code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ordinal Regression
# Some data are discrete but instrinsically **ordered**, these are called [**ordinal**](https://en.wikipedia.org/wiki/Ordinal_data) data. One example is the [likert scale](https://en.wikipedia.org/wiki/Likert_scale) for questionairs ("this is an informative tutorial": 1. strongly disagree / 2. disagree / 3. neither agree nor disagree / 4. agree / 5. strongly agree). Ordinal data is also ubiquitous in the medical world (e.g. the [Glasgow Coma Scale](https://en.wikipedia.org/wiki/Glasgow_Coma_Scale) for measuring neurological disfunctioning).
#
# This poses a challenge for statistical modeling as the data do not fit the most well known modelling approaches (e.g. linear regression). Modeling the data as [categorical](https://en.wikipedia.org/wiki/Categorical_distribution) is one possibility, but it disregards the inherent ordering in the data, and may be less statistically efficient. There are multiple appoaches for modeling ordered data. Here we will show how to use the OrderedLogistic distribution using cutpoints that are sampled from a Normal distribution with as additional constrain that the cutpoints they are ordered. For a more in-depth discussion of Bayesian modeling of ordinal data, see e.g. [Michael Betancour's blog](https://betanalpha.github.io/assets/case_studies/ordinal_regression.html)
# !pip install -q numpyro@git+https://github.com/pyro-ppl/numpyro
from jax import numpy as np, random
import numpyro
from numpyro import sample
from numpyro.distributions import (Categorical, ImproperUniform, Normal, OrderedLogistic,
TransformedDistribution, constraints, transforms)
from numpyro.infer import MCMC, NUTS
import pandas as pd
import seaborn as sns
assert numpyro.__version__.startswith('0.7.1')
# First, generate some data with ordinal structure
# +
simkeys = random.split(random.PRNGKey(1), 2)
nsim = 50
nclasses = 3
Y = Categorical(logits=np.zeros(nclasses)).sample(simkeys[0], sample_shape=(nsim,))
X = Normal().sample(simkeys[1], sample_shape = (nsim,))
X += Y
print("value counts of Y:")
df = pd.DataFrame({'X': X, 'Y': Y})
print(df.Y.value_counts())
for i in range(nclasses):
print(f"mean(X) for Y == {i}: {X[np.where(Y==i)].mean():.3f}")
# -
sns.violinplot(x='Y', y='X', data=df);
# We will model the outcomes Y as coming from an OrderedLogistic distribution, conditional on X. The `OrderedLogistic` distribution in numpyro requires ordered cutpoints. We can use the `ImproperUnifrom` distribution to introduce a parameter with an arbitrary support that is otherwise completely uninformative, and then add an `ordered_vector` constraint.
# +
def model1(X, Y, nclasses=3):
b_X_eta = sample('b_X_eta', Normal(0, 5))
c_y = sample('c_y', ImproperUniform(support=constraints.ordered_vector,
batch_shape=(),
event_shape=(nclasses-1,)))
with numpyro.plate('obs', X.shape[0]):
eta = X * b_X_eta
sample('Y', OrderedLogistic(eta, c_y), obs=Y)
mcmc_key = random.PRNGKey(1234)
kernel = NUTS(model1)
mcmc = MCMC(kernel, num_warmup=250, num_samples=750)
mcmc.run(mcmc_key, X,Y, nclasses)
mcmc.print_summary()
# -
# The `ImproperUniform` distribution allows us to use parameters with constraints on their domain, without adding any additional information e.g. about the location or scale of the prior distribution on that parameter.
#
# If we want to incorporate such information, for instance that the values of the cut-points should not be too far from zero, we can add an additional `sample` statement that uses another prior, coupled with an `obs` argument. In the example below we first sample cutpoints `c_y` from the `ImproperUniform` distribution with `constraints.ordered_vector` as before, and then `sample` a dummy parameter from a `Normal` distribution while conditioning on `c_y` using `obs=c_y`. Effectively, we've created an improper / unnormalized prior that results from restricting the support of a `Normal` distribution to the ordered domain
# +
def model2(X, Y, nclasses=3):
b_X_eta = sample('b_X_eta', Normal(0, 5))
c_y = sample('c_y', ImproperUniform(support=constraints.ordered_vector,
batch_shape=(),
event_shape=(nclasses-1,)))
sample('c_y_smp', Normal(0,1), obs=c_y)
with numpyro.plate('obs', X.shape[0]):
eta = X * b_X_eta
sample('Y', OrderedLogistic(eta, c_y), obs=Y)
kernel = NUTS(model2)
mcmc = MCMC(kernel, num_warmup=250, num_samples=750)
mcmc.run(mcmc_key, X,Y, nclasses)
mcmc.print_summary()
# -
# If having a proper prior for those cutpoints `c_y` is desirable (e.g. to sample from that prior and get [prior predictive](https://en.wikipedia.org/wiki/Posterior_predictive_distribution#Prior_vs._posterior_predictive_distribution)), we can use [TransformedDistribution](http://num.pyro.ai/en/stable/distributions.html#transformeddistribution) with an [OrderedTransform](http://num.pyro.ai/en/stable/distributions.html#orderedtransform) transform as follows.
# +
def model3(X, Y, nclasses=3):
b_X_eta = sample('b_X_eta', Normal(0, 5))
c_y = sample("c_y", TransformedDistribution(Normal(0, 1).expand([nclasses - 1]),
transforms.OrderedTransform()))
with numpyro.plate('obs', X.shape[0]):
eta = X * b_X_eta
sample('Y', OrderedLogistic(eta, c_y), obs=Y)
kernel = NUTS(model3)
mcmc = MCMC(kernel, num_warmup=250, num_samples=750)
mcmc.run(mcmc_key, X,Y, nclasses)
mcmc.print_summary()
| notebooks/source/ordinal_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from Utils import Utils
from Constants import Constants
from PS_Matching import PS_Matching
from Propensity_socre_network import Propensity_socre_network
from Utils import Utils
from GAN import Generator, Discriminator
from sklearn.neighbors import NearestNeighbors
from GAN_Manager import GAN_Manager
from Utils import Utils
from matplotlib import pyplot
from torch.autograd.variable import Variable
from collections import OrderedDict
# +
from DCN import DCN
class DCN_network_1:
def train(self, train_parameters, device, train_mode=Constants.DCN_TRAIN_PD):
epochs = train_parameters["epochs"]
treated_batch_size = train_parameters["treated_batch_size"]
control_batch_size = train_parameters["control_batch_size"]
lr = train_parameters["lr"]
shuffle = train_parameters["shuffle"]
model_save_path = train_parameters["model_save_path"].format(epochs, lr)
treated_set_train = train_parameters["treated_set_train"]
control_set_train = train_parameters["control_set_train"]
input_nodes = train_parameters["input_nodes"]
print("Saved model path: {0}".format(model_save_path))
treated_data_loader_train = torch.utils.data.DataLoader(treated_set_train,
batch_size=treated_batch_size,
shuffle=shuffle,
num_workers=1)
control_data_loader_train = torch.utils.data.DataLoader(control_set_train,
batch_size=control_batch_size,
shuffle=shuffle,
num_workers=1)
network = DCN(training_mode=train_mode,
input_nodes=input_nodes).to(device)
optimizer = optim.Adam(network.parameters(), lr=lr)
lossF = nn.MSELoss()
min_loss = 100000.0
dataset_loss = 0.0
print(".. Training started ..")
print(device)
for epoch in range(epochs):
network.train()
total_loss = 0
train_set_size = 0
if epoch % 2 == 0:
dataset_loss = 0
# train treated
network.hidden1_Y1.weight.requires_grad = True
network.hidden1_Y1.bias.requires_grad = True
network.hidden2_Y1.weight.requires_grad = True
network.hidden2_Y1.bias.requires_grad = True
network.out_Y1.weight.requires_grad = True
network.out_Y1.bias.requires_grad = True
network.hidden1_Y0.weight.requires_grad = False
network.hidden1_Y0.bias.requires_grad = False
network.hidden2_Y0.weight.requires_grad = False
network.hidden2_Y0.bias.requires_grad = False
network.out_Y0.weight.requires_grad = False
network.out_Y0.bias.requires_grad = False
for batch in treated_data_loader_train:
covariates_X, ps_score, y_f, y_cf = batch
covariates_X = covariates_X.to(device)
ps_score = ps_score.squeeze().to(device)
train_set_size += covariates_X.size(0)
treatment_pred = network(covariates_X, ps_score)
# treatment_pred[0] -> y1
# treatment_pred[1] -> y0
yf_predicted = treatment_pred[0]
if torch.cuda.is_available():
loss = lossF(yf_predicted.float().cuda(),
y_f.float().cuda()).to(device)
else:
loss = lossF(yf_predicted.float(),
y_f.float()).to(device)
optimizer.zero_grad()
loss.backward()
optimizer.step()
total_loss += loss.item()
dataset_loss = total_loss
elif epoch % 2 == 1:
# train controlled
network.hidden1_Y1.weight.requires_grad = False
network.hidden1_Y1.bias.requires_grad = False
network.hidden2_Y1.weight.requires_grad = False
network.hidden2_Y1.bias.requires_grad = False
network.out_Y1.weight.requires_grad = False
network.out_Y1.bias.requires_grad = False
network.hidden1_Y0.weight.requires_grad = True
network.hidden1_Y0.bias.requires_grad = True
network.hidden2_Y0.weight.requires_grad = True
network.hidden2_Y0.bias.requires_grad = True
network.out_Y0.weight.requires_grad = True
network.out_Y0.bias.requires_grad = True
for batch in control_data_loader_train:
covariates_X, ps_score, y_f, y_cf = batch
covariates_X = covariates_X.to(device)
ps_score = ps_score.squeeze().to(device)
train_set_size += covariates_X.size(0)
treatment_pred = network(covariates_X, ps_score)
# treatment_pred[0] -> y1
# treatment_pred[1] -> y0
yf_predicted = treatment_pred[1]
if torch.cuda.is_available():
loss = lossF(yf_predicted.float().cuda(),
y_f.float().cuda()).to(device)
else:
loss = lossF(yf_predicted.float(),
y_f.float()).to(device)
optimizer.zero_grad()
loss.backward()
optimizer.step()
total_loss += loss.item()
dataset_loss = dataset_loss + total_loss
if epoch % 10 == 9:
print("epoch: {0}, Treated + Control loss: {1}".format(epoch, dataset_loss))
torch.save(network.state_dict(), model_save_path)
def eval(self, eval_parameters, device, input_nodes, train_mode):
treated_set = eval_parameters["treated_set"]
control_set = eval_parameters["control_set"]
model_path = eval_parameters["model_save_path"]
network = DCN(training_mode=train_mode, input_nodes=input_nodes).to(device)
network.load_state_dict(torch.load(model_path, map_location=device))
network.eval()
treated_data_loader = torch.utils.data.DataLoader(treated_set,
shuffle=False, num_workers=1)
control_data_loader = torch.utils.data.DataLoader(control_set,
shuffle=False, num_workers=1)
err_treated_list = []
err_control_list = []
true_ITE_list = []
predicted_ITE_list = []
ITE_dict_list = []
for batch in treated_data_loader:
covariates_X, ps_score, y_f, y_cf = batch
covariates_X = covariates_X.to(device)
ps_score = ps_score.squeeze().to(device)
treatment_pred = network(covariates_X, ps_score)
predicted_ITE = treatment_pred[0] - treatment_pred[1]
true_ITE = y_f - y_cf
if torch.cuda.is_available():
diff = true_ITE.float().cuda() - predicted_ITE.float().cuda()
else:
diff = true_ITE.float() - predicted_ITE.float()
ITE_dict_list.append(self.create_ITE_Dict(covariates_X,
ps_score.item(), y_f.item(),
y_cf.item(),
true_ITE.item(),
predicted_ITE.item(),
diff.item()))
err_treated_list.append(diff.item())
true_ITE_list.append(true_ITE.item())
predicted_ITE_list.append(predicted_ITE.item())
for batch in control_data_loader:
covariates_X, ps_score, y_f, y_cf = batch
covariates_X = covariates_X.to(device)
ps_score = ps_score.squeeze().to(device)
treatment_pred = network(covariates_X, ps_score)
predicted_ITE = treatment_pred[0] - treatment_pred[1]
true_ITE = y_cf - y_f
if torch.cuda.is_available():
diff = true_ITE.float().cuda() - predicted_ITE.float().cuda()
else:
diff = true_ITE.float() - predicted_ITE.float()
ITE_dict_list.append(self.create_ITE_Dict(covariates_X,
ps_score.item(), y_f.item(),
y_cf.item(),
true_ITE.item(),
predicted_ITE.item(),
diff.item()))
err_control_list.append(diff.item())
true_ITE_list.append(true_ITE.item())
predicted_ITE_list.append(predicted_ITE.item())
# print(err_treated_list)
# print(err_control_list)
return {
"treated_err": err_treated_list,
"control_err": err_control_list,
"true_ITE": true_ITE_list,
"predicted_ITE": predicted_ITE_list,
"ITE_dict_list": ITE_dict_list
}
@staticmethod
def eval_semi_supervised(eval_parameters, device, input_nodes, train_mode, treated_flag):
eval_set = eval_parameters["eval_set"]
model_path = eval_parameters["model_save_path"]
network = DCN(training_mode=train_mode, input_nodes=input_nodes).to(device)
network.load_state_dict(torch.load(model_path, map_location=device))
network.eval()
treated_data_loader = torch.utils.data.DataLoader(eval_set,
shuffle=False, num_workers=1)
y_f_list = []
y_cf_list = []
for batch in treated_data_loader:
covariates_X, ps_score = batch
covariates_X = covariates_X.to(device)
ps_score = ps_score.squeeze().to(device)
treatment_pred = network(covariates_X, ps_score)
if treated_flag:
y_f_list.append(treatment_pred[0].item())
y_cf_list.append(treatment_pred[1].item())
else:
y_f_list.append(treatment_pred[1].item())
y_cf_list.append(treatment_pred[0].item())
return {
"y_f_list": np.array(y_f_list),
"y_cf_list": np.array(y_cf_list)
}
@staticmethod
def create_ITE_Dict(covariates_X, ps_score, y_f, y_cf, true_ITE,
predicted_ITE, diff):
result_dict = OrderedDict()
covariate_list = [element.item() for element in covariates_X.flatten()]
idx = 0
for item in covariate_list:
idx += 1
result_dict["X" + str(idx)] = item
result_dict["ps_score"] = ps_score
result_dict["factual"] = y_f
result_dict["counter_factual"] = y_cf
result_dict["true_ITE"] = true_ITE
result_dict["predicted_ITE"] = predicted_ITE
result_dict["diff"] = diff
return result_dict
# +
def get_propensity_scores(ps_train_set, iter_id, input_nodes, device):
prop_score_NN_model_path = Constants.PROP_SCORE_NN_MODEL_PATH \
.format(iter_id, Constants.PROP_SCORE_NN_EPOCHS, Constants.PROP_SCORE_NN_LR)
train_parameters_NN = {
"epochs": Constants.PROP_SCORE_NN_EPOCHS,
"lr": Constants.PROP_SCORE_NN_LR,
"batch_size": Constants.PROP_SCORE_NN_BATCH_SIZE,
"shuffle": True,
"train_set": ps_train_set,
"model_save_path": prop_score_NN_model_path,
"input_nodes": input_nodes
}
# ps using NN
ps_net_NN = Propensity_socre_network()
print("############### Propensity Score neural net Training ###############")
ps_net_NN.train(train_parameters_NN, device, phase="train")
# eval
eval_parameters_train_NN = {
"eval_set": ps_train_set,
"model_path": prop_score_NN_model_path,
"input_nodes": input_nodes
}
ps_score_list_train_NN = ps_net_NN.eval(eval_parameters_train_NN, device, phase="eval")
return ps_score_list_train_NN
# +
import numpy as np
from sklearn.neighbors import NearestNeighbors
from GAN_Manager import GAN_Manager
from Utils import Utils
def get_matched_and_unmatched_control_indices(ps_treated, ps_control):
nn = NearestNeighbors(n_neighbors=1)
nn.fit(ps_control)
distance, matched_control = nn.kneighbors(ps_treated)
matched_control_indices = np.array(matched_control).ravel()
# # remove duplicates
#matched_control_indices = list(dict.fromkeys(matched_control_indices))
set_matched_control_indices = set(matched_control_indices)
total_indices = list(range(len(ps_control)))
unmatched_control_indices = list(filter(lambda x: x not in set_matched_control_indices,
total_indices))
return matched_control_indices, unmatched_control_indices
def filter_control_groups(np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf, indices):
np_filter_control_df_X = np.take(np_control_df_X, indices, axis=0)
np_filter_control_ps_score = np.take(np_control_ps_score, indices, axis=0)
np_filter_control_df_Y_f = np.take(np_control_df_Y_f, indices, axis=0)
np_filter_control_df_Y_cf = np.take(np_control_df_Y_cf, indices, axis=0)
tuple_matched_control = (np_filter_control_df_X, np_filter_control_ps_score,
np_filter_control_df_Y_f, np_filter_control_df_Y_cf)
return tuple_matched_control
def filter_matched_and_unmatched_control_samples(np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf, matched_control_indices,
unmatched_control_indices):
tuple_matched_control = filter_control_groups(np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf,
matched_control_indices)
tuple_unmatched_control = filter_control_groups(np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf,
unmatched_control_indices)
return tuple_matched_control, tuple_unmatched_control
# -
iter_id = 10
# +
csv_path = "Dataset/ihdp_sample.csv"
from dataloader import DataLoader
split_size = 0.8
dL = DataLoader()
input_nodes = 25
device = Utils.get_device()
np_covariates_X_train, np_covariates_X_test, np_covariates_Y_train, np_covariates_Y_test \
= dL.preprocess_data_from_csv(csv_path, split_size)
ps_train_set = dL.convert_to_tensor(np_covariates_X_train, np_covariates_Y_train)
ps_score_list_train_NN = get_propensity_scores(ps_train_set, iter_id, input_nodes, device)
# -
data_loader_dict_train = dL.prepare_tensor_for_DCN(np_covariates_X_train,
np_covariates_Y_train,
ps_score_list_train_NN,
False)
# +
tuple_treated = data_loader_dict_train["treated_data"]
tuple_control = data_loader_dict_train["control_data"]
np_treated_df_X, np_treated_ps_score, np_treated_df_Y_f, np_treated_df_Y_cf = tuple_treated
np_control_df_X, np_control_ps_score, np_control_df_Y_f, np_control_df_Y_cf = tuple_control
# get unmatched controls
matched_control_indices, unmatched_control_indices = get_matched_and_unmatched_control_indices(
Utils.convert_to_col_vector(np_treated_ps_score),
Utils.convert_to_col_vector(np_control_ps_score))
print("Matched Control: {0}".format(len(matched_control_indices)))
print("Unmatched Control: {0}".format(len(unmatched_control_indices)))
tuple_matched_control, tuple_unmatched_control = filter_matched_and_unmatched_control_samples(
np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf, matched_control_indices,
unmatched_control_indices)
# generate matched treated for unmatched controls using variable
# tuple_unmatched_control
# create GAN code here
print("Matched Control: {0}".format(tuple_matched_control[0].shape))
print("Matched Treated: {0}".format(tuple_treated[0].shape))
tensor_treated = \
Utils.create_tensors_to_train_DCN(tuple_treated, dL)
# need to change for unmatched
tensor_matched_control = \
Utils.create_tensors_to_train_DCN(tuple_matched_control, dL)
tensor_unmatched_control = \
Utils.create_tensors_to_train_DCN(tuple_unmatched_control, dL)
# -
def draw(treated_ps_list, control_ps_list, bins1):
pyplot.hist(treated_ps_list, bins1, alpha=0.5, label='treated')
pyplot.hist(control_ps_list, bins1, alpha=0.5, label='control')
pyplot.legend(loc='upper right')
pyplot.show()
# +
ps_matched_control_list = tuple_matched_control[1].tolist()
ps_un_matched_control_list = tuple_unmatched_control[1].tolist()
ps_treated_list = tuple_treated[1].tolist()
print(len(ps_matched_control_list), len(ps_treated_list))
# -
bins1 = np.linspace(0, 1, 100)
bins2 = np.linspace(0, 0.2, 100)
bins3 = np.linspace(0.2, 0.5, 100)
bins4 = np.linspace(0.5, 1, 100)
# matched control and treated
draw(ps_treated_list, ps_matched_control_list, bins1)
# unmatched control and treated
draw(ps_treated_list, ps_un_matched_control_list, bins1)
class GAN_Module:
def __init__(self, discriminator_in_nodes, generator_out_nodes, device):
self.discriminator = Discriminator(in_nodes=discriminator_in_nodes).to(device)
self.discriminator.apply(self.weights_init)
self.generator = Generator(out_nodes=generator_out_nodes).to(device)
self.generator.apply(self.weights_init)
self.loss = nn.BCELoss()
def get_generator(self):
return self.generator
def __cal_propensity_loss(self, ps_score_control, prop_score_NN_model_path, gen_treated, device):
# Assign treated
Y = np.ones(gen_treated.size(0))
eval_set = Utils.convert_to_tensor(gen_treated.detach().numpy(), Y)
eval_parameters_ps_net = {
"eval_set": eval_set,
"model_path": prop_score_NN_model_path,
"input_nodes": 25
}
ps_net_NN = Propensity_socre_network()
ps_score_list_treated = ps_net_NN.eval(eval_parameters_ps_net, device,
phase="eval",
eval_from_GAN=True)
Tensor = torch.cuda.DoubleTensor if torch.cuda.is_available() else torch.DoubleTensor
ps_score_treated = Tensor(ps_score_list_treated)
prop_loss = torch.sum((torch.sub(ps_score_treated, ps_score_control)) ** 2)
return prop_loss
def noise(self, _size):
n = Variable(torch.normal(mean=0, std=1, size=(_size, 25)))
# print(n.size())
if torch.cuda.is_available(): return n.cuda()
return n
def weights_init(self, m):
if type(m) == nn.Linear:
nn.init.xavier_uniform_(m.weight)
torch.nn.init.zeros_(m.bias)
def real_data_target(self, size):
'''
Tensor containing ones, with shape = size
'''
data = Variable(torch.ones(size, 1))
if torch.cuda.is_available(): return data.cuda()
return data
def fake_data_target(self, size):
'''
Tensor containing zeros, with shape = size
'''
data = Variable(torch.zeros(size, 1))
if torch.cuda.is_available(): return data.cuda()
return data
def train_discriminator(self, optimizer, real_data, fake_data):
# Reset gradients
optimizer.zero_grad()
# 1.1 Train on Real Data
prediction_real = self.discriminator(real_data)
real_score = torch.mean(prediction_real).item()
# Calculate error and backpropagate
error_real = self.loss(prediction_real, self.real_data_target(real_data.size(0)))
error_real.backward()
# 1.2 Train on Fake Data
prediction_fake = self.discriminator(fake_data)
fake_score = torch.mean(prediction_fake).item()
# Calculate error and backpropagate
error_fake = self.loss(prediction_fake, self.fake_data_target(real_data.size(0)))
error_fake.backward()
# 1.3 Update weights with gradients
optimizer.step()
loss_D = error_real + error_fake
# Return error
return loss_D.item(), real_score, fake_score
def train_generator(self, optimizer, fake_data, ALPHA, ps_score_control,
prop_score_NN_model_path, device):
# 2. Train Generator
# Reset gradients
optimizer.zero_grad()
# Sample noise and generate fake data
predicted_D = self.discriminator(fake_data)
# Calculate error and back propagate
error_g = self.loss(predicted_D, self.real_data_target(predicted_D.size(0)))
prop_loss = self.__cal_propensity_loss(ps_score_control, prop_score_NN_model_path,
fake_data, device)
error = error_g + (ALPHA * prop_loss)
error.backward()
# Update weights with gradients
optimizer.step()
# Return error
return error_g.item(), prop_loss.item()
def train_GAN(self, train_parameters, device):
epochs = train_parameters["epochs"]
train_set = train_parameters["train_set"]
lr = train_parameters["lr"]
shuffle = train_parameters["shuffle"]
batch_size = train_parameters["batch_size"]
prop_score_NN_model_path = train_parameters["prop_score_NN_model_path"]
ALPHA = train_parameters["ALPHA"]
data_loader_train = torch.utils.data.DataLoader(train_set,
batch_size=batch_size,
shuffle=shuffle,
num_workers=1)
# generator = Generator(out_nodes=generator_out_nodes).to(device)
# discriminator = Discriminator(in_nodes=discriminator_in_nodes).to(device)
g_optimizer = optim.Adam(self.generator.parameters(), lr=lr)
d_optimizer = optim.Adam(self.discriminator.parameters(), lr=lr)
for epoch in range(epochs):
epoch += 1
# self.generator.train()
# self.discriminator.train()
total_G_loss = 0
total_D_loss = 0
total_prop_loss = 0
total_d_pred_real = 0
total_d_pred_fake = 0
for batch in data_loader_train:
covariates_X_control, ps_score_control, y_f, y_cf = batch
covariates_X_control = covariates_X_control.to(device)
covariates_X_control_size = covariates_X_control.size(0)
ps_score_control = ps_score_control.squeeze().to(device)
# 1. Train Discriminator
real_data = covariates_X_control
# Generate fake data
fake_data = self.generator(self.noise(covariates_X_control_size)).detach()
# Train D
d_error, d_pred_real, d_pred_fake = self.train_discriminator(d_optimizer,
real_data, fake_data)
total_D_loss += d_error
total_d_pred_real += d_pred_real
total_d_pred_fake += d_pred_fake
# 2. Train Generator
# Generate fake data
fake_data = self.generator(self.noise(covariates_X_control_size))
# Train G
error_g, prop_loss = self.train_generator(g_optimizer, fake_data, ALPHA, ps_score_control,
prop_score_NN_model_path, device)
total_G_loss += error_g
total_prop_loss += prop_loss
if epoch % 1000 == 0:
print("Epoch: {0}, D_loss: {1}, D_score_real: {2}, D_score_Fake: {3}, G_loss: {4}, "
"Prop_loss: {5}"
.format(epoch,
total_D_loss, total_d_pred_real, total_d_pred_fake, total_G_loss, total_prop_loss))
# +
# GAN Part from here
prop_score_NN_model_path = Constants.PROP_SCORE_NN_MODEL_PATH \
.format(iter_id, Constants.PROP_SCORE_NN_EPOCHS, Constants.PROP_SCORE_NN_LR)
gan = GAN_Module(discriminator_in_nodes=25, generator_out_nodes=25, device=device)
GAN_train_parameters = {
"epochs": 10000,
"lr": 0.0002,
"shuffle": True,
"train_set": tensor_unmatched_control,
"batch_size": 64,
"prop_score_NN_model_path": prop_score_NN_model_path,
"ALPHA": 1.2
}
gan.train_GAN(GAN_train_parameters, device=device)
# +
def noise(_size):
n = Variable(torch.normal(mean=0, std=1, size=(_size, 25)))
# print(n.size())
if torch.cuda.is_available(): return n.cuda()
return n
def eval_GAN(eval_size, generator, device):
treated_g = generator(noise(eval_size))
return treated_g
# -
eval_size = len(unmatched_control_indices)
gen_net = gan.get_generator()
treated_generated = eval_GAN(eval_size, gen_net, device)
print("eval_size: " + str(eval_size))
print("Treated G size")
print(treated_generated.size())
# +
Y = np.ones(treated_generated.size(0))
eval_set = Utils.convert_to_tensor(treated_generated.detach().numpy(), Y)
eval_parameters_ps_net = {
"eval_set": eval_set,
"model_path": prop_score_NN_model_path,
"input_nodes": 25
}
ps_net_NN = Propensity_socre_network()
ps_score_list_treated = ps_net_NN.eval(eval_parameters_ps_net, device,
phase="eval", eval_from_GAN=True)
len(ps_score_list_treated)
# -
# treated by GAN vs unmactched control
draw(ps_score_list_treated, ps_un_matched_control_list, bins1)
epochs = 100
lr = 0.001
def execute_DCN_train(tensor_treated_train, tensor_control_train, model_path, iter_id,
input_nodes, device, train_mode):
DCN_train_parameters = {
"epochs": epochs,
"lr": lr,
"treated_batch_size": 1,
"control_batch_size": 1,
"shuffle": True,
"treated_set_train": tensor_treated_train,
"control_set_train": tensor_control_train,
"model_save_path": model_path.format(iter_id,
epochs,
lr),
"input_nodes": input_nodes
}
# train DCN network
dcn = DCN_network_1()
dcn.train(DCN_train_parameters, device, train_mode=train_mode)
# +
print("### DCN semi supervised training using PS Matching No PD###")
model_path_no_dropout = "./DCNModel/NN_DCN_SEMI_SUPERVISED_NO_DROPOUT_PM_MATCH_FALSE_model_iter_id_{0}_epoch_{1}_lr_{2}.pth"
train_mode = Constants.DCN_TRAIN_NO_DROPOUT
execute_DCN_train(tensor_treated,
tensor_matched_control,
model_path_no_dropout, iter_id,
input_nodes, device, train_mode)
# t_treated = Utils.create_tensors_to_train_DCN(data_loader_dict_train["treated_data"], dL)
# t_control = Utils.create_tensors_to_train_DCN(data_loader_dict_train["control_data"], dL)
# train_mode = Constants.DCN_TRAIN_PD
# execute_DCN_train(t_treated,
# t_control,
# model_path_no_dropout,
# iter_id,
# input_nodes,
# device,
# train_mode)
# +
## to be added in the code
def convert_to_tensor_DCN_PS(tensor_x, ps_score):
tensor_ps_score = torch.from_numpy(ps_score)
# tensor_x = torch.stack([torch.Tensor(i) for i in X])
processed_dataset = torch.utils.data.TensorDataset(tensor_x, tensor_ps_score)
return processed_dataset
def eval_semi_supervised(eval_parameters, device, input_nodes, train_mode, treated_flag):
eval_set = eval_parameters["eval_set"]
model_path = eval_parameters["model_save_path"]
network = DCN(training_mode=train_mode, input_nodes=input_nodes).to(device)
network.load_state_dict(torch.load(model_path, map_location=device))
network.eval()
treated_data_loader = torch.utils.data.DataLoader(eval_set,
shuffle=False, num_workers=1)
y_f_list = []
y_cf_list = []
for batch in treated_data_loader:
covariates_X, ps_score = batch
covariates_X = covariates_X.to(device)
ps_score = ps_score.squeeze().to(device)
treatment_pred = network(covariates_X, ps_score)
if treated_flag:
y_f_list.append(treatment_pred[0].item())
y_cf_list.append(treatment_pred[1].item())
else:
y_f_list.append(treatment_pred[1].item())
y_cf_list.append(treatment_pred[0].item())
# print(err_treated_list)
# print(err_control_list)
return {
"y_f_list": np.array(y_f_list),
"y_cf_list": np.array(y_cf_list)
}
# +
ps_score_list_treated_np = np.array(ps_score_list_treated)
eval_set = convert_to_tensor_DCN_PS(treated_generated.detach(), ps_score_list_treated_np)
DCN_test_parameters = {
"eval_set": eval_set,
"model_save_path": model_path_no_dropout.format(iter_id,
epochs,
lr)
}
treated_gen_y = eval_semi_supervised(DCN_test_parameters, device, input_nodes,
Constants.DCN_EVALUATION, treated_flag=True)
# +
np_treated_generated = treated_generated.detach().numpy()
np_ps_score_list_gen_treated = ps_score_list_treated_np
np_treated_gen_f = Utils.convert_to_col_vector(treated_gen_y["y_f_list"])
np_treated_gen_cf = Utils.convert_to_col_vector(treated_gen_y["y_cf_list"])
print(np_treated_gen_f.shape)
np_original_X = tuple_treated[0]
np_original_ps_score = tuple_treated[1]
np_original_Y_f = tuple_treated[2]
np_original_Y_cf = tuple_treated[3]
print(np_original_Y_f.shape)
np_treated_x = np.concatenate((np_treated_generated, np_original_X), axis=0)
np_treated_ps = np.concatenate((np_ps_score_list_gen_treated, np_original_ps_score), axis=0)
np_treated_f = np.concatenate((np_treated_gen_f, np_original_Y_f), axis=0)
np_treated_cf = np.concatenate((np_treated_gen_cf, np_original_Y_cf), axis=0)
tensor_treated = Utils.convert_to_tensor_DCN(np_treated_x, np_treated_ps,
np_treated_f, np_treated_cf)
np_treated_x.shape
# +
np_control_unmatched_X = tuple_unmatched_control[0]
np_ps_score_list_control_unmatched = tuple_unmatched_control[1]
np_control_unmatched_f = tuple_unmatched_control[2]
np_control_unmatched_cf = tuple_unmatched_control[3]
np_control_matched_X = tuple_matched_control[0]
np_ps_score_list_control_matched = tuple_matched_control[1]
np_control_matched_f = tuple_matched_control[2]
np_control_matched_cf = tuple_matched_control[3]
print(np_control_unmatched_cf.shape)
print(np_control_matched_cf.shape)
np_control_x = np.concatenate((np_control_unmatched_X, np_control_matched_X), axis=0)
np_control_ps = np.concatenate((np_ps_score_list_control_unmatched, np_ps_score_list_control_matched), axis=0)
np_control_f = np.concatenate((np_control_unmatched_f, np_control_matched_f), axis=0)
np_control_cf = np.concatenate((np_control_unmatched_cf, np_control_matched_cf), axis=0)
tensor_control = Utils.convert_to_tensor_DCN(np_control_x, np_control_ps,
np_control_f, np_control_cf)
np_control_x.shape
# -
tuple_control_train = (np_control_x, np_control_ps, np_control_f, np_control_cf)
tuple_treated_train = (np_treated_x, np_treated_ps, np_treated_f, np_treated_cf)
# +
print("### DCN training using all dataset no PD ###")
train_mode = Constants.DCN_TRAIN_NO_DROPOUT
model_path = "./DCNModel/NN_DCN_NO_DROPOUT_PM_MATCH_FALSE_model_iter_id_{0}_epoch_{1}_lr_{2}.pth"
execute_DCN_train(tensor_treated,
tensor_control,
model_path,
iter_id,
input_nodes,
device,
train_mode)
# +
# testing
print("----------- Testing phase ------------")
dL = DataLoader()
ps_test_set = dL.convert_to_tensor(np_covariates_X_test,
np_covariates_Y_test)
is_synthetic = False
input_nodes = 25
# get propensity scores using NN
ps_net_NN = Propensity_socre_network()
ps_eval_parameters_NN = {
"eval_set": ps_test_set,
"model_path": Constants.PROP_SCORE_NN_MODEL_PATH
.format(iter_id, Constants.PROP_SCORE_NN_EPOCHS, Constants.PROP_SCORE_NN_LR),
"input_nodes": input_nodes
}
ps_score_list_NN = ps_net_NN.eval(ps_eval_parameters_NN, device, phase="eval")
data_loader_dict = dL.prepare_tensor_for_DCN(np_covariates_X_test,
np_covariates_Y_test,
ps_score_list_NN,
is_synthetic)
model_path = model_path.format(iter_id,
epochs,
lr)
print(model_path)
treated_group = data_loader_dict["treated_data"]
np_treated_df_X = treated_group[0]
np_treated_ps_score = treated_group[1]
np_treated_df_Y_f = treated_group[2]
np_treated_df_Y_cf = treated_group[3]
tensor_treated = Utils.convert_to_tensor_DCN(np_treated_df_X, np_treated_ps_score,
np_treated_df_Y_f, np_treated_df_Y_cf)
control_group = data_loader_dict["control_data"]
np_control_df_X = control_group[0]
np_control_ps_score = control_group[1]
np_control_df_Y_f = control_group[2]
np_control_df_Y_cf = control_group[3]
tensor_control = Utils.convert_to_tensor_DCN(np_control_df_X, np_control_ps_score,
np_control_df_Y_f, np_control_df_Y_cf)
DCN_test_parameters = {
"treated_set": tensor_treated,
"control_set": tensor_control,
"model_save_path": model_path
}
dcn = DCN_network_1()
response_dict = dcn.eval(DCN_test_parameters, device, input_nodes,
Constants.DCN_EVALUATION)
err_treated = [ele ** 2 for ele in response_dict["treated_err"]]
err_control = [ele ** 2 for ele in response_dict["control_err"]]
total_sum = sum(err_treated) + sum(err_control)
total_item = len(err_treated) + len(err_control)
MSE = total_sum / total_item
print("PSM")
print(MSE)
# +
print("### DCN training using all dataset DCN PD ###")
train_mode = Constants.DCN_TRAIN_PD
model_path = "./DCNModel/DCN_PD_model_iter_id_{0}_epoch_{1}_lr_{2}.pth"
t_treated = Utils.create_tensors_to_train_DCN(data_loader_dict_train["treated_data"], dL)
t_control = Utils.create_tensors_to_train_DCN(data_loader_dict_train["control_data"], dL)
execute_DCN_train(t_treated,
t_control,
model_path,
iter_id,
input_nodes,
device,
train_mode)
# +
# testing
print("----------- Testing phase ------------")
dL = DataLoader()
ps_test_set = dL.convert_to_tensor(np_covariates_X_test,
np_covariates_Y_test)
is_synthetic = False
input_nodes = 25
# get propensity scores using NN
ps_net_NN = Propensity_socre_network()
ps_eval_parameters_NN = {
"eval_set": ps_test_set,
"model_path": Constants.PROP_SCORE_NN_MODEL_PATH
.format(iter_id, Constants.PROP_SCORE_NN_EPOCHS, Constants.PROP_SCORE_NN_LR),
"input_nodes": input_nodes
}
ps_score_list_NN = ps_net_NN.eval(ps_eval_parameters_NN, device, phase="eval")
data_loader_dict = dL.prepare_tensor_for_DCN(np_covariates_X_test,
np_covariates_Y_test,
ps_score_list_NN,
is_synthetic)
model_path = model_path.format(iter_id,
epochs,
lr)
print(model_path)
treated_group = data_loader_dict["treated_data"]
np_treated_df_X = treated_group[0]
np_treated_ps_score = treated_group[1]
np_treated_df_Y_f = treated_group[2]
np_treated_df_Y_cf = treated_group[3]
tensor_treated = Utils.convert_to_tensor_DCN(np_treated_df_X, np_treated_ps_score,
np_treated_df_Y_f, np_treated_df_Y_cf)
control_group = data_loader_dict["control_data"]
np_control_df_X = control_group[0]
np_control_ps_score = control_group[1]
np_control_df_Y_f = control_group[2]
np_control_df_Y_cf = control_group[3]
tensor_control = Utils.convert_to_tensor_DCN(np_control_df_X, np_control_ps_score,
np_control_df_Y_f, np_control_df_Y_cf)
DCN_test_parameters = {
"treated_set": tensor_treated,
"control_set": tensor_control,
"model_save_path": model_path
}
dcn = DCN_network_1()
response_dict = dcn.eval(DCN_test_parameters, device, input_nodes,
Constants.DCN_EVALUATION)
err_treated = [ele ** 2 for ele in response_dict["treated_err"]]
err_control = [ele ** 2 for ele in response_dict["control_err"]]
total_sum = sum(err_treated) + sum(err_control)
total_item = len(err_treated) + len(err_control)
MSE = total_sum / total_item
print("DCN_PD")
print(MSE)
# -
# # TARNET
# +
class TARNetPhi(nn.Module):
def __init__(self, input_nodes, shared_nodes=200):
super(TARNetPhi, self).__init__()
# shared layer
self.shared1 = nn.Linear(in_features=input_nodes, out_features=shared_nodes)
nn.init.xavier_uniform_(self.shared1.weight)
nn.init.zeros_(self.shared1.bias)
self.shared2 = nn.Linear(in_features=shared_nodes, out_features=shared_nodes)
nn.init.xavier_uniform_(self.shared2.weight)
nn.init.zeros_(self.shared2.bias)
self.shared3 = nn.Linear(in_features=shared_nodes, out_features=shared_nodes)
nn.init.xavier_uniform_(self.shared3.weight)
nn.init.zeros_(self.shared3.bias)
def forward(self, x):
if torch.cuda.is_available():
x = x.float().cuda()
else:
x = x.float()
# shared layers
x = F.relu(self.shared1(x))
x = F.relu(self.shared2(x))
x = F.relu(self.shared3(x))
return x
class TARNetH_Y1(nn.Module):
def __init__(self, input_nodes=200, outcome_nodes=100):
super(TARNetH_Y1, self).__init__()
# potential outcome1 Y(1)
self.hidden1_Y1 = nn.Linear(in_features=input_nodes, out_features=outcome_nodes)
nn.init.xavier_uniform_(self.hidden1_Y1.weight)
nn.init.zeros_(self.hidden1_Y1.bias)
self.hidden2_Y1 = nn.Linear(in_features=outcome_nodes, out_features=outcome_nodes)
nn.init.xavier_uniform_(self.hidden2_Y1.weight)
nn.init.zeros_(self.hidden2_Y1.bias)
self.out_Y1 = nn.Linear(in_features=outcome_nodes, out_features=1)
nn.init.xavier_uniform_(self.out_Y1.weight)
nn.init.zeros_(self.out_Y1.bias)
def forward(self, x):
if torch.cuda.is_available():
x = x.float().cuda()
else:
x = x.float()
# potential outcome1 Y(1)
y1 = F.relu(self.hidden1_Y1(x))
y1 = F.relu(self.hidden2_Y1(y1))
y1 = self.out_Y1(y1)
return y1
class TARNetH_Y0(nn.Module):
def __init__(self, input_nodes=200, outcome_nodes=100):
super(TARNetH_Y0, self).__init__()
# potential outcome1 Y(0)
self.hidden1_Y0 = nn.Linear(in_features=input_nodes, out_features=outcome_nodes)
nn.init.xavier_uniform_(self.hidden1_Y0.weight)
nn.init.zeros_(self.hidden1_Y0.bias)
self.hidden2_Y0 = nn.Linear(in_features=outcome_nodes, out_features=outcome_nodes)
nn.init.xavier_uniform_(self.hidden2_Y0.weight)
nn.init.zeros_(self.hidden2_Y0.bias)
self.out_Y0 = nn.Linear(in_features=outcome_nodes, out_features=1)
nn.init.xavier_uniform_(self.out_Y0.weight)
nn.init.zeros_(self.out_Y0.bias)
def forward(self, x):
if torch.cuda.is_available():
x = x.float().cuda()
else:
x = x.float()
# potential outcome1 Y(0)
y0 = F.relu(self.hidden1_Y0(x))
y0 = F.relu(self.hidden2_Y0(y0))
y0 = self.out_Y0(y0)
return y0
# -
class PS_Matching:
def match_using_prop_score(self, tuple_treated, tuple_control):
matched_controls = []
# do ps match
np_treated_df_X, np_treated_ps_score, np_treated_df_Y_f, np_treated_df_Y_cf = tuple_treated
np_control_df_X, np_control_ps_score, np_control_df_Y_f, np_control_df_Y_cf = tuple_control
# get unmatched controls
matched_control_indices, unmatched_control_indices = self.get_matched_and_unmatched_control_indices(
Utils.convert_to_col_vector(np_treated_ps_score),
Utils.convert_to_col_vector(np_control_ps_score))
tuple_matched_control, tuple_unmatched_control = self.filter_matched_and_unmatched_control_samples(
np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf, matched_control_indices,
unmatched_control_indices)
return tuple_matched_control
def filter_matched_and_unmatched_control_samples(self, np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf, matched_control_indices,
unmatched_control_indices):
tuple_matched_control = self.filter_control_groups(np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf,
matched_control_indices)
tuple_unmatched_control = self.filter_control_groups(np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf,
unmatched_control_indices)
return tuple_matched_control, tuple_unmatched_control
@staticmethod
def filter_control_groups(np_control_df_X, np_control_ps_score,
np_control_df_Y_f,
np_control_df_Y_cf, indices):
np_filter_control_df_X = np.take(np_control_df_X, indices, axis=0)
np_filter_control_ps_score = np.take(np_control_ps_score, indices, axis=0)
np_filter_control_df_Y_f = np.take(np_control_df_Y_f, indices, axis=0)
np_filter_control_df_Y_cf = np.take(np_control_df_Y_cf, indices, axis=0)
tuple_matched_control = (np_filter_control_df_X, np_filter_control_ps_score,
np_filter_control_df_Y_f, np_filter_control_df_Y_cf)
return tuple_matched_control
@staticmethod
def get_matched_and_unmatched_control_indices(ps_treated, ps_control):
nn = NearestNeighbors(n_neighbors=1)
nn.fit(ps_control)
distance, matched_control = nn.kneighbors(ps_treated)
matched_control_indices = np.array(matched_control).ravel()
# remove duplicates
# matched_control_indices = list(dict.fromkeys(matched_control_indices))
set_matched_control_indices = set(matched_control_indices)
total_indices = list(range(len(ps_control)))
unmatched_control_indices = list(filter(lambda x: x not in set_matched_control_indices,
total_indices))
return matched_control_indices, unmatched_control_indices
@staticmethod
def get_unmatched_prop_list(tensor_unmatched_control):
control_data_loader_train = torch.utils.data.DataLoader(tensor_unmatched_control,
batch_size=1,
shuffle=False,
num_workers=1)
ps_unmatched_control_list = []
for batch in control_data_loader_train:
covariates_X, ps_score, y_f, y_cf = batch
ps_unmatched_control_list.append(ps_score.item())
return ps_unmatched_control_list
class InferenceNet:
def __init__(self, input_nodes, shared_nodes, outcome_nodes, device):
self.tarnet_phi = TARNetPhi(input_nodes, shared_nodes=shared_nodes).to(device)
self.tarnet_h_y1 = TARNetH_Y1(input_nodes=shared_nodes,
outcome_nodes=outcome_nodes).to(device)
self.tarnet_h_y0 = TARNetH_Y0(input_nodes=shared_nodes,
outcome_nodes=outcome_nodes).to(device)
def get_tarnet_phi(self):
return self.tarnet_phi
def get_tarnet_h_y1(self):
return self.tarnet_h_y1
def get_tarnet_h_y0_model(self):
return self.tarnet_h_y0
def train(self, train_parameters, device):
epochs = train_parameters["epochs"]
batch_size = train_parameters["batch_size"]
lr = train_parameters["lr"]
weight_decay = train_parameters["lambda"]
shuffle = train_parameters["shuffle"]
treated_tensor_dataset = train_parameters["treated_tensor_dataset"]
tuple_control = train_parameters["tuple_control_train"]
treated_data_loader_train = torch.utils.data.DataLoader(treated_tensor_dataset,
batch_size=batch_size,
shuffle=shuffle,
num_workers=1)
optimizer_W = optim.Adam(self.tarnet_phi.parameters(), lr=lr)
optimizer_V1 = optim.Adam(self.tarnet_h_y1.parameters(), lr=lr, weight_decay=weight_decay)
optimizer_V2 = optim.Adam(self.tarnet_h_y0.parameters(), lr=lr, weight_decay=weight_decay)
lossF = nn.MSELoss()
print(".. Training started ..")
print(device)
for epoch in range(epochs):
epoch += 1
total_loss_T = 0
total_loss_C = 0
for batch in treated_data_loader_train:
covariates_X_treated, ps_score_treated, y_f_treated, y_cf_treated = batch
covariates_X_treated = covariates_X_treated.to(device)
ps_score_treated = ps_score_treated.squeeze().to(device)
_tuple_treated = self.get_np_tuple_from_tensor(covariates_X_treated, ps_score_treated,
y_f_treated, y_cf_treated)
psm = PS_Matching()
tuple_matched_control = psm.match_using_prop_score(_tuple_treated, tuple_control)
covariates_X_control, ps_score_control, y_f_control, y_cf_control = \
self.get_tensor_from_np_tuple(tuple_matched_control)
y1_hat = self.tarnet_h_y1(self.tarnet_phi(covariates_X_treated))
y0_hat = self.tarnet_h_y0(self.tarnet_phi(covariates_X_treated))
if torch.cuda.is_available():
loss_T = lossF(y1_hat.float().cuda(),
y_f_treated.float().cuda()).to(device)
loss_C = lossF(y0_hat.float().cuda(),
y_f_control.float().cuda()).to(device)
else:
loss_T = lossF(y1_hat.float(),
y_f_treated.float()).to(device)
loss_C = lossF(y0_hat.float(),
y_f_control.float()).to(device)
optimizer_W.zero_grad()
optimizer_V1.zero_grad()
optimizer_V2.zero_grad()
loss_T.backward()
loss_C.backward()
optimizer_W.step()
optimizer_V1.step()
optimizer_V2.step()
total_loss_T += loss_T.item()
total_loss_C += loss_C.item()
if epoch % 100 == 0:
print("epoch: {0}, Treated + Control loss: {1}".format(epoch, total_loss_T + total_loss_C))
def eval(self, eval_parameters, device):
treated_set = eval_parameters["treated_set"]
control_set = eval_parameters["control_set"]
treated_data_loader = torch.utils.data.DataLoader(treated_set,
shuffle=False, num_workers=1)
control_data_loader = torch.utils.data.DataLoader(control_set,
shuffle=False, num_workers=1)
err_treated_list = []
err_control_list = []
true_ITE_list = []
predicted_ITE_list = []
ITE_dict_list = []
for batch in treated_data_loader:
covariates_X, ps_score, y_f, y_cf = batch
covariates_X = covariates_X.to(device)
y1_hat = self.tarnet_h_y1(self.tarnet_phi(covariates_X))
y0_hat = self.tarnet_h_y0(self.tarnet_phi(covariates_X))
predicted_ITE = y1_hat - y0_hat
true_ITE = y_f - y_cf
if torch.cuda.is_available():
diff = true_ITE.float().cuda() - predicted_ITE.float().cuda()
else:
diff = true_ITE.float() - predicted_ITE.float()
# ITE_dict_list.append(self.create_ITE_Dict(covariates_X,
# ps_score.item(), y_f.item(),
# y_cf.item(),
# true_ITE.item(),
# predicted_ITE.item(),
# diff.item()))
err_treated_list.append(diff.item())
true_ITE_list.append(true_ITE.item())
predicted_ITE_list.append(predicted_ITE.item())
for batch in control_data_loader:
covariates_X, ps_score, y_f, y_cf = batch
covariates_X = covariates_X.to(device)
y1_hat = self.tarnet_h_y1(self.tarnet_phi(covariates_X))
y0_hat = self.tarnet_h_y0(self.tarnet_phi(covariates_X))
predicted_ITE = y1_hat - y0_hat
true_ITE = y_cf - y_f
if torch.cuda.is_available():
diff = true_ITE.float().cuda() - predicted_ITE.float().cuda()
else:
diff = true_ITE.float() - predicted_ITE.float()
# ITE_dict_list.append(self.create_ITE_Dict(covariates_X,
# ps_score.item(), y_f.item(),
# y_cf.item(),
# true_ITE.item(),
# predicted_ITE.item(),
# diff.item()))
err_control_list.append(diff.item())
true_ITE_list.append(true_ITE.item())
predicted_ITE_list.append(predicted_ITE.item())
# print(err_treated_list)
# print(err_control_list)
return {
"treated_err": err_treated_list,
"control_err": err_control_list,
"true_ITE": true_ITE_list,
"predicted_ITE": predicted_ITE_list,
"ITE_dict_list": ITE_dict_list
}
@staticmethod
def get_np_tuple_from_tensor(covariates_X, ps_score, y_f, y_cf):
np_covariates_X = covariates_X.numpy()
ps_score = ps_score.numpy()
y_f = y_f.numpy()
y_cf = y_cf.numpy()
_tuple = (np_covariates_X, ps_score, y_f, y_cf)
return _tuple
@staticmethod
def get_tensor_from_np_tuple(_tuple):
np_df_X, np_ps_score, np_df_Y_f, np_df_Y_cf = _tuple
return torch.from_numpy(np_df_X), torch.from_numpy(np_ps_score), \
torch.from_numpy(np_df_Y_f), torch.from_numpy(np_df_Y_cf),
# +
tuple_control_train = (np_control_x, np_control_ps, np_control_f, np_control_cf)
tuple_treated_train = (np_treated_x, np_treated_ps, np_treated_f, np_treated_cf)
print("Actual Sizes")
print(np_treated_x.shape)
print(np_control_x.shape)
tensor_treated = Utils.convert_to_tensor_DCN(np_treated_x, np_treated_ps,
np_treated_f, np_treated_cf)
train_parameters = {
"epochs": 100,
"lr": 1e-3,
"lambda":1e-4,
"batch_size": 32,
"shuffle": True,
"treated_tensor_dataset": tensor_treated,
"tuple_control_train": tuple_control_train
}
inference = InferenceNet(input_nodes=25, shared_nodes=200,
outcome_nodes=100,
device=device)
inference.train(train_parameters, device)
# +
# testing
print("----------- Testing phase ------------")
dL = DataLoader()
ps_test_set = dL.convert_to_tensor(np_covariates_X_test,
np_covariates_Y_test)
is_synthetic = False
input_nodes = 25
# get propensity scores using NN
ps_net_NN = Propensity_socre_network()
ps_eval_parameters_NN = {
"eval_set": ps_test_set,
"model_path": Constants.PROP_SCORE_NN_MODEL_PATH
.format(iter_id, Constants.PROP_SCORE_NN_EPOCHS, Constants.PROP_SCORE_NN_LR),
"input_nodes": input_nodes
}
ps_score_list_NN = ps_net_NN.eval(ps_eval_parameters_NN, device, phase="eval")
data_loader_dict = dL.prepare_tensor_for_DCN(np_covariates_X_test,
np_covariates_Y_test,
ps_score_list_NN,
is_synthetic)
model_path = model_path.format(iter_id,
epochs,
lr)
print(model_path)
treated_group = data_loader_dict["treated_data"]
np_treated_df_X = treated_group[0]
np_treated_ps_score = treated_group[1]
np_treated_df_Y_f = treated_group[2]
np_treated_df_Y_cf = treated_group[3]
tensor_treated = Utils.convert_to_tensor_DCN(np_treated_df_X, np_treated_ps_score,
np_treated_df_Y_f, np_treated_df_Y_cf)
control_group = data_loader_dict["control_data"]
np_control_df_X = control_group[0]
np_control_ps_score = control_group[1]
np_control_df_Y_f = control_group[2]
np_control_df_Y_cf = control_group[3]
tensor_control = Utils.convert_to_tensor_DCN(np_control_df_X, np_control_ps_score,
np_control_df_Y_f, np_control_df_Y_cf)
print(np_treated_df_X.shape)
print(np_control_df_X.shape)
test_parameters = {
"treated_set": tensor_treated,
"control_set": tensor_control
}
response_dict = inference.eval(test_parameters, device)
err_treated = [ele ** 2 for ele in response_dict["treated_err"]]
err_control = [ele ** 2 for ele in response_dict["control_err"]]
total_sum = sum(err_treated) + sum(err_control)
total_item = len(err_treated) + len(err_control)
MSE = total_sum / total_item
print("Testing using Inference")
print(MSE)
# -
| RoughWork.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3]
# language: python
# name: conda-env-py3-py
# ---
# # Check River Flows
# +
import matplotlib.pyplot as plt
import netCDF4 as nc
import numpy as np
from salishsea_tools import nc_tools
# %matplotlib inline
# -
def find_points(flow):
for i in range(390, 501):
for j in range(280, 398):
if flow[0,i,j] != 0:
print (i,j, lat[i,j], lon[i,j], flow[0,i,j] * e1t[i,j] * e2t[i,j]/
(e1t[500, 395] * e2t[500, 395]))
def check_points(flow, tmask):
for i in range(898):
for j in range(398):
if tmask[0, i, j] !=1:
if flow[0, i, j] != 0:
print ('Land River', i ,j)
grid = nc.Dataset('/ocean/sallen/allen/research/MEOPAR/NEMO-forcing/grid/bathymetry_201702.nc')
lat = grid.variables['nav_lat'][:,:]
lon = grid.variables['nav_lon'][:,:]
depth = grid.variables['Bathymetry'][:]
grid.close()
mesh = nc.Dataset('../../../NEMO-forcing/grid/mesh_mask201702.nc')
e1t = mesh.variables['e1t'][0,:]
e2t = mesh.variables['e2t'][0,:]
tmask = mesh.variables['tmask'][0,:]
mesh.close()
river1 = nc.Dataset('/results/forcing/rivers/R201702DFraCElse_y2014m09d12.nc')
river2 = nc.Dataset('/ocean/sallen/allen/research/MEOPAR/NEMO-forcing/rivers/rivers_month_201702.nc')
print ('Sep 12, 2014')
print ('Daily')
find_points(river1.variables['rorunoff'][:,:,:])
print ('Monthly')
find_points(river2.variables['rorunoff'][:,:,:])
print ('Difference')
find_points(river2.variables['rorunoff'][:,:,:] - river1.variables['rorunoff'][:,:,:])
check_points(river2.variables['rorunoff'][:,:,:], tmask)
check_points(river2.variables['rorunoff'][:,:,:], tmask)
fig, ax = plt.subplots(1,1,figsize=(15,7.5))
imin = 780; imax = 830; jmin = 100; jmax = 150
cmap = plt.get_cmap('winter_r')
cmap.set_bad('burlywood')
mesh = ax.pcolormesh(depth[imin:imax,jmin:jmax], cmap=cmap)
ax.set_xlabel('Grid Points')
ax.set_ylabel('Grid Points')
cbar=fig.colorbar(mesh)
cbar.set_label('Depth (m)')
ax.plot(806+0.5-imin, 126+0.5-jmin,'ko');
| notebooks/Check River Files.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SPARQL
# language: sparql
# name: sparql
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # KEN3140: Lab 5
# -
# ### Writing and executing advanced SPARQL queries on remote RDF graphs (SPARQL endpoints)
#
# ##### Authors:
# + [<NAME>](https://www.maastrichtuniversity.nl/vincent.emonet): [<EMAIL>](mailto:<EMAIL>)
# + [<NAME>](https://www.maastrichtuniversity.nl/kody.moodley): [<EMAIL>](mailto:<EMAIL>)
#
# ##### Affiliation:
# [Institute of Data Science](https://www.maastrichtuniversity.nl/research/institute-data-science)
#
# ##### License:
# [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0)
#
# ##### Date:
# 2021-09-06
# #### In this lab you will learn:
#
# How to compose more advanced [SPARQL](https://www.w3.org/TR/2013/REC-sparql11-query-20130321/) queries to answer questions about the content of an RDF knowledge graph
#
# #### Specific learning goals:
#
# + How to select the appropriate SPARQL query form to answer a given question about a knowledge graph
# + How to select the appropriate SPARQL functions required to perform any calculations in a SPARQL query
# + How to refer to entities located in external SPARQL endpoints from a SPARQL query (querying multiple distributed endpoints)
# #### Prerequisite knowledge:
# + Lecture 4: Introduction to SPARQL
# + Lab 4: Introduction to SPARQL
# + Lecture 5: Advanced SPARQL
# + [SPARQL 1.1 language specification](https://www.w3.org/TR/sparql11-query/)
# + Chapters 1 - 3 of [Learning SPARQL](https://maastrichtuniversity.on.worldcat.org/external-search?queryString=SPARQL#/oclc/853679890)
# + Chapters 4 - 9, 11 of [Learning SPARQL](https://maastrichtuniversity.on.worldcat.org/external-search?queryString=SPARQL#/oclc/853679890)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Task information:
#
# # + In this lab, we will ask you to query the [DBpedia](https://dbpedia.org/) knowledge graph!
# # + [DBpedia](https://dbpedia.org/) is a crowd-sourced community effort to extract structured content in RDF from the information created in various [Wikimedia](https://www.wikimedia.org/) projects (e.g. [Wikipedia](https://www.wikipedia.org/)). DBpedia is similar in information content to [Wikidata](https://www.wikidata.org/wiki/Wikidata:Main_Page).
# # + **A word on data quality:** remember that DBpedia is crowd-sourced. This means that volunteers and members of the general public are permitted to add and maintain it's content. As a result, you may encounter inaccuracies / omissions in the content and inconsistencies in how the information is represented. Don't be alarmed by this, it is not critical that the content is accurate for the learning objectives of this lab.
#
# -
# #### Task information (contd):
#
# + The DBpedia SPARQL endpoint URL is: [https://dbpedia.org/sparql](https://dbpedia.org/sparql)
# + DBPedia has it's own SPARQL query interface at [https://dbpedia.org/sparql](https://dbpedia.org/sparql) which is built on OpenLink's [Virtuoso](https://virtuoso.openlinksw.com/) [RDF](https://www.w3.org/TR/rdf11-concepts/) triplestore management system.
# + In this lab, we will use an alternative SPARQL query interface to query DBPedia. It is called **[YASGUI](https://yasgui.triply.cc)**. The reason is that YASGUI has additional user-friendly features e.g. management of multiple SPARQL queries in separate tabs. It also allows one to query any publicly available SPARQL endpoint from the same interface.
# + [markdown] slideshow={"slide_type": "slide"}
# #### Tips 🔎
#
# # + How do I find vocabulary to use in my SPARQL query from DBpedia?
#
# > Search on google, e.g., if you want to know the term for "capital city" in DBpedia, search for: "**[dbpedia capital](https://www.google.com/search?&q=dbpedia+capital)**" In general, "dbpedia [approximate name of predicate or class you are looking for]"
#
# > Your search query does not have to exactly match the spelling of the DBpedia resource name you are looking for
#
# > Alternatively, you can formulate SPARQL queries to list properties and types in DBpedia Do you know what these queries might look like?
#
# # + Use [prefix.cc](http://prefix.cc/) to discover the full IRIs for unknown prefixes you may encounter
# # + First read the question carefully and determine what **query form** the question requires
# # + Do not try to formulate the full SPARQL query for each task from the get go. Rather build and test smaller queries first and add to it incrementally, testing each time to see if the results are expected
# # + Make use of the Lecture 4 & 5 slides and SPARQL specification page if you need to find additional functions and query features that may assist you in doing the tasks
#
# -
# # YASGUI interface
#
# <img src="yasgui-interface.png">
# + [markdown] slideshow={"slide_type": "subslide"}
# <!-- # Install the SPARQL kernel
#
# This notebook uses the SPARQL Kernel to define and **execute SPARQL queries in the notebook** codeblocks.
# To **install the SPARQL Kernel** in your JupyterLab installation:
#
# ```shell
# pip install sparqlkernel --user
# jupyter sparqlkernel install --user
# ```
#
# To start running SPARQL query in this notebook, we need to define the **SPARQL kernel parameters**:
# * 🔗 **URL of the SPARQL endpoint to query**
# * 🌐 Language of preferred labels
# * 📜 Log level -->
# + slideshow={"slide_type": "fragment"}
%endpoint http://dbpedia.org/sparql
# This is optional, it would increase the log level
%log debug
# Uncomment the next line to return label in english and avoid duplicates
# %lang en
# + [markdown] slideshow={"slide_type": "slide"}
# # Anatomy of a SPARQL query
#
# As we saw in Lecture 4, these are the main components of a SPARQL query:
#
# <img src="sparql_query_breakdown.png">
# + [markdown] slideshow={"slide_type": "slide"}
# # Task 1 [10min]:
# -
# Calculate and display the average [Gross Domestic Product (GDP)](https://en.wikipedia.org/wiki/Gross_domestic_product) of each country in DBpedia and aggregate the results by the currency that each country uses. Display the GDP averages from highest at the top of the list, to lowest at the bottom of the list
# + [markdown] slideshow={"slide_type": "slide"}
# # Task 2 [20min]:
# + [markdown] slideshow={"slide_type": "slide"}
# a) List the first 10 countries to have been dissolved (ceased to exist) and display the resulting countries in order from the most recently created, to the least recently created. **Hint:** use the properties **[dbo:dissolutionDate](https://dbpedia.org/ontology/dissolutionDate)** and **[dbo:foundingYear](https://dbpedia.org/ontology/foundingYear)** in your answer.
# -
# b) Display a set of triples indicating the country of origin of book authors in DBpedia. In the displayed triples:
# The type of book author instances should be indicated by **[schema:Person](https://schema.org/Person)** from **[Schema.org](http://schema.org)**
# The country of origin of book authors should be represented by **[schema:countryOfOrigin](https://schema.org/countryOfOrigin)**.
# **Note:** you can observe that in this CONSTRUCT query, we have basically created new triples using **new vocabulary** about the place of birth (country of origin) for book authors. This is sometimes called "mapping", "transforming" or "capturing" data in "another model". Here "model" refers to a new vocabulary. The old vocabulary used was DBpedia, and the new one is Schema.org.
#
# Suppose that I already had a knowledge graph using Schema.org vocabulary. If I wanted to integrate external information from another knowledge graph (e.g. DBpedia), this kind of transformation is helpful for importing those triples into the same "model" (vocabulary) of my knowledge graph. This allows me to query DBpedia information using my own vocabulary (Schema.org) without needing to use DBpedia terms in my queries.
#
# Of course, CONSTRUCT will not insert the triples to my graph. If I wanted to do that, I would need INSERT, and I would also need a federated query to access the SPARQL endpoint of my own knowledge graph from the DBpedia knowledge graph.
#
# The utility of CONSTRUCT here is to show us what these triples will look like and I can verify the correctness of these triples before I import them, if that is what I wanted to do.
# # Task 3 [20min]:
#
# **Background:** This task will ask you to create a federated SPARQL query to query information from two SPARQL distributed endpoints. In preparation for this task, change the SPARQL endpoint in YASGUI or this notebook (whichever you are using) to: https://bio2rdf.org/sparql/.
%endpoint http://bio2rdf.org/sparql
# **Note:** you may experience technical difficulties with this task such as long query execution times, errors in the execution of the federated query, or no results. This could be due to security access privileges changed in the external endpoint, the sheer size of the knowledge graphs, or changes to the graph vocabulary / content.
#
# **Facing major problems with this task?** If you do experience any of the above issues and would still like to see some results after executing a federated query, you can execute the SPARQL query provided [here](https://github.com/MaastrichtU-IDS/UM_KEN4256_KnowledgeGraphs/blob/master/resources/federated-query-genes-pathways.rq) (copy and paste it) at the following SPARQL endpoint: [http://sparql.wikipathways.org/sparql](http://sparql.wikipathways.org/sparql)
# **Task instructions:**
# List drugs that interact with 10 proteins and the label of the Genes that encode them 🧬
#
# + **Graph 1:** Get “gene encodes protein” information from the UniProt SPARQL endpoint (https://sparql.uniprot.org/)
# + **Graph 2:** Get “drug interacts with protein” from the IDS GraphDB installation (http://graphdb.dumontierlab.com/repositories/ncats-red-kg)
#
# Browse the biomedical vocabulary/ontology used in these graphs here: https://biolink.github.io/biolink-model/docs/classes.html
#
# ⚠️ Federated queries are much slower, use a subquery in the service call to avoid multiple calls between services
# + [markdown] slideshow={"slide_type": "slide"}
# # Examples of other public SPARQL endpoints 🔗
#
# * Wikidata, facts powering Wikipedia infobox: https://query.wikidata.org/sparql
# * Bio2RDF, linked data for the life sciences: https://bio2rdf.org/sparql
# * Disgenet, gene-disease association: http://rdf.disgenet.org/sparql
# * PathwayCommons, resource for biological pathways analysis: http://rdf.pathwaycommons.org/sparql
# * EU publications office, court decisions and legislative documents from the EU: http://publications.europa.eu/webapi/rdf/sparql
# * Finland legal open data, cases and legislation: https://data.finlex.fi/en/sparql
# * EU Knowledge Graph, open knowledge graph containing general information about the European Union: [SPARQL endpoint](https://query.linkedopendata.eu/#SELECT%20DISTINCT%20%3Fo1%20WHERE%20%7B%0A%20%20%3Chttps%3A%2F%2Flinkedopendata.eu%2Fentity%2FQ1%3E%20%3Chttps%3A%2F%2Flinkedopendata.eu%2Fprop%2Fdirect%2FP62%3E%20%3Fo1%20.%20%0A%7D%20%0ALIMIT%201000)
#
# # SPARQL applied to the COVID pandemic:
#
# * Wikidata SPARQL queries around the SARS-CoV-2 virus and pandemic: https://egonw.github.io/SARS-CoV-2-Queries
| lab05-sparql-advanced/Lab5_SPARQL_advanced.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#import urllib2
from io import StringIO
import os
import xmltodict
import pandas as pd
import geopandas as gpd
import windrose
from datetime import datetime
import statsmodels.api as sm
from scipy.stats import linregress
import matplotlib.pyplot as plt
import numpy as np
import sys
import requests
import glob
from pyproj import CRS
from pyproj import Transformer
from shapely.geometry import Point
import rasterstats as rs
import rasterio
from rasterio.plot import show
from rasterio.mask import mask
from rasterio.plot import show_hist
from shapely.geometry import box
from rasterstats import zonal_stats
from windrose import WindroseAxes
#import pycrs
#import earthpy as et
#import earthpy.plot as ep
from scipy.optimize import curve_fit
from sklearn import mixture
import itertools
from scipy import linalg
import matplotlib as mpl
#import pymannkendall as mk
from pylab import rcParams
# %matplotlib inline
rcParams['figure.figsize'] = 15, 10
# +
import matplotlib.pyplot as plt
from pylab import rcParams
# %matplotlib inline
rcParams['figure.figsize'] = 15, 10
from windrose import WindroseAxes
import geopandas as gpd
import windrose
import pandas as pd
import glob
import os
# -
xlfile = r"G:\Shared drives\UGS_Groundwater\Projects\Eddy_Covariance\Data_Downloads\BSF\Bonneville Salt Flats_Flux_AmeriFluxFormat.xlsx"
amflux = pd.read_excel(xlfile,skiprows=[1],index_col=0,parse_dates=True,na_values="NAN")
amflux[(amflux['CO2']>366)&(amflux['CO2']<405)]['CO2'].plot()
df = amflux.copy(deep=True)
df
roll_avg+2.5*roll_std
df['roll_med']
# +
window = 48
devs= 2.0
def filtdf(df, field, window=48, devs=2.0):
"""
df = dataframe with data
field = field you want to filter
window = rolling filter to apply
devs = std deviations to filter
returns filtered df
"""
df['roll_top'] = df[field].rolling(window).median().interpolate(method='bfill') + df[field].rolling(window).std().interpolate(method='bfill')*devs
df['roll_bot'] = df[field].rolling(window).median().interpolate(method='bfill') - df[field].rolling(window).std().interpolate(method='bfill')*devs
def noout(x):
if (x[0] <= x[1]) & (x[0] >= x[2]):
return x[0]
else:
return None
df[f"{field}_filt"] = df[[field,'roll_top','roll_bot']].apply(lambda x: noout(x),1)
df = df.drop(['roll_top','roll_bot'],axis=1)
return df
df = filtdf(df, 'CO2')
df = filtdf(df, 'ET')
#df['CO2_filt'].plot()
df['ET_filt'] = df['ET_filt'].mask(df['ET_filt'].lt(0))
#(df['ET_filt']*24).plot()
openetfile = r"G:\Shared drives\UGS_Groundwater\Projects\Eddy_Covariance\Data_Downloads\BSF\explore.etdata.org-shape.csv"
openet = pd.read_csv(openetfile, index_col='DateTime',parse_dates=True)
fig, ax = plt.subplots(2,1, sharex=True)
df.resample('1D').sum()['ET_filt'].plot(ax=ax[0])
ax[0].set_ylabel('ET (mm/day)')
df['CO2_filt'].groupby(pd.Grouper(freq='1D')).median().plot(ax=ax[1])
ax[1].set_ylabel('CO2 (ppm)')
# -
df['SWC_1_1_1'].plot()
openet['month'] = openet.index.month
openet.groupby('month')['PT-JPL ET'].median().plot(marker='o')
df['CO2_filt'].plot()
| BSF_AMERIFLUX_output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
from collections import Counter
# import spacy
# nlp = spacy.load('en_core_web_sm')
# from spacy.lang.en import English
# tokenizer = English().Defaults.create_tokenizer(nlp)
# def my_spacy_tokenizer(doc):
# # TODO: need to add a phase to break -- offsets and strip white space.
# tokens = tokenizer(doc)
# return([token.text for token in tokens])
# -
# # Dataset statistics on distribution of idf & vocab relative to number examples
#
# Datasets have fairly similar vocab size to number of example utterances ratios, but crisischats' is roughly double, indicating a large vocab relative to dataset size.
tasks = ['crisischatsmessages', 'cornell_movie', 'dailydialog', 'empathetic_dialogues', 'personachat']
for task in tasks:
tot_doc = open('/data/odemasi/packages/ParlAI/tmp/%s/dict_minfreq_2.tot_doc' % task, 'r').readline()
tot_doc = float(tot_doc.strip('\n'))
lines = open('/data/odemasi/packages/ParlAI/tmp/%s/dict_minfreq_2.doc_freq' % task, 'r').readlines()
doc_freq = [float(x.split('\t')[1].strip('\n')) for x in lines[4:]] # strip off special tokens at front
plt.hist([np.log(tot_doc/x) for x in doc_freq], bins=50)
plt.title('%s vocab: %s tot_doc: %s vocab/utters: %.2f' % (task, len(doc_freq), tot_doc, len(doc_freq)/float(tot_doc)))
# plt.xlim([-1, 11])
plt.xlabel('idf across utterances', fontsize=20)
plt.ylabel('Number of words', fontsize=20)
plt.show()
# # Load crowdworker input data
# +
batch_name = '../batches/Batch_3622867_batch_results.csv'
raw_df = pd.read_csv(batch_name)
turk_df = raw_df
# -
raw_df[:3]
list(raw_df.columns)
raw_df.loc[0,'Input.model_3_0']
# ## Look at evaluation of the warmup question:
for colname in ['Answer.int_warmup', 'Answer.rel_warmup',
'Answer.check_int_warmup', 'Answer.check_rel_warmup',
'Answer.rank_warmup_1', 'Answer.rank_warmup_2', 'Answer.rank_warmup_3']:
print(colname,'\t', Counter(raw_df[colname]))
# ## Justifications of warmup
print( '\n'.join(raw_df['Answer.Justified Answer'].values))
# +
# print( '\n'.join(map(str, raw_df['Answer.rank_warmup_3'].values)))
# -
# ## Justification of final
print(raw_df['Answer.Justified Answer Final'].values)
# ## Optional feedback:
print( raw_df['Answer.optionalfeedback'].values)
# 'A lot of these are non-sensical responses. It makes it hard to "rank" them'
# 'please define ~~ for future workers.'
# 'I see you got rid of the "ungrammatical" checkbox. Yeah, that wasn\'t working out too well before. Good decision.'
# ## check warmup question
# ## Look at justifications
# +
# [type(x) == str for x in raw_df['Answer.check_int_9']]
# -
raw_df[['Answer.check_int_9',
'Answer.check_int_warmup',
'Answer.check_rel_0',
'Answer.int_0',
'Answer.rank_0_1', # example_option
'Answer.rel_0']].loc[0]
# # Parse data into convenient data frame
# +
model_order = ['s2s', 'transformer', 'lm']
method_order = ['vanilla', 'idf', 'swapping']
choice_df = []
# grammar_df = []
for i in turk_df.index:
row = turk_df.loc[i]
hit_num = i
worker_id = row['WorkerId']
for ex in range(10): # given to each turker in a single HIT
option_to_method = {}
method_to_rank = {}
method_to_resp = {}
# method_to_gram = {}
for option in range(1, 4):
model_pieces = row['Input.model_%s_%s' % (option, ex)].split(' ')
if len(model_pieces) == 2:
dataset, second = model_pieces
if second not in ['s2s', 'transformer']:
method = second
model = 's2s'
else:
method = 'vanilla'
model = second
else:
dataset, model, method = model_pieces
option_to_method[option] = method
method_to_rank[method] = row['Answer.rank_%s_%s' % (ex, option)]
method_to_resp[method] = row['Input.resp_%s_%s' % (option, ex)]
# method_to_gram[method] = np.isnan(row['Answer.gram_%s_%s' % (ex, option)])
most_interesting = option_to_method[row['Answer.int_%s' % (ex,)]]
most_relevant = option_to_method[row['Answer.rel_%s' % (ex,)]]
none_int = type(row['Answer.check_int_%s'% (ex,)]) == str
none_rel = type(row['Answer.check_rel_%s'% (ex,)]) == str
message1 = row['Input.msg1_%s' % (ex,)]
message2 = row['Input.msg2_%s' % (ex,)]
choice_df.append([hit_num, worker_id, dataset, model, most_interesting, most_relevant, \
message1, message2, none_int, none_rel] + \
[method_to_rank[m] for m in method_order] + \
[method_to_resp[m] for m in method_order]
)
choice_df = pd.DataFrame(choice_df, columns = ['hit_num', 'worker_id', 'dataset', 'model', \
'most_int', 'most_rel', \
'message1', 'message2', \
'none_int', 'none_rel']\
+ method_order \
+ ['resp_%s' % m for m in method_order])
choice_df['context'] = choice_df['message1'] + choice_df['message2']
# -
choice_df[:3]
# # Summary statistics on all HIT input
# +
print('Most interesting: ', Counter(choice_df['most_int']))
print('Most relevant: ', Counter(choice_df['most_rel']))
print('Percent of examples where int==rel: %.2f%%' % (100.0*np.sum(choice_df['most_int'] == choice_df['most_rel']) \
/ float(choice_df['most_rel'].shape[0]),))
print('Fraction of examples where none interesting: %.2f' % choice_df['none_int'].mean())
print('Fraction of examples where none relevant: %.2f' % choice_df['none_rel'].mean())
# -
# # Consider times model gets a rank (consider all input individually)
# If a method gets a majority of rank 1 votes (for best message overall), it's usually vanilla. Idf and swapping split places 2 and 3
# +
colors = ['purple', 'grey', 'plum']
for (dataset, model), setting_df in choice_df.groupby(['dataset', 'model']):
some_relevant = ~setting_df['none_rel']
fig, ax = plt.subplots(1,2, figsize=(10,4))
for name, df in [('all obs', setting_df),
('w/o "none relevant"', setting_df[some_relevant])]:
N = float(df.shape[0])
if name == 'all obs':
use_ax = ax[0]
else:
use_ax = ax[1]
method_max = 0
for m, method in enumerate(method_order):
for r in range(3):
perc_rank = 100.*np.sum(df[method] == (r+1))/N
if r == 0:
method_max = np.max((method_max, perc_rank))
use_ax.bar(m*4+r, perc_rank, color=colors[m], label=method)
else:
use_ax.bar(m*4+r, perc_rank, color=colors[m])
use_ax.set_xticks(np.arange(12))
use_ax.set_xticklabels(1+np.mod(np.arange(12),4))
use_ax.plot([-1,12], [method_max, method_max],'k', linewidth=3, label='Rank 1 max: %d%%'%method_max)
use_ax.legend()
use_ax.set_ylim([0, 60])
use_ax.set_title(name, fontsize=14)
use_ax.set_ylabel('Percent of input', fontsize=14)
use_ax.set_xlabel('Rank in order of "best response"', fontsize=14)
plt.suptitle('%s %s' % (dataset, model), fontsize=14)
plt.show()
# -
# # Considering only examples that get a majority of "best" rank
#
# If we only consider examples where an method is ranked # 1 best overall, we see vanilla win and unclear difference between idf and swapping.
#
# +
colors = ['purple', 'grey', 'plum']
for (dataset, model), setting_df in choice_df.groupby(['dataset', 'model']):
best_choice = []
for context, df in setting_df.groupby('context'):
rank_votes = (df[['vanilla', 'idf', 'swapping']] == 1).sum()
if rank_votes.max() >= 2:
best_choice.append(rank_votes[rank_votes == rank_votes.max()].index[0])
c = Counter(best_choice)
for i in range(len(method_order)):
plt.bar(i, c[method_order[i]], label=method_order[i], color=colors[i])
plt.xticks(np.arange(3))
plt.legend(loc='best')
plt.xlabel('number chosen for best overall', fontsize=20)
plt.ylabel('number ex. (with majority)', fontsize=20)
plt.title('%s %s' % (dataset, model))
plt.show()
# -
# # Worker agreement on interestingness & relevance
# There is considerable disagreement amoung workers, but it varies between datasets.
for (dataset, model), setting_df in choice_df.groupby(['dataset', 'model']):
fig, ax = plt.subplots(1,2, figsize=(10,4))
num_most_int = []
num_most_rel = []
for context, df in setting_df.groupby('context'):
num_most_int.append(len(df['most_int'].unique()))
num_most_rel.append(len(df['most_rel'].unique()))
ax[0].hist(num_most_int, bins=np.arange(5)-.5)
ax[0].set_xticks(np.arange(3))
ax[0].set_xlabel('number chosen for most int', fontsize=20)
ax[0].set_ylabel('number examples', fontsize=20)
# ax[0].set_title('%s %s' % (dataset, model), fontsize=20)
# plt.show()
ax[1].hist(num_most_rel, bins=np.arange(5)-.5)
ax[1].set_xlabel('number chosen for most rel', fontsize=20)
ax[1].set_ylabel('number examples', fontsize=20)
# ax[1].set_title('%s %s' % (dataset, model), fontsize=20)
ax[1].set_xticks(np.arange(3))
plt.suptitle('%s %s' % (dataset, model), fontsize=20)
plt.show()
# # Look at how often a method is chosen for most relevant/interesting.
# +
for (dataset, model), setting_df in choice_df.groupby(['dataset', 'model']):
some_int = (~setting_df['none_int'])
some_rel = (~setting_df['none_rel'])
int_df = setting_df[some_int]
rel_df = setting_df[some_rel]
print(dataset, model)
print('=========')
data_model_key = '%s_%s' % (dataset, model)
fig, ax = plt.subplots(1, 2)
for m, method in enumerate(method_order):
ax[0].bar(m, np.sum(int_df['most_int'] == method))
ax[1].bar(m, np.sum(rel_df['most_rel'] == method))
ax[0].set_xticks(np.arange(3))
ax[0].set_xticklabels(method_order)
ax[0].set_title('Most interesting', fontsize=14)
ax[0].set_ylabel('Number input', fontsize=20)
# ax[0].set_title('%s %s' % (dataset, model), fontsize=20)
# plt.show()
ax[1].set_title('Most relevant', fontsize=14)
# ax[1].set_ylabel('Number examples', fontsize=20)
# ax[1].set_title('%s %s' % (dataset, model), fontsize=20)
ax[1].set_xticks(np.arange(3))
ax[1].set_xticklabels(method_order)
# cross_mat = np.NaN * np.zeros((len(method_order), len(method_order)))
# for i, m1 in enumerate(method_order):
# for j, m2 in enumerate(method_order):
# cross_mat[i,j] = np.sum((df['most_int'] == m1) & (df['most_rel'] == m2))
# plt.pcolor(cross_mat, vmin=0, vmax=50)
# plt.xticks(.5 + np.arange(len(method_order)), method_order)
# plt.yticks(.5 + np.arange(len(method_order)), method_order)
# plt.xlabel('Selected as most relevant')
# plt.ylabel('Selected as most interesting')
# plt.colorbar(label='Number of examples')
# ind = np.arange(len(method_order))
plt.suptitle('All input: %s' % (data_model_key, ), fontsize=14)
plt.show()
# -
# ## Exclude example without worker majority.
for (dataset, model), setting_df in choice_df.groupby(['dataset', 'model']):
some_int = (~setting_df['none_int'])
some_rel = (~setting_df['none_rel'])
int_df = setting_df[some_int]
rel_df = setting_df[some_rel]
rel_counts = np.zeros(len(method_order))
int_counts = np.zeros(len(method_order))
for context, ex_df in rel_df.groupby('context'):
c = Counter(ex_df['most_rel'])
if c.most_common(1)[0][1] > 1:
rel_counts[method_order.index(c.most_common(1)[0][0])] += 1
for context, ex_df in int_df.groupby('context'):
c = Counter(ex_df['most_int'])
if c.most_common(1)[0][1] > 1:
int_counts[method_order.index(c.most_common(1)[0][0])] += 1
print(dataset, model)
print('=========')
data_model_key = '%s_%s' % (dataset, model)
fig, ax = plt.subplots(1, 2)
for m, method in enumerate(method_order):
ax[0].bar(m, int_counts[m])
ax[1].bar(m, rel_counts[m])
ax[0].set_xticks(np.arange(3))
ax[0].set_xticklabels(method_order)
ax[0].set_title('Most interesting', fontsize=14)
ax[0].set_ylabel('Number examples', fontsize=20)
ax[1].set_title('Most relevant', fontsize=14)
ax[1].set_xticks(np.arange(3))
ax[1].set_xticklabels(method_order)
plt.suptitle('Majority only: %s' % (data_model_key,), fontsize=14)
plt.show()
# # Relationship of best and most relevant
# +
print('Percent inputs where most relevant response also chosen as best')
print('(not grouped by example, but "none relevant" removed)')
for (dataset, model), setting_df in choice_df.groupby(['dataset', 'model']):
some_rel = (~setting_df['none_rel'])
df = setting_df[some_rel]
print('=========')
print(dataset, model)
data_model_key = '%s_%s' % (dataset, model)
cor_mat = np.zeros((1, len(method_order)))
for ind in df.index:
most_rel_method = df.loc[ind, 'most_rel']
j = method_order.index(most_rel_method)
cor_mat[0, j] += df.loc[ind, most_rel_method] == 1 # ranked best overall.
print('\t\t %.2f' %(100.0*np.sum(cor_mat)/df.shape[0],))
# plt.pcolor(cor_mat/df.shape[0], vmin=0, vmax=50)
# plt.xticks(.5 + np.arange(len(method_order)), method_order)
# # plt.yticks(.5 + np.arange(len(method_order)), method_order)
# plt.xlabel('Selected as most relevant')
# # plt.ylabel('Selected as most interesting')
# plt.colorbar(label='Percent most relevant and mos i')
# ind = np.arange(len(method_order))
# plt.title('%s: Most int==most rel: %d%%' % (data_model_key, 100.0*np.sum(cross_mat[ind,ind])/np.sum(cross_mat)))
# plt.show()
# -
# # Relationship of when interested and when relevant
# There is little consistent relationship across datasets/models, but all have the most interesting match the most relevant < 25% of the time.
for (dataset, model), setting_df in choice_df.groupby(['dataset', 'model']):
some_int_relevant = ~(setting_df['none_rel'] & setting_df['none_int'])
df = setting_df[some_int_relevant]
print(dataset, model)
print('=========')
data_model_key = '%s_%s' % (dataset, model)
cross_mat = np.NaN * np.zeros((len(method_order), len(method_order)))
for i, m1 in enumerate(method_order):
for j, m2 in enumerate(method_order):
cross_mat[i,j] = np.sum((df['most_int'] == m1) & (df['most_rel'] == m2))
plt.pcolor(cross_mat, vmin=0, vmax=50)
plt.xticks(.5 + np.arange(len(method_order)), method_order)
plt.yticks(.5 + np.arange(len(method_order)), method_order)
plt.xlabel('Selected as most relevant')
plt.ylabel('Selected as most interesting')
plt.colorbar(label='Number of input')
ind = np.arange(len(method_order))
plt.title('%s: Most int==most rel: %d%%' % (data_model_key, 100.0*np.sum(cross_mat[ind,ind])/np.sum(cross_mat)))
plt.show()
method_order
# choice_df.columns
# task, method, model
choice_df['model'].unique()
# ## Look at distributions of idf generated
# We could run tests for distribution shifts, but everything is pretty overlapping, so I'd expect p-values to be very large.
# +
tasks = ['cornell_movie', 'dailydialog', 'empathetic_dialogues', 'personachat']
### Build dictionaries of idf for each dataset.
idf_dict = {t:{} for t in tasks}
for task in tasks:
tot_doc = open('/data/odemasi/packages/ParlAI/tmp/%s/dict_minfreq_2.tot_doc' % task, 'r').readline()
tot_doc = float(tot_doc.strip('\n'))
lines = open('/data/odemasi/packages/ParlAI/tmp/%s/dict_minfreq_2.doc_freq' % task, 'r').readlines()
# build idf dictionary for dataset:
for line in lines[4:]:
tok, ct = line.split('\t')
idf_dict[task][tok] = np.log(tot_doc/float(ct.strip('\n')))
### Look at distribution of generated idf for each model/dataset.
bin_edges = np.arange(0,13,.2)
generated_idf_dist = {t:{m:{} for m in ['seq2seq', 'transformer']} for t in tasks}
for (task, model), df in choice_df.groupby(['dataset', 'model']):
generated_idf_dist[task][model] = np.zeros((len(method_order), len(bin_edges)-1))
missing_toks = []
for m, method in enumerate(method_order):
responses = df['resp_%s'%method].values
idfs_generated = []
for resp in responses:
for tok in resp.split(' '):
try:
idfs_generated += [idf_dict[task][tok],]
except KeyError:
missing_toks += [tok,]
generated_idf_dist[task][model][m,:], _ = np.histogram(idfs_generated, bins=bin_edges)
print('Missing tokens: ', missing_toks)
# -
for task in tasks:
for model in choice_df['model'].unique():
fig, axs = plt.subplots(1,2, figsize=(10,5))
for m, method in enumerate(method_order):
axs[0].plot(bin_edges[:-1], generated_idf_dist[task][model][m,:], '-o', label=method)
axs[0].set_title('%s %s' % (task, model), fontsize=20)
axs[0].set_ylabel('number tokens generated', fontsize=20)
axs[0].legend(fontsize=18)
axs[0].set_xlabel('idf values', fontsize=20)
s = method_order.index('swapping')
v = method_order.index('vanilla')
axs[1].plot(bin_edges[:-1], 0*bin_edges[:-1], 'ko-')
axs[1].plot(bin_edges[:-1], generated_idf_dist[task][model][s,:] - generated_idf_dist[task][model][v,:], 'ro-')
plt.show()
# # Load relevance data
# +
elim_batch_name = '../batches/Batch_3629231_batch_results.csv'
elim_batch_name2 = '../batches/Batch_3636896_batch_results.csv'
# raw_elim_df = pd.read_csv(elim_batch_name)
raw_elim_df = pd.concat([pd.read_csv(elim_batch_name),
pd.read_csv(elim_batch_name2)], axis=0, sort=False).reset_index()
# -
raw_elim_df.shape
# +
# raw_elim_df['Input.model_1_3']
# +
elim_df = []
for i in raw_elim_df.index:
row = raw_elim_df.loc[i]
hit_num = i
worker_id = row['WorkerId']
for ex in range(10): # given to each turker in a single HIT
model_pieces = row['Input.model_1_%s' % (ex,)].split(' ')
dataset, model, method = model_pieces
well_formed_score = row['Answer.well_formed_%s' % (ex,)]
not_rel = type(row['Answer.check_rel_0_%s' % (ex,)]) == str
message1 = row['Input.msg1_%s' % (ex,)]
message2 = row['Input.msg2_%s' % (ex,)]
response = row['Input.resp_1_%s' % (ex,)]
elim_df.append([hit_num, worker_id, dataset, model, method, \
message1, message2, response, not_rel, well_formed_score])
elim_df = pd.DataFrame(elim_df, columns = ['hit_num', 'worker_id', 'dataset', 'model', 'method', \
'message1', 'message2', 'response', 'not_rel', 'well_score'])
elim_df['context'] = elim_df['message1'] + elim_df['message2']
# remove examples that had an empty message 2:
elim_df = elim_df[[type(x) == str for x in elim_df['message2']]]
# -
# ## Variability on warmup question:
for warmup_resp, df in raw_elim_df.groupby('Input.warmup_response'):
mean_score = df['Answer.well_formed_warmup'].mean()
print('Warmup response: ', warmup_resp)
print('Fraction flagged not-relevant: %.2f' % np.mean([type(x) == str for x in df['Answer.check_rel_1_warmup'].values]))
print('Mean well-formed score: ', mean_score)
plt.hist(df['Answer.well_formed_warmup'].values, bins = .5+np.arange(6))
plt.plot([mean_score, mean_score], [0, 30], linewidth=3)
plt.show()
raw_elim_df['Answer.optionalfeedback'].values
elim_df[:3]
# +
for (task, model), df in elim_df.groupby(['dataset', 'model']):
print(task, model)
for method, method_df in df.groupby('method'):
# perc_not_rel = 100.*method_df['not_rel'].mean()
perc_not_rel = 100.0 * np.mean(method_df.groupby('context')['not_rel'].agg(np.sum)>1)
well_score = method_df['well_score'].mean()
below_3 = np.mean(method_df.groupby('context')['well_score'].agg(np.mean) < 3)
well_std = method_df.groupby('context')['well_score'].agg(np.mean).std()
print('\t percent not relevant: %.2f \tmean well score (std): %.3f (%.3f) perc<3: %.2f \t %s' % \
(perc_not_rel, well_score, well_std, 100.0*below_3, method))
# +
method_order = ['vanilla', 'swapping', 'idf', 'face']
method_name = {'vanilla':'unweighted',
'swapping':'idf+swap',
'idf':'idf',
'face': 'FACE'}
dataset_name = {'cornell_movie':'Cornell Movie',
'dailydialog':'DailyDialog',
'empathetic': 'Empathetic Dialogues',
'empathetic_dialogues':'Empathetic Dialogues',
'personachat': 'Persona-Chat'}
print('Dataset & Method & Percent not relevant & Percent score $<$ 3\\\\ \hline \hline')
for (dataset, model), df in elim_df.groupby(['dataset', 'model']):
print('\multirow{4}{*}{\parbox{1.8cm}{\\vspace{.1cm} %s}}' % dataset_name[dataset])
method_grouped = df.groupby('method')
for method in method_order:
try:
method_df = method_grouped.get_group(method)
perc_not_rel = 100.0 * np.mean(method_df.groupby('context')['not_rel'].agg(np.sum)>1)
below_3 = 100.0*np.mean(method_df.groupby('context')['well_score'].agg(np.mean) < 3)
if method=='face':
print('\t & %s & %.1f & %.1f \\\\ \\hline' % \
(method_name[method], perc_not_rel, below_3))
else:
print('\t & %s & %.1f & %.1f \\\\ ' % \
(method_name[method], perc_not_rel, below_3))
except KeyError:
pass
# -
# +
off = 0
method_names = []
fig, ax = plt.subplots(figsize=(10, 10))
dataset_color = {'cornell_movie':'plum',
'dailydialog':'skyblue',
'empathetic': 'palevioletred',
'empathetic_dialogues':'palevioletred',
'personachat': 'lightsalmon'}
for method, method_df in elim_df.groupby('method'):
ind = 0
method_names.append(method)
for (model, task), df in method_df.groupby(['model', 'dataset']):
well_score = df['well_score'].mean()
# well_std = df['well_score'].std() # This is across all inputs, but we want across averged inputs?
well_std = df.groupby('context')['well_score'].agg(np.mean).std()
if method == 'vanilla':
plt.bar(ind + off*5, height=well_score, yerr=well_std, color=dataset_color[task], label=task)
else:
plt.bar(ind + off*5, height=well_score, yerr=well_std, color=dataset_color[task])
ind += 1
off += 1
plt.xticks(1+5*np.arange(4), method_names, fontsize=20)
plt.yticks(np.arange(6), fontsize=20)
plt.ylabel('Well-formed score (1-5)', fontsize=20)
plt.ylim([1,5.5])
plt.legend()
# Shrink current axis by 20%
box = ax.get_position()
ax.set_position([box.x0, box.y0-box.height *.3, box.width, box.height * 0.7])
# Put a legend to the right of the current axis
ax.legend(loc='lower left', bbox_to_anchor=(0, 1), ncol=2, fontsize=20)
plt.show()
# +
# df.groupby('context')['well_score'].agg([np.mean, np.size])#.std()
# -
for method in elim_df['method'].unique():
plt.hist(elim_df.groupby('method').get_group(method)['well_score'].values, bins = 5)
plt.title('%s (std: %s)' % (method, elim_df.groupby('method').get_group(method).groupby('context')['well_score'].agg(np.mean).std()))
plt.show()
# +
scored_untied_df = []
for (dataset, model, method), setting_df in elim_df.groupby(['dataset', 'model', 'method']):
for context, df in setting_df.groupby('context'):
if df.shape[0] !=3:
print(dataset, model, method, df.shape)
scored_untied_df.append([dataset, model, method, context, df['response'].values[0],
df['well_score'].mean(),
Counter(df['not_rel'].values).most_common(1)[0][0] == True])
scored_untied_df = pd.DataFrame(scored_untied_df, columns = ['dataset', 'model', 'method', 'context',
'response', 'avg_well_score', 'not_rel'])
# -
for (dataset, model, method), setting_df in elim_df.groupby(['dataset', 'model', 'method']):
print(dataset, model, method, 'missing: ', 100-len(setting_df.groupby('context').groups))
# if len(setting_df.groupby('context').groups) < 100:
# break
# for context, df in setting_df.groupby('context'):
len(setting_df['context'].unique())
# # Load Data that Duels Vanilla & IDF
# +
# raw_duel_df = pd.read_csv('batches/Batch_3630895_batch_results.csv')
raw_duel_df = pd.concat([pd.read_csv('../batches/Batch_3630895_batch_results.csv'),
pd.read_csv('../batches/Batch_3636922_batch_results.csv'),
pd.read_csv('../batches/Batch_3638556_batch_results.csv'),
pd.read_csv('../batches/Batch_3638681_batch_results.csv')], axis=0, sort=False).reset_index()
# -
raw_duel_df.shape
# +
dueling_methods = ['vanilla', 'idf']
duel_df = []
for i in raw_duel_df.index:
row = raw_duel_df.loc[i]
hit_num = i
worker_id = row['WorkerId']
for ex in range(10): # given to each turker in a single HIT
if np.isnan(row['Answer.best_%s' % ex]):
# This may be null from the re-run subset, so skip it (and count after!)
continue
option_to_method = {}
method_to_resp = {}
for option in range(1, 2+1):
model_pieces = row['Input.model_%s_%s' % (option, ex)].split(' ')
dataset, model, method = model_pieces
option_to_method[option] = method
method_to_resp[method] = row['Input.resp_%s_%s' % (option, ex)]
best_overall = option_to_method[row['Answer.best_%s' % (ex,)]]
most_interesting = option_to_method[row['Answer.int_%s' % (ex,)]]
most_relevant = option_to_method[row['Answer.rel_%s' % (ex,)]]
none_int = type(row['Answer.check_int_%s'% (ex,)]) == str
none_rel = type(row['Answer.check_rel_%s'% (ex,)]) == str
message1 = row['Input.msg1_%s' % (ex,)]
message2 = row['Input.msg2_%s' % (ex,)]
duel_df.append([hit_num, worker_id, dataset, model,
message1, message2] + \
[method_to_resp[x] for x in dueling_methods] + \
[best_overall, most_interesting, most_relevant, none_int, none_rel])
duel_df = pd.DataFrame(duel_df, columns = ['hit_num', 'worker_id', 'dataset', \
'model', 'message1', 'message2',] + \
['response_%s' % x for x in dueling_methods] + \
['best_overall', 'most_int', 'most_rel', 'none_int', 'none_rel'])
duel_df['context'] = duel_df['message1'] + duel_df['message2']
# remove examples (accidentially included) where message 2 was null
duel_df = duel_df[[type(x) == str for x in duel_df['message2']]]
# -
duel_df[[type(x) != str for x in duel_df['message2']]].shape
print(duel_df[[type(x) == str for x in duel_df['message2']]].shape)
print(duel_df['context'].unique().shape)
for (dataset, model), setting_df in duel_df.groupby(['dataset', 'model']):
print(dataset, model, method, 'missing: ', 100-len(setting_df.groupby('context').groups))
for context, setting_df in duel_df.groupby('context'):
if setting_df.shape[0] != 6:
print(setting_df.shape[0], context)
