code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# `openacc_interop` is an example program to demonstrate OpenACC interoperability
# features. The application reads a grayscale image in pgm format, applies an
# edge detector (combination of gaussian blur and sharpening filter) and writes
# the output as grayscale image in pgm format. The filters are applied in
# frequencies space and the necessary DFTs are carried out with using either a
# FFT library with a FFTW compatible interface, e.g. Intel MKL, in case a CPU is
# used for the compute or cuFFT in case a NVIDIA GPU is used.
# !make
#
# ### Usage
# ```
# ./openacc_interop input.pgm output.pgm
# ```
# E.g. using the example image from the subfolder images:
# !./openacc_interop images/617019_NVIDIA_HQ_bldg.pgm images/output.pgm
|
Code_Examples/Chapter_09/python-notebook/Chapter 9.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import sasoptpy as so
df = pd.read_csv('../data/input.csv')
df.head()
gk_data = df[df['Pos'] == 'G'].copy().reset_index()
gk_data.head()
model = so.Model(name='gk_model')
gk_data.set_index('index', inplace=True)
gk_data.head()
players = gk_data.index.tolist()
players
lineup = model.add_variables(players, name='lineup', vartype=so.binary)
lineup[7]
bench = model.add_variables(players, name='bench', vartype=so.binary)
total_xp = so.expr_sum(lineup[p] * gk_data.loc[p,'27_Pts'] for p in players) + 0.1 * so.expr_sum(bench[p] * gk_data.loc[p, '27_Pts'] for p in players)
total_xp
model.set_objective(total_xp, name='total_xp_obj', sense=so.maximize)
model.add_constraints((lineup[p] + bench[p] <= 1 for p in players), name='lineup_or_bench');
model.add_constraint(so.expr_sum(lineup[p] for p in players) == 1, name='single_lineup');
model.add_constraint(so.expr_sum(bench[p] for p in players) == 1, name='single_bench')
budget = 9.2
model.add_constraint(so.expr_sum((lineup[p] + bench[p]) * gk_data.loc[p, 'BV'] for p in players) <= budget, name='budget_con')
print(model.export_mps(fetch=True))
model.set_objective(-total_xp, name='total_xp_obj', sense='N')
model.export_mps(filename='gk.mps')
command = 'cbc gk.mps solve solu solution.txt'
# !{command}
with open('solution.txt', 'r') as f:
for v in model.get_variables():
v.set_value(0)
for line in f:
if 'objective value' in line:
continue
words = line.split()
var = model.get_variable(words[1])
var.set_value(float(words[2]))
print("LINEUP")
for p in players:
if lineup[p].get_value() > 0.5:
print(p, gk_data.loc[p])
print("BENCH")
for p in players:
if bench[p].get_value() > 0.5:
print(p, gk_data.loc[p])
|
notebooks/Tutorial 1 - Goalkeeper Problem.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:aparent]
# language: python
# name: conda-env-aparent-py
# ---
# +
import keras
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda
from keras.layers import Conv2D, MaxPooling2D, Conv1D, MaxPooling1D, LSTM, ConvLSTM2D, GRU, BatchNormalization, LocallyConnected2D, Permute
from keras.layers import Concatenate, Reshape, Softmax, Conv2DTranspose, Embedding, Multiply
from keras.callbacks import ModelCheckpoint, EarlyStopping, Callback
from keras import regularizers
from keras import backend as K
import keras.losses
import tensorflow as tf
from tensorflow.python.framework import ops
import isolearn.keras as iso
import numpy as np
import tensorflow as tf
import logging
logging.getLogger('tensorflow').setLevel(logging.ERROR)
import pandas as pd
import os
import pickle
import numpy as np
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
import isolearn.io as isoio
import isolearn.keras as isol
from genesis.visualization import *
from genesis.generator import *
from genesis.predictor import *
from genesis.optimizer import *
from definitions.generator.splirent_deconv_conv_generator_concat_alllayers import load_generator_network, get_shallow_copy_function
from definitions.predictor.splirent_only_random_regions_cuts import load_saved_predictor
import warnings
#Disable warnings
tf.logging.set_verbosity(tf.logging.ERROR)
from matplotlib.animation import FuncAnimation
from matplotlib import colors
import matplotlib.cm as cm
import matplotlib.colors as colors
import matplotlib as mpl
from matplotlib.text import TextPath
from matplotlib.patches import PathPatch, Rectangle
from matplotlib.font_manager import FontProperties
from matplotlib import gridspec
from matplotlib.ticker import FormatStrFormatter
#fp = FontProperties(family="Arial", weight="bold")
fp = FontProperties(family="Ubuntu", weight="bold")
globscale = 1.35
LETTERS = { "T" : TextPath((-0.305, 0), "T", size=1, prop=fp),
"G" : TextPath((-0.384, 0), "G", size=1, prop=fp),
"A" : TextPath((-0.35, 0), "A", size=1, prop=fp),
"C" : TextPath((-0.366, 0), "C", size=1, prop=fp),
"UP" : TextPath((-0.488, 0), '$\\Uparrow$', size=1, prop=fp),
"DN" : TextPath((-0.488, 0), '$\\Downarrow$', size=1, prop=fp),
"(" : TextPath((-0.25, 0), "(", size=1, prop=fp),
"." : TextPath((-0.125, 0), "-", size=1, prop=fp),
")" : TextPath((-0.1, 0), ")", size=1, prop=fp)}
COLOR_SCHEME = {'G': 'orange',
'A': 'red',
'C': 'blue',
'T': 'darkgreen',
'UP': 'green',
'DN': 'red',
'(': 'black',
'.': 'black',
')': 'black'}
def letterAt(letter, x, y, yscale=1, ax=None, color=None, alpha=1.0) :
text = LETTERS[letter]
chosen_color = COLOR_SCHEME[letter]
if color is not None :
chosen_color = color
t = mpl.transforms.Affine2D().scale(1*globscale, yscale*globscale) + \
mpl.transforms.Affine2D().translate(x,y) + ax.transData
p = PathPatch(text, lw=0, fc=chosen_color, alpha=alpha, transform=t)
if ax != None:
ax.add_artist(p)
return p
def plot_seqprop_logo(ax_logo, pwms, cut_preds, sequence_templates=None, logo_height=1.0, plot_start=0, plot_end=164) :
#Slice according to seq trim index
pwms = pwms[:, plot_start: plot_end, :]
cut_preds = cut_preds[:, plot_start: plot_end]
sequence_templates = [sequence_template[plot_start: plot_end] for sequence_template in sequence_templates]
pwms += 0.0001
for j in range(0, pwms.shape[1]) :
pwms[:, j, :] /= np.sum(pwms[:, j, :], axis=1).reshape(-1, 1)
entropies = np.zeros(pwms.shape)
entropies[pwms > 0] = pwms[pwms > 0] * -np.log2(pwms[pwms > 0])
entropies = np.sum(entropies, axis=2)
conservations = 2 - entropies
for k in range(pwms.shape[0]) :
pwm = pwms[k, :, :]
cut_pred = cut_preds[k, :]
sequence_template = sequence_templates[k]
conservation = conservations[k]
height_base = (1.0 - logo_height) / 2. + 4 * k * 2 + 2.5
for j in range(0, pwm.shape[0]) :
sort_index = np.argsort(pwm[j, :])
for ii in range(0, 4) :
i = sort_index[ii]
nt_prob = pwm[j, i] * conservation[j]
nt = ''
if i == 0 :
nt = 'A'
elif i == 1 :
nt = 'C'
elif i == 2 :
nt = 'G'
elif i == 3 :
nt = 'T'
color = None
if sequence_template[j] != 'N' :
color = 'black'
if ii == 0 :
letterAt(nt, j + 0.5, height_base, nt_prob * logo_height, ax_logo, color=color)
else :
prev_prob = np.sum(pwm[j, sort_index[:ii]] * conservation[j]) * logo_height
letterAt(nt, j + 0.5, height_base + prev_prob, nt_prob * logo_height, ax_logo, color=color)
plt.sca(ax_logo)
plt.xlim((-3, plot_end - plot_start))
plt.ylim((-0.1, 5 * 8))
plt.xticks([], [])
plt.yticks([], [])
plt.axis('off')
#ax_logo.axhline(y=0.01 + height_base, xmin=0, color='black', linestyle='-', linewidth=2)
ax_logo.plot([0, plot_end - plot_start], [0.01 + height_base, 0.01 + height_base], color='black', linestyle='-', linewidth=2)
l2, = ax_logo.plot(np.arange(plot_end - plot_start), 4 * k * 2 + 4.5 + cut_pred * 2., linewidth=2, linestyle='-', color='red', alpha=0.8)
# +
class GenesisMonitor(Callback):
def __init__(self, predictor_model, loss_model, track_mode='batch', batch_freq_dict=None, measure_funcs=[], batch_size=36, input_tensor_funcs=None, pwm_start=70 - 50, pwm_end=76 + 50) :
self.predictor_model = predictor_model
self.loss_model = loss_model
self.track_mode = track_mode
self.batch_freq_dict = batch_freq_dict
self.measure_funcs = measure_funcs
self.batch_size = batch_size
self.pwm_start = pwm_start
self.pwm_end = pwm_end
self.input_tensor_funcs = input_tensor_funcs
self.batch_history = []
self.loss_history = []
self.measure_history = []
self.entropy_history = []
self.nt_swap_history = []
self.pwm_history = []
self.input_history = []
self.prev_optimized_pwm = None
self.n_epochs = 0
self.n_batches = 0
self.batch_freq = 10
if self.batch_freq_dict is not None and 0 in self.batch_freq_dict :
self.batch_freq = self.batch_freq_dict[0]
input_tensors, optimized_loss, pred_bundle = self._predict_vals()
optimized_pwm = pred_bundle[3]
optimized_measures = [measure_func(pred_bundle[9:]) for measure_func in self.measure_funcs]
#Track metrics
self.batch_history.append(self.n_batches)
self._track_input_history(input_tensors)
self._track_pwm_history(optimized_pwm)
self._track_loss_history(optimized_loss)
self._track_measure_history(optimized_measures)
self._track_entropy_history(optimized_pwm)
self.prev_optimized_pwm = optimized_pwm
self.nt_swap_history.append(np.zeros((optimized_pwm.shape[0], 1)))
def _predict_vals(self) :
self.input_tensors = [self.input_tensor_funcs[i](i) for i in range(len(self.input_tensor_funcs))]
optimized_loss = self.loss_model.predict(x=self.input_tensors, batch_size=self.batch_size)
pred_bundle = self.predictor_model.predict(x=self.input_tensors, batch_size=self.batch_size)
return self.input_tensors, optimized_loss, pred_bundle
def _track_input_history(self, input_tensors) :
self.input_history.append(input_tensors)
def _track_pwm_history(self, optimized_pwm) :
self.pwm_history.append(optimized_pwm)
def _track_loss_history(self, optimized_loss) :
self.loss_history.append(optimized_loss)
def _track_measure_history(self, optimized_measures) :
self.measure_history.append(optimized_measures)
def _track_entropy_history(self, optimized_pwm) :
pwm_section = optimized_pwm[:, self.pwm_start:self.pwm_end, :, :]
entropy = pwm_section * -np.log(np.clip(pwm_section, 10**(-6), 1. - 10**(-6))) / np.log(2.0)
entropy = np.sum(entropy, axis=(2, 3))
conservation = 2.0 - entropy
mean_bits = np.expand_dims(np.mean(conservation, axis=-1), axis=-1)
self.entropy_history.append(mean_bits)
def _track_nt_swap_history(self, optimized_pwm) :
nt_swaps = np.zeros((optimized_pwm.shape[0], 1))
nt_swaps[:, 0] = self.nt_swap_history[-1][:, 0]
for i in range(optimized_pwm.shape[0]) :
for j in range(self.pwm_start, self.pwm_end) :
curr_max_nt = np.argmax(optimized_pwm[i, j, :, 0])
prev_max_nt = np.argmax(self.prev_optimized_pwm[i, j, :, 0])
if curr_max_nt != prev_max_nt :
nt_swaps[i, 0] += 1
self.nt_swap_history.append(nt_swaps)
def on_batch_end(self, batch, logs={}) :
self.n_batches += 1
if batch == 0 and self.batch_freq_dict is not None and self.n_epochs in self.batch_freq_dict :
self.batch_freq = self.batch_freq_dict[self.n_epochs]
if self.track_mode == 'batch' and batch % self.batch_freq == 0 :
input_tensors, optimized_loss, pred_bundle = self._predict_vals()
optimized_pwm = pred_bundle[3]
optimized_measures = [measure_func(pred_bundle[9:]) for measure_func in self.measure_funcs]
#Track metrics
self.batch_history.append(self.n_batches)
self._track_input_history(input_tensors)
self._track_pwm_history(optimized_pwm)
self._track_loss_history(optimized_loss)
self._track_measure_history(optimized_measures)
self._track_entropy_history(optimized_pwm)
self.prev_optimized_pwm = optimized_pwm
self.nt_swap_history.append(np.zeros((optimized_pwm.shape[0], 1)))
#Cache previous pwms
self.prev_optimized_pwm = optimized_pwm
def on_epoch_end(self, epoch, logs={}) :
self.n_epochs += 1
if self.track_mode == 'epoch' :
input_tensors, optimized_loss, pred_bundle = self._predict_vals()
optimized_pwm = pred_bundle[3]
optimized_measures = [measure_func(pred_bundle[9:]) for measure_func in self.measure_funcs]
#Track metrics
self._track_input_history(input_tensors)
self._track_pwm_history(optimized_pwm)
self._track_loss_history(optimized_loss)
self._track_measure_history(optimized_measures)
self._track_entropy_history(optimized_pwm)
self.prev_optimized_pwm = optimized_pwm
self.nt_swap_history.append(np.zeros((optimized_pwm.shape[0], 1)))
#Cache previous pwms
self.prev_optimized_pwm = optimized_pwm
# +
#Define target isoform loss function
def get_cleavage_loss(target_poses, region_1_start=10, region_1_end=35, region_1_target_bits=1.8, region_2_start=53, region_2_end=78, region_2_target_bits=1.8, entropy_weight=0.0, similarity_weight=0.0, similarity_margin=0.5) :
target_cuts = np.zeros((len(target_poses), 101))
for i, target_pos in enumerate(target_poses) :
target_cuts[i, target_pos] = 1.0
masked_entropy_mse_region_1 = get_target_entropy_sme_masked(pwm_start=region_1_start, pwm_end=region_1_end, target_bits=region_1_target_bits)
masked_entropy_mse_region_2 = get_target_entropy_sme_masked(pwm_start=region_2_start, pwm_end=region_2_end, target_bits=region_2_target_bits)
pwm_sample_entropy_func_region_1 = get_pwm_margin_sample_entropy_masked(pwm_start=region_1_start, pwm_end=region_1_end, margin=similarity_margin, shift_1_nt=True)
pwm_sample_entropy_func_region_2 = get_pwm_margin_sample_entropy_masked(pwm_start=region_2_start, pwm_end=region_2_end, margin=similarity_margin, shift_1_nt=True)
def loss_func(loss_tensors) :
_, _, _, sequence_class, pwm_logits_1, pwm_logits_2, pwm_1, pwm_2, sampled_pwm_1, sampled_pwm_2, mask, sampled_mask, hek_pred, hela_pred, mcf7_pred, cho_pred = loss_tensors
#Create target cuts with sample axis (ax=1)
cut_targets = K.constant(target_cuts)
cut_true = K.gather(cut_targets, sequence_class[:, 0])
cut_true = K.tile(K.expand_dims(cut_true, axis=1), (1, K.shape(sampled_pwm_1)[1], 1))
#Specify costs
cut_loss = 1.0 * K.mean(kl_divergence(cut_true, hek_pred), axis=1)
seq_loss = 0.0
entropy_loss = entropy_weight * (masked_entropy_mse_region_1(pwm_1, mask) + masked_entropy_mse_region_2(pwm_1, mask)) / 2.
entropy_loss += similarity_weight * (K.mean(pwm_sample_entropy_func_region_1(sampled_pwm_1, sampled_pwm_2, sampled_mask), axis=1) + K.mean(pwm_sample_entropy_func_region_2(sampled_pwm_1, sampled_pwm_2, sampled_mask), axis=1)) / 2.
#Compute total loss
total_loss = cut_loss + seq_loss + entropy_loss
return total_loss
def val_loss_func(loss_tensors) :
_, _, _, sequence_class, pwm_logits_1, pwm_logits_2, pwm_1, pwm_2, sampled_pwm_1, sampled_pwm_2, mask, sampled_mask, hek_pred, hela_pred, mcf7_pred, cho_pred = loss_tensors
#Create target cuts with sample axis (ax=1)
cut_targets = K.constant(target_cuts)
cut_true = K.gather(cut_targets, sequence_class[:, 0])
cut_true = K.tile(K.expand_dims(cut_true, axis=1), (1, K.shape(sampled_pwm_1)[1], 1))
#Specify costs
cut_loss = 1.0 * K.mean(kl_divergence(cut_true, hek_pred), axis=1)
seq_loss = 0.0
entropy_loss = entropy_weight * (masked_entropy_mse_region_1(pwm_1, mask) + masked_entropy_mse_region_2(pwm_1, mask)) / 2.
entropy_loss += similarity_weight * (K.mean(pwm_sample_entropy_func_region_1(sampled_pwm_1, sampled_pwm_2, sampled_mask), axis=1) + K.mean(pwm_sample_entropy_func_region_2(sampled_pwm_1, sampled_pwm_2, sampled_mask), axis=1)) / 2.
#Compute total loss
total_loss = cut_loss + seq_loss + entropy_loss
return total_loss
return loss_func, val_loss_func
class EpochVariableCallback(Callback):
def __init__(self, my_variable, my_func):
self.my_variable = my_variable
self.my_func = my_func
def on_epoch_end(self, epoch, logs={}):
K.set_value(self.my_variable, self.my_func(K.get_value(self.my_variable), epoch))
#Function for running GENESIS
def run_genesis(sequence_templates, loss_func, val_loss_func, val_fixed_tensor_funcs, val_random_tensor_funcs, library_contexts, model_path, batch_size=32, n_samples=1, n_epochs=10, steps_per_epoch=100, batch_freq_dict=None) :
#Build Generator Network
_, generator = build_generator(batch_size, len(sequence_templates[0]), load_generator_network, n_classes=len(sequence_templates), n_samples=n_samples, sequence_templates=sequence_templates, batch_normalize_pwm=False)
#Build Validation Generator Network
_, val_generator = get_generator_copier(generator)(batch_size, len(sequence_templates[0]), get_shallow_copy_function(generator), n_classes=len(sequence_templates), n_samples=n_samples, sequence_templates=sequence_templates, batch_normalize_pwm=False, validation_sample_mode='sample', supply_inputs=True)
#Build Predictor Network and hook it on the generator PWM output tensor
_, pwm_predictor = build_predictor(generator, load_saved_predictor(model_path, library_contexts=library_contexts), batch_size, n_samples=1, eval_mode='pwm')
_, sample_predictor = build_predictor(generator, load_saved_predictor(model_path, library_contexts=library_contexts), batch_size, n_samples=n_samples, eval_mode='sample')
for layer in pwm_predictor.layers :
if 'splirent' in layer.name :
layer.name += "_pwmversion"
_, val_predictor = build_predictor(val_generator, load_saved_predictor(model_path, library_contexts=library_contexts), batch_size, n_samples=n_samples, eval_mode='sample')
for layer in val_predictor.layers :
if 'splirent' in layer.name :
layer.name += "_valversion"
#Build Loss Model (In: Generator seed, Out: Loss function)
_, pwm_loss_model = build_loss_model(pwm_predictor, loss_func)
_, sample_loss_model = build_loss_model(sample_predictor, loss_func)
dual_loss_out = Lambda(lambda x: 0.5 * x[0] + 0.5 * x[1])([pwm_loss_model.outputs[0], sample_loss_model.outputs[0]])
loss_model = Model(inputs=pwm_loss_model.inputs, outputs=dual_loss_out)
_, val_loss_model = build_loss_model(val_predictor, val_loss_func)
#Specify Optimizer to use
#opt = keras.optimizers.SGD(lr=0.1)
opt = keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999)
#Compile Loss Model (Minimize self)
loss_model.compile(loss=lambda true, pred: pred, optimizer=opt)
get_hek_pred_func = lambda pred_outs: pred_outs[0]
fixed_genesis_monitor = GenesisMonitor(val_predictor, val_loss_model, track_mode='batch', batch_freq_dict=batch_freq_dict, measure_funcs=[get_hek_pred_func], batch_size=30, input_tensor_funcs=val_fixed_tensor_funcs)
random_genesis_monitor = GenesisMonitor(val_predictor, val_loss_model, track_mode='batch', batch_freq_dict=batch_freq_dict, measure_funcs=[get_hek_pred_func], batch_size=30, input_tensor_funcs=val_random_tensor_funcs)
#Specify callback entities
callbacks =[
fixed_genesis_monitor,
random_genesis_monitor
]
#Fit Loss Model
train_history = loss_model.fit(
[], np.ones((1, 1)), #Dummy training example
epochs=n_epochs,
steps_per_epoch=steps_per_epoch,
callbacks=callbacks
)
return generator, sample_predictor, train_history, fixed_genesis_monitor, random_genesis_monitor
# +
#Specfiy file path to pre-trained predictor network
save_dir = os.path.join(os.getcwd(), '../../../splirent/saved_models')
saved_predictor_model_name = 'aparent_splirent_only_random_regions_cuts_drop_02_sgd.h5'
saved_predictor_model_path = os.path.join(save_dir, saved_predictor_model_name)
# +
#Maximize isoform proportions for all native minigene libraries
sequence_templates = [
'AGGTGCTTGGNNNNNNNNNNNNNNNNNNNNNNNNNGGTCGACCCAGGTTCGTGNNNNNNNNNNNNNNNNNNNNNNNNNGAGGTATTCTTATCACCTTCGTGGCTACAGA',
'AGGTGCTTGGNNNNNNNNNNNNNNNNNNNNNNNNNGGTCGACCCAGGTTCGTGNNNNNNNNNNNNNNNNNNNNNNNNNGAGGTATTCTTATCACCTTCGTGGCTACAGA',
'AGGTGCTTGGNNNNNNNNNNNNNNNNNNNNNNNNNGGTCGACCCAGGTTCGTGNNNNNNNNNNNNNNNNNNNNNNNNNGAGGTATTCTTATCACCTTCGTGGCTACAGA',
'AGGTGCTTGGNNNNNNNNNNNNNNNNNNNNNNNNNGGTCGACCCAGGTTCGTGNNNNNNNNNNNNNNNNNNNNNNNNNGAGGTATTCTTATCACCTTCGTGGCTACAGA',
'AGGTGCTTGGNNNNNNNNNNNNNNNNNNNNNNNNNGGTCGACCCAGGTTCGTGNNNNNNNNNNNNNNNNNNNNNNNNNGAGGTATTCTTATCACCTTCGTGGCTACAGA'
]
library_contexts = [
'n/a',
'n/a',
'n/a',
'n/a',
'n/a'
]
cut_positions = [
0,
22,
44,
65,
79
]
# +
class_list = ([0] * 30) + ([1] * 30) + ([2] * 30) + ([3] * 30) + ([4] * 30)
#Fixed validation tensors
val_fixed_tensors = [
np.array(class_list).reshape(-1, 1),
np.random.uniform(-1, 1, (30 * 5, 100)),
np.random.uniform(-1, 1, (30 * 5, 100))
]
val_fixed_tensor_funcs = [
lambda i, val_fixed_tensors=val_fixed_tensors: val_fixed_tensors[0],
lambda i, val_fixed_tensors=val_fixed_tensors: val_fixed_tensors[1],
lambda i, val_fixed_tensors=val_fixed_tensors: val_fixed_tensors[2]
]
#Randomized validation tensors
val_random_tensor_funcs = [
lambda i: np.array(class_list).reshape(-1, 1),
lambda i: np.random.uniform(-1, 1, (30 * 5, 100)),
lambda i: np.random.uniform(-1, 1, (30 * 5, 100))
]
# +
#Train Splicing Cut GENESIS Network
print("Training GENESIS (multi-class)")
#Number of PWMs to generate per objective
batch_size = 30
#Number of One-hot sequences to sample from the PWM at each grad step
n_samples = 10
#Number of epochs per objective to optimize
n_epochs = 50
#Number of steps (grad updates) per epoch
steps_per_epoch = 100
batch_freq_dict = {
0 : 1,
1 : 5,
5 : 10,
25 : 20
}
save_name_suffix = 'hek'
loss, val_loss = get_cleavage_loss(
cut_positions,
region_1_start=10,
region_1_end=35,
region_1_target_bits=2.0,
region_2_start=53,
region_2_end=78,
region_2_target_bits=2.0,
entropy_weight=3.5,
similarity_weight=7.5,
similarity_margin=0.5
)
genesis_generator, genesis_predictor, train_history, fixed_genesis_monitor, random_genesis_monitor = None, None, None, None, None
with warnings.catch_warnings():
warnings.simplefilter("ignore")
genesis_generator, genesis_predictor, train_history, fixed_genesis_monitor, random_genesis_monitor = run_genesis(sequence_templates, loss, val_loss, val_fixed_tensor_funcs, val_random_tensor_funcs, library_contexts, saved_predictor_model_path, batch_size, n_samples, n_epochs, steps_per_epoch, batch_freq_dict=batch_freq_dict)
genesis_generator.get_layer('lambda_rand_sequence_class').function = lambda inp: inp
genesis_generator.get_layer('lambda_rand_input_1').function = lambda inp: inp
genesis_generator.get_layer('lambda_rand_input_2').function = lambda inp: inp
genesis_predictor.get_layer('lambda_rand_sequence_class').function = lambda inp: inp
genesis_predictor.get_layer('lambda_rand_input_1').function = lambda inp: inp
genesis_predictor.get_layer('lambda_rand_input_2').function = lambda inp: inp
# Save model and weights
save_dir = 'saved_models'
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
model_name = 'genesis_splicing_cnn_target_cuts_pwm_and_multisample_' + save_name_suffix + '_only_random_regions_' + str(n_epochs) + '_epochs_harderentropy_track_train_try_2_generator.h5'
model_path = os.path.join(save_dir, model_name)
genesis_generator.save(model_path)
print('Saved trained model at %s ' % model_path)
model_name = 'genesis_splicing_cnn_target_cuts_pwm_and_multisample_' + save_name_suffix + '_only_random_regions_' + str(n_epochs) + '_epochs_harderentropy_track_train_try_2_predictor.h5'
model_path = os.path.join(save_dir, model_name)
genesis_predictor.save(model_path)
print('Saved trained model at %s ' % model_path)
# +
#Deflate monitors into flat dictionaries
fixed_dict = {}
fixed_dict['batch_history'] = fixed_genesis_monitor.batch_history
fixed_dict['pwm_history'] = fixed_genesis_monitor.pwm_history
fixed_dict['seed_history'] = fixed_genesis_monitor.input_history
fixed_dict['loss_history'] = fixed_genesis_monitor.loss_history
fixed_dict['entropy_history'] = fixed_genesis_monitor.entropy_history
fixed_dict['nt_swap_history'] = fixed_genesis_monitor.nt_swap_history
fixed_dict['measure_history'] = fixed_genesis_monitor.measure_history
random_dict = {}
random_dict['batch_history'] = random_genesis_monitor.batch_history
random_dict['pwm_history'] = random_genesis_monitor.pwm_history
random_dict['seed_history'] = random_genesis_monitor.input_history
random_dict['loss_history'] = random_genesis_monitor.loss_history
random_dict['entropy_history'] = random_genesis_monitor.entropy_history
random_dict['nt_swap_history'] = random_genesis_monitor.nt_swap_history
random_dict['measure_history'] = random_genesis_monitor.measure_history
# +
#Store traced training data
import pickle
fixed_genesis_monitor.predictor_model = None
fixed_genesis_monitor.loss_model = None
random_genesis_monitor.predictor_model = None
random_genesis_monitor.loss_model = None
pickle.dump({'fixed_seeds' : fixed_dict, 'random_seeds' : random_dict}, open('genesis_splicing_cleavage_multiclass_monitors_try_2.pickle', 'wb'))
# +
#Load tracked training data
genesis_monitors = pickle.load(open('genesis_splicing_cleavage_multiclass_monitors_try_2.pickle', 'rb'))
fixed_genesis_monitor = genesis_monitors['fixed_seeds']
random_genesis_monitor = genesis_monitors['random_seeds']
# +
#Fixed seed GIF making
pwm_history = fixed_genesis_monitor['pwm_history']
flat_pwm_history = []
for step_index in range(0, len(pwm_history)) :
pwms = pwm_history[step_index]
flat_pwms = np.zeros((pwms.shape[0], 109))
for i in range(pwms.shape[0]) :
for j in range(109) :
max_nt_ix = np.argmax(pwms[i, j, :, 0])
flat_pwms[i, j] = max_nt_ix + 1
flat_pwms = flat_pwms[:, 0:85]
flat_pwm_history.append(flat_pwms)
batch_history = fixed_genesis_monitor['batch_history']
seed_history = fixed_genesis_monitor['seed_history']
loss_history = fixed_genesis_monitor['loss_history']
entropy_history = fixed_genesis_monitor['entropy_history']
nt_swap_history = fixed_genesis_monitor['nt_swap_history']
measure_history = fixed_genesis_monitor['measure_history']
min_loss = np.min(np.array([np.min(loss_history[i]) for i in range(loss_history[0].shape[0])]))
max_loss = np.max(np.array([np.max(loss_history[i]) for i in range(loss_history[0].shape[0])]))
sel_pwm_history = [
temp_pwms[[10, 45, 75, 100, 135][::-1], :, :, 0]
for temp_pwms in pwm_history
]
sel_cut_history = [
np.concatenate([np.zeros((5, 2)), temp_meas[0][[10, 45, 75, 100, 135][::-1], 0, 0:85 - 2]], axis=1)
for temp_meas in measure_history
]
# +
#Animation 1 (Fixed Seed): Loss and Diversity plot
n_classes = len(cut_positions)
n_frames = len(batch_history) - 1
cmap = colors.ListedColormap(['red', 'blue', 'orange', 'darkgreen'])
bounds=[0, 1, 2, 3, 4, 5]
norm = colors.BoundaryNorm(bounds, cmap.N)
w = flat_pwm_history[0].shape[1]
f, ax = plt.subplots(2, 1, figsize=(4, 9), gridspec_kw={'height_ratios': [1, 2.5]})
loss_lines = []
for i in range(loss_history[0].shape[0]) :
line, = ax[0].plot([], [], linewidth=2)
loss_lines.append(line)
plt.sca(ax[0])
plt.xlabel("Weight Updates", fontsize=14)
plt.ylabel("Loss", fontsize=14)
plt.title("Splicing DEN Training:\nTracking 30 Fixed Sequences\nPer Target Splice Site", fontsize=14)
plt.xticks([0, batch_history[n_frames-1]], [0, batch_history[n_frames-1]], fontsize=14)
plt.yticks(fontsize=14)
plt.xlim(0, batch_history[n_frames-1])
plt.ylim(min_loss - 0.02 * min_loss * np.sign(min_loss), max_loss + 0.02 * max_loss * np.sign(max_loss))
image = ax[1].imshow(flat_pwm_history[0][::-1, :], aspect='equal', interpolation='nearest', origin='lower', cmap=cmap, norm=norm)
plt.sca(ax[1])
plt.xlabel("Nucleotide Position", fontsize=14)
plt.ylabel("Generated Sequences", fontsize=14)
plt.xticks([25, 50, 75], ["-25", "CSE", "+25"], fontsize=14)
plt.yticks(np.arange(n_classes) * 30 + 15, ["SD1", "New", "SD2", "New", "SD3"][::-1], fontsize=14)
plt.xlim(0, w)
plt.ylim(0, 30 * n_classes)
plt.title("Weight Update 0\n1x Speedup >", fontsize=14)
plt.tight_layout()
for class_i in range(1, n_classes) :
ax[1].plot([0, w], [class_i * 30, class_i * 30], linewidth=3, color='black', linestyle='--')
#plt.show()
loss_data_x = [[0] for i in range(loss_history[0].shape[0])]
loss_data_y = [[loss_history[0][i]] for i in range(loss_history[0].shape[0])]
def init() :
for i in range(loss_history[0].shape[0]) :
loss_lines[i].set_data([], [])
image.set_data(flat_pwm_history[0][::-1, :])
return image,
def running_mean(x, N):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / float(N)
def animate(t) :
if t % 50 == 0 :
print("Grabbing frame " + str(t) + "...")
if t > 0 :
for i in range(loss_history[0].shape[0]) :
loss_data_x[i].append(batch_history[t])
loss_data_y[i].append(loss_history[t][i])
if t <= 100 :
loss_lines[i].set_data(loss_data_x[i], loss_data_y[i])
else :
loss_lines[i].set_data(loss_data_x[i], np.concatenate([loss_data_y[i][:99], running_mean(np.array(loss_data_y[i]), 100)]) )
image.set_data(flat_pwm_history[t][::-1, :])
curr_speed = 1
speed_sign = ">"
if t > 0 :
curr_speed = int(batch_history[t] - batch_history[t-1])
if curr_speed <= 1 :
speed_sign = ">"
elif curr_speed > 1 and curr_speed <= 5 :
speed_sign = ">>"
elif curr_speed > 5 :
speed_sign = ">>>"
ax[1].set_title("Weight Update " + str(batch_history[t]) + "\n" + str(curr_speed) + "x Speedup " + speed_sign, fontsize=14)
return image,
anim = FuncAnimation(f, animate, init_func=init, frames=n_frames+1, interval=20, blit=True)
anim.save('genesis_cleavage_multiclass_fixed_sequences.gif', writer='imagemagick')
# +
#Animation 2 (Fixed Seed): Seed, Loss and Diversity plot with PWMs
n_classes = len(cut_positions)
n_frames = len(batch_history) - 1
cmap = colors.ListedColormap(['red', 'blue', 'orange', 'darkgreen'])
bounds=[0, 1, 2, 3, 4, 5]
norm = colors.BoundaryNorm(bounds, cmap.N)
w = flat_pwm_history[0].shape[1]
f, ax = plt.subplots(2, 3, figsize=(14, 9), gridspec_kw={'width_ratios': [2, 4, 8], 'height_ratios': [1, 3]})
ax[0, 0].axis('off')
title_str = "Splicing DEN Training:\nTracking 30 Fixed Sequences\nPer Target Splice Site"
ax[0, 2].text(0.5, 0.5, title_str, {'color': 'black', 'fontsize': 20}, horizontalalignment='center', verticalalignment='center', transform=ax[0, 2].transAxes)
#Plot PWMs
ax[0, 2].axis('off')
ax[1, 2].axis('off')
ax[1, 2].get_xaxis().set_visible(False)
ax[1, 2].get_yaxis().set_visible(False)
plot_seqprop_logo(ax[1, 2], sel_pwm_history[500], sel_cut_history[0], sequence_templates=sequence_templates[::-1], logo_height=1.0, plot_start=0, plot_end=85)
loss_lines = []
for i in range(loss_history[0].shape[0]) :
line, = ax[0, 1].plot([], [], linewidth=2)
loss_lines.append(line)
plt.sca(ax[0, 1])
plt.ylabel("Loss", fontsize=14)
plt.xticks([0, batch_history[n_frames-1]], [0, batch_history[n_frames-1]], fontsize=14)
plt.yticks(fontsize=14)
plt.xlim(0, batch_history[n_frames-1])
plt.ylim(min_loss - 0.02 * min_loss * np.sign(min_loss), max_loss + 0.02 * max_loss * np.sign(max_loss))
seed_image = ax[1, 0].imshow(seed_history[0][1][::-1, :], aspect='auto', interpolation='nearest', origin='lower', cmap='Greys', vmin=-1.0, vmax=1.0)
plt.sca(ax[1, 0])
plt.xlabel("Seed Dimensions", fontsize=14)
plt.ylabel("Input Seeds", fontsize=14)
plt.xticks([0, 100], [0, 100], fontsize=14)
plt.yticks([], [], fontsize=14)
plt.xlim(0, 100)
plt.ylim(0, 30 * n_classes)
image = ax[1, 1].imshow(flat_pwm_history[0][::-1, :], aspect='equal', interpolation='nearest', origin='lower', cmap=cmap, norm=norm)
plt.sca(ax[1, 1])
plt.xlabel("Nucleotide Position", fontsize=14)
ax[1, 1].yaxis.set_label_position("right")
plt.ylabel("Generated Sequences", fontsize=14)
plt.xticks([25, 50, 75], ["-25", "CSE", "+25"], fontsize=14)
plt.yticks(np.arange(n_classes) * 30 + 15, ["SD1", "New", "SD2", "New", "SD3"][::-1], fontsize=14)
plt.xlim(0, w)
plt.ylim(0, 30 * n_classes)
plt.title("Weight Update 0\n1x Speedup >", fontsize=14)
ax[1, 2].annotate("", xytext=(-3, 4.5), xy=(0, 4.5), arrowprops=dict(arrowstyle="->"))
for class_i in range(1, n_classes) :
ax[1, 1].plot([0, w], [class_i * 30, class_i * 30], linewidth=3, color='black', linestyle='--')
ax[1, 2].annotate("", xytext=(-3, 4.5 + 4 * class_i * 2), xy=(0, 4.5 + 4 * class_i * 2), arrowprops=dict(arrowstyle="->"))
plt.tight_layout()
plt.subplots_adjust(wspace=0.15)
#plt.show()
loss_data_x = [[0] for i in range(loss_history[0].shape[0])]
loss_data_y = [[loss_history[0][i]] for i in range(loss_history[0].shape[0])]
def init() :
for i in range(loss_history[0].shape[0]) :
loss_lines[i].set_data([], [])
image.set_data(flat_pwm_history[0][::-1, :])
return image,
def running_mean(x, N):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / float(N)
def animate(t) :
if t % 10 == 0 :
print("Grabbing frame " + str(t) + "...")
if t > 0 :
for i in range(loss_history[0].shape[0]) :
loss_data_x[i].append(batch_history[t])
loss_data_y[i].append(loss_history[t][i])
if t <= 100 :
loss_lines[i].set_data(loss_data_x[i], loss_data_y[i])
else :
loss_lines[i].set_data(loss_data_x[i], np.concatenate([loss_data_y[i][:99], running_mean(np.array(loss_data_y[i]), 100)]) )
seed_image.set_data(seed_history[t][1][::-1, :])
image.set_data(flat_pwm_history[t][::-1, :])
curr_speed = 1
speed_sign = ">"
if t > 0 :
curr_speed = int(batch_history[t] - batch_history[t-1])
if curr_speed <= 1 :
speed_sign = ">"
elif curr_speed > 1 and curr_speed <= 5 :
speed_sign = ">>"
elif curr_speed > 5 :
speed_sign = ">>>"
ax[1, 1].set_title("Weight Update " + str(batch_history[t]) + "\n" + str(curr_speed) + "x Speedup " + speed_sign, fontsize=14)
ax[1, 2].clear()
ax[1, 2].axis('off')
plot_seqprop_logo(ax[1, 2], sel_pwm_history[t], sel_cut_history[t], sequence_templates=sequence_templates[::-1], logo_height=1.0, plot_start=0, plot_end=85)
for class_i in range(0, n_classes) :
ax[1, 2].annotate("", xytext=(-3, 4.5 + 4 * class_i * 2), xy=(0, 4.5 + 4 * class_i * 2), arrowprops=dict(arrowstyle="->"))
return image,
anim = FuncAnimation(f, animate, init_func=init, frames=n_frames+1, interval=50, blit=True)
anim.save('genesis_cleavage_multiclass_fixed_sequences_with_seeds_and_pwms.gif', writer='imagemagick')
# +
#Animation 3 (Fixed Seed): Loss and Diversity plot with PWMs
n_classes = len(cut_positions)
n_frames = len(batch_history) - 1
cmap = colors.ListedColormap(['red', 'blue', 'orange', 'darkgreen'])
bounds=[0, 1, 2, 3, 4, 5]
norm = colors.BoundaryNorm(bounds, cmap.N)
w = flat_pwm_history[0].shape[1]
f, ax = plt.subplots(2, 2, figsize=(12, 9), gridspec_kw={'width_ratios': [4, 8], 'height_ratios': [1, 3]})
ax = [
[None, ax[0, 0], ax[0, 1]],
[None, ax[1, 0], ax[1, 1]]
]
ax = np.array(ax, dtype=np.object)
title_str = "Splicing DEN Training:\nTracking 30 Fixed Sequences\nPer Target Splice Site"
ax[0, 2].text(0.5, 0.5, title_str, {'color': 'black', 'fontsize': 20}, horizontalalignment='center', verticalalignment='center', transform=ax[0, 2].transAxes)
#Plot PWMs
ax[0, 2].axis('off')
ax[1, 2].axis('off')
ax[1, 2].get_xaxis().set_visible(False)
ax[1, 2].get_yaxis().set_visible(False)
plot_seqprop_logo(ax[1, 2], sel_pwm_history[500], sel_cut_history[0], sequence_templates=sequence_templates[::-1], logo_height=1.0, plot_start=0, plot_end=85)
loss_lines = []
for i in range(loss_history[0].shape[0]) :
line, = ax[0, 1].plot([], [], linewidth=2)
loss_lines.append(line)
plt.sca(ax[0, 1])
plt.ylabel("Loss", fontsize=14)
plt.xticks([0, batch_history[n_frames-1]], [0, batch_history[n_frames-1]], fontsize=14)
plt.yticks(fontsize=14)
plt.xlim(0, batch_history[n_frames-1])
plt.ylim(min_loss - 0.02 * min_loss * np.sign(min_loss), max_loss + 0.02 * max_loss * np.sign(max_loss))
image = ax[1, 1].imshow(flat_pwm_history[0][::-1, :], aspect='equal', interpolation='nearest', origin='lower', cmap=cmap, norm=norm)
plt.sca(ax[1, 1])
plt.xlabel("Nucleotide Position", fontsize=14)
ax[1, 1].yaxis.set_label_position("right")
plt.ylabel("Generated Sequences", fontsize=14)
plt.xticks([25, 50, 75], ["-25", "CSE", "+25"], fontsize=14)
plt.yticks(np.arange(n_classes) * 30 + 15, ["SD1", "New", "SD2", "New", "SD3"][::-1], fontsize=14)
plt.xlim(0, w)
plt.ylim(0, 30 * n_classes)
plt.title("Weight Update 0\n1x Speedup >", fontsize=14)
ax[1, 2].annotate("", xytext=(-3, 4.5), xy=(0, 4.5), arrowprops=dict(arrowstyle="->"))
for class_i in range(1, n_classes) :
ax[1, 1].plot([0, w], [class_i * 30, class_i * 30], linewidth=3, color='black', linestyle='--')
ax[1, 2].annotate("", xytext=(-3, 4.5 + 4 * class_i * 2), xy=(0, 4.5 + 4 * class_i * 2), arrowprops=dict(arrowstyle="->"))
plt.tight_layout()
plt.subplots_adjust(wspace=0.15)
#plt.show()
loss_data_x = [[0] for i in range(loss_history[0].shape[0])]
loss_data_y = [[loss_history[0][i]] for i in range(loss_history[0].shape[0])]
def init() :
for i in range(loss_history[0].shape[0]) :
loss_lines[i].set_data([], [])
image.set_data(flat_pwm_history[0][::-1, :])
return image,
def running_mean(x, N):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / float(N)
def animate(t) :
if t % 10 == 0 :
print("Grabbing frame " + str(t) + "...")
if t > 0 :
for i in range(loss_history[0].shape[0]) :
loss_data_x[i].append(batch_history[t])
loss_data_y[i].append(loss_history[t][i])
if t <= 100 :
loss_lines[i].set_data(loss_data_x[i], loss_data_y[i])
else :
loss_lines[i].set_data(loss_data_x[i], np.concatenate([loss_data_y[i][:99], running_mean(np.array(loss_data_y[i]), 100)]) )
seed_image.set_data(seed_history[t][1][::-1, :])
image.set_data(flat_pwm_history[t][::-1, :])
curr_speed = 1
speed_sign = ">"
if t > 0 :
curr_speed = int(batch_history[t] - batch_history[t-1])
if curr_speed <= 1 :
speed_sign = ">"
elif curr_speed > 1 and curr_speed <= 5 :
speed_sign = ">>"
elif curr_speed > 5 :
speed_sign = ">>>"
ax[1, 1].set_title("Weight Update " + str(batch_history[t]) + "\n" + str(curr_speed) + "x Speedup " + speed_sign, fontsize=14)
ax[1, 2].clear()
ax[1, 2].axis('off')
plot_seqprop_logo(ax[1, 2], sel_pwm_history[t], sel_cut_history[t], sequence_templates=sequence_templates[::-1], logo_height=1.0, plot_start=0, plot_end=85)
for class_i in range(0, n_classes) :
ax[1, 2].annotate("", xytext=(-3, 4.5 + 4 * class_i * 2), xy=(0, 4.5 + 4 * class_i * 2), arrowprops=dict(arrowstyle="->"))
return image,
anim = FuncAnimation(f, animate, init_func=init, frames=n_frames+1, interval=50, blit=True)
anim.save('genesis_cleavage_multiclass_fixed_sequences_and_pwms.gif', writer='imagemagick')
# +
#Random seed GIF making
pwm_history = random_genesis_monitor['pwm_history']
flat_pwm_history = []
for step_index in range(0, len(pwm_history)) :
pwms = pwm_history[step_index]
flat_pwms = np.zeros((pwms.shape[0], 109))
for i in range(pwms.shape[0]) :
for j in range(109) :
max_nt_ix = np.argmax(pwms[i, j, :, 0])
flat_pwms[i, j] = max_nt_ix + 1
flat_pwms = flat_pwms[:, 0:85]
flat_pwm_history.append(flat_pwms)
batch_history = random_genesis_monitor['batch_history']
seed_history = random_genesis_monitor['seed_history']
loss_history = random_genesis_monitor['loss_history']
entropy_history = random_genesis_monitor['entropy_history']
nt_swap_history = random_genesis_monitor['nt_swap_history']
measure_history = random_genesis_monitor['measure_history']
min_loss = np.min(np.array([np.min(loss_history[i]) for i in range(loss_history[0].shape[0])]))
max_loss = np.max(np.array([np.max(loss_history[i]) for i in range(loss_history[0].shape[0])]))
sel_pwm_history = [
temp_pwms[[10, 45, 75, 100, 135][::-1], :, :, 0]
for temp_pwms in pwm_history
]
sel_cut_history = [
np.concatenate([np.zeros((5, 2)), temp_meas[0][[10, 45, 75, 100, 135][::-1], 0, 0:85 - 2]], axis=1)
for temp_meas in measure_history
]
# +
#Animation 1 (Random Seed): Loss and Diversity plot
n_classes = len(cut_positions)
n_frames = len(batch_history) - 1
cmap = colors.ListedColormap(['red', 'blue', 'orange', 'darkgreen'])
bounds=[0, 1, 2, 3, 4, 5]
norm = colors.BoundaryNorm(bounds, cmap.N)
w = flat_pwm_history[0].shape[1]
f, ax = plt.subplots(2, 1, figsize=(4, 9), gridspec_kw={'height_ratios': [1, 2.5]})
loss_lines = []
for i in range(loss_history[0].shape[0]) :
line, = ax[0].plot([], [], linewidth=2)
loss_lines.append(line)
plt.sca(ax[0])
plt.xlabel("Weight Updates", fontsize=14)
plt.ylabel("Loss", fontsize=14)
plt.title("Splicing DEN Training:\nRandomly Inspecting 30 Sequences\nPer Target Splice Site", fontsize=14)
plt.xticks([0, batch_history[n_frames-1]], [0, batch_history[n_frames-1]], fontsize=14)
plt.yticks(fontsize=14)
plt.xlim(0, batch_history[n_frames-1])
plt.ylim(min_loss - 0.02 * min_loss * np.sign(min_loss), max_loss + 0.02 * max_loss * np.sign(max_loss))
image = ax[1].imshow(flat_pwm_history[0][::-1, :], aspect='equal', interpolation='nearest', origin='lower', cmap=cmap, norm=norm)
plt.sca(ax[1])
plt.xlabel("Nucleotide Position", fontsize=14)
plt.ylabel("Generated Sequences", fontsize=14)
plt.xticks([25, 50, 75], ["-25", "CSE", "+25"], fontsize=14)
plt.yticks(np.arange(n_classes) * 30 + 15, ["SD1", "New", "SD2", "New", "SD3"][::-1], fontsize=14)
plt.xlim(0, w)
plt.ylim(0, 30 * n_classes)
plt.title("Weight Update 0\n1x Speedup >", fontsize=14)
plt.tight_layout()
for class_i in range(1, n_classes) :
ax[1].plot([0, w], [class_i * 30, class_i * 30], linewidth=3, color='black', linestyle='--')
#plt.show()
loss_data_x = [[0] for i in range(loss_history[0].shape[0])]
loss_data_y = [[loss_history[0][i]] for i in range(loss_history[0].shape[0])]
def init() :
for i in range(loss_history[0].shape[0]) :
loss_lines[i].set_data([], [])
image.set_data(flat_pwm_history[0][::-1, :])
return image,
def running_mean(x, N):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / float(N)
def animate(t) :
if t % 50 == 0 :
print("Grabbing frame " + str(t) + "...")
if t > 0 :
for i in range(loss_history[0].shape[0]) :
loss_data_x[i].append(batch_history[t])
loss_data_y[i].append(loss_history[t][i])
if t <= 100 :
loss_lines[i].set_data(loss_data_x[i], loss_data_y[i])
else :
loss_lines[i].set_data(loss_data_x[i], np.concatenate([loss_data_y[i][:99], running_mean(np.array(loss_data_y[i]), 100)]) )
image.set_data(flat_pwm_history[t][::-1, :])
curr_speed = 1
speed_sign = ">"
if t > 0 :
curr_speed = int(batch_history[t] - batch_history[t-1])
if curr_speed <= 1 :
speed_sign = ">"
elif curr_speed > 1 and curr_speed <= 5 :
speed_sign = ">>"
elif curr_speed > 5 :
speed_sign = ">>>"
ax[1].set_title("Weight Update " + str(batch_history[t]) + "\n" + str(curr_speed) + "x Speedup " + speed_sign, fontsize=14)
return image,
anim = FuncAnimation(f, animate, init_func=init, frames=n_frames+1, interval=20, blit=True)
anim.save('genesis_cleavage_multiclass_random_sequences.gif', writer='imagemagick')
# +
#Animation 2 (Random Seed): Seed, Loss and Diversity plot with PWMs
n_classes = len(cut_positions)
n_frames = len(batch_history) - 1
cmap = colors.ListedColormap(['red', 'blue', 'orange', 'darkgreen'])
bounds=[0, 1, 2, 3, 4, 5]
norm = colors.BoundaryNorm(bounds, cmap.N)
w = flat_pwm_history[0].shape[1]
f, ax = plt.subplots(2, 3, figsize=(14, 9), gridspec_kw={'width_ratios': [2, 4, 8], 'height_ratios': [1, 3]})
ax[0, 0].axis('off')
title_str = "Splicing DEN Training:\nRandomly Inspecting 30 Sequences\nPer Target Splice Site"
ax[0, 2].text(0.5, 0.5, title_str, {'color': 'black', 'fontsize': 20}, horizontalalignment='center', verticalalignment='center', transform=ax[0, 2].transAxes)
#Plot PWMs
ax[0, 2].axis('off')
ax[1, 2].axis('off')
ax[1, 2].get_xaxis().set_visible(False)
ax[1, 2].get_yaxis().set_visible(False)
plot_seqprop_logo(ax[1, 2], sel_pwm_history[500], sel_cut_history[0], sequence_templates=sequence_templates[::-1], logo_height=1.0, plot_start=0, plot_end=85)
loss_lines = []
for i in range(loss_history[0].shape[0]) :
line, = ax[0, 1].plot([], [], linewidth=2)
loss_lines.append(line)
plt.sca(ax[0, 1])
plt.ylabel("Loss", fontsize=14)
plt.xticks([0, batch_history[n_frames-1]], [0, batch_history[n_frames-1]], fontsize=14)
plt.yticks(fontsize=14)
plt.xlim(0, batch_history[n_frames-1])
plt.ylim(min_loss - 0.02 * min_loss * np.sign(min_loss), max_loss + 0.02 * max_loss * np.sign(max_loss))
seed_image = ax[1, 0].imshow(seed_history[0][1][::-1, :], aspect='auto', interpolation='nearest', origin='lower', cmap='Greys', vmin=-1.0, vmax=1.0)
plt.sca(ax[1, 0])
plt.xlabel("Seed Dimensions", fontsize=14)
plt.ylabel("Input Seeds", fontsize=14)
plt.xticks([0, 100], [0, 100], fontsize=14)
plt.yticks([], [], fontsize=14)
plt.xlim(0, 100)
plt.ylim(0, 30 * n_classes)
image = ax[1, 1].imshow(flat_pwm_history[0][::-1, :], aspect='equal', interpolation='nearest', origin='lower', cmap=cmap, norm=norm)
plt.sca(ax[1, 1])
plt.xlabel("Nucleotide Position", fontsize=14)
ax[1, 1].yaxis.set_label_position("right")
plt.ylabel("Generated Sequences", fontsize=14)
plt.xticks([25, 50, 75], ["-25", "CSE", "+25"], fontsize=14)
plt.yticks(np.arange(n_classes) * 30 + 15, ["SD1", "New", "SD2", "New", "SD3"][::-1], fontsize=14)
plt.xlim(0, w)
plt.ylim(0, 30 * n_classes)
plt.title("Weight Update 0\n1x Speedup >", fontsize=14)
ax[1, 2].annotate("", xytext=(-3, 4.5), xy=(0, 4.5), arrowprops=dict(arrowstyle="->"))
for class_i in range(1, n_classes) :
ax[1, 1].plot([0, w], [class_i * 30, class_i * 30], linewidth=3, color='black', linestyle='--')
ax[1, 2].annotate("", xytext=(-3, 4.5 + 4 * class_i * 2), xy=(0, 4.5 + 4 * class_i * 2), arrowprops=dict(arrowstyle="->"))
plt.tight_layout()
plt.subplots_adjust(wspace=0.15)
#plt.show()
loss_data_x = [[0] for i in range(loss_history[0].shape[0])]
loss_data_y = [[loss_history[0][i]] for i in range(loss_history[0].shape[0])]
def init() :
for i in range(loss_history[0].shape[0]) :
loss_lines[i].set_data([], [])
image.set_data(flat_pwm_history[0][::-1, :])
return image,
def running_mean(x, N):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / float(N)
def animate(t) :
if t % 10 == 0 :
print("Grabbing frame " + str(t) + "...")
if t > 0 :
for i in range(loss_history[0].shape[0]) :
loss_data_x[i].append(batch_history[t])
loss_data_y[i].append(loss_history[t][i])
if t <= 100 :
loss_lines[i].set_data(loss_data_x[i], loss_data_y[i])
else :
loss_lines[i].set_data(loss_data_x[i], np.concatenate([loss_data_y[i][:99], running_mean(np.array(loss_data_y[i]), 100)]) )
seed_image.set_data(seed_history[t][1][::-1, :])
image.set_data(flat_pwm_history[t][::-1, :])
curr_speed = 1
speed_sign = ">"
if t > 0 :
curr_speed = int(batch_history[t] - batch_history[t-1])
if curr_speed <= 1 :
speed_sign = ">"
elif curr_speed > 1 and curr_speed <= 5 :
speed_sign = ">>"
elif curr_speed > 5 :
speed_sign = ">>>"
ax[1, 1].set_title("Weight Update " + str(batch_history[t]) + "\n" + str(curr_speed) + "x Speedup " + speed_sign, fontsize=14)
ax[1, 2].clear()
ax[1, 2].axis('off')
plot_seqprop_logo(ax[1, 2], sel_pwm_history[t], sel_cut_history[t], sequence_templates=sequence_templates[::-1], logo_height=1.0, plot_start=0, plot_end=85)
for class_i in range(0, n_classes) :
ax[1, 2].annotate("", xytext=(-3, 4.5 + 4 * class_i * 2), xy=(0, 4.5 + 4 * class_i * 2), arrowprops=dict(arrowstyle="->"))
return image,
anim = FuncAnimation(f, animate, init_func=init, frames=n_frames+1, interval=50, blit=True)
anim.save('genesis_cleavage_multiclass_random_sequences_with_seeds_and_pwms.gif', writer='imagemagick')
# +
#Animation 3 (Random Seed): Loss and Diversity plot with PWMs
n_classes = len(cut_positions)
n_frames = len(batch_history) - 1
cmap = colors.ListedColormap(['red', 'blue', 'orange', 'darkgreen'])
bounds=[0, 1, 2, 3, 4, 5]
norm = colors.BoundaryNorm(bounds, cmap.N)
w = flat_pwm_history[0].shape[1]
f, ax = plt.subplots(2, 2, figsize=(12, 9), gridspec_kw={'width_ratios': [4, 8], 'height_ratios': [1, 3]})
ax = [
[None, ax[0, 0], ax[0, 1]],
[None, ax[1, 0], ax[1, 1]]
]
ax = np.array(ax, dtype=np.object)
title_str = "Splicing DEN Training:\nRandomly Inspecting 30 Sequences\nPer Target Splice Site"
ax[0, 2].text(0.5, 0.5, title_str, {'color': 'black', 'fontsize': 20}, horizontalalignment='center', verticalalignment='center', transform=ax[0, 2].transAxes)
#Plot PWMs
ax[0, 2].axis('off')
ax[1, 2].axis('off')
ax[1, 2].get_xaxis().set_visible(False)
ax[1, 2].get_yaxis().set_visible(False)
plot_seqprop_logo(ax[1, 2], sel_pwm_history[500], sel_cut_history[0], sequence_templates=sequence_templates[::-1], logo_height=1.0, plot_start=0, plot_end=85)
loss_lines = []
for i in range(loss_history[0].shape[0]) :
line, = ax[0, 1].plot([], [], linewidth=2)
loss_lines.append(line)
plt.sca(ax[0, 1])
plt.ylabel("Loss", fontsize=14)
plt.xticks([0, batch_history[n_frames-1]], [0, batch_history[n_frames-1]], fontsize=14)
plt.yticks(fontsize=14)
plt.xlim(0, batch_history[n_frames-1])
plt.ylim(min_loss - 0.02 * min_loss * np.sign(min_loss), max_loss + 0.02 * max_loss * np.sign(max_loss))
image = ax[1, 1].imshow(flat_pwm_history[0][::-1, :], aspect='equal', interpolation='nearest', origin='lower', cmap=cmap, norm=norm)
plt.sca(ax[1, 1])
plt.xlabel("Nucleotide Position", fontsize=14)
ax[1, 1].yaxis.set_label_position("right")
plt.ylabel("Generated Sequences", fontsize=14)
plt.xticks([25, 50, 75], ["-25", "CSE", "+25"], fontsize=14)
plt.yticks(np.arange(n_classes) * 30 + 15, ["SD1", "New", "SD2", "New", "SD3"][::-1], fontsize=14)
plt.xlim(0, w)
plt.ylim(0, 30 * n_classes)
plt.title("Weight Update 0\n1x Speedup >", fontsize=14)
ax[1, 2].annotate("", xytext=(-3, 4.5), xy=(0, 4.5), arrowprops=dict(arrowstyle="->"))
for class_i in range(1, n_classes) :
ax[1, 1].plot([0, w], [class_i * 30, class_i * 30], linewidth=3, color='black', linestyle='--')
ax[1, 2].annotate("", xytext=(-3, 4.5 + 4 * class_i * 2), xy=(0, 4.5 + 4 * class_i * 2), arrowprops=dict(arrowstyle="->"))
plt.tight_layout()
plt.subplots_adjust(wspace=0.15)
#plt.show()
loss_data_x = [[0] for i in range(loss_history[0].shape[0])]
loss_data_y = [[loss_history[0][i]] for i in range(loss_history[0].shape[0])]
def init() :
for i in range(loss_history[0].shape[0]) :
loss_lines[i].set_data([], [])
image.set_data(flat_pwm_history[0][::-1, :])
return image,
def running_mean(x, N):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / float(N)
def animate(t) :
if t % 10 == 0 :
print("Grabbing frame " + str(t) + "...")
if t > 0 :
for i in range(loss_history[0].shape[0]) :
loss_data_x[i].append(batch_history[t])
loss_data_y[i].append(loss_history[t][i])
if t <= 100 :
loss_lines[i].set_data(loss_data_x[i], loss_data_y[i])
else :
loss_lines[i].set_data(loss_data_x[i], np.concatenate([loss_data_y[i][:99], running_mean(np.array(loss_data_y[i]), 100)]) )
seed_image.set_data(seed_history[t][1][::-1, :])
image.set_data(flat_pwm_history[t][::-1, :])
curr_speed = 1
speed_sign = ">"
if t > 0 :
curr_speed = int(batch_history[t] - batch_history[t-1])
if curr_speed <= 1 :
speed_sign = ">"
elif curr_speed > 1 and curr_speed <= 5 :
speed_sign = ">>"
elif curr_speed > 5 :
speed_sign = ">>>"
ax[1, 1].set_title("Weight Update " + str(batch_history[t]) + "\n" + str(curr_speed) + "x Speedup " + speed_sign, fontsize=14)
ax[1, 2].clear()
ax[1, 2].axis('off')
plot_seqprop_logo(ax[1, 2], sel_pwm_history[t], sel_cut_history[t], sequence_templates=sequence_templates[::-1], logo_height=1.0, plot_start=0, plot_end=85)
for class_i in range(0, n_classes) :
ax[1, 2].annotate("", xytext=(-3, 4.5 + 4 * class_i * 2), xy=(0, 4.5 + 4 * class_i * 2), arrowprops=dict(arrowstyle="->"))
return image,
anim = FuncAnimation(f, animate, init_func=init, frames=n_frames+1, interval=50, blit=True)
anim.save('genesis_cleavage_multiclass_random_sequences_and_pwms.gif', writer='imagemagick')
# -
|
analysis/splicing/splicing_cleavage_genesis_hek_make_gifs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %config IPCompleter.greedy=True
# %matplotlib inline
# Import our dependencies
import numpy as np
import pandas as pd
# scikit-learn
# https://scikit-learn.org/stable/index.html
import sklearn as skl
from sklearn.datasets import make_blobs
# TensorFlow and tf.keras
# https://www.tensorflow.org/api_docs/python/tf/keras
import tensorflow as tf
# Helper libraries
import matplotlib.pyplot as plt
# +
# Generate dummy dataset
X, y = make_blobs(n_samples=1000, centers=2, n_features=2, random_state=78)
# Creating a DataFrame with the dummy data
df = pd.DataFrame(X, columns=["Feature 1", "Feature 2"])
df["Target"] = y
# Plotting the dummy data
df.plot.scatter(x="Feature 1", y="Feature 2", c="Target", colormap="winter")
# -
# Use scikit-learn to split dataset
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=78)
# Prepare the dataset for our neural network model. As with any machine learning algorithm, it is crucial to normalize or standardize our numerical variables to ensure that our neural network does not focus on outliers and can apply proper weights to each input. In most cases, the more that input variables are normalized to the same scale, the more stable the neural network model is, and the better the neural network model will generalize.
#
# https://www.youtube.com/watch?v=mnKm3YP56PY&t=21s
# +
# Create scaler instance
X_scaler = skl.preprocessing.StandardScaler()
# Fit the scaler
X_scaler.fit(X_train)
# Scale the data
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# -
# Create the Keras Sequential model
nn_model = tf.keras.models.Sequential()
# Add our first Dense layer, including the input layer
nn_model.add(tf.keras.layers.Dense(units=1, activation="relu", input_dim=2))
# In our previous layer, we used a ReLU activation function to enable nonlinear relationships; however, for our classification output, we want to use a sigmoid activation function to produce a probability output.
# Add the output layer that uses a probability activation function
nn_model.add(tf.keras.layers.Dense(units=1, activation="sigmoid"))
# Check the structure of the Sequential model
nn_model.summary()
# The process of informing the model how it should learn and train is called compiling the model.
#
# Depending on the function of the neural network, we'll have to compile the neural network using a specific optimization function and loss metric. The optimization function shapes and molds a neural network model while it is being trained to ensure that it performs to the best of its ability. The loss metric is used by machine learning algorithms to score the performance of the model through each iteration and epoch by evaluating the inaccuracy of a single input.
#
# To enhance the performance of our classification neural network, we'll use the adam optimizer, which uses a gradient descent approach to ensure that the algorithm will not get stuck on weaker classifying variables and features. As for the loss function, we'll use binary_crossentropy, which is specifically designed to evaluate a binary classification model.
# +
# There are two main types of evaluation metrics—the model predictive accuracy and model mean squared error (MSE).
# We use accuracy for classification models and msefor regression models. For model predictive accuracy,
# the higher the number the better, whereas for regression models, MSE should reduce to zero.
# Compile the Sequential model together and customize metrics
nn_model.compile(loss="binary_crossentropy", optimizer="adam", metrics=["accuracy"])
# -
# To train/fit our Keras model, we'll use the fit method and provide the x training values and y training values, as well as the number of epochs.
# Fit the model to the training data
fit_model = nn_model.fit(X_train_scaled, y_train, epochs=100)
# +
# Create a DataFrame containing training history
history_df = pd.DataFrame(fit_model.history, index=range(1,len(fit_model.history["loss"])+1))
# Plot the loss
history_df.plot(y="loss")
# -
# Plot the accuracy
history_df.plot(y="accuracy")
# Evaluate the model using the test data
model_loss, model_accuracy = nn_model.evaluate(X_test_scaled,y_test,verbose=2)
print(f"Loss: {model_loss}, Accuracy: {model_accuracy}")
print("---------------------------------------------------------------------")
print(nn_model.evaluate(X_test_scaled,y_test))
# Predict the classification of a new set of blob data
new_X, new_Y = make_blobs(n_samples=10, centers=2, n_features=2, random_state=42)
new_X_scaled = X_scaler.transform(new_X)
(nn_model.predict(new_X_scaled) > 0.5).astype("int32")
# +
# We'll generate some nonlinear moon-shaped data using Scikit-learn's make_moons method and visualize it using Pandas and Matplotlib.
from sklearn.datasets import make_moons
# Creating dummy nonlinear data
X_moons, y_moons = make_moons(n_samples=1000, noise=0.08, random_state=42)
# Transforming y_moons to a vertical vector
y_moons = y_moons.reshape(-1, 1)
# Creating a DataFrame to plot the nonlinear dummy data
df_moons = pd.DataFrame(X_moons, columns=["Feature 1", "Feature 2"])
df_moons["Target"] = y_moons
# Plot the nonlinear dummy data
df_moons.plot.scatter(x="Feature 1",y="Feature 2", c="Target",colormap="winter")
# +
# We need to split our nonlinear data into training and testing datasets and normalize our datasets.
# Create training and testing sets
X_moon_train, X_moon_test, y_moon_train, y_moon_test = train_test_split(
X_moons, y_moons, random_state=78
)
# Create the scaler instance
X_moon_scaler = skl.preprocessing.StandardScaler()
# Fit the scaler
X_moon_scaler.fit(X_moon_train)
# Scale the data
X_moon_train_scaled = X_moon_scaler.transform(X_moon_train)
X_moon_test_scaled = X_moon_scaler.transform(X_moon_test)
# +
# We'll train our neural network model using the fit method on the nonlinear training data.
# Training the model with the nonlinear data
model_moon = nn_model.fit(X_moon_train_scaled, y_moon_train, epochs=100, shuffle=True)
# +
# Create a DataFrame containing training history
history_df = pd.DataFrame(model_moon.history, index=range(1,len(model_moon.history["loss"])+1))
# Plot the loss
history_df.plot(y="loss")
# -
# Plot the loss
history_df.plot(y="accuracy")
# Generate our new Sequential model
new_model = tf.keras.models.Sequential()
# +
# Add the input and hidden layer
number_inputs = 2
number_hidden_nodes = 6
new_model.add(tf.keras.layers.Dense(units=number_hidden_nodes, activation="relu", input_dim=number_inputs))
# Add the output layer that uses a probability activation function
new_model.add(tf.keras.layers.Dense(units=1, activation="sigmoid"))
# +
# Compile the Sequential model together and customize metrics
new_model.compile(loss="binary_crossentropy", optimizer="adam", metrics=["accuracy"])
# Fit the model to the training data
new_fit_model = new_model.fit(X_moon_train_scaled, y_moon_train, epochs=100, shuffle=True)
# -
# Outlier Identification
df_moons.describe()
# Plotting a variable using Pandas' Series.plot method to look for outliers
plt.boxplot(df_moons["Feature 1"])
plt.show()
# Plotting a variable using Pandas' Series.plot method to look for outliers
plt.boxplot(df_moons["Feature 2"])
plt.show()
# For a neural network to understand and evaluate a categorical variable, we must preprocess the values using a technique called one-hot encoding.
#
# The process of reducing the number of unique categorical values in a dataset is known as bucketing or binning. Bucketing data typically follows one of two approaches:
#
# - Collapse all of the infrequent and rare categorical values into a single "other" category.
# - Create generalized categorical values and reassign all data points to the new corresponding values.
|
01-Keras-Intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# ___
# # Chapter 4 - Dimensionality Reduction
#
# ## Segment 2 - Explanatory factor analysis
# +
import pandas as pd
import numpy as np
import sklearn
from sklearn.decomposition import FactorAnalysis
from sklearn import datasets
# -
# ### Factor analysis on iris dataset
# +
iris = datasets.load_iris()
X = iris.data
variable_names = iris.feature_names
X[0:10,]
# +
factor = FactorAnalysis().fit(X)
pd.DataFrame(factor.components_, columns=variable_names)
# -
|
Ch04/04_02/04_02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # VQ-VAE
# (vector quantised variational auto-encoder)
#
# Based on DeepMind's [Neural Discrete Representation Learning
# ](https://arxiv.org/abs/1711.00937)
import tensorflow as tf
import os
# +
INPUT_SIZE = 3
BATCH_SIZE = 2
CODE_SIZE = 3 # size of a single vector in e (embedding space)
EMBEDDING_COUNT = 2 # no embedding vectors
# +
graph = tf.Graph()
with graph.as_default():
images = tf.placeholder(tf.float32, shape=[BATCH_SIZE, INPUT_SIZE], name='images')
with tf.variable_scope('encoder', reuse=tf.AUTO_REUSE):
encoder_out = images
with tf.variable_scope('embedding'):
embedding_space = tf.placeholder(tf.float32, shape=[EMBEDDING_COUNT, CODE_SIZE], name='embedding_space')
embedding_space_batch = tf.reshape(tf.tile(embedding_space, [BATCH_SIZE, 1]), [BATCH_SIZE, EMBEDDING_COUNT, CODE_SIZE])
encoder_tiled = tf.reshape(tf.tile(encoder_out, [1, EMBEDDING_COUNT]), [BATCH_SIZE, EMBEDDING_COUNT, CODE_SIZE])
differences = tf.subtract(embedding_space_batch, encoder_tiled)
l2_distances = tf.reduce_sum(tf.square(differences), axis=2, name='l2_distances')
e_index = tf.argmin(l2_distances, axis=1, name='e_index')
code = tf.gather(embedding_space, e_index, axis=0, name='lookup_result')
init = tf.group(tf.global_variables_initializer(), tf.local_variables_initializer())
summary_merged = tf.summary.merge_all()
# -
encoder_tiled.shape
# +
sess = tf.Session(graph=graph)
sess.run(init)
input_val = [[1,2,3],[4,5,6]]
embedding_space_val = [[0,2,4],[1,3,5]]
def get_val(tensor):
return sess.run(tensor, feed_dict={
images: input_val,
embedding_space: embedding_space_val
})
with sess.as_default():
print("input")
print(input_val)
print("embeddings")
print(embedding_space_val)
print("----------------")
print("1")
print("encoder tiled")
print(get_val(encoder_tiled))
print("----------------")
print("2")
print("embedding space batch")
print(get_val(embedding_space_batch))
print("----------------")
print("3")
print("distances")
print(get_val(l2_distances))
print("argmin indices")
print(get_val(e_index))
print("----------------")
print("4")
print("code; gather bs")
print(get_val(code))
# -
#
|
autoencoder/vq-vae/vq-vae-tiny.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 2. Neural Networks with Numpy
# In this notebook we will build our first neural network using only `numpy` as library.
# We will work on the same dataset as last week and try to predict which digit is shown on the given pixel values.
from sklearn.datasets import fetch_openml
X, y = fetch_openml('mnist_784', version=1, return_X_y=True, data_home="./data", cache=True)
# We know already from last time how the data looks:
X.head(3)
# The label is a number between 0-9 representing the digit shown on the pixels.
y.head(3)
# As we can see from above, the label is given as a digit. However, to calculate the loss function of the neural network, we need the label as a one-hot-encoded version, in which the label is encoded as `1` and the rest as `0`.
#
# For instance:
# - `3` -> `[0, 0, 0, 1, 0, 0, 0, 0, 0, 0]`
# - `9` -> `[0, 0, 0, 0, 0, 0, 0, 0, 0, 1]`
#
# This is done in the following:
import pandas as pd
y_categorical = pd.get_dummies(y).astype('float32').values
y_categorical[0:5]
# Before we start, we scale the data and divide it into train and test data:
# +
from sklearn.model_selection import train_test_split
X_scaled = (X/255).astype('float32').values
X_train, X_test, y_train, y_test = train_test_split(X_scaled, y_categorical, test_size=0.15, random_state=42)
# -
# ## Task 1: Implement the Forward pass
# We start with the following structure of a neural network with two hidden layers:
# - Input layer of size 784 with sigmoid activation
# - First hidden layer of size 128 with sigmoid activation
# - Second hidden layer of size 64 with sigmoid activation
# - Output layer of size 10 with softmax activation
#
# A skeleton code for this network is given in the following class. Your first task is to complete the method `forward_pass` to calculate the forward pass on one data point. After this class you can find a test to check whether your implementation is correct.
# +
import time
import numpy as np
class DeepNeuralNetwork():
def __init__(self):
# initialize weights randomly
np.random.seed(0)
self.w1 = np.random.randn(128, 784)
self.w2 = np.random.randn(64, 128)
self.w3 = np.random.randn(10, 64)
def forward_pass(self, x_train):
z1 = np.dot(self.w1, x_train)
a1 = 1/(1 + np.exp(-z1)) # sigmoid activation
z2 = np.dot(self.w2, a1)
a2 = 1/(1 + np.exp(-z2)) # sigmoid activation
z3 = np.dot(self.w3, a2)
a3 = np.exp(z3)/sum(np.exp(z3)) # softmax activation
# we need to remember all values for backpropagation
self.fwdpass = [x_train, z1, a1, z2, a2, z3, a3]
return a3
def backprop(self, y, y_hat):
# restore values from foward pass
a0, z1, a1, z2, a2, z3, a3 = self.fwdpass
# Calculate W3 update
exps = np.exp(z3 - z3.max())
softmax_derivative = exps / np.sum(exps, axis=0) * (1 - exps / np.sum(exps, axis=0))
error = 2 * (y_hat - y) / y_hat.shape[0] * softmax_derivative
gradient_w3 = np.outer(error, a2)
# Calculate W2 update
sigmoid_derivative = (np.exp(-z2))/((np.exp(-z2)+1)**2)
error = np.dot(self.w3.T, error) * sigmoid_derivative
gradient_w2 = np.outer(error, a1)
# Calculate W1 update
sigmoid_derivative = (np.exp(-z1))/((np.exp(-z1)+1)**2)
error = np.dot(self.w2.T, error) * sigmoid_derivative
gradient_w1 = np.outer(error, a0)
return [gradient_w1, gradient_w2, gradient_w3]
# +
# Test for task 1:
dnn = DeepNeuralNetwork()
# the network outputs a probability for every neuron in the last layer
y_hat = dnn.forward_pass(X_train[0])
print("The output of the last layer looks like this:\n", y_hat)
# to check if the network works correctly, check if the following condition is True
abs(y_hat[8] - 0.946) < 0.001
# -
print(dnn.forward_pass(X_train[0]))
# ### Task 2: Implement the training procedure
# We can now start training the network by implementing the training procedure. We train the network for 10 epochs as shown in the code below.
# In each epoch we go over every data point `x` in `X_train` and:
# 1. Calculate a forward pass on `x` as `y_hat`
# 2. Calculate the gradients for the weight in w1, w2 and w3 using the `backprop` function of the network
# 3. Update the weights w1, w2 and w3 of the network by moving into the negative direction of the gradient multiplied with the `learning_rate`
# 4. Bonus: Calculate the cross-entropy-loss after each epoch and plot it in relation to the epochs.
#
#
# +
dnn = DeepNeuralNetwork()
no_epochs = 10
learning_rate = 0.01
start_time = time.time()
losses = []
for iteration in range(no_epochs):
loss = 0
for x,y in zip(X_train, y_train):
y_hat = dnn.forward_pass(x)
gradients = dnn.backprop(y, y_hat)
loss += -np.log(np.dot(y, y_hat))
# one step stochastic gradient descent
dnn.w1 -= learning_rate * gradients[0]
dnn.w2 -= learning_rate * gradients[1]
dnn.w3 -= learning_rate * gradients[2]
print(f'Epoch: {iteration+1}, Time: {time.time() - start_time:.1f}s, Loss: {loss:.0f}')
losses.append(loss)
# -
import matplotlib.pyplot as plt
fig = plt.figure()
plt.plot(range(0, no_epochs), losses)
plt.xlabel('number of epochs')
plt.ylabel('loss')
# ### Task 3: Predict on the test data
# After the network is trained, we can use it to predict on the test data.
#
# __Task__:
# - Iterate over the test data and use the trained network the predict on every test data point.
# - Identify the index of the neuron which returned the highest probability.
# - Compare this value to the true label in the test data.
# - Compute the accuracy.
predictions = []
for x, y in zip(X_test, y_test):
output = dnn.forward_pass(x)
pred = np.argmax(output)
predictions.append(pred == np.argmax(y))
accuracy = np.mean(predictions)
print(accuracy)
# ### Bonus Task:
# - Remove the first hidden layer. Train the network and check the performance on the test data.
# +
import time
import numpy as np
class DeepNeuralNetworkSmall():
def __init__(self):
# initialize weights randomly
np.random.seed(0)
self.w1 = np.random.randn(64, 784)
self.w2 = np.random.randn(10, 64)
def forward_pass(self, x_train):
z1 = np.dot(self.w1, x_train)
a1 = 1/(1 + np.exp(-z1)) # sigmoid activation
z2 = np.dot(self.w2, a1)
a2 = np.exp(z2)/sum(np.exp(z2)) # softmax activation
# we need to remember all values for backpropagation
self.fwdpass = [x_train, z1, a1, z2, a2]
return a2
def backprop(self, y, y_hat):
# restore values from foward pass
a0, z1, a1, z2, a2 = self.fwdpass
# Calculate W2 update
exps = np.exp(z2 - z2.max())
softmax_derivative = exps / np.sum(exps, axis=0) * (1 - exps / np.sum(exps, axis=0))
error = 2 * (y_hat - y) / y_hat.shape[0] * softmax_derivative
gradient_w2 = np.outer(error, a1)
# Calculate W1 update
sigmoid_derivative = (np.exp(-z1))/((np.exp(-z1)+1)**2)
error = np.dot(self.w2.T, error) * sigmoid_derivative
gradient_w1 = np.outer(error, a0)
return [gradient_w1, gradient_w2]
# +
dnn = DeepNeuralNetworkSmall()
no_epochs = 10
learning_rate = 0.01
start_time = time.time()
losses = []
for iteration in range(no_epochs):
loss = 0
for x,y in zip(X_train, y_train):
y_hat = dnn.forward_pass(x)
gradients = dnn.backprop(y, y_hat)
loss += -np.log(np.dot(y, y_hat))
# one step stochastic gradient descent
dnn.w1 -= learning_rate * gradients[0]
dnn.w2 -= learning_rate * gradients[1]
print(f'Epoch: {iteration+1}, Time: {time.time() - start_time:.1f}s, Loss: {loss:.0f}')
losses.append(loss)
# -
predictions = []
for x, y in zip(X_test, y_test):
output = dnn.forward_pass(x)
pred = np.argmax(output)
predictions.append(pred == np.argmax(y))
accuracy = np.mean(predictions)
print(accuracy)
|
solutions/2_NeuralNets_Numpy-Solution.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + [markdown] id="SWzsE9omeSN1" colab_type="text"
# # Introduction to R
# + [markdown] id="THVk8YtgfLvF" colab_type="text"
# **DO NOT CHANGE THE RUNTIME BECAUSE YOU WON'T BE ABLE TO CHANGE IT BACK!**
# + [markdown] id="_H7u2UKDK9Hj" colab_type="text"
# **Run this segment of code first.**
# + id="be6urrizhGk9" colab_type="code" colab={}
install.packages("gcookbook")
install.packages("doParallel")
library(gcookbook)
library(datasets)
library(ggplot2)
library(lubridate)
# + [markdown] id="zQLKMs59rpvi" colab_type="text"
# # Outline
#
# * How to run R codes
# * Basic syntax
# * Data classes and objects in R
# * File operations
# * Useful functions
# * Managing R packages
# * Flow control structures
# * Graphics
# * Parallel processing
#
#
# + [markdown] id="lW8M-lb-5Sqm" colab_type="text"
# # Introduction
# + [markdown] id="QJGFuLmZsrCT" colab_type="text"
# ## What is R
#
# * R is a programming language for statistical computing
# * Importing, storing, exporting and manipulating data
# * Conducting statistical analyses
# * Displaying the results by tables, graphs, etc.
# * R is also a software environment for the development and implementation of new algorithms.
# * Many graphical user interface to R both free and commercial (e.g. Rstudio and Revolution R (now Microsoft R) ).
# + [markdown] id="OIeuD2OxtFph" colab_type="text"
# ## History of R
#
# * R is a dialect of the S language
# * S was created in 1976 at the Bell Labs as an internal statistical analysis environment
# * Goal of S was “to turn ideas into software, quickly and faithfully".
# * Most well known implementation is S-plus (most recent stable release was in 2010). S-Plus integrates S with a nice GUI interface and full customer support.
# * R was created by <NAME> and <NAME> at the University of Auckland, New Zealand.
# * The R core group was formed in 1997, who controls the source code of R (written in C)
# * The first stable version R 1.0.0 was released in 2000
# * Latest stable version is 4.0.0 released on Apr 24, 2020
# + id="vt-DUyoENrHF" colab_type="code" colab={}
R.Version()
# + [markdown] id="nOe0m3Gdteds" colab_type="text"
# ## Features of R
#
# * Designed for statistical analysis, with rich data analysis functionalities and sophisticated graphical capabilities
# * Available on most platform/OS
# * Active development and (very) active community
# * [CRAN](https://cran.r-project.org/): The Comprehensive R Archive Network
# * Source code and binaries, user contributed packages and documentation
# * More than 15,000 packages available on CRAN (as of May 2020)
# * 6,000 three years ago
# * Free to use!
# + [markdown] id="3xIyLZSX5ZYA" colab_type="text"
# # How to Run R Codes
# + [markdown] id="v_4TM4yIDHwM" colab_type="text"
# There are (at least) three options to run R codes:
# * Google Colaboratory
# * Rstudio
# * HPC clusters
# + [markdown] id="xYFRMtbvJXIb" colab_type="text"
# ## Colab
#
# This is what we are using right now. It is the most convenient - browser-based and no setup is needed. The only thing you need is a Google account. In the meanwhile, we don't have much control on the environment. For instance, we can't choose which version to use and have very limited control on where the data is stored.
# + [markdown] id="RUe1W-Y-DTE2" colab_type="text"
# ## Rstudio
#
# Rstudio is an intergrated development environment (IDE) for R.
#
# * Free to use: [Rstudio website](https://www.rstudio.com/)
# * Its user interface is similar to IDEs, dividing the screen into panes
# * Source code editor
# * Console
# * Workspace
# * Others (help message, plot etc.)
# * Rstudio in a desktop environment is better suited for code development and/or a limited number of small jobs.
#
# Rstudio also provides a collection of very useful cheat sheets [here](https://www.rstudio.com/resources/cheatsheets/).
# + [markdown] id="30bP-0GhLuwQ" colab_type="text"
# ## HPC Clusters
#
# The HPC clusters are good for resource‐demanding workloads, e.g. resource-intense tasks or many small tasks.
# + [markdown] id="2-a6403H6Pf_" colab_type="text"
# # Basic Syntax
# + [markdown] id="aMZx_TR9Dw2p" colab_type="text"
# ## Assignment
#
# For new users, the biggest difference from other languages is perhaps the assignment operator: **R uses "<-" instead of "="**.
# + id="Eu1cJEhrD_aT" colab_type="code" cellView="both" colab={}
x <- 2*4
# + [markdown] id="TFi6rURhD4uB" colab_type="text"
# The contents of the object "x" can be viewed by typing its name at the R prompt.
# + id="toTMccSpEClz" colab_type="code" colab={}
x
# + [markdown] id="xuOcTBQhY29q" colab_type="text"
# Actually, "=" works too, but there is some subtle differences, which are explained [here](https://renkun.me/2014/01/28/difference-between-assignment-operators-in-r/).
# + id="yKI9d3beMFkM" colab_type="code" colab={}
y = 2 * 4
y
# + [markdown] id="uiXEWIdZNLcF" colab_type="text"
# Again, "<-" is the recommended way of assigning a value to a variable.
# + [markdown] id="VmiULzr8sFGu" colab_type="text"
# ## Comment
# + [markdown] id="pnhHTCtQsI5z" colab_type="text"
# In R, any line staring with "#" will be interpreted as a comment.
# + id="oSwNxd-2sO_C" colab_type="code" colab={}
# z <- 2*4
# Nothing will happen.
# + [markdown] id="VMTnDgBtPP5O" colab_type="text"
# **Question #1**: What will happen if we execute the code block below?
# + id="lEFCvCPYPLXU" colab_type="code" colab={}
z
# + [markdown] id="aj5qv2UPaTSd" colab_type="text"
# ## Legal R Names
#
# Names for R objects can be any combination of letters, numbers, underscores (_) and periods (.), but must not start with a number or an underscore.
#
#
# + [markdown] id="xVbejwVyb1Ol" colab_type="text"
# These are legal names:
# + id="YSN3PuAFVJu1" colab_type="code" colab={}
num.Cats.2 <- 4
num.Cats.2
num_Cats <- 5
num_Cats
# + [markdown] id="sHxhm98ScLSp" colab_type="text"
# These are not:
# + id="kj6l6SZCbF4b" colab_type="code" colab={}
_num.cats <- 5
# + id="0LO-G6EPbVrX" colab_type="code" colab={}
num-cats <- 5
# + id="_zRGup_2bM-o" colab_type="code" colab={}
2cats <- 3
# + [markdown] id="ss3qglkYVg46" colab_type="text"
# R is case sensitive, e.g. X and x are different in R.
# + id="QKKCqeEnaokZ" colab_type="code" cellView="code" colab={}
x <- 4
print("The value of x is:")
x
print("The value of X is:")
X
# + [markdown] id="wkGt0NWQdUwj" colab_type="text"
# ## Arithematic operations
#
# Basic arithematic operators: +, -, *, /, ^
# + id="DXcHNQF9dmci" colab_type="code" colab={}
1 + 2*4^(3/5)
# + [markdown] id="88HfwU_QWEJO" colab_type="text"
# Scientific notation: 1e-2
# + id="I_wfRTY7WHUh" colab_type="code" colab={}
1e2 + 1e-2
# + [markdown] id="Um6jDvCUdnLu" colab_type="text"
# Special values: Inf (non-finite numeric values), NaN (not a number)
# + id="DeB4Mt5geDNH" colab_type="code" colab={}
1 / 0
-1 / 0
1/0 - 1/0
# + [markdown] id="SZd-QycYr8Pu" colab_type="text"
# ## Comparisons and logical operations
# + [markdown] id="fqU6sXBLsJi0" colab_type="text"
# Comparisons that will return a logical value:
# * Less than: ```<```
# * Less than or equal to: <=
# * Greater than: >
# * Greater than or equal to: >=
# * Equal to: ==
# * Not equal to: ```!=```
# + id="Axk5EW6KsPRF" colab_type="code" colab={}
1 > 2
# + id="Jczs9X8YsSvH" colab_type="code" colab={}
1 != 2
# + [markdown] id="boUXHSI_sf0N" colab_type="text"
# Logical operations:
# * NOT: !
# * AND (elementwise): &
# * OR (element wise): |
# + [markdown] id="xjWsuIi0P_Bi" colab_type="text"
# **Question #2** What is the value of the expression below?
# + id="xEQeIUZpsmrn" colab_type="code" colab={}
! 1 < 2
# + id="Ob7o4faBszo3" colab_type="code" colab={}
a <- 4
b <- 5
a < 10
b < 3
a < 10 & b < 3
a < 10 | b < 3
# + [markdown] id="8Y51nnhBst0y" colab_type="text"
# ## Getting help
# + [markdown] id="KlrtggX9dxfA" colab_type="text"
# Getting help is straightforward in R.
#
# For information about specific functions, use **?\<name of funciton>**.
# + id="OowbAimHd1ft" colab_type="code" colab={}
?class
# + [markdown] id="Hdxaxob7s5V2" colab_type="text"
# For a keyword search, use **??\<keyword>**
# + id="YyQSzEjCs4d1" colab_type="code" colab={}
??assignment
# + [markdown] id="a3ye6Yqt6r_d" colab_type="text"
# # Data Classes And Objects
# + [markdown] id="fA2WGM3oGiae" colab_type="text"
# ## Atomic Date Types
#
# R has five atomic classes:
#
#
# + [markdown] id="fd3ruUJvdWuQ" colab_type="text"
# * Numeric (double)
# * Numbers in R are treated as numeric unless specified otherwise.
#
# Note: the function **class()** reveals the class of a R object.
# + id="i2elyxrhWhPy" colab_type="code" colab={}
class(9.3)
class(3)
# + [markdown] id="McaKmgZMWl51" colab_type="text"
# * Integer
# + id="cvk20CBhWnj-" colab_type="code" colab={}
class(as.integer(3))
# + [markdown] id="yYB1TMMEWrcJ" colab_type="text"
# * Complex
# + id="SAoGwy9MWtM_" colab_type="code" colab={}
class(3+2i)
# + [markdown] id="lKY4G4AnW_8x" colab_type="text"
# * Character
# + id="ZSwPOLdTXCL6" colab_type="code" colab={}
class("a")
class("a cat")
class(a)
# + [markdown] id="PBfr5grGQ_vX" colab_type="text"
# **Question #3** Why `a` is the type returned by `class(a)`?
# + [markdown] id="MTbSNbhBXTL_" colab_type="text"
# * Logical (T, TRUE, F, FALSE)
# * Note that they must be upper case
#
#
# + id="xkfyJst0g9ju" colab_type="code" colab={}
class(TRUE)
class(T)
class(True)
# + [markdown] id="lqirzoNChx7X" colab_type="text"
# The **is.\<type>()** functions, which return logical values, can be used to check for the data classes too.
# + id="MA3mzcUQnb7Y" colab_type="code" colab={}
a <- 3
is.numeric(a)
is.logical(a)
# + [markdown] id="HTcdbbaHiDx9" colab_type="text"
# ## Derivative Data Types
#
# There are many derivative data types, built using the atomic ones. For exmple, the "Date" type.
# + id="B0xwwtZKiLCc" colab_type="code" colab={}
mydate <- today()
mydate
class(mydate)
# + [markdown] id="_HQcqdnMjDM6" colab_type="text"
# ## Data Objects
#
# Now let's look at the data objects in R. They are:
#
# * Vector: elements of same class, one dimension
# * Matrix: elements of same class, two dimensions
# * Array: elements of same class, 2+ dimensions
# * Lists: elements can be any objects
# * Data frames: “datasets” where columns are variables and rows are observations
# + [markdown] id="f_kFtIRtjW8S" colab_type="text"
# ### Vectors
#
#
# + [markdown] id="GpbKM_HNeirE" colab_type="text"
# Vectors contain elements of the **same** data type.
# + [markdown] id="SPX6G_1fcc4J" colab_type="text"
# Vectors can be constructed by
# * The **c()** funtion (concatenate):
# + id="CUfcZG3CAK9f" colab_type="code" colab={}
d <- c(1,2,3)
d
class(d)
# + id="s676HbyOAQER" colab_type="code" colab={}
d <- c("a","b","c")
d
# + [markdown] id="2is8Y5wgcsiO" colab_type="text"
# * The **vector()** function
# * The vector will be initiated to the default values.
# + id="xs233Ouie3MU" colab_type="code" colab={}
d <- vector("numeric", length = 10)
d
# + [markdown] id="fNSEYjM-cybd" colab_type="text"
# * The **seq()** and **rep()** functions, or the ":" operator
# + id="mLvAqfzaongH" colab_type="code" colab={}
d <- seq(from=2,to=10,by=2)
d
# + id="ANI4qAC6oplD" colab_type="code" colab={}
d <- seq(from=2,to=10,length=5)
d
# + id="fpWDtEAJorFK" colab_type="code" colab={}
d <- rep(5,6)
d
# + id="bQTIhhr-fHbK" colab_type="code" colab={}
d <- 1:15
d
# + [markdown] id="UZ5rhMVif3_l" colab_type="text"
# * Or a combination of all of them
# + [markdown] id="860Un_oua77-" colab_type="text"
# **Question #4** How many elements does `d` have?
# + id="z5940hpBf84c" colab_type="code" colab={}
a <- 2:4
d <- c(1,2,8:10,rep(a,3))
d
# + [markdown] id="699QJqLcgSuA" colab_type="text"
# You can convert an object to a different type using the **as.\<TYPE>()** functions.
#
# Note: the output will be an object of the specified type, while the input remains untouched.
# + id="3M78WQu8gfV0" colab_type="code" colab={}
# d is a numeric array.
d <- 1:3
d
# The output of as.character(d) is a character vector.
as.character(d)
# d is still a numeric array.
d
# + [markdown] id="mldhHejUp6vN" colab_type="text"
# When converting to logical values, a numeric "0" will be FALSE while all non-zeroes will be TRUE.
# + id="1n7dmSZ8qE72" colab_type="code" colab={}
as.logical(0:6)
# + [markdown] id="3sHvzlJcgeMY" colab_type="text"
# Coercion will occur when mixed objects are passed to the **c()** function, as if an **as.\<Type>()** function is explicitly called.
# + [markdown] id="mdQheBkRpS5a" colab_type="text"
# **Question #5** Which class will `d` be at the end of this segment?
# + id="8t53vPtSAQGd" colab_type="code" colab={}
d <- c(1e3,"a")
d
class(d)
# + [markdown] id="fNKo3ZSEphjc" colab_type="text"
# How about this one?
# + id="MvkIhdRNr79q" colab_type="code" colab={}
c(1.7,"a")
# + [markdown] id="rpuZSD3MplcP" colab_type="text"
# And this one?
# + id="SdTJOA57rrWH" colab_type="code" colab={}
c(T,2)
# + [markdown] id="Qc7agGbzoIwp" colab_type="text"
# Caution: type coercion may happen without you being aware of it and may have unintended results.
# + [markdown] id="2eI7whmShOZc" colab_type="text"
# One can use the [\<index>] operator to access individual element in a vector. Note that in R the indices start from 1.
# + id="SMkjHhcIhYZr" colab_type="code" colab={}
d <- 1:10
d[4]
# + [markdown] id="cjWS9E1_qijG" colab_type="text"
# For multiple elements with multiple indices, use the **c()** function.
# + id="0Y9hNAIgqh-N" colab_type="code" colab={}
d[1,4]
d[c(1,4)]
# + [markdown] id="1J2tOhUIrNac" colab_type="text"
# Logical values can be used to access individual elements too.
# + id="0Oub6edFrUYI" colab_type="code" colab={}
d[c(T,T,F)]
# + [markdown] id="RhFb7_QJrWc2" colab_type="text"
# Negative indices will drop the corresponding elements from the vector.
# + id="rXhbIOw3rbh-" colab_type="code" colab={}
d[-6:-2]
# + [markdown] id="CtYnVRNuBUGA" colab_type="text"
# **Important:** Lots of R operations process objects in a vectorized way
# * More efficient, concise, and easier to read.
# + id="1FAlkls2Ba3G" colab_type="code" colab={}
x <- 1:4
y <- 6:9
# + id="wRXPJis8ig0R" colab_type="code" colab={}
x + y
# + id="KTncXEdzipxM" colab_type="code" colab={}
x > 2
# + id="Ox4ry-10irS5" colab_type="code" colab={}
x > y
# + [markdown] id="YUVei3zyp8G9" colab_type="text"
# **Question #6** What do you expect to see?
# + id="dBUPBHF7isXg" colab_type="code" colab={}
x[x > 2]
# + [markdown] id="VKld5NEAAoih" colab_type="text"
# ### Matrices
#
# + [markdown] id="KhSI7T6UCPpk" colab_type="text"
# In R, matrices are merely vectors with a "dimension" attribute. Therefore, as in vectors, elements in matrices must be of the same type as well.
#
# + [markdown] id="oko-tFFqltHz" colab_type="text"
# Passing an "dim" attribute to a vector turns it into a matrix.
# + id="Qme15v9SlyJ2" colab_type="code" colab={}
m <- 1:12
m
dim(m) <- c(3,4)
m
# + [markdown] id="fvrNbJrMlmV2" colab_type="text"
# R matrices can be constructed by using the **matrix()** function as well.
# + id="GkZiv9n5lqA6" colab_type="code" colab={}
matrix(1:12,nrow=3,ncol=4)
# + [markdown] id="Uvl_J9sSlyVs" colab_type="text"
# Or by using the **cbind()** or **rbind()** functions.
# + id="h-iS3R5ZmBa6" colab_type="code" colab={}
cbind(1:3,4:6,7:9,10:12)
# + [markdown] id="1-AqG09RmBmO" colab_type="text"
# R matrices are constructed column‐wise.
#
# + id="XUMg5yZkmFVq" colab_type="code" colab={}
matrix(1:12,nrow=3,ncol=4)
matrix(1:12,nrow=3,ncol=4,byrow=T)
# + [markdown] id="8EjvH1f7mFkK" colab_type="text"
# • One can use [\<index>,\<index>] to access individual elements.
# + id="M6stfeDhm5G4" colab_type="code" colab={}
m[3,4]
# + [markdown] id="li5JUqAwCYNj" colab_type="text"
# ### Arrays
# + [markdown] id="JxDOrW-pCaIq" colab_type="text"
# Arrays consist of elements of same class with a number of dimensions. Vectors and matrices are arrays of 1 and 2 dimensions.
# + id="Qzhr0Nw4nH27" colab_type="code" colab={}
# a will be a three-dimensional array
a <- array(data = 1:12,dim = c(2,2,3))
a
a[1,1,2]
dim(a) <- c(3,4)
a[1,1,2]
# + [markdown] id="vhIuKcVJCi4v" colab_type="text"
# ### Lists
# + [markdown] id="OJ0yD7ucClYs" colab_type="text"
# Lists are an ordered collection of objects, which can be of **different types or classes**.
# + [markdown] id="HTzfaqMknnJ2" colab_type="text"
# Lists can be constructed by using the **list()** function.
#
# + id="S8Usv5C_nvQX" colab_type="code" colab={}
list(1,F,"a")
# + [markdown] id="aK6s7l1Mn-AY" colab_type="text"
# Members of a list do not have to be of atomic types, i.e. they can be vectors, matrices and even lists.
# + id="2GcZ0WG0n9Vl" colab_type="code" colab={}
mylist <- list(1,1:5,matrix(1:6,2,3),list(1,F,"a"))
mylist
# + [markdown] id="rzJ2Bx4UnuIO" colab_type="text"
# Lists can be indexed using the [[ ]] operator.
# + id="lMXnv97wocWl" colab_type="code" colab={}
mylist[[2]]
mylist[[4]][[2]]
# + [markdown] id="YL7uvFvXo3rR" colab_type="text"
# Elements of R objects can have names.
# + [markdown] id="_dDKckUGqxTv" colab_type="text"
# Names can be specified when an object is created.
# + id="XerI2_7Mq1N9" colab_type="code" colab={}
list(inst="LSU",location="Baton Rouge",state="LA")
# + [markdown] id="V-mAptaDrGSF" colab_type="text"
# Or they can be specified later when the **names()** function.
# + id="VVxgGiDno7kw" colab_type="code" colab={}
names(mylist) <- c("num","vec","mat","lst")
names(mylist)
# + [markdown] id="kHj7ByHqpEoM" colab_type="text"
# Names can be used to access elements in a data object using the $ operator.
# + id="6yRq-skdpes_" colab_type="code" colab={}
mylist$lst
# + [markdown] id="kNKYP4SbpkJm" colab_type="text"
# Indexing operations by names and indices can be nested and mixed.
# + id="HmOCuFmDpEFY" colab_type="code" colab={}
names(mylist$lst) <- c("c1","c2","c3")
mylist$lst$c2
mylist$lst[[2]]
mylist[[4]][[2]]
# + [markdown] id="iJPzTbBHCsVo" colab_type="text"
# ### Data frames
# + [markdown] id="TpXO9HkvCuKv" colab_type="text"
# Data frames are used to store tabular data.
# * They are a special type of lists where every element (i.e. column) has to be of the same length, but can be of different class.
# * Data frames can store different classes of objects in each column.
# * Data frames can have special attributes such as row.names.
# * Data frames can be created by reading data files, using functions such as **read.table()** or **read.csv()** (more on this later).
