize=(10, 6), rot=90)
plt.xlabel('Year')
plt.ylabel('Number of Immigrants')
plt.title('Icelandic Immigrants to Canada from 1980 to 2013')
# Annotate arrow
plt.annotate('', # s: str. will leave it blank for no text
xy=(32, 70), # place head of the arrow at point (year 2012 , pop 70)
xytext=(28, 20), # place base of the arrow at point (year 2008 , pop 20)
xycoords='data', # will use the coordinate system of the object being annotated
arrowprops=dict(arrowstyle='->', connectionstyle='arc3', color='blue', lw=2)
)
# Annotate Text
plt.annotate('2008 - 2011 Financial Crisis', # text to display
xy=(28, 30), # start the text at at point (year 2008 , pop 30)
rotation=72.5, # based on trial and error to match the arrow
va='bottom', # want the text to be vertically 'bottom' aligned
ha='left', # want the text to be horizontally 'left' algned.
)
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Horizontal Bar Plot**
#
# Sometimes it is more practical to represent the data horizontally, especially if you need more room for labelling the bars. In horizontal bar graphs, the y-axis is used for labelling, and the length of bars on the x-axis corresponds to the magnitude of the variable being measured. As you will see, there is more room on the y-axis to label categetorical variables.
#
# **Question:** Using the scripting layter and the `df_can` dataset, create a _horizontal_ bar plot showing the _total_ number of immigrants to Canada from the top 15 countries, for the period 1980 - 2013. Label each country with the total immigrant count.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Step 1: Get the data pertaining to the top 15 countries.
#
# + button=false new_sheet=false run_control={"read_only": false}
### type your answer here
df_can.sort_values(by='Total', ascending=True, inplace=True)
# get top 15 countries
df_top15 = df_can['Total'].tail(15)
df_top15
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <details><summary>Click here for a sample python solution</summary>
#
# ```python
# #The correct answer is:
#
# # sort dataframe on 'Total' column (descending)
# df_can.sort_values(by='Total', ascending=True, inplace=True)
#
# # get top 15 countries
# df_top15 = df_can['Total'].tail(15)
# df_top15
#
# ```
#
# </details>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Step 2: Plot data:
#
# 1. Use `kind='barh'` to generate a bar chart with horizontal bars.
# 2. Make sure to choose a good size for the plot and to label your axes and to give the plot a title.
# 3. Loop through the countries and annotate the immigrant population using the anotate function of the scripting interface.
#
# + button=false new_sheet=false run_control={"read_only": false}
### type your answer here
df_top15.plot(kind='barh', figsize=(12, 12), color='steelblue')
plt.xlabel('Number of Immigrants')
plt.title('Top 15 Conuntries Contributing to the Immigration to Canada between 1980 - 2013')
# annotate value labels to each country
for index, value in enumerate(df_top15):
label = format(int(value), ',') # format int with commas
# place text at the end of bar (subtracting 47000 from x, and 0.1 from y to make it fit within the bar)
plt.annotate(label, xy=(value - 47000, index - 0.10), color='white')
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <details><summary>Click here for a sample python solution</summary>
#
# ```python
# #The correct answer is:
#
# # generate plot
# df_top15.plot(kind='barh', figsize=(12, 12), color='steelblue')
# plt.xlabel('Number of Immigrants')
# plt.title('Top 15 Conuntries Contributing to the Immigration to Canada between 1980 - 2013')
#
# # annotate value labels to each country
# for index, value in enumerate(df_top15):
# label = format(int(value), ',') # format int with commas
#
# # place text at the end of bar (subtracting 47000 from x, and 0.1 from y to make it fit within the bar)
# plt.annotate(label, xy=(value - 47000, index - 0.10), color='white')
#
# plt.show()
#
# ```
#
# </details>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Thank you for completing this lab!
#
# ## Author
#
# <a href="https://www.linkedin.com/in/aklson/" target="_blank"><NAME></a>
#
# ### Other Contributors
#
# [<NAME>](https://www.linkedin.com/in/jayrajasekharan?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
# [<NAME>](https://www.linkedin.com/in/ehsanmkermani?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
# [<NAME>](https://www.linkedin.com/in/slobodan-markovic?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DV0101EN-SkillsNetwork-20297740&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
#
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ------------- | ---------------------------------- |
# | 2020-11-03 | 2.1 | <NAME> | Changed the URL of excel file |
# | 2020-08-27 | 2.0 | Lavanya | Moved lab to course repo in GitLab |
# | | | | |
# | | | | |
#
# ## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
#
| Data Science Certification/Data Visualization with Python/DV0101EN-2-2-1-Area-Plots-Histograms-and-Bar-Charts-py-v2.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit ('tf-workshop')
# metadata:
# interpreter:
# hash: 54c98f930adf7f56bf8f478a34adb9af0d8e994fc2bba90a19851c41a7ab1a64
# name: Python 3.6.9 64-bit ('tf-workshop')
# ---
# # Mejorando la Precisión a través de convoluciones
#
# En el anterior notebook hemos visto cómo implementar una red neuronal profunda con 3 capas, una capa de entrada, una capa oculta y la capa de salida. También hemos experimentado usando distinto número de unidades y capas ocultas para observar su efecto en la precisión del modelo entrenado.
#
# Para una referencia conveniente, a continuación tenemos el código completo para tener un valor inicial de precisión:
# + tags=[]
import tensorflow as tf
mnist = tf.keras.datasets.fashion_mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images / 255.0
test_images=test_images / 255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(training_images, training_labels, epochs=5)
test_loss = model.evaluate(test_images, test_labels)
# -
# La precisión alcanzada será aproximadamanete 89% en entrenamiento y 87% en validación, nada mal; pero, cómo podemos mejorar este rendimiento?
#
# Una forma es usar algo llamado Convolución. Una Convolución es una operación matemática que puede ser aplicada a señales de distinta naturaleza, en este caso, la señal a aplicar será nuestra imagen y sus correspondientes mapas de características en las capas ocultas.
#
# Pero antes de poder aplicar esta operación, debemos definir de mejor manera qué es y cómo se usa una convolución en imágenes.
#
# ## Convoluciones
# En escencia, tendremos un arreglo llamado **parche** o **kernel** (usualmente de 3x3 o 5x5) y lo *deslizaremos* sobre la imagen en la que queremos operar. Se puede observar una visualización de la misma operación en la siguiente figura:
#
# 
#
# en el caso anterior, un parche de 3x3 se desliza sobre una imagen original de tamaño 5x5 obteniendo como resultado una **nueva imagen** de tamaño 5x5, nótese que para que podamos tener una imagen del mismo tamaño original necesitamos agregar valores *extra* en los extremos de la imagen de entrada, esta técnica se denomina *padding*.
#
# ## Kernels de convolución
#
# En el contexto de las redes neuronales, intercambiaremos en nuestra arquitectura la estructura tradicional de las **neuronas** o unidades en las capas de la red por **filtros**. En la siguiente figura se puede visualizar el efecto de estos filtros.
#
# 
#
# Por tanto, durante el entrenamiento, la tarea es encontrar los valores más adecuados de cada elemento de los filtros para minimizar la función de costo o pérdida.
#
# La naturaleza y características de la operación de convolución son ideales para ser implementadas en tareas de visión artificial pues permiten a la red, a través de los filtros entrenados, *resaltar* las características más importantes para la predicción y mejora la eficiencia computacional pues nos enfocamos solamente en entrenar sobre ésas mismas características resaltadas.
#
# ## Fashion MNIST con una red convolucional
#
# Una vez entendido el concepto e importancia de la operación de convolución podemos aplicarlo a nuestra tarea de clasificación de prendas de vestir.
#
# Keras nos ofrece, mediante su modelo secuencial, definir una capa especial llamada **Conv2D** que implementa todas las operaciones.
#
# La implementación mejorada se vería así:
#
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.fashion_mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images.reshape(60000, 28, 28, 1)
training_images=training_images / 255.0
test_images = test_images.reshape(10000, 28, 28, 1)
test_images=test_images/255.0
# definición del modelo
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(64, (3,3), activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.summary()
model.fit(training_images, training_labels, epochs=5)
test_loss = model.evaluate(test_images, test_labels)
# Analice la diferencia en la precisión obtenida con nuestra nueva red convolucional.
#
#
# ## Ejercicios de exploración
#
# Incremente el número de épocas en el entrenamiento y analice cuidadosamente el efecto en la precisión final tanto en el conjunto de entrenamiento como en el conjunto de validación.
#
# ## Visualizando las capas convolucionales
#
# Visualizaremos los efectos de la convolución y los **mapas de características** de las capas ocultas de manera visual.
#
# De los 100 primeros ejemplos en el conjunto de pruebas, aquellos entre el índice 23 y 28 son zapatos. Analice la similaridad entre sus mapas de características.
#
# + tags=[]
print(test_labels[:100])
# -
import matplotlib.pyplot as plt
f, axarr = plt.subplots(3,4)
FIRST_IMAGE=0
SECOND_IMAGE=7
THIRD_IMAGE=26
CONVOLUTION_NUMBER = 1
from tensorflow.keras import models
layer_outputs = [layer.output for layer in model.layers]
activation_model = tf.keras.models.Model(inputs = model.input, outputs = layer_outputs)
for x in range(0,4):
f1 = activation_model.predict(test_images[FIRST_IMAGE].reshape(1, 28, 28, 1))[x]
axarr[0,x].imshow(f1[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[0,x].grid(False)
f2 = activation_model.predict(test_images[SECOND_IMAGE].reshape(1, 28, 28, 1))[x]
axarr[1,x].imshow(f2[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[1,x].grid(False)
f3 = activation_model.predict(test_images[THIRD_IMAGE].reshape(1, 28, 28, 1))[x]
axarr[2,x].imshow(f3[0, : , :, CONVOLUTION_NUMBER], cmap='inferno')
axarr[2,x].grid(False)
# ## Ejercicios
# 1. Modifique la cantidad de filtros de convolución, cambie de 32 a 16 o 64, cuál es el impacto en la precisión y/o el tiempo de entrenamiento?
# 2. Elimine la última capa de convolución, cuál es el impacto?
# 3. Y qué pasa si se agregan nuevas capas de convolución?
#
| 5.convnets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
import sys
import datetime
import os
import matplotlib.pyplot as plt
import matplotlib
import networkx as nx
import pickle
from collections import OrderedDict
import copy
from scipy.sparse import csr_matrix
from scipy import io
import seaborn as sns
import joblib
# from base import *
from joblib import Parallel, delayed
import random
import scipy
MNM_nb_folder = os.path.join('..', '..', '..', 'side_project', 'network_builder')
sys.path.append(MNM_nb_folder)
python_lib_folder = os.path.join('..', '..', 'pylib')
sys.path.append(python_lib_folder)
from MNMAPI import *
from MNM_mcnb import *
from mcDODE import *
data_folder = os.path.join('/home/lemma/Documents/MAC-POSTS/data/input_files_7link_multiclass')
nb = MNM_network_builder()
nb.load_from_folder(data_folder)
observed_link_list = [3, 4, 6]
ml_car = 2
ml_truck = 1
data_dict = dict()
num_interval = nb.config.config_dict['DTA']['max_interval']
true_car_x = np.random.rand(num_interval * len(observed_link_list)) * 100
true_truck_x = np.random.rand(num_interval * len(observed_link_list)) * 10
L_car_one = np.random.randint(2, size = (ml_car, len(observed_link_list)))
L_truck_one = np.random.randint(2, size = (ml_truck, len(observed_link_list)))
L_car = csr_matrix(scipy.linalg.block_diag(*[L_car_one for i in range(num_interval)]))
L_truck = csr_matrix(scipy.linalg.block_diag(*[L_truck_one for i in range(num_interval)]))
m_car = L_car.dot(true_car_x)
m_truck = L_truck.dot(true_truck_x)
data_dict['car_count_agg_L_list'] = [L_car]
data_dict['truck_count_agg_L_list'] = [L_truck]
data_dict['car_link_flow'] = [m_car]
data_dict['truck_link_flow'] = [m_truck]
# data_dict['car_link_tt'] = [m_spd_car]
# data_dict['truck_link_tt'] = [m_spd_truck]
config = dict()
config['use_car_link_flow'] = True
config['use_truck_link_flow'] = False
config['use_car_link_tt'] = False
config['use_truck_link_tt'] = False
config['car_count_agg'] = True
config['truck_count_agg'] = True
config['link_car_flow_weight'] = 1
config['link_truck_flow_weight'] = 1
config['link_tt_weight'] = 1
config['num_data'] = 1
config['observed_links'] = observed_link_list
config['paths_list'] = range(nb.config.config_dict['FIXED']['num_path'])
nb.config
dode = MCDODE(nb, config)
dode.add_data(data_dict)
(car_flow, truck_flow) = dode.init_path_flow(car_scale = 10, truck_scale = 1)
dode.estimate_path_flow(max_epoch = 10, car_init_scale = 100, truck_init_scale = 10, store_folder = '.')
from scipy.sparse import coo_matrix
| src/examples/mcDODE/DODE_7link.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
ds = pd.read_csv('data1.csv')
ds.head()
X = ds.iloc[:, 1:-1].values
y_temp = ds.iloc[:, -1].values
# +
Y = []
for i in y_temp:
if i == "M":
Y.append(1)
elif i == "B":
Y.append(0)
Y = np.array(Y).reshape(-1, 1)
# -
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.05, random_state = 101)
# +
# from sklearn.preprocessing import StandardScaler
# ScalerX = StandardScaler()
# ScalerY = StandardScaler()
# from sklearn.preprocessing import RobustScaler
# ScalerX = RobustScaler()
# ScalerY = RobustScaler()
from sklearn.preprocessing import MinMaxScaler
ScalerX = MinMaxScaler()
# ScalerY = MinMaxScaler()
# -
ScalerX.fit(X)
X_data = ScalerX.transform(X)
X_train = ScalerX.transform(X_train)
X_test = ScalerX.transform(X_test)
np.max(X_train)
def train(X, Y, lr = 0.05, epochs = 100):
W = np.random.normal(0, 1, 30).reshape((30, 1))
b = np.random.normal(0, 1, 1).reshape((1, 1))
for e in range(epochs):
Z = np.dot(X, W) + b
y_ = 1/(1 + np.exp(-Z))
# y_ = 0.78
loss = np.mean(-(Y * np.log(y_) + (1 - Y) * np.log(1 - y_)))
err = (Y - y_)
for i in range(len(W)):
W[i] = W[i] - (lr * np.mean(X[:, i].reshape(-1, 1) * -err))
b = b - (lr * np.mean(-err))
if e % 200 == 0:
print("epoch ", e, "loss: ", loss)
return W, b
W, b = train(X_train, Y_train, 0.5, 7500)
def test(X, Y, W, b):
Z = np.dot(X, W) + b
y_ = 1/(1 + np.exp(-Z))
y_ = [1 if i >= 0.5 else 0 for i in y_]
err = (Y - y_)
print("Err: ", np.sum(err)/len(Y))
test(X_test, Y_test, W, b)
W
b
y_pred = np.dot(X_data, W) + b
Z_pred = 1/(1 + np.exp(-y_pred))
Z_pred = np.array([1 if i >= 0.5 else 0 for i in Z_pred]).reshape(-1, 1)
print("Accuracy: ", 1 - np.mean(np.abs(Z_pred - Y)))
| Projects/.ipynb_checkpoints/Cancer_Prediction-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Training
# +
dataset_filepattern = 'data/*.bin'
_SQUARE_SIZE = 25
_CROP_SIZE = 24 # Divisible by 8.
_MARGIN = (_CROP_SIZE / 2) + 1
_BLUR = 1
# +
# Read data.
import glob
import msgpack
import numpy as np
dataset = []
dataset_golden = []
filenames = glob.glob(dataset_filepattern)
for filename in filenames:
unpacker = msgpack.Unpacker(open(filename, 'rb'))
for img, gt in unpacker:
blur = cv2.GaussianBlur(np.uint8(img), (_BLUR, _BLUR), 0)
if filename.endswith('golden.bin'):
dataset_golden.append((blur, np.int8(gt)))
else:
dataset.append((blur, np.int8(gt)))
print 'Loaded %d boards from %d files' % (len(dataset), len(filenames))
print 'Loaded %d golden boards from %d files' % (len(dataset_golden), len(filenames))
# +
# Show the first board.
import matplotlib.pylab as plt
plt.imshow(dataset[0][0], cmap='gray', vmin=0, vmax=255, interpolation='none')
plt.axis('off')
plt.show()
# +
# Crop relevant regions.
def CropLocalRegion(img, i, j, offset_x=0, offset_y=0):
x = _MARGIN + i * _SQUARE_SIZE - _CROP_SIZE / 2 + offset_x
y = _MARGIN + j * _SQUARE_SIZE - _CROP_SIZE / 2 + offset_y
if (x < 0 or x + _CROP_SIZE >= img.shape[1] or
y < 0 or y + _CROP_SIZE >= img.shape[0]):
return None
return img[y:y + _CROP_SIZE, x:x + _CROP_SIZE]
def BuildImageData(ds, jitter=0):
imgs = []
lbls = []
for img, gt in ds:
for i in range(gt.shape[0]):
for j in range(gt.shape[1]):
for k in range(-jitter, jitter + 1, 1):
for l in range(-jitter, jitter + 1, 1):
crop = CropLocalRegion(img, i, j, k, l)
if crop is not None:
imgs.append(crop)
lbls.append(gt[i][j])
lbls = np.array(lbls)
imgs = np.stack(imgs, axis=0)
return imgs, lbls
images, labels = BuildImageData(dataset, jitter=1)
images_golden, labels_golden = BuildImageData(dataset_golden)
print 'Generated %d samples' % len(labels)
print 'Generated %d golden samples' % len(labels_golden)
# +
# Show a few samples for each class.
import matplotlib.pylab as plt
num_samples = 10
plt.figure(figsize=(num_samples, 3))
for label in (0, 1, 2):
indices = (labels == label)
print 'There are %d samples for class %d.' % (np.sum(indices), label)
sample_images = images[indices][:num_samples]
for i, img in enumerate(sample_images):
plt.subplot(3, num_samples, label * num_samples + i + 1)
plt.imshow(img, cmap='gray', interpolation='none')
plt.axis('off')
plt.show()
# +
# Build features.
import cv2
def ComputeHoG(img):
gx = cv2.Sobel(img, cv2.CV_32F, 1, 0)
gy = cv2.Sobel(img, cv2.CV_32F, 0, 1)
mag, ang = cv2.cartToPolar(gx, gy)
bin_n = 16 # Number of orientation bins.
bin = np.int32(bin_n * ang / (2 * np.pi))
bin_cells = []
mag_cells = []
cellx = celly = 8
for i in range(img.shape[0] / celly):
for j in range(img.shape[1] / cellx):
bin_cells.append(bin[i * celly:i * celly + celly, j * cellx:j * cellx + cellx])
mag_cells.append(mag[i * celly:i * celly + celly, j * cellx:j * cellx + cellx])
hists = [np.bincount(b.ravel(), m.ravel(), bin_n) for b, m in zip(bin_cells, mag_cells)]
hist = np.hstack(hists)
# Transform to Hellinger kernel.
eps = 1e-7
hist /= hist.sum() + eps
hist = np.sqrt(hist)
hist /= np.linalg.norm(hist) + eps
return hist.astype(np.float32, copy=False)
def ComputeLBP(img):
v = ((1<<7) * (img[0:-2,0:-2] >= img[1:-1,1:-1]) +
(1<<6) * (img[0:-2,1:-1] >= img[1:-1,1:-1]) +
(1<<5) * (img[0:-2,2:] >= img[1:-1,1:-1]) +
(1<<4) * (img[1:-1,2:] >= img[1:-1,1:-1]) +
(1<<3) * (img[2:,2:] >= img[1:-1,1:-1]) +
(1<<2) * (img[2:,1:-1] >= img[1:-1,1:-1]) +
(1<<1) * (img[2:,:-2] >= img[1:-1,1:-1]) +
(1<<0) * (img[1:-1,:-2] >= img[1:-1,1:-1]))
bin_n = 256
hist = np.bincount(v.ravel(), minlength=bin_n)
hist = hist.astype(np.float32, copy=False)
eps = 1e-7
hist /= hist.sum() + eps
hist = np.sqrt(hist)
hist /= np.linalg.norm(hist) + eps
return hist
data = []
for img in images:
data.append(ComputeHoG(img))
data = np.array(data)
data_golden = []
for img in images_golden:
data_golden.append(ComputeHoG(img))
data_golden = np.array(data_golden)
print 'Generated data with shape:', data.shape
print 'Generated golden data with shape:', data_golden.shape
# +
# Train empty vs. rest.
from sklearn import svm
from sklearn import model_selection
new_labels = labels.copy()
new_labels[new_labels != 0] = 1
X_train, X_test, y_train, y_test = model_selection.train_test_split(data, new_labels, test_size=0.2, stratify=labels)
clf = svm.SVC(kernel='linear', class_weight='balanced')
clf.fit(X_train, y_train)
print 'Done'
# +
from sklearn import metrics
# Test.
fpr, tpr, _ = metrics.roc_curve(y_test, clf.decision_function(X_test))
auc = metrics.auc(fpr, tpr)
plt.plot(fpr, tpr, color='darkorange', lw=2, label='Test (area = %0.3f)' % auc)
# Train.
fpr, tpr, _ = metrics.roc_curve(y_train, clf.decision_function(X_train))
auc = metrics.auc(fpr, tpr)
plt.plot(fpr, tpr, color='gray', lw=2, label='Train (area = %0.3f)' % auc)
# Golden.
new_labels = labels_golden.copy()
new_labels[new_labels != 0] = 1
fpr, tpr, _ = metrics.roc_curve(new_labels, clf.decision_function(data_golden))
auc = metrics.auc(fpr, tpr)
plt.plot(fpr, tpr, color='blue', lw=2, label='Golden (area = %0.3f)' % auc)
plt.plot([0, 1], [0, 1], color='navy', lw=1, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc='lower right')
plt.show()
# +
# Happy? Save the model.
with open('models/empty_full.bin', 'wb') as fp:
fp.write(msgpack.packb(np.squeeze(clf.coef_).tolist()))
fp.write(msgpack.packb(clf.intercept_[0]))
# +
# Train white vs. black.
new_data = data[labels != 0]
new_labels = labels[labels != 0] - 1 # 0 is Black, 1 is White.
# Split.
X_train, X_test, y_train, y_test = model_selection.train_test_split(new_data, new_labels, test_size=0.2, stratify=new_labels)
clf = svm.SVC(kernel='linear', class_weight='balanced')
clf.fit(X_train, y_train);
print 'Done'
# +
# Test.
fpr, tpr, _ = metrics.roc_curve(y_test, clf.decision_function(X_test))
auc = metrics.auc(fpr, tpr)
plt.plot(fpr, tpr, color='darkorange', lw=2, label='Test (area = %0.3f)' % auc)
# Train.
fpr, tpr, _ = metrics.roc_curve(y_train, clf.decision_function(X_train))
auc = metrics.auc(fpr, tpr)
plt.plot(fpr, tpr, color='gray', lw=2, label='Train (area = %0.3f)' % auc)
# Golden.
new_data = data_golden[labels_golden != 0]
new_labels = labels_golden[labels_golden != 0] - 1 # 0 is Black, 1 is White.
fpr, tpr, _ = metrics.roc_curve(new_labels, clf.decision_function(new_data))
auc = metrics.auc(fpr, tpr)
plt.plot(fpr, tpr, color='blue', lw=2, label='Golden (area = %0.3f)' % auc)
plt.plot([0, 1], [0, 1], color='navy', lw=1, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc='lower right')
plt.show()
# +
# Happy? Save the model.
with open('models/black_white.bin', 'wb') as fp:
fp.write(msgpack.packb(np.squeeze(clf.coef_).tolist()))
fp.write(msgpack.packb(clf.intercept_[0]))
# -
| train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. Correct Syntax to read image in MATLAB in current folder
# ##### Ans: var_image = imread('my_image.jpg')
# #### 2. Select the correct option to crop top-left 50x50 section from the image read below.
#
# #### var_image = imread('my_image.jpg')
# ##### Ans: cropped_section = var_image(1:50,1:50)
# #### 3. What is initial data type of the image you read through imread function of MATLAB?
# ##### Ans: uint8
# #### 4. I1 = imread('my_image.jpg')
#
# #### I2 = im2double(I1)
# ##### Ans:
# - The array dimensions remain same
# - Converts the image from uint8 to double format
# - Scales the image intensity values from 0 to 1
# #### 5. Select the options which assigns height and width of an image correctly in MATLAB.
#
# #### var_image = imread('my_image.jpg')
# ##### Ans:
# - [height,width] = size(var_image );
# - image_dimension = size(var_image );
# - height = image_dimension(1)
# - width = image_dimension(2)
| Week 1/MATLAB_Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="kJJVSI8EMG93"
import pandas as pd
import sklearn as sk
# + [markdown] id="JwkxNXiXcmvH"
# Substitua `dataset_link` pelo dataset de sua escolha
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="wuj-QqY2mpt7" executionInfo={"status": "ok", "timestamp": 1606413324049, "user_tz": 180, "elapsed": 1912, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13898410217834619466"}} outputId="2bcd56e5-05fb-427e-ae9b-73ba7031be8f"
# Usa o dataset 'breast-cancer-wisconson'
dataset_link = 'http://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/wdbc.data'
# Carrega o dataset para um DataFrame do Pandas
df = pd.read_csv(dataset_link, header = None)
# Renomeia a coluna que será usada para a classificação
df.rename(columns={1: 'class'}, inplace=True)
# Impime as primeiras linhas para verificação
df.head()
# + [markdown] id="JLb4nUpKcs75"
# Remova as colunas que não deseja passando o indice delas para `drop` (use a váriavel boleana e a array para remover as colunas)
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="_4iTa25T3TN6" executionInfo={"status": "ok", "timestamp": 1606413325569, "user_tz": 180, "elapsed": 1029, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13898410217834619466"}} outputId="0882c66c-b210-438f-9ec6-6767202dec5e"
# Marca se os dados devem ser normalizados ou não
drop_col = True
cols_to_drop = [0]
# Remove colunas do DataFrame pois elas não são úteis ao modelo
if drop_col:
df = df.drop(cols_to_drop, axis=1)
# Impime as primeiras linhas para verificação
df.head()
# + [markdown] id="AkBqhCiPc3Rh"
# Separa a coluna que vai servir para a classificação, passe o índice dela considerando as colunas que já fora dropadas
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="5lWwufxgGVb8" executionInfo={"status": "ok", "timestamp": 1606413326773, "user_tz": 180, "elapsed": 814, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13898410217834619466"}} outputId="075b8547-a711-436a-ef4c-b58ff3b97a81"
# Separa a "classe" do resto do DataFrame
y = df[['class']]
# Impime as primeiras linhas para verificação
y.head()
# + [markdown] id="wQ9flQ8GdHf9"
# Remove a coluna que foi separada para ser a "classe"
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="FIb0h0L3snxE" executionInfo={"status": "ok", "timestamp": 1606413328160, "user_tz": 180, "elapsed": 1075, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13898410217834619466"}} outputId="27e91901-5eda-413a-d385-c5df8a23f0fe"
# Remove a coluna "class" que foi separada, mantendo somente as features
X = df.drop(['class'], axis = 1)
# Impime as primeiras linhas para verificação
X.head()
# + [markdown] id="Z6bSS1GWdMDI"
# Faz o pré-processamento dos dados necessários como por exemplo a normalização (marque as variáveis como true para fazer o pré-processamento)
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="F1V7ZnJzFbO3" executionInfo={"status": "ok", "timestamp": 1606413329783, "user_tz": 180, "elapsed": 1027, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13898410217834619466"}} outputId="a68fdfea-7256-48ba-891d-09591857f37b"
from sklearn import preprocessing
# Marca se os dados devem ser normalizados ou não
normalize = True
# Faz a normalização dos dados se necessário
if normalize:
min_max_scaler = preprocessing.MinMaxScaler()
X_normalized = min_max_scaler.fit_transform(X)
X = pd.DataFrame(X_normalized)
# Impime as primeiras linhas para verificação
X.head()
# + [markdown] id="ymK7FEDEdX9p"
# Cria a árvore de decisão e faz o 10-fold cross-validation
# + colab={"base_uri": "https://localhost:8080/"} id="YT3V_tOv8wqN" executionInfo={"status": "ok", "timestamp": 1606413332206, "user_tz": 180, "elapsed": 1558, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13898410217834619466"}} outputId="b9082142-e8f9-40f0-8a36-2e0e42f73699"
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import cross_validate
# Cria a árvore de decisão
dt = DecisionTreeClassifier()
# Faz a 10-fold cross-validation
scores = cross_validate(dt, X, y, cv=10, return_estimator=True)
print("Accuracy: %0.2f (+/- %0.2f)" % (scores['test_score'].mean(), scores['test_score'].std() * 2)) # 95% confidence
# + [markdown] id="I7M1KyTMdcFR"
# Mostra a melhor arvore de decisão na tela, lembre de substituir o nome das classes
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="yn90fnmrOKqi" executionInfo={"status": "ok", "timestamp": 1606416121819, "user_tz": 180, "elapsed": 3410, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13898410217834619466"}} outputId="b873b0a5-da2a-494f-ee2a-9dca2cbe84c7"
import matplotlib.pyplot as plt
from sklearn import tree
# Mostra a melhor árvore de decisão
max_score = scores['test_score'].max()
for i in range(len(scores['test_score'])):
if scores['test_score'][i] == max_score:
best_dt = scores['estimator'][i]
# Exibe a árvore
cn=['benign', 'malignant']
fig, ax = plt.subplots(figsize=(25, 25))
tree.plot_tree(best_dt, class_names=cn, filled = True);
plt.show()
| Trabalho 2/TrabalhoIA2-dataset1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # MCMC Exercises
# ### <NAME>, UC Irvine / LSSTC Data Science Fellows Program
# %pylab inline
# You will need to `pip install` these two packages, if you haven't already.
import emcee
import corner
# ## Introduction
# **This first section is a walk through to introduce you to the terminology and define some useful functions. Read it carefully, evaluate each cell, then move to the following sections where you will be repeating this basic study with variations.**
# We will study one of the simplest models, the straight line (which still has many [subtleties](https://arxiv.org/abs/1008.4686)):
# $$
# f(x) = m x + b
# $$
# This exercise is based on a more [complicated example](http://dan.iel.fm/emcee/current/user/line/) in the [emcee documentation](http://dan.iel.fm/emcee/current/).
# ### Initialization
# First, specify our true parameters. A dictionary is overkill with only two parameters, but is useful when tracking a larger number.
truth = dict(m=-1, b=5)
# Plot the true model over the range [0, 10]:
# +
x_lo, x_hi = 0, 10
x_plot = np.linspace(x_lo, x_hi, 100) # Really only need 2 points here
y_plot = truth['m'] * x_plot + truth['b']
plt.plot(x_plot, y_plot, 'r-', label='Truth')
plt.grid()
plt.legend();
# -
# Next, generate a "toy Monte Carlo" sample of `n_data = 50` observations using the true model:
# +
# Reproducible "randomness"
gen = np.random.RandomState(seed=123)
# Random x values, in increasing order.
n_data = 50
x_data = np.sort(gen.uniform(low=0, high=10, size=n_data))
# True y values without any noise.
y_true = truth['m'] * x_data + truth['b']
# Add Gaussian noise.
rms_noise = 0.5
y_error = np.full(n_data, rms_noise)
y_data = y_true + gen.normal(scale=y_error)
# -
# Plot the generated sample with the model superimposed:
plt.errorbar(x_data, y_data, yerr=y_error, fmt='.k', label='Toy MC')
plt.plot(x_plot, y_plot, 'r-', label='Truth')
plt.grid()
plt.legend();
# ### Least Squares Cross Check
# We chose a linear problem so that we would have an alternate method (weighted least squares linear regression) to calculate the expected answer, for cross checks. For details on this method, see:
#
# https://en.wikipedia.org/wiki/Linear_least_squares_(mathematics)#Weighted_linear_least_squares
#
# The appropriate weights here are "inverse variances":
weights = y_error ** -2
# scikit-learn provides a convenient class for (weighted) linear least squares:
# +
import sklearn.linear_model
linear_model = sklearn.linear_model.LinearRegression()
linear_model.fit(x_data.reshape(-1, 1), y_data, sample_weight=weights)
fit = dict(m=linear_model.coef_[0], b=linear_model.intercept_)
print fit
# -
# The weighted linear least squares method also provides a covariance matrix for the best-fit parameters, but unfortunately sklearn does not calculate this for you. However, we will soon have something even better using MCMC.
# Add this fit to our plot:
y_fit = fit['m'] * x_plot + fit['b']
plt.errorbar(x_data, y_data, yerr=rms_noise, fmt='.k', label='Toy MC')
plt.plot(x_plot, y_plot, 'r-', label='Truth')
plt.plot(x_plot, y_fit, 'b--', label='Fit')
plt.grid()
plt.legend();
# ### Markov Chain Monte Carlo
# Define our priors $P(\theta)$ on the two fit parameters $\theta$ with a function to calculate $logP(\theta)$. The function should return `-np.inf` for any parameter values that are forbidden.
#
# Start with simple "top hat" priors on both parameters:
def log_prior(theta):
m, b = theta
if -5 < m < 0.5 and 0 < b < 10:
return 0 # log(1) = 0
else:
return -np.inf # some parameter is outside its allowed range.
# Defined the likelihood $P(D|\theta)$ of our data $D$ given the parameters $\theta$ with a function that returns $\log P(D|\theta)$:
def log_likelihood(theta):
m, b = theta
# Calculate the predicted value at each x.
y_pred = m * x_data + b
# Calculate the residuals at each x.
dy = y_data - y_pred
# Assume Gaussian errors on each data point.
return -0.5 * np.sum(dy ** 2) / rms_noise ** 2
# Combine the priors and likelihood to calculate the log of the posterior $\log P(\theta|D)$:
def log_posterior(theta):
lp = log_prior(theta)
if np.isfinite(lp):
# Only calculate likelihood if necessary.
lp += log_likelihood(theta)
return lp
# Initialize a Markov chain sampler using the `emcee` package:
ndim = 2 # the number of parameters
nwalkers = 100 # bigger is better
sampler = emcee.EnsembleSampler(nwalkers, ndim, log_posterior)
# Give each "walker" a unique starting point in parameter space. The advice on this in the [docs](http://dan.iel.fm/emcee/current/user/faq/#how-should-i-initialize-the-walkers) is:
# > The best technique seems to be to start in a small ball around the a priori preferred position. Don’t worry, the walkers quickly branch out and explore the rest of the space.
#
# Pick something close but exactly the truth for our a priori preferred position.
a_priori = np.array([-1.1, 5.5])
ball_size = 1e-4
initial = [a_priori + ball_size * gen.normal(size=ndim) for i in range(nwalkers)]
# Generate 500 samples for each walker (and ignore the sampler return value).
sampler.run_mcmc(initial, 500);
# The `sampler` object has a `chain` attribute with our generated chains for each walker:
sampler.chain.shape
# Plot the chains from the first 3 walkers for each parameter. Notice how they all start from the same small ball around our a priori initial point.
plt.plot(sampler.chain[0, :, 0])
plt.plot(sampler.chain[1, :, 0])
plt.plot(sampler.chain[2, :, 0]);
plt.plot(sampler.chain[0, :, 1])
plt.plot(sampler.chain[1, :, 1])
plt.plot(sampler.chain[2, :, 1]);
# Combine all the walkers into a single array of generated samples. You could trim off some initial "burn-in" here, but I don't recommend it (see lecture notes):
samples = sampler.chain[:, :, :].reshape((-1, ndim))
samples.shape
# Consecutive samples are certainly correlated, but the ensemble of all samples should have the correct distribution for inferences. Make a corner plot to see this distribution.
fig = corner.corner(samples, labels=['m', 'b'], truths=[truth['m'], truth['b']])
# Each sample represents a possible model. Pick a few at random to superimpose on our data:
shuffled = samples[gen.permutation(len(samples))]
shuffled.shape
# +
plt.errorbar(x_data, y_data, yerr=rms_noise, fmt='.k', label='Toy MC')
plt.plot(x_plot, y_plot, 'r-', label='Truth')
for m, b in shuffled[:100]:
y_sample = m * x_plot + b
plt.plot(x_plot, y_sample, 'b', alpha=0.05)
plt.grid()
plt.legend();
# -
# ## Exercise: Marginal Distributions and Confidence Limits
# Create a function that uses the `samples` array to plot the distribution of predicted values for $y(x)^8$ given $x$ and prints the (central) 68% and 95% confidence limits on this prediction. (Hint: `np.percentile`)
def marginal(x):
# ...
# Note that you can use this technique to make inferences about any non-linear function of the parameters. I chose $y(x)^8$ here so that the result would have an asymmetric distribution.
# ## Exercise: Known Heteroscedastic Errors
# In the example above, the RMS error on each point is known and the same for each point. In this exercise, you will repeat the whole walk-through study above, but with **known but non-constant (aka heteroscedastic) errors** for each point. Copy each cell into this section then edit it, rather than editing the cells above directly, so you can compare later on.
# For your new errors use:
y_error = gen.uniform(low=0.1, high=1.0, size=n_data)
# Before starting, take a moment to review the results above and predict how they will change...
# ## Exercise: Uncertain Heteroscedastic Errors
# In this exercise, the errors reported for each point are uncertain: the true errors are different from the reported errors by an unknown constant scale factor. This introduces a new parameter, the scale factor, that we will call `s`:
truth = dict(m=-1, b=5, s=1.5)
# Regenerate the data using this larger error, while keeping the reported error the same:
y_data = y_true + gen.normal(scale=y_error * truth['s'])
# The model now has three parameters instead of two. Repeat the previous exercise with this additional parameter. Before starting, take a moment to review your results above and predict how they will change...
# ## Challenge Exercise: PyMC3
# Repeat the walk through above (constant known error) using the [pymc3](http://pymc-devs.github.io/pymc3/) package instead of `emcee`. Be sure to use `pymc3` and not `pymc2` (since they are quite different). You will find that `pymc3` has a different approach that takes some getting used to, but it is very helpful to have both packages in your toolkit.
| Sessions/Session02/Day3/MCMC-Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.neural_network import MLPRegressor
from sklearn import preprocessing as sklpp
import numpy as np
net = MLPRegressor(hidden_layer_sizes=(512,512,), verbose=True)
high_dim_data = np.load('exported-data/tsne_input_training.npy')
low_dim_data = np.load('exported-data/tsne_projected_training.npy')
rng_state = np.random.get_state()
np.random.shuffle(high_dim_data)
np.random.set_state(rng_state)
np.random.shuffle(low_dim_data)
scaler = sklpp.StandardScaler().fit(high_dim_data)
num_train = 15000
net.fit(scaler.transform(high_dim_data[:num_train]), low_dim_data[:num_train])
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.scatter(low_dim_data[:15000,0],low_dim_data[:15000,1],s=2)
ax.set_aspect('equal')
plt.show()
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.scatter(low_dim_data[15000:,0],low_dim_data[15000:,1],s=2)
ax.set_aspect('equal')
plt.show()
# +
foo = net.predict(scaler.transform(high_dim_data[:15000]))
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.scatter(foo[:,0],foo[:,1],s=2)
ax.set_aspect('equal')
plt.show()
# +
foo = net.predict(scaler.transform(high_dim_data[15000:]))
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.scatter(foo[:,0],foo[:,1],s=2)
ax.set_aspect('equal')
plt.show()
# -
epoch_data = np.load('exported-data/training_outputs_epoch_NEW.npy')
import torch
foo = torch.tensor(epoch_data)
foo.shape
bar = foo.transpose(0,1).flatten(start_dim=1,end_dim=2)
# +
foo = net.predict(scaler.transform(bar[0,::10]))
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
length = len(foo)//10
for i in range(10):
ax.scatter(
foo[i*length:(i+1)*length, 0],
foo[i*length:(i+1)*length, 1],
s=2
)
ax.set_aspect('equal')
plt.show()
| OLD_nn-oose.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import random
import numpy as np
from written_test_automation import pre, loc, result, nets, config
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from skimage.filters import sobel
def s(im):
plt.imshow(im,cmap="gray")
plt.show()
# -
# # Net trainer
weight_path = "./written_test_automation/answer_weights.pickle"
append_path = "./answer_images_np/"
folders_label =[(append_path + "0/*",[1,0,0,0,0,0]),
(append_path + "1/*",[0,1,0,0,0,0]),
(append_path + "2/*",[0,0,1,0,0,0]),
(append_path + "3/*",[0,0,0,1,0,0]),
(append_path + "4/*",[0,0,0,0,1,0]),
(append_path + "5/*",[0,0,0,0,0,1])]
paths = []
labels = []
for path,label in folders_label:
new_paths = glob.glob(path)
labels = labels + ([label] * len(new_paths))
paths = paths + new_paths
paths,p, labels,l = train_test_split(paths, labels, test_size=0.01)
print(p)
print(l)
# +
x = []
for path in paths:
x.append([np.load(path)])
x = np.array(x, dtype = np.float64 )
y = np.array(labels, dtype = np.float64)
x_test = []
for path in p:
x_test.append([np.load(path)])
x_test = np.array(x_test, dtype = np.float64 )
y_test = np.array(l, dtype = np.float64)
# -
print(x.shape)
print(x.min())
print(x.max())
print(x.mean())
s(x[0,0])
print(y[0])
net = nets.answer_model()
net.load_weights(weight_path)
# Train
# +
max_size = int(x.shape[0])
batch_size = 50
epochs = 15
i = 0
while True:
i += 1
if i > epochs:
break
print(i)
start = 0
fin = start + batch_size
while True:
if (start > max_size) or (fin > max_size):
break
net.numpy_train(x[start:fin],y[start:fin], epochs=1, print_loss=False)
start += batch_size
fin += batch_size
# -
net.numpy_train(x, y, epochs=10)
# Test
print(np.argmax(net.numpy_forward(x_test), axis=1))
print(np.argmax(y_test, axis=1))
print(np.sum(np.abs(net.numpy_forward(x_test) - y_test)))
net.save_weights(weight_path)
# # Config finder
# +
def pick(ims, to_print):
'''
threshold_kernel, blur_kernel, blob_size
'''
th_ker = random.uniform(0.00001,0.1)
blur_kernel = random.uniform(0.00001,0.1)
blob_size = 0.037#random.uniform(0.00001,0.1)
C = 7 #random.randint(2,7)
config.threshold_kernel_percent = th_ker
config.median_blur_kernel_percent = blur_kernel
config.blob_min_size_percent = blob_size
config.threshold_C = C
print(to_print ,th_ker, blur_kernel, blob_size, C)
try:
for im in ims:
result.predict(im)
print("yes")
return [th_ker,blur_kernel,blob_size, C]
except:
return None
def dir_pick(path, loops = 100):
paths = glob.glob(path)
ims = []
for i in paths:
print(i)
ims.append(pre.imread(i))
configs = []
for i in range(loops):
picked_config = pick(ims, str(i))
if picked_config == None:
pass
else:
configs = configs + [picked_config]
return configs
# -
con = dir_pick("./im/*", 100)
print(np.mean(con, axis=0))
print(np.median(con, axis=0))
con
| trainer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 64-bit (conda)
# name: python3
# ---
import os
import numpy as np
np.random.seed(0)
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import set_config
set_config(display="diagram")
DATA_PATH = os.path.abspath(
r"C:\Users\jan\Dropbox\_Coding\UdemyML\Chapter13_CaseStudies\CaseStudyIncome\adult.xlsx"
)
# ### Dataset
df = pd.read_excel(DATA_PATH)
idx = np.where(df["native-country"] == "Holand-Netherlands")[0]
# +
data = df.to_numpy()
x = data[:, :-1]
x = np.delete(x, idx, axis=0)
y = data[:, -1]
y = np.delete(y, idx, axis=0)
categorical_features = [1, 2, 3, 4, 5, 6, 7, 9]
numerical_features = [0, 8]
print(f"x shape: {x.shape}")
print(f"y shape: {y.shape}")
# -
# ### y-Data
def one_hot(y):
return np.array([0 if val == "<=50K" else 1 for val in y], dtype=np.int32)
# + tags=[]
y = one_hot(y)
# -
# ### Helper
def print_grid_cv_results(grid_result):
print(
f"Best model score: {grid_result.best_score_} "
f"Best model params: {grid_result.best_params_} "
)
means = grid_result.cv_results_["mean_test_score"]
stds = grid_result.cv_results_["std_test_score"]
params = grid_result.cv_results_["params"]
for mean, std, param in zip(means, stds, params):
mean = round(mean, 4)
std = round(std, 4)
print(f"{mean} (+/- {2 * std}) with: {param}")
# ### Sklearn Imports
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OrdinalEncoder
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.3)
# ### Classifier and Params
# +
params = {
"classifier__n_estimators": [50, 100, 200],
"classifier__max_depth": [None, 100, 200]
}
clf = RandomForestClassifier()
# -
# ### Ordinal Features
# +
numeric_transformer = Pipeline(
steps=[
('scaler', StandardScaler())
]
)
categorical_transformer = Pipeline(
steps=[
('ordinal', OrdinalEncoder())
]
)
preprocessor_odinal = ColumnTransformer(
transformers=[
('numeric', numeric_transformer, numerical_features),
('categorical', categorical_transformer, categorical_features)
]
)
# -
preprocessor_odinal
# +
preprocessor_odinal.fit(x_train)
x_train_ordinal = preprocessor_odinal.transform(x_train)
x_test_ordinal = preprocessor_odinal.transform(x_test)
print(f"Shape of odinal data: {x_train_ordinal.shape}")
print(f"Shape of odinal data: {x_test_ordinal.shape}")
# -
pipe_ordinal = Pipeline(
steps=[
('preprocessor_odinal', preprocessor_odinal),
('classifier', clf)
]
)
pipe_ordinal
grid_ordinal = GridSearchCV(pipe_ordinal, params, cv=3)
grid_results_ordinal = grid_ordinal.fit(x_train, y_train)
print_grid_cv_results(grid_results_ordinal)
# ### OneHot Features
# +
numeric_transformer = Pipeline(
steps=[
('scaler', StandardScaler())
]
)
categorical_transformer = Pipeline(
steps=[
('onehot', OneHotEncoder(handle_unknown="ignore", sparse=False))
]
)
preprocessor_onehot = ColumnTransformer(
transformers=[
('numeric', numeric_transformer, numerical_features),
('categorical', categorical_transformer, categorical_features)
]
)
# -
preprocessor_onehot
# +
preprocessor_onehot.fit(x_train)
x_train_onehot = preprocessor_onehot.transform(x_train)
x_test_onehot = preprocessor_onehot.transform(x_test)
print(f"Shape of onehot data: {x_train_onehot.shape}")
print(f"Shape of onehot data: {x_test_onehot.shape}")
# -
pipe_onehot = Pipeline(
steps=[
('preprocessor_onehot', preprocessor_odinal),
('classifier', clf)
]
)
pipe_onehot
grid_onehot = GridSearchCV(pipe_onehot, params, cv=3)
grid_results_onehot = grid_onehot.fit(x_train, y_train)
print_grid_cv_results(grid_results_onehot)
# ### TensorFlow Model
from tensorflow.keras.layers import Activation
from tensorflow.keras.layers import Dense
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import SGD
y_train = y_train.reshape(-1, 1)
y_test = y_test.reshape(-1, 1)
def build_model(input_dim, output_dim):
model = Sequential()
model.add(Dense(units=128, input_dim=input_dim))
model.add(Activation("relu"))
model.add(Dense(units=64))
model.add(Activation("relu"))
model.add(Dense(units=output_dim))
model.add(Activation("sigmoid"))
return model
# ### Neural Network with Ordinal Features
# +
model = build_model(
input_dim=x_test_ordinal.shape[1],
output_dim=y_train.shape[1]
)
model.compile(
loss="binary_crossentropy",
optimizer=SGD(learning_rate=0.001),
metrics=["binary_accuracy"]
)
history_ordinal = model.fit(
x=x_train_ordinal,
y=y_train,
epochs=20,
validation_data=(x_test_ordinal, y_test)
)
# +
val_binary_accuracy = history_ordinal.history["val_binary_accuracy"]
plt.plot(range(len(val_binary_accuracy)), val_binary_accuracy)
plt.show()
# -
# ### Neural Network with OneHot Features
# +
model = build_model(
input_dim=x_train_onehot.shape[1],
output_dim=y_train.shape[1]
)
model.compile(
loss="binary_crossentropy",
optimizer=SGD(learning_rate=0.001),
metrics=["binary_accuracy"]
)
history_onehot = model.fit(
x=x_train_onehot,
y=y_train,
epochs=20,
validation_data=(x_test_onehot, y_test)
)
# +
val_binary_accuracy = history_onehot.history["val_binary_accuracy"]
plt.plot(range(len(val_binary_accuracy)), val_binary_accuracy)
plt.show()
# -
# ### Pass in user-data
# +
pipe_ordinal.fit(x_train, y_train)
score = pipe_ordinal.score(x_test, y_test)
print(f"Score: {score}")
# +
x_sample = [
25,
"Private",
"11th",
"Never-married",
"Machine-op-inspct",
"Own-child",
"Black",
"Male",
40,
"United-States"
]
y_sample = 0
y_pred_sample = pipe_ordinal.predict([x_sample])
print(f"Pred: {y_pred_sample}")
| Chapter13_CaseStudies/CaseStudyIncome/USIncome_model4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('ml')
# language: python
# name: python3
# ---
# SOURCE : [URL](https://www.kaggle.com/karnikakapoor/rain-prediction-ann/notebook)
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import datetime
from sklearn.preprocessing import LabelEncoder
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from sklearn.metrics importconfusion_matrix
from keras.layers import Dense,Dropout
from keras.models import Sequential
from keras import callbacks
from tensorflow.keras.optimizers import Adam
from tensorflow import device
np.random.seed(0)
# -
# ## OBJECT
# - 내일 비가 올지 안올지를 예측하는 예측 모델(ANN)을 만들고자 함
# +
data = pd.read_csv('../data/weatherAUS.csv') # for local
# data = pd.read_csv("drive/MyDrive/Colab Notebooks/weatherAUS.csv") # for google
data.head()
# -
data.info()
data.isna().sum()
# #### check target value
sns.countplot(x = data['RainTomorrow'])
# +
corrmat = data.corr()
cmap = sns.diverging_palette(260,-10,s=50, l=75, n=6, as_cmap=True) # color map
plt.subplots(figsize=(15,15))
sns.heatmap(corrmat,cmap = cmap,annot=True,square=True)
# -
# ## data cleansing
# <br/>
#
# prefer the months and days in a cyclic continuous feature. As, date and time are inherently cyclical
# To let the ANN model know that a feature is cyclical I split it into periodic subsections. Namely, years, months and days
data['Date'].dtype
# +
data_c_date = data.copy()
data_c_date.loc[:,'Date'] = pd.to_datetime(data_c_date['Date'])
print(data_c_date['Date'])
print(data_c_date['Date'].isna().sum())
# -
# - add year, month, day columns
# +
data_a_col = data_c_date.copy()
data_a_col['year'] = data_a_col['Date'].dt.year
data_a_col['month'] = data_a_col['Date'].dt.month
data_a_col['day'] = data_a_col['Date'].dt.day
def encode_sin_cos(df,col_n,max_val):
df[col_n+'_sin'] = np.sin(2*np.pi*df[col_n]/max_val)
df[col_n+'_cos'] = np.cos(2*np.pi*df[col_n]/max_val)
return df
data_sin_cos = encode_sin_cos(data_a_col.copy(),'month',12)
data_sin_cos = encode_sin_cos(data_sin_cos,'day',31)
data_sin_cos[['year','month','day','month_sin','month_cos','day_sin','day_cos']]
# -
# - draw result
# +
subset = data_sin_cos[:400] # over 365, roughly
# month - day distribution
m_d = subset['day'].plot()
m_d.set_title("Distribution Of Days Over Year")
m_d.set_ylabel("Days In month")
m_d.set_xlabel("Days In Year")
# -
# - month cyclic
# +
c_m = sns.scatterplot(x="month_sin",y="month_cos",data=data_sin_cos)
c_m.set_title("Cyclic Encoding of Month")
c_m.set_ylabel("Cosine Encoded Months")
c_m.set_xlabel("Sine Encoded Months")
# -
# - day cyclic
# +
c_m = sns.scatterplot(x="day_sin",y="day_cos",data=data_sin_cos)
c_m.set_title("Cyclic Encoding of Days")
c_m.set_ylabel("Cosine Encoded Days")
c_m.set_xlabel("Sine Encoded Days")
# -
# ### process other type cols (object, float)
# +
# extract column names by type
df = data_sin_cos.copy()
s = df.dtypes == 'object'
str_cols = df.loc[:,df.dtypes == 'object'].columns
float_cols = df.loc[:,df.dtypes == 'float64'].columns
print(str_cols)
print(float_cols)
# +
# process na or null
# 1. str
df_c_str = df.copy()
for i in str_cols:
df_c_str[i].fillna(df_c_str[i].mode()[0],inplace=True)
# 2. float
df_c_float = df_c_str.copy()
for i in float_cols:
df_c_float[i].fillna(df_c_float[i].median(),inplace=True)
df_c_float.info()
# -
df_c_float.isna().sum()
# # Data Preprocessing
# +
# map objects to numeric
df_proc = df_c_float.copy()
label_encoder = LabelEncoder()
for i in str_cols:
df_proc.loc[:,i] = label_encoder.fit_transform(df_proc[i]) # fit then transform, return array
# +
# scaling targets
target = df_proc['RainTomorrow']
feats = df_proc.drop(['RainTomorrow','Date','day','month'],axis=1) # why excludes Date??
cols_orig = feats.columns
sd_scaler = preprocessing.StandardScaler()
feat_proc = pd.DataFrame(sd_scaler.fit_transform(feats),columns = cols_orig)
feat_proc.describe().to_json
# -
# ### check outliers
# boxplot
plt.figure(figsize=(20,10))
sns.boxenplot(data = feat_proc)
plt.xticks(rotation=90)
# ### remove outlier
# - use interquartile range (IQR) => lower acc..
# +
# get IQR
Q_l = feat_proc.quantile(0.25)
Q_r = feat_proc.quantile(0.75)
iqr = Q_r-Q_l
iqr
# +
# cond = ~((feat_proc < Q_l - 1.5*iqr)|(feat_proc > Q_r + 1.5*iqr))
# df_iqr = feat_proc[cond]
feat_proc = feat_proc[(feat_proc["MinTemp"]<2.3)&(feat_proc["MinTemp"]>-2.3)]
feat_proc = feat_proc[(feat_proc["MaxTemp"]<2.3)&(feat_proc["MaxTemp"]>-2)]
feat_proc = feat_proc[(feat_proc["Rainfall"]<4.5)]
feat_proc = feat_proc[(feat_proc["Evaporation"]<2.8)]
feat_proc = feat_proc[(feat_proc["Sunshine"]<2.1)]
feat_proc = feat_proc[(feat_proc["WindGustSpeed"]<4)&(feat_proc["WindGustSpeed"]>-4)]
feat_proc = feat_proc[(feat_proc["WindSpeed9am"]<4)]
feat_proc = feat_proc[(feat_proc["WindSpeed3pm"]<2.5)]
feat_proc = feat_proc[(feat_proc["Humidity9am"]>-3)]
feat_proc = feat_proc[(feat_proc["Humidity3pm"]>-2.2)]
feat_proc = feat_proc[(feat_proc["Pressure9am"]< 2)&(feat_proc["Pressure9am"]>-2.7)]
feat_proc = feat_proc[(feat_proc["Pressure3pm"]< 2)&(feat_proc["Pressure3pm"]>-2.7)]
feat_proc = feat_proc[(feat_proc["Cloud9am"]<1.8)]
feat_proc = feat_proc[(feat_proc["Cloud3pm"]<2)]
feat_proc = feat_proc[(feat_proc["Temp9am"]<2.3)&(feat_proc["Temp9am"]>-2)]
feat_proc = feat_proc[(feat_proc["Temp3pm"]<2.3)&(feat_proc["Temp3pm"]>-2)]
feat_proc['RainTomorrow'] = target
plt.figure(figsize=(20,10))
sns.boxenplot(data = feat_proc)
plt.xticks(rotation=90)
plt.show()
# -
# # MODELING
# +
# set dataset
x = feat_proc.drop(['RainTomorrow'],axis=1)
y = feat_proc['RainTomorrow']
x_train, x_test, y_train, y_test = train_test_split(x,y,test_size=0.2,random_state=42)
# -
# ## design and train model
# +
#Early stopping
early_stopping = callbacks.EarlyStopping(
min_delta=0.001, # minimium amount of change to count as an improvement
patience=20, # how many epochs to wait before stopping
restore_best_weights=True,
)
model = Sequential() # model init
# layers
model.add(Dense(units=32,kernel_initializer='uniform',activation='relu',input_dim=26))
model.add(Dense(units=32,kernel_initializer='uniform',activation='relu'))
model.add(Dense(units=16,kernel_initializer='uniform',activation='relu'))
model.add(Dropout(0.25))
model.add(Dense(units=8,kernel_initializer='uniform',activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(units = 1, kernel_initializer = 'uniform', activation = 'sigmoid'))
# compiling
optimizer = Adam(learning_rate=0.00009)
model.compile(optimizer = optimizer, loss = 'binary_crossentropy', metrics = ['accuracy'])
# training
with device('/device:GPU:0'):
model_fitted = model.fit(x_train, y_train, batch_size = 32, epochs = 150, callbacks=[early_stopping], validation_split=0.2)
# +
df_history = pd.DataFrame(model_fitted.history)
plt.plot(df_history.loc[:, ['loss']], "#BDE2E2", label='Training loss')
plt.plot(df_history.loc[:, ['val_loss']],"#C2C4E2", label='Validation loss')
plt.title('Training and Validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend(loc="best")
# -
# # CONCLUSION
y_pred = model.predict(x_test)
y_pred = (y_pred > 0.5)
# +
# confusion matrix
cmap1 = sns.diverging_palette(260,-10,s=50, l=75, n=5, as_cmap=True)
plt.subplots(figsize=(12,8))
cf_matrix = confusion_matrix(y_test, y_pred)
sns.heatmap(cf_matrix/np.sum(cf_matrix), cmap = cmap1, annot = True, annot_kws = {'size':15})
# -
loss, acc = model.evaluate(x_test,y_test,batch_size=32)
print(loss,acc)
| source/ANN_weather.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zTcd9LtZL28S" colab_type="text"
# ## Lista de Exercício 03 e 04 - Fundamentos de Matemática para Computação.
#
# # Universidade Federal do ABC - UFABC
# ## Centro de Matemática, Computação e Cognição - CMCC
#
# ## Disciplina: Fundamentos de Matemática para a Computação - CCM-007
#
# Responsável: Prof. Dr. <NAME>
#
# Estudantes: <NAME>.
#
# <NAME>, 07 de Março de 2019
#
# ### Primeira Parte do Curso
#
# #### Objetivos
#
# O objetivo dessa lista é explorar e aplicar os conceitos de sala de aula sobre **Interpolação Polinomial**.
#
# Para alcançar esse objetivo a resolução dos exercícios é necessária.
#
#
#
# + [markdown] id="1QJk7TpwQwCP" colab_type="text"
# Bibliotecas empregadas para resolução da Lista 03 e 04
# + id="nS9ls_eTLrzr" colab_type="code" colab={}
import matplotlib.pylab as plt
import sympy as sy
import numpy as np
x = sy.Symbol('x')
plt.style.use("ggplot")
# + [markdown] id="Yqn4ZXgoQ0gN" colab_type="text"
# ### Funções desenvolvidas para Lista 03
# ___
# + id="bFQUVMFEQupn" colab_type="code" colab={}
def coeficientes_newton(x_i,f_x):
#entrada: pares (x, y)
#saida: coeficientes
d = f_x.copy()
for k in range(1,len(f_x)):
for i in range(len(f_x), k, -1):
d[i-1] = (d[i-1]-d[i-2]) / (x_i[i-1] - x_i[i-(k+1)])
return d
# + id="3gYJHiuIQ-Fj" colab_type="code" colab={}
def funcao_polinomial(x_i, f_x):
coeficientes = coeficientes_newton(x_i, f_x)
n = len(x_i) - 1 # grau do polinomio
p = coeficientes[n]
for k in range(1,n+1):
p = coeficientes[n-k] + (x -x_i[n-k])*p
return p
# + [markdown] id="n-PMUYEmv_XU" colab_type="text"
# ___
# + [markdown] id="1F3iQdL6Rece" colab_type="text"
# # Exercício 01.
# ___
# + [markdown] id="6ecYNHuWSemg" colab_type="text"
# ### a. Determinar um valor aproximado para $\sqrt{1.12}$ usando o polinômio de interpolação de Newton sobre três pontos. (escolha pontos próximos a $x=1.12$)
#
# ___
#
# + id="iisVJeWq24bN" colab_type="code" colab={}
x_e1 = [1,1.1,1.15,1.25,1.30]
f_e1 = [1.000, 1.049, 1.072, 1.118, 1.140]
# + id="a6oVBySt3JFw" colab_type="code" colab={}
x_e1[:3]
p_e1 = funcao_polinomial(x_e1[:3].copy(), f_e1[:3].copy())
# + id="dJYEkKtPQum9" colab_type="code" colab={}
p_e1 = lambda x: (-0.199999999999964*x + 0.70999999999996)*(x - 1) + 1.0
# + [markdown] id="hlD-rL0FSAjI" colab_type="text"
# O valor estimado para $\sqrt{1.12}$ é:
# + id="_NDhJ18w39Od" colab_type="code" outputId="4e7dee76-ca5e-4067-c6e7-88cf5ea57e3f" colab={"base_uri": "https://localhost:8080/", "height": 34}
p_e1(1.12)
# + [markdown] id="uy1J1GW3SQg6" colab_type="text"
# Esse valor está com a precisão de $10^{-5}$
# + id="kYiR_2EJSPwZ" colab_type="code" outputId="6427aaf3-0581-4509-ebd5-8c0b48a3ef2e" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.abs(p_e1(1.12) - np.sqrt(1.12))
# + [markdown] id="-Q0lNxKcS1UJ" colab_type="text"
# ___
#
# ### b. Calcular um limitante superior para o erro de aproximação para valores próximos dos pontos de interpolação escolhidos (note que não são igualmente espaçado)
#
# ___
#
# Seja $f(x) = \sqrt{x}$, $f'(x) =\frac{1}{2} x^{-\frac{1}{2}}$, $f^{''}(x) = -\frac{1}{4}x^{-\frac{3}{2}}$, $f^{(3)}(x) =\frac{3}{8}x^{\frac{5}{8}}$
#
#
#
# Seja a função de erro para o polinômio de grau 3.
#
# >$|E_n(x)| = (x-x_0)(x-x_1)(x-x_2) \frac{f^{n+1}(\xi)}{(n+1)!}$
#
# >$|E_n(x)| \leq |(x-1)(x-1.1)(x-1.15) |\cdot \left| \frac{3}{8\cdot 3!} \right|$
#
# >$|E_n(x)| \leq \max|(x-1)(x-1.1)(x-1.15) |\cdot \left| \frac{1}{16} \right|$
#
# Estudando o polinômio temos que:
#
# >$(x-1)(x-1.1)(x-1.15) = -1.265 + 3.515 x - 3.25 x^2 + x^3$
#
# >$\frac{d(-1.265 + 3.515 x - 3.25 x^2 + x^3)}{dx} = 3.515 - 6.5 x + 3 x^2$
#
# >$3.515 - 6.5 x + 3 x^2 = 0 => r_1 = 1.03924, r_2 = 1.12743$
#
# Verificando computacionalmente temos que módulo que maximiza a função é:
#
#
# + id="IQBO_lEkolAx" colab_type="code" outputId="ac7bcfd4-5326-49d0-f403-d71a3d73bec8" colab={"base_uri": "https://localhost:8080/", "height": 34}
f_linha = lambda x: -1.265 + 3.515*x - 3.25*x**2 + x**3
np.abs(f_linha(1.03924)), np.abs(f_linha(1.12743))
# + [markdown] id="MCPJtU1xpB5p" colab_type="text"
# O valor que maximiza a função é: 0.00026407647302506554, logo, o limitante superior é:
# + id="qDwdMin6o7-P" colab_type="code" outputId="8c5c3b4e-1b5e-4958-c718-60e09ebf4349" colab={"base_uri": "https://localhost:8080/", "height": 34}
(0.00026407647302506554/16)
# + [markdown] id="9XvtjYus1Irh" colab_type="text"
# ___
# # Exercício 02.
#
# Mostre que a interpolação de um polinômio de grau $n$ sobre $n+k$ pontos, $k\geq1$, é exata.(Dica: use a fórmula para o erro de interpolação)
# ___
#
#
# Da forma do erro temos que a $(n+1)$-derivada de um polinômio de grau n será 0. Dessa forma, o erro será exato.
#
#
# + [markdown] id="Ud9CK_rOZaPQ" colab_type="text"
# ___
# # Exercício 03
#
# + [markdown] id="D6BlR7GaZfAE" colab_type="text"
# Determine o número de pontos necessários para aproximar a função $f(x) = xe^{3x}$ no intervalo $[0, 0.4]$ com erro inferior a $10^{−7}$ usando pontos igualmente espaçados. (Dica: mostre, usando indução, que $f^{(k)} = (k + 3x)3^{k−1} e^{3x}$, k ≥ 1.)
# ___
#
# ## **Caso base: **
# $k = 1$
#
# $\begin{align}
# f_1(x) & = xe^{3x} \\
# f'_1(x) & = e^{3x} + 3x e^{3x}\\
# \\
# \text{Exatamente como nossa fórmula nos diz.}\\
# \\
# f^{(1)}& = (k + 3x)3^{k−1} e^{3x}\\
# & = (1 + 3x)3^{1-1} e^{3x}\\
# & = (1 + 3x)1\cdot e^{3x}\\
# f^{(1)}& = e^{3x} +3x\cdot e^{3x}
# \end{align}$
#
# Passo base é verdadeiro :)
#
# ## **Para hipótese de indução**
#
# Assumimos que nosso caso $k-1$ é verdade. Ou seja:
#
# $\begin{align}
# f_{(k-1)}(x) & = (k-1+3x)3^{k-2} e^{3x}\\
# \end{align}$
#
#
# ## **Passo de indução**
#
#
# $\begin{align}
# f_{k-1}& =(k-1+3x)3^{k-2} e^{3x} \\
# \text{Ao derivarmos temos então:}\\
# f'_{(k-1)}(x)& = (k + 3x)\quad3^{k-1} e^{3 x} \Box \\
# \end{align}$
#
#
#
# Com a nossa $f^{n+1}$ temos:
#
# > $10^{-7} \leq \frac{h^{n+1}}{4(n+1)} \cdot \max_{t\in[0,0.4]} | f^{n+1}(t) |$
#
# > $10^{-7} \leq \frac{({\frac{0.4}{n}})^{n+1}}{4(n+1)} \cdot(n+1+3\cdot 0.4) 3^{n} e^{3\cdot 0.4} $
#
# + id="_borckAk0Ya-" colab_type="code" outputId="154352be-9bd0-4da9-d698-5d0282bf1b7b" colab={"base_uri": "https://localhost:8080/", "height": 156}
import numpy as np
condicao = True
n=1
while condicao:
print(n,"|",(.830029* 0.4**(n + 1)* (3**n)* (1/n)**(n + 1) *(n + 2.2))/(n + 1))
condicao = (10**(-7) <= ((0.4/n)**(n+1))/(4*(n+1)) * (n +1 + 3* 0.4)*(3**(n))*(np.e**(3*0.4)))
n=n +1
# + [markdown] id="AGn5rUrWbCqt" colab_type="text"
# Dessa forma, nós observamos que o número de pontos necesários é 9. Isso ocorre, pois, temos o erro para a função de grau $n+1$.
# + [markdown] id="4_7JTi0s4TLg" colab_type="text"
# ___
#
# # Exercício 04.
#
# + [markdown] id="ZjfGW35XPdFW" colab_type="text"
# Para gerar uma aproximação para a parte superior do pato, os pontos ilustrados no desenho foram selecionados e são apresentados na tabela que segue.
#
# 
#
# <table>
# <tr>
# <th>x</th>
# <th>0.9</th>
# <th>1.3</th>
# <th>1.9</th>
# <th>2.1</th>
# <th>2.6</th>
# <th>3.0</th>
# <th>3.9</th>
# <th>4.4</th>
# <th>4.7</th>
# <th>5.0</th>
# <th>6.0</th>
# <th>7.0</th>
# <th>8.0</th>
# <th>9.2</th>
# <th>10.5</th>
# <th>11.3</th>
# <th>11.6</th>
# <th>12.0</th>
# <th>12.6</th>
# <th>13.0</th>
# <th>13.3</th>
# </tr>
# <tr>
# <td>f(x)</td>
# <td>1.3</td>
# <td>1.5</td>
# <td>1.85</td>
# <td>2.1</td>
# <td>2.6</td>
# <td>2.7</td>
# <td>2.4</td>
# <td>2.15</td>
# <td>2.05</td>
# <td>2.1</td>
# <td>2.25</td>
# <td>2.3</td>
# <td>2.25</td>
# <td>1.95</td>
# <td>1.4</td>
# <td>0.9</td>
# <td>0.7</td>
# <td>0.6</td>
# <td>0.5</td>
# <td>0.4</td>
# <td>0.25</td>
# </tr>
# </table>
#
# Implemente em Python o polinômio interpolador de Newton para aproximar o desenho da parte superior do pato.
#
# ---
#
# + id="d0HXmowt0Odx" colab_type="code" colab={}
import numpy as np
import matplotlib.pylab as plt
# + id="3sW2rBcMP_d1" colab_type="code" colab={}
x_e4 = [0.9,1.3,1.9,2.1,2.6,3.0,3.9,4.4,4.7,5.0,6.0,7.0,8.0,9.2,10.5,11.3,11.6,12.0,12.6,13.0,13.3]
f_e4 = [1.3,1.5,1.85,2.1,2.6,2.7,2.4,2.15,2.05,2.1,2.25,2.3,2.25,1.95,1.4,0.9,0.7,0.6,0.5,0.4,0.25]
x_e5 = np.arange(np.min(x_e4),np.max(x_e4),0.1)
# + id="fDjjeYDIUhah" colab_type="code" outputId="bef3a335-ca39-42bf-e8a2-5009faf8c99e" colab={"base_uri": "https://localhost:8080/", "height": 54}
funcao_polinomial(x_e4.copy(),f_e4.copy())
# + id="CbPe6-mHVR2D" colab_type="code" colab={}
p = lambda x: (x - 0.9)*((x - 1.3)*((x - 1.9)*((x - 2.1)*((x - 2.6)*((x - 3.0)*((x - 3.9)*((x - 4.4)*((x - 4.7)*((x - 5.0)*((x - 6.0)*((x - 7.0)*((x - 8.0)*((x - 9.2)*((x - 10.5)*((x - 11.3)*((x - 11.6)*((x - 12.0)*((-3.074530780108e-11*x + 6.51542950918716e-10)*(x - 12.6) - 1.98371331929768e-9) + 1.46954842089078e-8) - 9.86036108443615e-8) + 6.08595004676425e-7) - 3.45353446933275e-6) + 1.57981716791882e-5) - 4.29023399481625e-5) - 6.34107592999165e-6) + 0.000572931763659381) - 0.00185867502994866) - 0.00254815041559561) + 0.0387469060492263) - 0.18391194861783) + 0.566835125658653) - 0.906324068088771) + 0.624999999999998) + 0.0833333333333337) + 0.5) + 1.3
# + id="vHy_Mr8SQOSe" colab_type="code" outputId="bf290055-40a4-4aca-8687-9ad610378d0d" colab={"base_uri": "https://localhost:8080/", "height": 595}
fig, ax = plt.subplots(figsize=(20,10))
ax.plot(x_e4, f_e4, 'o', c='r',label='pontos originais')
#ax.plot(x_e4, list(map(p,x_e4)), c='g', marker='+', label='interpolacao por Newton')
ax.plot(x_e5, list(map(p,x_e5)), c='b', marker='+', label='interpolando com mais pontos')
ax.legend(loc='lower left')
plt.show()
# + [markdown] id="MR9GP_yjcGwk" colab_type="text"
# Observamos no comportamento da função o fenômeno de Runge. Em suma, estamos empregando uma função de complexidade muito acima da necessária para descrever o nosso conjunto de dados. Uma das formas para contornar a situação consiste em amostrar o dataset. A técnica escolhida pode ser diversa. Nesse caso, optamos por escolher 1 ponto a cada 4 pontos.
# + id="HEgURHwFcEEZ" colab_type="code" colab={}
x_04_01 = x_e4[:5]
f_04_01 = f_e4[:5]
x_04_02 = x_e4[4:9]
f_04_02 = f_e4[4:9]
x_04_03 = x_e4[8:13]
f_04_03 = f_e4[8:13]
x_04_04 = x_e4[12:17]
f_04_04 = f_e4[12:17]
x_04_05 = x_e4[16:21]
f_04_05 = f_e4[16:21]
p_04_01 = funcao_polinomial(x_04_01,f_04_01)
p_04_02 = funcao_polinomial(x_04_02,f_04_02)
p_04_03 = funcao_polinomial(x_04_03,f_04_03)
p_04_04 = funcao_polinomial(x_04_04,f_04_04)
p_04_05 = funcao_polinomial(x_04_05,f_04_05)
# + id="va2Jfjd-Jw6W" colab_type="code" outputId="ad70c352-d58a-4ede-d11d-b62a2536eee7" colab={"base_uri": "https://localhost:8080/", "height": 104}
print(p_04_01)
print(p_04_02)
print(p_04_03)
print(p_04_04)
print(p_04_05)
# + id="Vzz3PijaKbTf" colab_type="code" colab={}
p_04_01 = lambda x: (x - 0.9)*((x - 1.3)*((-0.906324068088771*x + 2.52828054298642)*(x - 1.9) + 0.0833333333333337) + 0.5) + 1.3
p_04_02 = lambda x: (x - 2.6)*((x - 3.0)*((0.00448897507720958*x + 0.163398692810462)*(x - 3.9) - 0.448717948717949) + 0.25) + 2.6
p_04_03 = lambda x: (x - 4.7)*((x - 5.0)*((0.00489848315935264*x - 0.0504543765413322)*(x - 6.0) - 0.0128205128205137) + 0.166666666666668) + 2.05
p_04_04 = lambda x: (x - 8.0)*((x - 9.2)*((0.00901105067771695*x - 0.10998338081671)*(x - 10.5) - 0.0692307692307691) - 0.25) + 2.25
p_04_05 = lambda x: (x - 11.6)*((x - 12.0)*((-0.053867700926524*x + 0.581232492997194)*(x - 12.6) + 0.083333333333333) - 0.25) + 0.725
# + id="EVc8LQ4SLAZ4" colab_type="code" outputId="545e0b62-99d2-4933-8a3a-6dab4d564c60" colab={"base_uri": "https://localhost:8080/", "height": 612}
fig, ax = plt.subplots(figsize=(20,10))
ax.plot(x_e4, f_e4, 'o', c='r',label='pontos originais')
ax.plot(x_04_01, list(map(p_04_01,x_04_01)), c='b', marker='+',
label='1 divisao')
ax.plot(x_04_02, list(map(p_04_02,x_04_02)), c='g', marker='+',
label='2 divisao')
ax.plot(x_04_03, list(map(p_04_03,x_04_03)), c='y', marker='+',
label='3 divisao')
ax.plot(x_04_04, list(map(p_04_04,x_04_04)), c='turquoise', marker='+',
label='4 divisao')
ax.plot(x_04_05, list(map(p_04_05,x_04_05)), c='azure', marker='+',
label='5 divisao')
ax.legend(loc='lower left')
# + [markdown] id="mmnBIld-fElt" colab_type="text"
# ___
# ## Exercício 5
#
# Refazer o exercício **E4** da aula passada (do pato) usando splines
# ___
#
# + id="y3B3y9_XitST" colab_type="code" colab={}
from scipy.interpolate import interp1d
from scipy.interpolate import CubicSpline
# + id="xpdGz0Pvi92P" colab_type="code" outputId="b0565006-bb74-4747-a75a-3337c5a3e48b" colab={"base_uri": "https://localhost:8080/", "height": 612}
cs = CubicSpline(x=x_e4, y=f_e4)
fig, ax = plt.subplots(figsize=(15,10))
ax.plot(x_e4, f_e4, 'o', label='Dados')
ax.plot(x_e5, list(map(p,x_e5)), c='g', marker='+', label='Interpolação Newton')
ax.plot(x_e5, cs(x_e5), label="Splines Cúbicos", c='r')
ax.legend(loc='lower left')
# + id="3fqI9yjUoho8" colab_type="code" outputId="6e30790e-a116-4436-ea65-a90772b673ff" colab={"base_uri": "https://localhost:8080/", "height": 476}
fig, ax = plt.subplots(figsize=(15,7.5))
ax.plot(x_e4, f_e4, 'o', label='Dados')
ax.plot(x_e5, cs(x_e5), label="Splines Cúbicos")
for i in range(np.shape(cs.c)[0]):
ax.plot(x_e5, cs(x_e5,i), label="S^"+str(i))
ax.legend(loc='best', ncol=2)
| list_03-04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import re
import nltk
from nltk.tokenize import word_tokenize
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
from nltk.stem.wordnet import WordNetLemmatizer
from nltk.probability import FreqDist
from data_preprocesser import preprocessed_data_path
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
# -
df = pd.read_csv(preprocessed_data_path + 'categories_30000/yelp_academic_dataset_review_Auto Repair.csv')
df.head()
from textblob import TextBlob
| AG - Data Tests Algo Sentiment Analysis - YELP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Reading Files
# +
import pandas as pd
df1 = pd.read_csv('file1.csv')
print(df1,'\n')
# The first row becomes the columns header
print(df1['b'])
print(df1['message'])
# -
# Generating column's headers automatically
df2 = pd.read_csv('file1.csv',header=None)
print(df2,'\n')
print(df2[1])
print(df2[4])
# Specifying column's headers
df3 = pd.read_csv('file1.csv',names=['i','j','k','l','m'])
print(df3,'\n')
print(df3['j'])
print(df3['m'],'\n')
# +
# Using one of the columns to index the rows
df4 = pd.read_csv('file1.csv',names=['i','j','k','l','m'],index_col='m')
print(df4,'\n')
print(df4['k']['hello'])
print(df4['k'][['hello','foo']])
# -
# Pandas "read_table" also works, but the sep variable has to be properly settled
dt1 = pd.read_table('file1.csv',sep=',')
print(dt1)
# The default is a white space, which does not work for the content in file1.csv
dt2 = pd.read_table('file1.csv')
print(dt2)
# Columns in file3.txt are separated by white spaces, but the default assumes a single white spaces between columns
dt3 = pd.read_table('file3.txt')
print(dt3)
# The solution to handle multiple white spaces is
dt4 = pd.read_table('file3.txt', header=None, sep='\s+')
print(dt4)
# ## Missing Data
# Missing data
df5 = pd.read_csv('file2.csv',header=None)
print(df5)
# +
# removing columns with all elements = NaN
df6=df5.dropna(axis=1, how='all')
print(df6,'\n')
# removing columns with any element = NaN
df7=df5.dropna(axis=1, how='any')
print(df7,'\n')
# removing rows with at least k NON-NaN
df8 = df5.dropna(thresh=3)
print(df8)
# +
# filling missing values with a given value
df9 = df5.fillna(0)
print(df9,'\n')
# Filling by interpolate
df10 = df5.fillna(method='ffill')
print(df10,'\n')
# +
# filling each column independently
col_values = {0:0, 1:10, 2:20, 3:30, 4:40, 5:50}
df11 = df5.fillna(value=col_values)
print(df11,'\n')
# filling by adding
df12 = df10.add(df5,fill_value=0)
print(df12)
# -
# ## Massaging Data
# +
import numpy as np
dfm1 = pd.DataFrame(np.arange(12).reshape((3,4)),
index=['Ohio','Colorado','New York'],
columns=['one','two','three','four'])
print(dfm1)
# -
# replaces selected values (>7) with -1
dfm1[dfm1>7] = -1
print(dfm1)
# use .loc to access index along axis 0 and .apply to apply a function to all values in the series
print(dfm1.loc['Colorado'].apply(lambda x:x+2))
# ## Combining and Concatenating
dfc1 = pd.DataFrame({'c1':np.arange(5), 'key': ['<KEY>'], 'c2': np.arange(6,11)})
dfc2 = pd.DataFrame({'c3': np.arange(11,15),'key': ['<KEY>']})
print(dfc1,'\n')
print(dfc2)
dfc3 = pd.merge(dfc1,dfc2,on='key',how='inner')
#dfc3 = pd.merge(dfc1,dfc2,on='key',how='outer')
print(dfc3)
# merging based on the row index
dfc3 = pd.merge(dfc1,dfc2,left_index = True, right_index=True, how='outer')
print(dfc3)
dfc4 = pd.concat([dfc1,dfc2])
#dfc4 = pd.concat([dfc1,dfc2],ignore_index=True)
print(dfc4)
dfc5 = dfc1.stack()
print(dfc5)
print(dfc5[:][1],'\n')
print(dfc5[:][1,'c1'])
# ## Ploting and functions
dfp1 = pd.DataFrame(np.arange(40).reshape(8,5))
print(dfp1)
import matplotlib.pyplot as plt
dfp1.plot()
plt.show()
dfp2 = pd.DataFrame(dfp1[0].map(lambda x: np.sin(x)))
dfp2.plot()
plt.show()
dfp3 = pd.concat([dfp2,dfp1.loc[:,1:4]],axis=1)
print(dfp3)
dfp3.plot()
plt.show()
dfp3[0].plot(kind='bar')
plt.show()
| PDS_jupyter/Labs/L9 - Pandas II.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/R-Mosolov/sociology-scientometric-analysis/blob/main/analyze_article_texts.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="aCp-HDEDU_C3" colab={"base_uri": "https://localhost:8080/", "height": 37} outputId="4ef25fea-1753-4ed7-b6d8-fd973a868be8"
'''
ABOUT THE MODULE:
This module contains logic to analyze about 50000 article texts
from "CyberLeninka" as a Russian scientific database
'''
'''
NOTATIONS:
– df – a dataframe/dataset, or an array of initial data.
'''
# + id="Nr8B4aa7Vje6"
from bs4 import BeautifulSoup
import urllib.request
import requests
import csv
import pandas as pd
import random
import time
import os
import numpy as np
import statistics
# # !pip uninstall pandas_profiling
# # !pip install pandas_profiling
from pandas_profiling import ProfileReport
# # !pip uninstall nltk
# # !pip install nltk
import nltk
# nltk.download('punkt')
from nltk.tokenize import word_tokenize
from nltk.probability import FreqDist
# + id="bES3n-xuVtbW" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="a5c7a5a7-578a-475a-b619-b1a2acedf742"
'''
Initialize the dataframe
'''
df = pd.read_csv(
'/content/drive/MyDrive/Science/Datasets/cyberleninka-sociology-articles/cyberleninka-sociology-articles'
+ '__2-0_articles-content/integrated-dataframes/integrated-dataframe__actuality-09-01-2021-2.csv'
)
# + id="99ARLXbHVwt0"
'''
Open the dataframe
'''
df
# + id="T47wRMFSbAOs"
'''
Create decorder for texts from string to array type
'''
firstArticleText = df['article_text'][0]
def transformFromStrToArr(textAsStr):
textWithoutOuterWrapper = textAsStr.replace(']', '').replace('[', '')
textAsArr = textWithoutOuterWrapper.replace("'", '').split(', ')
return textAsArr
transformFromStrToArr(firstArticleText)
# + id="xWe1HKfrWdQ2"
'''
Open first article text
'''
firstArticleText
# + id="IOl4AszAeRIn"
'''
Restore text structure of an one random text
'''
textAsArr = transformFromStrToArr(firstArticleText)
SPACE = ' '
HYPHEN_1 = '-'
HYPHEN_2 = '–'
HYPHEN_3 = '—'
'''
Additional functions
'''
def isLastTextChunk(idx, textAsArr):
idx == len(textAsArr) - 1
def isEndsOnHyphen(textAsStr):
lastChunkSymbol = textAsArr[len(textAsArr) - 1]
if (
lastChunkSymbol == HYPHEN_1
or lastChunkSymbol == HYPHEN_2
or lastChunkSymbol == HYPHEN_3
):
return true
'''
Main function
'''
def restoreOneArticleText(textAsArr):
restoredText = ''
for idx, textChunk in enumerate(textAsArr):
# Pass last text chunk because it doesn't require handling for chunk end
if isLastTextChunk(idx, textChunk):
restoredText += textChunk
if isEndsOnHyphen(textChunk):
textChunkWithoutHyphen = textChunk[:-1]
restoredText += textChunkWithoutHyphen
# Handle other universal cases
else:
restoredText += textChunk + SPACE
return restoredText
restoreOneArticleText(textAsArr)
# + id="NIQhdBhIm9mg"
'''
Integrate all articles texts in one text
'''
all_article_texts = df['article_text']
integrated_text = '' # This variable integrates all 46957 articles texts in same one
SPACE = ' '
file_to_write = open(
'/content/drive/MyDrive/Science/Datasets/cyberleninka-sociology-articles/cyberleninka-sociology-articles'
+ '__2-0_articles-content/integrated-dataframes/all-article-texts-in-one-text__actuality-25-02-2021.txt',
'a'
)
for article_text in all_article_texts:
try:
textAsArr = transformFromStrToArr(article_text)
restoredText = restoreOneArticleText(textAsArr)
file_to_write.write(restoredText + SPACE)
file_to_write.close()
except:
pass
integrated_text
| analyze_article_texts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="i7AXtRi8jt2G"
# import imageio
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import scipy
from scipy import misc
import glob
import cv2
from numpy.random import seed
seed(1)
import matplotlib
matplotlib.use('Agg')
import os
import h5py
import scipy.io as sio
import gc
from keras.models import load_model, Model, Sequential
from keras.layers import (Input, Conv2D, MaxPooling2D, Flatten,
Activation, Dense, Dropout, ZeroPadding2D)
from keras.optimizers import Adam
from keras.layers.normalization import BatchNormalization
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras import backend as K
from sklearn.metrics import confusion_matrix, accuracy_score
from sklearn.model_selection import KFold, StratifiedShuffleSplit
from keras.layers.advanced_activations import ELU
import openvino
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="T4n9-o4NlaxT" outputId="5730f8fa-b504-4fd3-9087-c31893756406"
# from google.colab import drive
# drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="Kx26dn2MqevQ" outputId="5229fc5b-03c6-468b-acc4-b9e1b749685c"
#import os
#os.chdir("/content/drive")
#path = "/content/drive/My Drive/fall_detection"
#os.chdir(path)
#os.listdir(path)
# + colab={} colab_type="code" id="SyQZDgn7jt2M"
## fun
def rgb2grey(pic):
size = np.shape(pic)
pic = np.array(pic)
W = size[0]
H = size[1]
rim = pic[:,:,0]
gim = pic[:,:,1]
bim = pic[:,:,2]
return 0.299 * rim + 0.587 * gim + 0.114 * bim
# + [markdown] colab_type="text" id="p1NiE028puAv"
# **Test whether the image is correctly loaded.**
# + colab={"base_uri": "https://localhost:8080/", "height": 287} colab_type="code" id="hGfdUv6Kjt2P" outputId="757dd4a3-1d83-47d5-a6f4-7410512f01de"
# test
# %matplotlib inline
image_size=244
frame = 10
strid = 3
I = misc.imread("data/UR/fall-01-cam0-rgb/fall-01-cam0-rgb-001.png")
I = rgb2grey(I)
I = cv2.resize(I, (image_size,image_size))
# I.resize((image_size,image_size))
plt.imshow(I)
# + colab={} colab_type="code" id="EMy4aa2uHZUC"
file_path = 'data/UR'
fall_period = np.array([[93,135],[55,85],[161,200],[27,71],[92,140],[31,70],[96,127],[40,82],[134,175],[58,85],[61,105],[36,85],[54,78],[28,51],[36,71]])
#floders
im_folders1 = glob.glob('data/UR//fall*')
im_folders2 = glob.glob('data/UR//adl*')
im_folders3 = glob.glob('data/UR//*')
im_folders1.sort()
np.random.shuffle(im_folders3)
print(im_folders1)
print(im_folders2)
print(im_folders3)
final_data = []
final_label=[]
period_idx =0
# + [markdown] colab_type="text" id="eVpflKahk_dp"
# **Now, we start to upload the images from each file, and transfer them into optical flow.**
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="r0gLO9YOjt2U" outputId="79b28235-9d12-428c-fe02-1a9a1e9cd952"
# loop all the files
for i,folder in enumerate(im_folders1):
print (i,folder)
pics = glob.glob(folder+'//*')
image_set = []
for pic in pics:
I = misc.imread(pic)
I = rgb2grey(I)
I = cv2.resize(I, (image_size,image_size))
image_set.append(I)
image_set = np.array(image_set)
print("imgs size:",np.shape(image_set))
## cal the optical flow:
opflowx = []
opflowy = []
for k in range(len(image_set)-1):
flow = cv2.calcOpticalFlowFarneback(image_set[k] ,image_set[k+1], None, 0.5, 3, 15, 3, 5, 1.2, 0)
opflowx.append(flow[:,:,0])
opflowy.append(flow[:,:,1])
del image_set
opflow_set = []
for j in range(0,len(opflowx)-frame,strid):
xy = opflowx[j:j+frame]+opflowy[j:j+frame]
opflow_set.append(xy)
del opflowx,opflowy
final_data += opflow_set
opflow_set = np.array(opflow_set)
print("stacked size:",np.shape(opflow_set))
#opflow_set = [xxxxxxx(frame),yyyyyyy(frame),xxxxxxx,yyyyyyy...]
temp_label =[]
for patch_idx in range(0,len(opflow_set)):
#print("patch size:",np.shape(datax[patch_idx]))
patch =[]
patch = opflow_set[patch_idx]
if (folder in im_folders1 and patch_idx >= (fall_period[period_idx][0]-5) and
patch_idx <= (fall_period[period_idx][1]-5)): #label fall
temp_label.append(1)
else:
temp_label.append(0)
del opflow_set
final_label.append(temp_label)
print("label size:",np.shape(temp_label))
if folder in im_folders1:
period_idx +=1
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="fPK9qCn77gfX" outputId="1fecc2d1-abee-4c2f-c066-7959ca36b35b"
# above is label porcess
final_data = np.array(final_data)
#final_label = sum(final_label, []) ## flatten
print("finallebel",np.shape(final_label))
print("finaldata",np.shape(final_data))
# + [markdown] colab_type="text" id="z9ZN6zLcqDQS"
# **divide the data set into training set and test set**
# + colab={"base_uri": "https://localhost:8080/", "height": 87} colab_type="code" id="6o9tQPtojt2X" outputId="b736b59d-270d-4647-d8f6-8d7954e75750"
final_data_size = np.shape(final_label)
final_data_size = final_data_size[0]
trainx = final_data[:int(final_data_size*0.7)]
trainy = final_label[:int(final_data_size*0.7)]
testx = final_data[int(final_data_size*0.7):]
testy = final_label[int(final_data_size*0.7):]
trainx = trainx.transpose(0,3,2,1) #
testx = testx.transpose(0,3,2,1) #
print('trainx shape:',np.shape(trainx))
print('trainy shape:',np.shape(trainy))
print('testx shape:',np.shape(testx))
print('testny shape:',np.shape(testy))
# + [markdown] colab_type="text" id="IFw2ejJCqMPU"
# **Build the vgg model**
# + colab={} colab_type="code" id="kX0p_ZNEjt2a"
# hyper parameters
learning_rate = 0.0001
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="WfQPCaNJjt2e" outputId="883573d5-d1d7-4a2e-eb62-dd49b165283e"
num_features = num_features = 4096
model = Sequential()
model.add(ZeroPadding2D((1, 1), input_shape=(image_size, image_size, frame*2)))
model.add(Conv2D(64, (3, 3), activation='relu', name='conv1_1'))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(64, (3, 3), activation='relu', name='conv1_2'))
model.add(MaxPooling2D((2, 2), strides=(2, 2)))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(128, (3, 3), activation='relu', name='conv2_1'))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(128, (3, 3), activation='relu', name='conv2_2'))
model.add(MaxPooling2D((2, 2), strides=(2, 2)))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(256, (3, 3), activation='relu', name='conv3_1'))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(256, (3, 3), activation='relu', name='conv3_2'))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(256, (3, 3), activation='relu', name='conv3_3'))
model.add(MaxPooling2D((2, 2), strides=(2, 2)))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(512, (3, 3), activation='relu', name='conv4_1'))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(512, (3, 3), activation='relu', name='conv4_2'))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(512, (3, 3), activation='relu', name='conv4_3'))
model.add(MaxPooling2D((2, 2), strides=(2, 2)))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(512, (3, 3), activation='relu', name='conv5_1'))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(512, (3, 3), activation='relu', name='conv5_2'))
model.add(ZeroPadding2D((1, 1)))
model.add(Conv2D(512, (3, 3), activation='relu', name='conv5_3'))
model.add(MaxPooling2D((2, 2), strides=(2, 2)))
model.add(Flatten())
model.add(Dense(num_features, name='fc6', kernel_initializer='glorot_uniform'))
model.add(BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001))
model.add(Activation('relu'))
model.add(Dropout(0.9))
model.add(Dense(4096, name='fc2', kernel_initializer='glorot_uniform'))
model.add(BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001))
model.add(Activation('relu'))
model.add(Dropout(0.8))
model.add( Dense(1, name='predictions',kernel_initializer='glorot_uniform'))
model.add(Activation('sigmoid',name = 'Sigmoid'))
adam = Adam(lr=learning_rate, beta_1=0.9, beta_2=0.999,
epsilon=1e-08)
model.compile(optimizer=adam, loss='binary_crossentropy',
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/", "height": 416} colab_type="code" id="5rLhQp7Sjt2h" outputId="9cd27aeb-09ff-4cd9-ae21-283f817d74c6"
#from keras.utils import to_categorical
#print(np.shape(trainx))
#trainy = to_categorical(trainy) # ont-hot
#testy = to_categorical(testy)
model.fit(trainx,trainy,validation_data=(testx,testy),epochs=1)
#time()
model.evaluate(testx,testy)
# -
# ### now we start to import the model to openvino
## The next two cells is for save the model to h5 and convert it to .pb file, which the openvino needs
model.save("../saved.h5")
from keras.models import load_model
import tensorflow as tf
import os
import os.path as osp
import keras
#路径参数
input_path = '..'
weight_file = 'saved.h5'
weight_file_path = osp.join(input_path,weight_file)
output_graph_name = weight_file[:-3] + '.pb'
#转换函数
def h5_to_pb(h5_model,output_dir,model_name,out_prefix = "output_",log_tensorboard = True):
if osp.exists(output_dir) == False:
os.mkdir(output_dir)
out_nodes = []
for i in range(len(h5_model.outputs)):
out_nodes.append(out_prefix + str(i + 1))
tf.identity(h5_model.output[i],out_prefix + str(i + 1))
sess = tf.compat.v1.keras.backend.get_session()
from tensorflow.python.framework import graph_util,graph_io
init_graph = sess.graph.as_graph_def()
print(h5_model.outputs)
main_graph = graph_util.convert_variables_to_constants(sess,init_graph,out_nodes)
graph_io.write_graph(main_graph,output_dir,name = model_name,as_text = False)
if log_tensorboard:
from tensorflow.python.tools import import_pb_to_tensorboard
import_pb_to_tensorboard.import_to_tensorboard(osp.join(output_dir,model_name),output_dir)
#输出路径
output_dir = osp.join(os.getcwd(),"trans_model")
#加载模型
h5_model = load_model(weight_file_path)
h5_to_pb(h5_model,output_dir = output_dir,model_name = output_graph_name)
print('model saved')
## this is for comparison, to make sure that the out put is the same on or not on openvino
y = model.predict(testx)
print(y)
# ## Here we need to run the mo.py to convert the file to the IR model
# ### after that, the next cells load the generated ir model and output the result
from PIL import Image
import numpy as np
from openvino import inference_engine as ie
from openvino.inference_engine import IENetwork, IEPlugin
# except Exception as e:
# exception_type = type(e).__name__
# print("The following error happened while importing Python API module:\n[ {} ] {}".format(exception_type, e))
# sys.exit(1)
# Plugin initialization for specified device and load extensions library if specified.
plugin_dir = None
model_xml = './ir_model/saved.xml'
model_bin = './ir_model/saved.bin'
# Devices: GPU (intel), CPU, MYRIAD
plugin = IEPlugin("CPU", plugin_dirs=plugin_dir)
# Read IR
net = IENetwork(model=model_xml, weights=model_bin)
print(len(net.inputs.keys()) == 1)
print(len(net.outputs) == 1)
input_blob = next(iter(net.inputs))
out_blob = next(iter(net.outputs))
# Load network to the plugin
exec_net = plugin.load(network=net)
del net
# Run inference
predicted = []
fileName = 'data/UR/fall-01-cam0-rgb/fall-01-cam0-rgb-001.png'
for processedImg in testx:
# image, processedImg, imagePath = pre_process_image(fileName)
# processedImg = testx[0]
processedImg = processedImg.transpose((2, 0, 1))
processedImg = np.array([processedImg])
res = exec_net.infer(inputs={input_blob: processedImg})
# Access the results and get the index of the highest confidence score
output_node_name = list(res.keys())[-1]
res = res[output_node_name]
# idx = (res[0] >= 0.5)
predicted.append(res)
# print(res)
## acc:
print(predicted)
# #### as we can see the IR model's out put is the same
# +
### This cell is discarded
# force reset ipython namespaces
import tensorflow as tf
from tensorflow.python.framework import graph_io
from tensorflow.keras.models import load_model
# Clear any previous session.
tf.keras.backend.clear_session()
save_pb_dir = '..'
model_fname = '../saved.h5'
def freeze_graph(graph, session, output, save_pb_dir='.', save_pb_name='frozen_model.pb', save_pb_as_text=False):
with graph.as_default():
graphdef_inf = tf.graph_util.remove_training_nodes(graph.as_graph_def())
graphdef_frozen = tf.graph_util.convert_variables_to_constants(session, graphdef_inf, output)
graph_io.write_graph(graphdef_frozen, save_pb_dir, save_pb_name, as_text=save_pb_as_text)
return graphdef_frozen
# This line must be executed before loading Keras model.
tf.keras.backend.set_learning_phase(0)
model = load_model(model_fname)
session = tf.keras.backend.get_session()
INPUT_NODE = [t.op.name for t in model.inputs]
OUTPUT_NODE = [t.op.name for t in model.outputs]
print(INPUT_NODE, OUTPUT_NODE)
frozen_graph = freeze_graph(session.graph, session, [out.op.name for out in model.outputs], save_pb_dir=save_pb_dir)
# -
| OPENVINO/vgg_fall_detection_openvino.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <img src="../../images/banners/python-modules.png" width="600"/>
# # <img src="../../images/logos/python.png" width="23"/> YAML
#
# ## <img src="../../images/logos/toc.png" width="20"/> Table of Contents
# * [YAML Tutorial Quick Start: A Simple File](#yaml_tutorial_quick_start:_a_simple_file)
# * [Installation](#installation)
# * [Read YAML](#read_yaml)
# * [Introduction](#introduction)
# * [Rules for Creating YAML file](#rules_for_creating_yaml_file)
# * [Indentation of YAML](#indentation_of_yaml)
# * [Basics Components](#basics_components)
# * [Conventional Block Format](#conventional_block_format)
# * [Inline Format](#inline_format)
# * [Folded Text](#folded_text)
# * [YAML Syntax and format](#yaml_syntax_and_format)
# * [String](#string)
# * [Short texts](#short_texts)
# * [Long text](#long_text)
# * [Numbers](#numbers)
# * [Boolean](#boolean)
# * [Collections and Structures](#collections_and_structures)
# * [Lists](#lists)
# * [Maps](#maps)
# * [References](#references)
# * [Multiple documents](#multiple_documents)
# * [Tags](#tags)
#
# ---
# **Y**AML **A**in't **M**arkup **L**anguage (**YAML**) is a serialization language that has steadily increased in popularity over the last few years. It's often used as a format for configuration files, but its object serialization abilities make it a viable replacement for languages like JSON. This YAML tutorial will demonstrate the language syntax with a guide and some simple coding examples in Python. YAML has broad language support and maps easily into native data structures. It's also easy to for humans to read, which is why it's a good choice for configuration. The YAML acronym was shorthand for **Yet Another Markup Language**. But the maintainers renamed it to **YAML Ain't Markup Language** to place more emphasis on its data-oriented features.
