code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Problem 1 : Take a variable x and print "Even" if the number is divisible by 2, otherwise print "Odd".
# +
x=4
if(x%2==0):
print("Even")
else:
print("Odd")
# -
# # Problem 2 : Take a variable y and print "Grade A" if y is greater than 90, "Grade B" if y is greater than 60 but less than or equal to 90 and "Grade F" Otherwise.
# +
y=90
if(y>90):
print("Grade A")
elif(y>60):
print("Grade B")
else:
print("Grade F")
# -
| 13. conditional_statements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Tweepy
#
# *An easy-to-use Python library for accessing the Twitter API.*
#
# http://www.tweepy.org/
# + [markdown] slideshow={"slide_type": "slide"}
# ## Installing
#
# `pip install tweepy`
# + [markdown] slideshow={"slide_type": "slide"}
# ## Twitter App
#
# https://apps.twitter.com/
#
# Create an app and save:
#
# - Consumer key
# - Consumer secret
# - Access token
# - Access token secret
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Store Secrets Securely
#
# *Don't commit them to a public repo!*
#
# I set them via environment variables and then access via `os.environ`.
# + slideshow={"slide_type": "subslide"}
# Load keys, secrets, settings
import os
ENV = os.environ
CONSUMER_KEY = ENV.get('IOTX_CONSUMER_KEY')
CONSUMER_SECRET = ENV.get('IOTX_CONSUMER_SECRET')
ACCESS_TOKEN = ENV.get('IOTX_ACCESS_TOKEN')
ACCESS_TOKEN_SECRET = ENV.get('IOTX_ACCESS_TOKEN_SECRET')
USERNAME = ENV.get('IOTX_USERNAME')
USER_ID = ENV.get('IOTX_USER_ID')
print(USERNAME)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Twitter APIs
#
# ### REST vs Streaming
#
# https://dev.twitter.com/docs
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Twitter REST API
#
# * Search
# * Tweet
# * Get information
#
# https://dev.twitter.com/rest/public
# + slideshow={"slide_type": "slide"}
import tweepy
auth = tweepy.OAuthHandler(CONSUMER_KEY, CONSUMER_SECRET)
auth.set_access_token(ACCESS_TOKEN, ACCESS_TOKEN_SECRET)
api = tweepy.API(auth)
public_tweets = api.home_timeline(count=3)
for tweet in public_tweets:
print(tweet.text)
# + slideshow={"slide_type": "slide"}
# Models
user = api.get_user('clepy')
print(user.screen_name)
print(dir(user))
# + slideshow={"slide_type": "slide"}
# Tweet!
status = api.update_status("I'm at @CLEPY!")
print(status.id)
print(status.text)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Streaming API
#
# Real-time streaming of:
#
# * Searches
# * Mentions
# * Lists
# * Timeline
#
# https://dev.twitter.com/streaming/public
# + slideshow={"slide_type": "slide"}
# Subclass StreamListener and define on_status method
class MyStreamListener(tweepy.StreamListener):
def on_status(self, status):
print("@{0}: {1}".format(status.author.screen_name, status.text))
# + slideshow={"slide_type": "slide"}
myStream = tweepy.Stream(auth = api.auth, listener=MyStreamListener())
# + slideshow={"slide_type": "slide"}
try:
myStream.filter(track=['#clepy'])
except KeyboardInterrupt:
print('Interrupted...')
except tweepy.error.TweepError:
myStream.disconnect()
print('Disconnected. Try again!')
# + [markdown] slideshow={"slide_type": "slide"}
# See also:
#
# "Small Batch Artisanal Bots: Let's Make Friends" by <NAME> at PyCon & PyOhio
#
# https://www.youtube.com/watch?v=pDS_LWgjMgg
# + slideshow={"slide_type": "slide"}
| Tweepy/Tweepy.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
display(HTML("<style>.container { width:100% !important; }</style>"))
# #### MIT license (c) 2019 by <NAME>
# #### Jupyter notebook written in Julia 1.1.0. It implements on a CPUs the numerical program for computing the exercise boundary and the price of an American-style call option described in Sec. 18.4 of "Stochastic Methods in Asset Pricing" (pp. 514-516). The same program is implemented in Python in Appendix B.3 of SMAP.
# Note the formula $\int_a^bf(t)dt={1\over2}(b-a)\int_{-1}^{+1}f\bigl({1\over2}(b-a)s+{1\over2}(a+b)\bigr)d s$ to use with FastGaussQuadratures.
## the number of grid points over the time domain is set below
## 5000 is excessive and is meant to test Julia's capabilities
total_grid_no=5000 #this number must be divisible by pnmb
# Introduce the parameters in the model.
KK=40.0; # strike
sigma=0.3; # volatility
delta=0.07; # dividend
rr=0.02; # interest
TT=0.5; # time to maturity
DLT=TT/(total_grid_no) # distance between two grid points in the time domain
ABSC=range(0.0,length=total_grid_no+1,stop=TT) # the entire grid on the time domain
VAL=[max(KK,KK*(rr/delta)) for x in ABSC]; # first guess for the exercise boundary
# +
using SpecialFunctions
using Interpolations
using Roots
using FastGaussQuadrature
nodes,weights=gausslegendre( 200 );
function EUcall(S::Float64,t::Float64,K::Float64,σ::Float64,r::Float64,δ::Float64)
local v1,v2
v1=(δ-r+σ^2/2.0)*t+log(K/S)
v2=(-δ + r + σ^2/2.0)*t - log(K/S)
v3=σ*sqrt(2.0*t)
return -K*(exp(-t*r)/2)+K*(exp(-t*r)/2)*erf(v1/v3)+S*(exp(-t*δ)/2)*erf(v2/v3)+S*(exp(-t*δ)/2)
end;
function F(ϵ::Int64,t::Float64,u::Float64,v::Float64,r::Float64,δ::Float64,σ::Float64)
v1=(r-δ+ϵ*σ^2/2)*(u-t)-log(v)
v2=σ*sqrt(2*(u-t))
return 1.0+erf(v1/v2)
end;
function ah(t::Float64,z::Float64,r::Float64,δ::Float64,σ::Float64,f)
return (exp(-δ*(z-t))*(δ/2)*F(1,t,z,f(z)/f(t),r,δ,σ))
end;
function bh(t::Float64,z::Float64,r::Float64,δ::Float64,σ::Float64,K::Float64,f)
return (exp(-r*(z-t))*(r*K/2)*F(-1,t,z,f(z)/f(t),r,δ,σ))
end;
function make_grid0(step::Float64,size::Int64)
return 0.0:step:(step+(size-1)*step)
end;
# -
function mainF(start_iter::Int64,nmb_of_iter::Int64,conv_tol::Float64,K::Float64,σ::Float64,δ::Float64,r::Float64,T::Float64,Δ::Float64,nmb_grd_pnts::Int64,vls::Array{Float64,1},nds::Array{Float64,1},wghts::Array{Float64,1})
local no_iter,conv_check,absc,absc0,valPrev,val,loc,f
absc=range(0.0,length=nmb_grd_pnts+1,stop=T)
absc0=range(0.0,length=nmb_grd_pnts,stop=(T-Δ));
val=vls;
f=CubicSplineInterpolation(absc,val,extrapolation_bc = Interpolations.Line())
no_iter=start_iter;
conv_check=100.0
while no_iter<nmb_of_iter&&conv_check>conv_tol
no_iter+=1
loc=[max(K,K*(r/δ))]
for ttt=Iterators.reverse(absc0)
an=[(1/2)*(T-ttt)*ah(ttt,(1/2)*(T-ttt)*s+(1/2)*(ttt+T),r,δ,σ,f) for s in nodes]
bn=[(1/2)*(T-ttt)*bh(ttt,(1/2)*(T-ttt)*s+(1/2)*(ttt+T),r,δ,σ,K,f) for s in nodes]
aaa=weights'*an
bbb=weights'*bn;
LRT=find_zero(x->x-K-EUcall(x,T-ttt,K,σ,r,δ)-aaa*x+bbb,(K-10,K+20));
pushfirst!(loc,LRT)
end
valPrev=val
val=loc
f=CubicSplineInterpolation(absc,val,extrapolation_bc = Interpolations.Line())
conv_check=maximum(abs.(valPrev-val))
end
return absc,val,conv_check,no_iter
end
# Run the main routine. The third argument is an upper bound on the total iterations to run.
# The last two arguments control FastGaussQuadrature.
#
# masterF(prnmb,start_iter,nmb_of_iter,conv_tol,K,σ,δ,r,T,Δ,no_grd_pnts,vls,nds,wghts)
#
# mainF(start_iter, nmb_of_iter, conv_tol, $K$, $\sigma$, $\delta$, $r$, $T$, $\Delta$, no_grd_pnts, vls, nds, wghts)
@time ABSC,VAL,conv,iterations=mainF(0,100,1.0e-5,KK,sigma,delta,rr,TT,DLT,total_grid_no,VAL,nodes,weights);
# The call to 'masterF' can be repeated with the most recent VAL and the second argument set to the number of already performed iterations.
conv,iterations
f=CubicSplineInterpolation(ABSC,VAL,extrapolation_bc = Interpolations.Line());
using Plots
pyplot()
plotgrid=ABSC[1]:.001:ABSC[end];
pval=[f(x) for x in plotgrid];
plot(plotgrid,pval,label="exercise boundary")
xlabel!("real time")
ylabel!("underlying spot price")
f(0)
# The price of the EU call option at the money ($S_0=K$) with $T={1\over2}$:
EUcall(KK,TT,KK,sigma,rr,delta)
# The early exercise premium for at the money option at $t=0$ is:
an=[(0.5*(TT-0.0))*exp(-delta*((1/2)*(TT-0.0)*s+(1/2)*(0.0+TT)-0.0))*(KK*delta/2)*F(1,0.0,(1/2)*(TT-0.0)*s+(1/2)*(0.0+TT),f((1/2)*(TT-0.0)*s+(1/2)*(0.0+TT))/KK,rr,delta,sigma) for s in nodes]
bn=[(0.5*(TT-0.0))*exp(-rr*((1/2)*(TT-0.0)*s+(1/2)*(0.0+TT)-0.0))*(rr*KK/2)*F(-1,0.0,(1/2)*(TT-0.0)*s+(1/2)*(0.0+TT),f((1/2)*(TT-0.0)*s+(1/2)*(0.0+TT))/KK,rr,delta,sigma) for s in nodes]
aaa=weights'*an
bbb=weights'*bn;
EEP=aaa-bbb
EEP
# The price of an American at the money call with 6 months to expiry is:
EUcall(KK,TT,KK,sigma,rr,delta)+EEP
| Num_Program_US_Call_Julia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bellman Equation for MRPs
# In this exercise we will learn how to find state values for a simple MRPs using the scipy library.
# 
import numpy as np
from scipy import linalg
# Define the transition probability matrix
# define the Transition Probability Matrix
n_states = 3
P = np.zeros((n_states, n_states), np.float)
P[0, 1] = 0.7
P[0, 2] = 0.3
P[1, 0] = 0.5
P[1, 2] = 0.5
P[2, 1] = 0.1
P[2, 2] = 0.9
P
# Check that the sum over columns is exactly equal to 1, being a probability matrix.
# the sum over columns is 1 for each row being a probability matrix
assert((np.sum(P, axis=1) == 1).all())
# We can calculate the expected immediate reward for each state using the reward matrix and the transition probability
# define the reward matrix
R = np.zeros((n_states, n_states), np.float)
R[0, 1] = 1
R[0, 2] = 10
R[1, 0] = 0
R[1, 2] = 1
R[2, 1] = -1
R[2, 2] = 10
# calculate expected reward for each state by multiplying the probability matrix for each reward
R_expected = np.sum(P * R, axis=1, keepdims=True)
# The matrix R_expected
R_expected
# The R_expected vector is the expected immediate reward foe each state.
# State 1 has an expected reward of 3.7 that is exactly equal to 0.7 * 1 + 0.3*10.
# The same for state 2 and so on.
# define the discount factor
gamma = 0.9
# We are ready to solve the Bellman Equation
#
# $$
# (I - \gamma P)V = R_{expected}
# $$
#
# Casting this to a linear equation we have
# $$
# Ax = b
# $$
#
# Where
# $$
# A = (I - \gamma P)
# $$
# And
# $$
# b = R
# $$
# Now it is possible to solve the Bellman Equation
A = np.eye(n_states) - gamma * P
B = R_expected
# solve using scipy linalg
V = linalg.solve(A, B)
V
# The vector V represents the value for each state. The state 3 has the highest value.
| Chapter02/Exercise02_01/Exercise02_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Analyze_stable_points - evaluate biase in DEMs using selected unchanged points
#
# These points were picked on hopefully stable points in mostly flat places: docks, lawns, bare spots in middens. Also, the yurt roofs. Typically, 3 to 5 points were picked on most features.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
def pcoord(x, y):
"""
Convert x, y to polar coordinates r, az (geographic convention)
r,az = pcoord(x, y)
"""
r = np.sqrt( x**2 + y**2 )
az=np.degrees( np.arctan2(x, y) )
# az[where(az<0.)[0]] += 360.
az = (az+360.)%360.
return r, az
def xycoord(r, az):
"""
Convert r, az [degrees, geographic convention] to rectangular coordinates
x,y = xycoord(r, az)
"""
x = r * np.sin(np.radians(az))
y = r * np.cos(np.radians(az))
return x, y
# -
df=pd.read_csv("C:\\crs\\proj\\2019_DorianOBX\\Santa_Cruz_Products\\stable_points\\All_points_dunex_lidar.csv",header = 0)
df
col = df.loc[: , "Aug":"Nov"]
df['mean']=col.mean(axis=1)
df['std']=col.std(axis=1)
df['gnd50 anom']=df['Gnd_50']-df['mean']
df['first95 anom']=df['First_95']-df['mean']
df['Aug anom']=df['Aug']-df['mean']
df['Sep anom']=df['Sep']-df['mean']
df['Oct anom']=df['Oct']-df['mean']
df['Nov anom']=df['Nov']-df['mean']
r,az = pcoord( df['X'].values, df['Y'].values)
xr, yr = xycoord( r, az+42.)
df['alongshore']=xr-2870000.
df
df_anom = df.loc[:,"gnd50 anom":"Nov anom"].copy()
df_anom
print(df_anom.mean())
print(df_anom.std())
plt.plot(df['alongshore'],df['Aug anom'],'o',alpha=.5,label='Aug')
plt.plot(df['alongshore'],df['Sep anom'],'o',alpha=.5,label='Sep')
plt.plot(df['alongshore'],df['Oct anom'],'o',alpha=.5,label='Oct')
plt.plot(df['alongshore'],df['Nov anom'],'o',alpha=.5,label='Nov')
plt.legend()
plt.ylabel('Anomoly (m)')
plt.xlabel('Alongshore Distance (m)')
plt.plot(df['Aug'],df['Aug anom'],'o',alpha=.5,label='Aug')
plt.plot(df['Sep'],df['Sep anom'],'o',alpha=.5,label='Sep')
plt.plot(df['Oct'],df['Oct anom'],'o',alpha=.5,label='Oct')
plt.plot(df['Nov'],df['Nov anom'],'o',alpha=.5,label='Nov')
plt.legend()
plt.ylabel('Anomoly (m)')
plt.xlabel('Elevation (m NAVD88)')
plt.scatter(df['X'].values,df['Y'].values,s=50,c=df['Aug anom'].values,alpha=.4)
# boxplot of anomolies
boxprops = dict(linestyle='-', linewidth=3, color='k')
medianprops = dict(linestyle='-', linewidth=3, color='k')
bp=df_anom.boxplot(figsize=(6,5),grid=True,boxprops=boxprops, medianprops=medianprops)
plt.ylabel('Difference from Four SfM Map Mean (m)')
bp.set_xticklabels(['Gnd 50','First 95','Aug','Sep','Oct','Nov'])
plt.savefig('unchanged_pts_boxplot.png',dpi=200)
| Analyze_stable_points.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PyRosetta.notebooks]
# language: python
# name: conda-env-PyRosetta.notebooks-py
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains material from [PyRosetta](https://RosettaCommons.github.io/PyRosetta.notebooks);
# content is available [on Github](https://github.com/RosettaCommons/PyRosetta.notebooks.git).*
# <!--NAVIGATION-->
# < [Running Rosetta in Parallel](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.00-Running-PyRosetta-in-Parallel.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Distributed computation example: miniprotein design](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.02-PyData-miniprotein-design.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.01-PyData-ddG-pssm.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
# # Distributed analysis example: exhaustive ddG PSSM
#
# ## Notes
# This tutorial will walk you through how to generate an exhaustive ddG PSSM in PyRosetta using the PyData stack for analysis and distributed computing.
#
# This Jupyter notebook uses parallelization and is not meant to be executed within a Google Colab environment.
#
# ## Setup
# Please see setup instructions in Chapter 16.00
#
# ## Citation
# [Integration of the Rosetta Suite with the Python Software Stack via reproducible packaging and core programming interfaces for distributed simulation](https://doi.org/10.1002/pro.3721)
#
# <NAME>, <NAME>, <NAME>
#
# ## Manual
# Documentation for the `pyrosetta.distributed` namespace can be found here: https://nbviewer.jupyter.org/github/proteininnovation/Rosetta-PyData_Integration/blob/master/distributed_overview.ipynb
import sys
if 'google.colab' in sys.modules:
print("This Jupyter notebook uses parallelization and is therefore not set up for the Google Colab environment.")
sys.exit(0)
import logging
logging.basicConfig(level=logging.INFO)
import pandas
import seaborn
import matplotlib
import Bio.SeqUtils
import Bio.Data.IUPACData as IUPACData
import pyrosetta
import pyrosetta.distributed.io as io
import pyrosetta.distributed.packed_pose as packed_pose
import pyrosetta.distributed.tasks.rosetta_scripts as rosetta_scripts
import pyrosetta.distributed.tasks.score as score
# +
import os,sys,platform
platform.python_version()
# -
# ## Create test pose, initialize rosetta and pack
input_protocol = """
<ROSETTASCRIPTS>
<TASKOPERATIONS>
<RestrictToRepacking name="only_pack"/>
</TASKOPERATIONS>
<MOVERS>
<PackRotamersMover name="pack" task_operations="only_pack" />
</MOVERS>
<PROTOCOLS>
<Add mover="pack"/>
</PROTOCOLS>
</ROSETTASCRIPTS>
"""
input_relax = rosetta_scripts.SingleoutputRosettaScriptsTask(input_protocol)
# Syntax check via setup
input_relax.setup()
raw_input_pose = score.ScorePoseTask()(io.pose_from_sequence("TESTESTEST"))
input_pose = input_relax(raw_input_pose)
# ## Perform exhaustive point mutation and pack
def mutate_residue(input_pose, res_index, new_aa, res_label = None):
import pyrosetta.rosetta.core.pose as pose
work_pose = packed_pose.to_pose(input_pose)
# Annotate strucure with reslabel, for use in downstream protocol
# Add parameters as score, for use in downstream analysis
if res_label:
work_pose.pdb_info().add_reslabel(res_index, res_label)
pose.setPoseExtraScore(work_pose, "mutation_index", res_index)
pose.setPoseExtraScore(work_pose, "mutation_aa", new_aa)
if len(new_aa) == 1:
new_aa = str.upper(Bio.SeqUtils.seq3(new_aa))
assert new_aa in map(str.upper, IUPACData.protein_letters_3to1)
protocol = """
<ROSETTASCRIPTS>
<MOVERS>
<MutateResidue name="mutate" new_res="%(new_aa)s" target="%(res_index)i" />
</MOVERS>
<PROTOCOLS>
<Add mover_name="mutate"/>
</PROTOCOLS>
</ROSETTASCRIPTS>
""" % locals()
return rosetta_scripts.SingleoutputRosettaScriptsTask(protocol)(work_pose)
# +
refine = """
<ROSETTASCRIPTS>
<RESIDUE_SELECTORS>
<ResiduePDBInfoHasLabel name="mutation" property="mutation" />
<Not name="not_neighbor">
<Neighborhood selector="mutation" distance="12.0" />
</Not>
</RESIDUE_SELECTORS>
<TASKOPERATIONS>
<RestrictToRepacking name="only_pack"/>
<OperateOnResidueSubset name="only_repack_neighbors" selector="not_neighbor" >
<PreventRepackingRLT/>
</OperateOnResidueSubset>
</TASKOPERATIONS>
<MOVERS>
<PackRotamersMover name="pack_area" task_operations="only_pack,only_repack_neighbors" />
</MOVERS>
<PROTOCOLS>
<Add mover="pack_area"/>
</PROTOCOLS>
</ROSETTASCRIPTS>
"""
refine_mutation = rosetta_scripts.SingleoutputRosettaScriptsTask(refine)
# -
# # Mutation and pack
# ## Job distribution via `multiprocessing`
from multiprocessing import Pool
import itertools
with pyrosetta.distributed.utility.log.LoggingContext(logging.getLogger("rosetta"), level=logging.WARN):
with Pool() as p:
work = [
(input_pose, i, aa, "mutation")
for i, aa in itertools.product(range(1, len(packed_pose.to_pose(input_pose).residues) + 1), IUPACData.protein_letters)
]
logging.info("mutating")
mutations = p.starmap(mutate_residue, work)
# ## Job distribution via `dask`
if not os.getenv("DEBUG"):
import dask.distributed
cluster = dask.distributed.LocalCluster(n_workers=1, threads_per_worker=1)
client = dask.distributed.Client(cluster)
refinement_tasks = [client.submit(refine_mutation, mutant) for mutant in mutations]
logging.info("refining")
refinements = [task.result() for task in refinement_tasks]
client.close()
cluster.close()
# ## Analysis of delta score
if not os.getenv("DEBUG"):
result_frame = pandas.DataFrame.from_records(packed_pose.to_dict(refinements))
result_frame["delta_total_score"] = result_frame["total_score"] - input_pose.scores["total_score"]
result_frame["mutation_index"] = list(map(int, result_frame["mutation_index"]))
if not os.getenv("DEBUG"):
matplotlib.rcParams['figure.figsize'] = [24.0, 8.0]
seaborn.heatmap(
result_frame.pivot("mutation_aa", "mutation_index", "delta_total_score"),
cmap="RdBu_r", center=0, vmax=50)
# <!--NAVIGATION-->
# < [Running Rosetta in Parallel](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.00-Running-PyRosetta-in-Parallel.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Distributed computation example: miniprotein design](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.02-PyData-miniprotein-design.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.01-PyData-ddG-pssm.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
| notebooks/16.01-PyData-ddG-pssm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Announcments
#
# * Start the CP if you haven't already
# * We have a course tutor, <NAME> <EMAIL>
# +
# %matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (12, 9)
plt.rcParams["font.size"] = 18
# -
# ## Binary Nuclear Reactions
#
# ### Learning Objectives:
#
# - Connect concepts in particle collisions and decay to binary reactions
# - Categorize nuclear reactions using standard nomenclature
# - Apply conservation of nucleons to binary nuclear reactions
# - Formulate Q value equations for binary nuclear reactions
# - Apply conservation of energy and linear momentum to scattering
# - Apply coulombic threshold
# - Apply kinematic threshold
# - Determine when coulombic and kinematic thresholds apply or do not
# ## Recall from Weeks 3 & 4
#
# To acheive these objectives, we need to recall 3 major themes from weeks three and four.
#
# ### 1: Compare Exothermic and Endothermic reactions
#
# - In **_exothermic_** or **_exoergic_** reactions, energy is **emitted** ($Q>0$)
# - In **_endothermic_** or **_endoergic_** reactions, energy is **absorbed** ($Q<0$)
#
#
# 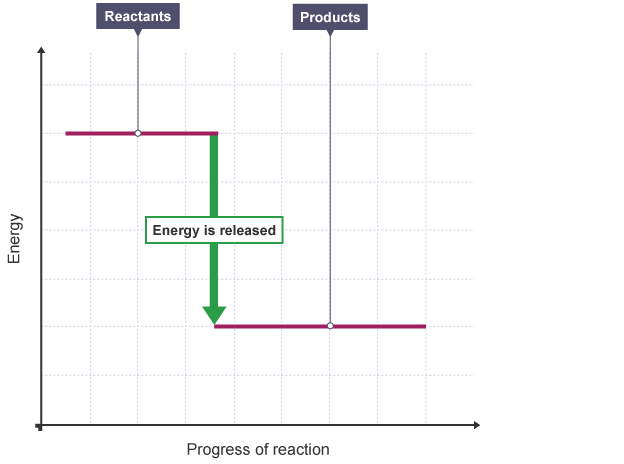
# 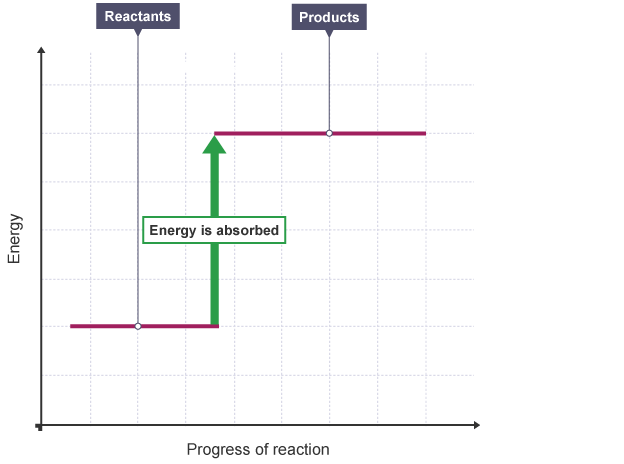
# <center>(credit: BBC)</center>
#
# ### 2: Relate energy and mass $E=mc^2$
#
# When the masses of reactions change, this is tied to a change in energy from whence we learn the Q value.
# This change in mass is equivalent to a change in energy because **$E=mc^2$**
#
# \begin{align}
# A + B + \cdots &\rightarrow C + D + \cdots\\
# \mbox{(reactants)} &\rightarrow \mbox{(products)}\\
# \implies \Delta M &= (\mbox{reactants}) - (\mbox{products})\\
# &= (M_A + M_B + \cdots) - (M_C + M_D + \cdots)\\
# \implies \Delta E &= \left[(M_A + M_B + \cdots) - (M_C + M_D + \cdots)\right]c^2\\
# \end{align}
#
#
# ### 3: Apply conservation of energy and momentum to scattering collisions
#
# Conservation of total energy and linear momentum can inform Compton scattering reactions. X-rays scattered from electrons had a change in wavelength $\Delta\lambda = \lambda' - \lambda$ proportional to $(1-\cos{\theta_s})$
#
# <a title="JabberWok [GFDL (http://www.gnu.org/copyleft/fdl.html) or CC-BY-SA-3.0 (http://creativecommons.org/licenses/by-sa/3.0/)], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Compton-scattering.svg"><img width="128" alt="Compton-scattering" src="https://upload.wikimedia.org/wikipedia/commons/thumb/e/e3/Compton-scattering.svg/128px-Compton-scattering.svg.png"></a>
#
# We used the law of cosines:
#
# \begin{align}
# p_e^2 &= p_\lambda^2 + p_{\lambda'}^2 - 2p_\lambda p_{\lambda'}\cos{\theta_s}
# \end{align}
#
#
# And we also used conservation of energy:
# \begin{align}
# p_\lambda c+m_ec^2 &= p_{\lambda'}c + mc^2\\
# \mbox{where }&\\
# m_e&=\mbox{rest mass of the electron}\\
# m &= \mbox{relativistic electron mass after scattering}
# \end{align}
#
# Combining these with our understanding of photon energy ($E=h\nu=pc$) gives:
#
# \begin{align}
# \lambda' - \lambda &= \frac{h}{m_ec}(1-\cos{\theta_s})\\
# \implies \frac{1}{E'} - \frac{1}{E} &= \frac{1}{m_ec^2}(1-\cos{\theta_s})\\
# \implies E' &= \left[\frac{1}{E} + \frac{1}{m_ec^2}(1-\cos{\theta_s})\right]^{-1}\\
# \end{align}
# ## More Types of Reactions
#
# Previously we were interested in fundamental particles striking one another (e.g. the electron and proton in Compton scattering) or nuclei emitting such particles (e.g. $\beta^\pm$ decay).
#
# **Today:** We are interested in myriad additional reactants and/or products. In particular, we're interested in:
#
# - neutron absorption and production reactions
# - _binary, two-product nuclear reactions_ in which two products emerge with new energies after the collision.
# ## Reaction Nomenclature
#
# **Transfer Reactions:** Nucleons (1 or 2) are transferred between the projectile and product.
#
# **Scattering reactions:** The projectile and product emerge from a collision with the same identities as when they started, exchanging only kinetic energy.
#
# **Knockout reactions:** The projectile directly interacts with the target nucleus and is re-emitted **along with** nucleons from the target nucleus.
#
# **capture reactions:** The projectile is absorbed, typically exciting the nucleus. The excited nucleus may emit that energy decaying via photon emission.
#
# **nuclear photoeffect:** A photon projectile liberates a nucleon from the target nucleus.
#
# ### Think Pair Share : categorize these reactions
#
# One example of each of the above appears below. Use the definitions to categorize them.
#
# - $(n, n)$
# - $(n, \gamma)$
# - $(n, 2n)$
# - $(\gamma, n)$
# - $(\alpha, n)$
#
# ## Binary, two-product nuclear reactions
#
# **Two initial nuclei collide to form two product nuclei.**
#
# \begin{align}
# ^{A_1}_{Z_1}X_1 + ^{A_2}_{Z_2}X_2 \longrightarrow ^{A_3}_{Z_3}X_3 + ^{A_4}_{Z_4}X_4
# \end{align}
#
# #### Applying Conservation of Neutrons and Protons
#
# The total number of nucleons is always conserved.
# If the `______________` force is not involved, we can also apply this conservation separately.
#
#
# In most binary, two-product nuclear reactions, this is the case, so the number of protons and neutrons are conserved. Thus:
#
# \begin{align}
# Z_1 + Z_2 = Z_3 + Z_4\\
# A_1 + A_2 = A_3 + A_4
# \end{align}
#
# Apply this to the following:
#
# \begin{align}
# ^{3}_{1}H + ^{16}_{8}O \longrightarrow \left(X\right)^* \longrightarrow ^{16}_{7}N + ^{A_4}_{Z_4}X_4
# \end{align}
#
# ### Think Pair Share:
#
# What are :
#
# - $A_4$
# - $Z_4$
# - $X_4$?
#
# - Bonus: What is $\left(X\right)^*$?
#
# #### Applying conservation of mass and energy.
#
# The Q-value calculation is the same as it has been before.
# The Q value represents the `________` in kinetic energy and, equivalently, a `________` in the rest masses.
#
# \begin{align}
# Q &= E_y + E_Y − E_x − E_X \\
# &= (m_x + m_X − m_y − m_Y )c^2\\
# &= \left(m\left(^{A_1}_{Z_1}X_1\right) + m\left(^{A_2}_{Z_2}X_2\right) - m\left(^{A_3}_{Z_3}X_3\right) - m\left(^{A_4}_{Z_4}X_4\right)\right)c^2\\
# \end{align}
#
# If proton numbers are conserved (true for everything but electron capture or reactions involving the weak force.), we can use the approximation that $m(X) = M(X)$.
#
# \begin{align}
# Q &= E_y + E_Y − E_x − E_X \\
# &= (m_x + m_X − m_y − m_Y )c^2\\
# &= (M_x + M_X − M_y − M_Y )c^2\\
# &= \left(M\left(^{A_1}_{Z_1}X_1\right) + M\left(^{A_2}_{Z_2}X_2\right) - M\left(^{A_3}_{Z_3}X_3\right) - M\left(^{A_4}_{Z_4}X_4\right)\right)c^2\\
# \end{align}
# +
def q(m_reactants, m_products):
"""Returns Q
Parameters
----------
m_reactants: list (of doubles)
the masses of the reactant atoms [amu]
m_products : list (of doubles)
the masses of the product atoms [amu]
"""
amu_to_mev = 931.5 # MeV/amu conversion
m_difference = sum(m_reactants) - sum(m_products)
return m_difference*amu_to_mev
# Look up the masses:
h_3_mass = 3.0160492675
o_16_mass = 15.9949146221
he_3_mass = 3.0160293097
n_16_mass = 16.0061014
m_react = [h_3_mass, o_16_mass]
m_prods = [he_3_mass, n_16_mass]
print("Q: ", q(m_react, m_prods))
# -
# #### Applying conservation of linear momentum
#
# Let's get back to collision kinematics.
#
# First, we'll assume the target nucleus ($X_2$) is initially at rest.
#
#
# 
# # Kinematic Threshold
#
# Relying on a combination of kinetic energies $E_i$ and corresponding linear momenta:
#
# \begin{align}
# p_i = \sqrt{2m_iE_i}
# \end{align}
#
# We can determine that some reactions aren't possible without a certain minimum quantity of kinetic energy.
#
# The solution to $E_3$ can become nonphysical if :
#
# - $\cos{\theta_3} < 0$
# - $Q < 0$
# - $m_4 - m_1 < 0$
#
# ## For Exoergic Reactions ($Q>0$)
#
# For $Q>0$ and $m_{4} > m_{1}$, $E_{3} = (a + \sqrt{a^2+b^2})^2$ is the only real, positive, meaningful solution.
#
# The kinetic energy of $E_3$ is, at minimum, the energy arrived at when $p_1 = 0$. Thus:
#
# \begin{align}
# E_3 \longrightarrow& \frac{m_4}{m_3 + m_4}Q\\
# &\mbox{ when } Q>0, p_1=0
# \end{align}
#
# So, no exoergic reactions are restricted by kinetics, as $Q = E_3 + E_4$, for the minimum linear momentum case, which is real and positive.
#
# ## For Endoergic Reactions ($Q<0$)
# Some $Q<0$ reactions aren't possible without a certain minimum quantity of kinetic energy.
#
#
# For $Q<0$ and $m_{4} > m_{1}$, some values of $E_{1}$ are too small to carry forward a real, positive solution. That is, the incident projectile must supply a minimum amount of kinetic energy before the reaction can occur. Without this energy, the solution for $E_3$ results in physically meaningless values. This minimum energy can be found from eqn 6.11 in your book and is :
#
# \begin{align}
# E_1^{th,k} = -\frac{m_3 + m_4}{m_3 + m_4 - m_1}Q.
# \end{align}
#
# One can often simplify this (assuming $m_i >> Q/c^2$ and $m_3 + m_4 - m_1 \simeq m_2$) :
#
#
# \begin{align}
# E_1^{th,k} \simeq - \left( 1 + \frac{m_1}{m_2} \right)Q.
# \end{align}
# +
def kinematic_threshold(m_1, m_3, m_4, Q):
"""Returns the kinematic threshold energy [MeV]
Parameters
----------
m_1: double
mass of incident projectile
m_3: double
mass of first product
m_3: double
mass of second product
Q : double
Q-value for the reaction [MeV]
"""
num = -(m_3 + m_4)*Q
denom = m_3 + m_4 - m_1
return num/denom
def kinematic_threshold_simple(m_1, m_2, Q):
"""Returns the coulombic threshold energy [MeV]
Parameters
----------
m_1: double
mass of incident projectile
m_2: double
mass of target
Q : double
Q-value for the reaction [MeV]
"""
to_return = -(1 + m_1/m_2)*Q
return to_return
# -
# # Coulombic Threshold
#
# Coulomb forces repel a projectile if it is:
#
# - a positively charged nucleus
# - a proton
#
# The force between the projectile (particle 1) and the target nucleus (particle 2) is :
#
# \begin{align}
# &F_C = \frac{Z_1Z_2e^2}{4\pi\epsilon_0r^2}\\
# \mbox{where}&&\\
# &\epsilon_0 = \mbox{the permittivity of free space.}
# \end{align}
#
# ### Think pair share:
# What are the other terms in the above equation:
#
# - $Z_1$ ?
# - $Z_2$ ?
# - $e$ ?
# - $r$ ?
#
# By evaluating the work function for approach to the nucleus with a coulomb barrier, we can establish that the coulombic threshold energy (in MeV) is :
#
# \begin{align}
# E_1^{th,C} \simeq 1.20 \frac{Z_1Z_2}{A_1^{1/3}+A_2^{1/3}}
# \end{align}
def colombic_threshold(z_1, z_2, a_1, a_2):
"""Returns the coulombic threshold energy [MeV]
Parameters
----------
z_1: int
proton number of incident projectile
z_2: int
proton number of target
a_1 : int or double
mass number of the incident projectile [amu]
a_2 : int or double
mass number of the target [amu]
"""
num = 1.20*z_1*z_2
denom = pow(a_1, 1/3) + pow(a_2, 1/3)
return num/denom
# ### Think Pair Share
#
# Which thresholds apply to the below situations:
#
# - A chargeless incident particle, reaction $Q>0$
# - A chargeless incident particle, reaction $Q<0$
# - A positively charged incident particle, reaction $Q>0$
# - A positively charged incident particle, reaction $Q<0$
# ## Overall threshold
#
# For the case where both thresholds apply, the minimum energy for the reaction to occur is the highest of the two thresholds.
#
# \begin{align}
# \min{\left(E_1^{th}\right)} = \max{\left(E^{th,C}_1,E_1^{th,k}\right)}.
# \end{align}
# ## Example
#
# Take the (p, n) reaction from $^{9}Be\longrightarrow^{9}B$. We will need to calculate:
#
# - The Q value
# - The kinematic threshold (if it applies)
# - The coulombic threshold (if it applies)
# - Determine which one is higher
# +
# Q value
# Look up the masses:
be_9_mass = 9.0121821
b_9_mass = 9.0133288
n_mass = 1.0086649158849
p_mass = 1.007825032 # hydrogen nucleus!
m_react = [be_9_mass, p_mass]
m_prods = [b_9_mass, n_mass]
q_example = q(m_react, m_prods)
print("Q: ", q_example)
# +
# Kinematic Threshold
# Which particles were which again?
m_1 = p_mass
m_2 = be_9_mass
m_3 = n_mass
m_4 = b_9_mass
# Calculate using both regular and simpler methods
E_k_th = kinematic_threshold(m_1, m_3, m_4, q_example)
E_k_th_simple = kinematic_threshold_simple(m_1, m_2, q_example)
print("E_k_th: ", E_k_th)
print("E_k_th (simplified): ", E_k_th_simple)
# +
# Coulombic Threshold
# Need some charge info and mass numbers
z_1 = 1 # proton
z_2 = 4 # Be
a_1 = 1 # proton
a_2 = 9 # Be
E_c_th = colombic_threshold(z_1, z_2, a_1, a_2)
print("E_c_th: ", E_c_th)
# +
## Which one is higher?
print("Total threshold: ", max(E_c_th, E_k_th))
# -
# # Applications: Neutron Detection
# Neutron's don't tend to directly ionize matter as they pass through. However, they can instigate nuclear reactions which produce charged products. These products, in turn, can be detected due to the ionization they create. The scheme for a Boron Trifluoride detector is below (hosted at https://www.orau.org/ptp/collection/proportional%20counters/bf3info.htm).
#
# 
#
# The wall effect results in the following spectrum (approximately):
# 
#
# In (n,p) reactions, for example, variation in emission angle of particle 3 can be used to determine the energy of the original incident neutron.
# # Applications: Neutron Production
# Specific neutron energies can be targetted by collecting them at a certain angle away from the production collision.
#
#
# 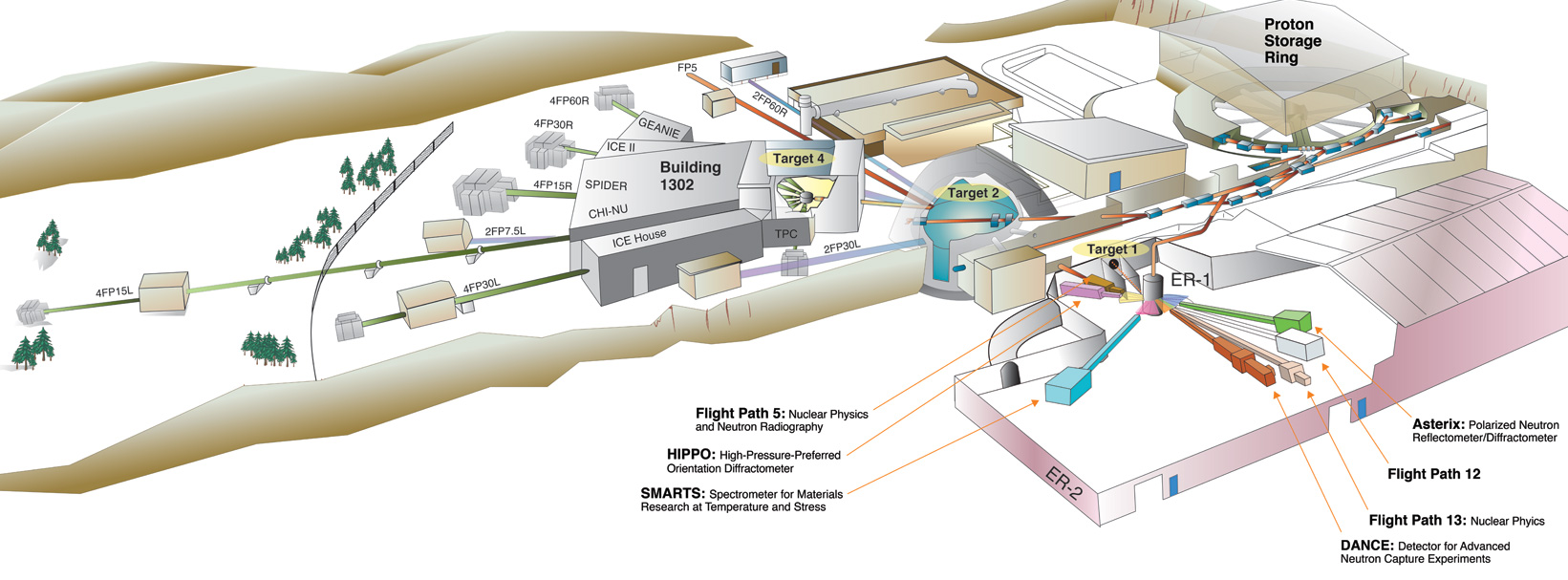
# <center>The accelerator and spallation target at LANSCE and other spallation experiments rely on this fact.</center>
#
#
# ## Two energies
#
# In (p,n) reactions, for example, certain proton energies may result in more than one neutron energy observed at a single angle. How?
#
# Recall the equation (Shultis and Faw 6.11):
#
# \begin{align}
# \sqrt{E_y}=&\sqrt{\frac{m_xm_yE_x}{(m_y + m_Y)^2}}\cos\theta_y \\
# &\pm \sqrt{\frac{m_xm_yE_x}{(m_y + m_Y)^2}\cos^2\theta_y + \left[\frac{m_Y-m_x}{(m_y + m_Y)}E_x + \frac{m_YQ}{(m_y + m_Y)}\right]}
# \end{align}
#
# Dr. Munk prefers this notation:
# \begin{align}
# \sqrt{E_3}=&\sqrt{\frac{m_1m_3E_1}{(m_3 + m_4)^2}}\cos\theta_3 \\
# &\pm \sqrt{\frac{m_1m_3E_1}{(m_3 + m_4)^2}\cos^2\theta_3 + \left[\frac{m_4-m_1}{(m_3 + m_4)}E_1 + \frac{m_4Q}{(m_3 + m_4)}\right]}
# \end{align}
# ## Heavy Particle scattering from an electron
#
# Much like the Compton reaction we saw between photons and electrons, we can see a similar reaction with heavy particles. Occaisionally, a heavy particle (e.g. a small nucleus, like an $\alpha$ particle) strikes the orbital electrons in atoms of a medium.
#
# Thus: particles 2 and 3 are the electron. So:
#
# \begin{align}
# m_2 &= m_3 = m_e = \mbox{(the electron mass)}\\
# E_3 &= E_e = \mbox{(the recoil electron energy)}\\
# m_1 &= m_4 = \mbox{(the mass of the heavy particle)}\\
# E_1 &= E_4 = \mbox{(the kinetic energy of the incident heavy particle)}
# \end{align}
#
# For this scattering process, there is no change in the rest masses of the reactants, so Q = 0.
#
# We can use the Shutlis and Faw 6.11 equation above to arrive at:
#
# \begin{align}
# \sqrt{E_e}=& \frac{2}{m_4 + m_e}\sqrt{m_4m_eE_4}\cos{\theta_e}
# \end{align}
#
# We can approximate that $m_4 >> m_e$ such that the electron recoil energy becomes:
#
# \begin{align}
# \implies E_e =& 4\frac{m_e}{m_4}E_4\cos^2{\theta_e}
# \end{align}
#
# ## Think Pair Share
# What angle, $\theta_e$, corresponds to the maximimum loss of kinetic energy by the incident heavy particle?
#
# 
#
# At $\theta_e=0$, we find that:
#
# \begin{align}
# (E_e)_{max} = 4\frac{m_e}{m_4}E_4
# \end{align}
import math
def recoil_energy(m_4, e_4, theta_e):
m_e = 0.0005486 # amu
num = 4*m_e*e_4*pow(math.cos(theta_e), 2)
return num/m_4
# +
th = [math.radians(-90),
math.radians(-75),
math.radians(-60),
math.radians(-45),
math.radians(-30),
math.radians(-15),
math.radians(0),
math.radians(15),
math.radians(30),
math.radians(45),
math.radians(60),
math.radians(75),
math.radians(90)]
m_4 = 4.003 # alpha particle
to_plot_4 = np.arange(0.,len(th))
to_plot_10 = np.arange(0.,len(th))
for k, v in enumerate(th):
to_plot_4[k] = (recoil_energy(m_4, 4, v))
to_plot_10[k] = (recoil_energy(m_4, 10, v))
plt.plot(th, to_plot_4, label="$4MeV$")
plt.plot(th, to_plot_10, label="$10MeV$")
plt.ylabel("Electron Recoil Energy ($MeV$)")
plt.xlabel("Angle (radians)")
plt.legend(loc=2)
# +
th = 0
m_4 = 4.003 # alpha particle
e_4 = 4 # MeV
print("Max (4MeV alpha): ", recoil_energy(m_4, e_4, th))
# -
# ## Neutron Scattering
#
# ### Neutron interactions with matter.
#
# \begin{align}
# ^1_0n + {^a_z}X \longrightarrow
# \begin{cases}
# ^1_0n + {^a_z}X & \mbox{Elastic Scattering}\\
# ^1_0n + \left({^a_z}X\right)^* & \mbox{Inlastic Scattering}
# \end{cases}
# \end{align}
#
# 
#
# Using the ubiquitous equation 6.11 for a neutron scatter:
#
# \begin{align}
# \sqrt{E_3}=&\sqrt{\frac{m_1m_3E_1}{(m_3 + m_4)^2}}\cos\theta_3 \\
# &\pm \sqrt{\frac{m_1m_3E_1}{(m_3 + m_4)^2}\cos^2\theta_3 + \left[\frac{m_4-m_1}{(m_3 + m_4)}E_1 + \frac{m_4Q}{(m_3 + m_4)}\right]}\\
# \end{align}
#
# We can define our particles as a neutron hitting a nucleus and changing in its energy.
#
# \begin{align}
# m_1 = m_3 = m_n\\
# E_1 = E_n\\
# E_3 = E_n'\\
# \end{align}
#
# Such that:
#
# \begin{align}
# \sqrt{E_n'} =&\sqrt{\frac{m_nm_nE_n}{(m_n + m_4)^2}}\cos\theta_s \\
# &\pm \sqrt{\frac{m_nm_nE_n}{(m_n + m_4)^2}\cos^2\theta_s + \left[\frac{m_4-m_n}{(m_n + m_4)}E_n + \frac{m_4Q}{(m_n + m_4)}\right]}
# \end{align}
#
# We can also agree that $m_2=m_4$, which is some nucleus with a mass that is approximately the same at the beginning and end of the scatter (approximate if the scattering is inelastic) . This gives, with some rearrangement:
#
# \begin{align}
# \sqrt{E_n'} &= \frac{1}{m_4 + m_n}\times\\ &\left[\sqrt{m_n^2E_n}\cos{\theta_s} \pm \sqrt{E(m_4^2 + m_n^2\cos^2{\theta_s} − m_n^2) + m_4 ( m_4 + m_n ) Q }\right]
# \end{align}
#
# And, for elastic scattering ($Q=0$):
#
# \begin{align}
# E' = \frac{1}{(A+1)^2}\left[\sqrt{E}\cos{\theta_s} + \sqrt{E(A^2 - 1 + \cos{\theta_s}^2)}\right]^2
# \end{align}
#
def scattered_neutron_energy(A, E, th):
"""Returns the energy of a scattered neutron [MeV]
Parameters
----------
A: int or double
mass number of medium
E: double
kinetic energy of the incident neutron [MeV]
th : double
scattering angle, in degrees
"""
cos_th = math.cos(math.radians(th))
term1 = 1/((A+1)**2)
term2 = math.sqrt(E)*cos_th
term3 = math.sqrt(E*(A**2 - 1 + cos_th**2))
return term1*((term2 + term3)**2)
# +
th = [math.radians(-90),
math.radians(-75),
math.radians(-60),
math.radians(-45),
math.radians(-30),
math.radians(-15),
math.radians(0),
math.radians(15),
math.radians(30),
math.radians(45),
math.radians(60),
math.radians(75),
math.radians(90)]
e_initial = 2.0 # 2 MeV is special
a_light = 4.003 # alpha particle
a_heavy = 235.0 # uranium atom
to_plot_light = np.arange(0.,len(th))
to_plot_heavy = np.arange(0.,len(th))
for k, v in enumerate(th):
to_plot_light[k] = (scattered_neutron_energy(a_light, e_initial, v))
to_plot_heavy[k] = (scattered_neutron_energy(a_heavy, e_initial, v))
plt.plot(th, to_plot_light, label="light")
plt.plot(th, to_plot_heavy, label="heavy")
plt.ylabel("Scattered Neutron Energy ($MeV$)")
plt.xlabel("Angle (radians)")
plt.legend(loc=2)
# -
# ## Average Energy Loss
#
# For elastic scattering (Q = 0), we see the minimum and maxium energies occur at the maximum and minimum angles.
#
# \begin{align}
# E'_{max} &= E'(\theta_{s,min})\\
# &= E'(\theta_{s}=0)\\
# &= E\\
# E'_{min} &= E'(\theta_{s,max})\\
# &= E'(\theta_{s}=\pi)\\
# &= \frac{(A-1)^2}{(A+1)^2} E\\
# &\equiv \alpha E\\
# \end{align}
#
# For isotropic scattering, we can find the average loss:
#
#
# \begin{align}
# (\Delta E)_{av} &\equiv E - E'_{av}\\
# &= E− 1(E+\alpha E)\\
# & = 1(1- \alpha)E
# \end{align}
# +
def alpha(a):
"""Returns the average energy loss of a
scattered neutron [MeV]
Parameters
----------
A: int or double
mass number of medium
"""
num = (a-1)**2
denom = (a+1)**2
return num/denom
def average_energy_loss(A, E):
"""Returns the average energy loss of a scattered neutron [MeV]
Parameters
----------
A: int or double
mass number of medium
E: double
kinetic energy of the incident neutron [MeV]
"""
return 1*(1-alpha(A))*E
# +
e_initial = np.arange(0, 2, 0.001)
to_plot_light = np.arange(0.,len(e_initial))
to_plot_heavy = np.arange(0.,len(e_initial))
for k, v in enumerate(e_initial):
to_plot_light[k] = (average_energy_loss(a_light, v))
to_plot_heavy[k] = (average_energy_loss(a_heavy, v))
plt.plot(e_initial, to_plot_light, label="light atom")
plt.plot(e_initial, to_plot_heavy, label="heavy atom")
plt.ylabel("Average Neutron Energy Loss ($MeV$)")
plt.xlabel("Initial neutron energy ($MeV$)")
plt.legend(loc=2)
# -
# ## Logarithmic Energy Loss
#
# It turns out, on a logarithmic energy scale, a neutron loses the same amount of logarithmic energy per elastic scatter, regardless of its initial energy. So, this is a helpful term, particularly since neutron energies can range by many orders of magnitude. So, we often use 'logarithmic energy loss' when discussing this downscattering. This is also called "lethargy".
#
# \begin{align}
# \left(\ln{(E)} - \ln{(E')}\right)_{av} & = \overline{\ln{\left(\frac{E}{E'}\right)}} \\
# &= 1 + \frac{\alpha}{1-\alpha}\\
# &= \xi\\
# &= \mbox{average logarithmic energy loss per elastic scatter}\\
# &= \mbox{lethargy}
# \end{align}
def lethargy(a):
"""Returns the average logarithmic energy
loss per elastic scatter
Parameters
----------
A: int or double
mass number of medium
"""
return 1.0 + alpha(a)/(1-alpha(a))
a = np.arange(1, 240)
plt.plot([lethargy(i) for i in a])
plt.ylabel("$\\xi$")
plt.xlabel("A($amu$)")
# ## Thermal Neutrons
#
# 1. a fast neutron slows down
# 2. may eventually come into thermal equilibrium with the medium
# 3. thermal motion of atoms in medium are in Maxwellian distribution
# 4. neutron may gain kinetic energy upon scattering from a rapidly moving nucleus
# 5. neutron may lose energy upon scattering from a slowly moving nucleus.
#
# <a title="By The original uploader was Pdbailey at English Wikipedia.
# Later versions were uploaded by Cryptic C62 at en.wikipedia.
# Convert into SVG by Lilyu from Image:MaxwellBoltzmann.gif. [Public domain], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:MaxwellBoltzmann-en.svg"><img width="512" alt="MaxwellBoltzmann-en" src="https://upload.wikimedia.org/wikipedia/commons/thumb/0/01/MaxwellBoltzmann-en.svg/512px-MaxwellBoltzmann-en.svg.png"></a>
#
#
# At room temperature, 293 K:
# - the most probable kinetic energy of thermal neutrons is 0.025 eV
# - 0.025 eV corresponds to a neutron speed of about 2200 m/s.
#
# ## Epithermal
#
# Neutrons that are faster than thermal neutrons, but aren't quite "fast" are called _epithermal_. ($0.2eV < E_{epi} < 1 MeV$)
#
# ## Fast
#
# $> 1MeV$
#
# ## Neutron Capture
#
# - Free neutrons will eventually be absorbed by a nucleus (or escape the domain of interest)
# - Neutron capture leaves the nucleus excited
# - Actually, very excited (Recall: what is a typical binding energy per nucleon?)
# - When it's released as a $\gamma$ that energy can be very hazardous
#
# Neutron slowing down can help us to reduce very high energy $\gamma$ emissions.
# ## Fission Reactions
#
# Some nuclei spontaneously fission (e.g. $^{252}Cf$). However, this isn't common.
#
#
#
# \begin{align}
# ^1_0n + ^{235}_{92}U \longrightarrow \left( ^{236}_{92}U \right)^*
# \begin{cases}
# ^{235}_{92}U + ^1_0n & \mbox{Elastic Scattering}\\
# ^{235}_{92}U + ^1_0n' + \gamma & \mbox{Inelastic Scattering}\\
# ^{236}_{92}U + \gamma & \mbox{Radiative Capture}\\
# ^{A_H}_{Z_H}X_H + ^{A_L}_{Z_L}X_L + ^1_0n + \cdots & \mbox{Fission}
# \end{cases}
# \end{align}
# # Announcements
#
# * I am modifying the schedule a bit. On Friday I will introduce different reactor types and then on Monday we are going to have a guest lecture on the nuclear fuel cycle by **<NAME>**.
# * The homework I assign on Friday will be due the Monday after spring break
# * If you wanted an adjustment on your midterm exam, you **need to email me** with the details so I can record it properly.
# * Exam 2 moved to 4/4. Syllabus updated on github.
# ### Recall: Cross sections
#
# The likelihood of each of these scattering events is captured by cross sections.
#
# - $\sigma_x = $ microscopic cross section $[cm^2]$
# - $\Sigma_x = $ macroscopic cross section $[1/length]$
# - $\Sigma_x = N\sigma_x $
# - $N = $ number density of target atoms $[\#/volume]$
#
#
# ### Cross sections are in units of area. Explain this to your neighbor.
# ### What energy neutron do we prefer for fission in $^{235}U$?
# 
# Nuclei that undergo neutron induced fission can be categorized into three types:
#
# - fissile: can fission with a slow neutron ($^{235}U$, $^{233}U$, $^{239}Pu$)
# - fissionable: require high energy (>1MeV) neutron ($^{238}U$, $^{240}Pu$)
# - fertile: can be converted into fissile or fissionable nuclide (breeding reactions)
#
# Key breeding reactions are :
#
# \begin{align}
# {^{232}_{90}}Th + ^1_0n \longrightarrow {^{233}_{90}}Th \overset{\beta^-}{\longrightarrow} {^{233}_{91}}Pa \overset{\beta^-}{\longrightarrow} {^{233}_{92}}U\\
# {^{238}_{92}}U + ^1_0n \longrightarrow {^{239}_{92}}U \overset{\beta^-}{\longrightarrow} {^{239}_{93}}Np \overset{\beta^-}{\longrightarrow} {^{239}_{94}}Pu\\
# \end{align}
# ## The fission process
#
# \begin{align}
# ^1_0n + ^{235}_{92}U \longrightarrow \left( ^{236}_{92}U \right)^* \longrightarrow X_H + X_L + \nu_p\left(^1_0n\right) + \gamma_p
# \end{align}
#
# Conserving neutrons and protons:
#
# \begin{align}
# A_L + A_H + \nu_p &= 236\\
# N_L + N_H + \nu_p &= 144\\
# Z_L + Z_H &= 92\\
# \end{align}
#
# <a title="JWB at en.wikipedia [CC BY 3.0
# (https://creativecommons.org/licenses/by/3.0
# ) or GFDL (http://www.gnu.org/copyleft/fdl.html)], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:ThermalFissionYield.svg"><img width="512" alt="ThermalFissionYield" src="https://upload.wikimedia.org/wikipedia/commons/thumb/6/68/ThermalFissionYield.svg/512px-ThermalFissionYield.svg.png"></a>
# ## Fission Product Decay
#
# The fission fragments end up very neutron rich.
#
# ### Think Pair Share
# Recall the chart of the nuclides. How will these fission products likely decay?
# ### Fission Spectrum
#
# $\chi(E)$ is an empirical probability density function describing the energies of prompt fission neutrons.
#
# \begin{align}
# \chi (E) &= 0.453e^{-1.036E}\sinh\left(\sqrt{2.29E}\right)\\
# \end{align}
# +
import numpy as np
import math
def chi(energy):
return 0.453*np.exp(-1.036*energy)*np.sinh(np.sqrt(2.29*energy))
energies = np.arange(0.0,10.0, 0.1)
plt.plot(energies, chi(energies))
plt.title(r'Prompt Neutron Energy Distribution $\chi(E)$')
plt.xlabel("Prompt Neutron Energy [MeV]")
plt.ylabel("probability")
# -
# #### Questions about this plot:
#
# - What is the most likely prompt neutron energy?
# - Can you write an equation for the average neutron energy?
# - Can you write an equation for the average neutron energy?
#
print(max([chi(e) for e in energies]), chi(0.7))
# #### Expectation Value
#
# Recall that the average energy will be the expectation value of the probability density function.
#
#
# \begin{align}
# <E> &= \int E\chi(E)dE\\
# &= E \chi(E)
# \end{align}
plt.plot(energies, [chi(e)*e for e in energies])
# ## Prompt and Delayed neutrons
#
# - Most of the neutrons in fission are emitted within $10^{-14}s$.
# - **prompt** neutrons
# - $\nu_p$
# - Some, ($<1\%$) are produced by delayed decay of fission products.
# - **delayed** neutrons
# - $\nu_d$
#
# We define the delayed neutron fraction as :
#
# \begin{align}
# \beta \equiv \frac{\nu_d}{\nu_d + \nu_p}
# \end{align}
# ## Energy from fission
# ### Reaction Rates
#
# - The microscopic cross section is just the likelihood of the event per unit area.
# - The macroscopic cross section is just the likelihood of the event per unit area of a certain density of target isotopes.
# - The reaction rate is the macroscopic cross section times the flux of incident neutrons.
#
# \begin{align}
# R_{i,j}(\vec{r}) &= N_j(\vec{r})\int dE \phi(\vec{r},E)\sigma_{i,j}(E)\\
# R_{i,j}(\vec{r}) &= \mbox{reactions of type i involving isotope j } [reactions/cm^3s]\\
# N_j(\vec{r}) &= \mbox{number of nuclei participating in the reactions } [\#/cm^3]\\
# E &= \mbox{energy} [MeV]\\
# \phi(\vec{r},E)&= \mbox{flux of neutrons with energy E at position i } [\#/cm^2s]\\
# \sigma_{i,j}(E)&= \mbox{cross section } [cm^2]\\
# \end{align}
#
#
# This can be written more simply as $R_x = \Sigma_x I N$, where I is intensity of the neutron flux.
#
# ### Source term
#
# The source of neutrons in a reactor are the neutrons from fission.
#
# \begin{align}
# s &=\nu \Sigma_f \phi
# \end{align}
#
# where
#
# \begin{align}
# s &= \mbox{neutrons available for next generation of fissions}\\
# \nu &= \mbox{the number born per fission}\\
# \Sigma_f &= \mbox{the number of fissions in the material}\\
# \phi &= \mbox{initial neutron flux}
# \end{align}
#
# This can also be written as:
#
# \begin{align}
# s &= \nu\Sigma_f\phi\\
# &= \nu\frac{\Sigma_f}{\Sigma_{a,fuel}}\frac{\Sigma_{a,fuel}}{\Sigma_a}{\Sigma_a} \phi\\
# &= \eta f {\Sigma_a} \phi\\
# \eta &= \frac{\nu\Sigma_f}{\Sigma_{a,fuel}} \\
# &= \mbox{number of neutrons produced per neutron absorbed by the fuel, "neutron reproduction factor"}\\
# f &= \frac{\Sigma_{a,fuel}}{\Sigma_a} \\
# &= \mbox{number of neutrons absorbed in the fuel per neutron absorbed anywhere, "fuel utilization factor"}\\
# \end{align}
#
# This absorption and flux term at the end seeks to capture the fact that some of the neutrons escape. However, if we assume an infinite reactor, we know that all the neutrons are eventually absorbed in either the fuel or the coolant, so we can normalize by $\Sigma_a\phi$ and therefore:
#
#
# \begin{align}
# k_\infty &= \frac{\eta f \Sigma_a\phi}{\Sigma_a \phi}\\
# &= \eta f
# \end{align}
| 10.06.1-binary_reactions/binary-reactions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Anaconda(Py3.6)
# language: python
# name: anaconda3
# ---
# # Find RA and Dec for the members in Reino+2018 Hyades study
#
# **Delete Gaia login credentials before publishing!**
import pandas as pd
import numpy as np
df = pd.read_csv('downloads/clusters_Reino/Hyades_Reino2018.txt', delimiter="\s+", skiprows=36, dtype={"source_id":str}).dropna(how='all')
df = df.rename(index=str, columns={'# HIP' : 'HIP',})
df = df.replace(' ""',-999)
df = df.replace(' ""',-999)
df.head()
df.columns = df.columns.str.replace(' ', '')
#df.source_id = df.source_id.astype(float).apply(np.rint).astype(int)
df = df.replace(-999, np.nan)
df.tail(10)
df.to_csv('downloads/clusters_Reino/hyades_reino.csv',index=False)
# # Find RA and Dec to each entry
pf = pd.read_csv('downloads/clusters_Reino/hyades_reino.csv',dtype={"source_id":str})
hip = pf.HIP.dropna().astype(int)
hip = hip.apply(lambda x: 'HIP {}'.format(str(x)))
hip.to_csv('downloads/clusters_Reino/hyades_reino_HIP.csv',index=False, header=False)
tyc = pf.TYC.dropna().str.strip()
tyc = tyc.apply(lambda x: 'TYC {}'.format(str(x)))
tyc.to_csv('downloads/clusters_Reino/hyades_reino_TYC.csv',index=False, header=False)
gaia = pf.source_id.dropna().astype(str).to_frame()
gaia.to_csv('downloads/clusters_Reino/hyades_reino_gaia.csv',index=False, header=False)
# ## HIP
hipsimbad = pd.read_table('downloads/clusters_Reino/simbad_HIP.tsv', delimiter='\t',skiprows=[0,1,2,3,4,6])
# +
hipsimbad.columns = hipsimbad.columns.str.replace(' ', '')
hipsimbad['RAJ2000'] = hipsimbad['coord1(ICRS,J2000/2000)'].apply(lambda x: str(x).split('+')[0].strip())
hipsimbad['DEJ2000'] = hipsimbad['coord1(ICRS,J2000/2000)'].apply(lambda x: str(x).split('+')[-1].strip())
hipsimbad.typedident = hipsimbad.typedident.apply(lambda x: str(x)[4:])
hipsimbad.columns.values
# -
hipsimbad.DEJ2000
excols = ['typedident','RAJ2000','DEJ2000']
hipmerge = hipsimbad[excols]
hipmerge = hipmerge.rename(index=str, columns=dict(zip(excols,['HIP','RAJ2000_HIP','DEJ2000_HIP'])))
hipmerge.DEJ2000_HIP = hipmerge.DEJ2000_HIP.apply(lambda a: a.replace(u'\N{MINUS SIGN}', '-').replace(u'\N{PLUS SIGN}', '+')).replace('','-999').astype(float)
hipmerge.RAJ2000_HIP = hipmerge.RAJ2000_HIP.apply(lambda a: a.replace(u'\N{MINUS SIGN}', '-').replace(u'\N{PLUS SIGN}', '+')).replace('','-999').astype(float)
hipmerge.HIP = hipmerge.HIP.replace("", "-999").astype(float)
hipmerge = hipmerge.replace(-999, np.nan)
hipmerge = hipmerge.dropna(how="all")
hipmerge.tail()
# ## TYC
tycsimbad = pd.read_table('downloads/clusters_Reino/simbad_TYC.tsv', delimiter='\t',skiprows=[0,1,2,3,4,6])
# +
tycsimbad.columns = tycsimbad.columns.str.replace(' ', '')
tycsimbad['RAJ2000'] = tycsimbad['coord1(ICRS,J2000/2000)'].apply(lambda x: str(x).split(' ')[0].strip())
tycsimbad['DEJ2000'] = tycsimbad['coord1(ICRS,J2000/2000)'].apply(lambda x: str(x).split(' ')[-1].strip())
tycsimbad.typedident = tycsimbad.typedident.apply(lambda x: str(x)[4:])
tycsimbad.columns.values
# -
excols = ['typedident','RAJ2000','DEJ2000']
tycmerge = tycsimbad[excols]
tycmerge = tycmerge.rename(index=str, columns=dict(zip(excols,['TYC','RAJ2000_TYC','DEJ2000_TYC'])))
tycmerge.DEJ2000_TYC = tycmerge.DEJ2000_TYC.apply(lambda a: a.replace(u'\N{MINUS SIGN}', '-').replace(u'\N{PLUS SIGN}', '+')).replace('','-999').astype(float)
tycmerge.RAJ2000_TYC = tycmerge.RAJ2000_TYC.apply(lambda a: a.replace(u'\N{MINUS SIGN}', '-').replace(u'\N{PLUS SIGN}', '+')).replace('','-999').astype(float)
tycmerge = tycmerge.replace(-999, np.nan)
tycmerge.head()
# ## Gaia
#
from astroquery.gaia import Gaia
Gaia.login(user='eilin', password='<PASSWORD>!')
#from astropy.table import Table
#filename="downloads/clusters_Reino/hyades_reino_gaia.vot"
#Table.from_pandas(gaia).write(filename, format='votable')
upload_resource = 'downloads/clusters_Reino/hyades_reino_gaia.vot'
job = "SELECT gaiadr1.gaia_source.source_id, gaiadr1.gaia_source.dec, gaiadr1.gaia_source.ra \
FROM gaiadr1.gaia_source \
JOIN user_eilin.table1 \
ON gaiadr1.gaia_source.source_id = user_eilin.table1.col1"
j = Gaia.launch_job(query=job, verbose=True, dump_to_file=True,
upload_resource=upload_resource, upload_table_name="table_test")
r = j.get_results()
r.pprint()
# +
gaiares = r.to_pandas().astype(str)
dd = gaiares.merge(gaia, on="source_id", how="outer")
# -
set(dd.source_id.dropna().tolist()) - set(gaiares.source_id.tolist())
# This is the first one in the table and it is lost in the process somehow... fix that:
gaiares = gaiares.append({"source_id":'67351752990540544', "ra" : 58.171521868568654, "dec" : 25.804222936867426 },ignore_index=True)
gaiares = gaiares.rename(index=str, columns=dict(zip(["source_id","ra","dec"],['source_id','RAJ2000_GaiaDR1','DEJ2000_GaiaDR1'])))
gaiares.to_csv("downloads/clusters_Reino/gaia_hyades_reino.csv", index=False)
gaiares.head()
# ### Now merge all tables together
hipmerge.HIP.head()
df = df.merge(hipmerge, on="HIP", how="left")
df = df.merge(tycmerge, on="TYC", how="left")
df = df.merge(gaiares, on="source_id", how="left")
df.head()
df.head(20)
# +
df = df.drop_duplicates()
df["RAJ2000"] = df[~df.DEJ2000_GaiaDR1.isnull()].RAJ2000_GaiaDR1
df["DEJ2000"] = df[~df.DEJ2000_GaiaDR1.isnull()].DEJ2000_GaiaDR1
df.loc[df.DEJ2000_GaiaDR1.isnull(),"RAJ2000"] = df.loc[df.DEJ2000_GaiaDR1.isnull(),["RAJ2000_TYC","RAJ2000_HIP"]].mean(axis=1)
df.loc[df.DEJ2000_GaiaDR1.isnull(),"DEJ2000"] = df.loc[df.DEJ2000_GaiaDR1.isnull(),["DEJ2000_TYC","DEJ2000_HIP"]].mean(axis=1)
df["ID"] = df[~df.DEJ2000_GaiaDR1.isnull()].source_id
df.loc[df.DEJ2000_GaiaDR1.isnull(),"ID"] = df.loc[df.DEJ2000_GaiaDR1.isnull(),"TYC"]#.apply(lambda x: "TYC {}".format(x))
df.loc[df.DEJ2000_GaiaDR1.isnull(),"ID"] = df.loc[df.DEJ2000_GaiaDR1.isnull(),"HIP"]#.apply(lambda x: "HIP {}".format(int(x)))
df.tail(20)
# -
# save the Gaia IDs to text
df.source_id.dropna().to_csv("downloads/clusters_Reino/gaiaidshyades_reino.txt", index=False, header=True)
# ### You need to go to the Gaia archive and propagate epoch 2015.5 coordinates to epoch 2000
#
# Here is the ADQL query (modify `user_eilin`, and other details as needed):
#
# SELECT source_id,
# COORD1(EPOCH_PROP_POS(ra, dec, parallax, pmra, pmdec, radial_velocity, 2015.5, 2000)),
# COORD2(EPOCH_PROP_POS(ra, dec, parallax, pmra, pmdec, radial_velocity, 2015.5, 2000))
#
# FROM gaiadr2.gaia_source as g
#
# INNER JOIN user_eilin.gaiachyades2 AS ei
#
# ON (g.source_id = ei.gch)
#
# ### After submitting the query, download results
#
# ...as csv file to `"downloads/clusters_Reino/hyades_reino_coords_propagated_to_epoch2000.csv"`
#
#
# ### Finally
#
# Merge new coordinates with Reino's table.
#
# Keep an eye on the `downloads/cluster_catalog_characteristics.csv` entry, update data format if needed (should be up to date).
corrected = pd.read_csv("downloads/clusters_Reino/hyades_reino_coords_propagated_to_epoch2000.csv")
corrected = corrected.rename(index=str, columns={"coord1":"RAJ2000_epoch2000",
"coord2":"DEJ2000_epoch2000"})
corrected.source_id = corrected.source_id.astype(str)
corrected.head()
newdf = df.merge(corrected, how="left", on="source_id")
newdf.loc[~newdf.RAJ2000_epoch2000.isnull(),"RAJ2000"] = newdf.RAJ2000_epoch2000.dropna()
newdf.loc[~newdf.DEJ2000_epoch2000.isnull(),"DEJ2000"] = newdf.DEJ2000_epoch2000.dropna()
newdf.to_csv("downloads/clusters_Reino/hyades_reino18.csv", index=False)
| Membership_Matching/02_PREP_Reino_Hyades_table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": true, "row": 72, "width": 4}, "report_default": {}}}}
# # Table of Contents
#
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# %matplotlib notebook
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
#plt.style.use('ggplot')
import ipywidgets as widgets
import sys, os, io, string, shutil, math
from hublib.ui import Submit
from hublib.ui import RunCommand
import hublib.use
# %use boost-1.62.0-mpich2-1.3-gnu-4.7.2
# %use lammps-31Mar17
# #%set_env OMP_NUM_THREADS=4
import numpy as np
import re
import time
from threading import Thread
from io import StringIO
from ipywidgets import Layout, Box, Label, Output
from IPython.display import display,HTML
#sys.path.append('../python/')
#import train as nn
import random
#from IPython.core.display import display, HTML
display(HTML("<style>.container { width:1450px !important; }</style>"))
style = {'description_width': 'initial'}
left_column='340px'
right_column='940px'
app_width='1280px'
form_item_layout = Layout(
display='flex',
flex_flow='row',
justify_content='space-between',
padding='5px'
)
form_item_layout_slider_text = Layout(
display='flex',
flex_flow='row',
justify_content='space-between',
padding='5px'
)
box_layout_image=Layout(
display='flex',
flex_flow='column',
border='solid 1px',
align_items='stretch',
#width='25%',
padding='5px',
#min_height='20px',
#min_width='200px'
width=left_column,
height='304.5px'
)
box_layout_phisical=Layout(
display='flex',
flex_flow='column',
border='solid 1px',
align_items='stretch',
#width='25%',
padding='5px',
#min_height='20px',
#min_width='200px'
width=left_column,
#height='180px'
)
box_layout_computing=Layout(
display='flex',
flex_flow='column',
border='solid 1px',
align_items='stretch',
padding='5px',
width=left_column,
#height='103px'
)
box_layout_progress=Layout(
display='flex',
flex_flow='column',
border='solid 1px',
align_items='stretch',
padding='5px',
width=left_column,
##height='55px'
)
box_layout_output=Layout(
display='flex',
flex_flow='column',
border='solid 1px',
align_items='stretch',
padding='5px',
width=left_column,
#height='150px'
)
box_layout_about=Layout(
display='flex',
flex_flow='column',
border='solid 1px',
align_items='stretch',
padding='5px',
width=right_column,
#height='136px'
)
form_item_layout_tab = Layout(
display='flex',
flex_flow='column',
align_items='stretch',
justify_content='space-between',
padding='5px'
)
box_layout_tabs=Layout(
display='flex',
flex_flow='column',
border='solid 1px',
align_items='stretch',
padding='5px',
width=right_column,
height='100%'
)
button_hide_layout=Layout(
visibility='hidden', #visible/hidden
padding='5px',
)
button_show_layout=Layout(
visibility='visible', #visible/hidden
padding='5px',
)
# +
'''
CSS = """
.output {
align-items: center;
}
div.output_area {
width: 100%;
}
div.output_area {
left-padding: 60px;
}
"""
display(HTML('<style>{}</style>'.format(CSS)))
'''
js_custom="""<script>
$('#appmode-leave').hide(); // Hides the edit app button.
$('#appmode-busy').hide(); // Hides the kernel busy indicator.
</script>
"""
display(HTML(js_custom))
# -
# %%capture
# This hides any output from this cell
# %set_env OMP_NUM_THREADS=4
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": true, "row": 66, "width": 4}, "report_default": {}}}} language="html"
# <style>
# .jupyter-widgets.widget-tab > .p-TabBar .p-TabBar-tab {
# flex: 0 1 175px
# }
# </style>
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 2, "hidden": true, "row": 29, "width": 4}, "report_default": {"hidden": false}}}}
npCharge = widgets.BoundedIntText(
value=-1000,
min=-1500,
max=-500,
step=1,
description='NP Charge (Q in e)',
style=style,
)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 2, "hidden": true, "row": 29, "width": 4}, "report_default": {"hidden": true}}}}
nLigand = widgets.Dropdown(
options={'25': 25, '50': 50, '75':75, '100':100},
value=50,
description="Linker Density (in units of NP density)",
style=style
)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 2, "hidden": true, "row": 29, "width": 4}, "report_default": {"hidden": true}}}}
salt = widgets.BoundedFloatText(
value=0.175,
min=0.005,
max=0.3,
step=0.005,
description='Salt Concentration (M):',
style=style
)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
imageSlider = widgets.IntSlider(
value=0,
min=0,
max=0,
step=10000,
description='Images',
style=style
)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
submit_str = ''
def onToggleBtnChange(b):
global submit_str
if clustorMode.value:
clustorMode.icon = 'check'
clustorMode.button_style='success'
if not submit_str:
output_image_warning_text.layout.visibility = 'visible'
else:
clustorMode.icon = ''
clustorMode.button_style=''
if not submit_str:
output_image_warning_text.layout.visibility = 'hidden'
'''
clustorMode =widgets.ToggleButton(
value=False,
description='Cluster mode',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Simulation will run on a computing cluster when this button is turned on but you will loose the access to dynamic simulation snapshot slider.',
icon='',
style=style
)
'''
clustorMode =widgets.Checkbox(
value=False,
description='Cluster mode',
disabled=False,
#button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Simulation will run on a computing cluster when this button is turned on but you will loose the access to dynamic simulation snapshot slider.',
icon='',
style=style
)
clustorMode.observe(onToggleBtnChange,'value')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
label_style = "style='background-color: #81c4fd; font-size:500; padding: 2px'"
label_style2 = "style='font-size:350; padding: 2px; font-weight: bold; align: center'"
def make_label(name, label_style, desc="",):
lval = '<p %s %s>%s</p>' % (desc, label_style, name)
return widgets.HTML(lval)
plabel = make_label('Input Parameters', label_style, '')
clabel = make_label('Computing Parameters',label_style, '')
olabel = make_label('Output Controllers', label_style, '')
imageSliderabout = make_label('Slide to navigate the simulation snapshots', label_style2, '')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 6, "hidden": true, "row": 43, "width": 12}, "report_default": {"hidden": false}}}}
'''
image_html = widgets.HTML(
value='<img src="../images/overview.jpg" style="width: 500px"/>',
)
'''
file = open("../images/overview.png", "rb")
image = file.read()
image=widgets.Image(
value=image,
format='jpg',
width=297,
height=297,
)
form_items = [
Box([image], layout=form_item_layout)
]
mainImage = Box(form_items, layout=box_layout_image)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 11, "hidden": true, "row": 59, "width": 4}, "report_default": {}}}}
form_items = [
Box([plabel], layout=form_item_layout),
Box([npCharge], layout=form_item_layout),
Box([nLigand], layout=form_item_layout),
Box([salt], layout=form_item_layout)
]
form_items_computing = [
Box([clabel], layout=form_item_layout),
Box([clustorMode], layout=form_item_layout)
]
form_items_output = [
Box([olabel], layout=form_item_layout),
Box([imageSlider], layout=form_item_layout),
Box([imageSliderabout], layout=form_item_layout)
]
physical_para = Box(form_items, layout=box_layout_phisical)
computing_para = Box(form_items_computing, layout=box_layout_computing)
output_para = Box(form_items_output, layout=box_layout_output)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 6, "hidden": true, "row": 43, "width": 8}, "report_default": {"hidden": false}}}}
about=widgets.Textarea(
value='This app simulates the self-assembly of charged nanoparticles (NPs) into aggregates mediated by smaller, oppositely-charged linkers under different ionic physiological conditions. Users can input control parameters such as NP charge (from -500e to -1500e), linker density (from 25X to 100X the NP density), and ionic strength (from 0.01 M to 0.3 M) to predict formation of NP aggregates. This information may be useful in designing NP features to produce desired effects when NPs interface with biological entities. Outputs are structural information such as pair correlation functions (often denoted as g(r)) and simulation snapshots (with only NPs shown for clarity). The NPs are modeled after P22 virus-like particles (VLPs) of diameter 56 nanometers, and linkers represent smaller nanoparticles (dendrimers) of 6.7 nanometer diameter. Linker charge is fixed to 35e. Simulations are performed using LAMMPS; pre- and postprocessing are done using C++ codes. Simulation results are tested and validated using SAXS and dynamic light scattering measurements of the VLP aggregates; experiments also guide the model design. After you click Run, the application pre-processes the input parameters and loads the input script in the LAMMPS engine; this may take a couple of minutes after which the output log should appear in the Output text pane. As the output shows, the images will start to populate the "Simulation Snapshot" tab; the slider can be used to navigate through the images. At the end of simulation run, which will take close to 60 minutes to run in interative mode(default) and 6 minutes to run in cluster, g(r) will be produced in the "Pair Correlation" tab. ',
placeholder='',
layout={'height': '119px','width': '100%'},
disabled=True,
style = {'font_weight': '150px'}
)
form_about = [
Box([about], layout=form_item_layout)
]
about_text = Box(form_about, layout=box_layout_about)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
def clear_All():
localOutput.value = ''
submit_str= ''
progressBar.value=0
imageSlider.max=0
imageSlider.value=0
output_image.value=imageEmptyBox
movie_download.layout = pair_core_download.layout = button_hide_layout
clustorMode.disabled = False
if clustorMode.value:
output_image_warning_text.layout.visibility = 'visible'
else:
output_image_warning_text.layout.visibility = 'hidden'
plt.figure('Pair correlation function')
plt.clf() # clear old plot
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
def runPreprocessor(npCharge_value, nLigand_value, salt_value):
preprocessingCMD = "./preprocessor -Q "+str(npCharge_value)+" -n "+str(nLigand_value)+" -c "+str(salt_value) + " > preprocessor.log"
try:
# !$preprocessingCMD
except Exception as e:
localOutput.value = localOutput.value + "Error occured during the execution of preprocessing executable\n"
sys.stderr.write('Error occured during the execution of preprocessing executable')
sys.stderr.write(str(e))
sys.exit(1)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
runFlag=False
working_dir=""
parameter_append=""
sim_bigin=False
build_Plots=True
simulation_params=""
def callExe(s):
global runFlag, working_dir, parameter_append, sim_bigin, ionsplot, build_Plots, simulation_params, nLigand_value, runName, simulationStepsToatal
global submit_str
if build_Plots:
showPlotUI()
build_Plots = False
clear_All()
#clustorMode.disabled = True
localOutput.value = localOutput.value + "Simulation begins...\n"
npCharge_value = npCharge.value
nLigand_value = nLigand.value
salt_value = salt.value
simulationStepsToatal = 3000000
total_processors = 64
walltime = 20
runName='npassemblylab'
#Adding standard outputs to string to be sent to output widget callback function
parameter_append= "NP Charge(Q in e) is "+ str(npCharge_value)+"\n"
parameter_append= parameter_append + "Number of Linkers is "+ str(nLigand_value*108)+"\n"
parameter_append= parameter_append + "Salt Concentration (M) is "+ str(salt_value)+"\n"
parameter_append= parameter_append + "requested walltime is "+ str(walltime)+" (mins)\n"
parameter_append= parameter_append + "requested total processors are "+ str(total_processors)+"\n"
simulation_params="_%d" % int(npCharge_value)+"_%d" % int(nLigand_value)+"_%.2f" % float(salt_value)
shutil.rmtree('outfiles',True)
if not os.path.exists('outfiles'):
os.makedirs('outfiles')
localOutput.value = localOutput.value + "Preprocessing is running...\n"
#preprocessing stage
runPreprocessor(npCharge_value, nLigand_value, salt_value)
localOutput.value = localOutput.value + "Preprocessing completed\n"
localOutput.value = localOutput.value + parameter_append;
runFlag=True
sim_bigin=True
#starting a thread for on the fly density plot
#t2 = Thread(target=iterative_density_plot)
#t2.start()
try:
#rname = s.make_rname(e_np_value, e_out_value, pion_value, c_out_value, np_radius_value, np_charge_value, sim_steps_value)
submit_str = ''
if clustorMode.value:
submit_str = '--venue standby@brown -n '+str(total_processors)+' -N 16'+' -w '+str(walltime)+' -e OMP_NUM_THREADS=1'+' --tailStdout 1000 ' +'--inputfile '+'infiles '+'--inputfile '+'outfiles '+'lammps-31Mar17-parallel -in in.lammps'
#submit_str = '-n '+str(total_processors)+' -w '+str(walltime)+' -N 20'+' -e OMP_NUM_THREADS=1'+' --tailStdout 1000 ' +'--inputfile '+'outfiles '+'lammps-31Mar17-parallel -in in.lammps'
#submit_str = '--venue <EMAIL> ' + '-n 32 -w 60 -N 16'+' --tailStdout 1000 ' +'--inputfile '+'outfiles '+'lammps-31Mar17-parallel -in in.lammps'
localOutput.value = localOutput.value + "LAMMPS input script designed and simulation submitted to the cluster, waiting for response...\n"
else:
submit_str = '--local stdbuf -o0 -e0 lmp_serial -in in.lammps'
localOutput.value = localOutput.value + "LAMMPS input script designed and simulation running...\n"
s.run(runName, submit_str)
time.sleep(1)
except Exception as e:
sys.stderr.write('Error occured during the execution of Lammps program')
sys.stderr.write(str(e))
sys.exit(1)
runFlag = False
#t2.join()
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
def doneFunc(s, rdir):
global submit_str
#this is a condtional variable for few things
submit_str = ''
progressBar.value=int(100)
localOutput.value = localOutput.value + "LAMMPS simulation finished\n"
submit_btn.but.disabled = True
clustorMode.disabled = False
localOutput.value = localOutput.value + "Postprocessing is running...\n"
#postprocessing stage
postprocessingCMD = "./postprocessor -n "+str(nLigand_value)+" -N 150 > postprocessor.log"
try:
# !$postprocessingCMD
except Exception as e:
localOutput.value = localOutput.value + "Error occured during the execution of postprocessing executable\n"
sys.stderr.write('Error occured during the execution of postprocessing executable')
sys.stderr.write(str(e))
sys.exit(1)
localOutput.value = localOutput.value + "Postprocessing is finished\n"
# Plotting the result
plot_data()
localOutput.value = localOutput.value + "Simulation ended\n"
submit_btn.but.disabled = False
#image slider fully ready
imageSlider.max=int(simulationStepsToatal/imageSlider.step)*imageSlider.step
imageSlider.value=int(simulationStepsToatal/imageSlider.step)*imageSlider.step
movie_download.layout = pair_core_download.layout = button_show_layout
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# after the run finished, optionally do something
# with the output
#def plot_data(s, rdir):
def plot_data():
global build_Plots
if build_Plots:
showPlotUI()
build_Plots = False
#We ignore rdir because submit puts non-parametric runs in the current directory
# When caching is enabled, they get put in a chached directory and rdit is always used.
#wait for 2 seconds before plotting errorBars.
time.sleep(2)
plt.figure('Pair correlation function')
plt.clf() # clear old plot
plt.title('NP-NP Pair correlation function')
plt.xlabel('Distance (in units of NP diameter)', fontsize=10)
plt.ylabel('g(r)', fontsize=10)
# our simulation writes to outfiles -> density_profile.dat
try:
density_profile = np.loadtxt(working_dir + 'outfiles/gr_VV_dr=0.005.out')
except:
sys.stderr.write('Can not find the pair correlation results file')
sys.exit(1)
#print(density_profile)
x = density_profile[:,0]
# droping data above 4.3 in x dim.
#x = numpy.select([ x < 4.3 ],x)
y = density_profile[:x.size,1]
#yrr = density_profile[:,2]
#plt.errorbar(x,y,xerr=0.0, yerr=yrr, c='b', ecolor='r', capsize=2)
plt.plot(x,y)
plt.show()
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
in_wrapper_progress = False
saved_data = ""
saved_data_enery = ""
saved_data_ions_pos = ""
stepNumber = 0
def stdoutput_handler(buffer):
global in_wrapper_progress, saved_data, parameter_append, sim_bigin, stepNumber, saved_data_enery, saved_data_ions_pos
output = []
#return buffer
if sim_bigin:
#output.append(parameter_append)
sim_bigin = False
data = buffer.split('\n')
last = len(data) - 1
for i, line in enumerate(data):
output.append(line)
if line.startswith('Loop time of '):
in_wrapper_progress = False
continue
if in_wrapper_progress:
number = line.split()
if number:
if number[0].isdigit():
#print(number[0])
percent = float(number[0])/simulationStepsToatal*100
progressBar.value=int(percent)
# This adjust the image slider dynamically
# dynamic update only happens for local VM run
if submit_str.startswith('--local '):
if(int(number[0])%imageSlider.step == 0):
imageSlider.max=number[0]
imageSlider.value=number[0]
continue
if line.startswith('Step Temp c_NanoparticlePE Volume'):
in_wrapper_progress = True
reduced_buffer = '\n'.join(output)
#appending new line at the end
if reduced_buffer:
if reduced_buffer[-1] != '\n':
reduced_buffer = reduced_buffer + '\n'
return reduced_buffer
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
#button_style = "style='background-color: #81c4fd; font-size:500; padding: 4px; -webkit-appearance: button; -moz-appearance: button; appearance: button; text-decoration: none; color: initial;'"
def make_downloadBtn(downFilename="", downloadURL="", displayText=""):
bval = "<a download='%s' href='%s'><button class='p-Widget jupyter-widgets jupyter-button widget-button mod-info'>%s</button></a>" % (downFilename, downloadURL, displayText)
return widgets.HTML(bval)
#from IPython.display import FileLink, FileLinks
#FileLinks('outfiles')
#FileLinks('data')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 59, "hidden": true, "row": 105, "width": 12}, "report_default": {"hidden": false}}}}
# %%capture
tab_layout=Layout(
width='100%',
height='588px',
justify_content='space-between',
)
# we want the plot to go here, so create an empty one initially
#create tabs
figsize_tup=(8.60, 5.05)
plt.ioff() #LINE ADDED, Interactive plot blur fix
figure_density_pos = plt.figure(num='Pair correlation function', figsize=figsize_tup)
out_desnity_pos = Output(layout=tab_layout)
plt.ion() #LINE ADDED
file = open("../images/image.emptybox.jpg", "rb")
imageEmptyBox = file.read()
output_image=widgets.Image(
format='jpg',
width=584,
height=584,
value=imageEmptyBox
)
#out_output_image = Output(layout=tab_layout)
output_image_layout=Layout(
display='flex',
flex_flow='row',
#border='solid 2px',
align_items='stretch',
margin = '0px 0px 0px 130px' #top, right, bottom and left
)
box_layout_warning_text=Layout(
display='flex',
flex_flow='row',
#border='solid 1px',
align_items='stretch',
padding='5px',
width='140px',
visibility = 'hidden'
#height='136px'
)
#label_style3 = "style='background-color: #eeeeee; color: #388e3c; font-size:350; padding: 2px; font-weight: bold; align: center'"
label_style3 = "style='background-color: #eeeeee; font-size:350; padding: 2px; font-weight: bold; align: center'"
output_image_warning_text = make_label('Cluster mode is on! You will lose access to the dynamic simulation snapshot data. Snapshot data will be available at the end of simulation.', label_style3, '')
form_output_image_sec = [
Box([output_image], layout=output_image_layout),
Box([output_image_warning_text], layout=box_layout_warning_text)
]
output_image_section = Box(form_output_image_sec, layout=tab_layout)
#data download tab
movie_download = make_downloadBtn(downFilename="movie.xyz", downloadURL="outfiles/dump.melt", displayText="Movie")
pair_core_download = make_downloadBtn(downFilename="Pair_correlation_data.out", downloadURL="outfiles/gr_VV_dr=0.005.out", displayText="Pair correlation data")
movie_download.layout = pair_core_download.layout = button_hide_layout
box_layout_outputs=Layout(
display='flex',
flex_flow='column',
align_items='stretch',
padding='5px'
)
form_output_content = [
Box([movie_download], layout=box_layout_outputs),
Box([pair_core_download], layout=box_layout_outputs)
]
downloadMovieSec = Box(form_output_content, layout=tab_layout)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
def onSliderChange(b):
value = imageSlider.value
valueText = str(value)
if value==0:
valueText = "000"
fileName="image."+valueText+".jpg"
fileName="outfiles/"+fileName;
try:
file = open(fileName, "rb")
selectedImage = file.read()
output_image.value=selectedImage
except:
sys.stderr.write('Image file is not there yet.')
imageSlider.observe(onSliderChange, names='value')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
#tabChange detection and plot data incase automatically not drawn
def onTabChange(b):
global build_Plots
if os.path.isfile('outfiles/gr_VV_dr=0.005.out') and not build_Plots:
plot_data()
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 27, "hidden": true, "row": 49, "width": 8}, "report_default": {}}}}
tab_contents = ['Pair Correlation', 'Simulation Snapshot', 'Downloads']
children = [out_desnity_pos, output_image_section, downloadMovieSec]
tabSpace = widgets.Tab()
tabSpace.children = children
for i in range(len(children)):
tabSpace.set_title(i, str(tab_contents[i]))
tabSpace.selected_index = 1
tabSpace.observe(onTabChange, names='selected_index')
form_about = [
Box([tabSpace], layout=form_item_layout_tab)
]
tab = Box(form_about, layout=box_layout_tabs)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {}}}}
def showPlotUI():
with out_desnity_pos:
display(figure_density_pos, layout=tab_layout)
#with out_output_image:
# display(output_image, layout=tab_layout)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": null, "height": 37, "hidden": true, "row": 105, "width": null}, "report_default": {"hidden": false}}}}
form_item_layout_output = Layout(
display='flex',
flex_flow='column',
justify_content='space-between',
padding='4px'
)
form_item_layout_run_btn = Layout(
display='flex',
flex_flow='column',
justify_content='space-between',
padding='4px'
)
#Submit(start_func=callExe, done_func=plot_data, cachename='SubmitTest1')
submit_btn=Submit(start_func=callExe, done_func=doneFunc, outcb=stdoutput_handler)
submit_btn.acc.set_title(0, "LAMMPS Output")
submit_btn.w.layout = form_item_layout_run_btn
#submit_btn.acc.layout = form_item_layout_output
localOutput = widgets.Textarea(layout={'width': '100%', 'height': '250px'})
localOutputWidget = widgets.Accordion(children=[localOutput],layout=form_item_layout_output)
localOutputWidget.set_title(0, "Output")
localOutputWidget.selected_index = None
localOutput.value = ""
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 2, "hidden": true, "row": 70, "width": 4}, "report_default": {}}}}
progressBar=widgets.IntProgress(
value=0,
min=0,
max=100,
step=1,
description='Progress:',
bar_style='', # 'success', 'info', 'warning', 'danger' or ''
orientation='horizontal',
layout={'width': '100%'},
style=style
)
form_items_progress = [
Box([progressBar], layout=form_item_layout)
]
progressBar_Ele = Box(form_items_progress, layout=box_layout_progress)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 29, "hidden": true, "row": 76, "width": 12}, "report_default": {}}}}
app_height='840px'
form_item_layout = Layout(
display='flex',
flex_flow='row',
justify_content='space-between',
padding='5px'
)
box_layout_column_left=Layout(
display='flex',
flex_flow='column',
#border='solid 2px',
align_items='stretch',
width=left_column,
height=app_height
#height='840px'
)
box_layout_column_right=Layout(
display='flex',
flex_flow='column',
#border='solid 2px',
align_items='stretch',
width=right_column,
height=app_height
#height='840px'
)
box_layout=Layout(
display='flex',
flex_flow='row',
#border='solid 2px',
align_items='stretch',
width=app_width,
#padding='5px',
height=app_height
)
box_layout_rows=Layout(
display='flex',
flex_flow='column',
#border='solid 2px',
align_items='stretch',
width=app_width,
#padding='5px',
height='1900px',
#overflow_x='scroll',
#overflow_y='scroll'
)
gui_left = [
Box([mainImage], layout=form_item_layout),
Box([physical_para], layout=form_item_layout),
Box([computing_para], layout=form_item_layout),
Box([progressBar_Ele], layout=form_item_layout),
Box([output_para], layout=form_item_layout)
]
gui_right = [
Box([about_text], layout=form_item_layout),
Box([tab], layout=form_item_layout)
]
form_left = Box(gui_left, layout=box_layout_column_left)
form_right = Box(gui_right, layout=box_layout_column_right)
gui_upper = [
Box([form_left], layout=box_layout_column_left),
Box([form_right], layout=box_layout_column_right)
]
form = Box(gui_upper, layout=box_layout)
gui = widgets.VBox([form, localOutputWidget, submit_btn.w],layout=box_layout_rows)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 78, "hidden": false, "row": 0, "width": 12}, "report_default": {}}}}
gui
| bin/npassemblylab-frontend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 7 - Time Evolution
# Exploring time evolution of quantum states. Run the usual imports, and use the spin-1/2 states as previously defined:
import matplotlib.pyplot as plt
from numpy import sqrt,pi,arange,cos,sin
from qutip import *
# %matplotlib inline
pz = Qobj([[1],[0]])
mz = Qobj([[0],[1]])
px = Qobj([[1/sqrt(2)],[1/sqrt(2)]])
mx = Qobj([[1/sqrt(2)],[-1/sqrt(2)]])
py = Qobj([[1/sqrt(2)],[1j/sqrt(2)]])
my = Qobj([[1/sqrt(2)],[-1j/sqrt(2)]])
Sx = 1/2.0*sigmax()
Sy = 1/2.0*sigmay()
Sz = 1/2.0*sigmaz()
# Define the Hamiltonian:
omega = 2*pi
H = -omega*Sz
t = arange(0,4*pi/omega,0.05)
# The next line calls a Schrödinger equation solver (sesolve). It's arguments are the Hamiltonian, the starting state $\lvert+x\rangle$ (px), the time values, and a list of operators. `sesolve` returns many things, but the `expect` method is most useful, it gives the expectation values of the three operators in the operator list.
expect_ops = [Sx,Sy,Sz]
result1 = sesolve(H, px, t, expect_ops)
expect_ops[0] # TODO get name of variable to use in label
# +
labels = ['x','y','z']
for r,l in zip(result1.expect,labels):
plt.plot(result1.times*omega/pi, r, label="$\langle S_%c \\rangle $" % l)
plt.xlabel("Time ($\Omega t/\pi$)", size=18)
plt.legend(fontsize=16) #TODO fix legend text size
# -
# Now what if the system starts in $\lvert+z\rangle$?
result2 = sesolve(H, pz, t, [Sx,Sy,Sz])
# +
for r,l in zip(result2.expect,labels):
plt.plot(result2.times*omega/pi, r, label="$\langle S_%c \\rangle $" % l)
plt.xlabel("Time ($\Omega t/\pi$)", size=18)
plt.legend(fontsize=16) #TODO fix legend text size
# -
# Spin-up stays spin-up (i.e. no prescession)
# ## Two particle systems:
# $\lvert\psi\rangle = \frac{1}{\sqrt{2}} \lvert+z,-z\rangle + \frac{1}{\sqrt{2}} \lvert-z,+z\rangle$
#
# Use the `tensor` QuTiP function to form multi-particle states
psi = 1/sqrt(2)*tensor(pz, mz) + 1/sqrt(2)*tensor(mz, pz)
# Hamiltonian is the same for both particles so we use the tensor to form $\hat{H}$ from individual operators
omega = 5
H = -omega*tensor(Sz,Sz)
expect_op = tensor(mz,pz)*tensor(mz,pz).dag()
result3 = sesolve(H, psi, t, expect_op)
# +
for r,l in zip(result3.expect,labels):
plt.plot(result3.times*omega/pi, r, label="$\langle -z,+z\\rangle$")
plt.xlabel("Time ($\Omega t/\pi$)", size=18)
plt.legend(fontsize=16) #TODO fix legend text size
# -
# The value is constant since the state is initially in an eigenstate of $\hat{H}$.
# ## What if the magnetic field is not along an axis?
# Notice the Hamiltonian has an $x$ and a $z$ component:
omega=2
H = -omega/sqrt(2)*(Sz + Sx)
t = arange(0,2*pi/omega,0.05)
result4 = sesolve(H, px, t, [Sx, Sy, Sz])
# +
for r,l in zip(result4.expect,labels):
plt.plot(result4.times*omega/pi, r, label="$\langle S_%c \\rangle $" % l)
plt.xlabel("Time ($\Omega t/\pi$)", size=18)
plt.legend(fontsize=16) #TODO fix legend text size
# -
# Harder to interpret, so we'll use the Bloch sphere:
sx, sy, sz = result4.expect
b = Bloch()
b.add_points([sx,sy,sz])
b.zlabel = ['$\\left|+z\\right>$', '$\\left|-z\\right>$']
b.view = [-45,20]
b.add_vectors([1/sqrt(2),0,1/sqrt(2)])
b.show()
# ## Time-dependent Hamiltonian:
# We'll explore the parameters of a spin in a time-varying magnetic field. This system is relevant to nuclear magnetic resonance (NMR) which is used in chemistry and as Magnetic Resonance Imaging (MRI) in medicine.
#
# Following Compliment 9.A the Hamiltonian is:
# $$\hat{H}= - \Omega_0 \hat{S}_z - \Omega_1 cos(\omega t)\hat{S}_x$$
# We then solve for a certain amount of time after the state starts in $|\psi(0)\rangle = |+z\rangle$
#
# We also use the definition of the Rabi frequency: $\Omega_R = \sqrt{(\omega - \Omega_0)^2 + (\Omega_1/2)^2}$ as in (9.A.28)
#
# Note that the time span is 3 units of $\frac{2\pi}{\Omega_R}$. Leave the scaling in place, but to plot a longer time period, change 3.0 to something larger. This lets us match the units in Fig. 9.A.1.
# +
omega0 = 2.0 * 2 * pi # pick a nice value for a frequency, note this is 1 Hz
omega1 = 0.25 * 2 * pi # 25% of omega0
w = 2.0 * 2 * pi # the driving frequency
H0 = - omega0 * Sz # the first term in H
H1 = - omega1 * Sx # the second term in H
omegaR = sqrt((w - omega0)**2 + (omega1/2.0)**2)
t = arange(0,3.0 * 2 * pi / omegaR,0.02) # scale the time by omegaR, plot 3 units of 2pi/omegaR
args = [H0, H1, w] # parts of the Hamiltonian
def H1_coeff(t, args):
return cos(w * t)
H = [H0, [H1, H1_coeff]]
# -
# The next line calls a Schrödinger equation solver (`sesolve`). The arguments are the Hamiltonian, the starting state $\lvert+z\rangle$ (`pz`), the time values, a list of operators, and the arguments to the function `H_t`. `sesolve` returns many things, but the `expect` method is most useful, it gives the expectation values of the four operators in the operator list. Notice the fourth operator is the $\lvert-z\rangle$ projection operator. It's expectation value is $P(\lvert-z\rangle,t)$
result5 = sesolve(H, pz, t, [Sx, Sy, Sz, mz*mz.dag()],args)
sx, sy, sz, Pmz = result5.expect
# Look at the Bloch sphere for this system:
b2 = Bloch()
b2.add_points([sx,sy,sz])
b2.zlabel = ['$\\left|+z\\right>$', '$\\left|-z\\right>$']
b2.show()
# Make a plot analogous to Fig 9.A.1:
plt.tick_params(labelsize=18)
plt.plot(result5.times*omegaR/pi,Pmz)
plt.xlabel("Time ($\Omega_R t/\pi$)", size=18)
plt.ylabel("$P(-z, t)$", size=18)
# ### Q) What happens in each unit of time ($\frac{2\pi}{\Omega_R}$)? Look at the plot of $P(-z,t)$ to interpret this. How is your figure different from the on in Fig. 9.A.1?
# ### Q) How does the evolution change if you double $\Omega_0$?
# ### Q) After doubling $\Omega_0$ what if you double the driving frequency ($\omega$) also? Interpret this observation in terms of Fig. 9.A.2. In practice, what experimental parameter changes $\Omega_0$?
# ### Q) How does $\Omega_1$ influence the dynamics? (Be careful reading the plots since the units are scaled by $\Omega_R$).
# ### Advanced topic: we can change the Hamiltonian so the applied field turns off at a certain time, and it is possible to get the spin to stay in a particular state. This is very useful in quantum optics where certain operations change the atomic state in a very specific way.
# +
omega0 = 1.0 * 2 * pi # pick a nice value for a frequency, note this is 1 Hz
omega1 = 0.05 * 2 * pi # 25% of omega0
w = 1.0 * 2 * pi # the driving frequency
H0 = - omega0 * Sz # the first term in H
H1 = - omega1 * Sx # the second term in H
omegaR = sqrt((w - omega0)**2 + (omega1/2.0)**2)
t = arange(0,3.0 * 2 * pi / omegaR,0.05) # scale the time by omegaR, plot 3 units of 2pi/omegaR
def H1_coeff2(t, args): # this function calculates H at each time step t
if t < 2*pi/omegaR * 0.5: # only add the H1 piece for the first chunk of time.
coeff = cos(w * t)
else:
coeff = 0
return coeff
H = [H0, [H1, H1_coeff2]]
# -
result6 = sesolve(H, pz, t, [Sx, Sy, Sz, mz*mz.dag()],args)
sx, sy, sz, Pz = result6.expect
plt.plot(result6.times,Pz)
plt.ylim(-0.1,1.1)
plt.xlim(-5,125)
plt.xlabel("Time ($\Omega_R t/\pi$)", size=18)
plt.ylabel("$P(-z, t)$", size=18)
| Lab 7 - Time Evolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import pickle
import matplotlib.pyplot as plt
import seaborn as sns
from utils import *
with open('../output/ncf_xnn_int_diff_v2_all_ut_0_test_score.p', 'rb') as f:
ncf = pickle.load(f)
ncf = ncf.loc[:,['session_id', 'step', 'item_id', 'score']]
ncf.head()
with open('../output/lgb_coocc_last_interact_action_v2_all_ut_test_score.p', 'rb') as f:
lgb = pickle.load(f)
lgb = lgb.loc[:,['session_id', 'step', 'item_id', 'score']]
print(lgb.shape)
lgb.head()
with open('../output/xgb_coo_lf200_lr002_v2_all_ut_test_score.p', 'rb') as f:
xgb = pickle.load(f)
xgb = xgb.loc[:,['session_id', 'step', 'item_id', 'score']]
xgb.head()
# +
# with open('../output/ncf_xnn_int_diff_v2_all_ut_3_test_score.p', 'rb') as f:
# zong_han = pickle.load(f)
# zong_han = zong_han.loc[:,['session_id', 'step', 'item_id', 'score']]
# zong_han.head()
# -
with open('../output/xgb_cc_img_local_img_dl_cc_v2_all_ut_test_score.p', 'rb') as f:
temp_xgb = pickle.load(f)
ncf['item_id'] = temp_xgb.item_id
ncf['step'] = temp_xgb.step
# zong_han['item_id'] = temp_xgb.item_id
# zong_han['step'] = temp_xgb.step
# ncf['item_id'] = item_encoders.reverse_transform(ncf.item_id.values)
# del lgb
ncf.head()
# +
# with open('../output/lgb_gic_lic_wosh_v2_weighted_140k_test_score.p', 'rb') as f:
# yifu = pickle.load(f)
# yifu = yifu.loc[:,['session_id', 'step', 'item_id', 'score']]
# print(yifu.shape)
# yifu.head()
# +
merged_df = ncf.rename(columns={'score':'ncf_score'})\
.merge(xgb.rename(columns={'score':'xgb_score'}), on=['session_id','step','item_id'])\
.merge(lgb.rename(columns={'score':'lgb_score'}), on=['session_id','step','item_id'])
# .merge(zong_han.rename(columns={'score':'zong_han_score'}), on=['session_id','step','item_id'])\
# .merge(yifu.rename(columns={'score':'yifu_score'}), on=['session_id','step','item_id'])\
merged_df.head()
# -
ncf.shape, lgb.shape, xgb.shape, merged_df.shape
# ncf['score'] = (ncf.score * 0.2 + lgb.score * 0.7 + xgb.score * 0.4)
# ncf.head()
merged_df['score'] = merged_df.ncf_score * 0.1 + merged_df.lgb_score * 0.7 + merged_df.xgb_score * 0.4
merged_df.head()
# +
# ncf.columns = ['row_index', 'step', 'item_id', 'score_1']
# ncf['score_2'] = lgb.score
# ncf.head()
# +
# ncf.score_2.mean()
# +
# ncf['step'] = lgb.step
# ncf['item_id'] = lgb.item_id
# # ncf['score'] = (lgb['score'] * 0.3 + ncf['score'] * 0.7)
# ncf.head()
# +
from scipy.special import softmax
predictions = []
session_ids = []
grouped_test = merged_df.groupby('session_id')
for session_id, group in grouped_test:
scores = group.score.values
# sm_score_1 = softmax(group['score_1'].values)
# sm_score_2 = softmax(group['score_2'].values)
# scores = sm_score_1 * 0.6 + sm_score_2 * 0.4
# rank_1 = compute_rank(group['score_1'].values, to_np=True)
# rank_2 = compute_rank(group['score_2'].values, to_np=True)
# scores = rank_1*0.6 + rank_2*0.4
sorted_arg = np.flip(np.argsort(scores))
sorted_item_ids = group['item_id'].values[sorted_arg]
# sorted_item_ids = cat_encoders['item_id'].reverse_transform(sorted_item_ids)
sorted_item_string = ' '.join([str(i) for i in sorted_item_ids])
predictions.append(sorted_item_string)
session_ids.append(session_id)
prediction_df = pd.DataFrame()
prediction_df['session_id'] = session_ids
prediction_df['item_recommendations'] = predictions
# +
sub_df = pd.read_csv('../input/submission_popular.csv')
sub_df.drop('item_recommendations', axis=1, inplace=True)
sub_df = sub_df.merge(prediction_df, on="session_id")
# sub_df['item_recommendations'] = predictions
sub_df.to_csv(f'../output/ncf_xnn_int_diff_lgb_coocc_last_interact_action_xgb_cc_img_local_img_dl_cc_v2_all_ut_1_7_4.csv', index=None)
# -
pd.read_csv(f'../output/ncf_xnn_int_diff_zonghan_lgb_coocc_xgb_gic_lic_wosh_lr002_05_05_9_4.csv').head()
pd.read_csv(f'../output/ncf_xnn_int_diff_lgb_new_hp_xgb_gic_lic_wosh_lr002_1_7_4.csv').head()
pd.read_csv(f'../output/ncf_xnn_int_diff_lgb_gic_lic_wosh_xgb_gic_lic_wosh_2_4_4.csv').head()
pd.read_csv(f'../output/ncf_xnn_int_diff_lgb_gic_lic_wosh_xgb_gic_lic_wosh_1_4_4.csv').head()
pd.read_csv(f'../output/ncf_xnn_int_diff_lgb_time_elapse_xgb_time_elapse_0_4_4.csv').head()
pd.read_csv(f'../output/ncf_xnn_int_diff_lgb_best_logloss_xgb_last_sc_2_4_4.csv').head()
pd.read_csv(f'../output/lgb_best_logloss_v2_all_ut.csv').head()
pd.read_csv(f'../output/lgb_best_logloss_v2_all_ut.csv').head()
pd.read_csv(f'../output/ncf_xnn_oic_oii_lgb_sum_v2_all_ut_test_score_xgb_2_7_4.csv').head()
pd.read_csv(f'../output/ncf_xnn_intpop_clickout_v2_all_ut_lgb_bf_beach_v2_all_ut_xgb_4_8_8.csv').head()
pd.read_csv(f'../output/ncf_xnn_intpop_clickout_v2_all_ut_lgb_cc_img_local_img_dl_cc_v2_all_ut_xgb_4_6_6.csv').head()
pd.read_csv(f'../output/lgb_cc_img_local_img_dl_cc_v2_all_ut.csv').head()
pd.read_csv(f'../output/ncf_all_xnn_action_v2_ncf_140k_multiclass_6_4.csv').head()
pd.read_csv('../input/submission_popular.csv').item_recommendations.values[:3]
sub = pd.read_csv('../input/submission_popular.csv')
sub.loc[sub.session_id== '1d688ec168932'].item_recommendations.values
sub.item_recommendations.apply(lambda x: x.split(' ')).apply(len).sum()
pd.read_csv('../output/ncf_all_xnn_dow_v2.csv').item_recommendations.apply(lambda x: x.split(' ')).apply(len).sum()
ncf['score'] = ncf.score / 2
pseudo_label = ncf.loc[:,['row_index','score']].groupby('row_index').score.apply(list)
pseudo_label = pseudo_label.reset_index()
pseudo_label.columns = ['session_id','pseudo_label']
pseudo_label.head()
# +
with open('../output/ncf_xnn_intpop_clickout_v2_all_ut_lgb_bf_beach_v2_all_ut_xgb_4_8_8_pseudo_label.p','wb') as f:
pickle.dump(pseudo_label, f)
# -
pseudo_label.shape, ncf.row_index.nunique()
| src/Merge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bitprj/DigitalHistory/blob/master/Week4-Introduction-to-data-visualization-and-graphs-with-matplotlib/Week4-homework/Week4-Homework.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_mwoK-OGd4Ie" colab_type="text"
# ## <div align="center">Lab 3</div>
#
#
# + [markdown] id="mTu386TreGYK" colab_type="text"
# # <div align="center">Malnutrition, A disease the world has forgotten</div>
# + id="UhAXLCCGd4If" colab_type="code" colab={}
# Run the Following Code
# !pip install descartes
# !pip install geopandas
# + [markdown] id="dlsnYt9Jd4Ik" colab_type="text"
# Welcome to Lab 3. In this lab, you will be using the techniques you learned throughout Week 3. The purpose of this lab for you to be able to practice visualization and the articulation of your observations. You will be asked to find answers to questions using ```matplotlib``` and ```pandas```. Because these topics are very new to you and can be challenging to absorb, we will be giving you hints and references to relevant sections in the week 3 lessons for some coding questions.