# * Can be converted to a matrix using the function **data.matrix()**.
# + [markdown] id="H3ReBPXnJ12R" colab_type="text"
# Data frames can be created directly by calling the **data.frame()** function.
# + id="oJNtp7X2J6nu" colab_type="code" colab={}
mydf <- data.frame(c(31,40,50), c("M","F","M"))
mydf
# + [markdown] id="21fJg0nOKCQ6" colab_type="text"
# We usually name the columns so that it's more meanful.
# + id="17B_Kr9yKBVI" colab_type="code" colab={}
names(mydf) <- c("age","sex")
mydf
# + [markdown] id="1yM746_VKccW" colab_type="text"
# Row names can be specified as well.
# + id="Xp5vrwERKfEd" colab_type="code" colab={}
row.names(mydf) <- c("obs1","obs2","obs3")
mydf
# + [markdown] id="rEbhKOExMal6" colab_type="text"
# To access individual elements in a data frame, there are a few options:
# + [markdown] id="xRgdLN_aKtQ8" colab_type="text"
# * Numeric indices
# + id="4cGXFHP2KwIG" colab_type="code" colab={}
mydf[1,2]
# + [markdown] id="3sWGcKObLA5Y" colab_type="text"
# * Row and column names
# + id="8bHbKVsCLDtj" colab_type="code" colab={}
mydf["obs1","sex"]
# + [markdown] id="MeKU0OXBLIjJ" colab_type="text"
# * Or a mix of indices and names
# + id="MstNHSCrLH_o" colab_type="code" colab={}
mydf["obs1",2]
# + [markdown] id="PsXKfQlXLWs2" colab_type="text"
# Since data frames are lists, both **[[ ]]** and **$** operators work.
# + id="TfBrE6PeLdf9" colab_type="code" colab={}
mydf[[2]]
# + id="HRI_YmX8LgF9" colab_type="code" colab={}
mydf$sex
# + [markdown] id="NBZf5ysyLkx2" colab_type="text"
# You could select rows by leaving the other index blank:
# + id="4Inl8zwxL6O7" colab_type="code" colab={}
mydf[1:2,]
# + [markdown] id="K6HhjQlJMAjd" colab_type="text"
# And vice versa:
# + id="XiA_kAQ6MB8T" colab_type="code" colab={}
mydf[,"sex"]
# + [markdown] id="15tZZUY7MH5N" colab_type="text"
# We can subset a data frame like this:
# + id="GAI5DLeLMHU8" colab_type="code" colab={}
mydf[c(1,3),c("age","sex")]
# + [markdown] id="VB0wM1JC51i_" colab_type="text"
# Or using a vector of logical values:
# + id="PnLyzVSr57CH" colab_type="code" colab={}
mydf[mydf$sex == "M",]
# + [markdown] id="HYy-JykcC7si" colab_type="text"
# ## Querying Object Attributes
# + [markdown] id="0iNRTLBU3Uos" colab_type="text"
# There are a few functions in R that help us obtain information about an object.
# + [markdown] id="q4Sohnrh3hVg" colab_type="text"
# We will work with the "mtcars" data frame in this section.
# + id="P5dwnwu33dVt" colab_type="code" colab={}
?mtcars
# + [markdown] id="kcQlMV6T2GNP" colab_type="text"
# We have already seen the **class()** function, which reveals the type of an object.
# + id="cMcVXZ3O2LNm" colab_type="code" colab={}
class(mtcars)
# + [markdown] id="aAGjyA4G2fVE" colab_type="text"
# The **length()** function shows the length of an object.
# + id="refsgm-J2m18" colab_type="code" colab={}
length(mtcars)
# + [markdown] id="pQSUwXw67Pe0" colab_type="text"
# The **nrow()** function counts the number of rows in a data frame.
# + id="XwL9Gvf57UfI" colab_type="code" colab={}
nrow(mtcars)
# + [markdown] id="M9pPtTER2eCj" colab_type="text"
# The **dim()** function reveals the dimension of an object.
# + id="G8UwvD7Y25oH" colab_type="code" colab={}
dim(mtcars)
# + [markdown] id="7Qfm9mwe3EOc" colab_type="text"
# The **attributes()** function reveals attributes of an object.
# + id="1FSg1jgJ3QXh" colab_type="code" colab={}
attributes(mtcars)
# + [markdown] id="MAIM2wbHC_Ra" colab_type="text"
# The **str()** function shows the internal strucutre of a R object.
#
# + id="XaDIpQZy3vAN" colab_type="code" colab={}
str(mtcars)
?str
# + [markdown] id="Gmt6NO9RGRuU" colab_type="text"
# ## Exercise 1
# + [markdown] id="7KLECRpoMhCi" colab_type="text"
# 1. Learn about the **airquality** data frame and answer the following quetions:
# * What is the source of the data?
# * How many rows and columns are there in the data frame?
# * What does each column represent?
# 2. Find the percentage of days when the high tempature measured at La Guardia Airport exceeded 70.
# 3. Find the number of days when the wind speed is between 10 and 20 miles per hour.
# + [markdown] id="4cWzF8Xw6bOH" colab_type="text"
# ### Solution
# + id="xvTL9soMkWz3" colab_type="code" cellView="both" colab={}
#@title
# question 1
?airquality
attach(airquality)
# question 2
nrow(airquality[Temp > 70,])/nrow(airquality)*100
# question 3
nrow(airquality[Wind > 10 & Wind < 20,])
detach(airquality)
# + [markdown] id="3yN1DHTr6xHi" colab_type="text"
# # Flow Control Structures
# + [markdown] id="jyWSGLWCNP2T" colab_type="text"
# Flow control structures in R, which allow one to control the flow of execution, are similar to other languages.
# + [markdown] id="5-VsMAtmdUye" colab_type="text"
# ## Condition testing
# + [markdown] id="4j0XFnjHcNlM" colab_type="text"
# Test a condition with the if...else structure:
#
# ```
# if (condition 1 is true) {
# do something
# } else if (condition 2 is true) {
# do something else
# } else {
# do something more
# }
# ```
#
#
# + id="HWAur64lb4gj" colab_type="code" colab={}
if (length(mtcars) > 3) {
print("We have more than 3 cars!")
}
# + [markdown] id="MZDqVv23da8F" colab_type="text"
# ## Loops
# + [markdown] id="XDfW1jMEc1Eg" colab_type="text"
# Loops with ```for```:
#
# ```
# for (variable in sequence) {
# statements
# }
# ```
# + id="-WRBrRHpejAD" colab_type="code" colab={}
for (i in 1:10) {
print(i^3)
}
# + [markdown] id="L_u6GzW2NZPr" colab_type="text"
# Loops are not very frequently used in R because many operations are inherently vectorized. This line of code does exactly the same thing:
# + id="jZSW9cgDesDM" colab_type="code" colab={}
(1:10)^3
# + [markdown] id="_X_MN4gY76rb" colab_type="text"
# The family of **apply()** functions are also very useful to perform operation over all elements of a vector/list (see next section).
# + [markdown] id="QYHSyiO264Lo" colab_type="text"
# # Useful Functions
# + [markdown] id="SSHcFWAE8d1O" colab_type="text"
# ## Simple Statistic Functions
# + [markdown] id="ANfns3j68jYV" colab_type="text"
# * **min()**: Minimum value
# * **max()**: Maximum value
# * **which.min()**: Location of minimum value
# * **which.max()**: Location of maximum value
# * **sum()**: Sum of the elements of a vector
# * **mean()**: Mean of the elements of a vector
# * **sd()**: Standard deviation of the elements of a vector
# * **quantile()**: Show quantiles of a vector
# * **summary()**: Display descriptive statistics
# + id="QDrwC_Ef9IbZ" colab_type="code" colab={}
mean(mtcars$mpg)
# + id="XZvXBHym9DkW" colab_type="code" colab={}
summary(mtcars)
# + [markdown] id="Tu3bLnYBmDHt" colab_type="text"
# ## Distributions and Random Variables
# + [markdown] id="buoMxY5DmLdF" colab_type="text"
# For each distribution R provides four functions: density (d),
# cumulative density (p), quantile (q), and random generation (r).
#
# + [markdown] id="D9gZxNN7mRij" colab_type="text"
# * Distrituion | Name in R
#
# * Uniform | ```unif```
# * Binomial | ```binom```
# * Poisson | ```pois```
# * Geometric | ```geom```
# * Gamma | ```gamma```
# * Normal | ```norm```
# * Log Normal | ```lnorm```
# * Exponential | ```exp```
# * Student's t | ```t```
# + [markdown] id="hcrAZzrGNibA" colab_type="text"
# The function name is of the form [d|p|q|r]\<name of
# distribution\>. For example, **qbinom()** gives the quantile of a binomial distribution.
# + [markdown] id="hVGzmYutJpuN" colab_type="text"
# Generate a random sample of 10 from the standard normal distribution:
# + id="hb2F99Jumg4H" colab_type="code" colab={}
rnorm(10,mean=0,sd=1)
# + [markdown] id="rRgDgUcHJxXQ" colab_type="text"
# The p-value for 1.96 and its inverse function (standard normal distribution):
# + id="iY-8nNGCmk8C" colab_type="code" colab={}
pnorm(1.96)
qnorm(pnorm(1.96))
# + [markdown] id="FRXeOROPSjqn" colab_type="text"
# When genrating random samples, setting the seed to the same value will
# + [markdown] id="24qZNXo0m0uI" colab_type="text"
# ## Sorting
# + [markdown] id="8pWcpOyMwHAJ" colab_type="text"
# Sort and order elements: sort(), rank() and order().
# + [markdown] id="P_msD9ddQSit" colab_type="text"
# By default, the **sort()** functions sorts the values in a vector into ascending order.
# + id="0i-6iE8VQF-J" colab_type="code" colab={}
sort(mtcars$mpg)
# + [markdown] id="Oc0PXkVNQ7pO" colab_type="text"
# The ```decreasing=T``` option will sort a vector into descending order.
# + id="ILBz51yeRDYp" colab_type="code" colab={}
sort(mtcars$mpg, decreasing=T)
# + [markdown] id="x5hR2mzowLyt" colab_type="text"
# In contrast, the **order()** functions returns the indices in order.
#
# + id="h_pfkmHRSqkP" colab_type="code" colab={}
order(mtcars$mpg)
order(mtcars$mpg, decreasing=T)
# + [markdown] id="qYsP8N_wS1lL" colab_type="text"
# Users can use the indices returned by **order()** to change the order of rows in a data frame.
# + id="gIPorrrfTOSk" colab_type="code" colab={}
mtcars
# + id="s8_7WeLDTSdP" colab_type="code" colab={}
mtcars[order(mtcars$mpg),]
# + [markdown] id="9ytChwxam-Wt" colab_type="text"
# ## Table
# + [markdown] id="3s7c6x5BwU6q" colab_type="text"
# The **table()** function tabulates factors or find the frequency of
# an object.
#
# + [markdown] id="LWpycIz8wYhN" colab_type="text"
# For instance, in the **mtcars** data frame, we can get the frequency table by the combination of the numbers of cylinders and gears:
# + id="OptkIM5FIZLN" colab_type="code" colab={}
table(mtcars[,c("cyl","gear")])
# + [markdown] id="bIradhn8nAsX" colab_type="text"
# ## The Apply() family of functions
# + [markdown] id="R6ffwwyUwjdc" colab_type="text"
# The **apply()** function evaluate a function over
# the margins of an array
# * More concise than the for loops (not necessarily
# faster)
# + [markdown] id="sUOJzrFxVODO" colab_type="text"
# Syntax:
#
# ```
# apply(data,dimension,function,function perimeters)
# ```
# + [markdown] id="viRlf7S2UACn" colab_type="text"
# For example, if we want to calculate the mean of each row in a matrix:
# + id="vJJ-dHYfT93H" colab_type="code" colab={}
x <- matrix(rnorm(200), 20, 10)
apply(x, 1, mean)
# + [markdown] id="RdDy2C9gUzDZ" colab_type="text"
# Column sum:
# + id="EP_XdvDvU1AS" colab_type="code" colab={}
apply(x,2,sum)
# + [markdown] id="I8rfXbF1VHh_" colab_type="text"
# It can perform multiple calculations in one function call:
# + id="3sg49uJ9VGDY" colab_type="code" colab={}
apply(x, 1, quantile, probs = c(0.25, 0.75))
# + [markdown] id="eXp5rsoOwkG-" colab_type="text"
# Other member of the **apply()** family include:
# * **lapply** - Loop over a list (data frame) and evaluate a function on each element
# * **sapply** - Same as **lapply** but simplifies the result to array
# * **tapply** - Apply a function over subsets of a vector
# * **mapply** - Multivariate version of **sapply**
# + [markdown] id="AzfcYspCY92P" colab_type="text"
# ## The ```plyr``` package
# + [markdown] id="zUfOsrGUZFk7" colab_type="text"
# The "split-apply-combine" pattern is very common in data analysis, where you solve a complex problem by breaking it down into small pieces, doing something to each piece and then combining the results back together again.
# + [markdown] id="_vTHA1NDZSXt" colab_type="text"
# For instance, in the **mtcars** data frame, we may want to know the average mileage-per-gallon for cars with 4, 6 and 8 cylinders. To do that, we will need to split the data into subsets according to the value of the "cyl" column, apply the mean function to the "mpg" columen, then combine the results from each subset.
# + [markdown] id="SXxY5EOxZmfU" colab_type="text"
# The ```plyr``` packages provide a group of functions that implement this split-apply-combine pattern.
#
# For example, the **ddply()** function takes a data frame, split it accorindg to, apply a function to each piece, then combine the result into a new data frame.
# + id="fNnHBd9HZb8O" colab_type="code" colab={}
library(plyr)
ddply(mtcars,"cyl",summarize,AverageMPG=mean(mpg))
# + [markdown] id="n1qNNxzhnFxc" colab_type="text"
# ## User-defined functions
# + [markdown] id="bmYphKDAogk7" colab_type="text"
# * Users can define their own functions in R by using the ```function()``` directives.
# * The return value is the last expression in the function body to be evaluated.
# * Functions can be nested.
# * Functions are R objects and can be passed as an argument to
# other functions.
# + [markdown] id="qGv2mlFFsksf" colab_type="text"
# Syntax to define a function:
#
#
# ```
# function_name <- function (arguments) {
# statements
# }
# ```
#
#
# + id="AorOHxmtsh7F" colab_type="code" colab={}
pow <- function(x, y) {
result <- x^y
}
# + [markdown] id="mlwcx99ptasP" colab_type="text"
# Then it can be called like any other function:
# + id="RZ2SQLz4s5kp" colab_type="code" colab={}
c <- pow(4,2)
c
# + [markdown] id="NydanKGDtfqM" colab_type="text"
# Functions can be used as an argument for other functions.
# + id="rdagOd6WtCWJ" colab_type="code" colab={}
myfunc <- function(func,a,b) {
result <- func(a,b) - 1
}
c <- myfunc(pow,4,2)
c
# + [markdown] id="QNnbwpbd8LdP" colab_type="text"
# ## Exercise 2
#
# Using the **airquality** data, find the average wind speed for the 10 hottest days.
#
# Hint: use the order() function.
# + [markdown] id="OhmXolDQ7cRv" colab_type="text"
# ### Solution
# + id="nybmcy0pbREI" colab_type="code" colab={}
#@title { display-mode: "form" }
mean(airquality[order(airquality$Temp,decreasing=T),"Wind"][1:10])
# + [markdown] id="ndYj2veJO_aI" colab_type="text"
# ## Exercise 3
# + [markdown] id="MzUgrKdNPCR3" colab_type="text"
# Using the **airquality** data, find the average high tempature of each month.
#
# Hint: use the ddply() function.
# + [markdown] id="Az9hNbGkhVMh" colab_type="text"
# ### Solution
# + id="2YqnhxbUhYuX" colab_type="code" cellView="both" colab={}
#@title
ddply(airquality,"Month",summarize,AvgTemp=mean(Temp))
# + [markdown] id="C-jdAOYX652m" colab_type="text"
# # Managing R Packages
# + [markdown] id="CVsBdnMJQ_hO" colab_type="text"
# To load a R package, use the **library()** or **require()** function:
# + id="Z1XbChLQRHlq" colab_type="code" colab={}
library(lubridate)
require(devtools)
# + [markdown] id="qZu1TqAb3lgX" colab_type="text"
# The main difference is that, if a package is not installed, **library()** will throw out an error message and the execution will stop, while **require()** throws out a warning and the execution continues.
# + id="v3TEHkDmRHpB" colab_type="code" colab={}
library(reshape)
print("End of code segment")
# + id="90_zQRz238RN" colab_type="code" colab={}
require(reshape)
print("End of code segment")
# + [markdown] id="pQCqbBgMRNn_" colab_type="text"
# If a package is not available, the **install.packages()** function can be used to install it.
# + id="vTPsokxQRYst" colab_type="code" colab={}
install.packages("reshape")
library(reshape)
# + [markdown] id="3TOGsHvIOLFW" colab_type="text"
# Multiple packages can be installed with one call of the **install.packages()** function.
# + id="ksi6rPshOWpS" colab_type="code" colab={}
require(datarium)
require(BiocManager)
install.packages(c("datarium","BiocManager"))
library(datarium)
library(BiocManager)
# + [markdown] id="vYQV6dMfRcFq" colab_type="text"
# Note that double quotation is **NOT** needed when loading packages, but necessary when installing them.
# + [markdown] id="ykBSP1N6TwTt" colab_type="text"
# Use the **remove.packages** function to remove installed packages.
# + id="DTGxWquhN4I-" colab_type="code" colab={}
remove.packages("datarium")
library(datarium)
# + [markdown] id="-iMVow7sTztO" colab_type="text"
# The **update.packages** function updates installed packages.
# + id="4r1GxL5LN0aK" colab_type="code" colab={}
update.packages("lubricate")
# + [markdown] id="fTdvZYEaT17B" colab_type="text"
# List installed packages.
# + id="SjAI5mpBT4et" colab_type="code" colab={}
installed.packages()
# + [markdown] id="mCFgcP7W8Rvm" colab_type="text"
# List all loaded packages and attached objects.
# + id="7I5HIa6C8VLD" colab_type="code" colab={}
search()
# + [markdown] id="m80UZgL1WfZY" colab_type="text"
# # File And Directory Operations
# + [markdown] id="_uv7F-KZOPNC" colab_type="text"
# ## Query working directory
# + [markdown] id="eiDBqpIwvxuO" colab_type="text"
# Each R session has a working directory. The **getwd()** function shows the current working directory.
# + id="n95VB47DOSZX" colab_type="code" colab={}
getwd()
# + [markdown] id="98aFasSSwAyR" colab_type="text"
# The **list.files()** and **list.dirs()** functions list the files and subdirectories.
# + id="dmJ_u7Nav-b9" colab_type="code" colab={}
list.files()
list.dirs()
# + [markdown] id="mBDmpEZbwMEI" colab_type="text"
# To change the working directory, use the **setwd()** functions.
# + id="BqouRHsLwLXd" colab_type="code" colab={}
setwd("/content/sample_data")
getwd()
# + [markdown] id="lx08L6_9WsGt" colab_type="text"
# ## Handling files
# + [markdown] id="8y1Ya7D5dqKS" colab_type="text"
# R has a set of file.\<opearation> functions.
# + [markdown] id="zpCtVCLrwTIC" colab_type="text"
# Check if a file exists:
# + id="CfC2sko8jDGA" colab_type="code" colab={}
file.exists("testfile")
# + [markdown] id="Sur7zWT7wXNS" colab_type="text"
# Create an empty file:
# + id="nICOYhxZjO-x" colab_type="code" colab={}
file.create("testfile")
file.exists("testfile")
# + [markdown] id="eNVnlzXsxw8I" colab_type="text"
# Copy and delete files:
# + id="2k3z6b6tjZl2" colab_type="code" colab={}
file.copy("testfile","anotherfile")
list.files()
file.remove("testfile","anotherfile")
# + [markdown] id="Ln_66oqI681p" colab_type="text"
# # Graphics
# + [markdown] id="Dn_0FhCwYGh_" colab_type="text"
# There are three plotting systems in R
# * base
# * Convenient, but hard to adjust after the plot is created
# * lattice
# * Good for creating conditioning plot
# * ggplot2
# * Powerful and flexible, many tunable feature, may require some time to master
#
# + [markdown] id="Z08zo1CzYbrL" colab_type="text"
# Each has its pros and cons, so it is up to the users which one to choose.
# + [markdown] id="_sh0dD7NYkY8" colab_type="text"
# ## base
# + [markdown] id="ERWkZMFVgFy_" colab_type="text"
# A few functions are avaible in the base plot systems
# * **plot()**: line and scatter plots
# * **boxplot()**: box plots
# * **hist()**: histograms
# + [markdown] id="oYA7Tm66HD6L" colab_type="text"
# A quick scatter plot example with the base plot system.
# + id="GjrJFgsff6uf" colab_type="code" colab={}
# Create the plot with title and axis labels.
plot(pressure,type="l",
main="Vapor Pressure of Mercury",
xlab="Temperature",
ylab="Vapor Pressure")
# Add points
points(pressure,col='red')
# Add annotation
text(150,700,"Source: Weast, R. C., ed. (1973) Handbook \n
of Chemistry and Physics. CRC Press.")
# + [markdown] id="MenwnCkuYoQt" colab_type="text"
# ## ggplot2
# + [markdown] id="9uKfPUvZHLsT" colab_type="text"
# The **qplot()** function is the ggplot2 version of **plot()**.
# + id="HFOpWRHxg5RW" colab_type="code" colab={}
qplot(weightLb, heightIn, data=heightweight, geom="point")
# + [markdown] id="mQ9kl7IZHZOn" colab_type="text"
# The **ggplot()** function is the main function in the ggplot2 package.
#
# Here is an example:
# + id="7wKAPUK7g6lo" colab_type="code" colab={}
ggplot(heightweight, aes(x=weightLb, y=heightIn, color=sex, shape=sex)) +
geom_point(size=3.5) +
ggtitle("School Children\nHeight ~ Weight") +
labs(y="Height (inch)", x="Weight (lbs)") +
stat_smooth(method=loess, se=T, color="black", fullrange=T) +
annotate("text",x=145,y=75,label="Locally weighted polynomial fit with 95% CI",color="Green",size=6) +
scale_color_brewer(palette = "Set1", labels=c("Female", "Male")) +
guides(shape=F) +
theme_bw() +
theme(plot.title = element_text(size=20, hjust=0.5),
legend.position = c(0.9,0.2),
axis.title.x = element_text(size=20), axis.title.y = element_text(size=20),
legend.title = element_text(size=15),legend.text = element_text(size=15))
# + [markdown] id="gRqDF8nIY5qa" colab_type="text"
# If you are interested to learn more, please visit the [Data Visualization in R](http://www.hpc.lsu.edu/training/weekly-materials/2018-Spring/Slides.html#(1)) tutorial from LSU HPC
# + [markdown] id="-1OjjS0p6_lt" colab_type="text"
# # Parallel Processing
# + [markdown] id="hkeDSGESZ_J0" colab_type="text"
# Modern computers are equipped with more than one CPU core and are capable of processing workloads in parallel, but base R is single‐threaded, i.e. not parallel.
#
# In other words, regardless how many cores are available, R can only
# use one of them.
#
# There are two options to run R in parallel: **implicit** and **explicit**.
#
# + [markdown] id="1Za3yZbWcIBm" colab_type="text"
# ## Implicit parallel processing
# + [markdown] id="SeE5pNzdcaqt" colab_type="text"
# Some functions in R can call parallel numerical libraries.
#
# For instance, on LONI QB2 cluster most linear algebraic and related functions (e.g. linear regression, matrix decomposition, computing inverse and determinant of a matrix) leverage the multi‐threaded Intel MKL library.
#
# In this case, no extra coding is needed to take advange of the multiple CPU cores - those functions will automatically use multiple cores when being called.
# + [markdown] id="KGFBQEzOcMlA" colab_type="text"
# ## Explicit parallel processing
# + [markdown] id="zhE2hltlfSpS" colab_type="text"
# If the implicit option is not available for what you'd like to do, some codes need to be written.
# + [markdown] id="FpyHuV4aBIxK" colab_type="text"
# Here is an example of using the **%dopar%** directive in the **doParallel** package.
# + [markdown] id="bX6HuHivBUxE" colab_type="text"
# The workload is to generate 100 random samples, each with
# 1,000,000 observations from a standard normal distribution, then take a summary for each sample.
# + id="VCru3RBK_fH1" colab_type="code" colab={}
iters <- 100
# + [markdown] id="KCxfQGZSBoSG" colab_type="text"
# Below is the sequential version with a for loop. The **system.time()** function is used to measure how long it takes to process the workload.
# + id="ViGrf8mZ9z2h" colab_type="code" colab={}
system.time(
for (i in 1:iters) {
to.ls <- rnorm(1e6)
to.ls <- summary(to.ls)
}
)
# + [markdown] id="W3HEm1xOB178" colab_type="text"
# This is the parallel example with the **doParallel** package.
# + id="wmk2YWK69l9J" colab_type="code" colab={}
library(doParallel)
system.time({
cl <- makeCluster(2)
registerDoParallel(cl)
ls<-foreach(icount(iters)) %dopar% {
to.ls<-rnorm(1e6)
to.ls<-summary(to.ls)
}
stopCluster(cl)
})
# + [markdown] id="vwM8hXRyZccR" colab_type="text"
# If you are interested to learn more, please visit the [Parallel Computing in R](http://www.hpc.lsu.edu/training/weekly-materials/2017-Fall/HPC_Parallel_R_Fall2017.pdf) tutorial from LSU HPC.
|
day3/Introduction to R.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading data in sktime
#
# Data for use with sktime should be stored in pandas DataFrame objects with cases represented by rows and series data for each dimension of a problem stored in columns (the specifics of the data structure are described in more detail in the section below). Data can be loaded into the sktime format through various means, such as loading directly from a bespoke sktime file format (.ts) or supported file formats provided by other existing data sources (such as ARFF and .tsv). Further, data can also be loaded through other means into a long-table format and then converted to the sktime format using a provided method.
#
# Below is a brief description of the .ts file format, an introduction of how data are stored in dataframes for sktime, and examples of loading data from a variety of file formats.
#
#
# ## Representing data with .ts files
#
# The most typical use case is to load data from a locally stored .ts file. The .ts file format has been created for representing problems in a standard format for use with sktime. These files include two main parts:
# * header information
# * data
#
# The header information is used to facilitate simple representation of the data through including metadata about the structure of the problem. The header contains the following:
#
# @problemName <problem name>
# @timeStamps <true/false>
# @univariate <true/false>
# @classLabel <true/false> <space delimted list of possible class values>
# @data
#
# The data for the problem should begin after the @data tag. In the simplest case where @timestamps is false, values for a series are expressed in a comma-separated list and the index of each value is relative to its position in the list (0, 1, ..., m). A _case_ may contain 1 to many dimensions, where cases are line-delimited and dimensions within a case are colon (:) delimited. For example:
#
# 2,3,2,4:4,3,2,2
# 13,12,32,12:22,23,12,32
# 4,4,5,4:3,2,3,2
#
# This example data has 3 _cases_, where each case has 2 _dimensions_ with 4 observations per dimension. Missing readings can be specified using ?, or for sparse datasets, readings can be specified by setting @timestamps to true and representing the data with tuples in the form of (timestamp, value). For example, the first case in the example above could be specified in this representation as:
#
# (0,2),(1,3)(2,2)(3,4):(0,4),(1,3),(2,2),(3,2)
#
# Equivalently,
#
# 2,5,?,?,?,?,?,5,?,?,?,?,4
#
# could be represnted with timestamps as:
#
# (0,2),(0,5),(7,5),(12,4)
#
# For classification problems, the class label for a case should be specified in the last dimension and @classLabel should be in the header information to specify the set of possible class values. For example, if a case consists of a single dimension and has a class value of 1 it would be specified as:
#
# 1,4,23,34:1
#
# ## Storing data in a pandas DataFrame
#
# The core data structure for storing datasets in sktime is a pandas DataFrame, where rows of the dataframe correspond to cases, and columns correspond to dimensions of the problem. The readings within each column of the dataframe are stored as pandas Series objects; the use of Series facilitates simple storage of sparse data or series with non-integer timestamps (such as dates). Further, if the loaded problem is a classification problem, the standard loading functionality within sktime will return the class values in a separate index-aligned numpy array (with an option to combine X and Y into a single dataframe for high-level task construction). For example, for a problem with n cases that each have data across c dimensions:
#
# DataFrame:
# index | dim_0 | dim_1 | ... | dim_c-1
# 0 | pd.Series | pd.Series | pd.Series | pd.Series
# 1 | pd.Series | pd.Series | pd.Series | pd.Series
# ... | ... | ... | ... | ...
# n | pd.Series | pd.Series | pd.Series | pd.Series
#
# And if the data is a classification problem, a separate (index-aligned) array will be returned with the class labels:
#
# index | class_val
# 0 | int
# 1 | int
# ... | ...
# n | int
#
#
# ## Loading from .ts file to pandas DataFrame
#
# A dataset can be loaded from a .ts file using the following method in sktime.utils.load_data.py:
#
# load_from_tsfile_to_dataframe(full_file_path_and_name, replace_missing_vals_with='NaN')
#
# This can be demonstrated using the Arrow Head problem that is included in sktime under sktime/datasets/data
# +
from sktime.utils.load_data import load_from_tsfile_to_dataframe
import os
import sktime
DATA_PATH = os.path.join(os.path.dirname(sktime.__file__), "datasets/data")
train_x, train_y = load_from_tsfile_to_dataframe(os.path.join(DATA_PATH, "ArrowHead/ArrowHead_TRAIN.ts"))
test_x, test_y = load_from_tsfile_to_dataframe(os.path.join(DATA_PATH, "ArrowHead/ArrowHead_TEST.ts"))
# -
# Train and test partitions of the ArrowHead problem have been loaded into dataframes with associated arrays for class values. As an example, below are the first 5 rows from the train_x and train_y:
train_x.head()
train_y[0:5]
# ## Loading from Weka ARFF files
#
# It is also possible to load data from Weka's attribute-relation file format (ARFF) files. This is the data format used by researchers at the University of East Anglia (available from www.timeseriesclassification.com ). The `load_from_arff_to_dataframe` method in `sktime.utils.load_data` supports reading both univariate and multivariate problems. Examples are shown below using the ArrowHead problem again (this time loading from ARFF) and also the multivariate BasicMotions problem.
#
# ### Loading the univariate ArrowHead problem from ARFF
# +
from sktime.utils.load_data import load_from_arff_to_dataframe
X, y = load_from_arff_to_dataframe(os.path.join(DATA_PATH, "ArrowHead/ArrowHead_TRAIN.arff"))
X.head()
# -
# ### Loading the multivariate BasicMotions problem from ARFF
X, y = load_from_arff_to_dataframe(os.path.join(DATA_PATH, "BasicMotions/BasicMotions_TRAIN.arff"))
X.head()
# ## Loading from UCR .tsv Format Files
#
# A further option is to load data into sktime from tab separated value (.tsv) files, as used by researchers at the University of Riverside, California (available at https://www.cs.ucr.edu/~eamonn/time_series_data_2018 ). The `load_from_ucr_tsv_to_dataframe` method in `sktime.utils.load_data` supports reading univariate problems. An example with ArrowHead is given below to demonstrate equivalence with loading from ARFF and .ts file formats.
#
# ### Loading the univariate ArrowHead problem from .tsv
# +
from sktime.utils.load_data import load_from_ucr_tsv_to_dataframe
X, y = load_from_ucr_tsv_to_dataframe(os.path.join(DATA_PATH, "ArrowHead/ArrowHead_TRAIN.tsv"))
X.head()
# -
# ## Using long-format data with sktime
#
# It is also possible to use data from sources other than .ts and .arff files by manually shaping the data into the format described above. For convenience, a helper function is also provided to convert long-format data into sktime-formatted data in the `from_long_to_nested` method in `sktime.utils.load_data` (with assumptions made on how the data is initially formatted).
#
# The method converts rows from a long-table schema data frame assuming each row contains information for:
#
# `case_id, dimension_id, reading_id, value`
#
# where `case_id` is an id to identify a specific case in the data, `dimension_id` is an integer between 0 and d-1 for d dimensions in the data, `reading_id` is the index of this observation for the associated `case_id` and `dimension_id`, and `value` is the actual value of the observation. E.g.:
#
# | case_id | dim_id | reading_id | value
# ------------------------------------------------
# 0 | int | int | int | double
# 1 | int | int | int | double
# 2 | int | int | int | double
# 3 | int | int | int | double
#
# To demonstrate this functionality the method below creates a dataset with a given number of cases, dimensions and observations:
# +
import numpy as np
import pandas as pd
def generate_example_long_table(num_cases=50, series_len=20, num_dims=2):
rows_per_case = series_len*num_dims
total_rows = num_cases*series_len*num_dims
case_ids = np.empty(total_rows, dtype=np.int)
idxs = np.empty(total_rows, dtype=np.int)
dims = np.empty(total_rows, dtype=np.int)
vals = np.random.rand(total_rows)
for i in range(total_rows):
case_ids[i] = int(i/rows_per_case)
rem = i%rows_per_case
dims[i] = int(rem/series_len)
idxs[i] = rem%series_len
df = pd.DataFrame()
df['case_id'] = pd.Series(case_ids)
df['dim_id'] = pd.Series(dims)
df['reading_id'] = pd.Series(idxs)
df['value'] = pd.Series(vals)
return df
# -
# The following example generates a long-format table with 50 cases, each with 4 dimensions of length 20:
X = generate_example_long_table(num_cases=50, series_len=20, num_dims=4)
X.head()
X.tail()
# As shown below, applying the `from_long_to_nested` method returns a sktime-formatted dataset with individual dimensions represented by columns of the output dataframe:
from sktime.utils.load_data import from_long_to_nested
X_nested = from_long_to_nested(X)
X_nested.head()
X_nested.iloc[0][0].head()
|
examples/loading_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Matplotlib-1</h1>
# <h6>Saket Tiwari</h6>
# Date: 17 June 2019
# +
import matplotlib.pyplot as plt
#from matplotlib import pyplot as plt
# %matplotlib inline
#magic function
# -
plt.plot([10,20,30],[5,2,4])
plt.show()
plt.plot([1,2,3,4],[10,30,20,40],'y')
plt.show()
plt.plot([1,2,3,4],[10,30,20,40],'--')
plt.show()
plt.plot([1,2,3,4],[10,30,20,40],'*')
plt.show()
plt.plot([1,2,3,4],[10,30,20,40],'r--')
plt.show()
plt.plot([1,2,3,4],[10,30,20,40],'r--',linewidth=4)
plt.show()
import numpy as np
x=np.arange(1,20)
y=x ** 2
plt.plot(x,y,'r--')
plt.show()
x=np.arange(1,20)
y=x ** 2
plt.plot(x,y)
plt.title('Square')
plt.xlabel('x-values')
plt.ylabel('x-squares')
plt.grid()
plt.xlim(-5,25) #used to set the x-limit
plt.ylim(20,500) #used to set the y-limit
plt.show()
# +
x=np.arange(-10,20)
y=x ** 2
x2=np.arange(0,30)
y2= x2+50
plt.plot(x,y,'g--',label='quadratic')
plt.plot(x2,y2,'r',label='linear')
plt.title('Square')
plt.xlabel('x-values')
plt.ylabel('x-squares')
plt.legend()
plt.text(8,60,'Hello',)
plt.show()
# matplotlib do not solve equations
#it only shows the points
# -
plt.plot(x,y,'g--',x2,y2,'r--')
x=np.arange(1,10)
y=np.log(x)
#np.log expects list as input
plt.plot(x,y)
plt.show()
# ! pip install math
import math
x=np.arange(1,10)
y=[math.log(i) for i in x]
#math.log expects a vector
plt.plot(x,y)
plt.show()
# +
import math
x=np.arange(1,10)
y=list(map(lambda i:math.log(i),x))
plt.plot(x,y)
plt.show()
# -
import math
x=np.arange(1,100)
y=np.vectorize(math.log)(x)
#vectorize makes the vector
plt.plot(x,y)
plt.show()
# # Scatter
x=np.random.rand(10)
x2=x=np.random.rand(10)
y=np.random.rand(10)
y2=np.random.rand(10)
plt.scatter(x,y)
plt.scatter(x2,y2)
plt.scatter(x,y,color='r')
plt.show()
# # Bar Graph
plt.bar([1,2,3,4],[10,20,30,40],width=.2)
plt.bar([1,2,3,4],[5,10,15,20],width=.2)
plat.
plt.bar(np.array([1,2,3,4])+0.2+[20,30,40,50])
plt.show()
#all the values between 1-2 is representd in 2nd bar
#all the values
# # Histogram
# +
#in histogram we send raw data as input,which automatically changes its height width and height
#it is mainly used to display frequencies
#buckets are nothing but range which is displayed by histogram
# -
vals=[1,2,3,4,5,6,7,8,9,8,5,3,2,2,3,4,5]
plt.hist(vals,3) #buckets can be decided
plt.show()
mu=100
sigma=15
s=np.random.normal(mu,sigma,100)
s
plt.hist(s,30)
plt.show()
# + active=""
#
# -
count, bins,ignored=plt.hist(s,30)
plt.show()
count
bins
# +
#sigmoid
#sigmoid(z)= 1/(1+e^(-z))
sigmoid = lambda x:1/(1+math.exp(-x))
x=np.arange(-10,10)
y=[sigmoid(i) for i in x] #we vectorized the values of y
plt.plot(x,y)
plt.show()
#sigmoid value is used for classification
|
Basics/MatplotLib-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/berthine/SIAM-Summer-School/blob/main/SIAM2021_Autoencoder_2_.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YSon7FLdIS1O"
# ## Practical: Autoencoders (MNIST)
# (26/July/2021)
#
# ### 2021 Gene Golub SIAM Summer School
# https://sites.google.com/aims.ac.za/g2s3/home
#
# Instructor
#
# <font color="green">***Dr. <NAME>***
#
# www.emmanueldufourq.com
#
# edufourq (['@']) gmail.com
#
# ***African Institute for Mathematical Sciences***
#
# ***Stellenbosch University***
#
# ***2021***
#
# Material adapted from: https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/generative/autoencoder.ipynb
#
# Code licensed under https://www.apache.org/licenses/LICENSE-2.0
#
#
#
# + [markdown] id="BvHfszboNuKI"
# ## <font color="green"> Learning outcomes:
#
# * Implement the autoencoder using CNNs.
#
# * The ```Conv2DTranspose``` layer
#
# ## <font color="green">Data information:
#
# * Features: (28x28) image
#
# * Output: (28x28) image
#
# ## <font color="green">Tasks for participants (boolean)?
#
# * No, follow along and make your own modifications and make sure you understand
#
# + [markdown] id="e1_Y75QXJS6h"
# ## Import TensorFlow and other libraries
# + id="YfIk2es3hJEd"
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras import backend as K
import tensorflow as tf
from sklearn.metrics import accuracy_score, precision_score, recall_score
from sklearn.model_selection import train_test_split
from tensorflow.keras import layers, losses
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Model
# + [markdown] id="sRD3m-MYT6qJ"
# ### Download the data
#
# Again this is an unsupervised problem so we don't need the labels
# + id="xqGTf1Q9i84Q"
(x_train, _), (x_test, _) = mnist.load_data()
# + [markdown] id="Y0J_uY7lT_X2"
# ### Normalise the data
#
# Divide by the maximum pixel value, 255.
# + id="b1cW0HfYi86v"
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
# + colab={"base_uri": "https://localhost:8080/"} id="_ZCVNc46UKB8" outputId="719ad0e3-937a-4999-fb69-9808bfd0167c"
x_train.shape
# + [markdown] id="rv1JLVAZUEhV"
# ### Reshape the data so that it has a 1 to indicate the depth of each image
# + id="YsG2DF21niEv"
x_train = x_train[..., tf.newaxis]
x_test = x_test[..., tf.newaxis]
# + colab={"base_uri": "https://localhost:8080/"} id="l4RIFtGxnY0D" outputId="426dedb5-67f6-41cc-e61c-5adee0460b70"
x_train.shape
# + [markdown] id="8dPOSQwgUdmc"
# Instead of working on the class directly like in the last prac, let's build the encoder and decoded using the functional API separately to make sure we get the shapes correct.
# + [markdown] id="ZjdkpnvbUOmb"
# ### Encoder
#
# Here we want to input an image of (28,28,1) and output a shape of (16) which is the encoded image.
#
# So we need to build an appropriate architecture which will produce that output.
#
#
# + id="OQx7glP75J_r"
# Here we use the Functional API which requires 3 things (Input, model and output)
# Specify the input layer here a shape of 28x28x1 (greyscale images)
encoder_input = tf.keras.Input(shape=(28, 28, 1))
# First conv layer which will reduce the shape by half given a stride of 2
encoder = tf.keras.layers.Conv2D(16, (3, 3), activation='relu', padding='same', strides=2)(encoder_input)
# Second conv layer which will reduce the shape by half given a stride of 2
encoder = tf.keras.layers.Conv2D(8, (3, 3), activation='relu', padding='same', strides=2)(encoder)
# Get the size of the current tensor - this will help us re-create this size when we create the decoder
# This will make more sense below.
tensor_shape = K.int_shape(encoder)
# Flatten
encoder = tf.keras.layers.Flatten()(encoder)
# Output layer has units based on latent dimension - here 16.
encoder_output = tf.keras.layers.Dense(16, activation = 'linear')(encoder)
# + id="hboDnAaW6tu8"
# Create a Model and specify the input and output
# This is normal Keras functional API like in the ANN prac
model = tf.keras.Model(encoder_input, encoder_output)
# + colab={"base_uri": "https://localhost:8080/"} id="hJ6pfo4j6z4N" outputId="43d4f03f-5b02-4994-b556-21f5fbf91b41"
model.summary()
# + [markdown] id="sN2W0JYXVK7A"
# here we see that we saved the volume as (7,7,8). So when we create the decoder, essentially we will go from 16 to 392 to (7,7,8). By saving this we can obtain two things: (1) the shape of 392 (7*7*8) and (2) the shape (7,7,8). We can do so without manually knowing these value and having to manually input them. See below from the two cells.
# + colab={"base_uri": "https://localhost:8080/"} id="hEHT9QR562Xk" outputId="a629fee7-dda6-4e23-d73d-78c2ff18b682"
tensor_shape
# + colab={"base_uri": "https://localhost:8080/"} id="OMkNou5vV3DF" outputId="2aee651e-4fd8-4aa3-ff2c-31b25ab4d720"
tensor_shape
# + id="kVcgKBz07y_h" colab={"base_uri": "https://localhost:8080/"} outputId="f5293888-bd83-46cf-f7de-6ab7e79d385e"
np.prod(tensor_shape[1:])
# + [markdown] id="XInYfe46Votj"
# ### Decoder
#
# Here we want to input a shape of (16) and output an image of (28,28,1) which is the decoded image.
#
# So we need to build an appropriate architecture which will produce that output.
#
# The decoder is essentially the inverse of the encoder.
# + id="TgmBipx15hrh"
# Here we use the Functional API which requires 3 things (Input, model and output)
# Specify the input layer here a shape of (16) (encoded images)
decoder_input = tf.keras.Input(shape=(16,))
# Now we create our first dense layer which will go from 16 -> 392. But we don't need to manually
# enter the value of 392 since we have a way to automatically calculate this (see above)
decoder = tf.keras.layers.Dense(np.prod(tensor_shape[1:]), activation = 'linear')(decoder_input)
# If we look at the encoder, the step before 392 unit layer was a tensor of shape (7,7,8). Again
# we don't have to manually enter these values.
decoder = tf.keras.layers.Reshape((tensor_shape[1], tensor_shape[2], tensor_shape[3]))(decoder)
# Working backwards, we can to the opposite of Conv2D which is Transpose convolution.
# API https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2DTranspose
decoder = layers.Conv2DTranspose(8, kernel_size=3, strides=2, activation='relu', padding='same')(decoder)
decoder = layers.Conv2DTranspose(16, kernel_size=3, strides=2, activation='relu', padding='same')(decoder)
# At this point our shape is (28,28,16) but we want (28,28,1) so we can remove the 16 features by
# applying a convolutional layer with 1 filter.
decoder_output = layers.Conv2D(1, kernel_size=(3, 3), activation='sigmoid', padding='same')(decoder)
# + id="TDiWLOig8HGN"
model = tf.keras.Model(decoder_input, decoder_output)
# + colab={"base_uri": "https://localhost:8080/"} id="jFyuGmSV5ht0" outputId="1288e7ff-41cb-4747-a3d1-f677e7bf55da"
model.summary()
# + [markdown] id="89MasJ8UlzLx"
# Now, let's take everything above and create our own Model object.
# + id="ef49wkt_i89Y"
# Define the latent dimension
latent_dim = 16
class Autoencoder(Model):
def __init__(self, latent_dim, name="autoencoder"):
super(Autoencoder, self).__init__()
# The latent dimension is a property of an autoencoder
self.latent_dim = latent_dim
# The Autoencoder has an encoder which we can make using normal layers
# Here we use the Functional API which requires 3 things (Input, model and output)
# ------------------------------------------
# Specify the input layer here a shape of 28x28x1 (greyscale images)
self.encoder_input = tf.keras.Input(shape=(28, 28, 1))
self.encoder = tf.keras.layers.Conv2D(16, (3, 3), activation='relu', padding='same', strides=2)(self.encoder_input)
self.encoder = tf.keras.layers.Conv2D(8, (3, 3), activation='relu', padding='same', strides=2)(self.encoder)
# Get this size
volumeSize = K.int_shape(self.encoder)
# Flatten
self.encoder = tf.keras.layers.Flatten()(self.encoder)
# Output
self.encoder_output = tf.keras.layers.Dense(latent_dim, activation = 'linear')(self.encoder)
self.encoder_model = tf.keras.Model(self.encoder_input, self.encoder_output)
# ------------------------------------------
# The Autoencoder has an decoder which we can make using normal layers
# Here we use the Functional API which requires 3 things (Input, model and output)
# ------------------------------------------
self.decoder_input = tf.keras.Input(shape=(latent_dim,))
self.decoder = tf.keras.layers.Dense(np.prod(tensor_shape[1:]), activation = 'linear')(self.decoder_input)
self.decoder = tf.keras.layers.Reshape((tensor_shape[1], tensor_shape[2], tensor_shape[3]))(self.decoder)
self.decoder = layers.Conv2DTranspose(8, kernel_size=3, strides=2, activation='relu', padding='same')(self.decoder)
self.decoder = layers.Conv2DTranspose(16, kernel_size=3, strides=2, activation='relu', padding='same')(self.decoder)
self.decoder_output = layers.Conv2D(1, kernel_size=(3, 3), activation='sigmoid', padding='same')(self.decoder)
self.decoder_model = tf.keras.Model(self.decoder_input, self.decoder_output)
# ------------------------------------------
# The forward pass
def call(self, x):
# Encode the inputs x using the propery defined above
# In the previous notebook we created the encoder and decoder as Sequential Models
# but now since we used the Functional API we need to call the Model objects we created above
encoded = self.encoder_model(x)
# Decode the encoded vector using the property defined above
decoded = self.decoder_model(encoded)
# Return the decoded (28,28) data
return decoded
autoencoder = Autoencoder(latent_dim)
# + id="MDsPp1USi8_-"
autoencoder.compile(optimizer='adam', loss=losses.MeanSquaredError())
# + colab={"base_uri": "https://localhost:8080/"} id="QLUPvT_Ii9Ck" outputId="3b59007b-f6ba-46a3-8e4f-6b374c5cbebc"
autoencoder.fit(x_train, x_train,
epochs=5,
shuffle=True,
validation_data=(x_test, x_test))
# + colab={"base_uri": "https://localhost:8080/"} id="4-MFPKXei9FL" outputId="3eeb3be3-568f-42af-9a22-ff09eb62bb76"
autoencoder.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="qV8kHSieonBg" outputId="9542896c-cf26-4e4f-f3e0-82c6680bc2fd"
x_test[0,:,:,0].shape
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="w8uZpo6Hi9Hz" outputId="4e830c13-980d-4a21-f3f9-7016eeae1782"
plt.imshow(x_test[0,:,:,0])
plt.gray()
# + id="zl8weMhci9Ku"
encoded_img = autoencoder.encoder_model(np.expand_dims(x_test[0],0)).numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="GOQOuPRri9Mu" outputId="57172373-da42-4099-d8e8-f4b6e848751c"
encoded_img
# + colab={"base_uri": "https://localhost:8080/"} id="YpyOM9t1o0hb" outputId="487073e2-e891-4f93-f1ab-502ba2c88ff5"
encoded_img.shape
# + id="Pm6sVs2_o4Hk"
decoded_img = autoencoder.decoder_model(encoded_img).numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="7dPKImAWo4KP" outputId="7833c68c-a064-402f-ba54-965a363072bd"
decoded_img.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="VAs8Xt5Ko4Rk" outputId="df534e7a-8394-408a-8e2e-c1babddc190d"
plt.imshow(decoded_img[0,:,:,0])
plt.gray()
# + [markdown] id="8BWCx-rOPWWl"
# ## Now let's compare the original images to the recontructed images on all of the testing data
# + id="lfZF_P7kPeyT"
encoded_imgs = autoencoder.encoder_model(x_test).numpy()
decoded_imgs = autoencoder.decoder_model(encoded_imgs).numpy()
# + id="s4LlDOS6FUA1" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="b56c444b-14e8-4ec3-cb09-8f929ce60b3a"
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test[i,:,:,0])
plt.title("original")
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i,:,:,0])
plt.title("reconstructed")
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# + [markdown] id="mll7JUsSeMrI"
# ## Task:
#
# * Explore modifications to the encoder and decoder (i.e. different conv layers in the encoder and decover).
#
# * Try different values for the latent dimensions
#
#
# + id="sVRPQaTQeY4f"
|
SIAM2021_Autoencoder_2_.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
"""
@author: <NAME>
Synergy of optical and synthetic aperture radar (SAR) data for early-stage crop yield estimation: A case study over a state of Germany
Methodology:
Step 1: Baseline Random Forest Regression Model
In this notebook we are importing ground truth yield data set along with Optical and SAR image features. 70% of the data is being used for taring and 30% for testing the model.
First a baseline random forest regression model with tuned hyper-paramets is being developed with all the input image features.
Step2: The performance of baseline model is being evaluated by correlation coeffifient (r), Mean Absolute Error (MAE) and Root Mean Square Error (RMSE) of obsered yield and predicted yield
Step 3: An omtimised random forest regression with genetic algorithm (GA) based feature selection is being developed.
This GA feature selection algorithm selected the best set of input image features for yied estimation with random forest regression model.
Step 4: The performance of omtimised random forest regression with genetic algorithm (GA) based feature selection is being evaluated by correlation coeffifient (r), Mean Absolute Error (MAE) and Root Mean Square Error (RMSE) of obsered yield and predicted yield
Step 5: Visualisation of baseline and optimised model performance
"""
###################################################################### Step 1: Baseline Random Forest Regression Model ####################################
######################
###################################################### Data Preparation ##############
# Pandas is used for data manipulation
import pandas as pd
features=pd.read_csv('WinterRapeseed_SampleData.csv')#Reading the Winter Rapeseed Ground Truth data in csv Format
features.head(5) #Display first 5 rows
# Use numpy to convert to arrays
import numpy as np
labels = np.array(features['Yield'])# Labels are the Crop Yield values
features= features.drop('Yield', axis = 1)# Remove the Crop Yield from the features
feature_list = list(features.columns)# Saving feature names for later use
features = np.array(features)# Convert features to numpy array
############################################## Tarin, Test data Preparation #########################
# Using Skicit-learn to split data into training and testing sets
from sklearn.model_selection import train_test_split
train_features, test_features, train_labels, test_labels = train_test_split(features, labels, test_size = 0.30, random_state = 42)# Split the data into 70% training and 30% testing sets
###################################### Baseline Random Forest Model Development and Hyper-Parameters Tuning with GridSearchCV#######################
#Importing required libraries
import numpy as np
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import accuracy_score
rfr=RandomForestRegressor(random_state=42)
param_grid = {
'bootstrap': [True],
'max_depth': [int(x) for x in np.linspace(10,100,10,endpoint=True,dtype = int)],
'min_samples_leaf': [int(x) for x in np.linspace(2,10,9,endpoint=True,dtype = int)],
'min_samples_split':[int(x) for x in np.linspace(2,15,14,endpoint=True,dtype = int)],
'n_estimators': [int(x) for x in np.linspace(50,1200,24,endpoint=True,dtype = int)]
} #Grid Space for Hyper-parameters tuning
CV_rfc = GridSearchCV(estimator=rfr, param_grid=param_grid, cv= 10,scoring='neg_mean_absolute_error',n_jobs = -1, verbose = 2)
CV_rfc.fit(train_features,train_labels)# Grid Search with 10-fold Cross Validation for Hyper-parameter tuning
####################Display Best set of Hyper-parameter
print(CV_rfc.best_params_)
##########Save the baseline random forest regression model with the results of hyper-parametrs tuning (CV_rfc.best_params_)
BaseLineRFR=RandomForestRegressor(bootstrap=True,
max_depth=80,
min_samples_leaf=3,
min_samples_split=8,
n_estimators=200)
import joblib
joblib.dump(BaseLineRFR, "./BaseLineModelrandom_forest_Regression.joblib",compress=3)# Saving the Baseline model for future use
###################################################### Step2: The performance Evaluation of baseline Random Forest Regression Model #################################
BaseLineModel = joblib.load("./BaseLineModelrandom_forest_Regression.joblib")# load the baseline model
BaseLineModel.fit(train_features,train_labels)
Bpredicted_labels_train_features=BaseLineModel.predict(train_features)#Predicting yield with training dataset
Bpredicted_labels_test_features=BaseLineModel.predict(test_features)#Predicting yield with testing dataset
############################################# Baseline Random Forest Regression Yield Model Performance Evaluation
from scipy.stats import pearsonr
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
print("Correlation Coefficient (r) of Baseline Random Forest Regression Model on Training Data: ",pearsonr(train_labels,Bpredicted_labels_train_features))
print("Correlation Coefficient (r) of Baseline Random Forest Regression Model on Testing Data: ",pearsonr(test_labels,Bpredicted_labels_test_features))
print("MAE of Baseline Random Forest Regression Model on Training Data: ",mean_absolute_error(train_labels,Bpredicted_labels_train_features))###t/ha
print("MAE of Baseline Random Forest Regression Model on Testing Data: ",mean_absolute_error(test_labels,Bpredicted_labels_test_features))###t/ha
print("RMSE of Baseline Random Forest Regression Model on Training Data: ",np.sqrt(mean_squared_error(train_labels,Bpredicted_labels_train_features)))###t/ha
print("RMSE of Baseline Random Forest Regression Model on Testing Data: ",np.sqrt(mean_squared_error(test_labels,Bpredicted_labels_test_features)))###t/ha
###################################################################################################################################################################
############################## Step 3: Opmtimised random forest regression with genetic algorithm (GA) based feature selection ################
from sklearn import linear_model
from genetic_selection import GeneticSelectionCV
Featureselector = GeneticSelectionCV(BaseLineModel,
cv=5,
verbose=1,
scoring="neg_mean_absolute_error",
max_features=20,
n_population=20,
crossover_proba=0.05,
mutation_proba=0.001,
n_generations=50,
crossover_independent_proba=0.5,
mutation_independent_proba=0.05,
tournament_size=3,
n_gen_no_change=10,
caching=True,
n_jobs=-1)#Feature Selection with Genetic Algorithm with Baseline Random ForestRegressor estimator with stopping criteria as NEGATIVE MSE
FeatureselectorModel = Featureselector.fit(train_features,train_labels)#Fitting the training data into Opmtimised random forest regression with genetic algorithm (GA) based feature selection
######################### Visualise the results of feature Selection ################
featurename= list(features.columns.values)#List of Input Feature Names
df = pd.DataFrame((featurename,Featureselector.support_,Featureselector.generation_scores_))### Feature Selection Result
Transpose=df.T
Transpose.columns =['Feature','Support','Score']
Transpose.head()#Showing the Selected Features
import seaborn as sns
import matplotlib.pyplot as plt
sns.factorplot(x= 'Feature', y= 'Support', data= Transpose, kind='bar', legend='True')
plt.title('Feature Selection',fontsize=15)
plt.show()#Ploting the selected features based on the Featureselector.support_
############################## Step 4: Performance evaluation of omtimised random forest regression with genetic algorithm (GA) based feature selection ############
predicted_labels_train_features=FeatureselectorModel .predict(train_features)
predicted_labels_test_features=FeatureselectorModel .predict(test_features)
print("Correlation Coefficient (r) of Optimised Random Forest Regression Model on Training Data: ",pearsonr(train_labels,predicted_labels_train_features))
print("Correlation Coefficient (r) of Optimised Random Forest Regression Model on Testing Data: ",pearsonr(test_labels,predicted_labels_test_features))
print("MAE of Optimised Random Forest Regression Model on Training Data: ",mean_absolute_error(train_labels,predicted_labels_train_features))#t/ha
print("MAE of Optimised Random Forest Regression Model on Testing Data: ",mean_absolute_error(test_labels,predicted_labels_test_features))#t/ha
print("RMSE of Optimised Random Forest Regression Model on Training Data: ",np.sqrt(mean_squared_error(train_labels,predicted_labels_train_features)))#t/ha
print("RMSE of Optimised Random Forest Regression Model on Testning Data: ",np.sqrt(mean_squared_error(test_labels,predicted_labels_test_features)))#t/ha
################ Step 5: Visualisation of baseline and optimised model performance ############
fig = plt.figure(figsize=(15,5))
###################################### Baseline Model##########################################
ax1 = plt.subplot(1, 2, 1)
ax1.scatter(train_labels,Bpredicted_labels_train_features,s=10, c='b', marker="o", label="Training Dataset = %.0f"%(train_labels.shape))
ax1.scatter(test_labels,Bpredicted_labels_test_features,s=10, c='r', marker="o", label="Testing Dataset = %.0f"%(test_labels.shape))
plt.xlabel('Observed Yield (t/ha)',fontsize=15)
plt.ylabel('Predicted Yield(t/ha)',fontsize=15)
plt.title('Baseline Model',fontsize=15)
plt.legend(loc="lower right")
############################ Optimised Model ###############
ax1 = plt.subplot(1, 2, 2)
ax1.scatter(train_labels,predicted_labels_train_features,s=10, c='b', marker="o",label="Training Dataset = %.0f"%(train_labels.shape))
ax1.scatter(test_labels,predicted_labels_test_features,s=10, c='r', marker="o", label="Testing Dataset = %.0f"%(test_labels.shape))
plt.xlabel('Observed Yield (t/ha)',fontsize=15)
plt.ylabel('Predicted Yield(t/ha)',fontsize=15)
plt.title('Optimised Model',fontsize=15)
plt.legend(loc="lower right")
plt.show()
|
WinterRapeseed_YieldEstimationModel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd> ErrorBars Element</dd>
# <dt>Dependencies</dt> <dd>Matplotlib</dd>
# <dt>Backends</dt> <dd><a href='./ErrorBars.ipynb'>Matplotlib</a></dd> <dd><a href='../bokeh/ErrorBars.ipynb'>Bokeh</a></dd>
# </dl>
# </div>
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
# ``ErrorBars`` provide a visual indicator for the variability of the plotted data on a graph. They are usually applied on top of other plots such as scatter, curve or bar plots to indicate the variability in each sample.
#
# ``ErrorBars`` may be used to represent symmetric error or asymmetric error. An ``ErrorBars`` Element must have one key dimensions representing the samples along the x-axis and two or three value dimensions representing the value of the sample and positive and negative error values associated with that sample. See the [Tabular Datasets](../../../user_guide/08-Tabular_Datasets.ipynb) user guide for supported data formats, which include arrays, pandas dataframes and dictionaries of arrays.
# #### Symmetric error
#
# By default the ``ErrorBars`` Element accepts x- and y-coordinates along with a symmetric error value:
np.random.seed(7)
errors = [(0.1*i, np.sin(0.1*i), np.random.rand()/2) for i in np.linspace(0, 100, 11)]
hv.Curve(errors) * hv.ErrorBars(errors)
# #### Assymetric error
# ``ErrorBars`` is a set of x-/y-coordinates with associated error values. Error values may be either symmetric or asymmetric, and thus can be supplied as an Nx3 or Nx4 array (or any of the alternative constructors Chart Elements allow).
errors = [(0.1*i, np.sin(0.1*i), np.random.rand()/2, np.random.rand()/4) for i in np.linspace(0, 100, 11)]
hv.Curve(errors) * hv.ErrorBars(errors, vdims=['y', 'yerrneg', 'yerrpos'])
# For full documentation and the available style and plot options, use ``hv.help(hv.ErrorBars).``
|
examples/reference/elements/matplotlib/ErrorBars.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"></ul></div>
# Everything:
# +
from fastai.vision.all import *
# paths
path = Path('/home/rory/data/coco2017')
train_im_dir, valid_im_dir = 'train2017', 'val2017'
train_json = 'annotations/instances_train2017.json'
valid_json = 'annotations/instances_val2017.json'
# get items & annos
def get_annos(path, anno_file, im_folder):
xs, ys = get_annotations(path/anno_file)
return L(xs).map(lambda x: path/im_folder/x), ys
train_paths, train_annos = get_annos(path, train_json, train_im_dir)
valid_paths, valid_annos = get_annos(path, valid_json, valid_im_dir)
paths = train_paths + valid_paths
annos = train_annos + valid_annos
bboxes = [a[0] for a in annos]
lbls = [a[1] for a in annos]
# create df and pkl
df = pd.DataFrame({
"path": paths,
"lbl": lbls,
"bbox": bboxes,
"is_valid": [0]*len(train_files) + [1]*len(valid_files),
"n_items": L(len(l) for l in lbls)
})
df.to_pickle(path/'20201027_coco_df.pkl')
df.head(1)
# -
# Subset:
# +
from fastai.vision.all import *
# paths
path = Path('/home/rory/data/coco2017')
train_im_dir, valid_im_dir = 'train2017', 'val2017'
train_json = 'annotations/instances_train2017.json'
valid_json = 'annotations/instances_val2017.json'
# get items & annos
def get_annos(path, anno_file, im_folder):
xs, ys = get_annotations(path/anno_file)
return L(xs).map(lambda x: path/im_folder/x), ys
train_files, train_annos = get_annos(path, train_json, train_im_dir)
valid_files, valid_annos = get_annos(path, valid_json, valid_im_dir)
annos = train_annos + valid_annos
bboxes = L(L(b) for b,l in annos)
lbls = L(L(l) for b,l in annos)
# get label subset
def flatten(l):
def _recur(l,res):
for o in l:
if isinstance(o,list): _recur(o,res)
elif isinstance(o,L) : _recur(o,res)
else: res.append(o)
return res
return _recur(l, L())
lbls_flat = flatten(lbls)
def get_count(lbl): return len(lbls_flat.filter(lambda x: x==lbl))
lbls_sorted = sorted(lbls_flat.unique(), key=get_count, reverse=True)
lbl_cts = [(l, get_count(l)) for l in lbls_sorted]
lbl_ss = [l for l,c in lbl_cts if 5000<c<30000]
lbl_ss.remove('traffic light')
lbl_ss.remove('motorcycle')
lbl_ss.remove('bus')
# get subset items & annos
ss_idxs = [[o in lbl_ss for o in l] for l in lbls]
ss_lbls = [list(o[i]) for o,i in zip(lbls,ss_idxs)]
ss_bboxes = [list(o[i]) for o,i in zip(bboxes,ss_idxs)]
ss_obj = [len(l) for l in ss_lbls]
# create df
df = pd.DataFrame({
"path" : train_files + valid_files,
"lbl" : ss_lbls,
"bbox" : ss_bboxes,
"n_obj" : ss_obj
})
# remove rows w/ 4+ objects per im
# reduce value counts to a max of 2*min_vc
too_many = 8
rng = list(range(too_many))
cts = list(df['n_obj'].value_counts())[:too_many]
minv = min(cts)
zipd = list(zip(*[rng,cts]))
zipd[0] = (0, minv) # limit ims w/zero objs
df = pd.concat([df[df['n_obj']==i].sample(n=min(s,minv*2)) for i,s in zipd])
# create pkl
df.to_pickle(path/'20201029_coco_ss_df.pkl')
# -
# Create dls:
# +
from fastai.vision.all import *
### Params ###
im_size = 224
batch_size = 64
path = Path('/home/rory/data/coco2017')
valid_pct = .10
### Items ###
df = pd.read_pickle(path/'20201029_coco_ss_df.pkl')
# get items
def get_cols(df,cols): return [df[c].to_list() for c in cols]
paths, bboxes, lbls = get_cols(df, ['path', 'bbox', 'lbl'])
p2b = {p:b for p,b in zip(paths,bboxes)}
p2l = {p:l for p,l in zip(paths,lbls)}
def get_bbox(p): return p2b[p]
def get_lbl(p): return p2l[p]
### DataBlock & DataLoaders ###
db = DataBlock(
blocks=[ImageBlock, BBoxBlock, BBoxLblBlock],
get_y=[get_bbox, get_lbl],
splitter=RandomSplitter(valid_pct),
item_tfms=Resize(im_size, method='squish'),
batch_tfms=Normalize.from_stats(*imagenet_stats),
n_inp=1)
dls = db.dataloaders(paths)
# -
### Inspection (IMPORTANT) ###
print("Size of train data:",len(dls.train.items))
print("Size of valid data:",len(dls.valid.items))
for i,t in enumerate(dls.one_batch()):
print(f"batch[{i}]:",'\t',t.dtype,'\t',t.shape)
|
_20201106_pickle_bug_multilabel_floats.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Steps 4 and 5: Analysing Data and Visualising Results
# ---
# ---
# Firstly, we need to reload the cleaned list of word tokens we were using in the [previous notebook](4-cleaning-and-exploring.ipynb) that we saved in a file. (You don't need to understand what is happening here in detail to follow the rest of the notebook.)
# +
# Import a module that helps with filepaths
from pathlib import Path
# Create a filepath for the file
tokens_file = Path('data', 'CLEAN-2199-0.txt')
# Create an empty list to hold the tokens
tokens = []
# Open the text file and append all the words to a list of tokens
with open(tokens_file, encoding='utf-8') as file:
for token in file.read().split():
tokens.append(token)
tokens[0:20]
# -
# ---
# ---
# ## Step 4: Analysing Data with Frequency Analysis
# Let's take a moment to remember our research question:
#
# > What are the top 10 words used in Homer's Iliad in English translation?
#
# In order to answer this question we need to count the number of each unique word in the text. Then we can see which are the most popular, or frequent, 10 words. This metric is called a *frequency distribution*.
#
# ### English Stopwords
# Before we start, we need to take a moment to think about what sort of words we are actually interested in counting.
#
# We are not interested in common words in English that carry little meaning, such as "the", "a" and "its". These are called *stopwords*. There is no definitive list of stopwords, but most Python packages used for Natural Language Processing provide one as a starting point, and spaCy is no exception.
# +
# Import the spaCy standard stopwords list
from spacy.lang.en.stop_words import STOP_WORDS
stopwords = [stop for stop in STOP_WORDS]
# Sort the stopwords in alphabetical order to make them easier to inspect
sorted(stopwords)
# +
# Write code here to count the number of stopwords
# -
# > **Exercise**: What do you notice about these stopwords?
#
# For your own research you will need to consider which stopwords are most appropriate:
# * Will standard stopword lists for modern languages be suitable for that language written 10, 50, 200 years ago?
# * Are there special stopwords specific to the topic or style of literature?
# * How might you find or create your own stopword list?
#
# Now we can filter out the stopwords that match this list:
tokens_nostops = [token for token in tokens if token not in stopwords]
tokens_nostops
# ### Creating a Frequency Distribution
# At last, we are ready to create a frequency distribution by counting the frequency of each unique word in the text.
#
# First, we create a frequency distribution:
# +
# Import a module that helps with counting
from collections import Counter
# Count the frequency of words
word_freq = Counter(tokens_nostops)
word_freq
# -
# This `Counter` maps each word to the number of times it appears in the text, e.g. `'coward': 17`. By scrolling down the list you can inspect what look like common and infrequent words.
#
# Now we can get precisely the 10 most common words using the function `most_common()`:
common_words = word_freq.most_common(10)
common_words
# > **Exercise**: Investigate what is further down the list of top words.
# ---
# ---
# ## Step 5: Presenting Results of the Analysis Visually
# There are many options for displaying simple charts, and very complex data, in Jupyter notebooks. We are going to use the most well-known library called [Matplotlib](https://matplotlib.org/), although it is perhaps not the easiest to use compared with some others.
#
# To create a Matplotlib plot we need to:
#
# * Import the matplotlib plot function
# * Arrange our data into a pair of lists: one for the x-axis, one for the y-axis
# * Set the appearance of titles, labels, ticks and gridlines
# * Pass the data into the plot function
#
# Let's display our results as a simple line plot:
# +
# Display the plot inline in the notebook with interactive controls
# %matplotlib notebook
# Import the matplotlib plot function
import matplotlib.pyplot as plt
# Get a list of the most common words
words = [word for word,_ in common_words]
# Get a list of the frequency counts for these words
freqs = [count for _,count in common_words]
# Set titles, labels, ticks and gridlines
plt.title("Top 10 Words used in Homer's Iliad in English translation")
plt.xlabel("Word")
plt.ylabel("Count")
plt.xticks(range(len(words)), [str(s) for s in words], rotation=90)
plt.grid(b=True, which='major', color='#333333', linestyle='--', alpha=0.2)
# Plot the frequency counts
plt.plot(freqs)
# Show the plot
plt.show()
# -
# With this interactive plot you can:
#
# * Resize the plot by dragging the bottom right-hand corner.
# * Pan across the plot to see values further to the right (if there are any to display).
# * Zoom into the plot.
#
# > **Exercise**: Change the code to explore different data and ways of displaying your data.
#
# There are also lots of other graphs that Matplotlib can create, and alternative plotting libraries to use instead, but these are beyond the scope of our course.
#
# ---
# ---
# ## Review and Reflection
# Now that you have seen the data and graph we have generated, no doubt you can see many ways we should improve.
#
# The process of text-mining a corpus (or individual text) is an iterative process. As you clean and explore the data, you will go back over your workflow again and again: from the collection stage, through to cleaning, analysis and presentation.
#
# > **Exercise**: List the ways you think we should improve the pipeline, from collection to plot.
#
# Fortunately, when you do your text-mining in code (and write explanatory text to document it) you know exactly what you did and can rerun and modify the process.
#
# ---
# ### Going Further: Libraries Libraries Libraries
#
# By now, you will be getting the idea that much of what you want to do in Python involves importing libraries to help you. Remember, libraries are _just code that someone else has written_.
#
# As reminder, here are some of the useful libraries we have used or mentioned in these notebooks:
# * [Requests](http://docs.python-requests.org/en/master/) - HTTP (web) requests library
# * [SpaCy](https://spacy.io/) - natural language processing library
# * [Matplotlib](https://matplotlib.org/) - 2D plotting library
#
# ---
# ---
# ---
# ## Summary
#
# Finally, we have:
#
# * **Loaded** clean token data from a file into a list.
# * Removed English **stopwords** from the list of tokens.
# * Created a **frequency distribution** and found the 10 most frequent words.
# * Visualised the frequency distribution in a **line plot**.
#
# ---
# ---
# ## What's Next?
# You will get the most out of this course if you can follow up on the learning over the next few days and weeks before you forget it all. This is particularly important when learning to code. Abstract concepts need to be reinforced little and often.
#
# ### Install Python on your Computer
#
# If you don't already have Python installed on your computer, perhaps the easiest way is with Anaconda:
#
# * [Installing Anaconda on Windows](https://www.datacamp.com/community/tutorials/installing-anaconda-windows)
# * [Installing Anaconda on Mac](https://www.datacamp.com/community/tutorials/installing-anaconda-mac-os-x)
#
# ### Running Jupyter Notebooks on your Computer
#
# Learn how to run and write Jupyter notebooks on your own computer (rather than using Binder): [Jupyter Notebook Tutorial: The Definitive Guide](https://www.datacamp.com/community/tutorials/tutorial-jupyter-notebook).
#
# ### Recommended books on Python
#
# * <NAME>. 2019. _Automate the Boring Stuff_ (2nd ed.) San Francisco: No Starch Press. [Available online](https://automatetheboringstuff.com/)
# * <NAME>. & <NAME>., 2016. _Data Wrangling with Python_. Sebastopol: O'Reilly Media.
#
# ### Text-mining and NLP in General
#
# * Work through this series of [Programming Historian tutorials](https://programminghistorian.org/en/lessons/working-with-text-files) to get some more practice with basic text files and basic text-mining techniques.
# * Follow a more in-depth set of Jupyter notebooks with [The Art of Literary Text Analysis](https://github.com/sgsinclair/alta/blob/master/ipynb/ArtOfLiteraryTextAnalysis.ipynb).
# * Read a practical and well-explained approach to Natural Language Processing (NLP) in Python: <NAME>., 2018. _Natural Language Processing and Computational Linguistics : A practical guide to text analysis with Python, Gensim, spaCy, and Keras._ Birmingham: Packt Publishing. [Available online](https://idiscover.lib.cam.ac.uk/primo-explore/fulldisplay?docid=44CAM_NPLD_MARC018975982&context=L&vid=44CAM_PROD&search_scope=SCOP_CAM_ALL&tab=cam_lib_coll&lang=en_US). This book has chapters on text pre-processing steps, various NLP techniques, and comes with Jupyter notebooks to follow.
#
# ### Python for Digital Humanities
#
# * Work through Chapters 1 - 4 (online Jupyter notebooks) of [Python Programming for the Humanities](http://www.karsdorp.io/python-course/).
# * Browse a big list of resources for [Teaching Yourself to Code in DH](http://scottbot.net/teaching-yourself-to-code-in-dh/).
|
5-analysis-and-visualisation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The .set_index() and .reset_index() Methods
import pandas as pd
bond_location = "/Volumes/GoogleDrive/My Drive/LIUZHICHENG/Udemy/Pandas/Data Source/pandas/jamesbond.csv"
bond = pd.read_csv(filepath_or_buffer=bond_location)
bond.head()
# set_index() for the "Film"
bond.set_index(keys=["Film"], inplace=True)
bond.head()
# set_index() to the "Year"
bond.set_index(keys=["Year"])
bond.head()
# reset_index() resets data into the original one
bond.reset_index()
bond.reset_index(drop=False)
bond.set_index(keys="Year", inplace=True)
bond.head()
# "Year" will be removed if drop=True
bond.reset_index(drop=True, inplace=False)
# reset the original one and "Year" comes back if previous inplace=False
bond.reset_index(drop=False, inplace=True)
bond.head()
# # Retrieve Rows by Index Label with .loc[] Accessor
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(inplace=True)
bond.head()
# +
# loc[] is the label location
# -
bond.loc["A View to a Kill"]
bond.loc["Goldfinger"]
# return a Series
type(bond.loc["Goldfinger"])
# the index lable must exist
bond.loc["HaHa"]
# index can be duplicate
bond.loc["Casino Royale"]
# return multiple rows
bond.loc[["<NAME>", "Goldfinger"]]
# in the list up to but not included
# in the dataframe it is inclusive
bond.loc["Diamonds Are Forever" : "From Russia with Love" ]
bond.loc["Diamonds Are Forever" : "From Russia with Love" : 2]
bond.loc["GoldenEye":]
bond.loc["<NAME>"::3, ["Actor", "Budget"]]
bond.loc[:"<NAME>"]
# check
"Gold Bond" in bond.index
"Dr. No" in bond.index
# # Retrieve Rows by Index Position with .iloc[] Accessor
bond = pd.read_csv(filepath_or_buffer=bond_location)
bond.head()
# +
# bond.iloc[, ] only accepts number
# -
bond.iloc[0]
bond.iloc[[15, 20]]
bond.iloc[[1, 20, 5], [5, 3, 1]]
# the index is out of the range
bond.loc[100]
# up to but not inclusive
# different from loc[]
bond.iloc[10:20]
bond.iloc[20::2]
bond[:10:4]
bond.iloc[[10, 7, 2]]
# # Second Argument to .loc[] and .iloc[] Accessors
# # the first one is index(row) and the second one is column
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col="Film")
bond.sort_index(ascending=True, inplace=True)
bond.head()
bond.loc[["Moonraker", "Dr. No"]]
# index then the column
bond.loc["Moonraker", "Actor"]
bond.loc["Dr. No", "Director"]
bond.loc["<NAME>", ["Year", "Box Office"]]
bond.loc[["Moonraker", "A View to a Kill"], ["Director", "Box Office"]]
bond.loc["Moonraker", "Director":"Budget"]
bond.loc["Moonraker":"Thunderball", "Director":"Budget"]
bond.loc["Moonraker":, :"Budget"]
bond.iloc[1, [2, 3]]
bond.iloc[10:, :5]
bond.iloc[:7, [2, 4]]
bond.iloc[1:7, [5, 3]]
# # Set New Value for a Specific Cell
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col="Film")
bond.sort_index(ascending=True, inplace=True)
bond.head()
# replace the value
bond.loc["Dr. No", "Actor"] = "<NAME> Connery"
bond.loc["Dr. No", "Actor"]
# replace the value
bond.loc["Dr. No", ["Box Office", "Budget", "Bond Actor Salary"]] = [7777, 7777, 7777]
bond.loc["Dr. No", ["Box Office", "Budget", "Bond Actor Salary"]]
# # Set Multiple Values in DataFrame
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col="Film")
bond.sort_index(ascending=True, inplace=True)
bond.head()
mask1 = bond["Actor"] == "Sean Connery"
bond[mask1]
# it doesn't modify the original one
# it changed the copy one
bond[mask1]["Actor"] = "Sir Sean Connery"
# it doesn't modify the original one
# it changed the copy one
bond[mask1].loc[:,"Actor"] = "Sir Sean Connery"
# it doesn't modify the original one
# it changed the copy one
bond.loc[mask1]["Actor"] = "Sir Sean Connery"
# below is correct and it returns a subset
bond.loc[mask1, "Actor"] = "Sir Sean Connery"
bond
# # .replace() also works to set multiple values
bond["Actor"].replace(to_replace="Sean Connery", value="<NAME>", inplace=True)
# # Rename Index Labels or Columns in a DataFrame
# # .rename()
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col="Film")
bond.sort_index(ascending=True, inplace=True)
bond.head()
# +
bond.rename(mapper={"GoldenEye":"Golden Eye",
"The World Is Not Enough":"Beset Bond Movie Ever"},
axis=0, inplace=False)
bond.rename(mapper={"GoldenEye":"Golden Eye",
"The World Is Not Enough":"Beset Bond Movie Ever"},
axis="rows", inplace=False)
bond.rename(mapper={"GoldenEye":"Golden Eye",
"The World Is Not Enough":"Beset Bond Movie Ever"},
axis="index", inplace=False)
bond.rename(index={"GoldenEye":"Golden Eye",
"The World Is Not Enough":"Beset Bond Movie Ever"}, inplace=False)
# +
bond.rename(mapper={"Year":"Release Date",
"Box Office":"Revenue"}, axis=1, inplace=False)
bond.rename(mapper={"Year":"Release Date",
"Box Office":"Revenue"}, axis="columns", inplace=False)
bond.rename(columns={"Year":"Release Date",
"Box Office":"Revenue"}, inplace=False)
# -
# if you want to replace all index or all columns
bond.columns = ["Release Year", "Cast", "Head", "Revenue", "Cost", "Bond Compensation"]
bond.columns
bond.index
# # Delete Rows or Columns from a DataFrame
# # .drop(), .pop() and del
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
# +
# drop a row
bond.drop(labels=["A View to a Kill"], axis=0, inplace=False)
bond.drop(labels=["A View to a Kill"], axis="rows", inplace=False)
bond.drop(labels=["A View to a Kill"], axis="index", inplace=False)
bond.drop(index=["A View to a Kill"], inplace=False)
# -
# you are drop a subset dataframe not the original one
bond[["Actor", "Director"]].drop(labels="Casino Royale", axis="index", inplace=True)
# +
# drop multiple columns
bond.drop(labels=["Director", "Budget"], axis=1, inplace=False)
bond.drop(labels=["Director", "Budget"], axis="columns", inplace=False)
bond.drop(columns=["Director", "Budget"], inplace=False)
# +
# .pop() will permanent remove Column (NO ROW) and return them
# .pop() only accepts one argument inside the brackets
# -
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
bond.pop("Actor")
# actor column has been removed
bond.head()
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
# it doesn't work for row (index)
bond.pop(["Casino Royale", "Die Another Day"])
# +
# del will permanent remove Column (NO ROW) and NO return them
# -
# del
del bond["Year"]
bond.head()
# remove whole DataFrame, bond
del bond
bond
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
# doesn't work for a row
del bond["A View to a Kill"]
# # Create A Random Sample using .sample()
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
# +
# random return the dataframe with no order
# +
# return 5 rows
bond.sample(n=5, axis=0)
bond.sample(n=5, axis="rows")
bond.sample(n=5, axis="index")
# -
# return the fraction of the rows, 25% of the rows will return
# by default, axis=None means 1, row or index
bond.sample(frac=0.25)
# +
# return the number of columns
bond.sample(n=1, axis=1)
bond.sample(n=1, axis="columns")
# -
bond.sample(frac=0.222, axis=1)
# # The .nsmallest() and .nlargest() Methods
# # It is available to the numeric columns
# # Similar to .sort_values()
# return the first three rows with largest Box Office
bond.sort_values(by=["Box Office"], ascending=False).head(3)
bond.nlargest(n=3, columns=["Box Office"])
# return the smallest budget rows
bond.sort_values(by=["Budget"], ascending=True, inplace=False).head(3)
bond.nsmallest(n=3, columns="Budget")
# it can work multiple column if having a tie
# keep parameter doesn't matter after experimenting
bond.nlargest(n=3, columns=["Box Office", "Bond Actor Salary"])
# you don't have to mention info for the columns if doing with Series
bond["Box Office"].nlargest(n=2)
bond["Year"].nsmallest(n=2)
# # Filtering with the .where() Method
# # However, .where() will retrieve the whole dataframe
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
mask1 = bond["Actor"] == "<NAME>"
bond[mask1]
# return the entire DataFrame
# cond -> condition
bond.where(cond=mask1)
bond.where(cond=bond["Box Office"] > 800, inplace=False)
mask2 = bond["Box Office"] > 800
bond[mask1 & mask2]
bond.where(cond = mask1 & mask2)
# # The .query() Method and it only works columns without spaces
# # SQL
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
[column_name for column_name in bond.columns]
[index_name for index_name in bond.index]
"ha ha".replace(" ", " cao ni ma ")
# replace " " into "_" so that it works
bond.columns = [column_name.replace(" ", "_") for column_name in bond.columns]
bond.head(1)
bond.query(expr="Actor == '<NAME>'")
bond.query(expr='Director != "<NAME>"')
bond.query("Actor == '<NAME>' and Director == '<NAME>'")
bond.query("Budget >= 150 or Year > 2015")
bond.query("Actor in ['<NAME>', '<NAME>']")
bond.query("Director not in ['<NAME>', '<NAME>', '<NAME>']")
# # A Review of the .apply() Method on Columns
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
bond.columns = [column_name.replace(" ", "_") for column_name in bond.columns]
bond.head(1)
bond["Year"].apply(func=lambda x:x*2)
def conver_to_string_and_add_millions(number):
return str(number) + " MILLIONS!"
bond["Box_Office"].apply(func=conver_to_string_and_add_millions)
# bond.loc[:, []]
bond.loc[:,"Budget"].apply(func=conver_to_string_and_add_millions)
columns = ["Box_Office", "Budget", "Bond_Actor_Salary"]
for column in columns:
bond[column] = bond[column].apply(conver_to_string_and_add_millions)
bond.head()
# # The .apply() Method with Row Values
bond.sort_index(ascending=True, inplace=True)
bond.head()
def good_movie(row):
actor = row[1]
budget = row[4]
if actor == "<NAME>":
return "The best"
elif actor == "<NAME>" and budget > 40:
return "Enjoyable"
else:
return "I have not clue"
# axis = 1 axis = "columns"
# a little bit confusing since using column not the row
bond.apply(func=good_movie, axis=1)
# # The .copy() Method and instead of impacting the original dataframe
bond.set_index(keys="Film", inplace=True)
bond.sort_index(ascending=True, inplace=True)
bond.head()
directors = bond["Director"]
directors.head()
# you are replacing values into the copy dataframe
directors["A View to a Kill"] = "<NAME>"
bond.head()
directors.head()
bond = pd.read_csv(filepath_or_buffer=bond_location, index_col=["Film"])
bond.sort_index(ascending=True, inplace=True)
bond.head()
# get an absolutely copy dataframe
directors = bond["Director"].copy()
directors.head()
directors["A View to a Kill"] = "<NAME>"
directors.head(3)
bond.head(3)
|
Section 6.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Computer Vision: CNN for CIFAR Dataset**
# +
import torch
import torch.nn as nn
import matplotlib.pyplot as plt
from torch.autograd import Variable
# Torchvision module contains various utilities, classes, models and datasets
# used towards computer vision usecases
from torchvision import datasets
from torchvision import transforms
import torchvision
# Functional module contains helper functions
import torch.nn.functional as F
# -
# **Create the DataLoaders**
# +
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
batch_size = 4
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=batch_size,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# -
# Resources that helped me understand CNN: <br>
# https://towardsdatascience.com/intuitively-understanding-convolutions-for-deep-learning-1f6f42faee1 <br>
# https://cs231n.github.io/convolutional-networks/#layers
# **Create the model**
# The Conv2d layer transforms a 3-channel image, and the MaxPool2d layer halves the height and width. The feature map gets smaller as we add more layers, until we are finally left with a small feature map, which can be flattened into a vector. We can then add some fully connected layers at the end to get vector of size 10 for each image.
# 
class CIFARNet(nn.Module):
def __init__(self):
super().__init__()
# Convolution Layer 1
self.conv1 = nn.Conv2d(3, 5, kernel_size=5)
self.pool = nn.MaxPool2d(2, 2)
# Convolution Layer 2
self.conv2 = nn.Conv2d(5, 20, kernel_size=5)
# Fully connected layers
self.fc1 = nn.Linear(20 * 5 * 5, 100)
self.fc2 = nn.Linear(100, 50)
self.fc3 = nn.Linear(50, 10)
def forward(self, x):
# Convolution Layer 1
x = self.conv1(x)
x = F.relu(x)
x = self.pool(x)
# Convolution Layer 2
x = self.conv2(x)
x = F.relu(x)
x = self.pool(x)
x = torch.flatten(x, 1) # flatten all dimensions except batch
# Fully connected layer 1
x = self.fc1(x)
x = F.relu(x)
# Fully connected layer 2
x = self.fc2(x)
x = F.relu(x)
# Fully connected layer 3
x = self.fc3(x)
return x
# **Create the objects**
# +
# The model
net = CIFARNet()
# Our loss function
criterion = nn.CrossEntropyLoss()
# Our optimizer
learning_rate = 0.001
optimizer = torch.optim.SGD(net.parameters(), lr=learning_rate, momentum=0.9)
# -
# **Train the Model**
# +
epoch_num = 2
train_loss = []
for epoch in range(epoch_num): # loop over the dataset multiple times
iter_loss = 0.0
iterations = 0
running_loss = 0.0
net.train() # Put the network into training mode
for i, data in enumerate(trainloader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
iter_loss += loss.data # Accumulate the loss
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
# Record the training loss
train_loss.append(running_loss/2000)
running_loss = 0.0
print('Finished Training')
# -
# **Calculate Accuracy of Entire Dataset**
# +
correct = 0
total = 0
# since we're not training, we don't need to calculate the gradients for our outputs
with torch.no_grad():
for data in testloader:
images, labels = data
# calculate outputs by running images through the network
outputs = net(images)
# the class with the highest energy is what we choose as prediction
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
# -
# **Calculate Accuracy of Each Class in Dataset**
# +
# prepare to count predictions for each class
correct_pred = {classname: 0 for classname in classes}
total_pred = {classname: 0 for classname in classes}
# again no gradients needed
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predictions = torch.max(outputs, 1)
# collect the correct predictions for each class
for label, prediction in zip(labels, predictions):
if label == prediction:
correct_pred[classes[label]] += 1
total_pred[classes[label]] += 1
# print accuracy for each class
for classname, correct_count in correct_pred.items():
accuracy = 100 * float(correct_count) / total_pred[classname]
print("Accuracy for class {:5s} is: {:.1f} %".format(classname,
accuracy))
# -
# **Visialize the loss**
# +
# Loss
f = plt.figure(figsize=(10, 8))
plt.plot(train_loss, label='training loss')
plt.legend()
plt.show()
# -
# **Standalone inference**
# +
import numpy as np
dataiter = iter(testloader)
images, labels = dataiter.next()
ind = 1
i = images[ind].unsqueeze(0)
#i = testset[image_index][0].resize_((1, 1, 224, 224))
label = labels[ind]
net.eval()
output = net(i)
# -
output.data
_, predicted = torch.max(output.data, 1)
print("Prediction is: ", classes[predicted])
print("Actual is is: ", classes[label])
|
cnnModel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
main_path = os.path.abspath(os.path.join(os.path.dirname("./quantity_search.ipynb"), '..'))
import sys
sys.path.insert(0, main_path)
import numpy as np
import pandas as pd
import seaborn as snb
import matplotlib.pyplot as plt
import json
plt.style.use('seaborn')
folder = os.path.join(main_path, "data", "quantity_search")
# +
def sort_quantities_by_acc(df):
quantities = np.unique(df["quantity"])
Q = []
acc = []
wl = []
win = []
for q in quantities:
df_i = df[df["quantity"] == q]
acc_i = df_i["cv_mean"].to_numpy()
wl_i = df_i["wl"].to_numpy()
win_i = df_i["win"].to_numpy()
idx = np.argmax(acc_i)
acc.append(float(acc_i[idx]))
wl.append(int(wl_i[idx]))
win.append(float(win_i[idx]))
Q.append(str(q))
return Q, acc, wl, win
def add_count_Qs(df):
df["count_Qs"] = [len(x.split("-")) for x in df["quantity"]]
return df
def comb_filter_only_bests(df, n_max):
df2 = df[df["count_Qs"] == 2]
cv_mean = df2["cv_mean"].to_numpy()
highest_idx = np.argmax(cv_mean)
line = df2.iloc[highest_idx]
df_filter = ((df["count_Qs"] == 2) & (df2["quantity"] == line.quantity))
for i in range(3,n_max):
df2 = df[df["count_Qs"] == i]
cv_mean = df2["cv_mean"].to_numpy()
highest_idx = np.argmax(cv_mean)
line = df2.iloc[highest_idx]
df_filter |= ((df["count_Qs"] == i) & (df2["quantity"] == line.quantity))
return df[df_filter]
def concat_single_best(df1, df2):
cv_mean = df1["cv_mean"].to_numpy()
highest_idx = np.argmax(cv_mean)
highest_Q = df1.iloc[highest_idx].quantity
df3 = df1[df1["quantity"] == highest_Q]
return pd.concat([df3, df2])
# -
# ## Single-Q results
filename = "single_quantity_lsa_20210903-044949"
df_single_q = pd.read_csv(os.path.join(folder, filename), index_col=None)
df_single_q = df_single_q[df_single_q["valid_cv"]]
add_count_Qs(df_single_q)
frames = [df_single_q[df_single_q["quantity"]=="(Tr)"],
df_single_q[df_single_q["quantity"]=="(Mm)"]]
df_resume = pd.concat(frames)
df_resume = df_resume[["quantity", "win", "wl", "cv_mean"]]
df_resume.head()
all_frames = [df_single_q]
# +
plt.figure(figsize=(16, 4))
b = snb.boxplot(x="quantity", y="cv_mean", data=df_single_q)
plt.ylabel("Cross-Validation accuracy", fontsize=16)
plt.xlabel("Statistical Quantity", fontsize=16)
plt.title("Single-quantity evaluation", fontsize=18)
plt.yticks(fontsize=14)
plt.xticks(fontsize=14)
plt.tight_layout()
plt.savefig("figs/single_q_evaluations.png", dpi=350)
# -
# ## Comb Single-Q results
filename = "comb_single_quantity_lsa_20210903-044949"
df_comb_single_q = pd.read_csv(os.path.join(folder, filename), index_col=None)
df_comb_single_q = df_comb_single_q[df_comb_single_q["valid_cv"]]
add_count_Qs(df_comb_single_q)
df_comb_single_q2 = comb_filter_only_bests(df_comb_single_q, 6)
df_comb_single_q2 = concat_single_best(df_single_q, df_comb_single_q2)
df_comb_single_q2.head()
frame = df_comb_single_q2[df_comb_single_q2["quantity"]=="(Tr-Mm)"][["quantity", "win", "wl", "cv_mean"]]
df_resume = pd.concat([df_resume, frame])
df_resume.head()
all_frames.append(df_comb_single_q2)
# +
plt.figure(figsize=(16, 4))
b = snb.boxplot(x="quantity", y="cv_mean", data=df_comb_single_q2)
loc, labels = plt.xticks()
# plt.xticks(loc, labels, rotation = 15, ha="right", fontsize=13)
plt.ylabel("Cross-Validation accuracy", fontsize=16)
plt.xlabel("Statistical Quantity", fontsize=16)
plt.title("Single-quantity combinations evaluation", fontsize=18)
plt.yticks(fontsize=14)
plt.xticks(fontsize=14)
plt.tight_layout()
plt.savefig("figs/single_q_comb_evaluations.png", dpi=350)
# -
# ## Double-Q results
# +
filename = "double_quantity_lsa_20210903-044949"
df_double_q = pd.read_csv(os.path.join(folder, filename), index_col=None)
df_double_q = df_double_q[df_double_q["valid_cv"]]
add_count_Qs(df_double_q)
df_double_q = df_double_q[df_double_q["quantity"] != "(MmTr)"]
# df_double_q.head()
frame = df_double_q[df_double_q["quantity"]=="(TrMm)"][["quantity", "win", "wl", "cv_mean"]]
df_resume = pd.concat([df_resume, frame])
all_frames.append(df_double_q)
# +
plt.figure(figsize=(16, 4))
b = snb.boxplot(x="quantity", y="cv_mean", data=df_double_q)
plt.ylabel("Cross-Validation accuracy", fontsize=16)
plt.xlabel("Statistical Quantity", fontsize=16)
plt.title("Double-quantity evaluation", fontsize=18)
plt.yticks(fontsize=14)
plt.xticks(fontsize=14)
plt.tight_layout()
plt.savefig("figs/double_q_evaluations.png", dpi=350)
# -
# ## Comb Double-Q results
# +
filename = "comb_double_quantity_lsa_20210904-024910"
df_comb_double_q = pd.read_csv(os.path.join(folder, filename), index_col=None)
df_comb_double_q = df_comb_double_q[df_comb_double_q["valid_cv"]]
add_count_Qs(df_comb_double_q)
df_comb_double_q2 = comb_filter_only_bests(df_comb_double_q, 6)
df_comb_double_q2 = concat_single_best(df_double_q, df_comb_double_q2)
# df_comb_double_q2.head()
frames = [df_resume,
df_comb_double_q2[df_comb_double_q2["quantity"]=="(MmTr-MmMn)"][["quantity", "win", "wl", "cv_mean"]],
df_comb_double_q2[df_comb_double_q2["quantity"]=="(MmTr-MmMn-MmMx)"][["quantity", "win", "wl", "cv_mean"]],
df_comb_double_q2[df_comb_double_q2["quantity"]=="(MmTr-MmMn-MmMx-TrMn)"][["quantity", "win", "wl", "cv_mean"]]]
df_resume = pd.concat(frames)
all_frames.append(df_comb_double_q2)
# +
plt.figure(figsize=(16, 4))
b = snb.boxplot(x="quantity", y="cv_mean", data=df_comb_double_q2)
plt.ylabel("Cross-Validation accuracy", fontsize=16)
plt.xlabel("Statistical Quantity", fontsize=16)
plt.title("Double-quantity combinations evaluation", fontsize=18)
plt.yticks(fontsize=13)
plt.xticks(fontsize=13)
plt.tight_layout()
plt.savefig("figs/double_q_comb_evaluations.png", dpi=350)
# -
# ## Triple-Q results
filename = "triple_quantity_lsa_20210904-024910"
df_triple_q = pd.read_csv(os.path.join(folder, filename), index_col=None)
df_triple_q = df_triple_q[df_triple_q["valid_cv"]]
add_count_Qs(df_triple_q)
df_triple_q = df_triple_q[df_triple_q["quantity"] != "(MmTr)"]
df_triple_q = df_triple_q.sort_values("cv_mean", ascending=False)
df_triple_q.head()
all_frames.append(df_triple_q)
# +
plt.figure(figsize=(16, 4))
b = snb.boxplot(x="quantity", y="cv_mean", data=df_triple_q)
plt.ylabel("Cross-Validation accuracy", fontsize=16)
plt.xlabel("Statistical Quantity", fontsize=16)
plt.title("Triple-quantity evaluation", fontsize=18)
plt.yticks(fontsize=14)
plt.xticks(fontsize=14)
plt.tight_layout()
# plt.savefig("figs/triple_q_evaluations.png", dpi=350)
# -
# ## Comb Triple-Q results
filename = "comb_triple_quantity_lsa_20210904-024910"
df_comb_triple_q = pd.read_csv(os.path.join(folder, filename), index_col=None)
df_comb_triple_q = df_comb_triple_q[df_comb_triple_q["valid_cv"]]
add_count_Qs(df_comb_triple_q)
df_comb_triple_q2 = comb_filter_only_bests(df_comb_triple_q, 5)
df_comb_triple_q2 = concat_single_best(df_triple_q, df_comb_triple_q2)
# df_comb_triple_q2.head()
frames = [df_resume,
df_comb_double_q2[df_comb_double_q2["quantity"]=="(MmTr-MmMn)"][["quantity", "win", "wl", "cv_mean"]],
df_comb_double_q2[df_comb_double_q2["quantity"]=="(MmTr-MmMn-MmMx)"][["quantity", "win", "wl", "cv_mean"]],
df_comb_double_q2[df_comb_double_q2["quantity"]=="(MmTr-MmMn-MmMx-TrMn)"][["quantity", "win", "wl", "cv_mean"]]]
df_resume = pd.concat(frames)
all_frames.append(df_comb_triple_q2)
# +
plt.figure(figsize=(16, 4))
b = snb.boxplot(x="quantity", y="cv_mean", data=df_comb_triple_q2)
plt.ylabel("Cross-Validation accuracy", fontsize=16)
plt.xlabel("Statistical Quantity", fontsize=16)
plt.title("Triple-quantity combinations evaluation", fontsize=18)
plt.yticks(fontsize=14)
plt.xticks(fontsize=14)
plt.tight_layout()
plt.savefig("figs/triple_q_comb_evaluations.png", dpi=350)
# -
# # RESUME
def resume_quantity_search(df1, df2, df3):
df1 = df1.reset_index()
df2 = df2.reset_index()
df3 = df3.reset_index()
qs1 = np.unique(df1["quantity"])
qs2 = np.unique(df2["quantity"])
qs3 = np.unique(df3["quantity"])
print(qs1)
idxs = []
for q in qs1:
df_tmp = df1[df1["quantity"] == q]
high_idx = np.argmax(df_tmp["cv_mean"])
idxs.append(high_idx)
print(idxs, len(df1))
df1_bests = df1.iloc[idxs]
df1_bests["type"] = np.array(["Single-Q"] * len(df1_bests))
idxs = []
for q in qs2:
high_idx = np.argmax(df2[df2["quantity"] == q]["cv_mean"])
idxs.append(high_idx)
df2_bests = df2.iloc[idxs]
df2_bests["type"] = np.array(["Double-Q"] * len(df2_bests))
idxs = []
for q in qs3:
high_idx = np.argmax(df3[df3["quantity"] == q]["cv_mean"])
idxs.append(high_idx)
df3_bests = df3.iloc[idxs]
df3_bests["type"] = np.array(["Triple-Q"] * len(df3_bests))
res_df = pd.concat([df1_bests, df2_bests, df3_bests])
return res_df[["quantity", "type", "cv_mean", "cv_std", "win", "wl", "alpha", "dropped", "bopf_shape", "cv_time"]]
df = resume_quantity_search(df_comb_single_q2, df_comb_double_q2, df_comb_triple_q2)
df = df.sort_values("cv_mean", ascending=False)
df
# +
plt.figure(figsize=(16, 4))
b = snb.boxplot(x="cv_mean", y="quantity", data=df_resume)
plt.xlabel("Cross-Validation accuracy", fontsize=16)
plt.ylabel("Statistical Quantity", fontsize=16)
plt.title("Multi-quantity search summary", fontsize=18)
plt.yticks(fontsize=14)
plt.xticks(fontsize=14)
plt.tight_layout()
plt.savefig("figs/multi-quantity-search-result.png", dpi=350)
# +
df_all = pd.concat(all_frames)
df_all = df_all.sort_values("cv_mean", ascending=False)
plt.figure(figsize=(14, 18))
b = snb.boxplot(x="cv_mean", y="quantity", data=df_all)
plt.xlabel("Cross-Validation accuracy", fontsize=16)
plt.ylabel("Statistical Quantity", fontsize=16)
plt.title("Multi-quantity search summary", fontsize=20)
plt.yticks(fontsize=15)
plt.xticks(fontsize=15)
plt.tight_layout()
plt.savefig("figs/multi-quantity-search-all-results.png", dpi=350)
# -
# # TIME COMPLEXITY BOPF METHOD
def time_complexity(dfs):
dfs_arr = []
for i, dfs_i in enumerate(dfs):
dfs_i["n_q"] = [i+1] * len(dfs_i)
dfs_arr.append(dfs_i)
df = pd.concat(dfs_arr)
df["features"] = [row.alpha ** (row.wl * row.n_q) for index, row in df.iterrows()]
df["x"] = [row.wl * row.n_q for index, row in df.iterrows()]
x = df["x"].to_numpy()
y = df["bopf_time"].to_numpy()
return x, y
x, y = time_complexity([df_single_q, df_double_q, df_triple_q])
plt.plot(x, y, ".")
plt.xlabel(r"$log_{\alpha}(N)$")
plt.ylabel("Time (sec.)")
|
notebooks/quantity_search.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
import random
# # Моделирование случайных процессов
#
# Нахождение числа $\pi$ методом Монте-Карло
# +
x = [] ; y = []
N = 100
for i in range(N):
x.append(random.random()); y.append(random.random())
plot(x,y,'ro')
# +
xr = [] ; yr = []
Nk = 0
for i in range(N):
if x[i] ** 2 + y[i] ** 2 <=1:
Nk = Nk + 1
xr.append(x[i]); yr.append(y[i])
4.0 * Nk / N # искомое пи
plot(x,y,'ob',xr,yr,'or')
# -
import random as rnd
# +
x = []
y = []
for i in range(180):
al=rnd.random()*2*math.pi
r=rnd.random()*10
x.append(r*cos(al))
y.append(r*sin(al))
plt.grid()
plt.plot(x,y,'+')
# +
x = []
y = []
for i in range(1000):
al=rnd.random()*2*math.pi
r=rnd.random()*10
x.append(r*cos(al))
y.append(r*sin(al))
plt.grid()
plt.plot(x,y,'+')
# +
x = []
y = []
for i in range(10000):
al=rnd.random()*2*math.pi
r=rnd.random()*10
x.append(r*cos(al))
y.append(r*sin(al))
plt.grid()
plt.plot(x,y,'+')
# +
x = []
y = []
for i in range(10000):
al=rnd.random()*0.5*math.pi
r=rnd.random()*10
x.append(r*cos(al))
y.append(r*sin(al))
plt.grid()
plt.plot(x,y,'+')
# -
plt.grid()
plt.xlim(-2,2)
plt.ylim(2,-2)
plt.plot(x,y,'+')
plt.grid()
plt.xlim(-5,15)
plt.ylim(15,-5)
plt.plot(x,y,'+')
plt.grid()
plt.xlim(-5,15)
plt.ylim(-5, 15)
plt.plot(x,y,'+')
|
visualisation_matplotlib/.ipynb_checkpoints/06_monte_carlo_rnd-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# #### Flow Control
#
# Python supports conditional logic and "branching" so "non-sequential" execution of commands are possible.
# There are two (2) types of iterations (or loops) available.
jack = ['work', 'work', 'work']
# #### While
#
# _while_ is discussed here.
dull = True
jackLength = len(jack)
jackLength
index = 0
# The "code block" is repeatedly executed until the terminitating condition becomes _True_. Generally, an iterator is "advanced" for it to end - otherwise, it will become an "infinite loop" and run indefinitely if it is not interrupted (i.e. either manually stopped by the programmer or until it uses up all the computing resources).