# print(dataset, model, method, 'missing: ', 100-len(setting_df.groupby('context').groups))
raw_duel_df['Answer.optionalfeedback'].values
np.sum(rel_and_int), rel_and_int.shape
# ### Look at counts of when rel & int, etc., overall.
# +
print('OVERALL:')
rel_and_int = ~(duel_df['none_rel'] | duel_df['none_int'])
a = duel_df['most_rel'] != duel_df['most_int']
b = duel_df['most_rel'] == duel_df['best_overall']
print('Fraction most_rel != most_int %.3f %d' % (np.mean(a[rel_and_int]), len(a[rel_and_int])))
print(sum(a[rel_and_int]))
print("Fraction most_rel == best %.3f %d" % (np.mean(b[~duel_df['none_rel']]), len(b[~duel_df['none_rel']])))
print(sum(b[~duel_df['none_rel']]))
print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f %d" % \
(np.sum(a[rel_and_int] & b[rel_and_int]) / np.sum(a[rel_and_int]), len(a[rel_and_int])))
# for (dataset, model), setting_df in duel_df.groupby(['dataset', 'model']):
# print("\n\n%s %s" % (dataset, model))
# print('Counts over all:')
# print('Best overall: ', Counter(setting_df['best_overall']))
# print('Most Interesting: ', Counter(setting_df['most_int']))
# print('Most relevant: ', Counter(setting_df['most_rel']))
# a = setting_df['most_rel'] != setting_df['most_int']
# b = setting_df['most_rel'] == setting_df['best_overall']
# print('Fraction most_rel != most_int %.3f' % np.mean(a))
# print("Fraction most_rel == best %.3f" % np.mean(b))
# print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f" % \
# (np.sum(a&b) / np.sum(a)))
print('\n\n\nUNTIED OVERALL:')
rel_and_int = ~(duel_untied_df['none_rel'] | duel_untied_df['none_int'])
a = duel_untied_df['most_rel'] != duel_untied_df['most_int']
b = duel_untied_df['most_rel'] == duel_untied_df['best_overall']
print('Fraction most_rel != most_int %.3f %d' % (np.mean(a[rel_and_int]), len(a)))
print()
print("Fraction most_rel == best %.3f %d" % (np.mean(b[~duel_untied_df['none_rel']]), len(b)))
print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f %d" % \
((np.sum(a[rel_and_int] & b[rel_and_int]) / np.sum(a[rel_and_int])), len(a)))
# -
duel_untied_df.columns
import nltk
def test(x, y):
if nltk.edit_distance(x,y) > 5:
return True
else:
return False
# +
no_similar_resps = duel_untied_df[duel_untied_df.apply(lambda x: test(x["response_vanilla"], x["response_idf"]), axis=1)]
a = no_similar_resps['most_rel'] != no_similar_resps['most_int']
b = no_similar_resps['most_rel'] == no_similar_resps['best_overall']
print("No similar responses")
print('Fraction most_rel != most_int %.3f' % np.mean(a[rel_and_int]))
print("Fraction most_rel == best %.3f" % np.mean(b[~duel_untied_df['none_rel']]))
print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f" % \
(np.sum(a[rel_and_int] & b[rel_and_int]) / np.sum(a[rel_and_int])))
print("\n\n\n")
no_short_resps = duel_untied_df[duel_untied_df['response_vanilla'].apply(lambda x: len(x.split(" "))>6) & duel_untied_df['response_idf'].apply(lambda x: len(x.split(" "))>6)]
a = no_short_resps['most_rel'] != no_short_resps['most_int']
b = no_short_resps['most_rel'] == no_short_resps['best_overall']
print("No short responses")
print('Fraction most_rel != most_int %.3f' % np.mean(a[rel_and_int]))
print("Fraction most_rel == best %.3f" % np.mean(b[~duel_untied_df['none_rel']]))
print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f" % \
(np.sum(a[rel_and_int] & b[rel_and_int]) / np.sum(a[rel_and_int])))
print("\n\n\n")
no_similar_or_short_resps = duel_untied_df[duel_untied_df.apply(lambda x: test(x["response_vanilla"], x["response_idf"]), axis=1) & (duel_untied_df['response_vanilla'].apply(lambda x: len(x.split(" "))>6) & duel_untied_df['response_idf'].apply(lambda x: len(x.split(" "))>6))]
a = no_similar_or_short_resps['most_rel'] != no_similar_or_short_resps['most_int']
b = no_similar_or_short_resps['most_rel'] == no_similar_or_short_resps['best_overall']
print("No short or similar responses")
print('Fraction most_rel != most_int %.3f' % np.mean(a[rel_and_int]))
print("Fraction most_rel == best %.3f" % np.mean(b[~duel_untied_df['none_rel']]))
print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f" % \
(np.sum(a[rel_and_int] & b[rel_and_int]) / np.sum(a[rel_and_int])))
print("\n\n\n")
one_long_resp = duel_untied_df[(duel_untied_df['response_vanilla'].apply(lambda x: len(x.split(" "))>6) & duel_untied_df['response_idf'].apply(lambda x: len(x.split(" "))<6) | duel_untied_df['response_vanilla'].apply(lambda x: len(x.split(" "))<6) & duel_untied_df['response_idf'].apply(lambda x: len(x.split(" "))>6))]
a = one_long_resp['most_rel'] != one_long_resp['most_int']
b = one_long_resp['most_rel'] == one_long_resp['best_overall']
print("One long and one short response")
print('Fraction most_rel != most_int %.3f' % np.mean(a[rel_and_int]))
print("Fraction most_rel == best %.3f" % np.mean(b[~duel_untied_df['none_rel']]))
print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f" % \
(np.sum(a[rel_and_int] & b[rel_and_int]) / np.sum(a[rel_and_int])))
print("\n\n\n")
no_both_short = duel_untied_df[~(duel_untied_df['response_vanilla'].apply(lambda x: len(x.replace(" .","").replace(" ,","").replace(" ?","").replace(" !","").split(" "))<6) & duel_untied_df['response_idf'].apply(lambda x: len(x.split(" "))<6))]
a = no_both_short['most_rel'] != no_both_short['most_int']
b = no_both_short['most_rel'] == no_both_short['best_overall']
print("One long and one short response")
print('Fraction most_rel != most_int %.3f' % np.mean(a[rel_and_int]))
print("Fraction most_rel == best %.3f" % np.mean(b[~duel_untied_df['none_rel']]))
print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f" % \
(np.sum(a[rel_and_int] & b[rel_and_int]) / np.sum(a[rel_and_int])))
print("\n\n\n")
# -
def is_short(string):
string = string.replace(" .","").replace(" ,","").replace(" ?","").replace(" !","")
if string.split(" ") > 5:
return True
return False
no_both_short = duel_untied_df[~(duel_untied_df['response_vanilla'].apply(lambda x: ~is_short(x)) & duel_untied_df['response_idf'].apply(lambda x: ~is_short(x)))]
a = no_both_short['most_rel'] != no_both_short['most_int']
b = no_both_short['most_rel'] == no_both_short['best_overall']
print("One long and one short response")
print('Fraction most_rel != most_int %.3f' % np.mean(a[rel_and_int]))
print("Fraction most_rel == best %.3f" % np.mean(b[~duel_untied_df['none_rel']]))
print("Fraction ((most_rel != most_int) & (most_rel == best))/(most_rel != most_int) %.3f" % \
(np.sum(a[rel_and_int] & b[rel_and_int]) / np.sum(a[rel_and_int])))
print("\n\n\n")
one_long_resp
# ### Look at counts when total crap is removed.
for (dataset, model), setting_df in duel_df.groupby(['dataset', 'model']):
print("\n\n%s %s" % (dataset, model))
print('Counts with crap removed:')
print('Best overall with not relevant and not interesting removed: ',
Counter(setting_df['best_overall'][~(setting_df['none_int'] & setting_df['none_rel'])]),
' (%s removed)' % np.sum(setting_df['none_int'] & setting_df['none_rel']))
print('Most int with not interesting removed: ',
Counter(setting_df['most_int'][~setting_df['none_int']]),
' (%s removed)' % np.sum(setting_df['none_int']))
print('Most relevant with not relevant removed: ',
Counter(setting_df['most_rel'][~setting_df['none_rel']]),
' (%s removed)' % np.sum(setting_df['none_rel']))
duel_df[:3]
# duel_df.columns
# df['response'][0]
df['response_vanilla'].values[0]
# +
duel_untied_df = []
for (dataset, model), setting_df in duel_df.groupby(['dataset', 'model']):
for context, df in setting_df.groupby('context'):
duel_untied_df.append([dataset, model, context,
df['response_vanilla'].values[0], df['response_idf'].values[0],
Counter(df['best_overall'].values).most_common(1)[0][0],
Counter(df['most_rel'].values).most_common(1)[0][0],
Counter(df['most_int'].values).most_common(1)[0][0],
Counter(df['none_rel'].values).most_common(1)[0][0] == True,
Counter(df['none_int'].values).most_common(1)[0][0] == True])
duel_untied_df = pd.DataFrame(duel_untied_df, columns = ['dataset', 'model', 'context',
'response_vanilla', 'response_idf',
'best_overall', 'most_rel', 'most_int',
'none_rel', 'none_int'])
# +
print('Counts over examples (i.e., tie-broken):')
for (dataset, model), setting_df in duel_untied_df.groupby(['dataset', 'model']):
print("\n\n%s %s" % (dataset, model))
print('Best overall: ', Counter(setting_df['best_overall']))
print('Most Interesting: ', Counter(setting_df['most_int']))
print('Most relevant: ', Counter(setting_df['most_rel']))
print('Fraction most_rel != most_int %.3f' % np.mean(setting_df['most_rel'] != setting_df['most_int']))
print("Fraction most_rel == best %.3f" % np.mean(setting_df['most_rel'] == setting_df['best_overall']))
# +
print('Counts with crap removed:')
for (dataset, model), setting_df in duel_untied_df.groupby(['dataset', 'model']):
print("\n\n%s %s" % (dataset, model))
print('Best overall with not relevant and not interesting removed: ',
Counter(setting_df['best_overall'][~(setting_df['none_int'] & setting_df['none_rel'])]),
' (%s removed)' % np.sum(setting_df['none_int'] & setting_df['none_rel']))
print('Most int with not interesting removed: ',
Counter(setting_df['most_int'][~setting_df['none_int']]),
' (%s removed)' % np.sum(setting_df['none_int']))
print('Most relevant with not relevant removed: ',
Counter(setting_df['most_rel'][~setting_df['none_rel']]),
' (%s removed)' % np.sum(setting_df['none_rel']))
# +
fig, ax = plt.subplots(figsize=(7,5))
for (model, dataset), setting_df in duel_untied_df.groupby(['model', 'dataset']):
c_best = Counter(setting_df['best_overall'])
c_int = Counter(setting_df['most_int'][~setting_df['none_int']])
c_rel = Counter(setting_df['most_rel'][~setting_df['none_rel']])
if model== 'transformer':
sym = '*'
mod='trans'
else:
sym='o'
mod='s2s'
ax.plot([1, 2], [c_int['vanilla'], c_int['idf']],'%s-'%sym, label='%s (%s)' % (dataset, mod), color=dataset_color[dataset])
ax.plot([4, 5], [c_rel['vanilla'], c_rel['idf']],'%s-'%sym, color=dataset_color[dataset])
ax.plot([7, 8], [c_best['vanilla'], c_best['idf']],'%s-'%sym, color=dataset_color[dataset])
# Shrink current axis by 20%
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
# Put a legend to the right of the current axis
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# plt.legend()
plt.xlabel('Most Interesting Most Relevant Best Overall',fontsize=10)
plt.ylabel('Number responses',fontsize=16)
plt.xticks([1,2, 4,5, 7,8], ['Unweighted', 'IDF', 'Unweighted', 'IDF','Unweighted', 'IDF'])
plt.ylim([30,70])
plt.show()
# +
dataset_abbrv = {'cornell_movie':'Cor',
'dailydialog':'Dai',
'empathetic': 'Emp',
'empathetic_dialogues':'Emp',
'personachat': 'Per'}
model_abbrv = {'seq2seq':'S2S',
'transformer': "Tfm"}
# dataset_color = {'cornell_movie':'plum',
# 'dailydialog':'skyblue',
# 'empathetic': 'palevioletred',
# 'empathetic_dialogues':'palevioletred',
# 'personachat': 'lightsalmon'}
fig, ax = plt.subplots(figsize=(4,10))
h = 0
plt.plot([50,50],[-1, 27], color='grey', linewidth=3)
ylabels = []
for (model, dataset), setting_df in duel_untied_df.groupby(['model', 'dataset']):
c_best = Counter(setting_df['best_overall'])
c_int = Counter(setting_df['most_int'][~setting_df['none_int']])
c_rel = Counter(setting_df['most_rel'][~setting_df['none_rel']])
d = float(sum(c_int.values()))
plt.barh(h, 100.*c_int['vanilla']/d, left=0, color='pink')
plt.barh(h, 100.*c_int['idf']/d, left=100.*c_int['vanilla']/d, color='mediumorchid')
d = float(sum(c_rel.values()))
plt.barh(h+9, 100.*c_rel['vanilla']/d, left=0, color='pink')
plt.barh(h+9, 100.*c_rel['idf']/d, left=100.*c_rel['vanilla']/d, color='mediumorchid')
d = float(sum(c_best.values()))
if model=='transformer' and dataset=='personachat':
plt.barh(h+18, 100.*c_best['vanilla']/d, left=0, color='pink', label='unweighted')
plt.barh(h+18, 100.*c_best['idf']/d, left=100.*c_best['vanilla']/d, color='mediumorchid', label='idf')
else:
plt.barh(h+18, 100.*c_best['vanilla']/d, left=0, color='pink')
plt.barh(h+18, 100.*c_best['idf']/d, left=100.*c_best['vanilla']/d, color='mediumorchid')
ylabels.append('%s:%s' % (dataset_abbrv[dataset], model_abbrv[model]))
# plt.plot([3, 4], [c_rel['vanilla'], c_rel['idf']],'o-')
# plt.plot([5, 6], [c_best['vanilla'], c_best['idf']],'o-')
h += 1
# plt.legend()
plt.ylabel('Most Interesting Most Relevant Best Overall', fontsize=16)
plt.xlabel('Percent of examples',fontsize=20)
# plt.xticks(1+np.arange(6), ['Unweighted', 'IDF', 'Unweighted', 'IDF','Unweighted', 'IDF'])
plt.yticks(np.arange(3*(len(ylabels)+1)), ylabels + [''] + ylabels + [''] + ylabels, fontsize=13)
plt.xlim([0,100])
plt.ylim([-1, 26])
# Shrink current axis by 20%
box = ax.get_position()
ax.set_position([box.x0, box.y0-box.height *.3, box.width, box.height * 0.7])
# Put a legend to the right of the current axis
ax.legend(loc='lower left', bbox_to_anchor=(0, 1), ncol=2, fontsize=14)
plt.show()
# -
# # Append additional metrics of diversity
idf_dict.keys()
# +
# scored_untied_df.columns
for ind in scored_untied_df.index:
if scored_untied_df.loc[ind, 'dataset'] == 'empathetic':
dataset_idf_dict = idf_dict['empathetic_dialogues']
else:
dataset_idf_dict = idf_dict[scored_untied_df.loc[ind, 'dataset']]
resp_a = scored_untied_df.loc[ind, 'response'].split(' ')
idf_a = [dataset_idf_dict[x] for x in resp_a if x in dataset_idf_dict.keys()]
tmp = [x for x in resp_a if x not in dataset_idf_dict.keys()]
if tmp != []:
print(tmp)
scored_untied_df.loc[ind, 'len'] = len(resp_a)
scored_untied_df.loc[ind, 'max_idf'] = np.max(idf_a)
scored_untied_df.loc[ind, 'avg_idf'] = np.mean(idf_a)
# -
scored_untied_df[:3]
for ind in duel_untied_df.index:
dataset_idf_dict = idf_dict[duel_untied_df.loc[ind, 'dataset']]
for suff in ['idf' ,'vanilla']:
resp_a = duel_untied_df.loc[ind, 'response_%s'%suff].split(' ')
idf_a = [dataset_idf_dict[x] for x in resp_a if x in dataset_idf_dict.keys()]
tmp = [x for x in resp_a if x not in dataset_idf_dict.keys()]
if tmp != []:
print(tmp)
duel_untied_df.loc[ind, 'len_%s'%suff] = len(resp_a)
duel_untied_df.loc[ind, 'max_idf_%s'%suff] = np.max(idf_a)
duel_untied_df.loc[ind, 'avg_idf_%s'%suff] = np.mean(idf_a)
# ### Look at how wellness score relates to auto-metrics
for (dataset, model), df in scored_untied_df.groupby(['dataset', 'model']):
fig, ax = plt.subplots(1,3, figsize=(15,4))
ax[0].plot(df['avg_idf'], df['avg_well_score'], 'o', alpha=.3)
ax[0].set_xlabel('avg_idf')
ax[1].plot(df['max_idf'], df['avg_well_score'], 'o', alpha=.3)
ax[1].set_xlabel('max_idf')
ax[2].plot(df['len'], df['avg_well_score'], 'o', alpha=.3)
ax[2].set_xlabel('response length')
ax[0].set_ylabel('avg. well-formed score')
plt.suptitle('%s %s' % (dataset, model))
plt.show()
# ### RELEVANCE:
# +
hist_info_rel = []
for (dataset, model), df in duel_untied_df.groupby(['dataset', 'model']):
fig, ax = plt.subplots(1,3, figsize=(15,4))
diffs = np.NaN*np.zeros((3, df.shape[0]))
for i, ind in enumerate(df.index):
# win on right
if df.loc[ind, 'most_rel'] == 'vanilla':
ax[0].plot([0, 1], [df.loc[ind, 'avg_idf_idf'], df.loc[ind, 'avg_idf_vanilla']], 'bo-', alpha=.3)
ax[1].plot([0, 1], [df.loc[ind, 'max_idf_idf'], df.loc[ind, 'max_idf_vanilla']], 'bo-', alpha=.3)
ax[2].plot([0, 1], [df.loc[ind, 'len_idf'], df.loc[ind, 'len_vanilla']], 'bo-', alpha=.3)
diffs[0, i] = df.loc[ind, 'avg_idf_vanilla'] - df.loc[ind, 'avg_idf_idf']
diffs[1, i] = df.loc[ind, 'max_idf_vanilla'] - df.loc[ind, 'max_idf_idf']
diffs[2, i] = df.loc[ind, 'len_vanilla'] - df.loc[ind, 'len_idf']
else: # idf won
ax[0].plot([0, 1], [df.loc[ind, 'avg_idf_vanilla'], df.loc[ind, 'avg_idf_idf']], 'b-o', alpha=.3)
ax[1].plot([0, 1], [df.loc[ind, 'max_idf_vanilla'], df.loc[ind, 'max_idf_idf']], 'b-o', alpha=.3)
ax[2].plot([0, 1], [df.loc[ind, 'len_vanilla'], df.loc[ind, 'len_idf']], 'b-o', alpha=.3)
diffs[0, i] = df.loc[ind, 'avg_idf_idf'] - df.loc[ind, 'avg_idf_vanilla']
diffs[1, i] = df.loc[ind, 'max_idf_idf'] - df.loc[ind, 'max_idf_vanilla']
diffs[2, i] = df.loc[ind, 'len_idf'] - df.loc[ind, 'len_vanilla']
hist_info_rel.append([dataset, model, diffs])
ax[0].set_ylabel('avg_idf')
ax[1].set_ylabel('max_idf')
ax[2].set_ylabel('response length')
ax[0].set_xticks([0,1])
ax[0].set_xticklabels(['lose', 'win'])
ax[1].set_xticks([0,1])
ax[1].set_xticklabels(['lose', 'win'])
ax[2].set_xticks([0,1])
ax[2].set_xticklabels(['lose', 'win'])
plt.suptitle('RELEVANCE: %s %s' % (dataset, model))
plt.show()
# +
fig, ax = plt.subplots(1,3, figsize=(15,4))
for i in range(len(hist_info)):
diffs = hist_info_rel[i][2]
y, bin_edges = np.histogram(diffs[0,:], bins=np.arange(-10, 10, 1))
ax[0].plot(bin_edges[:-1], y, 'o-', label='%s %s' % (hist_info_rel[0], hist_info_rel[1]))
y, bin_edges = np.histogram(diffs[1,:], bins=np.arange(-10, 10, 1))
ax[1].plot(bin_edges[:-1], y, 'o-', label='%s %s' % (hist_info_rel[0], hist_info_rel[1]))
y, bin_edges = np.histogram(diffs[2,:], bins=np.arange(-30, 30, 2))
ax[2].plot(bin_edges[:-1], y, 'o-', label='%s %s' % (hist_info_rel[0], hist_info_rel[1]))
ax[0].set_xlabel('difference of avg_idf of win - lose')
ax[1].set_xlabel('difference of max_idf of win - lose')
ax[2].set_xlabel('difference of length of win - lose')
ax[0].set_ylabel('number of examples')
plt.suptitle('RELEVANT')
plt.show()
# -
# ### INTERESTING
# +
hist_info = []
for (dataset, model), df in duel_untied_df.groupby(['dataset', 'model']):
fig, ax = plt.subplots(1,3, figsize=(15,4))
diffs = np.NaN*np.zeros((3, df.shape[0]))
for i, ind in enumerate(df.index):
# win on right
if df.loc[ind, 'most_int'] == 'vanilla':
ax[0].plot([0, 1], [df.loc[ind, 'avg_idf_idf'], df.loc[ind, 'avg_idf_vanilla']], 'o-', alpha=.3, color='purple')
ax[1].plot([0, 1], [df.loc[ind, 'max_idf_idf'], df.loc[ind, 'max_idf_vanilla']], 'o-', alpha=.3, color='purple')
ax[2].plot([0, 1], [df.loc[ind, 'len_idf'], df.loc[ind, 'len_vanilla']], 'o-', alpha=.3, color='purple')
diffs[0, i] = df.loc[ind, 'avg_idf_vanilla'] - df.loc[ind, 'avg_idf_idf']
diffs[1, i] = df.loc[ind, 'max_idf_vanilla'] - df.loc[ind, 'max_idf_idf']
diffs[2, i] = df.loc[ind, 'len_vanilla'] - df.loc[ind, 'len_idf']
else: # idf won
ax[0].plot([0, 1], [df.loc[ind, 'avg_idf_vanilla'], df.loc[ind, 'avg_idf_idf']], 'o-', alpha=.3, color='purple')
ax[1].plot([0, 1], [df.loc[ind, 'max_idf_vanilla'], df.loc[ind, 'max_idf_idf']], 'o-', alpha=.3, color='purple')
ax[2].plot([0, 1], [df.loc[ind, 'len_vanilla'], df.loc[ind, 'len_idf']], 'o-', alpha=.3, color='purple')
diffs[0, i] = df.loc[ind, 'avg_idf_idf'] - df.loc[ind, 'avg_idf_vanilla']
diffs[1, i] = df.loc[ind, 'max_idf_idf'] - df.loc[ind, 'max_idf_vanilla']
diffs[2, i] = df.loc[ind, 'len_idf'] - df.loc[ind, 'len_vanilla']
ax[0].set_ylabel('avg_idf')
ax[1].set_ylabel('max_idf')
ax[2].set_ylabel('response length')
ax[0].set_xticks([0,1])
ax[0].set_xticklabels(['lose', 'win'])
ax[1].set_xticks([0,1])
ax[1].set_xticklabels(['lose', 'win'])
ax[2].set_xticks([0,1])
ax[2].set_xticklabels(['lose', 'win'])
plt.suptitle('INTERESTING: %s %s' % (dataset, model))
plt.show()
hist_info.append([dataset, model, diffs])
# +
fig, ax = plt.subplots(1,3, figsize=(15,4))
for i in range(len(hist_info)):
diffs = hist_info[i][2]
y, bin_edges = np.histogram(diffs[0,:], bins=np.arange(-10, 10, 1))
ax[0].plot(bin_edges[:-1], y, 'o-', label='%s %s' % (hist_info[0], hist_info[1]))
y, bin_edges = np.histogram(diffs[1,:], bins=np.arange(-10, 10, 1))
ax[1].plot(bin_edges[:-1], y, 'o-', label='%s %s' % (hist_info[0], hist_info[1]))
y, bin_edges = np.histogram(diffs[2,:], bins=np.arange(-30, 30, 2))
ax[2].plot(bin_edges[:-1], y, 'o-', label='%s %s' % (hist_info[0], hist_info[1]))
ax[0].set_xlabel('difference of avg_idf of win - lose')
ax[1].set_xlabel('difference of max_idf of win - lose')
ax[2].set_xlabel('difference of length of win - lose')
ax[0].set_ylabel('number of examples')
plt.suptitle('INTERESTING')
plt.show()
# +
fig, ax = plt.subplots(figsize=(5,4))
for i in range(len(hist_info)):
diffs = hist_info[i][2]
y, bin_edges = np.histogram(diffs[1,:], bins=np.arange(-10, 10, 1))
ax.plot(bin_edges[:-1], y, 'o-g', label='%s %s' % (hist_info[0], hist_info[1]), alpha=.4)
ax.set_xlabel('max_idf of win - max_idf lose', fontsize=18)
ax.set_ylabel('Number of examples', fontsize=18)
# plt.suptitle('INTERESTING')
plt.show()
# -
# scored_untied_df.merge(duel_untied_df)
t2 = duel_untied_df
# t2
# +
t1_vanilla = scored_untied_df[scored_untied_df['method'] == 'vanilla'][['dataset', 'model',
'context', 'avg_well_score', 'not_rel']]
t1_vanilla = t1_vanilla.rename(columns = {'avg_well_score':'avg_well_score_vanilla', 'not_rel':'not_rel_vanilla'})
t1_idf = scored_untied_df[scored_untied_df['method'] == 'idf'][['dataset', 'model',
'context', 'avg_well_score', 'not_rel']]
t1_idf = t1_idf.rename(columns = {'avg_well_score':'avg_well_score_idf', 'not_rel':'not_rel_idf'})
# -
# t3 = t1.merge(t2, on=['dataset', 'model', 'context'], how='left')
t3 = t2.merge(t1_vanilla, on=['dataset', 'model', 'context'], how = 'right')
t3 = t3.merge(t1_idf, on=['dataset', 'model', 'context'], how = 'right')
# +
# t3.shape
# +
for (dataset, model), df in t3.groupby(['dataset', 'model']):
fig, ax = plt.subplots(1,2, figsize=(10,4))
diffs = np.NaN*np.zeros((3, df.shape[0]))
for i, ind in enumerate(df.index):
# win on right
if df.loc[ind, 'most_rel'] == 'vanilla':
ax[0].plot([0, 1], [df.loc[ind, 'avg_well_score_idf'], df.loc[ind, 'avg_well_score_vanilla']], 'bo-', alpha=.3)
else: # idf won
ax[0].plot([0, 1], [df.loc[ind, 'avg_well_score_vanilla'], df.loc[ind, 'avg_well_score_idf']], 'b-o', alpha=.3)
for i, ind in enumerate(df.index):
# win on right
if df.loc[ind, 'most_int'] == 'vanilla':
ax[1].plot([0, 1], [df.loc[ind, 'avg_well_score_idf'], df.loc[ind, 'avg_well_score_vanilla']], 'bo-', alpha=.3)
else: # idf won
ax[1].plot([0, 1], [df.loc[ind, 'avg_well_score_vanilla'], df.loc[ind, 'avg_well_score_idf']], 'b-o', alpha=.3)
ax[0].set_ylabel('avg_well_score')
ax[0].set_title('most relevant')
ax[1].set_title('most interesting')
ax[0].set_xticks([0,1])
ax[0].set_xticklabels(['lose', 'win'])
ax[1].set_xticks([0,1])
ax[1].set_xticklabels(['lose', 'win'])
# ax[2].set_xticks([0,1])
# ax[2].set_xticklabels(['lose', 'win'])
plt.suptitle('%s %s' % (dataset, model))
plt.show()
# fig, ax = plt.subplots(1,3, figsize=(15,4))
# ax[0].hist(diffs[0,:])
# ax[1].hist(diffs[1,:])
# ax[2].hist(diffs[2,:])
# ax[0].set_xlabel('avg_idf')
# ax[1].set_xlabel('max_idf')
# ax[2].set_xlabel('response length')
# ax[0].set_ylabel('number of examples')
# plt.suptitle('RELEVANCE: %s %s' % (dataset, model))
# plt.show()
# +
fig, ax = plt.subplots(1,3, figsize=(15,4))
for (dataset, model), df in t3.groupby(['dataset', 'model']):
rel_diffs = np.NaN*np.zeros(df.shape[0])
int_diffs = np.NaN*np.zeros(df.shape[0])
best_diffs = np.NaN*np.zeros(df.shape[0])
for i, ind in enumerate(df.index):
# win on right
if df.loc[ind, 'most_rel'] == 'vanilla':
rel_diffs[i] = df.loc[ind, 'avg_well_score_vanilla'] - df.loc[ind, 'avg_well_score_idf']
else: # idf won
rel_diffs[i] = df.loc[ind, 'avg_well_score_idf'] - df.loc[ind, 'avg_well_score_vanilla']
for i, ind in enumerate(df.index):
# win on right
if df.loc[ind, 'most_int'] == 'vanilla':
int_diffs[i] = df.loc[ind, 'avg_well_score_vanilla'] - df.loc[ind, 'avg_well_score_idf']
else: # idf won
int_diffs[i] = df.loc[ind, 'avg_well_score_idf'] - df.loc[ind, 'avg_well_score_vanilla']
for i, ind in enumerate(df.index):
# win on right
if df.loc[ind, 'best_overall'] == 'vanilla':
best_diffs[i] = df.loc[ind, 'avg_well_score_vanilla'] - df.loc[ind, 'avg_well_score_idf']
else: # idf won
best_diffs[i] = df.loc[ind, 'avg_well_score_idf'] - df.loc[ind, 'avg_well_score_vanilla']
y, bin_edges = np.histogram(rel_diffs)
ax[0].plot(bin_edges[:-1], y, '-o', label='%s %s' % (dataset, model))
y, bin_edges = np.histogram(int_diffs)
ax[1].plot(bin_edges[:-1], y, '-o', label='%s %s' % (dataset, model))
y, bin_edges = np.histogram(best_diffs)
ax[2].plot(bin_edges[:-1], y, '-o', label='%s %s' % (dataset, model))
ax[0].set_ylabel('number of examples', fontsize=18)
ax[0].set_xlabel('diff (win - losing response) \n in avg_well_score ', fontsize=18)
ax[1].set_xlabel('diff (win - losing response) \n in avg_well_score', fontsize=18)
ax[2].set_xlabel('diff (win - losing response) \n in avg_well_score', fontsize=18)
ax[0].set_title('most relevant', fontsize=18)
ax[1].set_title('most interesting', fontsize=18)
ax[2].set_title('best overall', fontsize=18)
# ax[0].set_xticks([0,1])
# ax[0].set_xticklabels(['lose', 'win'])
# ax[1].set_xticks([0,1])
# ax[1].set_xticklabels(['lose', 'win'])
# plt.suptitle('%s %s' % (dataset, model))
plt.show()
# +
auto_metrics_df = pd.DataFrame([['seq2seq', 'personachat', 'vanilla' ,2.067 ,4.528 ,10.963 ,0.005 ,0.021 ,0.299 ,3.663 ],
['seq2seq', 'personachat', 'swapping' ,2.417 ,5.032 ,13.110 ,0.008 ,0.033 ,0.340 ,3.563 ],
['seq2seq', 'personachat', 'idf' ,2.573 ,4.982 ,10.457 ,0.010 ,0.037 ,0.310 ,4.173 ],
['seq2seq', 'personachat', 'face' , 3.936 , 6.962 , 26.542 , 0.010 , 0.075 , 0.888 , 5.620 ],
['seq2seq', 'dailydialog', 'vanilla' ,2.426 ,4.948 ,7.917 ,0.020 ,0.090 ,0.320 ,4.269 ],
['seq2seq', 'dailydialog', 'swapping' ,2.990 ,5.811 ,9.938 ,0.033 ,0.145 ,0.512 ,5.143 ],
['seq2seq', 'dailydialog', 'idf' ,3.140 ,6.272 ,12.829 , 0.033 , 0.159 ,0.607 ,5.454 ],
['seq2seq', 'dailydialog', 'face' , 3.554 , 6.580 , 22.543 ,0.017 ,0.109 , 0.676 , 5.602 ],
['seq2seq', 'empathetic_dialogues', 'vanilla' ,2.279 ,4.451 ,11.812 ,0.005 ,0.018 ,0.193 ,3.153 ],
['seq2seq', 'empathetic_dialogues', 'swapping' ,2.699 ,5.115 ,9.313 ,0.011 ,0.043 ,0.343 ,4.154 ],
['seq2seq', 'empathetic_dialogues', 'idf' ,2.849 ,5.287 ,9.658 ,0.010 ,0.040 ,0.351 ,4.110 ],
['seq2seq', 'empathetic_dialogues', 'face' , 3.327 , 6.669 , 17.002 , 0.031 , 0.205 , 0.922 , 5.602 ],
['seq2seq', 'cornell_movie', 'vanilla' ,1.787 ,2.978 ,4.816 ,0.004 ,0.018 ,0.036 ,2.648 ],
['seq2seq', 'cornell_movie', 'swapping' ,2.347 ,4.274 ,9.517 ,0.004 ,0.018 ,0.093 ,3.034 ],
['seq2seq', 'cornell_movie', 'idf' ,2.184 ,3.841 ,6.990 ,0.006 ,0.021 ,0.084 ,3.517 ],
['seq2seq', 'cornell_movie', 'face' , 3.436 , 5.857 , 26.109 , 0.008 , 0.046 , 0.595 , 5.028 ],
['transformer', 'personachat', 'vanilla', 2.198 ,4.799 ,9.758 , 0.009 , 0.032 ,0.241 ,3.807 ],
['transformer', 'personachat', 'swapping' ,2.525 ,4.851 ,11.878 ,0.008 ,0.027 ,0.202 ,3.998 ],
['transformer', 'personachat', 'idf' , 2.576 , 5.027 , 12.383 ,0.009 ,0.027 , 0.251 , 4.041 ],
['transformer', 'dailydialog', 'vanilla', 2.449 ,5.020 ,6.935 ,0.027 ,0.101 ,0.277 ,4.348 ],
['transformer', 'dailydialog', 'swapping' , 3.081 ,5.799 ,10.848 , 0.029 , 0.119 , 0.445 , 5.286 ],
['transformer', 'dailydialog', 'idf' ,3.034 , 5.803 , 11.025 ,0.026 ,0.109 ,0.442 ,5.266 ],
['transformer', 'empathetic_dialogues', 'vanilla',2.346 ,4.663 , 9.957 ,0.011 ,0.043 ,0.323 ,3.957 ],
['transformer', 'empathetic_dialogues', 'swapping' , 2.812 , 5.289 ,9.498 , 0.020 , 0.075 , 0.410 , 4.429 ],
['transformer', 'empathetic_dialogues', 'idf' ,2.775 ,5.077 ,9.387 ,0.016 ,0.057 ,0.296 ,4.189 ],
['transformer', 'cornell_movie', 'vanilla', 1.908 ,3.293 ,4.678 ,0.006 ,0.019 ,0.034 ,2.779 ],
['transformer', 'cornell_movie', 'swapping', 2.435 ,4.120 ,6.898 ,0.006 ,0.016 ,0.060 ,3.750 ],
['transformer', 'cornell_movie', 'idf', 2.763 , 4.579 , 10.196 , 0.008 , 0.023 , 0.148 , 4.165 ]],
columns = ['model', 'dataset', 'method', 'avg_idf', 'max_idf', 'length', 'd1', 'd2', 'dN', 'entropy'])
# +
fig, ax = plt.subplots(1, 3, figsize=(15, 4))
ylabels = []
for (model, dataset), setting_df in duel_untied_df.groupby(['model', 'dataset']):
c_best = Counter(setting_df['best_overall'])
c_int = Counter(setting_df['most_int'][~setting_df['none_int']])
c_rel = Counter(setting_df['most_rel'][~setting_df['none_rel']])
d = float(sum(c_int.values()))
df = auto_metrics_df[(auto_metrics_df['dataset'] == dataset) & (auto_metrics_df['model'] == model)]
for method, marker in [('vanilla', 'ko'), ('idf', 'ro')]:
row = df[df['method'] == method]
ax[1].plot(row['d1'].values[0], 100.*c_int['vanilla']/d, marker)
# ax[1].plot(row['d2'].values[0], 100.*c_int['idf']/d, marker)
ax[2].plot(row['entropy'].values[0], 100.*c_int['idf']/d, marker)
ax[0].plot(auto_metrics_df['d1'], auto_metrics_df['d2'], 'o')
ax[0].set_ylabel('d2 of a dataset-model setting')
ax[0].set_xlabel('d1 of a dataset-model setting')
ax[1].set_ylabel('Percent won in duel')
ax[2].set_ylabel('Percent won in duel')
ax[1].set_xlabel('d-1')
# ax[1].set_xlabel('d-2')
ax[2].set_xlabel('unigram entropy')
plt.suptitle('Each point is a dataset-model setting (black=vanilla, red=idf)')
plt.show()
# -
plt.plot(auto_metrics_df['d1'], auto_metrics_df['entropy'], 'o')
plt.ylabel('unigram entropy of dataset-model setting')
plt.xlabel('d1 of a dataset-model setting')
# plt.plot(auto_metrics_df['d1'], auto_metrics_df['max_idf'], 'o')
plt.show()
# +
from scipy.stats import spearmanr
auto_metrics_df.columns
# +
# https://matplotlib.org/tutorials/colors/colormaps.html
auto_metrics = np.array([ 'max_idf', 'length', 'd1', 'd2', 'dN', 'entropy'])
auto_metric_names = np.array(['$idf_{max}$', 'len.', 'd-1', 'd-2', 'd-N', 'ent.'])
mat = np.NaN * np.zeros((len(auto_metrics)+2, len(auto_metrics)))
for j, metricj in enumerate(auto_metrics):
for i, metrici in enumerate(auto_metrics):
if i > j:
mat[i, j] = spearmanr(auto_metrics_df[metrici], auto_metrics_df[metricj])[0]
autos = {x:[] for x in auto_metrics}
humans_int = []
humans_rel = []
fig, ax = plt.subplots(figsize=(5,4))
for (model, dataset), setting_df in duel_untied_df.groupby(['model', 'dataset']):
c_best = Counter(setting_df['best_overall'])
c_int = Counter(setting_df['most_int'][~setting_df['none_int']])
c_rel = Counter(setting_df['most_rel'][~setting_df['none_rel']])
d = float(sum(c_int.values()))
humans_int.append(100.*c_int['vanilla']/d)
humans_rel.append(100.*c_rel['vanilla']/d)
df = auto_metrics_df[(auto_metrics_df['dataset'] == dataset) & (auto_metrics_df['model'] == model)]
for i, metrici in enumerate(auto_metrics):
row = df[df['method'] == 'vanilla']
autos[metrici].append(row[metrici].values[0])
for j, metricj in enumerate(auto_metrics):
mat[-2, j] = spearmanr(autos[metricj], humans_int)[0]
for j, metricj in enumerate(auto_metrics):
mat[-1, j] = spearmanr(autos[metricj], humans_rel)[0]
plt.pcolor(mat[::-1,:], vmin=-1, vmax=1, cmap='RdBu')
plt.xticks(.5 + np.arange(auto_metric_names[:].shape[0]), auto_metric_names[:], fontsize=12)#, rotation=90)
extended_metrics = np.concatenate([auto_metric_names, ['% int', '% rel']])
plt.yticks(.5 + np.arange(auto_metrics.shape[0]+2), extended_metrics[::-1], fontsize=16)
cbar = plt.colorbar()
cbar.ax.tick_params(labelsize=14)
cbar.set_label('Spearman correlation coeff.', size=16)
plt.show()
plt.pcolor(mat[::-1,:], vmin=-1, vmax=1, cmap='RdBu')
plt.xticks(.5 + np.arange(auto_metric_names[:].shape[0]), auto_metric_names[:], fontsize=12)#, rotation=90)
extended_metrics = np.concatenate([auto_metric_names, ['% int', '% rel']])
plt.yticks(.5 + np.arange(auto_metrics.shape[0]+2), extended_metrics[::-1], fontsize=16)
cbar = plt.colorbar()
cbar.ax.tick_params(labelsize=14)
cbar.set_label('Spearman correlation coeff.', size=16)
plt.ylim([2, len(auto_metrics)+1])
plt.xlim([0, len(auto_metrics)-1])
plt.show()
# -
mat
auto_metrics_df[:4]
# +
# https://matplotlib.org/tutorials/colors/colormaps.html
auto_metrics = np.array([ 'max_idf', 'length', 'd1', 'd2', 'dN', 'entropy'])
auto_metric_names = np.array(['$idf_{max}$', 'len.', 'd-1', 'd-2', 'd-N', 'ent.'])
diff_auto_metric_names = np.array(['$\Delta_{idf_{max}}$', '$\Delta_{len.}$', '$\Delta_{d-1}$',
'$\Delta_{d-2}$', '$\Delta_{d-N}$', '$\Delta_{ent.}$'])
diff_auto_metrics_df = []
for (model, dataset), df in auto_metrics_df.groupby(['model', 'dataset']):
idf_row = df[df['method'] == 'idf']
vanilla_row = df[df['method'] == 'vanilla']
diff_auto_metrics_df.append([model, dataset] + [vanilla_row[m].values[0] - idf_row[m].values[0] for m in auto_metrics])
col_names = ['model', 'dataset'] + list(auto_metrics)
diff_auto_metrics_df = pd.DataFrame(diff_auto_metrics_df, columns=col_names)
mat = np.NaN * np.zeros((len(auto_metrics)+2, len(auto_metrics)+1))
for j, metricj in enumerate(auto_metrics):
for i, metrici in enumerate(auto_metrics):
if i > j:
mat[i, j] = spearmanr(diff_auto_metrics_df[metrici], diff_auto_metrics_df[metricj])[0]
autos = {x:[] for x in auto_metrics}
humans_int = []
humans_rel = []
fig, ax = plt.subplots(figsize=(5,4))
for (model, dataset), setting_df in duel_untied_df.groupby(['model', 'dataset']):
c_best = Counter(setting_df['best_overall'])
c_int = Counter(setting_df['most_int'][~setting_df['none_int']])
c_rel = Counter(setting_df['most_rel'][~setting_df['none_rel']])
d = float(sum(c_int.values()))
humans_int.append(100.*c_int['vanilla']/d)
humans_rel.append(100.*c_rel['vanilla']/d)
df = diff_auto_metrics_df[(diff_auto_metrics_df['dataset'] == dataset) & (diff_auto_metrics_df['model'] == model)]
for i, metrici in enumerate(auto_metrics):
# row = df[df['method'] == 'vanilla']
# autos[metrici].append(row[metrici].values[0])
autos[metrici].append(df[metrici].values[0])
for j, metricj in enumerate(auto_metrics):
mat[-2, j] = spearmanr(autos[metricj], humans_int)[0]
for j, metricj in enumerate(auto_metrics):
mat[-1, j] = spearmanr(autos[metricj], humans_rel)[0]
mat[-1, -1] = spearmanr(humans_int, humans_rel)[0]
plt.pcolor(mat[::-1,:], vmin=-1, vmax=1, cmap='PRGn')
plt.xticks(.5 + np.arange(auto_metric_names[:].shape[0]+1), list(diff_auto_metric_names[:])+['% int',], fontsize=13)#, rotation=90)
extended_metrics = np.concatenate([diff_auto_metric_names, ['% int', '% rel']])
plt.yticks(.5 + np.arange(auto_metrics.shape[0]+2), extended_metrics[::-1], fontsize=16)
cbar = plt.colorbar()
cbar.ax.tick_params(labelsize=14)
cbar.set_label('Spearman correlation coeff.', size=16)
plt.ylim([0, len(auto_metrics)+1])
plt.show()
plt.subplots(figsize=(2, 4))
plt.pcolor(mat[:-3:-1,:].T, vmin=-1, vmax=1, cmap='RdBu')
plt.yticks(.5 + np.arange(auto_metric_names[:].shape[0]+1), list(diff_auto_metric_names[:])+['$\Delta_{int}$',], fontsize=20)#, rotation=90)
extended_metrics = np.concatenate([diff_auto_metric_names, ['$\Delta_{int}$', '$\Delta_{rel}$']])
plt.xticks(.5 + np.arange(auto_metrics.shape[0]+2), extended_metrics[::-1], fontsize=18)
cbar = plt.colorbar()
cbar.ax.tick_params(labelsize=14)
cbar.set_label('Spearman correlation coeff.', size=16)
plt.xlim([0, 2])
plt.show()
plt.subplots(figsize=(8, 2))
plt.pcolor(mat[:-3:-1,:], vmin=-1, vmax=1, cmap='RdBu')
plt.xticks(.5 + np.arange(auto_metric_names[:].shape[0]+1), list(diff_auto_metric_names[:])+['$\Delta_{int}$',], fontsize=20)#, rotation=90)
extended_metrics = np.concatenate([diff_auto_metric_names, ['$\Delta_{int}$', '$\Delta_{rel}$']])
plt.yticks(.5 + np.arange(auto_metrics.shape[0]+2), extended_metrics[::-1], fontsize=20)
cbar = plt.colorbar()
cbar.ax.tick_params(labelsize=14)
cbar.set_label('Spearman \n corr. coeff.', size=16)
plt.ylim([0, 2])
plt.show()
# -
len(humans_int)
duel_untied_df[duel_untied_df['response_vanilla'] == duel_untied_df['response_idf']]
vanilla_resps = duel_untied_df[duel_untied_df['dataset'] == "personachat"]["response_vanilla"]
idf_resps = duel_untied_df[duel_untied_df['dataset'] == "personachat"]["response_idf"]
# +
count = Counter(vanilla_resps)
# -
count.most_common()
idf_count = Counter(idf_resps)
idf_count.most_common()
duel_untied_df[duel_untied_df['best_overall'] == duel_untied_df['most_int'] & duel_untied_df['best_overall'] == duel_untied_df['most_int']]
[*raw_duel_df.columns]
with open("./just_with_id.csv", "w") as f:
f.write(raw_duel_df[['Input.msg1_9', 'Input.msg2_9', 'Input.resp_1_9', 'Input.resp_2_9', 'Answer.best_9', 'Answer.Justified Answer Final', 'WorkerId']].to_csv())
[*raw_duel_df.columns]
with open("./just_warmup_with_id.csv", "w") as f:
f.write(raw_duel_df[['Answer.best_warmup', 'Answer.Justified Answer', 'WorkerId']].to_csv())
inputs = []
import csv
with open("../../../face_turker_TEMP.csv", "r") as f:
csvreader = csv.reader(f, delimiter=',')
count = 0
for row in csvreader:
if count == 0:
count += 1
continue
for i in range(10):
inputs.append([row[i * 8], row[i * 8 + 1], row[i * 8 + 2], row[i * 8 + 3], row[i * 8 + 4], row[i * 8 + 5], row[i * 8 + 6], row[i * 8 + 7]])
inputs
not_in = []
old_conts = [*duel_df["context"].unique()]
stand_conts = []
for i in old_conts:
stand_conts.append(standardize_profanity(i))
for inp in inputs:
if standardize_profanity(inp[0] + inp[1]) not in stand_conts:
not_in.append(inp)
print(len(not_in))
duel_df.columns
duel_df[duel_df["response_vanilla"] == duel_df["response_idf"]]["dataset"]
len(not_in)
rel_inps = []
duel_inps = []
for inp in new_ones:
if "transformer" in inp[3]:
found = False
for i in not_in:
if "seq2seq" in i[3] and standardize_profanity(i[0]) == standardize_profanity(inp[0]) and standardize_profanity(i[1]) == standardize_profanity(inp[1]):
rel_inps.append(i)
duel_inps.append(i)
found = True
if not found:
print(inp)
duel_inps.append(inp)
else:
rel_inps.append(inp)
duel_inps.append(inp)
found = False
for i in not_in:
if "transformer" in i[3] and standardize_profanity(i[0]) == standardize_profanity(inp[0]) and standardize_profanity(i[1]) == standardize_profanity(inp[1]):
duel_inps.append(i)
found = True
if not found:
print(inp)
print(len(new_ones))
print("SHOULD BE 16 " + str(len(rel_inps)))
print("SHOULD BE 32 " + str(len(duel_inps)))
not_in
len(rel_inps)
len(duel_inps)
import re
def standardize_profanity(profane_str):
return re.sub("([#|\$|%|!|&|@])+([#|\$|%|!|&|@]).*([#|\$|%|!|&|@])+", "****", profane_str)
# +
# We need one empathetic, one personachat, and 14 cornell
new_ones = []
new_contexts = []
persona_counter = 0
emp_counter = 0
cornell_counter = 0
need_persona = 1
need_emp = 1
need_cornell = 14
for inp in not_in:
if "personachat" in inp[3] and persona_counter < need_persona and standardize_profanity(inp[0]) + " BREAK " + standardize_profanity(inp[1]) not in new_contexts:
new_ones.append(inp)
new_contexts.append(standardize_profanity(inp[0]) + " BREAK " + standardize_profanity(inp[1]))
persona_counter += 1
if "empathetic_dialogues" in inp[3] and emp_counter < need_emp and standardize_profanity(inp[0]) + " BREAK " + standardize_profanity(inp[1]) not in new_contexts:
new_ones.append(inp)
new_contexts.append(standardize_profanity(inp[0]) + " BREAK " + standardize_profanity(inp[1]))
emp_counter += 1
if "cornell_movie" in inp[3] and cornell_counter < need_cornell and standardize_profanity(inp[0]) + " BREAK " + standardize_profanity(inp[1]) not in new_contexts:
new_ones.append(inp)
new_contexts.append(standardize_profanity(inp[0]) + " BREAK " + standardize_profanity(inp[1]))
cornell_counter += 1
print(len(new_ones))
# -
new_ones
for inp in new_ones:
if inp[0] + inp[1] in duel_df["context"].unique():
print("ERROR")
duel_df["context"]
# +
from profanity import profanity
face_inputs = []
with open("../../../tempo/tmp_v5/tmp/personachat/face_minfreq_2_test.out", "r") as f:
data_name = "personachat seq2seq face"
new_data_name = "[personachat]:"
found_new = False
eoc = False
local_input = []
response = ""
counter = 0
examplenum = 0
for line in f.readlines():
#print(line)
if new_data_name in line:
found_new = True
if "[FACE]" in line:
#print("FOUND")
#print(examplenum)
response += line.replace("[FACE]:", "")
input1 = ""
input2 = ""
if len(local_input) > 1:
input1 = local_input[-2].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
input2 = local_input[-1].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
else:
input1 = "--"
input2 = local_input[-1].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
if input1 + " BREAK " + input2 not in new_contexts:
response = ""
local_input = []
examplenum += 1
input1 = ""
input2 = ""
continue
#else:
# import pdb; pdb.set_trace()
face_inputs.append([profanity.censor(input1), profanity.censor(input2), profanity.censor(response).replace("person2","").replace("\n","").replace("\t","").replace(",", u"\u002C").replace("__start__","").replace(data_name, "").replace("__unk__", "").replace("]","").replace("__SILENCE__", "").replace("[personachat:","").lower().strip(), data_name])
if len(face_inputs) > 99:
break
response = ""
local_input = []
examplenum += 1
input1 = ""
input2 = ""
if "eval_labels" in line or "[situation" in line or "[topic" in line or "[emotion" in line or "[prepend_ctx" in line or "elapsed: {'exs':" in line or "label_candidates" in line or "your persona:" in line or "[eval_labels:" in line or "eval_labels_choice" in line or "deepmoji_cand" in line or "emotion:" in line or "act_type" in line or "prepend_cand" in line or "deepmoji_ctx" in line:
continue
#if found_new:
local_input.append(line)
counter += 1
# -
len(face_inputs)
# +
from profanity import profanity
with open("../../../tempo/tmp_v5/tmp/empathetic_dialogues/face_minfreq_2_test.out", "r") as f:
data_name = "empathetic_dialogues seq2seq face"
new_data_name = "[empathetic_dialogues]:"
found_new = False
eoc = False
local_input = []
response = ""
counter = 0
examplenum = 0
for line in f.readlines():
#print(line)
if new_data_name in line:
found_new = True
if "[FACE]" in line:
#print("FOUND")
#print(examplenum)
response += line.replace("[FACE]:", "")
input1 = ""
input2 = ""
if len(local_input) > 1:
input1 = local_input[-2].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
input2 = local_input[-1].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
else:
input1 = "--"
input2 = local_input[-1].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
if input1 + " BREAK " + input2 not in new_contexts:
response = ""
local_input = []
examplenum += 1
input1 = ""
input2 = ""
continue
#else:
# import pdb; pdb.set_trace()
face_inputs.append([profanity.censor(input1), profanity.censor(input2), profanity.censor(response).replace("person2","").replace("\n","").replace("\t","").replace(",", u"\u002C").replace("__start__","").replace(data_name, "").replace("__unk__", "").replace("]","").replace("__SILENCE__", "").replace("[personachat:","").lower().strip(), data_name])
if len(face_inputs) > 99:
break
response = ""
local_input = []
examplenum += 1
input1 = ""
input2 = ""
if "eval_labels" in line or "[situation" in line or "[topic" in line or "[emotion" in line or "[prepend_ctx" in line or "elapsed: {'exs':" in line or "label_candidates" in line or "your persona:" in line or "[eval_labels:" in line or "eval_labels_choice" in line or "deepmoji_cand" in line or "emotion:" in line or "act_type" in line or "prepend_cand" in line or "deepmoji_ctx" in line:
continue
#if found_new:
local_input.append(line)
counter += 1
# -
len(face_inputs)
# +
from profanity import profanity
with open("../../../tempo/tmp_v5/tmp/cornell_movie/face_minfreq_2_test.out", "r") as f:
data_name = "cornell_movie seq2seq face"
new_data_name = "[cornell_movie]:"
found_new = False
eoc = False
local_input = []
response = ""
counter = 0
examplenum = 0
for line in f.readlines():
#print(line)
if new_data_name in line:
found_new = True
if "[FACE]" in line:
#print("FOUND")
#print(examplenum)
response += line.replace("[FACE]:", "")
input1 = ""
input2 = ""
#print(local_input)
if len(local_input) > 1:
input1 = local_input[-2].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
input2 = local_input[-1].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
else:
input1 = "~~"
input2 = local_input[-1].replace(new_data_name, "").replace("\n","").replace(",", u"\u002C").replace("__start__","").replace("__unk__", "").replace("\t","").replace("__SILENCE__", "").replace("]","").lower().strip()
if standardize_profanity(profanity.censor(input1)) + " BREAK " + standardize_profanity(profanity.censor(input2)) not in new_contexts:
response = ""
local_input = []
examplenum += 1
#print(input1 + " BREAK " + input2)
input1 = ""
input2 = ""
continue
#else:
# import pdb; pdb.set_trace()
face_inputs.append([profanity.censor(input1), profanity.censor(input2), profanity.censor(response).replace("person2","").replace("\n","").replace("\t","").replace(",", u"\u002C").replace("__start__","").replace(data_name, "").replace("__unk__", "").replace("]","").replace("__SILENCE__", "").replace("[personachat:","").lower().strip(), data_name])
if len(face_inputs) > 99:
break
response = ""
local_input = []
examplenum += 1
input1 = ""
input2 = ""
if "eval_labels" in line or "[situation" in line or "[topic" in line or "[emotion" in line or "[prepend_ctx" in line or "elapsed: {'exs':" in line or "label_candidates" in line or "your persona:" in line or "[eval_labels:" in line or "eval_labels_choice" in line or "deepmoji_cand" in line or "emotion:" in line or "act_type" in line or "prepend_cand" in line or "deepmoji_ctx" in line:
continue
#if found_new:
local_input.append(line)
counter += 1
# -
len(face_inputs)
len(new_contexts)
new_ones
final_rel_inps = []
for inp in rel_inps:
final_rel_inps.append([inp[0], inp[1], inp[2], inp[3]])
final_rel_inps.append([inp[0], inp[1], inp[4], inp[5]])
final_rel_inps.append([inp[0], inp[1], inp[6], inp[7]])
# +
final_master_inputs = final_rel_inps + face_inputs
import random
import csv
random.shuffle(final_master_inputs)
print(len(final_master_inputs))
warmup_questions = ["Cool is football", "That's cool!", "Yes, I played tennis last week! It was so much fun.", "Yes, football football football football football."]
with open('relevancy_turker_input_rerun_1.csv', 'w') as csvfile:
spamwriter = csv.writer(csvfile, delimiter=',')
with open('relevancy_turker_input_rerun_2.csv', 'w') as csvfile2:
spamwriter2 = csv.writer(csvfile2, delimiter=',')
msg_colnames = ["warmup_response"]
for x in range(10):
msg_colnames += ['msg1_%s' % (x), 'msg2_%s' % (x), 'resp_1_%s' % (x), 'model_1_%s' % (x)]
spamwriter.writerow(msg_colnames)
msg_colnames = ["warmup_response"]
for x in range(4):
msg_colnames += ['msg1_%s' % (x), 'msg2_%s' % (x), 'resp_1_%s' % (x), 'model_1_%s' % (x)]
spamwriter2.writerow(msg_colnames)
for i in range(int(len(final_master_inputs) / 10)):
row = [random.choice(warmup_questions)]
for j in range(10):
row.extend([final_master_inputs[i * 10 + j][0], final_master_inputs[i * 10 + j][1], final_master_inputs[i * 10 + j][2], final_master_inputs[i * 10 + j][3]])
spamwriter.writerow(row)
print(i)
row = [random.choice(warmup_questions)]
print("here")
i = 6
for j in range(4):
row.extend([final_master_inputs[i * 10 + j][0], final_master_inputs[i * 10 + j][1], final_master_inputs[i * 10 + j][2], final_master_inputs[i * 10 + j][3]])
spamwriter2.writerow(row)
# -
inputs = []
messages = []
duel_one = []
import random
random.seed(22)
for i in range(10):
msg1 = "msg1_" + str(i)
msg2 = "msg2_" + str(i)
resp1 = "resp_1_" + str(i)
model1 = "model_1_"+ str(i)
resp2 = "resp_2_" + str(i)
model2 = "model_2_"+ str(i)
resp3 = "resp_3_" + str(i)
model3 = "model_3_"+ str(i)
for j in range(len(raw_df)):
if "swapping" in raw_df[model1][j]:
if random.random() > 0.5:
duel_one.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp2][j], raw_df[model2][j], raw_df[resp3][j], raw_df[model3][j]])
else:
duel_one.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp3][j], raw_df[model3][j], raw_df[resp2][j], raw_df[model2][j]])
if "swapping" in raw_df[model2][j]:
if random.random() > 0.5:
duel_one.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp1][j], raw_df[model1][j], raw_df[resp3][j], raw_df[model3][j]])
else:
duel_one.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp3][j], raw_df[model3][j], raw_df[resp1][j], raw_df[model1][j]])
if "swapping" in raw_df[model3][j]:
if random.random() > 0.5:
duel_one.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp2][j], raw_df[model2][j], raw_df[resp1][j], raw_df[model1][j]])
else:
duel_one.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp1][j], raw_df[model1][j], raw_df[resp2][j], raw_df[model2][j]])
inputs.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp1][j], raw_df[model1][j]])
inputs.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp2][j], raw_df[model2][j]])
inputs.append([raw_df[msg1][j],raw_df[msg2][j], raw_df[resp3][j], raw_df[model3][j]])
#print(raw_df[msg][j])
messages.append(str(raw_df[msg1][j]) + " BREAK " + str(raw_df[msg2][j]))
# +
random.shuffle(final_duel_inps)
with open('duel_one_turker_input_rerun_1.csv', 'w') as csvfile:
with open('duel_one_turker_input_rerun_2.csv', 'w') as csvfile2:
spamwriter = csv.writer(csvfile, delimiter=',')
spamwriter2 = csv.writer(csvfile2, delimiter=',')
msg_colnames = []
for x in range(10):
msg_colnames += ['msg1_%s' % (x), 'msg2_%s' % (x), 'resp_1_%s' % (x), 'model_1_%s' % (x), 'resp_2_%s' % (x), 'model_2_%s' % (x)]
spamwriter.writerow(msg_colnames)
msg_colnames = []
for x in range(2):
msg_colnames += ['msg1_%s' % (x), 'msg2_%s' % (x), 'resp_1_%s' % (x), 'model_1_%s' % (x), 'resp_2_%s' % (x), 'model_2_%s' % (x)]
spamwriter2.writerow(msg_colnames)
for i in range(int(len(final_duel_inps) / 10)):
row = []
for j in range(10):
row.extend(final_duel_inps[i * 10 + j])
spamwriter.writerow(row)
i = 3
row = []
for j in range(2):
row.extend(final_duel_inps[i * 10 + j])
spamwriter2.writerow(row)
# -
final_duel_inps = []
already = []
import random
random.seed(22)
for i in duel_inps:
if "swapping" in i[3]:
if random.random() > 0.5:
final_duel_inps.append([i[0], i[1], i[4], i[5], i[6], i[7]])
else:
final_duel_inps.append([i[0], i[1], i[6], i[7], i[4], i[5]])
if "swapping" in i[5]:
if random.random() > 0.5:
final_duel_inps.append([i[0], i[1], i[2], i[3], i[6], i[7]])
else:
final_duel_inps.append([i[0], i[1], i[6], i[7], i[2], i[3]])
if "swapping" in i[7]:
if random.random() > 0.5:
final_duel_inps.append([i[0], i[1], i[4], i[5], i[2], i[3]])
else:
final_duel_inps.append([i[0], i[1], i[2], i[3], i[4], i[5]])
len(final_duel_inps)
final_master_inputs[-1]
duel_inps
rel_and_int = ~(duel_untied_df['none_rel'] | duel_untied_df['none_int'])
a = duel_untied_df['most_rel'] != duel_untied_df['most_int']
b = duel_untied_df['most_rel'] == duel_untied_df['best_overall']
with open("int_not_chosen.csv", "w") as f:
f.write(duel_untied_df[['context', 'response_vanilla', 'response_idf']].to_csv())
t3.columns
duel_df["model"]
fleiss_kappa
import statsmodels.stats.inter_rater as ir
kappa_dict = {}
kappa_table = []
for index, row in duel_df.iterrows():
if row["context"] + row["model"] not in kappa_dict:
kappa_dict[row["context"] + row["model"]] = [0,0]
if row["most_int"] == "vanilla":
kappa_dict[row["context"] + row["model"]][0] += 1
else:
kappa_dict[row["context"] + row["model"]][1] += 1
for key in kappa_dict.keys():
kappa_table.append(kappa_dict[key])
kappa_table = np.array(kappa_table)
ir.fleiss_kappa(kappa_table)
kappa_dict = {}
kappa_table = []
for index, row in duel_df.iterrows():
if row["context"] + row["model"] not in kappa_dict:
kappa_dict[row["context"] + row["model"]] = [0,0]
if row["most_rel"] == "vanilla":
kappa_dict[row["context"] + row["model"]][0] += 1
else:
kappa_dict[row["context"] + row["model"]][1] += 1
for key in kappa_dict.keys():
kappa_table.append(kappa_dict[key])
kappa_table = np.array(kappa_table)
ir.fleiss_kappa(kappa_table)
kappa_dict = {}
kappa_table = []
for index, row in duel_df.iterrows():
if row["context"] + row["model"] not in kappa_dict:
kappa_dict[row["context"] + row["model"]] = [0,0]
if row["best_overall"] == "vanilla":
kappa_dict[row["context"] + row["model"]][0] += 1
else:
kappa_dict[row["context"] + row["model"]][1] += 1
for key in kappa_dict.keys():
kappa_table.append(kappa_dict[key])
kappa_table = np.array(kappa_table)
ir.fleiss_kappa(kappa_table)
from sklearn.metrics import cohen_kappa_score
kappa_dict = {}
kappa_table = []
for index, row in duel_df.iterrows():
if row["context"] + row["model"] not in kappa_dict:
kappa_dict[row["context"] + row["model"]] = [-1,-1,-1]
if kappa_dict[row["context"] + row["model"]][0] == -1:
if row["best_overall"] == "vanilla":
kappa_dict[row["context"] + row["model"]][0] += 1
else:
kappa_dict[row["context"] + row["model"]][0] += 2
elif kappa_dict[row["context"] + row["model"]][1] == -1:
if row["best_overall"] == "vanilla":
kappa_dict[row["context"] + row["model"]][1] += 1
else:
kappa_dict[row["context"] + row["model"]][1] += 2
elif kappa_dict[row["context"] + row["model"]][2] == -1:
if row["best_overall"] == "vanilla":
kappa_dict[row["context"] + row["model"]][2] += 1
else:
kappa_dict[row["context"] + row["model"]][2] += 2
for key in kappa_dict.keys():
random.shuffle(kappa_dict[key])
kappa_table.append(kappa_dict[key])
kappa_table = np.array(kappa_table)
cohen_kappa_score(kappa_table[:,0], kappa_table[:,1])
from sklearn.metrics import cohen_kappa_score
kappa_dict = {}
kappa_table = []
for index, row in duel_df.iterrows():
if row["context"] + row["model"] not in kappa_dict:
kappa_dict[row["context"] + row["model"]] = [-1,-1,-1]
if kappa_dict[row["context"] + row["model"]][0] == -1:
if row["most_int"] == "vanilla":
kappa_dict[row["context"] + row["model"]][0] += 1
else:
kappa_dict[row["context"] + row["model"]][0] += 2
elif kappa_dict[row["context"] + row["model"]][1] == -1:
if row["most_int"] == "vanilla":
kappa_dict[row["context"] + row["model"]][1] += 1
else:
kappa_dict[row["context"] + row["model"]][1] += 2
elif kappa_dict[row["context"] + row["model"]][2] == -1:
if row["most_int"] == "vanilla":
kappa_dict[row["context"] + row["model"]][2] += 1
else:
kappa_dict[row["context"] + row["model"]][2] += 2
for key in kappa_dict.keys():
random.shuffle(kappa_dict[key])
kappa_table.append(kappa_dict[key])
kappa_table = np.array(kappa_table)
cohen_kappa_score(kappa_table[:,0], kappa_table[:,1])
from sklearn.metrics import cohen_kappa_score
kappa_dict = {}
kappa_table = []
for index, row in duel_df.iterrows():
if row["context"] + row["model"] not in kappa_dict:
kappa_dict[row["context"] + row["model"]] = [-1,-1,-1]
if kappa_dict[row["context"] + row["model"]][0] == -1:
if row["most_rel"] == "vanilla":
kappa_dict[row["context"] + row["model"]][0] += 2
else:
kappa_dict[row["context"] + row["model"]][0] += 3
elif kappa_dict[row["context"] + row["model"]][1] == -1:
if row["most_rel"] == "vanilla":
kappa_dict[row["context"] + row["model"]][1] += 2
else:
kappa_dict[row["context"] + row["model"]][1] += 3
elif kappa_dict[row["context"] + row["model"]][2] == -1:
if row["most_rel"] == "vanilla":
kappa_dict[row["context"] + row["model"]][2] += 2
else:
kappa_dict[row["context"] + row["model"]][2] += 3
avg = 0
for key in kappa_dict.keys():
avg += len(set(kappa_dict[key]))
random.shuffle(kappa_dict[key])
kappa_table.append(kappa_dict[key])
kappa_table = np.array(kappa_table)
print(cohen_kappa_score(kappa_table[:,0], kappa_table[:,1]))
print(avg / len(kappa_dict.keys()))
# ### Katie's new data for rerun of best
katies_raw_duel_df = pd.concat([pd.read_csv('../batches/Batch_3630895_batch_results.csv'),
pd.read_csv('../batches/Batch_3636922_batch_results.csv'),
pd.read_csv('../batches/Batch_3638556_batch_results.csv'),
pd.read_csv('../batches/Batch_3638681_batch_results.csv'),
pd.read_csv('../batches/Batch_3651082_batch_results.csv'),
pd.read_csv('../batches/Batch_3650478_batch_results.csv'),
pd.read_csv('../batches/Batch_3650472_batch_results.csv')
], axis=0, sort=False).reset_index()
# +
dueling_methods = ['vanilla', 'idf']
katies_duel_df = []
for i in katies_raw_duel_df.index:
row = katies_raw_duel_df.loc[i]
hit_num = i
worker_id = row['WorkerId']
for ex in range(10): # given to each turker in a single HIT
if np.isnan(row['Answer.best_%s' % ex]):
# This may be null from the re-run subset, so skip it (and count after!)
continue
option_to_method = {}
method_to_resp = {}
for option in range(1, 2+1):
model_pieces = row['Input.model_%s_%s' % (option, ex)].split(' ')
dataset, model, method = model_pieces
option_to_method[option] = method
method_to_resp[method] = row['Input.resp_%s_%s' % (option, ex)]
best_overall = option_to_method[row['Answer.best_%s' % (ex,)]]
most_interesting = option_to_method[row['Answer.int_%s' % (ex,)]]
most_relevant = option_to_method[row['Answer.rel_%s' % (ex,)]]
none_int = type(row['Answer.check_int_%s'% (ex,)]) == str
none_rel = type(row['Answer.check_rel_%s'% (ex,)]) == str
message1 = row['Input.msg1_%s' % (ex,)]
message2 = row['Input.msg2_%s' % (ex,)]
katies_duel_df.append([hit_num, worker_id, dataset, model,
message1, message2] + \
[method_to_resp[x] for x in dueling_methods] + \
[best_overall, most_interesting, most_relevant, none_int, none_rel])
katies_duel_df = pd.DataFrame(katies_duel_df, columns = ['hit_num', 'worker_id', 'dataset', \
'model', 'message1', 'message2',] + \
['response_%s' % x for x in dueling_methods] + \
['best_overall', 'most_int', 'most_rel', 'none_int', 'none_rel'])
katies_duel_df['context'] = katies_duel_df['message1'] + " BREAK " + katies_duel_df['message2']
# remove examples (accidentially included) where message 2 was null
katies_duel_df = katies_duel_df[[type(x) == str for x in katies_duel_df['message2']]]
# -
katies_duel_df.columns
# +
import nltk
def func(resp1, resp2):
return nltk.edit_distance(resp1, resp2) / ((len(resp1) + len(resp2)) / 2)
temp = katies_duel_df[['response_vanilla','response_idf']].apply(lambda x: func(*x), axis=1)
# +
katies_duel_untied_df = []
for (dataset, model), setting_df in katies_duel_df.groupby(['dataset', 'model']):
for context, df in setting_df.groupby('context'):
katies_duel_untied_df.append([dataset, model, context,
df['response_vanilla'].values[0], df['response_idf'].values[0],
Counter(df['best_overall'].values).most_common(1)[0][0],
Counter(df['most_rel'].values).most_common(1)[0][0],
Counter(df['most_int'].values).most_common(1)[0][0],
Counter(df['none_rel'].values).most_common(1)[0][0] == True,
Counter(df['none_int'].values).most_common(1)[0][0] == True])
katies_duel_untied_df = pd.DataFrame(katies_duel_untied_df, columns = ['dataset', 'model', 'context',
'response_vanilla', 'response_idf',
'best_overall', 'most_rel', 'most_int',
'none_rel', 'none_int'])
# -
katies_duel_untied_df.shape
temp = katies_duel_untied_df[['response_vanilla','response_idf']].apply(lambda x: func(*x), axis=1)
plt.hist(temp, bins=50)
plt.title("Edit distances for all datasets")
# plt.xlim([-1, 11])
plt.xlabel('Edit distance', fontsize=20)
plt.ylabel('Number of examples', fontsize=20)
plt.show()
# +
tasks = ['cornell_movie', 'dailydialog', 'empathetic_dialogues', 'personachat']
for task in tasks:
new_temp_df = katies_duel_df[katies_duel_df["dataset"] == task]
temp = new_temp_df[['response_vanilla','response_idf']].apply(lambda x: func(*x), axis=1)
plt.hist(temp, bins=50)
plt.title("Edit distances for %s" % task)
# plt.xlim([-1, 11])
plt.xlabel('Edit distance', fontsize=20)
plt.ylabel('Number of examples', fontsize=20)
plt.show()
# +
katies_duel_untied_df['edit_dist'] = katies_duel_untied_df[['response_vanilla','response_idf']].apply(lambda x: func(*x), axis=1)
# -
katies_duel_untied_df.columns
with open("sorted_df_s2s.csv", "w") as f:
f.write(katies_duel_untied_df[katies_duel_untied_df["model"]== "seq2seq"].sort_values(by=['edit_dist'], ascending=False).to_csv())
from statistics import median
for task in tasks:
new_temp_df = katies_duel_df[katies_duel_df["dataset"] == task]
temp = new_temp_df[['response_vanilla','response_idf']].apply(lambda x: func(*x), axis=1)
print("Median of %s is %d" %(task, median(temp)))
# +
def cutoff_func(resp1, resp2, cutoff):
return nltk.edit_distance(resp1, resp2) > cutoff
medians = [12, 28, 26, 26]
over_med = []
under_len = []
i = 0
for task in tasks:
new_temp_df = katies_duel_df[katies_duel_df["dataset"] == task]
over_med.append(new_temp_df[new_temp_df[['response_vanilla','response_idf']].apply(lambda x: cutoff_func(*x, medians[i]), axis=1)])
i += 1
over_med = pd.DataFrame(over_med, columns = ['dataset', 'model', 'context',
'response_vanilla', 'response_idf',
'best_overall', 'most_rel', 'most_int',
'none_rel', 'none_int', 'edit_dist'])
# -
over_med.shape
katies_duel_untied_df = katies_duel_untied_df.drop(columns="edit_dist")
# +
methods = ["char", "token"]
norms = ["none", "mean"]
def char_mean(resp1, resp2):
return nltk.edit_distance(resp1, resp2) / ((len(resp1) + len(resp2)) / 2)
def token_mean(resp1, resp2):
return nltk.edit_distance(resp1.split(" "), resp2.split(" ")) / ((len(resp1) + len(resp2)) / 2)
def char_none(resp1, resp2):
return nltk.edit_distance(resp1, resp2)
def token_none(resp1, resp2):
return nltk.edit_distance(resp1.split(" "), resp2.split(" "))
'''for met in methods:
for norm in norms:
method_name = met + "_" + norm
print(method_name)
katies_duel_untied_df[method_name] = katies_duel_untied_df[['response_vanilla','response_idf']].apply(lambda x: globals()[method_name](*x), axis=1)
with open("%s.csv" %(method_name), "w") as f:
f.write(katies_duel_untied_df[katies_duel_untied_df["model"]== "seq2seq"].sort_values(by=[method_name], ascending=False).to_csv())'''
# -
def ret_most_common(resp1, resp2):
#resp1 = resp1.replace("!", "").replace("?", "").replace(",","").replace(".","")
#resp2 = resp2.replace("!", "").replace("?", "").replace(",","").replace(".","")
return max(Counter(resp1.split(" ")).most_common(1)[0][1], Counter(resp2.split(" ")).most_common(1)[0][1])
# +
no_repeats = katies_duel_untied_df[['response_vanilla','response_idf']].apply(lambda x: ret_most_common(*x), axis=1) < 5
under_two = katies_duel_untied_df[['response_vanilla','response_idf']].apply(lambda x: char_none(*x), axis=1) > 2
to_run = katies_duel_untied_df[no_repeats & under_two]
# -
to_run.shape
sum(under_two)
sum(no_repeats)
to_run.columns
# +
import csv
import random
random.seed(22)
datasets = to_run["dataset"].to_list()
contexts = to_run["context"].to_list()
vanillas = to_run["response_vanilla"].to_list()
idfs = to_run["response_idf"].to_list()
models = to_run["model"].to_list()
with open('duel_one_turker_best_only.csv', 'w') as csvfile:
spamwriter = csv.writer(csvfile, delimiter=',')
msg_colnames = []
for x in range(10):
msg_colnames += ['msg1_%s' % (x), 'msg2_%s' % (x), 'resp_1_%s' % (x), 'model_1_%s' % (x), 'resp_2_%s' % (x), 'model_2_%s' % (x)]
spamwriter.writerow(msg_colnames)
inps = []
for i in range(len(to_run)):
msg1, msg2 = contexts[i].split("BREAK")
vanilla = vanillas[i ]
idf = idfs[i ]
dataset = datasets[i]
model = models[i ]
if random.random() < 0.5:
inps.append([msg1, msg2, vanilla, dataset + " " + model + " vanilla ", idf, dataset + " " + model + " idf "])
else:
inps.append([msg1, msg2, idf, dataset + " " + model + " idf ", vanilla, dataset + " " + model + " vanilla "])
random.shuffle(inps)
for i in range(int(len(to_run) / 10)):
row = []
for j in range(10):
row.extend(inps[i * 10 + j])
spamwriter.writerow(row)
# -
with open('duel_one_turker_best_only_short.csv', 'w') as csvfile:
spamwriter = csv.writer(csvfile, delimiter=',')
msg_colnames = []
for x in range(6):
msg_colnames += ['msg1_%s' % (x), 'msg2_%s' % (x), 'resp_1_%s' % (x), 'model_1_%s' % (x), 'resp_2_%s' % (x), 'model_2_%s' % (x)]
spamwriter.writerow(msg_colnames)
row = []
for i in range(6):
row.extend(inps[-i])
spamwriter.writerow(row)
# +
# Data from first weed-out best task
# -
best_raw_df = pd.concat([pd.read_csv('../batches/Batch_3652403_batch_results.csv'),
pd.read_csv('../batches/Batch_3652402_batch_results.csv'),
], axis=0, sort=False).reset_index()
# +
import re
def standardize_profanity(profane_str):
return re.sub("([#|\$|%|!|&|@])+([#|\$|%|!|&|@]).*([#|\$|%|!|&|@])+", "****", profane_str)
dueling_methods = ['vanilla', 'idf']
best_df = []
for i in best_raw_df.index:
row = best_raw_df.loc[i]
hit_num = i
worker_id = row['WorkerId']
for ex in range(10): # given to each turker in a single HIT
if np.isnan(row['Answer.best_%s' % ex]):
# This may be null from the re-run subset, so skip it (and count after!)
continue
option_to_method = {}
method_to_resp = {}
for option in range(1, 2+1):
model_pieces = row['Input.model_%s_%s' % (option, ex)].split(' ')
dataset, model, method = model_pieces
option_to_method[option] = method
method_to_resp[method] = row['Input.resp_%s_%s' % (option, ex)]
option_to_method[3] = "tie"
best_overall = option_to_method[row['Answer.best_%s' % (ex,)]]
message1 = row['Input.msg1_%s' % (ex,)]
message2 = row['Input.msg2_%s' % (ex,)]
best_df.append([hit_num, worker_id, dataset, model,
message1, message2] + \
[method_to_resp[x] for x in dueling_methods] + \
[best_overall])
best_df = pd.DataFrame(best_df, columns = ['hit_num', 'worker_id', 'dataset', \
'model', 'message1', 'message2',] + \
['response_%s' % x for x in dueling_methods] + \
['best_overall'])
best_df['context'] = best_df['message1'] + " BREAK " + best_df['message2']
# -
best_df
# +
best_untied_df = []
for_kappa = []
for_cohen = []
for (dataset, model), setting_df in best_df.groupby(['dataset', 'model']):
for context, df in setting_df.groupby('context'):
if df.shape[0] !=3:
print(dataset, model, method, df.shape)
continue
best_untied_df.append([dataset, model, context,
df['response_vanilla'].values[0], df['response_idf'].values[0],
Counter(df['best_overall'].values).most_common(1)[0][0]])
for_kappa.append([Counter(df['best_overall'].values)["vanilla"], Counter(df['best_overall'].values)["idf"], Counter(df['best_overall'].values)["tie"]])
best_untied_df = pd.DataFrame(best_untied_df, columns = ['dataset', 'model', 'context',
'response_vanilla', 'response_idf',
'best_overall'])
# -
best_untied_df.shape
# +
rates = []
for item in for_kappa:
if item[2] < 2:
rates.append(True)
else:
rates.append(False)
untied_for_rerun = best_untied_df[rates]
# -
untied_for_rerun.shape
from statsmodels.stats.inter_rater import fleiss_kappa
fleiss_kappa(for_kappa)
for_kappa
# +
from sklearn.metrics import cohen_kappa_score
tot_labs = []
for objs in for_kappa:
labs = []
for i in range(objs[0]):
labs.append(0)
for i in range(objs[1]):
labs.append(1)
for i in range(objs[2]):
labs.append(2)
tot_labs.append(labs)
r1 = [x[0] for x in tot_labs]
r2 = [x[1] for x in tot_labs]
r3 = [x[2] for x in tot_labs]
print((cohen_kappa_score(r1, r2) + cohen_kappa_score(r2, r3) + cohen_kappa_score(r1, r3)) / 3.0)
# +
tot_labs = []
for objs in for_kappa:
labs = []
for i in range(objs[0]):
labs.append(0)
for i in range(objs[1]):
labs.append(0)
for i in range(objs[2]):
labs.append(2)
tot_labs.append(labs)
r1 = [x[0] for x in tot_labs]
r2 = [x[1] for x in tot_labs]
r3 = [x[2] for x in tot_labs]
print("kappa for tie or not")
print((cohen_kappa_score(r1, r2) + cohen_kappa_score(r2, r3) + cohen_kappa_score(r1, r3)) / 3.0)
# -
| evaluation/katie_kappa_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os, re
data_path = "C:/Users/SpiffyApple/Documents/USC/OwnResearch/marketShare"
import matplotlib.pyplot as plt
# %matplotlib inline
jet = plt.get_cmap('jet')
import statsmodels.api as sm
import statsmodels.formula.api as smf
from statsmodels.sandbox.regression.predstd import wls_prediction_std
from statsmodels.regression.linear_model import WLS
import seaborn as sns
from statsmodels.stats.weightstats import ttest_ind
import itertools
# +
#convenience function
def toLatex(tmpDF, file_name, reg=False):
with open("/".join([data_path,file_name]), 'w+') as f:
if reg==False:
f.write(tmpDF.to_latex())
if reg:
f.write(tmpDF.as_latex())
return(tmpDF)
def toExcel(tmpDF, file_name):
tmpDF.to_excel("/".join([data_path,file_name]))
return(tmpDF)
# -
# # Milwaukee Tax Assessor data
#
# City-wide data were sourced from the Milwaukee city [website](https://city.milwaukee.gov/DownloadTabularData3496.htm). Two files are used, the 1) MPROP([documentation](https://itmdapps.milwaukee.gov/gis/mprop/Documentation/mprop.pdf)) and 2) MAI which contains data on the number of units, land use, residential vs commercial etc.
#
# Other potentially useful data sources can be obtained [here](https://city.milwaukee.gov/data)
#
# Information on Milwaukee residential [zoning](https://city.milwaukee.gov/ImageLibrary/Groups/ccClerk/Ordinances/Volume-2/CH295-sub5.pdf) but more succintly [here](https://city.milwaukee.gov/Designguidelines/UrbanDesignResources/Zoning-Districts.htm). For all zoning codes, come [here](https://city.milwaukee.gov/PlanningPermits/DCDzoninglink.htm)
#
# 2011-2015 5year sample ACS data for Milwaukee come from [Social Explorer](https://www.socialexplorer.com/tables/ACS2015_5yr/R11848037)
# ### Matching Assessor data to Tiger data
#
# The Assessor data make tract and block codes available but they do not make the county fips available. As a result, the tract and block number combinations are not uniquely matched to the Tiger file for Wisconsin blocks. In order to remedy this, I remove all counties that are not within Milwaukee city. Milwaukee City has 3 counties in it: Milwaukee, Washington, and Waukesha. Their respective [codes](https://www.disc.wisc.edu/Codebooks/FIPScodesWI.htm) are 079; 131; 133
tigers = pd.read_excel("/".join([data_path, 'milwaukee_blocks.xls']))
#tigers = pd.read_csv("/".join([data_path,"MilwaukeeCountyTiger.csv" ]))
tigers.loc[:,'tractblock'] = tigers.TRACTCE10.astype(str) +"."+tigers.BLOCKCE10.astype(str)
print("Number of unique tractblocks in tigers file: %d" %tigers.tractblock.unique().shape[0])
# +
# in order to improved matching between Tiger files and these data, I first exclude non-relevant counties
print("number of blocks in Wisconsin: %d" %tigers.shape[0])
print("number of blocks in relevant counties: %d" %tigers[(tigers.COUNTYFP10 == 79) | (tigers.COUNTYFP10 == 131) | (tigers.COUNTYFP10 == 133)].shape[0])
#tigers = tigers[(tigers.COUNTYFP10 == 79) | (tigers.COUNTYFP10 == 131) | (tigers.COUNTYFP10 == 133)]
print("Number of tract and block combinations in the Tiger data that are not unique: %d" %(tigers.tractblock).duplicated().sum())
# -
# # Data Processing
# +
## Load MPROP data
tax = pd.read_csv("/".join([data_path, 'mprop.csv']), low_memory=False) ## load assessor data
tax = tax.loc[:,~tax.isnull().all()] ## drop columns that are all NA
tax.drop(['TAX_DELQ','RAZE_STATUS','BI_VIOL'],axis=1, inplace=True) ## drop some of the columns with only one value
tax.columns = tax.columns.str.lower() ## make columns lower case
tax.dropna(subset=['geo_tract'],inplace=True)
tax.dropna(subset=['owner_name_1'],inplace=True)
tax.geo_tract = tax.geo_tract.astype(int)
tax.loc[:,'geo_blockgroup'] = tax.geo_block.str[:1]
tax.loc[:,'owner_name_1'] = tax.owner_name_1.str.strip()
tax.owner_name_1.replace(",","", regex=True, inplace=True)
tax.owner_name_1.replace("THE ", "", regex=True, inplace=True)
## combine the owner fields
tax.loc[:,'owner'] = (tax.owner_name_1+" "+tax.owner_name_2.astype(str)+" "+tax.owner_name_3.astype(str)).replace("nan",'',regex=True)
## LOAD MAI data - so far, there hasn't been any use for it
#mai = pd.read_csv("/".join([data_path, 'mai.csv']), low_memory=False)
#mai = mai.loc[:,~mai.isnull().all()]
#mai.columns = mai.columns.str.lower()
# +
zoningRepl = {'LB3':'local business-urban','LB2':'local business-midUrban','LB1':'local business-suburban',
'IM':'industrial mixed','C9C':'neighborhood retail','C9E':'major retail','C9F':'office and service',
'C9G':'mixed activity','RM1':'multifamily residential','RM2':'multifamily residential',
'RM3':'multifamily residential','RM4':'multifamily residential','RM5':'multifamily residential',
'RM6':'multifamily residential','RM7':'multifamily residential',
'RT1':'onetwo residential','RT2':'onetwo residential','RT3':'twofamily residential',
'RT4':'mixed residential','RO1':'residential office','RO2':'residential office',
'RS1':'singlefamily residential','RS2':'singlefamily residential','RS3':'singlefamily residential',
'RS4':'singlefamily residential','RS5':'singlefamily residential','RS6':'singlefamily residential',
'RB1':'regional business','RB2':'regional business'}
tax.zoning.replace(zoningRepl,inplace=True)
#tax.zoning.unique()
# +
print("Number of obs and proportion with unknown zoning. Num: %d, Share: %.3f" %(tax.zoning.isnull().sum(),tax.zoning.isnull().sum()/tax.shape[0]))
print("Number of units whose zoning is not known: %d" %tax.loc[tax.zoning.isnull(), 'nr_units'].sum())
tax.dropna(subset=['zoning'], inplace=True)
# -
tax.loc[:,'tractblock'] = tax.geo_tract.astype(str)+"."+tax.geo_block
print("Number of unique tract blocks before removal of non-residential properties: %d" %tax.tractblock.unique().shape[0])
tax.loc[:,'residential'] = tax.zoning.str.contains("residential") | (tax.land_use_gp <5) & (tax.land_use_gp>0) & (tax.land_use_gp.notnull())
tax.merge(tigers, on='tractblock',how='left').groupby('GEOID10').residential.sum().to_csv("/".join([data_path, 'milwaukee_props_w_commercial.csv']))
# +
## since we are not concerned with non-residential properties, I drop those here
print("All observations: %d" %tax.shape[0])
tax = tax.loc[tax.zoning.str.contains("residential") | (tax.land_use_gp <5) & (tax.land_use_gp>0) & (tax.land_use_gp.notnull())]
print("Residential observations: %d" %tax.shape[0])
# -
print("Number of units: %d" %tax.nr_units.sum())
# +
## remove properties that are owned by City of Milwaukee or its departments
#tax = tax.loc[~tax.owner_name_1.str.contains("CITY OF MILW|AUTH |AUTHORITY"),:]
# -
print("Number of unique owner addresses:")
tax.owner_mail_addr =tax.owner_mail_addr.str.replace("# \d+|#|%",'').str.replace("P O","PO").str.replace(" AV,"," AVE").str.strip()
tax.owner_mail_addr.unique().shape[0]
print("Number of unique owner The names:")
tax.owner_name_1.unique().shape[0]
# +
#tax.owner_name_1[~tax.owner_name_1.duplicated()].sort_values()
# -
ownernameRepl = {'W62N244 WASHINGTON AVE A105':'W62N244 WASHINGTON AVE','10 WHITE TRAIL LN':'10 WHITE TAIL LN',
'W7026 COTTONVILLEDR':'W7026 COTTONVILLE DR','W7026 COTTTONVILLE DR':'W7026 COTTONVILLE DR',
"DOOR4S DWELLINGS LLC":'DOORS DWELLINGS LLC'}
tax.owner_name_1.replace(ownernameRepl,inplace=True)
# # Data Descriptives
# +
## simple tabulation of owners and number of props each has
numPropsPerOwner = tax.loc[:,['owner_name_1','TAX_KEY']].groupby("owner_name_1").size()
numPropsPerOwner.name = 'numProperties'
numUnitsPerOwner = tax.loc[:,['owner_name_1','nr_units']].groupby("owner_name_1").sum()
pd.concat([numPropsPerOwner,numUnitsPerOwner],axis=1).sort_values('nr_units',ascending=False).head()
# +
#tax.loc[tax.owner_name_1.str.lower().str.contains("tarver"),['owner_name_1','owner_name_2','nr_units','own_ocpd','zoning','taxkey']]
# -
print("share of units not owner occupied: %.2f" %(tax.own_ocpd.isnull().sum()/tax.own_ocpd.shape[0]))
## share of units owner occupied and have more than one unit -- indeed
print("Share of owner-occupied properties with more than 1 unit: %.2f" %((tax.own_ocpd.notnull() & (tax.nr_units > 1)).sum()/tax.own_ocpd.shape[0]))
## per Milwaukee website, this building has 49 units; not 491
tax.nr_units.replace(491, 49, inplace=True)
tax.nr_units.replace(391,39,inplace=True)
## look at the distribution of units and distribution of units owner occupied
tax[tax.nr_units>0].nr_units.describe()
tax.loc[tax.nr_units>200,['owner','zoning','land_use_gp','house_nr_hi','street','nr_units']].head()
tax.loc[tax.nr_units> 0,['nr_units','owner_name_1','zoning']].groupby("owner_name_1").size().sort_values(ascending=False).head(n=20)
tax.nr_units[(tax.nr_units>2 ) & (tax.nr_units < 50)].hist()
plt.ylim(0,1000)
plt.title("Number of units owned by single owners")
plt.ylabel("count censored above 1,000")
print("Number of observations: %d" %tax.shape[0])
print("Number of duplicated owner names: %d" %tax.owner_name_1.duplicated().sum())
print("Number of duplicated owner addresses: %d" %tax.owner_mail_addr.duplicated().sum())
print("Number of duplicated owner addresses or names: %d" %(tax.owner_name_1.duplicated() | tax.owner_mail_addr.duplicated()).sum())
print("Smallest number of groups: %d" %(150510 - 42399))
# +
## map names to addresses assuming that first address is correct
#s = tax.drop_duplicates('owner_mail_addr').set_index("owner_mail_addr")['owner_name_1']
#tax.loc[:,'name'] = tax['owner_mail_addr'].map(s)
#tax.loc[:,'name'].unique().shape[0]
# +
## map names to addresses then addresses to names for consistency
s = tax.drop_duplicates('owner_name_1').set_index("owner_name_1")['owner_mail_addr']
tax.loc[:,'addr'] = tax['owner_name_1'].map(s)
k = tax.drop_duplicates('addr').set_index("addr")['owner_name_1']
tax.loc[:,'name'] = tax['addr'].map(k)
tax.loc[:,'addr'].unique().shape[0]
# -
# ### Method aside:
# Ideally, I would capture both duplicated addresses and duplicated names depending on which one is duplicated. The problem is that, that's hard. The corresponding [stack](https://stackoverflow.com/questions/52865020/groupby-this-or-that) solutions, one of which was implemented, don't solve the problems. They get pretty close but still 400 owners off from the ideal answer and the ideal answer isn't even ideal due to inevitable remaining errors. In fact, Abhi's answers perform better than JPPs on this dataset **but** Abhi's doesn't preserve index and is more difficult to follow so I opt for JPPs.
# +
# implementation of stack overflow answers:
### Abhi's modified answer from chat:
## map owner_mail_addresses to owner_name_1s
#mapper = tax.groupby(['owner_name_1','owner_mail_addr'],as_index=False).sum().drop_duplicates('owner_name_1',keep='first').set_index('owner_name_1').owner_mail_addr
#tax.loc[:,'name1'] = tax.owner_name_1.map(mapper)
### Abhi's initial answer:
#tax.loc[:,'group'] = tax.groupby('owner_mail_addr').grouper.group_info[0]
#d = {'owner_name_1':'first','nr_units':'sum'}
#tax1 = tax.groupby('owner_name_1',as_index=False).sum().groupby('group').agg(d)
#tax1.sort_values('owner_name_1')
#tax1
# -
# # Herfindahl–Hirschman Index(HHI)
#
# $$H = \sum_{i=1}^{N} s^2_i$$ where $i$ is a firm, $N$ is the total number of firms, and $H$ is the industry index. There are two ways to measure index: number of units and number of properties. In some cases, number of units is marked as 0 but number of properties owned by such owners (ie City of Milwaukee) can be quite large. The current preferred method is number of units.
# +
## city wide H index:
unitsSample = tax.loc[tax.nr_units>0,['nr_units','name']].groupby('name')
((unitsSample.sum()/unitsSample.ngroups)**2).sum()
# -
# ### Census Tract level index
print("Number of Census tracts in Milwaukee: %d" %tax.geo_tract.unique().shape[0])
## group by owner and tract -> sum by owner and divide by num of units in tract -> square then sum by tract
tractsSample = tax.loc[tax.nr_units>0,['nr_units','name','geo_tract']].groupby(['geo_tract','name'])
tractsSampleProps = tax.loc[tax.nr_units>0,['taxkey','name','geo_tract']].groupby(['geo_tract','name'])
## compute shares and sum squares by tract then select areas with more than 1 property
sharesByTract =(tractsSample.sum()/tractsSample.sum().groupby(level=0).transform('sum'))
tractHI = (sharesByTract**2).groupby(level=0).sum()[tractsSampleProps.size().groupby(level=0).sum()>=2]
# +
fig, axes = plt.subplots(1,3)
fig.set_figheight(5)
fig.set_figwidth(15)
plasma = plt.get_cmap("cool")
colors = iter(plasma(np.linspace(0,1,len(axes.flatten()))))
tractHI.hist(bins=15,ax=axes[0], color=next(colors))
tractHI[tractsSample.sum().groupby(level=0).sum()>10].hist(bins=15,ax=axes[1], color=next(colors))
tractHI[tractsSample.sum().groupby(level=0).sum()>20].hist(bins=15,ax=axes[2], color=next(colors))
for ax,title in zip(axes.flatten(),['all HHI','tracts w >10 units', 'tracts w >20 units']):
ax.set_ylim(0,20)
ax.set_ylabel("HHI censored above 20", fontsize = 18)
ax.set_title(title, fontsize = 20)
ax.set_yticks(np.arange(0,21,1))
print("Number of tracts with over 10 units: %d" %tractHI[tractsSample.sum().groupby(level=0).sum()>10].shape[0])
print("Number of tracts with concentration over .2: %d" %(tractHI[tractsSample.sum().groupby(level=0).sum()>10]>=.2).sum())
plt.savefig("/".join([data_path, 'HHI_tract.png']))
# -
# As can be seen between the 2 histograms immediately above, if there is any concentraion in market at the tract level, it seems it is due to census tracts with small number of units. However, once we exclude tracts with less than 10 units, it seems HHI stabilizes.
# ### Census Block Group level index
tax['tractblockgroup'] = tax.geo_tract.astype(str)+"."+tax.geo_blockgroup.astype(str)
print("Number of unique block groups: %d" %tax.tractblockgroup.unique().shape[0])
# +
groupSample = tax.loc[tax.nr_units>0,['nr_units','name','tractblockgroup']].groupby(['tractblockgroup','name'])
groupSampleProps = tax.loc[tax.nr_units>0,['taxkey','tractblockgroup']].groupby(['tractblockgroup']).size() #find number of props in block group
sharesByGroup =(groupSample.nr_units.sum()/groupSample.nr_units.sum().groupby(level=0).transform('sum'))
groupsHI = pd.concat([(sharesByGroup**2).groupby(level=0).sum(),groupSampleProps,groupSample.sum().groupby(level=0).sum()],axis=1,keys=['HHI','nrProps','nrUnits'])
groupsHI.columns = groupsHI.columns.droplevel(level=1)
# +
fig, axes = plt.subplots(1,3)
fig.set_figheight(5)
fig.set_figwidth(15)
plasma = plt.get_cmap("cool")
colors = iter(plasma(np.linspace(0,1,len(axes.flatten()))))
groupsHI.HHI.hist(bins=15,ax=axes[0], color=next(colors))
groupsHI.loc[(groupsHI.nrProps>1),'HHI'].hist(bins=15,ax=axes[1], color=next(colors))
groupsHI.loc[(groupsHI.nrUnits>10) & (groupsHI.nrProps>1),'HHI'].hist(bins=15,ax=axes[2], color=next(colors))
for ax,title in zip(axes.flatten(),['HHIs at block group level','block group w \n>1 props', 'block group w/ \n>10 units and >1 props']):
cens = 40
ax.set_ylim(0,cens)
ax.set_ylabel("HHI censored above %d" %cens, fontsize = 18)
ax.set_title(title, fontsize = 20)
ax.set_yticks(np.arange(0,42,2))
print("Number of block groups with more than 1 property: %d" %(groupsHI.nrProps>1).sum())
print("Number of groups with more than 10 units and 1 property with concentration over .2: %d" %(groupsHI[(groupsHI.nrUnits>10) & (groupsHI.nrProps>1)].HHI>=.2).sum())
plt.savefig("/".join([data_path, 'HHI_blockgroup.png']))
# -
# Census block group level is virtually the same as census tract level though not surprisingly since once is only twice as small as the other. As before, excluding small number of units seems to fix things a bit
# ### Census Block level index:
tax['tractblock'] = tax.geo_tract.astype(str)+"."+tax.geo_block.astype(str)
print("Number of unique blocks: %d" %tax.tractblock.unique().shape[0])
# +
## group by owner and block -> sum by owner and divide by num of units in tract -> square then sum by block
blocksSample = tax.loc[tax.nr_units>0,['nr_units','name','tractblock']].groupby(['tractblock','name'])
blocksSampleProps = tax.loc[tax.nr_units>0,['taxkey','tractblock']].groupby(['tractblock']).size()
sharesByBlock =(blocksSample.sum()/blocksSample.sum().groupby(level=0).transform('sum'))
blocksHI = pd.concat([(sharesByBlock**2).groupby(level=0).sum(), blocksSampleProps, blocksSample.sum().groupby(level=0).sum()], axis=1, keys = ['HHI','nrProps','nrUnits'])
blocksHI.columns = blocksHI.columns.droplevel(level=1)
# +
fig, axes = plt.subplots(1,3)
fig.set_figheight(5)
fig.set_figwidth(15)
plasma = plt.get_cmap("cool")
colors = iter(plasma(np.linspace(0,1,len(axes.flatten()))))
blocksHI.HHI.hist(bins=15,ax=axes[0], color=next(colors))
blocksHI[blocksHI.nrProps>1].HHI.hist(bins=15,ax=axes[1], color=next(colors))
blocksHI[(blocksHI.nrProps>1) & (blocksHI.nrUnits>10)].HHI.hist(bins=15,ax=axes[2], color=next(colors))
for ax,title in zip(axes.flatten(),['HHI at block level','blocks w \n>1 properties', 'blocks w \n>10 units & >1 properties']):
cens = 100
ax.set_ylim(0,cens)
ax.set_ylabel("HHI censored at over %d" %cens, fontsize =18)
ax.set_title(title, fontsize = 20)
ax.set_yticks(np.arange(0,110,10))
print("Number of blocks with over 10 units: %d" %((blocksHI.nrProps>1) & (blocksHI.nrUnits>10)).sum() )
print("Number of blocks with HHI over .2: %d" %((blocksHI.HHI>=.2) & (blocksHI.nrProps>1) & (blocksHI.nrUnits>10)).sum())
plt.savefig("/".join([data_path, 'HHI_block.png']))
# -
print("largest owners in high concentration blocks")
HighContrBlocks = blocksHI.HHI[blocksHI.HHI>.9].index
toLatex(blocksSample.sum().reset_index(level=1).loc[HighContrBlocks].sort_values('nr_units',ascending=False).head(n=15),"largestOwners.tex")
blocksHI.loc[(blocksHI.HHI>.9),:].head(n=15)
## Many of the high HHIs correspond with only 1 property on the block
print("Of HHIs equal to 1, proportion with only 1 property: %.2f" %(((blocksHI.HHI ==1) & (blocksHI.nrProps==1)).sum()/(blocksHI.HHI== 1).sum()))
print("Of HHIs greater than 20%%, proportion with only 1 property: %.2f" %(((blocksHI.HHI >.2) & (blocksHI.nrProps==1)).sum()/(blocksHI.HHI >.2).sum()))
# Same as the tract level, HHI is inflated when blocks with very few units is included but once we restrict to blocks with more than 10 units, HHI seems to stablize.
# ## Attach ACS data
# +
## remove block groups and blocks with less than 10 units
#groupsHI = groupsHI[groupSample.sum().groupby(level=0).sum()>=10]
#blocksHI = blocksHI[blocksSample.sum().groupby(level=0).sum()>=10]
# +
## load ACS data and examine tracts/block groups where there are high shares of low income households, blacks, or renters
block = pd.read_csv("/".join([data_path, '2010 Census Block Data Milwaukee.csv']))
group = pd.read_csv("/".join([data_path, '2011-2015 ACS block group Milwaukee.csv']))
groupRents = pd.read_csv("/".join([data_path, '2011-2015 ACS block group rents Milwaukee.csv']))
group.columns = group.columns.str.lower().str.replace(":|,",'')
block.columns = block.columns.str.lower().str.replace(":|,",'')
groupRents.columns = groupRents.columns.str.lower().str.replace(":|,|\$","")
block.drop(['area name-legal/statistical area description', 'qualifying name',
'area (land)', 'area (water)', 'summary level', 'geographic component',
'region', 'division', 'fips', 'state (fips)', 'county'],axis=1,inplace=True)
group.drop(['fips'],inplace=True,axis=1)
block.columns = block.columns.str.replace("\([\w\s\.]+\)|\.1",'').str.strip()
group.columns = group.columns.str.replace("\([\w\s\.]+\)|\.1",'').str.strip()
block.loc[:,'blockgroup'] = block.block.astype(str)[:1]
block = block.loc[:,~block.columns.duplicated()]
group = group.loc[:,~group.columns.duplicated()]
block.loc[:,'tractblock'] = block.loc[:,'census tract'].astype(str)+"."+block.loc[:,'block'].astype(str)
# -
print("Number of housing units according to Census blocks: %d" %block.loc[:,'housing units'].sum())
# +
## merge block level data to block level HHI
blocksHI = blocksHI.merge(block, left_index=True, right_on='tractblock',how='left')
# -
blocksHI['propBlack'] = np.divide(blocksHI.loc[:,'total population black or african american alone'],blocksHI.loc[:,'total population'])
blocksHI['propOver65'] = np.divide(blocksHI.loc[:,'total population 65 and over'],blocksHI.loc[:,'total population'])
blocksHI['propRent'] = np.divide(blocksHI.loc[:,'occupied housing units renter occupied'],blocksHI.loc[:,'occupied housing units'])
blocksHI['propVacant'] = np.divide(blocksHI.loc[:,'housing units vacant'],blocksHI.loc[:,'housing units'])
#drop blocks with unavialable census tract or block data
blocksHI = blocksHI[(blocksHI.block.notnull() & blocksHI.loc[:,'census tract'].notnull())]
blocksHI.loc[:,'housing units'].quantile([0,.1,.25,.5,.75,.9,1])
print("Number of properties with no housing units: %d" %(blocksHI.loc[:,'housing units'] == 0).sum())
print("HHI and population for blocks with 0 housing units:")
blocksHI.loc[(blocksHI.loc[:,'housing units'] == 0),['HHI','total population']].sort_values('total population',ascending=False).head()
print("Maybe remove HHIs with 0 population or housing units")
blocksHI.columns
# ## Analysis
# upload Tiger file, merge across tracts and blocks so my data have GEOID10 IDs that can be matched to shape files
toMap = blocksHI.merge(tigers, left_on = ['census tract','block'], right_on = ['TRACTCE10', 'BLOCKCE10'],how='outer')
toMap.columns = toMap.columns.str.title().str.replace(" ","")
toMap.to_csv("/".join([data_path,'blocksHHI.csv']))
toMap.to_excel("/".join([data_path, "blocksHHI.xlsx"]))
toMap.TotalPopulation.dropna().quantile([0,.25,.5,.75,1])
# +
## merge block group level data to group level HHI
group.loc[:,'tractgroup'] = group.loc[:,'census tract'].astype(str)+"."+group.loc[:,'block group'].astype(str)
groupRents.loc[:,'tractgroup'] = groupRents.loc[:,'census tract'].astype(str)+"."+groupRents.loc[:,'block group'].astype(str)
groupsHI = groupsHI.merge(group, left_index=True, right_on= 'tractgroup', how='left').merge(groupRents, how='left',on='tractgroup')
# +
#groupsHI.rename(columns = {"nr_units":'hhi'},inplace=True)
# -
groupsHI['propBlack'] = np.divide(groupsHI.loc[:,'total population black or african american alone'],groupsHI.loc[:,'total population'])
groupsHI['propOver65'] = np.divide(groupsHI.loc[:,'total population 65 and over'],groupsHI.loc[:,'total population'])
groupsHI['propRent'] = np.divide(groupsHI.loc[:,'occupied housing units renter occupied'],groupsHI.loc[:,'occupied housing units'])
groupsHI['propVacant'] = np.divide(groupsHI.loc[:,'housing units vacant'],groupsHI.loc[:,'housing units'])
groupsHI['unemployment'] = np.divide(groupsHI.loc[:,'population 16 years and over in labor force civilian unemployed'],groupsHI.loc[:,'population 16 years and over in labor force civilian',])
groupsHI['unitsPerPerson'] = np.divide(groupsHI.loc[:,'total population'],groupsHI.loc[:,'housing units'])
# +
rentDistr = np.divide(groupsHI[['renter-occupied housing units with cash rent less than 300',
'renter-occupied housing units with cash rent 300 to 599',
'renter-occupied housing units with cash rent 600 to 799',
'renter-occupied housing units with cash rent 800 to 999',
'renter-occupied housing units with cash rent 1000 to 1249',
'renter-occupied housing units with cash rent 1250 to 1499',
'renter-occupied housing units with cash rent 1500 to 1999',
'renter-occupied housing units with cash rent 2000 or more']],groupsHI['renter-occupied housing units with cash rent'].values.reshape(groupsHI.shape[0],1))
rentDistr.columns = rentDistr.columns.str.replace("renter-occupied housing units with cash ",'')
rentDistrCols = rentDistr.columns
#groupsHI = pd.concat([groupsHI, rentDistr],axis=1)
## most prevalent rent range in the data:
rentDistr.idxmax(axis=1).mode()
# -
groupsHousing = pd.read_excel("/".join([data_path, '2012-2016blockGroupACS.xlsx']))
groupsHousing.columns = groupsHousing.columns.str.replace("'|\$|,|\:|\-|Occupied|Housing|Units|Cash| Rent|with |\([\w\s]+\)","").str.replace(" "," ").str.strip().str.lower()
groupsHousing.columns = groupsHousing.columns.str.replace(" ","_")
groupsHI = groupsHI.merge(groupsHousing, left_on = ['census tract_x','block group_x'], right_on= ['census_tract', 'block_group'],how='left')
groupsHI.loc[:,'propCheap'] = np.divide(groupsHI['households_10000_to_14999']+groupsHI['households_less_than_10000'],groupsHI['households'])
groupsHI.rename(columns={'median gross rent':"medianGrossRent"},inplace=True)
# ## Relationship between HI index and Geographical Characteristics
# +
outDict = {}
regDict = {}
for x in ['np.log(median_value)','propCheap','propBlack','propRent','propVacant','medianGrossRent','propOver65','unitsPerPerson']:
form = "HHI~{}".format(x)
regDict[x] = smf.ols(formula=form, data=groupsHI.loc[(groupsHI.nrProps>1) & (groupsHI.nrUnits>9)]).fit()
cont = regDict[x]
inc = x
numObs = 'numObs'
outDict[x] = {'fitcoef': "%.04f (%.03f)" %(cont.params[inc].round(3), cont.pvalues[inc]),
'nObs':cont.nobs,
'R^2 adj':cont.rsquared_adj.round(3)}
regTbl = pd.DataFrame(outDict).loc[['fitcoef','R^2 adj','nObs']]
# -
toLatex(regTbl.transpose(),'blockGroupRegressions.tex')
# +
fig, axes = plt.subplots(4,2)
fig.set_figheight(15)
fig.set_figwidth(15)
plasma = plt.get_cmap("cool")
colors = iter(plasma(np.linspace(0,1,len(axes.flatten()))))
dotSize = 40
fontSize = 20
groupsHI= groupsHI.loc[groupsHI.nrUnits>1]
axes[0,0].scatter(x=groupsHI.propBlack,y=groupsHI.HHI, s=dotSize, color = next(colors))
axes[0,1].scatter(x=groupsHI.propOver65, y= groupsHI.HHI, s=dotSize, color = next(colors))
axes[1,0].scatter(x=groupsHI.propRent, y=groupsHI.HHI, s=dotSize, color=next(colors))
axes[1,1].scatter(x=groupsHI.propVacant, y=groupsHI.HHI, s=dotSize, color=next(colors))
axes[2,0].scatter(x=groupsHI.loc[:,'median household income'],y=groupsHI.HHI, s=dotSize, color=next(colors))
axes[2,1].scatter(x = groupsHI.unemployment, y=groupsHI.HHI,s=dotSize, color=next(colors) )
axes[3,0].scatter(x = groupsHI.loc[:,'medianGrossRent'], y = groupsHI.HHI, s=dotSize, color=next(colors))
axes[3,1].scatter(x = groupsHI.loc[:,'propCheap'], y = groupsHI.HHI, s=dotSize, color=next(colors))
for ax,xlab in zip(axes.flatten(),['proportion black','proportion over age 65','proportion who rent','vacancy rate','income','unemployment','median rent', 'unitsPerPerson']):
ax.set_ylim([0,.6])
#ax.set_xlim([-0.01,1.01])
ax.set_ylabel("HHI")
ax.set_xlabel(xlab, fontsize = fontSize)
#ax.set_title(xlab + " at block groups", fontsize = fontSize+1)
fig.subplots_adjust(hspace=.5)
print("For block groups with more than 1 property:")
#plt.savefig("/".join([data_path,'blockGroups_acs.png']))
# +
outDict = {}
regDict = {}
blocksHI = blocksHI.loc[(blocksHI.nrProps>1) & (blocksHI.nrUnits>9)]
for x in ['propBlack','propRent','propVacant','propOver65']:
form = "HHI~{}".format(x)
regDict[x] = smf.ols(formula=form, data=blocksHI.loc[(blocksHI.nrProps>1) & (blocksHI.nrUnits>9)]).fit()
cont = regDict[x]
inc = x
numObs = 'numObs'
outDict[x] = {'fitcoef': "%.04f (%.03f)" %(cont.params[inc], cont.pvalues[inc]),
'nObs':cont.nobs,
'R^2 adj':cont.rsquared_adj}
regTbl = pd.DataFrame(outDict).loc[['fitcoef','R^2 adj','nObs']]
toLatex(regTbl,'blockRegressions.tex')
# +
fig, axes = plt.subplots(2,2)
fig.set_figheight(15)
fig.set_figwidth(15)
plasma = plt.get_cmap("cool")
colors = iter(plasma(np.linspace(0,1,len(axes.flatten()))))
dotSize = 40
fontSize = 20
blocksHI = blocksHI.loc[(blocksHI.nrProps>1) & (blocksHI.nrUnits>9)]
axes[0,0].scatter(x=blocksHI.propBlack,y=blocksHI.HHI, s=dotSize, color = next(colors))
axes[0,1].scatter(x=blocksHI.propOver65, y= blocksHI.HHI, s=dotSize, color = next(colors))
axes[1,0].scatter(x=blocksHI.propRent, y=blocksHI.HHI, s=dotSize, color=next(colors))
axes[1,1].scatter(x=blocksHI.propVacant, y=blocksHI.HHI, s=dotSize, color=next(colors))
for ax,xlab in zip(axes.flatten(),['proportion black','proportion over age 65','proportion who rent','vacancy rate']):
ax.set_ylim([0,1])
ax.set_xlim([-0.01,1.01])
ax.set_ylabel("Herfindahl–Hirschman Index")
ax.set_xlabel(xlab, fontsize = fontSize)
ax.set_title(xlab + " at block level", fontsize = fontSize+1)
fig.subplots_adjust(hspace=.5)
print("For blocks with more than 1 property and 10 units")
plt.savefig("/".join([data_path, "blocks_acs.png"]))
# -
# # Geocode tax assessor
#
# arcGIS [geocoding](https://developers.arcgis.com/python/guide/batch-geocoding/) directions are a mess and don't work half the time but, nevertheless, do the job after persistent effort.
#
# Basically, have to geocode all addresses. On the other hand, addresses are already assigned census tracts and census block numbers which alleviates most of the need to geocode the addresses.
# +
#tax.lookupAddress.to_csv("/".join([data_path, 'addresses2geocode.csv']))
# +
from arcgis.gis import *
from arcgis.geocoding import geocode
#from arcgis.geometry import Point
from arcgis.gis import GIS
from arcgis.geocoding import get_geocoders, batch_geocode
gis = GIS("http://www.arcgis.com", "burinski_priceusc", "Disturb3")
# use the first of GIS's configured geocoders
geocoder = get_geocoders(gis)[0]
print("MaxBatchSize : " + str(geocoder.properties.locatorProperties.MaxBatchSize))
print("SuggestedBatchSize : " + str(geocoder.properties.locatorProperties.SuggestedBatchSize))
# +
def to_addrDict(x):
addrDict = {
'Street': " ".join([str(x['house_nr_hi']),x['street'],x['sttype']]).lower(),
'City':"Milwaukee",
'State':"WI",
'Zone':str(x['geo_zip_code'])[:5]}
return(addrDict)
def to_addrStr(x):
s= " ".join([str(x['house_nr_hi']),x['street'],str(x['sttype']) + ",","Milwaukee, WI", str(x['geo_zip_code'])[:5]]).title()
return(s)
def parse_geocode_results(results):
#topres = [r for r in results if r['score']>99][0]['location']
x = results[0]['location']['x']
y = results[0]['location']['y']
return((x,y))
def parse_batch_results(results):
coordDict = {}
coordDict['x'] = []
coordDict['y'] = []
for result in results:
coordDict['x'].append(result['location']['x'])
coordDict['y'].append(result['location']['y'])
return(pd.DataFrame(coordDict))
# +
#make address string for all addresses in the data set
tax.loc[:,'lookupAddress'] = tax.apply(lambda r: to_addrStr(r),axis=1)
## geocode single address
#results = geocode(addresses.iloc[0])
#parse_geocode_results(results)
## i can batch geocode the addresses OR code them individually. For one, I'll be charged, the other, I can get away with
#results = batch_geocode(list(addresses))
#parse_batch_results(results)
# -
## split my data frame into 200 parts
splitArray = np.array_split(tax.lookupAddress, int(tax.shape[0]/200))
len(splitArray)
## batch geocode them in 200s until done
chunkResults = []
for i,chunk in enumerate(splitArray[0:]):
if i%100 == 0:
print("Working on chunk: %d" %i)
#print("\tChunk size: %d" %chunk.shape[0])
try:
temp = parse_batch_results(batch_geocode(list(chunk)))
temp.index = chunk.index
chunkResults.append(temp)
except RuntimeError as e:
print(e)
print("stopped at chunk num: %d" %i)
break
codedBatch = pd.concat(chunkResults)
codedBatch.to_csv("/".join([data_path, 'geocodedAddresses_4thBatch.csv']))
# +
## batch stopped working so I am now turning to geocode.
chunkResults = []
for i,chunk in enumerate(splitArray[358:]):
if i%10 == 0:
print("Working on chunk: %d" %i)
print("\tChunk size: %d" %chunk.shape[0])
try:
temp = chunk.apply(lambda s: parse_geocode_results(geocode(s)))
temp.index = chunk.index
temp = temp.apply(pd.Series)
temp.columns = ['x','y']
chunkResults.append(temp)
except RuntimeError as e:
print(e)
print("stopped at chunk num: %d" %i)
break
# -
print("Number of chunks processed: %d" %len(chunkResults))
codedBatch = pd.concat(chunkResults)
codedBatch.to_csv("/".join([data_path, 'geocodedAddresses_4thBatch.csv']))
tempLoad = pd.read_csv("/".join([data_path, 'geocodedAddresses_3rdBatch.csv']))
tempLoad.shape[0]
9000/200
358+45
# +
batchDict = {}
for s in ['geocodedAddresses_1stBatch.csv','geocodedAddresses_2stBatch.csv','geocodedAddresses_3rdBatch.csv']:
batchDict[s] = pd.read_csv("/".join([data_path, s]))
batches = pd.concat(batchDict)
# -
batches.set_index('Unnamed: 0',inplace=True)
| marketConcentration_Milwaukee.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ml39
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Numpy Basics
# > A quick walk through vector and matrix multiplication with Numpy.
#
# - toc: true
# - badges: true
# #- title: Numpy Basics
# - hide: false
# - comments: true
# - author: <NAME>
# - image: images/numpy.png
# - tags: [python, numpy, matrix, vector]
# - permalink: /:title
# - categories: [python, numpy, matrix, vector]
#
#
# +
import numpy as np
from pprint import pprint
import matplotlib.pyplot as plt
# hadamard product = element-wise = a*b
# outer product = a*b.T
# inner product = dot product = a.T*b (Transpose here is only for distinction, np.dot() does not automagically Transpose, np.dot() expects proper dimensions for matrix math)
# a*b does a hadamard product
# dot(av,bv) of vectors does a hadamard product
# dot(am,bm) of matrix does matrix multiplication
# -
# # Hadamard Product
# Hadamard Product = (1*2), (1*2), (1*2)= <2,2,2>
a = np.array([1,1,1]) #this is not a column or row vector
b = np.array([2,2,2]) #this is not a column or row vector
print(f"\n(1)\na and b shape: {a.shape} --> dot(a,b) = \n{a*b}")
# # Outer Product
# +
# One way to force a row or column vector is to use double bracket [[]]
# where [[0], is a 2x1 column vector
# [0]]
# and [[0,0]] is a 1x2 row vector
ac = np.array([ [1],
[1],
[1]])
bc = np.array( [[2],
[2],
[2]])
print(f"\n(2)\nar and bc shape: {ac.shape} {bc.shape} --> \ndot(ac,bc.T) = \n{np.dot(ac,bc.T)} \ndot(bc,ac.T) = \n{np.dot(bc,ac.T)}")
# -
# # Inner Product
# inner product = dot product (1*2)+(1*2)+(1*2)=6
a = np.array([1,1,1]) #this is not a column or row vector
b = np.array([2,2,2]) #this is not a column or row vector
print(f"\n(1)\na and b shape: {a.shape} --> dot(a,b) = \n{np.dot(a,b)}")
# +
# One way to force a row or column vector is to use double bracket [[]]
# where [[0], is a 2x1 column vector
# [0]]
# and [[0,0]] is a 1x2 row vector
ac = np.array([ [1],
[1],
[1]])
bc = np.array( [[2],
[2],
[2]])
print(f"\n(2)\nar and bc shape: {ac.shape} {bc.shape} --> \ndot(ac.T,bc) = \n{np.dot(ac.T,bc)} \ndot(bc.T,ac) = \n{np.dot(bc.T,ac)}")
# Compare this with the Outer Product version above
# +
ar = np.array([[1,1,1 ]])
bc = np.array( [[2],
[2],
[2]])
print(f"\n(3)\nar and bc shape: {ar.shape} {bc.shape} --> \ndot(ar,bc) = \n{np.dot(ar,bc)} \ndot(bc,ar) = \n{np.dot(bc,ar)}")
# -
ar = np.array([[1,1],
[1,1]])
bc = np.array( [[1, 3],
[2, 2]
])
print(f"ar and bc shape: {ar.shape} {bc.shape} --> \ndot(ar,bc) = \n{np.dot(ar,bc)} \ndot(bc,ar) = \n{np.dot(bc,ar)} \n ar*bc = \n{ar*bc}")
# # Wrong Dimensions
# +
# Wrong Dimensions
am = np.array([[1,1,1,1]])
bm = np.array( [[2],
[2],
[2]])
print(f"ar and bc shape: {am.shape} {bm.shape} --> dot(a,b) = {np.dot(am,bm)}")
# Inner dimensions must match, ie 4 and 4 or 3 and 3 here.
# Error: ValueError: shapes (1,4) and (3,1) not aligned: 4 (dim 1) != 3 (dim 0)
| _notebooks/2022-02-21-Numpy-Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/moh2236945/pytorch_classification/blob/master/models/resnext50.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="0i1d6yW6xfNt" colab_type="code" colab={}
import torch
import torch.nn as nn
import torch.nn.functional as F
from collections import OrderedDict
from torch.autograd import Variable
"""
NOTICE:
BasicBlock_B is not implemented
BasicBlock_C is recommendation
The full architecture consist of BasicBlock_A is not implemented.