#
# By the end of the exercise, the deliverables are:
#
# | Questions | # of assignments|
# |---------------------------------------------------|-----------------|
# |Dataframe cleaning using pandas |1 |
# |Pandas Questions |4 |
# |Histogram plots |1 |
# |Scatter plots |4 |
# |Bar plots |4 |
# |Plot of the world map with the required features. |3 |
# |A comparison between two countries of your choice |3 |
# + [markdown] id="lcSP9Q-Sd4Ik" colab_type="text"
# ## About the Dataset
#
# ### Malnutrition EDA
# Malnutrition is a condition that results from eating a diet in which one or more nutrients are either not enough or are too much such that the diet causes health problems.
#
# Wasting: Also known as 'acute malnutrition', wasting is characterized by a rapid deterioration in nutritional status over a short time in children under five years of age. Wasted children are at higher risk of dying.
#
# Stunting is the impaired growth and development that children experience from poor nutrition, repeated infection, and inadequate psychosocial stimulation. Children are defined as stunted if their height-for-age is more than two standard deviations below the WHO Child Growth Standards median.
#
# **The data available**
#
# ```Severe Wasting``` - % of children aged 0–59 months who are below minus three standard deviations from median weight-for-height.
#
# ```Wasting``` – Moderate and severe: % of children aged 0–59 months who are below minus two standard deviations from median weight-for-height
#
# ```Overweight``` – Moderate and severe: % aged 0-59 months who are above two standard deviations from median weight-for-height
#
# ```Stunting``` – Moderate and severe:
# % of children aged 0–59 months who are below minus two standard deviations from median height-for-age
#
# ```Underweight``` – Moderate and severe: % of children aged 0–59 months who are below minus two standard deviations from median weight-for-age
#
# ```Income Classification``` - When it comes to income, the World Bank divides the world's economies into four income groups:
# - High
# - Upper-middle
# - Lower-middle
# - Low
#
# The income classification is based on a measure of national income per person, or GNI per capita, calculated using the Atlas method.
# + [markdown] id="_PEsPl-md4Im" colab_type="text"
# ## Import libraries
# + id="8w_mmDnbd4In" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] id="hBdbpiN9d4Ir" colab_type="text"
# ## Load the Dataset
# + id="K_UBZRDNd4Is" colab_type="code" colab={}
df = pd.read_csv('malnutrition/malnutrition.csv')
df
# + id="9VhFKkVKd4Iv" colab_type="code" colab={}
df = df.drop(columns = 'Unnamed: 0',axis = 1)
df
# + [markdown] id="cch8_GVGd4Iz" colab_type="text"
# ## Use pandas to display the basic information of the dataset
# + id="jYZ4YDUbd4I0" colab_type="code" colab={}
# head()
#INSERT CODE HERE
# + id="EMivw37Ed4I4" colab_type="code" colab={}
# tail()
#INSERT CODE HERE
# + id="zIP1psGFd4I9" colab_type="code" colab={}
# describe()
#INSERT CODE HERE
# + id="an8KHYV5d4JA" colab_type="code" colab={}
# info()
#INSERT CODE HERE
# + [markdown] id="SXxj5flxd4JD" colab_type="text"
# ### Q1. Check the total number of null values for each column and drop them [Graded]
# + id="bgKO_h7yd4JE" colab_type="code" colab={}
df.____().sum() #INSERT CODE HERE
# + id="SfdvGMUxd4JI" colab_type="code" colab={}
df = df.____() #INSERT CODE HERE
df
# + [markdown] id="oRrCuAz9d4JN" colab_type="text"
# ## Pandas Questions
# + [markdown] id="qeeM3IM-d4JO" colab_type="text"
# ### Q2. Which country shows the highest Underweight percentage? [Graded]
# + id="75j_MtPKd4JP" colab_type="code" colab={}
# + [markdown] id="ZZIenMzxd4JT" colab_type="text"
# ### Q3. Which country shows the highest Overweight percentage? [Graded]
# + id="oUR4b1-Ad4JU" colab_type="code" colab={}
# + [markdown] id="muzNndj6d4JX" colab_type="text"
# ### Q4. Top 5 Countries that are the most malnourished? [Graded]
# + id="sXqd1Hj0d4JX" colab_type="code" colab={}
# + [markdown] id="pZx68qDJd4Jd" colab_type="text"
# ### Q5. Top 5 Countries that are the most malnourished? [Graded]
# + id="LrnL8lB9d4Jd" colab_type="code" colab={}
# + [markdown] id="HWOVci7dd4Jg" colab_type="text"
# ## Histograms
# + [markdown] id="xy1gDau1d4Jh" colab_type="text"
# ### Q6. Unique Income Classification Histogram [Graded]
# + id="SlCJ2PjDd4Ji" colab_type="code" colab={}
# Check the number of Unique values int the 'Income Classification' column
df['INSERT COLUMN NAME'].______ #INSERT CODE HERE
# + id="5DNRiOPCd4Jo" colab_type="code" colab={}
# Display the histogram for the Income Classification column, set bins to 20
df["INSERT COLUMN NAME"].______ #INSERT CODE HERE
# + [markdown] id="G7uPj-Fbd4Jr" colab_type="text"
# ### Scatter Plots for Income Classification
#
# In this part, you will be drawing four scatterplots as subplots of a larger plot. The plot will have a figure size of (20,10).
#
# Throughout this exercise keep ```x``` as ```Income Classification```.
# + id="sXwI4Gz1d4Js" colab_type="code" colab={}
fig = plt.figure(figsize = (30,15))
ax1 = fig.add_subplot(2,2,1)
ax2 = fig.add_subplot(2,2,2)
ax3 = fig.add_subplot(2,2,3)
ax4 = fig.add_subplot(2,2,4)
# + [markdown] id="F9S8am6nd4Jv" colab_type="text"
# ### Q7. Wasting [Graded]
# + id="X3mAjqNpd4Jw" colab_type="code" colab={}
df.plot(# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
ax = # INSERT CODE HERE
_____='k') #INSERT CODE HERE
# + [markdown] id="7etIYyYPd4Jz" colab_type="text"
# ### Q8. Overweight [Graded]
# + id="l-8gRVuNd4Jz" colab_type="code" colab={}
df.plot(# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE,
# INSERT CODE HERE
)
# + [markdown] id="mza1wPpWd4J6" colab_type="text"
# ### Q9. Stunting [Graded]
# + id="yxoJIkfsd4J7" colab_type="code" colab={}
df.plot(# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE)
# + [markdown] id="soKBjprbd4KF" colab_type="text"
# ### Q10. U5 Population (\'000s) [Graded]
# + id="TEmaOYNYd4KG" colab_type="code" colab={}
df.plot(# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE)
# + id="pF1g59wed4KM" colab_type="code" colab={}
fig
# + [markdown] id="4SOQpQTUd4KQ" colab_type="text"
# ## Bar Graphs
# + [markdown] id="wf7uo-26d4KQ" colab_type="text"
# ### Q11. Plot a bar graph for the countries that have a waste percentage greater or equal to 15 [GRADED]
# + id="07gdjUMAd4KR" colab_type="code" colab={}
filt = ___________ >= ____ # INSERT CODE HERE
# find rows which match filt
max_waste = df._____(filt) # INSERT CODE HERE
# drop the null values
max_waste = max_waste.# INSERT CODE HERE
max_waste.plot(kind = 'bar',
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
figsize=(20,10))
# + [markdown] id="4HItFJUsd4KT" colab_type="text"
# ### Q12. Plot a bar graph for the countries that have a waste percentage less than or equal to 2 [GRADED]
# + id="2aGo1nKbd4KU" colab_type="code" colab={}
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
figsize=(20,10))
# + [markdown] id="2v7fj8-8d4KX" colab_type="text"
# ### Q13. Plot a bar graph showing all the countries with an income classification of 3.0
# + id="0NQwf352d4KY" colab_type="code" colab={}
filt = _________ == ____ # INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
HIGH_Income.plot(kind = 'bar',x = 'Country',
# INSERT CODE HERE,
figsize=(20,10))
# + [markdown] id="GH2MBMrbd4Kb" colab_type="text"
# ### Q14. Plot a bar graph showing all the countries with an Income Classification of 0 [GRADED]
# + id="1wKzg2mHd4Kc" colab_type="code" colab={}
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# + [markdown] id="mmkkbepid4Kg" colab_type="text"
# ## Plot the World Map
# + [markdown] id="aYHX82qBd4Kh" colab_type="text"
# As a bonus, I've added a feature that prints out the world map to make our visualizations more observable.
# + id="CDzzuhYrd4Kh" colab_type="code" colab={}
# Do not Modify
import descartes
import geopandas as gpd
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
# + [markdown] id="p_P8PLm6d4Kk" colab_type="text"
# ### 15. Plot the Longitude vs Latitude [GRADED]
#
# Set ```alpha``` to 0.6
# + id="ZiiIU0Xed4Kl" colab_type="code" colab={}
# Do not Modify the two lines below
fig,ax = plt.subplots(figsize=(30,15))
world.plot(ax=ax, color='lightgray')
df.plot(kind="scatter",
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HEREA
ax = ax)
plt.show()
# + [markdown] id="aa2i1_Xtd4Kn" colab_type="text"
# ### 16. Adding a color map for Income Classification [GRADED]
#
# Property: ```c : color, sequence, or sequence of color```
#
# Equate it to Income Classification.
#
# Also, add the following:
# - ```cmap: Colormap```
# - ```colorbar```
# - ```xlabel```
# - ```ylabel```
# - ```legend```
#
# **Note**: Features in the plot above must be present.
# + id="-_VQ4U1Bd4Ko" colab_type="code" colab={}
# Do not Modify the two lines below
fig, ax = plt.subplots(figsize=(20,10))
world.plot(ax=ax, color='lightgray')
df.plot(kind="scatter",
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
ax = ax)
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
plt.show()
# + [markdown] id="ekh8av45d4Ks" colab_type="text"
# ### 17. Adding a scalar pointer, U5-population
# Property: ```s : scalar or array_like, shape (n, )```
#
#
# Also, add the following:
# - Replace c with Stunting
# - Add a label for the feature
#
# **Note:** In order to get a viewable graph, you will have to divide the feature by a multiple of 10. Be careful, too high and your scalar points won't be distinguishable.
#
#
#
#
# + id="2RZ9XxKld4Kt" colab_type="code" colab={}
# Do not Modify the two lines below
fig, ax = plt.subplots(figsize=(30,15))
world.plot(ax=ax, color='lightgray')
df.plot(# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
ax = ax)
# INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
plt.show()
# + [markdown] id="OePrOur6d4Kw" colab_type="text"
# ## Compare two countries of your choice
#
#
# + id="v2U-u0qVd4Kw" colab_type="code" colab={}
fig = plt.figure(figsize=(20,10))
ax1 = fig.add_subplot(1,2,1)
ax2 = fig.add_subplot(1,2,2)
# We won't need the these columns
columns_to_remove = ['U5 Population (\'000s)','Longitude','Latitude']
# + [markdown] id="_Lrubv4ad4Kz" colab_type="text"
# ### 18. New DataFrame for Country 1 [GRADED]
#
# + id="sEp71RFrd4K0" colab_type="code" colab={}
filt = _________ == 'COUNTRY NAME'
# Make a new DataFrame using the filter
______ = df.____(____)
# Drop the columns in columns_to_remove
______ = # INSERT CODE HERE
# Drop all null values
______ = # INSERT CODE HERE
# view your newdataframe below
____
# + [markdown] id="pdRjvmGtd4K2" colab_type="text"
# ### 19. New DataFrame for Country 2 [GRADED]
# + id="L7wh-O6xd4K3" colab_type="code" colab={}
# Make a filter
# INSERT CODE HERE
# Make a new DataFrame using the filter
# INSERT CODE HERE
# Drop the columns in columns_to_remove
# INSERT CODE HERE
# Drop all null values
# INSERT CODE HERE
# view your newdataframe below
# INSERT CODE HERE
# + [markdown] id="q1du6Bded4K6" colab_type="text"
# ### 20. Plots bar charts for all features of both DataFrames [GRADED]
# + id="256-0rv1d4K6" colab_type="code" colab={}
# Country 1
_____.plot(# INSERT CODE HERE
# INSERT CODE HERE
ax = ax1)
# Country 2
#INSERT CODE HERE
# INSERT CODE HERE
# INSERT CODE HERE
)
# + id="3RDz_pM9d4LA" colab_type="code" colab={}
fig
# + [markdown] id="UynLVaoad4LE" colab_type="text"
# ## Appendix
#
#
# + [markdown] id="uEiHtr69d4LE" colab_type="text"
# ### How to Add latitude and Longitude
# + id="7zKYFLTBd4LF" colab_type="code" colab={}
from geopy.exc import GeocoderTimedOut
from geopy.geocoders import Nominatim
# declare an empty list to store
# latitude and longitude of values
# of city column
longitude = []
latitude = []
# function to find the coordinate
# of a given city
def findGeocode(city):
# try and catch is used to overcome
# the exception thrown by geolocator
# using geocodertimedout
try:
# Specify the user_agent as your
# app name it should not be None
geolocator = Nominatim(user_agent="your_app_name")
return geolocator.geocode(city)
except GeocoderTimedOut:
return findGeocode(city)
# each value from city column
# will be fetched and sent to
# function find_geocode
for i in (df["Country"]):
if findGeocode(i) != None:
loc = findGeocode(i)
# coordinates returned from
# function is stored into
# two separate list
latitude.append(loc.latitude)
longitude.append(loc.longitude)
# if coordinate for a city not
# found, insert "NaN" indicating
# missing value
else:
latitude.append(np.nan)
longitude.append(np.nan)
df['Latitude'] = latitude
df['Longitude'] = longitude
df.to_csv(r'../malnutrition/malnutrition.csv')
| Week3-Introduction-to-data-visualization-and-graphs-with-matplotlib/Week3-Lab/Week3-Lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
#
# # Lab 1: Generative Adversarial Networks
# In this lab, we will explore several architectures of generative adversarial networks. There are many different versions of GANs existing today. The break-up for today's lab is as follows:<ol>
# 1. Vanilla GAN
# 2. Conditional GAN
# 3. Cycle GAN
#
# These codes are modified from https://github.com/wiseodd/generative-models/tree/master/GAN
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Module 1: Vanilla GAN
# + slideshow={"slide_type": "slide"}
# Initialization of libraries
import torch
import torch.nn.functional as nn
import torch.autograd as autograd
import torch.optim as optim
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import os
from torch.autograd import Variable
import torchvision
import torchvision.transforms as transforms
device = torch.device('cuda')
# %matplotlib inline
# -
# defining parameters for the training
mb_size = 64 # Batch Size
Z_dim = 100 # Length of noise vector
X_dim = 784 # Input Length
h_dim = 128 # Hidden Dimension
lr = 1e-3 # Learning Rate
# + [markdown] slideshow={"slide_type": "notes"}
# Custom weights initialization function.
#
# (Read later about Xavier initialization: http://andyljones.tumblr.com/post/110998971763/an-explanation-of-xavier-initialization)
#
# (Link to paper: https://arxiv.org/abs/1406.2661)
# -
# Custom weight initialization
def xavier_init(size):
in_dim = size[0]
xavier_stddev = 1. / np.sqrt(in_dim / 2.)
vec = torch.randn(*size) * xavier_stddev
vec = vec.to(device)
vec.requires_grad = True
return vec
# + [markdown] slideshow={"slide_type": "slide"}
# We are using MNIST dataset for these experiments. Let us load the dataset. We will also define some functions that will be used for the other modules of this lab. Note that we are flattening the MNIST images into one dimensional vectors.
# + slideshow={"slide_type": "slide"}
dataroot = './data'
# Applying transformation to images
transform = transforms.Compose([transforms.ToTensor(), transforms.Normalize((0,), (1,))])
# defining dataloader for the images
trainset = torchvision.datasets.MNIST(root=dataroot , train=True, download=False,transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=mb_size, shuffle=True, num_workers=2)
# function to get the next batch
def mnist_next(dataiter):
try:
images, labels = dataiter.next()
images = images.view(images.numpy().shape[0],28*28)
except:
dataiter = iter(trainloader)
images, labels = dataiter.next()
images = images.view(images.numpy().shape[0],28*28)
return images.numpy(), labels
# Initialization of dataloader
def initialize_loader(trainset):
trainloader = torch.utils.data.DataLoader(trainset, batch_size=mb_size, shuffle=True, num_workers=2)
dataiter = iter(trainloader)
return dataiter
# -
# print some random training images
dataiter = iter(trainloader)
images, labels = dataiter.next()
img = torchvision.utils.make_grid(images)
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
# + [markdown] slideshow={"slide_type": "slide"}
# Now, let us define the network structure. For this experiment, we are not using deep networks. We are not even using torch.nn layers. Instead, we will use simple linear fully connected layers. The generator and discriminators are 2-layer networks. This is why we flattened the images in the block above.
# + slideshow={"slide_type": "slide"}
""" ==================== GENERATOR ======================== """
# generator weight initialization
Wzh = xavier_init(size=[Z_dim, h_dim])
Whx = xavier_init(size=[h_dim, X_dim])
# generator bias initialization
bzvar = torch.zeros(h_dim);
bhvar = torch.zeros(X_dim);
bzvar = bzvar.to(device)
bhvar = bhvar.to(device)
bzh = Variable(bzvar, requires_grad=True)
bhx = Variable(bhvar, requires_grad=True)
# trainable parameters of generator
G_params = [Wzh, bzh, Whx, bhx]
# -
# Network architecture of generator
def G(z):
h = nn.relu(torch.mm(z, Wzh) + bzh.repeat(z.size(0), 1))
X = nn.sigmoid(torch.mm(h, Whx) + bhx.repeat(h.size(0), 1))
return X
# + slideshow={"slide_type": "slide"}
""" ==================== DISCRIMINATOR ======================== """
# Discriminator weight initialization
Wxh = xavier_init(size=[X_dim, h_dim])
Why = xavier_init(size=[h_dim, 1])
# Discriminator bias initialization
bxvar = torch.zeros(h_dim)
bhvar = torch.zeros(1)
bxvar = bxvar.to(device)
bhvar = bhvar.to(device)
bxh = Variable(bxvar, requires_grad=True)
bhy = Variable(bhvar, requires_grad=True)
# trainable parameters of discriminator
D_params = [Wxh, bxh, Why, bhy]
# -
# Network architecture of discriminator
def D(X):
h = nn.relu(torch.mm(X, Wxh) + bxh.repeat(X.size(0), 1))
y = nn.sigmoid(torch.mm(h, Why) + bhy.repeat(h.size(0), 1))
return y
# + [markdown] slideshow={"slide_type": "slide"}
# Here, we will gather the parameters of the generator and the discriminator so that they can be given to the Adam optimizer to update the weights
# + slideshow={"slide_type": "slide"}
G_solver = optim.Adam(G_params, lr)
D_solver = optim.Adam(D_params, lr)
ones_label = torch.ones(mb_size,1)
zeros_label = torch.zeros(mb_size,1)
ones_label = ones_label.to(device)
zeros_label = zeros_label.to(device)
# -
# Reset the gradients to zero
params = G_params + D_params
def reset_grad():
for p in params:
p.grad.data.zero_()
# + [markdown] slideshow={"slide_type": "slide"}
# Now, we will start the actual training. The training alternates between updating the discriminator network's weights and updating the generator's weight.First, we update the discriminator's weight. We take a minibatch from the dataset and do a forward pass on the discriminator with the label '1'. Then, we feed noise into the generator and feed the generated images into the discriminator with the label '0'. We backpropagate the error and update the discriminator weights. To update the generator weights, we feed noise to the generator and feed the generated images into the discriminator with the label '1'. This error is backpropagated to update the weights of G.
# +
dataiter = initialize_loader(trainset)
for it in range(10000):
# Sample data
z = torch.randn(mb_size, Z_dim)
X, _ = mnist_next(dataiter)
if X.shape[0]!=mb_size:
dataiter = initialize_loader(trainset)
X,_ = mnist_next(dataiter)
X = torch.from_numpy(X)
X = X.to(device)
z = z.to(device)
# Dicriminator forward-loss-backward-update
#forward pass
G_sample = G(z)
D_real = D(X)
D_fake = D(G_sample)
# Calculate the loss
D_loss_real = nn.binary_cross_entropy(D_real, ones_label)
D_loss_fake = nn.binary_cross_entropy(D_fake, zeros_label)
D_loss = D_loss_real + D_loss_fake
# Calulate and update gradients of discriminator
D_loss.backward()
D_solver.step()
# reset gradient
reset_grad()
# Generator forward-loss-backward-update
z = torch.randn(mb_size, Z_dim)
z = z.to(device)
G_sample = G(z)
D_fake = D(G_sample)
G_loss = nn.binary_cross_entropy(D_fake, ones_label)
G_loss.backward()
G_solver.step()
# reset gradient
reset_grad()
# Print and plot every now and then
if it % 1000 == 0:
print('Iter-{}; D_loss: {}; G_loss: {}'.format(it, D_loss.data.cpu().numpy(), G_loss.data.cpu().numpy()))
# -
# Let us see the images generated by the generator:
z = torch.randn(mb_size, Z_dim)
z = z.to(device)
samples = G(z)
samples = samples.cpu()
img = samples.data
img = img.view([64,1,28,28])
img = torchvision.utils.make_grid(img)
img = img.permute(1,2,0)
plt.imshow(img.numpy())
# ### Image interpolation
#
# Let us try to interpolate between images
# +
# Generate 64 random noise vector by interpolating between two noise vector
a = torch.randn(Z_dim);
b = torch.randn(Z_dim);
noise = torch.Tensor(mb_size,Z_dim)
line = torch.linspace(0, 1, mb_size)
for i in range(0, mb_size):
noise.select(0, i).copy_(a * line[i] + b * (1 - line[i]))
noise = noise.to(device)
# Feed forward the interpolated noise vector to generator
samples = G(noise)
samples = samples.cpu()
# Plot the generated images
img = samples.data
img = img.view([mb_size,1,28,28])
img = torchvision.utils.make_grid(img)
img = img.permute(1,2,0)
plt.imshow(img.numpy())
# -
# ## Points to ponder
# 1. What happens if we don't do Xavier initialization?
# 2. What happens if we change the learning rate and other parameters?
# 3. What happens if we reduce the size of hidden layer to 10?
# 4. Is there any way to determine the class of the generated images by changing the input noise vector?
# 5. How long do we have to train the GAN to get good results? Can you plot the loss of the generator and discriminator and see if there is a correlation?
#
| Day 5 GAN/1-GANs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Tensor operations and GradientTape.
#
# In this graded assignment, you will perform different tensor operations as well as use [GradientTape](https://www.tensorflow.org/api_docs/python/tf/GradientTape). These are important building blocks for the next parts of this course so it's important to master the basics. Let's begin!
# + colab={} colab_type="code" id="jqev488WJ9-R"
import tensorflow as tf
import numpy as np
# -
# ## Exercise 1 - [tf.constant]((https://www.tensorflow.org/api_docs/python/tf/constant))
#
# Creates a constant tensor from a tensor-like object.
# + colab={} colab_type="code" id="MYdVyiSoLPgO"
# Convert NumPy array to Tensor using `tf.constant`
def tf_constant(array):
"""
Args:
array (numpy.ndarray): tensor-like array.
Returns:
tensorflow.python.framework.ops.EagerTensor: tensor.
"""
### START CODE HERE ###
tf_constant_array = tf.constant(array)
### END CODE HERE ###
return tf_constant_array
# +
tmp_array = np.arange(1,10)
x = tf_constant(tmp_array)
x
# Expected output:
# <tf.Tensor: shape=(9,), dtype=int64, numpy=array([1, 2, 3, 4, 5, 6, 7, 8, 9])>
# -
# Note that for future docstrings, the type `EagerTensor` will be used as a shortened version of `tensorflow.python.framework.ops.EagerTensor`.
# ## Exercise 2 - [tf.square](https://www.tensorflow.org/api_docs/python/tf/math/square)
#
# Computes the square of a tensor element-wise.
# + colab={} colab_type="code" id="W6BTwNJCLjV8"
# Square the input tensor
def tf_square(array):
"""
Args:
array (numpy.ndarray): tensor-like array.
Returns:
EagerTensor: tensor.
"""
# make sure it's a tensor
array = tf.constant(array)
### START CODE HERE ###
tf_squared_array = array ** 2
### END CODE HERE ###
return tf_squared_array
# +
tmp_array = tf.constant(np.arange(1, 10))
x = tf_square(tmp_array)
x
# Expected output:
# <tf.Tensor: shape=(9,), dtype=int64, numpy=array([ 1, 4, 9, 16, 25, 36, 49, 64, 81])>
# -
# ## Exercise 3 - [tf.reshape](https://www.tensorflow.org/api_docs/python/tf/reshape)
#
# Reshapes a tensor.
# + colab={} colab_type="code" id="7nzBSX8-L0Xt"
# Reshape tensor into the given shape parameter
def tf_reshape(array, shape):
"""
Args:
array (EagerTensor): tensor to reshape.
shape (tuple): desired shape.
Returns:
EagerTensor: reshaped tensor.
"""
# make sure it's a tensor
array = tf.constant(array)
### START CODE HERE ###
tf_reshaped_array = tf.reshape(array, shape)
### END CODE HERE ###
return tf_reshaped_array
# +
# Check your function
tmp_array = np.array([1,2,3,4,5,6,7,8,9])
# Check that your function reshapes a vector into a matrix
x = tf_reshape(tmp_array, (3, 3))
x
# Expected output:
# <tf.Tensor: shape=(3, 3), dtype=int64, numpy=
# [[1, 2, 3],
# [4, 5, 6],
# [7, 8, 9]]
# -
# ## Exercise 4 - [tf.cast](https://www.tensorflow.org/api_docs/python/tf/cast)
#
# Casts a tensor to a new type.
# + colab={} colab_type="code" id="VoT-jiAIL8x5"
# Cast tensor into the given dtype parameter
def tf_cast(array, dtype):
"""
Args:
array (EagerTensor): tensor to be casted.
dtype (tensorflow.python.framework.dtypes.DType): desired new type. (Should be a TF dtype!)
Returns:
EagerTensor: casted tensor.
"""
# make sure it's a tensor
array = tf.constant(array)
### START CODE HERE ###
tf_cast_array = tf.cast(array, dtype)
### END CODE HERE ###
return tf_cast_array
# +
# Check your function
tmp_array = [1,2,3,4]
x = tf_cast(tmp_array, tf.float32)
x
# Expected output:
# <tf.Tensor: shape=(4,), dtype=float32, numpy=array([1., 2., 3., 4.], dtype=float32)>
# -
# ## Exercise 5 - [tf.multiply](https://www.tensorflow.org/api_docs/python/tf/multiply)
#
# Returns an element-wise x * y.
# + colab={} colab_type="code" id="ivepGtD5MKP5"
# Multiply tensor1 and tensor2
def tf_multiply(tensor1, tensor2):
"""
Args:
tensor1 (EagerTensor): a tensor.
tensor2 (EagerTensor): another tensor.
Returns:
EagerTensor: resulting tensor.
"""
# make sure these are tensors
tensor1 = tf.constant(tensor1)
tensor2 = tf.constant(tensor2)
### START CODE HERE ###
product = tensor1 * tensor2
### END CODE HERE ###
return product
# +
# Check your function
tmp_1 = tf.constant(np.array([[1,2],[3,4]]))
tmp_2 = tf.constant(np.array(2))
result = tf_multiply(tmp_1, tmp_2)
result
# Expected output:
# <tf.Tensor: shape=(2, 2), dtype=int64, numpy=
# array([[2, 4],
# [6, 8]])>
# -
# ## Exercise 6 - [tf.add](https://www.tensorflow.org/api_docs/python/tf/add)
#
# Returns x + y element-wise.
# + colab={} colab_type="code" id="BVlntdYnMboh"
# Add tensor1 and tensor2
def tf_add(tensor1, tensor2):
"""
Args:
tensor1 (EagerTensor): a tensor.
tensor2 (EagerTensor): another tensor.
Returns:
EagerTensor: resulting tensor.
"""
# make sure these are tensors
tensor1 = tf.constant(tensor1)
tensor2 = tf.constant(tensor2)
### START CODE HERE ###
total = tensor1+tensor2
### END CODE HERE ###
return total
# +
# Check your function
tmp_1 = tf.constant(np.array([1, 2, 3]))
tmp_2 = tf.constant(np.array([4, 5, 6]))
tf_add(tmp_1, tmp_2)
# Expected output:
# <tf.Tensor: shape=(3,), dtype=int64, numpy=array([5, 7, 9])>
# + [markdown] colab_type="text" id="9EN0W15EWNjD"
# ## Exercise 7 - Gradient Tape
#
# Implement the function `tf_gradient_tape` by replacing the instances of `None` in the code below. The instructions are given in the code comments.
#
# You can review the [docs](https://www.tensorflow.org/api_docs/python/tf/GradientTape) or revisit the lectures to complete this task.
# + colab={} colab_type="code" id="p3K94BWZM6nW"
def tf_gradient_tape(x):
"""
Args:
x (EagerTensor): a tensor.
Returns:
EagerTensor: Derivative of z with respect to the input tensor x.
"""
with tf.GradientTape() as t:
### START CODE HERE ###
# Record the actions performed on tensor x with `watch`
t.watch(x)
# Define a polynomial of form 3x^3 - 2x^2 + x
y = 3*x**3 - 2*x**2 + x
# Obtain the sum of the elements in variable y
z = tf.reduce_sum(y)
# Get the derivative of z with respect to the original input tensor x
dz_dx = t.gradient(z, x)
### END CODE HERE
return dz_dx
# +
# Check your function
tmp_x = tf.constant(2.0)
dz_dx = tf_gradient_tape(tmp_x)
result = dz_dx.numpy()
result
# Expected output:
# 29.0
# -
# **Congratulations on finishing this week's assignment!**
#
# **Keep it up!**
| C2_custom_training/C2_W1_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# # Trading Platform Customer Attrition Risk Prediction using sklearn
#
# There are many users of online trading platforms and these companies would like to run analytics on and predict churn based on user activity on the platform. Since competition is rife, keeping customers happy so they do not move their investments elsewhere is key to maintaining profitability.
#
# In this notebook, we will leverage Watson Studio Local (that is a service on IBM Cloud Pak for Data) to do the following:
#
# 1. Ingest merged customer demographics and trading activity data
# 2. Visualize merged dataset and get better understanding of data to build hypotheses for prediction
# 3. Leverage sklearn library to build classification model that predicts whether customer has propensity to churn
# 4. Expose the classification model as RESTful API endpoint for the end-to-end customer churn risk prediction and risk remediation application
#
# <img src="https://github.com/burtvialpando/CloudPakWorkshop/blob/master/CPD/images/NotebookImage.png?raw=true" width="800" height="500" align="middle"/>
#
#
# <a id="top"></a>
# ## Table of Contents
#
# 1. [Load libraries](#load_libraries)
# 2. [Load and visualize merged customer demographics and trading activity data](#load_data)
# 3. [Prepare data for building classification model](#prepare_data)
# 4. [Train classification model and test model performance](#build_model)
# 5. [Save model to ML repository and expose it as REST API endpoint](#save_model)
# 6. [Summary](#summary)
# ### Quick set of instructions to work through the notebook
#
# If you are new to Notebooks, here's a quick overview of how to work in this environment.
#
# 1. The notebook has 2 types of cells - markdown (text) such as this and code such as the one below.
# 2. Each cell with code can be executed independently or together (see options under the Cell menu). When working in this notebook, we will be running one cell at a time because we need to make code changes to some of the cells.
# 3. To run the cell, position cursor in the code cell and click the Run (arrow) icon. The cell is running when you see the * next to it. Some cells have printable output.
# 4. Work through this notebook by reading the instructions and executing code cell by cell. Some cells will require modifications before you run them.
# <a id="load_libraries"></a>
# ## 1. Load libraries
# [Top](#top)
#
# Running the following cell will load all libraries needed to load, visualize, prepare the data and build ML models for our use case
#Uncomment and run once to install the package in your runtime environment
# !pip install sklearn-pandas==1.8.0
# If the following cell doesn't work, please un-comment out the next line and do upgrade the patplotlib package. When the upgrade is done, restart the kernal and start from the beginning again.
# !pip install --user --upgrade matplotlib
import brunel
import pandas as pd
import numpy as np
import sklearn.pipeline
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler, LabelEncoder, StandardScaler, LabelBinarizer, OneHotEncoder
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import f1_score, accuracy_score, roc_curve, roc_auc_score
from sklearn_pandas import DataFrameMapper
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
import json
import matplotlib.pyplot as plt
# %matplotlib inline
#Changed sk-learn version to be compatible with WML client4 on CPD v3.0.1
# !pip install scikit-learn==0.22
# # <a id="load_data"></a>
# ## 2. Load data example
# [Top](#top)
#
# Data can be easily loaded within ICPD using point-and-click functionality. The following image illustrates how to load a merged dataset assuming it is called "customer_demochurn_activity_analyze.csv". The file can be located by its name and inserted into the notebook as a **pandas** dataframe as shown below:
#
# <img src="https://github.com/burtvialpando/CloudPakWorkshop/blob/master/CPD/images/InsertPandasDataFrame.png?raw=true" width="300" height="300" align="middle"/>
#
# The interface comes up with a generic name, so it is good practice to rename the dataframe to match context of the use case. In this case, we will use df_churn.
df_churn_pd = pd.read_csv('/project_data/data_asset/customer_demochurn_activity_analyze.csv')
df_churn_pd.head()
df_churn_pd.dtypes
df_churn_pd.describe()
# %brunel data('df_churn_pd') stack polar bar x(CHURNRISK) y(#count) color(CHURNRISK) bar tooltip(#all)
# %brunel data('df_churn_pd') bar x(STATUS) y(#count) color(STATUS) tooltip(#all) | stack bar x(STATUS) y(#count) color(CHURNRISK: pink-orange-yellow) bin(STATUS) sort(STATUS) percent(#count) label(#count) tooltip(#all) :: width=1200, height=350
# %brunel data('df_churn_pd') bar x(TOTALUNITSTRADED) y(#count) color(CHURNRISK: pink-gray-orange) sort(STATUS) percent(#count) label(#count) tooltip(#all) :: width=1200, height=350
# %brunel data('df_churn_pd') bar x(DAYSSINCELASTTRADE) y(#count) color(CHURNRISK: pink-gray-orange) sort(STATUS) percent(#count) label(#count) tooltip(#all) :: width=1200, height=350
# <a id="prepare_data"></a>
# ## 3. Data preparation
# [Top](#top)
#
# Data preparation is a very important step in machine learning model building. This is because the model can perform well only when the data it is trained on is good and well prepared. Hence, this step consumes bulk of data scientist's time spent building models.
#
# During this process, we identify categorical columns in the dataset. Categories needed to be indexed, which means the string labels are converted to label indices. These label indices and encoded using One-hot encoding to a binary vector with at most a single one-value indicating the presence of a specific feature value from among the set of all feature values. This encoding allows algorithms which expect continuous features to use categorical features.
#
# Final step in the data preparation process is to assemble all the categorical and non-categorical columns into a feature vector. We use VectorAssembler for this. VectorAssembler is a transformer that combines a given list of columns into a single vector column. It is useful for combining raw features and features generated by different feature transformers into a single feature vector, in order to train ML models.
# #### Use the DataFrameMapper class to declare transformations and variable imputations.
#
# * LabelBinarizer - Converts a categorical variable into a dummy variable (aka binary variable)
# * StandardScaler - Standardize features by removing the mean and scaling to unit variance, z = (x - u) / s
#
# See docs:
# * https://github.com/scikit-learn-contrib/sklearn-pandas
# * https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html#sklearn.preprocessing.StandardScaler
# * https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelBinarizer.html#sklearn.preprocessing.LabelBinarizer
# * https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html
# Defining the categorical columns
categoricalColumns = ['GENDER', 'STATUS', 'HOMEOWNER', 'AGE_GROUP']
numericColumns = ['CHILDREN', 'ESTINCOME', 'TOTALDOLLARVALUETRADED', 'TOTALUNITSTRADED', 'LARGESTSINGLETRANSACTION', 'SMALLESTSINGLETRANSACTION',
'PERCENTCHANGECALCULATION', 'DAYSSINCELASTLOGIN', 'DAYSSINCELASTTRADE', 'NETREALIZEDGAINS_YTD', 'NETREALIZEDLOSSES_YTD']
mapper = DataFrameMapper([
(['GENDER'], LabelBinarizer()),
(['STATUS'], LabelBinarizer()),
(['HOMEOWNER'], LabelBinarizer()),
(['AGE_GROUP'], LabelBinarizer()),
(['CHILDREN'], StandardScaler()),
(['ESTINCOME'], StandardScaler()),
(['TOTALDOLLARVALUETRADED'], StandardScaler()),
(['TOTALUNITSTRADED'], StandardScaler()),
(['LARGESTSINGLETRANSACTION'], StandardScaler()),
(['SMALLESTSINGLETRANSACTION'], StandardScaler()),
(['PERCENTCHANGECALCULATION'], StandardScaler()),
(['DAYSSINCELASTLOGIN'], StandardScaler()),
(['DAYSSINCELASTTRADE'], StandardScaler()),
(['NETREALIZEDGAINS_YTD'], StandardScaler()),
(['NETREALIZEDLOSSES_YTD'], StandardScaler())], default=False)
df_churn_pd.columns
# Define input data to the model
X = df_churn_pd.drop(['ID','CHURNRISK','AGE','TAXID','CREDITCARD','DOB','ADDRESS_1', 'ADDRESS_2', 'CITY', 'STATE', 'ZIP', 'ZIP4', 'LONGITUDE',
'LATITUDE'], axis=1)
X.shape
# Define the target variable and encode with value between 0 and n_classes-1
le = LabelEncoder()
y = le.fit_transform(df_churn_pd['CHURNRISK'])
# split the data to training and testing set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=5)
# <a id="build_model"></a>
# ## 4. Build Random Forest classification model
# [Top](#top)
# We instantiate a decision-tree based classification algorithm, namely, RandomForestClassifier. Next we define a pipeline to chain together the various transformers and estimaters defined during the data preparation step before. Sklearn standardizes APIs for machine learning algorithms to make it easier to combine multiple algorithms into a single pipeline, or workflow.
#
# We split original dataset into train and test datasets. We fit the pipeline to training data and apply the trained model to transform test data and generate churn risk class prediction
import warnings
warnings.filterwarnings("ignore")
# +
# Instantiate the Classifier
random_forest = RandomForestClassifier(random_state=5)
# Define the steps in the pipeline to sequentially apply a list of transforms and the estimator, i.e. RandomForestClassifier
steps = [('mapper', mapper),('RandonForestClassifier', random_forest)]
pipeline = sklearn.pipeline.Pipeline(steps)
# train the model
model=pipeline.fit( X_train, y_train )
model
# -
### call pipeline.predict() on your X_test data to make a set of test predictions
y_prediction = model.predict( X_test )
# show first 10 rows of predictions
y_prediction[0:10,]
# show first 10 rows of predictions with the corresponding labels
le.inverse_transform(y_prediction)[0:10]
# ### Model results
#
# In a supervised classification problem such as churn risk classification, we have a true output and a model-generated predicted output for each data point. For this reason, the results for each data point can be assigned to one of four categories:
#
# 1. True Positive (TP) - label is positive and prediction is also positive
# 2. True Negative (TN) - label is negative and prediction is also negative
# 3. False Positive (FP) - label is negative but prediction is positive
# 4. False Negative (FN) - label is positive but prediction is negative
#
# These four numbers are the building blocks for most classifier evaluation metrics. A fundamental point when considering classifier evaluation is that pure accuracy (i.e. was the prediction correct or incorrect) is not generally a good metric. The reason for this is because a dataset may be highly unbalanced. For example, if a model is designed to predict fraud from a dataset where 95% of the data points are not fraud and 5% of the data points are fraud, then a naive classifier that predicts not fraud, regardless of input, will be 95% accurate. For this reason, metrics like precision and recall are typically used because they take into account the type of error. In most applications there is some desired balance between precision and recall, which can be captured by combining the two into a single metric, called the F-measure.
# display label mapping to assist with interpretation of the model results
label_mapping=le.inverse_transform([0,1,2])
print('0: ', label_mapping[0])
print('1: ', label_mapping[1])
print('2: ', label_mapping[2])
# +
### test your predictions using sklearn.classification_report()
report = sklearn.metrics.classification_report( y_test, y_prediction )
### and print the report
print(report)
# -
print('Accuracy: ',sklearn.metrics.accuracy_score( y_test, y_prediction ))
# #### Get the column names of the transformed features
m_step=pipeline.named_steps['mapper']
m_step.transformed_names_
features = m_step.transformed_names_
# Get the features importance
importances = pipeline.named_steps['RandonForestClassifier'][1].feature_importances_
indices = np.argsort(importances)
plt.figure(1)
plt.title('Feature Importances')
plt.barh(range(len(indices)), importances[indices], color='b',align='center')
plt.yticks(range(len(indices)), (np.array(features))[indices])
plt.xlabel('Relative Importance')
# <a id="save_model"></a>
# ## 5. Save the model into WML Deployment Space
# [Top](#top)
# Before we save the model we must create a deployment space. Watson Machine Learning provides deployment spaces where the user can save, configure and deploy their models. We can save models, functions and data assets in this space.
#
# The steps involved for saving and deploying the model are as follows:
#
# 1. Create a new deployment space. Enter the name of the space in the cell below. If a space with specified space_name already exists, existing space will be deleted before creating a new space.
# 2. Set this deployment space as the default space.
# 3. Store the model pipeline in the deployment space. Enter the name for the model in the cell below.
# 4. Deploy the saved model. Enter the deployment name in the cell below.
# 5. Retrieve the scoring endpoint to score the model with a payload
# 6. We will use the watson_machine_learning_client package to complete these steps.
# !pip install watson-machine-learning-client-v4
# +
# Specify a names for the space being created, the saved model and the model deployment
space_name = 'churnrisk_deployment_space'
model_name = 'churnrisk_model_nb'
deployment_name = 'churnrisk_model_deployment'
# +
from watson_machine_learning_client import WatsonMachineLearningAPIClient
import os
token = os.environ['USER_ACCESS_TOKEN']
from project_lib.utils import environment
url = environment.get_common_api_url()
wml_credentials = {
"token": token,
"instance_id" : "wml_local",
"url": url,
"version": "3.0.0"
}
client = WatsonMachineLearningAPIClient(wml_credentials)
# -
# If a space with specified space_name already exists, delete the existing space before creating a new one.
# +
for space in client.spaces.get_details()['resources']:
if space_name in space['entity']['name']:
client.spaces.delete(space['metadata']['guid'])
print(space_name, "is deleted")
# -
# ### 5.1 Create Deployment Space
# +
# create the space and set it as default
space_meta_data = {
client.spaces.ConfigurationMetaNames.NAME : space_name
}
stored_space_details = client.spaces.store(space_meta_data)
space_uid = stored_space_details['metadata']['guid']
# set the newly created deployment space as the default
client.set.default_space(space_uid)
# -
# fetching details of the space created
stored_space_details
# ### 5.2 Store the model in the deployment space
# list all supported software specs
client.software_specifications.list()
# run this line if you do not know the version of scikit-learn that was used to build the model
# !pip show scikit-learn
software_spec_uid = client.software_specifications.get_uid_by_name('scikit-learn_0.22-py3.6')
# +
metadata = {
client.repository.ModelMetaNames.NAME: model_name,
client.repository.ModelMetaNames.SOFTWARE_SPEC_UID: software_spec_uid,
client.repository.ModelMetaNames.TYPE: "scikit-learn_0.22"
}
stored_model_details = client.repository.store_model(pipeline,
meta_props=metadata,
training_data=X_train,
training_target=y_train)
# -
stored_model_details
# ### 5.3 Create a deployment for the stored model
# +
# deploy the model
meta_props = {
client.deployments.ConfigurationMetaNames.NAME: deployment_name,
client.deployments.ConfigurationMetaNames.ONLINE: {}
}
# deploy the model
model_uid = stored_model_details["metadata"]["guid"]
deployment_details = client.deployments.create( artifact_uid=model_uid, meta_props=meta_props)
# -
# ### 5.4 Score the model
# +
# retrieve the scoring endpoint
scoring_endpoint = client.deployments.get_scoring_href(deployment_details)
print('Scoring Endpoint: ',scoring_endpoint)
# -
scoring_deployment_id = client.deployments.get_uid(deployment_details)
client.deployments.get_details(scoring_deployment_id)
payload = [{"values": [ ['Young adult','M','S', 2,56000, 'N', 5030, 23, 2257, 125, 3.45, 2, 19, 1200, 251]]}]
payload_metadata = {client.deployments.ScoringMetaNames.INPUT_DATA: payload}
# score
predictions = client.deployments.score(scoring_deployment_id, payload_metadata)
predictions
# display label mapping to assist with interpretation of the model results
label_mapping=le.inverse_transform([0,1,2])
print('0: ', label_mapping[0])
print('1: ', label_mapping[1])
print('2: ', label_mapping[2])
# #### Write test data into .csv files for batch scoring and model evaluations
# Write the test data a .csv so that we can later use it for batch scoring
write_score_CSV=X_test
write_score_CSV.to_csv('/project_data/data_asset/model_batch_score.csv', sep=',', index=False)
# Write the test data to a .csv so that we can later use it for Evaluation
write_eval_CSV=X_test
write_eval_CSV.to_csv('/project_data/data_asset/model_eval.csv', sep=',', index=False)
# **Last updated:** 06/01/2020 - Original Notebook by <NAME>, updated in later versions by <NAME>. Final edits by <NAME> and <NAME> - IBM
| TradingCustomerChurnClassifier-CP4D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''CV'': conda)'
# name: python379jvsc74a57bd072595be4ea15549328747a62d7632306f334cff511fe79c2306ce58b7e7768b4
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import scipy.signal as ss
import gc
import copy
from PIL import Image
from matplotlib.pyplot import imshow
from sklearn.base import BaseEstimator, ClassifierMixin
from os import listdir, mkdir
from os.path import isfile, join
import math
from datetime import datetime
import time
from skimage.transform import radon
from skimage import color, data, restoration
from scipy import ndimage
from IPython.display import clear_output
from typing import Optional
import torch
# from method import momentum_method
import os.path
import seaborn as sns
# +
def FixImage(image):
'''
Returns image with values in [0, 1] segment
for normal output with possible negative elements
'''
min_value = image.min()
max_value = image.max()
if min_value < 0:
image -= min_value
return image / (max_value - min_value)
def images_out(class_elem):
'''
Relatively normal output
of _cur_image and _init_image
in element of FunctionalMinimisation class
'''
plt.figure(figsize=(35,35))
plt.subplot(1,2,1)
plt.imshow(FixImage(class_elem._cur_image), cmap='gray')
plt.subplot(1,2,2)
plt.imshow(FixImage(class_elem._init_image), cmap='gray')
def save_img(class_elem, p='my_stuff.png', p_b='blurred.png'):
plt.imsave(p, class_elem._cur_image, cmap='gray')
plt.imsave(p_b, class_elem._init_image, cmap='gray')
DEBUG = True
def rgb2gray(rgb):
r, g, b = rgb[:,:,0], rgb[:,:,1], rgb[:,:,2]
gray = 0.299 * r + 0.5870 * g + 0.1140 * b
return gray
def blend_images(orig, four, alpha = 0.8, colH = 10, w=1):
orig_img = Image.open(orig).convert('RGBA')
fourier_img = Image.open(four).convert('RGBA')
orig_img = orig_img.resize(((fourier_img.size[0], fourier_img.size[1])), Image.BICUBIC)
img = fourier_img
A = img.getchannel('A')
# Convert to HSV and save the V (Lightness) channel
V = img.convert('RGB').convert('HSV').getchannel('V')
# Synthesize new Hue and Saturation channels using values from colour picker
colpickerH, colpickerS = colH, 255
newH=Image.new('L',img.size,(colpickerH))
newS=Image.new('L',img.size,(colpickerS))
# Recombine original V channel plus 2 synthetic ones to a 3 channel HSV image
HSV = Image.merge('HSV', (newH, newS, V))
# Add original Alpha layer back in
R,G,B = HSV.convert('RGB').split()
RGBA = Image.merge('RGBA',(R,G,B,A))
new_img = Image.blend(orig_img, RGBA, alpha)
return new_img
def calculate_log(picture,threshold=0.5):
log = np.log(np.abs(np.fft.fft2(Cepstrum.hamming(picture))))
fourier_abs = np.abs(log)
return fourier_abs
def wiener_filter(img, kernel, K=1):
if np.shape(kernel)[0] == 0:
return img
kernel /= np.sum(kernel)
dummy = np.copy(img)
dummy = np.fft.fft2(dummy)
kernel = np.fft.fft2(kernel, s = img.shape)
kernel = np.conj(kernel) / (np.abs(kernel) ** 2 + K)
dummy = dummy * kernel
dummy = np.abs(np.fft.ifft2(dummy))
return dummy
def get_blur_len(img, angle, weight, w=1):
rotated_img = ndimage.rotate(img, -angle * 180/math.pi)
rotated_img[rotated_img < 4/255 * rotated_img.max()] = 0
r = radon(rotated_img, theta=[90], circle=False)
r[r > 0.6 * r.max()] = 0
r *= 1./max(r)
blur_len = 0
for i in range(len(r)):
if (r[i] > 0.7):
blur_len = len(r) // 2 - 1 - i
# if (blur_len > 2 * img.shape[0] // 5):
# blur_len = 0
break
if (DEBUG):
h = img.shape[0]
q = h // 2 - 1
k = -math.tan(angle)
b = (1 - k) * q
l = []
if abs(abs(angle * 180/math.pi) - 90) > 10:
for old_x in range(q - blur_len, q + blur_len):
old_y = round(k * old_x+b)
old_y = int((old_y if old_y >= 0 else 0) if old_y <= h-1 else h-1)
if (old_y <= 1 or old_y >= h-2 or old_x <= 1 or old_x >= h-2):
continue
for i in range(-w, w+1):
for j in range(-w, w+1):
x = old_x
y = old_y
y += i
y = (y if y >= 0 else 0) if y <= h-1 else h-1
x += j
x = (x if x >= 0 else 0) if x <= h-1 else h-1
if (y, x) not in l:
l.append((y, x))
else:
for y in range(q - blur_len, q + blur_len):
for i in range(-w, w+1):
if (y, q + i) not in l:
l.append((y, q + i))
p = np.zeros((h, h))
for t in l:
p[t] = weight
return (int(abs(blur_len)), p)
else:
return int(abs(blur_len))
def find_best_line(template_picture, dif=180):
h = template_picture.shape[0]
q = h // 2 - 1
theta = np.linspace(0., 180, dif, endpoint=False)
sinogram = radon(template_picture, theta=theta, circle=True)
max_values = sinogram.max(axis=0)
if DEBUG:
tmp.append(sinogram)
return (max(max_values), theta[np.argmax(max_values)] * math.pi/180 - math.pi/2)
def make_ker(ker_len, ker_angle):
# h = ker_len * 2
h = ker_len
ker_len = ker_len // 2
ker = np.zeros((h, h), dtype='float')
k = -math.tan(ker_angle)
b = (1 - k) * ker_len
if abs(abs(ker_angle * 180/math.pi) - 90) > 10:
for x in range(h):
y = round(k * x + b)
y = int((y if y >= 0 else 0) if y <= h-1 else h-1)
if (y == 0 or y == h - 1):
continue
ker[y, x] = 1
else:
for y in range(h):
ker[y, ker_len] = 1
ret_value = ker/ker.sum()
if np.isnan(np.sum(ret_value)):
return []
else:
return ret_value
def get_common_ker_len_angle(kers):
max_shape = max([a[0] for a in kers])
lenghts = [a[0] for a in kers]
angles = [a[1] for a in kers]
return (int(np.mean(lenghts)), np.mean(angles))
class Cepstrum:
def __init__(self, picture, batch_size=64, step=0.25):
gc.enable()
self.batch_size = batch_size
self.step = step
self.x_batches = int(picture.shape[1] // (batch_size * step) - 1)
self.y_batches = int(picture.shape[0] // (batch_size * step) - 1)
self.picture = copy.deepcopy(picture)
self.squared_image = [0] * self.x_batches * self.y_batches
self.MainProcess()
plt.imsave('orig_img.png', self.picture, cmap='gray')
def get_square(self):
pixel_step = int(self.batch_size * self.step)
for y in range(self.y_batches):
for x in range(self.x_batches):
square = self.picture[y * pixel_step : y * pixel_step + self.batch_size,
x * pixel_step : x * pixel_step + self.batch_size]
self.squared_image[y * self.x_batches + x] = square
yield self.swap_quarters(Cepstrum.calculate_cepstrum(square))
def ft_array(self):
# CALCULATE CEPSTRUMS
t = time.time()
self.count_ft()
if (DEBUG):
print("Counted cepstrums: ", time.time() - t)
self.count_angles()
if (DEBUG):
print("Counted angles: ", time.time() - t)
self.count_lengths()
if (DEBUG):
print("Counted lengths: ", time.time() - t)
self.make_kernels()
if (DEBUG):
print("Counted kernels: ", time.time() - t)
self.weight = self.weight.reshape((self.y_batches, self.x_batches))
self.weight /= self.weight.max()
self.angle = self.angle.reshape((self.y_batches, self.x_batches))
self.blur_len = self.blur_len.reshape((self.y_batches, self.x_batches))
if (np.max(self.blur_len) == 0) :
self.angle_value = 0
print("Unable to calculate blur lengths")
return
# self.kernels = np.reshape(self.kernels, (self.y_batches, self.x_batches, 2)) #here
self.blur_len_value, self.angle_value = get_common_ker_len_angle(self.kernels)
self.kernel_image = make_ker(self.blur_len_value, self.angle_value)
self.squared_image = np.reshape(self.squared_image, (self.y_batches, self.x_batches, self.batch_size, self.batch_size))
# self.restore1()
# self.restore() #here
def MainProcess(self):
self.ft_array()
try:
temp2 =[ 0 ] * self.y_squares
for y in range(self.y_squares):
temp2[y] = np.hstack(self.restored_image[y, :, :, :])
self.restored_image_full = np.vstack(temp2)
except AttributeError as error:
return
def hamming(picture):
hm_len = picture.shape[0]
bw2d = np.outer(ss.hamming(hm_len), np.ones(hm_len))
bw2d = bw2d * bw2d.T
return picture * bw2d
def calculate_cepstrum(picture,threshold=0.5):
log = np.log(1 + np.abs(np.fft.fft2(Cepstrum.hamming(picture))))
fourier_abs = np.abs(np.fft.ifft2(log))
fourier_abs[fourier_abs >= threshold * fourier_abs.max()] = 0
fourier_abs[fourier_abs >= threshold * fourier_abs.max()] = 0
return fourier_abs
def swap_quarters(self, picture):
out_pict = copy.deepcopy(picture)
batch_size = picture.shape[0]
temp_pict = copy.deepcopy(out_pict[: batch_size//2, : batch_size//2])
out_pict[: batch_size//2, : batch_size//2] = out_pict[batch_size//2 :, batch_size//2 :]
out_pict[batch_size//2 :, batch_size//2 :] = temp_pict
temp_pict = copy.deepcopy(out_pict[: batch_size//2, batch_size//2 :])
out_pict[: batch_size//2, batch_size//2 :] = out_pict[batch_size//2 :, : batch_size//2]
out_pict[batch_size//2 :, : batch_size//2] = temp_pict[:]
return out_pict
def restore(self):
self.cut_image = []
pixel_step = self.batch_size
self.y_squares = int(self.picture.shape[0] // self.batch_size)
self.x_squares = int(self.picture.shape[1] // self.batch_size)
for y in range(self.y_squares):
for x in range(self.x_squares):
square = self.picture[y * pixel_step : y * pixel_step + self.batch_size,
x * pixel_step : x * pixel_step + self.batch_size]
self.cut_image.append(square)
self.cut_image = np.reshape(self.cut_image, (self.y_squares, self.x_squares, pixel_step, pixel_step))
self.restored_image = np.copy(self.cut_image)
ker_divider = int(1. / self.step)
self.new_kernels = [[0] * self.x_squares] * self.y_squares
def tf(y, x):
new_y = int((y if y >= 0 else 0) if y <= self.y_batches - 1 else self.y_batches - 1)
new_x = int((x if x >= 0 else 0) if x <= self.x_batches - 1 else self.x_batches - 1)
return (new_y, new_x)
for y_orig in range(self.y_squares):
for x_orig in range(self.x_squares):
k_l = []
for y in range(-ker_divider + 1, ker_divider):
for x in range(-ker_divider + 1, ker_divider):
k_l.append(self.kernels[tf(y_orig * ker_divider + y, x_orig * ker_divider + x)])
self.new_kernels[y_orig][x_orig] = make_ker(get_common_ker_len_angle(k_l))
print(y_orig, x_orig)
self.restored_image[y_orig, x_orig] =\
self.restore_function(self.cut_image[y_orig, x_orig], self.new_kernels[y_orig][x_orig])
return self.restored_image
def restore1(self):
self.deb_ker = make_common_ker(self.kernels)
plt.imsave(save_dir + filename[:-4] + '_ker_'+ str(c.batch_size) +'.png', self.deb_ker, cmap='gray')
self.restored_image_full = self.restore_function(self.picture, self.deb_ker)
def count_ft(self):
self.cepstrum_picture = np.array(list(self.get_square()))
self.conc_cepstrum_picture = self.cepstrum_picture.reshape((self.y_batches, self.x_batches, self.batch_size, self.batch_size))
temp = [ 0 ] * self.y_batches
for y in range(self.y_batches):
temp[y] = np.hstack(self.conc_cepstrum_picture[y, :, :, :])
self.conc_cepstrum_picture = np.vstack(temp)
plt.imsave('big_img.png', self.conc_cepstrum_picture, cmap='gray')
def count_angles(self):
self.weight = np.ndarray((self.y_batches * self.x_batches), dtype='float')
self.angle = np.ndarray((self.y_batches * self.x_batches), dtype='float')
if (DEBUG):
self.lines_img = np.copy(self.cepstrum_picture)
for idx, q in enumerate(self.cepstrum_picture):
self.weight[idx], self.angle[idx] = find_best_line(q)
def count_lengths(self):
self.blur_len = np.ndarray((self.y_batches * self.x_batches), dtype='int')
for idx, q in enumerate(self.cepstrum_picture):
if (DEBUG):
self.blur_len[idx], self.lines_img[idx] = get_blur_len(q, self.angle[idx], self.weight[idx])
self.conc_lines_img = self.lines_img.reshape((self.y_batches, self.x_batches, self.batch_size, self.batch_size))
temp = [ 0 ] * self.y_batches
for y in range(self.y_batches):
temp[y] = np.hstack(self.conc_lines_img[y, :, :, :])
self.conc_lines_img = np.vstack(temp)
plt.imsave('lines_img.png', self.conc_lines_img, cmap='gray')
else:
self.blur_len[idx] = get_blur_len(q, self.angle[idx], self.weight[idx])
def make_kernels(self):
self.kernels = [0] * self.y_batches * self.x_batches
for idx, q in enumerate(self.cepstrum_picture):
self.kernels[idx] = (self.blur_len[idx], self.angle[idx])
def restore_function(self, img, kernel):
# img /= img.max()
# if (np.shape(kernel)[0] == 0):
# return img
# self.z_0 = np.zeros(img.shape)
# return momentum_method(self.z_0, 0.85, kernel, img)
# return wiener_filter(img, kernel)
# betas = [0.01/math.sqrt(i) for i in range(1, 60)]
# self.minimis = FunctionalMinimisation(img, kernel, betas)
# self.minimis.MainProcess()
# return self.minimis._cur_image
pass
# +
def predict_directory(img_pattern, weights_path, out_dir):
# script_descriptor = open("predict.py")
# a_script = script_descriptor.read()
# sys.argv = ["predict.py",
# os.path.join(img_pattern,"*"),
# "--weights_path",
# weights_path,
# #"./pretrained_models/last_fpn_l7.h5",
# "--out_dir",
# out_dir]
# # ".\\MyStuff\\results\\dates\\05_02\\a1\\"]
os.system('python predict.py ' + os.path.join(img_pattern,"*") + " --weights_path" + weights_path + ' --out_dir' + out_dir)
# exec(a_script)
return
def make_directory(dirname):
if (not os.path.exists(dirname)):
os.mkdir(dirname)
return
# +
work_dir = ".\imgs\l_03\\"
save_dir = './result/'
img = rgb2gray(plt.imread(work_dir + 'img_25.png'))
plt.figure(figsize=(10,230))
plt.imshow(img, cmap='gray')
# +
# %%time
DEBUG = False
tmp = []
c = Cepstrum(img, batch_size=256, step=0.5)
# -
plt.imshow(c.kernel_image, cmap='gray')
get_common_ker_len_angle(c.kernels)
# +
dir_x = './pict/b_test/'
fnames_X = listdir(dir_x)
dir_y = './pict/s_test/'
fnames_y = listdir(dir_y)
X = list([dir_x + item for item in fnames_X])[:30]
y = list([dir_y + item for item in fnames_y])[:30]
save_dir_x = './pict/b_test_rotated/'
save_dir_y = './pict/s_test_rotated/'
# -
from scipy import ndimage
angles = []
common_dir = "./imgs/"
l_dirs = fnames_X = listdir(common_dir)
for l_dir in l_dirs:
cur_d = os.path.join(common_dir, l_dir)
fnames = listdir(cur_d)
for name in fnames:
img = rgb2gray(plt.imread(os.path.join(cur_d, name)))
c = Cepstrum(img, batch_size=256, step=0.5)
angles.append(get_common_ker_len_angle(c.kernels)[1])
# +
angles1 = list([a * 180/math.pi for a in angles])
angles2 = list([a + 180 if a < -90 else a for a in angles1])plsns_plot = sns.distplot(angles2, bins=12)
fig = sns_plot.get_figure()
plfig.savefig("error_angle_distribution.png")
# +
def nn_restore_image():
pass
for name in fnames:
img = rgb2gray(plt.imread(os.path.join(cur_d, name)))
c = Cepstrum(img, batch_size=256, step=0.5)
angles.append(get_common_ker_len_angle(c.kernels)[1])
# -
# orig_img_dir = 'MyStuff/results/dates/04_28/orig/'
orig_img_dir = 'MyStuff/results/dates/05_14/a00/l_07_a_00/'
fnames = listdir(orig_img_dir)
save_dir = '.\MyStuff\\results\dates\\05_14\\rotated'
# +
calc_angles = []
for name in fnames:
img = rgb2gray(plt.imread(os.path.join(cur_d, name)))
c = Cepstrum(img, batch_size=256, step=0.5)
angles.append(get_common_ker_len_angle(c.kernels)[1])
# -
angles = ['10', '20', '30', '40']
# +
# %time
angles = ['10', '20', '30', '40']
for a in angles:
local_save_dir = os.path.join(save_dir, a)
make_directory(local_save_dir)
a = int(a)
for idx, p in enumerate(fnames):
img = plt.imread(os.path.join(orig_img_dir, p))
rotated_img = ndimage.rotate(img, a)
edge = (rotated_img.shape[0] - 600) // 2 + 1
print(rotated_img.shape, edge)
# plt.imsave(local_save_dir + p, np.clip(rotated_img[edge:edge + 600, edge:edge + 600], 0., 1.))
plt.imsave(os.path.join(local_save_dir,p), np.clip(rotated_img, 0., 1.))
# +
restore_dirs = os.listdir(save_dir)
for d in restore_dirs:
predict_directory(os.path.join(save_dir, d),
"./pretrained_models/last_fpn_l7.h5",
'./MyStuff/results/dates/05_14/nn_restored/' + d)
# -
# %time
save_dir1 = '.\MyStuff\\results\dates\\05_14\\rotated_back'
for a in angles:
local_save_dir = os.path.join(save_dir1, a)
make_directory(local_save_dir)
a = int(a)
for idx, p in enumerate(fnames):
img = plt.imread(os.path.join(save_dir, 'a' + str(a), p))
rotated_img = ndimage.rotate(img, -a)
edge = (rotated_img.shape[0] - 600) // 2 + 1
print(rotated_img.shape, edge)
plt.imsave(os.path.join(local_save_dir,p), np.clip(rotated_img[edge:edge + 600, edge:edge + 600], 0., 1.))
# plt.imsave(os.path.join(local_save_dir,p), np.clip(rotated_img, 0., 1.))
save_dir
| rotate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
## Import libraries
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import hypergeom
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats import binom
import matplotlib.pyplot as plt
import math
## Read in self alignment files
bt_cutoff = 90
gdsmp = {}
with open('gdlist') as g:
lines = g.readlines()
for line in lines:
sp = line[:-1]
gdsmp[sp] = []
fn = 'self-align/' + sp + '.out'
with open(fn) as gg:
nelines = gg.readlines()
for nline in nelines:
tmp = nline.split('_')[2]
tmp1 = tmp.split('/')[0]
tmp2 = tmp.split(':')[1]
tmp3 = tmp2.split('%')[0]
perc = float(tmp3)
if perc > bt_cutoff:
gdsmp[sp].append(tmp1)
## Read species metadata
species_taxa={}
with open('unique_clusters.csv') as g:
lines = g.readlines()
for line in lines:
tmp = line.split(',')
species_taxa[tmp[0]] = tmp[11]
# +
## Plot HGT sequence prevalence in both species
fig, ax = plt.subplots(1, figsize=(10, 10))
Mixing_ratio = 0.2
tot_sag = 20000 ## Total SAG numbers
cutoff = 500 ## Threshold for considering a SAG covers the HGT sequence
ct1 = 0.05
pvals = []
sizes = []
nums= []
mkl = 0
colors={}
colors['A'] = '#FF99CC'
colors['B'] = '#99FFFF'
colors['F'] = '#CC99FF'
colors['P'] = '#FFCC99'
colors['FS'] = '#99CC99'
colors['C'] = 'yellow'
with open('list2.txt') as g:
lines = g.readlines()
for line in lines:
mkl=mkl+1
fn1 = line[:-1]
fn2 = 'data/'+fn1+'.out'
a1 = 0
a2 = 0
b1 = 0
b2 = 0
s1 = fn1.split('_')[0] # Species A (strain) bin ID
s2 = fn1.split('_')[1] # Species B (strain) bin ID
sp1 = s1.split('-')[0]
sp2 = s2.split('-')[0]
phy1 = species_taxa[sp1]
phy2 = species_taxa[sp2]
with open('data/'+fn1+'.len') as abc:
ABC = abc.readlines()
contig_length = float(ABC[0].split(' ')[0]) - len(fn1)-2
with open(fn2) as g2: # Read the intermediary results for HGT sequence alignment
newlines = g2.readlines()
for nl in newlines[:-1]:
tmp0 = nl.split('_')[0]
tmp1 = tmp0.split(' ')
tmp2 = nl.split('_')[1]
tmp3 = tmp2.split('/')[0] # SAG ID
t1 = tmp1[-2] # Number of nucleotide from the HGT that is covered by the SAG
t2 = tmp1[-1] # The species/strain-level bin ID that corresponds to the SAG
if t2 in gdsmp:
if tmp3 in gdsmp[t2]: # If the SAG pass the purity threshold
if t2 == s1:
a1 = a1+1 # Total SAG from species/strain A
if int(t1)>cutoff or int(t1)>ct1*contig_length:
a2 = a2+1 # Total SAG from species/strain A that covers the HGT sequence
elif t2 == s2:
b1 = b1+1 # Total SAG from species/strain B
if int(t1)>cutoff or int(t1)>ct1*contig_length:
b2 = b2+1 # Total SAG from species/strain B that covers the HGT sequence
if a1>0 and b1>0:
x1 = float(a2)/float(a1)
x2 = float(b2)/float(b1)
prob1 = binom.sf(a2,a1,0.2*b1/tot_sag)
prob2 = binom.sf(b2,b1,0.2*a1/tot_sag)
if prob1>1e-150:
pvals.append(0-math.log10(prob1))
else:
pvals.append(150)
if prob2>1e-150:
pvals.append(0-math.log10(prob2))
else:
pvals.append(150)
sizes.append(math.log10(contig_length))
nums.append(a1)
if s1!='26-2' and s2!='26-2':
if phy1 == phy2:
if phy1 == 'firmicutes':
cat = 'F'
elif phy1 == 'bacteroidetes':
cat = 'B'
else:
cat = 'C'
else:
cat = 'C'
if x1>0 and x2>0:
plt.scatter(x1,x2,s=contig_length/50, color=colors[cat], alpha=0.5,edgecolors='black')
if x1<0.1 or x2<0.1:
# plt.text(x1,x2,fn1)
mixed_1 = 0 #Mixing_ratio*ab2/float(tot)
mixed_2 = 0 #Mixing_ratio*ab1/float(tot)
x = np.linspace(0,1, 100)
y = np.linspace(mixed_1,mixed_1, 100)
line1 = plt.plot(x, y,color='black')
y = np.linspace(mixed_2,mixed_2, 100)
line1 = plt.plot(y, x,color='black')
# Create the figure
from matplotlib.patches import Circle
from matplotlib.lines import Line2D
# legend_elements = [Patch(facecolor=colors['B'], edgecolor=colors['B'],label='Bacteroidetes-Bacteroidetes'),
# Patch(facecolor=colors['F'], edgecolor=colors['F'],label='Firmicutes-Firmicutes'),
# Patch(facecolor=colors['C'], edgecolor=colors['C'],label='Cross-phyla')]
# plt.legend(handles=legend_elements, loc='right')
plt.xlabel('Proportion of SAGs from Species A that covers the HGT sequence',fontsize=15)
plt.ylabel('Proportion of SAGs from Species B that covers the HGT sequence',fontsize=15)
# +
# Generate figure legend
fig, ax = plt.subplots(1, figsize=(10, 10))
x = np.linspace(0,1, 100)
y = np.linspace(mixed_1,mixed_1, 100)
line1 = plt.plot(x, y,color='black')
line1 = plt.plot(y, x,color='black')
plt.scatter(0.5,0.2,s=5000/50, color='gray', alpha=0.5,edgecolors='black')
plt.scatter(0.5,0.3,s=20000/50, color='gray', alpha=0.5,edgecolors='black')
plt.scatter(0.5,0.4,s=80000/50, color='gray', alpha=0.5,edgecolors='black')
plt.scatter(0.5,0.8,s=20000/50, color=colors['B'], alpha=0.5,edgecolors='black')
plt.scatter(0.5,0.7,s=20000/50, color=colors['F'], alpha=0.5,edgecolors='black')
# +
## Plot pvalues for the subset
import matplotlib.pyplot as plt
import numpy as np
x = np.random.normal(170, 10, 250)
plt.hist(pvals,200)
plt.xlabel('-log10(pval)',fontsize=15)
plt.ylabel('Counts of HGT sequences',fontsize=15)
plt.xlim([0,20])
plt.ylim([0,5])
plt.plot([0-math.log10(8e-5),0-math.log10(8e-5)],[0,20])
plt.show()
print(mkl)
# +
## Plot all p-values
import matplotlib.pyplot as plt
import numpy as np
x = np.random.normal(170, 10, 250)
plt.hist(pvals,20)
plt.xlabel('-log10(pval)',fontsize=15)
plt.ylabel('Counts of HGT sequences',fontsize=15)
plt.show()
# -
| 08_HGT_QC/08_HGT_QC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## [教學目標]
# 學習使用 sklearn 中的 train_test_split 等套件,進行資料的切分
# ## [範例重點]
# 了解函數中各個參數的意義
# ## 引入我們需要的套件
from sklearn.model_selection import train_test_split, KFold
import numpy as np
# ## 用 numpy 生成隨機資料
X = np.arange(50).reshape(10, 5) # 生成從 0 到 50 的 array,並 reshape 成 (10, 5) 的 matrix
y = np.zeros(10) # 生成一個全零 arrary
y[:5] = 1 # 將一半的值改為 1
print("Shape of X: ", X.shape)
print("Shape of y: ", y.shape)
print('X: shape: ' + str(X.shape))
print(X)
print("")
print('y: shape: ' + str(y.shape))
print(y)
# ## 使用 train_test_split 函數進行切分
# 請參考 train_test_split 函數的[說明](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html),了解函數裡的參數意義
# - test_size 一定只能小於 1 嗎?
# - random_state 不設置會怎麼樣呢?
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
X_train
y_train
# ## 使用 K-fold Cross-validation 來切分資料
# 請參考 kf 函數的[說明](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html),了解參數中的意義。K 可根據資料大小自行決定,K=5 是蠻常用的大小
# - 如果使用 shuffle=True 會怎麼樣?
kf = KFold(n_splits=5)
i = 0
for train_index, test_index in kf.split(X):
i +=1
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
print("FOLD {}: ".format(i))
print("X_test: ", X_test)
print("Y_test: ", y_test)
print("-"*30)
# ## [作業重點]
# 觀察函數說明,要如何切出固定大小的測試集?
# ## 作業
# 假設我們資料中類別的數量並不均衡,在評估準確率時可能會有所偏頗,試著切分出 y_test 中,0 類別與 1 類別的數量是一樣的 (亦即 y_test 的類別是均衡的)
X = np.arange(1000).reshape(200, 5)
y = np.zeros(200)
y[:40] = 1
y
# 可以看見 y 類別中,有 160 個 類別 0,40 個 類別 1 ,請試著使用 train_test_split 函數,切分出 y_test 中能各有 10 筆類別 0 與 10 筆類別 1 。(HINT: 參考函數中的 test_size,可針對不同類別各自作切分後再合併)
| D34_訓練_測試集切分的概念/Day_034_train_test_split.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Decision Trees Exercises
#
# 
# # Learning Objectives
#
# - Recognize Decision trees and how to use them for classification problems
# - Recognize how to identify the best split and the factors for splitting
# - Explain strengths and weaknesses of decision trees
# - Explain how regression trees help with classifying continuous values
# - Describe motivation for choosing Random Forest Classifier over Decision Trees
# - Apply Intel® Extension for Scikit-learn* to leverage underlying compute capabilities of hardware for Random Forest Classifier
#
# # scikit-learn*
#
# Frameworks provide structure that Data Scientists use to build code. Frameworks are more than just libraries, because in addition to callable code, frameworks influence how code is written.
#
# A main virtue of using an optimized framework is that code runs faster. Code that runs faster is just generally more convenient but when we begin looking at applied data science and AI models, we can see more material benefits. Here you will see how optimization, particularly hyperparameter optimization can benefit more than just speed.
#
# These exercises will demonstrate how to apply **the Intel® Extension for Scikit-learn*,** a seamless way to speed up your Scikit-learn application. The acceleration is achieved through the use of the Intel® oneAPI Data Analytics Library (oneDAL). Patching is the term used to extend scikit-learn with Intel optimizations and makes it a well-suited machine learning framework for dealing with real-life problems.
#
# To get optimized versions of many Scikit-learn algorithms using a patch() approach consisting of adding these lines of code PRIOR to importing sklearn:
#
# - **from sklearnex import patch_sklearn**
# - **patch_sklearn()**
#
# ## This exercise relies on installation of Intel® Extension for Scikit-learn*
#
# If you have not already done so, follow the instructions from Week 1 for instructions
# ## Introduction
#
# We will be using the wine quality data set for these exercises. This data set contains various chemical properties of wine, such as acidity, sugar, pH, and alcohol. It also contains a quality metric (3-9, with highest being better) and a color (red or white). The name of the file is `Wine_Quality_Data.csv`.
# +
from __future__ import print_function
import os
data_path = ['data']
# This listener will import Intel Extnsions for Scikit-learn optimized versions
# for any applicable imports from scikit-learn once this patch has been run
from sklearnex import patch_sklearn
patch_sklearn()
from io import StringIO
from IPython.display import Image, display
from sklearn.tree import export_graphviz
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.metrics import mean_squared_error
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
from sklearn.tree import DecisionTreeClassifier
# -
# ## Question 1
#
# * Import the data and examine the features.
# * We will be using all of them to predict `color` (white or red), but the colors feature will need to be integer encoded.
# ## Question 2
#
# * Use `StratifiedShuffleSplit` to split data into train and test sets that are stratified by wine quality. If possible, preserve the indices of the split for question 5 below.
# * Check the percent composition of each quality level for both the train and test data sets.
# ## Question 3
#
# * Fit a decision tree classifier with no set limits on maximum depth, features, or leaves.
# * Determine how many nodes are present and what the depth of this (very large) tree is.
# * Using this tree, measure the prediction error in the train and test data sets. What do you think is going on here based on the differences in prediction error?
# ## Question 4
#
# * Replace Decision Tree in the cell above with Random Forest algorithm
# * from sklearn.ensemble import RandomForestClassifier
# * patch sklearn to apply fast version from Intel Extensions for Sckit-learn*
# * Instantiate RandomForestClassifier(random_state=42)
# * Examine feature importance: dict(zip(X_train.columns, dt.feature_importances_))
# * Compare Metrics of Decision Tree to Random Forest
# +
# -
# ## Question 5
#
# * Using grid search with cross validation, find a decision tree that performs well on the test data set. Use a different variable name for this decision tree model than in question 3 so that both can be used in question 6.
# * Determine the number of nodes and the depth of this tree.
# * Measure the errors on the training and test sets as before and compare them to those from the tree in question 3.
# ## Question 6
#
# * Re-split the data into `X` and `y` parts, this time with `residual_sugar` being the predicted (`y`) data. *Note:* if the indices were preserved from the `StratifiedShuffleSplit` output in question 2, they can be used again to split the data.
# * Using grid search with cross validation, find a decision tree **regression** model that performs well on the test data set.
# * Measure the errors on the training and test sets using mean squared error.
# * Make a plot of actual *vs* predicted residual sugar.
# ## Question 7 *(Optional)*
#
# This question is optional as it requires an additional command line program (GraphViz) and Python library (PyDotPlus). GraphViz can be installed with a package manager on Linux and Mac. For PyDotPlus, either `pip` or `conda` (`conda install -c conda-forge pydotplus`) can be used to install the library.
#
# Once these programs are installed:
#
# * Create a visualization of the decision tree from question 3, where wine color was predicted and the number of features and/or splits are not limited.
# * Create a visualization of the decision tree from question 4, where wine color was predicted but a grid search was used to find the optimal depth and number of features.
#
# The decision tree from question 5 will likely have too many nodes to visualize.
| AI-and-Analytics/Jupyter/Introduction_to_Machine_Learning/07-Decision_Trees/Decision_Trees_Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
# values of x and y axes
x = [5, 10, 15, 20, 25, 30, 35, 40, 45, 50,55,56]
y = [1, 4, 3, 2, 7, 6, 9, 8, 10, 5,11,13]
plt.plot(x, y, 'b')
plt.xlabel('x')
plt.ylabel('y')
# 0 is the initial value, 51 is the final value
# (last value is not taken) and 5 is the difference
# of values between two consecutive ticks
plt.xticks(np.arange(0, 56, 5))
plt.yticks(np.arange(0, 12, 1))
plt.show()
# +
#subplots graph
import matplotlib.pyplot as plt
x=[1,2,3,4,5,6]
y=[10,30,40,55,67,80]
z=[0,2,4,6,8,10]
figure,axis=plt.subplots(1,4)
axis[0].set_xlabel("banglore")
axis[0].set_ylabel("temperature")
axis[0].plot(x, y, 'g--o')
axis[2].plot(x, y, 'y--o')
axis[3].plot(x, z, 'b--o')
axis[1].plot(y,x, 'r--o')
axis[1].set_xlabel("jaipur")
axis[1].set_ylabel("temperature")
figure.tight_layout()
# +
import matplotlib.pyplot as plt
import numpy as np
clark=np.array([1,2,3,4,5,6,7,8,9,10])
bellamy=np.array([5,10,15,20,25,30,35,40,45,50])
plt.scatter(clark,bellamy)
plt.plot([clark.mean()]*len(clark),bellamy,'r')
plt.plot(clark,[bellamy.mean()]*len(bellamy),'y')
for var in range(len(clark)):
plt.plot([clark[var],clark.mean()],[bellamy[var],bellamy[var]],'r:')
plt.plot([clark[var],clark[var]],[bellamy[var],bellamy.mean()],'b:')
# +
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from matplotlib import style
style.use('fivethirtyeight')
fig = plt.figure()
ax1 = fig.add_subplot(1,1,1)
def animate(i):
graph_data = open('example.txt','r').read()
print(graph_data)
lines = graph_data.split('\n')
xs =[]
ys = []
for line in lines:
if len(line) > 1:
x, y = line.split(',')
xs.append(float(x))
ys.append(float(y))
ax1.clear()
ax1.plot(xs, ys)
ani = animation.FuncAnimation(fig, animate, interval=1000)
plot.show()
# +
import datetime as dt
import matplotlib.pyplot as plt
import matplotlib.animation as animation
import tmp102
# Create figure for plotting
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
xs = []
ys = []
# Initialize communication with TMP102
tmp102.init()
# This function is called periodically from FuncAnimation
def animate(i, xs, ys):
# Read temperature (Celsius) from TMP102
temp_c = round(tmp102.read_temp(), 2)
# Add x and y to lists
xs.append(dt.datetime.now().strftime('%H:%M:%S.%f'))
ys.append(temp_c)
# Limit x and y lists to 20 items
xs = xs[-20:]
ys = ys[-20:]
# Draw x and y lists
ax.clear()
ax.plot(xs, ys)
# Format plot
plt.xticks(rotation=45, ha='right')
plt.subplots_adjust(bottom=0.30)
plt.title('TMP102 Temperature over Time')
plt.ylabel('Temperature (deg C)')
# Set up plot to call animate() function periodically
ani = animation.FuncAnimation(fig, animate, fargs=(xs, ys), interval=1000)
plt.show()
# +
import matplotlib.pyplot as plt
# draw the figure so the animations will work
fig = plt.gcf()
fig.show()
fig.canvas.draw()
while True:
# compute something
plt.plot([1], [2]) # plot something
# update canvas immediately
plt.xlim([0, 100])
plt.ylim([0, 100])
#plt.pause(0.01) # I ain't needed!!!
fig.canvas.draw()
# -
import matplotlib.pyplot as plt
import numpy as np
x=[1,2,3,5,7,8,9]
y=[1,2,3,5,6,7,8]
plt.figure(figsize=(10,12),facecolor='blue')
plt.plot(x,y)
plt.plot(y,x,'g')
plt.scatter(x,y)
plt.show()
import matplotlib.pyplot as plt
cap=plt.figure(figsize=(10,3))
#add_axes
nebula=cap.add_axes([0,0,.4,.5])
iron_man=cap.add_axes([.6,.0,.5,.5])
nebula.plot(x,y)
nebula.scatter(x,y)
#iron_man.plot(x,np.array(x)**2)
iron_man.plot(x,y)
# +
x = np.arange(0,4*np.pi,0.1)
y = np.sin(x)
z=np.cos(x)
plt.plot(x,y,x,z)
plt.show()
#another method
degree=np.arange(0,361,15)
radian=np.deg2rad(degree)
sine=np.sin(radian)
cose=np.cos(radian)
plt.plot(degree,sine)
plt.plot(degree,cose)
plt.scatter(radian,sine)
plt.scatter(radian,cose)
plt.show()
# -
#plot using radian only
plt.plot(radian)
plt.plot(radian**2)
plt.plot(radian**4)
# +
import matplotlib.pyplot as plt
import numpy as np
thor=plt.figure()
G=thor.add_axes([0,0,1,1])
y=thor.add_axes([.4,.7,.63,.64])
G.plot(radian,radian,'r--o',label='1D')
G.plot(radian,radian**2,'g--o',label='2D')
G.plot(radian,radian**3,'b--o',label='3D')
G.set_title("x x_square x_cube")
G.legend(loc=7)
G.set_xlabel('ridddhiiii')
G.set_ylabel('rupal')
G.grid()
#for second plot
y.plot(degree,sine,'r',label='sine graph')
y.plot(degree,cose,'y',label='cose graph')
y.set_title("sine and cose graph with degree")
y.legend()
y.set_xlabel('degree')
y.set_ylabel('radian')
#y.grid()
y.set_xticks(np.arange(0, 361, 30))
y.set_yticks(np.arange(0, 1, .2))
# +
import matplotlib.pyplot as plt
import numpy as np
#normal(mean,std,length)
x=np.random.normal(20,1.5,1000)
#plt.hist(x,bins=30)
#randint(range of x,,range of y under it lies)
y=np.random.randint(10,100,20)
plt.hist(y,bins=10)
plt.show()
# -
avengers=[100,200,300,400,550]
#Labels
powerof=['antman','spiderman','hawkeye','ironman','thor']
color=['r','pink','purple','g','yellow']
plt.pie(avengers,labels=powerof,colors=color,autopct='%.2s',explode=[0,0,0,0,.5]) #s for integer values
plt.title('End Game')
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
rafale=np.linspace(-5,10,1000)
r_sq=rafale**2
r_cube=rafale**3
plt.plot(rafale,rafale)
plt.plot(rafale,r_sq,'r')
plt.plot(rafale,r_cube,'g')
plt.show()
# -
avengers=[100,202,300,400,550]
#Labels
powerof=['antman','spiderman','hawkeye','ironman','thor']
#plt.bar(powerof,avengers)
plt.barh(powerof,avengers)
plt.show()
plt.boxplot([avengers])
plt.show()
# +
plt.scatter(np.arange(10),np.random.rand(10))
plt.show()
# -
import seaborn as sns
plt.style.use('seaborn')
plt.scatter(np.arange(10),np.random.rand(10))
plt.show()
year=[2,3,4,5,6]
growth=[20,30,40,55,79]
color=['blue','green','red','yellow','black']
plt.bar(year,height=growth,color=color)
yacht=np.random.normal(50,5,1000)
prince=plt.figure(figsize=(10,5))
ax=prince.add_axes([1,1,1,1])
sns.distplot(yacht,ax=ax)
plt.show()
library(help='datasets')
# +
from pydataset import data
titanic = data('titanic')
titanic.head(20)
#titanic.describe()
# -
| matplotlib 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:riboraptor]
# language: python
# name: conda-env-riboraptor-py
# ---
# %pylab inline
import pandas as pd
# +
import numpy as np
import matplotlib.gridspec as gridspec
import matplotlib as mpl
import matplotlib.pyplot as plt
import scipy
import seaborn as sns
import statsmodels.api as sm
from collections import OrderedDict
import sys
import time
import glob
from riboraptor.helpers import path_leaf
import pybedtools
from scipy.stats import poisson
from scipy.signal import savgol_filter
cds_bed = pybedtools.BedTool('/home/cmb-panasas2/skchoudh/github_projects/gencode_regions/data/GRCh38/v25/cds.bed.gz').to_dataframe()
PAPER_PRESET = {"style": "ticks", "font": "Arial", "context": "paper",
"rc": {"font.size":20, "axes.titlesize":22,
"axes.labelsize":28, 'axes.linewidth': 2,
"legend.fontsize":20, "xtick.labelsize": 28,
"ytick.labelsize":20, "xtick.major.size": 8.0,
"ytick.major.size": 8.0, "axes.edgecolor": "black",
"xtick.major.pad": 3.0, "ytick.major.pad": 3.0}}
PAPER_FONTSIZE = 20
sns.set(**PAPER_PRESET)
fontsize = PAPER_FONTSIZE
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
# -
cds_bed.head()
# +
def get_shifted_gene_profile(gene_profile, gene_offset_5p, metagene_offset, atg_location):
# How many upstream 5'UTr basese were counted?
assert gene_offset_5p >= 0
# Say -12 is the metagene offset
assert metagene_offset <= 0
total_length = len(gene_profile)
gene_profile = pd.Series(gene_profile, index=np.arange(-gene_offset_5p, len(gene_profile)-gene_offset_5p))
# Shoft -12 to zero by adding the - of metagene_offset
gene_profile = gene_profile.rename(lambda x: x-metagene_offset)
# For genes whivh are on negatve strand because of 1-offset
# we end up counting one extra base.
# So ATG does not start at 0-1-2
# but at 1-2-3
# So we need to again do offset
assert atg_location in [0, 1, 2]
shifted_gene_profile = gene_profile.rename(lambda x: x-atg_location)
return shifted_gene_profile
def avg_profiles(samples, gene_name):
samplewise_profiles = []
for sample in samples:
print(sample)
gene_coverage_tsv = '/staging/as/skchoudh/re-ribo-analysis/hg38/SRP098789/gene_coverages/{}_gene_coverages.tsv.gz'.format(sample)
gene_coverage = pd.read_csv(gene_coverage_tsv, compression='gzip', sep='\t').set_index('gene_name').loc[gene_name]
gene_counts = eval(gene_coverage['coverage'])
#gene_mean = eval(gene_coverage['mean'])
metagene_offset = -12#metagene_offsets[sample]
atg_location = 0#codon_map.loc[gene_name]['ATG_first_location']
#fasta_length = codon_map.loc[gene_name]['fasta_length'] - atg_location
gene_offset_5p = gene_coverage['offset_5p']
gene_profile = get_shifted_gene_profile(gene_counts, gene_offset_5p, metagene_offset, atg_location)
gene_profile = gene_profile#.get(np.arange(0, fasta_length))
gene_profile_normalized = gene_profile/gene_profile.mean(skipna=True)
samplewise_profiles.append(gene_profile_normalized)
samplewise_profiles = np.array(samplewise_profiles)
return np.nanmean(samplewise_profiles, axis=0)
def sum_profiles(samples, gene_name):
samplewise_profiles = []
for sample in samples:
print(sample)
gene_coverage_tsv = '/staging/as/skchoudh/re-ribo-analysis/hg38/SRP098789/gene_coverages/{}_gene_coverages.tsv.gz'.format(sample)
gene_coverage = pd.read_csv(gene_coverage_tsv, compression='gzip', sep='\t').set_index('gene_name').loc[gene_name]
gene_counts = eval(gene_coverage['coverage'])
#gene_mean = eval(gene_coverage['mean'])
metagene_offset = -12#metagene_offsets[sample]
atg_location = 0#codon_map.loc[gene_name]['ATG_first_location']
#fasta_length = codon_map.loc[gene_name]['fasta_length'] - atg_location
gene_offset_5p = gene_coverage['offset_5p']
gene_profile = get_shifted_gene_profile(gene_counts, gene_offset_5p, metagene_offset, atg_location)
samplewise_profiles.append(gene_profile)
samplewise_profiles = np.array(samplewise_profiles)
return np.nansum(samplewise_profiles, axis=0)
def collpase_gene_coverage_to_codon(gene_profile):
"""Assume the gene 0 is the true zero and exclude
trailing positions which are not 0 mod 3
"""
codon_profile = []
for i in range(0, len(gene_profile)-3, 3):
codon_profile.append(np.nansum(gene_profile[np.arange(i, i+3)]))
return pd.Series(codon_profile, index=np.arange(1, len(codon_profile)+1))
def g_transform(data):
data = np.array(data)
return np.log(np.log(data+1)+1)
def inverse_g_transform(y):
y = np.array(y)
return np.exp(np.exp(y)-1)-1
def baseline_correct(y, n_iterations=100):
z = g_transform(y)
z_copy = np.empty(len(z))
n = len(z)
for i in range(0, n_iterations):
for j in range(i, n-i):
mean_z = 0.5*(z[j-i]+[j+i])
mean_z = min(mean_z, z[j])
z_copy[j] = mean_z
for k in range(i, n-i):
z[k] = z_copy[k]
inv_z = inverse_g_transform(z)
return inv_z
def med_abs_dev(data):
"""Calculate Median absolute deviation
"""
return 1.4826*max(np.nanmedian(np.abs(data-np.nanmedian(data))), 1e-4)
def calculate_peaks(data, order=3, snr=2.5):
""" Calculate Peaks
"""
if isinstance(data, pd.Series):
index = data.index
else:
index = np.arange(0, len(data))
data = np.array(data)
data_rel_max_idx = signal.argrelmax(data, axis=0, order=order)[0]
noise = med_abs_dev(data)
#peaks_height = np.zeros(len(data))
peaks_idx = [x for x in data_rel_max_idx if data[x] > snr*noise]
peaks_x = index[peaks_idx]
#for x in peaks_idx:
# peaks_height[x] = data[x]
peaks_height = data[peaks_idx]
return peaks_x, peaks_height
def calculate_snr(data):
data = np.array(data)
sigma = med_abs_dev(data)
return data/sigma
def baseline_correct(y, n_iterations=100):
z = g_transform(y)
z_copy = np.empty(len(z))
n = len(z)
for i in np.arange(n_iterations, 0, -1):
for j in np.arange(i, n-i):
mean_z = 0.5*(z[j-i]+z[j+i])
mean_z = min(mean_z, z[j])
z_copy[j] = mean_z
for k in np.arange(i, n-i):
z[k] = z_copy[k]
inv_z = inverse_g_transform(z)
return inv_z
def remove_baseline(y, baseline):
return y-baseline
def Z_score(data):
return (data-np.nanmean(data))/np.std(data)
def get_poisson_lambda(data, method='ncg'):
res = sm.Poisson(data, np.ones_like(data)).fit(
disp=0, method=method)
l = res.predict()[0]
return l
def get_blockwise_poisson_lambdas(data, blocksize=3, method='ncg'):
data = list(data)
pointer = 0
index = 0
lambdas = []
gene_length = len(data)
while pointer != gene_length:
if (index + 1) * blocksize >= gene_length - 1:
end = gene_length - 1
pointer = gene_length
else:
end = (index + 1) * blocksize
start = index * blocksize
block_coverage = data[start:end]
lambdas.append(get_poisson_lambda(block_coverage, method))
index+=1
return lambdas
def load_gene_fasta(gene_name):
fasta_f = '/home/cmb-panasas2/skchoudh/genomes/hg38/cds_fasta/cds_gene_{}.fasta'.format(gene_name)
fasta = SeqIO.parse(open(fasta_f), 'fasta')
for record in fasta:
return str(record.seq)
def mark_extreme_points_poisson(data, lambdas, blocksize):
data = list(data)
pvalues = []
"""
for i in range(0, len(lambdas)):
for point in data[i:i+blocksize]:
pvalue = poisson.sf(point, lambdas[i])#1-poisson.cdf(point, lambdas[i])
pvalues.append(pvalue)
"""
data = list(data)
pointer = 0
index = 0
gene_length = len(data)
while pointer != gene_length:
if (index + 1) * blocksize >= gene_length - 1:
end = gene_length - 1
pointer = gene_length
else:
end = (index + 1) * blocksize
start = index * blocksize
for point in data[start:end]:
pvalue = poisson.sf(point, lambdas[index])#1-poisson.cdf(point, lambdas[i])
pvalues.append(pvalue)
index+=1
return pvalues
def get_position_wise_cdf(values):
values = list(values)
total = np.nansum(values)
cdf = np.cumsum(values)/total
return cdf
def get_poisson_difference(data, lambdas, blocksize):
data = list(data)
differences = []
j = 0
"""
for i in range(0, len(data)):
difference = data[i] - lambdas[i//blocksize]
differences.append(difference)
for i in range(0, len(lambdas)):
for point in data[i:i+blocksize]:
difference = point - lambdas[i]
differences.append(difference)
"""
data = list(data)
pointer = 0
index = 0
gene_length = len(data)
while pointer != gene_length:
if (index + 1) * blocksize >= gene_length - 1:
end = gene_length - 1
pointer = gene_length
else:
end = (index + 1) * blocksize
start = index * blocksize
for point in data[start:end]:
difference = point - lambdas[index]
differences.append(difference)
index+=1
return differences
def expand_poisson_lambda(lambdas, blocksize=3):
lambdas = list(lambdas)
expanded = []
for i in range(0, len(lambdas)):
lambda_points = [lambdas[i]]*blocksize
expanded += lambda_points
return expanded
def gaussian_pvalue(values):
values = np.array(values)
mean = np.nanmean(values)
std = np.std(values)
zscore = (values - mean)/std
pvalues = stats.norm.sf(zscore)
return pvalues
# -
def read_ribotrcer_samples(root_path):
ribotricer_output = OrderedDict()
for filepath in glob.glob('{}/*_translating_ORFs.tsv'.format(root_path)):
ribo_sample = path_leaf(filepath).replace('_translating_ORFs.tsv', '')
#filepath = os.path.join(root_path, ribo_sample+'_translating_ORFs.tsv')
ribotricer_output[ribo_sample] = pd.read_csv(filepath, sep='\t')
return ribotricer_output
samples_srx = OrderedDict([('1.5mu',
OrderedDict([('10min_1',
['SRX2536403', 'SRX2536404', 'SRX2536405']),
('60min_1',
['SRX2536412', 'SRX2536413', 'SRX2536414']),
('60min_2', ['SRX2536421', 'SRX2536423'])])),
('0.3mu',
OrderedDict([('10min_1',
['SRX2536406', 'SRX2536407', 'SRX2536408']),
('60min_1',
['SRX2536415', 'SRX2536416', 'SRX2536417'])])),
('vehicle',
OrderedDict([('10min_1',
['SRX2536409', 'SRX2536410', 'SRX2536411']),
('60min_1',
['SRX2536418', 'SRX2536419', 'SRX2536420']),
('60min_2', ['SRX2536422', 'SRX2536424'])]))])
gene_coverage_tsv = '/staging/as/skchoudh/re-ribo-analysis/hg38/SRP098789/gene_coverages/SRX2536406_gene_coverages.tsv.gz'.format(sample)
gene_coverage = pd.read_csv(gene_coverage_tsv, compression='gzip', sep='\t')
gene_coverage.head()
gene_name = 'ENSG00000169174'
mu15_10min_samples = samples_srx['1.5mu']['10min_1']
#ribotricer_output = read_ribotrcer_samples('/staging/as')
mu15_10min_avg = avg_profiles(mu15_10min_samples, gene_name)
mu15_10min_sum = sum_profiles(mu15_10min_samples, gene_name)
#sns.set_context('talk', font_scale=3)
sns.set_style('whitegrid')
fig, ax = plt.subplots(figsize=(10, 7))
ax.plot(collpase_gene_coverage_to_codon(mu15_10min_avg), color='#08519c', linewidth=1.5)
ax.set_title('PCSK9')
#ax.set_ylabel('Normalized RPF count')
#fig.tight_layout()
fig.savefig("plots_savgol/savgol_pcsk9_10min_Avg_profile.pdf", dpi="figure", bbox_inches="tight")
# +
PAPER_PRESET = {"style": "ticks", "font": "Arial", "context": "paper",
"rc": {"font.size":20, "axes.titlesize":22,
"axes.labelsize":28, 'axes.linewidth': 2,
"legend.fontsize":20, "xtick.labelsize": 28,
"ytick.labelsize":20, "xtick.major.size": 8.0,
"ytick.major.size": 8.0, "axes.edgecolor": "black",
"xtick.major.pad": 3.0, "ytick.major.pad": 3.0}}
PAPER_FONTSIZE = 20
sns.set(**PAPER_PRESET)
fontsize = PAPER_FONTSIZE
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
# -
from scipy import signal
from scipy import stats
import statsmodels.api as sm
sns.set_style('whitegrid')
# +
fig = plt.figure(figsize=(10, 7))
data = collpase_gene_coverage_to_codon(mu15_10min_sum)
ax1 = plt.subplot(211)
ax1.plot(data,color='#08519c', linewidth=1.5)
ax1.set_title('PCSK9 | blocksize = 30')
ax1.set_ylabel('RPF count')
lambdas = get_blockwise_poisson_lambdas(data, blocksize=30)
pvalues = mark_extreme_points_poisson(data, lambdas, blocksize=30)
ax2 = plt.subplot(212, sharex=ax1, sharey=ax1)
ax2.plot(expand_poisson_lambda(lambdas, blocksize=30),color='#08519c', linewidth=1.5)
ax2.set_ylabel('lambda')
fig.savefig('plots_savgol/pcsk9_10min_sum_gene_profile_poisson_blocksize30_top.pdf', dpi="figure", bbox_inches="tight")
fig = plt.figure(figsize=(10, 7))
ax3 = plt.subplot(211, sharex=ax1, sharey=ax1)
ax3.set_title('PCSK9 | blocksize = 30')
ax3.plot(get_poisson_difference(data, lambdas, 30),color='#08519c', linewidth=1.5)
ax3.set_ylabel('difference')
ax4 = plt.subplot(212, sharex=ax1)
ax4.plot(-np.log10(pvalues),color='#08519c', linewidth= 1.5)
ax4.axhline(y=2, color="#D55E00", linestyle='dashed', linewidth=3)
ax4.set_ylabel('-log(pval)')
#ax4.set_title('Poission mean | blocksize = 9')
#fig.tight_layout()
fig.savefig('plots_savgol/pcsk9_10min_sum_gene_profile_poisson_blocksize30_bottom.pdf', dpi="figure", bbox_inches="tight")
# +
fig = plt.figure(figsize=(10, 7))
data = collpase_gene_coverage_to_codon(mu15_10min_sum)
ax1 = plt.subplot(211)
ax1.plot(data,color='#08519c', linewidth=1.5)
ax1.set_title('PCSK9')
ax1.set_ylabel('RPF count')
filtered = savgol_filter(data, 15, 3)
ax2 = plt.subplot(212, sharex=ax1, sharey=ax1)
ax2.plot(filtered,color='#08519c', linewidth=1.5)
ax2.set_ylabel('RPF count')
fig.savefig('plots_savgol/pcsk9_10min_sum_gene_profile_zscore_golay_top.pdf', dpi="figure", bbox_inches="tight")
fig = plt.figure(figsize=(10, 7))
ax3 = plt.subplot(211, sharex=ax1)
peaks_x, peaks_height = calculate_peaks(filtered, order=9, snr=3)
ax3.set_title('PCSK9')
ax3.set_ylabel('SNR')
markerline, stemlines, baseline = ax3.stem(peaks_x, peaks_height)#, linewidth=1.5)
markerline.set_markerfacecolor('#08519c')
ax4 = plt.subplot(212, sharex=ax1)
pvalues = -np.log10(gaussian_pvalue(filtered))
ax4.plot(pvalues,color='#08519c', linewidth=1.5)
ax4.set_ylabel('-log(pval)')
ax4.axhline(y=2, color="#D55E00", linestyle='dashed', linewidth=3)
fig.savefig('plots_savgol//pcsk9_10min_sum_gene_profile_zscore_golay_bottom.pdf', dpi="figure", bbox_inches="tight")
# -
significant_peaks_lambdawise = []
blocksizes = []
for i in range(3, 151, 3):
lambdas = get_blockwise_poisson_lambdas(data, blocksize=i)
pvalues = mark_extreme_points_poisson(data, lambdas, blocksize=i)
pvalues = -np.log10(pvalues)
filtered_pvals = list(filter(lambda x: x>=2, pvalues))
blocksizes.append(i)
significant_peaks_lambdawise.append(len(filtered_pvals))
fig, ax = plt.subplots(figsize=(5, 5))
ax.plot(blocksizes, significant_peaks_lambdawise, marker='o',color='#08519c', linewidth=1.5)
ax.set_ylabel('# Significant Peaks')
ax.set_xlabel('Blocksize')
fig.tight_layout()
fig.savefig('plots_savgol//pcsk9_blocksizes_vs_peaks_poisson.pdf', dpi="figure", bbox_inches="tight")
ribotricer_output = read_ribotrcer_samples('/staging/as/skchoudh/re-ribo-analysis/hg38/SRP098789/ribotricer_results/')
ribotricer_output.keys()
mu15_10min_samples = samples_srx['1.5mu']['10min_1']
mu15_10min_samples
df = ribotricer_output['SRX2536426']
df_subset = df.query("gene_id == 'ENSG00000169174' & ORF_type=='annotated'")
df_subset
# # Poisson blocksize
# +
fig, ax = plt.subplots(figsize=(5,5))
sns.kdeplot(Z_score(collpase_gene_coverage_to_codon(mu15_10min_sum)), color='black', bw=.25, ax=ax, linewidth=3)
ax.set_label('Density')
ax.set_xlabel('Z score')
fig.tight_layout()
fig.savefig('plots_savgol//pcsk9_10min_sum_gene_profile_zscore.pdf', dpi="figure", bbox_inches="tight")
# +
fig, ax = plt.subplots(figsize=(8,8))
data = collpase_gene_coverage_to_codon(mu15_10min_sum)
sns.kdeplot(Z_score(savgol_filter(data, 15, 3)), color='black', bw=.25, ax=ax, linewidth=3)
ax.set_label('Density')
ax.set_xlabel('Z score')
fig.tight_layout()
fig.savefig('plots_savgol/pcsk9_10min_sum_gene_profile_zscore_savgol.pdf', dpi="figure", bbox_inches="tight")
# -
| notebooks/PCSK9-SRP098789.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.datasets import make_blobs
from sklearn.cluster import KMeans as SkKMeans
from dojo.cluster import KMeans
# -
K = 5
X, y = make_blobs(n_samples=500, n_features=2, centers=K, cluster_std=0.6)
def plot(y, centroids=None):
for cl, color in zip(range(K), ['k', 'blue', 'red', 'green', 'purple']):
plt.scatter(X[np.array(y) == cl, 0], X[np.array(y) == cl, 1], c=color, marker='^')
if centroids is not None:
centroids = np.array(centroids)
plt.scatter(centroids[:, 0], centroids[:, 1], c="yellow", marker='o', s=100)
plt.grid()
plt.show()
plot(y)
kmeans = KMeans(n_clusters=K).fit(X)
sk_kmeans = SkKMeans(n_clusters=K).fit(X)
# ## Dojo's KMeans
plot(kmeans.clusters, centroids=kmeans.centroids)
kmeans.distortion
# ## Sckit-learn's KMeans
plot(sk_kmeans.labels_, centroids=sk_kmeans.cluster_centers_)
sk_kmeans.inertia_
| examples/KMeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Db2 on Cloud Registration
# If you would like to try running the Db2 sample applications against a Db2 on Cloud database, then you will need to follow these steps to register for a Db2 on Cloud account.
# ## IBM Cloud Registration
# To get started you must have a userid and password on the IBM system. You may already have one based on previous product trails or downloads you may have registered for. To get access to the IBM cloud media, click on the following link: https://www.ibm.com/cloud/
#
# This will direct your browser to the main IBM Cloud page. **Note:** When you click on the above link it will open up another tab in the browser you are using.
#
# You should see a web page similar to the following (**Note:** Web pages have a tendency to change over time so these might be different than what you see.)
# 
# On the left hand side (and on the top right corner) of the welcome screen you will see the **Sign Up** link. This same link is used if you already have a user id, so start by clicking on that link. You will be directed to the account sign up / login window.
#
# 
# If you already have an account set up, click on the **Log in** link at the top. However, if you don't have an account, fill in the details on the right and Create a new account. The set up process should only take a few minutes. Once you do have an account, you will sign into the main IBM Cloud console. The screen may have various sections on it, but look for the menu item at the top that says **Cloud**. Once you click on the catalog you should see the following screen.
#
# 
# On the left hand side of the catalog is a list of categories of services that are available.
#
# 
# Selecting **Databases** will display the **Db2 on Cloud** plan.
#
# 
# When you select Db2 from the list, the following detailed description of the service will be displayed.
#
# 
# There are a number of fields that need to be filled in.
#
# * Region - What region the service should run in
# * Pricing Plan - Choose the plan that you want to use (Lite is highlighted)
# * Service Details (at the bottom of the web page)
# ### Region
# The **Region** field is particulary important because it will determine the type of performance that you will get from the database. The closer the region is to your location, the less network latency there is. The current regions that are available are shown in the image below.
#
# 
#
# Once you have determined the region you want to run in, select the type of plan that you want to use.
# ### Plan
# Since we are only using the Db2 on Cloud plan for demonstration purposes, select the **Lite** plan.
#
# 
#
# This plan is restricted to:
#
# * 200 MB of user data
# * 5 simultaneous users
# * 30 days of service (which can be extended monthly)
# * Shared resources and only one schema
#
# The next step is to scroll to the bottom of the screen and update the resource information.
# ### Resources
#
# 
# In the resources section, you should provide a name for your service (a default value is provided), add any tags which identify the service that you are using (for searching) and finally an email address that can be used to reach you in the event there are issues with the service, or to notify you when the plan needs to be updated.
#
# When you are done updating the fields, you will need to confirm your options by clicking on the create button on the right hand side of the screen.
# 
#
# This will create the Db2 service for you and then let you get started with the labs! You should see a screen similar to the following displayed when the service has been created.
#
# 
#
# There is a lot of helpful information on this screen on how to get started with Db2 on Cloud. The important section that you need to check out immediately is the Db2 Credentials.
# ## Db2 Credentials
# Once the access to a Db2 database has been confirmed, you can see the service in your dashboard. The dashboard is accessed by clicking on the ☰ symbol on the left side of the menu bar.
#
# 
#
# In the top left-hand box you will see the **Services** keyword. Click on that to view all of your services.
#
# 
#
# You will see that the Db2 service is there (called Db2-yk since we used the default name).
# When you click on the resource name **Db2-yk**, the system will display information about the Db2 connection.
# 
# From this screen you can access the console of the database and start loading and querying data.
#
# 
# In order to do the labs in this system, you must get the connection information. You can see some of the information by clicking on the **Connection Info (Connection Information)** section on the left side of the screen. This gives you some of the credentials needed to connect to the system.
#
# 
# What is missing from this screen is the password for your userid. **Note:** Your userid for connecting to Db2 is different than your IBM Cloud userid. There is a unique userid and schema created for you when you sign up for the Db2 on Cloud service. To get the password for your userid you must navigate back to the Service Details screen. When you clicked on the Service name, it would have opened up a separate browser tab for the Db2 on Cloud console. You should see the Service Details tab at the top of your browser:
#
# 
# On the Service Details screen you will see a menu with the **Service Credentials** as an option.
# 
# Selecting **Service Credentials** will display a list of authorized users for the Db2 on Cloud service.
# 
# If no services credentials are there, you will need to create one. To create a new service credential, click on the **New Credential** button. The following screen will be displayed.
#
# 
# Give the credential a meaningful name and then press **Add**. This will result in a new credential being added to the list.
#
# 
# Selecting **View credentials** will give you the necessary information to connect to Db2 on Cloud. The display will look similar to the following screen.
#
# 
# For connecting to Db2 on Cloud you will need a copy of these credentials, so click on the copy icon next to the credential information. This will copy all of the information found in the JSON document above and it will be used to establish a connection with Db2 without you having to remember all of the information on this screen.
#
# At this point you now have all of the information you need to get started running the labs.
# ### Skytap Considerations
# The JSON document that was copied in the previous step needs to be placed into the Db2 Connections notebook. If you copied the information to the clipboard **outside of the Skytap image** then the cut and paste operation into a Skytap image requires an intermediate step. If you have copied something to your computers clipboard, you must first place it into the Skytap clipboard. At the top of the browser that is running your Skytap image, you will see the Skytap toolbar:
#
# 
#
# Clicking on the clipboard icon will display the input area for anything you want to paste into the virtual machine.
#
# 
#
# Once you have copied your credentials into this box it will display a `Success` message and then you can then paste the information into the appropriate cell.