# YAML is a recently introduced data serialization format and is very comfortable for human reading and writing. **YAML is poised to replace XML and JSON**.
# <a class="anchor" id="yaml_tutorial_quick_start:_a_simple_file"></a>
# ## YAML Tutorial Quick Start: A Simple File
# Let's take a look at a YAML file for a brief overview.
# ```yaml
# ---
# quiz:
# description: >
# "This Quiz is to learn YAML."
# questions:
# - ["How many planets are there in the solar system?", "Name the non-planet"]
# - "Who is found more on the web?"
# - "What is the value of pi?"
# - "Is pluto related to platonic relationships?"
# - "How many maximum members can play TT?"
# - "Which value is no value?"
# - "Don't you know that the Universe is ever-expanding?"
#
# answers:
# - [8, "pluto"]
# - cats
# - 3.141592653589793
# - true
# - 4
# - null
# - no
# # explicit data conversion and reusing data blocks
# extra:
# refer: &id011 # give a reference to data
# x: !!float 5 # explicit conversion to data type float
# y: 8
# num1: !!int "123" # conversion to integer
# str1: !!str 120 # conversion to string
# again: *id011 # call data by giving the reference
# ```
# The identical `json` file is:
# ```json
# {
# "quiz": {
# "description": "\"This Quiz is to learn YAML.\"\n",
# "questions": [
# [
# "How many planets are there in the solar system?",
# "Name the non-planet"
# ],
# "Who is found more on the web?",
# "What is the value of pi?",
# "Is pluto related to platonic relationships?",
# "How many maximum members can play TT?",
# "Which value is no value?",
# "Don't you know that the Universe is ever-expanding?"
# ],
# "answers": [
# [
# 8,
# "pluto"
# ],
# "cats",
# 3.141592653589793,
# true,
# 4,
# null,
# false
# ]
# },
# "extra": {
# "refer": {
# "x": 5.0,
# "y": 8
# },
# "num1": 123,
# "str1": "120",
# "again": {
# "x": 5.0,
# "y": 8
# }
# }
# }
# ```
# JSON and YAML have similar capabilities, and you can convert most documents between the formats.
# <a class="anchor" id="installation"></a>
# ## Installation
# Install a Python module called the `pyyaml` to work with YAML files.
#
# ```bash
# pip install pyyaml
# ```
# <a class="anchor" id="read_yaml"></a>
# ## Read YAML
# Let's create a simple yaml file first:
# %%writefile learn_yaml.yaml
message: Hello World
# You can read a yaml file with `yaml.load`:
# +
import yaml
with open("learn_yaml.yaml", 'r') as f:
yaml_content = yaml.load(f, yaml.SafeLoader)
print("Key: Value")
for key, value in yaml_content.items():
print(f"{key}: {value}")
# -
# Also, `yaml.load` can read from a string:
yaml.load("this is", Loader=yaml.SafeLoader)
# Consider the following point number of “pi”, which has a value of 3.1415926. In YAML, it is represented as a floating number as shown below:
yaml.load('3.1415926536', Loader=yaml.SafeLoader)
# Suppose, multiple values are to be loaded in specific data structure as mentioned below:
#
# ```
# eggs
# ham
# spam
# French basil salmon terrine
# ```
yaml.load(
"""
- eggs
- ham
- spam
- French basil salmon terrine
""",
Loader=yaml.SafeLoader
)
# **Note:** You can read on why calling `yaml.load` without `Loader` is deprecated in [here](https://github.com/yaml/pyyaml/wiki/PyYAML-yaml.load(input)-Deprecation)
# To make it easier through this section, we define and use this function to parse yaml strings:
def read_yaml(string):
return yaml.load(string, Loader=yaml.FullLoader)
# <a class="anchor" id="introduction"></a>
# ## Introduction
# Now that you have an idea about YAML and its features, let us learn its basics with syntax and other operations. Remember that YAML includes a human readable structured format.
# <a class="anchor" id="rules_for_creating_yaml_file"></a>
# ### Rules for Creating YAML file
#
# When you are creating a file in YAML, you should remember the following basic rules:
#
# - YAML is case sensitive
# - The files should have `.yaml` or `.yml` as the extension
# - YAML does not allow the use of tabs while creating YAML files; spaces are allowed instead.
# <a class="anchor" id="indentation_of_yaml"></a>
# ### Indentation of YAML
#
# YAML does not include any mandatory spaces. Further, there is no need to be consistent. The valid YAML indentation is shown below.
read_yaml(
"""
a:
b:
- c
- d
- e
f:
"ghi"
"""
)
# You should remember the following rules while working with indentation in YAML:
#
# - Flow blocks must be intended with at least some spaces with surrounding current block level.
# - Flow content of YAML spans multiple lines. The beginning of flow content begins with `{` or `[`.
# - Block list items include same indentation as the surrounding block level because `-` is considered as a part of indentation.
# <a class="anchor" id="basics_components"></a>
# ### Basics Components
# The basic components of YAML are described below.
# <a class="anchor" id="conventional_block_format"></a>
# #### Conventional Block Format
# This block format uses hyphen+space to begin a new item in a specified list. Observe the example shown below:
read_yaml("""
# Favorite movies
- Casablanca
- North by Northwest
- The Man Who Wasn't There
"""
)
# **Note:** Comments in YAML begins with the `#` character. YAML does not support multi line comments.
# <a class="anchor" id="inline_format"></a>
# #### Inline Format
#
# Inline format is delimited with comma and space and the items are enclosed in `JSON`. Observe the example shown below:
read_yaml("""
# Shopping list
[milk, groceries, eggs, juice, fruits]
""")
read_yaml(yaml_string)
# <a class="anchor" id="folded_text"></a>
# #### Folded Text
#
# Folded text converts newlines to spaces and removes the leading whitespace. Observe the example shown below:
read_yaml("""
- {name: <NAME>, age: 33}
- name: <NAME>
age: 27
""")
# <a class="anchor" id="yaml_syntax_and_format"></a>
# ## YAML Syntax and format
# To begin with a YAML document, one needs to use `---`. All nested elements need to indented using two spaces but you can use one too. These three dashes are required only when there is more than one document within a single YAML file.
# <a class="anchor" id="string"></a>
# ### String
# <a class="anchor" id="short_texts"></a>
# #### Short texts
# Short texts are written, as shown below.
# ```yaml
# description: This is a brief description of something
# another description: "This is also a short description with ':' in it."
# 55.5 : keys need not be only strings
# ```
# <a class="anchor" id="long_text"></a>
# #### Long text
#
# Long texts are written using `|` and `>`. A block written using `|` is called a literal block, and a block written with `>` is called a folded block.
# ```yaml
# long description: |
# This text preserves the line breaks,
# as well as the indentation.
# folded long description: >
# This is also one way of writing the long text, but
# line breaks and indentations will be replaced with single spaces when read.
# ```
# The block style indicates how newlines inside the block should behave. If you would like them to be kept as **newlines**, use the **literal style**, indicated by a **pipe** (`|`). If instead you want them to be **replaced by spaces**, use the **folded style**, indicated by a **right angle bracket** (`>`). (To get a newline using the folded style, leave a blank line by putting two newlines in. Lines with extra indentation are also not folded.)
read_yaml("""
long description: |
This text preserves the line breaks,
as well as the indentation.
folded long description: >
This is also one way of writing the long text, but
line breaks and indentations will be replaced with single spaces when read.
"""
)
# <a class="anchor" id="numbers"></a>
# ### Numbers
#
# Integer numbers are written as shown below:
#
# ```yaml
# canonical: 12345
# decimal: +12,345
# octal: 014
# hexadecimal: 0xC
# ```
# Float numbers are written as shown below:
#
# ```yaml
# canonical: 1.23015e+3
# exponential: 12.3015e+02
# fixed: 1,230.15
# negative infinity: -.inf
# not a number: .NaN
# ```
# <a class="anchor" id="boolean"></a>
# ### Boolean
#
# Boolean is written as true and false:
#
# ```yaml
# flag: false
# not flag: true
# ```
# <a class="anchor" id="collections_and_structures"></a>
# ### Collections and Structures
# <a class="anchor" id="lists"></a>
# #### Lists
#
# YAML includes block collections which use indentation for scope. Here, each entry begins with a new line. Block sequences in collections indicate each entry with a dash and space (`-`). In YAML, block collections styles are not denoted by any specific indicator. Block collection in YAML can distinguished from other scalar quantities with an identification of key value pair included in them.
#
# Lists or arrays are written as shown below:
#
# ```yaml
# simple list: [1, 2, 3, four, five, “with quotes”]
# nested list:
# - item 1
# - item 2
# -
# - one
# - two
# - three
# - four
# ```
read_yaml(
"""
simple list: [1, 2, 3, four, five, “with quotes”]
nested list:
- item 1
- item 2
-
- one
- two
- three
- four
""")
# <a class="anchor" id="maps"></a>
# ### Maps
#
# Mappings are the representation of key value as included in JSON structure. It is used often in multi-lingual support systems and creation of API in mobile applications. Mappings use key value pair representation with the usage of colon and space (:).
#
# Maps can also be nested as shown below:
#
# ```yaml
# This:
# is a:
# nested_map:
# key: value
# ```
read_yaml(
"""
This:
is a:
nested_map:
key: value
"""
)
# <a class="anchor" id="references"></a>
# ### References
#
# References or anchors are useful when you want to copy an existing structure into a new form without repeating. An example is given below:
read_yaml(
"""
repeated text: &my_text This text need not be written again
an anchor: *my_text
# We can reference the maps too
person: &person
age: 10
# All members below are of the same age
member 1:
<<: *person
name: John
member 2:
<<: *person
name: Dave
"""
)
# <a class="anchor" id="multiple_documents"></a>
# ### Multiple documents
#
# Three dashes `---`, mark the beginning of a new document
# +
# read_yaml(
# """
# ---
# message: Message of document one
# ---
# message: Message of document two
# """
# )
# -
# <a class="anchor" id="tags"></a>
# ### Tags
#
# Tags, in YAML, are used to declare types. Examples are given below:
read_yaml(
"""
not a number: !!str 0.00035
python tuple: !!python/tuple [10,1]
# An example of storing a binary file
an image: !!binary |
R0lGODlhAQABAAAAACH5BAEKAAEALAAAAAABAAEAAAICTAEAOw==
"""
)
read_yaml(
"""
integer: !!str 3
key: !!python/tuple [1, 2]
""")
| Python/05. Modules/01.3 yaml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
myCat = {'size': 'fat', 'color': 'gray', 'disposition': 'loud'}
myCat['size']
'My cat has ' + myCat['color'] + ' fur.'
eggs = {'name': 'Zophie', 'species': 'cat', 'age': '8'}
ham = {'species': 'cat', 'age': '8', 'name': 'Zophie'}
eggs == ham
spam = ['cats', 'dogs', 'moose']
bacon = ['dogs', 'moose', 'cats']
spam == bacon
eggs['color']
birthdays = {'Alice': 'Apr 1', 'Bob': 'Dec 12', 'Carol': 'Mar 4'}
while True:
print('Enter a name: (blank to quit)')
name = input()
if name == '':
break
if name in birthdays:
print(birthdays[name] + ' is the birthday of ' + name)
else:
print('I do not have birthday information for ' + name)
print('What is their birthday?')
bday = input()
birthdays[name] = bday
print('Birthday database updated.')
list(eggs)
list(ham)
for key in eggs.keys():
print(key)
for value in eggs.values():
print(value)
for item in eggs.items():
print(item)
# unpacking the tuple
for key,value in eggs.items():
print(key, value)
'name' in eggs.keys()
'color' in eggs.keys()
picnicItems = {'apples': 5, 'cups': 2}
'I am bringing ' + str(picnicItems.get('cups', 0)) + ' cups.' # 0 is default if not found
'I am bringing ' + str(picnicItems.get('eggs', 0)) + ' eggs.'
spam = {'name': 'Pooka', 'age': 5}
if 'color' not in spam:
spam['color'] = 'black'
spam
spam = {'name': 'Pooka', 'age': 5}
spam.setdefault('color', 'black')
spam
spam.setdefault('color', 'white')
spam
message = 'It was a bright cold day in April, and the clocks were striking thirteen.'
count = {}
for character in message:
count.setdefault(character, 0)
count[character] = count[character] + 1
print(count)
import pprint
message = 'It was a bright cold day in April, and the clocks were striking thirteen.'
count = {}
for character in message:
count.setdefault(character, 0)
count[character] = count[character] + 1
pprint.pprint(count)
theBoard = {'top-L': ' ', 'top-M': ' ', 'top-R': ' ',
'mid-L': ' ', 'mid-M': ' ', 'mid-R': ' ',
'low-L': ' ', 'low-M': ' ', 'low-R': ' '}
from IPython.display import clear_output
def printBoard(board):
print(board['top-L'] + '|' + board['top-M'] + '|' + board['top-R'])
print('-+-+-')
print(board['mid-L'] + '|' + board['mid-M'] + '|' + board['mid-R'])
print('-+-+-')
print(board['low-L'] + '|' + board['low-M'] + '|' + board['low-R'])
turn = 'X'
for i in range(9):
clear_output(wait=True)
printBoard(theBoard)
print('Turn for ' + turn + '. Move on which space?')
move = input()
theBoard[move] = turn
if turn == 'X':
turn = 'O'
else:
turn = 'X'
printBoard(theBoard)
# +
allGuests = {'Alice': {'apples': 5, 'pretzels': 12},
'Bob': {'ham sandwiches': 3, 'apples': 2},
'Carol': {'cups': 3, 'apple pies': 1}}
def totalBrought(guests, item):
numBrought = 0
for k, v in guests.items():
numBrought = numBrought + v.get(item, 0)
return numBrought
print('Number of things being brought:')
print(' - Apples ' + str(totalBrought(allGuests, 'apples')))
print(' - Cups ' + str(totalBrought(allGuests,'cups')))
print(' - Cakes ' + str(totalBrought(allGuests, 'cakes')))
print(' - Ham Sandwiches ' + str(totalBrought(allGuests,'ham sandwiches')))
print(' - Apple Pies ' + str(totalBrought(allGuests,'apple pies')))
# +
def isValidChessBoard(board):
# check start with b or w and piece is of known time
pieceCounter = {}
for value in board.values():
if value[0] != 'b' and value[0] != 'w':
return False;
if value[1:] not in ['king', 'queen', 'bishop', 'rook', 'knight', 'pawn']:
return False;
pieceCounter[value] = pieceCounter.get(value, 0) + 1
# check for exactly one black king and one white king
if pieceCounter.get('bking') != 1 or pieceCounter.get('wking') != 1:
return False
if pieceCounter.get('bqueen', 0) > 1 or pieceCounter.get('wqueen', 0) > 1:
return False
# check for at most 16 pieces each, this get auto checked when we check count of all other pieces
# check for at most 8 pawns each
if pieceCounter.get('bpawn', 0) > 8 or pieceCounter.get('wpawn', 0) > 8:
return False
if pieceCounter.get('bbishop', 0) > 2 or pieceCounter.get('wbishop', 0) > 2:
return False
if pieceCounter.get('brook', 0) > 2 or pieceCounter.get('wrook', 0) > 2:
return false
if pieceCounter.get('bknight', 0) > 2 or pieceCounter.get('wknight', 0) > 2:
return False
# check key validity like 9z
for key in board.keys():
if int(key[0]) > 8 or key[1] > 'h' or key[1] < 'a':
return False
return True
board = {'1a': 'bking', '6c': 'wqueen',
'2g': 'bbishop', '5h': 'bqueen', '3e': 'wking'}
isValidChessBoard(board)
# -
stuff = {'rope': 1, 'torch': 6, 'gold coin': 42, 'dagger': 1, 'arrow': 12}
def displayInventory(inventory):
print("Inventory:")
item_total = 0
for k, v in inventory.items():
print(v, k, sep=" ")
item_total+=v
print("Total number of items: " + str(item_total))
displayInventory(stuff)
# +
dragonLoot = ['gold coin', 'dagger', 'gold coin', 'gold coin', 'ruby']
def addToInventory(inventory, addedItems):
for item in addedItems:
if item in inventory:
inventory[item] += 1
else :
inventory.setdefault(item, 1)
return inventory
inv = {'gold coin': 42, 'rope': 1}
dragonLoot = ['gold coin', 'dagger', 'gold coin', 'gold coin', 'ruby']
inv = addToInventory(inv, dragonLoot)
displayInventory(inv)
# -
| Chapter5/Chapter5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Answer Key to K-Means Quiz
# We might want to take a look at the distribution of the Title+Body length feature we used before and instead of using the raw number of words create categories based on this length: short, longer, and super long.
#
# In the questions below, I'll refer to length of the combined `Title` and `Body` fields as Description Length (and by length we mean the number of words when the text is tokenized with pattern=`"\\W"`).
from pyspark.sql import SparkSession
from pyspark.sql.functions import avg, col, concat, count, desc, explode, lit, min, max, split, stddev, udf
from pyspark.sql.types import IntegerType
from pyspark.ml.feature import RegexTokenizer, VectorAssembler
from pyspark.ml.regression import LinearRegression
from pyspark.ml.clustering import KMeans
spark = SparkSession.builder \
.master("local") \
.appName("Creating Features") \
.getOrCreate()
# ### Read Dataset
stack_overflow_data = 'Train_onetag_small.json'
df = spark.read.json(stack_overflow_data)
df.persist()
# ### Build Description Length Features
df = df.withColumn("Desc", concat(col("Title"), lit(' '), col("Body")))
regexTokenizer = RegexTokenizer(inputCol="Desc", outputCol="words", pattern="\\W")
df = regexTokenizer.transform(df)
body_length = udf(lambda x: len(x), IntegerType())
df = df.withColumn("DescLength", body_length(df.words))
assembler = VectorAssembler(inputCols=["DescLength"], outputCol="DescVec")
df = assembler.transform(df)
number_of_tags = udf(lambda x: len(x.split(" ")), IntegerType())
df = df.withColumn("NumTags", number_of_tags(df.Tags))
# # Question 1
# How many times greater is the Description Length of the longest question than the Description Length of the shortest question (rounded to the nearest whole number)?
#
# Tip: Don't forget to import Spark SQL's aggregate functions that can operate on DataFrame columns.
df.agg(min("DescLength")).show()
df.agg(max("DescLength")).show()
# # Question 2
# What is the mean and standard deviation of the Description length?
df.agg(avg("DescLength"), stddev("DescLength")).show()
# # Question 3
# Let's use K-means to create 5 clusters of Description lengths. Set the random seed to 42 and fit a 5-class K-means model on the Description length column (you can use KMeans().setParams(...) ). What length is the center of the cluster representing the longest questions?
kmeans = KMeans().setParams(featuresCol="DescVec", predictionCol="DescGroup", k=5, seed=42)
model = kmeans.fit(df)
df = model.transform(df)
df.head()
df.groupby("DescGroup").agg(avg(col("DescLength")), avg(col("NumTags")), count(col("DescLength"))).orderBy("avg(DescLength)").show()
| ml/8_k_means_quiz_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Camera Calibration
#
# Cameras introduce distortions into our data, these arise for various reasons such as the SSD, focal length, lens distortions, etc. Perhaps the most obvious is when using a fish-eye lens, this allows for a wider area to be captured however introduces large distortions near the image edges.
#
# Camera calibration is the process of determining the internal camera geometric and optical characteristics and/or the orientation of the camera frame relative to a certain world coordinate system. Physical camera parameters are commonly divided into extrinsic and intrinsic parameters.
#
# ## Extrinsic Parameters
# Extrinsic parameters are needed to transform object coordinates to a camera centered coordinate frame. To simplify this problem we use the camera pinhole model as shown in figure 1:
#
# <p style="text-align:center;"><img src="Camera_Calibration/pinhole.png" style="width:40%"/></p>
#
# This is based on the principle of collinearity, where each point in the object space (X,Y,Z) is projected by a straight line through the projection center into the image plane (x,y,z). so the problem can be simplified to a simple transformation:
#
# <p style="text-align:center;"><img src="Camera_Calibration/transformation.png" style="width:40%"/></p>
#
# Where matrix m carries out the translation and rotation which translates the coordinates of a 3D point to a coordinate system on the image plane.
#
# ## Intrinsic Parameters
# These are parameters specific to the camera and map the coordinates from the image-plane/SSD (x,y,z) to the digital image (pixel coordinate axis with origin commonly in the upper left corner of the iamge array). Parameters often included are the focal length $(f_x, f_y)$ ,optical centers $(c_x, c_y)$, etc. The values are often stored in what we call the camera matrix:
#
# <p style="text-align:center;"><img src="Camera_Calibration/matrix.png" style="width:40%"/></p>
#
# These are specific to the camera, so can be stored for future use.
#
#
# ## Extended model
# The pinhole model is only an approximation of the real camera projection! It is useful to establish simple mathematical formulations, however is not valid when high accuracy is required! Often the pinhole model is extended with some corrections, 2 common distortion corrections are:
#
# 1. radial -> straight lines will appear curved due to image points being displaced radially. More pronounced near the image edges. this distortion is solved as follows in our example below:
#
# $$x_{corrected} = x (1+k_1 r^2 +k_2 r^4 + k_3 r^6)$$
#
# $$y_{corrected} = y (1+k_1 r^2 +k_2 r^4 + k_3 r^6)$$
#
# 2. tangential -> occurs because the lense is not algined perfectly parallel to the image plane. Visualised as some areas in the image looking nearer than expected. This distortion is solved as follows in our example below:
#
# $$x_{corrected} = x + (2p_1xy+p_2(r^2+2x^2))$$
#
# $$y_{corrected} = y + (p_1(r^2+2y^2)+2p_2xy)$$
#
# So we are looking for the followig distortion coefficients:
#
# $$ (k_1, k_2, p_1, p_2, k_3)$$
#
# See image transformations notebook for further understanding of how these equations function
# ## Example Camera Calibration
# We can link the image coordinates and real world coordinates for a few points and hence we can solve for the translation matrix , with an empirical inverse model, to map between real world and image coordinates.
#
# Typically a checkerboard tends to be used to take images for easy point detection, from this the calibration matrixes are calculated. Then these matrixes can be stored and used to undistort other images. An example of how this process works is demonstrated below, images can be obtained from [here](https://github.com/DavidWangWood/Camera-Calibration-Python).
# import required libraries
import numpy as np
import cv2
import glob
import matplotlib.pyplot as plt
import pickle
# %matplotlib inline
# #%matplotlib qt
# +
# prepare object points, like (0,0,0), (1,0,0), (2,0,0) ....,(6,5,0)
objp = np.zeros((6*8,3), np.float32)
objp[:,:2] = np.mgrid[0:8, 0:6].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
# Make a list of calibration images
images = glob.glob('Camera_Calibration/GO*.jpg')
# Step through the list and search for chessboard corners
for idx, fname in enumerate(images):
img = cv2.imread(fname)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# Find the chessboard corners, set grid pattern that is looked for (8,6)
ret, corners = cv2.findChessboardCorners(gray, (8,6), None)
# If corners found, add object points, image points
if ret == True:
objpoints.append(objp)
imgpoints.append(corners)
# Draw and display the corners
cv2.drawChessboardCorners(img, (8,6), corners, ret)
#write_name = 'corners_found'+str(idx)+'.jpg'
#cv2.imwrite(write_name, img) #save image
cv2.imshow('img', img)
cv2.waitKey(500)
cv2.destroyAllWindows()
# +
# Test undistortion on an image
img = cv2.imread('Camera_Calibration/test_image.jpg')
img_size = (img.shape[1], img.shape[0])
# Do camera calibration given object points and image points
ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints, img_size,None,None)
#returns camera matrix, distortion coefficients, rotation and translation vectors
#undistort image
dst = cv2.undistort(img, mtx, dist, None, mtx)
cv2.imwrite('Camera_Calibration/test_undist.jpg',dst)
# Save the camera calibration result for later use (we won't worry about rvecs / tvecs)
dist_pickle = {}
dist_pickle["mtx"] = mtx
dist_pickle["dist"] = dist
pickle.dump( dist_pickle, open( "Camera_Calibration/wide_dist_pickle.p", "wb" ) )
#dst = cv2.cvtColor(dst, cv2.COLOR_BGR2RGB)
# + tags=["hide-input"]
# Visualize correction
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,10))
ax1.imshow(img)
ax1.set_title('Original Image', fontsize=30)
ax2.imshow(dst)
ax2.set_title('Undistorted Image', fontsize=30)
plt.show()
# -
# Now you can store the camera matrix and distortion coefficients using write functions in Numpy (np.savez, np.savetxt etc) for future uses.
#
# ## Errors
# Re-projection error gives a good estimation of just how exact the found parameters are. This should be as close to zero as possible.
tot_error=0
for i in range(len(objpoints)):
# transform object point to image point
imgpoints2, _ = cv2.projectPoints(objpoints[i], rvecs[i], tvecs[i], mtx, dist)
#calculate absolute norm between result of transformation and the corner finding algorithm
error = cv2.norm(imgpoints[i],imgpoints2, cv2.NORM_L2)/len(imgpoints2)
tot_error += error
#display the average error for all calibration images
print ("total error: ", tot_error/len(objpoints))
# to reduce errors a few measures can be taken:
# - make sure reflections on chessboard are minimized
# - use an checkerboard that cannot bend (print checkerboard and stick on a wooden/plastic board)
# - eliminate images with the largest calculated error as long as more than 10 images are remaining
#
# Ideal procedure to take pictures for calibration can be found [here](https://www.theeminentcodfish.com/gopro-calibration/).
#
# **NOTE**: If you change between wide angle/linear/super-wide or video/photo or resolution or any other function which may change image dimensions in px on the camera, you will require a new calibration matrix! Hence use the settings you are planning on using in your experiment from the start.
# example we obtained in the lab without cropping:
# <p style="text-align:center;"><img src="Camera_Calibration/Picture1.png" style="width:40%"/></p>
# <p style="text-align:center;"><img src="Camera_Calibration/Picture2.png" style="width:40%"/></p>
# # references
# - OpenCV functions: <br /> <NAME>. and <NAME>., 2017. Opencv-Python Tutorials Documentation. 1st ed. [ebook] pp.207 - 214. Available at: <https://buildmedia.readthedocs.org/media/pdf/opencv-python-tutroals/latest/opencv-python-tutroals.pdf> [Accessed 2 July 2020].
# - Code and checkerboard images obtained from: <br /> https://github.com/DavidWangWood/Camera-Calibration-Python
# - Paper with more advanced calibration technique: <br /> <NAME>. and <NAME>., n.d. A Four-step Camera Calibration Procedure with Implicit Image Correction. [online] Available at: <http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.94.7536&rep=rep1&type=pdf> [Accessed 2 July 2020].
# - pinhole image: <br /> Slideplayer.com. 2020. [online] Available at: <https://slideplayer.com/slide/5204330/16/images/1/The+Pinhole+Camera+Model.jpg> [Accessed 2 July 2020].
| geosciences/Camera_Calibration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Partial Reconfiguration
#
# Starting from image v2.4, the overlay class is able to handle
# partial bitstream downloading. After a partial bitstream is
# downloaded, the registers / parameters / hierarchies information
# will be exposed to users, so users can easily poke the registers
# associated to that partial region.
#
# As a reminder, users can always use the following code to download
# a full bitstream. Note that `BaseOverlay` is a wrapper for the
# underlying `Overlay` class.
#
# ```python
# from pynq.overlays.base import BaseOverlay
# base = BaseOverlay('base.bit')
# ```
# Then we can check the overlay information as follows:
# ```python
# print(base.ip_dict)
# print(base.interrupt_pins)
# print(base.gpio_dict)
# ```
#
# ## 1. Use `Overlay` class for static region
#
# The first step is to instantiate an `Overlay` object with the full bitstream.
# +
from pynq import Overlay
ol = Overlay('gpio_pr.bit')
# -
# After the full bitstream has been downloaded, the partial region will be loaded
# with the default Reconfigurable Module (RM). However, since the `hwh` file
# generated for the full bitstream ignores all the information in the partial
# region, no information on that partial region can be parsed.
#
# For the above reason, in the IP dictionary, internal IP blocks will be shown
# as memory interfaces (e.g. `gpio_0/S_AXI1`).
from pprint import pprint
pprint(ol.ip_dict)
# For this example, the `gpio_dict` of the overlay is empty,
# but you can check dictionaries for the interrupt, as well as
# the hierarchy dictionary in the next cell.
#
# We don't expect users to change the system interrupt controller, but
# the interrupt pins may get changed when downloading a partial bitstream
# onto a partial region.
#
# Note that before we download the partial bitstream, the interrupt pins have
# full paths only to the top level of the partial region (`gpio_0` in this example).
pprint(ol.interrupt_controllers)
pprint(ol.interrupt_pins)
# Use the following to check the hierarchy dictionary.
# ```python
# pprint(ol.hierarchy_dict)
# ```
#
# The hierarchy dictionary is quite a long dictionary so
# we will not show it in this notebook, but you
# can try it by yourself.
# ## 2. Interacting with the partial region
# Users have to specify the name of the partial region (hierarchical block name
# where you specified your partial region). In this example, it is `gpio_0`.
# Now we can reuse the `pr_download()` method with the following arguments -
# 1. the partial region name,
# 2. the name of the partial bitstream,
# 3. (optional) the name of the device tree blob file if any.
ol.pr_download('gpio_0', 'led_5.bit')
# Once the partial bitstream has been downloaded,
# you should be able to see the LEDs show the pattern `0101` (`0x5`)
# for LED 3 ~ LED 0.
# Also, all the relevant dictionaries are updated.
#
# We can also check which bitstream has been loaded.
pprint(ol.pr_dict)
# Notice that in the `ip_dict`, the internal IP block names have been
# updated.
pprint(ol.ip_dict)
# Pay attention to the following parameter; it indicates the default output
# value from the `leds_gpio` IP block.
ol.ip_dict['gpio_0/leds_gpio']['parameters']['C_DOUT_DEFAULT']
# You can also review the hierarchy dictionary by:
#
# ```python
# pprint(ol.hierarchy_dict)
# ```
#
# For interrupts, you can see that the interrupt pins are also updated.
# The full paths are now pointing to the internal GPIO blocks.
pprint(ol.interrupt_pins)
# You can also repeat the same process for another partial bitstream.
#
# For example, after running the next cell, you should be able to see the onboard LEDs show
# the pattern `1010` (`0xa`) for LED 3 ~ LED 0.
ol.pr_download('gpio_0', 'led_a.bit')
# Check the currently loaded bitstream.
pprint(ol.pr_dict)
# Now check the new value for the default output from `leds_gpio` IP block.
ol.ip_dict['gpio_0/leds_gpio']['parameters']['C_DOUT_DEFAULT']
# I hope you enjoy this example.
| boards/Pynq-Z1/gpio_pr/notebooks/partial_reconfig/partial_reconfiguration_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mappe interattive
# ## Introduzione
#
# Vediamo come controllare da Python delle mappe visualizzate in Jupyter con la libreria [ipyleaflet](https://ipyleaflet.readthedocs.io/) e [OpenStreetMap](https://www.openstreetmap.org), la mappa libera del mondo realizzata da volontari.
#
# <div class="alert alert-warning">
#
# **ATTENZIONE: Ciò che segue è solo una bozza MOLTO IN-PROGRESS !!!!!**
#
# </div>
#
# ### Prerequisiti
#
# Per proseguire è necessario prima aver letto il [tutorial sulle interfacce utente](https://it.softpython.org/gui/gui-sol.html) in Jupyter (che parla degli ipywidgets)
#
# ### Riferimenti
#
# * per una panoramica sul webmapping, la geolocalizzazione e OpenStreetMap, puoi vedere il tutorial [Integrazione](https://it.softpython.org/integration/integration-sol.html) dove si mappano gli agritur del Trentino.
# * per una rapida guida sull'HTML, vedere [tutorial CoderDojoTrento web 1 ](https://www.coderdojotrento.it/web1)
#
#
#
# ### Installazione ipyleaflet
#
# **Anaconda**:
#
# Apri Anaconda Prompt (per istruzioni su come trovarlo o se non hai idea di cosa sia, prima di proseguire [leggi sezione interprete Python nell'introduzione](https://it.softpython.org/intro/intro-sol.html#L'interprete-Python)) ed esegui:
#
# `conda install -c conda-forge ipyleaflet`
#
# Installare `ipyleaflet` con `conda` abiliterà automaticamente l'estensione per te in Jupyter
#
#
# **Linux/Mac**:
#
# * installa ipywidgets (`--user` installa nella propria home):
#
# ```bash
# python3 -m pip install --user ipyleaflet
# ```
#
# * abilita l'estensione così:
#
# ```bash
# jupyter nbextension enable --py ipyleaflet
# ```
# ### Proviamo OpenStreetMap
#
# Per prima cosa prova a navigare OpenStreetMap:
#
# [https://www.openstreetmap.org/#map=12/46.0849/11.1461](https://www.openstreetmap.org/#map=12/46.0849/11.1461)
#
# Spostati con la mappa, cambia lo zoom e nota cosa appare nella barra in alto dell'indirizzo del browser: dopo il `#` troverete le coordinate (latitudine e longitudine) e livello di zoom separati da una `/` :
#
# ### Mettiamo un palloncino
#
# Possiamo visualizzare una mappa in Jupyter sfruttando le coordinate trovate in OpenStreetMap. Inoltre, possiamo inserire dei palloncini (detti _marker_, [vedi documentazione](https://ipyleaflet.readthedocs.io/en/latest/api_reference/marker.html)), associando ad essi una descrizione formattata in linguaggio HTML - per una breve guida sull'HTML, vedere il [tutorial CoderDojoTrento web 1 ](https://www.coderdojotrento.it/web1)
#
# Nota che nella descrizione possiamo anche aggiungere immagini. In questo caso l'immagine d'esempio ([immagini/disi-unitn-it-logo.jpeg](immagini/disi-unitn-it-logo.jpeg)) risiede nella sottocartella `immagini` di questo foglio, ma volendo si potrebbero anche linkare foto da un sito qualunque usando il loro indirizzo per esteso, per esempio potresti provare a mettere questo indirizzo del logo CC BY che sta sul sito di softpython: [https://it.softpython.org/_images/cc-by.png](https://it.softpython.org/_images/cc-by.png)
#
#
# +
from ipywidgets import HTML, Layout, VBox, Button, Label, Image
from ipyleaflet import Map, Marker, Popup
# definiamo il centro della mappa su Trento (latitudine, longitudine)
centro_mappa = (46.0849,11.1461)
# per il livello di zoom giusto, puoi usare OpenStreetMap
mappa = Map(center=centro_mappa, zoom=12, close_popup_on_click=False)
# adesso andiamo a creare dei widget da mettere nel popup dei palloncini
# cominciamo con un bottone
bottone = Button(description="Cliccami")
def bottone_cliccato(b):
b.description="Mi hai cliccato !"
bottone.on_click(bottone_cliccato)
# per mostrare del testo formattato che contenga anche immagini, si può usare il linguaggio HTML
# per una breve guida sull'HTML, vedere il tutorial https://www.coderdojotrento.it/web1
# creiamo un widget HTML
# i tre doppi apici """ indicano che iniziamo una cosiddetta multistringa, cioè una stringa su più righe
html = HTML("""
Descrizione in <b>linguaggio HTML</b>,<br>
<a target="_blank" href="https://www.coderdojotrento.it/web1">vedi tutorial</a>. <br>
Questa è un immagine: <br/>
<img src="immagini/disi-unitn-it-logo.jpeg">
""")
# creaimo il pannello del popup come una VBox che contiene i due widget definiti precedentemente:
pannello_popup = VBox([bottone, html])
# il marcatore sarà un palloncino sul DISI, il Dipartimento di Informatica a Povo, Trento
marcatore = Marker(location=(46.06700,11.14985))
# associamo al marcatore un popup nella forma di un widget html
marcatore.popup = pannello_popup
# aggiungiamo il marcatore alla mappa
mappa.add_layer(marcatore)
# creiamo un widget che contenga titolo (in html) e la mappa
webapp = VBox([HTML("<h1>CLICCA SUI PALLONCINI</h1>"), mappa])
# infine forziamo Jupyter a mostrare il tutto:
display(webapp)
# +
# SOLUZIONE
# -
| gui/gui-maps-sol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Data Visualization I
# - Use the inbuilt dataset 'titanic'. The dataset contains 891 rows and contains information about the passengers who boarded the unfortunate Titanic ship. Use the Seaborn library to see if we can find any patterns in the data.
# - Write a code to check how the price of the ticket (column name: 'fare') for each
# passenger is distributed by plotting a histogram.
# # Titanic: Dataset
#
#
# Workflow: <br>
# * Exploratory Data Analysis. <br>
# - Surviving rate
# - Pclass
# - Name
# - Sex
# - Age
# - SibSp, Parch
# - Ticket
# - Fare
# - Cabin
# - Embarked
# * Feature Engineering <br>
# - Imputation on Embarked and Age columns
# - Title extraction
# - Ticket first letters
# - Cabin first letters
# - Encoding sex column
# - Family size
# - One Hot Encoding for all categorical variables
# * Machine Learning
# - Split data into train and test sets
# - Initialize a Random Forest Classifier
# - Hyperparameter Tuning with Grid Search
# - Prediction
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pickle
# I will keep the resulting plots
# %matplotlib inline
# Enable Jupyter Notebook's intellisense
# %config IPCompleter.greedy=True
# We want to see whole content (non-truncated)
pd.set_option('display.max_colwidth', None)
# -
# # 1. Exploratory Data Analysis
# +
train = pd.read_csv("train.csv")
display(train.head())
print(train.info())
print(train.info())
print(train.describe())
# -
# Notes: <br>
# * There are some missing values in Age, Embarked and Cabin columns. <br>
# * We do not need PassengerId column
# * The surviving rate is 38.3% in our dataset
# ## Survived
#
# Let's start with Survived column. It contains integer 1 or 0 which correspond to surviving ( 1 = Survived, 0 = Not Survived)
# +
# Visualize with a countplot
sns.countplot(x="Survived", data=train)
plt.show()
# Print the proportions
print(train["Survived"].value_counts(normalize=True))
# -
# ## Pclass
#
# Pclass column contains the socioeconomic status of the passengers. It might be predictive for our model <br>
# 1 = Upper <br>
# 2 = Middle <br>
# 3 = Lower
# +
# Visualize with a countplot
sns.countplot(x="Pclass", hue="Survived", data=train)
plt.show()
# Proportion of people survived for each class
print(train["Survived"].groupby(train["Pclass"]).mean())
# How many people we have in each class?
print(train["Pclass"].value_counts())
# -
# As I expected, first class passengers have higher surviving rate. We will use this information in our training data.
# ## Name
#
# At a first glance, I thought that I would use the titles.
# Display first five rows of the Name column
display(train[["Name"]].head())
# We can extract the titles from names.
# +
# Get titles
train["Title"] = train['Name'].str.split(', ', expand=True)[1].str.split('.', expand=True)[0]
# Print title counts
print(train["Title"].value_counts())
# -
# Is there any relationship between titles and surviving
# Print the Surviving rates by title
print(train["Survived"].groupby(train["Title"]).mean().sort_values(ascending=False))
# Apparently, there is relationship between titles and surviving rate. In feature engineering part, I will group title by their surviving rates like following <br>
#
# **higher** = the Countess, Mlle, Lady, Ms , Sir, Mme, Mrs, Miss, Master <br>
# **neutral** = Major, Col, Dr <br>
# **lower** = Mr, Rev, Jonkheer, Don, Capt <br>
# ## Age
# Print the missing values in Age column
print(train["Age"].isnull().sum())
# There are 177 missing values in Age column, we will impute them in Feature engineering part. Now, let's look at the distribution of ages by surviving
# +
# Survived by age
sns.distplot(train[train.Survived==1]["Age"],color="y", bins=7, label="1")
# Death by age
sns.distplot(train[train.Survived==0]["Age"], bins=7, label="0")
plt.legend()
plt.title("Age Distribution")
plt.show()
# -
# ## Sex
#
# Is sex important for surviving?
# +
# Visualize with a countplot
sns.countplot(x="Sex", hue="Survived", data=train)
plt.show()
# Proportion of people survived for each class
print(train["Survived"].groupby(train["Sex"]).mean())
# How many people we have in each class?
print(train["Sex"].value_counts())
# -
# Obviously, there is a relationship between sex and surviving.
# ## SibSp & Parch
# SibSp = Sibling or Spouse number <br>
# Parch = Parent or Children number <br>
#
# I decided to make a new feature called family size by summing the SibSp and Parch columns
# +
print(train["SibSp"].value_counts())
print(train["Parch"].value_counts())
train["family_size"] = train["SibSp"] + train["Parch"]
print(train["family_size"].value_counts())
# Proportion of people survived for each class
print(train["Survived"].groupby(train["family_size"]).mean().sort_values(ascending=False))
# -
# Apparently, family size is important to survive. I am going to group them in feature engineering step like following <br>
#
# **big family** = if family size > 3 <br>
# **small family** = if family size > 0 and family size < =3<br>
# **alone** = family size == 0 <br>
# ## Ticket
# At first, I thought that I would drop this column but after exploration I found useful features.
# Print the first five rows of the Ticket column
print(train["Ticket"].head(15))
# I extracted only first letters of the tickets because I thought that they would indicate the ticket type.
# +
# Get first letters of the tickets
train["Ticket_first"] = train["Ticket"].apply(lambda x: str(x)[0])
# Print value counts
print(train["Ticket_first"].value_counts())
# Surviving rates of first letters
print(train.groupby("Ticket_first")["Survived"].mean().sort_values(ascending=False))
# -
# The first letters of the tickets are correlated with surviving rate somehow. I am going to group them like following
#
# **higher surviving rate** = F, 1, P , 9 <br>
# **neutral** = S, C, 2 <br>
# **lower surviving rate** = else <br>
# ## Fare
# We can plot a histogram to see Fare distribution
# +
# Print 3 bins of Fare column
print(pd.cut(train['Fare'], 3).value_counts())
# Plot the histogram
sns.distplot(train["Fare"])
plt.show()
# Print binned Fares by surviving rate
print(train['Survived'].groupby(pd.cut(train['Fare'], 3)).mean())
# -
# There is also a correlation between ticket fares and surviving
# ## Cabin
# <img src="./images/titanic.png" width="1000" height="500">
# I found this figure [wikiwand.com](https://www.wikiwand.com/en/Sinking_of_the_Titanic). The figure shows us the most affacted parts of the Titanic and the Cabin locations. Although there are many missing value in Cabin column, I decided to extract the Cabin information to try whether it works or not.
# +
# Print the unique values in the Cabin column
print(train["Cabin"].unique())
# Get the first letters of Cabins
train["Cabin_first"] = train["Cabin"].apply(lambda x: str(x)[0])
# Print value counts of first letters
print(train["Cabin_first"].value_counts())
# Surviving rate of Cabin first letters
print(train.groupby("Cabin_first")["Survived"].mean().sort_values(ascending=False))
# -
# According to surviving rates. I will group the Cabins like following
#
# **higher surviving rate** = D, E, B, F, C <br>
# **neutral** = G, A<br>
# **lower surviving rate** else <br>
# ## Embarked
# Embarked is a categorical features which shows us the port of embarkation. <br>
#
# C = Cherbourg, Q = Queenstown, S = Southampton
# +
# Make a countplot
sns.countplot(x="Embarked", hue="Survived", data=train)
plt.show()
# Print the value counts
print(train["Embarked"].value_counts())
# Surviving rates of Embarked
print(train["Survived"].groupby(train["Embarked"]).mean())
# -
# No doubt, C has the higher surviving rate. We will definetely use this information.
# # 2. Feature Engineering
#
# We have learned a lot from exploratory data analysis. Now we can start feature engineering. Firstly, let's load the train and the test sets.
# +
# Load the train and the test datasets
train = pd.read_csv("train.csv")
test = pd.read_csv("test.csv")
print(test.info())
# -
# There is one missing value in the Fare column of the test set. I imputed it by using mean.
# Put the mean into the missing value
test['Fare'].fillna(train['Fare'].mean(), inplace = True)
# I have used two types of Imputer from sklearn. Iterative imputer for age imputation, and Simple imputer ( with most frequent strategy) for Embarked
# +
from sklearn.impute import SimpleImputer
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
# Imputers
imp_embarked = SimpleImputer(missing_values=np.nan, strategy="most_frequent")
imp_age = IterativeImputer(max_iter=100, random_state=34, n_nearest_features=2)
# Impute Embarked
train["Embarked"] = imp_embarked.fit_transform(train[["Embarked"]])
test["Embarked"] = imp_embarked.transform(test[["Embarked"]])
# Impute Age
train["Age"] = np.round(imp_age.fit_transform(train[["Age"]]))
test["Age"] = np.round(imp_age.transform(test[["Age"]]))
# -
# We also encode the sex column.
# +
from sklearn.preprocessing import LabelEncoder
# Initialize a Label Encoder
le = LabelEncoder()
# Encode Sex
train["Sex"] = le.fit_transform(train[["Sex"]].values.ravel())
test["Sex"] = le.fit_transform(test[["Sex"]].values.ravel())
# -
# In EDA, we decided to use family size feature
# Family Size
train["Fsize"] = train["SibSp"] + train["Parch"]
test["Fsize"] = test["SibSp"] + test["Parch"]
# Ticket first letters and Cabin first letters are also needed
# +
# Ticket first letters
train["Ticket"] = train["Ticket"].apply(lambda x: str(x)[0])
test["Ticket"] = test["Ticket"].apply(lambda x: str(x)[0])
# Cabin first letters
train["Cabin"] = train["Cabin"].apply(lambda x: str(x)[0])
test["Cabin"] = test["Cabin"].apply(lambda x: str(x)[0])
# -
# Extract the titles from the names
# Titles
train["Title"] = train['Name'].str.split(', ', expand=True)[1].str.split('.', expand=True)[0]
test["Title"] = test['Name'].str.split(', ', expand=True)[1].str.split('.', expand=True)[0]
# Now, we need some helper functions to group our categories
# +
# Group the family_size column
def assign_passenger_label(family_size):
if family_size == 0:
return "Alone"
elif family_size <=3:
return "Small_family"
else:
return "Big_family"
# Group the Ticket column
def assign_label_ticket(first):
if first in ["F", "1", "P", "9"]:
return "Ticket_high"
elif first in ["S", "C", "2"]:
return "Ticket_middle"
else:
return "Ticket_low"
# Group the Title column
def assign_label_title(title):
if title in ["the Countess", "Mlle", "Lady", "Ms", "Sir", "Mme", "Mrs", "Miss", "Master"]:
return "Title_high"
elif title in ["Major", "Col", "Dr"]:
return "Title_middle"
else:
return "Title_low"
# Group the Cabin column
def assign_label_cabin(cabin):
if cabin in ["D", "E", "B", "F", "C"]:
return "Cabin_high"
elif cabin in ["G", "A"]:
return "Cabin_middle"
else:
return "Cabin_low"
# -
# Apply the functions.
# +
# Family size
train["Fsize"] = train["Fsize"].apply(assign_passenger_label)
test["Fsize"] = test["Fsize"].apply(assign_passenger_label)
# Ticket
train["Ticket"] = train["Ticket"].apply(assign_label_ticket)
test["Ticket"] = test["Ticket"].apply(assign_label_ticket)
# Title
train["Title"] = train["Title"].apply(assign_label_title)
test["Title"] = test["Title"].apply(assign_label_title)
# Cabin
train["Cabin"] = train["Cabin"].apply(assign_label_cabin)
test["Cabin"] = test["Cabin"].apply(assign_label_cabin)
# -
# It's time to use One Hot Encoding
train = pd.get_dummies(columns=["Pclass", "Embarked", "Ticket", "Cabin","Title", "Fsize"], data=train, drop_first=True)
test = pd.get_dummies(columns=["Pclass", "Embarked", "Ticket", "Cabin", "Title", "Fsize"], data=test, drop_first=True)
# Drop the colums that are no longer needed
target = train["Survived"]
train.drop(["Survived", "SibSp", "Parch", "Name", "PassengerId"], axis=1, inplace=True)
test.drop(["SibSp", "Parch", "Name","PassengerId"], axis=1, inplace=True)
# Final look
# +
display(train.head())
display(test.head())
print(train.info())
print(test.info())
# -
# # 3. Machine Learning
# To evaluate our model's performance, we need to split our train data into training and test sets.
# +
from sklearn.model_selection import train_test_split
# Select the features and the target
X = train.values
y = target.values
# Split the data info training and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=34, stratify=y)
# -
# I have used GridSearchCV for tuning my Random Forest Classifier
# +
# Import Necessary libraries
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
from sklearn.metrics import accuracy_score, roc_auc_score, confusion_matrix, classification_report
# Initialize a RandomForestClassifier
rf = RandomForestClassifier(random_state=34)
params = {'n_estimators': [50, 100, 200, 300, 350],
'max_depth': [3,4,5,7, 10,15,20],
'criterion':['entropy', 'gini'],
'min_samples_leaf' : [1, 2, 3, 4, 5, 10],
'max_features':['auto'],
'min_samples_split': [3, 5, 10, 15, 20],
'max_leaf_nodes':[2,3,4,5],
}
# +
clf = GridSearchCV(estimator=rf,param_grid=params,cv=10, n_jobs=-1)
clf.fit(X_train, y_train.ravel())
print(clf.best_estimator_)
print(clf.best_score_)
rf_best = clf.best_estimator_
# Predict from the test set
y_pred = clf.predict(X_test)
# Print the accuracy with accuracy_score function
print("Accuracy: ", accuracy_score(y_test, y_pred))
# Print the confusion matrix
print("\nConfusion Matrix\n")
print(confusion_matrix(y_test, y_pred))
# -
# Save the model
pickle.dump(rf_best, open("model.pkl", 'wb'))
# We can look at the feature importances.
# +
# Create a pandas series with feature importances
importance = pd.Series(rf_best.feature_importances_,index=train.columns).sort_values(ascending=False)
sns.barplot(x=importance, y=importance.index)
# Add labels to your graph
plt.xlabel('Importance')
plt.ylabel('Feature')
plt.title("Important Features")
plt.show()
# -
# Train the model again with entire train data.
# +
last_clf = RandomForestClassifier(bootstrap=True, ccp_alpha=0.0, class_weight=None,
criterion='gini', max_depth=4, max_features='auto',
max_leaf_nodes=5, max_samples=None,
min_impurity_decrease=0.0, min_impurity_split=None,
min_samples_leaf=1, min_samples_split=15,
min_weight_fraction_leaf=0.0, n_estimators=350,
n_jobs=None, oob_score=True, random_state=34, verbose=0,
warm_start=False)
last_clf.fit(train, target)
print("%.4f" % last_clf.oob_score_)
# -
# Prepare the submission file
# +
# Store passenger ids
ids = pd.read_csv("test.csv")[["PassengerId"]].values
# Make predictions
predictions = last_clf.predict(test.values)
# Print the predictions
print(predictions)
# Create a dictionary with passenger ids and predictions
df = {'PassengerId': ids.ravel(), 'Survived':predictions}
# Create a DataFrame named submission
submission = pd.DataFrame(df)
# Display the first five rows of submission
display(submission.head())
# Save the file
submission.to_csv("submission_last.csv", index=False)
| 6th Sem/Data Science & Big Data/PR 8/Practical No 08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import cv2
import sys
sys.path.insert(0, '../')
import matplotlib.pyplot as plt
import tensorflow as tf
from sklearn.ensemble import IsolationForest
# +
def loadimg(img_path):
im = tf.keras.preprocessing.image.load_img(
img_path,
target_size=(300, 300, 3)
)
imarr = tf.keras.preprocessing.image.img_to_array(im)
imarr = tf.keras.applications.efficientnet.preprocess_input(imarr)
return np.array([imarr])
# -
# read NN features
df_features = pd.read_csv('../data/nn_features.csv', sep=',', header=None)
# subsample features
X_train = df_features.sample(5000, random_state = 42)
X_train.values.shape
# create outlier classifier
clf = IsolationForest(n_estimators=100, contamination = 0, max_samples = 500)
# train classifier on retrieval base features
clf = clf.fit(X_train)
# read model
model = tf.keras.models.load_model('../data/model.h5')
# read img to predict
img_path = '../test_imgs/dog.jpg'
# extract features
img_features_nn = model.predict(loadimg(img_path))[1][0]
# predict outlier
clf.predict([img_features_nn])
| notebooks/OutliersDetection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Planejamento da solução
# 1. Exploração dos dados visando encontrar inconsistências, tratamento de dados faltantes e possíveis insights que possam ser úteis ao time de negócios
#
# 2. Levantar hipóteses a partir da análise feita e validar as hipóteses em busca da correlações que possam ser úteis para o modelo de machine learning.
#
# 3. Preparação dos dados utlizando técnicas de rescaling e encodings.
#
# 4. Teste de vários algoritmos classificadores.
#
# 5. Avaliação dos modelos e tunning dos parametros.
#
# 6. Contabilizar a performance do modelo de Machine Learning e transformar em performance de negócio.
#
# 7. Fazer relatório respondedo as perguntas de negócio
#
# 8. Colocar modelo em produção e acessar as predições via Google Sheets
# # 0.0 Imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
import scikitplot as sckp
from IPython.core.display import display, HTML
# + [markdown] heading_collapsed=true
# ## 0.1 Helper Functions
# + hidden=true
def jupyter_settings():
# %matplotlib inline
# %pylab inline
plt.style.use( 'bmh' )
plt.rcParams['figure.figsize'] = [18, 9]
plt.rcParams['font.size'] = 24
display( HTML( '<style>.container { width:100% !important; }</style>') )
pd.options.display.max_columns = None
pd.options.display.max_rows = None
pd.set_option( 'display.expand_frame_repr', False )
pd.set_option('display.float_format', lambda x: '%.3f' % x)
warnings.filterwarnings('ignore')
sns.set()
jupyter_settings()
def ml_metrics(model_name, y_true, pred):
accuracy = m.balanced_accuracy_score(y_true, pred)
precision = m.precision_score(y_true, pred)
recall = m.recall_score(y_true, pred)
f1 = m.f1_score(y_true, pred)
kappa = m.cohen_kappa_score(y_true, pred)
return pd.DataFrame({'Balanced Accuracy': np.round(accuracy, 2),
'Precision': np.round(precision, 2),
'Recall': np.round(recall, 2),
'F1': np.round(f1, 2),
'Kappa': np.round(kappa, 2)}, index=[model_name])
def metrics_at_k(data, model_name, k=20000):
data = data.reset_index(drop=True)
# create ranking order
data['ranking'] = data.index + 1
# sort clients by propensity score
data = data.sort_values('score', ascending=False)
# precision
data['precision_at_k'] = data['response'].cumsum()/data['ranking']
# recall
data['recall_at_k'] = data['response'].cumsum()/data['response'].sum()
return pd.DataFrame({'Precision_at_k':data.loc[k, 'precision_at_k'],
'Recall_at_k':data.loc[k, 'recall_at_k']}, index=[model_name])
# -
# ## 0.2 Loading Data
df_raw = pd.read_csv('../data/insurance_cross_sell.csv')
df_raw_test = pd.read_csv('../data/test.csv')
# # 1.0 Data Description
df1 = df_raw.copy()
# - **Id**: identificador único do cliente.
#
# - **Gender**: gênero do cliente.
# - **Age**: idade do cliente.
# - **Driving License**: 0, o cliente não tem permissão para dirigir e 1, o cliente tem para dirigir ( CNH – Carteira Nacional de Habilitação )
# - **Region Code**: código da região do cliente.
# - **Previously Insured**: 0, o cliente não tem seguro de automóvel e 1, o cliente já tem seguro de automóvel.
# - **Vehicle Age**: idade do veículo.
# - **Vehicle Damage**: 0, cliente nunca teve seu veículo danificado no passado e 1, cliente já teve seu veículo danificado no passado.
# - **Anual Premium**: quantidade que o cliente pagou à empresa pelo seguro de saúde anual.
# - **Policy sales channel**: código anônimo para o canal de contato com o cliente.
# - **Vintage**: número de dias que o cliente se associou à empresa através da compra do seguro de saúde.
# - **Response**: 0, o cliente não tem interesse e 1, o cliente tem interesse.
# ## 1.1 Droping repeated columns
df1 = df1.drop(columns=['id.1', 'id.2'], axis=1)
df1.head()
# ## 1.2 Data Dimensions
df1.shape
# ## 1.3 Data Types
df1.dtypes
# ## 1.4 Check NA
df1.isna().sum()
# ## 1.5 Fillout Na
# There are not Nan values in the dataset
# ## 1.6 Change Types
# +
#changing the values 0 and 1 to 'yes' and 'no'. It'll help on the data description and analysis.
dic = {1:'yes', 0:'no'}
df1['driving_license'] = df1['driving_license'].map(dic)
df1['previously_insured'] = df1['previously_insured'].map(dic)
df1['response'] = df1['response'].map(dic)
# -
# ## 1.7 Descriptive Statistical
# + [markdown] heading_collapsed=true
# # 2.0 Feature Engineering
# + hidden=true
df2 = df1.copy()
# + [markdown] heading_collapsed=true
# # 3.0 Variables Filtering
# + hidden=true
df3 = df2.copy()
# + hidden=true
df3 = df3.drop(columns='vehicle_age', axis=1)
# + [markdown] heading_collapsed=true
# # 4.0 EDA
# + hidden=true
df4 = df3.copy()
# -
# # 5.0 Data preparation
df5 = df4.copy()
# ## 5.1 Split dataframe into training and validation dataset
from sklearn.model_selection import train_test_split
X = df5.drop(columns='response', axis=1)
y = df5['response'].copy()
X_train, X_valid, y_train, y_valid = train_test_split(X, y, test_size=0.2, random_state=42, stratify=y)
X_performance = X_valid.copy()
# ## 5.2 Rescaling
from sklearn.preprocessing import MinMaxScaler
# +
mm = MinMaxScaler()
X_train['age'] = mm.fit_transform(X_train[['age']].values)
X_valid['age'] = mm.fit_transform(X_valid[['age']].values)
X_train['region_code'] = mm.fit_transform(X_train[['region_code']].values)
X_valid['region_code'] = mm.fit_transform(X_valid[['region_code']].values)
X_train['policy_sales_channel'] = mm.fit_transform(X_train[['policy_sales_channel']].values)
X_valid['policy_sales_channel'] = mm.fit_transform(X_valid[['policy_sales_channel']].values)
X_train['annual_premium'] = mm.fit_transform(X_train[['annual_premium']].values)
X_valid['annual_premium'] = mm.fit_transform(X_valid[['annual_premium']].values)
X_train['vintage'] = mm.fit_transform(X_train[['vintage']].values)
X_valid['vintage'] = mm.fit_transform(X_valid[['vintage']].values)
# -
# ## 5.3 Encoding
# +
X_train['gender'] = X_train['gender'].map({'Female':0,'Male':1})
X_valid['gender'] = X_valid['gender'].map({'Female':0,'Male':1})
X_train['driving_license'] = X_train['driving_license'].map({'no':0,'yes':1})
X_valid['driving_license'] = X_valid['driving_license'].map({'no':0,'yes':1})
X_train['vehicle_damage'] = X_train['vehicle_damage'].map({'No':0,'Yes':1})
X_valid['vehicle_damage'] = X_valid['vehicle_damage'].map({'No':0,'Yes':1})
X_train['previously_insured'] = X_train['previously_insured'].map({'no':0,'yes':1})
X_valid['previously_insured'] = X_valid['previously_insured'].map({'no':0,'yes':1})
y_train = y_train.map({'no':0,'yes':1})
y_valid = y_valid.map({'no':0,'yes':1})
# -
# # 6.0 Feature Selection
df6 = df5.copy()
# # 7.0 Machine Learning Modeling
df7 = df6.copy()
# ## 7.1 Baseline Model
from sklearn.dummy import DummyClassifier
from sklearn import metrics as m
dummy = DummyClassifier()
dummy.fit(X_train, y_train)
pred = dummy.predict(X_valid)
pred_proba_dummy = dummy.predict_proba(X_valid)
# ### Model Performance
df_performance_dummy = X_performance.copy()
# +
df_performance_dummy['response'] = y_valid.copy()
# propensity score
df_performance_dummy['score'] = pred_proba_dummy[:, 1].tolist()
# compute metrics at K
df_metrics_dummy = metrics_at_k(df_performance_dummy, 'dummy', k=20000)
# -
dummy_results = ml_metrics('dummy', y_valid, pred)
dummy_results
# + [markdown] heading_collapsed=true
# ### Cumulative Gain Curve
# + hidden=true
sckp.metrics.plot_cumulative_gain(y_valid, pred_proba_dummy)
# -
# ## 7.2 Logistic Regression
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
lr.fit(X_train, y_train)
pred = lr.predict(X_valid)
pred_proba_lr = lr.predict_proba(X_valid)
# ### Model Performance
df_performance_lr = X_performance.copy()
df_performance_lr['response'] = y_valid.copy()
df_performance_lr['score'] = pred_proba_lr[:, 1].tolist()
df_metrics_lr = metrics_at_k(df_performance_lr, 'Logistic REegression', k=20000)
df_metrics_lr
logistic_regression_results = ml_metrics('Logistic Regression', y_valid, pred)
logistic_regression_results
# + [markdown] heading_collapsed=true
# ### Cumulative Gain Curve
# + hidden=true
sckp.metrics.plot_cumulative_gain(y_valid, pred_proba_lr)
# -
# ## 7.3 KNN
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
knn.fit(X_train, y_train)
pred = knn.predict(X_valid)
pred_proba_knn = knn.predict_proba(X_valid)
# ### Model Performance
df_performance_knn = X_performance.copy()
# +
df_performance_knn['response'] = y_valid.copy()
df_performance_knn['score'] = pred_proba_knn[:, 1].tolist()
df_metrics_knn = metrics_at_k(df_performance_knn, 'KNN', k=20000)
# -
knn_results = ml_metrics('KNN', y_valid, pred)
knn_results
# + [markdown] heading_collapsed=true
# ### Cumulative Gain Curve
# + hidden=true
sckp.metrics.plot_cumulative_gain(y_valid, pred_proba_knn)
# -
# ## 7.5 LightGBM
import lightgbm
lgb = lightgbm.LGBMClassifier(objective='binary', class_weight='balanced')
lgb.fit(X_train, y_train)
pred = lgb.predict(X_valid)
pred_proba_lgb = lgb.predict_proba(X_valid)
# ### Model Performance
df_performance_lgb = X_performance.copy()
# +
df_performance_lgb['response'] = y_valid.copy()
df_performance_lgb['score'] = pred_proba_lgb[:, 1].tolist()
df_metrics_lgb = metrics_at_k(df_performance_lgb, 'LGBM', k=20000)
# -
lgbm_results = ml_metrics('LGBM', y_valid, pred)
lgbm_results
# + [markdown] heading_collapsed=true
# ### Cumulative Gain Curve
# + hidden=true
sckp.metrics.plot_cumulative_gain(y_valid, pred_proba_lgb)
# -
# ## 7.6 Random Forest
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(class_weight='balanced', n_estimators=1000, n_jobs=-1)
rf.fit(X_train, y_train)
pred = rf.predict(X_valid)
pred_proba_rf = rf.predict_proba(X_valid)
# ### Model Performance
# +
df_performance_rf = X_performance.copy()
df_performance_rf['response'] = y_valid.copy()
df_performance_rf['score'] = pred_proba_rf[:, 1].tolist()
df_metrics_rf = metrics_at_k(df_performance_rf, 'Random Forest', k=20000)
# -
rf_results = ml_metrics('Random Forest', y_valid, pred)
rf_results
# + [markdown] heading_collapsed=true
# ### Cumulative Gain Curve
# + hidden=true
sckp.metrics.plot_cumulative_gain(y_valid, pred_proba_rf)
# -
# ## 7.7 XGBoost
from xgboost import XGBClassifier
xgb = XGBClassifier(objective='binary:logistic', verbosity=0)
xgb.fit(X_train, y_train)
pred = xgb.predict(X_valid)
pred_proba_xgb = xgb.predict_proba(X_valid)
# ### Model Performance
# +
df_performance_xgb = X_performance.copy()
df_performance_xgb['response'] = y_valid.copy()
df_performance_xgb['score'] = pred_proba_xgb[:, 1].tolist()
df_metrics_xgb = metrics_at_k(df_performance_xgb, 'XGBoost', k=20000)
# -
xgb_results = ml_metrics('XGBoost', y_valid, pred)
xgb_results
# + [markdown] heading_collapsed=true
# ### Cumulative Gain Curve
# + hidden=true
sckp.metrics.plot_cumulative_gain(y_valid, pred_proba_xgb)
# -
# ## 7.8 Results
# ### Results at K
df_results_at_k = pd.concat([df_metrics_dummy, df_metrics_lr, df_metrics_knn, df_metrics_lgb, df_metrics_rf, df_metrics_xgb])
df_results_at_k.style.highlight_max(color='lightgreen', axis=0)
# ### Results Regular Metrics
df_results = pd.concat([dummy_results, logistic_regression_results, knn_results,lgbm_results, rf_results, xgb_results])
df_results.style.highlight_max(color='lightgreen', axis=0)
# + [markdown] heading_collapsed=true
# # 8.0 Hyperparameter Fine Tuning
# + hidden=true
# + [markdown] heading_collapsed=true
# # 9.0 Conclusions
# + hidden=true
# -
# # 10.0 Deploy
| notebooks/cycle01-data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import dagstermill
# + tags=["parameters"]
from dagster import ModeDefinition, ResourceDefinition
from collections import namedtuple
url = "postgresql://{username}:{password}@{hostname}:5432/{db_name}".format(
username="test", password="<PASSWORD>", hostname="localhost", db_name="test"
)
DbInfo = namedtuple("DbInfo", "url")
context = dagstermill.get_context(
mode_def=ModeDefinition(resource_defs={"db_info": ResourceDefinition(lambda _: DbInfo(url))})
)
table_name = "delays_vs_fares"
# -
db_url = context.resources.db_info.url
# +
import os
import sqlalchemy as sa
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from dagster.utils import mkdir_p
# -
engine = sa.create_engine(db_url)
# +
from matplotlib.backends.backend_pdf import PdfPages
plots_path = os.path.join(os.getcwd(), "plots")
mkdir_p(plots_path)
pdf_path = os.path.join(plots_path, "fares_vs_delays.pdf")
pp = PdfPages(pdf_path)
# -
fares_vs_delays = pd.read_sql("select * from {table_name}".format(table_name=table_name), engine)
fares_vs_delays.head()
fares_vs_delays["avg_arrival_delay"].describe()
# +
plt.scatter(fares_vs_delays["avg_arrival_delay"], fares_vs_delays["avg_fare"])
try:
z = np.polyfit(fares_vs_delays["avg_arrival_delay"], fares_vs_delays["avg_fare"], 1)
f = np.poly1d(z)
x_fit = np.linspace(
fares_vs_delays["avg_arrival_delay"].min(), fares_vs_delays["avg_arrival_delay"].max(), 50
)
y_fit = f(x_fit)
plt.plot(x_fit, y_fit, "k--", alpha=0.5)
except:
pass
plt.title("Arrival Delays vs. Fares (Origin SFO)")
plt.xlabel("Average Delay at Arrival (Minutes)")
plt.ylabel("Average Fare ($)")
pp.savefig()
# +
fig, ax = plt.subplots(figsize=(10, 10))
for i, _ in enumerate(fares_vs_delays.index):
plt.text(
fares_vs_delays["avg_arrival_delay"][i],
fares_vs_delays["avg_fare_per_mile"][i],
fares_vs_delays["dest"][i],
fontsize=8,
)
plt.scatter(fares_vs_delays["avg_arrival_delay"], fares_vs_delays["avg_fare_per_mile"], alpha=0)
plt.title("Flight Delays (Origin SFO)")
plt.xlabel("Average Delay at Arrival (Minutes)")
plt.ylabel("Average Fare per Mile Flown($)")
pp.savefig()
# -
pp.close()
# +
from dagster import LocalFileHandle
dagstermill.yield_result(LocalFileHandle(pdf_path))
| examples/airline_demo/airline_demo/notebooks/Fares_vs_Delays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Visualization : seaborn
# In this tutorial we will explore how to create FacetGrid plots using a plotting tool called 'seaborn'. For that let's first import all the necessary packages:
import seaborn as sns
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Now suppose for example that you have a team in a company and you'd like to show the expenses of your team members in last week to your boss. How would that look like? Let's explore.
#name of team member:
name = ['Nilesh', 'Yoda', 'Kirk', 'Morpheus', 'Moriarty', 'Albert']
#days in a week
day = [0, 1, 2, 3, 4, 5, 6] #Sunday through Saturday
# Now for your team members, you need to track the expenses on each day. For that, you need to expand your name list, repeating every name 7 time and tiling every day 6 times(for each member)
name = np.repeat(name, 7).tolist() # repeat the names 7 times each and return a list
#print the contents of 'name'
name
# Just like we wanted, each name is repeated 7 times.
# Now let's tile the week days for each member i.e. 6 times
day = np.tile(day, 6).tolist() # tile the 'day's 6 times and return a list
#print day list
day
# Using these two lists let's create a dataframe. Before that we need to "zip" these two lists as follows:
np.c_[name, day]
# The above array will serve a content for the dataframe
df = pd.DataFrame(np.c_[name, day], columns=['Name','Day'])
#print dataframe
df
# But we don't have an expense list for every member on every week day. And we can't have because this is a hypothetical example. So let's just create one randomly
df.shape
expenses = []
for i in range(6):#for every member
expenses+=np.random.randint(0,100,7).tolist() #create an expense list of a week out of random numbers
len(expenses)
# This expenses list can be used to create a new column in dataframe
df['Expenses'] = expenses
df
# So, this is the dataframe you have at our disposal. Using this let's create some Facet plots.
#invoke the FacetGrid class using this dataframe
grid = sns.FacetGrid(df, col='Name', hue='Name', col_wrap=3, size=4.0)
# let's get a bit of insight into the parameters to this grid:
#
# * 'col' -> which column of dataset is to be considered for facets
# * 'hue' -> how to color the plot? By which column? Here a single subplot will have same color for all points but different subplots will have different colors
# * 'col_wrap' -> how many subplots to display in a horizontal strip of the plot
# * 'size' -> size of the subplot
#draw the horizontal axis at position y=0
grid.map(plt.axhline, y=0, ls="solid", c=".5")
# the parameters for above grid map specify:
#
# * 'y': the point at which the horizontal axis cuts the y axis
# * 'ls': also called linestyle, line design
# * 'c': is the color for horizontal line axis. A string of "0" means white while "1" means black. Any float between 0 and 1, as a string, is mixture of black and white.
#
# For more info, run:
help(plt.axhline)
df.columns
grid.map(plt.plot, "Day", "Expenses", marker="o", ms=6)
# For more info, run:
help(plt.plot)
# some more housekeeping; We encourage you to use the 'help' function in above manner to find out exactly what the following line is doing. It's very easy.
grid.set(xticks=np.arange(7), yticks=np.arange(-10,100,10), xlim=(-1,7), ylim=(-10,100))
# Again, run help on 'grid.fig.tight_layout' and try to figure out what w_pad = 1 means.
grid.fig.tight_layout(w_pad=1)
# ### Now, all the housekeeping is done; let's plot the grid
plt.show()
# Look at those subplots/facets!!
# Save the image:
# ### Now let's play a bit with grid parameters
#
# 1) change the col_wrap to 2
grid = sns.FacetGrid(df, col='Name', hue='Name',col_wrap=2, size=4.0)
grid.map(plt.axhline, y=10, ls="solid", c="0.5")
grid.map(plt.plot, "Day", "Expenses", marker="o", ms=6)
grid.set(xticks=np.arange(7), yticks=np.arange(-10,100,10), xlim=(-1,7), ylim=(-10,100))
grid.fig.tight_layout(w_pad=1)
plt.show()
# Did you notice the change?
# 2) change image size from 4 to 3
grid = sns.FacetGrid(df, col='Name', hue='Name',col_wrap=2, size=3.0)
grid.map(plt.axhline, y=10, ls="solid", c="0.5")
grid.map(plt.plot, "Day", "Expenses", marker="o", ms=6)
grid.set(xticks=np.arange(7), yticks=np.arange(-10,100,10), xlim=(-1,7), ylim=(-10,100))
grid.fig.tight_layout(w_pad=1)
plt.show()
# The size of the subplot/facet changes.
# 3) Now let's try to change the marker style *and* size, and see what happens
grid = sns.FacetGrid(df, col='Name', hue='Name',col_wrap=2, size=3.0)
grid.map(plt.axhline, y=10, ls="solid", c="0.5")
grid.map(plt.plot, "Day", "Expenses", marker="s", ms=10)
grid.set(xticks=np.arange(7), yticks=np.arange(-10,100,10), xlim=(-1,7), ylim=(-10,100))
grid.fig.tight_layout(w_pad=1)
plt.show()
# The reader is encouraged to change and play around with each and every parameter of the grid map and observe the effects which might prepare you for future requests.