"""
class ResBottleBlock(nn.Module):
def __init__(self, in_planes, bottleneck_width=4, stride=1, expansion=1):
super(ResBottleBlock, self).__init__()
self.conv0=nn.Conv2d(in_planes,bottleneck_width,1,stride=1,bias=False)
self.bn0 = nn.BatchNorm2d(bottleneck_width)
self.conv1=nn.Conv2d(bottleneck_width,bottleneck_width,3,stride=stride,padding=1,bias=False)
self.bn1=nn.BatchNorm2d(bottleneck_width)
self.conv2=nn.Conv2d(bottleneck_width,expansion*in_planes,1,bias=False)
self.bn2=nn.BatchNorm2d(expansion*in_planes)
self.shortcut=nn.Sequential()
if stride!=1 or expansion!=1:
self.shortcut=nn.Sequential(
nn.Conv2d(in_planes,in_planes*expansion,1,stride=stride,bias=False)
)
def forward(self, x):
out = F.relu(self.bn0(self.conv0(x)))
out = F.relu(self.bn1(self.conv1(out)))
out = self.bn2(self.conv2(out))
out += self.shortcut(x)
out = F.relu(out)
return out
class BasicBlock_A(nn.Module):
def __init__(self, in_planes, num_paths=32, bottleneck_width=4, expansion=1, stride=1):
super(BasicBlock_A,self).__init__()
self.num_paths = num_paths
for i in range(num_paths):
setattr(self,'path'+str(i),self._make_path(in_planes,bottleneck_width,stride,expansion))
# self.paths=self._make_path(in_planes,bottleneck_width,stride,expansion)
self.conv0=nn.Conv2d(in_planes*expansion,expansion*in_planes,1,stride=1,bias=False)
self.bn0 = nn.BatchNorm2d(in_planes * expansion)
self.shortcut = nn.Sequential()
if stride != 1 or expansion != 1:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, in_planes * expansion, 1, stride=stride, bias=False)
)
def forward(self, x):
out = self.path0(x)
for i in range(1,self.num_paths):
if hasattr(self,'path'+str(i)):
out+getattr(self,'path'+str(i))(x)
# out+=self.paths(x)
# getattr
# out = torch.sum(out, dim=1)
out = self.bn0(out)
out += self.shortcut(x)
out = F.relu(out)
return out
def _make_path(self, in_planes, bottleneck_width, stride, expansion):
layers = []
layers.append(ResBottleBlock(
in_planes, bottleneck_width, stride, expansion))
return nn.Sequential(*layers)
class BasicBlock_C(nn.Module):
"""
increasing cardinality is a more effective way of
gaining accuracy than going deeper or wider
"""
def __init__(self, in_planes, bottleneck_width=4, cardinality=32, stride=1, expansion=2):
super(BasicBlock_C, self).__init__()
inner_width = cardinality * bottleneck_width
self.expansion = expansion
self.basic = nn.Sequential(OrderedDict(
[
('conv1_0', nn.Conv2d(in_planes, inner_width, 1, stride=1, bias=False)),
('bn1', nn.BatchNorm2d(inner_width)),
('act0', nn.ReLU()),
('conv3_0', nn.Conv2d(inner_width, inner_width, 3, stride=stride, padding=1, groups=cardinality, bias=False)),
('bn2', nn.BatchNorm2d(inner_width)),
('act1', nn.ReLU()),
('conv1_1', nn.Conv2d(inner_width, inner_width * self.expansion, 1, stride=1, bias=False)),
('bn3', nn.BatchNorm2d(inner_width * self.expansion))
]
))
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != inner_width * self.expansion:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, inner_width * self.expansion, 1, stride=stride, bias=False)
)
self.bn0 = nn.BatchNorm2d(self.expansion * inner_width)
def forward(self, x):
out = self.basic(x)
out += self.shortcut(x)
out = F.relu(self.bn0(out))
return out
class ResNeXt(nn.Module):
def __init__(self, num_blocks, cardinality, bottleneck_width, expansion=2, num_classes=10):
super(ResNeXt, self).__init__()
self.cardinality = cardinality
self.bottleneck_width = bottleneck_width
self.in_planes = 64
self.expansion = expansion
self.conv0 = nn.Conv2d(3, self.in_planes, kernel_size=3, stride=1, padding=1)
self.bn0 = nn.BatchNorm2d(self.in_planes)
self.pool0 = nn.MaxPool2d(kernel_size=3, stride=2, padding=1)
self.layer1=self._make_layer(num_blocks[0],1)
self.layer2=self._make_layer(num_blocks[1],2)
self.layer3=self._make_layer(num_blocks[2],2)
self.layer4=self._make_layer(num_blocks[3],2)
self.linear = nn.Linear(self.cardinality * self.bottleneck_width, num_classes)
def forward(self, x):
out = F.relu(self.bn0(self.conv0(x)))
# out = self.pool0(out)
out = self.layer1(out)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
out = F.avg_pool2d(out, 4)
out = out.view(out.size(0), -1)
out = self.linear(out)
return out
def _make_layer(self, num_blocks, stride):
strides = [stride] + [1] * (num_blocks - 1)
layers = []
for stride in strides:
layers.append(BasicBlock_C(self.in_planes, self.bottleneck_width, self.cardinality, stride, self.expansion))
self.in_planes = self.expansion * self.bottleneck_width * self.cardinality
self.bottleneck_width *= 2
return nn.Sequential(*layers)
def resnext26_2x64d():
return ResNeXt(num_blocks=[2, 2, 2, 2], cardinality=2, bottleneck_width=64)
def resnext26_4x32d():
return ResNeXt(num_blocks=[2, 2, 2, 2], cardinality=4, bottleneck_width=32)
def resnext26_8x16d():
return ResNeXt(num_blocks=[2, 2, 2, 2], cardinality=8, bottleneck_width=16)
def resnext26_16x8d():
return ResNeXt(num_blocks=[2, 2, 2, 2], cardinality=16, bottleneck_width=8)
def resnext26_32x4d():
return ResNeXt(num_blocks=[2, 2, 2, 2], cardinality=32, bottleneck_width=4)
def resnext26_64x2d():
return ResNeXt(num_blocks=[2, 2, 2, 2], cardinality=32, bottleneck_width=4)
def resnext50_2x64d():
return ResNeXt(num_blocks=[3, 4, 6, 3], cardinality=2, bottleneck_width=64)
def resnext50_32x4d():
return ResNeXt(num_blocks=[3, 4, 6, 3], cardinality=32, bottleneck_width=4)
# def test():
# net = resnext50_2x64d()
# # print(net)
# data = Variable(torch.rand(1, 3, 32, 32))
# output = net(data)
# print(output.size())
# + id="MCnWUy2GxuZz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0e4d6f93-44d6-4dbd-e3af-5a8704febffa"
model = resnext50_2x64d()
device = torch.device("cuda") if torch.cuda.is_available() else torch.device("cpu")
model.to(device)
summary(model,(3, 32, 32))
| models/resnext50.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Stochastische Variabelen Genereren
# In het onderstaande Jupyter-verslag zal een standaard normaalverdeling worden gegenereerd door middel van uniform verdeelde psuedo-random getallen.
import scripts.RandomNumberGenerator as RNG
from scipy.special import ndtri
from scipy.stats import normaltest
import seaborn as sns
# Het aanmaken van een seed waarop de pseudo-random getallen worden gebaseerd.
seed = RNG.generate_seed()
seed
# 10.000 pseudo-random getallen genereren met de zojuist gegenereerde seed.
data = RNG.generator_endsquare(samples=10_000, seed=seed, show_seed=False)
print(data)
# Om een uniforme naar een normaalverdeling om te zetten dient van ieder getal de inverse van het CDF worden te berekend.
#
# [Inverse Transform Sampling](https://en.wikipedia.org/wiki/Inverse_transform_sampling)
#
#
# [Inverse van CDF in Python](https://stackoverflow.com/questions/20626994/how-to-calculate-the-inverse-of-the-normal-cumulative-distribution-function-in-p)
inverse_cdf = ndtri(data)
inverse_cdf
# Alle 10.000 waardes worden vervolgens geplot. De normaalverdeling hieronder is een standaard normaalverdeling. Deze heeft een $\mu$ van $0$ en een $\sigma$ van $1$. Tussen $-3$ en $3$ standaarddeviaties zullen zich dus >99% van de waardes bevinden.
sns.set(context="notebook")
sns.distplot(inverse_cdf)
# Om te testen of wat lijkt op een standaard normaalverdeling ook daadwerkelijk een standaard normaalverdeling is, moet dit worden getoetst. Dit wordt gedaan door middel van een hypothesetoets met de volgende twee hypotheses:
#
# H0: De verdeling is standaard normaalverdeeld
# H1: De verdeling is niet standaard normaalverdeeld
# +
_, p = normaltest(inverse_cdf)
alpha = 0.05
print(f"p = {p:.6f}")
if p < alpha:
print("H0 kan worden verworpen, H1 wordt aanvaard.")
else:
print("H0 kan niet worden verworpen, we blijven sceptisch.")
# -
| StochastischeVariabelenGenereren.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="yhyXAZv7SQCb"
# # LEDNet
# Instalación de la librería fast.ai (a continuación reiniciar el entorno de ejecución).
# + id="uLA8cBBNmFMD"
# !pip install fastai --upgrade
# + id="FTz7Hhm7qR1e" executionInfo={"status": "ok", "timestamp": 1623895128636, "user_tz": -120, "elapsed": 5209, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
from fastai.basics import *
from fastai.vision import models
from fastai.vision.all import *
from fastai.metrics import *
from fastai.data.all import *
from fastai.callback import *
from fastai.learner import defaults, Learner
from pathlib import Path
import random
# + [markdown] id="99sCgNTFSnBW"
# Descarga de la librería de arquitecturas.
# + id="2I2ze0KkSrNo"
# !wget https://www.dropbox.com/s/cmoblvx5icdifwl/architectures.zip?dl=1 -O architectures.zip
# !unzip architectures.zip
# + [markdown] id="93vZS9hOSybL"
# Descarga del dataset.
# + id="6tyw8cWTrJ8x"
# !wget https://www.dropbox.com/s/p92cw15pleunmqe/dataset.zip?dl=1 -O dataset.zip
# !unzip dataset.zip
# + [markdown] id="Z9wjeZ6dS5AR"
# Conexión con Drive para el almacenaje de los modelos.
# + id="RuUZJ8mmvQvW"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="I5biE_ZpTDhI"
# Rutas a los directorios del dataset.
# + id="FmLabxI3riWS" executionInfo={"status": "ok", "timestamp": 1623895166927, "user_tz": -120, "elapsed": 280, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
path=Path('dataset/')
path_images = path/"Images"
path_labels = path/"Labels"
test_name = "test"
# + [markdown] id="-gkexNyyT1Ou"
# Función que dada la ruta de una imagen devuelve el path de su anotación.
# + id="vGqcWni3roF2" executionInfo={"status": "ok", "timestamp": 1623895167732, "user_tz": -120, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
def get_y_fn (x):
return Path(str(x).replace("Images","Labels"))
# + [markdown] id="nS-mGv7MTQ4j"
# Clases: Background y Stoma.
# + id="jS74xiUkrvSc" executionInfo={"status": "ok", "timestamp": 1623895168804, "user_tz": -120, "elapsed": 273, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
codes = np.loadtxt(path/'codes.txt', dtype=str)
# + [markdown] id="P2DbR2nNUKFv"
# Función que permite partir el dataset entre entrenamiento y test.
# + id="QkER6uQkr_k5" executionInfo={"status": "ok", "timestamp": 1623895169941, "user_tz": -120, "elapsed": 3, "user": {"displayName": "<NAME> de clase", "photoUrl": "", "userId": "15591564048203513807"}}
def ParentSplitter(x):
return Path(x).parent.name==test_name
# + [markdown] id="1IYm4PFWUXDt"
# # Data augmentation
# Carga de la librería Albumentations.
# + id="TV1mWQWVsSgY" executionInfo={"status": "ok", "timestamp": 1623895173666, "user_tz": -120, "elapsed": 1154, "user": {"displayName": "<NAME> clase", "photoUrl": "", "userId": "15591564048203513807"}}
from albumentations import (
Compose,
OneOf,
ElasticTransform,
GridDistortion,
OpticalDistortion,
HorizontalFlip,
Flip,
Rotate,
Transpose,
CLAHE,
ShiftScaleRotate
)
class SegmentationAlbumentationsTransform(ItemTransform):
split_idx = 0
def __init__(self, aug):
self.aug = aug
def encodes(self, x):
img,mask = x
aug = self.aug(image=np.array(img), mask=np.array(mask))
return PILImage.create(aug["image"]), PILMask.create(aug["mask"])
# + [markdown] id="pljyxxaGUvBA"
# Transformación que aplica a las imagenes giros horizontales, rotaciones y una operación de distorsión.
# + id="LoPU-tVEstUE" executionInfo={"status": "ok", "timestamp": 1623895173668, "user_tz": -120, "elapsed": 5, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
transforms=Compose([HorizontalFlip(p=0.5),
Flip(p=0.5),
Rotate(p=0.40,limit=10)
],p=1)
# + id="8NbVeTzMsxSR" executionInfo={"status": "ok", "timestamp": 1623895174726, "user_tz": -120, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
transformPipeline=SegmentationAlbumentationsTransform(transforms)
# + [markdown] id="JqVYa4ns1-T8"
# Transformación que no aplica cambios a las imagenes.
# + id="Mi72sNDK1sB5" executionInfo={"status": "ok", "timestamp": 1623895175579, "user_tz": -120, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
transforms2=Compose([],p=1)
# + id="_Tz9tjPJ1sPz" executionInfo={"status": "ok", "timestamp": 1623895176450, "user_tz": -120, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
transform2Pipeline=SegmentationAlbumentationsTransform(transforms2)
# + [markdown] id="VdvYcbsYVEMb"
# Transformación que cambia todos los píxeles con valor 255 a valor 1 en las máscaras.
# + id="rzXohObls1_x" executionInfo={"status": "ok", "timestamp": 1623895177324, "user_tz": -120, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
class TargetMaskConvertTransform(ItemTransform):
def __init__(self):
pass
def encodes(self, x):
img,mask = x
#Convert to array
mask = np.array(mask)
mask[mask!=255]=0
# Change 255 for 1
mask[mask==255]=1
# Back to PILMask
mask = PILMask.create(mask)
return img, mask
# + [markdown] id="yTlxnkZKVQAv"
# # Dataloaders
# DataBlock de entrenamiento con aumento de datos.
# + id="EDxRc4LAtNfP" executionInfo={"status": "ok", "timestamp": 1623895189534, "user_tz": -120, "elapsed": 10735, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
trainDB = DataBlock(blocks=(ImageBlock, MaskBlock(codes)),
get_items=partial(get_image_files,folders=['train']),
get_y=get_y_fn,
splitter=RandomSplitter(valid_pct=0.2),
item_tfms=[Resize((50,50)), TargetMaskConvertTransform(), transformPipeline],
batch_tfms=Normalize.from_stats(*imagenet_stats)
)
# + [markdown] id="LGX4V5sTPNxs"
# DataBlock de entrenamiento sin aumento de datos.
# + id="DYKSbTQFPPYK" executionInfo={"status": "ok", "timestamp": 1623895189534, "user_tz": -120, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
train2DB = DataBlock(blocks=(ImageBlock, MaskBlock(codes)),
get_items=partial(get_image_files,folders=['train']),
get_y=get_y_fn,
splitter=RandomSplitter(valid_pct=0.2),
item_tfms=[Resize((50,50)), TargetMaskConvertTransform(), transform2Pipeline],
batch_tfms=Normalize.from_stats(*imagenet_stats)
)
# + [markdown] id="wfEn8J3rVWOx"
# DataBlock de test.
# + id="C64oABoRtRBV" executionInfo={"status": "ok", "timestamp": 1623895189535, "user_tz": -120, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
testDB = DataBlock(blocks=(ImageBlock, MaskBlock(codes)),
get_items=partial(get_image_files,folders=['train','test']),
get_y=get_y_fn,
splitter=FuncSplitter(ParentSplitter),
item_tfms=[Resize((50,50)), TargetMaskConvertTransform(), transformPipeline],
batch_tfms=Normalize.from_stats(*imagenet_stats)
)
# + [markdown] id="9ut0DwEMVeqA"
# Creación de los dataloaders.
# + id="y7fDTTLgtUZI" executionInfo={"status": "ok", "timestamp": 1623895190388, "user_tz": -120, "elapsed": 856, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
bs = 2
trainDLS = trainDB.dataloaders(path_images,bs=bs)
train2DLS = trainDB.dataloaders(path_images,bs=bs)
testDLS = testDB.dataloaders(path_images,bs=bs)
# + [markdown] id="svZDNg1fVhFx"
# Prueba de la carga de datos.
# + id="P27jeYx_tZUZ" colab={"base_uri": "https://localhost:8080/", "height": 335} executionInfo={"status": "ok", "timestamp": 1623895190393, "user_tz": -120, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="0ece36ff-0fd9-4bd6-d092-915ed99eb7cb"
trainDLS.show_batch(vmin=0,vmax=1,figsize=(12, 9))
# + [markdown] id="Sm5Nh5AqVpC9"
# # Modelos con aumento de datos
# Definición del modelo.
# + id="6kZnzPdUtkO5" executionInfo={"status": "ok", "timestamp": 1623895190897, "user_tz": -120, "elapsed": 510, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
from architectures import LEDNet
model = LEDNet(nclass=2)
# + [markdown] id="4VtE39t6EqJV"
# Creación del Learner con wd=1e-2 y definición del directorio de trabajo.
# + id="pdYLySprtotd" executionInfo={"status": "ok", "timestamp": 1623895190898, "user_tz": -120, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
learn = Learner(dls=trainDLS, model=model, metrics=[Dice(), JaccardCoeff()], wd=1e-2)
learn.model_dir = "/content/drive/MyDrive/Colab Notebooks/LEDNet"
# + [markdown] id="Y0uU2M9XFjRH"
# Freeze y elección de la tasa de aprendizaje.
# + id="EuxePLnJhmSe" colab={"base_uri": "https://localhost:8080/", "height": 300} executionInfo={"status": "ok", "timestamp": 1623309677412, "user_tz": -120, "elapsed": 10115, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="834d800d-6376-412b-fb84-c447e1601505"
learn.freeze()
learn.lr_find()
learn.recorder
# + [markdown] id="96r_xyeKFwk3"
# Entrenamiento del modelo usando EarlyStoppingCallback según el valid_loss (min_delta=0.0001 y patience=2).
# + id="5DTnmRK0ttep" colab={"base_uri": "https://localhost:8080/", "height": 566} executionInfo={"status": "ok", "timestamp": 1623312790827, "user_tz": -120, "elapsed": 3113420, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="e1049c56-a714-4d85-fece-44f844c89608"
name = "model_LEDNet_da_wd2"
learn.fit_one_cycle(100,slice(1e-5,1e-4),cbs=[
EarlyStoppingCallback(monitor='valid_loss', min_delta=0.0001, patience=2),
ShowGraphCallback(),
SaveModelCallback(monitor='valid_loss', min_delta=0.0001, fname=name, every_epoch=False)])
# + [markdown] id="VveKZbMdGGhn"
# Comprobación de los resultados obtenidos en el modelo almacenado.
# + id="xt7H1pqP0ejE" colab={"base_uri": "https://localhost:8080/", "height": 67} executionInfo={"status": "ok", "timestamp": 1623312829578, "user_tz": -120, "elapsed": 38763, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="5dcdc17a-cf3f-4d31-85b0-433576c97c6d"
learn.load("model_LEDNet_da_wd2")
learn.validate()
# + [markdown] id="Pg9TYtg4G7ps"
# Unfreeze y elección de la tasa de aprendizaje.
# + id="iVUWPqjv4Gfp" colab={"base_uri": "https://localhost:8080/", "height": 302} executionInfo={"status": "ok", "timestamp": 1623312838709, "user_tz": -120, "elapsed": 9135, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="9bd49bdb-d3c7-483c-ab5c-def01ce240bf"
learn.unfreeze()
learn.lr_find()
learn.recorder
# + [markdown] id="OLD76K0PHIVY"
# Entrenamiento del modelo usando EarlyStoppingCallback según el valid_loss (min_delta=0.0001 y patience=2).
# + id="aLWT2EAS4Ktv" colab={"base_uri": "https://localhost:8080/", "height": 967} executionInfo={"status": "ok", "timestamp": 1623315411161, "user_tz": -120, "elapsed": 383596, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="d60bd44a-6d7a-4394-deb4-8d16d38a32c9"
name = "model_LEDNet_da_wd2_unfreeze"
learn.fit_one_cycle(100,slice(1e-5,1e-4),cbs=[
EarlyStoppingCallback(monitor='valid_loss', min_delta=0.0001, patience=2),
ShowGraphCallback(),
SaveModelCallback(monitor='valid_loss', min_delta=0.0001, fname=name, every_epoch=False)])
# + [markdown] id="9NO06lrAHOU1"
# Comprobación de los resultados obtenidos en el modelo almacenado.
# + id="mo-mXxss4bJm" colab={"base_uri": "https://localhost:8080/", "height": 67} executionInfo={"status": "ok", "timestamp": 1623315449494, "user_tz": -120, "elapsed": 38346, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="8053fb18-5add-4ad4-8576-ab16bb590498"
learn.load("model_LEDNet_da_wd2_unfreeze")
learn.validate()
# + [markdown] id="A72AE009MC7K"
#
#
# ---
#
#
# Definición del modelo.
# + id="V-cgcM09DQpi"
del model, learn
model = LEDNet(nclass=2)
# + [markdown] id="SJU4N2LUMPd-"
# Creación del Learner con wd=1e-1 y definición del directorio de trabajo.
# + id="WsO8n5Ij3RhJ"
learn = Learner(dls=trainDLS, model=model, metrics=[Dice(), JaccardCoeff()], wd=1e-1)
learn.model_dir = "/content/drive/MyDrive/Colab Notebooks/LEDNet"
# + [markdown] id="1nanYEdmMVLc"
# Freeze y elección de la tasa de aprendizaje.
# + id="y3wEp_P93VUo" colab={"base_uri": "https://localhost:8080/", "height": 300} executionInfo={"status": "ok", "timestamp": 1623317255840, "user_tz": -120, "elapsed": 8939, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="919b1be1-49fc-4fa0-8174-3b73d28f15ab"
learn.freeze()
learn.lr_find()
learn.recorder
# + [markdown] id="7-UYzyzTMb6J"
# Entrenamiento del modelo usando EarlyStoppingCallback según el valid_loss (min_delta=0.0001 y patience=2).
# + id="vcT9ObOzRL-_" colab={"base_uri": "https://localhost:8080/", "height": 597} executionInfo={"status": "ok", "timestamp": 1623320918764, "user_tz": -120, "elapsed": 3662927, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="12f7ce02-0e96-4a4a-99e9-57d1967c18c4"
name = "model_LEDNet_da_wd1"
learn.fit_one_cycle(100,slice(1e-5,1e-4),cbs=[
EarlyStoppingCallback(monitor='valid_loss', min_delta=0.0001, patience=2),
ShowGraphCallback(),
SaveModelCallback(monitor='valid_loss', min_delta=0.0001, fname=name, every_epoch=False)])
# + [markdown] id="rVp7lbfEMk-p"
# Comprobación de los resultados obtenidos en el modelo almacenado.
# + id="F2wsdj4zRMMi" colab={"base_uri": "https://localhost:8080/", "height": 67} executionInfo={"status": "ok", "timestamp": 1623320957727, "user_tz": -120, "elapsed": 38969, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="5d97bd97-e4a8-4143-d591-fb0b7bf14487"
learn.load("model_LEDNet_da_wd1")
learn.validate()
# + [markdown] id="gQZx3gdRMppi"
# Unfreeze y elección de la tasa de aprendizaje.
# + id="NZkQkIyHR2K9" colab={"base_uri": "https://localhost:8080/", "height": 300} executionInfo={"status": "ok", "timestamp": 1623320967171, "user_tz": -120, "elapsed": 9450, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="2e997293-c5d9-458f-ff0d-e6d9fd61392e"
learn.unfreeze()
learn.lr_find()
learn.recorder
# + [markdown] id="dtgoRxSiMtVH"
# Entrenamiento del modelo usando EarlyStoppingCallback según el valid_loss (min_delta=0.0001 y patience=2).
# + id="-1Jy8yH0SAZ1" colab={"base_uri": "https://localhost:8080/", "height": 470} executionInfo={"status": "ok", "timestamp": 1623323055279, "user_tz": -120, "elapsed": 2088122, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="088b1aca-8240-489e-df01-b4fe6d5b78da"
name = "model_LEDNet_da_wd1_unfreeze"
learn.fit_one_cycle(100,slice(1e-5,1e-4),cbs=[
EarlyStoppingCallback(monitor='valid_loss', min_delta=0.0001, patience=2),
ShowGraphCallback(),
SaveModelCallback(monitor='valid_loss', min_delta=0.0001, fname=name, every_epoch=False)])
# + [markdown] id="wm2sOjfLMxsw"
# Comprobación de los resultados obtenidos en el modelo almacenado.
# + id="Fv61J8enSLZO" colab={"base_uri": "https://localhost:8080/", "height": 67} executionInfo={"status": "ok", "timestamp": 1623323094130, "user_tz": -120, "elapsed": 38861, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="db5c01bb-f79d-4f75-8ae9-37c4fa2f1f0c"
learn.load("model_LEDNet_da_wd1_unfreeze")
learn.validate()
# + [markdown] id="Awp47hCiM5gr"
# # Modelos sin aumento de datos
# Definición del modelo.
# + id="6Gq2k961SLjY"
del model, learn
model = LEDNet(nclass=2)
# + [markdown] id="0uBGjhDlOx-H"
# Creación del Learner con wd=1e-2 y definición del directorio de trabajo.
# + id="T9L8qgsROtj1"
learn = Learner(dls=train2DLS, model=model, metrics=[Dice(), JaccardCoeff()], wd=1e-2)
learn.model_dir = "/content/drive/MyDrive/Colab Notebooks/LEDNet"
# + [markdown] id="sPVbXh7vQBYB"
# Freeze y elección de la tasa de aprendizaje.
# + id="zLDqYHWSO3RB" colab={"base_uri": "https://localhost:8080/", "height": 302} executionInfo={"status": "ok", "timestamp": 1623395342453, "user_tz": -120, "elapsed": 10301, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="6ef2bab3-3b82-468d-c8f6-a711a34802a3"
learn.freeze()
learn.lr_find()
learn.recorder
# + [markdown] id="Eb2OufXiQKk4"
# Entrenamiento del modelo usando EarlyStoppingCallback según el valid_loss (min_delta=0.0001 y patience=2).
# + id="Uw8pvBaeQGJF" colab={"base_uri": "https://localhost:8080/", "height": 520} executionInfo={"status": "ok", "timestamp": 1623398031360, "user_tz": -120, "elapsed": 2688927, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="4dbb55fc-13bb-4e3c-dd62-c07d92e82e98"
name = "model_LEDNet_wd2"
learn.fit_one_cycle(100,slice(1e-5,1e-4),cbs=[
EarlyStoppingCallback(monitor='valid_loss', min_delta=0.0001, patience=2),
ShowGraphCallback(),
SaveModelCallback(monitor='valid_loss', min_delta=0.0001, fname=name, every_epoch=False)])
# + [markdown] id="cOZLre5KQasv"
# Comprobación de los resultados obtenidos en el modelo almacenado.
# + id="7AwsD0deQZq_" colab={"base_uri": "https://localhost:8080/", "height": 68} executionInfo={"status": "ok", "timestamp": 1623398071096, "user_tz": -120, "elapsed": 39773, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="b4606499-5542-4469-9f1d-bca0c985fe19"
learn.load("model_LEDNet_wd2")
learn.validate()
# + [markdown] id="-hmY9IgFRhYk"
# Unfreeze y elección de la tasa de aprendizaje.
# + id="Dyh8TPQTRbZB" colab={"base_uri": "https://localhost:8080/", "height": 334} executionInfo={"status": "ok", "timestamp": 1623399145842, "user_tz": -120, "elapsed": 9974, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="f2e62b03-73e2-4a3a-a3a4-020432c60aff"
learn.unfreeze()
learn.lr_find()
learn.recorder
# + [markdown] id="Dk5qFpyeRoqf"
# Entrenamiento del modelo usando EarlyStoppingCallback según el valid_loss (min_delta=0.0001 y patience=2).
# + id="qu4R8G4QRtfm" colab={"base_uri": "https://localhost:8080/", "height": 424} executionInfo={"status": "ok", "timestamp": 1623400746132, "user_tz": -120, "elapsed": 1600301, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="89ae610d-99b9-4274-b337-ff442aa35beb"
name = "model_LEDNet_wd2_unfreeze"
learn.fit_one_cycle(100,slice(1e-5,1e-4),cbs=[
EarlyStoppingCallback(monitor='valid_loss', min_delta=0.0001, patience=2),
ShowGraphCallback(),
SaveModelCallback(monitor='valid_loss', min_delta=0.0001, fname=name, every_epoch=False)])
# + [markdown] id="sb5sm60hR6TH"
# Comprobación de los resultados obtenidos en el modelo almacenado.
# + id="GYW3GP25R94V" colab={"base_uri": "https://localhost:8080/", "height": 68} executionInfo={"status": "ok", "timestamp": 1623400785892, "user_tz": -120, "elapsed": 39787, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="30bf6bc6-f328-46cf-abcf-06b30dae4101"
learn.load("model_LEDNet_wd2_unfreeze")
learn.validate()
# + [markdown] id="MbQapbl-SMRs"
#
#
# ---
#
#
# Definición del modelo.
# + id="syAmi8DjSJMn" executionInfo={"status": "ok", "timestamp": 1623895207743, "user_tz": -120, "elapsed": 265, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
del model, learn
model = LEDNet(nclass=2)
# + [markdown] id="QtpfSlJlSYxm"
# Creación del Learner con wd=1e-1 y definición del directorio de trabajo.
# + id="KbtGwbprSaUD" executionInfo={"status": "ok", "timestamp": 1623895208771, "user_tz": -120, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
learn = Learner(dls=train2DLS, model=model, metrics=[Dice(), JaccardCoeff()], wd=1e-1)
learn.model_dir = "/content/drive/MyDrive/Colab Notebooks/LEDNet"
# + [markdown] id="44xLotiKSkKE"
# Freeze y elección de la tasa de aprendizaje.
# + id="6Z7nQ77XSkpj" colab={"base_uri": "https://localhost:8080/", "height": 300} executionInfo={"status": "ok", "timestamp": 1623400940019, "user_tz": -120, "elapsed": 9809, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="a7539640-3bc0-4cf2-abb2-dd0cd644863b"
learn.freeze()
learn.lr_find()
learn.recorder
# + [markdown] id="7KO6VtMGSn-Q"
# Entrenamiento del modelo usando EarlyStoppingCallback según el valid_loss (min_delta=0.0001 y patience=2).
# + id="zlVTDlTqSob-" colab={"base_uri": "https://localhost:8080/", "height": 520} executionInfo={"status": "ok", "timestamp": 1623403630457, "user_tz": -120, "elapsed": 2690446, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="90262ae7-681f-44f2-c65b-36a3ac69ffd1"
name = "model_LEDNet_wd1"
learn.fit_one_cycle(100,slice(1e-5,1e-4),cbs=[
EarlyStoppingCallback(monitor='valid_loss', min_delta=0.0001, patience=2),
ShowGraphCallback(),
SaveModelCallback(monitor='valid_loss', min_delta=0.0001, fname=name, every_epoch=False)])
# + [markdown] id="DE8YB1tUS0PJ"
# Comprobación de los resultados obtenidos en el modelo almacenado.
# + id="iLxUqvjDS0sY" colab={"base_uri": "https://localhost:8080/", "height": 68} executionInfo={"status": "ok", "timestamp": 1623403671368, "user_tz": -120, "elapsed": 40938, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="e4b325fe-7dd2-4ebf-ad37-244996f1e2bc"
learn.load("model_LEDNet_wd1")
learn.validate()
# + [markdown] id="V6wt_10ITMcq"
# Unfreeze y elección de la tasa de aprendizaje.
# + id="XGaCuLCcTLvB" colab={"base_uri": "https://localhost:8080/", "height": 300} executionInfo={"status": "ok", "timestamp": 1623403681261, "user_tz": -120, "elapsed": 9904, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="7f649ccb-220c-4504-b6b8-52caff7380f8"
learn.unfreeze()
learn.lr_find()
learn.recorder
# + [markdown] id="VnMeHXStThS_"
# Entrenamiento del modelo usando EarlyStoppingCallback según el valid_loss (min_delta=0.0001 y patience=2).
# + id="diqadjmFTkSf" colab={"base_uri": "https://localhost:8080/", "height": 520} executionInfo={"status": "ok", "timestamp": 1623406403346, "user_tz": -120, "elapsed": 2722105, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="787baab0-f61e-4a2e-9358-658e89ef0258"
name = "model_LEDNet_wd1_unfreeze"
learn.fit_one_cycle(100,slice(1e-5,1e-4),cbs=[
EarlyStoppingCallback(monitor='valid_loss', min_delta=0.0001, patience=2),
ShowGraphCallback(),
SaveModelCallback(monitor='valid_loss', min_delta=0.0001, fname=name, every_epoch=False)])
# + [markdown] id="67QVoL6fTr4b"
# Comprobación de los resultados obtenidos en el modelo almacenado.
# + id="SAGtvEsiTo_4" colab={"base_uri": "https://localhost:8080/", "height": 68} executionInfo={"status": "ok", "timestamp": 1623406443524, "user_tz": -120, "elapsed": 40200, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="7c5c3757-8c98-4ad9-b5a5-0b98bc344d7a"
learn.load("model_LEDNet_wd1_unfreeze")
learn.validate()
# + [markdown] id="cj9X-MtbKRvn"
# # Evaluación de resultados
# ## Modelos con aumento de datos
# Carga del primer modelo en la CPU.
# + id="AKuyu1_yt5L0"
learn.load("model_LEDNet_da_wd2")
aux=learn.model
aux=aux.cpu()
# + [markdown] id="VDmKh0ZaKr1X"
# Asignación del dataloader de test y validación.
# + id="Q7LD1MXouZyn" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1623317196434, "user_tz": -120, "elapsed": 49245, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="c8af898f-dfc9-4635-bda1-bce34ba6ec13"
learn.dls = testDLS
learn.validate()
# + [markdown] id="PJKceuIkLJf2"
# Comparación de resultado buscado contra resultado obtenido.
# + id="uJ2vE-R6IPGB" colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"status": "ok", "timestamp": 1623317197071, "user_tz": -120, "elapsed": 642, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="c112e587-eabd-465d-90de-983888551048"
learn.show_results(vmin=0,vmax=1)
# + [markdown] id="1YLi_u2bLNJF"
# Carga del segundo modelo en la CPU.
# + id="ldxtJLe5LRja"
learn.load("model_LEDNet_da_wd2_unfreeze")
aux=learn.model
aux=aux.cpu()
# + [markdown] id="eUTJHuuKLb27"
# Asignación del dataloader de test y validación.
# + id="Df5FutG9LcmE" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1623317246073, "user_tz": -120, "elapsed": 49007, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="566af9d8-e925-4a41-c4fe-b142630c5569"
learn.dls = testDLS
learn.validate()
# + [markdown] id="QXuIkMg4Lg8z"
# Comparación de resultado buscado contra resultado obtenido.
# + id="nmsvN0UYLkDF" colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"status": "ok", "timestamp": 1623317246906, "user_tz": -120, "elapsed": 839, "user": {"displayName": "<NAME> clase", "photoUrl": "", "userId": "15591564048203513807"}} outputId="30db4ef7-b49f-4c3f-dddf-cf64a1a3392c"
learn.show_results(vmin=0,vmax=1)
# + [markdown] id="R1Oj5GcvNg1H"
#
#
# ---
#
#
# Carga del tercer modelo en la CPU.
# + id="7GEU209lNrk8"
learn.load("model_LEDNet_da_wd1")
aux=learn.model
aux=aux.cpu()
# + [markdown] id="RscvZwjBN0bY"
# Asignación del dataloader de test y validación.
# + id="JlwbedC1N1E-" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1623395202316, "user_tz": -120, "elapsed": 49382, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="66edecea-5fb4-4d9a-fd08-90108d9dff29"
learn.dls = testDLS
learn.validate()
# + [markdown] id="a-QCr_TBN5-v"
# Comparación de resultado buscado contra resultado obtenido.
# + id="bE5GUsVlN5RI" colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"status": "ok", "timestamp": 1623395203057, "user_tz": -120, "elapsed": 763, "user": {"displayName": "<NAME> clase", "photoUrl": "", "userId": "15591564048203513807"}} outputId="1d64f5e3-d3bc-408e-de0f-01cd2e8abd5c"
learn.show_results(vmin=0,vmax=1)
# + [markdown] id="3kex06amN-e0"
# Carga del cuarto modelo en la CPU.
# + id="Qby8KgwNOCSr"
learn.load("model_LEDNet_da_wd1_unfreeze")
aux=learn.model
aux=aux.cpu()
# + [markdown] id="1cQOr385OGgr"
# Asignación del dataloader de test y validación.
# + id="uCEBCHbtOG_B" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1623395254748, "user_tz": -120, "elapsed": 51359, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="74d6b467-1bca-48d5-e7ae-ac86acaa10d7"
learn.dls = testDLS
learn.validate()
# + [markdown] id="b3OibGP1OLvC"
# Comparación de resultado buscado contra resultado obtenido.
# + id="aVGBVu41OKvs" colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"status": "ok", "timestamp": 1623395255202, "user_tz": -120, "elapsed": 476, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="05d564eb-9454-445c-caf1-e192a22e0658"
learn.show_results(vmin=0,vmax=1)
# + [markdown] id="5CQTPz6fOUie"
# ## Modelos sin aumento de datos
# Carga del primer modelo en la CPU.
# + id="PxOvmT1TOTnt"
learn.load("model_LEDNet_wd2")
aux=learn.model
aux=aux.cpu()
# + [markdown] id="sf-vfeIAUPuy"
# Asignación del dataloader de test y validación.
# + id="6PWQrTtyUUiU" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1623400860045, "user_tz": -120, "elapsed": 49391, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="1b220a2e-de92-47d3-e558-c752f072b5b4"
learn.dls = testDLS
learn.validate()
# + [markdown] id="rMLK6UCoUZP3"
# Comparación de resultado buscado contra resultado obtenido.
# + id="nYhpZUxaUZwi" colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"status": "ok", "timestamp": 1623400860639, "user_tz": -120, "elapsed": 625, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="83f41a84-5c7b-45e8-947e-c7a2b2e5c1af"
learn.show_results(vmin=0,vmax=1)
# + [markdown] id="n4u8TaKgUirt"
# Carga del segundo modelo en la CPU.
# + id="RSc-hi3uUgV7" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1623400860643, "user_tz": -120, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="4ed81662-39a2-484d-a0b3-4595dfe82899"
learn.load("model_LEDNet_wd2_unfreeze")
aux=learn.model
aux=aux.cpu()
# + [markdown] id="WtnWlQCyUryx"
# Asignación del dataloader de test y validación.
# + id="aFb_gA2NUqsE" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1623400909877, "user_tz": -120, "elapsed": 49248, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="a82f7830-84aa-488d-dbf5-ab4bb861ece9"
learn.dls = testDLS
learn.validate()
# + [markdown] id="XzHJfjTxUwOc"
# Comparación de resultado buscado contra resultado obtenido.
# + id="9my6vzBvUwrS" colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"status": "ok", "timestamp": 1623400915179, "user_tz": -120, "elapsed": 926, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="2f392d0f-8b46-4b8f-e907-f4dca0815a04"
learn.show_results(vmin=0,vmax=1)
# + [markdown] id="POiIxe_pU3Ag"
#
#
# ---
#
#
# Carga del tercer modelo en la CPU.
# + id="zTw4lqSOU5Vu"
learn.load("model_LEDNet_wd1")
aux=learn.model
aux=aux.cpu()
# + [markdown] id="i1OsA0QBVAx9"
# Asignación del dataloader de test y validación.
# + id="b42uIHmrVADY" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1623406555253, "user_tz": -120, "elapsed": 49921, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="ebebb4a4-6762-487b-bc4e-b6369f0e1877"
learn.dls = testDLS
learn.validate()
# + [markdown] id="20HCrsACVEky"
# Comparación de resultado buscado contra resultado obtenido.
# + id="PylHxmH3VGK_" colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"status": "ok", "timestamp": 1623406555913, "user_tz": -120, "elapsed": 693, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="faf34f4f-82ab-40de-8321-5dcd2bd9b3e3"
learn.show_results(vmin=0,vmax=1)
# + [markdown] id="119dGxozVJQJ"
# Carga del cuarto modelo en la CPU.
# + id="u6_5zmVwVHnb"
learn.load("model_LEDNet_wd1_unfreeze")
aux=learn.model
aux=aux.cpu()
# + [markdown] id="QXiZRQyWVdte"
# Asignación del dataloader de test y validación.
# + id="vWzxfdLAVeLv" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1623406605247, "user_tz": -120, "elapsed": 49358, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="da3a0d39-cd24-425d-f82b-ed0327d002c1"
learn.dls = testDLS
learn.validate()
# + [markdown] id="1BmsnoxDVhk6"
# Comparación de resultado buscado contra resultado obtenido.
# + id="i5raKa_1ViNA" colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"status": "ok", "timestamp": 1623406605817, "user_tz": -120, "elapsed": 601, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="84c89d6d-5e49-4cbf-a1c4-d295f09eb8fa"
learn.show_results(vmin=0,vmax=1)
# + [markdown] id="zcZ8koWYsqc5"
# # Exportación del mejor modelo
# Carga del modelo en la CPU.
# + colab={"base_uri": "https://localhost:8080/", "height": 67} id="sPZxZOz7ss5W" executionInfo={"status": "ok", "timestamp": 1623895303338, "user_tz": -120, "elapsed": 51152, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}} outputId="eb271878-5f2d-486a-de06-1ff15f93cbb7"
learn.load("model_LEDNet_wd1_unfreeze")
learn.dls = testDLS
learn.validate()
# + id="k5qeFiL5sz-4" executionInfo={"status": "ok", "timestamp": 1623895307171, "user_tz": -120, "elapsed": 323, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
aux=learn.model
aux=aux.cpu()
# + [markdown] id="GVW-BfLjs1wB"
# Exportación del modelo mediante torch.jit.trace.
# + id="Rycumtz_s22v" executionInfo={"status": "ok", "timestamp": 1623895310978, "user_tz": -120, "elapsed": 2448, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15591564048203513807"}}
import torchvision.transforms as transforms
img = PILImage.create(path_images/'train/1D2_0.png')
transformer=transforms.Compose([transforms.Resize((50,50)),
transforms.ToTensor(),
transforms.Normalize(
[0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
img=transformer(img).unsqueeze(0)
img=img.cpu()
traced_cell=torch.jit.trace(aux, (img))
traced_cell.save("/content/drive/MyDrive/Colab Notebooks/LEDNet/model_LEDNet.pkl")
| E03 - Learning programs and models/Colab Notebooks/LEDNet/LEDNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 9 - Motor Control
# ### Introduction to modeling and simulation of human movement
# https://github.com/BMClab/bmc/blob/master/courses/ModSim2018.md
# * Tasks (for Lecture 9):
#
# Implement the Kelvin model. Set the parameters of the model so as to the response of the model is similar to the shown in the Figure 3(a) from <NAME>.; <NAME>.; <NAME>. Viscoelastic properties of passive skeletal muscle in compression: Stress-relaxation behaviour and constitutive modelling. Journal of Biomechanics, v. 41, n. 7, p. 1555–1566, 2008.
# +
import numpy as np
#import pandas as pd
#import pylab as pl
import matplotlib.pyplot as plt
import math
# %matplotlib notebook
# +
dt = 5e-2 # samples/s (time resolution)
t = np.arange(0,360,dt)
f = np.empty (t.shape)
# -- Parameters --
ks = 150 #kPa
kp = 350 #kPa
b = 800
l0 = 10e-3 #m
l = 9.99e-3 #m
# Samples were compressed up to 30% strain at rates of 0.5% s^-1
rate = 0.5e-2#/dt # m/ sample
#l = rate*l0
f[0] = 0
# -
# ## Kelvin model
x1 = f[0] - (kp+ks)*l
for i in range (1,len(t)):
if t[i]>=60:
l = 7e-3
#Equation
dx1dt = -(ks/b)*x1 -((ks**2)/b)*l -((ks*kp)/b)*l0
x1 = x1 + dt*dx1dt
f[i] = x1 + (kp+ks)*l
# ## Plot
# +
fig, ax = plt.subplots(1, 1, figsize=(5.2,5.2), sharex=True)
ax.plot(t,f,c='red',label='F0.5% $s^{-1}$')
plt.xlim([0,350])
plt.ylim([-3,0])
plt.grid()
plt.xlabel('time (s)')
plt.ylabel('Cauchy stress(kPa)')
plt.title('F = 0°')
ax.legend()
# -
| courses/modsim2018/tasks/Desiree/Task9_MotorControl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # What's in these flash-sorted files?
#
# The flash sorting process produces binary files. The purpose of this notebook is to demonstrate how to read and display data from these files. In practice, these files are no more difficult to work with than columns of ASCII data, because modern access libraries automate the annoying and tedious parts of traversing the file's internal hierarchy.
#
# The flash sorting analysis produces three kinds of files.
# - HDF5 files with two data tables
# - the VHF sources themselves
# - replicates ASCII data
# - retains single-point flashes
# - information about each flash
# - Gridded NetCDF files
# - Counts up things that are in the HDF5 files
# - source, flash, flash initiation density
# - average flash area (area-weighted flash density)
# - The grids are [CF-compliant](http://cfconventions.org)
# - uses standardized coordinate names and metadata
# - may be automatically read in and displayed in community tools
# - [IDV](http://www.unidata.ucar.edu/software/idv/index.html)
# - [McIDAS-V](http://www.ssec.wisc.edu/mcidas/software/v/)
# - [NOAA's Weather and Climate Toolkit](http://www.ncdc.noaa.gov/wct/)
# - Plots (PDF) of NetCDF grids
# - multi-panel per page, one panel for each time interval
# - very simple, with no maps, etc.
# - useful for a quick reference
#
#
# Additional data tables and gridded fields can be added to accomodate, for instance, VLF/LF stroke detection data, and an associated LMA `flash_id`.
#
#
# + language="bash"
#
# RESULTS="/data/GLM-wkshp/flashsort/results/"
# ls $RESULTS
# echo "-----"
# ls $RESULTS/h5_files/2009/Apr/10/
# echo "-----"
# ls $RESULTS/grid_files/2009/Apr/10/
# echo "-----"
# cat $RESULTS/h5_files/2009/Apr/10/input_params.py
# -
# # Inside the HDF5 files
#
# To look inside the HDF5 file, we'll use the Python library known as PyTables.
#
# The file object itself can be printed to describe the file's contents.
# +
import os
import tables, pandas
import numpy as np
results_dir = "/data/GLM-wkshp/flashsort/results/"
h5name = os.path.join(results_dir, "h5_files/2009/Apr/10/LYLOUT_090410_180000_3600.dat.flash.h5")
h5 = tables.open_file(h5name)
print h5
# -
# The file's internal hierarchy of groups and data tables is easily navigated by tab-completion. In the next cell, type
#
# `h5.root.`
#
# and then tab, and you'll get a list of group and table names.
#
# Let's look at events first. tab-complete to that group and then tab-complete again to get a reference to the data table. Note that a flash_id column has been added to the original source data.
h5.root.
event_table = h5.root.events.LMA_090410_180000_3600
event_table
# We can get all the altitudes ...
event_table.cols.alt[:]
# Or the first ten sources ...
event_table[:10]
# ... or the flash IDs for the first hundred sources.
event_table[:100]['flash_id']
# Let's look at flash #27. The indexing operation (colon) is what actually lets us access the data, instead of just the description of the data column.
print event_table.cols.flash_id
query = (event_table.cols.flash_id[:] == 27)
fl_27_events = event_table[query][:]
lats, lons, alts, times = fl_27_events['lat'], fl_27_events['lon'], fl_27_events['alt'], fl_27_events['time']
for lat, lon in zip(lats, lons):
print lat, lon
# +
import matplotlib
matplotlib.rc('font', size=12)
# %matplotlib inline
import matplotlib.pyplot as plt
plt.scatter(lons, lats, c=times)
plt_range = (lons.min(),lons.max(),lats.min(),lats.max())
print plt_range
plt.axis(plt_range)
# -
# Now, let's get the associated flash properties. Use pandas to make an appealing print of the the data table.
flash_table = h5.root.flashes.LMA_090410_180000_3600
fl_27 = flash_table[flash_table.cols.flash_id[:] == 27] # This is a numpy array with a named dtype
fl_27 = pandas.DataFrame(fl_27)
fl_27
fl_27_events = pandas.DataFrame(fl_27_events)
fl_27_events
# More [sophisticated queries](http://www.pytables.org/usersguide/condition_syntax.html) with the HDF5 data (and pandas) are possible.
# #Inside the NetCDF files
import netCDF4 as ncdf
nc_name = os.path.join(results_dir, "grid_files/2009/Apr/10/NALMA_20090410_180000_3600_10src_0.0109deg-dx_flash_extent.nc")
nc = ncdf.Dataset(nc_name)
print nc
print nc.variables['crs']
from lmatools.multiples_nc import centers_to_edges
matplotlib.rc('xtick', labelsize=10)
matplotlib.rc('ytick', labelsize=10)
lon, lat, fl_dens = nc.variables['longitude'][:], nc.variables['latitude'][:], nc.variables['flash_extent'][:]
lon_edge, lat_edge = centers_to_edges(lon), centers_to_edges(lat)
time_index = 0
plt.pcolormesh(lon_edge, lat_edge, fl_dens[time_index,:,:], cmap='gray_r')
cbar = plt.colorbar()
# ## Saving a custom animation
#
# The `lmatools` package includes a way to access these grids as a collection. We will use the collection to make a series of frames for an animation. This example is adapted from a notebook included in `lmatools`.
# +
import glob
from lmatools.grid_collection import LMAgridFileCollection
nc_filenames=glob.glob(os.path.join(results_dir, "grid_files/2009/Apr/10/*1[89]00*_flash_extent.nc"))
print nc_filenames
nc_field = 'flash_extent'
NCs = LMAgridFileCollection(nc_filenames, nc_field, x_name='longitude', y_name='latitude')
# -
from datetime import datetime
t = datetime(2009,4,10,18,30,00)
xedge, yedge, data = NCs.data_for_time(t)
limits = xedge.min(), xedge.max(), yedge.min(), yedge.max()
fig = plt.figure(figsize=(12,12))
if False: # set this to True to actually write files.
for t, lon, lat, data in NCs:
ax = plt.subplot(111)
mesh = ax.pcolormesh(lon, lat, np.log10(data), vmin=0, vmax=3, cmap='gray_r')
ax.axis(limits)
plt.colorbar(mesh)
outfile = "{0}_{1}.png".format(t.isoformat(), nc_field)
save_path = os.path.join(results_dir, outfile)
print(save_path)
fig.savefig(save_path)
fig.clf()
# ## Interactive animation
#
# Even better, we can use the notebook's widget system to make an interactive version of the above.
from IPython.html.widgets import interactive
from IPython.display import display
# +
from matplotlib.colors import LogNorm
def plot_for_frame(frame=0):
# get the data
t = NCs.times[frame]
xedge, yedge, data = NCs.data_for_time(t)
# plot a frame of data
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111)
mesh = ax.pcolormesh(xedge, yedge, data,
vmin=1, vmax=100, norm=LogNorm(), cmap='gray_r')
ax.axis(limits)
plt.colorbar(mesh, ax=ax)
title = "{1} at {0}".format(t.isoformat(), nc_field)
ax.set_title(title)
return fig
N_frames = len(NCs.times)
w = interactive(plot_for_frame, frame=(0, N_frames-1))
display(w)
# -
| GLM-AL-2015/Reading the flash-sorted files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import json
import requests
from keys import get_keys, get_endpoints
keys = get_keys()
endpoints = get_endpoints()
print(endpoints.keys())
# -
# # Text Analytics
# +
headers = {
'Ocp-Apim-Subscription-Key': keys['text'],
}
documents = { 'documents' : [
{'id': '1', 'language': 'en', 'text': 'I had a wonderful experience! The tuna tartare was divine!'},
{'id': '2', 'language': 'en', 'text': 'I had a terrible time at the hotel. The staff was rude and the food was awful.'},
{'id': '3', 'language': 'es', 'text': 'La verdad es que no me gusta mucho la psicina. Tod estaaba muy sucio.'}
]}
# -
response = requests.post(endpoints['keyphrases'], headers=headers, json=documents)
o = json.dumps(response.json(), indent=2, sort_keys=True)
print(o)
| notebooks/text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# First, create the model. This must match the model used in the interactive training notebook.
# +
import torch
import torchvision
CATEGORIES = ['apex']
device = torch.device('cuda')
model = torchvision.models.resnet18(pretrained=False)
model.fc = torch.nn.Linear(512, 2 * len(CATEGORIES))
model = model.cuda().eval().half()
# -
# Next, load the saved model. Enter the model path you used to save.
model.load_state_dict(torch.load('road_following_model.pth'))
# Convert and optimize the model using ``torch2trt`` for faster inference with TensorRT. Please see the [torch2trt](https://github.com/NVIDIA-AI-IOT/torch2trt) readme for more details.
#
# > This optimization process can take a couple minutes to complete.
# +
from torch2trt import torch2trt
data = torch.zeros((1, 3, 224, 224)).cuda().half()
model_trt = torch2trt(model, [data], fp16_mode=True)
# -
# Save the optimized model using the cell below
torch.save(model_trt.state_dict(), 'road_following_model_trt.pth')
# Load the optimized model by executing the cell below
# +
import torch
from torch2trt import TRTModule
model_trt = TRTModule()
model_trt.load_state_dict(torch.load('road_following_model_trt.pth'))
# -
# Create the racecar class
# +
from jetracer.nvidia_racecar import AIRacecar
car = AIRacecar()
# -
# Create the camera class.
# +
from jetcam.csi_camera import CSICamera
camera = CSICamera(width=224, height=224, capture_fps=65)
# -
# Finally, execute the cell below to make the racecar move forward, steering the racecar based on the x value of the apex.
#
# Here are some tips,
#
# * If the car wobbles left and right, lower the steering gain
# * If the car misses turns, raise the steering gain
# * If the car tends right, make the steering bias more negative (in small increments like -0.05)
# * If the car tends left, make the steering bias more postive (in small increments +0.05)
# +
from utils import preprocess
import numpy as np
STEERING_GAIN = 0.75
STEERING_BIAS = 0.00
car.throttle = 0.15
while True:
image = camera.read()
image = preprocess(image).half()
output = model_trt(image).detach().cpu().numpy().flatten()
x = float(output[0])
car.steering = x * STEERING_GAIN + STEERING_BIAS
| notebooks/road_following.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 2.889826, "end_time": "2021-10-14T06:50:10.304247", "exception": false, "start_time": "2021-10-14T06:50:07.414421", "status": "completed"} tags=[]
import pandas as pd
import numpy as np
import lightgbm as lgb
import optuna
import pickle
import os
import gc
from sklearn.preprocessing import MinMaxScaler
from matplotlib import pyplot as plt
import seaborn as sns
import time
import json
import optuna
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_auc_score
from utilities import (
RANDOM_STATE, TARGET_COL, N_FOLD,
)
INPUT_PATH = '../input/tabular-playground-series-oct-2021'
PATH_NOTEBOOK = '../input/preprocess'
# + papermill={"duration": 13.51891, "end_time": "2021-10-14T06:50:23.835952", "exception": false, "start_time": "2021-10-14T06:50:10.317042", "status": "completed"} tags=[]
train = pd.read_pickle(
os.path.join(PATH_NOTEBOOK, 'train_unscaled.pkl')
)
# + papermill={"duration": 0.024767, "end_time": "2021-10-14T06:50:23.870967", "exception": false, "start_time": "2021-10-14T06:50:23.846200", "status": "completed"} tags=[]
with open(os.path.join(PATH_NOTEBOOK, 'feature_dic.pkl'), 'rb') as file:
feature_dic = pickle.load(file)
# + papermill={"duration": 0.207445, "end_time": "2021-10-14T06:50:24.088679", "exception": false, "start_time": "2021-10-14T06:50:23.881234", "status": "completed"} tags=[]
#CONSTANT
FEATURE = feature_dic['feature']
CAT_COL = feature_dic['categorical']
NUMERIC_COL = feature_dic['numerical']
FOLD_LIST = list(range(N_FOLD))
gc.collect()
# + papermill={"duration": 6.121201, "end_time": "2021-10-14T06:50:30.220498", "exception": false, "start_time": "2021-10-14T06:50:24.099297", "status": "completed"} tags=[]
#train test split for optuna-study
train_x, test_x, train_y, test_y = train_test_split(
train[FEATURE], train[TARGET_COL], random_state = RANDOM_STATE,
stratify = train[TARGET_COL], test_size = .75
)
gc.collect()
# + papermill={"duration": 0.02679, "end_time": "2021-10-14T06:50:30.257938", "exception": false, "start_time": "2021-10-14T06:50:30.231148", "status": "completed"} tags=[]
def objective(trial):
dtrain = lgb.Dataset(
train_x, label=train_y,
categorical_feature=CAT_COL
)
dtest = lgb.Dataset(
test_x, label=test_y,
categorical_feature=CAT_COL
)
params_study = {
'objective': 'binary',
'boosting_type': 'gbdt',
'metric': 'auc',
'learning_rate': 0.1,
'random_state': RANDOM_STATE,
'verbose': -1,
'n_jobs': -1,
"num_leaves": trial.suggest_int("num_leaves", 2**6, 2**10),
"bagging_fraction": trial.suggest_float("bagging_fraction", .3, 1.0),
"bagging_freq": trial.suggest_int("bagging_freq", 1, 5),
"feature_fraction": trial.suggest_float("feature_fraction", .3, 1.0),
"lambda_l1": trial.suggest_float("lambda_l1", 1e-8, 10.0),
"lambda_l2": trial.suggest_float("lambda_l2", 1e-8, 10.0),
"min_data_in_leaf": trial.suggest_int("min_data_in_leaf", 1, 100),
"extra_trees": trial.suggest_categorical("extra_trees", [False, True]),
"min_gain_to_split": trial.suggest_float("min_gain_to_split", 1e-8, 10),
"path_smooth": trial.suggest_float("path_smooth", 0, 2),
}
pruning_callback = optuna.integration.LightGBMPruningCallback(
trial, "auc"
)
model = lgb.train(
params_study, dtrain,
valid_sets = dtest,
verbose_eval = False,
callbacks = [pruning_callback],
num_boost_round = 100000,
early_stopping_rounds = 100,
)
preds = model.predict(test_x)
auc = roc_auc_score(test_y, preds)
return auc
# + papermill={"duration": 30502.753184, "end_time": "2021-10-14T15:18:53.022230", "exception": false, "start_time": "2021-10-14T06:50:30.269046", "status": "completed"} tags=[]
study = optuna.create_study(
pruner=optuna.pruners.MedianPruner(n_warmup_steps=10),
direction="maximize",
)
study.optimize(objective, timeout=30500, show_progress_bar = False)
# + papermill={"duration": 1.324088, "end_time": "2021-10-14T15:18:55.675772", "exception": false, "start_time": "2021-10-14T15:18:54.351684", "status": "completed"} tags=[]
best_score = study.best_trial.values
print(best_score)
# + papermill={"duration": 1.337762, "end_time": "2021-10-14T15:18:58.349748", "exception": false, "start_time": "2021-10-14T15:18:57.011986", "status": "completed"} tags=[]
final_params = study.best_trial.params
print(final_params)
# + papermill={"duration": 1.329288, "end_time": "2021-10-14T15:19:00.998272", "exception": false, "start_time": "2021-10-14T15:18:59.668984", "status": "completed"} tags=[]
with open("final_xgb_param.pkl", "wb") as file_name:
pickle.dump(final_params, file_name)
| Tabular Playground Series - Oct 2021/optuna/optuna-lightgbm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Estimation of the number of quiescent cells
# Adding intra and interexperimental error. In the future maybe better do bootstrapping.
import pandas as pd
import scipy as sp
from scipy import stats
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
exec(open('settings.py').read(), globals())
cell_numbers = pd.read_csv('../data/cell_number_data.csv')
outgrowth = pd.read_csv('../data/outgrowth.csv')
lcell = 13.2
lcell_sem = 0.1
L0 = 800.0
cell_numbers = cell_numbers.query('pos > -@L0')
cell_numbers['quiescent'] = cell_numbers['SOX2'] - cell_numbers['PCNA']
cell_numbers_space_mean = cell_numbers[['ID', 'time', 'quiescent']].groupby('ID').agg({'time': ['mean'], 'quiescent': ['mean', 'sem']})
cell_numbers_mean = cell_numbers_space_mean.groupby([('time', 'mean')]).agg({('quiescent', 'mean'): ['mean', 'sem'], ('quiescent', 'sem'): ['mean']})
cell_numbers_mean['quiescent_delta'] = cell_numbers_mean['quiescent', 'mean', 'sem'] + cell_numbers_mean['quiescent', 'sem', 'mean']
cell_numbers_mean.index.name = 'time'
cell_numbers_mean['outgrowth'] = outgrowth.groupby('time').mean()['length']
cell_numbers_mean['outgrowth_sem'] = outgrowth.groupby('time').sem()['length']
for i, row in cell_numbers_space_mean.iterrows():
cell_numbers_space_mean.loc[i, 'outgrowth'] = float(outgrowth.groupby('time').mean().loc[int(row['time'])])
cell_numbers_space_mean.loc[i, 'outgrowth_sem'] = float(outgrowth.groupby('time').sem()['length'].loc[int(row['time'])])
cell_numbers_space_mean
Nq = pd.DataFrame(index = cell_numbers_mean.index)
Nq['mean'] = (cell_numbers_mean['outgrowth'] + L0) / lcell * cell_numbers_mean['quiescent', 'mean', 'mean']
Nq['delta'] = Nq['mean'] * (lcell_sem / lcell + cell_numbers_mean['outgrowth_sem'] / cell_numbers_mean['outgrowth'] + cell_numbers_mean['quiescent', 'mean', 'sem'] / cell_numbers_mean['quiescent', 'mean', 'mean'])
Nq.loc[0, 'delta'] = Nq.loc[0, 'mean'] * ((lcell_sem / lcell + float(cell_numbers_mean.loc[0, ('quiescent', 'mean', 'sem')]) / cell_numbers_mean.loc[0, ('quiescent', 'mean', 'mean')]))
Nq
# ## Check for significant differences
# Let $N_q$ be the total number of quiescent cells in the spatial zone reaching from $-800\mu m$ to the posterior tip of the spinal cord and $l_{cell}$ the anteroposterior length of the cells, $N_{q,s}$ the mean number of quiescent cells per cross section and $L$ the outgrowth of the spinal cord and $L_0$ the source zone length. Then, the following euqation holds:
# \begin{align}
# N_q = N_{q,s} \frac{L + L_0}{l_{cell}}
# \end{align}
# As $l_{cell}$ is constant (<NAME>, Tazaki et al., eLife, 2015) we test if
# \begin{align}
# N_q \cdot l_{cell} = N_{q,s} (L + L_0)
# \end{align}
# is significantly different to infer if $N_q$ is significantly different.
day = dict()
for time in Nq.index:
day[time] = sp.array(cell_numbers_space_mean[cell_numbers_space_mean['time', 'mean'] == time]['quiescent', 'mean'])
day[time] = day[time] * (float(outgrowth.groupby('time').mean().loc[time]) + L0)
# ttest vs day0
for time in Nq.index:
print 'Day 0 vs Day {0}: p = {1:.2f}'.format(time, sp.stats.ttest_ind(day[0], day[time]).pvalue)
# ## Plot
# +
from matplotlib.markers import TICKDOWN
def significance_bar(start,end,height,displaystring,linewidth = 1,markersize = 3,boxpad =0.3,fontsize = 12,color = 'k'):
# draw a line with downticks at the ends
plt.plot([start,end],[height]*2,'-',color = color,lw=linewidth,marker = TICKDOWN,markeredgewidth=linewidth,markersize = markersize)
# draw the text with a bounding box covering up the line
plt.text(0.5*(start+end),height,displaystring,ha = 'center',va='center',size = fontsize)
# +
fig, ax = plt.subplots(1, figsize = (1, 31.5/25.0))
ax.errorbar(Nq.index,\
Nq['mean'],
Nq['delta'],
fmt = 'ko', ls = '-')
significance_bar(0, 4, 305.0, '*')
significance_bar(0, 6, 340.0, '*')
ax.set_xlim(-0.7, 8.7)
ax.set_ylim(0, 370)
ax.set_yticks(sp.arange(0, 350, 100))
ax.set_xlabel('Time (days)')
ax.set_ylabel('# SOX2+/PCNA- cells', labelpad = 8)
plt.savefig('../figure_plots/Fig2_total_number_quiescent.svg')
plt.show()
# -
| calculations/number_quiescent_cells.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# 构建模型
from models.network_swinir import SwinIR as net
import torch
# 灰度图像降噪
model = net(
upscale=1,
in_chans=1,
img_size=128,
window_size=8,
depths=[6, 6, 6, 6, 6, 6],
embed_dim=180,
num_heads=[6, 6, 6, 6, 6, 6],
mlp_ratio=2
)
param_key_g = "params"
pretrained_model = torch.load(
# todo 这里替换成SwinIR_N2V.pth
"model_zoo/swinir/004_grayDN_DFWB_s128w8_SwinIR-M_noise15.pth"
)
model.load_state_dict(
pretrained_model[param_key_g]
if param_key_g in pretrained_model.keys()
else pretrained_model,
strict=True,
)
# +
import numpy as np
from skimage.metrics import structural_similarity, peak_signal_noise_ratio
def predict(img_lq, model, window_size=8,tile=256,tile_overlap=32):
# test the image tile by tile
# 格式NCHW
_,_, h, w = img_lq.size()
tile = min(tile, h, w)
assert tile % window_size == 0, "tile size should be a multiple of window_size"
tile_overlap = tile_overlap
stride = tile - tile_overlap
h_idx_list = list(range(0, h-tile, stride)) + [h-tile]
w_idx_list = list(range(0, w-tile, stride)) + [w-tile]
E = torch.zeros_like(img_lq)
W = torch.zeros_like(E)
for i,h_idx in enumerate(h_idx_list):
for j,w_idx in enumerate(w_idx_list):
# 进度条
print(f"\rprocessing {j+1+i*len(w_idx_list)}/{len(h_idx_list)*len(w_idx_list)} tile",end=" ")
in_patch = img_lq[..., h_idx:h_idx+tile, w_idx:w_idx+tile]
out_patch = model(in_patch)
out_patch_mask = torch.ones_like(out_patch)
E[..., h_idx:(h_idx+tile), w_idx:(w_idx+tile)].add_(out_patch)
W[..., h_idx:(h_idx+tile), w_idx:(w_idx+tile)].add_(out_patch_mask)
output = E.div_(W)
return output
def psnr_ssim(img_gt, img_lq):
assert img_lq.ndim==3, "输入格式应为NHW"
psnr_result=np.zeros(len(img_gt))
ssim_result=np.zeros(len(img_gt))
for i in range(len(img_gt)):
psnr_result[i]=peak_signal_noise_ratio(img_gt[i],img_lq[i])
ssim_result[i]=structural_similarity(img_gt[i],img_lq[i])
return psnr_result.mean(),ssim_result.mean()
# +
import skimage.io as io
from time import time
from math import ceil
UINT16=65535
# 读取数据
img_lq=io.imread("../FYP/dataset/actin-60x-noise1-lowsnr.tif")
# 必须写zeros_like才能正常保存成tif文件
output=np.zeros_like(img_lq)
# 格式NHW->NCHW
# dtype: uint16->float32
device=torch.device("cuda" if torch.cuda.is_available() else 'cpu')
img_lq=torch.from_numpy(img_lq.astype("float32")).unsqueeze(1).to(device)
# 预测、计时
model.eval()
model.to(device)
batch_size=1
batch_number=1
# batch_size=5
# batch_number=ceil(img_lq.shape[0]/batch_size)
with torch.no_grad():
# pad input image to be a multiple of window_size
window_size=8
_, _, h_old, w_old = img_lq.size()
h_pad = ceil(h_old/window_size) * window_size - h_old
w_pad = ceil(w_old/window_size) * window_size - w_old
# if not (h_pad==0 and w_pad==0):
# img_lq = torch.cat([img_lq, torch.flip(img_lq, [2])], 2)[:, :, :h_old + h_pad, :]
# img_lq = torch.cat([img_lq, torch.flip(img_lq, [3])], 3)[:, :, :, :w_old + w_pad]
print("开始".center(20,'-'))
print(f"batch_size:{batch_size}")
t0=time()
for i in range(batch_number):
print(f"\n#batch: {i+1}/{batch_number}")
batch=img_lq[i*batch_size:(i+1)*batch_size]
if h_pad!=0:
batch = torch.cat([batch, torch.flip(batch, [2])], 2)[:, :, :h_old + h_pad, :]
if w_pad!=0:
batch = torch.cat([batch, torch.flip(batch, [3])], 3)[:, :, :, :w_old + w_pad]
# !直接对uint16大小的数字做处理, 取值范围0-65535
output[i*batch_size:(i+1)*batch_size]=predict(batch,model,window_size).squeeze(1).round()[..., :h_old, :w_old].cpu().numpy()# 格式NCHW -> NHW
# 处理每张图片的平均耗时
# t=(time()-t0)/len(img_lq)
t=(time()-t0)
print("\n"+"结束".center(20,'-'))
print(f"处理每张图片的平均用时:{t:.2f}s")
# 保存图片
io.imsave("results/actin-60x-noise1-restored.tif",output[:batch_size*batch_number,...])
# 计算psnr和ssim
img_gt=io.imread("../FYP/dataset/actin-60x-noise1-highsnr.tif")
# psnr,ssim=psnr_ssim(img_gt,output)
# print(f"psnr:{psnr:.2f}")
# print(f"ssim:{ssim:.2f}")
# +
from torchvision.transforms import ToTensor
img_lq=img_lq[0].cpu().numpy().astype("uint16")
psnr,ssim=psnr_ssim(img_gt[0:1],output[0:1])
print("resotred:".center(20,'-'));print(f"psnr:{psnr:.2f}",end="\t");print(f"ssim:{ssim:.2f}")
psnr,ssim=psnr_ssim(img_gt[0:1],img_lq[0:1])
print("raw:".center(20,'-'));print(f"psnr:{psnr:.2f}",end="\t");print(f"ssim:{ssim:.2f}")
# +
import matplotlib.pyplot as plt
plt.figure(figsize=(14,7))
plt.subplot(131);plt.axis("off");plt.title("raw")
plt.imshow(img_lq[0],cmap="gray")
plt.subplot(132);plt.axis("off");plt.title("ground truth")
plt.imshow(img_gt[0],cmap="gray")
plt.subplot(133);plt.axis("off");plt.title("restored")
plt.imshow(output[0],cmap="gray")
plt.show()
plt.close()
| SwinIR_graydn_uint16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We have artificial data for 50 startup (R&D spendings, Adminstration spendings, Marketing spendings, location/state, profit).
# We have introduced a column with random values to simualte the effect of noise in data.
# The State feature has categorical data (Names of states).
# In this notebook, we will try:
# 1. Feature scaling.
# 2. Feature standardization.
# 3. Feature selection using backward elimination method.
# A linear regression model is fit for the data after applying each one of the above methods.
# Different models are comapred in terms of (MAE, MSE, RMSE, R-Sq, R-Sq-Adj).
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error,mean_squared_error,r2_score
from sklearn.preprocessing import LabelEncoder, OneHotEncoder, StandardScaler, minmax_scale
from sklearn.compose import ColumnTransformer
# %matplotlib inline
# -
df = pd.read_csv('50_Startups.csv', delimiter = ';')
df.head()
#Summary of all numerical features
df.describe()
#Summary of the categorical feature
df.State.describe()
#Check for Null value
df['R&D Spend'].isnull().values.any(),df['Administration'].isnull().values.any(), df['Marketing Spend'].isnull().values.any(),df['State'].isnull().values.any()
#Fill null values with the mean value of each column
df['R&D Spend'] = df['R&D Spend'].fillna(df['R&D Spend'].mean())
df['Administration'] = df['Administration'].fillna(df['Administration'].mean())
df['Marketing Spend'] = df['Marketing Spend'].fillna(df['Marketing Spend'].mean())
df.info()
sns.pairplot(df)
#Adding a new column with random numbers to simulate noise in the data
# X variable represents the features
# Y variable represents the response
rand = pd.DataFrame(np.random.randint(1,1000,(df.shape[0],1)),columns=['random'])
x = pd.concat([df.iloc[:,:-1] , rand], axis=1)
y = df[['Profit']]
x.shape, y.shape
x.head()
#Reordering the coulmns of x
cols = x.columns.tolist()
cols_reordered = [cols[3]] + cols[:3] + [cols[4]]
x = x[cols_reordered]
x.head()
#Converting categorical non-numeric feature to numerical feature
le = LabelEncoder()
le.fit(x.State)
le.classes_
x.State = le.transform(x.State)
x.head()
#Split the new categorical numeric feature into 3 binary columns (0, 1)
ohe = OneHotEncoder(categorical_features=[0])
x = ohe.fit_transform(x.values).toarray()
#Creating a new dataframe for the sparse matrix
cols2 = le.classes_.tolist() + cols_reordered[1:]
x = pd.DataFrame(x,columns=cols2)
x.shape, y.shape
#Drop one column from the three binary columns recently added (Dummy variable)
x = x.drop(columns=['California'])
# # Linear Regression with original data
# Data has features of different scales, but we will fit a linear regression model without scaling and calculate some metrics (MAE, MSE, RMSE, R-Sq, R-Sq-Adj).
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.2)
lm1 = LinearRegression()
lm1.fit(x_train,y_train)
lm1.intercept_, lm1.coef_
y_pred1 = lm1.predict(x_test)
MAE1, MSE1, R_Sq1, RMSE1 = mean_absolute_error(y_test,y_pred1), mean_squared_error(y_test,y_pred1), r2_score(y_test,y_pred1), np.sqrt(mean_squared_error(y_test,y_pred1))
# R-Sq-Adjusted formula:
# 
# where:
# n: number of examples.
# k: number of features.
n = y_test.shape[0]
p = y_test.shape[1]
R_Sq_adj1 = 1 - (1 - R_Sq1) * ((n - 1)/(n - p - 1))
print('MAE: {}, MSE: {}, R-Sq: {}, RMSE: {}, R-Sq-Adj: {}'.format(round(MAE1,3), round(MSE1,3), round(R_Sq1,3), round(RMSE1,3), round(R_Sq_adj1,3)))
# # Linear Regression with Features Scaling
x_sc = minmax_scale(x)
y_sc = minmax_scale(y)
x_train_sc, x_test_sc, y_train_sc, y_test_sc = train_test_split(x_sc, y_sc, test_size = 0.2)
lm2 = LinearRegression()
lm2.fit(x_train_sc,y_train_sc)
lm2.intercept_, lm2.coef_
y_pred2 = lm2.predict(x_test_sc)
MAE2, MSE2, R_Sq2, RMSE2 = mean_absolute_error(y_test_sc,y_pred2), mean_squared_error(y_test_sc,y_pred2), r2_score(y_test_sc,y_pred2), np.sqrt(mean_squared_error(y_test_sc,y_pred2))
R_Sq_adj2 = 1 - (1 - R_Sq2) * ((n - 1)/(n - p - 1))
print('MAE: {}, MSE: {}, R-squared: {}, RMSE: {}, R-Sq-Adj: {}'.format(round(MAE2,3), round(MSE2,3), round(R_Sq2,3), round(RMSE2,3), round(R_Sq_adj2,3)))
# # Linear Regression with Feature standardization
x_scaler = StandardScaler()
y_scaler = StandardScaler()
x_st = x_scaler.fit_transform(x)
y_st = y_scaler.fit_transform(y)
x_train_st, x_test_st, y_train_st, y_test_st = train_test_split(x_st, y_st, test_size = 0.2)
lm3 = LinearRegression()
lm3.fit(x_train_st,y_train_st)
lm3.intercept_, lm3.coef_
y_pred3 = lm3.predict(x_test_st)
MAE3, MSE3, R_Sq3, RMSE3 = mean_absolute_error(y_test_st,y_pred3), mean_squared_error(y_test_st,y_pred3), r2_score(y_test_st,y_pred3), np.sqrt(mean_squared_error(y_test_st,y_pred3))
R_Sq_adj3 = 1 - (1 - R_Sq3) * ((n - 1)/(n - p - 1))
print('MAE: {}, MSE: {}, R-squared: {}, RMSE: {}, R-Sq-Adj: {}'.format(round(MAE3,3), round(MSE3,3), round(R_Sq3,3), round(RMSE3,3), round(R_Sq_adj3,3)))
# # Features selection using Backward Elimination Method
#Standardized data will be used
#Create data frames for features and reponse data.
x_df = pd.DataFrame(x_st,columns= x.columns.tolist())
y_df = pd.DataFrame(y_st,columns=['Profit'])
x_df.head()
import statsmodels.api as sm
sl = 0.05
# Add the constant to features since statsmodels.OLS doesn't add them.
x_df = sm.add_constant(x_df)
x_df.head()
# Function to automate the feature elimination process
# In each iteration, the faeture with max p-value & its p-value is greater than significance level, 0.05
# is eliminated
#The function returns the statistically significant features and the fitted OLS model.
def feature_eliminate(x, y):
a = False
while a is False:
regressor = sm.OLS(endog= y , exog= x).fit()
for i in regressor.pvalues.values:
if regressor.pvalues.values.max() > 0.05:
if i == regressor.pvalues.values.max():
drop_feature = regressor.pvalues[regressor.pvalues == i].index[0]
x = x.drop(columns = drop_feature)
break
else:
a = True
break
return x, regressor
features_significant, model = feature_eliminate(x_df,y_df)
model.summary()
# # Linear Regression for statistically significant features.
x_train_selected, x_test_selected, y_train_selected, y_test_selected = train_test_split(features_significant, y_df, test_size = 0.2)
lm4 = LinearRegression()
lm4.fit(x_train_selected,y_train_selected)
lm4.intercept_, lm4.coef_
y_pred4 = lm4.predict(x_test_selected)
MAE4, MSE4, R_Sq4, RMSE4 = mean_absolute_error(y_test_selected,y_pred4), mean_squared_error(y_test_selected,y_pred4), r2_score(y_test_selected,y_pred4), np.sqrt(mean_squared_error(y_test_selected,y_pred4))
R_Sq_adj4 = 1 - (1 - R_Sq4) * ((n - 1)/(n - p - 1))
print('MAE: {}, MSE: {}, R-squared: {}, RMSE: {}, R-Sq-Adj: {}'.format(round(MAE4,3), round(MSE4,3), round(R_Sq4,3), round(RMSE4,3), round(R_Sq_adj4,3)))
# # Models Comparison
pd.DataFrame([[MAE1, MSE1, R_Sq1, RMSE1, R_Sq_adj1],[MAE2, MSE2, R_Sq2, RMSE2, R_Sq_adj2],[MAE3, MSE3, R_Sq3, RMSE3, R_Sq_adj3],[MAE4, MSE4, R_Sq4, RMSE4, R_Sq_adj4]], columns=['MAE','MSE','R-Sq','RMSE','R-Sq-Adj'], index= ['Original_Data','Scaled_Data','Standardized_Data', 'Stat_Sign_Features'])
| 50-Startups-Linear-Regression-Feature-Engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
output = 'This is an easy task :)'
print output
# -
text1 = """Why should you learn to write programs? 7746
12 1929 8827
Writing programs (or programming) is a very creative
7 and rewarding activity. You can write programs for
many reasons, ranging from making your living to solving
8837 a difficult data analysis problem to having fun to helping 128
someone else solve a problem. This book assumes that
everyone needs to know how to program ..."""
# +
import re
result_list = re.findall('[0-9]+', text1)
sum(map(lambda x: int(x), result_list))
# +
import urllib2 # the lib that handles the url stuff
import re
target_url = 'http://python-data.dr-chuck.net/regex_sum_42.txt'
data = urllib2.urlopen(target_url)
def lineToSum(line):
strList = re.findall('[0-9]+', line)
intList = map(lambda x: int(x), strList)
return sum(intList)
sum(map(lineToSum, data))
# +
import urllib2 # the lib that handles the url stuff
import re
target_url = 'http://python-data.dr-chuck.net/regex_sum_288663.txt'
data = urllib2.urlopen(target_url)
def lineToSum(line):
strList = re.findall('[0-9]+', line)
intList = map(lambda x: int(x), strList)
return sum(intList)
sum(map(lineToSum, data))
# -
import urllib2 as urllib
from bs4 import BeautifulSoup
url = 'https://docs.python.org/2/library/functions.html'
html = urllib.urlopen(url).read()
soup = BeautifulSoup(html, "lxml")
tags = soup('a')
for tag in tags:
print tag.get('href', None)
# +
# Homework week4 1:
import urllib2 as urllib
from bs4 import BeautifulSoup
url = 'http://python-data.dr-chuck.net/comments_288668.html'
html = urllib.urlopen(url).read()
soup = BeautifulSoup(html, "lxml")
tags = soup('span')
sum(map(lambda x1: int(x1.contents[0]), tags))
# +
# Homework week4 2:
import urllib2 as urllib
from bs4 import BeautifulSoup
url = 'http://python-data.dr-chuck.net/known_by_Mahdi.html'
for i in range(7):
html = urllib.urlopen(url).read()
soup = BeautifulSoup(html, "lxml")
tags = soup('a')
url = tags[17].get('href', None)
print url
# -
| Week1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from matplotlib import pyplot as plt
import numpy as np
import seaborn as sns
import pandas as pd
from scipy.stats.distributions import poisson,norm
def simulate_pixel(photon_rate, readout_noise, samples=1):
"""
Simulate the output of a detector pixel based on
- The `photon_rate` in photons per exposure time
- The pixel `readout_noise` in photons
This implements a very simple model where the counts
follow the Poisson distribution, readout noise is
normally distributed, and the outputs are digitized.
"""
#Draw the number of photon events from a Poisson distribution
true_counts = poisson.rvs(photon_rate, size=samples)
# Observed_counts before digitization with normal readout noise
observed_counts = norm.rvs(loc=true_counts, scale=readout_noise, size=samples)
return observed_counts
# +
rates = [10., 100., 1000.]
#rates = [0., 1.]
multiplicity = 5
readout_noises = [0., 2.]
df = None
for readout_noise in readout_noises:
for rate in rates:
_df = pd.DataFrame({
'Counts (photons / exposure)' : simulate_pixel(rate, readout_noise=readout_noise, samples=multiplicity),
'Readout Noise (photons)' : readout_noise,
'Ground Truth Rate' : rate,
})
df = pd.concat((df, _df))
sns.swarmplot(
x='Ground Truth Rate',
y='Counts (photons / exposure)',
hue='Readout Noise (photons)',
data=df,
palette='Set2',
dodge=True
)
| HW3/counting_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Classifying stars
#
# Processed and balanced data obtained from [Kaggle](https://www.kaggle.com/vinesmsuic/star-categorization-giants-and-dwarfs)
# I will used this data set to test out the XGBoost classifier. Information on the dataset is as follows:
#
# Stellar Classification uses the spectral data of stars to categorize them
# into different categories.
#
# The modern stellar classification system is known as the Morgan–Keenan (MK)
# classification system. It uses the old HR classification system to categorize
# stars with their chromaticity and uses Roman numerals to categorize the star’s
# size.
#
# In this Dataset, we will be using Absolute Magnitude and B-V Color Index to
# Identify Giants and Dwarfs (`TargetClass`).
# Classifying stars based on their `TargetClass` as being either:
# - Star is a Dwarf (0)
# - Star is a Giant (1)
# Note: This is a good dataset for learning about XGBoost, but it is not a good problem to solve with XGBoost as the number of features are relatively small and the problem is a simple binary classification problem.
# +
#load libraries
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OneHotEncoder
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import RandomForestClassifier
from sklearn.dummy import DummyClassifier
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
# +
#read in zip file
star_df = pd.read_csv('data/Star39552_balanced.csv.zip')
# -
# # Quick EDA
star_df.describe(include='all')
# All numeric apart from `SPtype` and `TargetClass` (which is numeric but only 0 or 1)
#check missing
star_df.isnull().sum()
# pairs plot to have a look at the data
sns.pairplot(star_df,
hue = 'TargetClass',
palette = 'flare',
corner = True)
# `Pix` and `e_Pix` look like they could do with being log transformed. For this initial run I am not going to do this.
#
# B-V is binomial for `TargetClass`= 1 (giants)
corr = star_df.corr()
corr_plot = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
corr_plot.set_xticklabels(
corr_plot.get_xticklabels(),
rotation=45,
horizontalalignment='right'
);
# `B-V` and `TargetClass` are strongly negatively correlated. `Amag` and `TargetClass` are moderately positively correlated. Other correlations appear fairly weak.
# # Create test train split
X = star_df.drop('TargetClass', axis=1)
y = star_df.TargetClass
# list of predictor names for later labelling
X_names = list(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# Now prepare categorical columns for One-hot Encoding.
# sort categorical columns from numeric
cat = (X_train.dtypes == 'object')
cat_cols = list(cat[cat].index)
# # Start building the pipeline
# +
# create one-hot encoder transformer
cat_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore', sparse='False'))
])
# create preprocessor
preprocessor = ColumnTransformer(
transformers=[
('cat', cat_transformer, cat_cols)
])
# -
# ## First model - Dummy classifier
# define initial dummy model
dummy_model = DummyClassifier(random_state=42)
dummy_pipe = Pipeline(steps=[('preprocessor', preprocessor),
('model', dummy_model)])
# run the dummy pipeline
dummy_pipe.fit(X_train, y_train)
# +
# def model metrics for use in all models tried
# assumes train test split defined with X and y
def model_metrics(input_pipe):
print("Training Accuracy", (input_pipe.score(X_train, y_train)) * 100)
print("Test Accuracy", (input_pipe.score(X_test, y_test)) * 100)
y_pred_test = input_pipe.predict(X_test)
y_pred_train = input_pipe.predict(X_train)
print("MAE train", mean_absolute_error(y_train.astype('int'),
y_pred_train.astype('int')))
print("MAE test", mean_absolute_error(y_test.astype('int'),
y_pred_test.astype('int')))
print("AUC train", roc_auc_score(y_train, y_pred_train))
print("AUC test", roc_auc_score(y_test, y_pred_test))
# -
model_metrics(dummy_pipe)
# As expected, the dummy classifier is no better than 50/50 guess!
# ## Second model - Random Forest
#
# Note - This model is for comparision with XGBoost.
# If I were training a random methodically, could use cross-validation (i.e. `GridSearchCV`) to maximise AUC and tune the hyper-parameters to determine the optimal value from the search grid.
#
# Three parameters are important to tweak to avoid overfitting a random forest:
#
# * `n_estimators` - more trees reduces overfitting
# * `max_depth` - the deeper the tree the more the risk of overfitting
# * `min_samples_leaves` - the smaller the number of training exmples in the leave the greater the risk of overfitting
# +
#use default settings for random forest
rf_model = RandomForestClassifier(random_state=42)
rf_pipe = Pipeline(steps=[('preprocessor', preprocessor),
('model', rf_model)])
# run the rf pipeline
rf_pipe.fit(X_train, y_train)
# -
# get model metrics
model_metrics(rf_pipe)
# Oh wow. This data is modelled extremely well with the random forest model. As I thought initially, this is not a good dataset for XGBoost. But it will give me good practice in setting up the pipelines to run the models.
# ## Third model - XGBoost
# Extreme Gradient Boosting (XGBoost) uses gradient boosting framework.
# Boosting uses weak learners (models), makes conclusions about feature importance and parameters and uses these to build better models by trying to reduce the error of the previous model.
#
# For XGBoost the weak learner is a tree ensemble.
# The model has an inbuilt cross-validation function.
# It can handle missing values.
#
# For this run I will use the scikit-learn API of XGBoost
#
# - note if use xgb version of XGBoost the data needs to be in DMatrices:
# - e.g. :
# - import xgboost as xgb
# - dtrain = xgb.DMatrix(X_train, label=y_train) etc.
# +
#use default settings for XGBoost, set eval_metric
xgb_model = XGBClassifier(use_label_encoder=False, eval_metric = 'mlogloss')
xgb_pipe = Pipeline(steps=[('preprocessor', preprocessor),
('model', xgb_model)])
# run the rf pipeline
xgb_pipe.fit(X_train, y_train)
# -
# get model metrics
model_metrics(xgb_pipe)
# XGBoost straight out of the box is performing reasonably well, but it is not as good as the random forest. Try tuning the hyperparameters to get a better model.
# Hyperparameters we can tune include:
#
# * `n_estimators` - same as the random forest this is the number of models in the ensemble
# * `learning_rate` - shrinks the feature weights on each step to prevent overfitting. Default is 0.1. Lowering this reduces how much the model retrains and can help prevent overfitting, however a lower learning rate also needs more boosting rounds and increases training time
# * `objective` - defines the loss function to be minimised
# * `early_stopping_rounds` - not used in the parameter tuning below. This helps prevent overfitting by assessing the performance of the model at each round and stopping the model if performance hasn't improved by the number of rounds set by this parameter
# To have a look I am going to change some of the hyperparameters discussed above, but a more robust way to tune hyperparameters is with the `cv` function from XGBoost. This lets you run cross-validation on the training dataset and returns a mean MAE score.<br /><br />
# To do this the data would need to be in XGBoost format and we would need to be running the `xgb` version of `xgboost`.
# +
# declare parameters to be tuned
param = {
'objective': 'binary:logistic',
'max_depth': 4,
'alpha': 10,
'learning_rate': 0.15,
'n_estimators': 1000,
'use_label_encoder': False,
'eval_metric': 'mlogloss'
}
# +
#use parameters for XGBoost, set eval_metric
xgb_param_model = XGBClassifier(**param)
xgb_param_pipe = Pipeline(steps=[('preprocessor', preprocessor),
('model', xgb_param_model)])
# run the rf pipeline
xgb_param_pipe.fit(X_train, y_train)
# -
# get model metrics
model_metrics(xgb_param_pipe)
# Adding the parameters has made the model worse! Another sign that the XGBoost model really isn't the model for this data.
# ## ROC comparison of models
#
# To compare the models run on the data so far, I will finish this off with a ROC comparison plot of all models.
# +
plt.figure(0).clf()
# add pipes for ROC
pipes = [
{
'label':'Dummy Classifier',
'pipe': dummy_pipe,
},
{
'label':'Random Forest',
'pipe': rf_pipe,
},
{
'label':'XGBoost',
'pipe': xgb_pipe,
},
{
'label':'XGBoost + parameters',
'pipe': xgb_param_pipe,
}
]
# iterate through pipelist
for p in pipes:
pipe = p['pipe']
y_pred=pipe.predict(X_test)
fpr, tpr, thresholds = roc_curve(y_test, pipe.predict_proba(X_test)[:,1])
auc = roc_auc_score(y_test, y_pred)
plt.plot(fpr, tpr, label='%s ROC (area = %0.2f)' % (p['label'], auc))
plt.plot([0, 1], [0, 1],'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('1-Specificity(False Positive Rate)')
plt.ylabel('Sensitivity(True Positive Rate)')
plt.title('Receiver Operating Characteristic')
plt.legend(loc="lower right")
plt.show()
# -
# # Summary
#
# * Initial run of XGBoost
# * Addition of parameter feature tweak to XGBoost
# * Random forest outperformed XGBoost on this data
# * Highlights the importance of selecting data-appropriate models
# # Future runs
#
# * Include CV (cross-validation) for hyperparameter tuning
# * Remember that if you are running predictions - once you have the best model, run it again on all the data
| xgboost/star_classifier_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Measuring a PSF for Spliner Analysis
#
# In this example we'll measure a microscope PSF for Spliner analysis.
# ### Configuration
#
# To measure the PSF you'll need a movie of sparse fluorescent beads being scanned through the microscope focus. We typically do this with small fluorescent beads on the order of 0.1um in diameter. The beads are fixed to the coverslip using a buffer with 100mM MgCl2. The coverslip is scanned through the focus in 10nm steps using a piezo z scanner. We assume that the drift in XYZ is neglible during the time it takes to take these movies (10s of seconds).
#
# In this example we're just going to simulate this using a theoritical PSF that is an idealized astigmatic PSF.
import os
os.chdir("/home/hbabcock/Data/storm_analysis/jy_testing/")
print(os.getcwd())
# Generate the sample data for this example.
import storm_analysis.jupyter_examples.spliner_measure_psf as spliner_measure_psf
spliner_measure_psf.configure()
# ### Prerequisites
# A text file containing the z-offset of each frame in the movie. This file contains two columns, the first is whether or not the data in this frame should be used (0 = No, 1 = Yes) and the second contains the z offset in microns.
# +
# Load sample z_offsets.txt file. In this example every frame is valid.
import numpy
z_offsets = numpy.loadtxt("z_offsets.txt")
print(z_offsets[:5,:])
# -
# A text file containing the approximate bead locations. One way to create this is by using the `visualizer` tool in the `storm_analysis.visualizer` folder. Another alternative might be using ImageJ. Here we are just going to use generated file.
bead_locs = numpy.loadtxt("bead_locs.txt")
print(bead_locs)
# Check that the specified positions align with the actual bead locations.
#
# spliner_measure.tif is the name of the simulated movie.
#
import storm_analysis.jupyter_examples.overlay_image as overlay_image
overlay_image.overlayImageBeads("spliner_measure.tif", "bead_locs.txt", 40)
# ### PSF Measurement
# Now that we have the 3 files that we need we can measure the PSF. You want to use an AOI that is a bit larger then the size you plan to use when creating the spline file.
#
# Notes:
#
# * The AOI size is the 1/2 size so using 12 pixels means that the final PSF will be 24 x 24 pixels.
# * The pixel size in this simulation was 100nm
# * The simulated movie covered a Z range of +-0.6um, so we're using exactly that. For real data you might want to use a Z range that is slightly smaller than range you scanned.
# * The Z step size is 50nm, which is fine enough for most applications, assuming a high NA objective with a depth of field on the order of 1um.
#
# Also:
#
# If you think that your coverslip might not be completely flat, or that your bead locations are not sufficiently accurate then you can set the `refine` parameter to `True`. With this setting each PSF will be aligned to the average of the remaining PSF using a correlation metric. You may see some warning messages like `Warning: Desired error not necessarily achieved due to precision loss.`, it is usually safe to ignore these.
# +
import storm_analysis.spliner.measure_psf_beads as measure_psf_beads
measure_psf_beads.measurePSFBeads("spliner_measure.tif",
"z_offsets.txt",
"bead_locs.txt",
"spliner_psf.psf",
aoi_size = 12,
pixel_size = 0.1,
refine = True,
z_range = 0.6,
z_step = 0.05)
# -
# ### Visualizing the measured PSF
# Make some images of the measured PSF.
# +
import storm_analysis.jupyter_examples.psf_images as psfImages
psfImages.psfImages("spliner_psf.psf")
# -
# Print some properties of the PSF.
# +
import storm_analysis.spliner.print_psf as printPSF
printPSF.printPSF("spliner_psf.psf")
# -
# Now you can use `storm_analysis.spliner.psf_to_spline` to convert the PSF into a spline that `Spliner` can use for SMLM movie analysis
# + active=""
# Reference printPSF.printPSF() results:
#
# pixel_size 0.1
# psf shape (25, 24, 24)
# type 3D
# version 2.0
# zmax 600.0
# zmin -600.0
# zvals [-600.0, -550.0, -500.0, -450.0, -400.0, -350.0, -300.0, -250.0, -200.0, -150.0, -100.0, -50.0, 0.0, 50.0, 100.0, 150.0, 200.0, 250.0, 300.0, 350.0, 400.0, 450.0, 500.0, 550.0, 600.0]
| jupyter_notebooks/spliner_measure_psf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demo of simple density-based imitation learning baselines
#
# This demo shows how to train a `Pendulum` agent (exciting!) with our simple density-based imitation learning baselines. `DensityTrainer` has a few interesting parameters, but the key ones are:
#
# 1. `density_type`: this governs whether density is measured on $(s,s')$ pairs (`db.STATE_STATE_DENSITY`), $(s,a)$ pairs (`db.STATE_ACTION_DENSITY`), or single states (`db.STATE_DENSITY`).
# 2. `is_stationary`: determines whether a separate density model is used for each time step $t$ (`False`), or the same model is used for transitions at all times (`True`).
# 3. `standardise_inputs`: if `True`, each dimension of the agent state vectors will be normalised to have zero mean and unit variance over the training dataset. This can be useful when not all elements of the demonstration vector are on the same scale, or when some elements have too wide a variation to be captured by the fixed kernel width (1 for Gaussian kernel).
# 4. `kernel`: changes the kernel used for non-parametric density estimation. `gaussian` and `exponential` are the best bets; see the [sklearn docs](https://scikit-learn.org/stable/modules/density.html#kernel-density) for the rest.
# +
# %matplotlib inline
# #%load_ext autoreload
# #%autoreload 2
import pprint
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from imitation import density_baselines as db
from imitation import util
# -
env_name = 'Pendulum-v0'
env = util.make_vec_env(env_name, 8)
rollouts = util.rollout.load_trajectories("../tests/data/expert_models/pendulum_0/rollouts/final.pkl")
imitation_trainer = util.init_rl(env, learning_rate=3e-4, nminibatches=32, noptepochs=10, n_steps=2048)
density_trainer = db.DensityTrainer(env,
rollouts=rollouts,
imitation_trainer=imitation_trainer,
density_type=db.STATE_ACTION_DENSITY,
is_stationary=False,
kernel='gaussian',
kernel_bandwidth=0.2, # found using divination & some palm reading
standardise_inputs=True)
# +
novice_stats = density_trainer.test_policy()
print('Novice stats (true reward function):')
pprint.pprint(novice_stats)
novice_stats_im = density_trainer.test_policy(true_reward=False)
print('Novice stats (imitation reward function):')
pprint.pprint(novice_stats_im)
for i in range(100):
density_trainer.train_policy(100000)
good_stats = density_trainer.test_policy()
print(f'Trained stats (epoch {i}):')
pprint.pprint(good_stats)
novice_stats_im = density_trainer.test_policy(true_reward=False)
print(f'Trained stats (imitation reward function, epoch {i}):')
pprint.pprint(novice_stats_im)
| notebooks/density_baseline_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Each Box is a "Cell" in Jupyter, when you click the Run button above itll go to the next Cell and run it, some of it is simply text, you can click next as you're reading things, feel free to skip by all the code if you don't need to see that, just hit run and itll jump to the next cell, no problems.<span style="color:green"><span style="font-family:Courier; font-size:medium;"> Green Text</span></span> is information<span style="color:blue"><span style="font-family:Courier; font-size:medium;"> Blue Text</span></span> is a Transition of speaking.<br><br>
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# Grabbing RotF Data from this pc, starting with basic imports.
# Using csv to parse the massive text files we make from the rotf discord bot channels.
# </span></span>
#
#
import csv
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# We use this function to read and clean the data we get from the rotf discord channel. Here is an example of what a line looks like from the csv.</span></span><br><br>
# ['deaths', '2019-07-18 21:16:42.498000', '<@&542343490600828930> : Pekson (6/8, 868) has been crushed by Thusala!']<br><br>
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# upon splitting this single line we get a very disgusting split on the main text that we want so we have to clean up each player, bossname, date and base fame. Basefame from the discord channel is your base fame and not your actual death fame, using a bracketed system we've made a crude estimate of each death (Feel tree to tweak these numbers as you see fit.)