# #### Credits: IBM 2019, <NAME> [<EMAIL>]
| Db2_on_Cloud_Registration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import sys, os, pdb, importlib
from matplotlib import rc
from matplotlib import rcParams
## Plotting
from matplotlib import pyplot as plt
from scipy.interpolate import interp1d
from scipy.signal import find_peaks
from scipy.ndimage.interpolation import rotate
## Project-specific notebook
sys.path.append('../src/')
import ast2050.lab4 as l4
rc('text', usetex=True)
rc('font', family='DejaVu Sans')
rc('font', serif='cm')
rc('font', size=16)
# -
data1 = l4.read_tiff('./data/April17/Neon1.tiff')[500:550,0:800]
fig = plt.figure( figsize=(15,15) )
plt.imshow(data1, vmin=0, vmax=255, origin='lower')
plt.show()
data_summed = np.zeros((800, 3))
for i in range(np.shape(data1)[1]):
temp = np.sum(data1[:,i])
data_summed[i] = temp
plt.plot(data_summed[:,2])
def envelope(wav, depth, numbins):
bins = np.linspace(min(wav), max(wav), numbins)
digitized = np.digitize(wav, bins)
bin_mins = [depth[digitized==i].min() for i in range(1, len(bins))]
idxs = [np.where(depth==mi)[0][0] for mi in bin_mins]
wavs = [wav[i] for i in idxs]
F = interp1d(wavs, bin_mins, fill_value='extrapolate')
return depth - F(wav)
test = envelope(np.arange(800), data_summed[:,0], 10)
peaks, _ = find_peaks(test, height=275)
fig = plt.figure(figsize=(16, 8))
plt.plot(test, color='k')
plt.plot(peaks, test[peaks], "x", color='red')
for peak in peaks:
plt.axvline(peak, linestyle='--', color='red', linewidth=0.5, alpha=0.6)
plt.xlim(200, 600)
plt.xlabel('Pixel Value', fontsize=18)
plt.ylabel('Flux [arbitrary units]', fontsize=18)
plt.ylim(-150, 2300)
plt.show()
neon_vals = np.asarray([7173.94, 7032.41, 6929.47, 6717.04, 6678.28, 6598.95, 6532.88, 6506.53, 6402.25, 6382.99, 6334.43, 6304.79, 6266.49, 6217.28, 6163.59, 6143.06, 6096.16, 6074.34, 6030.00, 5975.53, 5944.83, 5881.89, 5852.49])
cal = interp1d(peaks[1:], neon_vals, fill_value='extrapolate')
sol = cal(np.arange(200, 600))
fig = plt.figure(figsize=(6, 2))
plt.plot(np.arange(200, 600), sol, 'k')
plt.plot(peaks[1:], neon_vals, 'o', color='m')
plt.xlabel('Pixel Value')
plt.ylabel('Wavelength Value [Angstroms]')
data2 = l4.read_tiff('./data/18April/Hydrogen1.tiff')[500:550]
fig = plt.figure( figsize=(15,15) )
plt.imshow(data2, vmin=0, vmax=255, origin='lower')
plt.show()
data2_summed = np.zeros((1288, 3))
for i in range(np.shape(data2)[1]):
temp = np.sum(data2[:,i])
data2_summed[i] = temp
plt.plot(data2_summed[:,2])
test2 = envelope(np.arange(1288), data2_summed[:,0], 40)
peaks2, _ = find_peaks(test2, height=1200)
plt.plot(test2)
fig = plt.figure(figsize=(16, 8))
plt.plot(test2, color='k')
plt.plot(peaks2, test2[peaks2], "x", color='red')
for peak2 in peaks2:
plt.axvline(peak2, linestyle='--', color='red', linewidth=0.5, alpha=0.6)
plt.xlim(200, 1000)
plt.xlabel('Pixel Value', fontsize=18)
plt.ylabel('Flux [arbitrary units]', fontsize=18)
#plt.ylim(-150, 2300)
plt.show()
h_vals = np.asarray([6564.5377, 4861.3615, 4340.462])
cal_h = interp1d(peaks2, h_vals, fill_value='extrapolate')
sol_h = cal_h(np.arange(200, 1000))
fig = plt.figure(figsize=(6, 2))
plt.plot(np.arange(200, 1000), sol_h, 'k')
plt.plot(peaks2, h_vals, 'o', color='m')
plt.xlabel('Pixel Value')
plt.ylabel('Wavelength Value [Angstroms]')
fig = plt.figure(figsize=(10, 8))
plt.plot(np.arange(200, 600), sol, label='Neon Solution')
plt.plot(np.arange(200, 1000), sol_h, label='Hydrogen Solution')
plt.legend(loc='best', fancybox='True')
plt.xlabel('Pixel Value')
plt.ylabel('Wavelength Value [Angstroms]')
plt.show()
master_cal = interp1d(np.concatenate((peaks[1:], peaks2)), np.concatenate((neon_vals, h_vals)), fill_value='extrapolate')
master_sol = master_cal(np.arange(1288))
fig = plt.figure(figsize=(20, 6))
plt.plot(np.arange(1288), master_sol, color='k', label='Wavelength Solution', linestyle=':')
plt.plot(peaks[1:], neon_vals, 'o', label='Neon Peaks', markersize=10, alpha=0.9, mec='k')
plt.plot(peaks2, h_vals, 'o', label='Hydrogen Peaks', markersize=10, alpha=0.9, mec='k')
plt.legend(loc='best', fancybox='True', fontsize=16)
plt.xlabel('Pixel Value', fontsize=18)
plt.ylabel('Wavelength Value [Angstroms]', fontsize=18)
plt.show()
sun = l4.read_tiff('./data/April17/Sun1-1.tiff')[500:550]
fig = plt.figure( figsize=(15,15) )
plt.imshow(sun, vmin=0, vmax=255, origin='lower')
plt.show()
sun_peaks = np.asarray([np.argmax(sun[:,i,2]) for i in range(400, 800)])
fig = plt.figure( figsize=(30,30) )
plt.imshow(sun, vmin=0, vmax=255, origin='lower')
plt.plot(np.arange(1288), line[1] + np.arange(1288)*line[0], color='red', linestyle=':')
plt.xlabel('Pixel Value', fontsize=18)
plt.ylabel('Pixel Value', fontsize=18)
plt.show()
plt.plot(np.arange(400, 800), sun_peaks, '.')
plt.plot(np.arange(400, 800), line[1] + np.arange(400, 800)*line[0])
line = np.polyfit(np.arange(400, 800), sun_peaks, 1)
subtracted_sun = np.zeros(np.shape(sun)[:2])
for i in range(np.shape(sun)[1]):
subtracted_sun[:,i] = sun[:,i,2] - (line[1] + i*line[0])
plt.pcolormesh(np.arange(1288), np.arange(50), sun[:,:,2])
plt.plot(np.arange(1288), line[1] + line[0]*np.arange(1288))
yint = line[1]
yint2 = line[1] + 1288*line[0]
rot_angle = np.rad2deg(np.arctan((yint-yint2)/1288))
rotated_sun = rotate(sun, -rot_angle)
fig = plt.figure( figsize=(30,30) )
plt.imshow(rotated_sun, vmin=0, vmax=255, origin='lower')#, aspect='auto')
plt.xlabel('Pixel Value', fontsize=18)
plt.ylabel('Pixel Value', fontsize=18)
plt.show()
summed_sun = np.zeros((1288, 3))
for i in range(np.shape(rotated_sun)[1])[:-1]:
temp = np.sum(rotated_sun[30:34,i])
summed_sun[i] = temp
fig = plt.figure(figsize=(20, 12))
plt.plot(master_sol, summed_sun[:,2], color='k')
plt.xlabel('Wavelength [Angstroms]', fontsize=18)
plt.ylabel('Flux [Arbitrary Units]', fontsize=18)
plt.xlim(min(master_sol), max(master_sol))
plt.axvline(4861.3615, color='aqua', linestyle=':', label=r'H-$\beta$')
plt.axvline(5183.62, color='limegreen', linestyle=':', label=r'Mg b${}_1$ and b${}_2$')
plt.axvline(5172.70, color='limegreen', linestyle=':')
plt.axvline(5270.39, color='darkgreen', linestyle=':', label='Fe E${}_2$')
plt.axvline(5895.92, color='orange', linestyle=':')
plt.axvline(5889.95, color='orange', linestyle=':', label=r'Na D${}_1$ and D${}_2$ Doublet')
plt.axvline(6564.5377, color='red', linestyle=':', label=r'H-$\alpha$')
plt.axvline(6867.19, color='maroon', linestyle=':', label=r'O${}_2$ B')
plt.legend(loc='best', fancybox='True', fontsize=16)
plt.show()
| Lab4/Lab4-Calibration-Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from astropy import coordinates as coord
from astropy import units as u
from astropy.coordinates import Angle
from gala.coordinates import MagellanicStreamNidever08
import numpy as np
import sys
import pandas as pd
import pyregion
# ## Filter all sky tiles to just the GASKAP target set
#
# This notebook will take a set of all sky tiles and filter them down to a defined range. The range file is a csv with each row specifying a rule as follows:
#
# * min_l - The minimum Magellanic longitude of the rule (inclusive)
# * num_l - The number of 5 deg longitude bands that the rule covers
# * max_l - The maximum Magellanic longitude of the rule (inclusive)
# * min_b - The minimum Magellanic latitude (inclusive) to be included in this longitude range
# * max_b - The minimum Magellanic latitude (inclusive) to be included in this longitude range
# * num_b - The number of 5 deg tiles to be included
# +
def find_ms_b_range(tile_ranges, ms_l):
for row in tile_ranges:
if row[0] <= ms_l and row[2] >= ms_l:
#print (row[0], row[2], row[3], row[4])
return row[3], row[4]
return None
#find_ms_b_range(tile_ranges.to_numpy(), -15)
# -
def filter_regions(np_tile_ranges, regfile):
clippedarray=np.zeros(11)
#convert from gal to mag and check elements
for i in range(0,len(regfile),2):
#load in coordinates as j2000, references one of the tile vertices i think,should change to centre where the text is placed
c = coord.FK5(ra=float(regfile[i+1,1])*u.deg, dec=float(regfile[i+1,2])*u.deg)
#convert to mag coords
ms = c.transform_to(MagellanicStreamNidever08)
# Find the appropriate range
ms_b_range = find_ms_b_range(np_tile_ranges, ms.L.value)
if ms_b_range and ms_b_range[0] <= ms.B.value <= ms_b_range[1]:
print ("tile is inside at L={:.2f} B={:.2f}".format(ms.L.value, ms.B.value))
clippedarray=np.vstack((clippedarray,regfile[i]))
clippedarray=np.vstack((clippedarray,regfile[i+1]))
clippedarray[-1,-5:] = ""
# Remove the first, template row
clippedarray=clippedarray[1:,]
return clippedarray
# ## Produce region files for the pilot survey
regfileloc = 'gaskap_mag_2021.reg'
#regfileloc = 'croppedtilefield.reg'
importreg= pd.read_csv(regfileloc, delim_whitespace=True, header=None)
regfile=importreg.to_numpy(dtype='str')
regfile
tile_ranges = pd.read_csv('pilot_tile_range.csv')
tile_ranges
np_tile_ranges = tile_ranges.to_numpy()
clippedarray = filter_regions(np_tile_ranges, regfile)
# +
np.savetxt('pilottilefield_202110.reg',clippedarray,fmt='%s')
np.savetxt('pilottilefield_202110_notext.reg',clippedarray[0::2],fmt='%s')
# -
# ## Produce region files for the entire survey
# +
regfileloc = 'gaskap_mag_2021.reg'
#regfileloc = 'croppedtilefield.reg'
importreg= pd.read_csv(regfileloc, delim_whitespace=True, header=None)
regfile=importreg.to_numpy(dtype='str')
tile_ranges = pd.read_csv('tile_range.csv')
tile_ranges
# +
np_tile_ranges = tile_ranges.to_numpy()
clippedarray = filter_regions(np_tile_ranges, regfile)
np.savetxt('tilefield_202110.reg',clippedarray,fmt='%s')
np.savetxt('tilefield_202110_notext.reg',clippedarray[0::2],fmt='%s')
# -
| filter_tiles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="h5OEIzVgeM8C" colab_type="text"
# # Homework 2, *part 1* (40 points)
#
# This warm-up problem set is provided to help you get used to PyTorch.
#
# Please, only fill parts marked with "`Your code here`".
# + id="1sfI-m2-eM8E" colab_type="code" colab={}
import numpy as np
import math
import matplotlib.pyplot as plt
# %matplotlib inline
import torch
assert torch.__version__ >= '1.0.0'
import torch.nn.functional as F
import time
# + [markdown] id="SWJ9OjcEeM8K" colab_type="text"
# To learn best practices $-$ for example,
#
# * how to choose between `.sqrt()` and `.sqrt_()`,
# * when to use `.view()` and how is it different from `.reshape()`,
# * which `dtype` to use,
#
# $-$ you are expected to google a lot, read tutorials on the Web and study documentation.
#
# Quick documentation on functions and modules is available with `?` and `help()`, like so:
# + id="mnrSmmiLeM8M" colab_type="code" outputId="39c9034f-0a64-4379-b7d8-11246eedaadb" colab={"base_uri": "https://localhost:8080/", "height": 385}
help(torch.sqrt)
# + id="WMxpKoVOeM8Q" colab_type="code" colab={}
# to close the Jupyter help bar, press `Esc` or `q`
# ?torch.cat
# + [markdown] id="QOoBcpRaeM8T" colab_type="text"
# ## Task 1 (3 points)
#
# Use tensors only: **no lists, loops, numpy arrays etc.**
#
# *Clarification update:*
#
# 1. *you mustn't emulate PyTorch tensors with lists or tuples. Using a list for scaffolding utilities not provided by PyTorch core (e.g. to store model's layers or to group function arguments) is OK;*
# 2. *no loops*;
# 3. *you mustn't use numpy or other tensor libraries except PyTorch.*
#
# $\rho(\theta)$ is defined in polar coordinate system:
#
# $$\rho(\theta) = (1 + 0.9 \cdot \cos{8\theta} ) \cdot (1 + 0.1 \cdot \cos{24\theta}) \cdot (0.9 + 0.05 \cdot \cos {200\theta}) \cdot (1 + \sin{\theta})$$
#
# 1. Create a regular grid of 1000 values of $\theta$ between $-\pi$ and $\pi$.
# 2. Compute $\rho(\theta)$ at these values.
# 3. Convert it into Cartesian coordinates ([howto](http://www.mathsisfun.com/polar-cartesian-coordinates.html)).
# + id="uois9JLLeM8U" colab_type="code" outputId="b550eded-f923-4994-b6fc-b18d4afc9582" colab={"base_uri": "https://localhost:8080/", "height": 378}
# (1) Your code here
theta = torch.linspace(- np.pi, np.pi, steps=1000)
assert theta.shape == (1000,)
# (2) Your code here
rho = (1 + 0.9 * torch.cos(8*theta)) * (1 + 0.1 * torch.cos(24 theta)) * (0.9 + 0.05 * torch.cos(200*theta)) * (1 + torch.sin(theta))
assert torch.is_same_size(rho, theta)
# (3) Your code here
## having (r, theta):
## x = r*cos(theta)
## y = r*sin(theta)
x = rho*torch.cos(theta)
y = rho*torch.sin(theta)
plt.figure(figsize=[6,6])
plt.fill(x.data.numpy(), y.data.numpy(), color='green')
plt.grid()
# + [markdown] id="v1F_ap-FeM8Z" colab_type="text"
# ## Task 2 (7 points)
#
# Use tensors only: **no lists, loops, numpy arrays etc.**
#
# *Clarification update: see task 1.*
#
# We will implement [Conway's Game of Life](https://en.wikipedia.org/wiki/Conway's_Game_of_Life) in PyTorch.
#
# 
#
# If you skipped the URL above, here are the rules:
# * You have a 2D grid of cells, where each cell is "alive"(1) or "dead"(0)
# * At one step in time, the generation update happens:
# * Any living cell that has 2 or 3 neighbors survives, otherwise (0,1 or 4+ neighbors) it dies
# * Any cell with exactly 3 neighbors becomes alive if it was dead
#
# You are given a reference numpy implementation of the update step. Your task is to convert it to PyTorch.
# + id="IIG5upfEeM8Z" colab_type="code" colab={}
from scipy.signal import correlate2d as conv2d
def numpy_update(alive_map):
# Count neighbours with convolution
conv_kernel = np.array([[1,1,1],
[1,0,1],
[1,1,1]])
num_alive_neighbors = conv2d(alive_map, conv_kernel, mode='same')
# Apply game rules
born = np.logical_and(num_alive_neighbors == 3, alive_map == 0)
survived = np.logical_and(np.isin(num_alive_neighbors, [2,3]), alive_map == 1)
np.copyto(alive_map, np.logical_or(born, survived))
# + id="96aK1fOBeM8e" colab_type="code" colab={}
from torch.autograd import Variable
def torch_update(alive_map):
"""
Game of Life update function that does to `alive_map` exactly the same as `numpy_update`.
:param alive_map: `torch.tensor` of shape `(height, width)` and dtype `torch.float32`
containing 0s (dead) an 1s (alive)
"""
# Your code here
# Count neighbours with convolution
conv_kernel = torch.Tensor([[1,1,1],
[1,0,1],
[1,1,1]]).float().unsqueeze_(0).unsqueeze_(0)
num_alive_neighbors = torch.conv2d(alive_map[None, None, :], conv_kernel, padding = 1).squeeze_(0).squeeze_(0)
# Apply game rules
born = (num_alive_neighbors == 3) & (alive_map == 0)
survived = ((num_alive_neighbors == 2) | (num_alive_neighbors == 3)) & (alive_map == 1)
alive_map.copy_((born | survived))
# + id="hv9IxoXXeM8h" colab_type="code" outputId="9a074596-ec89-4470-8786-d87c5e4cf1b4" colab={"base_uri": "https://localhost:8080/", "height": 33}
# Generate a random initial map
alive_map_numpy = np.random.choice([0, 1], p=(0.5, 0.5), size=(100, 100))
alive_map_torch = torch.tensor(alive_map_numpy).float().clone()
numpy_update(alive_map_numpy)
torch_update(alive_map_torch)
# results should be identical
assert np.allclose(alive_map_torch.numpy(), alive_map_numpy), \
"Your PyTorch implementation doesn't match numpy_update."
print("Well done!")
# + id="CiHi2nZreM8l" colab_type="code" outputId="207e2752-f6b7-4ae2-fc32-5b2e48039d11" colab={"base_uri": "https://localhost:8080/", "height": 17}
# %matplotlib notebook
plt.ion()
# initialize game field
alive_map = np.random.choice([0, 1], size=(100, 100))
alive_map = torch.tensor(alive_map).float()
fig = plt.figure()
ax = fig.add_subplot(111)
fig.show()
for _ in range(100):
torch_update(alive_map)
# re-draw image
ax.clear()
ax.imshow(alive_map.view(100, 100).numpy(), cmap='gray')
fig.canvas.draw()
# + id="4IPKhwqAeM8o" colab_type="code" outputId="76e38e57-3564-4920-f67e-8bdf92c4f406" colab={"base_uri": "https://localhost:8080/", "height": 17}
# A fun setup for your amusement
alive_map = np.arange(100) % 2 + np.zeros([100, 100])
alive_map[48:52, 50] = 1
alive_map = torch.tensor(alive_map).float()
fig = plt.figure()
ax = fig.add_subplot(111)
fig.show()
for _ in range(150):
torch_update(alive_map)
ax.clear()
ax.imshow(alive_map.numpy(), cmap='gray')
fig.canvas.draw()
# + [markdown] id="K5P2teDHeM8r" colab_type="text"
# More fun with Game of Life: [video](https://www.youtube.com/watch?v=C2vgICfQawE)
# + [markdown] id="cbGcQ0SHeM8s" colab_type="text"
# ## Task 3 (30 points)
#
# You have to solve yet another character recognition problem: 10 letters, ~14 000 train samples.
#
# For this, we ask you to build a multilayer perceptron (*i.e. a neural network of linear layers*) from scratch using **low-level** PyTorch interface.
#
# Requirements:
# 1. at least 82% accuracy
# 2. at least 2 linear layers
# 3. use [softmax followed by categorical cross-entropy](https://gombru.github.io/2018/05/23/cross_entropy_loss/)
#
# **You are NOT allowed to use**
# * numpy arrays
# * `torch.nn`, `torch.optim`, `torch.utils.data.DataLoader`
# * convolutions
#
# ##### Clarification update:
#
# 1. *you mustn't emulate PyTorch tensors with lists or tuples. Using a list for scaffolding utilities not provided by PyTorch core (e.g. to store model's layers or to group function arguments) is OK;*
# 2. *you mustn't use numpy or other tensor libraries except PyTorch;*
# 3. *the purpose of part 1 is to make you google and read the documentation a LOT so that you learn which intrinsics PyTorch provides and what are their interfaces. This is why if there is some tensor functionality that is directly native to PyTorch, you mustn't emulate it with loops. Example:*
#
# ```
# x = torch.rand(1_000_000)
#
# # Wrong: slow and unreadable
# for idx in range(x.numel()):
# x[idx] = math.sqrt(x[idx])
#
# # Correct
# x.sqrt_()
# ```
#
# 4. *Loops are prohibited except for iterating over*
#
# * *parameters (and their companion tensors used by optimizer, e.g. running averages),*
# * *layers,*
# * *epochs (or "global" gradient steps if you don't use epoch logic),*
# * *batches in the dataset (using loops for collecting samples into a batch is not allowed).*
#
# Tips:
#
# * Pick random batches (either shuffle data before each epoch or sample each batch randomly).
# * Do not initialize weights with zeros ([learn why](https://stats.stackexchange.com/questions/27112/danger-of-setting-all-initial-weights-to-zero-in-backpropagation)). Gaussian noise with small variance will do.
# * 50 hidden neurons and a sigmoid nonlinearity will do for a start. Many ways to improve.
# * To improve accuracy, consider changing layers' sizes, nonlinearities, optimization methods, weights initialization.
# * Don't use GPU yet.
#
# **Reproducibility requirement**: you have to format your code cells so that `Cell -> Run All` on a fresh notebook **reliably** trains your model to the desired accuracy in a couple of minutes and reports the accuracy reached.
#
# Happy googling!
# + id="CNnQUkEHF9gP" colab_type="code" colab={}
import os
import numpy as np
from scipy.misc import imread,imresize
from sklearn.model_selection import train_test_split
from glob import glob
def load_notmnist(path='./notMNIST_small',letters='ABCDEFGHIJ',
img_shape=(28,28),test_size=0.25,one_hot=False):
# download data if it's missing. If you have any problems, go to the urls and load it manually.
if not os.path.exists(path):
print("Downloading data...")
assert os.system('curl http://yaroslavvb.com/upload/notMNIST/notMNIST_small.tar.gz > notMNIST_small.tar.gz') == 0
print("Extracting ...")
assert os.system('tar -zxvf notMNIST_small.tar.gz > untar_notmnist.log') == 0
data,labels = [],[]
print("Parsing...")
for img_path in glob(os.path.join(path,'*/*')):
class_i = img_path.split(os.sep)[-2]
if class_i not in letters: continue
try:
data.append(imresize(imread(img_path), img_shape))
labels.append(class_i,)
except:
print("found broken img: %s [it's ok if <10 images are broken]" % img_path)
data = np.stack(data)[:,None].astype('float32')
data = (data - np.mean(data)) / np.std(data)
#convert classes to ints
letter_to_i = {l:i for i,l in enumerate(letters)}
labels = np.array(list(map(letter_to_i.get, labels)))
if one_hot:
labels = (np.arange(np.max(labels) + 1)[None,:] == labels[:, None]).astype('float32')
#split into train/test
X_train, X_test, y_train, y_test = train_test_split(data, labels, test_size=test_size, random_state=42)
print("Done")
return X_train, y_train, X_test, y_test
# + id="FklzvacdeM8t" colab_type="code" outputId="8dd348ef-c924-4b18-b35f-cc817daa2053" colab={"base_uri": "https://localhost:8080/", "height": 217}
np.random.seed(666)
torch.manual_seed(666)
# from notmnist import load_notmnist
letters = 'ABCDEFGHIJ'
X_train, y_train, X_test, y_test = map(torch.tensor, load_notmnist(letters=letters))
X_train.squeeze_()
X_test.squeeze_();
# + id="slmcf2JdeM8y" colab_type="code" outputId="52645d2e-1829-43e3-fdf3-89a06a85b4ad" colab={"base_uri": "https://localhost:8080/", "height": 224}
fig, axarr = plt.subplots(2, 10, figsize=(15,3))
for idx, ax in enumerate(axarr.ravel()):
ax.imshow(X_train[idx].numpy(), cmap='gray')
ax.axis('off')
ax.set_title(letters[y_train[idx]])
# + [markdown] id="T9ljk3mCeM81" colab_type="text"
# The cell below has an example layout for encapsulating your neural network. Feel free to modify the interface if you need to (add arguments, add return values, add methods etc.). For example, you may want to add a method `do_gradient_step()` that executes one optimization algorithm (SGD / Adadelta / Adam / ...) step.
# + id="dE6PN3OXeM82" colab_type="code" colab={}
class NeuralNet:
def __init__(self, lr, batch_size):
# hyperparams
self.batch_size = batch_size
self.lr = lr
# first layer
self.W1 = torch.randn(28*28, 70, requires_grad=True)
self.b1 = torch.randn(1, 70, requires_grad=True)
# second layer
self.W2 = torch.randn(70, 10, requires_grad=True)
self.b2 = torch.randn(1, 10, requires_grad=True)
def _forward(self, x):
x = x.flatten(1)
x = x @ self.W1 + self.b1
x.sigmoid_()
x = x @ self.W2 + self.b2
x.exp_()
x = x / x.sum(dim=1, keepdim=True)
return x
def _loss(self, output, target):
log_softmax = output.log()
one_hot_target = torch.zeros(len(target), 10).scatter_(1, target.view(-1, 1), 1)
loss = -(log_softmax * one_hot_target).mean()
return loss
def _step(self, x, batch_target):
# forward
output = self._forward(x)
loss = self._loss(output, batch_target)
# backward
loss.backward()
# update weights
### first layer
self.W1.data -= self.lr * self.W1.grad.data
self.b1.data -= self.lr * self.b1.grad.data
### second layer
self.W2.data -= self.lr * self.W2.grad.data
self.b2.data -= self.lr * self.b2.grad.data
# zero grad
self.W1.grad.data.zero_()
self.b1.grad.data.zero_()
self.W2.grad.data.zero_()
self.b2.grad.data.zero_()
def train(self, X_train, y_train, n_epoch=150):
start_time = time.time()
for i in range(n_epoch):
t = time.time()
idx = torch.randperm(len(X_train))
n_batches = (len(X_train) - 1) // self.batch_size + 1
for j in range(n_batches):
X_batch = X_train[idx[j*self.batch_size : (j+1)*self.batch_size]]
y_batch = y_train[idx[j*self.batch_size : (j+1)*self.batch_size]]
self._step(X_batch, y_batch)
time_per_epoch = time.time() - t
train_acc = accuracy(model, X_train, y_train) * 100
test_acc = accuracy(model, X_test, y_test) * 100
print(f"EPOCH {i+1}: train acc: {train_acc:.2f} %")
end_time = time.time()
print(f'TOTAL TIME SPENT: {end_time - start_time:.1f}')
def predict(self, images):
"""
images: `torch.tensor` of shape `batch_size x height x width`
and dtype `torch.float32`.
returns: `output`, a `torch.tensor` of shape `batch_size x 10`,
where `output[i][j]` is the probability of `i`-th
batch sample to belong to `j`-th class.
"""
return self._forward(images)
# Your code here
# + id="mY-s9JPleM84" colab_type="code" colab={}
def accuracy(model, images, labels):
"""
model: `NeuralNet`
images: `torch.tensor` of shape `N x height x width`
and dtype `torch.float32`
labels: `torch.tensor` of shape `N` and dtype `torch.int64`. Contains
class index for each sample
returns:
fraction of samples from `images` correctly classified by `model`
"""
preds = model.predict(images).argmax(dim=1)
acc = torch.sum(preds == labels).float() / len(labels)
return acc
# + id="CfWFNCzCeM87" colab_type="code" outputId="59548a7f-862a-4942-a289-691229da7cad" colab={"base_uri": "https://localhost:8080/", "height": 3044}
model = NeuralNet(lr=0.055, batch_size=25)
# Your code here (train the model)
model.train(X_train, y_train, n_epoch= 180)
# + id="tXYsL9LNeM8-" colab_type="code" outputId="9285bb69-2c31-4e86-fa1b-1d0091e55010" colab={"base_uri": "https://localhost:8080/", "height": 33}
train_acc = accuracy(model, X_train, y_train) * 100
test_acc = accuracy(model, X_test, y_test) * 100
print("Train accuracy: %.2f, test accuracy: %.2f" % (train_acc, test_acc))
assert test_acc >= 82.0, "You have to do better"
# + id="IPjC8cQ3IEQ3" colab_type="code" colab={}
| skoltech_hw/hw02_p1_letters_recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution:
@staticmethod
def pdi_function(number: int, base: int = 10):
total = 0
while number > 0:
total += pow(number % base, 2)
number = number // base
return total
def isHappy(self, n: int) -> bool:
visited_nums = []
while n not in visited_nums:
visited_nums.append(n)
n = pdi_function(n)
if n == 1:
return True
return False
def pdi_function(number: int, base: int = 10):
total = 0
while number > 0:
total += pow(number % base, 2)
number = number // base
return total
assert pdi_function(19) == 82
assert pdi_function(82) == 68
a = (102 % 10)
a
102 // 10
s = Solution()
assert s.isHappy(19)
| HappyNumber.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
pd.set_option('display.max_columns', 500)
import numpy as np
from sklearn.neighbors import DistanceMetric
def haversine_vectorize(lon1, lat1, lon2, lat2):
lon1, lat1, lon2, lat2 = map(np.radians, [lon1, lat1, lon2, lat2])
newlon = lon2 - lon1
newlat = lat2 - lat1
haver_formula = np.sin(newlat/2.0)**2 + np.cos(lat1) * np.cos(lat2) * np.sin(newlon/2.0)**2
dist = 2 * np.arcsin(np.sqrt(haver_formula ))
km = 6367 * dist #6367 for distance in KM for miles use 3958
return km
d1 = pd.read_csv("0_VED_orig_data.csv")
d1.head(10)
mm = np.unique(d1['mm_edge_id'])
len(mm)
mm = np.unique(d1['trip'])
len(mm)
d1.shape
grouped_df = d1.groupby(["lat", "lon"])
first_values = grouped_df.first()
first_values = first_values.reset_index()
first_values.shape
first_values
grouped_df = first_values.groupby(["trip", "mm_edge_id"])
vals = grouped_df.first()
vals = vals.reset_index()
vals.shape
vals
def find_ratio(df):
t_max = np.max(pd.to_datetime(df['timestamp_ms']).astype('int64'))
t_min = np.min(pd.to_datetime(df['timestamp_ms']).astype('int64'))
dif = (t_max - t_min) / 1e3
return(dif)
def find_ratio_soc(df):
t_max = np.max(df['hvbattery_soc_per'])
t_min = np.min(df['hvbattery_soc_per'])
dif = (t_max - t_min)
return(dif)
tim = first_values[["trip", "mm_edge_id",'timestamp_ms']].groupby(["trip", "mm_edge_id"]).apply(find_ratio)
tim.to_numpy().shape
sped = first_values[["speed_kmh", "trip", "mm_edge_id"]].groupby(["trip", "mm_edge_id"]).apply(np.mean)
en = first_values[["trip", "mm_edge_id",'hvbattery_soc_per']].groupby(["trip", "mm_edge_id"]).apply(find_ratio_soc)
np.min(en), np.max(en)
vals['speed'] = sped['speed_kmh'].to_numpy()
vals['tim'] = tim.to_numpy()
vals['ev_kwh'] = en.to_numpy()
vals.head()
import matplotlib.pyplot as plt
plt.hist(vals['tim'])
def inf(data, col):
print(len(np.unique(data[col])), np.unique(data[col]))
print(data[col].describe())
d1.columns
np.unique(d1['mm_direction'])
np.sum(np.isnan(np.unique(d1['hourlydewpointtemperature'])))
inf(d1, 'trip')
# +
#'trip_id', -> trip
#'datekey', -> date
#'timekey', -> datetime
#'speed', -> speed_kmh
#'air_temperature', -> oat_degc
#'ev_kwh' -> hvbattery_soc_per
#'category', -> mm_edge_clazzs
#'wind_speed_ms', -> hourlywindspeed
#'wind_direction', -> hourlywinddirection
#'speedlimit_forward', -> mm_edge_kmh
#'speedlimit_backward', -> mm_edge_kmh
#'direction_x', -> mm_direction
#'trip_segmentno',
#'segmentkey',
#'segmentid',
#'meters_driven',
#'meters_segment',
#'seconds',
#'class',
#'segangle',
# -
data = vals[['trip', 'date', 'datetime', 'speed_kmh', 'oat_degc',
'hvbattery_soc_per', 'mm_direction', 'hourlywindspeed', 'hourlywinddirection',
'mm_edge_kmh', 'mm_edge_clazzs',
'lat','lon', 'speed', 'tim', 'ev_kwh']]
data.head()
trips = np.unique(data['trip'])
trips
idxes = np.ones(data.shape[0])
for i, trip in enumerate(trips):
if i % 10 == 0:
print(i)
ids = np.array([x for x in range(data.shape[0])])
tmp = (data['trip'] == trip)
tmp_idx = ids[tmp]
tmp_idx = tmp_idx[::10]
idxes[tmp] = False
idxes[tmp_idx] = True
sum(tmp), len(tmp_idx)
idxes = idxes.astype(np.bool)
data_filtered = data[idxes]
data_filtered.shape
data.shape
data_filtered = data
data_filtered.to_csv("./data/EVconsumption/0_filtered.csv")
data_ok = pd.DataFrame()
data_ok['trip_id'] = data_filtered['trip']
data_ok.shape
data_ok['trip_segmentno'] = -1
data_ok['segmentkey'] = -1
data_ok['segmentid'] = -1
data_ok['datekey'] = data_filtered['date'].str.replace('-','')
data_ok['direction_x'] = data_filtered['mm_direction']
data_ok['direction_x'] = data_ok['direction_x'].fillna(1)
data_ok['timekey'] = pd.to_datetime(data_filtered['datetime']).dt.strftime('%H%M')
data_ok['speed'] = data_filtered['speed']#data_filtered['speed_kmh']
data_ok['meters_driven'] = -1
data_ok['meters_segment'] = -1
data_ok['seconds'] = -1
data_ok['class'] = -1
data_ok['air_temperature'] = data_filtered['oat_degc']
data_ok['wind_direction'] = data_filtered['hourlywinddirection']
data_ok['wind_direction'] = data_ok['wind_direction'].fillna(0)
data_ok['wind_speed_ms'] = data_filtered['hourlywindspeed']
data_ok['wind_speed_ms'] = data_ok['wind_speed_ms'].fillna(0)
data_ok['category'] = data_filtered['mm_edge_clazzs'].str.replace('highway.','')
data_ok['segangle'] = 0
data_ok['speed_limit'] = data_filtered['mm_edge_kmh']
data_ok['speedlimit_forward'] = data_filtered['mm_edge_kmh']
data_ok['speedlimit_backward'] = data_filtered['mm_edge_kmh']
# +
data_ok['none'] = 0
data_ok['snow'] = 0
data_ok['thunder'] = 0
data_ok['fog'] = 0
data_ok['drifting'] = 0
data_ok['wet'] = 0
data_ok['dry'] = 0
data_ok['freezing'] = 0
# -
data_filtered['dist'] = haversine_vectorize(data_filtered['lon'],
data_filtered['lat'],
data_filtered['lon'].shift(),
data_filtered['lat'].shift())
data_filtered['ev_kwh'] = data_filtered['ev_kwh']#.shift() - data_filtered['hvbattery_soc_per']
data_filtered.head(10)
data_filtered['ev_kwh'][:10]
idxes = np.ones(data_filtered.shape[0])
for i, trip in enumerate(trips):
ids = np.array([x for x in range(data_filtered.shape[0])])
tmp = (data_filtered['trip'] == trip)
tmp_idx = ids[tmp]
idx = tmp_idx[0]
data_filtered['dist'][idx] = 0
data_filtered['ev_kwh'][idx] = 0
data_ok['seconds'] = data_filtered['tim']
data_ok.isnull().values.any()#, data_ok.isinf().values.any()
data_ok.head(20)
data_ok.to_csv("1_VED_orig_data.csv")
data = data_ok
import datetime
def to_weekend(vals):
weekDays = (0,0,0,0,0,1,1)
values = []
for v in vals:
thisXMas = datetime.date(int(str(v)[:4]), int(str(v)[4:6]), int(str(v)[6:]))
thisXMasDay = thisXMas.weekday()
thisXMasDayAsString = weekDays[thisXMasDay]
values.append(thisXMasDayAsString)
values = np.array(values)
return values
data['weekend'] = to_weekend(data['datekey'])
data['timekey'] = data['timekey'].astype(np.int)
data['time'] = 1
data['time'][data['timekey'] < 600] = 0
data['time'][data['timekey'] > 2200] = 0
data['time'][np.logical_and(data['timekey'] > 700, data['timekey'] < 900)] = 2
data['time'][np.logical_and(data['timekey'] > 1500, data['timekey'] < 1700)] = 2
data['category'][:14] = ['living_street', 'motorway', 'motorway_link', 'primary',
'residential', 'secondary', 'secondary_link', 'service',
'tertiary', 'track', 'trunk', 'trunk_link', 'unclassified',
'unpaved']
data = pd.concat([data, pd.get_dummies(data['category'])], axis=1)
d1 = data.drop(['datekey', 'direction_x', 'timekey', 'class', 'category',
'speedlimit_forward','meters_segment', 'speedlimit_backward'], axis=1)
d1.head()
d1.shape
d1.columns
a = ['Unnamed: 0', 'trip_id', 'trip_segmentno', 'segmentkey', 'segmentid',
'speed', 'meters_driven', 'seconds', 'air_temperature',
'wind_direction', 'wind_speed_ms', 'segangle', 'ev_kwh', 'weekend',
'speed_limit', 'time', 'drifting', 'dry', 'fog', 'freezing', 'none',
'snow', 'thunder', 'wet', 'living_street', 'motorway', 'motorway_link',
'primary', 'residential', 'secondary', 'secondary_link', 'service',
'tertiary', 'track', 'trunk', 'trunk_link', 'unclassified', 'unpaved']
# +
d1.dropna(axis = 0, how = 'any', inplace = True)
d2 = d1[d1['ev_kwh'].notna()]
d2 = d2[d2['wind_direction'].notna()]
d1.shape
# -
d2['speed_avg_week'] = 0
for seg in np.unique(d2['segmentkey']):
for val in np.unique(d2['weekend']):
idx = d2['segmentkey'] == seg
idx_w = d2['weekend'] == val
ids = idx & idx_w
speed = np.mean(d2[ids]['speed'])
d2.loc[ids, 'speed_avg_week'] = speed
#print(val)
d2['speed_avg_time'] = 0
for seg in np.unique(d2['segmentkey']):
for tim in np.unique(d2['time']):
idx = d2['segmentkey'] == seg
idx_w = d2['time'] == tim
ids = idx & idx_w
speed = np.mean(d2[ids]['speed'])
d2.loc[ids,'speed_avg_time'] = speed
#print(val)
d2['speed_avg_week_time'] = 0
for seg in np.unique(d2['segmentkey']):
for tim in np.unique(d2['time']):
for val in np.unique(d2['weekend']):
idx = d2['segmentkey'] == seg
idx_w = d2['weekend'] == val
idx_t = d2['time'] == tim
ids = idx & idx_w & idx_t
speed = np.mean(d2[ids]['speed'])
d2.loc[ids, 'speed_avg_week_time'] = speed
#print(val)
d2['speed_avg'] = 0
for seg in np.unique(d2['segmentkey']):
idx = d2['segmentkey'] == seg
ids = idx
speed = np.mean(d2[ids]['speed'])
d2.loc[ids, 'speed_avg'] = speed
#print(val)
d2.to_csv("2_VED_orig_data.csv")
df1 = d1.pop('trip_id')
df2 = d1.pop('ev_kwh')
df3 = d1.pop('trip_segmentno')
d1['trip_id'] = df1
d1['ev_kwh'] = df2
d1['trip_segmentno'] = df3
d1.head()
d1.shape
| prep_scripts/1_clean_VED.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="65958704b379a9676e5036c50e011036f7b71cc9"
# Thanks to this discussion for the observation: https://www.kaggle.com/c/santander-customer-transaction-prediction/discussion/84450
#
# In this notebook, I transform this column and re-order the train dataset using this column, and see what
# happens.
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import pandas as pd
from datetime import datetime
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
train_df = pd.read_csv('../input/train.csv')
# + [markdown] _uuid="f0fbcbdccd02eb8801be28f638709ef9b8771178"
# # Before you read
# + [markdown] _uuid="a02c4d6c07813386a5f1fdc6489f1c81ae63fb45"
# This exploration is an attempt to discover some hidden things behind the annonymization.
# Nothing is certain of course and this is so far specualative.
# Use this knowledge accordingly.
# + [markdown] _uuid="fe83208ccb4cc02d87bfa329c0d9a8a390709960"
# # Preliminary work
# + [markdown] _uuid="4b96e84ad022e750ec7229342a0fb5bf31f8f7aa"
# Two things to observe:
#
# - data has been annonymized
# - it comes from a business setting
#
# Thus, it is most likely (but not 100% sure) that some of the features
# contain date-like information (and also categorical features but that's
# for another day).
#
# How to find potential columns? Let's try to sort the columns using the number of unique
# values. What's the heurestic behind this choice?
# Well there shouldn't be a lot of dates, maybe few thousand top.
# + _uuid="7e1bac71a4dc7445485b3ad6537db0b823b53d1a"
train_df.drop(['ID_code', 'target'], axis=1).nunique().sort_values()
# + [markdown] _uuid="8e92a7d66f351654549f923a631ca0e6af9db2fe"
# ==> `var_68` has the least number of uniques, thus it **might** be a date-like column
# (it could also be a categorical column).
# There is also a possibility that this small number of uniques is a coincidence due to the rounding to 4 decimal numbers (bonus question: could you compute the probability of this event?)
# + _uuid="b46db78a4533a979abb56a020a6ddb215c296333"
f"Min: {train_df['var_68'].min()} and max: {train_df['var_68'].max()}"
# + [markdown] _uuid="bf3527b2dae50fa1ad6e07b10d80a5956c63aa7b"
# So how to extract a date?
# Well, first, get ride of the decimal values.
# Then transform to a datetime object supposing that it is an ordinal datetime.
# Try different offsets until you get a meaningful date range.
# That's it. Let's see this in action.
# + _uuid="f762e7cbcc37c0e393bc34ab6c42e91ea139beb7"
epoch_datetime = pd.datetime(1900, 1, 1)
trf_var_68_s = (train_df['var_68']*10000 - 7000 + epoch_datetime.toordinal()).astype(int)
date_s = trf_var_68_s.map(datetime.fromordinal)
train_df['date'] = date_s
sorted_train_df = train_df.drop('var_68', axis=1).sort_values('date')
# + [markdown] _uuid="2230f5057e675e5255a0d1f9cd09df8cce150a06"
# # Some plots
# + _uuid="8a1f6d85cab33728dc7dfe1ee2cf53fd5c84f6ac"
fig, ax = plt.subplots(1, 1, figsize=(20, 8))
sorted_train_df.set_index('date')['var_0'].plot(ax=ax)
# + _uuid="0b8be6bd2fd1fb50e89c413b24de8abf5484b744"
fig, ax = plt.subplots(1, 1, figsize=(20, 8))
sorted_train_df.set_index('date')['var_1'].plot(ax=ax)
# + _uuid="078b9c49ec2b60516b7c8a1970b585402854ca75"
fig, ax = plt.subplots(1, 1, figsize=(20, 8))
sorted_train_df.set_index('date')['var_2'].plot(ax=ax)
# + _uuid="81930d7b6346e4cd8d3ba1026d39574913be8d8c"
fig, ax = plt.subplots(1, 1, figsize=(20, 8))
sorted_train_df.set_index('date')['target'].plot(ax=ax)
# + [markdown] _uuid="fc1312306b437cb1b1d7f8562bf1bec82aae892c"
# # Date column exploration
# + [markdown] _uuid="2e9d7ed476185a3ea0f54cb725ddea22939cb97d"
# Alright, let's now explore this newly created column.
# + _uuid="de8e29cde62967093907a3ad3cdfd883781fed73"
date_s.nunique()
# + [markdown] _uuid="f3c8ce69df17e6fabc1bde16bef687cacd3b0be0"
# => I will thus use the `date` column to group rows.
# + _uuid="04e3499dc4ea5d559884619a866f74617baf1ea0"
f"Train starts: {date_s.min()}, ends: {date_s.max()}"
# + _uuid="dd474e0fa9e45ec3d95af56f91bcbeb1b839b015"
sorted_train_df['date'].dt.month.value_counts()
# + _uuid="835dc15122de3ae221d733dda793b2189e4508e6"
sorted_train_df['date'].dt.month.value_counts().plot(kind='bar')
# + _uuid="4ba4b06f589b01f175b86b5290a92a7ac6c11c4c"
sorted_train_df['date'].dt.year.value_counts()
# + _uuid="5005fb69937ba46654acf36b5d8fdd1420f1f239"
sorted_train_df['date'].dt.year.value_counts().plot(kind='bar')
# + _uuid="5ddec4e400c3b4872f74f8a23f3b52d1e910b495"
sorted_train_df['date'].dt.dayofweek.value_counts()
# + _uuid="99497b4303f6169813e4d4e5ce42e372a20f0b9e"
sorted_train_df['date'].dt.dayofweek.value_counts().plot(kind='bar')
# + [markdown] _uuid="2869825eb0833edc22894382057716c4a73c78c8"
# ==> Uniform day of week distribution. That's a good sign!
# + _uuid="43a225e930490c463f78b95ae430455c250c26c4"
fig, ax = plt.subplots(1, 1, figsize=(20, 8))
sorted_train_df.groupby('date')['target'].agg(['std', 'mean', 'max', 'min']).plot(ax=ax)
# + _uuid="fc19423b928de85817410754b5e5419cc0f5748d"
# In another cell signs the count is much bigger than the other statistics
fig, ax = plt.subplots(1, 1, figsize=(20, 8))
sorted_train_df.groupby('date')['target'].agg(['count']).plot(ax=ax)
# + [markdown] _uuid="5ecd7bf562cbd3bfec2f18fb70938306caeb6c3a"
# # What about the test?
# + [markdown] _uuid="87079ba51193aaa9dbbe66072366acc8f80bf661"
# Let's see if our observation transfers well to the test dataset.
# + _uuid="a0f57b3c8f2c43b3a0f98a11ca9e460c1fe9e995"
test_df = pd.read_csv('../input/test.csv')
epoch_datetime = pd.datetime(1900, 1, 1)
s = (test_df['var_68']*10000 - 7000 + epoch_datetime.toordinal()).astype(int)
test_df['date'] = s.map(datetime.fromordinal)
sorted_test_df = test_df.drop('var_68', axis=1).sort_values('date')
# + _uuid="aceb6820d181a9ba047b61f1a468de7d713a8d49"
f"Test starts: {test_df['date'].min()} and ends: {test_df['date'].max()}"
# + _uuid="341d02abba634e8dce820ab5dc5b384da05ce166"
test_df['date'].dt.year.value_counts().plot(kind='bar')
# + _uuid="91f2c924dd58e8ff1751c34e738a0048aa7f40b1"
test_df['date'].dt.month.value_counts().plot(kind='bar')
# + _uuid="739546ba42b9115417bcde1d7859937915ecc186"
test_df['date'].dt.dayofweek.value_counts().plot(kind='bar')
# + _uuid="f756c7d1e3a11e531f89db7167099288a8bd2293"
fig, ax = plt.subplots(1, 1, figsize=(20, 8))
sorted_test_df.groupby('date')['var_1'].agg(['count']).plot(ax=ax)
# + [markdown] _uuid="31ebeac4bba663d82ba054cc9fe883f3cc00e594"
# # Test and train date column comparaison
# + _uuid="67be6d4b27379449fab66f2222d7c27edce2af6c"
len(set(sorted_train_df['date']))
# + _uuid="9184b613c9d0512c869a5f13b65071ff3b0e6d01"
len(set(sorted_test_df['date']))
# + _uuid="0441e576db836cfbd0f8a84c03f8a81cb02d2fcc"
len(set(sorted_train_df['date']) & set(sorted_test_df['date']))
# + _uuid="6bd007e02fd16e584f0c48aa074fe7728fdb3304"
len(set(sorted_train_df['date']) - set(sorted_test_df['date']))
# + _uuid="6944eb219277463694bd2cad5a04b5f8b2255519"
len(set(sorted_test_df['date']) - set(sorted_train_df['date']))
# + _uuid="85ebfe8da69a7f16789cc6c52b9ec0739e200919"
set(sorted_test_df['date']) - set(sorted_train_df['date'])
# + [markdown] _uuid="83b27a4869b32ac0bef0ac4f1e9348bbea3f8e48"
# ==> Most of the dates overlap.
# + _uuid="c83b268440b75e5098688f1638f3b04baac54a3d"
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
sorted_train_df.groupby('date')['var_91'].count().plot(ax=ax, label="train")
sorted_test_df.groupby('date')['var_91'].count().plot(ax=ax, label="test")
ax.legend()
# + _uuid="494fd543f6efa3d59e260162347c7cc6b201fb53"
# Zoom on 2018
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
(sorted_train_df.loc[lambda df: df.date.dt.year == 2018]
.groupby('date')['var_91']
.count()
.plot(ax=ax, label="train"))
(sorted_test_df.loc[lambda df: df.date.dt.year == 2018]
.groupby('date')['var_91']
.count()
.plot(ax=ax, label="test"))
ax.legend()
# + _uuid="bb57f0e57b4f0ec2cb06a70ee00f42e8f7c24b98"
# Zoom on 2018-1
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
(sorted_train_df.loc[lambda df: (df.date.dt.year == 2018) & (df.date.dt.month == 1)]
.groupby('date')['var_91']
.count()
.plot(ax=ax, label="train"))
(sorted_test_df.loc[lambda df: (df.date.dt.year == 2018) & (df.date.dt.month == 1)]
.groupby('date')['var_91']
.count()
.plot(ax=ax, label="test"))
ax.legend()
# + _uuid="ad0bbb321b4c5377d519ed0eeabc21ab8b3a68d5"
# Zoom on 2018-1
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
(sorted_train_df.loc[lambda df: (df.date.dt.year == 2018) & (df.date.dt.month == 1)]
.groupby('date')['var_91']
.mean()
.plot(ax=ax, label="train"))
(sorted_test_df.loc[lambda df: (df.date.dt.year == 2018) & (df.date.dt.month == 1)]
.groupby('date')['var_91']
.mean()
.plot(ax=ax, label="test"))
ax.legend()
# + [markdown] _uuid="dce2f25f59e160624d545917fe796fd2467ed16d"
# Idea to try: predict the mean of the target (using the date
# for grouping) for the overlapping dates.
# + _uuid="95f8fd3be00036caed4f4c00038166894c0b47f3"
overlapping_dates = set(sorted_train_df['date']) & set(sorted_test_df['date'])
# + _uuid="baaf9a7049ffdce08203402336f2c054e339adf0"
grouped_df = (sorted_train_df.loc[lambda df: df.date.isin(overlapping_dates)]
.groupby('date')['target']
.mean())
# + _uuid="3bed40ffb34d276cbe4a9d43c53b15a60f3acfdf"
grouped_df.plot(kind='hist', bins=100)
# + _uuid="ee5c755bd3f10c7fb9e155c4639c5a3ab7bdc63d"
grouped_df.to_csv('grouped_df.csv', index=False)
# + [markdown] _uuid="0e3e3f61df9f5cc58a220769a262cb1c3957da72"
# # What to do now?
# + [markdown] _uuid="c5ca458c93c104beaf8a3455379a52a07674744c"
# Some of the things I will try to do:
# - Use this transformed column for a better temporal CV. Some ideas I have tried: stratification using years, day of weeks, and so on.
# - Transform other columns using this new one
#
# Stay tuned for more insights. :)
| 12 customer prediction/mystery-behind-var-68.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''dev'': venv)'
# name: python3
# ---
# We will start with an example using a randomly colored RGB image. Here we have a 16x16 pixel image with random 8-bit RGB colors. The first step is to convert the image into a vector of colors. We plot the first 20 colors from the vector to show how they correspond to the colors in the original image.
import colorsort as csort
import cv2
import matplotlib.pyplot as plt
import numpy as np
# +
num_x_pixels = 16
num_y_pixels = 16
num_channels = 3
image_shape = (num_x_pixels, num_y_pixels, num_channels)
np.random.seed(0)
image_rgb = np.random.randint(0,255,image_shape).astype(np.uint8)
plt.imshow(image_rgb)
plt.axis(False)
vec_rgb = csort.image_to_vec(image_rgb)
plt.figure()
plt.imshow(vec_rgb[None,:20], aspect=1)
plt.axis(False);
# -
# Next we will reduce the number of colors in the color vector. This leaves 10 colors that are representative of the original colors in the image. Color sorting can be done with a large list of colors, but the process is slow, and the results are unpredictable. If possible, reduce your color palette before sorting.
print(vec_rgb.shape)
vec_rgb_reduced = csort.reduce_colors(vec_rgb, 10)
print(vec_rgb_reduced.shape)
plt.imshow(vec_rgb_reduced[None], aspect=1)
plt.axis(False);
# Finally we run sort_colors on our reduced color vector. The colors are sorted in a pleasing order, and have the brightest color first, and the darkest color last.
vec_rgb_sorted = csort.sort_colors(vec_rgb_reduced)
print(vec_rgb_sorted.shape)
plt.imshow(vec_rgb_sorted[None], aspect=1)
plt.axis(False);
# Using real-world images, the color palette produced is even better.
# +
image_rgb = cv2.imread('saturn.png')[:,:,[2,1,0]]
plt.imshow(image_rgb)
plt.axis(False)
vec_rgb = csort.image_to_vec(image_rgb)
vec_rgb_reduced = csort.reduce_colors(vec_rgb, 10)
vec_rgb_sorted = csort.sort_colors(vec_rgb_reduced)
plt.figure()
plt.imshow(vec_rgb_sorted[None])
plt.axis(False);
# +
image_rgb = cv2.imread('macaws.png')[:,:,[2,1,0]]
plt.imshow(image_rgb)
plt.axis(False)
vec_rgb = csort.image_to_vec(image_rgb)
vec_rgb_reduced = csort.reduce_colors(vec_rgb, 8)
vec_rgb_sorted = csort.sort_colors(vec_rgb_reduced)
plt.figure()
plt.imshow(vec_rgb_sorted[None])
plt.axis(False);
| etc/examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import csv
import os
import numpy as np
# +
# Ler
dir = os.getcwd()
directory = '..\\datasets\\vehicle'
rows = []
for root,dirs,files in os.walk(directory):
for file in files:
if file.endswith(".dat"):
file_name = os.path.join(dir, directory, file)
csvfile = open(file_name, 'r')
readCSV = csv.reader(csvfile, delimiter=' ')
for row in readCSV:
rows.append(row)
csvfile.close()
# Gravar
#file = 'final.csv'
#with open(file, 'w', newline='') as f:
#writer = csv.writer(f)
#writer.writerows(row for row in rows if row)
# +
class custom_datasets():
@classmethod
def load_ionosphere(self):
return self.load_custom_dataset('..\\datasets\\ionosphere.data')
@classmethod
def load_wine(self):
return self.load_custom_dataset('..\\datasets\\wine_processed.data')
@classmethod
def load_live_disorders(self):
return self.load_custom_dataset('..\\datasets\\bupa.data')
@classmethod
def load_vehicle(self):
dataset = self.load_custom_dataset('..\\datasets\\vehicle.data', 'int')
dataset.data = dataset.data[1:]
dataset.target = dataset.target[1:]
return dataset
@classmethod
def load_balance_scale(self):
return self.load_custom_dataset('..\\datasets\\balance-scale.data')
@classmethod
def load_zoo(self):
#dataset = self.load_custom_dataset('..\\datasets\\skin-segmentation.data')
#dataset.data = dataset.data[1:]
#dataset.target = dataset.target[1:]
#return dataset
return self.load_custom_dataset('..\\datasets\\zoo.data')
@classmethod
def load_custom_dataset(self, path, dtype='float64'):
dir = os.getcwd()
file_name = os.path.join(dir, path)
data = np.genfromtxt(file_name, delimiter=',', dtype=dtype)
n_cols = data.shape[1]
X = data[:,0:n_cols-1]
y =np.genfromtxt(file_name, delimiter=',', usecols=(n_cols-1), dtype=None)
return Dataset(X, y)
class Dataset:
def __init__(self, X, y):
self.data = X
self.target = y
# +
np.set_printoptions(threshold=np.inf)
dataset = custom_datasets.load_zoo()
for i in range(dataset.data.shape[0]):
for j in range(dataset.data.shape[1]):
if(math.isnan(dataset.data[i][j])):
print((i,j))
# -
dataset.data[0]
dataset.data
np.all(np.isfinite(dataset.data))
np.any(np.isnan(dataset.data))
np.where(np.all(np.isnan(dataset.data), axis=1))[0]
| notebooks/Dataset Processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # About this Notebook
#
# Bayesian probabilistic matrix factorization (BPMF) is a classical model in the recommender system field. In the following, we will discuss:
#
# - What the BPMF is?
#
# - How to implement BPMF mainly using Python `Numpy` with high efficiency?
#
# - How to make data imputations with real-world spatiotemporal datasets?
#
# If you want to know more about BPMF, please read this article:
#
# > <NAME>, <NAME>, 2008. [**Bayesian probabilistic matrix factorization using Markov chain Monte Carlo**](https://www.cs.toronto.edu/~amnih/papers/bpmf.pdf). Proceedings of the 25th International Conference on Machine Learning (*ICML 2008*), Helsinki, Finland. [[Matlab code (official)](https://www.cs.toronto.edu/~rsalakhu/BPMF.html)]
#
# ## Quick Run
#
# This notebook is publicly available for any usage at our data imputation project. Please click [**transdim**](https://github.com/xinychen/transdim).
#
import numpy as np
from numpy.random import multivariate_normal as mvnrnd
from scipy.stats import wishart
from numpy.linalg import inv as inv
# # Part 1: Matrix Computation Concepts
#
# ## 1) Kronecker product
#
# - **Definition**:
#
# Given two matrices $A\in\mathbb{R}^{m_1\times n_1}$ and $B\in\mathbb{R}^{m_2\times n_2}$, then, the **Kronecker product** between these two matrices is defined as
#
# $$A\otimes B=\left[ \begin{array}{cccc} a_{11}B & a_{12}B & \cdots & a_{1m_2}B \\ a_{21}B & a_{22}B & \cdots & a_{2m_2}B \\ \vdots & \vdots & \ddots & \vdots \\ a_{m_11}B & a_{m_12}B & \cdots & a_{m_1m_2}B \\ \end{array} \right]$$
# where the symbol $\otimes$ denotes Kronecker product, and the size of resulted $A\otimes B$ is $(m_1m_2)\times (n_1n_2)$ (i.e., $m_1\times m_2$ columns and $n_1\times n_2$ rows).
#
# - **Example**:
#
# If $A=\left[ \begin{array}{cc} 1 & 2 \\ 3 & 4 \\ \end{array} \right]$ and $B=\left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10 \\ \end{array} \right]$, then, we have
#
# $$A\otimes B=\left[ \begin{array}{cc} 1\times \left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10\\ \end{array} \right] & 2\times \left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10\\ \end{array} \right] \\ 3\times \left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10\\ \end{array} \right] & 4\times \left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10\\ \end{array} \right] \\ \end{array} \right]$$
#
# $$=\left[ \begin{array}{cccccc} 5 & 6 & 7 & 10 & 12 & 14 \\ 8 & 9 & 10 & 16 & 18 & 20 \\ 15 & 18 & 21 & 20 & 24 & 28 \\ 24 & 27 & 30 & 32 & 36 & 40 \\ \end{array} \right]\in\mathbb{R}^{4\times 6}.$$
#
# ## 2) Khatri-Rao product (`kr_prod`)
#
# - **Definition**:
#
# Given two matrices $A=\left( \boldsymbol{a}_1,\boldsymbol{a}_2,...,\boldsymbol{a}_r \right)\in\mathbb{R}^{m\times r}$ and $B=\left( \boldsymbol{b}_1,\boldsymbol{b}_2,...,\boldsymbol{b}_r \right)\in\mathbb{R}^{n\times r}$ with same number of columns, then, the **Khatri-Rao product** (or **column-wise Kronecker product**) between $A$ and $B$ is given as follows,
#
# $$A\odot B=\left( \boldsymbol{a}_1\otimes \boldsymbol{b}_1,\boldsymbol{a}_2\otimes \boldsymbol{b}_2,...,\boldsymbol{a}_r\otimes \boldsymbol{b}_r \right)\in\mathbb{R}^{(mn)\times r}$$
# where the symbol $\odot$ denotes Khatri-Rao product, and $\otimes$ denotes Kronecker product.
#
# - **Example**:
#
# If $A=\left[ \begin{array}{cc} 1 & 2 \\ 3 & 4 \\ \end{array} \right]=\left( \boldsymbol{a}_1,\boldsymbol{a}_2 \right) $ and $B=\left[ \begin{array}{cc} 5 & 6 \\ 7 & 8 \\ 9 & 10 \\ \end{array} \right]=\left( \boldsymbol{b}_1,\boldsymbol{b}_2 \right) $, then, we have
#
# $$A\odot B=\left( \boldsymbol{a}_1\otimes \boldsymbol{b}_1,\boldsymbol{a}_2\otimes \boldsymbol{b}_2 \right) $$
#
# $$=\left[ \begin{array}{cc} \left[ \begin{array}{c} 1 \\ 3 \\ \end{array} \right]\otimes \left[ \begin{array}{c} 5 \\ 7 \\ 9 \\ \end{array} \right] & \left[ \begin{array}{c} 2 \\ 4 \\ \end{array} \right]\otimes \left[ \begin{array}{c} 6 \\ 8 \\ 10 \\ \end{array} \right] \\ \end{array} \right]$$
#
# $$=\left[ \begin{array}{cc} 5 & 12 \\ 7 & 16 \\ 9 & 20 \\ 15 & 24 \\ 21 & 32 \\ 27 & 40 \\ \end{array} \right]\in\mathbb{R}^{6\times 2}.$$
def kr_prod(a, b):
return np.einsum('ir, jr -> ijr', a, b).reshape(a.shape[0] * b.shape[0], -1)
A = np.array([[1, 2], [3, 4]])
B = np.array([[5, 6], [7, 8], [9, 10]])
print(kr_prod(A, B))
# ## 3) Computing Covariance Matrix (`cov_mat`)
#
# For any matrix $X\in\mathbb{R}^{m\times n}$, `cov_mat` can return a $n\times n$ covariance matrix for special use in the following.
def cov_mat(mat):
dim1, dim2 = mat.shape
new_mat = np.zeros((dim2, dim2))
mat_bar = np.mean(mat, axis = 0)
for i in range(dim1):
new_mat += np.einsum('i, j -> ij', mat[i, :] - mat_bar, mat[i, :] - mat_bar)
return new_mat
# # Part 2: Bayesian Probabilistic Matrix Factorization (BPMF)
#
#
def BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2):
"""Bayesian Probabilistic Matrix Factorization, BPMF."""
W = init["W"]
X = init["X"]
dim1, dim2 = sparse_mat.shape
dim = np.array([dim1, dim2])
pos = np.where((dense_mat != 0) & (sparse_mat == 0))
position = np.where(sparse_mat != 0)
binary_mat = np.zeros((dim1, dim2))
binary_mat[position] = 1
beta0 = 1
nu0 = rank
mu0 = np.zeros((rank))
W0 = np.eye(rank)
tau = 1
alpha = 1e-6
beta = 1e-6
W_plus = np.zeros((dim1, rank))
X_plus = np.zeros((dim2, rank))
mat_hat_plus = np.zeros((dim1, dim2))
for iters in range(maxiter1):
for order in range(2):
if order == 0:
mat = W.copy()
elif order == 1:
mat = X.copy()
mat_bar = np.mean(mat, axis = 0)
var_mu_hyper = (dim[order] * mat_bar + beta0 * mu0)/(dim[order] + beta0)
var_W_hyper = inv(inv(W0) + cov_mat(mat) + dim[order] * beta0/(dim[order] + beta0)
* np.outer(mat_bar - mu0, mat_bar - mu0))
var_Lambda_hyper = wishart(df = dim[order] + nu0, scale = var_W_hyper, seed = None).rvs()
var_mu_hyper = mvnrnd(var_mu_hyper, inv((dim[order] + beta0) * var_Lambda_hyper))
if order == 0:
var1 = X.T
mat0 = np.matmul(var1, sparse_mat.T)
elif order == 1:
var1 = W.T
mat0 = np.matmul(var1, sparse_mat)
var2 = kr_prod(var1, var1)
if order == 0:
mat1 = np.matmul(var2, binary_mat.T)
elif order == 1:
mat1 = np.matmul(var2, binary_mat)
var3 = tau * mat1.reshape(rank, rank, dim[order]) + np.dstack([var_Lambda_hyper] * dim[order])
var4 = tau * mat0 + np.dstack([np.matmul(var_Lambda_hyper, var_mu_hyper)] * dim[order])[0, :, :]
for i in range(dim[order]):
var_Lambda = var3[:, :, i]
inv_var_Lambda = inv((var_Lambda + var_Lambda.T)/2)
vec = mvnrnd(np.matmul(inv_var_Lambda, var4[:, i]), inv_var_Lambda)
if order == 0:
W[i, :] = vec.copy()
elif order == 1:
X[i, :] = vec.copy()
if iters + 1 > maxiter1 - maxiter2:
W_plus += W
X_plus += X
mat_hat = np.matmul(W, X.T)
if iters + 1 > maxiter1 - maxiter2:
mat_hat_plus += mat_hat
rmse = np.sqrt(np.sum((dense_mat[pos] - mat_hat[pos]) ** 2)/dense_mat[pos].shape[0])
var_alpha = alpha + 0.5 * sparse_mat[position].shape[0]
error = sparse_mat - mat_hat
var_beta = beta + 0.5 * np.sum(error[position] ** 2)
tau = np.random.gamma(var_alpha, 1/var_beta)
if (iters + 1) % 200 == 0 and iters < maxiter1 - maxiter2:
print('Iter: {}'.format(iters + 1))
print('RMSE: {:.6}'.format(rmse))
print()
W = W_plus/maxiter2
X = X_plus/maxiter2
mat_hat = mat_hat_plus/maxiter2
if maxiter1 >= 100:
final_mape = np.sum(np.abs(dense_mat[pos] - mat_hat[pos])/dense_mat[pos])/dense_mat[pos].shape[0]
final_rmse = np.sqrt(np.sum((dense_mat[pos] - mat_hat[pos]) ** 2)/dense_mat[pos].shape[0])
print('Imputation MAPE: {:.6}'.format(final_mape))
print('Imputation RMSE: {:.6}'.format(final_rmse))
print()
return mat_hat, W, X
# # Part 3: Data Organization
#
# ## 1) Matrix Structure
#
# We consider a dataset of $m$ discrete time series $\boldsymbol{y}_{i}\in\mathbb{R}^{f},i\in\left\{1,2,...,m\right\}$. The time series may have missing elements. We express spatio-temporal dataset as a matrix $Y\in\mathbb{R}^{m\times f}$ with $m$ rows (e.g., locations) and $f$ columns (e.g., discrete time intervals),
#
# $$Y=\left[ \begin{array}{cccc} y_{11} & y_{12} & \cdots & y_{1f} \\ y_{21} & y_{22} & \cdots & y_{2f} \\ \vdots & \vdots & \ddots & \vdots \\ y_{m1} & y_{m2} & \cdots & y_{mf} \\ \end{array} \right]\in\mathbb{R}^{m\times f}.$$
#
# ## 2) Tensor Structure
#
# We consider a dataset of $m$ discrete time series $\boldsymbol{y}_{i}\in\mathbb{R}^{nf},i\in\left\{1,2,...,m\right\}$. The time series may have missing elements. We partition each time series into intervals of predifined length $f$. We express each partitioned time series as a matrix $Y_{i}$ with $n$ rows (e.g., days) and $f$ columns (e.g., discrete time intervals per day),
#
# $$Y_{i}=\left[ \begin{array}{cccc} y_{11} & y_{12} & \cdots & y_{1f} \\ y_{21} & y_{22} & \cdots & y_{2f} \\ \vdots & \vdots & \ddots & \vdots \\ y_{n1} & y_{n2} & \cdots & y_{nf} \\ \end{array} \right]\in\mathbb{R}^{n\times f},i=1,2,...,m,$$
#
# therefore, the resulting structure is a tensor $\mathcal{Y}\in\mathbb{R}^{m\times n\times f}$.
# # Part 4: Experiments on Guangzhou Data Set
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.2
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = (np.round(random_tensor + 0.5 - missing_rate)
.reshape([random_tensor.shape[0], random_tensor.shape[1] * random_tensor.shape[2]]))
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 80
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.4
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = (np.round(random_tensor + 0.5 - missing_rate)
.reshape([random_tensor.shape[0], random_tensor.shape[1] * random_tensor.shape[2]]))
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 80
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.2
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(random_matrix[i1, i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1] * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.4
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(random_matrix[i1, i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1] * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# **Experiment results** of missing data imputation using BPMF:
#
# | scenario |`rank`|`maxiter1`|`maxiter2`| mape | rmse |
# |:----------|-----:|---------:|---------:|-------- --:|----------:|
# |**0.2, RM**| 80 | 1100 | 100 | **0.0954** | **4.0551**|
# |**0.4, RM**| 80 | 1100 | 100 | **0.0981** | **4.1659**|
# |**0.2, NM**| 10 | 1100 | 100 | **0.1028** | **4.2901**|
# |**0.4, NM**| 10 | 1100 | 100 | **0.1040** | **4.3994**|
#
# # Part 5: Experiments on Birmingham Data Set
#
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.1
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = (np.round(random_tensor + 0.5 - missing_rate)
.reshape([random_tensor.shape[0], random_tensor.shape[1] * random_tensor.shape[2]]))
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.3
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = (np.round(random_tensor + 0.5 - missing_rate)
.reshape([random_tensor.shape[0], random_tensor.shape[1] * random_tensor.shape[2]]))
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.1
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(random_matrix[i1, i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1] * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.3
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(random_matrix[i1, i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1] * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# **Experiment results** of missing data imputation using BPMF:
#
# | scenario |`rank`|`maxiter1`|`maxiter2`| mape | rmse |
# |:----------|-----:|---------:|---------:|-----------:|-----------:|
# |**10%, RM**| 30 | 1100 | 100 | **0.0787** | **81.593**|
# |**30%, RM**| 30 | 1100 | 100 | **0.0995** | **83.8159**|
# |**10%, NM**| 10 | 1100 | 100 | **0.1318** | **29.2774**|
# |**30%, NM**| 10 | 1100 | 100 | **0.1475** | **60.2924**|
#
# # Part 6: Experiments on Hangzhou Data Set
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.2
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = (np.round(random_tensor + 0.5 - missing_rate)
.reshape([random_tensor.shape[0], random_tensor.shape[1] * random_tensor.shape[2]]))
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 50
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.4
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = (np.round(random_tensor + 0.5 - missing_rate)
.reshape([random_tensor.shape[0], random_tensor.shape[1] * random_tensor.shape[2]]))
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 50
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.2
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(random_matrix[i1, i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1] * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.4
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(random_matrix[i1, i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1] * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# **Experiment results** of missing data imputation using BPMF:
#
# | scenario |`rank`|`maxiter1`|`maxiter2`| mape | rmse |
# |:----------|-----:|---------:|---------:|-----------:|-----------:|
# |**20%, RM**| 50 | 1100 | 100 | **0.2963** | **41.8653**|
# |**40%, RM**| 50 | 1100 | 100 | **0.3283** | **44.4621**|
# |**20%, NM**| 10 | 1100 | 100 | **0.3631** | **64.2751**|
# |**40%, NM**| 10 | 1100 | 100 | **0.3643** | **59.0373**|
#
# # Part 7: Experiments on Seattle Data Set
# +
import pandas as pd
dense_mat = pd.read_csv('../datasets/Seattle-data-set/mat.csv', index_col = 0)
RM_mat = pd.read_csv('../datasets/Seattle-data-set/RM_mat.csv', index_col = 0)
dense_mat = dense_mat.values
RM_mat = RM_mat.values
missing_rate = 0.2
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(RM_mat + 0.5 - missing_rate)
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 50
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import pandas as pd
dense_mat = pd.read_csv('../datasets/Seattle-data-set/mat.csv', index_col = 0)
RM_mat = pd.read_csv('../datasets/Seattle-data-set/RM_mat.csv', index_col = 0)
dense_mat = dense_mat.values
RM_mat = RM_mat.values
missing_rate = 0.4
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(RM_mat + 0.5 - missing_rate)
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 50
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import pandas as pd
dense_mat = pd.read_csv('../datasets/Seattle-data-set/mat.csv', index_col = 0)
NM_mat = pd.read_csv('../datasets/Seattle-data-set/NM_mat.csv', index_col = 0)
dense_mat = dense_mat.values
NM_mat = NM_mat.values
missing_rate = 0.2
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros((dense_mat.shape[0], 28, 288))
for i1 in range(binary_tensor.shape[0]):
for i2 in range(binary_tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(NM_mat[i1, i2] + 0.5 - missing_rate)
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_tensor.reshape([dense_mat.shape[0], dense_mat.shape[1]]))
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import pandas as pd
dense_mat = pd.read_csv('../datasets/Seattle-data-set/mat.csv', index_col = 0)
NM_mat = pd.read_csv('../datasets/Seattle-data-set/NM_mat.csv', index_col = 0)
dense_mat = dense_mat.values
NM_mat = NM_mat.values
missing_rate = 0.4
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros((dense_mat.shape[0], 28, 288))
for i1 in range(binary_tensor.shape[0]):
for i2 in range(binary_tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(NM_mat[i1, i2] + 0.5 - missing_rate)
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_tensor.reshape([dense_mat.shape[0], dense_mat.shape[1]]))
# -
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
init = {"W": 0.1 * np.random.rand(dim1, rank),
"X": 0.1 * np.random.rand(dim2, rank)}
maxiter1 = 1100
maxiter2 = 100
BPMF(dense_mat, sparse_mat, init, rank, maxiter1, maxiter2)
end = time.time()
print('Running time: %d seconds'%(end - start))
# **Experiment results** of missing data imputation using BPMF:
#
# | scenario |`rank`|`maxiter1`|`maxiter2`| mape | rmse |
# |:----------|-----:|---------:|---------:|-----------:|-----------:|
# |**20%, RM**| 50 | 1100 | 100 | **0.0651** | **4.0433** |
# |**40%, RM**| 50 | 1100 | 100 | **0.0703** | **4.2884** |
# |**20%, NM**| 10 | 1100 | 100 | **0.0912** | **5.2653** |
# |**40%, NM**| 10 | 1100 | 100 | **0.0919** | **5.3047** |
#
| experiments/Imputation-BPMF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # RNNs
# In this notebook you will learn how to build Recurrent Neural Networks (RNNs) for time series forecasting and sequence classification.
#
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/ageron/tf2_course/blob/master/06_rnns.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# </table>
# ## Imports
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import sklearn
import sys
import tensorflow as tf
from tensorflow import keras
import time
print("python", sys.version)
for module in mpl, np, pd, sklearn, tf, keras:
print(module.__name__, module.__version__)
assert sys.version_info >= (3, 5) # Python ≥3.5 required
assert tf.__version__ >= "2.0" # TensorFlow ≥2.0 required
# 
# # Exercise 1 – Time series forecasting
# ## 1.1) Load the data
# Let's start with a simple univariate time series: the daily temperatures in Melbourne from 1981 to 1990 ([source](https://datamarket.com/data/set/2324/daily-minimum-temperatures-in-melbourne-australia-1981-1990)).
# +
dataset_path = keras.utils.get_file(
"daily-minimum-temperatures-in-me.csv",
"https://raw.githubusercontent.com/ageron/tf2_course/master/datasets/daily-minimum-temperatures-in-me.csv")
temps = pd.read_csv(dataset_path,
parse_dates=[0], index_col=0)
# -
temps.info()
temps.head()
temps.plot(figsize=(10,5))
plt.show()
# ## 1.2) Prepare the data
# A few dates are missing, for example December 31st, 1984:
temps.loc["1984-12-29":"1985-01-02"]
# Let's ensure there's one row per day, filling missing values with the previous valid value:
temps = temps.asfreq("1D", method="ffill")
temps.loc["1984-12-29":"1985-01-02"]
# Alternatively, we could have interpolated using `temps.interpolate()`.
# ## 1.3) Add the shifted columns
# Next, let's create a function to add lag columns:
def add_lags(series, times):
cols = []
column_index = []
for time in times:
cols.append(series.shift(-time))
lag_fmt = "t+{time}" if time > 0 else "t{time}" if time < 0 else "t"
column_index += [(lag_fmt.format(time=time), col_name)
for col_name in series.columns]
df = pd.concat(cols, axis=1)
df.columns = pd.MultiIndex.from_tuples(column_index)
return df
# We will try to predict the temperature in 5 days (t+5) using the temperatures from the last 30 days (t-29 to t):
X = add_lags(temps, times=range(-30+1,1)).iloc[30:-5]
y = add_lags(temps, times=[5]).iloc[30:-5]
X.head()
y.head()
# Note: you may want to use `keras.preprocessing.sequence.TimeseriesGenerator` or `tf.data.Dataset.window()` instead.
# ## 1.4) Split the dataset
# Split this dataset into three periods: training (1981-1986), validation (1987-1988) and testing (1989-1990).
# +
#X_train, y_train = ...
#X_valid, y_valid = ...
#X_test, y_test = ...
# -
# ## 1.5) Reshape the inputs for the RNN
# Keras and TensorFlow expect a 3D NumPy array for any sequence. Its shape should be (number of instances, number of time steps, number of features per time step). Since this is a univariate time series, the last dimension is 1. Reshape the input features to get 3D arrays:
# +
#X_train_3D = ...
#X_valid_3D = ...
#X_test_3D = ...
# -
# ## 1.6) Build some baseline models
# Build some baseline models (at least one) and evaluate them on the validation set, using the Mean Absolute Error (MAE). For example:
#
# * a naive model, that just predicts the last known value.
# * an EMA model that predicts an exponential moving average of the last 48 hours (you can try to find the best span).
# * a linear model.
#
# Optional: plot the predictions.
# ## 1.7) Build a simple RNN
# Using Keras, build a simple 2-layer RNN with 100 neurons per layer, plus a dense layer with a single neuron. Train the model for 200 epochs with a batch size of 200, using Stochastic Gradient Descent with an learning rate of 0.005. Make sure to print the validation loss during training.
#
# Hints:
#
# * Create a `Sequential` model.
# * Add two `SimpleRNN` layers, with 100 units each. The first should return sequences but not the second. Indeed, in a Seq2Vec model, the last RNN layer should not return sequences. The first layer should specify the input shape (i.e., the shape of a single input sequence).
# * Use the MSE as the loss.
# * Call the model's `compile()` method, passing it an `SGD` instance with `lr=0.005`.
# * Call the model's `fit()` method, with the inputs and targets, number of epochs, batch size and validation data.
# +
#model1 = ...
# -
# ## 1.8) Plot the history
# Recall that you can simply use `pd.DataFrame(history.history).plot()`.
# ## 1.9) Evaluate the model
# Evaluate your RNN on the validation set, using the MAE. Try training your model again using the Huber loss and see if you get better performance.
def huber_loss(y_true, y_pred, max_grad=1.):
err = tf.abs(y_true - y_pred, name='abs')
mg = tf.constant(max_grad, name='max_grad')
lin = mg * (err - 0.5 * mg)
quad = 0.5 * err * err
return tf.where(err < mg, quad, lin)
# ## 1.10) Plot the predictions
# Make predictions on the validation set and plot them. Compare them to the targets and the baseline predictions.
# 
# # Exercise 1 – Solution
# ## 1.1) Load the data
# Let's start with a simple univariate time series: the daily temperatures in Melbourne from 1981 to 1990 ([source](https://datamarket.com/data/set/2324/daily-minimum-temperatures-in-melbourne-australia-1981-1990)).
# +
dataset_path = keras.utils.get_file(
"daily-minimum-temperatures-in-me.csv",
"https://raw.githubusercontent.com/ageron/tf2_course/master/datasets/daily-minimum-temperatures-in-me.csv")
temps = pd.read_csv(dataset_path,
parse_dates=[0], index_col=0)
# -
temps.info()
temps.head()
temps.plot(figsize=(10,5))
plt.show()
# ## 1.2) Prepare the data
# A few dates are missing, for example December 31st, 1984:
temps.loc["1984-12-29":"1985-01-02"]
# Let's ensure there's one row per day, filling missing values with the previous valid value:
temps = temps.asfreq("1D", method="ffill")
temps.loc["1984-12-29":"1985-01-02"]
# Alternatively, we could have interpolated using `temps.interpolate()`.
# ## 1.3) Add the shifted columns
# Next, let's create a function to add lag columns:
def add_lags(series, times):
cols = []
column_index = []
for time in times:
cols.append(series.shift(-time))
lag_fmt = "t+{time}" if time > 0 else "t{time}" if time < 0 else "t"
column_index += [(lag_fmt.format(time=time), col_name)
for col_name in series.columns]
df = pd.concat(cols, axis=1)
df.columns = pd.MultiIndex.from_tuples(column_index)
return df
add_lags(temps, times=(-2, -1, 0, +2)).head(10)
# We will try to predict the temperature in 5 days (t+5) using the temperatures from the last 30 days (t-29 to t):
X = add_lags(temps, times=range(-30+1,1)).iloc[30:-5]
y = add_lags(temps, times=[5]).iloc[30:-5]
X.head()
y.head()
# ## 1.4) Split the dataset
# Let's split this dataset into three periods: training, validation and testing:
train_slice = slice(None, "1986-12-25")
valid_slice = slice("1987-01-01", "1988-12-25")
test_slice = slice("1989-01-01", None)
X_train, y_train = X.loc[train_slice], y.loc[train_slice]
X_valid, y_valid = X.loc[valid_slice], y.loc[valid_slice]
X_test, y_test = X.loc[test_slice], y.loc[test_slice]
# ## 1.5) Reshape the inputs for the RNN
# Now let's create a function to reshape the multilevel DataFrames to 3D numpy arrays to feed to an RNN:
def multilevel_df_to_ndarray(df):
shape = [-1] + [len(level) for level in df.columns.remove_unused_levels().levels]
return df.values.reshape(shape)
X_train_3D = multilevel_df_to_ndarray(X_train)
X_valid_3D = multilevel_df_to_ndarray(X_valid)
X_test_3D = multilevel_df_to_ndarray(X_test)
X_train.shape
X_train_3D.shape
# ## 1.6) Build some baseline models
# Now let's evaluate some basic strategies, to get some baselines:
from sklearn.metrics import mean_absolute_error
def naive(X):
return X.iloc[:, -1]
y_pred_naive = naive(X_valid)
mean_absolute_error(y_valid, y_pred_naive)
def ema(X, span):
return X.T.ewm(span=span).mean().T.iloc[:, -1]
y_pred_ema = ema(X_valid, span=10)
mean_absolute_error(y_valid, y_pred_ema)
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(X_train, y_train)
y_pred_linear = lin_reg.predict(X_valid)
mean_absolute_error(y_valid, y_pred_linear)
# Let's plot these predictions:
def plot_predictions(*named_predictions, start=None, end=None, **kwargs):
day_range = slice(start, end)
plt.figure(figsize=(10,5))
for name, y_pred in named_predictions:
if hasattr(y_pred, "values"):
y_pred = y_pred.values
plt.plot(y_pred[day_range], label=name, **kwargs)
plt.legend()
plt.show()
plot_predictions(("Target", y_valid),
("Naive", y_pred_naive),
("EMA", y_pred_ema),
("Linear", y_pred_linear),
end=365)
# ## 1.7) Build a simple RNN
# Let's create a simple 2-layer RNN with 100 neurons per layer, plus a dense layer with a single neuron:
input_shape = X_train_3D.shape[1:]
input_shape
model1 = keras.models.Sequential()
model1.add(keras.layers.SimpleRNN(100, return_sequences=True, input_shape=input_shape))
model1.add(keras.layers.SimpleRNN(50))
model1.add(keras.layers.Dense(1))
model1.compile(loss="mse", optimizer=keras.optimizers.SGD(lr=0.005), metrics=["mae"])
history1 = model1.fit(X_train_3D, y_train, epochs=200, batch_size=200,
validation_data=(X_valid_3D, y_valid))
# ## 1.8) Plot the history
def plot_history(history, loss="loss"):
train_losses = history.history[loss]
valid_losses = history.history["val_" + loss]
n_epochs = len(history.epoch)
minloss = np.min(valid_losses)
plt.plot(train_losses, color="b", label="Train")
plt.plot(valid_losses, color="r", label="Validation")
plt.plot([0, n_epochs], [minloss, minloss], "k--",
label="Min val: {:.2f}".format(minloss))
plt.axis([0, n_epochs, 0, 20])
plt.legend()
plt.show()
plot_history(history1)
# ## 1.9) Evaluate the model
model1.evaluate(X_valid_3D, y_valid)
def huber_loss(y_true, y_pred, max_grad=1.):
err = tf.abs(y_true - y_pred, name='abs')
mg = tf.constant(max_grad, name='max_grad')
lin = mg * (err - 0.5 * mg)
quad = 0.5 * err * err
return tf.where(err < mg, quad, lin)
model1 = keras.models.Sequential()
model1.add(keras.layers.SimpleRNN(100, return_sequences=True, input_shape=input_shape))
model1.add(keras.layers.SimpleRNN(100))
model1.add(keras.layers.Dense(1))
model1.compile(loss=huber_loss, optimizer=keras.optimizers.SGD(lr=0.005), metrics=["mae"])
history1 = model1.fit(X_train_3D, y_train, epochs=200, batch_size=200,
validation_data=(X_valid_3D, y_valid))
model1.evaluate(X_valid_3D, y_valid)
# ## 1.10) Plot the predictions
y_pred_rnn1 = model1.predict(X_valid_3D)
plot_predictions(("Target", y_valid),
("Linear", y_pred_linear),
("RNN", y_pred_rnn1),
end=365)
# 
# # Exercise 2 – Forecasting the shifted sequence (Seq2Seq)
# Now let's predict temperatures for 30 days (from t-24 to t+5) instead of just one.
# ## 2.1) Define the 3D targets for training, validation and testing
# +
#Y_train_3D = ...
#Y_valid_3D = ...
#Y_test_3D = ...
# -
# ## 2.2) Define an `mae_last_step()` function
# For the final evaluation, we only want to look at the final time step (t+5). Create an `mae_last_step()` function that computes the MAE based on the final time step.
# ## 2.3) Build a Seq2Seq model
# Build a Seq2Seq model and compile it, using the Huber Loss, and using the last step MAE as the metric. Use SGD with a learning rate of 0.01. Hint: the layers are the same as earlier, except that the last RNN layer has `return_sequences=False`, and the `Dense` layer must be wrapped in a `keras.layers.TimeDistributed` layer.
# ## 2.4) Train the model
# Fit the model as earlier (but with the 3D targets). Again, evaluate the model and plot the predictions.
# 
# # Exercise 2 – Solution
# ## 2.1) Define the 3D targets for training, validation and testing
Y = add_lags(temps, times=range(-24, 5+1)).iloc[30:-5]
Y_train = Y.loc[train_slice]
Y_valid = Y.loc[valid_slice]
Y_test = Y.loc[test_slice]
Y_train_3D = multilevel_df_to_ndarray(Y_train)
Y_valid_3D = multilevel_df_to_ndarray(Y_valid)
Y_test_3D = multilevel_df_to_ndarray(Y_test)
# ## 2.2) Define an `mae_last_step()` function
# For the final evaluation, we only want to look at the final time step (t+5):
# +
K = keras.backend
def mae_last_step(Y_true, Y_pred):
return K.mean(K.abs(Y_pred[:, -1] - Y_true[:, -1]))
# -
# ## 2.3) Build a Seq2Seq model
model2 = keras.models.Sequential()
model2.add(keras.layers.SimpleRNN(100, return_sequences=True, input_shape=input_shape))
model2.add(keras.layers.SimpleRNN(100, return_sequences=True))
model2.add(keras.layers.TimeDistributed(keras.layers.Dense(1)))
model2.compile(loss=huber_loss, optimizer=keras.optimizers.SGD(lr=0.01),
metrics=[mae_last_step])
# ## 2.4) Train the model
history2 = model2.fit(X_train_3D, Y_train_3D, epochs=200, batch_size=200,
validation_data=(X_valid_3D, Y_valid_3D))
plot_history(history2, loss="mae_last_step")
model2.evaluate(X_valid_3D, Y_valid_3D)
y_pred_rnn2 = model2.predict(X_valid_3D)[:, -1]
plot_predictions(("Target", y_valid),
("Linear", y_pred_linear),
("RNN", y_pred_rnn2),
end=365)
# 
# # Exercise 3 – LSTM and GRU
# ## 3.1) Build, train and evaluate a Seq2Seq LSTM
# Train the same model as earlier but using `LSTM` or `GRU` instead of `SimpleRNN`. You can also try reducing the learning rate when the validation loss reaches a plateau, using the `ReduceLROnPlateau` callback.
# ## 3.2) Add $\ell_2$ regularization
# Add $\ell_2$ regularization to your RNN, using the layers' `kernel_regularizer` and `recurrent_regularizer` arguments, and the `l2()` function in `keras.regularizers`. Tip: use the `partial()` function in the `functools` package to avoid repeating the same arguments again and again.
# 
# # Exercise 3 – Solution
# ## 3.1) Build, train and evaluate a Seq2Seq LSTM
# You can try replacing `LSTM` with `GRU`.
model3 = keras.models.Sequential()
model3.add(keras.layers.LSTM(100, return_sequences=True, input_shape=input_shape))
model3.add(keras.layers.LSTM(100, return_sequences=True))
model3.add(keras.layers.TimeDistributed(keras.layers.Dense(1)))
model3.compile(loss=huber_loss, optimizer=keras.optimizers.SGD(lr=0.01),
metrics=[mae_last_step])
history3 = model3.fit(X_train_3D, Y_train_3D, epochs=200, batch_size=200,
validation_data=(X_valid_3D, Y_valid_3D),
callbacks=[keras.callbacks.ReduceLROnPlateau(verbose=1)])
model3.evaluate(X_valid_3D, Y_valid_3D)
plot_history(history3, loss="mae_last_step")
y_pred_rnn3 = model3.predict(X_valid_3D)[:, -1]
plot_predictions(("Target", y_valid),
("Linear", y_pred_linear),
("RNN", y_pred_rnn3),
end=365)
# ## 3.2) Add $\ell_2$ regularization
from functools import partial
RegularizedLSTM = partial(keras.layers.LSTM,
return_sequences=True,
kernel_regularizer=keras.regularizers.l2(1e-4),
recurrent_regularizer=keras.regularizers.l2(1e-4))
model3 = keras.models.Sequential()
model3.add(RegularizedLSTM(100, input_shape=input_shape))
model3.add(RegularizedLSTM(100))
model3.add(keras.layers.Dense(1))
model3.compile(loss=huber_loss, optimizer=keras.optimizers.SGD(lr=0.01),
metrics=[mae_last_step])
history3 = model3.fit(X_train_3D, Y_train_3D, epochs=200, batch_size=100,
validation_data=(X_valid_3D, Y_valid_3D))
model3.evaluate(X_valid_3D, Y_valid_3D)
plot_history(history3)
y_pred_rnn3 = model3.predict(X_valid_3D)[:, -1]
plot_predictions(("Target", y_valid),
("Linear", y_pred_linear),
("RNN", y_pred_rnn3),
end=365)
# 
# # Exercise 4 – Preprocessing with 1D-ConvNets
# At the beginning of your sequential model, add a `Conv1D` layer with 32 kernels of size 5, a `MaxPool1D` layer with pool size 5 and strides 2. Train and evaluate the model.
# 
# # Exercise 4 – Solution
model4 = keras.models.Sequential()
model4.add(keras.layers.Conv1D(32, kernel_size=5, input_shape=input_shape))
model4.add(keras.layers.MaxPool1D(pool_size=5, strides=2))
model4.add(keras.layers.LSTM(32, return_sequences=True))
model4.add(keras.layers.LSTM(32))
model4.add(keras.layers.Dense(1))
model4.compile(loss=huber_loss, optimizer=keras.optimizers.SGD(lr=0.005))
model4.summary()
history4 = model4.fit(X_train_3D, y_train, epochs=200, batch_size=100,
validation_data=(X_valid_3D, y_valid))
model4.evaluate(X_valid_3D, y_valid)
# 
# # Exercice 5 – Sequence classification
# Let's load the IMDB movie reviews, for binary sentiment analysis (positive review or negative review):
# We only want the 10,000 most common words:
num_words = 10000
(X_train, y_train), (X_test, y_test) = keras.datasets.imdb.load_data(num_words=num_words)
# Let's also get the word index (word to word id):
word_index = keras.datasets.imdb.get_word_index()
# And let's create a reverse index (word id to word). Three special word ids are added:
reverse_index = {word_id + 3: word for word, word_id in word_index.items()}
reverse_index[0] = "<pad>" # padding
reverse_index[1] = "<sos>" # start of sequence
reverse_index[2] = "<oov>" # out-of-vocabulary
reverse_index[3] = "<unk>" # unknown
# Let's write a little function to decode reviews:
def decode_review(word_ids):
return " ".join([reverse_index.get(word_id, "<err>") for word_id in word_ids])
# Let's look at a review:
decode_review(X_train[0])
# It seems very positive, let's look at the target (0=negative review, 1=positive review):
y_train[0]
# And another review:
decode_review(X_train[1])
# Very negative! Let's check the target:
y_train[1]
# ## 5.1) Train a baseline model
# Train and evaluate a baseline model using ScikitLearn. You will need to create a pipeline with a `CountVectorizer`, a `TfidfTransformer` and an `SGDClassifier`. The `CountVectorizer` transformer expects text as input, so let's create a text version of the training set and test set:
X_train_text = [decode_review(words_ids) for words_ids in X_train]
X_test_text = [decode_review(words_ids) for words_ids in X_test]
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.pipeline import Pipeline
from sklearn.linear_model import SGDClassifier
# ## 5.2) Create a sequence classifier
# Create a sequence classifier using Keras:
# * Use `keras.preprocessing.sequence.pad_sequences()` to preprocess `X_train`: this will create a 2D array of 25,000 rows (one per review) and `maxlen=500` columns. Reviews longer than 500 words will be cropped, while reviews shorter
# than 500 words will be padded with zeros.
# * The first layer in your model should be an `Embedding` layer, with `input_dim=num_words` and `output_dim=10`. The model will gradually learn to represent each of the 10,000 words as a 10-dimensional vector. So the next layer will receive 3D batchs of shape (batch size, 500, 10).
# * Add one or more LSTM layers with 32 neurons each.
# * The output layer should be a Dense layer with a sigmoid activation function, since this is a binary classification problem.
# * When compiling the model, you should use the `binary_crossentropy` loss.
# * Fit the model for 10 epochs, using a batch size of 128 and `validation_split=0.2`.
# 
# # Exercice 5 – Solution
# ## 5.1) Train a baseline model
X_train_text = [decode_review(words_ids) for words_ids in X_train]
X_test_text = [decode_review(words_ids) for words_ids in X_test]
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.pipeline import Pipeline
from sklearn.linear_model import SGDClassifier
pipeline = Pipeline([
('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', SGDClassifier(max_iter=50)),
])
pipeline.fit(X_train_text, y_train)
pipeline.score(X_test_text, y_test)
# We get 88.5% accuracy, that's not too bad. But don't forget to check the ratio of positive reviews:
y_test.mean()
# Let's try our model:
pipeline.predict(["this movie was really awesome"])
# ## 5.2) Create a sequence classifier
maxlen = 500
X_train_trim = keras.preprocessing.sequence.pad_sequences(X_train, maxlen=maxlen)
X_test_trim = keras.preprocessing.sequence.pad_sequences(X_test, maxlen=maxlen)
model = keras.models.Sequential()
model.add(keras.layers.Embedding(input_dim=num_words, output_dim=10))
model.add(keras.layers.LSTM(32))
model.add(keras.layers.Dense(1, activation="sigmoid"))
model.compile(loss="binary_crossentropy", optimizer="rmsprop", metrics=["accuracy"])
model.summary()
history = model.fit(X_train_trim, y_train,
epochs=10, batch_size=128, validation_split=0.2)
model.evaluate(X_test_trim, y_test)
# 
# # Exercise 6 – Bidirectional RNN
# Update the previous sequence classification model to use a bidirectional LSTM. For this, you just need to wrap the LSTM layer in a `Bidirectional` layer. If the model overfits, try adding a dropout layer.
# 
# # Exercise 6 – Solution
model = keras.models.Sequential()
model.add(keras.layers.Embedding(input_dim=num_words, output_dim=10))
model.add(keras.layers.Dropout(0.5))
model.add(keras.layers.Bidirectional(keras.layers.LSTM(32)))
model.add(keras.layers.Dense(1, activation="sigmoid"))
model.compile(loss="binary_crossentropy", optimizer="rmsprop", metrics=["accuracy"])
history = model.fit(X_train_trim, y_train,
epochs=10, batch_size=128, validation_split=0.2)
model.evaluate(X_test_trim, y_test)
| 06_rnns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="RUymE2l9GZfO"
# **Copyright 2019 The TensorFlow Hub Authors.**
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + cellView="code" id="JMyTNwSJGGWg"
# Copyright 2019 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] id="co7MV6sX7Xto"
# # Multilingual Universal Sentence Encoder를 사용한 교차 언어 유사성 및 의미론적 검색 엔진
#
# + [markdown] id="MfBg1C5NB3X0"
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/hub/tutorials/cross_lingual_similarity_with_tf_hub_multilingual_universal_encoder"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">}TensorFlow.org에서 보기</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/hub/tutorials/cross_lingual_similarity_with_tf_hub_multilingual_universal_encoder.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab에서 실행하기</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/hub/tutorials/cross_lingual_similarity_with_tf_hub_multilingual_universal_encoder.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub에서 소스 보기</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/hub/tutorials/cross_lingual_similarity_with_tf_hub_multilingual_universal_encoder.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">노트북 다운로드하기</a></td>
# </table>
# + [markdown] id="eAVQGidpL8v5"
# 이 노트북은 Multilingual Universal Sentence Encoder 모듈에 액세스하고 이 모듈을 다국어 문장 유사성에 사용하는 방법을 보여줍니다. 이 모듈은 [원래 Universal Encoder 모듈](https://tfhub.dev/google/universal-sentence-encoder/2)의 확장입니다.
#
# 이 노트북의 내용은 다음과 같이 구성됩니다.
#
# - 첫 번째 섹션에서는 언어 쌍 사이의 문장 시각화를 보여줍니다. 다소 학문적인 내용입니다.
# - 두 번째 섹션에서는 여러 언어로 된 Wikipedia Corpus 샘플에서 의미론적 검색 엔진을 빌드하는 방법을 보여줍니다.
# + [markdown] id="UvNRbHGarYeR"
# ## 인용
#
# *이 colab에서 탐구한 모델을 사용하는 연구 논문에는 다음 인용구를 넣어야 합니다.*
#
# ###
#
# [의미론적 검색을 위한 다국어 범용 문장 인코더](https://arxiv.org/abs/1907.04307)
#
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. 2019. arXiv preprint arXiv:1907.04307
# + [markdown] id="pOTzp8O36CyQ"
# ## 설정
#
# 이 섹션에서는 Multilingual Universal Sentence Encoder 모듈에 액세스할 수 있는 환경을 설정하고 일련의 영어 문장과 해당 번역을 준비합니다. 다음 섹션에서는 다국어 모듈을 사용하여 *교차 언어* 유사성을 계산합니다.
# + cellView="both" id="lVjNK8shFKOC"
# %%capture
#@title Setup Environment
# Install the latest Tensorflow version.
# !pip install tensorflow_text
# !pip install bokeh
# !pip install simpleneighbors[annoy]
# !pip install tqdm
# + cellView="both" id="MSeY-MUQo2Ha"
#@title Setup common imports and functions
import bokeh
import bokeh.models
import bokeh.plotting
import numpy as np
import os
import pandas as pd
import tensorflow.compat.v2 as tf
import tensorflow_hub as hub
from tensorflow_text import SentencepieceTokenizer
import sklearn.metrics.pairwise
from simpleneighbors import SimpleNeighbors
from tqdm import tqdm
from tqdm import trange
def visualize_similarity(embeddings_1, embeddings_2, labels_1, labels_2,
plot_title,
plot_width=1200, plot_height=600,
xaxis_font_size='12pt', yaxis_font_size='12pt'):
assert len(embeddings_1) == len(labels_1)
assert len(embeddings_2) == len(labels_2)
# arccos based text similarity (Yang et al. 2019; Cer et al. 2019)
sim = 1 - np.arccos(
sklearn.metrics.pairwise.cosine_similarity(embeddings_1,
embeddings_2))/np.pi
embeddings_1_col, embeddings_2_col, sim_col = [], [], []
for i in range(len(embeddings_1)):
for j in range(len(embeddings_2)):
embeddings_1_col.append(labels_1[i])
embeddings_2_col.append(labels_2[j])
sim_col.append(sim[i][j])
df = pd.DataFrame(zip(embeddings_1_col, embeddings_2_col, sim_col),
columns=['embeddings_1', 'embeddings_2', 'sim'])
mapper = bokeh.models.LinearColorMapper(
palette=[*reversed(bokeh.palettes.YlOrRd[9])], low=df.sim.min(),
high=df.sim.max())
p = bokeh.plotting.figure(title=plot_title, x_range=labels_1,
x_axis_location="above",
y_range=[*reversed(labels_2)],
plot_width=plot_width, plot_height=plot_height,
tools="save",toolbar_location='below', tooltips=[
('pair', '@embeddings_1 ||| @embeddings_2'),
('sim', '@sim')])
p.rect(x="embeddings_1", y="embeddings_2", width=1, height=1, source=df,
fill_color={'field': 'sim', 'transform': mapper}, line_color=None)
p.title.text_font_size = '12pt'
p.axis.axis_line_color = None
p.axis.major_tick_line_color = None
p.axis.major_label_standoff = 16
p.xaxis.major_label_text_font_size = xaxis_font_size
p.xaxis.major_label_orientation = 0.25 * np.pi
p.yaxis.major_label_text_font_size = yaxis_font_size
p.min_border_right = 300
bokeh.io.output_notebook()
bokeh.io.show(p)
# + [markdown] id="gk2IRjZFGDsK"
# 다음은 이 노트북 전체에서 텍스트를 인코딩하는 데 사용할 사전 훈련된 ML 모델을 가져오는 추가 상용구 코드입니다.
# + id="mkmF3w8WGLcM"
# The 16-language multilingual module is the default but feel free
# to pick others from the list and compare the results.
module_url = 'https://tfhub.dev/google/universal-sentence-encoder-multilingual/3' #@param ['https://tfhub.dev/google/universal-sentence-encoder-multilingual/3', 'https://tfhub.dev/google/universal-sentence-encoder-multilingual-large/3']
model = hub.load(module_url)
def embed_text(input):
return model(input)
# + [markdown] id="jhLPq6AROyFk"
# # 언어 간 텍스트 유사성 시각화하기
#
# 이제 문장 임베딩이 준비되었으므로 다양한 언어 사이에서 의미론적 유사성을 시각화할 수 있습니다.
# + [markdown] id="8xdAogbxJDTD"
# ## 텍스트 임베딩 계산하기
#
# 먼저 다양한 언어로 번역된 일련의 문장을 병렬로 정의합니다. 그런 다음 모든 문장에 대한 임베딩을 미리 계산합니다.
# + id="Q8F4LNGFqOiq"
# Some texts of different lengths in different languages.
arabic_sentences = ['كلب', 'الجراء لطيفة.', 'أستمتع بالمشي لمسافات طويلة على طول الشاطئ مع كلبي.']
chinese_sentences = ['狗', '小狗很好。', '我喜欢和我的狗一起沿着海滩散步。']
english_sentences = ['dog', 'Puppies are nice.', 'I enjoy taking long walks along the beach with my dog.']
french_sentences = ['chien', 'Les chiots sont gentils.', 'J\'aime faire de longues promenades sur la plage avec mon chien.']
german_sentences = ['Hund', 'Welpen sind nett.', 'Ich genieße lange Spaziergänge am Strand entlang mit meinem Hund.']
italian_sentences = ['cane', 'I cuccioli sono carini.', 'Mi piace fare lunghe passeggiate lungo la spiaggia con il mio cane.']
japanese_sentences = ['犬', '子犬はいいです', '私は犬と一緒にビーチを散歩するのが好きです']
korean_sentences = ['개', '강아지가 좋다.', '나는 나의 산책을 해변을 따라 길게 산책하는 것을 즐긴다.']
russian_sentences = ['собака', 'Милые щенки.', 'Мне нравится подолгу гулять по пляжу со своей собакой.']
spanish_sentences = ['perro', 'Los cachorros son agradables.', 'Disfruto de dar largos paseos por la playa con mi perro.']
# Multilingual example
multilingual_example = ["Willkommen zu einfachen, aber", "verrassend krachtige", "multilingüe", "compréhension du langage naturel", "модели.", "大家是什么意思" , "보다 중요한", ".اللغة التي يتحدثونها"]
multilingual_example_in_en = ["Welcome to simple yet", "surprisingly powerful", "multilingual", "natural language understanding", "models.", "What people mean", "matters more than", "the language they speak."]
# + id="weXZqLtTJY9b"
# Compute embeddings.
ar_result = embed_text(arabic_sentences)
en_result = embed_text(english_sentences)
es_result = embed_text(spanish_sentences)
de_result = embed_text(german_sentences)
fr_result = embed_text(french_sentences)
it_result = embed_text(italian_sentences)
ja_result = embed_text(japanese_sentences)
ko_result = embed_text(korean_sentences)
ru_result = embed_text(russian_sentences)
zh_result = embed_text(chinese_sentences)
multilingual_result = embed_text(multilingual_example)
multilingual_in_en_result = embed_text(multilingual_example_in_en)
# + [markdown] id="_3zGWuF-GhUm"
# ## 유사성 시각화하기
#
# 텍스트 임베딩이 준비되었으므로 내적을 사용하여 언어 간에 문장이 얼마나 유사한지 시각화할 수 있습니다. 어두운 색은 임베딩이 의미상 유사함을 나타냅니다.
# + [markdown] id="WOEIJA0mh70g"
# ### 다국어 유사성
# + id="R2hbCMhmiDWR"
visualize_similarity(multilingual_in_en_result, multilingual_result,
multilingual_example_in_en, multilingual_example, "Multilingual Universal Sentence Encoder for Semantic Retrieval (Yang et al., 2019)")
# + [markdown] id="h3TEhllsq3ax"
# ### 영어-아랍어 유사성
# + id="Q9UDpStmq7Ii"
visualize_similarity(en_result, ar_result, english_sentences, arabic_sentences, 'English-Arabic Similarity')
# + [markdown] id="QF9z48HMp4WL"
# ### 영어-러시아어 유사성
# + id="QE68UejYp86z"
visualize_similarity(en_result, ru_result, english_sentences, russian_sentences, 'English-Russian Similarity')
# + [markdown] id="BJkL6Az0QXNN"
# ### 영어-스페인어 유사성
# + id="CH_BXVGhQ0GL"
visualize_similarity(en_result, es_result, english_sentences, spanish_sentences, 'English-Spanish Similarity')
# + [markdown] id="imn28LCiQO7d"
# ### 영어-이탈리아어 유사성
# + id="X9uD3DirPIGd"
visualize_similarity(en_result, it_result, english_sentences, italian_sentences, 'English-Italian Similarity')
# + [markdown] id="m6ySvEGbQaTM"
# ### 이탈리아어-스페인어 유사성
# + id="irfwIeitQ7V6"
visualize_similarity(it_result, es_result, italian_sentences, spanish_sentences, 'Italian-Spanish Similarity')
# + [markdown] id="ueoRO8balwwr"
# ### 영어-중국어 유사성
# + id="xA7anofVlxL7"
visualize_similarity(en_result, zh_result, english_sentences, chinese_sentences, 'English-Chinese Similarity')
# + [markdown] id="8zV1BJc3mL3W"
# ### 영어-한국어 유사성
# + id="iqWy1e1UmQeX"
visualize_similarity(en_result, ko_result, english_sentences, korean_sentences, 'English-Korean Similarity')
# + [markdown] id="dfTj-JaunFTv"
# ### 중국어-한국어 유사성
# + id="MndSgKGPnJuF"
visualize_similarity(zh_result, ko_result, chinese_sentences, korean_sentences, 'Chinese-Korean Similarity')
# + [markdown] id="rRabHHQYQfLr"
# ### 기타 언어
#
# 위의 예는 **영어, 아랍어, 중국어, 네덜란드어, 프랑스어, 독일어, 이탈리아어, 일본어, 한국어, 폴란드어, 포르투갈어, 러시아어, 스페인어, 태국어 및 터키어**의 모든 언어 쌍으로 확장할 수 있습니다. 즐거운 코딩되세요!
# + [markdown] id="mxAFAJI9xsAU"
# # 다국어 의미론적 유사성 검색 엔진 만들기
#
# 이전 예제에서는 몇 개의 문장을 시각화했지만 이 섹션에서는 Wikipedia Corpus에서 약 200,000개의 문장에 대한 의미론적 검색 인덱스를 빌드합니다. Universal Sentence Encoder의 다국어 기능을 보여주기 위해 약 절반은 영어로, 나머지 절반은 스페인어로 제공됩니다.
#
# ## 인덱스로 데이터 다운로드하기
#
# 먼저 [News Commentary Corpus](http://opus.nlpl.eu/News-Commentary-v11.php) [1]에서 여러 언어로 된 뉴스 문장을 다운로드합니다. 이 접근 방식은 일반성을 잃지 않고, 지원되는 나머지 언어의 인덱싱에도 효과적으로 적용됩니다.
#
# 데모의 속도를 높이기 위해 언어당 문장을 1000개로 제한합니다.
# + id="587I9ye6yXEU"
corpus_metadata = [
('ar', 'ar-en.txt.zip', 'News-Commentary.ar-en.ar', 'Arabic'),
('zh', 'en-zh.txt.zip', 'News-Commentary.en-zh.zh', 'Chinese'),
('en', 'en-es.txt.zip', 'News-Commentary.en-es.en', 'English'),
('ru', 'en-ru.txt.zip', 'News-Commentary.en-ru.ru', 'Russian'),
('es', 'en-es.txt.zip', 'News-Commentary.en-es.es', 'Spanish'),
]
language_to_sentences = {}
language_to_news_path = {}
for language_code, zip_file, news_file, language_name in corpus_metadata:
zip_path = tf.keras.utils.get_file(
fname=zip_file,
origin='http://opus.nlpl.eu/download.php?f=News-Commentary/v11/moses/' + zip_file,
extract=True)
news_path = os.path.join(os.path.dirname(zip_path), news_file)
language_to_sentences[language_code] = pd.read_csv(news_path, sep='\t', header=None)[0][:1000]
language_to_news_path[language_code] = news_path
print('{:,} {} sentences'.format(len(language_to_sentences[language_code]), language_name))
# + [markdown] id="m3DIT9uT7Z34"
# ## 사전 훈련된 모델을 사용하여 문장을 벡터로 변환하기
#
# GPU의 RAM에 맞도록 여러 *배치*로 임베딩을 계산합니다.
# + id="yRoRT5qCEIYy"
# Takes about 3 minutes
batch_size = 2048
language_to_embeddings = {}
for language_code, zip_file, news_file, language_name in corpus_metadata:
print('\nComputing {} embeddings'.format(language_name))
with tqdm(total=len(language_to_sentences[language_code])) as pbar:
for batch in pd.read_csv(language_to_news_path[language_code], sep='\t',header=None, chunksize=batch_size):
language_to_embeddings.setdefault(language_code, []).extend(embed_text(batch[0]))
pbar.update(len(batch))
# + [markdown] id="oeBqoE8e-scg"
# ## 의미론적 벡터 인덱스 빌드하기
#
# [Annoy](https://pypi.org/project/simpleneighbors/) 라이브러리의 래퍼인 [SimpleNeighbors](https://github.com/spotify/annoy) 라이브러리를 사용하여 Corpus에서 결과를 효율적으로 조회합니다.