# # Challenge:
#
# There is a subtle difference between the first plot and the subsequent plots. Can you figure it out? I will give you 30 secs.
| Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy.stats import expon, truncexpon, uniform
# Based on Mondrian Forests: Efficient Online Random Forests
# https://arxiv.org/pdf/1406.2673.pdf
GAMMA = 20
from mondrianforest import MondrianTree
# def data_ranges(data):
# return np.min(data, axis=0), np.max(data, axis=0)
# class MondrianTree:
# def __init__(self, budget=np.inf, random_state=None): # TODO: use random state
# self.leaf_nodes = set()
# self.budget = budget
# self.classes = None
# self.class_indices = None
# self.root = None
# self.X = None
# self.y = None
# self.fitted = False
# # Algorithm 1 + fully online option
# def fit(self, X, y, online=False):
# self.X = X
# self.y = y
# self.classes = np.unique(y)
# self.class_indices = {cls: i for i, cls in enumerate(self.classes)}
# if not online:
# self.root = MondrianBlock(X, y, parent=None, budget=self.budget, tree=self)
# self.compute_predictive_posterior()
# else:
# self.root = MondrianBlock(X[:2], y[:2], parent=None, budget=self.budget, tree=self)
# for i in range(2, len(y)):
# self.extend(X[i], y[i])
# plot_2d_mondrian_tree(self, X, y)
# self.compute_predictive_posterior()
# self.fitted = True
# # Algorithm 7
# def compute_predictive_posterior(self):
# queue = [self.root]
# while queue:
# node = queue.pop()
# if node.parent is None:
# parent_posterior = np.ones_like(self.classes) / len(self.classes) # H
# else:
# parent_posterior = node.parent.posterior_predictive
# class_counts = node.class_counts
# tables = node.tables
# discount = node.discount
# node.posterior_predictive = (class_counts - discount * tables
# + discount * np.sum(tables) * parent_posterior) / np.sum(class_counts)
# if node.left:
# queue = [node.left] + queue
# if node.right:
# queue = [node.right] + queue
# # Algorithm 8
# def predict(self, x):
# assert len(x.shape) == 1 # prediction for single x for now
# x += 1e-12 # dirty hack in case x is included in the training
# current = self.root
# pnsy = 1.
# s = np.zeros_like(self.classes, dtype=np.float64)
# while True:
# cost_difference = current.cost - current._parent_cost()
# eta = (np.maximum(x - current.upper, 0) + np.maximum(current.lower - x, 0)).sum()
# psjx = -np.expm1(-eta * cost_difference)
# if psjx > 0:
# expected_discount = (eta / (eta + GAMMA)) * (-np.expm1(-(eta + GAMMA) * cost_difference)) \
# / (-np.expm1(-eta * cost_difference))
# class_counts = tables = np.minimum(current.class_counts, 1)
# if current.parent is None:
# tilde_parent_posterior = np.ones_like(self.classes) / len(self.classes)
# else:
# tilde_parent_posterior = current.parent.posterior_predictive
# posterior = (class_counts / np.sum(class_counts) - expected_discount * tables
# + expected_discount * tables.sum() * tilde_parent_posterior)
# s += pnsy * psjx * posterior
# if current.is_leaf:
# s += pnsy * (1 - psjx) * current.posterior_predictive
# return s
# else:
# pnsy *= 1 - psjx
# if x[current.delta] <= current.xi:
# current = current.left
# else:
# current = current.right
# def extend(self, X, y):
# self.root.extend(X, y)
# class MondrianBlock:
# def __init__(self, X, y, budget, parent=None, tree: MondrianTree = None, fit=True):
# assert tree
# self.tree = tree
# self.parent = parent
# self.left = None
# self.right = None
# self.budget = budget
# self.discount = 0
# self.lower = np.zeros(X.shape[1]) if X is not None else None
# self.upper = np.zeros_like(self.lower) if X is not None else None
# self.sides = np.zeros_like(self.lower) if X is not None else None
# self.class_counts = np.zeros_like(self.tree.classes) # not exactly _counts_
# self.tables = np.zeros_like(self.tree.classes) # see Chinese restaurants notation in the paper
# self.is_leaf = True # will be set to False when needed
# if fit:
# self._fit(X, y)
# def _parent_cost(self):
# if self.parent is None:
# return 0.
# else:
# return self.parent.cost
# # Algorithm 5
# def _initialize_posterior_counts(self, X, y):
# for i, cls in enumerate(self.tree.classes):
# self.class_counts[i] = np.count_nonzero(y == cls)
# current = self
# while True:
# if not current.is_leaf:
# l_tables = current.left.tables if current.left else np.zeros_like(current.class_counts)
# r_tables = current.right.tables if current.right else np.zeros_like(current.class_counts)
# current.class_counts = l_tables + r_tables
# current.tables = np.minimum(current.class_counts, 1)
# if current.parent is None:
# break
# else:
# current = current.parent
# # Algorithm 6
# def _update_posterior_counts(self, y):
# class_index = self.tree.class_indices[y]
# self.class_counts[class_index] += 1
# current = self
# while True:
# if current.tables[class_index] == 1:
# return
# else:
# if not current.is_leaf:
# l_table = current.left.tables[class_index] if current.left else 0
# r_table = current.right.tables[class_index] if current.right else 0
# current.class_counts[class_index] = l_table + r_table
# current.tables[class_index] = np.minimum(current.class_counts[class_index], 1)
# if current.parent is None:
# return
# else:
# current = current.parent
# # Algorithm 9
# def _fit(self, X, y):
# self.lower, self.upper = data_ranges(X)
# self.sides = self.upper - self.lower
# if len(y) <= 0 or np.all(y == y[0]): # all labels identical
# self.cost = self.budget
# else:
# split_cost = expon.rvs(scale=(1 / self.sides.sum()))
# self.cost = self._parent_cost() + split_cost
# if self.cost < self.budget:
# # choose split dimension delta and location xi
# self.delta = np.random.choice(np.arange(X.shape[1]), p=(self.sides / self.sides.sum()))
# self.xi = uniform.rvs(loc=self.lower[self.delta], scale=self.sides[self.delta])
# # perform an actual split
# left_indices = X[:, self.delta] <= self.xi
# X_left, y_left = X[left_indices], y[left_indices]
# X_right, y_right = X[~left_indices], y[~left_indices]
# # sample children
# self.is_leaf = False
# # we first create unfitted blocks and then fit because otherwise self.left and self.right
# # may be accessed in ._initialize_posterior_counts before being assigned
# self.left = MondrianBlock(X_left, y_left, budget=self.budget, parent=self, tree=self.tree, fit=False)
# self.left._fit(X_left, y_left)
# self.right = MondrianBlock(X_right, y_right, budget=self.budget, parent=self, tree=self.tree, fit=False)
# self.right._fit(X_right, y_right)
# else:
# self.cost = self.budget
# self.tree.leaf_nodes.add(self)
# self._initialize_posterior_counts(X, y)
# self.discount = np.exp(-GAMMA * (self.cost - self._parent_cost()))
# def _get_subset_indices(self):
# return np.all(self.tree.X >= self.lower, axis=1) & np.all(self.tree.X <= self.upper, axis=1)
# def _get_label_subset(self, indices=None):
# if indices is None:
# indices = self._get_subset_indices()
# return self.tree.y[indices]
# def _get_feature_subset(self, indices=None):
# if indices is None:
# indices = self._get_subset_indices()
# return self.tree.X[indices]
# def _get_feature_label_subset(self, indices=None):
# if indices is None:
# indices = self._get_subset_indices()
# return self._get_feature_subset(indices), self._get_label_subset(indices)
# # Algorithm 10
# def extend(self, x, y):
# labels = self._get_label_subset()
# if len(labels) <= 0 or np.all(labels == labels[0]): # all labels identical
# self.lower = np.minimum(self.lower, x)
# self.upper = np.maximum(self.upper, x)
# self.tree.X = np.vstack((self.tree.X, x)) # TODO: we possibly don't have to
# self.tree.y = np.hstack((self.tree.y, y))
# if y == labels[0]:
# self._update_posterior_counts(y)
# return
# else:
# self.tree.leaf_nodes.remove(self)
# X, y = self._get_feature_label_subset()
# self._fit(X, y)
# else:
# el = np.maximum(self.lower - x, 0)
# eu = np.maximum(x - self.upper, 0)
# sum_e = el + eu
# split_cost = expon.rvs(scale=(1 / sum_e.sum()))
# if self._parent_cost() + split_cost < self.cost:
# delta = np.random.choice(np.arange(len(x)), p=(sum_e / sum_e.sum()))
# if x[delta] > self.upper[delta]:
# xi = uniform.rvs(loc=self.upper[delta], scale=x[delta] - self.upper[delta])
# else:
# xi = uniform.rvs(loc=x[delta], scale=self.lower[delta] - x[delta])
# j_tilde = MondrianBlock(None, None, budget=self.budget, parent=self.parent, tree=self.tree, fit=False)
# j_tilde_attrs = {
# 'delta': delta,
# 'xi': xi,
# 'cost': self._parent_cost() + split_cost,
# 'lower': np.minimum(self.lower, x),
# 'upper': np.maximum(self.upper, x),
# 'sides': np.maximum(self.upper, x) - np.minimum(self.lower, x),
# 'is_leaf': False,
# }
# for attr, value in j_tilde_attrs.items():
# setattr(j_tilde, attr, value)
# if self.parent is None:
# self.tree.root = j_tilde
# print(x, 'changing root')
# else:
# if self is self.parent.left:
# self.parent.left = j_tilde
# elif self is self.parent.right:
# self.parent.right = j_tilde
# j_primes = MondrianBlock(X=np.array([x]), y=np.array([y]), budget=self.budget,
# parent=j_tilde, tree=self.tree)
# if x[delta] > xi:
# j_tilde.left = self
# j_tilde.right = j_primes
# else:
# j_tilde.left = j_primes
# j_tilde.right = self
# else:
# self.lower = np.minimum(self.lower, x)
# self.upper = np.maximum(self.upper, x)
# if not self.is_leaf:
# if x[self.delta] <= self.xi:
# child = self.left
# else:
# child = self.right
# child.extend(x, y)
# class MondrianRandomForest:
# def __init__(self, n_estimators=100, budget=np.inf, random_state=4):
# self.n_estimators = n_estimators
# self.estimators = []
# self.budget = budget
# self.random_state = random_state
# def fit(self, X, y, online=False):
# if not online:
# for i in range(self.n_estimators):
# self.estimators.append(MondrianTree(self.budget))
# self.estimators[-1].fit(X, y, online=False)
# def predict(self, x):
# assert len(x.shape) == 1
# predictions = np.zeros((self.n_estimators, len(self.estimators[0].classes)))
# return predictions.mean(axis=0)
# +
def plot_2d_mondrian_block(block):
plt.hlines([block.lower[1], block.upper[1]],
block.lower[0], block.upper[0],
linestyles='dashed', alpha=0.3)
plt.vlines([block.lower[0], block.upper[0]],
block.lower[1], block.upper[1],
linestyles='dashed', alpha=0.3)
if not block.is_leaf:
if block.delta == 0:
plt.vlines(block.xi, block.lower[1], block.upper[1], color='red', alpha=0.7)
elif block.delta == 1:
plt.hlines(block.xi, block.lower[0], block.upper[0], color='red', alpha=0.7)
if block.left:
plot_2d_mondrian_block(block.left)
if block.right:
plot_2d_mondrian_block(block.right)
def plot_2d_mondrian_tree(tree, X, y, xlim=None, ylim=None, **kwargs):
xmin, xmax = (np.min(X[:, 0]), np.max(X[:, 0])) if not xlim else xlim
xlen = xmax - xmin
x_margin = 0 if xlim else 0.05 * xlen
ymin, ymax = (np.min(X[:, 1]), np.max(X[:, 1])) if not ylim else ylim
ylen = ymax - ymin
y_margin = 0 if ylim else 0.05 * ylen
plt.figure(**kwargs)
plt.xlim(xmin - x_margin, xmax + x_margin)
plt.ylim(ymin - y_margin, ymax + y_margin)
plt.scatter(X[:, 0], X[:, 1], c=y)
plot_2d_mondrian_block(tree.root)
# -
import warnings
warnings.filterwarnings("error")
# +
np.random.seed(12)
n = 12
n_classes = 2
X = np.random.uniform(size=(n, 2))
y = np.random.randint(0, n_classes, size=n)
m_tree = MondrianTree()
m_tree.fit(X, y, online=True)
# +
from matplotlib import pyplot as plt
# %matplotlib inline
plot_2d_mondrian_tree(m_tree, X, y)
# -
m_tree.root.right.lower
np.argwhere(np.array([0, 0, 1, 0, 0]))
# +
np.random.seed(4)
X = np.array([[0., 0.], [0.1, 0.05], [0.7, 0.7], [2., 0.5]])
y = np.random.randint(0, n_classes, size=4)
m_tree = MondrianTree()
m_tree.fit(X[:2], y[:2], online=False)
plot_2d_mondrian_tree(m_tree, X[:2], y[:2])
# -
m_tree.extend(X[2], y[2])
plot_2d_mondrian_tree(m_tree, X, y)
m_tree.extend(X[3], y[3])
plot_2d_mondrian_tree(m_tree, X, y)
m_tree.root.xi
m_tree.predict(np.array([0.5, 0.2]))
from sklearn.datasets import load_iris
X, y = load_iris(return_X_y=True)
indices = np.arange(150)
np.random.shuffle(indices)
X = X[indices]
y = y[indices]
m_tree = MondrianTree()
m_tree.fit(X[:120], y[:120], online=False)
y
X
X[121]
m_tree.predict(np.array([7., 3., 5.5, 2.]))
m_tree.predict(np.array([0., 0.]))
m_tree.predict(np.array([0.03, 0.5]))
m_tree.root.class_counts
| mforests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import chemdataextractor as cde
from chemdataextractor.relex import Snowball, ChemicalRelationship
from chemdataextractor.model import BaseModel, StringType, ListType, ModelType, Compound
from chemdataextractor.parse import R, I, W, Optional, merge, join, OneOrMore, Any, ZeroOrMore, Start
from chemdataextractor.parse.cem import chemical_name, chemical_label
from chemdataextractor.parse.base import BaseParser
from chemdataextractor.parse.common import lrb, rrb, delim
from chemdataextractor.utils import first
from chemdataextractor.doc import Paragraph, Heading, Sentence
from lxml import etree
import re
# -
# # Abbreviations
#
# #### Resources:
# * https://en.wikiversity.org/wiki/Python_Concepts/Regular_Expressions#\d_and_\D
# * http://dev.lexalytics.com/wiki/pmwiki.php?n=Main.POSTags
#
# W = Word (case sensitive)
#
# I = IWord (case insensetive)
#
# R = Regex (regular expression)
#
# T = Tag (match tag exactly)
#
# H = Hide
#
# lrb = left parenthesis
#
# rrb = right parenthesis
#
# ^ = beginning of the string
#
# $ = end of the string
#
# | = or
#
# \* = any number of
#
# \+ = one or more of
#
# \? = 0 or 1 of
#
# \[ \] = any listed within the brackets
#
# \{ \} = within the listed range
#
# \( \) = parentheses define a group
#
# \d = any numeric character
#
# \D = any non-numeric character
# +
class CurieTemperature(BaseModel):
specifier = StringType()
value = StringType()
units = StringType()
Compound.curie_temperatures = ListType(ModelType(CurieTemperature))
# -
# Define a very basic entity tagger
specifier = (I('curie') + I('temperature') + Optional(lrb | delim) + Optional(R('^T(C|c)(urie)?')) + Optional(rrb) | R('^T(C|c)(urie)?'))('specifier').add_action(join)
units = (R('^[CFK]\.?$'))('units').add_action(merge)
value = (R('^\d+(\.\,\d+)?$'))('value')
# +
# Let the entities be any combination of chemical names, specifier values and units
entities = (chemical_name | specifier | value + units)
# Now create a very generic parse phrase that will match any combination of these entities
curie_temperature_phrase = (entities + OneOrMore(entities | Any()))('curie_temperature')
# List all the entities
curie_temp_entities = [chemical_name, specifier, value, units]
# -
| ipynb/cdesnowball_practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ezwyg4y5Z5S_" colab_type="code" colab={}
# %tensorflow_version 2.x
import tensorflow as tf
import matplotlib.pyplot as plt # data manipulation
import numpy as np # data manipulation
import pandas as pd # data visualisation
from sklearn.datasets import load_boston # dataset
from sklearn.model_selection import train_test_split # to split the data
# + id="NrAdbtAIaHDH" colab_type="code" colab={}
dataset = load_boston()
# + id="7JWQvYghaIGu" colab_type="code" outputId="e1b93339-e0b5-4156-8e5a-828699b79e20" executionInfo={"status": "ok", "timestamp": 1582098334032, "user_tz": -180, "elapsed": 630, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDfSH89H_io39Rxl1rTT_RIw29YUrP-dmPlq0Xi=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 904}
print(dataset.DESCR)
# + id="-TMgkGpXaJkY" colab_type="code" colab={}
columns = np.concatenate((np.copy(dataset.feature_names),['Median (Target)']))
data_df = pd.DataFrame(np.concatenate((dataset.data, dataset.target.reshape(-1,1)), axis = 1), columns = columns )
# + id="hHI55gmdbKoQ" colab_type="code" outputId="5c307d4a-004b-484a-f785-58e96f0d60db" executionInfo={"status": "ok", "timestamp": 1582098594005, "user_tz": -180, "elapsed": 607, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDfSH89H_io39Rxl1rTT_RIw29YUrP-dmPlq0Xi=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 314}
data_df.describe()
# + id="Sg7yOCg6bo7_" colab_type="code" colab={}
# Get the data
X, Y = dataset.data, dataset.target
N, D = X.shape
# + id="EAEbjNLucdUv" colab_type="code" outputId="31e36770-4eb0-42d4-c272-ef3bec9f8296" executionInfo={"status": "ok", "timestamp": 1582098994792, "user_tz": -180, "elapsed": 624, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDfSH89H_io39Rxl1rTT_RIw29YUrP-dmPlq0Xi=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Split the data into training and test set
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = 0.2)
print(X_train.shape, X_test.shape)
# + id="XmTw-4TRb3QD" colab_type="code" outputId="d066699e-a8b1-4297-9c8d-00b371594228" executionInfo={"status": "ok", "timestamp": 1582099834028, "user_tz": -180, "elapsed": 14120, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDfSH89H_io39Rxl1rTT_RIw29YUrP-dmPlq0Xi=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Create the model
model = tf.keras.models.Sequential([
tf.keras.layers.Dense(1, input_shape = (D,)) # we wont use an activation function since this is a regression problem
])
# compile the model with SGD and MSE
# optimizer = adam [CHECKED 3rd], nadam [2nd], rmsprop[1st]
model.compile(optimizer = tf.keras.optimizers.RMSprop(momentum = 0.8), loss = 'mse')
# scheduler
def schedule(epochs, lr):
if epochs >= 100:
return lr - (lr*0.1)
return lr
scheduler = tf.keras.callbacks.LearningRateScheduler(schedule)
# fit the model
r = model.fit(X_train, y_train, validation_data = (X_test, y_test), callbacks = [scheduler], epochs = 400, verbose = 2)
# plot the loss function
plt.plot(r.history['loss'], label = 'loss')
# + id="EbrTni3Ad8Y6" colab_type="code" outputId="6676ca49-fcff-4cee-9866-ed5b45154ef6" executionInfo={"status": "ok", "timestamp": 1582099835302, "user_tz": -180, "elapsed": 570, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDfSH89H_io39Rxl1rTT_RIw29YUrP-dmPlq0Xi=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 419}
preds = model.predict(X_test).flatten()
abs_error = abs(preds - y_test)
pd.DataFrame(np.concatenate((preds.reshape(-1,1),
y_test.reshape(-1,1),
abs_error.reshape(-1,1)), axis = 1), columns = ['Predictions', 'True Values', 'Absolute Error'])
| Tensorflow 2/Simple Regression/Boston Regression TF2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="d1rexij_dFSJ"
# # !pip install catboost
# + id="kclg-UoJcCnq" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="0a716c3c-5111-462f-8c68-fc263bcd3927"
#Standard libraries for data analysis:
import numpy as np
import pandas as pd
pd.options.display.max_columns = None
pd.options.display.max_rows = None
from scipy import stats
from scipy.stats import skew, norm
import statsmodels.api as sm
import joblib
import pickle
# sklearn modules for data preprocessing:
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import make_pipeline
#Modules for Model Selection:
from sklearn import svm, tree, linear_model, neighbors
from sklearn import naive_bayes, ensemble, discriminant_analysis
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
import xgboost as xgb
from xgboost import XGBRegressor
from lightgbm import LGBMRegressor
#from catboost import CatBoostRegressor
from sklearn.linear_model import LinearRegression, SGDRegressor, Lasso, Ridge, ElasticNet
from sklearn.svm import LinearSVR, SVR
from sklearn.ensemble import RandomForestRegressor, AdaBoostRegressor, GradientBoostingRegressor
#sklearn modules for Model Evaluation & Improvement:
from sklearn.metrics import mean_squared_log_error, mean_absolute_error, mean_squared_error, r2_score
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
from sklearn.model_selection import KFold
from sklearn import metrics
#Standard libraries for data visualization:
import seaborn as sns
from matplotlib import pyplot
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import matplotlib
# %matplotlib inline
color = sns.color_palette()
from IPython.display import display
import plotly.offline as py
py.init_notebook_mode(connected=True)
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
import plotly.graph_objects as go
import plotly.express as px
import plotly.tools as tls
import plotly.figure_factory as ff
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="9_tTuME3kRJn"
# # 7.1: Import the dataset 'insurance.csv'.
# - The column 'charges' should be considered the target label.
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 73} id="0pJfP-BvCq3t" outputId="d186cbd7-36c1-46f3-af1d-0144073d02ab"
from google.colab import files
uploaded = files.upload()
# + colab={"base_uri": "https://localhost:8080/", "height": 387} id="-goshUUpC6Zi" outputId="d087fc43-4560-409e-89b3-73635e76de66"
insurance = pd.read_csv('insurance.csv', encoding='Latin-1')
display(insurance.head(), insurance.tail())
# + [markdown] id="aJst7_i3GZI-"
# Columns
#
# - age: age of primary beneficiary
#
# - sex: insurance contractor gender, female, male
#
# - bmi: Body mass index, providing an understanding of body, weights that are relatively high or low relative to height,
# objective index of body weight (kg / m ^ 2) using the ratio of height to weight, ideally 18.5 to 24.9
#
# - children: Number of children covered by health insurance or number of dependents
#
# - smoker: Smoking
#
# - region: the beneficiary's residential area in the US, northeast, southeast, southwest, northwest.
#
# - charges: Individual medical costs billed by health insurance
# + [markdown] id="axrmp_uikrL5"
# # 7.2: Explore the data using at least 3 data exploratory tools of your choosing in pandas.
# + colab={"base_uri": "https://localhost:8080/"} id="sm13nX3TA9Rt" outputId="248481b0-a4a8-490f-9742-860f4e85f7ed"
print(f'The dataset has {insurance.shape[0]} rows and {insurance.shape[1]} columns')
# + colab={"base_uri": "https://localhost:8080/", "height": 277} id="xf_WONXuDy2D" outputId="f7fb8201-b806-4ef3-f07b-4cf35eb8a44b"
display(insurance.info())
# + colab={"base_uri": "https://localhost:8080/", "height": 156} id="fUFeshr30VJU" outputId="db6a9f6f-5acd-43cf-ed67-fd1027ac86bf"
display(insurance.isnull().any())
# + [markdown] id="gB7-XZ1wI7gL"
# P/N: No missing or undefined values
#
# + colab={"base_uri": "https://localhost:8080/", "height": 386} id="dttpgHbFI6-N" outputId="b764f4ae-7771-4f93-9c8c-6082ba88b593"
display(insurance.describe(include='all').round())
# + [markdown] id="aZBh4lrdKYMh"
# P/N:
# - The average age is 39: the youngest is 18, the oldest is 64.
# - Slightly more male participants than females.
# - There's a larger amount of participants from the southeast region.
# - The average BMI is 31: minimum BMI is 16, maximum BMI is 53.
# - Majority of the participants are non-smoker.
# - Average insurance charge is 13,270, with less than 50% participants paying above 9,382.
# + [markdown] id="WNRxYkwGi0xE"
# **Univariate Analysis**
#
# ---
#
# + colab={"base_uri": "https://localhost:8080/", "height": 69} id="4vps3MytmJ_N" outputId="f1ad9521-39a5-458b-cde3-73c45f594e16"
binary_features = insurance.nunique()[insurance.nunique() == 2].keys().tolist()
numeric_features = [col for col in insurance.select_dtypes(['float','int']).columns.tolist() if col not in binary_features]
categorical_features = [col for col in insurance.select_dtypes('object').columns.to_list() if col not in binary_features + numeric_features ]
display(binary_features, numeric_features, categorical_features)
# + [markdown] id="DId7JgTx0vsj"
# Visualising distribution of categorical variables
# + id="h0puKVMMenFW"
def countplot_ratio(x,data,hue=None,ax=None):
ax=sns.countplot(x,data=data,hue=hue,ax=ax)
ax.set_xticklabels(ax.get_xticklabels(),rotation=10)
ax.set_title(x + " Distributions")
ax.legend(bbox_to_anchor=(1.01,1))
total = float(len(data))
for p in ax.patches:
height = p.get_height()
ax.text(p.get_x()+p.get_width()/2.,height + 3, '{:.2f}%'.format((height/total)*100),fontsize=12, weight = 'bold',ha="center")
# + colab={"base_uri": "https://localhost:8080/", "height": 637} id="-ODEa7w6ereS" outputId="0bc13650-e1c4-4139-853f-d4367412daef"
features = binary_features + categorical_features
fig, axes =plt.subplots(1, 3, figsize=(15, 8),)
axes = axes.flatten()
#fig.suptitle('Distribution of Categorical Features', fontsize=16)
for ax, col in zip(axes, features ):
countplot_ratio(col, insurance[features], ax=ax)
plt.tight_layout()
plt.show()
# + [markdown] id="YgXFtUKr1CB0"
# Visualising distribution of numerical vaiables
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="pFPcGkRTh9PH" outputId="b421084f-b278-478e-d5dd-e355b3732892"
fig, axes =plt.subplots(2, 2, figsize=(15, 15),)
axes = axes.flatten()
#fig.suptitle('Distribution of Numerical Features', fontsize=16)
for ax, col in zip(axes, numeric_features):
sns.histplot(data=insurance, x=insurance[col], kde=True, ax=ax, )
ax.set_title(f'{col} Distribution')
plt.tight_layout()
plt.show()
# + [markdown] id="wyzrVviRFEtI"
# **Bivariate Analysis**
#
# ---
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 911} id="z-hRcsd9DPRU" outputId="0a0ec19a-fbf2-4dec-edfd-6e0671ae33ec"
# Visualising distribution of each variable against the target variable
fig, axes =plt.subplots(3, 2, figsize=(25, 20),)
axes = axes.flatten()
#fig.suptitle('Distribution Features Against the Target Variable(Charges)', fontsize=16)
for ax, col in zip(axes, insurance.columns[:-1]):
sns.barplot(y=insurance['charges'], x=insurance[col], ax=ax )
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="z3ehtaDTFKw_" outputId="de16eaf0-ff4a-44a1-c982-c66e5f073d34"
sns.pairplot(insurance,
x_vars=["age", "sex", "smoker", "bmi", "region", "children", "charges", ],
y_vars=["age", "sex", "smoker", "bmi", "region", "children", "charges", ])
# + [markdown] id="fAIw_Y-PNYll"
# - The pair plot is a quick view of the relationshib between the variables.
# - Based on the diagonal plots, a large amount of participants are in their 20s.
# - There are slightly more male participants than female.
# - More than a third of the participants are non-smokers.
# - BMI of all participants seems to be normally distributed, with a mean of approxtimately 30.
# - No. of participants from the different regions are almost uniformly distributed.
# - Most participants are charged below 20,000.
#
# - The Age-Charge graph indicates that as the participant's age increases, the insurance charge increases.
# - From the Gender-BMI graph, it shows that male participants has a wider range of BMI compared to female participants.
# - Smoker-Charges plot displays that smokers have a significantly higher insurance charge compared to non-smoker.
# - On the BMI-Region plot, participants from southeast has a higher BMI value.
# + [markdown] id="or3rxJAShG9V"
# **Correlation**
#
# ---
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 211} id="X8nQpkjwQJN1" outputId="374eed7e-6d74-46bc-b79b-8968311561e2"
# label encoding for binary features
encoded_df = insurance.copy()
le = LabelEncoder()
for feature in binary_features:
encoded_df[feature] = le.fit_transform(encoded_df[feature])
display(feature, np.unique(encoded_df[feature].values))
encoded_df = pd.get_dummies(encoded_df, columns=categorical_features)
display(encoded_df.head(3))
# + colab={"base_uri": "https://localhost:8080/", "height": 613} id="ppRpfvGXQtcR" outputId="8e04ceaf-de16-4cb9-d018-cb5c0ec1f132"
plt.figure(figsize=(20,10))
sns.heatmap(encoded_df.corr(), annot = True, cmap = 'Blues_r')
# + [markdown] id="MjCswDC4cTAv"
# P/N:
# - Number of children (dependents) and gender have correlation at minimal, which is 0.068 and 0.057.
# - As the number of children is the least influencing factor with all the other variables, a correlation range between 0.0077 to 0.068, this variable shall be excluded in most of the data analysis below.
# - Smoker is highly correlated to the insurance charge, with a correlation coefficient of 0.79.
# - Age and BMI are moderately correlated with insurance charge, a correlation coefficient of 0.3 and 0.2. This indicates that the insurance charge will be higher if the person is older or is a smoker or the person has a higher BMI value.
# + [markdown] id="u9CJDEpZrGUi"
# # 7.2(b): Interpret your observation of what form of predictive analysis that can be conducted on the data.
#
# Since the target variable is the continuous variable 'charges', we will need to perform a Regression analysis on it.
# + [markdown] id="jMbSmg6IlCBP"
# # 7.3: Visualize the age distribution for the column 'age' and comment on the results.
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="gFmshprYlPlI" outputId="ea243361-30b2-400e-c7b3-d44c6764e60e"
figure = px.histogram(insurance, x='age', color='smoker', hover_data=insurance.columns, color_discrete_sequence=['indianred', 'indigo'],
template='plotly_dark')
figure.show(renderer='colab')
# + [markdown] id="viEpKQJ0yOTc"
# - The analysis suggests that data in the age column is not normally distrubuted with most of the participants being young adults and therefore the data may be biased.
# + [markdown] id="uQMdeLV9oECE"
# # 7.4: Isolate all the continuous and discrete columns into their respective lists.
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="TBlrkiZURgoP" outputId="078b79e6-d7c4-4fc9-a53b-ad88f3815010"
numerical_discrete = [var for var in numeric_features if len(insurance[var].unique()) < 20]
numerical_continuous = [var for var in numeric_features if var not in numerical_discrete]
display(numerical_discrete, numerical_continuous)
# + [markdown] id="nJMn0NaRxUIG"
# # 7.5: Visually identify if there is presence of any outliers in the numerical_continuous columns and resolve them using a zscore test and a threshold of your choosing.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="KfHUV58c0_i9" outputId="a52a31a4-3584-4511-aeef-9df0150c59c4"
for col in insurance[numerical_continuous].columns:
fig = px.box(insurance, y=insurance[col], title=f'{col} Box Plot Distribution', color_discrete_sequence=['indianred'],
template='plotly_dark' )
fig.show(renderer='colab')
# + [markdown] id="mzeql0pOnZOQ"
# P/N: There are outliers
# + id="TOnWC7v-H9EX"
def outlier_detector(data):
outliers = []
zscore = []
threshold = 3
for i in data:
score = (i - np.mean(data)) / np.std(data)
zscore.append(score)
if np.abs(score) > threshold:
outliers.append(i)
return len(outliers), outliers
# + id="rBHFqh5UIYp6" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="2bd22005-5da7-4560-d617-b01d5f407f04"
num_bmi_outliers, bmi_outliers = outlier_detector(insurance['bmi'])
num_charges_outliers, charges_outliers = outlier_detector(insurance['charges'])
num_age_outliers, age_outliers = outlier_detector(insurance['age'])
display(num_bmi_outliers, bmi_outliers)
display(num_charges_outliers, charges_outliers)
display(num_age_outliers, age_outliers)
# + colab={"base_uri": "https://localhost:8080/"} id="AmwvSZwZ9kQY" outputId="e31706d4-dd59-4b89-fbcb-8ebc2f9dbafa"
for col in insurance[numerical_continuous].columns:
zscore = np.abs(stats.zscore(insurance[col]))
print(f'{col} outlier indexes: {np.where( z > 3 )}')
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="93ZnuyeJEqcT" outputId="d539d4c0-9f1d-412f-fa94-17bb16ec0acc"
zscore = np.abs(stats.zscore(insurance[numerical_continuous]))
display(np.where( zscore > 3 ))
# + id="niRxZ7OwFZh2"
insurance_clean = insurance[(zscore < 3).all(axis=1)]
# + colab={"base_uri": "https://localhost:8080/"} id="dQ2D4D2DATcI" outputId="b12c0023-e1be-408d-9e69-4cce19f5c715"
print(f'Dataset before outlier removal: {insurance.shape}')
print(f'Dataset after outlier removal: {insurance_clean.shape}')
# + [markdown] id="a3uVUqXVOBfs"
# #7.6: Validate that your analysis above was successful by visualizing the value distribution in the resulting columns using an appropriate visualization method.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Bqm34NfzsvSo" outputId="9f7d7a0c-ff37-4092-af5a-70d15aca83a6"
for col in insurance_clean[numerical_continuous].columns:
fig = px.box(insurance_clean, y=insurance_clean[col], title=f'{col} Box Plot Distribution After Cleaning', color_discrete_sequence=['gold'],
template='plotly_dark' )
fig.show(renderer='colab')
# + [markdown] id="niW6izU55YFB"
# - Using the Z-score method we can see that some outliers have been successfully removes however not all have been removed.
# + [markdown] id="_Dg1yheNOMNR"
# # 7.7: Isolate all the categorical column names into a list named 'categorical'.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="hsK3cx41O3in" outputId="a90f1d4b-6ed9-4fbd-a3e8-c967df418857"
categorical = [col for col in insurance.select_dtypes('object').columns]
display(categorical)
# + [markdown] id="rxn-VrBDPtDJ"
# # 7.8: Visually identify the outliers in the discrete and categorical features and resolve them using the combined rare levels method.
# + colab={"base_uri": "https://localhost:8080/", "height": 141} id="SbzFEJdFPsMu" outputId="93ac0930-8256-48df-f826-866596c01940"
features_df = pd.DataFrame()
for col in numerical_discrete + categorical:
features_df[col] = insurance[col]
features_df.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="3C5REYsz7pzS" outputId="ae072521-9c46-4731-fd5f-635fafab7c45"
for col in features_df.columns:
fig = px.bar(data_frame=features_df, x=features_df[col].unique(), y=features_df[col].value_counts(normalize=True),
color_discrete_sequence=['aqua'], template='plotly_dark', title=f'Frequency of {col} Distribution', )
fig.update_layout( xaxis_title = col, yaxis_title=f'Frequency of Observations per {col}',)
fig.show(renderer='colab')
# + id="q9_9jZR6laDF"
def rare_imputation(data, column):
#frequencies = data[column].value_counts(normalize=True, ascending=True)
#frequents = [x for x in frequencies.loc[frequencies > 0.03].index.values]
data[column] = data[column].mask(data[column].map(data[column].value_counts(normalize=True)) < 0.03, 'rare')
return data[column]
# + colab={"base_uri": "https://localhost:8080/"} id="Ng5sR8a2plFK" outputId="79d92a41-c8ff-492a-e512-8739b707fa5c"
for col in features_df.columns:
insurance_clean[col] = rare_imputation(insurance_clean, col)
print(f'{col}: \n {insurance_clean[col].unique()}')
# + [markdown] id="T-ysaQoKnyts"
# # 7.9: Encode the discrete and categorical features with one of the measures of central tendency of your choosing.
# + id="GCOD_jGsnBRV"
'''def target_encoder(df, column, target, index=None, method='median'):
"""
Target-based encoding is numerization of a categorical variables via the target variable. Main purpose is to deal
with high cardinality categorical features without exploding dimensionality. This replaces the categorical variable
with just one new numerical variable. Each category or level of the categorical variable is represented by a
summary statistic of the target for that level.
Args:
df (pandas df): Pandas DataFrame containing the categorical column and target.
column (str): Categorical variable column to be encoded.
target (str): Target on which to encode.
index (arr): Can be supplied to use targets only from the train index. Avoids data leakage from the test fold
method (str): Summary statistic of the target. Mean, median or std. deviation.
Returns:
arr: Encoded categorical column.
"""
index = df.index if index is None else index # Encode the entire input df if no specific indices is supplied
if method == 'mean':
encoded_column = df[column].map(df.iloc[index].groupby(column)[target].mean())
elif method == 'median':
encoded_column = df[column].map(df.iloc[index].groupby(column)[target].median())
elif method == 'std':
encoded_column = df[column].map(df.iloc[index].groupby(column)[target].std())
else:
raise ValueError("Incorrect method supplied: '{}'. Must be one of 'mean', 'median', 'std'".format(method))
return encoded_column'''
# + id="28vAEX8HTLFc"
def target_encoder(data, feature, target='charges'):
ordered_labels = data.groupby([feature])[target].mean().to_dict()
data[feature] = data[feature].map(ordered_labels)
return data[feature]
# + colab={"base_uri": "https://localhost:8080/", "height": 628} id="2nmuKLx5TuwE" outputId="d4a2467a-9d72-4dbb-aafb-814a3148a9c8"
for col in features_df.columns:
insurance_clean[col] = target_encoder(insurance_clean, col, target='charges')
print(f'{col}: \n {insurance_clean[col].value_counts()}')
display(insurance_clean.head(3), insurance_clean.tail(3))
# + [markdown] id="qX7P9miJ_hS4"
# # 7.10. Separate your features from the target appropriately. Narrow down the number of features to 5 using the most appropriate and accurate method. Which feature had to be dropped and what inference would you give as the main contributor of dropping the given feature.
# + [markdown] id="IM8g06JpAd_a"
# **Uncoverring Features Importance**
#
# ---
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 210} id="Epy-_rLZZEGr" outputId="3be737ae-1a51-4748-cbdd-de5223881c8c"
X = insurance_clean.drop(columns='charges')
y = insurance_clean['charges']
display(X.head(3), y.head(3))
# + colab={"base_uri": "https://localhost:8080/"} id="qhbvIDHZY3Wu" outputId="b2fd593c-1ba1-4c3c-f6fc-3814f252d0d7"
from sklearn.linear_model import RidgeCV
ridge = RidgeCV()
rr = ridge.fit(X, y)
r2_score = rr.score(X, y)
print(f'Best alpha : {rr.alpha_}')
print(f'Intercept c = {rr.intercept_}')
print(f'Coefficients m of each of the features = {rr.coef_}')
print(f'coefficient of determination = {r2_score}')
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="46VJdPZnJjN4" outputId="2a1bcbc1-fc81-4171-eb5a-bb91550ac2a2"
coefficients_df = pd.DataFrame({'Feature': X.columns, 'Coefficient':np.transpose(rr.coef_)}).sort_values(by='Coefficient',
ascending=False)
coefficients_df
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="iOl0s52EZ70A" outputId="a73beb03-e4e7-4b8c-fab1-171768786e3b"
figure = px.pie(coefficients_df, names='Feature', values='Coefficient', color='Feature', color_discrete_sequence=px.colors.sequential.RdBu,
title='Ridge Regression Coefficient Weights', template='plotly_dark')
figure.show(renderer='colab')
# + [markdown] id="fPvC438saykG"
# - Notice the absence of 'sex' due to it's negative coeffficients.
# - According to this analysis, 'bmi' has the most weight, while 'sex' has the least weight.
# - I will be dropping the sex feature since it has the least weights implying that it contributes least to the target variable and therefore is the least important feature.
#
# + [markdown] id="lhEFjoLs4J-E"
# #**Assignment 8**
# + [markdown] id="j6iwcoh54HAZ"
# #8.1: Convert the target labels to their respective log values and give 2 reasons why this step may be useful as we train the machine learning model.
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="Yin1o0e0AdAO" outputId="cecdf057-c61b-4934-f078-db34913b41c1"
X = X.drop(columns='sex')
y_log = np.log10(y)
display(X.head(2), y_log.head(2))
# + [markdown] id="ykVtxUizHRwt"
# 1. Reducing Skewness: Logarithmic transformations transform a highly skewed variable into a more normalized dataset. When modeling variables with non-linear relationships, the chances of producing errors may also be skewed negatively. In theory, we want to produce the smallest error possible when making a prediction, while also taking into account that we should not be overfitting the model. Using the logarithm of one or more variables improves the fit of the model by transforming the distribution of the features to a more normally-shaped bell curve i.e it reduces or removes the skewness in the dataset.
#
# 2. Convenience:
# - A transformed scale may be as natural as the original scale and more convenient for a specific purpose (e.g. percentages rather than original data, sines rather than degrees). One important example is standardization.
# - Linear relationships: When looking at relationships between variables, it is often far easier to think about patterns that are approximately linear than about patterns that are highly curved.
# - Additive relationships: Relationships are often easier to analyse when additive rather than multiplicative.
# - Equal spreads: A transformation may be used to produce approximately equal spreads, despite marked variations in level, which again makes data easier to handle and interpret.
# + [markdown] id="cslLV3M8HecS"
# # 8.2: Slice the selected feature columns and the labels into the training and testing set. Also ensure your features are normalized.
# + id="rg_cF44HIGQl"
X_train, X_test, y_train, y_test = train_test_split(X, y_log, test_size=0.2, random_state=42)
# + id="pGV7GvaTLIE2"
scaler = MinMaxScaler()
# transformer = ColumnTransformer([("scaler", scaler, X_train.columns)], remainder="passthrough")
Xtrain_scaled = scaler.fit_transform(X_train)
Xtest_scaled = scaler.transform(X_test)
# + [markdown] id="Z3HECntJMktF"
# # 8.3: Use at least 4 different regression based machine learning methods and use the training and testing cross accuracy and divergence to identify the best model.
# + id="NAUghM8xQRtB"
def val_model(X, y, regressor, quiet=False):
"""
Cross-validates a given model
# Arguments
X: DataFrame, feature matrix
y: Series, target vector
regressor: regression model from scikit-learn
quiet: Boolean, indicates if funcion should print the results
# Returns
Float, r2 validation scores
"""
X = np.array(X)
y = np.array(y)
#pipe = make_pipeline(MinMaxScaler(), regressor)
kfold = KFold(n_splits=5, shuffle=True, random_state=42)
scores = cross_val_score(regressor, X, y, cv=kfold, scoring='r2')
if quiet == False:
print(f"##### {regressor.__class__.__name__} #####")
print(f'Scores: {scores}')
print(f'R2: {scores.mean()} (+/- {scores.std()})')
return scores.mean()
# + id="Jj1rnxv-WoTn"
def getRegR2Scores(X_train, y_train, *reg_list):
"""
Provides the R2 scores for a given list of models
# Arguments
X_train: X_train
y_train: y_train
*reg_list: list of regressors
# Returns
DataFrame, r2 scores
"""
model = []
r2 = []
for reg in reg_list:
model.append(reg.__class__.__name__)
r2.append(val_model(X_train, y_train, reg))
return pd.DataFrame(data=r2, index=model, columns=['R2']).sort_values(by='R2', ascending=False)
# + id="BG7DTjYIZ_f4"
lr = LinearRegression()
lasso = Lasso()
ridge = Ridge()
elastic = ElasticNet()
random = RandomForestRegressor()
svr = SVR()
xgb = XGBRegressor()
sgd = SGDRegressor()
ada = AdaBoostRegressor()
#cat = CatBoostRegressor
#lg = LGBMRegressor
#gb = GradientBoostingRegressor
regressors = [lr, lasso, ridge, elastic, random, svr, xgb, sgd, ada]
# + colab={"base_uri": "https://localhost:8080/", "height": 879} id="R1Elq_YDg6Nl" outputId="828d0a0b-97dd-4ef8-c4aa-bafe1b5bb9aa"
scores_df = getRegR2Scores(Xtrain_scaled, y_train, *regressors)
display(scores_df)
# + [markdown] id="8SMigrgklTVS"
# - From the above analysis the Extreme Gradient Boosting Regressor is the model with the highest average R2 score and I will be using it for my predictive analysis
# + [markdown] id="NhpPr6bxJd8O"
# #8.4: After identifying the best model, train it with the training data again.
# + id="5Xte79w39TCB" colab={"base_uri": "https://localhost:8080/"} outputId="356ecb24-78f1-47ee-c5d0-9b20d1349dec"
model = XGBRegressor(objective='reg:squarederror', n_estimators=500, seed=42)
model.fit(Xtrain_scaled, y_train, eval_set=[(Xtest_scaled, y_test)], verbose=False)
# + [markdown] id="NxRzW6z_7pbA"
# b): Using at least 3 model evaluation metrics in regression, evaluate the models training and testing score. Also ensure as you test the models, the predicted and actual targets have been converted back to the original values.
# + id="Ys2zzss6HdlW"
def rmsle(y_test, y_preds):
return np.sqrt(mean_squared_log_error(y_test, y_preds))
def show_scores(model, X_train, X_test, y_train, y_test):
train_pred = model.predict(X_train)
y_pred = model.predict(X_test)
y_train_exp, y_test_exp, train_pred_exp, y_pred_exp = np.exp(y_train), np.exp(y_test), np.exp(train_pred), np.exp(y_pred)
scores = { "Train MAE": mean_absolute_error(y_train_exp, train_pred_exp),
"Test MAE": mean_absolute_error(y_test_exp, y_pred_exp),
"Train RMSLE": rmsle(y_train_exp, train_pred_exp),
"Test RMSLE": rmsle(y_test_exp, y_pred_exp),
"Train R^2": r2_score(y_train_exp, train_pred_exp),
"Test R^2": r2_score(y_test_exp, y_pred_exp) }
return scores
# + colab={"base_uri": "https://localhost:8080/"} id="VWMroYoOc2lF" outputId="91641372-d75c-42c8-c79f-740df1a400b1"
show_scores(model, Xtrain_scaled, Xtest_scaled, y_train, y_test)
| insurance_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#|hide
#|skip
! [ -e /content ] && pip install -Uqq fastai # upgrade fastai on colab
# +
#|default_exp collab
#default_class_lvl 3
# -
#|export
from __future__ import annotations
from fastai.tabular.all import *
#|hide
from nbdev.showdoc import *
# # Collaborative filtering
#
# > Tools to quickly get the data and train models suitable for collaborative filtering
# This module contains all the high-level functions you need in a collaborative filtering application to assemble your data, get a model and train it with a `Learner`. We will go other those in order but you can also check the [collaborative filtering tutorial](http://docs.fast.ai/tutorial.collab).
# ## Gather the data
#|export
class TabularCollab(TabularPandas):
"Instance of `TabularPandas` suitable for collaborative filtering (with no continuous variable)"
with_cont=False
# This is just to use the internal of the tabular application, don't worry about it.
# +
#|export
class CollabDataLoaders(DataLoaders):
"Base `DataLoaders` for collaborative filtering."
@delegates(DataLoaders.from_dblock)
@classmethod
def from_df(cls, ratings, valid_pct=0.2, user_name=None, item_name=None, rating_name=None, seed=None, path='.', **kwargs):
"Create a `DataLoaders` suitable for collaborative filtering from `ratings`."
user_name = ifnone(user_name, ratings.columns[0])
item_name = ifnone(item_name, ratings.columns[1])
rating_name = ifnone(rating_name, ratings.columns[2])
cat_names = [user_name,item_name]
splits = RandomSplitter(valid_pct=valid_pct, seed=seed)(range_of(ratings))
to = TabularCollab(ratings, [Categorify], cat_names, y_names=[rating_name], y_block=TransformBlock(), splits=splits)
return to.dataloaders(path=path, **kwargs)
@classmethod
def from_csv(cls, csv, **kwargs):
"Create a `DataLoaders` suitable for collaborative filtering from `csv`."
return cls.from_df(pd.read_csv(csv), **kwargs)
CollabDataLoaders.from_csv = delegates(to=CollabDataLoaders.from_df)(CollabDataLoaders.from_csv)
# -
# This class should not be used directly, one of the factory methods should be preferred instead. All those factory methods accept as arguments:
#
# - `valid_pct`: the random percentage of the dataset to set aside for validation (with an optional `seed`)
# - `user_name`: the name of the column containing the user (defaults to the first column)
# - `item_name`: the name of the column containing the item (defaults to the second column)
# - `rating_name`: the name of the column containing the rating (defaults to the third column)
# - `path`: the folder where to work
# - `bs`: the batch size
# - `val_bs`: the batch size for the validation `DataLoader` (defaults to `bs`)
# - `shuffle_train`: if we shuffle the training `DataLoader` or not
# - `device`: the PyTorch device to use (defaults to `default_device()`)
show_doc(CollabDataLoaders.from_df)
# Let's see how this works on an example:
path = untar_data(URLs.ML_SAMPLE)
ratings = pd.read_csv(path/'ratings.csv')
ratings.head()
dls = CollabDataLoaders.from_df(ratings, bs=64)
dls.show_batch()
show_doc(CollabDataLoaders.from_csv)
dls = CollabDataLoaders.from_csv(path/'ratings.csv', bs=64)
# ## Models
# fastai provides two kinds of models for collaborative filtering: a dot-product model and a neural net.
#|export
class EmbeddingDotBias(Module):
"Base dot model for collaborative filtering."
def __init__(self, n_factors, n_users, n_items, y_range=None):
self.y_range = y_range
(self.u_weight, self.i_weight, self.u_bias, self.i_bias) = [Embedding(*o) for o in [
(n_users, n_factors), (n_items, n_factors), (n_users,1), (n_items,1)
]]
def forward(self, x):
users,items = x[:,0],x[:,1]
dot = self.u_weight(users)* self.i_weight(items)
res = dot.sum(1) + self.u_bias(users).squeeze() + self.i_bias(items).squeeze()
if self.y_range is None: return res
return torch.sigmoid(res) * (self.y_range[1]-self.y_range[0]) + self.y_range[0]
@classmethod
def from_classes(cls, n_factors, classes, user=None, item=None, y_range=None):
"Build a model with `n_factors` by inferring `n_users` and `n_items` from `classes`"
if user is None: user = list(classes.keys())[0]
if item is None: item = list(classes.keys())[1]
res = cls(n_factors, len(classes[user]), len(classes[item]), y_range=y_range)
res.classes,res.user,res.item = classes,user,item
return res
def _get_idx(self, arr, is_item=True):
"Fetch item or user (based on `is_item`) for all in `arr`"
assert hasattr(self, 'classes'), "Build your model with `EmbeddingDotBias.from_classes` to use this functionality."
classes = self.classes[self.item] if is_item else self.classes[self.user]
c2i = {v:k for k,v in enumerate(classes)}
try: return tensor([c2i[o] for o in arr])
except KeyError as e:
message = f"You're trying to access {'an item' if is_item else 'a user'} that isn't in the training data. If it was in your original data, it may have been split such that it's only in the validation set now."
raise modify_exception(e, message, replace=True)
def bias(self, arr, is_item=True):
"Bias for item or user (based on `is_item`) for all in `arr`"
idx = self._get_idx(arr, is_item)
layer = (self.i_bias if is_item else self.u_bias).eval().cpu()
return to_detach(layer(idx).squeeze(),gather=False)
def weight(self, arr, is_item=True):
"Weight for item or user (based on `is_item`) for all in `arr`"
idx = self._get_idx(arr, is_item)
layer = (self.i_weight if is_item else self.u_weight).eval().cpu()
return to_detach(layer(idx),gather=False)
# The model is built with `n_factors` (the length of the internal vectors), `n_users` and `n_items`. For a given user and item, it grabs the corresponding weights and bias and returns
# ``` python
# torch.dot(user_w, item_w) + user_b + item_b
# ```
# Optionally, if `y_range` is passed, it applies a `SigmoidRange` to that result.
x,y = dls.one_batch()
model = EmbeddingDotBias(50, len(dls.classes['userId']), len(dls.classes['movieId']), y_range=(0,5)
).to(x.device)
out = model(x)
assert (0 <= out).all() and (out <= 5).all()
show_doc(EmbeddingDotBias.from_classes)
# `y_range` is passed to the main init. `user` and `item` are the names of the keys for users and items in `classes` (default to the first and second key respectively). `classes` is expected to be a dictionary key to list of categories like the result of `dls.classes` in a `CollabDataLoaders`:
dls.classes
# Let's see how it can be used in practice:
model = EmbeddingDotBias.from_classes(50, dls.classes, y_range=(0,5)
).to(x.device)
out = model(x)
assert (0 <= out).all() and (out <= 5).all()
# Two convenience methods are added to easily access the weights and bias when a model is created with `EmbeddingDotBias.from_classes`:
show_doc(EmbeddingDotBias.weight)
# The elements of `arr` are expected to be class names (which is why the model needs to be created with `EmbeddingDotBias.from_classes`)
mov = dls.classes['movieId'][42]
w = model.weight([mov])
test_eq(w, model.i_weight(tensor([42])))
show_doc(EmbeddingDotBias.bias)
# The elements of `arr` are expected to be class names (which is why the model needs to be created with `EmbeddingDotBias.from_classes`)
mov = dls.classes['movieId'][42]
b = model.bias([mov])
test_eq(b, model.i_bias(tensor([42])))
#|export
class EmbeddingNN(TabularModel):
"Subclass `TabularModel` to create a NN suitable for collaborative filtering."
@delegates(TabularModel.__init__)
def __init__(self, emb_szs, layers, **kwargs):
super().__init__(emb_szs=emb_szs, n_cont=0, out_sz=1, layers=layers, **kwargs)
show_doc(EmbeddingNN)
# `emb_szs` should be a list of two tuples, one for the users, one for the items, each tuple containing the number of users/items and the corresponding embedding size (the function `get_emb_sz` can give a good default). All the other arguments are passed to `TabularModel`.
emb_szs = get_emb_sz(dls.train_ds, {})
model = EmbeddingNN(emb_szs, [50], y_range=(0,5)
).to(x.device)
out = model(x)
assert (0 <= out).all() and (out <= 5).all()
# ## Create a `Learner`
# The following function lets us quickly create a `Learner` for collaborative filtering from the data.
#|export
@delegates(Learner.__init__)
def collab_learner(dls, n_factors=50, use_nn=False, emb_szs=None, layers=None, config=None, y_range=None, loss_func=None, **kwargs):
"Create a Learner for collaborative filtering on `dls`."
emb_szs = get_emb_sz(dls, ifnone(emb_szs, {}))
if loss_func is None: loss_func = MSELossFlat()
if config is None: config = tabular_config()
if y_range is not None: config['y_range'] = y_range
if layers is None: layers = [n_factors]
if use_nn: model = EmbeddingNN(emb_szs=emb_szs, layers=layers, **config)
else: model = EmbeddingDotBias.from_classes(n_factors, dls.classes, y_range=y_range)
return Learner(dls, model, loss_func=loss_func, **kwargs)
# If `use_nn=False`, the model used is an `EmbeddingDotBias` with `n_factors` and `y_range`. Otherwise, it's a `EmbeddingNN` for which you can pass `emb_szs` (will be inferred from the `dls` with `get_emb_sz` if you don't provide any), `layers` (defaults to `[n_factors]`) `y_range`, and a `config` that you can create with `tabular_config` to customize your model.
#
# `loss_func` will default to `MSELossFlat` and all the other arguments are passed to `Learner`.
learn = collab_learner(dls, y_range=(0,5))
learn.fit_one_cycle(1)
# ## Export -
#|hide
from nbdev.export import *
notebook2script()
| nbs/45_collab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="bYi4fvHr_brl"
# # [Siamese Neural Networks for One-shot Image Recognition](https://www.cs.cmu.edu/~rsalakhu/papers/oneshot1.pdf)
#
# A Pytorch Implementation
# + colab={"base_uri": "https://localhost:8080/"} id="7zdN5AIzBGim" outputId="7fd4e90f-974e-4ecb-b2ce-90fb206bca71"
# ! pip install torchscan
# + id="bs6uY9OIkbdb"
# https://www.cs.cmu.edu/~rsalakhu/papers/oneshot1.pdf
import torch
import torch.nn as nn
import warnings
warnings.filterwarnings('ignore')
# + id="hL7Y8VLVkPmt"
# To be used as the feature extractor, in order to compare the features
class FeatExt(nn.Module):
def __init__(self):
super(FeatExt, self).__init__()
# input.shape: (1, 105, 105)
self.cnn1 = nn.Conv2d(1, 64, kernel_size=10) # (1, 105, 105) -> (64, 96, 96)
self.relu1 = nn.ReLU()
# feature maps.shape: (64, 96, 96)
self.max_pool1 = nn.MaxPool2d(kernel_size=2) # (64, 96, 96) -> (64, 48, 48)
# feature maps.shape: (64, 48, 48)
self.cnn2 = nn.Conv2d(64, 128, kernel_size=7) # (64, 48, 48) -> (128, 42, 42)
self.relu2 = nn.ReLU()
# feature maps.shape: (128, 42, 42)
self.max_pool2 = nn.MaxPool2d(kernel_size=2) # (128, 42, 42) -> (128, 21, 21)
# feature maps.shape: (128, 21, 21)
self.cnn3 = nn.Conv2d(128, 128, kernel_size=4) # (128, 21, 21) -> (128, 18, 18)
self.relu3 = nn.ReLU()
# feature maps.shape: (128, 18, 18)
self.max_pool3 = nn.MaxPool2d(kernel_size=2) # (128, 18, 18) -> (128, 9, 9)
# feature maps.shape: (128, 9, 9)
self.cnn4 = nn.Conv2d(128, 256, kernel_size=4) # (128, 9, 9) -> (256, 6, 6)
self.relu4 = nn.ReLU()
# feature maps.shape: (256, 6, 6)
# torch.flatten: (256, 6, 6) -> (9216)
# feature maps.shape: (9216)
self.fc1 = nn.Linear(9216, 4096) # (9216) -> (4096)
self.sigmoid1 = nn.Sigmoid()
# # feature maps.shape: (4096)
# self.fc2 = nn.Linear(4096, 1)
# self.sigmoid2 = nn.Sigmoid()
def forward(self, x):
x = self.max_pool1(self.relu1(self.cnn1(x)))
x = self.max_pool2(self.relu2(self.cnn2(x)))
x = self.max_pool3(self.relu3(self.cnn3(x)))
x = self.relu4(self.cnn4(x))
x = torch.flatten(x, start_dim=1)
x = self.sigmoid1(self.fc1(x))
# x = self.sigmoid2(self.fc2(x))
return x
# if __name__ == '__main__':
# model = SiameseNet()
# batch_size = 5
# input = torch.rand(size=(batch_size, 1, 105, 105))
# out = model(input)
# print(input.shape, out.shape)
# assert tuple(out.shape) == (batch_size, 1)
# + id="InYxkyFZkalI"
class SiameseNet(nn.Module):
def __init__(self):
super(SiameseNet, self).__init__()
self.feat = FeatExt()
# feature maps.shape: (4096)
self.fc = nn.Linear(4096, 1)
self.sigmoid = nn.Sigmoid()
def forward(self, x1, x2):
x1 = self.feat(x1)
x2 = self.feat(x2)
return self.sigmoid(self.fc((x1-x2).abs()))
# + [markdown] id="WkrTC_d4HnM4"
# #### Weight Initilization
#
# * They initialized all network weights
# in the convolutional layers from a normal distribution with
# zero-mean and a standard deviation of `10−2`
# * Biases were
# also initialized from a normal distribution, but with mean `0.5` and standard deviation `10−2`
# * In the fully-connected
# layers, the biases were initialized in the same way as the
# convolutional layers, but the weights were drawn from a
# much wider normal distribution with zero-mean and standard deviation `2 × 10−1`
# .
# + id="RNq9JJaWHt4D"
def init_weights(m):
if type(m) == nn.Linear:
nn.init.normal_(m.weight, 0, 2e-1)
nn.init.normal_(m.bias, 0, 1e-2)
elif type(m) == nn.Conv2d:
nn.init.normal_(m.weight, 0, 1e-2)
nn.init.normal_(m.bias, 0.5, 1e-2)
# + colab={"base_uri": "https://localhost:8080/"} id="4Zp5n9_fAj5d" outputId="d5a9805b-16c5-4343-c2e4-dcb7b7ce1eb6"
# Just a quick check that every thing works as it should be
model = SiameseNet()
batch_size = 5
input = torch.rand(size=(batch_size, 1, 105, 105))
out = model(input, input)
print(input.shape, out.shape)
assert tuple(out.shape) == (batch_size, 1)
# + [markdown] id="0NJFd1Qko9_h"
# ## Getting & preparing the dataset
# + id="cBL3dmVL7qxG"
import os
working_dir = '/content/drive/MyDrive/siamese_net'
os.chdir(working_dir)
# + id="FkcuUDsFkd4_"
from torchvision.datasets import Omniglot
from torch.utils.data import Dataset, DataLoader
import numpy as np
import random
import logging
from tqdm.notebook import tqdm
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/"} id="vQrdco5Xkd3U" outputId="0dabd2be-141f-47ea-976f-eeac542d4757"
ds = Omniglot(root='./data', download=True, transform=np.array, background=True)
# + [markdown] id="QBUB0phidhJT"
# Note: For our training, we are creating dataset in such a way, that 1 means both images are similar, while 0 means they are dis-similar
# + id="AQH1IZjXCdCo"
# https://github.com/Rhcsky/siamese-one-shot-pytorch/blob/main/data_loader.py
from collections import defaultdict
class OmniglotDS(Dataset):
def __init__(self, ds):
super(OmniglotDS, self).__init__()
self.ds = ds
self.mean = 0.8444
self.std = 0.5329
self.imgs = defaultdict(list)
self.data = [ds[i] for i in range(1200)]
# self.data = ds
for img, label in tqdm(self.data, desc='Iterating over omniglot'):
self.imgs[label].append(img)
def __len__(self):
return len(self.data)
def __getitem__(self, idx):
img1, cls1 = random.choice(self.data)
# if they are from same class
if idx % 2 == 1:
label = 1.0
img2, cls2 = random.choice(self.imgs[cls1]), cls1
# they are from different class
else:
label = 0.0
options = list(set(self.imgs.keys()) - {cls1})
cls2 = random.choice(options)
img2 = random.choice(self.imgs[cls2])
img1, img2 = np.array(img1, dtype=np.float32) / 255, np.array(img2, dtype=np.float32) / 255
img1 = np.expand_dims(img1, axis=0)
img2 = np.expand_dims(img2, axis=0)
return {
'img1': torch.from_numpy(img1),
'img2': torch.from_numpy(img2),
'label': torch.from_numpy(np.array([label], dtype=np.float32))
}
# + id="--eNnp4ykd2t" colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["72dba319de29415cb0607ffea350a6f4", "081bf5d229214c4f97690aab5a020a66", "737f600da8e5493daca3457b9472a68f", "2fee5ad21a7e49f6a74aa0845d0c8919", "f75539acfa014a9d915fb31718c7f33b", "e12d08cb33b34485a73345dfffc6ae22", "51024be5133f4333bff1a8251e7759fd", "8c75c6fed11e4d81b992c6353784b91d"]} outputId="00ca792f-909b-4d25-dbf7-d841bc53e0a6"
dl = DataLoader(OmniglotDS(ds), batch_size=128)
# + id="U6BMJd6LNfG2" colab={"base_uri": "https://localhost:8080/"} outputId="41b9fb80-ebb5-4e5a-df25-3bcf08e46a20"
len(dl), len(dl.dataset)
# + colab={"base_uri": "https://localhost:8080/", "height": 83, "referenced_widgets": ["e2bda7323e874bb3bde83431dce43e45", "a3531156db644969a322d5828478bdfb", "a75c81c2825f43a1848dfe2bed8d5a88", "1a41fff52744463289976953c7ae1f8e", "2137ba65a20c47d4bae3ac3be3bd87de", "78bcd8aff1c3431e89746b75099a581b", "65c81581ad014e95a6949e97dde5a2f7", "5ca105301b9441dcadd8e07ef5ea524b"]} id="c_aPR_f5e0sL" outputId="4d9fec84-1007-496f-c6b9-269d34631cd9"
for x in OmniglotDS(ds):
print(x['img1'].shape, x['img2'].shape)
break
# + id="8VaMOZhHe0qn"
# + id="A4KkksbBe0pm"
# + [markdown] id="mizEVpSrDYse"
# ## Loss & optimizers
#
# - Binary Cross Entropy Loss with L2 Norm
# + id="0GoTQD7xE4FI"
from torch.optim import Adam
from torchsummary import summary
# + colab={"base_uri": "https://localhost:8080/"} id="bywb-EbVJ-jK" outputId="b8da8145-4066-470d-ae1d-369d7fd7103c"
model.apply(init_weights)
# + id="J8-LvR2DDfon"
device = 'cuda' if torch.cuda.is_available() else 'cpu'
criterian = nn.BCELoss()
optimizer = Adam(model.parameters(), weight_decay=0.1)
model = model.to(device)
from torch.optim.lr_scheduler import StepLR
scheduler = StepLR(optimizer, step_size=1, gamma=0.99)
# + colab={"base_uri": "https://localhost:8080/"} id="WQ8sKm4XKAaL" outputId="a5d42932-71d8-497c-cd8a-c5cadf512800"
summary(model, [(1, 105, 105), (1, 105, 105)])
# + [markdown] id="ocoh3dscKGng"
# ## Training Loop
#
# - LR Schedular
#
# To Do:
# (In Learning Rate Schedule Section): We fixed momentum to start at 0.5 in every layer,
# increasing linearly each epoch until reaching the value µj ,
# the individual momentum term for the jth layer.
# + id="DgcYuyNTKGUB"
NUM_EPOCH, print_every = 301, 50
# + id="RqLrTzPbKGTD" colab={"base_uri": "https://localhost:8080/", "height": 185, "referenced_widgets": ["ccb4dd1ceb51466e8aed132081bcf7b6", "af0b06c2012e4fb4bb2deeedef3d39c1", "b9df851a9ea54f8ebc40189410a2c526", "8013769a845944e68c64cc0363116dd9", "a1758a3c64a2490f85e4c776ff1cfbfe", "128b41ab86404d61979fabb35b00e467", "4ed81237e09649e3ac99401067f4301b", "bc0dc81dd9394da7914d25993978a376"]} outputId="392f0263-8617-4573-df72-d8e779c13cac"
losses = []
for epoch in tqdm(range(NUM_EPOCH), desc='Training...'):
running_loss = 0.0
for batch in dl:
img1, img2, label = batch['img1'].to(device), batch['img2'].to(device), batch['label'].to(device)
optimizer.zero_grad()
out = model(img1, img2)
loss = criterian(out, label)
running_loss += loss.item() * out.shape[0]
loss.backward()
optimizer.step()
scheduler.step()
running_loss = running_loss / len(dl.dataset)
losses.append(running_loss)
if epoch % print_every == 0:
print(f'Epoch: {epoch}, Loss: {running_loss:.2f}')
# + id="n2jwSpbshsSb" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="fb5db71c-6b62-438d-ca9c-c363c2ab7c06"
plt.plot(losses)
plt.title('Train Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
# + id="8WX72rqLcFNN"
torch.save({
'state_dict': model.state_dict()
}, './weights/dict.pth')
# + colab={"base_uri": "https://localhost:8080/"} id="uDWWmp_wb8b3" outputId="21a0e5d9-1eda-4701-e019-213c94a33d5f"
# ! pwd && ls weights
# + [markdown] id="g2dm32MZWPvp"
# ## Visualizations
# + id="tgmvJZR9WSta"
from PIL import Image
from matplotlib import pyplot as plot
from collections import defaultdict
# + id="MQobcrlrWXaQ"
images = defaultdict(list)
# + id="RaUUKM7gWXZk"
for img, label in ds:
images[label].append(img)
if len(images) > 4:
break
# + colab={"base_uri": "https://localhost:8080/", "height": 444} id="0mkOIKElWf2B" outputId="00a636b3-3e98-4f9b-998f-abbc79549c9f"
# display all images of diff class
# create figure
fig = plt.figure(figsize=(10, 7))
# setting values to rows and column variables
rows = 2
columns = 2
for i in range(4):
img = images[i][0]
# Adds a subplot at the 1st position
fig.add_subplot(rows, columns, i+1)
plt.imshow(img)
plt.title(f'Label: {i}')
plt.show()
# + id="av3BjjbPXelX"
assert len(images[0]) >= 4
# + colab={"base_uri": "https://localhost:8080/", "height": 444} id="YEMraZgvXmHx" outputId="8f085731-bea3-40b8-b576-886824521de9"
# display all images of same class
# create figure
fig = plt.figure(figsize=(10, 7))
# setting values to rows and column variables
rows = 2
columns = 2
for i in range(4):
img = images[0][i]
# Adds a subplot at the 1st position
fig.add_subplot(rows, columns, i+1)
plt.imshow(img)
plt.title(f'Label: 0')
plt.show()
# + [markdown] id="LdRHX2GFbEjZ"
# Note that, for our training, output of 1 means, both images are similar, and 0 means both are different (see the dataset we created)
# + colab={"base_uri": "https://localhost:8080/"} id="9kaJFhwGXvxh" outputId="bf1dfc76-1169-4068-df7e-7027fd02f724"
model = model.cpu()
x, y = np.array([images[0][0]], dtype=np.float32), np.array([images[0][1]], dtype=np.float32)
x, y = torch.from_numpy(np.expand_dims(x, axis=0)), torch.from_numpy(np.expand_dims(y, axis=0))
z = np.array([images[2][1]], dtype=np.float32)
z = torch.from_numpy(np.expand_dims(z, axis=0))
print(f'Scores for same class (0) is: {model(x, y).item(): .3f}, and for different classes (0, 2) is: {model(x, z).item(): .3f}')
# + colab={"base_uri": "https://localhost:8080/", "height": 228} id="GS8zW0kKbS0G" outputId="11f46f7b-af73-478c-f2bf-6ca49906d905"
# display all images of same class
# create figure
fig = plt.figure(figsize=(10, 7))
# setting values to rows and column variables
rows = 1
columns = 3
# Adds a subplot at the 1st position
fig.add_subplot(rows, columns, 1)
plt.imshow(images[0][0])
plt.title(f'Label: 0')
fig.add_subplot(rows, columns, 2)
plt.imshow(images[0][1])
plt.title(f'Label: 0')
fig.add_subplot(rows, columns, 3)
plt.imshow(images[2][1])
plt.title(f'Label: 2')
plt.show()
# + id="ONZDYv8Ib55s"
# + [markdown] id="Wbybo3VOERWY"
# ## Rough
# + colab={"base_uri": "https://localhost:8080/"} id="8RRgSTNkESrO" outputId="062d8758-4c39-4ef6-8875-94cf5588a289"
l = nn.BCELoss()
x, y = torch.tensor([[0.01]]), torch.tensor([[1.0]])
print(x, y, l(x, y))
# + id="6uu6avLFSnfA"
from PIL import Image
from matplotlib import pyplot as plot
from collections import defaultdict
# + id="TZy3WWJwTizZ"
images = defaultdict(list)
# + id="RCYSP0uSEU4h"
for img, label in ds:
images[label].append(img)
if len(images) > 4:
break
# + colab={"base_uri": "https://localhost:8080/"} id="pqUOdMX2UGzH" outputId="4c0ee353-78e4-479a-87ad-649b0e947945"
images.keys()
# + id="l3K4iIm5SZL0"
Image.fromarray(x[0]).show()
# + colab={"base_uri": "https://localhost:8080/"} id="wqx-0JHRS4MJ" outputId="26c78b7a-433b-4587-9e02-09a22b1e9ed6"
x[0]
# + id="IyQFg3aITQdL"
| implementations/SiameseNetwork/Siamese_Network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''spacytestenv'': conda)'
# language: python
# name: python3
# ---
# # Location Name Filter
# Processing of locations scraped from OneMap. Code is written to extract relevant names based on the raw data format in data/singapore-postal-codes. Import first JSON, then a function to read the relevant files.
# +
import json
data_path = '../data/singapore-postal-codes/'
def load_data(file):
with open(data_path + file, "r", encoding="utf-8") as f:
data = json.load(f)
return (data)
# -
# A function to properly capitalise location names is placed here to facilitate cleaning location names for all values later.
# Function to properly capitalise and clean values
def properly_capitalise(location_name):
properly_capitalised_name = location_name.lower().title()
return properly_capitalised_name
# # MRT/LRT Station Names Extraction
# Function created to extract station names, based on the raw format of the json they came in
# function to extract mrt & lrt station names
def stn_name_filter(data):
# filters out "possible locations" by zooming in on sub-list
list_of_stns= []
for i in range(len(data)):
list_of_stns.append(json.dumps(data[i]["Possible Locations"]))
# filters out actual station names (14 index after "SEARCHVAL": " to ", "X":) in a loop
count = 0
for each_stn in list_of_stns:
if each_stn != []:
final_name = each_stn[each_stn.find('"SEARCHVAL": "') + 14:each_stn.find('", "X":')]
#replace original entry with actual station name
list_of_stns[count] = final_name.lower()
count += 1
#filters out blank strings
while ('' in list_of_stns):
list_of_stns.remove('')
return list_of_stns
# Extraction, updating and cleaning of LRT station names
#load LRT station names and append missing stations
lrt_data0 = load_data("lrt_stations.json")
uncleaned_lrt_stns = stn_name_filter(lrt_data0) + ['senja lrt station', 'sengkang lrt station', 'punggol lrt station']
#Properly capitalise LRT station names and add to list
lrt_stns = []
for station in uncleaned_lrt_stns:
lrt_stn_name = properly_capitalise(station).replace("Lrt", "LRT")
lrt_stns.append(lrt_stn_name)
lrt_no_stn = lrt_stn_name.replace(" LRT Station", '')
lrt_stns.append(lrt_no_stn)
# Extraction, updating and cleaning of MRT station names
#load MRT station names and append missing stations
mrt_data0 = load_data("mrt_stations.json")
#append missing station names
uncleaned_mrt_stns = stn_name_filter(mrt_data0) + ['woodlands north mrt station', 'woodlands south mrt station', 'springleaf mrt station', 'lentor mrt station', 'mayflower mrt station', 'bright hill mrt station', 'upper thomson mrt station']
#Properly capitalise MRT station names and add to list
mrt_stns = []
for station in uncleaned_mrt_stns:
mrt_stn_name = properly_capitalise(station).replace("Mrt", "MRT")
mrt_stns.append(mrt_stn_name)
mrt_no_stn = mrt_stn_name.replace(" MRT Station", '')
mrt_stns.append(mrt_no_stn)
# # Building Names Extraction
#
# From buildings.json, several categories of location values were selected that are meaningful to everyday identification of building locations. The values for POSTAL, BUILDING, ROAD_NAME and BLK_NO were picked out, correctly capitalised and put together to give the following list of values:
#
# SIMPLE ADDRESS - created from the cleaned values of BLK_NO and ROAD_NAME
# POSTCODE - created by concat of "Singapore" to POSTAL
# BUILDING NAME - created by cleaning BUILDING
# ROAD NAME - created by cleaning ROAD NAME
#
# These lists are segragated to enable future provision to filter them as seperate entity tags, rather than a singular "LOC". For now, they will be concat together into a single JSON file.
#
# Previously, SIMPLE ADDRESS was created by taking SEARCHVAL, then deleting the postcode "SINGAPORE XXXXXX" via RegEx and building name if duplicated. However, this process was not precise enough - certain private estate names were input as the building name in the raw data, which result in filtering out even road names from SIMPLE ADDRESS. This left the filtered data with entries that are only numbers, which messed up the EntityRuler later on in the NER creation process.
buildings_data0 = load_data("buildings.json")
#creating three seperate lists for buildingname, address and postcode. not sure if it is meaningful to split but doing so for now if we need it in the future.
buildings_name_list = []
buildings_address_list = []
buildings_postcode = []
road_names_list = []
def extract_value(value):
value_result = properly_capitalise(json.dumps(buildings_data0[i][value], sort_keys = True).strip('"'))
return value_result
#running through each item in the scraped buildings list
for i in range(len(buildings_data0)):
onemap_postal = "Singapore " + extract_value("POSTAL")
buildings_postcode.append(onemap_postal)
onemap_building = extract_value('BUILDING')
buildings_name_list.append(onemap_building)
onemap_road_name = extract_value('ROAD_NAME')
road_names_list.append(onemap_road_name)
onemap_blk_no = extract_value("BLK_NO")
simple_buildings_address = onemap_blk_no + " " + onemap_road_name
buildings_address_list.append(simple_buildings_address)
#function to remove duplicate items in list
def duplicate_remover(thelist):
org_length = len(thelist)
the_new_list = sorted(list(set(thelist)))
final_length = len(the_new_list)
print(f"Number of unique items reduced from {org_length} to {final_length}")
return the_new_list
#putting unique items into their finalised lists
new_buildings_name_list = duplicate_remover(buildings_name_list)
new_buildings_address_list = duplicate_remover(buildings_address_list)
new_buildings_postcode = duplicate_remover(buildings_postcode)
new_road_names_list = duplicate_remover(road_names_list)
# # Transfer of Data to JSON
# +
save_data_path = "../../data/extracted_locations/"
def save_data(file, data):
with open (save_data_path + file, "w", encoding="utf-8") as f:
json.dump(data, f, indent=4)
save_data('extracted_mrt_stns.json', mrt_stns)
save_data('extracted_lrt_stns.json', lrt_stns)
save_data('extracted_buildings_address_list.json', new_buildings_address_list)
save_data('extracted_buildings_name_list.json', new_buildings_name_list)
save_data('extracted_buildings_postcode.json', new_buildings_postcode)
save_data('extracted_road_names_list.json', new_road_names_list)
| training_scripts/entity_ruler_base_training/onemap_names_filter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Everybody Loves Venn Diagrams
#
# In this notebook we look at the intersections between three catalogues: 20140111!Franzen, RGZ, and Norris. This incarnation of the Franzen was a prerelease catalogue used for RGZ, so everything in Franzen should be in RGZ. Additionally, the component IDs should all match up. We *expect* that for every Norris object, there should be a corresponding Franzen object, modulo the edges of the SWIRE field. If there is not, we need to look at the flux distributions of the different parts of the Venn diagram, as well as the position of the objects on the sky.
import astropy.io.ascii as asc
table = asc.read('one-table-to-rule-them-all.tbl')
table
# Venn diagram time!
rgz = {i['Key'] for i in table if i['Component Zooniverse ID (RGZ)']}
norris = {i['Key'] for i in table if i['Component # (Norris)']}
franzen = {i['Key'] for i in table if i['Component ID (Franzen)']}
# +
import matplotlib.pyplot as plt
from matplotlib_venn import venn3
import matplotlib.patches
# %matplotlib inline
def plot_venn():
plt.figure(figsize=(8, 8))
v = venn3([rgz, norris, franzen], set_labels=None, set_colors=('r', 'g', 'b'))
plt.legend(handles=[
matplotlib.patches.Patch(color='r', alpha=0.5, label='RGZ'),
matplotlib.patches.Patch(color='g', alpha=0.5, label='Norris'),
matplotlib.patches.Patch(color='b', alpha=0.5, label='Franzen'),
], loc='best')
plt.show()
plot_venn()
# -
# So RGZ is a proper(!) subset of Franzen, and Norris is *not* a subset of Franzen. Let's plot the integrated flux distributions for each subset.
import seaborn, numpy
# +
# Overall flux distribution.
# Columns: Component Int flux (Norris), Component S (Franzen)
fluxes = []
for nflux, fflux in zip(table['Component Int flux (Norris)'], table['Component S (Franzen)']):
if fflux:
fluxes.append(fflux)
elif nflux:
fluxes.append(nflux)
else:
continue
seaborn.distplot(numpy.log(fluxes))
plt.title('Overall Flux Distribution')
plt.xlabel('log $S$')
plt.ylabel('Number Density')
# +
# Norris & Franzen flux distribution.
norris_franzen_fluxes = []
for row in table:
if row['Key'] not in norris & franzen:
continue
nflux, fflux = row['Component Int flux (Norris)'], row['Component S (Franzen)']
if fflux:
norris_franzen_fluxes.append(fflux)
elif nflux:
norris_franzen_fluxes.append(nflux)
else:
continue
seaborn.distplot(numpy.log(norris_franzen_fluxes), color='brown')
plt.title('Norris $\cap$ Franzen Flux Distribution')
plt.xlabel('log $S$')
plt.ylabel('Number Density')
# +
# Norris - Franzen flux distribution.
norris_no_franzen_fluxes = []
for row in table:
if row['Key'] not in norris - franzen:
continue
nflux, fflux = row['Component Int flux (Norris)'], row['Component S (Franzen)']
if fflux:
norris_no_franzen_fluxes.append(fflux)
elif nflux:
norris_no_franzen_fluxes.append(nflux)
else:
continue
seaborn.distplot(numpy.log(norris_no_franzen_fluxes), color='green')
plt.title('Norris \\ Franzen Flux Distribution')
plt.xlabel('log $S$')
plt.ylabel('Number Density')
# +
# Franzen - Norris flux distribution.
franzen_no_norris_fluxes = []
for row in table:
if row['Key'] not in franzen - norris:
continue
nflux, fflux = row['Component Int flux (Norris)'], row['Component S (Franzen)']
if fflux:
franzen_no_norris_fluxes.append(fflux)
elif nflux:
franzen_no_norris_fluxes.append(nflux)
else:
continue
seaborn.distplot(numpy.log(franzen_no_norris_fluxes))
plt.title('Franzen \\ Norris Flux Distribution')
plt.xlabel('log $S$')
plt.ylabel('Number Density')
# +
# Franzen - RGZ flux distribution.
franzen_no_rgz_fluxes = []
for row in table:
if row['Key'] not in franzen - rgz:
continue
nflux, fflux = row['Component Int flux (Norris)'], row['Component S (Franzen)']
if fflux:
franzen_no_rgz_fluxes.append(fflux)
elif nflux:
franzen_no_rgz_fluxes.append(nflux)
else:
continue
seaborn.distplot(numpy.log(franzen_no_rgz_fluxes), color='blue')
plt.title('Franzen \\ RGZ Flux Distribution')
plt.xlabel('log $S$')
plt.ylabel('Number Density')
# -
# Let's plot the positions on the sky of each subset.
# +
def decimalify(ras, decs):
from astropy.coordinates import SkyCoord
coords = []
for ra, dec in zip(ras, decs):
sc = SkyCoord(ra=ra, dec=dec, unit=('hourangle', 'deg'))
coords.append((sc.ra.deg, sc.dec.deg))
return zip(*coords)
plt.scatter(table[sorted(franzen - rgz - norris)]['Component RA (Franzen)'],
table[sorted(franzen - rgz - norris)]['Component DEC (Franzen)'],
color='lightblue', marker='+', label='Franzen \\ (RGZ $\cup$ Norris)')
plt.scatter(table[sorted((franzen - rgz) & norris)]['Component RA (Franzen)'],
table[sorted((franzen - rgz) & norris)]['Component DEC (Franzen)'],
color='blue', marker='+', label='(Franzen \\ RGZ) $\cap$ Norris')
plt.scatter(table[sorted(rgz - norris)]['Component RA (Franzen)'],
table[sorted(rgz - norris)]['Component DEC (Franzen)'],
color='pink', marker='+', label='RGZ \\ Norris')
plt.scatter(table[sorted(rgz & norris)]['Component RA (Franzen)'],
table[sorted(rgz & norris)]['Component DEC (Franzen)'],
color='hotpink', marker='+', label='RGZ $\cap$ Norris')
plt.scatter(*decimalify(table[sorted(norris - franzen)]['Component Radio RA (Norris)'],
table[sorted(norris - franzen)]['Component Radio dec (Norris)']),
color='green', marker='+', label='Norris \\ RGZ')
plt.legend(loc=9, bbox_to_anchor=(0.5, -0.1), ncol=3)
# -
ras, decs = decimalify(table[sorted(norris - franzen)]['Component Radio RA (Norris)'], table[sorted(norris - franzen)]['Component Radio dec (Norris)'])
import astropy.table
norris_not_franzen = astropy.table.Table(data=[ras, decs, [0.05] * len(ras), [0.05] * len(ras)],
names=['ra', 'dec', 'w', 'h'])
for row in norris_not_franzen:
print('CROSS', row['ra'], row['dec'], row['w'], row['h'])
| notebooks/101_everybody_loves_venn_diagrams.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install memory_profiler
# %load_ext memory_profiler
# +
import random
import time
import math
import numpy as np
from sklearn.neighbors import NearestNeighbors
from scipy.spatial import KDTree
from scipy.stats import wasserstein_distance
import torch
import torch.nn as nn
from torch.nn import init
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from torch.autograd.variable import Variable
from torch.utils.data import DataLoader
# -
#device = torch.device('cuda') if torch.cuda.is_available else torch.device('cpu')
device = torch.device('cpu')
# +
set_dist = []
for i in range(50):
m = torch.distributions.multivariate_normal.MultivariateNormal(torch.zeros(2), torch.eye(2))
x = m.sample([250])
set_dist.append(x)
for i in range(50):
m = torch.distributions.multivariate_normal.MultivariateNormal(torch.tensor([0.0, 1.0]), torch.tensor([[1,.5],[.5,1]]))
x = m.sample([250])
set_dist.append(x)
for i in range(50):
m = torch.distributions.multivariate_normal.MultivariateNormal(torch.ones(2), covariance_matrix=torch.tensor([[.7,.1],[.1,1]]))
x = m.sample([250])
set_dist.append(x)
for i in range(50):
m = torch.distributions.multivariate_normal.MultivariateNormal(torch.tensor([1.0, 0.0]), torch.tensor([[.2, -.1], [-.1, 1]]))
x = m.sample([250])
set_dist.append(x)
for i in range(50):
m = torch.distributions.multivariate_normal.MultivariateNormal(torch.tensor([.5, .5]), torch.tensor([[.8,.4],[.4,1]]))
x = m.sample([250])
set_dist.append(x)
for i in range(50):
m = torch.distributions.multivariate_normal.MultivariateNormal(torch.tensor([-.25, -.5]), torch.eye(2)*.5)
x = m.sample([250])
set_dist.append(x)
# -
set_dist = torch.stack(set_dist)
set_dist.shape
class Set2Set(nn.Module):
def __init__(self, input_dim, hidden_dim, act_fn=nn.Tanh, num_layers=1):
'''
Args:
input_dim: input dim of Set2Set.
hidden_dim: the dim of set representation, which is also the INPUT dimension of
the LSTM in Set2Set.
This is a concatenation of weighted sum of embedding (dim input_dim), and the LSTM
hidden/output (dim: self.lstm_output_dim).
'''
super(Set2Set, self).__init__()
self.input_dim = input_dim
self.hidden_dim = hidden_dim
self.num_layers = num_layers
if hidden_dim <= input_dim:
print('ERROR: Set2Set output_dim should be larger than input_dim')
# the hidden is a concatenation of weighted sum of embedding and LSTM output
self.lstm_output_dim = hidden_dim - input_dim
self.lstm = nn.LSTM(hidden_dim, input_dim, num_layers=num_layers, batch_first=True)
# convert back to dim of input_dim
# self.pred = nn.Linear(hidden_dim, input_dim)
self.pred = nn.Linear(hidden_dim,4)
self.act = act_fn()
def forward(self, embedding):
'''
Args:
embedding: [batch_size x n x d] embedding matrix
Returns:
aggregated: [batch_size x d] vector representation of all embeddings
'''
batch_size = embedding.size()[0]
n = embedding.size()[1]
hidden = (torch.zeros(self.num_layers, batch_size, self.lstm_output_dim).cuda(),
torch.zeros(self.num_layers, batch_size, self.lstm_output_dim).cuda())
q_star = torch.zeros(batch_size, 1, self.hidden_dim).cuda()
for i in range(n):
# q: batch_size x 1 x input_dim
q, hidden = self.lstm(q_star, hidden)
# e: batch_size x n x 1
e = embedding @ torch.transpose(q, 1, 2)
a = nn.Softmax(dim=1)(e)
r = torch.sum(a * embedding, dim=1, keepdim=True)
q_star = torch.cat((q, r), dim=2)
q_star = torch.squeeze(q_star, dim=1)
out = self.act(self.pred(q_star))
return out
class DeepSet(nn.Module):
def __init__(self, in_features, set_features):
super(DeepSet, self).__init__()
self.in_features = in_features
self.out_features = set_features
self.feature_extractor = nn.Sequential(
nn.Linear(in_features, 50),
nn.ELU(inplace=True),
nn.Linear(50, 100),
nn.ELU(inplace=True),
nn.Linear(100, set_features)
)
self.regressor = nn.Sequential(
nn.Linear(set_features, 30),
nn.ELU(inplace=True),
nn.Linear(30, 30),
nn.ELU(inplace=True),
nn.Linear(30, 10),
nn.ELU(inplace=True),
nn.Linear(10, 2),
)
def forward(self, input):
x = input
x = self.feature_extractor(x)
x = x.sum(dim=1)
x = self.regressor(x)
return x
class Encoder(nn.Module):
""" Set Encoder
"""
def __init__(self, dim_Q, dim_K, dim_V, d_model, num_heads, ln=False, skip=True):
super(Encoder, self).__init__()
self.dim_V = dim_V
self.num_heads = num_heads
self.skip = skip
# self.s_max = s_max
#Maximum set size
self.d_model = d_model
self.fc_q = nn.Linear(dim_Q, d_model)
self.fc_k = nn.Linear(dim_K, d_model)
self.fc_v = nn.Linear(dim_K, d_model)
if ln:
self.ln0 = nn.LayerNorm(d_model)
self.ln1 = nn.LayerNorm(d_model)
#This is the classic pointwise feedforward in "Attention is All you need"
self.ff = nn.Sequential(
nn.Linear(d_model, 4 * d_model),
nn.ReLU(),
nn.Linear(4 * d_model, d_model))
# I have experimented with just a smaller version of this
# self.fc_o = nn.Linear(d_model,d_model)
# self.fc_rep = nn.Linear(s_max, 1)
#number of heads must divide output size = d_model
def forward(self, Q, K):
Q = self.fc_q(Q)
K, V = self.fc_k(K), self.fc_v(K)
dim_split = self.d_model // self.num_heads
Q_ = torch.cat(Q.split(dim_split, 2), 0)
K_ = torch.cat(K.split(dim_split, 2), 0)
V_ = torch.cat(V.split(dim_split, 2), 0)
A = torch.softmax(Q_.bmm(K_.transpose(-2,-1))/math.sqrt(self.d_model), dim=-1)
A_1 = A.bmm(V_)
O = torch.cat((A_1).split(Q.size(0), 0), 2)
O = torch.cat((Q_ + A_1).split(Q.size(0), 0), 2) if getattr(self, 'skip', True) else \
torch.cat((A_1).split(Q.size(0), 0), 2)
O = O if getattr(self, 'ln0', None) is None else self.ln0(O)
# O = O + F.relu(self.fc_o(O)) if getattr(self, 'skip', None) is None else F.relu(self.fc_o(O))
# For the classic transformers paper it is
O = O + self.ff(O)
O = O if getattr(self, 'ln1', None) is None else self.ln1(O)
O = torch.mean(O,dim=1)
# O = pad_sequence(O, batch_first=True, padding_value=0)
# O = O.transpose(-2,-1)
# O = F.pad(O, (0, self.s_max- O.shape[-1]), 'constant', 0)
# O = self.fc_rep(O)
# O = self.fc_rep(O.transpose(-2,-1))
# O = O.squeeze()
return O
class SelfAttention(nn.Module):
def __init__(self, dim_in=18, dim_out=8, num_heads=2, ln=True, skip=True):
super(SelfAttention, self).__init__()
self.Encoder = Encoder(dim_in, dim_in, dim_in, dim_out, num_heads, ln=ln, skip=skip)
def forward(self, X):
return self.Encoder(X, X)
# +
eps = 1e-15
"""Approximating KL divergences between two probability densities using samples.
It is buggy. Use at your own peril
"""
def knn_distance(point, sample, k):
""" Euclidean distance from `point` to it's `k`-Nearest
Neighbour in `sample` """
norms = np.linalg.norm(sample-point, axis=1)
return np.sort(norms)[k]
def verify_sample_shapes(s1, s2, k):
# Expects [N, D]
assert(len(s1.shape) == len(s2.shape) == 2)
# Check dimensionality of sample is identical
assert(s1.shape[1] == s2.shape[1])
def naive_estimator(s1, s2, k=1):
""" KL-Divergence estimator using brute-force (numpy) k-NN
s1: (N_1,D) Sample drawn from distribution P
s2: (N_2,D) Sample drawn from distribution Q
k: Number of neighbours considered (default 1)
return: estimated D(P|Q)
"""
verify_sample_shapes(s1, s2, k)
n, m = len(s1), len(s2)
D = np.log(m / (n - 1))
d = float(s1.shape[1])
for p1 in s1:
nu = knn_distance(p1, s2, k-1) # -1 because 'p1' is not in 's2'
rho = knn_distance(p1, s1, k)
D += (d/n)*np.log((nu/rho)+eps)
return D
def scipy_estimator(s1, s2, k=1):
""" KL-Divergence estimator using scipy's KDTree
s1: (N_1,D) Sample drawn from distribution P
s2: (N_2,D) Sample drawn from distribution Q
k: Number of neighbours considered (default 1)
return: estimated D(P|Q)
"""
verify_sample_shapes(s1, s2, k)
n, m = len(s1), len(s2)
d = float(s1.shape[1])
D = np.log(m / (n - 1))
nu_d, nu_i = KDTree(s2).query(s1, k)
rho_d, rhio_i = KDTree(s1).query(s1, k+1)
# KTree.query returns different shape in k==1 vs k > 1
if k > 1:
D += (d/n)*np.sum(np.log(nu_d[::, -1]/rho_d[::, -1]))
else:
D += (d/n)*np.sum(np.log(nu_d/rho_d[::, -1]))
return D
def skl_estimator(s1, s2, k=1):
""" KL-Divergence estimator using scikit-learn's NearestNeighbours
s1: (N_1,D) Sample drawn from distribution P
s2: (N_2,D) Sample drawn from distribution Q
k: Number of neighbours considered (default 1)
return: estimated D(P|Q)
"""
verify_sample_shapes(s1, s2, k)
n, m = len(s1), len(s2)
d = float(s1.shape[1])
D = np.log(m / (n - 1))
s1_neighbourhood = NearestNeighbors(k+1, 10).fit(s1)
s2_neighbourhood = NearestNeighbors(k, 10).fit(s2)
for p1 in s1:
s1_distances, indices = s1_neighbourhood.kneighbors([p1], k+1)
s2_distances, indices = s2_neighbourhood.kneighbors([p1], k)
rho = s1_distances[0][-1]
nu = s2_distances[0][-1]
D += (d/n)*np.log(nu/rho)
return D
# List of all estimators
Estimators = [naive_estimator, scipy_estimator, skl_estimator]
# -
class SinkhornDistance(nn.Module):
r"""
Given two empirical measures each with :math:`P_1` locations
:math:`x\in\mathbb{R}^{D_1}` and :math:`P_2` locations :math:`y\in\mathbb{R}^{D_2}`,
outputs an approximation of the regularized OT cost for point clouds.
Args:
eps (float): regularization coefficient
max_iter (int): maximum number of Sinkhorn iterations
reduction (string, optional): Specifies the reduction to apply to the output:
'none' | 'mean' | 'sum'. 'none': no reduction will be applied,
'mean': the sum of the output will be divided by the number of
elements in the output, 'sum': the output will be summed. Default: 'none'
Shape:
- Input: :math:`(N, P_1, D_1)`, :math:`(N, P_2, D_2)`
- Output: :math:`(N)` or :math:`()`, depending on `reduction`
"""
def __init__(self, eps, max_iter, reduction='none'):
super(SinkhornDistance, self).__init__()
self.eps = eps
self.max_iter = max_iter
self.reduction = reduction
def forward(self, x, y):
# The Sinkhorn algorithm takes as input three variables :
C = self._cost_matrix(x, y) # Wasserstein cost function
x_points = x.shape[-2]
y_points = y.shape[-2]
if x.dim() == 2:
batch_size = 1
else:
batch_size = x.shape[0]
# both marginals are fixed with equal weights
mu = torch.empty(batch_size, x_points, dtype=torch.float,
requires_grad=False).fill_(1.0 / x_points).to(device).squeeze()
nu = torch.empty(batch_size, y_points, dtype=torch.float,
requires_grad=False).fill_(1.0 / y_points).to(device).squeeze()
u = torch.zeros_like(mu).to(device)
v = torch.zeros_like(nu).to(device)
# To check if algorithm terminates because of threshold
# or max iterations reached
actual_nits = 0
# Stopping criterion
thresh = 1e-1
# Sinkhorn iterations
for i in range(self.max_iter):
u1 = u # useful to check the update
u = self.eps * (torch.log(mu+1e-8) - torch.logsumexp(self.M(C, u, v), dim=-1)) + u
v = self.eps * (torch.log(nu+1e-8) - torch.logsumexp(self.M(C, u, v).transpose(-2, -1), dim=-1)) + v
err = (u - u1).abs().sum(-1).mean()
actual_nits += 1
if err.item() < thresh:
break
U, V = u, v
# Transport plan pi = diag(a)*K*diag(b)
pi = torch.exp(self.M(C, U, V))
# Sinkhorn distance
cost = torch.sum(pi * C, dim=(-2, -1))
if self.reduction == 'mean':
cost = cost.mean()
elif self.reduction == 'sum':
cost = cost.sum()
# return cost, pi, C
return cost
def M(self, C, u, v):
"Modified cost for logarithmic updates"
"$M_{ij} = (-c_{ij} + u_i + v_j) / \epsilon$"
return (-C + u.unsqueeze(-1) + v.unsqueeze(-2)) / self.eps
@staticmethod
def _cost_matrix(x, y, p=2):
"Returns the matrix of $|x_i-y_j|^p$."
x_col = x.unsqueeze(-2)
y_lin = y.unsqueeze(-3)
C = torch.sum((torch.abs(x_col - y_lin)) ** p, -1)
return C
@staticmethod
def ave(u, u1, tau):
"Barycenter subroutine, used by kinetic acceleration through extrapolation."
return tau * u + (1 - tau) * u1
sinkhorn = SinkhornDistance(eps=0.1, max_iter=100, reduction=None).to(device)
class MyDataset(Dataset):
def __init__(self, data, transform=None):
self.data = data.float()
self.transform = transform
def __getitem__(self, index):
x = self.data[index]
if self.transform:
x = self.transform(x)
return x
def __len__(self):
return len(self.data)
dataset = MyDataset(set_dist)
loader = DataLoader(dataset, batch_size = 12, shuffle = True)
# +
model = DeepSet(2, 36).to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
checkpoint = torch.load('normal_2D_2condition1.pt')
model.load_state_dict(checkpoint['model_state_dict'])
optimizer.load_state_dict(checkpoint['optimizer_state_dict'])
loss = checkpoint['loss']
model.train()
# -
# Wasserstein distance has the following properties:
# 1) W(aX,aY) = |a|W(X,Y)
# 2) W(X+x, Y+x) = W(X,Y)
#
# Only implement these properties
# +
num_epochs = 500
running_loss = []
for t in range(num_epochs):
for n_batch, batch in enumerate(loader):
n_data = Variable(batch.to(device), requires_grad=True)
a = torch.rand(1).to(device)
b = torch.rand(2).to(device)
optimizer.zero_grad()
y = model(n_data)
y_a = model(a*n_data)
y_translate = model(n_data + b)
loss = 0
for i in range(len(batch)):
for j in range(i+1,len(batch)):
y_ij = torch.norm(y[i]-y[j], p=2)
w_ij = sinkhorn(n_data[i],n_data[j])
ya_ij = torch.norm(y_a[i]-y_a[j], p=2)
y_translate_ij = torch.norm(y_translate[i]-y_translate[j], p=2)
diff_translate_ij = torch.norm(y_translate[i]-y[j], p=2)**2
loss += torch.norm(y_ij-w_ij, p=2) + (ya_ij-a*y_ij)**2 + (y_translate_ij- y_ij)**2
del w_ij
#TODO FIX THE LAST TERMS WITH PAIRWISE DISTANCES (SEE PYTORCH CODE)
loss = loss/(len(batch)*(len(batch)-1)/2)
loss.backward()
optimizer.step()
running_loss.append(loss)
print(loss)
# -
# 196+41 epochs in
torch.save({
'model_state_dict': model.state_dict(),
'optimizer_state_dict': optimizer.state_dict(),
'loss': loss
}, 'normal_2D_2condition1.pt')
len(running_loss)
running_loss
# +
#Test ground truth
#Cov mat_1 = ID, Cov mat_2 = [[1,.5], [.5,1]], m_1 = (0,0) , m_2 = (0,1)
#Real Wass dist^2 = ||m_1 - m_2||^2 + (4-\sqrt(2)-\sqrt(6))
# -
m = torch.distributions.multivariate_normal.MultivariateNormal(torch.zeros(2), torch.eye(2))
m1 = m.sample([250]).view(1,-1,2)
m2 = m.sample([250]).view(1,-1,2)
n = torch.distributions.multivariate_normal.MultivariateNormal(torch.tensor([0.0, 1.0]), torch.tensor([[1,.5],[.5,1]]))
n1 = n.sample([250]).view(1,-1,2)
n2 = n.sample([250]).view(1,-1,2)
model(m1)
model(m2)
model(m1*.5)
model(n1*.5)
model(n1)
# +
#calculated distance = 1.336, scaling by .5 get distance to be .7 and moving them around got 1.323
# -
model(m1+.8)
model(n1+.8)
sinkhorn(m1+.5, n1+.5)
sinkhorn(m1,n1)
sinkhorn(m1*.5, n1*.5)
| experiments_with_wasserstein_metrics/normal_encoders-2D-2conditions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # K Nearest Neighbors Project - Solution
#
# Welcome to the KNN Project! This will be a simple project very similar to the lecture, except you'll be given another data set. Go ahead and just follow the directions below.