# _index_ (or _i_ for short) as the iterator since a loop is typically used to access the contents of an _array_ element
while (index < jackLength):
if (jack[index] == 'work'):
pass
else:
dull = False
index += 1
# _pass_ is equivelent to "no-op" where no action is taken - the _if_ could have been written the "opposite" way but would not have been able to illustrate this statement.
print('Index: ', index, "Dull: ", dull)
jack2 = ['play']
# + jupyter={"source_hidden": true}
jack += jack2
# -
jack
index2 = 0
jackLength2 = len(jack)
while (index2 < jackLength2):
if (jack[index2] != 'work'):
dull = False
index2 += 1
print('Index: ', index2, "Dull: ", dull)
# Note: _index_ was substituted with _index2_ and _jackLength_ with _jackLength2_ so that the loop would act as expected. Also. the _if_ statement was rewritten.
index2 = 0
while (index2 < jackLength2):
if (jack[index2] == 'work'):
print("Whistle! \n")
index2 += 1
current = 1
maximum = range(1, 4, 1)
while (current in maximum):
print('Current: ', current)
current += 1
# _range_ is used to define an iterator sequence. By default, it starts at 0 and increments by 1. The first parameter denotes the starting point, the next argument (and is the minimum required) specifies the final value minus the step (from what I know as other sites describe it differently), and the final value is the difference.
|
FlowControl/While.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Traffic Sign or Not Traffic Sign
# Reference: https://github.com/mzaradzki/neuralnets/tree/master/vgg_faces_keras
#
# Since we're brute-forcing some "art-sticker" to mislead the classifier, we may want to use AWS EC2 instance with GPU.
# Here's how to install Caffe: https://github.com/BVLC/caffe/wiki/Caffe-installing-script-for-ubuntu-16.04---support-Cuda-8
# +
import shutil
import requests
import tempfile
import os
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
pd.set_option('display.mpl_style', 'default')
# %matplotlib inline
caffe_root = '/opt/caffe/'
import sys
sys.path.insert(0, caffe_root + 'python')
import caffe
plt.rcParams['figure.figsize'] = (4, 4)
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# -
# Load the labels (so we know whether 242 means 'adorable puppy' or 'garbage can')
# n06794110 street sign
imagenet_labels_filename = caffe_root + '/data/ilsvrc12/synset_words.txt'
try:
labels = np.loadtxt(imagenet_labels_filename, str, delimiter='\t')
except:
# !/opt/caffe/data/ilsvrc12/get_ilsvrc_aux.sh
labels = np.loadtxt(imagenet_labels_filename, str, delimiter='\t')
# +
# Set Caffe to CPU mode because who even knows how to set up a GPU (not me)
caffe.set_mode_cpu()
# Load our model! trained by the GOOGLES! <3
def load_model():
BATCH_SIZE = 1
net = caffe.Net('/opt/caffe/models/bvlc_googlenet/deploy.prototxt',
'/models/bvlc_googlenet.caffemodel',
caffe.TEST)
# change batch size to 1 for faster processing
# this just means that we're only processing one image at a time instead of like 50
shape = list(net.blobs['data'].data.shape)
shape[0] = BATCH_SIZE
net.blobs['data'].reshape(*shape)
net.blobs['prob'].reshape(BATCH_SIZE, )
net.reshape()
return net
net = load_model()
# Caffe comes with a handy transformer pipeline so that
# we can make our images into the format it needs! Yay!
transformer = caffe.io.Transformer({'data': net.blobs['data'].data.shape})
transformer.set_transpose('data', (2,0,1))
transformer.set_raw_scale('data', 255) # the reference model operates on images in [0,255] range instead of [0,1]
transformer.set_channel_swap('data', (2,1,0)) # the reference model has channels in BGR order instead of RGB
# -
def get_png_image(url):
fd, path = tempfile.mkstemp()
tmpfile = os.fdopen(fd, "wb")
response = requests.get(url, stream=True)
shutil.copyfileobj(response.raw, tmpfile) # potential vulnerability in this function, but who cares?
if 'png' in url:
# !mv {path} {path}.png
else:
# !convert {path} {path}.png
path = path + ".png"
return transformer.preprocess('data', caffe.io.load_image(path))
art_sticker_05ft_data = transformer.preprocess('data', caffe.io.load_image('./experimental_attack_images/art-sticker-05ft_0deg.jpg'))
print art_sticker_05ft_data.shape
net.blobs['data'].data[...] = art_sticker_05ft_data
out = net.forward()
pred_class = out['prob'][0].argmax()
print("Predicted class is #{}.".format(pred_class))
plt.imshow(transformer.deprocess('data', net.blobs['data'].data[0]))
print labels[pred_class]
# So, we can't cheat ImageNet with art sticks that worked in Evtimov et al. (2017).
# Let's try something real. Image taken from https://www.upbeat.com/Images/blog/parkinglot1-09052013.jpg
# +
def display(data):
plt.imshow(transformer.deprocess('data', data))
def get_label_name(num):
options = labels[num].split(',')
# remove the tag
options[0] = ' '.join(options[0].split(' ')[1:])
return ','.join(options[:2])
def predict(data, n_preds=6, display_output=True):
net.blobs['data'].data[...] = data
if display_output:
display(data)
prob = net.forward()['prob']
probs = prob[0]
prediction = probs.argmax()
top_k = probs.argsort()[::-1]
for pred in top_k[:n_preds]:
percent = round(probs[pred] * 100, 2)
# display it compactly if we're displaying more than the top prediction
pred_formatted = "%03d" % pred
if n_preds == 1:
format_string = "label: {cls} ({label})\ncertainty: {certainty}%"
else:
format_string = "label: {cls} ({label}), certainty: {certainty}%"
if display_output:
print format_string.format(
cls=pred_formatted, label=get_label_name(pred), certainty=percent)
return prob
# -
# Convert PIL image to Caffe image
from PIL import Image
def conv_pil(im):
img = np.asarray(im).astype(np.float32) / 255.
if img.ndim == 2:
img = img[:, :, np.newaxis]
img = np.tile(img, (1, 1, 3))
elif img.shape[2] == 4:
img = img[:, :, :3]
return img
stop_im = Image.open('./stopsign-1.png')
stop_im = stop_im.crop((40, 30, 150, 180)) # 110 x 150
stop_caffe = conv_pil(stop_im)
stop_data = transformer.preprocess('data', stop_caffe)
probs = predict(stop_data)
plt.imshow(stop_im) # X: 25-90, Y: 40-125
# # Experiment 1
# Generate random stickers of black, white, and gray colors.
#
# Result: Failed after certain iterations.
# +
# Prepare "art stickers"
three_colors = [(240,245,245,0), (20,20,20,0), (60, 60, 60, 0)] # Black, white, gray stickers
widths = range(10, 11)
heights = range(10, 11)
num_stickers = 10
color_pallettes = [
(0, 152, 116, 0), # emerald
(221, 65, 36, 0), # tangerine tango
(214, 80, 118, 0), # honeysuckle
(68, 184, 172, 0), # turquoise
(239, 192, 80, 0), # mimosa
(91, 94, 166, 0), # blue izis
(155, 35, 53, 0), # chilli pepper
(223, 207, 190, 0), # sand dollar
(85, 180, 176, 0), # blue turquoise
(225, 93, 68, 0), # tigerlily
(127, 205, 205, 0), # aqua sky
(188, 36, 60, 0), # true red
(195, 68, 122, 0), # fuchsia rose
(152, 180, 212, 0), # cerulean blue
]
def paste_stickers(im, stickers, colors):
newim = im.copy()
draw = ImageDraw.Draw(newim)
for j in range(stickers):
w = choice(widths)
h = choice(heights)
x = randrange(25, 91-w)
y = randrange(40, 126-h)
color = choice(colors)
draw.rectangle((x, y, x+w, y+h), fill=color)
del draw
return newim
# +
def get_prob_label(prob):
probs = prob[0]
top_k = probs.argsort()[::-1]
return top_k[0]
def print_topk(prob):
probs = prob[0]
top_k = probs.argsort()[::-1]
for pred in top_k[:6]:
percent = round(probs[pred] * 100, 2)
pred_formatted = "%03d" % pred
format_string = "label: {cls} ({label}), certainty: {certainty}%"
print format_string.format(cls=pred_formatted, label=get_label_name(pred), certainty=percent)
# -
from PIL import ImageDraw
from random import randrange, choice
for i in range(1000):
print i,
newim = paste_stickers(stop_im, num_stickers, three_colors)
caffe_im = conv_pil(newim)
data = transformer.preprocess('data', caffe_im)
prob = predict(data, 1, False)
if get_prob_label(prob) != 919: # not a street sign
print '\nHit at iteration %d' % i
print_topk(prob)
break
del data
del caffe_im
del newim
display(data)
# # Experiment 2
# Use color pallettes here:
# https://i.pinimg.com/736x/ae/b0/01/aeb001b62989e505f22ebe0d6e0780b5--colour-hex-codes-colour-palettes.jpg
num_trials = 1000
for i in range(num_trials):
newim = paste_stickers(stop_im, num_stickers, color_pallettes)
caffe_im = conv_pil(newim)
data = transformer.preprocess('data', caffe_im)
prob = predict(data, 1, False)
if get_prob_label(prob) != 919: # not a street sign
print 'Hit at iteration %d' % i
print_topk(prob)
display(data)
break
del data
del caffe_im
del newim
|
neural-nets-are-weird/notebooks/Traffic Signs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import tensorflow as tf
import tensorflow.contrib.slim as slim
#List out our bandits. Currently bandit 4 (index#3) is set to most often provide a positive reward.
bandits = [0.2,0,-0.2,-5]
num_bandits = len(bandits)
def pullBandit(bandit):
#Get a random number.
result = np.random.randn(1)
if result > bandit:
#return a positive reward.
return 1
else:
#return a negative reward.
return -1
num_tests = 20
print([ pullBandit(bandits[0]) for _ in range(num_tests) ])
print([ pullBandit(bandits[1]) for _ in range(num_tests) ])
print([ pullBandit(bandits[2]) for _ in range(num_tests) ])
print([ pullBandit(bandits[3]) for _ in range(num_tests) ])
tf.reset_default_graph()
value = tf.constant([1, 2, 3, 4], dtype=tf.float32)
indices1 = tf.constant([0], dtype=tf.int32)
indices2 = tf.constant([2], dtype=tf.int32)
output1 = tf.slice(value, indices1, [2])
output2 = tf.slice(value, indices2, [1])
with tf.Session() as sess:
o1, o2 = sess.run([output1, output2])
print(o1)
print(o2)
# +
tf.reset_default_graph()
#These two lines established the feed-forward part of the network. This does the actual choosing.
weights = tf.Variable(tf.ones([num_bandits]))
chosen_action = tf.argmax(weights,0)
#The next six lines establish the training proceedure. We feed the reward and chosen action into the network
#to compute the loss, and use it to update the network.
reward_holder = tf.placeholder(shape=[1],dtype=tf.float32)
action_holder = tf.placeholder(shape=[1],dtype=tf.int32)
responsible_weight = tf.slice(weights,action_holder,[1])
loss = -(tf.log(responsible_weight)*reward_holder)
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001)
update = optimizer.minimize(loss)
# +
total_episodes = 1000 #Set total number of episodes to train agent on.
total_reward = np.zeros(num_bandits) #Set scoreboard for bandits to 0.
e = 0.1 #Set the chance of taking a random action.
init = tf.global_variables_initializer()
# Launch the tensorflow graph
with tf.Session() as sess:
sess.run(init)
i = 0
while i < total_episodes:
#Choose either a random action or one from our network.
if np.random.rand(1) < e:
action = np.random.randint(num_bandits)
else:
action = sess.run(chosen_action)
reward = pullBandit(bandits[action]) #Get our reward from picking one of the bandits.
#Update the network.
_,resp,ww = sess.run([update,responsible_weight,weights], feed_dict={reward_holder:[reward],action_holder:[action]})
#Update our running tally of scores.
total_reward[action] += reward
if i % 50 == 0:
print("Running reward for the " + str(num_bandits) + " bandits: " + str(total_reward))
i+=1
print("The agent thinks bandit " + str(np.argmax(ww)+1) + " is the most promising....")
if np.argmax(ww) == np.argmax(-np.array(bandits)):
print("...and it was right!")
else:
print("...and it was wrong!")
# -
class contextual_bandit():
def __init__(self):
self.state = 0
#List out our bandits. Currently arms 4, 2, and 1 (respectively) are the most optimal.
self.bandits = np.array([[0.2,0,-0.0,-5],[0.1,-5,1,0.25],[-5,5,5,5]])
self.num_bandits = self.bandits.shape[0]
self.num_actions = self.bandits.shape[1]
def getBandit(self):
self.state = np.random.randint(0,len(self.bandits)) #Returns a random state for each episode.
return self.state
def pullArm(self,action):
#Get a random number.
bandit = self.bandits[self.state,action]
result = np.random.randn(1)
if result > bandit:
#return a positive reward.
return 1
else:
#return a negative reward.
return -1
class agent():
def __init__(self, lr, s_size,a_size):
#These lines established the feed-forward part of the network. The agent takes a state and produces an action.
self.state_in= tf.placeholder(shape=[1],dtype=tf.int32)
state_in_OH = slim.one_hot_encoding(self.state_in,s_size)
output = slim.fully_connected(state_in_OH,a_size,\
biases_initializer=None,activation_fn=tf.nn.sigmoid,weights_initializer=tf.ones_initializer())
self.output = tf.reshape(output,[-1])
self.chosen_action = tf.argmax(self.output,0)
#The next six lines establish the training proceedure. We feed the reward and chosen action into the network
#to compute the loss, and use it to update the network.
self.reward_holder = tf.placeholder(shape=[1],dtype=tf.float32)
self.action_holder = tf.placeholder(shape=[1],dtype=tf.int32)
self.responsible_weight = tf.slice(self.output,self.action_holder,[1])
self.loss = -(tf.log(self.responsible_weight)*self.reward_holder)
optimizer = tf.train.GradientDescentOptimizer(learning_rate=lr)
self.update = optimizer.minimize(self.loss)
# +
tf.reset_default_graph() #Clear the Tensorflow graph.
cBandit = contextual_bandit() #Load the bandits.
myAgent = agent(lr=0.001,s_size=cBandit.num_bandits,a_size=cBandit.num_actions) #Load the agent.
weights = tf.trainable_variables()[0] #The weights we will evaluate to look into the network.
total_episodes = 10000 #Set total number of episodes to train agent on.
total_reward = np.zeros([cBandit.num_bandits,cBandit.num_actions]) #Set scoreboard for bandits to 0.
e = 0.1 #Set the chance of taking a random action.
init = tf.global_variables_initializer()
# Launch the tensorflow graph
with tf.Session() as sess:
sess.run(init)
i = 0
while i < total_episodes:
s = cBandit.getBandit() #Get a state from the environment.
#Choose either a random action or one from our network.
if np.random.rand(1) < e:
action = np.random.randint(cBandit.num_actions)
else:
action = sess.run(myAgent.chosen_action,feed_dict={myAgent.state_in:[s]})
reward = cBandit.pullArm(action) #Get our reward for taking an action given a bandit.
#Update the network.
feed_dict={myAgent.reward_holder:[reward],myAgent.action_holder:[action],myAgent.state_in:[s]}
_,ww = sess.run([myAgent.update,weights], feed_dict=feed_dict)
#Update our running tally of scores.
total_reward[s,action] += reward
if i % 500 == 0:
print("Mean reward for each of the " + str(cBandit.num_bandits) + " bandits: " + str(np.mean(total_reward,axis=1)))
i+=1
for a in range(cBandit.num_bandits):
print("The agent thinks action " + str(np.argmax(ww[a])+1) + " for bandit " + str(a+1) + " is the most promising....")
if np.argmax(ww[a]) == np.argmin(cBandit.bandits[a]):
print("...and it was right!")
else:
print("...and it was wrong!")
# -
|
notebooks/n_armed_bandit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SIGIR'21 DLG4NLP Tutorial Demo: Semantic Parsing
# In this tutorial demo, we will use the Graph4NLP library to build a GNN-based semantic parsing model. The model consists of
# - graph construction module (e.g., node embedding based dynamic graph)
# - graph embedding module (e.g., Bi-Sep GAT)
# - predictoin module (e.g., RNN decoder with attention, copy and coverage mechanisms)
#
# We will use the built-in Graph2Seq model APIs to build the model, and evaluate it on the Jobs dataset.
# ### Environment setup
# 1. Create virtual environment
# ```
# conda create --name graph4nlp python=3.7
# conda activate graph4nlp
# ```
#
# 2. Install [graph4nlp](https://github.com/graph4ai/graph4nlp) library
# - Clone the github repo
# ```
# git clone -b stable https://github.com/graph4ai/graph4nlp.git
# # cd graph4nlp
# ```
# - Then run `./configure` (or `./configure.bat` if you are using Windows 10) to config your installation. The configuration program will ask you to specify your CUDA version. If you do not have a GPU, please choose 'cpu'.
# ```
# ./configure
# ```
# - Finally, install the package
# ```
# python setup.py install
# ```
#
# 3. Set up StanfordCoreNLP (for static graph construction only, unnecessary for this demo because preprocessed data is provided)
# - Download [StanfordCoreNLP](https://stanfordnlp.github.io/CoreNLP/)
# - Go to the root folder and start the server
# ```
# java -mx4g -cp "*" edu.stanford.nlp.pipeline.StanfordCoreNLPServer -port 9000 -timeout 15000
# ```
# +
import os
import yaml
import numpy as np
import torch
import torch.optim as optim
from torch.utils.data import DataLoader
from graph4nlp.pytorch.datasets.jobs import JobsDataset
from graph4nlp.pytorch.models.graph2seq import Graph2Seq
from graph4nlp.pytorch.models.graph2seq_loss import Graph2SeqLoss
from graph4nlp.pytorch.modules.graph_construction import *
from graph4nlp.pytorch.modules.evaluation.base import EvaluationMetricBase
from graph4nlp.pytorch.modules.utils.config_utils import update_values
from graph4nlp.pytorch.modules.utils.copy_utils import prepare_ext_vocab
from graph4nlp.pytorch.modules.config import get_basic_args
# -
class Jobs:
def __init__(self, opt):
super(Jobs, self).__init__()
self.opt = opt
self.use_copy = self.opt["decoder_args"]["rnn_decoder_share"]["use_copy"]
self.use_coverage = self.opt["decoder_args"]["rnn_decoder_share"]["use_coverage"]
self._build_device(self.opt)
self._build_dataloader()
self._build_model()
self._build_loss_function()
self._build_optimizer()
self._build_evaluation()
def _build_device(self, opt):
seed = opt["seed"]
np.random.seed(seed)
if opt["use_gpu"] != 0 and torch.cuda.is_available():
print('[ Using CUDA ]')
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
from torch.backends import cudnn
cudnn.benchmark = True
device = torch.device('cuda' if opt["gpu"] < 0 else 'cuda:%d' % opt["gpu"])
else:
print('[ Using CPU ]')
device = torch.device('cpu')
self.device = device
def _build_dataloader(self):
if self.opt["graph_construction_args"]["graph_construction_share"]["graph_type"] == "dependency":
topology_builder = DependencyBasedGraphConstruction
graph_type = 'static'
dynamic_init_topology_builder = None
elif self.opt["graph_construction_args"]["graph_construction_share"]["graph_type"] == "constituency":
topology_builder = ConstituencyBasedGraphConstruction
graph_type = 'static'
dynamic_init_topology_builder = None
elif self.opt["graph_construction_args"]["graph_construction_share"]["graph_type"] == "node_emb":
topology_builder = NodeEmbeddingBasedGraphConstruction
graph_type = 'dynamic'
dynamic_init_topology_builder = None
elif self.opt["graph_construction_args"]["graph_construction_share"]["graph_type"] == "node_emb_refined":
topology_builder = NodeEmbeddingBasedRefinedGraphConstruction
graph_type = 'dynamic'
dynamic_init_graph_type = self.opt["graph_construction_args"]["graph_construction_private"][
"dynamic_init_graph_type"]
if dynamic_init_graph_type is None or dynamic_init_graph_type == 'line':
dynamic_init_topology_builder = None
elif dynamic_init_graph_type == 'dependency':
dynamic_init_topology_builder = DependencyBasedGraphConstruction
elif dynamic_init_graph_type == 'constituency':
dynamic_init_topology_builder = ConstituencyBasedGraphConstruction
else:
raise RuntimeError('Define your own dynamic_init_topology_builder')
else:
raise NotImplementedError("Define your topology builder.")
dataset = JobsDataset(root_dir=self.opt["graph_construction_args"]["graph_construction_share"]["root_dir"],
pretrained_word_emb_name=self.opt["pretrained_word_emb_name"],
pretrained_word_emb_url=self.opt["pretrained_word_emb_url"],
pretrained_word_emb_cache_dir=self.opt["pretrained_word_emb_cache_dir"],
merge_strategy=self.opt["graph_construction_args"]["graph_construction_private"][
"merge_strategy"],
edge_strategy=self.opt["graph_construction_args"]["graph_construction_private"][
"edge_strategy"],
seed=self.opt["seed"],
word_emb_size=self.opt["word_emb_size"],
share_vocab=self.opt["graph_construction_args"]["graph_construction_share"][
"share_vocab"],
graph_type=graph_type,
topology_builder=topology_builder,
topology_subdir=self.opt["graph_construction_args"]["graph_construction_share"][
"topology_subdir"],
thread_number=self.opt["graph_construction_args"]["graph_construction_share"][
"thread_number"],
dynamic_graph_type=self.opt["graph_construction_args"]["graph_construction_share"][
"graph_type"],
dynamic_init_topology_builder=dynamic_init_topology_builder,
dynamic_init_topology_aux_args=None)
self.train_dataloader = DataLoader(dataset.train, batch_size=self.opt["batch_size"], shuffle=True,
num_workers=1,
collate_fn=dataset.collate_fn)
self.test_dataloader = DataLoader(dataset.test, batch_size=self.opt["batch_size"], shuffle=False, num_workers=1,
collate_fn=dataset.collate_fn)
self.vocab = dataset.vocab_model
def _build_model(self):
self.model = Graph2Seq.from_args(self.opt, self.vocab).to(self.device)
def _build_loss_function(self):
self.loss = Graph2SeqLoss(ignore_index=self.vocab.out_word_vocab.PAD,
use_coverage=self.use_coverage, coverage_weight=0.3)
def _build_optimizer(self):
parameters = [p for p in self.model.parameters() if p.requires_grad]
self.optimizer = optim.Adam(parameters, lr=self.opt["learning_rate"])
def _build_evaluation(self):
self.metrics = [ExpressionAccuracy()]
def train(self):
max_score = -1
self._best_epoch = -1
for epoch in range(200):
self.model.train()
self.train_epoch(epoch, split="train")
self._adjust_lr(epoch)
if epoch >= 0:
score = self.evaluate(split="test")
if score >= max_score:
print("Best model saved, epoch {}".format(epoch))
self.save_checkpoint("best.pth")
self._best_epoch = epoch
max_score = max(max_score, score)
if epoch >= 30 and self._stop_condition(epoch):
break
return max_score
def _stop_condition(self, epoch, patience=20):
return epoch > patience + self._best_epoch
def _adjust_lr(self, epoch):
def set_lr(optimizer, decay_factor):
for group in optimizer.param_groups:
group['lr'] = group['lr'] * decay_factor
epoch_diff = epoch - self.opt["lr_start_decay_epoch"]
if epoch_diff >= 0 and epoch_diff % self.opt["lr_decay_per_epoch"] == 0:
if self.opt["learning_rate"] > self.opt["min_lr"]:
set_lr(self.optimizer, self.opt["lr_decay_rate"])
self.opt["learning_rate"] = self.opt["learning_rate"] * self.opt["lr_decay_rate"]
print("Learning rate adjusted: {:.5f}".format(self.opt["learning_rate"]))
def train_epoch(self, epoch, split="train"):
assert split in ["train"]
print("Start training in split {}, Epoch: {}".format(split, epoch))
loss_collect = []
dataloader = self.train_dataloader
step_all_train = len(dataloader)
for step, data in enumerate(dataloader):
graph, tgt, gt_str = data["graph_data"], data["tgt_seq"], data["output_str"]
graph = graph.to(self.device)
tgt = tgt.to(self.device)
oov_dict = None
if self.use_copy:
oov_dict, tgt = prepare_ext_vocab(graph, self.vocab, gt_str=gt_str, device=self.device)
prob, enc_attn_weights, coverage_vectors = self.model(graph, tgt, oov_dict=oov_dict)
loss = self.loss(logits=prob, label=tgt, enc_attn_weights=enc_attn_weights,
coverage_vectors=coverage_vectors)
loss_collect.append(loss.item())
if step % self.opt["loss_display_step"] == 0 and step != 0:
print("Epoch {}: [{} / {}] loss: {:.3f}".format(epoch, step, step_all_train,
np.mean(loss_collect)))
loss_collect = []
self.optimizer.zero_grad()
loss.backward()
self.optimizer.step()
def evaluate(self, split="val"):
self.model.eval()
pred_collect = []
gt_collect = []
assert split in ["val", "test"]
dataloader = self.val_dataloader if split == "val" else self.test_dataloader
for data in dataloader:
graph, tgt, gt_str = data["graph_data"], data["tgt_seq"], data["output_str"]
graph = graph.to(self.device)
if self.use_copy:
oov_dict = prepare_ext_vocab(batch_graph=graph, vocab=self.vocab, device=self.device)
ref_dict = oov_dict
else:
oov_dict = None
ref_dict = self.vocab.out_word_vocab
prob, _, _ = self.model(graph, oov_dict=oov_dict)
pred = prob.argmax(dim=-1)
pred_str = wordid2str(pred.detach().cpu(), ref_dict)
pred_collect.extend(pred_str)
gt_collect.extend(gt_str)
score = self.metrics[0].calculate_scores(ground_truth=gt_collect, predict=pred_collect)
print("Evaluation accuracy in `{}` split: {:.3f}".format(split, score))
return score
@torch.no_grad()
def translate(self):
self.model.eval()
pred_collect = []
gt_collect = []
dataloader = self.test_dataloader
for data in dataloader:
graph, tgt, gt_str = data["graph_data"], data["tgt_seq"], data["output_str"]
graph = graph.to(self.device)
if self.use_copy:
oov_dict = prepare_ext_vocab(batch_graph=graph, vocab=self.vocab, device=self.device)
ref_dict = oov_dict
else:
oov_dict = None
ref_dict = self.vocab.out_word_vocab
pred = self.model.translate(batch_graph=graph, oov_dict=oov_dict, beam_size=4, topk=1)
pred_ids = pred[:, 0, :] # we just use the top-1
pred_str = wordid2str(pred_ids.detach().cpu(), ref_dict)
pred_collect.extend(pred_str)
gt_collect.extend(gt_str)
score = self.metrics[0].calculate_scores(ground_truth=gt_collect, predict=pred_collect)
return score
def load_checkpoint(self, checkpoint_name):
checkpoint_path = os.path.join(self.opt["checkpoint_save_path"], checkpoint_name)
self.model.load_state_dict(torch.load(checkpoint_path))
def save_checkpoint(self, checkpoint_name):
checkpoint_path = os.path.join(self.opt["checkpoint_save_path"], checkpoint_name)
if not os.path.exists(self.opt["checkpoint_save_path"]):
os.makedirs(self.opt["checkpoint_save_path"], exist_ok=True)
torch.save(self.model.state_dict(), checkpoint_path)
# +
class ExpressionAccuracy(EvaluationMetricBase):
def __init__(self):
super(ExpressionAccuracy, self).__init__()
def calculate_scores(self, ground_truth, predict):
correct = 0
assert len(ground_truth) == len(predict)
for gt, pred in zip(ground_truth, predict):
if gt == pred:
correct += 1.
return correct / len(ground_truth)
def wordid2str(word_ids, vocab):
ret = []
assert len(word_ids.shape) == 2, print(word_ids.shape)
for i in range(word_ids.shape[0]):
id_list = word_ids[i, :]
ret_inst = []
for j in range(id_list.shape[0]):
if id_list[j] == vocab.EOS:
break
token = vocab.getWord(id_list[j])
ret_inst.append(token)
ret.append(" ".join(ret_inst))
return ret
# +
# config setup
config_file = '../config/jobs/gat_bi_sep_dynamic_node_emb.yaml'
config = yaml.load(open(config_file, 'r'), Loader=yaml.FullLoader)
opt = get_basic_args(graph_construction_name=config["graph_construction_name"],
graph_embedding_name=config["graph_embedding_name"],
decoder_name=config["decoder_name"])
update_values(to_args=opt, from_args_list=[config, config["other_args"]])
# -
# run the model
runner = Jobs(opt)
max_score = runner.train()
print("Train finish, best val score: {:.3f}".format(max_score))
runner.load_checkpoint("best.pth")
# runner.evaluate("test")
test_score = runner.translate()
print('Final test accuracy: {}'.format(test_score))
|
SIGIR2021_demo/semantic_parsing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (PyMC3 Dev)
# language: python
# name: pymc3-dev
# ---
# # Updating priors
# In this notebook, I will show how it is possible to update the priors as new data becomes available. The example is a slightly modified version of the linear regression in the [Getting started with PyMC3](https://github.com/pymc-devs/pymc3/blob/master/docs/source/notebooks/getting_started.ipynb) notebook.
# +
# %matplotlib inline
import warnings
import arviz as az
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pymc3 as pm
import theano.tensor as tt
from pymc3 import Model, Normal, Slice, sample
from pymc3.distributions import Interpolated
from scipy import stats
from theano import as_op
plt.style.use("seaborn-darkgrid")
print(f"Running on PyMC3 v{pm.__version__}")
# -
warnings.filterwarnings("ignore")
# ## Generating data
# +
# Initialize random number generator
np.random.seed(93457)
# True parameter values
alpha_true = 5
beta0_true = 7
beta1_true = 13
# Size of dataset
size = 100
# Predictor variable
X1 = np.random.randn(size)
X2 = np.random.randn(size) * 0.2
# Simulate outcome variable
Y = alpha_true + beta0_true * X1 + beta1_true * X2 + np.random.randn(size)
# -
# ## Model specification
# Our initial beliefs about the parameters are quite informative (sigma=1) and a bit off the true values.
# +
basic_model = Model()
with basic_model:
# Priors for unknown model parameters
alpha = Normal("alpha", mu=0, sigma=1)
beta0 = Normal("beta0", mu=12, sigma=1)
beta1 = Normal("beta1", mu=18, sigma=1)
# Expected value of outcome
mu = alpha + beta0 * X1 + beta1 * X2
# Likelihood (sampling distribution) of observations
Y_obs = Normal("Y_obs", mu=mu, sigma=1, observed=Y)
# draw 1000 posterior samples
trace = sample(1000)
# -
az.plot_trace(trace);
# In order to update our beliefs about the parameters, we use the posterior distributions, which will be used as the prior distributions for the next inference. The data used for each inference iteration has to be independent from the previous iterations, otherwise the same (possibly wrong) belief is injected over and over in the system, amplifying the errors and misleading the inference. By ensuring the data is independent, the system should converge to the true parameter values.
#
# Because we draw samples from the posterior distribution (shown on the right in the figure above), we need to estimate their probability density (shown on the left in the figure above). [Kernel density estimation](https://en.wikipedia.org/wiki/Kernel_density_estimation) (KDE) is a way to achieve this, and we will use this technique here. In any case, it is an empirical distribution that cannot be expressed analytically. Fortunately PyMC3 provides a way to use custom distributions, via `Interpolated` class.
def from_posterior(param, samples):
smin, smax = np.min(samples), np.max(samples)
width = smax - smin
x = np.linspace(smin, smax, 100)
y = stats.gaussian_kde(samples)(x)
# what was never sampled should have a small probability but not 0,
# so we'll extend the domain and use linear approximation of density on it
x = np.concatenate([[x[0] - 3 * width], x, [x[-1] + 3 * width]])
y = np.concatenate([[0], y, [0]])
return Interpolated(param, x, y)
# Now we just need to generate more data and build our Bayesian model so that the prior distributions for the current iteration are the posterior distributions from the previous iteration. It is still possible to continue using NUTS sampling method because `Interpolated` class implements calculation of gradients that are necessary for Hamiltonian Monte Carlo samplers.
traces = [trace]
for _ in range(10):
# generate more data
X1 = np.random.randn(size)
X2 = np.random.randn(size) * 0.2
Y = alpha_true + beta0_true * X1 + beta1_true * X2 + np.random.randn(size)
model = Model()
with model:
# Priors are posteriors from previous iteration
alpha = from_posterior("alpha", trace["alpha"])
beta0 = from_posterior("beta0", trace["beta0"])
beta1 = from_posterior("beta1", trace["beta1"])
# Expected value of outcome
mu = alpha + beta0 * X1 + beta1 * X2
# Likelihood (sampling distribution) of observations
Y_obs = Normal("Y_obs", mu=mu, sigma=1, observed=Y)
# draw 10000 posterior samples
trace = sample(1000)
traces.append(trace)
# +
print("Posterior distributions after " + str(len(traces)) + " iterations.")
cmap = mpl.cm.autumn
for param in ["alpha", "beta0", "beta1"]:
plt.figure(figsize=(8, 2))
for update_i, trace in enumerate(traces):
samples = trace[param]
smin, smax = np.min(samples), np.max(samples)
x = np.linspace(smin, smax, 100)
y = stats.gaussian_kde(samples)(x)
plt.plot(x, y, color=cmap(1 - update_i / len(traces)))
plt.axvline({"alpha": alpha_true, "beta0": beta0_true, "beta1": beta1_true}[param], c="k")
plt.ylabel("Frequency")
plt.title(param)
plt.tight_layout();
# -
# You can re-execute the last two cells to generate more updates.
#
# What is interesting to note is that the posterior distributions for our parameters tend to get centered on their true value (vertical lines), and the distribution gets thiner and thiner. This means that we get more confident each time, and the (false) belief we had at the beginning gets flushed away by the new data we incorporate.
# %load_ext watermark
# %watermark -n -u -v -iv -w
|
examples/pymc3_howto/updating_priors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Skew test
#
# <NAME>
#
# [MIT License](https://en.wikipedia.org/wiki/MIT_License)
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style='white')
from thinkstats2 import Pmf, Cdf
import thinkstats2
import thinkplot
decorate = thinkplot.config
# -
# Suppose you buy a loaf of bread every day for a year, take it
# home, and weigh it. You suspect that the distribution of weights is
# more skewed than a normal distribution with the same mean and
# standard deviation.
#
# To test your suspicion, write a definition for a class named
# `SkewTest` that extends `thinkstats.HypothesisTest` and provides
# two methods:
#
# * `TestStatistic` should compute the skew of a given sample.
#
# * `RunModel` should simulate the null hypothesis and return
# simulated data.
class HypothesisTest(object):
"""Represents a hypothesis test."""
def __init__(self, data):
"""Initializes.
data: data in whatever form is relevant
"""
self.data = data
self.MakeModel()
self.actual = self.TestStatistic(data)
self.test_stats = None
def PValue(self, iters=1000):
"""Computes the distribution of the test statistic and p-value.
iters: number of iterations
returns: float p-value
"""
self.test_stats = np.array([self.TestStatistic(self.RunModel())
for _ in range(iters)])
count = sum(self.test_stats >= self.actual)
return count / iters
def MaxTestStat(self):
"""Returns the largest test statistic seen during simulations.
"""
return np.max(self.test_stats)
def PlotHist(self, label=None):
"""Draws a Cdf with vertical lines at the observed test stat.
"""
plt.hist(self.test_stats, color='C4', alpha=0.5)
plt.axvline(self.actual, linewidth=3, color='0.8')
plt.xlabel('Test statistic')
plt.ylabel('Count')
plt.title('Distribution of the test statistic under the null hypothesis')
def TestStatistic(self, data):
"""Computes the test statistic.
data: data in whatever form is relevant
"""
raise UnimplementedMethodException()
def MakeModel(self):
"""Build a model of the null hypothesis.
"""
pass
def RunModel(self):
"""Run the model of the null hypothesis.
returns: simulated data
"""
raise UnimplementedMethodException()
# +
# Solution goes here
# -
# To test this class, I'll generate a sample from an actual Gaussian distribution, so the null hypothesis is true.
mu = 1000
sigma = 35
data = np.random.normal(mu, sigma, size=365)
# Now we can make a `SkewTest` and compute the observed skewness.
test = SkewTest(data)
test.actual
# Here's the p-value.
test = SkewTest(data)
test.PValue()
# And the distribution of the test statistic under the null hypothesis.
test.PlotHist()
# Most of the time the p-value exceeds 5%, so we would conclude that the observed skewness could plausibly be due to random sample.
#
# But let's see how often we get a false positive.
# +
iters = 100
count = 0
for i in range(iters):
data = np.random.normal(mu, sigma, size=365)
test = SkewTest(data)
p_value = test.PValue()
if p_value < 0.05:
count +=1
print(count/iters)
# -
# In the long run, the false positive rate is the threshold we used, 5%.
|
DSC 530 - Data Exploration and Analysis/ThinkStats2/examples/skew_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# セルに数式を入力する
import openpyxl
book = openpyxl.Workbook() # Excelブックを生成
sheet = book.active # アクティブなワークシートを取得
sheet['A1'] = 500 # A1セルに値を入力
sheet['A2'] = 300 # A2セルに値を入力
sheet['A3'] = '=SUM(A1:A2)' # A3セルにSUM()関数を入力
sheet['A3'].value # A3セルの数式を確認
# -
book.save('sample.xlsx') # ブックを保存
# +
# セルの幅と高さを設定する
import openpyxl
book = openpyxl.Workbook() # Workbookオブジェクトを生成
sheet = book.active # アクティブシートをストク
sheet['A1'] = 'column_dim' # A1セルに文字列を入力
sheet['B2'] = 'row_dim' # B2セルに文字列を入力
sheet.column_dimensions['A'].width = 30 # A列の幅を30に設定
sheet.row_dimensions[2].height = 100 # 2行目の高さを100に設定
sheet.column_dimensions['A'].width # A列の幅を確認
# -
sheet.row_dimensions[2].height # 2行目の高さを確認
book.save('dimensions.xlsx') # Excelブックとして保存
# +
# セルを結合する
import openpyxl
book = openpyxl.Workbook()
sheet = book.active
sheet.merge_cells('A1:A2') # セルを結合
sheet['A1'] = 'A1:A2結合' # セルに入力
sheet.merge_cells('B1:C1') # セルを結合
sheet['B1'] = 'B1:C1結合' # セルに入力
sheet.merge_cells('D1:G3') # セルを結合
sheet['D1'] = 'D1:G3結合' # セルに入力
book.save('merge_cells.xlsx')
# +
# セルの結合を解除する
import openpyxl
# カレントディレクトリのmerge_cells.xlsxを読み込む
book = openpyxl.load_workbook('merge_cells.xlsx')
sheet = book.active
sheet.unmerge_cells('A1:A2') # 結合を解除
sheet.unmerge_cells('B1:C1') # 結合を解除
sheet.unmerge_cells('D1:G3') # 結合を解除
book.save('merge_cells.xlsx')
# +
# ワークシートの1行目を固定する
import openpyxl
book = openpyxl.load_workbook('惣菜売上.xlsx')
sheet = book.active
sheet.freeze_panes = 'A2'
book.save('惣菜売上_freeze.xlsx')
# +
# レコード単位でデータを追加する
import openpyxl
def writeCell(rows, fname):
book = openpyxl.Workbook() # ブックを生成
sheet = book.active # アクティブなシートを取得
for row in rows: # 行数の数だけ繰り返す
sheet.append(row) # ワークシートに追加する
book.save(fname + '.xlsx') # ブックを保存
# -
# レコードのタプルをまとめたリストを作成
rows = [
('月', '商品A', '商品B'), # タイトル行
(1, 30, 35), # 12行のレコードデータ
(2, 10, 30),
(3, 40, 60),
(4, 50, 70),
(5, 20, 10),
(6, 30, 40),
(7, 50, 30),
(8, 65, 30),
(9, 70, 30),
(10, 50, 40),
(11, 60, 50),
(12, 65, 55),
]
# ファイル名
fname = '月別売上'
# writeCell()を実行
writeCell(rows, fname)
|
sample/Python_GOKUI/Python_GOKUI/chap06/sec03/Workbook4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Introduction
#
# This notebook demonstrates how to perform diffusivity and ionic conductivity analyses starting from a series of VASP AIMD simulations using Python Materials Genomics (pymatgen) and its add-on package pymatgen-analysis-diffusion. These notebooks are described in detail in
#
# <NAME>.; <NAME>.; <NAME>.; <NAME>. Data-Driven First-Principles Methods for the Study and Design of
# Alkali Superionic Conductors. Chem. Mater. 2017, 29 (1), 281–288 DOI: 10.1021/acs.chemmater.6b02648.
#
# If you find these notebooks useful and use the functionality demonstrated, please consider citing the above work.
#
# Let's start by importing some modules and classes that we will be using.
# +
from IPython.display import Image
# %matplotlib inline
import matplotlib.pyplot as plt
import json
import collections
from pymatgen.core import Structure
from pymatgen.analysis.diffusion.analyzer import DiffusionAnalyzer, \
get_arrhenius_plot, get_extrapolated_conductivity
from pymatgen.analysis.diffusion.aimd.pathway import ProbabilityDensityAnalysis
from pymatgen.analysis.diffusion.aimd.van_hove import VanHoveAnalysis
# -
# # Preparation
#
# The *DiffusionAnalyzer* class in pymatgen can be instantiated from a supplied list of sequential vasprun.xml output files from the AIMD simulations. An example code (commented out) is shown below.
# +
# files = ["run1/vasprun.xml", "run2/vasprun.xml", "run3/vasprun.xml"]
# analyzer = DiffusionAnalyzer.from_files(files, specie="Li", smoothed=False)
# -
# In this work, all trajectories are stored in an efficient document-based MongoDB database. The format of the documents in the database is a binary JSON format. Here, we will instead instantiate the *DiffusionAnalyzer* from a pre-serialized *DiffusionAnalyzer* for each temperature.
temperatures = [600, 800, 1000, 1200]
analyzers = collections.OrderedDict()
for temp in temperatures:
with open("aimd_data/%d.json" % temp) as f:
d = json.load(f)
analyzers[temp] = DiffusionAnalyzer.from_dict(d)
# # MSD vs time plot
#
# For each temperature, we can plot the mean square displacement against time as follows (only 1000 K shown).
plt = analyzers[1000].get_msd_plot()
title = plt.title("1000K", fontsize=24)
# # Activation energy and ionic conductivity
#
# From diffusivity at each temperature, we can obtain activation energy and room temperature ionic conductivity by constructing an Arrhenius plot.
# +
diffusivities = [d.diffusivity for d in analyzers.values()]
plt = get_arrhenius_plot(temperatures, diffusivities)
# -
# From the temperatures and diffusivities, one may obtained the extrapolated room-temperature conductivity as follows.
rts = get_extrapolated_conductivity(temperatures, diffusivities,
new_temp=300, structure=analyzers[800].structure,
species="Li")
print("The Li ionic conductivity for Li6PS5Cl at 300 K is %.4f mS/cm" % rts)
# # Probability density function analysis
#
# We can compute the probability density function from the AIMD trajectories using the *ProbabilityDensityAnalysis* class implemented in the pymatgen-diffusion add-on. We will use the calculation at 800K as an example. The probability density function can then be output to a CHGCAR-like file for visualization in VESTA.
structure = analyzers[800].structure
trajectories = [s.frac_coords for s in analyzers[800].get_drift_corrected_structures()]
pda = ProbabilityDensityAnalysis(structure, trajectories, species="Li")
pda.to_chgcar("aimd_data/CHGCAR.vasp") # Output to a CHGCAR-like file for visualization in VESTA.
# The VESTA visualization software can be used to visualize isosurfaces in the probability density. The 800K probability density function at an isosurface of 0.002 is shown below.
Image(filename='aimd_data/Isosurface_800K_0.png')
# # van Hove correlation function analysis
#
# We can compute the van Hove correlation function from a *DiffusionAnalyzer* using the *VanHoveAnalysis* class implemented in the pymatgen-diffusion add-on. We will use the calculation at 800K as an example.
vha = VanHoveAnalysis(analyzers[800])
# We can then plot the self ($G_s$) and distinct ($G_d$) parts of the van Hove correlation function as follows.
vha.get_3d_plot(mode="self")
vha.get_3d_plot(mode="distinct")
|
notebooks/2016-09-08-Data-driven First Principles Methods for the Study and Design of Alkali Superionic Conductors Part 3 - Diffusivity and Ionic Conductivity.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# <table>
# <tr>
# <td width=15%><img src="./img/UGA.png"></img></td>
# <td><center><h1>Introduction to Python for Data Sciences</h1></center></td>
# <td width=15%><a href="http://www.iutzeler.org" style="font-size: 16px; font-weight: bold"><NAME></a> </td>
# </tr>
# </table>
#
#
# <br/><br/>
#
# <center><a style="font-size: 40pt; font-weight: bold">Chap. 4 - Scikit Learn </a></center>
#
# <br/><br/>
#
#
# # 1- Scikit Learn
#
#
# Now that we explored data structures provided by the Pandas library, we will investigate how to learn over it using **Scikit-learn**.
#
# Scikit-learn is ont of the most celebrated and used machine learning library. It features a complete set of efficiently implemented machine learning algorithms for classification, regression, and clustering. Scikit-learn is designed to operate over Numpy, Scipy, and Pandas data structures.
#
# **Links:** [Scikit-learn webpage](http://scikit-learn.org) [Wikipedia article](https://en.wikipedia.org/wiki/Scikit-learn)
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Machine Learning problems
# *Machine learning* is the task of predicting properties out of some data. The *dataset* consists in several *examples* or *samples* and the associated target properties can be available, partially available, or not at all; we respectively call these setting *supervised*, *semi-supervised*, *unsupervised*. The examples are made out of one or several *features* or *attributes* that can be of different types (real number, discretes values, strings, booleans, etc.).
#
# Learning problems can be broadly divided in a few categories:
# * **supervised learning**
# * **classification:** Place incoming data into a finite number or classes by learning over labeled data. Example: Classifying iris into species based on recorded petal and sentil sizes from the 3 species.
# * **regression:** Predict a value from example data. To the difference of classification, the output value is continuous. Example: Predict the carbon monoxide concentration for next years based on previous measures.
# * **unsupervised learning**
# * **clustering:** Place the data (both new and the dataset) into a finite number of classes. To the difference with classification, no labeled data is provided. Example: Create market segments from customer information for targeted advertising.
# * **dimension reduction:** Discard uniformative features for the purpose of visualization or efficient storage. Example: Creation of eigenfaces in visage recognition.
#
#
# The following flowchart can be found on the [Scikit Learn website](http://scikit-learn.org/stable/tutorial/machine_learning_map/index.html):
#
# 
#
# ## Learning with Scikit Learn
#
# The process of learning and predicting with Scikit Learn follows three main steps:<br/>
# **1. Selecting and adjusting a model**<br/>
# **2. Fitting the model to the data**<br/>
# **3. Predicting from this fitted model**
# We will illustrate this process on a simple *linear model*
# $$ y = a x + b + \nu$$
# where
# * $(x,y)\in\mathbb{R}^m\times\mathbb{R}^m$ are the data points. $x$ contains the examples and $y$ the associated outputs
# * $a,b$ are the model coefficients to estimate
# * $\nu$ is a standard centered white Gaussian noise
# +
a = np.random.randn()*5 # Drawing randomly the slope
b = np.random.rand()*10 # Drawing randomly the initial point
m = 50 # number of points
x = np.random.rand(m,1)*10 # Drawing randomly abscisses
y = a*x + b + np.random.randn(m,1) # y = ax+b + noise
plt.scatter(x, y)
# -
# ### 1. Selecting and adjusting a model
#
# As we want to fit a linear model $y=ax+b$ through the data, we will import the `Linear Regression` module from scikit learn with `sklearn.linear_model import LinearRegression`.
#
# As our model has a non null coefficient at the origin, the model needs an *intercept*. This can be tuned, along with several other parameters, see Scikit Learn's [linear_model documentation](http://Scikit-Learn.org/stable/modules/linear_model.html).
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression(fit_intercept=True)
print(model)
# -
# This terminates our model tuning. Notice that we have described our model, but no learning or fitting has been done.
# ### 2. Fitting the model to the data
#
#
# Applying our model to the data $(x,y)$ is done using the `fit` method.
model.fit(x,y)
# Once the model is fitted, one can observe the learned coefficients:
# * `coef_` for the model coefficients ($a$ here)
# * `intercept_` foe the intercept ($b$ here)
print("Learned coefficients: a = {:.6f} \t b = {:.6f}".format(float(model.coef_),float(model.intercept_)))
print("True coefficients: a = {:.6f} \t b = {:.6f}".format(a,b))
# ### 3. Predicting from this fitted model
#
# From a feature matrix, the method `predict` returns the predicted output from the fitted model.
xFit = np.linspace(-2,12,21).reshape(-1, 1)
yFit = model.predict(xFit)
plt.scatter(x, y , label="data")
plt.plot(xFit, yFit , label="model",color="r")
plt.legend()
# ## Preprocessing Data
#
# ### Data format
#
# Scikit Learn can take as an input (*i.e.* passed to `fit` and `predict`) several format including:
# * Numpy arrays. **Warning:** the data *has* to be **2D** even if there is only one example or one feature.
# * Pandas dataframes.
# * SciPy sparse matrices.
#
# The *examples/samples* of the datasets are stored as *rows*.<br/>
# The *features* are the *columns*.
#
# ### Training/Testing sets
#
# In order to *cross-validate* our model, it is customary to split the dataset into training and testing subsets. It can be done manually but there is also a dedicated method.
# +
from sklearn.model_selection import train_test_split
xTrain, xTest, yTrain, yTest = train_test_split(x,y)
# -
print(xTrain.shape,yTrain.shape)
print(xTest.shape,yTest.shape)
# Let us use cross validation to compare linear model and linear model with intercept.
# +
from sklearn.linear_model import LinearRegression
model1 = LinearRegression(fit_intercept=True)
model2 = LinearRegression(fit_intercept=False)
model1.fit(xTrain,yTrain)
yPre1 = model1.predict(xTest)
error1 = np.linalg.norm(yTest-yPre1)
model2.fit(xTrain,yTrain)
yPre2 = model2.predict(xTest)
error2 = np.linalg.norm(yTest-yPre2)
print("Testing Error with intercept:", error1, "\t without intercept:" ,error2)
# -
plt.scatter(xTrain, yTrain , label="Train data")
plt.scatter(xTest, yTest , color= 'k' , label="Test data")
plt.plot(xTest, yPre1 , color='r', label="model w/ intercept (err = {:.1f})".format(error1))
plt.plot(xTest, yPre2 , color='m', label="model w/o intercept (err = {:.1f})".format(error2))
plt.legend()
# ### Performance metrics
#
# In order to quantitatively evaluate the models, Scikit Learn provide a wide range of [metrics](http://scikit-learn.org/stable/modules/classes.html#sklearn-metrics-metrics), we will see some of them in the following examples.
|
4-1_Scikit_Learn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import numpy as np
import pandas as pd
with open('analysis_dataset.json', 'r') as js:
games = json.load(js)
gameId = '4733146699'
games[gameId]
df_champion = pd.read_csv('../../data/csv/champion.csv')
df_champion = df_champion.drop(['Unnamed: 0'], axis=1)
column = ['champion','championId']
df_champion.tail()
champion_id_list = list(df_champion.key)
champion_id_list.sort()
# # Prior probability( = P(H))
def safe_divide(x, y):
"""
prevent from dividing by 0
:param x: numerator
:param y: denominator
:return: float
"""
try:
return round(x / y, 4)
except ZeroDivisionError:
return 0
# # Prior probability ( = P(H))
# ## Champion win rate
def champion_win_rate(champion_id):
"""
챔피언 하나의 승률.
:param champion_id: championId
:return: [승률, 챔피언이 플레이한 게임수, 챔피언이 플레이해서 이긴 게임수]
"""
game_count = 0
win_count = 0
teamIds = ['100', '200']
for gameId in games.keys():
for teamId in teamIds:
if champion_id in games[gameId]['pick'][teamId]:
game_count += 1
if games[gameId]['win'][teamId] == 'Win':
win_count += 1
return [safe_divide(win_count, game_count), game_count, win_count]
champion_win_rate(498)
champion_win_rates = pd.DataFrame()
for champion_id in champion_id_list:
champion_win_rates = champion_win_rates.append([[champion_id,
champion_win_rate(champion_id)[0],
champion_win_rate(champion_id)[1],
champion_win_rate(champion_id)[2]]])
champion_win_rates.columns = ['champion_id', 'champion_win_rate', 'game_count', 'champion_win_count']
champion_win_rates.reset_index(inplace=True)
del champion_win_rates['index']
champion_win_rates.sort_values('champion_win_rate', ascending=False)
# # Evidence( = P(E))
# ## P(E) = P(E|H) P(H) + P(E|~ H) P(~ H)
#
#
def without_pick_champion_win_rate(my_champion ,my_team_champions, opponent_team_champions):
"""
내가 고른 챔피언이 없는 게임들 중에서 우리 팀과 적 팀이 고른 챔피언이 속한 게임의 승률.
항상 우리 팀이 고른 챔피언 숫자가 적을 경우 픽하는 상황으로 한정
실 게임에서는 동시에 2명이 픽 하는 경우, 다른 summoner가 먼저 픽 할 경우 고른 챔피언 숫자가 우리팀과 적팀이 같으나 여기선 무시
:param my_champion: 내가 고른 championId int
:param my_team_champions: 우리 팀의 championId list
:param opponent_team_champions: 적 팀의 championId list
:return: [승률, 플레이한 게임수, 플레이해서 이긴 게임수]
"""
# 같은팀 두명이 동시에 고르는 경우 내가 픽 했더라도 팀원은 아직 고르지 않았다는 가정
if my_champion in my_team_champions:
print('다른 챔피언을 선택하세요. 이미 픽된 챔피언입니다.')
return [0, 0, 0]
else:
if my_champion in opponent_team_champions:
print('다른 챔피언을 선택하세요. 이미 픽된 챔피언입니다.')
return [0, 0, 0]
else:
teamIds = ['100', '200']
for gameId in games.keys():
# 내가 고른 챔피언이 없는 경우만 뽑음
if champion_id not in games[gameId]['pick'][teamIds[0]] + games[gameId]['pick'][teamIds[1]]:
if (len(my_team_champions), len(opponent_team_champions)) in [(0,1),(1,2),(2,3),(3,4),(4,5)]:
game_count = 0
win_count = 0
for gameId in games.keys():
for teamId in teamIds:
# 우리 팀이 뽑은 챔피언이 games안에 모두 있는지 확인
if len(set(games[gameId]['pick'][teamId]) - set(my_team_champions)) == (
5 - len(my_team_champions)):
# 적 팀이 뽑은 챔피언이 반대 팀에 있는지 확인
if len(set(games[gameId]['pick'][list(set(teamIds) - set([teamId]))[0]]) -
set(opponent_team_champions)) == (5 - len(opponent_team_champions)):
# 우리 팀과 적 팀이 뽑은 챔피언이 모두 있는 경우는 경기수 증가
game_count += 1
if games[gameId]['win'][teamId] == 'Win':
# 우리 팀이 이긴 경우 승수 추가
win_count += 1
return [safe_divide(win_count, game_count), game_count, win_count]
else:
return print('''올바른 Input 값을 넣으세요.
Only (len(my_team_champions), len(opponent_team_champions)) in [(0,1), (1,2), (2,3), (3,4), (4,5)] is allowed.''')
without_pick_champion_win_rate('498', [], [59])
without_pick_champion_win_rate('498', [24, 56], [59, 110])
without_pick_champion_win_rate('498', [24], [59, 110])
# 한계..
# # likelihood( = P(E|H))
def pick_champion_win_rate(my_champion ,my_team_champions, opponent_team_champions):
"""
내가 고른 챔피언이 있는 게임들 중에서 우리 팀과 적 팀이 고른 챔피언이 속한 게임의 승률
항상 우리 팀이 고른 챔피언 숫자가 적을 경우 픽하는 상황으로 한정
실 게임에서는 동시에 2명이 픽 하는 경우, 다른 summoner가 먼저 픽 할 경우 고른 챔피언 숫자가 우리팀과 적팀이 같으나 여기선 무시
:param my_champion: 내가 고른 championId int
:param my_team_champions: 우리 팀의 championId list
:param opponent_team_champions: 적 팀의 championId list
:return: [승률, 플레이한 게임수, 플레이해서 이긴 게임수]
"""
# 같은팀 두명이 동시에 고르는 경우 내가 픽 했더라도 팀원은 아직 고르지 않았다는 가정
if my_champion in my_team_champions:
print('다른 챔피언을 선택하세요. 이미 픽된 챔피언입니다.')
return [0, 0, 0]
else:
if my_champion in opponent_team_champions:
print('다른 챔피언을 선택하세요. 이미 픽된 챔피언입니다.')
return [0, 0, 0]
else:
if (len(my_team_champions) + 1, len(opponent_team_champions)) in [(1,1),(2,2),(3,3),(4,4),(5,5)]:
new_my_team_champions = my_team_champions + [my_champion]
game_count = 0
win_count = 0
teamIds = ['100', '200']
for gameId in games.keys():
for teamId in teamIds:
# 내가 새로 뽑을 챔피언과 우리 팀이 뽑은 챔피언이 games안에 모두 있는지 확인
if len(set(games[gameId]['pick'][teamId]) - set(new_my_team_champions)) == \
(5 - len(new_my_team_champions)):
# 적 팀이 뽑은 챔피언이 반대 팀에 있는지 확인
if len(set(games[gameId]['pick'][list(set(teamIds) - set([teamId]))[0]]) -
set(opponent_team_champions)) == (5 - len(opponent_team_champions)):
# 우리 팀과 적 팀이 뽑은 챔피언이 모두 있는 경우는 경기수 증가
game_count += 1
if games[gameId]['win'][teamId] == 'Win':
# 우리 팀이 이긴 경우 승수 추가
win_count += 1
return [safe_divide(win_count, game_count), game_count, win_count]
else:
return print('''올바른 Input 값을 넣으세요.
Only (len(my_team_champions), len(opponent_team_champions)) in [(0,1), (1,2), (2,3), (3,4), (4,5)] is allowed.''')
pick_champion_win_rate(498, [], [240])
pick_champion_win_rate(90, [113, 90, 63], [59, 110, 80, 134])
pick_champion_win_rate(59, [113, 90, 63], [59, 110, 80, 134])
pick_champion_win_rate(59, [113, 90, 63, 100], [110, 80, 134])
# # Posterior probability ( = P(H|E))
#
# ## Expect win rate
def expect_win_rate(my_champion, my_team_champions, opponent_team_champions):
"""
내가 특정 챔피언을 고를 경우 우리 팀과 적 팀이 선택한 챔피언들을 바탕으로 예상되는 승률
:param my_champion: 내가 고른 championId int
:param my_team_champions: 우리 팀의 championId list
:param opponent_team_champions: 적 팀의 championId list
:return: 예상 승률
"""
# P(E|H) * P(H)
upper = (pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
champion_win_rate(my_champion)[0])
# P(E) = P(E|H) * P(H) + P(E|~H) * P(~H)
lower = (pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
champion_win_rate(my_champion)[0]) + \
(without_pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
(1 - champion_win_rate(my_champion)[0]))
if np.logical_or(lower==0, upper==0):
return print('계산불가. 샘플수 부족...')
else:
# P(H|E)= P(E|H) * P(H) / P(E)
return safe_divide(upper, lower)
my_champion = 59
my_team_champions = [18]
opponent_team_champions = [498, 81]
print('prior probability:', champion_win_rate(my_champion)[0])
print('likelihood:', pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0])
print('evidence:', round((pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
champion_win_rate(my_champion)[0]) + \
(without_pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
(1 - champion_win_rate(my_champion)[0])), 4))
print('posterior probability:', expect_win_rate(my_champion, my_team_champions, opponent_team_champions))
my_champion = 266
my_team_champions = [18]
opponent_team_champions = [498, 81]
print('prior probability:', champion_win_rate(my_champion)[0])
print('likelihood:', pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0])
print('evidence:', round((pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
champion_win_rate(my_champion)[0]) + \
(without_pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
(1 - champion_win_rate(my_champion)[0])), 4))
print('posterior probability:', expect_win_rate(my_champion, my_team_champions, opponent_team_champions))
my_champion = 266
my_team_champions = []
opponent_team_champions = [1]
print('prior probability:', champion_win_rate(my_champion)[0])
print('likelihood:', pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0])
print('evidence:', round((pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
champion_win_rate(my_champion)[0]) + \
(without_pick_champion_win_rate(my_champion, my_team_champions, opponent_team_champions)[0] *
(1 - champion_win_rate(my_champion)[0])), 4))
print('posterior probability:', expect_win_rate(my_champion, my_team_champions, opponent_team_champions))
# # Recommend champion
df_champion.tail()
def championName_to_championId(championName):
# 챔피언 이름을 영문으로 넣으면 champion key 출력
return df_champion.set_index('id').to_dict()['key'][championName]
championName_to_championId('Zed')
def recommend_championId(my_team_champions, opponent_team_champions):
"""
내가 픽 할 상황에서 우리편과 상대편의 픽 된 챔피언 리스트를 넣으면 승률이 가장 높을 것으로 예상되는 상위 5개 챔피언을 보여준다.
:param my_team_champions: 우리 팀의 championId list
:param opponent_team_champions: 적 팀의 championId list
:return: df
"""
df = pd.DataFrame()
for champion_Id in champion_win_rates[champion_win_rates.game_count >= 300].champion_id:
df = df.append([[champion_Id, expect_win_rate(champion_Id, my_team_champions, opponent_team_champions)]])
df.columns = ['key', 'win_rate']
df.reset_index(inplace=True)
del df['index']
df.sort_values('win_rate', ascending=False, inplace=True)
df_champions = pd.read_csv('champion.csv')
result = pd.merge(df, df_champions, on ='key', how='inner')
result = result.drop(['Unnamed: 0'], axis=1)
return result.head()
recommend_championId([202], [497, 498])
|
src/ipynb/analysis_bayes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow_py3]
# language: python
# name: conda-env-tensorflow_py3-py
# ---
# +
"""
Create a model of classification SVM (linear)
base no example dataset 'Iris'
Refer to: https://www.analyticsvidhya.com/blog/2015/10/understaing-support-vector-machine-example-code/
"""
# import requred library and modlue
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm, datasets
# +
# print Iris dataset used in this project desc to screen
# print(iris.DESCR)
# -
# 导入虹膜数据集
iris = datasets.load_iris()
# 只使有前两列中的数据(avoid this ugly slicing by using a two-dim dataset)
X = iris.data[:, :2]
# 样本数据的标签(label)
y = iris.target
# +
# Classificador SVM com kernel Linear
svc = svm.SVC(kernel='linear', C=1,gamma='auto').fit(X, y)
"""
备注:
1. 在分类任务中,(C和γ)是提供模型决策边界的参数组合,使得构建的模型允许有更大的灵活性;
2.也可以了解其他的内核(比比如RBF,Sigmoid),他们允许在更复杂的数据集的情况下有更好的效果;
"""
# -
# 准备可视化工作
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
h = (x_max / x_min)/100
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
# +
# 画出边界,
plt.subplot(1, 1, 1)
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.title('SVC com kernel linear')
plt.show()
# -
|
supervisioned/SVM_classifier_used_scikitlearn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + dc={"key": "4"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 1. The World Bank's international debt data
# <p>It's not that we humans only take debts to manage our necessities. A country may also take debt to manage its economy. For example, infrastructure spending is one costly ingredient required for a country's citizens to lead comfortable lives. <a href="https://www.worldbank.org">The World Bank</a> is the organization that provides debt to countries.</p>
# <p>In this notebook, we are going to analyze international debt data collected by The World Bank. The dataset contains information about the amount of debt (in USD) owed by developing countries across several categories. We are going to find the answers to questions like: </p>
# <ul>
# <li>What is the total amount of debt that is owed by the countries listed in the dataset?</li>
# <li>Which country owns the maximum amount of debt and what does that amount look like?</li>
# <li>What is the average amount of debt owed by countries across different debt indicators?</li>
# </ul>
# <p><img src="https://assets.datacamp.com/production/project_754/img/image.jpg" alt></p>
# <p>The first line of code connects us to the <code>international_debt</code> database where the table <code>international_debt</code> is residing. Let's first <code>SELECT</code> <em>all</em> of the columns from the <code>international_debt</code> table. Also, we'll limit the output to the first ten rows to keep the output clean.</p>
# + dc={"key": "4"} tags=["sample_code"] language="sql"
# postgresql:///international_debt
#
# select* from international_debt limit 10
# + dc={"key": "12"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 2. Finding the number of distinct countries
# <p>From the first ten rows, we can see the amount of debt owed by <em>Afghanistan</em> in the different debt indicators. But we do not know the number of different countries we have on the table. There are repetitions in the country names because a country is most likely to have debt in more than one debt indicator. </p>
# <p>Without a count of unique countries, we will not be able to perform our statistical analyses holistically. In this section, we are going to extract the number of unique countries present in the table. </p>
# + dc={"key": "12"} tags=["sample_code"] language="sql"
# SELECT
# count(distinct(country_name)) AS total_distinct_countries
# FROM international_debt;
# + dc={"key": "20"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 3. Finding out the distinct debt indicators
# <p>We can see there are a total of 124 countries present on the table. As we saw in the first section, there is a column called <code>indicator_name</code> that briefly specifies the purpose of taking the debt. Just beside that column, there is another column called <code>indicator_code</code> which symbolizes the category of these debts. Knowing about these various debt indicators will help us to understand the areas in which a country can possibly be indebted to. </p>
# + dc={"key": "20"} tags=["sample_code"] language="sql"
# select distinct(indicator_code) as distinct_debt_indicators
# from international_debt order by distinct_debt_indicators;
# + dc={"key": "28"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 4. Totaling the amount of debt owed by the countries
# <p>As mentioned earlier, the financial debt of a particular country represents its economic state. But if we were to project this on an overall global scale, how will we approach it?</p>
# <p>Let's switch gears from the debt indicators now and find out the total amount of debt (in USD) that is owed by the different countries. This will give us a sense of how the overall economy of the entire world is holding up.</p>
# + dc={"key": "28"} tags=["sample_code"] language="sql"
# SELECT
# round(sum(debt)/1000000, 2) as total_debt
# FROM international_debt;
# + dc={"key": "36"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 5. Country with the highest debt
# <p>"Human beings cannot comprehend very large or very small numbers. It would be useful for us to acknowledge that fact." - <a href="https://en.wikipedia.org/wiki/Daniel_Kahneman"><NAME></a>. That is more than <em>3 million <strong>million</strong></em> USD, an amount which is really hard for us to fathom. </p>
# <p>Now that we have the exact total of the amounts of debt owed by several countries, let's now find out the country that owns the highest amount of debt along with the amount. <strong>Note</strong> that this debt is the sum of different debts owed by a country across several categories. This will help to understand more about the country in terms of its socio-economic scenarios. We can also find out the category in which the country owns its highest debt. But we will leave that for now. </p>
# + dc={"key": "36"} tags=["sample_code"] language="sql"
# SELECT
# country_name,
# sum(debt) as total_debt
# FROM international_debt
# GROUP BY 1
# ORDER BY 2 desc
# limit 1;
# + dc={"key": "44"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 6. Average amount of debt across indicators
# <p>So, it was <em>China</em>. A more in-depth breakdown of China's debts can be found <a href="https://datatopics.worldbank.org/debt/ids/country/CHN">here</a>. </p>
# <p>We now have a brief overview of the dataset and a few of its summary statistics. We already have an idea of the different debt indicators in which the countries owe their debts. We can dig even further to find out on an average how much debt a country owes? This will give us a better sense of the distribution of the amount of debt across different indicators.</p>
# + dc={"key": "44"} tags=["sample_code"] language="sql"
# SELECT
# indicator_code AS debt_indicator,
# indicator_name,
# avg(debt) as average_debt
# FROM international_debt
# GROUP BY debt_indicator,indicator_name
# ORDER BY 3 desc
# limit 10;
# + dc={"key": "52"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 7. The highest amount of principal repayments
# <p>We can see that the indicator <code>DT.AMT.DLXF.CD</code> tops the chart of average debt. This category includes repayment of long term debts. Countries take on long-term debt to acquire immediate capital. More information about this category can be found <a href="https://datacatalog.worldbank.org/principal-repayments-external-debt-long-term-amt-current-us-0">here</a>. </p>
# <p>An interesting observation in the above finding is that there is a huge difference in the amounts of the indicators after the second one. This indicates that the first two indicators might be the most severe categories in which the countries owe their debts.</p>
# <p>We can investigate this a bit more so as to find out which country owes the highest amount of debt in the category of long term debts (<code>DT.AMT.DLXF.CD</code>). Since not all the countries suffer from the same kind of economic disturbances, this finding will allow us to understand that particular country's economic condition a bit more specifically. </p>
# + dc={"key": "52"} tags=["sample_code"] language="sql"
# SELECT
# country_name,
# indicator_name
# FROM international_debt
# WHERE debt = (SELECT
# max(debt)
# FROM international_debt
# where indicator_code='DT.AMT.DLXF.CD' );
# + dc={"key": "60"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 8. The most common debt indicator
# <p>China has the highest amount of debt in the long-term debt (<code>DT.AMT.DLXF.CD</code>) category. This is verified by <a href="https://data.worldbank.org/indicator/DT.AMT.DLXF.CD?end=2018&most_recent_value_desc=true">The World Bank</a>. It is often a good idea to verify our analyses like this since it validates that our investigations are correct. </p>
# <p>We saw that long-term debt is the topmost category when it comes to the average amount of debt. But is it the most common indicator in which the countries owe their debt? Let's find that out. </p>
# + dc={"key": "60"} tags=["sample_code"] language="sql"
# select indicator_code ,count(indicator_code) as indicator_count
# from international_debt group by indicator_code order by 2 desc, 1 desc limit 20;
# + dc={"key": "68"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 9. Other viable debt issues and conclusion
# <p>There are a total of six debt indicators in which all the countries listed in our dataset have taken debt. The indicator <code>DT.AMT.DLXF.CD</code> is also there in the list. So, this gives us a clue that all these countries are suffering from a common economic issue. But that is not the end of the story, a part of the story rather. </p>
# <p>Let's change tracks from <code>debt_indicator</code>s now and focus on the amount of debt again. Let's find out the maximum amount of debt across the indicators along with the respective country names. With this, we will be in a position to identify the other plausible economic issues a country might be going through. By the end of this section, we will have found out the debt indicators in which a country owes its highest debt. </p>
# <p>In this notebook, we took a look at debt owed by countries across the globe. We extracted a few summary statistics from the data and unraveled some interesting facts and figures. We also validated our findings to make sure the investigations are correct.</p>
# + dc={"key": "68"} tags=["sample_code"] language="sql"
# select country_name,indicator_code,max(debt) as maximum_debt
# from international_debt group by country_name,indicator_code
# order by maximum_debt desc limit 10;
|
SQL Projects/Analyze International Debt Statistics/notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
foo = ['a', 'b', 'c', 'd', 'e']
print(random.choice(foo))
# -
print(random.choice(foo))
random.randint(1,100)
me=random.randint(1,100)
while True:
your_number = float(input("나는 얼마일까요 ?"))
print(your_number)
if your_number>me:
print("너무 커요 다시요")
elif your_number<me:
print("너무 작아요 다시요")
else:
print("잘했어요")
break;
|
python/pythoncodes/Untitled2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.1 64-bit (''env'': venv)'
# name: python36164bitenvvenv5ed069fb11d5433d9d9988a83a718c28
# ---
# # Some example
#
# > This is an example
# hide
# enable autoreload
# %load_ext autoreload
# %autoreload 2
from testnote import data
data.get_months()
data.get_names()
# Here's some code using the shared functions. Check how to use `get_names` and `get_months`.
for name in data.get_names():
for month in data.get_months():
print(f"{name} - {month}")
|
02_example1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def readfile(filename):
file = open(filename, 'rt')
original_text = file.read()
file.close()
return original_text
def writefile(filename, content):
file = open(filename, 'w')
file.write(str(content))
file.close()
def preprocess_data(text):
# remove punctuations !"#$%&'()*+,-./:;<=>?@[\]^_`{|}~
punctuation_list = ["!","\"","#","$","%","&","'","(",")","*","+",",","-",".","/",":",";","<","=",">","?","@","[","\\","]","^","_","`","{","|","}","~","–","€","•","«","»","’","“","”","£"]
for punctuation in punctuation_list:
text = text.replace(punctuation, "")
# remove numbers
RE_NUMBERS = re.compile('[0-9]+')
text = RE_NUMBERS.sub(r'', text)
# remove emojis
RE_EMOJI = re.compile('[\U00010000-\U0010ffff]', flags=re.UNICODE)
text = RE_EMOJI.sub(r'', text) # ⛈ corcra❄ gorm❄
# substitute multiple spaces with a single space
text = re.sub(' +', ' ', text)
# trim leading and trailing spaces
text = text.strip()
# normalize text to lower case
text = text.lower()
return text
# +
import re
text = readfile('train.txt')
# text = text.replace("\n", " ")
text = preprocess_data(text)
writefile("cleaned_train.txt", text)
# -
keyword_pair_dict = {
"a|á": { "keywords": ["a","á"], "probability": 0 },
"ais|áis": { "keywords": ["ais","áis"], "probability": 0 },
"aisti|aistí": { "keywords": ["aisti","aistí"], "probability": 0 },
"ait|áit": { "keywords": ["ait","áit"], "probability": 0 },
"ar|ár": { "keywords": ["ar","ár"], "probability": 0 },
"arsa|ársa": { "keywords": ["arsa","ársa"], "probability": 0 },
"ban|bán": { "keywords": ["ban","bán"], "probability": 0 },
"cead|céad": { "keywords": ["cead","céad"], "probability": 0 },
"chas|chás": { "keywords": ["chas","chás"], "probability": 0 },
"chuig|chúig": { "keywords": ["chuig","chúig"], "probability": 0 },
"dar|dár": { "keywords": ["dar","dár"], "probability": 0 },
"do|dó": { "keywords": ["do","dó"], "probability": 0 },
"gaire|gáire": { "keywords": ["gaire","gáire"], "probability": 0 },
"i|í": { "keywords": ["i","í"], "probability": 0 },
"inar|inár": { "keywords": ["inar","inár"], "probability": 0 },
"leacht|léacht": { "keywords": ["leacht","léacht"], "probability": 0 },
"leas|léas": { "keywords": ["leas","léas"], "probability": 0 },
"mo|mó": { "keywords": ["mo","mó"], "probability": 0 },
"na|ná": { "keywords": ["na","ná"], "probability": 0 },
"os|ós": { "keywords": ["os","ós"], "probability": 0 },
"re|ré": { "keywords": ["re","ré"], "probability": 0 },
"scor|scór": { "keywords": ["scor","scór"], "probability": 0 },
"te|té": { "keywords": ["te","té"], "probability": 0 },
"teann|téann": { "keywords": ["teann","téann"], "probability": 0 },
"thoir|thóir": { "keywords": ["thoir","thóir"], "probability": 0 },
}
# +
unigram_count_dict = {}
words = text.split()
num_all_words = len(words)
for word in words:
if word in unigram_count_dict:
unigram_count_dict[word] += 1
else:
unigram_count_dict[word] = 1
K = 3
V = len(unigram_count_dict)
N = len(words)
for key in keyword_pair_dict:
keyword1 = keyword_pair_dict[key]["keywords"][0]
keyword2 = keyword_pair_dict[key]["keywords"][1]
# +
# count bigram
bigram_count_dict = {}
for i in range(len(words)):
if i == 0:
# a word with nothing preceding
preceding_word = "^";
else:
# normal pairs of words
preceding_word = words[i-1];
bigram = preceding_word + " " + words[i]
if bigram in bigram_count_dict:
bigram_count_dict[bigram] += 1
else:
bigram_count_dict[bigram] = 1
# count when the word is the last word of the sentence
bigram = words[-1] + " $"
if bigram in bigram_count_dict:
bigram_count_dict[bigram] += 1
else:
bigram_count_dict[bigram] = 1
# +
# print(unigram_count_dict["thóir"] / (unigram_count_dict["thóir"] + unigram_count_dict["thoir"]))
# -
# store unigram probabilities to a dictionary
for key in keyword_pair_dict:
word_1 = keyword_pair_dict[key]["keywords"][0]
word_2 = keyword_pair_dict[key]["keywords"][1]
prob_word_1 = unigram_count_dict[word_1]
prob_word_2 = unigram_count_dict[word_2]
normalized_prob_word_1 = prob_word_1 / (prob_word_1 + prob_word_2)
keyword_pair_dict[key]["probability"] = normalized_prob_word_1
def generate_submission_unigram():
test_text = readfile('test.txt')
test_sentences = test_text.split("\n");
test_word_choices = []
answer_list = ["Id,Expected"]
running_number = 1
for sentence in test_sentences:
start_index = sentence.find("{")
end_index = sentence.find("}")
word_choice = sentence[start_index+1:end_index]
if(word_choice):
prob = keyword_pair_dict[word_choice]["probability"]
answer_list.append("{},{}".format(running_number, prob))
running_number += 1
answer = "\n".join(answer_list)
return answer
# +
# submission = generate_submission_unigram()
# writefile('submission7.csv', submission)
# -
def calculate_bigram_prob(sentence):
prob_word_choice_1 = 0
start_index = sentence.find("{")
end_index = sentence.find("}")
word_choice = sentence[(start_index+1):end_index] # "thoir|thóir"
if word_choice:
[word_choice_1, word_choice_2] = word_choice.split('|'); # word_choice_1 = "thoir" and word_choice_2 = "thóir"
words = sentence.split();
index = words.index("{" + word_choice + "}")
if index == 0:
prev_word = "^"
next_word = words[1]
elif index == len(words)-1:
prev_word = words[index - 1]
next_word = "$"
else:
prev_word = words[index - 1]
next_word = words[index + 1]
# E.g. <NAME> agus Bróigín ar {thoir|thóir} réalt-ainmhithe in éineacht le Pegaso.
# p1 = P(thoir|ar) * P(réaltainmhithe|thoir)
# c1 = count("ar thoir") * counbt("thoir réaltainmhithe")
# p2 = P(thòir|ar) * P(réaltainmhithe|thòir)
# c2 = count("ar thòir") * count("thòir réaltainmhithe")
# P(thoir) = p1/(p1+p2)
# P(thoir) = c1/(c1+c2)
word_1_bigram_1 = prev_word + " " + word_choice_1 # "ar thoir"
word_1_bigram_2 = word_choice_1 + " " + next_word # "thoir réaltainmhithe"
word_2_bigram_1 = prev_word + " " + word_choice_2 # "ar thòir"
word_2_bigram_2 = word_choice_2 + " " + next_word # "thòir réaltainmhithe"
count_word_1_bigram_1 = bigram_count_dict[word_1_bigram_1] if word_1_bigram_1 in bigram_count_dict else 0
count_word_1_bigram_2 = bigram_count_dict[word_1_bigram_2] if word_1_bigram_2 in bigram_count_dict else 0
count_word_2_bigram_1 = bigram_count_dict[word_2_bigram_1] if word_2_bigram_1 in bigram_count_dict else 0
count_word_2_bigram_2 = bigram_count_dict[word_2_bigram_2] if word_2_bigram_2 in bigram_count_dict else 0
count_prev_word = unigram_count_dict[prev_word] if prev_word in unigram_count_dict else 0
count_keyword_1 = unigram_count_dict[word_choice_1] if word_choice_1 in unigram_count_dict else 0
count_keyword_2 = unigram_count_dict[word_choice_2] if word_choice_2 in unigram_count_dict else 0
p_word_1_bigram_1 = (count_word_1_bigram_1 + K) / (count_prev_word + (K*V)) # "ar thoir"
p_word_1_bigram_2 = (count_word_1_bigram_2 + K) / (count_keyword_1 + (K*V)) # "thoir réaltainmhithe"
p_word_2_bigram_1 = (count_word_2_bigram_1 + K) / (count_prev_word + (K*V)) # "ar thòir"
p_word_2_bigram_2 = (count_word_2_bigram_2 + K) / (count_keyword_2 + (K*V)) # "thòir réaltainmhithe"
p1 = (p_word_1_bigram_1 * p_word_1_bigram_2)
p2 = (p_word_2_bigram_1 * p_word_2_bigram_2)
prob_word_choice_1 = p1 / (p1 + p2)
return prob_word_choice_1
def generate_submission_bigram():
test_text = readfile('test.txt')
test_sentences = test_text.split("\n");
test_word_choices = []
answer_list = ["Id,Expected"]
running_number = 1
for test_sentence in test_sentences:
if test_sentence:
prob = calculate_bigram_prob(test_sentence)
answer_list.append("{},{:.20f}".format(running_number, prob))
running_number += 1
answer = "\n".join(answer_list)
return answer
# +
# print(generate_submission_bigram())
writefile('submission_bigram_add_0.5_smoothing_11.csv', generate_submission_bigram())
|
irish-language-modeling/bigram_with_smoothing_irish_language_modeling .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Method: RC
# # Dataset: Lorenz-96, F = 8
# # Purpose: Uncertainty Quantification - Deep Ensemble
# # 1. Set-up
# +
# GPU
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "3"
# Package
import sys
sys.path.append("../..")
# +
from create_data import load_data
from utils import * # Number of testing samples
import numpy as np
import matplotlib.pyplot as plt
from time import time
from scipy import sparse
import jax
import jax.numpy as jnp
from jax import value_and_grad
from jax.numpy import tanh
from jax.example_libraries import optimizers
# -
train, test = load_data("Lorenz 96, F = 8", "../../data/lorenz8", 0.5)
np.random.seed(1)
train.data = train.data + np.random.normal(0, 1e-1, train.data.shape)
print(f"Train size: {train.data.shape}")
print(f"Test size: {test.data.shape}")
# **Create test set**
L_forecast_test = 400 # steps to forecast forward (when testing)
# +
np.random.seed(1)
data_test = test.data
T_test, data_dim = data_test.shape
possible_idx = T_test - (L_forecast_test + 1) # minus number of steps forward, and the warm-up period
T_indices = np.random.randint(0, possible_idx, size = NUM_TEST)
t_past_batch = np.repeat(T_indices[:, None], WARM_UP_TEST, axis = 1).astype(int) # 200 warmup
t_pred_batch = (T_indices[:, None] + np.arange(1, 1 + L_forecast_test)[None, :].astype(int))
X_test = data_test[t_past_batch]
y_test = data_test[t_pred_batch]
# -
print(f"Test input size: {X_test.shape}") # Number of test points x input length x dim
print(f"Test output size: {y_test.shape}") # Number of test points x horizon x dim
# # 2. RC Implementation
def get_parameters(nn_size, connectivity, spec_radius, lambd,
seed, batch_size, num_epoch, lr_schedule = [1e-4],
early_stopping = EARLY_STOPPING):
"""
Returns trained parameters (beta, intercept) and hidden layer values
"""
def initialize_coef():
"""
Initializes W_in and W.
W_in size = nn_size x data_dim
W size = nn_size x nn_size
"""
start = time()
# Generate input -> hidden unit weights
W_in = 2 * (np.random.rand(nn_size, data_dim) - 0.5)
W_in = W_in / (4 * np.sqrt(data_dim))
# Generate hidden -> hidden unit weights
# Considers connectivity to make the matrix sparse
start_mat = time()
rows = np.concatenate([np.full(connectivity, i) for i in range(nn_size)])
cols = np.concatenate([np.random.choice(range(nn_size), size = connectivity, replace = False) for _ in range(nn_size)])
vals = np.random.uniform(low = -omega, high = omega, size = (nn_size * connectivity))
W = sparse.csr_matrix((vals, (rows, cols)), shape = (nn_size, nn_size))
end_mat = time()
print(f"W generated. Time taken: {end_mat - start_mat:.2f}s")
# Calculate eigenvalues for scaling of matrix
print("Calculating eigenvalue")
e_start = time()
eigenvals = sparse.linalg.eigs(W, which = "LM", return_eigenvectors = False, k = 1)
max_eigen = np.abs(eigenvals)
e_end = time()
print(f"Eigenvalue calculated. Time taken: {e_end - e_start:.2f}s")
# Scale matrix by spectral radius
W = W / max_eigen * spec_radius # scale the matrix W by its spectral radius
W = sparse.csr_matrix(W)
end = time()
print(f"W and W_in generated. Time taken: {end-start:.2f}s")
print()
return W_in, W
def generate_hidden_states(W_in, W):
"""
Generate hidden states (z) values
hidden_states size = data_size x nn_size
"""
start = time()
print("Generating z values...")
indiv_z = np.zeros(shape = nn_size)
hidden_states = np.zeros((train_size, nn_size))
for t in range(train_size):
indiv_z = (1 - alpha) * indiv_z + \
alpha * np.tanh(W_in @ x[t] + W @ indiv_z)
hidden_states[t, :] = indiv_z
end = time()
print(f"z values generated. Time taken: {end-start:.2f}s")
return hidden_states
def mse(y, y_pred):
return jnp.mean((y_pred - y)**2)
@jax.jit
def mse_loss(params, x, y):
"""
returns mean squared error with ridge penalty
"""
beta, intercept = params
pred = x @ beta + intercept
return mse(pred, y) + np.mean(beta**2) * lambd / 2 + np.mean(intercept**2) * lambd / 2
def validation_loss(params, x_val, y_val):
beta, intercept = params
num_data_test, trans, data_dim = x_val.shape # testing ex, # steps used (transient), dim of data
def prediction(inp):
"""
Returns the mean of one of the testing input
mean will be a length_to_test x data_dim vector
"""
z = np.zeros((nn_size // 2, ))
for i in range(trans):
z = (1 - alpha) * z + alpha * np.tanh(W_in @ inp[i] + W @ z)
mus = []
stddevs = []
x = beta.T @ np.concatenate([z, z**2]) + intercept # output / input_of_next | size = dim_data
mus.append(x)
for _ in range(L_forecast_test - 1):
z = (1 - alpha) * z + alpha * np.tanh(W_in @ x + W @ z)
x = beta.T @ np.concatenate([z, z**2]) + intercept # output / input_of_next
mus.append(x)
return mus
mean_list = []
sd_list = []
for i in range(num_data_test):
pred = prediction(x_val[i])
mean_list.append(pred)
return mse(mean_list, y_val)
def training(x, y):
"""
Trains regression of y~x using SGD.
Returns parameters (beta, intercept) where
beta, intercept -> weights to determine the mean
beta size = nn_size x data_dim
intercept = data_dim (will be added for each training data)
should predict a mu with train_size x data_dim (\mu per dimension per datapoint)
and a sigma with train_size x 1 (single \sigma for all dimensions per datapoint)
"""
@jax.jit
def step(opt_state, x, y):
params = get_params(opt_state)
value, g = value_and_grad(mse_loss)(params, x, y)
opt_state = opt_update(0, g, opt_state)
return get_params(opt_state), opt_state, value
start = time()
# Plot loss
loss_train_traj = []
loss_train_all_traj = []
# Init parameters
beta = np.random.normal(0, 1 / np.sqrt(nn_size), size = (nn_size, data_dim))
intercept = np.random.normal(0, 1 / np.sqrt(nn_size * 2), size = (data_dim, ))
t_size = int(1. * train_size)
# v_size = train_size - t_size
# t_start_val = np.linspace(t_size, train_size - (L_forecast_test + 2), NUM_TEST // 2, dtype = int)
# t_past_batch_val = np.repeat(t_start_val[:, None], WARM_UP_TEST, axis = 1).astype(int)
# t_pred_batch_val = (t_start_val[:,None] + np.arange(1,1+L_forecast_test)[None,:]).astype(int)
# x_val = copy_x[t_past_batch_val]
# y_val = copy_y[t_pred_batch_val]
overall_best_mse = 9999999
for i, lr in enumerate(lr_schedule):
opt_init, opt_update, get_params = optimizers.adam(step_size = lr)
opt_state = opt_init([beta, intercept])
# For early stopping
best_state = opt_state
counter = 0
best_val_loss = 9999999
for epoch in range(num_epoch[i]):
e_start = time()
T_indices = np.arange(train_size)
np.random.shuffle(T_indices)
loss_epoch_train = []
for k in range(t_size // batch_size + 1):
t_start = T_indices[np.arange(k * batch_size, (k+1) * batch_size).astype(int) % len(T_indices)]
x_batch = x[t_start]
y_batch = y[t_start]
params, opt_state, l = step(opt_state, x_batch, y_batch)
loss_epoch_train.append(l)
loss_train_all_traj += loss_epoch_train
mse_train = np.mean(loss_epoch_train)
loss_train_traj.append(mse_train)
e_end = time()
if mse_train < best_val_loss:
best_val_loss = mse_train
counter = 0
best_state = opt_state
else:
counter += 1
if (epoch + 1) % 10 == 0:
print(f"Epoch {epoch + 1}: Train time = {e_end - e_start:.2f} | Train Loss = {mse_train:.7f}", end = " ")
# when_to_evaluate = 199 if i == 0 else 49
# if epoch >= when_to_evaluate:
# # Validation
# val_start = time()
# mse_val = validation_loss(params, x_val, y_val)
# if best_val_loss > mse_val: # Improvement
# counter = 0
# best_val_loss = mse_val
# best_state = opt_state
# else:
# counter += 1
# val_end = time()
# print(f"| Val time: {val_end - val_start:2f} | Val loss: {mse_val:.7f}", end = "")
print()
if counter == early_stopping:
print(f"EARLY STOPPING. Epoch {epoch + 1}: Train loss = {mse_train:.7f}")
break
print(f"Best Training MSE: {best_val_loss:.7f}")
if best_val_loss < overall_best_mse:
print("IMPROVED VALIDATION MSE")
overall_best_mse = best_val_loss
overall_best_state = best_state
beta, intercept = get_params(overall_best_state)
print()
end = time()
print(f"Total time: {end - start:.2f}")
return get_params(overall_best_state) # beta, intercept
start = time()
x, y = train.data[:-1], train.data[1:]
copy_x, copy_y = x, y
train_size, data_dim = x.data.shape
np.random.seed(seed)
W_in, W = initialize_coef()
z = generate_hidden_states(W_in, W)
# Want to regression Y ~ X ==> Y ~ [z, z**2]
final_y = y[transient:]
final_z = z[transient:]
print("Concatenating z with z**2", end = " "); concat_start = time()
final_z = np.concatenate([final_z, final_z**2], axis = 1) # shape: train_size x (nn_size*2)
concat_end = time()
print(f"Contenation complete. Time taken: {concat_end-concat_start:.2f}s", end = "\n\n")
train_size, nn_size = final_z.shape
params = training(final_z, final_y)
end = time()
print(f"Complete. Time taken: {end - start:.2f}s")
return params, (final_z, W_in, W)
def get_test_pred(data_test, nn_size, params, W_in, W):
beta, intercept = params
num_data_test, trans, data_dim = data_test.shape # testing ex, # steps used (transient), dim of data
def prediction(inp):
"""
Returns the mean of one of the testing input
mean will be a length_to_test x data_dim vector
"""
z = np.zeros((nn_size, ))
for i in range(trans):
z = (1 - alpha) * z + alpha * np.tanh(W_in @ inp[i] + W @ z)
mus = []
stddevs = []
x = beta.T @ np.concatenate([z, z**2]) + intercept # output / input_of_next | size = dim_data
mus.append(x)
for _ in range(L_forecast_test - 1):
z = (1 - alpha) * z + alpha * np.tanh(W_in @ x + W @ z)
x = beta.T @ np.concatenate([z, z**2]) + intercept # output / input_of_next
mus.append(x)
return mus
start = time()
mean_list = []
sd_list = []
for i in range(num_data_test):
pred = prediction(data_test[i])
mean_list.append(pred)
if (i+1) % 10 == 0:
print(f"{(i+1) / num_data_test * 100:.2f}% done")
end = time()
print(f"Testing complete. Time taken: {end - start:.2f}")
return np.array(mean_list)
# # 3. Best Parameters
# +
nn_size = 12000
ridge_penalty = 1e-6
spec_radius = 0.1
connectivity = 4
lr_list = [1e-4]
epoch_list = [300]
transient = 200 # points to ignore to allow system to stabilise
omega = 1 # scale of the values of matrix W
alpha = 1 # hidden state memory
b_size = 200
# -
# # 4. Ensemble
res_folder = os.path.join("results", "ensemble_noise")
def run_seed(seed):
"""
Runs the experiment with optimal parameters and saves the predictions into a file
"""
params, internal = get_parameters(nn_size, connectivity, spec_radius, lambd = ridge_penalty, seed = seed,
batch_size = b_size, num_epoch = epoch_list, lr_schedule = lr_list)
_, W_in, W = internal
mean_pred = get_test_pred(X_test, nn_size, params, W_in, W)
file_name = "mu_preds_" + str(seed) + ".pkl"
save_obj(mean_pred, res_folder, file_name)
# ## 4.1 Seed 2
# + tags=[]
run_seed(2)
# -
# ## 4.2 Seed 4
# + tags=[]
run_seed(4)
# -
# ## 4.3 Seed 6
run_seed(6)
# ## 4.4 Seed 8
run_seed(8)
# ## 4.5 Seed 42
# + tags=[]
run_seed(42)
# -
# ## 4.6 Compilation of means
mu_preds = []
for dirpath, dirnames, filenames in os.walk(res_folder):
for f in filenames:
mu_preds.append(load_obj(os.path.join(res_folder, f)))
mu_preds = np.array(mu_preds)
print(f"mean preds shape: {mu_preds.shape}")
# # 5. Analyze results
# ## 5.1 MSE
mixture_pred_all_mean = mu_preds.mean(axis = 0)
res_single = PointExperimentResultLyapunov(mixture_pred_all_mean - y_test, "lorenz")
res_single.plot_rmse(save_name = "RC Noise Deep Ensemble Horizon")
print()
res_single.get_loss()
# ## 5.2 Variance
# **Visualise for one dataset**
idx = 0
plt.plot(np.arange(L_forecast_test) / LORENZ_LT * 0.01, mu_preds.var(axis = 0)[idx].mean(axis = 1))
plt.grid("on")
plt.xlabel("Lyapunov Time")
plt.ylabel("Variance")
plt.savefig("RC Noise Deep Ensemble Variance.png", facecolor = "white", bbox_inches = "tight")
plt.show()
# ## 5.3 Negative Log LH
def neg_log_LH(mean_pred, sd_pred):
d = 40
constant_loss = d * np.log(2 * np.pi)
mu_loss = (mean_pred - y_test)**2
return 0.5 * (constant_loss + d * np.log(sd_pred) + (mu_loss / sd_pred**2)).mean(axis = (0, 2))
std_dev = mu_preds.std(axis = 0)
plt.plot(np.arange(L_forecast_test) / LORENZ_LT * 0.01, neg_log_LH(mixture_pred_all_mean, std_dev))
# plt.title("Negative Log LH against time")
plt.ylabel("Negative Log LH")
plt.xlabel("Lyapunov Time")
plt.grid("on")
plt.savefig("RC Noise Deep Ensemble NLL.png", facecolor = "white", bbox_inches = "tight")
plt.show()
print(f"Overall negative log LH: {neg_log_LH(mixture_pred_all_mean, std_dev).mean():.5f}")
|
3_uq/3_rc_noise/1_rc_deep_ensemble_noise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # API´s
# !pip install requests -q
import requests
# +
url = 'https://v6.exchangerate-api.com/v6/b30b64c2ef47f0501db155aa/latest/USD'
req = requests.get(url)
print(req.status_code)
# +
dados = req.json()
print(dados)
# -
valor_reais = int(input("Digite o valor em reais: "))
dolar = dados['conversion_rates']['BRL']
print(f'A conversão de {valor_reais}, equivale a : {(valor_reais / dolar):.2f} em dolar')
|
apis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Writing Valid Requests for NWIS
# The USGS National Water Information System (NWIS) is capable of handling a wide range of requests. A few features in Hydrofunctions are set up to help you write a successful request.
# First, import hydrofunctions.
import hydrofunctions as hf
# ## What can we specify?
# The NWIS can handle data requests that specify:
#
# - Where: we need to specify which stations we are interested in.
# - Service: the NWIS provides daily averages ('dv') and 'instantaneous values' ('iv')
# - When: we can specify a range of dates, a period of time before now, or just get the most recent observation.
# - What: we can specify which parameter we want, or just get everything collected at the site.
# - the data service we want.
#
# The only **required** element is a station:
minimum_request = hf.NWIS('01585200')
# Since we only specified the *where*, the NWIS will assume the following elements:
#
# - **Service**: if not specified, provide the daily average value ('dv')
# - **When**: if a `start_date` or `period` is not given, then provide the most recent reading.
# - **What**: if you don't ask for a specific parameter (`parameterCd`), you will get everything.
#
# Let's see what our request came back with:
minimum_request
# Here's what the data look like in table form:
minimum_request.df()
# ## Different ways to specify which site you want
# You can specify a site four different ways:
#
# - as a number or list of site numbers
# - using `stateCd` and a two letter postal code to retrieve every site in the state
# - using `countyCd` and a FIPS code to retrieve every site in a county or list of counties
# - using `bBox` to retrieve everything inside of a bounding box of latitudes and longitudes.
#
# You are required to set **one** of these parameters, but only one.
#
# All of these parameters are demonstrated in [Selecting Sites](Selecting_Sites.html)
# ## Different ways to specify time
# You can specify time in three different ways:
#
# - if you specify nothing, you'll get the most recent reading.
# - `period` will return up to 999 days of the most recent data: `period='P11D'`
# - `start_date` will return all of the data starting at this date: `start_date='2014-12-31'`
#
# If you specify a `start_date`, you can also specify an `end_date`, which is given in the same format.
# ## What happens when you make a bad request?
# The power of the NWIS also makes it easy to make mistakes.
# So, we've added a series of helpful error messages to let you know when something went wrong, and why it went wrong.
# For example, let's mistpye one of our parameters that worked so well above:
notSoGoodNWIS = hf.NWIS('01585200', 'xx', period='P200D')
# Okay, maybe I shouldn't have typed 'xx' for our service.
#
#
# Some errors get caught by hydrofunctions, but some don't. Sometimes we end up asking NWIS for something that doesn't make sense, or something that it doesn't have, or maybe NWIS isn't available. In this case, hydrofunctions will receive an error message from NWIS and help you figure out what went wrong.
# Let's ask for the impossible: the start date is AFTER the end date:
badRequest = hf.get_nwis('01585200', 'dv', '2017-12-31', '2017-01-01')
# ## Getting help
# I probably shouldn't have started with all of the things that go wrong! My point is that we've got ya.
#
# Where can you go to learn how to do things the RIGHT way?
#
# * [The User's Guide](https://hydrofunctions.readthedocs.io/en/latest/?badge=latest)
# * [The USGS guide to their waterservices](https://help.waterdata.usgs.gov/faq/automated-retrievals)
#
# **But we also have a few built-in helpers that you can use right here, right now:**
#
# * help() and ? will list the docstring for whatever object you are curious about
# * dir() and .\<TAB\> will tell you about available methods.
# Use the help() function to see all of the parameters for a function, their default values,
# and a short explanation of what it all means. Or you can type ?hf.NWIS to access the same information.
help(hf.NWIS)
# Use the dir() function to see what sort of methods you have available to you,
# or type hf.NWIS.<TAB> to see the same list.
dir(hf.NWIS)
|
docs/notebooks/Writing_Valid_Requests_for_NWIS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classifying data from samples
# ### Importing dependencies
# +
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, confusion_matrix
import pandas as pd
import numpy as np
# -
# ### Importing the dataset
# +
samples = pd.read_csv('./samples_data.csv', sep=",", index_col=0)
qttySamples = samples.shape[0]
qttyA = samples.loc[samples['maturity'] == 'A'].shape[0]
qttyB = samples.loc[samples['maturity'] == 'B'].shape[0]
qttyC = samples.loc[samples['maturity'] == 'C'].shape[0]
print('Number of samples: {0}'.format(qttySamples))
print('Number of A samples: {0} - {1}%'.format(qttyA, 100 * qttyA / qttySamples))
print('Number of B samples: {0} - {1}%'.format(qttyB, 100 * qttyB / qttySamples))
print('Number of C samples: {0} - {1}%'.format(qttyC, 100 * qttyC / qttySamples))
samples.head()
# -
# ### Preparing dataset
# +
x = samples.drop('maturity', axis=1) # Attributes
y = samples['maturity'] # Labels
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size = 0.20)
# -
# ### Preparing classifier
svclassifier = SVC(kernel='linear')
svclassifier.fit(X_train, y_train)
# ### Classifying and verifying results
# +
y_pred = svclassifier.predict(X_test)
print(confusion_matrix(y_test,y_pred))
print(classification_report(y_test,y_pred))
|
notebooks/ClassifyData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''anaconda3'': virtualenv)'
# name: python385jvsc74a57bd05b9c502b618e97131917a2f1409b4700bb639cdf99ce16cd88a0e27a90524386
# ---
# ### Import the libraries and the data
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import confusion_matrix, precision_score, accuracy_score, recall_score, f1_score, auc, roc_auc_score, plot_confusion_matrix, SCORERS, plot_roc_curve, classification_report, make_scorer
from sklearn.compose import ColumnTransformer
from sklearn.datasets import fetch_openml
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, OneHotEncoder, LabelEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split, GridSearchCV, cross_val_score
from sklearn.compose import make_column_selector as selector
np.random.seed(0)
# -
train_df = pd.read_csv("../data/01_raw/attrition_train.csv")
test_df = pd.read_csv("../data/01_raw/attrition_test.csv")
(train_df.Over18 != 'Y').sum()
(test_df.Over18 != 'Y').sum()
(train_df.EmployeeCount != 1).sum()
(test_df.EmployeeCount != 1).sum()
(train_df.StandardHours != 80).sum()
(test_df.StandardHours != 80).sum()
train_df.drop(columns=['Over18', 'EmployeeCount', 'EmployeeNumber', 'StandardHours'], inplace=True)
test_df.drop(columns=['Over18', 'EmployeeCount', 'EmployeeNumber', 'StandardHours', 'index1'], inplace=True)
# ### Get info about column names, missing values and Dtype
train_df.info()
test_df.info()
# ### Create X and y dataframes, split them into train and test, get the list of categorical and numeric features.
# +
y = train_df.Attrition
X = train_df.drop(['Attrition'], axis=1)
le = LabelEncoder()
y = le.fit_transform(y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
categorical_features = [cname for cname in X_train.columns if
X_train[cname].dtype == "object"]
numeric_features = [cname for cname in X_train.columns if
X_train[cname].dtype == "int64"]
# -
categorical_features
numeric_features
X.Age.describe()
# ### Preprocess the training data, create the classifier model and fit it with the data.
# +
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
categorical_transformer = OneHotEncoder(handle_unknown='ignore')
preprocessor = ColumnTransformer(transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)
])
clf = Pipeline(steps=[('preprocessor', preprocessor),
('classifier', LogisticRegression(random_state=0))])
clf.fit(X_train, y_train)
# -
# ### Get an extended list of numeric features, with the One Hot encoded categorical features.
onehot_columns = list(clf.named_steps['preprocessor'].named_transformers_['cat'].get_feature_names(input_features=categorical_features))
numeric_features_list = list(numeric_features)
numeric_features_list.extend(onehot_columns)
numeric_features_list
# ### Associate the features list to their coefficient in a dataframe, sorted in descending order.
coefs = best_clf.named_steps['classifier'].coef_
features_coef = pd.DataFrame(numeric_features_list)
features_coef['coef'] = pd.Series(coefs[0])
features_coef.rename(columns={0: "features"}, inplace=True)
features_coef.sort_values(by='coef', ascending=False)
SCORERS.keys()
# +
f1_scores = cross_val_score(clf, X, y, cv=5, scoring='f1')
roc_auc_scores = cross_val_score(clf, X_train, y_train, cv=5, scoring='roc_auc')
precision_scores = cross_val_score(clf, X_train, y_train, cv=5, scoring='precision')
recall_scores = cross_val_score(clf, X_train, y_train, cv=5, scoring='recall')
print("Average F1 scores: %.3f\n" % f1_scores.mean())
print("Average roc auc scores: %.3f\n" % roc_auc_scores.mean())
print("Average Precision scores: %.3f\n" % precision_scores.mean())
print("Average Recall scores: %.3f\n" % recall_scores.mean())
# -
# ### Recherche des paramètres du modèle de classification par une validation croisée
# + tags=["outputPrepend"]
param_grid = {
'classifier__C': [1, 3, 10, 30, 100],
'classifier__dual': [True, False],
'classifier__penalty' : ['l1', 'l2', 'elasticnet', 'none'],
'classifier__solver' : ['newton-cg', 'lbfgs', 'liblinear', 'sag'],
'classifier__l1_ratio' : [0.3, 0.5, 0.7],
'classifier__class_weight' : ['balanced', None]
}
scorers = {'f1': make_scorer(f1_score),
'roc_auc': make_scorer(roc_auc_score)}
grid_search = GridSearchCV(clf, param_grid, scoring=scorers, refit='roc_auc', cv=5)
grid_search.fit(X_train, y_train)
# -
grid_search.best_estimator_
grid_search.best_score_
grid_search.best_params_
clf.get_params().keys()
best_clf = grid_search.best_estimator_
best_clf
y_pred_proba = best_clf.predict_proba(X_test)
y_pred_proba[1][1]
attrition_score = []
for i in range(len(X_test)):
attrition_score.append(y_pred_proba[i][1])
y_pred = best_clf.predict(X_test)
y_pred_noparam = clf.predict(X_test)
y_pred
tn, fp, fn, tp = confusion_matrix(y_test, y_pred).ravel()
confusion_matrix(y_test, y_pred)
confusion_matrix(y_test, y_pred_noparam)
print("Sur les 220 salariés du jeu de données test :\n")
print("Le nombre de personnes qui ont quitté l'entreprise est : {}".format(fn + tp))
print("Le nombre de personnes dont on a prédit qu'il allaient quitter l'entreprise: {}".format(tp + fp))
print("Le nombre de personnes qui sont restées dans l'entreprise: {}".format(tn + fp))
print("Le nombre de cas total où on s'est trompé: {}".format(fn + fp))
print("Le nombre de cas où on s'est trompé sur des personnes qui avaient l'intention de partir: {}".format(fn))
print("Le nombre de cas où on s'est trompé sur des personnes qui voulaient rester: {}".format(fp))
# ### Calcul des métriques
# +
print("Métriques calculées par sklearn.metrics\n")
print("Précision : %.3f" % precision_score(y_test, y_pred))
print("Accuracy : %.3f" % accuracy_score(y_test, y_pred))
print("Rappel (ou sensibilité) : %.3f" % recall_score(y_test, y_pred))
print("F-measure : %.3f\n" % f1_score(y_test, y_pred))
print("Métriques recalculées\n")
print("Précision : {}".format(tp /(tp + fp)))
print("Accuracy : {}".format((tp + tn) /(tp + fp + tn + fn)))
print("Rappel (ou sensibilité) : {}".format(tp /(tp + fn)))
print("Spécificité : {}".format(tn /(fp + tn)))
print("F-measure : {}\n".format(2*tp /(2*tp + fp + fn)))
# -
plot_confusion_matrix(clf, X_test, y_test)
plt.show()
plot_confusion_matrix(best_clf, X_test, y_test)
plt.show()
# On veut améliorer le rappel (recall) ou sensibilité et la précision.
#
#
# Le rappel est intuitivement la capacité du classificateur à trouver tous les échantillons positifs.
# Un mauvais rappel signifie dans notre cas que nous avons sélectionné peu d'éléments pertinents et donc manqué de proposer le programme d'accompagnement à des personnes qui souhaitaient quitter l'entreprise.