# </span></span>
# +
def loot_data():
with open('Loot.txt', 'r') as file:
items = {} #Used to filter and find names for item blacklist
newFile = open('pandas_loot_P4.txt', 'w')
newFile.write('player;item;type;date\n')
blacklist = ['mike', 'miniguy', 'miniguy', 'skilly', 'tuckingfypo', 'blanks', 'lootaveli', 'kyooooh', 'letgay',
'freakbaby', 'byrooni', 'freeprem', 'darazakel', 'swan', 'chao', 'thotiana', 'kouhai', 'sendhelp',
'weebster', 'son', 'calena', 'fleaf', 'loganalt', 'mike', 'skilly', 'lilgary', 'myra', 'kotvkedax',
'zee', 'keep', 'shinx']
item_blacklist = ['pls video ;(((', 'pls video ;(((', 'autism', 'u aint gettin one', 'Depression',
'hah u thought', 'YOINKED UR 10K FAME - ARENA', 'gay', 'Dead Streamer', 'Gay', '0 0 0',
'A can of whoopass made for the hand.', 'yOU SMELL THIS TO0???', 'depression',
'Streetwise dumb bruh, but booksmart.', 'Fioreen said to', 'bruhmoment', 'Lol get rekt mate',
'Canhannon', 'LOL', 'looooooOOOOOOOOOOoooooooL', 'never getting a hc legit btw', '1',
'Hvis pose', 'Hentai', 'Short Dagger', 'breastplate of the big titan',
'Followers for Depression', 'u wish', 'Admin Sword', 'Omni Ring', 'oWocannon', ':)I',
'handcannon', 'LOOOOOOL', 'Omnipotence Ring', 'u like getting loot', 'Public Arena Key',
'64 Stacks of Cobblestone', ':)', 'this guy has 100% death ratio', 'this guy has swagger',
'U GOT IT AGAIN GROSS BOW', 'GROSS BOW!!!', 'Shattered Waraxe', 'Lucky Potio',
'he got hand & feet too', 'head', 'Lol you wish this was an Asura', 'MONEY MOVES!!!',
'Handcannon', "Tiive's Banhammer", "The Banhammer",'no',"Thusula's Slasher"]
loots = csv.reader(file, delimiter=' ', quoting=csv.QUOTE_NONE)
for eachDrop in loots:
if len(eachDrop) > 5:
# Get Date
date = eachDrop[1][1:]
# Get Player name
user = eachDrop[5][1:-1]
user = user.lower()
# Get Drop Type (Pimal/Lege) 200 tokens check
drop_type = eachDrop[9]
# primal=Primal, legendary=Legendary, Necrotic = 200 necrop tokens.
# Get item name
item_name = eachDrop[11:]
final_item = ''
for eachIndex in item_name:
if eachIndex[len(eachIndex)-1] == ']':
final_item += eachIndex
final_item = final_item[1:-1]
break
else:
final_item += eachIndex + ' '
if drop_type == 'primal' and final_item not in item_blacklist:
# Collect item
if final_item not in items and final_item not in item_blacklist:
newFile.write(user + ';' + final_item + ";" + drop_type + ';' + date + '\n')
elif final_item in items and final_item not in item_blacklist:
newFile.write(user + ';' + final_item + ";" + drop_type + ';' + date + '\n')
elif drop_type == 'legendary' and final_item not in item_blacklist:
#Item name collection
if final_item not in items and final_item not in item_blacklist:
newFile.write(user+';'+final_item+";"+drop_type+';'+date+'\n')
elif final_item in items and final_item not in item_blacklist:
newFile.write(user+';'+final_item+";"+drop_type+';'+date+'\n')
# Total lege addition
elif drop_type == 'Necrotic':
# Total Token addition
if drop_type =='Necrotic':
# Extra work to grab token names
token_item = final_item.split()
final_item = ''
for eachIndex in token_item[2:]:
if ']' in eachIndex:
final_item += eachIndex
final_item = final_item[1:-3]
#print(user, final_item, drop_type, date) working 100%
newFile.write(user+';'+final_item+";"+drop_type+';'+date+'\n')
break
else:
final_item += eachIndex + ' '
newFile.close()
return 'Completed, filename = pandas_loot.txt'
def death_data():
with open('Death.txt', 'r') as DeathFile:
blacklist = ['mike', 'miniguy', 'miniguy', 'skilly', 'tuckingfypo', 'blanks', 'lootaveli', 'kyooooh', 'letgay',
'freakbaby', 'byrooni', 'freeprem', 'darazakel', 'swan', 'chao', 'thotiana', 'kouhai', 'sendhelp',
'weebster', 'son', 'calena', 'fleaf', 'loganalt', 'mike', 'skilly', 'lilgary', 'myra', 'kotvkedax',
'zee', 'keep', 'lilpredato','nexuscrier','itscubic','vagrant','']
newFile = open('pandas_death_P4.txt', 'w')
newFile.write('player;basefame;monster;date\n')
deaths = csv.reader(DeathFile, delimiter=' ', quoting=csv.QUOTE_NONE)
for eachDeath in deaths:
if len(eachDeath) > 4 and eachDeath[3] != "'@Deaths:":
date = eachDeath[1][1:]
death = eachDeath[5:]
player = death[0]
player = player.lower()
max_deaths = death[1][1:-1]
#
monster = death[7:]
if monster[0] == 'Epic':
monster = monster[1:]
monster = ' '.join(monster)[:-3]
basefame = int(death[2][:-1])
# Collect Player that died and their fame gained
if True:
if basefame > 0 and basefame <= 450:
newFile.write(player + ';' + str(basefame*1) + ';' + monster + ';' + date + '\n')
elif basefame > 450 and basefame <= 1000:
newFile.write(player + ';' + str(basefame*1.5) + ';' + monster + ';' + date + '\n')
elif basefame > 1000 and basefame <= 3500:
newFile.write(player + ';' + str(basefame*2) + ';' + monster + ';' + date + '\n')
elif basefame > 3500 and basefame <= 5000:
newFile.write(player + ';' + str(basefame*2.5) + ';' + monster + ';' + date + '\n')
elif basefame > 5000 and basefame <= 7500:
newFile.write(player + ';' + str(basefame*3.5) + ';' + monster + ';' + date + '\n')
elif basefame > 7500 and basefame <= 9000:
newFile.write(player + ';' + str(basefame*4) + ';' + monster + ';' + date + '\n')
elif basefame > 9000 and basefame <= 9999:
newFile.write(player + ';' + str(basefame*4.5) + ';' + monster + ';' + date + '\n')
elif basefame >= 10000 and basefame <= 10500:
newFile.write(player + ';' + str(basefame*7) + ';' + monster + ';' + date + '\n')
elif basefame > 10500 and basefame <= 12500:
newFile.write(player + ';' + str(basefame*5.5) + ';' + monster + ';' + date + '\n')
elif basefame > 12500 and basefame <= 15000:
newFile.write(player + ';' + str(basefame*5.75) + ';' + monster + ';' + date + '\n')
elif basefame > 15000 and basefame <= 29999:
newFile.write(player + ';' + str(basefame*6) + ';' + monster + ';' + date + '\n')
elif basefame > 29999 and basefame <= 31000:
newFile.write(player + ';' + str(basefame*6.5) + ';' + monster + ';' + date + '\n')
elif basefame > 31000 and basefame <= 99999:
newFile.write(player + ';' + str(basefame*10) + ';' + monster + ';' + date + '\n')
elif basefame > 99999:
newFile.write(player + ';' + str(basefame*15) + ';' + monster + ';' + date + '\n')
newFile.close()
return 'Completed, filename = pandas_death.txt'
print("Give this part one moment, its cleaning every single death and loot drop from the discord channel.\nYou will see two 'Completed' messages below this and this block is considered finished.")
print(loot_data())
print(death_data())
# -
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# Nice The most confusing part is complete; Now we can actually begin exploring the data we've collected using 2 new libraries. Pandas and Seaborn, we'll use pandas to easily deal with our new text files as data frames and seaborn to better understand our data through visualization(graphs)</span></span>
# Each Import has that as there so that we don't have to write out pandas and seaborn when actively using each library.
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# Awesome, let's get a quick run down of what our data looks like from both the pandas_loot and pandas_death text files using pandas, When using the head() function from pandas it will print the first 5 rows of each dataframe, we can also grab a summary of each Data frame and look to see if anything interesting shows up immediately.</span></span>
# +
# Although we are using txt files we can still read it as a csv if we include a delimiter in our case ;
# Every column of data in each line of our text files was seperated with a ';'
loot_df = pd.read_csv('pandas_loot.txt', sep=';')
death_df = pd.read_csv('pandas_death.txt', sep=';')
print("Checking top 5 rows from Loots")
print(loot_df.head())
print("\n\nChecking Top 5 rows from Deaths")
print(death_df.head())
# -
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">From this we can see our 4 column names for each Dataframe, Loots uses players, item, type and date. Deaths uses player, basefame, monster and date. This data is currently not sorted in any way it simple shows the first 5 lines of each text file the pandas dataframe was created from.</span></span><br><br>
# <center><span style="color:blue"><span style="font-family:Courier; font-size:large;">
# Lets go a bit deeper with pandas function describe()
# </span></span></center>
#
# +
print("Using .describe on Loots DF\n")
print(loot_df.describe())
print("\n\nUsing .describe on Deaths DF\n")
print(death_df.describe())
# -
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# Lets start with Deaths here as its the simpler one to explain (ironic). Here you see it only shows one column basefame, thats because all of the other columns were all string type data pieces, Pandas will always work around one datatype for a dataset unless specified directly, in this example basefame is type float, so pandas prioritizes that and ends up with some massive numbers due to how its grabbing the sum of the basefame column. Quick spoilter there is well over 400 million fame from deaths alone on the server right now just from the data we get from this discord channel.</span></span>
#
# <br>
# <center><span style="color:red"><span style="font-family:Courier; font-size:x-large;">--Count--</span></span></center>
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# Count from each column is expectedly equal, there are 54,232 total drops logged on that bot that are legendary or primal and not absolute trolls by mike, we blacklisted a wonky amount of items and a couple of players that mike tends to troll drop most often to try and deal with those issues.</span></span>
#
# <br>
# <center><span style="color:red"><span style="font-family:Courier; font-size:x-large;">--Unique--</span></span></center>
#
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# Out of the 54,232 drops 10,864 players have dropped them all meaning that in an ideal world each player is dropping a mix of 5 lege/primal/tokens.<br><br>
# There are 154 unique items and 3 unique types Legendary, Primal or the 200 Necropolis Tokens turn in.<br><br>
# There are 159 total unique days of data from the discord channel.<br></span></span>
#
# <br>
# <center><span style="color:red"><span style="font-family:Courier; font-size:x-large;">--Top--</span></span></center>
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# Top is going to seem obvious as well, Pandas doesn't hide behind much confusing and obfuscation with the information it projects. <br><br>
# ehoniii has the most overall entries in the players column meaning he may have the most drops if we include all legendaries, primals and token turn ins. <br><br>
# The most common item is the planewalker, likely due to sprites mega spam for easy money starting off. <br><br>
# The most common type is Legendary, expectedly; primals only dropped from one dungeon until the yeti boy came around. <br><br>
# 6/16/2019 is the day with the most logged drops of them all, this was on a sunday dat 25% boost :^)<br></span></span>
#
# <br>
# <center><span style="color:red"><span style="font-family:Courier; font-size:x-large;">--Frequency--</span></span></center>
# <span style="color:green"><span style="font-family:Courier; font-size:medium;">
# FREQ is short for frequency, this line gives us a bit more information on the "TOP" row<br><br>
# Player "ehoniii" has dropped 291 notable times. <br><br>
# There have been 4724 Planewalker drops. <br> <br>
# Out of the 54,232 logged drops 49,753 of them are Legendary <br> <br>
# On 6/16/2019 802 Notable items dropped. <br><br>
# <br><br>
# Lets look at that last part a bit more, out of 159 total days if 54,232 drops were evenly split you would have Approximately
# <br><br>
# 342(rounded up) drops per unique day This Sunday doubled the average amount and then some. a near 200% Average drop of items.
# It's nice to break the data down like this. it becomes much easier to find interesting anomalies of sorts
# </span></span>
# <br><br>
# <center><span style="color:blue"><span style="font-family:Courier; font-size:large;">I want to look into the Deaths dataframe a bit more as it returned nothing insightful to us.</span></span></center>
#
# +
print("Getting the sum of all deaths logged\n")
print("{:,}".format(round(death_df.basefame.sum())) +'Total fame\n')
print("Getting the top 5 deaths\n")
sorted_basefame = death_df.sort_values('basefame', ascending=False)
print(sorted_basefame.head())
print("\nGetting the top 5 monster killers\n")
monster = death_df.monster.value_counts()
print(monster.head())
# -
# <span style="color:green"><span style="font-family:Courier; font-size:large;">From here we can see that there is approximately 428 million fame has entered the game since the start of this rotf discord bot. The top 5 deaths are ones from the legends leaderboard expectedly. Astonishingly in the top 5 monsters the "Angry Yeti" has surpassed all of the other monsters and he was released much later into the rotf notifier bot's existence! Actual hard content? :O</span></span>
#
# </span></span> <span style="color:blue"><span style="font-family:Courier; font-size:x-large;"> Let's look back at the loots Dataframe a bit more and prioritize the Date column</span></span>
#
print("Getting Dates information from Loots_DF")
dates = loot_df.date.value_counts()
print(dates.head())
# <span style="color:green"><span style="font-family:Courier; font-size:large;">That is very interesting the top 5 total drops days are ALL Sundays. The 25% Loot boost almsot certainly plays a role in that, however it may be more about getting MORE players online to play actively inflating the drops per day as well. I do wonder what happened on 6/16/2019 to make it stand out so much more than the rest. Lets try and take a peak deeper into that day.</span></span>
print("Checking one date")
selected_date = loot_df.loc[loot_df['date'] == '2019-06-16']
print(selected_date.describe())
# <span style="color:green"><span style="font-family:Courier; font-size:large;">Player named "baxter" got 9 drops <br> 78 of the drops this day were Shattered War Axes.
# <br>750 of the 802 drops were legendary <br>802 drops are split between 549 players and 102 different items drop, this number might be slightly slower since we seperated the primal tokens from the primal drops.<br> Lets dig deeper.</span></span>
#
# +
item_drops = selected_date.item.value_counts()
print("Top 5 dropped items")
print(item_drops.head())
print('\n\n')
print("Top 5 players (drop count)")
player_counts = selected_date.player.value_counts()
print(player_counts.head())
# -
# <span style="color:green"><span style="font-family:Courier; font-size:large;">This appears to be just another day; showing the top 10 players and dropped items we see nothing abnormal about this, our item and player blacklists are working as intended. This means that this sunday above all others was just a very lucky sunday or there were many staff events that would inflate the drop counter substantially (This would take some effort on their end).</span></span>
# <br>
# <center><span style="color:blue"><span style="font-family:Courier; font-size:x-large;"> Let's get more Visual</span></span></center>
#
#
#
print("Using the player counts we can get a better understanding on player 'luck'")
player_counts.plot.hist(title='x=Number of drops, y=#Number of players')
# <span style="color:green"><span style="font-family:Courier; font-size:large;">While there is a notion from the playerbase that they are "unlucky" for not getting more drops the reality is that 98% of the playerbase coasts at around 1-3 drops, 1 drop being the bulk of the playerbase. and this is on the LUCKIEST day we have logged so far. You are not unlucky that player is likely EXTREMELY lucky or more realistically they are actively killing valuable monsters much more frequently than you are in a given time span.</span></span> <br><br><br>
#
# <center><span style="color:blue"><span style="font-family:Courier; font-size:x-large;">Let's Look at primal drops</span></span></center>
# +
selected_type = loot_df.loc[loot_df['type'] == 'primal']
selected_type_tokens = loot_df.loc[loot_df['type']=='Necrotic']
combined = pd.merge(selected_type, selected_type_tokens, how='outer')
print(combined.head())
print('\n',combined.describe())
# -
# <span style="color:green"><span style="font-family:Courier; font-size:large;">Showing just primals Cocknibble is insane with 69 primal drops<br> the most common primal the "Orb of Destiny".<br>
# A very interesting thing to noticed here is that the 4,473 total drops are dropped by just 982 players, A massive increase
# to the average notable drop count we saw erlier at just 1 per player, this is where the good players dwell.<br>
# </span></span><br>
# <center><span style="color:blue"><span style="font-family:Courier; font-size:large;">Time for a better visual, lets see if there are larger discrepencies.</span></span></center>
player_counts = combined.player.value_counts()
print(player_counts.head())
player_counts.plot.hist()
# <span style="color:green"><span style="font-family:Courier; font-size:large;">I used the df.head function to show the top 5 players as far as dropped primals goes and then tossed ALL players onto a histogram, the X axist is the total Number of primals dropped by some player. The Y axis is the number of players that have dropped that many since there are so many people here, the scaling looks wonky, Cocknibble is alone up at 70 and you can't even see his bar he's that small of a portion of the playerbase. Playes are not "unlucky" they are simply skewed by seeing some of the more avid and non time-gated players go ham.<br><br></span></span> <center><span style="color:blue"><span style="font-family:Courier; font-size:large;">Let's look at specific items.</span></span></center>
# +
item_counts= combined.item.value_counts()
print("Printing top 5 most common primals")
print(item_counts.head())
print('\n')
print("Printing the rarest primal")
print(item_counts.tail(1))
print('\n')
item_counts.plot.barh(fontsize=7)
# -
# <span style="color:green"><span style="font-family:Courier; font-size:large;">Nothing out of the ordinary here tbh except the Amulet of Ascension, from what I understand only one has dropped an that guy is perm banned. The Annihilation Staff pieces sub labeled "TN" are very uncommon and rare as the staff is allegedlyop, Apex bulwark is actually VERY high up to where you would expectit to be, a large portion of them come from the necropolis tokens and not from the actual boss, its also not able to drop from chests now. All in all the drops are pretty straight forward, many of the items with 'more' drops are from the earlier bosses/chests where there are the most players applying for a chance at the drops. Currently the Amulet of Ascension is the rarest primal drop in the game, unless we include the staff of Annihilation and the banned amulet drop. </span></span><center><span style="color:blue"><span style="font-family:Courier; font-size:large;"> <br>Lets take another peak at the dates and see what we can learn from it.</span></span></center>
#
# +
#Adding the weekday into our data
loot_df['date'] = pd.to_datetime(loot_df['date'])
loot_df['weekday'] = loot_df['date'].dt.day_name()
print(loot_df.head(),'\n')
weekdays = loot_df.weekday.value_counts()
print(weekdays)
weekdays.plot.bar()
# -
# <span style="color:green"><span style="font-family:Courier; font-size:large;">Expectedly Sunday has quite a bit more drops than the rest of the days, a good portion may be due to the 25% boost, but I do believe that with the showing from Saturday here that the increase is likely due to the player count rising on these days, Sunday having that promotion of a loot boost bonus on that day is certainly inviting to all new players though I do feel it's likely "free" advertising where the 25% boost likely doesn't benefit the player as much, but its more likely the player is putting in much more time on that day actively playing and killing monsters. Friday is a bit lower in comparison to the rest of the weekdays, I wonder why that may be, there may be a valid time where Mike is updating the rotf bot on fridays? Some days are much lower in drop count and death count due to this.</span></span> <br><br><center> <span style="color:blue"><span style="font-family:Courier; font-size:large;"> Let's stop looking at loot for now and venture into visualizing Death information</span></span></center>
# +
print(death_df.head())
print('Total death fame, logged',death_df.basefame.sum())
print(death_df.date.value_counts().head())
sns.lineplot(x='date',y='basefame',data=death_df)
# -
# This will take a few lines to explain, ignoring that blob of text above date and just looking at the lines and y axis you'll see the average fame gained for a death on each day. The darket shaded line is the genuine average and then the line above that is an anomally that was noted but seemingly not directly included as the basefame never goes above 17500, its been weighted down to ensure a somewhat clean visual representation of the data. Each of those lighter shaded peaks is someone killing off a THICK death fame character. I am very curious to see what each day summed up looks like as well, it may give us a cleaner representation of the data.
# +
myData = {'date':[],'basefame':[]}
for eachDate in death_df.date.unique():
myDate = death_df[death_df.date == eachDate]
#print(round(myDate.basefame.sum()))
myData['date'].append(eachDate)
myData['basefame'].append(round(myDate.basefame.sum()))
date_death_df = pd.DataFrame.from_dict(data=myData)
print(date_death_df.head(),'\n')
print(date_death_df.sort_values('basefame', ascending=False).head())
sns.lineplot(x='date',y='basefame',data=date_death_df)
# -
# <span style="color:green"><span style="font-family:Courier; font-size:large;">when we create the total Sum of death fame for each day you can more cleanly see the larger death fame ticks when players like Wealthy, MrDuck and Eziiio killed off some very high grossing characters. You can also see one of the flaws with getting data from the discord bot, when it is down for any time you can see how the graph gets affected negatively. Some days the bots off for nearly the entire day could be due to server instability or someone updating/tweaking the bot. One thing we should not is that from left to the right time is progressing, the fame being attained via death is rising exponentially, likely due to players understanding the quest chest "meta" a bit more, many people are being funneled chests to get themselves and their guilds massive fame gains, with this persistant and clear increase to the sum of deathfame each day, it's safe to say that pure fame will become increasingly less valuable at a very spooky rate, I hope they have plans of sinking this fame some how I know quite a few ideas have been tossed around involving that.</span></span><br><br><center><span style="color:blue"><span style="font-family:Courier; font-size:large;">In Conclusion What can we learn from all of this?</span></span></center>
# <span style="color:green"><span style="font-family:Courier; font-size:large;">We can see that drop rates for items can be closely correlated to a players actual time played, and not so much about "luck" the more high profile monsters you kill the more chances at a valuable primal or legendary you have, with the addition of Necropolis Tokens this shines even more, kill more, profit more.<br><br>Sundays Loot Boost certainly does seem to actively increase the amount of primals and legendaries dropped each Sunday, whether its from the actual 25% Boost or if its a placebo of sorts that makes players actively go out and kill monsters much more on those days is not really conclusive. I think if they disabled the 25% boost, but kept the announcement there for a couple weeks we could get a better understanding of its actual use.<br><br>A players fame gain from deaths are becoming increasingly and rapidly higher on average, with the knowledge of chest funneling becoming more commong along with the addition of "epic" monster and chest proccing there appears to be a very linear and set in stone way for players to make fame. What was once spamming epic dungeons is now more along the lines of hopping into a necrop getting chests, maybe primals and tokens and funneling them all onto one character, you can either kill it off at 10k base where the final fame gain diminishing return hits or you can continue on to hit the legends leaderboard.</span></span><br><br>
# <center><span style="color:blue"><span style="font-family:Courier; font-size:large;">Necrop. Quest Chest Meta</span></span></center>
# <span style="color:orange"><span style="font-family:Courier; font-size:medium;">
# <br><br><tab>A personal opinion on this that comes from experience with predicting things in games like these is that this is very bad for the games longevity. Allow me to explain a bit more, The guild I've played with since these boosts became a thing was No, they were nice enough to actually hear me out on death fame gaining potential and become the first players to really push for efficiency and ultimately became first guild with 100% fame boost 100% loot boost and if I did my predictions right you will see 25/25 character slots around August some time.<br><br> The initial strategy involving death fame sorcerers was a lot of fun and gated behind someones ability to avidly and tirelessly grind out the character to die on it, but the player remains actively playing and attentive to the game and its surroundings. The Idea is that you make a sorcerer get it 82/82 put on some very strong wisdom bonus gear with a PD scepter and one shot gods in the godlands with rare obsession on get 10k base fame grab the non tedious fame bonuses and kill it, rinse repeat. The gods give solid EXP/Fame, with the Release of Necropolis and the Epic Update Death Fame Sorcerers did see a resurgance the buff was amazing, Elithor's soul bottle and the primal scepter reduced the time it took to hit 10k basefame by about 38% from my math done ages ago. You had to actually go back out and run around for a while to hit the 4million tiles for that fame bonus, before it was usuall 200-300k extra tiles and then you're good to suicide the character. <br><br> When Necropolis first hit of course our entire guild lived in there, closing realms as fast as possible to see this great dungeon once again! (The dungeon is a great addition difficulty and second secret memes aside) It did not take long for us to realize it's potential with how frequent quest chests dropped in there, Thus the initial Eziiio funnel began, this was before the epic update but he still rocketed up to 1million leaderboard fame in a couple weeks, this translates to 2m guild fame at 100% fame boost. This became the staple to fame farming, glands farming does not compete with the Quest chest strategy, which is ok. Issues arise when the epic update hit.<br><br> The "Epic" update made it so that monsters that spawn have a chance to become epic increasing their stats (hp, size) and their Experience gain, the latter is extremely powerful for anyone talking fame gains. Quest chests are considered monsters and an entity that spawns, they can become epic,What was a solidified +16base fame(normal chest) and +48 base fame(epic chest) was now jumping all over, epic procs on chests past 10k bf can benefit you 300+ Base Fame the rate at which they came into the game did not get touched and the players understanding of this mechanic is at full power in the games current meta.<br><br> This is ok if a fame sink comes into play, the real issue isn't the fame gained necesarily, its the fame being stagnant(not being deleted) and the ways in which players get the fame. Godlands farming is obsolete, quest chests can take that torch it's totally fine, but in the current state of the game you will see people log in for a Necropolis collect the chests and hop off either using them for themselves or funneling them to another player. <br><br>The Quest Chest drop rate for all other aspects of the game is abysmal in comparison. If keeping chest this way is a sticking plan, perhaps consider alternative farming methods for the chests; For example Move the higher drop rates to the epic dungeon family (UDL,SPRITE,SNAKE PIT, Just about any dungeon that isn't locked to 1 per realm). Make players go out into the game for longer periods of time, as of right now you can safely be optimal logging in checking for new realms hopping into a necrop grabbing chests/primals checking if a realm is worth closing for a necrop(no one does this anymore, its just what we did when the dungeon came out) and logging out, letting the rotf bot ping you if something worth mentioning is there. Redisbursement of the quest chest drop rate tiering would be a great way to relatively stress free resolve some player retention, there are great updates all the time, the "Epic" update has been added onto successfully on many different iterations we can get a full all epic dungeon now, thats amazing, but it's time to entice players to touch the rest of your content, before they write the game off as a borderline deca event 2.0 scene.</span></span>
| pandasRotf/.ipynb_checkpoints/RotF-Pandas-Jupyter-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import bertopic
#Define your data path
txtfolder = '/home/ngr/gdrive/simpletopics/data/processed/'
# The files have some utf-8 encoding issue so just resaving them
with open(os.path.join(txtfolder, 'Krammer20.txt'), encoding="latin-1") as fin:
with open(os.path.join(txtfolder, 'Krammer20_proc.txt'),'w',encoding='utf8') as fout:
fout.write(fin.read())
with open(os.path.join(txtfolder, 'Krammer20_proc.txt'), encoding="utf8") as f:
lines = [(line.rstrip()).split() for line in f]
#Bertopic does not like if there are non strings. Somehow lines seem to have some bad encodings
cleaned = []
for a in lines:
for b in a:
cleaned.extend([b for b in a if isinstance(b, str)])
topic_model = bertopic.BERTopic(verbose=True)
topics, _ = topic_model.fit_transform(cleaned)
freq = topic_model.get_topic_info(); freq.head(5)
# For more usage on using Bertopic for visualizations and get the topic probabilities check out https://github.com/MaartenGr/BERTopic and the colab notebooks in the github repo.
| notebooks/topic_model_BERT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="pOrSBEmKBmxF"
# 
#
#
#
# # SIT742: Modern Data Science
# **(Module: Python Programming)**
#
# ---
# - Materials in this module include resources collected from various open-source online repositories.
# - You are free to use, change and distribute this package.
# - If you found any issue/bug for this document, please submit an issue at [tulip-lab/sit742](https://github.com/tulip-lab/sit742/issues)
#
#
# Prepared by **SIT742 Teaching Team**
#
# ---
#
#
# ## Session 2G: Advanced Data Types
#
# we will learn about advanced data types in addition to the strings and number learnt before. We will examine lists, tuples and dictionaries used for a collection of related data.
# + [markdown] colab_type="text" id="8rxG_XNwBmxI"
# ## Table of Content
#
#
# 1 [List](#cell_list)
#
# 2 [Tuple](#cell_tuple)
#
# 3 [Dictionary](#cell_dict)
#
#
# + [markdown] colab_type="text" id="4Xu7rvCrBmyp"
# <a id = "cell_list"></a>
# + [markdown] colab_type="text" id="ZXxkKq8eBmyq"
#
# ## 1 List
#
# **List is a sequence**
#
# Like a string, a *list** is a sequence of values. The values in a list can be any type. We call the values *items** of the list. To create a list, enclose the items in square brackets.
#
# For example,
#
# + colab={} colab_type="code" id="E7e8UuwGBmyq"
shoplist = ['apple', 'mango', 'carrot', 'banana']
l = [10, 20, 30, 40]
empty = [ ] # Initialize an empty list
# + colab={} colab_type="code" id="aY2iD5kNBmys"
shoplist
# + colab={} colab_type="code" id="KE779JH2Bmyu"
l
# + colab={} colab_type="code" id="_IpjFmd4Bmyx"
empty
# + colab={} colab_type="code" id="XBrjhj1iBmyz"
shoplist
# + [markdown] colab_type="text" id="j7ruHjt3Bmy0"
# The elements of a list do not have to be the same type. An another list can also be nested inside a list.
#
# To access the element of a list, use the bracket operator to obtain the value available at the index. Note that the indices of list start at $0$. You can also use negative value as index, if you counts from right. For example, the negative index of last item is $-1$. Try the following examples:
# + colab={} colab_type="code" id="W9akILf7Bmy1"
l = [10, 20, 30, 40]
# + colab={} colab_type="code" id="Z5zPJWxpBmy3"
l[2]
# + colab={} colab_type="code" id="bNH-O27pBmy4"
l[-1]
# + colab={} colab_type="code" id="NtKexic5Bmy7"
l[-3]
# + [markdown] colab_type="text" id="Wpr1v6zrBmy9"
# Here is an example of nested list:
# + colab={} colab_type="code" id="r0xGwqeJBmy-"
l = ['apple', 2.0, 5, [10, 20]]
# + colab={} colab_type="code" id="SP-ntHwtBmy_"
l[1]
# + colab={} colab_type="code" id="WQsbJkhABmzC"
l[3]
# + colab={} colab_type="code" id="qDryScoCBmzD"
l[3][1]
# + [markdown] colab_type="text" id="z-zzr7ICBmzF"
# Unlike strings, lists are mutable, which means they can be altered. We use bracket on the left side of an assignment to assign a value to a list item.
# + colab={} colab_type="code" id="rX_ghZH_BmzF"
l = [10, 20, 30, 40]
l[1] = 200
# + colab={} colab_type="code" id="WaAO_Ir2BmzH"
l
# + [markdown] colab_type="text" id="bu8DphbXBmzI"
# **List operation**
#
# **In** is used to perform membership operation.
# The result of expression equals to **True** if a value exists in the list,
# and equals to **False** otherwise.
#
# + colab={} colab_type="code" id="XbCUkFPFBmzJ"
shoplist = ['apple', 'mango', 'carrot', 'banana']
# + colab={} colab_type="code" id="FLXYQQo9BmzK"
'apple' in shoplist
# + colab={} colab_type="code" id="IjdYHK1bBmzN"
'rice' in shoplist
# + [markdown] colab_type="text" id="UNLrFdonBmzO"
# Similarly, **In** operator also applies to **string** type. Here are some examples:
#
# + colab={} colab_type="code" id="GsfGjlPyBmzP"
'a' in 'banana'
# + colab={} colab_type="code" id="GBe4RSgOBmzQ"
'seed' in 'banana' # Test if 'seed' is a substring of 'banana'
# + [markdown] colab_type="text" id="HhQjXO2lBmzR"
# '**+**' is used for concatenation operation, which repeats a list a given number of times.
# + colab={} colab_type="code" id="X6RqwGxQBmzT"
[10, 20, 30, 40 ] + [50, 60]
# + [markdown] colab_type="text" id="F6tP9GVhBmzU"
# [50, 60]*3
#
#
# + [markdown] colab_type="text" id="Nw0vR9K_BmzV"
# ** List slices **
# Slicing operation allows to retrieve a slice of of the list. i.e. a part of the sequence. The sliding operation uses square brackets to enclose an optional pair of numbers separated by a colon. Again, to count the position of items from left(first item), start from $0$. If you count the position from right(last item), start from $-1$.
# + colab={} colab_type="code" id="GqB8ElRVBmzV"
l = [1, 2, 3, 4]
# + colab={} colab_type="code" id="qnnI1lXSBmzW"
l[1:3] # From position 1 to position 3 (excluded)
# + colab={} colab_type="code" id="T1T4ljvqBmzY"
l[:2] # From the beginning to position 2 (excluded)
# + colab={} colab_type="code" id="039r1EDMBmza"
l[-2:] # From the second right to the beginning
# + [markdown] colab_type="text" id="x9vjy8R6Bmzb"
# If you omit both the first and the second indices, the slice is a copy of the whole list.
# + colab={} colab_type="code" id="e6U4CrvhBmzc"
l[:]
# + [markdown] colab_type="text" id="0wMQJlYIBmzf"
# Since lists are mutable, above expression is often useful to make a copy before modifying original list.
# + colab={} colab_type="code" id="BA5iYESzBmzh"
l = [1, 2, 3, 4]
l_org = l[:]
l[0] = 8
# + colab={} colab_type="code" id="qTMISAbJBmzn"
l
# + colab={} colab_type="code" id="Smf9tzSSBmzp"
l_org # the original list is unchanged
# + [markdown] colab_type="text" id="2h-9tDg3Bmzq"
# **List methods**
#
# The methods most often applied to a list include:
# - append()
# - len()
# - sort()
# - split()
# - join()
# + [markdown] colab_type="text" id="qxDlX5huBmzr"
# **append()** method adds a new element to the end of a list.
# + colab={} colab_type="code" id="UUs8PmOiBmzs"
l= [1, 2, 3, 4]
# + colab={} colab_type="code" id="sr7rr13xBmzt"
l
# + colab={} colab_type="code" id="3BhyF427Bmzy"
l.append(5)
l.append([6, 7]) #list [6, 7] is nested in list l
# + colab={} colab_type="code" id="kHh85_Y6Bmzz"
l
# + [markdown] colab_type="text" id="XiLgDfWwBmz1"
# **len()** method returns the number of items of a list.
# + colab={} colab_type="code" id="5wRXT4WcBmz1"
l = [1, 2, 3, 4, 5]
len(l)
# + colab={} colab_type="code" id="LhWFPL_zBmz2"
# A list nested in another list is counted as a single item
l = [1, 2, 3, 4, 5, [6, 7]]
len(l)
# + [markdown] colab_type="text" id="xvuQnOPRBmz3"
# **sort()** arranges the elements of the list from low to high.
# + colab={} colab_type="code" id="v2Me8qqfBmz4"
shoplist = ['apple', 'mango', 'carrot', 'banana']
shoplist.sort()
# + colab={} colab_type="code" id="wNW-D9ifBmz6"
shoplist
# + [markdown] colab_type="text" id="YIPqgrbDBmz8"
#
# It is worth noted that **sort()** method modifies the list in place, and does not return any value. Please try the following:
# + colab={} colab_type="code" id="JauCHCv_Bmz8"
shoplist = ['apple', 'mango', 'carrot', 'banana']
shoplist_sorted = shoplist.sort()
# + colab={} colab_type="code" id="k1eeAAaxBmz_"
shoplist_sorted # No value is returned
# + [markdown] colab_type="text" id="IxkXo9AZBm0A"
#
# There is an alternative way of sorting a list. The build-in function **sorted()** returns a sorted list, and keeps the original one unchanged.
# + colab={} colab_type="code" id="SiS31NXcBm0A"
shoplist = ['apple', 'mango', 'carrot', 'banana']
shoplist_sorted = sorted(shoplist) #sorted() function return a new list
# + colab={} colab_type="code" id="K_eMR23aBm0F"
shoplist_sorted
# + colab={} colab_type="code" id="5c1BtFjFBm0G"
shoplist
# + [markdown] colab_type="text" id="gmabEBPqBm0H"
# There are two frequently-used string methods that convert between lists and strings:
#
# First, **split()** methods is used to break a string into words:
# + colab={} colab_type="code" id="NToLD-lOBm0I"
s = 'I love apples'
s.split(' ')
# + colab={} colab_type="code" id="1uH4tlgdBm0J"
s = 'spam-spam-spam'
# A delimiter '-' is specified here. It is used as word boundary
s.split('-')
# + [markdown] colab_type="text" id="XUuTMK7tBm0K"
# Second, **joint()** is the inverse of **split**. It takes a list of strings and concatenates the elements.
# + colab={} colab_type="code" id="HKZrUhUzBm0K"
l = ['I', 'love', 'apples']
s = ' '.join(l)
s
# + [markdown] colab_type="text" id="qFbGRFyoBm0M"
# How it works:
#
# Since **join** is a string method, you have to invoke it on the *delimiter*. In this case, the delimiter is a space character. So **' '.join()** puts a space between words. The list **l** is passed to **join()** as parameter.
#
# For more information on list methods, type "help(list)" in your notebook.
# + [markdown] colab_type="text" id="dOaAysQyBm0M"
# **Traverse a list**
#
# The most common way to traverse the items of a list is with a **for** loop. Try the following code:
# + colab={} colab_type="code" id="2pUz_BuABm0M"
shoplist = ['apple', 'mango', 'carrot', 'banana']
for item in shoplist:
print(item)
# + [markdown] colab_type="text" id="BdoviXGwBm0N"
# This works well if you only need to read the items of the list. However, you will need to use indices if you want to update the elements. In this case, you need to combine the function **range()** and **len()**.
# + colab={} colab_type="code" id="Hw-LhzXhBm0P"
l = [2, 3, 5, 7]
for i in range(len(l)):
l[i] = l[i] * 2
print(l)
# + [markdown] colab_type="text" id="6i0u44sWBm0Q"
# How it works:
#
# **len()** returns the number of items in the list, while **range(n)** returns a list from 0 to n - 1. By combining function **len()** and **range()**, **i** gets the index of the next element in each pass through the loop. The assignment statement then uses **i** to perform the operation.
# + [markdown] colab_type="text" id="PQIrW0NjBm0Q"
# <a id = "cell_tuple"></a>
# + [markdown] colab_type="text" id="kBsd3dhuBm0Q"
# ### 3.2 Tuple
#
# **Tuple are immutable **
#
# A **tuple** is also a sequence of values, and can be any type. Tuples and lists are very similar. The important difference is that tuples are immutable, which means they can not be changed.
#
# Tuples is typically used to group and organizing data into a single compound value. For example,
# + colab={} colab_type="code" id="wyr771UWBm0R"
year_born = ('<NAME>', 1995)
year_born
# + [markdown] colab_type="text" id="jS2uoyGvBm0S"
# To define a tuple, we use a list of values separated by comma.
# Although it is not necessary, it is common to enclose tuples in parentheses.
#
# Most list operators also work on tuples.
# The bracket operator indexes an item of tuples, and the slice operator works in similar way.
#
# Here is how to define a tuple:
# + colab={} colab_type="code" id="sSoZ9rIGBm0S"
t = ( ) # Empty tuple
t
# + colab={} colab_type="code" id="YFukci9YBm0T"
t = (1)
type(t) # Its type is int, since no comma is following
# + colab={} colab_type="code" id="DCYLnFmxBm0U"
t = (1,) # One item tuple; the item needs to be followed by a comma
type(t)
# + [markdown] colab_type="text" id="JL76QtuYBm0V"
# Here is how to access elements of a tuple:
# + colab={} colab_type="code" id="Tz1NCxjyBm0W"
t = ('a', 'b', 'c', 'd')
# + colab={} colab_type="code" id="2iTzh6wwBm0Y"
t[0]
# + colab={} colab_type="code" id="PlIiJg52Bm0Z"
t[1:3]
# + [markdown] colab_type="text" id="35hOHu21Bm0a"
# But if you try to modify the elements of the tuple, you get an error.
# + colab={} colab_type="code" id="_r_paxZnBm0b"
t = ('a', 'b', 'c', 'd')
t[1] = 'B'
# + [markdown] colab_type="text" id="Posmxo-aBm0c"
# ### Tuple assignment
#
# Tuple assignment allows a tuple of variables on the left of an assignment to be assigned values from a tuple on the right of the assignment.
# (We already saw this type of statements in the previous prac)
#
# For example,
#
# + colab={} colab_type="code" id="ChvTjx6qBm0d"
t = ('David', '0233', 78)
(name, id, score) = t
# + colab={} colab_type="code" id="-WDeVp8MBm0e"
name
# + colab={} colab_type="code" id="L4Ey2gDYBm0g"
id
# + colab={} colab_type="code" id="v6i5qFp7Bm0h"
score
# + [markdown] colab_type="text" id="aYAZYDZ3Bm0i"
# Naturally, the number of variables on the left and the number of values on the right have to be the same.Otherwise, you will have a system error.
# + colab={} colab_type="code" id="CZWKRkeZBm0j"
(a, b, c, d) = (1, 2, 3)
# + [markdown] colab_type="text" id="svAgKRIMBm0l"
# ### Lists and tuples
#
# It is common to have a list of tuples. For loop can be used to traverse the data. For example,
# + colab={} colab_type="code" id="erQ7rITYBm0l"
t = [('David', 90), ('John', 88), ('James', 70)]
for (name, score) in t:
print(name, score)
# + [markdown] colab_type="text" id="0N9Ozh__Bm0m"
# <a id = "cell_dict"></a>
# + [markdown] colab_type="text" id="P7Te1xa1Bm0o"
# ### 3 Dictionary
#
# A **dictionary** is like an address-book where you can find the address or contact details of a person by knowing only his/her name. The way of achieving this is to associate **keys**(names) with **values**(details). Note that the key in a dictionary must be unique. Otherwise we are not able to locate correct information through the key.
#
# Also worth noted is that we can only use immutable objects(strings, tuples) for the keys, but we can use either immutable or mutable objects for the values of the dictionary. This means we can use either a string, a tuple or a list for dictionary values.
#
# The following example defines a dictionary:
# + colab={} colab_type="code" id="uclLe92SBm0o"
dict = {'David': 70, 'John': 60, 'Mike': 85}
dict['David']
dict['Anne'] = 92 # add an new item in the dictionary
dict
# + [markdown] colab_type="text" id="oLay_eiFBm0q"
# **Traverse a dictionary**
#
# The key-value pairs in a dictionary are **not** ordered in any manner. The following example uses **for** loop to traversal a dictionary. Notice that the keys are in no particular order.
# + colab={} colab_type="code" id="QKfMs36lBm0t"
dict = {'David': 70, 'John': 60, 'Amy': 85}
for key in dict:
print(key, dict[key])
# + [markdown] colab_type="text" id="vLKSyg5OBm0u"
# However, we can sort the keys of dictionary before using it if necessary. The following example sorts the keys and stored the result in a list **sortedKey**. The **for** loop then iterates through list **sortedKey**. The items in the dictionary can then be accessed via the names in alphabetical order. Note that dictionary's **keys()** method is used to return a list of all the available keys in the dictionary.
# + colab={} colab_type="code" id="hLZi1CgdBm0u"
dict = {'David': 70, 'John': 60, 'Amy': 85}
sortedKeys = sorted(dict.keys())
for key in sortedKeys:
print(key, dict[key])
| Jupyter/M02-Python/M02G-AdvDataTypes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''dt-lab-01-JzneXQ0N'': pipenv)'
# name: python3
# ---
# # Лабораторные работы №1/2
#
# ## Изучение симплекс-метода решения прямой и двойственной задачи ЛП
# ## Условие задачи
#
# Требуется найти решение следующей задачи ЛП:
#
# $$F=\textbf{cx}\rightarrow max,$$
# $$\textbf{Ax}\leq\textbf{b},$$
# $$\textbf{x}\geq 0.$$
# +
import numpy as np
from IPython.display import display_markdown
from src.latex import array_to_latex, matrix_to_latex, equation_system, equation_body, check, Brackets
# Source data
c = np.array([3, 1, 4], dtype=np.float64)
A = np.array([2, 1, 1, 1, 4, 0, 0, 0.5, 1], dtype=np.float64).reshape(3, 3)
b = np.array([6, 4, 1], dtype=np.float64)
# c = np.array([7, 5, 3], dtype=np.float32)
# A = np.array([1, 2, 0, 4, 1, 1, 0, 0.5, 1], dtype=np.float32).reshape(3, 3)
# b = np.array([30, 40, 50], dtype=np.float32)
display_markdown(
"<center> <h2> Исходные данные согласно Варианту 2 </h2> </center>",
array_to_latex(c, 'c', 2, Brackets.square),
matrix_to_latex(A, 'A', 2, Brackets.square),
array_to_latex(b, 'b', 2, Brackets.square),
raw=True
)
# +
system = np.hstack([A, np.eye(A.shape[0])])
labels = [f"x_{idx + 1}" for idx in range(system.shape[1])]
eq = equation_body(c, labels[:A.shape[1]])
eq = f"$$F=-({eq}) \\rightarrow min$$"
cond = f"$${', '.join(labels)} \geq 0$$"
display_markdown(
"<center> <h2> Задача ЛП в канонической форме </h2> </center>",
eq, equation_system(system, b, labels), cond,
raw=True
)
# +
# Source data
c = np.array([3, 1, 4], dtype=np.float64).T
A = np.array([2, 1, 1, 1, 4, 0, 0, 0.5, 1], dtype=np.float64).reshape(3, 3).T
b = np.array([6, 4, 1], dtype=np.float64).T
# c = np.array([7, 5, 3], dtype=np.float32)
# A = np.array([1, 2, 0, 4, 1, 1, 0, 0.5, 1], dtype=np.float32).reshape(3, 3)
# b = np.array([30, 40, 50], dtype=np.float32)
display_markdown(
"<center> <h2> Исходные данные согласно Варианту 2 </h2> </center>",
array_to_latex(c, 'c^T', 2, Brackets.square),
matrix_to_latex(A, 'A^T', 2, Brackets.square),
array_to_latex(b, 'b^T', 2, Brackets.square),
raw=True
)
# +
system = np.hstack([A, np.eye(A.shape[0]) * (-1)])
labels = [f"y_{idx + 1}" for idx in range(system.shape[1])]
eq = equation_body(b, labels[:A.shape[1]])
eq = f"$$Ф={eq} \\rightarrow min$$"
cond = f"$${', '.join(labels)} \geq 0$$"
display_markdown(
"<center> <h2> Двойственная задача ЛП в канонической форме </h2> </center>",
eq, equation_system(system, c, labels), cond,
raw=True
)
# -
# ## Выполнение работы
# +
from IPython.display import display_markdown
from src.simplex import Simplex, Table
table = Table.create(b * - 1, A * - 1, c * - 1, sym='y', fsym='Ф')
display_markdown(
"## Изначальная таблица",
table.table_to_md(),
raw=True
)
solvable = Simplex.solve(table)
if solvable:
tgt, base = table.function_to_md()
display_markdown(
"## Результаты работы simplex-алгоритма",
table.table_to_md(),
"### Целевая функция (для исходной задачи)",
tgt,
raw=True
)
else:
display_markdown(
"## Результаты работы simplex-алгоритма",
table.table_to_md(),
"### Задача не может быть разрешена",
raw=True
)
# +
from src.simplex import Simplex, Table
def resolve(c, A, b, inverse: bool = False):
table = Table.create(b * -1, A * -1, c * -1, sym='y', fsym='Ф') if inverse else Table.create(c, A, b)
tgt, base = table.function_to_md()
display_markdown(
"### Изначальная симплекс-таблица",
table.table_to_md(),
raw=True
)
for idx, (table, pos, flag) in enumerate(Simplex.fix_gen(table)):
if pos and flag:
display_markdown(
"### Недопустимое решение",
base,
f"### Разрешающий элемент $x_{{{pos[0], pos[1]}}}={table.table[pos]}$",
"### Исправленная симплекс-таблица",
table.table_to_md(),
raw=True
)
else:
display_markdown(
"### Опорное решение",
base,
"### Целевая функция",
tgt,
raw=True
)
prev = table
for idx, (table, pos, flag) in enumerate(Simplex.solve_gen(table)):
tgt, base = table.function_to_md()
display_markdown(
f"### {idx + 1} шаг. Разрешающий элемент $x_{{{pos[0]},{pos[1]}}}={prev.table[pos]}$",
table.table_to_md(),
"### Опорное решение",
base,
"### Целевая функция",
tgt,
raw=True
)
prev = table
tgt, base = table.function_to_md(not inverse)
display_markdown(
f"### Результирующая симплекс-таблица",
table.table_to_md(),
"### Целевая функция для исходной задачи",
tgt,
raw=True,
)
return table
result = resolve(c, A, b, inverse=True)
# +
labels = {
f'y_{i + 1}' for i, _ in enumerate(result.h_labels[1:])
}
values = dict()
for value, label in zip(result.table[:-1, 0].flatten(), result.v_labels):
if label not in labels:
continue
values[label] = value
for label in labels.difference(values.keys()):
values[label] = 0
labels, values = map(
list, zip(
*sorted(values.items(), key=lambda x: x[0])
)
)
display_markdown(
"### Проверка решения",
f"$Ф={equation_body(values, labels)}={check(values, b)}$",
raw=True
)
| notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Assignment
#
# - Use the same data you chose for the previous lesson, or get new data.
# - Do train/test split. Report your mean absolute error on the train and test set, for these forecasts:
# - Mean Baseline
# - Naive Baseline (Last Observation)
# - Prophet forecast (you can choose the parameters or use the defaults)
# - Do time series cross-validation, using these Prophet functions:
# - cross_validation
# - performance_metrics
# - plot_cross_validation_metric
# - Commit your notebook to your fork of the GitHub repo.
#
# ### Stretch Challenges
# - **Share your visualizations on Slack!**
# - Use the Wikimedia Pageviews API to get data.
# - [Get daily weather station data](https://www.ncdc.noaa.gov/cdo-web/search) from the NOAA (National Oceanic and Atmospheric Administration). User Prophet to forecast the weather for your local area.
# - Adjust your forecasts with Prophet's [changepoints](https://facebook.github.io/prophet/docs/trend_changepoints.html) and [holidays](https://facebook.github.io/prophet/docs/seasonality,_holiday_effects,_and_regressors.html) options. In addition to Prophet's documentation, read <NAME>ehrsen's blog post, [Time Series Analysis in Python](https://towardsdatascience.com/time-series-analysis-in-python-an-introduction-70d5a5b1d52a). **How do these parameters affect your error metrics?**
# - Learn more about how Prophet works. Read the [tweestorm with animated GIFs](https://twitter.com/seanjtaylor/status/1123278380369973248) by Prophet developer <NAME>, or his [research paper](https://peerj.com/preprints/3190/).
# +
# %matplotlib inline
#import libs
import pandas as pd
import requests
import matplotlib.pyplot as plt
from sklearn.metrics import mean_absolute_error
import matplotlib
from fbprophet import Prophet
from fbprophet.plot import add_changepoints_to_plot
from fbprophet.diagnostics import cross_validation
from fbprophet.diagnostics import performance_metrics
from fbprophet.plot import plot_cross_validation_metric
# -
pd.__version__
matplotlib.__version__
pd.plotting.register_matplotlib_converters()
# ## Get Data
# +
#structure wikimedia pageview api request
project = 'en.wikipedia'
access = 'all-access'
agent = 'user'
article = 'Barkley_Marathons'
granularity = 'daily'
start = '2015070100'
end = '2019052700'
#make wikimedia api GET request
endpoint = f"/metrics/pageviews/per-article/{project}/{access}/{agent}/{article}/{granularity}/{start}/{end}"
url = 'https://wikimedia.org/api/rest_v1' + endpoint
resp = requests.get(url)
#ensure the request is successful
assert resp.status_code == 200
# +
#create dataframe from the wikimedia GET request's response body
bark_mara = pd.DataFrame(resp.json()['items'])
#rename timestamp and views columns for use with Prophet
bark_mara = bark_mara.rename(columns={'timestamp':'ds', 'views':'y'})
#cast ds column to datetime data type with formating
bark_mara['ds'] = pd.to_datetime(bark_mara['ds'], format='%Y%m%d%H')
# -
# ## Train / Test Split
#Set index to ds col
bark_mara = bark_mara.set_index('ds')
#Split train/test datasets by time window
train = bark_mara['2015':'2017']
test = bark_mara['2018':'2019']
#Reset indexes for both dataframes
train = train.reset_index()
test = test.reset_index()
#Plot train and test datasets
ax = train.plot(x='ds',y='y',color='blue',figsize=(16,4))
test.plot(x='ds',y='y',color='red',ax=ax)
plt.title('Daily Pageviews for The Barkley Marathons');
# ## Mean Baseline
# +
#create two new columns for the mean baseline of the train dataset
train['mean_baseline'] = train['y'].mean()
test['mean_baseline'] = train['y'].mean()
#print means
train['y'].mean(), test['y'].mean()
# -
#Plot train and test datasets with the mean baseline
ax = train.plot(x='ds',y='y',color='blue',figsize=(16,4))
test.plot(x='ds',y='y',color='red',ax=ax)
train.plot(x='ds',y='mean_baseline',color='black',ax=ax)
test.plot(x='ds',y='mean_baseline',color='black',ax=ax)
plt.title('Daily Pageviews for The Barkley Marathons');
# ## Naive Baseline (Last Observation)
train['naive_baseline'] = train['y'].shift(1).bfill()
train_last_observation = train['naive_baseline'].iloc[-1]
test['naive_baseline'] = train_last_observation
ax = train.plot(x='ds', y='y', color='blue', figsize=(16,4))
test.plot(x='ds', y='y', color='red', ax=ax)
train.plot(x='ds', y='naive_baseline', color='black', ax=ax)
test.plot(x='ds', y='naive_baseline', color='black', ax=ax);
# ## Prophet Forecast
bark_mara_model = Prophet(daily_seasonality=False)
bark_mara_model.fit(train)
train_forecast = bark_mara_model.predict(train)
test_forecast = bark_mara_model.predict(test)
train_forecast.head()
ax = train.plot(x='ds', y='y', color='blue', label='Train', figsize=(16,4))
test.plot(x='ds',y='y',color='red',label='Test',ax=ax)
train_forecast.plot(x='ds', y='yhat', color='black', label='Prophet forecast, train', ax=ax)
plt.title('Daily Pageviews for The Barkley Marathons');
# ## MAE for Each Forecast
# ### Mean Baseline
train_mae = mean_absolute_error(train['y'], train['mean_baseline'])
test_mae = mean_absolute_error(test['y'], test['mean_baseline'])
train_mae, test_mae
# ### Naive Baseline
train_mae = mean_absolute_error(train['y'], train['naive_baseline'])
test_mae = mean_absolute_error(test['y'], test['naive_baseline'])
train_mae, test_mae
# ### Prophet
train_mae = mean_absolute_error(train['y'], train_forecast['yhat'])
test_mae = mean_absolute_error(test['y'], test_forecast['yhat'])
train_mae, test_mae
# ## Cross Validation
#Cross Validation
df_cv = cross_validation(bark_mara_model, horizon='180 days')
#performance_metrics
performance_metrics(df_cv).head()
#plot_cross_validation_metric
fig = plot_cross_validation_metric(df_cv, metric='mae')
plt.title('Cross Validation MAE for The Barkley Marathons Pageviews');
fig = plot_cross_validation_metric(df_cv, metric='mape')
plt.title('Cross Validation MAPE for The Barkley Marathons Pageviews');
| module2-evaluating-forecasts/evaluating_forecasts_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Remember lists? They could contain anything, even other lists. Well, for dictionaries the same holds. Dictionaries can contain key:value pairs where the values are again dictionaries.
#
# As an example, have a look at the script where another version of europe - the dictionary you've been working with all along - is coded. The keys are still the country names, but the values are dictionaries that contain more information than just the capital.
#
# It's perfectly possible to chain square brackets to select elements. To fetch the population for Spain from europe, for example, you need:
#
# >europe['spain']['population']
# Dictionary of dictionaries
europe = { 'spain': { 'capital':'madrid', 'population':46.77 },
'france': { 'capital':'paris', 'population':66.03 },
'germany': { 'capital':'berlin', 'population':80.62 },
'norway': { 'capital':'oslo', 'population':5.084 } }
# ### Use chained square brackets to select and print out the capital of France.
# Print out the capital of France
print(europe['france']['capital'])
# ### Create a dictionary, named data, with the keys 'capital' and 'population'. Set them to 'rome' and 59.83, respectively.
# Create sub-dictionary data
data={'capital':'rome','population':59.83}
# ### Add a new key-value pair to europe; the key is 'italy' and the value is data, the dictionary you just built.
# Add data to europe under key 'italy'
# europe['italy']=data
europe.update({'italy':data})
# both works and data as usual can not in ''
# Print europe
print(europe)
| Intermediate Python for Data Science/Dictionaries - Pandas/06-Dictionariception.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Sun Feb 25 22:12:44 2018
@author: miaortizma
"""
import numpy as np
import itertools as it
import matplotlib.pyplot as plt
import random
'''
Returns the array after insertion
and the number of comparations, swaps and other instructions made
'''
def insert(arr, j):
i = j
while(i > 0 and arr[i] < arr[i-1]):
arr[i], arr[i-1] = arr[i-1],arr[i]
i = i - 1
if(i == 0):
comparations = 2*(j) + 1
others = 1 + (j - 1)
elif(i == j):
comparations = 2
else:
comparations = 2*(j - i)
swaps = j - i
others = 1 + (j - i)
return arr, (comparations, swaps, others)
def insertionSort(toSort):
n = len(toSort)
comparations = 0
swaps = 0
others = 0
for i in range(1,n):
toSort, temp = insert(toSort, i)
comparations = comparations + temp[0]
swaps = swaps + temp[1]
others = others + temp[2]
return (comparations, swaps, others)
# +
def graph(comparations, swaps, others):
N = len(comparations)
print("Average:")
print("Comparations:", sum(comparations)/N)
print("Swaps:", sum(swaps)/N)
print("Others:", sum(others)/N)
print("Distribucion:")
plt.hist(comparations,bins='auto')
plt.title("Histogram of Comparations")
plt.show()
plt.hist(swaps,bins='auto')
plt.title("Histogram of Swaps")
plt.show()
plt.hist(others,bins='auto')
plt.title("Histogram of Others")
plt.show()
allOps = comparations + swaps + others
plt.title("Histogram of all operations")
plt.hist(allOps, bins='auto')
plt.show()
'''
Referencia Heap's Algorithm - wikipedia
'''
def generate(n):
A = list(range(1,n+1))
c = [0]*n
yield A
i = 0
while i < n:
if c[i] < i:
if i%2 == 0:
A[0],A[i] = A[i], A[0]
else:
A[c[i]],A[i] = A[i], A[c[i]]
yield A
c[i] = c[i] + 1
i = 0
else:
c[i] = 0
i = i + 1
def shuffle(A):
A = list(A)
n = len(A)
for i in range(n-1):
j = random.randint(i,n-1)
A[j],A[i] = A[i], A[j]
return A
# +
def allPerms(n, perms):
if(n > 8):
print("Input too big, try a number like 8 or less")
comparations = []
swaps = []
others = []
for perm in perms:
temp = insertionSort(list(perm))
comparations = comparations + [temp[0]]
swaps = swaps + [temp[1]]
others = others + [temp[2]]
graph(comparations, swaps, others)
def samplePerms(n, m, rnd):
comparations = []
swaps = []
others = []
rng = range(n)
for i in range(m):
nxt = rnd(rng)
temp = insertionSort(nxt)
comparations = comparations + [temp[0]]
swaps = swaps + [temp[1]]
others = others + [temp[2]]
graph(comparations, swaps, others)
# -
n = 5
allPerms(n,it.permutations(range(n)))
#allPerms(n,generate(n))
# Se observa una distribución normal en cada una de las variables.
#
# El limite para una memoria RAM de 6Gb en la generación de todas las permutaciones es 11, más de 8 ya se demoran mucho.
n = 20
samples = 10000
samplePerms(n, samples, shuffle)
#samplePerms(n, samples, np.random.permutation)
| insertionSort/insertion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/meghanareddyappireddygari/calendar-in-python/blob/main/calendar.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="Y7_gQ-vGUdd_" outputId="842111a9-7157-41db-ab62-507cc2d79e15"
import calendar
yy = 2021
mm = 11
print(calendar.month(yy,mm))
# + colab={"base_uri": "https://localhost:8080/"} id="YFxjdUMpU8wx" outputId="ffcd5758-183d-4410-db01-5bc9d56f28b2"
yy = 1923
print(calendar.calendar(yy))
| calendar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stop Detection
#
# <img align="right" src="https://anitagraser.github.io/movingpandas/pics/movingpandas.png">
#
# [](https://mybinder.org/v2/gh/anitagraser/movingpandas/master?filepath=tutorials/4-stop-detection.ipynb)
#
# **<p style="color:#e31883">This notebook demonstrates the current development version of MovingPandas.</p>**
#
# For tutorials using the latest release visit https://github.com/anitagraser/movingpandas-examples.
#
# ## Setup
# +
import pandas as pd
import geopandas as gpd
from datetime import datetime, timedelta
import sys
sys.path.append("..")
import movingpandas as mpd
print(mpd.__version__)
import warnings
warnings.simplefilter("ignore")
# -
FSIZE = 350
# ## Loading Geolife Sample
# %%time
df = gpd.read_file('data/demodata_geolife.gpkg')
print("Finished reading {} rows".format(len(df)))
traj_collection = mpd.TrajectoryCollection(df, 'trajectory_id', t='t')
traj_collection
# ## Stop Detection with a SingleTrajectory
#
# There are no definitive answers when it comes to detecting / extracting stops from movement trajectories. Due to tracking inaccuracies, movement speed rarely goes to true zero. GPS tracks, for example, tend to keep moving around the object's stop location.
#
# Suitable stop definitions are also highly application dependent. For example, an application may be interested in analyzing trip purposes. To do so, analysts would be interested in stops that are longer than, for example, 5 minutes and may try to infer the purpose of the stop from the stop location and time. Shorter stops, such as delays at traffic lights, however would not be relevant for this appication.
#
# In the MovingPandas **TrajectoryStopDetector** implementation, a stop is detected if the movement stays within an area of specified size for at least the specified duration.
my_traj = traj_collection.trajectories[0]
my_traj
detector = mpd.TrajectoryStopDetector(my_traj)
traj_plot = my_traj.hvplot(title='Trajectory {}'.format(my_traj.id), line_width=7.0, tiles='CartoLight', color='slategray', frame_width=FSIZE, frame_height=FSIZE)
traj_plot
# ### Stop duration
# %%time
stop_durations = detector.get_stop_time_ranges(min_duration=timedelta(seconds=60), max_diameter=100)
for x in stop_durations:
print(x)
# ### Stop points
# %%time
stop_points = detector.get_stop_points(min_duration=timedelta(seconds=60), max_diameter=100)
stop_points
stop_point_plot = traj_plot * stop_points.hvplot(geo=True, size='duration_s', color='deeppink')
stop_point_plot
# ### Stop segments
# %%time
stops = detector.get_stop_segments(min_duration=timedelta(seconds=60), max_diameter=100)
stops
stop_segment_plot = stop_point_plot * stops.hvplot( size=200, line_width=7.0, tiles=None, color='orange')
stop_segment_plot
# ### Split at stops
# %%time
split = mpd.StopSplitter(my_traj).split(min_duration=timedelta(seconds=60), max_diameter=100)
split
for segment in split:
print(segment)
stop_segment_plot + split.hvplot(title='Trajectory {} split at stops'.format(my_traj.id), line_width=7.0, tiles='CartoLight', frame_width=FSIZE, frame_height=FSIZE)
# ## Stop Detection for TrajectoryCollections
#
# The process is the same as for individual trajectories.
# %%time
detector = mpd.TrajectoryStopDetector(traj_collection)
stops = detector.get_stop_segments(min_duration=timedelta(seconds=120), max_diameter=100)
len(stops)
traj_map = traj_collection.hvplot( frame_width=FSIZE, frame_height=FSIZE, line_width=7.0, tiles='CartoLight', color='slategray')
( traj_map *
stops.hvplot( size=200, line_width=7.0, tiles=None, color='deeppink') *
stops.get_start_locations().hvplot(geo=True, size=200, color='deeppink') )
stop_points = detector.get_stop_points(min_duration=timedelta(seconds=120), max_diameter=100)
# +
map1 = ( traj_map * stop_points.hvplot(geo=True, size=200, color='traj_id', title=f'Traj_id type {type(stop_points.iloc[0].traj_id)}'))
stop_points.traj_id = stop_points.traj_id.astype('str')
map2 = ( traj_map * stop_points.hvplot(geo=True, size=200, color='traj_id', title=f'Traj_id type {type(stop_points.iloc[0].traj_id)}'))
map1 + map2
# -
# ## Continue exploring MovingPandas
#
# 1. [Getting started](1-getting-started.ipynb)
# 1. [Reading data from files](2-reading-data-from-files.ipynb)
# 1. [Trajectory aggregation (flow maps)](3-generalization-and-aggregation.ipynb)
# 1. [Stop detection](4-stop-detection.ipynb)
| tutorials/4-stop-detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.6 64-bit
# language: python
# name: python38664bitdc5bcfc3f08f4be2983655d129be0c8b
# ---
import numpy as np
import seaborn as sns
import pandas as pd
import pandas_profiling
import matplotlib.pyplot as plt
data=pd.read_csv("/home/hemanth/Downloads/kc_house_data.csv(1)/kc_house_data.csv")
data.head()
#checking for categorical data
print(data.dtypes)
data.describe()
data.isna().sum()
#dropping the id and date column
dataset = data.drop(['id','date'], axis = 1)
#understanding the distribution with seaborn
with sns.plotting_context("notebook",font_scale=2.5):
g = sns.pairplot(dataset[['sqft_lot','sqft_above','price','sqft_living','bedrooms']],
hue='bedrooms', palette='tab20',size=6)
g.set(xticklabels=[]);
#separating independent and dependent variable
X = dataset.iloc[:,1:].values
y = dataset.iloc[:,0].values
X
y
#splitting dataset into training and testing dataset
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 1/3, random_state = 0)
# +
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
# Predicting the Test set results
y_pred = regressor.predict(X_test)
# +
#Backward Elimination
#import statsmodels.formula.api as sm
import statsmodels.api as sm
def backwardElimination(x, SL):
numVars = len(x[0])
temp = np.zeros((21613,19)).astype(int)
for i in range(0, numVars):
regressor_OLS = sm.OLS(y, x).fit()
maxVar = max(regressor_OLS.pvalues).astype(float)
adjR_before = regressor_OLS.rsquared_adj.astype(float)
if maxVar > SL:
for j in range(0, numVars - i):
if (regressor_OLS.pvalues[j].astype(float) == maxVar):
temp[:,j] = x[:, j]
x = np.delete(x, j, 1)
tmp_regressor = sm.OLS(y, x).fit()
adjR_after = tmp_regressor.rsquared_adj.astype(float)
if (adjR_before >= adjR_after):
x_rollback = np.hstack((x, temp[:,[0,j]]))
x_rollback = np.delete(x_rollback, j, 1)
print (regressor_OLS.summary())
return x_rollback
else:
continue
regressor_OLS.summary()
return x
SL = 0.05
X_opt = X[:, [0, 1, 2, 3, 4, 5,6,7,8,9,10,11,12,13,14,15,16,17]]
X_Modeled = backwardElimination(X_opt, SL)
# -
plt.figure(figsize=(16, 6))
heatmap = sns.heatmap(dataset.corr(), vmin=-1, vmax=1, annot=True, cmap='BrBG')
heatmap.set_title('Correlation Heatmap', fontdict={'fontsize':18}, pad=12);
plt.savefig('heatmap.png', dpi=300, bbox_inches='tight')
| Multiple_LR/Multiple_linear_Regression_kc_house_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 4) Preprocessing
# <h3>Table of Contents<span class="tocSkip"></span></h3>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Feature-Selection" data-toc-modified-id="Feature-Selection-1.1">Feature Selection</a></span></li><li><span><a href="#Scaling-and-one-hot-encoding-the-data" data-toc-modified-id="Scaling-and-one-hot-encoding-the-data-1.2">Scaling and one-hot encoding the data</a></span></li><li><span><a href="#Splitting-and-resampling" data-toc-modified-id="Splitting-and-resampling-1.3">Splitting and resampling</a></span></li></ul></div>
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from imblearn.over_sampling import SMOTE
pirate_data = pd.read_csv(f"data/pirate_data_visualized.csv", index_col = 0)
pirate_data['DATE (LT)'] = pd.to_datetime(pirate_data['DATE (LT)'])
pirate_data.head()
# ### Feature Selection
# +
# The variable "Incident Type" would cause data leakage.
# Others are too granular, have a lot of missing values or were created for visualization.
modelling_df = pirate_data.drop(columns=["DATE (LT)", "INCIDENT TYPE", "COUNTRY", "ATTACKS",
"YEAR_MONTH", "X", "Y", "datestring", "color", "REGION_NR"])
modelling_df["WEEK"] = modelling_df["WEEK"].astype(int)
modelling_df.dtypes
# -
# ### Scaling and one-hot encoding the data
# +
from sklearn.preprocessing import minmax_scale
modelling_df["LAT"] = minmax_scale(X=modelling_df["LAT"])
modelling_df["LONG"] = minmax_scale(X=modelling_df["LONG"])
OHE = pd.get_dummies(modelling_df[["REGION", "VESSEL TYPE", "VESSEL ACTIVITY LOCATION",
"TIME OF DAY", "YEAR", "MONTH", "WEEK", "DAY", "WEEKDAY"]])
modelling_df_merged = pd.concat([modelling_df, OHE], axis = 1, sort = False)
# -
modelling_df_merged = modelling_df_merged.drop(columns = ["REGION", "VESSEL TYPE", "VESSEL ACTIVITY LOCATION",
"TIME OF DAY", "YEAR", "MONTH", "WEEK", "DAY",
"WEEKDAY"])
modelling_df_merged.head()
# Now that all the variables are numeric, we can easily check for correlations between them.
# +
variables = modelling_df_merged
corr = round(variables.corr(),1)
mask = np.triu(np.ones_like(corr, dtype=bool))
fig, ax = plt.subplots(figsize=(15, 15))
cmap = sns.diverging_palette(230, 20, as_cmap=True)
sns.heatmap(corr, mask=mask, cmap= cmap, center=0,
square=True, linewidths=.05, cbar_kws={"shrink": .5}, annot = True);
# -
# Some of the regions are correlated with the longitude, but that was to be expected. Also, it makes sense that an unknown vessel type is correlated with an unknown vessel location of the attack. Missing information appears to affect several variables of the same rows at once. Since the correlations are not very strong, it is not necessary to drop any of the features.
# ### Splitting and resampling
# Add the date back in for the train/test split.
modelling_df_merged_date = pd.concat([modelling_df_merged, pirate_data["DATE (LT)"]], axis = 1, sort = False)
# +
# Split into independent and dependent variables.
X = modelling_df_merged_date.loc[:, modelling_df_merged_date.columns != 'ATTACK SUCCESS']
y = modelling_df_merged_date[["DATE (LT)", "ATTACK SUCCESS"]]
# Then assign data from the last three months to the test set. The rest is training data.
X_train = X[X["DATE (LT)"] < "2016-04-01"]
X_test = X[X["DATE (LT)"] >= "2016-04-01"]
y_train = y[y["DATE (LT)"] < "2016-04-01"]
y_test = y[y["DATE (LT)"] >= "2016-04-01"]
X_train.shape, X_test.shape
# -
# Drop the date column again.
X_train = X_train.drop(columns = "DATE (LT)")
X_test = X_test.drop(columns = "DATE (LT)")
y_train = y_train.drop(columns = "DATE (LT)")
y_test = y_test.drop(columns = "DATE (LT)")
y_train.value_counts(), y_test.value_counts()
# The data set is unbalanced.
## Use SMOTE to handle the class imbalance.
X_train_res, y_train_res = SMOTE().fit_resample(X_train, y_train)
X_train_res.shape, y_train_res.shape
y_train_res.value_counts(), y_test.value_counts()
X_train_res.to_csv("data/X_train_res.csv")
y_train_res.to_csv("data/y_train_res.csv")
X_test.to_csv("data/X_test.csv")
y_test.to_csv("data/y_test.csv")
| notebooks/4-Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from helpers import *
from implementations import ridge_regression
from cross_validation import cross_validation_ridge
from functions import abs_dif, mult
from feature_eng import build_x
from predictions import predict_labels
# +
# Loading the data sets
y_train, x_brute_train, _ = load_csv_data("train.csv")
_, x_brute_test, indices_test = load_csv_data("test.csv")
x_brute = np.concatenate((x_brute_train, x_brute_test))
train_size = x_brute_train.shape[0]
test_size = x_brute_test.shape[0]
# Constants
PHI_features = [15, 18, 20, 25, 28]
invalid_value = -999
# Mask to subdivide in different models
# Mask for the data (rows)
data_masks = [
x_brute[:, 22] == 0,
x_brute[:, 22] == 1,
x_brute[:, 22] > 1
]
num_models = len(data_masks)
# Mask for the features (columns)
features_masks = [(x_brute[m].std(axis=0) != 0) & np.any(x_brute[m] != -999., axis=0) & ~range_mask(30, PHI_features) for m in data_masks]
# Separate X and Y using the masks
ys_train = [y_train[mask[:train_size]] for mask in data_masks]
xs_brute_train = [x_brute_train[d_m[:train_size]][:, f_m] for d_m, f_m in zip(data_masks, features_masks)]
xs_brute_test = [x_brute_test[d_m[train_size:]][:, f_m] for d_m, f_m in zip(data_masks, features_masks)]
# +
lambdas = [1e-03, 3.16e-03, 1e-02]
k_fold = 4
num_models = len(data_masks)
# Models variables
degrees = [9, 11, 12]
roots = [3, 4, 3]
tanh_degrees = [3, 4, 3]
log_degrees = [3, 4, 3]
inv_log_degrees = [3, 4, 3]
fn_tanh = [True] * num_models
fn_log = [False] * num_models
fn_inv_log = [True] * num_models
functions = [[mult, abs_dif],] * num_models
def build_ith_x(i):
return build_x(xs_brute_train[i], xs_brute_test[i], degrees[i], roots[i], log_degree=log_degrees[i], tanh_degree=tanh_degrees[i],
inv_log_degree=inv_log_degrees[i], fn_log=fn_log[i], fn_inv_log=fn_inv_log[i], fn_tanh=fn_tanh[i],
functions=functions[i], print_=True)
# -
# # Cross Validation
# +
final_scores = []
ys_sub = []
iters = 2
for i in range(len(data_masks)):
x_train, x_test = build_ith_x(i)
print("x[{}] DONE".format(i))
w, _ = ridge_regression(ys_train[i], x_train, lambdas[i])
ys_sub.append(predict_labels(w, x_test))
scores = []
for it in range(iters):
score = cross_validation_ridge(ys_train[i], x_train, k_fold, lambdas[i], seed=100+it)
score *= 100
scores.extend(score)
final_scores.append(scores)
del x_train
del x_test
final_scores
# -
all_scores = np.array(final_scores)
def avg_score(scores):
return (np.array(scores) * np.array(data_masks).T.sum(axis=0) / (train_size + test_size)).sum()
# # SCORE
np.mean([avg_score(all_scores[:, i]) for i in range(all_scores.shape[1])])
# # STD
np.std([avg_score(all_scores[:, i]) for i in range(all_scores.shape[1])])
# # Submission
# +
y_submission = np.zeros(test_size)
for y, mask in zip(ys_sub, data_masks):
mask = mask[train_size:]
y_submission[mask] = y
create_csv_submission(indices_test, y_submission, "final_submission.csv")
| project01/src/cv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import mysql.connector
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from patsy import dmatrices
from sklearn.cross_validation import train_test_split
from sklearn import metrics
from sklearn.cross_validation import cross_val_score
from sklearn.tree import export_graphviz
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
# +
rds_host='mybike.c0jxuz6r8olg.us-west-2.rds.amazonaws.com'
name='hibike'
pwd='<PASSWORD>'
db_name='bike'
port=3306
conn = mysql.connector.connect(host=rds_host,user=name, password=<PASSWORD>, database=db_name)
df = pd.read_sql("select * from weather", con=conn)
df1 = pd.read_sql("select * from station_1", con=conn)
df.shape
# +
df['month'] = df['last_update'].dt.month
df['day'] = df['last_update'].dt.day
df['weekday'] = df['last_update'].dt.weekday_name
df['hour'] = df['last_update'].dt.hour
df['minute'] = df['last_update'].dt.minute
df1['month'] = df1['last_update'].dt.month
df1['day'] = df1['last_update'].dt.day
df1['weekday'] = df1['last_update'].dt.weekday_name
df1['hour'] = df1['last_update'].dt.hour
df1['minute'] = df1['last_update'].dt.minute
# -
df1.minute = (df1.minute > 30) *30
df2 = pd.merge(df, df1, left_on=['minute', 'hour', 'weekday', 'month', 'day'], right_on=['minute', 'hour', 'weekday', 'month', 'day'])
df2
df2 = df2.drop(['last_update_x'] , axis=1)
categorical_columns = df2[['weather_main', 'weekday', 'status']]
for column in categorical_columns:
df2[column] = df2[column].astype('category')
df2.select_dtypes(['category']).describe().T
# +
# Select columns containing continuous data
continuous_columns = []
for column in df2.columns.values:
if column not in categorical_columns:
continuous_columns.append(column)
df2['humidity'] = df2['humidity'].astype('float')
continuous_columns.pop(continuous_columns.index('last_update_y'))
print(df2[continuous_columns].dtypes)
# -
df_dummies_weather_main = pd.get_dummies(df2[['weather_main']])
print(df_dummies_weather_main.head())
df_dummies_weekday = pd.get_dummies(df2[['weekday']])
describe_columns = [x for x in continuous_columns if x not in ['available_bike_stands', 'bike_stands', 'available_bikes', 'visibility', 'day', 'month']]
describe_columns
X = pd.concat([df2[describe_columns], df_dummies_weather_main, df_dummies_weekday], axis =1)
y = df2.available_bikes
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
print("Training data:\n", pd.concat([X_train, y_train], axis=1))
print("\nTest data:\n", pd.concat([X_test, y_test], axis=1))
# -
# Train RF with 100 trees
rfc = RandomForestClassifier(n_estimators=100, max_features='auto', oob_score=True, random_state=1)
rfc.fit(X, y)
pd.DataFrame({'feature': X.columns, 'importance':rfc.feature_importances_})
rfc.fit(X_train, y_train)
rfc_predictions_test = rfc.predict(X_test)
df_true_vs_rfc_predicted_test = pd.DataFrame({'ActualClass': y_test, 'PredictedClass': rfc_predictions_test})
df_true_vs_rfc_predicted_test
print("Accuracy: ", metrics.accuracy_score(y_test, rfc_predictions_test))
# +
import mysql.connector
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from patsy import dmatrices
from sklearn.cross_validation import train_test_split
from sklearn import metrics
from sklearn.cross_validation import cross_val_score
from sklearn.tree import export_graphviz
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
rds_host='mybike.c0jxuz6r8olg.us-west-2.rds.amazonaws.com'
name='hibike'
pwd='<PASSWORD>'
db_name='bike'
port=3306
import pandas as pd
import requests
from datetime import datetime
# +
diCol = [x for x in X.columns]
diCol
# +
conn = mysql.connector.connect(host=rds_host,user=name, password=<PASSWORD>, database=db_name)
cur = conn.cursor()
cur.execute("select * from Bike_Stations")
stationData = cur.fetchall()
import datetime
PredictTime = -40
stationNum = 100
cur.execute('select * from station_1 ORDER BY last_update DESC limit 0, 1')
detailData = cur.fetchone()
cur.execute('select * from weather ORDER BY last_update DESC limit 0, 1')
weatherData = cur.fetchone()
print(weatherData)
diCol_pr = {}
global diCol
global di
futTime = ((datetime.datetime.now() - weatherData[0]).total_seconds() / 60 ) + float(PredictTime)
print(futTime)
if (futTime > 180) :
stationNum = int(stationNum)
print('>180')
try:
for i in diCol[stationNum]:
diCol_pr[i] = 0
r_w = requests.get('http://api.openweathermap.org/data/2.5/forecast?q=Dublin,IE&APPID=7b3e054e55cf81602b4298ca04a0fa18')
diCol_pr['hour'] = datetime.datetime.now().hour + PredictTime // 60
diCol_pr['minute'] = datetime.datetime.now().minute + datetime.datetime.now().hour * 60 + PredictTime
diCol_pr['humidity'] = r_w.json()['list'][0]['main']['humidity']
diCol_pr['temp'] = r_w.json()['list'][0]['main']['temp']
diCol_pr['wind'] = r_w.json()['list'][0]['wind']['speed']
weekday = datetime.datetime.now().weekday() + (datetime.datetime.now().hour + (datetime.datetime.now().minute + PredictTime) // 60) // 24
if weekday >= 7:
weekday = 0
weather = r_w.json()['list'][0]['weather'][0]['main']
print(weather)
weekdays = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
weekday = weekdays[weekday]
if 'weather_main_{}'.format(weatherData[1]) in diCol_pr.keys():
diCol_pr['weather_main_{}'.format(weather)] = 1
if 'weekday_{}'.format(weekday) in diCol_pr.keys():
diCol_pr['weekday_{}'.format(weekday)] = 1
predi = pd.DataFrame([diCol_pr])
rfc_predictions = "available_bikes : {}".format(int(rfx.predict(predi)))
print(rfc_predictions , diCol_pr, diCol[stationNum])
else:
rfc_predictions = "Something wrong, day information get error."
else:
rfc_predictions = "A new weather may appear, not enough data to make a prediction."
except Exception as e:
rfc_predictions = "We rebuild the prediction module, please try 10 minutes late."
else :
print('<180')
stationNum = int(stationNum)
for i in diCol:
diCol_pr[i] = 0
diCol_pr['hour'] = datetime.datetime.now().hour + (datetime.datetime.now().minute + PredictTime) // 60
diCol_pr['minute'] = (datetime.datetime.now().minute + PredictTime) % 60
diCol_pr['humidity'] = float(weatherData[2])
diCol_pr['temp'] = weatherData[3]
diCol_pr['wind'] = weatherData[5]
weekday = datetime.datetime.now().weekday() + (datetime.datetime.now().hour + (datetime.datetime.now().minute + PredictTime) // 60) // 24
if weekday >= 7:
weekday = 0
weekdays = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
weekday = weekdays[weekday]
if 'weather_main_{}'.format(weather) in diCol_pr.keys():
diCol_pr['weather_main_{}'.format(weatherData[1])] = 1
if 'weekday_{}'.format(weekday) in diCol_pr.keys():
diCol_pr['weekday_{}'.format(weekday)] = 1
predi = pd.DataFrame([diCol_pr])
print(rfc.predict(predi))
rfc_predictions = "available_bikes : {}".format(int(rfc.predict(predi)))
print(rfc_predictions , diCol_pr)
else:
rfc_predictions = "Something wrong, day information get error."
else:
rfc_predictions = "A new weather may appear, not enough data to make a prediction."
# -
| src/hibike/.ipynb_checkpoints/Untitled2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Map
# The map function applies a supplied function to the elements in a sequence. The syntax is as follows:
#
# map(function, sequence)
#
# In return, the map sends a list of the sequences transformed by the function sent in. An easy, practical example to cover is conversions. We can convert the items in a sequence to what we want to, without leaving the list, with the map() function. The function we write will not take the sequence as a whole. You must write the function as if you are taking a single element from the sequence.
# **Example 1** Transforming a sequence of day values into hours.
# +
# Function to apply
def days(hour):
return hour/24
# Sequence to transform
hour_list = [4.0, 24.0, 47, 72.0]
# Map
days_list = list(map(days, hour_list))
# -
print(days_list)
# It's a great time to start using lambdas. Lambda expressions were made for operations like the map() function. Let's try it out in the next example.
# **Example 2** You have a database of male names, and you need to send letters out to said people. Add the prefix "Mr." to every man's name.
# +
# Sequence to transform
male_names = ["Rick", "Rocky", "Charles", "Peter"]
# Map
mr_male_names = list(map(lambda name: "Mr. " + name, male_names))
# -
print(mr_male_names)
# ## Applying Map to Multiple Sequences
# You can apply map() to multiple sequences. A few good, immediate examples are combining sequences in some shape or form. Imagine you have the total scores of 3 players from 3 rounds, where each player X score is item X. If we want the total score, we have to add them all up using bracket notation.
# +
# [player1score, player2score, player3score]
round_1 = [8, 6, 9]
round_2 = [5, 7, 7]
round_3 = [8, 9, 6]
round_totals = list(map(lambda r_1, r_2, r_3: r_1 + r_2 + r_3, round_1, round_2, round_3))
# -
print(round_totals)
| Built-In Functions/map().ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
data_frame = pd.read_csv('dataset.csv')
data_frame.head(10)
sns.heatmap(data_frame.isnull())
Y = pd.DataFrame(data=data_frame,columns=['id','diagnosis'])
X = data_frame.drop(['id','diagnosis'],axis=1)
X_train, X_test, Y_train, Y_test= train_test_split(X,Y,test_size=0.3)
from sklearn.naive_bayes import MultinomialNB
from sklearn.tree import DecisionTreeClassifier,DecisionTreeRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.neighbors import KNeighborsClassifier
Y_test.drop('id',axis=1,inplace=True)
Y_train.drop('id',axis=1,inplace=True)
# # NB
from sklearn.naive_bayes import GaussianNB
nb=GaussianNB()
nb.fit(X_train,Y_train)
NB_predicted = nb.predict(X_test)
predicted
from sklearn.metrics import classification_report,confusion_matrix
print(confusion_matrix(Y_test,NB_predicted))
print(classification_report(Y_test,NB_predicted))
df = pd.DataFrame({'ID':id_pred,'NB_predictions':NB_predicted})
df['NB_predictions'] = df['NB_predictions'].map({1: 'M', 0: 'B'})
export_csv = df.to_csv ('NB_Prediction.csv', index = False, header=True)
# # Decision Tree Classifier
#
from sklearn.tree import DecisionTreeClassifier
dtree = DecisionTreeClassifier()
dtree.fit(X_train,Y_train)
DecisionTree_pred = dtree.predict(X_test)
print(confusion_matrix(Y_test,DecisionTree_pred))
print(classification_report(Y_test,DecisionTree_pred))
# # Decision Tree Regressor
from sklearn.tree import DecisionTreeRegressor
dtree = DecisionTreeClassifier()
dtree.fit(X_train,Y_train)
DecisionTree_pred = dtree.predict(X_test)
print(confusion_matrix(Y_test,DecisionTree_pred))
print(classification_report(Y_test,DecisionTree_pred))
# # KNN
# +
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=33)
knn.fit(X_train,Y_train)
KNN_pred = knn.predict(X_test)
print(confusion_matrix(Y_test,KNN_pred))
print(classification_report(Y_test,KNN_pred))
# -
| Breast Cancer/Breast Cancer Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.0
# language: julia
# name: julia-1.4
# ---
# +
using Plots
using Distributions
include("../src/MPLSH.jl")
using MPLSH
a = rand(MvNormal([5, 5], [[1, .9] [.9, 1]]), 100)'
b = a * [-1 0; 0 1]
c = a * [-5 .1; .1 -2]
a = rand(MvNormal([5, 5], [[1, .9] [.9, 1]]), 50)'
d = a * [-2 .5; .1 -1]
e = a * [-.5 0; 0 1] .- [-5 4]
a = rand(MvNormal([5, 5], [[1, .9] [.9, 1]]), 20)'
f = a * [-.2 .5; .5 -.5] .- [-1 3]
# println("shapes ", size(a), size(b))
data = vcat(a, b, c, d, e, f)
# scatter(data[:, 1], data[:, 2], legend=false)
lsh_base = LSHBase(2, 1., 5)
lsh = LSH(lsh_base, data)
query = [-6, -2.5]
# Get the 10 nearest neighbors.
nns = nearest_neighbors(lsh, query, 10)
print(typeof(nns))
print(nns)
scatter(data[:, 1], data[:, 2], alpha=.5, legend=false)
scatter!(nns[:, 1], nns[:, 2], color="red")
scatter!([query[1]], [query[2]], color="yellow")
# -
nns = nearest_neighbors(lsh, query, 1)
print(typeof(nns))
print(nns)
scatter(data[:, 1], data[:, 2], alpha=.5, legend=false)
scatter!(nns[:, 1], nns[:, 2], color="red")
scatter!([query[1]], [query[2]], color="yellow")
| test/test_MPLSH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github" slideshow={"slide_type": "-"}
# <a href="https://colab.research.google.com/github/cohmathonc/biosci670/blob/master/IntroductionComputationalMethods/00_CompWorkingEnv.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="kQws6d4Wan4s" slideshow={"slide_type": "-"}
# # Computational Environment
# + [markdown] slideshow={"slide_type": "-"}
# While simple mathematical models can be analyzed analytically, finding analytic solution becomes more difficult with increasing model complexity. Also, for many problems, analytic solutions simply do not exist.
#
# [*Numerical analysis*](https://www.britannica.com/science/numerical-analysis) is an area of mathematics and computer science that studies how *numerical solutions* to such problems can be obtained.
# These methods rely on [*algorithms*](https://www.britannica.com/science/algorithm), systematic procedures that produce the solution,or approximate the solution to some well-characterized level of accuracy.
# While those procedures can be carried out manually in principal, in practice this may not be feasible due to the large number of computational steps involved.
# Computers facilitate these repetitive arithmetic tasks.
#
# The application of computational tools and simulation to solve problems in various scientific disciplines is often called [*scientific computing*](https://en.wikipedia.org/wiki/Computational_science).
# -
# ## 'Programs' and 'Programming Languages'
# **A program is a sequence of instructions that specifies how to perform a computation.**
#
# Details will depend on the kind of computation, but typically the following basic instructions are involved:
#
# - *input*: Get data.
# - *output*: Display data.
# - *math*: Perform mathematical operations.
# - *conditional execution*: Check for conditions and selectively run specific code.
# - *repetition*: Perform some action repeatedly.
#
# Programs are specified in *formal languages*, that is, languages that are designed for specific applications.
# For example, the notation of mathematics is a formal language for denoting relationships among numbers and symbols.
#
# **Programming languages are formal languages designed to express computations.**
# + [markdown] slideshow={"slide_type": "-"}
# ## Common Programming Language Options for Scientific Computing
# -
#
# [Many](https://en.wikipedia.org/wiki/List_of_programming_languages) (!) [programming languages](https://en.wikipedia.org/wiki/Programming_language) have been designed, and most of them support the basic operations needed for numeric computing.
#
# When choosing a programming languages for scientific applications, computational speed during execution is often an important criterium. Other criteria may include whether libraries are available that facilitate specific computational tasks or whether the language implementation is open source or proprietary.
#
# Computationally intensive aspects of scientific computing typically rely on [*compiled*](https://en.wikipedia.org/wiki/Compiled_language) *general purpose languages*, such as [*C*](https://en.wikipedia.org/wiki/C_(programming_language)) or [*Fortran*](https://en.wikipedia.org/wiki/Fortran) and optimized software libraries for numeric computing.
# When computational speed is not the foremost concern, [*interpreted*](https://en.wikipedia.org/wiki/Interpreted_language) languages are usually preferred because they allow a more 'interactive' style of programming which facilitates program development.
#
# Various specialized interpreted languages have been developed for scientific applications.
# They typically originate from a specific user community and provide abstractions and functionalities tailored to its scientific domain.
# Often, they provide in-built functionality for data visualization.
# Common examples include:
#
# * [Matlab](https://www.mathworks.com/products/matlab.html): Focus on numerical computing for science & engineering. Proprietary, requires license.
# * [Octave](https://www.gnu.org/software/octave/): Numerical computation, high compatibility to Matlab, Open source.
# * [Scilab](https://www.scilab.org): Numerical computation, Open source.
# * [R](https://www.r-project.org): Focus on statistical computing. Open source.
# * [Wolfram Mathematica](https://www.wolfram.com/mathematica/): Many areas of technical computing. Proprietary, requires license.
#
# Also the following *general purpose languages* have become increasingly popular for scientific applications:
#
# * [Python](https://www.python.org): General purpose, Open source.
# * [Julia](https://julialang.org): General, some focus on scientific and numerical computing, Open source.
#
# These languages offer a similar interactive programming experience as the specialized languages listed above.
# Although designed as general purpose languages, a large ecosystem of 'external' libraries allows their functionality to be easily extended for specific aspects of scientific computing.
#
#
# ## Python
# We will use [Python](https://www.python.org) to introduce basic programming concepts and for most of the computational exercises in this course.
#
# Python is an **high-level**, **interpreted**, **general-purpose** programming language.
# Its reference implementation is **open source** software managed by the [Python Software Foundation](https://www.python.org/psf/).
# Development on Python started in the late 1980s and the very first release dates back to 1991.
# In case you wonder about the name, it has nothing to do with reptiles but is a tribute to the [Monty Python Flying Circus](https://en.wikipedia.org/wiki/Monty_Python) ([see FAQ](https://docs.python.org/2/faq/general.html#why-is-it-called-python)).
#
# Python has a large [standard library](https://docs.python.org/3/library/) that provides functionalities for a wide range of application domains.
# Most extensions and community-contributed software modules are indexed in the [Python Package Index](https://pypi.org), the official repository of third-party Python libraries.
# This rich ecosystem makes Python one of the [most popular](https://www.tiobe.com/tiobe-index/) programming languages today.
#
# Two major release versions of Python are commonly being used today, version `2.x` (typically `2.7.x`) and version `3.x` (latest `3.7.x`). We will be using version `3.7`. Refer to this [discussion](https://wiki.python.org/moin/Python2orPython3) about differences between those versions.
#
# ### Python Installation Options
# First, note that there is **no need to install Python** for this course; see *Python without Installation* below.
#
# The basic Python interpreter comes pre-installed on most linux systems and on MacOS. You can check whether it is available on your system by typing `python` in a terminal or command line window.
#
# If you do want to set up a local installation and this is your first Python experience, I would recommend to install [ANACONDA](https://www.anaconda.com/download) which provides a full featured Python environment that is [easy to install](https://docs.anaconda.com/anaconda/install/). Choose the `Python 3.x` installation option.
# ANACONDA bundles the basic python interpreter with additional tools, such as [IPython](https://ipython.org), an interactive interpreter, [Spyder](https://www.spyder-ide.org), an *integrated development environment* (IDE) for scientific python programming (similar to Matlab or RStudio for R), and [Jupyter](https://jupyter.org) for browser-based interactive programming notebooks.
# Jupyter Notebooks can contain live code, equations, visualizations and narrative text. They can be shared as static (see [nbviewer](https://nbviewer.jupyter.org) examples) or interactive (see this [example](https://mybinder.org/v2/gh/jupyterlab/jupyterlab-demo/master?urlpath=lab%2Ftree%2Fdemo%2FLorenz.ipynb)) documents containing text and executable code blocks.
#
# Information about installing the basic python interpreter is available [here](https://wiki.python.org/moin/BeginnersGuide/Download).
# ### Python without Installation
# + [markdown] colab_type="text" id="iKM56f47jzsN"
# One advantage of using a popular open-source programming language is the availability of free tools and services, including various options for editing and programming online.
#
# For example, [pythonanywhere](https://www.pythonanywhere.com) gives you access to an online Python or IPython console.
# [Colab](https://colab.research.google.com) is a Google service that is compatible to [Jupyter](https://jupyter.org) Notebooks and allows these notebooks to be opened, edited and executed online. Notebooks can be saved to Google Drive, be shared like other files on google drive and, of course, be downloaded.
# Microsoft offers a similar service called [Azure Notebooks](https://notebooks.azure.com).
# [Binder](https://mybinder.org) is another project that can turn Jupyter Notebooks into an executable online-environment.
#
# **We will be using Jupyter Notebooks in this course**, the **easiest way to access and edit** these Notebooks is probably via **Colab**. For more information, see the [Colab introduction](https://colab.research.google.com/notebooks/welcome.ipynb), particularly the links covering [basic features](https://colab.research.google.com/notebooks/basic_features_overview.ipynb), [text in Colab](https://colab.research.google.com/notebooks/markdown_guide.ipynb) and [plotting in Colab](https://colab.research.google.com/notebooks/charts.ipynb).
#
# + [markdown] colab_type="text" id="645xUBeqpHxr"
# ## Course Materials and Programming Exercises
# + [markdown] colab_type="text" id="Rn5wO3j6pN_1"
# Programming lectures and exercises for the course will be collected in this github repository:
#
# https://github.com/cohmathonc/biosci670coh
#
# The repository contains a readme file, folder structure and various [Jupyter](https://jupyter.org)-Notebook files (`*.ipynb`).
# You can download the entire repository or browse the content of the repository online.
#
# The notebook files can be edited and executed in a local installation, or online using [Google Colab](https://colab.research.google.com/notebooks/welcome.ipynb).
# To open them in Colab, simply click on the  badge that is displayed at the top of each file.
# -
# ## Other Resources
# + [markdown] slideshow={"slide_type": "-"}
# ### General Python
# - [Think Python](http://greenteapress.com/wp/think-python-2e/) is a free book on Python; interactive learning material based on this book, together with code examples is available [here](http://interactivepython.org/runestone/static/CS152f17/index.html).
# - The [official Python documentation](https://docs.python.org/3/), including a very detailed [tutorial](https://docs.python.org/3/tutorial/index.html).
# - [Python 101](https://python101.pythonlibrary.org) provides an in-depth introduction to general programming in python.
# - [The Hitchhiker’s Guide to Python!](https://docs.python-guide.org)
#
# ### Python for Scientific Computing
# - [SciPy](https://www.scipy.org/about.html), python for scientific computing.
# - [Getting Stared with SciPy](https://www.scipy.org/getting-started.html), and the [SciPy lecture notes](http://scipy-lectures.org/index.html).
#
# ### More ...
# See the [Beginner's Guide](https://wiki.python.org/moin/BeginnersGuide) and [Beginner's Guide for Non-Programmers](https://wiki.python.org/moin/BeginnersGuide/NonProgrammers) on the [PythonWiki](https://wiki.python.org) for a wider selection of learning resources.
# -
# ##### About
# This notebook is part of the *biosci670* course on *Mathematical Modeling and Methods for Biomedical Science*.
# See https://github.com/cohmathonc/biosci670coh for more information and material.
| IntroductionComputationalMethods/00_CompWorkingEnv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stochastic degradation model
#
# This example shows how the stochastic degradation model can be used.
# This model describes the stochastic process of a single chemical reaction, in which the concentration of a substance degrades over time as particles react.
# The substance degrades starting from an initial concentration, $n_0$, to 0 following a rate constant, $k$, according to the following model (Erban et al., 2007):
# $$A \xrightarrow{\text{k}} \emptyset$$
#
# The model is simulated according to the Gillespie stochastic simulation algorithm (Gillespie, 1976)
# 1. Sample a random value r from a uniform distribution: $r \sim \text{uniform}(0,1)$
# 2. Calculate the time ($\tau$) until the next single reaction as follows (Erban et al., 2007):
# $$ \tau = \frac{1}{A(t)k} \ln{\big[\frac{1}{r}\big]} $$
# 3. Update the molecule count at time t + $\tau$ as: $ A(t + \tau) = A(t) - 1 $
# 4. Return to step (1) until molecule count reaches 0
#
import pints
import pints.toy
import matplotlib.pyplot as plt
import numpy as np
import math
# Specify initial concentration, time points at which to record concentration values, and rate constant value (k)
# +
n_0 = 20
model = pints.toy.StochasticDegradationModel(n_0)
times = np.linspace(0, 100, 100)
k = [0.1]
values = model.simulate(k, times)
plt.step(times, values)
plt.xlabel('time')
plt.ylabel('concentration (A(t))')
plt.show()
# -
# Given the stochastic nature of this model, every iteration returns a different result. However, averaging the concentration values at each time step, produces a reproducible result which tends towards a deterministic function as the the number of iterations tends to infinity (Erban et al., 2007): $ n_0e^{-kt} $
#
# +
for i in range(10):
values = model.simulate(k, times)
plt.step(times, values)
mean = model.mean(k, times)
plt.plot(times, mean, label = 'deterministic mean of A(t)')
plt.title('stochastic degradation across different iterations')
plt.xlabel('time')
plt.ylabel('concentration (A(t))')
plt.legend(loc = 'upper right')
plt.show()
# -
# The deterministic mean (from above) is plotted with the deterministic standard deviation.
# The deterministic variance of this model is given by: $e^{-2kt}(-1 + e^{kt})n_0$
# +
mean = model.mean(k, times)
variance = model.variance(k, times)
std_dev = np.sqrt(variance)
plt.plot(times, mean, '-', label = 'deterministic mean of A(t)')
plt.plot(times, mean + std_dev, '--', label = 'standard deviation upper bound')
plt.plot(times, mean - std_dev, '--', label = 'standard deviation lower bound')
plt.legend(loc = 'upper right')
plt.xlabel('time')
plt.ylabel('concentration (A(t))')
plt.show()
| examples/toy/model-stochastic-degradation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:deep]
# language: python
# name: conda-env-deep-py
# ---
# # Remote resource monitor
# This ia a remote resource monitor in a notebook. It can be very frustrating to see no output for a long time: is your script actually running or did your kernel die in the background. This remote resource monitor will let you know by plotting the CPU, RAM and video memory.
#
# ## Requirements
# - Matplotlib
# - `pip install matplotlib`
# - PSUtil
# - `pip install psutil`
# - pynvml (for GPU)
# - `pip install py3nvml`
# +
show_gpu_mem = True
show_gpu_mem_max = False
cpu_color = "deepskyblue"
memory_color = "darkorange"
gpu_color = "yellowgreen"
# +
# %matplotlib inline
import psutil
import matplotlib.pyplot as plt
import time
import math
import numpy as np
from IPython import display
if show_gpu_mem:
from py3nvml.py3nvml import *
nvmlInit()
def convert_size(size_bytes):
if size_bytes == 0:
return "0B"
size_name = ("B", "KB", "MB", "GB", "TB", "PB", "EB", "ZB", "YB")
i = int(math.floor(math.log(size_bytes, 1024)))
p = math.pow(1024, i)
s = round(size_bytes / p, 2)
return "%s %s" % (s, size_name[i])
def pad_input(y_series, length):
padding = length - len(y_series)
y_series.extend([None for i in range(padding)])
return y_series
cpu_percent = []
mem_used = []
if show_gpu_mem:
gpu_used = []
gpu_total = nvmlDeviceGetMemoryInfo(nvmlDeviceGetHandleByIndex(0)).total
gpu_yticks = [0.1*gpu_total * i for i in range(0, 11)]
gpu_yticks_label = [""]+[convert_size(tick) for tick in gpu_yticks[1:]]
vm_total, _, _, vm_used, _ = list(psutil.virtual_memory())
cpu_yticks = range(0, 110, 20)
cpu_yticks_label = ["{}%".format(per) for per in cpu_yticks]
mem_yticks = [0.1*vm_total * i for i in range(0, 11)]
mem_yticks_label = ["+"]+[convert_size(tick) for tick in mem_yticks[1:]]
fig, ax1 = plt.subplots()
while True:
try:
# Update values
cpu_percent += [psutil.cpu_percent()]
if len(cpu_percent) > 60:
cpu_percent.pop(0)
mem_used += [list(psutil.virtual_memory())[3]]
if len(mem_used) > 60:
mem_used.pop(0)
if show_gpu_mem:
gpu_used += [nvmlDeviceGetMemoryInfo(nvmlDeviceGetHandleByIndex(0)).used]
if len(gpu_used) > 60:
gpu_used.pop(0)
ax1.set_xlabel("time (s)")
ax1.set_ylabel("CPU percentage", color=cpu_color)
ax1.set_yticks(cpu_yticks)
ax1.set_yticklabels(cpu_yticks_label)
ax1.set_ylim(bottom=0, top=100)
ax1.set_xticklabels([])
ax1.set_xlim(0, len(gpu_used))
ax2 = ax1.twinx()
ax2.set_ylabel("Memory usage", color=memory_color)
ax2.set_yticks(mem_yticks)
ax2.set_yticklabels(mem_yticks_label, )
ax2.set_ylim(bottom=0, top=vm_total)
ax1.plot(range(60), pad_input(cpu_percent, 60), color=cpu_color, label="CPU", ) # fit the line
ax2.plot(range(60), pad_input(mem_used, 60), color=memory_color, label="Memory") # fit the histogram
if show_gpu_mem:
ax2.plot(range(60), pad_input(gpu_used, 60), color=gpu_color, label="GPU")
if show_gpu_mem_max:
ax2.axhline(gpu_total, 0, 60, linestyle="dashed", color=gpu_color)
ax2.annotate("Max GPU", (30, gpu_total+500000000), color=gpu_color)
fig.legend(loc="lower left")
# Update display
display.display(fig)
ax1.clear()
ax1.set_yticks([])
ax2.clear()
ax2.set_yticks([])
time.sleep(0.5) #sleep
display.clear_output(wait=True)
except KeyboardInterrupt:
fig.clear()
display.set_matplotlib_close(True)
display.clear_output(wait=False)
display.display(fig)
break
print("")
# -
| resmon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["hide-input"]
from IPython.display import IFrame
IFrame(src="https://cdnapisec.kaltura.com/p/2356971/sp/235697100/embedIframeJs/uiconf_id/41416911/partner_id/2356971?iframeembed=true&playerId=kaltura_player&entry_id=1_x43aqvxh&flashvars[streamerType]=auto&flashvars[localizationCode]=en&flashvars[leadWithHTML5]=true&flashvars[sideBarContainer.plugin]=true&flashvars[sideBarContainer.position]=left&flashvars[sideBarContainer.clickToClose]=true&flashvars[chapters.plugin]=true&flashvars[chapters.layout]=vertical&flashvars[chapters.thumbnailRotator]=false&flashvars[streamSelector.plugin]=true&flashvars[EmbedPlayer.SpinnerTarget]=videoHolder&flashvars[dualScreen.plugin]=true&flashvars[hotspots.plugin]=1&flashvars[Kaltura.addCrossoriginToIframe]=true&&wid=1_k3es2um8" ,width='800', height='500')
# -
# # Definition of linearly separable sets
#
# In this section, we consider a special class of $k$ linearly separable
# sets for $k\ge 2$. Let us first introduce the following definition for
# binary classification.
#
# For $k=2$, there is a very simple geometric interpretation of two
# linearly separable sets.
#
# ```{admonition} Definition 1
# The two sets $A_1$, $A_2\subset \mathbb{R}^d$ are linearly separable if there exists a
# hyperplane $\label{2classH}
# H_0=\{x:wx+b=0\}$, such that $wx+b>0$ if $x\in A_1$ and $wx+b<0$ if
# $x\in A_2$.
# ```
# ```{figure} ../figures/LinearS1.png
# :name: LinearS1
# :height: 150px
# One linearly separable set
# ```
#
# ```{figure} ../figures/NLinearS2.png
# :name: NLinearS2
# :height: 150px
# Two non-linearly separable sets
# ```
#
# ```{admonition} Lemma 1
# The two sets $A_1$,
# $A_2\subset \mathbb{R}^d$ are linearly separable if there exists
# $W=
# \begin{pmatrix}
# w_1\\
# w_2
# \end{pmatrix} \in \mathbb{R}^{2\times d}$,
# $b=
# \begin{pmatrix}
# b_1\\
# b_2
# \end{pmatrix}$.
# $b\in R^{2\times d}$,
# such that,
# $w_1x+b_1 > w_2x+b_2,\ \forall x\in A_1$ and
# $w_1x+b_1 < w_2x+b_2,\ \forall x\in A_2$
# ```
#
# ```{admonition} Proof
# *Proof.* Here, we can just take $w = w_1 - w_2$ and $b = b_1 - b_2$,
# then we can check that the hyperplane $wx + b$ satisfies the definition
# as presented before. ◻
# ```
#
# Now let us consider multi-class classification. To begin with the
# definition, let us assume that the data space is divided into $k$
# classes represented by $k$ disjoint sets
# $A_1,A_2,\cdots,A_k\subset \mathbb{R}^d$, which means
# $A = A_1\cup A_2\cup \cdots \cup A_k, ~A_i\cap A_j = \emptyset, \forall i \neq j.$
#
# ```{admonition} Definition 2
# A collection of subsets $A_1,...,A_k\subset \mathbb{R}^d$ are linearly
# separable if there exist
# $W=
# \begin{pmatrix}
# w_1\\
# \vdots\\
# w_k
# \end{pmatrix}
# \in \mathbb{R}^{k\times d},
# b=
# \begin{pmatrix}
# b_1\\
# \vdots\\
# b_k
# \end{pmatrix}$
# $\in \mathbb{R}^{k\times d}$,
# such that, for each
# $1\le i\le k$ and
# $j \neq i$ $
# w_ix+b_i > w_jx+b_j,\ \forall x\in A_i.$ namely, each pairs of $A_i$
# and $A_j$ are linearly separable by the plane $
# H_{ij}=\{(w_i-w_j)\cdot x+(b_i-b_j) = 0\}, \quad \forall j\neq i.$
# ```
#
# The geometric interpretation for linearly separable sets is less obvious
# when $k>2$.
#
# ```{admonition} Lemma 2
# Assume that
# $A_1,...,A_k$ are linearly separable and $W\in
# \mathbb{R}^{k\times d}$ and $b\in\mathbb{R}^k$ satisfy
# . Define
# $
# \Gamma_i(W,b) = \{x\in\mathbb R^d: (Wx+b)_i > (Wx+b)_j,\ \forall j \neq i\}$
# Then for each $i$, $
# A_i \subset \Gamma_i(W,b)$
# ```
#
# We note that each $\Gamma_i(W,b)$ is a polygon whose boundary consists
# of hyperplanes given by [\[Hij\]](#Hij){reference-type="eqref"
# reference="Hij"}.
#
# ```{figure} ../figures/3-class.png
# :name: 3-class
# :height: 150px
# Linearly separable sets in 2-d space (k =
# 3)
# ```
#
#
# We next introduce two more definitions of linearly separable sets that
# have more clear geometric interpretation.
#
# ```{admonition} Definition 3
# A collection of subsets $A_1,...,A_k\subset \mathbb{R}^d$ is all-vs-one
# linearly separable if for each $i = 1,...,k$, $A_i$ and
# $\displaystyle \cup_{j\neq i} A_j$ are linearly separable.
# ```
# ```{figure} ../figures/MulLClassfication.png
# :name: MulLClassfication
# :height: 150px
# All-vs-One linearly separable sets (k =
# 3)
# ```
#
#
# ```{admonition} Definition 4
# A collection of subsets $A_1,...,A_k\subset \mathbb{R}^d$ is pairwise
# linearly separable if for each pair of indices $1\leq i <
# j\leq k$, $A_i$ and $A_j$ are linearly separable.
# ```
# ```{figure} ../figures/pairwise_linearly_separable.png
# :name: pairwise_linearly_separable
# :height: 150px
# Pairwise linearly separable sets in 2-d space (k =
# 3)
# ```
#
#
# We begin by comparing our notion of linearly separable to the two other
# previously introduced geometric definitions of all-vs-one linearly
# separable and pairwise lineaerly separable. Obviously, in the case of
# two classes, they are all equivalent, however, with more than two
# classes this is no longer the case. We do have the following
# implications, though.
#
# ```{admonition} Lemma 3
# If $A_1,...,A_k\subset \mathbb{R}^d$ are all-vs-one linearly separable,
# then they are linearly separable as well.
# ```
#
# ```{admonition} Proof
# *Proof.* Assume that $A_1,...,A_k$ are all-vs-one linearly separable.
# For each $i$, let $w_i$, $b_i$ be such that $w_ix + b_i$ separates $A_i$
# from $\cup_{j\neq i} A_j$, i.e. $w_ix + b_i > 0$ for $x\in A_i$ and
# $w_ix + b_i < 0$ for $x \in \cup_{j\neq i} A_j$.
#
# Set $W = (w_1^T,w_2^T,\cdots,w_k^T)^T$, $b = (b_1,b_2,\cdots,b_k)^T$ and
# observe that if $x\in A_i$, then $(Wx + b)_i > 0$ while $(Wx + b)_j < 0$
# for all $j\neq i$. ◻
# ```
#
# ```{admonition} Lemma 4
# If $A_1,...,A_k\subset \mathbb{R}^n$ are linearly separable, then they
# are pairwise linearly separable as well.
# ```
#
# ```{admonition} Proof
# *Proof.* If $A_1,...,A_k\subset \mathbb{R}^d$ are linearly separable,
# suppose that $W = (w_1^T,w_2^T,\cdots,w_k^T)^T$, $b =
# (b_1,b_2,\cdots,b_k)^T$. So we have $\begin{cases}
# w_i x+ b_i > w_j x + b_j & x\in A_i \\
# w_i x+ b_i < w_j x + b_j& x\in A_j \\
# \end{cases}$ Take $w_{i,j} = w_i - w_j, b_{i,j} = b_i-b_j$, then we
# have $w_{i,j}x + b_{i,j}\begin{cases}
# > 0 & x\in A_i \\
# < 0 & x\in A_j \\
# \end{cases}$ So $A_1,...,A_k$ are pairwise linearly separable. ◻
# ```
#
# However, the converses of both of these statements are false, as the
# following examples show.
#
# ```{admonition} example
# Consider the sets $A_1, A_2, A_3\subset \mathbb{R}$ given by
# $A_1 = [-4,-2]$, $A_2 = [-1,1]$, and $A_3 = [2,4]$. These sets are
# clearly not one-vs-all linearly separable because $A_2$ cannot be
# separated from both $A_1$ and $A_3$ by a single plane (in $\mathbb{R}$
# this is just cutting the real line at a given number, and $A_2$ is in
# the middle).
#
# However, these sets are linearly separated by $W = [-2,0,2]^T$ and
# $b = [-3,0,-3]^T$, for example.
# ```
#
# ```{admonition} example
# Consider the sets $A_1, A_2, A_3\subset \mathbb{R}^2$ shown in figure
# [6](#pairwise_separable_example){reference-type="ref"
# reference="pairwise_separable_example"}. Note that $A_i$ and $A_j$ are
# separated by hyperplane $H_{i,j}$ (drawn in the figure) and so these
# sets are pairwise linearly separable. We will show that they are not
# linearly separable.
#
# Assume to the contrary that $W\in \mathbb{R}^{3\times 2}$ and
# $b\in \mathbb{R}^2$ separate $A_1$, $A_2$, and $A_3$. Then
# $(w_i - w_j)x + (b_i - b_j)$ must be a plane which separates $A_i$ and
# $A_j$. Now consider a point $z$ bounded by $A_1$, $A_2$ and $A_3$ in
# figure [6](#pairwise_separable_example){reference-type="ref"
# reference="pairwise_separable_example"}. We see from the figure that
# given any plane separating $A_1$ from $A_2$, $z$ must be on the same
# side as $A_2$, given any plane separating $A_2$ from $A_3$, $z$ must be
# on the same side as $A_3$, and given any plane separating $A_3$ from
# $A_1$, $z$ must be on the same side as $A_1$.
#
# This means that $(w_2 - w_1)z + (b_2 - b_1) > 0$,
# $(w_3 - w_2)z + (b_3 - b_2) > 0$, and $(w_1 - w_3)z + (b_1 - b_3) > 0$.
# Adding these together, we obtain $0 > 0$, a contradiction.
#
# The essence behind this example is that although the sets $A_1$, $A_2$,
# and $A_3$ are pairwise linearly separable, no possible pairwise
# separation allows us to consistently classify arbitrary new points.
# However, a linear separation would give us a consistent scheme for
# classifying new points.
# ```
#
# So the notion of linear separability is sandwiched in between the more
# intuitive notions of all-vs-one and pairwise separability. It turns out
# that linear separability is the notion which is most useful for the
# $k$-class classification problem and so we focus on this notion of
# separability from now on.
# + active=""
#
| ch02/.ipynb_checkpoints/ch2_1-checkpoint.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// A Jupyter Notebook don't have a inbuilt kernel for java. So I used IJava NoteBook to create this. If you want to run any program or want to know more about IJava NoteBook then visit [GitHub](https://github.com/SpencerPark/IJava).
