code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" gradient={"editing": false, "id": "1fe2ae7d-6d6d-4216-bf57-e31a43ee07f8", "kernelId": ""} id="view-in-github"
# <a href="https://colab.research.google.com/github/bengsoon/lstm_lord_of_the_rings/blob/main/LOTR_LSTM_Character_Level_OneHot.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] gradient={"editing": false, "id": "3c8f437f-6d7a-404a-9173-88458dc55ec6", "kernelId": ""} id="sBQN-dtIGzPk"
# ## Creating a Language Model with LSTM using Lord of The Rings Corpus
# In this notebook, we will create a character-level language language model using LSTM using **one-hot encoding** on the input vectors
# + [markdown] gradient={"editing": false, "id": "d862b356-f5f3-461b-9c1b-1c86f24bc490", "kernelId": ""} id="BwvwLm9wnCwc"
# ### Imports
# + gradient={"editing": false, "id": "7552effa-3565-4e47-b1d4-80fac5b275be", "kernelId": ""}
# run this if you're running through paperspace
# !pip install -r requirements.txt
# + gradient={"editing": false, "id": "67c95a2c-4097-497f-9497-ab9626e8a18a", "kernelId": ""} id="6BvOxVto4pzg"
import tensorflow as tf
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
from tensorflow.keras.layers import Embedding, Input, LSTM, Flatten, Dense, Dropout
from tensorflow.keras.callbacks import LearningRateScheduler, ModelCheckpoint
from tensorflow.keras import Model
import numpy as np
from tensorflow.keras.models import load_model
from pprint import pprint as pp
from string import punctuation
import regex as re
import random
import os
from pathlib import Path
import math
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '2'
# + [markdown] gradient={"editing": false, "id": "78f9fc2f-3e38-418c-a7e3-230c49911bfe", "kernelId": ""} id="sMsUyeLHnFZC"
# ### Data Preprocessing & Pipeline
# + gradient={"editing": false, "id": "ae9df455-95a5-4c93-ade9-de60e919bc9d", "kernelId": ""} id="t4_puY1vH-Jv"
# get LOTR full text
# # !wget https://raw.githubusercontent.com/bengsoon/lstm_lord_of_the_rings/main/lotr_full.txt -P /content/drive/MyDrive/Colab\ Notebooks/LOTR_LSTM/data
# + [markdown] gradient={"editing": false, "id": "22a0e94d-2189-4c80-82b8-8de6416d8486", "kernelId": ""} id="s065iL9iBq_A"
# #### Loading Data
# + gradient={"editing": false, "id": "12cb95fe-b15a-4742-942c-c09ca8a76ac1", "kernelId": ""} id="SIYll8_1BycF"
path = Path(r"./")
# + colab={"base_uri": "https://localhost:8080/"} gradient={"editing": false, "id": "1ba16b44-906f-40c0-af14-8c7a06d39821", "kernelId": ""} id="8IVk7GxXn40r" outputId="a85132ac-4d97-4f97-e2c5-a7b360258a07"
with open(path/ "data/lotr_full.txt", "r", encoding="utf-8") as f:
text = f.read()
print(text[:1000])
# + colab={"base_uri": "https://localhost:8080/"} gradient={"editing": false, "id": "44cb50da-b565-4b39-9a25-c8b74adc9a31", "kernelId": ""} id="RjEFSBDEofqz" outputId="1d9943ee-adda-456b-c8a0-4e078493d337"
print(f"Corpus length: {int(len(text)) / 1000 } K characters")
# + [markdown] gradient={"editing": false, "id": "45b89d55-3d3b-406f-b5fb-77ee4fbde7b6", "kernelId": ""} id="utmzz69cqNJ6"
# ## One-Hot Encoding Model
# + gradient={"editing": false, "id": "7fdae940-8573-4659-b84b-f987d90af1c2", "kernelId": ""} id="b6KDl30qtgNe"
def standardize_text_string(text: str):
"""
create a custom standardization that:
1. Fixes whitespaces
2. Removes punctuations & numbers
3. Sets all texts to lowercase
4. Preserves the Elvish characters
"""
text = re.sub(r"[\s+]", " ", text)
text = re.sub(r"[0-9]", "", text)
text = re.sub(f"[{punctuation}–]", "", text)
return text.lower()
# + gradient={"editing": false, "id": "91f222d5-e339-4fe6-bae0-9d603ca30bee", "kernelId": ""} id="hEqgIvEtssNP"
# get unique characters in the text
chars = sorted(set(standardize_text_string(text)))
# + colab={"base_uri": "https://localhost:8080/"} gradient={"editing": false, "id": "757d1fc4-7294-4d10-add8-c5c4c1a7b306", "kernelId": ""} id="pQpwILNTtaXn" outputId="dc6358b9-3843-41ae-bb4f-084f33debdca"
print(chars, f"\n\nTotal unique characters: {len(chars)}")
# + gradient={"editing": false, "id": "780309e8-d5b1-4ce4-b925-197f2a4cb176", "kernelId": ""} id="6N5boivwtbF4"
# create dictionary mappings for chars to integers for vectorization & vice versa
char2int = {c: i for i, c in enumerate(chars)}
int2char = {i: c for c, i in char2int.items()}
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "530ccd9e-1f24-4c4b-8198-f36965c8a8ff", "kernelId": ""} id="ppDB2eqWujEi" outputId="813fbf60-fda2-4ece-e510-cfe6f6393f81"
print(char2int)
print(int2char)
# + gradient={"id": "cb39ef1c-48ed-4e82-8d50-8bd40609a57c", "kernelId": ""} id="mT7z5iWW35B2"
#let's standardize our original text
standardized_text = standardize_text_string(text)
# + gradient={"id": "e9f248c2-ec48-41d4-9ae2-5be9a2546dcf", "kernelId": ""} id="zklBFovrshdU"
# setting up sequence length and step to create dataset
MAX_SEQ_LEN = 20
step = 2
# + [markdown] id="x1MKin6j6RZX"
# Let's create our training examples from `standardized_text`. The input would be `sentences` where it is 'sampled' for `MAX_SEQ_LEN at every `step` from the length of the text.