# + id="lv_SOduAF1oi"
# %%time
# Takes about 8 minutes
num_index_trees = 40
language_name_to_index = {}
embedding_dimensions = len(list(language_to_embeddings.values())[0][0])
for language_code, zip_file, news_file, language_name in corpus_metadata:
print('\nAdding {} embeddings to index'.format(language_name))
index = SimpleNeighbors(embedding_dimensions, metric='dot')
for i in trange(len(language_to_sentences[language_code])):
index.add_one(language_to_sentences[language_code][i], language_to_embeddings[language_code][i])
print('Building {} index with {} trees...'.format(language_name, num_index_trees))
index.build(n=num_index_trees)
language_name_to_index[language_name] = index
# + id="0aqGwIuLGrtu"
# %%time
# Takes about 13 minutes
num_index_trees = 60
print('Computing mixed-language index')
combined_index = SimpleNeighbors(embedding_dimensions, metric='dot')
for language_code, zip_file, news_file, language_name in corpus_metadata:
print('Adding {} embeddings to mixed-language index'.format(language_name))
for i in trange(len(language_to_sentences[language_code])):
annotated_sentence = '({}) {}'.format(language_name, language_to_sentences[language_code][i])
combined_index.add_one(annotated_sentence, language_to_embeddings[language_code][i])
print('Building mixed-language index with {} trees...'.format(num_index_trees))
combined_index.build(n=num_index_trees)
# + [markdown] id="kg9cw0S2_ntQ"
# ## 의미론적 유사성 검색 엔진이 동작하는지 확인하기
#
# 이 섹션에서는 다음을 시연합니다.
#
# 1. 의미론적 검색 기능: Corpus에서 주어진 쿼리와 의미적으로 유사한 문장을 검색합니다.
# 2. 다국어 기능: 쿼리 언어와 인덱스 언어가 일치하면 여러 언어에서도 작업을 수행합니다.
# 3. 교차 언어 기능: 인덱싱된 Corpus와 다른 언어로 쿼리를 실행합니다.
# 4. 혼합 언어 Corpus: 모든 언어의 항목을 포함하는 단일 인덱스에 대해 위의 모든 항목을 수행합니다.
#
# + [markdown] id="Dxu66S8wJIG9"
# ### 의미론적 검색 교차 언어 기능
#
# 이 섹션에서는 샘플 영어 문장 세트와 관련된 문장을 검색하는 방법을 보여줍니다. 다음을 시도합니다.
#
# - 몇 가지 다른 샘플 문장을 시도합니다.
# - 반환된 결과의 수를 변경해 봅니다(유사성 순서로 반환됨).
# - 여러 언어로 결과를 반환하여 다국어 기능을 사용해 봅니다(일부 결과에 [구글 번역](http://translate.google.com)을 사용하여 모국어의 온전성을 확인할 수 있음).
#
# + cellView="form" id="_EFSd65B_mq8"
sample_query = 'The stock market fell four points.' #@param ["Global warming", "Researchers made a surprising new discovery last week.", "The stock market fell four points.", "Lawmakers will vote on the proposal tomorrow."] {allow-input: true}
index_language = 'English' #@param ["Arabic", "Chinese", "English", "French", "German", "Russian", "Spanish"]
num_results = 10 #@param {type:"slider", min:0, max:100, step:10}
query_embedding = embed_text(sample_query)[0]
search_results = language_name_to_index[index_language].nearest(query_embedding, n=num_results)
print('{} sentences similar to: "{}"\n'.format(index_language, sample_query))
search_results
# + [markdown] id="Ybgj9o7hKDZV"
# ### 혼합 코퍼스 기능
#
# 이제 영어로 쿼리를 제출하지만 결과는 인덱싱된 언어 중 하나에서 나옵니다.
# + cellView="form" id="MJeTzuj0KU41"
sample_query = 'The stock market fell four points.' #@param ["Global warming", "Researchers made a surprising new discovery last week.", "The stock market fell four points.", "Lawmakers will vote on the proposal tomorrow."] {allow-input: true}
num_results = 40 #@param {type:"slider", min:0, max:100, step:10}
query_embedding = embed_text(sample_query)[0]
search_results = language_name_to_index[index_language].nearest(query_embedding, n=num_results)
print('{} sentences similar to: "{}"\n'.format(index_language, sample_query))
search_results
# + [markdown] id="tqIRtHIL2jAw"
# 자신의 쿼리를 시도합니다.
# + cellView="form" id="ZK5ID6XF2n8k"
query = 'The stock market fell four points.' #@param {type:"string"}
num_results = 30 #@param {type:"slider", min:0, max:100, step:10}
query_embedding = embed_text(sample_query)[0]
search_results = combined_index.nearest(query_embedding, n=num_results)
print('{} sentences similar to: "{}"\n'.format(index_language, query))
search_results
# + [markdown] id="IPPwnhUNdOfc"
# # 추가 주제
#
# ## 다국어
#
# 마지막으로, 지원되는 언어(**영어, 아랍어, 중국어, 네덜란드어, 프랑스어, 독일어, 이탈리아어, 일본어, 한국어, 폴란드어, 포르투갈어, 러시아어, 스페인어, 태국어 및 터키어**로 쿼리를 시도해 볼 것을 권장합니다.
#
# 또한 여기서는 일부 언어로만 인덱싱했지만 지원되는 모든 언어로 콘텐츠를 인덱싱할 수도 있습니다.
#
# ## 모델 변형
#
# 메모리, 대기 시간 및/또는 품질과 같은 다양한 요소에 최적화된 다양한 Universal Encoder 모델이 제공됩니다. 자유롭게 시도해 보고 적절한 모델을 찾으세요.
#
# ## NN(Nearest neighbor) 라이브러리
#
# Annoy를 사용하여 NN을 효율적으로 검색했습니다. 트리 수(메모리 종속) 및 검색할 항목 수(대기 시간 종속)에 대해서는 [상충 관계 섹션](https://github.com/spotify/annoy/blob/master/README.rst#tradeoffs)을 참조하세요. SimpleNeighbors에서는 트리 수만 제어할 수 있지만 Annoy를 직접 사용하도록 코드를 간단하게 리팩토링할 수 있습니다. 일반적인 사용자를 위해 이 코드를 최대한 간단하게 유지하려고 했습니다.
#
# 해당 애플리케이션에 맞게 Annoy의 크기를 조정할 수 없으면 [FAISS](https://github.com/facebookresearch/faiss)도 확인해 보세요.
# + [markdown] id="5yj9VcfnbS-q"
# *멋진 다국어 의미론 애플리케이션을 빌드해 보세요!*
# + [markdown] id="X4oOfvSOKnjS"
# [1] <NAME>, 2012, [Parallel Data, Tools and Interfaces in OPUS](http://www.lrec-conf.org/proceedings/lrec2012/pdf/463_Paper.pdf). In Proceedings of the 8th International Conference on Language Resources and Evaluation (LREC 2012)
| site/ko/hub/tutorials/cross_lingual_similarity_with_tf_hub_multilingual_universal_encoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Qns9DRsxLdK_"
import numpy as np
import cv2
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
image = mpimg.imread('images/bridge_shadow.jpg')
# Edit this function to create your own pipeline.
def pipeline(img, s_thresh=(170, 255), sx_thresh=(20, 100)):
img = np.copy(img)
# Convert to HLS color space and separate the V channel
hls = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
l_channel = hls[:,:,1]
s_channel = hls[:,:,2]
# Sobel x
sobelx = cv2.Sobel(l_channel, cv2.CV_64F, 1, 0) # Take the derivative in x
abs_sobelx = np.absolute(sobelx) # Absolute x derivative to accentuate lines away from horizontal
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
# Threshold x gradient
sxbinary = np.zeros_like(scaled_sobel)
sxbinary[(scaled_sobel >= sx_thresh[0]) & (scaled_sobel <= sx_thresh[1])] = 1
# Threshold color channel
s_binary = np.zeros_like(s_channel)
s_binary[(s_channel >= s_thresh[0]) & (s_channel <= s_thresh[1])] = 1
# Stack each channel
color_binary = np.dstack(( np.zeros_like(sxbinary), sxbinary, s_binary)) * 255
return color_binary
result = pipeline(image)
# Plot the result
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
f.tight_layout()
ax1.imshow(image)
ax1.set_title('Original Image', fontsize=40)
ax2.imshow(result)
ax2.set_title('Pipeline Result', fontsize=40)
plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
| src/E2-CombineThresholds/CombiningColorGradientThresholds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import graphistry
graphistry.register(api=1, protocol="https", server="hub.graphistry.com", key="4f55eede43c4020fbe604f4bb159737568e7e890c202fc943f2164dd1bdc4adf")
# +
# load data
pd_data = pd.read_csv('./kg_final_with_temporal_data_and_validated_inconsistencies.txt', sep='\t')
pd_data = pd_data[['Subject', 'Predicate', 'Object']]
# filter positives only
neg_predicates = [
'confers no resistance to antibiotic',
'not upregulated by antibiotic',
'no represses',
'no activates',]
def _check_match(x, predicates):
flag = False
for predicate in predicates:
if predicate in x:
flag = True
return flag
pd_pos_data = pd_data[~pd_data['Predicate'].apply(lambda x: _check_match(x, neg_predicates))]
# assign predicate category
def map_func(label):
if 'confers resistance to antibiotic' in label:
return 'confers resistance to antibiotic'
elif 'upregulated by antibiotic' in label:
return 'upregulated by antibiotic'
elif 'represses' in label:
return 'represses'
elif 'activates' in label:
return 'activates'
elif 'has' in label:
return 'has'
elif 'is involved in' in label:
return 'is involved in'
elif 'is part of' in label:
return 'is part of'
elif 'targeted by' in label:
return 'targeted by'
else:
raise ValueError('Invalid label: {}'.format(label))
pd_pos_data['Predicate Category'] = pd_pos_data['Predicate'].apply(lambda x: map_func(x))
# +
g = graphistry.bind(
source='Subject',
destination='Object',
edge_label='Predicate',)
g.plot(pd_pos_data)
# -
| manuscript_preparation/kg_visualization/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
from torch.utils.data import DataLoader
from torch.utils.data.dataset import Dataset
from torchvision import datasets, transforms
# # Squares
class SquareDataset(Dataset):
def __init__(self, size):
self.size = size
self.X = torch.randint(255, (size, 9), dtype=torch.float)
real_w = torch.tensor([[1,1,1,0,0,0,0,0,0],
[0,0,0,1,1,1,0,0,0],
[0,0,0,0,0,0,1,1,1]],
dtype=torch.float)
y = torch.argmax(self.X.mm(real_w.t()), 1)
self.Y = torch.zeros(size, 3, dtype=torch.float) \
.scatter_(1, y.view(-1, 1), 1)
def __getitem__(self, index):
return (self.X[index], self.Y[index])
def __len__(self):
return self.size
squares = SquareDataset(256)
print(squares[34])
print(squares[254])
print(squares[25])
# +
dataloader = DataLoader(squares, batch_size=5)
for batch, (X, Y) in enumerate(dataloader):
print(X, '\n\n', Y)
break
# -
# # Digits
digits = datasets.MNIST('data', train=True, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Lambda(lambda x: x.view(28*28))
]),
target_transform=transforms.Compose([
transforms.Lambda(lambda y: torch.zeros(10, dtype=torch.float).scatter_(0, y, 1))
])
)
# +
dataloader = DataLoader(digits, batch_size=10, shuffle=True)
for batch, (X, Y) in enumerate(dataloader):
print(X, '\n\n', Y)
break
# -
| data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import sys, os
import pandas as pd
import numpy as np
from collections import defaultdict
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder, StandardScaler
from sklearn.pipeline import make_pipeline
# Use below for charts in dark jupyter theme
THEME_DARK = False
if THEME_DARK:
# This is used if Jupyter Theme dark is enabled.
# The theme chosen can be activated with jupyter theme as follows:
# >>> jt -t oceans16 -T -nfs 115 -cellw 98% -N -kl -ofs 11 -altmd
font_size = '20.0'
dark_theme_config = {
"ytick.color" : "w",
"xtick.color" : "w",
"text.color": "white",
'font.size': font_size,
'axes.titlesize': font_size,
'axes.labelsize': font_size,
'xtick.labelsize': font_size,
'ytick.labelsize': font_size,
'legend.fontsize': font_size,
'figure.titlesize': font_size,
'figure.figsize': [20, 7],
'figure.facecolor': "#384151",
'legend.facecolor': "#384151",
"axes.labelcolor" : "w",
"axes.edgecolor" : "w"
}
plt.rcParams.update(dark_theme_config)
sys.path.append("..")
import xai
import xai.data
# -
csv_path = 'data/adult.data'
categorical_cols = ["gender", "workclass", "education", "education-num", "marital-status",
"occupation", "relationship", "ethnicity", "loan"]
csv_columns = ["age", "workclass", "fnlwgt", "education", "education-num", "marital-status",
"occupation", "relationship", "ethnicity", "gender", "capital-gain", "capital-loss",
"hours-per-week", "loan"]
df = xai.data.load_census()
df.tail()
target = "loan"
protected = ["ethnicity", "gender", "age"]
df_groups = xai.imbalance_plot(df, "gender", categorical_cols=categorical_cols)
groups = xai.imbalance_plot(df, "gender", "loan", categorical_cols=categorical_cols)
bal_df = xai.balance(df, "gender", "loan", upsample=0.8, categorical_cols=categorical_cols)
groups = xai.group_by_columns(df, ["gender", "loan"], categorical_cols=categorical_cols)
for group, group_df in groups:
print(group)
print(group_df["loan"].head(), "\n")
_ = xai.correlations(df, include_categorical=True, plot_type="matrix")
_ = xai.correlations(df, include_categorical=True)
# +
proc_df = xai.normalize_numeric(bal_df)
proc_df = xai.convert_categories(proc_df)
x = proc_df.drop("loan", axis=1)
y = proc_df["loan"]
x_train, y_train, x_test, y_test, train_idx, test_idx = \
xai.balanced_train_test_split(
x, y, "gender",
min_per_group=300,
max_per_group=300,
categorical_cols=categorical_cols)
x_train_display = bal_df[train_idx]
x_test_display = bal_df[test_idx]
print("Total number of examples: ", x_test.shape[0])
df_test = x_test_display.copy()
df_test["loan"] = y_test
_= xai.imbalance_plot(df_test, "gender", "loan", categorical_cols=categorical_cols)
# +
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, mean_squared_error, roc_curve, auc
from keras.layers import Input, Dense, Flatten, \
Concatenate, concatenate, Dropout, Lambda
from keras.models import Model, Sequential
from keras.layers.embeddings import Embedding
def build_model(X):
input_els = []
encoded_els = []
dtypes = list(zip(X.dtypes.index, map(str, X.dtypes)))
for k,dtype in dtypes:
input_els.append(Input(shape=(1,)))
if dtype == "int8":
e = Flatten()(Embedding(X[k].max()+1, 1)(input_els[-1]))
else:
e = input_els[-1]
encoded_els.append(e)
encoded_els = concatenate(encoded_els)
layer1 = Dropout(0.5)(Dense(100, activation="relu")(encoded_els))
out = Dense(1, activation='sigmoid')(layer1)
# train model
model = Model(inputs=input_els, outputs=[out])
model.compile(optimizer="adam", loss='binary_crossentropy', metrics=['accuracy'])
return model
def f_in(X, m=None):
"""Preprocess input so it can be provided to a function"""
if m:
return [X.iloc[:m,i] for i in range(X.shape[1])]
else:
return [X.iloc[:,i] for i in range(X.shape[1])]
def f_out(probs, threshold=0.5):
"""Convert probabilities into classes"""
return list((probs >= threshold).astype(int).T[0])
# +
model = build_model(x_train)
model.fit(f_in(x_train), y_train, epochs=50, batch_size=512)
# -
score = model.evaluate(f_in(x_test), y_test, verbose=1)
print("Error %.4f: " % score[0])
print("Accuracy %.4f: " % (score[1]*100))
probabilities = model.predict(f_in(x_test))
pred = f_out(probabilities)
_= xai.metrics_plot(
y_test,
probabilities)
df.head()
_ = xai.metrics_plot(
y_test,
probabilities,
df=x_test_display,
cross_cols=["gender", "ethnicity"],
categorical_cols=categorical_cols)
_ = [xai.metrics_plot(
y_test,
probabilities,
df=x_test_display,
cross_cols=[p],
categorical_cols=categorical_cols) for p in protected]
xai.confusion_matrix_plot(y_test, pred)
xai.confusion_matrix_plot(y_test, pred, scaled=False)
_ = xai.roc_plot(y_test, probabilities)
_ = [xai.roc_plot(
y_test,
probabilities,
df=x_test_display,
cross_cols=[p],
categorical_cols=categorical_cols) for p in protected]
_= xai.pr_plot(y_test, probabilities)
_ = [xai.pr_plot(
y_test,
probabilities,
df=x_test_display,
cross_cols=[p],
categorical_cols=categorical_cols) for p in protected]
d = xai.smile_imbalance(
y_test,
probabilities)
d[["correct", "incorrect"]].sum().plot.bar()
d = xai.smile_imbalance(
y_test,
probabilities,
threshold=0.75,
display_breakdown=True)
display_bars = ["true-positives", "true-negatives",
"false-positives", "false-negatives"]
d[display_bars].sum().plot.bar()
d = xai.smile_imbalance(
y_test,
probabilities,
bins=9,
threshold=0.75,
manual_review=0.00001,
display_breakdown=False)
d[["correct", "incorrect", "manual-review"]].sum().plot.bar()
# +
def get_avg(x, y):
return model.evaluate(f_in(x), y, verbose=0)[1]
imp = xai.feature_importance(x_test, y_test, get_avg)
imp.head()
# -
| examples/XAI Tabular Data Example Usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/GbotemiB/Housing_Project/blob/main/concat_desciption_score.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5uzjYcqQPsm_"
# #This notebook was used to concatenating the description score that was done in batches across different notebooks and later used to retrain the model in another notebook.
# + id="Hi40ZkGwn3-a"
import pandas as pd
import numpy as np
# + id="EoLN3Q3Dn8Mr"
# Code to read csv file into Colaboratory:!pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials# Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + id="tcFn8Dpmo1br"
id7k = '<KEY>'
id14k = '<KEY>'
id21k = '<KEY>'
id28k = '<KEY>'
id33k = '1-9fDl9HQs_xjA7RH4X7i4M-_kQPwIo6k'
id28kk = '1YWSST7T6n0MSyKiiXmOEvXc4FYrp5dSO'
id666k = '1Iq4sadtZxuDh9VFNdoRDZyYU6-X7zMfP'
id999k = '1F3e-_yu7Vc7ieKnx45fukT9BrXNybTlC'
id77k = '1JVKTq_uheFaKdm-gA4cx21F86WYnHmCE'
id84k = '<KEY>-Z<KEY>'
id90k = '<KEY>'
id95k = '<KEY>'
id100k = '<KEY>'
id42k = '<KEY>jKk6mDKhorvH99aNfZEDO0'
id47k = '<KEY>'
id52k = '1ZYd_MYIhD7ejh9SJZ1ObTF6fwcyAIdbx'
id60k = '1yqVEX3ju6CsC5R<KEY>'
id666k = '1N1QpeIHQjcTi2Knrp2xUAtcDmOujwT2E'
# + id="1c5OCE7so90v"
ids = [id7k, id14k, id21k, id28k, id33k, id28kk, id666k, id999k, id77k, id84k, id90k, id95k, id100k, id42k, id47k, id52k, id60k, id666k]
df = {}
# + id="oX2UuenVpBRl"
for id in ids:
download = drive.CreateFile({'id':id})
data = ids.index(id)
download.GetContentFile(f'{data}.csv')
df[f'df_{data}'] = f'{data}.csv'
# + id="k2accaf6pEUY"
df0 = pd.read_csv('0.csv', header=None)
df1 = pd.read_csv('1.csv', header=None)
df2 = pd.read_csv('2.csv', header=None)
df3 = pd.read_csv('3.csv', header=None)
df4 = pd.read_csv('4.csv', header=None)
df5 = pd.read_csv('5.csv', header=None)
df6 = pd.read_csv('6.csv', header=None)
df7 = pd.read_csv('7.csv', header=None)
df8 = pd.read_csv('8.csv', header=None)
df9 = pd.read_csv('9.csv', header=None)
df10 = pd.read_csv('10.csv', header=None)
df11 = pd.read_csv('11.csv', header=None)
df12 = pd.read_csv('12.csv', header=None)
df12 = pd.read_csv('12.csv', header=None)
df13 = pd.read_csv('13.csv', header=None)
df14 = pd.read_csv('14.csv', header=None)
df15 = pd.read_csv('15.csv', header=None)
df16 = pd.read_csv('16.csv', header=None)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="RuAg-633xG18" outputId="5079f03a-b13b-47c3-a668-04c7c896d9ed"
df16.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="L3_QX1-PpPYE" outputId="fd1735d2-10e1-4c30-849b-1894b81a7077"
concat_data = pd.concat([df0, df1, df2, df3, df4, df5, df6, df7, df8, df9, df10, df11, df12, df13, df14, df15, df16], ignore_index=True)
len(concat_data)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="_xvTOtIDx-h3" outputId="9acb6a26-34e4-418d-db50-016dacb3c0a6"
concat_data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="2Ps3eV_4pQzM" outputId="7bf8a477-cbd0-4ab3-c17d-a94dfb4d977d"
concat_data.rename(columns = {0:'id', 1:'available_date', 2:'adDescription', 3:'detectLang', 4:'transDescription', 5:'transScore'}, inplace=True)
concat_data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="l3P1KeA9yM4h" outputId="d1b9f4e7-59d4-404b-9987-e9db5e5f19ba"
concat_data.info()
# + id="eI4yubqrygi7"
sorted_df = concat_data.sort_values('id')
# + colab={"base_uri": "https://localhost:8080/"} id="vtQ2WA7mytvo" outputId="92dbc8ce-389d-4b6d-d28e-90f3f47994dc"
len(sorted_df[sorted_df.duplicated()])
# + colab={"base_uri": "https://localhost:8080/"} id="5-FnveGnzHMK" outputId="3c812508-5cdf-4d8f-d43d-54904f0f1c11"
102701 - 10552
# + id="_hFyZxR1zdF7"
droped_duplicate = sorted_df.drop_duplicates(keep='first', ignore_index=True, inplace=False)
# + colab={"base_uri": "https://localhost:8080/"} id="tN7mHKDN0G7e" outputId="b9d347ed-9636-46d6-bda9-2ccfb9831276"
len(droped_duplicate)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="k_xMBMpK0MNe" outputId="8b2a868f-798a-4564-c102-6d99feb31a68"
droped_duplicate.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="x1rRRjb10kEe" outputId="ed3c983d-243d-47c2-dd6e-8754eb4441a5"
droped_duplicate[['id', 'transScore']]
# + [markdown] id="JqIqJ4AyQCMC"
# #The next line of code was used to output the merged files to be used in the modelling notebook
# + id="VzG9pd6Q0PyG"
droped_duplicate[['id', 'transScore']].to_csv('description_score.csv',index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="VjJTypj80n38" outputId="5c909822-141c-4bf0-93b0-2a2a0577c152"
99146-92149
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="3aCmoBgM23Yt" outputId="ce4f1173-a090-4854-b221-2fbd089dee94"
df0.head()
# + id="M1dHMwyk3Hii"
| concat_desciption_score.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Bayesian Structural Time Series: Forecasting and Decomposition Using PyMC3
# This is an advanced example of how a custom Bayesian time series forecasting/decomposition model can be built using PyMC3. The implementation is based on this [example](https://docs.pymc.io/notebooks/GP-MaunaLoa.html).
#
# ## Data
# The notebook uses datasets that are availbale in the `tensor-house-data` repository.
#
# ---
# +
import matplotlib as mpl
from matplotlib import pylab as plt
import matplotlib.dates as mdates
import seaborn as sns
import collections
import numpy as np
import pandas as pd
import pymc3 as pm
import arviz as az
print('Running on PyMC3 v{}'.format(pm.__version__))
demand_dates = np.arange('2014-01-01', '2014-02-26', dtype='datetime64[h]')
demand_loc = mdates.WeekdayLocator(byweekday=mdates.WE)
demand_fmt = mdates.DateFormatter('%a %b %d')
data = pd.read_csv('../resources/time-series/electricity-demand-victoria.csv', comment='#', header=None).T
data = pd.concat([data, pd.DataFrame(demand_dates)], axis=1)
data.columns = ["demand", "temperature", "date"]
num_forecast_steps = 24 * 7 * 2 # two weeks
data_training = data[:-num_forecast_steps]
data_test = data[-num_forecast_steps:]
colors = sns.color_palette()
c1, c2 = colors[0], colors[1]
fig = plt.figure(figsize=(12, 6))
ax = fig.add_subplot(2, 1, 1)
ax.plot(data_training.date,
data_training.demand, lw=2, label="training data")
ax.set_ylabel("Hourly demand (GW)")
ax = fig.add_subplot(2, 1, 2)
ax.plot(data_training.date,
data_training.temperature, lw=2, label="training data", c=c2)
ax.set_ylabel("Temperature (deg C)")
ax.set_title("Temperature")
ax.xaxis.set_major_locator(demand_loc)
ax.xaxis.set_major_formatter(demand_fmt)
fig.suptitle("Electricity Demand in Victoria, Australia (2014)", fontsize=15)
fig.autofmt_xdate()
# -
#
# plotting functions
#
def plot_components(dates,
component_means_dict,
component_stddevs_dict,
x_locator=None,
x_formatter=None):
colors = sns.color_palette()
c1, c2 = colors[0], colors[1]
axes_dict = collections.OrderedDict()
num_components = len(component_means_dict)
fig = plt.figure(figsize=(12, 2.5 * num_components))
for i, component_name in enumerate(component_means_dict.keys()):
component_mean = component_means_dict[component_name]
component_stddev = component_stddevs_dict[component_name]
ax = fig.add_subplot(num_components,1,1+i)
ax.plot(dates, component_mean, lw=2)
ax.fill_between(dates,
component_mean-2*component_stddev,
component_mean+2*component_stddev,
color=c2, alpha=0.5)
ax.set_title(component_name)
if x_locator is not None:
ax.xaxis.set_major_locator(x_locator)
ax.xaxis.set_major_formatter(x_formatter)
axes_dict[component_name] = ax
fig.autofmt_xdate()
fig.tight_layout()
return fig, axes_dict
# ## Model Specification and Fitting
#
# We use an additive Gaussian processes model with a sum of three GPs for the signal, and one GP for the noise:
# $$
# f(t) \sim \mathcal{GP}_{\text{slow}}(0,\, k_1(t, t')) +
# \mathcal{GP}_{\text{med}}(0,\, k_2(t, t')) +
# \mathcal{GP}_{\text{per}}(0,\, k_3(t, t')) +
# \mathcal{GP}_{\text{noise}}(0,\, k_n(t, t'))
# $$
#
# * A long term smooth rising trend represented by an exponentiated quadratic kernel.
# * A periodic term that decays away from exact periodicity. This is represented by the product of a Periodic covariance function and an exponentiated quadratic.
# * Small and medium term irregularities with a rational quadratic kernel.
# * The noise is modeled as the sum of an Exponential and a white noise kernel
#
# Lengthscale priors:
# * ℓ_pdecay: The periodic decay. The smaller this parameter is, the faster the periodicity goes away.
# * ℓ_psmooth: The smoothness of the periodic component. It controls how “sinusoidal” the periodicity is.
# * period: The period. We put a strong prior on a period of one day.
# * ℓ_med: This is the lengthscale for the short to medium long variations.
# * α: This is the shape parameter.
# * ℓ_trend: The lengthscale of the long term trend.
# * ℓ_noise: The lengthscale of the noise covariance.
#
# Scale priors:
# * η_per: Scale of the periodic or seasonal component.
# * η_med: Scale of the short to medium term component.
# * η_trend: Scale of the long term trend.
# * σ: Scale of the white noise.
# * η_noise: Scale of correlated, short term noise.
# +
#
# plot the priors
#
x = np.linspace(0, 10, 1000)
priors = [
("ℓ_pdecay", pm.Gamma.dist(alpha=4, beta=0.5)),
("ℓ_psmooth", pm.Gamma.dist(alpha=4, beta=3)),
("period", pm.Normal.dist(mu=1, sigma=0.5)),
("ℓ_med", pm.Gamma.dist(alpha=1, beta=0.75)),
("α", pm.Gamma.dist(alpha=5, beta=2)),
("ℓ_trend", pm.Gamma.dist(alpha=4, beta=0.1)),
("ℓ_noise", pm.Gamma.dist(alpha=4, beta=4))]
for i, prior in enumerate(priors):
plt.plot(x, np.exp(prior[1].logp(x).eval()), label=prior[0])
plt.legend(loc="upper right")
plt.xlabel("days")
plt.show();
x = np.linspace(0, 10, 1000)
priors = [
("η_per", pm.HalfCauchy.dist(beta=2)),
("η_med", pm.HalfCauchy.dist(beta=1.0)),
("η_trend", pm.HalfCauchy.dist(beta=3)),
("σ", pm.HalfNormal.dist(sigma=0.25)),
("η_noise", pm.HalfNormal.dist(sigma=0.5))]
for i, prior in enumerate(priors):
plt.plot(x, np.exp(prior[1].logp(x).eval()), label=prior[0])
plt.legend(loc="upper right")
plt.xlabel("days")
plt.show();
# +
def dates_to_idx(timelist):
reference_time = pd.to_datetime('2014-01-01')
t = (timelist - reference_time) / pd.Timedelta(1, "D")
return np.asarray(t)
t = dates_to_idx(data_training.date)[:,None]
y = data_training.demand.values
#
# define and fit the model
#
with pm.Model() as model:
# daily periodic component x long term trend
η_per = pm.HalfCauchy("η_per", beta=2, testval=1.0)
ℓ_pdecay = pm.Gamma("ℓ_pdecay", alpha=4, beta=0.5)
period = pm.Normal("period", mu=1, sigma=0.5)
ℓ_psmooth = pm.Gamma("ℓ_psmooth ", alpha=4, beta=3)
cov_seasonal = η_per**2 * pm.gp.cov.Periodic(1, period, ℓ_psmooth) \
* pm.gp.cov.Matern52(1, ℓ_pdecay)
gp_seasonal = pm.gp.Marginal(cov_func=cov_seasonal)
# small/medium term irregularities
η_med = pm.HalfCauchy("η_med", beta=0.5, testval=0.1)
ℓ_med = pm.Gamma("ℓ_med", alpha=2, beta=0.75)
α = pm.Gamma("α", alpha=5, beta=2)
cov_medium = η_med**2 * pm.gp.cov.RatQuad(1, ℓ_med, α)
gp_medium = pm.gp.Marginal(cov_func=cov_medium)
# long term trend
η_trend = pm.HalfCauchy("η_trend", beta=2, testval=2.0)
ℓ_trend = pm.Gamma("ℓ_trend", alpha=4, beta=0.1)
cov_trend = η_trend**2 * pm.gp.cov.ExpQuad(1, ℓ_trend)
gp_trend = pm.gp.Marginal(cov_func=cov_trend)
# noise model
η_noise = pm.HalfNormal("η_noise", sigma=0.5, testval=0.05)
ℓ_noise = pm.Gamma("ℓ_noise", alpha=4, beta=4)
σ = pm.HalfNormal("σ", sigma=0.25, testval=0.05)
cov_noise = η_noise**2 * pm.gp.cov.Matern32(1, ℓ_noise) + pm.gp.cov.WhiteNoise(σ)
# The Gaussian process is a sum of these three components
gp = gp_seasonal + gp_medium + gp_trend
# Since the normal noise model and the GP are conjugates, we use `Marginal` with the `.marginal_likelihood` method
y_ = gp.marginal_likelihood("y", X=t, y=y, noise=cov_noise)
# this line calls an optimizer to find the MAP
mp = pm.find_MAP(include_transformed=True)
# fitted model parameters
sorted([name + ":" + str(mp[name]) for name in mp.keys() if not name.endswith("_")])
# -
# ## Forecasting
# +
dates = pd.date_range(start='2014-02-12', end="2014-02-26", freq="1H")[:-1]
tnew = dates_to_idx(dates)[:,None]
first_y = 0
std_y = 1
print("Sampling gp predictions...")
mu_pred, cov_pred = gp.predict(tnew, point=mp)
# draw samples, and rescale
n_samples = 2000
samples = pm.MvNormal.dist(mu=mu_pred, cov=cov_pred).random(size=n_samples)
samples = samples * std_y + first_y
### plot mean and 2σ region of total prediction
fig = plt.figure(figsize=(16, 6))
# scale mean and var
mu_pred_sc = mu_pred * std_y + first_y
sd_pred_sc = np.sqrt(np.diag(cov_pred) * std_y**2 )
upper = mu_pred_sc + 2*sd_pred_sc
lower = mu_pred_sc - 2*sd_pred_sc
c = sns.color_palette()
plt.plot(data_test.date, mu_pred_sc, linewidth=2, color=c[0], label="Total fit")
plt.fill_between(data_test.date, lower, upper, color=c[0], alpha=0.4)
# some predictions
idx = np.random.randint(0, samples.shape[0], 10)
for i in idx:
plt.plot(data_test.date, samples[i,:], color=c[0], alpha=0.5, linewidth=0.5)
# true value
plt.plot(data_test.date, data_test.demand, linewidth=2, color=c[1], label="Observed data")
plt.ylabel("Demand")
plt.title("Demand forecast")
plt.legend(loc="upper right")
plt.show();
# -
# ## Decomposition
# +
# predict at a 1 hour granularity
dates = pd.date_range(start='2014-01-01', end="2014-02-12", freq="1H")[:-1]
tnew = dates_to_idx(dates)[:,None]
print("Predicting with gp ...")
mu, var = gp.predict(tnew, point=mp, diag=True)
mean_pred = mu*std_y + first_y
var_pred = var*std_y**2
# make dataframe to store fit results
fit = pd.DataFrame({"t": tnew.flatten(),
"mu_total": mean_pred,
"sd_total": np.sqrt(var_pred)},
index=dates)
print("Predicting with gp_trend ...")
mu, var = gp_trend.predict(tnew, point=mp,
given={"gp": gp, "X": t, "y": y, "noise": cov_noise},
diag=True)
fit = fit.assign(mu_trend = mu*std_y + first_y,
sd_trend = np.sqrt(var*std_y**2))
print("Predicting with gp_medium ...")
mu, var = gp_medium.predict(tnew, point=mp,
given={"gp": gp, "X": t, "y": y, "noise": cov_noise},
diag=True)
fit = fit.assign(mu_medium = mu*std_y + first_y,
sd_medium = np.sqrt(var*std_y**2))
print("Predicting with gp_seasonal ...")
mu, var = gp_seasonal.predict(tnew, point=mp,
given={"gp": gp, "X": t, "y": y, "noise": cov_noise},
diag=True)
fit = fit.assign(mu_seasonal = mu*std_y + first_y,
sd_seasonal = np.sqrt(var*std_y**2))
## plot the decompostion
fig = plt.figure(figsize=(16, 6))
# plot mean and 2σ region of total prediction
upper = fit.mu_total + 2*fit.sd_total
lower = fit.mu_total - 2*fit.sd_total
band_x = np.append(fit.index.values, fit.index.values[::-1])
band_y = np.append(lower, upper[::-1])
c = sns.color_palette()
# total fit
plt.plot(fit.index, fit.mu_total, linewidth=1, color=c[0], label="Total fit")
plt.fill_between(fit.index, lower, upper, color=c[0], alpha=0.4)
# trend
plt.plot(fit.index, fit.mu_trend, linewidth=1, color=c[1], label="Long term trend")
# medium
plt.plot(fit.index, fit.mu_medium, linewidth=1, color=c[2], label="Medium range variation")
# seasonal
plt.plot(fit.index, fit.mu_seasonal, linewidth=1, color=c[3], label="Seasonal process")
# true value
plt.plot(data_training.date, data_training.demand, linewidth=2, color=c[4], label="Observed data")
plt.ylabel("Demand")
plt.title("Demand decomposition")
plt.legend(loc="upper right")
plt.show();
#
# plot separate components of the decomposition
#
demand_component_means = {
'Ground truth': data_training.demand,
'Total fit': fit.mu_total,
'Trend': fit.mu_trend,
'Medium': fit.mu_medium,
'Seasonal': fit.mu_seasonal,
}
demand_component_stddevs = {
'Ground truth': np.zeros(len(data_training.demand)),
'Total fit': fit.sd_total,
'Trend': fit.sd_trend,
'Medium': fit.sd_medium,
'Seasonal': fit.sd_seasonal,
}
fig, axes = plot_components(
data_training.date,
demand_component_means,
demand_component_stddevs,
x_locator=demand_loc, x_formatter=demand_fmt)
| _basic-components/time-series/bsts-part-4-forecasting-pymc3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from numpy.linalg import inv
from scipy.stats import multivariate_normal
import matplotlib.pyplot as plt
from numpy import logical_and as land
from numpy import logical_or as lor
from numpy import invert as lin
# +
def get_theta(mu, cov, rho):
'''
Returns explicit threshold theta for a given percentage rho of anomalies in
data distributed as a Gaussian with mean mu and covariance matrix cov.
Parameters
mu mean of Gaussian distribution
cov covariance matrix of Gaussian distribution
rho percentage of anomalies, which must be between 0 and 100 inclusive
'''
# generate random variables (data)
X = multivariate_normal.rvs(mean=mu, cov=cov, size=5000000)
# center data (normalize) (for x_i - mu)
Z = X - mu
# calculate the mahalanobis distance
# d2M (xi, ˆμ) = (xi − ˆμ)T ˆΣ−1(xi − ˆμ)
d = np.sqrt(np.sum(Z.dot(inv(cov)) * Z, axis=1))
# tetha =
return np.percentile(d, 100-rho)
# get_theta([0, 0], [[1, 0], [0, 1]], 5)
# +
# plot settings and utility functions
plt.style.use('seaborn-dark')
plt.rcParams['figure.figsize']= 16, 10
def save_point_plot(data, outliers_indices, fpath):
'''
plot 2dim data and save them to file
data data plotted blue
outliers_indices if is not None, indices which are True will be plotted red
'''
outliers = data[outliers_indices, :]
data = data[lin(outliers_indices), :]
# create new figure
# fig, ax = plt.subplots()
# Plotting the generated samples
plt.plot(data[:,0], data[:,1], 'o', c='blue',
markeredgewidth = 0.5,
markeredgecolor = 'black')
plt.plot(outliers[:,0], outliers[:,1], 'o', c='red',
markeredgewidth = 0.5,
markeredgecolor = 'black')
# plt.title('covariance of distribution')
plt.xlabel('x1')
plt.ylabel('x2')
plt.axis('equal')
plt.gcf().savefig(fpath, bbox_inches='tight')
plt.close(plt.gcf())
# -
# # Assignment
# 1. Sample a data set D of size n from N (x; μ, Σ). Fix a percentage ρ.
# 2. Use the function get_theta(mu, cov, rho) provided by the notebook to
# obtain an explicit threshold θ given the percentage ρ. Note that θ is part
# of the ground-truth and therefore considered as unknown.
# 3. Determine the true anomalies of D. For this, use the explicit threshold θ
# together with the Mahalanobis distance d∗
# M defined by the true μ and Σ.
# 4. Use the data D to estimate μ and Σ. Construct the Mahalanobis distance
# dM defined by the estimates ˆμ and ˆΣ.
# 5. Predict the anomalies of D using the Mahalanobis distance dM and Eu-
# clidean distance dE . Anomalies are the ρ percent points xi ∈ D farthest
# from ˆμ (do not use θ). Assess precision and recall of both detectors.
# +
# function to evaluate the whole assignment and save the plotted figure
def evaluate(mu_T=np.array([0, 0]), covariance=.9, rho=3.5, size=2000, savefig=True):
# fixate groundtruth mean and covariance matrix for the bivariate gaussian distribution
# '_T' nominator stands for groundtruth variable
# '_E' nominator stands for estimated variable
sigma_T = np.array([[1, covariance], [covariance, 1]]) # covariance matrix
# 1. generate dataset (RandomVariableS)
D = multivariate_normal.rvs(mean=mu_T, cov=sigma_T, size=size)
# 2. use get_theta to get the 'groundtruth' explicit treshold theta
theta = get_theta(mu_T, sigma_T, rho)
# 3. determine subset of true anomalies of dataset D
# start by calculating the mahalanobis distance of each point from the mean
Z_T = D - mu_T
d_m_T = np.sqrt(np.sum(Z_T.dot(inv(sigma_T)) * Z_T, axis=1))
# filter out values (indices) over the groundtruth threshold theta (True / False array)
I_T = d_m_T > theta # indices of true anomalies with mahalanobis distance
# 4. Use the data D to estimate mu and sigma
mu_E = D.mean(axis=0)
sigma_E = np.cov(D.T)
# Construct the Mahalanobis distance d_m_E defined by the estimates mu_E and sigma_E
Z_E = D - mu_E
d_m_E = np.sqrt(np.sum(Z_E.dot(inv(sigma_E)) * Z_E, axis=1))
# construct euclidean distance d_e_E in the same manner (with mu_E and sigma_E)
d_e_E = np.sqrt(np.sum(Z_E ** 2, axis=1))
# 5. predict anomalies with estimated eucilidian (d_e_E) and mahalanobis distance (d_m_E)
# create list of indices (True / False array) (on axis 0 of dataset)
# estimated thresholds (eta) are rho percent points with the farthest distance from mu_E
eta_m = np.percentile(d_m_E, 100-rho)
eta_e = np.percentile(d_e_E, 100-rho)
I_m_E = d_m_E > eta_m
I_e_E = d_e_E > eta_e
# Comparison:
# Assess precision and recall of both detectors. (5)
# calculate tp, fp and fn for euclidean distance and for mahalanobis distance
# np.logical_and(I_m_T, I_m_E) [here: land] creates a logical AND mask over the two boolean arrays etc.
# (I_m_T * I_m_E)
tp_m = land(I_T, I_m_E).sum()
tp_e = land(I_T, I_e_E).sum()
fp_m = land(lin(I_T), I_m_E).sum()
fp_e = land(lin(I_T), I_e_E).sum()
fn_m = land(I_T, lin(I_m_E)).sum()
fn_e = land(I_T, lin(I_e_E)).sum()
# precisions and recalls mahalanobis (m) and euclidean (e) distance
precision_m = tp_m / (tp_m + fp_m)
recall_m = tp_m / (tp_m + fn_m)
precision_e = tp_e / (tp_e + fp_e)
recall_e = tp_e / (tp_e + fn_e)
print(f'precision euclidean : {precision_e}')
print(f'recall euclidean : {recall_e}')
print(f'precision mahalanobis : {precision_m}')
print(f'recall mahalanobis : {recall_m}')
# save figure
if savefig:
save_point_plot(D, I_T, f'./research_question_imgs/{rho}_groundtruth.png')
save_point_plot(D, I_m_E, f'./research_question_imgs/{rho}_mahalanobis_estimated.png')
save_point_plot(D, I_e_E, f'./research_question_imgs/{rho}_euclidean_estimated.png')
evaluate(rho=40)
# + [markdown] tags=[]
# ## Research question of choice : How do precision and recall depend on percentage __p__?
# -
# loop through the whole evaluation process with always
# augmenting values of rho
# save the anomalies plots for each percentage evaluation
# for groundtruth, estimated euclidean and mahalanobis distance
for i in range(1, 99, 3):
print(f'\nrho = {i}')
evaluate(rho=i)
# ## results:
#
# More outliers (higher rho) makes the euclidean distance work better than with lower rho. The mahalanobis distance works in any case equally good.
#
# **Why?**
#
# The euclidean distance measures unrelative (circular-shape) to the pdf, and since the nearer you get to the mean (mu) of the pdf the more points occur in a gaussian distribution, the less 'meaning' the co-dependence of the two axis gets. Since the mahalanobis distance works relatively (oval-shaped) to the density function, its performance wont change (oval). This only applies if the two variables are co-dependant (covariance not zero). Else neither distance measurement works better.
| assignment_5_1/research_question.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Running Python Scripts
# The first thing you want to be able to do is to learn how to run a Python script. Since you have installed Python, you
# can just call it from the command line and give the location of the Python file that we want to run.
#
# In this directory, we have a file called `example.py`. Without worrying about what's in the code, try to run it!
#
# ```bash
# python example.py
# ```
#
# or if you explicitly installed it as python3
#
# ```bash
# python3 example.py
# ```
run example.py # press the "Run" button to run this script!
# [Up Next: Lesson 2 - Variables](variables.ipynb)
#
# [Go Back: Lessons 2 - Python Basics](index.ipynb)
| lessons/lesson02-python-basics/running-python-scripts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
data = pd.read_csv('mercedesbenz.csv',usecols=['X1', 'X2', 'X3', 'X4', 'X5', 'X6', ])
data.head()
# lets look at how many labels each variables has
for col in data.columns:
print(col,':',len(data[col].unique()),'labels')
#lets examine how many columns we will obtain after one hot encoding these varibles.
pd.get_dummies(data, drop_first=True).shape
# lets find the top 10 most frequent categories for the variable x2
data.X2.value_counts().sort_values(ascending=False).head(20)
# +
#lets make a list with the most frequent categories of the varibles
top_10 = [x for x in data.X2.value_counts().sort_values(ascending=False).head(10).index]
top_10
# -
# and now we make the 10 binary variables
for label in top_10:
data[label] = np.where(data['X2'] == label, 1, 0)
data[['X2']+top_10].head(40)
# +
# get whole set of dummy variables, for all the categorical variables
def one_hot_top_x(df, variable, top_x_labels):
#functions to create the dummy varibles for the most frequent labels
#we can vary the number of most frequent labels that we encode
for label in top_x_labels:
df[variable+'_'+label] = np.where(data[variable]==label, 1, 0)
#read the data again
data = pd.read_csv('mercedesbenz.csv',usecols=['X1', 'X2', 'X3', 'X4', 'X5', 'X6', ])
#encode X2 into top 10 most frequent categories
one_hot_top_x(data,'X2',top_10)
data.head()
# +
# find the 10most frequet values for X1
top_10 = [x for x in data.X1.value_counts().sort_values(ascending=False).head(10).index]
#now create the 10 most frequet dummy varibes for x1
one_hot_top_x(data, 'X1', top_10)
data.head()
# -
| Feature Engineering/One Hot encoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def checknoinarr(li,x):
length = len(li)
if (length == 0):
return False
if(li[0] == x):
return True
smallerarr = checknoinarr(li[1:],x)
return smallerarr
# +
n = int(input())
li = [int(ele) for ele in input().split()]
x = int(input())
checknoinarr(li,x)
# -
| 02.Data-Structures-and-Algorithms/01.Recursion-1/09.Check-no-in-array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
@author: <NAME>
"""
import numpy as np
import tensorflow as tf
from tqdm import tqdm
import time
# Load words list
words = np.load('wordsList.npy').tolist()
words = [word.decode('UTF-8') for word in words]
word_vectors = np.load('wordVectors.npy')
# Load word2vec embeding
train_x = np.load('reviews_train_x.npy')
train_y = np.load('reviews_train_y.npy')
test_x = np.load('reviews_test_x.npy')
test_y = np.load('reviews_test_y.npy')
print(train_x.shape)
# Consts
TIME_STEPS = 250
BATCH_SIZE = 256
STATE_SIZE = 200
TRAIN_STEPS = 100
LSTM_LAYERS = 3
# Data loaders
x_input = tf.placeholder(tf.int64, shape = (None, TIME_STEPS))
y_input = tf.placeholder(tf.int64, shape = (None, 2))
x_embed = tf.gather(word_vectors, x_input)
print(x_embed)
x_unpack = tf.unstack(x_embed, TIME_STEPS, 1)
print(x_unpack)
# +
# print(words[2])
words[0] = ''
words[399999] = '???'
# 0 - nothing
# 399999 - unk
def vec2sent(vec):
return ' '.join(map(lambda i: words[i], vec)).strip()
def evaluate(pred, x, y):
count = 0
for i in range(len(pred)):
if pred[i] == 0:
count += 1
print(vec2sent(x[i]), 'POSITIVE' if y[i][0] else 'NEGATIVE')
if count >= 5:
break
# +
"""
@params input
@params learning<True/False> are we learning or testing?
@params params<Dictionary>
"""
# def RNN(x, learning):
# # lstm = tf.contrib.rnn.LSTMCell(STATE_SIZE)
# # lstm = tf.contrib.rnn.MultiRNNCell(
# # [tf.contrib.rnn.LSTMCell(STATE_SIZE) for _ in range(LSTM_LAYERS)])
# layers = []
# for _ in range(LSTM_LAYERS):
# cell = tf.contrib.rnn.GRUCell(STATE_SIZE)
# # variational_recurrent=True, input_size=STATE_SIZE, dtype=tf.float32 - pogarsza wyniki
# wrap = tf.contrib.rnn.DropoutWrapper(cell, output_keep_prob=1.0 - dropout, state_keep_prob=1.0 - 0.5*dropout)
# layers.append(wrap)
# lstm = tf.contrib.rnn.MultiRNNCell(layers)
# outputs, states = tf.contrib.rnn.static_rnn(lstm, x, dtype=tf.float32)
# dense = tf.layers.dense(outputs[-1], 2)
# return dense
# dropout = tf.placeholder(tf.float32)
# rnn = RNN(x_unpack, dropout)
# prediction = tf.argmax(tf.nn.softmax(rnn), 1)
# loss_op = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=rnn, labels=y_input))
# #optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01).minimize(loss_op)
# global_step = tf.Variable(0, trainable=False)
# starter_learning_rate = 1e-3
# learning_rate = tf.train.exponential_decay(starter_learning_rate, global_step,
# 90, 0.96, staircase=True)
# optimizer = tf.train.AdamOptimizer(learning_rate).minimize(loss_op, global_step = global_step)
# correct_pred = tf.equal(prediction, tf.argmax(y_input, 1))
# accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# init = tf.global_variables_initializer()
# print(rnn)
# +
# import sys
# verbose = False
# t0 = time.time()
# with tf.Session() as session:
# session.run(init)
# for i in range(100):
# idx = np.arange(len(train_x))
# np.random.shuffle(idx)
# idx = np.array_split(idx, (len(idx)+BATCH_SIZE-1)//BATCH_SIZE)
# accs = []
# for step in tqdm(idx, leave = False):
# # step = np.random.choice(len(train_x), BATCH_SIZE)
# x_batch, y_batch = train_x[step], train_y[step]
# _, acc = session.run([optimizer, accuracy], feed_dict = {x_input: x_batch, y_input: y_batch, dropout: 0.2})
# accs.append(acc)
# acc_train = 1.0*sum(accs)/len(accs)
# lr, pred, acc = session.run([learning_rate, correct_pred, accuracy], feed_dict = {x_input: test_x, y_input: test_y, dropout: 0.0})
# dt = time.time() - t0
# sys.stderr.flush()
# sys.stdout.flush()
# l = list(map(lambda i: 0 if pred[i] else np.count_nonzero(test_x[i]), range(len(pred))))
# l = 1.0*sum(l) / np.count_nonzero(l)
# print("{} acc: {:.2f} acc_train: {:.2f} time: {:.2f}, lr: {}, len: {}".format(i, 100*acc, 100*acc_train, dt, lr, l))
# # if i>5:
# # evaluate(pred, test_x, test_y)
# sys.stdout.flush()
# +
import itertools
tf.reset_default_graph()
# Factory for different LSTM cells
def cell(ctype, state_size, dropout, state_dropout, training):
if ctype != 'NORM':
if ctype == 'BASIC':
cell = tf.contrib.rnn.BasicLSTMCell(state_size)
# Gated Recurrent Unit cell (cf. http://arxiv.org/abs/1406.1078).
if ctype == 'GRU':
cell = tf.contrib.rnn.GRUCell(state_size)
# Long short-term memory unit (LSTM) recurrent network cell.
if ctype == 'LSTM':
cell = tf.contrib.rnn.LSTMCell(state_size)
# Long short-term memory unit (LSTM) recurrent network cell with peephholes
if ctype == 'LSTM-PEEP':
cell = tf.contrib.rnn.LSTMCell(state_size, use_peepholes=True)
return tf.contrib.rnn.DropoutWrapper(
cell,
output_keep_prob = tf.where(training, 1.0 - dropout, 1.0),
state_keep_prob = tf.where(training, 1.0 - state_dropout, 1.0)
)
else:
return tf.contrib.rnn.LayerNormBasicLSTMCell(
state_size,
dropout_keep_prob = tf.where(training, 1 - dropout, 1.0)
)
def network(input, output, training, params):
layers = []
for _ in range(params['layer_count']):
layers.append(cell(params['cell_type'], params['state_size'], params['dropout'], params['state_dropout'], training))
lstm = tf.contrib.rnn.MultiRNNCell(layers)
outputs, states = tf.contrib.rnn.static_rnn(lstm, input, dtype=tf.float32)
dense = tf.layers.dense(outputs[-1], 2)
# prediction = tf.argmax(tf.nn.softmax(dense), 1)
# loss_op = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=dense, labels=output))
return dense
def hyper_network(hparams):
for product in itertools.product(*hparams.values()):
tf.reset_default_graph()
x_input = tf.placeholder(tf.int64, shape = (None, TIME_STEPS))
y_input = tf.placeholder(tf.int64, shape = (None, 2))
x_embed = tf.gather(word_vectors, x_input)
x_unpack = tf.unstack(x_embed, TIME_STEPS, 1)
training = tf.placeholder(tf.bool)
params = {key: product[i] for i, key in enumerate(hparams.keys())}
print('Building: ', params)
yield (product, network(x_unpack, y_input, training, params), x_input, y_input, training)
def hyper_evaluate(network, x_input, y_input, training, steps=5):
# tf.reset_default_graph()
prediction = tf.argmax(tf.nn.softmax(network), 1)
loss_op = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=network, labels=y_input))
global_step = tf.Variable(0, trainable=False)
starter_learning_rate = 1e-3
learning_rate = tf.train.exponential_decay(starter_learning_rate, global_step,
90, 0.96, staircase=True)
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(loss_op, global_step = global_step)
correct_pred = tf.equal(prediction, tf.argmax(y_input, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
init = tf.global_variables_initializer()
with tf.Session() as session:
session.run(init)
# print('Init')
for i in range(steps):
#print('Step: ', i)
idx = np.arange(len(train_x))
np.random.shuffle(idx)
idx = np.array_split(idx, (len(idx)+BATCH_SIZE-1)//BATCH_SIZE)
accs = []
for step in tqdm(idx, leave = False):
# step = np.random.choice(len(train_x), BATCH_SIZE)
x_batch, y_batch = train_x[step], train_y[step]
_, acc = session.run([optimizer, accuracy], feed_dict = {x_input: x_batch, y_input: y_batch, training: True})
accs.append(acc)
acc_train = 1.0*sum(accs)/len(accs)
#print(acc_train)
_, _, acc = session.run([learning_rate, correct_pred, accuracy], feed_dict = {x_input: test_x, y_input: test_y, training: False})
return acc
# def evaluate(net):
# pass
params = {
'dropout': [0.0, 0.1, 0.15, 0.3],
'state_dropout': [0.0, 0.1, 0.2, 0.3],
'cell_type': ['GRU', 'LSTM', 'LSTM-PEEP', 'NORM'],
'layer_count': [1, 2, 3],
'state_size': [25, 50, 120, 300]
}
for round in range(5):
print('Round: ', round+1)
for p, net, x, y, t in hyper_network(params):
print('RESULTS: ', p, 'ACC=', hyper_evaluate(net, x, y, t))
# print(len(list(hyper_network(params))))
# for i, net in enumerate(hyper_network(params)):
# acc = evaluate(net)
# print(i, acc)
# -
| 05-sentimental/05 - Sentiment Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # IntroToHDF5
#
# Small introduction on how to work with the HDF5 format
# within Python, R and C++ programming languages.
#
# Documentation on [h5py] and [rhdf5].
#
# [h5py]: https://docs.h5py.org/en/stable/quick.html
# [rhdf5]: https://bioconductor.org/packages/release/bioc/vignettes/rhdf5/inst/doc/rhdf5.html
#
# ## Requirements
#
# To use HighFive library (for reading and writing of
# HDF5 files in C++) need to install (through `apt`
# tool in Ubuntu):
# * libboost-serialization1.71-dev
# * libboost-system1.71-dev
# * libboost1.71-dev
# * libhdf5-dev
# * hdf5-helpers
# * hdf5-tools
#
# To reproduce anything in this repository, first
# create the conda environment from `conda_env_simple.yml`:
# ```
# conda env create -n test_hdf5 -f conda_env_simple.yml
# ```
# If conda is not installed, follow the installation
# instructions [here](https://docs.conda.io/projects/conda/en/latest/user-guide/install/#regular-installation).
#
# Then activate the conda enviornment:
# ```
# conda activate test_hdf5
# ```
# and execute the following command:
# ```
# make all
# ```
# # h5py package (Python)
# +
# %load_ext rpy2.ipython
import h5py
import numpy as np
from os import path, makedirs
import pandas as pd
if not path.isdir("Output"): makedirs("Output")
# -
# Writing to an hfd5 file:
# +
fname = "Output/h5py_test.h5"
with h5py.File(fname, "w") as fh:
fh.create_dataset("int", data=1)
fh.create_dataset("string", data="test")
fh.create_dataset("array", data=[0, 10, 5])
fh.create_dataset("strings", data=["hello", "world"])
df = pd.DataFrame([["a", 1], ["b", 2]], columns=["letter", "number"])
df.to_hdf(fname, "df")
df
# -
# Reading from an hdf5 file:
with h5py.File(fname) as fh:
print(fh.keys())
print(fh["int"][()])
print(fh["string"][()].decode("UTF-8"))
print(fh["array"][:])
print([x.decode("UTF-8") for x in fh["strings"][()]])
pd.read_hdf(fname, "df")
# **Note**: Pandas dataframes can be saved and read from an hdf5 file, but through pandas' own API, not through h5py.
# # rhdf5 package (R)
# + language="R"
# library(rhdf5)
# -
# Writing to an hdf5 file:
# + language="R"
#
# fname <- "Output/rhdf5_test.h5"
# if (file.exists(fname)) { file.remove(fname) }
#
# h5createFile(fname)
# h5write(42, fname, "int")
# h5write("test", fname, "string")
# h5write(c(1.0, 2.0, 5.0), fname, "vector")
# strings <- c("hello", "world", "foofoofoo", "barbarbarbar")
# h5write(strings, fname, "strings")
#
# d <- rbind(c(FALSE, TRUE),
# c(TRUE, FALSE),
# c(TRUE, FALSE))
# h5write(d, fname, "rectangular_matrix")
#
# name <- c("Jon", "Bill", "Maria", "Ben", "Tina")
# age <- c(23, 41, 32, 58, 26)
# df <- data.frame(name, age)
# h5write(df, fname, "df")
#
# h5closeAll()
# -
# Reading from an hdf5 file:
# + language="R"
#
# fh = H5Fopen(fname)
# print(fh)
# print(fh$"int")
# print(fh$"string")
# print(fh$"vector")
# print(fh$"strings")
# print(fh$"rectangular_matrix")
# print(fh$"df")
# h5closeAll()
# -
# Data in an hdf5 file created in one language can be read in another language, but we need to be careful, as shown later
with h5py.File("Output/rhdf5_test.h5") as fh:
print(fh.keys())
print(fh["int"][0])
print(fh["string"][0].decode("UTF-8"))
print(fh["vector"][:])
print([x.decode("UTF-8") for x in fh["strings"]])
df = pd.DataFrame(np.array(fh["df"]))
df["name"] = df["name"].apply(lambda x : x.decode("UTF-8"))
print(df)
# **Important**: `rhdf5` package saves matrices transposed, which is explained in `rhdf5` docs [here](https://bioconductor.org/packages/release/bioc/vignettes/rhdf5/inst/doc/rhdf5.html#reading-hdf5-files-with-external-software). If we open matrix D in `rhdf5_test.h5` in Python:
with h5py.File("Output/rhdf5_test.h5") as fh:
print(fh["rectangular_matrix"][:])
# The matrix `rectangular_matrix`, when read in Python is transposed and boolean entries have been changed to integers. We can observe the same behaviour when opening a file created by `rhdf5` package within C++. This happens because `rhdf5` "This is due to the fact the fastest changing dimension on C is the last one, but on R it is the first one (as in Fortran)." based on `rhdf5` documentation.
# + language="R"
# fh = H5Fopen("Output/rhdf5_test.h5")
# print(fh$"rectangular_matrix")
# h5closeAll()
# -
# Lists and vectors from R saved in hdf5 file:
# + language="R"
#
# fname <- "Output/rhdf5_test2.h5"
# if (file.exists(fname)) { file.remove(fname) }
#
# h5createFile(fname)
#
# num <- 10
# a <- rep(1, num)
# names(a) <- paste0("M", 1:num)
# print(a) # names in a vector are not saved into hdf5 format (see below)
# h5write(a, fname, "a")
#
# b <- list(M1=1, M2=1)
# h5write(b, fname, "b")
#
# num <- 10
# c <- as.list(rep(1, num))
# names(c) <- paste0("M", 1:num)
# h5write(c, fname, "c")
# + language="R"
#
# fh = H5Fopen(fname)
# print("a = ")
# print(fh$"a")
# print("b = ")
# print(fh$"b")
# print("c = ")
# print(fh$"c")
# h5closeAll()
# -
# We can also read R's lists and vectors in python with `h5py` module:
with h5py.File("Output/rhdf5_test2.h5") as fh:
print(fh["a"][:])
print({x: fh["b"][x][0] for x in fh["b"].keys()})
print({x: fh["c"][x][0] for x in fh["c"].keys()})
# # HighFive package (C++)
#
# Below is the C++ code on how to read and write hdf5 format:
# +
from IPython.display import Markdown as md
with open("src/test_highfive.cpp") as fh:
cpp_file = fh.read()
md(f"```C++\n{cpp_file}```")
# -
# We can open `hdf5` files generated in C++, using `HighFive` package, like so:
with h5py.File("Output/test_highfive.h5") as fh:
print(fh.keys())
print(fh["int"][()])
print(fh["array"][:])
print([x.decode("UTF-8") for x in fh["strings"]])
| main.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Antibiotic Collateral Senstivity is Contingent on the Repeatability of Evolution
#
# Presented below is the Python code used to generate the results presented in the manuscript __Antibiotic Collateral Senstivity is Contingent on the Repeatability of Evolution__. This work appeared also as a preprint: __<NAME>, et al. "Collateral sensitivity is contingent on the repeatability of evolution." bioRxiv (2017): 185892.__
#
# This code is partially adapted from the Github repository with doi:10.5281/zenodo.27481 published alongside: __<NAME> al. "Steering evolution with sequential therapy to prevent the emergence of bacterial antibiotic resistance" PLoS computational biology 11.9 (2015): e1004493__. The details of the modelling are provided in this paper and are covered only briefly in this notebook.
# ## Prerequisites
# +
from copy import deepcopy
import time
import glob
import math
import random
import sys
import os
import matplotlib
import matplotlib.pyplot as plt
from matplotlib import rc
from mpl_toolkits.axes_grid1 import Grid
import numpy as np
import seaborn as sns
sns.set_style('white')
print ("Python version: ", sys.version)
print ("numpy version: ", np.__version__)
print ("matplotlib version: ", matplotlib.__version__)
# %matplotlib inline
# -
# ## Parameters
#
# The $r$ value determines the move rule use in the mathematical model (See Supplementary Note 1). Increasing this value biases the random walk towards genotypes with higher fitness.
r=0 #As used throughout the main text.
# ## Helper functions
#
# Here we represent genotypes as binary strings and landscapes as lists of values. As such, it is useful to define a collection of helper functions to translate between a binary string its equivalent index in the fitness landscape.
# +
#==============================================================================#
# Helper functions
#==============================================================================#
# Computes the hamming distance between two genotypes.
def hammingDistance(s1, s2):
assert len(s1) == len(s2)
return sum(ch1 != ch2 for ch1, ch2 in zip(s1, s2))
# Takes a genotype and converts it to an integer use indexing
# the fitness landscape list
def convertGenotypeToInt(genotype):
out = 0
for bit in genotype:
out = (out << 1) | bit
return out
# Converts an integer to a genotype by taking the binary value and
# padding to the left by 0s
def convertIntToGenotype(anInt, pad):
offset = 2**pad
return [int(x) for x in bin(offset+anInt)[3:]]
x = convertIntToGenotype(10, 20)
y = convertIntToGenotype(12, 20)
print hammingDistance(x, y)
# Function which returns all genotypes at Hamming distance 1 from a
# specified genotype
def getOneStepNeighbours(genotype):
neighbours = []
for x in range(0, len(genotype)):
temp = deepcopy(genotype)
temp[x] = 1 if temp[x] == 0 else 0 # my alternative to Dan's -- slightly less effecient
# temp[x] = (genotype[x]+1) %2 #There is some inefficiency here.
neighbours.append(temp)
return neighbours
def getOneStepNeighbours2(genotype):
neighbours = []
for x in range(0, len(genotype)):
temp = deepcopy(genotype)
# temp[x] = 1 if temp[x] == 0 else 0 # my alternative to Dan's -- unsure if more effecient
temp[x] = (genotype[x]+1) %2 #There is some inefficiency here.
neighbours.append(temp)
return neighbours
# -
# ## A Fitness Landscape Class
# +
#==============================================================================#
# Defining a fitness landscape class
#
# This class represents a fitness landscapes as a list of fitness values
# (self.landscape) and provdes a collection of useful methods of
# querying the landscape
#==============================================================================#
class FitnessLandscape:
def __init__(self, landscapeValues, name=None):
self.landscape = landscapeValues
self.name = name
def getFitness(self, genotype):
fitness = self.landscape[convertGenotypeToInt(genotype)]
return fitness
def genotypeLength(self):
return int(math.log(len(self.landscape), 2))
def numGenotypes(self):
return len(self.landscape)
def isPeak(self, g):
peak = True
for h in getOneStepNeighbours(g):
if self.getFitness(g) < self.getFitness(h):
peak = False
break
return peak
def getPeaks(self):
peaks = []
allGenotypes = []
N =self.genotypeLength()
for x in range(0, 2**N):
allGenotypes.append(convertIntToGenotype(x, self.genotypeLength()))
for g in allGenotypes:
if self.isPeak(g):
peaks.append(g)
return peaks
def getGlobalPeak(self):
return convertIntToGenotype(np.argmax(self.landscape), self.genotypeLength())
def getLowestFitnessPeak(self):
# Finds the peaks of the landscape
peak_genotypes = self.getPeaks()
lowest_peak_genotype = peak_genotypes[
np.argmin([self.getFitness(g) for g in peak_genotypes])]
return lowest_peak_genotype
# landscape.getFitness(x)
AMP = FitnessLandscape([1.851, 2.082, 1.948, 2.434, 2.024, 2.198, 2.033, 0.034,
1.57, 2.165, 0.051, 0.083, 2.186, 2.322, 0.088, 2.821],
"Ampicillin")
test_landscape = [1.851, 2.082, 1.948, 2.434, 2.024, 2.198, 2.033, 0.034,
1.57, 2.165, 0.051, 0.083, 2.186, 2.322, 0.088, 2.821]
print convertGenotypeToInt(x)
print test_landscape[convertGenotypeToInt(x)]
# -
# ## Markov Model
#
# Here we implement the Markov chain model defined in Equation 1.
#==============================================================================#
# Given two genotypes and a landscape, computes the transition probability
# Pr(g1->g2) in the markov chain transition matrix
#==============================================================================#
def transProbSSWM(g1, g2, landscape, r=0):
# Equation 3.1
# If the genotypes are more than one mutation apart, then 0
if hammingDistance(g1,g2) > 1:
return 0
#Else compute Pr(g1->g2) from Equation 3.2 and 3.3
elif hammingDistance(g1,g2) == 1:
#Equation 3.2
if landscape.getFitness(g1) >= landscape.getFitness(g2):
return 0
#Equation 3.3
else:
num = (landscape.getFitness(g2) - landscape.getFitness(g1))**r
den = 0.
for genotype in getOneStepNeighbours(g1):
fitDiff = (landscape.getFitness(genotype) - landscape.getFitness(g1))
if fitDiff > 0:
den += fitDiff**r
return num / den
#Finally add in those Pr(g1->g1)=1 for g1 a local optima (Equation 3.4)
else:
isPeak = landscape.isPeak(g1)
return int(isPeak)
#==============================================================================#
# Builds the transition matrix for a given landscape
#==============================================================================#
def buildTransitionMatrix(landscape, r=0):
genomeLen = landscape.genotypeLength()
matList = [[transProbSSWM(convertIntToGenotype(i,genomeLen),
convertIntToGenotype(j, genomeLen), landscape, r) \
for j in range(0, 2**genomeLen)] for i in range(0, 2**genomeLen)]
return np.matrix(matList)
#==============================================================================#
# Computes P* from P (Equation 3.8)
#
# Given a stochastic matrix P, finds the limit matrix
#==============================================================================#
def limitMatrix(P):
Q = np.identity(len(P))
while not np.array_equal(Q,P):
Q = deepcopy(P)
P = P*P #Square P until it no longer changes.
return P
# ## Model Parametrisation
# The following as the landscapes derived by Mira et al in the following paper:
#
# __Mira, <NAME>., et al. "Rational design of antibiotic treatment plans: a treatment strategy for managing evolution and reversing resistance." PLoS ONE 10.5 (2015): e0122283.__
# +
#==============================================================================#
# The landscapes reported by Mira et. al. (2015)
#==============================================================================#
AMP = FitnessLandscape([1.851, 2.082, 1.948, 2.434, 2.024, 2.198, 2.033, 0.034,
1.57, 2.165, 0.051, 0.083, 2.186, 2.322, 0.088, 2.821],
"Ampicillin")
AM = FitnessLandscape([1.778, 1.782, 2.042, 1.752, 1.448, 1.544, 1.184, 0.063,
1.72, 2.008, 1.799, 2.005, 1.557, 2.247, 1.768, 2.047],
"Amoxicillin")
CEC = FitnessLandscape([2.258, 1.996, 2.151, 2.648, 2.396, 1.846, 2.23, 0.214,
0.234, 0.172, 2.242, 0.093, 2.15, 0.095, 2.64, 0.516],
"Cefaclor")
CTX = FitnessLandscape([0.16, 0.085, 1.936, 2.348, 1.653, 0.138, 2.295, 2.269,
0.185, 0.14, 1.969, 0.203, 0.225, 0.092, 0.119, 2.412],
"Cefotaxime")
ZOX = FitnessLandscape([0.993, 0.805, 2.069, 2.683, 1.698, 2.01, 2.138, 2.688,
1.106, 1.171, 1.894, 0.681, 1.116, 1.105, 1.103, 2.591],
"Ceftizoxime")
CXM = FitnessLandscape([1.748, 1.7, 2.07, 1.938, 2.94, 2.173, 2.918, 3.272, 0.423,
1.578, 1.911, 2.754, 2.024, 1.678, 1.591, 2.923],
"Cefuroxime")
CRO = FitnessLandscape([1.092, 0.287, 2.554, 3.042, 2.88, 0.656, 2.732, 0.436,
0.83, 0.54, 3.173, 1.153, 1.407, 0.751, 2.74, 3.227],
"Ceftriaxone")
AMC = FitnessLandscape([1.435, 1.573, 1.061, 1.457, 1.672, 1.625, 0.073, 0.068,
1.417, 1.351, 1.538, 1.59, 1.377, 1.914, 1.307, 1.728],
"Amoxicillin+Clav")
CAZ = FitnessLandscape([2.134, 2.656, 2.618, 2.688, 2.042, 2.756, 2.924, 0.251,
0.288, 0.576, 1.604, 1.378, 2.63, 2.677, 2.893, 2.563],
"Ceftazidime")
CTT = FitnessLandscape([2.125, 1.922, 2.804, 0.588, 3.291, 2.888, 3.082, 3.508,
3.238, 2.966, 2.883, 0.89, 0.546, 3.181, 3.193, 2.543],
"Cefotetan")
SAM = FitnessLandscape([1.879, 2.533, 0.133, 0.094, 2.456, 2.437, 0.083, 0.094,
2.198, 2.57, 2.308, 2.886, 2.504, 3.002, 2.528, 3.453],
"Ampicillin+Sulbactam")
CPR = FitnessLandscape([1.743, 1.662, 1.763, 1.785, 2.018, 2.05, 2.042, 0.218,
1.553, 0.256, 0.165, 0.221, 0.223, 0.239, 1.811, 0.288],
"Cefprozil")
CPD = FitnessLandscape([0.595, 0.245, 2.604, 3.043, 1.761, 1.471, 2.91, 3.096,
0.432, 0.388, 2.651, 1.103, 0.638, 0.986, 0.963, 3.268],
"Cefpodoxime")
TZP = FitnessLandscape([2.679, 2.906, 2.427, 0.141, 3.038, 3.309, 2.528, 0.143,
2.709, 2.5, 0.172, 0.093, 2.453, 2.739, 0.609, 0.171],
"Piperacillin+Tazobactam")
FEP = FitnessLandscape([2.59, 2.572, 2.393, 2.832, 2.44, 2.808, 2.652, 0.611,
2.067, 2.446, 2.957, 2.633, 2.735, 2.863, 2.796, 3.203],
"Cefepime")
#==============================================================================#
# The limits of the Markov chain matrices corresponding to these landscapes
#==============================================================================#
L_AMP = limitMatrix(buildTransitionMatrix(AMP, r))
L_AM = limitMatrix(buildTransitionMatrix(AM, r))
L_CEC = limitMatrix(buildTransitionMatrix(CEC, r))
L_CTX = limitMatrix(buildTransitionMatrix(CTX, r))
L_ZOX = limitMatrix(buildTransitionMatrix(ZOX, r))
L_CXM = limitMatrix(buildTransitionMatrix(CXM, r))
L_CRO = limitMatrix(buildTransitionMatrix(CRO, r))
L_AMC = limitMatrix(buildTransitionMatrix(AMC, r))
L_CAZ = limitMatrix(buildTransitionMatrix(CAZ, r))
L_CTT = limitMatrix(buildTransitionMatrix(CTT, r))
L_SAM = limitMatrix(buildTransitionMatrix(SAM, r))
L_CPR = limitMatrix(buildTransitionMatrix(CPR, r))
L_CPD = limitMatrix(buildTransitionMatrix(CPD, r))
L_TZP = limitMatrix(buildTransitionMatrix(TZP, r))
L_FEP = limitMatrix(buildTransitionMatrix(FEP, r))
landscapes = [AMP, AM, CEC, CTX, ZOX, CXM, CRO, AMC,
CAZ, CTT, SAM, CPR, CPD, TZP, FEP]
limit_matrices = [L_AMP, L_AM, L_CEC, L_CTX, L_ZOX, L_CXM,
L_CRO, L_AMC, L_CAZ, L_CTT, L_SAM, L_CPR, L_CPD, L_TZP, L_FEP]
labs = ["AMP", "AM", "CEC", "CTX", "ZOX", "CXM",
"CRO", "AMC", "CAZ", "CTT", "SAM", "CPR", "CPD", "TZP", "FEP"]
# -
# ### Plotting the fitness landscapes as bar plots, highlighting fitness peaks:
# +
#==============================================================================#
# Supplementary Figure 1
#==============================================================================#
def lab(k):
gt = convertIntToGenotype(k, 4)
l = "".join(map(str,gt))
return l
def col(ls, k):
r = sns.xkcd_rgb['pale red']
b = sns.xkcd_rgb['denim blue']
if ls.isPeak(convertIntToGenotype(k, 4)):
return sns.xkcd_rgb['pale red']
else:
return sns.xkcd_rgb['denim blue']
fig = plt.figure(figsize=(18,12))
for k, ls in enumerate(landscapes):
plt.subplot(3,5,k+1)
cols = map(lambda k : col(ls, k), range(2**4))
barlist = plt.bar(range(2**4), ls.landscape, color=cols)
plt.ylim(0.0,3.0)
plt.title(labs[k], size=16)
plt.xticks(range(2**4), map(lab, range(2**4)), size=9, rotation='vertical')
plt.yticks(np.arange(0.0,4.1 ,0.5))
plt.subplots_adjust(hspace=0.6)
fig.add_subplot(111, frameon=False)
# hide tick and tick label of the big axes
plt.tick_params(labelcolor='none', top='off', bottom='off', left='off', right='off')
plt.ylabel("Avg. Growth Rate (x10e-3 cells per minute)", size=18)
plt.suptitle("Fitness Landscapes", size=28)
plt.tight_layout(rect=[0, 0.03, 1, 0.95]);
# -
# ## Collateral Sensitivity
#
# In our exploration of collateral sensitivity, we assume that the initial population is a "wild-type" population, mirroring experimental evolution of drug resistance. As such, we specify an initial population as follows
# g=0000 is the initial genotype
init_wt = np.array([1.]+[0. for i in range(15)])
# To determine the collateral response under a second drug $y$, first we randomly select a fitness peak genotype ($g_x^*$) in the first landscape, $x$, arising from evolution from the initial genotype $g_0$. Next, the collateral response is calculated as
#
# $\text{Collateral response of $Y$ to $X$} = \log_2\left( \frac{f_y(g_x^*)}{f_y(g_0)} \right)$
# +
#==============================================================================#
# Plotting functions used to display a table of collateral response
#==============================================================================#
# Displays the matrix/heatmap
def show_CSN(csmat):
fig = plt.figure(figsize=(5.3,4.5))
ax = fig.add_subplot(111)
cax=ax.matshow(csmat, cmap = plt.cm.RdBu_r, vmin = -5, vmax = 5)
cbar = fig.colorbar(cax)
cbar.set_ticks([-5,-4,-3,-2,-1,0,1,2,3,4,5])
cbar.ax.set_yticklabels(['32x','16x','8x','4x','2x','1x',
'2x','4x','8x','16x','32x'])
cbar.set_label('Change from wild-type sensitivity', size=14)
ax.set_xticks(range(len(labs)))
ax.set_yticks(range(len(labs)))
ax.set_xticklabels(labs, rotation=45, size=12)
ax.set_yticklabels(labs, size=12)
ax.axis('image')
ax.set_xlabel('First drug', size=16)
ax.xaxis.set_label_position('top')
ax.set_ylabel('Second drug', size=16)
# Displays a bar chart of showing number of CS/CR drug pairs
def bar_comparison(csmat):
bets, wors = [],[]
for d2 in range(len(landscapes)):
row = np.array(csmat).T[d2]
bets.append(sum(map(lambda x : x<-10**(-8), row)))
wors.append(sum(map(lambda x : x>10**(-8), row)))
bets = map(lambda x : -x, bets)
cmap = plt.cm.RdBu
res_bar_col = cmap(0.1)
sen_bar_col = cmap(0.9)
fig, ax = plt.subplots(figsize=(5,5))
ax = plt.subplot(1,2,1)
rects1 = ax.barh(np.arange(len(landscapes)), wors, color=res_bar_col)
rects2 = ax.barh(np.arange(len(landscapes)), bets, color=sen_bar_col)
plt.xlim(-15,15)
plt.ylim(-1,15)
plt.xticks([-15,-10,-5,0,5,10,15])
ax.set_xticklabels([15,10,5,0,5,10,15], size=12)
plt.xlabel('Number of second drugs', size=16)
plt.ylabel('First drug', size=16)
plt.yticks(np.arange(0.0,len(landscapes)+0.0,1.0), labs)
plt.axvline(0., lw=1.0, ls='-', c='k')
for v in [-10,-5,5,10]:
plt.axvline(v, lw=0.3, ls='-', c='k', zorder=0)
plt.gca().invert_yaxis()
# -
# ### Simulating a table of collateral response with one replicate per drug pair
# +
#==============================================================================#
# Simulates a single instance of determining collateral sensitivity
#
# As evolution is not necessarily repeatable, this is not guaranteed to generate
# the same matrix of collateral sensitivity on each instance.
#==============================================================================#
def col_sensitivity(d1, d2, init_pop, relative=True):
limit_d1 = limit_matrices[d1]
pop_dist = init_pop * limit_d1
pop_dist = np.array(pop_dist)[0]
peak = np.random.choice(np.array([i for i in range(16)]), p=pop_dist)
f_evolved = landscapes[d2].getFitness(convertIntToGenotype(peak,4))
f_wt = landscapes[d2].getFitness(convertIntToGenotype(0,4))
col_sens = np.log2(f_evolved / f_wt)
return col_sens
def generateCSN(init_pop):
network = [[col_sensitivity(d1,d2,init_pop) for d1 in range(len(landscapes))]
for d2 in range(len(landscapes))]
return network
# +
#==============================================================================#
# Example use.
#==============================================================================#
example_csn = generateCSN(init_wt)
with sns.axes_style("white"):
show_CSN(example_csn)
with sns.axes_style("ticks"):
bar_comparison(example_csn)
# -
# ### Exploring the space of collateral sensitivity tables
# The following methods are used to explore the space of collateral sensitivity tables to find the most likely outcome along with the best-case, worst-case and average-case outcome for a table of collateral sensitivity determined from a single round of experimental evolution.
# +
#==============================================================================#
# Determines the most likely CSM and the associated probability
#==============================================================================#
def most_likely(init_pop):
gs,ps = [],[]
for d in range(15):
limit = limit_matrices[d]
pop_dist = np.array(init_pop * limit)[0]
g = np.argmax(pop_dist)
gs.append(g)
ps.append(pop_dist[g])
network = [[np.log2(landscapes[d2].getFitness(convertIntToGenotype(g,4)) \
/landscapes[d2].getFitness(convertIntToGenotype(0,4))) \
for g in gs] \
for d2 in range(15)]
p = np.product(ps)
return p,network
#==============================================================================#
# Determines the best and worst case outcome by means of multiple
# trials.
#==============================================================================#
def worst_mat(init_pop):
def worst_outcome(d1,d2,init_pop):
limit_d1 = limit_matrices[d1]
pop_dist = init_pop * limit_d1
pop_dist = np.array(pop_dist)[0]
wt_f = landscapes[d2].getFitness([0,0,0,0])
#Determine the highest fitness outcome.
worst_f = -np.inf
for i in range(len(pop_dist)):
if pop_dist[i] > 10**(-8): #If its possible
i_f = landscapes[d2].getFitness(convertIntToGenotype(i,4))
if i_f > worst_f:
worst_f = i_f
return np.log2(worst_f/wt_f)
mat =[[worst_outcome(d1,d2,init_pop) \
for d1 in range(len(landscapes))] \
for d2 in range(len(landscapes))]
return mat
def best_mat(init_pop):
def best_outcome(d1,d2,init_pop):
limit_d1 = limit_matrices[d1]
pop_dist = init_pop * limit_d1
pop_dist = np.array(pop_dist)[0]
wt_f = landscapes[d2].getFitness([0,0,0,0])
#Determine the highest fitness outcome.
best_f = np.inf
for i in range(len(pop_dist)):
if pop_dist[i] > 10**(-8): #If its possible
i_f = landscapes[d2].getFitness(convertIntToGenotype(i,4))
if i_f < best_f:
best_f = i_f
return np.log2(best_f/wt_f)
mat =[[best_outcome(d1,d2,init_pop) \
for d1 in range(len(landscapes))] \
for d2 in range(len(landscapes))]
return mat
#==============================================================================#
# Determines the average collateral response for each drug pair
#==============================================================================#
def expected_network(init_pop):
def expected_sensitivity(d1,d2,init_pop):
limit_d1 = limit_matrices[d1]
pop_dist = init_pop * limit_d1
pop_dist = np.array(pop_dist)[0]
expected_evo_f = sum([pop_dist[i] * landscapes[d2].getFitness(convertIntToGenotype(i,4)) \
for i in range(16)])
wt_f = landscapes[d2].getFitness(convertIntToGenotype(0,4))
expected_fitness = np.log2(expected_evo_f/wt_f)
return expected_fitness
network =[[expected_sensitivity(d1,d2,init_pop) \
for d1 in range(len(landscapes))] \
for d2 in range(len(landscapes))]
return network
# -
exp = worst_mat(init_wt)
exp = np.array(exp).T
for ix,row in enumerate(exp):
print labs[ix],(np.array(row)>0).sum(), (np.array(row)<0).sum()
# +
#==============================================================================#
# Generating Figure 2:
#==============================================================================#
exp = expected_network(init_wt)
# best, worst = best_and_worst(1000, init_wt)
worst = worst_mat(init_wt)
best = best_mat(init_wt)
p,net = most_likely(init_wt)
print "The most likely CSM occurs with probability: ", p
directory = './figs'
if not os.path.exists(directory):
os.makedirs(directory)
# Uncomment below to build the parts of Figure 2
# show_CSN(exp)
# plt.savefig('./figs/expected_hm2.svg')
bar_comparison(exp)
plt.savefig('./figs/expected_bars.svg')
# show_CSN(best)
# plt.savefig('./figs/best_hm2.svg')
bar_comparison(best)
plt.savefig('./figs/best_bars.svg')
# show_CSN(worst)
# plt.savefig('./figs/worst_hm2.svg')
bar_comparison(worst)
plt.savefig('./figs/worst_bars.svg')
# show_CSN(net)
# plt.savefig('./figs/ml_hm2.svg')
bar_comparison(net)
plt.savefig('./figs/ml_bars.svg')
# -
# ### Simulating experimental evolution to find CS drugs
# In the manuscript we ask the following two questions:
# 1. What is the probability that a randomly chosen the drug pair exhibits cross resistance?
# 2. Given that a drug pair exhibits collateral sensitivity in a single simulation of experimental evolution, what is it the likelihood that it exhibits cross resistance in second such experiment?
#
# The following functions determine the answers to these questions.
# +
#==============================================================================#
# Computes the likelihood of CLR from a random drug pair
#
# Here we assume that the drug pair is chosen uniformly at random. As such,
# the probability of choosing a CR pair at random can be determined analytically by
# summing, over all possible pairs, the total probability mass of collateral
# resistance and then normalising appropriately.
#==============================================================================#
def random_cl():
tot = 0.
for d1 in range(15):
for d2 in range(15):
if d1!=d2:
temp = 0
wt_f = landscapes[d1].getFitness([0,0,0,0])
ev_pop = init_wt * limit_matrices[d1]
ev_pop = np.array(ev_pop)[0]
for k in range(len(ev_pop)):
ev_f = landscapes[d2].getFitness(convertIntToGenotype(k,4))
if ev_f > wt_f:
temp+=ev_pop[k]
tot+=temp
tot = tot / (15. * 14.)
return tot
#==============================================================================#
# Determines the probability that a pair identified as CS in a single simulation of
# experimental evolution is CR in a second simulation.
#
# This is determined by repeatedly sampling a random matrix of collateral response,
# choosing a CS pair at random and then determining the likelihood that the first of
# that pair induces CR in the second.
#==============================================================================#
def prob_clr():
#generate a random csn
csn = generateCSN(init_wt)
#Pick a pair with CS are random
flag = True
while flag:
d1 = random.randint(0,14)
d2 = random.randint(0,14)
if csn[d1][d2] < 0.0:
flag = False
#find the probability of clr.
prob_res = 0.0
wt_f = landscapes[d1].getFitness([0,0,0,0])
ev_pop = init_wt * limit_matrices[d1]
ev_pop = np.array(ev_pop)[0]
for k in range(len(ev_pop)):
ev_f = landscapes[d2].getFitness(convertIntToGenotype(k,4))
if ev_f > wt_f:
prob_res += ev_pop[k]
return prob_res
#10000 samples:
S = 10**3 #10**6 #In the paper
cr_ps = np.mean([prob_clr() for _ in range(S)])
print "The probability of CR in a random pair is: ", random_cl()
print "The probability of CR in a random pair identified as CS is: ", cr_ps
# -
# ## Collateral Sensitivity Likelihoods
# Here we derive *likelihood of collateral sensitivity* between pairs of drugs.
# +
#==============================================================================#
# Given two drugs, returns the probability that evolution
# under the first results in a genotype with fitness under the
# second that is lower than the WT fitness.
#==============================================================================#
def prob_better(d1,d2,init_pop):
limit_d1 = limit_matrices[d1]
pop_dist = init_pop * limit_d1
pop_dist = np.array(pop_dist)[0]
prob = 0
wt_f = landscapes[d2].getFitness([0,0,0,0])
for i in range(len(pop_dist)):
if landscapes[d2].getFitness(convertIntToGenotype(i,4)) < wt_f:
prob+=pop_dist[i]
return prob
#==============================================================================#
# Collateral Sensitivity Likelihood matrix
#==============================================================================#
def prob_matrix(init_pop):
mat = [[prob_better(d1,d2,init_pop) for d1 in range(len(landscapes))] \
for d2 in range(len(landscapes))]
return mat
#==============================================================================#
# Plots the table of collateral sensitivity likelihood
#==============================================================================#
def show_CSL(mat):
fig = plt.figure(figsize = (6,5))
ax = fig.add_subplot(111)
cax=ax.matshow(mat, cmap = plt.cm.Greens, vmin = 0.0, vmax = 1.0)
cbar = fig.colorbar(cax)
cbar.set_label('Probability of collateral sensitivity', size=14)
ax.set_xticks(range(len(labs)))
ax.set_yticks(range(len(labs)))
ax.set_xticklabels(labs, rotation=45, size=12)
ax.set_yticklabels(labs, size=12)
ax.axis('image')
ax.set_xlabel('First drug', size='16')
ax.xaxis.set_label_position('top')
ax.set_ylabel('Second drug', size='16')
#==============================================================================#
# Returns a CSL in which all entries <p are set to zero
#==============================================================================#
def cut_off_probs_CSL(network, p):
new_net = [[x if p-x<=10**(-8) else 0.0 for x in y] for y in network]
return new_net
# -
#==============================================================================#
# Example use. Figure 5 parts
#==============================================================================#
sns.set_style('white')
csl = prob_matrix(init_wt)
show_CSL(csl) #Uncomment for other figures.
plt.savefig('./figs/csls.svg')
show_CSL(cut_off_probs_CSL(csl, 1.0))
plt.savefig('./figs/csls_1.svg')
show_CSL(cut_off_probs_CSL(csl, 0.75))
plt.savefig('./figs/csls_075.svg')
| model/model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python for Psychologists - Session 3
# ## from Session2: sets
# ## later: handling data with dataframes & pandas
# ### Sets
# Sets are *unordered* and *unindexed* collections of things. Sets also differ from lists in that they cannot contain two identical elements. Sets can be created with the following syntax:
#
# ```python
# my_set = {"element1", element2, 4}
# ```
#
# Create a set of three random inputs.
my_set = {"hallo", 2, 9}
my_set
# Now try to create a set with a duplicate value. Take a look at the resulting set afterwards.
my_set = {"hallo", 2, 9, 9, 1}
my_set
# Try to access the first element of your set.
my_set[0]
# We can also convert lists to sets and vice versa:
#
# ```python
# my_list = [1,2,3,4]
# my_set = set(my_list)
# my_list = list(my_set)
# ```
#
# Take the following list, convert it to a set and back to a list again.
my_list = ["ich", "mag", "sets"]
my_set = set(my_list)
my_set
new_list = list(my_list)
new_list
# **Adding items to a set**
#
# We can add single items to a set using the following syntax:
#
# ```python
# my_set.add(new_item)
# ```
#
# We can also add several items at once:
#
# ```python
# my_set.update({new_item1, new_item2})
# ```
#
# Try to add some items to my_set.
my_set.update({"und", "Listen", "auch"})
my_set
# **Removing items from a set**
#
# Removing items from a set works similarly:
#
# ```python
# my_set.remove(item_to_be_removed)
# ```
#
# Delete one of the items that you have just added.
my_set.remove('Listen')
my_set
# ### handling data
#
# In the last two sessions you learned about the basic principles, data types, variables and how to handle them ... but **most** of the time we do not just work with single list, tuples or whatsoever, but with a bunch of data arranged in logfiles, tables, .csv files ..
#
#
# Today we learn about using **pandas** to ...
# 
#
#
# ... well to actually handle our data. **Pandas** is your data's home and we can get familiar with our data by cleaning, transforming and analyzing it through pandas.
# For getting started, we need to `import pandas as pd ` to use all its provided features. We use ***pd*** as an abbreviation, since we are a bit lazy here :)
import pandas as pd
# Pandas has two core components, i.e., ***series* and *dataframes***. A series is basically a single column, whereas a dataframes is a "multi-dimensional table" made out of a collection of series. Both can contains different kind of data types - for now we will use integers ..
# ----------
# **creating series**
#
# to create a series with any `element`, we can use:
#
# ```python
# s = pd.Series([element1, element2, element3], name="anynameyouwant")
# ```
#
# Try now to create two series representing your two favorite fruits and 6 random integers and check one of them:
#
# +
s1 = pd.Series([3,4,7,8,4,1], name="apples")
s2 = pd.Series([5,9,12,2,9,10], name="bananas")
s1
# -
# As we can see, there is one column (as described above) containing the assigned values, but wait .. why is there another column?
#
# The first column contains the index, in our case we just used the pandas default, that starts again with 0 (remember why?). Consequently, we can again use ```series[1] ``` for indexing the 2 value (row) in our series.
#
# Try to index the last element in one of your fruit series and think about what´s different when we index e.g. lists!
s2[5]
# ----------
# **create dataframes from scratch**
#
# Usually in data analysis we somehow end up with a .csv file from our experiment, but firstly we will learn how to create dataframes from scratch. There are many different ways and this notebook is certainly not exhaustive:
#
# - we can use a dictionary to combine our two fruit series s1 and s2 to get a dataframe "shoppinglist" by using the ```pd.Dataframe(some_data) ``` Dataframe Builder. Here each (key:value) corresponds to a column:
# +
fruits= {"apples" : s1, "bananas" : s2} # first we need to arrange our series in a dictionary
shoppinglist = pd.DataFrame(fruits) # pd.Dataframe(data) conveniently builds a nice looking dataframe for us
shoppinglist # show our shoppinglist
# -
# - another way to combine two series to get a dataframe is ``` pd.concat([seriesA, seriesB]) ``` which concatenates your series. Let´s try to recreate the result displayed above:
pd.concat([s1,s2])
# Oops, something went wrong! Do you have an idea what happened?
#
#
#
# **KEEP IN MIND!**
# (pandas) functions do have a default setting, which might sometimes behave different than expected.
#
# Remember? By checking ```pd.concat? ``` in a code cell we see, that the default option for concatenating two objects is along the axis=0, i.e. along the rows! However, we want to recreate the nice looking dataframe above, which means we need to concat the objects along the column axis (i.e., axis=1) and specify it respectively. Let's see whether this works:
shoppinglist = pd.concat([s1,s2], axis=1)
shoppinglist
# Right now, we are still using the pandas default for our index (i.e., numbers). Let´s say, we want to use customer names as an index:
#
# ```python
# dataframe.set_index([list_of_anything_with_equal_length_to_dataframe])
# ```
#
# Let´s create a list of 6 customers and replace the current indices with this list to see how many fruits each of them is buying at the Wochenmarkt:
customer=["Victoria", "Rhonda", "Elli", "Rebecca", "Lucie", "Isa"]
shoppinglist = shoppinglist.set_index([customer])
shoppinglist
# btw: if you want to check how long your dataframe is, just use ```len(dataframe)``` - pretty easy, huh?
# **Adding columns and rows**
#
# *Columns*
#
# The Wochenmarkt is about to close and all our customers are thrilled by all the last-minute sale offers. All of them are about to buy some plums.
#
# Again, many roads lead to Rome and we will just cover some of them:
#
# - declare a pd.series that is to be converted into a column by just creating a new ``` pd.Series ``` with an equal length and use ``` dataframe["new_column_name"] = pd.Series ```
#
# +
s3 = pd.Series([1,2,3,4,5,6], name="plums", index=customer) ## does not work if indeces do not correspond
shoppinglist["plums"]=s3
shoppinglist
# -
# Since series also contain a column that contains our index (if we don´t define it, pandas will use its default!) the index needs to correspond to the index in our dataframe, otherwise we will create a new column with undefined values (i.e. **N**ot **a** **N**umber, NaN values)
# - this also works with lists and might be a little bit more convenient ``` dataframe["new_column_name"] = [some_list_with_equal_length]``` since lists do not contain an index
#
# Try to add a new column "lemon" with random values for each customer!
shoppinglist["lemon"] = [2,4,5,1,7,9]
shoppinglist
# - if you want more flexibility, you could also use ```dataframe.insert``` to add a list of values to a new column at a specific position just like this:
#
# ```python
# dataframe.insert(position, "column_name", [some_list], True) ## omitting TRUE would raise an error when your
# ## column name already exists in your dataframe
# ```
#
# Try to add a new column "oranges" at the third position with any random integers for all our customers!
shoppinglist.insert(2, "oranges", [1,2,3,4,5,6], True)
shoppinglist
# **adding rows**
#
# Oh hey there, we just met Norbert, who is currently doing a smoothie-detox treatment and do you know what? He also likes apples, bananas, oranges, plums and lemons a lot! Let´s add him to our little dataframe!
# Again, we can use ```pd.DataFrame```to create a new, single-row dataframe for norbert, that contains values for each of our fruits. To combine our two dataframes, our column names in both dataframes need to be identical!
#
# ```python
#
# new_dataframe = pd.DataFrame([some_list_with_equal_length_to_old_df], columns=old_dataframe.columns.tolist())
#
# # list(old_dataframe) conveniently converts your column names into a list, that you can easily pass to your new
# # dataframe
#
# ```
# Try to create a new single-row dataframe called Norbert, that contains values for each fruit and uses the column name information of our shoppinglist dataframe!
norbert = pd.DataFrame([[5,5,4,6,3]], columns=shoppinglist.columns.tolist()) ## list(old_dataframe) works also!
norbert
# Let´s add Norbert to our shoppinglist dataframe! You are already familiar with ```.append ``` for adding new elements to list!
# We can do just the same in our case
# ```python
#
# dataframe.append(new_dataframe)
#
# ```
#
# Lets append Norbert to our dataframe and check our new dataframe!
shoppinglist = shoppinglist.append(norbert)
shoppinglist
# -------
#
# We already learned at the beginning of this session that we can use ```pd.concat([element1, element2])``` for combining two elements. We can use the same command to combine our two dataframes! Keep in mind, that you might have to specify the axis along which we want to add our new dataframe/row
pd.concat([shoppinglist, norbert])
# -------
# **renaming**
#
# What a pity! We forgot to update our index - Norberts name is missing - let´s better change that, before he gets any identity issues!
#
# Do you have an idea how to solve this issue? You essentially already know all the commands to beat the riddler!
#
# - let´s update our customer list
# - let´s set our index
# - let´s check our dataframe
customer.append("Norbert")
shoppinglist = shoppinglist.set_index([customer])
shoppinglist
# Ok, tbh this is probably not the most straight-forward way (some peope would maybe also say it´s not pythonic, btw. if you wanna know that pythonic means, check ```import this```).
#
# Let´s see how we can rename columns or indices in different ways:
#
# - Recap: we just used ```dataframe.column/index.tolist()``` to get a list of our columns/indices --> you already know how to change values in list --> by using ```dataframe.index/columns = your_changed_list``` you can assign new colum or indices
#
#
#
# Try to rename our colum "apples" with a specific kind of apple, e.g. GrannySmith by indexing:
columns =shoppinglist.columns.tolist()
columns[0] = "GrannySmith"
shoppinglist.columns = columns
shoppinglist
# - we can also use ```dataframe.rename(index/column = {"old_value:"new_value"}, inplace=True) ``` to solve the issue in just one single line of code. We define ```inplace=True``` which directly allows us to assign the modification to our dataframe. If we stick to the default (i.e. ```False ```) we would need to assign dataframe = dataframe to "save" our modifications
#
# Let´s try to change one of your customer names:
shoppinglist.rename(index = {"Victoria":"Bianca"}, inplace=True)
shoppinglist
# Besides adding and renaming stuff in our dataframe, we could also delete rows or columns by using ```drop``` :
# ```python
#
# dataframe.drop(index=["element1","element2"])
# dataframe.drop(columns=["element1","element2"])
# ```
#
#
# Try to delete the first customer in your list:
shoppinglist = shoppinglist.drop(index=["Bianca"])
shoppinglist
# **indexing**
#
# We already know from previous sessions, that we can use indexing to assess the first element of a list, the third letter of a string and so on ... in our dataframe universe we can just do the same
#
# *indexing columns or rows*
#
# - the easiest way to index a colum is by using ```dataframe["column"]``` for one column and ```dataframe[["column1", "column2"]] ```for two columns.
#
# Try to index your last two colums:
shoppinglist[["plums", "lemon"]]
# When the index operator ```[]``` is passed a str or int, it attempts to find a column with this particular name and return it as a series ... however if we pass a **slice** to the operator, it changes its behavior and selects rows instead. We can do this with *int* as well *str* !
#
# Try to index all rows expect the fist and last one by using an "int-slicing":
shoppinglist[1:5]
# Try to only show what one customer bought at the Wochenmarkt using "str-slicing":
#
shoppinglist["Elli":"Elli"]
# As the simple index operator ```[] ``` is not that flexible, we will have a look at two other ways to index rows and columns! Today we will get to know two different approaches
#
# - selecting rows and columns by **number** using ```dataframe.iloc[row_selection,column_selection]```
#
# Try to only select the first two rows and all columns:
#
#
shoppinglist.iloc[0:2] # you could als use shoppinglist.iloc[0:2,:] : --> "all"
# Try to select row # 2-4 and column # 3-5!
shoppinglist.iloc[1:4,2:5]
# - selecting rows and colums by label/index
# - selecting rows with a boolean
#
# using ```dataframe.loc[row_selection,column_selection]```
#
# Try to select two rows by using the (customer) index:
#
shoppinglist.loc[["Isa","Elli"]]
# Try to select three customers and two columns of your choice!
shoppinglist.loc[["Isa","Elli","Norbert"], ["bananas", "lemon"]]
# Let´s imagine that you are particularly interested in customers that bought more than 8 bananas or exactly 2 lemons. Such questions and row selecting can be easily done by using conditional selections with booleans in ```dataframe.loc[selection]```. Remember what booleans are about?
#
# If we want select only those customers who bought less than 8 bananas:
shoppinglist.loc[shoppinglist["bananas"] < 8]
# Let´s see how this works: if we use ```dataframe[selection] == some value``` we get a **Pandas Series** with TRUE or FALSE for all our rows:
shoppinglist["bananas"] < 8
# You can also combine two or more conditional statements:
shoppinglist.loc[(shoppinglist["bananas"] < 8) & (shoppinglist["lemon"] == 2)]
| session3/Session3_sets_pandas_dataframes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from luwiji.knn import illustration
# # Avoid the Data Leakage
illustration.data_leakage
# Data Leakage adalah bocornya informasi data test ketika kita training.
# Hal ini paling sering terjadi ketika:
# - imputation
# - kita malah impute menggunakan informasi dari seluruh data, harusnya hanya train data saja
# - solusi: fit_transform pada train, transform pada test
# - scaling
# - kita malah scaling menggunakan informasi dari seluruh data, harusnya hanya train data saja
# - solusi: fit_transform pada train, transform pada test
# - k-fold pada train-test split
# - k-fold menyebabkan kita menyentuh data test yang seharusnya tersembunyi
# - ibaratnya seperti kita boleh retake ujian, ya lama-lama kita jadi tahu soal ujiannya
# - solusi: train-val-test split
#
# Tips menghindari data leakage:
# - Split data sebelum melakukan imputation, scaling
# - Jangan pernah fit apapun selain data train, lalu simpan informasi hasil fit tersebut
# - Gunakan train-val-test split
# # Train-val-test Split
illustration.train_val_test
# # Siapkan data tanpa fillna maupun get_dummies
import pandas as pd
from sklearn.model_selection import train_test_split
df = pd.read_csv("data/titanic.csv", index_col="PassengerId")
df.drop(columns=["Name", "Ticket", "Age", "Cabin"], inplace=True)
# +
X = df.drop(columns="Survived")
y = df.Survived
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, stratify=y, random_state=42)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# -
# ### Preprocessor
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler,
# +
numerical_pipeline = Pipeline([
("imputer", SimpleImputer(strategy='mean')),
("scaler", MinMaxScaler())
])
categorical_pipeline = Pipeline([
("imputer", SimpleImputer(strategy="most_frequent")),
("onehot", OneHotEncoder())
])
# -
from sklearn.compose import ColumnTransformer
X_train.head()
preprocessor = ColumnTransformer([
("numeric", numerical_pipeline, ["SibSp", "Parch", "Fare"]),
("categoric", categorical_pipeline, ["Pclass", "Sex", "Embarked"])
])
# ### Pipeline
from sklearn.neighbors import KNeighborsClassifier
pipeline = Pipeline([
("prep", preprocessor),
("algo", KNeighborsClassifier())
])
# ### GridSearchCV
from sklearn.model_selection import GridSearchCV
# +
parameter = {
"algo__n_neighbors": range(1, 51, 2),
"algo__weights": ["uniform", "distance"],
"algo__p": [1, 2]
}
model = GridSearchCV(pipeline, parameter, cv=3, n_jobs=-1, verbose=1)
model.fit(X_train, y_train)
# -
model.best_params_
# # Summary
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler
# Import data dan drop kolom
df = pd.read_csv("data/titanic.csv", index_col="PassengerId")
df.drop(columns=["Name", "Ticket", "Age", "Cabin"], inplace=True)
# Dataset Splitting
X = df.drop(columns='Survived')
y = df.Survived
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, stratify=y, random_state=42)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# Preprocessor
numerical_pipeline = Pipeline([
("imputer", SimpleImputer(strategy='mean')),
("scaler", MinMaxScaler())
])
categorical_pipeline = Pipeline([
("imputer", SimpleImputer(strategy='most_frequent')),
("onehot", OneHotEncoder())
])
preprocessor = ColumnTransformer([
('numeric', numerical_pipeline, ["SibSp", "Parch", "Fare"]),
('categoric', categorical_pipeline, ["Pclass", "Sex", "Embarked"]),
])
# Pipeline
pipeline = Pipeline([
("prep", preprocessor),
("algo", KNeighborsClassifier())
])
# Parameter Tuning
parameter = {
"algo__n_neighbors": np.arange(1, 51, 2),
"algo__weights": ['uniform', 'distance'],
"algo__p": [1, 2]
}
model = GridSearchCV(pipeline, param_grid=parameter, cv=3, n_jobs=-1, verbose=1)
model.fit(X_train, y_train)
# Evaluation
print(model.best_params_)
print(model.score(X_train, y_train), model.best_score_, model.score(X_test, y_test))
# -
# # Prediction
illustration.jack_and_rose
X.iloc[:1]
data = [
[1, "female", 1, 1, 80, "S"],
[3, "male", 0, 0, 5, "S"]
]
X_pred = pd.DataFrame(data, columns=X.columns, index=["Rose", "Jack"])
X_pred
X_pred["Survived"] = model.predict(X_pred)
X_pred
# # Save Model
from jcopml.utils import save_model, load_model
save_model(model, "knn_titanic.pkl")
save_model(model.best_estimator_, "knn_titanic_small.pkl")
# # Load Model
model_yang_di_load = load_model("model/knn_titanic.pkl")
| 04 - KNN & Scikit-learn/Part 3 - Scikit-learn Pipeline and Workflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:geospatial]
# language: python
# name: conda-env-geospatial-py
# ---
# +
import rioxarray as rx
import geopandas as gpd
from pathlib import Path
import requests as r
import pandas as pd
data_path = Path("data-fire/imagery")
out_path = Path("data-fire/processed")
img_paths = list(data_path.glob("*.img"))
# -
# This throws out about 4 fires out of >45 that don't conform to the same filename type
metadata = sorted([str(path.name).split("_")[:-1] for path in img_paths if len(str(path).split("_")) == 4]) # getting metadata from filenames
df = pd.DataFrame(metadata, columns=['site', 'date', 'product'])
df["datetime"] = pd.to_datetime(df['date'])
df.index = df['datetime']
df = df.sort_index()
df
img_list = []
for i in img_paths:
x = rx.open_rasterio(i)
x.name = i.name
img_list.append(x)
nlcd_2016 = rx.open_rasterio(data_path / "external/")
# Need to check the present day (2016 NLCD) land cover classes for each burned area to make sure it has forest coverage type. Use this filter for selection before downloading gEDI data and store the majority land cover class within the df.
for i in img_list:
i.rio.reproject(4326).rio.to_raster(str(out_path / Path("WGS84_"+str(i.name))))
def bounds_request(bounds):
"""
Requests GEDI L2B data download links using the GEDI Finder service.
https://lpdaacsvc.cr.usgs.gov/services/gedifinder
Args:
bounds returned from rioxarray rAster Array.