// 1. What is the output of following program?
//
public class BitPuzzel {
public static void main(String[] args) {
int mask = 0x000F;
int value = 0x2222;
System.out.println(value & mask);
}
}
Answer: 2
// 2. What is the output of following program?
public class A {
int add(int i, int j) {
return i + j;
}
}
public class B extends A {
public static void main(String[] args) {
short s = 9;
System.out.println(add(s, 6));
}
}
Answer: Compilation Fails due to error at line 9, non-static method referenced from a static context
// 3. Which is the right answer to the following?
public interface Syrupable {
void getSugary();
}
abstract class Pancake implements Syrupable {}
class BuleBerryPancake implements Pancake {
public void getSugary() { ; }
}
class SourdoughBuleBerryPancake extends BuleBerryPancake {
void getSugary(int s) { ; }
}
Answer: Compilation fails due to an error on line 6
// 4. Find the Output
try {
Float f = new Float("3.0");
int x = f.intValue();
System.out.println("x : "+x);
byte b = f.byteValue();
System.out.println("b : "+b);
double d = f.doubleValue();
System.out.println("d : "+d);
System.out.println(x + b + d);
}
catch (NumberFormatException e) /* Line 9 */
{
System.out.println("bad number"); /* Line 11 */
}
Answer: 9.0
// 5. What is a covariant return type?
Answer: The overiding method can have base type as the return type instead of the derived type
| Certification/01. Java (Basic)/03. MCQs/Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
reviews = pd.read_csv('winemag-data-130k-v2.csv')
reviews.head()
# -
# ### Bar Chart
# plot countries against varities
reviews['country'].value_counts().head(10).plot.bar()
us_wines = reviews[reviews['country'] == 'US']
us_wines.head(5)
us_wines['province'].value_counts().head(10).plot.bar()
# #### to see the contribution of each province
# note: head value is given for neat display don't change it
(us_wines['province'].value_counts() / len(us_wines)).head(10).plot.bar()
# #### ordinal categories
# points and wines
reviews['points'].value_counts().sort_index().plot.bar()
# ### LIne Chart
# for these type of ordinal categories that has a large range of values Line chart can be useful
reviews['points'].value_counts().sort_index().plot.line()
# +
top_reviewers = reviews['taster_name'].value_counts().head(5)
top_reviewers
# -
# #### To see average scores given by <NAME>
# +
roger = reviews[reviews['taster_name'] == '<NAME>']
roger['points'].value_counts().sort_index().plot.bar()
# -
# ### Interval data
# Examples of interval variables are the wind speed in a hurricane, shear strength in concrete, and the temperature of the sun. An interval variable goes beyond an ordinal categorical variable: it has a meaningful order, in the sense that we can quantify what the difference between two entries is itself an interval variable.
#
# For example, if I say that this sample of water is -20 degrees Celcius, and this other sample is 120 degrees Celcius, then I can quantify the difference between them: 140 degrees "worth" of heat, or such-and-such many joules of energy.
#
# Line charts work for these type of data. Histrogram can also be applied
#
# ### Histograms
reviews[reviews['price'] < 100]['price'].value_counts().head(10).plot.hist()
# ## Bivarient Plots
#
# ### Scatter Plots
# These are simple bivarent plots that map variable of interset to two variables
reviews[reviews['price'] > 1200].plot.scatter(x="points",y="price")
reviews[reviews['price'] > 2000][['price','variety']].sort_index().head(8)
reviews[reviews['price'] < 150].sample(100).plot.scatter(x="price",y='points')
# Scatter plot can't be used for large amount of data It should be limited to small number of samples
# Below plot is the reason for it
reviews[reviews['price'] < 150].plot.scatter(x="price",y='points')
# ### Hexa Plot
# hexaplots can be used to see where the points are clustered
reviews[reviews['price'] < 100 ].plot.hexbin(x='price', y='points', gridsize=16)
# ### Stacked plots
# It is the one in which plots the variables on top of each other
top_wines = reviews['variety'].value_counts().head(5)
reviews[reviews['variety'] in top_wines]
# ### Pokemon Dataset
# It is used for exersices and to create some visulaizations.
# +
pokemon = pd.read_csv('Pokemon.csv',index_col=0)
pokemon.head()
# -
len(pokemon)
pokemon.plot.hexbin(x="Defense",y="Attack",gridsize=15)
pokemon.plot.hexbin(x="HP",y="Attack",gridsize=15)
# +
pokemon_stat_legendary = pokemon.groupby(['Legendary','Generation']).mean()[['Attack','Defense']]
pokemon_stat_legendary.head()
# -
pokemon_stat_legendary.plot.bar(stacked=True)
pokemon_stat_legendary.plot.line(figsize=(12,6))
# +
pokemon_stat_by_types = pokemon.groupby(['Generation','Type 1']).mean()[['HP','Attack','Defense','Sp. Atk','Speed']]
pokemon_stat_by_types.plot.line(figsize=(13,7))
# -
# ## Styling the Plot
#
# ### Introduction
# When we expose our plots to external audience Styling is a must.
# We will make this plot look better.
reviews['points'].value_counts().sort_index().plot.bar()
reviews['points'].value_counts().sort_index().plot.bar(figsize=(12,10))
reviews['points'].value_counts().sort_index().plot.bar(
figsize=(12,10),
color=(0.12,0.34,0.2)
)
reviews['points'].value_counts().sort_index().plot.bar(
figsize=(12,10),
color='mediumvioletred'
)
reviews['points'].value_counts().sort_index().plot.bar(
figsize=(12,6),
color=(0.12,0.34,0.2),
fontsize=12
)
reviews['points'].value_counts().sort_index().plot.bar(
figsize=(12,6),
color=(0.12,0.34,0.2),
title="Wine Ranking Distribution"
)
# +
import matplotlib.pyplot as plt
import seaborn as sb
ax = reviews['points'].value_counts().sort_index().plot.bar(
figsize=(12,5),
color=(0.12,0.34,0.2)
)
ax.set_title("Wine Ranking Distribution",fontsize=16)
# -
pokemon.head()
ax = pokemon['Type 1'].value_counts().head(12).plot.bar(
figsize=(12,6)
)
ax.set_title('Types of Pokemons and their numbers',fontsize=15)
# ## Subplots
#
# Subplotting is a method for creating plots that live side by side together.
#
# They are used to see two plots and how they match with each other.
#
# Subplots takes two arguments one for rows and secound for columns
# +
fig,axes = plt.subplots(2, 1,figsize=(10,6))
# To apply proper spacing between plots
plt.subplots_adjust(hspace=.35)
reviews['points'].value_counts().sort_index().plot.bar(
ax = axes[0],
title="Wine Price Distribution",
fontsize=12
)
reviews['variety'].value_counts().head(10).plot.bar(
ax = axes[1],
title="Wine Varieties",
fontsize=12
)
# -
fig, axes = plt.subplots(2,2,figsize=(12,6))
# +
fig, axes = plt.subplots(2,2,figsize=(12,6))
# To apply proper spacing between plots
plt.subplots_adjust(hspace=.35)
reviews['points'].value_counts().sort_index().plot.bar(
ax=axes[0][0],
color="mediumvioletred"
)
axes[0][0].set_title('Wine Ratings',fontsize=15)
reviews['price'].value_counts().sort_index().plot.hist(
ax=axes[0][1],
color="mediumvioletred"
)
axes[0][1].set_title('Wine Price',fontsize=15)
reviews['province'].value_counts().head(10).plot.bar(
ax=axes[1][0],
color="mediumvioletred"
)
axes[1][0].set_title('Wine Producing Provinces',fontsize=15)
reviews['variety'].value_counts().head(10).plot.bar(
ax=axes[1][1],
color="mediumvioletred"
)
axes[1][1].set_title('Wine Variety',fontsize=15)
# -
pokemon.head()
# +
gen1 = pokemon[pokemon['Generation'] == 1]
gen2 = pokemon[pokemon['Generation'] == 2]
gen3 = pokemon[pokemon['Generation'] == 3]
gen4 = pokemon[pokemon['Generation'] == 4]
plt.subplots_adjust(hspace=.5)
fig, axes = plt.subplots(2,4,figsize=(15,6))
gen1['HP'].value_counts().sort_index().plot.line(
ax = axes[0][0],
color="mediumvioletred",
title="Gen1"
)
gen2['HP'].value_counts().sort_index().plot.line(
ax = axes[0][1],
color="mediumvioletred",
title="Gen2"
)
gen3['HP'].value_counts().sort_index().plot.line(
ax = axes[0][2],
color="mediumvioletred",
title="Gen3"
)
gen4['HP'].value_counts().sort_index().plot.line(
ax = axes[0][3],
color="mediumvioletred",
title="Gen4"
)
# -
axes
| Machine Learning/visualization/Wine and Pokemons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# #! pip2 install opencv-python
# -
import cv2, glob, numpy, os
import pandas as pd
import matplotlib.pyplot as plt
scale=300
image_size = 540
source_dir='/media/carlos_bologna/WD_NTFS/Diabetic_Retinopathy/original_images/'
dest_dir='data/'
def scaleRadius(img,scale):
x=img[img.shape[0]/2,:,:].sum(1)
r=(x>x.mean()/10).sum() / 2
s=scale * 1.0 / r
return cv2.resize(img,(0,0),fx=s,fy=s)
# +
df_train = pd.read_csv('data/trainLabels.csv')
df_test = pd.read_csv('data/testLabels.csv') # Já temos os targets da base de teste, então vamos usar.
df_train['folder'] = 'train'
df_test['folder'] = 'test'
df = pd.concat([df_train.loc[:, ['folder', 'image', 'level']], df_test.loc[:, ['folder', 'image', 'level']]])
tot_lote = len(df) / 5000
# -
i=1
for index, row in df.iterrows():
try:
a=cv2.imread(os.path.join(source_dir, row.folder, row.image + '.jpeg'))
#scale img to a given radius
a=scaleRadius(a,scale)
#subtract local mean color
a=cv2.addWeighted(a, 4, cv2.GaussianBlur(a,(0,0),scale/30), -4, 128)
#remove outer 10%
b=numpy.zeros(a.shape)
cv2.circle(b,(a.shape[1]/2,a.shape[0]/2), int(scale * 0.9),(1,1,1), -1,8,0)
#a= a * b + 128 * (1 - b)
a = a * b
# Crop image removing black border
#r = calcRadius(a)
r = image_size / 2
h_half = a.shape[0] / 2
w_half = a.shape[1] / 2
if (r > h_half) | (r > w_half):
#Calc max padding to apply (toward h or w)
pad = max(r - h_half, r - w_half)
#Add padding
a = numpy.pad(a, ((pad,pad), (pad,pad), (0, 0)), 'constant')
h_half = a.shape[0] / 2
w_half = a.shape[1] / 2
a = a[h_half - r : h_half + r, w_half - r : w_half + r, :]
cv2.imwrite(os.path.join(dest_dir, str(row.level), row.folder + '_' + row.image + '.jpeg'), a)
if (index % 5000 == 0):
print 'Lote: ' + str(i) + '/' + str(tot_lote)
i+=1
except:
print 'Error in ' + row.image
| .ipynb_checkpoints/preprocess-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab_type="code" id="lCdA5A_4AawH" outputId="97b3c109-207d-43f8-c406-19f0a3492707" executionInfo={"status": "ok", "timestamp": 1586722256382, "user_tz": 240, "elapsed": 19403, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# + id="ho3k_anDWL09" colab_type="code" colab={}
GOOGLE_COLAB = True
# + colab_type="code" id="tBkHQA5CAYP2" colab={}
# %reload_ext autoreload
# %autoreload 2
# + colab_type="code" id="GmZLGhnTAYP7" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="30ce1d6c-cd23-42ea-ef18-2f0747bd33b2" executionInfo={"status": "ok", "timestamp": 1586722257310, "user_tz": 240, "elapsed": 911, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import random
import pickle
# + id="7ZW4u604PJOy" colab_type="code" outputId="4bf2b49e-0c5e-4f1f-e665-2a10a41c91da" executionInfo={"status": "ok", "timestamp": 1586722258857, "user_tz": 240, "elapsed": 2443, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorflow_version 2.x
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.callbacks import EarlyStopping
print(tf.__version__)
# + colab_type="code" id="vfTX4hByAYQE" colab={}
import sys
if GOOGLE_COLAB:
sys.path.append('drive/My Drive/yelp_sentiment_analysis')
else:
sys.path.append('../')
from yelpsent import data
from yelpsent import features
from yelpsent import metrics
from yelpsent import visualization
from yelpsent import models
# + id="rMagodlQ0CPW" colab_type="code" colab={}
import importlib
def reload():
importlib.reload(data)
importlib.reload(features)
importlib.reload(metrics)
importlib.reload(visualization)
importlib.reload(models)
# + [markdown] colab_type="text" id="5Eeg1CLlAYQH"
# # Load Dataset
# + colab_type="code" id="hGnUCDXLAYQI" colab={}
if GOOGLE_COLAB:
data_train, data_test = data.load_dataset("drive/My Drive/yelp_sentiment_analysis/data/yelp_train_balanced.json",
"drive/My Drive/yelp_sentiment_analysis/data/yelp_test.json")
data_train_clean, data_test_clean = data.load_dataset("drive/My Drive/yelp_sentiment_analysis/data/yelp_train_clean_nolemma.json",
"drive/My Drive/yelp_sentiment_analysis/data/yelp_test_clean_nolemma.json")
else:
data_train, data_test = data.load_dataset("../data/yelp_train.json",
"../data/yelp_test.json")
# + colab_type="code" id="qE4liC-yAYQX" colab={}
X_train = data_train['review'].tolist()
y_train = data_train['sentiment'].tolist()
X_train_clean = data_train_clean['review'].tolist()
# + colab_type="code" id="K-pw_ebWAYQZ" colab={}
X_test = data_test['review'].tolist()
y_test = data_test['sentiment'].tolist()
X_test_clean = data_test_clean['review'].tolist()
# + id="0U3d-SuCJJhl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1b019f27-7b73-49c8-9ca1-7e102b98bbdc" executionInfo={"status": "ok", "timestamp": 1586722269624, "user_tz": 240, "elapsed": 6497, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}}
from keras.utils.np_utils import to_categorical
# + id="4UZCExAUJAgj" colab_type="code" colab={}
y_train_one_hot = to_categorical(y_train, num_classes=3)
# + id="6mB4_nm1JY_d" colab_type="code" outputId="ec17b618-22ba-4941-9b12-7c60c6ba91b4" executionInfo={"status": "ok", "timestamp": 1586722269625, "user_tz": 240, "elapsed": 5801, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
y_train_one_hot.shape
# + colab_type="code" id="VGbHwAFpJh53" colab={}
y_test_one_hot = to_categorical(y_test, num_classes=3)
# + id="A295fPLiJk9K" colab_type="code" outputId="56bb8788-f3aa-419f-b176-b6bd67b0136a" executionInfo={"status": "ok", "timestamp": 1586722269808, "user_tz": 240, "elapsed": 4306, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
y_test_one_hot.shape
# + [markdown] id="sDORt1ygPcW7" colab_type="text"
# # Clean Reviews
# + id="zXCGsgP-PphU" colab_type="code" colab={}
import re
import string
import nltk
from nltk import WordNetLemmatizer, sent_tokenize, wordpunct_tokenize, pos_tag
from nltk.corpus import wordnet
# + id="nSNaOF2eP69i" colab_type="code" outputId="6cb894c4-30e7-42f3-c709-a67788e2c377" executionInfo={"status": "ok", "timestamp": 1586637366398, "user_tz": 240, "elapsed": 383, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
nltk.download('stopwords')
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
nltk.download('wordnet')
# + id="Udct0CkzPzU3" colab_type="code" colab={}
punct = set(string.punctuation)
# lemmatizer = WordNetLemmatizer()
# + id="bIUsS9YoPunS" colab_type="code" colab={}
# def lemmatize(token, tag):
# tag = {
# 'N': wordnet.NOUN,
# 'V': wordnet.VERB,
# 'R': wordnet.ADV,
# 'J': wordnet.ADJ
# }.get(tag[0], wordnet.NOUN)
# return lemmatizer.lemmatize(token, tag)
# + id="03ObyrW3wpys" colab_type="code" colab={}
# def clean_doc(doc):
# cleaned_tokens = []
# # Break the document into sentences
# for sent in sent_tokenize(doc):
# # Break the sentence into part of speech tagged tokens
# for token, tag in pos_tag(wordpunct_tokenize(sent)):
# # Lower case and strip spaces
# token = token.lower()
# token = token.strip()
# # If punctuation, continue
# if all(char in punct for char in token):
# continue
# # Lemmatize/stem the token
# token = lemmatize(token, tag)
# cleaned_tokens.append(token)
# return ' '.join(cleaned_tokens)
# + id="77HvuDww8YQ3" colab_type="code" colab={}
def clean_doc(doc):
cleaned_tokens = []
# Break the document into sentences
for sent in sent_tokenize(doc):
# Break the sentence into part of speech tagged tokens
for token in wordpunct_tokenize(sent):
# Lower case and strip spaces
token = token.lower()
token = token.strip()
# If punctuation, continue
if all(char in punct for char in token):
continue
# Add back token
cleaned_tokens.append(token)
return ' '.join(cleaned_tokens)
# + id="ev-6lFrB8iGt" colab_type="code" colab={}
X_train_clean = [clean_doc(doc) for doc in X_train]
X_test_clean = [clean_doc(doc) for doc in X_test]
# + id="aCQG_Jkk9ggz" colab_type="code" colab={}
data_train_clean = pd.DataFrame({'review':X_train_clean, 'sentiment':y_train})
data_train_test = pd.DataFrame({'review':X_test_clean, 'sentiment':y_test})
data_train_clean.to_json('drive/My Drive/yelp_sentiment_analysis/data/yelp_train_clean_nolemma.json', orient='records')
data_train_test.to_json('drive/My Drive/yelp_sentiment_analysis/data/yelp_test_clean_nolemma.json', orient='records')
# + id="kLK2DxRBQr85" colab_type="code" colab={}
# data_train_clean = pd.DataFrame({'review':X_train_clean, 'sentiment':y_train})
# data_train_test = pd.DataFrame({'review':X_test_clean, 'sentiment':y_test})
# + id="ET4rdiNHUVPV" colab_type="code" colab={}
# data_train_clean.to_json('drive/My Drive/yelp_sentiment_analysis/data/yelp_train_clean.json', orient='records')
# data_train_test.to_json('drive/My Drive/yelp_sentiment_analysis/data/yelp_test_clean.json', orient='records')
# + id="NpoApUJkArVO" colab_type="code" colab={}
import os
import urllib.request
# + id="vbO_sJzSsl6L" colab_type="code" outputId="5a0bfdce-14c3-4c1d-9e81-74715a5a2067" executionInfo={"status": "ok", "timestamp": 1586633418658, "user_tz": 240, "elapsed": 387979, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
urllib.request.urlretrieve('http://nlp.stanford.edu/data/glove.6B.zip', 'glove.6B.zip')
# + id="5E-WcP6wFrDs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 307} outputId="3f247253-1e64-45dd-baa8-23d2184bb366" executionInfo={"status": "error", "timestamp": 1586723493826, "user_tz": 240, "elapsed": 402, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}}
urllib.request.urlretrieve('https://www.floydhub.com/mhavelant/datasets/glove6b/1/glove.6B.200d.txt', 'glove.6B.200d.txt')
# + id="9xvuWLFOst25" colab_type="code" outputId="a63c45e4-3b33-48c6-abb0-16a30a551d29" executionInfo={"status": "ok", "timestamp": 1586633495424, "user_tz": 240, "elapsed": 24100, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
# !unzip glove.6B.zip
# + id="7UQPjw-tAk7x" colab_type="code" outputId="65a8c075-bbe0-4e82-ca6c-a7539147efcb" executionInfo={"status": "ok", "timestamp": 1586638982535, "user_tz": 240, "elapsed": 701992, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
urllib.request.urlretrieve('http://nlp.stanford.edu/data/glove.twitter.27B.zip', 'glove.twitter.27B.zip')
# + id="qe71c9Q1Avfh" colab_type="code" outputId="867204a1-aded-42f1-80c0-0a34ba20364a" executionInfo={"status": "ok", "timestamp": 1586639032399, "user_tz": 240, "elapsed": 44493, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
# !unzip glove.twitter.27B.zip
# + [markdown] id="XROCUaIoZcLc" colab_type="text"
# # Tokenize
# + id="uLmhn2bhYGhj" colab_type="code" colab={}
# vocab_size = 5000
# + id="G94yuRmdZNF_" colab_type="code" colab={}
tokenizer = Tokenizer()
# tokenizer = Tokenizer(num_words=vocab_size,
# oov_token=True)
# + id="6BsDh9CcbG_m" colab_type="code" outputId="7b0236f2-1ea4-438b-bd9b-bea4251dd0e7" executionInfo={"status": "ok", "timestamp": 1586724411546, "user_tz": 240, "elapsed": 12723, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# %time tokenizer.fit_on_texts(X_train_clean)
# + id="es56QVEFbDip" colab_type="code" colab={}
word_index = tokenizer.word_index
# + id="hOBSCGAiyeuu" colab_type="code" outputId="f84583e5-afff-449b-8ef2-64410dcb5950" executionInfo={"status": "ok", "timestamp": 1586724413108, "user_tz": 240, "elapsed": 358, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
len(word_index)
# + id="nEY8cCGwJpID" colab_type="code" colab={}
max_length = 200
# + id="f3anQBf1bSAr" colab_type="code" colab={}
X_train_sequences = tokenizer.texts_to_sequences(X_train_clean)
X_train_padded = pad_sequences(X_train_sequences,
maxlen=max_length,
padding='post',
truncating='post')
# + id="fXJfr332j4yS" colab_type="code" colab={}
X_test_sequences = tokenizer.texts_to_sequences(X_test_clean)
X_test_padded = pad_sequences(X_test_sequences,
maxlen=max_length,
padding='post',
truncating='post')
# + id="IwSHbyFHijPN" colab_type="code" outputId="6347ed1d-1c78-4c8f-dddc-51df3fd05e1b" executionInfo={"status": "ok", "timestamp": 1586724433164, "user_tz": 240, "elapsed": 3584, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 479}
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_article(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
print('---')
print(X_train[1])
print('---')
print(X_train_clean[1])
print('---')
print((X_train_padded[1]))
print('---')
print(decode_article(X_train_padded[1]))
# + [markdown] id="c_XOxPL97SGV" colab_type="text"
# Pre-Trained GloVe Embedding (https://nlp.stanford.edu/projects/glove/)
#
#
# + id="XmPfguc0uke-" colab_type="code" colab={}
embeddings_index = dict()
with open('glove.6B.200d.txt') as f:
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
# + id="Y_wptG0FwYoJ" colab_type="code" colab={}
embeddings_matrix = np.zeros((len(word_index), max_length))
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# print(embedding_vector.shape)
embeddings_matrix[i-1] = embedding_vector
# + [markdown] id="E6Ughqe7i6LX" colab_type="text"
# # Model
# + id="d_8Ds8eR1reV" colab_type="code" outputId="39e486f1-acbc-4d4c-d6d8-fd8c60bc0e37" executionInfo={"status": "ok", "timestamp": 1586724544780, "user_tz": 240, "elapsed": 5416, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 714}
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver() # TPU detection
print('Running on TPU ', tpu.cluster_spec().as_dict()['worker'])
except ValueError:
raise BaseException('ERROR: Not connected to a TPU runtime; please see the previous cell in this notebook for instructions!')
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
tpu_strategy = tf.distribute.experimental.TPUStrategy(tpu)
# + id="eyyKedSGh36V" colab_type="code" outputId="825c4967-a95f-4d41-fa3b-3a7a5974136c" executionInfo={"status": "ok", "timestamp": 1586724544780, "user_tz": 240, "elapsed": 2228, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
print((X_train_padded.shape, y_train_one_hot.shape, X_test_padded.shape, y_test_one_hot.shape))
# + colab_type="code" outputId="64977c62-3925-4751-937f-a9d0ff78c34b" executionInfo={"status": "ok", "timestamp": 1586727419584, "user_tz": 240, "elapsed": 2024, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} id="JCII2I_3C3Xx" colab={"base_uri": "https://localhost:8080/", "height": 357}
with tpu_strategy.scope():
model = tf.keras.Sequential([
# GloVe embedding
tf.keras.layers.Embedding(input_dim=len(word_index),
output_dim=max_length,
input_length=max_length,
weights=[embeddings_matrix],
trainable=False),
# Bidirectional LSTM
# tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(units=128,
# dropout=0.3,
# recurrent_dropout=0.3)),
tf.keras.layers.LSTM(units=64, return_sequences=True, recurrent_dropout=0.5),
tf.keras.layers.Dropout(rate=0.5),
tf.keras.layers.LSTM(units=64, recurrent_dropout=0.5),
# ReLU
tf.keras.layers.Dense(64, activation='relu'),
# SoftMax output
tf.keras.layers.Dense(3, activation='softmax')
])
model.compile(loss='categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(learning_rate=0.01,
decay=0.001),
metrics=['accuracy'])
es = EarlyStopping(monitor='val_loss',
mode='min',
verbose=1,
patience=2)
model.summary()
# + id="aq5MJP2ajNHV" colab_type="code" outputId="396f2538-8bf6-40ea-9573-362a933693a6" executionInfo={"status": "ok", "timestamp": 1586728478884, "user_tz": 240, "elapsed": 1054179, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 408}
history = model.fit(X_train_padded,
y_train_one_hot,
epochs=30,
# validation_data=(X_test_padded, y_test_one_hot),
validation_split=0.2,
verbose=2,
callbacks=[es])
# + id="6jyrV7csTL49" colab_type="code" outputId="6935fd5a-a477-4095-dc37-ab9d09b7844b" executionInfo={"status": "ok", "timestamp": 1586730595994, "user_tz": 240, "elapsed": 48675, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 85}
accr = model.evaluate(X_test_padded, y_test_one_hot)
print('Test set\n Loss: {:0.3f}\n Accuracy: {:0.3f}'.format(accr[0],accr[1]))
# + id="pdNnKEAPkJVk" colab_type="code" outputId="c21f2e35-cdc7-49e5-f186-2883d6e6fe3f" executionInfo={"status": "ok", "timestamp": 1586730941426, "user_tz": 240, "elapsed": 774, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 755}
def display_training_curves(training, validation, title, subplot):
ax = plt.subplot(subplot)
ax.plot(training)
ax.plot(validation)
ax.set_title('Model '+ title)
ax.set_ylabel(title)
ax.set_xlabel('epoch')
ax.legend(['Training', 'Validation'])
plt.subplots(figsize=(10,10))
plt.tight_layout()
display_training_curves(history.history['accuracy'], history.history['val_accuracy'], 'Accuracy', 211)
display_training_curves(history.history['loss'], history.history['val_loss'], 'Loss', 212)
# + id="TIKL-jDK1HHy" colab_type="code" colab={}
labels = [0,1,2]
# txt = [X_test_clean[0]]
# seq = tokenizer.texts_to_sequences(txt)
# padded = pad_sequences(seq, maxlen=max_length)
# pred = model.predict(padded)[:len(txt)]
# print(pred, labels[np.argmax(pred)])
# + id="9Jdb9QxnJV3c" colab_type="code" outputId="418163a8-aba6-44c1-973d-8c2c60b43497" executionInfo={"status": "ok", "timestamp": 1586730782873, "user_tz": 240, "elapsed": 49694, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# %time y_train_prob = model.predict(X_train_padded)[:len(X_train_padded)]
# + id="-AvVRC9Ezpnj" colab_type="code" outputId="fa959eb7-9d61-45f2-9811-77942b5b9420" executionInfo={"status": "ok", "timestamp": 1586730816360, "user_tz": 240, "elapsed": 82865, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# %time y_test_prob = model.predict(X_test_padded)[:len(X_test_padded)]
# + id="9fDLm7ZELccG" colab_type="code" outputId="54c06e3c-6c7a-43fa-8c85-1f0833ccff33" executionInfo={"status": "ok", "timestamp": 1586730816435, "user_tz": 240, "elapsed": 82589, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# %time y_train_pred = [labels[np.argmax(pred)] for pred in y_train_prob]
# + id="d5FvQ4NHLWYV" colab_type="code" outputId="cb98554f-5fb4-433b-b8c7-ab67edec49c3" executionInfo={"status": "ok", "timestamp": 1586730816811, "user_tz": 240, "elapsed": 82466, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# %time y_test_pred = [labels[np.argmax(pred)] for pred in y_test_prob]
# + id="E-q66FvuVDG3" colab_type="code" colab={}
import sklearn
# + id="xVfh2Qp5U_Bx" colab_type="code" outputId="ac991b01-7f17-4a62-8f87-868494be815a" executionInfo={"status": "ok", "timestamp": 1586730816922, "user_tz": 240, "elapsed": 81291, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
sklearn.metrics.f1_score(y_test, y_test_pred, average='macro')
# + id="P5p6o_CwEuWQ" colab_type="code" outputId="a8de29f4-65bc-4243-e767-c17dc9712833" executionInfo={"status": "ok", "timestamp": 1586730817767, "user_tz": 240, "elapsed": 79989, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg8PmDhUFm8THlQ0eP76DD2UF2SIKi7FuSRfOrD=s64", "userId": "11853385700800863483"}} colab={"base_uri": "https://localhost:8080/", "height": 314}
visualization.confusion_heat_map(y_test,
y_test_pred,
normalize='true',
fmt='.1%',
labels=set(y_test))
f1_train, f1_test = metrics.f1_score(y_train, y_train_pred), metrics.f1_score(y_test, y_test_pred)
print("Macro F1 Scores: \n Training: {0:.3f} \n Testing: {1:.3f}\n\n".format(f1_train, f1_test))
# + id="OM_pKOJr_GAO" colab_type="code" colab={}
from keras.models import load_model
# + id="zQ3qK4g3Riqf" colab_type="code" colab={}
model.save('drive/My Drive/yelp_sentiment_analysis/models/lstm_64units_10epochs.h5')
# model = load_model('drive/My Drive/yelp_sentiment_analysis/models/lstm.h5')
# + id="3wYc00u8OI8Y" colab_type="code" colab={}
| notebooks/model_lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
# example_solar_generation.py
## import needed packages
import pandas as pd
import pvlib
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from LESO import System
from LESO import PhotoVoltaic, PhotoVoltaicAdvanced, BifacialPhotoVoltaic
from LESO.feedinfunctions import PVlibweather
from LESO.feedinfunctions import bifacial
# +
## Set up the components and system
pv1 = PhotoVoltaic('simplePV')
pv2 = PhotoVoltaicAdvanced('advancedPV')
pv3 = PhotoVoltaicAdvanced('trackingPV-NS', tracking=True)
pv4 = PhotoVoltaicAdvanced('trackingPV2-EW', tracking=True, azimuth=-90)
pv5 = BifacialPhotoVoltaic('bifacialPV')
modelname = 'Trial to compare'
system = System(52, 5, model_name = modelname)
system.add_components([pv1, pv2, pv3, pv4, pv5])
system.fetch_input_data()
system.calculate_time_series()
for pv in system.components:
print(f'{pv.name}: {pv.state.power.sum()/pv.installed} kWh / ( year * kWp )')
# Note: bifacial not working ATM
# +
## plot yield over month
system.components[0].sum_by_month()
fig = plt.figure(figsize=(12,5))
i = 0.
r = np.arange(len(system.components[0].monthly_state.index))
barwidth = .2
colors = sns.color_palette('Set3', len(system.components))
for i, component in enumerate(system.components):
component.sum_by_month()
plt.bar(
r,
list(component.monthly_state.values.flatten()),
label = component.name,
width=barwidth,
alpha = 1,
color = colors[i]
)
plt.legend()
r = [x + barwidth for x in r]
plt.xticks([x - 3*barwidth for x in r], list(system.components[0].monthly_state.index), rotation = -90)
plt.tick_params(left=False, labelleft=False) #remove ticks
plt.box(False) #remove box
| examples/example_solar_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#==========Imports==========
import numpy as np
import matplotlib.pyplot as plt
import astropy.constants as const
import time
from scipy import interpolate
import Zach_OPTIMIZER.EBMFunctions as opt
import Bell_EBM as ebm
# -
#==========Set Up System==========
planet = ebm.Planet(rad=1.900*const.R_jup.value, mass=1.470*const.M_jup.value,
Porb=1.09142030, a=0.02340*const.au.value, inc=83.37, vWind=5e3, nlat = 8)
star = ebm.Star(teff=6300., rad=1.59, mass=1.20)
system = ebm.System(star, planet)
# +
#==========Set Up System==========
planet = ebm.Planet(rad=1.900*const.R_jup.value, mass=1.470*const.M_jup.value,
Porb=1.09142030, a=0.02340*const.au.value, inc=83.37, vWind=5e3, nlat=8)
star = ebm.Star(teff=6300., rad=1.59, mass=1.20)
system = ebm.System(star, planet)
#==========Baseline Creation==========
tUpdates = 500. #Amount of timesteps in baseline
tOrbits = 1 #Number of orbits in baseline
Teq = system.get_teq()
T0 = np.ones_like(system.planet.map.values)*Teq
t0 = 0.
t1 = t0+system.planet.Porb*tOrbits
dt = system.planet.Porb/tUpdates
testTimes, testMaps, testttc = system.run_model_tester(T0, t0, t1, dt, verbose=False)
testLightcurve = system.lightcurve()
# -
times = (t0 + np.arange(int(np.rint((t1-t0)/dt)))*dt)[:,np.newaxis]
TAs = system.planet.orbit.true_anomaly(times)[:,:,np.newaxis]
maps = T0[np.newaxis,:]
Teq = system.get_teq()
T0 = np.ones_like(system.planet.map.values)*Teq
t0 = 0.
t1 = t0+system.planet.Porb*1
dt = system.planet.Porb/1000
# %timeit system.run_model(T0, t0, t1, dt, verbose=False)
# %timeit system.ODE_EQ(times[1], maps[-1], dt, TAs[1])
# %timeit system.Fin(times[1], TAs[1])
# %timeit system.planet.Fout(maps)
# %timeit system.Firr(times[1], TAs[1], True, None, 4.5e-6)
# %timeit system.planet.weight(times[1], TAs[1])
# %timeit system.planet.orbit.get_ssp(times[1], TAs[1])
refLon, refLat = system.planet.orbit.get_ssp(times[1], TAs[1])
# %timeit (np.cos(system.planet.map.latGrid_radians)*np.cos(refLat*np.pi/180.)*np.cos((system.planet.map.lonGrid_radians-refLon*np.pi/180.))+ np.sin(system.planet.map.latGrid_radians)*np.sin(refLat*np.pi/180.))
# %timeit system.planet.map.latGrid_radians
# %timeit np.cos(273672179)
# %timeit np.pi
weight = (np.cos(system.planet.map.latGrid_radians)*np.cos(refLat*np.pi/180.)*np.cos((system.planet.map.lonGrid_radians-refLon*np.pi/180.))+ np.sin(system.planet.map.latGrid_radians)*np.sin(refLat*np.pi/180.))
# %timeit np.max(np.append(np.zeros_like(weight[np.newaxis,:]), weight[np.newaxis,:], axis=0), axis=0)
# %timeit system.star.Fstar(True, None, 4.5e-6)
temp, space = opt.Optimize(star, planet, 20., verbose=True)
space
| RunModelTiming_Tester.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAT281 - Laboratorio N°01
# <a id='p1'></a>
#
# ## Problema 01
#
# ### a) Calcular el número $\pi$
#
# En los siglos XVII y XVIII, <NAME> y <NAME> descubrieron una serie infinita que sirve para calcular $\pi$:
#
# $$\displaystyle \pi = 4 \sum_{k=1}^{\infty}\dfrac{(-1)^{k+1}}{2k-1} = 4(1-\dfrac{1}{3}+\dfrac{1}{5}-\dfrac{1}{7} + ...) $$
#
# Desarolle un programa para estimar el valor de $\pi$ ocupando el método de Leibniz, donde la entrada del programa debe ser un número entero $n$ que indique cuántos términos de la suma se utilizará.
#
#
# * **Ejemplo**: *calcular_pi(3)* = 3.466666666666667, *calcular_pi(1000)* = 3.140592653839794
#
# ### Definir Función
#
def calcular_pi(n:int)->float:
"""
calcular_pi(n)
Aproximacion del valor de pi mediante el método de Leibniz
Parameters
----------
n : int
Numero de terminos.
Returns
-------
output : float
Valor aproximado de pi.
Examples
--------
>>> calcular_pi(3)
3.466666666666667
>>> calcular_pi(1000)
3.140592653839794
"""
pi = 0 # valor incial
for k in range(1,n+1):
numerador = (-1)**(k+1) # numerador de la iteracion i
denominador = 2*k-1 # denominador de la iteracion i
pi+=numerador/denominador # suma hasta el i-esimo termino
return 4*pi
# Acceso a la documentación
help(calcular_pi)
# ### Verificar ejemplos
# ejemplo 01
assert calcular_pi(3) == 3.466666666666667, "ejemplo 01 incorrecto"
# ejemplo 02
assert calcular_pi(1000) == 3.140592653839794, "ejemplo 02 incorrecto"
# **Observación**:
#
# * Note que si corre la línea de comando `calcular_pi(3.0)` le mandará un error ... ¿ por qué ?
# * En los laboratorio, no se pide ser tan meticuloso con la documentacion.
# * Lo primero es definir el código, correr los ejemplos y luego documentar correctamente.
# ### b) Calcular el número $e$
#
# Euler realizó varios aportes en relación a $e$, pero no fue hasta 1748 cuando publicó su **Introductio in analysin infinitorum** que dio un tratamiento definitivo a las ideas sobre $e$. Allí mostró que:
#
#
# En los siglos XVII y XVIII, <NAME> y <NAME> descubrieron una serie infinita que sirve para calcular π:
#
# $$\displaystyle e = \sum_{k=0}^{\infty}\dfrac{1}{k!} = 1+\dfrac{1}{2!}+\dfrac{1}{3!}+\dfrac{1}{4!} + ... $$
#
# Desarolle un programa para estimar el valor de $e$ ocupando el método de Euler, donde la entrada del programa debe ser un número entero $n$ que indique cuántos términos de la suma se utilizará.
#
#
# * **Ejemplo**: *calcular_e(3)* =2.5, *calcular_e(1000)* = 2.7182818284590455
# ### Definir función
def calcular_e(n:int)->float:
"""
calcular_e(n)
Aproximacion del valor de e mediante el método de Euler
Parameters
----------
n : int
Numero de terminos.
Returns
-------
output : float
Valor aproximado de e.
Examples
--------
>>> calcular_e(3)
2.5
>>> calcular_e(1000)
2.7182818284590455
"""
if n==0:
return 0
else:
e = 1 # valor incial
factorial = 1
for k in range(1,n):
factorial *= k #computar el k-esimo factorial
e += 1/factorial # suma hasta el k-esimo termino
return e
# ### Verificar ejemplos
# ejemplo 01
assert calcular_e(3) == 2.5, "ejemplo 01 incorrecto"
# ejemplo 02
assert calcular_e(1000) == 2.7182818284590455, "ejemplo 02 incorrecto"
# <a id='p2'></a>
#
# ## Problema 02
#
#
# Sea $\sigma(n)$ definido como la suma de los divisores propios de $n$ (números menores que n que se dividen en $n$).
#
# Los [números amigos](https://en.wikipedia.org/wiki/Amicable_numbers) son enteros positivos $n_1$ y $n_2$ tales que la suma de los divisores propios de uno es igual al otro número y viceversa, es decir, $\sigma(n_1)=n_2$ y $\sigma(n_2)=n_1$.
#
#
# Por ejemplo, los números 220 y 284 son números amigos.
# * los divisores propios de 220 son 1, 2, 4, 5, 10, 11, 20, 22, 44, 55 y 110; por lo tanto $\sigma(220) = 284$.
# * los divisores propios de 284 son 1, 2, 4, 71 y 142; entonces $\sigma(284) = 220$.
#
#
# Implemente una función llamada `amigos` cuyo input sean dos números naturales $n_1$ y $n_2$, cuyo output sea verifique si los números son amigos o no.
#
# * **Ejemplo**: *amigos(220,284)* = True, *amigos(6,5)* = False
#
# ### Definir Función
def amigos(n1:int,n2:int)->bool:
"""
amigos(n1,n2)
Determina si n1 y n2 son numeros amigos, es decir si la suma de los
divisores de n1 es igual a n2 y viceversa.
Parameters
----------
n1,n2 : int
Numeros enteros positivos.
Returns
-------
output : bool
Valor lógico de la proposición "n1 y n2 son números amigos".
Examples
--------
>>> amigos(220,284)
True
>>> amigos(6,5)
False
"""
suma_n1 = 0 #sumas iniciales
suma_n2 = 0
for k in range(1,n1):#se itera para todos los k menor a n1
if n1%k == 0: #al encontrar un divisor se suma a la suma actual
suma_n1+=k
#se hace lo mismo para n2
for l in range (1,n2):
if n2%l== 0:
suma_n2+=l
#print(suma_n1, suma_n2) #opcional para visualizar la suma obtenida
return (suma_n1==n2 and suma_n2==n1)
# ### Verificar ejemplos
# ejemplo 01
assert amigos(220,284) == True, "ejemplo 01 incorrecto"
# ejemplo 02
assert amigos(6,5) == False, "ejemplo 02 incorrecto"
# <a id='p3'></a>
#
# ## Problema 03
#
# La [conjetura de Collatz](https://en.wikipedia.org/wiki/Collatz_conjecture), conocida también como conjetura $3n+1$ o conjetura de Ulam (entre otros nombres), fue enunciada por el matemático <NAME> en 1937, y a la fecha no se ha resuelto.
#
# Sea la siguiente operación, aplicable a cualquier número entero positivo:
# * Si el número es par, se divide entre 2.
# * Si el número es impar, se multiplica por 3 y se suma 1.
#
# La conjetura dice que siempre alcanzaremos el 1 (y por tanto el ciclo 4, 2, 1) para cualquier número con el que comencemos.
#
# Implemente una función llamada `collatz` cuyo input sea un número natural positivo $N$ y como output devulva la secuencia de números hasta llegar a 1.
#
# * **Ejemplo**: *collatz(9)* = [9, 28, 14, 7, 22, 11, 34, 17, 52, 26, 13, 40, 20, 10, 5, 16, 8, 4, 2, 1]
# ### Definir Función
def collatz(n:int)->list:
"""
collatz(n)
Encuentra la secuencia de Collatz de n.
Parameters
----------
n : int
Numero entero positivo.
Returns
-------
output : list[int]
Lista con los valores de la secuencia de números obtenidos al
realizar las operaciones de Collatz hasta llegar a 1.
Examples
--------
>>> collatz(9)
[9, 28, 14, 7, 22, 11, 34, 17, 52, 26, 13, 40, 20, 10, 5, 16, 8, 4, 2, 1]
>>> collatz(8)
[8, 4, 2, 1]
"""
Lista = [n] #lista inicial contienendo solo a la entrada
while n != 1: #condicion base
if n%2==0: #se detecta numero par
n//=2 #se usa cociente para que n no pase de int a float
else: #se detecta numero impar
n=3*n+1
Lista.append(n) #añadir el resultado a la lista
return Lista
# ### Verificar ejemplos
# ejemplo 01
assert collatz(9) == [9, 28, 14, 7, 22, 11, 34, 17, 52, 26, 13, 40, 20, 10, 5, 16, 8, 4, 2, 1], "ejemplo 01 incorrecto"
# <a id='p4'></a>
#
# ## Problema 04
#
# La [conjetura de Goldbach](https://en.wikipedia.org/wiki/Goldbach%27s_conjecture) es uno de los problemas abiertos más antiguos en matemáticas. Concretamente, <NAME>, en 1921, en su famoso discurso pronunciado en la Sociedad Matemática de Copenhague, comentó que probablemente la conjetura de Goldbach no es solo uno de los problemas no resueltos más difíciles de la teoría de números, sino de todas las matemáticas. Su enunciado es el siguiente:
#
# $$\textrm{Todo número par mayor que 2 puede escribirse como suma de dos números primos - <NAME> (1742)}$$
#
# Implemente una función llamada `goldbach` cuyo input sea un número natural positivo $N$ y como output devuelva la suma de dos primos ($N1$ y $N2$) tal que: $N1+N2=N$.
#
# * **Ejemplo**: goldbash(4) = (2,2), goldbash(6) = (3,3) , goldbash(8) = (3,5)
# ### Definir función
def esprimo(n:int)->bool:
"""
esprimo(n)
Determina si un numero entero positivo es primo.
Parameters
----------
n : int
Numero entero positivo.
Returns
-------
output : bool
Valor lógico de la proposición "n es primo".
Examples
--------
>>> esprimo(123)
False
>>> esprimo(83)
True
"""
primo=True #se asume inicialmente n primo
for i in range(2,n):#se busca un divisor de n
if n%i==0: #al econtrarlo se da el valor false al bool
primo=False
return primo
def goldbash(n:int)->tuple:
"""
goldbash(n)
Determina el par de numeros primos tal que su suma es igual a n
Parameters
----------
n : int
Numero entero positivo par mayor a 2.
Returns
-------
output : tuple
Tupla de numeros primos tales que su suma equivale a n
Examples
--------
>>> goldbash(4)
(2,2)
>>> goldbash(8)
(3,5)
"""
if esprimo(n//2): #caso base: n div 2 es primo
particion=(n//2,n//2)
else: #de lo contrario se asume que se cumple la conjetura de Goldbach
primer = n//2-1 #se busca la tupla desde los impares mas cerca de n div 2
while not (esprimo(primer) and esprimo(n-primer)): #DeMorgan
primer-=2 #se itera hacia el siguiente par de impares
particion =(primer,n-primer)
return particion
# ### Verificar ejemplos
# ejemplo 01
assert goldbash(4) == (2,2), "ejemplo 01 incorrecto"
# ejemplo 02
assert goldbash(6) == (3,3), "ejemplo 02 incorrecto"
# ejemplo 03
assert goldbash(8) == (3,5), "ejemplo 03 incorrecto"
| labs/lab_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import hail as hl
hl.init(app_name="make_finalised_mt")
# Some variables
path_to_mt = "file:///home/eriurn/Hail-joint-caller/data/reblock_key_alleles.mt"
path_to_vqsr_ht = "file:///directflow/ClinicalGenomicsPipeline/dev/2021-02-04-PIPELINE-1885-All-Hail/EricData/VQSR.filtered.final.ht"
out_mt_path = "file:////directflow/ClinicalGenomicsPipeline/dev/2021-02-04-PIPELINE-1885-All-Hail/EricData/vqsr_finalised.mt"
def annotate_vqsr(mt, vqsr_ht):
mt = mt.annotate_rows(**vqsr_ht[mt.row_key])
mt = mt.annotate_rows(info=vqsr_ht[mt.row_key].info)
mt = mt.annotate_rows(
filters=mt.filters.union(vqsr_ht[mt.row_key].filters),
)
mt = mt.annotate_globals(**vqsr_ht.index_globals())
return mt
mt = hl.read_matrix_table(path_to_mt)
vqsr_ht = hl.read_table(path_to_vqsr_ht)
mt = annotate_vqsr(mt, vqsr_ht)
for ht in [vqsr_ht]:
mt = mt.annotate_globals(**ht.index_globals())
mt.write(out_mt_path, overwrite=True)
| notebooks/make_finalised_mt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Animal():
def __init__(self):
print("Animal Created")
def who_am_i(self):
print("I am an Animal")
def eat(self):
print("I am Eating")
myanimal = Animal()
myanimal.eat()
myanimal.who_am_i()
class Dog(Animal):
def __init__(self):
Animal.__init__(self)
print("Dog Created")
def who_am_i(self):
print("I am a DOG")
def eat(self):
print("I am a Dog and eating")
def bark(self):
print("Woof")
mydog = Dog()
mydog.eat()
mydog.who_am_i()
mydog.bark()
# # Polymorphism
# +
class Dog():
def __init__(self, name):
self.name = name
def speak(self):
return self.name + " Says, Woof !!"
# +
class Cat():
def __init__(self, name):
self.name = name
def speak(self):
return self.name + " Says, Meow !!"
# -
niko = Dog("niko")
felix = Cat("Felix")
print(niko.speak())
print(felix.speak())
for pet_class in [niko, felix]:
print(type(pet_class))
print(type(pet_class.speak()))
def pet_speak(pet):
print(pet.speak())
pet_speak(niko)
pet_speak(felix)
class Animal():
def __init__(self, name):
self.name = name
def speak(self):
raise NotImplementedError("Subject Class muust implement this abstract method")
myanimal = Animal('Fred')
myanimal.speak()
class Dog(Animal):
def speak(self):
return self.name + "Says Woff"
class Cat(Animal):
def speak(self):
return self.name + "Says Meow"
fido = Dog('Fido')
isile = Cat('Isile')
print(fido.speak())
print(isile.speak())
| Jose Portilla/Python Bootcamp Go from Zero to Hero in Python 3/Python Object Oriented Programing/Inheritence & Polymorphism.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# 1. Iris 분류 문제를 Voting을 사용하여 풀어라. K=5인 교차 검증을 하였을 때 평균 성능을 구하라.
# +
from sklearn.datasets import load_iris
from sklearn.metrics import classification_report
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
iris = load_iris()
X = iris.data
y = iris.target
sumXy=pd.concat([pd.DataFrame(X),pd.DataFrame(y)],1)
# +
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.ensemble import VotingClassifier
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.datasets import load_iris
iris = load_iris()
X = iris.data
y = iris.target
model1 = LogisticRegression(random_state=1)
model2 = QuadraticDiscriminantAnalysis()
model3 = GaussianNB()
ensemble = VotingClassifier(estimators=[('lr', model1), ('qda', model2), ('gnb', model3)], voting='soft')
kfold = KFold(n_splits=5, shuffle=True, random_state=0)
ensemble.fit(X,y)
results = cross_val_score(ensemble, X, y, cv=kfold).mean()
results
# -
| machine_learning/07_model_combine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 💃 Classifying differential equations 💃
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# Questions:
# - What is a <mark>differential equation</mark>?
# - What is the difference between an ordinary (<mark>ODE</mark>) and partial (<mark>PDE</mark>) differential equation?
# - How do I classify the different types of differential equations?
# + [markdown] slideshow={"slide_type": "subslide"}
# Objectives:
# - Identify the dependent and independent variables in a differential equation
# - Distinguish between and ODE and PDE
# - Identify the <mark>order</mark> of a differential equation
# - Distinguish between <mark>linear</mark> and <mark>non-linear</mark> equations
# - Distinguish between <mark>heterogeneous</mark> and <mark>homogeneous</mark> equations
# - Identify a <mark>separable</mark> equation
# + [markdown] slideshow={"slide_type": "slide"}
# ### A differential equation is an equation that relates one or more functions and their derivatives
#
# - The functions usually represent physical quantities (e.g. $\mathbf{F}$))
# - The derivative represents a rate of change (e.g. acceleration)
# - The differential equation represents the relationship between the two.
# - For example, Newton's second law for $n$ particles of mass $m$:
#
# \begin{equation}
# \mathbf{F}(t,\mathbf{x},\mathbf{v}) = m\frac{d\mathbf{v}}{dt}
# \end{equation}
# + [markdown] slideshow={"slide_type": "slide"}
# ### An independent variable is... a quantity that varies independently...
#
# - An <mark>independent variable</mark> does not depend on other variables
# - A <mark> dependent variable</mark> depends on the independent variable
#
# \begin{equation}
# \mathbf{F}(t,\mathbf{x},\mathbf{v}) = m\frac{d\mathbf{v}}{dt}
# \end{equation}
#
# - $t$ is an <mark>independent</mark> variable
# - $x$ and $v$ are <mark>dependent</mark> variables
# - Writing $x = x(t)$ makes this relationship clear.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Differential equations can be classified in a variety of ways
#
# There are several ways to describe and classify differential equations. There are standard solution methods for each type, so it is useful to understand the classifications.
#
# 
#
# <small>Once you can cook a single piece of spaghetti, you can cook all pieces of spaghetti!</small>
# + [markdown] slideshow={"slide_type": "slide"}
# ### An ODE contains differentials with respect to only one variable
#
# For example, the following equations are ODEs:
#
# \begin{equation}
# \frac{d x}{d t} = at
# \end{equation}
# \begin{equation}
# \frac{d^3 x}{d t^3} + \frac{x}{t} = b
# \end{equation}
#
# As in each case the differentials are with respect to the single variable $t$.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Partial differential equations (PDE) contain differentials with respect to several independent variables.
#
# An example of a PDE is:
#
# \begin{equation}
# \frac{\partial x}{\partial t} = \frac{\partial x}{\partial y}
# \end{equation}
#
# As there is one differential with respect to $t$ and one differential with respect to $y$.
#
# Note also the difference in notation - <mark>for ODEs we use $d$ whilst for PDEs we use $\partial$</mark>.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### The order of a differential equation is the highest order of any differential contained in it.
#
# For example:
#
# $\frac{d x}{d t} = at$ is <mark>first order</mark>.
#
# $\frac{d^3 x}{d t^3} + \frac{x}{t} = b$ is <mark>third order</mark>.
#
# **Note:** $\frac{d^3 x}{d t^3}$ does not equal $\left(\frac{d x}{d t}\right)^3$!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Linear equations do not contain higher powers of either the dependent variable or its differentials
#
# For example:
#
# $\frac{d^3 x}{d t^3} = at$ and $\frac{\partial x}{\partial t} = \frac{\partial x}{\partial y} $ are <mark>linear</mark>.
#
# $(\frac{d x}{d t})^3 = at$ and $\frac{d^3 x}{d t^3} = x^2$ are <mark>non-linear</mark>.
#
# Non-linear equations can be particularly nasty to solve analytically, and so are often tackled numerically.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# ### Homogeneous equations do not contain any non-differential terms
#
# For example:
#
# $\frac{\partial x}{\partial t} = \frac{\partial x}{\partial y}$ is a <mark>homogeneous equation</mark>.
#
# $\frac{\partial x}{\partial t} - \frac{\partial x}{\partial y}=a$ is a <mark>heterogeneous equation</mark> (unless $a=0$!).
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# ### Separable equations can be written as a product of two functions of different variables
#
# A separable differential equation takes the form
#
# \begin{equation}
# f(x)\frac{d x}{d t} = g(t)
# \end{equation}
#
# Separable equations are some of the easiest to solve as we can split the equation into two independent parts with fewer variables, and solve each in turn - we will see an example of this in the next tutorial.
# + [markdown] slideshow={"slide_type": "slide"}
#
# Keypoints:
#
# - An independent variable is a quantity that varies independently
# - Differential equations can be classified in a variety of ways
# - An ODE contains differentials with respect to only one variable
# - The order is the highest order of any differential contained in it
# - Linear equations do not contain higher powers of either the dependent variable or its differentials
# - Homogeneous equations do not contain any non-differential terms
# + [markdown] slideshow={"slide_type": "slide"}
# # Radioactive Decay
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ----
# Questions:
# - How can I describe radioactive decay using a first-order ODE?
# - What are <mark>initial conditions</mark> and why are they important?
# ----
# + [markdown] slideshow={"slide_type": "subslide"}
# ----
# Objectives:
# - Map between physical notation for a particular problem and the more general notation for all differential equations
# - Solve a linear, first order, separable ODE using integration
# - Understand the physical importance of initial conditions
# ----
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Radioactive decay can be modelled a linear, first-order ODE
#
# As our first example of an ODE we will model radioactive decay using a differential equation.
#
# We know that the decay rate is proportional to the number of atoms present. Mathematically, this relationship can be expressed as:
#
# \begin{equation}
# \frac{d N}{d t} = -\lambda N
# \end{equation}
#
# Note that we could choose to use different variables, for example:
#
# \begin{equation}
# \frac{d y}{d x} = cy
# \end{equation}
#
# However we try to use variables connected to the context of the problem. For example $N$ for the Number of atoms.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# If we know that 10% of atoms will decay per second we could write:
#
# \begin{equation}
# \frac{d N}{d t} = -0.1 N
# \end{equation}
#
# where $N$ is the number of atoms and $t$ is time measured in seconds.
#
# This equation is linear and first-order.
#
# | physical notation | generic notation $\left(\frac{dy}{dx} = f(y)\right)$ |
# |-----|-----|
# |number of atoms $N$ | dependent variable $y$|
# | time $t$ | independent variable $x$|
# | decay rate $\frac{dN}{dt}$ | differential $\frac{dy}{dx}$|
# | constant of proportionality $\lambda=0.1 $ | parameter(s) |
# + [markdown] slideshow={"slide_type": "slide"}
# ### The equation for radioactive decay is separable and has an analytic solution
#
# The radioactive decay equation is <mark>separable</mark>:
#
# \begin{equation}
# \frac{1}{N}\frac{d N}{d t} = -\lambda
# \end{equation}
#
# \begin{equation}
# \frac{dN}{N} = -\lambda dt.
# \end{equation}
#
# We can then integrate each side:
#
# \begin{equation}
# \ln N = -\lambda t + const.
# \end{equation}
#
# and solve for N:
#
# \begin{equation}
# N = e^{-\lambda t}e^{\textrm{const.}}
# \end{equation}
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### To model a physical system an initial value has to be provided
#
# \begin{equation}
# N(t) = e^{-\lambda t}e^{\textrm{const.}}
# \end{equation}
#
# At the beginning (when $t=0$):
#
# \begin{equation}
# N (t_0) = e^{0}e^{\textrm{const.}} = e^{\textrm{const.}}
# \end{equation}
#
# So we can identify $e^{\textrm{const.}}$ as the amount of radioactive material that was present in the beginning. We denote this starting amount as $N_0$.
# + [markdown] slideshow={"slide_type": "subslide"}
# Substituting this back into Equation 4, the final solution can be more meaningfully written as:
#
# \begin{equation}
# N = N_0 e^{-\lambda t}
# \end{equation}
#
# We now have not just one solution, but a whole class of solutions that are dependent on the initial amount of radioactive material $N_0$.
#
# Remember that not all mathematical solutions make physical sense. <mark>To model a physical system, this initial value (also known as initial condition) has to be provided alongside the constant of proportionality $\lambda$.</mark>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### ODEs can have initial values or boundary values
#
# ODEs have either initial values or boundary values. Take Newton's second law as an example:
#
# \begin{equation}
# \frac{\mathrm{d}^2x}{\mathrm{d}t^2} = -g
# \end{equation}
#
# An initial value problem would be where we know the starting position and velocity. A boundary value problem would be where we specify the position of the ball at times $t=t_0$ and $t=t_1$.
#
# For second-order ODEs (such as acceleration under gravity) we need to provide two initial/boundary conditions, for third-order ODEs we would need to provide three, and so on. Our radioactive decay example was a first-order ODE and so we only had to provide a single initial condition.
#
# + [markdown] slideshow={"slide_type": "slide"}
# -----
# Keypoints:
# - Radioactive decay can be modelled a linear, first-order ODE
# - The equation for radioactive decay is separable and has an analytic solution
# - To model a physical system an initial value has to be provided
# - The number of initial conditions depends on the order of the differential equation
# -----
# + [markdown] slideshow={"slide_type": "slide"}
# # Euler's Method
# + [markdown] slideshow={"slide_type": "subslide"}
# -------
# Questions:
# - How do I use Euler's method to solve a first-order ODE?
#
#
#
# --------
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ---------
# Objectives:
# - Use Euler's method, implemented in Python, to solve a first-order ODE
# - Understand that this method is approximate and the significance of step size $h$
# - Compare results at different levels of approximation using the `matplotlib` library.
# ------
# + [markdown] slideshow={"slide_type": "slide"}
# ### There are a variety of ways to solve an ODE
#
# In the previous lesson we considered nuclear decay:
#
# \begin{equation}
# \frac{\mathrm{d} N}{\mathrm{d} t} = -\lambda N
# \end{equation}
#
# This is one of the simplest examples of am ODE - a first-order, linear, separable differential equation with one dependent variable. We saw that we could model the number of atoms $N$ by finding an analytic solution through integration:
#
# \begin{equation}
# N = N_0 e^{-\lambda t}
# \end{equation}
#
# However there is more than one way to crack an egg (or solve a differential equation). We could have, instead, used an approximate, numerical method. One such method - Euler's method - is this subject of this lesson.
# + [markdown] slideshow={"slide_type": "slide"}
# ### A function can be approximated using a Taylor expansion
#
# The Taylor series is a polynomial expansion of a function about a point.
# For example, the image below shows $\mathrm{sin}(x)$ and its Taylor approximation by polynomials of degree 1, 3, 5, 7, 9, 11, and 13 at $x = 0$.
# <figure>
# <img src="../images/Sintay_SVG.svg" width=500 />
# </figure>
# Credit: Image By IkamusumeFan - CC BY-SA 3.0, Commons Wikimedia
# + [markdown] slideshow={"slide_type": "subslide"}
#
# The Taylor series of $f(x)$ evaluated at point $a$ can be expressed as:
#
# \begin{equation}
# f(x) = f(a) + \frac{\mathrm{d} f}{\mathrm{d} x}(x-a) + \frac{1}{2!} \frac{\mathrm{d} ^2f}{\mathrm{d} x^2}(x-a)^2 + \frac{1}{3!} \frac{\mathrm{d} ^3f}{\mathrm{d} x^3}(x-a)^3
# \end{equation}
#
# Returning to our example of nuclear decay, we can use a <mark>Taylor expansion</mark> to write the value of $N$ a short interval $h$ later:
#
# \begin{equation}
# N(t+h) = N(t) + h\frac{\mathrm{d}N}{\mathrm{d}t} + \frac{1}{2}h^2\frac{\mathrm{d}^2N}{\mathrm{d}t^2} + \ldots
# \end{equation}
#
# \begin{equation}
# N(t+h) = N(t) + hf(N,t) + \mathcal{O}(h^2)
# \end{equation}
#
# If $h$ is small and $h^2$ is very small we can neglect the terms in $h^2$ and higher and we get:
#
# \begin{equation}
# N(t+h) = N(t) + hf(N,t).
# \end{equation}
# + slideshow={"slide_type": "skip"}
# https://gist.github.com/dm-wyncode/55823165c104717ca49863fc526d1354
"""Embed a YouTube video via its embed url into a notebook."""
from functools import partial
from IPython.display import display, IFrame
width, height = (560, 315, )
def _iframe_attrs(embed_url):
"""Get IFrame args."""
return (
('src', 'width', 'height'),
(embed_url, width, height, ),
)
def _get_args(embed_url):
"""Get args for type to create a class."""
iframe = dict(zip(*_iframe_attrs(embed_url)))
attrs = {
'display': partial(display, IFrame(**iframe)),
}
return ('YouTubeVideo', (object, ), attrs, )
def youtube_video(embed_url):
"""Embed YouTube video into a notebook.
Place this module into the same directory as the notebook.
>>> from embed import youtube_video
>>> youtube_video(url).display()
"""
YouTubeVideo = type(*_get_args(embed_url)) # make a class
return YouTubeVideo() # return an object
# + slideshow={"slide_type": "slide"}
youtube_video("https://www.youtube.com/embed/3d6DsjIBzJ4").display()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Euler's method can be used to approximate the solution of differential equations
#
# We can keep applying the equation above so that we calculate $N(t)$ at a succession of equally spaced points for as long as we want. If $h$ is small enough we can get a good approximation to the solution of the equation. This method for solving differential equations is called Euler's method, after <NAME>, its inventor.
#
# 
#
# **Note**: Although we are neglecting terms $h^2$ and higher, Euler's method typically has an error linear in $h$ as the error accumulates over repeated steps. This means that if we want to double the accuracy of our calculation we need to double the number of steps, and double the calcuation time.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Euler's method can be applied using the Python skills we have developed
#
# Let's use Euler's method to solve the differential equation for nuclear decay. We will model the decay process over a period of 10 seconds, with the decay constant $\lambda=0.1$ and the initial condition $N_0 = 1000$.
#
# \begin{equation}
# \frac{\mathrm{d}N}{\mathrm{d} t} = -0.1 N
# \end{equation}
#
# + [markdown] slideshow={"slide_type": "slide"}
# ----
#
# Keypoints:
#
# - There are a variety of ways to solve an ODE
# - A function can be approximated using a Taylor expansion
# - If the step size $h$ is small then higher order terms can be neglected
# - Euler's method can be used to approximate the solution of differential equations
# - Euler's method can be applied using the Python skills we have developed
# - We can easily visualise our results, and compare against the analytical solution, using the matplotlib plotting library
# -----
# + [markdown] slideshow={"slide_type": "slide"}
# # The Strange Attractor
# + [markdown] slideshow={"slide_type": "subslide"}
# Questions:
# - How do I solve simultaneous ODEs?
# - How do I solve second-order ODEs (and higher)?
#
# + [markdown] slideshow={"slide_type": "subslide"}
# Objectives:
# - Use Euler's method, implemented in Python, to solve a set of simultaneous ODEs
# - Use Euler's method, implemented in Python, to solve a second-order ODE
# - Understand how the same method could be applied to higher order ODEs
# + [markdown] slideshow={"slide_type": "slide"}
# ### Computers don't care so much about the type of differential equation
#
# - In the previous lesson we used Euler's method to model radioactive decay. But we have seen that this equation can also be solved analytically
#
# - However there are a large number of physical equations that cannot be solved analytically, and that rely on numerical methods for their modelling.
#
# - For example, the Lotka-Volterra equations for studying predator-prey interactions have multiple dependent variables and the Cahn-Hilliard equation for modelling phase separation in fluids is non-linear. These equations can be solved analytically for particular, special cases only.
#
# - Computers don't care whether an equation is linear or non-linear, multi-variable or single-variable, the numerical method for studying it is much the same
#
# - The caveat is that numerical methods are approximate and so we need to think about the accuracy of the methods used.
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# ### Simultaneous ODEs can be solved using numerical methods
#
# One class of problems that are difficult to solve analytically are <bold>simultaenous ODEs</bold>. These are equations where the derivative of each dependent variable can depend on any of the variables (dependent or independent). For example,
#
# \begin{eqnarray}
# \frac{\mathrm{d}x}{\mathrm{d}t} &=& xt + y \\
# \frac{\mathrm{d}y}{\mathrm{d}t} &=& \mathrm{sin}^2\omega t - xy
# \end{eqnarray}
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Euler's method is easily extended to simultaneous ODEs
#
# In the previous lesson we were introduced to Euler's method for the single variable case:
#
# \begin{equation}
# N(t+h) = N(t) + hf(N,t).
# \end{equation}
#
# This can be easily extended to the multi-variable case using vector notation:
#
# \begin{equation}
# \mathbf{r}(t+h) = \mathbf{r}(t) + h\mathbf{f}(\mathbf{r},t).
# \end{equation}
#
# We have seen that arrays can be easily represented in Python using the NumPy library. This allows us to do arithmetic with vectors directly (rather than using verbose workarounds such as `for` loops), so the code is not much more complicated than the one-variable case.
# + [markdown] slideshow={"slide_type": "slide"}
# ### The Lorenz equations are a famous set of simultaneous ODEs
#
# The Lorenz equations demonstrate that systems can be deterministic and yet be inherently unpredictable (even in the absence of quantum effects).
#
# \begin{eqnarray}
# \frac{\mathrm{d}x}{\mathrm{d}t} &=& \sigma(y-x) \\
# \frac{\mathrm{d}y}{\mathrm{d}t} &=& rx-y-xz \\
# \frac{\mathrm{d}z}{\mathrm{d}t} &=& xy-bz
# \end{eqnarray}
#
# There three dependent variables - $x$, $y$ and $z$, and one independent variable $t$. There are also three constants - $\sigma$, $r$ and $b$.
#
# For particular values of $\sigma$, $r$ and $b$ the Lorenz systems has <bold>chaotic behaviour</bold> (a strong dependence on the initial conditions) and for a subset of these, there is also fractal structure called the <bold>strange attractor</bold>.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Higher order ODEs can be re-cast as simultaneous ODEs and solved in the same way
#
# Many physical equations are second-order or higher. The general form for a second-order differential equation with one dependent variable is:
#
# \begin{equation}
# \frac{\mathrm{d}^2x}{\mathrm{d}t^2} = f\left(x,\frac{\mathrm{d}x}{\mathrm{d}t},t\right)
# \end{equation}
#
# Luckily, we can re-cast a higher order equation as a set of simultaneous equations, and then solve in the same way as above.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's use the non-linear pendulum as an example. For a pendulum with an art of length $l$ and a bob of mass $m$, Newton's second law ($F=ma$) provides the following equation of motion:
#
# \begin{equation}
# ml\frac{\mathrm{d}^2\theta}{\mathrm{d}t^2} = -mg\sin(\theta),
# \end{equation}
#
# where $\theta$ is the angle of displacement of the arm from the vertical and $l\frac{\mathrm{d}^2\theta}{\mathrm{d}t^2}$ is the acceleration of the mass in the tangential direction. The exact solution to this equation is unknown, but we now have the knowledge needed to find a numerical approximation.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# The equation can be re-written as:
#
# \begin{equation}
# \frac{\mathrm{d}^2\theta}{\mathrm{d}t^2} = -\frac{g}{l}\sin(\theta)
# \end{equation}
#
# We define a new variable $\omega$:
#
# \begin{equation}
# \frac{\mathrm{d}\theta}{\mathrm{d}t} = \omega
# \end{equation}
#
# and substitute this into the equation of motion:
#
# \begin{equation}
# \frac{\mathrm{d}\omega}{\mathrm{d}t} = -\frac{g}{l}\sin(\theta).
# \end{equation}
#
# These two expressions form a set of simultaneous equations that can be solved numerically using the method outlined above.
#
# + [markdown] slideshow={"slide_type": "slide"}
# -----
# Keypoints:
# - Computers don't care so much about the type of differential equation
# - Simulatenous ODEs can also be solved using numerical methods
# - Euler's method is easily extended to the multi-variable case
# - Higher order ODEs can be re-cast as simultaneous ODEs and solved the same way
# -----
# + [markdown] slideshow={"slide_type": "slide"}
# # The Runge-Kutta method
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# Questions:
# - How do I use the Runge-Kutta method for more accurate solutions?
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# Objectives:
# - Use the Runge-Kutta method, implemented in Python, to solve a first-order ODE
# - Compare results at different levels of approximation using the `matplotlib` library.
# + [markdown] slideshow={"slide_type": "slide"}
# ### The Runge-Kutta method is more accurate than Euler's method and runs just as fast
#
# So far we have used Euler's method for solving ODEs. We have learnt that, using this method, the final expression for the total error is linear in $h$.
#
# However for roughly the same compute time we can reduce the total error so it is of order $h^2$ by implementing another method - the <bold>Runge-Kutta method</bold>.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# > Note: It is common to use the Runge-Kutta method for solving ODEs given the improved accuracy over Euler's method. However Euler's method is still commonly used for PDEs (where there are other, larger, sources of error).
#
#
# > Note: The Runge-Kutta method is actually a family of methods. In fact, Euler's method is the first-order Runge-Kutta method. There is then the second-order Runge-Kutta method, third-order Runge-Kutta method, and so on..
# + [markdown] slideshow={"slide_type": "subslide"}
# Euler's method does not take into account the curvature of the solution, whilst Runge-Kutta methods do, by calculating the gradient at intermediate points in the (time-)step.
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ### Runge-Kutta methods are derived from Taylor expansion(s) around intermediate point(s)
#
# To derive the second-order Runge-Kutta method we:
#
# 1) estimate $x(t+h)$ using a Taylor expansion around $t+\frac{h}{2}$:
#
# \begin{equation}
# x(t+h) = x(t+\frac{h}{2}) + \frac{h}{2}\left(\frac{\mathrm{d}x}{\mathrm{d}t}\right)_{t+\frac{h}{2}} + \frac{h^2}{8}\left(\frac{\mathrm{d}^2x}{\mathrm{d}t^2}\right)_{t+\frac{h}{2}}+\mathcal{O}(h^3)
# \end{equation}
#
# 2) estimate $x(t)$ using a Taylor expansion around $t-\frac{h}{2}$:
#
# \begin{equation}
# x(t) = x(t+\frac{h}{2}) - \frac{h}{2}\left(\frac{\mathrm{d}x}{\mathrm{d}t}\right)_{t+\frac{h}{2}} + \frac{h^2}{8}\left(\frac{\mathrm{d}^2x}{\mathrm{d}t^2}\right)_{t+\frac{h}{2}}+\mathcal{O}(h^3)
# \end{equation}
#
# 3) Subtract Equation 2 from Equation 1 and re-arrange:
#
# \begin{eqnarray}
# x(t+h) &=& x(t) + h\left(\frac{\mathrm{d}x}{\mathrm{d}t}\right))_{t+\frac{h}{2}}+\mathcal{O}(h^3) \\
# x(t+h) &=& x(t) + hf(x(t+\frac{h}{2}),t+\frac{h}{2})+\mathcal{O}(h^3)
# \end{eqnarray}
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# > Note: the $h^2$ term has completely disappeared, and the error term is now order $h^3$. We can say that this approximation is now accurate to order $h^2$.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# The problem is that this requires knowledge of $x(t+\frac{h}{2})$ which we don't currently have. We can however estimate this using the Euler method!
#
# \begin{equation}
# x(t+\frac{h}{2}) = x(t) + \frac{h}{2}f(x,t).
# \end{equation}
#
# Substituting this into Equation 3 above, we can write the method for a single step as follows:
#
# \begin{eqnarray}
# k_1 &=& hf(x,t) \\
# k_2 &=& hf(x+\frac{k_1}{2},t+\frac{h}{2})\\
# x(t+h) &=& x(t) + k_2
# \end{eqnarray}
#
# See how $k_1$ is used to give an estimate for $x(t+\frac{h}{2})$ ($k_2$), which is then substituted into the third equation to give an estimate for $x(t+h)$.
#
# > Note: Higher orde Runge-Kutta methods can be derived in a similar way - by calculating the Taylor series around various points and then taking a linear combination of the resulting expansions. As we increase the number of intermediate points, we increase the accuracy of the method. The downside is that the equations get increasingly complicated.
#
# + [markdown] slideshow={"slide_type": "slide"}
# Keypoints:
# - The Runge-Kutta method is more accurate than Euler's method and runs just as fast
# - Runge-Kutta methods are derived from Taylor expansion(s) around intermediate point(s)
# - Runge-Kutta methods can be applied using the Python skills we have developed
# - We can easily compare our various models using the `matplotlib` plotting library
| slides/ODE-slides.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="m32JazI5pTmN"
# 1. Write a Python Program to print Prime Numbers between 2 numbers
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1550, "status": "ok", "timestamp": 1618459708182, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="VjdRJx49pqka" outputId="d33268b7-8bf1-4814-bc38-5cae3b89cf04"
start = 1
end = 20
print('Approach1')
for i in range(start, end+1):
#if i>1:
for j in range(2,i):
if(i % j==0):
break
else:
print(i)
print('Approach2')
for num in range(start,end):
if all(num % i != 0 for i in range(2, num)):
print(num)
# + [markdown] id="2G9C4AKjpVu1"
# 2. Write a Sort function to sort the elements in a list
#
# + id="u2PRHUN6tjTE"
l = [2,5,7,9,6,1,3,54,34,36, 76,21,99]
l.sort(reverse = True)
print(l)
# + [markdown] id="Tmz_Fs3upX9x"
# 3. Write a sorting function without using the list.sort function (descendring Order)
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1062, "status": "ok", "timestamp": 1618460181849, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="xD5Bg-leuUMx" outputId="2744d6a1-85a9-48af-ba38-0c6b0b8e2bdd"
data_list = [2,5,7,9,6,1,3,54,34,36, 76,21,99]
new_list = []
for a in data_list:
min = a
for x in data_list:
if x > min:
min = x
new_list.append(min)
print(new_list)
# + [markdown] id="tX9ncPrzpZMt"
# 4. Write a Python program to print Fibonacci Series
#
# ---
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2176, "status": "ok", "timestamp": 1618460496317, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="FdEJlWZFvrNl" outputId="204b1b2b-7550-4b37-a9ed-a8b527d83289"
def Fib(n):
if n == 0: return 0
elif n ==1: return 1
else: return Fib(n-1) + Fib(n-2)
for i in range(0,12):
print(Fib(i))
# + [markdown] id="MMMQWWj6pakf"
# 5. Write a Python program to print a list in reverse
#
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 855, "status": "ok", "timestamp": 1618460546751, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="hnvfdk4Cw2by" outputId="3517268c-c528-4f9b-d251-8b1116c4a2f1"
l = [2,5,7,9,6,1,3,54,34,36, 76,21,99]
def rev(n):
return n[:: -1]
rev(l)
# + [markdown] id="XFJvetPuvqbh"
#
# + [markdown] id="VH0K47Bkpb1v"
# 6. Write a Python program to check whether a string is a Palindrome or not
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2877, "status": "ok", "timestamp": 1618466115992, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="dvE6iWnxFrLk" outputId="c92e5e1c-9870-4169-9d12-201e5fa2dfac"
def isPalindrome(s):
return s == s[::-1]
def isPalindrome2(s):
rev = ''.join(reversed(s))
return s == rev
s = 'malayalam'
print(isPalindrome(s))
# + [markdown] id="SElfy-4wpeQu"
# 7. Write a Python program to print set of duplicates in a list
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 940, "status": "ok", "timestamp": 1618466197612, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="pWeLlMN4GWRE" outputId="2dcc4afc-0b25-4845-c6d9-b4f66ae69224"
l = [2,5,7,9,6,1,3,3,54,34,36, 76,21,8,8,99]
print(set([ x for x in l if l.count(x) > 1]))
# + [markdown] id="P1lr_Ilwpe0Y"
# 8. Write a Python program to print number of words in a given sentence
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 497, "status": "ok", "timestamp": 1618466246014, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="OL0Sb117Golh" outputId="c49d9f78-af20-4faa-998c-06aab16c6a22"
s = 'I am good boy'
print(len(s.split()))
# + [markdown] id="yPpBzSEWpgPq"
# 9. Given an array arr[] of n elements, write a Python function to search a given element x in arr[].
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 800, "status": "ok", "timestamp": 1618471501233, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="fAmKjBlwGxzQ" outputId="afef9b1f-efb8-46e5-f588-8e83920f898f"
def search(arr, x):
for i in range(len(arr)):
if arr[i] == x:
return i
print('Not present in the list')
l = [2,5,7,9,6,1,3,3,54,34,36,76,21,8,8,99]
search(l, 54)
# + [markdown] id="naVAROjiphdk"
# 10. Write a Python program to implement a Binary Search
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 887, "status": "ok", "timestamp": 1618471660691, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="A9R3ZtJnbDyx" outputId="578ee2aa-7a3c-4a72-c8e5-fec2dbb64275"
def binary_search(arr, x):
low = 0
high = len(arr) - 1
mid = 0
while low <= high:
mid = (high + low) // 2
# If x is greater, ignore left half
if arr[mid] < x:
low = mid + 1
# If x is smaller, ignore right half
elif arr[mid] > x:
high = mid - 1
# means x is present at mid
else:
return mid
# If we reach here, then the element was not present
return -1
# Test array
arr = [ 2, 3, 4, 10, 40 ]
x = 10
# Function call
result = binary_search(arr, x)
if result != -1:
print("Element is present at index", str(result))
else:
print("Element is not present in array")
# + [markdown] id="0FQ1OZDipi2K"
# 11. Write a Python program to plot a simple bar chart
#
# + [markdown] id="XGTkbcFzpkMc"
# 12. Write a Python program to join two strings (Hint: using join())
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 813, "status": "ok", "timestamp": 1618471788279, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="IH9djmWFbfcX" outputId="d2070194-fbad-439c-a38a-80f04e56c893"
str1 = 'Python'
str2 = 'is best'
res = '-'.join((str1, str2))
print(res)
# + [markdown] id="nxpN4bQepl_m"
# 13. Write a Python program to extract digits from given string
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 860, "status": "ok", "timestamp": 1618471932883, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="VzBDmh-pb7UG" outputId="bf87ab75-1808-4f76-bf2e-96021a6ae6bf"
str1 = 'r5m2a3n4'
res = ''.join(filter(lambda i: i.isdigit(), str1))
print(res)
# + [markdown] id="R9ADkyK6pmiQ"
# 14. Write a Python program to split strings using newline delimiter
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1040, "status": "ok", "timestamp": 1618472000967, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="ODFq_8o4cfaw" outputId="24983b01-6c03-40fb-b535-08ba7fc52d4e"
str1 = 'python is best language'
res = (str1.rstrip()).split(' ')
print(res)
# + [markdown] id="xqeUXGmSpoCh"
# 15. Given a string as your input, delete any reoccurring character, and return the new string.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"elapsed": 1063, "status": "ok", "timestamp": 1618472187340, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="hlQUgag8cwIo" outputId="42cba7d4-05c1-4f2f-a847-7c01d3cf1372"
def delete_recurring_char(str):
s = set()
output = ''
for c in str:
if c not in s:
s.add(c)
output+= c
return output
delete_recurring_char('venkataramana')
# + [markdown] id="OgsbmcJYzry9"
# 16. Split a string number into Number with comma sepertor
# + colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"elapsed": 942, "status": "ok", "timestamp": 1618495084204, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-xBOD0T0K43k/AAAAAAAAAAI/AAAAAAABJL0/st5MF0z46Vs/s64/photo.jpg", "userId": "00242112927648752124"}, "user_tz": -480} id="65VLdf9YztdX" outputId="b6d6d200-c823-46fc-cdfa-dd7f01536a38"
def reverse(string):
string = "".join(reversed(string))
return string
def format_integer(number, seperator=','):
s = reverse(str(number))
count = 0
result = ''
for char in s:
count = count+1
if count % 3 == 0:
if len(s) == count:
result = char + result
else:
result = seperator + char + result
else:
result = char + result
return result
input = '12345678'
format_integer(input)
# + id="Voctohkt0VHX"
| basic maths/Top 15 Python Coding Interview Questions with Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Basic of Machine Learning
#
# 1.Data:
# Image, Text, Audio, Video, Structured data
# 2.A model of how to transform the data
# 3.A loss function to measure how well we’re doing
# 4.An algorithm to tweak the model parameters such that the loss function is minimized
# ### ND Array in MXNet
import mxnet as mx
from mxnet import nd
import numpy as np
mx.random.seed(1)
x = nd.empty((3, 4))
print(x)
x = nd.ones((3, 4))
x
y = nd.random_normal(0, 1, shape=(3, 4))
print y
print y.shape
print y.size
x * y
nd.exp(y)
nd.dot(x, y.T)
# Memory Host
print "The current mem host y is {}".format(id(y))
y[:] = x + y
print "The current mem host after add + assigning y is {}".format(id(y))
y = x + y
print "The current mem host after add y is {}".format(id(y))
print y
print y[1:3]
print y[1:3,1:2]
print x
x[1,2] = 9
print x
x[1:2,1:3] = 5
print x
# #### Brodcasting
x = nd.ones(shape=(3,3))
print('x = ', x)
y = nd.arange(3)
print('y = ', y)
print('x + y = ', x + y)
# #### nd array <-> Numpy
a = x.asnumpy()
print "The type of a is {}".format(type(a))
y = nd.array(a)
print "The type of a is {}".format(type(y))
# ### Deal data with gpu
# +
# z = nd.ones(shape=(3,3), ctx=mx.gpu(0))
# z
# +
# x_gpu = x.copyto(mx.gpu(0))
# print(x_gpu)
# -
# ## Scala, Vector, Matrices, Tensors
# scalars
x = nd.array([3.0])
y = nd.array([2.0])
print 'x + y = ', x + y
print 'x * y = ', x * y
print 'x / y = ', x / y
print 'x ** y = ', nd.power(x,y)
# convert it to python scala
x.asscalar()
# +
# Vector
u = nd.arange(4)
print('u = ', u)
print u[3]
print len(u)
print u.shape
a = 2
x = nd.array([1,2,3])
y = nd.array([10,20,30])
print(a * x)
print(a * x + y)
# +
# Matrices
x = nd.arange(20)
A = x.reshape((5, 4))
print A
print 'A[2, 3] = ', A[2, 3]
print('row 2', A[2, :])
print('column 3', A[:, 3])
print A.T
# -
# Tensor
X = nd.arange(24).reshape((2, 3, 4))
print 'X.shape =', X.shape
print 'X =', X
u = nd.array([1, 2, 4, 8])
v = nd.ones_like(u) * 2
print 'v =', v
print 'u + v', u + v
print 'u - v', u - v
print 'u * v', u * v
print 'u / v', u / v
print nd.sum(u)
print nd.mean(u)
print nd.sum(u) / u.size
print nd.dot(u, v)
print nd.sum(u * v)
# Matrices multiple Vector
print nd.dot(A, u)
print nd.dot(A, A.T)
print nd.norm(u)
print nd.sqrt(nd.sum(u**2))
print nd.sum(nd.abs(u))
# ### Todo
# Additive axioms
# Multiplicative axioms
# Distributive axioms
#
| DS502-1704/MXNet-week1-part1/gluon-MNIST/Deep Learning Fundamentals-Gluon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importing libraries
import pickle
import itertools
import string
import re
import numpy as np
import pandas as pd
import multiprocessing
from collections import Counter
from scipy.sparse import csr_matrix
import gensim
from gensim.models import Doc2Vec
from gensim.models.doc2vec import TaggedDocument
from gensim.test.test_doc2vec import ConcatenatedDoc2Vec
import nltk
from nltk.corpus import stopwords
from nltk.stem.wordnet import WordNetLemmatizer
from tqdm import tqdm
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import seaborn as sn
from sklearn import utils
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn import svm
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.cluster import MiniBatchKMeans
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
# ### Load the preprocessed data from the data_directory
data_directory = "Generated_Files/data_after_preprocessing.csv"
# ### We devide the data into 3 groups:
# * Group 1: full data
# * Group 2: data with four large categories which have more than 1000 companies each
# * Group 3: seven categories of data, number of companies in each category is same but small
#
# ### In the function selectGroup, giving 1, 2 or 3 as input parameter to selet the relevant data for experiment
# read the data from directory, then select the group
# of data we want to process.
def selectGroup(directory, group_nr):
data = pd.read_csv(directory, sep='\t')
if group_nr == 1:
return data
if group_nr == 2:
df_healthcare_group=data[data['Category'] == 'HEALTHCARE GROUP'].sample(n=1041,replace=False)
df_business_financial_services=data[data['Category'] == 'BUSINESS & FINANCIAL SERVICES'].sample(n=1041,replace=False)
df_consumer_service_group=data[data['Category'] == 'CONSUMER SERVICES GROUP'].sample(n=1041,replace=False)
df_information_technology_group=data[data['Category'] == 'INFORMATION TECHNOLOGY GROUP'].sample(n=1041,replace=False)
df_clean = pd.concat([df_healthcare_group, df_business_financial_services,df_consumer_service_group,df_information_technology_group])
return df_clean.sample(frac=1)
if group_nr == 3:
df_healthcare_group=data[data['Category'] == 'HEALTHCARE GROUP'].sample(n=219,replace=False)
df_business_financial_services=data[data['Category'] == 'BUSINESS & FINANCIAL SERVICES'].sample(n=219,replace=False)
df_consumer_service_group=data[data['Category'] == 'CONSUMER SERVICES GROUP'].sample(n=219,replace=False)
df_information_technology_group=data[data['Category'] == 'INFORMATION TECHNOLOGY GROUP'].sample(n=219,replace=False)
df_industry_goods=data[data['Category'] == 'INDUSTRIAL GOODS & MATERIALS GROUP'].sample(n=219,replace=False)
df_consumer_goods=data[data['Category'] == 'CONSUMER GOODS GROUP'].sample(n=219,replace=False)
df_energy=data[data['Category'] == 'ENERGY & UTILITIES GROUP'].sample(n=219,replace=False)
df_clean = pd.concat([df_healthcare_group, df_business_financial_services,df_consumer_service_group,df_information_technology_group,df_industry_goods,df_consumer_goods,df_energy])
return df_clean.sample(frac=1)
# Select and Split the data
data = selectGroup(data_directory, 1)
train, test = train_test_split(data, test_size=0.2, random_state=42)
# ### Train and Build the doc2vec model
# +
def tokenize_text(text):
tokens = []
for sent in nltk.sent_tokenize(text):
for word in nltk.word_tokenize(sent):
if len(word) < 2:
continue
tokens.append(word.lower())
return tokens
train_tagged = train.apply(
lambda r: TaggedDocument(words=tokenize_text(r['clean']), tags=[r.Category]), axis=1)
test_tagged = test.apply(
lambda r: TaggedDocument(words=tokenize_text(r['clean']), tags=[r.Category]), axis=1)
cores = multiprocessing.cpu_count()
# Distributed Bag of Words (DBOW)
model_dbow = Doc2Vec(dm=0, vector_size=300, negative=5, hs=0, min_count=2, sample = 0, workers=cores)
model_dbow.build_vocab([x for x in tqdm(train_tagged.values)])
# -
# %%time
for epoch in range(30):
model_dbow.train(utils.shuffle([x for x in tqdm(train_tagged.values)]), total_examples=len(train_tagged.values), epochs=1)
model_dbow.alpha -= 0.002
model_dbow.min_alpha = model_dbow.alpha
# Distributed Memory (DM)
model_dmm = Doc2Vec(dm=1, dm_mean=1, vector_size=300, window=10, negative=5, min_count=1, workers=5, alpha=0.065, min_alpha=0.065)
model_dmm.build_vocab([x for x in tqdm(train_tagged.values)])
# %%time
for epoch in range(30):
model_dmm.train(utils.shuffle([x for x in tqdm(train_tagged.values)]), total_examples=len(train_tagged.values), epochs=1)
model_dmm.alpha -= 0.002
model_dmm.min_alpha = model_dmm.alpha
# Pair Distributed Bag of Words (DBOW) and Distributed Memory (DM)
model_dbow.delete_temporary_training_data(keep_doctags_vectors=True, keep_inference=True)
model_dmm.delete_temporary_training_data(keep_doctags_vectors=True, keep_inference=True)
new_model = ConcatenatedDoc2Vec([model_dbow, model_dmm])
# Infer vectors from the doc2vec model
def get_vectors(model, tagged_docs):
sents = tagged_docs.values
targets, regressors = zip(*[(doc.tags[0], model.infer_vector(doc.words, steps=20)) for doc in sents])
return targets, regressors
y_train, X_train = get_vectors(new_model, train_tagged)
print("Infer vector done for train data")
y_test, X_test = get_vectors(new_model, test_tagged)
# ### Use PCA and T-SNE to visualize the data distribution of different classes when using doc2vec to present them
# +
#Original Category Visualization
X = X_train + X_test
y = y_train + y_test
X = csr_matrix(pd.DataFrame(list(X)))
categories = ['BUSINESS & FINANCIAL SERVICES', 'CONSUMER GOODS GROUP', 'CONSUMER SERVICES GROUP', 'ENERGY & UTILITIES GROUP', 'HEALTHCARE GROUP', 'INDUSTRIAL GOODS & MATERIALS GROUP', 'INFORMATION TECHNOLOGY GROUP']
new_label = [0, 1, 2, 3, 4, 5, 6]
mydict=dict(zip(categories, new_label))
y_tsne = np.zeros(len(y))
for index, label in enumerate(y):
y_tsne[index] = mydict[label]
def plot_tsne_pca(data, labels):
max_label = max(labels+1)
max_items = np.random.choice(range(data.shape[0]), size=1000, replace=False)
pca = PCA(n_components=2).fit_transform(data[max_items,:].todense())
tsne = TSNE().fit_transform(PCA(n_components=50).fit_transform(data[max_items,:].todense()))
idx = np.random.choice(range(pca.shape[0]), size=300, replace=False)
label_subset = labels[max_items]
label_subset = [cm.hsv(i/max_label) for i in label_subset[idx]]
f, ax = plt.subplots(1, 2, figsize=(14, 6))
ax[0].scatter(pca[idx, 0], pca[idx, 1], c=label_subset)
ax[0].set_title('PCA Cluster Plot')
ax[1].scatter(tsne[idx, 0], tsne[idx, 1], c=label_subset)
ax[1].set_title('TSNE Cluster Plot')
plot_tsne_pca(X, y_tsne)
# -
# ### Use Unsupervised Learning method to classify the companies and visualize the clusters by PCA and T-SNE based on the vectors generated from Doc2Vec - Kmeans
# +
#K-Means Assigned Category Visualization
def find_optimal_clusters(data, max_k):
iters = range(2, max_k+1, 2)
sse = []
for k in iters:
sse.append(MiniBatchKMeans(n_clusters=k, init_size=128, batch_size=256, random_state=20).fit(data).inertia_)
print('Fit {} clusters'.format(k))
f, ax = plt.subplots(1, 1)
ax.plot(iters, sse, marker='o')
ax.set_xlabel('Cluster Centers')
ax.set_xticks(iters)
ax.set_xticklabels(iters)
ax.set_ylabel('SSE')
ax.set_title('SSE by Cluster Center Plot')
def plot_tsne_pca(data, labels):
max_label = max(labels+1)
max_items = np.random.choice(range(data.shape[0]), size=1000, replace=False)
pca = PCA(n_components=2).fit_transform(data[max_items,:].todense())
tsne = TSNE().fit_transform(PCA(n_components=50).fit_transform(data[max_items,:].todense()))
idx = np.random.choice(range(pca.shape[0]), size=300, replace=False)
label_subset = labels[max_items]
label_subset = [cm.hsv(i/max_label) for i in label_subset[idx]]
f, ax = plt.subplots(1, 2, figsize=(14, 6))
ax[0].scatter(pca[idx, 0], pca[idx, 1], c=label_subset)
ax[0].set_title('PCA Cluster Plot')
ax[1].scatter(tsne[idx, 0], tsne[idx, 1], c=label_subset)
ax[1].set_title('TSNE Cluster Plot')
find_optimal_clusters(X, 8)
clusters = MiniBatchKMeans(n_clusters=7, init_size=256, batch_size=256, random_state=20).fit_predict(X)
plot_tsne_pca(X, clusters)
# +
# Map category names to K-mean clusters based on data accuracy
def Calculate_accuracy(clusters, actual_label):
count = 0
for index, cluster in enumerate(clusters):
if cluster==actual_label[index]:
count+=1
accuracy = count/len(clusters)*1.0
return accuracy
def assign_clusters(original_label, permu):
categories = ['BUSINESS & FINANCIAL SERVICES', 'CONSUMER GOODS GROUP', 'CONSUMER SERVICES GROUP', 'ENERGY & UTILITIES GROUP', 'HEALTHCARE GROUP', 'INDUSTRIAL GOODS & MATERIALS GROUP', 'INFORMATION TECHNOLOGY GROUP']
mydict=dict(zip(categories, permu))
actual_label = np.zeros(len(original_label))
for index, label in enumerate(original_label):
actual_label[index] = mydict[label]
return actual_label
numbers=np.zeros(7)
for i in range(7):
numbers[i] = i
permu = list(itertools.permutations(numbers))
best_accuracy = 0
for i in range(len(permu)):
actual_label = assign_clusters(data['Category'].values, permu[i])
accuracy = Calculate_accuracy(clusters, actual_label)
if best_accuracy<accuracy:
best_accuracy=accuracy
final_label = actual_label
category = permu[i]
else: best_accuracy=best_accuracy
print(classification_report(final_label,clusters))
df_cm = pd.DataFrame(confusion_matrix(final_label,clusters), range(7),
range(7))
sn.set(font_scale=1.4)#for label size
ax = sn.heatmap(df_cm, cmap="Blues", annot=True,annot_kws={"size": 16}, fmt='g')# font size
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
print(confusion_matrix(final_label,clusters))
# -
# ### Use different Supervised Learning methods to classify the companies based on the vectors generated from Doc2Vec - Linear SVM, Logistic Regression, KNN
# +
#Linear SVM Classifier
def linear_svc_classifier(train_x, train_y, test_x, test_y):
print("start svm")
classifier_svm = svm.LinearSVC(verbose=1)
# classifier_svm = svm.SVC(kernel='linear')
classifier_svm.fit(train_x, train_y)
predictions = classifier_svm.predict(test_x)
confusion = confusion_matrix(test_y, predictions)
print(confusion)
y_true = ["BUSINESS & FINANCIAL SERVICES", "CONSUMER GOODS GROUP", "CONSUMER SERVICES GROUP", "ENERGY & UTILITIES GROUP", "HEALTHCARE GROUP", "INDUSTRIAL GOODS & MATERIALS GROUP", "INFORMATION TECHNOLOGY GROUP"]
y_pred = ["BUSINESS & FINANCIAL SERVICES", "CONSUMER GOODS GROUP", "CONSUMER SERVICES GROUP", "ENERGY & UTILITIES GROUP", "HEALTHCARE GROUP", "INDUSTRIAL GOODS & MATERIALS GROUP", "INFORMATION TECHNOLOGY GROUP"]
df_cm = pd.DataFrame(confusion, y_true, y_pred)
df_cm.index.name = 'Actual'
df_cm.columns.name = 'Predicted'
plt.figure(figsize = (10,7))
ax = sn.heatmap(df_cm, cmap="Blues", annot=True,annot_kws={"size": 16}, fmt='g')# font size
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
print(classification_report(test_y, predictions))
print(accuracy_score(test_y, predictions))
linear_svc_classifier(X_train, y_train, X_test, y_test)
# +
# Logistic Classifier
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
logistic_pred = logreg.predict(X_test)
confusion = confusion_matrix(y_test, logistic_pred)
print(confusion)
y_true = ["BUSINESS & FINANCIAL SERVICES", "CONSUMER GOODS GROUP", "CONSUMER SERVICES GROUP", "ENERGY & UTILITIES GROUP", "HEALTHCARE GROUP", "INDUSTRIAL GOODS & MATERIALS GROUP", "INFORMATION TECHNOLOGY GROUP"]
y_pred = ["BUSINESS & FINANCIAL SERVICES", "CONSUMER GOODS GROUP", "CONSUMER SERVICES GROUP", "ENERGY & UTILITIES GROUP", "HEALTHCARE GROUP", "INDUSTRIAL GOODS & MATERIALS GROUP", "INFORMATION TECHNOLOGY GROUP"]
df_cm = pd.DataFrame(confusion, y_true, y_pred)
df_cm.index.name = 'Actual'
df_cm.columns.name = 'Predicted'
plt.figure(figsize = (10,7))
ax = sn.heatmap(df_cm, cmap="Blues", annot=True,annot_kws={"size": 16}, fmt='g')# font size
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
print(classification_report(y_test, logistic_pred))
print(accuracy_score(y_test, logistic_pred))
# +
# KNN classifier
def grid_search(Max, X_data, y_data):
KNN_Best_Accuracy = 0
for n in range(5,Max):
modelknn = KNeighborsClassifier(n_neighbors=n)
modelknn.fit(X_data,y_data)
Accuracy = Calculate_accuracy(modelknn.predict(X_data), y_data)
if KNN_Best_Accuracy< Accuracy:
KNN_Best_Accuracy = Accuracy
Bestknnmodel = KNeighborsClassifier(n_neighbors=n)
Bestknnmodel.fit(X_data,y_data)
else: KNN_Best_Accuracy = KNN_Best_Accuracy
#print(n)
#print(KNN_Best_Accuracy)
return Bestknnmodel
Bestknnmodel = grid_search(6, X_train, y_train)
predicted_labels_knn = Bestknnmodel.predict(X_test)
print(classification_report(predicted_labels_knn,y_test))
y_true = ["BUSINESS & FINANCIAL SERVICES", "CONSUMER GOODS GROUP", "CONSUMER SERVICES GROUP", "ENERGY & UTILITIES GROUP", "HEALTHCARE GROUP", "INDUSTRIAL GOODS & MATERIALS GROUP", "INFORMATION TECHNOLOGY GROUP"]
y_pred = ["BUSINESS & FINANCIAL SERVICES", "CONSUMER GOODS GROUP", "CONSUMER SERVICES GROUP", "ENERGY & UTILITIES GROUP", "HEALTHCARE GROUP", "INDUSTRIAL GOODS & MATERIALS GROUP", "INFORMATION TECHNOLOGY GROUP"]
df_cm = pd.DataFrame(confusion_matrix(y_test, predicted_labels_knn), y_true, y_pred)
df_cm.index.name = 'Actual'
df_cm.columns.name = 'Predicted'
plt.figure(figsize = (10,7))
ax = sn.heatmap(df_cm, cmap="Blues", annot=True,annot_kws={"size": 16}, fmt='g')# font size
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
print(confusion_matrix(predicted_labels_knn,y_test))
# -
# ### Save the doc2vec model for future usage
# save the model to disk
filename = 'Generated_Files/doc2vec_model.sav'
pickle.dump(new_model, open(filename, 'wb'))
| Codes/4.Doc2Vec+K-means, SVM, Logistic Regression, KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 1. Create a program to help the user type faster. Give him a word and ask him to write it five times. check how many seconds it took him to type theh word at each round.
# ### 2. In the end, tell the user how many mistakes were made and show a chart with the typing speed evolution during the 5 rounds.
# +
import matplotlib.pyplot as plt
import time as t
times =[]
mistakes = 0
print("This program will help you to type faster. You will have to type the word 'programming' as fast as you can for five times.")
input("Press enter to continue.")
while len(times) < 5:
start = t.time()
word = input('Type the word: ')
end = t.time()
time_elapsed = end - start
times.append(time_elapsed)
if(word.lower() != 'programming') :
mistakes += 1
print('You made ' + str(mistakes) + ' mistake(s).')
print('Now let us see your evolution')
t.sleep(3)
x = [1, 2, 3, 4, 5]
y = times
plt.bar(x, y)
legend = ['1', '2', '3', '4', '5']
plt.xticks(x, legend)
plt.xlabel('Attempts')
plt.ylabel('Time in seconds')
plt.title('Your typing evolution')
plt.show()
# -
| Exercise-Time & pyplot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Copyright 2020 <NAME>, <NAME>
#
#Licensed under the Apache License, Version 2.0 (the "License");
#you may not use this file except in compliance with the License.