#
# The output would be `next_chars` where it is the 'supposed' character the model should predict during the training time.
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "3215a480-d35f-45b0-99c2-4d416f973cb4", "kernelId": ""} id="P7KPwJgw6IvB" outputId="ca6539c0-8f9d-4e5f-bcec-6d348dbba44b"
# create training examples: input (`sentences`) and output (`next_chars`)
sentences = []
next_chars = []
for i in range(0, len(standardized_text) - MAX_SEQ_LEN, step):
sentences.append(standardized_text[i: i + MAX_SEQ_LEN])
next_chars.append(standardized_text[i + MAX_SEQ_LEN])
print("Total number of training examples:", len(sentences))
# + gradient={"id": "7c841ce5-2f6e-4291-ba68-0769f2068862", "kernelId": ""} id="auq1fW8ZulLl"
# get the total number of unique chars
# these parameters will also be used later on in our model
N_UNIQUE_CHARS = len(chars)
m = len(sentences)
def vectorize_sentence(text, max_seq_len=MAX_SEQ_LEN, n_unique_chars=N_UNIQUE_CHARS):
""" Convert input sentence into one-hot encoding numpy vector of shape
(m, max_seq_len, n_unique_chars) """
if type(text) == str:
# if text is input as string
if len(text) > max_seq_len:
# if text is longer than max_seq_len it will be truncated
## and appended on the list
text_list = []
for i in range(0, len(text), max_seq_len):
text_list.append(text[i: i+max_seq_len])
text = text_list
else:
# if text is less than max_seq_len, convert str -> list(str)
text = [text]
m = len(text) # get total number of sentences
x = np.zeros((m, max_seq_len, n_unique_chars), dtype=np.bool)
for i, sentence in enumerate(text):
# for each sentence in the `text` list
for p, char in enumerate(sentence.lower()):
# p is the position of the letter in the sentence
# char is the character in the sentence
x[i, p, char2int[char]] = 1
return x
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "e21e0faf-afcb-442c-9b0f-06fd69bcd946", "kernelId": ""} id="nmGPT_egvTr7" outputId="03966279-b9e7-4ffb-9e35-651ae59901d3"
# try out sentence to ensure we get the right shape
text_test = "ABCDEFGHIJKLMNOPQRSTUVWXYZ" + "ABCDEFGHIJKLMNOPQRSTUVWXYZ".lower()
text_test_vector = vectorize_sentence(text_test)
print("Shape of text vector: {}".format(text_test_vector.shape))
print(f"Supposed shape:{(round(len(text_test) / MAX_SEQ_LEN), MAX_SEQ_LEN, N_UNIQUE_CHARS)}")
# + [markdown] id="fgCNwUaA2TP5"
# Nice! Now that we got the right output shape from the `vectorize_sentence`, let's vectorize our `sentences`
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "099a779f-ef31-4aa0-ae84-b8f8fde39dc4", "kernelId": ""} id="E4iRZLJL2Fs2" outputId="e85a57e6-c153-4d93-daf2-2a4627802182"
# vectorize input sentences
X_data = vectorize_sentence(sentences);
print(X_data.shape)
# + [markdown] id="FZ436LRL4qPc"
# > Supposed shape: `(len(sentences), MAX_SEQ_LEN, N_UNIQUE_CHARS)`
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "4d319e9e-6d9a-4261-bf68-74744d8bdec0", "kernelId": ""} id="2tHNxwSv2OfG" outputId="3450ec8e-6359-48e8-b321-69f613a78d8d"
# vectorize next_chars (output) -> shape: (m, N_UNIQUE_CHARS)
y_data = np.zeros((m, N_UNIQUE_CHARS), dtype=np.bool)
for i, char in enumerate(next_chars):
y_data[i, char2int[char]] = 1
print(y_data.shape)
# + gradient={"id": "d40660ab-da06-486a-85cc-48ec25bd610e", "kernelId": ""} id="sE7AfpwEW2b4"
EMBEDDING_DIM = 16
def char_LSTM_model(max_seq_len=MAX_SEQ_LEN, max_features=N_UNIQUE_CHARS, embedding_dim=EMBEDDING_DIM):
# Define input for the model (vocab indices)
inputs = tf.keras.Input(shape=(max_seq_len, max_features))
# No embedding for one-hot encoding
# X = Embedding((max_seq_len, max_features), (max_features, embedding_dim))(inputs)
X = LSTM(128, return_sequences=True)(inputs)
X = Flatten()(X)
outputs = Dense(max_features, activation="softmax")(X)
model = Model(inputs, outputs, name="model_LSTM")
return model
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "b72c64b6-3b7d-4380-a9af-22f83a7b148f", "kernelId": ""} id="8VglUcNoXeqO" outputId="606b9285-ee3b-4005-8bcf-692eadddfaa1"
# let's create our model
model = char_LSTM_model()
optimizer=tf.keras.optimizers.Adam(learning_rate=0.001)
model.compile(loss="categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
model.summary()
# + [markdown] id="6q4WasWHJM_B"
# ### Sampling Functions
# To pick a word from the model's prediction output, we can either use:
# 1. Greedy search: take the character with the highest probability (argmax). But this will mean that our sampling will be the same.