# ## Import Libraries
# **Import pandas,seaborn, and the usual libraries.**
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# ## Get the Data
# ** Read the 'KNN_Project_Data csv file into a dataframe **
df = pd.read_csv('KNN_Project_Data')
# **Check the head of the dataframe.**
df.head()
# # EDA
#
# Since this data is artificial, we'll just do a large pairplot with seaborn.
#
# **Use seaborn on the dataframe to create a pairplot with the hue indicated by the TARGET CLASS column.**
# THIS IS GOING TO BE A VERY LARGE PLOT
sns.pairplot(df,hue='TARGET CLASS',palette='coolwarm')
# # Standardize the Variables
#
# Time to standardize the variables.
#
# ** Import StandardScaler from Scikit learn.**
from sklearn.preprocessing import StandardScaler
# ** Create a StandardScaler() object called scaler.**
scaler = StandardScaler()
# ** Fit scaler to the features.**
scaler.fit(df.drop('TARGET CLASS',axis=1))
# **Use the .transform() method to transform the features to a scaled version.**
scaled_features = scaler.transform(df.drop('TARGET CLASS',axis=1))
# **Convert the scaled features to a dataframe and check the head of this dataframe to make sure the scaling worked.**
df_feat = pd.DataFrame(scaled_features,columns=df.columns[:-1])
df_feat.head()
# # Train Test Split
#
# **Use train_test_split to split your data into a training set and a testing set.**
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(scaled_features,df['TARGET CLASS'],
test_size=0.30)
# # Using KNN
#
# **Import KNeighborsClassifier from scikit learn.**
from sklearn.neighbors import KNeighborsClassifier
# **Create a KNN model instance with n_neighbors=1**
knn = KNeighborsClassifier(n_neighbors=1)
# **Fit this KNN model to the training data.**
knn.fit(X_train,y_train)
# # Predictions and Evaluations
# Let's evaluate our KNN model!
# **Use the predict method to predict values using your KNN model and X_test.**
pred = knn.predict(X_test)
# ** Create a confusion matrix and classification report.**
from sklearn.metrics import classification_report,confusion_matrix
print(confusion_matrix(y_test,pred))
print(classification_report(y_test,pred))
# # Choosing a K Value
# Let's go ahead and use the elbow method to pick a good K Value!
#
# ** Create a for loop that trains various KNN models with different k values, then keep track of the error_rate for each of these models with a list. Refer to the lecture if you are confused on this step.**
# +
error_rate = []
# Will take some time
for i in range(1,40):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train,y_train)
pred_i = knn.predict(X_test)
error_rate.append(np.mean(pred_i != y_test))
# -
# **Now create the following plot using the information from your for loop.**
plt.figure(figsize=(10,6))
plt.plot(range(1,40),error_rate,color='blue', linestyle='dashed', marker='o',
markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
# ## Retrain with new K Value
#
# **Retrain your model with the best K value (up to you to decide what you want) and re-do the classification report and the confusion matrix.**
# +
# NOW WITH K=30
knn = KNeighborsClassifier(n_neighbors=30)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
print('WITH K=30')
print('\n')
print(confusion_matrix(y_test,pred))
print('\n')
print(classification_report(y_test,pred))
# -
# # Great Job!
#
#
#
| Scikit/03-K-Nearest-Neighbors/03-K Nearest Neighbors Project - Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="sWQHUy3sA2Ir"
# This script fine-tunes and evaluates DA-RoBERTa, DA-BERT, DA-BART, and DA-T5 on the BABE dataset by 5-fold cross-validation:
#
# Required data to run this script:
# - BABE.xlsx
# - the pretrained model that should be evaluated (selected model from https://drive.google.com/drive/folders/1-A1hGKeu-27X9I4ySkja5vMlVscnF8GR?usp=sharing)
# + id="2FbVfplXA-Nf" outputId="b45a868e-b678-47f5-94ff-3063b37893c7"
# !pip install transformers
# !pip install openpyxl
import pandas as pd
import numpy as np
import torch
import torch.nn as nn
import json
import io
import sys
import random
import openpyxl
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split,StratifiedKFold
from sklearn.metrics import roc_auc_score,f1_score,precision_score,recall_score,accuracy_score,confusion_matrix
import transformers
from transformers import AdamW,BertTokenizer,BertModel,RobertaTokenizer,RobertaModel,T5EncoderModel,T5Tokenizer,BartModel,BartTokenizer
from torch.utils.data import DataLoader,TensorDataset,RandomSampler
# + [markdown] id="6VWxBejP6T3E"
# **Create model architecture** (Uncomment respective model which should be evaluated)
# -
# **RoBERTa**
# + id="k5OstJYpMw5I"
# class RobertaClass(torch.nn.Module):
# def __init__(self):
# super(RobertaClass, self).__init__()
# self.roberta = RobertaModel.from_pretrained("roberta-base")
# self.vocab_transform = torch.nn.Linear(768, 768)
# self.dropout = torch.nn.Dropout(0.2)
# self.classifier1 = torch.nn.Linear(768,2)
# def forward(self, input_ids, attention_mask):
# output_1 = self.roberta(input_ids=input_ids, attention_mask=attention_mask)
# hidden_state = output_1[0]
# pooler = hidden_state[:, 0]
# pooler = self.vocab_transform(pooler)
# pooler = self.dropout(pooler)
# output = self.classifier1(pooler)
# return output
# -
# **BERT**
# +
# class BertClass(torch.nn.Module):
# def __init__(self):
# super(BertClass, self).__init__()
# self.bert = BertModel.from_pretrained("bert-base-uncased")
# self.vocab_transform = torch.nn.Linear(768, 768)
# self.dropout = torch.nn.Dropout(0.2)
# self.classifier1 = torch.nn.Linear(768,2)
# def forward(self, input_ids, attention_mask):
# output_1 = self.bert(input_ids=input_ids, attention_mask=attention_mask)
# hidden_state = output_1[0]
# pooler = hidden_state[:, 0]
# pooler = self.vocab_transform(pooler)
# pooler = self.dropout(pooler)
# output = self.classifier1(pooler)
# return output
# -
# **T5**
# +
#create model
# class T5Class(torch.nn.Module):
# def __init__(self):
# super(T5Class, self).__init__()
# self.T5 = T5EncoderModel.from_pretrained("t5-base")
# self.vocab_transform = torch.nn.Linear(768, 768)
# self.dropout = torch.nn.Dropout(0.2)
# self.classifier1 = nn.Linear(768,2)
# def forward(self, input_ids, attention_mask):
# output_1 = self.T5(input_ids=input_ids, attention_mask=attention_mask)
# hidden_state = output_1[0]
# pooler = hidden_state[:, 0]
# pooler = self.vocab_transform(pooler)
# pooler = self.dropout(pooler)
# output = self.classifier1(pooler)
# return output
# -
# **BART**
# +
# #create model
# class BartClass(torch.nn.Module):
# def __init__(self):
# super(BartClass, self).__init__()
# self.bart = BartModel.from_pretrained("facebook/bart-base")
# self.vocab_transform = torch.nn.Linear(768, 768)
# self.dropout = torch.nn.Dropout(0.2)
# self.classifier1 = nn.Linear(768,2)
# def forward(self, input_ids, attention_mask):
# output_1 = self.bart(input_ids=input_ids, attention_mask=attention_mask)
# hidden_state = output_1[0]
# pooler = hidden_state[:, 0]
# pooler = self.vocab_transform(pooler)
# pooler = self.dropout(pooler)
# output = self.classifier1(pooler)
# return output
# + [markdown] id="uNRsqqn26YOu"
# **Connect to GPU**
# + id="CVAo1cx4J7RT" outputId="8a700457-85c8-4ba2-e1f1-fb2e72b3f277"
if torch.cuda.is_available():
device = torch.device("cuda")
print(f'There are {torch.cuda.device_count()} GPU(s) available.')
print('Device name:', torch.cuda.get_device_name(0))
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
# + [markdown] id="dC7ABPwV6jH5"
# **Load pre-trained domain-adapted weights/parameters for the model:** You might have to adapt the path pointing to the domain-adapted model
# + id="C7YkR_bhyd6P"
#load weights of pretrained news model
#weight_dict = torch.load('Roberta.bin')
#weight_dict = torch.load('BERT.bin')
#weight_dict = torch.load('T5.bin')
#weight_dict = torch.load('BART.bin')
#load saved classifier weights + classifier bias --> we use same parameters for the final classification of all models to achieve maximum comparability
classifier_weights = torch.load('../input/domainadaptivepretrainingjcdl/classifier.weights.pt')
classifier_bias = torch.load('../input/domainadaptivepretrainingjcdl/classifier.bias.pt')
#insert weights and bias into weight dict
weight_dict['classifier1.weight'] = classifier_weights
weight_dict['classifier1.bias'] = classifier_bias
# + [markdown] id="WktGY1UUmztM"
# **Load BABE Data:** You might have to adapt the path again
# + id="Aq7SkwILbrO9" outputId="36c41f76-6aa1-4fb3-a9b5-f29875ad2194"
df = pd.read_excel("BABE.xlsx")
df = df[df['label_bias']!= 'No agreement']
df['Label_bias_0-1'] = df['label_bias'].map({'Biased':1,'Non-biased':0})
df.head(3)
# + [markdown] id="HqbHOVdl7aJh"
# **Define Cross-Validation,Tokenizer,Batch Size,Epochs,Loss, and Seeds**
# + id="NOkpsCuGuQK3" outputId="626c1b1c-54ef-4248-a80f-26c03c502456"
np.random.seed(2018)
torch.manual_seed(2018)
random.seed(2018)
torch.cuda.manual_seed_all(2018)
random.seed(2018)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
kfold = StratifiedKFold(n_splits = 3,shuffle = True,random_state=2)
#tokenizer = RobertaTokenizer.from_pretrained('roberta-base')
#tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
#tokenizer = T5Tokenizer.from_pretrained('t5-base')
#tokenizer = BartTokenizer.from_pretrained('facebook/bart-base')
cross_entropy = nn.CrossEntropyLoss()
epochs = 10 #we implement an early stopping criterion. Fine-tuning is actually not done for 10 epochs
batch_size = 32
# + [markdown] id="GUoIFq7n7ekg"
# **Define functions for fine-tuning and validation**
# + id="oA6Np8VLxbuF"
def train(model):
model.train()
total_loss = 0
for batch in train_dataloader:
optim_dbert.zero_grad()
batch = [r.to(device) for r in batch]
sent_id, mask, labels = batch
outputs = model(sent_id, attention_mask=mask)
loss = cross_entropy(outputs,labels)
total_loss = total_loss+loss.item()
loss.backward()
optim_dbert.step()
avg_loss = total_loss / len(train_dataloader)
return avg_loss
# + id="7VvtdBe3xj0h"
def validate(model):
model.eval()
total_loss = 0
print("\n Validating...")
for batch in test_dataloader:
batch = [r.to(device) for r in batch]
sent_id, mask, labels = batch
with torch.no_grad():
outputs = model(sent_id, attention_mask=mask)
loss = cross_entropy(outputs,labels)
total_loss = total_loss+loss.item()
avg_loss = total_loss / len(test_dataloader)
return avg_loss
# + id="krCHuXqsxn5l"
#combine train and validate function: get train and validation loss for every cross-validation split, save best-performing model, and get predictions on the held-out test set to calculate evaluation metrics
def train_validate_pred(model):
best_valid_loss = float('inf')
# empty lists to store training and validation loss of each epoch
train_losses=[]
valid_losses=[]
#for each epoch
for epoch in range(epochs):
print('\n Epoch {} / {}'.format(epoch+1,epochs))
#train model
train_loss = train(model)
#evaluate model
valid_loss = validate(model)
#save the best model
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
global model_dbert
torch.save(model.state_dict(), 'saved_weights.pt')
#if validation loss increases, stop training
elif valid_loss >= best_valid_loss:
print("\n Validation loss not decreased, Model of previous epoch saved")
break
print(f'\n Training Loss: {train_loss:.3f}')
print(f' Validation Loss: {valid_loss:.3f}')
#predict
path = 'saved_weights.pt'
model.load_state_dict(torch.load(path))
with torch.no_grad():
preds = model(test_seq.to(device), test_mask.to(device))
preds = preds.detach().cpu().numpy()
preds = np.argmax(preds, axis = 1)
#save results
loss.append(best_valid_loss)
acc.append(accuracy_score(test_y,preds))
auc.append(roc_auc_score(test_y,preds))
micro_F1.append(f1_score(test_y,preds,average='micro'))
macro_F1_weighted.append(f1_score(test_y,preds,average='weighted'))
binary_F1.append(f1_score(test_y,preds,average='binary'))
precision.append(precision_score(test_y,preds))
recall.append(recall_score(test_y,preds))
conf_matrix = confusion_matrix(test_y, preds)
conf_matrices.append(conf_matrix)
# + id="GhKHZ2S7Vxhx" outputId="0d3218ce-c57f-437c-99c9-2b214e219f00"
#implement cross validation + train/validate/predict
loss = []
acc = []
auc = []
micro_F1 = []
macro_F1_weighted = []
binary_F1 = []
precision = []
recall = []
conf_matrices = []
for fold, (train_index, test_index) in enumerate(kfold.split(df['text'], df['Label_bias_0-1'])):
sys.stdout.write('\n \r Fold {} / {}\n'.format(fold+1,kfold.get_n_splits()))
#divide data into folds
train_text = df['text'].iloc[train_index]
test_text = df['text'].iloc[test_index]
train_labels = df['Label_bias_0-1'].iloc[train_index]
test_labels = df['Label_bias_0-1'].iloc[test_index]
#encode
train_encodings = tokenizer(train_text.tolist(), truncation=True, padding=True)
test_encodings = tokenizer(test_text.tolist(), truncation=True, padding=True)
#convert input to tensors
train_seq = torch.tensor(train_encodings['input_ids'])
train_mask = torch.tensor(train_encodings['attention_mask'])
train_y = torch.tensor(train_labels.tolist())
test_seq = torch.tensor(test_encodings['input_ids'])
test_mask = torch.tensor(test_encodings['attention_mask'])
test_y = torch.tensor(test_labels.tolist())
# wrap tensors into one dataset
train_data = TensorDataset(train_seq, train_mask, train_y)
test_data = TensorDataset(test_seq, test_mask, test_y)
#define dataloader
train_sampler = RandomSampler(train_data)
test_sampler = RandomSampler(test_data)
train_dataloader = DataLoader(train_data,sampler= train_sampler, batch_size=batch_size)
test_dataloader = DataLoader(test_data,sampler = test_sampler, batch_size=batch_size)
#create model instance with pre-trained weights and optimizer: insert respective model that is to be fine-tuned/evaluated
# model = BertClass()
# model = RobertaClass()
# model = BartClass()
# model = T5Class()
model.load_state_dict(weight_dict)
model.to(device)
optim_dbert = AdamW(model.parameters(), lr=1e-5)
#call train/validate/predict function
train_validate_pred(model)
# + id="2oVjkc1rwW85"
#compute cross-validated performance metrics
cv_loss = sum(loss)/len(loss)
cv_acc = sum(acc)/len(acc)
cv_auc = sum(auc)/len(auc)
cv_micro_f1 = sum(micro_F1)/len(micro_F1)
cv_macro_f1 = sum(macro_F1_weighted)/len(macro_F1_weighted)
sd = np.std(macro_F1_weighted)
cv_binary_f1 = sum(binary_F1)/len(binary_F1)
cv_prec = sum(precision)/len(precision)
cv_recall = sum(recall)/len(recall)
cv_conf_matrix = np.mean(conf_matrices, axis=0)
print("CV Accuracy = {}".format(round(cv_acc,4)))
print("CV AUC = {}".format(round(cv_auc,4)))
print("CV Micro F1 = {}".format(round(cv_micro_f1,4)))
print("CV Macro F1 weighted = {}".format(round(cv_macro_f1,4)))
print("SD = {}".format(round(sd,4)))
print("CV Binary F1 = {}".format(round(cv_binary_f1,4)))
print("CV Precision = {}".format(round(cv_prec,4)))
print("CV Recall = {}".format(round(cv_recall,4)))
print("CV Loss = {}".format(round(cv_loss,4)))
# + id="ANOV_r5ZGWyV"
#optionally save metrics in dict
#Roberta_DA_SG2_bs64_lr1e5_6ep = {"loss":cv_loss,"micro_f1":cv_micro_f1,"macro_f1":cv_macro_f1,"SD":sd,"binary_f1":cv_binary_f1,"prec":cv_prec,"recall":cv_recall}
#store metrics in json format
# with open('./Roberta_DA_SG2_bs64_lr1e5_6ep.json', 'w') as f:
# json.dump(Roberta_DA_SG2_bs64_lr1e5_6ep, f)
# + [markdown] id="bvJIY6pC-y0E"
# **McNemar test for statistical significance based on last cv split**
# + id="YzTY47jf-4Sb"
from mlxtend.evaluate import mcnemar,mcnemar_table
# + id="DTTUBpXxASAc"
#get predictions for model on test set. Insert the domain-adapted model you want to evaluate here. Predictions are provided in the repository and do not have to be computed separately
# with torch.no_grad():
# preds_DA = model(test_seq.to(device), test_mask.to(device))
# preds_DA = preds_DA.detach().cpu().numpy()
# preds_DA = np.argmax(preds_DA, axis = 1)
#optionally store predictions
#np.save("preds_T5_DA.npy",preds_T5_DA)
# +
# #get predictions for baseline model. Insert the baseline model you want to evaluate here.
# with torch.no_grad():
# preds_noDA = model(test_seq.to(device), test_mask.to(device))
# preds_noDA = preds_noDA.detach().cpu().numpy()
# preds_noDA = np.argmax(preds_noDA, axis = 1)
#optionally store predictions
# np.save("preds_T5_noDA.npy",preds_T5_noDA)
# + id="okzJKIcT-_Mh" outputId="03f2037a-630a-4ae2-a3bf-dd1be7fe8b6d"
# load predictions for baseline and domain-adapted model and get contingency table. Predictions are provided in the repository and do not have to be computed separately
preds_noDA = np.load("preds_T5_noDA.npy") #path might have to be adapted
preds__DA = np.load("preds_T5_DA.npy")
tb = mcnemar_table(y_target=np.array(test_labels),
y_model1=preds_roberta_noDA,
y_model2=preds_roberta_DA)
print(tb)
# + id="mSszecmS_gk-" outputId="1b78c5b7-1370-4be4-bc1f-e131398ff254"
#calculate McNemar test statistic
chi2, p = mcnemar(ary=tb, corrected=True)
print('chi-squared:', chi2)
print('p-value:', p)
# + id="V_s36EwTGkLJ"
| fine-tune-and-evaluate-domain-adaptive-pretraining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
import numpy as np
import pandas as pd
import pandas_profiling
import json
import os
import matplotlib
#matplotlib.use('TkAgg')
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.stats
from scipy.stats import norm
from datetime import datetime
# %matplotlib inline
pd.options.display.max_columns = None
import statsmodels.api as sm # Provides cross-sectional models and methods
from statsmodels.graphics.api import abline_plot # Plots line given intercept and slope
from sklearn.metrics import mean_squared_error, r2_score # Rates how close regression line is to data
from sklearn.model_selection import train_test_split # Splits data into training and test groups
from sklearn import linear_model, preprocessing # Ordinary least square regression and standardizes the data
import warnings # For handling error messages.
# +
os.chdir(r'C:\Users\2bogu\OneDrive\Desktop\Sringboard_Materials\capstone2\data\external')
df = pd.read_csv('Green_Jobs_-_Green_New_York__GJGNY__Residential_Loan_Portfolio___Beginning_November_2010.csv')
# -
# # Initial Cleaning
df.columns
df = df.drop(['LAST PAYMENT AMOUNT', 'LAST PAYMENT DATE', 'FIRST PAYMENT DATE',
'DELINQUENT AMOUNT', 'DAYS DELINQUENT', 'Reporting Period',
'CREDIT SCORE CURRENT HIGH', 'NEXT PAYMENT DUE DATE', 'PAYMENTS REMAINING',
'PROPERTY COUNTY', 'UTILITY', 'INSTALLATION CITY', 'INSTALLATION ZIP',
'Contractor Name', 'Georeference', 'UNAPPLIED CASH', 'TOTAL LATE CHARGE PAID',
'CURRENT BALANCE', 'TOTAL INTEREST PAID', 'Payments Made'], axis=1)
df.info()
# +
df['CANCEL REASON'].fillna('NONE', inplace = True)
df['TYPE OF BANKRUPTCY'].fillna('NONE', inplace = True)
df['SUCCESSOR NUMBER'].fillna('NONE', inplace = True)
# +
# indicates if loan has a cosigner
df['co-signed'] = np.where(pd.notnull(df['CREDIT SCORE NAME 2']), 1, 0)
# averages credit score feature to get rid of nans in credit score 2
df['avg_credit_score'] = df[['CREDIT SCORE', 'CREDIT SCORE NAME 2']].mean(axis=1)
# fills remaining na values with average of averages
df['avg_credit_score'].fillna(df['avg_credit_score'].mean(), inplace=True)
df.drop(['CREDIT SCORE','CREDIT SCORE NAME 2'], axis=1, inplace=True)
# -
df['ACCOUNT CODE'].value_counts()
# FILTERING FOR FINISHED LOANS
df = df.loc[df['ACCOUNT CODE'].str.contains('Hold|Canceled|Terminated|Bankruptcy|Full', regex=True)]
# creates dependent var
#df['bad'] = df['CANCEL REASON'].str.contains('NONE')
df['bad'] = df['ACCOUNT CODE'].str.contains('Hold|Canceled|Terminated|Bankruptcy', regex=True)
df['bad'] = df['bad'].apply(lambda x: 1 if x==True else 0)
df.describe()
# removing outlier
df = df[df['DEBT TO INCOME'] < 30]
df.describe()
df['bad'].sum()
df['CANCEL REASON'].value_counts()
# cancelations due to death
91 / 439
# no data collected on health
df = df[df['CANCEL REASON'] != 'DEATH']
df['bad'].sum()
df['SUCCESSOR NUMBER'].value_counts()
dfb = df[df['bad']==1]
dfb['SUCCESSOR NUMBER'].value_counts()
dfb[dfb['SUCCESSOR NUMBER']!='NONE'].count()
# proportion of original bad loans that are from successors
34/439
# ony dealing with people who went through the application process
df = df[df['SUCCESSOR NUMBER']=='NONE']
#all 'customer defaulted' loans were successors
df['CANCEL REASON'].value_counts()
# maybe needed
#df['Reporting Period'] = pd.to_datetime(df['Reporting Period'], format="%m/%d/%Y")
df['CONTRACT DATE'] = pd.to_datetime(df['CONTRACT DATE'], format="%m/%d/%Y")
df['SETUP DATE'] = pd.to_datetime(df['SETUP DATE'], format="%m/%d/%Y")
df['ORIGINAL MATURITY DATE'] = pd.to_datetime(df['ORIGINAL MATURITY DATE'], format="%m/%d/%Y")
df['CURRENT MATURITY DATE'] = pd.to_datetime(df['CURRENT MATURITY DATE'], format="%m/%d/%Y")
df['ACCOUNT CODE DATE'] = pd.to_datetime(df['ACCOUNT CODE DATE'], format="%m/%d/%Y")
# +
#df['s_y'] = df['SETUP DATE'].apply(lambda x: x.year)
# +
#df[df['s_y']==2020]
# -
# # A little EDA
# +
def diff_month(d1, d2):
return abs((d1.year - d2.year) * 12) + abs(d1.month - d2.month)
def to_quart(d):
qu = math.ceil(d.month/4)
# -
def time_active(df):
if (df['ACCOUNT CODE']=='Paid in Full') | (df['bad']==1):
return diff_month(df['ACCOUNT CODE DATE'], df['SETUP DATE'])
else:
return df['Months Since Origination']
# to find out how long it took for loans get current account code
# can only tell when a loan was cancelled, NOT how long a good loan has been active
# for age of good loans use Months Since Origination
df['active_m'] = df.apply(lambda x: time_active(x), axis=1)
# + [markdown] heading_collapsed=true
# ## All
# + hidden=true
df.describe()
# + hidden=true
sns.heatmap(df.corr())
# + hidden=true
df.hist(figsize=(10,10))
# + [markdown] heading_collapsed=true
# ## Good
# + hidden=true
df_g = df[df['bad']==0]
# + hidden=true
df_g.describe()
# + hidden=true
sns.heatmap(df_g.corr())
# + hidden=true
df_g.hist(figsize=(10,10))
# + [markdown] heading_collapsed=true
# ## Bad
# + hidden=true
df_b = df[df['bad']==1]
# + hidden=true
df_b.describe()
# + hidden=true
sns.heatmap(df_b.corr())
# + hidden=true
df_b.hist(figsize=(10,10))
# -
# # Defining the positive Class
import math
df_b['y'] = df_b['ACCOUNT CODE DATE'].apply(lambda x: x.year)
df_b['y'].hist()
#loans that have gone bad in 2020
df_b[df_b['y']==2020]['active_m'].hist()
#oans that have gone bad before 2020
df_b[df_b['y']!=2020]['active_m'].hist()
# ##### The economic impact of COVID could cause people who would otherwise had paid off their loans to slip into bad standing. This data represents only 5 months into 2020.
#
# ___H-null___ : There is no difference in how long it took for a loan to become 'bad' loans between 2020 and other years ___Rejected(?)___
#
# - doubling y_2020 to simulate a full year
# +
y_2020 = df_b[df_b['y']==2020]['active_m']
y_2020_ = y_2020.copy()
y_2020 = y_2020.append(y_2020_, ignore_index=True)
y_19 = df_b[df_b['y']!=2020]['active_m']
scipy.stats.ttest_ind(y_2020, y_19)
# -
# ##### Its right on the line. Doubling the amount of bad loans doubles both loans that have gone bad due to covid as well as loans that would have gone bad anyway. In theory this mean the t-stat and p-value would be stronger. They will be kept in this copy
#
#
# # Defining the Negative class
# + [markdown] heading_collapsed=true
# ## Determining threshold for loans of good standing
# + [markdown] heading_collapsed=true hidden=true
# ### DOES NOT APPLY TO THIS COPY
#
# ##### If we only include completed loans the model will not generalize well because it will be fit only on outliers. These are 10-20 year loans and we only have 9 years of data. The model must be able to distinguish between a typical loan and one that will fail. It won't be able to do this if it only has extraordinary cases to train on. When does it become safe to assume a loan will be paid off?
# + hidden=true
df_b = df[df['bad']==1]
# + hidden=true
import math
df_b['y'] = df_b['ACCOUNT CODE DATE'].apply(lambda x: x.year)
# + hidden=true
df_b['active_m'].hist()
#q = df[df['bad']==1]['active_m'].quantile(0.95)
# + hidden=true
bad_mean = np.mean(df_b.active_m)
bad_std = np.std(df_b.active_m, ddof=1)
bad_mean, bad_std
# + hidden=true
df_b.active_m.quantile(q=0.99)
# + hidden=true
1 - norm.cdf(95, loc=bad_mean, scale=bad_std)
# + hidden=true
len(df[(df['bad'] == 0) & (df['active_m'] >= 95)])
# + hidden=true
# + [markdown] hidden=true
# df = df[(df['bad'] == 1) |
# (df['ACCOUNT CODE'] == 'Paid in Full') |
# ((df['bad'] == 0) & (df['active_m'] >= 99))]
# + hidden=true
len(df)
# + [markdown] hidden=true
# ##### The number of months it takes for loans to fail is not normally distributed, so it was boostrapped with the expected mean falling within the confidence interval of the true mean. According to that re-sampled data, there is P=0.006 chance that a loan will be canceled at or after the 99th percentile, 95 months. We will keep loans that have been active for 95 months or more to give the model a sense of what a typical profile is, not just ones that are repaid quickly.
#
#
# ##### This ends up adding 1038 loans for a totla of 5503
#
#
# ##### Could potentially also exclude loans that were paid off a little too quickly...
#
# -
# ## Defining threshold for paid off loans
#
# ##### A ten year loan thats paid off in just a couple of months is generally atypical, except in this dataset. If there was 3rd party involvement to make this happen, and if this played a role in determining if the loan was approved or not is unknown. There is an abundance of paid off loans compared to canceled loans and 'safe' long standing loans so defining a threshold seems to make sense.
#
# ### Sample pop = Paid off loans
df_g = df[df['ACCOUNT CODE']=='Paid in Full']
df_g['active_m'].hist()
# +
good_mean = np.mean(df_g.active_m)
good_std = np.std(df_g.active_m, ddof=1)
good_mean, good_std
# -
df_g.active_m.quantile(0.025)
norm.cdf(0, loc=good_mean, scale=good_std)
len(df[((df['ACCOUNT CODE'] == 'Paid in Full') & (df['active_m'] < 1))])
# ##### The p value is just barely acceptable and removes 49 loans that were paid off before a month had passed
#
# ##### MAKING NO ALTERATIONS THIS COPY
# df = df[(df['ACCOUNT CODE']!= 'Paid in Full') |
# ((df['ACCOUNT CODE'] == 'Paid in Full') & (df['active_m'] >= 1))]
# # A little more EDA
df_b = df[(df['bad'] == 1)]
df_g = df[df['bad'] != 1]
# ##### Based on the histograms of each group bellow it seems it could be better to create a flag for original term length
# + [markdown] heading_collapsed=true
# ## Redoing the initial EDA, not relevent this copy
# + [markdown] hidden=true
# ### All
# + hidden=true
df.describe()
# + hidden=true
sns.heatmap(df.corr())
# + hidden=true
df.hist(figsize=(10,10))
# + [markdown] hidden=true
# ### Good
# + hidden=true
#df_g.describe()
# + hidden=true
sns.heatmap(df_g.corr())
# + hidden=true
df_g.hist(figsize=(10,10))
# + [markdown] hidden=true
# ### Bad
# + hidden=true
#df_b.describe()
# + hidden=true
sns.heatmap(df_b.corr())
# + hidden=true
df_b.hist(figsize=(10,10))
# + hidden=true
# -
# ## Category EDA
df.select_dtypes(include='object').columns
# +
objs = ['Purpose','Loan Type', 'Underwriting', 'Pledged']
for o in objs:
plt.figure()
sns.catplot(y=o, data=df, kind='count', hue='bad')
plt.show()
# -
for o in objs:
print(o)
print(df[o].value_counts())
print('')
# +
# one instance of a 30 debt to income ratio is removed
df = df[df['DEBT TO INCOME'] < 30]
# -
# ## Scatter plots
plt.figure(figsize=(10,10))
sns.scatterplot(x='avg_credit_score', y='DEBT TO INCOME', data=df, hue='bad')
plt.figure(figsize=(10,10))
sns.scatterplot(x='avg_credit_score', y='ORIGINAL LOAN AMOUNT', data=df, hue='bad')
plt.figure(figsize=(10,10))
sns.scatterplot(x='avg_credit_score', y='PAYMENT AMOUNT', data=df, hue='bad')
plt.figure(figsize=(10,10))
sns.violinplot(x='INTEREST RATE', y='avg_credit_score', data=df, hue='bad')
plt.figure(figsize=(10,10))
sns.violinplot(x='ORIGINAL TERM', y='avg_credit_score', data=df, hue='bad')
df.columns
# +
bx = ['ORIGINAL LOAN AMOUNT','ORIGINAL TERM','INTEREST RATE','DEBT TO INCOME','PAYMENT AMOUNT']
for b in bx:
pd.DataFrame(df[b]).boxplot()
plt.show()
# -
# # Category Binarification
# +
# CREATES BINARY COLUMN FOR PLEDGED VS UNPLEDGED LOANS
df['unpledged'] = df['Pledged'].apply(lambda x: 1 if x == 'Unpledged' else 0)
# DROPS ORIGINAL PLEDGED COLUMN
df.drop('Pledged', axis=1, inplace=True)
# +
# CREATES FLAG FOR UNDERWRITTING T1, ONLY T1 AND T2 IN SET
df['underwritten_t1'] = df['Underwriting'].apply(lambda x: 1 if x == 'Tier 1' else 0)
df.drop('Underwriting', axis=1, inplace=True)
# +
# CREATES BOOL FOR LOAN TYPE, 0 = ON BILL RECOVERY
df['loan_type_smart_energy'] = df['Loan Type'].apply(lambda x: 1 if x == 'Smart Energy' else 0)
df.drop('Loan Type', axis= 1, inplace=True)
# +
# CREATES BOOL FOR PURPOSE, DOUBLE ZERO IS OTHER
df['purpose_ee'] = df['Purpose'].apply(lambda x: 1 if x == 'Energy Efficiency (EE)' else 0)
df['purpose_sol'] = df['Purpose'].apply(lambda x: 1 if x == 'Solar (PV)' else 0)
df.drop('Purpose', axis=1, inplace=True)
# -
df.drop([
'CONTRACT DATE', 'SETUP DATE', 'ORIGINAL MATURITY DATE',
'CURRENT MATURITY DATE', 'ACCOUNT CODE', 'ACCOUNT CODE DATE',
'CANCEL REASON', 'TYPE OF BANKRUPTCY','active_m','Months Since Origination' #,'y'
],
axis=1,
inplace=True)
df['ORIGINAL TERM'].value_counts()
# +
# dtype int, but is a category. tricky tricky
df['term_180'] = df['ORIGINAL TERM'].apply(lambda x: 1 if x==180 else 0)
df['term_120'] = df['ORIGINAL TERM'].apply(lambda x: 1 if x==120 else 0)
df['term_60'] = df['ORIGINAL TERM'].apply(lambda x: 1 if x==60 else 0)
df.drop('ORIGINAL TERM',axis=1,inplace=True)
# -
df.info()
(sum(df.bad))
#/len(df)
#
len(df)- sum(df.bad)
# # Conclusion
#
# ##### Rows: 4813
# ##### Columns: 15 : 5 numeric, 10 binary categorical leave one out
# ##### Positive Class: 314(%6.5) : All loans that have been canceled by the original clients of GJGNY for a reason other than death or put on hold, and did not go bad in 2020
# ##### Negative Class: 4499: All loans paid off after one month had passed and loans in good standing that have been active for 95 months or longer
df.to_csv(r'C:\Users\2bogu\OneDrive\Desktop\Sringboard_Materials\capstone2\data\interim\fl', index = False)
| notebooks/Final Version/Cleaning&EDA_2_fl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decompose Prediction
#
# In this section, we will demonstrate how to visualize
#
# * time series forecasting
# * predicted components
#
# by using the plotting utilities that come with the Orbit package.
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import orbit
from orbit.models.dlt import DLTMAP, DLTFull
from orbit.diagnostics.plot import plot_predicted_data,plot_predicted_components
from orbit.utils.dataset import load_iclaims
from orbit.utils.plot import get_orbit_style
plt.style.use(get_orbit_style())
import warnings
warnings.filterwarnings('ignore')
# -
print(orbit.__version__)
# +
# load log-transformed data
df = load_iclaims()
train_df = df[df['week'] < '2017-01-01']
test_df = df[df['week'] >= '2017-01-01']
response_col = 'claims'
date_col = 'week'
regressor_col = ['trend.unemploy', 'trend.filling', 'trend.job']
# -
# ## Fit a model
# Here we use the `DLTFull` model as example.
# +
dlt = DLTFull(
response_col=response_col,
regressor_col=regressor_col,
date_col=date_col,
seasonality=52,
prediction_percentiles=[5, 95],
)
dlt.fit(train_df)
# -
# ## Plot Predictions
# First, we do the prediction on the training data before the year 2017.
# +
predicted_df = dlt.predict(df=train_df, decompose=True)
_ = plot_predicted_data(train_df, predicted_df,
date_col=dlt.date_col, actual_col=dlt.response_col)
# -
# Next, we do the predictions on the test data after the year 2017. This plot is useful to help check the overall model performance on the out-of-sample period.
# +
predicted_df = dlt.predict(df=test_df, decompose=True)
_ = plot_predicted_data(training_actual_df=train_df, predicted_df=predicted_df,
date_col=dlt.date_col, actual_col=dlt.response_col,
test_actual_df=test_df)
# -
# ## Plot Predicted Components
# `plot_predicted_components` is the utility to plot each component separately. This is useful when one wants to look into the model prediction results and inspect each component separately.
# +
predicted_df = dlt.predict(df=train_df, decompose=True)
_ = plot_predicted_components(predicted_df, date_col)
# -
# One can use `plot_components` to have more componets to be plotted if they are available in the supplied predicted_df.
_ = plot_predicted_components(predicted_df, date_col,
plot_components=['prediction', 'trend', 'seasonality', 'regression'])
| docs/tutorials/decompose_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ds-unit-4 (Python3)
# language: python
# name: ds-unit-4
# ---
# +
import spacy
# Load the "en_core_web_sm" model
nlp = spacy.load("en_core_web_sm")
text = "It's official: Apple is the first U.S. public company to reach a $1 trillion market value"
# Process the text
doc = nlp(text)
# Print the document text
print(doc.text)
# -
# Predicting linguistic annotations
# +
import spacy
nlp = spacy.load("en_core_web_sm")
text = "It's official: Apple is the first U.S. public company to reach a $1 trillion ,arket value"
# Process the text
doc = nlp(text)
for token in doc:
# Get the token text, part-of-speech tag and dependency label
token_text = token.text
token_pos = token.pos_
token_dep = token.dep_
# This is for formatting only
print(f"{token_text:<12}{token_pos:<10}{token_dep:<10}")
# +
import spacy
nlp = spacy.load("en_core_web_sm")
text = "It's official: Apple is the first U.S. public company to reach a $1 trillion market value."
# Process the text
doc = nlp(text)
# iterate over the predicted entities
for ent in doc.ents:
# Print the entity text and its label
print(ent.text, ent.label_)
# -
# Predicting Named entities in context
# +
import spacy
nlp = spacy.load("en_core_web_sm")
text = "Upcoming iPhone X release date leaked as Apple reveals pre-orders"
# Process the text
doc = nlp(text)
# Iterate over the entities
for ent in doc.ents:
# Print the entitiy text and label
print(ent.text, ent.label_)
# Get the span for "iPhone X"
iphone_x = doc[2:4]
# Print the span text
print("Missing entity:", iphone_x.text)
# +
#### left off 110 RULE-BASED MATCHING
| spaCy_107.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Porting of example from http://czep.net/16/session-ids-sql.html to Spark SQL
spark.sql("create or replace temporary view raw_events as ( \
select * from ( \
select 111 as user_id, timestamp '2016-05-01 17:00:00' as event_time union all \
select 111 as user_id, timestamp '2016-05-01 17:01:00' as event_time union all \
select 111 as user_id, timestamp '2016-05-01 17:02:00' as event_time union all \
select 111 as user_id, timestamp '2016-05-01 17:03:00' as event_time union all \
select 222 as user_id, timestamp '2016-05-01 18:00:00' as event_time union all \
select 333 as user_id, timestamp '2016-05-01 19:00:00' as event_time union all \
select 333 as user_id, timestamp '2016-05-01 19:10:00' as event_time union all \
select 333 as user_id, timestamp '2016-05-01 19:20:00' as event_time union all \
select 333 as user_id, timestamp '2016-05-01 19:30:00' as event_time union all \
select 333 as user_id, timestamp '2016-05-01 20:01:00' as event_time union all \
select 333 as user_id, timestamp '2016-05-01 20:02:00' as event_time union all \
select 444 as user_id, timestamp '2016-05-01 23:01:00' as event_time union all \
select 444 as user_id, timestamp '2016-05-01 23:21:00' as event_time union all \
select 444 as user_id, timestamp '2016-05-01 23:59:00' as event_time union all \
select 444 as user_id, timestamp '2016-05-02 00:01:00' as event_time union all \
select 444 as user_id, timestamp '2016-05-02 00:21:00' as event_time union all \
select 444 as user_id, timestamp '2016-05-02 23:59:00' as event_time union all \
select 444 as user_id, timestamp '2016-05-03 00:05:00' as event_time \
)\
)")
spark.sql("select * from raw_events").show()
spark.sql("create or replace temporary view lagged_events as ( \
select \
user_id, \
event_time, \
lag(event_time) over (partition by date(event_time), user_id order by event_time) as prev \
from \
raw_events \
)")
spark.sql("select * from lagged_events").show()
# changes:
# we use to_unix_timestamp to compare timestamps in seconds,
# 1800 represents 30 minutes which is used to group
# log events into sessions
spark.sql("create or replace temporary view new_sessions as ( \
select \
user_id, \
event_time, \
case \
when prev is null then 1 \
when to_unix_timestamp(event_time) - to_unix_timestamp(prev) > 1800 then 1 \
else 0 \
end as is_new_session \
from \
lagged_events \
)")
spark.sql("select * from new_sessions").show()
spark.sql("create or replace temporary view session_index as ( \
select \
user_id, \
event_time, \
is_new_session, \
sum(is_new_session) over (partition by user_id order by event_time rows between unbounded preceding and current row) as session_index \
from \
new_sessions \
)")
spark.sql("select * from session_index").show()
# changes:
# concat in place of || operator
# string in place of varchar
spark.sql("\
select \
concat(concat(cast(user_id as string), '.'), cast(session_index as string)) as user_id_session_index, \
user_id, \
event_time, \
is_new_session, \
session_index \
from \
session_index \
order by \
user_id, event_time \
").show()
| src/porting-czep.net-blog-post-to-spark-sql.ipynb |
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ### 練習問題2.15
;; Lemが正しいことを⽰せ。いろいろな数値演算の式についてこのシステムの挙動を調査せよ。
;; 何らかの区間AとB を作成し、それらを使って式A/AとA/Bを計算せよ。
;; 中央値に対して⼩さなパーセンテージの幅を持つ区間について調べると得られるところが⼤きいだろう。
;; 中央値-パーセント形式(練習問題 2.12参照)の計算結果を調べよ。
;; +
(define (make-interval a b) (cons a b))
(define (lower-bound x)(car x))
(define (upper-bound x)(cdr x))
(define (make-center-width c w)
(make-interval (- c w) (+ c w)))
(define (center i)
(/ (+ (lower-bound i) (upper-bound i)) 2))
(define (width i)
(/ (- (upper-bound i) (lower-bound i)) 2))
(define (make-center-percent c p)
(let ((w (* c (/ p 100.0))))
(make-interval (- c w) (+ c w)))
)
(define (percent i)
(* (/ (width i) (center i)) 100.0)
)
; 区間の和
(define (add-interval x y)
(make-interval (+ (lower-bound x) (lower-bound y))
(+ (upper-bound x) (upper-bound y)))
)
; 区間の差
(define (sub-interval x y)
(make-interval (- (lower-bound x) (lower-bound y))
(- (upper-bound x) (upper-bound y)))
)
; 区間の積
(define (mul-interval x y)
(let ((p1 (* (lower-bound x) (lower-bound y)))
(p2 (* (lower-bound x) (upper-bound y)))
(p3 (* (upper-bound x) (lower-bound y)))
(p4 (* (upper-bound x) (upper-bound y))))
(make-interval (min p1 p2 p3 p4)
(max p1 p2 p3 p4)))
)
; 区間の割り算
(define (div-interval x y)
(mul-interval
x
(make-interval (/ 1.0 (upper-bound y))
(/ 1.0 (lower-bound y))))
)
(define A (make-center-percent 32.0 2.0))
(define B (make-center-percent 16.0 1.0))
(define (print-i a)
(begin
(display "-----------------------")
(newline)
(display a)
(newline)
(display (center a))
(newline)
(display (width a))
(newline)
(display (percent a))
(newline)
)
)
(print-i A)
(print-i B)
(print-i (div-interval A A))
(print-i (div-interval A B))
(print-i (div-interval B B))
| exercises/2.14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbpresent={"id": "b6da1b81-9914-413e-9c40-e28c68081373"}
# # Monetary Policy Model
#
# **<NAME>, PhD**
#
# This demo is based on the original Matlab demo accompanying the <a href="https://mitpress.mit.edu/books/applied-computational-economics-and-finance">Computational Economics and Finance</a> 2001 textbook by <NAME> and <NAME>.
#
# Original (Matlab) CompEcon file: **demdp11.m**
#
# Running this file requires the Python version of CompEcon. This can be installed with pip by running
#
# # !pip install compecon --upgrade
#
# <i>Last updated: 2021-Oct-01</i>
# <hr>
# -
# ## About
#
# A central bank must set nominal interest rate so as to minimize deviations of inflation rate and GDP gap from established targets.
#
# A monetary authority wishes to control the nominal interest rate $x$ in order to minimize the variation of the inflation rate $s_1$ and the gross domestic product (GDP) gap $s_2$ around specified targets $s^∗_1$ and $s^∗_2$, respectively. Specifically, the authority wishes to minimize expected discounted stream of weighted squared deviations
#
# \begin{equation}
# L(s) = \frac{1}{2}(s − s^∗)'\Omega(s − s^∗)
# \end{equation}
#
# where $s$ is a $2\times 1$ vector containing the inflation rate and the GDP gap, $s^∗$ is a $2\times 1$ vector of targets, and $\Omega$ is a $2 \times 2$ constant positive definite matrix of preference weights. The inflation rate and the GDP gap are a joint controlled exogenous linear Markov process
#
# \begin{equation}
# s_{t+1} = \alpha + \beta s_t + \gamma x_t + \epsilon_{t+1}
# \end{equation}
#
# where $\alpha$ and $\gamma$ are $2 \times 1$ constant vectors, $\beta$ is a $2 \times 2$ constant matrix, and $\epsilon$ is a $2 \times 1$ random vector with mean zero. For institutional reasons, the nominal interest rate $x$ cannot be negative. **What monetary policy minimizes the sum of current and expected future losses?**
#
# This is an infinite horizon, stochastic model with time $t$ measured in years. The state vector $s \in \mathbb{R}^2$
# contains the inflation rate and the GDP gap. The action variable $x \in [0,\infty)$ is the nominal interest rate. The state transition function is $g(s, x, \epsilon) = \alpha + \beta s + \gamma x + \epsilon$
#
# In order to formulate this problem as a maximization problem, one posits a reward function that equals the negative of the loss function $f(s,x) = −L(s)$
#
# The sum of current and expected future rewards satisfies the Bellman equation
#
# \begin{equation}
# V(s) = \max_{0\leq x}\left\{-L(s) + \delta + E_\epsilon V\left(g(s,x,\epsilon)\right)\right\}
# \end{equation}
#
#
# Given the structure of the model, one cannot preclude the possibility that the nonnegativity constraint on the optimal nominal interest rate will be binding in certain states. Accordingly, the shadow-price function $\lambda(s)$ is characterized by the Euler conditions
#
# \begin{align}
# \delta\gamma'E_\epsilon \lambda\left(g(s,x,\epsilon)\right) &= \mu \\
# \lambda(s) &= -\Omega(s-s^*) + \delta\beta'E_\epsilon \lambda\left(g(s,x,\epsilon)\right)
# \end{align}
#
# where the nominal interest rate $x$ and the long-run marginal reward $\mu$ from increasing the nominal interest rate must satisfy the complementarity condition
# \begin{equation}
# x \geq 0, \qquad \mu \leq 0, \qquad x > 0 \Rightarrow \mu = 0
# \end{equation}
#
# It follows that along the optimal path
#
# \begin{align}
# \delta\gamma'E_\epsilon \lambda_{t+1} &= \mu_t \\
# \lambda_t &= -\Omega(s_t-s^*) + \delta\beta'E_\epsilon \lambda_{t+1}\\
# x \geq 0, \qquad \mu \leq 0, &\qquad x > 0 \Rightarrow \mu = 0
# \end{align}
#
# Thus, in any period, the nominal interest rate is reduced until either the long-run marginal reward or the nominal interest rate is driven to zero.
# + nbpresent={"id": "563ef60d-8931-476f-a6be-3e11d89cf68c"}
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from compecon import BasisChebyshev, DPmodel, BasisSpline, qnwnorm, demo
import pandas as pd
pd.set_option('display.float_format',lambda x: f'{x:.3f}')
# + [markdown] nbpresent={"id": "3422905c-443e-4ef5-9e5a-8dfef670e001"}
# ### Model Parameters
#
# + nbpresent={"id": "3292d111-905e-4878-b9e1-6aeef33328f2"}
α = np.array([[0.9, -0.1]]).T # transition function constant coefficients
β = np.array([[-0.5, 0.2], [0.3, -0.4]]) # transition function state coefficients
γ = np.array([[-0.1, 0.0]]).T # transition function action coefficients
Ω = np.identity(2) # central banker's preference weights
ξ = np.array([[1, 0]]).T # equilibrium targets
μ = np.zeros(2) # shock mean
σ = 0.08 * np.identity(2), # shock covariance matrix
δ = 0.9 # discount factor
# + [markdown] nbpresent={"id": "143615f9-8a38-4903-99d4-42f32c36b323"}
# ### State Space
# -
# There are two state variables: 'GDP gap' = $s_0\in[-2,2]$ and 'inflation'=$s_1\in[-3,3]$.
# + nbpresent={"id": "2eddfa9e-b88e-4103-a147-6ded864dfef2"}
n = 21
smin = [-2, -3]
smax = [ 2, 3]
basis = BasisChebyshev(n, smin, smax, method='complete',
labels=['GDP gap', 'inflation'])
# + [markdown] nbpresent={"id": "69027832-bca5-49f6-baf6-cb5a946eb9f9"}
# ### Action space
#
# There is only one action variable x: the nominal interest rate, which must be nonnegative.
# + nbpresent={"id": "9bd30725-21ec-40d4-9e3f-20555b1b3061"}
def bounds(s, i, j):
lb = np.zeros_like(s[0])
ub = np.full(lb.shape, np.inf)
return lb, ub
# + [markdown] nbpresent={"id": "b920851a-06ae-427a-8b78-9b44731c6145"}
# ### Reward Function
# + nbpresent={"id": "cfc887cb-9abb-4e8d-8f84-3caed2359bb2"}
def reward(s, x, i, j):
s = s - ξ
f = np.zeros_like(s[0])
for ii in range(2):
for jj in range(2):
f -= 0.5 * Ω[ii, jj] * s[ii] * s[jj]
fx = np.zeros_like(x)
fxx = np.zeros_like(x)
return f, fx, fxx
# + [markdown] nbpresent={"id": "cf4e628f-067a-4a1d-8881-5aaa191791cb"}
# ### State Transition Function
# + nbpresent={"id": "70c17a3c-56cf-40b5-8ae2-79e7fe6378c9"}
def transition(s, x, i, j, in_, e):
g = α + β @ s + γ @ x + e
gx = np.tile(γ, (1, x.size))
gxx = np.zeros_like(s)
return g, gx, gxx
# + [markdown] nbpresent={"id": "1d497e8a-4a8a-4094-b185-bf63e372aded"}
# The continuous shock must be discretized. Here we use Gauss-Legendre quadrature to obtain nodes and weights defining a discrete distribution that matches the first 6 moments of the Normal distribution (this is achieved with m=3 nodes and weights) for each of the state variables.
# + nbpresent={"id": "45a6bcad-9b3a-4106-93be-48132174588c"}
m = [3, 3]
[e,w] = qnwnorm(m,μ,σ)
# + [markdown] nbpresent={"id": "d721d6a3-1b64-46da-bfbe-e856c80186a0"}
# ### Model structure
# + nbpresent={"id": "26a4fcd3-66c9-494a-bc9a-fbba7f8c9dff"}
bank = DPmodel(basis, reward, transition, bounds,
x=['interest'], discount=δ, e=e, w=w)
# + [markdown] nbpresent={"id": "10890d3a-8e91-4166-9d00-f33822cc6994"}
# Compute Unconstrained Deterministic Steady-State
# +
bank_lq = bank.lqapprox(ξ,0)
sstar = bank_lq.steady['s']
xstar = bank_lq.steady['x']
# + [markdown] nbpresent={"id": "1b4facc1-d91f-4f52-8bbd-343d86c90e56"}
# If Nonnegativity Constraint Violated, Re-Compute Deterministic Steady-State
# + nbpresent={"id": "e72fdf32-dea3-4f55-9da9-ec130ff3c9dd"}
if xstar < 0:
I = np.identity(2)
xstar = 0.0
sstar = np.linalg.solve(np.identity(2) - β, α)
frmt = '\t%-21s = %5.2f'
print('Deterministic Steady-State')
print(frmt % ('GDP Gap', sstar[0]))
print(frmt % ('Inflation Rate', sstar[1]))
print(frmt % ('Nominal Interest Rate', xstar))
# + [markdown] nbpresent={"id": "e826616c-0049-42d4-bf90-6fc4e13c2b3f"}
# ### Solve the model
# We solve the model by calling the `solve` method in `bank`. On return, `sol` is a pandas dataframe with columns *GDP gap*, *inflation*, *value*, *interest*, and *resid*. We set a refined grid `nr=5` for this output.
# + nbpresent={"id": "9c138bd8-6d99-4cf9-87c6-96656670d3b1"}
S = bank.solve(nr=5)
# + [markdown] nbpresent={"id": "0fb51d07-9d77-4617-9da5-9cabfc7c85ee"}
# To make the 3D plots, we need to reshape the columns of `sol`.
# -
S3d = {x: S[x].values.reshape((5*n,5*n)) for x in S.columns}
# This function will make all plots
def makeplot(series,zlabel,zticks,title):
fig = plt.figure(figsize=[8,5])
ax = fig.gca(projection='3d')
ax.plot_surface(S3d['GDP gap'], S3d['inflation'], S3d[series], cmap=cm.coolwarm)
ax.set_xlabel('GDP gap')
ax.set_ylabel('Inflation')
ax.set_zlabel(zlabel)
ax.set_xticks(np.arange(-2,3))
ax.set_yticks(np.arange(-3,4))
ax.set_zticks(zticks)
ax.set_title(title)
# + [markdown] nbpresent={"id": "9d1a7459-9c0d-4397-bb41-c0c3cca75577"}
# ### Optimal policy
# + nbpresent={"id": "ff9ffca8-cfc8-44b9-95d6-bb1b415d4e35"}
fig1 = makeplot('interest', 'Nomianal Interest Rate',
np.arange(0,21,5),'Optimal Monetary Policy')
# + [markdown] nbpresent={"id": "425731c6-8616-4864-a3b8-0a134f04bed9"}
# ### Value function
# + nbpresent={"id": "dbc7d75d-4237-4301-86a1-457c8e7e4e13"}
fig2 = makeplot('value','Value',
np.arange(-12,S['value'].max(),4),'Value Function')
# + [markdown] nbpresent={"id": "0b07bbb4-e7e0-4177-8168-d7314c3804d9"}
# ### Residuals
# + nbpresent={"id": "04985aa7-71f9-4686-bbaa-7072fb9fefe4"}
fig3 = makeplot('resid','Residual',
[-1.5e-3, 0, 1.5e3],'Bellman Equation Residual')
plt.ticklabel_format(style='sci', axis='z', scilimits=(-1,1))
# -
# ## Simulating the model
#
# We simulate 21 periods of the model starting from $s=s_{\min}$, 10000 repetitions.
T = 21
nrep = 10_000
data = bank.simulate(T, np.tile(np.atleast_2d(smax).T,nrep))
subdata = data[data['time']==T][['GDP gap', 'inflation', 'interest']]
stats =pd.DataFrame({'Deterministic Steady-State': [*sstar.flatten(), xstar],
'Ergodic Means': subdata.mean(),
'Ergodic Standard Deviations': subdata.std()})
stats.T
# ### Simulated State and Policy Paths
# +
subdata = data[data['_rep'].isin(range(3))]
opts = dict(spec='r*', offset=(0, -15), fs=11, ha='right')
gdpstar, infstar, intstar = stats['Ergodic Means']
def simplot(series,ylabel,yticks,steady):
fig = demo.figure('Simulated and Expected ' + ylabel,'Period', ylabel,[0, T + 0.5])
plt.plot(data[['time',series]].groupby('time').mean())
plt.plot(subdata.pivot('time','_rep',series),lw=1)
plt.xticks(np.arange(0,24,4))
plt.yticks(yticks)
demo.annotate(T, steady, f'Expected {series}\n = {steady:.2f}', **opts)
return fig
# -
fig4 = simplot('GDP gap','GDP gap',np.arange(smin[0],smax[0]+1),gdpstar)
fig5 = simplot('inflation', 'Inflation Rate',np.arange(smin[1],smax[1]+1),infstar)
fig6 = simplot('interest','Nominal Interest Rate',np.arange(-2,5),intstar)
# +
#demo.savefig([fig1,fig2,fig3,fig4,fig5,fig6])
| _build/jupyter_execute/notebooks/dp/11 Monetary Policy Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from utilities_namespace import *
# ## CMap v2.0
# +
from config import DATA_DIR
from pathlib import Path
data_path = Path(DATA_DIR)
# -
metadata = pd.read_excel(data_path / 'cmap/cmap_instances_02.xls')
footnotes = metadata[metadata.isna().sum(axis=1) > 1].dropna(how='all').dropna(how='all', axis=1)
footnotes.head()
substances = metadata[metadata.isna().sum(axis=1) <= 1].dropna(how='all').set_index('instance_id')
substances.head()
substances
Series(substances.cmap_name.unique()).describe()
substances.cell2.value_counts()
substances['duration (h)'].value_counts()
| notebooks/Connectivity_Map_2.0/CMap_2.0_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
URL = "https://www.geeksforgeeks.org/data-structures/"
r = requests.get(URL)
print(r.content)
# +
import requests
file_url = "http://codex.cs.yale.edu/avi/db-book/db4/slide-dir/ch1-2.pdf"
r = requests.get(file_url, stream = True)
with open("python.pdf","wb") as pdf:
for chunk in r.iter_content(chunk_size=1024):
# writing one chunk at a time to pdf file
if chunk:
pdf.write(chunk)
# +
## Finding title of website using web scraping ##
import requests
import bs4
res = requests.get('https://learncodeonline.in')
type(res)
res.text
soup = bs4.BeautifulSoup(res.text,'lxml')
type(soup)
hi = soup.select('title')
hi[0].getText()
# +
## Web Scraping of Machine Learning wikipedia
import bs4
import requests
res = requests.get('http://www.mospi.gov.in/sites/default/files/publication_reports')
soup = bs4.BeautifulSoup(res.text,'lxml')
#soup.select('.mw-headline')
# -
import bs4
import requests
res = requests.get('http://www.mospi.gov.in/download-reports')
soup = bs4.BeautifulSoup(res.text,'lxml')
for link in soup.find_all('a',href=True):
if re.findall('.pdf$',link['href']):
print(link['href'])
# +
import bs4
import requests
import regex as re
for i in range(1,134):
print("Page : ", i)
url = "http://www.mospi.gov.in/download-reports?combine=&main_cat=All&publication_report_cat=All&sub_category=All&page="+str(i)
res = requests.get(url)
soup = bs4.BeautifulSoup(res.text,'lxml')
for link in soup.find_all('a',href=True):
if (re.findall('.PDF$',link['href'])) or (re.findall('.pdf$',link['href'])):
print(link['href'])
# -
import bs4
import requests
res = requests.get('http://www.mospi.gov.in/download-reports?combine=&main_cat=All&publication_report_cat=All&sub_category=All&page=6')
soup = bs4.BeautifulSoup(res.text,'lxml')
for link in soup.find_all('a',href=True):
if re.findall('.PDF$',link['href']):
print(link['href'])
| Untitled9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
raw_cat = pd.Categorical(["red", "red", "yellow", "green", "blue"],
categories=["red", "yellow", "green"], ordered=False)
s = pd.Series(raw_cat)
print(s)
# +
import numpy as np
df_size = 100_000
df1 = pd.DataFrame(
{
"float_1": np.random.rand(df_size),
"species": np.random.choice(["cat", "dog", "ape", "gorilla"], size=df_size),
}
)
df1_cat = df1.astype({"species": "category"})
# -
df1_cat.info()
display(df1_cat)
df1.memory_usage(deep=True)
df1_cat.memory_usage(deep=True)
| ecommerce/other/pandas-categorical-series-3.2.2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab
# %load_ext line_profiler
from numba import jit, njit
# +
class NTree:
"""Arbitrary dimensionality n-ant tree structure that stores arbitrary data associated with each point."""
def __init__(self, center, size, dim=3):
self.COM = None
self.center = center
self.size = size
self.data = None
self.IsLeaf = False
self.children = (1 << dim) * [None,]
self.dim = dim
def InsertPoint(self, x, data=None):
"""Inserts a point of position x and mass m into the tree."""
if self.COM is None: # no point already lives here, so let's make a leaf node and store the point there
self.COM = x
if data: self.data = data
self.IsLeaf = True
return
#otherwise we gotta split this up
if self.IsLeaf:
self.SpawnChildWithPoint(self.COM, data)
self.IsLeaf = False
self.SpawnChildWithPoint(x, data)
def SpawnChildWithPoint(self, x, data=None):
"""Spawns a child node for a point at position x and mass m to live in."""
signs = (x > self.center)
#sector, signs = Sector(x, self.center)
sector = SignsToSector(signs) # number from 0 to 2**dim - 1 deciding which n-ant
#print(sector is sector2)
if not self.children[sector]:
child_size = self.size/2
child_center = self.center + child_size*(signs-0.5)
self.children[sector] = NTree(child_center, child_size, dim=self.dim)
self.children[sector].InsertPoint(x, data)
def GetMoments(self):
"""Computes the mass and center of mass of a node recursively."""
if not self.IsLeaf: #: return self.mass, self.COM
self.data = 0.
self.COM = np.zeros(self.dim)
for c in self.children:
if c is None: continue
mc, xc = c.GetMoments()
self.data += mc
self.COM += mc*xc
self.COM /= self.data
return self.data, self.COM
@njit
def SignsToSector(signs):
"""Takes a boolean array and returns the integer given by those binary digits."""
sum = 0
for i in range(signs.shape[0]):
#sum += signs[i] * (1 << i)
if signs[i]: sum += 1 << i
return sum
@njit
def Sector(x, center):
"""Returns a number from 0 to 2**dim - 1 labeling which n-ant the point lives in"""
sum = 0
#signs = np.zeros(3, dtype=np.bool)
for i in range(center.shape[0]):
if x[i] > center[i]:
sum += 1 << i
#signs[i] = True
return sum
def ForceWalk(x, g, node, thetamax=0.7, eps=0.0):
dx = node.COM - x
#print(dx)
r = np.sqrt((dx**2).sum())
if r>0:
if node.IsLeaf or node.size/r < thetamax:
g += node.mass * dx / (r**2 + eps**2)**1.5
else:
for c in node.children:
if c: ForceWalk(x, g, c, thetamax, eps)
def Accel(points, tree, thetamax=0.7, G=1.0, eps=0.0):
accels = np.zeros_like(points)
for i in range(points.shape[0]):
ForceWalk(points[i], accels[i], tree, thetamax,eps)
return G*accels
@njit
def BruteForceAccel(x,m,eps=0., G=1.):
accel = zeros_like(x)
for i in range(x.shape[0]):
for j in range(i+1,x.shape[0]):
dx = x[j,0]-x[i,0]
dy = x[j,1]-x[i,1]
dz = x[j,2]-x[i,2]
r = sqrt(dx*dx + dy*dy + dz*dz + eps*eps)
mr3inv = m[i]/(r*r*r)
accel[j,0] -= mr3inv*dx
accel[j,1] -= mr3inv*dy
accel[j,2] -= mr3inv*dz
mr3inv = m[j]/(r*r*r)
accel[i,0] += mr3inv*dx
accel[i,1] += mr3inv*dy
accel[i,2] += mr3inv*dz
return G*accel
@jit
def BruteForcePotential(x,m,G=1., eps=0.):
potential = np.zeros_like(m)
for i in range(x.shape[0]):
for j in range(i+1,x.shape[0]):
dx = x[i,0]-x[j,0]
dy = x[i,1]-x[j,1]
dz = x[i,2]-x[j,2]
r = np.sqrt(dx*dx + dy*dy + dz*dz + eps*eps)
rinv = 1/r
potential[j] -= m[i]*rinv
potential[i] -= m[j]*rinv
return G*potential
def ConstructTree(points, data=None):
mins = np.min(points,axis=0)
maxes = np.max(points,axis=0)
center = (maxes+mins)/2
size = np.max(maxes-mins)
root = NTree(center, size, dim=points.shape[1])
if data:
for i in range(len(points)):
root.InsertPoint(points[i], data[i])#, masses[i])
else:
for i in range(len(points)):
root.InsertPoint(points[i])
#root.GetMoments()
return root
# -
x = 2*(np.random.rand(10**5,3) - 0.5)
#x = x[np.sum(x**2,axis=1)<1.]
#x[:,2] /= 10
masses = np.repeat(1/x.shape[0],x.shape[0])
#v = np.cross(x, np.array([0,0,1])) * 3
#v += np.random.normal(size=x.shape)*0.1
#v *= 0.
#plt.scatter(x[:,0], x[:,1]); plt.show()
# %lprun -f TreeNode.InsertPoint ConstructTree(x, masses)
# %time ConstructTree(x)
# %timeit ConstructTree(x, masses)
#g = np.zeros(3)
# %time root = ConstructTree(x, masses)
#a = Accel(x, root, thetamax=0.7,eps=0.1)
#a[0], BruteForceAccel(x,masses,eps)[0]
#plt.hist(a[:,1],100); plt.show()
#root.children#[2].center
#x[np.sum(a**2,axis=1).argmax()]
#BruteForceAccel(points, masses)
dt = 0.001
eps = 0.1
t = 0.
tmax = 1.
i = 0
#plt.ion()
#ion()
#fig = plt.figure()
#ax = fig.add_subplot(111)
#plt.axes().set_aspect('equal');
#plt.xlim(-1,1)
#plt.ylim(-1,1)
KE = []
PE = []
med = []
while t < tmax:
if not i%100: print(t)# ax.clear(); ax.scatter(x[:,0],x[:,1],s=0.3); plt.xlim(-1,1); plt.ylim(-1,1); plt.draw(); plt.pause(0.01)
#plt.savefig("%d.png"%i); plt.plt.clf()
x += v*dt #, v + BruteForceAccel(x, masses, eps=eps)*dt
#root = ConstructTree(x, masses)
v += BruteForceAccel(x, masses, eps=eps)*dt
i += 1
t += dt
KE.append((v**2).sum())
PE.append(BruteForcePotential(x,masses,1.,eps).sum())
med.append(np.percentile(np.sum(x**2,axis=1)**0.5, 50))
plt.plot(np.array(KE) + np.array(PE))
#plt.plot(PE)
plt.show()
plt.plot(med); plt.show()
np.packbits(np.array([True,True,True]))
signs = np.random.rand(3) > 0.5
# #%timeit sum(signs[i] * (1 << i) for i in range(3))
# %timeit SignsToIndex(signs)
# %timeit center + 0.5*(signs-0.5)
from numba import uint8, boolean
# +
@jit
def SignsToIndex(signs):
sum = 0
for i in range(signs.shape[0]):
sum += signs[i] * (1 << i)
return sum
@jit
# +
x = np.random.rand(3)*2 - 1
center = np.array([0.1,0.1,0.1])
# -
signs = (x > center)
# %timeit SignsToSector(signs) # number from 0 to 2**dim deciding which n-ant
| Quadtree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# code by <NAME> @graykode
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import matplotlib.pyplot as plt
def random_batch():
random_inputs = []
random_labels = []
random_index = np.random.choice(range(len(skip_grams)), batch_size, replace=False)
for i in random_index:
random_inputs.append(np.eye(voc_size)[skip_grams[i][0]]) # target
random_labels.append(skip_grams[i][1]) # context word
return random_inputs, random_labels
# Model
class Word2Vec(nn.Module):
def __init__(self):
super(Word2Vec, self).__init__()
# W and WT is not Traspose relationship
self.W = nn.Linear(voc_size, embedding_size, bias=False) # voc_size > embedding_size Weight
self.WT = nn.Linear(embedding_size, voc_size, bias=False) # embedding_size > voc_size Weight
def forward(self, X):
# X : [batch_size, voc_size]
hidden_layer = self.W(X) # hidden_layer : [batch_size, embedding_size]
output_layer = self.WT(hidden_layer) # output_layer : [batch_size, voc_size]
return output_layer
if __name__ == '__main__':
batch_size = 2 # mini-batch size
embedding_size = 2 # embedding size
sentences = ["apple banana fruit", "banana orange fruit", "orange banana fruit",
"dog cat animal", "cat monkey animal", "monkey dog animal"]
word_sequence = " ".join(sentences).split()
word_list = " ".join(sentences).split()
word_list = list(set(word_list))
word_dict = {w: i for i, w in enumerate(word_list)}
voc_size = len(word_list)
# Make skip gram of one size window
skip_grams = []
for i in range(1, len(word_sequence) - 1):
target = word_dict[word_sequence[i]]
context = [word_dict[word_sequence[i - 1]], word_dict[word_sequence[i + 1]]]
for w in context:
skip_grams.append([target, w])
model = Word2Vec()
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
# Training
for epoch in range(5000):
input_batch, target_batch = random_batch()
input_batch = torch.Tensor(input_batch)
target_batch = torch.LongTensor(target_batch)
optimizer.zero_grad()
output = model(input_batch)
# output : [batch_size, voc_size], target_batch : [batch_size] (LongTensor, not one-hot)
loss = criterion(output, target_batch)
if (epoch + 1) % 1000 == 0:
print('Epoch:', '%04d' % (epoch + 1), 'cost =', '{:.6f}'.format(loss))
loss.backward()
optimizer.step()
for i, label in enumerate(word_list):
W, WT = model.parameters()
x, y = W[0][i].item(), W[1][i].item()
plt.scatter(x, y)
plt.annotate(label, xy=(x, y), xytext=(5, 2), textcoords='offset points', ha='right', va='bottom')
plt.show()
| 1-2.Word2Vec/Word2Vec-Skipgram(Softmax).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
raw_data_path = os.path.join(os.path.pardir, 'data', 'raw')
train_file_path = os.path.join(raw_data_path, 'train.csv')
test_file_path = os.path.join(raw_data_path, 'test.csv')
train_df = pd.read_csv(train_file_path, index_col='PassengerId')
test_df = pd.read_csv(test_file_path, index_col='PassengerId')
type(train_df)
train_df.info()
test_df.info()
test_df['Survived'] = -888
test_df.info()
df = pd.concat((train_df, test_df), sort=True)
df.info()
df.head()
df.tail(10)
df.Name.head()
df[["Name", "Age"]][5:10]
# # Filtering with Pandas
df.loc[5:10,['Fare','Sex']]
df.loc[5:10,'Fare':'Sex']
df.iloc[5:10,1:5]
df.loc[df.Sex == 'male']
# # Numerical statistics
df.describe()
print('mean fare: {0}'.format(df.Fare.mean()))
print('fare standard deviation: {0}'.format(df.Fare.std()))
print('fare variance: {0}'.format(df.Fare.var()))
print('fare range: {0}'.format(df.Fare.max() - df.Fare.min()))
print('mean age: {0}'.format(df.Age.mean()))
print('median age: {0}'.format(df.Age.median()))
print('age standard deviation: {0}'.format(df.Age.std()))
print('age range: {0}'.format(df.Age.max() - df.Age.min()))
print('50 percentile: {0}'.format(df.Age.quantile(.50)))
print('50 percentile: {0}'.format(df.Age.quantile(.75)))
# +
# %matplotlib inline
#box-whisker plot
df.Fare.plot(kind='box')
# -
# # Categorical features for non-numerical data
df.describe(include='all')
df.Sex.value_counts()
df.Sex.value_counts(normalize=True)
df.groupby('Pclass').Sex.value_counts()
df[df.Survived != -888].Survived.value_counts(normalize=True)
df.Pclass.value_counts()
df.Pclass.value_counts().plot(kind='bar', rot=0, title='Class wise passenger count', color='b')
# # Distributions
df.Age.plot(kind='hist', title='histogram of passangers age', color='c')
df.Age.plot(kind='hist', title='histogram of passangers age', color='c', bins=20)
df.Age.plot(kind='kde', title='Density plot for passangers age', color='b')
#df.Age.plot.kde()
df.Fare.plot(kind='hist', title='historgram of fares', bins=20)
print('skewness for age: {0:.2f}'.format(df.Age.skew()))
print('skewness for fare: {0:.2f}'.format(df.Fare.skew()))
df.plot.scatter(x='Age', y='Fare', title='scatter plot: age vs fare', alpha=0.1)
df.plot.scatter(y='Fare', x='Pclass', title='scatter plot: Fare vs class')
# # Group by and aggregations
df.groupby('Sex').Age.median()
df.groupby('Pclass')['Fare', 'Age'].median()
df.groupby('Pclass').agg({'Fare': 'mean', 'Age': 'median'})
aggregation = {
'Fare': {
'mean_fare': 'mean',
'median_fare': 'median',
'max_fare': max,
'min_fare': min
},
'Age': {
'mean_age': 'mean',
'range_age': lambda x: max(x) - min(x)
}
}
df.groupby('Pclass').agg(aggregation)
df.groupby(['Pclass', 'Embarked']).Fare.median()
# # Crosstabs
pd.crosstab(df.Sex, df.Pclass)
pd.crosstab(df.Sex, df.Pclass).plot(kind='bar')
# # Pivot table
df.pivot_table(index='Sex', columns='Pclass', values='Age', aggfunc='mean')
# the same information by doing groupby:
df.groupby(['Sex', 'Pclass']).Age.mean()
df.groupby(['Sex', 'Pclass']).Age.mean().unstack()
# # Data munging - missing values
df.info()
# ## Feature: Embarked
df[df.Embarked.isnull()]
# which embarkment is the most common
df.Embarked.value_counts()
# which embarkement has the most survived passangers
df[df.Survived != -888].groupby('Embarked').Survived.value_counts(normalize=True)
# how many woman survived per embarkement
known_for_survival_df = df[df.Survived != -888]
df_female_survival = known_for_survival_df[known_for_survival_df.Sex == 'female']
df_female_survival.groupby('Embarked').Survived.value_counts(normalize=True)
# median age of woman per embarkement and survival
df_female_survival.pivot_table(index='Embarked', columns='Survived', values='Age', aggfunc='median')
pd.crosstab(df_female_survival.Survived, df_female_survival.Embarked, normalize=True)
df_female_high_fare = df_female_survival[df_female_survival.Fare > 70]
df_female_high_fare_class = df_female_high_fare[df_female_high_fare.Pclass == 1]
df_female_high_fare_class.info()
pd.crosstab(df_female_high_fare_class.Survived, df_female_high_fare_class.Embarked, normalize=True)
df_female_survival.groupby(['Pclass', 'Embarked']).Fare.median()
# impute missing values
#df.loc[df.Embarked.isnull(), 'Embarked'] = 'C'
df.Embarked.fillna('C', inplace=True)
df[df.Embarked.isnull()]
df.info()
# ## Feature: Fare
df[df.Fare.isnull()]
median_fare = df.loc[(df.Embarked == 'S') & (df.Pclass == 3), 'Fare'].median()
print(median_fare)
df.Fare.fillna(median_fare, inplace=True)
df.info()
# ## Feature: Age
pd.options.display.max_rows = 15
df[df.Age.isnull()]
df_age_filled = df[df.Age.isnull() == False]
df_age_filled_man = df_age_filled[df_age_filled.Sex == 'male']
df_age_filled_woman = df_age_filled[df_age_filled.Sex == 'female']
median_woman_age_c1 = df_age_filled_woman[df_age_filled_woman.Pclass == 1].Age.median()
woman_age_per_class = df_age_filled_woman.groupby('Pclass').Age.median()
man_age_per_class = df_age_filled_man.groupby('Pclass').Age.median()
print(woman_age_per_class)
print(man_age_per_class)
# visualize gender vs age
df_age_filled.boxplot('Age', 'Sex')
#quite similar, not worth distinguishing?
# ## Nice and quick way to fill with median age grouped by sex and passanger class
# +
#df.Age.fillna(df.groupby(['Sex', 'Pclass']).Age.transform('median'), inplace=True)
# -
df[df.Age.isnull()]
# ## Option 2: base age imputation on title in the name
def get_title(name):
first_name = name.split(',')[1]
return first_name.split('.')[0].strip().lower()
df_age_filled.Name.map(get_title)
df_age_filled.Name.map(get_title).unique()
df['Title'] = df.Name.map(get_title)
df.boxplot('Age', 'Title')
# NICE !!!
title_age_median = df.groupby('Title').Age.transform('median')
df.Age.fillna(title_age_median, inplace=True)
df.info()
# # Data munging - Handling outliers
df.Age.plot(kind='hist', bins=20, color='c')
df[df.Age > 70]
df.Fare.plot(kind='box')
df[df.Fare == df.Fare.max()]
# ## Outliers: Transform fare with logarithm function
df.Fare.plot(kind='hist', bins=20)
log_fare = np.log(df.Fare + 1)
log_fare.plot(kind='hist', bins=20)
# ## Outliers: binning/discretization
binned_fare = pd.qcut(df.Fare, 4, labels = ['small', 'medium', 'high', 'very high'])
binned_fare.value_counts().plot(kind='bar')
# add new bin fare feature
df['Fare_Bin'] = binned_fare
# # Feature engineering
# ## New feature: AgeState
df['AgeStatus'] = np.where(df.Age >= 18, 'Adult', 'Child')
pd.crosstab(df[df.Survived != -888].Survived, df.AgeStatus, normalize=True)
# ## New feature: FamilySize
df['FamilySize'] = df.Parch + df.SibSp + 1
pd.crosstab(df[df.Survived != -888].Survived, df.FamilySize)
# ## New feature: IsMother
df['IsMother'] = np.where(((df.Sex=='female') & (df.Parch>0) & (df.AgeStatus=='Adult') & (df.Title != 'miss')), 1,0)
pd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].IsMother)
# ## New feature: Deck
df.Cabin.unique()
df[df.Cabin=='T'] = np.nan
# +
#def get_deck(cabin):
# return str(np.where(pd.notnull(cabin), cabin[0].upper(), 'Z'))
def get_deck(cabin):
if pd.isnull(cabin):
return 'Z'
else:
return cabin[0].upper()
# -
df['Deck'] = df.Cabin.map(lambda x: get_deck(x))
df.Deck
pd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].Deck)
# # Categorical feature encoding
df['IsMale'] = df.Sex.map(lambda x: int(x == 'male'))
df.info()
df = pd.get_dummies(df, columns=['Deck', 'Pclass', 'Title', 'Fare_Bin', 'Embarked', 'AgeStatus'])
df.info()
# # Drop and reorder columns
df.drop(['Cabin', 'Name', 'Ticket', 'Parch', 'SibSp', 'Sex'], axis=1, inplace=True)
columns = [column for column in df.columns if column != 'Survived']
columns = ['Survived'] + columns
df = df[columns]
df.info()
# # Save processed dataframe
processed_data_path = os.path.join(os.path.pardir, 'data', 'processed')
train_path = os.path.join(processed_data_path, 'train.csv')
test_path = os.path.join(processed_data_path, 'test.csv')
df.loc[df.Survived != -888].to_csv(train_path)
test_columns = [col for col in df.columns if col != 'Survived']
df.loc[df.Survived == -888][test_columns].to_csv(test_path)
# # Create data processing script
script_path = os.path.join(os.path.pardir, 'src', 'data', 'process_data.py')
# +
# %%writefile $script_path
import numpy as np
import pandas as pd
import os
def read_data():
raw_data_path = os.path.join(os.path.pardir, 'data', 'raw')
train_file_path = os.path.join(raw_data_path, 'train.csv')
test_file_path = os.path.join(raw_data_path, 'test.csv')
train_df = pd.read_csv(train_file_path, index_col='PassengerId')
test_df = pd.read_csv(test_file_path, index_col='PassengerId')
test_df['Survived'] = -888
return pd.concat((train_df, test_df), sort=True)
def process_data(df):
return (df
.assign(Title = lambda df: df.Name.map(__get_title))
.pipe(__fill_missing_values)
.assign(FareBin = lambda df: pd.qcut(df.Fare, 4, labels=['low', 'medium', 'high', 'very_high']))
.assign(AgeStatus = lambda df: np.where(df.Age >= 18, 'adult', 'child'))
.assign(FamilySize = lambda df: df.Parch + df.SibSp + 1)
.assign(IsMother = lambda df: np.where(((df.Sex=='female') & (df.Parch>0) & (df.AgeStatus=='Adult') & (df.Title != 'miss')), 1, 0))
.assign(Cabin = lambda df: np.where(df.Cabin == 'T', np.nan, df.Cabin))
.assign(Deck = lambda df: df.Cabin.map(__get_deck))
.assign(IsMale = lambda df: df.Sex.map(lambda s: np.where(s == 'male', 1, 0)))
.pipe(pd.get_dummies, columns=['Deck', 'Pclass', 'Title', 'FareBin', 'Embarked', 'AgeStatus'])
.drop(['Cabin', 'Name', 'Ticket', 'Parch', 'SibSp', 'Sex'], axis=1)
.pipe(__reorder_columns)
)
def write_data(df):
processed_data_path = os.path.join(os.path.pardir, 'data', 'processed')
train_path = os.path.join(processed_data_path, 'train.csv')
test_path = os.path.join(processed_data_path, 'test.csv')
df.loc[df.Survived != -888].to_csv(train_path)
test_columns = [col for col in df.columns if col != 'Survived']
df.loc[df.Survived == -888][test_columns].to_csv(test_path)
def __get_title(name):
first_name = name.split(',')[1]
return first_name.split('.')[0].strip().lower()
def __fill_missing_values(df):
df.Embarked.fillna('C', inplace=True)
median_fare = df.loc[(df.Embarked == 'S') & (df.Pclass == 3), 'Fare'].median()
df.Fare.fillna(median_fare, inplace=True)
title_age_median = df.groupby('Title').Age.transform('median')
df.Age.fillna(title_age_median, inplace=True)
return df
def __get_deck(cabin):
if pd.isnull(cabin):
return 'Z'
else:
return cabin[0].upper()
def __reorder_columns(df):
columns = [column for column in df.columns if column != 'Survived']
columns = ['Survived'] + columns
return df[columns]
if __name__ == '__main__':
df = read_data()
df_processed = process_data(df)
write_data(df_processed)
# -
| notebooks/Exploring and processing data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
# df = pd.read_csv('DANIELS_data_engineering_dataset.csv')
# df.head()
# -
import sqlite3
conn = sqlite3.connect('cars.db')
c = conn.cursor()
# +
c.execute('''SELECT *
FROM Dim_Product
;''')
df_Dim_Product = pd.DataFrame(c.fetchall())
df_Dim_Product.columns = [x[0] for x in c.description]
df_Dim_Product.head()
# +
c.execute('''SELECT *
FROM Dim_Stores
;''')
df_Dim_Stores = pd.DataFrame(c.fetchall())
df_Dim_Stores.columns = [x[0] for x in c.description]
df_Dim_Stores.head()
# +
c.execute('''SELECT *
FROM Fact_Sales
;''')
df_Fact_Sales = pd.DataFrame(c.fetchall())
df_Fact_Sales.columns = [x[0] for x in c.description]
df_Fact_Sales
# +
c.execute('''SELECT *
FROM Fact_Sales
ORDER BY Store_ID ASC
LIMIT 100
;''')
df_Fact_Sales = pd.DataFrame(c.fetchall())
df_Fact_Sales.columns = [x[0] for x in c.description]
df_Fact_Sales
# -
df_Fact_Sales['Store_ID'][23]
df_Fact_Sales['PurchVal'][0]
type(df_Fact_Sales['PurchVal'][0])
type(df_Fact_Sales['Store_ID'][0])
type(df_Dim_Stores['Store_ID'][80])
df_Dim_Stores['Store_ID'][99]
type(df_Dim_Product['VIN_ID'][0])
df_Dim_Product['VIN_ID'][990]
c.execute('''SELECT *
FROM sqlite_master
WHERE TYPE = 'table'
ORDER BY name;''').fetchall()
| database/testing_area.ipynb |
try:
import openmdao.api as om
import dymos as dm
except ImportError:
# !python -m pip install openmdao[notebooks]
# !python -m pip install dymos[docs]
import openmdao.api as om
import dymos as dm
# (examples:minimium_time_climb)=
# # Supersonic Interceptor Minimum Time Climb
#
# This example is based on the _A/C Min Time to Climb_ example given in
# chapter 4 of Bryson {cite}`bryson1999dynamic`. It finds the
# angle-of-attack history required to accelerate a supersonic interceptor
# from near ground level, Mach 0.4 to an altitude of 20 km and Mach 1.0.
#
# 
#
# The vehicle dynamics are given by
#
# \begin{align}
# \frac{dv}{dt} &= \frac{T}{m} \cos \alpha - \frac{D}{m} - g \sin \gamma \\
# \frac{d\gamma}{dt} &= \frac{T}{m v} \sin \alpha + \frac{L}{m v} - \frac{g \cos \gamma}{v} \\
# \frac{dh}{dt} &= v \sin \gamma \\
# \frac{dr}{dt} &= v \cos \gamma \\
# \frac{dm}{dt} &= - \frac{T}{g I_{sp}}
# \end{align}
#
# The initial conditions are
#
# \begin{align}
# r_0 &= 0 \rm{\,m} \\
# h_0 &= 100 \rm{\,m} \\
# v_0 &= 135.964 \rm{\,m/s} \\
# \gamma_0 &= 0 \rm{\,deg} \\
# m_0 &= 19030.468 \rm{\,kg}
# \end{align}
#
# and the final conditions are
#
# \begin{align}
# h_f &= 20000 \rm{\,m} \\
# M_f &= 1.0 \\
# \gamma_0 &= 0 \rm{\,deg}
# \end{align}
#
# ## The ODE System: min_time_climb_ode.py
#
# The top level ODE definition is a _Group_ that connects several subsystems.
om.display_source("dymos.examples.min_time_climb.doc.aero_partial_coloring")
# +
import openmdao.api as om
from dymos.models.eom import FlightPathEOM2D
from dymos.examples.min_time_climb.prop import PropGroup
from dymos.models.atmosphere import USatm1976Comp
from dymos.examples.min_time_climb.doc.aero_partial_coloring import AeroGroup
class MinTimeClimbODE(om.Group):
def initialize(self):
self.options.declare('num_nodes', types=int)
self.options.declare('fd', types=bool, default=False, desc='If True, use fd for partials')
self.options.declare('partial_coloring', types=bool, default=False,
desc='If True and fd is True, color the approximated partials')
def setup(self):
nn = self.options['num_nodes']
self.add_subsystem(name='atmos',
subsys=USatm1976Comp(num_nodes=nn),
promotes_inputs=['h'])
self.add_subsystem(name='aero',
subsys=AeroGroup(num_nodes=nn,
fd=self.options['fd'],
partial_coloring=self.options['partial_coloring']),
promotes_inputs=['v', 'alpha', 'S'])
self.connect('atmos.sos', 'aero.sos')
self.connect('atmos.rho', 'aero.rho')
self.add_subsystem(name='prop',
subsys=PropGroup(num_nodes=nn),
promotes_inputs=['h', 'Isp', 'throttle'])
self.connect('aero.mach', 'prop.mach')
self.add_subsystem(name='flight_dynamics',
subsys=FlightPathEOM2D(num_nodes=nn),
promotes_inputs=['m', 'v', 'gam', 'alpha'])
self.connect('aero.f_drag', 'flight_dynamics.D')
self.connect('aero.f_lift', 'flight_dynamics.L')
self.connect('prop.thrust', 'flight_dynamics.T')
# -
# ## Building and running the problem
#
# In the following code we follow the following process to solve the
# problem:
# +
import matplotlib.pyplot as plt
import openmdao.api as om
import dymos as dm
from dymos.examples.plotting import plot_results
#
# Instantiate the problem and configure the optimization driver
#
p = om.Problem(model=om.Group())
p.driver = om.pyOptSparseDriver()
p.driver.options['optimizer'] = 'SLSQP'
p.driver.declare_coloring()
#
# Instantiate the trajectory and phase
#
traj = dm.Trajectory()
phase = dm.Phase(ode_class=MinTimeClimbODE,
transcription=dm.GaussLobatto(num_segments=15, compressed=False))
traj.add_phase('phase0', phase)
p.model.add_subsystem('traj', traj)
#
# Set the options on the optimization variables
# Note the use of explicit state units here since much of the ODE uses imperial units
# and we prefer to solve this problem using metric units.
#
phase.set_time_options(fix_initial=True, duration_bounds=(50, 400),
duration_ref=100.0)
phase.add_state('r', fix_initial=True, lower=0, upper=1.0E6, units='m',
ref=1.0E3, defect_ref=1.0E3,
rate_source='flight_dynamics.r_dot')
phase.add_state('h', fix_initial=True, lower=0, upper=20000.0, units='m',
ref=1.0E2, defect_ref=1.0E2,
rate_source='flight_dynamics.h_dot')
phase.add_state('v', fix_initial=True, lower=10.0, units='m/s',
ref=1.0E2, defect_ref=1.0E2,
rate_source='flight_dynamics.v_dot')
phase.add_state('gam', fix_initial=True, lower=-1.5, upper=1.5, units='rad',
ref=1.0, defect_ref=1.0,
rate_source='flight_dynamics.gam_dot')
phase.add_state('m', fix_initial=True, lower=10.0, upper=1.0E5, units='kg',
ref=1.0E3, defect_ref=1.0E3,
rate_source='prop.m_dot')
phase.add_control('alpha', units='deg', lower=-8.0, upper=8.0, scaler=1.0,
rate_continuity=True, rate_continuity_scaler=100.0,
rate2_continuity=False)
phase.add_parameter('S', val=49.2386, units='m**2', opt=False, targets=['S'])
phase.add_parameter('Isp', val=1600.0, units='s', opt=False, targets=['Isp'])
phase.add_parameter('throttle', val=1.0, opt=False, targets=['throttle'])
#
# Setup the boundary and path constraints
#
phase.add_boundary_constraint('h', loc='final', equals=20000, scaler=1.0E-3)
phase.add_boundary_constraint('aero.mach', loc='final', equals=1.0)
phase.add_boundary_constraint('gam', loc='final', equals=0.0)
phase.add_path_constraint(name='h', lower=100.0, upper=20000, ref=20000)
phase.add_path_constraint(name='aero.mach', lower=0.1, upper=1.8)
# Minimize time at the end of the phase
phase.add_objective('time', loc='final', ref=1.0)
p.model.linear_solver = om.DirectSolver()
#
# Setup the problem and set the initial guess
#
p.setup(check=True)
p['traj.phase0.t_initial'] = 0.0
p['traj.phase0.t_duration'] = 500
p.set_val('traj.phase0.states:r', phase.interp('r', [0.0, 50000.0]))
p.set_val('traj.phase0.states:h', phase.interp('h', [100.0, 20000.0]))
p.set_val('traj.phase0.states:v', phase.interp('v', [135.964, 283.159]))
p.set_val('traj.phase0.states:gam', phase.interp('gam', [0.0, 0.0]))
p.set_val('traj.phase0.states:m', phase.interp('m', [19030.468, 10000.]))
p.set_val('traj.phase0.controls:alpha', phase.interp('alpha', [0.0, 0.0]))
#
# Solve for the optimal trajectory
#
dm.run_problem(p)
#
# Get the explicitly simulated solution and plot the results
#
exp_out = traj.simulate()
plot_results([('traj.phase0.timeseries.time', 'traj.phase0.timeseries.states:h',
'time (s)', 'altitude (m)'),
('traj.phase0.timeseries.time', 'traj.phase0.timeseries.controls:alpha',
'time (s)', 'alpha (deg)')],
title='Supersonic Minimum Time-to-Climb Solution',
p_sol=p, p_sim=exp_out)
plt.show()
# +
from openmdao.utils.assert_utils import assert_near_equal
assert_near_equal(p.get_val('traj.phase0.t_duration'), 321.0, tolerance=1.0E-1)
# -
# ## References
#
# ```{bibliography}
# :filter: docname in docnames
# ```
| docs/examples/min_time_climb/min_time_climb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/micahks/DS-Unit-2-Linear-Models/blob/master/module2-regression-2/Micah_DS_212_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="zZq9uhZ-UyVm"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 2*
#
# ---
# + [markdown] id="7IXUfiQ2UKj6"
# # Regression 2
#
# ## Assignment
#
# You'll continue to **predict how much it costs to rent an apartment in NYC,** using the dataset from renthop.com.
#
# - [ ] Do train/test split. Use data from April & May 2016 to train. Use data from June 2016 to test.
# - [ ] Engineer at least two new features. (See below for explanation & ideas.)
# - [ ] Fit a linear regression model with at least two features.
# - [ ] Get the model's coefficients and intercept.
# - [ ] Get regression metrics RMSE, MAE, and $R^2$, for both the train and test data.
# - [ ] What's the best test MAE you can get? Share your score and features used with your cohort on Slack!
# - [ ] As always, commit your notebook to your fork of the GitHub repo.
#
#
# #### [Feature Engineering](https://en.wikipedia.org/wiki/Feature_engineering)
#
# > "Some machine learning projects succeed and some fail. What makes the difference? Easily the most important factor is the features used." — <NAME>, ["A Few Useful Things to Know about Machine Learning"](https://homes.cs.washington.edu/~pedrod/papers/cacm12.pdf)
#
# > "Coming up with features is difficult, time-consuming, requires expert knowledge. 'Applied machine learning' is basically feature engineering." — <NAME>, [Machine Learning and AI via Brain simulations](https://forum.stanford.edu/events/2011/2011slides/plenary/2011plenaryNg.pdf)
#
# > Feature engineering is the process of using domain knowledge of the data to create features that make machine learning algorithms work.