#You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
#Unless required by applicable law or agreed to in writing, software
#distributed under the License is distributed on an "AS IS" BASIS,
#WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
#See the License for the specific language governing permissions and
#limitations under the License.
from collections import Counter
from collections import defaultdict
from matplotlib import pyplot as plt
from sklearn.feature_extraction import DictVectorizer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.preprocessing import OneHotEncoder
from sklearn.metrics import confusion_matrix, accuracy_score
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.model_selection import KFold,StratifiedKFold
from sklearn import metrics
import pickle
import math
import re
import enchant
import pandas as pd
import os
import glob
import numpy as np
# np.random.seed(512)
from sklearn.linear_model import LogisticRegression, LogisticRegressionCV
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn import metrics
from sklearn.model_selection import KFold,StratifiedKFold
from sklearn.model_selection import GridSearchCV
from sklearn.feature_extraction.text import CountVectorizer
from sklearn import linear_model
import numpy as np # linear algebra
import pandas as pd
import pickle
import joblib
# +
xtrain = pd.read_csv('../../Benchmark-Labeled-Data/data_train.csv')
xtest = pd.read_csv('../../Benchmark-Labeled-Data/data_test.csv')
xtrain = xtrain.sample(frac=1,random_state=100).reset_index(drop=True)
print(len(xtrain))
y_train = xtrain.loc[:,['y_act']]
y_test = xtest.loc[:,['y_act']]
y_train
# +
dict_label = {
'numeric': 0,
'categorical': 1,
'datetime': 2,
'sentence': 3,
'url': 4,
'embedded-number': 5,
'list': 6,
'not-generalizable': 7,
'context-specific': 8
}
y_train['y_act'] = [dict_label[i] for i in y_train['y_act']]
y_test['y_act'] = [dict_label[i] for i in y_test['y_act']]
y_train
# -
def ProcessStats(data,y):
data1 = data[['total_vals', 'num_nans', '%_nans', 'num_of_dist_val', '%_dist_val', 'mean',
'std_dev', 'min_val', 'max_val','has_delimiters', 'has_url', 'has_email', 'has_date', 'mean_word_count',
'std_dev_word_count', 'mean_stopword_total', 'stdev_stopword_total',
'mean_char_count', 'stdev_char_count', 'mean_whitespace_count',
'stdev_whitespace_count', 'mean_delim_count', 'stdev_delim_count',
'is_list', 'is_long_sentence']]
data1 = data1.reset_index(drop=True)
data1 = data1.fillna(0)
def abs_limit(x):
if abs(x) > 10000:
return 10000*np.sign(x)
return x
column_names_to_normalize = ['total_vals', 'num_nans', '%_nans', 'num_of_dist_val', '%_dist_val', 'mean',
'std_dev', 'min_val', 'max_val','has_delimiters', 'has_url', 'has_email', 'has_date', 'mean_word_count',
'std_dev_word_count', 'mean_stopword_total', 'stdev_stopword_total',
'mean_char_count', 'stdev_char_count', 'mean_whitespace_count',
'stdev_whitespace_count', 'mean_delim_count', 'stdev_delim_count',
'is_list', 'is_long_sentence']
for col in column_names_to_normalize:
data1[col] = data1[col].apply(abs_limit)
print(column_names_to_normalize)
x = data1[column_names_to_normalize].values
x = np.nan_to_num(x)
x_scaled = StandardScaler().fit_transform(x)
df_temp = pd.DataFrame(x_scaled, columns=column_names_to_normalize, index=data1.index)
data1[column_names_to_normalize] = df_temp
y.y_act = y.y_act.astype(float)
print(f"> Data mean: {data1.mean()}\n")
print(f"> Data median: {data1.median()}\n")
print(f"> Data stdev: {data1.std()}")
return data1
# +
vectorizerName = CountVectorizer(ngram_range=(2, 2), analyzer='char')
vectorizerSample = CountVectorizer(ngram_range=(2, 2), analyzer='char')
def FeatureExtraction(data,data1,flag):
arr = data['Attribute_name'].values
arr = [str(x) for x in arr]
print(len(arr))
# data = data.fillna(0)
arr1 = data['sample_1'].values
arr1 = [str(x) for x in arr1]
arr2 = data['sample_2'].values
arr2 = [str(x) for x in arr2]
print(len(arr1),len(arr2))
if flag:
X = vectorizerName.fit_transform(arr)
X1 = vectorizerSample.fit_transform(arr1)
X2 = vectorizerSample.transform(arr2)
else:
X = vectorizerName.transform(arr)
X1 = vectorizerSample.transform(arr1)
X2 = vectorizerSample.transform(arr2)
attr_df = pd.DataFrame(X.toarray())
sample1_df = pd.DataFrame(X1.toarray())
sample2_df = pd.DataFrame(X2.toarray())
print(len(data1),len(attr_df),len(sample1_df),len(sample2_df))
data2 = pd.concat([data1, attr_df,sample1_df,sample2_df], axis=1, sort=False)
# data2 = pd.concat([attr_df, sample1_df], axis=1, sort=False)
# data2 = pd.concat([sample1_df, sample2_df, sample3_df, sample4_df], axis=1, sort=False)
# print(len(data2))
return data2
# return sample1_df
# +
xtrain1 = ProcessStats(xtrain,y_train)
xtest1 = ProcessStats(xtest,y_test)
X_train = FeatureExtraction(xtrain,xtrain1,1)
X_test = FeatureExtraction(xtest,xtest1,0)
X_train_new = X_train.reset_index(drop=True)
y_train_new = y_train.reset_index(drop=True)
X_train_new = X_train_new.values
y_train_new = y_train_new.values
k = 5
kf = KFold(n_splits=k, random_state = 100)
avg_train_acc, avg_test_acc = 0, 0
val_arr = [0.0001, 0.001, 0.01, 0.1, 1, 10, 100, 1000, 10000, 100000]
avgsc_lst, avgsc_train_lst, avgsc_hld_lst = [], [], []
avgsc, avgsc_train, avgsc_hld = 0, 0, 0
best_param_count = {'cval': {}}
for train_index, test_index in kf.split(X_train_new):
X_train_cur, X_test_cur = X_train_new[train_index], X_train_new[test_index]
y_train_cur, y_test_cur = y_train_new[train_index], y_train_new[test_index]
X_train_train, X_val, y_train_train, y_val = train_test_split(
X_train_cur, y_train_cur, test_size=0.25, random_state=100)
bestPerformingModel = LogisticRegression(
penalty='l2', multi_class='multinomial', solver='lbfgs', C=1)
bestscore = 0
print('='*10)
for val in val_arr:
clf = LogisticRegression(
penalty='l2', multi_class='multinomial', solver='lbfgs', C=val)
clf.fit(X_train_train, y_train_train)
sc = clf.score(X_val, y_val)
print(f"[C: {val}, accuracy: {sc}]")
if bestscore < sc:
bestcval = val
bestscore = sc
bestPerformingModel = clf
if str(bestcval) in best_param_count['cval']:
best_param_count['cval'][str(bestcval)] += 1
else:
best_param_count['cval'][str(bestcval)] = 1
bscr_train = bestPerformingModel.score(X_train_cur, y_train_cur)
bscr = bestPerformingModel.score(X_test_cur, y_test_cur)
bscr_hld = bestPerformingModel.score(X_test, y_test)
avgsc_train_lst.append(bscr_train)
avgsc_lst.append(bscr)
avgsc_hld_lst.append(bscr_hld)
avgsc_train = avgsc_train + bscr_train
avgsc = avgsc + bscr
avgsc_hld = avgsc_hld + bscr_hld
print()
print(f"> Best C: {bestcval}")
print(f"> Best training score: {bscr_train}")
print(f"> Best test score: {bscr}")
print(f"> Best held score: {bscr_hld}")
print('='*10)
# +
print(avgsc_train_lst)
print(avgsc_lst)
print(avgsc_hld_lst)
print(avgsc_train/k)
print(avgsc/k)
print(avgsc_hld/k)
y_pred = bestPerformingModel.predict(X_test)
bscr_hld = bestPerformingModel.score(X_test, y_test)
print(bscr_hld)
# -
bestPerformingModel.score(X_test, y_test)
| MLFeatureTypeInference/Source-Code/Models/Logistic_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from sklearn.metrics.pairwise import cosine_similarity
from sklearn.cluster import DBSCAN, KMeans
from sklearn.manifold import TSNE
from tqdm import tqdm
from spherecluster import SphericalKMeans
import matplotlib.pyplot as plt
# -
RANDOM_SEED = 42
library_embeddings = np.load('../models/libraries_embeddings.npy')
library_embeddings.shape
library_embeddings /= np.linalg.norm(library_embeddings, axis=1, keepdims=True)
libraries = [line.strip() for line in open('../models/libraries_embeddings_dependencies.txt')]
len(libraries)
def run_kmeans(n_clusters, embedding, init='k-means++', normalize=False):
if normalize:
embedding = embedding.copy() / np.linalg.norm(embedding, axis=1, keepdims=True)
clusterizer = KMeans(n_clusters, init=init, random_state=RANDOM_SEED).fit(embedding)
labels = clusterizer.predict(embedding)
return clusterizer, labels
def print_closest(ind, k=20):
print(f"Library: {libraries[ind]}")
for other in list(reversed(np.argsort(cosine_sims[ind])))[:k]:
print(f"{libraries[other]} | {cosine_sims[ind,other]:.4f}")
def compute_score(embeddings, cluster_centers):
return embeddings.dot(cluster_centers.T).max(axis=1).sum()
# +
kmeans_scores = []
uniform_scores = []
trials = np.array([2, 4, 8, 12, 16, 20, 24, 28, 32, 36, 40, 44, 48, 52, 56, 60, 64])
for n_clusters in tqdm(trials):
kmeans, clusters = run_kmeans(n_clusters, library_embeddings)
kmeans_scores.append(compute_score(library_embeddings, kmeans.cluster_centers_))
uniform_scores.append([])
for t in range(100):
uniform_centers = np.random.normal(size=(n_clusters, 32))
uniform_centers /= np.linalg.norm(uniform_centers, axis=1, keepdims=True)
uniform_scores[-1].append(compute_score(library_embeddings, uniform_centers))
# -
kmeans_scores = np.array(kmeans_scores)
uniform_scores = np.array(uniform_scores)
mean_scores = uniform_scores.mean(axis=1)
std_scores = uniform_scores.std(axis=1)
plt.figure(figsize=(20,10))
plt.xticks(trials)
plt.errorbar(x=trials, y=kmeans_scores-mean_scores, yerr=std_scores, fmt='o')
np.savetxt('gap_statistic.txt', np.column_stack([trials, kmeans_scores, mean_scores, std_scores]), fmt='%.3f')
N_CLUSTERS = 44
kmeans, labels = run_kmeans(N_CLUSTERS, library_embeddings)
# +
def get_idf(line):
lib, val = line.strip().split(';')
return lib, float(val)
idfs = [get_idf(line) for line in open('../models/idfs.txt')]
# -
idfs.sort(key=lambda lib: libraries.index(lib[0]))
idfs = np.array([idf[1] for idf in idfs])
labels = np.array(labels)
from analysis import get_name, get_year, read_dependencies
reqs = read_dependencies()
dep_index = {dep: i for i, dep in enumerate(libraries)}
# +
cluster_labels = [''] * N_CLUSTERS
cluster_repos = [''] * N_CLUSTERS
for c in range(N_CLUSTERS):
inds = np.where(labels == c)[0]
sorted_by_idf = np.argsort(idfs[inds])
closest_repos = []
edge = 4
while len(closest_repos) < 20:
edge -= 1
closest_repos = []
for repo, deps in reqs.items():
year = get_year(repo)
if year != '2020':
continue
name = get_name(repo)
total_c = 0
for dep in deps:
if labels[dep_index[dep]] == c:
total_c += 1
if total_c < edge:
continue
closest_repos.append((total_c / len(deps), name))
closest_repos.sort(reverse=True)
print('~'*20)
print(f'Cluster {c}:')
for ind, (dist, repo_name) in zip(sorted_by_idf[:20], closest_repos):
repo_under_ind = repo_name.rfind('_')
repo_name = f'https://github.com/{repo_name[:repo_under_ind]}/{repo_name[repo_under_ind+1:]}'
true_ind = inds[ind]
print(f'{repo_name} | {dist:.3f} | {libraries[true_ind]} | {cosine(library_embeddings[true_ind], kmeans.cluster_centers_[c]):.3f}')
cluster_labels[c] += " | " + libraries[true_ind]
print('~'*20)
print()
# -
def readfile(f):
labels = []
for line in open(f):
line = line.strip()
if line != '':
labels.append(line)
return labels
examples = np.array(readfile('examples.txt')).reshape((-1, 3))
common_labels = readfile('labels.txt')
print(len(examples), len(common_labels))
common_labels = [f"{label} | {', '.join(exs)}" for label, exs in zip(common_labels, examples)]
# +
import numpy as np
import pickle
from matplotlib import pyplot as plt
from scipy.cluster.hierarchy import dendrogram
from sklearn.cluster import AgglomerativeClustering
def plot_dendrogram(model, **kwargs):
# Create linkage matrix and then plot the dendrogram
# create the counts of samples under each node
counts = np.zeros(model.children_.shape[0])
n_samples = len(model.labels_)
for i, merge in enumerate(model.children_):
current_count = 0
for child_idx in merge:
if child_idx < n_samples:
current_count += 1 # leaf node
else:
current_count += counts[child_idx - n_samples]
counts[i] = current_count
linkage_matrix = np.column_stack([model.children_, model.distances_,
counts]).astype(float)
# Plot the corresponding dendrogram
dendrogram(linkage_matrix, **kwargs)
X = kmeans.cluster_centers_
n_clusters = len(X)
# setting distance_threshold=0 ensures we compute the full tree.
model = AgglomerativeClustering(distance_threshold=0, n_clusters=None, affinity='cosine', linkage='average')
model = model.fit(X / np.linalg.norm(X, keepdims=True, axis=1))
plt.title('Hierarchical Clustering Dendrogram')
# plot the top three levels of the dendrogram
# plt.xlabel("Number of points in node (or index of point if no parenthesis).")
# plt.show()
plt.figure(figsize=(10, max(10, n_clusters // 3)))
# fig = plt.figure()
# ax = fig.add_subplot(1, 1, 1)
plot_dendrogram(model, truncate_mode='level', p=40, labels=common_labels, color_threshold=.8,
orientation='left', leaf_font_size=16)
plt.savefig(f'dendrogram.pdf', bbox_inches='tight', dpi=100, format='pdf')
| analyze_library_embeddings/analyze_library_embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Load Dataset**
from keras.datasets import imdb
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(
num_words=10000)
word_index = imdb.get_word_index()
# **One Hot Encode Input Data**
import numpy as np
# +
def vectorize_sequences(sequences, dimension=10000):
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1
return results
# Vectorize training and test data
x_train = vectorize_sequences(train_data)
x_test = vectorize_sequences(test_data)
# Vectorize training and test labels
y_train = np.asarray(train_labels).astype('float32')
y_test = np.asarray(test_labels).astype('float32')
# -
# **Build Network**
# +
# Creat stack of the network
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(1, activation='sigmoid'))
# -
# Configure loss function
# and optimizer
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
# Create validation set
x_val = x_train[:10000]
partial_x_train = x_train[10000:]
y_val = y_train[:10000]
partial_y_train = y_train[10000:]
# Fit data to network
history = model.fit(partial_x_train,
partial_y_train,
epochs=4,
batch_size=512,
validation_data=(x_val, y_val))
# +
import matplotlib.pyplot as plt
# Access history of the fit session
history_dict = history.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
# Number of epochs trained
epochs = range(1, len(history_dict['acc']) + 1)
# Plot losses
plt.plot(epochs, loss_values, 'bo', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# +
# Plot accuracy training
plt.clf()
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
# Plot acc values for training and val
plt.plot(epochs, acc_values, 'bo', label='Training acc')
plt.plot(epochs, val_acc_values, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# Evaluate trained model on the test set
results = model.evaluate(x_test, y_test)
results
model.predict(x_test)
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise-2
# Add the specified code for each code cell, running the cells _in order_.
# Define a list **`names`** that includes your name and the names of at least 2 people close to you.
names = ["Elizabeth", "Viv", "Tom"]
# Use a **list comprehension** to _map_ the `names` list to a `shouted` list that includes all the names, but in upper case. Print out the `shouted` list.
shouted = list(map(str.upper, names))
print(shouted)
# Define a new list **`words`** that contains each of the words to the following [song lyric](https://www.youtube.com/watch?v=StTqXEQ2l-Y) (don't include the notes):
# ```
# ♫ everything is awesome everything is cool when you are part of a TEEEEAM ♫
# didn't have a dime, though I always had a vision, had to have high, high hopes
# ```
#
# _Hint:_ use the `split()` string method.
lyric = "didn't have a dime, though I always had a vision, had to have high, high hopes"
words = lyric.split()
print(words)
# Use a **list comprehension** to filter the list of words to only include words that have (strictly) more than 4 letters.
more_than_four = [i for i in words if len(i) > 4]
print(more_than_four)
# Use a **list comprehension** to create a list of initials for each word in the lyric that is _not_ 3 letters or shorter.
initials = [i[0].upper() + "." for i in words if len(i) > 3]
print(initials)
# Use a **list comprehension** (with nested `for` clauses) to produce a list of _tuples_ representing the different combinations of numbers found on two rolled six-sided dice:
# ```
# [(1,1), (1,2), (1,3), ...]
# ```
# Note that there should be 36 different combinations!
rolls = [(d1, d2) for d1 in range(1,7) for d2 in range(1,7)]
print(rolls)
# Use a **list comprehension** to produce a list of tuples representing the rolls on two dice that sum up to 7 OR 11.
rolls2 = [(d1, d2) for d1 in range(1,7) for d2 in range(1,7) if d1+d2 == 7 or d1+d2 == 11]
print(rolls2)
| exercise-2/exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Remove input cells at runtime (nbsphinx)
import IPython.core.display as d
d.display_html('<script>jQuery(function() {if (jQuery("body.notebook_app").length == 0) { jQuery(".input_area").toggle(); jQuery(".prompt").toggle();}});</script>', raw=True)
# # Particle classification (DL2)
# **Author(s):**
#
# - Dr. <NAME> (CEA-Saclay/IRFU/DAp/LEPCHE), 2020
#
# based on previous work by <NAME>.
#
# **Description:**
#
# This notebook contains benchmarks for the _protopipe_ pipeline regarding particle classification at the DL2 level.
# The plots shown here are performed also by `protopipe.scripts.model_diagnostics` which will be discontinued.
#
# **NOTES:**
# - these benchmarks will be cross-validated and migrated in cta-benchmarks/ctaplot,
# - follow [this](https://www.overleaf.com/16933164ghbhvjtchknf) document by adding new benchmarks or proposing new ones.
#
# **Requirements:**
#
# To run this notebook you will 1 merged files of DL2 data per particle type produced on the grid with protopipe.
#
# When comparing the performance of protopipe with that of CTAMARS (2019), the MC production to be used and the appropriate set of files to use for this notebook can be found [here](https://forge.in2p3.fr/projects/step-by-step-reference-mars-analysis/wiki#The-MC-sample ).
#
# The data format required to run the notebook is the current one used by _protopipe_ .
#
# **Development and testing:**
#
# As with any other part of _protopipe_ and being part of the official repository, this notebook can be further developed by any interested contributor.
# The execution of this notebook is not currently automatic: it must be done locally by the user - preferably _before_ pushing a pull-request.
#
# Please, if you wish to contribute to this notebook, before pushing clear all the output and remove local directory paths (leave empty strings).
#
# **TODO:**
# * check any missing model diagnostics product
# * add remaining benchmarks from CTA-MARS comparison
# * same for EventDisplay
# ## Table of contents
# - [Estimate distribution](#Estimate-distribution)
# - [ROC curve](#ROC-curve)
# + [markdown] nbsphinx="hidden"
# ## Imports
# -
# %matplotlib inline
import os
from pathlib import Path
import yaml
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from sklearn.metrics import auc, roc_curve
# + [markdown] nbsphinx="hidden"
# ## Functions
# +
def load_config(name):
"""Load YAML configuration file."""
try:
with open(name, "r") as stream:
cfg = yaml.load(stream, Loader=yaml.FullLoader)
except FileNotFoundError as e:
print(e)
raise
return cfg
def plot_hist(ax, data, nbin, limit, norm=False, yerr=False, hist_kwargs={}, error_kw={}):
"""Utility function to plot histogram"""
bin_edges = np.linspace(limit[0], limit[-1], nbin + 1, True)
y, tmp = np.histogram(data, bins=bin_edges)
weights = np.ones_like(y)
if norm is True:
weights = weights / float(np.sum(y))
if yerr is True:
yerr = np.sqrt(y) * weights
else:
yerr = np.zeros(len(y))
centers = 0.5 * (bin_edges[1:] + bin_edges[:-1])
width = bin_edges[1:] - bin_edges[:-1]
ax.bar(centers, y * weights, width=width, yerr=yerr, error_kw=error_kw, **hist_kwargs)
return ax
def plot_roc_curve(ax, model_output, y, **kwargs):
"""Plot ROC curve for a given set of model outputs and labels"""
fpr, tpr, _ = roc_curve(y_score=model_output, y_true=y)
roc_auc = auc(fpr, tpr)
label = '{} (area={:.2f})'.format(kwargs.pop('label'), roc_auc) # Remove label
ax.plot(fpr, tpr, label=label, **kwargs)
return ax
def plot_evt_roc_curve_variation(ax, data_test, cut_list, model_output_name):
"""
Parameters
----------
ax: `~matplotlib.axes.Axes`
Axis
data_test: `~pd.DataFrame`
Test data
cut_list: `list`
Cut list
Returns
-------
ax: `~matplotlib.axes.Axes`
Axis
"""
color = 1.
step_color = 1. / (len(cut_list))
for i, cut in enumerate(cut_list):
c = color - (i + 1) * step_color
data = data_test.query(cut)
if len(data) == 0:
continue
opt = dict(color=str(c), lw=2, label='{}'.format(cut.replace('reco_energy', 'E')))
plot_roc_curve(ax, data[model_output_name], data['label'], **opt)
ax.plot([0, 1], [0, 1], color='navy', lw=2, linestyle='--')
return ax
# + [markdown] nbsphinx="hidden"
# ## Load data
# -
# First we check if a _plots_ folder exists already.
# If not, we create it.
Path("./plots").mkdir(parents=True, exist_ok=True)
# EDIT ONLY THIS CELL WITH YOUR SETUP INFORMATION
parentDir = "" # path to 'shared_folder'
analysisName = ""
# +
configuration = os.path.join(parentDir,
"shared_folder/analyses",
analysisName,
"configs/classifier.yaml")
cfg = load_config(configuration)
if cfg["Method"]["use_proba"] is True:
model_output = 'gammaness'
output_range = [0, 1]
else:
model_output = 'score'
output_range = [-1, 1]
indir = os.path.join(parentDir,
"shared_folder/analyses",
analysisName,
"data/DL2")
data_gamma = pd.read_hdf(os.path.join(indir, 'DL2_tail_gamma_merged.h5'), "/reco_events").query('NTels_reco >= 2')
data_electron = pd.read_hdf(os.path.join(indir, 'DL2_tail_electron_merged.h5'), "/reco_events").query('NTels_reco >= 2')
data_proton = pd.read_hdf(os.path.join(indir, 'DL2_tail_proton_merged.h5'), "/reco_events").query('NTels_reco >= 2')
data_gamma['label'] = np.ones(len(data_gamma))
data_electron['label'] = np.zeros(len(data_electron))
data_proton['label'] = np.zeros(len(data_proton))
# + [markdown] nbsphinx="hidden"
# ## Benchmarks
# -
# ### Estimate distribution
# [back to top](#Table-of-contents)
# +
energy_bounds = np.logspace(np.log10(cfg["Diagnostic"]["energy"]["min"]),
np.log10(cfg["Diagnostic"]["energy"]["max"]),
cfg["Diagnostic"]["energy"]["nbins"] + 1)
ncols = int(cfg["Diagnostic"]["energy"]["nbins"] / 2)
n_ax = len(energy_bounds) - 1
nrows = int(n_ax / ncols) if n_ax % ncols == 0 else int((n_ax + 1) / ncols)
nrows = nrows
fig, axes = plt.subplots(nrows=nrows, ncols=ncols, figsize=(5 * ncols, 3 * nrows))
if nrows == 1 and ncols == 1:
axes = [axes]
else:
axes = axes.flatten()
for idx in range(len(energy_bounds) - 1):
ax = axes[idx]
# Data selection
query = 'true_energy >= {} and true_energy < {}'.format(energy_bounds[idx], energy_bounds[idx + 1])
gamma = data_gamma.query(query)
proton = data_proton.query(query + ' and offset < {}'.format(1.))
electron = data_electron.query(query + ' and offset < {}'.format(1.))
data_list = [gamma, proton]
# Graphical stuff
color_list = ['blue', 'red', 'green']
edgecolor_list = ['black', 'black', 'green']
fill_list = [True, True, False]
ls_list = ['-', '-', '--']
lw_list = [2, 2, 2]
alpha_list = [0.2, 0.2, 1]
label_list = ['Gamma', 'Proton', 'Electron']
opt_list = []
err_list = []
for jdx, data in enumerate(data_list):
opt_list.append(dict(edgecolor=edgecolor_list[jdx], color=color_list[jdx], fill=fill_list[jdx], ls=ls_list[jdx], lw=lw_list[jdx], alpha=alpha_list[jdx], label=label_list[jdx]))
err_list.append(dict(ecolor=color_list[jdx], lw=lw_list[jdx], alpha=alpha_list[jdx], capsize=3, capthick=2,))
for jdx, data in enumerate(data_list):
ax = plot_hist(
ax=ax, data=data[model_output], nbin=50, limit=output_range,
norm=True, yerr=False,
hist_kwargs=opt_list[jdx],
error_kw=err_list[jdx],
)
ax.set_title('E=[{:.3f},{:.3f}] TeV'.format(energy_bounds[idx], energy_bounds[idx + 1]), fontdict={'weight': 'bold'})
ax.set_xlabel(model_output)
ax.set_ylabel('Arbitrary units')
ax.set_xlim(output_range)
ax.legend(loc='upper left')
ax.grid()
plt.tight_layout()
fig.savefig(f"./plots/particle_classification_estimate_distribution_protopipe_{analysisName}.png")
# -
# ### ROC curve
# [back to top](#Table-of-contents)
# +
data = pd.concat([data_gamma, data_electron, data_proton])
plt.figure(figsize=(5,5))
ax = plt.gca()
cut_list = ['reco_energy >= {:.2f} and reco_energy <= {:.2f}'.format(
energy_bounds[i],
energy_bounds[i+1]
) for i in range(len(energy_bounds) - 1)]
plot_evt_roc_curve_variation(ax, data, cut_list, model_output)
ax.legend(loc='lower right', fontsize='small')
ax.set_xlabel('False positive rate')
ax.set_ylabel('True positive rate')
ax.grid()
plt.tight_layout()
fig.savefig(f"./plots/particle_classification_roc_curve_protopipe_{analysisName}.png")
# -
| docs/contribute/benchmarks/DL2/benchmarks_DL2_particle-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="TERQPzGCB6Y6"
# # Installing the latest python 3 version and its necessary packages
#
# We strongly recommend installing Miniconda. There are installers for the 3 main Operating Systems (Linux, Windows & MacOS) https://docs.conda.io/en/latest/miniconda.html.
#
# To use the full 'power' of `python` you'll need a package manager to download and install all the necessary . The default one is `pip`, but in our current configuration we recommend you use `conda` (for the interested people among you, [here](https://www.anaconda.com/using-pip-in-a-conda-environment/) is a link to a blog post explaining why mixing both is bad idea).
#
# Once you've installed miniconda, open a terminal and install the following packages:
#
# `sklearn`, `numpy`, `pandas`, `matplotlib` by doing:
#
# `conda install <packages>`
# + [markdown] colab_type="text" id="c-Abe7wFhGmi"
# # What is a Jupyter Notebook?
#
# Python can be executed via the command line (by calling `python <program>.py` or inside a so called [`notebook`](https://jupyter.org/install).
#
# A notebook is a file composed of `cells` which can be executed seperately (`ctrl`+`enter` or `shift`+`enter` to also go to the next one) but that **share** a same memory system. This means that variables initialized in a certain cell can be accessed from any other cell as long as the former is executed *before* the latter. Try executing the 2 following cells
# + colab={} colab_type="code" id="Hi998DpvXYfV"
a = 5
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="uzx3DFjdi-_C" outputId="0343db04-49cc-48ad-afdf-9e8fc7431d54"
print(a)
# + [markdown] colab_type="text" id="kDAMgk7iviDI"
# A `cell` can thus be used to write and execute code or to write text. The default text format is [Markdown](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet). Text mode also supports $\LaTeX$ math environments which can be defined with the `$`-sign: $\forall b \in \mathcal{B}: b \ge \frac{5}{2}$
#
# + [markdown] colab_type="text" id="8OgNt5EPXZU0"
# # The Fundamentals
#
# The python language syntax decided to do away with curly brackets and decided on seperating code-blocks by line indentations. The colon (`:`) indicates the start of such a new indentation
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="da85iq-i6QDn" outputId="f6821f7d-b6b6-4baa-f854-a88a06c67b84"
a = 5
if a == 5:
print("a is equal to 5")
else:
print("Sadly a is not a beautiful 5 anymore :'(")
# + [markdown] colab_type="text" id="4O7kABW56MKf"
# Python is strongly **dynamically** typed language. This means that a variable's `type` is only checked at runtime (= when executing the code), the same was already true for `R` and `Matlab`. If there is a term you don't understand, you'll find a handy glossary [here](https://docs.python.org/3.1/glossary.html).
#
# The base types are the following:
# * `int`: integer values
# * `float`: floating point values
# * `str`: strings of characters
# * `bool`: `True` or `False`
#
#
# But also more complex ones:
# * `list`: A list of items
# * `tuple`: An immutable list of items
# * `dict`: A `key`-`value` store, the keys can be of various types as long as they are hashable
# * `set`: A hashset of unique values
#
# The number of elements in thos data structures can be obtained by calling the `len` function. Note that, contrary to some other languages (i.e. `Matlab`), **the indexation starts at zero**.
# + colab={"base_uri": "https://localhost:8080/", "height": 109} colab_type="code" id="_FupfXyAngZY" outputId="6dd4bf04-0aa0-41e7-8fb7-0dd2b54af6a3"
# (Text preceded by a '#' is ignored)
# Base types
a = 5
print(f"Example of integer division: {a//2} vs floating point division: {a/2}")
b = 3.5
c = "Hey I'm a suite of characters"
always_true = True or False
always_false = True and False
always_false = not True
# Advanced types
li = [1,2,3]
li.append(5)
print(f"I'm a list of length {len(li)}")
tu = (7,9,8)
t1, t2, t3 = tu
print(f"Does t2 contain the same values as the second element in the tuple? {t2==tu[1]}")
dico = {
"hey": 1,
56897: 6
}
print(f"At key 'hey' in dico the value is: {dico['hey']}")
se = set([1,1,1,1,2,3,4,5,5,5,6])
print(f"This set only has {len(se)} elements")
# + [markdown] colab_type="text" id="aBFXkLGqpVf6"
# (By now you'll have noted that we used a strange `string` format called `f-string` to print the values behind the variables, [here](https://realpython.com/python-f-strings/) you can find a more detailed explanation on the inner workings.)
#
# There is also a whole slew of even more versatile data structures in the `collections` [package](https://docs.python.org/3/library/collections.html). A handy one is the `defaultdict` which gives a default `value` to a previously unseen `key`.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="54TsRpFhpnYT" outputId="feb285ad-f903-48b5-a93e-93c973b6b42e"
# Importing a specific function or class from another package follows this syntax
# from <package> import <function>
from collections import defaultdict
a = defaultdict(int)
print(f"The default value is {a['whatever']}")
# + [markdown] colab_type="text" id="Odv_5ZT_toOd"
# # Iterators
#
# There are a few easy ways to iterate over data structures in python, the most important ones are listed below, but this is far from an exhaustive list.
# + colab={"base_uri": "https://localhost:8080/", "height": 109} colab_type="code" id="ARSWs7_VXe7J" outputId="ecf08e5c-65c9-421a-a455-dac1c2349d5d"
# The basic iterator or for loop
print("Range 1")
for i in range(3):
print(i, end=', ')
print("\n\nRange 2")
for i in range(4,9,2):
print(i, end=', ')
# + colab={"base_uri": "https://localhost:8080/", "height": 274} colab_type="code" id="8Qb1MSm3xE2h" outputId="7ba8174e-43c8-46cd-e302-a57c4a7a619c"
# Iterating over a list
print("Iterating over list")
for elem in li:
print(elem, end=', ')
print("\n\nIterating over list with indices")
for index, elem in enumerate(li):
print(f"At index {index}: {elem}")
print("\nIterating over 2 list of the same size simultaneously")
li2 = li.copy() # duplicates a list
li2[1] = 150
for elem, elem2 in zip(li, li2):
print(elem, elem2)
# + colab={"base_uri": "https://localhost:8080/", "height": 201} colab_type="code" id="lQtvyTlg2D8Y" outputId="8e45c75b-b22c-48fd-99af-281656312e01"
# Iterating over a dictionary
print("Simple dict iteration")
for key in dico: # An equivalent formulation would be: for key in dico.keys()
print(f"Value {dico[key]} at key {key}")
print("\nIterating over values only")
for value in dico.values():
print(value, end=', ')
print("\n\nIterating over both at once")
for key, value in dico.items():
print(f"Value {value} at key {key}")
# + [markdown] colab_type="text" id="g8OS_CkNyMM3"
# # Functions and classes
#
# Python supports both functional programming and object oriented programming with inheritance (like in Java).
#
# As was already stated, python is dynamically typed, so during function definition you don't need to specify an argument's type. You can however give it a default value.
# + colab={"base_uri": "https://localhost:8080/", "height": 109} colab_type="code" id="So55Rcht3Y32" outputId="438f1503-9205-42ce-fdd1-7b763f7e5a0a"
# Example of function definition
def line(x, alpha=1, c=0):
""" Returns the y-value of the line with coefficients:
- alpha (steepness)
- c (offset)
at point x
"""
return alpha*x + c
# OK
print(line(5))
print(line(5, 2, 1))
print(line(5, c=1, alpha=2)) # If you name the arguments, order is not important
print(line(x=5, c=1, alpha=2))
# Not ideal
print(line(c=1, alpha=2, x=5)) # But moving the arguments without defaults like
# this, is not really good practice
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="N4iO8gpr79n8" outputId="2f7683a6-d12b-4d9f-d502-bc9cdae4dae5"
# Example of a class definition
class Line():
def __init__(self, alpha=1, c=0): # This is the object's constructor
""" Creates the Line objectwith coefficients:
- alpha (steepness)
- c (offset)
"""
self.alpha = alpha
self.c = c
def get_y(self, x):
""" Returns the value of the line at point x
"""
return self.alpha*x + self.c
simple_line = Line(alpha=2, c=1)
print(simple_line.get_y(5))
# + [markdown] colab_type="text" id="0vYo54YlxSn-"
# # Pitfalls
#
# Python is not without its flaws, and one you should really look out for is the scope of variables. For example in most languages the following example would throw an error, but not here. This is due to python not really limiting the scope of certain variables inside a function
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="LwHUjVpvxa9G" outputId="1df5f813-e3eb-4e4c-91e6-347ac42e114c"
for i in range(6):
pass
print("i after for loop = ", i) # i still exists!!
def test():
print("i in test = ", i) # i is still acessible here too!!
j = 5
test()
try:
print(j) # But j is not accessible here
except:
print("j is not declared")
# + [markdown] colab_type="text" id="HZSOrA6QX5jz"
# # Final words
# Do not hesitate to read the excellent official [documentation](https://docs.python.org/3/) or the Q&A's on StackOverflow to find new ways to write better code, but never forget to cite your sources and have fun!
# -
# Je teste le code
# ...
# Jo
#
# Je df
#
print("Salut")
| Exercises/0 - Introduction Jupiter/PythonIntro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# SML-17 Workshop #2: 2b Evaluation of classifiers
# This week we'll be using the *SPAM base* dataset from the UCI machine learning dataset repository. This dataset comprises a few thousand emails that have been annotated as being spam or not, and the several features have been created from the email text (e.g., presence of certain important words and characters, too many CAPITALS etc.) Note that for many practical applications defining the features is one of the hardest and most important steps. Thankfully this has already been done for us, so we can deploy and evaluate our machine learning algorithms directly.
#
# Please see http://archive.ics.uci.edu/ml/datasets/Spambase for a full description of the dataset and feature definitions.
# Please follow the instructions from the previous lab for importing numpy etc, e.g.,
# %pylab inline
import numpy as np
import matplotlib.pyplot as plt
# ### Loading the SPAM data
# The first step is to load the data, which we'll download from
# http://archive.ics.uci.edu/ml/machine-learning-databases/spambase/spambase.data
# +
# download the datafile
import sys
if sys.version_info[0] >= 3:
from urllib.request import urlretrieve
else:
from urllib import urlretrieve
#url = "https://staffwww.dcs.shef.ac.uk/people/T.Cohn/campus_only/mlai13/spambase.data.data"
url = "http://archive.ics.uci.edu/ml/machine-learning-databases/spambase/spambase.data"
urlretrieve(url, 'spambase.data.data')
# load the CSV file as an array
Xandt = np.loadtxt('spambase.data.data', delimiter=',')
# randomly shuffle the rows, so as to remove any order bias
np.random.shuffle(Xandt)
# the last column are the response labels (targets), 0 = not spam, 1 = spam
# remap into -1 and +1 and take only the first 500 examples
t = Xandt[:500,-1] * 2 - 1
# and the remaining columns are the data
X = Xandt[:500,:-1]
print("Loaded", X.shape, "data points, and", t.shape, "labels,", t.min() , t.max())
# -
# The data has many different types of features, operating on different ranges and with overall very different distributions. Therefore, it is important to standardise each feature. Also, assumptions about the distributions of the input features are central to proofs of the generalisation bounds of several machine learning algorithms.
#
# Inspect the minimum, maximum, mean and standard deviation for each column, using `np.mean` and `np.std`. Note that you'll need to supply an axis if you're just interested in columns, e.g.,
tmp = np.mean(X, 0)
print(np.round(tmp,2))
print('Num of features : ' , len(tmp))
# Note that the last three features have mean orders of magnitude greater than the other features. (Take a look at their definitions.) Thus, we will standardise all features to have mean 0 and standard deviation 1.
X = (X - np.mean(X, 0)) / np.std(X, 0)
# Note that in previous labs we've plotted the data to get a feeling for how easily the data might be modelled, e.g., if it's linearly separable or requires polynomial basis functions etc. This technique is fine for data with 1 or 2 dimensions, but isn't so straight-forward for 57 dimensional data. (Note that dimensionality reduction methods, such as PCA, can be used to find the most important dimensions for viewing or exploiting in other learning algorithms.).
# ### K-nearest neighbour classifier
# To mix things up, for this session we will use a k-NN classifier. This classifier is incredibly simple: for each test point find the closest few ($k$) points in the training sets and return the majority label from these points. We will compare the setting of $k$, to try and find the *best* classifier.
# Let's start by defining a function *euclidean* to calculate the euclidean distance, $d(\mathbf{x}, \mathbf{z}) = \sqrt{(\mathbf{x} - \mathbf{z})^T (\mathbf{x} - \mathbf{z})}$.
def euclidean(x, z):
d = x - z
if len(d.shape) > 1 and d.shape[1] > 1:
return np.sqrt(np.diag(np.dot(d, d.T)))
else:
return np.sqrt(np.dot(d, d))
# Next, define a function to find nearby training points for a given test point under the euclidean distance. You may want to use `np.argsort` which returns the sort order (as an array indices) for an input array.
def neighbours(x, train_x, k):
dists = euclidean(train_x, x)
return np.argsort(dists)[:k]
# Note that this returns the indices of the training points, which can be used to look up the label. Now define the *k-NN* prediction algorithm, which processes each test point and finds the majority class of its neighbours in the training set.
def knn(test_x, train_x, train_t, k):
predict = np.zeros(test_x.shape[0])
for i in range(test_x.shape[0]):
ns = neighbours(train_x, test_x[i], k)
predict[i] = np.sign(np.sum(train_t[ns]))
return predict
# ### Heldout evaluation
# We have a large dataset of 500 examples. If we were to use all of these for training, we would have nothing left with which to evaluate the *generalisation error*. Recall that models often *overfit* the training sample, and therefore their performance on this set is often misleading: we have no way of telling if this is due to modelling the true problem or just fitting noise and other idiosyncracies of the training sample.
# To illustrate, compute the *training error* of the approach with a few different values of $k$, `[1,3,9,15,33,77]`. We'll later see if other evaluation methods lead to different conclusions.
for k in [1,3,9,15,33,77]:
print('%d-nn' % k, np.sum(knn(X, X, t, k) != t) / float(X.shape[0]))
# What is going on? Which value of k is best?
# #### Attempt 1: fixed validation set
# Now we'll try evaluating on *heldout validation* data. This has been excluded from training, so the model cannot *overfit*. Instead this serves as a fresh data sample, which reflects the intended usage of the classifier in a live scenario (i.e., processing new emails as they arrive in your inbox).
#
# One of the easiest methods is to slice the data into two parts. We'll half for training and half for testing. Note that these numbers are fairly arbitrary, but we do want enough test data to get a reliable error estimate.
N = X.shape[0]
cut = int(N/2)
Xtrain = X[:cut,:]
ttrain = t[:cut]
Xtest = X[cut:,:]
ttest = t[cut:]
print("There are", Xtrain.shape, "training samples, and", Xtest.shape, "heldout test samples")
print("Test split: for class label 1: ", np.sum(ttest ==1), "and class label -1", np.sum(ttest ==-1))
print("Train split: for class label 1: ", np.sum(ttrain ==1), "and class label -1", np.sum(ttrain ==-1))
# Now your job is to apply $k$-NN on this data for the various values of $k$, and evaluate their training and heldout error.
# IMPLEMENT ME
for k in [1,3,9,15,33,77]:
print('%d-nn' % k ,'training error',
np.sum(knn(Xtrain, Xtrain, ttrain, k) != ttrain) / float(Xtrain.shape[0]),'heldout error',
np.sum(knn(Xtest, Xtrain, ttrain, k) != ttest) / float(Xtest.shape[0]))
# Is the heldout error similar to the training error, or do you notice consistent differences? Can you explain why? What classifier might you select now, and does this match your earlier choice?
# Good going. We now have a decent spam classifier, similar to the ones in Google and other email providers (which are trained on *much* more data). You could fairly easily take this approach and the feature definitions and plug this into your own email inbox to classify incoming mails. Note that scaling this up to the full dataset would require a bit of engineering (or patience!), but otherwise would be straightforward.
# #### Attempt 2: Leave-one-out cross validation
# Another validation method is *cross-validation*, which is particularly suitable when you have only a small amount of data. This technique divides the data into parts, called *folds*. Then each fold is used for evaluation, and the model is trained on all other folds. This is repeated, such that we have heldout predictions for the entire dataset.
#
# The leave-one-out method is the most extreme version of cross-validation, and defines each fold as a single data-point.
# Implement k-NN to work using the leave-one-out method. This only requires a small change from the `knn()` method in order exclude the current point from consideration in the inner loop. Hint: start by taking the $k+1$ nearest neighbours, and then correct the result.
def knn_LOO(x, t, k):
predict = np.zeros(x.shape[0])
for i in range(x.shape[0]):
ns = neighbours(x, x[i], k+1)
if i in ns:
predict[i] = np.sign(np.sum(t[ns]) - t[i])
else:
ns = ns[:k]
predict[i] = np.sign(np.sum(t[ns]))
return predict
# Another Implementation by Yasmeen
def knn_LOO1(x, t, k):
predict = np.zeros(x.shape[0])
for i in range(x.shape[0]):
xtrain = np.delete(x, (i), axis=0)
ttrain = np.delete(t, (i), axis=0)
xtest = x[i]
ttest = t[i]
ns = neighbours(xtrain, xtest, k)
#print(xtrain.shape , ttrain.shape,xtest.shape, ttest, np.sign(np.sum(t[ns])))
predict[i] = np.sign(np.sum(ttrain[ns]))
return predict
# Try this out for a few values of k, and compare their error rates. How do these values compare to your results from using a fixed validation set? Are they more reliable, and would you expect the cross-validation error to be higher or lower?
for k in [1,3,9,15,33,77]:
print('%d-nn' % k,
'LOO error', np.sum(knn_LOO1(X, t, k) != t) / float(X.shape[0]))
for k in [1,3,9,15,33,77]:
print('%d-nn' % k,
'LOO error', np.sum(knn_LOO(X, t, k) != t) / float(X.shape[0]))
# ## Error types and ROC analysis
# Spam classification is an interesting test case. Consider the impact on an email user of various errors:
#
# 1. misclassifying a spam email as good
# 1. misclassifying a good email as spam
#
# We need to balance annoyance (1) with missing potentially important information (2).
#
# These two errors are referred to as Type I and Type II errors, and are treated uniformly when measuring accuracy (and 0/1 loss). In this case it is more informative to evaluate the outputs by counting each of the four categories: true positives (true spam which was classified as spam), false positives (good classified as spam), true negatives (good classified as good), and false negatives (spam classified as good).
pred = knn_LOO(X, t, 9)
print('true positives ', np.sum(t[pred == 1] == 1))
print('false positives', np.sum(t[pred == 1] == -1))
print('true negatives ', np.sum(t[pred == -1] == -1))
print('false negatives', np.sum(t[pred == -1] == 1))
# This shows that the data are skewed, i.e. there are more negatives than positives. Are the numbers of the two types of error are closely balanced?
# The ROC curve gives a better view of the compromise between the two error types. This graph shows the relationship between the *false positive rate* and *true positive rate*. Each point on the curve represents a different classifier. In our case we can vary the value of k, recording the FPR and TPR at each point. Randomly guessing will give us a straight line in ROC space, and we seek to do better by being above and left (higher TPR for a given FPR; lower FPR for a given TPR). For a thorough description please read the wikipedia page on ROC analysis http://en.wikipedia.org/wiki/Receiver_operating_characteristic
# +
fprs = []
tprs = []
for k in [1,2,3,5,9,15,33,77]:
pred = knn_LOO(X, t, k)
tp = np.sum(t[pred==1] == 1)
fp = np.sum(t[pred==1] == -1)
tn = np.sum(t[pred==-1] == -1)
fn = np.sum(t[pred==-1] == 1)
# now compute the TPR and FPR
fpr = fp / float(fp + tn)
tpr = tp / float(tp + fn)
plt.plot([0,fpr,1],[0,tpr,1],label='k = %d'%k)
fprs.append(fpr)
tprs.append(tpr)
plt.legend()
xlabel('FPR')
ylabel('TPR')
# +
plot(fprs, tprs, 'r-')
xlabel('FPR')
ylabel('TPR')
print (fprs , tprs)
# -
# Depending on our appetite for false positive errors, we can read off the corresponding accuracy for predictions (TPR) with respect to false negatives. Plotting the ROC curve for several classifiers can help us choose which is best while incorporating unbalanced loss functions.
| 1_KNN_CrossHalidation_LeaveOneOut_HeldOut/notebook/2b_performance_evaluation_answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + language="javascript"
# var kernel = IPython.notebook.kernel;
# var body = document.body,
# attribs = body.attributes;
# var command = "__filename__ = " + "'" + decodeURIComponent(attribs['data-notebook-name'].value) + "'";
# kernel.execute(command);
# -
print(__filename__)
# +
import os, sys, numpy as np, tensorflow as tf
from pathlib import Path
import time
try:
print(__file__)
__current_dir__ = str(Path(__file__).resolve().parents[0])
__filename__ = os.path.basename(__file__)
except NameError:
# jupyter notebook automatically sets the working
# directory to where the notebook is.
__current_dir__ = str(Path(os.getcwd()))
module_parent_dir = str(Path(__current_dir__).resolve().parents[0])
sys.path.append(module_parent_dir)
import LeNet_plus
__package__ = 'LeNet_plus'
from . import network
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
BATCH_SIZE = 250
SCRIPT_DIR = __current_dir__
FILENAME = __filename__
SUMMARIES_DIR = SCRIPT_DIR
SAVE_PATH = SCRIPT_DIR + "/network.ckpt"
### configure devices for this eval script.
USE_DEVICE = '/gpu:0'
session_config = tf.ConfigProto(log_device_placement=True)
session_config.gpu_options.allow_growth = True
# this is required if want to use GPU as device.
# see: https://github.com/tensorflow/tensorflow/issues/2292
session_config.allow_soft_placement = True
# -
with tf.Graph().as_default() as g, tf.device(USE_DEVICE):
# inference()
input, deep_features = network.inference()
labels, logits, loss_op = network.loss(deep_features)
eval = network.evaluation(logits, labels)
init = tf.initialize_all_variables()
with tf.Session(config=session_config) as sess:
saver = tf.train.Saver()
saver.restore(sess, SAVE_PATH)
# TEST_BATCH_SIZE = np.shape(mnist.test.labels)[0]
logits_output, deep_features_output, loss_value, accuracy = \
sess.run(
[logits, deep_features, loss_op, eval], feed_dict={
input: mnist.test.images[5000:],
labels: mnist.test.labels[5000:]
})
print("MNIST Test accuracy is ", accuracy)
import matplotlib.pyplot as plt
# %matplotlib inline
color_palette = ['#507ba6', '#f08e39', '#e0595c', '#79b8b3', '#5ca153',
'#edc854', '#af7ba2', '#fe9fa9', '#9c7561', '#bbb0ac']
from IPython.display import FileLink, FileLinks
# +
bins = {}
for logit, deep_feature in zip(logits_output, deep_features_output):
label = np.argmax(logit)
try:
bins[str(label)].append(list(deep_feature))
except KeyError:
bins[str(label)] = [list(deep_feature)]
plt.figure(figsize=(5, 5))
for numeral in map(str, range(10)):
try:
features = np.array(bins[numeral])
except KeyError:
print(numeral + " does not exist")
features = []
plt.scatter(
features[:, 0],
features[:, 1],
color=color_palette[int(numeral)],
label=numeral
)
plt.legend(loc=(1.1, 0.1), frameon=False)
title = 'MNIST LeNet++ with 2 Deep Features (PReLU)'
plt.title(title)
plt.xlabel('activation of hidden neuron 1')
plt.ylabel('activation of hidden neuron 2')
fname = './figures/' + title + '.png'
plt.savefig(fname, dpi=300, bbox_inches='tight')
FileLink(fname)
# -
| misc/deep_learning_notes/Proj_Centroid_Loss_LeNet/LeNet_plus/2D visualization of learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (oat-use)
# language: python
# name: oat-use
# ---
# %run "Common setup.ipynb"
from SALib.analyze.radial_ee import analyze
def incremental_radialee_analysis(lower, upper, step=1):
res, idx = [], []
for reps in range(lower, upper, step):
results = analyze(CIM_SPEC, numeric_vals[:reps], np_res[:reps], int(reps/step), seed=101)
total = results.to_df()
res.append(total.loc[tgt_param, ['mu_star', 'sigma']].tolist())
idx.append(int(reps))
# End for
return res, idx
# End incremental_radialee_analysis()
numeric_samples = pd.read_csv(f'{DATA_DIR}radial_10_numeric_samples.csv', index_col=0)
numeric_vals = numeric_samples[perturbed_cols].values
# +
cols = ['$\mu*$', '$\sigma$']
# Coupling disabled
radial_no_irrigation_results = pd.read_csv(f'{DATA_DIR}radial_no_irrigation_10_results.csv', index_col=0)
radial_no_irrigation_results['Avg. $/ML'].fillna(radial_no_irrigation_results["Avg. Annual Profit ($M)"], inplace=True)
np_res = radial_no_irrigation_results.loc[:, tgt_metric].values
runs = np_res.shape[0]
res, idx = incremental_radialee_analysis(54, runs+1, 54)
# -
disabled = pd.DataFrame(data=res, columns=cols, index=idx)
# +
# Coupling enabled
radial_with_irrigation_results = pd.read_csv(f'{DATA_DIR}radial_with_irrigation_10_results.csv', index_col=0)
radial_with_irrigation_results['Avg. $/ML'].fillna(radial_with_irrigation_results["Avg. Annual Profit ($M)"], inplace=True)
np_res = radial_with_irrigation_results.loc[:, tgt_metric].values
runs = np_res.shape[0]
res, idx = incremental_radialee_analysis(54, runs+1, 54)
# -
# plot_incremental_results(res, idx)
enabled = pd.DataFrame(data=res, columns=cols, index=idx)
# +
fig, axes = plt.subplots(1,2, figsize=(12,4), sharey=True, sharex=True)
disabled.plot(kind='bar',
legend=None,
use_index=True,
title='Disabled',
ax=axes[0],
rot=45)
enabled.plot(kind='bar',
legend=None,
use_index=True,
title='Enabled',
ax=axes[1],
rot=45).legend(
bbox_to_anchor=(1.45, 0.65)
)
axes[0].axhline(0.1, c='k', ls='--')
axes[1].axhline(0.1, c='k', ls='--')
fig.suptitle("Radial OAT", x=0.51, y=1.05, fontsize=22)
plt.xlabel("$N$", x=-0.1, labelpad=15);
# -
fig.savefig(f'{FIG_DIR}radial_oat_results.png', dpi=300, bbox_inches='tight')
# ---
#
# With full results
# +
tgt_result_idx = all_outputs.columns.tolist().index("SW Allocation Index")
numeric_samples = to_numeric_samples(all_inputs)[perturbed_cols]
numeric_vals = numeric_samples.values
np_res = all_outputs.values[:, tgt_result_idx]
rows = np_res.shape[0]
res, idx = incremental_radialee_analysis(54, rows+1, 54)
# -
pd.DataFrame(data=res[::2], columns=cols, index=idx[::2]).plot(kind='bar', figsize=(16,4));
| notebooks/2 Radial OAT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
import re
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.cross_validation import train_test_split
from sklearn.linear_model import LogisticRegression
df= pd.read_csv("ASTD.csv")
df2= pd.read_csv("ASTD.csv")
df.head()
# +
df.info()
# -
def cleaner(text):
text = text.lower()
text = re.sub("@[^\s]+","",text)
text = text.replace(":)","")
text = text.replace("@","")
text = text.replace("#","")
text = text.replace(":(","")
return text
def remove_stop_words(text):
sw = stopwords.words("arabic")
clean_words = []
text = text.split()
for word in text:
if word not in sw:
clean_words.append(word)
return " ".join(clean_words)
def stemming(text):
ps = PorterStemmer()
text = text.split()
stemmed_words = []
for word in text :
stemmed_words.append(ps.stem(word))
return " ".join(stemmed_words)
def run(text):
text = cleaner(text)
text = remove_stop_words(text)
text = stemming(text)
return text
df['txt'] = df['txt'].apply(run)
df2.iloc[1,1]
tfidf = TfidfVectorizer()
x = tfidf.fit_transform(df["txt"]).toarray()
y = df['sentiment'].values
y
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.33)
lr = LogisticRegression()
lr.fit(x_train,y_train)
y_pred = lr.predict(x_test)
lr.score(x_test,y_test)
test = "عادة لا اعلن عن مواقفي السياسيةلكن كله الا الدستور في كلمة واحدة:مهزلةو لو وصلنا لمهزلة الاستفتاء حقول لا"
test = run(test)
test = tfidf.transform([test]).toarray()
lr.predict(test)
from sklearn.metrics import classification_report
print(classification_report(y_pred,y_test))
from sklearn.metrics import confusion_matrix
confusion_matrix(y_pred,y_test)
| Sentiment Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cxbxmxcx/GenReality/blob/master/GEN_3_variation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="XhB46Q8JTpBd"
from sklearn.model_selection import train_test_split
import numpy as np
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
from scipy.stats import norm
# + id="9ITk0PPQb67p"
def function(x):
return norm.logpdf(x)
# + id="tNHQgiXUUFIa" outputId="9c533170-e951-4c48-cb01-21398e01a1fa" colab={"base_uri": "https://localhost:8080/", "height": 285}
X = np.random.rand(50,1) * 2 - 1
y = function(X)
inputs = X.shape[1]
y = y.reshape(-1, 1)
plt.plot(X, y, 'o', color='black')
# + id="z05ZyUrmU7is" outputId="5bdfae48-87a3-431c-90b6-7940269b19ab" colab={"base_uri": "https://localhost:8080/", "height": 34}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
num_train = X_train.shape[0]
X_train[:2], y_train[:2]
num_train
# + id="QLshQolGVUlz"
torch.set_default_dtype(torch.float64)
net = nn.Sequential(
nn.Linear(inputs, 50, bias = True), nn.ReLU(),
nn.Linear(50, 50, bias = True), nn.ReLU(),
nn.Linear(50, 50, bias = True), nn.Sigmoid(),
nn.Linear(50, 1)
)
loss_fn = nn.MSELoss()
optimizer = torch.optim.Adam(net.parameters(), lr = .001)
# + id="ZhR9h8Y3WCWM"
num_epochs = 20000
y_train_t = torch.from_numpy(y_train).clone().reshape(-1, 1)
x_train_t = torch.from_numpy(X_train).clone()
y_test_t = torch.from_numpy(y_test).clone().reshape(-1, 1)
x_test_t = torch.from_numpy(X_test).clone()
history = []
# + id="Ef62jrN3WP0d" outputId="5e888732-2bdb-4376-da9f-701f592a0f71" colab={"base_uri": "https://localhost:8080/", "height": 173}
for i in range(num_epochs):
y_pred = net(x_train_t)
loss = loss_fn(y_train_t,y_pred)
history.append(loss.data)
loss.backward()
optimizer.step()
optimizer.zero_grad()
test_loss = loss_fn(y_test_t,net(x_test_t))
if i > 0 and i % 2000 == 0:
print(f'Epoch {i}, loss = {loss:.9f}, test loss {test_loss:.9f}')
# + id="IifIajH-XFrl" outputId="80eac69b-0db2-47ac-ebc0-110ab5f5b0c7" colab={"base_uri": "https://localhost:8080/", "height": 283}
plt.plot(history)
# + id="ZJXQixjLhDkt" outputId="ae28d9c3-acf0-4d41-de95-392e30100b04" colab={"base_uri": "https://localhost:8080/", "height": 285}
X_a = torch.rand(25,1).clone() * 2 - 1
y_a = net(X_a)
y_a = y_a.detach().numpy()
plt.plot(X_a, y_a, 'o', color='black')
| GEN_3_variation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Combine district CSVs into a timeseries file
# %load_ext lab_black
import pandas as pd
import glob
import os
import json
# ### Get all the district json files
# + active=""
#
# files = glob.glob(os.path.join(path, "data/districts-2021*.csv"))
# + active=""
# charter_files = glob.glob(os.path.join(path, "data/charters-2021*.csv"))
# -
def concat_output(name):
path = ""
# get files
files = glob.glob(os.path.join(path, f"data/{name}-2021*.csv"))
# create date field
file_df = (
pd.read_csv(f, low_memory=False).assign(date=os.path.basename(f)) for f in files
)
# concat
concatenated_df = pd.concat(
file_df,
ignore_index=True,
)
# create date column
concatenated_df["date"] = (
concatenated_df["date"]
.str.replace(f"{name}-", "", regex=False)
.str.replace(".csv", "", regex=False)
)
# export
concatenated_df.to_csv(f"data/{name}-timeseries.csv", index=False)
concat_output("districts")
concat_output("charters")
concat_output("private")
# + active=""
# ### Read them and create a date field
# + active=""
# file_df = (
# pd.read_csv(f, low_memory=False).assign(date=os.path.basename(f)) for f in files
# )
# + active=""
# charter_df = (
# pd.read_csv(f, low_memory=False).assign(date=os.path.basename(f))
# for f in charter_files
# )
# + active=""
# ### Concatenate all the files
# + active=""
# concatenated_df = pd.concat(
# file_df,
# ignore_index=True,
# )
# + active=""
# concatenated_charters_df = pd.concat(
# charter_df,
# ignore_index=True,
# )
# + active=""
# ### Clean up the date field and column names
# + active=""
# concatenated_df["date"] = (
# concatenated_df["date"]
# .str.replace("districts-", "", regex=False)
# .str.replace(".csv", "", regex=False)
# )
# + active=""
# concatenated_df.columns = (
# concatenated_df.columns.str.replace(".", "", regex=False)
# .str.lower()
# .str.replace("attributes", "", regex=False)
# )
# + active=""
# concatenated_charters_df["date"] = (
# concatenated_charters_df["date"]
# .str.replace("charters-", "", regex=False)
# .str.replace(".csv", "", regex=False)
# )
# + active=""
# concatenated_charters_df.columns = (
# concatenated_charters_df.columns.str.replace(".", "", regex=False)
# .str.lower()
# .str.replace("attributes", "", regex=False)
# )
# + active=""
# ### Cleaning up the unix dates
# + active=""
# district_dates = [
# "creationdate",
# "editdate",
# "nearterm_reopening_date",
# "elem_reopen_dt",
# "sec_reopen_dt",
# ]
# + active=""
# charter_dates = [
# "creationdate",
# "editdate",
# "reopening_date",
# ]
# + active=""
# def parse_dates(df, dates):
# for d in dates:
# df[f"{d}"] = pd.to_datetime(df[f"{d}"], unit="ms")
# df[f"{d}_clean"] = df[f"{d}"].dt.date
# return df
# + active=""
# df = parse_dates(concatenated_df, district_dates)
# + active=""
# charters_final = parse_dates(concatenated_charters_df, charter_dates)
# + active=""
# ---
# + active=""
# ## Export
# + active=""
# df.to_csv("data/districts-all-timeseries.csv", index=False)
# + active=""
# charters_final.to_csv("data/charters-all-timeseries.csv", index=False)
| school-reopenings/concatenate_districts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import pandas as pd
import pickle
import time
import itertools
import matplotlib
matplotlib.rcParams.update({'font.size': 17.5})
import matplotlib.pyplot as plt
# %matplotlib inline
import sys
import os.path
sys.path.append( os.path.abspath(os.path.join( os.path.dirname('..') , os.path.pardir )) )
# -
from FLAMEdb import *
from FLAMEbit import *
from data_generation import *
# +
# data generation, set exponential to be True or False for exponential decay and power-law decay respectively
d = data_generation_gradual_decrease( 15000 , 15000 , 20, exponential=False )
df = d[0]
holdout,_ = data_generation_gradual_decrease( 15000 , 15000, 20, exponential=False )
res = run_bit(df, holdout, range(20), [2]*20, tradeoff_param = 0.1)
# -
def bubble_plot(res):
sizes = []
effects = []
for i in range(min(len(res),21)):
r = res[i]
if (r is None):
effects.append([0])
sizes.append([0])
continue
effects.append(list( r['effect'] ) )
sizes.append(list(r['size'] ) )
return sizes, effects
# +
# plot the CATE on each level, figure 3
ss, es = bubble_plot(res[1])
plt.figure(figsize=(5,5))
for i in range(len(ss)):
plt.scatter([i]*len(es[i]), es[i], s = ss[i], alpha = 0.6 , color = 'blue')
plt.xticks(range(len(ss)), [str(20-i) if i%5==0 else '' for i in range(20) ] )
plt.ylabel('estimated treatment effect')
plt.xlabel('number of covariates remaining')
#plt.ylim([8,14])
plt.tight_layout()
#plt.savefig('tradeoff01.png', dpi = 300)
# -
| experiments/plot_importance_decay.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Lab 5: Higgs Classification
#
# #### The Large Hadron Collider
#
# The Large Hadron Collider (LHC) is used in experiments where particles are accelerated to relativistic speeds and then collided. Upon collision, atomic particles release their subatomic components. One of these subatomic particles is the Higgs boson, which is created when the energy released by the collision is sufficiently large. However, due to large background contamination of other quantum chromodynamic particles (QCD), a direct 5-sigma observation of Higgs boson jets has not yet been accomplished. In order to improve our ability to detect Higgs boson jets, we will use a series of training datasets which mimic the data output of the Large Hadron Collider to optimize a discriminator and select events to enhance discovery sensitivity.
#
# #### How the LHC Differentiates Particles
#
# The LHC has four sections of detectors. The tracking chamber detects charged particles, the electromagnetic calorimeter interacts with protons and electrons, the hadronic calorimeter interacts with hadrons like pions and protons, and the muon chamber is reserved for particles which pass through all other detectors. The two calorimeters essentially filter out high energy particles and only allow relatively low energy particles to pass through. By counting the particles in these decay chains, the total energy of the original particle can be uncovered.
#
# #### What is a Jet?
#
# A jet is a collection of particles that go towards the same direction in the detector. Intuitively, they come from 'showering' and the fragmentation of a primordial hard quark or gluon. In practice, jets must be defined from experimental observables, such as the 4-momenta of observed particles in the event.
#
# When particles are recorded by the calorimeters, they create detection clusters which depend on their original quark or gluon. In order to determine the original quark or gluon these detections came from, we must group them together. These jets can then be given characterstic values like mass, angular information, transverse momentum, and substructure variables which describe the shape of the jet.
#
# #### Sub-Jetting
#
# Sometimes a jet can split into multiple distinct jets corresponding to seperate cluons or quarks. This is caused by the decay of a boson which creates highly collimated particles which in turn create seperate jets that are close to one another. We can identify this phenomenon numerically by grouping a single jet into multiple sub-jets and finding the average distance to each sub-jet from the detector. By increasing the number of sub-jets, the average distance will always decrease, so by taking the ratio of the average distance for N sub-jets and the average distance for n-1 sub-jets, the number of independant sub-jets can be found.
#
# #### Understanding the Data
#
# | pt | eta | phi | mass | ee2 | ee3 | d2 | angularity | t1 | t2 | t3 | t21 | t32 | KtDeltaR |
# |-----|-----|-----|------|-----|-----|-----|------------|-----|-----|-----|-----|-----|----------|
# | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
#
#
# -pt: The transverse momentum of a jet
# -eta: Rapidity, which is a function of angle theta which goes to plus or minus infinity along the beam axis
# -phi: The Azimuthal angle of the jet
# -mass: The total mass of the jet
# -ee2: A 2 point correlation function measuring the energy of the jet.
# -ee3: A 2 point correlation function measuring the energy of the jet.
# -d2: The ratio of ee3:ee2
# -angularity: Describes how symetric the decay is, where a smaller angularity indicates higher symmetry.
# -t1: Average distance from jets with 1 sub-jet
# -t2: Average distance from jets with 2 sub-jets
# -t3: Average distance from jets with 3 sub-jets
# -t21: The ratio of t2:t1
# -t32 The ratio of t3:t2
# -KtDeltaR: The radius of the jets, which measures how clustered or close together the detected jets are.
import pickle
import math
import pandas as pd
import numpy as np
import scipy
from scipy import stats
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (10,5)
# +
qcd_file = open("qcd_100000_pt_1000_1200.pkl", 'rb')
qcd_d = pickle.load(qcd_file)
higgs_file = open("higgs_100000_pt_1000_1200.pkl", 'rb')
higgs_d = pickle.load(higgs_file)
features = [key for key in qcd.keys()]
print(features)
# -
# Our first step will be to determine which features provide a high discrimination potential between the theoretical Higgs boson jet and background QCD contamination. In order to discover this, we will plot the two via comparative histograms for each feature.
for i in range(0,14):
plt.hist(qcd[features[i]], density=True, bins=100, alpha=0.7, label='QCD')
plt.hist(higgs[features[i]], density=True, bins=100, alpha=0.7, label='Higgs')
plt.xlabel(features[i])
plt.show()
# The histograms above compare each feature of the Higgs jets vs. the feature of the QCD jets. By visual inspection, it appears that mass, ee2, ee3, d2, angularity, t2, t3, t21, and KtDeltaR provide strong discrimination power between signal and background. By comparison, pt, eta, and t1 provide weak discrimination power, and phi and t32 provide no discrimination power.
#
# To discover the correlations among the features, we will create a correlation table, and then summarize it to identify the most highly correlated features.
ctable = qcd_d.corr()
ctable
a = (ctable-np.eye(len(ctable))).max()
b = (ctable-np.eye(len(ctable))).idxmax()
pd.DataFrame([a,b]).T
# As we can see, mass and ee2, t2 and t3, and t21 and t2 are all highly correlated.
#
# In order to determine the signifcance of the deviation in the Higgs observations vs. the background, we will use CLT to find the population mean and standard deviation for each.
# +
samplesize = 2000
samples = 200
qcdSamples = {}
higgsSamples = {}
bgMean = {}
bgStd = {}
for feature in features:
for s in range(0,samples):
qcdsample = np.random.choice(qcd[feature], samplesize, replace=True)
higgssample = np.random.choice(higgs[feature], samplesize, replace=True)
if feature not in qcdSamples:
qcdSamples[feature] = {"mean": [np.mean(qcdsample)], "std": [np.std(qcdsample)]}
higgsSamples[feature] = {"mean": [np.mean(higgssample)], "std": [np.std(higgssample)]}
else:
qcdSamples[feature]["mean"].append(np.mean(qcdsample))
qcdSamples[feature]["std"].append(np.std(qcdsample))
qcdSamples[feature]["mean"].append(np.mean(higgssample))
qcdSamples[feature]["std"].append(np.std(higgssample))
bgMean[feature] = np.mean(qcdSamples[feature]["mean"])
bgStd[feature] = np.std(qcdSamples[feature]["std"])/sqrt(samplesize)
# -
sigmas = {}
for feature in features:
p = stats.norm.cdf(higgsSamples[feature]["mean"], loc=bgMean[feature], scale =bgStd[feature])
# print(feature , "p = ", p)
sigmas[feature] = stats.norm.ppf(p)
print(feature , "s = ", stats.norm.ppf(p))
# Discovery Sensitivities
# It seems that our best strategy to discriminate between QCD particles and the Higgs boson will be to examine the features t2 and t21, which have the highest sensitivity by far compared to the other features. In order to use these to better differentiate the two types of particles, we may want to focus on detecting a subset of all detections which contains the average t2 value for a Higgs boson jet (approx 0.2) but excludes the average value for the background (approx 0.6).
| Lab-5-Higgs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Navigating TTFs via FontTools#
#
# by [<NAME>](https://www.lynneyun.com)
#
# If you've ever tried to parse font files like TTFs, you'll know that it's no simple task. However, there is a powerful python module called FontTools that can help you! In this mini-tutorial, I'll go over how to use some functions of `fontTools`, and eventually illustrate what we can do with them via `drawBot`. This post was inspired by [Allison Parrish](https://www.decontextualize.com/)'s Notebook on [Manipulating Font Data](https://github.com/aparrish/material-of-language/blob/master/manipulating-font-data.ipynb), so check that out if you want more context!
# This first cell demonstrates installing drawBot and fonttools if you don't have them already. Uncomment and install if necessary:
# +
# # !pip install git+https://github.com/typemytype/drawbot
# # !pip install fonttools
# -
# ## fontTools ##
# What is fontTools you say? You can check out the library here: [fontTools Python library](https://rsms.me/fonttools-docs/). For our first example, let's try to grab basic information, such as the GlyphID. I'm importing an open-source typeface here, called Noto Sans.
#
# ### Calling the GlyphID ###
from fontTools.ttLib import TTFont
font = TTFont("./NotoSans-Regular.ttf")
print('glyphID is: ' + str(font.getGlyphID('a')))
# Now, that's the `glyphID`, which is not the Unicode Codepoint. Glyph IDs are the order in which the glyphs have been arranged in a font file, and may not be the same across different fonts.
#
# ### Calling the Unicode Codepoint ###
# In case you want to grab the unicode codepoint represented as an integer, you can use `ord`.
print(ord('a'))
# ### Calling the Glyph Width ###
# In font files, there are 'widths' of each glyph. They include the left and right sidebearings. Access the advances with the font object's `.width` attribute. You will have to call `getGlyphSet()` in order to do so.
glyph = font.getGlyphSet()['a']
print('width of glyph is ' + str(glyph.width))
# ### Units Per Em Value ###
#
# You can also get the `unitsPerEm` attribute of the font object from the [`ttLib` package](https://rsms.me/fonttools-docs/ttLib/index.html?highlight=unitsperem).
units_per_em = font['head'].unitsPerEm
print('units per em is: ' + str(units_per_em))
# ### Glyph Names ###
#
# Keep in mind that `Glyph Names` are how a certain glyph is named, even if that's not how the character itself is represented. For example, when you want to get information about the `"ã"` chracter, you'll have to point to it as `"atilde"`.
# You can also get all the Glyph Names in a font file, using `getGlyphNames()`. Using this, we can also get how many characters are in this font file:
len(font.getGlyphNames())
# Let's get the first 10 glyph names to see what this list looks like.
font.getGlyphNames()[:10]
# ### Mapping Unicode Values to Glyph Names ###
# So considering what we just learned about `Glyph Names` you might be scratching your head about how to convert unicode values to glyph names, and vice versa. Thanks to [<NAME>](https://twitter.com/justvanrossum) for letting me know about this super useful feature!
#
# Here's a handy method for doing just that! Remember in an earlier example we got a unicode value using `ord`? This will return a dict that maps unicode to Glyph Names.
font.getBestCmap()
# So, using this, here is an example of converting `ã` to a unicode value and getting the `Glyphname` from it:
font.getBestCmap()[ord('ã')]
# ### Getting Glyph Outlines via RecordingPen ###
# Let's try to do something a bit more interesting, perhaps — let's grab all the points from the glyph using a `RecordingPen`.
from fontTools.pens.recordingPen import RecordingPen
glyph = font.getGlyphSet()['a']
p = RecordingPen()
glyph.draw(p)
p.value
# If you're not sure about what `moveTo`, `qCurveTo`, `lineTo` is doing refer to the [DrawBot BezierPath Documentation](https://www.drawbot.com/content/shapes/bezierPath.html).
#
# I'm going to make a function to make grabbing the curve information easier:
def get_outline(font, ch):
glyph = font.getGlyphSet()[ch]
p = RecordingPen()
glyph.draw(p)
return p.value
# you can see that now this one line is all we need:
get_outline(font, "a")
# If you are playing with the above code though, you'll notice that glyphs that are made out of composites like the 'atilde' give us components, not the actual outlines:
get_outline(font, "atilde")
# ### Decomposing Recording Pen ###
#
# To grab these, we will need the `Decomposing Pen` to decompose them. Here is the modified function from above to grab all outlines. Now trying to grab `atilde` should work!
# +
from fontTools.pens.recordingPen import DecomposingRecordingPen
def get_all_outlines(font, ch):
glyphset = font.getGlyphSet()
glyph = glyphset[ch]
p = DecomposingRecordingPen(glyphset)
glyph.draw(p)
return p.value
# print(get_all_outlines(font, "atilde"))
# -
# ## Let's do fun things with Drawbot! ##
#
# Okay, what can we do with all this information besides just plain information, you say? Let's do some fun things, illustrated with [DrawBot](https://drawbot.com). I'm going to use `Drawbot` as a python module here, so let's import it.
import drawBot as draw
# I'm using drawbot as a module here, so I need to set it up so I can see it on my Jupyter Notebook. I'm setting a `startdraw` function and `show` so I can quickly call it while I'm drawing things, since drawbot needs the same lines to start and end a drawing.
#
# This incorporates the IPython module, so you may need to install it if you don't already have it.
# +
from IPython.display import Image, display
def startdraw(canvas_width,canvas_height):
draw.newDrawing()
draw.newPage(canvas_width, canvas_height)
def show():
draw.saveImage("drawBotImage.png")
draw.endDrawing()
drawing = Image(filename = "drawBotImage.png")
display(drawing)
# -
# ### Drawing fontTools Glyphs in Drawbot ###
# Thanks to [@Drawbotapp](https://twitter.com/drawbotapp), I learned that a `BezierPath()` is also a pen! It makes it super easy to draw the fontTools glyphs.
# +
def draw_outline(font, ch, scale_num):
path = draw.BezierPath()
glyph = font.getGlyphSet()[ch]
glyph.draw(path)
path.scale(scale_num)
return path
startdraw(200,200)
draw.drawPath(draw_outline(font,"a",0.3))
show()
# -
# ### Modifying Outline Paths ###
# Using the `get_outline` function we made before, let's draw the letter 'a'. I'm parsing information from `get_outline`, making a `BezierPath` object, and adding all the paths in there. Take a look at the [Drawbot Documentation for BezierPath](http://www.drawbot.com/content/shapes/bezierPath.html) if you need a refresher. You'll notice that I'm using the `.scale` to reduce the size before doing the line too.
# +
path_orig = get_outline(font, "a")
startdraw(200,200)
paths = []
for i in path_orig:
if i[0] == 'moveTo':
path = draw.BezierPath()
path.moveTo((i[1][0]))
if i[0] == 'lineTo':
path.lineTo((i[1][0]))
if i[0] == 'qCurveTo':
path.qCurveTo(*(i[1]))
if i[0] == 'closePath':
path.closePath()
paths.append(path)
finalpath = paths[0].difference(paths[1])
finalpath.scale(0.3) #let's scale it down!
draw.fill(0.5)
draw.stroke(0)
draw.drawPath(finalpath)
show()
# -
# Ta-da! We made just one single letter, no big deal, you might say. Let's start manipulating all the points to give it a distorted filter:
import random
path_orig = get_outline(font, "a")
startdraw(300,300)
paths = []
for i in path_orig:
disturbance = random.randint(-35, 35)
if i[0] == 'moveTo':
x = i[1][0][0] + disturbance
y = i[1][0][1] + disturbance
path = draw.BezierPath()
path.moveTo((x,y))
if i[0] == 'lineTo':
x = i[1][0][0] + disturbance
y = i[1][0][1]+ disturbance
path.lineTo((x,y))
if i[0] == 'qCurveTo':
temp = []
for cord in i[1]:
x = cord[0] + disturbance
y = cord[1] + disturbance
temp.append((x,y))
path.qCurveTo(*temp)
if i[0] == 'closePath':
path.closePath()
paths.append(path)
finalpath = paths[0].difference(paths[1])
finalpath.scale(0.3) #let's scale it down!
draw.fill(0.5)
draw.stroke(0)
draw.drawPath(finalpath)
show()
# ## Parting words ##
# See, lots of fun things! :-) That's it from me (at least for now.)
#
# Check out the wiki Page on Github for fontTools if you would like to dig deeper:
# [Wiki Page on Github for FontTools](https://github.com/fonttools/fonttools/wiki)
#
# Wish you the best of luck on your font-navigating journey!
| FontTools & DrawBot/.ipynb_checkpoints/Navigating TTFs with fontTools-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Data" data-toc-modified-id="Data-1"><span class="toc-item-num">1 </span>Data</a></span><ul class="toc-item"><li><span><a href="#Load-genotype" data-toc-modified-id="Load-genotype-1.1"><span class="toc-item-num">1.1 </span>Load genotype</a></span></li><li><span><a href="#Load-Pedigree" data-toc-modified-id="Load-Pedigree-1.2"><span class="toc-item-num">1.2 </span>Load Pedigree</a></span></li><li><span><a href="#Training-dataset" data-toc-modified-id="Training-dataset-1.3"><span class="toc-item-num">1.3 </span>Training dataset</a></span></li></ul></li><li><span><a href="#Build-GAN" data-toc-modified-id="Build-GAN-2"><span class="toc-item-num">2 </span>Build GAN</a></span><ul class="toc-item"><li><span><a href="#Generator" data-toc-modified-id="Generator-2.1"><span class="toc-item-num">2.1 </span>Generator</a></span></li><li><span><a href="#Discriminator" data-toc-modified-id="Discriminator-2.2"><span class="toc-item-num">2.2 </span>Discriminator</a></span></li><li><span><a href="#Loss" data-toc-modified-id="Loss-2.3"><span class="toc-item-num">2.3 </span>Loss</a></span></li><li><span><a href="#Optimizer" data-toc-modified-id="Optimizer-2.4"><span class="toc-item-num">2.4 </span>Optimizer</a></span></li></ul></li><li><span><a href="#Training" data-toc-modified-id="Training-3"><span class="toc-item-num">3 </span>Training</a></span></li></ul></div>
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="7heGNeSW4Lta" outputId="342a7962-441c-43fe-fe51-0c38fbc15530"
import os
import glob
import time
import PIL
import imageio
import pickle
from IPython import display
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
import matplotlib.cm as cm
from matplotlib.colors import ListedColormap
from sklearn.preprocessing import MinMaxScaler
from sklearn.utils import resample
import tensorflow as tf
from tensorflow.keras import datasets, layers, Sequential, Model
from tensorflow.keras.utils import to_categorical
from tensorflow.keras import initializers, activations, regularizers, constraints
from tensorflow.keras.layers import InputSpec
import tensorflow.keras.backend as K
tf.__version__
# +
# Set environment variables
os.environ['CUDA_VISIBLE_DEVICES'] = '6'
# On DGX-1 you might need to set the following environment variable for maximum performance:
os.environ['NCCL_TOPOLOGY'] = 'CUBEMESH'
# -
gpus = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(gpus[0], True)
# + [markdown] colab_type="text" id="DwON8HAsxVDl"
# # Data
# + [markdown] colab_type="text" id="tTu9kSqI4Ltz"
# ## Load genotype
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="_2oS9Ve9026g" outputId="d884dee1-1b73-496f-e8d3-f41d330aef5f"
# load data
vcf_file = 'data/HLA.recode.vcf'
# get header
with open(vcf_file,'r') as f_in:
# skip info
for line_num in range(252):
f_in.readline()
line=f_in.readline()
# print(line.strip().split('\t'))
# load genotype
genotype = pd.read_csv(vcf_file, comment='#', sep='\t', names=line.strip().split('\t'))
print('genotype_file shape:', genotype.shape)
# -
X_train = genotype.iloc[:, 9:].replace({
'0|0': 0,
'0|1': 1,
'1|0': 2,
'1|1': 3
}).transpose()
X_train.head()
# + [markdown] colab_type="text" id="UL-Yi-Tz4Lt9"
# ## Load Pedigree
# + colab={} colab_type="code" id="ZUZjS-eI4Lt-"
ped_file = 'data/integrated_call_samples.20130502.ALL.ped'
pedigree = pd.read_csv(ped_file, sep='\t', index_col='Individual ID')
# + colab={} colab_type="code" id="dwtwBB0T4LuA"
pedigree['Super Population'] = pedigree['Population']
pedigree['Super Population'] = pedigree['Super Population'].replace({
'YRI': 'AFR', 'CEU': 'EUR', 'GWD': 'AFR', 'ESN': 'AFR',
'CHS': 'EAS', 'IBS': 'EUR', 'PJL': 'SAS', 'PUR': 'AMR',
'CLM': 'AMR', 'BEB': 'SAS', 'CHB': 'EAS', 'PEL': 'AMR',
'STU': 'SAS', 'MSL': 'AFR', 'JPT': 'EAS', 'KHV': 'EAS',
'ACB': 'AMR', 'LWK': 'AFR', 'ITU': 'SAS', 'GIH': 'SAS',
'ASW': 'AMR', 'TSI': 'EUR', 'CDX': 'EAS', 'CHD': 'EAS',
'GBR': 'EUR', 'MXL': 'AMR', 'FIN': 'EUR' })
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="PVSPnAXO4LuC" outputId="cec4f008-96cf-439c-fef7-09633ba8e824"
pedigree['Super Population'] = pedigree['Super Population'].replace({
'EAS': 0,
'EUR': 1,
'AFR': 2,
'AMR': 3,
'SAS': 4
})
# -
# ## Training dataset
# only use EAS, EUR and AFR population
Y_train = pedigree.loc[genotype.iloc[:, 9:].columns, :]['Super Population']
Y_train
X_train = X_train.loc[Y_train.index, :7159]
print(X_train.shape)
X_train.head()
X_train = to_categorical(X_train)
X_train.shape
dataset = (X_train, Y_train)
# + [markdown] colab_type="text" id="iITgqDhW4LuN"
# # Build GAN
# + colab={} colab_type="code" id="LsfBCC0N4LuO"
noise_dim = 100
genotype_len = X_train.shape[1]
channels = X_train.shape[2]
genotype_shape = (genotype_len, channels)
num_classes = 5
lrelu_alpha = 0.2
# + [markdown] colab_type="text" id="QNl6b_qn4LuQ"
# ## Generator
# + colab={} colab_type="code" id="YbIht4ck4LuS"
def make_generator_model():
# input
noise = layers.Input(shape=(noise_dim, ))
label = layers.Input(shape=(1, ))
label_embedding = layers.Flatten()(layers.Embedding(num_classes, noise_dim)(label))
model_input = layers.multiply([noise, label_embedding])
# network
x = layers.Dense(1790 * 256, use_bias=False, input_shape=(noise_dim, ))(model_input)
x = layers.Reshape((1790, 256))(x)
x = layers.BatchNormalization(epsilon=2e-5, momentum=9e-1)(x)
x = layers.LeakyReLU()(x)
x = layers.Conv1D(256,
7,
strides=1,
padding='same',
use_bias=False,
kernel_initializer='glorot_uniform')(x)
x = layers.UpSampling1D(2)(x)
x = layers.BatchNormalization(epsilon=2e-5, momentum=9e-1)(x)
x = layers.LeakyReLU()(x)
x = layers.Conv1D(128,
5,
strides=1,
padding='same',
use_bias=False,
kernel_initializer='glorot_uniform')(x)
x = layers.UpSampling1D(2)(x)
x = layers.BatchNormalization(epsilon=2e-5, momentum=9e-1)(x)
x = layers.LeakyReLU()(x)
x = layers.Conv1D(channels,
5,
strides=1,
padding='same',
activation='softmax')(x)
return Model([noise, label], x)
# + colab={"base_uri": "https://localhost:8080/", "height": 156} colab_type="code" id="C0VKq2GE4LuW" outputId="5ffe6d1b-96d5-4b68-8795-7975850de2d0"
generator = make_generator_model()
noise = tf.random.normal([1, 100])
label = np.random.randint(0, 3, (1, ))
generated_genotype = generator([noise, label], training=False)
plt.imshow(np.argmax(generated_genotype, axis=2).reshape(40, 179),
cmap=ListedColormap(["red", "blue", "yellow", "green"]))
plt.title('label: {}'.format(label))
# -
generator.summary()
# + [markdown] colab_type="text" id="TzyrknUx4LuZ"
# ## Discriminator
# + colab={} colab_type="code" id="7lsQ-3p9hoCS"
class ClipConstraint(constraints.Constraint):
# set clip value when initialized
def __init__(self, clip_value):
self.clip_value = clip_value
# clip model weights to hypercube
def __call__(self, weights):
return K.clip(weights, -self.clip_value, self.clip_value)
# get the config
def get_config(self):
return {'clip_value': self.clip_value}
const = ClipConstraint(0.01)
# + colab={} colab_type="code" id="P38rAwEc4LuZ"
def make_discriminator_model():
genotype = layers.Input(shape=genotype_shape)
label = layers.Input(shape=(1, ), dtype='int32')
flat_genotype = layers.Flatten()(genotype)
label_embedding = layers.Flatten()(layers.Embedding(num_classes, genotype_len * channels)(label))
model_input = layers.multiply([flat_genotype, label_embedding])
model_input = layers.Reshape((genotype_len, channels))(model_input)
x = layers.LeakyReLU()(model_input)
x = layers.Dropout(0.3)(x)
x = layers.Conv1D(64,
5,
strides=2,
padding='same',
kernel_constraint=const,
kernel_initializer='glorot_uniform')(x)
x = layers.LeakyReLU()(x)
x = layers.Dropout(0.3)(x)
x = layers.Conv1D(128,
7,
strides=2,
padding='same',
kernel_constraint=const,
kernel_initializer='glorot_uniform')(x)
x = layers.LeakyReLU()(x)
x = layers.Dropout(0.3)(x)
x = layers.Flatten()(x)
x = layers.Dense(512, kernel_initializer='glorot_uniform')(x)
x = layers.LeakyReLU()(x)
x = layers.Dense(1, kernel_initializer='glorot_uniform')(x)
return Model([genotype, label], x)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="f1JhXwQR4Lub" outputId="f704bfe0-cb76-425a-db5c-d596463b7b91"
# Use the (as yet untrained) discriminator
# to classify the generated images as real or fake.
discriminator = make_discriminator_model()
decision = discriminator([generated_genotype, label])
print(decision)
# -
discriminator.summary()
# + [markdown] colab_type="text" id="JPkRMOdi4Luc"
# ## Loss
# +
# wasserstein_loss
def generator_loss(fake_output):
return -tf.reduce_mean(fake_output)
def discriminator_loss(real_output, fake_output):
return -tf.reduce_mean(real_output) + tf.reduce_mean(fake_output)
# +
# # hinge_loss
# hinge_loss = tf.keras.losses.Hinge()
# def discriminator_loss(real_output, fake_output):
# real_loss = hinge_loss(tf.ones_like(real_output), real_output)
# fake_loss = hinge_loss(tf.math.negative(tf.ones_like(fake_output)),
# fake_output)
# total_loss = real_loss + fake_loss
# return total_loss
# def generator_loss(fake_output):
# return hinge_loss(tf.ones_like(fake_output), fake_output)
# + [markdown] colab_type="text" id="Y0ZSxcba4Lui"
# ## Optimizer
# + colab={} colab_type="code" id="tlSE1u3f4Lui"
# RMSprop optimizer
generator_optimizer = tf.keras.optimizers.RMSprop(5e-5)
discriminator_optimizer=tf.keras.optimizers.RMSprop(5e-5)
# +
# # Adam optimizer
# BETA_1 = 0.5
# BETA_2 = 0.9
# generator_optimizer = tf.keras.optimizers.Adam(1e-4,
# beta_1=BETA_1,
# beta_2=BETA_2,
# amsgrad=True)
# discriminator_optimizer = tf.keras.optimizers.Adam(1e-4,
# beta_1=BETA_1,
# beta_2=BETA_2,
# amsgrad=True)
# + [markdown] colab_type="text" id="vUWXkWh_4Luk"
# # Training
# + colab={} colab_type="code" id="Cxz_HIJt4Lun"
num_examples_to_generate = num_classes * 2
# We will reuse this seed overtime (so it's easier)
# to visualize progress in the animated GIF)
noise = tf.random.normal([num_examples_to_generate, noise_dim])
label = np.array([0, 0, 1, 1, 2, 2, 3, 3, 4, 4])
seed = [noise, label]
# + colab={} colab_type="code" id="wTD1ZtuD4Lup"
# larger value to make a better discriminator in training process.
num_repeat_disc = 4
# Notice the use of `tf.function`
# This annotation causes the function to be "compiled".
@tf.function
def train_step(batch_size, genotypes, labels):
# train disc
for _ in range(num_repeat_disc):
noise = tf.random.normal([batch_size, noise_dim])
with tf.GradientTape() as disc_tape:
generated_genotypes = generator([noise, labels], training=False)
real_output = discriminator([genotypes, labels], training=True)
fake_output = discriminator([generated_genotypes, labels], training=True)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
# train both
noise = tf.random.normal([batch_size, noise_dim])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_genotypes = generator([noise, labels], training=True)
real_output = discriminator([genotypes, labels], training=True)
fake_output = discriminator([generated_genotypes, labels], training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
return gen_loss, disc_loss
# + colab={} colab_type="code" id="XqPV9xsJ4Luq"
checkpoint_dir = ".training_checkpoints/HLA/PG-cGAN"
checkpoint_prefix = os.path.join(checkpoint_dir, "wgan")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
# + colab={} colab_type="code" id="6WcnFaei4Lus"
def generate_and_save_images(model, epoch, test_input):
# Notice `training` is set to False.
# This is so all layers run in inference mode (batchnorm).
predictions = model(test_input, training=False)
fig = plt.figure(figsize=(8, 12))
for i in range(predictions.shape[0]):
plt.subplot(5, 2, i + 1)
plt.imshow(np.argmax(predictions[i], axis=1).reshape(40, 179),
ListedColormap(["red", "blue", "yellow", "green"]))
plt.title('label: {}'.format(test_input[1][i]))
plt.axis('off')
fig.tight_layout(h_pad=-35)
plt.show()
# + colab={} colab_type="code" id="A-v3TqYs4Lut"
def train(dataset, epochs, batch_size):
gen_loss_log, disc_loss_log = [], []
for epoch in range(epochs):
start = time.time()
# shuffle and batch the data
train_dataset = tf.data.Dataset.from_tensor_slices(dataset)\
.shuffle(X_train.shape[0])\
.batch(batch_size, drop_remainder=True)
for genotype_batch, label_batch in train_dataset:
gen_loss, disc_loss = train_step(batch_size, genotype_batch, label_batch)
# append loss of last batch
gen_loss_log.append(gen_loss.numpy())
disc_loss_log.append(disc_loss.numpy())
# Produce images for the GIF as we go
if (epoch + 1) % 10 == 0:
display.clear_output(wait=True)
generate_and_save_images(generator, epoch + 1, seed)
# Save the model every 50 epochs
if (epoch + 1) % 50 == 0:
checkpoint.save(file_prefix=checkpoint_prefix)
# print loss of last batch
print('gen_loss: {}, disc_loss: {}'.format(gen_loss.numpy(), disc_loss.numpy()))
print('Time for epoch {} is {} sec'.format(epoch + 1, time.time() - start))
return gen_loss_log, disc_loss_log
# + colab={"base_uri": "https://localhost:8080/", "height": 949} colab_type="code" id="-Vl8G0lP4Luu" outputId="e9a04e45-84ea-4176-9a80-d80fe8591072"
EPOCHS = 50
# EPOCHS = 5000
gen_loss_log, disc_loss_log = train(dataset, epochs=EPOCHS, batch_size=64)
# + colab={"base_uri": "https://localhost:8080/", "height": 265} colab_type="code" id="DoCcCVCt4Luw" outputId="0275b936-b515-42a9-ff2f-62eae73f33c9"
plt.plot(range(EPOCHS), gen_loss_log, 'r')
plt.plot(range(EPOCHS), disc_loss_log, 'b')
plt.show()
# -
| PG-cGAN code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import requests
import re
from datetime import date
from datetime import datetime, timedelta
import json
import locale
import csv
import pandas as pd
from csv import DictWriter
from collections import defaultdict
front_page = 'https://www.estrepublicain.fr/sante/coronavirus'
def fetch_links(source_website):
"""this function fetches links from the front page
and returns a list"""
list_of_links = []
#pagination on the website follows a simple pattern
#'https://www.estrepublicain.fr/sante/coronavirus?page=NUMBER_OF_PAGE
#thus, we run a loop
for i in range(101):
url = source_website + '?page=' + str(i)
page = requests.get(url)
data = page.text
soup = BeautifulSoup(data)
#catching bugs if a link doesn't contain a substring
for link in soup.find_all('a'):
try:
if link.get('href').startswith('/sante/2'):
list_of_links.append('https://www.estrepublicain.fr'+link.get('href'))
except AttributeError:
continue
return list_of_links
big_list = fetch_links('https://www.estrepublicain.fr/sante/coronavirus')
big_list[0]
with open('list_of_links.txt', 'w') as f:
for item in big_list:
f.write("%s\n" % item)
# +
def grab_info(link):
"""this function scrapes information
from a single web page, including text, title, and date"""
#headers = { 'User-Agent': 'Mozilla/5.0 (Windows NT 6.0; WOW64; rv:24.0) Gecko/20100101 Firefox/24.0' }
page_single = requests.get(link).text
soup_single = BeautifulSoup(page_single)
#check if the article is not behind a paywall
if soup_single.find_all('div', class_='preview non-paywall2'):
return None
#fetching the date from the link string
date_from_link = link.split('/sante/')[1].split('/')
date_fin = '/'.join(date_from_link[:3])
#putting together all the text from the page
text = []
for i in soup_single.find_all('p', class_=None)[8:]:
text.append(i.get_text())
text = text[1:]
text = ' '.join(text)
text = text
#fetching the title
#two options depending on the page structure
try:
title = soup_single.find('title').get_text()
except:
title = soup_single.find("span",{"class":"headline"}).next_sibling.lstrip()
#getting reference summaries
meta_tag = soup_single.find('meta', attrs={'name':'Description'})
summary = meta_tag['content']
dictionary = {'date': date_fin,
'summary':summary,
'text':text,
'title':title,
'topic':'',
'url': link}
return dictionary
#json.dumps(dictionary, indent = 6, ensure_ascii=False)
# -
grab_info('https://www.estrepublicain.fr/sante/2021/11/17/covid-19-les-contaminations-en-forte-hausse-espoir-autour-du-traitement-de-pfizer')
def fetch_summaries(li_of_links):
d = defaultdict()
for i in li_of_links:
try:
page_single = requests.get(i).text
soup_single = BeautifulSoup(page_single)
meta_tag = soup_single.find('meta', attrs={'name':'Description'})
summary = meta_tag['content']
d[i]=summary
except:
print(i)
pass
return dict(d)
def clean_text(row):
search='Ce contenu est bloqué car vous n\'avez pas accepté les traceurs. En cliquant sur « J’accepte », les traceurs seront déposés et vous pourrez visualiser les contenus . En cliquant sur « J’accepte tous les traceurs », vous autorisez des dépôts de traceurs pour le stockage de vos données sur nos sites et applications à des fins de personnalisation et de ciblage publicitaire.'
if search in str(row):
print('yes')
spl = row.split(search)
new_row = ' '.join(spl)
return new_row
return row
def delete_cas(row):
search = 'CAS N°1 : Vous (ou une autre personne) utilisez plus de $this.View.AuthResponse.DeviceLimit appareils et/ou navigateurs en même temps →\r\n\t\t\t\t\t\t\t\tDéconnectez-vous des appareils et/ou navigateurs que vous n\'utilisez pas CAS N°2 : Vous naviguez en mode privé →\r\n\t\t\t\t\t\t\t\tDéconnectez-vous systématiquement avant de fermer la fenêtre du navigateur CAS N°3 : Vous refusez les cookies de connexion dans les paramètres de votre navigateur (ou une mise à jour a modifié vos paramètres) →\r\n\t\t\t\t\t\t\t\tChanger les paramètres d\'acceptation des cookies de votre navigateur DANS TOUS LES CAS → cliquer sur "continuer sur cet appareil" résout le problème Que se passe-t-il si je clique sur "continuer sur cet appareil" ?\r\n\t\t\t\t\t\t\t\tVous pourrez profiter de votre compte sur cet appareil et tous vos autres appareils\r\n\t\t\t\t\t\t\t\tseront déconnectés. Vous pourrez toujours vous y reconnecter, dans la limite de\r\n\t\t\t\t\t\t\t\t$this.View.AuthResponse.DeviceLimit appareils. Comment puis-je voir les appareils connectés ?\r\n\t\t\t\t\t\t\t\tRendez-vous dans votre espace client puis cliquez sur "gérer les équipements". Si vous y allez après avoir cliqué sur "Continuer sur cet appareil", il ne devrait y en avoir qu\'un seul. Retrouvez tous nos contenus et notre journal en numériqueTéléchargez gratuitement l\'app'
if search in str(row):
return row[len(search):]
return row
def list_of_dic(source):
list_of_dic = []
with open(source, "r") as a_file:
for idx,line in enumerate(a_file):
try:
stripped_line = line.strip()
dic=grab_info(stripped_line)
print(idx)
if dic['text']:
list_of_dic.append(dic)
except ConnectionError:
continue
return list_of_dic
def write_csv(li_of_li, file_name):
"""this function writes a csv file
from a list of lists"""
with open(file_name, "wb") as f:
writer = csv.writer(f)
writer.writerows(li_of_li)
# +
#ensure_ascii=False makes sure that french special characters are read correctly
with open('lest_republicain_summaries.json', 'w') as f:
json.dump(li, f, ensure_ascii=False)
# +
#converting a json to a csv file for convenience
data_file = open('csv_les_republicain_summ.csv', 'w', newline='')
csv_writer = csv.writer(data_file)
with open('lest_republicain_summaries.json') as json_file:
jsondata = json.load(json_file)
count = 0
for data in jsondata:
if count == 0:
header = data.keys()
csv_writer.writerow(header)
count += 1
csv_writer.writerow(data.values())
data_file.close()
# +
#use a set or hashmap
# -
def update_csv(source_link):
links_file = open('list_of_links-Copy1.txt', 'r')
reader = csv.reader(links_file)
allRows = [row[0] for row in reader]
li_of_links = []
for i in range(101):
find = False
url = source_link + '?page=' + str(i)
page = requests.get(url)
data = page.text
soup = BeautifulSoup(data)
#catching bugs if a link doesn't contain a substring
for link in soup.find_all('a'):
if link.get('href')and link.get('href').startswith('/sante/2'):
link_full = 'https://www.estrepublicain.fr'+link.get('href')
if link_full in allRows and link_full != 'https://www.estrepublicain.fr/sante/2020/09/21/coronavirus-suivez-l-evolution-du-taux-d-incidence-dans-le-grand-est-et-en-bourgogne-franche-comte':
print(link_full)
find = True
li_of_links.append(link_full+'\n')
dic = grab_info(link_full)
headersCSV = dic.keys()
with open('csv_les_republicain-Copy1.csv', 'a', newline='') as f_object:
# Pass the CSV file object to the Dictwriter() function
# Result - a DictWriter object
dictwriter_object = DictWriter(f_object, fieldnames=headersCSV)
# Pass the data in the dictionary as an argument into the writerow() function
dictwriter_object.writerow(dic)
# Close the file object
f_object.close()
if find:
break
with open('list_of_links-Copy1.txt','a') as out:
out.writelines(li_of_links)
def get_new(source='https://www.estrepublicain.fr/sante/coronavirus'):
url = source
page = requests.get(url)
data = page.text
soup = BeautifulSoup(data)
#catching bugs if a link doesn't contain a substring
li_of_dict = []
for link in soup.find_all('a'):
date_today = str(datetime.today().strftime('%Y/%m/%d'))
if link.get('href') and link.get('href').startswith('/sante/2'):
link_full = 'https://www.estrepublicain.fr'+link.get('href')
date_from_link = link_full.split('/sante/')[1].split('/')
date_fin = '/'.join(date_from_link[:3])
if date_fin == date_today:
info = grab_info(link_full)
if info:
li_of_dict.append(info)
keys = list(li_of_dict[0].keys())
a_file = open("lest_rep_today.csv", "w")
dict_writer = csv.DictWriter(a_file, keys)
dict_writer.writeheader()
dict_writer.writerows(li_of_dict)
a_file.close()
get_new(source='https://www.estrepublicain.fr/sante/coronavirus')
data = pd.read_csv("lest_rep_today.csv")
data
| data/scraping_est_republicain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MOC computation
# If you would like to try to repeat examples from this notebook, you can download FESOM2 data and mesh. The data are quite heavy, about 15 Gb.
#
# Link: https://swiftbrowser.dkrz.de/public/dkrz_c719fbc3-98ea-446c-8e01-356dac22ed90/PYFESOM2/
#
# You have to download both archives (`LCORE.tar` and `LCORE_MESH.tar`) and extract them.
#
# Alternative would be to use very light weight mesh that comes with pyfesom2 in the `tests/data/pi-grid/` and example data on this mesh in `tests/data/pi-results`.
import pyfesom2 as pf
import matplotlib.cm as cm
import matplotlib.pylab as plt
import numpy as np
mesh = pf.load_mesh('/Users/nkolduno/PYTHON/DATA/LCORE_MESH/',
abg=[50, 15, -90])
data = pf.get_data('../../DATA/LCORE/', 'w', range(1950,1959), mesh, how="mean", compute=True )
data.shape
# Minimum nessesary input is mesh and one 3D field of w.
lats, moc = pf.xmoc_data(mesh, data)
# You get latitides and the MOC values (global by default). They can be used to plot the MOC using hofm_plot function:
plt.figure(figsize=(10, 3))
pf.plot_xyz(mesh, moc, xvals=lats, maxdepth=7000, cmap=cm.seismic, levels = np.linspace(-30, 30, 51),
facecolor='gray')
# Increase the number of latitude bins:
lats, moc = pf.xmoc_data(mesh, data, nlats=200)
plt.figure(figsize=(10, 3))
pf.plot_xyz(mesh, moc, xvals=lats, maxdepth=7000, cmap=cm.seismic, levels = np.linspace(-30, 30, 51),
facecolor='gray')
# You can define the region for MOC computation.
lats, moc = pf.xmoc_data(mesh, data, mask="Atlantic_MOC")
plt.figure(figsize=(10, 3))
pf.plot_xyz(mesh, moc, xvals=lats, maxdepth=7000, cmap=cm.seismic, levels = np.linspace(-30, 30, 51),
facecolor='gray')
# Regions are listed in documentation of pf.get_mask. Currently they are:
#
# Ocean Basins:
# "Atlantic_Basin"
# "Pacific_Basin"
# "Indian_Basin"
# "Arctic_Basin"
# "Southern_Ocean_Basin"
# "Mediterranean_Basin"
# "Global Ocean"
# "Global Ocean 65N to 65S"
# "Global Ocean 15S to 15N"
# MOC Basins:
# "Atlantic_MOC"
# "IndoPacific_MOC"
# "Pacific_MOC"
# "Indian_MOC"
# Nino Regions:
# "Nino 3.4"
# "Nino 3"
# "Nino 4"
# You can combine masks:
mask1 = pf.get_mask(mesh, "Atlantic_Basin")
mask2 = pf.get_mask(mesh, "Arctic_Basin")
mask3 = mask1|mask2
# Check how the masked data looks like:
pf.plot(mesh, data[:,0]*mask3)
# You can then provide combined mask to `xmoc_data`:
lats, moc = pf.xmoc_data(mesh, data, mask=mask3)
plt.figure(figsize=(10, 3))
pf.plot_xyz(mesh, moc, xvals=lats, maxdepth=7000, cmap=cm.seismic, levels = np.linspace(-30, 30, 51),
facecolor='gray')
# If you work with large meshes, it makes sence to compute some data beforehand and provide them to the function:
meshdiag = pf.get_meshdiag(mesh)
el_area = meshdiag['elem_area'][:]
nlevels = meshdiag['nlevels'][:]-1
face_x, face_y = pf.compute_face_coords(mesh)
mask = pf.get_mask(mesh, 'Global Ocean')
lats, moc = pf.xmoc_data(mesh, data, mask = mask,
el_area = el_area, nlevels=nlevels,
face_x=face_x, face_y=face_y)
plt.figure(figsize=(10, 3))
pf.plot_xyz(mesh, moc, xvals=lats, maxdepth=7000, cmap=cm.seismic, levels = np.linspace(-30, 30, 51),
facecolor='gray')
# ## Compute AMOC at 26.5N
# Open data as time series
data = pf.get_data('../../DATA/LCORE/', 'w', range(1950,1959), mesh, how="ori", compute=False )
data
# We have 9 time steps
data.shape
# Compute AMOC for each time step, increase the number of latitudes if you want to be preciselly at 26.5:
# +
moc = []
for i in range(data.shape[0]):
lats, moc_time = pf.xmoc_data(mesh, data[i,:,:], mask='Atlantic_MOC', nlats=361)
moc.append(moc_time)
# -
# Find which index corresponds to 26.5
lats[233]
# Compute maximum at this point for each time step:
amoc26 = []
for i in range(len(moc)):
amoc26.append(moc[i][233,:].max())
amoc26
# You can use time information from original dataset for plotting:
plt.plot(data.time, amoc26)
| newver_202009/notebooks/compute_MOC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.6 64-bit (''anime_python-NngzkPi4'': pipenv)'
# name: python386jvsc74a57bd049c31735898bcc7a7f9475ec39f9101bc3fe8f729d94da83e0f39c0054a8cb73
# ---
# +
import pandas as pd
people = {
"first": ["Corey", "Jane", "John"],
"last": ["Schafer", "Doe", "Doe"],
"email": ["<EMAIL>", "<EMAIL>", "<EMAIL>"]
}
df = pd.DataFrame(people)
df
# -
# Add one new column
df['full_name'] = df['first'] + " " + df['last']
df
# Drop columns, can do inplace operations
df.drop(columns=['first', 'last'], inplace=True)
# Add multiple columns
df[['first', 'last']] = df['full_name'].str.split(' ', expand=True)
df
# Add a single row of data, makes a new DataFrame (no inplace)
df.append({'first': 'Tony'}, ignore_index=True)
new_people = {
'first': ['Tony', 'Steve'],
'last': ['Stark', 'Rogers'],
'email': ['<EMAIL>', '<EMAIL>']
}
df2 = pd.DataFrame(new_people)
df2
# Add dataframes together
df = df.append(df2, ignore_index=True)
df
df.drop(index=4)
df.drop(index=df[df['last'] == 'Doe'].index)
| video_6/snippets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 10. Convolutional Neural Networks II
# ## Large Convolutional Networks
#
# There are several architectures in the field of Convolutional Networks that have a name. The most common are:
#
# + **LeNet**, 1990’s.
# <center>
# <img src="images/conv2.png" alt="" style="width: 600px;"/>
# </center>
#
#
#
# + **AlexNet**. 2012.
# <center>
# <img src="images/alexnet.png" alt="" style="width: 700px;"/>
# (Source: http://papers.nips.cc/paper/4824-imagenet-classification-with-deep-convolutional-neural-networks.pdf)
# </center>
#
#
# + **ZF Net**. The ILSVRC 2013 winner was a Convolutional Network from <NAME> and <NAME>. It became known as the ZFNet (short for Zeiler & Fergus Net). It was an improvement on AlexNet by tweaking the architecture hyperparameters, in particular by expanding the size of the middle convolutional layers and making the stride and filter size on the first layer smaller.
# <center>
# <img src="images/zfnet.png" alt="" style="width: 700px;"/>
# (Source: https://www.cs.nyu.edu/~fergus/papers/zeilerECCV2014.pdf)
# </center>
#
#
# + **VGGNet**. The runner-up in ILSVRC 2014 was the network from <NAME> and <NAME> that became known as the VGGNet. Its main contribution was in showing that the depth of the network is a critical component for good performance. Their final best network contains 16 CONV/FC layers and, appealingly, features an **extremely homogeneous architecture that only performs 3x3 convolutions and 2x2 pooling from the beginning to the end**. A downside of the VGGNet is that it is more expensive to evaluate and uses a lot more memory and parameters. Most of these parameters are in the first fully connected layer, and it was since found that these FC layers can be removed with no performance downgrade, significantly reducing the number of necessary parameters.
#
#
# <center>
# <img src="images/vgg16.png" alt="" style="width: 600px;"/>
# (Source: https://blog.heuritech.com/2016/02/29/a-brief-report-of-the-heuritech-deep-learning-meetup-5/)
# </center>
# +
# Small VGG-like convnet in Keras
import numpy as np
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras.optimizers import SGD
# Generate dummy data
def to_categorical(y, num_classes=None):
"""
Converts a class vector (integers) to binary class matrix.
"""
y = np.array(y, dtype='int').ravel()
if not num_classes:
num_classes = np.max(y) + 1
n = y.shape[0]
categorical = np.zeros((n, num_classes))
categorical[np.arange(n), y] = 1
return categorical
x_train = np.random.random((100, 100, 100, 3))
y_train = to_categorical(np.random.randint(10, size=(100, 1)), num_classes=10)
x_test = np.random.random((20, 100, 100, 3))
y_test = to_categorical(np.random.randint(10, size=(20, 1)), num_classes=10)
model = Sequential()
# input: 100x100 images with 3 channels -> (100, 100, 3) tensors.
# this applies 32 convolution filters of size 3x3 each.
model.add(Conv2D(32, 3, 3, activation='relu', input_shape=(100, 100, 3)))
model.add(Conv2D(32, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, 3, 3, activation='relu'))
model.add(Conv2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax'))
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', optimizer=sgd)
print(model.summary())
model.fit(x_train, y_train, batch_size=32, nb_epoch=10)
score = model.evaluate(x_test, y_test, batch_size=32)
# -
# **Exercise**
#
# + Why do we have 896 parameters in the ``convolution2d_1`` layer of the previous example?
#
# + Compute the number of parameters of the original VGG16 (all CONV layers are 3x3).
# > The VGG16 architecture is: INPUT: [224x224x3] $\rightarrow$ CONV3-64: [224x224x64] $\rightarrow$ CONV3-64: [224x224x64] $\rightarrow$ POOL2: [112x112x64] $\rightarrow$ CONV3-128: [112x112x128] $\rightarrow$ CONV3-128: [112x112x128] $\rightarrow$ POOL2: [56x56x128] $\rightarrow$ CONV3-256: [56x56x256] $\rightarrow$ CONV3-256: [56x56x256] $\rightarrow$ CONV3-256: [56x56x256] $\rightarrow$ POOL2: [28x28x256] $\rightarrow$ CONV3-512: [28x28x512] $\rightarrow$ CONV3-512: [28x28x512] $\rightarrow$ CONV3-512: [28x28x512] $\rightarrow$ POOL2: [14x14x512] $\rightarrow$ CONV3-512: [14x14x512] $\rightarrow$ CONV3-512: [14x14x512] $\rightarrow$ CONV3-512: [14x14x512] $\rightarrow$ POOL2: [7x7x512] $\rightarrow$ FC: [1x1x4096] $\rightarrow$ FC: [1x1x4096] $\rightarrow$ FC: [1x1x1000].
#
# + The largest bottleneck to be aware of when constructing ConvNet architectures is the memory bottleneck. What is the necessary memory size (supposing that we need 4 bytes for each element) to store intermediate data?
#
#
# +
# your code here
# -
# ## More Large Convolutional Networks
#
#
# + **GoogLeNet**. The ILSVRC 2014 winner was a Convolutional Network from Sze<NAME> al. from Google. Its main contribution was the development of an **Inception Module** that dramatically reduced the number of parameters in the network (4M, compared to AlexNet with 60M). Additionally, this paper uses Average Pooling instead of Fully Connected layers at the top of the ConvNet, eliminating a large amount of parameters that do not seem to matter much. There are also several followup versions to the GoogLeNet, most recently Inception-v4.
#
#
# <center>
# <img src="images/googlenet.jpg" alt="" style="width: 700px;"/>
# GoogLeNet Architecture. Source: https://arxiv.org/pdf/1409.4842v1.pdf
# </center>
#
# > Blue Box: Convolution | Red Box: Pooling | Yelow Box: Softmax | Green Box: Normalization
#
# <center>
# <img src="images/inception.png" alt="" style="width: 400px;"/>
# Inception Layer. Source: https://arxiv.org/pdf/1409.4842v1.pdf
# </center>
#
# <center>
# <img src="images/googlenet2.png" alt="" style="width: 600px;"/>
# GoogLeNet parameters and ops. Source: https://arxiv.org/pdf/1409.4842v1.pdf
# </center>
#
# > What is the role of 1x1 convolutions?
#
# + **ResNet**. Residual Network developed by <NAME> al. was the winner of ILSVRC 2015. It features special skip connections and a heavy use of batch normalization. A Residual Network, or ResNet is a neural network architecture which solves the problem of vanishing gradients in the simplest way possible. If there is trouble sending the gradient signal backwards, why not provide the network with a shortcut at each layer to make things happen more smoothly? The architecture is also missing fully connected layers at the end of the network.