# 2. Sampling: sampling from the distribution by picking a random character, but with the argmax values being the highest chance to be picked.
#
# References:
# 1. https://stackoverflow.com/questions/58764619/why-should-we-use-temperature-in-softmax
# 2. https://datascience.stackexchange.com/questions/72770/why-we-sample-when-predicting-with-recurent-neural-network
# + gradient={"id": "66a4aaae-04d4-4125-81bb-1b04852d33ef", "kernelId": ""} id="OeI1aOEbX2Vw"
def generate_text(model, original_sentence, step, temperature):
"""
Generates text from the `model` for `step` number
of times (equivalent to total characters sampled),
given the `original_sentence` (seed) and `temperature` value.
Args:
- model: LSTM model (Keras model)
- original_sentence: text to be used as the starting seed for sampling (str)
- step: number times that you'd want to sample.
... translates to total chars to be sampled (int)
- temperature: Temperature parameter for softmax function in `sample` (int)
"""
# get the original sentence
sentence = original_sentence
print(f"Generating with this sentence... '{original_sentence}'")
print("Temperature/Diversity value:", temperature)
generated_sentence = ""
for i in range(step):
seed = vectorize_sentence(sentence) # shape-> (1,20,36)
# get the softmax prediction
predictions=model.predict(seed)[0] # shape -> (20, 36)
# sample the softmax prediction
next_index = sample(predictions, temperature)
# convert next_index into character
next_char = int2char[next_index]
# append on our generated sentence
generated_sentence += next_char
# move the "sentence" (input) to the right by one char
## and append the predicted next_char
sentence = sentence[1:] + next_char
print(f"Generated: {generated_sentence}")
print()
def sample(predictions, temperature=0.2):
"""
Function to sample from the LSTM Softmax distribution
(as opposed to greedy search - argmax)
Args:
- predictions: LSTM softmax output of shape (MAX_SEQ_LEN, N_UNIQUE_CHARS)
- temperature: temperature parameter for softmax function.
... Provides diversity to the sample
... the higher the temperature, the less confident the model
about its pred (int)
Returns:
- max value from the probability distribution of softmax sampling (int)
"""
# convert into numpy array
predictions = np.asarray(predictions).astype("float64")
# perform softmax sampling
## the higher the temperature, the less confident the model about its pred
predictions = np.log(predictions) / temperature
exp_preds = np.exp(predictions)
predictions = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, predictions, 1)
return np.argmax(probas)
# + [markdown] id="nYXfGrto8JtS"
# Let's test out our sampling functions to see if they work
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "d7c64ca4-aa19-48ef-a20a-2944b3602f36", "kernelId": ""} id="AnTJ8zDw8OMB" outputId="092caed4-999c-46ce-a647-88561a0ec508"
BATCH_SIZE = 16
# fit only 1 epoch
model.fit(x=X_data, y=y_data, batch_size=BATCH_SIZE, epochs=1)
# + gradient={"id": "0838456e-d10a-4c7d-9500-f27de845aecd", "kernelId": ""} id="8CViiRyRMC2H"
SAMPLING_STEPS = 100
# -
# ### Training from Scratch
# _start here if you'd like to train from scratch_
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "428bcb78-e0a4-4bcc-9768-8e8c3a84d7db", "kernelId": ""} id="JspPhZ0UK_Jc" outputId="c21c15e5-b45c-429e-ef29-96362caf90d4"
def generate_and_sample(model, corpus, sequence_length, step, diversity_list):
"""
generate & sample characters from model's prediction output
Args:
- model: LSTM model (Keras model)
- corpus: Text to be used as a starting point / seed for sampling (str)
- sequence_length: Maximum sequence length to be for starting point (int)
- step: number times that you'd want to sample.