#
# #### Feature Ideas
# - Does the apartment have a description?
# - How long is the description?
# - How many total perks does each apartment have?
# - Are cats _or_ dogs allowed?
# - Are cats _and_ dogs allowed?
# - Total number of rooms (beds + baths)
# - Ratio of beds to baths
# - What's the neighborhood, based on address or latitude & longitude?
#
# ## Stretch Goals
# - [ ] If you want more math, skim [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapter 3.1, Simple Linear Regression, & Chapter 3.2, Multiple Linear Regression
# - [ ] If you want more introduction, watch [<NAME>, Statistics 101: Simple Linear Regression](https://www.youtube.com/watch?v=ZkjP5RJLQF4)
# (20 minutes, over 1 million views)
# - [ ] Add your own stretch goal(s) !
# + [markdown] id="58R20nunbyfM"
# #Data Wrangling
# + id="o9eSnDYhUGD7"
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + id="cvrw-T3bZOuW"
import numpy as np
import pandas as pd
# Read New York City apartment rental listing data
df = pd.read_csv(DATA_PATH+'apartments/renthop-nyc.csv')
assert df.shape == (49352, 34)
# Remove the most extreme 1% prices,
# the most extreme .1% latitudes, &
# the most extreme .1% longitudes
df = df[(df['price'] >= np.percentile(df['price'], 0.5)) &
(df['price'] <= np.percentile(df['price'], 99.5)) &
(df['latitude'] >= np.percentile(df['latitude'], 0.05)) &
(df['latitude'] < np.percentile(df['latitude'], 99.95)) &
(df['longitude'] >= np.percentile(df['longitude'], 0.05)) &
(df['longitude'] <= np.percentile(df['longitude'], 99.95))]
# + id="p7fGUrnPWIMZ" outputId="33010984-1125-4781-fc77-e0a02f8951b5" colab={"base_uri": "https://localhost:8080/", "height": 501}
df.head()
# + id="hRNv3ffKYYlL" outputId="26cde851-5dbd-4989-9561-9b817014cdbc" colab={"base_uri": "https://localhost:8080/", "height": 745}
#change the created column to at date_time object and removed the time.
df['created'] = pd.to_datetime(df['created']).dt.date
df['created'] = pd.to_datetime(df['created'])
df.info()
# + id="DcpK6wiIYgG6" outputId="6ee8b17d-6bc5-43e2-8659-269f4bd74bb9" colab={"base_uri": "https://localhost:8080/", "height": 484}
df.head()
# + id="116mVbT5aHF3"
#set index to the created column
df = df.set_index('created').sort_values('created')
# + id="GoHywxd8a_g4" outputId="33131759-37cc-464a-bc00-12a3f7d04727" colab={"base_uri": "https://localhost:8080/", "height": 564}
df.head()
# + [markdown] id="Ei5GimOPnm08"
# ## Create new features
# + id="IWXFXjhPfESY" outputId="349aa559-989c-4292-ff54-8816cbe54c94" colab={"base_uri": "https://localhost:8080/", "height": 798}
#creating new columns for feature engineering
df['amenity_score'] = df[['elevator',
'cats_allowed',
'hardwood_floors',
'dogs_allowed',
'doorman',
'dishwasher',
'no_fee',
'laundry_in_building',
'fitness_center',
# 'pre-war',
'laundry_in_unit',
'roof_deck',
'outdoor_space',
'dining_room',
'high_speed_internet',
'balcony',
'swimming_pool',
'new_construction',
'terrace',
'exclusive',
'loft',
'garden_patio',
'wheelchair_access',
'common_outdoor_space']].sum(axis=1)
df['interest_score'] = df['interest_level'].map({'low': 1, 'medium': 2, 'high':3})
df['description_char'] = df['description'].str.len()
df['description_char'] = df['description_char'].fillna(0)
df['lat/lon'] = df['latitude']*df['longitude']
df.info()
# + [markdown] id="nnaQ-uBupPBK"
# ## Plots
# + id="Tklqe0mygKAI"
bathrooms = 'bathrooms'
bedrooms = 'bedrooms'
description = 'description_char'
lat = 'latitude'
lon = 'longitude'
interest = 'interest_score'
amenities = 'amenity_score'
price = 'price'
# + id="j4Z4j_AWyhUs" outputId="8099ccd3-a292-400d-89d3-5bc46aef5b21" colab={"base_uri": "https://localhost:8080/", "height": 247}
df[description]
# + id="eWGKpH0mjl5R" outputId="6c59f326-f911-4c9f-ce9a-6701e4d2b73e" colab={"base_uri": "https://localhost:8080/", "height": 280}
import matplotlib.pyplot as plt
plt.scatter(df[bathrooms], df[price], alpha=.1)
plt.xlabel('baths')
plt.ylabel('price');
# + id="RsqGe7rvqytq" outputId="4c2a27dc-593f-41df-f3a5-46c9db37c30b" colab={"base_uri": "https://localhost:8080/", "height": 280}
plt.scatter(df[bedrooms], df[price], alpha=.1)
plt.xlabel('beds')
plt.ylabel('price');
# + id="-s18ra_qq3gd" outputId="43e0ebe1-36c8-40b0-a5d0-f33c481e11a2" colab={"base_uri": "https://localhost:8080/", "height": 280}
plt.scatter(df[description], df[price], alpha=.1)
plt.xlabel('description chars')
plt.ylabel('price');
# + id="IPB6GEAXrFen" outputId="d22bfe27-a3ba-433e-a33c-58e4405f7b24" colab={"base_uri": "https://localhost:8080/", "height": 280}
plt.scatter(df[lat], df[price], alpha=.1)
plt.xlabel('lattitude')
plt.ylabel('price');
# + id="3sZX7MmSt4Vb" outputId="ecca46af-3c8e-4e33-a7f6-dfd86269cdd8" colab={"base_uri": "https://localhost:8080/", "height": 280}
plt.scatter(df[lon], df[price], alpha=.1)
plt.xlabel('longitude')
plt.ylabel('price');
# + id="4kfrrII11tb7" outputId="df31593f-5361-4e93-81e4-5881c7561c1f" colab={"base_uri": "https://localhost:8080/", "height": 280}
plt.scatter(df[latlon], df[price], alpha=.1)
plt.xlabel('lat/lon')
plt.ylabel('price');
# + id="61ChOM5crNJH" outputId="747b8010-02f9-461d-e76f-7a2e97b59a93" colab={"base_uri": "https://localhost:8080/", "height": 278}
#for my own curiosity, I wanted to see where the expesive rentals are in relation to eachother.
plt.scatter(df[lat], df[lon], c=df[price])
plt.xlabel('lat')
plt.ylabel('long');
# + id="HmnH1hI1rbg6" outputId="6b853295-64f7-4f08-e6fd-5fcf828d8823" colab={"base_uri": "https://localhost:8080/", "height": 280}
plt.scatter(df[interest], df[price], alpha=.01)
plt.xlabel('interest score')
plt.ylabel('price');
# + id="yfIGoH1Hrl3o" outputId="28718fde-df9c-4721-fa10-759d7a57651c" colab={"base_uri": "https://localhost:8080/", "height": 280}
plt.scatter(df[amenities], df[price], alpha=.1)
plt.xlabel('amenity score')
plt.ylabel('price');
# + [markdown] id="eYMNPeK9eJjc"
# # Specify Target vector and feature maxtix
# + id="kietpehPeiz_"
y = df[price]
X = df[[bathrooms,
bedrooms,
description,
lat,
lon,
interest,
amenities
]]
# + [markdown] id="QvzK-_ywdKzJ"
# #Split Data between **train set** and **test set**
# + id="T3Z2JSgWbBlX"
#train on data before june 2016.
cutoff = "2016-06-01"
mask = X.index < '2016-06-01'
X_train, y_train = X.loc[mask], y.loc[mask]
X_test, y_test = X.loc[~mask], y.loc[~mask]
# + id="N_Ovy13yumQ_"
assert len(X) == len(X_train) + len(X_test)
# + id="lpUR-JgqxS8F" outputId="bdacf529-35de-414d-fec5-0d98fd4cdaf1" colab={"base_uri": "https://localhost:8080/", "height": 265}
X_train.info()
# + [markdown] id="mMrHgF50wbh9"
# ##Baseline
# + id="_ek2AMKVvfRa" outputId="4b0178c6-b9ec-4b2d-d951-25613fca2a20" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.metrics import mean_absolute_error
y_pred = [y_train.mean()] * len(y_train)
print('Baseline MAE:', mean_absolute_error(y_train, y_pred))
# + [markdown] id="5BKa1_tSwrFX"
# ##Build Model
# + id="zBN8qcocwIIN" outputId="5edb2678-43b9-4687-ec4c-694a369fd67c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Step 1: Import our predictor
from sklearn.linear_model import LinearRegression
# Step 2: Instantiate our predictor
model = LinearRegression()
# Step 3: Fit our model to the TRAINING DATA
model.fit(X_train, y_train)
# Step 4: Make predictions... see below
# + id="S5A3fdC1w0zw" outputId="229bde19-55ad-4d24-c43d-929f48d0fc80" colab={"base_uri": "https://localhost:8080/", "height": 52}
print('Training MAE:', mean_absolute_error(y_train, model.predict(X_train)))
print('Testing MAE:', mean_absolute_error(y_test, model.predict(X_test)))
# + id="MkiFyfKzzS2F" outputId="ead5ca6c-e131-457e-a4f1-2d12bdc31dec" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.metrics import mean_squared_error
print('Training RMSE:', mean_squared_error(y_train, model.predict(X_train), squared=False))
print('Testing RMSE:', mean_squared_error(y_test, model.predict(X_test), squared=False))
# + id="YwxIx5x4zhEG" outputId="9bfbc6c1-f988-4cbf-89a0-c5453fc66b8e" colab={"base_uri": "https://localhost:8080/", "height": 52}
print('Training R^2:', model.score(X_train, y_train))
print('Testing R^2:', model.score(X_test, y_test))
# + id="OJ52IUqNzwn_"
| module2-regression-2/Micah_DS_212_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Introduction to [Plotly](https://plotly.com/)
#
# Plotly is a versatile interactive plotting package that can be used with Python and Javascript and also through an online editor (without the need for coding).
# ## Why/When to use Plotly (my 2 cents)
#
# If you already know Python and you don't really want to learn another coding language, but you do want to create interactive figures (e.g., within a Jupyter notebook and/or for use on a website), you should look into Plotly.
#
# In particular, [Plotly express](https://plotly.com/python/plotly-express/) is a fantastic tool for generating quick interactive figures without much code. Plotly express covers a good amount of ground, and you may be able to do all/most your work within Plotly express, depending on your specific needs. In this workshop, I'll show you Plotly express, but then move beyond it for the majority of the content.
#
# Though you can do a lot with Plotly, it definitely has limitations (some of which we'll see in this workshop). Also, as with all of the ready-made interactive plot solutions (e.g., [Bokeh](https://docs.bokeh.org/en/latest/), [Altair](https://altair-viz.github.io/), [Glue](https://glueviz.org/), etc.), Plotly has a specific look, which can only be tweaked to a certain extent. If you like the look well enough and you don't mind the limitations, then it's a good choice.
# ## In this tutorial...
#
# We will explore the basics of the Python version, using COVID-19 data from the following sources:
#
# - COVID-19 data from the WHO: https://covid19.who.int/info/
# - GDP Data from the World Bank: https://data.worldbank.org/indicator/NY.GDP.MKTP.CD
#
# I will make two plots, one comparing COVID-19 data to GDPs and another showing COVID-19 data as a function of time.
# ## Installation
#
# I recommend installing Python using [Anaconda](https://www.anaconda.com/products/individual). Then you can create and activate a new environment for this workshop by typing the following commands into your (bash) terminal.
#
# ```
# $ conda create -n plotly-env python=3.9 jupyter pandas plotly statsmodels
# $ conda activate plotly-env
# ```
# ## Import the relevant packages that we will use.
# +
import pandas as pd
import numpy as np
import scipy.stats
import plotly.graph_objects as go
from plotly.subplots import make_subplots
import plotly.express as px
# -
# ## 1. Create a plot showing COVID-19 and GDP data.
# ### 1.1. Read in the data.
#
# I will join multiple data tables together, using the *pandas* package so that I have one DataFrame containing all values for a given country.
# Current cumulative COVID-19 data from the WHO.
# dfCT = pd.read_csv('data/WHO-COVID/WHO-COVID-19-global-table-data.csv') # in case the WHO server goes down
dfCT = pd.read_csv('https://covid19.who.int/WHO-COVID-19-global-table-data.csv', index_col=False)
dfCT
# Current vaccination data from the WHO
# dfV = pd.read_csv('data/WHO-COVID/vaccination-data.csv') # in case the WHO server goes down
dfV = pd.read_csv('https://covid19.who.int/who-data/vaccination-data.csv')
dfV
# +
# Vaccination metadata from the WHO; this file contains the start dates (and end dates) for vaccines for each country.
# dfVM = pd.read_csv('data/WHO-COVID/vaccination-metadata.csv') # in case the WHO server goes down
dfVM = pd.read_csv('https://covid19.who.int/who-data/vaccination-metadata.csv')
# drop columns without a start date
dfVM.dropna(subset = ['START_DATE'], inplace = True)
# convert the date columns to datetime objects for easier plotting and manipulation later on
dfVM['AUTHORIZATION_DATE'] = pd.to_datetime(dfVM['AUTHORIZATION_DATE'])
dfVM['START_DATE'] = pd.to_datetime(dfVM['START_DATE'])
dfVM['END_DATE'] = pd.to_datetime(dfVM['END_DATE'])
# I will simplify this table to just take the earliest start date for a given country
# sort by the start date and country code
dfVM.sort_values(['START_DATE', 'ISO3'], ascending = (True, True), inplace = True)
# take only the first entry for a given country
dfVM.drop_duplicates(subset = 'ISO3', keep = 'first', inplace = True)
dfVM
# -
# GDP data from the World Bank (the first three rows do not contain data)
# I don't think there's a direct link to this data on their server (but I didn't look very hard)
dfM = pd.read_csv('data/WorldBank/API_NY.GDP.MKTP.CD_DS2_en_csv_v2_3469429.csv', skiprows = 3)
dfM
# +
# Join these 4 tables so that I have one DataFrame with all values for a given country.
# I will start by joining the two vaccination data tables.
dfJ1 = dfV.join(dfVM.set_index('ISO3'), on = 'ISO3', how = 'left', rsuffix = '_meta')
# Next I will join this with the COVID-19 data table.
# First rename this column in the COVID-19 data so that it is the same as the vaccine data. Then I will join on that column.
dfCT.rename(columns = {'Name':'COUNTRY'}, inplace = True)
dfJ2 = dfJ1.join(dfCT.set_index('COUNTRY'), on = 'COUNTRY', how = 'left')
# Finally, I will join in the GDP data from the World Bank.
# I will rename a column in the World Bank data to match a column in the joined data above.
dfM.rename(columns = {'Country Code':'ISO3'}, inplace = True)
dfJoinedCOVID = dfJ2.join(dfM.set_index('ISO3'), on = 'ISO3', how = 'left')
dfJoinedCOVID
# -
# ### 1.2. Create a couple simple Plotly figures using [Plotly express](https://plotly.com/python/plotly-express/).
#
# Plotly express is a simplified version of the Plotly interface for Python that allows users to create many types of Plotly figures with single lines of code. This greatly simplifies the workflow for some kinds of Plotly figures. We will start with Plotly express (and for some of your use cases, that may be enough), but we will move on to full blown Plotly for the rest of this workshop.
# In this plot, I will show total vaccinations vs. GDP with the point size scaled by the total cumulative COVID-19 cases.
# Note: We imported plotly.express as px
# I will create a scatter plot using the DataFrame I created above, define the keys for the x and y data,
# plot the y axis in the log, and also size each data point by the number of COVID-19 cases.
# A nice part of plotly express is that you can add a trend line very easily.
fig = px.scatter(dfJoinedCOVID, x = 'TOTAL_VACCINATIONS_PER100', y = '2020', log_y = True,
size = np.clip(np.nan_to_num(dfJoinedCOVID['Cases - cumulative total per 100000 population']/500.), 5, 100),
trendline = 'ols', trendline_options = dict(log_y = True)
)
fig.show()
# Lets also plot the first vaccination start date vs. GDP, with the size based on the total vaccionations. In this example, I will also modify the hover and axis attributes.
# +
# The command is similar to that from the previous cell, but here I'm also defining the data shown on hover in the tooltips.
# (It's not quite as easy to add a trendline here when plotting dates, though it is possible.)
fig = px.scatter(dfJoinedCOVID, x = 'START_DATE', y = '2020', log_y = True,
size = np.nan_to_num(dfJoinedCOVID['TOTAL_VACCINATIONS_PER100']),
hover_name = 'COUNTRY',
hover_data = ['2020',
'START_DATE',
'TOTAL_VACCINATIONS_PER100',
'Cases - cumulative total per 100000 population'
]
)
# a few manipulations to the axes
fig.update_xaxes(title = 'Vaccine Start Date', range = [np.datetime64('2020-07-01'), np.datetime64('2021-07-01')])
fig.update_yaxes(title = '2020 GDP (USD)')
fig.show()
# -
# ### *Exercise 1: Create your own plot using Plotly express.*
#
# Use the data we read in above (or your own data). You can start with one of the commands above or choose a different style of plot. Whichever format you use, choose different columns to plot than above. Try to also add a new option to the command to change the plot.
#
# Hint: Go to the [Plotly express homepage](https://plotly.com/python/plotly-express/), and click on a link to see many examples (e.g., [here's the page for the scatter plot](https://plotly.com/python/line-and-scatter/))
# Create a plot using Plotly express
# ### 1.4. Create the plot using the standard Plotly [Graph Object](https://plotly.com/python/graph-objects/).
#
# For the remainder of the workshop we will use Graph Objects for our Plotly figures. One motivation here is so that I can create multiple panels in one figure, which can be downloaded to an html file. (Plotly express will only make an individual figure, and does not support arbitrary subplots.)
#
# First you create a <b>"trace"</b>, which holds the data. There are many kinds of traces available in Plotly. (e.g., bar, scatter, etc.). For this example, we will use a scatter trace. (Interestingly, the scatter trace object also includes line traces, accessed by changing the "mode" key. I will show the line version later on.)
#
# Then you create a figure and add the trace to that figure. A single figure can have multiple traces.
# +
# Create a plot using Plotly Graph Objects(s)
# Note: We imported the plotly.graph_objects as go.
# create the trace
trace1 = go.Scatter(x = dfJoinedCOVID['TOTAL_VACCINATIONS_PER100'], y = dfJoinedCOVID['2020'], # x and y values for the plot
mode = 'markers', # setting mode to markers produces a typical scatter plot
)
# create the figure
fig = go.Figure()
# add the trace and update a few parameters for the axes
fig.add_trace(trace1)
fig.update_xaxes(title = 'Total Vaccionations Per 100 People', range=[0,300])
fig.update_yaxes(title = 'GDP (USD)', type = 'log')
fig.show()
# -
# Re-create this figure with more customizations.
# +
# Note: We imported the plotly.graph_objects as go.
# create the trace and set various parameters
trace1 = go.Scatter(x = dfJoinedCOVID['TOTAL_VACCINATIONS_PER100'], y = dfJoinedCOVID['2020'], # x and y values for the plot
mode = 'markers', # setting mode to markers produces a typical scatter plot
showlegend = False, # since I will only have one trace, I don't need a legend
name = 'COVID Vaccines', # name for the legend and tooltips (this is not strictly necessary here)
# set various parameters for the markers in the following dict, e.g., color, opacity, size, outline, etc.
marker = dict(
color = 'rgba(0, 0, 0, 0.2)',
opacity = 1,
size = np.nan_to_num(np.clip(dfJoinedCOVID['Cases - cumulative total per 100000 population']/1000., 5, 100)),
line = dict(
color = 'rgba(0, 0, 0, 1)',
width = 1
),
),
# set a template for the tooltips below.
# hovertemplate can accept the x and y data and additional "text" as defined by a separate input
# Note, the "<extra></extra>" is included to remove some formatting that plotly imposes on tooltips
hovertemplate = '%{text}' +
'Total Vaccinations / 100 people: %{x}<br><extra></extra>' +
'GDP: $%{y}<br>',
# additional text to add to the hovertemplate. This needs to be a list with the same length and the x and y data.
text = ['Country: {}<br>Total COVID Cases / 100,000 people: {}<br>Vaccine start date: {}<br>'.format(x1, x2, x3)
for (x1, x2, x3) in zip(dfJoinedCOVID['COUNTRY'],
dfJoinedCOVID['Cases - cumulative total per 100000 population'],
dfJoinedCOVID['START_DATE'].dt.strftime('%b %Y'))
],
# style the tooltip as desired
hoverlabel = dict(
bgcolor = 'white',
)
)
# Add a trendline
# I will use scipy.stats.linregress (and fit to the log of the GDP)
dfFit1 = dfJoinedCOVID.dropna(subset = ['TOTAL_VACCINATIONS_PER100', '2020'])
slope1, intercept1, r1, p1, se1 = scipy.stats.linregress(dfFit1['TOTAL_VACCINATIONS_PER100'], np.log10(dfFit1['2020']))
xFit1 = np.linspace(0, 300, 100)
yFit1 = 10.**(slope1*xFit1 + intercept1)
trace1F = go.Scatter(x = xFit1, y = yFit1,
mode = 'lines', # Set the mode the lines (rather than markers) to show a line.
opacity = 1,
marker_color = 'black',
showlegend = False,
hoverinfo='skip' # Don't show anything on hover. (We could show the trendline info, but I'll leave that out for now.)
)
# create the figure
fig = go.Figure()
# add the trace and update a few parameters for the axes
fig.add_trace(trace1)
fig.add_trace(trace1F)
fig.update_xaxes(title = 'Total Vaccionations Per 100 People', range=[0,300])
fig.update_yaxes(title = 'GDP (USD)', type = 'log')
fig.show()
# -
# ### *Exercise 2: Create your own plot using Plotly Graph Object(s).*
#
# Use the data we read in above (or your own data). You can start with one of the commands above or choose a different style of plot. Whichever format you use, choose different columns to plot than above. Try to also add a new option to the command to change the plot.
#
# Hint: The Plotly help pages usually contain examples for both Plotly express and Graph Object. If you go to the [Plotly express homepage](https://plotly.com/python/plotly-express/) and click on a link (e.g., [the page for the scatter plot](https://plotly.com/python/line-and-scatter/)), you can scroll down to see Graph Object examples.
# +
# Create a plot using Plotly Graph Objects(s)
# First, create the trace
# Second, create the figure and show it
# -
# ### 1.5. Show two plots side-by-side sharing the y axis.
# +
# Create the trace for the 2nd figure (similar method to above).
trace2 = go.Scatter(x = dfJoinedCOVID['START_DATE'], y = dfJoinedCOVID['2020'],
mode = 'markers',
showlegend = False,
name = 'COVID Vaccines',
marker = dict(
color = 'rgba(0, 0, 0, 0.2)',
opacity = 1,
size = np.nan_to_num(np.clip(dfJoinedCOVID['TOTAL_VACCINATIONS_PER100']/7., 5, 100)),
line = dict(
color = 'rgba(0, 0, 0, 1)',
width = 1
),
),
hovertemplate = '%{text}' +
'Vaccine start date: %{x}<br><extra></extra>' +
'GDP: $%{y}<br>',
text = ['Country: {}<br>Total COVID Cases / 100,000 people: {}<br>Total Vaccinations / 100 people: {}<br>'.format(x1, x2, x3)
for (x1, x2, x3) in zip(dfJoinedCOVID['COUNTRY'],
dfJoinedCOVID['Cases - cumulative total per 100000 population'],
dfJoinedCOVID['TOTAL_VACCINATIONS_PER100'])
],
hoverlabel=dict(
bgcolor = 'white',
)
)
# Add trendlines
dfFit2 = dfJoinedCOVID.dropna(subset = ['START_DATE', '2020'])
delta = (dfFit2['START_DATE'] - dfFit2['START_DATE'].min())/np.timedelta64(1,'D')
slope2, intercept2, r2, p2, se2 = scipy.stats.linregress(delta, np.log10(dfFit2['2020']))
xx2 = np.linspace(0, 500, 100)
yFit2 = 10.**(slope2*xx2 + intercept2)
xFit2 = xx2*np.timedelta64(1,'D') + dfFit2['START_DATE'].min()
trace2F = go.Scatter(x = xFit2, y = yFit2,
mode = 'lines',
opacity = 1,
marker_color = 'black',
showlegend = False,
hoverinfo='skip'
)
# Create the figure and add the traces
# I will use Plotly's "make_subplots" method (imported above).
# Define the number of rows and columns, the column_widths, spacing, and here I will share the y axis.
# Sharing the y axis means that if you zoom/pan on one plot, the other will also zoom/pan.
fig = make_subplots(rows = 1, cols = 2, column_widths = [0.5, 0.5], horizontal_spacing = 0.01, shared_yaxes = True)
# Add the first trace and update the axes.
# Note that I specify which row and column within each of these commands.
fig.add_trace(trace1, row = 1, col = 1)
fig.add_trace(trace1F, row = 1, col = 1)
fig.update_xaxes(title = 'Total Vaccionations Per 100 People', range=[0,280], row = 1, col = 1)
fig.update_yaxes(title = 'GDP (USD)', type = 'log', row = 1, col = 1)
# Add the second trace and update the axes.
# Note that I am using numpy's datetime64 data types in order to set the axis range here
fig.add_trace(trace2, row = 1, col = 2)
fig.add_trace(trace2F, row = 1, col = 2)
fig.update_xaxes(title = 'Vaccine Start Date', range = [np.datetime64('2020-07-02'),
np.datetime64('2021-07-01')], row = 1, col = 2)
fig.update_yaxes(type = 'log', row = 1, col = 2)
# Provide an overall title to the figure.
fig.update_layout(title_text = 'COVID-19 Vaccine Equity')
# Add annotations to tell what the symbol sizes mean.
# I will position these relative to the data domain, and therefore they will not move around when zooming and panning.
fig.add_annotation(x = 0.01, y = 0.99, row = 1, col = 1, showarrow = False,
xref = 'x domain', yref = 'y domain',
text = 'Symbol size indicates total COVID-19 cases.')
fig.add_annotation(x = 0.01, y = 0.99, row = 1, col = 2, showarrow = False,
xref = 'x domain', yref = 'y domain',
text = 'Symbol size indicates total vaccinations.')
# Show the final result
fig.show()
# -
# #### You can save the figure in html format to use on a website.
fig.write_html('plotly_graph.html')
# ## 2. Create a plot showing COVID-19 cases and deaths vs. time for a given country.
#
# I will also include [custom buttons](https://plotly.com/python/custom-buttons/) to toggle between various ways of viewing the data.
# ### 2.1. Read in the data
# +
# COVID-19 cases and deaths as a function of time for multiple countries
# dfC = pd.read_csv('data/WHO-COVID/WHO-COVID-19-global-data.csv') # in case the WHO server goes down
dfC = pd.read_csv('https://covid19.who.int/WHO-COVID-19-global-data.csv')
# convert the date column to datetime objects for easier plotting and manipulation later on
dfC['Date_reported'] = pd.to_datetime(dfC['Date_reported'])
dfC
# -
# ### 2.2. Choose a country, and then create the plot,
country = 'United States of America'
# Select only the data that is from the country.
use3 = dfC.loc[dfC['Country'] == country]
# +
# Create the trace.
# In this example I will use a bar chart.
trace3 = go.Bar(x = use3['Date_reported'], y = use3['New_cases'],
opacity = 1,
marker_color = 'black',
showlegend = False,
name = 'COVID Cases'
)
# Create the figure.
fig = go.Figure()
# Add the trace and update a few parameters for the axes.
fig.add_trace(trace3)
fig.update_xaxes(title = 'Date')
fig.update_yaxes(title = 'Total COVID-19 Cases')
fig.show()
# -
# #### Let's improve this plot.
#
# - I want to take a rolling average (this is easily done with *pandas*).
# - I'd prefer a filled region rather than bars.
# Define the number of days to use for the rolling average.
rollingAve = 7
# +
# Create the trace, using Scatter to create lines and fill the region between the line and y=0.
trace3 = go.Scatter(x = use3['Date_reported'], y = use3['New_cases'].rolling(rollingAve).mean(),
mode = 'lines', # Set the mode the lines (rather than markers) to show a line.
opacity = 1,
marker_color = 'black',
fill = 'tozeroy', # This will fill between the line and y=0.
showlegend = False,
name = 'COVID Count',
hovertemplate = 'Date: %{x}<br>Number: %{y}<extra></extra>', #Note: the <extra></extra> removes the trace label.
)
# Create the figure.
fig = go.Figure()
# Add the trace and update a few parameters for the axes.
fig.add_trace(trace3)
fig.update_xaxes(title = 'Date')
fig.update_yaxes(title = 'Total COVID-19 Cases')
fig.show()
# -
# ### *Exercise 3: Create your own plot showing COVID-19 deaths vs time.*
#
# You can use either Plotly express or Graph Objects. Try to pick a different country than I used above. Also try to use a different style than I plotted above.
# +
# Create a Plotly figure showing COVID-19 deaths vs. time
# -
# ### 2.3. Add some buttons to interactively change the plot.
#
# I want to be able to toggle between cumulative vs. total as well as cases vs. death. We can do this with [custom buttons](https://plotly.com/python/custom-buttons/) that will "restyle" the plot.
#
# You can also create interactions with buttons and other "widgets" using [dash](https://plotly.com/dash/), but we won't go there in this workshop.
# +
# Create the figure.
fig = go.Figure()
# For this scenario, I am going to add each of the 4 traces to the plot but only show one at a time
# Add traces for each column
columns = ['New_cases', 'New_deaths', 'Cumulative_cases', 'Cumulative_deaths']
for i, c in enumerate(columns):
visible = False
if (i == 0):
visible = True
# Create the trace, using Scatter to create lines and fill the region between the line and y=0.
trace = go.Scatter(x = use3['Date_reported'], y = use3[c].rolling(rollingAve).mean(),
mode = 'lines', # Set the mode the lines (rather than markers) to show a line.
opacity = 1,
marker_color = 'black',
fill = 'tozeroy', # This will fill between the line and y=0.
showlegend = False,
name = 'COVID Count',
hovertemplate = 'Date: %{x}<br>Number: %{y}<extra></extra>', #Note: the <extra></extra> removes the trace label.
visible = visible
)
# Add that trace to the figure
fig.add_trace(trace)
# Update a few parameters for the axes.
# Note: I added a margin to the top ('t') of the plot within fig.update_layout to make room for the buttons.
fig.update_xaxes(title = 'Date')#, range = [np.datetime64('2020-03-01'), np.datetime64('2022-01-12')])
fig.update_yaxes(title = 'COVID-19 Count')
fig.update_layout(title_text = 'COVID-19 Data Explorer : '+ country + '<br>(' + str(rollingAve) +'-day rolling average)',
margin = dict(t = 150)
)
# Add buttons (this can also be easily done with dash, but then you can't export easily to html).
fig.update_layout(
updatemenus = [
# Buttons for choosing the data to plot.
dict(
type = 'buttons',
direction = 'left', # This defines what orientation to include all buttons. 'left' shows them in one row.
buttons = list([
dict(
# 'args' tells the button what to do when clicked.
# In this case it will change the visibility of the traces
# 'label' is the text that will be displayed on the button
# 'method' is the type of action the button will take.
# method = 'restyle' allows you to redefine certain preset plot styles (including the visible key).
# See https://plotly.com/python/custom-buttons/ for different methods and their uses
args = [{'visible': [True, False, False, False]}],
label = 'Daily Cases',
method = 'restyle'
),
dict(
args = [{'visible': [False, True, False, False]}],
label = 'Daily Deaths',
method = 'restyle'
),
dict(
args = [{'visible': [False, False, True, False]}],
label = 'Cumulative Cases',
method = 'restyle'
),
dict(
args = [{'visible': [False, False, False, True]}],
label = 'Cumulative Deaths',
method = 'restyle'
)
]),
showactive = True, # Highlight the active button
# Below is for positioning
x = 0.0,
xanchor = 'left',
y = 1.13,
yanchor = 'top'
),
]
)
fig.show()
# -
# ### *Exercise 4: Convert the buttons into a dropdown menu.*
#
# Using the code that I wrote above, create a plot of COVID-19 cases vs. time (for a single country) that uses a dropdown menu to choose between "Daily Cases", "Cumulative Cases", "Daily Deaths" and "Cumulative Deaths".
#
# Hint: [This website gives examples of dropdown menus in Plotly](https://plotly.com/python/dropdowns/).
# +
# Create a plot of COVID-19 cases vs. time with a dropdown menu to change the data that is plotted.
# You can use the same trace and figure from above or create new ones.
# The code will be VERY similar to above. In fact, you could solve this exercise with only 3 lines of code...
# -
# ## 3. Put all of these plots together into one "dashboard".
#
# ### 3.1. I will put all the plotting commands (from above) into a single function.
#
# That way I can reuse this later on for the final step (#4 below). This is mostly copying and pasting, but with some additions that I will point out below in the comments.
# +
# In order to reduce the lines of code, I created a function that generates the vaccine trace, given inputs
def generateVaccineTrace(xData, yData, size, color, hovertemplate, text, hoverbg = 'white'):
'''
xData : the x data for the trace
yData : the y data for the trace
size : sizes for the data points
color : color for the markers
hovertemplate : the template for the tooltip
text : the additional text to include in the tooltip
hoverbg : optional parameter to set the background color of the tooltip (defaut is white)
'''
trace = go.Scatter(x = xData, y = yData,
mode = 'markers',
showlegend = False,
name = 'COVID Vaccines',
marker = dict(
color = color,
opacity = 1,
size = size,
line = dict(
color = 'rgba(0, 0, 0, 1)',
width = 1
),
),
hovertemplate = hovertemplate,
text = text,
hoverlabel = dict(
bgcolor = hoverbg,
),
)
return trace
# This is a large function that will generate the entire figure with all the subplots
def generateFigure(co):
'''
co : the country that we want to plot
'''
##################################
# First, create the traces.
##################################
# cases over time
useC = dfC.loc[dfC['Country'] == co]
traces1 = []
columns = ['New_cases', 'New_deaths', 'Cumulative_cases', 'Cumulative_deaths']
for i, c in enumerate(columns):
visible = False
if (i == 0):
visible = True
# Create the trace, using Scatter to create lines and fill the region between the line and y=0.
trace = go.Scatter(x = use3['Date_reported'], y = use3[c].rolling(rollingAve).mean(),
mode = 'lines', # Set the mode the lines (rather than markers) to show a line.
opacity = 1,
marker_color = 'black',
fill = 'tozeroy', # This will fill between the line and y=0.
showlegend = False,
name = 'COVID Count',
hovertemplate = 'Date: %{x}<br>Number: %{y}<extra></extra>', #Note: the <extra></extra> removes the trace label.
visible = visible
)
traces1.append(trace)
# vaccine fraction vs. GDP (using the function that I wrote above)
trace2 = generateVaccineTrace(dfJoinedCOVID['TOTAL_VACCINATIONS_PER100'], dfJoinedCOVID['2020'],
np.nan_to_num(np.clip(dfJoinedCOVID['Cases - cumulative total per 100000 population']/1000., 5, 100)),
'rgba(0, 0, 0, 0.2)',
'%{text}' + \
'Total Vaccinations / 100 people: %{x}<br><extra></extra>' + \
'GDP: $%{y}<br>',
['Country: {}<br>Total COVID Cases / 100,000 people: {}<br>Vaccine start date: {}<br>'.format(x1, x2, x3)
for (x1, x2, x3) in zip(dfJoinedCOVID['COUNTRY'],
dfJoinedCOVID['Cases - cumulative total per 100000 population'],
dfJoinedCOVID['START_DATE'].dt.strftime('%b %Y'))
],
)
# vaccine start date vs. GDP (using the function that I wrote above)
trace3 = generateVaccineTrace(dfJoinedCOVID['START_DATE'], dfJoinedCOVID['2020'],
np.nan_to_num(np.clip(dfJoinedCOVID['TOTAL_VACCINATIONS_PER100']/7., 5, 100)),
'rgba(0, 0, 0, 0.2)',
'%{text}' + \
'Vaccine start date: %{x}<br><extra></extra>' + \
'GDP: $%{y}<br>',
['Country: {}<br>Total COVID Cases / 100,000 people: {}<br>Total Vaccinations / 100 people: {}<br>'.format(x1, x2, x3)
for (x1, x2, x3) in zip(dfJoinedCOVID['COUNTRY'],
dfJoinedCOVID['Cases - cumulative total per 100000 population'],
dfJoinedCOVID['TOTAL_VACCINATIONS_PER100'])
],
)
# Add trendlines
# This is simply copied from above
dfFit1 = dfJoinedCOVID.dropna(subset = ['TOTAL_VACCINATIONS_PER100', '2020'])
slope1, intercept1, r1, p1, se1 = scipy.stats.linregress(dfFit1['TOTAL_VACCINATIONS_PER100'], np.log10(dfFit1['2020']))
xFit1 = np.linspace(0, 300, 100)
yFit1 = 10.**(slope1*xFit1 + intercept1)
trace2F = go.Scatter(x = xFit1, y = yFit1,
mode = 'lines', # Set the mode the lines (rather than markers) to show a line.
opacity = 1,
marker_color = 'black',
showlegend = False,
hoverinfo='skip' # Don't show anything on hover. (We could show the trendline info, but I'll leave that out for now.)
)
dfFit2 = dfJoinedCOVID.dropna(subset = ['START_DATE', '2020'])
delta = (dfFit2['START_DATE'] - dfFit2['START_DATE'].min())/np.timedelta64(1,'D')
slope2, intercept2, r2, p2, se2 = scipy.stats.linregress(delta, np.log10(dfFit2['2020']))
xx2 = np.linspace(0, 500, 100)
yFit2 = 10.**(slope2*xx2 + intercept2)
xFit2 = xx2*np.timedelta64(1,'D') + dfFit2['START_DATE'].min()
trace3F = go.Scatter(x = xFit2, y = yFit2,
mode = 'lines',
opacity = 1,
marker_color = 'black',
showlegend = False,
hoverinfo='skip'
)
# Add 2 more traces for the vaccine plots to highlight the selected country (using the function that I wrote above).
# These are nearly identical to the 2 traces from above but using the limitted useH dataset (below) and colored red.
useH = dfJoinedCOVID.loc[dfJoinedCOVID['COUNTRY'] == co]
trace2H = generateVaccineTrace(useH['TOTAL_VACCINATIONS_PER100'], useH['2020'],
np.nan_to_num(np.clip(useH['Cases - cumulative total per 100000 population']/500., 5, 100)),
'rgba(255, 0, 0, 1)',
'%{text}' + \
'Total Vaccinations / 100 people: %{x}<br><extra></extra>' + \
'GDP: $%{y}<br>',
['Country: {}<br>Total COVID Cases / 100,000 people: {}<br>Vaccine start date: {}<br>'.format(x1, x2, x3)
for (x1, x2, x3) in zip(useH['COUNTRY'],
useH['Cases - cumulative total per 100000 population'],
useH['START_DATE'].dt.strftime('%b %Y'))
],
hoverbg = 'red'
)
trace3H = generateVaccineTrace(useH['START_DATE'], useH['2020'],
np.clip(useH['TOTAL_VACCINATIONS_PER100']/7., 5, 100),
'rgba(255, 0, 0, 1)',
'%{text}' + \
'Vaccine start date: %{x}<br><extra></extra>' + \
'GDP: $%{y}<br>',
['Country: {}<br>Total COVID Cases / 100,000 people: {}<br>Total Vaccinations / 100 people: {}<br>'.format(x1, x2, x3)
for (x1, x2, x3) in zip(useH['COUNTRY'],
useH['Cases - cumulative total per 100000 population'],
useH['TOTAL_VACCINATIONS_PER100'])
],
hoverbg = 'red'
)
##################################
# Second, create the figure and add the traces.
##################################
# I will create a subplot object where
# - the top will have 1 column and contain the cases over time,
# - the bottom will be split in two columns for the vaccine plots,
# - and the bottom two columns will share the y axis.
fig = make_subplots(rows = 2, cols = 2, shared_yaxes = True,
column_widths = [0.5, 0.5],
row_heights = [0.35, 0.65],
specs = [ [{"colspan": 2}, None], [{}, {}] ], # here is where I define that the first row only has one column
horizontal_spacing = 0.01,
vertical_spacing = 0.08
)
# Add in the traces and update the axes (specifying with row and column they below to)
for t in traces1:
fig.add_trace(t, row = 1, col = 1)
fig.update_xaxes(title = 'Date')#, range = [np.datetime64('2020-03-01'), np.datetime64('2022-01-12')], row = 1, col = 1)
fig.update_yaxes(title = 'COVID-19 Count', row = 1, col = 1, rangemode = 'nonnegative')
fig.add_trace(trace2, row = 2, col = 1)
fig.add_trace(trace2F, row = 2, col = 1)
fig.add_trace(trace2H, row = 2, col = 1)
fig.update_xaxes(title = 'Total Vaccionations Per 100 People', range=[0,280], row = 2, col = 1)
fig.update_yaxes(title = 'GDP (USD)', type = 'log', row = 2, col = 1)
fig.add_trace(trace3, row = 2, col = 2)
fig.add_trace(trace3F, row = 2, col = 2)
fig.add_trace(trace3H, row = 2, col = 2)
fig.update_xaxes(title = 'Vaccine Start Date', range = [np.datetime64('2020-07-02'),
np.datetime64('2021-07-01')], row = 2, col = 2)
fig.update_yaxes(type = 'log', row = 2, col = 2)
# Add a title and define the size and margin.
fig.update_layout(title_text = 'COVID-19 Data Explorer : '+ co + '<br>(' + str(rollingAve) +'-day rolling average)',
title_y = 0.97,
height = 1000,
width = 1000,
margin = dict(t = 120))
# Add the annotations to tell what the symbol sizes mean.
fig.add_annotation(x = 0.01, y = 0.99, row = 2, col = 1, showarrow = False,
xref = 'x domain', yref = 'y domain',
text = 'Symbol size indicates total COVID-19 cases.')
fig.add_annotation(x = 0.01, y = 0.99, row = 2, col = 2, showarrow = False,
xref = 'x domain', yref = 'y domain',
text = 'Symbol size indicates total vaccinations.')
return fig
# -
def addButtons(fig):
##################################
# Third, add the buttons.
##################################
# Note that here in 'args' I need to provide values for all the traces (even though only one plot will change).
fig.update_layout(
updatemenus = [
dict(
type = 'buttons',
direction = 'left',
buttons = list([
dict(
args = [{'visible': [True, False, False, False, True, True, True, True, True, True]}],
label = 'Daily Cases',
method = 'restyle'
),
dict(
args = [{'visible': [False, True, False, False, True, True, True, True, True, True]}],
label = 'Daily Deaths',
method = 'restyle'
),
dict(
args = [{'visible': [False, False, True, False, True, True, True, True, True, True]}],
label = 'Cumulative Cases',
method = 'restyle'
),
dict(
args = [{'visible': [False, False, False, True, True, True, True, True, True, True]}],
label = 'Cumulative Deaths',
method = 'restyle'
)
]),
showactive = True,
x = 0.0,
xanchor = 'left',
y = 1.05,
yanchor = 'top'
),
]
)
return fig
# Use the functions to create the figure.
fig = generateFigure(country)
fig = addButtons(fig)
fig.show()
# ## 4. *Bonus:* Add a dropdown to choose the country.
#
# I'd like to be able to interactively choose the country to plot using a dropdown. Plotly allows for [dropdown menus](https://plotly.com/python/dropdowns/) within a figure, in a similar method to how we added the buttons. This code will become a bit combersome, but we'll walk through it together.
#
# In general, we will use the "update" mode (rather than "restyle") for the dropdown menu. Update will allow us to change the data being plotted, using an external function (often called a "callback" function). The rest of the syntax for the dropdown menu will be very similar to the buttons.
#
# Again, this could also be accomplished in [dash](https://plotly.com/dash/), but we're not going there in this workshop. (Note that with dash you can build an app that would do this, but hosting it online is much harder than simply downloading the Plotly html file and sticking it on your website.)
# +
# Define functions that will update the data being shows in the time plot
# For the time series plot
# Since there are actually 4 traces in the time plot (only 1 visible), I will need to send 4 data sets back from each function
# This one I can just call 4 times
def updateTimePlotX(co):
use = dfC.loc[dfC['Country'] == co]
return use['Date_reported']
# There may be a smarter way to do this, but I will write 4 functions here
def updateTimePlotY1(co):
use = dfC.loc[dfC['Country'] == co]
return use['New_cases'].rolling(rollingAve).mean()
def updateTimePlotY2(co):
use = dfC.loc[dfC['Country'] == co]
return use['New_deaths'].rolling(rollingAve).mean()
def updateTimePlotY3(co):
use = dfC.loc[dfC['Country'] == co]
return use['Cumulative_cases'].rolling(rollingAve).mean()
def updateTimePlotY4(co):
use = dfC.loc[dfC['Country'] == co]
return use['Cumulative_deaths'].rolling(rollingAve).mean()
# For the vaccine data highlights
def updateVaccinePlotX1(co):
use = dfJoinedCOVID.loc[dfJoinedCOVID['COUNTRY'] == co]
return use['TOTAL_VACCINATIONS_PER100']
def updateVaccinePlotX2(co):
use = dfJoinedCOVID.loc[dfJoinedCOVID['COUNTRY'] == co]
return use['START_DATE']
def updateVaccinePlotY(co):
use = dfJoinedCOVID.loc[dfJoinedCOVID['COUNTRY'] == co]
return use['2020']
# Functions to help with the vaccine tooltip text
def getVaccineTextAll():
return ['Country: {}<br>Total COVID Cases / 100,000 people: {}<br>Vaccine start date: {}<br>'.format(x1, x2, x3)
for (x1, x2, x3) in zip(dfJoinedCOVID['COUNTRY'],
dfJoinedCOVID['Cases - cumulative total per 100000 population'],
dfJoinedCOVID['START_DATE'].dt.strftime('%b %Y'))
]
def updateVaccineHText(co):
use = dfJoinedCOVID.loc[dfJoinedCOVID['COUNTRY'] == co]
return ['Country: {}<br>Total COVID Cases / 100,000 people: {}<br>Total Vaccinations / 100 people: {}<br>'.format(x1, x2, x3)
for (x1, x2, x3) in zip(use['COUNTRY'],
use['Cases - cumulative total per 100000 population'],
use['TOTAL_VACCINATIONS_PER100'])
]
# Functions to help with the vaccine marker size
def getVaccineMarkersize1():
return np.nan_to_num(np.clip(dfJoinedCOVID['Cases - cumulative total per 100000 population']/1000., 5, 100))
def getVaccineMarkersize2():
return np.clip(dfJoinedCOVID['TOTAL_VACCINATIONS_PER100']/7., 5, 100)
def getVaccineMarkersize1H(co):
use = dfJoinedCOVID.loc[dfJoinedCOVID['COUNTRY'] == co]
return np.nan_to_num(np.clip(use['Cases - cumulative total per 100000 population']/1000., 5, 100))
def getVaccineMarkersize2H(co):
use = dfJoinedCOVID.loc[dfJoinedCOVID['COUNTRY'] == co]
return np.clip(use['TOTAL_VACCINATIONS_PER100']/7., 5, 100)
# Copying these fits here so that it's all in one place
slope1, intercept1, r1, p1, se1 = scipy.stats.linregress(dfFit1['TOTAL_VACCINATIONS_PER100'], np.log10(dfFit1['2020']))
xFit1 = np.linspace(0, 300, 100)
yFit1 = 10.**(slope1*xFit1 + intercept1)
dfFit2 = dfJoinedCOVID.dropna(subset = ['START_DATE', '2020'])
delta = (dfFit2['START_DATE'] - dfFit2['START_DATE'].min())/np.timedelta64(1,'D')
slope2, intercept2, r2, p2, se2 = scipy.stats.linregress(delta, np.log10(dfFit2['2020']))
xx2 = np.linspace(0, 500, 100)
yFit2 = 10.**(slope2*xx2 + intercept2)
xFit2 = xx2*np.timedelta64(1,'D') + dfFit2['START_DATE'].min()
# +
# I am going to create the dropdown list here and then add it to the figure below
# I will need to update the x and y data for the time series plot
# AND also update the text for the tooltips for the red circles in the bottom panel
# AND also update the marker size for the red circles in the bottom panel
# Even though some data will not change, I will need to specify everything in this dropdown menu
# Identify the countries to use
# I will but The United States of America first so that it can be the default country on load (the first button)
availableCountries = dfC['Country'].unique().tolist()
availableCountries.insert(0, availableCountries.pop(availableCountries.index('United States of America')))
# Create the dropdown buttons
dropdown = []
for c in availableCountries:
if (c in dfJoinedCOVID['COUNTRY'].tolist()):
dropdown.append(dict(
args = [{'x': [updateTimePlotX(c), updateTimePlotX(c), updateTimePlotX(c), updateTimePlotX(c), # time plot
dfJoinedCOVID['TOTAL_VACCINATIONS_PER100'], # full scatter plot on the left
xFit1, # fit line 1
updateVaccinePlotX1(c), # red circle in left scatter plot
dfJoinedCOVID['START_DATE'], # full scatter plot on the right
xFit2, # fit line 2
updateVaccinePlotX2(c) # red circle on right scatter plot
],
'y': [updateTimePlotY1(c), updateTimePlotY2(c), updateTimePlotY3(c), updateTimePlotY4(c),
dfJoinedCOVID['2020'],
yFit1,
updateVaccinePlotY(c),
dfJoinedCOVID['2020'],
yFit2,
updateVaccinePlotY(c)
],
'text': ['', '', '', '',
getVaccineTextAll(), '', updateVaccineHText(c),
getVaccineTextAll(), '', updateVaccineHText(c)
],
'marker.size': ['', '', '', '',
getVaccineMarkersize1(), '',getVaccineMarkersize1H(c),
getVaccineMarkersize2(), '',getVaccineMarkersize2H(c),
]
}],
label = c,
method = 'update'
))
# -
# A Function to add the dropdown menu and buttons
# Note: I've seen odd behavior with adding the dropdown first and then the buttons. (e.g., the dropdown turns into many buttons)
def addButtonsAndDropdown(fig):
fig.update_layout(
updatemenus = [
# Copy the buttons from above
dict(
type = 'buttons',
direction = 'left',
buttons = list([
dict(
args = [{'visible': [True, False, False, False, True, True, True, True, True, True]}],
label = 'Daily Cases',
method = 'restyle'
),
dict(
args = [{'visible': [False, True, False, False, True, True, True, True, True, True]}],
label = 'Daily Deaths',
method = 'restyle'
),
dict(
args = [{'visible': [False, False, True, False, True, True, True, True, True, True]}],
label = 'Cumulative Cases',
method = 'restyle'
),
dict(
args = [{'visible': [False, False, False, True, True, True, True, True, True, True]}],
label = 'Cumulative Deaths',
method = 'restyle'
)
]),
showactive = True,
x = 0.0,
xanchor = 'left',
y = 1.05,
yanchor = 'top'
),
# Add the dropdown
dict(
buttons = dropdown,
direction = 'down',
showactive = True,
x = 0.0,
xanchor = 'left',
y = 1.1,
yanchor = 'top'
),
]
)
return fig
# +
# I can't find a way to initialize the dropdown menu to anything other than the first in the list
# (I resorted the list to put the United States of America on top, before creating the dropdowns, so that is the default on load)
country = availableCountries[0]
# Use the function from above
fig = generateFigure(country)
# Now add in the buttons and dropdown menu
fig = addButtonsAndDropdown(fig)
# Move the title up a bit more, and remove the country from the name
fig.update_layout(title_text = 'COVID-19 Data Explorer <br>(' + str(rollingAve) +'-day rolling average)',
title_y = 0.97,
margin = dict(t = 140)
)
fig.show()
# -
# You can save the plotly figure as an html file to use on your website.
fig.write_html('plotly_graph.html')
| introToPlotly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # IOOS QARTOD software (ioos_qc)
#
# This post will demonstrate how to [run ``ioos_qc``](https://github.com/ioos/ioos_qc) on a time-series dataset. ``ioos_qc`` implements the [Quality Assurance / Quality Control of Real Time Oceanographic Data (QARTOD)](https://ioos.noaa.gov/project/qartod/).
#
# We will [use `bokeh`](https://docs.bokeh.org/en/latest/) for interactive plots, so let's start by loading the interactive notebook output.
from bokeh.plotting import output_notebook
output_notebook()
# We will be using the water level data from a [fixed station in Kotzebue, AK](https://www.google.com/maps?q=66.895035,-162.566752).
#
# Below we create a simple Quality Assurance/Quality Control (QA/QC) configuration that will be used as input for ``ioos_qc``. All the interval values are in the same units as the data.
#
# For more information on the tests and recommended values for QA/QC check the documentation of each test and its inputs:
# https://ioos.github.io/ioos_qc/api/ioos_qc.html#module-ioos_qc.qartod
# +
variable_name = "sea_surface_height_above_sea_level_geoid_mhhw"
qc_config = {
"qartod": {
"gross_range_test": {
"fail_span": [-10, 10],
"suspect_span": [-2, 3]
},
"flat_line_test": {
"tolerance": 0.001,
"suspect_threshold": 10800,
"fail_threshold": 21600
},
"spike_test": {
"suspect_threshold": 0.8,
"fail_threshold": 3,
}
}
}
# -
# Now we are ready to load the data, run tests and plot results!
#
# We will get the data from the [AOOS ERDDAP server](http://erddap.aoos.org/erddap/). Note that the data may change in the future. For reproducibility's sake we will save the data downloaded into a CSV file.
# +
from pathlib import Path
import pandas as pd
from erddapy import ERDDAP
path = Path().absolute()
fname = path.joinpath("data", "water_level_example.csv")
if fname.is_file():
data = pd.read_csv(fname, parse_dates=["time (UTC)"])
else:
e = ERDDAP(
server="http://erddap.aoos.org/erddap/",
protocol="tabledap"
)
e.dataset_id = "kotzebue-alaska-water-level"
e.constraints = {
"time>=": "2018-09-05T21:00:00Z",
"time<=": "2019-07-10T19:00:00Z",
}
e.variables = [
variable_name,
"time",
"z",
]
data = e.to_pandas(
index_col="time (UTC)",
parse_dates=True,
)
data["timestamp"] = data.index.astype("int64") // 1e9
data.to_csv(fname)
data.head()
# +
from ioos_qc.config import QcConfig
qc = QcConfig(qc_config)
qc_results = qc.run(
inp=data["sea_surface_height_above_sea_level_geoid_mhhw (m)"],
tinp=data["timestamp"],
zinp=data["z (m)"],
)
qc_results
# -
# The results are returned in a dictionary format, similar to the input configuration, with a mask for each test. While the mask **is** a masked array it should not be applied as such. The results range from 1 to 4 meaning:
#
# 1. data passed the QA/QC
# 2. did not run on this data point
# 3. flag as suspect
# 4. flag as failed
#
# Now we can write a plotting function that will read these results and flag the data.
# +
# %matplotlib inline
from datetime import datetime
import numpy as np
import matplotlib.pyplot as plt
def plot_results(data, var_name, results, title, test_name):
time = data["time (UTC)"]
obs = data[var_name]
qc_test = results["qartod"][test_name]
qc_pass = np.ma.masked_where(qc_test != 1, obs)
qc_suspect = np.ma.masked_where(qc_test != 3, obs)
qc_fail = np.ma.masked_where(qc_test != 4, obs)
qc_notrun = np.ma.masked_where(qc_test != 2, obs)
fig, ax = plt.subplots(figsize=(15, 3.75))
fig.set_title = f"{test_name}: {title}"
ax.set_xlabel("Time")
ax.set_ylabel("Observation Value")
kw = {"marker": "o", "linestyle": "none"}
ax.plot(time, obs, label="obs", color="#A6CEE3")
ax.plot(time, qc_notrun, markersize=2, label="qc not run", color="gray", alpha=0.2, **kw)
ax.plot(time, qc_pass, markersize=4, label="qc pass", color="green", alpha=0.5, **kw)
ax.plot(time, qc_suspect, markersize=4, label="qc suspect", color="orange", alpha=0.7, **kw)
ax.plot(time, qc_fail, markersize=6, label="qc fail", color="red", alpha=1.0, **kw)
ax.grid(True)
title = "Water Level [MHHW] [m] : Kotzebue, AK"
# -
# The gross range test test should fail data outside the $\pm$ 10 range and suspect data below -2, and greater than 3. As one can easily see all the major spikes are flagged as expected.
plot_results(
data,
"sea_surface_height_above_sea_level_geoid_mhhw (m)",
qc_results,
title,
"gross_range_test"
)
# An actual spike test, based on a data increase threshold, flags similar spikes to the gross range test but also indetifies other suspect unusual increases in the series.
plot_results(
data,
"sea_surface_height_above_sea_level_geoid_mhhw (m)",
qc_results,
title,
"spike_test"
)
# The flat line test identifies issues with the data where values are "stuck."
#
# `ioos_qc` succefully identified a huge portion of the data where that happens and flagged a smaller one as suspect. (Zoom in the red point to the left to see this one.)
plot_results(
data,
"sea_surface_height_above_sea_level_geoid_mhhw (m)",
qc_results,
title,
"flat_line_test"
)
# This notebook was adapt from <NAME> and <NAME>'s [original ioos_qc examples](https://github.com/ioos/ioos_qc/blob/b34b3762d659362fb3af11f52d8905d18cd6ec7b/docs/source/examples/QartodTestExample_WaterLevel.ipynb). Please [see the ``ioos_qc`` documentation](https://ioos.github.io/ioos_qc/) for more examples.
| jupyterbook/content/Code Gallery/data_analysis_and_visualization_notebooks/2020-02-14-QARTOD_ioos_qc_Water-Level-Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="SKRXKtDGS6iM"
import cv2
import numpy as np
from matplotlib import pyplot as plt
import glob
import os
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from numpy import linalg
# from google.colab import drive
# + colab={} colab_type="code" id="72YGbrkFS6iZ"
def mean(Xtrain):
return np.mean(Xtrain, axis = 1)
# + colab={} colab_type="code" id="-yLK4fUjS6in"
def X(Xtrain, mean):
Xdef = []
for col in Xtrain.T:
Xdef.append(col - mean)
Xdef = np.array(Xdef)
return Xdef.T
# + colab={} colab_type="code" id="DLWtgXsRS6it"
def covMat(X):
C1 = np.dot(X, X.T)
return C1
# + colab={} colab_type="code" id="DPjaD60SS6iy"
def eigenVectors(X, C):
eigVal1, eigVect1 = linalg.eig(C)
return eigVect1
# + colab={} colab_type="code" id="wHW-qkT_S6i3"
def eigenWeights(E, X):
weights = np.matmul(E.T, X)
return weights
# + colab={} colab_type="code" id="5Lt3U2JjS6i-"
def face_detection(image):
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
cascade_image = cv2.CascadeClassifier('FaceDetHaarXML/haarcascade_frontalface_alt2.xml')
faces = cascade_image.detectMultiScale(image, scaleFactor = 1.3, minNeighbors = 5)
if (len(faces) == 0):
return None, None
(x, y, l, b) = faces[0]
return image[y:y+b, x:x+l], faces[0]
def oval(img):
gray = cv2.cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
face_cascade = cv2.CascadeClassifier('FaceDetHaarXML/haarcascade_frontalface_alt.xml')
faces = face_cascade.detectMultiScale(gray, 1.3, 5)
face_x =0
face_y =0
final_img = None
for (x,y,w,h) in faces:
face_x = x
face_y = y
roi_gray = gray[y:y+h, x:x+w]
roi_color =img[y:y+h, x:x+w]
mask = np.zeros_like(roi_color)
rows, cols,_ = mask.shape
mask=cv2.ellipse(mask,(int(rows/2), int(cols/2)), (250,300), angle=0, startAngle=0, endAngle=360, color=(255,255,255), thickness=-1)
oval = np.bitwise_and(roi_color,mask)
final_img=np.zeros_like(img)
final_img[y:y+h, x:x+w]= oval
final_img = cv2.cvtColor(final_img, cv2.COLOR_BGR2GRAY)
return final_img
# + colab={} colab_type="code" id="rhI-0VzyS6jE"
def loadDataLabels(data_path):
faces = []
labels = []
enc_labels = []
directories = os.listdir(data_path)
for i, direc in enumerate(directories):
sub_dir_path = data_path + "/" + direc
sub_dir_images = os.listdir(sub_dir_path)
label = direc
for j, image_name in enumerate(sub_dir_images):
image_path = sub_dir_path + "/" + image_name
img = cv2.imread(image_path)
# image = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
face_detected, shape = face_detection(img)
# face_detected = oval(img)
# face_detected = cv2.cvtColor(face_detected, cv2.COLOR_RGB2GRAY)
if face_detected is not None:
face_detected = cv2.resize(face_detected, (64, 64))
faces.append(face_detected)
labels.append(label)
labels = np.array(labels)
label_encoder = preprocessing.LabelEncoder()
enc_labels = label_encoder.fit_transform(labels)
labels_unique = label_encoder.classes_
faces = np.array(faces)
labels_unique = np.array(labels_unique)
enc_labels = np.array(enc_labels)
return faces, labels, labels_unique, enc_labels
# + colab={} colab_type="code" id="m7k6OMWkS6jP"
def weightMatrix(X_data):
print ('X_data shape: ', X_data.shape)
mean_ = mean(X_data)
print ('mean shape: ', mean_.shape)
X_ = X(X_data, mean_)
print ('X shape: ', X_.shape)
C = covMat(X_)
print ('C shape: ', C.shape)
E = eigenVectors(X_, C)
print ('E shape: ', E.shape)
weightMatrix = np.matmul(E.T, X_data)
print ('wtMat shape: ', weightMatrix.shape)
return E, weightMatrix
# + colab={} colab_type="code" id="3WbrYp1fS6ja"
def flattenData(X):
X_data = []
for image in X:
X_data.append(image.flatten())
X_data = np.array(X_data)
return X_data.T
# + colab={} colab_type="code" id="WS_HiMcFS6ji"
def PCAfamilyCoeff(X_train):
X_data = flattenData(X_train)
eigenVect, eigenCoeff = weightMatrix(X_data)
return eigenVect, eigenCoeff
def PCAfamilyEigen(X_data):
X_data = flattenData(X_data)
mean_ = mean(X_data)
print ('mean shape: ', mean_.shape)
X_ = X(X_data, mean_)
print ('X shape: ', X_.shape)
C = covMat(X_)
print ('C shape: ', C.shape)
E = eigenVectors(X_, C)
print ('E shape: ', E.shape)
return E
# + colab={} colab_type="code" id="7FXp1BcmS6jo"
def drawEigenFaces(eigenVect, index):
eigenFace = np.real(eigenVect)
eigenFace = ((eigenFace.T)[index]).reshape(64, 64)
print(eigenFace)
plt.imshow(eigenFace, cmap = 'jet')
plt.show()
# + colab={} colab_type="code" id="MvuHwJ1mS6ju"
def euclideanDistance(vector1, vector2):
return np.linalg.norm(vector1 - vector2)
# + colab={} colab_type="code" id="q8qEdsXtS6jz"
def leastEucDist(matrix, vector):
min_ind = 1000000
min_ed = 1000000
for i in range(((matrix.T).shape)[0]):
euc_dist = euclideanDistance(vector, (matrix.T)[i])
if(euc_dist < min_ed):
min_ed = euc_dist
min_ind = i
return min_ind
def leastEucDistN(matrix, vector, N):
min_ind = 1000000
min_ed = 1000000
l = []
for i in range(((matrix.T).shape)[0]):
euc_dist = euclideanDistance(vector, (matrix.T)[i])
l.append([euc_dist, i])
l = sorted(l)
min_ind_list = []
for i in range(N):
min_ind_list.append(l[i][1])
# print (min_ind_list)
return min_ind_list
# + colab={} colab_type="code" id="NESPO6TSS6j4"
def PCA_prediction(X_train, y_train, X_test, E):
# train_eigenVect, train_eigenCoeff = PCAfamilyCoeff(X_train)
# test_eigenVect, test_eigenCoeff = PCAfamilyCoeff(X_test)
eigenVect_train = E
X_train = flattenData(X_train)
X_test = flattenData(X_test)
print('X_train: ', X_train.shape)
train_eigenCoeff = eigenWeights(eigenVect_train, X_train)
print('Eigen Coeff train shape: ', train_eigenCoeff.shape)
test_eigenCoeff = eigenWeights(eigenVect_train, X_test)
print('Eigen Coeff test shape: ', test_eigenCoeff.shape)
predictions = []
for test_col in test_eigenCoeff.T:
predictions.append(y_train[leastEucDist(train_eigenCoeff, test_col)])
return predictions
def PCA_predictionN(X_train, y_train, X_test, E, N):
# train_eigenVect, train_eigenCoeff = PCAfamilyCoeff(X_train)
# test_eigenVect, test_eigenCoeff = PCAfamilyCoeff(X_test)
eigenVect_train = E
X_train = flattenData(X_train)
X_test = flattenData(X_test)
# print('X_train: ', X_train.shape)
train_eigenCoeff = eigenWeights(eigenVect_train, X_train)
# print('Eigen Coeff train shape: ', train_eigenCoeff.shape)
test_eigenCoeff = eigenWeights(eigenVect_train, X_test)
# print('Eigen Coeff test shape: ', test_eigenCoeff.shape)
predictions = []
for test_col in test_eigenCoeff.T:
l1 = leastEucDistN(train_eigenCoeff, test_col, N)
l2 = []
for i in range(len(l1)):
l2.append(y_train[l1[i]])
predictions.append(l2)
return predictions
# + colab={} colab_type="code" id="D2pjDNIQS6j9"
def PCA_accuracy(predictions, y_test):
count = 0
len_y_test = len(y_test)
if (len(predictions) != len(y_test)):
print("Error...Mismatch in Lenghts!")
len_y_test = len_y_test - 1
for i in range(len(predictions)):
if (predictions[i] == y_test[i]):
count += 1
print("PCA implementation Accuracy: ", (count/len_y_test)*100)
def PCA_accuracyN(predictions, y_test, N):
count = 0
len_y_test = len(y_test)
if (len(predictions) != len(y_test)):
print("Error...Mismatch in Lenghts!")
len_y_test = len_y_test - 1
for i in range(len(y_test)):
if ((y_test[i]) in predictions[i]):
count += 1
print("PCA implementation top-", N, "Accuracy: ", (count/len_y_test)*100)
# + colab={} colab_type="code" id="MdGxY8UJS6kF"
def Example(index, predictions, y_test, unique_labels):
plt.imshow(X_test[index], cmap = 'gray')
plt.show()
print("Prediction made by Model: ", unique_labels[predictions[index]])
print("Ground Truth: ", unique_labels[y_test[index]])
# + colab={} colab_type="code" id="FUlbGbNuS6kJ"
faces, labels, unique_labels, enc_labels = loadDataLabels("./Dataset")
(X_train, X_test, y_train, y_test) = train_test_split(faces, enc_labels, test_size = 0.25, random_state = 42)
# + colab={} colab_type="code" id="a_G4OkNHS6kV"
eigenVect = PCAfamilyEigen(X_train)
# -
drawEigenFaces(eigenVect, 6)
predictions = PCA_prediction(X_train, y_train, X_test, eigenVect)
# +
# print(predictions)
# print(y_test)
# -
PCA_accuracy(predictions, y_test)
Example(100, predictions, y_test, unique_labels)
predictions_top1 = PCA_predictionN(X_train, y_train, X_test, eigenVect, 1)
PCA_accuracyN(predictions_top1, y_test, 1)
predictions_top3 = PCA_predictionN(X_train, y_train, X_test, eigenVect, 3)
PCA_accuracyN(predictions_top3, y_test, 3)
predictions_top10 = PCA_predictionN(X_train, y_train, X_test, eigenVect, 10)
PCA_accuracyN(predictions_top10, y_test, 10)
| PCA_scratchFR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import h5py
import json
import sys
sys.path.append('F:/Linux')
sys.path.append("C:/Users/qq651/OneDrive/Codes/A2project/")
import illustris_python as il
import matplotlib.pyplot as plt
from plotTools.plot import *
il1_A2list = np.load('f:/Linux/localRUN/il1_A2dict(135-68_21part).npy', allow_pickle=1).item()
tng_A2list = np.load('f:/Linux/localRUN/tng_A2dict(99-33_21part).npy', allow_pickle=1).item()
tng_zformation = np.load('f:/Linux/localRUN/tng_zformation.npy', allow_pickle=1).item()
il1_zformation = np.load('f:/Linux/localRUN/il1_zformation.npy', allow_pickle=1).item()
il1_barID = np.load('f:/Linux/localRUN/barredID_il1.npy')
il1_diskID = np.load('f:/Linux/localRUN/diskID_il1.npy')
tng_barID = np.load('f:/Linux/localRUN/barredID_4WP_TNG.npy')
tng_diskID = np.load('f:/Linux/localRUN/diskID_4WP.npy')
'''
Rs il1 TNG
0.0: [135, 99],
0.1: [127, 91],
0.2: [120, 84],
0.3: [113, 78],
0.4: [108, 72],
0.5: [103, 67],
0.6: [99, 63],
0.7: [95, 59],
0.8: [92, 56],
0.9: [89, 53],
1.0: [85, 50],
1.1: [82, 47],
1.2: [80, 45],
1.3: [78, 43],
1.4: [76, 41],
1.5: [75, 40],
1.6: [73, 38],
1.7: [71, 36],
1.8: [70, 35],
1.9: [69, 34],
2.0: [68, 33],
2.5: [64, 29],
3.0: [60, 25]
'''
il1_snap = np.array([135, 127, 120, 113, 108, 103, 99, 95, 92, 89, 85, 82, 80, 78, 76, 75, 73, 71, 70, 69, 68])
tng_snap = np.array([99, 91, 84, 78, 72, 67, 63, 59, 56, 53, 50, 47, 45, 43, 41, 40, 38, 36, 35, 34, 33])
Redshift = np.array([0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 1.1, 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8, 1.9, 2.0])
il1_tbar = []
tng_tbar = []
il1_id = []
tng_id = []
for i in il1_A2list.keys():
il1_tbar.append(zbar(i, il1_A2list))
prog = LoadMergHist('il1', i)[0]
try:
il1_id.append(prog[68])
except KeyError:
continue
for i in tng_A2list.keys():
prog = LoadMergHist('TNG', i)[0]
try:
tng_id.append(prog[33])
tng_tbar.append(zbar(i, tng_A2list))
except KeyError:
continue
mas = il.func.loadSubhalos('TNG', 33, 'SubhaloMassInHalfRadType')
Gf = mas[:, 0] / (mas[:, 4] + mas[:, 0])
Gf[np.isnan(Gf)] = 0
tng_gf = Gf
mas = il.func.loadSubhalos('il1', 68, 'SubhaloMassInHalfRadType')
Gf = mas[:, 0] / (mas[:, 4] + mas[:, 0])
Gf[np.isnan(Gf)] = 0
il1_gf = Gf
il1_tbar = np.array(il1_tbar)
tng_tbar = np.array(tng_tbar)
# -
plt.scatter(tng_gf[tng_id], tng_tbar, s=4, label='TNG-100')
plt.scatter(il1_gf[il1_id], il1_tbar, s=5, color='r', label='Illustris-1')
plt.xlabel('Gas fraction at $z=2$', fontsize = 20)
plt.ylabel(r'$z_{bar}$', fontsize = 20)
plt.xlim(-0.02, 1)
plt.legend(fontsize = 13)
plt.savefig('F:/Linux/local_result/zbar/zbar.pdf')
tng_dots = np.vstack((tng_gf[tng_id], tng_tbar))
il1_dots = np.vstack((il1_gf[il1_id], il1_tbar))
bins = np.linspace(0, 1, 11)
tng_plotdata = [[], [], []]
for i in range(10):
mask = (tng_dots[0, :] >= bins[i]) & (tng_dots[0, :] < bins[i + 1])
tmp = tng_dots[1, :][mask]
d0, d1, d2 = ErrorBarMedian(tmp)
tng_plotdata[0].append(d0)
tng_plotdata[1].append(d1)
tng_plotdata[2].append(d2)
tng_plotdata = np.array(tng_plotdata)
tng_Err = np.vstack((tng_plotdata[1, :] - tng_plotdata[0, :], tng_plotdata[2, :] - tng_plotdata[1, :]))
il1_plotdata = [[], [], []]
for i in range(10):
mask = (il1_dots[0, :] >= bins[i]) & (il1_dots[0, :] < bins[i + 1])
tmp = il1_dots[1, :][mask]
d0, d1, d2 = ErrorBarMedian(tmp)
il1_plotdata[0].append(d0)
il1_plotdata[1].append(d1)
il1_plotdata[2].append(d2)
il1_plotdata = np.array(il1_plotdata)
il1_Err = np.vstack((il1_plotdata[1, :] - il1_plotdata[0, :], il1_plotdata[2, :] - il1_plotdata[1, :]))
plt.errorbar(bins[:-1] + 0.05, tng_plotdata[1, :], yerr = tng_Err, elinewidth=2, capthick=2, capsize=3, color='c', fmt='o', label='TNG-100')
plt.errorbar(bins[:-1] + 0.05, il1_plotdata[1, :], yerr=il1_Err, elinewidth=2, capthick=2, capsize=3, color='r', fmt='o', label='Illustris-1', alpha=0.75)
plt.xlim(-0.02, 1)
#plt.title('Statistics of zbar in each bin')
plt.xlabel('Gas fraction at $z=2$', fontsize = 20)
plt.ylabel(r'$z_{bar}$', fontsize = 20)
plt.xticks(bins[:-1])
plt.legend(fontsize = 13)
plt.savefig('F:/Linux/local_result/zbar/zbar_err.pdf')
tng_gf = {}
for snap in tng_snap:
mas = il.func.loadSubhalos('TNG', snap, 'SubhaloMassInHalfRadType')
Gf = mas[:, 0] / (mas[:, 4] + mas[:, 0])
Gf[np.isnan(Gf)] = 0
tng_gf[snap] = Gf
il1_gf = {}
for snap in il1_snap:
mas = il.func.loadSubhalos('il1', snap, 'SubhaloMassInHalfRadType')
Gf = mas[:, 0] / (mas[:, 4] + mas[:, 0])
Gf[np.isnan(Gf)] = 0
il1_gf[snap] = Gf
tng_plotdata = [[], []]
for haloID in tng_barID:
ind = np.where(Redshift == zbar(haloID, tng_A2list))[0][0]
snap = tng_snap[ind]
prog = LoadMergHist('TNG', haloID)[0]
try:
subID = prog[snap]
except KeyError:
continue
tng_plotdata[0].append(tng_gf[snap][subID])
tng_plotdata[1].append(zbar(haloID, tng_A2list))
il1_plotdata = [[], []]
for haloID in il1_barID:
ind = np.where(Redshift == zbar(haloID, il1_A2list))[0][0]
snap = il1_snap[ind]
prog = LoadMergHist('il1', haloID)[0]
try:
subID = prog[snap]
except KeyError:
continue
il1_plotdata[0].append(il1_gf[snap][subID])
il1_plotdata[1].append(zbar(haloID, il1_A2list))
plt.scatter(tng_plotdata[0], tng_plotdata[1], s=4, label='TNG-100')
plt.scatter(il1_plotdata[0], il1_plotdata[1], s=6, color='r', label='Illustris-1')
plt.xlabel(r'Gas fraction at $z_{bar}$', fontsize = 20)
plt.ylabel(r'$z_{bar}$', fontsize = 20)
plt.xlim(-0.02, 1.0)
plt.ylim(-0.1, 2.1)
plt.legend(fontsize = 13)
plt.savefig('F:/Linux/local_result/zbar/zBarFormed.pdf')
| JpytrNb/pdfPlot/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import urllib.request
import urllib.parse
import urllib.error
from bs4 import BeautifulSoup
import ssl
import json
import ast
import json
import os
from urllib.request import Request, urlopen
#importing library
import requests
import csv
import re
import pandas as pd
# +
##Getting channel Title
def channelTitle(content):
title = content.h3.a.text
return title
youtubeIDs=[]
youtubeChannel=[]
videoCate=[]
def mainFunction(searchValue,channel=1):
youtubeUrl="https://www.youtube.com/results?search_query="
page = "&page="
count=1
pages = 1
searchQuery=searchValue
for category in searchQuery:
count=1
while count <= pages:
scrapeURL = youtubeUrl + str(category) + page + str(count)
print(category)
source = requests.get(scrapeURL).text
soup = BeautifulSoup(source, 'lxml')
#getting the div yt-lockup-content
for content in soup.find_all('div', class_= "yt-lockup-content"):
try:
ID=content.h3.a
matching=bool('/watch' in ID.get('href'))
if(matching):
youtubeIDs.append(ID.get('href'))
videoCate.append(category)
else:
if(channel):
youtubeChannel.append(channelTitle(content))
except Exception as e:
print(e)
print("Exception")
description = None
#increasing the count
count=count+1
searchValue=['FordvFerrari']
mainFunction(searchValue)
#Getting video of youtubeChannel
mainFunction(youtubeChannel,channel=0)
df = {'Videourl': youtubeIDs,'Category':videoCate}
df2=pd.DataFrame(df)
#storing Youtube videos link into csv file
df2.to_csv("Videourl.csv",index=False)
link = (df2['Videourl'][1])
# +
ctx = ssl.create_default_context()
ctx.check_hostname = False
ctx.verify_mode = ssl.CERT_NONE
# Input from user
url = ("https://www.youtube.com.{}" .format(link))
import webbrowser
webbrowser.open(url)
# -
| Youtube.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sudoggie/sas-visualanalytics-thirdpartyvisualizations/blob/master/Perceptron_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="zGrQyZxaxzW-"
# # Perceptron 1
# + [markdown] id="5hNUHxiAx2XE"
# 
# + [markdown] id="0Zy1NxIgOCnX"
# ## Inputs and weights
# + id="z-5dEcggSO_S"
inputs = [35, 25]
# + id="iTpWAQDJSf-e" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d5c98858-598e-43e7-820e-5fb8d0e1935d"
type(inputs)
# + id="FaQu-mJgSo6I" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4af4a6e7-da17-4058-fff8-c55586971838"
inputs[0]
# + id="ZYmiXB3ESrXO" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f3a547f4-d136-4924-ed70-54e151f06a64"
inputs[1]
# + id="sUegRBmjSxU_"
weights = [-0.8, 0.1]
# + [markdown] id="b-FFtbMFOHPO"
# ## Sum function
# + id="1ZFnRkgWS6tF"
def sum(inputs, weights):
s = 0
for i in range(2):
#print(i)
#print(inputs[i])
#print(weights[i])
s += inputs[i] * weights[i]
return s
# + id="-nCHELAjTQw1"
s = sum(inputs, weights)
# + id="Yp7Eo5o5UC9n" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6dcbc477-7dd3-40ec-9342-7fa505f3b6b6"
s
# + [markdown] id="HnggNLkiOJj2"
# ## Step function
# + id="tyGCL1jYUS52"
def step_function(sum):
if (sum >= 1):
return 1
return 0
# + [markdown] id="pybCG4h8ONSR"
# ## Final result
# + id="nISd_PZSUzvI" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="56302a9b-2d9f-4f6a-b5d5-273072b71cd2"
step_function(s)
| Perceptron_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Tentativa 1
# Render do problema do taxi
import gym
env = gym.make("Taxi-v3").env
env.render()
# # Tentativa 2
# Teste aplicando as funções predeterminadas da lib para rodar. rodando os valores dentro do notebook, no cria a janela com o modelo e não é possível acompanahar a animação
import gym
env = gym.make('Acrobot-v1')
env.reset()
for _ in range(500):
env.render()
env.step(env.action_space.sample())
def show_state(env, step=0, info=""):
plt.figure(3)
plt.clf()
plt.imshow(env.render(mode='rgb_array'))
plt.title("%s | Step: %d %s" % (env._spec.id,step, info))
plt.axis('off')
display.clear_output(wait=True)
display.display(plt.gcf())
from gym import envs
all_envs = envs.registry.all()
env_ids = [env_spec.id for env_spec in all_envs]
print(env_ids)
# # Tentativa 3
# Salvando como video e criando uma janela para rodar
# !apt-get install ffmpeg
# !pip install imageio-ffmpeg
# +
import gym
from gym import wrappers
env = gym.make('BipedalWalker-v3')
env = wrappers.Monitor(env, "./gym-results", force=True)
env.reset()
for _ in range(1000):
action = env.action_space.sample()
observation, reward, done, info = env.step(action)
if done: break
env.close()
# +
import io
import base64
from IPython.display import HTML
video = io.open('./gym-results/openaigym.video.%s.video000000.mp4' % env.file_infix, 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''
<video width="360" height="auto" alt="test" controls><source src="data:video/mp4;base64,{0}" type="video/mp4" /></video>'''
.format(encoded.decode('ascii')))
# -
# # Tentativa 4
#
# Aplicando visualização com o display do ipython. demora bastante em renderizar o resultado.
import gym
import numpy as np
import matplotlib.pyplot as plt
from IPython import display as ipythondisplay
# +
env = gym.make('MountainCar-v0')
env.reset()
prev_screen = env.render(mode='rgb_array')
plt.imshow(prev_screen)
for _ in range(50):
action = env.action_space.sample()
obs, reward, done, info = env.step(action)
screen = env.render(mode='rgb_array')
plt.imshow(screen)
ipythondisplay.clear_output(wait=True)
ipythondisplay.display(plt.gcf())
if done:
break
ipythondisplay.clear_output(wait=True)
env.close()
# -
# # Tentativa 5
# Método otimizado para rodar. Atualização da imagem.
# +
import gym
from IPython import display as ipythondisplay
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
env = gym.make('MountainCar-v0')
env.reset()
img = plt.imshow(env.render(mode='rgb_array')) # only call this once
for _ in range(100):
img.set_data(env.render(mode='rgb_array')) # just update the data
ipythondisplay.display(plt.gcf())
ipythondisplay.clear_output(wait=True)
action = env.action_space.sample()
env.step(action)
ipythondisplay.clear_output(wait=True)
env.close()
# -
# !apt-get install ffmpeg
# !apt-get install libav-tools
# +
import gym
from IPython import display
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
env = gym.make('MountainCar-v0')
env.reset()
plt.figure(figsize=(9,9))
img = plt.imshow(env.render(mode='rgb_array')) # only call this once
for _ in range(500):
img.set_data(env.render(mode='rgb_array')) # just update the data
display.display(plt.gcf())
display.clear_output(wait=True)
action = env.action_space.sample()
env.step(action)
env.close()
| notebooks/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Allow us to load `open_cp` without installing
import sys, os.path
sys.path.insert(0, os.path.abspath(os.path.join("..", "..")))
# # Uncensored chicago data
#
# After asking around the research community, it appears that the chicago dataset has been re-geocoded sometime in the last year (or so). In the past, each event was geocoded to a fairly exact address (perhaps building entrance).
#
# A little searching can find copies of extracts of this data. As there must have been a good reason for changing the geocoding (presumably privacy based) we will not release a copy of this data, and we will be careful when drawing maps etc. not to show more detail than can already by found in the literature.
#
# However, it is interesting to compare the data, and gain some insight into how the new geocoding is performed.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import geopandas as gpd
import pandas as pd
import numpy as np
import open_cp.sources.chicago as chicago
import os, lzma
filename = os.path.join("..", "..", "..", "..", "Data", "two.csv")
file_all = os.path.join("..", "..", "open_cp", "sources", "chicago_all.csv.xz")
# -
# ## Load the uncensored data
#
# - As usual, we'll drop the "HOMICIDE" crimes.
# - The case number "HV612588" is reported twice, and one looks like an error. We'll drop in
# Load the uncensored data
data = chicago.load_to_geoDataFrame(filename, type="all_other")
data = data[data.case != "HV612588"]
data = data[data.crime != "HOMICIDE"]
len(data)
# So every row has a unique case
assert len(data.case.unique()) == len(data)
data = data.set_index("case")
# ## Load matching "new" data
#
# We load the same case numbers from the dataset (downloaded in May 2017), perform a little manual cleaning, and check that we then have a unique set of case numbers.
#
# We are now mising about 1000 cases from the "old" data.
# +
cases = set(data.index)
with lzma.open(file_all, "rt") as file:
data_all = [event for event in chicago.generate_GeoJSON_Features(file, type="all")
if event["properties"]["case"] in cases]
data_all = gpd.GeoDataFrame.from_features(data_all)
# -
len(data_all), len(data_all.case.unique())
data_all = data_all[(data_all.case != "") |
((data_all.case == "") & (data_all.crime == "BURGLARY"))]
data_all = data_all[(data_all.case != "HT155546") |
((data_all.case == "HT155546") & (data_all.location == "TAVERN/LIQUOR STORE"))]
len(data_all), len(data_all.case.unique())
data_all = data_all.set_index("case")
len(set(data.index) - set(data_all.index)), len(set(data_all.index) - set(data.index))
# ## Check the time range
#
# We now look at the current file, and find all events in the same time range. There are about 1000 more; so very good agreement overall.
times = data.timestamp.map(chicago._date_from_iso)
start, end = min(times), max(times)
start, end
with lzma.open(file_all, "rt") as file:
data_all_in_time = []
for event in chicago.generate_GeoJSON_Features(file, type="all"):
ts = chicago._date_from_iso(event["properties"]["timestamp"])
if ts >= start and ts <= end:
data_all_in_time.append(event)
data_all_in_time = gpd.GeoDataFrame.from_features(data_all_in_time)
len(set(data_all_in_time.case))
data_all_in_time = None
# # Check for any differences, except in geometry
#
# Turns out there are quite a few differences.
#
# - A _lot_ of differences in "crime", and a huge number in "type". I have not looked closely, but from a brief look, it seems that the differences are mostly down to reasonable re-classification.
both = pd.merge(data, data_all, left_index=True, right_index=True)
print("'address' differences", len(both[~(both.address_x == both.address_y)]))
print("'crime' differences", len(both[~(both.crime_x == both.crime_y)]))
print("'type' differences", len(both[~(both.type_x == both.type_y)]))
print("'location' differences", len(both[~(both.location_x == both.location_y)]))
print("'timestamp' differences", len(both[~(both.timestamp_x == both.timestamp_y)]))
# # Compute difference in distance between the points
#
# In lon/lat space, which is areal, but not unreasonable on a small scale like this.
#
# We then visualise, with a log scale. So almost all points are between 10e-3 and 10e-4 distance apart. I.e. most move a bit, but not too much.
dists = []
for _, row in both.iterrows():
if row.geometry_x is not None and row.geometry_y is not None:
dists.append(row.geometry_x.distance(row.geometry_y))
else:
dists.append(-1)
both["distance"] = dists
# +
x = np.linspace(2, 5, 100)
xx = np.exp(-x * np.log(10))
y = [len(both[both["distance"] > t]) for t in xx]
fig, ax = plt.subplots()
ax.scatter(x,y)
# -
# ## Assemble the points into a geoDataFrame
# +
tmp = both.ix[both[both["distance"] < 0.001].index]
compare = gpd.GeoDataFrame()
compare["geo1"] = tmp["geometry_x"]
compare["geo2"] = tmp["geometry_y"]
compare["name1"] = tmp["address_x"]
compare["name2"] = tmp["address_y"]
compare.set_index(tmp.index)
tmp = None
compare = compare[compare.geo1.map(lambda pt : pt is not None)]
compare = compare[compare.geo2.map(lambda pt : pt is not None)]
# +
import shapely.geometry
compare["geometry"] = [ shapely.geometry.LineString([ row.geo1, row.geo2 ])
for _, row in compare.iterrows() ]
# -
compare = compare.set_geometry("geometry")
compare.crs = {"init": "epsg:4326"}
import os
tmp = compare.drop(["geo1", "geo2"], axis=1)
tmp.to_file( os.path.join("..", "..", "..", "..", "Data", "Old_Chicago_SHP") )
tmp = None
bds = compare.total_bounds
lines = []
for geo in compare.geometry:
xs, ys = geo.xy
lines.append(list(zip(xs, ys)))
# +
from matplotlib.collections import LineCollection
lc = LineCollection(lines, linewidths=0.1)
fig, ax = plt.subplots(figsize=(16,16))
ax.add_collection(lc)
ax.set(xlim=[bds[0], bds[2]], ylim=[bds[1], bds[3]])
None
# -
# # Look at one block
#
# We'll pick some blocks at random (without giving their names, to protect privacy) and plot the difference in the geocoding as lines joining the old and new points.
#
# What we see is that:
# - Each point appears to be projected (orthogonally) to a line. (Later we'll see that this line comes from the TIGER/Line dataset from the US Census bureau.)
# - Then the points are squashed together.
# - This has the effect of preserving the ordering and relative position along the block
# - As you can see the last two plots, a single block might be split into more than one "edge", and this procedure appears to be applied to each "edge" independently.
#
# We hence very clearly see how the pattern observed in the new/current dataset is generated: we will naturally end up with clumps of points in, roughly, the middle of the centre of the road of each block.