#
# <center>
# <img src="images/res1.png" alt="" style="width: 400px;"/>
# (Source: https://arxiv.org/pdf/1512.03385.pdf)
# </center>
#
# <center>
# <img src="images/resnet.png" alt="" style="width: 400px;"/>
# (Source: https://arxiv.org/pdf/1512.03385.pdf)
# </cente
# ## Deeper is better?
#
# When it comes to neural network design, the trend in the past few years has pointed in one direction: deeper.
#
# Whereas the state of the art only a few years ago consisted of networks which were roughly twelve layers deep, it is now not surprising to come across networks which are hundreds of layers deep.
#
# This move hasn’t just consisted of greater depth for depths sake. For many applications, the most prominent of which being object classification, the deeper the neural network, the better the performance.
#
# So the problem is to design a network in which the gradient can more easily reach all the layers of a network which might be dozens, or even hundreds of layers deep. This is the goal behind some of state of the art architectures: ResNets, HighwayNets, and DenseNets.
#
# **HighwayNets** builds on the ResNet in a pretty intuitive way. The Highway Network preserves the shortcuts introduced in the ResNet, but augments them with a learnable parameter to determine to what extent each layer should be a skip connection or a nonlinear connection. Layers in a Highway Network are defined as follows:
#
# $$ y = H(x, W_H) \cdot T(x,W_T) + x \cdot C(x, W_C) $$
#
# In this equation we can see an outline of two kinds of layers discussed: $y = H(x,W_H)$ mirrors the traditional layer, and $y = H(x,W_H) + x$ mirrors our residual unit.
#
# The traditional layer can be implemented as:
#
# ```python
# def dense(x, input_size, output_size, activation):
# W = tf.Variable(tf.truncated_normal([input_size, output_size], stddev=0.1), name="weight")
# b = tf.Variable(tf.constant(0.1, shape=[output_size]), name="bias")
# y = activation(tf.matmul(x, W) + b)
# return y
# ```
#
# What is new is the $T(x,W_t)$, the transform gate function and $C(x,W_C) = 1 - T(x,W_t)$, the carry gate function. What happens is that when the transform gate is 1, we pass through our activation (H) and suppress the carry gate (since it will be 0). When the carry gate is 1, we pass through the unmodified input (x), while the activation is suppressed.
#
# ```python
# def highway(x, size, activation, carry_bias=-1.0):
# W_T = tf.Variable(tf.truncated_normal([size, size], stddev=0.1), name="weight_transform")
# b_T = tf.Variable(tf.constant(carry_bias, shape=[size]), name="bias_transform")
#
# W = tf.Variable(tf.truncated_normal([size, size], stddev=0.1), name="weight")
# b = tf.Variable(tf.constant(0.1, shape=[size]), name="bias")
#
# T = tf.sigmoid(tf.matmul(x, W_T) + b_T, name="transform_gate")
# H = activation(tf.matmul(x, W) + b, name="activation")
# C = tf.sub(1.0, T, name="carry_gate")
#
# y = tf.add(tf.mul(H, T), tf.mul(x, C), "y")
# return y
# ```
#
# With this kind of network you can train models with hundreds of layers.
#
# **DenseNet** takes the insights of the skip connection to the extreme. The idea here is that if connecting a skip connection from the previous layer improves performance, why not connect every layer to every other layer? That way there is always a direct route for the information backwards through the network.
#
# <center>
# <img src="images/densenet.png" alt="" style="width: 700px;"/>
# (Source: https://arxiv.org/abs/1608.06993)
# </center>
#
# Instead of using an addition however, the DenseNet relies on stacking of layers. Mathematically this looks like:
#
# $$ y = f(x, x-1, x-2, \dots, x-n) $$
#
# This architecture makes intuitive sense in both the feedforward and feed backward settings. In the feed-forward setting, a task may benefit from being able to get low-level feature activations in addition to high level feature activations. In classifying objects for example, a lower layer of the network may determine edges in an image, whereas a higher layer would determine larger-scale features such as presence of faces. There may be cases where being able to use information about edges can help in determining the correct object in a complex scene. In the backwards case, having all the layers connected allows us to quickly send gradients to their respective places in the network easily.
#
#
# ## Fully Convolutional Networks
#
# (Source: http://cs231n.github.io/convolutional-networks/#convert)
#
# The only difference between **Fully Connected (FC)** and **Convolutional (CONV)** layers is that the neurons in the CONV layer are connected only to a local region in the input, and that many of the neurons in a CONV volume share parameters.
#
# However, the neurons in both layers still compute dot products, so their functional form is identical.
#
# Then, it is easy to see that for any CONV layer there is an FC layer that implements the same forward function. The weight matrix would be a large matrix that is mostly zero except for at certain blocks (due to local connectivity) where the weights in many of the blocks are equal (due to parameter sharing).
#
# <center>
# <img src="images/t10.png" alt="" style="width: 800px;"/>
# <center>
#
#
# Conversely, any FC layer can be converted to a CONV layer.
#
# Let $F$ be the receptive field size of the CONV layer neurons, $S$ the stride with which they are applied, $P$ the amount of zero padding used on the border, and $K$ the depth (number of bands) of the CONV layer.
#
# For example, an FC layer with $K=4096$ that is looking at some input volume of size $7×7×512$ can be equivalently expressed as a CONV layer with $F=7,P=0,S=1,K=4096$.
#
# In other words, we are setting the filter size to be exactly the size of the input volume, and hence the output will simply be 1×1×4096 since only a single depth column “fits” across the input volume, giving identical result as the initial FC layer.
#
# This can be very useful!
# Consider a ConvNet architecture that takes a $224x224x3$ image, and then uses a series of CONV layers and POOL layers to reduce the image to an activations volume of size $7x7x512$. From there, two FC layers of size $4096$ and finally the last FC layers with $1000$ neurons that compute the class scores. We can convert each of these three FC layers to CONV layers as described above:
#
# + Replace the first FC layer that looks at $[7x7x512]$ volume with a CONV layer that uses filter size $F=7$, giving output volume $[1x1x4096]$.
# + Replace the second FC layer with a CONV layer that uses filter size $F=1$, giving output volume $[1x1x4096]$.
# + Replace the last FC layer similarly, with $F=1$, giving final output $[1x1x1000]$.
#
# It turns out that this conversion allows us to “slide” the original ConvNet very efficiently across many spatial positions in a larger image, in a single forward pass.
#
# For example, if $224x224$ image gives a volume of size $[7x7x512]$ - i.e. a reduction by 32, then forwarding an image of size 384x384 through the converted architecture would give the equivalent volume in size $[12x12x512]$, since $384/32 = 12$. Following through with the next 3 CONV layers that we just converted from FC layers would now give the final volume of size $[6x6x1000]$, since $(12 - 7)/1 + 1 = 6$. Note that instead of a single vector of class scores of size $[1x1x1000]$, we’re now getting and entire $6x6$ array of class scores across the $384x384$ image.
#
# > Evaluating the original ConvNet (with FC layers) independently across $224x224$ crops of the $384x384$ image in strides of 32 pixels gives an identical result to forwarding the converted ConvNet one time. Forwarding the converted ConvNet a single time is much more efficient than iterating the original ConvNet over all those 36 locations, since the 36 evaluations share computation.
#
#
# ## Object Detection and Segmentation
# (Source: https://blog.athelas.com/a-brief-history-of-cnns-in-image-segmentation-from-r-cnn-to-mask-r-cnn-34ea83205de4)
#
# In classification, there’s generally an image with a single object as the focus and the task is to say what that image is. But when we look at the world around us, we see complicated sights with multiple overlapping objects, and different backgrounds and we not only classify these different objects but also identify their boundaries, differences, and relations to one another!
#
# To what extent do CNN generalize to object detection? Object detection is the task of finding the different objects in an image and classifying them.
#
# ### R-CNN
#
# A team, comprised of <NAME> (a name we’ll see again), <NAME>, and <NAME> found that this problem can be solved with AlexNet by testing on the PASCAL VOC Challenge, a popular object detection challenge akin to ImageNet.
#
# The goal of R-CNN is to take in an image, and correctly identify where the main objects (via a bounding box) in the image.
#
# >Inputs: Image
#
# >Outputs: Bounding boxes + labels for each object in the image.
#
# But how do we find out where these bounding boxes are? R-CNN proposes a bunch of boxes in the image and see if any of them actually correspond to an object.
#
# R-CNN creates these bounding boxes, or region proposals, using a process called Selective Search (see http://www.cs.cornell.edu/courses/cs7670/2014sp/slides/VisionSeminar14.pdf).
#
# At a high level, Selective Search (shown in the image above) looks at the image through windows of different sizes, and for each size tries to group together adjacent pixels by texture, color, or intensity to identify objects.
#
# Once the proposals are created, R-CNN warps the region to a standard square size and passes it through to a modified version of AlexNet.
#
# On the final layer of the CNN, R-CNN adds a Support Vector Machine (SVM) that simply classifies whether this is an object, and if so what object.
#
# <center>
# <img src="images/rcnn.png" alt="" style="width: 800px;"/>
# </center>
#
# Now, having found the object in the box, can we tighten the box to fit the true dimensions of the object? We can, and this is the final step of R-CNN. R-CNN runs a simple linear regression on the region proposal to generate tighter bounding box coordinates to get our final result. Here are the inputs and outputs of this regression model:
#
# > Inputs: sub-regions of the image corresponding to objects.
#
# > Outputs: New bounding box coordinates for the object in the sub-region.
#
#
#
# ### Fast R-CNN
#
# R-CNN works really well, but is really quite slow for a few simple reasons:
# + It requires a forward pass of the CNN (AlexNet) for every single region proposal for every single image (that’s around 2000 forward passes per image!).
# + It has to train three different models separately - the CNN to generate image features, the classifier that predicts the class, and the regression model to tighten the bounding boxes. This makes the pipeline extremely hard to train.
#
# In 2015, <NAME>, the first author of R-CNN, solved both these problems, leading to Fast R-CNN.
#
# For the forward pass of the CNN, Girshick realized that for each image, a lot of proposed regions for the image invariably overlapped causing us to run the same CNN computation again and again (~2000 times!). His insight was simple — Why not run the CNN just once per image and then find a way to share that computation across the ~2000 proposals?
#
# This is exactly what Fast R-CNN does using a technique known as **RoIPool** (Region of Interest Pooling). At its core, RoIPool shares the forward pass of a CNN for an image across its subregions. In the image below, notice how the CNN features for each region are obtained by selecting a corresponding region from the CNN’s feature map. Then, the features in each region are pooled (usually using max pooling). So all it takes us is one pass of the original image as opposed to ~2000!
#
# <center>
# <img src="images/fastrcnn.png" alt="" style="width: 600px;"/>
# (Source: Stanford’s CS231N slides by <NAME>, <NAME>, and <NAME>)
# </center>
#
#
# The second insight of Fast R-CNN is to jointly train the CNN, classifier, and bounding box regressor in a single model. Where earlier we had different models to extract image features (CNN), classify (SVM), and tighten bounding boxes (regressor), Fast R-CNN instead used a single network to compute all three.
#
# <center>
# <img src="images/fastrcnn2.png" alt="" style="width: 400px;"/>
# (Source: https://www.slideshare.net/simplyinsimple/detection-52781995)
# </center>
#
# ### Faster R-CNN
#
# Even with all these advancements, there was still one remaining bottleneck in the Fast R-CNN process — the region proposer. As we saw, the very first step to detecting the locations of objects is generating a bunch of potential bounding boxes or regions of interest to test. In Fast R-CNN, these proposals were created using Selective Search, a fairly slow process that was found to be the bottleneck of the overall process.
#
# In the middle 2015, a team at Microsoft Research composed of <NAME>, <NAME>, <NAME>, and <NAME>, found a way to make the region proposal step almost cost free through an architecture they (creatively) named Faster R-CNN.
#
# The insight of Faster R-CNN was that region proposals depended on features of the image that were already calculated with the forward pass of the CNN (first step of classification). So why not reuse those same CNN results for region proposals instead of running a separate selective search algorithm?
#
# <center>
# <img src="images/fasterrcnn.png" alt="" style="width: 400px;"/>
# (Source: https://arxiv.org/abs/1506.01497)
# </center>
#
# Here are the inputs and outputs of their model:
#
# > Inputs: Images (Notice how region proposals are not needed).
#
# > Outputs: Classifications and bounding box coordinates of objects in the images.
#
# ### Mask R-CNN
#
# So far, we’ve seen how we’ve been able to use CNN features in many interesting ways to effectively locate different objects in an image with bounding boxes.
#
# Can we extend such techniques to go one step further and locate exact pixels of each object instead of just bounding boxes? This problem, known as image segmentation, is what <NAME> and a team of researchers, including Girshick, explored at Facebook AI using an architecture known as Mask R-CNN.
#
# Given that Faster R-CNN works so well for object detection, could we extend it to also carry out pixel level segmentation?
#
# Mask R-CNN does this by adding a branch to Faster R-CNN that outputs a binary mask that says whether or not a given pixel is part of an object. The branch (in white in the above image), as before, is just a Fully Convolutional Network on top of a CNN based feature map. Here are its inputs and outputs:
#
# > Inputs: CNN Feature Map.
# > Outputs: Matrix with 1s on all locations where the pixel belongs to the object and 0s elsewhere (this is known as a binary mask).
#
# <center>
# <img src="images/pixelrcnn.png" alt="" style="width: 700px;"/>
# (Source: https://arxiv.org/abs/1703.06870)
# </center>
# ## 1D-Conv for text classification
#
# **IMDB Movie reviews sentiment classification**: Dataset of 25,000 movies reviews from IMDB, labeled by sentiment (positive/negative). Reviews have been preprocessed, and each review is encoded as a sequence of word indexes (integers). For convenience, words are indexed by overall frequency in the dataset, so that for instance the integer "3" encodes the 3rd most frequent word in the data. This allows for quick filtering operations such as: "only consider the top 10,000 most common words, but eliminate the top 20 most common words".
#
# The seminal research paper on this subject was published by <NAME> on 2014. In this paper Yoon Kim has laid the foundations for how to model and process text by convolutional neural networks for the purpose of sentiment analysis. He has shown that by simple one-dimentional convolutional networks, one can develops very simple neural networks that reach 90% accuracy very quickly.
#
# Here is the text of an example review from our dataset:
#
# <center>
# <img src="images/review1.png" alt="" style="width: 600px;"/>
# <center>
# +
'''
This example demonstrates the use of Convolution1D for text classification.
'''
from __future__ import print_function
import numpy as np
np.random.seed(1337) # for reproducibility
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation, Flatten
from keras.layers.embeddings import Embedding
from keras.layers.convolutional import Convolution1D, MaxPooling1D
from keras.datasets import imdb
# set parameters:
max_features = 5000
maxlen = 100
batch_size = 32
embedding_dims = 100
nb_filter = 250
filter_length = 3
hidden_dims = 250
nb_epoch = 10
print('Loading data...')
(X_train, y_train), (X_test, y_test) = imdb.load_data(nb_words=max_features)
print(len(X_train), ' train sequences \n')
print(len(X_test), ' test sequences \n')
print('Pad sequences (samples x time)')
X_train = sequence.pad_sequences(X_train, maxlen=maxlen)
X_test = sequence.pad_sequences(X_test, maxlen=maxlen)
print('X_train shape:', X_train.shape)
print('X_test shape:', X_test.shape)
print('Build model...')
model = Sequential()
# we start off with an efficient embedding layer which maps
# our vocab indices into embedding_dims dimensions
model.add(Embedding(max_features, embedding_dims, input_length=maxlen))
model.add(Dropout(0.25))
# we add a Convolution1D, which will learn nb_filter
# word group filters of size filter_length:
model.add(Convolution1D(nb_filter=nb_filter,
filter_length=filter_length,
border_mode='valid',
activation='relu',
subsample_length=1))
# we use standard max pooling (halving the output of the previous layer):
model.add(MaxPooling1D(pool_length=2))
model.add(Convolution1D(nb_filter=nb_filter,
filter_length=filter_length,
border_mode='valid',
activation='relu',
subsample_length=1))
model.add(MaxPooling1D(pool_length=2))
# We flatten the output of the conv layer,
# so that we can add a vanilla dense layer:
model.add(Flatten())
# We add a vanilla hidden layer:
model.add(Dense(hidden_dims))
model.add(Dropout(0.25))
model.add(Activation('relu'))
# We project onto a single unit output layer, and squash it with a sigmoid:
model.add(Dense(1))
model.add(Activation('sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
model.fit(X_train, y_train,
batch_size=batch_size,
nb_epoch=nb_epoch,
validation_data=(X_test, y_test))
| 10. Convolutional Neural Networks II.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="0Knx0FNjNIfv"
# ### Questions about Airbnb Data
#
# Which features correlate with an Airbnb homestay price? <br>
# What are the best neighbourhoods to stay in Seattle? <br>
# What is the best time of the year to visit Seattle?
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 890} id="aatbK1h0NE1B" executionInfo={"status": "ok", "timestamp": 1639821177846, "user_tz": -60, "elapsed": 3506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="30d4c941-ad97-424d-d609-065b078c3523"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import plotly.express as px
from scipy.stats import pearsonr
from plotly.subplots import make_subplots
import plotly.graph_objects as go
# %matplotlib inline
data = pd.read_csv('calendar.csv')
df = pd.read_csv('listings.csv')
df.head()
# + [markdown] id="aiqvwo7hO6ra"
# # Correlation Coefficients of Airbnb Pricings
#
# **listings.csv**
# - neighborhood_overview
# - transit description (0 or 1)
# - neighborhood
# - property_type
# - room_type
# - accommodates
# - bathrooms
# - bedrooms
# - beds
# - bed_type
# - number of amenities
# - free parking
# - wireless internet
# - square_feets
# - price
# - number_of_reviews
# + [markdown] id="s1-lrRNXW2ND"
# ### Data Cleaning
# + id="QJkGPjZQO6I2" executionInfo={"status": "ok", "timestamp": 1639818065875, "user_tz": -60, "elapsed": 3051, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} colab={"base_uri": "https://localhost:8080/"} outputId="cc97c4e2-257e-41a7-e7f3-07aa7548e797"
def clean_data(df):
'''
INPUT
df - pandas dataframe
OUTPUT
X - A matrix holding all of the variables you want to consider when predicting the response
y - the corresponding response vector
Function to obtain the correct X and y objects
This function cleans df using the following steps to produce X and y:
1. Drop all the rows with no salaries
2. Create X as all the columns that are to be investigated
3. Create y as the Price column
4. create new columns for "number of amenities", "Wifi", "Free Parking", "transit_description"
5. transfer number_of_amenities to numeric
6. y: Convert string to float
7. For each numeric variable in X, fill the column with the mean value of the column.
8. X: drop 'transit', 'amenities'
9. Create dummy columns for all the categorical variables in X, drop the original columns
'''
# 1. 'Price' is the target we want to analysis, therefore we can only calculate each row with valid data in this column
df.dropna(subset=['price'], axis=0, inplace=True)
# 2.
X = df.loc[:, ('transit', 'neighbourhood', 'property_type', 'room_type', 'accommodates',
'bathrooms', 'bedrooms', 'beds', 'bed_type', 'amenities', 'square_feet', 'number_of_reviews')]
# 3.
y = df.price
# 4.
X['number_of_amenities'] = ""
X['wifi'] = 0
X['free_parking'] = 0
X['transit_description'] = 0
for idx in X.index:
# count amenitites by ',' character
X.number_of_amenities.loc[idx] = X.amenities.loc[idx].count(',') + 1
# check if wifi is available
if "Wireless" in X.amenities.loc[idx]:
X.wifi.loc[idx] = 1
# check if free parking is available
if "Free Parking" in X.amenities.loc[idx]:
X.free_parking.loc[idx] = 1
# check if transit entry is empty
entry = X.transit.loc[idx]
if pd.isna(entry) == False:
X.transit_description.loc[idx] = 1
# 5.
X.number_of_amenities = pd.to_numeric(X.number_of_amenities)
# 6.
y = y.str.replace('$', '')
y = y.str.replace(',', '')
y = pd.to_numeric(y)
# 7.
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
num_cols = X.select_dtypes(include=numerics).columns
X[num_cols] = X[num_cols].fillna(value=X[num_cols].mean())
# 8.
X.drop(labels=['transit', 'amenities'], inplace=True, axis=1)
# 9.
cat_cols = X.select_dtypes(include=['object']).columns
for col in cat_cols:
try:
# for each cat add dummy var, drop original column
X = pd.concat([X.drop(col, axis=1), pd.get_dummies(X[col], prefix=col, prefix_sep='_')], axis=1)
except:
continue
return X, y
#Use the function to create X and y
X, y = clean_data(df)
# + colab={"base_uri": "https://localhost:8080/"} id="eDx2DyU4UIJ6" executionInfo={"status": "ok", "timestamp": 1639814433823, "user_tz": -60, "elapsed": 209, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="375c6ff8-029e-459d-ffb2-f452d7b7bb6e"
print(f"Missing values in X:\n{X.isnull().sum()} \n")
print(f"Number of Rows of X: {X.shape[0]}\n")
print(f"Is there any NaN in X: {X.isnull().values.any()}")
# + colab={"base_uri": "https://localhost:8080/", "height": 317} id="kFLktbeiVG1e" executionInfo={"status": "ok", "timestamp": 1639818075462, "user_tz": -60, "elapsed": 234, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="86f2667d-d61d-492b-e42e-bc49f5ba52d6"
X.head()
# + [markdown] id="P7MJvPpX_FLw"
# ### Check Column 'Square Feet'
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="AC11BFA4x-9H" executionInfo={"status": "ok", "timestamp": 1639814450837, "user_tz": -60, "elapsed": 243, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="47f2a507-473a-4833-f6d6-74db10bb5542"
print(f"{df.square_feet.isna().values.sum()} of {df.shape[0]} are NaN!")
# + [markdown] id="_s76VTt5XcC1"
# ### Create Correlation Coefficients
# + id="nguj7tgcXbnx"
# concatenate X and y
df_clean = pd.concat([X, y], axis=1)
# + id="3vM6Pp11Y2yD"
from _plotly_utils.basevalidators import DataArrayValidator
df_corr = df_clean.corr(method='pearson')
#create DataFrame
corr_coef = pd.DataFrame(df_corr.price)
# drop last row (price)
corr_price = corr_coef.drop('price')
#create column with absolute value
corr_price['abs'] = df_corr.price.abs()
# sort by absolute value
corr_sort = corr_price.sort_values(by='abs', ascending=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="Nq49ezqnZM9n" executionInfo={"status": "ok", "timestamp": 1639818086267, "user_tz": -60, "elapsed": 568, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="6521556e-b855-4b7b-e093-b3c6283df85f"
fig = px.bar(corr_sort, x=corr_sort.index[:20], y=corr_sort.price[:20], labels={'y':'Value', 'x': 'Correlation Coefficients'})
fig.update_layout(title_text='Top 20 Correlation Coefficients for Airbnb Prices')
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="fIsIVpGchFKt" executionInfo={"status": "ok", "timestamp": 1639818091286, "user_tz": -60, "elapsed": 864, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="422a1b53-ee39-4120-bf33-2ec44a018ff9"
fig = px.bar(corr_sort, x=corr_sort.index[:10], y=corr_sort.price[:10],
labels={'y':'Value', 'x': 'Correlation Coefficients'},
width=600, height=400)
fig.update_layout(title_text='Top 10 Correlation Coefficients for Airbnb Prices')
fig.show()
# + [markdown] id="kZ24RhqYjzYB"
# ### Analysis the correlation coefficients
#
# I use plotly express subplots to show the correlation between the price and the top 10 features:
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="wlKUZzfkvZQa" executionInfo={"status": "ok", "timestamp": 1639817535173, "user_tz": -60, "elapsed": 248, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="f635e641-d918-45d4-d898-8691c9c73c63"
corr_sort[:10]
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="rpYSGZvdqV1I" executionInfo={"status": "ok", "timestamp": 1639818113400, "user_tz": -60, "elapsed": 951, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="29b4b1c9-efa0-4556-921d-8bbc4477fb11"
trace1 = go.Scatter(x=df_clean.accommodates, y=df_clean.price, mode="markers")
trace2 = go.Scatter(x=df_clean.bedrooms, y=df_clean.price, mode="markers")
trace3 = go.Scatter(x=df_clean.beds, y=df_clean.price, mode="markers")
trace4 = go.Scatter(x=df_clean.bathrooms, y=df_clean.price, mode="markers")
trace5 = go.Scatter(x=df_clean['room_type_Entire home/apt'], y=df_clean.price, mode="markers")
trace6 = go.Scatter(x=df_clean['room_type_Private room'], y=df_clean.price, mode="markers")
trace7 = go.Scatter(x=df_clean['number_of_amenities'], y=df_clean.price, mode="markers")
trace8 = go.Scatter(x=df_clean['room_type_Shared room'], y=df_clean.price, mode="markers")
trace9 = go.Scatter(x=df_clean['bed_type_Real Bed'], y=df_clean.price, mode="markers")
trace10 = go.Scatter(x=df_clean['number_of_reviews'], y=df_clean.price, mode="markers")
fig = make_subplots(rows=2, cols=5, subplot_titles=("Accommodates","Bedrooms",
"Beds", "Bathrooms", "Entire Home/Apt", "Private room", "Amenities", "Shared room",
"Real Bed", "Reviews"))
fig.append_trace(trace1, row=1, col=1)
fig.append_trace(trace2, row=1, col=2)
fig.append_trace(trace3, row=1, col=3)
fig.append_trace(trace4, row=1, col=4)
fig.append_trace(trace5, row=1, col=5)
fig.append_trace(trace6, row=2, col=1)
fig.append_trace(trace7, row=2, col=2)
fig.append_trace(trace8, row=2, col=3)
fig.append_trace(trace9, row=2, col=4)
fig.append_trace(trace10, row=2, col=5)
fig.update_layout(title_text="Top 10 correlation coefficients")
fig.update_layout(showlegend=False)
fig.show()
# + [markdown] id="Sch2hZwPzx7j"
# Overall the coefficients make sense. E.g. an entire home has a positive influence to the price, where a private room or shared room correlate to an lower price. Also the correlation to the different bed types is reasonable. The coefficient for the number of reviews is negative. Looking at the scatter plot above, shows that the expensive homestay only have a few reviews, where the more affordable ones tend to have more.
#
# Most of the plots don't show clear linear relation: Accommodates, Beds, Bathrooms, Amenities and Reviews. Therefore the correlation coefficients are not close to ±1.
#
# I calculate the p-value for the top 20 correlation coefficients to validate the linear relationship.
# + id="kvRq3lux6ZER"
# calculate the p-value
def calculate_pvalues(df):
dfcols = pd.DataFrame(columns=df.columns)
pvalues = dfcols.transpose().join(dfcols, how='outer')
for r in df.columns:
for c in df.columns:
pvalues[r][c] = round(pearsonr(df[r], df[c])[1], 4)
return pvalues
p_values = calculate_pvalues(df_clean).price
# + colab={"base_uri": "https://localhost:8080/"} id="66WlavVD-Knl" executionInfo={"status": "ok", "timestamp": 1639822437712, "user_tz": -60, "elapsed": 310, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="1ca74e66-5942-43df-8d3d-23f5ec910738"
# show the p_values of the top 20 correlation coefficients
p_values[['accommodates', 'bedrooms', 'beds', 'bathrooms', 'room_type_Entire home/apt',
'room_type_Private room', 'number_of_amenities', 'room_type_Shared room',
'bed_type_Real Bed', 'number_of_reviews', 'neighbourhood_Queen Anne',
'neighbourhood_Belltown', 'bed_type_Futon', 'square_feet', 'free_parking',
'property_type_Boat', 'neighbourhood_Portage Bay', 'neighbourhood_Magnolia',
'bed_type_Pull-out Sofa', 'neighbourhood_North Beacon Hill']]
# + [markdown] id="ra8YoHUoA0Wu"
# All p-values are below 0.05 (=5%). Therefore a correlation between the features and the price clearly exists.
# + [markdown] id="cVq73F3m3t5Z"
# # Popular Neighbourhoods in Seattle
# + id="VXAzqvzw34uE"
pop_nbh = df.groupby('neighbourhood').count().id
top20_nbh = pop_nbh.sort_values(ascending=False)[:20]
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="PmAE0abj4536" executionInfo={"status": "ok", "timestamp": 1639546425675, "user_tz": -60, "elapsed": 426, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="ece3532b-d1f4-47a6-f01c-8893ff05f8ff"
fig = px.bar(top20_nbh, x=top20_nbh.index, y=top20_nbh.values, labels={'y':'Counts', 'x': 'Neighbourhood'})
fig.update_layout(title_text='Top 20 Most Popular Neighbourhoods in Seattle')
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="oEJZyTDW5b1H" executionInfo={"status": "ok", "timestamp": 1639546425676, "user_tz": -60, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="665c389b-5dc1-4571-dc53-d33c38c067e7"
fig = px.bar(top20_nbh, x=top20_nbh.index[:10], y=top20_nbh.values[:10],
labels={'y':'Counts', 'x': 'Neighbourhood'},
width=600, height=400)
fig.update_layout(title_text='Top 10 Most Popular Neighbourhoods in Seattle')
fig.show()
# + [markdown] id="tMl4ZkQC10Sh"
# The most popular neighborhood is by fare Capitol Hill with more than 350 counts. Followed by Ballard, Belltwon, Minor and Queen Anne which have around 200 counts.
# The popularity is of course also comparable with the population density.
# + [markdown] id="H8u_4oJI6Owr"
# # Best time of the year to visit Seattle
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="2DwXkdDv6ePp" executionInfo={"status": "ok", "timestamp": 1639546426991, "user_tz": -60, "elapsed": 1326, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="ba6e62d2-18c7-4ee1-fe4f-71a3b66f2131"
# convert price column to float:
data.price = data.price.str.replace('$', '')
data.price = data.price.str.replace(',', '')
data.price = pd.to_numeric(data.price)
data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="k34anbSN7Aa7" executionInfo={"status": "ok", "timestamp": 1639546426992, "user_tz": -60, "elapsed": 13, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="b03389dc-9feb-421a-b8df-348578c74a6c"
print(f"{data.price.isna().values.sum()} of {data.shape[0]} are NaN ({data.price.isna().values.sum()*100/data.shape[0]:.3}%)")
# + id="Ks7YmGJ97g7Q"
# convert date column to datetime
data.date = pd.to_datetime(data.date)
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="toBOjYxCBLgv" executionInfo={"status": "ok", "timestamp": 1639545528673, "user_tz": -60, "elapsed": 2190, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16820290495133761291"}} outputId="5a91145e-47f0-405a-b6e2-b52ffb163ffd"
mean_price = data.groupby(data.date.dt.month).price.mean()
month_avail = data[data.available=='t'].groupby(data.date.dt.month).available.value_counts()
month_avail
px.bar(month_avail, x=month_avail.index.get_level_values(0), y=month_avail.values, color=mean_price)
fig = px.bar(month_avail, x=month_avail.index.get_level_values(0), y=month_avail.values, color=mean_price,
labels={'y':'Availability', 'x': 'Months of 2016', 'color': 'Mean Price [$]'},
width=600, height=400)
fig.update_layout(title_text='Availability and Pricing in Seattle in 2016')
fig.update_xaxes(tickvals=[1,2,3,4,5,6,7,8,9,10,11,12])
fig.show()
# + [markdown] id="SN7XGN7I2vEP"
# During the summer the prices are the highest and the availability is low. At the begin of the year 2016 the availability was low as well, but the prices were affordable. At the end of the year towards Christmas the prices went a bit higher, but the availability was high too. So the best months to travel are probably April and May where prices are good and availability high.
| listing_prices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Descriptive and Exploratory Answers
# The below cells describe some Magic numbers and column indices. They offer very rudimentary insight into what columns map on to what data. While we're at it let us also define a handful of helpful functions.
# **PLEASE FIX THE FILE PATH TO SOMETHING THAT POINTS TO A FILE ON YOUR COMPUTER**
# +
# I've gone ahead and used "=" in place of "<-" because the latest R doesn't
# care, also makes the code more (Pythonic)readable IMHO
# Legend of MAGIC Numbers, File Paths and Arguments
CORRELATION_STRATEGY = "pearson"
PATH_TO_DATA = "~/Documents/datascience/"
PRIMARY_FILE = "Data_Adults_1.csv"
INFO_START = 2
INFO_END = 14
DISORDER_START = 15
DISORDER_END = 76
SURVEY_START = 77
SURVEY_END = 378
RCBF_RAW_START = 379
RCBF_RAW_END = 636
RCBF_SCALED_START = 637
RCBF_SCALED_END = 754
# +
# Computes Correlation Between 2 quantities and returns a correlation vector.
# Param: quant_a : The first data frame to correlate
# Param: quant_b : The second data frame to correlate with
# Return: A vector of column correlation
correlate = function(quant_a, quant_b = NA) {
if (is.na(quant_b)){
cor_val = cor(quant_a, method = CORRELATION_STRATEGY)
} else {
cor_val = cor(quant_a, quant_b, method = CORRELATION_STRATEGY)
}
return (cor_val)
}
# This function returns a matrix with 2 rows, one with n smallest values and the other with n largest.
# Param: vec : The vector from which the values need to be extracted
# Param: n : The number of values needed
# Param: req_plot : A switch which will optionally plot the data if required
# Return: A row bound matrix of n smallest and largest values respectively
sort_plot = function(vec, n, req_plot = FALSE) {
smallest_n = sort(vec)[1:n]
largest_n = sort(vec, TRUE)[1:n]
if (req_plot) {
plot(vec)
}
return (rbind(smallest_n, largest_n))
}
# Function prefixes the patient ID column to a data fram for kicks, actually it
# does this so that we can find out who is where.
# Param: start_index : The beginning of the columns to bind the patient ID
# ahead of
# Param: end_index : The end of the columns
# Return: a data frame with the required columns prefixed by patient ID
prefix_id = function(start_index, end_index) {
return (cbind(cleansed_data[2], cleansed_data[, start_index : end_index]))
}
# Function prints the min, max and median of each column in a data set.
# Param: dataset : The data set to crunch
min_max_mean_extract = function(dataset) {
print("Minimum Values")
#print(sapply(dataset[-1], min))
print("Maximum Values")
#print(sapply(dataset[-1], max))
print("Mean Values")
#print(sapply(dataset[-1], mean))
# not printing the values for the sake of brevity
}
## Function to discern levels of nominal variables.
# Param: quantity : The quanttity to potentially factorize
discern_levels = function(quantity) {
char_vals = quantity[sapply(quantity, function(quantity) !is.numeric(quantity))]
apply(char_vals, 2, unique)
}
# -
# ## Descriptive Answers
# <hr>
# ### Descriptive Question - 1
# What is the size of our training data? What is the length of each feature vector and how many features does each vector have?
#
# ### Descriptive Question - 2
# What does the presence of Null values indicate? How should they be dealt with?
spect_data = data.frame(read.csv(paste0(PATH_TO_DATA, PRIMARY_FILE), as.is = TRUE))
cleansed_data = na.omit(spect_data) # deal with Null values
cat("The dataset as is has", nrow(spect_data), "rows and", ncol(spect_data), "columns\n")
cat("The downsampled dataset, omitting rows that have NA values has", nrow(cleansed_data),
"rows and", ncol(cleansed_data), "columns\n")
# **Answer 1**: As evidenced by the code above even when we elect to throw out any row that has a NA value we still have a sizeable datset. This doesn't mean however that we will not use the data thrown out. We will simply downsample till we have a complete yet sizeable dataset and draw corellation conclusions etc. from it. Once we have these theories we will try to apply them to less downsampled datasets to see if we can fill in missing data and verify if our theories still hold.
#
# **Answer 2**: As stated in the answer above, NA values were thrown out. We still had a considerably large dataset after doing so.
# ### Descriptive Question - 3
# What do the features in the vector indicate? What are they for? How are the feature values arrived at?
cleansed_data_patient_info = cleansed_data[,INFO_START : INFO_END]
cleansed_data_disorder_info = prefix_id(DISORDER_START, DISORDER_END)
cleansed_data_survey_info = prefix_id(SURVEY_START, SURVEY_END)
cleansed_data_RCBF_raw_info = prefix_id(RCBF_RAW_START, RCBF_RAW_END)
cleansed_data_RCBF_scaled_info = prefix_id(RCBF_SCALED_START, RCBF_SCALED_END)
# **Answer 3**: By manually examining the data, we realized that certain portions of the dataset contain specific data that can be
# examined mutually exclusively from the other portions of the dataset. We made note of the column indices that allow the data to be split into these mutually exclusive portions and went ahead and split the data set according to those indices. This also allows for easier correlation experiments.
#
# - The columns in *patient_info* represent information about the patient like gender, age, location etc
# - The columns in *disorder_info* represent a series of boolean values that indicate whether the patient ails
# from a particular disorder
# - The columns in *survey_info* represent the patient's responses to the BSC, GSC and LDS survey questions(assuming they are surveys)
# - The columns in *RCBF_raw* represent the raw cerebral blood flow values
# - The columns in *RCBF_scaled* represent the scaled cerebral blood flow values
# ## Exploratory Answers
# <hr/>
# ### Exploratory Question - 1
# What is the mean, maximum and minimum for the various numerical features? Also how do these features scale?
min_max_mean_extract(cleansed_data_patient_info)
min_max_mean_extract(cleansed_data_disorder_info)
min_max_mean_extract(cleansed_data_survey_info)
min_max_mean_extract(cleansed_data_RCBF_raw_info)
min_max_mean_extract(cleansed_data_RCBF_scaled_info)
# **Answer 1**: Though max and min values don't really make sense for data like boolean disorder flags we run them anyway because we might've missed something.
# ### Exploratory Question - 2
# Is there a metric that can be used to separate healthy people from unhealthy ones?
unhealthy_vector = apply(cleansed_data_disorder_info[-1], 1, function(row) any(row[] == 1 ))
cat("There are ", sum(unhealthy_vector == FALSE), "healthy people")
# **Answer 2**: There are 50 healthy people
# ### Exploratory Question - 3
# What is the range of levels nominal values can take?
discern_levels(cleansed_data_patient_info)
discern_levels(cleansed_data_disorder_info)
discern_levels(cleansed_data_survey_info)
discern_levels(cleansed_data_RCBF_raw_info)
discern_levels(cleansed_data_RCBF_scaled_info)
| Code/Assignment-3/Descriptive_Exploratory_Answers_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="../images/26-weeks-of-data-science-banner.jpg"/>
# + [markdown] slideshow={"slide_type": "slide"}
# # Getting Started with Python
# -
# ## About Python
# <img src="../images/python-logo.png" alt="Python" style="width: 500px;"/>
#
#
# Python is a
#
# - general purpose programming language
# - interpreted, not compiled
# - both **dynamically typed** _and_ **strongly typed**
# - supports multiple programming paradigms: object oriented, functional
# - comes in 2 main versions in use today: 2.7 and 3.x
#
# ## Why Python for Data Science?
# ***
#
# Python is great for data science because:
#
# - general purpose programming language (as opposed to R)
# - faster idea to execution to deployment
# - battle-tested
# - mature ML libraries
#
# <div class="alert alert-block alert-success">And it is easy to learn !</div>
#
# <img src="../images/icon/Concept-Alert.png" alt="Concept-Alert" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Python's Interactive Console : The Interpreter
#
# ***
# - The Python interpreter is a console that allows interactive development
# - We are currently using the Jupyter notebook, which uses an advanced Python interpreter called IPython
# - This gives us much more power and flexibility
#
# **Let's try it out !**
#
#
#
#
#
#
#
print("Hello World!") #As usual with any language we start with with the print function
# + [markdown] slideshow={"slide_type": "slide"}
# # What are we going to learn today?
# ***
# - CHAPTER 1 - **Python Basics**
# - **Strings**
# - Creating a String, variable assignments
# - String Indexing & Slicing
# - String Concatenation & Repetition
# - Basic Built-in String Methods
# - **Numbers**
# - Types of Numbers
# - Basic Arithmetic
#
#
#
# - CHAPTER 2 - **Data Types & Data Structures**
# - Lists
# - Dictionaries
# - Sets & Booleans
#
#
# - CHAPTER 3 - **Python Programming Constructs**
# - Loops & Iterative Statements
# - if,elif,else statements
# - for loops, while loops
# - Comprehensions
# - Exception Handling
# - Modules, Packages,
# - File I/O operations
#
# -
# # CHAPTER - 1 : Python Basics
# ***
# Let's understand
# - Basic data types
# - Variables and Scoping
# - Modules, Packages and the **`import`** statement
# - Operators
#
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Concept-Alert" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Strings
# ***
#
# Strings are used in Python to record text information, such as name. Strings in Python are actually a *sequence*, which basically means Python keeps track of every element in the string as a sequence. For example, Python understands the string "hello' to be a sequence of letters in a specific order. This means we will be able to use indexing to grab particular letters (like the first letter, or the last letter).
#
# This idea of a sequence is an important one in Python and we will touch upon it later on in the future.
#
# In this lecture we'll learn about the following:
#
# 1.) Creating Strings
# 2.) Printing Strings
# 3.) String Indexing and Slicing
# 4.) String Properties
# 5.) String Methods
# 6.) Print Formatting
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Creating a String
# ***
# To create a string in Python you need to use either single quotes or double quotes. For example:
# + slideshow={"slide_type": "fragment"}
# Single word
print('hello')
print() # Used to have a line space between two sentences. Try deleting this line & seeing the difference.
# Entire phrase
print('This is also a string')
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Concept-Alert" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Variables : Store your Value in me!
# ***
#
# In the code below we begin to explore how we can use a variable to which a string can be assigned. This can be extremely useful in many cases, where you can call the variable instead of typing the string everytime. This not only makes our code clean but it also makes it less redundant.
# Example syntax to assign a value or expression to a variable,
#
# variable_name = value or expression
#
# Now let's get coding!!. With the below block of code showing how to assign a string to variable.
#
#
# + slideshow={"slide_type": "subslide"}
s = 'New York'
print(s)
print(type(s))
print(len(s)) # what's the string length
# -
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### String Indexing
# ***
# We know strings are a sequence, which means Python can use indexes to call parts of the sequence. Let's learn how this works.
#
# In Python, we use brackets [] after an object to call its index. We should also note that indexing starts at 0 for Python. Let's create a new object called s and the walk through a few examples of indexing.
# Assign s as a string
s = '<NAME>'
# +
# Print the object
print(s)
print()
# Show first element (in this case a letter)
print(s[0])
print()
# Show the second element (also a letter)
print(s[1])
# -
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## String Concatenation and Repetition
#
# ***
# **String Concatenation** is a process to combine two strings. It is done using the '+' operator.
#
# **String Repetition** is a process of repeating a same string multiple times
#
# The examples of the above concepts is as follows.
# + slideshow={"slide_type": "fragment"}
# concatenation (addition)
s1 = 'Hello'
s2 = "World"
print(s1 + " " + s2)
# + slideshow={"slide_type": "fragment"}
# repetition (multiplication)
print("Hello_" * 3)
print("-" * 10)
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## String Slicing & Indexing
# ***
# **String Indexing** is used to to select the letter at a particular index/position.
#
# **String Slicing** is a process to select a subset of an entire string
#
# The examples of the above stated are as follows
# + slideshow={"slide_type": "subslide"}
s = "Namaste World"
# print sub strings
print(s[1]) #This is indexing.
print(s[6:11]) #This is known as slicing.
print(s[-5:-1])
# test substring membership
print("Wor" in s)
# -
# Note the above slicing. Here we're telling Python to grab everything from 6 up to 10 and from fifth last to second last. You'll notice this a lot in Python, where statements and are usually in the context of "up to, but not including".
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Basic Built-in String methods
#
# ***
# Objects in Python usually have built-in methods. These methods are functions inside the object (we will learn about these in much more depth later) that can perform actions or commands on the object itself.
#
# We call methods with a period and then the method name. Methods are in the form:
#
# object.method(parameters)
#
# Where parameters are extra arguments we can pass into the method. Don't worry if the details don't make 100% sense right now. Later on we will be creating our own objects and functions!
#
# Here are some examples of built-in methods in strings:
# + slideshow={"slide_type": "fragment"}
s = "Hello World"
print(s.upper()) ## Convert all the element of the string to Upper case..!!
print(s.lower()) ## Convert all the element of the string to Lower case..!!
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Print Formatting
#
# We can use the .format() method to add formatted objects to printed string statements.
#
# The easiest way to show this is through an example:
# + slideshow={"slide_type": "fragment"}
name = "Einstein"
age = 22
married = True
print("My name is %s, my age is %s, and it is %s that I am married" % (name, age, married))
print("My name is {}, my age is {}, and it is {} that I am married".format(name, age, married))
# + [markdown] slideshow={"slide_type": "skip"}
# My Practise
# +
s = 'a quick brown fox'
print(type(s), s)
# +
s_lead = ' a quick brown fox'
s_trail = 'a quick brown fox '
s_both = ' a quick brown fox '
print(s_lead.lstrip(), type(s_lead), s_lead)
print(s_trail.rstrip(), type(s_lead))
print(s_both.strip(), type(s_lead))
# -
s_1 = '4$564%%'
print(s_1.rstrip('%'))
s2 = '$4%'
print(s2.lstrip('$'))
'abcddcbaabcd'.replace('abcd', '', 1)
'abcddcbaabcd'.replace('abcd', '', 2)
'abcddcbaabcdvamabcdefghabcdwxyz'.replace('abcd', '%')
# <img src="../images/icon/Concept-Alert.png" alt="Concept-Alert" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Numbers
# ***
#
# Having worked with string we will turn our attention to numbers
# We'll learn about the following topics:
#
# 1.) Types of Numbers in Python
# 2.) Basic Arithmetic
# 3.) Object Assignment in Python
# <img src="../images/icon/Concept-Alert.png" alt="Concept-Alert" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Types of numbers
# ***
# Python has various "types" of numbers (numeric literals). We'll mainly focus on integers and floating point numbers.
#
# Integers are just whole numbers, positive or negative. For example: 2 and -2 are examples of integers.
#
# Floating point numbers in Python are notable because they have a decimal point in them, or use an exponential (e) to define the number. For example 2.0 and -2.1 are examples of floating point numbers. 4E2 (4 times 10 to the power of 2) is also an example of a floating point number in Python.
#
# Throughout this course we will be mainly working with integers or simple float number types.
#
# Here is a table of the two main types we will spend most of our time working with some examples:
#
# <table>
# <tr>
# <th>Examples</th>
# <th>Number "Type"</th>
# </tr>
#
# <tr>
# <td>1,2,-5,1000</td>
# <td>Integers</td>
# </tr>
#
# <tr>
# <td>1.2,-0.5,2e2,3E2</td>
# <td>Floating-point numbers</td>
# </tr>
# </table>
# Now let's start with some basic arithmetic.
# + [markdown] slideshow={"slide_type": "-"}
# ## Basic Arithmetic
# +
# Addition
print(2+1)
# Subtraction
print(2-1)
# Multiplication
print(2*2)
# Division
print(3/2)
# -
# ## Arithmetic continued
# Powers
2**3
# Order of Operations followed in Python
2 + 10 * 10 + 3
# Can use parenthesis to specify orders
(2+10) * (10+3)
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Variable Assignments
# ***
# Now that we've seen how to use numbers in Python as a calculator let's see how we can assign names and create variables.
#
# We use a single equals sign to assign labels to variables. Let's see a few examples of how we can do this.
# Let's create an object called "a" and assign it the number 5
a = 5
# Now if I call *a* in my Python script, Python will treat it as the number 5.
# Adding the objects
a+a
# What happens on reassignment? Will Python let us write it over?
# Reassignment
a = 10
# Check
a
# +
a = 5
print(a)
print(type(a))
# +
b = 10.5
print(b)
print(type(b))
# -
c = b - a
print(c)
print(type(c))
d = b ** a
print(d, type(d))
e = b * a
print(e, type(e))
f = b/a
print(f, type(f))
g = b//a
print(g, type(g))
# <img src="../images/icon/ppt-icons.png" alt="ppt-icons" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Mini Challenge - 1
# ***
#
# Its your turn now!! store the word `hello` in my_string. print the my_string + name.
# +
my_string = 'hello'
my_name = input('Enter your name')
print(my_string + ' ' + my_name)
# -
# <img src="../images/icon/ppt-icons.png" alt="ppt-icons" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Mini Challenge - 2
# ***
#
# **Its your turn now!!!** given the numbers stored in variables `a` and `b`. Can you write a simple code to compute the mean of these two numbers and assign it to a variable `mean`.
# +
a = int(input('Enter Value - '))
b = int(input('Enter another Value - '))
mean = (a+b)/2
print(mean)
# +
a = int(input('Enter Value - '))
b = int(input('Enter another Value - '))
mean = (a+b)/2
print(mean)
# -
# <img src="../images/icon/Pratical-Tip.png" alt="Pratical-Tip" style="width: 100px;float:left; margin-right:15px"/>
# <br />
# The names you use when creating these labels need to follow a few rules:
#
# 1. Names can not start with a number.
# 2. There can be no spaces in the name, use _ instead.
# 3. Can't use any of these symbols :'",<>/?|\()!@#$%^&*~-+
#
#
# Using variable names can be a very useful way to keep track of different variables in Python. For example:
# ## From Sales to Data Science
# ***
# Discover the story of <NAME> who made a successful transition from Sales to Data Science. Making a successful switch to Data Science is a game of Decision and Determenination. But it's a long road from Decision to Determination. To read more, click <a href="https://greyatom.com/blog/2018/03/career-transition-decision-to-determination/">here</a>
# # CHAPTER - 2 : Data Types & Data Structures
# ***
# - Everything in Python is an "object", including integers/floats
# - Most common and important types (classes)
# - "Single value": None, int, float, bool, str, complex
# - "Multiple values": list, tuple, set, dict
#
#
# - Single/Multiple isn't a real distinction, this is for explanation
# - There are many others, but these are most frequently used
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Identifying Data Types
#
# + slideshow={"slide_type": "fragment"}
a = 42
b = 32.30
print(type(a))#gets type of a
print(type(b))#gets type of b
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Single Value Types
# ***
# - int: Integers
# - float: Floating point numbers
# - bool: Boolean values (True, False)
# - complex: Complex numbers
# - str: String
# -
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Lists
# ***
# Lists can be thought of the most general version of a *sequence* in Python. Unlike strings, they are mutable, meaning the elements inside a list can be changed!
#
# In this section we will learn about:
#
# 1.) Creating lists
# 2.) Indexing and Slicing Lists
# 3.) Basic List Methods
# 4.) Nesting Lists
# 5.) Introduction to List Comprehensions
#
# Lists are constructed with brackets [] and commas separating every element in the list.
#
# Let's go ahead and see how we can construct lists!
# Assign a list to an variable named my_list
my_list = [1,2,3]
# We just created a list of integers, but lists can actually hold different object types. For example:
my_list = ['A string',23,100.232,'o']
# Just like strings, the len() function will tell you how many items are in the sequence of the list.
len(my_list)
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Adding New Elements to a list
# ***
# We use two special commands to add new elements to a list. Let's make a new list to remind ourselves of how this works:
my_list = ['one','two','three',4,5]
# + slideshow={"slide_type": "fragment"}
# append a value to the end of the list
l = [1, 2.3, ['a', 'b'], 'New York']
l.append(3.1)
print(l)
# + slideshow={"slide_type": "fragment"}
# extend a list with another list.
l = [1, 2, 3]
l.extend([4, 5, 6])
print(l)
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Slicing
# ***
# Slicing is used to access individual elements or a rage of elements in a list.
#
# Python supports "slicing" indexable sequences. The syntax for slicing lists is:
#
# - `list_object[start:end:step]` or
# - `list_object[start:end]`
#
# start and end are indices (start inclusive, end exclusive). All slicing values are optional.
# + slideshow={"slide_type": "fragment"}
lst = list(range(10)) # create a list containing 10 numbers starting from 0
print(lst)
print("elements from index 4 to 7:", lst[4:7])
print("alternate elements, starting at index 0:", lst[0::2]) # prints elements from index 0 till last index with a step of 2
print("every third element, starting at index 1:", lst[1::3]) # prints elements from index 1 till last index with a step of 3
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-block alert-success">**Other `list` operations**</div>
#
# ***
# - **`.append`**: add element to end of list
# - **`.insert`**: insert element at given index
# - **`.extend`**: extend one list with another list
# +
print(my_list)
my_list.append('a')
print(my_list)
popper = my_list.pop()
print('popped element - {0}'.format(popper))
print(my_list)
my_list.insert(3, "three1")
print(my_list)
my_list.append("oneMore")
print(my_list)
popper1 = my_list.pop()
popper2 = my_list.pop()
print(popper1, ' - ', popper2)
# -
# # Did you know?
#
# **Did you know that Japanese Anime Naruto is related to Data Science. Find out how**
#
# <img src="https://greyatom.com/blog/wp-content/uploads/2017/06/naruto-1-701x321.png">
#
#
# Find out here https://medium.com/greyatom/naruto-and-data-science-how-data-science-is-an-art-and-data-scientist-an-artist-c5f16a68d670
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# # Dictionaries
# ***
# Now we're going to switch gears and learn about *mappings* called *dictionaries* in Python. If you're familiar with other languages you can think of these Dictionaries as hash tables.
#
# This section will serve as a brief introduction to dictionaries and consist of:
#
# 1.) Constructing a Dictionary
# 2.) Accessing objects from a dictionary
# 3.) Nesting Dictionaries
# 4.) Basic Dictionary Methods
#
# A Python dictionary consists of a key and then an associated value. That value can be almost any Python object.
# ## Constructing a Dictionary
# ***
# Let's see how we can construct dictionaries to get a better understanding of how they work!
# Make a dictionary with {} and : to signify a key and a value
my_dict = {'key1':'value1','key2':'value2'}
# Call values by their key
my_dict['key2']
# We can effect the values of a key as well. For instance:
my_dict['key1']=123
my_dict
# Subtract 123 from the value
my_dict['key1'] = my_dict['key1'] - 123
#Check
my_dict['key1']
# A quick note, Python has a built-in method of doing a self subtraction or addition (or multiplication or division). We could have also used += or -= for the above statement. For example:
# Set the object equal to itself minus 123
my_dict['key1'] -= 123
my_dict['key1']
# Now its your turn to get hands-on with Dictionary, create a empty dicts. Create a new key calle animal and assign a value 'Dog' to it..
#
# Create a new dictionary
d = {}
# Create a new key through assignment
d['animal'] = 'Dog'
# +
dict1 = {}
print(type(dict1))
# -
set1 = {'a', 'b', 'c'}
print(type(set1))
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# # Set and Booleans
# ***
# There are two other object types in Python that we should quickly cover. Sets and Booleans.
#
# ## Sets
# Sets are an unordered collection of *unique* elements. We can construct them by using the set() function. Let's go ahead and make a set to see how it works
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Set Theory
# <img src="../images/sets2.png" width="60%"/>
# +
x = set()
# We add to sets with the add() method
x.add(1)
#Show
x
# -
# Note the curly brackets. This does not indicate a dictionary! Although you can draw analogies as a set being a dictionary with only keys.
#
# We know that a set has only unique entries. So what happens when we try to add something that is already in a set?
# +
# Add a different element
x.add(2)
#Show
x
# +
# Try to add the same element
x.add(1)
#Show
x
# -
# Notice how it won't place another 1 there. That's because a set is only concerned with unique elements! We can cast a list with multiple repeat elements to a set to get the unique elements. For example:
# Create a list with repeats
l = [1,1,2,2,3,4,5,6,1,1]
# Cast as set to get unique values
set(l)
set1 = set('abracadabra')
print(set1)
set2 = set('zipzabzoo')
print(set2)
set1 and set2
set1 - set2
set1 | set2
set1 ^ set2
# <img src="../images/icon/ppt-icons.png" alt="ppt-icons" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Mini Challenge - 3
# ***
# Can you access the last element of a l which is a list and find the last element of that list.
l[-1]
# + [markdown] slideshow={"slide_type": "subslide"}
# # CHAPTER - 3 : Python Programming Constructs
# ***
# We'll be talking about
# - Looping
# - Conditional Statements
# - Comprehensions
# -
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
# ## Loops and Iterative Statements
# ## If,elif,else Statements
# ***
# if Statements in Python allows us to tell the computer to perform alternative actions based on a certain set of results.
#
# Verbally, we can imagine we are telling the computer:
#
# "Hey if this case happens, perform some action"
#
# We can then expand the idea further with elif and else statements, which allow us to tell the computer:
#
# "Hey if this case happens, perform some action. Else if another case happens, perform some other action. Else-- none of the above cases happened, perform this action"
#
# Let's go ahead and look at the syntax format for if statements to get a better idea of this:
#
# if case1:
# perform action1
# elif case2:
# perform action2
# else:
# perform action 3
# + slideshow={"slide_type": "fragment"}
a = 5
b = 4
if a > b:
# we are inside the if block
print("a is greater than b")
elif b > a:
# we are inside the elif block
print("b is greater than a")
else:
# we are inside the else block
print("a and b are equal")
# Note: Python doesn't have a switch statement
# -
# <img src="../images/icon/Warning.png" alt="Warning" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Indentation
# ***
# It is important to keep a good understanding of how indentation works in Python to maintain the structure and order of your code. We will touch on this topic again when we start building out functions!
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# # For Loops
# ***
# A **for** loop acts as an iterator in Python, it goes through items that are in a *sequence* or any other iterable item. Objects that we've learned about that we can iterate over include strings,lists,tuples, and even built in iterables for dictionaries, such as the keys or values.
#
# We've already seen the **for** statement a little bit in past lectures but now lets formalize our understanding.
#
# Here's the general format for a **for** loop in Python:
#
# for item in object:
# statements to do stuff
#
# The variable name used for the item is completely up to the coder, so use your best judgment for choosing a name that makes sense and you will be able to understand when revisiting your code. This item name can then be referenced inside you loop, for example if you wanted to use if statements to perform checks.
#
# Let's go ahead and work through several example of **for** loops using a variety of data object types.
#
# +
#Simple program to find the even numbers in a list
list_1 = [2,4,5,6,8,7,9,10] # Initialised the list
for number in list_1: # Selects one element in list_1
if number % 2 == 0: # Checks if it is even. IF even, only then, goes to next step else performs above step and continues iteration
print(number,end=' ') # prints no if even. end=' ' prints the nos on the same line with a space in between. Try deleting this command & seeing the difference.
# +
lst1 = [4, 7, 13, 11, 3, 11, 15]
lst2 = []
for index, e in enumerate(lst1):
if e == 10:
break
if e < 10:
continue
lst2.append((index, e*e))
else:
print("out of loop without using break statement")
lst2
# -
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# # While loops
# ***
# The **while** statement in Python is one of most general ways to perform iteration. A **while** statement will repeatedly execute a single statement or group of statements as long as the condition is true. The reason it is called a 'loop' is because the code statements are looped through over and over again until the condition is no longer met.
#
# The general format of a while loop is:
#
# while test:
# code statement
# else:
# final code statements
#
# Let’s look at a few simple while loops in action.
#
# +
x = 0
while x < 10:
print ('x is currently: ',x,end=' ') #end=' ' to put print below statement on the same line after thsi statement
print (' x is still less than 10, adding 1 to x')
x+=1
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Comprehensions
# ***
# - Python provides syntactic sugar to write small loops to generate lists/sets/tuples/dicts in one line
# - These are called comprehensions, and can greatly increase development speed and readability
#
# Syntax:
# ```
# sequence = [expression(element) for element in iterable if condition]
# ```
#
# The brackets used for creating the comprehension define what type of object is created.
#
# Use **[ ]** for lists, **()** for _generators_, **{}** for sets and dicts
# + [markdown] slideshow={"slide_type": "subslide"}
# ### `list` Comprehension
# + slideshow={"slide_type": "fragment"}
names = ["Ravi", "Pooja", "Vijay", "Kiran"]
hello = ["Hello " + name for name in names]
print(hello)
# + slideshow={"slide_type": "fragment"}
numbers = [55, 32, 87, 99, 10, 54, 32]
even = [num for num in numbers if num % 2 == 0]
print(even)
odd_squares = [(num, num * num) for num in numbers if num % 2 == 1]
print(odd_squares)
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Technical-Stuff" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Exception Handling
# ***
# #### try and except
#
# The basic terminology and syntax used to handle errors in Python is the **try** and **except** statements. The code which can cause an exception to occue is put in the *try* block and the handling of the exception is the implemented in the *except* block of code. The syntax form is:
#
# try:
# You do your operations here...
# ...
# except ExceptionI:
# If there is ExceptionI, then execute this block.
# except ExceptionII:
# If there is ExceptionII, then execute this block.
# ...
# else:
# If there is no exception then execute this block.
#
# We can also just check for any exception with just using except: To get a better understanding of all this lets check out an example: We will look at some code that opens and writes a file:
# + slideshow={"slide_type": "fragment"}
try:
x = 1 / 0
except ZeroDivisionError:
print('divided by zero')
print('executed when exception occurs')
else:
print('executed only when exception does not occur')
finally:
print('finally block, always executed')
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="../images/icon/Concept-Alert.png" alt="Concept-Alert" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## Modules, Packages, and `import`
# ***
# A module is a collection of functions and variables that have been bundled together in a single file. Module helps us:
# - Used for code organization, packaging and reusability
# - Module: A Python file
# - Package: A folder with an ``__init__.py`` file
# - Namespace is based on file's directory path
#
# Module's are usually organised around a theme. Let's see how to use a module. To access our module we will import it using python's import statement. Math module provides access to the mathematical functions.
# + slideshow={"slide_type": "subslide"}
# import the math module
import math
# use the log10 function in the math module
math.log10(123)
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="../images/icon/Technical-Stuff.png" alt="Concept-Alert" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ## File I/O : Helps you read your files
# ***
# - Python provides a `file` object to read text/binary files.
# - This is similar to the `FileStream` object in other languages.
# - Since a `file` is a resource, it must be closed after use. This can be done manually, or using a context manager (**`with`** statement)
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-block alert-info">Create a file in the current directory</div>
# + slideshow={"slide_type": "fragment"}
with open('myfile.txt', 'w') as f:
f.write("This is my first file!\n")
f.write("Second line!\n")
f.write("Last line!\n")
# let's verify if it was really created.
# For that, let's find out which directory we're working from
import os
print(os.path.abspath(os.curdir))
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-block alert-info">Read the newly created file</div>
# + slideshow={"slide_type": "fragment"}
# read the file we just created
with open('myfile.txt', 'r') as f:
for line in f:
print(line)
# -
# <img src="../images/icon/ppt-icons.png" alt="ppt-icons" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Mini Challenge - 4
# ***
# Can you compute the square of a number assigned to a variable a using the math module?
# +
import math
a = int(input('Enter a number to compute square'))
val = math.pow(a, 2)
print(val)
# -
# <img src="../images/icon/ppt-icons.png" alt="ppt-icons" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# ### Mini Challenge - 5
# ***
# Can you create a list of 10 numbers iterate through the list and print the square of each number ?
# +
numList = list(range(1,11))
print(numList)
squareList = [(num*num) for num in numList]
print(squareList)
# + [markdown] slideshow={"slide_type": "slide"}
# # Further Reading
#
# - Official Python Documentation: https://docs.python.org/
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="../images/icon/Recap.png" alt="Recap" style="width: 100px;float:left; margin-right:15px"/>
# <br />
#
# # In-session Recap Time
# ***
# * Python Basics
# * Variables and Scoping
# * Modules, Packages and Imports
# * Data Types & Data Structures
# * Python Programming Constructs
# * Data Types & Data Structures
# * Lists
# * Dictionaries
# * Sets & Booleans
# * Python Prograamming constructs
# * Loops and Conditional Statements
# * Exception Handling
# * File I/O
# + [markdown] slideshow={"slide_type": "slide"}
# # Thank You
# ***
# ### Coming up next...
#
# - **Python Functions**: How to write modular functions to enable code reuse
# - **NumPy**: Learn the basis of most numeric computation in Python
# -
| Week-01-Git_&_Python_intro/notebooks/python_getting_started_lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
def calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch):
t=days_until_maturity/(365*time_stretch)
T=days_until_maturity/365
return y_reserves*(-(2/((1-T*APY/100)**(1/t)-1))-2)
def calc_price(x_reserves,y_reserves,total_supply,days_until_maturity,time_stretch):
t=days_until_maturity/(365*time_stretch)
return 1/((y_reserves + total_supply)/x_reserves)**t
def calc_k(in_reserves,out_reserves,t):
return pow(in_reserves,1-t) + pow(out_reserves,1-t)
def calc_apy(price,days_until_maturity):
T=days_until_maturity/365
return (1-price)/T * 100
def is_trade_valid(in_,in_reserves,out_reserves,t):
k = pow(in_reserves,1-t) + pow(out_reserves,1-t)
check = math.log(k)/math.log(in_reserves + in_) + t
return True if check >= 1 else False
def calc_max_trade(in_reserves,out_reserves,t):
k = pow(in_reserves,1-t) + pow(out_reserves,1-t)
return k**(1/(1-t)) - in_reserves
def calc_out_given_in(in_,in_reserves,out_reserves,token_out,g,t):
k=pow(in_reserves,1-t) + pow(out_reserves,1-t)
without_fee = out_reserves - pow(k-pow(in_reserves+in_,1-t),1/(1-t))
if token_out == "base":
fee = (in_-without_fee)*g
with_fee = without_fee-fee
elif token_out == "fyt":
fee = (without_fee-in_)*g
with_fee = without_fee-fee
return (with_fee,without_fee,fee)
# +
APY = 25
days_until_maturity = 90
time_stretch = 4.43
y_reserves = 100
x_reserves = calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch)
print("(APY,y_reserves,days_until_maturity,time_stretch): "+str((APY,y_reserves,days_until_maturity,time_stretch)))
print("required x_reserves: " + str(x_reserves))
total_supply=x_reserves+y_reserves
price=calc_price(x_reserves,y_reserves,total_supply,days_until_maturity,time_stretch)
print("resulting price: " + str(price))
print("check apy: " + str(calc_apy(price,days_until_maturity)))
amount=106
g=0.1
t=days_until_maturity/(365*time_stretch)
(with_fee,without_fee,fee)=calc_out_given_in(amount,y_reserves+total_supply,x_reserves,"base",g,t)
without_slippage_or_fee = price * amount
slippage = abs(without_slippage_or_fee-without_fee)
print("\nTrade size: " + str(amount))
print("Price w/o slippage: " + str(without_slippage_or_fee))
print("Price with slippage: " + str(without_fee))
print("Price with slippage and fee: " + str(with_fee))
print("Total fee: " + str(fee))
print("Percent slippage: " + str(slippage/without_slippage_or_fee * 100))
print("check apy: " + str(calc_apy(with_fee/amount,days_until_maturity)))
print("x_reserves = {:} vs price_with_slippage_fee = {:}".format(x_reserves, with_fee))
print("is valid trade = {:}".format(is_trade_valid(amount,y_reserves+total_supply,x_reserves,t)))
print("max valid trade = {:}".format(calc_max_trade(y_reserves+total_supply,x_reserves,t)))
amount=95
g=0.1
t=days_until_maturity/(365*time_stretch)
(with_fee,without_fee,fee)=calc_out_given_in(amount,x_reserves,y_reserves+total_supply,"fyt",g,t)
price=calc_price(x_reserves+amount,y_reserves-with_fee,total_supply,days_until_maturity,time_stretch)
without_slippage_or_fee = price * amount
slippage = abs(without_slippage_or_fee-without_fee)
print("\nTrade size: " + str(amount))
print("Price w/o slippage: " + str(without_slippage_or_fee))
print("Price with slippage: " + str(without_fee))
print("Price with slippage and fee: " + str(with_fee))
print("Total fee: " + str(fee))
print("Percent slippage: " + str(slippage/without_slippage_or_fee * 100))
print("check apy: " + str(calc_apy(price,days_until_maturity)))
print("x_reserves = {:} vs price_with_slippage_fee = {:}".format(x_reserves, with_fee))
print("is valid trade = {:}".format(is_trade_valid(amount,y_reserves+total_supply,x_reserves,t)))
# -
APY = 1
tranche_length = 30
days_until_maturity = tranche_length
time_stretch = 5
y_reserves = 50
x_reserves = calc_x_reserves(APY,y_reserves,tranche_length,time_stretch)
total_supply=x_reserves+y_reserves
print("required x_reserves: " + str(x_reserves))
g=0.05
t=days_until_maturity/(365*time_stretch)
k=calc_k(y_reserves+total_supply,x_reserves,t)
print("k: " + str(k))
print("k^(1/(1-t)): " + str(k**(1/(1-t))))
max_trade = calc_max_trade(y_reserves+total_supply,x_reserves,t) - .0001
print("max valid trade = {:}".format(max_trade))
(with_fee,without_fee,fee)=calc_out_given_in(max_trade,y_reserves+total_supply,x_reserves,"base",g,t)
print("Total Price after max trade: " + str(without_fee))
unit_price = without_fee/max_trade
print("Unit Price after max trade: " + str(unit_price))
resulting_apy=calc_apy(unit_price,days_until_maturity)
print("resulting apy: " + str(resulting_apy))
(resulting_apy-APY)/APY * 100
# +
import matplotlib.pyplot as plt
import numpy as np
fig, ax = plt.subplots(figsize=(12,12))
time_stretch=1
for tranche_length in [30,60,120]:
apy_data = []
x_reserves_data = []
for APY in np.arange(1, 51, 1):
days_until_maturity = tranche_length
y_reserves=100
x_reserves = calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch)
apy_data.append(APY)
x_reserves_data.append(x_reserves/y_reserves)
x=apy_data
y=x_reserves_data
ax.set_xlabel("PT APY",fontsize=18)
ax.set_ylabel("Base Reserves to PT Reserves Ratio",fontsize=18)
ax.set_xticks(np.arange(0, 51, 1))
ax.set_ylim(0,50)
ax.set_yticks(np.arange(0, 51, 5))
#ax.set_yticks(np.arange(0, 105, 5))
ax.plot(x, y,label=tranche_length)
ax.legend(title="Term Length")
plt.title("Reserves Ratio Vs PT APY",fontsize=20 )
plt.grid(b=True, which='major', color='#666666', linestyle='-',alpha=0.2)
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
fig, ax = plt.subplots(figsize=(12,12))
for time_stretch in [1,3,5,10,20,30,40]:
apy_data = []
x_reserves_data = []
for APY in np.arange(1, 51, 1):
days_until_maturity = 30
y_reserves=100
x_reserves = calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch)
apy_data.append(APY)
x_reserves_data.append(x_reserves/y_reserves)
x=apy_data
y=x_reserves_data
ax.set_xlabel("PT APY",fontsize=18)
ax.set_ylabel("Base Reserves to PT Reserves Ratio",fontsize=18)
ax.set_xticks(np.arange(0, 51, 1))
ax.set_ylim(0,10)
ax.set_yticks(np.arange(0, 11, 1))
ax.plot(x, y,label=time_stretch)
ax.legend(title="Time Stretch")
plt.title("Reserves Ratio Vs PT APY",fontsize=20 )
plt.grid(b=True, which='major', color='#666666', linestyle='-',alpha=0.2)
plt.show()
# +
import numpy as np
import matplotlib.pyplot as plt
APY = 20
days_until_maturity = 90
time_stretch = 5
y_reserves = 100
x_reserves = calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch)
total_supply=x_reserves+y_reserves
epsilon=.0000001
times = []
max_trade_amounts = []
for t in np.arange(328, 0, -.1):
times.append(t/365)
max_trade = calc_max_trade(y_reserves+total_supply,x_reserves,t/365)-epsilon
max_trade_amounts.append(max_trade)
fig, ax = plt.subplots(figsize=(12,12))
x=times
y=max_trade_amounts
ax.set_title("Max Allowable Input FYT Trade Size vs t-Param\n(base_reserves,fyt_reserves)=({:.2f},{:.2f})".format(x_reserves,y_reserves),fontsize=18)
ax.set_xlabel("t",fontsize=18)
ax.set_ylabel("Max Input FYT Trade Size",fontsize=18)
ax.invert_xaxis()
ax.plot(x, y)
# +
import matplotlib.pyplot as plt
import numpy as np
APY = 20
days_until_maturity = 90
time_stretch = 5
y_reserves = 100
x_reserves = calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch)
total_supply=x_reserves+y_reserves
epsilon=.0000001
g=.1
times = []
days_until_maturity_list = []
max_trade_amounts = []
resulting_apys = []
for t in np.arange(days_until_maturity, 0, -.1):
times.append(t/(365*time_stretch))
days_until_maturity_list.append(t)
max_trade = calc_max_trade(y_reserves+total_supply,x_reserves,t/(365*time_stretch))-epsilon
max_trade_amounts.append(max_trade)
(with_fee,without_fee,fee)=calc_out_given_in(max_trade,y_reserves+total_supply,x_reserves,"base",g,t/(365*time_stretch))
resulting_apy=calc_apy(without_fee/max_trade,t)
resulting_apys.append((resulting_apy-APY)/APY * 100)
fig, ax1 = plt.subplots(figsize=(12,12))
ax2 = ax1.twinx()
x = days_until_maturity_list
y1 = max_trade_amounts
ax1.set_title("Days Until Maturity vs Max Allowable Input FYT Trade Size vs Max % Change in Implied APY\n\n(base_reserves, fyt_reserves)=({:.2f}, {:.2f})\ntime_stretch={:}".format(x_reserves,y_reserves,time_stretch),fontsize=18)
ax1.set_xlabel("Days Until Maturity",fontsize=18)
ax1.set_ylabel("Max Input FYT Trade Size",fontsize=18,color='b')
ax1.invert_xaxis()
ax1.plot(x, y1,'b-')
y2 = resulting_apys
ax2.set_ylabel("Max % Change in Implied APY",fontsize=18,color='g')
ax2.plot(x, y2,'g-')
# +
import matplotlib.pyplot as plt
import numpy as np
for time_stretch in [20,30,40]:
fig, ax1 = plt.subplots(figsize=(12,12))
APY = 1.7
days_until_maturity = 180
y_reserves = 50
x_reserves = calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch)
total_supply=x_reserves+y_reserves
epsilon=.0000001
g=.1
times = []
days_until_maturity_list = []
max_trade_amounts = []
resulting_apys = []
for t in np.arange(days_until_maturity, 0, -.1):
times.append(t/(365*time_stretch))
days_until_maturity_list.append(t)
max_trade = calc_max_trade(y_reserves+total_supply,x_reserves,t/(365*time_stretch))-epsilon
max_trade_amounts.append(max_trade)
(with_fee,without_fee,fee)=calc_out_given_in(max_trade,y_reserves+total_supply,x_reserves,"base",g,t/(365*time_stretch))
resulting_apy=calc_apy(without_fee/max_trade,t)
resulting_apys.append((resulting_apy-APY)/APY * 100)
ax2 = ax1.twinx()
x = days_until_maturity_list
y1 = max_trade_amounts
ax1.set_title("Days Until Maturity vs Max Allowable Input FYT Trade Size vs Max % Change in Implied APY\n\nAPY={:.2f}%\n(base_reserves, fyt_reserves)=({:.2f}, {:.2f})\ntime_stretch={:}".format(APY,x_reserves,y_reserves,time_stretch),fontsize=18)
ax1.set_xlabel("Days Until Maturity",fontsize=18)
ax1.set_ylabel("Max Input FYT Trade Size",fontsize=18,color='b')
ax1.invert_xaxis()
y2 = resulting_apys
ax2.set_ylabel("Max % Change in Implied APY",fontsize=18,color='g')
if time_stretch == 20:
ax1.plot(x, y1,'b-')
ax2.plot(x, y2,'g-')
elif time_stretch == 30:
ax1.plot(x, y1,'b--')
ax2.plot(x, y2,'g--')
elif time_stretch == 40:
ax1.plot(x, y1,'b.')
ax2.plot(x, y2,'g.')
# +
import matplotlib.pyplot as plt
import numpy as np
fig, ax1 = plt.subplots(figsize=(12,12))
min_apy_changes = []
max_apy_changes = []
time_stretches = []
for time_stretch in np.arange(1, 51, 1):
APY = 1.7
days_until_maturity = 180
y_reserves = 50
x_reserves = calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch)
total_supply=x_reserves
epsilon=.0000001
g=.1
times = []
days_until_maturity_list = []
max_trade_amounts = []
resulting_apys = []
for t in np.arange(days_until_maturity, 0, -.1):
times.append(t/(365*time_stretch))
days_until_maturity_list.append(t)
max_trade = calc_max_trade(y_reserves+total_supply,x_reserves,t/(365*time_stretch))-epsilon
max_trade_amounts.append(max_trade)
(with_fee,without_fee,fee)=calc_out_given_in(max_trade,y_reserves+total_supply,x_reserves,"base",g,t/(365*time_stretch))
resulting_apy=calc_apy(without_fee/max_trade,t)
resulting_apys.append((resulting_apy-APY)/APY * 100)
time_stretches.append(time_stretch)
min_apy_changes.append(min(resulting_apys))
max_apy_changes.append(max(resulting_apys))
ax1.scatter(time_stretches,min_apy_changes,color='g')
ax1.scatter(time_stretches,max_apy_changes,color='r')
#ax1.set_ylim(0,1000)
# this plot tells me that the difference between the min and max % change apy isnt very much
# -
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
cols = ["apy","tranche_length","time_stretch",\
"x_reserves", "y_reserves",\
"min_k","max_k",\
"min_trade_amount","max_trade_amount",\
"min_resulting_fyt_unit_prices","max_resulting_fyt_unit_prices",\
"min_resulting_apy","max_resulting_apy",\
"min_apy_change","max_apy_change"]
tests = []
g=.1
for y_reserves in np.arange(50, 1010, 50):
for tranche_length in np.arange(30, 181, 30):
for APY in np.arange(1, 51, 1):
for time_stretch in np.arange(1, 40.1, .1):
days_until_maturity = tranche_length
x_reserves = calc_x_reserves(APY,y_reserves,days_until_maturity,time_stretch)
total_supply=x_reserves+y_reserves
epsilon=.0000001
times = []
days_until_maturity_list = []
ks = []
max_trade_amounts = []
resulting_fyt_unit_prices = []
resulting_apys = []
resulting_apy_changes = []
for day in np.arange(days_until_maturity, 0, -1):
times.append(day/(365*time_stretch))
days_until_maturity_list.append(day)
k=calc_k(y_reserves+total_supply,x_reserves,day/(365*time_stretch))
ks.append(k)
max_trade = calc_max_trade(y_reserves+total_supply,x_reserves,day/(365*time_stretch))-epsilon
max_trade_amounts.append(max_trade)
(with_fee,without_fee,fee)=calc_out_given_in(max_trade,y_reserves+total_supply,x_reserves,"base",g,day/(365*time_stretch))
resulting_fyt_unit_price = without_fee/max_trade
resulting_fyt_unit_prices.append(resulting_fyt_unit_price)
resulting_apy=calc_apy(resulting_fyt_unit_price,day)
resulting_apys.append(resulting_apy)
resulting_apy_changes.append((resulting_apy-APY)/APY * 100)
test = [APY,tranche_length,time_stretch,x_reserves,y_reserves,\
min(ks),max(ks),\
min(max_trade_amounts),max(max_trade_amounts),\
min(resulting_fyt_unit_prices),max(resulting_fyt_unit_prices),\
min(resulting_apys), max(resulting_apys),\
min(resulting_apy_changes),max(resulting_apy_changes)]
tests.append(test)
df = pd.DataFrame(tests,columns=cols)
#df.to_csv('apy_change.csv')
df
# -
df.to_csv('apy_change_new.csv')
import pandas as pd
df = pd.read_csv('apy_change_new.csv')
df = df.drop(columns=['Unnamed: 0']).reset_index()
cols = df.columns
for col in cols:
df[col] = df[col].astype(float)
df.head()
# +
import numpy as np
#df.to_csv('apy_change.csv')
pd.set_option('display.max_rows', None)
df['reserve_ratio']=df['x_reserves']/df['y_reserves']
df['time_stretch'] = round(df['time_stretch'],5)
reserve_ratio_filter=(df['reserve_ratio']>=.1)&(df['reserve_ratio']<=1)
price_discovery_filter=(df['max_apy_change']>=50)
df_filtered = df[reserve_ratio_filter & price_discovery_filter].reset_index()
apy_t_stretches=[]
for APY in np.arange(3, 51, 1):
min_ts=df_filtered[df_filtered['apy']==APY]['time_stretch'].min()
max_ts=df_filtered[df_filtered['apy']==APY]['time_stretch'].max()
min_tl=df_filtered[(df_filtered['apy']==APY)&(df_filtered['time_stretch']>=min_ts)&(df_filtered['time_stretch']<=max_ts)]['tranche_length'].min()
max_tl=df_filtered[(df_filtered['apy']==APY)&(df_filtered['time_stretch']>=min_ts)&(df_filtered['time_stretch']<=max_ts)]['tranche_length'].max()
apy_t_stretches.append((APY,min_ts,max_ts,min_tl,max_tl))
# +
import numpy as np
pd.reset_option("display.max_rows")
df['reserve_ratio']=df['x_reserves']/df['y_reserves']
df['spot_price']=(df['x_reserves']/(2*df['y_reserves']+df['x_reserves']))**(df['tranche_length']/(365*df['time_stretch']))
df['spot_apy']=(1-df['spot_price'])/(df['tranche_length']/365) * 100
# we want to target the time stretch that allows the reserve ratio to be equal to the spot price
reserve_ratio_filter=(df['reserve_ratio']-df['reserve_ratio']*0.05<=df['spot_price'])&(df['spot_price']<=df['reserve_ratio']+df['reserve_ratio']*0.05)&(df['reserve_ratio']<=1)
df_filtered = df[reserve_ratio_filter].reset_index()
apy_t_stretches=[]
for APY in np.arange(3, 51, 1):
min_ts=df_filtered[df_filtered['apy']==APY]['time_stretch'].min()
max_ts=df_filtered[df_filtered['apy']==APY]['time_stretch'].max()
apy_t_stretches.append((APY,min_ts,max_ts))
df_filtered
# +
#df['time_stretch'] = round(df['time_stretch'],1)
#df[(df['apy']==9)&(df['tranche_length']==90)&(df['time_stretch']==9)]
#df_filtered[(df_filtered['apy']==10)]
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
apys = [apy for apy,min_ts,max_ts in apy_t_stretches]
mean_tss= [(min_ts+max_ts)/2 for apy,min_ts,max_ts in apy_t_stretches]
err_tss= [max_ts-(min_ts+max_ts)/2 for apy,min_ts,max_ts in apy_t_stretches]
plt.yticks(np.arange(0,41, 1))
plt.xticks(np.arange(0,51, 1))
#plt.errorbar(apys, mean_tss, yerr=err_tss, fmt='o', color='black',
# ecolor='red', elinewidth=3, capsize=0);
plt.scatter(apys,mean_tss,color='black')
plt.grid(True)
plt.title('Suggested Time Stretch vs PT APY', fontsize=14)
plt.xlabel('PT APY', fontsize=14)
plt.ylabel('Time Stretch', fontsize=14)
# -
apy_t_stretches
# +
from scipy.optimize import curve_fit
def objective(x,a,b):
return a/(b*x)
x = apys
y = mean_tss
# curve fit
popt, _ = curve_fit(objective, x, y)
# summarize the parameter values
a, b = popt
print('y = %.5f /( %.5f * x)' % (a, b))
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
plt.yticks(np.arange(0,41, 1))
plt.xticks(np.arange(0,51, 1))
#plt.errorbar(apys, mean_tss, yerr=err_tss, fmt='o', color='black',
# ecolor='red', elinewidth=3, capsize=0);
plt.scatter(apys,mean_tss,color='black')
plt.grid(True)
plt.title('Suggested Time Stretch vs PT APY', fontsize=14)
plt.xlabel('PT APY', fontsize=14)
plt.ylabel('Time Stretch', fontsize=14)
x = np.arange(3,51,1)
y = 3.09396 /( 0.02789 * x)
plt.plot(x, y, '--', color="green")
# -
APY = 1
tranche_length = 30
days_until_maturity = 1
time_stretch = 5
y_reserves = 50
x_reserves = calc_x_reserves(APY,y_reserves,tranche_length,time_stretch)
total_supply=x_reserves
print("required x_reserves: " + str(x_reserves))
g=0.05
t=days_until_maturity/(365*time_stretch)
k=calc_k(y_reserves+total_supply,x_reserves,t)
print("k: " + str(k))
print("k^(1/(1-t)): " + str(k**(1/(1-t))))
max_trade = calc_max_trade(y_reserves+total_supply,x_reserves,t) - .0001
print("max valid trade = {:}".format(max_trade))
(with_fee,without_fee,fee)=calc_out_given_in(max_trade,y_reserves+total_supply,x_reserves,"base",g,t)
print("Total Price after max trade: " + str(without_fee))
unit_price = without_fee/max_trade
print("Unit Price after max trade: " + str(unit_price))
resulting_apy=calc_apy(unit_price,1)
print("resulting apy: " + str(resulting_apy))
(resulting_apy-APY)/APY * 100
# +
#df = df.drop(columns=['Unnamed: 0']).reset_index()
#df_filtered = df[(df['time_stretch']>=3)]
#df_filtered.head()
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
df_filtered = df[(df['time_stretch']==3) & (df['tranche_length']<=180)]
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], color='orange',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[(df['time_stretch']==5) & (df['tranche_length']<=180)]
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], color='green',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[(df['time_stretch']==10) & (df['tranche_length']<=180)]
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], color='yellow',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[(df['time_stretch']==20) & (df['tranche_length']<=180)]
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], color='blue',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[(df['time_stretch']==30) & (df['tranche_length']<=180)]
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], color='red',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[(df['time_stretch']==40) & (df['tranche_length']<=180)]
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], color='brown',label= df_filtered['time_stretch'].iloc[0])
plt.title('PT APY Vs Max Resulting APY', fontsize=14)
plt.xlabel('PT APY', fontsize=14)
plt.ylabel('Max Resulting APY', fontsize=14)
plt.grid(True)
plt.legend(title="Time Stretch")
x = np.arange(0,51,1)
y = x
plt.plot(x, y, '--', color="black")
plt.yticks(np.arange(0,101, 2))
plt.xticks(np.arange(0,51, 2))
plt.ylim(0,100)
plt.show()
### This tells me that yield bearing asset APY does have an affect on Max % APY Change
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
df_filtered = df[(df['time_stretch']==3) & (df['tranche_length']==30)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='<', color='orange',label=label)
df_filtered = df[(df['time_stretch']==3) & (df['tranche_length']==90)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='o', color='orange',label=label)
df_filtered = df[(df['time_stretch']==3) & (df['tranche_length']==180)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='>', color='orange',label=label)
df_filtered = df[(df['time_stretch']==5) & (df['tranche_length']==30)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='<', color='green',label=label)
df_filtered = df[(df['time_stretch']==5) & (df['tranche_length']==90)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='o', color='green',label=label)
df_filtered = df[(df['time_stretch']==5) & (df['tranche_length']==180)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='>', color='green',label=label)
df_filtered = df[(df['time_stretch']==10) & (df['tranche_length']==30)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='<', color='red',label=label)
df_filtered = df[(df['time_stretch']==10) & (df['tranche_length']==90)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='o', color='red',label=label)
df_filtered = df[(df['time_stretch']==10) & (df['tranche_length']==180)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='>', color='red',label=label)
df_filtered = df[(df['time_stretch']==20) & (df['tranche_length']==30)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='<', color='blue',label=label)
df_filtered = df[(df['time_stretch']==20) & (df['tranche_length']==90)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='o', color='blue',label=label)
df_filtered = df[(df['time_stretch']==20) & (df['tranche_length']==180)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='>', color='blue',label=label)
df_filtered = df[(df['time_stretch']==40) & (df['tranche_length']==30)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='<',color='brown',alpha=.25,label=label)
df_filtered = df[(df['time_stretch']==40) & (df['tranche_length']==90)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='o',color='brown',alpha=.25,label=label)
df_filtered = df[(df['time_stretch']==40) & (df['tranche_length']==180)]
label=(df_filtered['time_stretch'].iloc[0],df_filtered['tranche_length'].iloc[0])
plt.scatter(df_filtered['apy'], df_filtered['max_resulting_apy'], marker='>',color='brown',alpha=.25,label=label)
plt.title('PT APY Vs Max Resulting APY', fontsize=14)
plt.xlabel('PT APY', fontsize=14)
plt.ylabel('Max Resulting APY', fontsize=14)
plt.grid(True)
plt.legend(title="Time Stretch, Tranche Length")
x = np.arange(0,51,1)
y = x
plt.plot(x, y, '--', color="black")
plt.yticks(np.arange(0,101, 2))
plt.ylim(0,100)
plt.xticks(np.arange(0,51, 2))
plt.show()
### This tells me that yield bearing asset APY does have an affect on Max % APY Change
# -
x = np.arange(0,50,1)
y = x
plt.plot(x, y)
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
df_filtered = df[df['time_stretch']==5]
plt.scatter(df_filtered['apy'], df_filtered['max_apy_change'], color='green',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[df['time_stretch']==25]
plt.scatter(df_filtered['apy'], df_filtered['max_apy_change'], color='blue',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[df['time_stretch']==50]
plt.scatter(df_filtered['apy'], df_filtered['max_apy_change'], color='red',label= df_filtered['time_stretch'].iloc[0])
plt.title('APY Vs Max Percent APY Change', fontsize=14)
plt.xlabel('APY', fontsize=14)
plt.ylabel('Max Percent APY Change', fontsize=14)
plt.grid(True)
plt.legend(title="Time Stretch")
plt.show()
### This tells me that yield bearing asset APY does have an affect on Max % APY Change
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
df_filtered = df[df['apy']==1]
plt.scatter(df_filtered['time_stretch'], df_filtered['max_apy_change'], color='green',label= df_filtered['apy'].iloc[0])
df_filtered = df[df['apy']==5]
plt.scatter(df_filtered['time_stretch'], df_filtered['max_apy_change'], color='blue',label= df_filtered['apy'].iloc[0])
df_filtered = df[df['apy']==10]
plt.scatter(df_filtered['time_stretch'], df_filtered['max_apy_change'], color='red',label= df_filtered['apy'].iloc[0])
plt.title('Time Stretch Vs Max Percent APY Change', fontsize=14)
plt.xlabel('Time Stretch', fontsize=14)
plt.ylabel('Max Percent APY Change', fontsize=14)
plt.grid(True)
plt.legend(title="APY")
plt.show()
### This tells me that time stretch does have an affect on Max % APY Change
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
df_filtered = df[df['apy']==1]
plt.scatter(df_filtered['tranche_length'], df_filtered['max_apy_change'], color='green',label= df_filtered['apy'].iloc[0])
df_filtered = df[df['apy']==5]
plt.scatter(df_filtered['tranche_length'], df_filtered['max_apy_change'], color='blue', label= df_filtered['apy'].iloc[0])
df_filtered = df[df['apy']==10]
plt.scatter(df_filtered['tranche_length'], df_filtered['max_apy_change'], color='red',label= df_filtered['apy'].iloc[0])
plt.title('Tranche Length Vs Max Percent APY Change', fontsize=14)
plt.xlabel('Tranche Length', fontsize=14)
plt.ylabel('Max Percent APY Change', fontsize=14)
plt.grid(True)
plt.legend(title="APY", loc='upper right')
plt.show()
## This tells me that Tranche Length has virtually no affect on Max % APY Change
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
df_filtered = df[df['apy']==1]
plt.scatter(df_filtered['y_reserves'], df_filtered['max_apy_change'], color='green',label= df_filtered['apy'].iloc[0])
df_filtered = df[df['apy']==5]
plt.scatter(df_filtered['y_reserves'], df_filtered['max_apy_change'], color='blue',label= df_filtered['apy'].iloc[0])
df_filtered = df[df['apy']==10]
plt.scatter(df_filtered['y_reserves'], df_filtered['max_apy_change'], color='red',label= df_filtered['apy'].iloc[0])
plt.title('FYT Reserves Vs Max Percent APY Change', fontsize=14)
plt.xlabel('FYT Reserves', fontsize=14)
plt.ylabel('Max Percent APY Change', fontsize=14)
plt.grid(True)
plt.legend(title="APY", loc='upper right')
plt.show()
## This tells me that reserve size has virtually no affect on Max % APY Change
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
#df_filtered = df[(df['time_stretch']==5) & (df['y_reserves']==50) & (df['tranche_length']==30)]
df_filtered = df[(df['time_stretch']==5) & (df['tranche_length']==30)]
plt.scatter(df_filtered['apy'], df_filtered['max_trade_amount'], color='green',label= df_filtered['tranche_length'].iloc[0])
#df_filtered = df[(df['time_stretch']==5) & (df['y_reserves']==500) & (df['tranche_length']==30)]
#plt.scatter(df_filtered['apy'], df_filtered['max_apy_change'], color='blue',label= df_filtered['y_reserves'].iloc[0])
#df_filtered = df[(df['time_stretch']==5) & (df['y_reserves']==1000) & (df['tranche_length']==30)]
#plt.scatter(df_filtered['apy'], df_filtered['max_apy_change'], color='red',label= df_filtered['y_reserves'].iloc[0])
plt.title('APY Vs Max Trade Amount', fontsize=14)
plt.xlabel('APY', fontsize=14)
plt.ylabel('Max Trade Amount', fontsize=14)
plt.grid(True)
plt.legend(title="Tranche Length")
plt.ylim(0,100)
plt.show()
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
df_filtered = df[df['time_stretch']==5]
plt.scatter(df_filtered['apy'], df_filtered['min_resulting_fyt_unit_prices'], color='green',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[df['time_stretch']==25]
plt.scatter(df_filtered['apy'], df_filtered['min_resulting_fyt_unit_prices'], color='blue',label= df_filtered['time_stretch'].iloc[0])
df_filtered = df[df['time_stretch']==50]
plt.scatter(df_filtered['apy'], df_filtered['min_resulting_fyt_unit_prices'], color='red',label= df_filtered['time_stretch'].iloc[0])
plt.title('APY Vs Max Percent APY Change', fontsize=14)
plt.xlabel('APY', fontsize=14)
plt.ylabel('Min Unit FYT Price, fontsize=14)
plt.grid(True)
plt.legend(title="Time Stretch")
plt.show()
### This tells me that yield bearing asset APY does have an affect on Max % APY Change
# +
import pandas as pd
import matplotlib.pyplot as plt
plt.subplots(figsize=(12,12))
df_filtered = df[df['apy']==5]
plt.scatter(df_filtered['time_stretch'], df_filtered['min_resulting_fyt_unit_prices'], color='green',label= df_filtered['apy'].iloc[0])
df_filtered = df[df['apy']==25]
plt.scatter(df_filtered['time_stretch'], df_filtered['min_resulting_fyt_unit_prices'], color='blue',label= df_filtered['apy'].iloc[0])
df_filtered = df[df['apy']==50]
plt.scatter(df_filtered['time_stretch'], df_filtered['min_resulting_fyt_unit_prices'], color='red',label= df_filtered['apy'].iloc[0])
plt.title('APY Vs Max Percent APY Change', fontsize=14)
plt.xlabel('Time Stretch', fontsize=14)
plt.ylabel('Min Unit FYT Price', fontsize=14)
plt.grid(True)
plt.legend(title="APY")
plt.show()
# -
df.to_csv('apy_change.csv')
APY = 8.66
term_length = 7
days_until_maturity = 4
time_stretch = 9
y_reserves = 177.52
x_reserves = 2995.64 #calc_x_reserves(APY,y_reserves,term_length-days_until_maturity,time_stretch)
print("(APY,y_reserves,days_until_maturity,time_stretch): "+str((APY,y_reserves,days_until_maturity,time_stretch)))
print("required x_reserves: " + str(x_reserves))
total_supply=x_reserves
price=calc_price(x_reserves,y_reserves,total_supply,days_until_maturity,time_stretch)
print("resulting price: " + str(price))
print("check apy: " + str(calc_apy(price,days_until_maturity)))
# +
x_reserves = 1856051.71
y_reserves = 1425357.41
total_supply = x_reserves+1425357.41
APY = 11.4
unit_price = 0.9728
t = 85/(365*9)
(x_reserves/(y_reserves+total_supply))**t
# -
3284587796090403818248754/10**18
1856051.71 + 1425357.41
3.09396 /( 0.02789 * 20)
5.11436 /( 0.04610 * 10)
| limits_to_price_discovery.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Self-Driving Car Engineer Nanodegree
#
# ## Deep Learning
#
# ## Project: Build a Traffic Sign Recognition Classifier
#
# In this notebook, a template is provided for you to implement your functionality in stages, which is required to successfully complete this project. If additional code is required that cannot be included in the notebook, be sure that the Python code is successfully imported and included in your submission if necessary.
#
# > **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
#
# In addition to implementing code, there is a writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) that can be used to guide the writing process. Completing the code template and writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/481/view) for this project.
#
# The [rubric](https://review.udacity.com/#!/rubrics/481/view) contains "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. The stand out suggestions are optional. If you decide to pursue the "stand out suggestions", you can include the code in this Ipython notebook and also discuss the results in the writeup file.
#
#
# >**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
# ---
# ## Step 0: Load The Data
# +
# Load pickled data
import os
import glob
import pickle
import pandas as pd
import numpy as np
import cv2
# TODO: Fill this in based on where you saved the training and testing data
training_file = 'dataset/train.p'
validation_file= 'dataset/valid.p'
testing_file = 'dataset/test.p'
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
print()
print("Image Shape: {}".format(X_train[0].shape))
print()
print("Training Set: {} samples".format(len(X_train)))
print("validation set: {} samples".format(len(X_valid)))
print("Test Set: {} samples".format(len(X_test)))
# -
# ---
#
# ## Step 1: Dataset Summary & Exploration
#
# The pickled data is a dictionary with 4 key/value pairs:
#
# - `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
# - `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
# - `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
# - `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
#
# Complete the basic data summary below. Use python, numpy and/or pandas methods to calculate the data summary rather than hard coding the results. For example, the [pandas shape method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.shape.html) might be useful for calculating some of the summary results.
# ### Provide a Basic Summary of the Data Set Using Python, Numpy and/or Pandas
# +
### Replace each question mark with the appropriate value.
n_train = X_train.shape[0]
n_validate = X_valid.shape[0]
n_test = X_test.shape[0]
image_shape = (X_train.shape[1], X_train.shape[2], X_train.shape[3])
n_classes = y_train.max() + 1
print("Number of training examples =", n_train)
print("Number of validation examples =", n_validate)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
# -
# ### Include an exploratory visualization of the dataset
# Visualize the German Traffic Signs Dataset using the pickled file(s). This is open ended, suggestions include: plotting traffic sign images, plotting the count of each sign, etc.
#
# The [Matplotlib](http://matplotlib.org/) [examples](http://matplotlib.org/examples/index.html) and [gallery](http://matplotlib.org/gallery.html) pages are a great resource for doing visualizations in Python.
#
# **NOTE:** It's recommended you start with something simple first. If you wish to do more, come back to it after you've completed the rest of the sections. It can be interesting to look at the distribution of classes in the training, validation and test set. Is the distribution the same? Are there more examples of some classes than others?
# +
### Data exploration visualization code goes here.
### Feel free to use as many code cells as needed.
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
# %matplotlib inline
hist, bins = np.histogram(y_train, bins=n_classes)
plt.hist(y_train, bins=bins)
plt.show()
hist, bins = np.histogram(y_test, bins=n_classes)
plt.hist(y_test, bins=bins)
plt.show()
hist, bins = np.histogram(y_valid, bins=n_classes)
plt.hist(y_valid, bins=bins)
plt.show()
# +
rows = 5
cols = 9
unique_index = np.unique(y_train, return_index=True)[1]
fig, axs = plt.subplots(rows, cols, figsize=(18, 14))
fig.subplots_adjust(hspace=.001, wspace=.001)
axs = axs.ravel()
for i in range(rows*cols):
axs[i].axis('off')
axs[i].imshow(X_train[unique_index[i]])
# -
# ----
#
# ## Step 2: Design and Test a Model Architecture
#
# Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
#
# The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
#
# With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
#
# There are various aspects to consider when thinking about this problem:
#
# - Neural network architecture (is the network over or underfitting?)
# - Play around preprocessing techniques (normalization, rgb to grayscale, etc)
# - Number of examples per label (some have more than others).
# - Generate fake data.
#
# Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
# ### Pre-process the Data Set (normalization, grayscale, etc.)
# Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - 128)/ 128` is a quick way to approximately normalize the data and can be used in this project.
#
# Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
#
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
# +
import skimage as sk
from skimage import transform
from skimage import util
from numpy.random import randint
import pandas as pd
augmentation = ['flip', 'noise', 'rotate']
MIN_IMG_COUNT = 1000
values, total = np.unique(y_train, return_counts=True)
augmented_X_train = np.copy(X_train)
augmented_y_train = np.copy(y_train)
def augment(image, randn):
technique = augmentation[randn]
if technique == 'flip':
return np.flip(image, axis=randn%2).astype(np.uint8)
elif technique == 'noise':
return (sk.util.random_noise(image, clip=True)*255).astype(np.uint8)
elif technique == 'rotate':
random_degree = randint(-90, 90)*randn
return np.rot90(image, randint(1, randn*randn)).astype(np.uint8)
# +
augmented_classes = np.where(total < MIN_IMG_COUNT)[0]
augmented_index = np.where([item==augmented_classes for item in y_train])[0]
print('Number of classes required to go through augmentation: %d' % len(augmented_classes))
# -
# image data augmentation
# if image class has less than 1000 images, it gets proceeded with augmentation
for item in augmented_classes:
augmented_index = np.where(item == y_train)[0]
class_count = total[item]
print('--------------------------------------')
print('%s has %d count.' % (item, class_count))
while (class_count < MIN_IMG_COUNT):
for index in augmented_index:
randn = randint(0, 3)
new_img = augment(X_train[index], randn)
augmented_X_train = np.append(augmented_X_train, np.reshape(new_img, (1, 32, 32, 3)), axis=0)
augmented_y_train = np.append(augmented_y_train, y_train[index])
class_count += 1
print('%s done!' % item)
# +
# save augmented image data into pickle
train = {'features': augmented_X_train, 'labels': augmented_y_train}
with open(os.path.join('dataset', 'augmented_train.p'), 'wb') as p:
pickle.dump(train, p)
# +
# load augmented image data from pickle
with open(os.path.join('dataset', 'augmented_train.p'), 'rb') as f:
augmented_train = pickle.load(f)
aug_X_train = augmented_train['features'].astype(np.float32)
aug_y_train = augmented_train['labels']
print('Length of augmented X and Y are: %d | %d ' % (len(aug_X_train), len(aug_y_train)))
# +
### visualize non augmented and augmented classes distribution
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
# %matplotlib inline
hist, bins = np.histogram(y_train, bins=n_classes)
plt.hist(y_train, bins=bins)
plt.show()
hist, bins = np.histogram(aug_y_train, bins=n_classes)
plt.hist(aug_y_train, bins=bins)
plt.show()
# +
### Preprocess the data here. It is required to normalize the data. Other preprocessing steps could include
### converting to grayscale, etc.
### Feel free to use as many code cells as needed.
# create grayscale image
def grayscale(img):
return cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
orig = aug_X_train[0].shape
print(orig)
X_train_gray = [np.reshape(grayscale(img), (orig[0], orig[1], 1)) for img in X_train]
X_valid_gray = [np.reshape(grayscale(img), (orig[0], orig[1], 1)) for img in X_valid]
X_test_gray = [np.reshape(grayscale(img), (orig[0], orig[1], 1)) for img in X_test]
print("after grayed Shape: {}".format(X_train_gray[0].shape) + '\n')
plt.figure()
plt.imshow(X_train_gray[0].squeeze())
# +
# normalize image to zero mean
def normalize(img):
mean = np.mean(img)
std = np.std(img)
return (img-mean)/std
X_train_normalized = [np.reshape(normalize(img), (32, 32, 1)) for img in X_train_gray]
X_valid_normalized = [np.reshape(normalize(img), (32, 32, 1)) for img in X_valid_gray]
X_test_normalized = [np.reshape(normalize(img), (32, 32, 1)) for img in X_test_gray]
plt.figure()
plt.imshow(X_train_normalized[0].squeeze())
# +
from sklearn.utils import shuffle
# shuffle training data
X_train_shuffle, y_train_shuffle = shuffle(X_train_normalized, y_train)
# -
# ### Model Architecture
# +
### Define your architecture here.
### Feel free to use as many code cells as needed.
import tensorflow as tf
from tensorflow.contrib.layers import flatten
EPOCHS = 60
BATCH_SIZE = 128
def ConvNet(x, is_train):
# arguments
mu = 0
sigma = 0.1
## input 32 x 32 x 1
# layer 1 ## 32 x 32 x 1
conv1_w = tf.Variable(tf.truncated_normal(shape=(5, 5, 1, 6), mean=mu, stddev=sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_w, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
## conv1 now has 28 x 28 x 6
# activation
conv1 = tf.nn.relu(conv1)
# pooling 1
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
## conv1 now has 14 x 14 x 6
# dropout and bn
conv1 = tf.layers.batch_normalization(conv1, training=is_train)
conv1 = tf.nn.dropout(conv1, keep_prob)
# layer 2 ## 14 x 14 x 6
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
## conv2 now has 10 x 10 x 16
# activation
conv2 = tf.nn.relu(conv2)
# pooling
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
## 5 x 5 x 16
# dropout and bn
conv2 = tf.layers.batch_normalization(conv2, training=is_train)
conv2 = tf.nn.dropout(conv2, keep_prob)
# layer 3 ## 5 x 5 x 16
conv3_W = tf.Variable(tf.truncated_normal(shape=(3, 3, 16, 100), mean = mu, stddev = sigma))
conv3_b = tf.Variable(tf.zeros(100))
conv3 = tf.nn.conv2d(conv2, conv3_W, strides=[1, 1, 1, 1], padding='VALID') + conv3_b
## 3 x 3 x 100
# activation
conv3 = tf.nn.relu(conv3)
# dropout and bn
conv3 = tf.layers.batch_normalization(conv3, training=is_train)
conv3 = tf.nn.dropout(conv3, keep_prob)
# flatten conv2 output
fc0 = flatten(conv3)
# layer 4, fully connected layer
fc1_W = tf.Variable(tf.truncated_normal(shape=(900, 400), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(400))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# activation
fc1 = tf.nn.relu(fc1)
#fc1 = tf.layers.batch_normalization(fc1, training=is_train)
# layer 5, fully connected layer
fc2_W = tf.Variable(tf.truncated_normal(shape=(400, 200), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(200))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# activation
fc2 = tf.nn.relu(fc2)
# layer 5, fully connected layer
fc3_W = tf.Variable(tf.truncated_normal(shape=(200, n_classes), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(n_classes))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
# -
# ### Train, Validate and Test the Model
# A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
# sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
# +
### Train your model here.
### Feel free to use as many code cells as needed.
is_train = tf.placeholder(tf.bool, name="is_train")
x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, (None))
keep_prob = tf.placeholder(tf.float32)
one_hot_y = tf.one_hot(y, n_classes)
rate = 0.0005
logits = ConvNet(x, is_train=True)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=one_hot_y)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(update_ops):
training_operation = optimizer.minimize(loss_operation, global_step=tf.train.get_global_step())
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y, keep_prob: 1.0, is_train: True})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
# -
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train_gray)
print("Training...")
print()
for i in range(EPOCHS):
X_train_shuffle, y_train_shuffle = shuffle(X_train_normalized, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train_shuffle[offset:end], y_train_shuffle[offset:end]
sess.run([training_operation], feed_dict={x: batch_x, y: batch_y, keep_prob: 0.5, is_train: True})
validation_accuracy = evaluate(X_valid_normalized, y_valid)
print("EPOCH {} ...".format(i+1))
print("Validation Accuracy = {:.3f}".format(validation_accuracy))
print()
test_accuracy = evaluate(X_test_normalized, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
print()
try:
saver
except NameError:
saver = tf.train.Saver()
saver.save(sess, 'lenet')
print("Model saved")
# ---
#
# ## Step 3: Test a Model on New Images
#
# To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
#
# You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
# ### Load and Output the Images
# +
### Load the images and plot them here.
### Feel free to use as many code cells as needed.
import matplotlib.image as mpimg
pic_dir = os.path.join('images', 'test_images')
# creating list of images
images = [mpimg.imread(img) for img in sorted(glob.glob(os.path.join(pic_dir, '*.png')))]
rows = 2
cols = 4
fig, axs = plt.subplots(rows, cols, figsize=(5, 5))
fig.subplots_adjust(hspace=.001, wspace=.001)
axs = axs.ravel()
for i in range(rows*cols):
axs[i].axis('off')
axs[i].imshow(images[i])
print('Images have shape of %s and type %s' % (images[-1].shape, type(images[-1])))
# -
# ### Predict the Sign Type for Each Image
# +
### Run the predictions here and use the model to output the prediction for each image.
### Make sure to pre-process the images with the same pre-processing pipeline used earlier.
### Feel free to use as many code cells as needed.
orig = images[0].shape
print("Original X prediction shape: {}".format(images[0].shape) + '\n')
X_pred_gray = [np.reshape(grayscale(cv2.cvtColor(img, cv2.COLOR_RGB2BGR)), (orig[0], orig[1], 1)) for img in images]
X_pred_normalized = [np.reshape(normalize(img), (32, 32, 1)) for img in X_pred_gray]
print("After grayed Shape: {}".format(X_train_gray[0].shape) + '\n')
plt.figure()
plt.imshow(X_pred_gray[0].squeeze())
plt.figure()
plt.imshow(X_pred_normalized[0].squeeze())
# -
# ### Analyze Performance
# +
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
labels = [11, 1, 12, 38, 34, 18, 25, 3]
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
saver.restore(sess, 'lenet')
#prediction = sess.run(logits, feed_dict={x: X_pred_normalized, keep_prob : 1.0})
pred_accuracy = evaluate(X_pred_normalized, labels)
print("Prediction Set Accuracy = {:.3f}".format(pred_accuracy))
# -
# ### Output Top 5 Softmax Probabilities For Each Image Found on the Web
# For each of the new images, print out the model's softmax probabilities to show the **certainty** of the model's predictions (limit the output to the top 5 probabilities for each image). [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here.