... translates to total chars to be sampled (int)
- diversity_list: List of temperature parameters for softmax function
... in `sample` (list)
Output:
prints generated text at different diversity/temperature values
"""
# set a random starting point in the text
start_index = random.randint(0, len(corpus) - sequence_length - 1)
# create a seed
original_sentence = corpus[start_index : start_index + MAX_SEQ_LEN]
for diversity in diversity_list:
generate_text(model, original_sentence, step, diversity)
print()
generate_and_sample(model, standardized_text, MAX_SEQ_LEN, SAMPLING_STEPS, [0.2, 0.5, 1.0, 1.2])
# + [markdown] id="tquKmXL7MLBv"
# Of course, our model's prediction output won't make sense it because it's only been trained for 1 epoch, but hey, our sampling functions worked!
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "df19e572-35e8-4db9-b8f7-0c4c1945bea4", "kernelId": ""} id="NSkWhRAM5xQa" outputId="6bb8efdc-eaf0-462f-9051-a73d07dde414"
# Create a callback that saves the model's weights
checkpoint_path = path / "models/one_hot/model_cp.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
cp_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_path,
save_weights_only=True,
verbose=1)
# Train the model
epochs = 30
BATCH_SIZE = 64
SAMPLING_STEPS = 100
diversity_list = [0.2, 0.5, 1.0, 1.2]
for epoch in range(epochs):
print("-"*40 + f" Epoch: {epoch}/{epochs} " + "-"*40)
model.fit(X_data, y_data, batch_size=BATCH_SIZE, epochs=1, callbacks=[cp_callback])
print()
print("*"*30 + f" Generating text after epoch #{epoch} " + "*"*30)
generate_and_sample(model, standardized_text, MAX_SEQ_LEN, SAMPLING_STEPS,
diversity_list)
# + gradient={"id": "ad1a71e6-6418-4bd3-bb2d-cfe0dde37176", "kernelId": ""} id="rq68WrcQYDyh"
model.save(path / "models/Char_LSTM_LOTR_OneHot.h5")
# -
# ### Load Saved Model
# _Start here if you'd like to use the saved model_
# + gradient={"id": "a9d4aa50-ffe9-4735-9b6a-f20121fdeb46", "kernelId": ""} id="1jRj-VpENVPv"
model = load_model(path / "models/Char_LSTM_LOTR_OneHot.h5")
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "e009cd9a-e35b-4568-9b99-9cb3c1bf2399", "kernelId": ""} id="41ddr6Ikw1BM" outputId="b8ea2b70-031d-4c56-85e5-a039b99d53aa"
# model.evaluate(X_data, y_data)
# + [markdown] id="1vtjx6ip_ASf"
# ### Test out different learning rates
#
# Let's test out different learning rates on the model to see if we can squeeze better accuracy than 0.64 - 0.66
#
# Using the equation:
# $1e^{-3} \times 1000^{\frac{epoch}{total\_epoch}}$
#
# we should get the following range:
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "4beac8ca-c860-45d9-a214-187e499e354e", "kernelId": ""} id="w-grF12wEI6s" outputId="1beb1db5-7b09-4809-d7f5-63345c5e6a92"
total_epochs = 10
for i in range(total_epochs + 1):
print("*"*20 + f" Epoch: {i} " + "*"*20)
print(1e-3 * 1000 ** (i/total_epochs))