# +
fig, axes = plt.subplots(ncols=5, figsize=(18,10))
blocks = np.random.randint(low=0, high=10000, size=5)
blocks = [3243, 7016, 6951, 5475, 804]
for i, ax in zip(blocks, axes):
block = compare.ix[i].name1
compare[compare.name1 == block].plot(ax=ax, color="black")
ax.set_aspect(1)
ax.axes.xaxis.set_visible(False)
ax.axes.yaxis.set_visible(False)
# -
# ## Look at the TIGER data
#
# The [TIGER/Line](https://www.census.gov/geo/maps-data/data/tiger-line.html) dataset from the US Census bureau provides a great resource for getting block-level address data. See the other notebook for more details.
filename = os.path.join("/media", "disk", "TIGER Data", "tl_2013_17031_edges", "tl_2013_17031_edges.shp")
edges = gpd.read_file(filename)
want = {"geometry", "FULLNAME", "LFROMADD", "LTOADD", "RFROMADD", "RTOADD"}
edges = gpd.GeoDataFrame({key:edges[key] for key in want})
edges.crs={'init': 'epsg:4269'}
# +
import rtree
gap = 0.001
def gen():
for i, row in edges.iterrows():
bds = list(row.geometry.bounds)
bds = [bds[0]-gap, bds[1]-gap, bds[2]+gap, bds[3]+gap]
yield i, bds, None
idx = rtree.index.Index(gen())
# -
def find_match_via_rtree(point):
possibles = edges.ix[list(idx.intersection(point.coords[0]))]
if len(possibles) == 0:
#raise ValueError("Found no candidates for {}".format(point))
from collections import namedtuple
Error = namedtuple("Error", ["name"])
return Error(name=-1)
i = possibles.geometry.distance(point).argmin()
return edges.ix[i]
# +
block = compare.ix[804].name1
block = compare[compare.name1 == block]
rows = set( find_match_via_rtree(row.geometry).name for _, row in block.iterrows() )
rows = edges.ix[rows]
rows = rows[~rows.FULLNAME.map(lambda x : x is None or x == "Alley")]
fig, ax = plt.subplots(figsize=(16,10))
rows.plot(ax=ax)
block.plot(color="black", ax=ax)
xcs, ycs = [], []
for geo in rows.geometry:
x, y = geo.coords[0]
xcs.append(x)
ycs.append(y)
x, y = geo.coords[-1]
xcs.append(x)
ycs.append(y)
ax.scatter(xcs, ycs, color="blue", marker="o")
ax.set_aspect(1)
ax.axes.xaxis.set_visible(False)
ax.axes.yaxis.set_visible(False)
# -
# I have plotted here:
#
# - Edge distinct edge from the TIGER dataset, in a different color.
# - The start and end node of each edge, for easier visualistion.
#
# What I see here is no particularly clear pattern.
#
# - Visually, if we ignore the edge plot, then there seem to be two distinct "blocks" with points being mapped to one or the other
# - But there are 5 edges.
#
# If I cheat, and look with a base map, then in this case there are actually two blocks. But this hard to recognise automatically from the edge data.
| examples/Chicago/Old Chicago Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import sys
cwd = os.getcwd()
qiskit_dir = os.path.abspath(os.path.dirname(os.path.dirname(os.path.dirname(cwd))))
sys.path.append(qiskit_dir)
from qiskit import LegacySimulators, QuantumRegister, ClassicalRegister, QuantumCircuit, execute, compile
from qiskit.tools.parallel import parallel_map
from qiskit.tools.monitor import job_monitor
from qiskit.tools.events import TextProgressBar
from qiskit.tools.jupyter import *
sim_backend = LegacySimulators.get_backend('qasm_simulator')
import time
def func(_):
time.sleep(0.1)
return 0
# ## Test HTMLProgressBar
HTMLProgressBar()
parallel_map(func, list(range(10)));
# ## Test magic with HTMLProgessBar
# %%qiskit_progress_bar
parallel_map(func, list(range(10)));
# ## Test TextProgressBar
TextProgressBar()
parallel_map(func, list(range(10)));
# ## Test magic with TextProgessBar
# %%qiskit_progress_bar -t text
parallel_map(func, list(range(10)));
# ## Check compiler spawns progress bar
# +
q = QuantumRegister(2)
c = ClassicalRegister(2)
qc = QuantumCircuit(q, c)
qc.h(q[0])
qc.cx(q[0], q[1])
qc.measure(q, c)
HTMLProgressBar()
qobj = compile([qc]*20, backend=sim_backend)
# -
# ## Test job status magic
# +
# %%qiskit_job_status
q = QuantumRegister(2)
c = ClassicalRegister(2)
qc = QuantumCircuit(q, c)
qc.h(q[0])
qc.cx(q[0], q[1])
qc.measure(q, c)
job_sim = execute([qc]*10, backend=sim_backend)
# -
# ## Test job_monitor
job_sim2 = execute([qc]*10, backend=sim_backend)
job_monitor(job_sim2, monitor_async=False)
job_sim3 = execute([qc]*10, backend=sim_backend)
job_monitor(job_sim3)
| test/python/notebooks/test_pbar_status.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as stats
pd.set_option("display.max_columns", None)
# # Week 3
#
# I hope you're getting the hang of things. Today we're going on with the prinicples of data visualization!
# ## Overview
#
# Once again, the lecture has three parts:
#
# * First you will watch a video on visualization and solve a couple of exercises.
# * After that, we'll be reading about *scientific data visualization*, and the huge number of things you can do with just one variable. Naturally, we'll be answering questions about that book.
# * And finally reproducing some of the plots from that book.
# ## Part 1: Fundamentals of data visualization
# Last week we had a small introduction of data visualization. Today, we are going to be a bit more specific on data analysis and visualization. Digging a bit more into the theory with the next video.
#
# <mark>*It's important to highlight that these lectures are quite important. We don't have a formal book on data visualization. So the only source of knowledge about the **principles**, **theories**, and **ideas**, that are the foundation for good data viz, comes from the videos*. So watch them 🤓 </mark>
#
# [](https://www.youtube.com/watch?v=yiU56codNlI)
# > *Excercise 1.1:* Questions for the lecture
# > * As mentioned earlier, visualization is not the only way to test for correlation. We can (for example) calculate the Pearson correlation. Explain in your own words how the Pearson correlation works and write down it's mathematical formulation. Can you think of an example where it fails (and visualization works)?
#
# Pearson doesn't detect non-linear relationships, and also does not convey steepness of the slope (only the direction).
#
# > * What is the difference between a bar-chart and a histogram?
#
# Histogram has value intervals on the x-axis (and counts on y-axis), essentially displaying some distribution of data. Bar chart usually shows a specific value for a given entry, category etc.
#
# > * I mention in the video that it's important to choose the right bin-size in histograms. But how do you do that? Do a Google search to find a criterion you like and explain it.
#
# Experimental is one approach. More defined methods include Freedman-Diaconis and Sturges rule.
# http://www.jtrive.com/determining-histogram-bin-width-using-the-freedman-diaconis-rule.html
def freedman_diaconis(data, returnas="width"):
"""
Use Freedman Diaconis rule to compute optimal histogram bin width.
``returnas`` can be one of "width" or "bins", indicating whether
the bin width or number of bins should be returned respectively.
Parameters
----------
data: np.ndarray
One-dimensional array.
returnas: {"width", "bins"}
If "width", return the estimated width for each histogram bin.
If "bins", return the number of bins suggested by rule.
"""
data = np.asarray(data, dtype=np.float_)
IQR = stats.iqr(data, rng=(25, 75), scale="raw", nan_policy="omit")
N = data.size
bw = (2 * IQR) / np.power(N, 1/3)
if returnas=="width":
result = bw
else:
datmin, datmax = data.min(), data.max()
datrng = datmax - datmin
result = int((datrng / bw) + 1)
return result
# Ok, now that we've talked a bit about correlation and distributions, we are going to compute/visualize them while also testing some hypotheses along the way. Until now, we have analysed data at an explorative level, but we can use statistics to verify whether relationships between variables are significant. We'll do this in the following exercise.
#
# > *Exercise 1.2:* Hypothesis testing. We will look into correlations between number of steps and BMI, and differences between two data samples (Females vs Males). Follow the steps below for success:
# >
# > * First, we need to get some data. Download and read the data from the Female group [here](https://raw.githubusercontent.com/suneman/socialdata2022/main/files/data9b_f.csv) and the one from the Male group [here](https://raw.githubusercontent.com/suneman/socialdata2022/main/files/data9b_m.csv).
# > * Next, we are going to verify the following hypotheses:
# > 1. <mark>*H1: there is a statistically significant difference in the average number of steps taken by men and women*</mark>. Is there a statistically significant difference between the two groups? What is the difference between their mean number of steps? Plot two histograms to visualize the step-count distributions, and use the criterion you chose in Ex.1.1 to define the right bin-size.
# **Hint** you can use the function `ttest_ind()` from the `stats` package to test the hypothesis and consider a significance level $\alpha=0.05$.
# > 2. <mark>*H2: there is a negative correlation between the number of steps and the BMI for women*.</mark> We will use Pearson's correlation here. Is there a negative correlation? How big is it?
# > 3. <mark>*H3: there is a positive correlation between the number of steps and the BMI for men*.</mark> Is there a positive correlation? Compare it with the one you found for women.
# > * We have now gathered the results. Can you find a possible explanation for what you observed? You don't need to come up with a grand theory about mobility and gender, just try to find something (e.g. theory, news, papers, further analysis etc.) to support your conclusions and write down a couple of sentences.
#
# +
females = pd.read_csv('https://raw.githubusercontent.com/suneman/socialdata2022/main/files/data9b_f.csv')
males = pd.read_csv('https://raw.githubusercontent.com/suneman/socialdata2022/main/files/data9b_m.csv')
plt.subplot(1, 2, 1)
plt.hist(females.steps, bins=freedman_diaconis(females.steps, 'bins'))
plt.title('Females')
plt.subplot(1, 2, 2)
plt.title('Males')
plt.hist(males.steps, bins=freedman_diaconis(males.steps, 'bins'))
plt.tight_layout()
# -
stats.ttest_ind(females.steps, males.steps)
print(stats.pearsonr(females.steps, females.bmi))
print(stats.pearsonr(males.steps, males.bmi))
# > *Exercise 1.3:* scatter plots. We're now going to fully visualize the data from the previous exercise.
# >
# > * Create a scatter plot with both data samples. Use `color='#f6756d'` for one <font color=#f6756d>sample</font> and `color='#10bdc3'` for the other <font color=#10bdc3>sample</font>. The data is in front of you, what do you observe? Take a minute to think about these exercises: what do you think the point is?
# * After answering the questions above, have a look at this [paper](https://genomebiology.biomedcentral.com/track/pdf/10.1186/s13059-020-02133-w.pdf) (in particular, read the *Not all who wander are lost* section).
# > * The scatter plot made me think of another point we often overlook: *color-vision impairments*. When visualizing and explaining data, we need to think about our audience:
# > * We used the same colors as in the paper, try to save the figure and use any color-blindness simulator you find on the web ([this](https://www.color-blindness.com/coblis-color-blindness-simulator/) was the first that came out in my browser). Are the colors used problematic? Explain why, and try different types of colors. If you are interested in knowing more you can read this [paper](https://www.tandfonline.com/doi/pdf/10.1179/000870403235002042?casa_token=<KEY> <KEY>).
# > * But, are colors the only option we have? Find an alternative to colors, explain it, and change your scatter plot accordingly.
plt.scatter(females.steps, females.bmi, color='#f6756d')
plt.scatter(males.steps, males.bmi, color='#10bdc3')
# ## Part 2: Reading about the theory of visualization
#
# Since we can go deeper with the visualization this year, we are going to read the first couple of chapters from [*Data Analysis with Open Source Tools*](http://shop.oreilly.com/product/9780596802363.do) (DAOST). It's pretty old, but I think it's a fantastic resource and one that is pretty much as relevant now as it was back then. The author is a physicist (like Sune) so he likes the way he thinks. And the books takes the reader all the way from visualization, through modeling to computational mining. Anywho - it's a great book and well worth reading in its entirety.
#
# As part of this class we'll be reading the first chapters. Today, we'll read chaper 2 (the first 28 pages) which supports and deepens many of the points we made during the video above.
#
# To find the text, you will need to go to **DTU Learn**. It's under "Course content" $\rightarrow$ "Content" $\rightarrow$ "Lecture 3 reading".
# > *Excercise 2*: Questions for DAOST
# > * Explain in your own words the point of the jitter plot.
# > ** To identify multiple dots with same x/y values in a dot/scatter plot
# > * Explain in your own words the point of figure 2-3. (I'm going to skip saying "in your own words" going forward, but I hope you get the point; I expect all answers to be in your own words).
# > ** The two histograms suggest two different distributions - uniform vs normal
# > * The author of DAOST (<NAME>) likes KDEs (and think they're better than histograms). And we don't. Sune didn't give a detailed explanation in the video, but now that works to our advantage. We'll ask you to think about this and thereby create an excellent exercise: When can KDEs be misleading?
# > ** If the bandwidth is quite high, the smoothing can hide certain patterns. While KDE is more visually appealing, the reader might just try to unwrap the KDE into a regular histogram when interpreting it.
# > * Sune discussed some strengths of the CDF - there are also weaknesses. Janert writes "CDFs have less intuitive appeal than histograms or KDEs". What does he mean by that?
# > ** Histograms and KDEs allow very quick interpretation for the human eye i guess?
# > * What is a *Quantile plot*? What is it good for.
# > ** Detecting distributions
# > * How is a *Probablity plot* defined? What is it useful for? Have you ever seen one before?
# > * One of the reasons we like DAOST is that Janert is so suspicious of mean, median, and related summary statistics. Explain why one has to be careful when using those - and why visualization of the full data is always better.
# > * Sune loves box plots (but not enough to own one of [these](https://twitter.com/statisticiann/status/1387454947143426049) 😂). When are box plots most useful?
# > * The book doesn't mention [violin plots](https://en.wikipedia.org/wiki/Violin_plot). Are those better or worse than box plots? Why?
# ## Part 3: *Finally*! Let's create some visualizations
# > *Excercise 3.1*: Connecting the dots and recreating plots from DAOST but using our own favorite dataset.
# > * Let's make a jitter-plot (that is, code up something like **Figure 2-1** from DAOST from scratch), but based on *SF Police data*. My hunch from inspecting the file is that the police-folks might be a little bit lazy in noting down the **exact** time down to the second. So choose a crime-type and a suitable time interval (somewhere between a month and 6 months depending on the crime-type) and create a jitter plot of the arrest times during a single hour (like 13-14, for example). So let time run on the $x$-axis and create vertical jitter.
# > * Last time, we did lots of bar-plots. Today, we'll play around with histograms (creating two crime-data based versions of the plot-type shown in DAOST **Figure 2-2**). I think the GPS data could be fun to see this way.
# > * This time, pick two crime-types with different geographical patterns **and** a suitable time-interval for each (you want between 1000 and 10000 points in your histogram)
# > * Then take the latitude part of the GPS coordinates for each crime and bin the latitudes so that you have around 50 bins across the city of SF. You can use your favorite method for binning. I like `numpy.histogram`. This function gives you the counts and then you do your own plotting.
# > * Next up is using the plot-type shown in **Figure 2-4** from DAOST, but with the data you used to create Figure 2.1. To create the kernel density plot, you can either use `gaussian_kde` from `scipy.stats` ([for an example, check out this stackoverflow post](https://stackoverflow.com/questions/4150171/how-to-create-a-density-plot-in-matplotlib)) or you can use [`seaborn.kdeplot`](https://seaborn.pydata.org/generated/seaborn.kdeplot.html).
# > * Now grab 25 random timepoints from the dataset (of 1000-10000 original data) you've just plotted and create a version of Figure 2-4 based on the 25 data points. Does this shed light on why I think KDEs can be misleading?
# >
# > Let's take a break. Get some coffee or water. Stretch your legs. Talk to your friends for a bit. Breathe. Get relaxed so you're ready for the second part of the exercise.
#
# +
from datetime import timedelta
df = pd.read_csv('data/crimedata.csv')
df['Date'] = pd.to_datetime(df['Date'])
df['Year'] = df['Date'].dt.year
df = df[df['Year'] != '2018']
df.head(5)
# -
set(df.Hour)
df['Datetime'] = df['Date'].dt.strftime('%Y-%m-%d')
df['Datetime'] = df['Datetime'] + ' ' + df['Time']
df['Datetime'] = pd.to_datetime(df['Datetime'])
df.head(5)
df['Month'] = df['Date'].dt.month
df['Day'] = df['Date'].dt.dayofweek
df['Hour'] = df['Datetime'].dt.hour
category = 'ASSAULT'
df_assault = df[df['Category'] == category]
df_assault = df_assault[(df_assault['Year'] == 2012) & (df_assault['Month'] == 6) & (df_assault['Hour'] == 12)]
df_assault_jitter = df_assault
df_assault_jitter['Y'] = np.random.uniform(-0.5, 0.5, size=len(df_assault_jitter))
fig, ax = plt.subplots()
ax.scatter(df_assault_jitter['Date'], df_assault_jitter['Y'])
ax.set_ylim(-10,10)
ax.get_yaxis().set_visible(False)
plt.xticks(rotation=90);
# +
cats = ['WARRANTS', 'BURGLARY']
df1 = df[df['Category'].isin(cats)]
df1 = df1[(df1['Year'] == 2012) & (df1['Month'].isin([5, 6, 7]))]
print(len(df1[df1['Category'] == cats[0]]), len(df1[df1['Category'] == cats[1]]))
plt.hist(df1[df1['Category'] == cats[0]]['Y'], bins=50);
plt.hist(df1[df1['Category'] == cats[1]]['Y'], bins=50);
# -
df.Category.value_counts()
#
# > *Exercise 3.2*. Ok. Now for more plots 😊
# > * Now we'll work on creating two versions of the plot in **Figure 2-11**, but using the GPS data you used for your version of Figure 2-2. Comment on the result. It is not easy to create this plot from scracth.
# **Hint:** Take a look at the `scipy.stats.probplot` function.
# > * OK, we're almost done, but we need some box plots. Here, I'd like you to use the box plots to visualize fluctuations of how many crimes happen per day. We'll use data from the 15 focus crimes defined last week.
# > * For the full time-span of the data, calulate the **number of crimes per day** within each category for the entire duration of the data.
# > * Create a box-and whiskers plot showing the mean, median, quantiles, etc for all 15 crime-types side-by-side. There are many ways to do this. I like to use [matplotlibs's built in functionality](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.boxplot.html), but you can also achieve good results with [seaborn](https://seaborn.pydata.org/generated/seaborn.boxplot.html) or [pandas](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.boxplot.html).
# > * What does this plot reveal that you can't see in the plots from last time?
# > * Also I want to show you guys another interesting use of box plots. To get started, let's calculate another average for each focus-crime, namely what time of day the crime happens. So this time, the distribution we want to plot is the average time-of-day that a crime takes place. There are many ways to do this, but let me describe one way to do it.
# * For datapoint, the only thing you care about is the time-of-day, so discard everything else.
# * You also have to deal with the fact that time is annoyingly not divided into nice units that go to 100 like many other numbers. I can think of two ways to deal with this.
# * For each time-of-day, simply encode it as seconds since midnight.
# * Or keep each whole hour, and convert the minute/second count to a percentage of an hour. So 10:15 $\rightarrow$ 10.25, 8:40 $\rightarrow$ 8.67, etc.
# * Now you can create box-plots to create an overview of *when various crimes occur*. Note that these plot have quite a different interpretation than ones we created in the previous exercise. Cool, right?
stats.probplot(df1[df1['Category'] == cats[0]]['Y'], plot=plt)
#stats.probplot(df1[df1['Category'] == cats[1]]['Y'], plot=plt)
| exercises/.ipynb_checkpoints/Week3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Rz6ODflA987g"
# **PRIMER PARCIAL**
#
#
#
#
#
#
#
#
# + id="Z2xziFRT98iz"
fecha = "18/03/2021"
nombre_apellido = "" # Debe ser un String
CI = "" # Debe ser un String
# OBS:
# MODIFICAR SOLO LO QUE TENGA 'None'
# + id="-uIWojN_8C9a"
# Actividad 1- Importar las librerías numpy( como np ), pandas( como pd ), matplotlib( como plt )
## MODIFICAR AQUI
import None
## =============
# + id="w-99LamtAYbD"
# Actividad 2- Importar el dataset 'datosfintrados.xls'
## MODIFICAR AQUI
datos = None
## =============
# + id="8H6HYH6BphCw"
# NO MODIFICAR
grouped = datos.groupby(datos.Asignatura)
datos_new = grouped.get_group("CALCULO 2")
data = datos_new[['Primer.Par','Segundo.Par']].values
labels = datos_new['Aprobado'].values
labels = np.array([1.0 if i=='S' else 0.0 for i in labels])
# + id="nBTYT_-a-y1k"
# Actividad 3- Imprimir la dimensión de los datos y las etiquetas
## MODIFICAR AQUI
data_shape = None
label_shape = None
## =============
print("Dimensión de los datos = ",data_shape)
print("Dimensión de las etiquetas = ",label_shape)
# + colab={"base_uri": "https://localhost:8080/"} id="MOrxNg4ceVkI" outputId="cb8324ac-fbf7-4d0e-ef99-82a5319df1af"
# Actividad 4- Separar los datos en conjunto de entrenamiento y de prueba con el "train_test_split"
# usar el test_size = 0.1
from sklearn.model_selection import train_test_split
## MODIFICAR AQUI
data_train, data_test, labels_train, labels_test = None
## =============
print("Dimensión de los datos entrenamiento = ",data_train.shape)
print("Dimensión de las etiquetas entrenamiento = ",data_test.shape)
print("Dimensión de los datos prueba = ",labels_train.shape)
print("Dimensión de las etiquetas prueba = ",labels_test.shape)
# + id="DRZiuHule0k4"
import keras
import keras.backend as K
from keras.models import Sequential
from keras.layers import Dense
# Actividad 5- Crear el modelo de una Red Neuronal Densa:
# * Usar la clase Sequential
# * 1 Capa de entrada de 7 neuronas, con función de activación lineal 'linear' e incluir la dimensión de entrada - input_shape
# * 2 Capas ocultas de 5 neuronas cada una y con función de activación tangente hiperbólico 'tanh'
# * 1 Capa Oculta de 10 neuronas y con función de activación sigmoide 'sigmoid'
## MODIFICAR AQUI
model = None
model.add(None)
model.add(None)
model.add(None)
model.add(None)
## =============
# + id="k9EfRMIEe1nT"
# Actividad 6- Completar la variable loss_func con la función de pérdida que creas que sea
# Lineal => 'mse'
# Binaria => 'binary_crossentropy'
## MODIFICAR AQUI
loss_func = 'binary_crossentropy'
## =============
model.compile(optimizer = 'adam', loss = loss_func, metrics = 'accuracy')
model.summary()
# + id="o_E7h93pe8Eu"
# NO MODIFICAR
# Entrenar el modelo
history = model.fit(data_train, labels_train, epochs=1000, verbose=1, validation_split=0.1)
print(history.history.keys())
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper right')
plt.show()
# + id="6m-zMRH_e-BO"
# Actividad 7- Evaluar el modelo usando model.predict sobre los datos de prueba (data_test)
## MODIFICAR AQUI
predictions = model.predict(data_test)
## =============
print(model.evaluate(data_test, labels_test))
print(model.metrics_names)
# + id="E1u5cBqKfAJK"
# NO MODIFICAR
from sklearn.metrics import confusion_matrix, classification_report
predict_label = predictions.round()
print('Cantidad de predicciones = {} \n'.format(predict_label.shape))
cf_matrix = confusion_matrix(predict_label, labels_test)
print(cf_matrix)
import seaborn as sns
sns.heatmap(cf_matrix/np.sum(cf_matrix), annot=True,
fmt='.2%', cmap='Blues')
# + id="NkYgg9hE82CV"
## =============
# + id="fUTeAaJH82NC"
## =============
# + id="WGjrnBCU82Yf"
## =============
# + id="ZDqJRL9oAjVA"
# NO MODIFICAR ESTE CÓDIGO
# Código de evaluación - Deberá copiar el texto generado en este campo y pegar en la evaluacion del aula virtual
print("Fecha = \'{}\'".format(fecha),end =";")
print("nombre_apellido = \'{}\'".format(nombre_apellido),end =";")
print("CI = \'{}\'".format(CI), end =";")
# Tema 1
print( 'Tema1 = {0:b}'.format(('np' in vars() or 'np' in globals())<<2|('pd' in vars() or 'pd' in globals())<<1|('plt' in vars() or 'plt' in globals())<<0), end =";")
# Tema 2
print( 'Tema2 = \'{}{}\''.format(int('datos' in vars() or 'datos' in globals()), datos['id_anony'].iloc[0]), end =";")
# Tema 3
print( 'Tema3 = {}{}{}'.format(data_shape[0],data_shape[1],label_shape[0]), end =";")
# Tema 4
print( 'Tema4 = {}{}'.format(data_train.size, labels_test.size), end =";")
# Tema 5
print( 'Tema5 = {}'.format(model.count_params()), end =";")
# Tema 6
print( 'Tema6 = \'{}\''.format(loss_func), end =";")
# Tema 6
print( 'Tema6 = {}'.format(predictions.size), end =";")
| 1erparcial/.ipynb_checkpoints/PrimerParcial_IA2021_FilaB-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + dc={"key": "5"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 1. Scala's real-world project repository data
# <p>With almost 30k commits and a history spanning over ten years, Scala is a mature programming language. It is a general-purpose programming language that has recently become another prominent language for data scientists.</p>
# <p>Scala is also an open source project. Open source projects have the advantage that their entire development histories -- who made changes, what was changed, code reviews, etc. -- are publicly available. </p>
# <p>We're going to read in, clean up, and visualize the real world project repository of Scala that spans data from a version control system (Git) as well as a project hosting site (GitHub). We will find out who has had the most influence on its development and who are the experts.</p>
# <p>The dataset we will use, which has been previously mined and extracted from GitHub, is comprised of three files:</p>
# <ol>
# <li><code>pulls_2011-2013.csv</code> contains the basic information about the pull requests, and spans from the end of 2011 up to (but not including) 2014.</li>
# <li><code>pulls_2014-2018.csv</code> contains identical information, and spans from 2014 up to 2018.</li>
# <li><code>pull_files.csv</code> contains the files that were modified by each pull request.</li>
# </ol>
# + dc={"key": "5"} jupyter={"outputs_hidden": true} tags=["sample_code"]
# Importing pandas
# ... YOUR CODE FOR TASK 1 ...
import pandas as pd
# Loading in the data
pulls_one = pd.read_csv('datasets/pulls_2011-2013.csv')
pulls_two = pd.read_csv('datasets/pulls_2014-2018.csv')
pull_files = pd.read_csv('datasets/pull_files.csv')
# + dc={"key": "12"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 2. Preparing and cleaning the data
# <p>First, we will need to combine the data from the two separate pull DataFrames. </p>
# <p>Next, the raw data extracted from GitHub contains dates in the ISO8601 format. However, <code>pandas</code> imports them as regular strings. To make our analysis easier, we need to convert the strings into Python's <code>DateTime</code> objects. <code>DateTime</code> objects have the important property that they can be compared and sorted.</p>
# <p>The pull request times are all in UTC (also known as Coordinated Universal Time). The commit times, however, are in the local time of the author with time zone information (number of hours difference from UTC). To make comparisons easy, we should convert all times to UTC.</p>
# + dc={"key": "12"} tags=["sample_code"]
# Append pulls_one to pulls_two
pulls = pd.concat([pulls_one, pulls_two])
# Convert the date for the pulls object
pulls['date'] = pd.to_datetime(pulls['date'], utc = True)
# + dc={"key": "19"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 3. Merging the DataFrames
# <p>The data extracted comes in two separate files. Merging the two DataFrames will make it easier for us to analyze the data in the future tasks.</p>
# + dc={"key": "19"} tags=["sample_code"]
# Merge the two DataFrames
data = pulls.merge(pull_files, on = 'pid')
data.head()
# + dc={"key": "26"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 4. Is the project still actively maintained?
# <p>The activity in an open source project is not very consistent. Some projects might be active for many years after the initial release, while others can slowly taper out into oblivion. Before committing to contributing to a project, it is important to understand the state of the project. Is development going steadily, or is there a drop? Has the project been abandoned altogether?</p>
# <p>The data used in this project was collected in January of 2018. We are interested in the evolution of the number of contributions up to that date.</p>
# <p>For Scala, we will do this by plotting a chart of the project's activity. We will calculate the number of pull requests submitted each (calendar) month during the project's lifetime. We will then plot these numbers to see the trend of contributions.</p>
# <ul>
# <li><p>A helpful reminder of how to access various components of a date can be found in <a href="https://campus.datacamp.com/courses/data-manipulation-with-pandas/slicing-and-indexing?ex=12">this exercise of Data Manipulation with pandas</a></p></li>
# <li><p>Additionally, recall that you can group by multiple variables by passing a list to <code>groupby()</code>. This video from <a href="https://campus.datacamp.com/courses/data-manipulation-with-pandas/aggregating-data-ad6d4643-0e95-470c-8299-f69cc4c83de8?ex=9">Data Manipulation with pandas</a> should help!</p></li>
# </ul>
# + dc={"key": "26"} tags=["sample_code"]
# %matplotlib inline
# Create a column that will store the month
data['month'] = pd.DatetimeIndex(data['date']).month
# Create a column that will store the year
data['year'] = pd.DatetimeIndex(data['date']).year
# Group by the month and year and count the pull requests
counts = data.groupby(['month', 'year']).count()
# print(counts)
# Plot the results
counts.plot(kind='bar', figsize = (12,4))
# + dc={"key": "33"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 5. Is there camaraderie in the project?
# <p>The organizational structure varies from one project to another, and it can influence your success as a contributor. A project that has a very small community might not be the best one to start working on. The small community might indicate a high barrier of entry. This can be caused by several factors, including a community that is reluctant to accept pull requests from "outsiders," that the code base is hard to work with, etc. However, a large community can serve as an indicator that the project is regularly accepting pull requests from new contributors. Such a project would be a good place to start.</p>
# <p>In order to evaluate the dynamics of the community, we will plot a histogram of the number of pull requests submitted by each user. A distribution that shows that there are few people that only contribute a small number of pull requests can be used as in indicator that the project is not welcoming of new contributors. </p>
# + dc={"key": "33"} tags=["sample_code"]
# Required for matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# Group by the submitter
by_user = data.groupby('user').agg({'pid':'sum'})
# Plot the histogram
by_user.hist(bins=10)
plt.xlabel('Number of Contributions')
plt.ylabel('Number of Contributor')
plt.title('Is the project welcoming to the new Contributors ?')
# + dc={"key": "40"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 6. What files were changed in the last ten pull requests?
# <p>Choosing the right place to make a contribution is as important as choosing the project to contribute to. Some parts of the code might be stable, some might be dead. Contributing there might not have the most impact. Therefore it is important to understand the parts of the system that have been recently changed. This allows us to pinpoint the "hot" areas of the code where most of the activity is happening. Focusing on those parts might not the most effective use of our times.</p>
# + dc={"key": "40"} tags=["sample_code"]
# Identify the last 10 pull requests
last_10 = pulls.nlargest(10, 'pid', keep='last')
# Join the two data sets
joined_pr = last_10.merge(pull_files, on = 'pid')
# Identify the unique files
files = set(joined_pr['file'].unique())
# Print the results
files
# + dc={"key": "47"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 7. Who made the most pull requests to a given file?
# <p>When contributing to a project, we might need some guidance. We might find ourselves needing some information regarding the codebase. It is important direct any questions to the right person. Contributors to open source projects generally have other day jobs, so their time is limited. It is important to address our questions to the right people. One way to identify the right target for our inquiries is by using their contribution history.</p>
# <p>We identified <code>src/compiler/scala/reflect/reify/phases/Calculate.scala</code> as being recently changed. We are interested in the top 3 developers who changed that file. Those developers are the ones most likely to have the best understanding of the code.</p>
# + dc={"key": "47"} tags=["sample_code"]
# This is the file we are interested in:
file = 'src/compiler/scala/reflect/reify/phases/Calculate.scala'
# Identify the commits that changed the file
file_pr = data[data['file'] == file]
# Count the number of changes made by each developer
author_counts = file_pr.groupby('user').count()
# Print the top 3 developers
print(list(author_counts.nlargest(3, 'pid').index))
# author_counts.sort_values(by='pid', ascending=False).head(3)
# + dc={"key": "54"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 8. Who made the last ten pull requests on a given file?
# <p>Open source projects suffer from fluctuating membership. This makes the problem of finding the right person more challenging: the person has to be knowledgeable <em>and</em> still be involved in the project. A person that contributed a lot in the past might no longer be available (or willing) to help. To get a better understanding, we need to investigate the more recent history of that particular part of the system. </p>
# <p>Like in the previous task, we will look at the history of <code>src/compiler/scala/reflect/reify/phases/Calculate.scala</code>.</p>
# + dc={"key": "54"} tags=["sample_code"]
file = 'src/compiler/scala/reflect/reify/phases/Calculate.scala'
# Select the pull requests that changed the target file
file_pr = pull_files[pull_files['file'] == file]
# Merge the obtained results with the pulls DataFrame
joined_pr = file_pr.merge(pulls, on = 'pid')
# Find the users of the last 10 most recent pull requests
users_last_10 = set(joined_pr.nlargest(10, 'pid', keep = 'last')['user'])
# Printing the results
users_last_10
# + dc={"key": "61"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 9. The pull requests of two special developers
# <p>Now that we have identified two potential contacts in the projects, we need to find the person who was most involved in the project in recent times. That person is most likely to answer our questions. For each calendar year, we are interested in understanding the number of pull requests the authors submitted. This will give us a high-level image of their contribution trend to the project.</p>
# + dc={"key": "61"} tags=["sample_code"]
# %matplotlib inline
# The developers we are interested in
authors = ['xeno-by', 'soc']
# Get all the developers' pull requests
by_author = pulls[pulls['user'].isin(authors)]
# Count the number of pull requests submitted each year
counts = by_author.groupby(['user', by_author.date.dt.year]).agg({'pid': 'count'}).reset_index()
# Convert the table to a wide format
counts_wide = counts.pivot_table(index='date', columns='user', values='pid', fill_value=0)
# Plot the results
# ... YOUR CODE FOR TASK 9 ...
counts_wide.plot.bar()
# + dc={"key": "68"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 10. Visualizing the contributions of each developer
# <p>As mentioned before, it is important to make a distinction between the global expertise and contribution levels and the contribution levels at a more granular level (file, submodule, etc.) In our case, we want to see which of our two developers of interest have the most experience with the code in a given file. We will measure experience by the number of pull requests submitted that affect that file and how recent those pull requests were submitted.</p>
# + dc={"key": "68"} tags=["sample_code"]
authors = ['xeno-by', 'soc']
file = 'src/compiler/scala/reflect/reify/phases/Calculate.scala'
# Select the pull requests submitted by the authors, from the `data` DataFrame
by_author = data[(data.user == authors[0]) | (data.user == authors[1])]
# Select the pull requests that affect the file
by_file = by_author[by_author.file == file]
# Group and count the number of PRs done by each user each year
grouped = by_file.groupby(['user', by_file['date'].dt.year]).count()['pid'].reset_index()
# Transform the data into a wide format
by_file_wide = grouped.pivot_table(index='date', columns='user', values='pid', fill_value=0)
# Plot the results
by_file_wide.plot(kind='bar')
| The GitHub History of the Scala Language/The GitHub History of the Scala Language/The GitHub History of the Scala Language.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Zinnenlauf/github-slideshow/blob/master/transfer_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="xp81GSFN7uHY"
# # Deep Learning week - Day 3 - Exercise 3: Transfer Learning
#
# This notebook is dedicated to **transfer learning** which corresponds to using an existing model that has been already trained on a particular task, and fine-tuning it for a different task.
#
# To that end, we will use the [VGG-16 Neural Network](https://neurohive.io/en/popular-networks/vgg16/), a well-known architecture that has been trained on ImageNet which is a very large database of images of different categories.
#
# The idea is that we will load the existing VGG16 network, remote the last fully connected layers, replace them by new connected layers whose weights are randomly set, and train these last layers on a specific classification task - here, separate types of flower. The underlying idea is that the first convolutional layers of VGG-16, that has already been trained, corresponds to filters that are able to extract meaning features from images.
#
# # Google Colab
#
# In the following, the computations are computationally heavy and may be too long on your computer. For this reason, we will work in a new environment: **Google Colab** which provides free access to GPU that accelerate the computational time.
#
# To do so, you have to :
# - Open Google Colab [here](https://colab.research.google.com/)
# - Import a Notebook and select this file, the one you are currently reading.
# - Once open, you a running a similar Notebook but in Google Colab.
# - Change the runtime type to GPU - default is CPU.
#
# # Data loading & Preprocessing
#
# You first have to download the [data here](https://wagon-public-datasets.s3.amazonaws.com/flowers-dataset.zip) and add them to your Google Drive in a folder called `Deep_learning_data`.
#
# Then, run the following code. It will ask you to go to a given webpage where you copy the link and past it in the Colab form that will appear:
# + [markdown] colab_type="text" id="riTyl7uf3YDC"
# Now, Google Colab has access to your Google Drive. You can therefore load the data you just added to your drive.
#
# ❓ **Question** ❓ Use the following code to load your data. Be patient, it can take several minutes to load.
# + id="XnNdWnWPr21T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="1e1ca37c-ab05-4d4a-8e3f-be9d494bd914"
from google.colab import drive
drive.mount('/content/drive')
# + id="hB0XEe6y7uH7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 73} outputId="507477e2-7c6f-43c0-d2f3-b1c98d762ba3"
from tqdm import tqdm
import numpy as np
import os
from PIL import Image
from tensorflow.keras.utils import to_categorical
def load_flowers_data():
data_path = '/content/drive/My Drive/Deep_learning_data/flowers'
classes = {'daisy':0, 'dandelion':1, 'rose':2}
imgs = []
labels = []
for (cl, i) in classes.items():
images_path = [elt for elt in os.listdir(os.path.join(data_path, cl)) if elt.find('.jpg')>0]
for img in tqdm(images_path[:300]):
path = os.path.join(data_path, cl, img)
if os.path.exists(path):
image = Image.open(path)
image = image.resize((256, 256))
imgs.append(np.array(image))
labels.append(i)
X = np.array(imgs)
num_classes = len(set(labels))
y = to_categorical(labels, num_classes)
# Finally we shuffle:
p = np.random.permutation(len(X))
X, y = X[p], y[p]
first_split = int(len(imgs) /6.)
second_split = first_split + int(len(imgs) * 0.2)
X_test, X_val, X_train = X[:first_split], X[first_split:second_split], X[second_split:]
y_test, y_val, y_train = y[:first_split], y[first_split:second_split], y[second_split:]
return X_train, y_train, X_val, y_val, X_test, y_test, num_classes
X_train, y_train, X_val, y_val, X_test, y_test, num_classes = load_flowers_data()
# + [markdown] id="kn_6QB6SroGO" colab_type="text"
# First things first, let's start by some preprocessing
#
# ❓ **Question** ❓ First check that all your images have the same size. How many colors do these images have?
# + id="dM0tilAiroGQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 73} outputId="b7b06ec2-9ec1-4b83-c98c-611cc3ef4f25"
print(f"X_train - {X_train.shape}")
print(f"X_val - {X_val.shape}")
print(f"X_test - {X_val.shape}")
# + id="zP9A1Zh-0np8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="ff04a82a-2800-4d8d-bdae-a2b3a78c819d"
print(f"y_train - {y_train.shape}")
# + id="cbYJnWZP0tgb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 148} outputId="9b5e8d2b-90ef-4040-dc2b-4787c9b04021"
y_train
# + [markdown] id="lObWeTgQroGZ" colab_type="text"
# ❓ **Question** ❓ Resize your images
# + id="LGVtAXhBroGa" colab_type="code" colab={}
import tensorflow as tf
X_train_resize = tf.image.resize(X_train, [224,224])
# + id="OUaWnGxt2VrK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="73e534cd-de9f-454c-f2eb-b3c0e77c0b56"
X_train_resize.shape
# + [markdown] colab_type="text" id="OOF7fVp27uI2"
# ❓ **Question** ❓ Plot some images with the `imshow` function of matplotlib. If you obtained an ugly image, figure out why and correct the problem !
# + id="tud67n_43A-V" colab_type="code" colab={}
X_train_resize = X_train / 255
X_test_resize = X_test / 255
X_val_resize = X_val / 255
# + id="9AxH4ew93R0_" colab_type="code" colab={}
import matplotlib.pyplot as plt
# %matplotlib inline
# + id="dXzHmPMW3Tmd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 288} outputId="72fa9337-08bf-4713-ea02-022a921bb760"
plt.imshow(X_train[1])
# + id="unZ0dY1SroGh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 288} outputId="07eebbc3-cf08-42bd-dad6-bd705b9d9a03"
plt.imshow(X_train_resize[1])
# + [markdown] colab_type="text" id="nr6m5eKs9s54"
# # The model
#
# Let's now build our model.
#
# ❓ **Question** ❓ Write a first function `load_model()` that loads the pretrained VGG-16 model from `tensorflow.keras.applications.vgg16`. Especially, look at the [documentation](https://www.tensorflow.org/api_docs/python/tf/keras/applications/VGG16) to load the model where:
# - the `weights` have been learnt on `imagenet`
# - the `input_shape` corresponds to the input shape of any of your images (which all should have the same shape as see previously)
# - the `include_top` argument is set to `False` in order not to load the fully-connected layers of the VGG-16
#
# ❗ **Remark** ❗ Do not change the default value of the other arguments
# + id="vdhoMaURroGo" colab_type="code" colab={}
from tensorflow.keras.applications.vgg16 import VGG16
def load_model():
model = VGG16(weights='imagenet', input_shape=(256, 256, 3),
include_top = False) # pooling=None, classes=1000, classifier_activation='softmax'
return model
# + [markdown] id="6xHeTjinroGu" colab_type="text"
# ❓ **Question** ❓ Look at the architecture of the model thanks to the summary method
# + id="9mepwmVproGv" colab_type="code" colab={}
model = load_model()
# + id="DimGKEUp47Uv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 877} outputId="e3d014d7-5f54-4a24-d67a-009d3baf7d3c"
model.summary()
# + [markdown] id="eZSUHJW9roG0" colab_type="text"
# Impressive, right? Two things to notice:
# - It ends with convolution. The flattening of the output and the fully connected layers are not here yet! We will add them
# - There are more than 14.000.000 parameters, which is a lot. We could fine-tune them, meaning still updates them in the model algorithm, but it will take a lot of time, therefore we will fix them so they are not trainable. And only the last layers (that we will add) will be trained and updates
#
# ❓ **Question** ❓ Write a function that takes the previous model as input, and does what we just described:
# - set the first layers to be not-trainable, by applying `model.trainable = False`
# - adding new layers that by default will be trainable.
#
# To do the second part (adding new layers), we will use another way to define the model (instead of `model.sequential()`). Look at the following code and try to apply it in your scenario:
#
# ```
# base_model = load_model()
# dense_layer = layers.Dense(SOME_NUMBER_1, activation='relu')
# prediction_layer = layers.Dense(SOME_NUMBER_2, activation='APPROPRIATE_ACTIVATION')
#
# model = tf.keras.Sequential([
# base_model,
# dense_layer,
# prediction_layer
# ])
#
# ```
#
# ❗ **Remark** ❗ Let's consider a dense layer with 500 neurons ; on the other hand, the prediction layer should be related to this specific task, so just replace `SOME_NUMBER_2` and `APPROPRIATE_ACTIVATION` with the correct values.
#
#
# ❗ **Remark** ❗ Do not forget to Flatten the convoluted outputs first, before the fully connected layers ;)
# + id="ZQyst9-36PWr" colab_type="code" colab={}
from tensorflow.keras import layers
from tensorflow.keras import models
# + id="zIVWE_jk7UFt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="0b76e5f1-d627-4a8a-ebcd-33a79edd4727"
y_train.shape
# + id="MpNMS1uuroG0" colab_type="code" colab={}
from tensorflow.keras import layers
from tensorflow.keras import models
def update_model():
base_model = load_model()
# Set the first layers to be untrainable
base_model.trainable = False
# Add layers to the model
flattened = layers.Flatten()
dense_layer = layers.Dense(500, activation='relu')
prediction_layer = layers.Dense(3, activation='softmax')
model = tf.keras.Sequential([
base_model,
flattened,
dense_layer,
prediction_layer
])
return model
# + [markdown] id="njiSznTxroG6" colab_type="text"
# ❓ **Question** ❓ Now look at the layers and parameters of your model. Note that there is a distinction, at the end, between the trainable and non-trainable parameters
# + id="vjgX4qb2roG6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 316} outputId="4ab87892-718d-4eca-cb78-6c7908acc3d6"
model_new = update_model()
model_new.summary()
# + [markdown] id="OIAICCTQroG_" colab_type="text"
# ❓ **Question** ❓ Write a function to compile your model - we advise Adam with `learning_rate=1e-4`.
# + id="7Zic50lFroHA" colab_type="code" colab={}
from tensorflow.keras import optimizers
def compile_model(model):
adam_opt = optimizers.Adam(learning_rate=0.1e-4)
model.compile(loss='categorical_crossentropy',
optimizer=adam_opt,
metrics=['accuracy'])
return model
model_compile = compile_model(model_new)
# + [markdown] id="dgLn5D3hroHE" colab_type="text"
# ❓ **Question** ❓ Write an overall function that :
# - load the model
# - update the layers
# - compiles it
# + id="tfLnAP3eroHF" colab_type="code" colab={}
def build_model():
model_updated = update_model()
model_compiled = compile_model(model_updated)
return model_compiled
# + id="UCP8pKrUDLYa" colab_type="code" colab={}
model = build_model()
# + id="WKgHUgoyB_nP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 316} outputId="17360ee0-ca31-4f3a-b55c-47089246afd8"
model.summary()
# + [markdown] id="IMfmWWX5roHK" colab_type="text"
# # Back to the data
#
# The VGG16 model was trained on images which were preprocessed in a specific way.
#
# ❓ **Question** ❓ Apply this processing to the images here using the method `preprocess_input` that you can import from `tensorflow.keras.applications.vgg16`.
# + id="uvoT6Yyb-BTP" colab_type="code" colab={}
# preprocess_input?
# + colab_type="code" id="B--Gyb-23YDb" colab={}
from tensorflow.keras.applications.vgg16 import preprocess_input
x_train_pre = preprocess_input(X_train)
x_test_pre = preprocess_input(X_test)
x_val_pre = preprocess_input(X_val)
# + id="rBXtd3A6KjLF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="830f6e24-23eb-4125-e1e5-69085eda4332"
y_val.shape
# + id="8YQ9E9jCGON1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="2168a347-d6bd-4a07-dbb6-1ce1654aac03"
X_train_resize.shape
# + id="5HdgX-_2GJ_p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="59741471-dd59-4d4c-9624-33fc03de8cff"
x_train_pre.shape
# + id="nCZD1LX8GmlM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="9185faa7-c703-493f-b672-20ec8642399f"
y_train.shape
# + id="dQgrP2g_GqC3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="b5d25a1f-ff9a-4e12-dbb1-55e025240965"
x_test_pre.shape
# + id="FE58ljqlGuqn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="1fc62d48-61a1-4cd3-c13b-910d47682d80"
x_val_pre.shape
# + id="VBKVvIUiKbvD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="669ff3a8-2043-48dd-e499-b757692f58ac"
y_val.shape
# + [markdown] colab_type="text" id="Wu2H0KZF-EoI"
# # Run the model
#
# ❓ **Question** ❓ Now estimate the model, with an early stopping criterion on the validation accuracy - here, the validation data are provided, therefore use `validation_data` instead of `validation_split`.
#
# ❗ **Remark** ❗ Store the results in a `history` variable
# + id="531EWQJCG98B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="80627f58-5ac3-4b23-9f8f-c5ef34cc67d1"
x_val_pre.shape
# + id="ZLQyGqUWG8To" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="dd8f0c10-6b4f-4a62-846a-8d4f0e513cc3"
y_val.shape
# + id="8KJY8I2u-8Nc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 879} outputId="db7c62c6-3591-482f-947a-95d04a22a4be"
from tensorflow.keras import callbacks
es = callbacks.EarlyStopping(patience=5, monitor = "val_acc", restore_best_weights = True, min_delta=1e-2)
history = model.fit(x_train_pre, y_train,
epochs=15,
batch_size=32,
verbose=1,
validation_data=[x_val_pre, y_val],
callbacks=[es])
# + [markdown] colab_type="text" id="z97kx9yUAas5"
# # YOUR CODE HERE
# + [markdown] colab_type="text" id="ec_I9JpiAm-W"
# ❓ **Question** ❓ Plot the accuracy for the test and validation set.
# + id="tzxurXw1IozQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="a9128071-260c-4c6c-b948-bd965e7220d7"
history.history.keys()
# + id="lKdwWKF0roHT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="b008da11-ef30-41b8-debe-836503aad5ab"
plt.plot(history.history["accuracy"])
plt.plot(history.history["val_accuracy"])
# + [markdown] colab_type="text" id="y3plexlQAtcC"
# ❓ **Question** ❓ Evaluate the model accuracy on the test set. What is the chance level on this classification task (i.e. accuracy of a random classifier).
# + colab_type="code" id="BzU0wCXlB6UI" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="c27de122-7b20-439a-8c53-a5d6e5a026f2"
model.evaluate(X_test_resize, y_test)
# + [markdown] colab_type="text" id="vzetiM3XA2fu"
# # Data augmentation
#
# The next question are a less guided but really on what you did in the previous exercise - don't hesitate to come back to what you have done.
#
# ❓ **Question** ❓ Use some data augmentation techniques for this task - you can store the fitting in a `history_data_aug` variable that you can plot. Do you see an improvement ? Don't forget to evaluate it on the test set
# + id="uUJAERGlroHc" colab_type="code" colab={}
from tensorflow.keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(
featurewise_center=False,
featurewise_std_normalization=False,
rotation_range=10,
width_shift_range=0.2,
height_shift_range=0.2,
horizontal_flip=True,
brightness_range=(0.8, 1.),
zoom_range=(0.8, 1.2),
rescale=1./255.)
datagen.fit(x_train_pre)
# + colab_type="code" id="IvsCub2JBre7" colab={}
train_flow = datagen.flow(x_train_pre, y_train, batch_size=16)
# + id="GFmM5jgvLcKh" colab_type="code" colab={}
val_flow = datagen.flow(x_val_pre, y_val, batch_size=16)
# + [markdown] colab_type="text" id="oF39HIb7BSOy"
# # Advanced [Optional]
#
# You can here try to improve the model test accuracy. To do that, here are some options you can consider
#
# 1) Is my model overfitting ? If yes, you can try more data augmentation. If no, try a more complex model (unlikely the case here)
#
# 2) Perform precise grid search on all the hyper-parameters: learning_rate, batch_size, data augmentation etc...
#
# 3) Change the base model to more modern one (resnet, efficient nets) available in the keras library
#
# 4) Curate the data: maintaining a sane data set is one of the keys to success.
#
# 5) Obtain more data
#
#
# ❗ **Remark** ❗ Note also that it is good practice to perform a real cross-validation. You can also try to do that here to be sure of your results.
# + colab_type="code" id="7IyqGWzGBN0Y" colab={}
| transfer_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importing packages
# +
import warnings
warnings.filterwarnings('ignore')
import xml.etree.ElementTree as ET
import sys
import os
import pandas as pd
# -
# ## Loading RAW DATASET
# +
DATA_DIR = "/data/user/tr27p/Courses/CS762-NLP/FinalProject/nlp-group-project-fall-2020-deepbiocomp/dataset/raw/1_CancerGov_QA"
DATA_FILES = sorted(os.listdir(DATA_DIR))
# -
# ## Parsing XML to Data Frame with necessary tags
# +
data_column_names = ["Answer"]
TXT_GEN_SIZE = 0
Answers = []
for each_file in DATA_FILES:
tree = ET.parse(DATA_DIR+'/'+each_file)
root = tree.getroot()
for each_passage in root.iter('Document'):
for each_Answer in each_passage.iter('Answer'):
Answers.append(each_Answer.text)
TXT_GEN_SIZE +=1
TXT_GEN_DATAFRAME = pd.DataFrame(Answers, columns = data_column_names)
# -
# ## Spliting Data Frames to TRAIN(80%), DEV(10%), TEST(10%)
# +
TRAIN_TXT_GEN_DATAFRAME = TXT_GEN_DATAFRAME.sample(frac=0.8,random_state=200)
temp_df = TXT_GEN_DATAFRAME.drop(TRAIN_TXT_GEN_DATAFRAME.index)
DEV_TXT_GEN_DATAFRAME = temp_df.sample(frac=0.5,random_state=200)
TEST_TXT_GEN_DATAFRAME = temp_df.drop(DEV_TXT_GEN_DATAFRAME.index)
TRAIN_TXT_GEN_DATAFRAME = TRAIN_TXT_GEN_DATAFRAME.reset_index(drop=True)
DEV_TXT_GEN_DATAFRAME = DEV_TXT_GEN_DATAFRAME.reset_index(drop=True)
TEST_TXT_GEN_DATAFRAME = TEST_TXT_GEN_DATAFRAME.reset_index(drop=True)
print(" Train datasize (80%) :", len(TRAIN_TXT_GEN_DATAFRAME),
"\n+ Validation datasize (10%): ",len(DEV_TXT_GEN_DATAFRAME),
"\n+ Test datasize (10%) : ", len(TEST_TXT_GEN_DATAFRAME),
"\n", "-"*32, "\n",
" Total QA size :", TXT_GEN_SIZE)
# -
# ## Added $\lt$BOS$\gt$ and $\lt$EOS$\gt$ tags
def dataFrame_to_text_generate(DataFrame_name, OutPut_json_file_name):
with open(OutPut_json_file_name, 'w') as outfile:
for index in range(len(DataFrame_name)):
temp_answer = DataFrame_name.Answer[index].split('\n')
for line in temp_answer:
line = line.strip()
if line:
temp_line = "<BOS>"+line+"<EOS>\n"
outfile.write(temp_line)
# ## Saving data to file
# +
PREPARED_DATA_PATH ='/data/user/tr27p/Courses/CS762-NLP/FinalProject/nlp-group-project-fall-2020-deepbiocomp/dataset/prepared-data/text-generation/'
TRAIN_TXT_GEN_TXT = PREPARED_DATA_PATH + 'train_text-generation.txt'
DEV_TXT_GEN_TXT = PREPARED_DATA_PATH + 'dev_text-generation.txt'
TEST_TXT_GEN_TXT = PREPARED_DATA_PATH + 'test_text-generation.txt'
dataFrame_to_text_generate(TRAIN_TXT_GEN_DATAFRAME, TRAIN_TXT_GEN_TXT)
dataFrame_to_text_generate(DEV_TXT_GEN_DATAFRAME, DEV_TXT_GEN_TXT)
dataFrame_to_text_generate(TEST_TXT_GEN_DATAFRAME, TEST_TXT_GEN_TXT)
# -
| dataset/data-prepare-script/data-prepare-script_text-generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import numpy as np
import pandas as pd
import pickle
from src.features.sqr import calculate_sqi
# -
# # DESARROLLO
sessions = pd.read_parquet('../data/raw/flex_desarrollo_sesiones.parquet')
sessions.head()
sessions = pd.read_parquet('../data/raw/flex_desarrollo_sesiones.parquet')
movements = pd.read_parquet('../data/raw/flex_desarrollo_movimientos.parquet')
def rename_movement_columns(df):
old_names = ['fecha', 'presion0', 'presion1', 'presion2', 'presion3',
'presion4', 'presion5', 'presion6', 'presion7', 'presion8',
'presion9', 'presion10', 'presion11', 'presion12',
'tipoMovimiento']
new_names = ['timestamp', 'pressure0', 'pressure1', 'pressure2',
'pressure3', 'pressure4', 'pressure5', 'pressure6',
'pressure7', 'pressure8','pressure9', 'pressure10',
'pressure11', 'pressure12', 'type']
return df.rename(columns=dict(zip(old_names, new_names)))
# +
movements = rename_movement_columns(movements)
def calculate_new_sqr(session):
session_movements = movements[movements['idSesion'] == session['idSesiones']]
return calculate_sqi(session['fechaInicio'], session['fechaInicio'], session_movements)
sessions['sqr'] = sessions.apply(calculate_new_sqr, axis=1)
# -
sessions.to_parquet('../data/processed/sessions_new_sqr_flex_desarrollo.parquet')
sessions.shape
sessions['sqr'][sessions['sqr'] < 100].shape
sessions['sqr'][sessions['sqr'] < 100].hist()
sessions_gt_7 = sessions[(sessions['fechaFin'] - sessions['fechaEntrada']) > pd.Timedelta(hours=7)]
sessions_gt_7.shape
sessions_gt_7['sqr'][sessions_gt_7['sqr'] < 100].hist()
# # PRO
sessions = pd.read_parquet('../data/raw/flex_sesiones.parquet')
movements = pd.read_parquet('../data/raw/flex_movimientos.parquet')
movements = rename_movement_columns(movements)
# sessions_gt_7 = sessions[(sessions['fechaFin'] - sessions['fechaInicio']) > pd.Timedelta(hours=7)]
sessions_gt_7.shape
# +
def calculate_new_sqr(session):
session_movements = movements[movements['idSesion'] == session['idSesiones']]
return calculate_sqi(session['fechaEntrada'], session['fechaInicio'], session_movements)
sessions_gt_7['sqr'] = sessions_gt_7.apply(calculate_new_sqr, axis=1)
# -
sessions_gt_7.to_parquet('../data/processed/sessions_new_sqr_flex.parquet')
# sessions_gt_7 = pd.read_parquet('../data/processed/sessions_new_sqr_flex.parquet')
sessions_gt_7['sqr'][sessions_gt_7['sqr'] < 100].hist()
sessions_gt_7['sqr'].hist()
print(sessions_gt_7['sqr'][sessions_gt_7['sqr'] < 100].shape)
print(sessions_gt_7['sqr'].shape)
sessions_gt_7[sessions_gt_7['sqr'] > 100]
sessions_gt_7[sessions_gt_7['idSesiones'] == 26732]
movements[movements['idSesion'] == 26732]
sessions = pd.read_parquet('../data/raw/flex_sesiones.parquet')
movements = pd.read_parquet('../data/raw/flex_movimientos.parquet')
movements = rename_movement_columns(movements)
# +
def calculate_new_sqr(session):
session_movements = movements[movements['idSesion'] == session['idSesiones']]
return calculate_sqi(session['fechaEntrada'], session['fechaInicio'], session_movements)
sessions['sqr'] = sessions.apply(calculate_new_sqr, axis=1)
# -
sessions.to_parquet('../data/processed/sessions_new_sqr_flex_not_filtered.parquet')
# sessions_gt_7 = pd.read_parquet('../data/processed/sessions_new_sqr_flex.parquet')
sessions[(sessions['fechaFin'] - sessions['fechaEntrada']) < pd.Timedelta(hours=7)]
| notebooks/new_sqr_data_preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# Previously, the majority of transport reactions were removed as there were quite some incorrect reactions present. However, now it is worth revisiting if we should reintroduce some transports.
#
# For example, various amino acids can also be produced and exported in a sort of overflow metabolism mechanism. Now this is prevented because there is no transport reaction. So here I will go through the list of amino acid transports and check what type of transport should be added and add them if it makes sense.
#
import cameo
import pandas as pd
import cobra.io
import escher
from escher import Builder
from cobra import Metabolite, Reaction
model = cobra.io.read_sbml_model('../model/p-thermo.xml')
df = pd.read_csv('../databases/transport_aa.csv', sep = ';')
df[0:5]
#add all extracellular metabolites
for index,react in df.iterrows():
met = react['Amino acid']
ex_met_id = met[:-2] + '_e'
model.add_metabolites(Metabolite(id=ex_met_id))
model.metabolites.get_by_id(ex_met_id).name = model.metabolites.get_by_id(met).name
model.metabolites.get_by_id(ex_met_id).formula = model.metabolites.get_by_id(met).formula
model.metabolites.get_by_id(ex_met_id).charge = model.metabolites.get_by_id(met).charge
model.metabolites.get_by_id(ex_met_id).annotation = model.metabolites.get_by_id(met).annotation
model.metabolites.get_by_id(ex_met_id).compartment = 'e'
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
#add all reactions to the model
for index,react in df.iterrows():
rct = react['ID']
model.add_reaction(Reaction(id=rct))
c_met = react['Amino acid']
e_met = c_met[:-2] + '_e'
model.reactions.get_by_id(rct).name = 'Transport of ' + model.metabolites.get_by_id(c_met).name
model.reactions.get_by_id(rct).annotation['sbo'] = 'SBO:0000185'
t_type = react['Transport type']
direction = react['Direction']
if t_type in 'passive':
model.reactions.get_by_id(rct).add_metabolites({
model.metabolites.get_by_id(c_met): -1,
model.metabolites.get_by_id(e_met): 1
})
if direction in 'rev':
model.reactions.get_by_id(rct).bounds = (-1000,1000)
elif direction in 'out':
model.reactions.get_by_id(rct).bounds = (0,1000)
elif direction in 'in':
model.reactions.get_by_id(rct).bounds = (-1000,0)
else:
print (rct)
elif t_type in 'sym_proton':
model.reactions.get_by_id(rct).add_metabolites({
model.metabolites.get_by_id(c_met): -1,
model.metabolites.get_by_id(e_met): 1,
model.metabolites.h_c:-1,
model.metabolites.h_e: 1
})
if direction in 'rev':
model.reactions.get_by_id(rct).bounds = (-1000,1000)
elif direction in 'out':
model.reactions.get_by_id(rct).bounds = (0,1000)
elif direction in 'in':
model.reactions.get_by_id(rct).bounds = (-1000,0)
else:
print (rct)
elif t_type in 'anti_proton':
model.reactions.get_by_id(rct).add_metabolites({
model.metabolites.get_by_id(c_met): 1,
model.metabolites.get_by_id(e_met): -1,
model.metabolites.h_c:-1,
model.metabolites.h_e: 1
})
if direction in 'rev':
model.reactions.get_by_id(rct).bounds = (-1000,1000)
elif direction in 'out':
model.reactions.get_by_id(rct).bounds = (-1000,0)
elif direction in 'in':
model.reactions.get_by_id(rct).bounds = (0,1000)
else:
print (rct)
elif t_type in 'abc':
model.reactions.get_by_id(rct).add_metabolites({
model.metabolites.get_by_id(c_met): 1,
model.metabolites.get_by_id(e_met): -1,
model.metabolites.atp_c:-1,
model.metabolites.h2o_c:-1,
model.metabolites.adp_c:1,
model.metabolites.h_c:1,
model.metabolites.pi_c:1
})
if direction in 'rev':
model.reactions.get_by_id(rct).bounds = (-1000,1000)
elif direction in 'out':
model.reactions.get_by_id(rct).bounds = (-1000,0)
elif direction in 'in':
model.reactions.get_by_id(rct).bounds = (0,1000)
else:
print (rct)
else:
print(rct.id)
#add exchange where they are not present already
ex_id = 'EX_'+ e_met
try:
model.add_boundary(model.metabolites.get_by_id(e_met),type = 'exchange', reaction_id = ex_id, sbo_term = 'SBO:0000627')
model.reactions.get_by_id(ex_id).bounds = (0,1000)
except ValueError: #for the exchanges already present, just be sure bounds only allow production
model.reactions.get_by_id(ex_id).bounds = (0,1000)
#check number of rcts added
len(model.reactions)
model.optimize().objective_value
# Biomass prediction doesn't change, as one would expect.
#save&commit
cobra.io.write_sbml_model(model,'../../model/p-thermo.xml')
| notebooks/36. Transport reactions AA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import ipywidgets as widgets
from ipywidgets import Button, HBox, VBox
# +
speed_f = 10.0
speed_s = 1.0
x = ['X {}'.format(-speed_f), 'X {}'.format(-speed_s), 'X {}'.format(speed_s), 'X {}'.format(speed_f)]
items = [Button(description=w) for w in x]
left_box = VBox([items[0], items[1]])
right_box = VBox([items[2], items[3]])
buttons = HBox([left_box, right_box])
output = widgets.Output()
display(buttons, output)
'''
b_x_fast_neg.on_click(x_fast_neg)
b_x_slow_neg.on_click(x_slow_neg)
b_x_slow_pos.on_click(x_slow_pos)
b_x_fast_pos.on_click(x_fast_pos)
def x_fast_neg(b):
print('x_fast_neg')
def x_slow_neg(b):
print('x_slow_neg')
def x_slow_pos(b):
print('x_slow_pos')
def x_fast_pos(b):
print('x_fast_neg')
'''
# -
| src/processes/alignToolsGUI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#<NAME> 2
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import multivariate_normal
class LinearReg:
#linear reg equ. 3.50, 3.51
def __init__(self, alpha, beta):
self.alpha = alpha
self.beta = beta
self.mean = None
self.prec = None
#update parameter
def fit(self, X, t):
if self.mean is not None:
mean_prev = self.mean
prec_prev = self.prec
else:
mean_prev = np.zeros(np.size(X, 1))
prec_prev = self.alpha * np.eye(np.size(X, 1))
self.prec = prec_prev + self.beta * X.T @ X
self.varr = np.linalg.inv(self.prec)
mean = self.varr@(prec_prev@mean_prev+self.beta * X.T @ t)
self.mean = mean
# + pycharm={"name": "#%%\n"}
n_iters = 20
# generate data space & shuffle
grid_res = 100
x1 = np.linspace(-1,1,n_iters)
np.random.shuffle(x1)
# generate target t as described in pg.154
y = 0.5*x1 - 0.3
g_noise = np.random.normal(0,0.2,n_iters)
t = y + g_noise
x2 = [np.ones(np.shape(x1)),x1]
x2 = np.transpose(x2)
x = np.linspace(-1, 1, n_iters)
X = [np.ones(np.shape(x)),x]
X = np.transpose(X)
#parameter space
w0, w1 = np.meshgrid(np.linspace(-1, 1, grid_res),np.linspace(-1, 1, grid_res))
w = np.array([w0, w1]).transpose(1, 2, 0)
LR = LinearReg(alpha=2., beta=25.)
llh = np.ones(np.shape(w0))
for begin,end in [[0, 0], [0, 1], [1, 2], [2, 20]]: #4 rows
LR.fit(x2[begin: end], t[begin: end])
#generating likelihood, build upon previous likelihood
plt.subplot(1, 3, 1)
for ind in range(begin, end):
mean_vec = w0+w1*x1[ind]
for i in range(1,grid_res):
for j in range(1,grid_res):
llh[i][j] *= multivariate_normal.pdf(t[ind], mean=mean_vec[i][j], cov=1/25)
plt.imshow(llh,cmap='plasma')
plt.xlabel("w0")
plt.ylabel("w1")
plt.title("Likelihood")
#plot prior/posterior
plt.subplot(1, 3, 2)
plt.scatter((-0.3+1)*grid_res/2, (0.5+1)*grid_res/2, s=100, marker="x")
plt.imshow(multivariate_normal.pdf(w, mean=LR.mean, cov=LR.varr),cmap='plasma')
plt.xlabel("w0")
plt.ylabel("w1")
plt.title("prior/posterior")
#plot data space
plt.subplot(1, 3, 3)
plt.scatter(x1[:end], t[:end], s=50, facecolor="none", edgecolor="blue")
plt.plot(x, X @ np.random.multivariate_normal(LR.mean, LR.varr, size=6).T, c="red")
plt.xlim(-1, 1)
plt.ylim(-1, 1)
plt.title("data space")
plt.gca().set_aspect('equal')
plt.show()
# + pycharm={"name": "#%%\n"}
n_iters = 25
x1 = np.linspace(-1,1,n_iters)
np.random.shuffle(x1)
t = np.sin(2 * np.pi * x1)
g_noise = np.random.normal(0,0.2,n_iters)
t = t + g_noise
x = np.linspace(0, 1, n_iters)
y_test = np.sin(2 * np.pi * x)
#generate w/ gaussian basis, 9 gaussians
m0 = np.linspace(0,1,9)
x2 = [np.ones(np.shape(x1))]
x_test2 = [np.ones(np.shape(x))]
for m in m0:
x2.append(multivariate_normal.pdf(x1,m,0.2))
x_test2.append(multivariate_normal.pdf(x,m,0.2))
x2 = np.transpose(x2)
x_test2 = np.transpose(x_test2)
LR = LinearReg(alpha=1e-3, beta=2.)
for begin, end in [[0, 1], [1, 3], [2, 7], [7, 25]]:
LR.fit(x2[begin: end], t[begin: end])
y = x_test2 @ LR.mean
std = np.sqrt(1 / LR.beta + np.sum(x_test2 @ LR.varr * x_test2,axis=1))
plt.scatter(x1[:end], t[:end], s=50, facecolor="none", edgecolor="blue")
plt.fill_between(x, y - std, y + std, color="red", alpha=0.3)#confidence
plt.plot(x, y_test)#group truth
plt.plot(x, y)#approx
plt.ylim(-1.8, 1.8)
plt.xlim(0, 1)
plt.show()
| proj2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#写函数,给定符号和行数,如’*’,5,可打印相应行数的如下图形:
def a(b,n):
for i in range(1,n+1):
print(' '*(n-i)+(b+' ')*i+' '*(n-i))
b=input("请输入符号:")
n=int(input("请输入行数:"))
a(b,n)
# +
#打印如下9*9 乘法口诀表,注意每列左侧竖向对齐
for i in range(1,10):
for j in range(1,10):
if i>=j:
print(i,'*',j,'=',i*j,end=' ')
print()
# +
#写函数,可检查一个数(2-100000之间整数)能不能表示成两个质数之和,如果能,则打印这两个质数。主程序用18及93887分别做测试
def check(n):
n=int(input"请输入一个数:")
# +
#有一个列表:[1, 2, 3, 4…n],n=20;请编写代码打印如下规律的输出(不允许直接拷贝并直接打印作为答案,想象n可能是变量):
numbers=[]
number=[]
for i in range(20):
numbers.append(i)
i += 1
for i in range(1,4):
print('[')
for j in range(i-1,i+5):
if j==i-1:
numbers[j]=(j+'*')
else:
print(',',numbers[j])
print(number)
j=0
print(']')
for i in range(4,19):
print('[')
for j in range(i-1,i+5):
if j==i+1:
print(numbers[j],'*')
else:
print(',',numbers[j])
print(number)
j=0
print(']')
for i in range(19,21):
print('[')
for j in range(i-1,i+5):
if j==i+4:
print(numbers[j],'*')
else:
print(',',numbers[j])
print(number)
j=0
print(']')
# -
| chapter2/homework/computer/middle/201611680696 .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Prepare Dataset for Model Training and Evaluating
# # Amazon Customer Reviews Dataset
#
# https://s3.amazonaws.com/amazon-reviews-pds/readme.html
#
# ## Schema
#
# - `marketplace`: 2-letter country code (in this case all "US").
# - `customer_id`: Random identifier that can be used to aggregate reviews written by a single author.
# - `review_id`: A unique ID for the review.
# - `product_id`: The Amazon Standard Identification Number (ASIN). `http://www.amazon.com/dp/<ASIN>` links to the product's detail page.
# - `product_parent`: The parent of that ASIN. Multiple ASINs (color or format variations of the same product) can roll up into a single parent.
# - `product_title`: Title description of the product.
# - `product_category`: Broad product category that can be used to group reviews (in this case digital videos).
# - `star_rating`: The review's rating (1 to 5 stars).
# - `helpful_votes`: Number of helpful votes for the review.
# - `total_votes`: Number of total votes the review received.
# - `vine`: Was the review written as part of the [Vine](https://www.amazon.com/gp/vine/help) program?
# - `verified_purchase`: Was the review from a verified purchase?
# - `review_headline`: The title of the review itself.
# - `review_body`: The text of the review.
# - `review_date`: The date the review was written.
# +
import boto3
import sagemaker
import pandas as pd
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
# -
# %store -r personalize_user_item_s3_uri
print(personalize_user_item_s3_uri)
# %store -r personalize_user_s3_uri
print(personalize_user_s3_uri)
# %store -r personalize_item_s3_uri
print(personalize_item_s3_uri)
# # TODO: Train the Model
# + language="javascript"
# Jupyter.notebook.save_checkpoint();
# Jupyter.notebook.session.delete();
| 02_usecases/personalize_recommendations/wip/05_Train_Model_Recommend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CS146 Problem Set 1
# ## Call center data modeling & other exercises
# ## Problem 1: Call center data modeling
# ### LOs: #PythonImplementation, #interpretingProbabilities
# Task1 : Compute a 98% posterior confidence interval over the number of calls per minute (the call rate lambda) for each hour of the data which will lead in 24 confidence intervals.
#
# Task2 : Compute the posterior mean of lambda for each hour of the day.
#
# Task 3 : Present the results graphically.
#
# Task 4 : Write a paragraph to accompany you plot and present your findings to the client. (Summarize how many calls you expect during different parts of the day, and how much uncertainty there is in your estimates.)
#import the libraries that we need for the task
import scipy as sp
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as sts
# %matplotlib inline
# From the preclass and class activities of session 4, load the data and re-form the data for us to use (Codes Reused)
# +
#Let's load the data
waiting_times_day = np.loadtxt('https://course-resources.minerva.kgi.edu/uploaded_files/mke/'
'00124343-8397/call-center.csv')
#Display the basics of the Data
print('Size of data set:', len(waiting_times_day))
print('First 3 values in data set:', waiting_times_day[:3])
print('Sum of data set:', sum(waiting_times_day))
#data pre-processing
#from the preclass work from before, make 24 hrs list of waiting times per hour
waiting_times_per_hour = [[] for _ in range(24)]
# Split the data into 24 separate series, one for each hour of the day.
current_time = 0
for t in waiting_times_day:
current_hour = int(current_time // 60)
current_time += t
waiting_times_per_hour[current_hour].append(t)
table_time = []
table_calls = []
for hour, calls_in_hour in enumerate(waiting_times_per_hour):
print(f'{hour:02}:00-{hour + 1:02}:00 - {len(calls_in_hour)} calls')
table_time.append([f'{hour:02}:00-{hour + 1:02}:00'])
table_calls.append([len(calls_in_hour)])
#show the data
#from above task, we have the data. Plot the graph of numbers of calls in specific time intervals
plt.bar(list(range(24)), [int(i) for i in np.array(table_calls)])
plt.xticks(list(range(24)),table_time, rotation = 90)
plt.show()
# Plot histogram of waiting times for one hour
hour_index = 11
waiting_times_hour = waiting_times_per_hour[hour_index]
plt.figure(figsize=(8, 6))
plt.hist(waiting_times_hour, bins=20)
plt.xlabel('Time between calls [minutes]')
plt.ylabel('Count')
plt.title(f'Histogram of waiting times for hour {hour_index}')
#fit the likelihood
# Exponential distribution with maximum likelihood fit to the data
lambda_ = 1 / np.mean(waiting_times_hour)
distribution = sts.expon(scale=1/lambda_)
plt.figure(figsize=(8, 6))
plt.hist(waiting_times_hour, bins=20, density=True, color='#c0c0c0') #to make the distribution to have 1 as the total area, density = true
x = np.linspace(0, max(waiting_times_hour), 200)
y = distribution.pdf(x)
plt.plot(x, y)
plt.xlabel('Time between calls [minutes]')
plt.ylabel('Density')
plt.title(f'Fit of maximum likelihood estimate to the data for hour {hour_index}')
plt.show()
# -
# For it to be done correctly, we have to define what distributions that we would like to use for likelihood and prior probability. Here, our task is to see the number of calls in certain period of time, which is always positive and always a natural number (positive integer). From the class, we have chosen exponential distribution for the likelihood because of this characteristic of the variable, but also because we can use a single variable lambda to represent the rate of the calls(lambda) for the distribution as exponential distribution is used for modeling the time between the events(calls). From the class, we have discussed about this by looking at the certain time duration's histogram we saw the exponential distribution to back up with the idea.(We could have done gamma, but that would mean that we need to consider two hyperparameters when we can just live with estimating only one)
#
# Now considering that the likelihood is exponential distribution, we also in the class, have used gamma distribution as the conjugte prior distribution so that we can easily define the posterior distribution which will become gamma distribution with a new hyper parameters that prior and the data makes (such as alpha for posterior will be alpha of prior + size of the data).
#
# As I have no idea what will the hyperparameter of the prior will be, I would like to stick with the value that we have done during the class. Alpha = 1, beta = 0.25
# +
#Hyperparameter_prior
a_0 = 1
b_0 = 0.25
#Make an empty list to save the values of the posterior parameter
a_post = list()
b_post = list()
#from given data and parameter of prior, we can calculate the parameter of the posterior (gamma dist)
#calculations are based from given link : https://en.wikipedia.org/wiki/Conjugate_prior
for i in range(len(waiting_times_per_hour)):
a_post.append(a_0 + len(waiting_times_per_hour[i])) #a_post = a_0 + n
b_post.append(b_0 + sum(waiting_times_per_hour[i])) #b_post = b_0 + sum of x from 1 to n
#gamma distribution for posterior (as we now have the parameter, we can draw the distribution)
#we need mean, 98% intervals to plot them.
posterior_dist = [] #have graphs as elements
post_mean = [] #mean of each time section gonna be saved as lists
post_upper = [] #upperbound of 98% interval
post_lower = [] #lowerbound of 98% interval
#draw graph using parts of the codes from class session 4
plt.figure(figsize = (8,6))
plt.xlabel('Hrs of the day')
plt.ylabel('Calls per minute')
plt.title('Mean & 98% interval error bar graph over call rate')
for i in range(len(a_post)):
posterior = sts.gamma(a = a_post[i], scale = 1/b_post[i]) #for each hyperparameter of that time slot, draw a gamma dist
posterior_dist.append(posterior)
post_mean.append(posterior.mean())
#calculating 98%interval from sample data: https://stackoverflow.com/questions/15033511/compute-a-confidence-interval-from-sample-data
post_upper.append(posterior.ppf((1+0.98)/2) - posterior.mean())
post_lower.append(posterior.mean()-posterior.ppf((1-0.98)/2))
plt.xticks(list(range(24)),table_time, rotation = 90)
plt.yticks(list(range(17)))
plt.errorbar(list(range(24)), post_mean, yerr = (post_upper, post_lower), color = 'green')
plt.show()
# -
# Now we know that there are two modes where the call rate surges around 11am and 6pm, it is evident that there needs to be more call respondents for this time duration. But as the confidence interval can vary up to a bit more than +- 1, having the 'mean' as the criteria for number of respondents are not ideal, which might cause some customers to wait if the call rates are bigger than the mean at the expected moment. So, by considering 98% interval as the boundary, we need to see the upper bound for making the choice of allocating the number of respondents at the time for the sake of saving customers' waiting time.
# ## Problem 2: Bent coin inference
# ### LOs: #BayesInference
#
# A normal coin has a probability of 0.5 of coming up heads when it is flipped. In this problem, we consider a bent coin that has an unknown probability of coming up heads. You flip the coin 20 times and the coin comes up heads 6 times. Now calculate the probability that the coin is biased in favor of heads by addressing all the questions and tasks below.
#
# 1. Let pi denote the probability that the bent coin comes up heads when flipped.
#
# 2. Use a binomial distribution for the likelihood function and a beta distribution for the prior over pi (likelihood - binomial dist, prior - beta, according to the wikipiedia[https://en.wikipedia.org/wiki/Conjugate_prior], I will calculate)
#
# 3. Select and motivate for your choice of parameters for the prior distribution (as I have no idea how bent the coin is, I want to use uniform distribution so that the data can dictate the outcome, which means that I am giving equal chance for any pi to be possible from prior distribution. This results alpha = 1, beta = 1 for prior to have under the x_axis interval of 0 to 1 as pi is going to be a probability.)
#
# 4. Using the observations above (6 heads out of 20 coin flips), compute the exact posterior distribution over pi
#
# 5. Plot the posterior distribution over pi and show a 95% credible interval of pi on the plot.
#
# 6. Compute the probability that the coin is biased in favor of heads - that it is more probable for the coin to come up heads rather than tails.
#for calculating 95% percentile, from the class, this was from professor's code
def compute_percentile(parameter_values, distribution_values, percentile):
'''
Compute the parameter value at a particular percentile of the given
probability distribution values. This function uses the cumulative trapezoid
integrator in SciPy.
Inputs:
parameter_values (array of float) This is the list of parameter values
at which the probability distribution has been evaluated.
distribution_values (array of float) This is the list of values of the
probability density function evaluated at the parameter values above.
percentile (float) This is the value between 0 and 1 of the percentile
to compute.
Returns: (float) The parameter value at the given percentile.
'''
cumulative_distribution = sp.integrate.cumtrapz(
distribution_values, parameter_values)
percentile_index = np.searchsorted(cumulative_distribution, percentile)
return lambdas[percentile_index]
# +
#calculate and plot posterior
#parameters of prior
a0 = 1 #alpha of beta dist
b0 = 1 #beta of beta dist
#posterior distribution calc
a1 = a0 + 6 #new alpha for posterior given the data : 6heads out of 20 trials
b1 = b0 + 20 - 6 #new beta for posterior given the data
posterior = sts.beta.pdf(np.linspace(0,1,50), a1, b1)
#calculate 95% interval using the codes from the class
percentiles = [compute_percentile(np.linspace(0,1,50), posterior, p ) for p in [0.025, 0.975]]
#plot the posterior and 95% credible interval
plt.title('posterior distribution and 95% credible interval')
plt.axvline(percentiles[0], color = 'orange')
plt.axvline(percentiles[1], color = 'orange', label = '95% credible interval')
plt.plot(np.linspace(0,1,50), posterior, color = 'green', label = 'posterior distribution')
plt.xlabel('pi')
plt.ylabel('probability density')
plt.legend()
plt.show()
print(percentiles)
# -
# This means that for the probability to have the coin biased to have more heads than tails is less than 2.5% from given data as the 95% interval upper boundary given above states 0.52 which is near 0.5 that is when heads and tails are both equally likely to happen. (bigger the pi, higher the probability for heads to come. right side of the green graph's area from \[0.52, 1\] is less than 2.5% is what I mean)
#
| CS146 Assignment 2 Problem set 1 Jeong woo Park.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# here's a file that has no bottom row on the first page
import pdfplumber
path = "../srcpdfs/UFOReports2006WholeoftheUK.pdf"
pdf = pdfplumber.open(path)
# on the first page the bottom row of the table isn't identified by default settings of tablefinder
p0 = pdf.pages[0]
im = p0.to_image()
im.reset().debug_tablefinder()
# the last row returned by the tablefinder is not Basildon by Oldbury
p0.extract_table()[-1]
# we can get the debug information of the tablefinder() and use the last set of decimal values
# to determine where the bottom row should be
db = p0.debug_tablefinder()
starts_at =db.cells[-1][1] # the position the border top
ends_at =db.cells[-1][3] # the position of border bottom
diff = ends_at - starts_at # the row height
new_ends_at = ends_at + diff # border bottom plus row height
print(new_ends_at)
# we can pass a setting to the table finder to add a horizontal line to close off the table
settings={
"explicit_horizontal_lines":[
new_ends_at
],
}
# and then we can re-try the tablefinder
im.reset().debug_tablefinder(settings)
# now we should get Basildon as our last row of data
p0.extract_table(settings)[-1]
| notebooks/PDFPlumber handling tables with no bottom rows.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="YohSkewix2HX"
# # Homework 3: Exploratory Data Analysis
#
# *In this homework, you are going to perform exploratory data analysis (EDA) on a dataset compiled by a research group from Harvard University.*
#
# **Submission Instructions**
#
# ---
# It is important that you follow the submission instructions.
# 1. Copy this assignment notebook to your Drive. <font color = 'red'> `File` --> `Save a copy in Drive`</font>. Rename it as <font color = 'green'>`Lastname_Firstname_hw3`</font>.
#
# 2. Write your solutions in the cells marked <font color = 'green'>`# your code`</font>.
#
# 3. **Do not delete your outputs. They are essential for the grading. Make sure that cells containing your solutions are executed, and the results are displayed on the notebook.**
#
# 4. When you're done please submit your solutions as an <font color="red">`.ipynb`</font> file. To do so:
#
# - Click on <font color="red">`File`</font> at the top left on the Colab screen, then click on <font color = 'red'>`Download .ipynb`</font>.
# - Then submit the downloaded <font color="red">`.ipynb`</font> version of your work on SUCourse.
#
#
# For any question, you may send an email to your TAs and LAs.
#
# ---
#
#
# + [markdown] id="IRa9WpBUyaFK"
# ## Income Segregation by Education Level
#
# In 2017, Chetty et al. compiled an anonymous data from USA Federal Government, in which they recorded the earnings of students born between 1980 and 1990 in their thirties. In addition, they recorded the earnings of their parents as well. In their study, they analyze the future financial status of students coming from different parental income levels and display how colleges help students progress.
#
# More information and the paper itself can be obtained from here: https://opportunityinsights.org/paper/undermatching/
#
# In this homework, you are going to analyze the dataset compiled in this study, `mrc_table3.csv`. In addition to the dataset, we also shared a PDF document, named `Codebook-MRC-Table-3.pdf`, prepared by the research team as the data dictionary that displays the columns and their explanations.
#
# The dataset is indexed by `cohorts`. In this context, a cohort is a group of students of the same age and college. In the image below a snippet from the dataset can be observed. Although `cohort` is a column name for the students' age; the actual cohort can be conceptualized as of age + college. For instance, the first row in the image below captures the attributes of the students born in 1980 and attended Brown University.
#
# 
#
# As stated above, the dataset stores the estimated financial status of the students in their 30s and their parents. In addition to storing mean income values to represent financial status, such as `par_mean` and `k_mean`, the researchers also provide a set of attributes to capture the relative information. To this end, they utilize *quintiles* and *percentiles* to represent fractions of the cohort.
#
# Below, you may find some of the column patterns that utilize quintiles and percentiles, along with their explanations.
#
# - **par_q[PARQUINT]**: Fraction of parents in income quintile [PARQUINT]. 1 is the bottom quintile and 5 is the top.
# - Remember that each row stores the financial status of that cohort's students and their families financial attributes. The value in this attribute captures the fraction of parents that reside in the [PARQUINT] quintile.
# - Since, with quintiles we basically divide the data into 5 different regions, [PARQUINT] can take values between 1 and 5.
# - 1 -> bottom quintile, in other words, lowest income level
# - 5 -> top quintile, or the highest income level
# - *So, there are five columns that store the fraction of parents in that quintile, e.g. `par_q5` stores the percentage of the parents that are in the top quintile.*
#
# - **k_top[PCTILE]pc**: Fraction of students in the top [PCTILE] percentile. For instance, `top1pc` refers to children in the top 1% of the income
# distribution.
# - The columns that contains the [PCTILE] tag captures the fractions with respect to `percentiles`.
# - As stated in the these attributes store the percentage of students that reside in the top [PCTILE]% of the income.
# - *e.g. If `k_top1pc` is set to 0.56, then we can conclude that 56% of the students in that cohort are in the top 1% of the income distribution in their 30s.*
#
# And lastly, the researchers provide conditional probabilities as a financial projection for the students.
#
# - **ktop1pc_cond_parq[PARQUINT]**: Probability of student in top 1%, conditional on parent in quintile [PARQUINT].
# - *e.g. ktop1pc_cond_parq1 stores the probability of a student being in the top 1% income level given that his/her parents were in the bottom quintile.*
#
# - **kq[KIDQUINT]_cond_parq[PARQUINT]**: Probability of kid in quintile [KIDQUINT], conditional on parent in quintile [PARQUINT].
# - *e.g. kq5_cond_parq1 stores the probability of a student being in the top income quintile given that his/her parents were in the bottom quintile.*
#
# *p.s. In this notebook, the terms `students`, `child` and `children` are used interchangeably. Each usage refers to a cohort.*
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="2ChTXnRVx8v4" outputId="74e04013-30c8-406e-a6c0-e637c8baa790"
from google.colab import drive
drive.mount("./drive")
path_prefix = "./drive/My Drive"
# + id="6h1nTH78yIvc"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from os.path import join
import matplotlib.ticker as mtick
from scipy.stats import iqr
import seaborn as sns
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 580} id="x3F7ycZnyY1-" outputId="e253b132-cd03-4ec0-e35f-08bc6edc6fe8"
fname = "mrc_table3.csv"
df = pd.read_csv(join(path_prefix, fname))
df.head()
# + [markdown] id="p2i_C80cNaEk"
# ## Get to Know the Dataset
#
# In this section, you are going to display the distribution of some attributes to better understand the data you are dealing with.
# + [markdown] id="rGyEj6s9TlDa"
# ### Q1: NaN Values
#
# In this notebook, we are not interested in all of the attributes. From the non-financial attributes, we are only interested in the `cohort`, `name`, `tier_name` and `type` columns. You need to make sure that there is no NaN value in these columns. And for the financial attributes we're interested in the all of the columns begining with `par_mean` (index 17 in the columns) till the end.
#
# 1. Check and print the NaN distributions in `cohort`, `name`, `tier_name` and `type` columns. If there are any NaN values in those columns, remove the corresponding rows.
# 2. Directly remove the rows where at least one NaN value exists in the financial attributes. *Notice that the columns starting from index 17 till the end are all financial attributes.*
#
# + colab={"base_uri": "https://localhost:8080/"} id="CTreFkXRb_FP" outputId="cac3dff8-e91f-44d1-9477-a3ce247c509d"
# NaN values in part 1
print('cohort: {}'.format(df['cohort'].isnull().sum()),
'\nname: {}'.format(df['name'].isnull().sum()),
'\ntier_name: {}'.format(df['tier_name'].isnull().sum()),
'\ntype: {}'.format(df['type'].isnull().sum()))
df = df.dropna(subset =['type'], axis=0 ) # 24 rows with NaN val in 'type' removed
# + colab={"base_uri": "https://localhost:8080/"} id="y-HgUPHKZEu-" outputId="e838cd37-4886-449a-9eeb-63278682919c"
pd.set_option('display.max_rows',1000) # in order to show all the financial cols
df_c = df.copy()
df_c = df[pd.notnull(df['k_married_cond_parq5'])]
df_c.iloc[:,17:].isnull().sum()
# + [markdown] id="r6eR2fpjPSpW"
# ### Q2: College Tier Distribution
#
# In the dataset, colleges are categorized into types and tiers at differing granularities. In the `type` column, the colleges are categorized as `public`, `private non-profit` and `for-profit`.
#
# Find the number of colleges in each type and display their percentages as a bar chart.
#
# The result should look like the figure below.
#
# 
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="urBrJUQeMIWK" outputId="3d2ef52f-2b50-43ac-f73a-1dff243161f6"
public = 0
private = 0
profit = 0
for i in df_c['type']:
if i == 3:
profit += 1
elif i == 2:
private += 1
elif i == 1:
public +=1
sum = profit + private + public
Rpublic = public / sum * 100
Rprivate = private / sum * 100
Rprofit = profit / sum * 100
collegeTier = pd.DataFrame( {'type': ['for-profit', 'private non-profit', 'public'],'College Distribution by Type': [Rprofit,Rprivate,Rpublic] })
ax = collegeTier.plot.barh(x = 'type', y = 'College Distribution by Type' ,alpha = 0.75 ,color = 'red', figsize=(8,5), fontsize = 12)
plt.title('College Distribution by Type', fontsize = 14)
ax.set_xlabel('Share', fontsize = 12)
fmt = '{x:,.0f}%'
tick = mtick.StrMethodFormatter(fmt)
ax.xaxis.set_major_formatter(tick)
# + [markdown] id="gU1udO6vRSrC"
# ### Q3: Student & Parent Income Distribution
#
# Columns `par_mean` and `k_mean` store the mean income for the parents and students in a cohort. In order to understand the overall distribution, display the `par_mean` and `k_mean` attributes as boxplots on the same figure.
#
# However, the mean distributions are highly skewed. So, in order to better evaluate the distributions, we can remove the outliers.
#
# - Create a 2x1 grid layout. Display the boxplots of the original distributions on the left.
#
# - Remove the outliers from both of the distributions by utilizing 1.5xIQR rule.
#
# - Generate the boxplots for the resulting distributions on the right axes.
#
# The result should look like the figure below.
#
# 
# + colab={"base_uri": "https://localhost:8080/", "height": 655} id="gGama5zVcbyG" outputId="a92f35d3-4739-45cc-c1ae-8d6917303242"
mean_income = pd.DataFrame({'Parent Mean Income':df_c['par_mean'] /1000,'Student Mean Income': df_c['k_mean']/1000})
boxplot = mean_income.boxplot(column=['Parent Mean Income','Student Mean Income'],figsize=(9,5))
boxplot.set_title('Mean Income Distribution per Parent and Students')
boxplot.set_yticks(np.arange(0,mean_income.max().max() , 146.24))
boxplot.set_ylabel('Income in ($) Thousands')
boxplot.grid(True)
fmt = '${x:,.0f}K'
tick = mtick.StrMethodFormatter(fmt)
boxplot.yaxis.set_major_formatter(tick)
plt.show()
Q1 = mean_income.quantile(0.25)
Q3 = mean_income.quantile(0.75)
IQR = Q3 - Q1
outliers = mean_income[~((mean_income < (Q1 - 1.5 * IQR)) |(mean_income > (Q3 + 1.5 * IQR))).any(axis = 1)]
boxplot2 = outliers.boxplot(column=['Parent Mean Income','Student Mean Income'],figsize=(9,5))
boxplot2.set_title('Mean Income Distribution per Parent and Students- Outliers Removed')
boxplot2.set_yticks(np.arange(0,outliers.max().max() , 22.65))
boxplot2.set_ylabel('Income in ($) Thousands')
boxplot2.grid(True)
fmt = '${x:,.0f}K'
tick = mtick.StrMethodFormatter(fmt)
boxplot2.yaxis.set_major_formatter(tick)
plt.show()
# + [markdown] id="HSBrIqgmdfR-"
# ## Bivariate Analysis
#
# In this section, you are going to perform bivariate analysis on different attribute pairs.
# + [markdown] id="YXeg-MLgdt5O"
# ### Q1: Parent Income Distribution by College Tier
#
# The income distribution is highly skewed as it can be observed in the previous question. With the generated charts, we see how the overall distribution is shaped with the help of boxplots. However, we can not observe how this distribution changes with respect to college tiers.