# Or, c'est précisément ce que nous ne voulons pas pour éviter le turn-over.
#
# La précision est intuitivement la capacité du classificateur à ne pas étiqueter comme positif un échantillon qui est négatif.
# Dans notre cas, une mauvaise précision nous fera inscrire au programme d'accompagnement des personnes qui ne souhaitaient pas quitter l'entreprise, ce qui occasionera un coût non nécessaire, en plus de prendre des places qui auraient été plus utiles si elles avaient étaient attribuées aux personnes susceptibles de partir.
#
#
# Le score F1 peut être interprété comme une moyenne pondérée de la précision et du rappel, où un score F1 atteint sa meilleure valeur à 1 et son pire score à 0. La contribution relative de la précision et du rappel au score F1 est égale.
#
plot_roc_curve(clf, X_test, y_test)
plot_roc_curve(best_clf, X_test, y_test)
plt.show()
print(classification_report(y_test, y_pred))
# ### Prédiction du taux d'attrition pour les employés de l'entreprise (fichier attrition_test.csv)
#
best_clf.fit(X, y)
# +
f1_scores = cross_val_score(best_clf, X, y, cv=5, scoring='f1')
roc_auc_scores = cross_val_score(best_clf, X, y, cv=5, scoring='roc_auc')
precision_scores = cross_val_score(clf, X, y, cv=5, scoring='precision')
recall_scores = cross_val_score(clf, X, y, cv=5, scoring='recall')
print("Average F1 scores: %.3f\n" % f1_scores.mean())
print("Average roc auc scores: %.3f\n" % roc_auc_scores.mean())
print("Average Precision scores: %.3f\n" % precision_scores.mean())
print("Average Recall scores: %.3f\n" % recall_scores.mean())
# -
attrition_score_pred = best_clf.predict_proba(test_df)
attrition_score_pred
attrition_score = []
for i in range(len(test_df)):
attrition_score.append(int(100*attrition_score_pred[i][1]))
attrition_pred = best_clf.predict(test_df)
attrition_pred
test_df
test_df['PredictWillLeave'] = attrition_pred
test_df['AttritionScore%'] = attrition_score
(test_df.PredictWillLeave == 1).sum()
will_leave = test_df.loc[(test_df.PredictWillLeave == 1)]
will_leave
(will_leave['AttritionScore%'] >= 60).sum()
(will_leave['AttritionScore%'] >= 70).sum()
(will_leave['AttritionScore%'] >= 80).sum()
will_leave.loc[(will_leave['AttritionScore%'] >= 90)]
thresh70 = will_leave.loc[(will_leave['AttritionScore%'] >= 70)]
np.array(will_leave.Age)
test_df.to_csv('../data/01_raw/predictions.csv')
|
notebooks/2021-04-14-at-cs-logistic_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from singlezone_gen import main
zone_area=8.30
zone_ratio=0.904
zone_height=2.5
azimuth=90
absorptance=.5
wall_u=4.083
wall_ct=165.6
ground=1
roof=1
shading=[0.5,0,0,0]
living_room = False
exp=[1,0,0,1]
wwr=[0.198,0,0,0]
open_fac=[0.45,0,0,0]
glass_fs=.87
equipment=0
lights = 5
bldg_ratio=0.85
floor_height=0
door=False #por limitação do código só é possível utilizar False para door
has_hive=True
input_file='seed.json'
output= 'dorm2_hive_floor1_roof1.epJSON'
main(zone_area,zone_ratio,zone_height,azimuth,absorptance,wall_u,wall_ct,ground,roof,shading,living_room,exp,wwr,open_fac,glass_fs,equipment,lights,bldg_ratio,floor_height,door,has_hive,input_file,output)
|
singlezone_gen/run_dorm2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 6 - Other Popular Machine Learning Methods
# ## Segment 4 - Decision tree models with CART
#
# ### Decision Trees
import os
os.environ["path"]
os.environ["path"]=os.environ["path"]+";C:\\Program Files (x86)\\Graphviz2.38\\bin"
os.environ["path"]
import sklearn.datasets as datasets
import pandas as pd
from sklearn import metrics
# ### Preparing the data
# +
iris=datasets.load_iris()
df = pd.DataFrame(iris.data, columns= iris.feature_names)
y = pd.DataFrame(iris.target)
y.columns = ['labels']
df.head()
# -
y.labels.value_counts()
# ### The decision tree model
# +
from sklearn.tree import DecisionTreeClassifier
dtree = DecisionTreeClassifier()
dtree.fit(df, y)
# + tags=[]
# ! pip install pydotplus
# + tags=[]
# ! pip install graphviz
# +
from sklearn.externals.six import StringIO
from IPython.display import Image
from sklearn.tree import export_graphviz
import pydotplus
dot_data = StringIO()
export_graphviz(dtree, out_file=dot_data, filled=True, rounded=True, special_characters=True)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())
# -
|
Pt_2/06_04_Decision_tree_models_with_CART/6_04_part1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #Fire up GraphLab Create
#
# We always start with this line before using any part of GraphLab Create
import graphlab
# #Load a tabular data set
sf = graphlab.SFrame('people-example.csv')
# #SFrame basics
sf #we can view first few lines of table
sf.tail() # view end of the table
# #GraphLab Canvas
# .show() visualizes any data structure in GraphLab Create
sf.show()
# If you want Canvas visualization to show up on this notebook,
# rather than popping up a new window, add this line:
graphlab.canvas.set_target('ipynb')
sf['age'].show(view='Categorical')
# #Inspect columns of dataset
sf['Country']
sf['age']
# Some simple columnar operations
sf['age'].mean()
sf['age'].max()
# #Create new columns in our SFrame
sf
sf['Full Name'] = sf['First Name'] + ' ' + sf['Last Name']
sf
sf['age'] * sf['age']
# #Use the apply function to do a advance transformation of our data
sf['Country']
sf['Country'].show()
def transform_country(country):
if country == 'USA':
return 'United States'
else:
return country
transform_country('Brazil')
transform_country('Brasil')
transform_country('USA')
sf['Country'].apply(transform_country)
sf['Country'] = sf['Country'].apply(transform_country)
sf
|
Course1/Getting Started with SFrames.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Structures and Algorithms in Python
# ## Python Primer
# +
# R-1.1
def is_multiple(n, m):
return n % m == 0
is_multiple(12,3)
# +
# R-1.2
def is_even(k):
return k % 2 == 0
is_even(2)
# +
# R-1.3
def minmax(data):
min = data[0]
max = data[0]
for i in data:
if i < min:
min = i
if i > max:
max = i
return min, max
minmax([1,3,4,5,6,2,233,3,7])
# +
# R-1.4
def sum_of_squares(n):
sum = 0
for i in range(n,0, -1):
sum += i * i
return sum
sum_of_squares(4)
# -
# R-1.5
sum(i * i for i in range(4, 0, -1))
# +
# R-1.6
def sum_of_squares_odd(n):
sum = 0
for i in range(n-1,0, -1):
if i % 2 != 0:
sum += i * i
return sum
sum_of_squares_odd(4)
# -
# R-1.7
sum(i * i for i in range(4-1, 0, -1) if i % 2 != 0)
# +
# R-1.8
s = "jakob"
s_length = len(s)
print(s[-3])
print(s[s_length - 3])
# -
# R-1.9
for i in range(50, 90, 10):
print(i)
# R-1.10
for i in range(8, -10, -2):
print(i)
# R-1.11
print([2**i for i in range(0, 9)])
# +
# R-1.12
import random
def choice(data):
data_length = len(data)
return data[random.randrange(data_length)]
choice([0,2,3,4,5,6])
# -
# C-1.13
data = [1,2,3,4,5]
print(data[::-1])
print(list(reversed(data)))
# +
# C-1.14
def distinct_product(data):
nums = set(data)
for i in nums:
for j in nums:
if i * j % 2 != 0 and i != j:
return True
return False
distinct_product([1, 4, 5])
# +
# C-1.15
def is_distinct(data):
nums = set(data)
if len(nums) != len(data):
return False
return True
is_distinct([1,2])
# +
# C-1.16
def scale(data, factor):
for i in range(len(data)):
data[i] *= factor
data = [1,2]
scale(data, 2)
print(data)
# This works because we refer to elements in the list
# +
# C-1.17
def scale(data, factor):
for val in data:
val *= factor
data = [1,2]
scale(data, 2)
print(data)
# This does not work because we are just taking the values of the elements of the list,
# but aren't referencing them directly
# -
# C-1.18
# 0 2 6 12 20 30 ...
[k*(k-1) for k in range(1, 11)]
# C-1.19
[chr(k) for k in range(97, 123)]
# +
# C-1.20
def shuffle(data):
used = []
count = 0
number_of_occurences_data = 0
number_of_occurences_used = 0
while len(data) > len(used):
num = data[random.randint(0, len(data)-1)]
for i in data:
if i == num:
number_of_occurences_data += 1
for i in used:
if i == num:
number_of_occurences_used += 1
if num not in used or number_of_occurences_data > number_of_occurences_used:
used.append(num)
count += 1
number_of_occurences_data = 0
number_of_occurences_used = 0
return used
shuffle([1,2,5,23,3, 2])
# -
# C-1.21
inputArray = []
try:
while True:
inputArray.append(input("Please enter something: "))
except EOFError:
for i in reversed(inputArray):
print(i)
# C-1.22
def dot_product(a, b):
result = []
for i in range(len(a)):
result.append(a[i]*b[i])
return result
dot_product([2,3], [3,4])
# +
# C-1.23
def write_index(data, value, index):
if index > len(data):
print("Don't try to buffer overflow attacks in Python!")
return
data[index] = value
data = [3,2,1]
write_index(data, 5, 6)
print(data)
# +
# C-1.24
def count_vowels(word):
count = 0
for i in word:
if i in "aeiou":
count += 1
return count
count_vowels("jakobus")
# +
# C-1.25
def remove_punctuation(sentence):
result = ""
for i in sentence:
if i not in ".,;:'":
result += i
return result
remove_punctuation("Let's try, Mike.")
# +
# C-1.26
def can_be_used(a, b, c):
if a * b == c:
return True
if a - b == c:
return True
if a + b == c:
return True
return False
can_be_used(1,2,4)
# +
# C-1.27 # TODO
from math import sqrt
def factors(n):
k=1
while k * k <= n:
if n % k == 0:
yield k
if sqrt(n) % 1 == 0:
yield int(sqrt(n))
yield n // k
k += 1
if k * k == n:
break
for i in factors(6):
print(i)
# +
# C-1.28
def norm(v, p=2):
sum = 0
for num in v:
sum += num**2
return sum**(1/p)
norm([1,1], 4)
# +
# P-1.29
def permute(data, i, length):
if i==length:
print(''.join(data) )
else:
for j in range(i,length):
#swap
data[i], data[j] = data[j], data[i]
permute(data, i+1, length)
data[i], data[j] = data[j], data[i]
string = "catdog"
n = len(string)
data = list(string)
permute(data, 0, n)
# +
# P-1.30
def by2(n, count=0):
if n < 2:
# Why can't I return count here?
print(count)
else:
by2(n//2, count+1)
by2(16)
# +
# P-1.31
def change(betrag, gezahlt):
euro_1 = 0
euro_2 = 0
cent_50 = 0
cent_20 = 0
cent_10 = 0
cent_5 = 0
cent_2 = 0
cent_1 = 0
change = gezahlt - betrag
euro_2 = change // 2
remaining = change % 2
euro_1 = remaining // 1
remaining = change % 1
cent_50 = remaining // 0.5
remaining = remaining % 0.5
cent_20 = remaining // 0.2
remaining = remaining % 0.2
cent_10 = remaining // 0.1
remaining = remaining % 0.1
cent_5 = remaining // 0.05
remaining = remaining % 0.05
cent_2 = remaining // 0.02
remaining = remaining % 0.02
cent_1 = round(remaining, 2) // 0.01
return euro_2, euro_1, cent_50, cent_20, cent_10, cent_5, cent_2, cent_1
change(2.20, 4.29)
# +
# P-1.35
import random
def same_birthday(n):
birthdays = []
for i in range(n):
birthdays.append((random.randrange(1,31), random.randrange(1,12)))
print(birthdays)
for i in range(len(birthdays)):
for j in range(len(birthdays)):
if i != j:
if birthdays[i][0] == birthdays[j][0] and birthdays[i][1] == birthdays[j][1]:
return True
return False
same_birthday(24)
# +
# P-1.36
def count_words(words):
data = words.split(" ")
print(data)
result = {}
for word in data:
if word in result:
result[word] += 1
if word not in result:
result[word] = 1
print(result)
words = "Ich bin Jakob und ich bin"
count_words(words)
# -
# ## OOP
# +
# R-2.1
# Rocket software
# Car software
# Medical software
# +
# R-2.2
# Overflow on a bank account
# +
# R-2.3
# Pen
# change_color()
# change_thickness()
# +
# R-2.4
class Flower:
def __init__(self, name, number_of_petals, price):
self.name = name
self.number_of_petals = number_of_petals
self.price = price
def set_name(self, name):
self.name = name
def get_name(self):
return self.name
def set_number_of_petals(self, number_of_petals):
self.number_of_petals = number_of_petals
def get_number_of_petals(self):
return self.number_of_petals
def set_price(self):
self.price = price
def get_price(self):
return self.price
flower = Flower("rose", 12, 2)
print(flower.get_price())
# +
# R-2.9
class Vector:
def __init__(self, d):
# Create d-dimensional vectors of zeros
self._coords = [0] * d
def __len__(self):
# Return the dimension of the vector
return len(self._coords)
def __getitem__(self, j):
# Return the jth coordinate of the vector
return self._coords[j]
def __setitem__(self, j, val):
# Set jth coordinate of vector to given value
self._coords[j] = val
def __add__(self, other):
# Retun sum of two vectors
if len(self) != len(other):
raise ValueError('dimensions must match')
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] + other[j]
return result
def __sub__(self, other):
# Return difference of two vectors
if len(self) != len(other):
raise ValueError('dimensions must watch')
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] - other[j]
return result
def __eq__(self, other):
# Return true if vector has same coordinates as other
return self._coords == other._coords
def __ne__(self, other):
# Return true if vector differs from other
return not self == other
def __str__(self):
# Produce string representation of vector
return '<' + str(self._coords)[1:-1] + '>'
v = Vector(3)
v[1] = 3
str(v)
v =
# -
|
data_structures_and_algorithms_in_python_exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
def testing(model, dataset):
result = []
if torch.cuda.is_available():
model.cuda()
for data in dataset:
inputs, labels = data
if torch.cuda.is_available():
inputs, labels = inputs.cuda(), labels.cuda()
output = model(inputs)
_, preds = torch.max(output, 1)
# testing data is sequential, so label is unnecessary thing
result += [int(element) for element in preds.tolist()]
return result
|
testing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %cd ..
from dev.retina_copie import Retina, RetinaWhiten
#args = init(filename='../data/2019-07-23')
from easydict import EasyDict
args = EasyDict({'w': 28,
'minibatch_size': 10,
'train_batch_size': 1000,
'test_batch_size': 126,
'noise_batch_size': 1000,
'mean': 0.1307,
'std': 0.3081,
'N_pic': 1718,
'N_X': 1718,
'N_Y': 2444,
'N_X2': 98,
'N_Y2': 140,
'offset_std': 30,
'offset_max': 34,
'noise': 0.75,
'contrast': 0.7,
'sf_0': 0.1,
'B_sf': 0.1,
'N_theta': 6,
'N_azimuth': 48,
'N_eccentricity': 24,
'N_phase': 2,
'rho': 1.41,
'bias_deconv': True,
'p_dropout': 0.5,
'dim1': 1000,
'dim2': 1000,
'lr': 0.005,
'do_adam': 'adam',
'bn1_bn_momentum': 0,
'bn2_bn_momentum': 0,
'momentum': 0,
'epochs': 60,
'num_processes': 1,
'no_cuda': True,
'log_interval': 20,
'verbose': 1,
'filename': '../data/2019-07-23',
'seed': 2019,
'N_cv': 10,
'do_compute': True,
'save_model': True,
'zoomW': 300})
print('N_X :', args.N_X)
print('N_Y :', args.N_Y)
print('N_theta :', args.N_theta)
print('N_azimuth :', args.N_azimuth)
print('N_eccentricity :', args.N_eccentricity)
print('N_phase :', args.N_phase)
args.N_X, args.N_Y = 768, 1024
args_N_pic = args.N_Y
retina = Retina(args)
retina.retina_dico.keys() # theta
retina.retina_dico[0].keys() # phi
retina.retina_dico[0][0].keys() # eccentricity
retina.retina_dico[0][0][16].shape # central filter
retina.retina_dico[0][0][23].shape # periphery filter
plt.imshow(retina.retina_dico[2][0][16].reshape((6, 6)))
plt.imshow(retina.retina_dico[0][0][23].reshape((76, 76)))
plt.imshow(retina.retina_dico[2][0][23].reshape((76, 76)))
plt.imshow(retina.retina_dico[2][1][23].reshape((76, 76)))
from PIL import Image
impath = 'data/i05june05_static_street_boston_p1010764.jpeg'
im = Image.open(impath)
im
im_color_npy = np.asarray(im)
im_color_npy
im_bw = im.convert("L")
im_bw
im_npy = np.asarray(im_bw)
im_npy
args.N_X, args.N_Y = im_npy.shape
args_N_pic = args.N_Y
# +
##
# -
whiten_transform = RetinaWhiten(args)
im_whiten = whiten_transform(im_npy)
plt.figure(figsize=(20,20))
plt.imshow(im_whiten, cmap='gray')
pixel_fullfield, retina_features = retina.transform(im_whiten)
retina_features.shape
plt.plot(retina_features)
img_trans = retina.inverse_transform(retina_features)
plt.figure(figsize=(20,20))
plt.imshow(img_trans, cmap='gray')
# +
from LogGabor import LogGabor
N_theta=6
rho = 1.41
ecc_max = .8 # self.args.ecc_max
sf_0_r = 0.03 # self.args.sf_0_r
sf_0_max = 0.45
B_theta = np.pi / N_theta / 2 # self.args.B_theta
B_sf = .4
pe = {'N_image': 100, 'seed': None, 'N_X': 512, 'N_Y': 512, 'noise':
0.1, 'do_mask': True, 'mask_exponent': 3.0, 'do_whitening': True,
'white_name_database': 'kodakdb', 'white_n_learning': 0, 'white_N':
0.07, 'white_N_0': 0.0, 'white_f_0': 0.4, 'white_alpha': 1.4,
'white_steepness': 4.0, 'white_recompute': False, 'base_levels':
1.618, 'n_theta': 24, 'B_sf': 0.4, 'B_theta': 0.17453277777777776,
'use_cache': True, 'figpath': 'results', 'edgefigpath':
'results/edges', 'matpath': 'cache_dir', 'edgematpath':
'cache_dir/edges', 'datapath': 'database/', 'ext': '.pdf', 'figsize':
14.0, 'formats': ['pdf', 'png', 'jpg'], 'dpi': 450, 'verbose': 0}
lg = LogGabor(pe=pe)
i_eccentricity = 15
N_eccentricity = 20
ecc = ecc_max * (1 / rho) ** (N_eccentricity - i_eccentricity)
i_theta = 5
N_theta = 6
theta_ref = i_theta * np.pi / N_theta
sf_0 = 0.5 * sf_0_r / ecc
sf_0 = np.min((sf_0, sf_0_max))
N = 200
dimension_filtre = int( N /2 * ecc)
#if dimension_filtre % 2 == 1:
# dimension_filtre += 1
#dimension_filtre = 100
# print("dimension_filtre", dimension_filtre)
lg.set_size((dimension_filtre, dimension_filtre))
i_phase = 1
phase = i_phase * np.pi / 2
x = dimension_filtre // 2
y = dimension_filtre // 2
params = {'sf_0': sf_0,
'B_sf': B_sf,
'theta': theta_ref,
'B_theta': B_theta}
filter = lg.normalize(lg.invert(lg.loggabor(x, y, **params) *
np.exp(-1j * phase)))
# -
plt.imshow(filter)
plt.plot(ecc_max * (1 / rho) ** (N_eccentricity - np.arange(N_eccentricity)))
plt.plot(ecc_max * (1 / 1.21) ** (N_eccentricity - np.arange(N_eccentricity)))
plt.plot(ecc_max * (1 / 1.11) ** (N_eccentricity - np.arange(N_eccentricity)))
plt.plot(ecc_max * (1 / rho) ** (N_eccentricity - np.arange(10)))
|
dev/2019-12-03_retina-tests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
df=pd.read_csv("C:\\Users\\HP\\Desktop\\LGMVIP Task2\\data.csv")
df.head()
df.tail()
df1=df.reset_index()["Close"]
df1
import matplotlib.pyplot as plt
plt.plot(df1)
import numpy as np
df1
from sklearn.preprocessing import MinMaxScaler
Scaler = MinMaxScaler (feature_range=(0,1))
df1=Scaler.fit_transform(np.array(df1).reshape(-1,1))
print(df1)
training_size= int(len(df1)*0.65)
test_size= len(df1)-training_size
train_data, test_data= df1[0:training_size,:],df1[training_size:len(df1),:1]
training_size, test_size
train_data
import numpy
def create_dataset(dataset, time_step=1):
dataX, dataY = [],[]
for i in range(len(dataset)-time_step-1):
a = dataset[i:(i+time_step),0]
dataX.append(a)
dataY.append(dataset[i+ time_step, 0])
return numpy.array(dataX), numpy.array(dataY)
time_step = 100
X_train, y_train = create_dataset(train_data, time_step)
X_test, ytest = create_dataset(test_data, time_step)
print(X_train.shape), print(y_train.shape)
X_train= X_train.reshape(X_train.shape[0], X_train.shape[1], 1)
X_train= X_train.reshape(X_train.shape[0], X_train.shape[1], 1)
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import LSTM
model=Sequential()
model.add(LSTM(50,return_sequences = True, input_shape=(100,1)))
model.add(LSTM(50,return_sequences = True))
model.add(LSTM(50))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.summary()
model.fit(X_train, y_train, validation_data=(X_test,ytest),epochs=100,batch_size=64,verbose=1)
# +
import tensorflow as tf
train_predict=model.predict(X_train)
test_predict=model.predict(X_test)
import sklearn
X = sklearn.preprocessing.standardScaler().fit_transform(X)
train_predict=x_scaler.inverse_transform(train_predict)
test_predict=x_scaler.inverse_transform(test_predict)
# -
import tensorflow as tf
train_predict=model.predict(X_train)
test_predict=model.predict(X_test)
import math
from sklearn.metrics import mean_squared_error
math.sqrt(mean_squared_error(y_train,train_predict))
### Test Data RMSE
math.sqrt(mean_squared_error(ytest,test_predict))
### Plotting
# shift train predictions for plotting
look_back=100
trainPredictPlot = numpy.empty_like(df1)
trainPredictPlot[:, :] = np.nan
trainPredictPlot[look_back:len(train_predict)+look_back, :] = train_predict
# shift test predictions for plotting
testPredictPlot = numpy.empty_like(df1)
testPredictPlot[:, :] = numpy.nan
testPredictPlot[len(train_predict)+(look_back*2)+1:len(df1)-1, :] = test_predict
# plot baseline and predictions
plt.plot(scaler.inverse_transform(df1))
plt.plot(trainPredictPlot)
plt.plot(testPredictPlot)
plt.show()
len(test_data)
x_input=test_data[613:].reshape(1,-1)
x_input.shape
temp_input=list(x_input)
temp_input=temp_input[0].tolist()
# +
# demonstrate prediction for next 10 days
from numpy import array
lst_output=[]
n_steps=100
i=0
while(i<30):
if(len(temp_input)>100):
#print(temp_input)
x_input=np.array(temp_input[1:])
print("{} day input {}".format(i,x_input))
x_input=x_input.reshape(1,-1)
x_input = x_input.reshape((1, n_steps, 1))
#print(x_input)
yhat = model.predict(x_input, verbose=0)
print("{} day output {}".format(i,yhat))
temp_input.extend(yhat[0].tolist())
temp_input=temp_input[1:]
#print(temp_input)
lst_output.extend(yhat.tolist())
i=i+1
else:
x_input = x_input.reshape((1, n_steps,1))
yhat = model.predict(x_input, verbose=0)
print(yhat[0])
temp_input.extend(yhat[0].tolist())
print(len(temp_input))
lst_output.extend(yhat.tolist())
i=i+1
print(lst_output)
# -
day_new=np.arange(1,101)
day_pred=np.arange(101,131)
len(df1)
|
Stock market prediction and Forecasting using Stacked LSTM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example of constructing an environment using Python
import gym
#creating cartpole-v0 environment
env = gym.make('CartPole-v0')
#intializing the environment
env.reset()
for _ in range(1000):
env.render()#render the environment for visual representation
env.step(env.action_space.sample())
env.close()#closing the environment for necessary cleanup
# # EXample of construction an agent using Python
import gym
#Creating cartpole-v0 environment
env = gym.make('CartPole-v0')
for _ in range(20):
obs = env.reset()
for i in range(100):
env.render()
print(obs)
action = env.action_space.sample()
obs, reward, done, info = env.step(action)
if done:
print("Episode finished after {} timesteps".format(i+1))
break
|
Chapter 09/Chapter09.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Face Mask Detection using PaddlePaddle
//
// In this tutorial, we will be using pretrained PaddlePaddle model from [PaddleHub](https://github.com/PaddlePaddle/PaddleHub/tree/release/v1.5/demo/mask_detection/cpp) to do mask detection on the sample image. To complete this procedure, there are two steps needs to be done:
//
// - Recognize face on the image (no matter wearing mask or not) using Face object detection model
// - classify the face is wearing mask or not
//
// These two steps will involve two paddle models. We will implement the corresponding preprocess and postprocess logic to it.
//
// ## Import dependencies and classes
//
// PaddlePaddle is one of the Deep Engines that requires DJL hybrid mode to run inference. Itself does not contains NDArray operations and needs a supplemental DL framework to help with that. So we import Pytorch DL engine as well in here to do the processing works.
// +
// // %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
// %maven ai.djl:api:0.14.0
// %maven ai.djl.paddlepaddle:paddlepaddle-model-zoo:0.14.0
// %maven org.slf4j:slf4j-api:1.7.32
// %maven org.slf4j:slf4j-simple:1.7.32
// second engine to do preprocessing and postprocessing
// %maven ai.djl.pytorch:pytorch-engine:0.14.0
// +
import ai.djl.*;
import ai.djl.inference.*;
import ai.djl.modality.*;
import ai.djl.modality.cv.*;
import ai.djl.modality.cv.output.*;
import ai.djl.modality.cv.transform.*;
import ai.djl.modality.cv.translator.*;
import ai.djl.modality.cv.util.*;
import ai.djl.ndarray.*;
import ai.djl.ndarray.types.Shape;
import ai.djl.repository.zoo.*;
import ai.djl.translate.*;
import java.io.*;
import java.nio.file.*;
import java.util.*;
// -
// ## Face Detection model
//
// Now we can start working on the first model. The model can do face detection and require some additional processing before we feed into it:
//
// - Resize: Shrink the image with a certain ratio to feed in
// - Normalize the image with a scale
//
// Fortunatly, DJL offers a `Translator` interface that can help you with these processing. The rough Translator architecture looks like below:
//
// 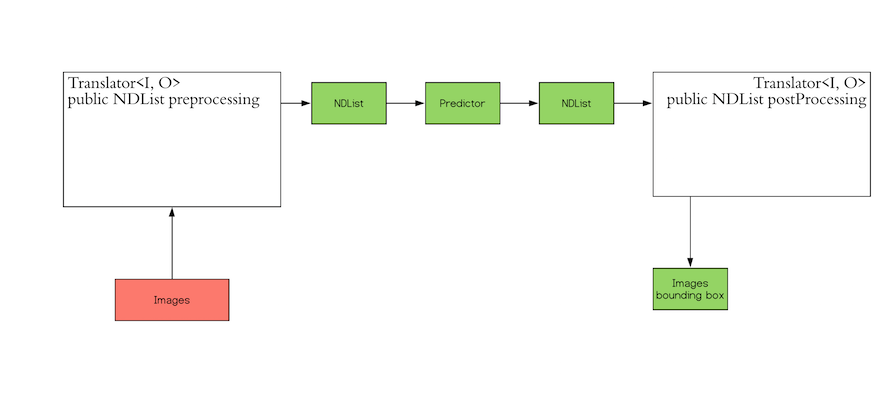
//
// In the following sections, we will implement a `FaceTranslator` class to do the work.
//
// ### Preprocessing
//
// In this stage, we will load an image and do some preprocessing work to it. Let's load the image first and take a look at it:
String url = "https://raw.githubusercontent.com/PaddlePaddle/PaddleHub/release/v1.5/demo/mask_detection/python/images/mask.jpg";
Image img = ImageFactory.getInstance().fromUrl(url);
img.getWrappedImage();
// Then, let's try to apply some transformation to it:
// +
NDList processImageInput(NDManager manager, Image input, float shrink) {
NDArray array = input.toNDArray(manager);
Shape shape = array.getShape();
array = NDImageUtils.resize(
array, (int) (shape.get(1) * shrink), (int) (shape.get(0) * shrink));
array = array.transpose(2, 0, 1).flip(0); // HWC -> CHW BGR -> RGB
NDArray mean = manager.create(new float[] {104f, 117f, 123f}, new Shape(3, 1, 1));
array = array.sub(mean).mul(0.007843f); // normalization
array = array.expandDims(0); // make batch dimension
return new NDList(array);
}
processImageInput(NDManager.newBaseManager(), img, 0.5f);
// -
// As you can see above, we convert the image to a NDArray with shape following (number_of_batches, channel (RGB), height, width). This is the required input for the model to run object detection.
//
// ### Postprocessing
//
// For postprocessing, The output is in shape of (number_of_boxes, (class_id, probability, xmin, ymin, xmax, ymax)). We can store them into the prebuilt DJL `DetectedObjects` classes for further processing. Let's assume we have an inference output of ((1, 0.99, 0.2, 0.4, 0.5, 0.8)) and try to draw this box out.
// +
DetectedObjects processImageOutput(NDList list, List<String> className, float threshold) {
NDArray result = list.singletonOrThrow();
float[] probabilities = result.get(":,1").toFloatArray();
List<String> names = new ArrayList<>();
List<Double> prob = new ArrayList<>();
List<BoundingBox> boxes = new ArrayList<>();
for (int i = 0; i < probabilities.length; i++) {
if (probabilities[i] >= threshold) {
float[] array = result.get(i).toFloatArray();
names.add(className.get((int) array[0]));
prob.add((double) probabilities[i]);
boxes.add(
new Rectangle(
array[2], array[3], array[4] - array[2], array[5] - array[3]));
}
}
return new DetectedObjects(names, prob, boxes);
}
NDArray tempOutput = NDManager.newBaseManager().create(new float[]{1f, 0.99f, 0.1f, 0.1f, 0.2f, 0.2f}, new Shape(1, 6));
DetectedObjects testBox = processImageOutput(new NDList(tempOutput), Arrays.asList("Not Face", "Face"), 0.7f);
Image newImage = img.duplicate();
newImage.drawBoundingBoxes(testBox);
newImage.getWrappedImage();
// -
// ### Create Translator and run inference
//
// After this step, you might understand how process and postprocess works in DJL. Now, let's do something real and put them together in a single piece:
class FaceTranslator implements NoBatchifyTranslator<Image, DetectedObjects> {
private float shrink;
private float threshold;
private List<String> className;
FaceTranslator(float shrink, float threshold) {
this.shrink = shrink;
this.threshold = threshold;
className = Arrays.asList("Not Face", "Face");
}
@Override
public DetectedObjects processOutput(TranslatorContext ctx, NDList list) {
return processImageOutput(list, className, threshold);
}
@Override
public NDList processInput(TranslatorContext ctx, Image input) {
return processImageInput(ctx.getNDManager(), input, shrink);
}
}
// To run inference with this model, we need to load the model from Paddle model zoo. To load a model in DJL, you need to specify a `Crieteria`. `Crieteria` is used identify where to load the model and which `Translator` should apply to it. Then, all we need to do is to get a `Predictor` from the model and use it to do inference:
// +
Criteria<Image, DetectedObjects> criteria = Criteria.builder()
.setTypes(Image.class, DetectedObjects.class)
.optModelUrls("djl://ai.djl.paddlepaddle/face_detection/0.0.1/mask_detection")
.optFilter("flavor", "server")
.optTranslator(new FaceTranslator(0.5f, 0.7f))
.build();
var model = criteria.loadModel();
var predictor = model.newPredictor();
DetectedObjects inferenceResult = predictor.predict(img);
newImage = img.duplicate();
newImage.drawBoundingBoxes(inferenceResult);
newImage.getWrappedImage();
// -
// As you can see above, it brings you three faces detections.
//
// ## Mask Classification model
//
//
// So, once we have the image location ready, we can crop the image and feed it to the Mask Classification model for further processing.
//
// ### Crop the image
//
// The output of the box location is a value from 0 - 1 that can be mapped to the actual box pixel location if we simply multiply by width/height. For better accuracy on the cropped image, we extend the detection box to square. Let's try to get a cropped image:
// +
int[] extendSquare(
double xmin, double ymin, double width, double height, double percentage) {
double centerx = xmin + width / 2;
double centery = ymin + height / 2;
double maxDist = Math.max(width / 2, height / 2) * (1 + percentage);
return new int[] {
(int) (centerx - maxDist), (int) (centery - maxDist), (int) (2 * maxDist)
};
}
Image getSubImage(Image img, BoundingBox box) {
Rectangle rect = box.getBounds();
int width = img.getWidth();
int height = img.getHeight();
int[] squareBox =
extendSquare(
rect.getX() * width,
rect.getY() * height,
rect.getWidth() * width,
rect.getHeight() * height,
0.18);
return img.getSubImage(squareBox[0], squareBox[1], squareBox[2], squareBox[2]);
}
List<DetectedObjects.DetectedObject> faces = inferenceResult.items();
getSubImage(img, faces.get(2).getBoundingBox()).getWrappedImage();
// -
// ### Prepare Translator and load the model
//
// For the face classification model, we can use DJL prebuilt `ImageClassificationTranslator` with a few transformation. This Translator brings a basic image translation process and can be extended with additional standard processing steps. So in our case, we don't have to create another `Translator` and just leverage on this prebuilt one.
// +
var criteria = Criteria.builder()
.setTypes(Image.class, Classifications.class)
.optModelUrls("djl://ai.djl.paddlepaddle/mask_classification/0.0.1/mask_classification")
.optFilter("flavor", "server")
.optTranslator(
ImageClassificationTranslator.builder()
.addTransform(new Resize(128, 128))
.addTransform(new ToTensor()) // HWC -> CHW div(255)
.addTransform(
new Normalize(
new float[] {0.5f, 0.5f, 0.5f},
new float[] {1.0f, 1.0f, 1.0f}))
.addTransform(nd -> nd.flip(0)) // RGB -> GBR
.build())
.build();
var classifyModel = criteria.loadModel();
var classifier = classifyModel.newPredictor();
// -
// ### Run inference
//
// So all we need to do is to apply the previous implemented functions and apply them all together. We firstly crop the image and then use it for inference. After these steps, we create a new DetectedObjects with new Classification classes:
// +
List<String> names = new ArrayList<>();
List<Double> prob = new ArrayList<>();
List<BoundingBox> rect = new ArrayList<>();
for (DetectedObjects.DetectedObject face : faces) {
Image subImg = getSubImage(img, face.getBoundingBox());
Classifications classifications = classifier.predict(subImg);
names.add(classifications.best().getClassName());
prob.add(face.getProbability());
rect.add(face.getBoundingBox());
}
newImage = img.duplicate();
newImage.drawBoundingBoxes(new DetectedObjects(names, prob, rect));
newImage.getWrappedImage();
|
jupyter/paddlepaddle/face_mask_detection_paddlepaddle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # REINFORCE on CartPole-v0
#
# > In this post, We will take a hands-on-lab of Monte Carlo Policy Gradient (also known as REINFORCE) on openAI gym CartPole-v0 environment. This is the coding exercise from udacity Deep Reinforcement Learning Nanodegree.
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Python, Reinforcement_Learning, PyTorch, Udacity]
# - image: images/CartPole-v0.gif
# ## REINFORCE
# ---
# In this notebook, you will implement REINFORCE agent on OpenAI Gym's CartPole-v0 environment. For summary, The **REINFORCE** algorithm ([Williams, 1992](https://link.springer.com/content/pdf/10.1007/BF00992696.pdf)) is a monte carlo variation of policy gradient algorithm in RL. The agent collects the trajectory of an episode from current policy. Usually, this policy depends on the policy parameter which denoted as $\theta$. Actually, REINFORCE is acronym for "**RE**ward **I**ncrement = **N**onnegative **F**actor * **O**ffset **R**einforcement * **C**haracteristic **E**ligibility"
#
# ### Import the Necessary Packages
# +
import gym
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (16, 10)
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.distributions import Categorical
torch.manual_seed(0)
import base64, io
# For visualization
from gym.wrappers.monitoring import video_recorder
from IPython.display import HTML
from IPython import display
import glob
# -
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
device
# ### Instantiate the Environment and Agent
#
# CartPole environment is very simple. It has discrete action space (2) and 4 dimensional state space.
# +
env = gym.make('CartPole-v0')
env.seed(0)
print('observation space:', env.observation_space)
print('action space:', env.action_space)
# -
# ### Define Policy
# Unlike value-based method, the output of policy-based method is the probability of each action. It can be represented as policy. So activation function of output layer will be softmax, not ReLU.
class Policy(nn.Module):
def __init__(self, state_size=4, action_size=2, hidden_size=32):
super(Policy, self).__init__()
self.fc1 = nn.Linear(state_size, hidden_size)
self.fc2 = nn.Linear(hidden_size, action_size)
def forward(self, state):
x = F.relu(self.fc1(state))
x = self.fc2(x)
# we just consider 1 dimensional probability of action
return F.softmax(x, dim=1)
def act(self, state):
state = torch.from_numpy(state).float().unsqueeze(0).to(device)
probs = self.forward(state).cpu()
model = Categorical(probs)
action = model.sample()
return action.item(), model.log_prob(action)
# ### REINFORCE
def reinforce(policy, optimizer, n_episodes=1000, max_t=1000, gamma=1.0, print_every=100):
scores_deque = deque(maxlen=100)
scores = []
for e in range(1, n_episodes):
saved_log_probs = []
rewards = []
state = env.reset()
# Collect trajectory
for t in range(max_t):
# Sample the action from current policy
action, log_prob = policy.act(state)
saved_log_probs.append(log_prob)
state, reward, done, _ = env.step(action)
rewards.append(reward)
if done:
break
# Calculate total expected reward
scores_deque.append(sum(rewards))
scores.append(sum(rewards))
# Recalculate the total reward applying discounted factor
discounts = [gamma ** i for i in range(len(rewards) + 1)]
R = sum([a * b for a,b in zip(discounts, rewards)])
# Calculate the loss
policy_loss = []
for log_prob in saved_log_probs:
# Note that we are using Gradient Ascent, not Descent. So we need to calculate it with negative rewards.
policy_loss.append(-log_prob * R)
# After that, we concatenate whole policy loss in 0th dimension
policy_loss = torch.cat(policy_loss).sum()
# Backpropagation
optimizer.zero_grad()
policy_loss.backward()
optimizer.step()
if e % print_every == 0:
print('Episode {}\tAverage Score: {:.2f}'.format(e, np.mean(scores_deque)))
if np.mean(scores_deque) >= 195.0:
print('Environment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(e - 100, np.mean(scores_deque)))
break
return scores
# ### Run
policy = Policy().to(device)
optimizer = optim.Adam(policy.parameters(), lr=1e-2)
scores = reinforce(policy, optimizer, n_episodes=2000)
# ### Plot the learning progress
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# ### Animate it with Video
# +
def show_video(env_name):
mp4list = glob.glob('video/*.mp4')
if len(mp4list) > 0:
mp4 = 'video/{}.mp4'.format(env_name)
video = io.open(mp4, 'r+b').read()
encoded = base64.b64encode(video)
display.display(HTML(data='''<video alt="test" autoplay
loop controls style="height: 400px;">
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii'))))
else:
print("Could not find video")
def show_video_of_model(policy, env_name):
env = gym.make(env_name)
vid = video_recorder.VideoRecorder(env, path="video/{}.mp4".format(env_name))
state = env.reset()
done = False
for t in range(1000):
vid.capture_frame()
action, _ = policy.act(state)
next_state, reward, done, _ = env.step(action)
state = next_state
if done:
break
vid.close()
env.close()
# -
show_video_of_model(policy, 'CartPole-v0')
show_video('CartPole-v0')
|
_notebooks/2021-05-12-REINFORCE-CartPole.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="bTry4ZMD2859"
# # What-If Tool Challenge Lab
#
# In this notebook, you will use mortgage data from NY in 2017 to create two binary classifiers to determine if a mortgage applicant will be granted a loan.
#
# You will train classifiers on two datasets. One will be trained on the complete dataset, and the other will be trained on a subset of the dataset, where 90% of the female applicants that were granted a loan were removed from the training data (so the dataset has 90% less females that were granted loans).
#
# You will then compare and examine the two models using the What-If Tool.
#
# In this notebook, you will be exepcted to:
# * Understand how the data is processed
# * Write TensorFlow code to build and train two models
# * Write code to deploy the the models to AI Platform
# * Examine the models in the What-If Tool
# + [markdown] colab_type="text" id="zU9bzX-VWQCb"
# # Download and import the data
#
# Here, you'll import some modules and download some data from the Consumer Finance public [datasets](https://www.consumerfinance.gov/data-research/hmda/historic-data/?geo=ny&records=all-records&field_descriptions=labels).
# + colab={} colab_type="code" id="nhmYvLmUxSqU"
import pandas as pd
import numpy as np
import collections
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.utils import shuffle
from witwidget.notebook.visualization import WitWidget, WitConfigBuilder
# + colab={} colab_type="code" id="oVhFQBvggsio"
# !wget https://files.consumerfinance.gov/hmda-historic-loan-data/hmda_2017_ny_all-records_labels.zip
# !unzip hmda_2017_ny_all-records_labels.zip
# + [markdown] colab_type="text" id="uFyKHeHZD1e6"
# # Process the Data
#
# In this section, you **don't need to write any code**. We suggest you read through the cells to understand how the dataset is processed.
#
# Here, we start by importing the dataset into a Pandas dataframe. Then we process the data to exclude incomplete information and make a simple binary classification of loan approvals. We then create two datasets, one complete and one where 90% of female applicants are removed.
# + colab={} colab_type="code" id="LSsrdPdyCVYn"
# Set column dtypes for Pandas
column_names = collections.OrderedDict({
'as_of_year': np.int16,
'agency_abbr': 'category',
'loan_type': 'category',
'property_type': 'category',
'loan_purpose': 'category',
'owner_occupancy': np.int8,
'loan_amt_000s': np.float64,
'preapproval': 'category',
'county_code': np.float64,
'applicant_income_00s': np.float64,
'purchaser_type': 'category',
'hoepa_status': 'category',
'lien_status': 'category',
'population': np.float64,
'ffiec_median_fam_income': np.float64,
'tract_to_msamd_income': np.float64,
'num_of_owner_occupied_units': np.float64,
'number_of_1_to_4_family_units': np.float64,
'approved': np.int8,
'applicant_race_name_3': 'category',
'applicant_race_name_4': 'category',
'applicant_race_name_5': 'category',
'co_applicant_race_name_3': 'category',
'co_applicant_race_name_4': 'category',
'co_applicant_race_name_5': 'category'
})
# Import the CSV into a dataframe
data = pd.read_csv('hmda_2017_ny_all-records_labels.csv', dtype=column_names)
data = shuffle(data, random_state=2)
# + [markdown] colab_type="text" id="5fMc5a2eY3Kh"
# ## Extract columns and create dummy dataframes
#
# We first specify which columns to keep then drop the columns that don't have `loan originated` or `loan denied`, to make this a simple binary classification.
#
# We then create two dataframes `binary_df` and `bad_binary_df`. The first will include all the data, and the second will have 90% of female applicants removed, respectively. We then convert them into "dummy" dataframes to turn categorical string features into simple 0/1 features and normalize all the columns.
# + colab={} colab_type="code" id="qWNJwq2-Htxz"
# Only use a subset of the columns for these models
text_columns_to_keep = [
'agency_name',
'loan_type_name',
'property_type_name',
'loan_purpose_name',
'owner_occupancy_name',
'applicant_ethnicity_name',
'applicant_race_name_1',
'applicant_sex_name',
]
numeric_columns_to_keep = [
'loan_amount_000s',
'applicant_income_000s',
'population',
'minority_population',
'hud_median_family_income'
]
columns_to_keep = text_columns_to_keep + numeric_columns_to_keep + ['action_taken_name']
# Drop columns with incomplete information and drop columns that don't have loan orignated or denied, to make this a simple binary classification
df = data[columns_to_keep].dropna()
binary_df = df[df.action_taken_name.isin(['Loan originated', 'Application denied by financial institution'])].copy()
binary_df.loc[:,'loan_granted'] = np.where(binary_df['action_taken_name'] == 'Loan originated', 1, 0)
binary_df = binary_df.drop(columns=['action_taken_name'])
# Drop 90% of loaned female applicants for a "bad training data" version
loaned_females = (binary_df['applicant_sex_name'] == 'Female') & (binary_df['loan_granted'] == 1)
bad_binary_df = binary_df.drop(binary_df[loaned_females].sample(frac=.9).index)
# + colab={} colab_type="code" id="ic6mWTvENrLd"
# Now lets' see the distribution of approved / denied classes (0: denied, 1: approved)
print(binary_df['loan_granted'].value_counts())
# + colab={} colab_type="code" id="6h3kQmIqMLYr"
# Turn categorical string features into simple 0/1 features (like turning "sex" into "sex_male" and "sex_female")
dummies_df = pd.get_dummies(binary_df, columns=text_columns_to_keep)
dummies_df = dummies_df.sample(frac=1).reset_index(drop=True)
bad_dummies_df = pd.get_dummies(bad_binary_df, columns=text_columns_to_keep)
bad_dummies_df = bad_dummies_df.sample(frac=1).reset_index(drop=True)
# + colab={} colab_type="code" id="3VfdY4PzWOoI"
# Normalize the numeric columns so that they all have the same scale to simplify modeling/training
def normalize():
min_max_scaler = preprocessing.MinMaxScaler()
column_names_to_normalize = ['loan_amount_000s', 'applicant_income_000s', 'minority_population', 'hud_median_family_income', 'population']
x = dummies_df[column_names_to_normalize].values
x_scaled = min_max_scaler.fit_transform(x)
df_temp = pd.DataFrame(x_scaled, columns=column_names_to_normalize, index = dummies_df.index)
dummies_df[column_names_to_normalize] = df_temp
x = bad_dummies_df[column_names_to_normalize].values
x_scaled = min_max_scaler.fit_transform(x)
bad_df_temp = pd.DataFrame(x_scaled, columns=column_names_to_normalize, index = bad_dummies_df.index)
bad_dummies_df[column_names_to_normalize] = bad_df_temp
normalize()
# + [markdown] colab_type="text" id="m20NBqsMaMkx"
# ## Get the Train & Test Data
#
# Now, let's get the train and test data for our models.
#
# For the **first** model, you'll use `train_data` and `train_labels`.
#
# For the **second** model, you'll use `limited_train_data` and `limited_train_labels`.
# + colab={} colab_type="code" id="Np8JM4KINnKC"
# Get the training data & labels
test_data_with_labels = dummies_df
train_data = dummies_df
train_labels = train_data['loan_granted']
train_data = train_data.drop(columns=['loan_granted'])
# Get the bad (limited) training data and labels
limited_train_data = bad_dummies_df
limited_train_labels = limited_train_data['loan_granted']
limited_train_data = bad_dummies_df.drop(columns=['loan_granted'])
# Split the data into train / test sets for Model 1
x,y = train_data,train_labels
train_data,test_data,train_labels,test_labels = train_test_split(x,y)
# Split the bad data into train / test sets for Model 2
lim_x,lim_y=limited_train_data,limited_train_labels
limited_train_data,limited_test_data,limited_train_labels,limited_test_labels = train_test_split(lim_x,lim_y)
# + [markdown] colab_type="text" id="MyUxXszu0Mp0"
# # Create and train your TensorFlow models
#
# In this section, you will write code to train two TensorFlow Keras models.
# + [markdown] colab_type="text" id="K685pKOMUPQD"
# ## Train your first model on the complete dataset.
#
# * **Important**: your first model should be named **model**.
# * The data will come from `train_data` and `train_labels`.
#
# If you get stuck, you can view the documentation [here](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential).
# + colab={} colab_type="code" id="PvgHgZ-agsi_"
# import TF modules
from tensorflow.keras import layers
from tensorflow.keras import initializers
from tensorflow.keras import optimizers
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
# + colab={} colab_type="code" id="l4qrBBr5bUSK"
# This is the size of the array you'll be feeding into our model for each example
input_size = len(train_data.iloc[0])
# Train the first model on the complete dataset. Use `train_data` for your data and `train_labels` for you labels.
# ---- TODO ---------
# create the model = Sequential()
# model.add (your layers)
# model.compile
# model.fit
model = Sequential()
model.add(layers.Dense(8, input_dim=input_size))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(optimizer='sgd', loss='mse')
model.fit(train_data, train_labels, batch_size=32, epochs=10)
# + colab={} colab_type="code" id="CWGEGaxPgsjD"
# Save your model
# !mkdir -p saved_model
model.save('saved_model/my_model')
# + colab={} colab_type="code" id="hg0bnNVwgsjF"
# Get predictions on the test set and print the accuracy score (Model 1)
y_pred = model.predict(test_data)
acc = accuracy_score(test_labels, y_pred.round())
print("Model 1 Accuracy: %.2f%%" % (acc * 100.0))
# + [markdown] colab_type="text" id="U2hPhuA-UXTT"
# ## Train your second model on the limited datset.
#
# * **Important**: your second model should be named **limited_model**.
# * The data will come from `limited_train_data` and `limited_train_labels`.
#
#
# If you get stuck, you can view the documentation [here](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential).
# + colab={} colab_type="code" id="NP8cr7JvgsjH"
# Train your second model on the limited dataset. Use `limited_train_data` for your data and `limited_train_labels` for your labels.
# Use the same input_size for the limited_model
# ---- TODO ---------
# create the limited_model = Sequential()
# limited_model.add (your layers)
# limited_model.compile
# limited_model.fit
limited_model = Sequential()
limited_model.add(layers.Dense(8, input_dim=input_size))
limited_model.add(layers.Dense(1, activation='sigmoid'))
limited_model.compile(optimizer='sgd', loss='mse')
limited_model.fit(limited_train_data, limited_train_labels, batch_size=32, epochs=10)
# + colab={} colab_type="code" id="5UauXNlMgsjK"
# Save your model
# !mkdir -p saved_limited_model
limited_model.save('saved_limited_model/my_limited_model')
# + colab={} colab_type="code" id="n0UxiCcygsjM"
# Get predictions on the test set and print the accuracy score (Model 2)
limited_y_pred = limited_model.predict(limited_test_data)
acc = accuracy_score(limited_test_labels, limited_y_pred.round())
print("Model 2 Accuracy: %.2f%%" % (acc * 100.0))
# + [markdown] colab_type="text" id="-5X33HRf0b2C"
# # Deploy your models to the AI Platform
#
# In this section, you will first need to create a Cloud Storage bucket to store your models, then you will use gcloud commands to copy them over.
#
# You will then create two AI Platform model resources and their associated versions.
# + colab={} colab_type="code" id="Jfp8H0esC6k_"
# ---- TODO ---------
# Fill out this information:
GCP_PROJECT = 'qwiklabs-gcp-01-526d2b6d39ec'
MODEL_BUCKET = 'gs://qwiklabs-gcp-01-526d2b6d39ec'
MODEL_NAME = 'complete_model' #do not modify
LIM_MODEL_NAME = 'limited_model' #do not modify
VERSION_NAME = 'v1'
REGION = 'us-central1'
# + colab={} colab_type="code" id="NJOTCAsLDjcF"
# Copy your model files to Cloud Storage (these file paths are your 'origin' for the AI Platform Model)
# !gsutil cp -r ./saved_model $MODEL_BUCKET
# !gsutil cp -r ./saved_limited_model $MODEL_BUCKET
# + colab={} colab_type="code" id="dbGP-3qIDoza"
# Configure gcloud to use your project
# !gcloud config set project $GCP_PROJECT
# + [markdown] colab_type="text" id="V1RF5Ga_HAva"
# ## Create your first AI Platform model: **complete_model**
#
# Here's what you will need to create your AI Platform model:
#
# * Version (`VERSION_NAME`)
# * Model (`MODEL_NAME`=`complete_model`)
# * Framework (`TensorFlow`)
# * Runtime version (`2.1`)
# * Origin (directory path to your model in the Cloud Storage bucket)
# * Staging-bucket (`MODEL_BUCKET`)
# * Python version (`3.7`)
#
# + [markdown] colab_type="text" id="LGtP39EAgd9N"
# 1. You will first need to create a model resource with the name `$MODEL_NAME` and region `$REGION`.
#
# 2. Then you will create a version for your model with the information specified above.
#
# Be sure to name your first model **complete_model**.
#
# If you get stuck, you can always find the documentation for this [here](https://cloud.google.com/ai-platform/prediction/docs/deploying-models#gcloud).
#
# To use bash in the code cells, you can put a `!` before the command (as seen in cells above) and use a `$` in front of your environment variables.
# + colab={} colab_type="code" id="BSfwEaE8DpOP"
# 1. Create an AI Platform model resource for your COMPLETE model
# ---- TODO ---------
# !gcloud ai-platform models create $MODEL_NAME --regions $REGION
# + colab={} colab_type="code" id="QN6oEh1TX0Bf"
# 2. Now create a version. This will take a couple of minutes to deploy.
# ---- TODO ------
# !gcloud ai-platform versions create $VERSION_NAME \
# --model=$MODEL_NAME \
# --framework='TensorFlow' \
# --runtime-version=2.1 \
# --origin=$MODEL_BUCKET/saved_model/my_model \
# --staging-bucket=$MODEL_BUCKET \
# --python-version=3.7 \
# --project=$GCP_PROJECT
# + [markdown] colab_type="text" id="TNCuzUbsKuUv"
# ## Create your second AI Platform model: **limited_model**
#
# Here's what you will need to create your AI Platform model:
#
# * Version (`VERSION_NAME`)
# * Model (`LIM_MODEL_NAME`)
# * Framework (`TensorFlow`)
# * Runtime version (`2.1`)
# * Origin (directory path to your second model in the Cloud Storage bucket)
# * Staging-bucket (`MODEL_BUCKET`)
# * Python version (`3.7`)
# + [markdown] colab_type="text" id="kHCNMxEtgVII"
# 1. You will first need to create a model resource with the name `$LIM_MODEL_NAME` and region `$REGION`.
#
# 2. Then you will create a version for your model with the information specified above.
#
# Be sure to name your second model **limited_model**.
#
# If you get stuck, you can always find the documentation for this [here](https://cloud.google.com/ai-platform/prediction/docs/deploying-models#gcloud_1).
#
# To use bash in the code cells, you can put a `!` before the command (as seen in cells above) and use a `$` in front of your environment variables.
# + colab={} colab_type="code" id="xuPue_4Mgsjd"
# 1. Create an AI Platform model resource for your LIMITED model
# ---- TODO ---------
# !gcloud ai-platform models create $LIM_MODEL_NAME --regions $REGION
# + colab={} colab_type="code" id="IMF5kzNjYBys"
# 2. Now create a version. This will take a couple of minutes to deploy.
# ---- TODO ------
# !gcloud ai-platform versions create $VERSION_NAME \
# --model=$LIM_MODEL_NAME \
# --framework='TensorFlow' \
# --runtime-version=2.1 \
# --origin=$MODEL_BUCKET/saved_limited_model/my_limited_model \
# --staging-bucket=$MODEL_BUCKET \
# --python-version=3.7 \
# --project=$GCP_PROJECT
# + [markdown] colab_type="text" id="<KEY>"
# # Using the What-if Tool to interpret your model
# Once your models have deployed, you're now ready to connect them to the What-if Tool using the WitWidget.
#
# We've provided the Config Builder code and a couple of functions to get the class predictions from the models, which are necessary inputs for the WIT. If you've successfully deployed and saved your models, **you won't need to modify any code in this cell**.
# + colab={} colab_type="code" id="bQrAb7lbOhvI"
#@title Show model results in WIT
num_datapoints = 1000 #@param {type: "number"}
# Column indices to strip out from data from WIT before passing it to the model.
columns_not_for_model_input = [
test_data_with_labels.columns.get_loc('loan_granted'),
]
# Return model predictions.
def custom_predict(examples_to_infer):
# Delete columns not used by model
model_inputs = np.delete(
np.array(examples_to_infer), columns_not_for_model_input, axis=1).tolist()
# Get the class predictions from the model.
preds = model.predict(model_inputs)
preds = [[1 - pred[0], pred[0]] for pred in preds]
return preds
# Return 'limited' model predictions.
def limited_custom_predict(examples_to_infer):
# Delete columns not used by model
model_inputs = np.delete(
np.array(examples_to_infer), columns_not_for_model_input, axis=1).tolist()
# Get the class predictions from the model.
preds = limited_model.predict(model_inputs)
preds = [[1 - pred[0], pred[0]] for pred in preds]
return preds
examples_for_wit = test_data_with_labels.values.tolist()
column_names = test_data_with_labels.columns.tolist()
config_builder = (WitConfigBuilder(
examples_for_wit[:num_datapoints],feature_names=column_names)
.set_custom_predict_fn(limited_custom_predict)
.set_target_feature('loan_granted')
.set_label_vocab(['denied', 'accepted'])
.set_compare_custom_predict_fn(custom_predict)
.set_model_name('limited')
.set_compare_model_name('complete'))
WitWidget(config_builder, height=800)
|
Google Cloud Data Science and Machine Learning/Explore Machine Learning Models with Explainable AI/Challenge Lab/what-if-tool-challenge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # for Loops
#
# A **for** loop acts as an iterator in Python, it goes through items that are in a *sequence* or any other iterable item. Objects that we've learned about that we can iterate over include strings,lists,tuples, and even built in iterables for dictionaries, such as the keys or values.
#
# We've already seen the **for** statement a little bit in past lectures but now lets formalize our understanding.
#
# Here's the general format for a **for** loop in Python:
#
# for item in object:
# statements to do stuff
#
# The variable name used for the item is completely up to the coder, so use your best judgment for choosing a name that makes sense and you will be able to understand when revisiting your code. This item name can then be referenced inside you loop, for example if you wanted to use if statements to perform checks.
#
# Let's go ahead and work through several example of **for** loops using a variety of data object types. we'll start simple and build more complexity later on.
#
# ##Example 1
# Iterating through a list.
# We'll learn how to automate this sort of list in the next lecture
l = [1,2,3,4,5,6,7,8,9,10]
for num in l:
print num
# Great! Hopefully this makes sense. Now lets add a if statement to check for even numbers. We'll first introduce a new concept here--the modulo.
# ### Modulo
# The modulo allows us to get the remainder in a division and uses the % symbol. For example:
17 % 5
# This makes sense since 17 divided by 5 is 3 remainder 2. Let's see a few more quick examples:
# 3 Remainder 1
10 % 3
# 2 Remainder 4
18 % 7
# 2 no remainder
4 % 2
# Notice that if a number is fully divisible with no remainder, the result of the modulo call is 0. We can use this to test for even numbers, since if a number modulo 2 is equal to 0, that means it is an even number!
#
# Back to the **for** loops!
#
# ##Example 2
# Let's print only the even numbers from that list!
for num in l:
if num % 2 == 0:
print num
# We could have also put in else statement in there:
for num in l:
if num % 2 == 0:
print num
else:
print 'Odd number'
# ## Example 3
# Another common idea during a **for** loop is keeping some sort of running tally during the multiple loops. For example, lets create a for loop that sums up the list:
# +
# Start sum at zero
list_sum = 0
for num in l:
list_sum = list_sum + num
print list_sum
# -
# Great! Read over the above cell and make sure you understand fully what is going on. Also we could have implemented a += to to the addition towards the sum. For example:
# +
# Start sum at zero
list_sum = 0
for num in l:
list_sum += num
print list_sum
# -
# ## Example 4
# We've used for loops with lists, how about with strings? Remember strings are a sequence so when we iterate through them we will be accessing each item in that string.
for letter in 'This is a string.':
print letter
# ##Example 5
# Let's now look at how a for loop can be used with a tuple:
# +
tup = (1,2,3,4,5)
for t in tup:
print t
# -
# ## Example 6
# Tuples have a special quality when it comes to **for** loops. If you are iterating through a sequence that contains tuples, the item can actually be the tuple itself, this is an example of *tuple unpacking*. During the **for** loop we will be unpacking the tuple inside of a sequence and we can access the individual items inside that tuple!
l = [(2,4),(6,8),(10,12)]
for tup in l:
print tup
# Now with unpacking!
for (t1,t2) in l:
print t1
# Cool! With tuples in a sequence we can access the items inside of them through unpacking! The reason this is important is because many object will deliver their iterables through tuples. Let's start exploring iterating through Dictionaries to explore this further!
# ##Example 7
d = {'k1':1,'k2':2,'k3':3}
for item in d:
print item
# Notice how this produces only the keys. So how can we get the values? Or both the keys and the values?
#
# Here is where we are going to have a Python 3 Alert!
# ### <font color='red'>Python 3 Alert!</font>
#
# ### Python 2: Use .iteritems() to iterate through
#
# In Python 2 you should use .iteritems() to iterate through the keys and values of a dictionary. This basically creates a generator (we will get into generators later on in the course) that will generate the keys and values of your dictionary. Let's see it in action:
# Creates a generator
d.iteritems()
# Calling the items() method returns a list of tuples. Now we can iterate through them just as we did in the previous examples.
# Create a generator
for k,v in d.iteritems():
print k
print v
# ### Python 3: items()
# In Python 3 you should use .items() to iterate through the keys and values of a dictionary. For example:
# For Python 3
for k,v in d.items():
print(k)
print(v)
# You might be wondering why this worked in Python 2. This is because of the introduction of generators to Python during its earlier years. (We will go over generators and what they are in a future section, but the basic notion is that generators don't store data in memory, but instead just yield it to you as it goes through an iterable item).
#
# Originally, Python items() built a real list of tuples and returned that. That could potentially take a lot of extra memory.
#
# Then, generators were introduced to the language in general, and that method was reimplemented as an iterator-generator method named iteritems(). The original remains for backwards compatibility.
#
# One of Python 3’s changes is that items() now return iterators, and a list is never fully built. The iteritems() method is also gone, since items() now works like iteritems() in Python 2.
#
# ## Conclusion
#
# We've learned how to use for loops to iterate through tuples,lists,strings, and dictionaries. It will be an important tool for us, so make sure you know it well and understood the above examples.
#
# [More resources](http://www.tutorialspoint.com/python/python_for_loop.htm)
|
others/resources/python/intro-to-python-jupyter-notebooks-master/11-For Loops.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="19ft5ldGPWjL"
# # Load the pre-trained model and generate text, measuring its perplexity
#
# ## Pieces of code were borrowed from the following public repositories and tutorials:
# ### https://mccormickml.com/2019/07/22/BERT-fine-tuning/
# ### https://snappishproductions.com/blog/2020/03/01/chapter-9-text-generation-with-gpt-2-and-only-pytorch.html.html
# ### Hugging Face Language model fine-tuning script https://huggingface.co/transformers/v2.0.0/examples.html#language-model-fine-tuning
# ### <NAME>: Easy GPT2 fine-tuning with Hugging Face and PyTorch https://colab.research.google.com/drive/13dZVYEOMhXhkXWfvSMVM1TTtUDrT6Aeh?usp=sharing#scrollTo=EKOTlwcmxmej
# ### Code for measuring perplexity is based on https://huggingface.co/transformers/perplexity.html and taken from https://github.com/huggingface/transformers/issues/4147
# + colab={"base_uri": "https://localhost:8080/"} id="3C0184vGvIxj" outputId="53a3f5d3-bdb1-4197-a437-19ba0d600f85"
# !pip install transformers
# + id="CJqPMqXjwK0p"
import os
os.environ['CUDA_LAUNCH_BLOCKING'] = "1"
import time
import datetime
from google.colab import drive
import pandas as pd
import seaborn as sns
import numpy as np
import random
import matplotlib.pyplot as plt
% matplotlib inline
import torch
from torch.utils.data import Dataset, DataLoader, random_split, RandomSampler, SequentialSampler
torch.manual_seed(42)
from transformers import GPT2LMHeadModel, GPT2Tokenizer, GPT2Config, GPT2LMHeadModel
from transformers import AdamW, get_linear_schedule_with_warmup
# + colab={"base_uri": "https://localhost:8080/"} id="VxPLfb7ry6Li" outputId="df6e19c8-c0fa-4c3e-aff1-db2244a4858b"
# mount my Google Drive directory and access the training data located there
gdrive_dir = '/content/drive/'
data_dir = os.path.join(gdrive_dir, "'My Drive'","'Colab Notebooks'")
drive.mount(gdrive_dir, force_remount=True)
# + colab={"base_uri": "https://localhost:8080/"} id="9zpXin9OUZv9" outputId="2f09cfab-aeb3-4c90-b8be-dc4c7f21efab"
# cd '/content/drive/MyDrive/Colab Notebooks/model_save'
# + colab={"base_uri": "https://localhost:8080/"} id="f-_EegXOVLsd" outputId="efea0111-a617-400d-cfa8-0b8527f32ef2"
import os
os.chdir("/content/drive/MyDrive/Colab Notebooks/model_save")
# !ls
# + id="A7_PDwRWwa9I"
# Tell pytorch to run this model on the GPU.
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + colab={"base_uri": "https://localhost:8080/"} id="f2wJM9kSRpep" outputId="7ef9ead6-6895-4a62-b9c2-1f266fd4f2e3"
# !ls -l --block-size=K "/content/drive/MyDrive/Colab Notebooks/model_save"
# + colab={"base_uri": "https://localhost:8080/"} id="VpVptlPQQopR" outputId="37a070b7-ddf2-4078-976c-611512362055"
# !ls -l --block-size=M "/content/drive/MyDrive/Colab Notebooks/model_save/pytorch_model.bin"
# Copy the model files to a directory in your Google Drive.
# #!cp -r '/content/drive/MyDrive/Colab Notebooks/model_save' $data_dir
output_dir = '/content/drive/MyDrive/Colab Notebooks/model_save'
# # Load a trained model and vocabulary that you have fine-tuned
model = GPT2LMHeadModel.from_pretrained(output_dir)
tokenizer = GPT2Tokenizer.from_pretrained(output_dir)
model.to(device)
# + colab={"base_uri": "https://localhost:8080/"} id="F5DyZ3OeWOcm" outputId="133fd9c6-e039-4123-85fc-437dc586403b"
model.eval()
sentence = 'CSS Dimensions'
input_ids = tokenizer.encode(sentence, return_tensors='pt')
#greedy output
#output= model.generate(input_ids, max_length=1024)
#best-possible output achieved by adding more parameters
output = model.generate(input_ids, min_length=1024, max_length=1024, num_beams=5, no_repeat_ngram_size=2, top_k=50, top_p=0.95, early_stopping=True).to('cpu')
# average result param
#output = model.generate(input_ids, do_sample=True, max_length=50, top_p=0.92, top_k=0)
# + colab={"base_uri": "https://localhost:8080/"} id="yQHBSzUtXahA" outputId="00c9105b-1445-4add-d7bb-a8d1e2f67bb4"
output
# + colab={"base_uri": "https://localhost:8080/"} id="fpvRYmXlXoBR" outputId="b5d41b52-250d-48ee-90e2-84ae1c5a2803"
print(tokenizer.decode(output[0], skip_special_tokens=True))
# + [markdown] id="g_yy-KM7RW7y"
# # Perplexity
# + id="Cx1d22-8ZlB7"
import math
# + id="dKCNAexCRWlW"
def calculatePerplexity(sentence,model,tokenizer):
input_ids = torch.tensor(tokenizer.encode(sentence)).unsqueeze(0)
input_ids = input_ids.to('cpu')
with torch.no_grad():
outputs = model(input_ids, labels=input_ids)
loss, logits = outputs[:2]
return math.exp(loss)
# + colab={"base_uri": "https://localhost:8080/"} id="_eFcR6CYZc7B" outputId="e22af51e-44c5-47cc-b08f-218185a9cf86"
print(calculatePerplexity(sentence, model, tokenizer))
|
Run_Pre_Trained_GPT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib as plt
#reading data from csv files
df = pd.read_csv("com.csv")
df
#printing top two rows
df.head(2)
#only columns
df['Salary']
df.iloc[:,0:2] #index location
df.iloc[:,0:2].values
l_col = df.iloc[:,-1]
l_col.max()
l_col.mean()
l_col.min()
#any random columns
ab = df[['Name','Salary']]
ab
df.plot(x='Name', y='Salary', kind='bar')
|
PandasDF.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''pytorch'': conda)'
# name: python38364bitpytorchconda70fdc7f787194f4c972bb3207dd25917
# ---
# http://preview.d2l.ai/d2l-en/master/chapter_convolutional-modern/vgg.html
# +
from d2l import torch as d2l
import torch
from torch import nn
def vgg_block(num_convs, in_channels, out_channels):
layers=[]
for _ in range(num_convs):
layers.append(nn.Conv2d(in_channels, out_channels,
kernel_size=3, padding=1))
layers.append(nn.ReLU())
in_channels = out_channels
layers.append(nn.MaxPool2d(kernel_size=2,stride=2))
return nn.Sequential(*layers)
# -
conv_arch = ((1, 64), (1, 128), (2, 256), (2, 512), (2, 512))
# +
def vgg(conv_arch):
# The convolutional part
conv_blks=[]
in_channels=1
for (num_convs, out_channels) in conv_arch:
conv_blks.append(vgg_block(num_convs, in_channels, out_channels))
in_channels = out_channels
return nn.Sequential(
*conv_blks, nn.Flatten(),
# The fully-connected part
nn.Linear(out_channels * 7 * 7, 4096), nn.ReLU(), nn.Dropout(0.5),
nn.Linear(4096, 4096), nn.ReLU(), nn.Dropout(0.5),
nn.Linear(4096, 10))
net = vgg(conv_arch)
# + tags=[]
X = torch.randn(size=(1, 1, 224, 224))
for blk in net:
X = blk(X)
print(blk.__class__.__name__,'output shape:\t',X.shape)
# -
ratio = 4
small_conv_arch = [(pair[0], pair[1] // ratio) for pair in conv_arch]
net = vgg(small_conv_arch)
# + tags=[]
lr, num_epochs, batch_size = 0.05, 10, 128
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size, resize=224)
d2l.train_ch6(net, train_iter, test_iter, num_epochs, lr)
|
Ch07_Modern_CNN/7-2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><img src="http://i.imgur.com/sSaOozN.png" width="500"></center>
# ## Course: Computational Thinking for Governance Analytics
#
# ### Prof. <NAME>, PhD
# * Visiting Professor of Computational Policy at Evans School of Public Policy and Governance, and eScience Institute Senior Data Science Fellow, University of Washington.
# * Professor of Government and Political Methodology, Pontificia Universidad Católica del Perú.
#
# _____
#
# # Session 1: Programming Fundamentals
#
# ## Part B: Control of Execution in Python
# <a id='beginning'></a>
#
# You can not be an effective programmer, if you can not master the concept of control of execution when writing a code. I will introduce three main schemes:
#
# 1. [Conditional Execution.](#part1)
# 2. [Loops.](#part2)
# 3. [Error Handling.](#part3)
#
# I will also introduce the concept of **[comprehensions](#comprehension)** that Python supports (but not R).
#
# ----
# <a id='part1'></a>
# ## Conditional Execution
# This is how you tell the computer what part of a code to execute depending if an event is true or false.
# +
from math import sqrt
#math is the library, sqrt is the function
value=100
#condition
if value >= 0:
# what to do if condition is true:
rootValue=sqrt(value)
print (rootValue)
else:
# what to do if condition is false:
print('Sorry, I do not compute square roots of negative numbers')
# -
# Notice the condition follows *if* immediately. Notice also the use of **indentation** to indicate a group of instructions under the effect of the condition. This is very different from *R*. If you omitted the whole **else** section, the program will still run, but it will neither send any message nor value when the input is invalid.
#
# When condition is complex, besides using **&**/**|**/**~** as in pandas, you can use **and** / **or** / **not**:
# +
value=8
if (value <= 10) & (value%2==0) :
print('This is an even number less than 11')
elif (value <= 10) & (value%2>0) :
print('This is an odd number less than 11')
elif (value > 10) & (value%2>0) :
print('This is an odd number greater than 10')
else:
print('This is an even number greater than 10')
# +
#'and' does not strictly require parethesis
value=8
if value <= 10 and value%2==0 :
print('This is an even number less than 11')
elif value <= 10 and value%2>0 :
print('This is an odd number less than 11')
elif value > 10 and value%2>0 :
print('This is an odd number greater than 10')
else:
print('This is an even number greater than 10')
# -
# Notice what happens if you do not use parenthesis with the '&' (or that family)
# +
#the '&' strictly requires paranthesis!!!!
value=8
if value <= 10 & value%2==0 :
print('This is an even number less than 11')
elif value <= 10 & value%2>0:
print('This is an odd number less than 11')
elif value > 10 & value%2>0:
print('This is an odd number greater than 10')
else:
print('This is an even number greater than 10')
# -
# [Go to page beginning](#beginning)
#
# ----
# <a id='part2'></a>
#
# ## Loops
# This is how you tell the computer to do something many times (and to stop when it has to):
# +
from math import sqrt # no need for this in R
perfsqs=[9,25,100]
for cat in perfsqs: # for each value in values...
print(sqrt(cat)) # do this
#'cat' was originally value, and perfsqs was values - the names are unimportant
#for needs a container - either a list or tuple - if it were a value, for wouldn't work, but a list of a single value does
# -
# Notice that Python does not have a *sqrt* function in its base. The package **math** took care of that.
#
# You do not need to show each result, you could save the results.
# +
values=[9,25,100]
rootValues=[] # empty list, we will populate it later!
for value in values:
rootValues.append(sqrt(value)) # appending an element to the list (populating the list)
# This list started empty, now see what its elements are:
rootValues
# -
# It is evident that combining *loops* and *conditonals* we can make better programs. This code is NOT controlling well the process:
# +
values=[9,25,-100]
rootValues=[]
for value in values:
rootValues.append(sqrt(value))
# to see the results:
rootValues
# -
# Above, you saw that Python gives an error ('ValueError'), it is because _sqrt_ is not defined for negative values; then the process ended abruptly. The code below controls the execution better:
# +
values=[9,25,-100, 16, 24, -2]
rootValues=[]
for value in values:
if value >=0:
rootValues.append(sqrt(value))
else:
print('We added a missing value (None) when we received a negative input')
rootValues.append(None) #'None' will still allow us to do some mathematics
# to see the results:
rootValues
# -
# We are producing an output with the same size as input. If we omit the **else** structure, we will produce an output with smaller size than the input.
#
# You can also use **break** when you consider the execution should stop:
# +
values=[9,25,-100,144,-72]
rootValues=[]
for value in values:
# checking the value:
if value <0:
print('We need to stop, invalid value detected')
break
# you will get here if the value is not negative
rootValues.append(sqrt(value))
# to see the results:
rootValues
# -
# The code above halted the program.
# You can use **continue** when you consider the execution should not halt:
# +
import numpy as np
values=[9,None,np.nan, '1000',-100, 144,-72]
for value in values: # notice the order of 'IFs'
if value==None: # condition1
print ('missing values as input')
continue
if isinstance(value, str): #condition2
print ('string as input')
continue
if value < 0: # condition3
print ('negative value as input')
continue
print (sqrt(value), 'is the root of ',value)
#nan means missing
# -
# The _None_ and _NAN_ have a different nature:
type(None),type(np.nan)
# You use both values to denote a missing value, but NAN is common in structures containing only numbers, while None in any structure. Becareful when doing math:
10 + None
# In the previous case, Python complains because '+' can not be used to add those two different data types. It is like trying this:
10 + '10'
# As previously mentioned, nan is used with numerical data to denote missing values, so this operation is allowed:
10 + np.nan
#
# _Loops_ are also needed when you want to count the presence of a particular value:
# +
values=[9,25,-100,144,-72]
counterOfInvalids=0 # counter
for value in values:
if value <0:
counterOfInvalids +=1 #updating counter #increase by one
# to see the results:
counterOfInvalids
# -
# You may want to save particular positions (here is another difference with R):
# +
values=[9,25,-100,144,-72]
positionInvalids=[]
currentPosition=0 # this is the 'accumulator' initial position
for value in values:
if value <0:
positionInvalids.append(currentPosition)
currentPosition+=1 # becareful where you put the 'accumulator'
#So, if the value is neg, then you append the "position" to the list;
#it checks each value if there should be a position appended, and then adds 1 to the counter before checking the next number
# to see the results:
positionInvalids
# +
# testing:
for pos in positionInvalids:
print (values[pos])
#guessing that "pos" means "position" in Python
# -
# If you have boolean values, you can profit by using boolean operators:
# +
bvalues=[True,False,True,True]
for element in bvalues:
if element:
print('this guy is True')
# +
bvalues=[True,False,True,True]
for element in bvalues:
print (element)
if element:
print('this guy is True',type(element))
# -
# Notice this are not boolean:
# this is wrong
for element in bvalues:
if ~element:
print('this guy is True')
for element in bvalues:
print (element)
if ~element:
print('this guy is True',~element,type(~element))
# this is wrong
for element in bvalues:
if !element:
print('this guy is True')
#
# [Go to page beginning](#beginning)
#
# ----
# <a id='part3'></a>
#
# ## Error Handling
# We have controlled errors before, using *if-else*; however, Python has particular functions to take care of that:
# what kind of error you get:
print (sqrt(-10))
# what kind of error you get:
print (sqrt('10'))
# Python is giving different types of **errors** (*Type* and *Value*), let's use that:
values=[10,-10,'10']
for value in values:
try:
print (sqrt(value))
except ValueError:
print (value,'is a Wrong number!')
except TypeError:
print (value,'is Not even a number!!')
# [Go to page beginning](#beginning)
# ____
# <a id='comprehension'></a>
# ### Comprehensions
#
# Python has implemented ways to create data structures using a technique called comprehensions (R can not do that).
# As lists are mutable, this operation is creating a list on the run.
# +
from math import sqrt
values=[9,25,49,121]
rootsInList=[sqrt(value) for value in values] #List comprehension
rootsInList
# -
# As tuples are immutable, this operation is not creating a tuple on the run. We are in fact generating values that will later become a tuple.
values=[9,25,49,-121]
rootsInTuple=tuple(sqrt(value) for value in values if value > 0) #tuple comprehension
rootsInTuple
# Dicts can also be created that way:
values=[9,25,49,-121]
rootsInDict={value:(sqrt(value) if value > 0 else None) for value in values} #Dic comprehension
rootsInDict
# When you have a dict as input in comprehensions you can visit its values using _items()_ like this:
newDict={'name':'John', 'age':40, 'State':'WA'}
[[key,value] for key,value in newDict.items()]
# The function **zip** allows you to create tuples using parallel association:
letters=['a','b','c']
numbers=[10,20,30]
list(zip(letters,numbers))
# _Zipped_ lists are common in comprehensions:
[(number,double) for number,double in zip(numbers,np.array(numbers)**2)]
# ## Class exercises:
#
# Make a function that:
#
# 1. Create a data frame with this:
import pandas
names=["Tomás", "Pauline", "Pablo", "Bjork","Alan","Juana"]
woman=[False,True,False,False,False,True]
ages=[32,33,28,30,32,27]
country=["Chile", "Senegal", "Spain", "Norway","Peru","Peru"]
education=["Bach", "Bach", "Master", "PhD","Bach","Master"]
data={'names':names, 'woman':woman, 'ages':ages, 'country':country, 'education':education}
data
friends=pandas.DataFrame.from_dict(data)
friends
# 2. Create a list of tuples, where each tuple is a pair (name,country), using comprehensions
# 3. Implement a _for_ loop to count how many peruvian there are in the data frame. Try using **not** in one solution and **~** in another one.
# 4. Implement a _for_ loop to get the count of men. Try using **not** in one solution and **~** in another one.
# Solve this in a new Jupyter notebook, and then upload it to GitHub. Name the notebook as 'ex_controlOfEx'.
# ## Homework
# 1. Implement a _for_ loop to get the count of men that have a Bach degree in the data frame. I recommend the use of **zip** (somwehere)
# 2. Implement a _for_ loop to get the count of people whose current age is an even number.
# Solve this in a new Jupyter notebook, and then upload it to GitHub. Name the notebook as 'hw_controlOfEx'.
# _____
#
# * [Go to page beginning](#beginning)
# * [Go to REPO in Github](https://github.com/EvansDataScience/ComputationalThinking_Gov_1)
# * [Go to Course schedule](https://evansdatascience.github.io/GovernanceAnalytics/)
|
.ipynb_checkpoints/S1_B_Py_controlOfExecution-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Credit Risk Resampling Techniques
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
from pathlib import Path
from collections import Counter
# # Read the CSV into DataFrame
# Load the data
file_path = Path('Resources/lending_data.csv')
df = pd.read_csv(file_path)
df.head()
# # Split the Data into Training and Testing
# +
# Create our features
X_columns = [i for i in df.columns if i not in ('homeowner','loan_status')]
X= df[X_columns]
# Create our target
target= ['loan_status']
y = df.loc [:, target]
# -
X.describe()
# Check the balance of our target values
y['loan_status'].value_counts()
# +
# Create X_train, X_test, y_train, y_test
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test= train_test_split(X, y, random_state=1, stratify=y)
X_train.shape
# -
# ## Data Pre-Processing
#
# Scale the training and testing data using the `StandardScaler` from `sklearn`. Remember that when scaling the data, you only scale the features data (`X_train` and `X_testing`).
# Create the StandardScaler instance
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
# +
# Fit the Standard Scaler with the training data
# When fitting scaling functions, only train on the training dataset
X_scaler = scaler.fit(X_train)
# +
# Scale the training and testing data
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# -
# # Simple Logistic Regression
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(solver='lbfgs', random_state=1)
model.fit(X_train, y_train)
# Calculated the balanced accuracy score
from sklearn.metrics import balanced_accuracy_score
y_pred = model.predict(X_test)
balanced_accuracy_score(y_test, y_pred)
# Display the confusion matrix
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test, y_pred)
# Print the imbalanced classification report
from imblearn.metrics import classification_report_imbalanced
print(classification_report_imbalanced(y_test, y_pred))
# # Oversampling
#
# In this section, you will compare two oversampling algorithms to determine which algorithm results in the best performance. You will oversample the data using the naive random oversampling algorithm and the SMOTE algorithm. For each algorithm, be sure to complete the folliowing steps:
#
# 1. View the count of the target classes using `Counter` from the collections library.
# 3. Use the resampled data to train a logistic regression model.
# 3. Calculate the balanced accuracy score from sklearn.metrics.
# 4. Print the confusion matrix from sklearn.metrics.
# 5. Generate a classication report using the `imbalanced_classification_report` from imbalanced-learn.
#
# Note: Use a random state of 1 for each sampling algorithm to ensure consistency between tests
# ### Naive Random Oversampling
# +
# Resample the training data with the RandomOversampler
from imblearn.over_sampling import RandomOverSampler
ros = RandomOverSampler(random_state=1)
X_random, y_random = ros.fit_resample(X_train, y_train)
# View the count of target classes with Counter
Counter(y_random)
# -
# Train the Logistic Regression model using the resampled data
model_random = LogisticRegression(solver='lbfgs', random_state=1)
model.fit(X_random, y_random)
# Calculated the balanced accuracy score
y_random_pred = model.predict(X_test)
balanced_accuracy_score(y_test, y_random_pred)
# Display the confusion matrix
confusion_matrix(y_test, y_random_pred)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_random_pred))
# ### SMOTE Oversampling
# +
# Resample the training data with SMOTE
from imblearn.over_sampling import SMOTE
X_smote, y_smote = SMOTE(random_state=1, sampling_strategy=1.0).fit_resample(X_train, y_train)
# View the count of target classes with Counter
Counter(y_smote)
# -
# Train the Logistic Regression model using the resampled data
model_smote = LogisticRegression(solver='lbfgs', random_state=1)
model_smote.fit(X_smote, y_smote)
# Calculated the balanced accuracy score
y_smote_pred = model_smote.predict(X_test)
balanced_accuracy_score(y_test, y_smote_pred)
# Display the confusion matrix
confusion_matrix(y_test, y_smote_pred)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_smote_pred))
# # Undersampling
#
# In this section, you will test an undersampling algorithm to determine which algorithm results in the best performance compared to the oversampling algorithms above. You will undersample the data using the Cluster Centroids algorithm and complete the folliowing steps:
#
# 1. View the count of the target classes using `Counter` from the collections library.
# 3. Use the resampled data to train a logistic regression model.
# 3. Calculate the balanced accuracy score from sklearn.metrics.
# 4. Display the confusion matrix from sklearn.metrics.
# 5. Generate a classication report using the `imbalanced_classification_report` from imbalanced-learn.
#
# Note: Use a random state of 1 for each sampling algorithm to ensure consistency between tests
# +
# Resample the data using the ClusterCentroids resampler
from imblearn.under_sampling import ClusterCentroids
cc = ClusterCentroids(random_state=1)
X_cluster, y_cluster = cc.fit_resample(X_train, y_train)
# View the count of target classes with Counter
Counter(y_cluster)
# -
# Train the Logistic Regression model using the resampled data
model_cluster = LogisticRegression(solver='lbfgs', random_state=1)
model_cluster.fit(X_cluster, y_cluster)
# Calculate the balanced accuracy score
y_cluster_pred = model_cluster.predict(X_test)
balanced_accuracy_score(y_test, y_cluster_pred)
# Display the confusion matrix
confusion_matrix(y_test, y_cluster_pred)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_cluster_pred))
# # Combination (Over and Under) Sampling
#
# In this section, you will test a combination over- and under-sampling algorithm to determine if the algorithm results in the best performance compared to the other sampling algorithms above. You will resample the data using the SMOTEENN algorithm and complete the folliowing steps:
#
# 1. View the count of the target classes using `Counter` from the collections library.
# 3. Use the resampled data to train a logistic regression model.
# 3. Calculate the balanced accuracy score from sklearn.metrics.
# 4. Display the confusion matrix from sklearn.metrics.
# 5. Generate a classication report using the `imbalanced_classification_report` from imbalanced-learn.
#
# Note: Use a random state of 1 for each sampling algorithm to ensure consistency between tests
# +
# Resample the training data with SMOTEENN
from imblearn.combine import SMOTEENN
sm = SMOTEENN(random_state=1)
X_smoteenn, y_smoteenn = sm.fit_resample(X, y)
# View the count of target classes with Counter
Counter(y_smoteenn)
# -
# Train the Logistic Regression model using the resampled data
model_smoteenn = LogisticRegression(solver='lbfgs', random_state=1)
model_smoteenn.fit(X_smoteenn, y_smoteenn)
# Calculate the balanced accuracy score
y_smoteenn_pred = model.predict(X_test)
balanced_accuracy_score(y_test, y_smoteenn_pred)
# Display the confusion matrix
confusion_matrix(y_test, y_smoteenn_pred)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_smoteenn_pred))
# # Final Questions
#
# 1. Which model had the best balancedaccuracy score?
#
# Naive Oversampling, SMOTE and SMOTEENN had the same balanced accuracy scores.
#
#
# 2. Which model had the best recall score?
#
# All models had the same recall scores
#
#
# 3. Which model had the best geometric mean score?
#
# All models had the same geometric mean score.
|
Starter_Code/credit_risk_resampling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Double-slit model
# ## AMCDawes
# A model of the interference between a plane-wave LO and the far-field double-slit output. The FFT is computed to model what we expect to measure in our experimental setup. Physically accurate parameters have been chosen.
#
# Comments:
# - The three-peak Fourier output is fairly consistent over a wide range of parameters
# - LO is a plane-wave at first, then a Gaussian beam later on in the notebook.
# - LO is at an angle, signal is normal-incident. This is not the same as
# the experiment, but easier to treat numerically.
#
# June 2021: fixed sign error in E_LO for plane wave, verified values for experimental data. Next: simulate multiple frames of data collection with phase variation.
# +
import matplotlib.pyplot as plt
from numpy import pi, sin, cos, linspace, exp, real, imag, abs, conj, meshgrid, log, log10, angle
from numpy.fft import fft, fftshift, ifft
from mpl_toolkits.mplot3d import axes3d
import BeamOptics as bopt
# -
# %matplotlib inline
# +
b=.08*1e-3 # the slit width
a=.5*1e-3 # the slit spacing
k=2*pi/(795*1e-9) # longitudinal wavenumber
wt=0 # let time be zero
C=1 # unit amplitude
L=1.8 # distance from slits to CCD
d=.016 # distance from signal to LO at upstream end (used to calculate k_perp)
ccdwidth = 1300 # number of pixels
pixwidth = 20e-6 # pixel width (in meters)
y = linspace(-pixwidth*ccdwidth/2,pixwidth*ccdwidth/2,ccdwidth)
# -
def alpha(y,a):
return k*a*y/(2*L)
def beta(y,b):
return k*b*y/(2*L)
def E_ds(y,a,b):
""" Double-slit field """
# From Hecht p 458:
#return b*C*(sin(beta(y)) / beta(y)) * (sin(wt-k*L) + sin(wt-k*L+2*alpha(y)))
# drop the time-dep term as it will average away:
return 2*b*C*(sin(beta(y,b)) / beta(y,b)) * cos(alpha(y,a)) #* sin(wt - k*L + alpha(y))
def E_dg(y,a,b):
""" Double gaussian field """
# The width needs to be small enough to see interference
# otherwise the beam doesn't diffract and shows no interference.
# We're using b for the gaussian width (i.e. equal to the slit width)
w=b
#return C*exp(1j*k*0.1*d*y/L)
return 5e-3*(bopt.gaussian_beam(0,y-a/2,L,E0=1,wavelambda=795e-9,w0=w,k=[0,0,k]) +
bopt.gaussian_beam(0,y+a/2,L,E0=1,wavelambda=795e-9,w0=w,k=[0,0,k]))
def E_lo(y,d):
"""Plane-wave LO beam incident at small angle, transverse wavenumber k*d*y/L"""
return C*exp(-1j*k*d*y/L)
# ### Sanity check: plot the field:
plt.plot(y,abs(E_ds(y,a,b)))
plt.title("Double slit field")
# + inputHidden=false outputHidden=false
plt.plot(y,abs(E_dg(y,a,b)))
plt.title("Double-Gaussian field")
# -
# ## Define a single function to explore the FFT:
# + inputHidden=false outputHidden=false
def plotFFT(d,a,b):
"""Single function version of generating the FFT output"""
TotalField = E_dg(y,a,b)+E_lo(y,d)
TotalIntensity=TotalField*TotalField.conj()
plt.plot(abs(fft(TotalIntensity)),".-")
plt.ylim([0,1e-2])
plt.xlim([0,650])
plt.title("FFT output")
# + inputHidden=false outputHidden=false
plotFFT(d=0.046,a=0.5e-3,b=0.08e-3)
# -
# This agrees well with Matt's code using symbolic calculations. The main difference I see is in the size of the low-frequency peak. It's much smaller here than in his version.
# ## d=0.035
plotFFT(d=0.035,a=0.5e-3,b=0.08e-3)
# ## d=0.02
plotFFT(d=0.02,a=0.5e-3,b=0.08e-3)
# ## Double slit is still very different:
# + inputHidden=false outputHidden=false
def plotFFTds(d,a,b):
"""Single function version of generating the FFT output"""
TotalField = E_ds(y,a,b)+E_lo(y,d)
TotalIntensity=TotalField*TotalField.conj()
plt.plot(abs(fft(TotalIntensity)),".-")
plt.ylim([0,0.1])
plt.xlim([400,500])
plt.title("FFT output")
# + inputHidden=false outputHidden=false
plotFFTds(d=0.025,a=0.5e-3,b=0.08e-3)
# -
# This does not agree with experimental results.
# ### Replace with Gaussian LO: import gaussian beam function, and repeat:
# +
# bopt.gaussian_beam?
# -
# bopt.gaussian_beam(x, y, z, E0, wavelambda, w0, k)
# set to evaluate gaussian at L (full distance to CCD) with waist width of 2 cm
# using d=0.046 for agreement with experiment
d=0.046
E_lo_gauss = bopt.gaussian_beam(0,y,L,E0=1,wavelambda=795e-9,w0=0.02,k=[0,k*d/L,k])
plt.plot(y,abs(E_lo_gauss))
TotalIntensity=(E_dg(y,a,b)+E_lo_gauss) * (E_dg(y,a,b)+E_lo_gauss).conj()
# +
plt.figure(figsize=(14,4))
plt.plot(y,TotalIntensity,".-")
#plt.xlim([-.002,0])
# +
plt.plot(abs(fft(TotalIntensity)),".-")
plt.ylim([0,0.01]) # Had to lower the LO power quite a bit, and then zoom way in.
plt.xlim([0,650])
# -
|
Double Slit Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Examples</h1>
# <strong>Example #1: Basic If Else Example</strong>
# +
age = 25
if age > 21:
print('You can serve this person beer.')
elif 21 > age >=18:
print('No beer. Mark hand with X.')
else:
print('Call the bouncer and have this person thrown out.')
# -
# <strong>Example #2: Nested If Else Example</strong>
# +
age = 21
all_ages_show = True
birthday_on_id_equals_today = True
if age >= 21:
print('You can serve this person beer.')
if birthday_on_id_equals_today:
print('HAPPY BIRTHDAY! FREE SHOTS!')
elif 21 > age >=18:
if all_ages_show:
print('No beer. Mark hand with X.')
else:
print('Call the bouncer and have this person thrown out.')
else:
print('Why is there a child in the bar?!')
# -
# Copyright © 2020, Mass Street Analytics, LLC. All Rights Reserved.
|
02 Basics/08-control-flow-if-else.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Vantage-AI case project
#
# For this project we'll be using both R and python.
# R is used for more exploratory analyses and less automated analysis work. Once we understand the data and have
# a clear idea of which rules to implement for data cleaning, then we can continue working with Python to set up pipelines with the `scikit-learn` library.
#
# The goal of this case is to predict the need of repair of water pumps, effectively a fairly standard classification problem.
# What makes it hard is that there are many features that consists of an intractable amount of labels, meaning we can't just use them as categories since many categories will only have 1 or 2 observations; there won't be enough observations per 'category' to train a model
#
# ## Reticulate package
#
# The `reticulate` package is used to run python code. This way we can do some basic computation steps by writing python code within our R script, making it easier to port the resulting code to dedicated Python scripts to automate the analysis from raw data to the final result.
#
# After a LOT of trial and error the reticulate package now works pretty cleanly with `conda`. You can use `conda` as a package manager for both R and Python, as opposed to R package manager `packrat` or the widely used `virtualenv` for python.
#
# ## Project directory structure
#
# The directory structure was taken from the BigData Republic GitHub! By doing this I hope to learn more about the preferred project structure at BDR.
#
# The cookiecutter-data-science repository can be found here https://github.com/BigDataRepublic/cookiecutter-data-science. Now, what makes this truly come full circle is that...
#
# - The Tanzania Water Pump data was originally hosted by **DrivenData** for a data science competition
# * Hosted here https://www.drivendata.org/competitions/7/pump-it-up-data-mining-the-water-table/
# * Original data made available by [Taarifa](http://taarifa.org) and [Tanzania Ministry of Water](http://maji.go.tz/)
# - The Github repository from BDR is actually a fork from https://github.com/drivendata/cookiecutter-data-science !
# * Changes include getting rid of `virtualenv` , thus only using `conda`
# * Source code directory named after the project instead of just `src`
#
# While the directory structure is completely overkill for a project like this, it still serves its purpose as a standardized data science project directory. Even if project are small, finding the code isn't usually straightforward... Unless you already know the file structure, then it becomes a piece of cake!
#
# # Exploratory analysis
#
# Before we move to modelling and actual data cleaning, let's explore the data to get a feeling for the data.
# Load libraries
library(tidyverse)
library(broom)
library(ggthemes)
library(DT)
# Library to use python code using our conda environment
library(reticulate)
py_config()
# # Predicting Tanzania water pump maintenance
#
# ### The data
#
# The case presented here is about water pumps in Tanzania. The data set originates from the Tanzanian Ministry of Water. Currently, this ministry maintains its pumps based on a maintenance schedule or, of course, when they break down. We feel that the maintenance of the Tanzanian water pumps could improve in both the cost of maintenance and the prevention of break downs by introducing machine learning to predict if a water pump is in need of repair or even the moment of failure of each water pump.
#
# ### Objective
#
# Our objective is to develop a reproducible model that can predict which pumps will fail in the future, either the moment or just whether they fail. The resulting model is to be used by the Tanzanian government to effectively maintain their water pumps.
#
# Normally for this type of data you would expect that a single water pump was measured
# over-time and using this data we could predict the need for repairs for a given pump. In this case however, at
# every practically every timepoint a different water point was measured! So we have to make some assumptions about the
# data and the approach to how we create a model.
#
# #### Assumptions
#
# **We should know in advance which water point should be predicted**, there must be something like a
# maintenance schedule at least 3 days ahead. We make this assumption because we have to able to take
# into account the measured variables about the water pumps.!
#
# **Measured variable should all be assumed to be known in advance**, otherwise we should choose
# to leave out some variables that may have only become known at the time of determining the functioning of a water pump.
# Variables like the `funder` and `district_code` are clearly known ahead of maintenance, but other variables like
# `water_quality` and `amount_tsh` (total amount water available to waterpoint) may not be known before determining the state of the pump.
#
# # Getting a feeling for the data
#
# Before we start modelling the data we should understand the data. In this dataset
# we find a total of 40 measured variables to be used as features in our models. In this
# section we will aim to understand these features and make motivated choices to keep, remove or transform these
# features from our prediction model.
#
# ## A closer look at our features
#
# Our data consists of,
# - `r nrow(df)` observations
# - `r ncol(df)` features
# - `r length(unique(labels[,2]))` types of labels to predict
#
# Not all of the features are necessarily useful for predicting the moment of failure of water pumps,
# for instance the measurement `id` won't be of much use at it is unique.
#
# First we'll take a look at the raw data to fully understand the structure of our data,
# especially how exactly the data was measured over time. We have `r nrow(df)` observations.
#
# ### The raw data
#
# The raw data is not considered big-data in the sense that it will fit on a single machine; we do not have to worry about batch processing because of sheer file size with 'only' `r nrow(df)` observations. The number of features is also a mere `r ncol(df)`! Enough to make a interpretable predictive model and this will allow us to dive deeper into the
# individual variables.
# Read local data
# Expect to be synced with data hosten on Amazon S3
data <- read.csv("../../data/raw/water_pump_set.csv")
labels <- read.csv("../../data/raw/water_pump_labels.csv")
features <- read.csv("../../data/raw/water_pump_features.csv")
# Merge labels and features into single dataset by id
df <- merge(data, labels, by="id")
head(df)
# 39 features,to use for modelling a unique id and the status_group labels
# Total of 59400 observations
dim(df)
# Show the feature descriptions
# This excludes the 'status'group'
head(features)
# +
# Let's add some more info to the feature info using tidyverse piping
feature_desc <- features %>% filter(Feature %in% colnames(data)) %>%
mutate(example = sapply(data[1, -1], as.character), # First row of the data without the 'id' column
unique = map_int(data[,-1],function(x) length(unique(x))),
type = map_chr(data[,-1], function(x) class(unlist(x))), #
nonzero = map_int(data[,-1], function(x) sum(as.character(x) != "0")), # Nonzero values, assuming 0 usually means NA
max_single = map_int(data[,-1], function(x) max(table(x))), # Observations of most occuring value
max_occurence = map_chr(data[,-1], function(x) names(which.max(table(x))))) # Highest occuring value
# Let's glance at the numeric values in the dataset
feature_desc %>% filter(type %in% c("numeric", "integer"))
feature_desc %>% filter(type %in% c("numeric", "integer")) %>% dim()
# -
# The number of unique values for each numerical feature is surprising in some cases. Continuous values like,
# - `gps_height`
# - `population`
# - `amount_tsh`
#
# all have a very low number of unique values. So, even though we have not seen any `NA` values in the data, but there must be some missing values. These values are simply set to `0`, it is unclear to me whether we can distinguish true `0` values from missing values.