# + [markdown] id="SNdulE5FGhJS"
# Let's train our model for another 10 epochs
# + gradient={"id": "33983fd4-1248-4a53-b3a0-cefd1b906f99", "kernelId": ""}
# class GenerateSampleCallback(tf.keras.callbacks.Callback):
# """
# generate & sample characters from model's prediction output
# Args:
# - corpus: Text to be used as a starting point / seed for sampling (str)
# - sequence_length: Maximum sequence length to be for starting point (int)
# - step: number times that you'd want to sample.
# ... translates to total chars to be sampled (int)
# - diversity_list: List of temperature parameters for softmax function
# ... in `sample` (list)
# Output:
# generates text at different diversity/temperature at epoch_end
# """
# def __init__(self, corpus, sequence_length, step, diversity_list = [0.2, 0.5, 1.0, 1.2]):
# self.corpus = corpus
# self.sequence_length = sequence_length
# self.step = step
# self.diversity_list = diversity_list
# def on_epoch_end(self, epoch, logs=None):
# start_index = random.randint(0, len(self.corpus) - self.sequence_length - 1)
# # create a seed
# original_sentence = self.corpus[start_index : start_index + self.sequence_length]
# for diversity in self.diversity_list:
# self.generate_text(self.model, original_sentence, self.step, diversity)
# print()
# def generate_text(self, model, original_sentence, step, temperature):
# """
# Generates text from the `model` for `step` number
# of times (equivalent to total characters sampled),
# given the `original_sentence` (seed) and `temperature` value.
# Args:
# - model: LSTM model (Keras model)
# - original_sentence: text to be used as the starting seed for sampling (str)
# - step: number times that you'd want to sample.
# ... translates to total chars to be sampled (int)
# - temperature: Temperature parameter for softmax function in `sample` (int)
# """
# # get the original sentence
# sentence = original_sentence
# print(f"Generating with this sentence... '{original_sentence}'")
# print("Temperature/Diversity value:", temperature)
# generated_sentence = ""
# for i in range(step):
# seed = vectorize_sentence(sentence) # shape-> (1,20,36)
# # get the softmax prediction
# predictions=model.predict(seed)[0] # shape -> (20, 36)
# # sample the softmax prediction
# next_index = self.sample(predictions, temperature)
# # convert next_index into character
# next_char = int2char[next_index]
# # append on our generated sentence
# generated_sentence += next_char
# # move the "sentence" (input) to the right by one char
# ## and append the predicted next_char
# sentence = sentence[1:] + next_char
# print(f"Generated: {generated_sentence}")
# print()
# def sample(predictions, temperature=0.2):
# """
# Function to sample from the LSTM Softmax distribution
# (as opposed to greedy search - argmax)
# Args:
# - predictions: LSTM softmax output of shape (MAX_SEQ_LEN, N_UNIQUE_CHARS)
# - temperature: temperature parameter for softmax function.
# ... Provides diversity to the sample
# ... the higher the temperature, the less confident the model
# about its pred (int)
# Returns:
# - max value from the probability distribution of softmax sampling (int)
# """
# # convert into numpy array
# predictions = np.asarray(predictions).astype("float64")
# # perform softmax sampling
# ## the higher the temperature, the less confident the model about its pred
# predictions = np.log(predictions) / temperature
# exp_preds = np.exp(predictions)
# predictions = exp_preds / np.sum(exp_preds)
# probas = np.random.multinomial(1, predictions, 1)
# return np.argmax(probas)
# + colab={"base_uri": "https://localhost:8080/"} gradient={"id": "e05a5f13-8d76-44cb-b906-57a0e6011fa2", "kernelId": ""} id="Y_pPmk1VDZ66" outputId="d4789d49-49c4-41c2-8e6e-90f04f9c08b8"
total_epochs = 10
BATCH_SIZE=64
# callback function that sets different learning rate at each epoch
lr_callback = LearningRateScheduler(lambda epoch: 1e-3 * 1000 ** (epoch / total_epochs))
# callback function to save model checkpoints
checkpoint_path = path / "models/one_hot/model_cp_lr.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
cp_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_path,
save_weights_only=True,
verbose=1)
lr_history = model.fit(X_data, y_data, epochs = total_epochs, verbose = 1, batch_size=BATCH_SIZE, callbacks=[lr_callback, cp_callback])
model.save(path / "models/Char_LSTM_LOTR_OneHot_LR.h5")
# + gradient={"id": "59a5f6ad-5fd4-45c6-9711-e4ec90a16dac", "kernelId": ""} id="NKUmNt2kGl4A"
|
LOTR_LSTM_Character_Level_OneHot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ClOkvz0yb-gs" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 2, Module 3*
#
# ---
# + [markdown] id="gTKIlwkWb-gv" colab_type="text"
# # Cross-Validation
#
#
# ## Assignment
# - [ ] [Review requirements for your portfolio project](https://lambdaschool.github.io/ds/unit2), then submit your dataset.
# - [ ] Continue to participate in our Kaggle challenge.
# - [ ] Use scikit-learn for hyperparameter optimization with RandomizedSearchCV.
# - [ ] Submit your predictions to our Kaggle competition. (Go to our Kaggle InClass competition webpage. Use the blue **Submit Predictions** button to upload your CSV file. Or you can use the Kaggle API to submit your predictions.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
#
# You won't be able to just copy from the lesson notebook to this assignment.
#
# - Because the lesson was ***regression***, but the assignment is ***classification.***
# - Because the lesson used [TargetEncoder](https://contrib.scikit-learn.org/categorical-encoding/targetencoder.html), which doesn't work as-is for _multi-class_ classification.
#
# So you will have to adapt the example, which is good real-world practice.
#
# 1. Use a model for classification, such as [RandomForestClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html)
# 2. Use hyperparameters that match the classifier, such as `randomforestclassifier__ ...`
# 3. Use a metric for classification, such as [`scoring='accuracy'`](https://scikit-learn.org/stable/modules/model_evaluation.html#common-cases-predefined-values)
# 4. If you’re doing a multi-class classification problem — such as whether a waterpump is functional, functional needs repair, or nonfunctional — then use a categorical encoding that works for multi-class classification, such as [OrdinalEncoder](https://contrib.scikit-learn.org/categorical-encoding/ordinal.html) (not [TargetEncoder](https://contrib.scikit-learn.org/categorical-encoding/targetencoder.html))
#
#
#
# ## Stretch Goals
#
# ### Reading
# - <NAME>, [Python Data Science Handbook, Chapter 5.3](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html), Hyperparameters and Model Validation
# - <NAME>, [Statistics for Hackers](https://speakerdeck.com/jakevdp/statistics-for-hackers?slide=107)
# - <NAME>, [A Programmer's Guide to Data Mining, Chapter 5](http://guidetodatamining.com/chapter5/), 10-fold cross validation
# - <NAME>, [A Basic Pipeline and Grid Search Setup](https://github.com/rasbt/python-machine-learning-book/blob/master/code/bonus/svm_iris_pipeline_and_gridsearch.ipynb)
# - <NAME>, [A Comparison of Grid Search and Randomized Search Using Scikit Learn](https://blog.usejournal.com/a-comparison-of-grid-search-and-randomized-search-using-scikit-learn-29823179bc85)