#
# As you can see from the shared data dictionary, there are 14 different college types. Instead of putting all of the tiers into account, in this question, you are going to focus on 6 of them: `Ivy Plus, Other elite schools (public and private), Highly selective public, Highly selective private, Selective public, Selective private`. Display the `par_mean` distribution for each of the selected tiers.
#
# - Group the dataframe with respect to the selected tier types.
# - For each group, display the `par_mean` attribute on the same figure as a boxplot.
# - Sort the boxplots with respect to their medians.
#
# The result should look like the figure below.
#
# 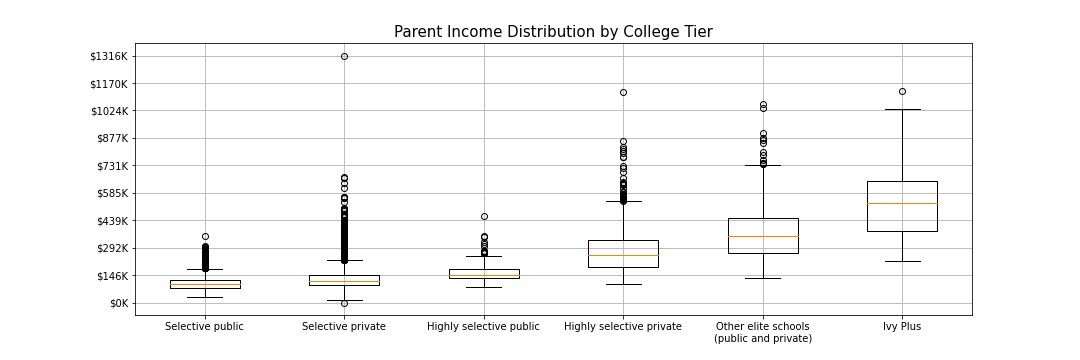
#
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="7MMiuQTd2ceb" outputId="4b515722-7850-46b8-eb1d-2d381bc3cb9e"
df_x = df_c.copy()
colleges = df_x.loc[(df_x['tier_name'] == "Selective public") | (df_x['tier_name'] == "Selective private") |
(df_x['tier_name'] == "Highly selective public") | (df_x['tier_name'] == "Highly selective private")|
(df_x['tier_name'] == "Ivy Plus") | (df_x['tier_name'] == "Other elite schools (public and private)")]
colleges = colleges[['tier_name','par_mean']]
colleges['par_mean'] = colleges['par_mean'] / 1000
grouped = colleges.groupby(['tier_name'])
df_grp = pd.DataFrame({col:val['par_mean'] for col,val in grouped})
meds = df_grp.median()
meds.sort_values(ascending=True, inplace=True)
df_grp = df_grp[meds.index]
multiBox= df_grp.boxplot(figsize=(18,5))
multiBox.set_title('Parent Income Distribution by College Tier', fontsize = 14)
multiBox.set_yticks(np.arange(0,mean_income.max().max() , 146.24))
fmt = '${x:,.0f}K'
tick = mtick.StrMethodFormatter(fmt)
multiBox.yaxis.set_major_formatter(tick)
plt.show()
# + [markdown] id="Io8lIi9Y2yyq"
# ### Q2: Mean Child Rank vs. Age at Income Measurement by College Tier
#
# In this question, you are going to display how the mean student income rank changes as the age of income measurement changes for the selected college tiers. In the dataset, we have students born between 1980 and 1991. In **2014**, their income level is measured. In the dataset, the `k_rank` column stores the student income rank.
#
# - First, find the age of each cohort by subtracting the birth years from the year of measurement and store them in a new column named `measurement_age`.
#
# - Group the dataframe by `tier_name` and `age`, and find the mean student income rank for each group.
#
# - For the listed tier names below, display the change of mean student income rank with respect to the age of measurement as a line chart.
#
# `Ivy Plus, Other elite schools (public and private), Highly selective public, Highly selective private, Selective public, Selective private`
#
# The result should look like the figure below.
#
# 
#
# *Hint: You may use the unstack function alongise transposition.*
#
# *Please visit the [documentation](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.unstack.html) for the details on unstack, and [this link](https://cmdlinetips.com/2020/05/fun-with-pandas-groupby-aggregate-multi-index-and-unstack/) for the use cases.*
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 590} id="NSz5Ooi42rhP" outputId="0e42aab2-bf38-4d4d-b39b-a0c4d63d7fc7"
df_krank = df_c.copy()
age = []
for i in df_krank.itertuples():
age.append(2014-int(i[2]))
df_krank['measurement_age'] = age
colleges = df_krank.loc[(df_krank['tier_name'] == "Selective public") | (df_krank['tier_name'] == "Selective private") |
(df_krank['tier_name'] == "Highly selective public") | (df_krank['tier_name'] == "Highly selective private")|
(df_krank['tier_name'] == "Ivy Plus") | (df_krank['tier_name'] == "Other elite schools (public and private)")]
colleges = colleges[['tier_name','k_rank','measurement_age']]
plt.figure(figsize=(15,9))
plt.grid()
plt.xlabel('Age of Income Measurement',size = 14)
plt.ylabel('Mean Student Income Rank',size = 14)
plt.title('Mean Child Rank vs Age at Income Measurement By College Tier',fontsize=14)
sns.lineplot(x = 'measurement_age', y ='k_rank', hue = 'tier_name', data = colleges)
# + [markdown] id="nsDT_bBC7367"
# ## Mobility Rate
#
# The researchers analyzed the role of colleges for students to progress their income level, especially the students coming from lower quintiles that end up in higher quintiles in their adulthoods. To this end, they derive a new metric named `mobility rate`.
#
# > "The fraction of students who come from bottom quintile and end up in top quintile"
#
# 
#
# In the dataset `kq5_cond_parq1` column stores the success rate of each cohort; while `par_q1` column stores the access rates.
#
# In addition to defining the success rate as P(Child in Q5 | Parent in Q1), the researchers also developed the same rate with respect to student income percentiles: P(Child in **P1** | Parent in Q1) stands for the students who come from bottom quintile and end up in top 1% percentile. And `ktop1pc_cond_parq1` column stores those values for each cohort.
# + [markdown] id="sK0XwYtQ78Pf"
# ### Q1: Calculating the Mobility Rate
#
# In this question, you are going to calculate the mobility rate for each college and then find the top 10 colleges with the highest mobility rates.
#
# - For each cohort, in other words each row, calculate the mobility rate with both `kq5_cond_parq1` and `ktop1pc_cond_parq1` and store them in columns named `mobility_rate_q5` and `mobility_rate_p1`, respectively.
# - `kq5_cond_parq1` * `par_q1` -> `mobility_rate_q5`
# - `ktop1pc_cond_parq1` * `par_q1` -> `mobility_rate_p1`
#
# - Group the dataframe with respect to the colleges and find the mean of `mobility_rate_q5, mobility_rate_p1, kq5_cond_parq1, par_q1` columns.
#
# - First, sort the resulting groups, i.e. colleges, with respect to `mobility_rate_q5` and display the top 10 rows as a dataframe.
#
# - And lastly, sort the resulting groups with respect to `mobility_rate_p1` and display the top 10 rows as a dataframe.
# + id="6Sla1Jcc1fRe"
dfNew = df_c.copy()
dfNew['mobility_rate_q5'] = dfNew['kq5_cond_parq1'] * dfNew['par_q1']
dfNew['mobility_rate_p1'] = dfNew['ktop1pc_cond_parq1'] * dfNew['par_q1']
# + colab={"base_uri": "https://localhost:8080/", "height": 373} id="aIUPVBnaqVDd" outputId="6347721f-54e8-40db-dde3-0032a4033ab5"
df1 = dfNew.copy()
df1 = df1.groupby('name').mean()[['mobility_rate_q5','mobility_rate_p1','kq5_cond_parq1','par_q1']]
df1 = df1.sort_values(by = ['mobility_rate_q5'], ascending=False).head(10)
df1.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 373} id="e1vE0N8SAu3Y" outputId="3e89de4d-b594-4251-b0cc-ca4c6771c721"
df2 = dfNew.copy()
df2 = df2.groupby('name').mean()[['mobility_rate_q5','mobility_rate_p1','kq5_cond_parq1','par_q1']]
df2 = df2.sort_values(by = ['mobility_rate_p1'], ascending=False).head(10)
df2.head(10)
# + [markdown] id="8ynG8WdQEGuk"
# ### Q2: Success vs. Access Rates by College Tier
#
# And finally, you are going to check how success and access rates change for different college tiers. In this question, you are going to focus on two college tiers: `Ivy Plus` and `Two-year for-profit`. In addition, you are going to display this relationship using only the success rate definition 2: P(Child in P1 | Parent in Q1).
#
# - Group the dataframe by college and find the mean of success and access rates for each college.
# - Success rate:
# - Definition 2: P(Child in P1 | Par in Q1) -> ktop1pc_cond_parq1
# - Access rate: P(Par in Q1) -> par_q1
#
# - Display a scatter plot in which access rates are encoded in x-axis and success rates in y-axis.
#
# - At the end, each dot on the figure would represent a college. Highlight `Ivy Plus` and `Two-year for-profit` with distinct color for separability.
#
# The result should look like the figure below.
#
# 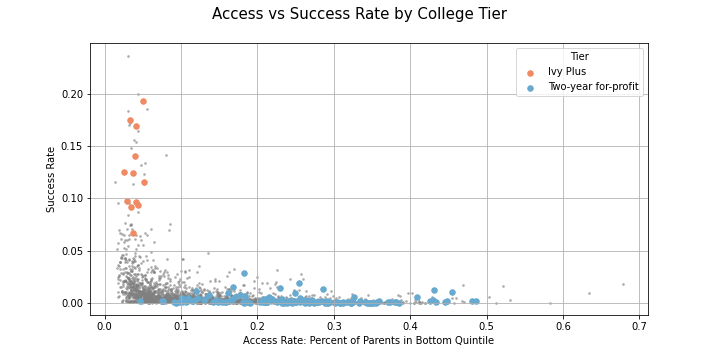
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="3LQ_-aZk_Io0" outputId="410fd336-aec6-4e48-af2d-a6a29261bc9b"
dfX = dfNew.copy()
ivyPlus = dfX.loc[dfX['tier_name'] == 'Ivy Plus']
ivyPlus = ivyPlus.groupby(['name'])
twoYear = dfX.loc[dfX['tier_name'] == 'Two-year for-profit']
twoYear = twoYear.groupby(['name'])
dfX = dfX.groupby('name')
successAll = dfX['ktop1pc_cond_parq1'].mean()
accessAll = dfX['par_q1'].mean()
ivySucess = ivyPlus['ktop1pc_cond_parq1'].mean()
ivyAccess = ivyPlus['par_q1'].mean()
twoSuccess = twoYear['ktop1pc_cond_parq1'].mean()
twoAccess = twoYear['par_q1'].mean()
plt.figure(figsize= (10,5))
plt.grid()
plt.scatter(accessAll, successAll, c = 'gray', s = 2)
plt.scatter(ivyAccess, ivySucess, c = 'orange')
plt.scatter(twoAccess, twoSuccess, c = 'blue')
sns.scatterplot().set_xlabel('Access Rate: Percent of Parents in Bottom Quintile', fontsize = 12)
sns.scatterplot().set_ylabel('Success Rate',fontsize = 12)
plt.title('Access vs Success Rate by College Tier',fontsize= 14)
import matplotlib.patches as mpatch
orangePatch = mpatch.Patch(color = "orange",label = "Ivy Plus")
bluePatch = mpatch.Patch(color = "blue",label = "Two-year for-profit")
plt.legend(handles =[orangePatch,bluePatch])
plt.show()
| EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matplotlib Practice
#
# This notebook offers a set of exercises to different tasks with Matplotlib.
#
# It should be noted there may be more than one different way to answer a question or complete an exercise.
#
# Different tasks will be detailed by comments or text.
#
# For further reference and resources, it's advised to check out the [Matplotlib documentation](https://matplotlib.org/3.1.1/contents.html).
#
# If you're stuck, don't forget, you can always search for a function, for example if you want to create a plot with `plt.subplots()`, search for [`plt.subplots()`](https://www.google.com/search?q=plt.subplots()).
# Import the pyplot module from matplotlib as plt and make sure
# plots appear in the notebook using '%matplotlib inline'
# %matplotlib inline
import matplotlib.pyplot as plt
# Create a simple plot using plt.plot()
plt.plot();
# Plot a single Python list
plt.plot([1,2,3,4]);
# Create two lists, one called X, one called y, each with 5 numbers in them
x = [1, 2, 3, 4, 5]
y = [1, 4, 9, 16, 25]
# Plot X & y (the lists you've created)
plt.plot(x,y);
# There's another way to create plots with Matplotlib, it's known as the object-orientated (OO) method. Let's try it.
# Create a plot using plt.subplots()
fig, ax = plt.subplots();
# Create a plot using plt.subplots() and then add X & y on the axes
fig, ax = plt.subplots()
ax.plot(x, y);
# Now let's try a small matplotlib workflow.
# +
# Import and get matplotlib ready
# %matplotlib inline
import matplotlib.pyplot as plt
# Prepare data (create two lists of 5 numbers, X & y)
X = [1, 2, 3, 4, 5]
y = [11, 22, 33, 44, 55]
# Setup figure and axes using plt.subplots()
fig, ax = plt.subplots()
# Add data (X, y) to axes
ax.plot(X, y)
# Customize plot by adding a title, xlabel and ylabel
ax.set(title='Practice Plot',
xlabel='Practice X',
ylabel='Practice y');
# Save the plot to file using fig.savefig()
fig.savefig('practice-plot.png');
# -
# Okay, this is a simple line plot, how about something a little different?
#
# To help us, we'll import NumPy.
# Import NumPy as np
import numpy as np
# Create an array of 100 evenly spaced numbers between 0 and 100 using NumPy and save it to variable X
X = np.linspace(0, 100, 100)
X
# Create a plot using plt.subplots() and plot X versus X^2 (X squared)
fig, ax = plt.subplots()
ax.plot(X, X**2);
# We'll start with scatter plots.
# Create a scatter plot of X versus the exponential of X (np.exp(X))
fig, ax = plt.subplots()
ax.scatter(X, np.exp(X));
# Create a scatter plot of X versus np.sin(X)
fig, ax = plt.subplots()
ax.scatter(X, np.sin(X));
# How about we try another type of plot? This time let's look at a bar plot. First we'll make some data.
# Create a Python dictionary of 3 of your favourite foods with
# The keys of the dictionary should be the food name and the values their price
fav_food = {'Amla': 100,
'Almond': 500,
'Cashew': 700}
# +
# Create a bar graph where the x-axis is the keys of the dictionary
# and the y-axis is the values of the dictionary
fig, ax = plt.subplots()
ax.bar(fav_food.keys(), fav_food.values())
# Add a title, xlabel and ylabel to the plot
ax.set(title='Food Prices',
xlabel='Favourite Food',
ylabel='Price');
# -
# Make the same plot as above, except this time make the bars go horizontal
fig, ax = plt.subplots()
ax.barh(list(fav_food.keys()), list(fav_food.values()))
ax.set(title='Food Prices',
xlabel='Prices',
ylabel='Fav Food');
# All this food plotting is making me hungry. But we've got a couple of plots to go.
#
# Let's see a histogram.
# +
# Create a random NumPy array of 1000 normally distributed numbers using NumPy and save it to X
X = np.random.randn(1000)
# Create a histogram plot of X
fig, ax = plt.subplots()
ax.hist(X);
# +
# Create a NumPy array of 1000 random numbers and save it to X
X = np.random.rand(1000)
# Create a histogram plot of X
fig, ax = plt.subplots()
ax.hist(X);
# -
# Notice how the distributions (spread of data) are different. Why do they differ?
#
# What else can you find out about the normal distribution?
#
# Can you think of any other kinds of data which may be normally distributed?
#
# These questions aren't directly related to plotting or Matplotlib but they're helpful to think of.
#
# Now let's try make some subplots. A subplot is another name for a figure with multiple plots on it.
# Create an empty subplot with 2 rows and 2 columns (4 subplots total)
fig, ax = plt.subplots(nrows=2, ncols=2);
# Notice how the subplot has multiple figures. Now let's add data to each axes.
# +
# Create the same plot as above with 2 rows and 2 columns and figsize of (10, 5)
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2, figsize=(10, 5))
# Plot X versus X/2 on the top left axes
ax1.plot(X, X/2)
# Plot a scatter plot of 10 random numbers on each axis on the top right subplot
x = np.random.rand(10)
y = np.random.rand(10)
ax2.scatter(x, y)
# Plot a bar graph of the favourite food keys and values on the bottom left subplot
ax3.bar(fav_food.keys(), fav_food.values())
# Plot a histogram of 1000 random normally distributed numbers on the bottom right subplot
nx = np.random.randn(1000)
ax4.hist(nx);
# -
# Woah. There's a lot going on there.
#
# Now we've seen how to plot with Matplotlib and data directly. Let's practice using Matplotlib to plot with pandas.
#
# First we'll need to import pandas and create a DataFrame work with.
# Import pandas as pd
import pandas as pd
# Import the '../data/car-sales.csv' into a DataFame called car_sales and view
car_sales=pd.read_csv('../pandas/car-sales.csv')
car_sales
# Try to plot the 'Price' column using the plot() function
car_sales['Price'].plot();
# Why doesn't it work?
#
# Hint: It's not numeric data.
#
# In the process of turning it to numeric data, let's create another column which adds the total amount of sales and another one which shows what date the car was sold.
#
# Hint: To add a column up cumulatively, look up the cumsum() function. And to create a column of dates, look up the date_range() function.
# Remove the symbols, the final two numbers from the 'Price' column and convert it to numbers
car_sales['Price'] = car_sales['Price'].str.replace('[\$\,]|\.\d*', '').apply(int)
car_sales
# +
# Add a column called 'Total Sales' to car_sales which cumulatively adds the 'Price' column
car_sales['Total Sales'] = np.cumsum(car_sales['Price'])
# Add a column called 'Sale Date' which lists a series of successive dates starting from today (your today)
car_sales['Sale Date'] = pd.date_range(start='16/09/2020', periods=10)
# View the car_sales DataFrame
car_sales
# -
# Now we've got a numeric column (`Total Sales`) and a dates column (`Sale Date`), let's visualize them.
# Use the plot() function to plot the 'Sale Date' column versus the 'Total Sales' column
car_sales.plot(x='Sale Date', y='Total Sales');
# +
# Convert the 'Price' column to the integers
########## Already Done
# Create a scatter plot of the 'Odometer (KM)' and 'Price' column using the plot() function
car_sales.plot.scatter(x='Odometer (KM)', y='Price');
# +
# Create a NumPy array of random numbers of size (10, 4) and save it to X
X = np.random.rand(10, 4)
# Turn the NumPy array X into a DataFrame with columns called ['a', 'b', 'c', 'd']
df = pd.DataFrame(X, columns=['a', 'b', 'c', 'd'])
# Create a bar graph of the DataFrame
df.plot.bar();
# -
# Create a bar graph of the 'Make' and 'Odometer (KM)' columns in the car_sales DataFrame
car_sales.plot.bar(x='Make', y='Odometer (KM)');
# Create a histogram of the 'Odometer (KM)' column
car_sales['Odometer (KM)'].plot(kind='hist');
# Create a histogram of the 'Price' column with 20 bins
car_sales['Price'].plot(kind='hist', bins=20);
# Now we've seen a few examples of plotting directly from DataFrames using the `car_sales` dataset.
#
# Let's try using a different dataset.
# Import "../data/heart-disease.csv" and save it to the variable "heart_disease"
heart_disease = pd.read_csv('../heart-disease.csv')
heart_disease.head()
# View the first 10 rows of the heart_disease DataFrame
heart_disease.head(10)
# Create a histogram of the "age" column with 50 bins
heart_disease['age'].plot.hist(bins=50);
# Call plot.hist() on the heart_disease DataFrame and toggle the
# "subplots" parameter to True
heart_disease.plot.hist(subplots=True);
# That plot looks pretty squished. Let's change the figsize.
# Call the same line of code from above except change the "figsize" parameter
# to be (10, 30)
heart_disease.plot.hist(subplots=True, figsize=(10, 30));
# Now let's try comparing two variables versus the target variable.
#
# More specifially we'll see how age and cholesterol combined effect the target in **patients over 50 years old**.
#
# For this next challenge, we're going to be replicating the following plot:
#
# <img src="../images/matplotlib-heart-disease-chol-age-plot.png"/>
# +
# Replicate the above plot in whichever way you see fit
# Note: The method below is only one way of doing it, yours might be
# slightly different
# Create DataFrame with patients over 50 years old
over_50 = heart_disease[heart_disease['age'] > 50]
# Create the plot
fig, ax = plt.subplots(figsize=(10, 5))
# Plot the data
scattey = ax.scatter(x=over_50['age'],
y=over_50['chol'],
c=over_50['target'])
# Customize the plot
ax.set(title='Practice Plot 3',
xlabel='Age',
ylabel='Cholesterol')
ax.legend(*scattey.legend_elements(), title='Target')
# Add a meanline
ax.axhline(y=over_50['chol'].mean(), linestyle='--');
# -
# Beatiful, now you've created a plot of two different variables, let's change the style.
# Check what styles are available under plt
plt.style.available
# Change the style to use "seaborn-whitegrid"
plt.style.use('seaborn-whitegrid')
# Now the style has been changed, we'll replot the same figure from above and see what it looks like.
#
# If you've changed the style correctly, it should look like the following:
# <img src="../images/matplotlib-heart-disease-chol-age-plot-seaborn-whitegrid.png"/>
#
# +
# Reproduce the same figure as above with the "seaborn-whitegrid" style
# Create the plot
fig, ax = plt.subplots(figsize=(10, 5))
# Plot the data
scattey = ax.scatter(x=over_50['age'],
y=over_50['chol'],
c=over_50['target'])
# Customize the plot
ax.set(title='Practice Plot 4',
xlabel='Age',
ylabel='Cholesterol')
ax.legend(*scattey.legend_elements(), title='Target')
ax.set_xlim([50, 80])
# Add a meanline
ax.axhline(y=over_50['chol'].mean(), linestyle='--');
# -
# Wonderful, you've changed the style of the plots and the figure is looking different but the dots aren't a very good colour.
#
# Let's change the `cmap` parameter of `scatter()` as well as the `color` parameter of `axhline()` to fix it.
#
# Completing this step correctly should result in a figure which looks like this:
# <img src="../images/matplotlib-heart-disease-chol-age-plot-cmap-change.png"/>
# +
# Replot the same figure as above except change the "cmap" parameter
# of scatter() to "winter"
# Also change the "color" parameter of axhline() to "red"
# Create the plot
fig, ax = plt.subplots(figsize=(10, 5))
# Plot the data
scattey = ax.scatter(x=over_50['age'],
y=over_50['chol'],
c=over_50['target'],
cmap='winter')
# Customize the plot
ax.set(title='Practice Plot 4',
xlabel='Age',
ylabel='Cholesterol')
ax.legend(*scattey.legend_elements(), title='Target')
ax.set_xlim([50, 80])
# Add a meanline
ax.axhline(y=over_50['chol'].mean(), linestyle='--', c='red');
# -
# Beautiful! Now our figure has an upgraded color scheme let's save it to file.
# Save the current figure using savefig(), the file name can be anything you want
fig.savefig('practice-plot-final.png');
# Reset the figure by calling plt.subplots()
fig, ax = plt.subplots()
# ## Extensions
#
# For more exercises, check out the [Matplotlib tutorials page](https://matplotlib.org/3.1.1/tutorials/index.html). A good practice would be to read through it and for the parts you find interesting, add them into the end of this notebook.
#
# The next place you could go is the [Stack Overflow page for the top questions and answers for Matplotlib](https://stackoverflow.com/questions/tagged/matplotlib?sort=MostVotes&edited=true). Often, you'll find some of the most common and useful Matplotlib functions here. Don't forget to play around with the Stack Overflow filters! You'll likely find something helpful here.
#
# Finally, as always, remember, the best way to learn something new is to try it. And try it relentlessly. Always be asking yourself, "is there a better way this data could be visualized so it's easier to understand?"
| matplotlib/matplotlib-exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
###########IMPORTS############
from tensorflow.python.framework import constant_op
from tensorflow.python.framework import dtypes
from tensorflow.python.framework import ops
from tensorflow.python.ops import array_ops
from tensorflow.python.ops import control_flow_ops
from tensorflow.python.ops import functional_ops
from tensorflow.python.ops import math_ops
from tensorflow.python.ops import tensor_array_ops
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
###########INTEGRATOR FRAMEWORK############
#1 Constraint Checks
def _check_input_types(t, y0): # Ensure input is Correct
if not (y0.dtype.is_floating or y0.dtype.is_complex):
raise TypeError('`y0` must have a floating point or complex floating point dtype')
if not t.dtype.is_floating:
raise TypeError('`t` must have a floating point dtype')
def _assert_increasing(t): # Check Time is Monotonous
assert_increasing = control_flow_ops.Assert(math_ops.reduce_all(t[1:] > t[:-1]), ['`t` must be monotonic increasing'])
return ops.control_dependencies([assert_increasing])
#2 Integrator Class
class _Integrator():
def integrate(self, evol_func, y0, time_grid): # iterator
time_delta_grid = time_grid[1:] - time_grid[:-1]
scan_func = self._make_scan_func(evol_func)
y_grid = functional_ops.scan(scan_func, (time_grid[:-1], time_delta_grid),y0)
return array_ops.concat([[y0], y_grid], axis=0)
def _make_scan_func(self, evol_func): # stepper function
def scan_func(y, t_dt):
if n_>0:
t,dt = t_dt
dy = self._step_func(evol_func, t, dt, y)
dy = math_ops.cast(dy, dtype=y.dtype)
out = y + dy
## Operate on non-integral
ft = y[-n_:]
l = tf.zeros(tf.shape(ft),dtype=ft.dtype)
l_ = t-ft
z = tf.less(y[:n_],F_b)
z_ = tf.greater_equal(out[:n_],F_b)
df = tf.where(tf.logical_and(z,z_),l_,l)
ft_ = ft+df
return tf.concat([out[:-n_],ft_],0)
else:
t, dt = t_dt
dy = self._step_func(evol_func, t, dt, y)
dy = math_ops.cast(dy, dtype=y.dtype)
return y + dy
return scan_func
def _step_func(self, evol_func, t, dt, y):
k1 = evol_func(y, t)
half_step = t + dt / 2
dt_cast = math_ops.cast(dt, y.dtype)
k2 = evol_func(y + dt_cast * k1 / 2, half_step)
k3 = evol_func(y + dt_cast * k2 / 2, half_step)
k4 = evol_func(y + dt_cast * k3, t + dt)
return math_ops.add_n([k1, 2 * k2, 2 * k3, k4]) * (dt_cast / 6)
#3 Integral Caller
def odeint_fixed(func, y0, t):
t = ops.convert_to_tensor(t, preferred_dtype=dtypes.float64, name='t')
y0 = ops.convert_to_tensor(y0, name='y0')
_check_input_types(t, y0)
with _assert_increasing(t):
return _Integrator().integrate(func, y0, t)
# +
#### ALL EQUATIONS AND PARAMETERS TAKEN FROM <NAME> et. al. ####
#### Values for PNs are used here ####
T = 26 # Temperature
n_n = 4 # number of neurons
p_n = 2 # number of PNs
l_n = 2 # number of LNs
t = np.arange(0.0, 500, 0.01) # duration of simulation
C_m = [1.0]*n_n # n_n x 1 vector for capacitance
g_Na = [100.0]*n_n # n_n x 1 vector for sodium conductance
g_K = [10.0]*n_n # n_n x 1 vector for potassium conductance
g_L = [0.15]*n_n # n_n x 1 vector for leak conductance
g_KL = [0.05]*n_n # n_n x 1 vector for K leak conductance
g_A = [10.0]*n_n # n_n x 1 vector for Transient K conductance
E_Na = [50.0]*n_n # n_n x 1 vector for Na Potential
E_K = [-95.0]*n_n # n_n x 1 vector for K Potential
E_L = [-55.0]*n_n # n_n x 1 vector for Leak Potential
E_KL = [-95.0]*n_n # n_n x 1 vector for K Leak Potential
E_A = [-95.0]*n_n # n_n x 1 vector for Transient K Potential
F_b = [0.0]*n_n # n_n x 1 vector for fire potential
inp = [0.0,15.0,0.0] # External Current Inputs
# ACETYLCHOLINE
ach_mat = np.array([[0.0,0.0,0.0],
[0.0,0.0,0.0],
[0.0,1.0,0.0]])
n_syn_ach = int(np.sum(ach_mat)) # Number of Acetylcholine (Ach) Synapses
alp_ach = [10.0]*n_syn_ach # Alpha for Ach Synapse
bet_ach = [0.2]*n_syn_ach # Beta for Ach Synapse
t_max = 0.3 # Maximum Time for Synapse
t_delay = 0 # Axonal Transmission Delay
A = [0.5]*n_n # Synaptic Response Strength
g_ach = [1.0]*n_n # Ach Conductance
E_ach = [0.0]*n_n # Ach Potential
# FAST GABA
fgaba_mat = np.array([[0.0,0.0,0.0],
[0.0,0.0,1.0],
[0.0,0.0,0.0]])
n_syn_fgaba = int(np.sum(fgaba_mat)) # Number of Fast GABA (fGABA) Synapses
alp_fgaba = [10.0]*n_syn_fgaba # Alpha for fGABA Synapse
bet_fgaba = [0.16]*n_syn_fgaba # Beta for fGABA Synapse
V0 = [-20.0]*n_n # Decay Potential
sigma = [1.5]*n_n # Decay Time Constant
g_fgaba = [0.8]*n_n # fGABA Conductance
E_fgaba = [-70.0]*n_n # fGABA Potential
phi = 3.0**((22-T)/10)
def Na_prop(V):
V_ = V-(-50)
alpha_m = 0.32*(13.0 - V_)/(tf.exp((13.0 - V_)/4.0) - 1.0)
beta_m = 0.28*(V_ - 40.0)/(tf.exp((V_ - 40.0)/5.0) - 1.0)
alpha_h = 0.128*tf.exp((17.0 - V_)/18.0)
beta_h = 4.0/(tf.exp((40.0 - V_)/5.0) + 1.0)
t_m = 1.0/((alpha_m+beta_m)*phi)
t_h = 1.0/((alpha_h+beta_h)*phi)
return alpha_m*t_m, t_m, alpha_h*t_h, t_h
def K_prop(V):
V_ = V-(-50)
alpha_n = 0.02*(15.0 - V_)/(tf.exp((15.0 - V_)/5.0) - 1.0)
beta_n = 0.5*tf.exp((10.0 - V_)/40.0)
t_n = 1.0/((alpha_n+beta_n)*phi)
return alpha_n*t_n, t_n
def m_a_inf(V):
return 1/(1+tf.exp(-(V+60.0)/8.5))
def h_a_inf(V):
return 1/(1+tf.exp((V+78.0)/6.0))
def tau_m_a(V):
return 1/(tf.exp((V+35.82)/19.69) + tf.exp(-(V+79.69)/12.7) + 0.37) / phi
def tau_h_a(V):
return tf.where(tf.less(V,-63),1/(tf.exp((V+46.05)/5) + tf.exp(-(V+238.4)/37.45)) / phi,19.0 / phi * tf.ones(tf.shape(V),dtype=V.dtype))
def m_Ca_inf(V):
return 1/(1+tf.exp(-(V+20.0)/6.5))
def h_Ca_inf(V):
return 1/(1+tf.exp((V+25.0)/12))
def tau_m_Ca(V):
return 1.5
def tau_h_Ca(V):
return 0.3*tf.exp((V-40.0)/13.0) + 0.002*tf.exp((60.0-V)/29)
# NEURONAL CURRENTS
def I_Na(V, m, h):
return g_Na * m**3 * h * (V - E_Na)
def I_K(V, n):
return g_K * n**4 * (V - E_K)
def I_L(V):
return g_L * (V - E_L)
def I_KL(V):
return g_KL * (V - E_KL)
def I_A(V, m, h):
return g_A * m**4 * h * (V - E_A)
# SYNAPTIC CURRENTS
def I_ach(o,V):
o_ = tf.Variable([0.0]*n_n**2,dtype=tf.float64)
ind = tf.boolean_mask(tf.range(n_n**2),ach_mat.reshape(-1) == 1)
o_ = tf.scatter_update(o_,ind,o)
o_ = tf.reshape(o_,(n_n,n_n))
return tf.reduce_sum(g_ach*o_*(V-E_ach),1)
def I_fgaba(o,V):
o_ = tf.Variable([0.0]*n_n**2,dtype=tf.float64)
ind = tf.boolean_mask(tf.range(n_n**2),fgaba_mat.reshape(-1) == 1)
o_ = tf.scatter_update(o_,ind,o)
o_ = tf.reshape(o_,(n_n,n_n))
return tf.reduce_sum(g_fgaba*o_*(V-E_fgaba),1)
def I_inj_t(t):
return tf.where(tf.logical_and(tf.greater(t,100),tf.less(t,400)),tf.constant(inp,dtype=tf.float64),tf.constant([0]*n_n,dtype=tf.float64))
# -
def dAdt(X, t):
V = X[0:n_n]
m = X[n_n:2*n_n]
h = X[2*n_n:3*n_n]
n = X[3*n_n:4*n_n]
m_a = X[4*n_n:5*n_n]
h_a = X[5*n_n:6*n_n]
m_a = X[6*n_n:5*n_n]
h_a = X[5*n_n:6*n_n]
o_ach = X[6*n_n:6*n_n+n_syn_ach]
o_fgaba = X[6*n_n+n_syn_ach:6*n_n+n_syn_ach+n_syn_fgaba]
fire_t = X[-n_n:]
dVdt = (I_inj_t(t) - I_Na(V, m, h) - I_K(V, n) - I_A(V, m_a, h_a) - I_L(V) - I_KL(V) - I_ach(o_ach,V) - I_fgaba(o_fgaba,V)) / C_m
m0,tm,h0,th = Na_prop(V)
n0,tn = K_prop(V)
dmdt = - (1.0/tm)*(m-m0)
dhdt = - (1.0/th)*(h-h0)
dndt = - (1.0/tn)*(n-n0)
dm_adt = - (1.0/tau_m_a(V))*(m_a-m_a_inf(V))
dh_adt = - (1.0/tau_h_a(V))*(h_a-h_a_inf(V))
A_ = tf.constant(A,dtype=tf.float64)
T_ach = tf.where(tf.logical_and(tf.greater(t,fire_t+t_delay),tf.less(t,fire_t+t_max+t_delay)),A_,tf.zeros(tf.shape(A_),dtype=A_.dtype))
T_ach = tf.multiply(tf.constant(ach_mat,dtype=tf.float64),T_ach)
T_ach = tf.boolean_mask(tf.reshape(T_ach,(-1,)),ach_mat.reshape(-1) == 1)
do_achdt = alp_ach*(1.0-o_ach)*T_ach - bet_ach*o_ach
T_fgaba = 1.0/(1.0+tf.exp(-(V-V0)/sigma))
T_fgaba = tf.multiply(tf.constant(fgaba_mat,dtype=tf.float64),T_fgaba)
T_fgaba = tf.boolean_mask(tf.reshape(T_fgaba,(-1,)),fgaba_mat.reshape(-1) == 1)
do_fgabadt = alp_fgaba*(1.0-o_fgaba)*T_fgaba - bet_fgaba*o_fgaba
dfdt = tf.zeros(tf.shape(fire_t),dtype=fire_t.dtype)
out = tf.concat([dVdt,dmdt,dhdt,dndt,dm_adt,dh_adt,do_achdt,do_fgabadt,dfdt],0)
return out
global n_
n_ = n_n
state_vector = [-65]*n_n + [0.05]*n_n + [0.6]*n_n + [0.32]*n_n+ [0.05]*n_n + [0.6]*n_n + [0]*(n_syn_ach) + [0]*(n_syn_fgaba) +[-500]*n_n
init_state = tf.constant(state_vector, dtype=tf.float64)
tensor_state = odeint_fixed(dAdt, init_state, t)
# %%time
with tf.Session() as sess:
tf.global_variables_initializer().run()
state = sess.run(tensor_state)
fig, ax = plt.subplots(3, 1,figsize=(5,5),sharex=True,sharey=True)
for n,i in enumerate(ax):
i.plot(t[:],state[:,n])
plt.tight_layout()
plt.show()
| archive/NetworkFlow - PN only.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### summary
# - pandas
# - 데이터 분석 : 데이터 전처리 파트에서 많이 사용하는 패키지
# - 테이블 형테의 데이터를 처리할때 사용하는 python 라이브러리
# - Series : index, value 로 이루어지며, 한가지 데이터 타입만 가질수 있음
# - 생성, 선택, 수정 방법
# - DataFrame : index, value, column으로 이루어지며 Series들의 집합
# - 생성 방법 1 : 딕셔너리의 리스트 {[],[]} -> 리스트 -> 컬럼 데이터
# - 생성 방법 2 : 리스트의 딕셔너리[{},{}] -> 딕셔너리 -> 로우 데이터
# - row 선택 : df.loc[idx]
# - column 선택 : df[column name]
# - row, column 선택 : df.loc[idx, column]
# - 함수
# - apply, append, concat,
# - groupby, merge
import makedata
makedata.get_age(), makedata.get_name()
makedata.make_data()
# #### quiz
# - makedata 모듈을 이용해서 데이터 프레임 만들기
# - user_df
# - 8명의 데이터
# - UserID
# - Name : makedata.get_name()
# - Age : makedata.get_age()
# - 중복되는 Name 값이 없도록
# +
# 딕셔너리의 리스트 : UserID, Name,Age
datas = {}
datas["UserID"] = list(range(1, 9))
datas["Age"] = [makedata.get_age() for _ in range(8)]
names = []
while True:
name = makedata.get_name()
if name not in names:
names.append(name)
if len(names) >= 8:
break
datas["Name"] = names
user_df = pd.DataFrame(datas)
user_df
# +
# 리스트의 딕셔너리
# row데이터를 넣기위해 컬럼부터 생성
user_df = pd.DataFrame(columns = ["UserID", "Name", "Age"])
datas = []
for idx in range(1, 9):
name = makedata.get_name()
# user_df["Name"]에 생성된 name이 있으면 다시 돌림.
while name in list(user_df["Name"]) :
name = makedata.get_name()
data = {"Name": name, "UserID": idx, "Age": makedata.get_age()}
# 딕셔너리로 맨 마지막 행을 넣는 법
#loc[] 에 idx를 넣어야 하는데
#loc[len(user_df)]를 하면 user_df 갯수의 인덱스에 생성된 데이터가 들어감
user_df.loc[len(user_df)] = data
datas.append(data)
user_df
# -
# ### quiz
# - money_df 만들기
# - 15개의 데이터가 있습니다.
# - ID : 1 ~ 8 까지의 랜덤한 숫자 데이터
# - Money : 1000원 단위로 1000원 ~ 20000원 까지의 숫자가 저장
# +
# 딕셔너리 데이터를 데이터 프레임에 하나씩 추가
money_df = pd.DataFrame(columns=["ID", "Money"])
np.random.randint(1,9)
for _ in range(15) :
money_df.loc[len(money_df)] = {
"ID" : np.random.randint(1,9),
"Money" : np.random.randint(1,21) * 1000,
}
# 컬럼데이터에서 Unique값 확인 # 1번과 4번이없게 만듬
# 일부러 데이터가 빠질때까지 생성
ids = money_df["ID"].unique()
# 보기좋게 내림차순
ids.sort()
ids
# -
money_df.tail(1)
user_df.tail(1)
# ### 1. Merge
# - 1 : 1로 테이블을 만든 뒤, 같은 데이터들끼리 필터링 해줌
user_df.merge(money_df, left_on = "UserID", right_on="ID")
# 컬럼명 변경
user_df.rename(columns={"UserID" : "ID"}, inplace = True)
user_df.tail(1)
# 컬럼명이 동일하면 left_on right_on 설정을 안해주어도 된다.
user_df.merge(money_df).tail(2)
# pd.객체의 merge함수를 사용하여 생성 가능
result_df = pd.merge(money_df, user_df) # 순서에 따라 붙음
result_df.tail(2)
# groupby : sum, size, min .. 함수를 사용하여 그룹바이 해줌, Series
# inner merge 이기에 없는건 안나옴
# Series 이기에 더해주는 값 등은 1개만 올수있음 ex) money
money_list = result_df.groupby("Name").sum()["Money"].reset_index()
money_list
# groupby : agg("sum"), agg("mean"), ... : DataFrame
# 데이터프레임이기에 더해주는 값은 여러개 가능함 money뒤에 "Age" 넣어주면됨
money_list = result_df.groupby("Name").agg("sum").reset_index()[["Name", "Money"]]
money_list
# +
# merge : mnoney_list, user_df : outer
# outer 조인으로 없는 데이터도 조인이 되게 할수있음
# -
result = pd.merge(user_df, money_list, how = "outer")
result
# +
# fillna : Nan을 특정 데이터로 채워줌 (결측치 처리)
# -
# fillna를 사용하여 결측치를 0으로 변경해줌
# 데이터 프레임을 연산하는 과정을 거치면 실수타입으로 변환이됨
result.fillna(value= 0, inplace=True)
result
# money 컬럼을 정수 데이터 타입으로 변경
result.dtypes # money가 float64임
# money 컬럼을 정수 데이터 타입으로 변경
result["Money"] = result["Money"].astype("int")
result
# Money가 많은 사람으로 정렬
result.sort_values("Money", ascending=False)
# 상위 3명의 평균값
np.average(result.sort_values("Money", ascending=False)[:3]["Money"])
| python/numpy_pandas/04_pandas2_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import matplotlib.pyplot as plt
# + pycharm={"name": "#%%\n"}
with open('../data/southwest_images_new_test.pkl', 'rb') as train_f:
saved_southwest_dataset_train = pickle.load(train_f)
print(saved_southwest_dataset_train.shape)
# + pycharm={"name": "#%%\n"}
for i in range(20):
plt.figure()
plt.imshow(saved_southwest_dataset_train[i])
plt.show()
| plots/preprocessing/load_southwest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv_pyadlml
# language: python
# name: venv_pyadlml
# ---
# ## UCI ADL Binary
#
# <NAME>.; <NAME>.; <NAME>. Activity Recognition Using Hybrid Generative/Discriminative Models on Home Environments Using Binary Sensors. Sensors 2013, 13, 5460-5477.
#
# ### Description
# This dataset comprises information regarding the ADLs performed by two users on a daily basis in their
# own homes. This dataset is composed by two instances of data, each one corresponding to a different
# user and summing up to 35 days of fully labelled data. Each instance of the dataset is described by
# three text files, namely: description, sensors events (features), activities of the daily living (labels).
# Sensor events were recorded using a wireless sensor network and data were labelled manually.
#
# ### House
# Home setting: 4 rooms house
# Number of labelled days: 14 days
# Labels (ADLs included): Leaving, Toileting, Showering, Sleeping, Breakfast, Lunch, Dinner, Snack, Spare_Time/TV, Grooming
# Number of sensors: 12 sensors
# Sensors: PIR: Shower, Basin, Cooktop
# Magnetic: Maindoor, Fridge, Cabinet, Cupboard
# Flush: Toilet
# Pressure: Seat, Bed
# Electric: Microwave, Toaster
#
# +
import sys
sys.path.append("../..")
import pandas as pd
import matplotlib.pyplot as plt
from pyadlml.dataset import set_data_home, fetch_uci_adl_binary
set_data_home('/home/chris/code/adlml/data_home')
subject = 'OrdonezA'
data = fetch_uci_adl_binary(keep_original=True, cache=True, subject=subject)
# + active=""
# import plotly
# plotly.offline.init_notebook_mode(connected=True)
# -
data.df_activities.head()
data.df_devices.head()
# ## Activities
print('There were ', len(data.df_activities['activity'].unique()), 'activities recorded')
# + active=""
# from pyadlml.dataset.plotly.activities import gantt
# gantt(data.df_activities).show()
# +
from pyadlml.dataset.plot.activities import hist_counts
idle = True
hist_counts(data.df_activities, idle=idle);
# +
from pyadlml.dataset.plot.activities import boxplot_duration
idle = False
boxplot_duration(data.df_activities, idle=idle);
# +
from pyadlml.dataset.plot.activities import hist_cum_duration
idle = True
y_scale = 'log'
hist_cum_duration(data.df_activities, y_scale=y_scale, idle=idle);
# + active=""
# from pyadlml.dataset.plot.activities import ridge_line
# #Parameters
# n = 100 # number of samples to draw to approximate distribution
# # TODO change the calculation by mapping each interval onto a second grid with 1's for stuff
# # in the interval and 0's outside. Add all intervals on each other to get the histogram
# idle = False
# ridge_line(data.df_activities, n=n, idle=idle)
# -
# #### Transition Heatmap
# $x_{72}$ should be read as first used the toilet and after that went to bed for a total of 46 times
# +
from pyadlml.dataset.plot.activities import heatmap_transitions
idle = False
z_scale = None #'log'
heatmap_transitions(data.df_activities, z_scale=z_scale, idle=idle);
# -
# ## Devices
print('There are', len(data.df_devices['device'].unique()), 'devices recorded.')
from pyadlml.dataset.devices import device_rep1_2_rep2
rep2_nodrop, syn_acts = device_rep1_2_rep2(data.df_devices, drop=False)
rep2_drop = device_rep1_2_rep2(data.df_devices, drop=True)
from pyadlml.dataset.plot.devices import hist_on_off
hist_on_off(rep2_nodrop);
# As we can see a lot of devices are off the whole time.
from pyadlml.dataset.plot.devices import boxplot_on_duration
boxplot_on_duration(rep2_nodrop);
from pyadlml.dataset.plot.devices import heatmap_trigger_one_day
heatmap_trigger_one_day(data.df_devices, t_res='5m');
# +
from pyadlml.dataset.plot.devices import heatmap_trigger_time
t_window = '5s'
z_scale = 'log'
heatmap_trigger_time(data.df_devices, t_window=t_window, z_scale=z_scale);
# -
from pyadlml.dataset.plot.devices import heatmap_cross_correlation
heatmap_cross_correlation(data.df_devices);
from pyadlml.dataset.plot.devices import hist_trigger_time_diff
hist_trigger_time_diff(data.df_devices);
# + tags=[]
from pyadlml.dataset.plot.devices import hist_counts
hist_counts(data.df_devices);
# -
# # Activites ~ Devices
#
#
# ### Triggers
# +
from pyadlml.dataset.plot import heatmap_contingency_triggers
idle = True
z_scale = None#'log'
heatmap_contingency_triggers(data.df_devices, data.df_activities, idle=idle, z_scale=z_scale);
# +
from pyadlml.dataset.plot import heatmap_contingency_triggers_01
idle = True
z_scale = None#'log'
heatmap_contingency_triggers_01(data.df_devices, data.df_activities, idle=idle, z_scale=z_scale);
# -
# ## Duration
#
# the overlap of the intervals is also interesting.
# +
from pyadlml.dataset.plot import heatmap_contingency_overlaps
z_scale='log'
idle=True
heatmap_contingency_overlaps(data.df_devices, data.df_activities, z_scale=z_scale, idle=idle)
# -
# # Representations
#
# There are different ways to represent our data. One way is to discretize the data into intervals.
# - RAW representation _____|^^^^^^|______
# is 1 whenever the sensor is 'on' and 0 otherwise
#
# - Changepoint representation ______|________|______
# sensor gives a 1 to timeslices where the sensor reading changes
#
# - LastFired representation ______|^^^^^^^^^
# last sensor that changed state gives cont. 1
#
# - Lagged representation
# the
# +
import sys
sys.path.append("../..")
import pandas as pd
import matplotlib.pyplot as plt
from pyadlml.dataset import set_data_home, fetch_uci_adl_binary
set_data_home('/home/chris/code/adlml/data_home')
subject = 'OrdonezA'
data = fetch_uci_adl_binary(keep_original=True, cache=True, subject=subject)
# -
# ## RAW - representation
# +
from pyadlml.preprocessing import DiscreteEncoder
enc_raw = DiscreteEncoder(rep='raw')
raw = enc_raw.fit_transform(data.df_devices)
# +
from pyadlml.preprocessing import LabelEncoder
enc_lbl = LabelEncoder(raw)
lbls = enc_lbl.fit_transform(data.df_activities)
# + active=""
# raw
# + active=""
# t_res = '60s'
# idle = True
# st='int_coverage'
#
# enc_raw = RawEncoder(t_res=t_res, sample_strat=st)
# raw = enc_raw.fit_transform(data.df_devices)
#
# enc_lbl = LabelEncoder(raw, idle=idle)
# lbls = enc_lbl.fit_transform(data.df_activities)
# +
def check_raw(raw):
# should not contain any nan values
assert not raw.isnull().values.any()
check_raw(raw)
# + active=""
# raw
# + tags=[]
from pyadlml.dataset.plot.raw import hist_activities
hist_activities(enc_lbl.inverse_transform(lbls), scale='log', figsize=(10,8))
# +
from pyadlml.dataset.plot.raw import heatmap_contingency_01
z_scale = 'log'
rep = 'raw'
heatmap_contingency_01(raw, enc_lbl.inverse_transform(lbls), rep=rep, z_scale=z_scale);
# +
from pyadlml.dataset.plot.raw import heatmap_cross_correlation
heatmap_cross_correlation(raw);
# +
# TODO sth. is wrong here
#from pyadlml.dataset.plot.raw import corr_devices_01
#corr_devices_01(raw)
# -
# ## Changepoint - representation
# Raw: _____|^^^^^^|______
#
# CP : ______|________|______
#
# sensor gives a 1 to timeslices where the sensor reading changes
# +
idle = True
t_res = '60s'
rep = 'changepoint'
cp_enc = DiscreteEncoder(rep='changepoint', t_res=t_res)
cp = cp_enc.fit_transform(data.df_devices)
# +
from pyadlml.preprocessing import LabelEncoder
enc_lbl = LabelEncoder(cp, idle=idle)
lbls = enc_lbl.fit_transform(data.df_activities)
# +
def check_changepoint(cp):
# should not contain any nan values
assert not cp.isnull().values.any()
check_changepoint(cp)
# +
from pyadlml.dataset.plot.raw import heatmap_contingency_01
z_scale = 'log'
rep = 'Changepoint'
heatmap_contingency_01(cp, enc_lbl.inverse_transform(lbls), rep=rep, z_scale=z_scale);
# +
from pyadlml.dataset.plot.raw import heatmap_cross_correlation
heatmap_cross_correlation(cp);
# + active=""
# from pyadlml.dataset.plot.raw import corr_devices_01
# corr_devices_01(cp)
# -
# ## LastFired - representation
# ______|^^^^^^^^^
#
# last sensor that changed state gives cont. 1
# +
t_res = '60s'
idle=True
rep = 'lastfired'
lf_enc = DiscreteEncoder(rep=rep, t_res=t_res)
lf = lf_enc.fit_transform(data.df_devices)
# +
from pyadlml.preprocessing import LabelEncoder
enc_lbl = LabelEncoder(lf, idle=idle)
lbls = enc_lbl.fit_transform(data.df_activities)
# +
def check_lastfired(lf):
# should not contain any nan values
assert not lf.isnull().values.any()
# each row has to have exactly one device being on
for row in lf.iterrows():
assert 1 == row[1].values.sum()
check_lastfired(lf)
# +
from pyadlml.dataset.plot.raw import heatmap_contingency_01
z_scale = 'log'
rep = 'LastFired'
heatmap_contingency_01(lf, enc_lbl.inverse_transform(lbls), rep=rep, z_scale=z_scale);
# +
from pyadlml.dataset.plot.raw import heatmap_cross_correlation
heatmap_cross_correlation(lf);
# + active=""
# from pyadlml.dataset.plot.raw import corr_devices_01
# corr_devices_01(lf)
# -
# ## Image - representation
#
# a lot of timeseries can be represented as images. There are good algorithms for images available. There are Recurrence Plots (RP), Gramian Angular Field (GAF) (bad for binary data), Markov Transition Field (MTF): https://doi.org/10.3390/electronics9010068.
#
#
# Gramian Angular field:
# https://medium.com/analytics-vidhya/encoding-time-series-as-images-b043becbdbf3
#
# https://pyts.readthedocs.io/en/stable/auto_examples/image/plot_mtf.html#sphx-glr-auto-examples-image-plot-mtf-py
# ### Lagged RAW
#
# The lagged raw representation generates a 3D tensor from the raw input. A state at time $t$ is $1D$ vector. The window size determines how big the $2D$ picture is. Stack the windows in the third dimension.
#
# Also see last part of
# https://towardsdatascience.com/ml-approaches-for-time-series-4d44722e48fe
# +
from pyadlml.preprocessing import ImageEncoder, ImageLabelEncoder
window_size = 20
t_res = '2min'
sample_strat = 'int_coverage'
idle = True
rep = 'raw'
enc_lgd_raw = ImageEncoder(rep, window_size,
t_res=t_res,
sample_strat=sample_strat)
images = enc_lgd_raw.fit_transform(data.df_devices)
# -
images.shape
# +
enc_lgd_lbl = ImageLabelEncoder(data.df_devices,
window_size,
t_res=t_res,
idle=idle)
labels = enc_lgd_lbl.fit_transform(data.df_activities)
labels.shape
# +
from pyadlml.dataset.plot.image import mean_image
mean_image(images, list(data.df_devices['device'].unique()))
# +
from pyadlml.dataset.plot.image import mean_image_per_activity
X = images
y = enc_lgd_lbl.inverse_transform(labels)
mean_image_per_activity(X, y, list(data.df_devices['device'].unique()))
# -
# # Lagged Changepoint representation
# +
from pyadlml.preprocessing import LaggedChangepointEncoder
window_size = 20
t_res = '2min'
sample_strat = 'int_coverage'
idle = True
enc_lgd_cp = LaggedChangepointEncoder(window_size, t_res=t_res)
images = enc_lgd_cp.fit_transform(data.df_devices)
images.shape
# +
from pyadlml.dataset.plot.image import mean_image
mean_image(images, data.devices)
# +
enc_lgd_lbl = LaggedLabelEncoder(data.df_devices,
window_size,
t_res=t_res,
idle=idle)
labels = enc_lgd_lbl.fit_transform(data.df_activities)
labels.shape
# +
from pyadlml.dataset.plot.image import mean_image_per_activity
X = images
y = enc_lgd_lbl.inverse_transform(labels)
mean_image_per_activity(X, y, data.devices)
# -
# ## Lagged Last Fired Encoder
# +
from pyadlml.preprocessing import LaggedLastFiredEncoder
window_size = 20
t_res = '2min'
sample_strat = 'int_coverage'
idle=True
enc_lgd_lf = LaggedLastFiredEncoder(window_size, t_res=t_res)
images = enc_lgd_lf.fit_transform(data.df_devices)
images.shape
# +
from pyadlml.dataset.plot.image import mean_image
mean_image(images, data.devices)
# +
enc_lgd_lbl = LaggedLabelEncoder(data.df_devices,
window_size,
t_res=t_res,
idle=idle)
labels = enc_lgd_lbl.fit_transform(data.df_activities)
labels.shape
# +
from pyadlml.dataset.plot.image import mean_image_per_activity
X = images
y = enc_lgd_lbl.inverse_transform(labels)
mean_image_per_activity(X, y, data.devices)
# -
| notebooks/datasets/uci_adl_binary_subjectA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.0 Intro to Financial Concepts using Python
import numpy as np
import scipy as sp
# ## ROI: Return on investment (% gain)
#
# $ return = \frac{v_{t2}-v_{t1}}{v_{t1}} = r $
#
# - $v_{t1}$: investment initial value
# - $v_{t2}$: investment final value
def percent_gain(pv, fv, displayPercent=True):
"""
Returns the percent gain from investment
Parameters
----------
pv: int / float
Investment's initial value
fv: int / float
Investment's final value
displayPercent: bool
0 returns result in float format, while 1 returns result in percent format
"""
if not all(isinstance(i, (int, float)) for i in [pv,fv]):
raise TypeError('pv and fv must be int/float datatype')
elif not isinstance(displayPercent, (bool)):
raise TypeError('displayPercent value must be bool datatype')
elif displayPercent is True:
return 'The return on investment is {0}%'.format(str((fv-pv)/pv * 100))
elif displayPercent is not True:
return 'The return on investment is {0}'.format((fv-pv)/pv)
else:
raise Exception('Unknown error, please check the function and values')
# You invest `$10,000` and it's worth `$11,000` after 1 year, what's the interest rate?
percent_gain(10000,11000)
percent_gain(10000,11000,False)
# ## ROI: Return on investment (Dollar Value)
#
# $ v_{t2} = v_{t1} * (1+r) $
#
# - `r`: the rate of return of the investment per period t
# Assuming you invest `$10,000` and the ARR=`10%`, what do you get after 1 year?
np.fv(0.1, 1, 0, -10000)
# ## Cumulative growth / Depreciation
#
# Investment value = $ v_{t_0} * (1+r)^t $
#
# - `r`: growth rate
# Assuming you invest `$10,000` and the ARR=`10%`, what do you get after 10 years?
np.fv(0.1, 10, 0, -10000)
# ## Discount factors
#
# $ df = \frac{1}{(1+r)^t} $
#
# $ v = fv * df $
#
# - `df`: discount factor
# - `v`: investment's initial value
# - `fv`: investment future value
# ## Compound interest
#
# Investment value = $ v_{t_0} * (1 + \frac{r}{c})^{t*c} $
#
# - `r`: annual growth rate
# - `c`: # of compounding periods per year
# Consider a `$1,000` investment with a `10%` annual return, compounded quarterly (every 3 months, 4 times per year)
np.fv(0.1/4, 1*4, 0, -1000)
# Compounded over `30` years
np.fv(0.1/4, 30*4, 0, -1000)
# # 1.1 Present and future values
# Calculate the present value of `$100` received 3 years from now at a `1.0%` inflation rate.
np.pv(0.01, 3, 0, -100)
# Calculate the future value of `$100` invested for 3 years at a `5.0%` average annual rate of return.
np.fv(0.05, 3, 0, -100)
# # 1.2 Net present value & cash flows
# ## Net present value (NPV)
#
# NPV is equal to the sum of all discounted cash flows. NPV is a simple cash flow valuation measure that does not allow for the comparison of different sized projects or lengths.
#
# $ NPV = \sum^T_{t=1} \frac{C_t}{(1+r)^t} - C_0 $
#
# - $C_t$: cashflow at time t
# - `r`: discount rate
# ### Discount cash flow
# https://www.investopedia.com/terms/d/dcf.asp
# Year 1-5
50.93 + 51.87 + 50.43 + 49.03 + 47.67
# Include 0 as we don't have cashflow for year 0
# Don't include terminal from the example
cf = np.array([0, 55, 60.5, 63.53, 66.7, 70.04])
np.npv(rate=0.08, values=cf)
# # 2.0 A tale of 2 project proposals
# ## Internal rate of return (IRR)
#
# The internal rate of return must be computed by solving for IRR in the NPV equation when set equal to 0. IRR can be used to compare projects of different sizes and lengths but requires an algorithmic solution and does not measure total value.
#
# $ NPV = \sum^T_{t=1} \frac{C_t}{(1+IRR)^t} - C_t = 0 $
#
# - `IRR`: internal rate of return
#
# https://en.wikipedia.org/wiki/Internal_rate_of_return#Example
cf = np.array([-123400,36200,54800,48100])
np.irr(cf)
# # 2.1 The weighted average cost of capital (WACC)
#
# $ WACC = F_{equity}*C_{equity} + F_{debt}*C_{debt} * (1-TR) $
#
# - `F`: financing
# - `C`: cost
# - `TR`: corporate tax rate
#
# $ F_{equity} = \frac{M_{equity}}{M_{total}} $
#
# $ F_{debt} = \frac{M_{debt}}{M_{total}} $
#
# $ M_{total} = M_{equity} + M_{debt}$
#
# **WACC vs discount rate**: https://www.investopedia.com/ask/answers/052715/what-difference-between-cost-capital-and-discount-rate.asp
# Calculate the WACC of a company with 12% cost of debt, 14% cost of equity, 20% debt financing and 80% equity financing, with a corporate tax rate of 35%
# +
financing_equity = 0.8
cost_equity = 0.14
financing_debt = 0.2
cost_debt = 0.12
corporate_tax_rate = 0.35
WACC = financing_equity*cost_equity + financing_debt*cost_debt * (1-corporate_tax_rate)
WACC
# -
# Calculate the NPV of a project that provides $100 in cashflow every year for 5 years. With WACC of 13%
cf_project1 = np.repeat(100,5)
npv_project1 = np.npv(0.13, cf_project1)
print(npv_project1)
# # 2.2 Comparing 2 projects of different life spans
# +
project1_cf = np.array([-100,200,300])
project2_cf = np.array([-125,100,100,100,100,100,100,100])
print('project1 length:', len(project1_cf))
print('project2 length:', len(project2_cf))
# NPV: Assume 5% discount rate for both projects
project1_npv = np.npv(0.05, project1_cf)
project2_npv = np.npv(0.05, project2_cf)
print('project1_npv:', project1_npv)
print('project2_npv:', project2_npv)
# IRR
project1_irr = np.irr(project1_cf)
project2_irr = np.irr(project2_cf)
print('project1_irr:', project1_irr)
print('project2_irr:', project2_irr)
# -
# You can undertake multiple project 1 (3 years) over 8 years (which project 2 requires). Even though project 2 has a higher NPV, it has a smaller IRR.
# ## Equivalent annual annuity
#
# Use to compare 2 projects of different lifespans in present value terms
project1_eea = np.pmt(rate=0.05, nper=3, pv=-1*project1_npv, fv=0)
project2_eea = np.pmt(rate=0.05, nper=8, pv=-1*project2_npv, fv=0)
print('project1_eea:', project1_eea)
print('project2_eea:', project2_eea)
# project 1 has higher EEA than project 2
# # 3.0 Mortgage basics
#
# A mortgage is a loan that covers the remaining cost of a home after paying a percentage of the home value as downpayment.
#
# ## Converting annual rate to periodic rate
# $ R_{periodic} = (1 + R_{annual})^{\frac{1}{N}} - 1 $
#
# - `R`: rate of return
# - `N`: number of payment periods per year
# Convert a 12% annual interest rate to the equivalent monthly rate
(1 + 0.12)**(1/12) - 1
def annual_to_periodic_rate(annualRate, periods=12):
"""
Returns a periodic rate
Parameters
----------
annualRate: float
Represents the annual interest rate, where 0 = 0% and 1 = 100%
periods: int
Represents the number of payment periods per year
"""
if annualRate is None or not isinstance(annualRate, (float)):
raise ValueError('Please provide a valid annual rate with float datatype')
if periods is None or not isinstance(periods, (int)):
raise ValueError('Please provide a valid payment periods with int datatype')
periodicRate = (1 + annualRate)**(1/periods) - 1
return periodicRate
# Calculate the monthly mortgage payment of a $400k 30 year loan at 3.8% interest
monthlyRate = annual_to_periodic_rate(.038)
np.pmt(rate=monthlyRate, nper=12*30, pv=400000)
# # 3.1 Amortization, principal and interest
#
# ## Amortization
# `Principal` (equity): The amount of your mortgage paid that counts towards the value of the house itself
#
# `Interest payment`: $ IP_{periodic} = RMB * R_{periodic} $
#
# `Principal payment`: $ PP_{periodic} = MP_{periodic} - IP_{periodic} $
#
# `PP`: prinicpal payment
#
# `MP`: mortgage payment
#
# `IP`: interest payment
#
# `R`: mortgage periodic interest rate
#
# `RMB`: remaining mortgage balance
# # 3.2 Home ownership, equity and forecasting
#
# `Home equity`: percent of the home you actually owned
#
# $ Percent\ equity\ owned = P_{downpayment} + \frac{E_{cumulative,t}}{V_{home\ value}} $
#
# $ E_{cumulative,t} = \sum^T_{t=1} P_{principal,t} $
#
# $E_{cumulative,t}$: Cumulative home equity at time t
#
# $P_{principal,t}$: Principal payment at time t
# ## Underwater mortgage
#
# It is when the remaining amount you owe on your mortgage is actually higher than the value of the house itself
np.cumsum(np.array([1,2,3]))
np.cumprod(np.array([1,2,3]))
# What's the cumulative value at each point in time of a $100 investment that grows by 3% in period 1, then 3% again in period 2, and then 5% in period 3?
np.cumprod(1 + np.array([0.03,0.03,0.05]))
# # 4.0 Budgeting project proposal
# ## Constant cumulative growth forecast
#
# What's the cumulative growth of an investment that grows by 3% per year for 3 years?
np.cumprod(1 + np.repeat(0.03, 3)) - 1
# ## Forecasting values from growth rates
#
# Compute the value at each point in time of an initial $100 investment that grows by 3% per year for 3 years
100 * np.cumprod(1 + np.repeat(0.03, 3))
# # 4.1 Net worth and valuation in your personal financial life
#
# Net worth = Assets - liabilities = equity
#
# ## Valuation
# - NPV (discount rate, cash flows)
# - Future cash flows, salary and expenses
# - Adjust for inflation
# # 4.2 The power of time and compound interest
#
# Save $1 million over 40 years. Assume an average 7% rate of return per year
monthlyRate = annual_to_periodic_rate(0.07)
np.pmt(rate=monthlyRate, nper=12*40, pv=0, fv=1000000)
# If ROI is 5% per year?
monthlyRate = annual_to_periodic_rate(0.05)
np.pmt(rate=monthlyRate, nper=12*40, pv=0, fv=1000000)
# Save $1 million over 25 years. Assume an average 7% rate of return per year
monthlyRate = annual_to_periodic_rate(0.07)
np.pmt(rate=monthlyRate, nper=12*25, pv=0, fv=1000000)
# If ROI is 5% per year?
monthlyRate = annual_to_periodic_rate(0.05)
np.pmt(rate=monthlyRate, nper=12*25, pv=0, fv=1000000)
# Save $1 million over 10 years, assume an average 9% rate of return per year
monthlyRate = annual_to_periodic_rate(0.09)
np.pmt(rate=monthlyRate, nper=12*10, pv=0, fv=1000000)
# Assume an average rate of inflation of 3% per year, what will be present value of $1 million be worth in 25 years?
np.fv(rate=-0.03, nper=25, pv=-1000000, pmt=0)
# # 4.3 Summary
#
# - The time value of money
# - Compound interest
# - Discounting and projecting cash flows
# - Making rational economic decisions
# - Mortgage structures
# - Interest and equity
# - The cost of capital
# - Wealth accumulation
| intro-to-financial-concepts-using-python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Perform Phone Segmentation in the Notebook
# Copyright (c) 2021 <NAME>, MIT License
# ## Preliminaries
# +
# %matplotlib inline
from pathlib import Path
from tqdm import tqdm
import numpy as np
import pylab as plt
import sys
sys.path.append("..")
from utils.eval_segmentation import boundaries_to_intervals, intervals_to_boundaries
from vqwordseg.algorithms import get_segment_intervals, dp_penalized, neg_log_gamma
# -
# ## Read spectrograms, codes, segmentation
# +
# Data set
dataset = "buckeye"
split = "val"
# model = "vqcpc"
model = "cpc_big"
# Utterances
utterances = [
"s22_02b_027666-027761",
"s17_01a_001056-001166",
]
# Directories
indices_dir = Path("../exp/")/model/dataset/split/"indices"
z_dir = Path("../exp/")/model/dataset/split/"prequant"
audio_dir = Path("../../VectorQuantizedCPC/datasets/")/dataset/split
phoneref_dir = Path("../data/")/dataset/"phone_intervals"
# Filenames
embedding_fn = Path("../exp/")/model/dataset/"embedding.npy"
# +
# Embedding matrix
embedding = np.load(embedding_fn)
# Read indices
indices_dict = {}
for utterance_key in sorted(utterances):
indices_fn = (indices_dir/utterance_key).with_suffix(".txt")
indices_dict[utterance_key] = np.asarray(np.loadtxt(indices_fn), dtype=int)
# Read pre-quantisation
z_dict = {}
for utterance_key in sorted(utterances):
z_fn = (z_dir/utterance_key).with_suffix(".txt")
z_dict[utterance_key] = np.loadtxt(z_fn)
# Audio
audio_dict = {}
for utterance_key in sorted(utterances):
fn = (audio_dir/utterance_key.split("_")[0]/utterance_key).with_suffix(".mel.npy")
audio_dict[utterance_key] = np.load(fn).T
# Read reference
reference_interval_dict = {}
for utterance_key in sorted(utterances):
reference_interval_dict[utterance_key] = []
fn = (phoneref_dir/utterance_key).with_suffix(".txt")
for start, end, label in [i.split() for i in fn.read_text().strip().split("\n")]:
start = int(start)
end = int(end)
reference_interval_dict[utterance_key].append((start, end, label))
# Convert intervals to boundaries
reference_boundaries_dict = {}
for utterance_key in reference_interval_dict:
reference_boundaries_dict[utterance_key] = intervals_to_boundaries(
reference_interval_dict[utterance_key]
)
# -
# ## Plot codes on top of spectrograms
# +
downsample_factor = 1
for utt_key in utterances[1:]:
features = audio_dict[utt_key].T
indices = indices_dict[utt_key]
phones = reference_interval_dict[utt_key]
# Codes
plt.figure(figsize=(15, 4))
plt.imshow(features, origin="bottom", aspect="auto", interpolation="nearest")
for i in range(len(indices)):
plt.text(i*downsample_factor + 0.5, features.shape[0] - 3, str(indices[i]), color="w", va="center", ha="center", rotation=90, size=9)
plt.hlines(features.shape[0] - 7, -0.5, features.shape[1] - 0.5, colors="w", lw=1)
# Reference
for start, end, label in phones:
mid_frame = start + (end - start)/2.0
plt.text(mid_frame - 0.5, 3, label, color="w", va="center", ha="center")
plt.vlines(
np.where(reference_boundaries_dict[utt_key])[0] + 0.5, ymin=-0.5, ymax=73, colors="w", lw=1
) # reference boundaries
# -
# ## Segmentation
# Segmentation
boundaries_dict = {}
code_indices_dict = {}
for utt_key in utterances[:1]:
# Segment
z = z_dict[utt_key]
boundaries, code_indices = dp_penalized(embedding, z, dur_weight=3)
# boundaries, code_indices = dp_penalized(
# embedding, z, dur_weight=1,
# dur_weight_func=neg_log_gamma,
# model_eos=True
# )
# Convert boundaries to same frequency as reference
if downsample_factor > 1:
boundaries_upsampled = np.zeros(
len(boundaries)*downsample_factor, dtype=bool
)
for i, bound in enumerate(boundaries):
boundaries_upsampled[i*downsample_factor + 1] = bound
boundaries = boundaries_upsampled
code_indices_upsampled = []
for start, end, index in code_indices:
code_indices_upsampled.append((
start*downsample_factor,
end*downsample_factor,
index
))
code_indices = code_indices_upsampled
boundaries_dict[utt_key] = boundaries
code_indices_dict[utt_key] = code_indices
for utt_key in utterances[:1]:
features = audio_dict[utt_key].T
indices = indices_dict[utt_key]
phones = reference_interval_dict[utt_key]
# Codes
plt.figure(figsize=(15, 4))
plt.imshow(features, origin="bottom", aspect="auto", interpolation="nearest")
for i in range(len(indices)):
plt.text(i*downsample_factor + 0.5, features.shape[0] - 3, str(indices[i]), color="w", va="center", ha="center", rotation=90, size=9)
plt.hlines(features.shape[0] - 7, -0.5, features.shape[1] - 0.5, colors="w", lw=1)
# Reference
for start, end, label in phones:
mid_frame = start + (end - start)/2.0
plt.text(mid_frame - 0.5, 3, label, color="w", va="center", ha="center")
plt.vlines(
np.where(reference_boundaries_dict[utt_key])[0] + 0.5, ymin=-0.5, ymax=73, colors="w", lw=1
) # reference boundaries
# Segmentation
c = "orange"
for start, end, label in code_indices_dict[utt_key]:
mid_frame = start + (end - start)/2.0
plt.text(mid_frame - 0.5, features.shape[0] - 11, label, color=c, va="center", ha="center", rotation=90, size=9)
plt.vlines(
np.where(boundaries_dict[utt_key][:-1])[0] + 0.5, ymin=-0.5, ymax=features.shape[0] - 7, colors=c, lw=1, linestyles="dashed"
) # predicted boundaries
plt.hlines(features.shape[0] - 16, -0.5, features.shape[1] - 0.5, colors=c, lw=1, linestyles="dashed")
for utt_key in utterances[:1]:
features = audio_dict[utt_key].T
indices = indices_dict[utt_key]
phones = reference_interval_dict[utt_key]
# Codes
plt.figure(figsize=(15, 4))
plt.imshow(features, origin="bottom", aspect="auto", interpolation="nearest")
for i in range(len(indices)):
plt.text(i*downsample_factor + 0.5, features.shape[0] - 3, str(indices[i]), color="w", va="center", ha="center", rotation=90, size=9)
plt.hlines(features.shape[0] - 7, -0.5, features.shape[1] - 0.5, colors="w", lw=1)
# Reference
for start, end, label in phones:
mid_frame = start + (end - start)/2.0
plt.text(mid_frame - 0.5, 3, label, color="w", va="center", ha="center")
plt.vlines(
np.where(reference_boundaries_dict[utt_key])[0] + 0.5, ymin=-0.5, ymax=73, colors="w", lw=1
) # reference boundaries
# Segmentation
c = "orange"
for start, end, label in code_indices_dict[utt_key]:
mid_frame = start + (end - start)/2.0
plt.text(mid_frame - 0.5, features.shape[0] - 11, label, color=c, va="center", ha="center", rotation=90, size=9)
plt.vlines(
np.where(boundaries_dict[utt_key][:-1])[0] + 0.5, ymin=-0.5, ymax=features.shape[0] - 7, colors=c, lw=1, linestyles="dashed"
) # predicted boundaries
plt.hlines(features.shape[0] - 16, -0.5, features.shape[1] - 0.5, colors=c, lw=1, linestyles="dashed")
| notebooks/phoneseg_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
from main import Name, Bet, DataBase, Result
db = DataBase('db')
# +
def reg_user(name, bet):
user = (Name(name), Bet(bet))
db.users.append(user)
reg_user('Guilherme', 1500)
reg_user('Fábio', 1000)
reg_user('André', 2000)
reg_user('Murilo', 3000)
reg_user('Pedro', 1000)
# -
for user in db.users:
name = user[0].user_name
bet = user[1].user_bet
print(f'Nome: {name} - Bet: {bet}')
names = [c[0].user_name for c in db.users]
bets = [c[1].user_bet for c in db.users]
result = Result(db)
result.result
| Python/Alura/TDD/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Jupyter notebooks
#
# This is a [Jupyter](http://jupyter.org/) notebook using Python. You can install Jupyter locally to edit and interact with this notebook.
#
# # Linear Algebra
#
# You have all seen basic linear algebra before, but this will summarize some different ways of thinking about the fundamental operations. It also presents concepts in a different order than Sauer's book.
#
# Linear algebra is the study of linear transformations on vectors, which represent points in a finite dimensional space. The matrix-vector product $y = A x$ is a linear combination of the columns of $A$. The familiar definition,
#
# $$ y_i = \sum_j A_{i,j} x_j $$
#
# can also be viewed as
#
# $$ y = \Bigg[ A_{:,0} \Bigg| A_{:,1} \Bigg| \dotsm \Bigg] \begin{bmatrix} x_0 \\ x_1 \\ \vdots \end{bmatrix}
# = \Bigg[ A_{:,0} \Bigg] x_0 + \Bigg[ A_{:,1} \Bigg] x_1 + \dotsb . $$
#
# The notation $A_{i,j}$ corresponds to the Python syntax `A[i,j]` and the colon `:` means the entire range (row or column). So $A_{:,j}$ is the $j$th column and $A_{i,:}$ is the $i$th row. The corresponding Python syntax is `A[:,j]` and `A[i,:]`.
# +
# %matplotlib notebook
import numpy
from matplotlib import pyplot
def matmult1(A, x):
"""Entries of y are dot products of rows of A with x"""
y = numpy.zeros_like(A[:,0])
for i in range(len(A)):
row = A[i,:]
for j in range(len(row)):
y[i] += row[j] * x[j]
return y
A = numpy.array([[1,2],[3,5],[7,11]])
x = numpy.array([10,20])
matmult1(A, x)
# +
def matmult2(A, x):
"""Same idea, but more compactly"""
y = numpy.zeros_like(A[:,0])
for i,row in enumerate(A):
y[i] = row.dot(x)
return y
matmult2(A, x)
# +
def matmult3(A, x):
"""y is a linear expansion of the columns of A"""
y = numpy.zeros_like(A[:,0])
for j,col in enumerate(A.T):
y += col * x[j]
return y
matmult3(A, x)
# -
# We will use this version
A.dot(x)
# ### Some common terminology
#
# * The **range** of $A$ is the space spanned by its columns. This definition coincides with the range of a function $f(x)$ when $f(x) = A x$.
# * The **nullspace** of $A$ is the space of vectors $x$ such that $A x = 0$.
# * The **rank** of $A$ is the dimension of its range.
# * A matrix has **full rank** if the nullspace of either $A$ or $A^T$ is empty (only the 0 vector). Equivalently, if all the columns of $A$ (or $A^T$) are linearly independent.
# * A **nonsingular** (or **invertible**) matrix is a square matrix of full rank. We call the inverse $A^{-1}$ and it satisfies $A^{-1} A = A A^{-1} = I$.
#
# $\DeclareMathOperator{\rank}{rank} \DeclareMathOperator{\null}{null} $
# If $A \in \mathbb{R}^{m\times m}$, which of these doesn't belong?
# 1. $A$ has an inverse $A^{-1}$
# 2. $\rank (A) = m$
# 3. $\null(A) = \{0\}$
# 4. $A A^T = A^T A$
# 5. $\det(A) \ne 0$
# 6. $A x = 0$ implies that $x = 0$
#
# When we write $x = A^{-1} y$, we mean that $x$ is the unique vector such that $A x = y$.
# (It is rare that we explicitly compute a matrix $A^{-1}$, though it's not as "bad" as people may have told you.)
# A vector $y$ is equivalent to $\sum_i e_i y_i$ where $e_i$ are columns of the identity.
# Meanwhile, $x = A^{-1} y$ means that we are expressing that same vector $y$ in the basis of the columns of $A$, i.e., $\sum_i A_{:,i} x_i$.
#
B = numpy.array([[2, 3],[0, 4]])
print(B)
print(B.dot(B.T), B.T.dot(B))
Binv = numpy.linalg.inv(B)
Binv.dot(B), B.dot(Binv)
# ### Vandermonde matrices
#
# A Vandermonde matrix is one whose columns are functions evaluated at discrete points.
x = numpy.linspace(-1,1)
A = numpy.array([x**3, x**2, x, 1+0*x]).T
print('shape =', A.shape) # This is a tall matrix with 4 columns
pyplot.style.use('ggplot')
pyplot.figure()
pyplot.plot(x, A)
pyplot.ylim((-1.1,1.1))
pyplot.show()
# * This type of matrix is very common; we can also create it with `numpy.vander`.
#
# We can evaluate polynomials using matrix-vector multiplication.
# For example,
# $$ 5x^3 - 3x = \Bigg[ x^3 \Bigg|\, x^2 \Bigg|\, x \,\Bigg|\, 1 \Bigg] \begin{bmatrix}5 \\ 0 \\ -3 \\ 0 \end{bmatrix} . $$
pyplot.figure()
p = numpy.array([5,0,-3,0])
pyplot.plot(x, A.dot(p))
# Now suppose we know the value of a polynomial at a few points.
# We can use the Vandermonde matrix to find a polynomial through those points.
x1 = numpy.array([-0.9, 0.1, 0.5, 0.8]) # points where we know values
y = numpy.array([1, 2.4, -0.2, 1.3]) # values at those points
pyplot.figure()
pyplot.plot(x1, y, '*')
B = numpy.vander(x1) # Vandermonde matrix at the known points
p = numpy.linalg.solve(B, y) # Compute the polynomial coefficients
print(p)
pyplot.plot(x, A.dot(p)) # Plot the polynomial evaluated at all points
print('B =', B, '\np =', p)
# Evidently $p(x) = 12.983 x^3 - 1.748 x^2 - 9.476 x + 3.352$ is the unique cubic polynomial that interpolates those points.
# Applying $B^{-1}$ converted from the values at the marked points to the polynomial coefficients.
# ## Inner products and orthogonality
#
# The **inner product**
# $$ x^T y = \sum_i x_i y_i $$
# of vectors (or columns of a matrix) tell us about their magnitude and about the angle.
# The **norm** is induced by the inner product,
# $$ \lVert x \rVert = \sqrt{x^T x} $$
# and the angle $\theta$ is defined by
# $$ \cos \theta = \frac{x^T y}{\lVert x \rVert \, \lVert y \rVert} . $$
# Inner products are **bilinear**, which means that they satisfy some convenient algebraic properties
# $$ \begin{split}
# (x + y)^T z &= x^T z + y^T z \\
# x^T (y + z) &= x^T y + x^T z \\
# (\alpha x)^T (\beta y) &= \alpha \beta x^T y \\
# \end{split} . $$
# The pairwise inner products between two sets of vectors can be expressed by collecting the sets as columns in matrices and writing $A = X^T Y$ where $A_{i,j} = x_i^T y_j$.
# It follows from this definition that
# $$ (X^T Y)^T = Y^T X .$$
#
# ### Orthogonal matrices
#
# If $x^T y = 0$ then we say $x$ and $y$ are **orthogonal** (or "$x$ is orthogonal to $y$").
# A vector is said to be **normalized** if $\lVert x \rVert = 1$.
# If $x$ is orthogonal to $y$ and $\lVert x \rVert = \lVert y \rVert = 1$ then we say $x$ and $y$ are **orthonormal**.
# A matrix with orthonormal columns is said to be an **orthogonal matrix**.
# We typically use $Q$ or $U$ and $V$ for matrices that are known/constructed to be orthogonal.
# Orthogonal matrices are always full rank -- the columns are linearly independent.
# The inverse of a *square* orthogonal matrix is its transpose:
# $$ Q^T Q = Q Q^T = I . $$
# Orthogonal matrices are a powerful building block for robust numerical algorithms.
# +
# Make some polynomials
q0 = A.dot(numpy.array([0,0,0,.5])) # .5
q1 = A.dot(numpy.array([0,0,1,0])) # x
q2 = A.dot(numpy.array([0,1,0,0])) # x^2
pyplot.figure()
pyplot.plot(x, numpy.array([q0, q1, q2]).T)
# +
# Inner products of even and odd functions
q0 = q0 / numpy.linalg.norm(q0)
q1.dot(q0), q2.dot(q0), q2.dot(q1)
# +
# What is the constant component of q2?
pyplot.figure()
pyplot.plot(x, q2.dot(q0)*q0)
# +
# Let's project that away so that q2 is orthogonal to q0
q2 = q2 - q2.dot(q0)*q0
Q = numpy.array([q0, q1, q2]).T
print(Q.T.dot(Q))
pyplot.figure()
pyplot.plot(x, Q)
# -
# ## Gram-Schmidt Orthogonalization
#
# Given a collection of vectors (columns of a matrix), we can find an orthogonal basis by applying the above procedure one column at a time.
# +
def gram_schmidt_naive(X):
Q = numpy.zeros_like(X)
R = numpy.zeros((len(X.T),len(X.T)))
for i in range(len(Q.T)):
v = X[:,i].copy()
for j in range(i):
r = v.dot(Q[:,j])
R[j,i] = r
v -= r * Q[:,j] # "modified Gram-Schmidt" - remove each component before next dot product
R[i,i] = numpy.linalg.norm(v)
Q[:,i] = v / R[i,i]
return Q, R
Q, R = gram_schmidt_naive(A)
print(Q.T.dot(Q))
print(numpy.linalg.norm(Q.dot(R)-A))
pyplot.figure()
pyplot.plot(x, Q)
# -
Q, R = gram_schmidt_naive(numpy.vander(x, 4, increasing=True))
pyplot.figure()
pyplot.plot(x, Q)
# ### Theorem: all full-rank $m\times n$ matrices ($m \ge n$) have a unique $Q R$ factorization with $R_{j,j} > 0$.
#
# ### Orthogonal polynomials
#
# We used `x = numpy.linspace(-1,1)` which uses $m=50$ points by default. The number 50 is arbitrary and as we use more points, our columns become better approximations of continuous functions and the vector inner product becomes an integral (up to scaling):
# $$ \frac 2 m \sum_{i=1}^m p_i q_i \approx \int_{-1}^1 p(x) q(x) . $$
#
# When we orthogonalize the monomials using this inner product, we get the [Legendre Polynomials](https://en.wikipedia.org/wiki/Legendre_polynomials) (up to scaling). These polynomials have important applications in physics and engineering, as well as playing an important role in approximation (which we will go into in more detail).
#
# ### Solving equations using QR
#
# To solve
# $$ A x = b $$
# we can compute $A = QR$ and then
# $$ x = R^{-1} Q^T b . $$
#
# This also works for non-square systems!
x1 = numpy.array([-0.9, 0.1, 0.5, 0.8]) # points where we know values
y = numpy.array([1, 2.4, -0.2, 1.3]) # values at those points
pyplot.figure()
pyplot.plot(x1, y, '*')
B = numpy.vander(x1, 2) # Vandermonde matrix at the known points
Q, R = gram_schmidt_naive(B)
p = numpy.linalg.solve(R, Q.T.dot(y)) # Compute the polynomial coefficients
print(p)
pyplot.plot(x, numpy.vander(x,2).dot(p)) # Plot the polynomial evaluated at all points
print('B =', B, '\np =', p)
# +
m = 20
V = numpy.vander(numpy.linspace(-1,1,m), increasing=False)
Q, R = gram_schmidt_naive(V)
def qr_test(qr, V):
Q, R = qr(V)
m = len(Q.T)
print(qr.__name__, numpy.linalg.norm(Q.dot(R) - V), numpy.linalg.norm(Q.T.dot(Q) - numpy.eye(m)))
qr_test(gram_schmidt_naive, V)
qr_test(numpy.linalg.qr, V)
# +
def gram_schmidt_classical(X):
Q = numpy.zeros_like(X)
R = numpy.zeros((len(X.T),len(X.T)))
for i in range(len(Q.T)):
v = X[:,i].copy()
R[:i,i] = Q[:,:i].T.dot(v)
v -= Q[:,:i].dot(R[:i,i])
R[i,i] = numpy.linalg.norm(v)
Q[:,i] = v / R[i,i]
return Q, R
qr_test(gram_schmidt_classical, V)
# -
# Classical Gram-Schmidt is highly parallel, but unstable, as evidenced by the lack of orthogonality in $Q$.
#
# ### Right-looking algorithms
#
# The implementations above have been "left-looking"; when working on column $i$, we compare it only to columns to the left (i.e., $j < i$). We can reorder the algorithm to look to the right by projecting $q_i$ out of all columns $j > i$. This algorithm is stable while being just as parallel as `gram_schmidt_classical`.
# +
def gram_schmidt_modified(X):
Q = X.copy()
R = numpy.zeros((len(X.T), len(X.T)))
for i in range(len(Q.T)):
R[i,i] = numpy.linalg.norm(Q[:,i])
Q[:,i] /= R[i,i]
R[i,i+1:] = Q[:,i+1:].T.dot(Q[:,i])
Q[:,i+1:] -= numpy.outer(Q[:,i], R[i,i+1:])
return Q, R
qr_test(gram_schmidt_modified, V)
# -
# ### Householder triangularization
#
# Gram-Schmidt methods perform triangular transformations to build an orthogonal matrix. As we have seen, $X = QR$ is satisfied accurately, but $Q$ may not be orthogonal when $X$ is ill-conditioned. Householder triangularization instead applies a sequence of orthogonal transformations to build a triangular matrix.
#
# $$ \underbrace{Q_{n-1} \dotsb Q_0}_{Q^T} A = R $$
#
# The structure of the algorithm is
#
# $$ \underbrace{\begin{bmatrix} * & * & * \\ * & * & * \\ * & * & * \\ * & * & * \\ * & * & * \\ \end{bmatrix}}_{A} \to
# \underbrace{\begin{bmatrix} * & * & * \\ 0 & * & * \\ 0 & * & * \\ 0 & * & * \\ 0 & * & * \\ \end{bmatrix}}_{Q_0 A} \to
# \underbrace{\begin{bmatrix} * & * & * \\ 0 & * & * \\ 0 & 0 & * \\ 0 & 0 & * \\ 0 & 0 & * \\ \end{bmatrix}}_{Q_1 Q_0 A} \to
# \underbrace{\begin{bmatrix} * & * & * \\ 0 & * & * \\ 0 & 0 & * \\ 0 & 0 & 0 \\ 0 & 0 & 0 \\ \end{bmatrix}}_{Q_2 Q_1 Q_0 A}
# $$
#
# where the elementary orthogonal matrices $Q_i$ chosen to introduce zeros below the diagonal in the $i$th column of $R$.