It's unpacked to ul_lat,ul_lon,lr_lat,lr_lon
"""
ul_lon = bounds[0]
ul_lat = bounds[-1]
lr_lon = bounds[2]
lr_lat = bounds[1]
result = r.get(f"https://lpdaacsvc.cr.usgs.gov/services/gedifinder?product=GEDI01_B&version=001&bbox=[{ul_lat},{ul_lon},{lr_lat},{lr_lon}]")
print(result)
return result.json()['data']
all_urls = []
for i in img_list:
url_list = bounds_request(i.rio.bounds())
if len(url_list) > 0:
all_urls.extend(url_list)
with open("fire_gedi_url_list.txt", 'w') as output:
for row in all_urls:
output.write(str(row) + '\n')
| fire_regrowth_structure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
pd.set_option('display.max_columns',500)
# ## SIEDU
# ! ls ../
# +
import glob, os
os.chdir("/Users/pipe/Documents/Spike/CR2/datos/INDICADORES_SIEDU/")
siedu_files = {}
siedu_sheetnames = {}
#xl.sheet_names
for file in glob.glob("*.xlsx"):
nom_file = file.split('.')[0]
siedu_sheetnames[nom_file] = pd.ExcelFile('../INDICADORES_SIEDU/'+file)#, sheet_name=nom_file.split(' ')[0]+'_INDICADOR')
print(f'nombre archivo: {nom_file}, sheets {siedu_sheetnames[nom_file].sheet_names}')
for element in siedu_sheetnames[nom_file].sheet_names:
siedu_files[element] = siedu_sheetnames[nom_file].parse(element)
# print(nom_file, siedu[nom_file].sheet_names)
# +
lista_indicadores = [m for m in siedu_files.keys() if 'INDICADOR' in m or 'RESULTADOS' in m]
columnas_comun = ['Región', 'Nombre Ciudad', 'Cod_Ciudad', 'Provincia', 'Área', 'Comuna', 'CUT']
# -
indicadores_raros = []
indicadores_buenis = []
for indicador in lista_indicadores:
if set(columnas_comun).difference(set(siedu_files[indicador].columns)) == set():
siedu_files[indicador].dropna(subset=columnas_comun, inplace=True)
indicadores_buenis += [indicador]
else:
indicadores_raros += [indicador]
for indicador in indicadores_buenis:
if 'Brecha' in siedu_files[indicador].columns:
if 'INDICADOR' in indicador:
tipo = indicador.split('_INDICADOR')[0]
siedu_files[indicador].rename(columns={'Brecha':'Brecha_'+tipo}, inplace=True)
df_aux = siedu_files[indicadores_buenis[0]]
for indicador in indicadores_buenis[1:]:
df_aux = pd.merge(df_aux, siedu_files[indicador], on=columnas_comun,how='left')
df_aux.drop(columns = [m for m in df_aux.columns if 'Unnamed' in m], inplace=True)
df_aux.head(3)
# ### Pasamos todos los campos a minuscula
# +
df_aux.columns = df_aux.columns.str.lower()
for coli in ['región', 'nombre ciudad', 'provincia', 'área', 'comuna']:
df_aux[coli] = df_aux[coli].str.lower()
df_aux.head(3)
# -
df = pd.read_csv('/Users/pipe/Documents/Spike/CR2/datos/Censo2017_16R_ManzanaEntidad_CSV/Censo2017_Manzanas.csv',
sep=';')
numero_habitantes = df.groupby('COMUNA')['PERSONAS'].sum()
numero_habitantes = numero_habitantes.reset_index()
numero_habitantes = numero_habitantes.rename(columns={'COMUNA':'cut', 'PERSONAS': 'num_personas'})
df_aux['cut'] = df_aux['cut'].astype('int')
df_aux.shape[0]
numero_habitantes.shape[0]
df_aux = df_aux.merge(numero_habitantes, on='cut', how='right')
df_aux.shape[0]
df_aux.to_csv('/Users/pipe/Documents/Spike/CR2/calidad_aire_2050_cr2/datos_streamlit/socioeconomicos_por_comuna.csv', index=False)
df_aux.tail()
df_aux.comuna.unique()
df_aux.num_personas.isna().sum()
# +
lista_variables_socio = [ 'num_personas',
'is_34 porcentaje de viviendas con situación de allegamiento externo',
'población total comunal luc_x', 'población usuaria de plazas',
'bpu_28 porcentaje de población atendida por plazas públicas',
'población total comunal luc_y', 'población usuaria de parques',
'bpu_28 porcentaje de población atendida por parques públicas',
'pob_urb_2017', 'pob_urb_afectada',
'ea_48 porcentaje de población expuesta a inundación por tsunami',
'superficie plazas (m2)', 'superficie de area verde (m2)',
'poblacion cpv 2017',
'bpu_29 superficie de area verde por habitantes (m2/hab)',
'bpu_4 razón entre disponibilidad efectiva de matrículas y demanda potencial por educación',
'bpu_3 distancia a e. basica (m)',
'ea_22 consumo per capita residencial (kwh/persona)',
'is_31 porcentaje de viviendas particulares que requieren mejoras de materialidad y/o servicios básicos',
'is_40 porcentaje de manzanas con veredas con buena calidad de pavimento',
'is_33 porcentaje de hacinamiento', 'población área de servicio_x',
'superficie parques m²', 'bpu_23 superficie de parques por habitante',
'bpu_20 distancia a plazas publicas (m)',
'bpu_1 distancia a e. inicial (m)', 'población área de servicio_y',
'superficie plazas m²', 'bpu_21 superficie de plazas por habitantes',
'población 2017', 'lesionados 2018',
'de_31 n° de lesionados en siniestros de tránsito por cada 100.000 habitantes',
'poblacióm 2017', 'fallecidos 2018',
'de_28 n° de víctimas mortales en siniestros de tránsito por cada 100.000 habitantes',
'ig_90 porcentaje de participiación']
porcentajenan = {}
for var_socio in lista_variables_socio:
porcentajenan[var_socio] = df_aux[var_socio].isna().sum()/df_aux.shape[0]
pd.DataFrame().from_dict(porcentajenan, orient='index')
# -
# ### Rellenamos
import geopandas as gpd
comunas = gpd.read_file("/Users/pipe/Documents/Spike/CR2/datos/mapas_censo/Comunas/comunas.shp")
comunas.columns = [m.lower() for m in comunas.columns]
comunas = comunas.to_crs({'init': 'epsg:4326'})
comunas.head(2)
for col in ['comuna']:
comunas[col] = comunas[col].str.lower()
comunas.head(2)
df = df_aux.merge(comunas[['geometry', 'comuna', 'codregion']], on='comuna', how='left')
df.head(1)
lista_variables_numericas = df[list(df.columns)[7:-2]].fillna(0).dtypes[df[list(df.columns)[7:-2]].fillna(0).dtypes == 'float64'].index
lista_variables_numericas
gdf_to_json = gpd.GeoDataFrame(df, geometry='geometry')
# +
gdf_to_json.isna().sum()[gdf_to_json.isna().sum() > 0]
# -
import tqdm
import json
gdf_to_json.query('codregion==@reg')
'R'+str(int(reg))
import numpy as np
gdf_to_json.query('codregion==13').comuna.sort_values().unique()
# +
regiones = df.codregion.unique()
datos_por_region_json = {}
for reg in np.arange(1,17):
datos_por_region_json['R'+str(int(reg))] = json.loads(gdf_to_json.query('codregion==@reg').to_json())
for key in datos_por_region_json.keys():
with open('/Users/pipe/Documents/Spike/CR2/calidad_aire_2050_cr2/datos_git/json_socioeconomicos_'+key+'.json', 'w') as f:
json.dump(datos_por_region_json[key], f)
# -
# !pwd
df.dropna(subset=['codregion'], inplace=True)
df['codregion'] = df['codregion'].astype('int')
df_aux.columns = df_aux.columns.str.lower()
df_aux.to_csv('/Users/pipe/Documents/Spike/CR2/datos/calidad_aire_2050_cr2/datos_streamlit/indicadores_siedu_por_comuna.csv', index=False)
# ### PENDIENTE: seguir con los otros indicadores: indicadores_raros
# # CENSO 2017
# # División del territorio:
# ## División político-administrativa
# Pais > Región > Provincia > Comuna
#
# ## División censal (límites censales definidos por el INE)
# ### Área urbana:
# Comuna > Distrito Censal > Zona > Manzana
# ### Área rural:
# Comuna > Localidad > Entidad
# ! pwd
import pandas as pd
df = pd.read_csv('/Users/pipe/Documents/Spike/CR2/datos/Censo2017_16R_ManzanaEntidad_CSV/Censo2017_Manzanas.csv',
sep=';')
df.head(2)
df.replace({'*':0}, inplace=True)
df['PUEBLO'] = df['PUEBLO'].astype('int')
df['PERSONAS'] = df['PERSONAS'].astype('int')
df['INMIGRANTES'] = df['INMIGRANTES'].astype('int')
numero_habitantes = df.groupby('COMUNA')['PERSONAS'].sum()
datos_socioec = df.groupby('COMUNA')[['PERSONAS','PUEBLO','INMIGRANTES']].agg({'PERSONAS':'sum', 'PUEBLO':'sum', 'INMIGRANTES':'sum',}).reset_index()
datos_socioec.head(2)
datos_socioec.to_csv('/Users/pipe/Documents/Spike/CR2/datos/calidad_aire_2050_cr2/datos_streamlit/indicadores_CENSO17_por_comuna.csv', index=False)
# diccionario_columnas = {'REGION': 'código de cada región, de 1 a 16',
# 'PROVINCIA': 'concatena región + número de provincia',
# 'COMUNA': 'concatena región, provincia y comuna',
# 'DC' : 'distrito censal' (99 = missing value),
# 'AREA' : '1: urbana, 2: rural',
# 'ZC_LOC' : 'zona censal' o 'localidad' (999=missingf_value),
# 'MZ_ENT' : 'manzana' o 'entidad',
# 'ID_ZONA_LOC' : 'el id de la zona o de la localidad',
# 'ID_MANZENT' : 'el id de la manzana o la entidad',
# 'PERSONAS' : 'cantidad de personas',
# 'HOMBRES' : 'cantidad de hombres',
# 'MUJERES' : 'cantidad de mujeres',
# 'EDAD_xAy' : 'cantidad de gente con edad entre x e y',
# 'INMIGRANTES' : 'Total de personas migrantes',
# 'PUEBLO' : 'Total de personas que se consideran pertenecientes a un pueblo indígena u originario',
# 'VIV_PART' : 'Total de viviendas particulares',
# 'VIV_COL' : 'Total de viviendas colectivas',
# 'VPOMP' : 'Total de viviendas particulares ocupadas con moradores presentes',
# 'TOTAL_VIV' : 'Total de viviendas',
# 'CANT_HOG' : 'cantidad de hogares por vivienda',
# 'P01' : 'tipo de vivienda (1 casa, 2 depto, etc)'
# 'P03A' : 'material muros exteriores'
# 'P03B' : 'material cubierta techo'
# 'P03C' : 'material construcción del piso'
# 'P05' : 'origen del agua'
# 'MATACEP' : 'Total de viviendas con materialidad aceptable',
# 'MATREC' : 'Total de viviendas con materialidad aceptable',
# 'MATIRREC' : 'total de viviendas con materialidad irrecuperable',
# }
#
| old_notebooks/1.2 Generar_datos_socioeconomicos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from ase.build import bulk
from jsonextended import edict
from ipyatom import repeat_cell
from ipyatom import render_ivol
from ipyatom import bonds
from ipyatom import plot_mpl
atoms = bulk("Fe").repeat((2,1,1))
atoms
vstruct = repeat_cell.atoms_to_dict(atoms)
vstruct["elements"][0]["sites"][0]["label"] = "S"
vstruct = repeat_cell.change_site_variable(
vstruct, {"color_fill": "green"}, {"label": "S"})
edict.pprint(vstruct)
new_struct, fig, controls = render_ivol.create_ivol(vstruct)
render_ivol.ipy_style(fig)
vstruct2 = bonds.add_bonds(vstruct, "Fe", "S", 2.5)
edict.pprint(vstruct2)
new_struct, fig, controls = render_ivol.create_ivol(vstruct2)
render_ivol.ipy_style(fig)
vstruct3 = bonds.add_bonds(vstruct, "Fe", "S", 2.5, color_by_dist=True, drange=(2,4))
edict.pprint(vstruct3)
new_struct, fig, controls = render_ivol.create_ivol(vstruct3)
render_ivol.ipy_style(fig)
# %matplotlib inline
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1)
plot_mpl.create_colorbar(ax, (2, 4), "jet", "Bond Length")
fig.set_size_inches((5,1))
plot_mpl.plot_atoms_top(vstruct)
| example/Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # A Simple Image Classification Problem using Keras (dog_vs_cat)
# import the necessary packages
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from keras.models import Sequential
from keras.layers import Activation
from keras.optimizers import SGD
from keras.layers import Dense
from keras.utils import np_utils
from imutils import paths
import numpy as np
import argparse
import os
import cv2
import pandas as pd
import numpy as np
def image_to_feature_vector(image, size=(32, 32)):
# resize the image to a fixed size, then flatten the image into
# a list of raw pixel intensities
return cv2.resize(image, size).flatten()
# +
import glob
print("[INFO] describing images...")
train_image_path = "data/train/"
image_paths = glob.glob(os.path.join(train_image_path, '*.jpg'))
# initialize the data matrix and labels list
data = []
labels = []
# -
# loop over the input images
for (i, imagePath) in enumerate(image_paths):
# load the image and extract the class label (assuming that our
# path as the format: /path/to/dataset/{class}.{image_num}.jpg
image = cv2.imread(imagePath)
label = imagePath.split(os.path.sep)[-1].split(".")[0]
# construct a feature vector raw pixel intensities, then update
# the data matrix and labels list
features = image_to_feature_vector(image)
data.append(features)
labels.append(label)
# show an update every 1,000 images
if i > 0 and i % 1000 == 0:
print("[INFO] processed {}/{}".format(i, len(image_paths)))
# ### Data Preprocessing
# +
# encode the labels, converting them from strings to integers
le = LabelEncoder()
encoded_labels = le.fit_transform(labels)
pd.DataFrame(encoded_labels).head(5)
print(pd.DataFrame(labels).describe())
normalized_data = np.array(data) / 255.0
categorical_labels = np_utils.to_categorical(encoded_labels, 2)
# -
# partition the data into training and testing splits, using 75%
# of the data for training and the remaining 25% for testing
print("[INFO] constructing training/testing split...")
labels = categorical_labels.tolist
(trainData, testData, trainLabels, testLabels) = train_test_split(data, categorical_labels, test_size=0.25, random_state=42)
# ### Define an architecture - > Feed Forward Network of dimension "3072-768-384-2"
model = Sequential()
model.add(Dense(768, input_dim=3072, kernel_initializer="uniform", activation="relu"))
model.add(Dense(384, kernel_initializer="uniform", activation="relu"))
model.add(Dense(2))
model.add(Activation("softmax"))
# train the model using SGD
print("[INFO] compiling model...")
sgd = SGD(lr=0.001)
model.compile(loss="binary_crossentropy", optimizer=sgd, metrics=["accuracy"])
model.fit(np.array(trainData), np.array(trainLabels), epochs=50, batch_size=128)
# show the accuracy on the testing set
print("[INFO] evaluating on testing set...")
(loss, accuracy) = model.evaluate(np.array(testData), np.array(testLabels), batch_size=150, verbose=1)
print("[INFO] loss={:.4f}, accuracy: {:.4f}%".format(loss, accuracy * 100))
| notebook_gallery/jupyter/deep_learning/image_classification_dog_vs_cat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import datetime
from sklearn import preprocessing
from datetime import datetime
from sklearn.ensemble import RandomForestClassifier
from sklearn import neighbors
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.svm import SVC
import operator
from pandas_datareader import data, wb
# from sklearn.qda import QDA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from dateutil import parser
# from backtest import Strategy, Portfolio
# +
def getStock(symbol, start, end):
df = data.get_data_yahoo(symbol, start, end)
df.columns = df.columns + '_' + symbol
df['Return_%s' %symbol] = df['Adj Close_%s' %symbol].pct_change()
return df
# -
start = '2014-1-2'
end = '2016-2-4'
goog = getStock('GOOG',start,end)
goog.head()
def addFeatures(dataframe, adjclose, returns, n):
"""
operates on two columns of dataframe:
- n >= 2
- given Return_* computes the return of day i respect to day i-n.
- given AdjClose_* computes its moving average on n days
"""
return_n = adjclose[9:] + "Time" + str(n)
dataframe[return_n] = dataframe[adjclose].pct_change(n)
roll_n = returns[7:] + "RolMean" + str(n)
dataframe[roll_n] = pd.rolling_mean(dataframe[returns], n)
def applyRollMeanDelayedReturns(datasets, delta):
"""
applies rolling mean and delayed returns to each dataframe in the list
"""
for dataset in datasets:
columns = dataset.columns
adjclose = columns[-2]
returns = columns[-1]
for n in delta:
addFeatures(dataset, adjclose, returns, n)
return datasets
def mergeDataframes(datasets, index, cut):
"""
merges datasets in the list
"""
subset = []
subset = [dataset.iloc[:, index:] for dataset in datasets[1:]]
first = subset[0].join(subset[1:], how = 'outer')
finance = datasets[0].iloc[:, index:].join(first, how = 'left')
finance = finance[finance.index > cut]
return finance
def applyTimeLag(dataset, lags, delta):
"""
apply time lag to return columns selected according to delta.
Days to lag are contained in the lads list passed as argument.
Returns a NaN free dataset obtained cutting the lagged dataset
at head and tail
"""
dataset.Return_Out = dataset.Return_Out.shift(-1)
maxLag = max(lags)
columns = dataset.columns[::(2*max(delta)-1)]
for column in columns:
for lag in lags:
newcolumn = column + str(lag)
dataset[newcolumn] = dataset[column].shift(lag)
return dataset.iloc[maxLag:-1,:]
import tensorflow as tf
tf.__version__
import tensorflow.keras as keras
import tensorflow.contrib.keras as keras
model = keras.models.Sequential()
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Dependences
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# +
# Take in all of our CitiBike data and read it into pandas
jan_2019 = "../Resources/JC-201901-citibike-tripdata 4 2.csv"
feb_2019 = "../Resources/JC-201902-citibike-tripdata.csv"
mar_2019 = "../Resources/JC-201903-citibike-tripdata.csv"
apr_2019 = "../Resources/JC-201904-citibike-tripdata.csv"
may_2019 = "../Resources/JC-201905-citibike-tripdata.csv"
june_2019 = "../Resources/JC-201906-citibike-tripdata.csv"
july_2019 = "../Resources/JC-201907-citibike-tripdata.csv"
aug_2019 = "../Resources/JC-201908-citibike-tripdata 2.csv"
#Create DataFrames for all the data files
jan_2019_df = pd.read_csv(jan_2019)
feb_2019_df = pd.read_csv(feb_2019)
mar_2019_df = pd.read_csv(mar_2019)
apr_2019_df = pd.read_csv(apr_2019)
may_2019_df = pd.read_csv(may_2019)
june_2019_df = pd.read_csv(june_2019)
july_2019_df = pd.read_csv(july_2019)
aug_2019_df = pd.read_csv(aug_2019)
# +
#Merge all data from 2019 into one big data frame
combined_CityBike_df = pd.merge(jan_2019_df, feb_2019_df, how='outer')
combined_CityBike_df3 = pd.merge(combined_CityBike_df,mar_2019_df, how = "outer")
combined_CityBike_df4 = pd.merge(combined_CityBike_df3,apr_2019_df, how = "outer")
combined_CityBike_df5 = pd.merge(combined_CityBike_df4,may_2019_df, how = "outer")
combined_CityBike_df6 = pd.merge(combined_CityBike_df5,june_2019_df, how = "outer")
combined_CityBike_df7 = pd.merge(combined_CityBike_df6,july_2019_df, how = "outer")
combined_CityBike_df8 = pd.merge(combined_CityBike_df7,aug_2019_df, how = "outer")
combined_CityBike_df8
# -
| CitiBike/CitiBike 2019 Merge Data.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# ## Requirements
#
# * set directory in the next cell
datadir = "../Datasets/"
using LightGraphs
using DataFrames
using CSV
using Statistics
using PyPlot
using Chain
using StatsBase
using GraphPlot
using Random
# ## Useful functions
#
## k_nn^{mode1,mode2}(l) : average mode2-degree of mode1-neighbours of nodes with mode1-degree = l
# normally mode1 and mode2 should be: degree, indegree or outdegree
deg_corr_directed(G::SimpleDiGraph, mode1::Function, mode2::Function) =
@chain edges(G) begin
DataFrame
transform(:src => ByRow(x -> mode1(G, x)) => :src_deg,
:dst => ByRow(x -> mode2(G, x)) => :dst_deg)
groupby(:src_deg, sort=true)
combine(:dst_deg => mean)
end
## degree correlation for neutral graphs: <k^2>/<k>
function deg_corr_neutral(G, mode::Function)
x = mode(G)
return mean(x .^ 2)/mean(x)
end
# undirected, or default mode=='all' if G is directed
deg_corr(G::SimpleGraph) =
@chain edges(G) begin
DataFrame
append!(_, select(_, :dst => :src, :src => :dst)) # add edge in reverse
transform(:src => ByRow(x -> degree(G, x)) => :src_deg,
:dst => ByRow(x -> degree(G, x)) => :dst_deg)
groupby(:src_deg, sort=true)
combine(:dst_deg => mean)
end
assortativity(G) =
@chain edges(G) begin
DataFrame
append!(_, select(_, :dst => :src, :src => :dst)) # add edge in reverse
transform(:src => ByRow(x -> degree(G, x)) => :src_deg,
:dst => ByRow(x -> degree(G, x)) => :dst_deg)
cov(_.src_deg, _.dst_deg, corrected=false) /
(std(_.src_deg, corrected=false) * std(_.dst_deg, corrected=false))
end
## Correlation exponent via linear regression (taking logs)
function corr_exp(G)
## compute knn's
knn = deg_corr(G)
# Fit the regression
x = log.(knn.src_deg)
y = log.(knn.dst_deg_mean)
return ([ones(length(x)) x] \ y)[2]
end
function richClub(g, l=1)
l_max = maximum(degree(g))
c = countmap(degree(g))
n = nv(g)
moment = sum(k * ck / n for (k, ck) in pairs(c)) ^ 2
S = [k * ck / n for (k, ck) in pairs(c) if k >= l]
phi_hat = sum(x * y for x in S, y in S) * ne(g) / moment
G = induced_subgraph(g, findall(>=(l), degree(g)))[1]
phi = ne(G)
return phi / phi_hat
end
function cm_simple(ds)
@assert iseven(sum(ds))
stubs = reduce(vcat, fill(i, ds[i]) for i in 1:length(ds))
shuffle!(stubs)
local_edges = Set{Tuple{Int, Int}}()
recycle = Tuple{Int,Int}[]
for i in 1:2:length(stubs)
e = minmax(stubs[i], stubs[i+1])
if (e[1] == e[2]) || (e in local_edges)
push!(recycle, e)
else
push!(local_edges, e)
end
end
# resolve self-loops and duplicates
last_recycle = length(recycle)
recycle_counter = last_recycle
while !isempty(recycle)
recycle_counter -= 1
if recycle_counter < 0
if length(recycle) < last_recycle
last_recycle = length(recycle)
recycle_counter = last_recycle
else
break
end
end
p1 = popfirst!(recycle)
from_recycle = 2 * length(recycle) / length(stubs)
success = false
for _ in 1:2:length(stubs)
p2 = if rand() < from_recycle
used_recycle = true
recycle_idx = rand(axes(recycle, 1))
recycle[recycle_idx]
else
used_recycle = false
rand(local_edges)
end
if rand() < 0.5
newp1 = minmax(p1[1], p2[1])
newp2 = minmax(p1[2], p2[2])
else
newp1 = minmax(p1[1], p2[2])
newp2 = minmax(p1[2], p2[1])
end
if newp1 == newp2
good_choice = false
elseif (newp1[1] == newp1[2]) || (newp1 in local_edges)
good_choice = false
elseif (newp2[1] == newp2[2]) || (newp2 in local_edges)
good_choice = false
else
good_choice = true
end
if good_choice
if used_recycle
recycle[recycle_idx], recycle[end] = recycle[end], recycle[recycle_idx]
pop!(recycle)
else
pop!(local_edges, p2)
end
success = true
push!(local_edges, newp1)
push!(local_edges, newp2)
break
end
end
success || push!(recycle, p1)
end
g = SimpleGraph(length(ds))
for e in local_edges
add_edge!(g, e...)
end
return g
end
# ## US Airport Volume of Passengers
#
# same data as in previous chapter, directed weighted graph (passenger volumes)
# +
## read edges and build weighted directed graph
D = CSV.read(datadir * "Airports/connections.csv", DataFrame)
id2name = sort!(unique(union(D.orig_airport, D.dest_airport)))
name2id = Dict(id2name .=> axes(id2name, 1))
g = SimpleDiGraph(length(id2name))
for row in eachrow(D)
from = name2id[row.orig_airport]
to = name2id[row.dest_airport]
from == to || add_edge!(g, from, to)
end
g
# -
# ## Directed Degree Correlation Functions (4 cases)
#
# We consider the 4 combinations in/out vs in/out degrees
#
# Dashed lines are for neutral graphs
knn = deg_corr_directed(g,indegree,indegree)
r = deg_corr_neutral(g,indegree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
ylabel("k_nn(l)",fontsize=12);
knn = deg_corr_directed(g,indegree,outdegree)
r = deg_corr_neutral(g,outdegree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
ylabel("k_nn(l)",fontsize=12);
knn = deg_corr_directed(g,outdegree,indegree)
r = deg_corr_neutral(g,indegree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
ylabel("k_nn(l)",fontsize=12);
knn = deg_corr_directed(g,outdegree,outdegree)
r = deg_corr_neutral(g,outdegree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
ylabel("k_nn(l)",fontsize=12);
# # We consider the undirected airport graph from now on
#
## Undirected graph
g = SimpleGraph(g)
# ## Degree correlation: also look via log scale
knn = deg_corr(g)
r = deg_corr_neutral(g, degree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
xlabel("degree l", fontsize=12)
ylabel("k_nn(l)",fontsize=12);
loglog(eachcol(knn)...,"o",c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
xlabel("degree l", fontsize=12);
# ## State by state assortativity and correlation exponent
A = CSV.read(datadir * "Airports/airports_loc.csv", DataFrame)
A.id = [name2id[a] for a in A.airport]
@assert A.id == axes(A, 1)
@assert A.airport == id2name
first(A, 5)
## for each state compute degree assortativity (r)
## note that we drop airports w/o in-state edge
## also estimate correlation exponent (mu) via regression (taking the logs)
## Show assortativity and mu for states with 5+ vertices
P = DataFrame(state=String[], nodes=Int[], edges=Int[], assortativity=Float64[], mu=Float64[])
for s in unique(A.state)
hva = findall(==(s), A.state)
G = induced_subgraph(g, hva)[1]
G = induced_subgraph(G, findall(>(0), degree(G)))[1]
if nv(G) > 5
mu = corr_exp(G)
push!(P, [s, nv(G), ne(G), assortativity(G), mu])
end
end
sort!(P, :assortativity)
first(P, 5)
last(P, 5)
## some states are quite small,
## but we still see good correlation between r and mu
plot(P.assortativity,P.mu,"o",color="black")
xlabel("degree correlation coefficient (r)",fontsize=12)
ylabel("correlation exponent (mu)",fontsize=12)
println("Person correlation: ",cor(P.assortativity,P.mu))
ident = [-1.0, 1.0]
plot(ident,ident,":",c="gray");
# ## Looking at a few states with high/low assortativity
## positive case: the Dakotas (ND+SD)
hva = findall(in(["SD", "ND"]), A.state)
G_D = induced_subgraph(g, hva)[1]
G_D = induced_subgraph(G_D, findall(>(0), degree(G_D)))[1]
println("r = ", assortativity(G_D))
Random.seed!(4)
gplot(G_D,
NODESIZE=0.03, nodefillc="black",
EDGELINEWIDTH=0.2, edgestrokec="gray")
## compare r and mu vs random models for the Dakotas: G = SD+ND
## here we use the configuration model
## we also report the proportion of nodes above the structural cutoff
## given the degree distribution.
r = Float64[]
mu = Float64[]
for i in 1:1000
cm = cm_simple(degree(G_D))
push!(r, assortativity(cm))
push!(mu, corr_exp(cm))
end
## structural cutoff
sc = sqrt(2*ne(G_D))
p = count(>(sc), degree(G_D)) / nv(G_D)
println("r = ",assortativity(G_D), " mu = ",corr_exp(G_D), " P(edges above structural cutoff) = ", p)
boxplot([r,mu],labels=["assortativity (r)", "correlation exponent (mu)"],widths=.7,sym="");
## degree correlation function for the Dakotas graph
knn = deg_corr(G_D)
r = deg_corr_neutral(G_D, degree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
xlabel("degree (k)",fontsize=12)
ylabel("k_nn(k)",fontsize=12);
## degree correlation function for a configuration model random graph used above
## quite different!
cm = cm_simple(degree(G_D))
knn = deg_corr(cm)
r = deg_corr_neutral(cm, degree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
xlabel("degree (k)",fontsize=12)
ylabel("k_nn(k)",fontsize=12);
## negative case: the MO graph
hva = findall(==("MO"), A.state)
G_D = induced_subgraph(g, hva)[1]
G_D = induced_subgraph(G_D, findall(>(0), degree(G_D)))[1]
println("r = ", assortativity(G_D))
Random.seed!(4)
gplot(G_D,
NODESIZE=0.03, nodefillc="black",
EDGELINEWIDTH=0.2, edgestrokec="gray")
## r and mu vs random configuration model for MO graph
## compare r and mu vs random models for the Dakotas: G = SD+ND
## here we use the configuration model
## we also report the proportion of nodes above the structural cutoff
## given the degree distribution.
r = Float64[]
mu = Float64[]
for i in 1:1000
cm = cm_simple(degree(G_D))
push!(r, assortativity(cm))
push!(mu, corr_exp(cm))
end
## structural cutoff
sc = sqrt(2*ne(G_D))
p = count(>(sc), degree(G_D)) / nv(G_D)
println("r = ",assortativity(G_D), " mu = ",corr_exp(G_D), " P(edges above structural cutoff) = ", p)
boxplot([r,mu],labels=["assortativity (r)", "correlation exponent (mu)"],widths=.7,sym="");
## degree correlation function for MO graph
knn = deg_corr(G_D)
r = deg_corr_neutral(G_D, degree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
xlabel("degree (k)",fontsize=12)
ylabel("k_nn(k)",fontsize=12);
## degree correlation function for a configuration random graph
## quite similar!
cm = cm_simple(degree(G_D))
knn = deg_corr(cm)
r = deg_corr_neutral(cm, degree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
xlabel("degree (k)",fontsize=12)
ylabel("k_nn(k)",fontsize=12);
## state with r = -1 (NE)
hva = findall(==("NE"), A.state)
G_D = induced_subgraph(g, hva)[1]
G_D = induced_subgraph(G_D, findall(>(0), degree(G_D)))[1]
println("r = ", assortativity(G_D))
Random.seed!(4)
gplot(G_D,
NODESIZE=0.03, nodefillc="black",
EDGELINEWIDTH=0.2, edgestrokec="gray")
## state with r = +1 (AR)
hva = findall(==("AR"), A.state)
G_D = induced_subgraph(g, hva)[1]
G_D = induced_subgraph(G_D, findall(>(0), degree(G_D)))[1]
println("r = ", assortativity(G_D))
Random.seed!(4)
gplot(G_D,
NODESIZE=0.03, nodefillc="black",
EDGELINEWIDTH=0.2, edgestrokec="gray")
# ## Back to Overall US Airport graph
#
# - friendship paradox illustration
# - looking for rich club phenomenon
#
## plot degree vs avg neighbour degree
## friendship 'paradox' US Airport graph (overall)
deg = degree(g)
nad = [mean(degree(g, neighbors(g, v))) for v in 1:nv(g)]
scatter(deg,nad,c="black",marker=".")
xlim((0,200))
ylim((0,200))
xlabel("node degree", fontsize=14)
ylabel("average neighbour degree", fontsize=14);
plot([0,200],[0,200], "--", c="gray")
print("r = ", assortativity(g));
## looking for rich club -- not here!
d = unique(degree(g))
rc = richClub.(Ref(g), d)
semilogx(d,rc,".",c="black")
xlabel("degree l",fontsize=12)
ylabel("rich club coefficient rho(l)");
# ## A quick look: Europe electric grid network
#
# We notice:
#
# - degree distribution quite uniform
# - positive assortativity, also seen with degree correlation function
# - friendship paradox not obvious
## Europe Electric Grid
edge_list = split.(readlines(datadir * "GridEurope/gridkit_europe-highvoltage.edges"))
vertex_ids = unique(reduce(vcat, edge_list))
vertex_map = Dict(vertex_ids .=> 1:length(vertex_ids))
grid = SimpleGraph(length(vertex_ids))
foreach(((from, to),) -> add_edge!(grid, vertex_map[from], vertex_map[to]), edge_list)
# +
## plot degree vs avg neighbour degree
deg = degree(grid)
nad = [mean(degree(grid, neighbors(grid, v))) for v in 1:nv(grid)]
scatter(deg,nad,c="black",marker=".")
xlim((0,18))
ylim((0,18))
xlabel("node degree", fontsize=14)
ylabel("average neighbour degree", fontsize=14);
plot([0,18],[0,18], "--", c="gray")
print("r = ", assortativity(grid));
# +
## Degree correlation function
knn = deg_corr(grid)
r = deg_corr_neutral(grid, degree)
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
xlabel("degree (k)",fontsize=12)
ylabel("k_nn(k)",fontsize=12);
# -
# ## Quick look: GitHub Developers Graph
#
# - negative assortativity
# - strong friendship paradox phenomenon
## GitHub Developers (undirected)
D = CSV.read(datadir * "GitHubDevelopers/musae_git_edges.csv", DataFrame) .+ 1
max_node_id = max(maximum(D.id_1), maximum(D.id_2))
git = SimpleGraph(max_node_id)
foreach(row -> add_edge!(git, row...), eachrow(D))
## plot degree vs avg neighbour degree
## zoom in on nodes with degree < LIM
LIM = 1000
deg = degree(git)
nad = [mean(degree(git, neighbors(git, v))) for v in 1:nv(git)]
scatter(deg,nad,c="black",marker=".")
xlim((0,LIM))
ylim((0,LIM))
xlabel("node degree", fontsize=14)
ylabel("average neighbour degree", fontsize=14);
plot([0,LIM],[0,LIM], "--", c="gray")
print("r = ", assortativity(git));
## degree correlation function
knn = deg_corr(git)
r = deg_corr_neutral(git, degree)
xlim((0,LIM))
scatter(eachcol(knn)..., c="black")
hlines(y=r,xmin=minimum(knn.src_deg),xmax=maximum(knn.src_deg),linestyles=":")
xlabel("degree (k)",fontsize=12)
ylabel("k_nn(k)",fontsize=12);
## still no rich club group here
d = unique(degree(git))
rc = richClub.(Ref(git), d)
semilogx(d,rc,".",c="black")
xlabel("degree l",fontsize=12)
ylabel("rich club coefficient rho(l)");
# ## Showing a rich club: the actors graph
#
# This data set is part of the accompanying material of the book "Complex Networks: Principles, Methods and Applications", <NAME>, <NAME>, <NAME>, Cambridge University Press (2017)
#
# 248,243 nodes (actors) and 8,302,734 edges (co-appearing in at least 1 movie)
#
# +
D = CSV.read(datadir * "Actors/movie_actors.net", header=[:id_1, :id_2], DataFrame) .+ 1
max_node_id = max(maximum(D.id_1), maximum(D.id_2))
g = SimpleGraph(max_node_id)
foreach(row -> add_edge!(g, row...), eachrow(D))
g = induced_subgraph(g, findall(>(0), degree(g)))[1]
d = sample(unique(degree(g)), 200, replace=false)
rc = richClub.(Ref(g), d)
semilogx(d,rc,".",c="black")
xlabel("degree l",fontsize=12)
ylabel("rich club coefficient rho(l)");
# -
# ## Figures 4.1 and 4.2: Xulvi-Brunet and Sokolov algorithm
#
# Version in book obtained with faster Julia code. We show a smaller scale example here.
#
# +
## Naive Xulvi-Brunet and Sokolov algorithm
function XBS(n, λ, q, assortative, seed)
p = λ / n
Random.seed!(seed)
g = erdos_renyi(n, p)
q == 0 && return g
e = [(x.src, x.dst) for x in edges(g)]
touched = falses(length(e))
count_touched = 0
while count_touched < length(e)
i, j = rand(axes(e, 1)), rand(axes(e, 1))
if i != j
vs = [e[i]..., e[j]...]
if allunique(vs)
if rand() < q
sort!(vs, by=x -> degree(g, x))
if !assortative
vs[2], vs[4] = vs[4], vs[2]
end
else
shuffle!(vs)
end
a1, a2, b1, b2 = vs
if ((a1, a2) == e[i] || (a1, a2) == e[j])
count_touched += !touched[i] + !touched[j]
touched[i] = true
touched[j] = true
else
if !has_edge(g, a1, a2) && !has_edge(g, b1, b2)
@assert rem_edge!(g, e[i]...)
@assert rem_edge!(g, e[j]...)
e[i] = (a1, a2)
e[j] = (b1, b2)
add_edge!(g, a1, a2)
add_edge!(g, b1, b2)
count_touched += !touched[i] + !touched[j]
touched[i] = true
touched[j] = true
end
end
end
end
end
@assert all(touched)
return g
end
# +
## Fig 4.2 with N nodes and averaging Rep results
N = 2^9 ## we use 2^16 and Rep=64 in book
Rep = 8
seeds = rand(UInt64, Rep)
df = DataFrame()
for seed in seeds
for (q, a) in ((0.0, true), (1/3, true), (2/3, true), (1/3, false), (2/3, false))
for d in 0.25:0.25:3
g = XBS(2^9, d, q, a, seed)
c = maximum(length, connected_components(g)) / nv(g)
push!(df, (seed=seed, q=q, a=a, d=d, component=c))
end
end
end
@chain df begin
groupby([:q, :a, :d])
combine(:component => mean => :component)
groupby([:q, :a])
foreach((c, s, sdf) -> plot(sdf.d, sdf.component, color=c, linestyle=s),
["black", "gray", "black", "gray", "black"],
["-", "-", ":", "--", "--"], _[[3,2,1,4,5]])
end
ylim([0.0,1.0])
xlim([0.0,3.0])
xlabel("average degree")
ylabel("fraction of nodes")
legend(["assortative, q=2/3", "assortative, q=1/3", "q=0",
"dissortative, q=1/3", "dissortative, q=2/3"])
# +
## Fig 4.1
Random.seed!(1234)
seeds = rand(UInt64, Rep)
df = DataFrame()
for seed in seeds
for (q, a) in ((0.0, true), (1/3, true), (2/3, true), (1/3, false), (2/3, false))
g = XBS(2^9, 4, q, a, seed)
g = induced_subgraph(g, findall(>(0), degree(g)))[1]
ds, knns = eachcol(deg_corr(g))
append!(df, DataFrame(seed=seed, q=q, a=a, ds=ds, knns=knns))
end
end
@chain df begin
groupby([:q, :a, :ds])
combine(:knns => mean => :knns)
sort(:ds)
groupby([:q, :a])
foreach((c, s, sdf) -> plot(sdf.ds, sdf.knns, color=c, linestyle=s),
["black", "gray", "black", "gray", "black"],
["-", "-", ":", "--", "--"], _[[3,2,1,4,5]])
end
xlabel("degree (\$\\ell\$)")
ylabel("\$k_{nn} (\\ell)\$")
xticks(1:2:17)
xlim([0.0,10.0])
ylim([3, 8.0])
legend(["assortative, q=2/3", "assortative, q=1/3", "q=0",
"dissortative, q=1/3", "dissortative, q=2/3"],
loc="upper center", ncol=2)
| Julia_Notebooks/Chapter_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xarray as xr
import datetime
import os
from shapely.geometry import Point
import gdal
import geopandas
from geopandas.tools import sjoin
import rasterio
import numpy as np
from affine import Affine
from pyproj import Proj, transform
import pandas as pd
import affine
from geopandas import GeoDataFrame
# +
coords = pd.read_csv('jt_dataset.txt' , index_col=['Event ID'])
coords = coords.rename(columns={'Longitude': 'longitude', 'Latitude': 'latitude'})
def RetrieveRasterAppAttr(global_coords, rasterfile, newattr):
# retrieve pixel values for lat, lon
t1 = datetime.datetime.now()
with rasterio.open(rasterfile) as src:
global_coords[newattr] = [x[0] for x in src.sample(zip(global_coords.longitude, global_coords.latitude))]
t2 = datetime.datetime.now()
print('It took ', t2-t1)
# -
geom = coords.apply(lambda x : Point([x['longitude'],x['latitude']]), axis=1)
coords = geopandas.GeoDataFrame(coords, geometry=geom) #geom is a Series
coords.crs = {'init' :'epsg:4326'}
shapefile=r'\\pngssvmh01\hpuzzang\Geo Data\global_urban_extent_polygons_v1.01.shp'
poly = geopandas.GeoDataFrame.from_file(shapefile)
coords = sjoin(coords, poly.loc[:, ['SCHNM', 'geometry']], how='left').drop(['index_right'], axis = 1)
shapefile=r'\\pngssvmh01\hpuzzang\Geo Data\TM_WORLD_BORDERS_SIMPL-0.3.shp'
poly = geopandas.GeoDataFrame.from_file(shapefile)
coords = sjoin(coords, poly, how='left').drop(['geometry', 'index_right'], axis = 1)
coords[['Location Name', 'Country',]] = coords[['Location Name', 'Country']].fillna('')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_population_density_adjusted_to_2015_unwpp_country_totals_rev10_2015_2pt5_min.tif', 'GPW_pop_2015')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a000_014bt_2010_cntm_30_sec.tif', 'age0to14cnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a000_014ft_2010_cntm_30_sec.tif', 'age0to14femalecnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a000_014mt_2010_cntm_30_sec.tif', 'age0to14malecnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a015_049bt_2010_cntm_30_sec.tif', 'age15to49cnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a015_049ft_2010_cntm_30_sec.tif', 'age15to49femalecnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a015_049mt_2010_cntm_30_sec.tif', 'age15to49malecnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a015_064bt_2010_cntm_30_sec.tif', 'age15to64cnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a015_064ft_2010_cntm_30_sec.tif', 'age15to64femalecnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a015_064mt_2010_cntm_30_sec.tif', 'age15to64malecnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a065plusbt_2010_cntm_30_sec.tif', 'age65pluscnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a065plusft_2010_cntm_30_sec.tif', 'age65plusfemalecnt')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_basic_demographic_characteristics_rev10_a065plusmt_2010_cntm_30_sec.tif', 'age65plusmalecnt')
# +
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_land_water_area_rev10_landareakm_30_sec.tif', 'landareakm')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\gpw_v4_land_water_area_rev10_waterareakm_30_sec.tif', 'waterareakm')
RetrieveRasterAppAttr(coords, r'H:\Geo Data\final_trr.tif', 'epopTerro')
# -
# replace the missing values
coords._get_numeric_data()[coords._get_numeric_data() < -100000] = np.NaN
gcoords = coords.fillna(0)
# create smart variables: ratio
# water ratio
gcoords.eval("ratioWater = (waterareakm + 0.000001)/(waterareakm + landareakm + 0.000001)", inplace=True)
# female ratio
gcoords.eval("ratioFemale0to14 = (age0to14femalecnt + 0.000001)/(age0to14cnt + 0.000001)", inplace=True)
gcoords.eval("ratioFemale15to49 = (age15to49femalecnt + 0.000001)/(age15to49cnt + 0.000001)", inplace=True)
gcoords.eval("ratioFemale15to64 = (age15to64femalecnt + 0.000001)/(age15to64cnt + 0.000001)", inplace=True)
gcoords.eval("ratioFemale65plus = (age65plusfemalecnt + 0.000001)/(age65pluscnt + 0.000001)", inplace=True)
# kids ratio
gcoords.eval("ratioKid = (age0to14cnt + 0.000001)/(age0to14cnt + age15to64cnt + age65pluscnt + 0.000001)", inplace=True)
# senior ratio
gcoords.eval("ratioSenior = (age65pluscnt + 0.000001)/(age0to14cnt + age15to64cnt + age65pluscnt + 0.000001)", inplace=True)
gcoords.to_csv(r'jt_dataset_appended.txt', sep=',')
| Terrorism Data/Adding Raster Features to Coordinates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/DenysNunes/data-examples/blob/main/spark/advanced/DynamicPartitionInserts.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="zR-_l_KMqNyi"
# ## Dynamic Partition Inserts
#
# Example using Dynamic Partition Inserts. <br>
# This technique is used to overwrite a single partition instead of all data.
# Read more about [here](https://jaceklaskowski.gitbooks.io/mastering-spark-sql/content/spark-sql-dynamic-partition-inserts.html).
# + [markdown] id="KslJXmDbovN4"
# # Init spark
# + id="4zf-u_ukobIK" colab={"base_uri": "https://localhost:8080/"} outputId="90592921-3860-4ef6-9cca-aab91ca7d933"
# !pip install -q pyspark==3.1.1
# !sudo apt install tree
# !rm -rf /tmp/dynpartition/df1/
from pyspark.sql import SparkSession
spark = SparkSession \
.builder \
.master('local[*]') \
.appName("New Session Example") \
.config("spark.serializer", "org.apache.spark.serializer.KryoSerializer") \
.enableHiveSupport() \
.getOrCreate()
spark.sql("drop table if exists tb_parquet_persons")
# + [markdown] id="q2bajL8VotYq"
# # Saving a first dataFrame table
# + colab={"base_uri": "https://localhost:8080/"} id="6hZL6Mibo36b" outputId="2b6fd35a-610f-439f-fd04-5dd2a8e49888"
from pyspark.sql.types import Row
raw_rows = [
Row(id=1, name='Jonh', partition_id=1),
Row(id=2, name='Maria', partition_id=1),
Row(id=3, name='Ben', partition_id=2)
]
df = spark.createDataFrame(raw_rows)
df.show()
# + id="zwtp2ELhpD06"
df.write.saveAsTable(path='/tmp/dynpartition/df1/', name='tb_parquet_persons', partitionBy='partition_id')
# + colab={"base_uri": "https://localhost:8080/"} id="NnTBH4YWpQ6x" outputId="578639b7-da64-473b-b855-5ef099c43dbc"
# !tree /tmp/dynpartition/df1/
# + [markdown] id="PGpAuF9Zp2hd"
# ## Saving a new df over partition 2
# + id="ttp---VnpiaL"
raw_rows_2 = [
Row(id=4, name='Oliver', partition_id=2),
Row(id=5, name='Agata', partition_id=2)
]
df_2 = spark.createDataFrame(raw_rows_2)
df_2.registerTempTable("tb_parquet_new_persons")
# + colab={"base_uri": "https://localhost:8080/"} id="WyOD5sMTq8pB" outputId="26390e11-bc07-4b72-e11b-0499e52f17bb"
spark.sql("""
INSERT OVERWRITE TABLE tb_parquet_persons
PARTITION(partition_id = 2)
SELECT id, name FROM tb_parquet_new_persons
""")
# + colab={"base_uri": "https://localhost:8080/"} id="xux-qEL_rLC6" outputId="2df42e7b-4788-4476-b2f5-6accf0b46ba6"
# !tree /tmp/dynpartition/df1/
# + [markdown] id="la9HljfwsAZh"
# ## Verifying a new data source
#
# Notice that partition 2 was overwritten
# + colab={"base_uri": "https://localhost:8080/"} id="d54O2x5SrMq7" outputId="25d41aff-a0b9-4d08-c7f3-a7786bbe7015"
spark.sql("""
SELECT * FROM tb_parquet_persons
order by id, partition_id
""").show()
# + id="6yvJUuSXrS5P"
| spark/3 - advanced/dynamic_partitions_insert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# This is a note how to start a [data algebra](https://github.com/WinVector/data_algebra) pipeline with user supplied SQL.
# The idea is: in addition to starting query work from tables and materialzied views, one can
# start a calculation from a SQL query.
#
# Let's show how this works.
#
# We import our libraries and set up some example data which we push into
# our notional example database.
# + pycharm={"name": "#%%\n"}
import pandas
from data_algebra.data_ops import *
import data_algebra.SQLite
import data_algebra.test_util
# + pycharm={"name": "#%%\n"}
d1 = pandas.DataFrame({
'g': ['a', 'a', 'b', 'b'],
'v1': [1, 2, 3, 4],
'v2': [5, 6, 7, 8],
})
# + pycharm={"name": "#%%\n"}
sqlite_handle = data_algebra.SQLite.example_handle()
sqlite_handle.insert_table(d1, table_name='d1')
# + [markdown] pycharm={"name": "#%% md\n"}
# Now suppose we have a "by hand" or "user SQL" query we want to start
# with.
# + pycharm={"name": "#%%\n"}
user_sql = """
SELECT
*,
v1 * v2 AS v3
FROM
d1
"""
# + [markdown] pycharm={"name": "#%% md\n"}
# The idea is: we pretend we have a table with the columns `[v1, v2, v3]` and
# realize that by starting our data algebra pipeline with the query instead
# of a table reference or description.
#
# For example, if we wanted to create a pipeline that incorporates
# the user supplied SQL and creates a new column `v4 := v3 + v1`, we
# would do it as so.
# + pycharm={"name": "#%%\n"}
ops = (
SQLNode(
sql=user_sql,
column_names=['g', 'v1', 'v2', 'v3'],
view_name='derived_results'
)
.extend({'v4': 'v3 + v1'})
)
# + [markdown] pycharm={"name": "#%% md\n"}
# This entire combined pipeline can be translated into SQL.
# + pycharm={"name": "#%%\n"}
print(sqlite_handle.to_sql(ops))
# + [markdown] pycharm={"name": "#%% md\n"}
# And it can, of course, be executed in the database.
# + pycharm={"name": "#%%\n"}
res_sqllite = sqlite_handle.read_query(ops)
res_sqllite
# + [markdown] pycharm={"name": "#%% md\n"}
# The returned table matches our expectations.
# + pycharm={"name": "#%%\n"}
expect = d1.copy()
expect['v3'] = expect['v1'] * expect['v2']
expect['v4'] = expect['v3'] + expect['v1']
assert data_algebra.test_util.equivalent_frames(res_sqllite, expect)
# + [markdown] pycharm={"name": "#%% md\n"}
# We can also run the same query in Pandas if we supply the expected *result* of
# the user supplied SQL (not the input).
# + pycharm={"name": "#%%\n"}
dr = d1.copy()
dr['v3'] = dr['v1'] * dr['v2']
res_pandas = ops.eval({'derived_results': dr})
res_pandas
# + pycharm={"name": "#%%\n"}
assert data_algebra.test_util.equivalent_frames(res_pandas, expect)
# + [markdown] pycharm={"name": "#%% md\n"}
# And there we have it: starting data algebra pipelines from arbitrary SQL instead of
# merely from a table reference.
# + pycharm={"name": "#%%\n"}
sqlite_handle.close()
| Examples/GettingStarted/User_SQL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pretraining_env]
# language: python
# name: conda-env-pretraining_env-py
# ---
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "3"
import torchfly
torchfly.set_random_seed(1)
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import DataLoader, Dataset
from torch.nn.utils.rnn import pad_sequence
import numpy as np
import regex as re
import random
import itertools
import tqdm
import time
import json
try:
from torch.utils.tensorboard import SummaryWriter
except:
from tensorboardX import SummaryWriter
from apex import amp
from allennlp.training.checkpointer import Checkpointer
from transformers import AdamW
from transformers import WarmupLinearSchedule
from torchfly.text.tokenizers import UnifiedBPETokenizer
from torchfly.modules.losses import SequenceFocalLoss, SequenceCrossEntropyLoss
from torchfly.modules.transformers import GPT2SimpleLM, UnifiedGPT2SmallConfig
from text_utils import recoverText, normalize
# -
# set tokenizer
tokenizer = UnifiedBPETokenizer()
tokenizer.sep_token = "None"
# add speicial tokens in the same order as Roberta
# tokenizer.add_tokens(["<s>", "<pad>", "</s>", "<unk>", "<mask>"])
'''
class GPT2SmallConfig:
vocab_size = 50257 + len(tokenizer.added_tokens_encoder)
n_special = len(tokenizer.added_tokens_encoder)
n_positions = 1024
n_ctx = 1024
n_embd = 768
n_layer = 12
n_head = 12
resid_pdrop = 0.1
embd_pdrop = 0.1
attn_pdrop = 0.1
layer_norm_epsilon = 1e-5
initializer_range = 0.02
gradient_checkpointing = False
class GPT2MediumConfig:
vocab_size = len(tokenizer.added_tokens_encoder)
n_special = len(tokenizer.added_tokens_encoder)
n_positions = 1024
n_ctx = 1024
n_embd = 1024
n_layer = 24
n_head = 16
resid_pdrop = 0.1
embd_pdrop = 0.1
attn_pdrop = 0.1
layer_norm_epsilon = 1e-5
initializer_range = 0.02
gradient_checkpointing = True
'''
model_A = GPT2SimpleLM(UnifiedGPT2SmallConfig)
model_B = GPT2SimpleLM(UnifiedGPT2SmallConfig)
model_A.load_state_dict(torch.load("../../Checkpoint/best.th"))
model_B.load_state_dict(torch.load("../../Checkpoint/best.th"))
# +
def align_keep_indices(batch_keep_indices):
prev = batch_keep_indices[1]
new_batch_keep_indices = [prev]
for i in range(1, len(batch_keep_indices)):
curr = batch_keep_indices[i]
new = []
for idx in curr:
new.append(prev.index(idx))
new_batch_keep_indices.append(new)
prev = curr
return new_batch_keep_indices
class MultiWOZDataset(Dataset):
def __init__(self, data, tokenizer):
self.data = data
self.tokenizer = tokenizer
self.bos = tokenizer.encode("<s>")
self.user_bos = tokenizer.encode("A:")
self.system_bos = tokenizer.encode("B:")
self.eos = [628, 198]
def __len__(self):
return len(self.data)
def __getitem__(self, index):
full_dialog = self.data[index]['log']
full_dialog_tokens = []
cur_pos = 0
for turn_dialog in full_dialog:
# cur_pos = 0
# user
user = recoverText(turn_dialog['user_delex'])
# user = recoverText(turn_dialog['user_delex'])
user_tokens = self.user_bos + tokenizer.encode(user) + self.eos
user_pos = torch.arange(cur_pos, cur_pos + len(user_tokens))
cur_pos = user_pos[-1] + 1
# belief span
# belief_tokens = self.bos + \
# tokenizer.encode(";".join(turn_dialog['bspan_inform'][1:])) + \
# self.eos
# belief_pos = torch.arange(cur_pos, cur_pos + len(belief_tokens))
# cur_pos = belief_pos[-1]
# Database
if eval(turn_dialog['pointer'])[-2:] == (1, 0):
booked = "book"
elif eval(turn_dialog['pointer'])[-2:] == (0, 1):
booked = "fail"
else:
booked = "none"
if len(turn_dialog['match']) > 0:
num_match = int(turn_dialog['match']) if int(turn_dialog['match']) < 4 else 4
else:
num_match = 0
database = str(num_match) + ";" + booked + ";" + turn_dialog['turn_domain'].strip("[]") + ";"
database_tokens = tokenizer.encode(database)
database_pos = torch.arange(cur_pos, cur_pos + len(database_tokens))
cur_pos = database_pos[-1] + 1
# System
system = recoverText(process_text(turn_dialog['resp'], turn_dialog['turn_domain'].strip("[]")))
system_tokens = self.system_bos + tokenizer.encode(system) + self.eos
system_pos = torch.arange(cur_pos, cur_pos + len(system_tokens))
cur_pos = system_pos[-1] + 1
user_tokens = torch.LongTensor(user_tokens)
system_tokens = torch.LongTensor(system_tokens)
database_tokens = torch.LongTensor(database_tokens)
full_dialog_tokens.append((user_tokens,
user_pos,
system_tokens,
system_pos,
database_tokens,
database_pos))
# if system_pos[-1] > 1:
# break
return full_dialog_tokens
class Collate_Function:
"""This function handles batch collate.
"""
def __init__(self, tokenizer):
self.tokenizer = tokenizer
self.pad = self.tokenizer.encode("<pad>")[0]
def __call__(self, unpacked_data):
breakpoint()
max_turn_len = max([len(item) for item in unpacked_data])
batch_dialogs = []
batch_keep_indices = []
for turn_num in range(max_turn_len):
keep_indices = []
for batch_idx in range(len(unpacked_data)):
if turn_num < len(unpacked_data[batch_idx]):
keep_indices.append(batch_idx)
user_tokens = pad_sequence([unpacked_data[idx][turn_num][0] for idx in keep_indices],
batch_first=True,
padding_value=self.pad)
user_pos = pad_sequence([unpacked_data[idx][turn_num][1] for idx in keep_indices],
batch_first=True,
padding_value=0)
system_tokens = pad_sequence([unpacked_data[idx][turn_num][2] for idx in keep_indices],
batch_first=True,
padding_value=self.pad)
system_pos = pad_sequence([unpacked_data[idx][turn_num][3] for idx in keep_indices],
batch_first=True,
padding_value=0)
database_tokens = pad_sequence([unpacked_data[idx][turn_num][4] for idx in keep_indices],
batch_first=True,
padding_value=self.pad)
database_pos = pad_sequence([unpacked_data[idx][turn_num][5] for idx in keep_indices],
batch_first=True,
padding_value=0)
user_mask = (user_tokens != self.pad).byte()
system_mask = (system_tokens != self.pad).byte()
database_mask = (database_tokens != self.pad).byte()
batch_dialogs.append((user_tokens, user_pos, user_mask,
system_tokens, system_pos, system_mask,
database_tokens, database_pos, database_mask))
batch_keep_indices.append(keep_indices)
# align keep indices
# batch_keep_indices = align_keep_indices(batch_keep_indices)
return batch_dialogs, batch_keep_indices
# +
def calculate_loss(logits, target, mask):
logits = logits[:, :-1].contiguous()
target = target[:, 1:].contiguous()
mask = mask[:, 1:].contiguous().float()
loss = criterion(logits, target, mask, label_smoothing=0.01, reduce=True)
return loss
def filter_past(past, keep_indices):
past = [item[:, keep_indices] for item in past]
return past
def replace_punc(x):
x = x.replace("<", "").replace(">", "")
return x.replace(".", " .").replace(",", " .").replace("?", " ?").replace("?", " ?")
# -
def process_text(text, domain):
text = text.replace("[value_choice]", "[value_count]")
text = text.replace("[value_people]", "[value_count]")
text = text.replace("[value_starts]", "[value_count]")
text = text.replace("[value_car]", '[taxi_type]')
text = text.replace("[value_leave]", "[value_time]")
text = text.replace("[value_arrive]", "[value_time]")
text = text.replace("[value_price]", "[value_pricerange]")
text = text.replace('[value_postcode]', f'[{domain}_postcode]')
text = text.replace('[value_reference]', f'[{domain}_reference]')
text = text.replace('[value_address]', f'[{domain}_address]')
text = text.replace('[value_phone]', f'[{domain}_phone]')
text = text.replace('[value_name]', f'[{domain}_name]')
text = text.replace('[value_id]', f'[{domain}_id]')
return text
# +
# TODO should be clean_train_data.json
with open("yichi_data/train_data.json") as f:
train_data = json.load(f)
with open("yichi_data/val_data.json") as f:
val_data = json.load(f)
with open("yichi_data/test_data.json") as f:
test_data = json.load(f)
indices = np.arange(len(train_data))
np.random.shuffle(indices)
# use all data
indices = indices
train_data = [train_data[idx] for idx in indices]
# +
train_dataset = MultiWOZDataset(train_data, tokenizer)
val_dataset = MultiWOZDataset(val_data, tokenizer)
test_dataset = MultiWOZDataset(test_data, tokenizer)
train_batch_size = 1
collate_func = Collate_Function(tokenizer)
train_dataloader = DataLoader(dataset=train_dataset,
shuffle=True,
batch_size=train_batch_size,
collate_fn=collate_func)
eval_batch_size = 4
val_dataloader = DataLoader(dataset=val_dataset,
shuffle=False,
batch_size=eval_batch_size,
collate_fn=collate_func)
test_dataloader = DataLoader(dataset=test_dataset,
shuffle=False,
batch_size=eval_batch_size,
collate_fn=collate_func)
# -
criterion = SequenceFocalLoss(gamma=0.0, beta=0.0)
device = torch.device("cuda")
model_A = model_A.to(device)
model_B = model_B.to(device)
# ## Training
if not os.path.isdir("Checkpoint"):
os.makedirs("Checkpoint")
checkpointer = Checkpointer(serialization_dir="Checkpoint",
keep_serialized_model_every_num_seconds=3600*2,
num_serialized_models_to_keep=10)
# +
# optimizer
num_epochs = 20
num_gradients_accumulation = 4
num_train_optimization_steps = num_train_optimization_steps = len(train_dataset) * num_epochs // train_batch_size // num_gradients_accumulation
param_optimizer = list(model_A.named_parameters()) + list(model_B.named_parameters())
no_decay = ['ln', 'bias', 'LayerNorm']
optimizer_grouped_parameters = [
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)], 'weight_decay': 0.01},
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)], 'weight_decay': 0.0}
]
optimizer = AdamW(optimizer_grouped_parameters,
lr=3e-5,
correct_bias=False)
scheduler = WarmupLinearSchedule(optimizer,
warmup_steps=1000,
t_total=num_train_optimization_steps)
# +
# [model_A, model_B], optimizer = amp.initialize([model_A, model_B], optimizer, opt_level="O1")
# +
user_weight = 1.0
def train_one_iter(batch_dialogs, batch_keep_indices, update_count, fp16=False):
aligned_batch_keep_indices = align_keep_indices(batch_keep_indices)
mask = torch.ByteTensor([]).to(device)
prev_batch_size = batch_dialogs[0][0].shape[0]
past = None
all_logits = []
target = []
total_loss = 0
for turn_num in range(len(batch_keep_indices)):
# data send to gpu
dialogs = batch_dialogs[turn_num]
dialogs = [item.to(device) for item in dialogs]
user_tokens, user_pos, user_mask, \
system_tokens, system_pos, system_mask, \
database_tokens, database_pos, database_mask = dialogs
# filtering algorithm
keep_indices = aligned_batch_keep_indices[turn_num]
if len(keep_indices) != prev_batch_size:
past = filter_past(past, keep_indices)
mask = mask[keep_indices, :]
# User Utterance
mask = torch.cat([mask, user_mask], dim=-1)
logits, past = model_A(user_tokens, position_ids=user_pos, mask=mask, past=past)
A_loss = calculate_loss(logits, user_tokens, user_mask)
# Database Tokens
mask = torch.cat([mask, database_mask], dim=-1)
logits, past = model_B(database_tokens, position_ids=database_pos, mask=mask, past=past)
database_loss = calculate_loss(logits, database_tokens, database_mask)
# System Response
mask = torch.cat([mask, system_mask], dim=-1)
logits, past = model_B(system_tokens, position_ids=system_pos, mask=mask, past=past)
B_loss = calculate_loss(logits, system_tokens, system_mask)
# tail
total_loss = total_loss + user_weight * A_loss + B_loss + database_loss
prev_batch_size = user_tokens.shape[0]
# breakpoint
# all_logits = torch.cat(all_logits, dim=1)
# all_logits = all_logits[:, :-1].contiguous()
# target = torch.cat(target, dim=1)
# target = target[:, 1:].contiguous()
# target_mask = torch.ones_like(target).float()
# total_loss = criterion(all_logits, target, target_mask, label_smoothing=0.02, reduce=True)
# gradient accumulation
total_loss /= len(batch_keep_indices)
total_loss /= num_gradients_accumulation
if fp16:
with amp.scale_loss(total_loss, optimizer) as scaled_loss:
scaled_loss.backward()
else:
total_loss.backward()
record_loss = total_loss.item() * num_gradients_accumulation
perplexity = np.exp(record_loss)
return record_loss, perplexity
# -
def calculate_length(batch_dialogs):
total_sum = 0
for turn_num in range(len(batch_keep_indices)):
total_sum += batch_dialogs[turn_num][2].sum(-1) + \
batch_dialogs[turn_num][5].sum(-1) + \
batch_dialogs[turn_num][8].sum(-1)
return total_sum
# +
update_count = 0
progress_bar = tqdm.tqdm_notebook
start = time.time()
for ep in range(num_epochs):
"Training"
pbar = progress_bar(train_dataloader)
model_A.train()
model_B.train()
for batch_dialogs, batch_keep_indices in pbar:
if calculate_length(batch_dialogs).item() > 900:
print("exceed limit")
continue
if len(batch_keep_indices) < 2:
continue
record_loss, perplexity = train_one_iter(batch_dialogs, batch_keep_indices, update_count, fp16=False)
update_count += 1
if update_count % num_gradients_accumulation == num_gradients_accumulation - 1:
# update for gradient accumulation
scheduler.step()
torch.nn.utils.clip_grad_norm_(model_A.parameters(), 5.0)
torch.nn.utils.clip_grad_norm_(model_B.parameters(), 5.0)
optimizer.step()
optimizer.zero_grad()
# speed measure
end = time.time()
speed = train_batch_size * num_gradients_accumulation / (end - start)
start = end
# show progress
pbar.set_postfix(loss=record_loss, perplexity=perplexity, speed=speed)
# "Evaluation"
# print(f"Epoch {ep} Validation")
# eval_res = validate(val_dataloader, val_data)
# print(eval_res)
# print(f"Epoch {ep} Test")
# eval_res = validate(test_dataloader, test_data)
# print(eval_res)
checkpointer.save_checkpoint(ep,
[model_A.state_dict(), model_B.state_dict()],
{"None": None},
True
)
# +
res = []
for batch_dialogs, batch_keep_indices in pbar:
length = calculate_length(batch_dialogs).item()
res.append(length)
# -
res = np.array(res)
def validate(dataloader, data):
model_A.eval()
model_B.eval()
temperature = 0.5
all_response = []
for batch_dialogs, batch_keep_indices in tqdm.tqdm_notebook(dataloader):
aligned_batch_keep_indices = align_keep_indices(batch_keep_indices)
past = None
generated_responses = [[] for i in range(batch_dialogs[0][0].shape[0])]
mask = torch.ByteTensor([]).to(device)
prev_batch_size = batch_dialogs[0][0].shape[0]
with torch.no_grad():
for turn_num in range(len(batch_keep_indices)):
# data send to gpu
dialogs = batch_dialogs[turn_num]
dialogs = [item.to(device) for item in dialogs]
user_tokens, user_pos, user_mask, \
system_tokens, system_pos, system_mask, \
belief_tokens, belief_pos, belief_mask = dialogs
# batch filtering algorithm
keep_indices = aligned_batch_keep_indices[turn_num]
if len(keep_indices) != prev_batch_size:
past = filter_past(past, keep_indices)
mask = mask[keep_indices, :]
# define some initials
cur_batch_size = user_tokens.shape[0]
flags = np.ones(cur_batch_size)
generated_tokens = [[] for i in range(cur_batch_size)]
# feed in user
mask = torch.cat([mask, user_mask], dim=-1)
_, past = model_A(user_tokens, position_ids=user_pos, mask=mask, past=past)
# response generation
response = []
# first three tokens
prev_input = system_tokens[:, :3]
cur_pos = system_pos[:, :3]
temp_past = past
temp_mask = F.pad(mask, pad=(0,3), value=1)
# feed into B
logits, temp_past = model_B(prev_input, position_ids=cur_pos, mask=temp_mask, past=temp_past)
# set current position
cur_pos = cur_pos[:, -1].unsqueeze(1) + 1
for i in range(50):
logits = logits[:, -1, :] / temperature
prev_tokens = torch.argmax(logits, dim=-1)
np_prev_tokens = prev_tokens.cpu().numpy()
# nucleus sampling
# logits = top_filtering(logits, top_k=100, top_p=0.7)
# probs = F.softmax(logits, -1)
# prev_input = torch.multinomial(probs, num_samples=1)
# add to generated tokens list
count = 0
for idx, value in enumerate(flags):
if value != 0:
generated_tokens[idx].append(np_prev_tokens[count])
count += 1
# filtering algorithm
if np.any(np_prev_tokens == 628):
# set flags 0
count = 0
for idx, value in enumerate(flags):
if value == 1:
if np_prev_tokens[count] == 628:
flags[idx] = 0
count += 1
# compute which one to keep
keep_indices = np.argwhere(np_prev_tokens != 628).squeeze(1)
# filter
prev_tokens = prev_tokens[keep_indices.tolist()]
cur_pos = cur_pos[keep_indices.tolist(), :]
temp_mask = temp_mask[keep_indices.tolist(), :]
temp_past = [item[:, keep_indices.tolist()] for item in temp_past]
np_prev_tokens = np_prev_tokens[keep_indices.tolist()]
if np.all(flags == 0):
break
# prepare for the next token
temp_mask = F.pad(temp_mask, pad=(0, 1), value=1)
logits, temp_past = model_B(prev_tokens.view(-1, 1),
position_ids=cur_pos,
mask=temp_mask,
past=temp_past)
cur_pos = cur_pos + 1
# real system_tokens feed in
mask = torch.cat([mask, system_mask], dim=-1)
_, past = model_B(system_tokens, position_ids=system_pos, mask=mask, past=past)
# inject into generated_responses_list
decoded_responses = [tokenizer.decode(item).replace("\n", "") for item in generated_tokens]
count = 0
for idx in batch_keep_indices[turn_num]:
generated_responses[idx].append(decoded_responses[count])
count += 1
# add to the final responses
for item in generated_responses:
all_response.extend(item)
# Stage 2
# prepare for metric eval
dialog_data = []
count = 0
all_results = []
for i in range(len(data)):
raw_dialog = data[i]
for turn_num in range(len(raw_dialog)):
replaced_response = clean_sentence(
replace_punc(raw_dialog[turn_num]["replaced_response"].lower().replace("slot", "SLOT")), entity_dict)
generated_response = clean_sentence(replace_punc(all_response[count].lower().replace("slot", "SLOT")), entity_dict)
dialog_data.append({"dial_id": raw_dialog[turn_num]["dial_id"],
"turn_num": raw_dialog[turn_num]["turn_num"],
"response": replaced_response,
"generated_response":generated_response
})
count += 1
sccuess_f1 = success_f1_metric(dialog_data)
bleu = bleu_metric(dialog_data)
return {"bleu": bleu,
"sccuess_f1": sccuess_f1
}
| experiement/multiwoz/MultiWOZ Multi-Turn Train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to pass class names to ConfusionMatrix
#
# This is a follow up to issue [#244](https://github.com/DistrictDataLabs/yellowbrick/issues/244) and PR [#253](https://github.com/DistrictDataLabs/yellowbrick/pull/253), to document how to pass class names to `ConfusionMatrix`
# +
from sklearn.datasets import load_iris
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split as tts
from yellowbrick.classifier import ConfusionMatrix
# +
iris = load_iris()
X = iris.data
y = iris.target
X_train, X_test, y_train, y_test = tts(X, y, test_size=0.2)
# -
## target names are a list of strings corresponding to the classes
classes = iris.target_names
classes
# +
model = LogisticRegression()
cm = ConfusionMatrix(model, classes=classes)
cm.fit(X_train, y_train)
cm.score(X_test, y_test)
cm.poof()
# -
# :(
#
# Workaround:
cm = ConfusionMatrix(
model, classes=classes,
label_encoder={0: 'setosa', 1: 'versicolor', 2: 'virginica'}
)
cm.fit(X_train, y_train)
cm.score(X_test, y_test)
cm.poof()
| examples/rebeccabilbro/conf_matrix_class_names.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Topomap visualizations with MNE
#
# ### Content
#
# + [1. Notebook description](#1.-Notebook-Description)
# + [2. Load Data](#2.-Load-Data)
# + [3. Plots](#3.-Plots)
#
# ---
#
# # 1. Notebook Description
#
# This notebook is a plotting helper for my thesis. It generates example time series and topographic projections using an MNE montage.
#
# ---
#
# **Imports:**
# %load_ext autoreload
# %autoreload 2
# %pylab inline
# +
# digits related code
from digits.data import matimport
from digits.data import select
# system libraries
import matplotlib.pyplot as plt
import numpy as np
from mne import create_info, EvokedArray
from mne.viz import plot_topomap, plot_layout, plot_montage
from mne.channels import make_eeg_layout
from IPython.display import HTML
# -
# # 2. Load Data
imp = matimport.Importer(dataroot='../../data/thomas/artcorr/')
imp.open('3130.h5')
samples = imp.store.samples
targets = imp.store.targets
# And select a small portion of the samples.
samples = select.fromtimerange(samples, 't_0200', 't_0600')
samples = select.fromchannelblacklist(samples, ['LHEOG', 'RHEOG', 'IOL'])
samples, targets = select.fromtargetlist(samples, targets, [0,1])
# # 3. Plots
info = create_info(select.getchannelnames(samples),
sfreq=1000,
ch_types='eeg',
montage='standard_1005')
layout = make_eeg_layout(info)
arr = EvokedArray(samples.ix[0].reshape(61, -2), info, tmin=0)
_ = arr.plot_topomap(times=[0, 10/1000, 100/1000, 200/1000, 300/1000], size=2, colorbar=False)
rcParams['figure.figsize'] = (2,2)
_ = plot_layout(layout)
# Print some sample time series:
rcParams['figure.figsize'] = (14, 3)
# 10-20 list
es = ['Fp1', 'Fpz', 'Fp2',
'O1', 'Oz', 'O2']
for e in es:
plot(samples.iloc[0][e].values)
plt.legend(es, title='Electrodes', loc='lower right', ncol=3)
plt.xlabel('time (ms)')
_ = plt.ylabel('voltage')
# Use html5 features for an animation:
fig, anim = arr.animate_topomap('eeg', times = np.linspace(0,0.4,50),
frame_rate=1, blit=False, show=False)
anim.repeat = False
_ = HTML(anim.to_html5_video())
# ---
| results/mne_topomap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
Imputing missing values through various strategies
# +
from sklearn import datasets
import numpy as np
iris = datasets.load_iris()
iris_X = iris.data
# -
masking_array = np.random.binomial(1, .25,iris_X.shape).astype(bool)
iris_X[masking_array] = np.nan
masking_array[:5]
iris_X [:5]
from sklearn import preprocessing
impute = preprocessing.Imputer()
iris_X_prime = impute.fit_transform(iris_X)
iris_X_prime[:5]
iris_X_prime[0, 0]
iris_X[0, 0]
impute = preprocessing.Imputer(strategy='median')
iris_X_prime = impute.fit_transform(iris_X)
iris_X_prime[:5]
iris_X[np.isnan(iris_X)] = -1
iris_X[:5]
impute = preprocessing.Imputer(missing_values=-1)
iris_X_prime = impute.fit_transform(iris_X)
iris_X_prime[:5]
import pandas as pd
iris_X_prime = np.where(pd.DataFrame(iris_X).isnull(),-1,iris_X)
iris_X_prime[:5]
pd.DataFrame(iris_X).fillna(-1)[:5].values
| Chapter02/Imputing missing values through various strategies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Science Pipeline
# ## EDA
# %matplotlib inline
import pandas as pd
iris_filename = 'datasets-uci-iris.csv'
iris = pd.read_csv(iris_filename, header=None,
names= ['sepal_length', 'sepal_width',
'petal_length', 'petal_width', 'target'])
iris.head()
iris.describe()
boxes = iris.boxplot(return_type='axes')
iris.quantile([0.1, 0.9])
iris.target.unique()
pd.crosstab(iris['petal_length'] > 3.758667, iris['petal_width'] > 1.198667)
scatterplot = iris.plot(kind='scatter', x='petal_width', y='petal_length',
s=64, c='blue', edgecolors='white')
distr = iris.petal_width.plot(kind='hist', alpha=0.5, bins=20)
# ## Feature Creation
import numpy as np
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
cali = datasets.california_housing.fetch_california_housing()
X = cali['data']
Y = cali['target']
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2)
from sklearn.neighbors import KNeighborsRegressor
regressor = KNeighborsRegressor()
regressor.fit(X_train, Y_train)
Y_est = regressor.predict(X_test)
print ("MAE=", mean_squared_error(Y_test, Y_est))
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
regressor = KNeighborsRegressor()
regressor.fit(X_train_scaled, Y_train)
Y_est = regressor.predict(X_test_scaled)
print ("MAE=", mean_squared_error(Y_test, Y_est))
from sklearn.preprocessing import RobustScaler
scaler2 = RobustScaler()
X_train_scaled = scaler2.fit_transform(X_train)
X_test_scaled = scaler2.transform(X_test)
regressor = KNeighborsRegressor()
regressor.fit(X_train_scaled, Y_train)
Y_est = regressor.predict(X_test_scaled)
print ("MAE=", mean_squared_error(Y_test, Y_est))
non_linear_feat = 5 # AveOccup
X_train_new_feat = np.sqrt(X_train[:,non_linear_feat])
X_train_new_feat.shape = (X_train_new_feat.shape[0], 1)
X_train_extended = np.hstack([X_train, X_train_new_feat])
X_test_new_feat = np.sqrt(X_test[:,non_linear_feat])
X_test_new_feat.shape = (X_test_new_feat.shape[0], 1)
X_test_extended = np.hstack([X_test, X_test_new_feat])
scaler = StandardScaler()
X_train_extended_scaled = scaler.fit_transform(X_train_extended)
X_test_extended_scaled = scaler.transform(X_test_extended)
regressor = KNeighborsRegressor()
regressor.fit(X_train_extended_scaled, Y_train)
Y_est = regressor.predict(X_test_extended_scaled)
print ("MAE=", mean_squared_error(Y_test, Y_est))
# ## Dimensionality Reduction
# ### Covariance matrix
from sklearn import datasets
import numpy as np
iris = datasets.load_iris()
cov_data = np.corrcoef(iris.data.T)
print (iris.feature_names)
print (cov_data)
import matplotlib.pyplot as plt
img = plt.matshow(cov_data, cmap=plt.cm.rainbow)
plt.colorbar(img, ticks=[-1, 0, 1], fraction=0.045)
for x in range(cov_data.shape[0]):
for y in range(cov_data.shape[1]):
plt.text(x, y, "%0.2f" % cov_data[x,y],
size=12, color='black', ha="center", va="center")
plt.show()
from sklearn.decomposition import PCA
pca_2c = PCA(n_components=2)
X_pca_2c = pca_2c.fit_transform(iris.data)
X_pca_2c.shape
plt.scatter(X_pca_2c[:,0], X_pca_2c[:,1], c=iris.target, alpha=0.8,
s=60, marker='o', edgecolors='white')
plt.show()
pca_2c.explained_variance_ratio_.sum()
pca_2c.components_
pca_2cw = PCA(n_components=2, whiten=True)
X_pca_1cw = pca_2cw.fit_transform(iris.data)
plt.scatter(X_pca_1cw[:,0], X_pca_1cw[:,1], c=iris.target, alpha=0.8, s=60, marker='o', edgecolors='white')
plt.show()
pca_2cw.explained_variance_ratio_.sum()
pca_1c = PCA(n_components=1)
X_pca_1c = pca_1c.fit_transform(iris.data)
plt.scatter(X_pca_1c[:,0], np.zeros(X_pca_1c.shape), c=iris.target, alpha=0.8, s=60, marker='o', edgecolors='white')
plt.show()
pca_1c.explained_variance_ratio_.sum()
pca_95pc = PCA(n_components=0.95)
X_pca_95pc = pca_95pc.fit_transform(iris.data)
print (pca_95pc.explained_variance_ratio_.sum())
print (X_pca_95pc.shape)
# ### A variation of PCA for big data: randomized PCA
rpca_2c = PCA(svd_solver='randomized', n_components=2)
X_rpca_2c = rpca_2c.fit_transform(iris.data)
plt.scatter(X_rpca_2c[:,0], X_rpca_2c[:,1], c=iris.target, alpha=0.8, s=60, marker='o', edgecolors='white')
plt.show()
rpca_2c.explained_variance_ratio_.sum()
# ### Latent Factor Analysis
from sklearn.decomposition import FactorAnalysis
fact_2c = FactorAnalysis(n_components=2)
X_factor = fact_2c.fit_transform(iris.data)
plt.scatter(X_factor[:,0], X_factor[:,1], c=iris.target, alpha=0.8, s=60, marker='o', edgecolors='white')
plt.show()
# ### Linear Discriminant Analysis, LDA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
lda_2c = LDA(n_components=2)
X_lda_2c = lda_2c.fit_transform(iris.data, iris.target)
plt.scatter(X_lda_2c[:,0], X_lda_2c[:,1], c=iris.target, alpha=0.8, s=60, marker='o', edgecolors='white')
plt.show()
# ### Latent Semantical Analysis, LSA
from sklearn.datasets import fetch_20newsgroups
categories = ['sci.med', 'sci.space']
twenty_sci_news = fetch_20newsgroups(categories=categories)
from sklearn.feature_extraction.text import TfidfVectorizer
tf_vect = TfidfVectorizer()
word_freq = tf_vect.fit_transform(twenty_sci_news.data)
from sklearn.decomposition import TruncatedSVD
tsvd_2c = TruncatedSVD(n_components=50)
tsvd_2c.fit(word_freq)
np.array(tf_vect.get_feature_names())[tsvd_2c.components_[20].argsort()[-10:][::-1]]
# ### Kernel PCA
def circular_points (radius, N):
return np.array([[np.cos(2*np.pi*t/N)*radius, np.sin(2*np.pi*t/N)*radius] for t in range(N)])
N_points = 50
fake_circular_data = np.vstack([circular_points(1.0, N_points), circular_points(5.0, N_points)])
fake_circular_data += np.random.rand(*fake_circular_data.shape)
fake_circular_target = np.array([0]*N_points + [1]*N_points)
plt.scatter(fake_circular_data[:,0], fake_circular_data[:,1], c=fake_circular_target, alpha=0.8,
s=60, marker='o', edgecolors='white')
plt.show()
from sklearn.decomposition import KernelPCA
kpca_2c = KernelPCA(n_components=2, kernel='rbf')
X_kpca_2c = kpca_2c.fit_transform(fake_circular_data)
plt.scatter(X_kpca_2c[:,0], X_kpca_2c[:,1], c=fake_circular_target, alpha=0.8, s=60, marker='o', edgecolors='white')
plt.show()
# ### T-NSE
# +
from sklearn.manifold import TSNE
from sklearn.datasets import load_iris
iris = load_iris()
X, y = iris.data, iris.target
X_tsne = TSNE(n_components=2).fit_transform(X)
plt.scatter(X_tsne[:, 0], X_tsne[:, 1], c=y, alpha=0.8, s=60, marker='o', edgecolors='white')
plt.show()
# -
# ### Restricted Boltzmann Machine, RBM
from sklearn import preprocessing
from sklearn.neural_network import BernoulliRBM
n_components = 64 # Try with 64, 100, 144
olivetti_faces = datasets.fetch_olivetti_faces()
X = preprocessing.binarize(preprocessing.scale(olivetti_faces.data.astype(float)), 0.5)
rbm = BernoulliRBM(n_components=n_components, learning_rate=0.01, n_iter=100)
rbm.fit(X)
plt.figure(figsize=(4.2, 4))
for i, comp in enumerate(rbm.components_):
plt.subplot(int(np.sqrt(n_components+1)), int(np.sqrt(n_components+1)), i + 1)
plt.imshow(comp.reshape((64, 64)), cmap=plt.cm.gray_r, interpolation='nearest')
plt.xticks(()); plt.yticks(())
plt.suptitle(str(n_components) + ' components extracted by RBM', fontsize=16)
plt.subplots_adjust(0.08, 0.02, 0.92, 0.85, 0.08, 0.23)
plt.show()
# ## Detection and Treatment of Outliers
# ### Univariate outlier detection
from sklearn.datasets import load_boston
boston = load_boston()
continuous_variables = [n for n in range(boston.data.shape[1]) if n!=3]
import numpy as np
from sklearn import preprocessing
normalized_data = preprocessing.StandardScaler().fit_transform(boston.data[:,continuous_variables])
outliers_rows, outliers_columns = np.where(np.abs(normalized_data)>3)
print (len(outliers_rows))
print (outliers_rows)
print (list(zip(outliers_rows, outliers_columns)))
# ### EllipticEnvelope
# Create an artificial distribution made of blobs
from sklearn.datasets import make_blobs
blobs = 1
blob = make_blobs(n_samples=100, n_features=2, centers=blobs, cluster_std=1.5, shuffle=True, random_state=5)
# Robust Covariance Estimate
from sklearn.covariance import EllipticEnvelope
robust_covariance_est = EllipticEnvelope(contamination=.1).fit(blob[0])
detection = robust_covariance_est.predict(blob[0])
outliers = np.where(detection==-1)[0]
inliers = np.where(detection==1)[0]
# Draw the distribution and the detected outliers
from matplotlib import pyplot as plt
# Just the distribution
plt.scatter(blob[0][:,0],blob[0][:,1], c='blue', alpha=0.8, s=60, marker='o', edgecolors='white')
plt.show()
# The distribution and the outliers
in_points = plt.scatter(blob[0][inliers,0],blob[0][inliers,1], c='blue', alpha=0.8, s=60, marker='o', edgecolors='white')
out_points = plt.scatter(blob[0][outliers,0],blob[0][outliers,1], c='red', alpha=0.8, s=60, marker='o', edgecolors='white')
plt.legend((in_points,out_points),('inliers','outliers'), scatterpoints=1, loc='lower right')
plt.show()
from sklearn.decomposition import PCA
# Normalized data relative to continuos variables
continuous_variables = [n for n in range(boston.data.shape[1]) if n!=3]
normalized_data = preprocessing.StandardScaler().fit_transform(boston.data[:,continuous_variables])
# Just for visualization purposes pick the first 2 PCA components
pca = PCA(n_components=2)
Zscore_components = pca.fit_transform(normalized_data)
vtot = 'PCA Variance explained ' + str(round(np.sum(pca.explained_variance_ratio_),3))
v1 = str(round(pca.explained_variance_ratio_[0],3))
v2 = str(round(pca.explained_variance_ratio_[1],3))
# Robust Covariance Estimate
robust_covariance_est = EllipticEnvelope(store_precision=False, assume_centered = False, contamination=.05)
robust_covariance_est.fit(normalized_data)
detection = robust_covariance_est.predict(normalized_data)
outliers = np.where(detection==-1)
regular = np.where(detection==1)
# Draw the distribution and the detected outliers
from matplotlib import pyplot as plt
in_points = plt.scatter(Zscore_components[regular,0],Zscore_components[regular,1], c='blue', alpha=0.8, s=60, marker='o', edgecolors='white')
out_points = plt.scatter(Zscore_components[outliers,0],Zscore_components[outliers,1], c='red', alpha=0.8, s=60, marker='o', edgecolors='white')
plt.legend((in_points,out_points),('inliers','outliers'), scatterpoints=1, loc='best')
plt.xlabel('1st component ('+v1+')')
plt.ylabel('2nd component ('+v2+')')
plt.xlim([-7,7])
plt.ylim([-6,6])
plt.title(vtot)
plt.show()
# ### OneClassSVM
from sklearn.decomposition import PCA
from sklearn import preprocessing
from sklearn import svm
# Normalized data relative to continuos variables
continuous_variables = [n for n in range(boston.data.shape[1]) if n!=3]
normalized_data = preprocessing.StandardScaler().fit_transform(boston.data[:,continuous_variables])
# Just for visualization purposes pick the first 5 PCA components
pca = PCA(n_components=5)
Zscore_components = pca.fit_transform(normalized_data)
vtot = 'PCA Variance explained ' + str(round(np.sum(pca.explained_variance_ratio_),3))
# OneClassSVM fitting and estimates
outliers_fraction = 0.02 #
nu_estimate = 0.95 * outliers_fraction + 0.05
machine_learning = svm.OneClassSVM(kernel="rbf", gamma=1.0/len(normalized_data), degree=3, nu=nu_estimate)
machine_learning.fit(normalized_data)
detection = machine_learning.predict(normalized_data)
outliers = np.where(detection==-1)
regular = np.where(detection==1)
# Draw the distribution and the detected outliers
from matplotlib import pyplot as plt
for r in range(1,5):
in_points = plt.scatter(Zscore_components[regular,0],Zscore_components[regular,r], c='blue', alpha=0.8, s=60, marker='o', edgecolors='white')
out_points = plt.scatter(Zscore_components[outliers,0],Zscore_components[outliers,r], c='red', alpha=0.8, s=60, marker='o', edgecolors='white')
plt.legend((in_points,out_points),('inliers','outliers'), scatterpoints=1, loc='best')
plt.xlabel('Component 1 ('+str(round(pca.explained_variance_ratio_[0],3))+')')
plt.ylabel('Component '+str(r+1)+'('+str(round(pca.explained_variance_ratio_[r],3))+')')
plt.xlim([-7,7])
plt.ylim([-6,6])
plt.title(vtot)
plt.show()
# ## Scoring functions
# ### Multilabel classification
from sklearn import datasets
iris = datasets.load_iris()
# No crossvalidation for this dummy notebook
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(iris.data, iris.target, test_size=0.50, random_state=4)
# Use a very bad multiclass classifier
from sklearn.tree import DecisionTreeClassifier
classifier = DecisionTreeClassifier(max_depth=2)
classifier.fit(X_train, Y_train)
Y_pred = classifier.predict(X_test)
iris.target_names
from sklearn import metrics
cm = metrics.confusion_matrix(Y_test, Y_pred)
print (cm)
import matplotlib.pyplot as plt
img = plt.matshow(cm, cmap=plt.cm.autumn)
plt.colorbar(img, fraction=0.045)
for x in range(cm.shape[0]):
for y in range(cm.shape[1]):
plt.text(x, y, "%0.2f" % cm[x,y],
size=12, color='black', ha="center", va="center")
plt.show()
print ("Accuracy:", metrics.accuracy_score(Y_test, Y_pred))
print ("Precision:", metrics.precision_score(Y_test, Y_pred, average='weighted'))
print ("Recall:", metrics.recall_score(Y_test, Y_pred, average='weighted'))
print ("F1 score:", metrics.f1_score(Y_test, Y_pred, average='weighted'))
from sklearn.metrics import classification_report
print (classification_report(Y_test, Y_pred, target_names=iris.target_names))
# ### Regression
from sklearn.metrics import mean_absolute_error
mean_absolute_error([1.0, 0.0, 0.0], [0.0, 0.0, -1.0])
from sklearn.metrics import mean_squared_error
mean_squared_error([-10.0, 0.0, 0.0], [0.0, 0.0, 0.0])
# ## Testing and Validating
from sklearn.datasets import load_digits
digits = load_digits()
print (digits.DESCR)
X = digits.data
y = digits.target
X[0]
from sklearn import svm
h1 = svm.LinearSVC(C=1.0) # linear SVC
h2 = svm.SVC(kernel='rbf', degree=3, gamma=0.001, C=1.0) # Radial basis SVC
h3 = svm.SVC(kernel='poly', degree=3, C=1.0) # 3rd degree polynomial SVC
h1.fit(X,y)
print (h1.score(X,y))
from sklearn import model_selection
chosen_random_state = 1
X_train, X_test, y_train, y_test = model_selection.train_test_split(X, y, test_size=0.30, random_state=chosen_random_state)
print ("(X train shape %s, X test shape %s, \ny train shape %s, y test shape %s" \
% (X_train.shape, X_test.shape, y_train.shape, y_test.shape))
h1.fit(X_train,y_train)
print (h1.score(X_test,y_test)) # Returns the mean accuracy on the given test data and labels
chosen_random_state = 1
X_train, X_validation_test, y_train, y_validation_test = model_selection.train_test_split(X, y, test_size=.40,
random_state=chosen_random_state)
X_validation, X_test, y_validation, y_test = model_selection.train_test_split(X_validation_test, y_validation_test,
test_size=.50, random_state=chosen_random_state)
print ("X train shape, %s, X validation shape %s, X test shape %s, \ny train shape %s, y validation shape %s, y test shape %s\n" % \
(X_train.shape, X_validation.shape, X_test.shape, y_train.shape, y_validation.shape, y_test.shape))
for hypothesis in [h1, h2, h3]:
hypothesis.fit(X_train,y_train)
print ("%s -> validation mean accuracy = %0.3f" % (hypothesis, hypothesis.score(X_validation,y_validation)) )
h2.fit(X_train,y_train)
print ("\n%s -> test mean accuracy = %0.3f" % (h2, h2.score(X_test,y_test)))
# ### Cross Validation
choosen_random_state = 1
cv_folds = 10 # Try 3, 5 or 20
eval_scoring='accuracy' # Try also f1
workers = 1
X_train, X_test, y_train, y_test = model_selection.train_test_split(X, y, test_size=0.30, random_state=choosen_random_state)
for hypothesis in [h1, h2, h3]:
scores = model_selection.cross_val_score(hypothesis, X_train, y_train, cv=cv_folds, scoring= eval_scoring, n_jobs=workers)
print ("%s -> cross validation accuracy: mean = %0.3f std = %0.3f" % (hypothesis, np.mean(scores), np.std(scores)))
scores
# ### Using cross-validation iterators
kfolding = model_selection.KFold(n_splits=10, shuffle=True, random_state=1)
for train_idx, validation_idx in kfolding.split(range(100)):
print (train_idx, validation_idx)
h1.fit(X[train_idx],y[train_idx])
h1.score(X[validation_idx],y[validation_idx])
# ### Sampling and Bootstrapping
subsampling = model_selection.ShuffleSplit(n_splits=10, test_size=0.1, random_state=1)
for train_idx, validation_idx in subsampling.split(range(100)):
print (train_idx, validation_idx)
# +
import random
def Bootstrap(n, n_iter=3, random_state=None):
"""
Random sampling with replacement cross-validation generator.
For each iter a sample bootstrap of the indexes [0, n) is
generated and the function returns the obtained sample
and a list of all theexcluded indexes.
"""
if random_state:
random.seed(random_state)
for j in range(n_iter):
bs = [random.randint(0, n-1) for i in range(n)]
out_bs = list({i for i in range(n)} - set(bs))
yield bs, out_bs
boot = Bootstrap(n=100, n_iter=10, random_state=1)
for train_idx, validation_idx in boot:
print (train_idx, validation_idx)
# -
# ## Hyper-parameters Optimization
from sklearn.datasets import load_digits
digits = load_digits()
X, y = digits.data, digits.target
from sklearn import svm
h = svm.SVC()
hp = svm.SVC(probability=True, random_state=1)
from sklearn import model_selection
search_grid = [
{'C': [1, 10, 100, 1000], 'kernel': ['linear']},
{'C': [1, 10, 100, 1000], 'gamma': [0.001, 0.0001], 'kernel': ['rbf']},
]
scorer = 'accuracy'
search_func = model_selection.GridSearchCV(estimator=h, param_grid=search_grid, scoring=scorer, n_jobs=-1, iid=False, refit=True, cv=10)
# %timeit search_func.fit(X,y)
print (search_func.best_estimator_)
print (search_func.best_params_)
print (search_func.best_score_)
# ### Building custom scoring functions
from sklearn.metrics import log_loss, make_scorer
Log_Loss = make_scorer(log_loss, greater_is_better=False, needs_proba=True)
search_func = model_selection.GridSearchCV(estimator=hp, param_grid=search_grid, scoring=Log_Loss, n_jobs=-1, iid=False, refit=True, cv=3)
search_func.fit(X,y)
print (search_func.best_score_)
print (search_func.best_params_)
import numpy as np
from sklearn.preprocessing import LabelBinarizer
def my_custom_log_loss_func(ground_truth, p_predictions, penalty = list(), eps=1e-15):
adj_p = np.clip(p_predictions, eps, 1 - eps)
lb = LabelBinarizer()
g = lb.fit_transform(ground_truth)
if g.shape[1] == 1:
g = np.append(1 - g, g, axis=1)
if penalty:
g[:,penalty] = g[:,penalty] * 2
summation = np.sum(g * np.log(adj_p))
return summation * (-1.0/len(ground_truth))
my_custom_scorer = make_scorer(my_custom_log_loss_func, greater_is_better=False, needs_proba=True, penalty = [4,9])
from sklearn import model_selection
search_grid = [{'C': [1, 10, 100, 1000], 'kernel': ['linear']},
{'C': [1, 10, 100, 1000], 'gamma': [0.001, 0.0001], 'kernel': ['rbf']}]
search_func = model_selection.GridSearchCV(estimator=hp, param_grid=search_grid, scoring=my_custom_scorer, n_jobs=1, iid=False, cv=3)
search_func.fit(X,y)
print (search_func.best_score_)
print (search_func.best_params_)
# ### Reducing grid search run time
search_dict = {'kernel': ['linear','rbf'],'C': [1, 10, 100, 1000], 'gamma': [0.001, 0.0001]}
scorer = 'accuracy'
search_func = model_selection.RandomizedSearchCV(estimator=h,
param_distributions=search_dict,
n_iter=7,
scoring=scorer,
n_jobs=-1,
iid=False,
refit=True,
cv=10,
return_train_score=False)
# %timeit search_func.fit(X,y)
print (search_func.best_estimator_)
print (search_func.best_params_)
print (search_func.best_score_)
res = search_func.cv_results_
for el in zip(res['mean_test_score'], res['std_test_score'], res['params']):
print(el)
# ## Feature Selection
# ### Selection based on variance
# +
from sklearn.datasets import make_classification
X, y = make_classification(n_samples=10, n_features=5, n_informative=3, n_redundant=0, random_state=101)
# +
print ("Variance:", np.var(X, axis=0))
from sklearn.feature_selection import VarianceThreshold
X_selected = VarianceThreshold(threshold=1.0).fit_transform(X)
print ("Before:", X[0, :])
print ("After: ", X_selected[0, :])
# -
# ### Univariate selection
X, y = make_classification(n_samples=800, n_features=100, n_informative=25, n_redundant=0, random_state=101)
from sklearn.feature_selection import SelectPercentile
from sklearn.feature_selection import chi2, f_classif
from sklearn.preprocessing import Binarizer, scale
Xbin = Binarizer().fit_transform(scale(X))
# if you use chi2, input X must be non-negative: X must contain booleans or frequencies
# hence the choice to binarize after the normalization if the variable if above the average
Selector_chi2 = SelectPercentile(chi2, percentile=25).fit(Xbin, y)
Selector_f_classif = SelectPercentile(f_classif, percentile=25).fit(X, y)
chi_scores = Selector_chi2.get_support()
f_classif_scores = Selector_f_classif.get_support()
selected = chi_scores & f_classif_scores # use the bitwise and operator
# ### Recursive elimination
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_classification
X, y = make_classification(n_samples=100, n_features=100, n_informative=5, n_redundant=2, random_state=101)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=101)
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(random_state=101)
classifier.fit(X_train, y_train)
print ('In-sample accuracy: %0.3f' % classifier.score(X_train, y_train))
print ('Out-of-sample accuracy: %0.3f' % classifier.score(X_test, y_test))
from sklearn.feature_selection import RFECV
selector = RFECV(estimator=classifier, step=1, cv=10, scoring='accuracy')
selector.fit(X_train, y_train)
print('Optimal number of features : %d' % selector.n_features_)
X_train_s = selector.transform(X_train)
X_test_s = selector.transform(X_test)
classifier.fit(X_train_s, y_train)
print ('Out-of-sample accuracy: %0.3f' % classifier.score(X_test_s, y_test))
# ### Stability and L1 based selection
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(C=0.1, penalty='l1', random_state=101) # the smaller C the fewer features selected
classifier.fit(X_train, y_train)
print ('Out-of-sample accuracy: %0.3f' % classifier.score(X_test, y_test))
from sklearn.linear_model import RandomizedLogisticRegression
selector = RandomizedLogisticRegression(n_resampling=300, random_state=101)
selector.fit(X_train, y_train)
print ('Variables selected: %i' % sum(selector.get_support()!=0))
X_train_s = selector.transform(X_train)
X_test_s = selector.transform(X_test)
classifier.fit(X_train_s, y_train)
print ('Out-of-sample accuracy: %0.3f' % classifier.score(X_test_s, y_test))
# +
from sklearn.linear_model import RandomizedLasso
from sklearn.datasets import make_regression
X, y = make_regression(n_samples=100, n_features=10, n_informative=4, random_state=101)
rlasso = RandomizedLasso()
rlasso.fit(X, y)
list(enumerate(rlasso.scores_))
# -
# ## Wrapping data into pipelines
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline
from sklearn.pipeline import FeatureUnion
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_classification
X, y = make_classification(n_samples=100, n_features=100, n_informative=5, n_redundant=2, random_state=101)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=101)
classifier = LogisticRegression(C=0.1, penalty='l1', random_state=101)
# +
from sklearn.decomposition import PCA
from sklearn.decomposition import KernelPCA
from sklearn.preprocessing import FunctionTransformer
def identity(x):
return x
def inverse(x):
return 1.0 / x
parallel = FeatureUnion(transformer_list=[
('pca', PCA()),
('kernelpca', KernelPCA()),
('inverse', FunctionTransformer(inverse)),
('original',FunctionTransformer(identity))], n_jobs=1)
# +
from sklearn.preprocessing import RobustScaler
from sklearn.linear_model import RandomizedLogisticRegression
from sklearn.feature_selection import RFECV
selector = RandomizedLogisticRegression(n_resampling=300, random_state=101, n_jobs=1)
pipeline = Pipeline(steps=[('parallel_transformations', parallel),
('random_selection', selector),
('logistic_reg', classifier)])
# -
from sklearn import model_selection
search_dict = {'logistic_reg__C':[10,1,0.1], 'logistic_reg__penalty':['l1','l2']}
search_func = model_selection.GridSearchCV(estimator=pipeline, param_grid =search_dict,
scoring='accuracy', n_jobs=1, iid=False, refit=True, cv=10)
search_func.fit(X_train,y_train)
print (search_func.best_estimator_)
print (search_func.best_params_)
print (search_func.best_score_)
from sklearn.metrics import classification_report
print (classification_report(y_test, search_func.predict(X_test)))
# ## Building custom transformers
# +
from sklearn.base import BaseEstimator, TransformerMixin
class filtering(BaseEstimator, TransformerMixin):
def __init__(self, columns):
self.columns = columns
def fit(self, X, y=None):
return self
def transform(self, X):
if len(self.columns) == 0:
return X
else:
return X[:,self.columns]
# -
ff = filtering([1,2,3])
ff.fit_transform(X_train)
| Chapter3/Data Pipeline-V3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Get started to MONAI bundle
#
# A MONAI bundle usually includes the stored weights of a model, TorchScript model, JSON files which include configs and metadata about the model, information for constructing training, inference, and post-processing transform sequences, plain-text description, legal information, and other data the model creator wishes to include.
#
# For more information about MONAI bundle read the description: https://docs.monai.io/en/latest/bundle_intro.html.
#
# This notebook is a step-by-step tutorial to help get started to develop a bundle package, which contains a config file to construct the training pipeline and also has a `metadata.json` file to define the metadata information.
#
# This notebook mainly contains the below sections:
# - Define a training config with `JSON` or `YAML` format
# - Execute training based on bundle scripts and configs
# - Hybrid programming with config and python code
#
# You can find the usage examples of MONAI bundle key features and syntax in this tutorial, like:
# - Instantiate a python object from a dictionary config with `_target_` indicating class or function name or module path.
# - Execute python expression from a string config with the `$` syntax.
# - Refer to other python object with the `@` syntax.
# - Macro text replacement with the `%` syntax to simplify the config content.
# - Leverage the `_disabled_` syntax to tune or debug different components.
# - Override config content at runtime.
# - Hybrid programming with config and python code.
#
# [](https://colab.research.google.com/github/Project-MONAI/tutorials/blob/master/modules/bundle/get_started.ipynb)
# ## Setup environment
# !python -c "import monai" || pip install -q "monai-weekly[nibabel]"
# ## Setup imports
# + tags=[]
# Copyright (c) MONAI Consortium
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import os
from monai.apps import download_and_extract
from monai.config import print_config
from monai.bundle import ConfigParser
print_config()
# -
# ## Setup data directory
#
# Here specify a directory with the `MONAI_DATA_DIRECTORY` environment variable to save downloaded dataset and outputs.
# + tags=[]
root_dir = os.environ.get("MONAI_DATA_DIRECTORY")
print(f"root dir is: {root_dir}")
# -
# ## Download dataset
#
# Downloads and extracts the dataset.
# The dataset comes from http://medicaldecathlon.com/.
# + tags=[]
resource = "https://msd-for-monai.s3-us-west-2.amazonaws.com/Task09_Spleen.tar"
md5 = "410d4a301da4e5b2f6f86ec3ddba524e"
compressed_file = os.path.join(root_dir, "Task09_Spleen.tar")
data_dir = os.path.join(root_dir, "Task09_Spleen")
if not os.path.exists(data_dir):
download_and_extract(resource, compressed_file, root_dir, md5)
# -
# ## Define train config - Set imports and input / output environments
#
# Now let's start to define the config file for a regular training task. MONAI bundle support both `JSON` and `YAML` format, here we use `JSON` as the example.
#
# According to the predefined syntax of MONAI bundle, `$` indicates an expression to evaluate, `@` refers to another object in the config content. For more details about the syntax in bundle config, please check: https://docs.monai.io/en/latest/config_syntax.html.
#
# Please note that a MONAI bundle doesn't require any hard-coded logic in the config, so users can define the config content in any structure.
#
# For the first step, import `os` and `glob` to use in the expressions (start with `$`), then define input / output environments and enable `cudnn.benchmark` for better performance.
# ```json
# {
# "imports": [
# "$import glob",
# "$import os",
# "$import ignite"
# ],
# "device": "$torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')",
# "ckpt_path": "/workspace/data/models/model.pt",
# "dataset_dir": "/workspace/data/Task09_Spleen",
# "images": "$list(sorted(glob.glob(@dataset_dir + '/imagesTr/*.nii.gz')))",
# "labels": "$list(sorted(glob.glob(@dataset_dir + '/labelsTr/*.nii.gz')))"
# }
# ```
# ## Define train config - Define network, optimizer, loss function
#
# Define `UNet` of MONAI as the training network, and use the `Adam` optimizer of PyTorch, `DiceCELoss` of MONAI.
#
# An instantiable config component uses `_target_` keyword to define the class / function name or module path, other keys are args for the component.
# ```json
# "network_def": {
# "_target_": "UNet",
# "spatial_dims": 3,
# "in_channels": 1,
# "out_channels": 2,
# "channels": [16, 32, 64, 128, 256],
# "strides": [2, 2, 2, 2],
# "num_res_units": 2,
# "norm": "batch"
# }
# ```
# Move the network to the expected `device`
# ```json
# "network": <EMAIL>)"
# ```
# Define optimizer and loss function, for MONAI classes, we can use the class name directly, other classes should provide the module path (like `Adam`).
# ```json
# "loss": {
# "_target_": "DiceCELoss",
# "to_onehot_y": true,
# "softmax": true,
# "squared_pred": true,
# "batch": true
# },
# "optimizer": {
# "_target_": "torch.optim.Adam",
# "params": "$@network.<EMAIL>()",
# "lr": 1e-4
# }
# ```
# ## Define train config - Define data loading and preprocessing logic
#
# Define `transforms` and `dataset`, `dataloader` to generate training data for network.
#
# To make the config stucture clear, here we split the `train` and `validate` related components into 2 sections:
# ```json
# "train": {...},
# "validate": {...}
# ```
# The composed transforms are for preprocessing.
# ```json
# "train": {
# "preprocessing": {
# "_target_": "Compose",
# "transforms": [
# {
# "_target_": "LoadImaged",
# "keys": ["image", "label"]
# },
# {
# "_target_": "EnsureChannelFirstd",
# "keys": ["image", "label"]
# },
# {
# "_target_": "Orientationd",
# "keys": ["image", "label"],
# "axcodes": "RAS"
# },
# {
# "_target_": "Spacingd",
# "keys": ["image", "label"],
# "pixdim": [1.5, 1.5, 2.0],
# "mode": ["bilinear", "nearest"]
# },
# {
# "_target_": "ScaleIntensityRanged",
# "keys": "image",
# "a_min": -57,
# "a_max": 164,
# "b_min": 0,
# "b_max": 1,
# "clip": true
# },
# {
# "_target_": "RandCropByPosNegLabeld",
# "keys": ["image", "label"],
# "label_key": "label",
# "spatial_size": [96, 96, 96],
# "pos": 1,
# "neg": 1,
# "num_samples": 4,
# "image_key": "image",
# "image_threshold": 0
# },
# {
# "_target_": "EnsureTyped",
# "keys": ["image", "label"]
# }
# ]
# }
# }
# ```
# The train and validation image file names are organized into a list of dictionaries.
#
# Here we use `dataset` instance as 1 argument of `dataloader` by the `@` syntax, and please note that `"#"` in the reference id are interpreted as special characters to go one level further into the nested config structures. For example: `"dataset": "@train#dataset"`.
# ```json
# "dataset": {
# "_target_": "CacheDataset",
# "data": "$[{'image': i, 'label': l} for i, l in zip(@images[:-9], @labels[:-9])]",
# "transform": "@train#preprocessing",
# "cache_rate": 1.0,
# "num_workers": 4
# },
# "dataloader": {
# "_target_": "DataLoader",
# "dataset": "@train#dataset",
# "batch_size": 2,
# "shuffle": false,
# "num_workers": 4
# }
# ```
# ## Define train config - Define inference method, post-processing and event-handlers
#
# Here we use `SimpleInferer` to execute `forward()` computation for the network and add post-processing methods like `activation`, `argmax`, `one-hot`, etc. And logging into stdout and TensorBoard based on event handlers.
# ```json
# "inferer": {
# "_target_": "SimpleInferer"
# },
# "postprocessing": {
# "_target_": "Compose",
# "transforms": [
# {
# "_target_": "Activationsd",
# "keys": "pred",
# "softmax": true
# },
# {
# "_target_": "AsDiscreted",
# "keys": ["pred", "label"],
# "argmax": [true, false],
# "to_onehot": 2
# }
# ]
# },
# "handlers": [
# {
# "_target_": "StatsHandler",
# "tag_name": "train_loss",
# "output_transform": "$monai.handlers.from_engine(['loss'], first=True)"
# },
# {
# "_target_": "TensorBoardStatsHandler",
# "log_dir": "eval",
# "tag_name": "train_loss",
# "output_transform": "$monai.handlers.from_engine(['loss'], first=True)"
# }
# ]
# ```
# ## Define train config - Define Accuracy metric for training data to avoid over-fitting
#
# Here we define the `Accuracy` metric to compute on training data to help check whether the converge is expected and avoid over-fitting. Note that it's not validation step during the training.
# ```json
# "key_metric": {
# "train_accuracy": {
# "_target_": "ignite.metrics.Accuracy",
# "output_transform": "$monai.handlers.from_engine(['pred', 'label'])"
# }
# }
# ```
# ## Define train config - Define the trainer
#
# Here we use MONAI engine `SupervisedTrainer` to execute a regular training.
#
# If users have customized logic, then can put the logic in the `iteration_update` arg or implement their own `trainer` in python code and set `_target_` to the class directly.
# ```json
# "trainer": {
# "_target_": "SupervisedTrainer",
# "max_epochs": 100,
# "device": "@device",
# "train_data_loader": "@train#dataloader",
# "network": "@network",
# "loss_function": "@loss",
# "optimizer": "@optimizer",
# "inferer": "@train#inferer",
# "postprocessing": "@train#postprocessing",
# "key_train_metric": "@train#key_metric",
# "train_handlers": "@train#handlers",
# "amp": true
# }
# ```
# ## Define train config - Define the validation section
#
# Usually we need to execute validation for every N epochs during training to verify the model and save the best model.
#
# Here we don't define the `validate` section step by step as it's similar to the `train` section. The full config is available:
# https://github.com/Project-MONAI/tutorials/blob/master/modules/bundle/spleen_segmentation/configs/train.json
#
# Just show an example of `macro text replacement` to simplify the config content and avoid duplicated text. Please note that it's just token text replacement of the config content, not refer to the instantiated python objects.
# ```json
# "validate": {
# "preprocessing": {
# "_target_": "Compose",
# "transforms": [
# "%train#preprocessing#transforms#0",
# "%train#preprocessing#transforms#1",
# "%train#preprocessing#transforms#2",
# "%train#preprocessing#transforms#3",
# "%train#preprocessing#transforms#4",
# "%train#preprocessing#transforms#6"
# ]
# }
# }
# ```
# ## Define metadata information
#
# We can define a `metadata` file in the bundle, which contains the metadata information relating to the model, including what the shape and format of inputs and outputs are, what the meaning of the outputs are, what type of model is present, and other information. The structure is a dictionary containing a defined set of keys with additional user-specified keys.
#
# A typical `metadata` example is available:
# https://github.com/Project-MONAI/tutorials/blob/master/modules/bundle/spleen_segmentation/configs/metadata.json
# ## Execute training with bundle script - `run`
#
# There are several predefined scripts in MONAI bundle module to help execute `regular training`, `metadata verification base on schema`, `network input / output verification`, `export to TorchScript model`, etc.
#
# Here we leverage the `run` script and specify the ID of trainer in the config.
#
# Just define the entry point expressions in the config to execute in order, and specify the `runner_id` in CLI script.
# ```json
# "training": [
# "$monai.utils.set_determinism(seed=123)",
# "$setattr(torch.backends.cudnn, 'benchmark', True)",
# "$@train#trainer.run()"
# ]
# ```
# `python -m monai.bundle run training --config_file configs/train.json`
# ## Execute training with bundle script - Override config at runtime
#
# To override some config items at runtime, users can specify the target `id` and `value` at command line, or override the `id` with some content in another config file. Here we set the device to `cuda:1` at runtime.
#
# Please note that "#" and "$" may be meaningful syntax for some `shell` and `CLI` tools, so may need to add escape character or quotes for them in the command line, like: `"\$torch.device('cuda:1')"`. For more details: https://github.com/google/python-fire/blob/v0.4.0/fire/parser.py#L60.
# `python -m monai.bundle run training --config_file configs/train.json --device "\$torch.device('cuda:1')"`
# Override content from another config file.
# `python -m monai.bundle run training --config_file configs/train.json --network "%configs/test.json#network"`
# ## Hybrid programming with config and python code
#
# A MONAI bundle supports flexible customized logic, there are several ways to achieve this:
#
# - If defining own components like transform, loss, trainer, etc. in a python file, just use its module path in `_target_` within the config file.