#
# ### Doing
# - Add your own stretch goals!
# - Try other [categorical encodings](https://contrib.scikit-learn.org/categorical-encoding/). See the previous assignment notebook for details.
# - In additon to `RandomizedSearchCV`, scikit-learn has [`GridSearchCV`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html). Another library called scikit-optimize has [`BayesSearchCV`](https://scikit-optimize.github.io/notebooks/sklearn-gridsearchcv-replacement.html). Experiment with these alternatives.
# - _[Introduction to Machine Learning with Python](http://shop.oreilly.com/product/0636920030515.do)_ discusses options for "Grid-Searching Which Model To Use" in Chapter 6:
#
# > You can even go further in combining GridSearchCV and Pipeline: it is also possible to search over the actual steps being performed in the pipeline (say whether to use StandardScaler or MinMaxScaler). This leads to an even bigger search space and should be considered carefully. Trying all possible solutions is usually not a viable machine learning strategy. However, here is an example comparing a RandomForestClassifier and an SVC ...
#
# The example is shown in [the accompanying notebook](https://github.com/amueller/introduction_to_ml_with_python/blob/master/06-algorithm-chains-and-pipelines.ipynb), code cells 35-37. Could you apply this concept to your own pipelines?
#
# + [markdown] id="6r4YK65gb-gw" colab_type="text"
# ### BONUS: Stacking!
#
# Here's some code you can use to "stack" multiple submissions, which is another form of ensembling:
#
# ```python
# import pandas as pd
#
# # Filenames of your submissions you want to ensemble
# files = ['submission-01.csv', 'submission-02.csv', 'submission-03.csv']
#
# target = 'status_group'
# submissions = (pd.read_csv(file)[[target]] for file in files)
# ensemble = pd.concat(submissions, axis='columns')
# majority_vote = ensemble.mode(axis='columns')[0]
#
# sample_submission = pd.read_csv('sample_submission.csv')
# submission = sample_submission.copy()
# submission[target] = majority_vote
# submission.to_csv('my-ultimate-ensemble-submission.csv', index=False)
# ```
# + id="2OUcbYBSb-gx" colab_type="code" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="8wva3b1Xb-g0" colab_type="code" colab={}
import pandas as pd
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# + id="2-8QxX9Pb-g2" colab_type="code" colab={}
# Split train into train & val
from sklearn.model_selection import train_test_split
import numpy as np
train, val = train_test_split(train, train_size=0.80, test_size=0.20,
stratify=train['status_group'], random_state=42)
def wrangle(X):
"""Wrangle train, validate, and test sets in the same way"""
# Prevent SettingWithCopyWarning
X = X.copy()
# About 3% of the time, latitude has small values near zero,
# outside Tanzania, so we'll treat these values like zero.
X['latitude'] = X['latitude'].replace(-2e-08, 0)