# Each of these transformations will have the form
# $$Q_i = \begin{bmatrix} I_i & 0 \\ 0 & F \end{bmatrix}$$
# where $F$ is a "reflection" that achieves
# $$ F x = \begin{bmatrix} \lVert x \rVert \\ 0 \\ 0 \\ \vdots \end{bmatrix} $$
# where $x$ is the column of $R$ from the diagonal down.
# This transformation is a reflection across a plane with normal $v = Fx - x = \lVert x \rVert e_1 - x$.
#
# 
#
# The reflection, as depected above by Trefethen and Bau (1999) can be written $F = I - 2 \frac{v v^T}{v^T v}$.
# +
def householder_Q_times(V, x):
"""Apply orthogonal matrix represented as list of Householder reflectors"""
y = x.copy()
for i in reversed(range(len(V))):
y[i:] -= 2 * V[i] * V[i].dot(y[i:])
return y
def qr_householder1(A):
"Compute QR factorization using naive Householder reflection"
m, n = A.shape
R = A.copy()
V = []
for i in range(n):
x = R[i:,i]
v = -x
v[0] += numpy.linalg.norm(x)
v = v/numpy.linalg.norm(v) # Normalized reflector plane
R[i:,i:] -= 2 * numpy.outer(v, v.dot(R[i:,i:]))
V.append(v) # Storing reflectors is equivalent to storing orthogonal matrix
Q = numpy.eye(m, n)
for i in range(n):
Q[:,i] = householder_Q_times(V, Q[:,i])
return Q, numpy.triu(R[:n,:])
qr_test(qr_householder1, numpy.array([[1.,2],[3,4],[5,6]]))
# -
qr_test(qr_householder1, V)
qr_test(numpy.linalg.qr, V)
# ### Choice of two projections
#
# It turns out our implementation has a nasty deficiency.
qr_test(qr_householder1, numpy.eye(1))
qr_test(qr_householder1, numpy.eye(3,2))
# Inside `qr_householder1`, we have the lines
# ```
# x = R[i:,i]
# v = -x
# v[0] += numpy.linalg.norm(x)
# v = v/numpy.linalg.norm(v) # Normalized reflector plane
# ```
# What happens when $$x = \begin{bmatrix}1 \\ 0 \end{bmatrix}$$
# (i.e., the column of $R$ is already upper triangular)?
#
# We are trying to define a reflector plane (via its normal vector) from the zero vector,
# $$v = \lVert x \rVert e_0 - x .$$
# When we try to normalize this vector, we divide zero by zero and the algorithm breaks down (`nan`). Maybe we just need to test for this special case and "skip ahead" when no reflection is needed? And if so, how would we define $Q$?
qr_test(qr_householder1, numpy.array([[1.,1], [2e-8,1]]))
print(qr_householder1(numpy.array([[1.,1], [2e-8,1]])))
# The error $QR - A$ is still $10^{-8}$ for this very well-conditioned matrix so something else must be at play here.
#
# 
# +
def qr_householder2(A):
"Compute QR factorization using Householder reflection"
m, n = A.shape
R = A.copy()
V = []
for i in range(n):
v = R[i:,i].copy()
v[0] += numpy.sign(v[0])*numpy.linalg.norm(v) # Choose the further of the two reflections
v = v/numpy.linalg.norm(v) # Normalized reflector plane
R[i:,i:] -= 2 * numpy.outer(v, v.dot(R[i:,i:]))
V.append(v) # Storing reflectors is equivalent to storing orthogonal matrix
Q = numpy.eye(m, n)
for i in range(n):
Q[:,i] = householder_Q_times(V, Q[:,i])
return Q, numpy.triu(R[:n,:])
qr_test(qr_householder2, numpy.eye(3,2))
qr_test(qr_householder2, numpy.array([[1.,1], [1e-8,1]]))
print(qr_householder2(numpy.array([[1.,1], [1e-8,1]])))
qr_test(qr_householder2, V)
# -
# We now have a usable implementation of Householder QR. There are some further concerns for factoring rank-deficient matrices. We will visit the concept of pivoting later, in the context of LU and Cholesky factorization.
#
# ## Condition number of a matrix
#
# We may have informally referred to a matrix as "ill-conditioned" when the columns are nearly linearly dependent, but let's make this concept for precise. Recall the definition of (relative) condition number from the Rootfinding notes,
#
# $$ \kappa = \max_{\delta x} \frac{|\delta f|/|f|}{|\delta x|/|x|} . $$
#
# We understood this definition for scalar problems, but it also makes sense when the inputs and/or outputs are vectors (or matrices, etc.) and absolute value is replaced by vector (or matrix) norms. Let's consider the case of matrix-vector multiplication, for which $f(x) = A x$.
#
# $$ \kappa(A) = \max_{\delta x} \frac{\lVert A (x+\delta x) - A x \rVert/\lVert A x \rVert}{\lVert \delta x\rVert/\lVert x \rVert}
# = \max_{\delta x} \frac{\lVert A \delta x \rVert}{\lVert \delta x \rVert} \, \frac{\lVert x \rVert}{\lVert A x \rVert} = \lVert A \rVert \frac{\lVert x \rVert}{\lVert A x \rVert} . $$
#
# There are two problems here:
#
# * I wrote $\kappa(A)$ but my formula depends on $x$.
# * What is that $\lVert A \rVert$ beastie?
#
# ### Stack push: Matrix norms
#
# Vector norms are built into the linear space (and defined in term of the inner product). Matrix norms are *induced* by vector norms, according to
#
# $$ \lVert A \rVert = \max_{x \ne 0} \frac{\lVert A x \rVert}{\lVert x \rVert} . $$
#
# * This equation makes sense for non-square matrices -- the vector norms of the input and output spaces may differ.
# * Due to linearity, all that matters is direction of $x$, so it could equivalently be written
#
# $$ \lVert A \rVert = \max_{\lVert x \rVert = 1} \lVert A x \rVert . $$
#
# ### Stack pop
#
# Now we understand the formula for condition number, but it depends on $x$. Consider the matrix
#
# $$ A = \begin{bmatrix} 1 & 0 \\ 0 & 0 \end{bmatrix} . $$
#
# * What is the norm of this matrix?
# * What is the condition number when $x = [1,0]^T$?
# * What is the condition number when $x = [0,1]^T$?
#
# The condition number of matrix-vector multiplication depends on the vector. The condition number of the matrix is the worst case (maximum) of the condition number for any vector, i.e.,
#
# $$ \kappa(A) = \max_{x \ne 0} \lVert A \rVert \frac{\lVert x \rVert}{\lVert A x \rVert} .$$
#
# If $A$ is invertible, then we can rephrase as
#
# $$ \kappa(A) = \max_{x \ne 0} \lVert A \rVert \frac{\lVert A^{-1} (A x) \rVert}{\lVert A x \rVert} =
# \max_{A x \ne 0} \lVert A \rVert \frac{\lVert A^{-1} (A x) \rVert}{\lVert A x \rVert} = \lVert A \rVert \lVert A^{-1} \rVert . $$
#
# Evidently multiplying by a matrix is just as ill-conditioned of an operation as solving a linear system using that matrix.
# +
def R_solve(R, b):
"""Solve Rx = b using back substitution."""
x = b.copy()
m = len(b)
for i in reversed(range(m)):
x[i] -= R[i,i+1:].dot(x[i+1:])
x[i] /= R[i,i]
return x
Q, R = numpy.linalg.qr(A)
b = Q.T.dot(A.dot(numpy.array([1,2,3,4])))
numpy.linalg.norm(R_solve(R, b) - numpy.linalg.solve(R, b))
R_solve(R, b)
# -
# ### Cost of Householder factorization
#
# The dominant cost comes from the line
# ```Python
# R[i:,i:] -= 2 * numpy.outer(v, v.dot(R[i:,i:]))
# ```
# were `R[i:,i:]` is an $(m-i)\times(n-i)$ matrix.
# This line performs $2(m-i)(n-i)$ operations in `v.dot(R[i:,i:])`, another $(m-i)(n-i)$ in the "outer" product and again in subtraction. As written, multiplication by 2 would be another $(m-i)(n-i)$ operations, but is only $m-i$ operations if we rewrite as
# ```Python
# w = 2*v
# R[i:,i:] -= numpy.outer(w, v.dot(R[i:,i:]))
# ```
# in which case the leading order cost is $4(m-i)(n-i)$. To compute the total cost, we need to sum over all columns $i$,
# $$\begin{split} \sum_{i=1}^n 4(m-i)(n-i) &= 4 \Big[ \sum_{i=1}^n (m-n)(n-i) + \sum_{i=1}^n (n-i)^2 \Big] \\
# &= 4 (m-n) \sum_{i=1}^n i + 4 \sum_{i=1}^n i^2 \\
# &\approx 2 (m-n) n^2 + 4 n^3/3 \\
# &= 2 m n^2 - \frac 2 3 n^3 .
# \end{split}$$
# Recall that Gram-Schmidt QR cost $2 m n^2$, so Householder costs about the same when $m \gg n$ and is markedly less expensive when $m \approx n$.
#
# ## Least squares and the normal equations
#
# A **least squares problem** takes the form: given an $m\times n$ matrix $A$ ($m \ge n$), find $x$ such that
# $$ \lVert Ax - b \rVert $$
# is minimized. If $A$ is square and full rank, then this minimizer will satisfy $A x - b = 0$, but that is not the case in general because $b$ is not in the range of $A$.
# The residual $A x - b$ must be orthogonal to the range of $A$.
#
# * Is this the same as saying $A^T (A x - b) = 0$?
# * If $QR = A$, is it the same as $Q^T (A x - b) = 0$?
#
# In HW2, we showed that $QQ^T$ is an orthogonal projector onto the range of $Q$. If $QR = A$,
# $$ QQ^T (A x - b) = QQ^T(Q R x - b) = Q (Q^T Q) R x - QQ^T b = QR x - QQ^T b = A x - QQ^T b . $$
# So if $b$ is in the range of $A$, we can solve $A x = b$. If not, we need only *orthogonally* project $b$ into the range of $A$.
#
# ### Solution by QR (Householder)
#
# Solve $R x = Q^T b$.
#
# * QR factorization costs $2 m n^2 - \frac 2 3 n^3$ operations and is done once per matrix $A$.
# * Computing $Q^T b$ costs $4 (m-n)n + 2 n^2 = 4 mn - 2n^2$ (using the elementary reflectors, which are stable and lower storage than naive storage of $Q$).
# * Solving with $R$ costs $n^2$ operations. Total cost per right hand side is thus $4 m n - n^2$.
#
# This method is stable and accurate.
#
# ### Solution by Cholesky
#
# The mathematically equivalent form $(A^T A) x = A^T b$ are called the **normal equations**. The solution process involves factoring the symmetric and positive definite $n\times n$ matrix $A^T A$.
#
# * Computing $A^T A$ costs $m n^2$ flops, exploiting symmetry.
# * Factoring $A^T A = R^T R$ costs $\frac 1 3 n^3$ flops. The total factorization cost is thus $m n^2 + \frac 1 3 n^3$.
# * Computing $A^T b$ costs $2 m n$.
# * Solving with $R^T$ costs $n^2$.
# * Solving with $R$ costs $n^2$. Total cost per right hand side is thus $2 m n + 2 n^2$.
#
# The product $A^T A$ is ill-conditioned: $\kappa(A^T A) = \kappa(A)^2$ and can reduce the accuracy of a least squares solution.
#
# ### Solution by Singular Value Decomposition
#
# Next, we will discuss a factorization
# $$ U \Sigma V^T = A $$
# where $U$ and $V$ have orthonormal columns and $\Sigma$ is diagonal with nonnegative entries.
# The entries of $\Sigma$ are called **singular values** and this decomposition is the **singular value decomposition** (SVD).
# It may remind you of an eigenvalue decomposition $X \Lambda X^{-1} = A$, but
# * the SVD exists for all matrices (including non-square and deficient matrices)
# * $U,V$ have orthogonal columns (while $X$ can be arbitrarily ill-conditioned).
# Indeed, if a matrix is symmetric and positive definite (all positive eigenvalues), then $U=V$ and $\Sigma = \Lambda$.
# Computing an SVD requires a somewhat complicated iterative algorithm, but a crude estimate of the cost is $2 m n^2 + 11 n^3$. Note that this is similar to the cost of $QR$ when $m \gg n$, but much more expensive for square matrices.
# Solving with the SVD involves
# * Compute $U^T b$ at a cost of $2 m n$.
# * Solve with the diagonal $n\times n$ matrix $\Sigma$ at a cost of $n$.
# * Apply $V$ at a cost of $2 m n$. The total cost per right hand side is thus $4 m n$.
#
# ### Pseudoinverse
#
# An alternative is to explicitly form the $n\times m$ pseudoinverse $A^\dagger = R^{-1} Q^T$ (at a cost of $mn^2$) at which point each right hand side costs $2 mn$. Why might we do this?
#
# * Lots of right hand sides
# * Real-time solution
#
# +
# Test accuracy of solver for an ill-conditioned square matrix
x = numpy.linspace(-1,1,19)
A = numpy.vander(x)
print('cond(A) = ',numpy.linalg.cond(A))
Q, R = numpy.linalg.qr(A)
print('cond(R^{-1} Q^T A) =', numpy.linalg.cond(numpy.linalg.solve(R, Q.T.dot(A))))
L = numpy.linalg.cholesky(A.T.dot(A))
print('cond(L^{-T} L^{-1} A^T A) =', numpy.linalg.cond(numpy.linalg.solve(L.T, numpy.linalg.solve(L, A.T.dot(A)))))
# -
# ## The [Singular Value Decomposition](https://en.wikipedia.org/wiki/Singular_value_decomposition)
#
# The SVD is the decomposition
# $$ U \Sigma V^T = A $$
# where $U$ and $V$ have orthonormal columns and $\Sigma$ is diagonal and nonnegative.
# Evidenly an arbitrary matrix consists of
# 1. Orthogonal matrix -- rotation and reflection
# 2. Diagonal scaling (transforms a sphere into an ellipsoid aligned to the coordinate axes)
# 3. Orthogonal matrix -- rotation and reflection
#
# It is typical to order the singular values in descending order.
#
# The inverse has the same behavior:
#
# $$ A^{-1} = V \Sigma^{-1} U^T $$
#
# The matrix norm is the largest singular value
#
# $$ \lVert A \rVert = \sigma_{\max} . $$
#
# The largest value in $\Sigma^{-1}$ is $\sigma_\min^{-1}$, so
#
# $$ \lVert A^{-1} \rVert = \sigma_\min^{-1} . $$
#
# We showed previously that
#
# $$\kappa(A) = \lVert A \rVert \, \lVert A^{-1} \rVert $$
#
# so now we can also write
#
# $$ \kappa(A) = \frac{\sigma_\max}{\sigma_\min} . $$
#
# The SVD is a crucial tool in statistics and dimensionality reduction, often under names like
#
# * [Principal Component Analysis (PCA)](https://en.wikipedia.org/wiki/Principal_component_analysis)
# * Proper Orthogonal Decomposition (POD)
# * Karhunen-Loeve Expansion
#
# In this context, one computes an SVD of a data matrix and finds that the spectrum $\Sigma$ decays rapidly and only the $k \ll n$ components larger than some threshold are "important". The dense matrix can thus be approximated as
#
# $$ \hat U \hat \Sigma \hat V^T $$
#
# by keeping only the first (largest) $k$ columns. This reduces the storage from $mn$ entries to $mk + kn$ entries.
# ## Tinkering in class 2016-10-10
m = 10
x = numpy.cos(numpy.linspace(0,numpy.pi,m))
f = 1.0*(x > 0) + (x < 0.5)
A = numpy.vander(x)
Q, R = numpy.linalg.qr(A)
p = numpy.linalg.solve(R, Q.T.dot(f))
y = numpy.linspace(-1,1,50)
g = numpy.vander(y, m).dot(p)
pyplot.figure()
pyplot.plot(x, f, '*')
pyplot.plot(y, g)
print(numpy.linalg.cond(A))
'%10e' % numpy.linalg.cond(numpy.vander(numpy.linspace(-1,1,100),20))
| LinearAlgebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stacks, Queues, and Deques Overview
# In this section of the course we will be learning about Stack, Queues, and Deques. These are linear structures. They are similar to arrays, but each of these structures differs by how it adds and removes items.
#
# Here's what to expect in this section:
#
# 1.) A Brief Overview of the Linear Structures
# 2.) An Overview of Stacks
# 3.) An Implementation of a Stack class
# 4.) An Overview of Queues
# 5.) An Implementation of a Queue class
# 6.) An Overview of Deques
# 7.) An Implementation of a Deque class
#
# Then finally a variety of interview questions based on Stacks, Queues, and Deques!
#
# **See the lecture video for a complete breakdown of this Section of the course!**
| code/algorithms/course_udemy_1/Stacks, Queues and Deques/Stacks, Queues, and Deques Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import nltk
import nltk.corpus
# +
###tokenizetion
engtxt ="""Perhaps one of the most significant advances made by Arabic mathematics began at this time with the work of al-Khwarizmi, namelythe beginnings of algebra. It is important to understand just how significant this new idea was. It was a revolutionary move away from the Greek concept of mathematics which was essentially geometry. Algebra was a unifying theory which allowed rational numbers, irrational numbers, geometrical magnitudes, etc., to all be treated as \"algebraic objects\". It gave mathematics a whole new development path so much broader in concept to that which had existed before, and provided a vehicle for future development of the subject. Another important aspect of the introduction of algebraic ideas was that it allowed mathematics to be applied to itself in a way which had not happened before."""
arabtxt="""ربما كانت أحد أهم التطورات التي قامت بها الرياضيات العربية التي بدأت في هذا الوقت بعمل الخوارزمي وهي بدايات الجبر, ومن المهم فهم كيف كانت هذه الفكرة الجديدة مهمة, فقد كانت خطوة نورية بعيدا عن المفهوم اليوناني للرياضيات التي هي في جوهرها هندسة, الجبر کان نظرية موحدة تتيح الأعداد الكسرية والأعداد اللا كسرية, والمقادير الهندسية وغيرها, أن تتعامل على أنها أجسام جبرية, وأعطت الرياضيات ككل مسارا جديدا للتطور بمفهوم أوسع بكثير من الذي كان موجودا من قبل, وقم وسيلة للتنمية في هذا الموضوع مستقبلا. وجانب آخر مهم لإدخال أفكار الجبر وهو أنه سمح بتطبيق الرياضيات على نفسها بطريقة لم تحدث من قبل"""
# -
from nltk.tokenize import word_tokenize
nltk.download('punkt')
Eng_tokens = word_tokenize(engtxt)
Eng_tokens
Arab_tokens = word_tokenize(arabtxt)
Arab_tokens
len(Eng_tokens)
### words_Frequency
from nltk.probability import FreqDist
fdict = FreqDist()
for word in Eng_tokens:
fdict[word.lower()]+=1
fdict
nltk.download("stopwords")
from nltk.corpus import stopwords
Engwords = set(stopwords.words("english"))
Arwords = set(stopwords.words("arabic"))
clair_txt = []
for word in Eng_tokens:
if word.casefold() not in Engwords:
clair_txt.append(word)
clair_txt
###paragraph tokenizetionf
from nltk.tokenize import blankline_tokenize
eng_blank = blankline_tokenize(engtxt)
eng_blank
len(eng_blank)
from nltk.util import bigrams, trigrams, ngrams
quotes_bigrams = list(nltk.bigrams(Eng_tokens))
quotes_bigrams
#####################STEMMING########################
from nltk.stem import PorterStemmer
pst = PorterStemmer()
d = dict()
for w in Eng_tokens:
if pst.stem(w) in d:
d[pst.stem(w)]+=1
else:
d[pst.stem(w)] = 1
print(d)
##################lemmatization#########################
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
from nltk.stem import wordnet
lemmatizer = WordNetLemmatizer()
lemmatizer
for w in Eng_tokens:
print(w," : ",lemmatizer.lemmatize(w))
##################################### POS tagging ########################################################################
nltk.download('averaged_perceptron_tagger')
for w in Eng_tokens:
print(nltk.pos_tag([w]))
################################## entity recognition ###########################################################
nltk.download('words')
from nltk import ne_chunk
tags = nltk.pos_tag(Eng_tokens)
chnuks = ne_chunk(tags)
print(chnuks)
################################# chunking #######################################################################
Grammar = "NP: {<DT>?<JJ>*<NN>}"
chunk_parser = nltk.RegexpParser(Grammar)
# !pip install ghostscript
t = chunk_parser.parse(tags)
t.draw()
# +
####################################" end " ################################################################""""""
| NLTK nlp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/timrocar/DS-Unit-2-Linear-Models/blob/master/module4-logistic-regression/LS_DS_214_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="m4d2dQbCliRi" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 4*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Logistic Regression
#
#
# ## Assignment 🌯
#
# You'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
#
# > We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
# - [ ] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# - [ ] Begin with baselines for classification.
# - [ ] Use scikit-learn for logistic regression.
# - [ ] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
# - [ ] Get your model's test accuracy. (One time, at the end.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
#
# ## Stretch Goals
#
# - [ ] Add your own stretch goal(s) !
# - [ ] Make exploratory visualizations.
# - [ ] Do one-hot encoding.
# - [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html).
# - [ ] Get and plot your coefficients.
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="J5Juq7BCliR0" colab_type="code" colab={}
# Load data downloaded from https://srcole.github.io/100burritos/
import pandas as pd
df = pd.read_csv(DATA_PATH+'burritos/burritos.csv')
# + id="_HrjG4kHliSD" colab_type="code" colab={}
# Derive binary classification target:
# We define a 'Great' burrito as having an
# overall rating of 4 or higher, on a 5 point scale.
# Drop unrated burritos.
df = df.dropna(subset=['overall'])
df['Great'] = df['overall'] >= 4
# + id="8JIhuCBTliSL" colab_type="code" colab={}
# Clean/combine the Burrito categories
df['Burrito'] = df['Burrito'].str.lower()
california = df['Burrito'].str.contains('california')
asada = df['Burrito'].str.contains('asada')
surf = df['Burrito'].str.contains('surf')
carnitas = df['Burrito'].str.contains('carnitas')
df.loc[california, 'Burrito'] = 'California'
df.loc[asada, 'Burrito'] = 'Asada'
df.loc[surf, 'Burrito'] = 'Surf & Turf'
df.loc[carnitas, 'Burrito'] = 'Carnitas'
df.loc[~california & ~asada & ~surf & ~carnitas, 'Burrito'] = 'Other'
# + id="FYOYOrGLliST" colab_type="code" colab={}
# Drop some high cardinality categoricals
df = df.drop(columns=['Notes', 'Location', 'Reviewer', 'Address', 'URL', 'Neighborhood'])
# + id="D8AjCnAzliSe" colab_type="code" colab={}
# Drop some columns to prevent "leakage"
df = df.drop(columns=['Rec', 'overall'])
# + [markdown] id="hVzjUY_rG3dT" colab_type="text"
# ## EDA
# + id="8pjQeLrTliSo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 241} outputId="e5875814-02e8-40f5-c073-e3daa7ac91f6"
df.head()
# + id="MXm7MQPgCgIl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="a2b693d6-61b5-47c0-e3b9-560fadfc3e24"
df.info()
# + id="QERs7ShsFX0V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 334} outputId="861bfbf3-9201-4ab6-97ff-5a12e0041b00"
df.describe()
# + id="vWATmV9OChye" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 241} outputId="cc417343-432c-4444-f16d-b544337435b5"
df.select_dtypes('object').head()
# + id="u9cFKxgfViBH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 719} outputId="f82d7743-02d2-45dd-8574-99d9642a2328"
list(df.select_dtypes('object').columns)
# + [markdown] id="WxAtH5LQAgM_" colab_type="text"
# We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
# - [ ] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# - [ ] Begin with baselines for classification.
# - [ ] Use scikit-learn for logistic regression.
# - [ ] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
# - [ ] Get your model's test accuracy. (One time, at the end.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
# + id="pXlbVcV4JJZh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 241} outputId="816df4da-bfc5-4f23-cfa5-af8504788c7d"
df.head()
# + id="pQPzC0RaFf7S" colab_type="code" colab={}
## Setting the date column as my index
df['Date'] = pd.to_datetime(df['Date'], format='%m/%d/%Y')
df.set_index('Date', inplace=True)
# + id="RQyTCDqSH1Bv" colab_type="code" colab={}
# Target vector
y = df.Great
# Feature Matrix
X = df.drop(['Queso', 'NonSD', 'Unreliable', 'Burrito', 'Mass (g)', 'Density (g/mL)', 'Great'], axis=1)
# + id="YM_gIh2nKbDw" colab_type="code" colab={}
## Splitting Data
# Training w data from 2016 and earlier
cutoff = '2017-01-01'
mask = X.index < cutoff
X_train, y_train = X.loc[mask], y.loc[mask]
## Validation data from 2017
end_date = '2018-01-01'
valmask = (X.index >= cutoff) & (X.index < end_date)
X_val, y_val = X.loc[valmask], y.loc[valmask]
## Test data
testmask = X.index > end_date
X_test, y_test = X.loc[testmask], y.loc[testmask]
# + id="dvxeul1NLcXD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="c7aac6bf-3eb6-469d-c500-0633d7e739f9"
print(X.shape)
print(X_train.shape)
print(X_val.shape)
print(X_test.shape)
# + [markdown] id="gbmAKPf7U-Pz" colab_type="text"
# ## Setting Baselines!
# + id="BxJDwFWhLl14" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6b47e241-ad75-4ab3-f222-bfc2caf08b74"
## Classification baseline - (Majority Classifier ---> Accuracy score)
print('Baseline Accuracy Score:', y_train.value_counts(normalize=True).max())
# + [markdown] id="iZnMmabaWKpD" colab_type="text"
# ## Model Building
# + id="tTYxwfc1VSHY" colab_type="code" colab={}
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
log_model = Pipeline([
('imputer', SimpleImputer()),
('regressor', LogisticRegression())
])
log_model.fit(X_train[['Tortilla']], y_train)
y_pred_log = log_model.predict(X_train[['Tortilla']])
# + id="hcENjjcyXuCa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="324d3807-37bb-4b92-8fd5-ef1e5df2b728"
X_train.head()
# + id="deSPcCB8XySM" colab_type="code" colab={}
from category_encoders import OneHotEncoder
# + id="UlfPjJ3_X3QP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 575} outputId="3b27411e-81bb-4ca8-bc31-7b838bf957c9"
full_model = Pipeline([
('encoder', OneHotEncoder(cols=['Chips', 'Beef', 'Pico', 'Guac',
'Cheese', 'Fries', 'Sour cream', 'Pork', 'Chicken', 'Shrimp', 'Fish',
'Rice', 'Beans', 'Lettuce', 'Tomato', 'Bell peper', 'Carrots',
'Cabbage', 'Sauce', 'Salsa.1', 'Cilantro', 'Onion', 'Taquito',
'Pineapple', 'Ham', 'Chile relleno', 'Nopales', 'Lobster', 'Egg',
'Mushroom', 'Bacon', 'Sushi', 'Avocado', 'Corn', 'Zucchini'])),
('imputer', SimpleImputer()),
('regressor', LogisticRegression())
])
full_model.fit(X_train, y_train)
# + id="I-HPpc4lZEMt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="9a4835a8-0e38-4324-b509-87ae205f503c"
print('Training Accuracy:', full_model.score(X_train, y_train))
print('Validation Accuracy:', full_model.score(X_val, y_val))
print('Testing Accuracy:', full_model.score(X_test, y_test))
# + id="a9pkKF6FZjQ7" colab_type="code" colab={}
| module4-logistic-regression/LS_DS_214_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%capture
## compile PyRossTSI for this notebook
import os
owd = os.getcwd()
os.chdir('../..')
# %run setup.py install
os.chdir(owd)
import pyross.tsi as pyrosstsi
import numpy as np
import nlopt
import matplotlib.pyplot as plt
plt.rcParams.update({'font.size': 12})
# +
T = 15 # Longest infectious duration
Td = 5 # Doubling time in linear growth regime
Tf = 150 # Duration of simulation
tsi = np.array([0, 3, 5, 10, T]) # Time since infection (days)
beta = np.array([0, 0.5, 1, .5, 0]) # Mean infectiousness
M = 2 # Number of age groups to model
Ni = 10**6*np.ones(M) # Number of people in each age group
Np = sum(Ni) # Total population size
#how many 'stages' to resolve in time since infection?
Nk = 10
#define a time-dependent contact matrix.
def contactMatrix_0(t):
return np.array([[4, 1],[1, 2]]);
subclasses = ['Recovered', 'Hospitalized', 'Mortality']
pR = 0.99*np.ones(M); #probability of eventually recovering for each age class
pH = 0.05*np.ones(M); #probability of needing hospitalization for each age class
pD = 1-pR; #probability of death for each age class
#prepare for a linear interpolating function evaluated at times:
tsi_sc = np.array([0, 3., 6., 9., 12, T])
phiR = np.array([0, 0, 0.5, 3, 2, 0]) #rate of transferring to 'recovered' (arbitrary units)
phiH_in = np.array([0, 0, 1, 1, 0, 0]) #rate that people enter hospital (arbitrary units)
phiH_out = np.array([0, 0, 0, 1, 1, 0]) #rate that people enter hospital (arbitrary units)
phiD = np.array([0, 0, 0, 1, 1, .5]) #times at which a person dies (arbitrary units)
#combine hospital in/out to a single function for net change in hospitalized cases
phiH = np.add(-phiH_out/np.trapz(phiH_out,tsi_sc),phiH_in/np.trapz(phiH_in,tsi_sc))
#normalize all to one -- can then be rescaled by approprate pR, pH, pD, etc. at a later time
phiR, phiD = phiR/np.trapz(phiR,tsi_sc), phiD/np.trapz(phiD,tsi_sc)
#group them all together for later processing
phi_alpha, p_alpha = np.array([phiR, phiH, phiD]), np.array([pR, pH, pD])
# +
#Reference simulation: No control
parameters = {'M':M, 'Ni':Ni, 'Nc':len(subclasses), 'Nk':Nk, 'Tf':Tf, 'Tc':(T/2), 'T':T, 'Td':Td,
'tsi':tsi,'beta':beta,'tsi_sc':tsi_sc, 'phi_alpha':phi_alpha, 'p_alpha':p_alpha,
'contactMatrix':contactMatrix_0}
#Set up and Solve
model = pyrosstsi.deterministic.Simulator(parameters)
IC = model.get_IC()
data = model.simulate(IC)
#unpack and rescale simulation output
t = data['t']; S_t = data['S_t']; I_t = data['I_t']; Ic_t = data['Ic_t']
plt.figure(figsize=(12, 4)); plt.subplot(121)
plt.plot(t,np.sum(S_t,0), color="#348ABD", lw=2, label = 'Susceptible') #all susceptible
plt.plot(t,np.sum(I_t,0), color="#A60628", lw=2, label = 'Infected') #all Infected
plt.plot(t,np.sum(Ic_t[0,:,:],0), color='green', lw=2, label = 'Recovered') #all Recovered
plt.xlabel('time (days)'); plt.xlim(0,Tf);
plt.ylabel('Fraction of compartment value'); plt.legend()
plt.subplot(122)
for i in (1 + np.arange(len(subclasses)-1)):
plt.plot(t,np.sum(Ic_t[i,:,:],0), lw=2, label = subclasses[i])
plt.legend(); plt.xlabel('time (days)'); plt.xlabel('time (days)'); plt.xlim(0,Tf);
# -
# ## Set up a Function for Cost Evaluations
# +
#####################################
## ##
## Generic cost fn Evaluation ##
## ##
#####################################
#Define the particular cost function
def evaluate_cost(t,u_t,S_t,I_t,Ic_t,cost_params):
#unpack cost params
Omega = cost_params
#evaluate costs from infections/death
nt = len(t)
pos = 2
cost_health = np.sum(Ic_t[pos,:,nt-1])
#evaluate costs from implementing controls:
cost_NPI = Omega*np.trapz(np.sum(((1-u_t)**2)*S_t,0),t)
return (cost_health + cost_NPI)
#Set a function to compute outcomes and evaluate cost function
def get_cost(x, gradx, IC, cost_params, evaluate_cost,print_figs = False):
#set up the time-dependent contact matrix based on control scheme
nu = int((len(x) + 2)/(M + 1))
t_con = np.append(np.append(0, x[:(nu - 2)]),Tf)
u_con = np.reshape(x[(nu - 2):],(M,nu))
def contactMatrix(t):
u = np.zeros(M)
if t == 0:
return contactMatrix_0(t) #this is needed because rescalings are done inside pyrosstsi based on this
else:
for i in range(M):
u[i] = np.interp(t,t_con,u_con[i,:])
return np.outer(u,u)*contactMatrix_0(t)
parameters['contactMatrix'] = contactMatrix
#let's not messs around with poorly sorted lists
for i in range(nu - 1):
if t_con[i + 1] < t_con[i]:
return float('inf')
#set up and solve
model = pyrosstsi.deterministic.Simulator(parameters)
data = model.simulate(IC)
#unpack and rescale simulation output
t = data['t']; S_t = data['S_t']; I_t = data['I_t']; Ic_t = data['Ic_t']
#compute cost function
nt = len(t)
u_t = np.zeros((M,nt))
for j in range(M):
u_t[j,:] = np.interp(t,t_con,u_con[j,:])
#Standard SIR plot
if print_figs:
plt.figure()
plt.plot(t,np.sum(S_t,0),'r', label = 'Susceptible') #all susceptible
plt.plot(t,np.sum(I_t,0),'b', label = 'Infected') #all Infected
plt.plot(t,np.sum(Ic_t[0,:,:],0),'g', label = subclasses[0]) #all recovered
plt.legend()
plt.xlabel('time (days)'); plt.xlim(0,Tf)
plt.ylabel('number of people'); plt.ylim(0,1)
return evaluate_cost(t,u_t,S_t,I_t,Ic_t,cost_params)
# +
###################################
## ##
## Optimize Controls ##
## ##
###################################
###################################
## Herd Immunity Example ##
###################################
#Define input parameters to the cost function:
Omega = 10**-3 #input parameter to trial cost function. Omega scales cost of lockdown to statistical value of life.
#low values represent cases where lockdowns are cheap (or disease is deadly)
#Generalizations should include age-structuring.
#bundle all input parameters together
cost_params = Omega
nu = 5 #how many interpolating points (including endpoints)
#set an initial guess (evenly spaced points, no intervention)
x0 = np.append(np.linspace(Tf/(nu - 1), Tf*(nu-2)/(nu-1),nu-2), np.ones(nu*M))
nx = len(x0)
#We use nlopt for optimization.
opt = nlopt.opt(nlopt.LN_COBYLA, nx)
opt.set_lower_bounds(np.zeros(nx))
opt.set_upper_bounds(np.append(Tf*np.ones(nu - 2),np.ones(nx - (nu - 2))))
opt.set_min_objective(lambda x, gradx: get_cost(x, gradx, IC, cost_params,evaluate_cost))
opt.set_xtol_rel(1e-3)#-4
x_opt = opt.optimize(x0)
minf = opt.last_optimum_value()
#group results and plot
t_con = np.append(np.append(0, x_opt[:(nu - 2)]),Tf)
u_con = np.reshape(x_opt[(nu - 2):],(M,nu))
plt.figure()
for i in range(M):
lbl = ['age group ', str(i)]
lbl = ''.join(lbl)
plt.plot(t_con,u_con[i,:], label = lbl)
plt.legend()
plt.xlabel('time (days)'); plt.xlim(0,Tf)
plt.ylabel('fraction of normal contacts'); plt.ylim(0,1)
print(get_cost(x_opt, [], IC, cost_params,evaluate_cost,True))
# +
###################################
## ##
## Optimize Controls ##
## ##
###################################
###################################
## Lockdown Example ##
###################################
#Define input parameters to the cost function:
Omega = 10**-3*2/T #input parameter to trial cost function. Omega scales cost of lockdown to statistical value of life.
#low values represent cases where lockdowns are cheap (or disease is deadly)
#Generalizations should include age-structuring.
#bundle all input parameters together
cost_params = Omega
nu = 5 #how many interpolating points (including endpoints)
#set an initial guess (evenly spaced points, no intervention)
x0 = np.append(np.linspace(Tf/(nu - 1), Tf*(nu-2)/(nu-1),nu-2), np.ones(nu*M))
nx = len(x0)
#We use nlopt for optimization.
opt = nlopt.opt(nlopt.LN_COBYLA, nx)
opt.set_lower_bounds(np.zeros(nx))
opt.set_upper_bounds(np.append(Tf*np.ones(nu - 2),np.ones(nx - (nu - 2))))
opt.set_min_objective(lambda x, gradx: get_cost(x, gradx, IC, cost_params,evaluate_cost))
opt.set_xtol_rel(1e-3)#-4
x_opt = opt.optimize(x0)
minf = opt.last_optimum_value()
#group results and plot
t_con = np.append(np.append(0, x_opt[:(nu - 2)]),Tf)
u_con = np.reshape(x_opt[(nu - 2):],(M,nu))
plt.figure()
for i in range(M):
lbl = ['age group ', str(i)]
lbl = ''.join(lbl)
plt.plot(t_con,u_con[i,:], label = lbl)
plt.legend()
plt.xlabel('time (days)'); plt.xlim(0,Tf)
plt.ylabel('fraction of normal contacts'); plt.ylim(0,1)
print(get_cost(x_opt, [], IC, cost_params,evaluate_cost,True))
# -
Omega*T
| examples/tsi/ex02_control.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import Image
# # Step 1: Browse the Ocean Marketplace Using the App
# To work with Ocean Protocol you need to set up a digital wallet. First, create a metamask account. There are many guides for doing this available online (e.g. [here](https://docs.oceanprotocol.com/tutorials/metamask-setup/)). Switch from the Ethereum Mainnet to the Rinkeby Test Network from your MetaMask wallet. Rinkeby is where you can test with no real transaction fees. Instead we use Rinkeby tokens that can be requested from faucets. Ocean tokens are needed to purchase datasets on the Ocean Marketplace. You can request Rinkeby Ocean from the Ocean faucet [here](https://faucet.rinkeby.oceanprotocol.com/). Making transactions on Ocean Marketplace (e.g. purchasing a dataset) also costs gas in ETH. We can request Rinkeby ETH from the faucet [here](https://faucets.chain.link/rinkeby).
#
# Now that we have some Rinkeby Ocean and ETH in our wallet, we can browse and purchase datasets on the [Ocean Marketplace](https://market.oceanprotocol.com/). When you enter a Web3 app in the browser you may need to sign in with your wallet. Make sure that you are browsing datasets on the Rinkeby network (see image below).
display(Image(filename='images/marketplace-network.png', width = 400))
# We will be working with a BCI dataset. You can find more information about the dataset [here](https://phas3.notion.site/EEG-dataset-details-680893f310d54fcaa85deb8b02f59c03). While the dataset is freely available online, we have uploaded it as a private dataset to practice the workflow. In future, we hope that the use of private datasets with BCI algorithms opens up new use cases. You can see the dataset on the Ocean marketplace [here](https://market.oceanprotocol.com/asset/did:op:3819B03ADeaBE19e14b8e514544Ba1a6E61ea510).
#
# In the traditional data science workflow, a data scientist downloads a dataset locally before running their code on it. In this scenario, the data comes to the code running on it. In contrast, private datasets on the marketplace cannot be downloaded. Instead, a data scientist can send code to the data itself where it runs the computations before returning the results. This is called Compute-to-Data (C2D), which is similar to Federated Learning. On the Ocean Marketplace, data providers should provide a sample of the data to give an idea of the quality of the data as well as the data interface through which it can be accessed.
#
# Download the sample data for the BCI dataset through the Marketplace GUI and inspect it (always make sure to only download samples from data providers that you trust!).
display(Image(filename='images/download-sample.png', width = 800))
# # Step 2: Browse the Ocean Marketplace Using the Ocean Python Library
# Now lets do the same through the Ocean Python library. We need to connect to the Ethereum network via an Ethereum node. We have set the config parameters for you in a config file. We are currently using [Infura](https://infura.io) for this but will be migrating to a full Ethereum Erigon node asap for increased decentralization.
# +
from ocean_lib.ocean.ocean import Ocean
from ocean_lib.config import Config
config = Config('config.ini')
ocean = Ocean(config)
print(f"config.network_url = '{config.network_url}'")
print(f"config.metadata_cache_uri = '{config.metadata_cache_uri}'")
print(f"config.provider_url = '{config.provider_url}'")
# -
# Next, export your private key from your metamask wallet. We highly recommend doing this with a wallet that has no real tokens in it (only Rinkeby tokens). For more info on private keys, see [this](https://github.com/oceanprotocol/ocean.py/blob/main/READMEs/wallets.md) from the ocean.py documentation:
#
# *The whole point of crypto wallets is to store private keys. Wallets have various tradeoffs of cost, convienence, and security. For example, hardware wallets tend to be more secure but less convenient and not free. It can also be useful to store private keys locally on your machine, for testing, though only with a small amount of value at stake (keep the risk down). Do not store your private keys on anything public, unless you want your tokens to disappear. For example, don't store your private keys in GitHub or expose them on frontend webpage code.*
#
# With this in mind, you can directly load your private key into the notebook. We use an envvar rather than storing it in code that might be pushed to a repo. We copy this in for a new session (you may need to restart the notebook server). Here's how we export an environmental variable using an example key (replace this with your actual private key.). From your console:
#
# ```console
# export MY_TEST_KEY=<KEY>
# ```
#
# Now initialize your wallet:
# +
import os
from ocean_lib.web3_internal.wallet import Wallet
wallet = Wallet(ocean.web3, private_key=os.getenv('MY_TEST_KEY'), transaction_timeout=20, block_confirmations=config.block_confirmations)
print(f"public address = '{wallet.address}'")
# -
# This should print out the public key of your metamask wallet. Check that it matches the one displayed in your metamask. Let's check the balances in our wallet. These should match the amount you received from the faucets (minus any you've since spent).
# +
from ocean_lib.web3_internal.currency import from_wei # wei is the smallest denomination of ether e.g. like cents
from ocean_lib.models.btoken import BToken #BToken is ERC20
OCEAN_token = BToken(ocean.web3, ocean.OCEAN_address)
print(f"ETH balance = '{from_wei(ocean.web3.eth.get_balance(wallet.address))}'")
print(f"OCEAN balance = '{from_wei(OCEAN_token.balanceOf(wallet.address))}'")
# -
# Now let's find the dataset. For the BCI dataset [here](https://market.oceanprotocol.com/asset/did:op:3819B03ADeaBE19e14b8e514544Ba1a6E61ea510), copy the decentralized identifier (DID).
display(Image(filename='images/did.png', width = 400))
# +
did = "did:op:3819B03ADeaBE19e14b8e514544Ba1a6E61ea510"
asset = ocean.assets.resolve(did)
print(f"Data token info = '{asset.values['dataTokenInfo']}'")
print(f"Dataset name = '{asset.metadata['main']['name']}'")
# -
# We can get the URL to the sample data from the associated metadata:
from pathlib import Path
sample_link = asset.metadata['additionalInformation']['links'][0]['url']
dataset_name = Path(sample_link).parts[2]
print(f"Sample link = '{sample_link}'")
data_dir = Path('data')
sample_dir = data_dir / dataset_name
# !mkdir -p {data_dir}
# Download the data from the command line:
# !wget {sample_link} -O {sample_dir}
# Now lets inspect the sample data. The data provider should provide this in the same format as the whole dataset. This helps us as data scientists to write scripts that run on both the sample data and the whole dataset. We call this the **interface** of the data.
import zipfile
with zipfile.ZipFile(sample_dir, 'r') as zip_ref:
zip_ref.extractall(str(data_dir))
print("Listing files...")
data_path = []
for root, dirs, files in os.walk(data_dir):
path = root.split(os.sep)
print((len(path) - 1) * '---', os.path.basename(root))
for file in files:
fn = os.path.join(root,file)
if fn.split('.')[-1] in ['feather']:
data_path.append(fn)
print(len(path) * '---', file)
import numpy as np
import pandas as pd
print("Reading files...")
datas = []
for path in data_path:
datas.append(np.array(pd.read_feather(data_path[0])))
data = np.stack(datas)
print("Data shape:", data.shape)
# There are 5 different sessions, where participants completed five tasks:
#
# 1. Resting-state: eyes open
# 2. Resting-state: eyes closed
# 3. Cognitive: subtraction task
# 4. Cognitive: listening to music
# 5. Cognitive: memory task.
#
# We included 61 sensors from the EEG net.
#
# You can find more information about the dataset [here](https://phas3.notion.site/EEG-dataset-details-680893f310d54fcaa85deb8b02f59c03).
# The next step is to write some code to run on the sample data before sending it to run on the full dataset.
| algorithms/eeg-classification/notebooks/1-download-sample-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #Introduction
# We're going to explore Pizza Franshise data set from http://college.cengage.com/mathematics/brase/understandable_statistics/7e/students/datasets/slr/frames/frame.html
#
# We want to know if we should be opening the next pizza franshise or not.
#
# In the following data
# X = annual franchise fee ($1000)
# Y = start up cost ($1000)
# for a pizza franchise
# %matplotlib inline
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
# #Data Exploring
df = pd.read_csv('slr12.csv', names=['annual', 'cost'], header=0)
df.describe()
df.head()
df.annual.plot()
df.cost.plot()
df.plot(kind='scatter', x='X', y='Y');
slope, intercept, r_value, p_value, std_err = stats.linregress(df['X'], df['Y'])
plt.plot(df['X'], df['Y'], 'o', label='Original data', markersize=2)
plt.plot(df['X'], slope*df['X'] + intercept, 'r', label='Fitted line')
plt.legend()
plt.show()
# So from this trend we can predict that if you annual fee is high then you need your startup cost will be high as well.
| pizza-franchise/Pizza Franchise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Explore those patients who have had repeat RRT events during an encounter
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime as datetime
from impala.util import as_pandas
from impala.dbapi import connect
plt.style.use('ggplot')
# %matplotlib notebook
conn = connect(host="mycluster.domain.com", port=my_impala_port_number)
cur = conn.cursor()
cur.execute("use my_db")
query_rrt_counts = """
SELECT enc.encntr_id, COUNT(1) AS count
FROM encounter enc
INNER JOIN clinical_event ce
ON enc.encntr_id = ce.encntr_id
WHERE enc.loc_facility_cd='633867'
AND enc.encntr_complete_dt_tm < 4e12
AND ce.event_cd='54411998'
AND ce.result_status_cd NOT IN ('31', '36')
AND ce.valid_until_dt_tm > 4e12
AND ce.event_class_cd not in ('654645')
AND enc.admit_type_cd != '0'
AND enc.encntr_type_class_cd = '391'
GROUP BY enc.encntr_id
ORDER BY enc.encntr_id;
"""
cur.execute(query_rrt_counts)
df = as_pandas(cur)
df.head()
# Number of encounters with multiple RRTs:
len(df[df['count']>1])
# to get just the encounter ids with counts >1
encs = df[df['count']>1]['encntr_id'].get_values()
encs
len(df)
df.sort_values(by='count', ascending=False).head(10)
df[df['count']>1].hist()
# ## Of the all the people with valid & complete RRT events, some of them have multiple RRT events.
# ## BUT: how many are truly multiple events, or mistaken entry, or bad data?
#
# ### For each encoutner with multiple RRT events:
# ### Check that the RRT events are reasonably spaced out
# ### Check that the RRT events are all after arrival & before enc.departure
# Print out the data related to the encounter with multiple RRT events, if the duration beteween RRT events is less than 1 hour,
# or if the start & end time of the encounter doesn't make sense related to the time of the RRT event.
count = 0
for enc in encs:
count+=1
print "count: {0}".format(count)
query = """
SELECT
ce.encntr_id
, ce.event_id
, ce.valid_until_dt_tm
, from_unixtime(CAST(ce.event_end_dt_tm / 1000 as bigint)) AS event_end
, ce.event_end_dt_tm
, from_unixtime(CAST(ce.valid_from_dt_tm / 1000 as bigint)) AS valid_from
, from_unixtime(CAST(enc.arrive_dt_tm/1000 as bigint)) AS enc_arrive
, enc.arrive_dt_tm
, COALESCE(tci.checkin_dt_tm, enc.arrive_dt_tm) AS check_in_time
, from_unixtime(CAST(COALESCE(tci.checkin_dt_tm
, enc.arrive_dt_tm)/1000 as bigint)) AS check_in
, from_unixtime(CAST(enc.depart_dt_tm/1000 as bigint)) AS enc_depart
, enc.depart_dt_tm
FROM clinical_event ce
INNER JOIN encounter enc
ON ce.encntr_id = enc.encntr_id
LEFT OUTER JOIN (
SELECT
ti.encntr_id AS encntr_id
, MIN(tc.checkin_dt_tm) AS checkin_dt_tm
FROM tracking_item ti
JOIN tracking_checkin tc ON ti.tracking_id = tc.tracking_id
GROUP BY ti.encntr_id
) tci
ON tci.encntr_id = enc.encntr_id
WHERE ce.event_cd = '54411998' AND ce.encntr_id='{0}'
AND ce.valid_until_dt_tm>4e12;
""".format(enc)
cur.execute(query)
dfenc = as_pandas(cur)
dfenc['dep-perf_hrs'] = ((dfenc['depart_dt_tm']-dfenc['event_end_dt_tm'])/3600000)
dfenc['perf-arr_hrs'] = ((dfenc['event_end_dt_tm']-dfenc['check_in_time'])/3600000)
# if there's a problem, print the dfenc
if any(dfenc['dep-perf_hrs']<0):
print dfenc
if any(dfenc['perf-arr_hrs']<0):
print dfenc
RRTtimes = sorted(dfenc['event_end_dt_tm'].get_values())
time_lastloop = 0
for RRTtime in RRTtimes:
if time_lastloop == 0:
time_lastloop = RRTtime
else:
if (RRTtime-time_lastloop)/3600000. < 1:
print dfenc
# +
# Several of these have elapsed intervals under 1 hour. ==> Very unlikely.
# Sometimes, info can get entered twice into the system, esp if there is a shift change.
# -
dfenc
| notebooks/EDA/multi_rrts[EDA].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Ontology
# +
# import libraries
import rdflib, pathlib, uuid
# +
# define graph and namespace
graph = rdflib.Graph()
name_wb = rdflib.Namespace('http://wikibas.se/ontology')
name_fiaf = rdflib.Namespace('https://www.fiafnet.org/')
# +
# useful functions
def make_claim(s, p, o):
claim_id = name_fiaf[f"resource/claim/{uuid.uuid4()}"]
graph.add((s, name_wb['#claim'], claim_id))
graph.add((claim_id, p, o))
return claim_id
# +
# basic construction
graph.add((name_fiaf['ontology/property/instance_of'], rdflib.RDFS.label, rdflib.Literal('instance of', lang='en')))
graph.add((name_fiaf['ontology/property/external_id'], rdflib.RDFS.label, rdflib.Literal('external id', lang='en')))
print(len(graph))
# +
# EN15907 framework
graph.add((name_fiaf['ontology/property/work'], rdflib.RDFS.label, rdflib.Literal('work', lang='en')))
graph.add((name_fiaf['ontology/item/work'], rdflib.RDFS.label, rdflib.Literal('Work', lang='en')))
graph.add((name_fiaf['ontology/property/manifestation_of'], rdflib.RDFS.label, rdflib.Literal('manifestation of', lang='en')))
graph.add((name_fiaf['ontology/property/manifestation'], rdflib.RDFS.label, rdflib.Literal('manifestation', lang='en')))
graph.add((name_fiaf['ontology/item/manifestation'], rdflib.RDFS.label, rdflib.Literal('Manifestation', lang='en')))
graph.add((name_fiaf['ontology/property/item_of'], rdflib.RDFS.label, rdflib.Literal('item of', lang='en')))
graph.add((name_fiaf['ontology/property/item'], rdflib.RDFS.label, rdflib.Literal('item', lang='en')))
graph.add((name_fiaf['ontology/item/item'], rdflib.RDFS.label, rdflib.Literal('Item', lang='en')))
print(len(graph))
# +
# titles
graph.add((name_fiaf['ontology/property/title'], rdflib.RDFS.label, rdflib.Literal('title', lang='en')))
graph.add((name_fiaf['ontology/property/title_type'], rdflib.RDFS.label, rdflib.Literal('title type', lang='en')))
graph.add((name_fiaf['ontology/item/title_type'], rdflib.RDFS.label, rdflib.Literal('Title Type', lang='en')))
make_claim(name_fiaf['ontology/item/original_title'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/title_type'])
graph.add((name_fiaf['ontology/item/original_title'], rdflib.RDFS.label, rdflib.Literal('Original Title', lang='en')))
graph.add((name_fiaf['ontology/item/original_title'], rdflib.RDFS.label, rdflib.Literal('Originaltitel', lang='sv')))
graph.add((name_fiaf['ontology/item/original_title'], rdflib.RDFS.label, rdflib.Literal('Nazev Origin', lang='cs')))
graph.add((name_fiaf['ontology/item/original_title'], rdflib.RDFS.label, rdflib.Literal('Originele Titel', lang='nl')))
graph.add((name_fiaf['ontology/item/original_title'], rdflib.RDFS.label, rdflib.Literal('Originaltitel', lang='de')))
make_claim(name_fiaf['ontology/item/work_title'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/title_type'])
graph.add((name_fiaf['ontology/item/work_title'], rdflib.RDFS.label, rdflib.Literal('Work Title', lang='en')))
print(len(graph))
# +
# events
graph.add((name_fiaf['ontology/property/event'], rdflib.RDFS.label, rdflib.Literal('event', lang='en')))
graph.add((name_fiaf['ontology/property/event_type'], rdflib.RDFS.label, rdflib.Literal('event type', lang='en')))
graph.add((name_fiaf['ontology/item/event_type'], rdflib.RDFS.label, rdflib.Literal('Event Type', lang='en')))
make_claim(name_fiaf['ontology/item/birth'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/event_type'])
graph.add((name_fiaf['ontology/item/birth'], rdflib.RDFS.label, rdflib.Literal('Birth', lang='en')))
graph.add((name_fiaf['ontology/item/birth'], rdflib.RDFS.label, rdflib.Literal('Geboorte', lang='nl')))
make_claim(name_fiaf['ontology/item/death'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/event_type'])
graph.add((name_fiaf['ontology/item/death'], rdflib.RDFS.label, rdflib.Literal('Death', lang='en')))
graph.add((name_fiaf['ontology/item/death'], rdflib.RDFS.label, rdflib.Literal('Sterf', lang='nl')))
make_claim(name_fiaf['ontology/item/publication'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/event_type'])
graph.add((name_fiaf['ontology/item/publication'], rdflib.RDFS.label, rdflib.Literal('Publication', lang='en')))
make_claim(name_fiaf['ontology/item/decision_copyright'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/event_type'])
graph.add((name_fiaf['ontology/item/decision_copyright'], rdflib.RDFS.label, rdflib.Literal('Decision (Copyright)', lang='en')))
make_claim(name_fiaf['ontology/item/decision_censorship'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/event_type'])
graph.add((name_fiaf['ontology/item/decision_censorship'], rdflib.RDFS.label, rdflib.Literal('Decision (Censorship)', lang='en')))
graph.add((name_fiaf['ontology/property/certificate'], rdflib.RDFS.label, rdflib.Literal('certificate', lang='en')))
print(len(graph))
# +
# location data
graph.add((name_fiaf['ontology/property/contributed_by'], rdflib.RDFS.label, rdflib.Literal('contributed by', lang='en')))
graph.add((name_fiaf['ontology/property/institution'], rdflib.RDFS.label, rdflib.Literal('institution', lang='en')))
graph.add((name_fiaf['ontology/property/held_at'], rdflib.RDFS.label, rdflib.Literal('held at', lang='en')))
graph.add((name_fiaf['ontology/property/located_in'], rdflib.RDFS.label, rdflib.Literal('located in', lang='en')))
graph.add((name_fiaf['ontology/item/holding_institution'], rdflib.RDFS.label, rdflib.Literal('Institution', lang='en')))
graph.add((name_fiaf['ontology/property/production_country'], rdflib.RDFS.label, rdflib.Literal('production country', lang='en')))
graph.add((name_fiaf['ontology/property/production_country'], rdflib.RDFS.label, rdflib.Literal('productieland', lang='nl')))
graph.add((name_fiaf['ontology/property/country'], rdflib.RDFS.label, rdflib.Literal('country', lang='en')))
graph.add((name_fiaf['ontology/property/country'], rdflib.RDFS.label, rdflib.Literal('vertaling', lang='nl')))
graph.add((name_fiaf['ontology/item/country'], rdflib.RDFS.label, rdflib.Literal('Country', lang='en')))
graph.add((name_fiaf['ontology/item/country'], rdflib.RDFS.label, rdflib.Literal('Vertaling', lang='nl')))
make_claim(name_fiaf['ontology/item/netherlands'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/country'])
graph.add((name_fiaf['ontology/item/netherlands'], rdflib.RDFS.label, rdflib.Literal('Netherlands', lang='en')))
graph.add((name_fiaf['ontology/item/netherlands'], rdflib.RDFS.label, rdflib.Literal('Nederland', lang='nl')))
make_claim(name_fiaf['ontology/item/germany'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/country'])
graph.add((name_fiaf['ontology/item/germany'], rdflib.RDFS.label, rdflib.Literal('Germany', lang='en')))
graph.add((name_fiaf['ontology/item/germany'], rdflib.RDFS.label, rdflib.Literal('Duitsland', lang='nl')))
make_claim(name_fiaf['ontology/item/usa'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/country'])
graph.add((name_fiaf['ontology/item/usa'], rdflib.RDFS.label, rdflib.Literal('United States of America', lang='en')))
graph.add((name_fiaf['ontology/item/usa'], rdflib.RDFS.label, rdflib.Literal('Verenigde Staten', lang='nl')))
make_claim(name_fiaf['ontology/item/uk'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/country'])
graph.add((name_fiaf['ontology/item/uk'], rdflib.RDFS.label, rdflib.Literal('United Kingdom', lang='en')))
make_claim(name_fiaf['ontology/item/czech_republic'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/country'])
for a,b in [('Česko', 'cs'),('Tschechien', 'de'),('Czech Republic', 'en'),('Tsjechië', 'nl'),('Tjeckien', 'sv')]:
graph.add((name_fiaf['ontology/item/czech_republic'], rdflib.RDFS.label, rdflib.Literal(a, lang=b)))
make_claim(name_fiaf['ontology/item/sweden'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/country'])
for a,b in [('Švédsko', 'cs'),('Schweden', 'de'),('Sweden', 'en'),('Zweden', 'nl'),('Sverige', 'sv')]:
graph.add((name_fiaf['ontology/item/sweden'], rdflib.RDFS.label, rdflib.Literal(a, lang=b)))
print(len(graph))
# +
# agent
graph.add((name_fiaf['ontology/property/agent'], rdflib.RDFS.label, rdflib.Literal('agent', lang='en')))
graph.add((name_fiaf['ontology/item/agent'], rdflib.RDFS.label, rdflib.Literal('Agent', lang='en')))
graph.add((name_fiaf['ontology/property/forename'], rdflib.RDFS.label, rdflib.Literal('forename', lang='en')))
graph.add((name_fiaf['ontology/property/forename'], rdflib.RDFS.label, rdflib.Literal('voornamen', lang='nl')))
graph.add((name_fiaf['ontology/property/surname'], rdflib.RDFS.label, rdflib.Literal('surname', lang='en')))
graph.add((name_fiaf['ontology/property/surname'], rdflib.RDFS.label, rdflib.Literal('familienaam', lang='nl')))
graph.add((name_fiaf['ontology/property/gender'], rdflib.RDFS.label, rdflib.Literal('gender', lang='en')))
graph.add((name_fiaf['ontology/property/gender'], rdflib.RDFS.label, rdflib.Literal('geslacht', lang='nl')))
graph.add((name_fiaf['ontology/item/gender'], rdflib.RDFS.label, rdflib.Literal('Gender', lang='en')))
graph.add((name_fiaf['ontology/item/gender'], rdflib.RDFS.label, rdflib.Literal('Geslacht', lang='nl')))
make_claim(name_fiaf['ontology/item/male'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/gender'])
graph.add((name_fiaf['ontology/item/male'], rdflib.RDFS.label, rdflib.Literal('Male', lang='en')))
graph.add((name_fiaf['ontology/item/male'], rdflib.RDFS.label, rdflib.Literal('Man', lang='nl')))
make_claim(name_fiaf['ontology/item/female'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/gender'])
graph.add((name_fiaf['ontology/item/female'], rdflib.RDFS.label, rdflib.Literal('Female', lang='en')))
graph.add((name_fiaf['ontology/item/female'], rdflib.RDFS.label, rdflib.Literal('Vrouw', lang='nl')))
print(len(graph))
# +
# agent type
graph.add((name_fiaf['ontology/property/agent_type'], rdflib.RDFS.label, rdflib.Literal('agent type', lang='en')))
graph.add((name_fiaf['ontology/item/agent_type'], rdflib.RDFS.label, rdflib.Literal('Agent Type', lang='en')))
make_claim(name_fiaf['ontology/item/cast'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/agent_type'])
graph.add((name_fiaf['ontology/item/cast'], rdflib.RDFS.label, rdflib.Literal('Cast', lang='en')))
graph.add((name_fiaf['ontology/item/cast'], rdflib.RDFS.label, rdflib.Literal('Roll', lang='sv')))
graph.add((name_fiaf['ontology/item/cast'], rdflib.RDFS.label, rdflib.Literal('Obsazení', lang='cs')))
graph.add((name_fiaf['ontology/item/cast'], rdflib.RDFS.label, rdflib.Literal('Acteur', lang='nl')))
graph.add((name_fiaf['ontology/item/cast'], rdflib.RDFS.label, rdflib.Literal('Darsteller', lang='de')))
make_claim(name_fiaf['ontology/item/director'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/agent_type'])
graph.add((name_fiaf['ontology/item/director'], rdflib.RDFS.label, rdflib.Literal('Director', lang='en')))
graph.add((name_fiaf['ontology/item/director'], rdflib.RDFS.label, rdflib.Literal('Regie', lang='nl')))
make_claim(name_fiaf['ontology/item/producer'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/agent_type'])
graph.add((name_fiaf['ontology/item/producer'], rdflib.RDFS.label, rdflib.Literal('Producer', lang='en')))
graph.add((name_fiaf['ontology/item/producer'], rdflib.RDFS.label, rdflib.Literal('Producent', lang='nl')))
make_claim(name_fiaf['ontology/item/cinematographer'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/agent_type'])
graph.add((name_fiaf['ontology/item/cinematographer'], rdflib.RDFS.label, rdflib.Literal('Cinematographer', lang='en')))
graph.add((name_fiaf['ontology/item/cinematographer'], rdflib.RDFS.label, rdflib.Literal('Camera', lang='nl')))
make_claim(name_fiaf['ontology/item/editor'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/agent_type'])
graph.add((name_fiaf['ontology/item/editor'], rdflib.RDFS.label, rdflib.Literal('Editor', lang='en')))
graph.add((name_fiaf['ontology/item/editor'], rdflib.RDFS.label, rdflib.Literal('Montage', lang='nl')))
make_claim(name_fiaf['ontology/item/screenwriter'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/agent_type'])
graph.add((name_fiaf['ontology/item/screenwriter'], rdflib.RDFS.label, rdflib.Literal('Screenwriter', lang='en')))
graph.add((name_fiaf['ontology/item/screenwriter'], rdflib.RDFS.label, rdflib.Literal('Scenarioschrijver', lang='nl')))
make_claim(name_fiaf['ontology/item/composer'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/agent_type'])
graph.add((name_fiaf['ontology/item/composer'], rdflib.RDFS.label, rdflib.Literal('Composer', lang='en')))
graph.add((name_fiaf['ontology/item/composer'], rdflib.RDFS.label, rdflib.Literal('Componist', lang='nl')))
print(len(graph))
# +
# item access
graph.add((name_fiaf['ontology/property/access'], rdflib.RDFS.label, rdflib.Literal('access', lang='en'))) # property
graph.add((name_fiaf['ontology/item/access_type'], rdflib.RDFS.label, rdflib.Literal('Access Type', lang='en'))) # item
make_claim(name_fiaf['ontology/item/moma_only'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/access_type'])
graph.add((name_fiaf['ontology/item/moma_only'], rdflib.RDFS.label, rdflib.Literal('MoMA use only', lang='en')))
make_claim(name_fiaf['ontology/item/viewing'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/access_type'])
graph.add((name_fiaf['ontology/item/viewing'], rdflib.RDFS.label, rdflib.Literal('Viewing', lang='en')))
make_claim(name_fiaf['ontology/item/master'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/access_type'])
graph.add((name_fiaf['ontology/item/master'], rdflib.RDFS.label, rdflib.Literal('Master', lang='en')))
make_claim(name_fiaf['ontology/item/restricted'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/access_type'])
graph.add((name_fiaf['ontology/item/restricted'], rdflib.RDFS.label, rdflib.Literal('Restricted', lang='en')))
make_claim(name_fiaf['ontology/item/lending'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/access_type'])
graph.add((name_fiaf['ontology/item/lending'], rdflib.RDFS.label, rdflib.Literal('Lending', lang='en')))
print(len(graph))
# +
# item carrier
graph.add((name_fiaf['ontology/property/carrier'], rdflib.RDFS.label, rdflib.Literal('carrier', lang='en'))) # property
graph.add((name_fiaf['ontology/item/carrier_type'], rdflib.RDFS.label, rdflib.Literal('Carrier Type', lang='en'))) # item
make_claim(name_fiaf['ontology/item/film'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/carrier_type'])
graph.add((name_fiaf['ontology/item/film'], rdflib.RDFS.label, rdflib.Literal('Film', lang='en')))
make_claim(name_fiaf['ontology/item/sound_tape'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/carrier_type'])
graph.add((name_fiaf['ontology/item/sound_tape'], rdflib.RDFS.label, rdflib.Literal('Soundtape', lang='en')))
make_claim(name_fiaf['ontology/item/video_tape'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/carrier_type'])
graph.add((name_fiaf['ontology/item/video_tape'], rdflib.RDFS.label, rdflib.Literal('Videotape', lang='en')))
make_claim(name_fiaf['ontology/item/digital'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/carrier_type'])
graph.add((name_fiaf['ontology/item/digital'], rdflib.RDFS.label, rdflib.Literal('Digital', lang='en')))
make_claim(name_fiaf['ontology/item/disc'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/carrier_type'])
graph.add((name_fiaf['ontology/item/disc'], rdflib.RDFS.label, rdflib.Literal('Disc', lang='en')))
print(len(graph))
# +
# item specific carrier
graph.add((name_fiaf['ontology/property/specific_carrier'], rdflib.RDFS.label, rdflib.Literal('specific carrier', lang='en'))) # property
graph.add((name_fiaf['ontology/item/specific_carrier_type'], rdflib.RDFS.label, rdflib.Literal('Specific Carrier Type', lang='en'))) # item
make_claim(name_fiaf['ontology/item/16mm'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/16mm'], rdflib.RDFS.label, rdflib.Literal('16mm', lang='en')))
make_claim(name_fiaf['ontology/item/35mm'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/35mm'], rdflib.RDFS.label, rdflib.Literal('35mm', lang='en')))
make_claim(name_fiaf['ontology/item/9mm'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/9mm'], rdflib.RDFS.label, rdflib.Literal('9.5mm', lang='en')))
make_claim(name_fiaf['ontology/item/quarter-inch'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/quarter-inch'], rdflib.RDFS.label, rdflib.Literal('1/4-inch', lang='en')))
make_claim(name_fiaf['ontology/item/digibeta'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/digibeta'], rdflib.RDFS.label, rdflib.Literal('Digibeta', lang='en')))
make_claim(name_fiaf['ontology/item/vhs'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/vhs'], rdflib.RDFS.label, rdflib.Literal('VHS', lang='en')))
make_claim(name_fiaf['ontology/item/umatic'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/umatic'], rdflib.RDFS.label, rdflib.Literal('U-matic', lang='en')))
make_claim(name_fiaf['ontology/item/hdcam'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/hdcam'], rdflib.RDFS.label, rdflib.Literal('HDcam', lang='en')))
make_claim(name_fiaf['ontology/item/hi8'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/hi8'], rdflib.RDFS.label, rdflib.Literal('Hi8', lang='en')))
make_claim(name_fiaf['ontology/item/d2'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/d2'], rdflib.RDFS.label, rdflib.Literal('D2', lang='en')))
make_claim(name_fiaf['ontology/item/d5'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/d5'], rdflib.RDFS.label, rdflib.Literal('D5', lang='en')))
make_claim(name_fiaf['ontology/item/betamax'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/betamax'], rdflib.RDFS.label, rdflib.Literal('Betamax', lang='en')))
make_claim(name_fiaf['ontology/item/betacamsp'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/betacamsp'], rdflib.RDFS.label, rdflib.Literal('Betacam SP', lang='en')))
make_claim(name_fiaf['ontology/item/dvd'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/dvd'], rdflib.RDFS.label, rdflib.Literal('DVD', lang='en')))
make_claim(name_fiaf['ontology/item/mxf'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/specific_carrier_type'])
graph.add((name_fiaf['ontology/item/mxf'], rdflib.RDFS.label, rdflib.Literal('MXF', lang='en')))
print(len(graph))
# +
# item element
graph.add((name_fiaf['ontology/property/element'], rdflib.RDFS.label, rdflib.Literal('element', lang='en'))) # property
graph.add((name_fiaf['ontology/item/element_type'], rdflib.RDFS.label, rdflib.Literal('Element Type', lang='en'))) # item
make_claim(name_fiaf['ontology/item/print'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/print'], rdflib.RDFS.label, rdflib.Literal('Print', lang='en')))
make_claim(name_fiaf['ontology/item/duplicate_negative'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/duplicate_negative'], rdflib.RDFS.label, rdflib.Literal('Duplicate Negative', lang='en')))
make_claim(name_fiaf['ontology/item/duplicate_positive'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/duplicate_positive'], rdflib.RDFS.label, rdflib.Literal('Duplicate Positive', lang='en')))
make_claim(name_fiaf['ontology/item/negative'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/negative'], rdflib.RDFS.label, rdflib.Literal('Negative', lang='en')))
make_claim(name_fiaf['ontology/item/duplicate_reversal'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/duplicate_reversal'], rdflib.RDFS.label, rdflib.Literal('Duplicate reversal', lang='en')))
make_claim(name_fiaf['ontology/item/track_negative'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/track_negative'], rdflib.RDFS.label, rdflib.Literal('Track negative', lang='en')))
make_claim(name_fiaf['ontology/item/fine_grain_master'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/fine_grain_master'], rdflib.RDFS.label, rdflib.Literal('Fine grain master', lang='en')))
make_claim(name_fiaf['ontology/item/magnetic_track'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/magnetic_track'], rdflib.RDFS.label, rdflib.Literal('Magnetic track', lang='en')))
make_claim(name_fiaf['ontology/item/sound_negative'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/sound_negative'], rdflib.RDFS.label, rdflib.Literal('Sound Negative', lang='en')))
make_claim(name_fiaf['ontology/item/original_negative'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/original_negative'], rdflib.RDFS.label, rdflib.Literal('Original Negative', lang='en')))
make_claim(name_fiaf['ontology/item/work_print'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/element_type'])
graph.add((name_fiaf['ontology/item/work_print'], rdflib.RDFS.label, rdflib.Literal('Work Print', lang='en')))
print(len(graph))
# +
# item base
graph.add((name_fiaf['ontology/property/base'], rdflib.RDFS.label, rdflib.Literal('base', lang='en'))) # property
graph.add((name_fiaf['ontology/item/base_type'], rdflib.RDFS.label, rdflib.Literal('Base Type', lang='en'))) # item
make_claim(name_fiaf['ontology/item/acetate'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/base_type'])
graph.add((name_fiaf['ontology/item/acetate'], rdflib.RDFS.label, rdflib.Literal('Acetate', lang='en')))
make_claim(name_fiaf['ontology/item/polyester'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/base_type'])
graph.add((name_fiaf['ontology/item/polyester'], rdflib.RDFS.label, rdflib.Literal('Polyester', lang='en')))
make_claim(name_fiaf['ontology/item/diacetate'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/base_type'])
graph.add((name_fiaf['ontology/item/diacetate'], rdflib.RDFS.label, rdflib.Literal('Diacetate', lang='en')))
make_claim(name_fiaf['ontology/item/nitrate'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/base_type'])
graph.add((name_fiaf['ontology/item/nitrate'], rdflib.RDFS.label, rdflib.Literal('Nitrate', lang='en')))
print(len(graph))
# +
# item colour
graph.add((name_fiaf['ontology/property/colour'], rdflib.RDFS.label, rdflib.Literal('colour', lang='en'))) # property
graph.add((name_fiaf['ontology/item/colour_type'], rdflib.RDFS.label, rdflib.Literal('Colour Type', lang='en'))) # item
make_claim(name_fiaf['ontology/item/black_and_white'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/colour_type'])
graph.add((name_fiaf['ontology/item/black_and_white'], rdflib.RDFS.label, rdflib.Literal('Black and White', lang='en')))
make_claim(name_fiaf['ontology/item/colour'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/colour_type'])
graph.add((name_fiaf['ontology/item/colour'], rdflib.RDFS.label, rdflib.Literal('Colour', lang='en')))
make_claim(name_fiaf['ontology/item/black_and_white_and_colour'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/colour_type'])
graph.add((name_fiaf['ontology/item/black_and_white_and_colour'], rdflib.RDFS.label, rdflib.Literal('(Black and White) and Colour', lang='en')))
make_claim(name_fiaf['ontology/item/tinted'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/colour_type'])
graph.add((name_fiaf['ontology/item/tinted'], rdflib.RDFS.label, rdflib.Literal('Tinted', lang='en')))
print(len(graph))
# +
# item sound
graph.add((name_fiaf['ontology/property/sound'], rdflib.RDFS.label, rdflib.Literal('sound', lang='en'))) # property
graph.add((name_fiaf['ontology/item/sound_type'], rdflib.RDFS.label, rdflib.Literal('Sound Type', lang='en'))) # item
make_claim(name_fiaf['ontology/item/silent'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/sound_type'])
graph.add((name_fiaf['ontology/item/silent'], rdflib.RDFS.label, rdflib.Literal('Silent', lang='en')))
make_claim(name_fiaf['ontology/item/sound'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/sound_type'])
graph.add((name_fiaf['ontology/item/sound'], rdflib.RDFS.label, rdflib.Literal('Sound', lang='en')))
print(len(graph))
# +
# item sound format
graph.add((name_fiaf['ontology/property/sound_format'], rdflib.RDFS.label, rdflib.Literal('sound format', lang='en'))) # property
graph.add((name_fiaf['ontology/item/sound_format_type'], rdflib.RDFS.label, rdflib.Literal('Sound Format Type', lang='en'))) # item
make_claim(name_fiaf['ontology/item/optical'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/sound_format_type'])
graph.add((name_fiaf['ontology/item/optical'], rdflib.RDFS.label, rdflib.Literal('Optical', lang='en')))
make_claim(name_fiaf['ontology/item/magnetic'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/sound_format_type'])
graph.add((name_fiaf['ontology/item/magnetic'], rdflib.RDFS.label, rdflib.Literal('Magnetic', lang='en')))
make_claim(name_fiaf['ontology/item/analogue'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/sound_format_type'])
graph.add((name_fiaf['ontology/item/analogue'], rdflib.RDFS.label, rdflib.Literal('Analogue', lang='en')))
print(len(graph))
# +
# item languages
graph.add((name_fiaf['ontology/property/intertitles_language'], rdflib.RDFS.label, rdflib.Literal('intertitles language', lang='en'))) # property
graph.add((name_fiaf['ontology/property/maintitles_language'], rdflib.RDFS.label, rdflib.Literal('maintitles language', lang='en'))) # property
graph.add((name_fiaf['ontology/item/language'], rdflib.RDFS.label, rdflib.Literal('Language', lang='en'))) # item
make_claim(name_fiaf['ontology/item/english'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/language'])
graph.add((name_fiaf['ontology/item/english'], rdflib.RDFS.label, rdflib.Literal('English', lang='en')))
make_claim(name_fiaf['ontology/item/german'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/language'])
graph.add((name_fiaf['ontology/item/german'], rdflib.RDFS.label, rdflib.Literal('German', lang='en')))
make_claim(name_fiaf['ontology/item/french'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/language'])
graph.add((name_fiaf['ontology/item/french'], rdflib.RDFS.label, rdflib.Literal('French', lang='en')))
make_claim(name_fiaf['ontology/item/dutch'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/language'])
graph.add((name_fiaf['ontology/item/dutch'], rdflib.RDFS.label, rdflib.Literal('Dutch', lang='en')))
print(len(graph))
# +
# item extent
graph.add((name_fiaf['ontology/property/extent_feet'], rdflib.RDFS.label, rdflib.Literal('extent (feet)', lang='en')))
graph.add((name_fiaf['ontology/property/extent_metres'], rdflib.RDFS.label, rdflib.Literal('extent (metres)', lang='en')))
graph.add((name_fiaf['ontology/property/duration'], rdflib.RDFS.label, rdflib.Literal('duration (minutes)', lang='en')))
print(len(graph))
# +
graph.serialize(destination=str(pathlib.Path.cwd() / 'fiaf.ttl'), format="turtle")
print(len(graph))
| 1-ontology/.ipynb_checkpoints/fiaf-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import syft as sy
import numpy as np
from syft.core.adp.entity import Entity
# ### Loading the dataset
data = pd.read_csv("../datasets/it - feb 2021.csv")[0:100]
data.head()
# ### Logging into the domain
it = sy.login(email="<EMAIL>",
password="<PASSWORD>",
port=8082)
# ### Upload the dataset to Domain node
# +
# We will upload only the first few rows
# All these three columns are of `int` type
# NOTE: casting this tensor as np.int32 is REALLY IMPORTANT. We need to create flags for this or something
data_batch = ((np.array(list(data['Trade Value (US$)'])) / 100000)[0:10]).astype(np.int32)
trade_partners = ((list(data['Partner'])))[0:10]
entities = list()
for i in range(len(trade_partners)):
entities.append(Entity(name="Other Asia, nes"))
# Upload a private dataset to the Domain object, as the root owner
sampled_italy_dataset = sy.Tensor(data_batch)
sampled_italy_dataset.public_shape = sampled_italy_dataset.shape
sampled_italy_dataset = sampled_italy_dataset.private(0, 3, entities="bob").tag("trade_flow")
# -
it.load_dataset(
assets={"Italy Trade": sampled_italy_dataset},
name="Italy Trade Data - First few rows",
description="""A collection of reports from iStat's statistics
bureau about how much it thinks it imports and exports from other countries.""",
)
it.datasets
# ### Create a Data Scientist User
it.users.create(
**{
"name": "<NAME>",
"email": "<EMAIL>",
"password": "<PASSWORD>",
"budget":200
}
)
# ### Accept/Deny Requests to the Domain
it.requests.pandas
it.requests[-1].accept()
| notebooks/trade_demo/demo/Data Owner - Italy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ml-gpu
# language: python
# name: ml-gpu
# ---
# # Capstone Project - Car Accident Severity
# ## 1. Introduction / Business Problem
# ### Stakeholders:
# * Seattle Department of Transportation
# * Seattle Police Department - Traffic Enforcement Section
# * Car Drivers in Seattle
# Car accident is serious problem in Seattle. During the first six months of 2019, 101 people were seriously injured or killed in 98 collisions on Seattle streets, which is the highest number of crashes in the first half of a year since 2010, according to the data provided by the Seattle Department of Transportation (SDOT). The accident victims and their families, the insurance conpanies, health care personal and even the ordinary people, are affected by traffic accidents in many ways. Therefore, predicting the possibility and severity of a car accident based on the weather, road conditions and other factors is important. For drivers, they would drive more carefully or even change their travel if they are able to. For the police and traffic departments, they can put up some warning signs when there is a high traffic accident risk.
# ## 2. Data
# The dataset used in this analysis is provided by SDOT Traffic Management Division, Traffic Records Group. It includes all types of collisions at the intersection or mid-block of a segment recorded by the Traffic Records from 2004 to Present.
#
# The dataset contains 194,673 records with 38 attributes: 16 numerical and 22 categorical (see the figures below for the detailed descriptions). Among these 38 features, 6 of them have a large portion of null value, which are INTKEY, EXCEPTRSNCODE, EXCEPTRSNDESC, INATTENTIONIND, PEDROWNOTGRNT, SPEEDING. We will perform imputation on these feature during the feature engineering step. The target for this prediction task is the "Accident Severity", which are represented by three attributes: SEVERITYCODE, SEVERITYCODE.1, SEVERITYDESC, containing two sets of values: 1, 1, Property Damage Only Collision and 2, 2, Injury Collision. The target is unevenly distributed, with ~70% Property Damage Collisions Only and ~30% Injury Collisions. Therefore, target weights will be applied in the modeling step.
# 
#
# 
#
# 
# ### 2.1 Feature Selection
# 1. Target: Choose the 'SEVERITYCODE' (A code that corresponds to the severity of the collision: 1-Property Damage; 2-Injury), or 'SEVERITYCODE.1', 'SEVERITYDESC', as the target.
# 2. Features: Choose features related to the weather, road and drivers' condition.
# * 'INATTENTIONIND': Whether or not collision was due to inattention.
# * 'UNDERINFL': Whether or not a driver involved was under the influence of drugs or alcohol.
# * 'WEATHER': A description of the weather conditions during the time of the collision.
# * 'ROADCOND': The condition of the road during the collision.
# * 'LIGHTCOND': The light conditions during the collision.
# * 'SPEEDING': Whether or not speeding was a factor in the collision.
# ## 3. Methodology
# ### 3.1 Exploratory Data Analysis
# +
# import python library
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
pd.set_option('display.max_columns', None)
# load dataset
df = pd.read_csv('Data-Collisions.csv', low_memory=False)
# +
# target distribution
target_feature = 'SEVERITYCODE'
plt.figure(figsize=(20, 4))
sns.countplot(x=target_feature, data=df).set_title(target_feature)
plt.show()
# +
# count distributions
selected_features = ['INATTENTIONIND', 'UNDERINFL', 'WEATHER', 'ROADCOND', 'LIGHTCOND', 'SPEEDING']
for col in selected_features:
plt.figure(figsize=(20, 4))
sns.countplot(x=col, data=df.fillna('nan')).set_title(col)
plt.show()
# -
# mean response on target
for col in selected_features:
plt.figure(figsize=(20, 4))
sns.barplot(x=col, y=target_feature, data=df.fillna('nan')).set_title(col)
plt.show()
# Insights:
# * The target is unevenly distributed, with ~70% Property Damage (1) and ~30% Injury (2).
# * Inattention ('INATTENTIONIND'='Y) will increase the collision severity.
# * Drivers under the influence of drugs or alcohol ('UNDERINFL'='Y' or '1') will increase the collision severity.
# * Speeding ('SPEEDING'='Y') will increase the collision severity.
# * Bad weather ('WEATHER'), road ('ROADCOND') and light ('LIGHTCOND') conditions will also slightly increase the collision severity.
# ### 3.2 Feature Encoding
# +
# select features
df_feature = df[selected_features + [target_feature]].copy()
# binary featurtes
df_feature['SEVERITYCODE'] = df['SEVERITYCODE'].map({1: 0, 2: 1})
df_feature['INATTENTIONIND'] = df['INATTENTIONIND'].map({'Y': 1, np.nan: 0})
df_feature['SPEEDING'] = df['SPEEDING'].map({'Y': 1, np.nan: 0})
# label encoding
df_feature['UNDERINFL'] = df['UNDERINFL'].map({'N': 0, '0': 0, 'Y': 2, '1': 2, np.nan: 1})
# one hot encoding
dummy_feature = ['WEATHER', 'ROADCOND', 'LIGHTCOND']
df_feature = pd.concat([df_feature.drop(columns=dummy_feature), pd.get_dummies(df[dummy_feature].fillna('Unknown'))], axis=1)
# show processed dataset
print(f"shape: {df_feature.shape}")
df_feature.head()
# -
# ### 3.3 Feature Correlation Analysis
# plot heatmap for feature correlation matrix
plt.figure(figsize=(10, 8))
sns.heatmap(df_feature.drop(columns=target_feature).corr(), vmin=-1, vmax=+1)
plt.show()
# High correlation observed between weather, road and light conditions, which is reasonable since the road and light conditions are both related to the weather conditon.
# ### 3.4 Model
# #### 3.4.1 Model Selection
# 1. Logistic Regression: simple baseline model
# 2. Decision Tree: simple tree based model
# 3. Random Forest: advanced tree based model using the bagging method
# 4. XGBoost: advanced tree based model using the boosting method
# #### 3.4.2 Feature Preprocessing
# +
from imblearn.over_sampling import SMOTE
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
# extract feature & target
X = df_feature.drop(columns=target_feature)
y = df_feature[target_feature]
# feature standardization
X = StandardScaler().fit_transform(X)
# oversampling
X, y = SMOTE().fit_resample(X, y)
# train & test splitting
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, test_size=0.20, random_state=2020)
# show the processed data
print(f"training set:\n{y_train.value_counts(normalize=True)}")
print(f"test set:\n{y_test.value_counts(normalize=True)}")
# -
# #### 3.4.3 Logistic Regression
# +
from sklearn.metrics import roc_auc_score, plot_roc_curve
from sklearn.linear_model import LogisticRegression
model_lr = LogisticRegression(C=0.01, max_iter=1000, solver='lbfgs', class_weight='balanced', random_state=2020)
model_lr.fit(X_train, y_train)
y_pred = model_lr.predict_proba(X_test)[:, 1]
print(f"Test ROC-AUC: {roc_auc_score(y_test, y_pred)}")
plot_roc_curve(model_lr, X_test, y_test)
# -
# #### 3.4.4 Decision Tree
# +
from sklearn.tree import DecisionTreeClassifier
model_dt = DecisionTreeClassifier(criterion='entropy', max_depth=7, min_weight_fraction_leaf=0.001, min_samples_split=10,class_weight='balanced', random_state=2020)
model_dt.fit(X_train, y_train)
y_pred = model_dt.predict_proba(X_test)[:, 1]
print(f"Test ROC-AUC: {roc_auc_score(y_test, y_pred)}")
plot_roc_curve(model_dt, X_test, y_test)
# -
# #### 3.4.5 Random Forest
# +
from sklearn.ensemble import RandomForestClassifier
model_rf = RandomForestClassifier(criterion='entropy', n_estimators=100, max_depth=9, min_weight_fraction_leaf=0.000005, class_weight='balanced', random_state=2020, n_jobs=4)
model_rf.fit(X_train, y_train)
y_pred = model_rf.predict_proba(X_test)[:, 1]
print(f"Test ROC-AUC: {roc_auc_score(y_test, y_pred)}")
plot_roc_curve(model_rf, X_test, y_test)
# -
# #### 3.4.6 XGBoost
# +
from xgboost import XGBClassifier
params = {
"tree_method" : "gpu_hist",
"predictor" : "gpu_predictor",
"booster" : "gbtree",
"objective" : "binary:logistic",
"eval_metric" : "auc",
"n_estimators" : 200,
"max_depth" : 6,
"min_child_weight" : 15,
"learning_rate" : 0.04,
"gamma" : 0,
"subsample" : 1.00,
"colsample_bytree" : 0.60,
"scale_pos_weight" : (y == 0).sum() / (y == 1).sum(),
"verbosity" : 1,
"random_state" : 2020,
}
model_xgb = XGBClassifier(**params)
model_xgb.fit(X_train, y_train)
y_pred = model_xgb.predict_proba(X_test)[:, 1]
print(f"Test ROC-AUC: {roc_auc_score(y_test, y_pred)}")
plot_roc_curve(model_xgb, X_test, y_test)
# -
# ## 4. Results
# +
from sklearn.metrics import precision_score, recall_score, f1_score, accuracy_score, roc_auc_score, log_loss
from sklearn.metrics import confusion_matrix, classification_report
models = {
'Logistic Regression' : model_lr,
'DecisionTree' : model_dt,
'Random Forest' : model_rf,
'XGBoost' : model_xgb,
}
df_result = pd.DataFrame(index=models.keys())
for key, val in models.items():
print(f"{key}\n{classification_report(y_test, val.predict(X_test))}")
df_result.loc[key, 'Precision'] = precision_score(y_test, val.predict(X_test))
df_result.loc[key, 'Recall'] = recall_score(y_test, val.predict(X_test))
df_result.loc[key, 'F1-score'] = f1_score(y_test, val.predict(X_test))
df_result.loc[key, 'Accuracy'] = accuracy_score(y_test, val.predict(X_test))
df_result.loc[key, 'ROC-AUC'] = roc_auc_score(y_test, val.predict_proba(X_test)[:, 1])
df_result.loc[key, 'logloss'] = log_loss(y_test, val.predict_proba(X_test)[:, 1])
df_result
# -
# * Overall, the 4 models performs similarly. But the models metrics are not very good.
# * The Random Forest and XGBoost perform slightly better than the Logistic Regression and Decision Tree.
# ## 5. Summary
# * We have performed Exploratory Data Analysis, Feature Engineering and Modeling for the project. Use 6 features related to the weather, road and driver's conditions to predict the severity of an accident.
# * We examined 6 metrics: Precision, Recall ,F1-score, Accuracy, ROC-AUC, logloss, on 4 models: Logistic Regression, Decision Tree, Random Forest and XGBoost, where the XGboost model performs the best, with a Precision score of 0.548157, a Recall score of 0.839616, a F1 score of 0.663281, an accuracy score of 0.573763, a ROC-AUC score of 0.598076, and a logloss score of 0.661718.
# * The overall performance of all the models are not very good. For improvement, we can add more features from the original dataset to our models.
# ## 6. Recommendations
# * To the transportation department:
# * Improve the road and light conditions for those roads where most severe accidents take place.
# * Put up signs to remind driver to stay conscious, and slow down when weather condition is not optimal.
# * Send our more police officers to monitor the speeding issue.
# * To the car drivers:
# * Pay attention to the change of road and weather condition.
# * Always be conscious and focused, and obey the traffic rules.
| capstone3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Download Ros na Rún season 2
#
# > "Dataset"
#
# - toc: false
# - branch: master
# - badges: false
# - comments: true
# - hidden: true
# - categories: [kaggle, irish, rosnarun, dataset, unlabelled]
# Original on [Kaggle](https://www.kaggle.com/jimregan/ros-na-run-s2)
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 13.566819, "end_time": "2021-08-01T21:28:51.217282", "exception": false, "start_time": "2021-08-01T21:28:37.650463", "status": "completed"} tags=[]
# %%capture
# !pip install youtube-dl
# + papermill={"duration": 2337.861561, "end_time": "2021-08-01T22:07:49.083260", "exception": false, "start_time": "2021-08-01T21:28:51.221699", "status": "completed"} tags=[]
# !youtube-dl -f bestaudio PLtVSQEQG0xVHeyao6vZyaY3kXGfbFFiAk
| _notebooks/2021-08-01-ros-na-run-s2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Quadrupole Example
#
# Simple quadrupole example
#
# +
# Useful for debugging
# %load_ext autoreload
# %autoreload 2
import numpy as np
import os
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
# +
# locate the drift template
from impact import Impact
ifile = '../templates/quadrupole/ImpactT.in'
os.path.exists(ifile)
# -
# calculate gamma*beta
mec2 = 0.51099895000e6 # eV
Etot = 6e6 #eV
gamma = Etot/mec2
GB = np.sqrt(gamma**2 -1)
GB
# # Use Impact's built-in Gaussian particle generator
# +
I = Impact(ifile)
I.header['Np'] = 100000
I.header['Nx'] = 32
I.header['Ny'] = 32
I.header['Nz'] = 32
I.header['Dt'] = 10e-12
I.header['Bcurr'] = 0
I.header['zmu2'] = GB
# set normal and skew quads
I.ele['CQ01']['b1_gradient'] = 0.00714 # T/m
I.ele['SQ01']['b1_gradient'] = 0
# -
# # Single particle tracking
# Track
I2 = I.copy()
I2.configure()
ele = I2.ele['CQ01']
ele
# Estimate for angle change for a 6 MeV/c momentum particle, offset by 1 mm.
ele['b1_gradient']*ele['L_effective']*299792458 / 6e6 * .001
P2 = I2.track1(s=2.2, z0 = 0, x0=0.001, pz0=6e6)
P2.xp
I2.plot('mean_x')
# # Track beam
# Regular and Skew quads
I.run()
I.output['stats'].keys()
PI = I.particles['initial_particles']
PF = I.particles['final_particles']
PI['sigma_y']
# Compare these.
key1 = 'mean_z'
key2 = 'sigma_x'
units1 = str(I.units(key1))
units2 = str(I.units(key2))
plt.xlabel(key1+f' ({units1})')
plt.ylabel(key2+f' ({units2})')
plt.plot(I.stat(key1), I.stat(key2))
plt.scatter(
[I.particles[name][key1] for name in I.particles],
[I.particles[name][key2] for name in I.particles], color='red')
# Compare these.
key1 = 'mean_z'
key2 = 'sigma_x'
units1 = str(I.units(key1))
units2 = str(I.units(key2))
plt.xlabel(key1+f' ({units1})')
plt.ylabel(key2+f' ({units2})')
plt.plot(I.stat(key1), I.stat(key2))
plt.scatter(
[I.particles[name][key1] for name in I.particles],
[I.particles[name][key2] for name in I.particles], color='red')
key2 = 'sigma_y'
plt.plot(I.stat(key1), I.stat(key2))
plt.scatter(
[I.particles[name][key1] for name in I.particles],
[I.particles[name][key2] for name in I.particles], color='green')
PF.plot('x', 'y')
PF.plot('delta_z', 'delta_pz')
| examples/elements/quadrupole.ipynb |