# - Parse the config in your own python program and do lazy instantiation with customized logic.
#
# Here we show an example to parse the config in python code and execute the training.
parser = ConfigParser()
parser.read_config(f="configs/train.json")
parser.read_meta(f="configs/metadata.json")
# `get`/`set` configuration content, the `set` method should happen before calling `parse()`.
# original input channels 1
print(parser["network_def"]["in_channels"])
# change input channels to 4
parser["network_def"]["in_channels"] = 4
print(parser["network_def"]["in_channels"])
# Parse the config content and instantiate components.
# +
# parse the structured config content
parser.parse()
# instantiate the network component and print the network structure
net = parser.get_parsed_content("network")
print(net)
# execute training
trainer = parser.get_parsed_content("train#trainer")
trainer.run()
| modules/bundle/get_started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python API for Table Display
#
# In addition to APIs for creating and formatting BeakerX's interactive table widget, the Python runtime configures pandas to display tables with the interactive widget instead of static HTML.
import pandas as pd
from beakerx import *
from beakerx.object import beakerx
pd.read_csv('../resources/data/interest-rates.csv')
table = TableDisplay(pd.read_csv('../resources/data/interest-rates.csv'))
table.setAlignmentProviderForColumn('m3', TableDisplayAlignmentProvider.CENTER_ALIGNMENT)
table.setRendererForColumn("y10", TableDisplayCellRenderer.getDataBarsRenderer(False))
table.setRendererForType(ColumnType.Double, TableDisplayCellRenderer.getDataBarsRenderer(True))
table
# +
df = pd.read_csv('../resources/data/interest-rates.csv')
df['time'] = df['time'].str.slice(0,19).astype('datetime64[ns]')
table = TableDisplay(df)
table.setStringFormatForTimes(TimeUnit.DAYS)
table.setStringFormatForType(ColumnType.Double, TableDisplayStringFormat.getDecimalFormat(4,6))
table.setStringFormatForColumn("m3", TableDisplayStringFormat.getDecimalFormat(0, 0))
table
# +
table = TableDisplay(pd.read_csv('../resources/data/interest-rates.csv'))
table
#freeze a column
table.setColumnFrozen("y1", True)
#hide a column
table.setColumnVisible("y30", False)
table.setColumnOrder(["m3", "y1", "y5", "time", "y2"])
def config_tooltip(row, column, table):
return "The value is: " + str(table.values[row][column])
table.setToolTip(config_tooltip)
table.setDataFontSize(16)
table.setHeaderFontSize(18)
table
# +
mapListColorProvider = [
{"a": 1, "b": 2, "c": 3},
{"a": 4, "b": 5, "c": 6},
{"a": 7, "b": 8, "c": 5}
]
tabledisplay = TableDisplay(mapListColorProvider)
colors = [
[Color.LIGHT_GRAY, Color.GRAY, Color.RED],
[Color.DARK_GREEN, Color.ORANGE, Color.RED],
[Color.MAGENTA, Color.BLUE, Color.BLACK]
]
def color_provider(row, column, table):
return colors[row][column]
tabledisplay.setFontColorProvider(color_provider)
tabledisplay
# +
mapListFilter = [
{"a":1, "b":2, "c":3},
{"a":4, "b":5, "c":6},
{"a":7, "b":8, "c":5}
]
display = TableDisplay(mapListFilter)
def filter_row(row, model):
return model[row][1] == 8
display.setRowFilter(filter_row)
display
# +
table = TableDisplay(pd.read_csv('../resources/data/interest-rates.csv'))
table.addCellHighlighter(TableDisplayCellHighlighter.getHeatmapHighlighter("m3", TableDisplayCellHighlighter.FULL_ROW))
table
# -
# ### Display mode: Pandas default
beakerx.pandas_display_default()
pd.read_csv('../resources/data/interest-rates.csv')
# ### Display mode: TableDisplay Widget
beakerx.pandas_display_table()
pd.read_csv('../resources/data/interest-rates.csv')
# ## Recognized Formats
TableDisplay([{'y1':4, 'm3':2, 'z2':1}, {'m3':4, 'z2':2}])
TableDisplay({"x" : 1, "y" : 2})
# ## Programmable Table Actions
# +
mapList4 = [
{"a":1, "b":2, "c":3},
{"a":4, "b":5, "c":6},
{"a":7, "b":8, "c":5}
]
display = TableDisplay(mapList4)
def dclick(row, column, tabledisplay):
tabledisplay.values[row][column] = sum(map(int,tabledisplay.values[row]))
display.setDoubleClickAction(dclick)
def negate(row, column, tabledisplay):
tabledisplay.values[row][column] = -1 * int(tabledisplay.values[row][column])
def incr(row, column, tabledisplay):
tabledisplay.values[row][column] = int(tabledisplay.values[row][column]) + 1
display.addContextMenuItem("negate", negate)
display.addContextMenuItem("increment", incr)
display
# +
mapList4 = [
{"a":1, "b":2, "c":3},
{"a":4, "b":5, "c":6},
{"a":7, "b":8, "c":5}
]
display = TableDisplay(mapList4)
#set what happens on a double click
display.setDoubleClickAction("runDoubleClick")
display
# + tags=["runDoubleClick"]
print("runDoubleClick fired")
print(display.details)
# -
# ## Set index to DataFrame
df = pd.read_csv('../resources/data/interest-rates.csv')
df.set_index(['m3'])
df = pd.read_csv('../resources/data/interest-rates.csv')
df.index = df['time']
df
# # Update cell
# +
dataToUpdate = [
{'a':1, 'b':2, 'c':3},
{'a':4, 'b':5, 'c':6},
{'a':7, 'b':8, 'c':9}
]
tableToUpdate = TableDisplay(dataToUpdate)
tableToUpdate
# -
tableToUpdate.values[0][0] = 99
tableToUpdate.sendModel()
tableToUpdate.updateCell(2,"c",121)
tableToUpdate.sendModel()
# ## HTML format
#
# HTML format allows markup and styling of the cell's content. Interactive JavaScript is not supported however.
table = TableDisplay({
'w': '$2 \\sigma$',
'x': '<em style="color:red">italic red</em>',
'y': '<b style="color:blue">bold blue</b>',
'z': 'strings without markup work fine too',
})
table.setStringFormatForColumn("Value", TableDisplayStringFormat.getHTMLFormat())
table
# ## Auto linking of URLs
#
# The normal string format automatically detects URLs and links them. An underline appears when the mouse hovers over such a string, and when you click it opens in a new window.
TableDisplay({'Two Sigma': 'http://twosigma.com', 'BeakerX': 'http://BeakerX.com'})
| doc/python/TableAPI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
# %matplotlib inline
from dipy.reconst.dti import fractional_anisotropy, color_fa
from argparse import ArgumentParser
from scipy import ndimage
import os
import re
import numpy as np
import nibabel as nb
import sys
import matplotlib
# matplotlib.use('Agg') # very important above pyplot import
import matplotlib.pyplot as plt
# + deletable=true editable=true
from dipy.reconst.dti import from_lower_triangular
# + deletable=true editable=true
img = nb.load('fibers/dogsigma_0gausigma_0tensorfsl.nii')
# + deletable=true editable=true
data = img.get_data()
output = from_lower_triangular(data)
output_ds = output[:, :, :, :, :]
print output.shape
print output_ds.shape
# + deletable=true editable=true
FA = fractional_anisotropy(output_ds)
FA = np.clip(FA, 0, 1)
FA[np.isnan(FA)] = 0
print FA.shape
# + deletable=true editable=true
from dipy.reconst.dti import decompose_tensor
# + deletable=true editable=true
evalues, evectors = decompose_tensor(output_ds)
print evectors[..., 0, 0].shape
print evectors.shape[-2:]
# + deletable=true editable=true
print FA[:, :, :, 0].shape
# + deletable=true editable=true
RGB = color_fa(FA[:, :, :, 0], evectors)
nb.save(nb.Nifti1Image(np.array(255 * RGB, 'uint8'), img.get_affine()), 'tensor_rgb_upper.nii.gz')
# + deletable=true editable=true
def plot_rgb(im):
plt.rcParams.update({'axes.labelsize': 'x-large',
'axes.titlesize': 'x-large'})
if im.shape == (182, 218, 182):
x = [78, 90, 100]
y = [82, 107, 142]
z = [88, 103, 107]
else:
shap = im.shape
x = [int(shap[0]*0.35), int(shap[0]*0.51), int(shap[0]*0.65)]
y = [int(shap[1]*0.35), int(shap[1]*0.51), int(shap[1]*0.65)]
z = [int(shap[2]*0.35), int(shap[2]*0.51), int(shap[2]*0.65)]
coords = (x, y, z)
labs = ['Sagittal Slice (YZ fixed)',
'Coronal Slice (XZ fixed)',
'Axial Slice (XY fixed)']
var = ['X', 'Y', 'Z']
idx = 0
for i, coord in enumerate(coords):
for pos in coord:
idx += 1
ax = plt.subplot(3, 3, idx)
ax.set_title(var[i] + " = " + str(pos))
if i == 0:
image = ndimage.rotate(im[pos, :, :], 90)
elif i == 1:
image = ndimage.rotate(im[:, pos, :], 90)
else:
image = im[:, :, pos]
if idx % 3 == 1:
ax.set_ylabel(labs[i])
ax.yaxis.set_ticks([0, image.shape[0]/2, image.shape[0] - 1])
ax.xaxis.set_ticks([0, image.shape[1]/2, image.shape[1] - 1])
plt.imshow(image)
fig = plt.gcf()
fig.set_size_inches(12.5, 10.5, forward=True)
return fig
# + deletable=true editable=true
affine = img.get_affine()
fa = nb.Nifti1Image(np.array(255 * RGB, 'uint8'), affine)
im = fa.get_data()
# + deletable=true editable=true
# print np.asarray(fa)
# + deletable=true editable=true
fig = plot_rgb(im)
# + deletable=true editable=true
import os
# + deletable=true editable=true
from PIL import Image
im = plt.imread('fibers/v100/ch0/luke40.tiff')
plt.imshow(im)
# + deletable=true editable=true
import dipy.reconst.dti as dti
from dipy.reconst.dti import fractional_anisotropy
from dipy.tracking.local import (ThresholdTissueClassifier, LocalTracking)
classifier = ThresholdTissueClassifier(FA[0], .2)
# + deletable=true editable=true
from dipy.data import default_sphere
from dipy.direction import DeterministicMaximumDirectionGetter
from dipy.io.trackvis import save_trk
from dipy.reconst.csdeconv import ConstrainedSphericalDeconvModel
from dipy.data import get_sphere
sphere = get_sphere('symmetric724')
from dipy.reconst.dti import quantize_evecs
peak_indices = quantize_evecs(evectors, sphere.vertices)
# + deletable=true editable=true
from dipy.tracking.eudx import EuDX
eu = EuDX(FA.astype('f8')[0], peak_indices, seeds=50000, odf_vertices = sphere.vertices, a_low=0.2)
tensor_streamlines = [streamline for streamline in eu]
# + deletable=true editable=true
tensor_streamlines_trk = ((sl, None, None) for sl in tensor_streamlines)
# + deletable=true editable=true
from argparse import ArgumentParser
from dipy.viz import window, actor
import numpy as np
def visualize(fibers, outf=None):
"""
Takes fiber streamlines and visualizes them using DiPy
Required Arguments:
- fibers:
fiber streamlines in a list as returned by DiPy
Optional Arguments:
- save:
flag indicating whether or not you want the image saved
to disk after being displayed
"""
# Initialize renderer
renderer = window.Renderer()
# Add streamlines as a DiPy viz object
stream_actor = actor.line(fibers)
# Set camera orientation properties
# TODO: allow this as an argument
renderer.set_camera() # args are: position=(), focal_point=(), view_up=()
# Add streamlines to viz session
renderer.add(stream_actor)
# Display fibers
# TODO: allow size of window as an argument
window.show(renderer, size=(600, 600), reset_camera=False)
# Saves file, if you're into that sort of thing...
if outf is not None:
window.record(renderer, out_path=outf, size=(600, 600))
# + deletable=true editable=true
visualize(tensor_streamlines)
# -
import vtk
# + deletable=true editable=true
import vtk
from IPython.display import Image
def vtk_show(renderer, width=400, height=300):
"""
Takes vtkRenderer instance and returns an IPython Image with the rendering.
"""
renderWindow = vtk.vtkRenderWindow()
renderWindow.SetOffScreenRendering(1)
renderWindow.AddRenderer(renderer)
renderWindow.SetSize(width, height)
renderWindow.Render()
windowToImageFilter = vtk.vtkWindowToImageFilter()
windowToImageFilter.SetInput(renderWindow)
windowToImageFilter.Update()
writer = vtk.vtkPNGWriter()
writer.SetWriteToMemory(1)
writer.SetInputConnection(windowToImageFilter.GetOutputPort())
writer.Write()
data = str(buffer(writer.GetResult()))
return Image(data)
# + deletable=true editable=true
renderer = window.Renderer()
# Add streamlines as a DiPy viz object
stream_actor = actor.line(tensor_streamlines)
# Set camera orientation properties
# TODO: allow this as an argument
renderer.set_camera() # args are: position=(), focal_point=(), view_up=()
# Add streamlines to viz session
renderer.add(stream_actor)
vtk_show(renderer)
# + deletable=true editable=true
| Jupyter/fibers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introducing AI Platform Training Service
# **Learning Objectives:**
# - Learn how to make code compatible with AI Platform Training Service
# - Train your model using cloud infrastructure via AI Platform Training Service
# - Deploy your model behind a production grade REST API using AI Platform Training Service
#
# ## Introduction
#
# In this notebook we'll make the jump from training and predicting locally, to do doing both in the cloud. We'll take advantage of Google Cloud's [AI Platform Training Service](https://cloud.google.com/ai-platform/).
#
# AI Platform Training Service is a managed service that allows the training and deployment of ML models without having to provision or maintain servers. The infrastructure is handled seamlessly by the managed service for us.
# +
# Uncomment and run if you need to update your Google SDK
# # !sudo apt-get update && sudo apt-get --only-upgrade install google-cloud-sdk
# -
# ## Make code compatible with AI Platform Training Service
# In order to make our code compatible with AI Platform Training Service we need to make the following changes:
#
# 1. Upload data to Google Cloud Storage
# 2. Move code into a Python package
# 3. Modify code to read data from and write checkpoint files to GCS
# ### Upload data to Google Cloud Storage (GCS)
#
# Cloud services don't have access to our local files, so we need to upload them to a location the Cloud servers can read from. In this case we'll use GCS.
#
# Specify your project name and bucket name in the cell below.
PROJECT = "qwiklabs-gcp-00-34ffb0f0dc65" # Replace with your PROJECT
BUCKET = "cloud-training-bucket" # Replace with your BUCKET
REGION = "us-central1" # Choose an available region for AI Platform Training Service
TFVERSION = "1.14" # TF version for AI Platform Training Service to use
# Jupyter allows the subsitution of python variables into bash commands when using the `!<cmd>` format.
# It is also possible using the `%%bash` magic but requires an [additional parameter](https://stackoverflow.com/questions/19579546/can-i-access-python-variables-within-a-bash-or-script-ipython-notebook-c).
# !gcloud config set project {PROJECT}
# !gsutil mb -l {REGION} gs://{BUCKET}
# !gsutil -m cp *.csv gs://{BUCKET}/taxifare/smallinput/
# ### Move code into a python package
#
# When you execute a AI Platform Training Service training job, the service zips up your code and ships it to the Cloud so it can be run on Cloud infrastructure. In order to do this AI Platform Training Service requires your code to be a Python package.
#
# A Python package is simply a collection of one or more `.py` files along with an `__init__.py` file to identify the containing directory as a package. The `__init__.py` sometimes contains initialization code but for our purposes an empty file suffices.
# #### Create Package Directory and \_\_init\_\_.py
#
# The bash command `touch` creates an empty file in the specified location.
# + language="bash"
# mkdir taxifaremodel
# touch taxifaremodel/__init__.py
# -
# #### Paste existing code into model.py
#
# A Python package requires our code to be in a .py file, as opposed to notebook cells. So we simply copy and paste our existing code for the previous notebook into a single file.
#
# The %%writefile magic writes the contents of its cell to disk with the specified name.
# #### **Exercise 1**
#
# In the cell below, write the content of the `model.py` to the file `taxifaremodel/model.py`. This will allow us to package the model we
# developed in the previous labs so that we can deploy it to AI Platform Training Service. You'll also need to reuse the input functions and the `EvalSpec`, `TrainSpec`, `RunConfig`, etc. that we implemented in the previous labs.
#
# Complete all the TODOs in the cell below by copy/pasting the code we developed in the previous labs. This will write all the necessary components we developed in our notebook to a single `model.py` file.
#
# Once we have the code running well locally, we will execute the next cells to train and deploy your packaged model to AI Platform Training Service.
# +
# %%writefile taxifaremodel/model.py
# TODO: Your code goes here. Import the necessary libraries (e.g. tensorflow, etc)
CSV_COLUMN_NAMES = # TODO: Your code goes here
CSV_DEFAULTS = # TODO: Your code goes here
FEATURE_NAMES = # TODO: Your code goes here
def parse_row(row):
# TODO: Your code goes here
return features, label
def read_dataset(csv_path):
# TODO: Your code goes here
return dataset
def train_input_fn(csv_path, batch_size = 128):
# TODO: Your code goes here
return dataset
def eval_input_fn(csv_path, batch_size = 128):
# TODO: Your code goes here
return dataset
def serving_input_receiver_fn():
# TODO: Your code goes here
return tf.estimator.export.ServingInputReceiver(features = features, receiver_tensors = receiver_tensors)
def my_rmse(labels, predictions):
# TODO: Your code goes here
return {"rmse": tf.metrics.root_mean_squared_error(labels = labels, predictions = pred_values)}
def create_model(model_dir, train_steps):
# TODO: Your code goes here
return model
def train_and_evaluate(params):
OUTDIR = params["output_dir"]
TRAIN_DATA_PATH = params["train_data_path"]
EVAL_DATA_PATH = params["eval_data_path"]
TRAIN_STEPS = params["train_steps"]
model = # TODO: Your code goes here.
train_spec = # TODO: Your code goes here
exporter = # TODO: Your code goes here
eval_spec = # TODO: Your code goes here
tf.logging.set_verbosity(tf.logging.INFO)
shutil.rmtree(path = OUTDIR, ignore_errors = True)
tf.estimator.train_and_evaluate(estimator = model, train_spec = train_spec, eval_spec = eval_spec)
# -
# ### Modify code to read data from and write checkpoint files to GCS
#
# If you look closely above, you'll notice two changes to the code
#
# 1. The input function now supports reading a list of files matching a file name pattern instead of just a single CSV
# - This is useful because large datasets tend to exist in shards.
# 2. The train and evaluate portion is wrapped in a function that takes a parameter dictionary as an argument.
# - This is useful because the output directory, data paths and number of train steps will be different depending on whether we're training locally or in the cloud. Parametrizing allows us to use the same code for both.
#
# We specify these parameters at run time via the command line. Which means we need to add code to parse command line parameters and invoke `train_and_evaluate()` with those params. This is the job of the `task.py` file.
#
# Exposing parameters to the command line also allows us to use AI Platform Training Service's automatic hyperparameter tuning feature which we'll cover in a future lesson.
# #### **Exercise 2**
#
# Add two additional command line parameter parsers to the list we've started below. You should add code to parse command line parameters for the `output_dir` and the `job-dir`. Look at the examples below to make sure you have the correct format, including a `help` description and `required` specification.
# +
# %%writefile taxifaremodel/task.py
import argparse
import json
import os
from . import model
if __name__ == "__main__":
parser = argparse.ArgumentParser()
parser.add_argument(
"--train_data_path",
help = "GCS or local path to training data",
required = True
)
parser.add_argument(
"--train_steps",
help = "Steps to run the training job for (default: 1000)",
type = int,
default = 1000
)
parser.add_argument(
"--eval_data_path",
help = "GCS or local path to evaluation data",
required = True
)
parser.add_argument(
# TODO: Your code goes here
)
parser.add_argument(
# TODO: Your code goes here
)
args = parser.parse_args().__dict__
model.train_and_evaluate(args)
# -
# ## Train using AI Platform Training Service (local)
#
# AI Platform Training Service comes with a local test tool ([`gcloud ai-platform local train`](https://cloud.google.com/sdk/gcloud/reference/ml-engine/local/train)) to ensure we've packaged our code directly. It's best to first run that for a few steps before trying a Cloud job.
#
# The arguments before `-- \` are for AI Platform Training Service
# - package-path: speficies the location of the Python package
# - module-name: specifies which `.py` file should be run within the package. `task.py` is our entry point so we specify that
#
# The arguments after `-- \` are sent to our `task.py`.
# %%time
# !gcloud ai-platform local train \
# --package-path=taxifaremodel \
# --module-name=taxifaremodel.task \
# -- \
# --train_data_path=taxi-train.csv \
# --eval_data_path=taxi-valid.csv \
# --train_steps=1 \
# --output_dir=taxi_trained
# ## Train using AI Platform Training Service (Cloud)
#
# To submit to the Cloud we use [`gcloud ai-platform jobs submit training [jobname]`](https://cloud.google.com/sdk/gcloud/reference/ml-engine/jobs/submit/training) and simply specify some additional parameters for AI Platform Training Service:
# - jobname: A unique identifier for the Cloud job. We usually append system time to ensure uniqueness
# - job-dir: A GCS location to upload the Python package to
# - runtime-version: Version of TF to use. Defaults to 1.0 if not specified
# - python-version: Version of Python to use. Defaults to 2.7 if not specified
# - region: Cloud region to train in. See [here](https://cloud.google.com/ml-engine/docs/tensorflow/regions) for supported AI Platform Training Service regions
#
# Below the `-- \` note how we've changed our `task.py` args to be GCS locations
OUTDIR = "gs://{}/taxifare/trained_small".format(BUCKET)
# !gsutil -m rm -rf {OUTDIR} # start fresh each time
# !gcloud ai-platform jobs submit training taxifare_$(date -u +%y%m%d_%H%M%S) \
# --package-path=taxifaremodel \
# --module-name=taxifaremodel.task \
# --job-dir=gs://{BUCKET}/taxifare \
# --python-version=3.5 \
# --runtime-version={TFVERSION} \
# --region={REGION} \
# -- \
# --train_data_path=gs://{BUCKET}/taxifare/smallinput/taxi-train.csv \
# --eval_data_path=gs://{BUCKET}/taxifare/smallinput/taxi-valid.csv \
# --train_steps=1000 \
# --output_dir={OUTDIR}
# You can track your job and view logs using [cloud console](https://console.cloud.google.com/mlengine/jobs). It will take 5-10 minutes to complete. **Wait until the job finishes before moving on.**
# ## Deploy model
#
# Now let's take our exported SavedModel and deploy it behind a REST API. To do so we'll use AI Platform Training Service's managed TF Serving feature which auto-scales based on load.
# !gsutil ls gs://{BUCKET}/taxifare/trained_small/export/exporter
# AI Platform Training Service uses a model versioning system. First you create a model folder, and within the folder you create versions of the model.
#
# Note: You will see an error below if the model folder already exists, it is safe to ignore
VERSION='v1'
# !gcloud ai-platform models create taxifare --regions us-central1
# !gcloud ai-platform versions delete {VERSION} --model taxifare --quiet
# !gcloud ai-platform versions create {VERSION} --model taxifare \
# --origin $(gsutil ls gs://{BUCKET}/taxifare/trained_small/export/exporter | tail -1) \
# --python-version=3.5 \
# --runtime-version {TFVERSION}
# ## Online prediction
#
# Now that we have deployed our model behind a production grade REST API, we can invoke it remotely.
#
# We could invoke it directly calling the REST API with an HTTP POST request [reference docs](https://cloud.google.com/ml-engine/reference/rest/v1/projects/predict), however AI Platform Training Service provides an easy way to invoke it via command line.
# ### Invoke prediction REST API via command line
# First we write our prediction requests to file in json format
# %%writefile ./test.json
{"dayofweek": 1, "hourofday": 0, "pickuplon": -73.885262, "pickuplat": 40.773008, "dropofflon": -73.987232, "dropofflat": 40.732403}
# Then we use [`gcloud ai-platform predict`](https://cloud.google.com/sdk/gcloud/reference/ml-engine/predict) and specify the model name and location of the json file. Since we don't explicitly specify `--version`, the default model version will be used.
#
# Since we only have one version it is already the default, but if we had multiple model versions we can designate the default using [`gcloud ai-platform versions set-default`](https://cloud.google.com/sdk/gcloud/reference/ml-engine/versions/set-default) or using [cloud console](https://pantheon.corp.google.com/mlengine/models)
# !gcloud ai-platform predict --model=taxifare --json-instances=./test.json
# ### Invoke prediction REST API via python
# #### **Exercise 3**
#
# In the cell below, use the Google Python client library to query the model you just deployed on AI Platform Training Service. Find the estimated taxi fare for a ride with the following properties
# - ride occurs on Monday
# - at 8:00 am
# - pick up at (40.773, -73.885)
# - drop off at (40.732, -73.987)
#
# Have a look at this post and examples on ["Using the Python Client Library"](https://cloud.google.com/ml-engine/docs/tensorflow/python-client-library) and ["Getting Online Predictions"](https://cloud.google.com/ml-engine/docs/tensorflow/online-predict) from Google Cloud.
# +
from googleapiclient import discovery
from oauth2client.client import GoogleCredentials
import json
credentials = # TODO: Your code goes here
api = # TODO: Your code goes here
request_data = {"instances":
[
{
# TODO: Your code goes here
}
]
}
parent = # TODO: Your code goes here
response = # TODO: Your code goes here
print("response = {0}".format(response))
# -
# ## Challenge exercise
#
# Modify your solution to the challenge exercise in e_traineval.ipynb appropriately. Make sure that you implement training and deployment. Increase the size of your dataset by 10x since you are running on the cloud. Does your accuracy improve?
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/02_tensorflow/labs/f_ai_platform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## There is some problem with the previous code
# So I made modification
# +
import random
from random import seed
from random import random
from numpy import where
from collections import Counter
from sklearn.datasets import make_blobs
from matplotlib import pyplot
import numpy as np
import matplotlib.pyplot as plt
from sklearn import mixture
from sklearn.neighbors import KernelDensity
import random
# -
def my_circle(center, rx, ry, Nmax):
R = 1
x2=[]
y2=[]
for i in range(Nmax):
r2=np.sqrt(random.random())
theta2=2 * np.pi * random.random()
x2.append(rx*r2 * np.cos(theta2) + center[0])
y2.append(ry*r2 * np.sin(theta2) + center[1])
return np.transpose([x2,y2])
def my_plot(data,lab,counter):
for label, _ in counter.items():
row_ix = where(lab == label)[0]
pyplot.scatter(data[row_ix, 0], data[row_ix, 1], label=str(label))
pyplot.legend()
pyplot.xlim([0, 1])
pyplot.ylim([0, 1])
pyplot.show()
# +
def data1a():
seed(30)
X1=my_circle([0.3,0.5],0.25,0.4,750)
X2=my_circle([0.7,0.5],0.25,0.4,750)
y1=[0] * 750
y2=[1] * 750
X_1=np.concatenate((X1,X2))
y_1=np.concatenate((y1, y2))
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
def data1b():
seed(30)
X1=my_circle([0.4,0.2],0.25,0.15,375)
X2=my_circle([0.6,0.5],0.25,0.15,750)
X3=my_circle([0.4,0.8],0.25,0.15,375)
y1=[0] * 375
y2=[1] * 750
y3=[0] * 375
X_1=np.concatenate((X1,X2,X3))
y_1=np.concatenate((y1, y2,y3))
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
def data1c():
seed(30)
X1=my_circle([0.3,0.2],0.25,0.12,375)
X2=my_circle([0.7,0.4],0.25,0.12,375)
X3=my_circle([0.3,0.6],0.25,0.12,375)
X4=my_circle([0.7,0.8],0.25,0.12,375)
y1=[0] * 375
y2=[1] * 375
y3=[0] * 375
y4=[1] * 375
X_1=np.concatenate((X1,X2,X3,X4))
y_1=np.concatenate((y1, y2,y3,y4))
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
def data2a():
seed(30)
X1=my_circle([0.25,0.25],0.2,0.2,375)
X2=my_circle([0.25,0.75],0.2,0.2,375)
X3=my_circle([0.75,0.75],0.2,0.2,375)
X4=my_circle([0.75,0.25],0.2,0.2,375)
y1=[0] * 375
y2=[1] * 375
y3=[0] * 375
y4=[1] * 375
X_1=np.concatenate((X1,X2,X3,X4))
y_1=np.concatenate((y1, y2,y3,y4))
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
def data2b():
seed(30)
r=0.12
n=166
X1=my_circle([0.2,0.2],r,r,n)
X2=my_circle([0.5,0.2],r,r,n)
X3=my_circle([0.8,0.2],r,r,n)
X4=my_circle([0.2,0.5],r,r,n)
X5=my_circle([0.5,0.5],r,r,n)
X6=my_circle([0.8,0.5],r,r,n)
X7=my_circle([0.2,0.8],r,r,n)
X8=my_circle([0.5,0.8],r,r,n)
X9=my_circle([0.8,0.8],r,r,n)
y1=[0] * n
y2=[1] * n
y3=[0] * n
y4=[1] * n
y5=[0] * n
y6=[1] * n
y7=[0] * n
y8=[1] * n
y9=[0] * n
X_1=np.concatenate((X1,X2,X3,X4,X5,X6,X7,X8,X9))
y_1=np.concatenate((y1, y2,y3,y4,y5,y6,y7,y8,y9))
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
def data2c():
seed(30)
r=0.1
n=93
X1=my_circle([0.15,0.15],r,r,n)
X2=my_circle([0.38,0.15],r,r,n)
X3=my_circle([0.62,0.15],r,r,n)
X4=my_circle([0.85,0.15],r,r,n)
X5=my_circle([0.15,0.38],r,r,n)
X6=my_circle([0.38,0.38],r,r,n)
X7=my_circle([0.62,0.38],r,r,n)
X8=my_circle([0.85,0.38],r,r,n)
X9=my_circle([0.15,0.62],r,r,n)
X10=my_circle([0.38,0.62],r,r,n)
X11=my_circle([0.62,0.62],r,r,n)
X12=my_circle([0.85,0.62],r,r,n)
X13=my_circle([0.15,0.85],r,r,n)
X14=my_circle([0.38,0.85],r,r,n)
X15=my_circle([0.62,0.85],r,r,n)
X16=my_circle([0.85,0.85],r,r,n)
y1=[0] * n
y2=[1] * n
y3=[0] * n
y4=[1] * n
y5=[1] * n
y6=[0] * n
y7=[1] * n
y8=[0] * n
y9=[0] * n
y10=[1] * n
y11=[0] * n
y12=[1] * n
y13=[1] * n
y14=[0] * n
y15=[1] * n
y16=[0] * n
X_1=np.concatenate((X1,X2,X3,X4,X5,X6,X7,X8,X9,X10,X11,X12,X13,X14,X15,X16))
y_1=np.concatenate((y1, y2,y3,y4,y5,y6,y7,y8,y9,y10,y11,y12,y13,y14,y15,y16))
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
def data3a():
seed(30)
X1 = []
radius = 0.25
n=750
index = 0
while index < n:
x=random.random()
y=random.random()
if (x-0.5)**2 + (y-0.5)**2 > radius**2:
X1 = X1 + [[x,y]]
index = index + 1
y1=[1] *750
X2=my_circle([0.5,0.5],radius,radius,750)
y2=[0] *750
X_1=np.concatenate((X1,X2))
y_1=np.concatenate((y1, y2))
# Generate uniform noise
counter = Counter(y_1)
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
def data3b():
seed(30)
X1 = []
radius = 0.15
n=750
index = 0
while index < n:
x=random.random()
y=random.random()
if ((x-0.25)**2 + (y-0.25)**2 > radius**2 and (x-0.75)**2 + (y-0.75)**2 > radius**2):
X1 = X1 + [[x,y]]
index = index + 1
y1=[1] *750
X2=my_circle([0.25,0.25],radius,radius,375)
X3=my_circle([0.75,0.75],radius,radius,375)
y2=[0] *750
X_1=np.concatenate((X1,X2,X3))
y_1=np.concatenate((y1, y2))
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
def data3c():
seed(30)
X1 = []
radius = 0.15
n=750
n2=187
index = 0
while index < n:
x=random.random()
y=random.random()
if ((x-0.25)**2 + (y-0.25)**2 > radius**2 and (x-0.75)**2 + (y-0.75)**2 > radius**2 and (x-0.75)**2 + (y-0.25)**2 > radius**2 and (x-0.25)**2 + (y-0.75)**2 > radius**2 ):
X1 = X1 + [[x,y]]
index = index + 1
y1=[1] *750
X2=my_circle([0.25,0.25],radius,radius,n2)
X3=my_circle([0.75,0.75],radius,radius,n2)
X4=my_circle([0.75,0.25],radius,radius,n2)
X5=my_circle([0.25,0.75],radius,radius,n2)
y2=[0] *748
X_1=np.concatenate((X1,X2,X3,X4,X5))
y_1=np.concatenate((y1, y2))
# Generate uniform noise
counter = Counter(y_1)
# Generate uniform noise
counter = Counter(y_1)
return X_1, y_1, counter
# +
# Plot
my_plot(data1a()[0],data1a()[1],data1a()[2])
# Plot
my_plot(data1b()[0],data1b()[1],data1b()[2])
# Plot
my_plot(data1c()[0],data1c()[1],data1c()[2])
# Plot
my_plot(data2a()[0],data2a()[1],data2a()[2])
# Plot
my_plot(data2b()[0],data2b()[1],data2b()[2])
# Plot
my_plot(data2c()[0],data2c()[1],data2c()[2])
# Plot
my_plot(data3a()[0],data3a()[1],data3a()[2])
# Plot
my_plot(data3b()[0],data3b()[1],data3b()[2])
# Plot
my_plot(data3c()[0],data3c()[1],data3c()[2])
| dataset/.ipynb_checkpoints/Data Generation Draft 3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > This is one of the 100 recipes of the [IPython Cookbook](http://ipython-books.github.io/), the definitive guide to high-performance scientific computing and data science in Python.
#
# # 5.5. Ray tracing: Cython array buffers
# In this example, we will render a sphere with a diffuse and specular material. The principle is to model a scene with a light source and a camera, and use the physical properties of light propagation to calculate the light intensity and color of every pixel of the screen.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext cythonmagic
# ## Take 1
# + language="cython"
# import numpy as np
# cimport numpy as np
# from numpy import dot
# from libc.math cimport sqrt
#
# DBL = np.double
# ctypedef np.double_t DBL_C
# INT = np.int
# ctypedef np.int_t INT_C
# cdef INT_C w, h
#
# w, h = 200, 200 # Size of the screen in pixels.
#
# def normalize(np.ndarray[DBL_C, ndim=1] x):
# # This function normalizes a vector.
# x /= np.linalg.norm(x)
# return x
#
# def intersect_sphere(np.ndarray[DBL_C, ndim=1] O, np.ndarray[DBL_C, ndim=1] D,
# np.ndarray[DBL_C, ndim=1] S, DBL_C R):
# # Return the distance from O to the intersection
# # of the ray (O, D) with the sphere (S, R), or
# # +inf if there is no intersection.
# # O and S are 3D points, D (direction) is a
# # normalized vector, R is a scalar.
#
# cdef DBL_C a, b, c, disc, distSqrt, q, t0, t1
# cdef np.ndarray[DBL_C, ndim=1] OS
#
# a = dot(D, D)
# OS = O - S
# b = 2 * dot(D, OS)
# c = dot(OS, OS) - R*R
# disc = b*b - 4*a*c
# if disc > 0:
# distSqrt = np.sqrt(disc)
# q = (-b - distSqrt) / 2.0 if b < 0 \
# else (-b + distSqrt) / 2.0
# t0 = q / a
# t1 = c / q
# t0, t1 = min(t0, t1), max(t0, t1)
# if t1 >= 0:
# return t1 if t0 < 0 else t0
# return np.inf
#
# def trace_ray(np.ndarray[DBL_C, ndim=1] O, np.ndarray[DBL_C, ndim=1] D,
# np.ndarray[DBL_C, ndim=1] position,
# np.ndarray[DBL_C, ndim=1] color,
# np.ndarray[DBL_C, ndim=1] L,
# np.ndarray[DBL_C, ndim=1] color_light):
#
# cdef DBL_C t
# cdef np.ndarray[DBL_C, ndim=1] M, N, toL, toO, col
#
# # Find first point of intersection with the scene.
# t = intersect_sphere(O, D, position, radius)
# # No intersection?
# if t == np.inf:
# return
# # Find the point of intersection on the object.
# M = O + D * t
# N = normalize(M - position)
# toL = normalize(L - M)
# toO = normalize(O - M)
# # Ambient light.
# col = ambient * np.ones(3)
# # Lambert shading (diffuse).
# col += diffuse * max(dot(N, toL), 0) * color
# # Blinn-Phong shading (specular).
# col += specular_c * color_light * \
# max(dot(N, normalize(toL + toO)), 0) \
# ** specular_k
# return col
#
# def run():
# cdef np.ndarray[DBL_C, ndim=3] img
# img = np.zeros((h, w, 3))
# cdef INT_C i, j
# cdef DBL_C x, y
# cdef np.ndarray[DBL_C, ndim=1] O, Q, D, col, position, color, L, color_light
#
# # Sphere properties.
# position = np.array([0., 0., 1.])
# color = np.array([0., 0., 1.])
# L = np.array([5., 5., -10.])
# color_light = np.ones(3)
#
# # Camera.
# O = np.array([0., 0., -1.]) # Position.
# Q = np.array([0., 0., 0.]) # Pointing to.
#
# # Loop through all pixels.
# for i, x in enumerate(np.linspace(-1., 1., w)):
# for j, y in enumerate(np.linspace(-1., 1., h)):
# # Position of the pixel.
# Q[0], Q[1] = x, y
# # Direction of the ray going through the optical center.
# D = normalize(Q - O)
# # Launch the ray and get the color of the pixel.
# col = trace_ray(O, D, position, color, L, color_light)
# if col is None:
# continue
# img[h - j - 1, i, :] = np.clip(col, 0, 1)
# return img
#
# cdef DBL_C radius, ambient, diffuse, specular_k, specular_c
#
# # Sphere and light properties.
# radius = 1.
# diffuse = 1.
# specular_c = 1.
# specular_k = 50.
# ambient = .05
# -
img = run()
plt.imshow(img);
plt.xticks([]); plt.yticks([]);
# %timeit -n1 -r1 run()
# ## Take 2
# In this version, we rewrite normalize in pure C.
# + language="cython"
# import numpy as np
# cimport numpy as np
# from numpy import dot
# from libc.math cimport sqrt
#
# DBL = np.double
# ctypedef np.double_t DBL_C
# INT = np.int
# ctypedef np.int_t INT_C
# cdef INT_C w, h
#
# w, h = 200, 200 # Size of the screen in pixels.
#
# # normalize is now a pure C function that does not make
# # use NumPy for the computations
# cdef normalize(np.ndarray[DBL_C, ndim=1] x):
# cdef DBL_C n
# n = sqrt(x[0] * x[0] + x[1] * x[1] + x[2] * x[2])
# x[0] /= n
# x[1] /= n
# x[2] /= n
# return x
#
# def intersect_sphere(np.ndarray[DBL_C, ndim=1] O, np.ndarray[DBL_C, ndim=1] D,
# np.ndarray[DBL_C, ndim=1] S, DBL_C R):
# # Return the distance from O to the intersection
# # of the ray (O, D) with the sphere (S, R), or
# # +inf if there is no intersection.
# # O and S are 3D points, D (direction) is a
# # normalized vector, R is a scalar.
#
# cdef DBL_C a, b, c, disc, distSqrt, q, t0, t1
# cdef np.ndarray[DBL_C, ndim=1] OS
#
# a = dot(D, D)
# OS = O - S
# b = 2 * dot(D, OS)
# c = dot(OS, OS) - R*R
# disc = b*b - 4*a*c
# if disc > 0:
# distSqrt = np.sqrt(disc)
# q = (-b - distSqrt) / 2.0 if b < 0 \
# else (-b + distSqrt) / 2.0
# t0 = q / a
# t1 = c / q
# t0, t1 = min(t0, t1), max(t0, t1)
# if t1 >= 0:
# return t1 if t0 < 0 else t0
# return np.inf
#
# def trace_ray(np.ndarray[DBL_C, ndim=1] O, np.ndarray[DBL_C, ndim=1] D,
# np.ndarray[DBL_C, ndim=1] position,
# np.ndarray[DBL_C, ndim=1] color,
# np.ndarray[DBL_C, ndim=1] L,
# np.ndarray[DBL_C, ndim=1] color_light):
#
# cdef DBL_C t
# cdef np.ndarray[DBL_C, ndim=1] M, N, toL, toO, col
#
# # Find first point of intersection with the scene.
# t = intersect_sphere(O, D, position, radius)
# # No intersection?
# if t == np.inf:
# return
# # Find the point of intersection on the object.
# M = O + D * t
# N = normalize(M - position)
# toL = normalize(L - M)
# toO = normalize(O - M)
# # Ambient light.
# col = ambient * np.ones(3)
# # Lambert shading (diffuse).
# col += diffuse * max(dot(N, toL), 0) * color
# # Blinn-Phong shading (specular).
# col += specular_c * color_light * \
# max(dot(N, normalize(toL + toO)), 0) \
# ** specular_k
# return col
#
# def run():
# cdef np.ndarray[DBL_C, ndim=3] img
# img = np.zeros((h, w, 3))
# cdef INT_C i, j
# cdef DBL_C x, y
# cdef np.ndarray[DBL_C, ndim=1] O, Q, D, col, position, color, L, color_light
#
# # Sphere properties.
# position = np.array([0., 0., 1.])
# color = np.array([0., 0., 1.])
# L = np.array([5., 5., -10.])
# color_light = np.ones(3)
#
# # Camera.
# O = np.array([0., 0., -1.]) # Position.
# Q = np.array([0., 0., 0.]) # Pointing to.
#
# # Loop through all pixels.
# for i, x in enumerate(np.linspace(-1., 1., w)):
# for j, y in enumerate(np.linspace(-1., 1., h)):
# # Position of the pixel.
# Q[0], Q[1] = x, y
# # Direction of the ray going through the optical center.
# D = normalize(Q - O)
# # Launch the ray and get the color of the pixel.
# col = trace_ray(O, D, position, color, L, color_light)
# if col is None:
# continue
# img[h - j - 1, i, :] = np.clip(col, 0, 1)
# return img
#
# cdef DBL_C radius, ambient, diffuse, specular_k, specular_c
#
# # Sphere and light properties.
# radius = 1.
# diffuse = 1.
# specular_c = 1.
# specular_k = 50.
# ambient = .05
# -
img = run()
plt.imshow(img);
plt.xticks([]); plt.yticks([]);
# %timeit -n1 -r1 run()
# > You'll find all the explanations, figures, references, and much more in the book (to be released later this summer).
#
# > [IPython Cookbook](http://ipython-books.github.io/), by [<NAME>](http://cyrille.rossant.net), Packt Publishing, 2014 (500 pages).
| notebooks/chapter05_hpc/05_ray_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load and Visualize FashionMNIST
# ---
# In this notebook, we load and look at images from the [Fashion-MNIST database](https://github.com/zalandoresearch/fashion-mnist).
#
# The first step in any classification problem is to look at the dataset you are working with. This will give you some details about the format of images and labels, as well as some insight into how you might approach defining a network to recognize patterns in such an image set.
#
# PyTorch has some built-in datasets that you can use, and FashionMNIST is one of them; it has already been dowloaded into the `data/` directory in this notebook, so all we have to do is load these images using the FashionMNIST dataset class *and* load the data in batches with a `DataLoader`.
# ### Load the [data](http://pytorch.org/docs/master/torchvision/datasets.html)
#
# #### Dataset class and Tensors
#
# ``torch.utils.data.Dataset`` is an abstract class representing a
# dataset. The FashionMNIST class is an extension of this Dataset class and it allows us to 1. load batches of image/label data, and 2. uniformly apply transformations to our data, such as turning all our images into Tensor's for training a neural network. *Tensors are similar to numpy arrays, but can also be used on a GPU to accelerate computing.*
#
# Let's see how to construct a training dataset.
#
# +
# our basic libraries
import torch
import torchvision
# data loading and transforming
from torchvision.datasets import FashionMNIST
from torch.utils.data import DataLoader
from torchvision import transforms
# The output of torchvision datasets are PILImage images of range [0, 1].
# We transform them to Tensors for input into a CNN
## Define a transform to read the data in as a tensor
data_transform = transforms.ToTensor()
# choose the training and test datasets
train_data = FashionMNIST(root='./data', train=True,
download=False, transform=data_transform)
# Print out some stats about the training data
print('Train data, number of images: ', len(train_data))
# -
# #### Data iteration and batching
#
# Next, we'll use ``torch.utils.data.DataLoader`` , which is an iterator that allows us to batch and shuffle the data.
#
# In the next cell, we shuffle the data and load in image/label data in batches of size 20.
# +
# prepare data loaders, set the batch_size
## TODO: you can try changing the batch_size to be larger or smaller
## when you get to training your network, see how batch_size affects the loss
batch_size = 20
train_loader = DataLoader(train_data, batch_size=batch_size, shuffle=True)
# specify the image classes
classes = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# -
# ### Visualize some training data
#
# This cell iterates over the training dataset, loading a random batch of image/label data, using `dataiter.next()`. It then plots the batch of images and labels in a `2 x batch_size/2` grid.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(batch_size):
ax = fig.add_subplot(2, batch_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title(classes[labels[idx]])
# -
# ### View an image in more detail
#
# Each image in this dataset is a `28x28` pixel, normalized, grayscale image.
#
# #### A note on normalization
#
# Normalization ensures that, as we go through a feedforward and then backpropagation step in training our CNN, that each image feature will fall within a similar range of values and not overly activate any particular layer in our network. During the feedfoward step, a network takes in an input image and multiplies each input pixel by some convolutional filter weights (and adds biases!), then it applies some activation and pooling functions. Without normalization, it's much more likely that the calculated gradients in the backpropagaton step will be quite large and cause our loss to increase instead of converge.
#
# +
# select an image by index
idx = 2
img = np.squeeze(images[idx])
# display the pixel values in that image
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
# -
| nn-review/Load and Visualize FashionMNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: html
# language: python
# name: html
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
from sklearn.datasets import load_diabetes
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split, GridSearchCV
# +
# if you want more information about the dataset for this demo:
# scikit-learn dataset
# https://scikit-learn.org/stable/datasets/toy_dataset.html#diabetes-dataset
# in short, regression problem, trying to predict disease progression based on
# a bunch of variables
# load dataset
diabetes_X, diabetes_y = load_diabetes(return_X_y=True)
X = pd.DataFrame(diabetes_X)
y = diabetes_y
X.head()
# +
# random forests
rf_model = RandomForestRegressor(
n_estimators=100, max_depth=3, random_state=0, n_jobs=4)
# hyperparameter space
rf_param_grid = dict(
n_estimators=[2, 5, 10, 20, 50, 100, 200, 300],
# max_depth=[1, 2, 3, 4],
# min_samples_split=[2,4,8,12,24]
)
# search
reg = GridSearchCV(rf_model, rf_param_grid,
scoring='r2', cv=5)
search = reg.fit(X, y)
# best hyperparameters
search.best_params_
# +
# plot the resuls
results = pd.DataFrame(search.cv_results_)[['params', 'mean_test_score', 'std_test_score']]
results.index = rf_param_grid['n_estimators']
results['mean_test_score'].plot(yerr=[results['std_test_score'], results['std_test_score']], subplots=True)
plt.ylim(0.2, 0.6)
plt.ylabel('Mean r2 score')
plt.xlabel('Number of trees')
# +
# random forests
rf_model = RandomForestRegressor(
n_estimators=200, max_depth=3, random_state=0, n_jobs=4)
# hyperparameter space
rf_param_grid = dict(
# n_estimators=[2, 5, 10, 20, 50, 100, 200],
max_depth=[1, 2, 3, 4, 5, 6],
# min_samples_split=[2,4,8,12,24]
)
# search
reg = GridSearchCV(rf_model, rf_param_grid,
scoring='r2', cv=5)
search = reg.fit(X, y)
# best hyperparameters
search.best_params_
# +
results = pd.DataFrame(search.cv_results_)[['params', 'mean_test_score', 'std_test_score']]
results.index = rf_param_grid['max_depth']
results['mean_test_score'].plot(yerr=[results['std_test_score'], results['std_test_score']], subplots=True)
plt.ylim(0.2, 0.6)
plt.ylabel('Mean r2 score')
plt.xlabel('Tree depth')
# +
# random forests
rf_model = RandomForestRegressor(
n_estimators=200, max_depth=3, random_state=0, n_jobs=4)
# hyperparameter space
rf_param_grid = dict(
# n_estimators=[2, 5, 10, 20, 50, 100, 200],
# max_depth=[1, 2, 3, 4, 5, 6],
min_samples_split=[2,4,8,12,24,48]
)
# search
reg = GridSearchCV(rf_model, rf_param_grid,
scoring='r2', cv=5)
search = reg.fit(X, y)
# best hyperparameters
search.best_params_
# +
results = pd.DataFrame(search.cv_results_)[['params', 'mean_test_score', 'std_test_score']]
results.index = rf_param_grid['min_samples_split']
results['mean_test_score'].plot(yerr=[results['std_test_score'], results['std_test_score']], subplots=True)
plt.ylim(0.2, 0.6)
plt.ylabel('Mean r2 score')
plt.xlabel('Minimum samples at split')
# -
| Section-02-Hyperparamter-Overview/02-02-Low-Effective-Dimension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Phonetic similarity experiment
#
# This notebook contains the code necessary to compare the phonetic similarity figures in [Vitz and Winkler (1973)](https://www.researchgate.net/publication/232418589_Predicting_the_Judged_Similarity_of_Sound_of_English_words) to the cosine similarity obtained between items in the vector embedding described in my paper. CSV files with the experimental data are included with this repository. There is an additional experiment in Vitz and Winkler that I didn't include (haven't had time to transcribe the data yet!).
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# The `adjustText` module adds the nice arrows from the labels to the points in the scatterplots. Recommended!
from adjustText import adjust_text
# The cosine similarity function...
# +
from numpy import dot
from numpy.linalg import norm
# cosine similarity
def cosine(v1, v2):
if norm(v1) > 0 and norm(v2) > 0:
return dot(v1, v2) / (norm(v1) * norm(v2))
else:
return 0.0
# -
# The "space" is defined as a dictionary, whose keys are the words and whose values are the vectors. These are parsed from a pre-computed set of vectors that were hard-coded to include the nonce words used in Vitz and Winkler.
space = dict()
for line in open("cmudict-0.7b-simvecs-vitz", encoding="latin1"):
line = line.strip()
word, vec_raw = line.split(" ")
word = word.lower()
space[word] = np.array([float(x) for x in vec_raw.split()])
# Just to make sure everything loaded correctly:
space["cheese"]
# The `runexperiment()` function takes a CSV file with data from the Vitz and Winkler experiment, along with a vector space (as loaded above) and a "standard word" (i.e., the word against which phonetic similarity is being tested). It returns a Pandas dataframe.
def runexperiment(csv_filename, space, exp_word):
df = pd.read_csv(csv_filename)
df["embedding_cosine"] = [cosine(space[exp_word], space[x]) for x in df["word"]]
df["vw_predicted"] = [1-x for x in df["vw_predicted"]]
df.sort_values(by="embedding_cosine")
return df
# The `getplot()` function takes a dataframe as returned by `runexperiment` and plots it as a scatterplot. Note: The labels need to be changed manually in the function call (see the implementation below for details). To save the plot as a PDF, specify the `fname` parameter with the desired filename.
def getplot(df, x="embedding_cosine", y="obtained", labelx="Vector cosine similarity",
labely="Obtained (Vitz and Winkler)",
title="Standard word: sit",
fname=None):
plt.figure(figsize=(6, 6), dpi=75) # change DPI here for print-ready
plt.scatter(df[x], df[y], s=4.0)
texts = []
for i, text in enumerate(df["word"]):
row = df.iloc[i]
texts.append(plt.text(row[x], row[y], row["word"]))
adjust_text(texts, arrowprops=dict(arrowstyle="->", lw=0.5, alpha=0.5))
plt.xlabel(labelx)
plt.ylabel(labely)
plt.title(title)
if fname:
plt.savefig(fname)
plt.show()
# ## Experiment 1: sit
#
# Results from the vector space:
df = runexperiment("./vitz-1973-experiment-sit.csv", space, "sit")
getplot(df, labelx="Cosine similarity", labely="Obtained (Vitz and Winkler)",
title="Standard word: sit")
# Correlation:
np.corrcoef(df["obtained"], df["embedding_cosine"])[0, 1]
# Results from Vitz and Winkler's "PPD" metric ("predicted phonemic distance"):
getplot(df, "vw_predicted", "obtained", labelx="PPD (Vitz and Winkler)",
labely="Obtained (Vitz and Winkler)",
title="Standard word: sit")
# Correlation:
np.corrcoef(df["obtained"], df["vw_predicted"])[0, 1]
# ## Experiment 2: plant
# Vector space results:
df = runexperiment("./vitz-1973-experiment-plant.csv", space, "plant")
getplot(df, labelx="Cosine similarity", labely="Obtained (Vitz and Winkler)",
title="Standard word: plant")
# Correlation:
np.corrcoef(df["obtained"], df["vw_predicted"])[0, 1]
# Vitz and Winkler PPD results:
df = runexperiment("./vitz-1973-experiment-plant.csv", space, "plant")
getplot(df, "vw_predicted", "obtained", labelx="PPD (Vitz and Winkler)",
labely="Obtained (Vitz and Winkler)",
title="Standard word: plant")
# Correlation:
np.corrcoef(df["obtained"], df["embedding_cosine"])[0, 1]
# ## Experiment 3: wonder
# Vector space results:
df = runexperiment("./vitz-1973-experiment-wonder.csv", space, "wonder")
getplot(df, labelx="Cosine similarity", labely="Obtained (Vitz and Winkler)",
title="Standard word: wonder")
# Correlation:
np.corrcoef(df["obtained"], df["embedding_cosine"])[0, 1]
# Vitz and Winkler results:
getplot(df, "vw_predicted", "obtained", labelx="Inverse PPD (Vitz and Winkler)", labely="Obtained (Vitz and Winkler)",
title="Standard word: wonder")
# Correlation:
np.corrcoef(df["obtained"], df["vw_predicted"])[0, 1]
| experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json, operator, os, re, requests, string, pandas as pd
from bs4 import BeautifulSoup
from pandas.io.json import json_normalize
with open('../data/processed/hamilton_data.json', 'r') as f:
haml = json.load(f)
# ## Basestats
for key in haml[0]:
print(key)
for song in haml:
song['act#'] = int(song['act#'])
song['track#'] = int(song['track#'])
haml.sort(key=operator.itemgetter('act#','track#'))
#for song in haml:
# print('Act:', song['act#'], '\tTrack#:', song['track#'], '\tTrack:', song['track'])
basestats = json_normalize(haml)[['act#', 'track#','track','lyrics']]
line_length, word_length = [], []
for song in haml:
line_length.append(len(song['lyrics']))
words_in_song = 0
for line in song['lyrics']:
words_in_song += len(line['tokenized'])
word_length.append(words_in_song)
basestats['#lines'] = line_length
basestats['#words'] = word_length
r = requests.get('https://en.wikipedia.org/w/index.php?title=Hamilton_(album)&oldid=751292662')
soup = BeautifulSoup(r.content, 'html.parser')
tbl1, tbl2 = soup.find_all('table')[3], soup.find_all('table')[4]
rows1, rows2 = tbl1.find_all('tr'), tbl2.find_all('tr')
song_length = []
for row in rows1[:-1]: # Exclude last line, cause it's just a summary
if len(row.find_all('td'))==0:
continue # Exclude empty rows
row_compl = row.find_all('td') # Find all cells
song_length.append(row_compl[3].text) # Get row giving the length of song
for row in rows2[:-1]: # Exclude last line, cause it's just a summary
if len(row.find_all('td'))==0:
continue # Exclude empty rows
row_compl = row.find_all('td') # Find all cells
song_length.append(row_compl[3].text) # Get row giving the length of song
basestats['len_seconds'] = [int(m[0])*60+int(m[2:4]) for m in song_length]
all_speakers = []
speakers_song_ls = []
for song in haml:
speakers_song = []
for line in song['lyrics']:
all_speakers.append(line['speakers'])
speakers_song.append(line['speakers'])
#speakers_song_ls.append(set(speakers_song))
speakers_song_ls.append(set([item for sublist in speakers_song for item in sublist]))
#print(set([item for sublist in speakers_song for item in sublist]))
all_speakers = set([item for sublist in all_speakers for item in sublist])
basestats['speakers'] = speakers_song_ls
num_speakers = []
for spkls in speakers_song_ls:
num_speakers.append(len(spkls))
basestats['#speakers'] = num_speakers
basestats['#words/sec'] = basestats['#words']/basestats['len_seconds']
basestats.drop('lyrics', axis=1, inplace=False).to_csv('../data/processed/basestats.csv', sep=',')
basestats
# ## Count words per cast member
r = requests.get('https://en.wikipedia.org/w/index.php?title=Hamilton_(musical)&oldid=751273119')
soup = BeautifulSoup(r.content, 'html.parser')
cast = soup.find_all('table')[4]
castr = cast.find_all('tr')
main_cast = []
for row in castr:
if len(row.find_all('th'))==0:
continue # Exclude empty rows
row_compl = row.find_all('th') # Find all cells
cur_speaker_row = row_compl[0].text.split(' / ')
for whole_name in cur_speaker_row:
whole_name_list = whole_name.split(' ')
if 'Character' in whole_name_list:
continue
# Women, children, and kings are referred to by first name
elif 'Schuyler' in whole_name_list:
main_cast.append(whole_name_list[0].upper())
elif 'Reynolds' in whole_name_list:
main_cast.append(whole_name_list[0].upper())
elif 'Philip' in whole_name_list:
main_cast.append(whole_name_list[0].upper())
elif 'King' in whole_name_list:
main_cast.append('KING GEORGE'.upper())
else:
main_cast.append(whole_name_list[-1].upper())
# +
all_corpora = dict()
for speaker in main_cast:
all_corpora[speaker] = []
for song in haml:
for line in song['lyrics']:
for speaker in line['speakers']:
if speaker in main_cast:
all_corpora[speaker].append(line['normalized'])
word_c = []
for speaker in main_cast:
all_corpora[speaker] = [item for sublist in all_corpora[speaker] for item in sublist]
print(speaker, ': \t', len(all_corpora[speaker]))
word_c.append(len(all_corpora[speaker]))
word_counts = pd.DataFrame(word_c,index=main_cast,columns=['#words'])
word_counts.to_csv('../data/processed/word_counts.csv', sep=',')
# -
# ## Calculate similarity
def rem_stopw_punct(string_):
# Remove common stopwords
stopwords = ["a", "about", "above", "above", "across", "after", "afterwards", "again", "against", "all",
"almost", "alone", "along", "already", "also", "although", "always", "am", "among", "amongst",
"amoungst", "amount", "an", "and", "another", "any", "anyhow", "anyone", "anything", "anyway",
"anywhere", "are", "around", "as", "at", "back", "be", "became", "because", "become", "becomes",
"becoming", "been", "before", "beforehand", "behind", "being", "below", "beside", "besides",
"between", "beyond", "bill", "both", "bottom", "but", "by", "call", "can", "cannot", "cant", "co",
"con", "could", "couldnt", "cry", "de", "describe", "detail", "do", "done", "down", "due",
"during", "each", "eg", "eight", "either", "eleven", "else", "elsewhere", "empty", "enough",
"etc", "even", "ever", "every", "everyone", "everything", "everywhere", "except", "few",
"fifteen", "fify", "fill", "find", "fire", "first", "five", "for", "former", "formerly", "forty",
"found", "four", "from", "front", "full", "further", "get", "give", "go", "had", "has", "hasnt",
"have", "he", "hence", "her", "here", "hereafter", "hereby", "herein", "hereupon", "hers",
"herself", "him", "himself", "his", "how", "however", "hundred", "i", "ie", "if", "in", "inc",
"indeed", "interest", "into", "is", "it", "its", "itself", "keep", "last", "latter", "latterly",
"least", "less", "ltd", "made", "many", "may", "me", "meanwhile", "might", "mill", "mine", "more",
"moreover", "most", "mostly", "move", "much", "must", "my", "myself", "name", "namely", "neither",
"never", "nevertheless", "next", "nine", "no", "nobody", "none", "noone", "nor", "not", "nothing",
"now", "nowhere", "of", "off", "often", "on", "once", "one", "only", "onto", "or", "other",
"others", "otherwise", "our", "ours", "ourselves", "out", "over", "own", "part", "per", "perhaps",
"please", "put", "rather", "re", "same", "see", "seem", "seemed", "seeming", "seems", "serious",
"several", "she", "should", "show", "side", "since", "sincere", "six", "sixty", "so", "some",
"somehow", "someone", "something", "sometime", "sometimes", "somewhere", "still", "such",
"system", "take", "ten", "than", "that", "the", "their", "them", "themselves", "then", "thence",
"there", "thereafter", "thereby", "therefore", "therein", "thereupon", "these", "they", "thickv",
"thin", "third", "this", "those", "though", "three", "through", "throughout", "thru", "thus",
"to", "together", "too", "top", "toward", "towards", "twelve", "twenty", "two", "un", "under",
"until", "up", "upon", "us", "very", "via", "was", "we", "well", "were", "what", "whatever",
"when", "whence", "whenever", "where", "whereafter", "whereas", "whereby", "wherein", "whereupon",
"wherever", "whether", "which", "while", "whither", "who", "whoever", "whole", "whom", "whose",
"why", "will", "with", "within", "without", "would", "yet", "you", "your", "yours", "yourself",
"yourselves", "the"]
pattern1 = re.compile(r'\b(' + r'|'.join(stopwords) + r')\b\s*')
string_ = pattern1.sub('', string_)
# Remove punctuation, i.e. anything that's not a letter or space
pattern2 = re.compile(r'[^A-Za-z ]')
string_ = pattern2.sub(' ', string_).strip()
# Also, reduce any sequence of 2+ white spaces to a single one, while also stripping trailing blanks
pattern3 = re.compile(r' +')
string_ = pattern3.sub(' ', string_).strip()
return string_.translate(string.punctuation)
# Use other bool sim formula?
def bool_sim(st1, st2, df):
return (int(df[[st1]].multiply(df[st2], axis="index").sum()))/(int(df[[st1]].sum()) * int(df[[st2]].sum()))
#return float((df[[st1]].multiply(df[st2], axis="index")).sum()) / (int(df[[st1]].sum()) * int(df[[st2]].sum()))
### STOPPED HERE
# Have to split it up again, right?
for speaker in all_corpora:
all_corpora[speaker] = rem_stopw_punct(str(' '.join(all_corpora[speaker])).lower())
# +
# Get vocabulary
corp_flat = []
for speaker in all_corpora:
corp_flat.append(all_corpora[speaker].split(' '))
corp_flat = [item for sublist in corp_flat for item in sublist]
voc = set(corp_flat)
# Create term-document matrix
tdm = pd.DataFrame(0,index=voc,columns=main_cast)
for speaker in all_corpora:
tokens = all_corpora[speaker].split(' ')
for word in tokens:
tdm.at[word,speaker] = 1
tdm.to_csv('../data/processed/tdm.csv', sep=',')
# -
bool_sim(main_cast[0], main_cast[2], tdm)
# +
# Do boolean, td-idf similarity
# Create document similarity matrix
dsm = pd.DataFrame(0,index=main_cast,columns=main_cast)
counter = 0
for cast1 in main_cast:
for cast2_index in range(counter,len(main_cast)):
if (main_cast == main_cast[cast2_index]):
dsm.loc[main_cast, main_cast[cast2_index]] = 1
else:
dsm.loc[main_cast, main_cast[cast2_index]] = \
dsm.loc[main_cast[cast2_index], main_cast] = \
bool_sim(main_cast, main_cast[cast2_index], tdm)
counter += 1
dsm.to_csv('../data/processed/dsm.csv', sep=' ')
| notebooks/2.3-j-hamilton-stats-relationships.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PaddlePaddle 2.0.0b0 (Python 3.5)
# language: python
# name: py35-paddle1.2.0
# ---
# # 1. DCGAN改LSGAN
# * DCGAN与LSGAN主要区别在于损失函数
# 可以看下有提示的地方。
#导入一些必要的包
import os
import random
import paddle
import paddle.nn as nn
import paddle.optimizer as optim
import paddle.vision.datasets as dset
import paddle.vision.transforms as transforms
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
# +
dataset = paddle.vision.datasets.MNIST(mode='train',
transform=transforms.Compose([
# resize ->(32,32)
transforms.Resize((32,32)),
# 归一化到-1~1
transforms.Normalize([127.5], [127.5])
]))
dataloader = paddle.io.DataLoader(dataset, batch_size=32,
shuffle=True, num_workers=4)
# +
#参数初始化的模块
@paddle.no_grad()
def normal_(x, mean=0., std=1.):
temp_value = paddle.normal(mean, std, shape=x.shape)
x.set_value(temp_value)
return x
@paddle.no_grad()
def uniform_(x, a=-1., b=1.):
temp_value = paddle.uniform(min=a, max=b, shape=x.shape)
x.set_value(temp_value)
return x
@paddle.no_grad()
def constant_(x, value):
temp_value = paddle.full(x.shape, value, x.dtype)
x.set_value(temp_value)
return x
def weights_init(m):
classname = m.__class__.__name__
if hasattr(m, 'weight') and classname.find('Conv') != -1:
normal_(m.weight, 0.0, 0.02)
elif classname.find('BatchNorm') != -1:
normal_(m.weight, 1.0, 0.02)
constant_(m.bias, 0)
# +
# Generator Code
class Generator(nn.Layer):
def __init__(self, ):
super(Generator, self).__init__()
self.gen = nn.Sequential(
# input is Z, [B, 100, 1, 1] -> [B, 64 * 4, 4, 4]
nn.Conv2DTranspose(100, 64 * 4, 4, 1, 0, bias_attr=False),
nn.BatchNorm2D(64 * 4),
nn.ReLU(True),
# state size. [B, 64 * 4, 4, 4] -> [B, 64 * 2, 8, 8]
nn.Conv2DTranspose(64 * 4, 64 * 2, 4, 2, 1, bias_attr=False),
nn.BatchNorm2D(64 * 2),
nn.ReLU(True),
# state size. [B, 64 * 2, 8, 8] -> [B, 64, 16, 16]
nn.Conv2DTranspose( 64 * 2, 64, 4, 2, 1, bias_attr=False),
nn.BatchNorm2D(64),
nn.ReLU(True),
# state size. [B, 64, 16, 16] -> [B, 1, 32, 32]
nn.Conv2DTranspose( 64, 1, 4, 2, 1, bias_attr=False),
nn.Tanh()
)
def forward(self, x):
return self.gen(x)
netG = Generator()
# Apply the weights_init function to randomly initialize all weights
# to mean=0, stdev=0.2.
netG.apply(weights_init)
# Print the model
print(netG)
# +
class Discriminator(nn.Layer):
def __init__(self,):
super(Discriminator, self).__init__()
self.dis = nn.Sequential(
# input [B, 1, 32, 32] -> [B, 64, 16, 16]
nn.Conv2D(1, 64, 4, 2, 1, bias_attr=False),
nn.LeakyReLU(0.2),
# state size. [B, 64, 16, 16] -> [B, 128, 8, 8]
nn.Conv2D(64, 64 * 2, 4, 2, 1, bias_attr=False),
nn.BatchNorm2D(64 * 2),
nn.LeakyReLU(0.2),
# state size. [B, 128, 8, 8] -> [B, 256, 4, 4]
nn.Conv2D(64 * 2, 64 * 4, 4, 2, 1, bias_attr=False),
nn.BatchNorm2D(64 * 4),
nn.LeakyReLU(0.2),
# state size. [B, 256, 4, 4] -> [B, 1, 1, 1]
nn.Conv2D(64 * 4, 1, 4, 1, 0, bias_attr=False),
# 这里为需要改变的地方
# nn.Sigmoid() # DCGAN
nn.LeakyReLU() # LSGAN
)
def forward(self, x):
return self.dis(x)
netD = Discriminator()
netD.apply(weights_init)
print(netD)
# +
# Initialize BCELoss function
# 这里为需要改变的地方
# loss = nn.BCELoss() # DCGAN
loss=nn.MSELoss() # LSGAN
# Create batch of latent vectors that we will use to visualize
# the progression of the generator
fixed_noise = paddle.randn([32, 100, 1, 1], dtype='float32')
# Establish convention for real and fake labels during training
real_label = 1.
fake_label = 0.
# Setup Adam optimizers for both G and D
optimizerD = optim.Adam(parameters=netD.parameters(), learning_rate=0.0002, beta1=0.5, beta2=0.999)
optimizerG = optim.Adam(parameters=netG.parameters(), learning_rate=0.0002, beta1=0.5, beta2=0.999)
# -
losses = [[], []]
#plt.ion()
now = 0
for pass_id in range(100):
for batch_id, (data, target) in enumerate(dataloader):
############################
# (1) Update D network: maximize log(D(x)) + log(1 - D(G(z)))
###########################
optimizerD.clear_grad()
real_img = data
bs_size = real_img.shape[0]
label = paddle.full((bs_size, 1, 1, 1), real_label, dtype='float32')
real_out = netD(real_img)
errD_real = loss(real_out, label)
errD_real.backward()
noise = paddle.randn([bs_size, 100, 1, 1], 'float32')
fake_img = netG(noise)
label = paddle.full((bs_size, 1, 1, 1), fake_label, dtype='float32')
fake_out = netD(fake_img.detach())
errD_fake = loss(fake_out,label)
errD_fake.backward()
optimizerD.step()
optimizerD.clear_grad()
errD = errD_real + errD_fake
losses[0].append(errD.numpy()[0])
############################
# (2) Update G network: maximize log(D(G(z)))
###########################
optimizerG.clear_grad()
noise = paddle.randn([bs_size, 100, 1, 1],'float32')
fake = netG(noise)
label = paddle.full((bs_size, 1, 1, 1), real_label, dtype=np.float32,)
output = netD(fake)
errG = loss(output,label)
errG.backward()
optimizerG.step()
optimizerG.clear_grad()
losses[1].append(errG.numpy()[0])
############################
# visualize
###########################
if batch_id % 100 == 0:
generated_image = netG(noise).numpy()
imgs = []
plt.figure(figsize=(15,15))
try:
for i in range(10):
image = generated_image[i].transpose()
image = np.where(image > 0, image, 0)
image = image.transpose((1,0,2))
plt.subplot(10, 10, i + 1)
plt.imshow(image[...,0], vmin=-1, vmax=1)
plt.axis('off')
plt.xticks([])
plt.yticks([])
plt.subplots_adjust(wspace=0.1, hspace=0.1)
msg = 'Epoch ID={0} Batch ID={1} \n\n D-Loss={2} G-Loss={3}'.format(pass_id, batch_id, errD.numpy()[0], errG.numpy()[0])
print(msg)
plt.suptitle(msg,fontsize=20)
plt.draw()
plt.savefig('{}/{:04d}_{:04d}.png'.format('work', pass_id, batch_id), bbox_inches='tight')
plt.pause(0.01)
except IOError:
print(IOError)
paddle.save(netG.state_dict(), "work/generator.params")
# 训练最后的结果:
# 
#
# # 2.参考资料
#
# 【PaddleGAN的Github地址】:https://github.com/PaddlePaddle/PaddleGAN
#
# 【PaddleGAN的Gitee地址】:https://gitee.com/PaddlePaddle/PaddleGAN
#
# 【生成对抗网络七日打卡营】课程链接:https://aistudio.baidu.com/aistudio/course/introduce/16651
#
# 【生成对抗网络七日打卡营】项目合集:https://aistudio.baidu.com/aistudio/projectdetail/1807841
#
# 【图像分割7日打卡营常见问题汇总】
# https://aistudio.baidu.com/aistudio/projectdetail/1100155
#
# 【PaddlePaddle使用教程】
# https://www.paddlepaddle.org.cn/documentation/docs/zh/develop/guides/index_cn.html
#
# 【本地安装PaddlePaddle的常见错误】
# https://aistudio.baidu.com/aistudio/projectdetail/697227
#
# 【API文档】
# https://www.paddlepaddle.org.cn/documentation/docs/zh/develop/api/index_cn.html
#
# 【PaddlePaddle/hapi Github】
# https://github.com/PaddlePaddle/hapi
#
# 【Github使用】
# https://guides.github.com/activities/hello-world/
#
#
# # 3.个人介绍
# > 中南大学 机电工程学院 机械工程专业 2019级 研究生 雷钢
#
# > 百度飞桨官方帮帮团成员
#
# > Github地址:https://github.com/leigangblog
#
# > B站:https://space.bilibili.com/53420969
#
# 来AI Studio互关吧,等你哦~ https://aistudio.baidu.com/aistudio/personalcenter/thirdview/118783
# 欢迎大家fork喜欢评论三连,感兴趣的朋友也可互相关注一下啊~
| education/leigangblog/homework2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
"""
@Author: <NAME>
@Date: Arg 8, 2020
Usage Notice:
1. It's just a demo, which I don't recommend to run because of selenium's slow time
(5 - 20 seconds for one website, typically at least 5 seconds, TOO SLOW!!!)
2. I'd like to recommend "fast-crawler-NSTL-data.ipynb" script instead
3. If you have few websites, e.g., 100 - 1000, you can definitely use selenium to capture the
source info (contents) of the website.
4. If you want fast capturing, don't use too many "print()" functions since they take a lot time
5. Download the Chrome Driver first by using the below code
driver_path = ChromeDriverManager().install()
Then, comment this line and change the blow to your Driver's path:
driver_path = '/Users/shuyuej/.wdm/drivers/chromedriver/mac64/84.0.4147.30/chromedriver'
6. I don't recommend you to download the Google Chrome Driver from the website by yourself,
as it might lead to the problem of version compatibility
So, just use "driver_path = ChromeDriverManager().install()"
to download it automatically by the system
Then, you comment it and change the path of the driver
7. If you have new dynamic webs to capture, you can change the string matching rules to fit the needs
"""
# import necessary packages
import os
import requests
import time
import numpy as np
import sys
import re
from selenium import webdriver
from webdriver_manager.chrome import ChromeDriverManager
from selenium.webdriver.common.desired_capabilities import DesiredCapabilities
import xml
import pandas as pd
# -
def read_url(url: str, driver_path: str):
"""
Read the website and return the contents of the website
:param url: The url of the website
:param driver_path: The path of the Google Chrome Driver
:return soup.text: The contents of the website
"""
start_time = time.time()
option = webdriver.ChromeOptions()
option.add_argument(
'user-agent="MQQBrowser/26 Mozilla/5.0 (Linux; U; Android 2.3.7; zh-cn; MB200 Build/GRJ22; CyanogenMod-7)'
'AppleWebKit/533.1 (KHTML, like Gecko) Version/4.0 Mobile Safari/533.1"')
option.add_argument(
'user-agent="Mozilla/5.0 (iPhone; CPU iPhone OS 9_1 like Mac OS X) AppleWebKit/601.1.46 (KHTML, like Gecko)'
'Version/9.0 Mobile/13B143 Safari/601.1"')
option.add_argument('--disable-infobars')
option.add_argument('--incognito')
option.add_argument('headless')
option.add_argument('blink-settings=imagesEnabled=false')
prefs = {
"profile.managed_default_content_settings.images": 2,
'profile.default_content_settings.popups': 0
}
option.add_experimental_option("prefs", prefs)
# If you don't have Google Chrome Driver installed, uncomment this line
# driver_path = ChromeDriverManager().install()
try:
driver = webdriver.Chrome(driver_path, chrome_options=option)
except Exception as e:
driver_path = ChromeDriverManager().install()
print('You have successfully installed the Google Chrome Driver!')
print('Now change the path of driver to your own path in your laptop!')
driver = webdriver.Chrome(driver_path, chrome_options=option)
driver.get(url)
contents = driver.page_source
START = contents.find('serverContent')
END = contents.find('QRcodebox')
contents_cut = contents[START:END]
end_time = time.time()
print('Time used for get the website was %7f' % (end_time - start_time))
return contents, contents_cut, driver
def find_English_term(content: str):
"""
Find the English Term from the contents
:param content: The contents of the website
:return Eng_term: The found English term
:return content: The contents that cut the English term part
"""
mark = content.find('detail_content')
temp_cont = content[mark-100:mark]
START = temp_cont.find('">')
END = temp_cont.find('</a></h3>')
Eng_term = temp_cont[START+2:END]
content = content[mark+len('detail_content'):]
return Eng_term, content
def find_Chinese_term(content: str):
"""
Find the Chinese Term from the contents
:param content: The contents of the website
:return Chi_term: The found Chinese Term
:return content: The contents that cut the Chinese term part
"""
if '中文名称' not in content:
Chi_term = ''
else:
mark = content.find('target')
temp_cont = content[mark:mark+100]
START = temp_cont.find('target')
END = temp_cont.find('</a>')
Chi_term = temp_cont[START+len('target="_blank">'):END]
chi_loc = content.find(Chi_term)
content = content[chi_loc+len(Chi_term):]
return Chi_term, content
def find_English_definition(content: str):
"""
Find the English Definition from the content
:param content: The contents of the website
:return Eng_def: The found English definition
:return content: The contents that cut the English definition part
"""
if '释义' not in content:
Eng_def = ''
else:
START = content.find('释义')
END = content.find('</i>')
Eng_def = content[START+len('释义:<span><i>'):END]
content = content[END+len('</i></span></div>'):]
return Eng_def, content
def synonym(content: str):
"""
Find all the Synonym words w.r.t. the English term
:param content: The contents of the website
:return synonym_words: The found synonym words
"""
if '同义词' not in content:
synonym_words = ''
else:
START = content.find('同义词')
END = content.find('范畴')
main_content = content[START:END]
key_word = 'target'
synonym_words = []
cur_content = main_content
while key_word in cur_content:
start = cur_content.find('target') + len('target')
ite_content = cur_content[start:start+100]
new_start = ite_content.find(">")
end = ite_content.find('</a></span>')
synonym_word = ite_content[new_start+1:end]
synonym_words.append(synonym_word)
cur_content = cur_content[start+1:]
synonym_words = np.array(synonym_words)
synonym_words = np.squeeze(synonym_words)
synonym_words = str(synonym_words).replace('[', '')
synonym_words = [str(synonym_words).replace(']', '')]
content = content[END:]
return synonym_words, content
def field(content: str):
"""
Find and save all the Fields of this particular term
:param content: The contents of the website
:return content: The Fields contents
"""
if '范畴' not in content:
field = ''
else:
content.replace("title=""", '')
START = content.find('target') + len('target')
content = content[START:]
field = []
new_content = content
while 'title' in new_content:
start = new_content.find('title=') + len('title=')
end = new_content.find('><span')
temp_field = new_content[start+1:end-1]
if temp_field != '':
field.append(temp_field)
new_content = new_content[start:]
field = np.array(field)
field = np.squeeze(field)
field = str(field).replace('[', '')
field = [str(field).replace(']', '')]
return field
# The main function
if __name__ == "__main__":
# Set an index to observe the final number of captured webs
index = 0
# Initialize the saved contents
English_terms = []
Chinese_terms = []
English_definition = []
Synonym_words = []
Fileds_summary = []
# If you want to capture 0 - 100 webs
start = '0'
end = '100'
save_file = start + '-' + end
# Change this if you want to capture more websites
start_index = int(0)
end_index = int(100)
for i in range(start_index, end_index):
if i < 10:
i = '00000' + str(i)
elif 10 <= i < 100:
i = '0000' + str(i)
elif 100 <= i < 1000:
i = '000' + str(i)
elif 1000 <= i < 10000:
i = '00' + str(i)
elif 10000 <= i < 100000:
i = '0' + str(i)
else:
i = str(i)
url = 'https://www.nstl.gov.cn/stkos_detail.html?id=C019' + i
driver_path = '/Users/shuyuej/.wdm/drivers/chromedriver/mac64/84.0.4147.30/chromedriver'
save_path = 'NSTD_data/'
if not os.path.exists(save_path):
os.mkdir(save_path)
# Read the URL and see whether there was contents
contents, contents_cut, driver = read_url(url=url, driver_path=driver_path)
# If there was no contents in this website, skip and continue
if '暂无相关资源' in contents:
print('There is no data in this webpage! Skip and continue......')
continue
else:
Eng_term, con_cut_eng = find_English_term(content=contents_cut)
English_terms.append(Eng_term)
Chi_term, con_cut_chi = find_Chinese_term(content=con_cut_eng)
Chinese_terms.append(Chi_term)
Eng_def, con_cut_def = find_English_definition(content=con_cut_chi)
English_definition.append(Eng_def)
synonym_word, synonym_cut_con = synonym(content=con_cut_def)
Synonym_words.append(synonym_word)
fields = field(content=synonym_cut_con)
Fileds_summary.append(fields)
index += 1
print('It\'s ' + str(i) + ' Website, saved its data, and continue......')
rows = np.shape(English_terms)[0]
English_terms = np.reshape(English_terms, [rows, 1])
Chinese_terms = np.reshape(Chinese_terms, [rows, 1])
English_definition = np.reshape(English_definition, [rows, 1])
Synonym_words = np.reshape(Synonym_words, [rows, 1])
Fileds_summary = np.reshape(Fileds_summary, [rows, 1])
save_data = np.concatenate([English_terms, Chinese_terms, English_definition, Synonym_words, Fileds_summary], axis=1)
save_data = pd.DataFrame(save_data)
save_data.to_csv(save_path + '%s.csv' % save_file, sep=',', index=False, header=['English Term', 'Chinese Term', 'English Definition', 'Synonym', 'Field'])
driver.close()
print('Cheers! %s\'s NSTL data (%s terms) has been successfully saved!' % (save_file, str(index)))
driver.quit()
| selenium-crawler-NSTL-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: python
# ---
# # Creating and grading assignments
# This guide walks an instructor through the workflow for generating an assignment and preparing it for release to students.
# + active=""
# .. contents:: Table of Contents
# :depth: 2
# + active=""
# .. versionadded:: 0.5.0
#
# Much of the core functionality of nbgrader can now be accessed through the "formgrader" extension.
# -
# ## Accessing the formgrader extension
# + active=""
# .. seealso::
#
# :doc:`installation`
# Instructions on how to install the formgrader extension.
# -
# The formgrader extension provides the core access to nbgrader's instructor tools. After the extension has been installed, you can access it through the tab in the notebook list:
#
# 
# ## Creating a new assignment
# + active=""
# .. seealso::
#
# :doc:`managing_the_database`
# Instructions on how to manage assignments in the database from the command line
#
# :doc:`/command_line_tools/nbgrader-db-assignment-add`
# Command line options for ``nbgrader db assignment add``
# -
# ### From the formgrader
# To create a new assignment, open the formgrader extension and click the "Add new assignment..." button at the bottom of the page. This will ask you to provide some information such as the name of the assignment and its due date. Then, you can add files to the assignment and edit them by clicking the name of the assignment:
#
# 
# ### From the command line
# + active=""
# If you are not using the formgrader extension, you can add a new assignment simply by creating a folder in your course directory with the name of the assignment. You can specify the assignment metadata (such as the duedate) using the `nbgrader db assignment` command (see :doc:`managing_the_database`).
# -
# To simplify this example, two notebooks of the assignment have already been stored in the `source/ps1` folder:
#
# * [source/ps1/problem1.ipynb](source/ps1/problem1.ipynb)
# * [source/ps1/problem2.ipynb](source/ps2/problem2.ipynb)
# ## Developing assignments with the assignment toolbar
# **Note**: As you are developing your assignments, you should save them
# into the `source/{assignment_id}/` folder of the nbgrader hierarchy,
# where `assignment_id` is the name of the assignment you are creating
# (e.g. "ps1").
# + active=""
# .. seealso::
#
# :doc:`philosophy`
# More details on how the nbgrader hierarchy is structured.
#
# :doc:`/configuration/student_version`
# Instructions for customizing how the student version of the assignment looks.
#
# Before you can begin developing assignments, you will need to actually
# install the nbgrader toolbar. If you do not have it installed, please
# first follow the :doc:`installation instructions <installation>`.
# -
# Once the toolbar has been installed, you should see it in the drop down "View -> Cell Toolbar" menu:
#
# 
#
# Selecting the "Create Assignment" toolbar will create a separate toolbar
# for each cell which by default will be a dropdown menu with the "-" item
# selected. For markdown cells, there are two additional options to choose
# from, either "Manually graded answer" or "Read-only":
#
# 
#
# For code cells, there are four options to choose from, including
# "Manually graded answer", "Autograded answer", "Autograder tests", and
# "Read-only":
#
# 
#
# The following sections go into detail about the different cell types,
# and show cells that are taken from a complete example of an assignment
# generated with the nbgrader toolbar extension:
#
# - [source/ps1/problem1.ipynb](source/ps1/problem1.html)
# - [source/ps1/problem2.ipynb](source/ps1/problem2.html)
# + active=""
# .. _manually-graded-cells:
# -
# ### "Manually graded answer" cells
# If you select the "Manually graded answer" option (available for both
# markdown and code cells), the nbgrader extension will mark that cell as
# a cell that contains an answer that must be manually graded by a human
# grader. Here is an example of a manually graded answer cell:
#
# 
#
# The most common use case for this type of cell is for written
# free-response answers (for example, which interpret the results of code
# that may have been written and/or executed above).
# + active=""
# When you specify a manually graded answer, you must additionally tell nbgrader how many points the answer is worth, and an id for the cell. Additionally, when creating the release version of the assignment (see :ref:`assign-and-release-an-assignment`), the bodies of answer cells will be replaced with a code or text stub indicating to the students that they should put their answer or solution there. Please see :doc:`/configuration/student_version` for details on how to customize this behavior.
# -
# *Note: the blue border only shows up when the nbgrader extension toolbar
# is active; it will not be visible to students.*
# + active=""
# .. _manually-graded-task-cells:
# -
# ### “Manually graded task” cells
# + active=""
# .. versionadded:: 0.6.0
# -
# If you select the “Manually graded task” option (available for markdown cells),
# the nbgrader extension will mark that cell as
# a cell that contains the description of a task that students have to perform.
# They must be manually graded by a human
# grader. Here is an example of a manually graded answer cell:
#
# 
#
# The difference with a manually graded answer is that the manually graded tasks cells are not edited by the student. A manually or automatically graded cell ask students to perform a task *in* one cell. A manually graded task asks students to perform a task *with* cells.
#
# The common use case for this type of cell is for tasks that require the
# student to create several cells such as "Process the data and create a plot to illustrate your results."
# or to contain notebook-wide tasks such as "adhere to the PEP8 style convention."
#
# *Note: the blue border only shows up when the nbgrader extension toolbar
# is active; it will not be visible to students.*
# + active=""
# .. _manually-graded-task-cell-mark-scheme:
# -
# ### “Manually graded task” cells with mark scheme
# + active=""
# .. versionadded:: 0.6.0
# -
# A mark scheme can be created through the use of a
# special syntax such as ``=== BEGIN MARK SCHEME ===`` and
# ``=== END MARK SCHEME ===``. The section of text between the two markers will be removed from the student version,
# but will be visible at the grading stage and in the feedback.
# + active=""
# .. _autograded-answer-cells:
# -
# ### "Autograded answer" cells
# If you select the "Autograded answer" option (available only for code
# cells), the nbgrader extension will mark that cell as a cell that
# contains an answer which will be autograded. Here is an example of an
# autograded graded answer cell:
#
# 
# + active=""
# As shown in the image above, solutions can be specified inline, through the use of a special syntax such as ``### BEGIN SOLUTION`` and ``### END SOLUTION``. When creating the release version (see :ref:`assign-and-release-an-assignment`), the region between the special syntax lines will be replaced with a code stub. If this special syntax is not used, then the entire contents of the cell will be replaced with the code stub. Please see :doc:`/configuration/student_version` for details on how to customize this behavior.
# -
# Unlike manually graded answers, autograded answers aren't worth any
# points: instead, the points for autograded answers are specified for the
# particular tests that grade those answers. See the next section for
# further details.
#
# *Note: the blue border only shows up when the nbgrader extension toolbar
# is active; it will not be visible to students.*
# ### "Autograder tests" cells
# If you select the "Autograder tests" option (available only for code
# cells), the nbgrader extension will mark that cell as a cell that
# contains tests to be run during autograding. Here is an example of two
# test cells:
#
# 
# + active=""
# Test cells should contain ``assert`` statements (or similar). When run through ``nbgrader autograde`` (see :ref:`autograde-assignments`), the cell will pass if no errors are raised, and fail otherwise. You must specify the number of points that each test cell is worth; then, if the tests pass during autograding, students will receive the specified number of points, and otherwise will receive zero points.
# -
# The lock icon on the left side of the cell toolbar indicates that the
# tests are "read-only". See the next section for further details on what
# this means.
# + active=""
# For tips on writing autograder tests, see :ref:`autograding-resources`.
# -
# *Note: the blue border only shows up when the nbgrader extension toolbar
# is active; it will not be visible to students.*
# + active=""
# .. _autograder-tests-cell-hidden-tests:
# -
# ### "Autograder tests" cells with hidden tests
# + active=""
# .. versionadded:: 0.5.0
# -
# Tests in "Autograder tests" cells can be hidden through the use of a special syntax such as ``### BEGIN HIDDEN TESTS`` and ``### END HIDDEN TESTS``, for example:
#
# 
# + active=""
# When creating the release version (see :ref:`assign-and-release-an-assignment`), the region between the special syntax lines will be removed. If this special syntax is not used, then the contents of the cell will remain as is. Please see :doc:`/configuration/student_version` for details on how to customize this behavior.
#
# .. note::
#
# Keep in mind that wrapping all tests (for an "Autograder tests" cell) in this special syntax will remove all these tests in
# the release version and the students will only see a blank cell. It is recommended to have at least one or more visible
# tests, or a comment in the cell for the students to see.
#
# These hidden tests are placed back into the "Autograder tests" cells when running ``nbgrader autograde``
# (see :ref:`autograde-assignments`).
# + active=""
# .. _read-only-cells:
# -
# ### "Read-only" cells
# If you select the "Read-only" option (available for both code and
# markdown cells), the nbgrader extension will mark that cell as one that
# cannot be modified. This is indicated by a lock icon on the left side of
# the cell toolbar:
#
# 
# + active=""
# However, this doesn't actually mean that it is truly read-only when opened in the notebook. Instead, what it means is that during the ``nbgrader generate_assignment`` step (see :ref:`assign-and-release-an-assignment`), the source of these cells will be recorded into the database. Then, during the ``nbgrader autograde`` step (see :ref:`autograde-assignments`), nbgrader will check whether the source of the student's version of the cell has changed. If it has, it will replace the cell's source with the version in the database, thus effectively overwriting any changes the student made.
#
# .. versionadded:: 0.4.0
# Read-only cells (and test cells) are now truly read-only! However, at the moment this functionality will only work on the master version of the notebook (5.0.0.dev).
# -
# This functionality is particularly important for test cells, which are
# always marked as read-only. Because the mechanism for autograding is
# that students receive full credit if the tests pass, an easy way to get
# around this would be to simply delete or comment out the tests. This
# read-only functionality will reverse any such changes made by the
# student.
# ## Validating the instructor version
# + active=""
# .. seealso::
#
# :doc:`/command_line_tools/nbgrader-validate`
# Command line options for ``nbgrader validate``
# -
# ### From the validate extension
# Ideally, the solutions in the instructor version should be correct and pass all the test cases to ensure that you are giving your students tests that they can actually pass. To verify this is the case, you can use the validate extension:
#
# 
#
# If your assignment passes all the tests, you'll get a success pop-up:
#
# 
#
# If it doesn't pass all the tests, you'll get a message telling you which cells failed:
#
# 
# ### From the command line
# You can also validate assignments on the command line using the `nbgrader validate` command:
# + language="bash"
#
# nbgrader validate source/ps1/*.ipynb
# + active=""
# .. _assign-and-release-an-assignment:
# -
# ## Generate and release an assignment
# + active=""
# .. seealso::
#
# :doc:`/command_line_tools/nbgrader-generate-assignment`
# Command line options for ``nbgrader generate_assignment``
#
# :doc:`philosophy`
# Details about how the directory hierarchy is structured
#
# :doc:`/configuration/config_options`
# Details on ``nbgrader_config.py``
# -
# ### From the formgrader
# After an assignment has been created with the assignment toolbar, you will want to generate the version that students will receive. You can do this from the formgrader by clicking the "generate" button:
#
# 
#
# This should succeed with a pop-up window containing log output:
#
# 
# ### From the command line
# + active=""
# As described in :doc:`philosophy`, you need to organize your files in a particular way. For releasing assignments, you should have the master copy of your files saved (by default) in the following source directory structure:
# -
# ```
# {course_directory}/source/{assignment_id}/{notebook_id}.ipynb
# ```
#
# Note: The `student_id` is not included here because the source and release versions of the assignment are the same for all students.
#
# After running `nbgrader generate_assignment`, the release version of the notebooks will be:
#
# ```
# {course_directory}/release/{assignment_id}/{notebook_id}.ipynb
# ```
#
# As a reminder, the instructor is responsible for distributing this release version to their students using their institution's existing student communication and document distribution infrastructure.
# When running `nbgrader generate_assignment`, the assignment name (which is "ps1") is passed. We also specify a *header* notebook (`source/header.ipynb`) to prepend at the beginning of each notebook in the assignment. By default, this command should be run from the root of the course directory:
# + language="bash"
#
# nbgrader generate_assignment "ps1" --IncludeHeaderFooter.header=source/header.ipynb --force
# -
# ## Preview the student version
# After generating the student version of assignment, you should preview it to make sure that it looks correct. You can do this from the formgrader extension by clicking the "preview" button:
#
# 
# Under the hood, there will be a new folder called `release` with the same structure as `source`. The `release` folder contains the actual release version of the assignment files:
#
# * [release/ps1/problem1.ipynb](release/ps1/problem1.ipynb)
# * [release/ps1/problem2.ipynb](release/ps1/problem2.ipynb)
# If you are working on the command line, you may want to formally verify the student version as well. Ideally, all the tests should fail in the student version if the student hasn't implemented anything. To verify that this is in fact the case, we can use the `nbgrader validate --invert` command:
# + language="bash"
#
# nbgrader validate --invert release/ps1/*.ipynb
# -
# If the notebook fails all the test cases, you should see the message "Success! The notebook does not pass any tests."
# ## Releasing files to students and collecting submissions
# + active=""
# .. seealso::
#
# :doc:`managing_assignment_files`
# Guide to releasing and collecting submissions.
#
# :doc:`/command_line_tools/nbgrader-release-assignment`
# Command line options for ``nbgrader release_assignment``
#
# :doc:`/command_line_tools/nbgrader-collect`
# Command line options for ``nbgrader collect``
#
# :doc:`philosophy`
# Details about how the directory hierarchy is structured
#
# :doc:`/configuration/config_options`
# Details on ``nbgrader_config.py``
# + active=""
# Note: the :doc:`Managing Assignment Files Guide <managing_assignment_files>` goes into greater depth on how to release and collect assignments, and the various options that are available to do you for doing so.
# + active=""
# At this point you will be able to take the files in the ``release`` folder and distribute them to students. If you are using nbgrader with JupyterHub, you can do this with either with the formgrader extension or with the ``nbgrader release_assignment`` command (see :doc:`managing_assignment_files`). Otherwise, you will need to do this manually.
#
# Similarly, to collect submissions, you can do this either with the formgrader extension or with the ``nbgrader collect`` command. Otherwise, you will need to manually place submitted files into the ``submitted`` directory. As described in :doc:`philosophy`, you need to organize your files in a particular way. For submitted assignments, you should have the submitted versions of students' assignments organized as follows:
# -
# ```
# submitted/{student_id}/{assignment_id}/{notebook_id}.ipynb
# ```
# **Please note**: Students must use version 3 or greater of the IPython/Jupyter notebook for nbgrader to work properly. If they are not using version 3 or greater, it is possible for them to delete cells that contain important metadata for nbgrader. With version 3 or greater, there is a feature in the notebook that prevents cells from being deleted. See [this issue](https://github.com/jupyter/nbgrader/issues/424) for more details.
#
# To ensure that students have a recent enough version of the notebook, you can include a cell such as the following in each notebook of the assignment:
#
# ```python
# import IPython
# assert IPython.version_info[0] >= 3, "Your version of IPython is too old, please update it."
# ```
# + active=""
# .. _autograde-assignments:
# -
# ## Autograde assignments
# + active=""
# .. seealso::
#
# :doc:`/command_line_tools/nbgrader-autograde`
# Command line options for ``nbgrader autograde``
#
# :doc:`philosophy`
# Details about how the directory hierarchy is structured
#
# :doc:`/configuration/config_options`
# Details on ``nbgrader_config.py``
# -
# In the following example, we have an assignment with two notebooks. There are two submissions of the assignment:
#
# Submission 1:
#
# * [submitted/bitdiddle/ps1/problem1.ipynb](submitted/bitdiddle/ps1/problem1.ipynb)
# * [submitted/bitdiddle/ps1/problem2.ipynb](submitted/bitdiddle/ps1/problem2.ipynb)
#
# Submission 2:
#
# * [submitted/hacker/ps1/problem1.ipynb](submitted/hacker/ps1/problem1.ipynb)
# * [submitted/hacker/ps1/problem2.ipynb](submitted/hacker/ps1/problem2.ipynb)
# ### From the formgrader
# You can autograde individual submissions from the formgrader directly. To do so, click on the the number of submissions in the "Manage Assignments" view:
#
# 
#
# This will take you to a new page where you can see all the submissions. For a particular submission, click the "autograde" button to autograde it:
#
# 
#
# After autograding completes, you will see a pop-up window with log output:
#
# 
#
# And back on the submissions screen, you will see that the status of the submission has changed to "needs manual grading" and there is now a reported score as well:
#
# 
# ### From the command line
# We can run the autograder for all students at once from the command line:
# + language="bash"
#
# nbgrader autograde "ps1" --force
# -
# When grading the submission for `Bitdiddle`, you'll see some warnings that look like "Checksum for grade cell correct_squares has changed!". What's happening here is that nbgrader has recorded what the *original* contents of the grade cell `correct_squares` (when `nbgrader generate_assignment` was run), and is checking the submitted version against this original version. It has found that the submitted version changed (perhaps this student tried to cheat by commenting out the failing tests), and has therefore overwritten the submitted version of the tests with the original version of the tests.
#
# You may also notice that there is a note saying "ps1 for Bitdiddle is 21503.948203 seconds late". What is happening here is that nbgrader is detecting a file in Bitdiddle's submission called `timestamp.txt`, reading in that timestamp, and saving it into the database. From there, it can compare the timestamp to the duedate of the problem set, and compute whether the submission is at all late.
#
# Once the autograding is complete, there will be new directories for the autograded versions of the submissions:
#
# ```
# autograded/{student_id}/{assignment_id}/{notebook_id}.ipynb
# ```
#
# Autograded submission 1:
#
# * [autograded/bitdiddle/ps1/problem1.ipynb](autograded/bitdiddle/ps1/problem1.ipynb)
# * [autograded/bitdiddle/ps1/problem2.ipynb](autograded/bitdiddle/ps1/problem2.ipynb)
#
# Autograded submission 2:
#
# * [autograded/hacker/ps1/problem1.ipynb](autograded/hacker/ps1/problem1.ipynb)
# * [autograded/hacker/ps1/problem2.ipynb](autograded/hacker/ps1/problem2.ipynb)
# ## Manual grading
# + active=""
# .. seealso::
#
# :doc:`philosophy`
# More details on how the nbgrader hierarchy is structured.
#
# :doc:`/configuration/config_options`
# Details on ``nbgrader_config.py``
# + active=""
# After assignments have been autograded, they will saved into an ``autograded`` directory (see :doc:`philosophy` for details):
# -
# After running `nbgrader autograde`, the autograded version of the
# notebooks will be:
#
# autograded/{student_id}/{assignment_id}/{notebook_id}.ipynb
#
# We can manually grade assignments through the formgrader as well, by clicking on the "Manual Grading" navigation button. This will provide you with an interface for hand grading assignments that it finds in the directory listed above. Note that this applies to *all* assignments as well -- as long as the autograder has been run on the assignment, it will be available for manual grading via the formgrader.
# ## Generate feedback on assignments
# + active=""
# .. seealso::
#
# :doc:`/command_line_tools/nbgrader-generate-feedback`
# Command line options for ``nbgrader generate_feedback``
#
# :doc:`/command_line_tools/nbgrader-release-feedback`
# Command line options for ``nbgrader release_feedback``
#
# :doc:`philosophy`
# Details about how the directory hierarchy is structured
#
# :doc:`/configuration/config_options`
# Details on ``nbgrader_config.py``
# + active=""
# As mentioned before, after assignments have been autograded and/or manually graded, they will located in the `autograded` directory (see :doc:`philosophy` for details):
# -
# ```
# autograded/{student_id}/{assignment_id}/{notebook_id}.ipynb
# ```
#
# Creating feedback for students is divided into two parts:
#
# * generate feedback
# * release feedback
#
# Generating feedback will create HTML files in the local instructor directory. Releasing feedback will copy those HTML files to the nbgrader exchange.
#
# We can generate feedback based on the graded notebooks by running the `nbgrader generate_feedback` command, which will produce HTML versions of these notebooks at the following location:
#
# ```
# feedback/{student_id}/{assignment_id}/{notebook_id}.html
# ```
#
# The `nbgrader generate_feedback` is available by clicking the Generate Feedback button on either the Manage Assignments view (to generate feedback for all graded submissions), or on the individual student's Manage Submission page (to generate feedback for that specific individual).
# We can release the generated feedback by running the `nbgrader release_feedback` command, which will send the generated HTML files to the nbgrader exchange.
#
# The `nbgrader release_feedback` is available by clicking the Release Feedback button on either the Manage Assignments view (to release feedback for all generated feedback), or on the individual student's Manage Submission page (to release feedback for that specific individual).
# ### Workflow example: Instructor returning feedback to students
# In some scenarios, you may not want to (or be able to) use the exchange to deliver student feedback. This sections describes a workflow for manually returning generated feedback.
#
# In the following example, we have an assignment with two notebooks. There are two submissions of the assignment that have been graded:
#
# Autograded submission 1:
#
# * [autograded/bitdiddle/ps1/problem1.ipynb](autograded/bitdiddle/ps1/problem1.ipynb)
# * [autograded/bitdiddle/ps1/problem2.ipynb](autograded/bitdiddle/ps1/problem2.ipynb)
#
# Autograded submission 2:
#
# * [autograded/hacker/ps1/problem1.ipynb](autograded/hacker/ps1/problem1.ipynb)
# * [autograded/hacker/ps1/problem2.ipynb](autograded/hacker/ps1/problem2.ipynb)
# Generating feedback is fairly straightforward (and as with the other nbgrader commands for instructors, this must be run from the root of the course directory):
# + language="bash"
#
# nbgrader generate_feedback "ps1"
# -
# Once the feedback has been generated, there will be new directories and HTML files corresponding to each notebook in each submission:
#
# Feedback for submission 1:
#
# * [feedback/bitdiddle/ps1/problem1.html](feedback/bitdiddle/ps1/problem1.html)
# * [feedback/bitdiddle/ps1/problem2.html](feedback/bitdiddle/ps1/problem2.html)
#
# Feedback for submission 2:
#
# * [feedback/hacker/ps1/problem1.html](feedback/hacker/ps1/problem1.html)
# * [feedback/hacker/ps1/problem2.html](feedback/hacker/ps1/problem2.html)
#
# If the exchange is available, one would of course use `nbgrader release_feedback`. However if not available, you can now deliver these generated HTML feedback files via whatever mechanism you wish.
# ## Getting grades from the database
# + active=""
# .. versionadded:: 0.4.0
#
# .. seealso::
#
# :doc:`/command_line_tools/nbgrader-export`
# Command line options for ``nbgrader export``
#
# :doc:`/plugins/export-plugin`
# Details on how to write your own custom exporter.
# -
# In addition to creating feedback for the students, you may need to upload grades to whatever learning management system your school uses (e.g. Canvas, Blackboard, etc.). nbgrader provides a way to export grades to CSV out of the box, with the `nbgrader export` command:
# + language="bash"
#
# nbgrader export
# -
# After running `nbgrader export`, you will see the grades in a CSV file called `grades.csv`:
# + language="bash"
#
# cat grades.csv
# + active=""
# If you need to customize how the grades are exported, you can :doc:`write your own exporter </plugins/export-plugin>`.
| nbgrader/docs/source/user_guide/creating_and_grading_assignments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 64-bit
# name: python3
# ---
# # Interpolation
import numpy as np
import matplotlib.pyplot as plt
import math
# ### Linear Interpolation
# Suppose we are given a function $f(x)$ at just two points, $x=a$ and $x=b$, and you want to know the function at another point in between. The simplest way to find an estimate of this value is using linear interpolation. Linear interpolation assumes the function follows a straight line between two points. The slope of the straight line approximate is:
# $$ m = \frac{f(b) - f(a)}{b - a} $$
#
# Then the value $f(x)$ can be approximated by:
# $$ f(x) \approx \frac{f(b) - f(a)}{b-a} (x-a) + f(a) $$
# #### Step 1: Define a linear function
#
# Create a linear function $f(x) = ax + b$. Linear interpolation will yield an accurate answer for a a linear function. This is how we will test our linear interpolation.
def my_function(x):
return (4*x + 5)
pass
# #### Step 2: Implement the linear interpolation
# Using the equations given above, implement the linear interpolation function
def linear_interpolation(x, a, fa, b, fb):
"""
Fits a line to points (a, f(a)) and (b, f(b)) and returns an
approximation for f(x) for some value x between a and b from
the equation of the line.
Parameters:
x (float): the point of interest between a and b
a (float): known x value
fa (float): known f(a) value
b (float): known x value (b > a)
fb (float): known f(b) value
Returns:
(float): an approximation of f(x) using linear interpolation
"""
m = (fb - fa) / (b - a) # transferring equation above into code
return m*(x-a) + fa
# To Do: Implement the linear interpolation function
pass
# #### Step 3: Test your linear interpolation
# Using the linear function you created and your linear interpolation function, write at least three assert statements.
# +
# To DO: Create at least three assert statements using my_function and linear_interpolation
# To Do: Write at least three assert statements
assert(linear_interpolation(2, 3, my_function(3), 4, my_function(4) > 1))
assert(linear_interpolation(2, math.pi, my_function(math.pi), 8, my_function(8) > 1))
assert(linear_interpolation(5, 12, my_function(12), 45, my_function(45)> 1))
# -
# #### Step 4: Visualization your results
# Plot your function. Using a scatter plot, plot at least three x, y points generated using your linear_interpolation function.
x = np.linspace(1, 100, 10)
y = linear_interpolation(x, 1, my_function(1), 7, my_function(7))
plt.plot(x,y)
plt.scatter([1, 7], [my_function(1), my_function(7)])
# ### 2nd Order Lagrangian Interpolation
# If we have more than two points, a better way to get an estimate of "in between" points is using a Lagrangian Interpolation. Lagrangian Interpolation fits a nth order polynomial to a number of points. Higher order polynomials often introduce unnecessary "wiggles" that introduce error. Using many low-order polynomials often generate a better estimate. For this example, let's use a quadratic (i.e. a 2nd order polynomial).
#
# $$f(x) = \frac{(x-b)(x-c)}{(a - b)(a-c)}f(a) + \frac{(x-a)(x-c)}{(b-a)(b-c)}f(b) + \frac{(x - a)(x-b)}{(c - a)(c - b)} f(c) $$
# #### Step 1: Define a quadratic function
#
# Create a quadratic function $f(x) = ax^2 + bx + c$. 2nd Order Lagrangian Interpolation will yield an accurate answer for a 2nd order polynomial (i.e. a quadratic). This is how we will test our interpolation.
def my_function1(x):
#To Do: Create a quadratic function
return 1*(x**2) + 2*x + 3
pass
# #### Step 2: Implement the 2nd Order Lagrangian Interpolation Function
# Using the equations given above, implement the 2nd order lagrangian interpolation function
def lagrangian_interpolation(x, a, fa, b, fb, c, fc):
"""
Fits a quadratic to points (a, f(a)), (b, f(b)), and (c, f(c)) and returns an
approximation for f(x) for some value x between a and c from the
equation of a quadratic.
Parameters:
x (float): the point of interest between a and b
a (float): known x value
fa (float): known f(a) value
b (float): known x value (b > a)
fb (float): known f(b) value
c (float): known x value (c > b)
fc (float): known f(c) value
Returns:
(float): an approximation of f(x) using linear interpolation
"""
return ((x-b)*(x-c))/((a-b)*(a-c))*fa + ((x-a)*(x-c))/((b-a)*(b-c))*fb + ((x-a)*(x-b))/((c-a)*(c-b))*fc #transfers equation above into code
pass
# #### Step 3: Test your results
# Using the quadratic function you created and your 2nd order lagrangian interpolation function, write at least three assert statements.
# +
# To Do: Write at least three assert statements
assert(lagrangian_interpolation(2, 3, my_function(3), 4, my_function(4), 5, my_function(5) > 1))
assert(lagrangian_interpolation(2, 3, my_function(math.pi), 8, my_function(8), 100, my_function(100) > 1))
assert(lagrangian_interpolation(5, 12, my_function(12), 45, my_function(45), 100, my_function(100) > 1))
# -
# #### Step 4: Visualize your results
# Plot your function and using a scatter plot, plot at least three x, y points generated from your lagrangian_interpolation function.
x = np.linspace(1, 100, 100)
y = lagrangian_interpolation(x, 1, my_function1(1), 7, my_function1(7), 13, my_function1(13))
plt.plot(x,y)
plt.scatter([1, 7,13], [my_function1(1), my_function1(7), my_function1(13)])
# ### Application
# Also contained in this file is a text file called `Partial_Data.txt`. This contains sparse data. In this application section we're going to import the data and approximate the curve using linear and 2nd order lagranging interpolation.
# #### Step 1: Import the data
# Take a look at the file and see what data it contains. I suggest using `np.loadtxt` to import this data. Using the argument `unpack = True` will allow you to easily assign each column of data to an individual variable. For more information on the `loadtxt` function and its allowed arguments, see: https://numpy.org/doc/stable/reference/generated/numpy.loadtxt.html
# +
# To Do: Import the data
x_cord,y_cord= np.loadtxt("Partial_Data.txt", unpack = True)
plt.plot(x_cord, y_cord)
len(x)
# To Do: Scatter plot the data
# -
# #### Step 2: Linear Interpolation
# Using your linear interpolation function above, iterate through the sparse data and generate interpolated value.
#
# Here's one method to get you started:
#
# Starting at the 2nd data point, interate through the data, using the current value (let this value be $b$) and the previous data point (let this be $a$ where $b$ > $a$). Interpolate 100 points between the values of ($a, b$) and plot these values. Move onto the next data point and repeat.
# +
x_cordin = []
y_cordin = []
for i in range(1, len(x_cord)):
a = x_cord[i-1]
ay = y_cord[i-1]
b = x_cord[i]
by = y_cord[i]
# gets i+1th point and calculates linear interpolation between ith and (i+1)th point
x_between = np.linspace(a,b, num=100) # iterates 100 times between "a" and "b"
for j in x_between:
x_cordin.append(j)
y_cordin.append(linear_interpolation(j, a, ay, b, by)) # linear interpolation
#plots
plt.scatter(x_cord, y_cord)
plt.plot(x_cordin, y_cordin)
# -
# #### Step 3: 2nd Order Lagrangian Interpolation
# Using your 2nd Order Lagrangian Interpolation function above, iterate through the sparse data and generate interpolated value.
#
# Here's one method to get you started:
#
# Starting at the 3rd data point, interate through the data, using the current value (let this value be $c$) and the previous two (let these be $a$ and $b$ where $b$ > $a$). Interpolate 100 points between the values of ($a, b$) and plot these values. Move onto the next data point and repeat.
# +
# To Do: Generate and plot interpolated data
x_cordin = []
y_cordin = []
for i in range(2, len(x_cord)):
a = x_cord[i-2]
ay = y_cord[i-2]
b = x_cord[i-1]
by = y_cord[i-1]
c = x_cord[i]
cy = y_cord[i]
# gets i+2th point and calculates linear interpolation between ith and (i+1)th point
x_between = np.linspace(a,b, num=100) # iterates 100 times between i and i+1
for j in x_between:
x_cordin.append(j)
y_cordin.append(lagrangian_interpolation(j, a, ay, b, by, c, cy)) #lagragian interpolation
plt.scatter(x_cord, y_cord)
plt.plot(x_cordin, y_cordin)
| Assignment8/Interpolation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Skantastico/DS-Unit-2-Linear-Models/blob/master/LS_DSPT3_Unit2_Sprint1_assignment_regression_classification_4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="KWtX78I5MDyt" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 4*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Logistic Regression
#
#
# ## Assignment 🌯
#
# You'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
#
# > We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
# - [ ] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# - [ ] Begin with baselines for classification.
# - [ ] Use scikit-learn for logistic regression.
# - [ ] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
# - [ ] Get your model's test accuracy. (One time, at the end.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
# - [ ] Watch Aaron's [video #1](https://www.youtube.com/watch?v=pREaWFli-5I) (12 minutes) & [video #2](https://www.youtube.com/watch?v=bDQgVt4hFgY) (9 minutes) to learn about the mathematics of Logistic Regression.