#
# ### Non-numeric summaries
# Show non-numeric value summary
feature_desc %>% filter(!type %in% c("numeric", "integer"))
feature_desc %>% filter(!type %in% c("numeric", "integer")) %>% dim()
# ### Handling feature groups
#
# The variables in the data are far from independent, some variables practically mean the same thing.
# Some feature groups are so painfully obvious that we should clean these manually before moving on to
# more generic analyses.
#
# In the `Description` column, we can see some overlap in values. These features are likely to represent the same
# variable, so we can reduce the dimensionality of the dataset just by smartly combining some variables.
#
# Let's start with common sense and group some of the variables manually to prepare the data
# for more automated pipelines. Here are some logical groups we can check,
#
# - scheme_name, scheme_management, installer, funder
# - extraction_type, extraction_type_group, extraction_type_class
# - management, management_group
# - scheme_management, scheme_name
# - payment, payment_type
# - water_quality, quality_group
# - quantity_quantity_group
# - source, source_class
# - waterpoint_type, waterpoint_type_group
#
# #### Scheme_name, scheme_management, installer, funder
#
#
data %>%
select(scheme_name, scheme_management, installer, funder) %>%
mutate_all(as.character) %>%
mutate_all(tolower) %>% # To lowercase since there are many typing 'errors' or inconsistencies
group_by(scheme_name, scheme_management, installer, funder) %>%
tally() %>%
arrange(desc(n)) %>%
head(10)
# Scheme names
data %>% select(scheme_name) %>% table() %>% sort(decreasing = T) %>% head()
# Half the data is missing data, and the rest is extremely scattered
# Furthermore there are many unambiguous names and spelling errors like 'Adra', 'ADRA', 'WOULD BANK', etc..
df %>%
filter(scheme_name != "") %>%
group_by(scheme_name, status_group) %>%
tally() %>% head(20)
# Based on this we remove the scheme_name entirely
# +
# Scheme management
data %>% select(scheme_management) %>% table() %>% sort(decreasing = T) %>% head()
# Plot label per scheme_management
df %>% select(scheme_management, status_group) %>%
ggplot(aes(x = scheme_management, fill = status_group)) +
geom_bar() + coord_flip() +
ggtitle("Water pomp status voor management_scheme") +
theme_fivethirtyeight()
# Most of the water pumps are from a handful of installers.
# We could consider grouping all the low occurences into a group 'other'
# Plot label per scheme_management
scheme_management_groups = df %>% select(scheme_management, status_group) %>%
group_by(scheme_management) %>%
tally()
cat(sprintf("Scheme mangement in 10 groups accounts for %1.2f%% of the data. And just VWC is %1.2f%%",
100 * sum(scheme_management_groups$n) / nrow(df),
100 * max(scheme_management_groups$n) / nrow(df)))
# +
# installer
data %>% select(installer) %>% table() %>% sort(decreasing = T) %>% head()
length(unique(data$installer))
# Most of the water pumps are from a handful of installers.
# We could consider grouping all the low occurences into a group 'other'
# Plot label per scheme_management
installer_groups = df %>% select(installer, status_group) %>%
group_by(installer) %>%
tally() %>%
top_n(20, n)
# Plot installer versus status_group to verify
df %>% select(installer, status_group) %>%
mutate(installer = ifelse(installer %in% installer_groups$installer, as.character(installer), "other"),
installer = ifelse(installer %in% c("", "0"), "unknown", installer)) %>%
ggplot(aes(x = installer, fill = status_group)) +
geom_bar() + coord_flip() +
ggtitle("Water pomp status voor installer", subtitle = "Kleine groepen samengevoegd in 'other'") +
theme_fivethirtyeight()
# Show counts of installer labes, only top20
# Notice that many of these have about the same number of observations and grouping all the others into
# 'other' is quite arbitrary since the 'other' group will still be half the data.
# The number of 'larger' groups and the number of spelling errors in the data require more detailed feature engineering
# like some kind of string matching- which we will not spend our time on.
# THIS FEATURE WILL BE REMOVED FROM THE MODELLING DATA
# +
# funder
data %>% select(funder) %>% table() %>% sort(decreasing = T) %>% head()
cat(sprintf("Of the %i unique funder names, %i overlap with the installer names!",
length(unique(data$funder)),
length(intersect(tolower(as.character(data$funder)), tolower(as.character(data$installer))))))
# Due to the high overlap between funder and installer, we can assume that these two are heavily related and
# will probably give us the same difficulty as the funder names by themselves.
# WE REMOVE THIS FEATURE FROM THE DATASET!
# +
# Groups for each extraction type to determine which has the most information
df %>% select(extraction_type, extraction_type_group, extraction_type_class) %>%
apply(2, function(x) length(unique(x)))
# Count label co-occurences to potentially remove some features as we have a
# tractable number of features (not 100s this time!)
df %>%
group_by(extraction_type, extraction_type_group, extraction_type_class) %>%
tally()
# -
# By far the most prevalent type of waterpump is the gravity water pump type. This type of pump is eco-friendly as it does not use energy other than gravity.
#
# Notice that there are some very small groups in the data, mostly due to the inclusion of `extraction_type`. The `extraction_type` has some values that we could combine with other categories to reduce the dimensionality.
#
# - **india mark ii** and **india mark iii** can be combined to just **india mark**
# - **cemo** and **climax** are just two types of motorpumps, let's not differentiate these two
# - **other - mkulima/shinyanga** is very specific, we can just classify this as handpump together with **play pump** and **walimi**
# - Lastly the **swn 81** and **swn 80** are both handpumps, we can classifiy these as **swn**
# * The SWN 80 and SWN 81 type of handle pumps only differ in their pump head so that the SWN 81 can reach deeper wells. [Read more about handpumps here:)](https://www.ircwash.org/sites/default/files/232.2-13249.pdf)
# * Fun fact, the SWN stands for 'Sociale Werkplaats Nunspeet'! The SWN family of pumps was introduced partly by a workshop for the handicapped in the Netherlands sine 1976.
#
# Let's manually combine some of the feature groups into related groups.
# +
# Temporary data about the extraction type
df_extraction <- df %>%
mutate(
extraction_type = as.character(extraction_type),
extraction_type = ifelse(extraction_type %in% c("india mark ii", "india mark iii"), "india mark", extraction_type),
extraction_type = ifelse(extraction_type %in% c("cemo", "climax"), "motorpump", extraction_type),
extraction_type = ifelse(extraction_type %in% c("other - swn 81", "swn 80"), "swn", extraction_type),
extraction_type = ifelse(extraction_type %in% c('other - mkulima/shinyanga','other - play pump', 'walimi'), "handpump", extraction_type)
) %>%
select(id, extraction_type, extraction_type_class)
df_extraction %>% group_by(extraction_type, extraction_type_class) %>% tally()
# Make a new table of group counts
df_extraction %>%
mutate(extraction_group = paste(extraction_type, extraction_type_class, sep="-")) %>%
group_by(extraction_group, extraction_type, extraction_type_class) %>%
tally() %>%
arrange(n) %>%
ungroup() %>%
mutate(extraction_type = factor(extraction_type, levels=extraction_type)) %>% # Relevel to reorder plot
ggplot(aes(x = extraction_type, y = n, label = n, fill = extraction_type_class)) +
geom_bar(stat="identity", position="dodge") +
ggtitle("Water pomp types na combineren van groepen") +
geom_text() + coord_flip() + theme_fivethirtyeight()
# -
# Using two out of three extraction types, after some adjustments, in the data we can classify all the data in a way that none of the groups is truly underrepresented.
# #### management, management_group
#
# These two variables seem to mean the exact same thing.
# Cross table of only two variables
table(df %>% select(management, management_group))
# The management variable has more information than management_group
# We will select the column with *more* information, so we keep the `management` variable and drop the `management_group`.
#
# #### Payment and payment_type
#
# The payment and payment type columns also mean the exact same thing.
# Cross table of only two variables
df %>% group_by(payment, payment_type) %>% tally()
# The payments are exactly the same, we can drop either one.
# The payment_type variables are more concise so we will drop the `payment` column
# +
# Cross table of only two variables
df %>% group_by(water_quality, quality_group) %>% tally()
# -
# The water quality intuitively seems like it would be predictive of the water pump status.
# +
# Is water quality correlated to the water pump status?
df %>% group_by(water_quality, status_group) %>%
tally() %>%
group_by(water_quality) %>% mutate(n = n / sum(n)) %>%
filter(water_quality != "soft") %>%
ggplot(aes(x = water_quality, y = n, fill = status_group)) +
geom_bar(stat="identity") + coord_flip() +
ggtitle("Water kwaliteit versus pomp status ratios") +
theme_fivethirtyeight()
# Is water amount available correlated to the water quality?
df %>% group_by(water_quality) %>%
filter(amount_tsh > 100) %>%
ggplot(aes(x = water_quality, y = amount_tsh, col = water_quality)) +
geom_boxplot() + coord_flip() +
ggtitle("Water kwaliteit versus pomp status ratios") +
theme_fivethirtyeight()
# -
# The `water_quality` contains more information so we will drop the `quality_group`.
#
# The `water_quality` is also related to the `amount_tsh` in the data, even though we are removing `amount_tsh`(!) The only pumps where a significant amount of water is available is the ones where 'soft' water was found. So even though we are leaving out `amount_tsh`, we will still capture some of its information by including the `water_quality`.
#
# We could improve the visualisation above but it will likely not affect the choices we already made, so let's move on...
#
# #### Quantity and quantity_group
#
# The `quantity` and `quantity_group` sounds like they have the same relation as the water quality. Let's check!
#
# +
# Cross table of only two variables
df %>% group_by(quantity, quantity_group) %>% tally()
# -
# welp.. we checked, and we'll just leave out the `quantity_group` column.
#
# #### Source, source_class and source_type
# Cross table of only two variables
df %>% group_by(source, source_class, source_type) %>% tally()
# The water `source` is more detailed than `source_class` and contains the exact same values as `source_type`. So let's keep only the `source` and `source_type`.
# However, the `source_class` contains only 3 categories and this more general distinction may in fact help us make predictions.
# We can calculate the feature importance later, so let's keep both variables in our data and verify whether inclusion of `source_class` improves the model at all.
#
# Alternatively,
# Is water quality correlated to the water pump status?
df %>% group_by(source, source_class, status_group) %>%
tally() %>%
group_by(source) %>% mutate(n = n / sum(n)) %>%
ggplot(aes(x = source, y = n, fill = status_group)) +
geom_bar(stat="identity") + coord_flip() +
ggtitle("Water bron versus pomp status ratios") +
theme_fivethirtyeight()
# The source is definitely not a perfect prediction of water pump status, but at least a distinction between classes could be beneficial. It remains to be seen whether the water source is truly indicative of the water pump status, why would a pump near a lake be more likely to be non-functional than that near a spring? Unless the water source has dried up, but this would rather affect multiple pumps in the same area.
#
# If there was more time available we could find if there is just a cluster of water pumps at a single lake that are non-functional because of regional differences, for example. The location of a pump probably determines the water source, and since we will already take the gps-coordinates into account we might not be including any new information with the `source` feature.
#
# #### Waterpoint_type and waterpoint_type_group
#
# Finally, this group of two variables may be reduced to a single variable.
# Cross table of only two variables
df %>% group_by(waterpoint_type, waterpoint_type_group) %>% tally()
# They are the exact same variable. So we remove the waterpoint_type_group from our data, later.
# #### Conclusion about dealing with grouped data variables
#
# **Remove 9 features**
# - `scheme_name`, `scheme_management` , too many unique
# - `extraction_type_group`, overlaps with `extraction_type` and `extraction_class`
# - `funder` and `installer`, many missing values, unique values and data-entry inconsistencies
# - `source_class`, `payment`, `waterpoint_type_group`, `management_group` because they are duplicates of other variables
#
# **Edit 1 feature**
# - `extraction_type`, rename some similar categories with very low occurences
#
# Let's make a note of these findings. We will cleanly integrate these steps into our data preprocessing pipeline.
#
# +
df_combined_groups <- df %>%
select(-c(scheme_name, scheme_management, extraction_type_group, funder,
installer, source_class, payment, waterpoint_type_group, management_group)) %>%
mutate(
extraction_type = as.character(extraction_type),
extraction_type = ifelse(extraction_type %in% c("india mark ii", "india mark iii"), "india mark", extraction_type),
extraction_type = ifelse(extraction_type %in% c("cemo", "climax"), "motorpump", extraction_type),
extraction_type = ifelse(extraction_type %in% c("other - swn 81", "swn 80"), "swn", extraction_type),
extraction_type = ifelse(extraction_type %in% c('other - mkulima/shinyanga','other - play pump', 'walimi'), "handpump", extraction_type)
)
dim(df_combined_groups)
# Write df_combined_groups to csv
# NOTE THIS IS ACTUALLY HAPPENING IN DEDICATED SCRIPTS, NOT FROM THIS NOTEBOOK
#df %>% write_csv("../../data/interim/data_grouped_vars.csv")
# This part was rewritten into vantage-project/vantage/data/clean_data.py
# The exploration is done in R but final code execution in Python
# -
# # Missing values
#
# Now that we have removed some categorical values that we have checked because they seemed grouped with other variables, we still have to take a look at the remaining variables.
#
# While there are no true `NA` values in the data, we have already seen cases where missing values are indicated by empty values, zeroes, or labels like 'none'. Let us look at the feature descriptions again, and take note of the number of unique values and maximum occuring value.
feature_desc %>% filter(Feature %in% colnames(df_combined_groups))
# ## amount_tsh
#
# We'll **remove** this column as it consists mostly of zeroes, and there is no way for us to distinguish whether these are missing values or just dried up water pumps. I expect that not every included water pump was actually measured as detailed as every other pump. By far the most values are zeroes.
table(df_combined_groups$amount_tsh)[1:10]
# ## wpt_name
#
# The `wpt_name`, or waterpump name, many unique values and the most occuring values is 'none'.
# Now, even the pumps with the same exact name don't actually seem to guarantee the same pump.
#
# This is reason enough to **remove** this feature as unique labels like this will only aid to overfit the model.
# Show some values with wpt_name==Zahanati
# Notice that these are not the same waterpoint at all!
# So wpt_name is not a variable we'll be able to use
df %>% filter(wpt_name == "Zahanati") %>% select(wpt_name, everything())
# ## num_private
# The `num_private`, which came without a detailed description, has a loooooottt of zero values.
# Again, we cannot differentiate between true 0 values and missing values, since 0 does seem like possible value for `num_private`. We remove this column from further analysis.
# Ratio of values in the data
table(df$num_private)[1:5] / nrow(df) * 100
# ## construction_year
#
# Missing values of `construction_year` are indicated by zero-values. How to replace these is not necessarily straightforward since we treat this as a continuous variable! Since the difference between 1955 and 2000 is much smaller than 1955 and 0 we should be careful here! If the construction year is predictive of the water pump status then zero-values will likely introduce a bias.
#
# For missing values we should definitely impute some value. Traditionally the most straightforward way is to just impute the mean or median value. We may correct for region or pump type to improve this imputation. A quick glance at factors that may indicate a missing construction_year we don't find any. So let us use a very simple imputation method to save time.
# +
df %>% filter(construction_year > 0) %>%
ggplot(aes(x = construction_year, color = construction_year)) + geom_bar() +
ggtitle("Distribution of non-missing construction years",
sprintf("Number of excluded missing construction years: %i", sum(df$construction_year == 0))) +
theme_fivethirtyeight()
median(df$construction_year[df$construction_year > 0])
# Replace construction_year with median of non-zero construction years
df_combined_groups %>%
mutate(construction_year = replace(construction_year,
construction_year==0,
median(construction_year[construction_year>0])))
# -
# ### GPS coordinates
#
# There are three types of gps coordinates in the data,
#
# - gps_height
# - longitude
# - latitude
#
# We expect these to be continuous values.
#
#
# See which gps coordinates occur more than once to reveal missing values
# Maybe some duplicate gps coordinates?? Guess not..
df %>% group_by(gps_height, longitude, latitude) %>%
tally() %>% filter(n > 2)
#
#
# Of all the observations we only find that 1812 observations are missing values, where
# gps_height, longitude and latitude are all 0.
#
# We could try to deal with these values in a number of ways,
#
# - Impute values based on other location data (region_code, district, etc.)
# - Ignore missing values as it only comprises small part of the data
# - At least center the water pumps to somewhere in Tanzania
#
# Of course this has varying levels of complexity and for now we simply ignore missing values
# and leave them as is(!) These are fairly hard to deal with and they will probably not have any more value than the other location data such as `region`, `basin` and `district`.
#
# **We leave these as-is.**
#
#
|
notebooks/dev/data-exploration-r.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # HSV Color Space, Balloons
# ### Import resources and display image
import numpy as np
import matplotlib.pyplot as plt
import cv2
# +
# %matplotlib inline
# Read in the image
image = cv2.imread('images/water_balloons.jpg')
# Make a copy of the image
image_copy = np.copy(image)
# Change color to RGB (from BGR)
image = cv2.cvtColor(image_copy, cv2.COLOR_BGR2RGB)
plt.imshow(image)
# -
# ### Plot color channels
# +
# RGB channels
r = image[:,:,0]
g = image[:,:,1]
b = image[:,:,2]
f, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,10))
ax1.set_title('Red')
ax1.imshow(r, cmap='gray')
ax2.set_title('Green')
ax2.imshow(g, cmap='gray')
ax3.set_title('Blue')
ax3.imshow(b, cmap='gray')
# +
# Convert from RGB to HSV
hsv = cv2.cvtColor(image, cv2.COLOR_RGB2HSV)
# HSV channels
h = hsv[:,:,0]
s = hsv[:,:,1]
v = hsv[:,:,2]
f, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,10))
ax1.set_title('Hue')
ax1.imshow(h, cmap='gray')
ax2.set_title('Saturation')
ax2.imshow(s, cmap='gray')
ax3.set_title('Value')
ax3.imshow(v, cmap='gray')
# -
print(h[400:410,300:310])
# ### Define pink and hue selection thresholds
# Define our color selection criteria in HSV values
lower_hue = np.array([160,0,0])
upper_hue = np.array([180,255,255])
# Define our color selection criteria in RGB values
lower_pink = np.array([180,0,100])
upper_pink = np.array([255,255,230])
# ### Mask the image
# +
# Define the masked area in RGB space
mask_rgb = cv2.inRange(image, lower_pink, upper_pink)
# mask the image
masked_image = np.copy(image)
masked_image[mask_rgb==0] = [0,0,0]
# Vizualize the mask
plt.imshow(masked_image)
# -
print(hsv[300:305,400:405,:])
# +
# Now try HSV!
# Define the masked area in HSV space
mask_hsv = cv2.inRange(hsv, lower_hue, upper_hue)
# plt.imshow(mask_hsv,cmap='gray')
# mask the image
masked_image = np.copy(image)
masked_image[mask_hsv ==0 ] = [0,0,0]
# Vizualize the mask
plt.imshow(masked_image)
# -
|
1_1_Image_Representation/5_1. HSV Color Space, Balloons.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="eJGtmni-DezY" colab_type="text"
# # Lambda School Data Science Module 141
# ## Statistics, Probability, and Inference
# + [markdown] id="FMhDKOFND0qY" colab_type="text"
# ## Prepare - examine what's available in SciPy
#
# As we delve into statistics, we'll be using more libraries - in particular the [stats package from SciPy](https://docs.scipy.org/doc/scipy/reference/tutorial/stats.html).
# + id="fQ9rkLJmEbsk" colab_type="code" outputId="937d6c40-d775-4016-9b69-70a82cc8b4c0" colab={"base_uri": "https://localhost:8080/", "height": 4427}
from scipy import stats
dir(stats)
# + id="bxW4SG_gJGlZ" colab_type="code" outputId="e715ad1a-883f-41e2-b070-a1106316f4e7" colab={"base_uri": "https://localhost:8080/", "height": 70}
# As usual, lots of stuff here! There's our friend, the normal distribution
norm = stats.norm()
print(norm.mean())
print(norm.std())
print(norm.var())
# + id="RyNKPt_tJk86" colab_type="code" outputId="db64f558-1945-4fef-f7d7-3184212d8237" colab={"base_uri": "https://localhost:8080/", "height": 70}
# And a new friend - t
t1 = stats.t(5) # 5 is df "shape" parameter
print(t1.mean())
print(t1.std())
print(t1.var())
# + [markdown] id="SRn1zMuaKgxX" colab_type="text"
# 
#
# *(Picture from [Wikipedia](https://en.wikipedia.org/wiki/Student's_t-distribution#/media/File:Student_t_pdf.svg))*
#
# The t-distribution is "normal-ish" - the larger the parameter (which reflects its degrees of freedom - more input data/features will increase it), the closer to true normal.
# + id="seQv5unnJvpM" colab_type="code" outputId="b2f84397-b204-4864-84a1-2b29eb926bbf" colab={"base_uri": "https://localhost:8080/", "height": 70}
t2 = stats.t(30) # Will be closer to normal
print(t2.mean())
print(t2.std())
print(t2.var())
# + [markdown] id="FOvEGMysLaE2" colab_type="text"
# Why is it different from normal? To better reflect the tendencies of small data and situations with unknown population standard deviation. In other words, the normal distribution is still the nice pure ideal in the limit (thanks to the central limit theorem), but the t-distribution is much more useful in many real-world situations.
#
# History sidenote - this is "Student":
#
# 
#
# *(Picture from [Wikipedia](https://en.wikipedia.org/wiki/File:William_Sealy_Gosset.jpg))*
#
# His real name is <NAME>, and he published under the pen name "Student" because he was not an academic. He was a brewer, working at Guinness and using trial and error to determine the best ways to yield barley. He's also proof that, even 100 years ago, you don't need official credentials to do real data science!
# + [markdown] id="1yx_QilAEC6o" colab_type="text"
# ## Live Lecture - let's perform and interpret a t-test
#
# We'll generate our own data, so we can know and alter the "ground truth" that the t-test should find. We will learn about p-values and how to interpret "statistical significance" based on the output of a hypothesis test. We will also dig a bit deeper into how the test statistic is calculated based on the sample error, and visually what it looks like to have 1 or 2 "tailed" t-tests.
# + id="BuysRPs-Ed0v" colab_type="code" colab={}
# TODO - during class, but please help!
# + [markdown] id="egXb7YpqEcZF" colab_type="text"
# ## Assignment - apply the t-test to real data
#
# Your assignment is to determine which issues have "statistically significant" differences between political parties in this [1980s congressional voting data](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records). The data consists of 435 instances (one for each congressperson), a class (democrat or republican), and 16 binary attributes (yes or no for voting for or against certain issues). Be aware - there are missing values!
#
# Your goals:
#
# 1. Load and clean the data (or determine the best method to drop observations when running tests)
# 2. Using hypothesis testing, find an issue that democrats support more than republicans with p < 0.01
# 3. Using hypothesis testing, find an issue that republicans support more than democrats with p < 0.01
# 4. Using hypothesis testing, find an issue where the difference between republicans and democrats has p > 0.1 (i.e. there may not be much of a difference)
#
# Note that this data will involve *2 sample* t-tests, because you're comparing averages across two groups (republicans and democrats) rather than a single group against a null hypothesis.
#
# Stretch goals:
#
# 1. Refactor your code into functions so it's easy to rerun with arbitrary variables
# 2. Apply hypothesis testing to your personal project data (for the purposes of this notebook you can type a summary of the hypothesis you formed and tested)
# + id="nstrmCG-Ecyk" colab_type="code" colab={}
# TODO - your code here!
# + [markdown] id="wiq83guLcuAE" colab_type="text"
# # Resources
#
# - https://homepage.divms.uiowa.edu/~mbognar/applets/t.html
# - https://rpsychologist.com/d3/tdist/
# - https://gallery.shinyapps.io/tdist/
# - https://en.wikipedia.org/wiki/Standard_deviation#Sample_standard_deviation_of_metabolic_rate_of_northern_fulmars
|
module1-statistics-probability-and-inference/LS_DS_141_Statistics_Probability_and_Inference.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit
# name: python3
# ---
# Using scikit-learn to perform K-Means clustering
from sklearn.cluster import KMeans
from sklearn.datasets import load_digits
digits = load_digits()
digits.data.shape
kmeans = KMeans(n_clusters=10, random_state=77)
prediction = kmeans.fit_predict(digits.data)
prediction
kmeans.cluster_centers_.shape
# Scatter plot of the data point
import matplotlib.pyplot as plt
fig, ax = plt.subplots(2, 5, figsize=(8, 3))
centers = kmeans.cluster_centers_.reshape(10, 8, 8)
for axi, center in zip(ax.flat, centers):
axi.set(xticks=[], yticks=[])
axi.imshow(center, interpolation='nearest', cmap=plt.cm.binary)
import numpy as np
from scipy.stats import mode
labels = np.zeros_like(prediction)
for i in range(10):
mask = (prediction == i)
labels[mask] = mode(digits.target[mask])[0]
from sklearn.metrics import accuracy_score
accuracy_score(digits.target, labels)
from sklearn.metrics import confusion_matrix
import seaborn as sns
mat = confusion_matrix(digits.target, labels)
ax = sns.heatmap(mat.T, square=True, annot=True, fmt='d', cbar=False,xticklabels=digits.target_names,yticklabels=digits.target_names)
ax.set_ylim(10.0,0)
plt.xlabel('true label')
plt.ylabel('predicted label');
|
lab8/KMeans_Clustering_LabeledData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:universe]
# language: python
# name: conda-env-universe-py
# ---
# # Building a Video Game Bot using OpenAI Universe
# Let us learn how to build a video game bot which plays car racing game. Our objective is
# that car has to move forward without getting stuck by any obstacles and hitting other cars.
# First, we import necessary libraries,
import gym
import universe
import random
# Then we simulate our car racing environment by make function.
env = gym.make('flashgames.NeonRace-v0')
env.configure(remotes=1)
# And let us create variables for moving the car,
# +
# Move left
left = [('KeyEvent', 'ArrowUp', True), ('KeyEvent', 'ArrowLeft', True),
('KeyEvent', 'ArrowRight', False)]
# Move right
right = [('KeyEvent', 'ArrowUp', True), ('KeyEvent', 'ArrowLeft', False),
('KeyEvent', 'ArrowRight', True)]
# Move forward
forward = [('KeyEvent', 'ArrowUp', True), ('KeyEvent', 'ArrowRight', False),
('KeyEvent', 'ArrowLeft', False), ('KeyEvent', 'n', True)]
# -
# Followed by, we will initialize some other variables
# +
# We use turn variable for deciding whether to turn or not
turn = 0
# We store all the rewards in rewards list
rewards = []
# we will use buffer as some kind of threshold
buffer_size = 100
# We set our initial action has forward i.e our car moves just forward without making any turns
action = forward
# -
# Now, let us begin our game agent to play in an infinite loop which continuously performs an action based on interaction with the environment.
while True:
turn -= 1
# Let us say initially we take no turn and move forward.
# First, We will check the value of turn, if it is less than 0
# then there is no necessity for turning and we just move forward
if turn <= 0:
action = forward
turn = 0
action_n = [action for ob in observation_n]
# Then we use env.step() to perform an action (moving forward for now) one-time step
observation_n, reward_n, done_n, info = env.step(action_n)
# store the rewards in the rewards list
rewards += [reward_n[0]]
# We will generate some random number and if it is less than 0.5 then we will take right, else
# we will take left and we will store all the rewards obtained by performing each action and
# based on our rewards we will learn which direction is the best over several timesteps.
if len(rewards) >= buffer_size:
mean = sum(rewards)/len(rewards)
if mean == 0:
turn = 20
if random.random() < 0.5:
action = right
else:
action = left
rewards = []
env.render()
|
Chapter02/2.09 Building a Video Game Bot .ipynb
|
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
#skip
! [ -e /content ] && pip install -Uqq fastai # upgrade fastai on colab
#export
from fastai.basics import *
#export
_all_ = ['SuggestionMethod']
#hide
from nbdev.showdoc import *
# +
#default_exp callback.schedule
# -
# # Hyperparam schedule
#
# > Callback and helper functions to schedule any hyper-parameter
from fastai.test_utils import *
# ## Annealing
#export
class _Annealer:
def __init__(self, f, start, end): store_attr('f,start,end')
def __call__(self, pos): return self.f(self.start, self.end, pos)
#export
def annealer(f):
"Decorator to make `f` return itself partially applied."
@functools.wraps(f)
def _inner(start, end): return _Annealer(f, start, end)
return _inner
# This is the decorator we will use for all of our scheduling functions, as it transforms a function taking `(start, end, pos)` to something taking `(start, end)` and return a function depending of `pos`.
# +
#export
#TODO Jeremy, make this pickle
#@annealer
#def SchedLin(start, end, pos): return start + pos*(end-start)
#@annealer
#def SchedCos(start, end, pos): return start + (1 + math.cos(math.pi*(1-pos))) * (end-start) / 2
#@annealer
#def SchedNo (start, end, pos): return start
#@annealer
#def SchedExp(start, end, pos): return start * (end/start) ** pos
#
#SchedLin.__doc__ = "Linear schedule function from `start` to `end`"
#SchedCos.__doc__ = "Cosine schedule function from `start` to `end`"
#SchedNo .__doc__ = "Constant schedule function with `start` value"
#SchedExp.__doc__ = "Exponential schedule function from `start` to `end`"
# +
#export
def sched_lin(start, end, pos): return start + pos*(end-start)
def sched_cos(start, end, pos): return start + (1 + math.cos(math.pi*(1-pos))) * (end-start) / 2
def sched_no (start, end, pos): return start
def sched_exp(start, end, pos): return start * (end/start) ** pos
def SchedLin(start, end): return _Annealer(sched_lin, start, end)
def SchedCos(start, end): return _Annealer(sched_cos, start, end)
def SchedNo (start, end): return _Annealer(sched_no, start, end)
def SchedExp(start, end): return _Annealer(sched_exp, start, end)
SchedLin.__doc__ = "Linear schedule function from `start` to `end`"
SchedCos.__doc__ = "Cosine schedule function from `start` to `end`"
SchedNo .__doc__ = "Constant schedule function with `start` value"
SchedExp.__doc__ = "Exponential schedule function from `start` to `end`"
# -
#hide
tst = pickle.dumps(SchedCos(0, 5))
annealings = "NO LINEAR COS EXP".split()
p = torch.linspace(0.,1,100)
fns = [SchedNo, SchedLin, SchedCos, SchedExp]
#export
def SchedPoly(start, end, power):
"Polynomial schedule (of `power`) function from `start` to `end`"
def _inner(pos): return start + (end - start) * pos ** power
return _inner
for fn, t in zip(fns, annealings):
plt.plot(p, [fn(2, 1e-2)(o) for o in p], label=t)
f = SchedPoly(2,1e-2,0.5)
plt.plot(p, [f(o) for o in p], label="POLY(0.5)")
plt.legend();
show_doc(SchedLin)
sched = SchedLin(0, 2)
test_eq(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.5, 1., 1.5, 2.])
show_doc(SchedCos)
sched = SchedCos(0, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.29289, 1., 1.70711, 2.])
show_doc(SchedNo)
sched = SchedNo(0, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0., 0., 0., 0.])
show_doc(SchedExp)
sched = SchedExp(1, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [1., 1.18921, 1.41421, 1.68179, 2.])
show_doc(SchedPoly)
sched = SchedPoly(0, 2, 2)
test_close(L(map(sched, [0., 0.25, 0.5, 0.75, 1.])), [0., 0.125, 0.5, 1.125, 2.])
# +
p = torch.linspace(0.,1,100)
pows = [0.5,1.,2.]
for e in pows:
f = SchedPoly(2, 0, e)
plt.plot(p, [f(o) for o in p], label=f'power {e}')
plt.legend();
# -
#export
def combine_scheds(pcts, scheds):
"Combine `scheds` according to `pcts` in one function"
assert sum(pcts) == 1.
pcts = tensor([0] + L(pcts))
assert torch.all(pcts >= 0)
pcts = torch.cumsum(pcts, 0)
pct_lim = len(pcts) - 2
def _inner(pos):
idx = min((pos >= pcts).nonzero().max(), pct_lim)
actual_pos = (pos-pcts[idx]) / (pcts[idx+1]-pcts[idx])
return scheds[idx](actual_pos.item())
return _inner
# `pcts` must be a list of positive numbers that add up to 1 and is the same length as `scheds`. The generated function will use `scheds[0]` from 0 to `pcts[0]` then `scheds[1]` from `pcts[0]` to `pcts[0]+pcts[1]` and so forth.
p = torch.linspace(0.,1,100)
f = combine_scheds([0.3,0.7], [SchedCos(0.3,0.6), SchedCos(0.6,0.2)])
plt.plot(p, [f(o) for o in p]);
p = torch.linspace(0.,1,100)
f = combine_scheds([0.3,0.2,0.5], [SchedLin(0.,1.), SchedNo(1.,1.), SchedCos(1., 0.)])
plt.plot(p, [f(o) for o in p]);
#hide
test_close([f(0.), f(0.15), f(0.3), f(0.4), f(0.5), f(0.7), f(1.)],
[0., 0.5, 1., 1., 1., 0.65451, 0.])
#export
def combined_cos(pct, start, middle, end):
"Return a scheduler with cosine annealing from `start`→`middle` & `middle`→`end`"
return combine_scheds([pct,1-pct], [SchedCos(start, middle), SchedCos(middle, end)])
# This is a useful helper function for the [1cycle policy](https://sgugger.github.io/the-1cycle-policy.html). `pct` is used for the `start` to `middle` part, `1-pct` for the `middle` to `end`. Handles floats or collection of floats. For example:
f = combined_cos(0.25,0.5,1.,0.)
plt.plot(p, [f(o) for o in p]);
#hide
test_close([f(0.), f(0.1), f(0.25), f(0.5), f(1.)], [0.5, 0.67275, 1., 0.75, 0.])
f = combined_cos(0.25, np.array([0.25,0.5]), np.array([0.5,1.]), np.array([0.,0.]))
for a,b in zip([f(0.), f(0.1), f(0.25), f(0.5), f(1.)],
[[0.25,0.5], [0.33638,0.67275], [0.5,1.], [0.375,0.75], [0.,0.]]):
test_close(a,b)
# ## ParamScheduler -
#export
@docs
class ParamScheduler(Callback):
"Schedule hyper-parameters according to `scheds`"
order,run_valid = 60,False
def __init__(self, scheds): self.scheds = scheds
def before_fit(self): self.hps = {p:[] for p in self.scheds.keys()}
def before_batch(self): self._update_val(self.pct_train)
def _update_val(self, pct):
for n,f in self.scheds.items(): self.opt.set_hyper(n, f(pct))
def after_batch(self):
for p in self.scheds.keys(): self.hps[p].append(self.opt.hypers[-1][p])
def after_fit(self):
if hasattr(self.learn, 'recorder') and hasattr(self, 'hps'): self.recorder.hps = self.hps
_docs = {"before_fit": "Initialize container for hyper-parameters",
"before_batch": "Set the proper hyper-parameters in the optimizer",
"after_batch": "Record hyper-parameters of this batch",
"after_fit": "Save the hyper-parameters in the recorder if there is one"}
# `scheds` is a dictionary with one key for each hyper-parameter you want to schedule, with either a scheduler or a list of schedulers as values (in the second case, the list must have the same length as the the number of parameters groups of the optimizer).
learn = synth_learner()
sched = {'lr': SchedLin(1e-3, 1e-2)}
learn.fit(1, cbs=ParamScheduler(sched))
n = len(learn.dls.train)
test_close(learn.recorder.hps['lr'], [1e-3 + (1e-2-1e-3) * i/n for i in range(n)])
#hide
#test discriminative lrs
def _splitter(m): return [[m.a], [m.b]]
learn = synth_learner(splitter=_splitter)
sched = {'lr': combined_cos(0.5, np.array([1e-4,1e-3]), np.array([1e-3,1e-2]), np.array([1e-5,1e-4]))}
learn.fit(1, cbs=ParamScheduler(sched))
show_doc(ParamScheduler.before_fit)
show_doc(ParamScheduler.before_batch)
show_doc(ParamScheduler.after_batch)
show_doc(ParamScheduler.after_fit)
#export
@patch
def fit_one_cycle(self:Learner, n_epoch, lr_max=None, div=25., div_final=1e5, pct_start=0.25, wd=None,
moms=None, cbs=None, reset_opt=False):
"Fit `self.model` for `n_epoch` using the 1cycle policy."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr_max is None else lr_max)
lr_max = np.array([h['lr'] for h in self.opt.hypers])
scheds = {'lr': combined_cos(pct_start, lr_max/div, lr_max, lr_max/div_final),
'mom': combined_cos(pct_start, *(self.moms if moms is None else moms))}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
# The 1cycle policy was introduced by <NAME> et al. in [Super-Convergence: Very Fast Training of Neural Networks Using Large Learning Rates](https://arxiv.org/abs/1708.07120). It schedules the learning rate with a cosine annealing from `lr_max/div` to `lr_max` then `lr_max/div_final` (pass an array to `lr_max` if you want to use differential learning rates) and the momentum with cosine annealing according to the values in `moms`. The first phase takes `pct_start` of the training. You can optionally pass additional `cbs` and `reset_opt`.
#Integration test: training a few epochs should make the model better
learn = synth_learner(lr=1e-2)
xb,yb = learn.dls.one_batch()
init_loss = learn.loss_func(learn.model(xb), yb)
learn.fit_one_cycle(2)
xb,yb = learn.dls.one_batch()
final_loss = learn.loss_func(learn.model(xb), yb)
assert final_loss < init_loss
#Scheduler test
lrs,moms = learn.recorder.hps['lr'],learn.recorder.hps['mom']
test_close(lrs, [combined_cos(0.25,1e-2/25,1e-2,1e-7)(i/20) for i in range(20)])
test_close(moms, [combined_cos(0.25,0.95,0.85,0.95)(i/20) for i in range(20)])
#export
@patch
def plot_sched(self:Recorder, keys=None, figsize=None):
keys = self.hps.keys() if keys is None else L(keys)
rows,cols = (len(keys)+1)//2, min(2, len(keys))
figsize = figsize or (6*cols,4*rows)
_, axs = plt.subplots(rows, cols, figsize=figsize)
axs = axs.flatten() if len(keys) > 1 else L(axs)
for p,ax in zip(keys, axs):
ax.plot(self.hps[p])
ax.set_ylabel(p)
#hide
#test discriminative lrs
def _splitter(m): return [[m.a], [m.b]]
learn = synth_learner(splitter=_splitter)
learn.fit_one_cycle(1, lr_max=slice(1e-3,1e-2))
#n = len(learn.dls.train)
#est_close(learn.recorder.hps['lr'], [1e-3 + (1e-2-1e-3) * i/n for i in range(n)])
learn = synth_learner()
learn.fit_one_cycle(2)
learn.recorder.plot_sched()
#export
@patch
def fit_flat_cos(self:Learner, n_epoch, lr=None, div_final=1e5, pct_start=0.75, wd=None,
cbs=None, reset_opt=False):
"Fit `self.model` for `n_epoch` at flat `lr` before a cosine annealing."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr is None else lr)
lr = np.array([h['lr'] for h in self.opt.hypers])
scheds = {'lr': combined_cos(pct_start, lr, lr, lr/div_final)}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
learn = synth_learner()
learn.fit_flat_cos(2)
learn.recorder.plot_sched()
#export
@patch
def fit_sgdr(self:Learner, n_cycles, cycle_len, lr_max=None, cycle_mult=2, cbs=None, reset_opt=False, wd=None):
"Fit `self.model` for `n_cycles` of `cycle_len` using SGDR."
if self.opt is None: self.create_opt()
self.opt.set_hyper('lr', self.lr if lr_max is None else lr_max)
lr_max = np.array([h['lr'] for h in self.opt.hypers])
n_epoch = cycle_len * (cycle_mult**n_cycles-1)//(cycle_mult-1)
pcts = [cycle_len * cycle_mult**i / n_epoch for i in range(n_cycles)]
scheds = [SchedCos(lr_max, 0) for _ in range(n_cycles)]
scheds = {'lr': combine_scheds(pcts, scheds)}
self.fit(n_epoch, cbs=ParamScheduler(scheds)+L(cbs), reset_opt=reset_opt, wd=wd)
# This schedule was introduced by I<NAME>. in [SGDR: Stochastic Gradient Descent with Warm Restarts](https://arxiv.org/abs/1608.03983). It consists of `n_cycles` that are cosine annealings from `lr_max` (defaults to the `Learner` lr) to 0, with a length of `cycle_len * cycle_mult**i` for the `i`-th cycle (first one is `cycle_len`-long, then we multiply the length by `cycle_mult` at each epoch). You can optionally pass additional `cbs` and `reset_opt`.
# +
#slow
learn = synth_learner()
with learn.no_logging(): learn.fit_sgdr(3, 1)
test_eq(learn.n_epoch, 7)
iters = [k * len(learn.dls.train) for k in [0,1,3,7]]
for i in range(3):
n = iters[i+1]-iters[i]
#The start of a cycle can be mixed with the 0 of the previous cycle with rounding errors, so we test at +1
test_close(learn.recorder.lrs[iters[i]+1:iters[i+1]], [SchedCos(learn.lr, 0)(k/n) for k in range(1,n)])
learn.recorder.plot_sched()
# -
#export
@patch
@delegates(Learner.fit_one_cycle)
def fine_tune(self:Learner, epochs, base_lr=2e-3, freeze_epochs=1, lr_mult=100,
pct_start=0.3, div=5.0, **kwargs):
"Fine tune with `Learner.freeze` for `freeze_epochs`, then with `Learner.unfreeze` for `epochs`, using discriminative LR."
self.freeze()
self.fit_one_cycle(freeze_epochs, slice(base_lr), pct_start=0.99, **kwargs)
base_lr /= 2
self.unfreeze()
self.fit_one_cycle(epochs, slice(base_lr/lr_mult, base_lr), pct_start=pct_start, div=div, **kwargs)
learn.fine_tune(1)
# ## LRFind -
#export
@docs
class LRFinder(ParamScheduler):
"Training with exponentially growing learning rate"
def __init__(self, start_lr=1e-7, end_lr=10, num_it=100, stop_div=True):
if num_it < 6: num_it = 6
self.scheds = {'lr': [SchedExp(s, e) for (s,e) in zip(start_lr,end_lr)
] if is_listy(start_lr) else SchedExp(start_lr, end_lr)}
self.num_it,self.stop_div = num_it,stop_div
def before_fit(self):
super().before_fit()
path = self.path/self.model_dir
path.mkdir(parents=True, exist_ok=True)
self.tmp_d = tempfile.TemporaryDirectory(dir=path)
self.tmp_p = Path(self.tmp_d.name).stem
self.learn.save(f'{self.tmp_p}/_tmp')
self.best_loss = float('inf')
def before_batch(self): self._update_val(self.train_iter/self.num_it)
def after_batch(self):
super().after_batch()
if self.smooth_loss < self.best_loss: self.best_loss = self.smooth_loss
if self.smooth_loss > 4*self.best_loss and self.stop_div: raise CancelFitException()
if self.train_iter >= self.num_it: raise CancelFitException()
def before_validate(self): raise CancelValidException()
def after_fit(self):
self.learn.opt.zero_grad() # Needed before detaching the optimizer for future fits
tmp_f = self.path/self.model_dir/self.tmp_p/'_tmp.pth'
if tmp_f.exists():
self.learn.load(f'{self.tmp_p}/_tmp', with_opt=True)
self.tmp_d.cleanup()
_docs = {"before_fit": "Initialize container for hyper-parameters and save the model",
"before_batch": "Set the proper hyper-parameters in the optimizer",
"after_batch": "Record hyper-parameters of this batch and potentially stop training",
"after_fit": "Save the hyper-parameters in the recorder if there is one and load the original model",
"before_validate": "Skip the validation part of training"}
# +
#cuda
from fastai.vision.all import *
set_seed(99, True)
path = untar_data(URLs.PETS)/'images'
image_files = get_image_files(path)
if sys.platform == "win32" and IN_NOTEBOOK:
image_files = random.choices(image_files, k=int(len(image_files)/8))
print("Randomly select 1/8 files in NOTEBOOK on Windows to save time")
# pickle can't serializer lamda function.
def _label_func(x):
return x[0].isupper()
dls = ImageDataLoaders.from_name_func(
path, image_files, valid_pct=0.2,
label_func=_label_func, item_tfms=Resize(224))
learn = cnn_learner(dls, resnet18)
learn.fit(1)
learn.opt.state_dict()['state'][1]['grad_avg']
# -
#slow
with tempfile.TemporaryDirectory() as d:
learn = synth_learner(path=Path(d))
init_a,init_b = learn.model.a,learn.model.b
with learn.no_logging(): learn.fit(20, cbs=LRFinder(num_it=100))
assert len(learn.recorder.lrs) <= 100
test_eq(len(learn.recorder.lrs), len(learn.recorder.losses))
#Check stop if diverge
if len(learn.recorder.lrs) < 100: assert learn.recorder.losses[-1] > 4 * min(learn.recorder.losses)
#Test schedule
test_eq(learn.recorder.lrs, [SchedExp(1e-7, 10)(i/100) for i in range_of(learn.recorder.lrs)])
#No validation data
test_eq([len(v) for v in learn.recorder.values], [1 for _ in range_of(learn.recorder.values)])
#Model loaded back properly
test_eq(learn.model.a, init_a)
test_eq(learn.model.b, init_b)
test_eq(learn.opt.state_dict()['state'], [{}, {}])
show_doc(LRFinder.before_fit)
show_doc(LRFinder.before_batch)
show_doc(LRFinder.after_batch)
show_doc(LRFinder.before_validate)
# ### Suggestion Methods
# There are a few methodologies for suggesting a learning rate automatically and these as we will see can further be passed into `lr_find`. Currently four methods are supported, however to write your own it should look like a function that can accept `LRFinder`'s returned `lrs`, `losses`, as well as the `num_it`.
# Your function should return an `x,y` coordinate that can be plotted, such as below:
#
#
# ```python
# def myfunc(lrs:list, losses:list, num_it:int) -> tuple(float, tuple(float,int)):
# ...
# return suggestion, (suggestion,loss_idx)
# ```
#
# If there are any more parameters to be passed in, you should pass in your `func` as a partial and specify them yourself, such as:
#
# ```python
# def myfunc(lrs:list, losses:list, num_it:int, pct_reduction:float) -> tuple(float, tuple(float,int)):
# ...
# return suggestion, (suggestion,loss_idx)
# ```
#
# ```python
# f = partial(myfunc, pct_reduction=.2)
# ```
# +
#hide
learn = synth_learner()
with learn.no_logging(): learn.fit(20, cbs=LRFinder(num_it=100))
lrs,losses = tensor(learn.recorder.lrs[100//10:-5]),tensor(learn.recorder.losses[100//10:-5])
# -
#export
def valley(lrs:list, losses:list, num_it:int):
"Suggests a learning rate from the longest valley and returns its index"
n = len(losses)
max_start, max_end = 0,0
# find the longest valley
lds = [1]*n
for i in range(1,n):
for j in range(0,i):
if (losses[i] < losses[j]) and (lds[i] < lds[j] + 1):
lds[i] = lds[j] + 1
if lds[max_end] < lds[i]:
max_end = i
max_start = max_end - lds[max_end]
sections = (max_end - max_start) / 3
idx = max_start + int(sections) + int(sections/2)
return float(lrs[idx]), (float(lrs[idx]), losses[idx])
doc(valley)
# The `valley` algorithm was developed by [ESRI](https://forums.fast.ai/t/automated-learning-rate-suggester/44199/30) and takes the steepest slope roughly 2/3 through the longest valley in the LR plot, and is also the default for `Learner.lr_find`
#hide
valley(lrs, losses, 100)
#export
def slide(lrs:list, losses:list, num_it:int, lr_diff:int=15, thresh:float=.005, adjust_value:float=1.):
"Suggests a learning rate following an interval slide rule and returns its index"
losses = to_np(losses)
loss_grad = np.gradient(losses)
r_idx = -1
l_idx = r_idx - lr_diff
local_min_lr = lrs[l_idx]
while (l_idx >= -len(losses)) and (abs(loss_grad[r_idx] - loss_grad[l_idx]) > thresh):
local_min_lr = lrs[l_idx]
r_idx -= 1
l_idx -= 1
suggestion = float(local_min_lr) * adjust_value
idx = np.interp(np.log10(suggestion), np.log10(lrs), losses)
return suggestion, (suggestion, idx)
doc(slide)
# The `slide` rule is an algorithm developed by <NAME> out of Novetta, and is detailed [here](https://forums.fast.ai/t/automated-learning-rate-suggester/44199?u=muellerzr).
#hide
slide(lrs, losses, 100)
#export
def minimum(lrs:list, losses:list, num_it:int):
"Suggests a learning rate one-tenth the minumum before divergance and returns its index"
lr_min = lrs[losses.argmin()].item()
loss_idx = losses[min(range(len(lrs)), key=lambda i: abs(lrs[i]-lr_min))]
return lr_min/10, (lr_min, loss_idx)
#hide
minimum(lrs, losses, 100)
doc(minimum)
#export
def steep(lrs:list, losses:list, num_it:int) -> (float, tuple):
"Suggests a learning rate when the slope is the steepest and returns its index"
grads = (losses[1:]-losses[:-1]) / (lrs[1:].log()-lrs[:-1].log())
lr_steep = lrs[grads.argmin()].item()
loss_idx = losses[min(range(len(lrs)), key=lambda i: abs(lrs[i]-lr_steep))]
return lr_steep, (lr_steep, loss_idx)
doc(steep)
#hide
steep(lrs, losses, 100)
#export
@patch
def plot_lr_find(self:Recorder, skip_end=5, return_fig=True, suggestions=None, nms=None, **kwargs):
"Plot the result of an LR Finder test (won't work if you didn't do `learn.lr_find()` before)"
lrs = self.lrs if skip_end==0 else self.lrs [:-skip_end]
losses = self.losses if skip_end==0 else self.losses[:-skip_end]
fig, ax = plt.subplots(1,1)
ax.plot(lrs, losses)
ax.set_ylabel("Loss")
ax.set_xlabel("Learning Rate")
ax.set_xscale('log')
if suggestions:
colors = plt.rcParams['axes.prop_cycle'].by_key()['color'][1:]
for (val, idx), nm, color in zip(suggestions, nms, colors):
ax.plot(val, idx, 'o', label=nm, c=color)
ax.legend(loc='best')
#export
mk_class("SuggestionMethod", **{o.__name__.capitalize():o for o in [valley,slide,minimum,steep]},
doc="All possible suggestion methods as convience attributes to get tab-completion and typo-proofing")
#export
@patch
def lr_find(self:Learner, start_lr=1e-7, end_lr=10, num_it=100, stop_div=True, show_plot=True, suggest_funcs=(SuggestionMethod.Valley)):
"Launch a mock training to find a good learning rate and return suggestions based on `suggest_funcs` as a named tuple"
n_epoch = num_it//len(self.dls.train) + 1
cb=LRFinder(start_lr=start_lr, end_lr=end_lr, num_it=num_it, stop_div=stop_div)
with self.no_logging(): self.fit(n_epoch, cbs=cb)
if suggest_funcs is not None:
lrs, losses = tensor(self.recorder.lrs[num_it//10:-5]), tensor(self.recorder.losses[num_it//10:-5])
nan_idxs = torch.nonzero(torch.isnan(losses.view(-1)))
if len(nan_idxs) > 0:
drop_idx = min(nan_idxs)
lrs = lrs[:drop_idx]
losses = losses[:drop_idx]
_suggestions, nms = [], []
for func in tuplify(suggest_funcs):
nms.append(func.__name__ if not isinstance(func, partial) else func.func.__name__) # deal with partials
_suggestions.append(func(lrs, losses, num_it))
SuggestedLRs = collections.namedtuple('SuggestedLRs', nms)
lrs, pnts = [], []
for lr, pnt in _suggestions:
lrs.append(lr)
pnts.append(pnt)
if show_plot: self.recorder.plot_lr_find(suggestions=pnts, nms=nms)
return SuggestedLRs(*lrs)
elif show_plot: self.recorder.plot_lr_find()
# First introduced by <NAME> in [Cyclical Learning Rates for Training Neural Networks](https://arxiv.org/pdf/1506.01186.pdf), the LR Finder trains the model with exponentially growing learning rates from `start_lr` to `end_lr` for `num_it` and stops in case of divergence (unless `stop_div=False`) then plots the losses vs the learning rates with a log scale.
#
# A variety of learning rate suggestion algorithms can be passed into the function, by default we use the `valley` paradigm.
#slow
with tempfile.TemporaryDirectory() as d:
learn = synth_learner(path=Path(d))
weights_pre_lr_find = L(learn.model.parameters())
lr_min, lr_steep, lr_valley, lr_slide = learn.lr_find(suggest_funcs=(minimum, steep, valley, slide))
weights_post_lr_find = L(learn.model.parameters())
test_eq(weights_pre_lr_find, weights_post_lr_find)
print(f"Minimum/10:\t{lr_min:.2e}\nSteepest point:\t{lr_steep:.2e}\nLongest valley:\t{lr_valley:.2e}\nSlide interval:\t{lr_slide:.2e}")
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
|
nbs/14_callback.schedule.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Direct imaging
# * GPIES ([Nielsen+2019]())
# +
import chronos as cr
tabs = cr.get_vizier_tables("J/AJ/158/13", tab_index=0, row_limit=-1)
tabs
# + active=""
# !mkdir /home/jp/github/research/project/tql/tests/Nielsen2019
# -
oudir = "/home/jp/github/research/project/tql/tests/Nielsen2019/"
fp_out = oudir+'nielsen2019.txt'
tabs["Name"].to_csv(fp_out, index=False)
|
notebooks/Nielsen2019_direct_imaging.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Imports
from time import time
from os.path import join as join_path
import numpy as np
import pandas as pd
import multiprocessing
cores = multiprocessing.cpu_count()
from gensim.models import Word2Vec
from gensim.models.callbacks import CallbackAny2Vec
import logging # Setting up the loggings to monitor gensim
logging.basicConfig(format="%(levelname)s - %(asctime)s: %(message)s", datefmt= '%H:%M:%S', level=logging.INFO)
import nltk
nltk.download('punkt')
from utils import clean_text, EpochSaver
from tqdm.notebook import tqdm
from sklearn.metrics.pairwise import cosine_similarity
# -
# ## Load and prepare data
# Constants
cord_data_dir = 'data'
cord_data_path = join_path(cord_data_dir, 'cord-19-data.csv')
w2v_saved_models_dir = 'models-word2vec'
saved_models_prefix = 'model'
cord_data = pd.read_csv(cord_data_path)
cord_data_eng = cord_data[cord_data['language'] == 'en']
eng_texts = cord_data_eng['body_text'].values
cord_num_sentences = 0
for text in tqdm(eng_texts):
sentences = nltk.tokenize.sent_tokenize(text)
cord_num_sentences += len(sentences)
print(f'Total number of CORD-19 sentences: {cord_num_sentences}')
class CORDDataIteratorWord2Vec():
def __init__(self, texts: np.ndarray):
self.texts = texts
def __iter__(self):
for text in self.texts:
sentences = nltk.tokenize.sent_tokenize(text)
cleaned_sentences = [clean_text(sent) for sent in sentences]
for sentence in cleaned_sentences:
yield sentence
cord_sentences = CORDDataIteratorWord2Vec(eng_texts)
# ## Learn word embeddings using Word2vec
class EpochSaver(CallbackAny2Vec):
'''Callback to save model after each epoch.'''
def __init__(self, output_dir: str, prefix: str, start_epoch: int = 1):
self.output_dir = output_dir
self.prefix = prefix
self.epoch = start_epoch
def on_epoch_end(self, model):
output_path = join_path(self.output_dir, f'{self.prefix}_epoch_{self.epoch}.model')
model.save(output_path)
self.epoch += 1
# Setup initial model
w2v_model = Word2Vec(
min_count=20,
window=2,
size=300,
negative=5,
workers=cores-1,
callbacks=[EpochSaver(w2v_saved_models_dir, saved_models_prefix)]
)
# Build vocabulary
t = time()
w2v_model.build_vocab(tqdm(cord_sentences, total=cord_num_sentences), progress_per=int(cord_num_sentences / 100))
print(f'Time to build vocab: {round((time() - t) / 60, 2)} mins')
# Train model
t = time()
w2v_model.train(
cord_sentences,
total_examples=w2v_model.corpus_count,
epochs=20,
report_delay=30
)
print(f'Time to train the model: {round((time() - t) / 60, 2)} mins')
|
code/learn_word_embeddings_word2vec.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# Dependencies
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# Load in csv
pyber_ride_df = pd.read_csv("Resources/PyBer_ride_data.csv")
pyber_ride_df
pyber_ride_df.plot(x="Month", y="Avg. Fare ($USD)")
plt.show()
# Set x-axis and tick locations.
x_axis = np.arange(len(pyber_ride_df))
tick_locations = [value for value in x_axis]
# Plot the data.
pyber_ride_df.plot(x="Month", y="Avg. Fare ($USD)")
plt.xticks(tick_locations, pyber_ride_df["Month"])
plt.show()
pyber_ride_df.plot.bar(x="Month", y="Avg. Fare ($USD)")
plt.show()
pyber_ride_df.plot(x="Month", y="Avg. Fare ($USD)", kind='bar')
plt.errorbar(x="Month", y="Avg. Fare ($USD)", yerr=stdev,capsize=3)
plt.show()
import statistics
stdev = statistics.stdev(pyber_ride_df['Avg. Fare ($USD)'])
stdev
pyber_ride_df.plot(x="Month", y="Avg. Fare ($USD)", kind='bar')
# plt.errorbar(x="Month", y="Avg. Fare ($USD)", yerr=stdv,capsize=3)
plt.show()
x_axis = pyber_ride_df["Month"]
y_axis = pyber_ride_df["Avg. Fare ($USD)"]
plt.bar(x_axis, y_axis, yerr=stdev, capsize=3, color="skyblue")
plt.yticks(np.arange(0, 51, step=5))
plt.legend(["Avg. Fare ($USD)"])
plt.xlabel("Month")
plt.show()
|
PyBer_ride_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
unames = ['user_id', 'gender', 'age', 'occupation', 'zip']
users = pd.read_table('ml-1m/users.dat', sep='::', header=None, names=unames, engine='python')
rnames = ['user_id', 'movie_id', 'rating', 'timestamp']
ratings = pd.read_table('ml-1m/ratings.dat', sep='::', header=None, names=rnames,engine='python')
mnames = ['movie_id', 'title', 'genres']
movies = pd.read_table('ml-1m/movies.dat', sep='::', header=None,names=mnames, engine='python')
users[:5]
ratings[:5]
movies[:5]
data = pd.merge(pd.merge(ratings, users), movies)
data
mean_ratings = data.pivot_table('rating', index='title', columns='gender',aggfunc='mean')
mean_ratings[:5]
ratings_by_title = data.groupby('title').size()
ratings_by_title[:10]
active_titles = ratings_by_title.index[ratings_by_title >= 250]
active_titles
mean_ratings = mean_ratings.ix[active_titles]
mean_ratings[:10]
top_female_ratings = mean_ratings.sort_values(by='F', ascending=False)
top_female_ratings[:10]
mean_ratings['diff'] = mean_ratings['M'] - mean_ratings['F']
sorted_by_diff = mean_ratings.sort_values(by='diff')
sorted_by_diff[:15]
sorted_by_diff[::-1][:15]
# Standard deviation of rating grouped by title
rating_std_by_title = data.groupby('title')['rating'].std()
rating_std_by_title = rating_std_by_title.ix[active_titles]
rating_std_by_title.sort_values(ascending=False)[:10]
|
DA Filmes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/VindhyaHV/AppliedAI_Assignments/blob/main/Assignment_6_KNN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="3ZFoCP20ClQe"
# + colab={"base_uri": "https://localhost:8080/"} id="breSw6aYkgd8" outputId="fe97b948-f3d5-4106-caad-baeda6894e64"
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import numpy
from tqdm import tqdm
import numpy as np
from sklearn.metrics.pairwise import euclidean_distances
x,y = make_classification(n_samples=10000, n_features=2, n_informative=2, n_redundant= 0, n_clusters_per_class=1, random_state=60)
X_train, X_test, Y_train, Y_test = train_test_split(x,y,stratify=y,random_state=42)
# del X_train,X_test
X_train.ndim
type(X_train)
len(X_train)
len(Y_train)
print(type(X_train),type(X_train[0]))
print(np.shape(Y_train))
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="i1jM4vjwkgeF" outputId="1e2c708f-92c7-49dd-f64c-0d6988e823af"
# %matplotlib inline
import matplotlib.pyplot as plt
colors = {0:'red', 1:'blue'}
plt.scatter(X_test[:,0], X_test[:,1],c=Y_test)
plt.show()
# + [markdown] id="jJM-3AzTkgeQ"
# # Implementing Custom RandomSearchCV
# + [markdown] id="gXpMFxiUkgeS"
# <pre>
# def RandomSearchCV(x_train,y_train,classifier, param_range, folds):
# # x_train: its numpy array of shape, (n,d)
# # y_train: its numpy array of shape, (n,) or (n,1)
# # classifier: its typically KNeighborsClassifier()
# # param_range: its a tuple like (a,b) a < b
# # folds: an integer, represents number of folds we need to devide the data and test our model
#
#
# #1.generate 10 unique values(uniform random distribution) in the given range "param_range" and store them as "params"
# # ex: if param_range = (1, 50), we need to generate 10 random numbers in range 1 to 50
# #2.devide numbers ranging from 0 to len(X_train) into groups= folds
# # ex: folds=3, and len(x_train)=100, we can devide numbers from 0 to 100 into 3 groups
# group 1: 0-33, group 2:34-66, group 3: 67-100
# #3.for each hyperparameter that we generated in step 1:
# # and using the above groups we have created in step 2 you will do cross-validation as follows
#
# # first we will keep group 1+group 2 i.e. 0-66 as train data and group 3: 67-100 as test data, and find train and
# test accuracies
#
# # second we will keep group 1+group 3 i.e. 0-33, 67-100 as train data and group 2: 34-66 as test data, and find
# train and test accuracies
#
# # third we will keep group 2+group 3 i.e. 34-100 as train data and group 1: 0-33 as test data, and find train and
# test accuracies
# # based on the 'folds' value we will do the same procedure
#
# # find the mean of train accuracies of above 3 steps and store in a list "train_scores"
# # find the mean of test accuracies of above 3 steps and store in a list "test_scores"
# #4. return both "train_scores" and "test_scores"
#
# #5. call function RandomSearchCV(x_train,y_train,classifier, param_range, folds) and store the returned values into "train_score", and "cv_scores"
# #6. plot hyper-parameter vs accuracy plot as shown in reference notebook and choose the best hyperparameter
# #7. plot the decision boundaries for the model initialized with the best hyperparameter, as shown in the last cell of reference notebook
#
# </pre>
# + [markdown] id="7gP11D7g4Ofx"
# function to split x_train in to n chunks n = len(x_train)/folds
# + id="R8tPoLx5h8IQ"
def split_list_into_chunks(x_train,folds):
interval = int(len(x_train)/folds)
return [x_train[i:i+interval] for i in range(0,len(x_train),interval)]
#https://stackoverflow.com/questions/312443/how-do-you-split-a-list-into-evenly-sized-chunks
# + [markdown] id="asaKjzP-7o3Z"
#
# + [markdown] id="Z9orv3h0RtZ4"
# function to shuffle the x_train folds to split into train and test set , returns list of all the train and test sets seperatedly
#
# example for training (5000,2) , (5000,) for testing (2500,2),(2500,)
# + id="PZJ2yAQvV8EN"
def list_shuffler(input):
ls_out = []
for i in range(-len(input),0):
st = i
ls_in = []
for i in range(len(input)):
ls_in.append(input[st])
st+=1
ls_out.append(ls_in)
#print(len(ls_out[0][0][0][0]))
train_list = []
test_list = []
for i in range(len(ls_out)):
train = []
test = []
for j in range(len(ls_out[0])-1):
# print(ls_out[i][j])
train.extend(ls_out[i][j])
test.extend(ls_out[i][-1])
train_list.append(np.array(train))
#print(len(train_list),len(train_list[i]),len(train_list[i][j]))
test_list.append(np.array(test))
#print(len(test_list),len(test_list[i]),len(test_list[i][j]))
#print(type(train_list))
#print(type(train_list))
#print(len(train_list),len(train_list[i]),len(train_list[i][j]))
#print(len(test_list),len(test_list[i]),len(test_list[i][j]))
#print(type(train_list),type(train_list[0]),type(train_list[0][0]))
return train_list,test_list
#print(t[0][0])
#list_shuffler([[(1,1),(1,1),(1,1)],[(2,2),(2,2),(2,2)],[(3,3),(3,3),(3,3)]])
# + id="wEG0tqi_BBZt"
# it will take classifier and set of values for hyper parameter in dict type dict({hyper parameter: [list of values]})
# we are implementing this only for KNN, the hyper parameter should n_neighbors
def RandomSearchCV(x_train,y_train,classifier, param_range, folds):
trainscores = []
testscores = []
for k in tqdm(params['n_neighbors']):
trainscores_folds = []
testscores_folds = []
x_split = split_list_into_chunks(x_train,folds)
y_split = split_list_into_chunks(y_train,folds)
train_x,test_x = list_shuffler(x_split)
train_y,test_y = list_shuffler(y_split)
classifier.n_neighbors = k
for i in range(len(train_x)):
classifier.n_neighbors = k
classifier.fit(train_x[i],train_y[i])
y_predicted = classifier.predict(test_x[i])
testscores_folds.append(accuracy_score(test_y[i],y_predicted))
y_predicted = classifier.predict(train_x[i])
trainscores_folds.append(accuracy_score(train_y[i], y_predicted))
trainscores.append(np.mean(np.array(trainscores_folds)))
testscores.append(np.mean(np.array(testscores_folds)))
return trainscores,testscores
# check this out: https://stackoverflow.com/a/9755548/40840
# + id="Ok_eC_6uF-gg"
# + colab={"base_uri": "https://localhost:8080/", "height": 333} id="vxxj1iIyBEGd" outputId="ddbbddc8-7a70-4844-8b38-0cb7d76b2e96"
from sklearn.metrics import accuracy_score
from sklearn.neighbors import KNeighborsClassifier
import matplotlib.pyplot as plt
import random
import warnings
warnings.filterwarnings("ignore")
neigh = KNeighborsClassifier()
p1 , p2 = input('enter the range for parameters').split()
params = {}
params['n_neighbors'] = sorted(random.sample(range(int(p1),int(p2)),10))
print(params)
#params = {'n_neighbors':[3,5,7,9,11,13,15,17,19,21,23]}
folds = 3
trainscores,testscores = RandomSearchCV(X_train, Y_train, neigh, params, folds)
plt.plot(params['n_neighbors'],trainscores, label='train cruve')
plt.plot(params['n_neighbors'],testscores, label='test cruve')
plt.title('Hyper-parameter VS accuracy plot')
plt.legend()
plt.show()
# + id="rlePbF-eBHgd"
# understanding this code line by line is not that importent
def plot_decision_boundary(X1, X2, y, clf):
# Create color maps
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
x_min, x_max = X1.min() - 1, X1.max() + 1
y_min, y_max = X2.min() - 1, X2.max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.02), np.arange(y_min, y_max, 0.02))
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=cmap_light)
# Plot also the training points
plt.scatter(X1, X2, c=y, cmap=cmap_bold)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.title("2-Class classification (k = %i)" % (clf.n_neighbors))
plt.show()
# + id="F8_lHCHcBJ8b" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="0202356b-3149-43d5-f146-7857190e60bf"
from matplotlib.colors import ListedColormap
neigh = KNeighborsClassifier(n_neighbors = 38 )
neigh.fit(X_train, Y_train)
plot_decision_boundary(X_train[:, 0], X_train[:, 1], Y_train, neigh)
# + id="A_MywoDCj2KQ"
|
Assignment_6_KNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="P8oDbrY3lpSS"
# # Dados de Internações COVID-19
# + [markdown] id="RxEEJpXil5-n"
# # Bibliotecas
# + colab={"base_uri": "https://localhost:8080/", "height": 595} id="Hp4otIHnlzPr" outputId="9bd6f21c-09c7-4d28-fe6e-f607efb6ba45"
# Manipulação e tratamento das bases
import pandas as pd
import numpy as np
from numpy import mean
from numpy import std
pd.set_option('display.max_rows', 50)
pd.set_option('display.max_columns', 100)
pd.set_option('display.width', 1000)
from scipy import stats
# #!pip install -U scikit-learn
#Visualização gráfica
# %matplotlib inline
import seaborn as sns
from matplotlib import pyplot
import matplotlib.pyplot as plt
import plotly.express as px
import io
from sklearn import tree
#Pré-Processamento das bases
# !pip install imblearn
from imblearn.under_sampling import RandomUnderSampler
from imblearn.over_sampling import SMOTE
from imblearn.under_sampling import TomekLinks
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler, StandardScaler, RobustScaler, OneHotEncoder, OrdinalEncoder
from sklearn.compose import ColumnTransformer
# #!pip install Boruta
from boruta import BorutaPy
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import mutual_info_classif
from sklearn.feature_selection import SelectFromModel
#Modelagem de Dados
from sklearn.datasets import make_classification
from sklearn.ensemble import GradientBoostingClassifier, BaggingClassifier, AdaBoostClassifier, VotingClassifier, StackingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier, ExtraTreesClassifier
from sklearn.naive_bayes import GaussianNB, BernoulliNB
from sklearn.svm import LinearSVC
from sklearn.linear_model import RidgeClassifier, LogisticRegression, SGDClassifier, PassiveAggressiveClassifier, Perceptron
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis, QuadraticDiscriminantAnalysis
from sklearn.neighbors import KNeighborsClassifier, NearestCentroid
from sklearn.neural_network import MLPClassifier, BernoulliRBM
from sklearn.calibration import CalibratedClassifierCV
from sklearn.dummy import DummyClassifier
from sklearn.semi_supervised import LabelSpreading, LabelPropagation
# ! pip install XGboost
from xgboost import XGBClassifier
#from lightgbm import LGBMClassifier
# ! pip install catboost
from catboost import CatBoostClassifier
from sklearn.tree import DecisionTreeClassifier, plot_tree, ExtraTreeClassifier
from sklearn.model_selection import cross_val_score, KFold
from sklearn.model_selection import RepeatedStratifiedKFold
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import RandomizedSearchCV
from sklearn.metrics import plot_confusion_matrix
from sklearn.metrics import classification_report
from sklearn.metrics import f1_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import roc_curve
from yellowbrick.classifier import ROCAUC
from yellowbrick.classifier import ConfusionMatrix
# + [markdown] id="HSNCFr_oo8Qk"
# # 1- Leitura de Dados e Pré- Processamento
#
#
# + [markdown] id="IwJz1VYXLUUm"
# ## **A-** Premissas:
#
#
# + [markdown] id="yaChLS_lN2RL" endofcell="--"
# Data Frame de Brasil:(**B-** df_Brasil)
# - Convertendo as variáveis com datas para datetime
# - Filtro: Criação da coluna IDADE
# - DATA LIMITE: 16/01/2021(Primeiro dia de vacinação no Brasil/sp)
# - Remoção de duplicadas
# - Remoção de variáveis não relevantes à análise (combinadas em reunião) e das que não agregam valor nenhum à Análise (nunique = 1)
# - Filtro: HOSPITAL = 1
#
# Data Frame da cidade de São Paulo:(**C-**df_sp)
# - Reset no INDEX
# - Remoção de colunas sobre o cód da cidade, estado, UF.. ou seja, com unique() = 1
# - Redução de categorias nas colunas 'UTI' de 1,2,3,9 e missing foram para 1 ou 0.
# -
# --
# + [markdown] id="lnFesWO93MXZ"
# ## **B**- Análises dos Internados Brasil
# + [markdown] id="2pte25iPeR90"
# Inclusão da Base "antes" do período da Vacina - Primeira parte do estudo
# + colab={"base_uri": "https://localhost:8080/"} id="dM-Uyl2txDmo" outputId="afe6c46b-f4f6-4b87-918a-55715c4ec1de"
df_brasil_antes= pd.read_csv('INFLUD-13-09-2021.csv', sep= ';')
# + [markdown] id="iKbqDVSsdn20"
# Filtro: Criação da coluna IDADE e drop nas outras .
# OBS: No campo de 'NU_IDADE_N','TP_IDADE','COD_IDADE',devemos dar atenção maior se as idades forem menores que 1 ano:
#
# Ex: Paciente apresenta, na coluna da idade, o valor 8, porém na coluna seguinte (que classifica em 1:dias, 2:meses, 3: anos) apresenta o valor 2 para este paciente. Então não são 8 anos e sim, 8 meses.( 0,67 anos)
#
# Vamos então criar uma coluna com idade em float e não int.
# + id="gnzjHh2apCju"
idade_aux = []
for i,j in enumerate (df_brasil_antes.TP_IDADE):
if j == 2:
idade_aux.append(df_brasil_antes.NU_IDADE_N[i]/12)
elif j == 1:
idade_aux.append(df_brasil_antes.NU_IDADE_N[i]/360)
else:
idade_aux.append(df_brasil_antes.NU_IDADE_N[i])
df_brasil_antes["IDADE_ANOS"] = idade_aux
# + [markdown] id="L7olKLKad1NK"
# Seleção das 20 maiores cidades para o período "antes da vacina"
#
# Aplicação do filtro para seleção somente dos pacientes internados
#
# Seleção somente dos pacientes que foram curados ou vieram a óbito por Covid
# + colab={"base_uri": "https://localhost:8080/"} id="3hGw773sWRzo" outputId="8228e898-acce-43e7-ec52-47acbf8491af"
df_mcid1= df_brasil_antes.loc[df_brasil_antes['CO_MUN_NOT'].isin([355030, 330455, 530010, 292740, 230440, 310620, 130260, 410690,261160, 431490, 520870, 150140, 351880, 350950, 211130, 330490,270430, 330170, 240810, 500270])]
df_mcid1=df_mcid1[(df_mcid1['HOSPITAL']==1.0)]
df_mcid1= df_mcid1.loc[(df_mcid1['EVOLUCAO']<=2.0)]
df_mcid1=df_mcid1.assign(Periodo=1)
df_mcid1.info()
# + [markdown] id="smdoXvLNORSO"
# - Convertendo as variáveis com datas para datetime
# + colab={"base_uri": "https://localhost:8080/"} id="3bRP_n3eI829" outputId="3ef863af-147e-4a0d-bb6d-24980a0a6bdd"
df_mcid1['DT_NOTIFIC'] = pd.to_datetime(df_mcid1['DT_NOTIFIC'], format= "%d/%m/%Y")
df_mcid1['DT_NASC'] = pd.to_datetime(df_mcid1['DT_NASC'], format= "%d/%m/%Y")
df_mcid1['DT_EVOLUCA'] = pd.to_datetime(df_mcid1['DT_EVOLUCA'], format= "%d/%m/%Y")
df_mcid1['DT_ENCERRA'] = pd.to_datetime(df_mcid1['DT_ENCERRA'], format= "%d/%m/%Y")
df_mcid1['DT_ENTUTI'] = pd.to_datetime(df_mcid1['DT_ENCERRA'], format= "%d/%m/%Y")
df_mcid1['DT_SAIDUTI'] = pd.to_datetime(df_mcid1['DT_ENCERRA'], format= "%d/%m/%Y")
df_mcid1[['DT_NOTIFIC','DT_NASC', 'DT_EVOLUCA','DT_ENCERRA','DT_ENTUTI', 'DT_SAIDUTI']].info()
# + colab={"base_uri": "https://localhost:8080/"} id="qMneMrpuDZwR" outputId="251ffad7-2e88-4161-ee64-f79785c4c165"
df_mcid1['DT_NOTIFIC'].max()
# + colab={"base_uri": "https://localhost:8080/"} id="HDIRlT9YDciO" outputId="7757fb1d-7a4e-43fa-b985-789129401006"
df_mcid1['DT_NOTIFIC'].min()
# + [markdown] id="E2TolkiPeFqW"
# Inclusão da Base "durante" o período da Vacina - Segunda parte do estudo
# + colab={"base_uri": "https://localhost:8080/"} id="k3KzM2v2eTql" outputId="1d1bc3bb-a7a9-414d-a171-0b4eb3103690" tags=[]
df_brasil_durante= pd.read_csv('INFLUD21-13-09-2021.csv', sep= ';')
# + [markdown] id="tptFdsWgmcrw"
# - Filtro: Criação da coluna IDADE e drop nas outras .
#
# OBS: No campo de 'NU_IDADE_N','TP_IDADE','COD_IDADE',devemos dar atenção maior se as idades forem menores que 1 ano:
#
# Ex: Paciente apresenta, na coluna da idade, o valor 8, porém na coluna seguinte (que classifica em 1:dias, 2:meses, 3: anos) apresenta o valor 2 para este paciente. Então não são 8 anos e sim, 8 meses.( 0,67 anos)
#
# Vamos então criar uma coluna com idade em float e não int.
#
#
# + id="VlKHW6DwVRBH"
idade_aux = []
for i,j in enumerate (df_brasil_durante.TP_IDADE):
if j == 2:
idade_aux.append(df_brasil_durante.NU_IDADE_N[i]/12)
elif j == 1:
idade_aux.append(df_brasil_durante.NU_IDADE_N[i]/360)
else:
idade_aux.append(df_brasil_durante.NU_IDADE_N[i])
df_brasil_durante["IDADE_ANOS"] = idade_aux
# + [markdown] id="wNroq_Rl8rvb"
# Seleção das 20 maiores cidades para o período "durante a vacina"
#
# Aplicação do filtro para seleção somente dos pacientes internados
#
# Seleção somente dos pacientes que foram curados ou vieram a óbito por Covid
# + colab={"base_uri": "https://localhost:8080/", "height": 813} id="VUMFctZLW5L2" outputId="421700d4-c8cf-4cf6-bd3e-40db95a680b0"
df_mcid2= df_brasil_durante.loc[df_brasil_durante['CO_MUN_NOT'].isin([355030, 330455, 530010, 292740, 230440, 310620, 130260, 410690,261160, 431490, 520870, 150140, 351880, 350950, 211130, 330490,270430, 330170, 240810, 500270])]
df_mcid2=df_mcid2[(df_mcid2['HOSPITAL']==1.0)]
df_mcid2= df_mcid2.loc[(df_mcid2['EVOLUCAO']<=2.0)]
df_mcid2=df_mcid2.assign(Periodo=2)
df_mcid2
# + [markdown] id="sXchL2gLeTqm"
# - Convertendo as variáveis com datas para datetime
# + colab={"base_uri": "https://localhost:8080/"} id="DVHbdQeTeTqm" outputId="797f9e3a-1742-4e3b-b6c1-a6639426d8c3"
df_mcid2['DT_NOTIFIC'] = pd.to_datetime(df_mcid2['DT_NOTIFIC'], format= "%d/%m/%Y")
df_mcid2['DT_NASC'] = pd.to_datetime(df_mcid2['DT_NASC'], format= "%d/%m/%Y")
df_mcid2['DT_EVOLUCA'] = pd.to_datetime(df_mcid2['DT_EVOLUCA'], format= "%d/%m/%Y")
df_mcid2['DT_ENCERRA'] = pd.to_datetime(df_mcid2['DT_ENCERRA'], format= "%d/%m/%Y")
df_mcid2['DT_ENTUTI'] = pd.to_datetime(df_mcid2['DT_ENCERRA'], format= "%d/%m/%Y")
df_mcid2['DT_SAIDUTI'] = pd.to_datetime(df_mcid2['DT_ENCERRA'], format= "%d/%m/%Y")
df_mcid2[['DT_NOTIFIC','DT_NASC', 'DT_EVOLUCA','DT_ENCERRA','DT_ENTUTI', 'DT_SAIDUTI']].info()
# + colab={"base_uri": "https://localhost:8080/"} id="vCJZYMgDeTqn" outputId="2d2a42fe-424d-4cf0-c2ce-43fb28fdc3d2"
df_mcid2['DT_NOTIFIC'].max()
# + colab={"base_uri": "https://localhost:8080/"} id="41_vKAnjeTqn" outputId="8c81bb6a-9749-4eec-bf82-0f6eb41073fb"
df_mcid2['DT_NOTIFIC'].min()
# + [markdown] id="ZK31Gchv2jCJ"
# - DATA LIMITE: 16/01/2021
# df_mcid1 = Base antes da Vacina
#
# df_mcid2 = Base durante a Vacina
# + colab={"base_uri": "https://localhost:8080/"} id="dZlttxoW2wf5" outputId="458620cb-d939-4309-a259-4848e4302ff0"
df_mcid1=df_mcid1.loc[df_mcid1['DT_NOTIFIC']<='2021-01-16']
df_mcid1.info()
# + colab={"base_uri": "https://localhost:8080/"} id="8gky4TBBeTqo" outputId="cf71fd14-a28d-466a-b5aa-8429ad61c2bb"
df_mcid2=df_mcid2.loc[df_mcid2['DT_NOTIFIC']>'2021-01-16']
df_mcid2.info()
# + colab={"base_uri": "https://localhost:8080/"} id="GUjWgotTeTqo" outputId="22dc4790-4232-419c-ad4c-fc0fe488bbcf"
df_mcid3=pd.concat([df_mcid1, df_mcid2])
df_mcid3.info()
# + [markdown] id="yQLQkBCstHZd"
# - Variáveis : Combinamos de manter as variáveis que tenham pelo menos um SIM [nesta tabela](https://docs.google.com/spreadsheets/d/17-jccz8DWl_fW7NpA0d6A3wQ3Ntc8qKX/edit#gid=1202247063) a partir da interpretação do Dicionário.
#
# OBS: Nesta base atualizada existem 8 variáveis a mais do que a antiga:
#
# 'ESTRANG', 'VACINA_COV', 'DOSE_1_COV', 'DOSE_2_COV', 'LAB_PR_COV', 'LOTE_1_COV', 'LOTE_2_COV', 'FNT_IN_COV'.
#
# Essas colunas também foram dropadas.
#
#
#
# + id="2Ng3X3x94V6Q"
df_mcid3.drop(columns=['DT_NOTIFIC','SG_UF_NOT','NU_IDADE_N','TP_IDADE','COD_IDADE','DT_NASC','SEM_NOT','DT_SIN_PRI','SEM_PRI','ID_REGIONA','CO_REGIONA', 'ID_UNIDADE', 'CO_UNI_NOT','ID_PAIS','CO_PAIS',
'SG_UF', 'ID_RG_RESI','CO_RG_RESI','ID_MN_RESI','CO_MUN_RES','CS_ZONA', 'HOSPITAL',
'CS_ETINIA', 'PAC_COCBO','PAC_DSCBO','PAIS_VGM','CO_PS_VGM','LO_PS_VGM','DT_VGM','DT_RT_VGM',
'OUTRO_DES','MORB_DESC','DT_UT_DOSE','MAE_VAC', 'DT_VAC_MAE', 'ID_MUNICIP',
'M_AMAMENTA','DT_DOSEUNI', 'DT_1_DOSE','DT_2_DOSE','TP_ANTIVIR','OUT_ANTIV','DT_ANTIVIR','DT_INTERNA',
'ID_RG_INTE','CO_RG_INTE', 'ID_MN_INTE','RAIOX_OUT','DT_RAIOX','TOMO_OUT','DT_TOMO', 'DT_ENTUTI', 'DT_SAIDUTI',
'AMOSTRA','DT_COLETA','TP_AMOSTRA','OUT_AMOST', 'DT_PCR','POS_PCRFLU','TP_FLU_PCR','PCR_FLUASU','FLUASU_OUT',
'PCR_FLUBLI','FLUBLI_OUT','POS_PCROUT','PCR_SARS2', 'PCR_VSR','PCR_PARA1','PCR_PARA2','PCR_PARA3','PCR_PARA4',
'PCR_ADENO','PCR_METAP','PCR_BOCA','PCR_RINO','PCR_OUTRO', 'DS_PCR_OUT','CLASSI_FIN','CLASSI_OUT','CRITERIO', 'AVE_SUINO','OUTRO_SIN','OUT_MORBI','CO_MU_INTE','RAIOX_RES','TOMO_RES',
'TP_TES_AN','DT_RES_AN','RES_AN','POS_AN_FLU','TP_FLU_AN','POS_AN_OUT', 'AN_SARS2','AN_VSR','AN_PARA1', 'SG_UF_INTE', 'CO_MU_INTE', 'DT_ENCERRA',
'AN_PARA2','AN_PARA3','AN_ADENO','AN_OUTRO','DS_AN_OUT','TP_AM_SOR','SOR_OUT','DT_CO_SOR', 'TP_SOR','OUT_SOR', 'RES_IGG', 'RES_IGM', 'RES_IGA', 'DT_EVOLUCA',
'DT_RES','DT_DIGITA','OBES_IMC', 'OUT_ANIM', 'ESTRANG', 'VACINA_COV', 'DOSE_1_COV', 'DOSE_2_COV', 'LAB_PR_COV', 'LOTE_1_COV', 'LOTE_2_COV', 'FNT_IN_COV'], inplace=True)
# -
df_mcid3.info()
# + [markdown] id="8-GAQYw0-0Cv"
# - Linhas duplicadas
#
# + colab={"base_uri": "https://localhost:8080/"} id="tuJfUnfd-N4H" outputId="3a6302e4-5dd2-4e88-cad5-704adea92807"
print(f'Temos {df_mcid3.duplicated().sum()} linhas duplicadas.')
# + id="A7K9B7S6XNIa"
df_mcid3=df_mcid3.drop_duplicates()
# + [markdown] id="0KPcAJ1C8HT3"
# Foi decidido pelo grupo dropar as 3407 linhas duplicadas. Que corresponde a menos de 1%
#
#
# + [markdown] id="CdO_T2aQXsX9"
# - Missing
# + id="QSjrw55UkEnB"
# Porcentagem de missing
#df_mcid3.isnull().sum()/len(df_mcid3)
# + [markdown] id="AX_0rHApaxtB"
# - Avaliação de Variáveis com unique()= 1
# + colab={"base_uri": "https://localhost:8080/"} id="wLBLIRFdoFrg" outputId="02e152a5-ccfa-4440-eab8-ac08b80d5900"
# Analise das colunas que provavelmente vamos dropar:
print(list(df_mcid3.nunique()))
#As que estão com 1 são constantes em todo df_sp.
# -
df_mcid3.to_csv(r'Base_Tratada.csv',index = False)
# + [markdown] id="ZWj9qJdu7WUx"
# ## **C-** PRÉ-PROCESSAMENTO
# + [markdown] id="BKOOToD5C2nf"
# ###- ADEQUAÇÃO DAS CATEGORIAS E SUBSTITUIÇÃO DOS MISSINGS
#
# + id="muIubcW0mbSu"
df_mcid3['CS_GESTANT'].replace({1.0: 1, 2.0: 1, 3.0 :1, 4.0 : 1}, inplace= True)
df_mcid3['CS_GESTANT'].replace({5.0: 0, 6.0:0, 9.0:0}, inplace= True)
df_mcid3['CS_RACA'].fillna(9,inplace= True)
df_mcid3['CS_ESCOL_N'].fillna(9,inplace= True)
df_mcid3['SURTO_SG'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['SURTO_SG'].fillna(0,inplace= True)
df_mcid3['NOSOCOMIAL'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['NOSOCOMIAL'].fillna(0,inplace= True)
df_mcid3['FEBRE'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['FEBRE'].fillna(0,inplace= True)
df_mcid3['TOSSE'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['TOSSE'].fillna(0,inplace= True)
df_mcid3['GARGANTA'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['GARGANTA'].fillna(0,inplace= True)
df_mcid3['DISPNEIA'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['DISPNEIA'].fillna(0,inplace= True)
df_mcid3['DESC_RESP'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['DESC_RESP'].fillna(0,inplace= True)
df_mcid3['SATURACAO'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['SATURACAO'].fillna(0,inplace= True)
df_mcid3['DIARREIA'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['DIARREIA'].fillna(0,inplace= True)
df_mcid3['VOMITO'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['VOMITO'].fillna(0,inplace= True)
df_mcid3['PUERPERA'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['PUERPERA'].fillna(0,inplace= True)
df_mcid3['CARDIOPATI'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['CARDIOPATI'].fillna(0,inplace= True)
df_mcid3['HEMATOLOGI'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['HEMATOLOGI'].fillna(0,inplace= True)
df_mcid3['SIND_DOWN'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['SIND_DOWN'].fillna(0,inplace= True)
df_mcid3['HEPATICA'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['HEPATICA'].fillna(0,inplace= True)
df_mcid3['ASMA'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['ASMA'].fillna(0,inplace= True)
df_mcid3['DIABETES'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['DIABETES'].fillna(0,inplace= True)
df_mcid3['NEUROLOGIC'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['NEUROLOGIC'].fillna(0,inplace= True)
df_mcid3['PNEUMOPATI'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['PNEUMOPATI'].fillna(0,inplace= True)
df_mcid3['IMUNODEPRE'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['IMUNODEPRE'].fillna(0,inplace= True)
df_mcid3['RENAL'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['RENAL'].fillna(0,inplace= True)
df_mcid3['OBESIDADE'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['OBESIDADE'].fillna(0,inplace= True)
df_mcid3['ASMA'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['ASMA'].fillna(0,inplace= True)
df_mcid3['ANTIVIRAL'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['ANTIVIRAL'].fillna(0,inplace= True)
df_mcid3['UTI'].replace({2.0: 0, 9.0: 0}, inplace= True)
df_mcid3['UTI'].fillna(0,inplace= True)
df_mcid3['SUPORT_VEN'].replace({3.0: 0, 9.0: 0}, inplace= True)
df_mcid3['SUPORT_VEN'].fillna(0,inplace= True)
df_mcid3['PCR_RESUL'].fillna(4,inplace= True)
df_mcid3['HISTO_VGM'].replace({0: 2}, inplace= True)
df_mcid3['DOR_ABD'].replace({9.0: 0, 2.0 :0}, inplace= True)
df_mcid3['DOR_ABD'].fillna(0,inplace= True)
df_mcid3['FADIGA'].replace({9.0: 0, 2.0 :0}, inplace= True)
df_mcid3['FADIGA'].fillna(0,inplace= True)
df_mcid3['PERD_OLFT'].replace({9.0: 0, 2.0 :0}, inplace= True)
df_mcid3['PERD_OLFT'].fillna(0,inplace= True)
df_mcid3['PERD_PALA'].replace({9.0: 0, 2.0 :0}, inplace= True)
df_mcid3['PERD_PALA'].fillna(0,inplace= True)
df_mcid3['VACINA'].fillna(0,inplace= True)
df_mcid3['FATOR_RISC'].replace({'S': 1, 'N':2, '1':1, '2':2}, inplace= True)
df_mcid3['FATOR_RISC'].fillna(0,inplace= True)
# + [markdown] id="Mxqje8B3CKr1"
# Conferência das categorias
# + [markdown] id="IqkLiKmCM5H3"
# - Resetando o Index novamente.
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="6O-EIaVrM8GU" outputId="575f3ea0-ab30-4bda-b423-82ec0439d017"
df_mcid3= df_mcid3.reset_index(drop=True)
df_mcid3.head()
# + id="XGHk-bWHWBNp"
#df_mcid3.isnull().sum()/len(df_mcid3)
# + [markdown] id="pi0o69qkcHYH"
# Aplicação da Dummy nas Features Categóricas
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="qAf4fQgkcFzx" outputId="cf5b3abd-a202-4576-e7a0-7a4bb6a6e6b3"
df_mcid3=pd.get_dummies(df_mcid3, columns=['CS_SEXO', 'CS_GESTANT', 'CS_RACA', 'CS_ESCOL_N',
'SURTO_SG', 'NOSOCOMIAL', 'FEBRE', 'TOSSE', 'GARGANTA', 'DISPNEIA',
'DESC_RESP', 'SATURACAO', 'DIARREIA', 'VOMITO', 'PUERPERA',
'FATOR_RISC', 'CARDIOPATI', 'HEMATOLOGI', 'SIND_DOWN', 'HEPATICA',
'ASMA', 'DIABETES', 'NEUROLOGIC', 'PNEUMOPATI', 'IMUNODEPRE', 'RENAL',
'OBESIDADE', 'VACINA', 'ANTIVIRAL', 'UTI', 'SUPORT_VEN', 'PCR_RESUL',
'HISTO_VGM', 'DOR_ABD', 'FADIGA', 'PERD_OLFT', 'PERD_PALA'], drop_first=True)
df_mcid3.head()
# + [markdown] id="l5D4Xr01t6Wf"
# CONFERÊNCIA DO BALANEAMENTO DA BASE
# -
df_mcid3.info()
# + colab={"base_uri": "https://localhost:8080/"} id="XGaMX56Ar0Nt" outputId="90f2dc2c-d90c-414c-9a55-c65c827641f2"
df_mcid3["EVOLUCAO"].value_counts(normalize=True)
# + [markdown] id="ZoLmafi2eTrS"
# Split da Base
# + id="5_8M7X-SeTrT"
X = df_mcid3.drop(columns=["CO_MUN_NOT", "Periodo", 'EVOLUCAO', 'CS_SEXO_I', 'CS_RACA_9.0', 'CS_ESCOL_N_9.0', 'VACINA_9.0',
'PCR_RESUL_9.0', 'HISTO_VGM_9'])
y = df_mcid3['EVOLUCAO']
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, test_size=0.3, random_state=42)
# + [markdown] id="D7FlOjVb1DPH"
# BALANCEAMENTO DA BASE
# + colab={"base_uri": "https://localhost:8080/"} id="xfP7clSmvS6n" outputId="210fe63d-a5b1-4979-d8f1-5160fabf9dad"
Xtrain.shape, Xtest.shape, ytrain.shape, ytest.shape
# + colab={"base_uri": "https://localhost:8080/"} id="JhPicRsA1H3R" outputId="e02bc667-90f9-4fbc-d8be-eee86025de72"
smote = SMOTE(sampling_strategy = 'minority', random_state = 42)
Xtrain_over, ytrain_over = smote.fit_resample(Xtrain,ytrain)
Xtest_over, ytest_over = smote.fit_resample(Xtest,ytest)
Xtrain_over.shape, ytrain_over.shape, Xtest_over.shape, ytest_over.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 163} id="ZSC48tLpE8Gj" outputId="ae6c13c7-b4aa-4337-d477-1043fa4d9ff7"
Xtest_over.head()
# + [markdown] id="bHsyLlESERFn"
# Seleção de Features via Feature Importance
# + colab={"base_uri": "https://localhost:8080/"} id="n2NVKuqFhs6B" outputId="0b3a5461-e05c-4266-8627-a3362a952596"
random_forest = RandomForestClassifier(random_state=42)
random_forest.fit(Xtrain_over, ytrain_over)
# + colab={"base_uri": "https://localhost:8080/"} id="9CayjL7ths3t" outputId="d98872b6-3125-4616-b2e2-c1abc771caf8"
previsoes = random_forest.predict(Xtest_over)
previsoes
# + colab={"base_uri": "https://localhost:8080/"} id="DuQEQqlWhszp" outputId="56f2f42e-d230-43ab-d319-0284b39cfff5"
accuracy_score(ytest_over, previsoes)
# -
Xtest_over.columns
# + colab={"base_uri": "https://localhost:8080/"} id="4NVAPzsXhsxl" outputId="0007d9dc-6869-4573-b282-bd055d51be2f"
x = random_forest.feature_importances_
np.set_printoptions(suppress=True)
print(x)
# -
# Seleção e teste de performance das 13 features
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="Lga7aKC122yC" outputId="e66a63a9-ff62-41ac-c36e-a1183a1993ad"
X = Xtrain_over[['IDADE_ANOS','CS_SEXO_M','CS_RACA_4.0','FEBRE_1.0','DISPNEIA_1.0','SATURACAO_1.0','UTI_1.0',
'SUPORT_VEN_1.0', 'SUPORT_VEN_2.0', 'PCR_RESUL_2.0','TOSSE_1.0','DESC_RESP_1.0', 'FATOR_RISC_2']]
y = ytrain_over
X_test = Xtest_over[['IDADE_ANOS','CS_SEXO_M','CS_RACA_4.0','FEBRE_1.0','DISPNEIA_1.0','SATURACAO_1.0','UTI_1.0',
'SUPORT_VEN_1.0', 'SUPORT_VEN_2.0', 'PCR_RESUL_2.0','TOSSE_1.0','DESC_RESP_1.0', 'FATOR_RISC_2']]
y_test = ytest_over
# -
random_forest = RandomForestClassifier(random_state=42)
random_forest.fit(X, y)
previsoes = random_forest.predict(X_test)
previsoes
accuracy_score(y_test, previsoes)
print(classification_report(y_test, previsoes))
# # Seleção da Base Pré-Processada com as Features Selecionadas
Base_Final = df_mcid3[['Periodo','CO_MUN_NOT','IDADE_ANOS','CS_SEXO_M','CS_RACA_4.0','FEBRE_1.0','DISPNEIA_1.0',
'SATURACAO_1.0','UTI_1.0','SUPORT_VEN_1.0','SUPORT_VEN_2.0','PCR_RESUL_2.0','TOSSE_1.0',
'DESC_RESP_1.0', 'FATOR_RISC_2', 'EVOLUCAO']]
# + id="gdcIvDaPTtUo"
Base_Final.to_csv(r'Base_Final.csv',index = False)
|
Notebooks gerais/Notebook_Pre_Processamento.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from tqdm import tqdm
from utils import Kernel
from dataset.dodd import load_dodd
from onlineKernels.klmsRFF import *
from onlineKernels.knlmsRFF import *
from onlineKernels.krlsRFF import *
from syncFL import fl_sync
from asyncFL import fl_async
from cent_learn import cent_learn
# Multiprocessing
import multiprocessing as mp
print("Number of processors: ", mp.cpu_count())
# +
# Parameters
num_data = 3000
kernel = Kernel(3.73)
step_size = 0.8
reg_coeff = 0.1
threshold = 0.5
# Federated learning: Synchronous updating rule
plt.figure(figsize=(7, 6))
K = 10 # K edge clients
c = np.ones(K).reshape(K,1)/K # edge weighting
D = 50 # Dictionary size
L = [20, 100, 150]
iteration = 10000
num_experiments = 10
step_size = 0.8
KLMS_RFF = klms_rff(step_size,D)
# KLMS_RFF = KLMS(u,d,kernel,step_size,threshold,alpha_0=np.array(0).reshape(1,1))
KLMS_RFF_cent = klms_rff(0.2,D)
KNLMS_RFF = knlms_rff(step_size,reg_coeff,D)
KRLS_RFF = krls_rff(D)
for l in L:
amse = np.zeros(iteration)
amse_cent = np.zeros(iteration)
for epoch in tqdm(range(num_experiments)):
# Process data and kernel
u,d,d_true = load_dodd(num_data)
sigma = 1/np.sqrt(2*kernel.sigma)
W = (1/sigma) * np.random.normal(size=(2,D))
b = np.random.uniform(0,2*np.pi,(D,1))
h = np.sqrt(2/D) * np.cos(W.T @ u.T + b)
FLsync = fl_sync(K,h,l)
# Federated learning
mse_cent = cent_learn(iteration,K,KLMS_RFF_cent,h,u,d)
amse_cent += mse_cent/num_experiments
mse = FLsync.train(iteration,KLMS_RFF,u,d)
amse += mse/num_experiments
plt.semilogy(range(len(amse)),amse,label="l=" + str(l))
plt.semilogy(range(len(amse_cent)),amse_cent,label="centralized")
plt.legend()
plt.ylim([1e-2,1e0])
plt.xlim([0,3000])
# + tags=[]
# Synchronous Federated learning using different kernels
num_data = 3000
kernel = Kernel(3.73)
step_size = 0.8
reg_coeff = 1e-6
plt.figure(figsize=(7, 6))
K = 10 # K edge clients
c = np.ones(K).reshape(K,1)/K # edge weighting
D = 100 # Dictionary size
L = [100]
iteration = 10000
num_experiments = 10
KLMS_RFF = klms_rff(step_size,D)
KNLMS_RFF = knlms_rff(step_size,reg_coeff,D)
KRLS_RFF = krls_rff(D,beta=0.01,l=1.0)
KLMS_RFF_cent = klms_rff(0.1,D)
KNLMS_RFF_cent = knlms_rff(0.1,1e-8,D)
KRLS_RFF_cent = krls_rff(D,beta=4,l=1.0)
for l in L:
amse_KLMS = np.zeros(iteration)
amse_KNLMS = np.zeros(iteration)
amse_KRLS = np.zeros(iteration)
amse_KLMS_cent = np.zeros(iteration)
amse_KNLMS_cent = np.zeros(iteration)
amse_KRLS_cent = np.zeros(iteration)
for epoch in tqdm(range(num_experiments)):
# Process data and kernel
u,d,d_true = load_dodd(num_data)
sigma = 1/np.sqrt(2*kernel.sigma)
W = (1/sigma) * np.random.normal(size=(2,D))
b = np.random.uniform(0,2*np.pi,(D,1))
h = np.sqrt(2/D) * np.cos(W.T @ u.T + b)
FLsync = fl_sync(K,h,l)
mse_KLMS_cent = cent_learn(iteration,K,KLMS_RFF_cent,h,u,d)
amse_KLMS_cent += mse_KLMS_cent/num_experiments
mse_KNLMS_cent = cent_learn(iteration,K,KNLMS_RFF_cent,h,u,d)
amse_KNLMS_cent += mse_KNLMS_cent/num_experiments
mse_KRLS_cent = cent_learn(iteration,K,KRLS_RFF_cent,h,u,d)
amse_KRLS_cent += mse_KRLS_cent/num_experiments
mse_KLMS = FLsync.train(iteration,KLMS_RFF,u,d)
amse_KLMS += mse_KLMS/num_experiments
mse_KNLMS = FLsync.train(iteration,KNLMS_RFF,u,d)
amse_KNLMS += mse_KNLMS/num_experiments
mse_KRLS = FLsync.train(iteration,KRLS_RFF,u,d)
amse_KRLS += mse_KRLS/num_experiments
plt.semilogy(range(len(amse_KLMS)),amse_KLMS,label="$KLMS_{" + str(l)+"}$")
plt.semilogy(range(len(amse_KNLMS)),amse_KNLMS,label="$KNLMS_{" + str(l)+"}$")
plt.semilogy(range(len(amse_KRLS)),amse_KRLS,label="$KRLS_{" + str(l)+"}$")
plt.semilogy(range(len(amse_KLMS_cent)),amse_KLMS_cent,label="$KLMS_{centralized}$")
plt.semilogy(range(len(amse_KNLMS_cent)),amse_KNLMS_cent,label="$KNLMS_{centralized}$")
plt.semilogy(range(len(amse_KRLS_cent)),amse_KRLS_cent,label="$KRLS_{centralized}$")
plt.legend()
plt.ylim([1e-2,1e0])
plt.xlim([0,3000])
plt.ylabel("MSE")
plt.xlabel("iteration")
# -
plt.semilogy(range(len(amse_KLMS)),amse_KLMS,label="$KLMS_{" + str(l)+"}$")
plt.semilogy(range(len(amse_KNLMS)),amse_KNLMS,label="$KNLMS_{" + str(l)+"}$")
plt.semilogy(range(len(amse_KRLS)),amse_KRLS,label="$KRLS_{" + str(l)+"}$")
plt.semilogy(range(len(amse_KLMS_cent)),amse_KLMS_cent,label="$KLMS_{centralized}$")
plt.semilogy(range(len(amse_KNLMS_cent)),amse_KNLMS_cent,label="$KNLMS_{centralized}$")
plt.semilogy(range(len(amse_KRLS_cent)),amse_KRLS_cent,label="$KRLS_{centralized}$")
plt.legend()
# plt.ylim([1e-2,2e-2])
plt.xlim([0,3000])
plt.ylabel("MSE")
# +
num_experiments = 25
D = 100
# Experiment
mse_KNLMS = np.zeros(num_data)
mse_KNLMS_RFF = np.zeros(num_data)
for i in tqdm(range(num_experiments)):
# print('experiment',i+1)
u,d,d_true = generate_data(num_data)
sigma = 1/np.sqrt(2*kernel.sigma)
W = (1/sigma) * np.random.normal(size=(2,D))
b = np.random.uniform(0,np.pi,(D,1))
h = np.sqrt(2/D) * np.cos(W.T @ u.T + b)
KLMS = KLMS_RFF(d,h,step_size,D,alpha_0 = np.zeros((D,1)))
err_KNLMS,_,_ = KNLMS(u,d,kernel,step_size,threshold,0.5)
mse_KNLMS += ((np.array(d_true) - np.array(d) + err_KNLMS)**2)/num_experiments
err_KNLMS_RFF,_ = KLMS.train()
mse_KNLMS_RFF += ((np.array(d_true) - np.array(d) + err_KNLMS_RFF)**2)/num_experiments
mse_KNLMS_smooth = np.convolve(mse_KNLMS,np.ones(20)/20,'valid')
mse_KNLMS_RFF_smooth = np.convolve(mse_KNLMS_RFF,np.ones(20)/20,'valid')
plt.figure(figsize=(10, 6))
plt.semilogy(range(len(mse_KNLMS_smooth)),mse_KNLMS_smooth, label='KLMS')
plt.semilogy(range(len(mse_KNLMS_RFF_smooth)),mse_KNLMS_RFF_smooth, label='KLMS')
plt.ylim([0.005,1e0])
plt.xlim([0,3000])
plt.legend()
plt.show()
# +
import numpy as np
D = 100
step_size = 0.2
reg_coeff = 0.03
threshold = 0.5
num_data = 3000
kernel = Kernel(3.73)
class krls_rff:
def __init__(self,D,beta=1.0,l=1.0):
self.D = D
self.beta = beta
self.l = l
self.P = np.eye(D)/l
def train(self,h,d,alpha_0,P=None):
D = self.D
beta = self.beta
l = self.l
lambd = 0
alpha = alpha_0
err = []
if (P.any()):
P = self.P
k = h[0,0]
# print(h.shape)
K_inv = np.matrix(1/(lambd + k))
alpha = alpha_0
for n in range(1,len(d)):
k = h[n,n]
# print(k)
h_n = h.T[n].reshape((D,1))
a = P @ h
# print(lambd + k - h.T @ a)
delta = (lambd + k - h.T @ a).item()
P_top = np.c_[delta*P + a @ a.T,-a]
P_bot = np.c_[-a.T, 1]
P = np.r_[P_top,P_bot]
P /= delta
err.append((d[n] - h_n.T @ alpha).item())
alpha = np.array(alpha - ((a * err[-1])/delta)).reshape(m,1)
alpha = np.r_[alpha,[[err[-1]/delta]]]
return err,alpha,P
# Experiment
mse = np.zeros(num_data)
num_experiments = 25
KRLS_RFF = krls_rff(D)
for ii in tqdm(range(num_experiments)):
u,d,d_true = load_dodd(num_data)
sigma = 1/np.sqrt(2*kernel.sigma)
W = (1/sigma) * np.random.normal(size=(2,D))
b = np.random.uniform(0,2*np.pi,D).reshape(D,1)
h = np.sqrt(2/D) * np.cos(W.T @ u.T + b)
err,_,_ = KRLS_RFF.train(h,d,np.zeros((D,1)),KRLS_RFF.P)
mse = mse + (np.square((d_true - d) + err))/num_experiments
mse_smooth = np.convolve(mse,np.ones(20)/20,'valid')
plt.figure(figsize=(10, 6))
plt.semilogy(range(len(mse_smooth)),mse_smooth)
plt.ylim([0.005,1e0])
plt.xlim([0,3000])
plt.show()
# -
# +
def KRLS_RFF2(u,d,kernel,l,beta,D):
err = np.array([])
W = np.random.normal(loc=0, scale=2.25, size=(2,D))
b = np.random.uniform(0,2*np.pi,D).reshape(D,1)
# Initalization
P = np.eye(D)/l
alpha = np.zeros((D,1))
for n in range(1, len(d)):
u_n = u[n].reshape(1,2)
d_n = d[n]
k = kernel(u_n,u_n)
h = np.sqrt(2/D) * np.cos(W.T @ u_n + b)
a = K_inv @ h
delta = (k - h.T @ a).item()
err = np.append(err,d_n - h.T @ alpha)
if delta > threshold:
dictionary = np.r_[dictionary, u_n]
K_inv_num = np.c_[delta*K_inv + a @ a.T,-a]
K_inv_den = np.c_[-a.T, 1]
K_inv = np.r_[K_inv_num,K_inv_den]
K_inv = K_inv/delta
P_num = np.c_[P,np.zeros((m,1))]
P_den = np.c_[np.zeros((m,1)).T, 1]
P = np.r_[P_num,P_den]
alpha = np.array(alpha - ((a * err[-1])/delta)).reshape(m,1)
alpha = np.r_[alpha,[[err[-1]/delta]]]
m = m + 1
q_t = (P @ a)/(1 + a.T @ P @ a)
P = P - ((P @ a @ a.T @ P)/(1 + a.T @ P @ a))
alpha = alpha + K_inv @ q_t * err[-1]
# print('number of SVs',len(dictionary))
return err
# Experiment
mse = np.zeros(num_data)
for i in tqdm(range(num_experiments)):
# print('experiment',i+1)
u,d,d_true = generate_data(num_data)
startTime = time.time()
err = KRLS_RFF2(u,d,kernel,0.1,1,20)
# mse = mse + (((d_true - d) + err)**2)/num_experiments
# mse_smooth = np.convolve(mse,np.ones(20)/20,'valid')
# +
d_true =[0.1 , 0.1]
d = [d_true[0], d_true[1]]
# Grab new data
new_d_true = lambda x: x.append((0.8 - 0.5 * np.exp(-(x[-1]**2)))*x[-1] - (0.3 + 0.9*np.exp(-(x[-1]**2)))*x[-2] + 0.1*np.sin(np.pi*x[-1]))
for i in range(2,num_data+2):
print(new_d_true(d_true))
d.append(d_true[-1] + np.random.normal(0.0,0.1))
# -
|
FederatedLearing/playground.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
data={
"marksA": np.random.randint(0,100,5),
"marksB":np.random.randint(0,100,5),
"marksC":np.random.randint(0,100,5),
}
data
df=pd.DataFrame(data)
print(df)
df.head()
df.head(n=3)
df.to_csv("pandasBasics.csv",index=False)
mydata=pd.read_csv("pandasBasics.csv")
mydata
# # loading MNIST dataset using python
import pandas as pd
df = pd.read_csv("mnist_train.csv")
df.shape
df.columns
df.head(n=5)
data=df.values
print(type(data))
import numpy as np
import matplotlib.pyplot as plt
X=data[:,1:]
Y=data[:,0]
print(X)
print(Y)
img=X[245].reshape(28,28)
print(img)
plt.imshow(img,cmap='gray')
# ## train and test
split=int(0.8*X.shape[0])
X_train , Y_train= X[:split,:],Y[:split]
X_test, Y_test =X[split:,:],Y[split:]
print(X_train.shape,Y_train.shape)
print(X_test.shape,Y_test.shape)
# # scikit split
from sklearn.model_selection import train_test_split
XT ,YT ,xt ,yt =train_test_split?