#
# The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
#
# `tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
#
# Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. `tf.nn.top_k` is used to choose the three classes with the highest probability:
#
# ```
# # (5, 6) array
# a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
# 0.12789202],
# [ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
# 0.15899337],
# [ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
# 0.23892179],
# [ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
# 0.16505091],
# [ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
# 0.09155967]])
# ```
#
# Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
#
# ```
# TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
# [ 0.28086119, 0.27569815, 0.18063401],
# [ 0.26076848, 0.23892179, 0.23664738],
# [ 0.29198961, 0.26234032, 0.16505091],
# [ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
# [0, 1, 4],
# [0, 5, 1],
# [1, 3, 5],
# [1, 4, 3]], dtype=int32))
# ```
#
# Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
# +
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
### Feel free to use as many code cells as needed.
img = [img.split(os.sep)[-1] for img in sorted(glob.glob(os.path.join(pic_dir, '*.png')))]
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
saver.restore(sess, 'lenet')
prediction = sess.run(logits, feed_dict={x: X_pred_normalized, keep_prob : 1.0})
predicts = sess.run(tf.nn.top_k(prediction, k=5, sorted=True))
for i in range(len(predicts[0])):
print('Probabilities of Image', img[i], ':', predicts[0][i], '\nPredicted classes:', predicts[1][i], '\n-----')
# -
# ### Project Writeup
#
# Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
# ---
#
# ## Step 4 (Optional): Visualize the Neural Network's State with Test Images
#
# This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
#
# Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
#
# For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
#
# <figure>
# <img src="visualize_cnn.png" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> Your output should look something like this (above)</p>
# </figcaption>
# </figure>
# <p></p>
#
# +
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
| traffic-sign-classifier/Traffic_Sign_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <center><h1>Domain Expertise in Data Analysis</h1></center>
# <center><h3><NAME></h3></center>
# <center><h3>Twitter: @gnfrazier</h3></center>
# <center><h3>https://github.com/gnfrazier/Analytics-Forward-Domain-Expertise</h3></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><h2>Venn Diagram of Data Science</h2></center>
#
# <center><img src="data_venn_diagram.jpg"></center>
#
# <center>http://www.prooffreader.com/2016/09/battle-of-data-science-venn-diagrams.html</center>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## What is Domain Expertise?
#
# ### Domain Knowledge
# ### Subject Matter Expert
# ### Professional
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why is it important?
# - Domain Specific Terms and Acronyms
# - Metric Parameters
# - Definitions
# - Criteria
# - Variation from company to company
# - Understanding Field Names and Data Structures
# - All important Gut Check
# - Allows faster iteration and discovery
# - Communication and Presentation
# + [markdown] slideshow={"slide_type": "slide"}
# ## When?
#
# # Always and Constantly
#
# "Not growing your career you are slowing your career" - <NAME>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Digging into a Vertical - Passive
#
# - YouTube
# - Podcasts
# - Forums
# - Social Media
# + [markdown] slideshow={"slide_type": "slide"}
# ## Digging into a Vertical - Active
#
# - Other Data Analysts and Data Scientists in the company
# - Sales or Marketing folks down the hall
# - Meetups
# - Conferences
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## What do I get out of it?
# - Understanding of strategy that may be out of scope
# - Trust within the organization
# - Ability to communicate to executives
# - Ability to rephrase narratives into terms people understand
# - You know who to ask
# + [markdown] slideshow={"slide_type": "slide"}
# ## Disscussion Points
# - Industry Survey
# - Challenges in transitioning from Academia to Private Sector
# - More resources to learn various segments
#
# #### Slides on my github https://github.com/gnfrazier/Analytics-Forward-Domain-Expertise
# + [markdown] slideshow={"slide_type": "slide"}
# ## Disscussion Points Surfaced by the Group
#
# #### Additional Important Points
# - Knowing Buzzwords for a company or industry
# - Become an expert at working with domain experts
# - Feeling of current expertise not being valuable
# - Transitioning from thinking/thought to application
# - Contrast of research in academia vs private sector
# - Keep yourself in scope/balance (don't get too deep in other departments domain expertise)
# - Remember that companies lie
# - Remember that companies are not uniform
#
#
# #### Additional Sources or Resourses to gain Domain Expertise
# - Mooc - Coursera, Pluralsite, etc
# - Books and printed material
# - Journals and Papers
# - Google Scholar
# - Friends of the NCSU library (full article access)
# - JOSS
# - Stack Exchange (even for non-software questions)
# - Industry Professional Groups and Meetups
# - Industry Magazines
# - University Department Seminars
# - Safari Books Online
# - O'reilly and Pearson
# - Mentor or Mentee
# - Talk to real customers!!
# - Company Mission Statement
# - Glassdoor - Previous Employee Reviews
# - Social Media Feeds for the company (full immersion)
# - Linked In
#
| Domain-Expertise-in-Data-Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Logistic Regression with Bert encodings
import numpy as np
import pandas as pd
# for model:
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
from sklearn.multiclass import OneVsRestClassifier
from sklearn.model_selection import train_test_split
# for scoring:
from sklearn import metrics
from sklearn.metrics import f1_score
x = np.loadtxt("toxic_bert_matrix.out", delimiter=",")
df = pd.read_csv('toxic-train-clean.csv')
y = df.iloc[:, 2:8]
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size= 0.2)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# +
# liblinear used because of onevsrest:
pipe = make_pipeline(OneVsRestClassifier(LogisticRegression(max_iter=500, class_weight='balanced')))
param_grid = {'onevsrestclassifier__estimator__solver': ['liblinear']}
grid = GridSearchCV(pipe, param_grid, cv=15, scoring='roc_auc', verbose=3)
grid3 = grid.fit(X_train, y_train)
# -
grid3.best_score_
predicted_y_test = grid3.predict(X_test)
predicted_y_test[:1]
y_pred_prob = grid3.predict_proba(X_test)
y_pred_prob[:1]
auc_score = metrics.roc_auc_score(y_test, y_pred_prob)
auc_score
f1_score(y_test, predicted_y_test, average='micro')
| bert_log_reg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="tAOl3D-XPbxW" outputId="621c0d63-4706-4aa9-960c-bf266da56680"
x=input('Enter 1st no.')
# + colab={"base_uri": "https://localhost:8080/"} id="TGG0Aj0JPs7N" outputId="81d9c0b8-d093-49c7-c1e7-9041a3b3f164"
y=input('Enter 2nd no.')
# + colab={"base_uri": "https://localhost:8080/"} id="S5uMJ2quPyvr" outputId="bb005b91-8280-4126-c26e-8e1adcf377c4"
int(x)+int(y)
# + colab={"base_uri": "https://localhost:8080/"} id="S0w8E9keP8ym" outputId="2c1e0df1-8b5f-4b9a-b537-133251a50ed2"
int(x)-int(y)
# + colab={"base_uri": "https://localhost:8080/"} id="RxfAxPOaQAwm" outputId="b47d9bfb-e1cb-49bb-9f4e-0e86c89ca750"
int(x)*int(y)
# + colab={"base_uri": "https://localhost:8080/"} id="vLSGmwWPQEze" outputId="6511df3d-2a48-4c80-c521-6f38e9e440a4"
int(x)/int(y)
# + colab={"base_uri": "https://localhost:8080/"} id="aMd3vOEnQIgW" outputId="653866e7-2104-439a-d071-29f6c8e132a5"
x=int(input('Enter 1st no.'))
# + colab={"base_uri": "https://localhost:8080/"} id="jd5VeEV0R8st" outputId="877677e2-ee9d-40b7-9f0a-56d45bd5136d"
y=int(input('Enter 2nd no.'))
# + colab={"base_uri": "https://localhost:8080/"} id="80kByhqrSGxp" outputId="bd570bfe-a4cb-4678-e8cd-6e6510e365e6"
x+y
# + colab={"base_uri": "https://localhost:8080/"} id="A8M3CGaVSK45" outputId="f0ea1f69-a119-412b-b426-ebf99f84578b"
x-y
# + colab={"base_uri": "https://localhost:8080/"} id="gDbBO3Z8SMf5" outputId="606fa6fb-6a09-436b-8d68-44da47072ec0"
x*y
# + colab={"base_uri": "https://localhost:8080/"} id="2FYRprnTSNtA" outputId="d93ca2e6-6a43-4253-f4f0-7b515ae6ee47"
x/y
# + colab={"base_uri": "https://localhost:8080/"} id="N5OoPPjxSOuF" outputId="2a5241ad-ca3c-4a69-acf4-a3cbd5e0b6c7"
p=int(input('Enter principle amount'))
# + colab={"base_uri": "https://localhost:8080/"} id="ZCV2BP2YTvE1" outputId="411862c5-dea6-4b3e-fad2-ee94a22012d3"
n=int(input('Enter no. of years'))
# + colab={"base_uri": "https://localhost:8080/"} id="4HsIi06JT28B" outputId="84f5d4b4-6fff-4292-94ce-5715984f055f"
r=int(input('Enter rate'))
# + id="TUmcVJUVUEkw"
i=p*n*r/100
# + colab={"base_uri": "https://localhost:8080/"} id="xhH-eTauUSqg" outputId="8a3bb946-1238-42f5-fdaa-1066944059a2"
print(i)
# + colab={"base_uri": "https://localhost:8080/"} id="clzSlRo0UWHP" outputId="dc851144-358c-462f-b9f0-b6da9fd59e69"
x=int(input('Enter your marks'))
# + colab={"base_uri": "https://localhost:8080/"} id="BW6LYFcSYW2I" outputId="1c499bf8-f5c0-4c8d-d195-b207d2b7c450"
if x > 50 :
print('You have passed')
if x < 50 :
print('You have failed')
# + colab={"base_uri": "https://localhost:8080/"} id="aUP-17HfY7kC" outputId="bd210e35-b5c1-4195-a72e-0155f5c9ac3b"
x=int(input('Enter your salary'))
# + colab={"base_uri": "https://localhost:8080/"} id="BFnKdsaEarfK" outputId="057b232e-59e5-41fc-f858-f64597302387"
if x>=250000:
print('You are taxable')
if x<250000:
print('You are not taxable')
# + colab={"base_uri": "https://localhost:8080/"} id="G1oGw1gibQ97" outputId="25de2e3a-fbb9-4c9d-9bb6-fa35e09e3620"
x=int(input('Enter your salary'))
# + colab={"base_uri": "https://localhost:8080/"} id="28eun0xpbtt_" outputId="0b166cde-ab12-4d20-b9e4-f65d8f3e15bd"
if x>=250000:
print('You are taxable')
print('Your tax amount is',x*10/100)
if x<250000:
print('You are not taxable')
# + colab={"base_uri": "https://localhost:8080/"} id="w86Fjjk-cQm2" outputId="137c0286-9f6c-4a6d-d528-c1af81eb1555"
x=int(input('Enter your salary'))
# + colab={"base_uri": "https://localhost:8080/"} id="a2V2qVI-eLnJ" outputId="5606d4b5-c032-4c2d-ef7f-920587568707"
if x>=250000:
print('You are taxable')
print('Your tax amount is',(x-250000)*10/100)
if x<250000:
print('You are not taxable')
# + colab={"base_uri": "https://localhost:8080/"} id="WbgaT2Wre2H2" outputId="9f6a6716-b08f-41fb-c4cf-3069725a0adc"
x=int(input('Enter your salary'))
# + colab={"base_uri": "https://localhost:8080/"} id="6nbPx8B4jmm9" outputId="b788e237-a08e-423f-a263-4f229bddb533"
if x>250000 and x<500000:
print('You are taxable')
print('Your tax amount is',(x-250000)*10/100)
if x>500000:
print('You are taxable')
print('Your tax amount is',(x-250000)*20/100)
if x<250000:
print('You are not taxable')
# + id="tIWa8EGnkFYk"
| PythonTax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.1 64-bit (''campaign'': conda)'
# language: python
# name: python38164bitcampaignconda511bbd138feb405e9bb06c17f1890a1c
# ---
# +
from matplotlib import pyplot as plt
from matplotlib import cm
import pandas as pd
from pprint import pprint
from random import randint
import numpy as np
#import matplotlib as mpl
#mpl.rcParams['text.usetex'] = True
#mpl.rcParams['text.latex.unicode'] = True
blues = cm.get_cmap(plt.get_cmap('Blues'))
greens = cm.get_cmap(plt.get_cmap('Greens'))
reds = cm.get_cmap(plt.get_cmap('Reds'))
oranges = cm.get_cmap(plt.get_cmap('Oranges'))
purples = cm.get_cmap(plt.get_cmap('Purples'))
greys = cm.get_cmap(plt.get_cmap('Greys'))
set1 = cm.get_cmap(plt.get_cmap('Set1'))
tab20 = cm.get_cmap(plt.get_cmap('tab20'))
def tableau20(color):
# Use coordinated colors. These are the "Tableau 20" colors as
# RGB. Each pair is strong/light. For a theory of color
tableau20 = [(31 , 119, 180), (174, 199, 232), # blue [ 0,1 ]
(255, 127, 14 ), (255, 187, 120), # orange [ 2,3 ]
(44 , 160, 44 ), (152, 223, 138), # green [ 4,5 ]
(214, 39 , 40 ), (255, 152, 150), # red [ 6,7 ]
(148, 103, 189), (197, 176, 213), # purple [ 8,9 ]
(140, 86 , 75 ), (196, 156, 148), # brown [10,11]
(227, 119, 194), (247, 182, 210), # pink [12,13]
(188, 189, 34 ), (219, 219, 141), # yellow [14,15]
(23 , 190, 207), (158, 218, 229), # cyan [16,17]
(65 , 68 , 81 ), (96 , 99 , 106), # gray [18,19]
(127, 127, 127), (143, 135, 130), # gray [20,21]
(165, 172, 175), (199, 199, 199), # gray [22,23]
(207, 207, 207)] # gray [24]
# Scale the RGB values to the [0, 1] range, which is the format
# matplotlib accepts.
r, g, b = tableau20[color]
return (round(r/255.,1), round(g/255.,1), round(b/255.,1))
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import warnings
warnings.filterwarnings('ignore')
# +
StHomoCampaigns_4StHomoResourcesGA = pd.read_csv('../Data/ga/perc_100/StHomoCampaigns_4DynHomoResourcesGA00_new.csv')
StHomoCampaigns_4StHomoResourcesGA25 = pd.read_csv('../Data/ga/perc_075/StHomoCampaigns_4DynHomoResourcesGA25_new.csv')
StHomoCampaigns_4StHomoResourcesGA50 = pd.read_csv('../Data/ga/perc_050/StHomoCampaigns_4DynHomoResourcesGA50_new.csv')
#StHomoCampaigns_4StHomoResourcesGA.drop(['plan', 'time'], axis=1, inplace=True)
#StHomoCampaigns_4StHomoResourcesGA25.drop(['plan', 'time'], axis=1, inplace=True)
#StHomoCampaigns_4StHomoResourcesGA50.drop(['plan', 'time'], axis=1, inplace=True)
# -
StHomoCampaigns_4StHomoResourcesGA
# +
ConvGA = pd.DataFrame(columns=['count', 'total'])
ConvGA25 = pd.DataFrame(columns=['count', 'total'])
ConvGA50 = pd.DataFrame(columns=['count', 'total'])
for i in [4,8,16,32,64,128,256,512,1024, 2048]:
ConvGA.loc[i] = [0,0]
ConvGA25.loc[i] = [0,0]
ConvGA50.loc[i] = [0,0]
for idx, row in StHomoCampaigns_4StHomoResourcesGA.iterrows():
if row['expected'] == (75000 * row['size'] / 4):
ConvGA.loc[row['size']]['count'] += 1
ConvGA.loc[row['size']]['total'] += 1
for idx, row in StHomoCampaigns_4StHomoResourcesGA25.iterrows():
if row['expected'] == (75000 * row['size'] / 4):
ConvGA25.loc[row['size']]['count'] += 1
ConvGA25.loc[row['size']]['total'] += 1
for idx, row in StHomoCampaigns_4StHomoResourcesGA50.iterrows():
if row['expected'] == (75000 * row['size'] / 4):
ConvGA50.loc[row['size']]['count'] += 1
ConvGA50.loc[row['size']]['total'] += 1
# -
ConvGA
fig,axis = plt.subplots(nrows=1,ncols=1,figsize=(15,7.5))
_ = axis.errorbar(ConvGA.index, ConvGA['count'] / ConvGA['total'], color=tableau20(4), marker='^', label='GA', linewidth=2)
_ = axis.errorbar(ConvGA25.index, ConvGA25['count'] / ConvGA25['total'], color=tableau20(6),marker='^', label='GA-25',linewidth=2)
_ = axis.errorbar(ConvGA50.index, ConvGA50['count'] / ConvGA50['total'], color=tableau20(8),marker='^', label='GA-50',linewidth=2)
#_ = axis.errorbar([4, 8, 16, 32, 64, 128, 256, 512, 1024], min_vals, linestyle='--',label='Expected', color=greys(150),linewidth=2)
# _ = axis.fill_between(workflows_sizes, min_vals, max_vals, color=greens(250), alpha=0.2, label='RANDOM Max-Min')
_ = axis.set_xscale('symlog', basex=2)
_ = axis.set_ylabel('Convergence Rate',fontsize=20)
_ = axis.set_xlabel('Number of workflows',fontsize=20)
_ = axis.set_xticklabels(axis.get_xticks().astype('int'),fontsize=18)
_ = axis.set_yticklabels(np.around(axis.get_yticks().astype('float'), decimals=2),fontsize=18)
_ = axis.grid('on')
#_ = axis.set_title('As a function of the number of workflows', fontsize=22)
_ = axis.legend(fontsize=20)
#_ = fig.savefig('../Figures/StHomoCampaigns_4StHomoResources.png',bbox_inches='tight')
_ = fig.savefig('../Figures/StHomoCampaigns_4StHomoResourcesGAconv.png',bbox_inches='tight')
_ = fig.savefig('../Figures/StHomoCampaigns_4StHomoResourcesGAconv.pdf',bbox_inches='tight')
# +
StHomoCampaigns_4StHomoResourcesGA = pd.read_csv('../Data/ga/perc_100/DynHomoResources_StHomoCampaignsGA00_new.csv')
StHomoCampaigns_4StHomoResourcesGA25 = pd.read_csv('../Data/ga/perc_075/DynHomoResources_StHomoCampaignsGA25_new.csv')
StHomoCampaigns_4StHomoResourcesGA50 = pd.read_csv('../Data/ga/perc_050/DynHomoResources_StHomoCampaignsGA50_new.csv')
StHomoCampaigns_4StHomoResourcesGA.drop(['plan', 'time'], axis=1, inplace=True)
StHomoCampaigns_4StHomoResourcesGA25.drop(['plan', 'time'], axis=1, inplace=True)
StHomoCampaigns_4StHomoResourcesGA50.drop(['plan', 'time'], axis=1, inplace=True)
# +
ConvGA = pd.DataFrame(columns=['count', 'total'])
ConvGA25 = pd.DataFrame(columns=['count', 'total'])
ConvGA50 = pd.DataFrame(columns=['count', 'total'])
for i in [4, 8, 16, 32, 64, 128, 256]:
ConvGA.loc[i] = [0,0]
ConvGA25.loc[i] = [0,0]
ConvGA50.loc[i] = [0,0]
for idx, row in StHomoCampaigns_4StHomoResourcesGA.iterrows():
if row['expected'] == (75000 * 1024 / row['size']):
ConvGA.loc[row['size']]['count'] += 1
ConvGA.loc[row['size']]['total'] += 1
for idx, row in StHomoCampaigns_4StHomoResourcesGA25.iterrows():
if row['expected'] == (75000 * 1024 / row['size']):
ConvGA25.loc[row['size']]['count'] += 1
ConvGA25.loc[row['size']]['total'] += 1
for idx, row in StHomoCampaigns_4StHomoResourcesGA50.iterrows():
if row['expected'] == (75000 * 1024 / row['size']):
ConvGA50.loc[row['size']]['count'] += 1
ConvGA50.loc[row['size']]['total'] += 1
# -
fig,axis = plt.subplots(nrows=1,ncols=1,figsize=(15,7.5))
_ = axis.errorbar(ConvGA.index, ConvGA['count'] / ConvGA['total'], color=tableau20(4), marker='^', label='GA', linewidth=2)
_ = axis.errorbar(ConvGA25.index, ConvGA25['count'] / ConvGA25['total'], color=tableau20(6),marker='^', label='GA-25',linewidth=2)
_ = axis.errorbar(ConvGA50.index, ConvGA50['count'] / ConvGA50['total'], color=tableau20(8),marker='^', label='GA-50',linewidth=2)
#_ = axis.errorbar([4, 8, 16, 32, 64, 128, 256, 512, 1024], min_vals, linestyle='--',label='Expected', color=greys(150),linewidth=2)
# _ = axis.fill_between(workflows_sizes, min_vals, max_vals, color=greens(250), alpha=0.2, label='RANDOM Max-Min')
_ = axis.set_xscale('symlog', basex=2)
_ = axis.set_ylabel('Convergence Rate',fontsize=20)
_ = axis.set_xlabel('Number of resources',fontsize=20)
_ = axis.set_xticklabels(axis.get_xticks().astype('int'),fontsize=18)
_ = axis.set_yticklabels(np.around(axis.get_yticks().astype('float'), decimals=2),fontsize=18)
_ = axis.grid('on')
#_ = axis.set_title('As a function of the number of resources', fontsize=22)
_ = axis.legend(fontsize=20)
#_ = fig.savefig('../Figures/StHomoCampaigns_4StHomoResources.png',bbox_inches='tight')
_ = fig.savefig('../Figures/HomogeResources_StHomogeCampaignsGAconv.png',bbox_inches='tight')
_ = fig.savefig('../Figures/HomogeResources_StHomogeCampaignsGAconv.pdf',bbox_inches='tight')
| Notebooks/GAConvergenceRate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
letters = ['a', 'b', 'c']
numbers = [1, 2, 3]
np_arr = np.array(numbers)
dict = {'a':1, 'b':2, 'c':3}
pd.Series(data = numbers)
pd.Series(data=numbers, index=letters)
pd.Series(numbers, letters)
letters
numbers
np_arr
dict
pd.Series(np_arr)
pd.Series(np_arr, letters)
dict
pd.Series(dict)
pd.Series(data=letters)
life_long_average = pd.Series([84.7, 84.5, 83.7], ['Hong Kong', 'Japan', 'Singapore'])
life_long_average
life_long_average['Hong Kong']
life_long_average1 = pd.Series([84.7, 84.5, 83.7], ['USA', 'Japan', 'Singapore'])
life_long_average1 + life_long_average
| notebooks/Python3-DataScience/03-Pandas/01-Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pylab as pl
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
from sklearn.utils import shuffle
from sklearn.svm import SVC
from sklearn.metrics import confusion_matrix,classification_report
from sklearn.model_selection import cross_val_score, GridSearchCV
from collections import Counter
from matplotlib import pyplot
from sklearn.model_selection import StratifiedKFold
final_df = pd.read_csv('final_Suleyman', index_col = 0)
final_df
final_df = final_df.drop(['DOB', 'DOD','ADMITTIME'], axis = 1)
X = final_df.iloc[:,:-1]
y = final_df.iloc[:,-1]
counter = Counter(y)
for k,v in counter.items():
per = v / len(y) * 100
print('Class=%d, n=%d (%.3f%%)' % (k, v, per))
# plot the distribution
pyplot.bar(counter.keys(), counter.values())
pyplot.show()
from imblearn.over_sampling import SMOTE
# transform the dataset
oversample = SMOTE()
X, y = oversample.fit_resample(X, y)
# summarize distribution
counter = Counter(y)
for k,v in counter.items():
per = v / len(y) * 100
print('Class=%d, n=%d (%.3f%%)' % (k, v, per))
# plot the distribution
pyplot.bar(counter.keys(), counter.values())
pyplot.show()
final_df = final_df.reset_index()
X = final_df.iloc[:,:-1]
y = final_df.iloc[:,-1]
skf = StratifiedKFold(n_splits=2, shuffle = True, random_state = 3)
for train_index, test_index in skf.split(np.zeros(len(y)), y):
print("TRAIN:", train_index, "TEST:", test_index)
train_X, test_X = X.loc[train_index], X.loc[test_index]
train_y,test_y = y.loc[train_index],y.loc[test_index]
# training a Naive Bayes classifier
from sklearn.naive_bayes import GaussianNB
gnb = GaussianNB()
from yellowbrick.classifier import ClassificationReport
visualizer = ClassificationReport(gnb, classes=y.unique(), support=True)
visualizer.fit(train_X, train_y) # Fit the visualizer and the model
visualizer.score(test_X, test_y) # Evaluate the model on the test data
visualizer.show() # Plot the result
scores = cross_val_score(gnb, X, y, cv=10)
print(scores)
scores.mean()
| GausianNB (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://www.kaggle.com/drjohnwagner/heart-disease-prediction-with-xgboost?scriptVersionId=85327390" target="_blank"><img align="left" alt="Kaggle" title="Open in Kaggle" src="https://kaggle.com/static/images/open-in-kaggle.svg"></a>
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 11.441957, "end_time": "2022-01-15T20:33:26.09305", "exception": false, "start_time": "2022-01-15T20:33:14.651093", "status": "completed"} tags=[]
import json
import random
import numpy as np
import pandas as pd
from igraph import Graph
import igraph
from pprint import pprint
#
import plotly.io as pio
import plotly.graph_objects as go
#
from xgboost import XGBClassifier
from xgboost import XGBModel
from xgboost import Booster
#
from sklearn.compose import ColumnTransformer
from sklearn.feature_selection import SelectKBest, chi2
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import make_scorer
from sklearn.metrics import plot_confusion_matrix
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OrdinalEncoder
from sklearn.preprocessing import MinMaxScaler
from sklearn.utils import shuffle
try:
from PDSUtilities.xgboost import plot_importance
from PDSUtilities.xgboost import plot_tree
except ImportError as e:
try:
# !pip install PDSUtilities --upgrade
from PDSUtilities.xgboost import plot_importance
from PDSUtilities.xgboost import plot_tree
except ImportError as e:
raise ImportError("You must install PDSUtilities to plot importance and trees...") from e
from plotly.offline import init_notebook_mode, iplot
init_notebook_mode(connected=True)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk("/kaggle/input"):
for filename in filenames:
print(os.path.join(dirname, filename))
# # Colorblindness friendly colours...
# # It is important to make our work
# # as accessible as possible...
# COLORMAP = ["#005ab5", "#DC3220"]
# Labels for plotting...
LABELS = {
"Sex": "Sex",
"Age": "Age",
"MaxHR": "Max HR",
"OldPeak": "Old Peak",
"STSlope": "ST Slope",
"RestingBP": "Rest. BP",
"FastingBS": "Fast. BS",
"RestingECG": "Rest. ECG",
"Cholesterol": "Cholesterol",
"HeartDisease": "Heart Disease",
"ChestPainType": "Chest Pain",
"ExerciseAngina": "Ex. Angina",
}
# Random seed for determinism...
SEED = 395147
# # You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# # You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + [markdown] papermill={"duration": 0.012985, "end_time": "2022-01-15T20:33:26.121184", "exception": false, "start_time": "2022-01-15T20:33:26.108199", "status": "completed"} tags=[]
# ## Description
# Simple notbook showing how to use `xgboost` to analyse the Heart Disease dataset.
#
# Notebook also shows how to use **`PDSUtilities.xgboost.plot_importance()`** and **`PDSUtilities.xgboost.plot_tree()`**, my Plotly-based replacements for `xgboost.plot_importance()` and `xgboost.plot_tree()`.
#
# Check them out at [https://github.com/DrJohnWagner/PDSUtilities](https://github.com/DrJohnWagner/PDSUtilities).
# + [markdown] papermill={"duration": 0.012923, "end_time": "2022-01-15T20:33:26.147567", "exception": false, "start_time": "2022-01-15T20:33:26.134644", "status": "completed"} tags=[]
# ## Load the Data
# + papermill={"duration": 0.053872, "end_time": "2022-01-15T20:33:26.214856", "exception": false, "start_time": "2022-01-15T20:33:26.160984", "status": "completed"} tags=[]
# Loading the data from the csv file...
df = pd.read_csv("/kaggle/input/heart-failure-prediction/heart.csv")
df.head()
# + [markdown] papermill={"duration": 0.013358, "end_time": "2022-01-15T20:33:26.242343", "exception": false, "start_time": "2022-01-15T20:33:26.228985", "status": "completed"} tags=[]
# ## Fix the Column Names
# Columns `ST_Slope` and `Oldpeak` do not use the same naming convention as the other columns.
#
# We also convert `HeartDisease` to a categorical variable (`int64` to `int8`)...
# + papermill={"duration": 0.034808, "end_time": "2022-01-15T20:33:26.29097", "exception": false, "start_time": "2022-01-15T20:33:26.256162", "status": "completed"} tags=[]
# Fix the egregious column naming error...
df = df.rename(columns = {"ST_Slope": "STSlope", "Oldpeak": "OldPeak"})
# Always test these things...
assert len(df["STSlope"]) > 0, "Ruh roh! ST_Slope is still terribly mistaken!"
assert len(df["OldPeak"]) > 0, "Ruh roh! Oldpeak is still terribly mistaken!"
# Convert target to categorical
target = pd.Categorical(df["HeartDisease"])
df["HeartDisease"] = target.codes
print("Datatypes")
print("---------")
print(df.dtypes)
# + [markdown] papermill={"duration": 0.014682, "end_time": "2022-01-15T20:33:26.32007", "exception": false, "start_time": "2022-01-15T20:33:26.305388", "status": "completed"} tags=[]
# ## Grab the Column Names
# + papermill={"duration": 0.024584, "end_time": "2022-01-15T20:33:26.35897", "exception": false, "start_time": "2022-01-15T20:33:26.334386", "status": "completed"} tags=[]
# Break the columns into two groupings...
categorical_columns = [column for column in df.columns if df[column].dtypes == np.object]
numerical_columns = [column for column in df.columns if df[column].dtypes != np.object]
if "HeartDisease" in numerical_columns:
numerical_columns.remove("HeartDisease")
assert "HeartDisease" not in numerical_columns, "Ruh roh! HeartDisease is still in numerical_columns!"
print("Categorical Columns: ", categorical_columns)
print(" Numerical Columns: ", numerical_columns)
# + [markdown] papermill={"duration": 0.01406, "end_time": "2022-01-15T20:33:26.387572", "exception": false, "start_time": "2022-01-15T20:33:26.373512", "status": "completed"} tags=[]
# ## Randomly Redistribute Missing Values
# The columns `RestingBP` and `Cholesterol` have records with value 0. Hypothesising that these
# represent missing values, we set them to random values drawn from a normal distribution fit to
# the rest of the values in each of those columns.
# + papermill={"duration": 0.033939, "end_time": "2022-01-15T20:33:26.435705", "exception": false, "start_time": "2022-01-15T20:33:26.401766", "status": "completed"} tags=[]
def set_column_value_to_normal_distribution(df, column, value):
# Compute the column's mean and standard deviation
# after removing rows whose column matches value...
mean_value = df[df[column] != value][column].mean()
std_value = df[df[column] != value][column].std()
# Create a random number generator...
rng = np.random.default_rng(SEED)
# Now set the column of those rows to a
# random sample from a normal distribution...
df[column] = df[column].apply(
lambda x : rng.normal(mean_value, std_value) if x == value else x
)
return df
df = set_column_value_to_normal_distribution(df, "RestingBP" , 0)
df = set_column_value_to_normal_distribution(df, "Cholesterol", 0)
# Always test...
assert len(df[df["RestingBP" ] == 0]) == 0, "Ruh roh! One or more patients has crashed again!"
assert len(df[df["Cholesterol"] == 0]) == 0, "Ruh roh! One or more patients has crashed again!"
# + [markdown] papermill={"duration": 0.014046, "end_time": "2022-01-15T20:33:26.464193", "exception": false, "start_time": "2022-01-15T20:33:26.450147", "status": "completed"} tags=[]
# ## Build the model...
#
# Run this cell if you want to perform a very limited grid search:
# + papermill={"duration": 0.022014, "end_time": "2022-01-15T20:33:26.500547", "exception": false, "start_time": "2022-01-15T20:33:26.478533", "status": "completed"} tags=[]
PERFORM_GRID_SEARCH = True
# + [markdown] papermill={"duration": 0.014159, "end_time": "2022-01-15T20:33:26.529322", "exception": false, "start_time": "2022-01-15T20:33:26.515163", "status": "completed"} tags=[]
# Run this cell if you **do not** want to perform grid search:
# + papermill={"duration": 0.02146, "end_time": "2022-01-15T20:33:26.565113", "exception": false, "start_time": "2022-01-15T20:33:26.543653", "status": "completed"} tags=[]
PERFORM_GRID_SEARCH = False
# + [markdown] papermill={"duration": 0.014069, "end_time": "2022-01-15T20:33:26.593734", "exception": false, "start_time": "2022-01-15T20:33:26.579665", "status": "completed"} tags=[]
# Change `PERFORM_HUGE_GRID_SEARCH` to `True` if you want to do an extensive grid search.
#
# Be aware: this takes many hours on a four-core CPU!
# + papermill={"duration": 0.020183, "end_time": "2022-01-15T20:33:26.628512", "exception": false, "start_time": "2022-01-15T20:33:26.608329", "status": "completed"} tags=[]
PERFORM_HUGE_GRID_SEARCH = False
# + papermill={"duration": 0.036838, "end_time": "2022-01-15T20:33:26.679613", "exception": false, "start_time": "2022-01-15T20:33:26.642775", "status": "completed"} tags=[]
# Split the dataset into training and test...
xt, xv, yt, yv = train_test_split(
df.drop("HeartDisease", axis = 1),
df["HeartDisease"],
test_size = 0.2,
random_state = 42,
shuffle = True,
stratify = df["HeartDisease"]
)
# Define the data preparation, feature
# selection and classification pipeline
pipeline = Pipeline(steps = [
("transform", ColumnTransformer(
transformers = [
("cat", OrdinalEncoder(), categorical_columns),
("num", MinMaxScaler(), numerical_columns)
]
)
),
("features", SelectKBest()),
("classifier", XGBClassifier(
objective = "binary:logistic", eval_metric = "auc", use_label_encoder = False
)
)
])
if PERFORM_GRID_SEARCH:
# Define our search space for grid search...
# Short search over gamma as a quick example...
search_space = [{
"classifier__n_estimators": [60],
"classifier__learning_rate": [0.1],
"classifier__max_depth": [4],
"classifier__colsample_bytree": [0.2],
"classifier__gamma": [i / 10.0 for i in range(3, 7)],
"features__score_func": [chi2],
"features__k": [10],
}]
if PERFORM_HUGE_GRID_SEARCH:
# Define our search space for grid search...
# This is a real search but takes hours...
search_space = [{
"classifier__n_estimators": [i*10 for i in range(1, 10)],
"classifier__learning_rate": [0.01, 0.05, 0.1, 0.2],
"classifier__max_depth": range(1, 10, 1),
"classifier__colsample_bytree": [i/20.0 for i in range(7)],
"classifier__gamma": [i / 10.0 for i in range(3, 7)],
"features__score_func": [chi2],
"features__k": [10],
}]
# Define grid search...
grid = GridSearchCV(
pipeline,
param_grid = search_space,
# Define cross validation...
cv = KFold(n_splits = 10, random_state = 917, shuffle = True),
# Define AUC and accuracy as score...
scoring = {
"AUC": "roc_auc",
"Accuracy": make_scorer(accuracy_score)
},
refit = "AUC",
verbose = 1,
n_jobs = -1
)
# Fit grid search
grid_model = grid.fit(xt, yt)
yp = grid_model.predict(xv)
#
print(f"Best AUC Score: {grid_model.best_score_}")
print(f"Accuracy: {accuracy_score(yv, yp)}")
print("Confusion Matrix: ", confusion_matrix(yv, yp))
print("Best Parameters: ", grid_model.best_params_)
# + papermill={"duration": 0.302103, "end_time": "2022-01-15T20:33:26.996264", "exception": false, "start_time": "2022-01-15T20:33:26.694161", "status": "completed"} tags=[]
# Use the new best parameters if they were computed
# else use the previously computed best parameters.
# These produced an AUC score of 0.9244531360448315
# and an accuracy of 0.8858695652173914.
parameters = {
"classifier__colsample_bytree": 0.2,
"classifier__gamma": 0.4,
"classifier__learning_rate": 0.1,
"classifier__max_depth": 4,
"classifier__n_estimators": 60,
"features__k": 10,
"features__score_func": chi2
}
if PERFORM_GRID_SEARCH:
parameters = grid_model.best_params_
pipeline.set_params(**parameters)
model = pipeline.fit(xt, yt)
yp = model.predict(xv)
print(f"Accuracy: {accuracy_score(yv, yp)}")
print("Confusion Matrix: ", confusion_matrix(yv, yp))
print("Prediction: ", yp)
# + [markdown] papermill={"duration": 0.014281, "end_time": "2022-01-15T20:33:27.025307", "exception": false, "start_time": "2022-01-15T20:33:27.011026", "status": "completed"} tags=[]
# ## Plotting the Importance and First Five Trees
# Here I demonstrate the use of **`PDSUtilities.xgboost.plot_importance()`** and **`PDSUtilities.xgboost.plot_tree()`** on the `xgboost` model above...
# + papermill={"duration": 1.912424, "end_time": "2022-01-15T20:33:28.95229", "exception": false, "start_time": "2022-01-15T20:33:27.039866", "status": "completed"} tags=[]
classifier = pipeline["classifier"]
trees = [tree for tree in classifier.get_booster()]
# print(pio.templates["presentation"])
features = [LABELS[column] for column in df.drop("HeartDisease", axis = 1).columns]
fig = plot_importance(classifier, features = features)
fig.update_layout(template = "presentation")
fig.update_layout(width = 700, height = 600)
fig.update_layout(
margin = { 'l': 150 }, #, 'r': 10, 't': 50, 'b': 10 },
)
# This is literally the dumbest thing I've seen in years...
# This puts space between the ticks and tick labels. SMFH.
fig.update_yaxes(ticksuffix = " ")
fig.show()
booster = classifier.get_booster()
print("Plotting the first five trees:")
for tree in range(0, np.minimum(5, len(trees))):
title = f"Tree {tree}"
grayscale = tree % 2 == 0
edge_labels = { 'Yes/Missing': "Yes" }
fig = plot_tree(booster, tree, features = features, grayscale = grayscale, edge_labels = edge_labels)
fig.update_layout(
margin = { 'l': 10, 'r': 10, 't': 50, 'b': 10 },
title = { 'text': title, 'x': 0.5, 'xanchor': "center" },
)
fig.show()
# + papermill={"duration": 0.080827, "end_time": "2022-01-15T20:33:29.115468", "exception": false, "start_time": "2022-01-15T20:33:29.034641", "status": "completed"} tags=[]
| heart-disease-prediction-with-xgboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy.stats
inverse_cdf = scipy.stats.norm.ppf
cdf = scipy.stats.norm.cdf
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("notebook", font_scale=2.0)
# # Multivariate normal PDF
# $$
# \text{MVNormalPDF}(x | \mu, \Sigma) = \frac{1}{(2\pi)^{D/2}} \frac{1}{\text{det}(\Sigma)^{1/2}} \exp{-\frac{1}{2} (x-\mu)^T \Sigma^{-1} (x-\mu) }
# $$
# # Plot the contours of the PDF in 2D space
# +
def plotGauss2DContour(
mu_D, cov_DD,
color='b',
prob_mass_inside_contour_grid=[0.25, 0.75, 0.99],
markersize=3.0,
unit_circle_radian_step_size=0.03,
ax_handle=None,
):
''' Plot elliptical contours for provided mean mu, covariance Sigma.
Args
----
mu_D : 1D numpy array, shape (D,) = (2,)
Must be a 2-d mean vector
cov_DD : 2D numpy array, shape (D,D) = (2,2)
Must be a 2x2 symmetric, positive definite covariance
prob_mass_inside_contour_grid : list
How much probability mass should lie inside each contour
Post Condition
--------------
Plot created on current axes (or the provided axis)
'''
# If user provided an axis, use that
if ax_handle is not None:
plt.sca(ax_handle)
mu_D = np.squeeze(np.asarray(mu_D))
cov_DD = np.asarray(cov_DD)
assert mu_D.shape == (2,)
assert cov_DD.shape == (2,2)
# Decompose cov matrix into eigenvalues "lambda[d]" and eigenvectors "V[:,d]"
lambda_D, V_DD = np.linalg.eig(cov_DD)
sqrtCov_DD = np.dot(V_DD, np.sqrt(np.diag(lambda_D)))
# Prep for plotting elliptical contours
# by creating grid of G different (x,y) points along perfect circle
t_G = np.arange(-np.pi, np.pi, unit_circle_radian_step_size)
x_G = np.sin(t_G)
y_G = np.cos(t_G)
Zcirc_DG = np.vstack([x_G, y_G])
# Warp circle into ellipse defined by Sigma's eigenvectors
Zellipse_DG = np.dot(sqrtCov_DD, Zcirc_DG)
# Plot contour lines across several radius lengths
for prob_mass in sorted(prob_mass_inside_contour_grid):
# How large is the radius r?
# Need to set r such that area from (-r, r) for 1D Gaussian equals p
# Equivalently by symmetry we set: 2 * ( CDF(r) - CDF(0) ) = p
r = inverse_cdf(prob_mass/2 + cdf(0))
Z_DG = r * Zellipse_DG + mu_D[:, np.newaxis]
plt.plot(
Z_DG[0], Z_DG[1], '.',
markersize=markersize,
markerfacecolor=color,
markeredgecolor=color)
# +
mu_D = np.asarray([0, 0])
cov_DD = np.asarray([[1, 0 ], [0, 1]])
# First, plot N samples
N = 5000
x_N2 = scipy.stats.multivariate_normal.rvs(mu_D, cov_DD, N)
plt.plot(x_N2[:,0], x_N2[:,1], 'r.')
# Second, plot the contours
plotGauss2DContour(mu_D, cov_DD)
plt.gca().set_aspect('equal', 'box');
plt.gca().set_xlim([-5, 5]);
plt.gca().set_ylim([-5, 5]);
# +
mu_D = np.asarray([0, 0])
cov_DD = np.asarray([[3, 0 ], [0, 1]])
# First, plot N samples
N = 5000
x_N2 = scipy.stats.multivariate_normal.rvs(mu_D, cov_DD, N)
plt.plot(x_N2[:,0], x_N2[:,1], 'r.')
# Second, plot the contours
plotGauss2DContour(mu_D, cov_DD)
plt.gca().set_aspect('equal', 'box');
plt.gca().set_xlim([-5, 5]);
plt.gca().set_ylim([-5, 5]);
# +
mu_D = np.asarray([0, 0])
cov_DD = np.asarray([[1, -1.5], [-1.5, 4]])
# First, plot N samples
N = 5000
x_N2 = scipy.stats.multivariate_normal.rvs(mu_D, cov_DD, N)
plt.plot(x_N2[:,0], x_N2[:,1], 'r.')
# Second, plot the contours
plotGauss2DContour(mu_D, cov_DD)
plt.gca().set_aspect('equal', 'box');
plt.gca().set_xlim([-5, 5]);
plt.gca().set_ylim([-5, 5]);
# -
| notebooks/MultivariateNormalContourVisualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets('MNIST_data', one_hot=True)
sess = tf.InteractiveSession()
# # MNIST
#
# MNIST is a database of handwritten digits. The images are 28x28 greyscale images (encoded as 784-dimensional vectors in row-major order). There are 60,000 images in the training set, and there are 10,000 images in the test set.
#
# Why is there a train-test split? We care about how our function generalizes, and so we want to benchmark its performance on a set of data that it hasn't seen before. Otherwise, a "perfect" learning algorithm could just memorize all the data points, but this algorithm wouldn't generalize well.
#
# Let's see what one of the MNIST images looks like.
plt.imshow(mnist.test.images[0].reshape(28, 28), cmap='gray')
# Let's check the label of this image. The MNIST labels are encoded in one-hot format. There are 10 possible labels, and the vector with label $i$ is the $i$-dimensional vector that has the entry $1$ in the $i$th position and $0$s elsewhere.
mnist.test.labels[0]
np.argmax(mnist.test.labels[0])
# # Fully-connected neural network
# Let's design a fully-connected neural network to classify MNIST digits. It should take a 784-dimensional input and give a 10-dimensional one-hot encoded probability distribution as output.
x = tf.placeholder(tf.float32, (None, 28*28)) # batch of inputs
y_ = tf.placeholder(tf.float32, (None, 10)) # batch of corresponding labels
# +
# this corresponds with the model of a neuron in [ 02-01-notes ]
# except this is describing an entire layer, not a single neuron
# and we're not including the activation function inside here
def fully_connected(x, input_dimension, output_dimension):
raise NotImplementedError
# -
# Let's start with a really simple neural network with only one fully-connected layer (the output layer) with 10 neurons.
#
# See [ 02-04-notes ] for an architecture diagram.
# +
# TODO
# y = ???
# -
# Above, $y$ is a 10-dimensional vector, but it's not a probability distribution. We can fix that by applying the softmax function to the logits $y$:
#
# $$\sigma(y)_i = \frac{e^{y_i}}{\sum_j e^{y_j}}$$
# And we can define loss as the cross entropy between the true probability distribution (the labels) $p$ and the predicted probability distribution $q$:
#
# $$H(p, q) = - \sum_i p(x) \log q(x)$$
# In TensorFlow, we can do both of these in a single step (also needed for numerical stability):
loss = tf.nn.softmax_cross_entropy_with_logits(logits=y, labels=y_)
# ## Training
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.5).minimize(loss)
# We have 60000 training points, so we'll be doing minibatch stochastic gradient descent to train our network (instead of computing gradients over all 60000 data points).
# +
BATCH_SIZE = 100
ITERATIONS = 1000
sess.run(tf.global_variables_initializer())
for i in range(ITERATIONS):
x_batch, y_batch = mnist.train.next_batch(BATCH_SIZE)
sess.run(optimizer, {x: x_batch, y_: y_batch})
# -
# ## Evaluation
# Let's evaluate the accuracy of our network over the test set.
def accuracy(predictions, labels):
return np.mean(np.argmax(predictions, 1) == np.argmax(labels, 1))
predictions = y.eval({x: mnist.test.images})
accuracy(predictions, mnist.test.labels)
# # Deep fully-connected network
#
# Will adding a ton of parameters help us find a better solution? Let's use a deep fully-connected network using layers with 2000, 1000, and 100 neurons in the hidden layers and then 10 neurons in the output layer. Let's use ReLU activation for all the hidden layers. See [ 02-05-notes ] for an architecture diagram.
# +
# TODO
# y = ???
# -
# Our initial network had ~8,000 parameters. The above network has ~3.5 million parameters, which is over 400x the capacity of the first one.
loss = tf.nn.softmax_cross_entropy_with_logits(logits=y, labels=y_)
# ## Training
# Let's use a fancier optimizer this time.
optimizer = tf.train.AdamOptimizer().minimize(loss)
# +
BATCH_SIZE = 50
ITERATIONS = 5000 # this takes ~ 2 minutes on my laptop
sess.run(tf.global_variables_initializer())
for i in range(ITERATIONS):
x_batch, y_batch = mnist.train.next_batch(BATCH_SIZE)
l, _ = sess.run([loss, optimizer], {x: x_batch, y_: y_batch})
if (i+1) % 100 == 0:
print('iteration %d, batch loss %f' % (i+1, np.mean(l)))
# -
# ## Evaluation
predictions = y.eval({x: mnist.test.images})
accuracy(predictions, mnist.test.labels)
# # Convolutional neural network
#
# Let's design a convolutional neural network to classify MNIST digits.
x_image = tf.reshape(x, (-1, 28, 28, 1)) # turn our 784-dimensional vector into a 28x28 image
# +
# a convolutional layer
def convolve(x, kernel_height, kernel_width, input_channels, output_channels):
w = tf.Variable(tf.random_normal((kernel_height, kernel_width, input_channels, output_channels)))
b = tf.Variable(tf.random_normal((output_channels,)))
return tf.nn.conv2d(x, w, strides=(1, 1, 1, 1), padding='SAME')
# +
# a 2x2 max pooling layer
def pool(x):
return tf.nn.max_pool(x, (1, 2, 2, 1), (1, 2, 2, 1), padding='SAME')
# -
# ## Network architecture
#
# Let's design an architecture with the following layers:
#
# * convolution layer with 25 3x3 filters, relu activation
# * 2x2 max pooling layer
# * convolution layer with 50 3x3 filters, relu activation
# * 2x2 max pooling layer
# * fully-connected layer with 1000 neurons, relu activation
# * fully-connected output layer (10 neurons)
#
# See [ 02-06-notes ] for an architecture diagram.
# +
# TODO
# y = ???
# -
# This network has approximately 2.5 million parameters. Note that this is about 1 million parameters _fewer_ than the deep fully-connected network.
loss = tf.nn.softmax_cross_entropy_with_logits(logits=y, labels=y_)
# ## Training
optimizer = tf.train.AdamOptimizer().minimize(loss)
# +
BATCH_SIZE = 50
ITERATIONS = 5000 # this takes ~ 3.5 minutes on my laptop
sess.run(tf.global_variables_initializer())
for i in range(ITERATIONS):
x_batch, y_batch = mnist.train.next_batch(BATCH_SIZE)
l, _ = sess.run([loss, optimizer], {x: x_batch, y_: y_batch})
if (i+1) % 100 == 0:
print('iteration %d, batch loss %f' % (i+1, np.mean(l)))
# -
# ## Evaluation
predictions = y.eval({x: mnist.test.images})
accuracy(predictions, mnist.test.labels)
# ## Visualization
# Let's see what some intermediate activations look like
conv1_, conv2_ = sess.run([conv1, conv2], {x: mnist.test.images[0:1]})
# +
fig, ax = plt.subplots(2, 2)
ax[0, 0].imshow(conv1_[0,:,:,0], cmap='gray')
ax[0, 1].imshow(conv1_[0,:,:,1], cmap='gray')
ax[1, 0].imshow(conv1_[0,:,:,2], cmap='gray')
ax[1, 1].imshow(conv1_[0,:,:,3], cmap='gray')
# +
fig, ax = plt.subplots(2, 2)
ax[0, 0].imshow(conv2_[0,:,:,0], cmap='gray')
ax[0, 1].imshow(conv2_[0,:,:,1], cmap='gray')
ax[1, 0].imshow(conv2_[0,:,:,2], cmap='gray')
ax[1, 1].imshow(conv2_[0,:,:,3], cmap='gray')
| 02-03-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sys import path
path.append("../")
# %matplotlib inline
import matplotlib.pyplot as plt
# # Coastal Boundary Classifier algorithm
# This coastal boundary algorithm is used to classify a given pixel as either coastline or not coastline using a simple binary format like in the table before.
#
# <br>
#
# $\begin{array}{|c|c|}
# \hline
# 1& Coastline \\ \hline
# 0& Not Coastline \\ \hline
# \end{array}$
#
# <br>
#
#
# The algorithm makes a classification by examining surrounding pixels and making a determination based on how many pixels around it are water
#
# <br>
#
# 
#
# <br>
#
# If the count of land pixels surrounding a pixel exceeds 5, then it's likely not coastline.
# If the count of land pixels surrounding a pixel does not exceed 1, then it's likely not a coastline
#
# <br>
#
# $$
# Classification(pixel) = \begin{cases}
# 1 & 2\le count\_water\_surrounding(pixel) \leq 5 \\
# 0 &
# \end{cases}
# $$
#
# <br>
#
#
# ### Counting by applying a convolutional kernel
#
# A convolution applies a `kernel` to a point and it's surrounding pixels. Then maps the product to a new grid.
#
#
# 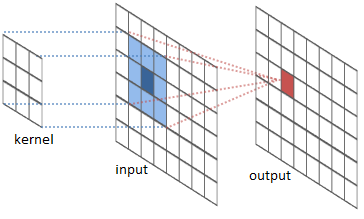
#
#
#
#
#
# In the case of coastal boundary classification, A convolution the following kernel is applied to a grid of `water`, `not-water` pixels.
#
# <br>
#
#
# $$
# Kernel =
# \begin{bmatrix}
# 1 & 1 & 1\\
# 1 & 0 & 1\\
# 1 & 1 & 1\\
# \end{bmatrix}
# $$
#
# <br>
# There exist more complicated differential kernels that would also work( see [sobel operator](https://en.wikipedia.org/wiki/Sobel_operator)).
# The one used in this notebooks however, operates on binary variables, and is easier to work with and easy to debug.
# ### Defined in code
# +
import scipy.ndimage.filters as conv
def _coastline_classification(dataset, water_band='wofs'):
kern = np.array([[1, 1, 1], [1, 0.001, 1], [1, 1, 1]])
convolved = conv.convolve(dataset[water_band], kern, mode='constant') // 1
ds = dataset.where(convolved > 0)
ds = ds.where(convolved < 6)
ds.wofs.values[~np.isnan(ds.wofs.values)] = 1
ds.wofs.values[np.isnan(ds.wofs.values)] = 0
ds.rename({"wofs": "coastline"}, inplace=True)
return ds
# -
# <br>
# # Test Algorithm
# ## Togo
lon = (1.1919, 1.4509)
lat = (6.0809, 6.2218)
#
# International agencies like World Bank, UNEP, and USAID are currently reporting and addressing the problem of coastal erosion near Lomé, Togo. The links listed below are references from these agencies regarding coastal erosion in Togo and coastal erosion as a world wide phenomena.
#
# - *"West Africa Coastal Erosion Project launched in Togo" (2016) * [- link](http://www.ndf.fi/news/west-africa-coastal-erosion-project-launched-togo)
# - *Agreement to Erosion Adaptation Project* (2016) [- link](http://pubdocs.worldbank.org/en/493191479316551864/WACA-Lom%C3%A9-Communiqu%C3%A9-2016-English.pdf)
# - World Bank WACA program brochure (2015) [- link](http://pubdocs.worldbank.org/en/622041448394069174/1606426-WACA-Brochure.pdf)
# - UNEP - Technologies for climate change adaption (2010) [- link](http://www.unep.org/pdf/TNAhandbook_CoastalErosionFlooding.pdf)
# - USAID - Adapting to Coastal Climate Change (2009) - [- link](http://www.crc.uri.edu/download/CoastalAdaptationGuide.pdf)
# - UNEP - Coastal Erosion and Climate Change in Western Africa(2002) - [- link](http://www.unep.org/dewa/africa/publications/aeo-1/121.htm)
#
from utils.data_cube_utilities.dc_display_map import display_map
display_map(latitude = lat, longitude = lon)
# ### Loading Togo imagery
# The following lines are needed to pull Togo imagery from our data-cube.
platform = 'LANDSAT_7'
product_type = 'ls7_ledaps_togo'
# +
from datetime import datetime
params = dict(platform=platform,
product=product_type,
time=(datetime(2007,1,1), datetime(2007,12,31)) ,
lon= lon,
lat= lat,
measurements = ['red', 'green', 'blue', 'nir', 'swir1', 'swir2', 'pixel_qa'] )
# -
import datacube
dc = datacube.Datacube(app = "Coastline classification", config = "/home/localuser/.datacube.conf")
# +
dataset = dc.load(**params)
# -
# ### Create a usable composite of 2017 dataset
# The imagery displayed below is noisy, cloudy, and bares artifacts of a sensor malfunction(scanlines)
#
# <br>
# +
def figure_ratio(ds, fixed_width = 10):
width = fixed_width
height = len(ds.latitude) * (fixed_width / len(ds.longitude))
return (width, height)
dataset.isel(time = 1).swir1.plot(cmap = "Greys", figsize = figure_ratio(dataset, fixed_width = 20))
# -
# The following code creates a composite that reduces all acquisitions in 2017, to a single cloud free statistical representation.
# +
import numpy as np
def mask_water_and_land(dataset):
#Create boolean Masks for clear and water pixels
clear_pixels = dataset.pixel_qa.values == 2 + 64
water_pixels = dataset.pixel_qa.values == 4 + 64
a_clean_mask = np.logical_or(clear_pixels, water_pixels)
return a_clean_mask
# +
from utils.data_cube_utilities.dc_mosaic import create_median_mosaic
def mosaic(dataset):
cloud_free_boolean_mask = mask_water_and_land(dataset)
return create_median_mosaic(dataset, clean_mask = cloud_free_boolean_mask)
# -
composited_dataset = mosaic(dataset)
# **Visualize Composited imagery**
composited_dataset.swir1.plot(cmap = "Greys", figsize = figure_ratio(dataset, fixed_width = 20))
# # water classification
# +
from utils.data_cube_utilities.dc_water_classifier import wofs_classify
water_classification = wofs_classify(composited_dataset, mosaic = True)
# -
water_classification
water_classification.wofs.plot(cmap = "Blues", figsize = figure_ratio(dataset, fixed_width = 20))
# <br>
# # run coastline classifier on water classification
coast = _coastline_classification(water_classification, water_band='wofs')
# <br>
coast
# <br>
coast.coastline.plot(cmap = "Blues", figsize = figure_ratio(dataset, fixed_width = 20))
| Coastal Change/Coastline_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Title
#
# Current Issue
#
# 1. ~~def revise_with_hand_edits(label_dict, row): [called with inherit_changes()] breaks because row isn't defined~~
# 2. ~~Does revised_xml include all metadata as well?~~
# 3. ~~Generate download button after revising (with download capability, of course)~~
# 4. Do I need to supply metadata (below, it's being erased):
#
# \<?xml version="1.0" encoding="UTF-8"?\>
# \<?xml-model href="http://www.masshist.org/publications/pub/schema/codem-0.2-djqa.rng" type="application/xml" schematypens="http://relaxng.org/ns/structure/1.0"?\>
#
# To Do
# 1. Ensure suggested encoding does not duplicate encoding already there (I believe up conversion handles this)
# 2. Work on KWIC View (this might be something that happens in the first application)
# 3. Expand NER search beyond \<p>
# +
import warnings, re, glob, datetime, csv, sys, os, base64, io, spacy
import pandas as pd
import numpy as np
from io import BytesIO
from lxml import etree
import dash, dash_table
import dash_core_components as dcc
from dash.dependencies import Input, Output, State
import dash_html_components as html
from jupyter_dash import JupyterDash
from urllib.parse import quote
from flask import Flask, send_from_directory, send_file
# Import spaCy language model.
nlp = spacy.load('en_core_web_sm')
# Ignore simple warnings.
warnings.simplefilter('ignore', DeprecationWarning)
# Declare directory location to shorten filepaths later.
abs_dir = "/Users/quinn.wi/Documents/SemanticData/"
# -
# ## Functions
# +
# %%time
"""
XML Parsing Function: Get Namespaces
"""
def get_namespace(root):
namespace = re.match(r"{(.*)}", str(root.tag))
ns = {"ns":namespace.group(1)}
return ns
"""
XML Parsing Function: Retrieve XPaths
"""
def get_abridged_xpath(child):
while child.getparent().get('{http://www.w3.org/XML/1998/namespace}id') is None:
elem = child.getparent()
if elem.getparent().get('{http://www.w3.org/XML/1998/namespace}id') is not None:
ancestor = elem.getparent().tag
xml_id = elem.getparent().get('{http://www.w3.org/XML/1998/namespace}id')
abridged_xpath = f'.//ns:body//{ancestor}[@xml:id="{xml_id}"]/{child.tag}'
return abridged_xpath
"""
XML Parsing Function: Get Encoded Content
"""
def get_encoding(elem):
encoding = etree.tostring(elem, pretty_print = True).decode('utf-8')
encoding = re.sub('\s+', ' ', encoding) # remove additional whitespace
return encoding
"""
XML & Regex: Up Conversion
Function replaces all spaces between beginning and end tags with underscores.
Then, function wraps each token (determined by whitespace) with word tags (<w>...</w>)
"""
def up_convert_encoding(column):
# Regularize spacing & store data as new variable ('converted_encoding').
converted_encoding = re.sub('\s+', ' ', column, re.MULTILINE)
# Create regex that replaces spaces with underscores if spaces occur within tags.
# This regex treats tags as a single token later.
tag_regex = re.compile('<(.*?)>')
# Accumulate underscores through iteration
for match in re.findall(tag_regex, column):
replace_space = re.sub('\s', '_', match)
converted_encoding = re.sub(match, replace_space, converted_encoding)
# Up-Converstion
# Tokenize encoding and text, appending <w> tags, and re-join.
converted_encoding = converted_encoding.split(' ')
for idx, item in enumerate(converted_encoding):
item = '<w>' + item + '</w>'
converted_encoding[idx] = item
converted_encoding = ' '.join(converted_encoding)
return converted_encoding
"""
XML Parsing Function: Suggest New Encoding with Hand Edits
Similar to make_ner_suggestions(), this function folds in revision using regular expressions.
The outcome is the previous encoding with additional encoded information determined by user input.
Expected Columns:
previous_encoding
entities
accept_changes
make_hand_edits
"""
def revise_with_hand_edits(label_dict, make_hand_edits,
label, entity, previous_encoding, new_encoding):
label = label_dict[label]
# Up convert PREVIOUS ENCODING: assumes encoder will supply new encoding and attribute with value.
converted_encoding = up_convert_encoding(previous_encoding)
converted_entity = ' '.join(['<w>' + e + '</w>' for e in entity.split(' ')])
# If there is a unique id to add & hand edits...
if make_hand_edits != '':
entity_regex = re.sub('<w>(.*)</w>', '(\\1)(.*?</w>)', converted_entity)
entity_match = re.search(entity_regex, converted_encoding)
identifier_regex = re.search('<(.+)>(.+)</.+>', make_hand_edits, re.VERBOSE|re.DOTALL)
print (converted_entity, '\t', identifier_regex)
revised_encoding = re.sub(f'{entity_match.group(0)}',
f'<{label}>{entity_match.group(1)}</{label}>{entity_match.group(2)}',
converted_encoding)
revised_encoding = re.sub(f'<{label}>', f'<{identifier_regex.group(1)}>', revised_encoding)
# Clean up any additional whitespace and remove word tags.
revised_encoding = re.sub('\s+', ' ', revised_encoding, re.MULTILINE)
revised_encoding = re.sub('<[/]?w>', '', revised_encoding)
revised_encoding = re.sub('_', ' ', revised_encoding) # Remove any remaining underscores in tags.
revised_encoding = re.sub('“', '"', revised_encoding) # Change quotation marks to correct unicode.
revised_encoding = re.sub('”', '"', revised_encoding)
return revised_encoding
else:
pass
"""
XML & NER: Update/Inherit Accepted Changes
Expects a dataframe (from a .csv) with these columns:
file
abridged_xpath
previous_encoding
entities
new_encoding
accept_changes
make_hand_edits
"""
def inherit_changes(label_dict, dataframe):
dataframe = dataframe.fillna('')
for index, row in dataframe.iterrows():
# If HAND changes are accepted...
if row['make_hand_edits'] != '':
revised_by_hand = revise_with_hand_edits(label_dict,
row['make_hand_edits'],
row['label'], row['entity'],
row['previous_encoding'], row['new_encoding'])
dataframe.loc[index, 'new_encoding'] = revised_by_hand
try:
if dataframe.loc[index + 1, 'abridged_xpath'] == row['abridged_xpath'] \
and dataframe.loc[index + 1, 'descendant_order'] == row['descendant_order']:
dataframe.loc[index + 1, 'previous_encoding'] = row['new_encoding']
else:
dataframe.loc[index, 'new_encoding'] = revised_by_hand
except KeyError as e:
dataframe.loc[index, 'new_encoding'] = revised_by_hand
# If NER suggestions are accepted as-is...
elif row['accept_changes'] == 'y' and row['make_hand_edits'] == '':
try:
if dataframe.loc[index + 1, 'abridged_xpath'] == row['abridged_xpath'] \
and dataframe.loc[index + 1, 'descendant_order'] == row['descendant_order']:
dataframe.loc[index + 1, 'previous_encoding'] = row['new_encoding']
else:
dataframe.loc[index, 'new_encoding'] = row['new_encoding']
except KeyError as e:
dataframe.loc[index, 'new_encoding'] = row['new_encoding']
# If changes are rejected...
else:
try:
if dataframe.loc[index + 1, 'abridged_xpath'] == row['abridged_xpath'] \
and dataframe.loc[index + 1, 'descendant_order'] == row['descendant_order']:
dataframe.loc[index + 1, 'previous_encoding'] = dataframe.loc[index, 'previous_encoding']
except KeyError as e:
dataframe.loc[index, 'new_encoding'] = dataframe.loc[index, 'previous_encoding']
# Subset dataframe with finalized revisions.
dataframe = dataframe.groupby(['abridged_xpath', 'descendant_order']).tail(1)
return dataframe
"""
XML & NER: Write New XML File with Accepted Revisions
Expects:
XML File with Original Encoding
CSV File with Accepted Changes
Label Dictionary
"""
def revise_xml(xml_file, csv_df, label_dict):
# First, update data to reflect accepted changes.
new_data = inherit_changes(label_dict, csv_df)
root = etree.fromstring(xml_file)
ns = get_namespace(root)
tree_as_string = etree.tostring(root, pretty_print = True).decode('utf-8')
tree_as_string = re.sub('\s+', ' ', tree_as_string) # remove additional whitespace
# Write accepted code into XML tree.
for index, row in new_data.iterrows():
original_encoding_as_string = row['previous_encoding']
# Remove namespaces within tags to ensure regex matches accurately.
original_encoding_as_string = re.sub('^<(.*?)( xmlns.*?)>(.*)$',
'<\\1>\\3',
original_encoding_as_string)
accepted_encoding_as_string = row['new_encoding']
accepted_encoding_as_string = re.sub('<(.*?)( xmlns.*?)>(.*)$',
'<\\1>\\3',
accepted_encoding_as_string) # Remove namespaces within tags.
tree_as_string = re.sub(original_encoding_as_string,
accepted_encoding_as_string,
tree_as_string)
# Check well-formedness (will fail if not well-formed)
doc = etree.fromstring(tree_as_string)
et = etree.ElementTree(doc)
# Convert to string.
et = etree.tostring(et, encoding='unicode', method='xml', pretty_print = True)
return et
"""
App Function: Write Filename when Uploaded
"""
def parse_contents(contents, filename):
return html.Div([
html.H4(f'{filename} succesfully uploaded')
])
"""
App Function: Make XML Revisions and Provide Download Link
"""
def commit_xml_revisions(xml_contents, csv_contents, xml_filename, csv_filename):
try:
# XML Contents
xml_content_type, xml_content_string = xml_contents.split(',')
xml_decoded = base64.b64decode(xml_content_string).decode('utf-8')
xml_file = xml_decoded.encode('utf-8')
except Exception as e:
return html.Div([f'There was an error processing {xml_filename}: {e}.'])
try:
# CSV Contents
csv_content_type, csv_content_string = csv_contents.split(',')
csv_decoded = base64.b64decode(csv_content_string).decode('utf-8')
csv_df = pd.read_csv(io.StringIO(csv_decoded))
except Exception as e:
return html.Div([f'There was an error processing {csv_filename}: {e}.'])
try:
revised_data = revise_xml(xml_file, csv_df, label_dict)
return revised_data
except Exception as e:
return html.P(f'Error with Revision Process: {e}')
# -
# ## APP
# +
# %%time
label_dict = {
'PERSON':'persName',
'LOC':'placeName', # Non-GPE locations, mountain ranges, bodies of water.
'GPE':'placeName', # Countries, cities, states.
'FAC':'placeName', # Buildings, airports, highways, bridges, etc.
'ORG':'orgName', # Companies, agencies, institutions, etc.
'NORP':'name', # Nationalities or religious or political groups.
'EVENT':'name', # Named hurricanes, battles, wars, sports events, etc.
'WORK_OF_ART':'name', # Titles of books, songs, etc.
'LAW':'name', # Named documents made into laws.
}
server = Flask(__name__)
app = JupyterDash(__name__, server = server)
# Layout.
app.layout = html.Div([
# Title
html.H1('XML & NER Revision'),
# Upload Data Area.
html.H2('Upload Original XML File'),
dcc.Upload(
id = 'upload-xml',
children = html.Div([
'Drag and Drop or ', html.A('Select XML File')
]),
style={
'width': '95%',
'height': '60px',
'lineHeight': '60px',
'borderWidth': '1px',
'borderStyle': 'dashed',
'borderRadius': '5px',
'textAlign': 'center',
'margin': '10px'
},
multiple = True # Allow multiple files to be uploaded
),
html.Div(id = 'xml-data-upload'),
# Upload Data Area.
html.H2('Upload CSV File with Accepted Changes'),
dcc.Upload(
id = 'upload-csv',
children = html.Div([
'Drag and Drop or ', html.A('Select CSV File')
]),
style={
'width': '95%',
'height': '60px',
'lineHeight': '60px',
'borderWidth': '1px',
'borderStyle': 'dashed',
'borderRadius': '5px',
'textAlign': 'center',
'margin': '10px'
},
multiple = True # Allow multiple files to be uploaded
),
html.Div(id = 'csv-data-upload'),
html.Div(id = 'text-area', children = []),
html.Div(id = 'button-area', n_clicks = 0, children = []),
html.Div(id = 'download-area', children = [])
])
# App Callbacks
# Write XML Filename when Uploaded
@app.callback(
Output('xml-data-upload', 'children'),
Input('upload-xml', 'contents'),
State('upload-xml', 'filename')
)
def update_xml_file(xml_contents, xml_filename):
if xml_contents is not None:
xml_children = [
parse_contents(xc, xf) for xc, xf in zip(xml_contents, xml_filename)
]
return xml_children
# Write CSV Filename when Uploaded
@app.callback(
Output('csv-data-upload', 'children'),
Input('upload-csv', 'contents'),
State('upload-csv', 'filename')
)
def update_csv_file(csv_contents, csv_filename):
if csv_contents is not None:
csv_children = [
parse_contents(cc, cf) for cc, cf in zip(csv_contents, csv_filename)
]
return csv_children
"""
Show Revisions
"""
# Print Revisions.
@app.callback(
Output('text-area', 'children'),
[Input('upload-xml', 'contents'), Input('upload-csv', 'contents')],
[State('upload-xml', 'filename'), State('upload-csv', 'filename')]
)
def print_revisions(xml_contents, csv_contents, xml_filename, csv_filename):
if xml_contents and csv_contents:
# Incoporate Revisions
revisions = [
commit_xml_revisions(xc, cc, xf, cf) for xc, cc, xf, cf in
zip(xml_contents, csv_contents, xml_filename, csv_filename)
]
return revisions
else:
return 'Waiting for Required Files'
"""
Generate Download Option When Possible
"""
# Generates a download button for the resource
def build_download_button(uri):
button = html.Form(
id = 'download-button',
action = uri,
method="get",
children=[
html.Button(
className="button",
type="submit",
children=["Download Revised File"]
)]
)
return button
# Downloads with Button Click
@app.callback(
Output("button-area", "children"),
[Input("text-area", "children"), Input('upload-xml', 'filename')]
)
def show_download_button(text, filename):
if text != 'Waiting for Required Files':
path = f"{filename[0]}"
return [build_download_button(path)]
@app.callback(
Output('download-area', 'children'),
[Input('button-area', 'n_clicks'), Input('text-area', 'children'), Input('upload-xml', 'filename')]
)
def download_with_button_click(n_clicks, text, filename):
if n_clicks:
revised_content = ' '.join(text)
path = f"{filename[0]}"
with open(path, "w") as file:
file.write(revised_content)
return f'{filename[0]} downloaded!'
if __name__ == "__main__":
app.run_server(mode = 'inline', debug = True) # mode = 'inline' for JupyterDash
# -
# ## Test Area
# +
# %%time
test = pd.DataFrame({'file':[0,0,0,0,0,0,0,0,0,0],
'abridged_xpath':[1, 1, 1, 2, 2, 2, 3, 3, 4, 4],
'previous_encoding':['a', 'a', 'a', 'b', 'b', 'b', 'c', 'c', 'd', 'Going to Boston for the holiday'],
'new_encoding':['k','l','m','n', 'o','p','q <LOC>c</LOC>', 'r', 's', 't'],
'accept_changes':['y','n','y','n','y','y','y','y','y','y'],
'entities':["('a','LOC')", "('a','LOC')", "('a','LOC')", "('b','LOC')", "('b','LOC')", "('b','LOC')", "('c','LOC')", "('c','LOC')", "('d','LOC')", "('Boston','LOC')"],
'entity':['a', 'a', 'a', 'b', 'b', 'b', 'c', 'c', 'd', 'Boston'],
'label':['LOC','LOC','LOC','LOC','LOC','LOC','LOC','LOC','LOC','LOC'],
'make_hand_edits':['','','','','','','','','','<placeName>Boston</placeName>'],
'add_unique_identifier':['','','','','','','c-2','','','boston-1']})
for index, row in test.iterrows():
new = inherit_changes(label_dict, test)
print (new)
test
# +
# %%time
cf = abs_dir + 'Data/TestEncoding/EditingData/Data.csv'
csv_df = pd.read_csv(cf)
new_data = inherit_changes(label_dict, csv_df)
# filename = abs_dir + "Data/TestEncoding/EditingData/test_xml-before.xml"
# xml_file = open(filename).read()
# root = etree.fromstring(xml_file.encode('utf-8'))
# ns = get_namespace(root)
# tree_as_string = etree.tostring(r oot, pretty_print = True).decode('utf-8')
# tree_as_string = re.sub('\s+', ' ', tree_as_string) # remove additional whitespace
# -
new_data
csv_df
for index, row in csv_df.iterrows():
try:
if csv_df.loc[index + 1, 'abridged_xpath'] == row['abridged_xpath'] \
and csv_df.loc[index + 1, 'descendant_order'] == row['descendant_order']:
print ('Descendant orders equivalent')
else:
print ('Recognizes different elements')
except KeyError as e:
print ('Last row behavior')
| Jupyter_Notebooks/Interfaces/NER_Application/Drafts/App_NER-XML-Updater.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## basic use
#
# the next cells show how the %cache magic should be used
#
# Note: This notebook requires the the packages scikit-learn, numpy and cache_magic to be installed
# +
import cache_magic
# delete everthing currently cached
# %cache --reset
# store a new value for a
# %cache a = "111"
# fetch the cached value for a
# %cache a = "111"
# +
# an examle for an actual use-case
import cache_magic
import numpy as np
from sklearn import svm
# %cache --reset
# %timeit -n 1 -r 5 %cache -v 1 clf = svm.LinearSVC().fit(np.random.randint(5, size=(5000, 40)), np.random.randint(5, size=(5000)))
# +
# the following 4 cases use the same version
# %cache -r
# without explicit version, the expression (=right hand site of assignment) is used as version
# %cache a = 0
# if parameter is an integer, it will be the version
# %cache -v 0 a = 1
# if parameter is a variable name, it's value is used as version
my_version = 0
# %cache -v my_version a = 1
# new and old version are converted into a string before comparing them
my_version_2 = "0"
# %cache -v my_version_2 a = 1
# +
# show everything, that is cached
# %cache
# generate some variables
# %cache b=3
def fun(x):
return x+1
# %cache c = fun(b)
# %cache -v c d = fun(1.1)
# show the new cache
# %cache
# -
# ## power use
#
# the next cells show how the %cache magic can be used
# +
import cache_magic
import numpy as np
from sklearn import svm
# %cache --reset
# even if the expression changes, but not the version, the old value will still be loaded
# in which case there will be a warning
# %cache -v 1 clf = svm.LinearSVC().fit(np.random.randint(5, size=(1000, 40)), np.random.randint(5, size=(1000)))
# %cache -v 1 clf = "not a classifier"
print(clf.predict(np.random.randint(5,size=(1,40)))[0])
# without an expression, it will always try to reload the cached value
del clf
# %cache -v 1 clf
print(clf.predict(np.random.randint(5,size=(1,40)))[0])
# you can store the current value of a var without an actual statement by assigning it to itself
clf="not a classifier"
# %cache -v 2 clf=clf
print(clf)
# +
# while the cache still exists in the file system, the cell can be executed alone
import cache_magic
# %cache clf
print(clf)
# +
# the cache is stored in the directory where the kernel was first started in
import cache_magic
import os
# %cache -r
# %cache a=1
# %cache b=1
# %cache c=1
# %cache
for root, dirs, files in os.walk(".cache_magic"):
# there is one folder per cache variable
print(root)
# if the working dir changes, the .cache-dir stays where it is
# %cd ..
# %cache
for root, dirs, files in os.walk(".cache_magic"):
# no output, because no .cache-dir
print(root)
# %cd -
# %cache
for root, dirs, files in os.walk(".cache_magic"):
# now we see the cache directory againg
print(root)
# +
# always store a new value and never read from cache
# %cache -r a=1
# remove a single variable from cache
# %cache -r a
# Error:
# %cache a
# load last value if possible, and store new value on miss
# %cache a = a
# +
# load last value if possible, but don't store new value on miss
import cache_magic
del a
# %cache a
# +
# You can use this magic-module as a regular python module
from cache_magic import CacheCall
cache = CacheCall(get_ipython().kernel.shell)
# setting all parameter by name
cache(
version="*",
reset=False,
var_name="aaa",
var_value="1+1",
show_all=False,
set_debug=True)
# setting all parameter by ordering
cache("1",False,"bbb","1+1",False, False)
# setting parameter selectivly
cache(show_all=True)
# -
# ## development tests
#
# the next cells show how the %cache magic should not be used
#
# these examples are for debug-purposes only
# +
#testing successfull calls
import cache_magic
# Dev-Note: use reload, so you don't have to restart the kernel everytime you change the
from imp import reload
reload(cache_magic)
my_version = 3
# %cache --reset
print(" exptecting: new values")
# %cache -v 2 a = "ex3"
# %cache -v my_version c = "ex3"
# %cache --version my_version sadsda = "ex3"
# %cache -v 3 a=""
# %cache -v 3 -r a=""
# %cache -v 3 -r a=""
print(" exptecting: warnings")
# %cache -v 3 a= " _ "
# %cache -v 3 sadsda = "ex4"
print(" exptecting: stored values")
# %cache -v my_version sadsda = "ex3"
# %cache -v 3 sadsda = "ex3"
# +
# testing errors
import cache_magic
reload(cache_magic)
# %cache -v "a" a = "ex3"
# %cache -v a 1=a
# +
# testing loading without storing
import cache_magic
reload(cache_magic)
# %cache --reset
a=1
del a
# error:
# %cache a
# %cache a=1
del a
# %cache a
del a
# error:
# %cache -v '1' a
# error
# %cache -v 1 a
# %cache --reset
a=1
del a
# Error
# %cache -v 0 a
# %cache -v * a=1
# Error:
# %cache -v * a
# +
# %cache -v "1" a
# +
# %cache -v 213 a = "1"
# get stored version via error message
# %cache -v * a
# testing debug flag '-d'
# %cache -d -v 1 -r a = "1"
# %cache -d a = "1"
# +
import cache_magic
from imp import reload
reload(cache_magic)
# %cache -r a=1
# %cache -r a
# %cache -r a=1
# +
import cache_magic
from imp import reload
reload(cache_magic)
# %cache -r a = 1
# %cache a
print (a)
# +
import cache_magic
from imp import reload
reload(cache_magic)
def foo(x):
return x+1
# %cache --reset
# %cache -v * a= foo(3)
# %cache -v * a= foo(3)
# %cache -v * a= 2
# +
# #!pip install -e .
| example.ipynb |
// # Automatic generation of Notebook using PyCropML
// This notebook implements a crop model.
// ### Model Preciprec
#include "..\..\src\cpp\STICS_SNOW\SnowState.cpp"
#include "..\..\src\cpp\STICS_SNOW\SnowRate.cpp"
#include "..\..\src\cpp\STICS_SNOW\SnowAuxiliary.cpp"
#include "..\..\src\cpp\STICS_SNOW\SnowExogenous.cpp"
#include "..\..\src\cpp\STICS_SNOW\Preciprec.cpp"
class Test
{
private:
SnowState s;
SnowState s1;
SnowRate r;
SnowAuxiliary a;
SnowExogenous ex;
Preciprec mod;
public:
//check snow model
// test_snow1
void test_snow1()
{
this->mod.setrho(100.0);
this->s.setSdepth(0.087);
this->s1.setSdepth(0.085);
this->r.setMrf(0.27);
this->r.setSnowaccu(0.23);
this->a.setprecip(5.2);
this->s1.setSdry(10.0);
this->s.setSdry(10.0);
this->s.setSwet(0.03);
this->s1.setSwet(5.0);
this->mod.Calculate_Model(s,s1, r, a, ex);
//preciprec: 4.5;
cout << "preciprec estimated :\n";
cout << "\t" << this->s.getpreciprec() << "\n";
};
}
Test t;
t.test_snow1();
| test/cpp/Preciprec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: IPython (Python 3)
# language: python
# name: python3
# ---
# +
import matplotlib
import pandas as pd
# %matplotlib inline
# -
import aggregate_mummer_results
import mummer_all_bins
# +
import numpy as np
import os
import sys
import glob
# -
sys.path.append('./support_files/')
import summarise_bins
sample_path = './mummer_results/Methylophilus_methylotrophus-79_Ga0081650_to_Methylophilus_methylotrophus-55_Ga0081633.tsv'
result = aggregate_mummer_results.prepare_result(sample_path)
result.head(3)
result.shape
result['% IDY'].plot.hist()
aggregate_mummer_results.plot_identity_vs_len(sample_path, save_path=None)
# + active=""
# aggregate_mummer_results.plot_IDY_vs_len(
# './mummer_results/Methylophilus_methylotrophus-79_Ga0081650_to_Methylophilus_methylotrophus-55_Ga0081633.tsv')
# +
#for fp in glob.glob('./mummer_results/*.tsv'):
# aggregate_mummer_results.plot_IDY_vs_len(fp)
#glob.glob('/path/to/dir/*.csv')
# -
result.head(2)
aggregate_mummer_results.percent_identity(result)
# +
test_samples = glob.glob('./mummer_results/*.tsv')[1:20]
i_res = aggregate_mummer_results.percent_idty_all_results(test_samples)
# -
i_res.head()
i_res.shape
i_res[['% identity']]
res_piv = i_res.pivot(index='query name', columns='ref name', values='% identity')
res_piv.fillna(value=0, inplace = True)
res_piv
import seaborn as sns
# This is just a toy for proof of principle of the pivot stuff.
sns.heatmap(res_piv)
pid = pd.read_csv('./percent_identities.tsv', sep='\t')
pid.shape
pid.head(2)
pid[['query name', 'ref name']].drop_duplicates().shape
# + active=""
# pid[pid[['query name', 'ref name']].duplicated() == True]['mummer file'].astype(str)
# -
pd.set_option('display.height', 1000)
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
pd.options.display.max_colwidth = 110
pid[pid[['query name', 'ref name']].duplicated() == True]['mummer file'].astype(str).shape
# ## This notebook and data dir was developed before I discovered naming problems.
#
# I am going to drop the problem name from analysis hereafter because I'm not willing to spend the time to really fix an old broken dir for a lab presentation:
#
# drop Methylotenera mobilis-49
# Drop Methylotenera mobilis-49 from all the analysis.
pid = pid[pid['query name'] != 'Methylotenera mobilis-49']
pid = pid[pid['ref name'] != 'Methylotenera mobilis-49']
pid[pid[['query name', 'ref name']].duplicated() == True]['mummer file'].astype(str)
# all files contain Methylotenera_mobilis-49
pid[pid[['query name', 'ref name']].duplicated() == True]['mummer file'].str.contains('Methylotenera_mobilis-49')
res_piv = pid[['query name', 'ref name','% identity']
].pivot(index='query name',
columns = 'ref name', values = '% identity')
res_piv
p = sns.heatmap(res_piv)
p.figure.set_size_inches(15, 15)
p.figure.tight_layout()
p.figure.savefig("160528_fauzi_bin_ANIs.pdf")
| compare_fauzi_bins/Develop_mummer_aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Notebook written by [<NAME>](https://github.com/zhedongzheng)
#
# <img src="transformer.png" width="250">
#
# [Attention Is All You Need](https://arxiv.org/abs/1706.03762)
# +
from bunch import Bunch
from collections import Counter
import json
import numpy as np
import tensorflow as tf
# -
args = Bunch({
'source_max_len': 10,
'target_max_len': 20,
'min_freq': 50,
'hidden_units': 128,
'num_blocks': 2,
'num_heads': 8,
'dropout_rate': 0.1,
'batch_size': 64,
'position_encoding': 'non_param',
'activation': 'relu',
'tied_proj_weight': True,
'tied_embedding': True,
'label_smoothing': False,
'lr_decay_strategy': 'exp',
})
class DataLoader:
def __init__(self, source_path, target_path):
self.source_words = self.read_data(source_path)
self.target_words = self.read_data(target_path)
self.source_word2idx = self.build_index(self.source_words)
self.target_word2idx = self.build_index(self.target_words, is_target=True)
def read_data(self, path):
with open(path, 'r', encoding='utf-8') as f:
return f.read()
def build_index(self, data, is_target=False):
chars = [char for line in data.split('\n') for char in line]
chars = [char for char, freq in Counter(chars).items() if freq > args.min_freq]
if is_target:
symbols = ['<pad>','<start>','<end>','<unk>']
return {char: idx for idx, char in enumerate(symbols + chars)}
else:
symbols = ['<pad>','<unk>'] if not args.tied_embedding else ['<pad>','<start>','<end>','<unk>']
return {char: idx for idx, char in enumerate(symbols + chars)}
def pad(self, data, word2idx, max_len, is_target=False):
res = []
for line in data.split('\n'):
temp_line = [word2idx.get(char, word2idx['<unk>']) for char in line]
if len(temp_line) >= max_len:
if is_target:
temp_line = temp_line[:(max_len-1)] + [word2idx['<end>']]
else:
temp_line = temp_line[:max_len]
if len(temp_line) < max_len:
if is_target:
temp_line += ([word2idx['<end>']] + [word2idx['<pad>']]*(max_len-len(temp_line)-1))
else:
temp_line += [word2idx['<pad>']] * (max_len - len(temp_line))
res.append(temp_line)
return np.array(res)
def load(self):
source_idx = self.pad(self.source_words, self.source_word2idx, args.source_max_len)
target_idx = self.pad(self.target_words, self.target_word2idx, args.target_max_len, is_target=True)
return source_idx, target_idx
# +
def layer_norm(inputs, epsilon=1e-8):
mean, variance = tf.nn.moments(inputs, [-1], keep_dims=True)
normalized = (inputs - mean) / (tf.sqrt(variance + epsilon))
params_shape = inputs.get_shape()[-1:]
gamma = tf.get_variable('gamma', params_shape, tf.float32, tf.ones_initializer())
beta = tf.get_variable('beta', params_shape, tf.float32, tf.zeros_initializer())
outputs = gamma * normalized + beta
return outputs
def embed_seq(inputs, vocab_size, embed_dim, zero_pad=False, scale=False):
lookup_table = tf.get_variable('lookup_table', dtype=tf.float32, shape=[vocab_size, embed_dim])
if zero_pad:
lookup_table = tf.concat((tf.zeros([1, embed_dim]), lookup_table[1:, :]), axis=0)
outputs = tf.nn.embedding_lookup(lookup_table, inputs)
if scale:
outputs = outputs * np.sqrt(embed_dim)
return outputs
def multihead_attn(queries, keys, q_masks, k_masks, future_binding, is_training):
"""
Args:
queries: A 3d tensor with shape of [N, T_q, C_q]
keys: A 3d tensor with shape of [N, T_k, C_k]
"""
num_units = args.hidden_units
num_heads = args.num_heads
dropout_rate = args.dropout_rate
T_q = queries.get_shape().as_list()[1] # max time length of query
T_k = keys.get_shape().as_list()[1] # max time length of key
Q = tf.layers.dense(queries, num_units, name='Q') # (N, T_q, C)
K_V = tf.layers.dense(keys, 2*num_units, name='K_V')
K, V = tf.split(K_V, 2, -1)
Q_ = tf.concat(tf.split(Q, num_heads, axis=2), axis=0) # (h*N, T_q, C/h)
K_ = tf.concat(tf.split(K, num_heads, axis=2), axis=0) # (h*N, T_k, C/h)
V_ = tf.concat(tf.split(V, num_heads, axis=2), axis=0) # (h*N, T_k, C/h)
# Scaled Dot-Product
align = tf.matmul(Q_, tf.transpose(K_, [0,2,1])) # (h*N, T_q, T_k)
align = align / np.sqrt(K_.get_shape().as_list()[-1]) # scale
# Key Masking
paddings = tf.fill(tf.shape(align), float('-inf')) # exp(-large) -> 0
key_masks = k_masks # (N, T_k)
key_masks = tf.tile(key_masks, [num_heads, 1]) # (h*N, T_k)
key_masks = tf.tile(tf.expand_dims(key_masks, 1), [1, T_q, 1]) # (h*N, T_q, T_k)
align = tf.where(tf.equal(key_masks, 0), paddings, align) # (h*N, T_q, T_k)
if future_binding:
lower_tri = tf.ones([T_q, T_k]) # (T_q, T_k)
lower_tri = tf.linalg.LinearOperatorLowerTriangular(lower_tri).to_dense() # (T_q, T_k)
masks = tf.tile(tf.expand_dims(lower_tri,0), [tf.shape(align)[0], 1, 1]) # (h*N, T_q, T_k)
align = tf.where(tf.equal(masks, 0), paddings, align) # (h*N, T_q, T_k)
# Softmax
align = tf.nn.softmax(align) # (h*N, T_q, T_k)
# Query Masking
query_masks = tf.to_float(q_masks) # (N, T_q)
query_masks = tf.tile(query_masks, [num_heads, 1]) # (h*N, T_q)
query_masks = tf.tile(tf.expand_dims(query_masks, -1), [1, 1, T_k]) # (h*N, T_q, T_k)
align *= query_masks # (h*N, T_q, T_k)
align = tf.layers.dropout(align, dropout_rate, training=is_training) # (h*N, T_q, T_k)
# Weighted sum
outputs = tf.matmul(align, V_) # (h*N, T_q, C/h)
# Restore shape
outputs = tf.concat(tf.split(outputs, num_heads, axis=0), axis=2) # (N, T_q, C)
# Residual connection
outputs += queries # (N, T_q, C)
# Normalize
outputs = layer_norm(outputs) # (N, T_q, C)
return outputs
def pointwise_feedforward(inputs, activation=None):
num_units = [4*args.hidden_units, args.hidden_units]
# Inner layer
outputs = tf.layers.conv1d(inputs, num_units[0], kernel_size=1, activation=activation)
# Readout layer
outputs = tf.layers.conv1d(outputs, num_units[1], kernel_size=1, activation=None)
# Residual connection
outputs += inputs
# Normalize
outputs = layer_norm(outputs)
return outputs
def learned_position_encoding(inputs, mask, embed_dim):
T = inputs.get_shape().as_list()[1]
outputs = tf.range(tf.shape(inputs)[1]) # (T_q)
outputs = tf.expand_dims(outputs, 0) # (1, T_q)
outputs = tf.tile(outputs, [tf.shape(inputs)[0], 1]) # (N, T_q)
outputs = embed_seq(outputs, T, embed_dim, zero_pad=False, scale=False)
return tf.expand_dims(tf.to_float(mask), -1) * outputs
def sinusoidal_position_encoding(inputs, mask, repr_dim):
T = inputs.get_shape()[1].value
pos = tf.reshape(tf.range(0.0, tf.to_float(T), dtype=tf.float32), [-1, 1])
i = np.arange(0, repr_dim, 2, np.float32)
denom = np.reshape(np.power(10000.0, i / repr_dim), [1, -1])
enc = tf.expand_dims(tf.concat([tf.sin(pos / denom), tf.cos(pos / denom)], 1), 0)
return tf.tile(enc, [tf.shape(inputs)[0], 1, 1]) * tf.expand_dims(tf.to_float(mask), -1)
def label_smoothing(inputs, epsilon=0.1):
C = inputs.get_shape().as_list()[-1] # number of channels
return ((1 - epsilon) * inputs) + (epsilon / C)
# +
def forward_pass(sources, targets, mode, params):
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
pos_enc = _get_position_encoder()
# ENCODER
en_masks = tf.sign(sources)
with tf.variable_scope('encoder_embedding'):
encoded = embed_seq(inputs = sources,
vocab_size = params['source_vocab_size'],
embed_dim = args.hidden_units,
zero_pad = True,
scale = True)
with tf.variable_scope('encoder_position_encoding'):
encoded += pos_enc(sources, en_masks, args.hidden_units)
with tf.variable_scope('encoder_dropout'):
encoded = tf.layers.dropout(encoded, args.dropout_rate, training=is_training)
for i in range(args.num_blocks):
with tf.variable_scope('encoder_attn_%d'%i):
encoded = multihead_attn(queries = encoded,
keys = encoded,
q_masks = en_masks,
k_masks = en_masks,
future_binding = False,
is_training = is_training,)
with tf.variable_scope('encoder_feedforward_%d'%i):
encoded = pointwise_feedforward(encoded,
activation = params['activation'])
# DECODER
decoder_inputs = _shift_right(targets, params['start_symbol'])
de_masks = tf.sign(decoder_inputs)
if args.tied_embedding:
vs = tf.variable_scope('encoder_embedding', reuse=True)
else:
vs = tf.variable_scope('decoder_embedding')
with vs:
decoded = embed_seq(decoder_inputs,
params['target_vocab_size'],
args.hidden_units,
zero_pad = True,
scale = True)
with tf.variable_scope('decoder_position_encoding'):
decoded += pos_enc(decoder_inputs, de_masks, args.hidden_units)
with tf.variable_scope('decoder_dropout'):
decoded = tf.layers.dropout(decoded, args.dropout_rate, training=is_training)
for i in range(args.num_blocks):
with tf.variable_scope('decoder_self_attn_%d'%i):
decoded = multihead_attn(queries = decoded,
keys = decoded,
q_masks = de_masks,
k_masks = de_masks,
future_binding = True,
is_training = is_training)
with tf.variable_scope('decoder_attn_%d'%i):
decoded = multihead_attn(queries=decoded,
keys = encoded,
q_masks = de_masks,
k_masks = en_masks,
future_binding = False,
is_training = is_training)
with tf.variable_scope('decoder_feedforward_%d'%i):
decoded = pointwise_feedforward(decoded,
activation = params['activation'])
# OUTPUT LAYER
if args.tied_proj_weight:
b = tf.get_variable('bias', [params['target_vocab_size']], tf.float32)
_scope = 'encoder_embedding' if args.tied_embedding else 'decoder_embedding'
with tf.variable_scope(_scope, reuse=True):
shared_w = tf.get_variable('lookup_table')
decoded = tf.reshape(decoded, [-1, args.hidden_units])
logits = tf.nn.xw_plus_b(decoded, tf.transpose(shared_w), b)
logits = tf.reshape(logits, [tf.shape(sources)[0], -1, params['target_vocab_size']])
else:
with tf.variable_scope('output_layer'):
logits = tf.layers.dense(decoded, params['target_vocab_size'], reuse=reuse)
return logits
def _model_fn_train(features, mode, params):
logits = forward_pass(features['source'], features['target'], mode, params)
targets = features['target']
masks = tf.to_float(tf.not_equal(targets, 0))
if args.label_smoothing:
loss_op = label_smoothing_sequence_loss(logits = logits,
targets = targets,
weights = masks,
label_depth = params['target_vocab_size'])
else:
loss_op = tf.contrib.seq2seq.sequence_loss(logits = logits,
targets = targets,
weights = masks)
if args.lr_decay_strategy == 'noam':
step_num = tf.train.get_global_step() + 1 # prevents zero global step
lr = _get_noam_lr(step_num)
elif args.lr_decay_strategy == 'exp':
lr = tf.train.exponential_decay(1e-3, tf.train.get_global_step(), 100000, 0.1)
else:
raise ValueError("lr decay strategy must be one of 'noam' and 'exp'")
log_hook = tf.train.LoggingTensorHook({'lr': lr}, every_n_iter=100)
train_op = tf.train.AdamOptimizer(lr).minimize(loss_op,
global_step = tf.train.get_global_step())
return tf.estimator.EstimatorSpec(mode = mode,
loss = loss_op,
train_op = train_op,
training_hooks = [log_hook])
def _model_fn_predict(features, mode, params):
def cond(i, x, temp):
return i < args.target_max_len
def body(i, x, temp):
logits = forward_pass(features['source'], x, mode, params)
ids = tf.argmax(logits, -1)[:, i]
ids = tf.expand_dims(ids, -1)
temp = tf.concat([temp[:, 1:], ids], -1)
x = tf.concat([temp[:, -(i+1):], temp[:, :-(i+1)]], -1)
x = tf.reshape(x, [tf.shape(temp)[0], args.target_max_len])
i += 1
return i, x, temp
_, res, _ = tf.while_loop(cond, body, [tf.constant(0), features['target'], features['target']])
return tf.estimator.EstimatorSpec(mode=mode, predictions=res)
def tf_estimator_model_fn(features, labels, mode, params):
if mode == tf.estimator.ModeKeys.TRAIN:
return _model_fn_train(features, mode, params)
if mode == tf.estimator.ModeKeys.PREDICT:
return _model_fn_predict(features, mode, params)
def _shift_right(targets, start_symbol):
start_symbols = tf.cast(tf.fill([tf.shape(targets)[0], 1], start_symbol), tf.int64)
return tf.concat([start_symbols, targets[:, :-1]], axis=-1)
def _get_position_encoder():
if args.position_encoding == 'non_param':
pos_enc = sinusoidal_position_encoding
elif args.position_encoding == 'param':
pos_enc = learned_position_encoding
else:
raise ValueError("position encoding has to be either 'param' or 'non_param'")
return pos_enc
def _get_noam_lr(step_num):
return tf.rsqrt(tf.to_float(args.hidden_units)) * tf.minimum(
tf.rsqrt(tf.to_float(step_num)),
tf.to_float(step_num) * tf.convert_to_tensor(args.warmup_steps ** (-1.5)))
# +
def greedy_decode(test_words, tf_estimator, dl):
test_indices = []
for test_word in test_words:
test_idx = [dl.source_word2idx[c] for c in test_word] + \
[dl.source_word2idx['<pad>']] * (args.source_max_len - len(test_word))
test_indices.append(test_idx)
test_indices = np.atleast_2d(test_indices)
zeros = np.zeros([len(test_words), args.target_max_len], np.int64)
pred_ids = tf_estimator.predict(tf.estimator.inputs.numpy_input_fn(
x={'source':test_indices, 'target':zeros}, batch_size=len(test_words), shuffle=False))
pred_ids = list(pred_ids)
target_idx2word = {i: w for w, i in dl.target_word2idx.items()}
for i, test_word in enumerate(test_words):
ans = ''.join([target_idx2word[id] for id in pred_ids[i]])
print(test_word, '->', ans.replace('<end>', ''))
def prepare_params(dl):
if args.activation == 'relu':
activation = tf.nn.relu
elif args.activation == 'elu':
activation = tf.nn.elu
elif args.activation == 'lrelu':
activation = tf.nn.leaky_relu
else:
raise ValueError("acitivation fn has to be 'relu' or 'elu' or 'lrelu'")
params = {
'source_vocab_size': len(dl.source_word2idx),
'target_vocab_size': len(dl.target_word2idx),
'start_symbol': dl.target_word2idx['<start>'],
'activation': activation}
return params
def main():
print(json.dumps(args, indent=4))
dl = DataLoader(
source_path='../temp/letters_source.txt',
target_path='../temp/letters_target.txt')
sources, targets = dl.load()
tf_estimator = tf.estimator.Estimator(
tf_estimator_model_fn, params=prepare_params(dl))
for epoch in range(5):
tf_estimator.train(tf.estimator.inputs.numpy_input_fn(
x = {'source':sources, 'target':targets},
batch_size = args.batch_size,
num_epochs = None,
shuffle = True), steps=1000)
greedy_decode(['apple', 'common', 'zhedong'], tf_estimator, dl)
if __name__ == '__main__':
main()
# -
| src_nlp/tensorflow/attn_is_all_u_need/train_letters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Load original model
import tensorflow as tf
import pathlib
import os
import numpy as np
from matplotlib.pyplot import imshow
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score
root_dir = '../train_base_model'
model_dir = 'trained_resnet_vector-unquantized/save_model'
saved_model_dir = os.path.join(root_dir, model_dir)
trained_model = tf.saved_model.load(saved_model_dir)
list(trained_model.signatures.keys())
# ## Converting full model to TFLite hybrid quantization model
# Create a converter object from a SavedModel directory.
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_dir)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
tflite_model = converter.convert()
# Save TFLite model
# +
root_dir = '' # Leave it empty, we want to stay in the current directory as we create a subdirectory with next line.
tflite_models_dir = 'tflite_hybrid_model'
to_save_tflite_model_dir = os.path.join(root_dir, tflite_models_dir)
saved_tflite_models_dir = pathlib.Path(to_save_tflite_model_dir) #convert string to pathlib object
saved_tflite_models_dir.mkdir(exist_ok=True, parents=True) # make directory
# -
# Create a pathlib object to save quantized model with path and file name.
tgt = pathlib.Path(to_save_tflite_model_dir, 'converted_model_hybrid.tflite')
# Write quantized model to the file.
tgt.write_bytes(tflite_model)
# We wrote a TFLite hybrid quantization model to `tflite_hybrid_model` folder. Mode name is `converted_model_hybrid.tflite`. It's size is 23 MB.
# ## Preparing test data from TFRecord
# Loading TFRecord test data
# +
root_dir = '../train_base_model/tf_datasets/flower_photos'
test_pattern = "{}/image_classification_builder-test.tfrecord*".format(root_dir)
test_all_files = tf.data.Dataset.list_files( tf.io.gfile.glob(test_pattern))
test_all_ds = tf.data.TFRecordDataset(test_all_files, num_parallel_reads=tf.data.experimental.AUTOTUNE)
# -
sample_size = 0
for raw_record in test_all_ds:
sample_size += 1
print('Sample size: ', sample_size)
# Helper function
# +
def decode_and_resize(serialized_example):
# resized image should be [224, 224, 3] and normalized to value range [0, 255]
# label is integer index of class.
parsed_features = tf.io.parse_single_example(
serialized_example,
features = {
'image/channels' : tf.io.FixedLenFeature([], tf.int64),
'image/class/label' : tf.io.FixedLenFeature([], tf.int64),
'image/class/text' : tf.io.FixedLenFeature([], tf.string),
'image/colorspace' : tf.io.FixedLenFeature([], tf.string),
'image/encoded' : tf.io.FixedLenFeature([], tf.string),
'image/filename' : tf.io.FixedLenFeature([], tf.string),
'image/format' : tf.io.FixedLenFeature([], tf.string),
'image/height' : tf.io.FixedLenFeature([], tf.int64),
'image/width' : tf.io.FixedLenFeature([], tf.int64)
})
image = tf.io.decode_jpeg(parsed_features['image/encoded'], channels=3)
label = tf.cast(parsed_features['image/class/label'], tf.int32)
label_txt = tf.cast(parsed_features['image/class/text'], tf.string)
label_one_hot = tf.one_hot(label, depth = 5)
resized_image = tf.image.resize(image, [224, 224], method='nearest')
return resized_image, label_one_hot
def normalize(image, label):
#Convert `image` from [0, 255] -> [0, 1.0] floats
image = tf.cast(image, tf.float32) / 255.
return image, label
# -
# Apply transformation and normalization to TFRecord:
decoded = test_all_ds.map(decode_and_resize)
normed = decoded.map(normalize)
# Convert TFRecord to numpy array for scoring:
np_img_holder = np.empty((0, 224, 224,3), float)
np_lbl_holder = np.empty((0, 5), int)
for img, lbl in normed:
r = img.numpy() # image value extracted
rx = np.expand_dims(r, axis=0) # expand by adding a dimension for batching images.
lx = np.expand_dims(lbl, axis=0) # expand by adding a dimension for batching labels.
np_img_holder = np.append(np_img_holder, rx, axis=0) # append each image to create a batch of images.
np_lbl_holder = np.append(np_lbl_holder, lx, axis=0) # append each one-hot label to create a batch of labels.
# Now test data is in numpy format with standardized dimension, pixel value between 0 and 1, and batched. So are labels.
# %matplotlib inline
plt.figure()
for i in range(len(np_img_holder)):
plt.subplot(10, 5, i+1)
plt.axis('off')
imshow(np.asarray(np_img_holder[i]))
# For single tet file scoring, we need to add a dimension to one sample. Since a given image is of shape (224, 224, 3), we need to make it into (1, 224, 224, 3) so it will be accepted by the model for scoring, as the model is built to handle batch scoring, it is expecting four dimensions with the first dimension being sample size. So let's use the first image in the batch as an example:
x = np.expand_dims(np_img_holder[0], axis=0)
x.shape
# Convert it to a tensor with type of float32:
xf = tf.dtypes.cast(x, tf.float32)
type(xf)
# ## Mapping prediction to class name
# I need to create a reverse lookup dictionary to map probability back to label. In other words, I want to find the index where maximum probability is positioned in the array. Map this position index to flower type. To create the lookup dictionary, I need to parse the TFRecord with feature descriptions to extract label indices and names.
# +
feature_description = {
'image/channels' : tf.io.FixedLenFeature([], tf.int64),
'image/class/label' : tf.io.FixedLenFeature([], tf.int64),
'image/class/text' : tf.io.FixedLenFeature([], tf.string),
'image/colorspace' : tf.io.FixedLenFeature([], tf.string),
'image/encoded' : tf.io.FixedLenFeature([], tf.string),
'image/filename' : tf.io.FixedLenFeature([], tf.string),
'image/format' : tf.io.FixedLenFeature([], tf.string),
'image/height' : tf.io.FixedLenFeature([], tf.int64),
'image/width' : tf.io.FixedLenFeature([], tf.int64)
}
def _parse_function(example_proto):
return tf.io.parse_single_example(example_proto, feature_description)
parsd_ds = test_all_ds.map(_parse_function)
val_label_map = {}
# getting label mapping
for image_features in parsd_ds.take(30):
label_idx = image_features['image/class/label'].numpy()
label_str = image_features['image/class/text'].numpy().decode()
if label_idx not in val_label_map:
val_label_map[label_idx] = label_str
# -
val_label_map
# ## Scoring using quantized model
# Earlier, we created a variable `tgt` to hold the file path object that points to the path where our reduced float16 TFLite is saved.
str(tgt)
##Load the TFLite model and allocate tensors.
## tgt is the pathlib object referring to file path and <TFLITE_MODEL_NAME>.tflite
interpreter = tf.lite.Interpreter(model_path=str(tgt))
interpreter.allocate_tensors()
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
input_details[0]['shape']
output_details[0]['shape']
input_details
output_details
# ### Scoring single image
# Test the model
input_data = np.array(np.expand_dims(np_img_holder[0], axis=0), dtype=np.float32)
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
# The function `get_tensor()` returns a copy of the tensor data.
# Use `tensor()` in order to get a pointer to the tensor.
output_data = interpreter.get_tensor(output_details[0]['index'])
print(output_data)
# The last position has the highest probability. This position is index 4. We need to use `lookup` function to map this index to our label map.
def lookup(np_entry, dictionary):
class_key = np.argmax(np_entry)
return dictionary.get(class_key)
# `lookup` functio returns the class name:
lookup(output_data, val_label_map)
# The model predicts this image is `tulips` class.
# ### Scoring image batch
# We need to get the actual labels and put these true labels in a list.
actual = []
for i in range(len(np_lbl_holder)):
class_key = np.argmax(np_lbl_holder[i])
actual.append(val_label_map.get(class_key))
# Then we create a function `batch_predict` for batch scoring.
def batch_predict(input_raw, input_tensor, output_tensor, dictionary):
input_data = np.array(np.expand_dims(input_raw, axis=0), dtype=np.float32)
interpreter.set_tensor(input_tensor[0]['index'], input_data)
interpreter.invoke()
interpreter_output = interpreter.get_tensor(output_tensor[0]['index'])
plain_text_label = lookup(interpreter_output, dictionary)
return plain_text_label
# We call `batch_predict` to score our test images and store predictions in a list `batch_quantized_prediction`.
batch_quantized_prediction = []
for i in range(sample_size):
plain_text_label = batch_predict(np_img_holder[i], input_details, output_details, val_label_map)
batch_quantized_prediction.append(plain_text_label)
# Then we use `sklearn`'s `accuracy_score` to compare the prediction with the actual label list.
quantized_accuracy = accuracy_score(actual, batch_quantized_prediction)
print(quantized_accuracy)
# The accuracy of our hybrid quantization model is 0.8.
| Chapter07/train_hybrid_quantized_model/Hybrid_Quantization.ipynb |