#
#
# ## Stretch Goals
#
# - [ ] Add your own stretch goal(s) !
# - [ ] Make exploratory visualizations.
# - [ ] Do one-hot encoding.
# - [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html).
# - [ ] Get and plot your coefficients.
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="CvhF3cZeMDzE" colab_type="code" colab={}
# Load data downloaded from https://srcole.github.io/100burritos/
import pandas as pd
df = pd.read_csv(DATA_PATH+'burritos/burritos.csv')
# + id="QR7QE_gVMDzN" colab_type="code" colab={}
# Derive binary classification target:
# We define a 'Great' burrito as having an
# overall rating of 4 or higher, on a 5 point scale.
# Drop unrated burritos.
df = df.dropna(subset=['overall'])
df['Great'] = df['overall'] >= 4
# + id="uNtLaJlbMDzV" colab_type="code" colab={}
# Clean/combine the Burrito categories
df['Burrito'] = df['Burrito'].str.lower()
california = df['Burrito'].str.contains('california')
asada = df['Burrito'].str.contains('asada')
surf = df['Burrito'].str.contains('surf')
carnitas = df['Burrito'].str.contains('carnitas')
df.loc[california, 'Burrito'] = 'California'
df.loc[asada, 'Burrito'] = 'Asada'
df.loc[surf, 'Burrito'] = 'Surf & Turf'
df.loc[carnitas, 'Burrito'] = 'Carnitas'
df.loc[~california & ~asada & ~surf & ~carnitas, 'Burrito'] = 'Other'
# + id="mvQ93kIgMDzc" colab_type="code" colab={}
# Drop some high cardinality categoricals
df = df.drop(columns=['Notes', 'Location', 'Reviewer', 'Address', 'URL', 'Neighborhood'])
# + id="eoQmgGfjMDzi" colab_type="code" colab={}
# Drop some columns to prevent "leakage"
df = df.drop(columns=['Rec', 'overall'])
# + [markdown] id="LEdysd1nSuoJ" colab_type="text"
# ## Explore Dataset a Little
# + id="w5EWfqs8MDzr" colab_type="code" outputId="2a246edb-2099-4010-b154-494119f34f1d" colab={"base_uri": "https://localhost:8080/", "height": 244}
df.head()
# + id="kp50buS4SGM6" colab_type="code" outputId="536aa5f6-9aab-4100-c73b-2351b7272e31" colab={"base_uri": "https://localhost:8080/", "height": 683}
df.describe().T
# + id="oqup1_uHSgDb" colab_type="code" outputId="a41555c0-2574-423e-bd15-11c842a23325" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df.nunique()
# + id="7wcnrQQcS0oC" colab_type="code" outputId="9e512549-9f50-47d3-9023-da1a2f096b72" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df.isna().sum()
# + id="QBImFBZlUvkv" colab_type="code" colab={}
# Change Great to numerical
df['Great'] = df['Great'].astype(int)
# + id="U4oSoN5-VmuL" colab_type="code" outputId="ce86b8c6-925c-4341-b546-825c383ddbdd" colab={"base_uri": "https://localhost:8080/", "height": 69}
df['Great'].value_counts()
# + id="ygOym3E-Vpes" colab_type="code" outputId="82e9405d-375b-4b2c-8a1d-e36ddfa560aa" colab={"base_uri": "https://localhost:8080/", "height": 244}
df['Date'] = pd.to_datetime(df['Date'], infer_datetime_format=True)
df.head()
# + [markdown] id="AxavYCqWW1VX" colab_type="text"
# ## Create Train/Validate/Test
# + id="KYuKJd3qWurm" colab_type="code" outputId="56c2e563-fb5f-457a-d6b1-0eb4aea210b0" colab={"base_uri": "https://localhost:8080/", "height": 34}
train = df[df['Date'].dt.year.isin([2016])]
validate = df[df['Date'].dt.year.isin([2017])]
test = df[df['Date'].dt.year.isin([2018])]
print(train.shape,validate.shape,test.shape)
# + id="HxURnEeUXZ5c" colab_type="code" colab={}
# remove target from test!
test = df.drop(columns=['Great'])
# + id="lcnSqcaYZi3K" colab_type="code" outputId="8c797148-c379-4b06-8e5f-19ce14c2da88" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(train.shape, validate.shape, test.shape)
# + id="tR7b6NgYZtJH" colab_type="code" outputId="23382c73-7cc3-465f-a89a-ef6de46c3e47" colab={"base_uri": "https://localhost:8080/", "height": 34}
set(train.columns) - set(test.columns)
# + [markdown] id="ANhYyHOKbCCn" colab_type="text"
# ## Create Baselines
# + id="kGorQPGWZyeQ" colab_type="code" outputId="2a0c4c8f-1331-42e9-c672-12a35db617e0" colab={"base_uri": "https://localhost:8080/", "height": 69}
# See how data does as is
target = 'Great'
y_train = train[target]
y_train.value_counts(normalize=True)
# + id="FxgMGGZFbATd" colab_type="code" outputId="4c27c263-6bb4-4e26-ab04-b83657ac380b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
majority_class = y_train.mode()[0]
y_pred = [majority_class] * len(y_train)
y_pred
# + id="HvCleEmMaUbU" colab_type="code" outputId="718e288d-0e0c-40f6-943e-b913cd5a0f0c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Accuracy score for train
from sklearn.metrics import accuracy_score
accuracy_score(y_train, y_pred)
# + id="2R_8Lsm4bTa8" colab_type="code" outputId="7efd232a-b255-4abf-9d71-80c42733706e" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Accuracy Score for validate set
y_val = validate[target]
y_pred = [majority_class] * len(y_val)
accuracy_score(y_val, y_pred)
# + [markdown] id="TOdUwBbxTetX" colab_type="text"
# ## Logistic Regression
# + id="Rs6fYDlS9UCA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="089d2633-b971-428e-de68-0726ccf53344"
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
target = 'Great'
features = ['Yelp', 'Google', 'Volume', 'Temp', 'Uniformity',
'Cost', 'Tortilla', 'Salsa', 'Synergy']
x_train = train[features]
y_train = train[target]
x_val = validate[features]
y_val = validate[target]
encoder = ce.OneHotEncoder(use_cat_names=True)
x_train_encoded = encoder.fit_transform(x_train)
x_val_encoded = encoder.transform(x_val)
imputer = SimpleImputer()
x_train_imputed = imputer.fit_transform(x_train_encoded)
x_val_imputed = imputer.transform(x_val_encoded)
scaler = StandardScaler()
x_train_scaled = scaler.fit_transform(x_train_imputed)
x_val_scaled = scaler.transform(x_val_imputed)
model = LogisticRegression(random_state=42)
model.fit(x_train_scaled, y_train)
print('Validation Accuracy', model.score(x_val_scaled, y_val))
# + [markdown] id="OAVYr1K_iDYW" colab_type="text"
# ### Use Model on test set, having trouble getting this to work
# + id="u9mQtQLhUPoY" colab_type="code" outputId="2d273e90-b0d2-43b7-b21b-40881ad18bb3" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Do an accuracy check on Test
X_test = test[features]
X_test_encoded = encoder.transform(X_test)
X_test_imputed = imputer.transform(X_test)
X_test_scaled = scaler.transform(X_test_imputed)
y_pred = model.predict(X_test_scaled)
print('Validation accuracy: ', model.score(X_test_scaled, y_pred))
# + id="_MOgE4FqaX-e" colab_type="code" colab={}
| LS_DSPT3_Unit2_Sprint1_assignment_regression_classification_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.061424, "end_time": "2021-10-02T11:31:11.027444", "exception": false, "start_time": "2021-10-02T11:31:10.966020", "status": "completed"} tags=[]
# # **Getting Started**
#
# **Title : Lung Cancer Prediction**
#
# **Lung Cancer Status :**
#
# 0 -- > Yes
#
# 1 -- > NO
#
# + [markdown] papermill={"duration": 0.069029, "end_time": "2021-10-02T11:31:11.164974", "exception": false, "start_time": "2021-10-02T11:31:11.095945", "status": "completed"} tags=[]
# ### **DataFraming**
#
# **Read .csv file into pandas**
# + papermill={"duration": 0.099575, "end_time": "2021-10-02T11:31:11.327845", "exception": false, "start_time": "2021-10-02T11:31:11.228270", "status": "completed"} tags=[]
data = pd.read_csv('survey lung cancer.csv')
data.head()
# + [markdown] papermill={"duration": 0.070397, "end_time": "2021-10-02T11:31:11.472150", "exception": false, "start_time": "2021-10-02T11:31:11.401753", "status": "completed"} tags=[]
# ## **Exploratory data analysis**
# + papermill={"duration": 0.071973, "end_time": "2021-10-02T11:31:11.609402", "exception": false, "start_time": "2021-10-02T11:31:11.537429", "status": "completed"} tags=[]
#Shape of data
print(data.shape)
#dtypes of data
print(data.dtypes)
# + papermill={"duration": 0.094191, "end_time": "2021-10-02T11:31:11.768819", "exception": false, "start_time": "2021-10-02T11:31:11.674628", "status": "completed"} tags=[]
# Info of data
data.info()
# + papermill={"duration": 0.072553, "end_time": "2021-10-02T11:31:11.908018", "exception": false, "start_time": "2021-10-02T11:31:11.835465", "status": "completed"} tags=[]
# Checking for null values
data.isnull().sum()
# + [markdown] papermill={"duration": 0.063232, "end_time": "2021-10-02T11:31:12.034710", "exception": false, "start_time": "2021-10-02T11:31:11.971478", "status": "completed"} tags=[]
# # **Transformation**
# + papermill={"duration": 0.08226, "end_time": "2021-10-02T11:31:12.183417", "exception": false, "start_time": "2021-10-02T11:31:12.101157", "status": "completed"} tags=[]
# label encoding
data.replace({"LUNG_CANCER":{'YES':0,'NO':1}},inplace=True)
# printing the first 5 rows of the dataframe
data.head(5)
# + papermill={"duration": 0.074691, "end_time": "2021-10-02T11:31:12.321686", "exception": false, "start_time": "2021-10-02T11:31:12.246995", "status": "completed"} tags=[]
# Value_counts of loan_status
data['LUNG_CANCER'].value_counts()
# + papermill={"duration": 0.083068, "end_time": "2021-10-02T11:31:12.469633", "exception": false, "start_time": "2021-10-02T11:31:12.386565", "status": "completed"} tags=[]
# label encoding
data.replace({"GENDER":{'M':0,'F':1}},inplace=True)
# printing the first 5 rows of the dataframe
data.head(5)
# + [markdown] papermill={"duration": 0.064546, "end_time": "2021-10-02T11:31:12.599224", "exception": false, "start_time": "2021-10-02T11:31:12.534678", "status": "completed"} tags=[]
# # Data Visualization
# + papermill={"duration": 1.116112, "end_time": "2021-10-02T11:31:13.781860", "exception": false, "start_time": "2021-10-02T11:31:12.665748", "status": "completed"} tags=[]
# education & Loan Status
import seaborn as sns
sns.countplot(x='LUNG_CANCER',hue='LUNG_CANCER',data=data)
# + papermill={"duration": 0.24632, "end_time": "2021-10-02T11:31:14.094263", "exception": false, "start_time": "2021-10-02T11:31:13.847943", "status": "completed"} tags=[]
# education & Loan Status
import seaborn as sns
sns.countplot(x='GENDER',hue='LUNG_CANCER',data=data)
# + papermill={"duration": 1.810272, "end_time": "2021-10-02T11:31:15.972694", "exception": false, "start_time": "2021-10-02T11:31:14.162422", "status": "completed"} tags=[]
# let's see how data is distributed for every column
import matplotlib.pyplot as plt
plt.figure(figsize = (20, 25))
plotnumber = 1
for column in data:
if plotnumber <= 9:
ax = plt.subplot(3, 3, plotnumber)
sns.distplot(data[column])
plt.xlabel(column, fontsize = 15)
plotnumber += 1
plt.show()
# + [markdown] papermill={"duration": 0.073622, "end_time": "2021-10-02T11:31:16.121349", "exception": false, "start_time": "2021-10-02T11:31:16.047727", "status": "completed"} tags=[]
# **We can see that the distribution of data is normal ! lets move for the Model preparation.** 🚀
# + [markdown] papermill={"duration": 0.07183, "end_time": "2021-10-02T11:31:16.266208", "exception": false, "start_time": "2021-10-02T11:31:16.194378", "status": "completed"} tags=[]
# # **Model Preparation**
#
# Spilt into X & Y
# + papermill={"duration": 0.080964, "end_time": "2021-10-02T11:31:16.419712", "exception": false, "start_time": "2021-10-02T11:31:16.338748", "status": "completed"} tags=[]
# separating the data and target
X = data.drop(columns=['LUNG_CANCER'],axis=1)
y = data['LUNG_CANCER']
# + papermill={"duration": 0.081128, "end_time": "2021-10-02T11:31:16.574218", "exception": false, "start_time": "2021-10-02T11:31:16.493090", "status": "completed"} tags=[]
print("The shape of X is " , X.shape)
print("The shape of Y is " , y.shape)
# + papermill={"duration": 0.24632, "end_time": "2021-10-02T11:31:16.894893", "exception": false, "start_time": "2021-10-02T11:31:16.648573", "status": "completed"} tags=[]
from sklearn.model_selection import train_test_split
# separating into train and testing
X_train, X_test,Y_train,Y_test = train_test_split(X,y,test_size=0.2,stratify=y,random_state=42)
print("Shape of X_train is " ,X_train.shape)
print("Shape of X_test is " ,X_test.shape)
print("Shape of Y_train is " ,Y_train.shape)
print("Shape of Y_test is " ,Y_test.shape)
# + papermill={"duration": 0.085183, "end_time": "2021-10-02T11:31:17.054504", "exception": false, "start_time": "2021-10-02T11:31:16.969321", "status": "completed"} tags=[]
# After statify Y train & test values
print(Y_train.value_counts())
print(Y_test.value_counts())
# + [markdown] papermill={"duration": 0.07314, "end_time": "2021-10-02T11:31:17.202612", "exception": false, "start_time": "2021-10-02T11:31:17.129472", "status": "completed"} tags=[]
# # **Feature Scalilng**
# + papermill={"duration": 0.086397, "end_time": "2021-10-02T11:31:17.362873", "exception": false, "start_time": "2021-10-02T11:31:17.276476", "status": "completed"} tags=[]
# scaling the data
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# + papermill={"duration": 0.083302, "end_time": "2021-10-02T11:31:17.520223", "exception": false, "start_time": "2021-10-02T11:31:17.436921", "status": "completed"} tags=[]
X_train
# + [markdown] papermill={"duration": 0.0797, "end_time": "2021-10-02T11:31:17.675506", "exception": false, "start_time": "2021-10-02T11:31:17.595806", "status": "completed"} tags=[]
# # **Model Training**
#
# **We will train different model after the evaluation of model we will select out best model for production.**
#
# 1. Logistic Regression
# 2. KNN
# 3. SVC
# 3. Decision Tree
# 4. Random Forest Regressor
# 5. XgBoost
# 6. Ada Boost
# 7. Gradient Boosting
# 8. Stochascated Gradient Boosting
# 9. Stacking
# + [markdown] papermill={"duration": 0.074265, "end_time": "2021-10-02T11:31:17.824394", "exception": false, "start_time": "2021-10-02T11:31:17.750129", "status": "completed"} tags=[]
# ## **Logistic Regression**
# + papermill={"duration": 0.165392, "end_time": "2021-10-02T11:31:18.064687", "exception": false, "start_time": "2021-10-02T11:31:17.899295", "status": "completed"} tags=[]
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
lr = LogisticRegression()
lr.fit(X_train, Y_train)
y_pred = lr.predict(X_test)
lr_train_acc = accuracy_score(Y_train, lr.predict(X_train))
lr_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of Logistic Regression Model is {lr_train_acc}")
print(f"Test Accuracy of Logistic Regression Model is {lr_test_acc}")
# + papermill={"duration": 0.084961, "end_time": "2021-10-02T11:31:18.224750", "exception": false, "start_time": "2021-10-02T11:31:18.139789", "status": "completed"} tags=[]
# confusion matrix
confusion_matrix(Y_test, y_pred)
# + papermill={"duration": 0.08672, "end_time": "2021-10-02T11:31:18.385344", "exception": false, "start_time": "2021-10-02T11:31:18.298624", "status": "completed"} tags=[]
# classification report
print(classification_report(Y_test, y_pred))
# + [markdown] papermill={"duration": 0.075141, "end_time": "2021-10-02T11:31:18.536295", "exception": false, "start_time": "2021-10-02T11:31:18.461154", "status": "completed"} tags=[]
# ## **KNeighborsClassifier**
# + papermill={"duration": 0.159961, "end_time": "2021-10-02T11:31:18.770812", "exception": false, "start_time": "2021-10-02T11:31:18.610851", "status": "completed"} tags=[]
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
knn.fit(X_train, Y_train)
y_pred = knn.predict(X_test)
knn_train_acc = accuracy_score(Y_train, knn.predict(X_train))
knn_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of KNN Model is {knn_train_acc}")
print(f"Test Accuracy of KNN Model is {knn_test_acc}")
# + papermill={"duration": 0.089526, "end_time": "2021-10-02T11:31:18.936112", "exception": false, "start_time": "2021-10-02T11:31:18.846586", "status": "completed"} tags=[]
# confusion matrix
confusion_matrix(Y_test, y_pred)
# + papermill={"duration": 0.08812, "end_time": "2021-10-02T11:31:19.103374", "exception": false, "start_time": "2021-10-02T11:31:19.015254", "status": "completed"} tags=[]
# classification report
print(classification_report(Y_test, y_pred))
# + [markdown] papermill={"duration": 0.07577, "end_time": "2021-10-02T11:31:19.255964", "exception": false, "start_time": "2021-10-02T11:31:19.180194", "status": "completed"} tags=[]
# ## **SVC**
#
# + papermill={"duration": 0.090449, "end_time": "2021-10-02T11:31:19.422865", "exception": false, "start_time": "2021-10-02T11:31:19.332416", "status": "completed"} tags=[]
from sklearn.svm import SVC
svc = SVC()
svc.fit(X_train, Y_train)
y_pred = svc.predict(X_test)
svc_train_acc = accuracy_score(Y_train, svc.predict(X_train))
svc_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of SVC Model is {svc_train_acc}")
print(f"Test Accuracy of SVC Model is {svc_test_acc}")
# + papermill={"duration": 0.086247, "end_time": "2021-10-02T11:31:19.587191", "exception": false, "start_time": "2021-10-02T11:31:19.500944", "status": "completed"} tags=[]
# confusion matrix
confusion_matrix(Y_test, y_pred)
# + papermill={"duration": 0.090064, "end_time": "2021-10-02T11:31:19.754922", "exception": false, "start_time": "2021-10-02T11:31:19.664858", "status": "completed"} tags=[]
# classification report
print(classification_report(Y_test, y_pred))
# + [markdown] papermill={"duration": 0.077376, "end_time": "2021-10-02T11:31:19.909996", "exception": false, "start_time": "2021-10-02T11:31:19.832620", "status": "completed"} tags=[]
# ## **DecisionTreeClassifier**
# + papermill={"duration": 0.115986, "end_time": "2021-10-02T11:31:20.103618", "exception": false, "start_time": "2021-10-02T11:31:19.987632", "status": "completed"} tags=[]
from sklearn.tree import DecisionTreeClassifier
dtc = DecisionTreeClassifier()
dtc.fit(X_train, Y_train)
y_pred = dtc.predict(X_test)
dtc_train_acc = accuracy_score(Y_train, dtc.predict(X_train))
dtc_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of Decision Tree Model is {dtc_train_acc}")
print(f"Test Accuracy of Decision Tree Model is {dtc_test_acc}")
# + papermill={"duration": 0.086122, "end_time": "2021-10-02T11:31:20.267160", "exception": false, "start_time": "2021-10-02T11:31:20.181038", "status": "completed"} tags=[]
# confusion matrix
confusion_matrix(Y_test, y_pred)
# + papermill={"duration": 0.089446, "end_time": "2021-10-02T11:31:20.434006", "exception": false, "start_time": "2021-10-02T11:31:20.344560", "status": "completed"} tags=[]
# classification report
print(classification_report(Y_test, y_pred))
# + [markdown] papermill={"duration": 0.078879, "end_time": "2021-10-02T11:31:20.591732", "exception": false, "start_time": "2021-10-02T11:31:20.512853", "status": "completed"} tags=[]
# ## **Hyper parameter tuning**
# + papermill={"duration": 4.954645, "end_time": "2021-10-02T11:31:25.626276", "exception": false, "start_time": "2021-10-02T11:31:20.671631", "status": "completed"} tags=[]
# hyper parameter tuning
from sklearn.model_selection import GridSearchCV
grid_params = {
'criterion' : ['gini', 'entropy'],
'max_depth' : [3, 5, 7, 10],
'min_samples_split' : range(2, 10, 1),
'min_samples_leaf' : range(2, 10, 1)
}
grid_search = GridSearchCV(dtc, grid_params, cv = 5, n_jobs = -1, verbose = 1)
grid_search.fit(X_train, Y_train)
# + papermill={"duration": 0.089268, "end_time": "2021-10-02T11:31:25.796807", "exception": false, "start_time": "2021-10-02T11:31:25.707539", "status": "completed"} tags=[]
# best parameters and best score
print(grid_search.best_params_)
print(grid_search.best_score_)
# + papermill={"duration": 0.092374, "end_time": "2021-10-02T11:31:25.970627", "exception": false, "start_time": "2021-10-02T11:31:25.878253", "status": "completed"} tags=[]
dtc = grid_search.best_estimator_
y_pred = dtc.predict(X_test)
dtc_train_acc = accuracy_score(Y_train, dtc.predict(X_train))
dtc_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of Decesion Tree Model is {dtc_train_acc}")
print(f"Test Accuracy of Decesion Tree Model is {dtc_test_acc}")
# + [markdown] papermill={"duration": 0.081609, "end_time": "2021-10-02T11:31:26.135313", "exception": false, "start_time": "2021-10-02T11:31:26.053704", "status": "completed"} tags=[]
# **Visualization the DTC tree.**
# + papermill={"duration": 1.80162, "end_time": "2021-10-02T11:31:28.018149", "exception": false, "start_time": "2021-10-02T11:31:26.216529", "status": "completed"} tags=[]
from sklearn import tree
plt.figure(figsize=(15,10))
tree.plot_tree(dtc,filled=True)
# + [markdown] papermill={"duration": 0.084498, "end_time": "2021-10-02T11:31:28.187573", "exception": false, "start_time": "2021-10-02T11:31:28.103075", "status": "completed"} tags=[]
# ## **Random Forest Classifier**
# + papermill={"duration": 0.471868, "end_time": "2021-10-02T11:31:28.744148", "exception": false, "start_time": "2021-10-02T11:31:28.272280", "status": "completed"} tags=[]
from sklearn.ensemble import RandomForestClassifier
rand_clf = RandomForestClassifier(criterion = 'gini', max_depth = 3, max_features = 'sqrt', min_samples_leaf = 2, min_samples_split = 4, n_estimators = 180)
rand_clf.fit(X_train, Y_train)
y_pred = rand_clf.predict(X_test)
rand_clf_train_acc = accuracy_score(Y_train, rand_clf.predict(X_train))
rand_clf_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of Random Forest Model is {rand_clf_train_acc}")
print(f"Test Accuracy of Random Forest Model is {rand_clf_test_acc}")
# + [markdown] papermill={"duration": 0.084613, "end_time": "2021-10-02T11:31:28.914226", "exception": false, "start_time": "2021-10-02T11:31:28.829613", "status": "completed"} tags=[]
# ## **KNN**
# + papermill={"duration": 0.116222, "end_time": "2021-10-02T11:31:29.116965", "exception": false, "start_time": "2021-10-02T11:31:29.000743", "status": "completed"} tags=[]
from sklearn.neighbors import KNeighborsClassifier
k_model = KNeighborsClassifier(n_neighbors=16)
kfitModel = k_model.fit(X_train, Y_train)
# accuracy score on training data
kX_train_prediction = kfitModel.predict(X_train)
training_data_accuray = accuracy_score(kX_train_prediction,Y_train)
print('Accuracy on training data : ', training_data_accuray)
# accuracy score on testing data
kX_test_prediction = kfitModel.predict(X_test)
kx_lgr_test_data_accuray = accuracy_score(kX_test_prediction,Y_test)
print('Accuracy on test data : ', kx_lgr_test_data_accuray)
# + [markdown] papermill={"duration": 0.084693, "end_time": "2021-10-02T11:31:29.289565", "exception": false, "start_time": "2021-10-02T11:31:29.204872", "status": "completed"} tags=[]
# # **Boosting**
# + papermill={"duration": 25.020369, "end_time": "2021-10-02T11:31:54.394684", "exception": false, "start_time": "2021-10-02T11:31:29.374315", "status": "completed"} tags=[]
#Ada Boost Classifie
from sklearn.ensemble import AdaBoostClassifier
ada = AdaBoostClassifier(base_estimator = dtc)
parameters = {
'n_estimators' : [50, 70, 90, 120, 180, 200],
'learning_rate' : [0.001, 0.01, 0.1, 1, 10],
'algorithm' : ['SAMME', 'SAMME.R']
}
grid_search = GridSearchCV(ada, parameters, n_jobs = -1, cv = 5, verbose = 1)
grid_search.fit(X_train, Y_train)
# + papermill={"duration": 0.097409, "end_time": "2021-10-02T11:31:54.582875", "exception": false, "start_time": "2021-10-02T11:31:54.485466", "status": "completed"} tags=[]
print(grid_search.best_params_)
print(grid_search.best_score_)
# + papermill={"duration": 0.475837, "end_time": "2021-10-02T11:31:55.147990", "exception": false, "start_time": "2021-10-02T11:31:54.672153", "status": "completed"} tags=[]
ada = AdaBoostClassifier(base_estimator = dtc, algorithm = 'SAMME.R', learning_rate = 0.1, n_estimators = 180)
ada.fit(X_train, Y_train)
ada_train_acc = accuracy_score(Y_train, ada.predict(X_train))
ada_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of Ada Boost Model is {ada_train_acc}")
print(f"Test Accuracy of Ada Boost Model is {ada_test_acc}")
# + papermill={"duration": 0.097939, "end_time": "2021-10-02T11:31:55.335669", "exception": false, "start_time": "2021-10-02T11:31:55.237730", "status": "completed"} tags=[]
# confusion matrix
confusion_matrix(Y_test, y_pred)
# + papermill={"duration": 0.106649, "end_time": "2021-10-02T11:31:55.531451", "exception": false, "start_time": "2021-10-02T11:31:55.424802", "status": "completed"} tags=[]
# classification report
print(classification_report(Y_test, y_pred))
# + [markdown] papermill={"duration": 0.090846, "end_time": "2021-10-02T11:31:55.714844", "exception": false, "start_time": "2021-10-02T11:31:55.623998", "status": "completed"} tags=[]
# ## **GradientBoostingClassifier**
# + papermill={"duration": 8.057705, "end_time": "2021-10-02T11:32:03.863723", "exception": false, "start_time": "2021-10-02T11:31:55.806018", "status": "completed"} tags=[]
#Gradient Boosting Classifier
from sklearn.ensemble import GradientBoostingClassifier
gb = GradientBoostingClassifier()
parameters = {
'loss': ['deviance', 'exponential'],
'learning_rate': [0.001, 0.1, 1, 10],
'n_estimators': [100, 150, 180, 200]
}
grid_search = GridSearchCV(gb, parameters, cv = 5, n_jobs = -1, verbose = 1)
grid_search.fit(X_train, Y_train)
# + papermill={"duration": 0.102348, "end_time": "2021-10-02T11:32:04.058848", "exception": false, "start_time": "2021-10-02T11:32:03.956500", "status": "completed"} tags=[]
# best parameter and best score
print(grid_search.best_params_)
print(grid_search.best_score_)
# + papermill={"duration": 0.272347, "end_time": "2021-10-02T11:32:04.423589", "exception": false, "start_time": "2021-10-02T11:32:04.151242", "status": "completed"} tags=[]
gb = GradientBoostingClassifier(learning_rate = 1, loss = 'exponential', n_estimators = 180)
gb.fit(X_train, Y_train)
y_pred = gb.predict(X_test)
gb_train_acc = accuracy_score(Y_train, gb.predict(X_train))
gb_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of Gradient Boosting Classifier Model is {gb_train_acc}")
print(f"Test Accuracy of Gradient Boosting Classifier Model is {gb_test_acc}")
# + papermill={"duration": 0.100659, "end_time": "2021-10-02T11:32:04.616706", "exception": false, "start_time": "2021-10-02T11:32:04.516047", "status": "completed"} tags=[]
# confusion matrix
confusion_matrix(Y_test, y_pred)
# + papermill={"duration": 0.107779, "end_time": "2021-10-02T11:32:04.818207", "exception": false, "start_time": "2021-10-02T11:32:04.710428", "status": "completed"} tags=[]
# classification report
print(classification_report(Y_test, y_pred))
# + [markdown] papermill={"duration": 0.093229, "end_time": "2021-10-02T11:32:05.006960", "exception": false, "start_time": "2021-10-02T11:32:04.913731", "status": "completed"} tags=[]
# ## **Stochastic Gradient Boosting (SGB)**
# + papermill={"duration": 0.188164, "end_time": "2021-10-02T11:32:05.289598", "exception": false, "start_time": "2021-10-02T11:32:05.101434", "status": "completed"} tags=[]
sgbc = GradientBoostingClassifier(learning_rate = 0.1, subsample = 0.9, max_features = 0.75, loss = 'deviance',
n_estimators = 100)
sgbc.fit(X_train, Y_train)
y_pred = sgbc.predict(X_test)
sgbc_train_acc = accuracy_score(Y_train, sgbc.predict(X_train))
sgbc_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of SGB Model is {sgbc_train_acc}")
print(f"Test Accuracy of SGB Model is {sgbc_test_acc}")
# + papermill={"duration": 0.103311, "end_time": "2021-10-02T11:32:05.485993", "exception": false, "start_time": "2021-10-02T11:32:05.382682", "status": "completed"} tags=[]
# confusion matrix
confusion_matrix(Y_test, y_pred)
# + papermill={"duration": 0.108185, "end_time": "2021-10-02T11:32:05.688519", "exception": false, "start_time": "2021-10-02T11:32:05.580334", "status": "completed"} tags=[]
# classification report
print(classification_report(Y_test, y_pred))
# + [markdown] papermill={"duration": 0.09441, "end_time": "2021-10-02T11:32:05.880093", "exception": false, "start_time": "2021-10-02T11:32:05.785683", "status": "completed"} tags=[]
# ## **Cat Boost**
# + papermill={"duration": 0.462794, "end_time": "2021-10-02T11:32:06.438685", "exception": false, "start_time": "2021-10-02T11:32:05.975891", "status": "completed"} tags=[]
#Cat boost
from catboost import CatBoostClassifier
cat = CatBoostClassifier(iterations = 30, learning_rate = 0.1)
cat.fit(X_train, Y_train)
y_pred = cat.predict(X_test)
# + papermill={"duration": 0.105387, "end_time": "2021-10-02T11:32:06.638703", "exception": false, "start_time": "2021-10-02T11:32:06.533316", "status": "completed"} tags=[]
cat_train_acc = accuracy_score(Y_train, cat.predict(X_train))
cat_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of Cat Boost Classifier Model is {cat_train_acc}")
print(f"Test Accuracy of Cat Boost Classifier Model is {cat_test_acc}")
# + [markdown] papermill={"duration": 0.094352, "end_time": "2021-10-02T11:32:06.829186", "exception": false, "start_time": "2021-10-02T11:32:06.734834", "status": "completed"} tags=[]
# ## **XGB Classifier**
# + papermill={"duration": 0.212621, "end_time": "2021-10-02T11:32:07.137371", "exception": false, "start_time": "2021-10-02T11:32:06.924750", "status": "completed"} tags=[]
from xgboost import XGBClassifier
xgb = XGBClassifier(booster = 'gblinear', learning_rate = 1, n_estimators = 10)
xgb.fit(X_train, Y_train)
y_pred = xgb.predict(X_test)
xgb_train_acc = accuracy_score(Y_train, xgb.predict(X_train))
xgb_test_acc = accuracy_score(Y_test, y_pred)
print(f"Training Accuracy of XGB Model is {xgb_train_acc}")
print(f"Test Accuracy of XGB Model is {xgb_test_acc}")
# + [markdown] papermill={"duration": 0.094472, "end_time": "2021-10-02T11:32:07.327925", "exception": false, "start_time": "2021-10-02T11:32:07.233453", "status": "completed"} tags=[]
# # **Stacking**
# + papermill={"duration": 0.105958, "end_time": "2021-10-02T11:32:07.527773", "exception": false, "start_time": "2021-10-02T11:32:07.421815", "status": "completed"} tags=[]
# let's divide our dataset into training set and holdout set by 50%
from sklearn.model_selection import train_test_split
train, val_train, test, val_test = train_test_split(X, y, test_size = 0.5, random_state = 355)
# + papermill={"duration": 0.104764, "end_time": "2021-10-02T11:32:07.728154", "exception": false, "start_time": "2021-10-02T11:32:07.623390", "status": "completed"} tags=[]
# let's split the training set again into training and test dataset
X_train, X_test, y_train, y_test = train_test_split(train, test, test_size = 0.2, random_state = 355)
# + papermill={"duration": 0.107778, "end_time": "2021-10-02T11:32:07.934350", "exception": false, "start_time": "2021-10-02T11:32:07.826572", "status": "completed"} tags=[]
svm = SVC()
svm.fit(X_train, y_train)
# + papermill={"duration": 0.138413, "end_time": "2021-10-02T11:32:08.168622", "exception": false, "start_time": "2021-10-02T11:32:08.030209", "status": "completed"} tags=[]
# using Logistic Regression and SVM algorithm as base models.
# Let's fit both of the models first on the X_train and y_train data.
lr = LogisticRegression(solver='liblinear')
lr.fit(X_train, y_train)
# + papermill={"duration": 0.109991, "end_time": "2021-10-02T11:32:08.380545", "exception": false, "start_time": "2021-10-02T11:32:08.270554", "status": "completed"} tags=[]
predict_val1 = lr.predict(val_train)
predict_val2 = svm.predict(val_train)
# + papermill={"duration": 0.104201, "end_time": "2021-10-02T11:32:08.582148", "exception": false, "start_time": "2021-10-02T11:32:08.477947", "status": "completed"} tags=[]
predict_val = np.column_stack((predict_val1, predict_val2))
# + papermill={"duration": 0.108999, "end_time": "2021-10-02T11:32:08.788780", "exception": false, "start_time": "2021-10-02T11:32:08.679781", "status": "completed"} tags=[]
#Let's get the prediction of all the base models on test set X_set
predict_test1 = lr.predict(X_test)
predict_test2 = svm.predict(X_test)
# + papermill={"duration": 0.105714, "end_time": "2021-10-02T11:32:08.993283", "exception": false, "start_time": "2021-10-02T11:32:08.887569", "status": "completed"} tags=[]
# Let's stack the prediction values for validation set together as 'predict_set'
predict_test = np.column_stack((predict_test1, predict_test2))
# + papermill={"duration": 0.282984, "end_time": "2021-10-02T11:32:09.373686", "exception": false, "start_time": "2021-10-02T11:32:09.090702", "status": "completed"} tags=[]
rand_clf = RandomForestClassifier()
rand_clf.fit(predict_val, val_test)
# + papermill={"duration": 0.116239, "end_time": "2021-10-02T11:32:09.586534", "exception": false, "start_time": "2021-10-02T11:32:09.470295", "status": "completed"} tags=[]
stacking_acc = accuracy_score(y_test, rand_clf.predict(predict_test))
print(stacking_acc)
# + papermill={"duration": 0.127649, "end_time": "2021-10-02T11:32:09.813517", "exception": false, "start_time": "2021-10-02T11:32:09.685868", "status": "completed"} tags=[]
# confusion matrix
confusion_matrix(y_test, rand_clf.predict(predict_test))
# + papermill={"duration": 0.127712, "end_time": "2021-10-02T11:32:10.051992", "exception": false, "start_time": "2021-10-02T11:32:09.924280", "status": "completed"} tags=[]
# classification report
print(classification_report(y_test, rand_clf.predict(predict_test)))
# + papermill={"duration": 0.122791, "end_time": "2021-10-02T11:32:10.292791", "exception": false, "start_time": "2021-10-02T11:32:10.170000", "status": "completed"} tags=[]
models = ['Logistic Regression', 'KNN', 'SVC', 'Decision Tree', 'Random Forest','Ada Boost', 'Gradient Boosting', 'SGB', 'XgBoost', 'Stacking', 'Cat Boost']
scores = [lr_test_acc, knn_test_acc, svc_test_acc, dtc_test_acc, rand_clf_test_acc, ada_test_acc, gb_test_acc, sgbc_test_acc, xgb_test_acc, stacking_acc, cat_test_acc]
models = pd.DataFrame({'Model' : models, 'Score' : scores})
models.sort_values(by = 'Score', ascending = False)
# + papermill={"duration": 0.330856, "end_time": "2021-10-02T11:32:10.723072", "exception": false, "start_time": "2021-10-02T11:32:10.392216", "status": "completed"} tags=[]
plt.figure(figsize = (18, 8))
sns.barplot(x = 'Model', y = 'Score', data = models)
plt.show()
# + [markdown] papermill={"duration": 0.098307, "end_time": "2021-10-02T11:32:10.921409", "exception": false, "start_time": "2021-10-02T11:32:10.823102", "status": "completed"} tags=[]
# ### ***Logistic Regression gives us the best result so we will save this model for production.***
| Contributions/Lung_cancer_prdiction/lung-cancer-different-models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] azdata_cell_guid="d495f8be-74c3-4658-b897-ad69e6ed88ac"
# <img src="https://github.com/Microsoft/sqlworkshops/blob/master/graphics/solutions-microsoft-logo-small.png?raw=true" alt="Microsoft">
# <br>
#
# # SQL Server 2019 big data cluster Tutorial - CTP 3.2
# ## 00 - Scenario Overview and System Setup
#
# In this set of tutorials you'll work with an end-to-end scenario that uses SQL Server 2019's big data clusters to solve real-world problems.
#
# + [markdown] azdata_cell_guid="3815241f-e81e-4cf1-a48e-c6e67b0ccf7c"
# ## Wide World Importers
#
# Wide World Importers (WWI) is a traditional brick and mortar business that makes specialty items for other companies to use in their products. They design, sell and ship these products worldwide.
#
# WWI corporate has now added a new partnership with a company called "AdventureWorks", which sells bicycles both online and in-store. The AdventureWorks company has asked WWI to produce super-hero themed baskets, seats and other bicycle equipment for a new line of bicycles. WWI corporate has asked the IT department to develop a pilot program with these goals:
#
# - Integrate the large amounts of data from the AdventureWorks company including customers, products and sales
# - Allow a cross-selling strategy so that current WWI customers and AdventureWorks customers see their information without having to re-enter it
# - Incorporate their online sales information for deeper analysis
# - Provide a historical data set so that the partnership can be evaluated
# - Ensure this is a "framework" approach, so that it can be re-used with other partners
#
# WWI has a typical N-Tier application that provides a series of terminals, a Business Logic layer, and a Database back-end. They use on-premises systems, and are interested in linking these to the cloud.
#
# In this series of tutorials, you will build a solution using the scale-out features of SQL Server 2019, Data Virtualization, Data Marts, and the Data Lake features.
# + [markdown] azdata_cell_guid="1c3e4b5e-fef4-43ef-a4e3-aa33fe99e25d"
# ## Running these Tutorials
#
# - You can read through the output of these completed tutorials if you wish - or:
#
# - You can follow along with the steps you see in these tutorials by copying the code into a SQL Query window and Spark Notebook using the Azure Data Studio tool, or you can click here to download these Jupyter Notebooks and run them in Azure Data Studio for a hands-on experience.
#
# - If you would like to run the tutorials, you'll need a SQL Server 2019 big data cluster and the client tools installed. If you want to set up your own cluster, <a href="https://docs.microsoft.com/en-us/sql/big-data-cluster/deploy-get-started?view=sqlallproducts-allversions" target="_blank">click this reference and follow the steps you see there for the server and tools you need</a>.
#
# - You will need to have the following:
# - Your **Knox Password**
# - The **Knox IP Address**
# - The `sa` **Username** and **Password** to your Master Instance
# - The **IP address** to the SQL Server big data cluster Master Instance
# - The **name** of your big data cluster
#
# For a complete workshop on SQL Server 2019's big data clusters, <a href="https://github.com/Microsoft/sqlworkshops/tree/master/sqlserver2019bigdataclusters" target="_blank">check out this resource</a>.
# + [markdown] azdata_cell_guid="5220c555-f819-409e-b206-de9a2dd6d434"
# ## Copy Database backups to the SQL Server 2019 big data cluster Master Instance
#
# The first step for the solution is to copy the database backups from WWI from their location on the cloud and then up to your cluster.
#
# These commands use the `curl` program to pull the files down. [You can read more about curl here](https://curl.haxx.se/).
#
# The next set of commands use the `kubectl` command to copy the files from where you downloaded them to the data directory of the SQL Server 2019 bdc Master Instance. [You can read more about kubectl here](https://kubernetes.io/docs/reference/kubectl/overview/).
#
# Note that you will need to replace the section of the script marked with `<ReplaceWithClusterName>` with the name of your SQL Server 2019 bdc. (It does not need single or double quotes, just the letters of your cluster name.)
#
# Notice also that these commands assume a `c:\temp` location, if you want to use another drive or directory, edit accordingly.
#
# Once you have edited these commands, you can open a Command Prompt *(not PowerShell)* on your system and copy and paste each block, one at a time and run them there, observing the output.
#
# In the next tutorial you will restore these databases on the Master Instance.
# + azdata_cell_guid="3e1f2304-cc0a-4e0e-96e2-333401b52036"
REM Create a temporary directory for the files
md c:\temp
# cd c:\temp
REM Get the database backups
curl "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/WWI.bak" -o c:\temp\WWI.bak
curl "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/AdventureWorks.bak" -o c:\temp\AdventureWorks.bak
curl "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/AdventureWorksDW.bak" -o c:\temp\AdventureWorksDW.bak
curl "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/Analysis.bak" -o c:\temp\Analysis.bak
curl "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/sales.bak" -o c:\temp\sales.bak
curl "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/NYC.bak" -o c:\temp\NYC.bak
curl "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/WWIDW.bak" -o c:\temp\WWIDW.bak
# + azdata_cell_guid="19106890-7c6c-4631-9acc-3dda4d2a50ab"
REM Copy the backups to the data location on the SQL Server Master Instance
# cd c:\temp
kubectl cp WWI.bak master-0:/var/opt/mssql/data -c mssql-server -n <ReplaceWithClusterName>
kubectl cp WWIDW.bak master-0:/var/opt/mssql/data -c mssql-server -n <ReplaceWithClusterName>
kubectl cp sales.bak master-0:/var/opt/mssql/data -c mssql-server -n <ReplaceWithClusterName>
kubectl cp analysis.bak master-0:/var/opt/mssql/data -c mssql-server -n <ReplaceWithClusterName>
kubectl cp AdventureWorks.bak master-0:/var/opt/mssql/data -c mssql-server -n <ReplaceWithClusterName>
kubectl cp AdventureWorksDW.bak master-0:/var/opt/mssql/data -c mssql-server -n <ReplaceWithClusterName>
kubectl cp NYC.bak master-0:/var/opt/mssql/data -c mssql-server -n <ReplaceWithClusterName>
# + [markdown] azdata_cell_guid="2c426b35-dc57-4dc8-819d-6642deb69110"
# ## Copy Exported Data to Storage Pool
#
# Next, you'll download a few text files that will form the external data to be ingested into the Storage Pool HDFS store. In production environments, you have multiple options for moving data into HDFS, such as Spark Streaming or the Azure Data Factory.
#
# The first code block creates directories in the HDFS store. The second block downloads the source data from a web location. And in the final block, you'll copy the data from your local system to the SQL Server 2019 big data cluster Storage Pool.
#
# You need to replace the `<ReplaceWithHDFSGatewayPassword>`, `<ReplaceWithHDFSGatewayEndpoint>`, and potentially the drive letter and directory values with the appropriate information on your system.
# > (You can use **CTL-H** to open the Find and Replace dialog in the cell)
# + azdata_cell_guid="f2143d4e-6eb6-4bbc-864a-b417398adc21"
REM Make the Directories in HDFS
curl -i -L -k -u root:<ReplaceWithKnoxPassword> -X PUT "https://<ReplaceWithHDFSGatewayEndpoint>:30443/gateway/default/webhdfs/v1/product_review_data?op=MKDIRS"
curl -i -L -k -u root:<ReplaceWithKnoxPassword> -X PUT "https://<ReplaceWithHDFSGatewayEndpoint>:30443/gateway/default/webhdfs/v1/partner_customers?op=MKDIRS"
curl -i -L -k -u root:<ReplaceWithKnoxPassword> -X PUT "https://<ReplaceWithHDFSGatewayEndpoint>:30443/gateway/default/webhdfs/v1/partner_products?op=MKDIRS"
curl -i -L -k -u root:<ReplaceWithKnoxPassword> -X PUT "https://<ReplaceWithHDFSGatewayEndpoint>:30443/gateway/default/webhdfs/v1/web_logs?op=MKDIRS"
# + azdata_cell_guid="c8a74514-2e0d-4f3c-99dd-4c541c11e15e"
REM Get the textfiles
curl -G "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/product_reviews_sample.csv" -o product_reviews.csv
curl -G "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/customers.csv" -o customers.csv
curl -G "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/stockitemholdings.csv" -o products.csv
curl -G "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/web_clickstreams.csv" -o web_clickstreams.csv
curl -G "https://cs7a9736a9346a1x44c6xb00.blob.core.windows.net/backups/training-formatted.csv" -o training-formatted.csv
# + azdata_cell_guid="9c9e49ef-ef0d-47c4-92fd-b7e7bfa2d2f2"
REM Copy the text files to the HDFS directories
curl -i -L -k -u root:<ReplaceWithHDFSGatewayPassword> -X PUT "https://<ReplaceWithHDFSGatewayEndpoint>:30443/gateway/default/webhdfs/v1/product_review_data/product_reviews.csv?op=create&overwrite=true" -H "Content-Type: application/octet-stream" -T "product_reviews.csv"
curl -i -L -k -u root:<ReplaceWithHDFSGatewayPassword> -X PUT "https://<ReplaceWithHDFSGatewayEndpoint>:30443/gateway/default/webhdfs/v1/partner_customers/customers.csv?op=create&overwrite=true" -H "Content-Type: application/octet-stream" -T "customers.csv"
curl -i -L -k -u root:<ReplaceWithHDFSGatewayPassword> -X PUT "https://<ReplaceWithHDFSGatewayEndpoint>:30443/gateway/default/webhdfs/v1/partner_products/products.csv?op=create&overwrite=true" -H "Content-Type: application/octet-stream" -T "products.csv"
curl -i -L -k -u root:<ReplaceWithHDFSGatewayPassword> -X PUT "https://<ReplaceWithHDFSGatewayEndpoint>:30443/gateway/default/webhdfs/v1/web_logs/web_clickstreams.csv?op=create&overwrite=true" -H "Content-Type: application/octet-stream" -T "web_clickstreams.csv"
# + [markdown] azdata_cell_guid="519aa112-47e0-443b-9b27-05fc02349b09"
# ## Next Step: Working with the SQL Server 2019 big data cluster Master Instance
#
# Now you're ready to open the next Python Notebook - [bdc_tutorial_01.ipynb](bdc_tutorial_01.ipynb) - to learn how to work with the SQL Server 2019 bdc Master Instance.
| sqlserver2019bigdataclusters/SQL2019BDC/notebooks/bdc_tutorial_00.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="GfqUrXjRZDxM" outputId="91bf4429-f677-40d9-ea96-a2ee3f517845" colab={"base_uri": "https://localhost:8080/"}
import os
# Find the latest version of spark 3.0 from http://www-us.apache.org/dist/spark/ and enter as the spark version
# For example:
spark_version = 'spark-3.0.1'
# spark_version = 'spark-3.<enter version>'
os.environ['SPARK_VERSION']=spark_version
# Install Spark and Java
# !apt-get update
# !apt-get install openjdk-11-jdk-headless -qq > /dev/null
# !wget -q http://www-us.apache.org/dist/spark/$SPARK_VERSION/$SPARK_VERSION-bin-hadoop2.7.tgz
# !tar xf $SPARK_VERSION-bin-hadoop2.7.tgz
# !pip install -q findspark
# Set Environment Variables
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-11-openjdk-amd64"
os.environ["SPARK_HOME"] = f"/content/{spark_version}-bin-hadoop2.7"
# Start a SparkSession
import findspark
findspark.init()
# + id="2X3OQPfUvGX9" outputId="2b7b0431-ff55-422b-e213-37b9e854694d" colab={"base_uri": "https://localhost:8080/"}
# !wget https://jdbc.postgresql.org/download/postgresql-42.2.9.jar
# + id="iD4WphUqZF2L"
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("Sports").config("spark.driver.extraClassPath","/content/postgresql-42.2.9.jar").getOrCreate()
# + [markdown] id="A_dg78ZuswA7"
# Extraction
# + id="701kGYE3ZJo0" outputId="ec544518-f22a-4296-9cb1-6235dd52c529" colab={"base_uri": "https://localhost:8080/"}
from pyspark import SparkFiles
# Connecting to Amazon public dataset for sports
url = "https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_us_Sports_v1_00.tsv.gz"
spark.sparkContext.addFile(url)
sports_df = spark.read.option("header", "true").csv(SparkFiles.get("amazon_reviews_us_Sports_v1_00.tsv.gz"), inferSchema=True, sep="\t")
sports_df.show(10)
# + id="HpZNQIBxl0O3" outputId="d5e166d2-9e8c-4b36-ed07-2374bc6574fa" colab={"base_uri": "https://localhost:8080/"}
sports_df.count()
# + id="qAfExIVrbwss" outputId="a7826670-4900-4abd-95e9-4b3f670bf059" colab={"base_uri": "https://localhost:8080/"}
# Print our schema
sports_df.printSchema()
# + [markdown] id="2dl0jddNs2yO"
# Transformation
# + id="O5InU_t1ksgD"
# Import struct fields that we can use
from pyspark.sql.types import StructField, StringType, IntegerType, StructType, DateType
# + id="v_krpxAmlYsJ"
# Next we need to create the list of struct fields
schema = [StructField("marketplace", StringType(), True),
StructField("customer_id", IntegerType(), True),
StructField("review_id", StringType(), True),
StructField("product_id", StringType(), True),
StructField("product_parent", IntegerType(), True),
StructField("product_title", StringType(), True),
StructField("product_category", StringType(), True),
StructField("star_rating", IntegerType(), True),
StructField("helpful_votes", IntegerType(), True),
StructField("total_votes", IntegerType(), True),
StructField("vine", StringType(), True),
StructField("verified_purchase", StringType(), True),
StructField("review_headline", StringType(), True),
StructField("review_body", StringType(), True),
StructField("review_date", DateType(), True)]
# + id="GIW7nAapoYXw" outputId="98e39e10-6eb0-46ab-c84c-d7f0c2f51e14" colab={"base_uri": "https://localhost:8080/"}
final = StructType(fields=schema)
sports_df = spark.read.option("header", "true").csv(SparkFiles.get("amazon_reviews_us_Sports_v1_00.tsv.gz"), schema=final, sep="\t")
sports_df.show(10)
# + id="7jMudECjoLfM" outputId="d1dae665-e71f-4db4-fd5b-82d18e9a497a" colab={"base_uri": "https://localhost:8080/"}
# Print our schema
sports_df.printSchema()
# + id="vD14aFYloR4W"
# Constructing Review table
review_id_table = sports_df.select(['review_id', 'customer_id', 'product_id', 'product_parent', 'review_date'])
# + id="u9e1e46Cqerz"
# Constructing Product table
products = sports_df.select(['product_id', 'product_title']).drop_duplicates()
# + id="aSPr6NYCqyh-"
# Constructing Customers table
customers = sports_df.groupby("customer_id").agg({"customer_id":"count"}).withColumnRenamed("count(customer_id)","customer_count")
# + id="aWp-GY4crwY_"
# Constructing Vine table
vine_table = sports_df.select(["review_id", "star_rating", "helpful_votes", "total_votes", "vine"])
# + [markdown] id="DwqtynLts9Mb"
# **Loading**
# + id="e6-8QbAJsm9B"
# Configure settings for RDS
mode = "append"
jdbc_url="jdbc:postgresql://bigdata-hw.cr5bt5kg46tf.us-west-1.rds.amazonaws.com:5432/BigData"
config = {"user":"postgres",
"password": "password",
"driver":"org.postgresql.Driver"}
# + id="CfSriAMytYw9" outputId="c4b709b5-0dfb-46e1-a1b4-a5373e4509bb" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Writing tables in RDS
# review_id_table.write.jdbc(url=jdbc_url, table='review_id_table', mode=mode, properties=config)
products.write.jdbc(url=jdbc_url, table='products', mode=mode, properties=config)
# customers.write.jdbc(url=jdbc_url, table='customers', mode=mode, properties=config)
# vine_table.write.jdbc(url=jdbc_url, table='vine_table', mode=mode, properties=config)
# + id="xLM68eBguP08"
| sports.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><center>IAML LAB 1: NAIVE BAYES AND DECISION TREE CLASSIFICATION</h1></center>
# +
import os
import itertools
from scipy.io import arff
import pandas as pd
import numpy as np
from utils import numeric2bool, str2cat, object2str, update_plot_params
from sklearn.naive_bayes import BernoulliNB
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn.metrics import confusion_matrix, roc_auc_score, accuracy_score
import plotly.plotly as py
import plotly.graph_objs as go
from plotly.tools import set_credentials_file
from plotly.offline import init_notebook_mode
set_credentials_file(username='mynameistony', api_key='M<KEY>')
init_notebook_mode(connected=True)
import matplotlib.pyplot as mpl_plt
import seaborn as sns
sns.set(palette="Set2")
# -
update_plot_params()
# #### Part 1
# Load data in .arff to pd.DataFrame (1.2)
data, _ = arff.loadarff('./data/raw/spambase.arff.txt')
spambase = pd.DataFrame(data)
spambase.head()
# Drop columns (1.4)
spambase.drop(['capital_run_length_average', 'capital_run_length_longest', 'capital_run_length_total'], 1, inplace=True)
# Convert column types (1.5)
spambase = numeric2bool(spambase)
spambase = object2str(spambase, False, True)
spambase.head()
# Save data with bag of words (1.6)
spambase.to_csv('./data/preprocessed/spambase_bag_of_words.csv', index=False)
# #### Part 2 (unfinished)
# Create labels (classes) to fit Naive Bayes (train and test sets in fact)
y = spambase.pop('is_spam')
X = spambase
# Fit Naive Bayes for Bernoulli destribution (because all columns are boolean now)
clf = BernoulliNB(fit_prior=False)
clf.fit(X, y)
# Predict for random sample
clf.predict(X.loc[5:6])
# Compare execution time on whole dataset (4601 obs.) and sample (2000 obs.)
# %%timeit
clf.fit(X.loc[:1999], y[:2000])
# %%timeit
clf.fit(X, y)
# #### Part 3
# Load data in .arff to pd.DataFrame (3.1)
data, _ = arff.loadarff('./data/raw/credit.arff.txt')
credit = pd.DataFrame(data)
credit.head()
# Preprocessing
credit = object2str(credit)
credit, cat_labels = str2cat(credit)
credit.head()
# Plot Age and Duration (3.2)
def age_duration_plt(df, age_column, duration_column):
plt = [go.Scatter(x=credit.index, y=df[age_column], name='Age'),
go.Scatter(x=credit.index, y=df[duration_column], name='Duration')]
return plt
plt = age_duration_plt(credit, 'Age', 'Duration')
py.iplot(plt)
# Drop observations where Age < 0 (3.4)
credit = credit.loc[credit.Age > 0]
plt = age_duration_plt(credit, 'Age', 'Duration')
py.iplot(plt)
y = credit.pop('Approve')
X = credit
# Check preprocessing if we have a binary classification
assert len(y.value_counts()) == 2
# Split on train and test sets (3.5)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1)
# Define built-in Decision Tree Classifier which is more advanced version of C4.5 algorithm (3.6)
model = DecisionTreeClassifier()
# Train classifier and predict (3.7)
clf = model.fit(X_train, y_train)
y_pred = clf.predict(X_test)
conf_matrix = confusion_matrix(y_test, y_pred)
sns.heatmap(conf_matrix, annot=True, fmt="d")
mpl_plt.ylabel('True label', size=20)
mpl_plt.xlabel('Predicted label', size=20)
mpl_plt.title('Confusion matrix', size=26)
mpl_plt.show()
# Due to an unbalanced dataset, the best metric is ROC_AUC Curve (it will be much more accurate than accuracy)
print("ROC_AUC score: {:.4f}".format(roc_auc_score(y_test, y_pred)))
print("Accuracy score: {:.4f}".format(accuracy_score(y_test, y_pred)))
# Print feature importances
def print_feature_importance(df, clf):
importances = clf.feature_importances_
indices = np.argsort(importances)[::-1]
for f in range(X.shape[1]):
print("{}. {} ({:.4f})".format(f + 1, df.columns[f], importances[indices[f]]))
print_feature_importance(X, clf)
# Create decision tree graph and visualize it
graph_name = 'credit_decision_tree'
export_graphviz(clf, out_file='./plots/{}.dot'.format(graph_name)) # save graph in .dot format
os.system("dot -Tpng ./plots/{}.dot -o ./plots/{}.png".format(graph_name, graph_name)) # convert graph to png
os.remove("./plots/{}.dot".format(graph_name)) # remove .dot file
os.system("eog ./plots/{}.png".format(graph_name)) # open graph
| Introductory Applied Machine Learning/Problems/Problem 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas Week Day 1 All Activities
# # Instructor Turn Activity 1: Jupyter Introduction
# ====================================
# Activity 1 Instructor
# Running the basic "Hello World" code
# ====================================
hello = "Hello World"
print(hello)
# Doing simple math
4 + 4
# Storing results in variable
a = 5
# Using those variables elsewhere in the code
a
# Variables will hold the values most recently run
# This means that, if we run the code above, it will now print 2
a = 2
# # Students Turn Activity 2: Netflix Remix
#
# ## Instructions
#
# * Using `Netflix.py` as a jumping off point, convert the application so that it runs properly within a Jupyter Notebook.
#
# * Make sure to have the application print out the user's input, the path to `Netflix_Ratings.csv`, and the final rating/review for the film in different cells.
#
# ## Bonus
#
# * Go through any of the activities from last week and attempt to convert them to run within a Jupyter Notebook. While doing this, try to split up the code into cells and print out the outputs.
# ====================================
# Activity 2 Student
# ====================================
# Code Here
#Modules
# Prompt user for video lookup
# +
# Set path for file
# Set variable to check if we found the video
# -
# # Instructor Turn Activity 3: Creating Data Frames
# Dependencies importing pandas and storing it to a variable called pd
import pandas as pd
# Then create a Pandas Series from a raw list
data_series = pd.Series(["UC Irvine", "UCLA", "UC Berkley", "UC Riverside", "UC Davis"])
data_series
# Convert a list of dictionarys into a dataframe
states_dicts = [{"STATE": "California", "ABBREVIATION" : "CA"},
{"STATE": "New York", "ABBREVIATION": "NY"}]
dataframe_states = pd.DataFrame(states_dicts)
dataframe_states
# Convert a single dictionary containing list into dataframe
dataframe = pd.DataFrame(
{
"Dynasty": ["Early Dynastic Period", "Old Kingdom"],
"Pharoh": ["Thinis", "Memphis"]
}
)
dataframe
# # Students Turn Activity 4: Data-Frame Shop
#
# ## Instructions
#
# * Create a DataFrame for a frame shop that contains three columns - "Frame", "Price", and "Sales" - and has five rows of data stored within it.
#
# * Using an alternate method from that used before, create a DataFrame for an art gallery that contains three columns - "Painting", "Price", and "Popularity" - and has four rows of data stored within it.
#
# ## Bonus
#
# * Once both of the DataFrames have been created, discuss with those around you which method you prefer to use and why.
# Import Dependencies
# DataFrame should have 3 columns: Frame, Price, and Sales AND 5 rows of data# DataF
# Use a different method of creating DataFrames to
# Create a DataFrame for an art gallery that contains three columns - "Painting", "Price", and "Popularity"
# and has 4 rows of data
# # Instructor Turn Activity 5: Data-Frame Functions
# Dependencies
import os
import pandas as pd
# Save path to data set in a variable
data_file = os.path.join("Resources", "dataSet.csv")
print(data_file)
# Use Pandas to read data
data_file_pd = pd.read_csv(data_file)
data_file_pd.head()
# Dislay a statistical overview of the DataFrame
data_file_pd.describe()
# Reference a single column within a DataFrame
data_file_pd["Amount"].head()
# Reference multiple columns within a DataFrame
data_file_pd[["Amount", "Gender"]].head()
# The mean method averages the series
average = data_file_pd["Amount"].mean()
average
# The sum method adds every entry in the series
total = data_file_pd["Amount"].sum()
total
# The unique method shows every element of the series that appears only once
unique = data_file_pd["Last Name"].unique()
unique
# The value_counts method counts unique values in a column
count = data_file_pd["Gender"].value_counts()
count
# +
# Calculations can also be performed on Series and added into DataFrames as new columns
thousands_of_dollars = data_file_pd["Amount"]/1000
data_file_pd["Thousands of Dollars"] = thousands_of_dollars
data_file_pd.head()
# -
# # Students Turn: Activity 6 Training Grounds
#
# ## Instructions
#
# * Using the DataFrame provided, perform all of the following actions...
#
# * Provide a simple, analytical overview of the dataset's numeric columns
#
# * Collect all of the names of the trainers within the dataset
#
# * Figure out how many students each trainer has
#
# * Find the average weight of the students at the gym
#
# * Find the combined weight of all of the students at the gym
#
# * Convert the "Membership (Days)" column into weeks and then add this new series into the DataFrame
# ====================================
# Activity 6 Student
# ====================================
# Import Dependencies
# A gigantic DataFrame of individuals' names, their trainers, their weight, and their days as gym members
# Collecting a summary of all numeric data
# Finding the names of the trainers
# Finding how many students each trainer has
# Finding the average weight of all students
# Finding the combined weight of all students
# Converting the membership days into weeks and then adding a column to the DataFrame
# # Instructor Turn: Activity 7 Column Manipulation
# Import Dependencies
import pandas as pd
# A gigantic DataFrame of individuals' names, their trainers, their weight, and their days as gym members
training_data = pd.DataFrame({
"Name":["<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","Junita Dogan","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","Evel<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","Astrid Duffer","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","China Truett","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","<NAME>","P<NAME>","<NAME>"],
"Trainer":['<NAME>','<NAME>','<NAME>','<NAME>','Blanch Victoria','Ald<NAME>ler','Ald<NAME>ler','<NAME>','Junie Ritenour','<NAME>','<NAME>','<NAME>','Aldo Byler','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','Aldo Byler','Brittani Brin','<NAME>','Phyliss Houk','<NAME>avory','Junie Ritenour','Aldo Byler','Calvin North','Brittani Brin','Junie Ritenour','Blanch Victoria','Brittani Brin','Bettyann Savory','Blanch Victoria','<NAME>','Bettyann Savory','Blanch Victoria','Brittani Brin','Junie Ritenour','Pa Dargan','Gordon Perrine','Phyliss Houk','Pa Dargan','<NAME>','Ph<NAME>k','Ph<NAME>k','Calvin North','Williams Camire','Brittani Brin','Gordon Perrine','Bettyann Savory','Bettyann Savory','<NAME>','Phyliss Houk','Barton Stecklein','Blanch Victoria','<NAME>','Phyliss Houk','Blanch Victoria','<NAME>','Harland Coolidge','Calvin North','Bettyann Savory','Phyliss Houk','Bettyann Savory','Harland Coolidge','Gordon Perrine','Junie Ritenour','Harland Coolidge','Blanch Victoria','<NAME>io','<NAME>','Aldo Byler','Bettyann Savory','Gordon Perrine','Bettyann Savory','Barton Stecklein','Harland Coolidge','Aldo Byler','Aldo Byler','<NAME>an','Junie Ritenour','Brittani Brin','Junie Ritenour','Gordon Perrine','<NAME>','<NAME>io','<NAME>io','Bet<NAME>ory','Brittani Brin','Aldo Byler','Phyliss Houk','Blanch Victoria','Pa Dargan','Phyliss Houk','Brittani Brin','Barton Stecklein','<NAME>','<NAME>','<NAME>','Gordon Perrine','Blanch Victoria','Junie Ritenour','Phyliss Houk','<NAME>','Williams Camire','Harland Coolidge','Williams Camire','Aldo Byler','Harland Coolidge','Gordon Perrine','Brittani Brin','<NAME>','Calvin North','Phyliss Houk','Brittani Brin','Aldo Byler','Bettyann Savory','Brittani Brin','Gordon Perrine','Calvin North','Harland Coolidge','Coleman Dunmire','Harland Coolidge','Aldo Byler','Junie Ritenour','Blanch Victoria','Harland Coolidge','Blanch Victoria','Junie Ritenour','Harland Coolidge','Junie Ritenour','<NAME>','B<NAME>','<NAME>','Williams Camire','Junie Ritenour','B<NAME>in','Cal<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','B<NAME>','<NAME>','<NAME>','<NAME>','Junie Ritenour','<NAME>','Williams Camire','<NAME>','Calvin North','Williams Camire','<NAME>','Ald<NAME>ler','<NAME>','<NAME>','Blanch Victoria','<NAME>','<NAME>','Har<NAME>idge','<NAME>','Ph<NAME>','<NAME>','<NAME>','<NAME>','Harland Coolidge','Junie Ritenour','<NAME>','<NAME>','Blanch Victoria','Williams Camire','Phyliss Houk','Phyliss Houk','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','Blanch Victoria','Co<NAME>','<NAME>','<NAME>','Aldo Byler','Aldo Byler','<NAME>','Williams Camire','Phyliss Houk','Aldo Byler','Williams Camire','Aldo Byler','Williams Camire','Co<NAME>','Phyliss Houk'],
"Weight":[128,180,193,177,237,166,224,208,177,241,114,161,162,151,220,142,193,193,124,130,132,141,190,239,213,131,172,127,184,157,215,122,181,240,218,205,239,217,234,158,180,131,194,171,177,110,117,114,217,123,248,189,198,127,182,121,224,111,151,170,188,150,137,231,222,186,139,175,178,246,150,154,129,216,144,198,228,183,173,129,157,199,186,232,172,157,246,239,214,161,132,208,187,224,164,177,175,224,219,235,112,241,243,179,208,196,131,207,182,233,191,162,173,197,190,182,231,196,196,143,250,174,138,135,164,204,235,192,114,179,215,127,185,213,250,213,153,217,176,190,119,167,118,208,113,206,200,236,159,218,168,159,156,183,121,203,215,209,179,219,174,220,129,188,217,250,166,157,112,236,182,144,189,243,238,147,165,115,160,134,245,174,238,157,150,184,174,134,134,248,199,165,117,119,162,112,170,224,247,217],
"Membership(Days)":[52,70,148,124,186,157,127,155,37,185,158,129,93,69,124,13,76,153,164,161,48,121,167,69,39,163,7,34,176,169,108,162,195,86,155,77,197,200,80,142,179,67,58,145,188,147,125,15,13,173,125,4,61,29,132,110,62,137,197,135,162,174,32,151,149,65,18,42,63,62,104,200,189,40,38,199,1,12,8,2,195,30,7,72,130,144,2,34,200,143,43,196,22,115,171,54,143,59,14,52,109,115,187,185,26,19,178,18,120,169,45,52,130,69,168,178,96,22,78,152,39,51,118,130,60,156,108,69,103,158,165,142,86,91,117,77,57,169,86,188,97,111,22,83,81,177,163,35,12,164,21,181,171,138,22,107,58,51,38,128,19,193,157,13,104,89,13,10,26,190,179,101,7,159,100,49,120,109,56,199,51,108,47,171,69,162,74,119,148,88,32,159,65,146,140,171,88,18,59,13]
})
training_data.head(10)
# Collecting a list of all columns within the DataFrame
training_data.columns
# Reorganizing the columns using double brackets
organized_df = training_data[["Name","Trainer","Weight","Membership(Days)"]]
organized_df.head()
# Using .rename(columns={}) in order to rename columns
renamed_df = organized_df.rename(columns={"Membership(Days)":"Membership in Days", "Weight":"Weight in Pounds"})
renamed_df.head()
# # Students Turn: Activity 8 Hey, Arnold!
#
# * This assignment will give you experience creating DataFrames from scratch.
#
# * You will create a pandas DataFrame of characters from this TV show: [https://en.wikipedia.org/wiki/Hey_Arnold!](https://en.wikipedia.org/wiki/Hey_Arnold!)
#
# ## Instructions
#
# 1. First, use Pandas to create a DataFrame with the following columns and values:
#
# * `Character_in_show`: Arnold, Gerald, Helga, Phoebe, Harold, Eugene
#
# * `color_of_hair`: blonde, black, blonde, black, unknown, red
#
# * `Height`: average, tallish, tallish, short, tall, short
#
# * `Football_Shaped_Head`: True, False, False, False, False, False
#
# 2. You'll note that the above column names are inconsistent and difficult to work with. Rename them to the following, respectively:
#
# * `Character`, `Hair Color`, `Height`, `Football Head`
#
# 3. Next, create a new table that contains all of the columns in the following order...
#
# * `Character`, `Football Head`, `Hair Color`, `Height`
#
# 4. Finally, save the file in Excel format.
# import dependencies
# Create a data frame with given columns and value
# Rename columns to clean up the look
# Organize columns into a more logical order
# Export new data
# # Instructor Turn: Activity 9 Reading and Writing CSVs
# Dependencies
import pandas as pd
# Store filepath in a variable
file_one = "Resources/DataOne.csv"
# Read our Data file with the pandas library
# Not every CSV requires an encoding, but be aware this can come up
file_one_df = pd.read_csv(file_one, encoding="ISO-8859-1")
# Show just the header
file_one_df.head()
# Show a single column
file_one_df["first_name"].head()
# Show mulitple specific columns--note the extra brackets
file_one_df[["first_name", "email"]].head()
# Head does not change the DataFrame--it only displays it
file_one_df.head()
# Export file as a CSV, without the Pandas index, but with the header
file_one_df.to_csv("Output/fileOne.csv", index=False, header=True)
# # Students Turn Activity 10: GoodReads - Part 1
#
# ## Instructions
#
# * Read in the GoodReads CSV using Pandas
#
# * Remove unnecessary columns from the DataFrame so that only the following columns remain: `isbn`, `original_publication_year`, `original_title`, `authors`, `ratings_1`, `ratings_2`, `ratings_3`, `ratings_4`, and `ratings_5`
#
# * Rename the columns to the following: `ISBN`, `Publication Year`, `Original Title`, `Authors`, `One Star Reviews`, `Two Star Reviews`, `Three Star Reviews`, `Four Star Reviews`, and `Five Star Reviews`
#
# * Write the DataFrame into a new CSV file
#
# ## Hints
#
# * The base CSV file uses UTF-8 encoding. Trying to read in the file using some other kind of encoding could lead to strange characters appearing within the dataset.
# Import Dependencies
# +
# Make a reference to the books.csv file path
# Import the books.csv file as a DataFrame
# -
# Remove unecessary columns from the DataFrame and save the new DataFrame
# Only keep: "isbn", "original_publication_year", "original_title", "authors",
# "ratings_1", "ratings_2", "ratings_3", "ratings_4", "ratings_5"
# Rename the headers to be more explanatory
# Push the remade DataFrame to a new CSV file
# # Students Turn Activity 11: GoodReads - Part II
#
# ## Instructions
#
# * Using the modified DataFrame that was created earlier, create a summary table for the dataset that includes the following pieces of information...
#
# * The count of unique authors within the DataFrame
#
# * The year of the earliest published book in the DataFrame
#
# * The year of the latest published book in the DataFrame
#
# * The total number of reviews within the DataFrame
# Import Dependencies
# +
# File to Load
# Read the modified GoodReads csv and store into Pandas DataFrame
# +
# Calculate the number of unique authors in the DataFrame
# Calculate the earliest/latest year a book was published
# Calculate the total reviews for the entire dataset
# -
# Place all of the data found into a summary DataFrame
| Activities Week 4 (pandas)/Pandas_Week/.ipynb_checkpoints/Day1-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 第1章: 数と式
#
# - - -
#
# ## 1節: 式の展開と因数分解
#
# 問1〜5は単純に同一の式の内の同類項を揃えることを扱っているため, ここでは扱われない
import numpy as np
# - - -
#
# ### 問6
#
# $$
# [a, b, c] = a \times x + b \times y + c \times z \space \space (a, b, cは全て定数)
# $$
def prob6(A: np.poly1d, B: np.poly1d):
print("-"*40)
print(f"A + B = \n{(A + B).c}")
print(f"A - B = \n{(A - B).c}")
print(f"3A = \n{(3*A).c}")
print("-"*40)
# +
# (1)
## [x, y, z]
A = np.poly1d([1, -2, 4])
B = np.poly1d([3, 1, -2])
prob6(A, B)
# (2)
## [x^2, x, x^0]
A = np.poly1d([2, -3, -1])
B = np.poly1d([-3, 7, -2])
prob6(A, B)
# (3)
## [x^3, x^2, x, x^0]
A = np.poly1d([1, 0, -2, -3])
B = np.poly1d([-3, 2, -5, -1])
prob6(A, B)
# (4)
## [x^2, xy, y^2]
A = np.poly1d([2, -1, -1])
B = np.poly1d([1, 1, -3])
prob6(A, B)
# -
# - - -
#
# ### 問7
#
# $$
# A = 3x^2 - x + 1 \\
# B = -x^2 + 4x - 3
# $$
#
# これらを次の式で計算せよ
# +
def prob7(prob: str, ans: np.poly1d):
print('-'*40)
print(f"{prob} = \n{ans.c}")
print('-'*40)
A = np.poly1d([3, -1, 1])
B = np.poly1d([-1, 4, -3])
# +
# (1)
prob7("3A - 2B", 3*A - 2*B)
# (2)
prob7("2B - (B - 2A)", 2*B - (B - 2*A))
# (3)
prob7("2(A - 3B) - 3(A + B)", 2*(A - 3*B) - 3*(A + B))
# -
# - - -
#
# ### 問8
#
# $$
# A = x^2 - 2xy - 3y^2 \\
# B = -x^2 + 3xy - 2y^2 \\
# C = x^2 - xy + y^2
# $$
#
# これらを次の式で計算せよ
prob8 = prob7
A = np.poly1d([1, -2, -3])
B = np.poly1d([-1, 3, -2])
C = np.poly1d([1, -1, 1])
# +
# (1)
prob8("2A + B - C", 2*A+B-C)
# (2)
prob8("A + 2B - 2(B - C)", A+2*B-2*(B-C))
# (3)
prob8("3(A - 2B) - 2(C - 3B)", 3*(A-2*B)-2*(C-3*B))
# -
# - - -
#
# ### 問9
#
# $$
# A = 3x^2 + 2x - 1 \\
# B = -x^2 + 2x + 4
# $$
#
# のとき, 次の式を満たす整式_X_を求めよ
# +
def prob9(X_1: list, X_2: list): # [X_scala, np.array([x^2, x, x^0])]
# converting [int, np.array] into np.array(list of ints)
tmpX_1 = [X_1[0]]
for i in X_1[1]: tmpX_1.append(i)
tmpX_1 = np.array(tmpX_1)
tmpX_2 = [X_2[0]]
for i in X_2[1]: tmpX_2.append(i)
tmpX_2 = np.array(tmpX_2)
# ======================================================
ans = tmpX_1 - tmpX_2
ans = 1/ans[0]*ans[1:] if abs(ans[0]) > 1 else ans[1:]
ans = np.poly1d(ans)
print(f"{'-'*40}\nX = \n{ans}\n{'-'*40}")
A = np.array([3, 2, -1])
B = np.array([-1, 2, 4])
# +
# (1)
## X-2A=2X-B
prob9([1, -2*A], [2, -1*B])
# (2)
## 3X-2B=X+4A
prob9([3, -2*B], [1, 4*A])
# -
# - - -
#
# ## 2節: 整式の乗法
#
# 問10〜13は単項式, 多項式同士の展開であるため, ここでは扱わない
#
# ### Pythonを用いた単項式・多項式同士の演算方法
# - Numpyを用いて単項式, 多項式の演算を行う際はpoly1dメソッドを用いると乗算を行うことができる
# - しかし, 出力結果が$$ax^n + bx^{n-1} + \dots + cx^0$$のようになってしまう。<br/>これは複数種の項を含む式 (αx+βy+γzなど) を計算する際には向かない。そのため, poly1dメソッドのあとに.cをつけることで$$[a, b, \dots, c]$$のような出力結果となり, 項を出力しなくすることができる
#
# - - -
#
# ### 問14
#
# 次の式を計算せよ。
def prob14(prob: str, ans: np.poly1d):
print('-'*40)
print(f"{prob} =\n{ans.c}")
print('-'*40)
# +
# (1)
## (x+y)^2 + (x-y)^2
prob14_1 = np.poly1d([1, 1])**2 + np.poly1d([1, -1])**2
prob14("(x + y)^2 + (x - y)^2", prob14_1)
# (2)
## (x + y)^2 - (x - y)^2
prob14_2 = np.poly1d([1, 1])**2 - np.poly1d([1, -1])**2
prob14("(x + y)^2 - (x - y)^2", prob14_2)
# (3)
## (x - 1)(x + 2) - x(x + 1)
prob14_3 = np.poly1d([1, -1])*np.poly1d([1, 2]) - np.poly1d([1, 0])*np.poly1d([1, 1])
prob14("(x - 1)(x + 2) - x(x + 1)", prob14_3)
# (4)
## (a + 2b)^2 - (a - 2b)(a + 2b)
prob14_4 = np.poly1d([1, 2])**2 - np.poly1d([1, -2])*np.poly1d([1, 2])
prob14("(a + 2b)^2 - (a - 2b)(a + 2b)", prob14_4)
# -
| chapters/algebra_1A/chapter1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome to Binder
#
# This is where will do all our Python, Shell and Git live coding.
#
#
# ## Jupyter Lab
#
# Let's quickly familiarise ourselves with the enironment ...
#
# - the overal environment (ie your entire browser tab) is called:
#
# *Jupyter Lab*
#
# it contains menus, tabs, toolbars and a file browser
#
# - Jupyter Lab allows you to *launch* files and application into the *Work Area*. Right now you probably have two tabs in the *Work Area* - this document an another tab called *Launcher*.
#
#
#
# ## Juptyer Notebooks
#
# - this document is document is called an:
#
# *Interactive Python Notebook* (or Notebook for short)
#
# Notebooks are a documents (files) made up of a sequence of *cells* that contain either code (python in our case) or documentation (text, or formatted text called *markdown*).
#
# ### Cells
#
# The three types of Cells are:
#
# - *Markdown* - formatted text like this cell (with *italics*, **bold** and tables etc ...)
# - *Raw* - like the following cell, and
# - *Code* - (in our case) python code
#
# Cells can be modified by simply clicking inside them and typing.
#
# See if you can change the cell below by replacing *boring* with something more exciting.
# + active=""
# RAW CELL
#
# This Cell is a 'Raw' Cell - it's just boring text.
# -
# #### Executing Cells
#
# Both *markdown* and *code* cells can be excuted,
#
# Excuting a *markdown* causes the *formatted* version of the cell to be displayed. Excuting a *code* cell causes the code to run and any results are displayed below the cell.
#
# Any cell can be executed by pressing the play icon at the top of the document while the cell is highlighted.
#
# You can also press **CTL-ENTER** to excute the active cell.
#
# Go ahead and make some more changes to the cells above and execute them - what happens when you execute a *Raw* cell ?
# #### Adding a Removing Cells
#
# You can use the `+` (plus icon) at the top of the docuement to add a new cell, and the cell type drop down the change the type.
#
# You can also use the A key to add cell *above* the current cell and the B key to add *below* the current cell.
#
#
# Now add a couple of cell of your own ...
# + active=""
#
#
# see if you can delete this cell - tip, checkout the *Edit* menu ... or even try right-click
#
#
# -
# #### Code Cells
#
# Code cells allow us to write (in our case Python) and run our code and see the results right inside the notebook.
#
# The next cell is a code cell that contains Python code to add 4 numbers.
#
# Try executing the cell and if get the right result, try some more/different numbers
#
1 + 2 + 3 + 4
# ## Let's save our work so for and Push to our changes to GitHub
#
# ### Saving
#
# By pressing the save icon on the document (or File -> Save Notebook) we can save our work to our Binder environment.
#
#
# ### But what about Version Control and Git (wasn't that in the Workshop syllabus)
#
# Since our binder environment will dissappear when we are no longer using is, we need cause our new version to be saved some where permanent.
#
# Luckly we have a GitHub respository already connected to our current environment - however there are couple steps required to make our GitHub repository match the copy of the respository inside our Binder environment.
#
# #### Git likes to know how you are
#
# ... otherwise it keeps complaining it cannot track who made what commits (changes).
#
# To tell Git who you are, we need to do the following:
#
# - Launch a Terminal sesson (File -> New -> Terminal, or you can use the *Laucher* tab
# - At the command prompt, type: `git-setup`
#
# This operation only needs to be done once per binder session.
#
# #### Add your changed files to git's list of files to track
#
# - at the same terminal prompt type: `git add .`
#
# #### Tell Git to commit (record) this state as a version
#
# - at the same terminal prompt type: `git commit -m "changes made inside binder"`
#
# at this point git has added an additional version of your files to your repository inside your curren Binder environment. However, your repository on GitHub remains unchanges (you might like to go check).
#
# #### Tell Git to push the new commit (version) to GitHub
#
# - again at the same prompt type: `git push`
#
# once you supply the correct GitHub usename and password, all your changes will be pushed.
#
# Go check out your respository on github.com ...
#
#
| welcome.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 99} colab_type="code" id="EUhlAxLGezgk" outputId="019a1063-6e0f-4f17-fc40-8bb0819c1f48"
import tensorflow.keras.layers as Layers
import tensorflow.keras.activations as Actications
import tensorflow.keras.models as Models
import tensorflow.keras.optimizers as Optimizer
import tensorflow.keras.metrics as Metrics
import tensorflow.keras.utils as Utils
from keras.utils.vis_utils import model_to_dot
from keras.utils.vis_utils import plot_model
import os
import matplotlib.pyplot as plot
import cv2
import numpy as np
from sklearn.utils import shuffle
from sklearn.metrics import confusion_matrix as CM
from random import randint
from IPython.display import SVG
import matplotlib.gridspec as gridspec
# + colab={} colab_type="code" id="Z3Ryd55gezgv"
Images = np.load('./dataset/Image_1.npy') #converting the list of images to numpy array.
Labels = np.load('./dataset/Label_1.npy')
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="I_SnPdLNezgx" outputId="9281c49c-c2e0-4f7c-d595-a35bc84d6fb6"
print("Shape of Images:",Images.shape)
print("Shape of Labels:",Labels.shape)
# -
from sklearn.model_selection import train_test_split
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="tacXBg_rezgz" outputId="97048603-1a85-4654-a87f-edd70d498bf1"
len(Images)
# -
names = {0: 'Tomato_H', 1: 'Tomato_D01', 2: 'Tomato_D04', 3: 'Tomato_D05',
4: 'Tomato_D07', 5: 'Tomato_D08', 6: 'Tomato_D09', 7: 'Tomato_P03',
8: 'Tomato_P05'}
X_train, X_test ,y_train, y_test = train_test_split(Images, Labels,test_size=0.1, stratify=Labels, random_state=1, shuffle=True)
print(X_train.shape,X_test.shape,y_train.shape,y_test.shape)
np.bincount(y_train),np.bincount(y_test)
y_test[0:20]
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="uPTxW0Huezg4" outputId="8cba049b-a0ab-4ea8-e051-e2b2506988cc"
import tensorflow as tf
model = Models.Sequential()
model.add(Layers.Conv2D(200,kernel_size=(3,3),activation='relu',input_shape=(128,128,3),
kernel_regularizer = tf.keras.regularizers.l2(0.001),
kernel_initializer=tf.contrib.layers.variance_scaling_initializer(uniform=False)))
model.add(Layers.BatchNormalization())
model.add(Layers.Conv2D(180,kernel_size=(3,3), activation='relu',))
model.add(Layers.MaxPool2D(5,5))
model.add(Layers.BatchNormalization())
model.add(Layers.Conv2D(180,kernel_size=(3,3 ), activation='relu'))
model.add(Layers.BatchNormalization())
model.add(Layers.Conv2D(140,kernel_size=(3,3), activation='relu'))
model.add(Layers.BatchNormalization())
model.add(Layers.Conv2D(100,kernel_size=(3,3), activation='relu'))
model.add(Layers.BatchNormalization())
model.add(Layers.Conv2D(50,kernel_size=(3,3), activation='relu'))
model.add(Layers.BatchNormalization())
model.add(Layers.MaxPool2D(5,5))
model.add(Layers.Flatten())
model.add(Layers.Dense(180, activation='relu'))
model.add(Layers.Dense(100, activation='relu'))
model.add(Layers.Dense(50, activation='relu'))
model.add(Layers.Dropout(rate=0.3))
model.add(Layers.Dense(9,activation='softmax'))
model.compile(optimizer=Optimizer.Adam(lr=0.001),loss='sparse_categorical_crossentropy',metrics=['accuracy'])
model.summary()
#SVG(model_to_dot(model).create(prog='dot', format='svg'))
#Utils.plot_model(model,to_file='model.png',show_shapes=True)
# +
from sklearn.utils import class_weight
class_weights = class_weight.compute_class_weight('balanced',
np.unique(y_train),
y_train)
class_weights
# +
class_weight = {}
for i in range(9):
class_weight[i] = class_weights[i]
class_weight
# -
class_weight = [np.sqrt(np.sqrt(i)) for i in class_weights]
class_weight
# + colab={"base_uri": "https://localhost:8080/", "height": 702} colab_type="code" id="bLP9hwRTtQkf" outputId="5d35a414-7a66-4eb6-a88b-7c85041c8461"
fig = plot.figure(figsize=(30, 30))
outer = gridspec.GridSpec(5, 5, wspace=0.2, hspace=0.2)
for i in range(10):
inner = gridspec.GridSpecFromSubplotSpec(2, 1,subplot_spec=outer[i], wspace=0.1, hspace=0.1)
rnd_number = randint(0,len(test_images)) # 0~18 중 하나를 뽑는 난수 생성
pred_image = np.array([test_images[rnd_number]]) # test_images[random] 하나 뽑아서 넘파이로 변경\
result = ['H', 'D01', 'D04', 'D05', 'D07', 'D08', 'D09', 'P03', 'P05'][test_labels[rnd_number]]
pred_class = get_classlabel(model.predict_classes(pred_image)[0]) # 추측해서 가장 높은 값이 나온 거 위치 반환
pred_prob = model.predict(pred_image).reshape(9)
for j in range(2):
if (j%2) == 0:
ax = plot.Subplot(fig, inner[j])
ax.imshow(pred_image[0])
ax.set_title('result = {}, predict = {}'.format(result, pred_class))
ax.set_xticks([])
ax.set_yticks([])
fig.add_subplot(ax)
else:
ax = plot.Subplot(fig, inner[j])
ax.bar(['H', 'D01', 'D04', 'D05', 'D07', 'D08', 'D09', 'P03', 'P05'],pred_prob)
fig.add_subplot(ax)
fig.show()
# -
X_train_1, X_val, y_train_1, y_val = train_test_split(X_train, y_train, test_size=0.1, stratify=y_train, shuffle=True, random_state=1 )
trained = model.fit(X_train_1, y_train_1, epochs=60, batch_size=64, validation_data=(X_val,y_val), \
class_weight = class_weight)
# +
plot.plot(trained.history['acc'])
plot.plot(trained.history['val_acc'])
plot.title('Model accuracy')
plot.ylabel('Accuracy')
plot.xlabel('Epoch')
plot.legend(['Train', 'Test'], loc='upper left')
plot.show()
plot.plot(trained.history['loss'])
plot.plot(trained.history['val_loss'])
plot.title('Model loss')
plot.ylabel('Loss')
plot.xlabel('Epoch')
plot.legend(['Train', 'Test'], loc='upper left')
plot.show()
# -
test_images, test_labels = shuffle_Images[16063:].copy(),\
shuffle_Labels[16063:].copy()
# test_images,test_labels = get_images('./test/')
# test_images = np.array(test_images)
# test_labels = np.array(test_labels)
model.evaluate(test_images,test_labels, verbose=1)
model.save('90per_2root.h5')
| Project/CNN_tomato/128_128 resize 10per.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + code_folding=[] init_cell=true
# #%reload_ext autoreload
# #%autoreload 2
'''
Ctrl + Enter: run the cell
Enter: switch to edit mode
Up or k: select the previous cell
Down or j: select the next cell
y / m: change the cell type to code cell/Markdown cell
a / b: insert a new cell above/below the current cell
x / c / v: cut/copy/paste the current cell
dd: delete the current cell
z: undo the last delete operation
Shift + =: merge the cell below
h: display the help menu with the list of keyboard shortcuts
'''
import export_env_variables
from export_env_variables import *
from visualizations import *
import defs
from defs import *
import demo_modes
from demo_modes import *
import utils
from utils import *
import write_prototxts
from write_prototxts import *
from uber_script import call_predict_on_val_txts, demo_average_results_plot_averages
import sys
import os
from colorama import Fore, Back, Style
import subprocess
#from IPython.display import Image, display
from skimage import io
# plot inline
# %matplotlib inline
# ----------- GLOBALS -----------------
convolution_vizualizations_img_dir = os.path.join(demo_modes_path, "img_convs")
makedirs_ok(convolution_vizualizations_img_dir)
convolution_vizualizations_img_path = os.path.join(convolution_vizualizations_img_dir, "img_conv.png")
healthy_label = 0
kc_label = 1
ast_label = 2
EXIT = -1
SUCCESS = 0
hkc_label_to_name = {healthy_label: "Healthy", kc_label: "KC", ast_label:"Astigmatism"}
hkc_label_to_color = {healthy_label: Fore.BLUE, kc_label: Fore.RED, ast_label:Fore.YELLOW}
interactive_prediction_user_ans_to_real_world = {1:healthy_label, 2:kc_label, 3:ast_label, 4:EXIT}
FROM_SCRATCH = "from_scratch"
FINE_TUNE = "fine_tune"
train_from_scratch_mode = {"mode": recycle_healthy_vs_kc_vs_cly_from_scratch, "train_method":FROM_SCRATCH}
train_fine_tune_mode = {"mode": recycle_healthy_vs_kc_vs_cly_best_iter_cv, "train_method":FINE_TUNE}
# -------------------------------------------------- FUNCTIONS ---------------------------------------------------
def plot_image_from_file(img_path):
im = io.imread(img_path)
plt.figure(figsize=(8,8))
plt.imshow(im)
plt.xticks(())
plt.yticks(())
plt.show()
# -------------------------------------------------------------------------------------------------------
def plot_filtered_image():
plot_image_from_file(convolution_vizualizations_img_path)
# -------------------------------------------------------------------------------------------------------
# ----------------- Interactive Game ----------------
def interactive_predict_val_txts(mode, statistics):
caffe = import_caffe()
net = caffe.Net(mode.deploy_prototxt,
caffe.TEST,
weights=mode.weights
)
# create transformer for the input called 'data'
transformer = get_caffenet_transformer(caffe, net, mode.mean_binaryproto)
with open(mode.val_txt) as f:
val_images = f.readlines()
imgs_and_labels = set()
for image_name_n_label in val_images:
if len(image_name_n_label.split(' ')) != 2:
continue
image_basename, true_label = image_name_n_label.split(' ')[0], int(image_name_n_label.split(' ')[1])
imgs_and_labels.add((image_basename, true_label))
for image_basename, true_label in imgs_and_labels:
image_file = os.path.join(my_model_data, image_basename) # ALWAYS PREDICT ON ORIGINAL IMAGES
image = caffe.io.load_image(image_file)
# image shape is (3, 256, 256). we want it (3, 227, 227) for caffenet.
# asking about shape[0] and shape[1] because I can't know if the image is (chan, h, w) or (h, w, chan)
if image.shape[0] == TRAINING_IMAGE_SIZE or image.shape[1] == TRAINING_IMAGE_SIZE or image.shape[2] == TRAINING_IMAGE_SIZE:
# I'm cropping the numpy array on the fly so that I don't have to mess with resizing
# the actual images in a separate folder each time.
image = center_crop_image(image, CLASSIFICATION_IMAGE_SIZE, CLASSIFICATION_IMAGE_SIZE)
# show image to user
plt.figure(figsize=(5,5))
plt.imshow(image)
plt.xticks(())
plt.yticks(())
plt.show()
# get user prediction
user_ans = input("What's your diagnosis? \n" +
hkc_label_to_color[healthy_label] + "1: Healthy \n" + \
hkc_label_to_color[kc_label] + "2: KC \n" + \
hkc_label_to_color[ast_label] + "3: Astigmatism \n" + \
Fore.BLACK + "4: exit \nans: ")
print("\n")
user_ans = interactive_prediction_user_ans_to_real_world[user_ans]
if user_ans == EXIT:
return EXIT
# transform image
try:
transformed_image = transformer.preprocess('data', image)
except:
# try to transpose and again
image = image.transpose(2,0,1) # (height, width, chan) -> (chan, height, width)
transformed_image = transformer.preprocess('data', image)
# copy the image data into the memory allocated for the net
net.blobs['data'].data[...] = transformed_image
### perform classification
output = net.forward(start='conv1')
# save conv
feat = net.blobs['conv1'].data[0, :9]
vis_square(feat, filename=convolution_vizualizations_img_path)
output_prob = output['prob'][0] # the output probability vector for the first image in the batch
max_prob = max(output_prob)
predicted_label = output_prob.argmax()
if predicted_label == true_label:
statistics["net_correct"] += 1
if user_ans == true_label:
statistics["user_correct"] += 1
net_accuracy = ((100. * statistics["net_correct"]) / (statistics["current_image_i"]))
user_accuracy = ((100. * statistics["user_correct"]) / (statistics["current_image_i"]))
print("You said the image is " + hkc_label_to_color[user_ans] + hkc_label_to_name[user_ans])
print(Style.RESET_ALL)
print("The net said the image is " + hkc_label_to_color[predicted_label] + hkc_label_to_name[predicted_label] + " with probability {:.0f}%".format(100*max_prob))
print(Style.RESET_ALL)
print("The real label is " + hkc_label_to_color[true_label] + hkc_label_to_name[true_label])
print(Style.RESET_ALL)
print("The image name is " + hkc_label_to_color[true_label] + image_basename.replace("cly_", "ast_"))
print(Style.RESET_ALL)
if user_ans == true_label and predicted_label == true_label:
print(Fore.GREEN + "You're both right!!! :-)")
elif user_ans == true_label:
print(Fore.GREEN + "You're right and net is wrong!!! :-)")
else:
print(Fore.RED + "Net is right...")
print(Style.RESET_ALL)
print(Fore.MAGENTA + "net accuracy is " + "{:.0f}%".format(net_accuracy))
print(Fore.CYAN + "your accuracy is " + "{:.0f}%".format(user_accuracy))
if net_accuracy < user_accuracy:
print(Fore.GREEN + "You're winning!!! :-)")
print(Style.RESET_ALL)
print("\n\n\n----------------------------------------------------------------")
statistics["current_image_i"] += 1
return SUCCESS
# -------------------------------------------------------------------------------------------------------
def interactive_prediction_game():
mode = healthy_vs_kc_vs_cly_best_iter_cv
snapshot_iter = mode.solver_net_parameters.max_iter
statistics = {"current_image_i":1, "user_correct":0, "net_correct":0}
for sub_mode in mode.get_sub_modes():
sub_mode.weights = sub_mode.resume_from_iter(snapshot_iter)
out_code = interactive_predict_val_txts(sub_mode, statistics)
if out_code == EXIT:
break
# -------------------------------------------------------------------------------------------------------
# --------------------- Training ----------------------------
def plot_misclassified_after_training():
with open(misclassified_images_file) as f:
lines = f.readlines()
for i, line in enumerate(lines):
if not ".jpg" in line:
continue
if "False" in line:
print("\n---------------------------------------------------------")
image_name, true_label, predicted_label, max_prob = get_imagename__true_label__pred_label__and_max_prob_from_line(line)
print("Image: " + image_name)
print("Is: " + hkc_label_to_color[true_label] + hkc_label_to_name[true_label] + Style.RESET_ALL)
print("But Was classified as: " + hkc_label_to_color[predicted_label] + \
hkc_label_to_name[predicted_label] + " with probability {:.0f}%".format(100*max_prob))
print(Style.RESET_ALL)
image_file = os.path.join(my_model_data, image_name)
plot_image_from_file(image_file)
# -------------------------------------------------------------------------------------------------------
def print_confusion_matrix():
with open(misclassified_images_file) as f:
lines = f.readlines()
start_printing = False
for line in lines:
line = line.replace("\n", "")
if "confusion matrix" in line.lower():
start_printing = True
if start_printing and line == "":
break
elif start_printing:
print(line)
# -------------------------------------------------------------------------------------------------------
def get_classifications_and_plot(mode):
mode.snapshot_iters = mode.get_snapshots_iters_by_solver_params(include_zero=False)
demo_average_results_plot_averages(mode, mode.snapshot_iters)
# ---------------------------------------------------------------------
def train_predict_from_uber(train_method=FINE_TUNE, print_summary=False, last_set_i='1'):
"""
train_method: string which is "fine_tune" for fine tuning pre trained caffenet or "from_scratch" to train caffenet from scratch
"""
if train_method == FROM_SCRATCH:
mode = recycle_healthy_vs_kc_vs_cly_from_scratch
else:
mode = recycle_healthy_vs_kc_vs_cly_best_iter_cv
clean_mode(mode)
proc = subprocess.Popen(['python','uber_script.py','demo_train_predict', train_method, 'last_set_i', last_set_i],stdout=subprocess.PIPE)
iter = 0
while True:
line = proc.stdout.readline()
if line != '':
if print_summary:
if "Test net output #0: accuracy" not in line and "Test net output #1: loss" not in line and "Train net output #0: loss" not in line:
continue
line = line.partition("]")[2]
if "Test net output #0: accuracy" in line:
print("\nIteration " + str(iter) + ":")
iter += mode.solver_net_parameters.display_iter
#the real code does filtering here
if "Test net output #0: accuracy" in line:
color = Fore.GREEN
elif "Test net output #1: loss" in line:
color = Fore.RED
elif "Train net output #0: loss" in line:
color = Fore.BLUE
else:
color = ""
print (color + line.rstrip() + Style.RESET_ALL)
else:
break
# don't call in subprocess. call here to get plot.
get_classifications_and_plot(mode)
#plot_misclassified_after_training()
# print_confusion_matrix()
# ---------------------------------------------------------------------
# ---------------------------------------------- MAIN ----------------------------------------
mode = recycle_healthy_vs_kc_vs_cly_best_iter_cv
# --- Train and predict ---
#train_predict_from_uber(train_method=FINE_TUNE, print_summary=False, last_set_i='1')
# --- Interactive Prediction Game ---
#interactive_prediction_game()
# --- Plot and get misclassifiedfrom last training w/o trainging ---
#plot_misclassified_after_training()
# --- Get Classifications and Learning Curve plot ---
#get_classifications_and_plot(mode)
# --- Print Confusion Matrix after training ---
print_confusion_matrix()
| scripts/run_demos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dev_pytorch18]
# language: python
# name: conda-env-dev_pytorch18-py
# ---
# + colab={} colab_type="code" id="qdB23YiUA02F"
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# %load_ext line_profiler
# -
import os
# os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID" # see issue #152
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
# +
import sys, os, warnings, time, logging
from functools import partial
import copy
import torch
import torch.nn as nn
import pytorch_lightning as pl
from pytorch_lightning.core.decorators import auto_move_data
from pytorch_lightning.callbacks import EarlyStopping, LearningRateMonitor, ModelCheckpoint
from einops import rearrange, repeat
import numpy as np
import matplotlib.pyplot as plt
import torchvision
from tqdm.autonotebook import tqdm
from lightning_addons.lightprogressbar import LightProgressBar
from lightning_addons.progressplotter import ProgressPlotter
from lightning_addons.progressprinter import ProgressPrinter
warnings.filterwarnings("ignore", category=UserWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
# this makes lightning reports not look like errors
pl._logger.handlers = [logging.StreamHandler(sys.stdout)]
print("python", sys.version, sys.executable)
print("pytorch", torch.__version__)
print("torchvision", torchvision.__version__)
print("pytorch-lightning", pl.__version__)
# -
def to_3channel(image):
return image.repeat((3, 1, 1))
# ## mnist
# +
class MnistDataModule(pl.LightningDataModule):
def __init__(self, batch_size, n_workers, train_fraction=0.7, extra_transforms=None, data_dir="data/mnist"):
super().__init__()
self.batch_size = batch_size
self.n_workers = n_workers
self.train_fraction = train_fraction
self.data_dir = data_dir
transforms = [torchvision.transforms.ToTensor(),]
if extra_transforms is not None:
transforms += extra_transforms
self.transform = torchvision.transforms.Compose(transforms)
def prepare_data(self):
# this is intended to just run the download
torchvision.datasets.MNIST(self.data_dir, download=True, train=True)
torchvision.datasets.MNIST(self.data_dir, download=True, train=False)
def setup(self, stage=None):
# this can potentially be run on different machines and GPUs hence the actual preperation should be done here
if stage == "fit" or stage is None:
trainval_data = torchvision.datasets.MNIST(self.data_dir, train=True, transform=self.transform)
n_trainval = len(trainval_data)
n_train = int(self.train_fraction * n_trainval)
self.train_data = torch.utils.data.Subset(trainval_data, range(0, n_train))
self.val_data = torch.utils.data.Subset(trainval_data, range(n_train, n_trainval))
# if we set this here we get the actual transformed size, which is nice at it does not require prior knowledge as in __init__()
self.dims = tuple(self.train_data[0][0].shape)
if stage == "test" or stage is None:
self.test_data = torchvision.datasets.MNIST(self.data_dir, train=False, transform=self.transform)
self.dims = tuple(self.test_data[0][0].shape)
def train_dataloader(self):
return torch.utils.data.DataLoader(self.train_data, batch_size=self.batch_size, num_workers=self.n_workers, shuffle=True)
def val_dataloader(self):
return torch.utils.data.DataLoader(self.val_data, batch_size=self.batch_size, num_workers=self.n_workers)
def test_dataloader(self):
return torch.utils.data.DataLoader(self.test_data, batch_size=self.batch_size, num_workers=self.n_workers)
output_size = 28
# data standard normalization parameters:
# to compute mean and std
# norm_mean = 0.
# norm_std = 1.
# default
# norm_mean = 0.5
# norm_std = 0.5
# imagenet
# norm_mean = [0.485, 0.456, 0.406]
# norm_std = [0.229, 0.224, 0.225]
# mnist
norm_mean = [0.1311, 0.1311, 0.1311]
norm_std = [0.3087, 0.3087, 0.3087]
# TODO: we can do better by actually computing these values from mnist
extra_transforms = [
torchvision.transforms.Resize(output_size),
to_3channel, # we use 3 channels to directly use (pretrained) ResNets
torchvision.transforms.Normalize(norm_mean, norm_std),
]
mnist_data = MnistDataModule(32, 8, extra_transforms=extra_transforms)
mnist_data.prepare_data()
mnist_data.setup()
mnist_data.size(), len(mnist_data.train_dataloader().dataset), len(mnist_data.val_dataloader().dataset), len(mnist_data.test_dataloader().dataset)
# -
# ## select dataset
data = mnist_data
# ## compute normalization stats
# +
# # compute mean and std etc
# input_key = 0
# def collect_stats(sample):
# image = sample[input_key]
# return image.sum((1, 2)), (image ** 2).sum((1, 2))#, image.mean((1, 2)), image.min((1, 2)), image.max((1, 2))
# all_stats = [collect_stats(sample) for sample in tqdm(data.train_dataloader().dataset)]
# sums, squared_sums= [torch.stack(stats, 0) for stats in (zip(*all_stats))]
# # # , means, mins, maxs
# input_shape = data.train_dataloader().dataset[0][input_key].shape
# input_area = input_shape[1] * input_shape[2]
# ds_mean = sums.mean(0) / input_area
# ds_std = (squared_sums.mean(0) / input_area - ds_mean ** 2).sqrt()
# ds_mean, ds_std
# -
# ## check samples
dataloader = data.train_dataloader()
batch = next(iter(dataloader))
# 0: input, 1: target
n_samples = 8
f, axs = plt.subplots(1, n_samples, figsize=(10, 5))
for i, ax in enumerate(axs):
sample_image = batch[0][i]
sample_target = batch[1][i]
ax.imshow(sample_image.permute(1, 2, 0).mul(torch.as_tensor(norm_std)).add(torch.as_tensor(norm_mean)).clamp(0.0, 1.0))
ax.set_title(f"label: {sample_target.numpy()}");
ax.axis(False);
# ## resnet model
# +
class ResNetModel(pl.LightningModule):
def __init__(self, data_shape, num_classes,
learning_rate, momentum=0.9, weight_decay=5e-4,
pretrained=False, activation="softmax", sigmoid_soft_factor=1.0, scheduler="one_cycle"):
super().__init__()
self.save_hyperparameters()
# torchvision's resnet18 does not allow pretraining for num_classes != 1000
assert pretrained == False
self.resnet = torchvision.models.resnet18(num_classes=num_classes)
# # to use pretraining simply replace last linear layer with one mathcing num_classes
# self.resnet = torchvision.models.resnet18(pretrained=pretrained)
# self.resnet.fc = torch.nn.Linear(512, 10)
# this allows the trainer to show input and output sizes in the report (1 is just a sample batch size)
self.example_input_array = torch.zeros(1, *data_shape)
self.train_acc = pl.metrics.Accuracy(compute_on_step=False)
self.val_acc = pl.metrics.Accuracy(compute_on_step=False)
self.test_acc = pl.metrics.Accuracy(compute_on_step=False)
self.test_confmatrix = pl.metrics.ConfusionMatrix(num_classes, compute_on_step=False)
def configure_optimizers(self):
optimizer = torch.optim.SGD(self.parameters(), lr=self.hparams.learning_rate, momentum=self.hparams.momentum, weight_decay=self.hparams.weight_decay)
schedulers = []
if self.hparams.scheduler == "one_cycle":
steps_per_epoch = int(self.trainer.limit_train_batches * len(self.train_dataloader())) # same as batches per epoch
max_epochs = self.trainer.max_epochs
scheduler = torch.optim.lr_scheduler.OneCycleLR(
optimizer,
max_lr=self.hparams.learning_rate,
epochs=max_epochs,
steps_per_epoch=steps_per_epoch,
anneal_strategy="cos", # can be "linear" or "cos"(default)
three_phase=False,
)
# "interval: step" is required to let the scheduler update per step rather than epoch
schedulers = [{"scheduler": scheduler, "interval": "step", "frequency": 1}]
elif self.hparams.scheduler == "cosine_annealing":
steps_per_epoch = int(self.trainer.limit_train_batches * len(self.train_dataloader())) # same as batches per epoch
max_epochs = self.trainer.max_epochs
max_steps = max_epochs * steps_per_epoch
scheduler = torch.optim.lr_scheduler.CosineAnnealingLR(
optimizer,
T_max=max_steps,
)
schedulers = [{"scheduler": scheduler, "interval": "step", "frequency": 1}]
elif isinstance(self.hparams.scheduler, type) or isinstance(self.hparams.scheduler, partial):
schedulers = [self.hparams.scheduler(optimizer)]
return [optimizer], schedulers
def class_activation(self, x):
if self.hparams.activation == "softmax":
return x.softmax(1)
elif self.hparams.activation == "sigmoid":
return x.sigmoid()
def class_loss(self, y_hat, y):
# y_hat: prediction, y: target
if self.hparams.activation == "softmax":
return torch.nn.functional.cross_entropy(y_hat, y)
elif self.hparams.activation == "sigmoid":
y = y.float()
y = y * self.hparams.sigmoid_soft_factor
y[y == 0] = (1.0 - self.hparams.sigmoid_soft_factor) / (self.hparams.num_classes - 1)
return torch.nn.functional.binary_cross_entropy_with_logits(y_hat, y)
def _forward(self, x):
return self.resnet(x)
def training_step(self, batch, batch_nb):
images, targets = batch
if self.hparams.activation == "sigmoid":
targets = torch.nn.functional.one_hot(targets, num_classes=self.hparams.num_classes)
preds = self._forward(images)
loss = self.class_loss(preds, targets)
self.log("loss", loss)
self.train_acc.update(self.class_activation(preds), targets)
# the accuracies are updated here, but only computed and reported at the epoch, see "compute_on_step=False", "on_step=False", "on_epoch=True"
self.log('train_acc', self.train_acc, prog_bar=False, on_step=False, on_epoch=True)
return loss
def validation_step(self, batch, batch_nb):
images, targets = batch
if self.hparams.activation == "sigmoid":
targets = torch.nn.functional.one_hot(targets, num_classes=self.hparams.num_classes)
preds = self._forward(images)
loss = self.class_loss(preds, targets)
self.log("val_loss", loss)
self.val_acc.update(self.class_activation(preds), targets)
self.log('val_acc', self.val_acc, prog_bar=False, on_step=False, on_epoch=True)
return loss
def test_step(self, batch, batch_nb):
images, targets = batch
if self.hparams.activation == "sigmoid":
targets = torch.nn.functional.one_hot(targets, num_classes=self.hparams.num_classes)
preds = self._forward(images)
loss = self.class_loss(preds, targets)
self.log("test_loss", loss)
self.test_acc.update(self.class_activation(preds), targets)
self.log('test_acc', self.test_acc, prog_bar=False, on_step=False, on_epoch=True)
self.test_confmatrix.update(self.class_activation(preds), targets)
return loss
# this moves data to gpu when forward is called
@auto_move_data
def forward(self, x):
# in pl forward() is intended for inference, hence the activation is done here
logits = self._forward(x)
return self.class_activation(logits)
num_classes = 10
learning_rate = 1e-2
momentum = 0.9
weight_decay = 5e-4
pretrained = False
activation = "softmax"
# activation = "sigmoid"
# soft label: this is a multiplier to reduce the one hot 1s in case of sigmoid to a smaller value
# TODO: handle with care, this is very likely not correctly implemented
sigmoid_soft_factor = 1.0
scheduler = "one_cycle" # "cosine_annealing" or None
# one can also pass a partial scheduler class like the following to use other schedulers
# scheduler = partial(torch.optim.lr_scheduler.MultiStepLR, milestones=[3, 6], gamma=0.1)
rn_model = ResNetModel(
data.size(),
num_classes,
learning_rate,
momentum=momentum,
weight_decay=weight_decay,
pretrained=pretrained,
activation=activation,
sigmoid_soft_factor=sigmoid_soft_factor,
scheduler=scheduler,
)
rn_model.hparams
# -
# ## train model
# +
# run this to show very detailed model information and if model has example_input_array also detailed feature sizes
# pl.core.memory.ModelSummary(model, mode="full")
# + tags=[]
tb_logger = pl.loggers.TensorBoardLogger("/home/mtadmin/projects/tensorboard_logs")
plotter = ProgressPlotter()
callbacks = [
LightProgressBar(),
ProgressPrinter(),
plotter,
LearningRateMonitor(log_momentum=True),
]
# early_stop_callback = EarlyStopping(
# monitor="val_loss",
# mode="min",
# patience=5,
# verbose=True,
# )
# callbacks = [early_stop_callback] + callbacks
# model_checker = ModelCheckpoint(
# monitor="val_loss",
# mode="min",
# save_last=True,
# save_top_k=1,
# filename="best_{epoch}",
# )
# callbacks += [model_checker]
model = rn_model
max_epochs = 10
limit_train_batches = 1.0
limit_train_batches = 0.1 # 1.0 by default/full set, use this to check training on a smaller train set, also exists for val and test set
trainer = pl.Trainer(
max_epochs=max_epochs,
gpus=1,
callbacks=callbacks,
logger=[tb_logger],
precision=16,
# fast_dev_run=True,
limit_train_batches=limit_train_batches,
)
start_time = time.time()
trainer.fit(model, data)
print(f"runtime: {time.time() - start_time:.0f} s")
# -
# # we aim for ~99 %
# +
# trainer.logger.log_dir
# trainer.checkpoint_callback.best_model_path
# -
# static plot after training, useful when coming back after training finished
plotter.static_plot()
trainer.test(verbose=False)
plt.imshow(model.test_confmatrix.compute().cpu())
# ## compute accuracy the hard way
# +
model.eval()
all_targets = []
all_scores = []
for images, targets in tqdm(data.test_dataloader()):
with torch.no_grad():
scores = model(images).cpu().float()
all_scores.append(scores)
all_targets.append(targets)
scores = torch.cat(all_scores)
targets = torch.cat(all_targets)
# -
accuracy = (scores.max(1)[1] == targets).sum() / float(len(targets))
accuracy
| tutorial-vision_transformer-baseline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Morphology of the (s)RGB owing to Boundary Conditions
#
# A quick exploration of how different surface boundary conditions affect the morphology of the sub-giant-branch and red-giant-branch.