# When columns have zeros and shouldn't, they are like null values.
# So we will replace the zeros with nulls, and impute missing values later.
# Also create a "missing indicator" column, because the fact that
# values are missing may be a predictive signal.
cols_with_zeros = ['longitude', 'latitude', 'construction_year',
'gps_height', 'population']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
X[col+'_MISSING'] = X[col].isnull()
# Drop duplicate columns
duplicates = ['quantity_group', 'payment_type']
X = X.drop(columns=duplicates)
# Drop recorded_by (never varies) and id (always varies, random)
unusable_variance = ['recorded_by', 'id']
X = X.drop(columns=unusable_variance)
# Convert date_recorded to datetime
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.day
X = X.drop(columns='date_recorded')
# Engineer feature: how many years from construction_year to date_recorded
X['years'] = X['year_recorded'] - X['construction_year']
X['years_MISSING'] = X['years'].isnull()
# return the wrangled dataframe
return X
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
# + id="GvjaKyf88BPd" colab_type="code" colab={}
# The status_group column is the target
target = 'status_group'
# Get a dataframe with all train columns except the target
train_features = train.drop(columns=[target])
# Get a list of the numeric features
numeric_features = train_features.select_dtypes(include='number').columns.tolist()
# Get a series with the cardinality of the nonnumeric features
cardinality = train_features.select_dtypes(exclude='number').nunique()
# Get a list of all categorical features with cardinality <= 50
categorical_features = cardinality[cardinality <= 50].index.tolist()
# Combine the lists
features = numeric_features + categorical_features
# + id="Ss4nfmo68Maq" colab_type="code" colab={}
# Arrange data into X features matrix and y target vector
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test[features]
# + id="w2JbiyVk8QXR" colab_type="code" colab={}
import category_encoders as ce
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import cross_val_score
# + id="5yhOcNi8JDvT" colab_type="code" colab={}
pipeline = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(strategy='median'),
RandomForestClassifier(n_estimators=100, n_jobs=-1, random_state=42)
)
# + id="739_qyRBK9c8" colab_type="code" colab={}
encoder = ce.OrdinalEncoder()
X_train_encoded = encoder.fit_transform(X_train, y_train)
X_val_encoded = encoder.transform(X_val, y_val)
# + id="zrTGCZBSLXkL" colab_type="code" colab={}
imputer = SimpleImputer(strategy='median')
X_train_imputed = imputer.fit_transform(X_train_encoded)
X_val_imputed = imputer.transform(X_val_encoded)
# + id="m0hHObQ6MIfs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="ef3e0725-6b79-4057-c905-1766d9a2662a"
model = RandomForestClassifier(n_estimators=10, n_jobs=-1, random_state=42)
model.fit(X_train_imputed, y_train)
# + id="8uZdNH8mPRmE" colab_type="code" colab={}
X_train.dtypes
# + id="WwOtrFBE9IMA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="88ccf403-91ef-4c2d-f260-a9f2372ba8f3"
k = 3
scores = cross_val_score(pipeline, X_train, y_train, cv=k) #scoring='neg_mean_absolute_error')
print(f'MAE for {k} folds:', -scores)
# + id="QAf3o_AuPNWk" colab_type="code" colab={}
|
module3/NB_LS_DS_223_assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
pd.set_option('display.max_columns', None)
# Taking the mean values for each government policy, grouping by the cluster
top_df = pd.read_csv("top_5_per_cluster.csv")
bottom_df = pd.read_csv("bottom_5_per_cluster.csv")
for column in top_df.columns[13:]:
top_df[column] = top_df.groupby(['cluster'])[column].transform('mean')
bottom_df[column] = bottom_df.groupby(['cluster'])[column].transform('mean')
top_df = top_df.iloc[0:20,12:52].drop_duplicates()
top_df
# -
bottom_df = bottom_df.iloc[0:20,12:52].drop_duplicates()
bottom_df
|
04. Code/Best Gov Policies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="jG7ZEc_982io"
# # StyleGAN2-ADA-PyTorch
#
# **Notes**
# * Training section should be fairly stable. I’ll slowly add features but it should work for most mainstream use cases
# * Inference section is a work in progress. If you come across bug or have feature requests please post them in [Slack](https://ml-images.slack.com/archives/CLJGF384R) or on [Github](https://github.com/dvschultz/stylegan2-ada-pytorch/issues)
#
# ---
#
# If you find this notebook useful, consider signing up for my [Patreon](https://www.patreon.com/bustbright) or [YouTube channel](https://www.youtube.com/channel/UCaZuPdmZ380SFUMKHVsv_AA/join). You can also send me a one-time payment on [Venmo](https://venmo.com/Derrick-Schultz).
# + [markdown] id="Vj4PG4_i9Alt"
# ## Setup
# + [markdown] id="qGEXPcFJ9UTY"
# Let’s start by checking to see what GPU we’ve been assigned. Ideally we get a V100, but a P100 is fine too. Other GPUs may lead to issues.
# + id="9giNAy_VqBku" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619747859060, "user_tz": -540, "elapsed": 2650, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="41b08177-d2a8-4252-e17f-e5ea63cefe47"
# !nvidia-smi
# + id="_zedqBUQXdim" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619616474945, "user_tz": -540, "elapsed": 911, "user": {"displayName": "SHOT", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gh5vvPwI0wxV86goecWvaz89UBqVzkx1WnnD5ovSU8=s64", "userId": "16892596591524153853"}} outputId="9db01dc6-e5f8-4994-db2d-f1a0712a209f"
# !ls
# + [markdown] id="rSV_HEoD9dxo"
# Next let’s connect our Google Drive account. This is optional but highly recommended.
# + id="IuVPuJmbigRs" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619747957264, "user_tz": -540, "elapsed": 57748, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="a45a85d0-f957-4c12-a21f-e611a28f7711"
from google.colab import drive
drive.mount('/content/drive')
# + id="z2_B1aKxX2Eg"
# !ls '/content/drive/My Drive'
# + colab={"base_uri": "https://localhost:8080/"} id="c_JZrWZcEjXD" executionInfo={"status": "ok", "timestamp": 1619747961660, "user_tz": -540, "elapsed": 957, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="21a22b88-d7e1-45ce-d337-ab7840bacb51"
# !pwd
# + [markdown] id="nTjVmfSK9CYa"
# ## Install repo
#
# The next cell will install the StyleGAN repository in Google Drive. If you have already installed it it will just move into that folder. If you don’t have Google Drive connected it will just install the necessary code in Colab.