XT ,YT ,xt ,yt =train_test_split
XT ,xt ,YT,yt =train_test_split(X,Y,test_size=0.2)
print(XT.shape,YT.shape)
print(xt.shape,yt.shape)
|
MachineLearning/pandasBasics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Clean And Analyze Employee Exit Surveys
#
# In this project we are going to clean and analyze surveys from employees of the Department of Education, Training and Employment (DETE) and the Technical and Further Education (TAFE) institute in Queensland, Australia. Data sets provided for both of them in the project directory
#
# There is two main question that we are going to answer:
#
# 1- Are employees who only worked for the institutes for a short period of time resigning due to some kind of dissatisfaction? What about employees who have been there longer?
#
# 2- Are younger employees resigning due to some kind of dissatisfaction? What about older employees?
#
# They want us to answer these question for both of data sets combined.
#
# ## Opening and viewing data set
#
# We first start by importing essential libraries and opening the data set
#
# Then we are going to explore the data set by showing first 2 rows for each data sets and print information about them and also explore missing data counts in separate cells
#
# +
import pandas as pd
import numpy as np
dete_survey = pd.read_csv("dete_survey.csv")
tafe_survey = pd.read_csv("tafe_survey.csv")
# %matplotlib inline
# -
# For dete_survey data set:
#
print(dete_survey.head(2))
print(dete_survey.isnull().sum())
dete_survey.info()
# As we can see in blow the data set have 822 rows and 56 columns. Some columns have a lot of missing values like:
#
# `Business Unit`
# `Aboriginal`
# `Torres Strait`
# `South Sea`
# `Disability`
# `NESB`
#
# Also this data set contains 'Not Stated' values that indicate values are missing, but they aren't represented as Nan.
#
# Let's do the same for tafe_survey data set
#
print(tafe_survey.head(2))
tafe_survey.info()
print(tafe_survey.isnull().sum())
# As we can see in blow the data set have 702 rows and 72 columns. There are a lot columns with a very long string names. Also there is some columns with a lot of missing data.
#
# ## Data cleaning
# In this step we are going to apply some data cleaning techniques to our data sets
#
# ### "Not Stated" to Nan
#
# We are going to read the dete_survey data set again but this time we set the 'Not Stated' to NaN
#
#
dete_survey = pd.read_csv("dete_survey.csv", na_values = 'Not Stated')
dete_survey.head(2)
# ### Dropping non-necessary columns
#
# Let's drop non necessary columns for each data set. We are going to drop 28 to 49 columns from dete_survey data set and 17 to 66 columns for tafe_survey data set
#
dete_survey = dete_survey.drop(dete_survey.columns[28:49], axis = 1)
tafe_survey = tafe_survey.drop(tafe_survey.columns[17:66], axis = 1)
# ### Modifying columns name
#
# in this part we are going to make some changes to data sets columns name to make them statndard and same.
#
# We first start by making modification to dete_survey data set. We are going to:
#
# Make all capitalization lowercase
#
# Remove any trailing whitespace from the end of the strings
#
# Replace spaces with underscores ('_')
#
dete_survey.columns = dete_survey.columns.str.lower().str.replace("\s+", " ") .str.replace(" ", "_")
# Then we are going to make changes to tafe_survey columns and make them same as dete_survey columns like blow:
#
# `Record ID`: `id`
# `CESSATION YEAR`: `cease_date`
# `Reason for ceasing employment`: `separationtype`
# `Gender. What is your Gender?`: `gender`
# `CurrentAge. Current Age`: `age`
# `Employment Type. Employment Type`: `employment_status`
# `Classification. Classification`: `position`
# `LengthofServiceOverall. Overall Length of Service at Institute (in years)`: `institute_service`
#
# +
changes = {'Record ID': 'id',
'CESSATION YEAR': 'cease_date',
'Reason for ceasing employment': 'separationtype',
'Gender. What is your Gender?': 'gender',
'CurrentAge. Current Age': 'age',
'Employment Type. Employment Type': 'employment_status',
'Classification. Classification': 'position',
'LengthofServiceOverall. Overall Length of Service at Institute (in years)': 'institute_service',
'LengthofServiceCurrent. Length of Service at current workplace (in years)': 'role_service'
}
tafe_survey = tafe_survey.rename(changes, axis = 1)
# -
# Let's take a look at first rows for each data sets
dete_survey.head(2)
tafe_survey.head(2)
# ### Include only resignation types.
#
# If we remember goal of this project it is aimed to explore only for separation type of resignation. Then we are going to exclude other type of separation. To see the unique type of separations for each data set we showed them in the following two cells blow. As you can see for the dete_survey data set there are three type of resignation and we should include all three of them.
#
dete_survey["separationtype"].value_counts()
tafe_survey["separationtype"].value_counts()
# We are going to find the row that their separation type contain 'Resignation' then save them to new copies
dete_resignations = dete_survey[dete_survey["separationtype"].str.contains("Resignation", na = False)].copy()
tafe_resignations = tafe_survey[tafe_survey["separationtype"].str.contains("Resignation", na = False)].copy()
dete_resignations["separationtype"].value_counts()
# ### Cleaning date columns
#
# First let's take a look at unique values in `cease_date` `dete_resignations` columns in dete_resignations data set and `cease_date` column in tafe_registrations data set to explore any nonsense value
#
dete_resignations["cease_date"].value_counts()
# As it is clear all dates are logical in `cease_date` column of dete_resignations data set
tafe_resignations["cease_date"].value_counts()
# Also all dates are logical in `cease_date` column of tafe_resignations data set
dete_resignations["dete_start_date"].value_counts()
# Also in this column there are no nonsense value
#
# Now were are going to extract the years from `cease_date` column of dete_resignations data set and convert to float type
#
pattern = r"([1-2][0-9]{3})"
dete_resignations["cease_date"] = dete_resignations["cease_date"].str.extract(pattern)
dete_resignations["cease_date"] = dete_resignations["cease_date"].astype("float")
dete_resignations["cease_date"].value_counts()
boxplot1 = dete_resignations.boxplot(column = ['cease_date', 'dete_start_date'])
boxplot2 = tafe_resignations.boxplot(column = ['cease_date'])
# As you can see for better view of dates in these data sets we plotted the box plot for date columns as in above.
#
# for the `dete_resignations` data set most of the start dates are in between 200 and 2010 but there are some outliers which are below 1980. but for cease date most of them happened after 2010. Also for `tafe_resignations` data set cease dates happened mostly between 2011 and 2012
#
# ### New column of institute_service
#
# According to the project goal we need service time to be calculated. For `tafe_resignations` these time actually calculated and saved in the column `institute_service`. We need to add new column with this name in `dete_resignations` data set. We only need to subtract the `dete_start_date` from the `cease_date`.
#
#
dete_resignations["institute_service"] = dete_resignations['cease_date'] - dete_resignations['dete_start_date']
dete_resignations["institute_service"].value_counts().sort_index(ascending = False)
dete_resignations["institute_service"].value_counts()
dete_resignations["institute_service"].plot.hist(bins = 5)
# as you can see base on above results most of employees had between 0 and 10 years of service.
#
#
# ### Identify Dissatisfied Employees
#
# In this step we are going to identify dissatisfied employees. In each data set there are some columns that we can imply if the employee resigned because of dissatisfaction or not then we use them to create new column to indicate our imply. We first start with tafe data set.
# In this data set there are two columns that we can use them for this task"
# `Contributing Factors. Dissatisfaction`
# `Contributing Factors. Job Dissatisfaction`
#
# in the code blow we just showed the value counts for one of them
#
tafe_resignations['Contributing Factors. Dissatisfaction'].value_counts(dropna=False)
# We need to replace every '-' and NaN tyope False value and the rest of them to True value. For this we first define a faction for this task and then we will pass this function to applymap method to apply this function to all of the values in these two columns.
# Here is the update_vals function in the code blow
#
# +
def update_vals(val):
if pd.isnull(val):
return np.nan
elif val == "-":
return False
else:
return True
# -
# Then we pass the function to applymap method as in code blow
# +
factors_tafe_dissatisfaction = ['Contributing Factors. Dissatisfaction', 'Contributing Factors. Job Dissatisfaction']
tafe_resignations[factors_tafe_dissatisfaction] = tafe_resignations[factors_tafe_dissatisfaction].applymap(update_vals)
# -
# We can see the changes for `Contributing Factors. Dissatisfaction` column
tafe_resignations['Contributing Factors. Dissatisfaction'].value_counts(dropna=False)
# And the changes for `Contributing Factors. Job Dissatisfaction` column for tafe data set
tafe_resignations['Contributing Factors. Job Dissatisfaction'].value_counts(dropna=False)
# now we are going to use df.any() method to creat a new column `dissatisfied`. as in code blow
tafe_resignations['dissatisfied'] = tafe_resignations[factors_tafe_dissatisfaction].any(axis=1, skipna=False)
tafe_resignations_up = tafe_resignations.copy()
tafe_resignations_up['dissatisfied'].value_counts(dropna=False)
# As you can see there are 91 employee that resigned because of dissatisfaction in some way in tafe data set.
#
# Now we are going to identify dissatisfaction for dete data set in this data set we are going to use blow data sets to imply dissatisfaction and we are going to do same approaches as we did for tafe data set
#
# Columns used for dissatisfaction identification:
#
# `job_dissatisfaction`
# `dissatisfaction_with_the_department`
# `physical_work_environment`
# `lack_of_recognition`
# `lack_of_job_security`
# `work_location`
# `employment_conditions`
# `work_life_balance`
# `workload`
#
# But if we look at some we will know that in this data set the values are True or False so we don't need to change them
#
# For example:
#
dete_resignations['job_dissatisfaction'].value_counts(dropna=False)
# we only need to creat new column of `dissatisfied` by using any() method
# +
factors_dete_dissatisfaction =['job_dissatisfaction',
'dissatisfaction_with_the_department',
'physical_work_environment',
'lack_of_recognition',
'lack_of_job_security',
'work_location',
'employment_conditions',
'work_life_balance',
'workload'
]
dete_resignations['dissatisfied'] = dete_resignations[factors_dete_dissatisfaction].any(axis = 1, skipna = False)
dete_resignations_up = dete_resignations.copy()
dete_resignations_up['dissatisfied'].value_counts(dropna = False)
# -
# AS we can see in above there are 149 employee in dete data set that resigned because of dissatisfaction.
#
# ## Combining the data
#
# Now it is the time to combine data sets.
#
# we are going to add a column to each data frame named `institute` witch for dete_resignations_up is `DETE` and for tafe_resignations_up is `TAFE` to make both of them distingushible in combined datafarme
#
dete_resignations_up['institute'] = 'DETE'
tafe_resignations_up['institute'] = 'TAFE'
# Now in the code blow we are going to combine two data sets
combined = pd.concat([dete_resignations_up, tafe_resignations_up], ignore_index = True)
combined.head()
# ## Dropping missing values
#
# In this part we are going to deal with missing values. First we are going to see number of missing values in each column by the code blow
#
combined.notnull().sum()
# For this task we are going to drop columns with more than 500 missing values. Then we are going to filter columns with 151 or less non null values.
#
# in the code blow we are going to drop these columns
#
combined_update = combined.dropna(thresh = 151, axis = 1).copy()
# lets check the number of missing values in combined_update dataframe
combined_update.isnull().sum()
# As you can see there is no column with more than 500 missing values
#
# ## Cleaning the service column
#
# In this part we are going to first extract numbers from this column then categorize them and assign them to new column named `service_cat`.
#
# We are going to use the definition blow for categorizing
#
# New: Less than 3 years at a company
#
# Experienced: 3-6 years at a company
#
# Established: 7-10 years at a company
#
# Veteran: 11 or more years at a company
#
# First lets convert each value in `institute_service` column to string then extract numbers then convert to float as in code blow
#
# +
combined_update['institute_service_num'] = combined_update['institute_service'].astype('str').str.extract(r"(\d+)")
combined_update['institute_service_num'] = combined_update['institute_service_num'].astype('float')
combined_update['institute_service_num'].value_counts()
# -
# Now we are going to define a function to categorize numbers as we mentioned before then apply it to `institute_service_num` and assign it to new `service_cat` column
# +
def categorize(val):
if pd.isnull(val):
return np.nan
elif val < 3:
return 'New'
elif 3 <= val < 6:
return 'Experienced'
elif 6 <= val < 10:
return 'Established'
elif val >= 10:
return 'Veteran'
combined_update['service_cat'] = combined_update['institute_service_num'].apply(categorize)
combined_update['service_cat'].value_counts(dropna = False)
# -
# ## Analyzing the data
#
# In this part we are going to do our final analysis and show results
# But first we need to deal with some missing data in `dissatisfied` column as you can see there is 8 missing data
#
combined_update['dissatisfied'].value_counts(dropna = False)
# To deal with this 8 missing values we are going to replace these values with the value that occurs most frequently in this column True or False
combined_update['dissatisfied'] = combined_update['dissatisfied'].fillna(False)
combined_update['dissatisfied'].value_counts(dropna = False)
# Now it is the time to calculate the dissatisfied employees in each `service_cat` group
#
# We are going to use pivot_table() method
#
# True values are going to be considered 1 and false values considered 0 by calculating mean we could have the percentage.
#
pv_dissatisfied = combined_update.pivot_table(values = 'dissatisfied', index = 'service_cat')
pv_dissatisfied
# Let's plot the above table in a bar plot using matplotlib
# %matplotlib inline
pv_dissatisfied.plot(kind ='bar')
# ### First question
#
# lets remind the first question:
#
# Are employees who only worked for the institutes for a short period of time resigning due to some kind of dissatisfaction? What about employees who have been there longer?
#
# now we can answer this question
#
# As you can see employees who worked for long time resigning due to some kind of dissatisfaction more than employees who worked short period of time
# ### Second question
#
# The second question was:
#
# Are younger employees resigning due to some kind of dissatisfaction? What about older employees?
#
# *"This project is ongoing project and we will add the rest of it as soon as possible"*
#
combined_update['age'].value_counts(dropna=False)
# Lets categorize the ages for this data set. we are going to this as instruction in this [link](https://www.statcan.gc.ca/eng/concepts/definitions/age2) but just a little modification by our self
#
# blow 24 = Young
# 24-55 = Adult
# over 56 = old
#
# but we first find the numbers in each cell of this column then, if there are upper and lower boundaries for the age we calculate the average and if not we only use the single number to categorize. after that we categorize this column and put in the new column named `age_cat`
#
# +
def categorize_age(val):
pattern = r"(\d+)"
if pd.isnull(val):
return np.nan
else:
x_string = re.findall(pattern, val)
x = list(map(int, x_string))
if len(x) > 1:
age = (x[0] + x[1]) / 2
else:
age = x[0]
if age <= 24:
return "Young"
elif 24 < age <= 55:
return "Adult"
elif age > 55:
return "old"
combined_update['age_cat'] = combined_update['age'].apply(categorize_age)
combined_update['age_cat'].value_counts(dropna = False)
# -
pv_dissatisfied_age = combined_update.pivot_table(values = 'dissatisfied', index = 'age_cat')
pv_dissatisfied_age
pv_dissatisfied_age.plot.bar()
# ## conclusion
#
# - in this project we tried to answer some question about the employee exit for two data set
# - we first did some data cleaning task for both data sets then we combined two of them
# - we categorized the dissatisfied employees in terms of their experience and age
# - we conclude employees who worked for long time resigning due to some kind of dissatisfaction more than employees who worked short period of time
# - also we saw mostly old employees resigning due to some kind of dissatisfaction in compare to young employees
|
clean_analyze_emp_exit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Fine-tuning: 通过微调来迁移学习
#
#
# 在前面的章节里我们展示了如何训练神经网络来识别小图片里的问题。我们也介绍了ImageNet这个学术界默认的数据集,它有超过一百万的图片和一千类的物体。这个数据集很大的改变计算机视觉这个领域,展示了很多事情虽然在小的数据集上做不到,但在数GB的大数据上是可能的。事实上,我们目前还不知道有什么技术可以在类似的但小图片数据集上,例如一万张图片,训练出一个同样强大的模型。
#
# 所以这是一个问题。尽管深度卷积神经网络在ImageNet上有了很惊讶的结果,但大部分人不关心Imagenet这个数据集本身。他们关心他们自己的问题。例如通过图片里面的人脸识别身份,或者识别图片里面的10种不同的珊瑚。通常大部分在非BAT类似大机构里的人在解决计算机视觉问题的时候,能获得的只是相对来说中等规模的数据。几百张图片很正常,找到几千张图片也有可能,但很难同Imagenet一样获得上百万张图片。
#
# 于是我们会有一个很自然的问题,如何使用在百万张图片上训练出来的强大的模型来帮助提升在小数据集上的精度呢?这种在源数据上训练,然后将学到的知识应用到目标数据集上的技术通常被叫做**迁移学习**。幸运的是,我们有一些有效的技术来解决这个问题。
#
# 对于深度神经网络来首,最为流行的一个方法叫做微调(fine-tuning)。它的想法很简单但有效:
#
#
# * 在源数据 $S$ 上训练一个神经网络。
# * 砍掉它的头,将它的输出层改成适合目标数据 $S$ 的大小
# * 将输出层的权重初始化成随机值,但其它层保持跟原先训练好的权重一致
# * 然后开始在目标数据集开始训练
#
# 下图图示了这个算法:
#
# 
#
# ## 热狗识别
#
# 这一章我们将通过[ResNet](../chapter_convolutional-neural-networks/resnet-gluon.md)来演示如何进行微调。因为通常不会每次从0开始在ImageNet上训练模型,我们直接从Gluon的模型园下载已经训练好的。然后将其迁移到一个我们感兴趣的问题上:识别热狗。
#
# 
#
#
# 热狗识别是一个二分类问题。我们这里使用的热狗数据集是从网上抓取的,它有$1400$张正类和同样多的负类,负类主要是食品相关图片。我们将各类的$1000$张作为训练集合,其余的作为测试集合。
#
# ### 获取数据
#
# 我们首先从网上下载数据并解压到`../data/hotdog`。每个文件夹下会有对应的`png`文件。
# + attributes={"classes": [], "id": "", "n": "17"}
from mxnet import gluon
import zipfile
data_dir = '../data'
fname = gluon.utils.download(
'https://apache-mxnet.s3-accelerate.amazonaws.com/gluon/dataset/hotdog.zip',
path=data_dir, sha1_hash='fba480ffa8aa7e0febbb511d181409f899b9baa5')
with zipfile.ZipFile(fname, 'r') as f:
f.extractall(data_dir)
# -
# 我们使用[图片增强](../image-augmentation.md)里类似的方法来处理图片。
# + attributes={"classes": [], "id": "", "n": "18"}
from mxnet import nd
from mxnet import image
from mxnet import gluon
train_augs = [
image.HorizontalFlipAug(.5),
image.RandomCropAug((224,224))
]
test_augs = [
image.CenterCropAug((224,224))
]
def transform(data, label, augs):
data = data.astype('float32')
for aug in augs:
data = aug(data)
data = nd.transpose(data, (2,0,1))
return data, nd.array([label]).asscalar().astype('float32')
# -
# 读取文件夹下的图片,并且画出一些图片
# + attributes={"classes": [], "id": "", "n": "20"}
# %matplotlib inline
import sys
sys.path.append('..')
import utils
train_imgs = gluon.data.vision.ImageFolderDataset(
data_dir+'/hotdog/train',
transform=lambda X, y: transform(X, y, train_augs))
test_imgs = gluon.data.vision.ImageFolderDataset(
data_dir+'/hotdog/test',
transform=lambda X, y: transform(X, y, test_augs))
data = gluon.data.DataLoader(train_imgs, 32, shuffle=True)
for X, _ in data:
X = X.transpose((0,2,3,1)).clip(0,255)/255
utils.show_images(X, 4, 8)
break
# -
# ### 模型和训练
#
# 这里我们将使用Gluon提供的ResNet18来训练。我们先从模型园里获取改良过ResNet。使用`pretrained=True`将会自动下载并加载从ImageNet数据集上训练而来的权重。
# + attributes={"classes": [], "id": "", "n": "21"}
from mxnet.gluon.model_zoo import vision as models
pretrained_net = models.resnet18_v2(pretrained=True)
# -
# 通常预训练好的模型由两块构成,一是`features`,二是`classifier`。后者主要包括最后一层全连接层,前者包含从输入开始的大部分层。这样的划分的一个主要目的是为了更方便做微调。我们先看下`classifer`的内容:
# + attributes={"classes": [], "id": "", "n": "22"}
pretrained_net.classifier
# -
# 【注意】未来几天我们可能会将`classifier`重命名成`output`,并在里面只保留最后的Dense层。
#
# 我们可以看一下第一个卷积层的部分权重。
# + attributes={"classes": [], "id": "", "n": "23"}
pretrained_net.features[1].params.get('weight').data()[0][0]
# -
# 在微调里,我们一般新建一个网络,它的定义跟之前训练好的网络一样,除了最后的输出数等于当前数据的类别数。新网络的`features`被初始化前面训练好网络的权重,而`classfier`则是从头开始训练。
# + attributes={"classes": [], "id": "", "n": "24"}
from mxnet import init
finetune_net = models.resnet18_v2(classes=2)
finetune_net.features = pretrained_net.features
finetune_net.classifier.initialize(init.Xavier())
# -
# 我们先定义一个可以重复使用的训练函数。
# + attributes={"classes": [], "id": "", "n": "25"}
def train(net, ctx, batch_size=64, epochs=10, learning_rate=0.01, wd=0.001):
train_data = gluon.data.DataLoader(train_imgs, batch_size, shuffle=True)
test_data = gluon.data.DataLoader(test_imgs, batch_size)
# 确保net的初始化在ctx上
net.collect_params().reset_ctx(ctx)
net.hybridize()
loss = gluon.loss.SoftmaxCrossEntropyLoss()
# 训练
trainer = gluon.Trainer(net.collect_params(), 'sgd', {
'learning_rate': learning_rate, 'wd': wd})
utils.train(train_data, test_data, net, loss, trainer, ctx, epochs)
# -
# 现在我们可以训练了。
# + attributes={"classes": [], "id": "", "n": "10"}
ctx = utils.try_all_gpus()
train(finetune_net, ctx)
# -
# 对比起见我们尝试从随机初始值开始训练一个网络
# + attributes={"classes": [], "id": "", "n": "11"}
scratch_net = models.resnet18_v2(classes=2)
scratch_net.initialize(init=init.Xavier())
train(scratch_net, ctx)
# -
# 可以看到,微调版本收敛比从随机值开始的要快很多。
#
# ### 图片预测
# + attributes={"classes": [], "id": "", "n": "12"}
import matplotlib.pyplot as plt
def classify_hotdog(net, fname):
with open(fname, 'rb') as f:
img = image.imdecode(f.read())
data, _ = transform(img, -1, test_augs)
plt.imshow(data.transpose((1,2,0)).asnumpy()/255)
data = data.expand_dims(axis=0)
out = net(data.as_in_context(ctx[0]))
out = nd.SoftmaxActivation(out)
pred = int(nd.argmax(out, axis=1).asscalar())
prob = out[0][pred].asscalar()
label = train_imgs.synsets
return 'With prob=%f, %s'%(prob, label[pred])
# -
# 接下来我们用训练好的图片来预测几张图片:
# + attributes={"classes": [], "id": "", "n": "13"}
classify_hotdog(finetune_net, '../img/real_hotdog.jpg')
# + attributes={"classes": [], "id": "", "n": "14"}
classify_hotdog(finetune_net, '../img/leg_hotdog.jpg')
# + attributes={"classes": [], "id": "", "n": "15"}
classify_hotdog(finetune_net, '../img/dog_hotdog.jpg')
# -
# ## 结论
#
# 我们看到通过一个预先训练好的模型,我们可以在即使小的数据集上训练得到很好的分类器。这是因为这两个任务里面的数据表示有很多共通性。例如他们都需要如何识别纹理,形状,边等等。而这些通常被在靠近数据的层有效的处理。因此,如果你有一个相对较小的数据在手,而且当心它可能不够训练出很好的模型,你可以寻找跟你数据类似的大数据集来先训练你的模型,然后再在你手上的数据集上微调。
#
# ## 练习
#
# - 多跑几个`epochs`直到收敛(你可以也需要调调参数),看看`scratch_net`和`finetune_net`最后的精度是不是有区别
# - 这里`finetune_net`重用了`pretrained_net`除最后全连接外的所有权重,试试少重用些权重,有会有什么区别
# - 事实上`ImageNet`里也有`hotdog`这个类,它的index是713。例如它对应的weight可以这样拿到。试试如何重用这个权重
# + attributes={"classes": [], "id": "", "n": "16"}
weight = pretrained_net.classifier[4].params.get('weight')
hotdog_w = nd.split(weight.data(), 1000, axis=0)[713]
hotdog_w.shape
# -
# - 试试不让`finetune_net`里重用的权重参与训练,就是不更新权重
# - 如果图片预测这一章里我们训练的模型没有分对所有的图片,如何改进?
#
#
# **吐槽和讨论欢迎点**[这里](https://discuss.gluon.ai/t/topic/2272)
|
chapter_computer-vision/fine-tuning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Rover Project Test Notebook
# This notebook contains the functions from the lesson and provides the scaffolding you need to test out your mapping methods. The steps you need to complete in this notebook for the project are the following:
#
# * First just run each of the cells in the notebook, examine the code and the results of each.
# * Run the simulator in "Training Mode" and record some data. Note: the simulator may crash if you try to record a large (longer than a few minutes) dataset, but you don't need a ton of data, just some example images to work with.
# * Change the data directory path (2 cells below) to be the directory where you saved data
# * Test out the functions provided on your data
# * Write new functions (or modify existing ones) to report and map out detections of obstacles and rock samples (yellow rocks)
# * Populate the `process_image()` function with the appropriate steps/functions to go from a raw image to a worldmap.
# * Run the cell that calls `process_image()` using `moviepy` functions to create video output
# * Once you have mapping working, move on to modifying `perception.py` and `decision.py` to allow your rover to navigate and map in autonomous mode!
#
# **Note: If, at any point, you encounter frozen display windows or other confounding issues, you can always start again with a clean slate by going to the "Kernel" menu above and selecting "Restart & Clear Output".**
#
# **Run the next cell to get code highlighting in the markdown cells.**
# %%HTML
<style> code {background-color : orange !important;} </style>
# + outputExpanded=false
# %matplotlib inline
# #%matplotlib qt # Choose %matplotlib qt to plot to an interactive window (note it may show up behind your browser)
# Make some of the relevant imports
import cv2 # OpenCV for perspective transform
import numpy as np
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import scipy.misc # For saving images as needed
import glob # For reading in a list of images from a folder
import imageio
imageio.plugins.ffmpeg.download()
# -
# ## Quick Look at the Data
# There's some example data provided in the `test_dataset` folder. This basic dataset is enough to get you up and running but if you want to hone your methods more carefully you should record some data of your own to sample various scenarios in the simulator.
#
# Next, read in and display a random image from the `test_dataset` folder
# + outputExpanded=false
path = '../test_dataset/IMG/*'
img_list = glob.glob(path)
# Grab a random image and display it
idx = np.random.randint(0, len(img_list)-1)
image = mpimg.imread(img_list[idx])
plt.imshow(image)
# -
# ## Calibration Data
# Read in and display example grid and rock sample calibration images. You'll use the grid for perspective transform and the rock image for creating a new color selection that identifies these samples of interest.
# +
# In the simulator you can toggle on a grid on the ground for calibration
# You can also toggle on the rock samples with the 0 (zero) key.
# Here's an example of the grid and one of the rocks
example_grid = '../calibration_images/example_grid1.jpg'
example_rock = '../calibration_images/example_rock1.jpg'
grid_img = mpimg.imread(example_grid)
rock_img = mpimg.imread(example_rock)
fig = plt.figure(figsize=(12,3))
plt.subplot(121)
plt.imshow(grid_img)
plt.subplot(122)
plt.imshow(rock_img)
# -
# ## Perspective Transform
#
# Define the perspective transform function from the lesson and test it on an image.
# +
# Define a function to perform a perspective transform
# I've used the example grid image above to choose source points for the
# grid cell in front of the rover (each grid cell is 1 square meter in the sim)
# Define a function to perform a perspective transform
def perspect_transform(img, src, dst):
M = cv2.getPerspectiveTransform(src, dst)
warped = cv2.warpPerspective(img, M, (img.shape[1], img.shape[0]))# keep same size as input image
return warped
# Define calibration box in source (actual) and destination (desired) coordinates
# These source and destination points are defined to warp the image
# to a grid where each 10x10 pixel square represents 1 square meter
# The destination box will be 2*dst_size on each side
dst_size = 5
# Set a bottom offset to account for the fact that the bottom of the image
# is not the position of the rover but a bit in front of it
# this is just a rough guess, feel free to change it!
bottom_offset = 6
source = np.float32([[14, 140], [301 ,140],[200, 96], [118, 96]])
destination = np.float32([[image.shape[1]/2 - dst_size, image.shape[0] - bottom_offset],
[image.shape[1]/2 + dst_size, image.shape[0] - bottom_offset],
[image.shape[1]/2 + dst_size, image.shape[0] - 2*dst_size - bottom_offset],
[image.shape[1]/2 - dst_size, image.shape[0] - 2*dst_size - bottom_offset],
])
warped = perspect_transform(grid_img, source, destination)
plt.imshow(warped)
#scipy.misc.imsave('../output/warped_example.jpg', warped)
# -
def find_rocks():
# Create an array of zeros same xy size as img, but single channel
color_select = np.zeros_like(img[:,:,0])
# Require that each pixel be above all three threshold values in RGB
# above_thresh will now contain a boolean array with "True"
# where threshold was met
rock_pix = (img[:,:,0] > rgb_thresh[0]) \
& (img[:,:,1] > rgb_thresh[1]) \
& (img[:,:,2] > rgb_thresh[2])
# Index the array of zeros with the boolean array and set to 1
color_select[rock_pix] = 1
# Return the binary image
return color_select
# ## Color Thresholding
# Define the color thresholding function from the lesson and apply it to the warped image
#
# **TODO:** Ultimately, you want your map to not just include navigable terrain but also obstacles and the positions of the rock samples you're searching for. Modify this function or write a new function that returns the pixel locations of obstacles (areas below the threshold) and rock samples (yellow rocks in calibration images), such that you can map these areas into world coordinates as well.
# **Hints and Suggestion:**
# * For obstacles you can just invert your color selection that you used to detect ground pixels, i.e., if you've decided that everything above the threshold is navigable terrain, then everthing below the threshold must be an obstacle!
#
#
# * For rocks, think about imposing a lower and upper boundary in your color selection to be more specific about choosing colors. You can investigate the colors of the rocks (the RGB pixel values) in an interactive matplotlib window to get a feel for the appropriate threshold range (keep in mind you may want different ranges for each of R, G and B!). Feel free to get creative and even bring in functions from other libraries. Here's an example of [color selection](http://opencv-python-tutroals.readthedocs.io/en/latest/py_tutorials/py_imgproc/py_colorspaces/py_colorspaces.html) using OpenCV.
#
# * **Beware However:** if you start manipulating images with OpenCV, keep in mind that it defaults to `BGR` instead of `RGB` color space when reading/writing images, so things can get confusing.
# +
# Identify pixels above the threshold
# Threshold of RGB > 160 does a nice job of identifying ground pixels only
def color_thresh(img, rgb_thresh=(160, 160, 160)):
# Create an array of zeros same xy size as img, but single channel
color_select = np.zeros_like(img[:,:,0])
# Require that each pixel be above all three threshold values in RGB
# above_thresh will now contain a boolean array with "True"
# where threshold was met
above_thresh = (img[:,:,0] > rgb_thresh[0]) \
& (img[:,:,1] > rgb_thresh[1]) \
& (img[:,:,2] > rgb_thresh[2])
# Index the array of zeros with the boolean array and set to 1
color_select[above_thresh] = 1
# Return the binary image
return color_select
threshed = color_thresh(warped)
plt.imshow(threshed, cmap='gray')
#scipy.misc.imsave('../output/warped_threshed.jpg', threshed*255)
# -
def obstacle_thresh(img, rgb_thresh=(130,130,130)):
# Create an array of zeros same xy size as img, but single channel
color_select = np.zeros_like(img[:,:,0])
# Require that each pixel be above all three threshold values in RGB
# above_thresh will now contain a boolean array with "True"
# where threshold was met
below_thresh = (img[:,:,0] < rgb_thresh[0]) \
& (img[:,:,1] < rgb_thresh[1]) \
& (img[:,:,2] < rgb_thresh[2])
# Index the array of zeros with the boolean array and set to 1
color_select[below_thresh] = 1
# Return the binary image
return color_select
obstacles = obstacle_thresh(warped)
plt.imshow(obstacles, cmap='gray')
def rock_thresh(img):
# Convert RGB to HSV using openCV
hsv = cv2.cvtColor(img, cv2.COLOR_RGB2HSV, 3)
#Define range of yellow colors in HSV
lower_yellow = np.array([20,150, 100], dtype='uint8')
upper_yellow = np.array([50, 255, 255], dtype='uint8')
# Threshhold the HSV image to get only yellow colors
mask = cv2.inRange(hsv,lower_yellow, upper_yellow)
return mask
rock = rock_thresh(rock_img)
plt.imshow(rock, cmap='gray')
# ## Coordinate Transformations
# Define the functions used to do coordinate transforms and apply them to an image.
# + outputExpanded=false
# Define a function to convert from image coords to rover coords
def rover_coords(binary_img):
# Identify nonzero pixels
ypos, xpos = binary_img.nonzero()
# Calculate pixel positions with reference to the rover position being at the
# center bottom of the image.
x_pixel = -(ypos - binary_img.shape[0]).astype(np.float)
y_pixel = -(xpos - binary_img.shape[1]/2 ).astype(np.float)
return x_pixel, y_pixel
# Define a function to convert to radial coords in rover space
def to_polar_coords(x_pixel, y_pixel):
# Convert (x_pixel, y_pixel) to (distance, angle)
# in polar coordinates in rover space
# Calculate distance to each pixel
dist = np.sqrt(x_pixel**2 + y_pixel**2)
# Calculate angle away from vertical for each pixel
angles = np.arctan2(y_pixel, x_pixel)
return dist, angles
# Define a function to map rover space pixels to world space
def rotate_pix(xpix, ypix, yaw):
# Convert yaw to radians
yaw_rad = yaw * (np.pi / 180)
xpix_rotated = (xpix * np.cos(yaw_rad)) - (ypix * np.sin(yaw_rad))
ypix_rotated = (xpix * np.sin(yaw_rad)) + (ypix * np.cos(yaw_rad))
# Return the result
return xpix_rotated, ypix_rotated
def translate_pix(xpix_rot, ypix_rot, xpos, ypos, scale):
# Apply a scaling and a translation
xpix_translated = (xpix_rot / scale) + xpos
ypix_translated = (ypix_rot / scale) + ypos
# Return the result
return xpix_translated, ypix_translated
# Define a function to apply rotation and translation (and clipping)
# Once you define the two functions above this function should work
def pix_to_world(xpix, ypix, xpos, ypos, yaw, world_size, scale):
# Apply rotation
xpix_rot, ypix_rot = rotate_pix(xpix, ypix, yaw)
# Apply translation
xpix_tran, ypix_tran = translate_pix(xpix_rot, ypix_rot, xpos, ypos, scale)
# Perform rotation, translation and clipping all at once
x_pix_world = np.clip(np.int_(xpix_tran), 0, world_size - 1)
y_pix_world = np.clip(np.int_(ypix_tran), 0, world_size - 1)
# Return the result
return x_pix_world, y_pix_world
# Grab another random image
idx = np.random.randint(0, len(img_list)-1)
image = mpimg.imread(img_list[idx])
warped = perspect_transform(image, source, destination)
threshed = color_thresh(warped)
# Calculate pixel values in rover-centric coords and distance/angle to all pixels
xpix, ypix = rover_coords(threshed)
dist, angles = to_polar_coords(xpix, ypix)
mean_dir = np.mean(angles)
# Do some plotting
fig = plt.figure(figsize=(12,9))
plt.subplot(221)
plt.imshow(image)
plt.subplot(222)
plt.imshow(warped)
plt.subplot(223)
plt.imshow(threshed, cmap='gray')
plt.subplot(224)
plt.plot(xpix, ypix, '.')
plt.ylim(-160, 160)
plt.xlim(0, 160)
arrow_length = 100
x_arrow = arrow_length * np.cos(mean_dir)
y_arrow = arrow_length * np.sin(mean_dir)
plt.arrow(0, 0, x_arrow, y_arrow, color='red', zorder=2, head_width=10, width=2)
# -
# ## Read in saved data and ground truth map of the world
# The next cell is all setup to read your saved data into a `pandas` dataframe. Here you'll also read in a "ground truth" map of the world, where white pixels (pixel value = 1) represent navigable terrain.
#
# After that, we'll define a class to store telemetry data and pathnames to images. When you instantiate this class (`data = Databucket()`) you'll have a global variable called `data` that you can refer to for telemetry and map data within the `process_image()` function in the following cell.
#
# + inputHidden=false outputHidden=false
# Import pandas and read in csv file as a dataframe
import pandas as pd
# Change the path below to your data directory
# If you are in a locale (e.g., Europe) that uses ',' as the decimal separator
# change the '.' to ','
df = pd.read_csv('../test_dataset/robot_log.csv', delimiter=';', decimal='.')
csv_img_list = df["Path"].tolist() # Create list of image pathnames
# Read in ground truth map and create a 3-channel image with it
ground_truth = mpimg.imread('../calibration_images/map_bw.png')
ground_truth_3d = np.dstack((ground_truth*0, ground_truth*255, ground_truth*0)).astype(np.float)
# Creating a class to be the data container
# Will read in saved data from csv file and populate this object
# Worldmap is instantiated as 200 x 200 grids corresponding
# to a 200m x 200m space (same size as the ground truth map: 200 x 200 pixels)
# This encompasses the full range of output position values in x and y from the sim
class Databucket():
def __init__(self):
self.images = csv_img_list
self.xpos = df["X_Position"].values
self.ypos = df["Y_Position"].values
self.yaw = df["Yaw"].values
self.count = 0 # This will be a running index
self.worldmap = np.zeros((200, 200, 3)).astype(np.float)
self.ground_truth = ground_truth_3d # Ground truth worldmap
# Instantiate a Databucket().. this will be a global variable/object
# that you can refer to in the process_image() function below
data = Databucket()
# -
# ## Write a function to process stored images
#
# Modify the `process_image()` function below by adding in the perception step processes (functions defined above) to perform image analysis and mapping. The following cell is all set up to use this `process_image()` function in conjunction with the `moviepy` video processing package to create a video from the images you saved taking data in the simulator.
#
# In short, you will be passing individual images into `process_image()` and building up an image called `output_image` that will be stored as one frame of video. You can make a mosaic of the various steps of your analysis process and add text as you like (example provided below).
#
#
#
# To start with, you can simply run the next three cells to see what happens, but then go ahead and modify them such that the output video demonstrates your mapping process. Feel free to get creative!
# +
# Define a function to pass stored images to
# reading rover position and yaw angle from csv file
# This function will be used by moviepy to create an output video
def process_image(img):
# Example of how to use the Databucket() object defined above
# to print the current x, y and yaw values
# print(data.xpos[data.count], data.ypos[data.count], data.yaw[data.count])
# TODO:
# 1) Define source and destination points for perspective transform
dst_size = 5
bottom_offset = 6
source = np.float32([[14, 140], [301 ,140],[200, 96], [118, 96]])
destination = np.float32([[image.shape[1]/2 - dst_size, image.shape[0] - bottom_offset],
[image.shape[1]/2 + dst_size, image.shape[0] - bottom_offset],
[image.shape[1]/2 + dst_size, image.shape[0] - 2*dst_size - bottom_offset],
[image.shape[1]/2 - dst_size, image.shape[0] - 2*dst_size - bottom_offset],
])
# 2) Apply perspective transform
warped= perspect_transform(img, source, destination)
# 3) Apply color threshold to identify navigable terrain/obstacles/rock samples
navigable_terrain = color_thresh(warped, (160,160,160))
obstacles = obstacle_thresh (warped, (130,130,130))
rock_samples = rock_thresh(warped)
# 4) Convert thresholded image pixel values to rover-centric coords
navigable_xpix,navigable_ypix = rover_coords(navigable_terrain)
obstacles_xpix,obstacles_ypix = rover_coords(obstacles)
rocks_xpix,rocks_ypix = rover_coords(rock_samples)
# 5) Convert rover-centric pixel values to world coords
scale = dst_size * 2
try:
# this will fail for the last frame index
xpos, ypos = data.xpos[data.count], data.ypos[data.count]
yaw = data.yaw[data.count-1]
worldmap_size = data.worldmap.shape[0]
navigable_x_world, navigable_y_pix_world = pix_to_world(
navigable_xpix, navigable_ypix,
xpos, ypos, yaw, worldmap_size, scale)
obstacles_x_world, obstacles_y_pix_world = pix_to_world(
obstacles_xpix, obstacles_ypix,
xpos, ypos, yaw, worldmap_size, scale)
rock_x_world, rock_y_pix_world = pix_to_world(
rocks_xpix, rocks_ypix,
xpos, ypos, yaw, worldmap_size, scale)
except IndexError:
xpos, ypos = data.xpos[data.count-1], data.ypos[data.count-1]
yaw = data.yaw[data.count]
# 6) Update worldmap (to be displayed on right side of screen)
data.worldmap[obstacle_y_world, obstacle_x_world, 0] =255
data.worldmap[rock_y_world, rock_x_world, 1] =255
data.worldmap[navigable_y_world, navigable_x_world, 2] =255
# 7) Make a mosaic image, below is some example code
# First create a blank image (can be whatever shape you like)
output_image = np.zeros((img.shape[0] + data.worldmap.shape[0], img.shape[1]*2, 3))
# Next you can populate regions of the image with various output
# Here I'm putting the original image in the upper left hand corner
output_image[0:img.shape[0], 0:img.shape[1]] = img
# Let's create more images to add to the mosaic, first a warped image
warped = perspect_transform(img, source, destination)
# Add the warped image in the upper right hand corner
output_image[0:img.shape[0], img.shape[1]:] = warped
# Overlay worldmap with ground truth map
map_add = cv2.addWeighted(data.worldmap, 1, data.ground_truth, 0.5, 0)
# Flip map overlay so y-axis points upward and add to output_image
output_image[img.shape[0]:, 0:data.worldmap.shape[1]] = np.flipud(map_add)
# Then putting some text over the image
cv2.putText(output_image,"Populate this image with your analyses to make a video!", (20, 20),
cv2.FONT_HERSHEY_COMPLEX, 0.4, (255, 255, 255), 1)
if data.count < len(data.images) - 1:
data.count += 1 # Keep track of the index in the Databucket()
return output_image
# -
# ## Make a video from processed image data
# Use the [moviepy](https://zulko.github.io/moviepy/) library to process images and create a video.
#
# +
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from moviepy.editor import ImageSequenceClip
# Define pathname to save the output video
output = '../output/test_mapping.mp4'
data = Databucket() # Re-initialize data in case you're running this cell multiple times
clip = ImageSequenceClip(data.images, fps=60) # Note: output video will be sped up because
# recording rate in simulator is fps=25
new_clip = clip.fl_image(process_image) #NOTE: this function expects color images!!
# %time new_clip.write_videofile(output, audio=False)
# -
# ### This next cell should function as an inline video player
# If this fails to render the video, try running the following cell (alternative video rendering method). You can also simply have a look at the saved mp4 in your `/output` folder
# +
from IPython.display import HTML
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(output))
# -
# ### Below is an alternative way to create a video in case the above cell did not work.
import io
import base64
video = io.open(output, 'r+b').read()
encoded_video = base64.b64encode(video)
HTML(data='''<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded_video.decode('ascii')))
|
code/Rover_Project_Test_Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Copyright 2021 The Cirq Developers
# +
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# -
# ### Cirq-web 3D circuit visualization
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/quantumlib/Cirq/blob/master/cirq-web/circuit_example.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/colab_logo_1x.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/quantumlib/Cirq/blob/master/cirq-web/circuit-example.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/github_logo_1x.png" />View source on GitHub</a>
# </td>
# </table>
# !pip install --quiet cirq
# +
import cirq
qubits = [cirq.GridQubit(x, y) for x in range(10) for y in range(10)]
import cirq.testing
from cirq.protocols import circuit_diagram_info
circuit = cirq.testing.random_circuit(cirq.GridQubit.square(10), n_moments=5, op_density=.7)
# +
"""
This cell builds a 3D circuit diagram using a big list of operations provided TS Circuit class.
"""
import cirq_web
from typing import Optional
from cirq_web.circuits.symbols import SymbolInfo
class FunkyHadamard(cirq_web.circuits.symbols.SymbolResolver):
def resolve(self, operation: cirq.Operation) -> Optional[SymbolInfo]:
if isinstance(operation.gate, cirq.HPowGate):
return SymbolInfo(['Hello!'], ['yellow'])
else:
return None
resolvers = list(cirq_web.circuits.symbols.DEFAULT_SYMBOL_RESOLVERS) + [
FunkyHadamard()
]
from cirq_web import Circuit3D
c3d = Circuit3D(circuit, resolvers, 2.5)
display(c3d)
# -
|
cirq-web/circuit_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''myenv'': venv)'
# language: python
# name: python37364bitmyenvvenv357140d6b8384aec8a2ca97fa481d04d
# ---
# ライブラリのインポート
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# 先程は$f(x_1,x_2)=x_1^2 + x_2^2$の最小化を行いましたが、以下の関数(Rosenbrock function)を最小化してみましょう。
# $$
# f(x_1,x_2)=(1-x_1)^2 + 100(x_2-x_1^2)^2
# $$
# この関数は$(x_1,x_2)=(1, 1)$で最小値を取ります。
# 微分は
# $$
# \begin{align*}
# \frac{\partial f}{\partial x_1} &= -2(1-x_1) - 400(x_2-x_1^2)x_1 \\
# \frac{\partial f}{\partial x_2} &= 200(x_2-x_1^2) \\
# \end{align*}
# $$
# となります。これを最急降下法(GD)で最小化すると、
# +
# 目的関数の等高線プロット
x1, x2 = np.meshgrid(np.linspace(-0.8, 1.2, num=100), np.linspace(-0.1, 1.1, num=100))
y = np.square(1-x1) + 100 * np.square(x2-np.square(x1)) # Rosenbrock関数
plt.contour(x1, x2, np.log(y), linestyles='dashed', levels=10) # 見やすさのため、z軸をlog-scaleにしています
x1History = []
x2History = []
x1 = -0.5 # 初期値
x2 = 0.5 # 初期値
x1History.append(x1)
x2History.append(x2)
# 最適化
learning_rate = 0.005 # ステップ幅
num_steps = 100 # 繰り返し回数
for i in range(num_steps):
# 一次微分
grad_x1 = -2 * (1 - x1) - 400 * (x2 - x1 * x1) * x1
grad_x2 = 200 * (x2 - x1 * x1)
# Gradient descent
x1 = x1 - learning_rate * grad_x1
x2 = x2 - learning_rate * grad_x2
x1History.append(x1)
x2History.append(x2)
# 更新値履歴のプロット
plt.plot(x1History, x2History,
color='black',
marker='o', markersize=5, markerfacecolor='None', markeredgecolor='black') # プロット
plt.xlabel('x1')
plt.ylabel('x2')
plt.xlim([-0.8, 1.2])
plt.ylim([-0.1, 1.1])
plt.show()
# -
# 学習率やステップ回数を変化させてどのように学習が変わるかを見てみてください。最小値までたどり着くためにはどの程度のステップ数が必要でしょうか?
# 最急降下法は
#
# $$
# \mathbf{x}^{(k+1)} = \mathbf{x}^{(k)} - \epsilon \cdot \left. \frac{\partial f}{\partial \mathbf{x}} \right|_{\mathbf{x}=\mathbf{x}^{(k)}}
# $$
# のように値を更新するアルゴリズムでした。このアルゴリズムでは、その場その場の傾きに従って降下を行うため、ジグザクのパターンで無駄な動きをすることがあります。また、変数ごとに微分の大きさが大きく異なる時、うまく最適化ができません。
#
# この問題を解決するためにさまざまな最適化手法が提案されています。最適化手法について調べて、実装し、上の関数(Rosenbrock function)を最適化してください。
#
# [Kerasに実装されている最適化手法のドキュメント](https://keras.io/ja/optimizers/)に有名な最適化手法のリストと参考文献があります。
#
# 他の最適化手法の一例としては
# - Momentum
# - AdaGrad
# - RMSprop
# - Adam
#
# 等があります。この内特にAdamが深層学習でよく使われています。
|
Optimization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # plotting the p-mode power spectrum
# In this notebook, we will be plotting a power spectrum of the p-modes using mean values of both full-disk Doppler velocity and continuum intensity data from the Helioseismic and Magnetic Imager instrument on NASA's Solar Dynamics Observatory (SDO). </br>
#
# Some of this code is borrowed from <NAME>'s tutorial on making power spectra from Doppler velocity data and analyzing acoustic travel times through the Sun.
# First, we'll import some modules.
import drms
import json, urllib, numpy as np, matplotlib.pylab as plt
from datetime import datetime as dt_obj
from matplotlib.dates import *
from math import pi
import os
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
# ### Step 1: Plot the data
# Grab two days of Doppler Velocity data:
c = drms.Client()
# +
# keys_V = c.query('hmi.V_45s[2015.05.14_08:00_TAI/2d][? QUALITY=0 ?]', key='T_REC, OBS_VR, DATAMEA2')
# -
# Convert T_REC from a string to a datetime object:
def parse_tai_string(tstr,datetime=True):
year = int(tstr[:4])
month = int(tstr[5:7])
day = int(tstr[8:10])
hour = int(tstr[11:13])
minute = int(tstr[14:16])
second = int(tstr[17:19])
if datetime: return dt_obj(year, month, day, hour, minute, second)
else: return year,month,day,hour,minute,second
t_rec_V = np.array([parse_tai_string(keys_V.T_REC[i],datetime=True) for i in range(keys_V.T_REC.size)])
nrecs_V = keys_V.shape[0]
# +
# subtract the radial component of the spacecraft's orbital velocity
# from the Doppler Velocity data (both in units of m/s)
corrected_data_V = keys_V.DATAMEA2 - keys_V.OBS_VR
# +
fig, ax = plt.subplots()
# ascribe the data to the axes
ax.plot(t_rec_V, corrected_data_V)
# format the x-axis with international atomic time
locator = AutoDateLocator()
locator.intervald[HOURLY] = [24] # only show every day
formatter = DateFormatter('%d %b %Y')
ax.xaxis.set_major_locator(locator)
ax.xaxis.set_major_formatter(formatter)
# label the axes and the plot
ax.set_xlabel('time in TAI starting at '+str(t_rec_V[0]))
ax.set_ylabel('DATAMEA2 - OBS_VR (m/s)')
ax.set_title('Raw Doppler Velocity data')
# -
# Grab four days of continuum intensity data:
keys_I = c.query('hmi.Ic_45s[2015.05.13_08:00_TAI/4d][? QUALITY=0 ?]', key='T_REC, OBS_VR, DATAMEA2')
t_rec_I = np.array([parse_tai_string(keys_I.T_REC[i],datetime=True) for i in range(keys_I.T_REC.size)])
nrecs_I = keys_I.shape[0]
# remove the effects of orbital velocity from the continuum intensity data
chunk = []
for i in range(1920, nrecs_I-1921, 1920):
before_chunk = keys_I.DATAMEA2[i-1920:i]
after_chunk = keys_I.DATAMEA2[i+1920:i+3840]
avg_chunk = (before_chunk + after_chunk) / 2.0
chunk.append(keys_I.DATAMEA2[i:i+1920] - avg_chunk)
# Plot the continuum intensity data for the same date range as the Doppler velocity data:
t_start_index = np.where(t_rec_I == t_rec_V[0])[0][0]
t_end_index = np.where(t_rec_I == t_rec_V[-1])[0][0]
# +
fig, ax = plt.subplots()
# ascribe the data to the axes
ax.plot(t_rec_I[t_start_index : t_end_index+1], keys_I.DATAMEA2[t_start_index : t_end_index+1])
# format the x-axis with international atomic time
locator = AutoDateLocator()
locator.intervald[HOURLY] = [24] # only show every day
formatter = DateFormatter('%d %b %Y')
ax.xaxis.set_major_locator(locator)
ax.xaxis.set_major_formatter(formatter)
# label the axes and the plot
ax.set_xlabel('time in TAI starting at '+str(t_rec_V[0]))
ax.set_ylabel('DATAMEA2 (DN/s) [with orbital velocity removed]')
ax.set_title('Raw Continuum Intensity Data')
# -
# ### Step 2: Remove the low-frequency components
# Perform a high pass filter by removing all the low-frequency components. The data spans 2 days so the step in frequency in the Fourier transform is 1/2d. Therefore, removing all power to 10/2d, 5/day, will remove the 12 and 24 hour oscillations and leave oscillations in the 5-minute range.
# zeroMean(thisArray) - sets the mean of the array to zero and replaces NaNs with 0.0
def zeroMean(thisArray):
avg = np.nanmean(thisArray)
thisArray[np.isnan(thisArray)] = avg
np.nan_to_num(thisArray)
thisArray - avg
return thisArray
final_V = zeroMean(corrected_data_V)
final_I = zeroMean(keys_I.DATAMEA2[t_start_index : t_end_index+1])
days = 2
filtf = 5 * days
x_V = np.fft.rfft(final_V)
x_V[0:filtf] = 0
y_V = np.fft.irfft(x_V)
y_V[len(y_V)-3:] = 0
# +
# plot the Doppler Velocity data
fig, ax = plt.subplots()
# ascribe the data to the axes
ax.plot(t_rec_V, y_V)
# format the x-axis with international atomic time
locator = AutoDateLocator()
locator.intervald[HOURLY] = [24] # only show every day
formatter = DateFormatter('%d %b %Y')
ax.xaxis.set_major_locator(locator)
ax.xaxis.set_major_formatter(formatter)
# label the axes and the plot
ax.set_xlabel('time in TAI starting at '+str(t_rec_V[0]))
ax.set_ylabel('DATAMEA2 - OBS_VR (m/s)')
ax.set_title('Detrended Doppler Velocity data')
# -
x_I = np.fft.rfft(final_I)
x_I[0:filtf] = 0
y_I = np.fft.irfft(x_I)
y_I[len(y_I)-3:] = 0
# +
# plot the Continuum Intensity data
fig, ax = plt.subplots()
# ascribe the data to the axes
ax.plot(t_rec_I[t_start_index : t_end_index+1], y_I)
# format the x-axis with international atomic time
locator = AutoDateLocator()
locator.intervald[HOURLY] = [24] # only show every day
formatter = DateFormatter('%d %b %Y')
ax.xaxis.set_major_locator(locator)
ax.xaxis.set_major_formatter(formatter)
# label the axes and the plot
ax.set_xlabel('time in TAI starting at '+str(t_rec_V[0]))
ax.set_ylabel('DATAMEA2 (DN/s) [with orbital velocity removed]')
ax.set_title('Detrended Continuum Intensity Data')
# -
# ### Step 3: Compute the power spectrum of the filtered data
# +
# the function below returns the power spectrum of real array data
def powSpec(thisData):
data = np.array(thisData)
averaged_data = zeroMean(data)
fft = np.fft.rfft(averaged_data)
powspec = fft * np.conj(fft)
return np.real(powspec)
# +
# Now compute power spectrum of the filtered Doppler Velocity data
ps_V = powSpec(y_V)
# Now compute power spectrum of the filtered continuum intensity data
ps_I = powSpec(y_I)
# We can zoom into the p-mode part of the spectrum
lowf = 150 * days
highf = 400 * days
# We can get an array of frequencies for the power spectrum
ifreqs = np.fft.rfftfreq(len(y_V),45) # len(y_V) equals len(y_I)
# Convert from Hertz to microHz
mfr = ifreqs * 1e6
# +
# Plot the p-mode power spectrum from the Doppler Velocity data
# and the continuum intensity data
fig, ax = plt.subplots()
# ascribe the data to the axes
ax.plot(mfr, ps_I, label='Continuum Intensity')
ax.plot(mfr, ps_V, label='Doppler Velocity')
# add some limits
plt.xlim(1500,5500)
plt.ylim(0,150000)
# label the axes, plot, and legend
ax.set_xlabel('Freq (microHz)')
ax.set_ylabel('Power (arb units)')
ax.set_title('P mode power spectrum')
legend = plt.legend(loc='upper right', fontsize=12, framealpha=0.0,title='')
legend.get_frame().set_linewidth(0.0)
# -
period_seconds = (1 / ifreqs)
period_minutes = period_seconds / 60.
# +
# Plot the p-mode power spectrum from the Doppler Velocity data
# and the continuum intensity data
fig, ax = plt.subplots()
# ascribe the data to the axes
ax.plot(period_minutes, ps_I, label='Continuum Intensity')
ax.plot(period_minutes, ps_V, label='Doppler Velocity')
# add some limits
plt.xlim(0,10)
plt.ylim(0,150000)
# label the axes, plot, and legend
ax.set_xlabel('Period (minutes)')
ax.set_ylabel('Power (arb units)')
ax.set_title('P mode power spectrum')
legend = plt.legend(loc='upper right', fontsize=12, framealpha=0.0,title='')
legend.get_frame().set_linewidth(0.0)
# -
# ***
# # Reproducing the above power spectrum with Brett's detrended SDO continuum intensity photometry
t18, f18 = np.load('detrended_photometry2018.npy').T
t17, f17 = np.load('detrended_photometry2017.npy').T
t16, f16 = np.load('detrended_photometry2016.npy').T
t15, f15 = np.load('detrended_photometry2015.npy').T
# +
fig, ax = plt.subplots(1, 4, figsize=(14, 4), sharey=True)
ax[0].plot(t15, f15)
ax[1].plot(t16, f16)
ax[2].plot(t17, f17)
ax[3].plot(t18, f18)
for axis, year in zip(ax, [2015, 2016, 2017, 2018]):
axis.set_title(year)
axis.set_xlabel('Cadence')
ax[0].set_ylabel('Relative Flux')
plt.show()
# +
# We can get an array of frequencies for the power spectrum
ifreqs_15 = np.fft.rfftfreq(len(t15), 45)
ifreqs_16 = np.fft.rfftfreq(len(t16), 45)
ifreqs_17 = np.fft.rfftfreq(len(t17), 45)
ifreqs_18 = np.fft.rfftfreq(len(t18), 45)
power_15 = powSpec(f15)
power_16 = powSpec(f16)
power_17 = powSpec(f17)
power_18 = powSpec(f18)
period_seconds_15 = (1 / ifreqs_15)
period_seconds_16 = (1 / ifreqs_16)
period_seconds_17 = (1 / ifreqs_17)
period_seconds_18 = (1 / ifreqs_18)
fig, ax = plt.subplots(1, 4, figsize=(14, 4), sharey=True)
ax[0].plot(period_seconds_15 / 60, power_15)
ax[1].plot(period_seconds_16 / 60, power_16)
ax[2].plot(period_seconds_17 / 60, power_17)
ax[3].plot(period_seconds_18 / 60, power_18)
for axis, year in zip(ax, [2015, 2016, 2017, 2018]):
axis.set_xlim([0, 10])
axis.set_title(year)
axis.set_ylim([0, 4e-5])
axis.set_xlabel('Period [min]')
ax[0].set_ylabel('Power')
plt.show()
# -
|
examples/generate_powerspectrum.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pylab import rcParams
rcParams['figure.figsize'] = 15, 10
from ast import literal_eval
from sklearn.feature_extraction.text import CountVectorizer
data = pd.read_csv("../data/interim/trigrams3.csv")
data = data.drop(['Unnamed: 0'], axis=1)
data['TRIGRAMS'] = data['TRIGRAMS'].apply(literal_eval)
data.head()
# +
# In this histogram, the y-axis is capped at 400 to show finer detail.
# The first bar actually represents a freqeuency of about 2,000 records.
data['FINE'].hist(bins = 30)
plt.xlabel('Amount of Fine ($)')
plt.ylabel('Frequency')
plt.ylim([0, 400])
plt.title('Distribution of Healthcare Fines')
plt.rcParams.update({'font.size': 16})
plt.show()
# -
data['FINE'].describe()
# +
# Build a corpus of strings out of the trigram column in the main database
corpus = [' '.join(data.loc[row, 'TRIGRAMS']) for row in range(len(data))]
corpus[0][0:100], corpus[1][0:100], corpus[2][0:100]
# +
# Convert the corpus to a vectorized bag of words
vectorizer = CountVectorizer(min_df=0)
vectorizer.fit(corpus)
x = vectorizer.transform(corpus)
x = x.toarray()
y = vectorizer.get_feature_names()
# +
# Build a list of the most frequently appearing trigrams, not including trigrams that are
# merely a number, e.g. "20".
# Can set the generate value to 'False' to save time in constructing the frequency list, which is
# stored to disc.
generate = True
if generate:
def generate_row(current_row_num, y):
current_row = [(y[i], x[current_row_num][i]) for i in range(len(y))]
return current_row
freq = {}
for current_row_num in range(len(data)):
for word in generate_row(current_row_num, y):
if (word[0] in freq):
freq[word[0]] += word[1]
else:
freq[word[0]] = word[1]
freq_df = pd.DataFrame.from_dict(freq, orient='index')
freq_df.columns=['frequency']
freq_df.index.name='word'
freq_df.to_csv("../data/interim/wordfreqs.csv")
# -
freq_df
freq_df = pd.read_csv("../data/interim/wordfreqs.csv")
numbers = freq_df['word'].str.contains('^[0-9]*$')
top_words = freq_df[~numbers].sort_values('frequency', ascending=False).head(50)
top_words
bigrams = freq_df['word'].str.contains('_')
top_bigrams = freq_df[bigrams].sort_values('frequency', ascending=False).head(20)
top_bigrams
trigrams = [t for t in freq_df['word'] if t.count('_') == 2]
trigrams[0:5]
# +
#freq_df['word'].in(trigrams)
# -
trigrams = freq_df['word'].str.contains('^[^_]+_([^_]+)_[^_]+$')
top_trigrams = freq_df[trigrams].sort_values('frequency', ascending=False).head(20)
top_trigrams
# +
# Illustrate the most frequently appearing words
from pylab import rcParams
rcParams['figure.figsize'] = 25, 12
plt.bar(top_words['word'], top_words['frequency'])
plt.xticks(
rotation=90,
horizontalalignment='center',
fontweight='light',
fontsize='small'
)
plt.title('Most commonly used words in California Healthcare Incident Reports')
plt.ylabel('Frequency of word in entire corpus', fontsize='small')
fig1 = plt.gcf()
fig1.savefig('../reports/figures/wordfreqs.png')
plt.show()
# +
# Illustrate most common bigrams
plt.bar(top_bigrams['word'], top_bigrams['frequency'])
plt.xticks(
rotation=90,
horizontalalignment='center',
fontweight='light',
fontsize='small'
)
plt.title('Most commonly used bigrams in California Healthcare Incident Reports')
plt.ylabel('Frequency of bigram in entire corpus', fontsize='small')
fig1 = plt.gcf()
fig1.savefig('../reports/figures/bigramfreqs.png')
plt.show()
# +
# Illustrate most common trigrams
plt.bar(top_trigrams['word'], top_trigrams['frequency'])
plt.xticks(
rotation=90,
horizontalalignment='center',
fontweight='light',
fontsize='small'
)
plt.title('Most commonly used trigrams in California Healthcare Incident Reports')
plt.ylabel('Frequency of trigram in entire corpus', fontsize='small')
fig1 = plt.gcf()
fig1.savefig('../reports/figures/trigramfreqs.png')
plt.show()
# +
# Break it up by fine amount
nofine = data[data['FINE'] == 0]
smallfine = data[(data['FINE'] > 0) & (data['FINE'] < 2000)]
medfine = data[(data['FINE'] >= 2000) & (data['FINE'] < 10000)]
largefine = data[data['FINE'] > 10000]
nofine.head()
# -
len(nofine), len(smallfine), len(medfine), len(largefine)
# +
# This step is very time-consuming, so I have saved the results to disk. Only set 'generateLists' to True if you want
# to re-generate each lsit instead of importing them from disk.
generateLists = False
if generateLists:
no_fine_words = pd.Series()
for row in range(len(nofine)):
nextentry = pd.Series(nofine.iloc[row, 5]).value_counts()
no_fine_words = no_fine_words.add(nextentry, fill_value=0)
no_fine_words = no_fine_words[no_fine_words > 100] / len(nofine)
no_fine_words = no_fine_words.sort_values()
small_fine_words = pd.Series()
for row in range(len(smallfine)):
nextentry = pd.Series(smallfine.iloc[row, 5]).value_counts()
small_fine_words = small_fine_words.add(nextentry, fill_value=0)
small_fine_words = small_fine_words[small_fine_words > 100] / len(smallfine)
small_fine_words = small_fine_words.sort_values()
med_fine_words = pd.Series()
for row in range(len(medfine)):
nextentry = pd.Series(medfine.iloc[row, 5]).value_counts()
med_fine_words = med_fine_words.add(nextentry, fill_value=0)
med_fine_words = med_fine_words[med_fine_words > 100] / len(medfine)
med_fine_words = med_fine_words.sort_values()
large_fine_words = pd.Series()
for row in range(len(largefine)):
nextentry = pd.Series(largefine.iloc[row, 5]).value_counts()
large_fine_words = large_fine_words.add(nextentry, fill_value=0)
large_fine_words = large_fine_words[large_fine_words > 100] / len(largefine)
large_fine_words = large_fine_words.sort_values()
all_words = pd.Series()
for row in range(len(data)):
nextentry = pd.Series(data.iloc[row, 5]).value_counts()
all_words = all_words.add(nextentry, fill_value=0)
all_words = all_words[all_words > 100] / len(data)
all_words = all_words * -1
all_words = all_words.sort_values()
no_fine_words.to_csv("../data/interim/nofine.csv")
small_fine_words.to_csv("../data/interim/smallfine.csv")
med_fine_words.to_csv("../data/interim/medfine.csv")
large_fine_words.to_csv("../data/interim/largefine.csv")
all_words.to_csv("../data/interim/allfine.csv")
# +
# This function converts each saved spreadsheet into a series while dropping unnecessary information.
# The function is inelegant but seems needed now that pd.Series.from_csv has been deprecated.
def process_csv_series(filename):
path = "../data/interim/" + filename
temp = pd.read_csv(path, squeeze=True, header=None, index_col=False)
temp.index = temp.loc[:, 0]
temp.index.name = None
output = temp.drop(0, axis=1)[1]
return output
# +
# Import each list of characteristic words from disk
no_fine_words = process_csv_series("nofine.csv")
small_fine_words = process_csv_series("smallfine.csv")
med_fine_words = process_csv_series("medfine.csv")
large_fine_words = process_csv_series("largefine.csv")
all_words = process_csv_series("allfine.csv")
# -
no_fine_diff = no_fine_words.add(all_words, fill_value=0).sort_values(ascending=False).head(50)
no_fine_diff.head(10)
small_fine_diff = small_fine_words.add(all_words, fill_value=0).sort_values(ascending=False).head(50)
small_fine_diff.head(10)
med_fine_diff = med_fine_words.add(all_words, fill_value=0).sort_values(ascending=False).head(50)
med_fine_diff.head(10)
large_fine_diff = large_fine_words.add(all_words, fill_value=0).sort_values(ascending=False).head(50)
large_fine_diff.head(10)
all_words.head(10)
top_words_by_fine_size = pd.DataFrame({"no_fine": no_fine_diff.index,
"small_fine":small_fine_diff.index,
"med_fine": med_fine_diff.index,
"large_fine": large_fine_diff.index})
top_words_by_fine_size.head(20)
|
notebooks/03 Data Exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
from astropy.io import fits
import matplotlib.pyplot as plt
from lightkurve import TessLightCurveFile
from astropy.coordinates import SkyCoord
import astropy.units as u
import lightkurve
import theano
import exoplanet as xo
import pymc3 as pm
import theano.tensor as tt
from astropy.stats import BoxLeastSquares
print(xo.__version__)
print(theano.__version__)
print(pm.__version__)
target = 'l 98-59'
tpf = lightkurve.search_targetpixelfile(target).download()
# +
texp = tpf.hdu[1].header['FRAMETIM'] * tpf.hdu[1].header["NUM_FRM"]
texp /= 60.0 * 60.0 * 24.0
ref_time = 0.5 * (tpf.time.min() + tpf.time.max())
flux = tpf.flux
pix_mask = tpf.create_threshold_mask(threshold=3)
tpf.plot(aperture_mask=pix_mask, )
plt.gcf().savefig('pixmask.pdf', dpi=200)
# +
lc = tpf.extract_aperture_photometry(aperture_mask=pix_mask, )
lc = lc.normalize()
lc.flux = (lc.flux - 1.0) * 1e3
# Build the first order PLD basis
X_pld = np.reshape(flux[:, pix_mask], (len(flux), -1))
X_pld = X_pld / np.sum(flux[:, pix_mask], axis=-1)[:, None]
# Build the second order PLD basis and run PCA to reduce the number of dimensions
X2_pld = np.reshape(X_pld[:, None, :] * X_pld[:, :, None], (len(flux), -1))
U, _, _ = np.linalg.svd(X2_pld, full_matrices=False)
X2_pld = U[:, :X_pld.shape[1]]
# Construct the design matrix and fit for the PLD model
X_pld = np.concatenate((np.ones((len(flux), 1)), X_pld, X2_pld), axis=-1)
XTX = np.dot(X_pld.T, X_pld)
w_pld = np.linalg.solve(XTX, np.dot(X_pld.T, lc.flux))
pld_flux = np.dot(X_pld, w_pld)
# Plot the de-trended light curve
plt.figure(figsize=(10, 5))
plt.plot(lc.time, lc.flux-pld_flux, "k")
plt.xlabel("time [days]")
plt.ylabel("de-trended flux [ppt]")
plt.title("initial de-trended light curve")
plt.xlim(lc.time.min(), lc.time.max());
# +
period_grid = np.exp(np.linspace(np.log(0.1), np.log(15), 50000))
duration = 0.03
bls_periods = np.zeros([3])
bls_t0s = np.zeros([3])
bls_depths = np.zeros([3])
time = lc.time - ref_time
# planet 1
bls = BoxLeastSquares(time, lc.flux - pld_flux)
bls_power = bls.power(period_grid, duration, oversample=20)
bls_power0 = bls_power
# Save the highest peak as the planet candidate
index = np.argmax(bls_power.power)
bls_period = bls_power.period[index]
bls_t0 = bls_power.transit_time[index]
bls_depth = bls_power.depth[index]
transit_mask = bls.transit_mask(time, bls_period, 3*duration, bls_t0)
bls_periods[0] = bls_period
bls_t0s[0] = bls_t0
bls_depths[0] = bls_depth
# planet 2
bls = BoxLeastSquares(time[~transit_mask], lc.flux[~transit_mask] - pld_flux[~transit_mask])
bls_power = bls.power(period_grid, duration, oversample=20)
# Save the highest peak as the planet candidate
index = np.argmax(bls_power.power)
bls_period = bls_power.period[index]
bls_t0 = bls_power.transit_time[index]
bls_depth = bls_power.depth[index]
transit_mask = np.logical_or((transit_mask), (bls.transit_mask(time, bls_period, 3*duration, bls_t0)))
bls_periods[1] = bls_period
bls_t0s[1] = bls_t0
bls_depths[1] = bls_depth
# planet 3
bls = BoxLeastSquares(time[~transit_mask], lc.flux[~transit_mask] - pld_flux[~transit_mask])
bls_power = bls.power(period_grid, duration, oversample=20)
# Save the highest peak as the planet candidate
index = np.argmax(bls_power.power)
bls_period = bls_power.period[index]
bls_t0 = bls_power.transit_time[index]
bls_depth = bls_power.depth[index]
transit_mask = np.logical_or((transit_mask), (bls.transit_mask(time, bls_period, 3*duration, bls_t0)))
bls_periods[2] = bls_period
bls_t0s[2] = bls_t0
bls_depths[2] = bls_depth
# +
fig, axes = plt.subplots(4, 1, figsize=(10, 10))
# Plot the periodogram
ax = axes[0]
ax.axvline(np.log10(bls_periods[0]), color="C1", lw=5, alpha=0.8)
ax.axvline(np.log10(bls_periods[1]), color="C1", lw=5, alpha=0.8)
ax.axvline(np.log10(bls_periods[2]), color="C1", lw=5, alpha=0.8)
ax.plot(np.log10(bls_power0.period), bls_power0.power, "k")
ax.annotate("periods = {0:.4f}, {1:.4f}, {2:.4f} d".format(*bls_periods),
(0, 1), xycoords="axes fraction",
xytext=(5, -5), textcoords="offset points",
va="top", ha="left", fontsize=12)
ax.set_ylabel("bls power")
ax.set_yticks([])
ax.set_xlim(np.log10(period_grid.min()), np.log10(period_grid.max()))
ax.set_xlabel("log10(period)")
# Plot the folded transits
bls = BoxLeastSquares(time, lc.flux - pld_flux)
for i in range(3):
ax = axes[i+1]
x_fold = (time - bls_t0s[i] + 0.5*bls_periods[i])%bls_periods[i] - 0.5*bls_periods[i]
m = np.abs(x_fold) < 0.4
ax.plot(x_fold[m], lc.flux[m] - pld_flux[m], ".k")
inds = np.argsort(x_fold[m])
ax.plot(x_fold[m][inds], bls.model(time, bls_periods[i], duration, bls_t0s[i])[m][inds])
# Overplot the phase binned light curve
bins = np.linspace(-0.11, 0.11, 52)
denom, _ = np.histogram(x_fold, bins)
num, _ = np.histogram(x_fold, bins, weights=lc.flux - pld_flux)
denom[num == 0] = 1.0
ax.plot(0.5*(bins[1:] + bins[:-1]), num / denom, color="C1")
ax.set_xlim(-0.1, 0.1)
ax.set_ylabel("de-trended flux [ppt]")
ax.set_xlabel("time since transit");
# +
m = ~transit_mask
XTX = np.dot(X_pld[m].T, X_pld[m])
w_pld = np.linalg.solve(XTX, np.dot(X_pld[m].T, lc.flux[m]))
pld_flux = np.dot(X_pld, w_pld)
x = np.ascontiguousarray(lc.time-ref_time, dtype=np.float64)
y = np.ascontiguousarray(lc.flux-pld_flux, dtype=np.float64)
plt.figure(figsize=(10, 5))
plt.plot(x, y, "k")
plt.xlabel("time [days]")
plt.ylabel("de-trended flux [ppt]")
plt.title("final de-trended light curve")
plt.xlim(x.min(), x.max());
# +
plt.figure(figsize=(10, 5))
x_fold = (x - bls_t0 + 0.5*bls_period) % bls_period - 0.5*bls_period
m = np.abs(x_fold) < 0.3
plt.plot(x_fold[m], pld_flux[m], ".k", ms=4)
bins = np.linspace(-0.5, 0.5, 60)
denom, _ = np.histogram(x_fold, bins)
num, _ = np.histogram(x_fold, bins, weights=pld_flux)
denom[num == 0] = 1.0
plt.plot(0.5*(bins[1:] + bins[:-1]), num / denom, color="C1", lw=2)
plt.xlim(-0.2, 0.2)
plt.xlabel("time since transit")
plt.ylabel("PLD model flux");
# +
def build_model(mask=None, start=None):
if mask is None:
mask = np.ones(len(x), dtype=bool)
with pm.Model() as model:
# Parameters for the stellar properties
mean = pm.Normal("mean", mu=0.0, sd=10.0)
u_star = xo.distributions.QuadLimbDark("u_star",
testval=np.array([0.3, 0.2]))
# Stellar parameters from Kostov et al (2018)
R_star_kostov = 0.291, 0.025
# M_star_kostov = 0.32, 0.025
Rho_star_kostov = 18.0
# m_star = pm.Normal("m_star", mu=M_star_kostov[0], sd=M_star_kostov[1])
r_star = pm.Normal("r_star", mu=R_star_kostov[0], sd=R_star_kostov[1])
logrho_star = pm.Normal("logrho_star", mu=np.log(Rho_star_kostov), sd=0.1)
rho_star = pm.Deterministic("rho_star", tt.exp(logrho_star))
# Prior to require physical parameters
# pm.Potential("m_star_prior", tt.switch(m_star > 0, 0, -np.inf))
pm.Potential("r_star_prior", tt.switch(r_star > 0, 0, -np.inf))
# pm.Potential("rho_star_prior", tt.switch(rho_star > 0, 0, -np.inf))
# Orbital parameters for the planets
logP = pm.Normal("logP", mu=np.log(bls_periods), sd=0.1, shape=3)
t0 = pm.Normal("t0", mu=bls_t0s, sd=0.1, shape=3)
logror = pm.Normal("logror", mu=np.log(np.sqrt(1e-3)*np.sqrt(bls_depths)),
sd=2, shape=3)
ror = pm.Deterministic("r", tt.exp(logror))
b_param = pm.Uniform("b_param", lower=[0,0.3,0], upper=[0.8,1,0.8], shape=3)
b = pm.Deterministic("b", b_param * (1 + ror))
# ror, b = xo.distributions.get_joint_radius_impact(
# min_radius=0.01, max_radius=0.3,
# testval_r=np.sqrt(1e-3)*np.sqrt(bls_depths),
# testval_b=[0.1,0.8,0.1])
ecc = pm.Bound(pm.Beta, lower=0.0, upper=0.8)("ecc", alpha=0.867, beta=3.03,
testval=[0.1,0.1,0.1], shape=3)
# ecc = pm.Beta('ecc', alpha=0.867, beta=3.03, testval=[0.03,0.02,0.04], shape=3)
omega = xo.distributions.Angle("omega", shape=3)
# Log-uniform prior on ror
pm.Potential("ror_prior_lo", tt.switch(tt.all(0.005 < ror), 0.0, -np.inf))
pm.Potential("ror_prior_hi", tt.switch(tt.all(ror < 0.3), 0.0, -np.inf))
# pm.Potential("ror_prior", -tt.log(ror))
# Transit jitter & GP parameters
logs2 = pm.Normal("logs2", mu=np.log(np.var(y[mask])), sd=10)
logS0 = pm.Normal("logS0", mu=np.log(np.var(y[mask])), sd=10)
logw0 = pm.Normal("logw0", mu=np.log(2*np.pi/10), sd=10)
# Tracking planet parameters
period = pm.Deterministic("period", tt.exp(logP))
r_pl = pm.Deterministic("r_pl", r_star * ror)
r_ple = pm.Deterministic("r_ple", (r_star * ror / 0.009155))
# Orbit model
orbit = xo.orbits.KeplerianOrbit(
r_star=r_star, #m_star=m_star,
period=period, t0=t0, b=b,
ecc=ecc, omega=omega,
rho_star=rho_star,
)
a = pm.Deterministic("a", orbit.a)
aAU = pm.Deterministic("aAU", orbit.a * r_star / 215.9)
pm.Potential("ecc_prior", tt.switch((ecc < 1 - (1/a)), 0, -np.inf))
# Compute the model light curve using starry
light_curves = xo.StarryLightCurve(u_star).get_light_curve(
orbit=orbit, r=r_pl, t=x[mask], texp=texp)*1e3
light_curve = pm.math.sum(light_curves, axis=-1)
pm.Deterministic("light_curve", light_curve)
model.light_curves = light_curves
# GP model for the light curve
kernel = xo.gp.terms.SHOTerm(log_S0=logS0, log_w0=logw0, Q=1/np.sqrt(2))
gp = xo.gp.GP(kernel, x[mask], tt.exp(logs2) + tt.zeros(mask.sum()), J=2)
pm.Potential("transit_obs", gp.log_likelihood(y[mask] - light_curve - mean))
pm.Deterministic("gp_pred", gp.predict())
# Fit for the maximum a posteriori parameters, I've found that I can get
# a better solution by trying different combinations of parameters in turn
if start is None:
start = model.test_point
map_soln = pm.find_MAP(start=start, vars=[logs2, logS0, logw0])
# map_soln = pm.find_MAP(start=map_soln, vars=[model.rb])
map_soln = pm.find_MAP(start=map_soln, vars=[model.logror, model.b_param])
map_soln = pm.find_MAP(start=map_soln)
return model, map_soln
model0, map_soln0 = build_model()
# +
def plot_light_curve(model, soln, mask=None):
if mask is None:
mask = np.ones(len(x), dtype=bool)
fig, axes = plt.subplots(3, 1, figsize=(10, 7), sharex=True)
ax = axes[0]
ax.plot(x[mask], y[mask], "k", label="data")
gp_mod = xo.utils.eval_in_model(model.gp_pred, soln, model=model)
gp_mod += soln["mean"]
ax.plot(x[mask], gp_mod, color="C2", label="gp model")
ax.legend(fontsize=10)
ax.set_ylabel("relative flux [ppt]")
ax = axes[1]
ax.plot(x[mask], y[mask] - gp_mod, "k", label="de-trended data")
light_curves = xo.utils.eval_in_model(model.light_curves, soln, model=model)
for i, l in enumerate(["b", "c", 'd']):
mod = light_curves[:, i]
ax.plot(x[mask], mod, label="planet {0}".format(l))
ax.legend(fontsize=10, loc=3)
ax.set_ylabel("de-trended flux [ppt]")
ax = axes[2]
mod = gp_mod + np.sum(light_curves, axis=-1)
ax.plot(x[mask], y[mask] - mod, "k")
ax.axhline(0, color="#aaaaaa", lw=1)
ax.set_ylabel("residuals [ppt]")
ax.set_xlim(x[mask].min(), x[mask].max())
ax.set_xlabel("time [days]")
return fig
plot_light_curve(model0, map_soln0);
# +
mod = map_soln0["gp_pred"] + map_soln0["mean"] + map_soln0["light_curve"]
resid = y - mod
rms = np.sqrt(np.median(resid**2))
mask = np.abs(resid) < 4.5 * rms
plt.figure(figsize=(10, 5))
plt.plot(x, resid, "k", label="data")
plt.plot(x[~mask], resid[~mask], "xr", label="outliers")
plt.axhline(0, color="#aaaaaa", lw=1)
plt.ylabel("residuals [ppt]")
plt.xlabel("time [days]")
plt.legend(fontsize=12, loc=3)
plt.xlim(x.min(), x.max());
# -
model, map_soln = build_model(mask, map_soln0)
plot_light_curve(model, map_soln, mask);
sampler = xo.PyMC3Sampler(window=200, start=200, finish=200)
with model:
burnin = sampler.tune(tune=6000, start=map_soln, step_kwargs=dict(target_accept=0.9))
start, step = sampler._get_start_and_step()
with model:
step = pm.NUTS(potential=step.potential, target_accept=0.9)
trace = pm.sample(draws=3000, tune=200, step=step, start=start)
pm.summary(trace, varnames= ['b_param', 'logror', 't0', 'logP', 'ecc', 'omega',
'logrho_star', 'r_star', 'u_star', 'logw0', 'logS0',
'logs2', 'mean', 'r_ple', 'rho_star'], )
pm.traceplot(trace, varnames=["b_param"])
fig = plt.figure(figsize=[12,8])
pm.forestplot(trace, varnames=["logrho_star", "r_star", "u_star", "mean", "logw0", "logS0", "logs2",
"t0", "logP", "b_param", "logror", "ecc", "omega"])
trace["b"][:,0]
q = pm.summary(trace, varnames= ['logrho_star', 'u_star', 'r_star',
't0', 'logP', 'period', 'b', 'logror', 'ecc', 'omega', 'r',
'a', 'r_ple'], alpha=0.1, )
# +
$\ln{Period [days]}$ & 0.81208 & 0.81232 & 0.81255 \\
Impact parameter & 0.00 & 0.36 & 0.63 \\
$\ln{R_p/R_s}$ & -3.90 & -3.80 & -3.70 \\
ecentricity & 0.00 & 0.17 & 0.38 \\
$\omega$ & -2.6 & 0.0 & 3.2 \\
$\ln{Period [days]}$ & 1.30569 & 1.30578 & 1.30587 \\
Impact parameter & 0.00 & 0.32 & 0.60 \\
$\ln{R_p/R_s}$ & -3.27 & -3.20 & -3.14 \\
ecentricity & 0.00 & 0.17 & 0.39 \\
$\omega$ & -3.1 & -0.5 & 2.3 \\
$\ln{Period [days]}$ & 2.0083 & 2.0084 & 2.0086 \\
Impact parameter & 0.78 & 0.90 & 1.04 \\
$\ln{R_p/R_s}$ & -3.3 & -3.1 & -2.9 \\
ecentricity & 0.00 & 0.19 & 0.47 \\
$\omega$ & -3.1 & 0.1 & 2.6 \\
# -
q.loc[['r__1', 'r_ple__1', 'a__1', ], ['hpd_5', 'mean', 'hpd_95']]
ref_time + q.loc[['t0__1', ], ['hpd_5', 'mean', 'hpd_95']]
# +
# Compute the GP prediction
gp_mod = np.median(trace["gp_pred"] , axis=0)
fig, axes = plt.subplots(3,1, figsize=[10, 8])
for i, pnum in enumerate([2,0,1]):
ax1 = axes[i]
# Get the posterior median orbital parameters
p = np.median(trace["period"][:,pnum])
t0 = np.median(trace['t0'][:,pnum])
# Plot the folded data
x_fold = (x[mask] - t0 + 0.5*p) % p - 0.5*p
ax1.plot(x_fold * 24., y[mask] - gp_mod, ".k", label="data",
alpha=0.5, zorder=-1000, mec="none")
# Plot the folded model
inds = np.argsort(x_fold)
inds = inds[np.abs(x_fold)[inds] < 0.3]
pred = np.add(trace['mean'], trace["light_curve"][:, inds].T ).T
pred = np.percentile(pred, [5, 50, 95], axis=0)
ax1.plot(x_fold[inds] * 24., pred[1], color="C1", label="model")
art = ax1.fill_between(x_fold[inds] * 24., pred[0], pred[2], color="C1", alpha=0.3,
zorder=1000)
art.set_edgecolor("none")
# Annotate the plot with the planet's period
txt = "period = {0:.5f} +/- {1:.5f} days".format(
np.mean(trace["period"][:,pnum]), np.std(trace["period"][:,pnum]))
ax1.annotate(txt, (0, 0.8), xycoords="axes fraction",
xytext=(5, 5), textcoords="offset points",
ha="left", va="bottom", fontsize=12)
txt = "radius = {0:.2f} +/- {1:.2f} $R_\oplus$".format(
np.mean(trace["r_ple"][:,pnum]), np.std(trace["r_ple"][:,pnum]))
ax1.annotate(txt, (0.6, 0.8), xycoords="axes fraction",
xytext=(5, 5), textcoords="offset points",
ha="left", va="bottom", fontsize=12)
ax1.legend(fontsize=10, loc=4)
ax1.set_xlabel("time since transit [hours]")
ax1.set_ylabel("de-trended flux (ppt)")
ax1.set_xlim(-4, 4);
ax1.set_ylim(-4, 4);
plt.tight_layout()
plt.savefig('l98-transit-plot2.png', dpi=200)
# -
np.add(trace['mean'], trace["light_curve"][:, inds].T ).T
np.log(np.sqrt(1e-3)*np.sqrt(bls_depths))
start, step = sampler._get_start_and_step()
with model:
step = pm.NUTS(potential=step.potential, target_accept=0.9)
trace = pm.sample(draws=5000, tune=200, step=step, start=start)
pm.summary(trace, varnames= ['b_param', 'logror', 't0', 'logP', 'ecc', 'omega',
'logrho_star', 'r_star', 'u_star', 'logw0', 'logS0',
'logs2', 'mean', 'r_ple', 'rho_star'], )
pm.save_trace(trace)
|
code/l98-tb1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kentokura/python_medical_images/blob/master/chapter1/Chapter_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LZwtWZNT3Z_H" colab_type="text"
# ## はじめにインストールされているライブラリとバージョンの確認
# ### Linuxのコマンドを利用!!
#
# + id="TCzwAxX24ies" colab_type="code" outputId="ceea3bdf-43eb-4216-8eb4-0f24248048ff" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# ! pip freeze
# + id="q2REuBPH4Xas" colab_type="code" outputId="2dcd5866-ebe5-4714-833f-3c0a98623608" colab={"base_uri": "https://localhost:8080/", "height": 122}
# !pip install pydicom
# + id="ZczV2XYI8GHx" colab_type="code" outputId="5ca4efbb-37e6-4e25-bcd0-159e4c7c3e3c" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/drive')
# + id="L52379Rw9ClL" colab_type="code" colab={}
|
chapter1/Chapter_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import json
overpass_url = "http://overpass-api.de/api/interpreter"
overpass_query = """
[out:json];
area["ISO3166-1"="DE"][admin_level=2];
(node["amenity"="biergarten"](area);
way["amenity"="biergarten"](area);
rel["amenity"="biergarten"](area);
);
out center;
"""
response = requests.get(overpass_url,
params={'data': overpass_query})
data = response.json()
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# Collect coords into list
coords = []
for element in data['elements']:
if element['type'] == 'node':
lon = element['lon']
lat = element['lat']
coords.append((lon, lat))
elif 'center' in element:
lon = element['center']['lon']
lat = element['center']['lat']
coords.append((lon, lat))
# Convert coordinates into numpy array
X = np.array(coords)
plt.plot(X[:, 0], X[:, 1], 'o')
plt.title('Biergarten in Germany')
plt.xlabel('Longitude')
plt.ylabel('Latitude')
plt.axis('equal')
plt.show()
import geopandas
import osmnx as ox
place_name = "Leeds, Leeds, great britain"
graph = ox.graph_from_place(place_name)
type(graph)
fig, ax = ox.plot_graph(graph)
plt.tight_layout()
area = ox.gdf_from_place(place_name)
nodes, edges = ox.graph_to_gdfs(graph)
nodes.head()
edges.highway.tolist()
# +
import osmium as osm
import pandas as pd
class OSMHandler(osm.SimpleHandler):
def __init__(self):
osm.SimpleHandler.__init__(self)
self.osm_data = []
def tag_inventory(self, elem, elem_type):
for tag in elem.tags:
self.osm_data.append([elem_type,
elem.id,
elem.version,
elem.visible,
pd.Timestamp(elem.timestamp),
elem.uid,
elem.user,
elem.changeset,
len(elem.tags),
tag.k,
tag.v])
def node(self, n):
self.tag_inventory(n, "node")
def way(self, w):
self.tag_inventory(w, "way")
def relation(self, r):
self.tag_inventory(r, "relation")
# -
osmhandler = OSMHandler()
osmhandler.apply_file("map.osm")
data_colnames = ['type', 'id', 'version', 'visible', 'ts', 'uid',
'user', 'chgset', 'ntags', 'tagkey', 'tagvalue']
df_osm = pd.DataFrame(osmhandler.osm_data, columns=data_colnames)
#df_osm = tag_genome.sort_values(by=['type', 'id', 'ts'])
df_osm.head()
df_osm.tagkey.unique()
df_osm[df_osm.tagkey=='railway']
|
GPS_data.ipynb
|
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// +
#include <iostream>
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
using namespace std;
#define MAX 500
// -
typedef struct {
int sifra;
char naziv[20 + 1];
}zapis;
void print(zapis tmp){
printf("%d, %s\n", tmp.sifra, tmp.naziv);
}
int adresa(int sifra) {
return sifra % MAX;
}
void print(zapis tablica[]){
// Ispis tablice
for (int i=0; i<MAX;i++)
if(tablica[i].sifra != 0){
printf("%d -> ", i);
print(tablica[i]);
}
}
// **Memorijski rezidentna tablica**
zapis tablica[MAX];
// ### 3. Zadatak
// Napisati funkciju za upis **šifre** (cijeli broj) i **naziva** (20+1) u memorijski rezidentnu tablicu raspršenih adresa s **500 pretinaca**. Pretinac sadrži jedan zapis. Ako je pretinac popunjen, prelazi se **ciklički** na susjedni. Ulazni argumenti su već izračunata adresa pretinca, šifra i naziv. Funkcija vraća vrijednost 1 ako je upis obavljen, 0 ako podatak već postoji, a -1 ako je tablica popunjena pa se podatak nije mogao upisati.
//
int upis(int adr, int sifra, char *naziv)
{
// Ovdje napisati kod
zapis tmp;
tmp.sifra = sifra;
strcpy(tmp.naziv, naziv);
tablica[adr] = tmp;
return 1;
}
upis(6, 102, "artikal");
print(tablica);
// ### 4. Zadatak
//
// Napisati funkciju za pronalaženje **šifre** (cijeli broj) i **naziva ** (20+1) iz memorijski rezidentne tablice raspršenih adresa s **200 pretinaca**. Pretinac sadrži jedan zapis. Ako je pretinac popunjen, a ne sadrži traženu vrijednost ključa, prelazi se ciklički na susjedni. Ulazni argumenti su već izračunata adresa pretinca i šifra. Izlazni argument je naziv poduzeća. Funkcija vraća vrijednost 1 ako je zapis pronađen, a 0 ako nije.
//
int pronadji(int adr, int sifra);
{
// Ovdje napisati kod
}
|
notebooks/rasprsenoAdresiranje/RasprsenoAdresiranjev2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix, classification_report, r2_score, roc_auc_score, accuracy_score, auc
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import GridSearchCV
df = pd.read_csv('breast_cancer.csv')
df.head()
df.diagnosis.value_counts()
df.diagnosis = (df.diagnosis.astype('category')).cat.codes
df.head()
df = df.drop(['id','Unnamed: 32'],1)
df.head()
# # KNN Classification
knn = KNeighborsClassifier()
X = df.drop(['diagnosis'],1)
y = df['diagnosis']
param = {
'n_neighbors' : np.arange(3,12,1),
'metric' : ['manhattan','jaccard','minkowski','euclidean']
}
from sklearn.preprocessing import MinMaxScaler
scalar = MinMaxScaler()
X = pd.DataFrame(scalar.fit_transform(X),columns= list(X))
model = GridSearchCV(cv=10, estimator=knn, param_grid=param)
model = model.fit(X,y)
model.best_estimator_
model.best_score_
# # Logistic Regression
log_reg = LogisticRegression()
param_log = {
'C': [0.001,0.01,0.1,1,10,100],
'penalty': ['l1','l2']
}
model_log = GridSearchCV(cv=10, estimator=log_reg, param_grid=param_log)
model_log = model_log.fit(X,y)
model_log.best_estimator_
model_log.best_score_
model_log.best_params_
# # KNN Classification
# +
# Dividing data into Train, test, and validation data
# -
df.shape
train_test = df[:500]
validation_test = df[500:]
train_test.shape
validation_test.shape
X_train_test = train_test.drop(['diagnosis'],1)
y_train_test = train_test['diagnosis']
X_train_test = pd.DataFrame(scalar.fit_transform(X_train_test),columns= list(X_train_test))
X_validation_test = validation_test.drop(['diagnosis'],1)
y_validation_test = validation_test['diagnosis']
X_validation_test = pd.DataFrame(scalar.fit_transform(X_validation_test),columns= list(X_validation_test))
params = {"n_neighbors":np.arange(2,15,1),
"metric":["euclidean", "minkowski", "manhattan", "jaccard", "cosine"]}
model_knn = GridSearchCV(knn, param_grid= param, cv = 10)
model_knn.fit(X_train_test,y_train_test)
model_knn.best_estimator_
model_knn.best_params_
model_knn.best_score_
y_pred = model_knn.predict(X_validation_test)
accuracy_score(y_validation_test,y_pred)
confusion_matrix(y_validation_test,y_pred)
roc_auc_score(y_validation_test,y_pred)
# # Logistic Regression
train_test_lr = df[:500]
validation_test_lr = df[500:]
X_train_test_lr = train_test_lr.drop(['diagnosis'],1)
y_train_test_lr = train_test_lr['diagnosis']
X_train_test_lr = pd.DataFrame(scalar.fit_transform(X_train_test_lr),columns= list(X_train_test_lr))
X_validation_test_lr = validation_test_lr.drop(['diagnosis'],1)
y_validation_test_lr = validation_test_lr['diagnosis']
X_validation_test_lr = pd.DataFrame(scalar.fit_transform(X_validation_test_lr),columns= list(X_validation_test_lr))
param = {
'C': [0.001,0.01,0.1,1,10,100],
'penalty': ['l1','l2']
}
model_lr = GridSearchCV(cv = 10 , estimator=log_reg, param_grid= param)
model_lr = model_lr.fit(X_train_test_lr,y_train_test_lr)
model_lr.best_score_
y_pred_lr = model_lr.predict(X_validation_test_lr)
model_lr.best_estimator_
accuracy_score(y_pred_lr,y_validation_test_lr)
confusion_matrix(y_pred_lr,y_validation_test_lr)
|
C14_Logistic Regression/Logistic Regression and KNN Classification on Breast Cancer dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Profiling
# Sometimes computing the likelihood is not as fast as we would like. Theano provides handy profiling tools which are wrapped in PyMC3 by `model.profile`. This function returns a `ProfileStats` object conveying information about the underlying Theano operations. Here we'll profile the likelihood and gradient for the stochastic volatility example.