#
# ## GS98 solar abundance distribution
#
# All the models adopt the Grevesse & Sauval (1998) solar abundance distribution. Boundary conditions considered include:
# 1. Eddington grey $T(\tau)$ approximation (Eddington 1926).
# 2. Krishna-Swamy (1966) solar-calibrated grey $T(\tau)$ approximation.
# 3. Phoenix NextGen non-grey prescribed where $T(\tau) = T_{\rm eff}$ (Hauschildt 1999a,b; Dotter et al. 2007, 2008).
# 4. Phoenix NextGen non-grey prescribed where $\tau_{\rm ross} = 1$ (Feiden, _this note_)
# 5. Phoenix NextGen non-grey prescribed where $\tau_{\rm ross} = 10$ (Feiden, _this note_)
# 6. Phoenix NextGen non-grey prescribed where $\tau_{\rm ross} = 100$ (Feiden, _this note_)
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
def loadTrack(filename):
trk = np.genfromtxt(filename, usecols=(0,1,2,3,4))
bools = [(point[0] > 3.0e7) for point in trk]
return np.compress(bools, trk, axis=0)
#m1500_edd = loadTrack('files/trk/edd/m1550_GAS07_p000_p0_y26_mlt2.202.trk')
#m1500_ks66 = loadTrack('files/trk/ks/m1550_GAS07_p000_p0_y26_mlt2.202.trk')
m1500_dsep = loadTrack('files/trk/m150fehp00afep0.jc2mass')
m1500_teff = loadTrack('files/trk/m1500_GS98_p0_p0_T60.iso')
m1500_t010 = loadTrack('files/trk/m1500_GS98_p000_p0_y28_mlt1.884.trk')
# +
fig, ax = plt.subplots(1, 1, figsize=(8., 8.))
ax.set_xlabel('${\\rm Effective\\ Temperature\\ (K)}$', fontsize=22.)
ax.set_ylabel('$\\log(L / L_{\\odot})$', fontsize=22.)
ax.tick_params(which='major', axis='both', length=10., labelsize=16.)
ax.grid(True)
ax.set_xlim(8000., 2000.)
ax.plot(10**m1500_dsep[:,1], m1500_dsep[:,3], '-', dashes=(5.0, 5.0), lw=3, c='#069F74')
ax.plot(10**m1500_teff[:,1], m1500_teff[:,3], '-', dashes=(15.0, 10.0), lw=3, c='#800000')
ax.plot(10**m1500_t010[:,1], m1500_t010[:,3], '-', lw=3, c='#0473B3')
# -
# ## GAS07 solar abundance distribution
#
# All models below adopt the Grevesse et al. (2007) solar abundance distribution.
#
# We first begin with models that are identical with the exception of the surface boundary conditions. This means that tracks computed with the grey approximation _do not_ adopt the solar-calibrated mixing length parameter.
m1550_edd = loadTrack('files/trk/edd/m1550_GAS07_p000_p0_y26_mlt2.202.trk')
m1550_ks66 = loadTrack('files/trk/ks/m1550_GAS07_p000_p0_y26_mlt2.202.trk')
m1550_t050 = loadTrack('files/trk/m1550_GAS07_p000_p0_y26_mlt2.202.trk')
# +
fig, ax = plt.subplots(1, 1, figsize=(8., 8.))
ax.set_xlabel('${\\rm Effective\\ Temperature\\ (K)}$', fontsize=22.)
ax.set_ylabel('$\\log(L / L_{\\odot})$', fontsize=22.)
ax.tick_params(which='major', axis='both', length=10., labelsize=16.)
ax.grid(True)
ax.set_xlim(8000., 2000.)
ax.plot(10**m1550_edd[:,1], m1550_edd[:,3], '-', dashes=(5.0, 5.0), lw=3, c='#069F74')
ax.plot(10**m1550_ks66[:,1], m1550_ks66[:,3], '-', dashes=(15.0, 10.0), lw=3, c='#800000')
ax.plot(10**m1550_t050[:,1], m1550_t050[:,3], '-', lw=3, c='#0473B3')
# -
| Daily/20150812_rgb_morphology_bcs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="qmY71e44TPCs" colab={"base_uri": "https://localhost:8080/"} outputId="f43418aa-5611-4354-c803-aef198e935a9"
# !pip install git+https://github.com/AI4Finance-LLC/FinRL-Library.git
# + [markdown] id="gXaoZs2lh1hi"
# # Deep Reinforcement Learning for Stock Trading from Scratch: Multiple Stock Trading
#
# Tutorials to use OpenAI DRL to trade multiple stocks in one Jupyter Notebook | Presented at NeurIPS 2020: Deep RL Workshop
#
# * This blog is based on our paper: FinRL: A Deep Reinforcement Learning Library for Automated Stock Trading in Quantitative Finance, presented at NeurIPS 2020: Deep RL Workshop.
# * Check out medium blog for detailed explanations: https://towardsdatascience.com/finrl-for-quantitative-finance-tutorial-for-multiple-stock-trading-7b00763b7530
# * Please report any issues to our Github: https://github.com/AI4Finance-LLC/FinRL-Library/issues
# * **Pytorch Version**
#
#
# + [markdown] id="lGunVt8oLCVS"
# # Content
# + [markdown] id="HOzAKQ-SLGX6"
# * [1. Problem Definition](#0)
# * [2. Getting Started - Load Python packages](#1)
# * [2.1. Install Packages](#1.1)
# * [2.2. Check Additional Packages](#1.2)
# * [2.3. Import Packages](#1.3)
# * [2.4. Create Folders](#1.4)
# * [3. Download Data](#2)
# * [4. Preprocess Data](#3)
# * [4.1. Technical Indicators](#3.1)
# * [4.2. Perform Feature Engineering](#3.2)
# * [5.Build Environment](#4)
# * [5.1. Training & Trade Data Split](#4.1)
# * [5.2. User-defined Environment](#4.2)
# * [5.3. Initialize Environment](#4.3)
# * [6.Implement DRL Algorithms](#5)
# * [7.Backtesting Performance](#6)
# * [7.1. BackTestStats](#6.1)
# * [7.2. BackTestPlot](#6.2)
# * [7.3. Baseline Stats](#6.3)
# * [7.3. Compare to Stock Market Index](#6.4)
# + [markdown] id="sApkDlD9LIZv"
# <a id='0'></a>
# # Part 1. Problem Definition
# + [markdown] id="HjLD2TZSLKZ-"
# This problem is to design an automated trading solution for single stock trading. We model the stock trading process as a Markov Decision Process (MDP). We then formulate our trading goal as a maximization problem.
#
# The algorithm is trained using Deep Reinforcement Learning (DRL) algorithms and the components of the reinforcement learning environment are:
#
#
# * Action: The action space describes the allowed actions that the agent interacts with the
# environment. Normally, a ∈ A includes three actions: a ∈ {−1, 0, 1}, where −1, 0, 1 represent
# selling, holding, and buying one stock. Also, an action can be carried upon multiple shares. We use
# an action space {−k, ..., −1, 0, 1, ..., k}, where k denotes the number of shares. For example, "Buy
# 10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or −10, respectively
#
# * Reward function: r(s, a, s′) is the incentive mechanism for an agent to learn a better action. The change of the portfolio value when action a is taken at state s and arriving at new state s', i.e., r(s, a, s′) = v′ − v, where v′ and v represent the portfolio
# values at state s′ and s, respectively
#
# * State: The state space describes the observations that the agent receives from the environment. Just as a human trader needs to analyze various information before executing a trade, so
# our trading agent observes many different features to better learn in an interactive environment.
#
# * Environment: Dow 30 consituents
#
#
# The data of the single stock that we will be using for this case study is obtained from Yahoo Finance API. The data contains Open-High-Low-Close price and volume.
#
# + [markdown] id="Ffsre789LY08"
# <a id='1'></a>
# # Part 2. Getting Started- ASSUMES USING DOCKER, see readme for instructions
# + [markdown] id="Uy5_PTmOh1hj"
# <a id='1.1'></a>
# ## 2.1. Add FinRL to your path. You can of course install it as a pipy package, but this is for development purposes.
#
# + id="mPT0ipYE28wL"
import sys
sys.path.append("..")
# + id="0J6czZ7R3fEV" colab={"base_uri": "https://localhost:8080/"} outputId="54188ee8-d587-4f40-ae3a-4f33d8d90c7a"
import pandas as pd
print(pd.__version__)
# + [markdown] id="osBHhVysOEzi"
#
# <a id='1.2'></a>
# ## 2.2. Check if the additional packages needed are present, if not install them.
# * Yahoo Finance API
# * pandas
# * numpy
# * matplotlib
# * stockstats
# * OpenAI gym
# * stable-baselines
# * tensorflow
# * pyfolio
# + [markdown] id="nGv01K8Sh1hn"
# <a id='1.3'></a>
# ## 2.3. Import Packages
# + id="lPqeTTwoh1hn" colab={"base_uri": "https://localhost:8080/"} outputId="91d812fd-b82e-468d-b357-9e41964347b1"
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
matplotlib.use('Agg')
# %matplotlib inline
import datetime
from finrl.apps import config
from finrl.neo_finrl.preprocessor.yahoodownloader import YahooDownloader
from finrl.neo_finrl.preprocessor.preprocessors import FeatureEngineer, data_split
from finrl.neo_finrl.env_stock_trading.env_stocktrading_cashpenalty import StockTradingEnvCashpenalty
from finrl.drl_agents.stablebaselines3.models import DRLAgent
from finrl.plot import backtest_stats, backtest_plot, get_daily_return, get_baseline,convert_daily_return_to_pyfolio_ts
import sys
sys.path.append("../FinRL-Library")
# + [markdown] id="T2owTj985RW4"
# <a id='1.4'></a>
# ## 2.4. Create Folders
# + id="w9A8CN5R5PuZ"
import os
if not os.path.exists("./" + config.DATA_SAVE_DIR):
os.makedirs("./" + config.DATA_SAVE_DIR)
if not os.path.exists("./" + config.TRAINED_MODEL_DIR):
os.makedirs("./" + config.TRAINED_MODEL_DIR)
if not os.path.exists("./" + config.TENSORBOARD_LOG_DIR):
os.makedirs("./" + config.TENSORBOARD_LOG_DIR)
if not os.path.exists("./" + config.RESULTS_DIR):
os.makedirs("./" + config.RESULTS_DIR)
# + [markdown] id="A289rQWMh1hq"
# <a id='2'></a>
# # Part 3. Download Data
# Yahoo Finance is a website that provides stock data, financial news, financial reports, etc. All the data provided by Yahoo Finance is free.
# * FinRL uses a class **YahooDownloader** to fetch data from Yahoo Finance API
# * Call Limit: Using the Public API (without authentication), you are limited to 2,000 requests per hour per IP (or up to a total of 48,000 requests a day).
#
# + [markdown] id="NPeQ7iS-LoMm"
#
#
# -----
# class YahooDownloader:
# Provides methods for retrieving daily stock data from
# Yahoo Finance API
#
# Attributes
# ----------
# start_date : str
# start date of the data (modified from config.py)
# end_date : str
# end date of the data (modified from config.py)
# ticker_list : list
# a list of stock tickers (modified from config.py)
#
# Methods
# -------
# fetch_data()
# Fetches data from yahoo API
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="h3XJnvrbLp-C" outputId="4468b4e2-c14e-4e89-981d-38c6f8bd0ee8"
# from config.py start_date is a string
config.START_DATE
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="FUnY8WEfLq3C" outputId="2950fd55-3dfb-4f69-d86e-2c75a501736e"
# from config.py end_date is a string
config.END_DATE
# + colab={"base_uri": "https://localhost:8080/"} id="JzqRRTOX6aFu" outputId="f286c49a-8667-425f-93eb-f461e5e1a8b2"
print(config.DOW_30_TICKER)
# + colab={"base_uri": "https://localhost:8080/"} id="yCKm4om-s9kE" outputId="1250f26a-197d-4727-825e-0cc5fcc10071"
tickers = ['IBM', 'JNJ', 'CSCO', 'NKE', 'MCD']
df = YahooDownloader(start_date = '2009-01-01',
end_date = '2021-01-01',
ticker_list = tickers).fetch_data()
# + [markdown] id="uqC6c40Zh1iH"
# # Part 4: Preprocess Data
# Data preprocessing is a crucial step for training a high quality machine learning model. We need to check for missing data and do feature engineering in order to convert the data into a model-ready state.
# * Add technical indicators. In practical trading, various information needs to be taken into account, for example the historical stock prices, current holding shares, technical indicators, etc. In this article, we demonstrate two trend-following technical indicators: MACD and RSI.
# * Add turbulence index. Risk-aversion reflects whether an investor will choose to preserve the capital. It also influences one's trading strategy when facing different market volatility level. To control the risk in a worst-case scenario, such as financial crisis of 2007–2008, FinRL employs the financial turbulence index that measures extreme asset price fluctuation.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} id="xzijKkDF3fEY" colab={"base_uri": "https://localhost:8080/"} outputId="543bdd99-edd8-4d8d-ede2-7af3aa51fcf9"
fe = FeatureEngineer(
use_technical_indicator=True,
tech_indicator_list = config.TECHNICAL_INDICATORS_LIST,
use_turbulence=False,
user_defined_feature = False)
processed = fe.preprocess_data(df)
# + id="BbYbbrZA3fEY" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="cac54cdb-72ff-43d9-8fe5-8e339a0c6e7c"
processed['log_volume'] = np.log(processed.volume*processed.close)
processed['change'] = (processed.close-processed.open)/processed.close
processed['daily_variance'] = (processed.high-processed.low)/processed.close
processed = processed.sort_values(['date','tic'],ignore_index=True)
processed.head()
# + id="6Sgw_v3SUqCw" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="d2129a21-24b3-4764-fa88-46aa44f0879e"
processed.tail()
# + [markdown] id="-QsYaY0Dh1iw"
# <a id='4'></a>
# # Part 5. Design Environment
# Considering the stochastic and interactive nature of the automated stock trading tasks, a financial task is modeled as a **Markov Decision Process (MDP)** problem. The training process involves observing stock price change, taking an action and reward's calculation to have the agent adjusting its strategy accordingly. By interacting with the environment, the trading agent will derive a trading strategy with the maximized rewards as time proceeds.
#
# Our trading environments, based on OpenAI Gym framework, simulate live stock markets with real market data according to the principle of time-driven simulation.
#
# The action space describes the allowed actions that the agent interacts with the environment. Normally, action a includes three actions: {-1, 0, 1}, where -1, 0, 1 represent selling, holding, and buying one share. Also, an action can be carried upon multiple shares. We use an action space {-k,…,-1, 0, 1, …, k}, where k denotes the number of shares to buy and -k denotes the number of shares to sell. For example, "Buy 10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or -10, respectively. The continuous action space needs to be normalized to [-1, 1], since the policy is defined on a Gaussian distribution, which needs to be normalized and symmetric.
# + [markdown] id="5TOhcryx44bb"
# ## Training data split: 2009-01-01 to 2016-01-01
# ## Trade data split: 2016-01-01 to 2021-01-01
#
# DRL model needs to update periodically in order to take full advantage of the data, ideally we need to retrain our model yearly, quarterly, or monthly. We also need to tune the parameters along the way, in this notebook I only use the in-sample data from 2009-01 to 2016-01 to tune the parameters once, so there is some alpha decay here as the length of trade date extends.
#
# Numerous hyperparameters – e.g. the learning rate, the total number of samples to train on – influence the learning process and are usually determined by testing some variations.
# + colab={"base_uri": "https://localhost:8080/"} id="W0qaVGjLtgbI" outputId="dba01e86-62a0-494f-b132-3bee3aa047a9"
train = data_split(processed, '2009-01-01','2019-01-01')
trade = data_split(processed, '2019-01-01','2021-01-01')
print(len(train))
print(len(trade))
# + [markdown] id="8ckaw35dW5eF"
# ## Create the stock trading environment
# + id="k2XdqNUdW909"
import numpy as np
import pandas as pd
from gym.utils import seeding
import gym
from gym import spaces
import matplotlib
from copy import deepcopy
matplotlib.use("Agg")
import matplotlib.pyplot as plt
import pickle
from stable_baselines3.common.vec_env import DummyVecEnv
import numpy as np
import pandas as pd
from gym.utils import seeding
import gym
from gym import spaces
import matplotlib
import random
matplotlib.use("Agg")
import matplotlib.pyplot as plt
import pickle
from stable_baselines3.common.vec_env import DummyVecEnv, SubprocVecEnv
#from stable_baselines3.common import logger
import time
class MyStockTradingEnvCashpenalty(gym.Env):
"""
A stock trading environment for OpenAI gym
This environment penalizes the model for not maintaining a reserve of cash.
This enables the model to manage cash reserves in addition to performing trading procedures.
Reward at any step is given as follows
r_i = (sum(cash, asset_value) - initial_cash - max(0, sum(cash, asset_value)*cash_penalty_proportion-cash))/(days_elapsed)
This reward function takes into account a liquidity requirement, as well as long-term accrued rewards.
Parameters:
df (pandas.DataFrame): Dataframe containing data
buy_cost_pct (float): cost for buying shares
sell_cost_pct (float): cost for selling shares
hmax (int, array): maximum cash to be traded in each trade per asset. If an array is provided, then each index correspond to each asset
discrete_actions (bool): option to choose whether perform dicretization on actions space or not
shares_increment (int): multiples number of shares can be bought in each trade. Only applicable if discrete_actions=True
turbulence_threshold (float): Maximum turbulence allowed in market for purchases to occur. If exceeded, positions are liquidated
print_verbosity(int): When iterating (step), how often to print stats about state of env
initial_amount: (int, float): Amount of cash initially available
daily_information_columns (list(str)): Columns to use when building state space from the dataframe. It could be OHLC columns or any other variables such as technical indicators and turbulence index
cash_penalty_proportion (int, float): Penalty to apply if the algorithm runs out of cash
patient (bool): option to choose whether end the cycle when we're running out of cash or just don't buy anything until we got additional cash
RL Inputs and Outputs
action space: [<n_assets>,] in range {-1, 1}
state space: {start_cash, [shares_i for in in assets], [[indicator_j for j in indicators] for i in assets]]}
TODO:
Organize functions
Write README
Document tests
"""
metadata = {"render.modes": ["human"]}
def __init__(
self,
df,
buy_cost_pct=3e-3,
sell_cost_pct=3e-3,
date_col_name="date",
hmax=10,
discrete_actions=False,
shares_increment=1,
turbulence_threshold=None,
print_verbosity=10,
initial_amount=1e6,
daily_information_cols=['open', 'high', 'low', 'close', 'volume'],
cache_indicator_data=True,
cash_penalty_proportion=0.1,
random_start=True,
patient=False,
currency="$",
):
self.df = df
self.stock_col = "tic"
self.assets = df[self.stock_col].unique()
self.dates = df[date_col_name].sort_values().unique()
self.random_start = random_start
self.discrete_actions = discrete_actions
self.patient = patient
self.currency = currency
self.df = self.df.set_index(date_col_name)
self.shares_increment = shares_increment
self.hmax = hmax
self.initial_amount = initial_amount
self.print_verbosity = print_verbosity
self.buy_cost_pct = buy_cost_pct
self.sell_cost_pct = sell_cost_pct
self.turbulence_threshold = turbulence_threshold
self.daily_information_cols = daily_information_cols
self.state_space = (
1 + len(self.assets) + len(self.assets) * len(self.daily_information_cols)
)
self.action_space = spaces.Box(low=-1, high=1, shape=(len(self.assets),))
self.observation_space = spaces.Box(
low=-np.inf, high=np.inf, shape=(self.state_space,)
)
self.turbulence = 0
self.episode = -1 # initialize so we can call reset
self.episode_history = []
self.printed_header = False
self.cache_indicator_data = cache_indicator_data
self.cached_data = None
self.cash_penalty_proportion = cash_penalty_proportion
if self.cache_indicator_data:
print("caching data")
self.cached_data = [
self.get_date_vector(i) for i, _ in enumerate(self.dates)
]
print("data cached!")
def seed(self, seed=None):
if seed is None:
seed = int(round(time.time() * 1000))
random.seed(seed)
@property
def current_step(self):
return self.date_index - self.starting_point
@property
def cash_on_hand(self):
# amount of cash held at current timestep
return self.state_memory[-1][0]
@property
def holdings(self):
# Quantity of shares held at current timestep
return self.state_memory[-1][1 : len(self.assets) + 1]
@property
def closings(self):
return np.array(self.get_date_vector(self.date_index, cols=["close"]))
def reset(self):
self.seed()
self.sum_trades = 0
if self.random_start:
starting_point = random.choice(range(int(len(self.dates) * 0.5)))
self.starting_point = starting_point
else:
self.starting_point = 0
self.date_index = self.starting_point
self.turbulence = 0
self.episode += 1
self.actions_memory = []
self.transaction_memory = []
self.state_memory = []
self.account_information = {
"cash": [],
"asset_value": [],
"total_assets": [],
"reward": [],
}
init_state = np.array(
[self.initial_amount]
+ [0] * len(self.assets)
+ self.get_date_vector(self.date_index)
)
self.state_memory.append(init_state)
return init_state
def get_date_vector(self, date, cols=None):
if (cols is None) and (self.cached_data is not None):
return self.cached_data[date]
else:
date = self.dates[date]
if cols is None:
cols = self.daily_information_cols
trunc_df = self.df.loc[[date]]
v = []
for a in self.assets:
subset = trunc_df[trunc_df[self.stock_col] == a]
v += subset.loc[date, cols].tolist()
assert len(v) == len(self.assets) * len(cols)
return v
def return_terminal(self, reason="Last Date", reward=0):
state = self.state_memory[-1]
self.log_step(reason=reason, terminal_reward=reward)
# Add outputs to logger interface
gl_pct = self.account_information["total_assets"][-1] / self.initial_amount
# logger.record("environment/GainLoss_pct", (gl_pct - 1) * 100)
# logger.record(
# "environment/total_assets",
# int(self.account_information["total_assets"][-1]),
# )
reward_pct = self.account_information["total_assets"][-1] / self.initial_amount
# logger.record("environment/total_reward_pct", (reward_pct - 1) * 100)
# logger.record("environment/total_trades", self.sum_trades)
# logger.record(
# "environment/avg_daily_trades",
# self.sum_trades / (self.current_step),
# )
# logger.record(
# "environment/avg_daily_trades_per_asset",
# self.sum_trades / (self.current_step) / len(self.assets),
# )
# logger.record("environment/completed_steps", self.current_step)
# logger.record(
# "environment/sum_rewards", np.sum(self.account_information["reward"])
# )
# logger.record(
# "environment/cash_proportion",
# self.account_information["cash"][-1]
# / self.account_information["total_assets"][-1],
# )
return state, reward, True, {}
def log_step(self, reason, terminal_reward=None):
if terminal_reward is None:
terminal_reward = self.account_information["reward"][-1]
cash_pct = (
self.account_information["cash"][-1]
/ self.account_information["total_assets"][-1]
)
gl_pct = self.account_information["total_assets"][-1] / self.initial_amount
rec = [
self.episode,
self.date_index - self.starting_point,
reason,
f"{self.currency}{'{:0,.0f}'.format(float(self.account_information['cash'][-1]))}",
f"{self.currency}{'{:0,.0f}'.format(float(self.account_information['total_assets'][-1]))}",
f"{terminal_reward*100:0.5f}%",
f"{(gl_pct - 1)*100:0.5f}%",
f"{cash_pct*100:0.2f}%",
]
self.episode_history.append(rec)
print(self.template.format(*rec))
def log_header(self):
if self.printed_header is False:
self.template = "{0:4}|{1:4}|{2:15}|{3:15}|{4:15}|{5:10}|{6:10}|{7:10}" # column widths: 8, 10, 15, 7, 10
print(
self.template.format(
"EPISODE",
"STEPS",
"TERMINAL_REASON",
"CASH",
"TOT_ASSETS",
"TERMINAL_REWARD_unsc",
"GAINLOSS_PCT",
"CASH_PROPORTION",
)
)
self.printed_header = True
def get_reward(self):
if self.current_step == 0:
return 0
else:
assets = self.account_information["total_assets"][-1]
cash = self.account_information["cash"][-1]
cash_penalty = max(0, (assets * self.cash_penalty_proportion - cash))
assets -= cash_penalty
reward = (assets / self.initial_amount) - 1
reward /= self.current_step
return reward
def get_transactions(self, actions):
"""
This function takes in a raw 'action' from the model and makes it into realistic transactions
This function includes logic for discretizing
It also includes turbulence logic.
"""
# record actions of the model
self.actions_memory.append(actions)
# multiply actions by the hmax value
actions = actions * self.hmax
# Do nothing for shares with zero value
actions = np.where(self.closings > 0, actions, 0)
# discretize optionally
if self.discrete_actions:
# convert into integer because we can't buy fraction of shares
actions = actions // self.closings
actions = actions.astype(int)
# round down actions to the nearest multiplies of shares_increment
actions = np.where(
actions >= 0,
(actions // self.shares_increment) * self.shares_increment,
((actions + self.shares_increment) // self.shares_increment)
* self.shares_increment,
)
else:
actions = actions / self.closings
# can't sell more than we have
actions = np.maximum(actions, -np.array(self.holdings))
# deal with turbulence
if self.turbulence_threshold is not None:
# if turbulence goes over threshold, just clear out all positions
if self.turbulence >= self.turbulence_threshold:
actions = -(np.array(self.holdings))
self.log_step(reason="TURBULENCE")
return actions
def step(self, actions):
# let's just log what we're doing in terms of max actions at each step.
self.sum_trades += np.sum(np.abs(actions))
self.log_header()
# print if it's time.
if (self.current_step + 1) % self.print_verbosity == 0:
self.log_step(reason="update")
# if we're at the end
if self.date_index == len(self.dates) - 1:
# if we hit the end, set reward to total gains (or losses)
return self.return_terminal(reward=self.get_reward())
else:
"""
First, we need to compute values of holdings, save these, and log everything.
Then we can reward our model for its earnings.
"""
# compute value of cash + assets
begin_cash = self.cash_on_hand
assert min(self.holdings) >= 0
asset_value = np.dot(self.holdings, self.closings)
# log the values of cash, assets, and total assets
self.account_information["cash"].append(begin_cash)
self.account_information["asset_value"].append(asset_value)
self.account_information["total_assets"].append(begin_cash + asset_value)
# compute reward once we've computed the value of things!
reward = self.get_reward()
self.account_information["reward"].append(reward)
"""
Now, let's get down to business at hand.
"""
transactions = self.get_transactions(actions)
# compute our proceeds from sells, and add to cash
sells = -np.clip(transactions, -np.inf, 0)
proceeds = np.dot(sells, self.closings)
costs = proceeds * self.sell_cost_pct
coh = begin_cash + proceeds
# compute the cost of our buys
buys = np.clip(transactions, 0, np.inf)
spend = np.dot(buys, self.closings)
costs += spend * self.buy_cost_pct
# if we run out of cash...
if (spend + costs) > coh:
if self.patient:
# ... just don't buy anything until we got additional cash
self.log_step(reason="CASH SHORTAGE")
transactions = np.where(transactions > 0, 0, transactions)
spend = 0
costs = 0
else:
# ... end the cycle and penalize
return self.return_terminal(
reason="CASH SHORTAGE", reward=self.get_reward()
)
self.transaction_memory.append(
transactions
) # capture what the model's could do
# verify we didn't do anything impossible here
assert (spend + costs) <= coh
# update our holdings
coh = coh - spend - costs
holdings_updated = self.holdings + transactions
self.date_index += 1
if self.turbulence_threshold is not None:
self.turbulence = self.get_date_vector(
self.date_index, cols=["turbulence"]
)[0]
# Update State
state = (
[coh] + list(holdings_updated) + self.get_date_vector(self.date_index)
)
self.state_memory.append(state)
return state, reward, False, {}
def get_sb_env(self):
def get_self():
return deepcopy(self)
e = DummyVecEnv([get_self])
obs = e.reset()
return e, obs
def get_multiproc_env(self, n=10):
def get_self():
return deepcopy(self)
e = SubprocVecEnv([get_self for _ in range(n)], start_method="fork")
obs = e.reset()
return e, obs
def save_asset_memory(self):
if self.current_step == 0:
return None
else:
self.account_information["date"] = self.dates[
-len(self.account_information["cash"]) :
]
return pd.DataFrame(self.account_information)
def save_action_memory(self):
if self.current_step == 0:
return None
else:
return pd.DataFrame(
{
"date": self.dates[-len(self.account_information["cash"]) :],
"actions": self.actions_memory,
"transactions": self.transaction_memory,
}
)
# + [markdown] id="iyuPzwP7XAdS"
#
# + id="FCqTEpfh3fEZ" colab={"base_uri": "https://localhost:8080/"} outputId="05616dc9-0758-4e16-ed4c-abc69e817211"
print(MyStockTradingEnvCashpenalty.__doc__)
# + [markdown] id="Q2zqII8rMIqn" outputId="8a2c943b-1be4-4b8d-b64f-666e0852b7e6"
# #### state space
# The state space of the observation is as follows
#
# `start_cash, <owned_shares_of_n_assets>, <<indicator_i_for_asset_j> for j in assets>`
#
# indicators are any daily measurement you can achieve. Common ones are 'volume', 'open' 'close' 'high', 'low'.
# However, you can add these as needed,
# The feature engineer adds indicators, and you can add your own as well.
#
# + id="Ac5vErdj3fEa" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="a6c83b68-36ef-4ded-c98a-74eb937577b9"
processed.head()
# + id="AWyp84Ltto19" colab={"base_uri": "https://localhost:8080/"} outputId="e020821d-25ca-46d7-8603-9f5a41d8f51c"
information_cols = ['daily_variance', 'change', 'log_volume', 'close','day',
'macd', 'rsi_30', 'cci_30', 'dx_30']
e_train_gym = MyStockTradingEnvCashpenalty(df = train,initial_amount = 1e6,hmax = 5000,
turbulence_threshold = None,
currency='$',
buy_cost_pct=3e-3,
sell_cost_pct=3e-3,
cash_penalty_proportion=0.1,
cache_indicator_data=True,
daily_information_cols = information_cols,
print_verbosity = 100,
patient = True,
random_start = True)
e_trade_gym = MyStockTradingEnvCashpenalty(df = trade,initial_amount = 1e6,hmax = 5000,
turbulence_threshold = None,
currency='$',
buy_cost_pct=3e-3,
sell_cost_pct=3e-3,
cash_penalty_proportion=0.1,
cache_indicator_data=True,
daily_information_cols = information_cols,
print_verbosity = 100,
patient = True,
random_start = False)
# + [markdown] id="64EoqOrQjiVf"
# ## Environment for Training
# There are two available environments. The multiprocessing and the single processing env.
# Some models won't work with multiprocessing.
#
# ```python
# # single processing
# env_train, _ = e_train_gym.get_sb_env()
#
#
# #multiprocessing
# env_train, _ = e_train_gym.get_multiproc_env(n = <n_cores>)
# ```
#
# + colab={"base_uri": "https://localhost:8080/"} id="xwSvvPjutpqS" outputId="7777af40-370f-4c93-91a8-589dbaf176f4"
# for this example, let's do multiprocessing with n_cores-2
import multiprocessing
n_cores = multiprocessing.cpu_count() - 2
n_cores = 24
print(f"using {n_cores} cores")
#this is our training env. It allows multiprocessing
env_train, _ = e_train_gym.get_multiproc_env(n = n_cores)
# env_train, _ = e_train_gym.get_sb_env()
#this is our observation environment. It allows full diagnostics
env_trade, _ = e_trade_gym.get_sb_env()
# + [markdown] id="HMNR5nHjh1iz"
# <a id='5'></a>
# # Part 6: Implement DRL Algorithms
# * The implementation of the DRL algorithms are based on **OpenAI Baselines** and **Stable Baselines**. Stable Baselines is a fork of OpenAI Baselines, with a major structural refactoring, and code cleanups.
# * FinRL library includes fine-tuned standard DRL algorithms, such as DQN, DDPG,
# Multi-Agent DDPG, PPO, SAC, A2C and TD3. We also allow users to
# design their own DRL algorithms by adapting these DRL algorithms.
# + id="364PsqckttcQ"
agent = DRLAgent(env = env_train)
# + [markdown] id="uijiWgkuh1jB"
# ### Model PPO
#
# + id="Lc7V2tWp3fEb" colab={"base_uri": "https://localhost:8080/"} outputId="daa89b13-9512-49af-e6f1-53a7b7e7cf2a"
# from torch.nn import Softsign, ReLU
ppo_params ={'n_steps': 256,
'ent_coef': 0.0,
'learning_rate': 0.000005,
'batch_size': 1024,
'gamma': 0.99}
policy_kwargs = {
# "activation_fn": ReLU,
"net_arch": [1024 for _ in range(10)],
# "squash_output": True
}
model = agent.get_model("ppo",
model_kwargs = ppo_params,
policy_kwargs = policy_kwargs, verbose = 0)
# model = model.load("scaling_reward.model", env = env_train)
# + id="zTwet1oZ3fEb" colab={"base_uri": "https://localhost:8080/"} outputId="df0774da-4a23-4fca-b229-fb01d347e700"
model.learn(total_timesteps = 20000,
eval_env = env_trade,
eval_freq = 500,
log_interval = 1,
tb_log_name = 'env_cashpenalty_highlr',
n_eval_episodes = 1)
# + id="9Jbf8TQq3fEc"
model.save("different1_24.model")
# + [markdown] id="U5mmgQF_h1jQ"
# ### Trade
#
# DRL model needs to update periodically in order to take full advantage of the data, ideally we need to retrain our model yearly, quarterly, or monthly. We also need to tune the parameters along the way, in this notebook I only use the in-sample data from 2009-01 to 2018-12 to tune the parameters once, so there is some alpha decay here as the length of trade date extends.
#
# Numerous hyperparameters – e.g. the learning rate, the total number of samples to train on – influence the learning process and are usually determined by testing some variations.
# + id="gABFy6u03fEc" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="cae04044-d3df-4411-d849-92cdcf26a996"
trade.head()
# + id="eLOnL5eYh1jR"
e_trade_gym.hmax = 5000
# + id="Bq9BaTCQ3fEc" colab={"base_uri": "https://localhost:8080/"} outputId="0d9ed052-99ff-4319-fa2e-8ed0a6b13518"
print(len(e_trade_gym.dates))
# + id="WlDYBI9e3fEd" colab={"base_uri": "https://localhost:8080/"} outputId="4e44f5d7-730e-4b01-ddfc-70be33da65f0"
df_account_value, df_actions = DRLAgent.DRL_prediction(model=model,environment = e_trade_gym)
# + id="PzEigeLB3fEd" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="64aaacf3-acd5-4b69-ae48-30df8911b2d0"
df_actions.head()
# + colab={"base_uri": "https://localhost:8080/"} id="ERxw3KqLkcP4" outputId="ff819e29-6cf3-4732-b2a7-118b8af69ce1"
df_account_value.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="2yRkNguY5yvp" outputId="da2d4b18-2553-4424-ee2c-e1207843ef38"
df_account_value.head(50)
# + [markdown] id="W6vvNSC6h1jZ"
# <a id='6'></a>
# # Part 7: Backtest Our Strategy
# Backtesting plays a key role in evaluating the performance of a trading strategy. Automated backtesting tool is preferred because it reduces the human error. We usually use the Quantopian pyfolio package to backtest our trading strategies. It is easy to use and consists of various individual plots that provide a comprehensive image of the performance of a trading strategy.
# + [markdown] id="Lr2zX7ZxNyFQ"
# <a id='6.1'></a>
# ## 7.1 BackTestStats
# pass in df_account_value, this information is stored in env class
#
# + colab={"base_uri": "https://localhost:8080/"} id="Nzkr9yv-AdV_" outputId="704e0979-4129-46d4-82ba-4799c9cc1e58"
print("==============Get Backtest Results===========")
perf_stats_all = backtest_stats(account_value=df_account_value, value_col_name = 'total_assets')
# + [markdown] id="9U6Suru3h1jc"
# <a id='6.2'></a>
# ## 7.2 BackTestPlot
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="lKRGftSS7pNM" outputId="72d5d595-5235-4f4b-d9c6-e04d1a5a30ef"
print("==============Compare to DJIA===========")
# %matplotlib inline
# S&P 500: ^GSPC
# Dow Jones Index: ^DJI
# NASDAQ 100: ^NDX
backtest_plot(df_account_value,
baseline_ticker = '^DJI',
baseline_start = '2019-01-01',
baseline_end = '2021-01-01', value_col_name = 'total_assets')
# + id="-AXVQiMZ3fEe"
| tutorials/tutorial_env_multistock_cashpenaltyMYVERSION222.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={}
import json
import time
import os
import requests
import fml_manager
from fml_manager import *
manager = fml_manager.FMLManager()
# -
# ## For more details about the FMLManager, please refer to this [document](https://kubefate.readthedocs.io/README.html)
# +
response = manager.load_data(url='./data/breast_b.csv', namespace='fate_flow_test_breast', table_name='breast_b', work_mode=1, head=1, partition=10)
output = json.loads(response.content)
print(output)
guest_job_id = output['jobId']
guest_query_condition = QueryCondition(job_id=guest_job_id)
manager.query_job_status(guest_query_condition, max_tries=20)
# + pycharm={}
# Pipeline
data_io = ComponentBuilder(name='dataio_0',
module='DataIO')\
.add_input_data('args.train_data')\
.add_output_data('train')\
.add_output_model('dataio').build()
hetero_feature_binning = ComponentBuilder(name='hetero_feature_binning_0',
module='HeteroFeatureBinning')\
.add_input_data('dataio_0.train')\
.add_output_data('train')\
.add_output_model('hetero_feature_binning').build()
hetero_feature_selection = ComponentBuilder(name='hetero_feature_selection_0',
module='HeteroFeatureSelection')\
.add_input_data('hetero_feature_binning_0.train')\
.add_input_isometric_model('hetero_feature_binning_0.hetero_feature_binning')\
.add_output_data('train')\
.add_output_model('selected').build()
hetero_lr = ComponentBuilder(name='hetero_lr_0',
module='HeteroLR')\
.add_input_train_data('hetero_feature_selection_0.train')\
.add_output_data('train')\
.add_output_model('hetero_lr').build()
evaluation = ComponentBuilder(name='evaluation_0',
module='Evaluation',
need_deploy=False)\
.add_input_data('hetero_lr_0.train')\
.add_output_data('evaluate').build()
pipeline = Pipeline(
data_io,
hetero_feature_selection,
hetero_feature_binning,
hetero_lr,
evaluation)
# Configuration
initiator = Initiator(role='guest', party_id=10000)
job_parameters = JobParameters(work_mode=1)
role = RoleBuilder()\
.add_guest(party_id=10000)\
.add_host(party_id=9999)\
.add_arbiter(party_id=9999).build()
guest_data_io_config = {
'with_label': [True],
'label_name': ['y'],
'label_type': ['int'],
'output_format': ['dense']
}
host_data_io_config = {
'with_label': [False],
'output_format': ['dense']
}
role_parameters = RoleParametersBuilder()\
.add_guest_train_data(namespace='fate_flow_test_breast', name='breast_b')\
.add_guest_module_config(module='dataio_0', config=guest_data_io_config)\
.add_host_train_data(namespace='fate_flow_test_breast', name='breast_a')\
.add_host_module_config(module='dataio_0', config=host_data_io_config).build()
hetero_lr_params = {
'penalty': 'L2',
'optimizer': 'rmsprop',
'eps': 1e-5,
'alpha': 0.01,
'max_iter': 3,
'converge_func': 'diff',
'batch_size': 320,
'learning_rate': 0.15,
'init_param': {
'init_method': 'random_uniform'
}
}
algorithm_parameters = AlgorithmParametersBuilder()\
.add_module_config(module='hetero_lr_0', config=hetero_lr_params).build()
config = Config(initiator, job_parameters, role, role_parameters, algorithm_parameters)
# +
response = manager.submit_job(pipeline.to_dict(),config.to_dict())
manager.prettify(response, verbose=True)
stdout = json.loads(response.content)
job_id = stdout['jobId']
query_condition = QueryCondition(job_id)
model_id, model_version = '', ''
manager.query_job_status(query_condition, max_tries=20)
manager.prettify(response, verbose=True)
output = json.loads(response.content)
model_id, model_version = output['data']['model_info']['model_id'], output['data']['model_info']['model_version']
# -
# ## Click [here](/fateboard-10000/) to view jobs in FATE Board
# + pycharm={}
# Pipline
data_io = ComponentBuilder(name='dataio_0',
module='DataIO')\
.add_input_data('args.train_data')\
.add_output_data('train')\
.add_output_model('dataio').build()
hetero_feature_binning = ComponentBuilder(name='hetero_feature_binning_0',
module='HeteroFeatureBinning')\
.add_input_data('dataio_0.train')\
.add_output_data('train')\
.add_output_model('hetero_feature_binning').build()
hetero_feature_selection = ComponentBuilder(name='hetero_feature_selection_0',
module='HeteroFeatureSelection')\
.add_input_data('hetero_feature_binning_0.train')\
.add_input_isometric_model('hetero_feature_binning_0.hetero_feature_binning')\
.add_output_data('eval')\
.add_output_model('selected').build()
evaluation = ComponentBuilder(name='evaluation_0',
module='Evaluation',
need_deploy=False)\
.add_input_data('hetero_feature_selection_0.eval')\
.add_output_data('evaluate').build()
pipeline = Pipeline(
data_io,
hetero_feature_selection,
hetero_feature_binning,
evaluation)
# Configuration
initiator = Initiator(role='guest', party_id=10000)
job_parameters = JobParameters(work_mode=1,
job_type='predict',
model_id='arbiter-9999#guest-10000#host-9999#model',
model_version='2020060802475836992436')
role = RoleBuilder()\
.add_guest(party_id=10000)\
.add_host(party_id=9999)\
.add_arbiter(party_id=9999).build()
role_parameters = RoleParametersBuilder()\
.add_guest_eval_data(namespace='fate_flow_test_breast', name='breast_b')\
.add_host_eval_data(namespace='fate_flow_test_breast', name='breast_a').build()
config = Config(initiator, job_parameters, role, role_parameters)
# -
response = manager.submit_job(pipeline.to_dict(), config.to_dict())
manager.prettify(response, verbose=True)
stdout = json.loads(response.content)
job_id = stdout['jobId']
query_condition = QueryCondition(job_id)
print(query_condition)
# +
model_id, model_version = '', ''
manager.query_job_status(query_condition, max_tries=20)
manager.prettify(response, verbose=True)
output = json.loads(response.content)
model_id, model_version = output['data']['model_info']['model_id'], output['data']['model_info']['model_version']
# + pycharm={}
# data is a pandas.DataFrame
data = manager.track_component_output_data(job_id='2020060806580950942753', role='guest', party_id='10000', component_name='hetero_lr_0')
print(data.head(4))
| fml_manager/Examples/Hetero_LR/hetero_lr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/larsaars/pgki_uebungen/blob/main/klausur/klausur_gedankenprotokoll.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="cwKaejnSHg2P"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="4xRsp-ZiH4hf" outputId="89b87da5-9e10-4777-ed70-269b5cd5df4e"
# 5
index = ['Abdul-Jabbar', 'Malone', 'James', 'Bryant', 'Jordan']
data = {
'points': [38.4, 37., 35.4, 33.6, 32.2],
'games': [1560, 1476, 1310, 1346, 1072],
'team': ['Lakers', 'Jazz', 'Chavaliers', 'Lakers', 'Bulls'],
}
pd.DataFrame(data, index=index)
# + colab={"base_uri": "https://localhost:8080/", "height": 267} id="CsZ0tYJmJbIP" outputId="75a626ee-0ac9-4234-8212-138fdb08bbd3"
# 4
X = np.random.uniform(-1, 1, (30, 3))
fig, axes = plt.subplots(3, 3)
for i, row in enumerate(axes):
for j, ax in enumerate(row):
x = X[:, i]
y = X[:, j]
ax.scatter(x, y, c='blue')
plt.show()
# + id="xYiq9iCZRnDO"
| klausur/klausur_gedankenprotokoll.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Nanostring Hack 2021
# language: python
# name: nanostring-hack-2021
# ---
# # Cell-cell communication analysis
#
# ## Performing differential expression (DE) analysis of the NanoString ROIs with `limma`
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import scanpy as sc
import anndata
import matplotlib as mpl
import scipy
import matplotlib.pyplot as plt
sc.settings.verbosity = 3 # verbosity: errors (0), warnings (1), info (2), hints (3)
#sc.logging.print_versions()
sc.settings.set_figure_params(dpi=80) # low dpi (dots per inch) yields small inline figures
# -
nanostring_data_path = '../KidneyDataset/'
# ## Read in and prepare NanoString data
# target QC’d, filtered, and quantile normalized (Q3) target level count data
# going to feed this into limma
Kidney_Q3Norm_TargetCountMatrix = pd.read_csv(nanostring_data_path + 'Kidney_Q3Norm_TargetCountMatrix.txt',
sep='\t', index_col=0)
# ROI metadata
Kidney_Sample_Annotations = pd.read_csv(nanostring_data_path + 'Kidney_Sample_Annotations.txt',
sep='\t')
# +
# here in Kidney_Sample_Annotations column 'region' there is no indication of whether tubule is proximal or distal
# need to add this depending on the SegmentLabel - neg means proximal tubules, PanCK - distal
def add_exact_region(idx, table):
curr_SegmentLabel = table.loc[idx,'SegmentLabel']
if curr_SegmentLabel == 'Geometric Segment':
return('glomerulus')
elif curr_SegmentLabel == 'neg':
return('proximal tubule')
elif curr_SegmentLabel == 'PanCK':
return('distal tubule')
else:
return('error')
Kidney_Sample_Annotations['idx'] = Kidney_Sample_Annotations.index
Kidney_Sample_Annotations['region_exact'] = Kidney_Sample_Annotations['idx'].apply(lambda x: add_exact_region(x,Kidney_Sample_Annotations))
Kidney_Sample_Annotations.set_index('SegmentDisplayName', inplace=True)
# -
# getting rid of NaNs
Kidney_Sample_Annotations['pathology'] = [elem if str(elem) != 'nan' else 'NA' for elem in Kidney_Sample_Annotations['pathology']]
# We will be comparing all abnormal ROIs to all healthy ROIs (acc to pathology)
# and then this 1 list of DE I will artificially copy for each of the single cell clusters
comparison_cluster = 'pathology'
Kidney_Q3Norm_TargetCountMatrix = Kidney_Q3Norm_TargetCountMatrix.T
# make sure indices match
Kidney_Sample_Annotations = Kidney_Sample_Annotations.loc[Kidney_Q3Norm_TargetCountMatrix.index]
# object with all features that passed QC (.raw.X)
adata_full = anndata.AnnData(X = Kidney_Q3Norm_TargetCountMatrix,
obs = Kidney_Sample_Annotations,
var = pd.DataFrame(index = Kidney_Q3Norm_TargetCountMatrix.columns)
)
# _____________________________________________________________________________________________________________________________________________________________
# # DE with limma
#
# - all abnormal ROIs vs all healthy ROIs according to pathology annotation
# discard the ROIs that are not glomeruli (aka tubules)
adata_full = adata_full[adata_full.obs['pathology'].isin(['abnormal', 'healthy'])].copy()
# +
# marker calling
t = adata_full.X.T
df = pd.DataFrame(data=t, columns= adata_full.obs.index, index=adata_full.var_names)
meta_df = pd.DataFrame(data={'Cell':list(adata_full.obs.index),
'cell_type':[ str(i) for i in adata_full.obs['pathology']],
#'sample':[ str(i) for i in adata_full.obs['sample']]
})
meta_df.set_index('Cell', inplace=True)
meta_df.reset_index(inplace=True)
# -
# %load_ext rpy2.ipython
outpath = './'
# + language="R"
# library(limma)
# library(edgeR)
# +
# because R replaces things
meta_df['Cell'] = [elem.replace(' | ','...') for elem in meta_df['Cell']]
meta_df['Cell'] = [elem.replace(' ','.') for elem in meta_df['Cell']]
#meta_df.head()
df.columns = [elem.replace(' | ','...') for elem in df.columns]
# -
np.unique(meta_df['cell_type'], return_counts=True)
case = 'abnormal'
ctrl = 'healthy'
# + magic_args="-i df -i meta_df -i outpath -i ctrl -i case " language="R"
#
# library(limma)
# library(edgeR)
#
# # Format
# ex_mat=as.matrix(df)
# rownames(meta_df) = meta_df$Cell
#
# # subset meta
# meta_df = subset(meta_df, cell_type %in% unlist(c(ctrl, case)) )
#
# print(unique(meta_df$cell_type))
#
# # Shared cells
# shared_cells = intersect(rownames(meta_df), colnames(ex_mat))
# message(length(shared_cells), ' shared cells')
# ex_mat = ex_mat[, shared_cells]
# meta_df = meta_df[shared_cells,]
#
# # Filter lowly expressed genes
# keep = rowSums(ex_mat, na.rm=T) > 0.1
# ex_mat = ex_mat[ keep, ]
# keep = aveLogCPM(ex_mat) > 0.1
# ex_mat = ex_mat[ keep, ]
#
# # Extract celltypes
# cells = rownames(meta_df)
# celltypes = unique(meta_df$cell_type)
# covariates = meta_df$covariate
#
# # Extract cells in cluster and rest
# cells_case = rownames(subset(meta_df, cell_type == case))
# cells_ctrl = rownames(subset(meta_df, cell_type == ctrl)) # changed from control to ctrl
#
# # build cluster_type vector
# cluster_type = rep(0, length(cells))
# names(cluster_type) = cells
# cluster_type[ cells_case ] = 'case'
# cluster_type[ cells_ctrl ] = 'ctrl'
#
# print(unique(cluster_type))
#
# #design.matrix <- model.matrix(~ 0 + cluster_type + covariates)
# design.matrix <- model.matrix(~ 0 + cluster_type)
#
# # Now tell limma how do you want to compare (i.e. case vs control)
# contrast.matrix <- makeContrasts(caseVScontrol = cluster_typecase - cluster_typectrl, levels = design.matrix)
#
# # Make model and run contrasts
# fit <- lmFit(ex_mat, design.matrix)
# fit <- contrasts.fit(fit, contrast.matrix)
# fit <- eBayes(fit)
#
# # Make a dataframe containing the important data
# results = topTable(fit, adjust="fdr", number = nrow(ex_mat), coef = 'caseVScontrol')
#
# # Add and filter needed data
# results$Gene = rownames(results)
# results = results[ , c('Gene', 'logFC', 'P.Value', 'adj.P.Val')]
# results$AveExpr_cluster = apply(ex_mat[ results$Gene, cells_case], 1, mean)
# results$AveExpr_rest = apply(ex_mat[ results$Gene, cells_ctrl], 1, mean)
# results$percentExpr_cluster = apply(ex_mat[ results$Gene, cells_case], 1, function(x) sum(c(x > 0)+0) ) / length(cells_case)
# results$percentExpr_rest = apply(ex_mat[ results$Gene, cells_ctrl], 1, function(x) sum(c(x > 0)+0) ) / length(cells_ctrl)
#
# results$AveExpr_cluster = round(results$AveExpr_cluster, 6)
# results$AveExpr_rest = round(results$AveExpr_rest, 6)
# results$percentExpr_cluster = round(results$percentExpr_cluster, 6)
# results$percentExpr_rest = round(results$percentExpr_rest, 6)
# # and store it as csv file
# write.csv(results, file = paste0(outpath, case, '_vs_', ctrl, '_limma_DEGs.csv'), row.names = F, col.names = T, quote = F)
# -
# let's have a look at the DEGs
DE_table = pd.read_csv('./abnormal_vs_healthy_limma_DEGs.csv', index_col=0)
DE_table
# how many significant DEGs?
DE_table_significant = DE_table[DE_table['adj.P.Val'] < 0.05]
DE_table_significant
# # Creating a joint DE table out of limma outputs from NanoString data
# Because we don't have cell type resolution here, let's copy this DE table for every cell type artificially to then intersect with the expressed L/R from Young et al. data
# from Young et al.
meta = pd.read_csv('./cellphonedb_meta.tsv', sep='\t')
cell_types = np.unique(meta['cell_type'])
# get rid of 'celltype_'
cell_types = [elem[9:] for elem in cell_types]
cell_types
# let's have a look at the DEGs
DE_table = pd.read_csv('./abnormal_vs_healthy_limma_DEGs.csv')
DE_table
# +
# artificially copying it for all the cell types
DE_tables = {}
for ct in cell_types:
#print(ct)
DE_tables[ct] = DE_table
# -
DE_table
# +
cluster_list = []
for ct in cell_types:
#print(ct)
cluster_list.append([ct]*len(DE_table))
cluster_list = [item for sublist in cluster_list for item in sublist]
# -
# without any filtering
joint_DE_table = pd.concat(DE_tables.values())
joint_DE_table['cluster'] = cluster_list
joint_DE_table
joint_DE_table.to_csv('./joint_DEGs_list_all_cell_types_for_cellphone.csv')
| Analysis/3_Cell-cell_comm_analysis/S4_DE_analysis_limma_ROIs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Stimuli delivery
#
# This interface use [Brython](https://brython.info) and the [Radiant framework](https://radiant-framework.readthedocs.io/en/latest/) as backend for do the web development in a Python style, this interface inherits all features from [Radiant](https://radiant-framework.readthedocs.io/en/latest/) and extends the utilities with an specific ones.
# -
# ## Bare minimum
# +
from bci_framework.extensions.stimuli_delivery import StimuliAPI
# Brython modules
from browser import document, html
from browser.widgets.dialog import InfoDialog
# Main class must inherit `StimuliAPI`
class StimuliDelivery(StimuliAPI):
def __init__(self, *args, **kwargs):
# Initialize `StimuliAPI` class
super().__init__(*args, **kwargs) # very importante line ;)
document.clear()
# -------------------------------------------------------------
# main brython code
document.select_one('body') <= html.H3('Hello world')
button = html.BUTTON('click me')
button.bind('click', lambda evt: InfoDialog('Hello', 'world'))
document.select_one('body') <= button
# -------------------------------------------------------------
if __name__ == '__main__':
# Create and run the server
StimuliDelivery()
# -
# <img src='images/stimuli_bare_minimum.png'></img>
# ## Stimuli area and Dashboard
#
# One of the main features is the possibility to make configurable experiments, in favor of this philosophy, by default they are builded both areas `self.stimuli_area` and `self.dashboard`.
# +
# -------------------------------------------------------------
# main brython code
# Create a division for the stimuli_area and the dashboard
self.stimuli_area <= html.H3('Stimuli area')
self.dashboard <= html.H3('Dashboard')
# Insert a cross in the middle of the stimuli area
self.show_cross()
# This area is used for external event processord
self.show_synchronizer()
# -------------------------------------------------------------
# -
# The `self.stimuli_area` at left attemp to be used to display stimuli and the `self.dashboard` on the right is for widgets and configurations.
# <img src='images/areas.png'></img>
# ## Widgets
#
# All widgets and styles they are part of [material-components-web](https://github.com/material-components/material-components-web) with a custom framework implementation designed to display widgets and get values.
#
# All widgets are available trought the `Widgets` submodule located in the module `bci_framework.extensions.stimuli_delivery.utils`.
#
# ``` python
# from bci_framework.extensions.stimuli_delivery.utils import Widgets as w
# ```
# The following styles are used for all examples
flex = {'margin-bottom': '15px', 'display': 'flex', }
flex_title = {'margin-top': '50px', 'margin-bottom': '10px', 'display': 'flex', }
# ### Typography
# +
# -------------------------------------------------------------
# main brython code
self.dashboard <= w.label('headline1', typo='headline1', style=flex)
self.dashboard <= w.label('headline2', typo='headline2', style=flex)
self.dashboard <= w.label('headline3', typo='headline3', style=flex)
self.dashboard <= w.label('headline4', typo='headline4', style=flex)
self.dashboard <= w.label('headline5', typo='headline5', style=flex)
self.dashboard <= w.label('headline6', typo='headline6', style=flex)
self.dashboard <= w.label('body1', typo='body1', style=flex)
self.dashboard <= w.label('body2', typo='body2', style=flex)
self.dashboard <= w.label('subtitle1', typo='subtitle1', style=flex)
self.dashboard <= w.label('subtitle2', typo='subtitle2', style=flex)
self.dashboard <= w.label('caption', typo='caption', style=flex)
self.dashboard <= w.label('button', typo='button', style=flex)
self.dashboard <= w.label('overline', typo='overline', style=flex)
# -------------------------------------------------------------
# -
# <img src='images/stimuli_typography.png'></img>
# ### Buttons
# +
# -------------------------------------------------------------
# main brython code
self.dashboard <= w.label('Buttons', typo='headline4', style=flex_title)
self.dashboard <= w.button('Button 1', style=flex, on_click=lambda: setattr(document.select_one('#for_button'), 'html', 'Button 1 pressed!'))
self.dashboard <= w.button('Button 2', style=flex, on_click=self.on_button2)
self.dashboard <= w.label(f'', id='for_button', typo=f'body1', style=flex)
# -------------------------------------------------------------
def on_button2(self):
document.select_one('#for_button').html = 'Button 2 pressed!'
# -
# <img src='images/stimuli_buttons.png'></img>
# ### Switch
# +
# -------------------------------------------------------------
# main brython code
self.dashboard <= w.label('Switch', typo='headline4', style=flex_title)
self.dashboard <= w.switch('Switch 1', checked=True, on_change=self.on_switch, id='my_swicth')
self.dashboard <= w.label(f'', id='for_switch', typo=f'body1', style=flex)
# -------------------------------------------------------------
def on_switch(self, value):
# value = self.widgets.get_value('my_swicth')
document.select_one('#for_switch').html = f'Switch Changed: {value}'
# -
# <img src='images/stimuli_switch.png'></img>
# ### Checkbox
# +
# -------------------------------------------------------------
# main brython code
self.dashboard <= w.label('Checkbox', typo='headline4', style=flex_title)
self.dashboard <= w.checkbox('Checkbox', options=[[f'chb-{i}', False] for i in range(4)], on_change=self.on_checkbox, id='my_checkbox')
self.dashboard <= w.label(f'', id='for_checkbox', typo=f'body1', style=flex)
# -------------------------------------------------------------
def on_checkbox(self):
value = w.get_value('my_checkbox')
document.select_one('#for_checkbox').html = f'Checkbox Changed: {value}'
# -
# <img src='images/stimuli_checkbox.png'></img>
# ### Radios
# +
# -------------------------------------------------------------
# main brython code
self.dashboard <= w.label('Radios', typo='headline4', style=flex_title)
self.dashboard <= w.radios('Radios', options=[[f'chb-{i}', f'chb-{i}'] for i in range(4)], on_change=self.on_radios, id='my_radios')
self.dashboard <= w.label(f'', id='for_radios', typo=f'body1', style=flex)
# -------------------------------------------------------------
def on_radios(self):
value = w.get_value('my_radios')
document.select_one('#for_radios').html = f'Radios Changed: {value}'
# -
# <img src='images/stimuli_radios.png'></img>
# ### Select
# +
# -------------------------------------------------------------
# main brython code
self.dashboard <= w.label('Select', typo='headline4', style=flex)
self.dashboard <= w.select('Select', [[f'sel-{i}', f'sel-{i}'] for i in range(4)], on_change=self.on_select, id='my_select')
self.dashboard <= w.label(f'', id='for_select', typo=f'body1', style=flex)
# -------------------------------------------------------------
def on_select(self, value):
# value = w.get_value('my_select')
document.select_one('#for_select').html = f'Select Changed: {value}'
# -
# <img src='images/stimuli_select.png'></img>
# ### Sliders
# +
# -------------------------------------------------------------
# main brython code
# Slider
self.dashboard <= w.label('Slider', typo='headline4', style=flex)
self.dashboard <= w.slider('Slider', min=1, max=10, step=0.1, value=5, on_change=self.on_slider, id='my_slider')
self.dashboard <= w.label(f'', id='for_slider', typo=f'body1', style=flex)
# Slider range
self.dashboard <= w.label('Slider range', typo='headline4', style=flex)
self.dashboard <= w.range_slider('Slider range', min=0, max=20, value_lower=5, value_upper=15, step=1, on_change=self.on_slider_range, id='my_range')
self.dashboard <= w.label(f'', id='for_range', typo=f'body1', style=flex)
# -------------------------------------------------------------
def on_slider(self, value):
# value = w.get_value('my_slider')
document.select_one('#for_slider').html = f'Slider Changed: {value}'
def on_slider_range(self, value):
# value = w.get_value('my_slider')
document.select_one('#for_range').html = f'Range Changed: {value}'
# -
# <img src='images/stimuli_sliders.png'></img>
# ## Sound
# ### Tones
#
# The `Tone` library allows playing single notes using the javascript `AudioContext` backend, the `duration` and the `gain` can also be configured.
from bci_framework.extensions.stimuli_delivery.utils import Tone as t
# -------------------------------------------------------------
# main brython code
duration = 100
gain = 0.5
self.dashboard <= w.button('f#4', on_click=lambda: t('f#4', duration, gain), style={'margin': '0 15px'})
self.dashboard <= w.button('D#0', on_click=lambda: t('D#0', duration, gain), style={'margin': '0 15px'})
self.dashboard <= w.button('B2', on_click=lambda: t('B2', duration, gain), style={'margin': '0 15px'})
# -------------------------------------------------------------
# <img src='images/stimuli_tones1.png'></img>
# ### Audio files
# Not implemented yet
# ## Pipelines
#
# Pipelines consist of the controlled execution of methods with asynchronous timeouts.
#
# Let's assume that we have 4 view methods, each method could be a step needed to build a trial.
# +
def view1(self, s1, r1):
print(f'On view1: {s1=}, {r1=}')
def view2(self, s1, r1):
print(f'On view2: {s1=}, {r1=}')
def view3(self, s1, r1):
print(f'On view3: {s1=}, {r1=}')
def view4(self, s1, r1):
print(f'On view4: {s1=}, {r1=}\n')
# -
# The first rule is that all methods will have the same arguments, these arguments consist of the needed information to build a single trial.
# Now, we need the `trials`, for example, here we define 3 trials (notice the arguments):
trials = [
{'s1': 'Hola', # Trial 1
'r1': 91,
},
{'s1': 'Mundo', # Trial 2
'r1': 85,
},
{'s1': 'Python', # Trial 3
'r1': 30,
},
]
# And the `pipeline` consists of a list of sequential methods with a respective timeout `(method, timeout)`, if the `timeout` is a number then this will indicate the milliseconds until the next method call. If the `timeout` is a list, then a random (with uniform distribution) number between that range will be generated on each trial.
pipeline_trial = [
(self.view1, 500),
(self.view2, [500, 1500]),
(self.view3, w.get_value('slider')),
(self.view4, w.get_value('range')),
]
# Finally, our pipeline can be executed with the method `self.run_pipeline`:
self.run_pipeline(pipeline_trial, trials)
# +
On view1: s1=Hola, r1=91
On view2: s1=Hola, r1=91
On view3: s1=Hola, r1=91
On view4: s1=Hola, r1=91
On view1: s1=Mundo, r1=85
On view2: s1=Mundo, r1=85
On view3: s1=Mundo, r1=85
On view4: s1=Mundo, r1=85
On view1: s1=Python, r1=30
On view2: s1=Python, r1=30
On view3: s1=Python, r1=30
On view4: s1=Python, r1=30
# -
# ## Recording EEG automatically
#
# If there is a current EEG streaming, the stimuli delivery can be configured to automatically start and stop the EEG recording with the methods `self.start_record()` and `self.stop_record()` respectively.
# ## Send Markers
#
# The markers are used to synchronize events. The `self.send_marker` method is available all the time to stream markers through the streaming platform.
self.send_marker("MARKER")
# <div class="alert alert-warning">
#
# The method _self.send_marker_ **works on the delivery views**, so, if you have not an active remote presentation the markers will never send.
#
# </div>
# ## Hardware-based event synchonization
#
# In case that there is needed maximum precision about markers synchronization is possible to [attach an external input](https://doi.org/10.3389/fninf.2020.00002) directly to the analog inputs of OpenBCI.
# The method `self.show_synchronizer()` do the trick, and is possible to configure the duration of the event with the `blink` argument in `self.send_marker`:
self.send_marker('RIGHT', blink=100)
self.send_marker('LEFT', blink=200)
# ## Fixation cross
#
# The fixation cross merely serves to center the subject's gaze at the center of the screen. We can control the presence of this mark with the methods:
self.show_cross()
self.hide_cross()
# ## Send annotations
#
# In the same way that `markers`, the anotations (as defined in the EDF file format) can be streamed with the `self.send_annotation` method.
self.send_annotationn('Data record start')
self.send_annotationn('The subject yawn', duration=5)
# ## Feedbacks
#
# The `feedbacks`are used to comunicate the [Data analysis](03-data_analysis.ipynb) and [Data visualizations](04-data_visualizations.ipynb) with the Stimuli Delivery platform. For this purpose, there is a predefined stream channel called `feedback`. This is useful to develop [Neurofeedback applications](07-neurofeedback.ipynb).
#
# The asynchronous handler can be configured with the `Feedback` class:
from bci_framework.extensions.stimuli_delivery import Feedback
# This class needs an `ID` and bind a method:
self.feedback = Feedback(self, 'my_feedback_id') # ID
self.feedback.on_feedback(self.on_input_feedback)
# So, this method will be executed asynchronously on each feedback call.
def on_input_feedback(self, **feedback):
...
# The feedbacks are bidirectional comunications, so, is possible to write messages back to the data rocessor with the method `Feedback.write`:
self.feedback.write(data)
| docs/source/notebooks/06-stimuli_delivery.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.7 64-bit
# metadata:
# interpreter:
# hash: 09d279961bf9625707cb254620cad507eb2d4a44de356bc3dcf8e99132d7a735
# name: python3
# ---
import psycopg2 as db
con=db.connect(host="PTPGS0016.ikeadt.com",
database="rdc",
user="rcmpc",
password="<PASSWORD>")
cur=con.cursor()
# + tags=["outputPrepend"]
cur.execute("select item_no ,iip_insert_date ,iip_update_date from rcmpc_cem.cem_item_bu_range_t cibrt")
rows=cur.fetchall()
for r in rows:
print(f"item_no{r[0]} iip_insert_date {r[1]} iip_update_date {r[2]}")
cur.close()
con.close()
# -
cur.execute("select 'Item Sales Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_range_t ccibrt union select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_range_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD');")
cur.fetchone()
cur.execute("select 'Item Sales Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_range_t ccibrt union select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_range_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item CCL Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_comclass_t ccibct union select item_no,iip_update_date from rcmpc_cem.cem_item_bu_comclass_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item ROC Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_rangeoffer_t ccirt union select item_no,iip_update_date from rcmpc_cem.cem_item_rangeoffer_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item ROCC Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_rangeo_cri_t ccirct union select item_no,iip_update_date from rcmpc_cem.cem_item_rangeoffer_cri_t circt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD');")
cur.fetchall()
# +
import matplotlib.pyplot as plt
import numpy as np
cur.execute("select 'Item Sales Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_range_t ccibrt union select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_range_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item CCL Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_comclass_t ccibct union select item_no,iip_update_date from rcmpc_cem.cem_item_bu_comclass_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item ROC Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_rangeoffer_t ccirt union select item_no,iip_update_date from rcmpc_cem.cem_item_rangeoffer_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item ROCC Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_rangeo_cri_t ccirct union select item_no,iip_update_date from rcmpc_cem.cem_item_rangeoffer_cri_t circt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD');")
attribute = []
cnt = []
for rec in cur:
attribute.append(rec[0])
cnt.append(rec[1])
np.asarray(attribute,dtype='S')
np.asarray(cnt,dtype='float')
plt.plot(attribute,cnt,color='blue', label='User Count')
# +
import matplotlib.pyplot as plt
import numpy as np
cur.execute("select 'Item Sales Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_range_t ccibrt union select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_range_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item CCL Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_bu_comclass_t ccibct union select item_no,iip_update_date from rcmpc_cem.cem_item_bu_comclass_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item ROC Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_rangeoffer_t ccirt union select item_no,iip_update_date from rcmpc_cem.cem_item_rangeoffer_t ccibrt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD') UNION select 'Item ROCC Delta' as attribute,count(1) as cnt from (select item_no,iip_update_date from rcmpc_cem.cem_cre_item_rangeo_cri_t ccirct union select item_no,iip_update_date from rcmpc_cem.cem_item_rangeoffer_cri_t circt2 ) a where iip_update_date::date between to_Date('2019-06-04','YYYY-MM-DD') and to_Date('2019-06-05','YYYY-MM-DD');")
item_no = []
iip_insert_date = []
iip_update_date=[]
for rec in cur:
item_no.append(rec[0])
iip_insert_date.append(rec[1])
iip_update_date.append(rec[2])
np.asarray(item_no,dtype='float')
np.asarray(iip_insert_date,dtype='float')
np.asarray(iip_update_date,dtype='float')
plt.plot(item_no, iip_insert_date,iip_update_date, color='blue', label='User Count')
# +
x = np.linspace(0, 10, 30)
y = np.sin(x)
plt.plot(x, y, 'o', color='black');
# +
cur = con.cursor()
query=("select item_no ,iip_insert_date ,iip_update_date from rcmpc_cem.cem_item_bu_range_t cibrt")
outputquery = "COPY ({0}) TO STDOUT WITH CSV HEADER".format(query)
with open('resultsfile', 'w') as f:
cur.copy_expert(outputquery, f)
con.close()
# -
# +
from tkinter import *
root=Tk()
w = Label(root, text="RDC JOB SCRIPT!")
w.pack()
root.mainloop()
# -
from PIL import ImageTk, Image
import matplotlib.pyplot as plt
try:
import tkinter as tk
from tkinter import ttk
except ImportError:
import Tkinter as tk
import ttk
# +
from tkcalendar import Calendar, DateEntry
def date_pic():
top = tk.Toplevel(root)
ttk.Label(top, text='Choose date').pack(padx=10, pady=10)
cal = DateEntry(top, width=12, background='darkblue',
foreground='white',font={"courier",50}, borderwidth=2)
cal.pack(padx=10, pady=10)
root = tk.Tk()
s = ttk.Style(root)
s.theme_use('clam')
ttk.Button(root, text='DateEntry', command=date_pic).pack(padx=10, pady=10)
root.mainloop()
# +
from tkinter import *
from tkcalendar import Calendar,DateEntry
root = Tk()
cal = DateEntry(root,width=30,bg="darkblue",fg="white",year=2010)
cal.grid()
root.mainloop()
# +
from tkinter import *
from tkinter import messagebox
from PIL import ImageTk, Image
root = Tk()
C = Canvas(root, bg="blue", height=250, width=300)
filename = ImageTk.PhotoImage(Image.open(r"C:\Users\dibha2\download.jpg"),master=root)
background_label = Label(root, image=filename)
background_label.image=filename
background_label.place(x=0, y=0, relwidth=1, relheight=1)
cal = DateEntry(root,width=30,bg="darkblue",fg="white",year=2010)
cal.grid()
C.pack()
root.mainloop()
# +
from tkinter import *
from PIL import ImageTk, Image
root = Tk()
canv =Canvas(root, bg="blue", height=750, width=600)
canv.grid(row=3, column=3)
img = ImageTk.PhotoImage(Image.open(r"C:\Users\dibha2\download.jpg"),master=root) # PIL solution
canv.create_image(50, 50, anchor=NW, image=img)
mainloop()
# +
from flask import Flask
app = Flask(__name__)
@app.route('/')
def index():
return 'RDC JOB AUTOMATION'
app.run(host='0.0.0.0', port=81)
# -
| python/Untitled-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''momo'': conda)'
# name: python3710jvsc74a57bd0f8af005c6536e801c34ed1a329b6361f501640f22aff193c57cfc4ad0e1dbb64
# ---
# # 6 - Transformers for Sentiment Analysis
#
# In this notebook we will be using the transformer model, first introduced in [this](https://arxiv.org/abs/1706.03762) paper. Specifically, we will be using the BERT (Bidirectional Encoder Representations from Transformers) model from [this](https://arxiv.org/abs/1810.04805) paper.
#
# Transformer models are considerably larger than anything else covered in these tutorials. As such we are going to use the [transformers library](https://github.com/huggingface/transformers) to get pre-trained transformers and use them as our embedding layers. We will freeze (not train) the transformer and only train the remainder of the model which learns from the representations produced by the transformer. In this case we will be using a multi-layer bi-directional GRU, however any model can learn from these representations.
# ## Preparing Data
#
# First, as always, let's set the random seeds for deterministic results.
# +
import torch
import random
import numpy as np
SEED = 1234
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
# -
# The transformer has already been trained with a specific vocabulary, which means we need to train with the exact same vocabulary and also tokenize our data in the same way that the transformer did when it was initially trained.
#
# Luckily, the transformers library has tokenizers for each of the transformer models provided. In this case we are using the BERT model which ignores casing (i.e. will lower case every word). We get this by loading the pre-trained `bert-base-uncased` tokenizer.
# +
from transformers import BertTokenizer
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
# -
# The `tokenizer` has a `vocab` attribute which contains the actual vocabulary we will be using. We can check how many tokens are in it by checking its length.
len(tokenizer.vocab)
# Using the tokenizer is as simple as calling `tokenizer.tokenize` on a string. This will tokenize and lower case the data in a way that is consistent with the pre-trained transformer model.
# +
tokens = tokenizer.tokenize('Hello WORLD how ARE yoU?')
print(tokens)
# -
# We can numericalize tokens using our vocabulary using `tokenizer.convert_tokens_to_ids`.
# +
indexes = tokenizer.convert_tokens_to_ids(tokens)
print(indexes)
# -
# The transformer was also trained with special tokens to mark the beginning and end of the sentence, detailed [here](https://huggingface.co/transformers/model_doc/bert.html#transformers.BertModel). As well as a standard padding and unknown token. We can also get these from the tokenizer.
#
# **Note**: the tokenizer does have a beginning of sequence and end of sequence attributes (`bos_token` and `eos_token`) but these are not set and should not be used for this transformer.
# +
init_token = tokenizer.cls_token
eos_token = tokenizer.sep_token
pad_token = tokenizer.pad_token
unk_token = tokenizer.unk_token
print(init_token, eos_token, pad_token, unk_token)
# -
# We can get the indexes of the special tokens by converting them using the vocabulary...
# +
init_token_idx = tokenizer.convert_tokens_to_ids(init_token)
eos_token_idx = tokenizer.convert_tokens_to_ids(eos_token)
pad_token_idx = tokenizer.convert_tokens_to_ids(pad_token)
unk_token_idx = tokenizer.convert_tokens_to_ids(unk_token)
print(init_token_idx, eos_token_idx, pad_token_idx, unk_token_idx)
# -
# ...or by explicitly getting them from the tokenizer.
# +
init_token_idx = tokenizer.cls_token_id
eos_token_idx = tokenizer.sep_token_id
pad_token_idx = tokenizer.pad_token_id
unk_token_idx = tokenizer.unk_token_id
print(init_token_idx, eos_token_idx, pad_token_idx, unk_token_idx)
# -
# Another thing we need to handle is that the model was trained on sequences with a defined maximum length - it does not know how to handle sequences longer than it has been trained on. We can get the maximum length of these input sizes by checking the `max_model_input_sizes` for the version of the transformer we want to use. In this case, it is 512 tokens.
# +
max_input_length = tokenizer.max_model_input_sizes['bert-base-uncased']
print(max_input_length)
# -
# Previously we have used the `spaCy` tokenizer to tokenize our examples. However we now need to define a function that we will pass to our `TEXT` field that will handle all the tokenization for us. It will also cut down the number of tokens to a maximum length. Note that our maximum length is 2 less than the actual maximum length. This is because we need to append two tokens to each sequence, one to the start and one to the end.
def tokenize_and_cut(sentence):
tokens = tokenizer.tokenize(sentence)
tokens = tokens[:max_input_length-2]
return tokens
# Now we define our fields. The transformer expects the batch dimension to be first, so we set `batch_first = True`. As we already have the vocabulary for our text, provided by the transformer we set `use_vocab = False` to tell torchtext that we'll be handling the vocabulary side of things. We pass our `tokenize_and_cut` function as the tokenizer. The `preprocessing` argument is a function that takes in the example after it has been tokenized, this is where we will convert the tokens to their indexes. Finally, we define the special tokens - making note that we are defining them to be their index value and not their string value, i.e. `100` instead of `[UNK]` This is because the sequences will already be converted into indexes.
#
# We define the label field as before.
# +
from torchtext import data
TEXT = data.Field(batch_first = True,
use_vocab = False,
tokenize = tokenize_and_cut,
preprocessing = tokenizer.convert_tokens_to_ids,
init_token = init_token_idx,
eos_token = eos_token_idx,
pad_token = pad_token_idx,
unk_token = unk_token_idx)
LABEL = data.LabelField(dtype = torch.float)
# -
# We load the data and create the validation splits as before.
# +
from torchtext import datasets
train_data, test_data = datasets.IMDB.splits(TEXT, LABEL)
train_data, valid_data = train_data.split(random_state = random.seed(SEED))
# -
print(f"Number of training examples: {len(train_data)}")
print(f"Number of validation examples: {len(valid_data)}")
print(f"Number of testing examples: {len(test_data)}")
# We can check an example and ensure that the text has already been numericalized.
print(vars(train_data.examples[6]))
# We can use the `convert_ids_to_tokens` to transform these indexes back into readable tokens.
# +
tokens = tokenizer.convert_ids_to_tokens(vars(train_data.examples[6])['text'])
print(tokens)
# -
# Although we've handled the vocabulary for the text, we still need to build the vocabulary for the labels.
LABEL.build_vocab(train_data)
print(LABEL.vocab.stoi)
# As before, we create the iterators. Ideally we want to use the largest batch size that we can as I've found this gives the best results for transformers.
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print ('Available devices ', torch.cuda.device_count())
print ('Current cuda device ', torch.cuda.current_device())
print(torch.cuda.get_device_name(device))
# GPU 할당 변경하기
GPU_NUM = 1 # 원하는 GPU 번호 입력
device = torch.device(f'cuda:{GPU_NUM}' if torch.cuda.is_available() else 'cpu')
torch.cuda.set_device(device) # change allocation of current GPU
# +
print ('Current cuda device ', torch.cuda.current_device()) # check
# Additional Infos
if device.type == 'cuda':
print(torch.cuda.get_device_name(GPU_NUM))
print('Memory Usage:')
print('Allocated:', round(torch.cuda.memory_allocated(GPU_NUM)/1024**3,1), 'GB')
print('Cached: ', round(torch.cuda.memory_cached(GPU_NUM)/1024**3,1), 'GB')
# +
BATCH_SIZE = 128
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE,
device = device)
# -
# ## Build the Model
#
# Next, we'll load the pre-trained model, making sure to load the same model as we did for the tokenizer.
# +
from transformers import BertTokenizer, BertModel
bert = BertModel.from_pretrained('bert-base-uncased')
# -
# Next, we'll define our actual model.
#
# Instead of using an embedding layer to get embeddings for our text, we'll be using the pre-trained transformer model. These embeddings will then be fed into a GRU to produce a prediction for the sentiment of the input sentence. We get the embedding dimension size (called the `hidden_size`) from the transformer via its config attribute. The rest of the initialization is standard.
#
# Within the forward pass, we wrap the transformer in a `no_grad` to ensure no gradients are calculated over this part of the model. The transformer actually returns the embeddings for the whole sequence as well as a *pooled* output. The [documentation](https://huggingface.co/transformers/model_doc/bert.html#transformers.BertModel) states that the pooled output is "usually not a good summary of the semantic content of the input, you’re often better with averaging or pooling the sequence of hidden-states for the whole input sequence", hence we will not be using it. The rest of the forward pass is the standard implementation of a recurrent model, where we take the hidden state over the final time-step, and pass it through a linear layer to get our predictions.
# +
import torch.nn as nn
class BERTGRUSentiment(nn.Module):
def __init__(self,
bert,
hidden_dim,
output_dim,
n_layers,
bidirectional,
dropout):
super().__init__()
self.bert = bert
embedding_dim = bert.config.to_dict()['hidden_size']
self.rnn = nn.GRU(embedding_dim,
hidden_dim,
num_layers = n_layers,
bidirectional = bidirectional,
batch_first = True,
dropout = 0 if n_layers < 2 else dropout)
self.out = nn.Linear(hidden_dim * 2 if bidirectional else hidden_dim, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, text):
#text = [batch size, sent len]
with torch.no_grad():
embedded = self.bert(text)[0]
#embedded = [batch size, sent len, emb dim]
_, hidden = self.rnn(embedded)
#hidden = [n layers * n directions, batch size, emb dim]
if self.rnn.bidirectional:
hidden = self.dropout(torch.cat((hidden[-2,:,:], hidden[-1,:,:]), dim = 1))
else:
hidden = self.dropout(hidden[-1,:,:])
#hidden = [batch size, hid dim]
output = self.out(hidden)
#output = [batch size, out dim]
return output
# -
# Next, we create an instance of our model using standard hyperparameters.
# +
HIDDEN_DIM = 256
OUTPUT_DIM = 1
N_LAYERS = 2
BIDIRECTIONAL = True
DROPOUT = 0.25
model = BERTGRUSentiment(bert,
HIDDEN_DIM,
OUTPUT_DIM,
N_LAYERS,
BIDIRECTIONAL,
DROPOUT)
# -
# We can check how many parameters the model has. Our standard models have under 5M, but this one has 112M! Luckily, 110M of these parameters are from the transformer and we will not be training those.
# +
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
# -
# In order to freeze paramers (not train them) we need to set their `requires_grad` attribute to `False`. To do this, we simply loop through all of the `named_parameters` in our model and if they're a part of the `bert` transformer model, we set `requires_grad = False`.
for name, param in model.named_parameters():
if name.startswith('bert'):
param.requires_grad = False
# We can now see that our model has under 3M trainable parameters, making it almost comparable to the `FastText` model. However, the text still has to propagate through the transformer which causes training to take considerably longer.
# +
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
# -
# We can double check the names of the trainable parameters, ensuring they make sense. As we can see, they are all the parameters of the GRU (`rnn`) and the linear layer (`out`).
for name, param in model.named_parameters():
if param.requires_grad:
print(name)
# ## Train the Model
#
# As is standard, we define our optimizer and criterion (loss function).
# +
import torch.optim as optim
optimizer = optim.Adam(model.parameters())
# -
criterion = nn.BCEWithLogitsLoss()
# Place the model and criterion onto the GPU (if available)
model = model.to(device)
criterion = criterion.to(device)
# Next, we'll define functions for: calculating accuracy, performing a training epoch, performing an evaluation epoch and calculating how long a training/evaluation epoch takes.
def binary_accuracy(preds, y):
"""
Returns accuracy per batch, i.e. if you get 8/10 right, this returns 0.8, NOT 8
"""
#round predictions to the closest integer
rounded_preds = torch.round(torch.sigmoid(preds))
correct = (rounded_preds == y).float() #convert into float for division
acc = correct.sum() / len(correct)
return acc
def train(model, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
optimizer.zero_grad()
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
def evaluate(model, iterator, criterion):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# +
import time
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# -
# Finally, we'll train our model. This takes considerably longer than any of the previous models due to the size of the transformer. Even though we are not training any of the transformer's parameters we still need to pass the data through the model which takes a considerable amount of time on a standard GPU.
# +
N_EPOCHS = 5
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut6-model.pt')
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
# -
# We'll load up the parameters that gave us the best validation loss and try these on the test set - which gives us our best results so far!
# +
model.load_state_dict(torch.load('tut6-model.pt'))
test_loss, test_acc = evaluate(model, test_iterator, criterion)
print(f'Test Loss: {test_loss:.3f} | Test Acc: {test_acc*100:.2f}%')
# -
# ## Inference
#
# We'll then use the model to test the sentiment of some sequences. We tokenize the input sequence, trim it down to the maximum length, add the special tokens to either side, convert it to a tensor, add a fake batch dimension and then pass it through our model.
def predict_sentiment(model, tokenizer, sentence):
model.eval()
tokens = tokenizer.tokenize(sentence)
tokens = tokens[:max_input_length-2]
indexed = [init_token_idx] + tokenizer.convert_tokens_to_ids(tokens) + [eos_token_idx]
tensor = torch.LongTensor(indexed).to(device)
tensor = tensor.unsqueeze(0)
prediction = torch.sigmoid(model(tensor))
return prediction.item()
predict_sentiment(model, tokenizer, "This film is terrible")
predict_sentiment(model, tokenizer, "This film is great")
| 6 - Transformers for Sentiment Analysis.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
using LinearAlgebra
using SymPy
# Creates arrays with orthogonal states via simply slicing up a diagonal matrix of size 2^numQubits ⊗ 2^numQubits
function ortho_states(numQubits::Int64)
States=2^numQubits
V = [zeros(Int64, 1, States ) for _ in 1:States]
count=1
for i in V
V[count][count]=1
count +=1
end
V
end
#Convert the indexing values to actual symbols for manipulation.
# Requires the number of qubits in the data set, the data set, and a prefix to track the name
function indexNameSymbols(Dat_qubits::Int64, Dat::Array{Array{Int64,2},1}, Prefix::String)
Sym=repeat([""],2^Dat_qubits)
for i in 1:2^Dat_qubits
for j in 1:2^Dat_qubits
Sym[i]*=string(Dat[j][i])
end
end
Sym
[symbols(Prefix*string(i)) for i in Sym]
end
Nouns = ["John","James","Bill","Bob"];
Verbs = ["met","saw","likes",""];
Ns_qubits = ceil(Int64,log2(length(Nouns)));
Ns = ortho_states(Ns_qubits);
No_qubits = ceil(Int64,log2(length(Nouns)));
No = ortho_states(No_qubits);
V_qubits = ceil(Int64,log2(length(Verbs)));
V = ortho_states(V_qubits);
#Create the arrays of unqique symbols, each corresponding to the coefficient of a specific state
# eg SymS = s_{10}*|0> + s_{01}*|1>, where |0> = (1 0)' and |1> = (0 1)'
SymNs = indexNameSymbols(Ns_qubits, Ns, "Ns");
SymNo = indexNameSymbols(Ns_qubits, No, "No");
SymV = indexNameSymbols(V_qubits, V, "V");
kron(Nouns,kron(Verbs,Nouns))
| modules/py/nb/TensorOps_jl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''base'': conda)'
# language: python
# name: python38364bitbaseconda97cd5720aa194725b86ca3d3cdd6281a
# ---
# # Debugging the baseline SAKT model
# - There is seemingly a bug in the `iter_env`.
#
# ### TO-DO:
# features encoding:
# - how to address the problem with previous answers correctly not uniformly predicted
# - question tags
# +
import os
import gc
import sys
import pickle
from time import time
import datatable as dt
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import torch
import torch.nn as nn
from torch import optim
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import train_test_split
from torch.autograd import Variable
import torch.optim as optim
from torch.optim import Optimizer
from torch.optim.lr_scheduler import (CosineAnnealingWarmRestarts, CyclicLR, OneCycleLR,
ReduceLROnPlateau)
from torch.utils.data import DataLoader, Dataset
from tqdm import tqdm
sns.set()
DEFAULT_FIG_WIDTH = 20
sns.set_context("paper", font_scale=1.2)
# WORKSPACE_FOLDER=/home/scao/Documents/kaggle-riiid-test
# PYTHONPATH=${WORKSPACE_FOLDER}:${WORKSPACE_FOLDER}/sakt:${WORKSPACE_FOLDER}/transformer
HOME = os.path.abspath(os.path.join('.', os.pardir))
print(HOME, '\n\n')
# HOME = "/home/scao/Documents/kaggle-riiid-test/"
MODEL_DIR = os.path.join(HOME, 'model')
DATA_DIR = os.path.join(HOME, 'data')
sys.path.append(HOME)
from utils import *
get_system()
from sakt import *
from iter_env import *
# +
# set-up
DEBUG = True
TRAIN = False
PREPROCESS = False
TEST_SIZE = 0.05
NUM_SKILLS = 13523 # number of problems
MAX_SEQ = 180
ACCEPTED_USER_CONTENT_SIZE = 4
EMBED_SIZE = 128
NUM_HEADS = 8
BATCH_SIZE = 64
VAL_BATCH_SIZE = 2048
DEBUG_TEST_SIZE = 2500
DROPOUT = 0.1
SEED = 1127
get_seed(SEED)
'''
Columns placeholder and preprocessing params
'''
CONTENT_TYPE_ID = "content_type_id"
CONTENT_ID = "content_id"
TARGET = "answered_correctly"
USER_ID = "user_id"
PRIOR_QUESTION_TIME = 'prior_question_elapsed_time'
PRIOR_QUESTION_EXPLAIN = 'prior_question_had_explanation'
TASK_CONTAINER_ID = "task_container_id"
TIMESTAMP = "timestamp"
ROW_ID = 'row_id'
FILLNA_VAL = 14_000 # for prior question elapsed time, rounded average in train
TIME_SCALING = 1000 # scaling down the prior question elapsed time
TRAIN_COLS = [TIMESTAMP, USER_ID, CONTENT_ID, CONTENT_TYPE_ID, TARGET]
TRAIN_DTYPES = {TIMESTAMP: 'int64',
USER_ID: 'int32',
CONTENT_ID: 'int16',
CONTENT_TYPE_ID: 'bool',
TARGET:'int8',
PRIOR_QUESTION_TIME: np.float32,
PRIOR_QUESTION_EXPLAIN: 'boolean'}
if DEBUG:
NROWS_TEST = 25_000
NROWS_TRAIN = 5_000_000
NROWS_VAL = 500_000
else:
NROWS_TEST = 250_000
NROWS_TRAIN = 50_000_000
NROWS_VAL = 2_000_000
# +
if PREPROCESS:
with timer("Loading train from parquet"):
train_df = pd.read_parquet(os.path.join(DATA_DIR, 'cv2_train.parquet'),
columns=list(TRAIN_DTYPES.keys())).astype(TRAIN_DTYPES)
valid_df = pd.read_parquet(os.path.join(DATA_DIR, 'cv2_valid.parquet'),
columns=list(TRAIN_DTYPES.keys())).astype(TRAIN_DTYPES)
if DEBUG:
train_df = train_df[:NROWS_TRAIN]
valid_df = valid_df[:NROWS_VAL]
with timer("Processing train"):
train_group = preprocess(train_df)
valid_group = preprocess(valid_df, train_flag=2)
else:
with open(os.path.join(DATA_DIR, 'sakt_group_cv2.pickle'), 'rb') as f:
group = pickle.load(f)
train_group, valid_group = train_test_split(group, test_size = TEST_SIZE, random_state=SEED)
print(f"valid users: {len(valid_group.keys())}")
print(f"train users: {len(train_group.keys())}")
# +
class SAKTDataset(Dataset):
def __init__(self, group, n_skill, max_seq=MAX_SEQ):
super(SAKTDataset, self).__init__()
self.samples, self.n_skill, self.max_seq = {}, n_skill, max_seq
self.user_ids = []
for i, user_id in enumerate(group.index):
content_id, answered_correctly = group[user_id]
if len(content_id) >= ACCEPTED_USER_CONTENT_SIZE:
if len(content_id) > self.max_seq:
total_questions = len(content_id)
last_pos = total_questions // self.max_seq
for seq in range(last_pos):
index = f"{user_id}_{seq}"
self.user_ids.append(index)
start = seq * self.max_seq
end = (seq + 1) * self.max_seq
self.samples[index] = (content_id[start:end],
answered_correctly[start:end])
if len(content_id[end:]) >= ACCEPTED_USER_CONTENT_SIZE:
index = f"{user_id}_{last_pos + 1}"
self.user_ids.append(index)
self.samples[index] = (content_id[end:],
answered_correctly[end:])
else:
index = f'{user_id}'
self.user_ids.append(index)
self.samples[index] = (content_id, answered_correctly)
def __len__(self):
return len(self.user_ids)
def __getitem__(self, index):
user_id = self.user_ids[index]
content_id, answered_correctly = self.samples[user_id]
seq_len = len(content_id)
content_id_seq = np.zeros(self.max_seq, dtype=int)
answered_correctly_seq = np.zeros(self.max_seq, dtype=int)
if seq_len >= self.max_seq:
content_id_seq[:] = content_id[-self.max_seq:]
answered_correctly_seq[:] = answered_correctly[-self.max_seq:]
else:
content_id_seq[-seq_len:] = content_id
answered_correctly_seq[-seq_len:] = answered_correctly
target_id = content_id_seq[1:] # question till the current one
label = answered_correctly_seq[1:]
x = content_id_seq[:-1].copy() # question till the previous one
# encoded answers till the previous one
x += (answered_correctly_seq[:-1] == 1) * self.n_skill
return x, target_id, label
train_dataset = SAKTDataset(train_group, n_skill=NUM_SKILLS, max_seq=MAX_SEQ)
train_dataloader = DataLoader(train_dataset,
batch_size=BATCH_SIZE,
shuffle=True,
drop_last=True)
val_dataset = SAKTDataset(valid_group, n_skill=NUM_SKILLS, max_seq=MAX_SEQ)
val_dataloader = DataLoader(val_dataset,
batch_size=VAL_BATCH_SIZE,
shuffle=False)
sample_batch = next(iter(train_dataloader))
sample_batch[0].shape, sample_batch[1].shape, sample_batch[2].shape
# +
class FFN(nn.Module):
def __init__(self, state_size = MAX_SEQ,
forward_expansion = 1,
bn_size=MAX_SEQ - 1,
dropout=0.2):
super(FFN, self).__init__()
self.state_size = state_size
self.lr1 = nn.Linear(state_size, forward_expansion * state_size)
self.relu = nn.ReLU()
self.bn = nn.BatchNorm1d(bn_size)
self.lr2 = nn.Linear(forward_expansion * state_size, state_size)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
x = self.relu(self.lr1(x))
x = self.bn(x)
x = self.lr2(x)
return self.dropout(x)
def future_mask(seq_length):
future_mask = (np.triu(np.ones([seq_length, seq_length]), k = 1)).astype('bool')
return torch.from_numpy(future_mask)
class TransformerBlock(nn.Module):
def __init__(self, embed_dim,
heads = 8,
dropout = DROPOUT,
forward_expansion = 1):
super(TransformerBlock, self).__init__()
self.multi_att = nn.MultiheadAttention(embed_dim=embed_dim,
num_heads=heads, dropout=dropout)
self.dropout = nn.Dropout(dropout)
self.layer_normal = nn.LayerNorm(embed_dim)
self.ffn = FFN(embed_dim,
forward_expansion = forward_expansion,
dropout=dropout)
self.layer_normal_2 = nn.LayerNorm(embed_dim)
def forward(self, value, key, query, att_mask):
att_output, att_weight = self.multi_att(value, key, query, attn_mask=att_mask)
att_output = self.dropout(self.layer_normal(att_output + value))
att_output = att_output.permute(1, 0, 2)
# att_output: [s_len, bs, embed] => [bs, s_len, embed]
x = self.ffn(att_output)
x = self.dropout(self.layer_normal_2(x + att_output))
return x.squeeze(-1), att_weight
class Encoder(nn.Module):
def __init__(self, n_skill, max_seq=100,
embed_dim=128,
dropout = DROPOUT,
forward_expansion = 1,
num_layers=1,
heads = 8):
super(Encoder, self).__init__()
self.n_skill, self.embed_dim = n_skill, embed_dim
self.embedding = nn.Embedding(2 * n_skill + 1, embed_dim)
self.pos_embedding = nn.Embedding(max_seq - 1, embed_dim)
self.e_embedding = nn.Embedding(n_skill+1, embed_dim)
self.layers = nn.ModuleList([TransformerBlock(embed_dim, heads=heads,
forward_expansion = forward_expansion) for _ in range(num_layers)])
self.dropout = nn.Dropout(dropout)
def forward(self, x, question_ids):
device = x.device
x = self.embedding(x)
pos_id = torch.arange(x.size(1)).unsqueeze(0).to(device)
pos_x = self.pos_embedding(pos_id)
x = self.dropout(x + pos_x)
x = x.permute(1, 0, 2) # x: [bs, s_len, embed] => [s_len, bs, embed]
e = self.e_embedding(question_ids)
e = e.permute(1, 0, 2)
for layer in self.layers:
att_mask = future_mask(e.size(0)).to(device)
x, att_weight = layer(e, x, x, att_mask=att_mask)
x = x.permute(1, 0, 2)
x = x.permute(1, 0, 2)
return x, att_weight
class SAKTModel(nn.Module):
def __init__(self,
n_skill,
max_seq=MAX_SEQ,
embed_dim=EMBED_SIZE,
dropout = DROPOUT,
forward_expansion = 1,
enc_layers=1,
heads = NUM_HEADS):
super(SAKTModel, self).__init__()
self.encoder = Encoder(n_skill,
max_seq,
embed_dim,
dropout,
forward_expansion,
num_layers=enc_layers,
heads=heads)
self.pred = nn.Linear(embed_dim, 1)
def forward(self, x, question_ids):
x, att_weight = self.encoder(x, question_ids)
x = self.pred(x)
return x.squeeze(-1), att_weight
class TestDataset(Dataset):
def __init__(self, samples, test_df, n_skill, max_seq=100):
super(TestDataset, self).__init__()
self.samples = samples
self.user_ids = [x for x in test_df["user_id"].unique()]
self.test_df = test_df
self.n_skill, self.max_seq = n_skill, max_seq
def __len__(self):
return self.test_df.shape[0]
def __getitem__(self, index):
test_info = self.test_df.iloc[index]
user_id = test_info['user_id']
target_id = test_info['content_id']
content_id_seq = np.zeros(self.max_seq, dtype=int)
answered_correctly_seq = np.zeros(self.max_seq, dtype=int)
if user_id in self.samples.index:
content_id, answered_correctly = self.samples[user_id]
seq_len = len(content_id)
if seq_len >= self.max_seq:
content_id_seq = content_id[-self.max_seq:]
answered_correctly_seq = answered_correctly[-self.max_seq:]
else:
content_id_seq[-seq_len:] = content_id
answered_correctly_seq[-seq_len:] = answered_correctly
x = content_id_seq[1:].copy()
x += (answered_correctly_seq[1:] == 1) * self.n_skill
questions = np.append(content_id_seq[2:], [target_id])
return x, questions
# +
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(f'\nUsing device: {device}')
model_file = MODEL_DIR+'sakt_seq_180_auc_0.7689.pth'
model = SAKTModel(n_skill=NUM_SKILLS,
max_seq=MAX_SEQ,
embed_dim=EMBED_SIZE,
forward_expansion=1,
enc_layers=1,
heads=NUM_HEADS,
dropout=DROPOUT)
n_params = get_num_params(model)
print(f"Current model has {n_params} parameters.")
model = model.to(device)
model.load_state_dict(torch.load(model_file, map_location=device))
# +
# mock test
with timer("Loading private simulated test set"):
all_test_df = pd.read_parquet(DATA_DIR+'cv2_valid.parquet')
all_test_df = all_test_df[:DEBUG_TEST_SIZE]
all_test_df['answer_correctly_true'] = all_test_df[TARGET]
predicted = []
def set_predict(df):
predicted.append(df)
# reload all user group for cv2
with timer('loading cv2'):
with open(os.path.join(DATA_DIR, 'sakt_group_cv2.pickle'), 'rb') as f:
group = pickle.load(f)
# -
def iter_env_run(all_test_df, n_iter=1):
'''
Running mock test for n_iter iterations using tito's iter_env simulator and cv2_train user group.
'''
iter_test = Iter_Valid(all_test_df, max_user=1000)
prev_test_df = None
prev_group = None
batch_user_ids = []
# reload all user group for cv2
with open(os.path.join(DATA_DIR, 'sakt_group_cv2.pickle'), 'rb') as f:
group = pickle.load(f)
for _ in range(n_iter):
test_df, sample_prediction_df = next(iter_test)
if prev_test_df is not None:
prev_test_df['answered_correctly'] = eval(test_df['prior_group_answers_correct'].iloc[0])
prev_test_df = prev_test_df[prev_test_df.content_type_id == False]
prev_group = prev_test_df[['user_id', 'content_id', 'answered_correctly']]\
.groupby('user_id').apply(lambda r: (
r['content_id'].values,
r['answered_correctly'].values))
for prev_user_id in prev_group.index:
prev_group_content = prev_group[prev_user_id][0]
prev_group_answered_correctly = prev_group[prev_user_id][1]
if prev_user_id in group.index:
group[prev_user_id] = (np.append(group[prev_user_id][0], prev_group_content),
np.append(group[prev_user_id][1], prev_group_answered_correctly))
else:
group[prev_user_id] = (prev_group_content, prev_group_answered_correctly)
if len(group[prev_user_id][0]) > MAX_SEQ:
new_group_content = group[prev_user_id][0][-MAX_SEQ:]
new_group_answered_correctly = group[prev_user_id][1][-MAX_SEQ:]
group[prev_user_id] = (new_group_content, new_group_answered_correctly)
prev_test_df = test_df.copy()
test_df = test_df[test_df.content_type_id == False]
batch_user_ids.append(test_df.user_id.unique())
test_dataset = TestDataset(group, test_df, NUM_SKILLS, max_seq=MAX_SEQ)
test_dataloader = DataLoader(test_dataset, batch_size=len(test_df), shuffle=False)
item = next(iter(test_dataloader))
x = item[0].to(device).long()
target_id = item[1].to(device).long()
with torch.no_grad():
output, _ = model(x, target_id)
output = torch.sigmoid(output)
preds = output[:, -1]
test_df['answered_correctly'] = preds.cpu().numpy()
set_predict(test_df.loc[test_df['content_type_id'] == 0,
['row_id', 'answered_correctly']])
return test_df, prev_test_df, output, item, group, prev_group, batch_user_ids
# ## Debugging notes
#
# Current set up, cv2_valid first 25k rows
# first 4 batches common `user_id`: 143316232, 1089397940, 1140583044 (placeholder user?)
#
group[1089397940]
# Iteration number 1 in the `iter_env`, the model gives the correct preds.
test_df, prev_test_df, output, item, group_updated, _, _ = iter_env_run(all_test_df, n_iter=1)
u_idx_loc = test_df.index.get_loc(test_df[test_df.user_id==1089397940].index[0])
print(f"local index of user 1089397940: {u_idx_loc}", '\n')
test_df.iloc[u_idx_loc]
test_df
# +
print(item[1][u_idx_loc, -12:]) # user 1089397940 first batch in example_test (question sequence)
print(item[0][u_idx_loc, -12:]) # user 1089397940 first batch in example_test: skill sequence = prev_content_id * (correct or not) + 13523
print(output[u_idx_loc, -12:].cpu().numpy(),'\n') # user 1089397940 probability prediction
print(group_updated[1089397940][0][:12]) # in the first iteration the length is only 11
print(group_updated[1089397940][1][:12])
# -
# Iteration number 2
test_df, prev_test_df, output, item, group_updated, _, _ = iter_env_run(all_test_df, n_iter=2)
u_idx_loc = test_df.index.get_loc(test_df[test_df.user_id==1089397940].index[0])
print(f"local index of user 1089397940: {u_idx_loc}", '\n')
test_df.iloc[u_idx_loc]
test_df
# +
print(item[1][u_idx_loc, -12:]) # user 1089397940 2nd batch in example_test (question sequence)
print(item[0][u_idx_loc, -12:]) # user 1089397940 2nd batch in example_test: skill sequence = prev_content_id * (correct or not) + 13523
print(output[u_idx_loc, -12:].cpu().numpy(),'\n') # user 1089397940 probability prediction
print(group_updated[1089397940][0][:12]) # in the 2nd iteration the length is only 11
print(group_updated[1089397940][1][:12])
# -
# Iteration number 3
test_df, prev_test_df, output, item, group_updated, _, _ = iter_env_run(all_test_df, n_iter=3)
u_idx_loc = test_df.index.get_loc(test_df[test_df.user_id==1089397940].index[0])
print(f"local index of user 1089397940: {u_idx_loc}", '\n')
test_df.iloc[u_idx_loc]
prev_test_df
prev_test_df['prior_group_answers_correct'].iloc[0]
# +
print(item[1][u_idx_loc, -12:]) # user 1089397940 first batch in example_test (question sequence)
print(item[0][u_idx_loc, -12:]) # user 1089397940 first batch in example_test: skill sequence = prev_content_id * (correct or not) + 13523
print(output[u_idx_loc, -12:].cpu().numpy(),'\n') # user 1089397940 probability prediction
print(group_updated[1089397940][0][:12]) # in the first iteration the length is only 11
print(group_updated[1089397940][1][:12])
# -
# Iteration number 4
test_df, prev_test_df, output, item, group_updated, _, _ = iter_env_run(all_test_df, n_iter=4)
u_idx_loc = test_df.index.get_loc(test_df[test_df.user_id==1089397940].index[0])
print(f"local index of user 1089397940: {u_idx_loc}", '\n')
test_df.iloc[u_idx_loc]
test_df
# +
print(item[1][u_idx_loc, -12:]) # user 1089397940 first batch in example_test (question sequence)
print(item[0][u_idx_loc, -12:]) # user 1089397940 first batch in example_test: skill sequence = prev_content_id * (correct or not) + 13523
print(output[u_idx_loc, -12:].cpu().numpy(),'\n') # user 1089397940 probability prediction
print(group_updated[1089397940][0][:12]) # in the first iteration the length is only 11
print(group_updated[1089397940][1][:12])
# -
prev_test_df
| sakt/debug_sakt_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: GeoPython
# language: python
# name: geopython
# ---
# # Geospatial Analysis for the Smart City
#
# Big Data BBQ 9/24/2021
#
# * https://github.com/gboeing/osmnx
# * https://osmnx.readthedocs.io/en/stable/
#
import folium
import osmnx as ox
import networkx as nx
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
import seaborn as sns
ox.config(use_cache=True, log_console=False)
# ### Retrieve Street Network from OSM Data
#
# * drive - get drivable public streets (but not service roads)
# * drive_service - get drivable streets, including service roads
# * walk - get all streets and paths that pedestrians can use (this network type ignores one-way directionality)
# * bike - get all streets and paths that cyclists can use
# * all - download all non-private OSM streets and paths
# * all_private - download all OSM streets and paths, including private-access ones
place = "Mannheim, Germany"
graph = ox.graph_from_place(place, network_type='drive')
fig, ax = ox.plot_graph(graph)
# #### Export als ESRI Shapefile
ox.save_graph_shapefile(graph, filepath='network-shape_mannheim')
nodes, streets = ox.graph_to_gdfs(graph)
len(streets)
streets.head()
street_types = pd.DataFrame(streets["highway"].apply(pd.Series)[0].value_counts().reset_index())
street_types.columns = ["type", "count"]
street_types
fig, ax = plt.subplots(figsize=(8,5))
sns.barplot(y="type", x="count", data=street_types, ax=ax)
plt.tight_layout()
plt.show();
# ### Display Street Network on a Map
# +
m = folium.Map([49.473, 8.475],zoom_start=13,tiles="CartoDB dark_matter")
jsondata = streets.to_json()
style = {'color': '#FFDD66',
'weight':'1'}
folium.GeoJson(jsondata, style_function=lambda x: style).add_to(m)
m
# -
# ### Routing
#
# Train Station
pos_1 = ox.geocode('Mannheim HBF, Mannheim, Germany')
pos_1
# MAFINEX
pos_2 = ox.geocode('Julius-Hatry-Straße 1, 68163 Mannheim, Germany')
pos_2
# #### Display on Map
# +
m = folium.Map([49.473, 8.475], zoom_start=13,tiles="CartoDB dark_matter")
#jsondata = streets.to_json()
style = {'color': '#FFDD66',
'weight':'1'}
folium.GeoJson(jsondata, style_function=lambda x: style).add_to(m)
folium.Marker(pos_1,
popup="Mannheim HBF",
icon=folium.Icon(color="green", prefix="fa", icon="train")).add_to(m)
folium.Marker(pos_2,
popup="MAFINEX",
icon=folium.Icon(color="red", prefix="fa", icon="star")).add_to(m)
m
# -
origin_node = ox.get_nearest_node(graph, pos_1)
destination_node = ox.get_nearest_node(graph, pos_2)
# #### Route finden
route = nx.shortest_path(graph, origin_node, destination_node)
fig, ax = ox.plot_graph_route(graph, route)
# +
from shapely.geometry import LineString
line = []
for i in route:
point = (graph.nodes[i]['x'],graph.nodes[i]['y'])
line.append(point)
print(point)
ls = LineString(line)
# -
# #### GeoDataFrame from LineString
#
gdf_route = pd.DataFrame(data= [['route 1',ls]],columns=['route','geometry'])
gdf_route
gdf_route = gpd.GeoDataFrame(gdf_route,geometry='geometry')
gdf_route
gdf_route.plot();
# Convert this route to GeoJSON
jsonroute = gdf_route.to_json()
# +
m = folium.Map([49.473, 8.475], zoom_start=13,tiles="CartoDB dark_matter")
#jsondata = streets.to_json()
style = {'color': '#FFDD66',
'weight':'1'}
route_style = {'color': '#FF0000',
'weight' : '4'}
folium.GeoJson(jsondata, style_function=lambda x: style).add_to(m)
folium.GeoJson(jsonroute, style_function=lambda x: route_style).add_to(m)
folium.Marker(pos_1,
popup="Mannheim HBF",
icon=folium.Icon(color="green", prefix="fa", icon="train")).add_to(m)
folium.Marker(pos_2,
popup="MAFINEX",
icon=folium.Icon(color="red", prefix="fa", icon="star")).add_to(m)
m
# -
# ### Building Footprints
# +
buildings = ox.geometries_from_place("Mannheim, Germany", tags={'building':True})
buildings.shape
# -
buildings.head()
ox.plot_footprints(buildings, figsize=(16,15));
# ### Retrieve some data
museum = buildings.query("tourism == 'museum'")
museum = museum[['name', 'geometry']]
museum.head(20)
ox.plot_footprints(museum, figsize=(16,15));
# ### Darstellung auf Karte
# +
import html
m = folium.Map([49.473, 8.475],zoom_start=15)
style = {'color': '#FF0000',
'fillColor': '#FFFF00',
'weight':'2'
}
def building(building):
jsondata = gpd.GeoSeries([building["geometry"]]).to_json()
name = html.escape(building["name"])
folium.GeoJson(jsondata, style_function=lambda x: style).add_child(folium.Popup(name)).add_to(m)
museum.apply(building, axis=1)
m.save("museums.html")
m
# -
# ### Marker from Buildings
# +
def get_geometry(x):
g = x["geometry"]
c = g.centroid
return c
centroid = museum.apply(get_geometry, axis=1)
# -
museum["centroid"] = centroid
museum.head()
# +
import html
m = folium.Map([49.473, 8.475],zoom_start=15)
def building(building):
lng = building["centroid"].x
lat = building["centroid"].y
name = html.escape(building["name"])
folium.Marker([lat,lng],
popup=name,
icon=folium.Icon(color="red", prefix="fa", icon="university")).add_to(m)
museum.apply(building, axis=1)
m
# -
# ### Get Points of interest from OSM
museum = ox.pois_from_place("Mannheim, Germany", tags={'tourism': 'museum'})
len(museum)
museum.head()
# +
import html
import shapely
m = folium.Map([49.473, 8.475],zoom_start=15)
def mus2poi(x):
if type(x["geometry"]) == shapely.geometry.point.Point:
lng = x["geometry"].x
lat = x["geometry"].y
folium.Marker([lat,lng],
popup=x["name"],
icon=folium.Icon(color="green", prefix="fa", icon="university")).add_to(m)
museum.apply(mus2poi, axis=1)
m
# -
| BigDataBBQMannheim.ipynb |