#
# First we build the model.
# +
import numpy as np
import pandas as pd
import pymc3 as pm
print("Running on PyMC3 v{}".format(pm.__version__))
# -
RANDOM_SEED = 8927
np.random.seed(RANDOM_SEED)
# Load the data
returns = pd.read_csv(pm.get_data("SP500.csv"), index_col=0, parse_dates=True)
# Stochastic volatility example
with pm.Model() as model:
sigma = pm.Exponential("sigma", 1.0 / 0.02, testval=0.1)
nu = pm.Exponential("nu", 1.0 / 10)
s = pm.GaussianRandomWalk("s", sigma ** -2, shape=returns.shape[0])
r = pm.StudentT("r", nu, lam=np.exp(-2 * s), observed=returns["change"])
# Then we call the `profile` function and summarize its return values.
# Profiling of the logp call
model.profile(model.logpt).summary()
# Profiling of the gradient call dlogp/dx
model.profile(pm.gradient(model.logpt, model.vars)).summary()
# %load_ext watermark
# %watermark -n -u -v -iv -w
|
docs/source/notebooks/profiling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import folium
print(folium.__version__)
# -
# **Note** : The examples presented below are the copy of the ones presented on https://github.com/bbecquet/Leaflet.PolylineOffset
# ## Basic Demo
# - The dashed line is the "model", with no offset applied.
# - The Red line is with a -5px offset,
# - The Green line is with a 10px offset.
# The three are distinct Polyline objects but uses the same coordinate array
# +
from folium import plugins
m = folium.Map(location=[58.0, -11.0], zoom_start=4, tiles="Mapbox Bright")
coords = [
[58.44773, -28.65234],
[53, -23.33496],
[53, -14.32617],
[58.1707, -10.37109],
[59, -13],
[57, -15],
[57, -18],
[60, -18],
[63, -5],
[59, -7],
[58, -3],
[56, -3],
[60, -4],
]
plugins.PolyLineOffset(
coords, weight=2, dash_array="5,10", color="black", opacity=1
).add_to(m)
plugins.PolyLineOffset(coords, color="#f00", opacity=1, offset=-5).add_to(m)
plugins.PolyLineOffset(coords, color="#080", opacity=1, offset=10).add_to(m)
m.save(os.path.join('results', "PolyLineOffset_simple.html"))
m
# -
# ## Bus Lines
# A more complex demo.
# Offsets are computed automatically depending on the number of bus lines using the same segment.
# Other non-offset polylines are used to achieve the white and black outline effect.
# +
m = folium.Map(location=[48.868, 2.365], zoom_start=15)
geojson = {
"type": "FeatureCollection",
"features": [
{
"type": "Feature",
"properties": {"lines": [0, 1]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.357919216156006, 48.87621773324153],
[2.357339859008789, 48.874834693731664],
[2.362983226776123, 48.86855408432749],
[2.362382411956787, 48.86796126699168],
[2.3633265495300293, 48.86735432768131],
],
},
},
{
"type": "Feature",
"properties": {"lines": [2, 3]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.351503372192383, 48.86443950493823],
[2.361609935760498, 48.866775611250205],
[2.3633265495300293, 48.86735432768131],
],
},
},
{
"type": "Feature",
"properties": {"lines": [1, 2]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.369627058506012, 48.86619159489603],
[2.3724031448364253, 48.8626397112042],
[2.3728322982788086, 48.8616233285001],
[2.372767925262451, 48.86080456075567],
],
},
},
{
"type": "Feature",
"properties": {"lines": [0]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.3647427558898926, 48.86653565369396],
[2.3647642135620117, 48.86630981023694],
[2.3666739463806152, 48.86314789481612],
[2.3673176765441895, 48.86066339254944],
],
},
},
{
"type": "Feature",
"properties": {"lines": [0, 1, 2, 3]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.3633265495300293, 48.86735432768131],
[2.3647427558898926, 48.86653565369396],
],
},
},
{
"type": "Feature",
"properties": {"lines": [1, 2, 3]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.3647427558898926, 48.86653565369396],
[2.3650002479553223, 48.86660622956524],
[2.365509867668152, 48.866987337550164],
[2.369627058506012, 48.86619159489603],
],
},
},
{
"type": "Feature",
"properties": {"lines": [3]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.369627058506012, 48.86619159489603],
[2.372349500656128, 48.865702850895744],
],
},
},
],
}
# manage overlays in groups to ease superposition order
outlines = folium.FeatureGroup("outlines")
line_bg = folium.FeatureGroup("lineBg")
bus_lines = folium.FeatureGroup("busLines")
bus_stops = folium.FeatureGroup("busStops")
line_weight = 6
line_colors = ["red", "#08f", "#0c0", "#f80"]
stops = []
for line_segment in geojson["features"]:
# Get every bus line coordinates
segment_coords = [[x[1], x[0]] for x in line_segment["geometry"]["coordinates"]]
# Get bus stops coordinates
stops.append(segment_coords[0])
stops.append(segment_coords[-1])
# Get number of bus lines sharing the same coordinates
lines_on_segment = line_segment["properties"]["lines"]
# Width of segment proportional to the number of bus lines
segment_width = len(lines_on_segment) * (line_weight + 1)
# For the white and black outline effect
folium.PolyLine(
segment_coords, color="#000", weight=segment_width + 5, opacity=1
).add_to(outlines)
folium.PolyLine(
segment_coords, color="#fff", weight=segment_width + 3, opacity=1
).add_to(line_bg)
# Draw parallel bus lines with different color and offset
for j, line_number in enumerate(lines_on_segment):
plugins.PolyLineOffset(
segment_coords,
color=line_colors[line_number],
weight=line_weight,
opacity=1,
offset=j * (line_weight + 1) - (segment_width / 2) + ((line_weight + 1) / 2),
).add_to(bus_lines)
# Draw bus stops
for stop in stops:
folium.CircleMarker(
stop,
color="#000",
fill_color="#ccc",
fill_opacity=1,
radius=10,
weight=4,
opacity=1,
).add_to(bus_stops)
outlines.add_to(m)
line_bg.add_to(m)
bus_lines.add_to(m)
bus_stops.add_to(m)
m.save(os.path.join('results', "PolyLineOffset_bus.html"))
m
|
examples/plugin-PolyLineOffset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
from sklearn.preprocessing import MinMaxScaler, LabelEncoder
from sklearn.svm import SVR
from sklearn.ensemble import RandomForestRegressor
from sklearn.linear_model import LinearRegression, Ridge
from sklearn.metrics import r2_score
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
sns.set_style("white")
# -
years = [2012, 2013, 2014, 2015, 2016]
# ## Preparing Weather Data
# +
weather_data = pd.DataFrame()
for year in years:
year_data = pd.read_csv('../data/{}.csv'.format(year), index_col=0)
weather_data = pd.concat([weather_data, year_data])
weather_data.shape
# -
weather_data.columns
weather_data.head()
weather_data.drop(weather_data.columns[[0, 1,4,6,8,10,12,14,16]], axis=1, inplace=True)
weather_data['year'] = weather_data['yearmoda'] // 10000
weather_data['month'] = weather_data['yearmoda'] // 100 % 100
weather_data['day'] = weather_data['yearmoda'] % 100
weather_data['sndp'].replace(999.9,0, inplace = True)
weather_data['yearmoda1'] = weather_data['yearmoda'].astype(str).apply(lambda x: x[:9])
weather_data.index = pd.to_datetime(weather_data['yearmoda1'], format='%Y%m%d').values
weather_data.drop(weather_data.columns[[0,16]], axis=1, inplace=True)
for x in weather_data[weather_data['wdsp'] == 999.9].index:
weather_data.loc[x,'wdsp'] = np.NaN
weather_data['wdsp'] = weather_data['wdsp'].interpolate(method='time')
for x in weather_data[weather_data['mxspd'] == 999.9].index:
weather_data.loc[x,'mxspd'] = np.NaN
weather_data['mxspd'] = weather_data['mxspd'].interpolate(method='time')
for x in weather_data[weather_data['prcp'] == '99.99'].index:
weather_data.loc[x,'prcp'] = np.NaN
for x in weather_data.index:
if (str(weather_data.loc[x,'prcp'])[-1] == 'I') or (str(weather_data.loc[x,'prcp'])[-1] == 'H'):
weather_data.loc[x,'prcp'] = np.NaN
for x in weather_data[weather_data['prcp'].notnull()].index:
weather_data.loc[x,'prcp'] = weather_data.loc[x,'prcp'][:-1]
for x in weather_data.index:
if (str(weather_data.loc[x,'max'])[-1] == '*'):
weather_data.loc[x,'max'] = weather_data.loc[x,'max'][:-1]
if (str(weather_data.loc[x,'min'])[-1] == '*'):
weather_data.loc[x,'min'] = weather_data.loc[x,'min'][:-1]
weather_data['max']=weather_data['max'].astype(np.float64)
weather_data['min']=weather_data['min'].astype(np.float64)
weather_data['prcp']=weather_data['prcp'].astype(np.float64)
weather_data['prcp'] = weather_data['prcp'].interpolate(method='time')
weather_data['fog'] = weather_data['frshtt'] // 100000
weather_data['rain'] = weather_data['frshtt'] // 10000 % 10
weather_data['snow'] = weather_data['frshtt'] // 1000 % 10
weather_data['hail'] = weather_data['frshtt'] // 100 % 10
weather_data['thunder'] = weather_data['frshtt'] // 10 % 10
weather_data['tornado'] = weather_data['frshtt'] % 10
weather_data.drop(['frshtt'],axis = 1, inplace = True)
weather_data.head()
columns = [col for col in weather_data.columns if col not in ['month', 'day']]
weather_data = weather_data[columns]
weather_data['week'] = weather_data.index.week
weather_data_week = weather_data.groupby(['year', 'week']).mean()
weather_data_week.head()
weather_data_week.reset_index(inplace=True)
weather_data_week.head()
# ## Preparing Yield Data
def get_year_culture_week_ndvi():
data = pd.DataFrame(columns = ['year', 'culture', 'field', 'week', 'ndvi', 'yields'])
for year in years:
df = pd.read_excel('../Сводная вегетация.xlsx', sheetname=str(year), header=1)
ndvi_columns = [col for col in df.columns if 'неделя' in col]
culture_column = 'Культура ' + str(year)
field_column = 'Поле'
yields_column = 'Урожайность, т./га.'
interesting_columns = [culture_column, field_column] + ndvi_columns + [yields_column]
df = df[interesting_columns]
data_array = []
for i in range(df.shape[0]):
for j in range(2, df.shape[1] - 1):
culture = df.iloc[i][culture_column]
field = df.iloc[i][field_column]
week = df.columns[j].replace('неделя ', '')
ndvi = df.iloc[i, j]
yields = df.iloc[i][yields_column]
row = [year, culture, field, week, ndvi, yields]
data_array.append(row)
data_array = np.array(data_array)
data_frame = pd.DataFrame(data_array, columns=data.columns)
data = pd.concat([data, data_frame[data_frame['ndvi'] != 'nan']])
return data
yield_data = get_year_culture_week_ndvi()
yield_data.head()
yield_data.dtypes
yield_data['year'] = yield_data['year'].astype(int)
yield_data['week'] = yield_data['week'].astype(int)
yield_data['ndvi'] = yield_data['ndvi'].astype(float)
yield_data['yields'] = yield_data['yields'].astype(float)
le = LabelEncoder()
yield_data['field'] = le.fit_transform(yield_data['field'])
yield_data.dropna(inplace=True)
yield_data_culture = yield_data[yield_data['culture'] == 'Подсолнечник'][['year', 'field', 'week', 'ndvi', 'yields']]
yield_data_week = yield_data_culture.groupby(['field', 'year', 'week']).mean()
yield_data_week.head()
yield_data_week.reset_index(inplace=True)
yield_data_week.head()
# ## Combining the datasets
data_week = pd.merge(weather_data_week, yield_data_week, on=['year', 'week'])
data_week.head()
data_week.dropna(inplace=True)
data_week.reset_index(inplace=True)
data_week.iloc[:,1:].head()
data_week.iloc[:,1:].to_csv('~/Desktop/ndvi-weather.csv', index=False)
data_week.shape
# ## Train / test split
train_data = data_week[data_week['year'] != 2015]
test_data = data_week[data_week['year'] == 2015]
# +
x_cols = list(data_week.columns.copy())
x_cols.remove('yields')
X_train = train_data[x_cols]
X_test = test_data[x_cols]
y_train = train_data['yields']
y_test = test_data['yields']
# -
print('X_train:', X_train.shape)
print('X_test:', X_test.shape)
print('y_train:', y_train.shape)
print('y_test:', y_test.shape)
# ## Scaling input values
# +
columns = X_train.columns
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.fit_transform(X_test)
X_train = pd.DataFrame(X_train, columns=columns)
X_test = pd.DataFrame(X_test, columns=columns)
# -
X_train.head()
X_test.head()
# ## Model training
def r2_of(model):
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
return r2_score(y_test, y_pred)
r2_of(RandomForestRegressor())
r2_of(LinearRegression())
r2_of(SVR())
# ## Visualizations
weather_data.temp.plot()
sns.distplot(weather_data[weather_data.year == 2015].temp)
sns.jointplot(x="ndvi", y="temp", data=data_week, kind="kde", color='#4baf9c')
plt.savefig('../img/ndvi-temp.png', transparent=True, dpi=150)
temp_ndvi_yields = data_week[['field', 'temp', 'ndvi', 'yields']].groupby('field').mean()
sns.jointplot(x="temp", y="yields", data=temp_ndvi_yields, kind="kde", color='#4baf9c')
plt.savefig('../img/temp-yields.png', dpi=150)
sns.jointplot(x="ndvi", y="yields", data=temp_ndvi_yields, kind="kde", color='#4baf9c')
plt.savefig('../img/ndvi-yields.png', dpi=150)
# +
x_vars = ['prcp', 'slp', 'stp', 'temp', 'max', 'min']
y_vars = ['ndvi', 'yields']
f = sns.pairplot(data_week, x_vars=x_vars, y_vars=y_vars, palette=['#4baf9c'], kind='reg', diag_kind='kde', diag_kws=dict(shade=True))
sns.despine(left=True, bottom=True)
for i in range(len(y_vars)):
for j in range(len(x_vars)):
f.axes[i, j].get_xaxis().set_visible(False)
f.axes[i, j].get_yaxis().set_visible(False)
# -
data_week.columns
from scipy.stats import spearmanr
spearmanr(data_week['ndvi'], data_week['yields'])
spearmanr(data_week['temp'], data_week['yields'])
spearmanr(data_week['ndvi'], data_week['temp'])
|
notebooks/LongWeatherNDVI.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Example of XOR gate using an MLP and MLP class
#
# ## Author: <NAME> <NAME>
#
# ##### Reference: <http://iamtrask.github.io/2015/07/12/basic-python-network/>
#
import numpy as np
import datetime
import sys
sys.path.append("../")
from mlp import mlp
# ## Examples
#
# #### Using sigmoid
clf = mlp(seed=1, activation="sigmoid", max_iter=10000,
hidden_layer_size=4, alpha=0.1, momentum=0.9,tol=1e-3, weight_range=(-1,1), bias=True)
# %time clf.example_run(show="static")
# #### Using tanh
clf = mlp(seed=1, activation="tanh", max_iter=10000,
hidden_layer_size=4, alpha=0.1, momentum=0.9, tol=1e-3, weight_range=(-1,1), bias=True)
# %time clf.example_run(show="static")
# #### Using softplus
clf = mlp(seed=1, activation="softplus", max_iter=10000,
hidden_layer_size=4, alpha=0.1, momentum=0.9, tol=1e-3, weight_range=(-1,1), bias=True)
# %time clf.example_run(show="static")
# #### Using linear (it doesn't work, and it shouldn't)
clf = mlp(seed=1, activation="linear", max_iter=10000,
hidden_layer_size=4, alpha=0.1, momentum=0.1, tol=1e-3, weight_range=(-1,1), bias=True)
# %time clf.example_run(show="static")
|
xor/xor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-b4b05ac4d498e337", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Analyzing the World Happiness Data
#
#
# ### Grouping subsets of data by cuts
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-8475747c3578a636", "locked": true, "schema_version": 3, "solution": false, "task": false}
# In this exercise, we will continue to examine how to group subsets of data using the split-apply-combine methodology, this time by making various cuts of the data.
#
# As before, we'll repeat here some of the code developed in previous exercises so that we can continue to work with data in this exercise. Execute the next few code cells to load and reconfigure the data. In addition to the full dataset, we will also want to examine the data just from 2015-2017, which we will store in a dataframe named `df1517`.
# + nbgrader={"grade": false, "grade_id": "cell-8d6f09be2e4cabf0", "locked": true, "schema_version": 3, "solution": false, "task": false}
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
pd.options.display.float_format = '{:.2f}'.format
# + nbgrader={"grade": false, "grade_id": "cell-79b10ee0db1c989a", "locked": true, "schema_version": 3, "solution": false, "task": false}
dfraw = pd.read_excel('WHR2018Chapter2OnlineData.xls', sheet_name='Table2.1')
# + nbgrader={"grade": false, "grade_id": "cell-79a1546346ff2981", "locked": true, "schema_version": 3, "solution": false, "task": false}
cols_to_include = ['country', 'year', 'Life Ladder',
'Positive affect','Negative affect',
'Log GDP per capita', 'Social support',
'Healthy life expectancy at birth',
'Freedom to make life choices',
'Generosity', 'Perceptions of corruption']
renaming = {'Life Ladder': 'Happiness',
'Log GDP per capita': 'LogGDP',
'Social support': 'Support',
'Healthy life expectancy at birth': 'Life',
'Freedom to make life choices': 'Freedom',
'Perceptions of corruption': 'Corruption',
'Positive affect': 'Positive',
'Negative affect': 'Negative'}
df = dfraw[cols_to_include].rename(renaming, axis=1)
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-010d6899a146d267", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Step 1
#
# The `groupby` method on a dataframe puts entries into groups that share a common label. In the previous exercise, we were able to group the WHR data by "country" or "year" to get statistics aggregated over those variables. Sometimes, however, we have continuous floating point data that we want to group together into discrete bins. Fortunately, pandas provides mechanisms for converting continuous data into categorical bins.
#
# In the video, we considered the example shown in the code cell below (execute the cell and examine the output):
# + nbgrader={"grade": false, "grade_id": "cell-a03e4de0590fb84a", "locked": true, "schema_version": 3, "solution": false, "task": false}
df.groupby(pd.cut(df.Happiness, bins=list(range(0,11)))).mean()
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-86bef3152ad238a3", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Step 2
#
# Note the index of the dataframe in the output above. These represent Happiness bins: from 0 to 1, 1 to 2, etc., on up to 9 to 10. These bins, or intervals, are regions defined by two endpoints: a lower value and an upper value. But the notation used above to describe these intervals is a little funny, with a parenthesis at the left (lower) end and a square bracket at the right (upper) end. This notation is used to indicate whether the endpoint itself is *included* in the interval: a parenthesis indicates an open interval boundary that does not include the endpoint, whereas a square bracket indicates a closed boundary that does include the endpoint. For example, the interval labeled as `(2, 3]` refers to an interval where Happiness is *greater than* 2 and *less than or equal to* 3. The reason for constructing asymmetric intervals like this is so that every number is contained within one and only one bin: if we instead had intervals such as ```[2,3]``` and ```[3,4]```, then the number ```3``` would belong to both of those bins.
#
# To understand what is going on here, it is useful to tease apart the expression above. The inner part of the code above involves a call to `pd.cut`. The `pd.cut` function is documented [here](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.cut.html). Execute the code cell below and examine the output.
# + nbgrader={"grade": false, "grade_id": "cell-d83049d315f05826", "locked": true, "schema_version": 3, "solution": false, "task": false}
pd.cut(df.Happiness, bins=list(range(0,11)))
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-6d9a3b0757b8c045", "locked": true, "schema_version": 3, "solution": false, "task": false}
# The ```pd.cut``` function puts data into bins based on the values of the data and the location of the bin edges. There are different ways of specifying how you want the bins constructed. One way is to pass an integer, such as 4, and ```pd.cut``` creates that number of equal-size bins that can contain all the data from the minimum to maximum values.
#
# Alternatively, you can specify bins by providing a list of bin edges rather than having pandas calculate them from the data. That is what we have done here when we have specified ```bins=list(range(0,11))```: we have said we want the bin edges to be given by the numbers in the list ```[0,1,2,3,4,5,6,7,8,9,10]```. The ```pd.cut``` function has another option ```right``` with a default value of ```True```: this indicates whether the bins include the rightmost edge or not. Since ```right=True``` by default, we have specified that we want the bins to start at ```0``` and end at ```10```, so that we get the following set of bins:
#
# ```
# [(0, 1] < (1, 2] < (2, 3] < (3, 4] ... (6, 7] < (7, 8] < (8, 9] < (9, 10]]
# ```
# We can see that the result of this call to `pd.cut` is a Series where each row of the `df.Happiness` data is mapped to a particular bin within this collection of 10 bins starting at ```0``` and ending at ```10```.
#
# In other words, the continuous floating point data in `df.Happiness` have been discretized into a set of categories. And now that we have a set of discrete categories, we can group on those as we would with any other label.
#
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-37f3ca8ab60a6fb6", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Step 3
#
# Let's try binning on something other than the Happiness column, say, the LogGDP data instead. In this case, instead of providing a specific list of bin edges, let's just specify the total number of bins we want and let pandas construct the bins based on the data.
#
# In the code cell below, group the WHR data into 10 bins in LogGDP, and compute the total number of entries in each bin by applying the `size` method. Assign the result to the variable `LogGDP_bins`. Examine the result. You should notice that the bin edges are not integers as in the previous example, since pandas has decided to subdivide the full interval ranging from 6.372 to 11.77 into 10 equal-size bins.
# -
# ## Graded Cell
#
# This cell is worth 50% of the grade for this assignment.
# + nbgrader={"grade": false, "grade_id": "cell-e8d6d83c35d859dc", "locked": false, "schema_version": 3, "solution": true, "task": false}
LogGDP_bins = df.groupby(pd.cut(df.LogGDP, bins=10)).size()
# -
# ## Self-Check
#
# Run the cell below to test the correctness of your code above before submitting for grading.
# +
# Run this self-test cell to check your code; do not add code or delete code in this cell
from jn import testBins
try:
print(testBins(LogGDP_bins))
except Exception as e:
print("Error!\n" + str(e))
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-975cbb72578a4d39", "locked": true, "schema_version": 3, "solution": false, "task": false}
# If you want, you can see what the distribution of LogGDP values produced by this cut looks like by plotting ```LogGDP_bins```. Execute the code cell below.
# + nbgrader={"grade": false, "grade_id": "cell-61eb38341ed4781e", "locked": true, "schema_version": 3, "solution": false, "task": false}
LogGDP_bins.plot(rot=90)
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-e24df81846d227fa", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Step 4
#
# In addition to `pd.cut`, which cuts the data into prescribed bins, there is also `pd.qcut`, which cuts data into prescribed quantiles, such that each quantile contains (approximately) the same number of entries. We specify how many bins we want to cut the data into, and pandas decides where the bin edges need to be located to divide into bins with the same number of entries. If we wanted to cut into 4 bins, we could pass the parameter `q=4` to `pd.qcut`: this would group the values into bins for the ranges 0%-25%, 25%-50%, 50%-75%, and 75%-100%. The term "quantile" is general, and the specific division depends on the value of ```q```. But for a few particular values of ```q```, special names are typically used in the statistics community: for example, ```q=4``` produces "quartiles", ```q=5``` produces "quintiles", and ```q=10``` produces "deciles".
#
# In the code cell below, group the WHR data into quartiles in LogGDP, and compute the total number of entries in each bin by applying the `size` method. Assign the result to the variable `LogGDP_quartiles`. Examine the result. Note that the number of entries in each bin are approximately equal (as desired), and that the values defining each bin are now chosen to accomplish the split into 4 bins with the same number of elements. The bins will contain exactly the same number of entries only if the total size of the dataset is evenly divisible by ```q``` (without remainder). If the dataset is not evenly divisible, then the number of entries in bins can differ by 1.
# -
# ## Graded Cell
#
# This cell is worth 50% of the grade for this assignment.
# + nbgrader={"grade": false, "grade_id": "cell-6d43d8c714862019", "locked": false, "schema_version": 3, "solution": true, "task": false}
LogGDP_quartiles = df.groupby(pd.qcut(df.LogGDP, q=4)).size()
# -
# ## Self-Check
#
# Run the cell below to test the correctness of your code above before submitting for grading.
# +
# Run this self-test cell to check your code; do not add code or delete code in this cell
from jn import testQuartiles
try:
print(testQuartiles(LogGDP_quartiles))
except Exception as e:
print("Error!\n" + str(e))
|
GroupingCuts.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# +
# %cd ..
from nsopy_stoch.benchmark_problems.generate_caroe_schultz_datasets import generate_caroe_schultz_cor_file
from nsopy_stoch.benchmark_problems.generate_caroe_schultz_datasets import generate_caroe_schultz_sto_and_tim_files
# -
# # Examine the norm of dual optimizers and convergence rates @ different $|S|$
from nsopy.subgradient_methods import SubgradientMethod as SG
from nsopy.universal_gradient_methods import UniversalPGM, OneNormUPGM
from nsopy.quasi_monotone_subgradient_methods import SGMTripleAveraging as TA
from nsopy.method_loggers import EnhancedDualMethodLogger
from nsopy_stoch.oracles import TwoStage_SMPS_InnerProblem
# +
# Setup
BENCHMARKS_PATH = './nsopy_stoch/benchmark_problems/2_caroe_schultz/'
n_S_exp = [10, 50, 100, 500, 1000]
GAMMA_0_TA = 0.1
# +
# Generate IP and all methods
inner_problems = []
methods = []
methods_nonscaled = []
method_loggers = []
method_loggers_nonscaled = []
for n_S in n_S_exp: # [100, ]:
ip = TwoStage_SMPS_InnerProblem(BENCHMARKS_PATH+'caroe_schultz_{}'.format(n_S))
inner_problems.append(ip)
# upgm = UniversalPGM(oracle=ip.oracle, projection_function=ip.projection_function, dimension=ip.dimension)
# one_norm_upgm = OneNormUPGM(oracle=ip.oracle, projection_function=ip.projection_function, dimension=ip.dimension)
ta = TA(oracle=ip.oracle, projection_function=ip.projection_function, dimension=ip.dimension, gamma=GAMMA_0_TA*n_S)
ta_nonscale_stepsize = TA(oracle=ip.oracle, projection_function=ip.projection_function, dimension=ip.dimension, gamma=GAMMA_0_TA)
# logger_upgm = EnhancedDualMethodLogger(upgm)
# logger_one_norm_upgm = EnhancedDualMethodLogger(one_norm_upgm)
logger_ta = EnhancedDualMethodLogger(ta)
logger_ta_nonscaled = EnhancedDualMethodLogger(ta_nonscale_stepsize)
# methods.append(upgm)
# methods.append(one_norm_upgm)
methods.append(ta)
methods_nonscaled.append(ta_nonscale_stepsize)
# method_loggers.append(logger_upgm)
# method_loggers.append(logger_one_norm_upgm)
method_loggers.append(logger_ta)
method_loggers_nonscaled.append(logger_ta_nonscaled)
# +
for method in methods:
for step in range(1000):
if not step % 100:
print('step: '+str(step)+' of method'+str(method))
method.dual_step()
for method in methods_nonscaled:
for step in range(1000):
if not step % 100:
print('step: '+str(step)+' of (nonscaled) method'+str(method))
method.dual_step()
# -
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib notebook
# +
# small hack to find d*
d_stars = []
# STANDARD
# for method_logger in method_loggers:
# d_stars.append(max(method_logger.d_k_iterates))
# Scaled vs NONSCALED
for logger_ix, method_logger in enumerate(method_loggers):
d_stars.append(max(max(method_loggers[logger_ix].d_k_iterates), max(method_loggers_nonscaled[logger_ix].d_k_iterates)))
# Version where I compare UPGM with 1-norm UPGM
# # 10
# ds = max(max(method_loggers[0].d_k_iterates), max(method_loggers[1].d_k_iterates))
# d_stars.append(ds)
# d_stars.append(ds)
# # 50
# ds = max(max(method_loggers[2].d_k_iterates), max(method_loggers[3].d_k_iterates))
# d_stars.append(ds)
# d_stars.append(ds)
# # 100
# ds = max(max(method_loggers[4].d_k_iterates), max(method_loggers[5].d_k_iterates))
# d_stars.append(ds)
# d_stars.append(ds)
# -
d_stars
# +
# PLOT PROGRESS OF METHODS
plt.figure(facecolor='w')
for logger_ix, logger in enumerate(method_loggers):
# plt.plot(logger.oracle_calls, logger.d_k_iterates,
# label='N scen = {}, {}'.format(methods[logger_ix].dimension/2, methods[logger_ix].desc))
# plt.plot(logger.oracle_calls, [d_stars[logger_ix] - item for item in logger.d_k_iterates],
# label='N scen = {}, {}'.format(methods[logger_ix].dimension/2, methods[logger_ix].desc))
plt.loglog([call for call in logger.oracle_calls], [d_stars[logger_ix] - item + 0.001 for item in logger.d_k_iterates],
label='N scen = {}, {}'.format(float(methods[logger_ix].dimension)/2.0, methods[logger_ix].desc))
# plt.loglog([call+1 for call in logger.oracle_calls], [-item for item in logger.d_k_iterates],
# label='N scen = {}'.format(n_S_exp[logger_ix]))
# plt.loglog([call+1 for call in logger.oracle_calls], [-item for item in logger.d_k_iterates],
# label='N scen = {}, {}'.format(methods[logger_ix].dimension/2, methods[logger_ix].desc))
# # PLOT UPPER BOUND
# INSTANCE = 6
# GAMMA = 1.0
# P_MAX_UB = 0
# D_0 = method_loggers[INSTANCE].d_k_iterates[0]
# RHO = 2.5
# DIAM_X_SQR = 50
# R_D = float(P_MAX_UB - D_0)/float(RHO)
# upper_bound = [-method_loggers[INSTANCE].d_k_iterates[-1] + 1.0/np.sqrt(1+k)*(2*GAMMA*R_D*R_D + 1.0/(float(GAMMA))*DIAM_X_SQR) for k in method_loggers[0].oracle_calls]
# Plot stuff
# plt.loglog([call+1 for call in method_loggers[0].oracle_calls], upper_bound, label='Uppber bound (N={})'.format(n_S_exp[INSTANCE]))
plt.xlabel('iteration $k$')
plt.ylabel('$d^\star - d(\lambda_k)$')
plt.legend()
# +
# PLOT PROGRESS OF METHODS
plt.figure(facecolor='w')
for logger_ix, logger in enumerate(method_loggers_nonscaled):
# plt.plot(logger.oracle_calls, logger.d_k_iterates,
# label='N scen = {}, {}'.format(methods[logger_ix].dimension/2, methods[logger_ix].desc))
# plt.plot(logger.oracle_calls, [d_stars[logger_ix] - item for item in logger.d_k_iterates],
# label='N scen = {}, {}'.format(methods[logger_ix].dimension/2, methods[logger_ix].desc))
plt.loglog([call for call in logger.oracle_calls], [d_stars[logger_ix] - item + 0.001 for item in logger.d_k_iterates],
label='N scen = {}, {}'.format(float(methods[logger_ix].dimension)/2.0, methods_nonscaled[logger_ix].desc))
# plt.loglog([call+1 for call in logger.oracle_calls], [-item for item in logger.d_k_iterates],
# label='N scen = {}'.format(n_S_exp[logger_ix]))
# plt.loglog([call+1 for call in logger.oracle_calls], [-item for item in logger.d_k_iterates],
# label='N scen = {}, {}'.format(methods[logger_ix].dimension/2, methods[logger_ix].desc))
# # PLOT UPPER BOUND
# INSTANCE = 6
# GAMMA = 1.0
# P_MAX_UB = 0
# D_0 = method_loggers[INSTANCE].d_k_iterates[0]
# RHO = 2.5
# DIAM_X_SQR = 50
# R_D = float(P_MAX_UB - D_0)/float(RHO)
# upper_bound = [-method_loggers[INSTANCE].d_k_iterates[-1] + 1.0/np.sqrt(1+k)*(2*GAMMA*R_D*R_D + 1.0/(float(GAMMA))*DIAM_X_SQR) for k in method_loggers[0].oracle_calls]
# Plot stuff
# plt.loglog([call+1 for call in method_loggers[0].oracle_calls], upper_bound, label='Uppber bound (N={})'.format(n_S_exp[INSTANCE]))
plt.xlabel('iteration $k$')
plt.ylabel('$d^\star - d(\lambda_k)$')
plt.legend()
# -
# +
plt.figure(facecolor='w')
plt.semilogx(n_S_exp, [np.linalg.norm(logger.lambda_k_iterates[-1],2) for logger in method_loggers], '.-')
plt.xlabel('scenarios #')
plt.ylabel('$\| \lambda_{k=\mathrm{end}} \|_2$')
plt.legend()
# +
plt.figure(facecolor='w')
for logger_ix, logger in enumerate(method_loggers):
plt.loglog([call+1 for call in logger.oracle_calls], [np.linalg.norm(iterate,np.inf) for iterate in logger.lambda_k_iterates], label='N scen = {}'.format(n_S_exp[logger_ix]))
plt.xlabel('iteration $k$')
plt.ylabel('$\| \lambda_{k} \|_2$')
plt.legend()
# -
# # Generate Caroe & Schultz Datasets
# desired number of scenarios
# n_S_exp = [10, 50, 100, 500, 1000, 5000, 10000]
n_S_exp = [10, 50, 100, 500, 1000]
for n_S in n_S_exp:
generate_caroe_schultz_cor_file(n_S=n_S)
generate_caroe_schultz_sto_and_tim_files(n_S=n_S)
# Uppber Bound on d(0):
3.0/2.0*5+20+16+19+23+28
|
notebooks/0. Caroe - Schultz Investigation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import scipy.spatial.distance as dist
import matplotlib.pyplot as plt
import sklearn.svm as svm
import sklearn.metrics as metrics
# +
num_train_samples = 400
num_test_samples = 1000
a1 = np.random.randn(2, np.int(num_test_samples*0.025))*0.2
a2 = np.random.randn(2, np.int(num_test_samples*0.025))*0.2
a3 = np.random.randn(2, np.int(num_test_samples*0.025))*0.2
d1 = np.random.randn(2, np.int(num_test_samples*0.6))
n = num_test_samples - d1.shape[1] - a1.shape[1] - a2.shape[1] - a3.shape[1]
d2 = np.random.randn(2, n)*0.5
a1_ = np.random.randn(2, np.int(num_train_samples*0.025))*0.2
d1_ = np.random.randn(2, np.int(num_train_samples*0.6))
n = num_train_samples - d1_.shape[1] - a1_.shape[1]
d2_ = np.random.randn(2, n)*0.5
d2[0, :] += 4.
a1[1, :] += 4.
a2[0, :] += 4.
a3[0, :] -= 3.
a2[1, :] -= 3.
a3[1, :] -= 3.
d2_[0, :] += 4.
a1_[1, :] += 4.
num_train_anomalies = a1_.shape[1]
num_test_anomalies = a1.shape[1] + a2.shape[1] + a3.shape[1]
polluted_train_data = np.concatenate((a1_, d1_, d2_), axis=1)
polluted_labels = np.zeros(polluted_train_data.shape[1], dtype=np.int)
polluted_labels[:num_train_anomalies] = 1
semi_supervised_train_data = np.concatenate((d1_, d2_), axis=1)
test_data = np.concatenate((a1, a2, a3, d1, d2), axis=1)
test_labels = np.zeros(test_data.shape[1], dtype=np.int)
test_labels[:num_test_anomalies] = 1
colors = np.array(['b', 'r'])
fig = plt.figure(figsize=(16, 6), dpi= 80, facecolor='w', edgecolor='k')
plt.subplot(1, 2, 1)
plt.title('Training data', fontsize=16)
plt.scatter(polluted_train_data[0, :], polluted_train_data[1, :], 60, colors[polluted_labels], alpha=0.6)
plt.grid()
plt.xticks([], [], fontsize=14)
plt.yticks([], [], fontsize=14)
plt.subplot(1, 2, 2)
plt.scatter(test_data[0, :], test_data[1, :], 60, colors[test_labels], alpha=0.6)
plt.title('Test data', fontsize=16)
plt.grid()
plt.xticks([], [], fontsize=14)
plt.yticks([], [], fontsize=14)
fig.tight_layout()
plt.savefig('anomaly_detection_learning_setting.pdf')
# +
binary_classifier = svm.SVC()
binary_classifier.fit(polluted_train_data.T, polluted_labels)
oneclass_classifier = svm.OneClassSVM()
oneclass_classifier.fit(polluted_train_data.T)
train_pred = binary_classifier.predict(polluted_train_data.T)
print(metrics.roc_auc_score(polluted_labels, train_pred))
train_pred = oneclass_classifier.predict(polluted_train_data.T)
print(metrics.roc_auc_score(polluted_labels, -train_pred))
test_pred = binary_classifier.predict(test_data.T)
svm_auc = metrics.roc_auc_score(test_labels, test_pred)
test_pred = oneclass_classifier.predict(test_data.T)
ocsvm_auc = metrics.roc_auc_score(test_labels, -test_pred)
delta = 0.025
x = np.arange(-6.0, 6.0, delta)
y = np.arange(-6.0, 6.0, delta)
X, Y = np.meshgrid(x, y)
dx = np.reshape(X, (1, X.size))
dy = np.reshape(Y, (1, Y.size))
binary_dz = binary_classifier.decision_function(np.concatenate((dx, dy), axis=0).T)
oneclass_dz = oneclass_classifier.decision_function(np.concatenate((dx, dy), axis=0).T)
bZ = np.reshape(binary_dz, X.shape)
ocZ = np.reshape(oneclass_dz, X.shape)
fig = plt.figure(figsize=(16, 8), dpi= 80, facecolor='w', edgecolor='k')
plt.subplot(1, 2, 1)
CS = plt.contourf(X, Y, bZ, 20, cmap='bone')
# CS = plt.contour(X, Y, Z, [0.0, 0.5, 1.0], cmap='gray')
plt.scatter(test_data[0, :], test_data[1, :], 60, colors[test_labels], alpha=0.6)
plt.title('Supervised SVM (AUC={0:1.2f})'.format(svm_auc), fontsize=16)
plt.grid()
plt.xticks([], [], fontsize=14)
plt.yticks([], [], fontsize=14)
plt.subplot(1, 2, 2)
CS = plt.contourf(X, Y, ocZ, 20, cmap='bone')
# CS = plt.contour(X, Y, Z, [0.0, 0.5, 1.0], cmap='gray')
plt.scatter(test_data[0, :], test_data[1, :], 60, colors[test_labels], alpha=0.6)
plt.title('One-class SVM (AUC={0:1.2f})'.format(ocsvm_auc), fontsize=16)
plt.grid()
plt.xticks([], [], fontsize=14)
plt.yticks([], [], fontsize=14)
fig.tight_layout()
plt.savefig('supervised_vs_unsupervised.pdf')
# -
|
notebooks/anomaly_detection_setting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cross-validated MANOVA in Python
# I've worked quite a bit on 'decoding (machine-learning) analyses' of fMRI data. A while back, I came across the article by [<NAME> (2014)](http://www.sciencedirect.com/science/article/pii/S1053811913011920) on 'cross-validated MANOVA', an multivariate encoding-type of analysis. In this notebook, I've tried to implement this analysis and annotated some of the steps.
# # MGLM
# The MANOVA is a test derived from the multivariate general linear model (MGLM). Conceptually, a MANOVA tests how much variance in a set of target variables (dependent variables; $\mathbf{y}$) by a (set of) predictor(s) ($\mathbf{X}$); in other words, it's a "multivariate analysis of variance", as the name implies. In my understanding (which is often more conceptual than mathematical), the first step in the analysis described by Allefeld & Haynes is to simply calculate the parameters of the GLM ($\beta$) for all target variables. The GLM, here, is defined as:
#
# \begin{align}
# y = \beta\mathbf{X} + \epsilon
# \end{align}
#
# in which $\beta$ represents the GLM's parameter(s) and $\epsilon$ the model's errors. Now, the parameter(s) $\beta$ can be found by the GLM's analytical solution:
#
# \begin{align}
# \hat{\beta} = (\mathbf{X}'\mathbf{X})^{-1}y
# \end{align}
#
# In which $X$ represents the design-matrix (predictors), a $N\ (samples) \times P\ (predictors)$ matrix, and $y$ represents a $N \times 1$ column-vector with the target variable. I recently found at that you can estimate the parameters for different target variables (so different $y$ variables) corresponding to the model:
#
# \begin{align}
# \mathbf{y} = \beta\mathbf{X} + \Xi
# \end{align}
#
# in which $\mathbf{y}$ is now a $N \times K$ (number of target variables) matrix and $\Xi$ a $N \times K$ matrix with the model errors. As such, the parameters of this model can be found by vectorizing the previously outlined formula for the (univariate) GLM as follows:
#
# \begin{align}
# \hat{\mathbf{\beta}} = (\mathbf{X}'\mathbf{X})^{-1}\mathbf{y}
# \end{align}
#
# in which, now, $\mathbf{y}$ is an $N \times K$ (target-variables) matrix and $\beta$ is an $P \times K$ matrix.
#
# Let's check out an example. We'll use the 'canonical' Iris-dataset, which contains data of three types of Iris flowers (let's say, class C1, C2, and C3) associated with four types of variables (let's call them V1, V2, V3, and V4). In the context of a MANOVA, we could investigate whether the factor 'flower type' has a multivariate effect on the four measured variables. Thus, here flower type represents the design-matrix ($\mathbf{X}$) and the measured variables represent the target variables ($\mathbf{y}$). Note, this may seem counterintuitive for people that are familiar with this dataset in the context of machine learning, in which it is used to predict flower type ($y$) as a function of the flower features ($\mathbf{X}$). This is, essentially, just a manifestation of a different question about the data. In neuroimaging, however, this difference is quite important (check out the awesome article by [Naselaris and colleagues](http://www.sciencedirect.com/science/article/pii/S1053811910010657) on this topic).
#
# Anyway, let's check out how the parameters are calculated in the MGLM. We'll first load the data:
# +
from sklearn.datasets import load_iris
import numpy as np
y, flower_types = load_iris(return_X_y=True)
print("Shape of y (flower properties): %s" % (y.shape,))
print("Shape of flower types variable: %s" % (flower_types.shape,), end='\n\n')
print(flower_types)
# -
# To investigate the effect of flower type on the target variables in the context of the GLM, we need to create a separate predictor for each class (C1, C2, C3), which contains all zeros except for the samples that correspond to that particular class. This process is known (albeit in a different context) as 'one-hot encoding'. Let's do this:
from sklearn.preprocessing import OneHotEncoder
ohe = OneHotEncoder(sparse=False)
X = ohe.fit_transform(flower_types[:, np.newaxis]).astype(float)
print("Shape of X (flower type): %s" % (X.shape,))
print(X)
# Sweet! Exactly what we wanted. Now, let's solve the parameters of this GLM-model:
betas = np.linalg.pinv(X.T.dot(X)).dot(X.T).dot(y)
print("Shape of betas: %s" % (betas.shape,))
print(betas)
# In fact, the betas (given this design-matrix) represent the mean of each class (rows) for each variable (columns).
#
# ## The MANOVA
# Now, the Allefeld & Haynes paper define a 'pattern-distinctness' statistic $\hat{D}$, which is based on the Barlett-Lawley-Hotelling Trace $T_{BLH}$, which represents the trace of the product of the between-class covariance (I call it $B$) and the inverse of the within-class covariance (they call it $\Sigma$). Thus, the formula for the Barlett-Lawley-Hotelling trace become:
#
# \begin{align}
# T_{BLH} = \mathrm{trace}(B\Sigma^{-1})
# \end{align}
#
# In the paper, $B$ is calculated using the following formula (this becomes a bit hairy):
#
# \begin{align}
# B = \hat{\beta}_{\Delta}'\mathbf{X}'\mathbf{X}\hat{\beta}_{\Delta}
# \end{align}
#
# in which $\hat{\beta}_{\Delta}$ is calculated using the contrast-matrix $C$ as follows:
#
# \begin{align}
# \hat{\beta}_{\Delta} = (C'C)^{-1}C\hat{\beta}
# \end{align}
#
# Fortunately, $\Sigma$ is simply the inner product of the residuals from the model:
#
# \begin{align}
# \Sigma = (\mathbf{y} - \hat{\beta}\mathbf{X})'(\mathbf{y} - \hat{\beta}\mathbf{X}) = \hat{\Xi}'\hat{\Xi}
# \end{align}
#
# Alright, but what contrast should we use? Well, that of course depends on your question. In our case, we said we were interested in the effect of flower type on the target variables. Given that we have three flower types (C1, C2, C3), an F-test of $K - 1$ pairwise difference contrasts (e.g. C1 - C2 and C1 - C3) should do the trick. As such, we could define $C$ as:
#
# \begin{align}
# C =
# \begin{bmatrix}
# 1 & -1 & 0\\1 & 0 & -1
# \end{bmatrix}
# \end{align}
#
# Let's define it below
C = np.array([
[1, 0, -1],
[0, 1, -1]
])
# Now, let's calculate $\hat{\beta}_{\Delta}$ ...
beta_delta = C.T.dot(np.linalg.pinv(C.T.dot(C)).dot(C.T).T).dot(betas)
print("Shape B-delta: %s" % (beta_delta.shape,))
# ... and $B$:
B = beta_delta.T.dot(X.T.dot(X)).dot(beta_delta)
# ... $\Sigma$:
sigma = (y - X.dot(betas)).T.dot((y - X.dot(betas)))
# ... and finally $T_{BLH}$!
T_blh = np.trace(B.dot(np.linalg.inv(sigma)))
print(T_blh)
# I always check whether I got the right result by checking it against some existing implementation. Fortunately, there is a MANOVA implementation in the master branch (not yet op PyPI) of the statsmodels package. So, let's check whether it matches:
# +
from statsmodels.multivariate.manova import MANOVA
mglm = MANOVA(endog=y, exog=X)
fitted_model = mglm.mv_test([
('One-way MANOVA', C)
])
fitted_model.summary()
# -
# Nice! Exactly the same (the value in statsmodels is called *Hotelling-Layley trace*).
# ## The *cross-validated* MANOVA
# Importantly, the paper suggests to cross-validate the MANOVA procedure (and thus the calculation of the statistic of interest, $\hat{D}$), because $\hat{D}$ is a distance-measure -- essentially an extension of the mahalanobis distance for more than two groups (levels) -- which are inherently positively biased because distances cannot be negative. Cross-validated makes sure the estimate becomes unbiased.
# +
from sklearn.model_selection import StratifiedKFold
M = 5
cv = StratifiedKFold(n_splits=5)
T_blh_cv = np.zeros(5)
for i, (train_idx, test_idx) in enumerate(cv.split(X=y, y=flower_types)):
X_train, X_test = X[train_idx], X[test_idx]
y_train, y_test = y[train_idx], y[test_idx]
betas_train = np.linalg.pinv(X_train.T.dot(X_train)).dot(X_train.T).dot(y_train)
betas_test = np.linalg.pinv(X_test.T.dot(X_test)).dot(X_test.T).dot(y_test)
beta_delta_train = C.T.dot(np.linalg.pinv(C.T.dot(C)).dot(C.T).T).dot(betas_train)
beta_delta_test = C.T.dot(np.linalg.pinv(C.T.dot(C)).dot(C.T).T).dot(betas_test)
B_cv = beta_delta_train.T.dot(X_test.T.dot(X_test)).dot(beta_delta_test)
sigma_cv = (y_train - X_train.dot(betas_train)).T.dot((y_train - X_train.dot(betas_train)))
T_blh_cv[i] = np.trace(B_cv.dot(np.linalg.inv(sigma_cv)))
print(T_blh_cv.mean())
N = X.shape[0]
((M - 1)*(N - np.linalg.matrix_rank(X))-y.shape[1]-1) / ((M - 1) * N)
# -
# Hmm, this is much lower than the uncrossvalidated statistic. Need to check up on this.
|
cv_MANOVA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Passionate Investors and Cyberpunks. ERC20 Analysis
# + pycharm={"name": "#%%\n"}
import pandas as pd
from math import log10
from tqdm.notebook import tqdm_notebook
from config import PROJECT_ID, INITIAL_TS, SNAPSHOT_TS, \
ETH_ANALYSIS_DATASET_NAME, ETH_ANALYSIS_DISTRIBUTION_TABLE_NAME, \
ERC20_ANALYSIS_DATASET_NAME, ERC20_ROW_BALANCES_TABLE_NAME, ERC20_REVISED_BALANCES_TABLE_NAME, \
ERC20_BALANCES_TABLE_NAME, ERC20_ANALYSIS_DISTRIBUTION_TABLE_NAME, ERC20_TOKEN_METADATA_TABLE_NAME, \
STABLECOINS_LOVERS_TABLE_NAME, INVESTORS_AUDIENCE, CYBERPUNKS_AUDIENCE
from src.utils_bigquery import drop_table, create_table, create_table_from_df, get_df
from src.utils_charts import grade_boundaries_analysis
from src.extractor_balances import get_balances
DROP_TABLES = True
CREATE_TABLES = True
REVISE_TOKEN_BALANCES = False
# + pycharm={"name": "#%%\n"}
investors_min_number_of_tokens = 2
erc20_balance_initial_boundary = 0.01
erc20_tokens_dict = {
# name: [address, type, ETH price, decimals]
'YFI': ['0x0bc529c00c6401aef6d220be8c6ea1667f6ad93e', 'investment', 7.6, 18],
'FTM': ['0x4e15361fd6b4bb609fa63c81a2be19d873717870', 'investment', 0.00064, 18],
'AMPL': ['0xd46ba6d942050d489dbd938a2c909a5d5039a161', 'investment', 0.00034, 9],
'renBTC': ['0xeb4c2781e4eba804ce9a9803c67d0893436bb27d', 'investment', 13.8, 8],
'OCEAN': ['0x967da4048cd07ab37855c090aaf366e4ce1b9f48', 'investment', 0.000215, 18],
'RPL': ['0xb4efd85c19999d84251304bda99e90b92300bd93', 'investment', 0.012, 18],
'UNI': ['0x1f9840a85d5af5bf1d1762f925bdaddc4201f984', 'investment', 0.0057, 18],
'KEEP3R': ['0x1ceb5cb57c4d4e2b2433641b95dd330a33185a44', 'investment', 0.072, 18],
'GST2': ['0x0000000000b3f879cb30fe243b4dfee438691c04', 'investment', 0.017, 2],
'CHI': ['0x0000000000004946c0e9f43f4dee607b0ef1fa1c', 'investment', 0.0007, 0],
'MLN': ['0xec67005c4e498ec7f55e092bd1d35cbc47c91892', 'investment', 0.031, 18],
'AAVE': ['0x7fc66500c84a76ad7e9c93437bfc5ac33e2ddae9', 'investment', 0.076, 18],
'GLM': ['0x7dd9c5cba05e151c895fde1cf355c9a1d5da6429', 'investment', 0.000126, 18],
'ZRX': ['0xe41d2489571d322189246dafa5ebde1f4699f498', 'investment', 0.00026, 18],
'SNX': ['0xc011a73ee8576fb46f5e1c5751ca3b9fe0af2a6f', 'investment', 0.0023, 18],
'SUSHI': ['0x6b3595068778dd592e39a122f4f5a5cf09c90fe2', 'investment', 0.0028, 18],
'COMP': ['0xc00e94cb662c3520282e6f5717214004a7f26888', 'investment', 0.081, 18],
'BAL': ['0xba100000625a3754423978a60c9317c58a424e3d', 'investment', 0.0062, 18],
'GRT': ['0xc944e90c64b2c07662a292be6244bdf05cda44a7', 'investment', 0.00025, 18],
'MKR': ['0x9f8f72aa9304c8b593d555f12ef6589cc3a579a2', 'investment', 0.64, 18],
'CRV': ['0xd533a949740bb3306d119cc777fa900ba034cd52', 'investment', 0.00091, 18],
'LPT': ['0x58b6a8a3302369daec383334672404ee733ab239', 'investment', 0.0075, 18],
'ANT': ['0xa117000000f279d81a1d3cc75430faa017fa5a2e', 'investment', 0.00114, 18],
'GEN': ['0x543ff227f64aa17ea132bf9886cab5db55dcaddf', 'investment', 0.0000137, 18],
'PNK': ['0x93ed3fbe21207ec2e8f2d3c3de6e058cb73bc04d', 'investment', 0.00003, 18],
'REP': ['0x221657776846890989a759ba2973e427dff5c9bb', 'investment', 0.0055, 18],
'SNT': ['0x744d70fdbe2ba4cf95131626614a1763df805b9e', 'investment', 0.000021, 18],
'2KEY': ['0xE48972fCd82a274411c01834e2f031D4377Fa2c0', 'investment', 0.0000036, 18],
'NU': ['0x4fE83213D56308330EC302a8BD641f1d0113A4Cc', 'investment', 0.0002, 18],
'MANA': ['0x0f5d2fb29fb7d3cfee444a200298f468908cc942', 'investment', 0.00056, 18],
'TORN': ['0x77777feddddffc19ff86db637967013e6c6a116c', 'investment', 0.013, 18],
'XRT': ['0x7de91b204c1c737bcee6f000aaa6569cf7061cb7', 'investment', 0.0043, 9],
'FOAM': ['0x4946fcea7c692606e8908002e55a582af44ac121', 'cyberpunk', 0.000015, 18],
'GOLD': ['0x150b0b96933b75ce27af8b92441f8fb683bf9739', 'cyberpunk', 0.000007, 18],
'Tether': ['0xdac17f958d2ee523a2206206994597c13d831ec7', 'stablecoin', 0.00022, 6],
'USD Coin': ['0xa0b86991c6218b36c1d19d4a2e9eb0ce3606eb48', 'stablecoin', 0.00022, 6],
'Binance USD': ['0x4fabb145d64652a948d72533023f6e7a623c7c53', 'stablecoin', 0.00022, 18],
'DAI': ['0x6b175474e89094c44da98b954eedeac495271d0f', 'stablecoin', 0.00022, 18],
'Fei Protocol': ['0x956f47f50a910163d8bf957cf5846d573e7f87ca', 'stablecoin', 0.00022, 18],
'TerraUSD': ['0xa47c8bf37f92abed4a126bda807a7b7498661acd', 'stablecoin', 0.00022, 18],
'Liquity USD': ['0x5f98805a4e8be255a32880fdec7f6728c6568ba0', 'stablecoin', 0.00022, 18],
'Paxos Standard': ['0x8e870d67f660d95d5be530380d0ec0bd388289e1', 'stablecoin', 0.00022, 18],
'TrueUSD': ['0x0000000000085d4780b73119b644ae5ecd22b376', 'stablecoin', 0.00022, 18],
'HUSD': ['0xdf574c24545e5ffecb9a659c229253d4111d87e1', 'stablecoin', 0.00022, 8],
'Neutrino USD': ['0x674c6ad92fd080e4004b2312b45f796a192d27a0', 'stablecoin', 0.00022, 18],
'sUSD': ['0x57ab1ec28d129707052df4df418d58a2d46d5f51', 'stablecoin', 0.00022, 18]
}
erc20_tokens_dict = {k: [v[0].lower(), v[1], v[2], v[3]] for k, v in erc20_tokens_dict.items()}
erc20_tokens_inverse_dict = {v[0].lower():k for k, v in erc20_tokens_dict.items()}
erc20_tokens_df = pd.DataFrame(erc20_tokens_dict.values(),
index=erc20_tokens_dict.keys(),
columns=['address', 'type', 'eth_price', 'decimals']).reset_index()
MINT_EVENT_HASH = '0x0f6798a560793a54c3bcfe86a93cde1e73087d944c0ea20544137d4121396885'
BURN_EVENT_HASH = '0xcc16f5dbb4873280815c1ee09dbd06736cffcc184412cf7a71a0fdb75d397ca5'
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Set Token Data
# + pycharm={"name": "#%%\n"}
investors_erc20_tokens_addresses_tuple = tuple(erc20_tokens_df[erc20_tokens_df.type=='investment'].address.values.tolist())
cyberpunks_erc20_tokens_addresses_tuple = tuple(erc20_tokens_df[erc20_tokens_df.type=='cyberpunk'].address.values.tolist())
stablecoins_erc20_tokens_addresses_tuple = tuple(erc20_tokens_df[erc20_tokens_df.type=='stablecoin'].address.values.tolist())
erc20_tokens_addresses_tuple = investors_erc20_tokens_addresses_tuple + \
cyberpunks_erc20_tokens_addresses_tuple + \
stablecoins_erc20_tokens_addresses_tuple
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Create Token Metadata Table
# + pycharm={"name": "#%%\n"}
if CREATE_TABLES:
r = create_table_from_df(source_df=erc20_tokens_df,
table_name=ERC20_TOKEN_METADATA_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME,
drop_existing_table=DROP_TABLES)
elif DROP_TABLES:
drop_table(table_name=ERC20_ANALYSIS_DISTRIBUTION_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Get Balances
# + pycharm={"name": "#%%\n"}
query_1 = f'''
WITH incoming_transfers AS (
SELECT
transaction_hash,
token_address,
to_address as address,
safe_cast(value as float64) as value
FROM `bigquery-public-data.crypto_ethereum.token_transfers`
WHERE token_address IN {erc20_tokens_addresses_tuple}
AND to_address NOT IN ('0x0000000000000000000000000000000000000000')
AND block_timestamp >= '{INITIAL_TS}'
AND block_timestamp <= '{SNAPSHOT_TS}'
),
outgoing_transfers AS (
SELECT
transaction_hash,
token_address,
from_address as address,
- safe_cast(value as float64) as value
FROM `bigquery-public-data.crypto_ethereum.token_transfers`
WHERE token_address IN {erc20_tokens_addresses_tuple}
AND from_address NOT IN ('0x0000000000000000000000000000000000000000')
AND block_timestamp >= '{INITIAL_TS}'
AND block_timestamp <= '{SNAPSHOT_TS}'
),
mint_burn AS (
SELECT
transaction_hash,
address as token_address,
REPLACE(topics[ORDINAL(2)], '0x000000000000000000000000', '0x') as address,
IF(topics[ORDINAL(1)] = '{MINT_EVENT_HASH}', 'mint', 'burn') as event_name,
safe_cast(data AS float64) as value
FROM `bigquery-public-data.crypto_ethereum.logs`
WHERE block_timestamp >= '{INITIAL_TS}'
AND block_timestamp <= '{SNAPSHOT_TS}'
AND address IN {erc20_tokens_addresses_tuple}
AND topics[ORDINAL(1)] IN ('{MINT_EVENT_HASH}', '{BURN_EVENT_HASH}')
),
token_transfers AS (
SELECT
token_address,
address,
sum(value) as balance
FROM (
SELECT
token_address,
address,
IF((mint_value > COALESCE(transfer_value, 0) * 1.01) OR (mint_value < COALESCE(transfer_value, 0) * 0.99), mint_value, 0) as value
FROM (
SELECT
transaction_hash,
token_address,
address,
sum(value) as mint_value
FROM mint_burn
WHERE event_name = 'mint'
GROUP BY transaction_hash, token_address, address
)
LEFT JOIN (
SELECT
transaction_hash,
token_address,
address,
sum(value) as transfer_value
FROM incoming_transfers
GROUP BY transaction_hash, token_address, address
)
USING (transaction_hash, token_address, address)
UNION ALL
SELECT
token_address,
address,
value
FROM incoming_transfers
UNION ALL
SELECT
token_address,
address,
value
FROM outgoing_transfers
UNION ALL
SELECT
token_address,
address,
IF((burn_value < COALESCE(transfer_value, 0)*1.01) OR (burn_value > COALESCE(transfer_value, 0)*0.99), burn_value, 0) as value
FROM (
SELECT
transaction_hash,
token_address,
address,
- sum(value) as burn_value
FROM mint_burn
WHERE event_name = 'burn'
GROUP BY transaction_hash, token_address, address
)
LEFT JOIN (
SELECT
transaction_hash,
token_address,
address,
sum(value) as transfer_value
FROM outgoing_transfers
GROUP BY transaction_hash, token_address, address
)
USING (transaction_hash, token_address, address)
)
GROUP BY token_address, address
)
SELECT
token_address,
token_transfers.address as address,
balance,
token_metadata.eth_price as eth_price,
token_metadata.decimals as decimals,
balance * token_metadata.eth_price / pow(10, token_metadata.decimals) as eth_balance
FROM token_transfers
LEFT JOIN `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_TOKEN_METADATA_TABLE_NAME}` as token_metadata
ON token_transfers.token_address = token_metadata.address
'''
if DROP_TABLES:
drop_table(table_name=ERC20_ROW_BALANCES_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
if CREATE_TABLES:
create_table(query=query_1,
table_name=ERC20_ROW_BALANCES_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Check Balances
# + pycharm={"name": "#%%\n"}
query_2 = f'''
SELECT
token_address,
SUM(IF(balance > 0, eth_balance, 0)) as eth_positive_balances,
SUM(IF(balance < 0, - eth_balance, 0)) as eth_negative_balances,
COUNTIF(balance > 0) as number_positive_balances,
COUNTIF(balance = 0) as number_zero_balances,
COUNTIF(balance < 0) as number_negative_balances
FROM `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_ROW_BALANCES_TABLE_NAME}`
GROUP BY token_address
HAVING eth_negative_balances > 0.01
AND number_negative_balances > 5
ORDER BY number_negative_balances DESC
'''
negative_balances_df = get_df(query_2)
print(f'Total addresses with negative balance {sum(negative_balances_df.number_negative_balances):>,}')
print(f'Total addresses with this tokens {sum(negative_balances_df.number_negative_balances) + sum(negative_balances_df.number_positive_balances):>,}')
negative_balances_df
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Revise Balances
# + pycharm={"name": "#%%\n"}
if REVISE_TOKEN_BALANCES:
revise_token_list = tuple(negative_balances_df['token_address'].values)
revised_balances_df = pd.DataFrame(columns=('address', 'token_address', 'balance'))
for revise_token in tqdm_notebook(revise_token_list):
print(erc20_tokens_inverse_dict[revise_token], revise_token)
balances_df = get_balances(token_address=revise_token, threads_number=15)
revised_balances_df = revised_balances_df.append(balances_df)
balances_df.to_csv('data/balances_'+erc20_tokens_inverse_dict[revise_token]+'.csv')
revised_balances_df = revised_balances_df.astype({'balance': 'float'})
if CREATE_TABLES:
create_table_from_df(source_df=revised_balances_df,
table_name=ERC20_REVISED_BALANCES_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME,
drop_existing_table=DROP_TABLES)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Update Balances after Revising
# + pycharm={"name": "#%%\n"}
query_3 = f'''
WITH revised_tokens as (
SELECT DISTINCT
token_address
FROM `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_REVISED_BALANCES_TABLE_NAME}`
),
balances as (
SELECT
address,
token_address,
balance
FROM `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_REVISED_BALANCES_TABLE_NAME}`
UNION ALL
SELECT
address,
token_address,
balance
FROM `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_ROW_BALANCES_TABLE_NAME}`
WHERE token_address NOT IN (SELECT token_address FROM revised_tokens)
OR balance = 0
)
SELECT
token_address,
balances.address as address,
balance,
token_metadata.eth_price as eth_price,
token_metadata.decimals as decimals,
balance * token_metadata.eth_price / pow(10, token_metadata.decimals) as eth_balance
FROM balances
LEFT JOIN `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_TOKEN_METADATA_TABLE_NAME}` as token_metadata
ON balances.token_address = token_metadata.address
'''
if DROP_TABLES:
drop_table(table_name=ERC20_BALANCES_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
if CREATE_TABLES:
create_table(query=query_3,
table_name=ERC20_BALANCES_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Get Stablecoin Lovers
# + pycharm={"name": "#%%\n"}
query_4 = f'''
WITH erc20_balances AS (
SELECT
address,
sum(selected_erc20_balance) as sum_selected_erc20_balance,
sum(stablecoin_balance) as sum_stablecoin_balance
FROM (
SELECT
address,
if(token_address IN {investors_erc20_tokens_addresses_tuple + cyberpunks_erc20_tokens_addresses_tuple}, eth_balance, 0) as selected_erc20_balance,
if(token_address IN {stablecoins_erc20_tokens_addresses_tuple}, eth_balance, 0) as stablecoin_balance
FROM `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_BALANCES_TABLE_NAME}`)
GROUP BY address
HAVING sum_stablecoin_balance > sum_selected_erc20_balance
AND sum_stablecoin_balance > 0.1),
eth_balances AS (
SELECT
address,
eth_balance
FROM `{PROJECT_ID}.{ETH_ANALYSIS_DATASET_NAME}.{ETH_ANALYSIS_DISTRIBUTION_TABLE_NAME}`
)
SELECT
address,
sum_selected_erc20_balance,
sum_stablecoin_balance,
eth_balance
FROM erc20_balances
LEFT JOIN eth_balances
USING (address)
WHERE sum_stablecoin_balance > sum_selected_erc20_balance + eth_balance
'''
if DROP_TABLES:
drop_table(table_name=STABLECOINS_LOVERS_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
if CREATE_TABLES:
create_table(query=query_4,
table_name=STABLECOINS_LOVERS_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Analysis of Grade Boundaries. ERC20 balance
# + pycharm={"name": "#%%\n"}
query_5 = f'''
SELECT
if(round(sum_eth_balance, 1) < 0.1, 0.05, round(sum_eth_balance, 1)) as erc20_balance_class,
count(address) as number_of_addresses
FROM (
SELECT
address,
count(distinct token_address) as number_of_tokens,
sum(eth_balance) as sum_eth_balance
FROM `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_BALANCES_TABLE_NAME}`
WHERE balance > {erc20_balance_initial_boundary}
AND token_address IN {investors_erc20_tokens_addresses_tuple}
GROUP BY address
HAVING number_of_tokens >= {investors_min_number_of_tokens}
AND sum_eth_balance > {erc20_balance_initial_boundary})
GROUP BY erc20_balance_class
'''
distribution_by_erc20_balance_df = get_df(query_5)
# + pycharm={"name": "#%%\n"}
investors_boundary_erc20_balance = \
grade_boundaries_analysis(
distribution_df=distribution_by_erc20_balance_df,
value_column = 'erc20_balance_class',
value_chart_label = 'ERC20 Balance by address, Log10',
value_name = 'ERC20 Balance',
value_transform_func=lambda x: log10(x),
chart_title = 'Distribution of ERC20 Balance',
initial_boundary=erc20_balance_initial_boundary,
max_show_value=10000,
level_line_shift=0)
# -
# ### Distribution Rules. Passionate Investors
#
# <table style="text-align: left">
# <thead style="text-align: center">
# <tr>
# <th rowspan=2></th>
# <th colspan=3>Grade</th>
# </tr>
# <tr>
# <th>1</th>
# <th>2</th>
# <th>3</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td style="text-align: left"> Number of tokens >= 2 and ERC20 balance </td>
# <td> > 0.01 ETH </td>
# <td> > 3 ETH </td>
# <td> > 55 ETH </td>
# </tr>
# </tbody>
# </table>
# ### Distribution Rule. Cyberpunks
#
# <table style="text-align: left">
# <thead style="text-align: center">
# <tr>
# <th rowspan=2></th>
# <th colspan=3>Grade</th>
# </tr>
# <tr>
# <th>1</th>
# <th>2</th>
# <th>3</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td style="text-align: left"> Owners of the Selected tokens </td>
# <td> - </td>
# <td> - </td>
# <td> here </td>
# </tr>
# </tbody>
# </table>
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Create Distribution Table
# + pycharm={"name": "#%%\n"}
query_6 = f'''
SELECT
'{INVESTORS_AUDIENCE}' as audience,
'ERC20 Balance' as segment,
address,
CASE
WHEN sum_eth_balance > {investors_boundary_erc20_balance[2]} THEN 3
WHEN sum_eth_balance > {investors_boundary_erc20_balance[1]} THEN 2
WHEN sum_eth_balance > {investors_boundary_erc20_balance[0]} THEN 1
END
AS grade,
sum_eth_balance,
number_of_tokens
FROM (
SELECT
address,
count(distinct token_address) as number_of_tokens,
sum(eth_balance) as sum_eth_balance
FROM `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_BALANCES_TABLE_NAME}`
WHERE balance > 0
AND token_address IN {investors_erc20_tokens_addresses_tuple}
GROUP BY address)
WHERE number_of_tokens >= {investors_min_number_of_tokens}
UNION ALL
SELECT
'{CYBERPUNKS_AUDIENCE}' as audience,
'Owners of the Selected ERC20 Tokens' as segment,
address,
3 AS grade,
sum_eth_balance,
number_of_tokens
FROM (
SELECT
address,
count(distinct token_address) as number_of_tokens,
sum(eth_balance) as sum_eth_balance
FROM `{PROJECT_ID}.{ERC20_ANALYSIS_DATASET_NAME}.{ERC20_BALANCES_TABLE_NAME}`
WHERE balance > 0
AND token_address IN {cyberpunks_erc20_tokens_addresses_tuple}
GROUP BY address)
'''
if DROP_TABLES:
drop_table(table_name=ERC20_ANALYSIS_DISTRIBUTION_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
if CREATE_TABLES:
create_table(query=query_6,
table_name=ERC20_ANALYSIS_DISTRIBUTION_TABLE_NAME,
dataset_name=ERC20_ANALYSIS_DATASET_NAME)
# + pycharm={"name": "#%%\n"}
|
erc20__investors_and_cyberpunks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Sequence to Sequence 네트워크와 Attention을 이용한 번역
# *************************************************************
# **Author**: `<NAME> <https://github.com/spro/practical-pytorch>`_
# **번역**: `황성수 <https://github.com/adonisues>`_
#
# 이 프로젝트에서 신경망이 불어를 영어로 번역하도록 가르칠 예정입니다.
#
# ::
#
# [KEY: > input, = target, < output]
#
# > il est en train de peindre un tableau .
# = he is painting a picture .
# < he is painting a picture .
#
# > pourquoi ne pas essayer ce vin delicieux ?
# = why not try that delicious wine ?
# < why not try that delicious wine ?
#
# > elle n est pas poete mais romanciere .
# = she is not a poet but a novelist .
# < she not not a poet but a novelist .
#
# > vous etes trop maigre .
# = you re too skinny .
# < you re all alone .
#
# ... 성공율은 변할 수 있습니다.
#
# 하나의 시퀀스를 다른 시퀀스로 바꾸는 두개의 RNN이 함께 동작하는
# `sequence to sequence network <http://arxiv.org/abs/1409.3215>`__ 의 간단하지만 강력한 아이디어가
# 이것(번역)을 가능하게 합니다. 인코더 네트워크는 입력 시퀀스를 벡터로 압축하고,
# 디코더 네트워크는 해당 벡터를 새로운 시퀀스로 펼칩니다.
#
# .. figure:: /_static/img/seq-seq-images/seq2seq.png
# :alt:
#
# 이 모델을 개선하기 위해 `Attention Mechanism <https://arxiv.org/abs/1409.0473>`__ 을
# 사용하면 디코더가 입력 시퀀스의 특정 범위에 초점을 맞출 수 있도록 합니다.
#
# **추천 자료:**
#
# 최소한 Pytorch 를 설치했고, Python을 알고, Tensor 를 이해한다고 가정합니다:
#
# - http://pytorch.org/ 설치 안내
# - :doc:`/beginner/deep_learning_60min_blitz` 전반적인 PyTorch 시작을 위한 자료
# - :doc:`/beginner/pytorch_with_examples` 넓고 깊은 통찰을 위한 자료
# - :doc:`/beginner/former_torchies_tutorial` 이전 Lua Torch 사용자를 위한 자료
#
#
# Sequence to Sequence 네트워크와 동작 방법에 관해서 아는 것은 유용합니다:
#
# - `Learning Phrase Representations using RNN Encoder-Decoder for
# Statistical Machine Translation <http://arxiv.org/abs/1406.1078>`__
# - `Sequence to Sequence Learning with Neural
# Networks <http://arxiv.org/abs/1409.3215>`__
# - `Neural Machine Translation by Jointly Learning to Align and
# Translate <https://arxiv.org/abs/1409.0473>`__
# - `A Neural Conversational Model <http://arxiv.org/abs/1506.05869>`__
#
# 이전 튜토리얼에 있는
# :doc:`/intermediate/char_rnn_classification_tutorial`
# 와 :doc:`/intermediate/char_rnn_generation_tutorial` 는
# 각각 인코더와 디코더 모델과 비슷한 컨센을 가지기 때문에 도움이 됩니다.
#
# 추가로 이 토픽들을 다루는 논문을 읽어 보십시오:
#
# - `Learning Phrase Representations using RNN Encoder-Decoder for
# Statistical Machine Translation <http://arxiv.org/abs/1406.1078>`__
# - `Sequence to Sequence Learning with Neural
# Networks <http://arxiv.org/abs/1409.3215>`__
# - `Neural Machine Translation by Jointly Learning to Align and
# Translate <https://arxiv.org/abs/1409.0473>`__
# - `A Neural Conversational Model <http://arxiv.org/abs/1506.05869>`__
#
#
# **요구 사항**
#
#
# +
from __future__ import unicode_literals, print_function, division
from io import open
import unicodedata
import string
import re
import random
import torch
import torch.nn as nn
from torch import optim
import torch.nn.functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# -
# 데이터 파일 로딩
# ==================
#
# 이 프로젝트의 데이터는 수천 개의 영어-프랑스어 번역 쌍입니다.
#
# `Open Data Stack Exchange <http://opendata.stackexchange.com/questions/3888/dataset-of-sentences-translated-into-many-languages>`__
# 에 관한 이 문의는 http://tatoeba.org/eng/downloads 에서 다운 로드가 가능한
# 공개 번역 사이트 http://tatoeba.org/ 를 알려 주었습니다. 더 나은 방법으로
# 언어 쌍을 개별 텍스트 파일로 분할하는 추가 작업을 수행한
# http://www.manythings.org/anki/ 가 있습니다:
#
# 영어-프랑스어 쌍이 너무 커서 저장소에 포함 할 수 없기 때문에
# 계속하기 전에 ``data/eng-fra.txt`` 로 다운로드하십시오.
# 이 파일은 탭으로 구분된 번역 쌍 목록입니다:
#
# ::
#
# I am cold. J'ai froid.
#
# .. Note::
# `여기 <https://download.pytorch.org/tutorial/data.zip>`_
# 에서 데이터를 다운 받고 현재 디렉토리에 압축을 푸십시오.
#
#
# 문자 단위 RNN 튜토리얼에서 사용된 문자 인코딩과 유사하게, 언어의 각
# 단어들은 One-Hot 벡터 또는 그 단어의 주소에만 단 하나의 1을 제외하고
# 모두 0인 큰 벡터로 표현합니다. 한 가지 언어에 있는 수십 개의 문자와
# 달리 번역에는 아주 많은 단어들이 있기 때문에 인코딩 벡터는 매우 더 큽니다.
# 그러나 우리는 약간의 속임수를 써서 언어 당 수천 단어 만
# 사용하도록 데이터를 다듬을 것입니다.
#
# .. figure:: /_static/img/seq-seq-images/word-encoding.png
# :alt:
#
#
#
#
# 나중에 네트워크의 입력 및 목표로 사용하려면 단어 당 고유 번호가
# 필요합니다. 이 모든 것을 추적하기 위해 우리는
# 단어→색인(``word2index``)과 색인→단어(``index2word``) 사전,
# 그리고 나중에 희귀 단어를 대체하는데 사용할 각 단어의 빈도
# ``word2count`` 를 가진 ``Lang`` 이라는 헬퍼 클래스를 사용합니다.
#
#
#
# +
SOS_token = 0
EOS_token = 1
class Lang:
def __init__(self, name):
self.name = name
self.word2index = {}
self.word2count = {}
self.index2word = {0: "SOS", 1: "EOS"}
self.n_words = 2 # SOS 와 EOS 단어 숫자 포함
def addSentence(self, sentence):
for word in sentence.split(' '):
self.addWord(word)
def addWord(self, word):
if word not in self.word2index:
self.word2index[word] = self.n_words
self.word2count[word] = 1
self.index2word[self.n_words] = word
self.n_words += 1
else:
self.word2count[word] += 1
# -
# 파일은 모두 유니 코드로 되어있어 간단하게하기 위해 유니 코드 문자를
# ASCII로 변환하고, 모든 문자를 소문자로 만들고, 대부분의 구두점을
# 지워줍니다.
#
#
#
# +
# 유니 코드 문자열을 일반 ASCII로 변환하십시오.
# http://stackoverflow.com/a/518232/2809427 에 감사드립니다.
def unicodeToAscii(s):
return ''.join(
c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn'
)
# 소문자, 다듬기, 그리고 문자가 아닌 문자 제거
def normalizeString(s):
s = unicodeToAscii(s.lower().strip())
s = re.sub(r"([.!?])", r" \1", s)
s = re.sub(r"[^a-zA-Z.!?]+", r" ", s)
return s
# -
# 데이터 파일을 읽으려면 파일을 줄로 나눈 다음 줄을 쌍으로 나눕니다.
# 파일은 모두 영어 → 기타 언어이므로 만약 다른 언어 → 영어로
# 번역한다면 쌍을 뒤집을 수 있도록 ``reverse`` 플래그를 추가했습니다.
#
#
#
def readLangs(lang1, lang2, reverse=False):
print("Reading lines...")
# 파일을 읽고 줄로 분리
lines = open('data/%s-%s.txt' % (lang1, lang2), encoding='utf-8').\
read().strip().split('\n')
# 모든 줄을 쌍으로 분리하고 정규화
pairs = [[normalizeString(s) for s in l.split('\t')] for l in lines]
# 쌍을 뒤집고, Lang 인스턴스 생성
if reverse:
pairs = [list(reversed(p)) for p in pairs]
input_lang = Lang(lang2)
output_lang = Lang(lang1)
else:
input_lang = Lang(lang1)
output_lang = Lang(lang2)
return input_lang, output_lang, pairs
# *많은* 예제 문장이 있고 신속하게 학습하기를 원하기 때문에
# 비교적 짧고 간단한 문장으로만 데이터 셋을 정리할 것입니다. 여기서
# 최대 길이는 10 단어 (종료 문장 부호 포함)이며 "I am" 또는
# "He is" 등의 형태로 번역되는 문장으로 필터링됩니다.(이전에
# 아포스트로피는 대체 됨)
#
#
#
# +
MAX_LENGTH = 10
eng_prefixes = (
"i am ", "i m ",
"he is", "he s ",
"she is", "she s",
"you are", "you re ",
"we are", "we re ",
"they are", "they re "
)
def filterPair(p):
return len(p[0].split(' ')) < MAX_LENGTH and \
len(p[1].split(' ')) < MAX_LENGTH and \
p[1].startswith(eng_prefixes)
def filterPairs(pairs):
return [pair for pair in pairs if filterPair(pair)]
# -
# 데이터 준비를 위한 전체 과정:
#
# - 텍스트 파일을 읽고 줄로 분리하고, 줄을 쌍으로 분리합니다.
# - 텍스트를 정규화 하고 길이와 내용으로 필터링 합니다.
# - 쌍의 문장들에서 단어 리스트를 생성합니다.
#
#
#
# +
def prepareData(lang1, lang2, reverse=False):
input_lang, output_lang, pairs = readLangs(lang1, lang2, reverse)
print("Read %s sentence pairs" % len(pairs))
pairs = filterPairs(pairs)
print("Trimmed to %s sentence pairs" % len(pairs))
print("Counting words...")
for pair in pairs:
input_lang.addSentence(pair[0])
output_lang.addSentence(pair[1])
print("Counted words:")
print(input_lang.name, input_lang.n_words)
print(output_lang.name, output_lang.n_words)
return input_lang, output_lang, pairs
input_lang, output_lang, pairs = prepareData('eng', 'fra', True)
print(random.choice(pairs))
# -
# Seq2Seq 모델
# =================
#
# Recurrent Neural Network(RNN)는 시퀀스에서 작동하고 후속 단계의
# 입력으로 자신의 출력을 사용하는 네트워크입니다.
#
# `Sequence to Sequence network <http://arxiv.org/abs/1409.3215>`__, 또는
# Seq2Seq 네트워크, 또는 `Encoder Decoder
# network <https://arxiv.org/pdf/1406.1078v3.pdf>`__ 는 인코더 및
# 디코더라고 하는 두 개의 RNN으로 구성된 모델입니다.
# 인코더는 입력 시퀀스를 읽고 단일 벡터를 출력하고,
# 디코더는 해당 벡터를 읽어 출력 시퀀스를 생성합니다.
#
# .. figure:: /_static/img/seq-seq-images/seq2seq.png
# :alt:
#
# 모든 입력에 해당하는 출력이 있는 단일 RNN의 시퀀스 예측과 달리
# Seq2Seq 모델은 시퀀스 길이와 순서를 자유롭게하기 때문에
# 두 언어 사이의 번역에 이상적입니다.
#
# 다음 문장 "Je ne suis pas le chat noir" → "I am not the black cat"
# 를 살펴 봅시다. 입력 문장의 단어 대부분은 출력 문장에서
# 직역("chat noir" 와 "black cat")되지만 약간 다른 순서도 있습니다.
# "ne/pas" 구조로 인해 입력 문장에 단어가 하나 더 있습니다.
# 입력 단어의 시퀀스에서 직접적으로는 정확한 번역을 만드는
# 것은 어려울 것입니다.
#
# Seq2Seq 모델을 사용하면 인코더는 하나의 벡터를 생성합니다.
# 이상적인 경우에 입력 시퀀스의 "의미"를 문장의 N 차원 공간에 있는
# 단일 지점인 단일 벡터으로 인코딩합니다.
#
#
#
# 인코더
# -----------
#
# Seq2Seq 네트워크의 인코더는 입력 문장의 모든 단어에 대해 어떤 값을
# 출력하는 RNN입니다. 모든 입력 단어에 대해 인코더는 벡터와
# 은닉 상태를 출력하고 다음 입력 단어에 그 은닉 상태를 사용합니다.
#
# .. figure:: /_static/img/seq-seq-images/encoder-network.png
# :alt:
#
#
#
#
class EncoderRNN(nn.Module):
def __init__(self, input_size, hidden_size):
super(EncoderRNN, self).__init__()
self.hidden_size = hidden_size
self.embedding = nn.Embedding(input_size, hidden_size)
self.gru = nn.GRU(hidden_size, hidden_size)
def forward(self, input, hidden):
embedded = self.embedding(input).view(1, 1, -1)
output = embedded
output, hidden = self.gru(output, hidden)
return output, hidden
def initHidden(self):
return torch.zeros(1, 1, self.hidden_size, device=device)
# 디코더
# -----------
#
# 디코더는 인코더 출력 벡터를 받아서 번역을 생성하는 단어 시퀀스를
# 출력합니다.
#
#
#
# 간단한 디코더
# ^^^^^^^^^^^^^^
#
# 가장 간단한 Seq2Seq 디코더에서 인코더의 마지막 출력만을 이용합니다.
# 이 마지막 출력은 전체 시퀀스에서 문맥을 인코드하기 때문에
# *문맥 벡터(context vector)* 로 불립니다. 이 문맥 벡터는 디코더의 초기 은닉 상태로
# 사용 됩니다.
#
# 디코딩의 매 단계에서 디코더에게 입력 토큰과 은닉 상태가 주어집니다.
# 초기 입력 토큰은 문자열-시작 (start-of-string) ``<SOS>`` 토큰이고,
# 첫 은닉 상태는 문맥 벡터(인코더의 마지막 은닉 상태) 입니다.
#
# .. figure:: /_static/img/seq-seq-images/decoder-network.png
# :alt:
#
#
#
#
class DecoderRNN(nn.Module):
def __init__(self, hidden_size, output_size):
super(DecoderRNN, self).__init__()
self.hidden_size = hidden_size
self.embedding = nn.Embedding(output_size, hidden_size)
self.gru = nn.GRU(hidden_size, hidden_size)
self.out = nn.Linear(hidden_size, output_size)
self.softmax = nn.LogSoftmax(dim=1)
def forward(self, input, hidden):
output = self.embedding(input).view(1, 1, -1)
output = F.relu(output)
output, hidden = self.gru(output, hidden)
output = self.softmax(self.out(output[0]))
return output, hidden
def initHidden(self):
return torch.zeros(1, 1, self.hidden_size, device=device)
# 이 모델의 결과를 학습하고 관찰하는 것을 권장하지만,
# 공간을 절약하기 위해 최종 목적지로 바로 이동해서
# 어텐션(attention) 메카니즘을 소개 할 것입니다.
#
#
#
# 어텐션 디코더
# ^^^^^^^^^^^^^^^^^
#
# 문맥 벡터만 인코더와 디코더 사이로 전달 된다면, 단일 벡터가 전체 문장을
# 인코딩 해야하는 부담을 가지게 됩니다.
# 어텐션은 디코더 네트워크가 자기 출력의 모든 단계에서 인코더 출력의
# 다른 부분에 "집중" 할 수 있게 합니다. 첫째 *어텐션 웨이트* 의 세트를
# 계산합니다. 이것은 가중치 조합을 만들기 위해서 인코더 출력 벡터와
# 곱해집니다. 그 결과(코드에서 ``attn_applied``)는 입력 시퀀스의
# 특정 부분에 관한 정보를 포함해야하고 따라서 디코더가 알맞은 출력
# 단어를 선택하는 것을 도와줍니다.
#
# .. figure:: https://i.imgur.com/1152PYf.png
# :alt:
#
# 어텐션 가중치 계산은 디코더의 입력 및 은닉 상태를 입력으로
# 사용하는 다른 feed-forwad layer 인 ``attn`` 으로 수행됩니다.
# 학습 데이터에는 모든 크기의 문장이 있기 때문에 이 계층을 실제로
# 만들고 학습시키려면 적용 할 수 있는 최대 문장 길이 (인코더 출력을 위한 입력 길이)를
# 선택해야 합니다. 최대 길이의 문장은 모든 어텐션 가중치를 사용하지만
# 더 짧은 문장은 처음 몇 개만 사용합니다.
#
# .. figure:: /_static/img/seq-seq-images/attention-decoder-network.png
# :alt:
#
#
#
#
class AttnDecoderRNN(nn.Module):
def __init__(self, hidden_size, output_size, dropout_p=0.1, max_length=MAX_LENGTH):
super(AttnDecoderRNN, self).__init__()
self.hidden_size = hidden_size
self.output_size = output_size
self.dropout_p = dropout_p
self.max_length = max_length
self.embedding = nn.Embedding(self.output_size, self.hidden_size)
self.attn = nn.Linear(self.hidden_size * 2, self.max_length)
self.attn_combine = nn.Linear(self.hidden_size * 2, self.hidden_size)
self.dropout = nn.Dropout(self.dropout_p)
self.gru = nn.GRU(self.hidden_size, self.hidden_size)
self.out = nn.Linear(self.hidden_size, self.output_size)
def forward(self, input, hidden, encoder_outputs):
embedded = self.embedding(input).view(1, 1, -1)
embedded = self.dropout(embedded)
attn_weights = F.softmax(
self.attn(torch.cat((embedded[0], hidden[0]), 1)), dim=1)
attn_applied = torch.bmm(attn_weights.unsqueeze(0),
encoder_outputs.unsqueeze(0))
output = torch.cat((embedded[0], attn_applied[0]), 1)
output = self.attn_combine(output).unsqueeze(0)
output = F.relu(output)
output, hidden = self.gru(output, hidden)
output = F.log_softmax(self.out(output[0]), dim=1)
return output, hidden, attn_weights
def initHidden(self):
return torch.zeros(1, 1, self.hidden_size, device=device)
# <div class="alert alert-info"><h4>Note</h4><p>상대 위치 접근을 이용한 길이 제한을 하는 다른 형태의 어텐션
# 이 있습니다. "local attention"에 관한 자료
# `Effective Approaches to Attention-based Neural Machine Translation <https://arxiv.org/abs/1508.04025>`__
# 를 읽으십시오</p></div>
#
# 학습
# ========
#
# 학습 데이터 준비
# -----------------------
#
# 학습을 위해서, 각 쌍마다 입력 Tensor(입력 문장의 단어 주소)와
# 목표 Tensor(목표 문장의 단어 주소)가 필요합니다. 이 벡터들을
# 생성하는 동안 두 시퀀스에 EOS 토큰을 추가 합니다.
#
#
#
# +
def indexesFromSentence(lang, sentence):
return [lang.word2index[word] for word in sentence.split(' ')]
def tensorFromSentence(lang, sentence):
indexes = indexesFromSentence(lang, sentence)
indexes.append(EOS_token)
return torch.tensor(indexes, dtype=torch.long, device=device).view(-1, 1)
def tensorsFromPair(pair):
input_tensor = tensorFromSentence(input_lang, pair[0])
target_tensor = tensorFromSentence(output_lang, pair[1])
return (input_tensor, target_tensor)
# -
# 모델 학습
# ------------------
#
# 학습을 위해서 인코더에 입력 문장을 넣고 모든 출력과 최신 은닉 상태를
# 추적합니다. 그런 다음 디코더에 첫 번째 입력으로 ``<SOS>`` 토큰과
# 인코더의 마지막 은닉 상태가 첫번쩨 은닉 상태로 제공됩니다.
#
# "Teacher forcing"은 다음 입력으로 디코더의 예측을 사용하는 대신
# 실제 목표 출력을 다음 입력으로 사용하는 컨셉입니다.
# "Teacher forcing"을 사용하면 수렴이 빨리되지만 `학습된 네트워크가
# 잘못 사용될 때 불안정성을 보입니다
# <http://minds.jacobs-university.de/sites/default/files/uploads/papers/ESNTutorialRev.pdf>`__
#
# Teacher-forced 네트워크의 출력이 일관된 문법으로 읽지만 정확한
# 번역과는 거리가 멀다는 것을 볼 수 있습니다. 직관적으로 출력 문법을
# 표현하는 법을 배우고 교사가 처음 몇 단어를 말하면 의미를 "선택" 할 수 있지만,
# 번역에서 처음으로 문장을 만드는 법은 잘 배우지 못합니다.
#
# PyTorch의 autograd 가 제공하는 자유 덕분에 간단한 If 문으로
# Teacher Forcing을 사용할지 아니면 사용하지 않을지를 선택할 수 있습니다.
# 더 많이 사용하려면 ``teacher_forcing_ratio`` 를 확인하십시오.
#
#
#
# +
teacher_forcing_ratio = 0.5
def train(input_tensor, target_tensor, encoder, decoder, encoder_optimizer, decoder_optimizer, criterion, max_length=MAX_LENGTH):
encoder_hidden = encoder.initHidden()
encoder_optimizer.zero_grad()
decoder_optimizer.zero_grad()
input_length = input_tensor.size(0)
target_length = target_tensor.size(0)
encoder_outputs = torch.zeros(max_length, encoder.hidden_size, device=device)
loss = 0
for ei in range(input_length):
encoder_output, encoder_hidden = encoder(
input_tensor[ei], encoder_hidden)
encoder_outputs[ei] = encoder_output[0, 0]
decoder_input = torch.tensor([[SOS_token]], device=device)
decoder_hidden = encoder_hidden
use_teacher_forcing = True if random.random() < teacher_forcing_ratio else False
if use_teacher_forcing:
# Teacher forcing 포함: 목표를 다음 입력으로 전달
for di in range(target_length):
decoder_output, decoder_hidden, decoder_attention = decoder(
decoder_input, decoder_hidden, encoder_outputs)
loss += criterion(decoder_output, target_tensor[di])
decoder_input = target_tensor[di] # Teacher forcing
else:
# Teacher forcing 미포함: 자신의 예측을 다음 입력으로 사용
for di in range(target_length):
decoder_output, decoder_hidden, decoder_attention = decoder(
decoder_input, decoder_hidden, encoder_outputs)
topv, topi = decoder_output.topk(1)
decoder_input = topi.squeeze().detach() # 입력으로 사용할 부분을 히스토리에서 분리
loss += criterion(decoder_output, target_tensor[di])
if decoder_input.item() == EOS_token:
break
loss.backward()
encoder_optimizer.step()
decoder_optimizer.step()
return loss.item() / target_length
# -
# 이것은 현재 시간과 진행률%을 고려해 경과된 시간과 남은 예상
# 시간을 출력하는 헬퍼 함수입니다.
#
#
#
# +
import time
import math
def asMinutes(s):
m = math.floor(s / 60)
s -= m * 60
return '%dm %ds' % (m, s)
def timeSince(since, percent):
now = time.time()
s = now - since
es = s / (percent)
rs = es - s
return '%s (- %s)' % (asMinutes(s), asMinutes(rs))
# -
# 전체 학습 과정은 다음과 같습니다:
#
# - 타이머 시작
# - optimizers와 criterion 초기화
# - 학습 쌍의 세트 생성
# - 도식화를 위한 빈 손실 배열 시작
#
# 그런 다음 우리는 여러 번 ``train`` 을 호출하며 때로는 진행률
# (예제의 %, 현재까지의 예상 시간)과 평균 손실을 출력합니다.
#
#
#
def trainIters(encoder, decoder, n_iters, print_every=1000, plot_every=100, learning_rate=0.01):
start = time.time()
plot_losses = []
print_loss_total = 0 # 매 print_every 마다 초기화
plot_loss_total = 0 # 매 plot_every 마다 초기화
encoder_optimizer = optim.SGD(encoder.parameters(), lr=learning_rate)
decoder_optimizer = optim.SGD(decoder.parameters(), lr=learning_rate)
training_pairs = [tensorsFromPair(random.choice(pairs))
for i in range(n_iters)]
criterion = nn.NLLLoss()
for iter in range(1, n_iters + 1):
training_pair = training_pairs[iter - 1]
input_tensor = training_pair[0]
target_tensor = training_pair[1]
loss = train(input_tensor, target_tensor, encoder,
decoder, encoder_optimizer, decoder_optimizer, criterion)
print_loss_total += loss
plot_loss_total += loss
if iter % print_every == 0:
print_loss_avg = print_loss_total / print_every
print_loss_total = 0
print('%s (%d %d%%) %.4f' % (timeSince(start, iter / n_iters),
iter, iter / n_iters * 100, print_loss_avg))
if iter % plot_every == 0:
plot_loss_avg = plot_loss_total / plot_every
plot_losses.append(plot_loss_avg)
plot_loss_total = 0
showPlot(plot_losses)
# 결과 도식화
# ----------------
#
# matplotlib로 학습 중에 저장된 손실 값 ``plot_losses`` 의 배열을
# 사용하여 도식화합니다.
#
#
#
# +
import matplotlib.pyplot as plt
plt.switch_backend('agg')
import matplotlib.ticker as ticker
import numpy as np
def showPlot(points):
plt.figure()
fig, ax = plt.subplots()
# 주기적인 간격에 이 locator가 tick을 설정
loc = ticker.MultipleLocator(base=0.2)
ax.yaxis.set_major_locator(loc)
plt.plot(points)
# -
# 평가
# ==========
#
# 평가는 대부분 학습과 동일하지만 목표가 없으므로 각 단계마다 디코더의
# 예측을 되돌려 전달합니다.
# 단어를 예측할 때마다 그 단어를 출력 문자열에 추가합니다.
# 만약 EOS 토큰을 예측하면 거기에서 멈춥니다.
# 나중에 도식화를 위해서 디코더의 어텐션 출력을 저장합니다.
#
#
#
def evaluate(encoder, decoder, sentence, max_length=MAX_LENGTH):
with torch.no_grad():
input_tensor = tensorFromSentence(input_lang, sentence)
input_length = input_tensor.size()[0]
encoder_hidden = encoder.initHidden()
encoder_outputs = torch.zeros(max_length, encoder.hidden_size, device=device)
for ei in range(input_length):
encoder_output, encoder_hidden = encoder(input_tensor[ei],
encoder_hidden)
encoder_outputs[ei] += encoder_output[0, 0]
decoder_input = torch.tensor([[SOS_token]], device=device) # SOS
decoder_hidden = encoder_hidden
decoded_words = []
decoder_attentions = torch.zeros(max_length, max_length)
for di in range(max_length):
decoder_output, decoder_hidden, decoder_attention = decoder(
decoder_input, decoder_hidden, encoder_outputs)
decoder_attentions[di] = decoder_attention.data
topv, topi = decoder_output.data.topk(1)
if topi.item() == EOS_token:
decoded_words.append('<EOS>')
break
else:
decoded_words.append(output_lang.index2word[topi.item()])
decoder_input = topi.squeeze().detach()
return decoded_words, decoder_attentions[:di + 1]
# 학습 세트에 있는 임의의 문장을 평가하고
# 입력, 목표 및 출력을 출력하여 주관적인 품질 판단을 내릴 수 있습니다:
#
#
#
def evaluateRandomly(encoder, decoder, n=10):
for i in range(n):
pair = random.choice(pairs)
print('>', pair[0])
print('=', pair[1])
output_words, attentions = evaluate(encoder, decoder, pair[0])
output_sentence = ' '.join(output_words)
print('<', output_sentence)
print('')
# 학습과 평가
# =======================
#
# 이러한 모든 헬퍼 함수를 이용해서 (추가 작업처럼 보이지만 여러 실험을
# 더 쉽게 수행 할 수 있음) 실제로 네트워크를 초기화하고 학습을
# 시작할 수 있습니다.
#
# 입력 문장이 많이 필터링되었음을 기억하십시오. 이 작은 데이터 세트의
# 경우 256 크기의 은닉 노드(hidden node)와 단일 GRU 계층 같은 상대적으로 작은
# 네트워크를 사용할 수 있습니다. MacBook CPU에서 약 40분 후에
# 합리적인 결과를 얻을 것입니다.
#
# .. Note::
# 이 노트북을 실행하면 학습, 커널 중단, 평가를 할 수 있고 나중에
# 이어서 학습을 할 수 있습니다. 인코더와 디코더가 초기화 된 행을
# 주석 처리하고 ``trainIters`` 를 다시 실행하십시오.
#
#
#
# +
hidden_size = 256
encoder1 = EncoderRNN(input_lang.n_words, hidden_size).to(device)
attn_decoder1 = AttnDecoderRNN(hidden_size, output_lang.n_words, dropout_p=0.1).to(device)
trainIters(encoder1, attn_decoder1, 75000, print_every=5000)
# -
evaluateRandomly(encoder1, attn_decoder1)
# 어텐션 시각화
# ---------------------
#
# 어텐션 메커니즘의 유용한 속성은 하나는 해석 가능성이 높은 출력입니다.
# 입력 시퀀스의 특정 인코더 출력에 가중치를 부여하는 데 사용되므로
# 각 시간 단계에서 네트워크가 가장 집중되는 위치를 파악할 수 있습니다.
#
# 어텐션 출력을 행렬로 표시하기 위해 ``plt.matshow(attentions)`` 를
# 간단하게 실행할 수 있습니다. 열은 입력 단계와 행이 출력 단계입니다:
#
#
#
output_words, attentions = evaluate(
encoder1, attn_decoder1, "je suis trop froid .")
plt.matshow(attentions.numpy())
# 더 나은 보기를 위해 축과 라벨을 더하는 추가 작업을 수행합니다:
#
#
#
# +
def showAttention(input_sentence, output_words, attentions):
# colorbar로 그림 설정
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(attentions.numpy(), cmap='bone')
fig.colorbar(cax)
# 축 설정
ax.set_xticklabels([''] + input_sentence.split(' ') +
['<EOS>'], rotation=90)
ax.set_yticklabels([''] + output_words)
# 매 틱마다 라벨 보여주기
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
def evaluateAndShowAttention(input_sentence):
output_words, attentions = evaluate(
encoder1, attn_decoder1, input_sentence)
print('input =', input_sentence)
print('output =', ' '.join(output_words))
showAttention(input_sentence, output_words, attentions)
evaluateAndShowAttention("elle a cinq ans de moins que moi .")
evaluateAndShowAttention("elle est trop petit .")
evaluateAndShowAttention("je ne crains pas de mourir .")
evaluateAndShowAttention("c est un jeune directeur plein de talent .")
# -
# 연습
# =========
#
# - 다른 데이터 셋을 시도해 보십시오
#
# - 다른 언어쌍
# - 사람 → 기계 (e.g. IOT 명령어)
# - 채팅 → 응답
# - 질문 → 답변
#
# - word2vec 또는 GloVe 같은 미리 학습된 word embedding 으로
# embedding 을 교체하십시오
#
# - 더 많은 레이어, 은닉 유닛, 더 많은 문장을 사용하십시오.
# 학습 시간과 결과를 비교해 보십시오
# - 만약 같은 구문 두개의 쌍으로 된 번역 파일을 이용한다면,
# (``I am test \t I am test``), 이것을 오토인코더로
# 사용할 수 있습니다.
# 이것을 시도해 보십시오:
#
# - 오토인코더 학습
# - 인코더 네트워크 저장하기
# - 그 상태에서 번역을 위한 새로운 디코더 학습
#
#
#
|
docs/_downloads/7e2473184677c1b15004ef4807d5ca05/seq2seq_translation_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## EEG Sleep states
# In this tutorial, we just give a practical example how to use NLD properties in biomedical signals properties. In particular, we want to test whether NLD properties are better in the EEG sleep states classification compared to Powerband EEG.
#
# Refer to the following tutorial to know more about the data and how to import them in Python
#
# https://mne.tools/stable/auto_tutorials/clinical/60_sleep.html
#
# We are going to follow closely this tutorial and we are goign to add a part related DFA and Sampel Entropy to test whether these features have better predictive power that EEG powerband.
# ## Import packages
import numpy as np
import scipy as sp
from src import py_msent
import matplotlib.pyplot as plt
import fathon
from fathon import fathonUtils as fu
import math
import os
import numpy as np
from scipy.integrate import solve_ivp
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import nolds
from scipy.signal import welch
# ## Import MNE
# +
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne.datasets.sleep_physionet.age import fetch_data
from mne.time_frequency import psd_welch
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import FunctionTransformer
# -
# ## Polisomongraphy data
# +
ALICE, BOB = 0, 1
[alice_files, bob_files] = fetch_data(subjects=[ALICE, BOB], recording=[1])
mapping = {'EOG horizontal': 'eog',
'Resp oro-nasal': 'resp',
'EMG submental': 'emg',
'Temp rectal': 'misc',
'Event marker': 'misc'}
raw_train = mne.io.read_raw_edf(alice_files[0])
annot_train = mne.read_annotations(alice_files[1])
raw_train.set_annotations(annot_train, emit_warning=False)
raw_train.set_channel_types(mapping)
# plot some data
# scalings were chosen manually to allow for simultaneous visualization of
# different channel types in this specific dataset
raw_train.plot(start=60, duration=60,
scalings=dict(eeg=1e-4, resp=1e3, eog=1e-4, emg=1e-7,
misc=1e-1))
# -
# ## Extract 30s events from annotations
# +
annotation_desc_2_event_id = {'Sleep stage W': 1,
'Sleep stage 1': 2,
'Sleep stage 2': 3,
'Sleep stage 3': 4,
'Sleep stage 4': 4,
'Sleep stage R': 5}
# keep last 30-min wake events before sleep and first 30-min wake events after
# sleep and redefine annotations on raw data
annot_train.crop(annot_train[1]['onset'] - 30 * 60,
annot_train[-2]['onset'] + 30 * 60)
raw_train.set_annotations(annot_train, emit_warning=False)
events_train, _ = mne.events_from_annotations(
raw_train, event_id=annotation_desc_2_event_id, chunk_duration=30.)
# create a new event_id that unifies stages 3 and 4
event_id = {'Sleep stage W': 1,
'Sleep stage 1': 2,
'Sleep stage 2': 3,
'Sleep stage 3/4': 4,
'Sleep stage R': 5}
# plot events
fig = mne.viz.plot_events(events_train, event_id=event_id,
sfreq=raw_train.info['sfreq'],
first_samp=events_train[0, 0])
# keep the color-code for further plotting
stage_colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
# -
# ## Create Epochs from the data based on the events found in the annotations
# +
tmax = 30. - 1. / raw_train.info['sfreq'] # tmax in included
epochs_train = mne.Epochs(raw=raw_train, events=events_train,
event_id=event_id, tmin=0., tmax=tmax, baseline=None)
print(epochs_train)
# -
# ## Applying the same steps to the test data from Bob
# +
raw_test = mne.io.read_raw_edf(bob_files[0])
annot_test = mne.read_annotations(bob_files[1])
annot_test.crop(annot_test[1]['onset'] - 30 * 60,
annot_test[-2]['onset'] + 30 * 60)
raw_test.set_annotations(annot_test, emit_warning=False)
raw_test.set_channel_types(mapping)
events_test, _ = mne.events_from_annotations(
raw_test, event_id=annotation_desc_2_event_id, chunk_duration=30.)
epochs_test = mne.Epochs(raw=raw_test, events=events_test, event_id=event_id,
tmin=0., tmax=tmax, baseline=None)
print(epochs_test)
# -
# ## Feature Engineering
# +
# visualize Alice vs. Bob PSD by sleep stage.
fig, (ax1, ax2) = plt.subplots(ncols=2)
# iterate over the subjects
stages = sorted(event_id.keys())
for ax, title, epochs in zip([ax1, ax2],
['Alice', 'Bob'],
[epochs_train, epochs_test]):
for stage, color in zip(stages, stage_colors):
epochs[stage].plot_psd(area_mode=None, color=color, ax=ax,
fmin=0.1, fmax=20., show=False,
average=True, spatial_colors=False)
ax.set(title=title, xlabel='Frequency (Hz)')
ax2.set(ylabel='µV^2/Hz (dB)')
ax2.legend(ax2.lines[2::3], stages)
plt.tight_layout()
plt.show()
# -
# ## Feature Engineering -- DFA
nMin = 16
nMax = 2**10
nScales = 32
exponents = np.linspace(np.log2(nMin), np.log2(nMax), nScales)
scales = np.round(np.power(2.0, exponents)).astype(np.int64)
q_list = [-5, -3, -1, 1, 3, 5]
def get_idxs(vec, scales):
idxs = []
for s in scales:
idxs.append(np.where(vec==s)[0][0])
return idxs
def test_mat_dfa(x):
x_dfa = fathon.DFA(fu.toAggregated(x))
n_w, F_w = x_dfa.computeFlucVec(scales, revSeg=False, polOrd=1)
idxs = get_idxs(n_w, scales)
n_w = n_w[idxs]
F_w = F_w[idxs]
H, _ = np.polyfit(np.log2(n_w), np.log2(F_w), 1)
return H,n_w,F_w
# +
# visualize Alice vs. Bob DFA by sleep stage.
fig, (ax1, ax2) = plt.subplots(ncols=2)
# iterate over the subjects
stages = sorted(event_id.keys())
for ax, title, epochs in zip([ax1, ax2],
['Alice', 'Bob'],
[epochs_train, epochs_test]):
stg_idx = 0
for stage, color in zip(stages, stage_colors):
x = epochs[stage].get_data()
eeg_x = x[0,0,:]
H,n_w,F_w = test_mat_dfa(eeg_x)
ax.scatter(np.log2(n_w), np.log2(F_w),label=stages[stg_idx])
stg_idx +=1
ax.set(title=title, xlabel='Scales')
ax2.set(ylabel='F(n)')
# ax2.legend(ax2.lines[2::3], stages)
ax2.legend()
plt.tight_layout()
plt.show()
# -
# ## Create NLD classifier and Powerband classifier
#
def eeg_nld(epochs):
"""EEG relative power band feature extraction.
This function takes an ``mne.Epochs`` object and creates EEG features based
on relative power in specific frequency bands that are compatible with
scikit-learn.
Parameters
----------
epochs : Epochs
The data.
Returns
-------
X : numpy array of shape [n_samples, 5]
Transformed data.
"""
x = epochs.get_data()
X = np.zeros([x.shape[0],3])
for ep_idx in range(x.shape[0]):
# print(['EPOCH-->',str(ep_idx)])
eeg_x = x[ep_idx,0,:]
H,n_w,F_w = test_mat_dfa(eeg_x)
sd_x = np.std(eeg_x, ddof=1)
samp_en = nolds.sampen(eeg_x, emb_dim=3, tolerance= 0.2*sd_x)
NSEG = (x.shape[0] + 1)/10
NSEG_OV = np.round(NSEG*0.5)
f,PX = welch(eeg_x, fs=1, window='hann', nperseg=NSEG, noverlap=NSEG_OV, nfft=None)
f_loc = np.where(f<0.1)[0]
beta, _ = np.polyfit(np.log10(f[f_loc[1:]]),np.log10(PX[f_loc[1:]]), 1)
X[ep_idx,0] = H
X[ep_idx,1] = samp_en
X[ep_idx,2] = beta
return X
def eeg_power_band(epochs):
"""EEG relative power band feature extraction.
This function takes an ``mne.Epochs`` object and creates EEG features based
on relative power in specific frequency bands that are compatible with
scikit-learn.
Parameters
----------
epochs : Epochs
The data.
Returns
-------
X : numpy array of shape [n_samples, 5]
Transformed data.
"""
# specific frequency bands
FREQ_BANDS = {"delta": [0.5, 4.5],
"theta": [4.5, 8.5],
"alpha": [8.5, 11.5],
"sigma": [11.5, 15.5],
"beta": [15.5, 30]}
psds, freqs = psd_welch(epochs, picks='eeg', fmin=0.5, fmax=30.)
# Normalize the PSDs
psds /= np.sum(psds, axis=-1, keepdims=True)
X = []
for fmin, fmax in FREQ_BANDS.values():
psds_band = psds[:, :, (freqs >= fmin) & (freqs < fmax)].mean(axis=-1)
X.append(psds_band.reshape(len(psds), -1))
print(np.concatenate(X, axis=1).shape)
return np.concatenate(X, axis=1)
# +
pipe = make_pipeline(FunctionTransformer(eeg_power_band, validate=False),
RandomForestClassifier(n_estimators=100, random_state=42))
# Train
y_train = epochs_train.events[:, 2]
pipe.fit(epochs_train, y_train)
# Test
y_pred = pipe.predict(epochs_test)
# Assess the results
y_test = epochs_test.events[:, 2]
acc = accuracy_score(y_test, y_pred)
print("Accuracy score: {}".format(acc))
# +
pipe = make_pipeline(FunctionTransformer(eeg_nld, validate=False),
RandomForestClassifier(n_estimators=100, random_state=42))
# Train
y_train = epochs_train.events[:, 2]
pipe.fit(epochs_train, y_train)
# Test
y_pred = pipe.predict(epochs_test)
# Assess the results
y_test = epochs_test.events[:, 2]
acc = accuracy_score(y_test, y_pred)
print("Accuracy score: {}".format(acc))
|
notebook/.ipynb_checkpoints/05_EEG_signals-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interpret topic-model results
#
# We ran ```topicmodel_gridsearch.Rmd``` five times, producing five different versions of ```topicmodel_deltas.csv.``` (All uploaded to github.)
#
# The code below averages the five runs, and then pairs the averaged results with a data frame called ```interpret``` that stores the topic categorizes and sizes (the percentage of the corpus in each topic). These are used to produce a weighted average of delta across the corpus.
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
from scipy.stats import pearsonr, spearmanr, ttest_ind, linregress
from pathlib import Path
# ### Load the results of the R scripts.
# +
root = 'mainresults/topicmodel_deltas'
deltas = dict()
for i in range(1, 6):
suffix = str(i) + '.csv'
deltas[i] = pd.read_csv(Path(root + suffix))
print(len(deltas), ' files loaded.')
# -
# There is some continuity but also significant divergence between different runs.
deltas[4].head()
# +
# How much stability is there between runs?
# Note: I'm using arctanh and tanh to average correlation coefficients
# because r is not normally distributed; it's bounded at 1 and -1, so
# a Fisher's z-transform is needed.
deltalist = []
deltaooslist = []
for i in range(1, 6):
for j in range(i + 1, 6):
deltalist.append(spearmanr(deltas[i].delta, deltas[j].delta)[0])
deltaooslist.append(spearmanr(deltas[i].delta_oos, deltas[j].delta_oos)[0])
def average_rs(rlist):
r_z = [np.arctanh(x) for x in rlist]
return np.tanh(np.mean(r_z))
print('Avg spearman in-sample: ', average_rs(deltalist))
print('Avg spearman oos: ', average_rs(deltaooslist))
# -
# ### Construct a data frame that has average values
# +
smoothed = dict()
cols = ['delta', 'adjdelta', 'delta_oos', 'cmse', 'pmse', 'cmse_oos', 'pmse_oos',
'totalr2', 'r2_oos', 'bydf', 'fpdf', 'agemse']
for c in cols:
if c not in smoothed:
smoothed[c] = []
for rownum in range(200):
values = []
for i in range(1, 6):
if c in deltas[i].columns: # the only exception is 'agemse' which got added late
# and won't be in all five runs
values.append(deltas[i].loc[rownum, c])
smoothed[c].append(np.mean(values))
avgdf = pd.DataFrame(smoothed)
# -
avgdf.head()
# +
# How close are different runs to the average?
for i in range(1, 6):
print('In sample: ', spearmanr(deltas[i].delta, avgdf.delta)[0])
print('Out of sample: ', spearmanr(deltas[i].delta_oos, avgdf.delta_oos)[0])
# -
# ### Load the coding of the 200 topics
interpret = pd.read_csv('../interrater/k200standardcoding.tsv', sep = '\t')
interpret.drop(columns = ['next5books, biggestauth, datequartiles', 'comments'], inplace = True)
interpret.head()
# For our present purposes, we don't need all this information, and we could use shorter category labels for visualization.
# +
# FUNCTIONS THAT SIMPLIFY AND CLEAN THE CODING FRAME
def getpct(astring):
firstpart = astring.split('%')[0]
number = float(firstpart.split('= ')[1])
return number
def shorten(astring):
shortversion = astring[0:6]
if shortversion == 'human ':
shortversion = 'institu' # for human institution, practice, or relationship
return shortversion
interpret['pct'] = interpret.topicstats.map(getpct)
interpret['category'] = interpret.category.map(shorten)
interpret.drop(columns = ['keywords', 'top7books', 'topicstats'], inplace = True)
interpret.head()
# -
# ### Concatenate the topic labels with all five frames of numeric results.
topics = dict()
for i in range(1, 6):
topics[i] = pd.concat([deltas[i], interpret], axis = 1)
# ### And also with the average of all five.
avgdf = pd.concat([avgdf, interpret], axis = 1)
# +
def weighted_avg(aframe):
avg = sum(aframe.pct * aframe.cmse) / (sum(aframe.pct * aframe.cmse) + sum(aframe.pct * aframe.pmse))
return avg
def weighted_avg_oos(aframe):
cmseoos = [0 if i < 0 else i for i in aframe.cmse_oos]
pmseoos = [0 if i < 0 else i for i in aframe.pmse_oos]
avg = sum(aframe.pct * cmseoos) / (sum(aframe.pct * cmseoos) + sum(aframe.pct * pmseoos))
return avg
# -
# ### Calculate average delta as per our pre-registered plan.
#
# The columns that matter most are the "weighted" ones; we've already decided to care more about variables where the model is strong than about ones where r2 is low and no chronological variables are very predictive--also more about large topics than small ones.
print('raw in-sample\traw oos\t\tweighted in-sample\tweighted oos')
for i in range(1, 6):
print(round(np.mean(topics[i].delta), 4), '\t\t', round(np.mean(topics[i].delta_oos), 4), '\t\t',
round(weighted_avg(topics[i]), 4), '\t\t', round(weighted_avg_oos(topics[i]), 4))
print("Overall, weighted in-sample is", round(weighted_avg(avgdf), 4))
print("And out-of-sample: ", round(weighted_avg_oos(avgdf), 4))
# Actually, technically we proposed to exclude 'accidents of transcription' and 'author-dominated' categories from the average. This makes only a small difference.
legit = avgdf.loc[(avgdf.category != 'accide') & (avgdf.category != 'author'), :]
print("Overall, weighted in-sample is", round(weighted_avg(legit), 4))
print("And out-of-sample: ", round(weighted_avg_oos(legit), 4))
# ### Differences between topic categories
#
# Our article focuses on the way r2 is partitioned by ANOVA (in-sample). In theory there are advantages to calculating on out-of-sample data, but I suspect in practice it's noisier, as seen by the low Spearman correlations in cells above.
#
# Here are category means (not weighted by topic size or r2):
cats = avgdf.category.unique()
deltadict = dict()
for c in cats:
deltadict[c] = avgdf.loc[avgdf.category == c, 'delta'].tolist()
print(c, np.mean(deltadict[c]))
# And a visualization.
# +
def widen (astring):
mapper = {'event': 'event', 'genre': 'genre', 'dialec': 'dialect or language', 'accide': 'accident of transcription', 'author': 'author-dominated topic',
'techno': 'technological change', 'uncate': 'uncategorized', 'institu': 'institution or practice', 'nation': 'nationality or region',
'physic': 'physical description'}
return mapper[astring]
avgdf['longcategory'] = avgdf.category.map(widen)
# -
cm = 1/2.54
plt.figure(figsize = (11, 11))
res = sns.swarmplot(data = avgdf, x = 'delta', y = 'longcategory', order = ['event', 'accident of transcription', 'genre',
'dialect or language', 'uncategorized',
'institution or practice',
'author-dominated topic', 'technological change',
'physical description', 'nationality or region'], orient = 'h', size = 12)
plt.rcParams.update({'font.size': 20})
plt.ylabel('')
plt.gcf().set_facecolor('white')
# plt.savefig('/Users/tunder/Dropbox/python/cohort/figures/swarmdeltas.png', bbox_inches = 'tight', dpi = 300)
plt.show()
# The most dramatic and stable pattern is that 'event' topics have much lower delta, and are more dominated by period factors. Here they are; as you can see, they're mostly wars.
avgdf.loc[avgdf.category == 'event', : ]
# This pattern holds just as strongly if we measure delta out of sample.
deltadict = dict()
for c in cats:
deltadict[c] = avgdf.loc[avgdf.category == c, 'delta_oos'].tolist()
print(c, np.mean(deltadict[c]))
plt.figure(figsize = (10, 6))
sns.boxplot(data = avgdf, x = 'category', y = 'delta_oos', order = ['event', 'genre','dialec', 'accide', 'author', 'techno', 'uncate',
'institu', 'nation', 'physic'])
plt.rcParams.update({'font.size': 12})
plt.show()
# ### Testing our pre-registered hypothesis about categories
#
# We specified that we didn't expect *p* < .05 kind of significance here, because n is very low (n for the three categories is 5, 4, and 9)!
#
# Instead we were going to look at effect size. For instance, Cohen's *d*. And any way you measure this (in or out of sample), there is in fact a non-trivial effect.
#
# But this is mostly the ```event``` (and to some extent ```accident```) topics pulling down the ```technology``` category. If we just considered the technology topics by themselves, they wouldn't seem notably divergent from the rest of the model.
#
# So our hypothesis is confirmed, but with some internal variation that shows we're not seeing exactly what we might have expected.
target_cat_deltas = avgdf.loc[(avgdf.category == 'event') | (avgdf.category == 'accide') | (avgdf.category == 'techno'), 'delta']
np.mean(target_cat_deltas)
other_deltas = avgdf.loc[(avgdf.category != 'event') & (avgdf.category != 'accide') & (avgdf.category != 'techno'), 'delta']
np.mean(other_deltas)
# +
## Calculate Cohen's d.
(np.mean(other_deltas) - np.mean(target_cat_deltas)) / np.std(avgdf.delta)
# -
# ### Where is r2 strong?
#
# This is a bit of a surprise. "Genre" and "physical description" are the categories where chronological variables have most explanatory power.
#
# I might have expected ```technology``` and ```event``` to have higher r2s.
plt.figure(figsize = (10, 6))
sns.boxplot(data = avgdf, x = 'category', y = 'totalr2')
plt.show()
# ### Calculate average r2 weighted by topic size
weightedavgr2 = np.average(avgdf.totalr2, weights = avgdf.pct)
print('r2: ', weightedavgr2)
naiver2 = np.mean(avgdf.totalr2)
naiver2
pearsonr(avgdf.totalr2, avgdf.delta)
pearsonr(avgdf.totalr2, avgdf.pct)
# So, there's a significant and moderately strong pattern that r2 is larger when a topic contains more words. Not very surprising.
plt.figure(figsize = (10, 10))
sns.swarmplot(data = avgdf, y = 'category', x = 'pct', orient = 'h', size = 7)
plt.show()
# Categories are quite strongly grouped, but this is not altogether surprising since r2 correlates with topic size and topic size is one of the things we looked at in categorizing.
# ### Examine particular topics to understand patterns better
#
# Let's look at the trajectories of topics over time to understand why they are associated with period or cohort.
bookdata = pd.read_csv('../topicdata/bookleveltopicdata.tsv', sep = '\t', low_memory = False)
data = bookdata.loc[bookdata.us_national & (bookdata.firstpub > 1889) & (bookdata.firstpub < 1990), : ]
data.shape
def extract_trajectories(depvar, data):
avgbyfirstpub = []
avgbybirthyear = []
firstpubtimeline = []
birthyeartimeline = []
for fp, df in data.groupby('firstpub'):
if fp > 1889 and fp < 1990:
firstpubtimeline.append(fp)
avgbyfirstpub.append(np.mean(df[depvar]))
for by, df in data.groupby('birthyear'):
if by > 1850 and by < 1960:
birthyeartimeline.append(by)
avgbybirthyear.append(np.mean(df[depvar]))
return firstpubtimeline, avgbyfirstpub, birthyeartimeline, avgbybirthyear
fpxaxis, avgbyfirstpub, byxaxis, avgbybirthyear = extract_trajectories('t0', data)
sns.lineplot(x = fpxaxis, y = avgbyfirstpub)
plt.title('Topic 0, War mostly WWII, by firstpub')
plt.show()
# You can see why models of that trajectory invariably choose a 4-year granularity. It's very tightly responsive to events.
sns.lineplot(x = byxaxis, y = avgbybirthyear)
plt.title('Topic 0, War mostly WWII, by birth year')
plt.show()
# Although there is one crazy spike there, models of birthyear choose a 16-year granularity on average, because it's a smoother pattern on the whole.
fpxaxis, avgbyfirstpub, byxaxis, avgbybirthyear = extract_trajectories('t141', data)
sns.lineplot(x = byxaxis, y = avgbybirthyear)
plt.title('Topic 141, late 20c cities, by birth year')
plt.show()
sns.lineplot(x = fpxaxis, y = avgbyfirstpub)
plt.title('Topic 141, late 20c cities, by firstpub')
plt.show()
# +
x = []
y = []
variable = []
topicnumber = []
for tn in ['t0', 't141']:
fpxaxis, avgbyfirstpub, byxaxis, avgbybirthyear = extract_trajectories(tn, data)
if tn == 't0':
topicname = "War"
else:
topicname = "Urban life"
x.extend(fpxaxis)
y.extend(avgbyfirstpub)
variable.extend(['pub. year'] * len(fpxaxis))
topicnumber.extend([topicname] * len(fpxaxis))
x.extend(byxaxis)
y.extend(avgbybirthyear)
variable.extend(["birth year"] * len(byxaxis))
topicnumber.extend([topicname] * len(byxaxis))
facetdf = pd.DataFrame({'year': x, 'prominence': y, 'x': variable, 'topic': topicnumber})
# +
plt.rcParams.update({'font.sans-serif':'Avenir'})
plt.rcParams.update({'font.size': 14})
g = sns.FacetGrid(facetdf, row = 'topic', col = 'x', hue = 'x', height = 4, aspect = 1.2,
sharex = False, sharey = False, palette = 'deep')
g.map(sns.lineplot, 'year', 'prominence')
g.axes[1,0].set_xlabel('year of publication')
g.axes[1,1].set_xlabel("year of author's birth")
g.axes[1,0].set_ylabel("Topic 141: late 20c urban life")
g.axes[0,0].set_ylabel('Topic 0: war, mostly WWII')
plt.gcf().set_facecolor('white')
# plt.savefig('/Users/tunder/Dropbox/python/cohort/figures/lineplotgrid.png', bbox_inches = 'tight', dpi = 300)
# -
# ### A slope hypothesis producing null results
#
# Patterns like the one below give us some reason to think that period-driven changes cause steep upslopes, whereas cohorts become more determinative when topics are in decline. (Writers "hold onto the practices they learned in youth.")
#
# This seems to be true of crime fiction.
sns.lineplot(x = byxaxis, y = avgbybirthyear)
plt.title('Topic 50, mid 20c crime fiction, by birth year')
plt.show()
sns.lineplot(x = fpxaxis, y = avgbyfirstpub)
plt.title('Topic 50, mid20c crime fiction, by firstpub')
plt.show()
# It's also supported by a pattern we find in the "technology" category. Generally speaking, emergent technologies, with a positive slope, are more period-driven than residual technologies, with a net negative slope across the century.
# +
techno = avgdf.loc[avgdf.category == 'techno', : ].copy()
techno.sort_values(by = 'delta', inplace = True)
slopes = []
for idx, row in techno.iterrows():
fpxaxis, avgbyfirstpub, byxaxis, avgbybirthyear = extract_trajectories('t' + str(idx), data)
slope_0, intercept, r_value, p_value, std_err = linregress(fpxaxis, avgbyfirstpub)
slopes.append(slope_0)
techno = techno.assign(slope = slopes)
techno.loc[ : , ['delta', 'slope', 'totalr2', 'bydf', 'fpdf', 'label']]
# -
plt.scatter(techno.delta, techno.slope)
# However we don't find a lot of evidence that this is true across all 200 topics.
# +
slopes = []
for topnum in range(200):
fpxaxis, avgbyfirstpub, byxaxis, avgbybirthyear = extract_trajectories('t' + str(topnum), data)
slope_0, intercept, r_value, p_value, std_err = linregress(fpxaxis, avgbyfirstpub)
# r2, p = spearmanr(byxaxis, avgbybirthyear)
slopes.append(slope_0)
slopes = np.array(slopes)
# -
pearsonr(slopes, avgdf.delta)
# ### Comparison to the SEM experiment
# How do the results of structural equation modeling compare to the deltas we inferred from our regression experiment?
#
# First we load the SEM results.
sem = pd.read_csv('../sem-topics/sem_topic_preferred.csv')
sem
# Then we separate two categories of topics: those where the active updating model was preferred, and those where the settled disposition model was preferred.
aumdeltas = avgdf.loc[sem.verdict == 'AUM', 'delta']
sdmdeltas = avgdf.loc[sem.verdict == 'SDM', 'delta']
# A naive approach would be to directly calculate the difference between these deltas by taking the means and using a t-test to evaluate the difference of means.
print('AUM: ', round(np.mean(aumdeltas), 4))
print('SDM: ', round(np.mean(sdmdeltas), 4))
ttest_ind(aumdeltas, sdmdeltas, equal_var = False) # Welch's t-test
# The difference would be significant, if we took that naive approach.
#
# #### A better approach
#
# However, recall that in reality we weight deltas both by $r^2$ and by the size of the topic when calculating overall delta. It's reasonable to apply the same weights to calculate the means of the two groups here, and apply them also to the t-test.
avgdf = avgdf.assign(weights = avgdf.pct * avgdf.totalr2)
avgdf = avgdf.assign(weights = avgdf.weights * (200 / sum(avgdf.weights)))
aumpct = avgdf.loc[sem.verdict == 'AUM', 'pct']
sdmpct = avgdf.loc[sem.verdict == 'SDM', 'pct']
aumr2 = avgdf.loc[sem.verdict == 'AUM', 'totalr2']
sdmr2 = avgdf.loc[sem.verdict == 'SDM', 'totalr2']
aumweights = aumpct * aumr2
sdmweights = sdmpct * sdmr2
allweights = np.concatenate([aumweights, sdmweights])
allweights = allweights * (len(allweights) / sum(allweights))
print(sum(allweights))
# That's a sanity check; the numbers should add up to the length of the vectors.
aumweights = allweights[0 : len(aumweights)]
sdmweights = allweights[len(aumweights) : ]
from statsmodels.stats.weightstats import ttest_ind # a more flexible implementation
ttest_ind(x1 = aumdeltas, x2 = sdmdeltas, weights = (aumweights, sdmweights), usevar = 'unequal') # returns t statistic, pval, dfs
np.average(aumdeltas, weights = aumweights)
np.average(sdmdeltas, weights = sdmweights)
# Those are the means and statistics we report in the article.
sum((sem.verdict == 'AUM') & (avgdf.delta > 0.5)) # I quote this figure at one point
# #### visualizing the difference of means described above
avgdf['semtype'] = sem.verdict.values
sns.swarmplot(data = avgdf, x = 'semtype', y = 'delta')
|
regression/InterpretTopicResults.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What is machine learning?
#
# The art and science of:
#
# giving computer the ability to learn to make decisions from data... without being explicitly programmed!!
#
# Examples:
#
# Learning to predict whether an email is spam or not (the most typical example).
#
# Clustering wikipedia entries into different categories.
# Unsupervised learning: Uncovering hidden patterns from unlabeled data.
#
# Exmple:
#
# Grouping customers into distinct categories (Clustering)
# Reinforcement learning: Software agents interact with an enviroment.
# Learn how to optimize their behavior.
# Given a system of rewards and punishments.
#
# note: In 2016, reinforcement learning was used to train Google DeepMind's Alpha Go, which was the first computer program to beat the world champion in Go.
# Supervised learning : predictor variables/features and a target variable. The goal is predict the target variable, given the predictor variables.
#
# In classification the target variable consist of categories.
#
# note: Here, we will use scikit-learn/sklearn. Also, there are others like TensorFlow, Keras and Pytorch.
#
#
# Now, we will import the libraries that we will need to code.
# Also, we will set the plotting style to 'ggplot' using plt dot style dot use. Firstl, because it looks great and secondly, in order to help all you R aficionados feel at home XD!!!
import sklearn.datasets
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# Then, we load the dataset and assign the data to a variable iris. In the second line, we print the keys, where DESCR provides a description of the dataset; the target names which contains the values features; and the target, which is the target data.
iris = sklearn.datasets.load_iris()
print(iris.keys())
type(iris.data), type(iris.target)
# In data, there are 150 rows and 4 columns. (samples are in rows and columns are in columns). The categories consist en 'setosa','vesicolor','virginica' which we can see them printing the target names (or categories).
iris.data.shape
iris.target_names
# In order to realize EDA (Exploratory data analysis), we wil assign the feature and target data to x and y, respectively. So, we construct a DataFrame of Pandas (in this case as pd) with the feature names.
# +
X = iris.data
Y = iris.target
df = pd.DataFrame(X, columns = iris.feature_names)
print(df)
# -
# we will visualize the data using scatter. We pass it the our DataFrame, along with our target variables as argument to the parameter c (thus, ensuring that each species will be colored in direfent way). figsize specifies the size of figure. S determinates the marker size and the marker stablish the type.
pd.plotting.scatter_matrix(df, c = Y, figsize = [5, 10], s = 100, marker = 'A')
# In the figure, the diagonal are histograms of the features. The off-diagonal figures are scatter plots of the column feature vs row feature colored by the target variable. How we can see in the sub-figure (row 3, column 4), the petal width and length are highly correlated, as you may expect, and that flowers are clustered according to species.
#
# note: you should use the next methods in order to get data information.
print(df.info())
print(df.describe())
|
Machine learning basics/SupervisedLearningWithScikitLearn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
# %pylab inline
# %matplotlib inline
# + [markdown] deletable=true editable=true
# ## Preliminaries
# + deletable=true editable=true
wheretosave='/home/ubuntu/Notebooks/xcell_trainingdata.mat'
no_data_ex=590
no_data_validation=0
no_data_test=0
total_data_ex=590
# + deletable=true editable=true
import scipy.io as sio
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import tensorflow as tf
from __future__ import division
run_opts = tf.RunOptions(report_tensor_allocations_upon_oom = True)
# + deletable=true editable=true
datapath='/home/ubuntu/Notebooks/randflicker_20191212_preconv_data.mat'
data=sio.loadmat(datapath)
## Handle training data: Stimuli
input_bip1_0 = data['b1_input']
input_bip2_0 = data['b2_input']
input_bip3_0 = data['b3_input']
input_bip4_0 = data['b4_input']
input_bip5_0 = data['b5_input']
input_bip6_0 = data['b6_input']
input_bip7_0 = data['b7_input']
input_bip8_0 = data['b8_input']
input_bip9_0 = data['b9_input']
input_bip10_0 = data['b10_input']
input_bip11_0 = data['b11_input']
input_bip12_0 = data['b12_input']
input_bip13_0 = data['b13_input']
input_bip14_0 = data['b14_input']
numpix=21
data_duration=input_bip1_0.shape[1]
print(data_duration)
def rearrange_bip_input(input_bip_0, startind, endind):
input_bip_1 = reshape(input_bip_0, [1, total_data_ex, data_duration, numpix])
input_bip_11 = input_bip_1[:, startind:endind, :, :]
input_bip_2 = np.swapaxes(input_bip_11, 0, 3)
input_bip_3 = reshape(input_bip_2, [numpix, no_data_ex, data_duration])
return input_bip_3
startind = 0
endind = no_data_ex
input_bip1_train = rearrange_bip_input(input_bip1_0, startind, endind)
input_bip2_train = rearrange_bip_input(input_bip2_0, startind, endind)
input_bip3_train = rearrange_bip_input(input_bip3_0, startind, endind)
input_bip4_train = rearrange_bip_input(input_bip4_0, startind, endind)
input_bip5_train = rearrange_bip_input(input_bip5_0, startind, endind)
input_bip6_train = rearrange_bip_input(input_bip6_0, startind, endind)
input_bip7_train = rearrange_bip_input(input_bip7_0, startind, endind)
input_bip8_train = rearrange_bip_input(input_bip8_0, startind, endind)
input_bip9_train = rearrange_bip_input(input_bip9_0, startind, endind)
input_bip10_train = rearrange_bip_input(input_bip10_0, startind, endind)
input_bip11_train = rearrange_bip_input(input_bip11_0, startind, endind)
input_bip12_train = rearrange_bip_input(input_bip12_0, startind, endind)
input_bip13_train = rearrange_bip_input(input_bip13_0, startind, endind)
input_bip14_train = rearrange_bip_input(input_bip14_0, startind, endind)
# + deletable=true editable=true
# SET NUMBER OF NEURONS IN EACH LAYER
no_filters=14
no_bipolars=21
no_bipolar_types=14
no_relu=0
no_am_types = 5
no_am1=7
no_am2=21
no_am3=21
no_gc=8
# + [markdown] deletable=true editable=true
# ## load and handle filters
# + deletable=true editable=true
filter_data=sio.loadmat('/home/ubuntu/Notebooks/dc_filterbank.mat')
bip_filters=filter_data['franke_dc_filters']
ds=2 #downsample factor
bip_filters=bip_filters[::ds, :] #just take every 10th sample.
bip_filtersize=shape(bip_filters)[0]
bip_filters=np.transpose(bip_filters)
print(bip_filtersize)
bip_filters=np.reshape(bip_filters, [14, bip_filtersize, 1, 1, 1])
filter1=np.reshape(bip_filters[0, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter2=np.reshape(bip_filters[1, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter3=np.reshape(bip_filters[2, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter4=np.reshape(bip_filters[3, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter5=np.reshape(bip_filters[4, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter6=np.reshape(bip_filters[5, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter7=np.reshape(bip_filters[6, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter8=np.reshape(bip_filters[7, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter9=np.reshape(bip_filters[8, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter10=np.reshape(bip_filters[9, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter11=np.reshape(bip_filters[10, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter12=np.reshape(bip_filters[11, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter13=np.reshape(bip_filters[12, :, :, :, :], [bip_filtersize, 1, 1, 1])
filter14=np.reshape(bip_filters[13, :, :, :, :], [bip_filtersize, 1, 1, 1])
# + [markdown] deletable=true editable=true
# ## Helper Functions
# + deletable=true editable=true
#Helper Functions
def bias_var(shape, initial_val):
initial = tf.constant(initial_val, shape=shape)
# initial = tf.random_uniform(shape, minval=0.5, maxval=3.5, dtype=tf.float32) #2.9
return tf.Variable(initial)
def synapse_var(shape, initial_val):
initial=tf.constant(initial_val, shape=shape)
# initial = tf.random_uniform(shape, minval=0.1, maxval=0.8, dtype=tf.float32) #2.9
return tf.Variable(initial) #initial
def ag_synapse_var(shape, true_initial_val, train_initial_val):
initial=tf.constant(true_initial_val, shape=shape)
# initial=tf.constant(train_initial_val, shape=shape)
# initial = tf.random_uniform(shape, minval=0.1, maxval=0.8, dtype=tf.float32) #2.9
return tf.Variable(initial) #initial
def pbconv2d(x, W):
padsize=175 #200 #W.shape[0]
paddedx=tf.pad(x, [[0, 0], [padsize, 0], [0, 0], [0, 0]], 'CONSTANT')
outconv=tf.nn.conv2d(paddedx, W, strides=[1, 1, 1, 1], padding='SAME') #250 for movingdot and noise
#return tf.reshape(outconv[:, np.round(padsize/2).astype(int):np.round(padsize/2).astype(int)+250, 0, 0], [-1, 250, 1, 1])
return tf.reshape(outconv[:, np.round(padsize/2).astype(int):np.round(padsize/2).astype(int)+x_train.shape[1], 0, 0], [-1, x_train.shape[1], 1, 1])
# + [markdown] deletable=true editable=true
# # Trained Network
# + deletable=true editable=true
input_bip1_ = tf.placeholder("float32", name="input_bip1")
input_bip2_ = tf.placeholder("float32", name="input_bip2")
input_bip3_ = tf.placeholder("float32", name="input_bip3")
input_bip4_ = tf.placeholder("float32", name="input_bip4")
input_bip5_ = tf.placeholder("float32", name="input_bip5")
input_bip6_ = tf.placeholder("float32", name="input_bip6")
input_bip7_ = tf.placeholder("float32", name="input_bip7")
input_bip8_ = tf.placeholder("float32", name="input_bip8")
input_bip9_ = tf.placeholder("float32", name="input_bip9")
input_bip10_ = tf.placeholder("float32", name="input_bip10")
input_bip11_ = tf.placeholder("float32", name="input_bip11")
input_bip12_ = tf.placeholder("float32", name="input_bip12")
input_bip13_ = tf.placeholder("float32", name="input_bip13")
input_bip14_ = tf.placeholder("float32", name="input_bip14")
# + deletable=true editable=true
# Define Variables
b1g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 5.5]
b2g = [0.0, 0.5, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b3g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 5.5, 0.0]
b4g = [0.0, 0.0, 0.5, 0.0, 0.0, 0.0, 0.0, 0.0]
b5g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.5, 0.0, 0.0]
b6g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b7g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b8g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b9g = [0.0, 0.0, 0.0, 0.0, 0.5, 0.0, 0.0, 0.0]
b10g = [0.0, 0.0, 0.0, 0.5, 0.0, 0.0, 0.0, 0.0]
b11g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b12g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b13g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b14g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b1copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b2copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b3copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b4copyg = [0.5, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b5copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b6copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b7copyg = [0.5, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b8copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b9copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b10copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b11copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b12copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b13copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b14copyg = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
b1b = 0.0
b2b = 0.0
b3b = -29.0
b4b = 0.0
b5b = 0.0
b6b = 0.0
b7b = 0.0
b8b = 0.0
b9b = 0.0
b10b = -0.0
b11b = -0.0
b12b = 0.0
b13b = 0.0
b14b = 0.0
b1a1 = 0.0
b2a1 = 0.0
b3a1 = 0.0
b4a1 = 0.5
b5a1 = 0.0
b6a1 = 0.0
b7a1 = 0.5
b8a1 = 0.0
b9a1 = 0.0
b10a1 = 0.0
b11a1 = 0.0
b12a1 = 0.0
b13a1 = 0.0
b14a1 = 0.0
b1a2 = 0.0
b2a2 = 0.0
b3a2 = 0.0
b4a2 = 0.0
b5a2 = 0.0
b6a2 = 0.0
b7a2 = 0.0
b8a2 = 0.0
b9a2 = 0.0
b10a2 = 1.7
b11a2 = 0.0
b12a2 = 0.0
b13a2 = 0.0
b14a2 = 0.0
b1a3 = 0.0
b2a3 = 0.0
b3a3 = 0.0
b4a3 = 0.0
b5a3 = 0.0
b6a3 = 0.0
b7a3 = 0.0
b8a3 = 0.0
b9a3 = 0.0
b10a3 = 0.0
b11a3 = 1.0
b12a3 = 0.0
b13a3 = 0.0
b14a3 = 0.0
a1g = [1.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]
a2g = [0.0, 0.0, 0.0, 0.0, 0.0, 100.0, 0.0, 0.0]
a3g = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 1.5, 0.0]
a1b3copy = 0.0
a1b4copy = 0.1
a1b5copy = 0.0
a1b6copy = 0.0
a1b7copy = 0.1
a1b8copy = 0.0
a1b9copy = 0.0
bip1_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in [5, 6, 7, 8]:
for gc_i in range(no_gc):
bip1_gc_initial[bip_i, gc_i]=b1g[gc_i]
bip1_gc_initial=bip1_gc_initial.astype(float32)
bip2_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip2_gc_initial[bip_i, gc_i]=b2g[gc_i]
bip2_gc_initial=bip2_gc_initial.astype(float32)
bip3_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in [5, 6, 7, 8]:
for gc_i in range(no_gc):
bip3_gc_initial[bip_i, gc_i]=b3g[gc_i]
bip3_gc_initial=bip3_gc_initial.astype(float32)
bip4_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip4_gc_initial[bip_i, gc_i]=b4g[gc_i]
bip4_gc_initial=bip4_gc_initial.astype(float32)
bip5_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip5_gc_initial[bip_i, gc_i]=b5g[gc_i]
bip5_gc_initial=bip5_gc_initial.astype(float32)
bip6_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip6_gc_initial[bip_i, gc_i]=b6g[gc_i]
bip6_gc_initial=bip6_gc_initial.astype(float32)
bip7_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip7_gc_initial[bip_i, gc_i]=b7g[gc_i]
bip7_gc_initial=bip7_gc_initial.astype(float32)
bip8_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip8_gc_initial[bip_i, gc_i]=b8g[gc_i]
bip8_gc_initial=bip8_gc_initial.astype(float32)
bip9_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip9_gc_initial[bip_i, gc_i]=b9g[gc_i]
bip9_gc_initial=bip9_gc_initial.astype(float32)
bip10_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip10_gc_initial[bip_i, gc_i]=b10g[gc_i]
bip10_gc_initial=bip10_gc_initial.astype(float32)
bip11_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(3):
bip11_gc_initial[bip_i, gc_i]=b11g[gc_i]
bip11_gc_initial=bip11_gc_initial.astype(float32)
bip12_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip12_gc_initial[bip_i, gc_i]=b12g[gc_i]
bip12_gc_initial=bip12_gc_initial.astype(float32)
bip13_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip13_gc_initial[bip_i, gc_i]=b13g[gc_i]
bip13_gc_initial=bip13_gc_initial.astype(float32)
bip14_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip14_gc_initial[bip_i, gc_i]=b14g[gc_i]
bip14_gc_initial=bip14_gc_initial.astype(float32)
bip1_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip1_copy_gc_initial[bip_i, gc_i]=b1copyg[gc_i]
bip1_copy_gc_initial=bip1_copy_gc_initial.astype(float32)
bip2_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip2_copy_gc_initial[bip_i, gc_i]=b2copyg[gc_i]
bip2_copy_gc_initial=bip2_copy_gc_initial.astype(float32)
bip3_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip3_copy_gc_initial[bip_i, gc_i]=b3copyg[gc_i]
bip3_copy_gc_initial=bip3_copy_gc_initial.astype(float32)
bip4_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip4_copy_gc_initial[bip_i, gc_i]=b4copyg[gc_i]
bip4_copy_gc_initial=bip4_copy_gc_initial.astype(float32)
bip5_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip5_copy_gc_initial[bip_i, gc_i]=b5copyg[gc_i]
bip5_copy_gc_initial=bip5_copy_gc_initial.astype(float32)
bip6_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip6_copy_gc_initial[bip_i, gc_i]=b6copyg[gc_i]
bip6_copy_gc_initial=bip6_copy_gc_initial.astype(float32)
bip7_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(4, 12):
for gc_i in range(no_gc):
bip7_copy_gc_initial[bip_i, gc_i]=b7copyg[gc_i]
bip7_copy_gc_initial=bip7_copy_gc_initial.astype(float32)
bip8_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip8_copy_gc_initial[bip_i, gc_i]=b8copyg[gc_i]
bip8_copy_gc_initial=bip8_copy_gc_initial.astype(float32)
bip9_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip9_copy_gc_initial[bip_i, gc_i]=b9copyg[gc_i]
bip9_copy_gc_initial=bip9_copy_gc_initial.astype(float32)
bip10_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip10_copy_gc_initial[bip_i, gc_i]=b10copyg[gc_i]
bip10_copy_gc_initial=bip10_copy_gc_initial.astype(float32)
bip11_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(3):
bip11_gc_initial[bip_i, gc_i]=b11copyg[gc_i]
bip11_copy_gc_initial=bip11_copy_gc_initial.astype(float32)
bip12_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip12_gc_initial[bip_i, gc_i]=b12copyg[gc_i]
bip12_copy_gc_initial=bip12_copy_gc_initial.astype(float32)
bip13_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip13_copy_gc_initial[bip_i, gc_i]=b13copyg[gc_i]
bip13_copy_gc_initial=bip13_copy_gc_initial.astype(float32)
bip14_copy_gc_initial=np.zeros([no_bipolars, no_gc])
for bip_i in range(8):
for gc_i in range(no_gc):
bip14_copy_gc_initial[bip_i, gc_i]=b14copyg[gc_i]
bip14_copy_gc_initial=bip14_copy_gc_initial.astype(float32)
# + deletable=true editable=true
am1_b3copy_initial=np.zeros([no_am1, no_bipolars])
for am_i in range(3):
for bip_i in range(8):
am1_b3copy_initial[am_i, bip_i]=a1b3copy
am1_b3copy_initial=am1_b3copy_initial.astype(float32)
am1_b4copy_initial=np.zeros([no_am1, no_bipolars])
for am_i in range(1):
for bip_i in range(4, 12):
am1_b4copy_initial[am_i, bip_i]=a1b4copy
am1_b4copy_initial=am1_b4copy_initial.astype(float32)
am1_b5copy_initial=np.zeros([no_am1, no_bipolars])
for am_i in range(3):
for bip_i in range(8):
am1_b5copy_initial[am_i, bip_i]=a1b5copy
am1_b5copy_initial=am1_b5copy_initial.astype(float32)
am1_b6copy_initial=np.zeros([no_am1, no_bipolars])
for am_i in range(3):
for bip_i in range(8):
am1_b6copy_initial[am_i, bip_i]=a1b6copy
am1_b6copy_initial=am1_b6copy_initial.astype(float32)
am1_b7copy_initial=np.zeros([no_am1, no_bipolars])
for am_i in range(1):
for bip_i in range(4, 12):
am1_b7copy_initial[am_i, bip_i]=a1b7copy
am1_b7copy_initial=am1_b7copy_initial.astype(float32)
am1_b8copy_initial=np.zeros([no_am1, no_bipolars])
for am_i in range(3):
for bip_i in range(8):
am1_b8copy_initial[am_i, bip_i]=a1b8copy
am1_b8copy_initial=am1_b8copy_initial.astype(float32)
am1_b9copy_initial=np.zeros([no_am1, no_bipolars])
for am_i in range(3):
for bip_i in range(8):
am1_b9copy_initial[am_i, bip_i]=a1b9copy
am1_b9copy_initial=am1_b9copy_initial.astype(float32)
am1_gc_initial=np.zeros([no_am1, no_gc])
for am_i in range(3):
for gc_i in range(no_gc):
am1_gc_initial[am_i, gc_i]=a1g[gc_i]
am1_gc_initial=am1_gc_initial.astype(float32)
am1_gc_train_initial=np.zeros([no_am1, no_gc])
for am_i in range(no_am1):
am1_gc_train_initial[am_i, 0]=0.0*np.random.uniform()
am1_gc_train_initial[am_i, 1]=0.0*np.random.uniform()
am1_gc_train_initial[am_i, 2]=0.0*np.random.uniform()
am1_gc_train_initial[am_i, 3]=0.0*np.random.uniform()
am1_gc_train_initial[am_i, 4]=0.0*np.random.uniform()
am1_gc_train_initial=am1_gc_train_initial.astype(float32)
am2_gc_initial=np.zeros([no_am2, no_gc])
for am_i in range(4, 12):
for gc_i in range(no_gc):
am2_gc_initial[am_i, gc_i]=a2g[gc_i]
am2_gc_initial=am2_gc_initial.astype(float32)
am2_gc_train_initial=np.zeros([no_am2, no_gc])
for am_i in range(no_am1):
am2_gc_train_initial[am_i, 0]=0.0*np.random.uniform()
am2_gc_train_initial[am_i, 1]=0.0*np.random.uniform()
am2_gc_train_initial[am_i, 2]=0.0*np.random.uniform()
am2_gc_train_initial[am_i, 3]=0.0*np.random.uniform()
am2_gc_train_initial[am_i, 4]=0.0*np.random.uniform()
am2_gc_train_initial=am2_gc_train_initial.astype(float32)
am3_gc_initial=np.zeros([no_am3, no_gc])
for am_i in [1, 2, 3, 4, 9, 10, 11, 12]:
for gc_i in range(no_gc):
am3_gc_initial[am_i, gc_i]=a3g[gc_i]
am3_gc_initial=am3_gc_initial.astype(float32)
am3_gc_train_initial=np.zeros([no_am3, no_gc])
for am_i in range(no_am1):
am3_gc_train_initial[am_i, 0]=0.0*np.random.uniform()
am3_gc_train_initial[am_i, 1]=0.0*np.random.uniform()
am3_gc_train_initial[am_i, 2]=0.0*np.random.uniform()
am3_gc_train_initial[am_i, 3]=0.0*np.random.uniform()
am3_gc_train_initial[am_i, 4]=0.0*np.random.uniform()
am3_gc_train_initial=am3_gc_train_initial.astype(float32)
bip1_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip1_am1_initial[bip_i, am_i]=b1a1
bip1_am1_initial=bip1_am1_initial.astype(float32)
bip2_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip2_am1_initial[bip_i, am_i]=b2a1
bip2_am1_initial=bip2_am1_initial.astype(float32)
bip3_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip3_am1_initial[bip_i, am_i]=b3a1
bip3_am1_initial=bip3_am1_initial.astype(float32)
bip4_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(0, 4):
bip4_am1_initial[bip_i, am_i]=b4a1
for am_i in range(1):
for bip_i in range(12, 16):
bip4_am1_initial[bip_i, am_i]=b4a1
bip4_am1_initial=bip4_am1_initial.astype(float32)
bip5_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip5_am1_initial[bip_i, am_i]=b5a1
bip5_am1_initial=bip5_am1_initial.astype(float32)
bip6_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip6_am1_initial[bip_i, am_i]=b6a1
bip6_am1_initial=bip6_am1_initial.astype(float32)
bip7_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(0, 4):
bip7_am1_initial[bip_i, am_i]=b7a1
for am_i in range(1):
for bip_i in range(12, 16):
bip7_am1_initial[bip_i, am_i]=b7a1
bip7_am1_initial=bip7_am1_initial.astype(float32)
bip8_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip8_am1_initial[bip_i, am_i]=b8a1
bip8_am1_initial=bip8_am1_initial.astype(float32)
bip9_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip9_am1_initial[bip_i, am_i]=b9a1
bip9_am1_initial=bip9_am1_initial.astype(float32)
bip10_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip10_am1_initial[bip_i, am_i]=b10a1
bip10_am1_initial=bip10_am1_initial.astype(float32)
bip11_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip11_am1_initial[bip_i, am_i]=b11a1
bip11_am1_initial=bip11_am1_initial.astype(float32)
bip12_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip12_am1_initial[bip_i, am_i]=b12a1
bip12_am1_initial=bip12_am1_initial.astype(float32)
bip13_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip13_am1_initial[bip_i, am_i]=b13a1
bip13_am1_initial=bip13_am1_initial.astype(float32)
bip14_am1_initial=np.zeros([no_bipolars, no_am1])
for am_i in range(1):
for bip_i in range(8, 16):
bip14_am1_initial[bip_i, am_i]=b14a1
bip14_am1_initial=bip14_am1_initial.astype(float32)
bip1_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip1_am2_initial[bip_i, am_i]=b1a2
bip1_am2_initial=bip1_am2_initial.astype(float32)
bip2_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip2_am2_initial[bip_i, am_i]=b2a2
bip2_am2_initial=bip2_am2_initial.astype(float32)
bip3_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip3_am2_initial[bip_i, am_i]=b3a2
bip3_am2_initial=bip3_am2_initial.astype(float32)
bip4_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip4_am2_initial[bip_i, am_i]=b4a2
bip4_am2_initial=bip4_am2_initial.astype(float32)
bip5_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip5_am2_initial[bip_i, am_i]=b5a2
bip5_am2_initial=bip5_am2_initial.astype(float32)
bip6_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip6_am2_initial[bip_i, am_i]=b6a2
bip6_am2_initial=bip6_am2_initial.astype(float32)
bip7_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip7_am2_initial[bip_i, am_i]=b7a2
bip7_am2_initial=bip7_am2_initial.astype(float32)
bip8_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip8_am2_initial[bip_i, am_i]=b8a2
bip8_am2_initial=bip8_am2_initial.astype(float32)
bip9_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip9_am2_initial[bip_i, am_i]=b9a2
bip9_am2_initial=bip9_am2_initial.astype(float32)
bip10_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(4, 12):
bip10_am2_initial[am_i, am_i]=b10a2
bip10_am2_initial=bip10_am2_initial.astype(float32)
bip11_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip11_am2_initial[bip_i, am_i]=b11a2
bip11_am2_initial=bip11_am2_initial.astype(float32)
bip12_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip12_am2_initial[bip_i, am_i]=b12a2
bip12_am2_initial=bip12_am2_initial.astype(float32)
bip13_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip13_am2_initial[bip_i, am_i]=b13a2
bip13_am2_initial=bip13_am2_initial.astype(float32)
bip14_am2_initial=np.zeros([no_bipolars, no_am2])
for am_i in range(1):
for bip_i in range(4, 12):
bip14_am2_initial[bip_i, am_i]=b14a2
bip14_am2_initial=bip14_am2_initial.astype(float32)
bip1_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip1_am3_initial[bip_i, am_i]=b1a3
bip1_am3_initial=bip1_am3_initial.astype(float32)
bip2_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip2_am3_initial[bip_i, am_i]=b2a3
bip2_am3_initial=bip2_am3_initial.astype(float32)
bip3_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip3_am3_initial[bip_i, am_i]=b3a3
bip3_am3_initial=bip3_am3_initial.astype(float32)
bip4_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip4_am3_initial[bip_i, am_i]=b4a3
bip4_am3_initial=bip4_am3_initial.astype(float32)
bip5_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip5_am3_initial[bip_i, am_i]=b5a3
bip5_am3_initial=bip5_am3_initial.astype(float32)
bip6_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip6_am3_initial[bip_i, am_i]=b6a3
bip6_am3_initial=bip6_am3_initial.astype(float32)
bip7_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip7_am3_initial[bip_i, am_i]=b7a3
bip7_am3_initial=bip7_am3_initial.astype(float32)
bip8_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip8_am3_initial[bip_i, am_i]=b8a3
bip8_am3_initial=bip8_am3_initial.astype(float32)
bip9_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip9_am3_initial[bip_i, am_i]=b9a3
bip9_am3_initial=bip9_am3_initial.astype(float32)
bip10_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(4, 12):
bip10_am3_initial[am_i, am_i]=b10a3
bip10_am3_initial=bip10_am3_initial.astype(float32)
bip11_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in [1, 2, 3, 4, 9, 10, 11, 12, 13, 14, 15, 16]:
# for bip_i in range(4, 12):
bip11_am3_initial[am_i, am_i]=b11a3
bip11_am3_initial=bip11_am3_initial.astype(float32)
bip12_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip12_am3_initial[bip_i, am_i]=b12a3
bip12_am3_initial=bip12_am3_initial.astype(float32)
bip13_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip13_am3_initial[bip_i, am_i]=b13a3
bip13_am3_initial=bip13_am3_initial.astype(float32)
bip14_am3_initial=np.zeros([no_bipolars, no_am3])
for am_i in range(1):
for bip_i in range(4, 12):
bip14_am3_initial[bip_i, am_i]=b14a3
bip14_am3_initial=bip14_am3_initial.astype(float32)
# + deletable=true editable=true
bip1_gc_syn=synapse_var([no_bipolars, no_gc], bip1_gc_initial)
bip2_gc_syn=synapse_var([no_bipolars, no_gc], bip2_gc_initial)
bip3_gc_syn=synapse_var([no_bipolars, no_gc], bip3_gc_initial)
bip4_gc_syn=synapse_var([no_bipolars, no_gc], bip4_gc_initial)
bip5_gc_syn=synapse_var([no_bipolars, no_gc], bip5_gc_initial)
bip6_gc_syn=synapse_var([no_bipolars, no_gc], bip6_gc_initial)
bip7_gc_syn=synapse_var([no_bipolars, no_gc], bip7_gc_initial)
bip8_gc_syn=synapse_var([no_bipolars, no_gc], bip8_gc_initial)
bip9_gc_syn=synapse_var([no_bipolars, no_gc], bip9_gc_initial)
bip10_gc_syn=synapse_var([no_bipolars, no_gc], bip10_gc_initial)
bip11_gc_syn=synapse_var([no_bipolars, no_gc], bip11_gc_initial)
bip12_gc_syn=synapse_var([no_bipolars, no_gc], bip12_gc_initial)
bip13_gc_syn=synapse_var([no_bipolars, no_gc], bip13_gc_initial)
bip14_gc_syn=synapse_var([no_bipolars, no_gc], bip14_gc_initial)
bip1_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip1_copy_gc_initial)
bip2_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip2_copy_gc_initial)
bip3_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip3_copy_gc_initial)
bip4_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip4_copy_gc_initial)
bip5_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip5_copy_gc_initial)
bip6_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip6_copy_gc_initial)
bip7_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip7_copy_gc_initial)
bip8_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip8_copy_gc_initial)
bip9_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip9_copy_gc_initial)
bip10_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip10_copy_gc_initial)
bip11_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip11_copy_gc_initial)
bip12_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip12_copy_gc_initial)
bip13_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip13_copy_gc_initial)
bip14_copy_gc_syn=synapse_var([no_bipolars, no_gc], bip14_copy_gc_initial)
bip1_am1_syn = synapse_var([no_bipolars, no_am1], bip1_am1_initial)
bip2_am1_syn = synapse_var([no_bipolars, no_am1], bip2_am1_initial)
bip3_am1_syn = synapse_var([no_bipolars, no_am1], bip3_am1_initial)
bip4_am1_syn = synapse_var([no_bipolars, no_am1], bip4_am1_initial)
bip5_am1_syn = synapse_var([no_bipolars, no_am1], bip5_am1_initial)
bip6_am1_syn = synapse_var([no_bipolars, no_am1], bip6_am1_initial)
bip7_am1_syn = synapse_var([no_bipolars, no_am1], bip7_am1_initial)
bip8_am1_syn = synapse_var([no_bipolars, no_am1], bip8_am1_initial)
bip9_am1_syn = synapse_var([no_bipolars, no_am1], bip9_am1_initial)
bip10_am1_syn = synapse_var([no_bipolars, no_am1], bip10_am1_initial)
bip11_am1_syn = synapse_var([no_bipolars, no_am1], bip11_am1_initial)
bip12_am1_syn = synapse_var([no_bipolars, no_am1], bip12_am1_initial)
bip13_am1_syn = synapse_var([no_bipolars, no_am1], bip13_am1_initial)
bip14_am1_syn = synapse_var([no_bipolars, no_am1], bip14_am1_initial)
bip1_am2_syn = synapse_var([no_bipolars, no_am2], bip1_am2_initial)
bip2_am2_syn = synapse_var([no_bipolars, no_am2], bip2_am2_initial)
bip3_am2_syn = synapse_var([no_bipolars, no_am2], bip3_am2_initial)
bip4_am2_syn = synapse_var([no_bipolars, no_am2], bip4_am2_initial)
bip5_am2_syn = synapse_var([no_bipolars, no_am2], bip5_am2_initial)
bip6_am2_syn = synapse_var([no_bipolars, no_am2], bip6_am2_initial)
bip7_am2_syn = synapse_var([no_bipolars, no_am2], bip7_am2_initial)
bip8_am2_syn = synapse_var([no_bipolars, no_am2], bip8_am2_initial)
bip9_am2_syn = synapse_var([no_bipolars, no_am2], bip9_am2_initial)
bip10_am2_syn = synapse_var([no_bipolars, no_am2], bip10_am2_initial)
bip11_am2_syn = synapse_var([no_bipolars, no_am2], bip11_am2_initial)
bip12_am2_syn = synapse_var([no_bipolars, no_am2], bip12_am2_initial)
bip13_am2_syn = synapse_var([no_bipolars, no_am2], bip13_am2_initial)
bip14_am2_syn = synapse_var([no_bipolars, no_am2], bip14_am2_initial)
bip1_am3_syn = synapse_var([no_bipolars, no_am3], bip1_am3_initial)
bip2_am3_syn = synapse_var([no_bipolars, no_am3], bip2_am3_initial)
bip3_am3_syn = synapse_var([no_bipolars, no_am3], bip3_am3_initial)
bip4_am3_syn = synapse_var([no_bipolars, no_am3], bip4_am3_initial)
bip5_am3_syn = synapse_var([no_bipolars, no_am3], bip5_am3_initial)
bip6_am3_syn = synapse_var([no_bipolars, no_am3], bip6_am3_initial)
bip7_am3_syn = synapse_var([no_bipolars, no_am3], bip7_am3_initial)
bip8_am3_syn = synapse_var([no_bipolars, no_am3], bip8_am3_initial)
bip9_am3_syn = synapse_var([no_bipolars, no_am3], bip9_am3_initial)
bip10_am3_syn = synapse_var([no_bipolars, no_am3], bip10_am3_initial)
bip11_am3_syn = synapse_var([no_bipolars, no_am3], bip11_am3_initial)
bip12_am3_syn = synapse_var([no_bipolars, no_am3], bip12_am3_initial)
bip13_am3_syn = synapse_var([no_bipolars, no_am3], bip13_am3_initial)
bip14_am3_syn = synapse_var([no_bipolars, no_am3], bip14_am3_initial)
am1_gc_syn = ag_synapse_var([no_am1, no_gc], am1_gc_initial, am1_gc_train_initial)
am2_gc_syn = ag_synapse_var([no_am2, no_gc], am2_gc_initial, am2_gc_train_initial)
am3_gc_syn = ag_synapse_var([no_am3, no_gc], am3_gc_initial, am3_gc_train_initial)
am1_b3copy_syn = synapse_var([no_am1, no_bipolars], am1_b3copy_initial)
am1_b4copy_syn = synapse_var([no_am1, no_bipolars], am1_b4copy_initial)
am1_b5copy_syn = synapse_var([no_am1, no_bipolars], am1_b5copy_initial)
am1_b6copy_syn = synapse_var([no_am1, no_bipolars], am1_b6copy_initial)
am1_b7copy_syn = synapse_var([no_am1, no_bipolars], am1_b7copy_initial)
am1_b8copy_syn = synapse_var([no_am1, no_bipolars], am1_b8copy_initial)
am1_b9copy_syn = synapse_var([no_am1, no_bipolars], am1_b9copy_initial)
b1_bias_initial=b1b*np.ones([no_bipolars, 1])
b1_bias_initial=b1_bias_initial.astype(float32)
b2_bias_initial=b2b*np.ones([no_bipolars, 1])
b2_bias_initial=b2_bias_initial.astype(float32)
b3_bias_initial=b3b*np.ones([no_bipolars, 1])
b3_bias_initial=b3_bias_initial.astype(float32)
b4_bias_initial=b4b*np.ones([no_bipolars, 1])
b4_bias_initial=b4_bias_initial.astype(float32)
b5_bias_initial=b5b*np.ones([no_bipolars, 1])
b5_bias_initial=b5_bias_initial.astype(float32)
b6_bias_initial=b6b*np.ones([no_bipolars, 1])
b6_bias_initial=b6_bias_initial.astype(float32)
b7_bias_initial=b7b*np.ones([no_bipolars, 1])
b7_bias_initial=b7_bias_initial.astype(float32)
b8_bias_initial=b8b*np.ones([no_bipolars, 1])
b8_bias_initial=b8_bias_initial.astype(float32)
b9_bias_initial=b9b*np.ones([no_bipolars, 1])
b9_bias_initial=b9_bias_initial.astype(float32)
b10_bias_initial=b10b*np.ones([no_bipolars, 1])
b10_bias_initial=b10_bias_initial.astype(float32)
b11_bias_initial=b11b*np.ones([no_bipolars, 1])
b11_bias_initial=b11_bias_initial.astype(float32)
b12_bias_initial=b12b*np.ones([no_bipolars, 1])
b12_bias_initial=b12_bias_initial.astype(float32)
b13_bias_initial=b13b*np.ones([no_bipolars, 1])
b13_bias_initial=b13_bias_initial.astype(float32)
b14_bias_initial=b14b*np.ones([no_bipolars, 1])
b14_bias_initial=b14_bias_initial.astype(float32)
b1_bias=bias_var([no_bipolars, 1], b1_bias_initial)
b2_bias=bias_var([no_bipolars, 1], b2_bias_initial)
b3_bias=bias_var([no_bipolars, 1], b3_bias_initial)
b4_bias=bias_var([no_bipolars, 1], b4_bias_initial)
b5_bias=bias_var([no_bipolars, 1], b5_bias_initial)
b6_bias=bias_var([no_bipolars, 1], b6_bias_initial)
b7_bias=bias_var([no_bipolars, 1], b7_bias_initial)
b8_bias=bias_var([no_bipolars, 1], b8_bias_initial)
b9_bias=bias_var([no_bipolars, 1], b9_bias_initial)
b10_bias=bias_var([no_bipolars, 1], b10_bias_initial)
b11_bias=bias_var([no_bipolars, 1], b11_bias_initial)
b12_bias=bias_var([no_bipolars, 1], b12_bias_initial)
b13_bias=bias_var([no_bipolars, 1], b13_bias_initial)
b14_bias=bias_var([no_bipolars, 1], b14_bias_initial)
am1_bias=-0.0*np.ones([no_am1, 1])
am1_bias=am1_bias.astype(float32)
am2_bias=-1.0*np.ones([no_am2, 1])
am2_bias=am2_bias.astype(float32)
am3_bias=-1.0*np.ones([no_am3, 1])
am3_bias=am3_bias.astype(float32)
gc_bias = np.array([[-0.5], [-0.5], [-0.5], [-0.5], [-0.5], [-1.7], [-1.7], [0.0]])
gc_bias=gc_bias.astype(float32)
# + deletable=true editable=true
dur=data_duration
batchsize=32
no_bip=no_bipolars
batchsize_ = tf.placeholder("int32", name="batch_size")
@tf.function
def biplayer(b_input, b_bias, bip_gc_syn, no_bip, no_gc, batchsize, dur):
b_bias_expand=tf.broadcast_to(tf.reshape(b_bias, [no_bipolars, 1, 1]), [no_bipolars, batchsize_, dur], name="bro1")
b_bias_add=tf.add(b_input, b_bias_expand)
b_relu=tf.nn.relu(b_bias_add)
bip_layer_expand=tf.broadcast_to(tf.reshape(b_relu, [no_bip, batchsize_, 1, dur]), [no_bip, batchsize_, no_gc, dur], name="bro2")
bg_syn_expand=tf.broadcast_to(tf.reshape(tf.abs(bip_gc_syn), [no_bip, 1, no_gc, 1]), [no_bip, batchsize_, no_gc, dur], name="bro3")
bg_mult=tf.math.multiply(bip_layer_expand, bg_syn_expand)
bg_sum=tf.reduce_sum(bg_mult, 0)
return b_relu, bg_sum
@tf.function
def linear_biplayer(b_input, b_bias, bip_gc_syn, no_bip, no_gc, batchsize, dur):
b_bias_expand=tf.broadcast_to(tf.reshape(b_bias, [no_bipolars, 1, 1]), [no_bipolars, batchsize_, dur], name="bro1")
b_bias_add=tf.add(b_input, b_bias_expand)
b_relu=b_bias_add
bip_layer_expand=tf.broadcast_to(tf.reshape(b_relu, [no_bip, batchsize_, 1, dur]), [no_bip, batchsize_, no_gc, dur], name="bro2")
bg_syn_expand=tf.broadcast_to(tf.reshape(tf.abs(bip_gc_syn), [no_bip, 1, no_gc, 1]), [no_bip, batchsize_, no_gc, dur], name="bro3")
bg_mult=tf.math.multiply(bip_layer_expand, bg_syn_expand)
bg_sum=tf.reduce_sum(bg_mult, 0)
return b_relu, bg_sum
b1_relu, b1g_sum = linear_biplayer(input_bip1_, b1_bias, bip1_gc_syn, no_bip, no_gc, batchsize_, dur)
b2_relu, b2g_sum = biplayer(input_bip2_, b2_bias, bip2_gc_syn, no_bip, no_gc, batchsize_, dur)
b3_relu, b3g_sum = biplayer(input_bip3_, b3_bias, bip3_gc_syn, no_bip, no_gc, batchsize_, dur)
b4_relu, b4g_sum = biplayer(input_bip4_, b4_bias, bip4_gc_syn, no_bip, no_gc, batchsize_, dur)
b5_relu, b5g_sum = biplayer(input_bip5_, b5_bias, bip5_gc_syn, no_bip, no_gc, batchsize_, dur)
b6_relu, b6g_sum = biplayer(input_bip6_, b6_bias, bip6_gc_syn, no_bip, no_gc, batchsize_, dur)
b7_relu, b7g_sum = biplayer(input_bip7_, b7_bias, bip7_gc_syn, no_bip, no_gc, batchsize_, dur)
b8_relu, b8g_sum = biplayer(input_bip8_, b8_bias, bip8_gc_syn, no_bip, no_gc, batchsize_, dur)
b9_relu, b9g_sum = biplayer(input_bip9_, b9_bias, bip9_gc_syn, no_bip, no_gc, batchsize_, dur)
b10_relu, b10g_sum = biplayer(input_bip10_, b10_bias, bip10_gc_syn, no_bip, no_gc, batchsize_, dur)
b11_relu, b11g_sum = biplayer(input_bip11_, b11_bias, bip11_gc_syn, no_bip, no_gc, batchsize_, dur)
b12_relu, b12g_sum = biplayer(input_bip12_, b12_bias, bip12_gc_syn, no_bip, no_gc, batchsize_, dur)
b13_relu, b13g_sum = biplayer(input_bip13_, b13_bias, bip13_gc_syn, no_bip, no_gc, batchsize_, dur)
b14_relu, b14g_sum = biplayer(input_bip14_, b14_bias, bip14_gc_syn, no_bip, no_gc, batchsize_, dur)
@tf.function
def bip_to_am_input(b_relu, bip_am_syn, no_bip, no_am, batchsize, dur):
bip_layer_am_expand=tf.broadcast_to(tf.reshape(b_relu, [no_bip, batchsize, 1, dur]), [no_bip, batchsize, no_am, dur], name="bro10")
ba_syn_expand = tf.broadcast_to(tf.reshape(tf.abs(bip_am_syn), [no_bip, 1, no_am, 1]), [no_bip, batchsize, no_am, dur], name="bro11")
ba_mult = tf.math.multiply(bip_layer_am_expand, ba_syn_expand)
ba_sum = tf.reduce_sum(ba_mult, 0)
return ba_sum
b4a1_sum = bip_to_am_input(b4_relu, bip4_am1_syn, no_bip, no_am1, batchsize_, dur)
b7a1_sum = bip_to_am_input(b7_relu, bip7_am1_syn, no_bip, no_am1, batchsize_, dur)
b10a2_sum = bip_to_am_input(b10_relu, bip10_am2_syn, no_bip, no_am2, batchsize_, dur)
b11a3_sum = bip_to_am_input(b11_relu, bip11_am3_syn, no_bip, no_am3, batchsize_, dur)
am1_activation = tf.add_n([b4a1_sum, b7a1_sum])
am2_activation = tf.add_n([b10a2_sum])
am3_activation = tf.add_n([b11a3_sum])
am1_bias_expand = tf.broadcast_to(am1_bias, [batchsize_, no_am1, dur], name="bro20")
am2_bias_expand = tf.broadcast_to(am2_bias, [batchsize_, no_am2, dur], name="bro21")
am3_bias_expand = tf.broadcast_to(am3_bias, [batchsize_, no_am3, dur], name="bro21")
am1_bias_add = tf.add(am1_activation, am1_bias_expand)
am2_bias_add = tf.add(am2_activation, am2_bias_expand)
am3_bias_add = tf.add(am3_activation, am3_bias_expand)
am1_output = tf.nn.relu(am1_bias_add)
am2_output = tf.nn.relu(am2_bias_add)
am3_output = tf.nn.relu(am3_bias_add)
am1_reshape = tf.reshape(am1_output, [batchsize_, no_am1, 1, dur])
am1_expand=tf.broadcast_to(am1_reshape, [batchsize_, no_am1, no_gc, dur], name="bro22")
am1g_syn_expand=tf.broadcast_to(tf.reshape(tf.abs(am1_gc_syn), [1, no_am1, no_gc, 1]), [batchsize_, no_am1, no_gc, dur], name="bro23")
am1g_mult=tf.math.multiply(am1_expand, am1g_syn_expand)
am1g_sum=tf.reduce_sum(am1g_mult, 1)
am2_reshape = tf.reshape(am2_output, [batchsize_, no_am2, 1, dur])
am2_expand=tf.broadcast_to(am2_reshape, [batchsize_, no_am2, no_gc, dur], name="bro24")
am2g_syn_expand=tf.broadcast_to(tf.reshape(tf.abs(am2_gc_syn), [1, no_am2, no_gc, 1]), [batchsize_, no_am2, no_gc, dur], name="bro25")
am2g_mult=tf.math.multiply(am2_expand, am2g_syn_expand)
am2g_sum=tf.reduce_sum(am2g_mult, 1)
am3_reshape = tf.reshape(am3_output, [batchsize_, no_am3, 1, dur])
am3_expand=tf.broadcast_to(am3_reshape, [batchsize_, no_am3, no_gc, dur], name="bro24")
am3g_syn_expand=tf.broadcast_to(tf.reshape(tf.abs(am3_gc_syn), [1, no_am3, no_gc, 1]), [batchsize_, no_am3, no_gc, dur], name="bro25")
am3g_mult=tf.math.multiply(am3_expand, am3g_syn_expand)
am3g_sum=tf.reduce_sum(am3g_mult, 1)
am1_bcopy_expand=tf.broadcast_to(am1_reshape, [batchsize_, no_am1, no_bip, dur], name="bro26")
@tf.function
def biplayer_copy_input(b_input,am_bcopy_expand, am_bcopy_syn, b_bias, bip_copy_gc_syn, no_bip, no_am, no_gc, batchsize, dur):
ambcopy_syn_expand=tf.broadcast_to(tf.reshape(tf.abs(am_bcopy_syn), [1, no_am, no_bip, 1]), [batchsize, no_am, no_bip, dur], name="bro33")
ambcopy_mult=tf.math.multiply(am_bcopy_expand, ambcopy_syn_expand)
ambcopy_sum1=tf.squeeze(tf.reduce_sum(ambcopy_mult, 1))
ambcopy_sum=tf.transpose(ambcopy_sum1, [1, 0, 2])
b_bias_expand=tf.broadcast_to(tf.reshape(b_bias, [no_bipolars, 1, 1]), [no_bipolars, batchsize, dur], name="bro1")
b_bias_add=tf.add_n([b_input,-1.0*ambcopy_sum, b_bias_expand])
b_relu=tf.nn.relu(b_bias_add)
bip_layer_expand=tf.broadcast_to(tf.reshape(b_relu, [no_bip, batchsize_, 1, dur]), [no_bip, batchsize, no_gc, dur], name="bro2")
bg_syn_expand=tf.broadcast_to(tf.reshape(tf.abs(bip_copy_gc_syn), [no_bip, 1, no_gc, 1]), [no_bip, batchsize, no_gc, dur], name="bro3")
bg_mult=tf.math.multiply(bip_layer_expand, bg_syn_expand)
bg_sum=tf.reduce_sum(bg_mult, 0)
return bg_sum
b3copyg_sum = biplayer_copy_input(input_bip3_,am1_bcopy_expand, am1_b3copy_syn, b3_bias, bip3_copy_gc_syn, no_bip, no_am1, no_gc, batchsize_, dur)
b4copyg_sum = biplayer_copy_input(input_bip4_,am1_bcopy_expand, am1_b4copy_syn, b4_bias, bip4_copy_gc_syn, no_bip, no_am1, no_gc, batchsize_, dur)
b5copyg_sum = biplayer_copy_input(input_bip5_,am1_bcopy_expand, am1_b5copy_syn, b5_bias, bip5_copy_gc_syn, no_bip, no_am1, no_gc, batchsize_, dur)
b6copyg_sum = biplayer_copy_input(input_bip6_,am1_bcopy_expand, am1_b6copy_syn, b6_bias, bip6_copy_gc_syn, no_bip, no_am1, no_gc, batchsize_, dur)
b7copyg_sum = biplayer_copy_input(input_bip7_,am1_bcopy_expand, am1_b7copy_syn, b7_bias, bip7_copy_gc_syn, no_bip, no_am1, no_gc, batchsize_, dur)
b8copyg_sum = biplayer_copy_input(input_bip8_,am1_bcopy_expand, am1_b8copy_syn, b8_bias, bip8_copy_gc_syn, no_bip, no_am1, no_gc, batchsize_, dur)
b9copyg_sum = biplayer_copy_input(input_bip9_,am1_bcopy_expand, am1_b9copy_syn, b9_bias, bip9_copy_gc_syn, no_bip, no_am1, no_gc, batchsize_, dur)
gc_activation=tf.add_n([b3copyg_sum, b4copyg_sum, b5copyg_sum, b6copyg_sum, b7copyg_sum, b8copyg_sum, b9copyg_sum, b1g_sum, b2g_sum, b3g_sum, b4g_sum, b5g_sum, b6g_sum, b7g_sum, b8g_sum, b9g_sum, b10g_sum, b11g_sum, b12g_sum, b13g_sum, b14g_sum, -1.0*am1g_sum, -1.0*am2g_sum, -1.0*am3g_sum])
gc_bias_expand=tf.broadcast_to(gc_bias, [batchsize_, no_gc, dur])
gc_bias_add=tf.add(gc_activation, gc_bias_expand)
output1=tf.nn.relu(gc_bias_add)
# + deletable=true editable=true
outputfeedback1=tf.zeros([batchsize_, no_gc-1, dur])
feedbackx = np.linspace(0.0, 100.0, 500)
feedbackb=3.0
feedbacky=np.exp(-1.0*(feedbackx-feedbackx[200])/feedbackb)
feedbacky[0:200]=0.0
feedbacky=(np.reshape(feedbacky, [500, 1]))
feedbackfilt=np.reshape(feedbacky, [500, 1, 1, 1])
outforconv=tf.reshape(tf.slice(output1, [0, 5, 0], [batchsize_, 1, dur]), [batchsize_, dur, 1, 1])
padsize=250
paddedx=tf.pad(outforconv, [[0, 0], [padsize, 0], [0, 0], [0, 0]], 'CONSTANT')
outconv=tf.nn.conv2d(paddedx, feedbackfilt, strides=[1, 1, 1, 1], padding='SAME') #250 for movingdot and noise
outconv1=tf.reshape(outconv[:, np.round(padsize/2).astype(int):np.round(padsize/2).astype(int)+dur, 0, 0], [batchsize_, 1, dur])
outputfeedback=tf.nn.relu(tf.concat([outputfeedback1, outconv1], 1))
output=tf.nn.relu(tf.add(output1, -0.0*outputfeedback))
# + deletable=true editable=true
sess=tf.Session()
sess.run(tf.global_variables_initializer())
# + deletable=true editable=true
oo_output=np.zeros([576,8, 5000])
for bbatch in range(36):
startind=(bbatch)*16
endind=(bbatch+1)*16
fd={batchsize_: 16, input_bip1_: input_bip1_train[:, startind:endind, :], input_bip2_: input_bip2_train[:, startind:endind, :], input_bip3_: input_bip3_train[:, startind:endind, :], input_bip4_: input_bip4_train[:, startind:endind, :], input_bip5_: input_bip5_train[:, startind:endind, :], input_bip6_: input_bip6_train[:, startind:endind, :], input_bip7_: input_bip7_train[:, startind:endind, :], input_bip8_: input_bip8_train[:, startind:endind, :], input_bip9_: input_bip9_train[:, startind:endind, :], input_bip10_: input_bip10_train[:, startind:endind, :], input_bip11_: input_bip11_train[:, startind:endind, :], input_bip12_: input_bip12_train[:, startind:endind, :], input_bip13_: input_bip13_train[:, startind:endind, :], input_bip14_: input_bip14_train[:, startind:endind, :]}
[o_output, o_output1, o_b1_relu, o_b3_relu, o_b4_relu, o_b5_relu, o_b7_relu, o_b10_relu, o_b11_relu, o_am1_output, o_am2_output, o_am3_output, o_b1g_sum, o_b3g_sum, o_b4g_sum, o_b5g_sum, o_b7g_sum, o_b10a2_sum, o_b4copyg_sum, o_b7copyg_sum, o_am1g_sum, o_am1_reshape, o_am1_expand, o_am1g_mult, o_am2g_sum, o_am3g_sum, o_gc_activation] = sess.run([output, output1, b1_relu, b3_relu, b4_relu, b5_relu, b7_relu, b10_relu, b11_relu, am1_output, am2_output, am3_output, b1g_sum, b3g_sum, b4g_sum, b5g_sum, b7g_sum, b10a2_sum, b4copyg_sum, b7copyg_sum, am1g_sum, am1_reshape, am1_expand, am1g_mult, am2g_sum, am3g_sum, gc_activation], feed_dict=fd, options = run_opts)
oo_output[startind:endind,:, :]=o_output
# + deletable=true editable=true
db={}
db['y_train']=oo_output
db['b1_relu']=o_b1_relu
db['b4_relu']=o_b4_relu
db['b5_relu']=o_b5_relu
db['b7_relu']=o_b7_relu
db['b10_relu']=o_b10_relu
db['b11_relu']=o_b11_relu
db['b1g_sum']=o_b1g_sum
db['b4g_sum']=o_b4g_sum
db['b5g_sum']=o_b5g_sum
db['b7g_sum']=o_b7g_sum
db['b10a2_sum']=o_b10a2_sum
db['b4copyg_sum']=o_b4copyg_sum
db['b7copyg_sum']=o_b7copyg_sum
db['am1g_sum']=o_am1g_sum
db['am1_expand']=o_am1_expand
db['am1_reshape']=o_am1_reshape
db['am1g_mult']=o_am1g_mult
db['am2g_sum']=o_am2g_sum
db['am1_output']=o_am1_output
db['am2_output']=o_am2_output
db['gc_activation']=o_gc_activation
db['bip1_gc_syn']=bip1_gc_initial
db['bip2_gc_syn']=bip2_gc_initial
db['bip3_gc_syn']=bip3_gc_initial
db['bip4_gc_syn']=bip4_gc_initial
db['bip5_gc_syn']=bip5_gc_initial
db['bip6_gc_syn']=bip6_gc_initial
db['bip7_gc_syn']=bip7_gc_initial
db['bip8_gc_syn']=bip8_gc_initial
db['bip9_gc_syn']=bip9_gc_initial
db['bip10_gc_syn']=bip10_gc_initial
db['bip11_gc_syn']=bip11_gc_initial
db['bip12_gc_syn']=bip12_gc_initial
db['bip13_gc_syn']=bip13_gc_initial
db['bip14_gc_syn']=bip14_gc_initial
db['bip1_am1_syn']=bip1_am1_initial
db['bip2_am1_syn']=bip2_am1_initial
db['bip3_am1_syn']=bip3_am1_initial
db['bip4_am1_syn']=bip4_am1_initial
db['bip5_am1_syn']=bip5_am1_initial
db['bip6_am1_syn']=bip6_am1_initial
db['bip7_am1_syn']=bip7_am1_initial
db['bip8_am1_syn']=bip8_am1_initial
db['bip9_am1_syn']=bip9_am1_initial
db['bip10_am1_syn']=bip10_am1_initial
db['bip11_am1_syn']=bip11_am1_initial
db['bip12_am1_syn']=bip12_am1_initial
db['bip13_am1_syn']=bip13_am1_initial
db['bip14_am1_syn']=bip14_am1_initial
db['bip1_am2_syn']=bip1_am2_initial
db['bip2_am2_syn']=bip2_am2_initial
db['bip3_am2_syn']=bip3_am2_initial
db['bip4_am2_syn']=bip4_am2_initial
db['bip5_am2_syn']=bip5_am2_initial
db['bip6_am2_syn']=bip6_am2_initial
db['bip7_am2_syn']=bip7_am2_initial
db['bip8_am2_syn']=bip8_am2_initial
db['bip9_am2_syn']=bip9_am2_initial
db['bip10_am2_syn']=bip10_am2_initial
db['bip11_am2_syn']=bip11_am2_initial
db['bip12_am2_syn']=bip12_am2_initial
db['bip13_am2_syn']=bip13_am2_initial
db['bip14_am2_syn']=bip14_am2_initial
db['am1_b3copy_syn']=am1_b3copy_initial
db['am1_b4copy_syn']=am1_b4copy_initial
db['am1_b5copy_syn']=am1_b5copy_initial
db['am1_b6copy_syn']=am1_b6copy_initial
db['am1_b7copy_syn']=am1_b7copy_initial
db['am1_b8copy_syn']=am1_b8copy_initial
db['am1_b9copy_syn']=am1_b9copy_initial
db['am1_gc_syn']=am1_gc_initial
db['am2_gc_syn']=am2_gc_initial
db['b1_bias']=b1_bias_initial
db['b2_bias']=b2_bias_initial
db['b3_bias']=b3_bias_initial
db['b4_bias']=b4_bias_initial
db['b5_bias']=b5_bias_initial
db['b6_bias']=b6_bias_initial
db['b7_bias']=b7_bias_initial
db['b8_bias']=b8_bias_initial
db['b9_bias']=b9_bias_initial
db['b10_bias']=b10_bias_initial
db['b11_bias']=b11_bias_initial
db['b12_bias']=b12_bias_initial
db['b13_bias']=b13_bias_initial
db['b14_bias']=b14_bias_initial
sio.savemat(wheretosave, db)
print(wheretosave)
|
Figure 5 Simulations/Oracles/KDD_XCell_Oracle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: base
# language: python
# name: base
# ---
# +
# %load_ext autoreload
# %autoreload 2
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# +
import sys
from itertools import groupby
import numpy as np
import cv2
from tqdm.notebook import tqdm
tqdm.pandas()
import pandas as pd
import os
from glob import glob
from multiprocessing import Pool
import matplotlib.pyplot as plt
# import cupy as cp
import ast
from pathlib import Path
import pickle5 as pickle
import torch
import shutil
import sys
sys.path.append("../src/")
sys.path.append("../yolov5/")
import util
from joblib import Parallel, delayed
from IPython.display import display, HTML
from sahi.model import Yolov5DetectionModel
from sahi.utils.cv import read_image
from sahi.utils.file import download_from_url
from sahi.predict import get_prediction, get_sliced_prediction, predict
from ensemble_boxes import weighted_boxes_fusion
from IPython.display import Image
from matplotlib import animation, rc
rc('animation', html='jshtml')
# -
CONF = 0
IOU = 0.4
AUGMENT = True # True
SIZE = int(1280 * 1.3) # 1280 * 1.3
#USE_CLACHE = True
USE_SAHI = False
# # Load Data
INPUT_DIR = Path("../../data/tensorflow-great-barrier-reef/")
df = pd.read_csv(INPUT_DIR / "train.csv")
#folds = pd.read_csv("../input/train_folds_balanced.csv") #_CPF
#df = pd.merge(df, folds[["video_id","video_frame","fold"]], on=["video_id",'video_frame'])
df["fold"] = df["video_id"]
df.shape
data_param = {'root_dir':INPUT_DIR,'label_dir':INPUT_DIR / "labels"}
df = df.progress_apply(lambda x: util.get_path(x, data_param, infer=True), axis=1)
df['annotations'] = df['annotations'].progress_apply(lambda x: ast.literal_eval(x))
df["real_bbox"] = df["annotations"].apply(lambda annots: [list(annot.values()) for annot in annots])
df['num_bbox'] = df['annotations'].progress_apply(lambda x: len(x))
data = (df.num_bbox>0).value_counts(normalize=True)*100
print(f"No BBox: {data[0]:0.2f}% | With BBox: {data[1]:0.2f}%")
# COTS per frame
df.groupby("fold").apply(lambda df: df["num_bbox"].sum() / df.shape[0])
# # Model
meta_params = {}
#model_versions = ['0123_yolov5s', "0124_yolov5s_f0","0124_yolov5s_f1","0125_yolov5s_f2","0125_yolov5s_f3"]
#model_versions = ['0125_yolov5s_f0_cpf', "0125_yolov5s_f1_cpf","0125_yolov5s_f2_cpf","0125_yolov5s_f3_cpf","0125_yolov5s_f4_cpf"]
#model_versions = ['0125_yolov5l_wpre_f0', "0125_yolov5l_wpre_f1","0125_yolov5l_wpre_f2","0126_yolov5l_wpre_f3","0125_yolov5l_wpre_f4"]
model_versions = ['0127_yolov5l_wpre_v0_FP', "0127_yolov5l_wpre_v1_FP","0127_yolov5l_wpre_v2_FP"]
for model_version in model_versions:#, '0110_yolov5l_fold0_new_hyp_remove_noaug']:
model_folder = Path(f"../output/{model_version}/")
try:
params_path = model_folder / "config" / "params.pkl"
params = pickle.load(open(params_path, 'rb'))
except:
params_path = model_folder / "config" / "params.yaml"
params = util.load_yaml(params_path)
for key, val in params.items():
if "dir" in key or "path" in key or "file" in key:
params[key] = Path(val)
meta_params[model_version] = params
fig, axes = plt.subplots(1, len(meta_params),figsize=(5*len(meta_params), 5))
target_score = {}
for i,(model_version, params) in enumerate(meta_params.items()):
if params["tools"] == "mmdetection":
continue
logging_dir = params["ckpt_path"].parent.parent
res_df = pd.read_csv(logging_dir / "results.csv")
res_df[[' metrics/score',' metrics/precision', ' metrics/recall', ' metrics/mAP_0.5', 'metrics/mAP_0.5:0.95', ]].plot(ax=axes[i])
axes[i].legend()
axes[i].set_title(f"{model_version} score")
target_score[model_version] = res_df[' metrics/score'].max()
plt.show()
target_score
models = {}
for model_version, params in meta_params.items():
if params['tools'] == "yolov5":
params["repo"] = Path("../yolov5/").resolve()
else:
ckp = glob(str(params['output_dir']) + "/" + "best*")[0]
params['ckpt_path'] = ckp
params["augment"] = AUGMENT
params["conf"] = CONF
params["iou"] = IOU
if USE_SAHI:
model = Yolov5DetectionModel(
model_path=str(params['ckpt_path']),
confidence_threshold=CONF,
image_size = params['img_size'],
device="cuda:0", # or 'cuda:0'
)
else:
if params['tools'] == "yolov5":
model = util.load_model(params)
device = torch.device("cuda")
model.to(device)
elif params['tools'] == "mmdetection":
model = init_detector(str(params['cfg_dir'] / "config.py"), ckp, device='cuda:0')
models[model_version] = model
# # Prediction
meta_params[model_version]["fold"][0]
df["pred"] = None
for model_version in meta_params.keys():
fold = meta_params[model_version]["fold"][0]
df["pred_" + model_version + f"_fold{fold}"] = None
df.columns
SH = 640
SW = 640
OHR = 0.2
OWR = 0.2
PMT = 0.2
clahe = cv2.createCLAHE(clipLimit=5, tileGridSize=(14, 14))
for idx in tqdm(range(len(df))):
row = df.loc[idx]
target_fold = row['fold']
img_path = row["image_path"]
img = cv2.imread(str(img_path))
for model_version, model in models.items():
model_fold = meta_params[model_version]["fold"][0]
if model_fold != target_fold:
continue
USE_CLACHE = meta_params[model_version].get("use_clahe", True)
img2 = img.copy()
if USE_CLACHE:
for i in range(3):
img2[:, :, i] = clahe.apply((img2[:, :, i]))
if meta_params[model_version]["tools"] == "yolov5":
img2 = img2[...,::-1]
if USE_SAHI:
result_sliced = get_sliced_prediction(
img2,
model,
slice_height = SH,
slice_width = SW,
overlap_height_ratio = OHR,
overlap_width_ratio = OWR,
postprocess_match_threshold = PMT,
verbose = False)
object_prediction_list = result_sliced.object_prediction_list
confs = [obj_pred.score.value for obj_pred in object_prediction_list]
pred_bbox = np.array([obj_pred.bbox.to_coco_bbox() for obj_pred in object_prediction_list])
else:
pred_bbox, confs = util.predict(model, img2, size=SIZE, augment=AUGMENT, use_sahi=USE_SAHI)
elif meta_params[model_version]["tools"] == "mmdetection":
result = inference_detector(model, img2)
pred_bbox = result[0][:,:4]
pred_bbox[:,2:] = pred_bbox[:,2:] - pred_bbox[:,:2]
pred_confs = result[0][:,4]
confs = pred_confs.tolist()
df.at[idx, "pred_" + model_version + f"_fold{target_fold}"] = [ [conf] + pred_bbox[i].tolist() for i, conf in enumerate(confs)]
df.at[idx, "pred"] = df.at[idx, "pred_" + model_version + f"_fold{target_fold}"]
#save it
util.save_pickle(df, "pred_all_large_wpre_video_FP.pkl")
# # Pick Conf Thresh
def run_wbf(bboxes, confs, image_size=1280, iou_thr=0.4, skip_box_thr=0.7, weights=None, conf_type='avg'):
if len(bboxes) == 1:
return bboxes[0], confs[0], []
boxes = [bbox / (image_size-1) for bbox in bboxes]
scores = [conf for conf in confs]
labels = [np.ones(len(conf)) for conf in confs]
boxes, scores, labels = weighted_boxes_fusion(boxes, scores, labels, weights=weights, iou_thr=iou_thr, skip_box_thr=skip_box_thr,conf_type=conf_type)
boxes = boxes * (image_size-1)
return boxes, scores, labels
pred_col = "pred"
gt_bboxs_list = []
for idx in range(len(df)):
row = df.iloc[idx]
if row[pred_col] is None:
continue
gt_bboxs = row["real_bbox"]
gt_bboxs_list.append(np.array(gt_bboxs))
#conf_ths = [10, 15, 20, 25, 30, 35, 40, 45, 50, 55]
#conf_ths = [12, 16, 18, 20, 22, 24]
conf_ths = [6, 8, 10, 12, 14]
#conf_ths = [20, 24, 26, 28, 30]
#conf_ths = [30, 32, 35, 38, 45, 50, 55]
#conf_ths = [32, 34, 40, 45, 50]
iou_ths = np.arange(0.3, 0.85, 0.05)
for conf_th in conf_ths:
conf_th /= 100
prd_bboxs_list = []
for index in df.index:
if df.loc[index, "pred"] is None:
continue
prd_bboxs = df.loc[index, pred_col]
prd_bboxs = [p for p in prd_bboxs if p[0] >= conf_th]
#prd_bboxs = [[confis[i]] + bboxes[i] for i in range(len(prd_bboxs)) if prd_bboxs[i][0] >= conf_th]
prd_bboxs_list.append(np.array(prd_bboxs))
tps, fps, fns = util.calc_f2_score(gt_bboxs_list, prd_bboxs_list, verbose=False)
#df["tp"], df["fp"], df["fn"] = tps, fps, fns
precision = util.f_beta(np.sum(tps), np.sum(fps), np.sum(fns), beta=0)
recall = util.f_beta(np.sum(tps), np.sum(fps), np.sum(fns), beta=100)
score = util.f_beta(np.sum(tps), np.sum(fps), np.sum(fns), beta=2)
#score_only_bbox = util.f_beta(*df.query("has_annotations")[["tp", "fp", "fn"]].sum(axis=0), beta=2)
print(f"conf_th {conf_th:.2f}, pred: f2 {score:.4f}, precision {precision:.2f}, recall {recall:.2f} ")
# # Tracking
# +
from norfair import Detection, Tracker
# Helper to convert bbox in format [x_min, y_min, x_max, y_max, score] to norfair.Detection class
def to_norfair(detects, frame_id):
result = []
for x_min, y_min, x_max, y_max, score in detects:
xc, yc = (x_min + x_max) / 2, (y_min + y_max) / 2
w, h = x_max - x_min, y_max - y_min
result.append(Detection(points=np.array([xc, yc]), scores=np.array([score]), data=np.array([w, h, frame_id])))
return result
def euclidean_distance(detection, tracked_object):
return np.linalg.norm(detection.points - tracked_object.estimate)
def tracking_function(tracker, frame_id, bboxes, scores, best_conf):
detects = []
predictions = []
if len(scores)>0:
for i in range(len(bboxes)):
# remember to check
if scores[i] <= best_conf:
continue
box = bboxes[i]
score = scores[i]
x_min = int(box[0])
y_min = int(box[1])
bbox_width = int(box[2])
bbox_height = int(box[3])
detects.append([x_min, y_min, x_min+bbox_width, y_min+bbox_height, score])
predictions.append('{:.2f} {} {} {} {}'.format(score, x_min, y_min, bbox_width, bbox_height))
# print(predictions[:-1])
# Update tracks using detects from current frame
tracked_objects = tracker.update(detections=to_norfair(detects, frame_id))
for tobj in tracked_objects:
bbox_width, bbox_height, last_detected_frame_id = tobj.last_detection.data
if last_detected_frame_id == frame_id: # Skip objects that were detected on current frame
continue
# Add objects that have no detections on current frame to predictions
xc, yc = tobj.estimate[0]
x_min, y_min = int(round(xc - bbox_width / 2)), int(round(yc - bbox_height / 2))
score = tobj.last_detection.scores[0]
predictions.append('{:.2f} {} {} {} {}'.format(score, x_min, y_min, bbox_width, bbox_height))
return predictions
# -
tracker = Tracker(
distance_function=euclidean_distance,
distance_threshold=30,
hit_inertia_min=3,
hit_inertia_max=6,
initialization_delay=1,
)
# +
BEST_CONF = 0.12
frame_id = 0
prd_bboxs_tracking_list = []
for idx in range(len(df)):
row = df.iloc[idx]
prd_bboxs = row["pred"]
if prd_bboxs is None:
continue
bboxes, confis = [p[1:] for p in prd_bboxs], [p[0] for p in prd_bboxs]
predictions = tracking_function(tracker, frame_id, bboxes, confis, BEST_CONF)
prd_bboxs_tracking = [[float(p) for p in pred.split(" ")] for pred in predictions]
prd_bboxs_tracking_list.append(np.array(prd_bboxs_tracking))
frame_id += 1
tps, fps, fns = util.calc_f2_score(gt_bboxs_list, prd_bboxs_list, verbose=False)
#df["tp"], df["fp"], df["fn"] = tps, fps, fns
precision = util.f_beta(np.sum(tps), np.sum(fps), np.sum(fns), beta=0)
recall = util.f_beta(np.sum(tps), np.sum(fps), np.sum(fns), beta=100)
score = util.f_beta(np.sum(tps), np.sum(fps), np.sum(fns), beta=2)
print(f"Adding tracking, pred: f2 {score:.4f}, precision {precision:.2f}, recall {recall:.2f}")
# -
# # check image
|
notebook/5fold-infer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.1 64-bit
# language: python
# name: python3
# ---
# # Binary search tree (BST)
#
# ### Definition
#
# A binary search tree (BST), also called an ordered or sorted binary tree, is a rooted binary tree data structure whose internal nodes each store a key greater than all the keys in the node’s left subtree and less than those in its right subtree.
#
# A binary tree is a type of data structure for storing data such as numbers in an organized way. Binary search trees allow binary search for fast lookup, addition and removal of data items, and can be used to implement dynamic sets and lookup tables.
#
# The order of nodes in a BST means that each comparison skips about half of the remaining tree, so the whole lookup takes time proportional to the binary logarithm of the number of items stored in the tree. This is much better than the linear time required to find items by key in an (unsorted) array, but slower than the corresponding operations on hash tables. Several variants of the binary search tree have been studied.
#
# 
#
# +
# A class to store a BST node
class Node:
def __init__(self, data, left=None, right=None):
self.data = data
self.left = left
self.right = right
# Function to perform inorder traversal on the BST
def inorder(root):
if root is None:
return
inorder(root.left)
print(root.data, end=" ")
inorder(root.right)
# -
# ## Operations and time complexity
#
# Binary search trees support three main operations:
#
# 1. Searching: For searching element 1, we have to traverse all elements (in order 3, 2, 1). Therefore, searching in binary search tree has worst case complexity of O(n). In general, time complexity is O(h) where h is height of BST.
# 2. Creation: If you do not know all the elements of BST in advance (online algorithm) then you have to insert each of n elements one after another. If you are extremely unlucky, the complexity of insert is O(n) and thus it deteriorates to O(n^2).
# Notice that this situation is highly unlikely, but still possible.
# But you can still achieve O(nlog(n)) if you know all the elements in advance. You can sort them O(nlog(n)) and then insert the elements in the following order. Take the middle element and insert it as a root, then recursively do the same for both parts that are left. You will end up creating balanced BST, inserting n elements using log(n).
# 3. Insertion: For inserting element 0, it must be inserted as left child of 1. Therefore, we need to traverse all elements (in order 3, 2, 1) to insert 0 which has worst case complexity of O(n). In general, time complexity is O(h).
# 4. Deletion: For deletion of element 1, we have to traverse all elements to find 1 (in order 3, 2, 1). Therefore, deletion in binary tree has worst case complexity of O(n). In general, time complexity is O(h).
#
#
# The latter two possibly change the structural arrangement of the nodes in the tree, whereas the first one is a navigating and read-only operation. Other read-only operations are traversal, verification, etc.
# ## Insertion
#
# A new key is always inserted at the leaf. We start searching a key from the root until we hit a leaf node. Once a leaf node is found, the new node is added as a child of the leaf node.
#
#
# 
#
# +
# Recursive function to insert a key into a BST
# Space Complexity: O(h), no extra space is required (not counting [keys] array).
def insert(root, key):
# if the root is None, create a new node and return it
if root is None:
return Node(key)
# if the given key is less than the root node, recur for the left subtree
if key < root.data:
root.left = insert(root.left, key)
# if the given key is more than the root node, recur for the right subtree
else:
root.right = insert(root.right, key)
return root
keys = [21, 28, 14, 32, 25, 18, 11, 30, 19, 15, 23, 27]
root = None
for key in keys:
root = insert(root, key)
inorder(root)
# +
# Iterative function to insert a key into a BST
# Space Complexity: O(1), no extra space is required (not counting [keys] array).
def insert(root, key):
# Create a new Node containing
# the new element
newnode = Node(key)
# Pointer to start traversing from root
# and traverses downward path to search
# where the new node to be inserted
x = root
# Pointer y maintains the trailing
# pointer of x
y = None
while (x != None):
y = x
if (key < x.data):
x = x.left
else:
x = x.right
# If the root is None i.e the tree is
# empty. The new node is the root node
if (y == None):
y = newnode
# If the new key is less then the leaf node key
# Assign the new node to be its left child
elif (key < y.data):
y.left = newnode
# else assign the new node its
# right child
else:
y.right = newnode
# Returns the pointer where the
# new node is inserted
return y
keys = [21, 28, 14, 32, 25, 18, 11, 30, 19, 15, 23, 27]
root = None
started = False
for key in keys:
if (not started):
root = insert(root, key)
started = True
else:
insert(root, key)
inorder(root)
# -
# ## Searching
#
# Searching in a binary search tree for a specific key can be programmed recursively or iteratively.
#
# We begin by examining the root node. If the tree is empty, the key we are searching for does not exist in the tree.
# Otherwise, if the key equals that of the root, the search is successful and we return the node.
#
# If the key is less than that of the root, we search the left subtree. Similarly, if the key is greater than that of the root, we search the right subtree.
#
# This process is repeated until the key is found or the remaining subtree is empty. If the searched key is not found after a empty subtree is reached, then the key is not present in the tree.
#
#
# 
#
# This algorithm searches from the tree’s root to the leaf farthest from the root in the worst-case. The search operation takes time proportional to the tree’s height. On average, binary search trees with n nodes have O(log(n)) height. However, in the worst case, binary search trees can have O(n) height (for skewed trees where all the nodes except the leaf have one and only one child) when the unbalanced tree resembles a linked list.
#
# The space used by the call stack is also proportional to the tree’s height. The algorithm can be implemented iteratively to avoid use of extra space.
#
#
# +
# Recursive function to search in a given BST (BFS)
def search(root, key, parent):
# if the key is not present in the key
if root is None:
print('Key not found')
return
# if the key is found
if root.data == key:
if parent is None:
print(f'The node with key {key} is root node')
elif key < parent.data:
print('The given key is the left node of the node with key', parent.data)
else:
print('The given key is the right node of the node with key', parent.data)
return
# if the given key is less than the root node, recur for the left subtree;
# otherwise, recur for the right subtree
if key < root.data:
search(root.left, key, root)
else:
search(root.right, key, root)
search(root, 27, None)
# +
# Iterative function to search in a given BST (DFS)
def searchIterative(root, key):
# start with the root node
curr = root
# pointer to store the parent of the current node
parent = None
# traverse the tree and search for the key
while curr and curr.data != key:
# update the parent to the current node
parent = curr
# if the given key is less than the current node, go to the left subtree;
# otherwise, go to the right subtree
if key < curr.data:
curr = curr.left
else:
curr = curr.right
# if the key is not present in the key
if curr is None:
print('Key not found')
return
if parent is None:
print(f'The node with key {key} is root node')
elif key < parent.data:
print('The given key is the left node of the node with key', parent.data)
else:
print('The given key is the right node of the node with key', parent.data)
searchIterative(root, 27)
# -
# ## Deletion
#
# Here are three possible cases to consider deleting a node from BST:
#
# Case 1: Deleting a node with no children: remove the node from the tree.
#
# 
#
#
# Case 2: Deleting a node with two children: call the node to be deleted N. Do not delete N. Instead, choose either its inorder successor (The smallest number in the right subtree) node or its inorder predecessor node (The biggest number in the left subtree), R. Copy the value of R to N, then recursively call delete on R until reaching one of the first two cases. If we choose the inorder successor of a node (The smallest number in the right subtree), as the right subtree is not NULL (our present case is a node with 2 children), then its inorder successor is a node with the least value in its right subtree, which will have at a maximum of 1 subtree, so deleting it would fall in one of the first 2 cases.
#
# 
#
# Case 3: Deleting a node with one child: remove the node and replace it with its child.
#
# 
#
# Broadly speaking, nodes with children are harder to delete. As with all binary trees, a node’s inorder successor is its right subtree’s leftmost child, and a node’s inorder predecessor is the left subtree’s rightmost child. In either case, this node will have zero or one child. Delete it according to one of the two simpler cases above.
#
#
# +
# Helper function to find minimum value node in the subtree rooted at `curr`
def getMinimumKey(curr):
while curr.left:
curr = curr.left
return curr
# Function to delete a node from a BST
def deleteNode(root, key):
# pointer to store the parent of the current node
parent = None
# start with the root node
curr = root
# search key in the BST and set its parent pointer
while curr and curr.data != key:
# update the parent to the current node
parent = curr
# if the given key is less than the current node, go to the left subtree;
# otherwise, go to the right subtree
if key < curr.data:
curr = curr.left
else:
curr = curr.right
# return if the key is not found in the tree
if curr is None:
return root
# Case 1: node to be deleted has no children, i.e., it is a leaf node
if curr.left is None and curr.right is None:
# if the node to be deleted is not a root node, then set its
# parent left/right child to None
if curr != root:
if parent.left == curr:
parent.left = None
else:
parent.right = None
# if the tree has only a root node, set it to None
else:
root = None
# Case 2: node to be deleted has two children
elif curr.left and curr.right:
# find its inorder successor node
successor = getMinimumKey(curr.right)
# store successor value
val = successor.data
# recursively delete the successor. Note that the successor
# will have at most one child (right child)
deleteNode(root, successor.data)
# copy value of the successor to the current node
curr.data = val
# Case 3: node to be deleted has only one child
else:
# choose a child node
if curr.left:
child = curr.left
else:
child = curr.right
# if the node to be deleted is not a root node, set its parent
# to its child
if curr != root:
if curr == parent.left:
parent.left = child
else:
parent.right = child
# if the node to be deleted is a root node, then set the root to the child
else:
root = child
return root
root = deleteNode(root, 16)
inorder(root)
|
notes/data_structures/binary_search_tree/binary_search_tree.ipynb
|