# + id="B8ADVNpBh8Ox" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619756911129, "user_tz": -540, "elapsed": 6100, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="38d21b8f-13eb-4070-f2da-14e0aaf3c374"
# !mkdir colab-sg2-ada-pytorch
# %cd colab-sg2-ada-pytorch
# !git clone https://github.com/cordob/stylegan2-ada-pytorch
# %cd stylegan2-ada-pytorch
# !mkdir downloads
# !mkdir datasets
# !mkdir pretrained
# !mkdir training_runs
# !pip install ninja opensimplex
# + id="-cyV-0x2etdf" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619672310862, "user_tz": -540, "elapsed": 533, "user": {"displayName": "SHOT", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gh5vvPwI0wxV86goecWvaz89UBqVzkx1WnnD5ovSU8=s64", "userId": "16892596591524153853"}} outputId="4bda0037-2eb3-4e9d-ad65-2ce17d3490f4"
# !pwd
# + id="fYTc-YQ8hWGk" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619756915415, "user_tz": -540, "elapsed": 844, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="fd4a6366-c39c-48f8-ef92-2430f08abc46"
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="6qYvmln-_3b8" executionInfo={"status": "ok", "timestamp": 1619750015398, "user_tz": -540, "elapsed": 20215, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="7376cd64-036b-48a6-becf-47e9fc99de3f"
from google.colab import drive
drive.mount('/content/drive')
# + id="JhHwUgMQ_7Xv" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619750215312, "user_tz": -540, "elapsed": 447, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="77b46ce9-ab0e-482d-be9a-0a4d531efadb"
# !ls '/content/drive/My Drive'
# + [markdown] id="zMl-oOoe64it"
# generate images from prtained
#
# + id="wwLlE-wc3DJI" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619748056912, "user_tz": -540, "elapsed": 71877, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "<KEY>", "userId": "07447657114463313189"}} outputId="5578d074-6e94-4e6b-ffc1-d48e3815826f"
# !python generate.py --outdir=out --trunc=0.7 --seeds=8100-8125 --network=https://nvlabs-fi-cdn.nvidia.com/stylegan2-ada-pytorch/pretrained/ffhq.pkl
# + id="Frahw_kxfLhr"
# !sudo apt install imagemagick-6.q16
# + id="p9en6vtSeqFD"
# !montage -mode concatenate -tile 4x4 out/*.png out/result.jpg
# + id="jMiV3ga97UkH"
# !ls out
# + id="PrIFWaXz8cah"
from google.colab import files
from IPython import display
display.Image("out/result-0.jpg",
width=1600)
# + id="csqA7SOj-Joe"
# !ffmpeg -framerate 2 -pattern_type glob -i 'out/*.png' \
# -c:v libx264 out3.mp4
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="BWcWH4xgDmaF" executionInfo={"status": "ok", "timestamp": 1619656758594, "user_tz": -540, "elapsed": 823, "user": {"displayName": "SHOT", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gh5vvPwI0wxV86goecWvaz89UBqVzkx1WnnD5ovSU8=s64", "userId": "16892596591524153853"}} outputId="dc1a6716-d10f-4342-daf2-9691b55ec8f9"
from google.colab import files
files.download('out3.mp4')
# + colab={"base_uri": "https://localhost:8080/"} id="AUsf_JY42kxW" executionInfo={"status": "ok", "timestamp": 1619653909635, "user_tz": -540, "elapsed": 29873, "user": {"displayName": "SHOT", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gh5vvPwI0wxV86goecWvaz89UBqVzkx1WnnD5ovSU8=s64", "userId": "16892596591524153853"}} outputId="7896f7b9-114d-4294-9075-657b7be16881"
# !python render.py --mp4_fps 30 --filename test --duration_sec 5 --network_pkl https://nvlabs-fi-cdn.nvidia.com/stylegan2-ada-pytorch/pretrained/ffhq.pkl
# + id="-7rZnvjU55J7" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619757178611, "user_tz": -540, "elapsed": 642, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="ab3ed87c-5962-4f0b-9bcf-2e8db54bb8e7"
# !pwd
# + [markdown] id="YCaJJNjf6whX"
# video download
#
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="-61iJwqs5sBT" executionInfo={"status": "ok", "timestamp": 1619654092842, "user_tz": -540, "elapsed": 461, "user": {"displayName": "SHOT", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gh5vvPwI0wxV86goecWvaz89UBqVzkx1WnnD5ovSU8=s64", "userId": "16892596591524153853"}} outputId="acad5061-b035-4a0f-e739-8bad7fdc90c0"
from google.colab import files
files.download('videos/seed1619653880.mp4')
# + id="cK0ESEo06L3h"
# !rm videos/*.*
# + [markdown] id="9jMmUpn4DWRe"
# local file upload !!
# + id="OzAYBwowYMi8" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "Ly8gQ29weXJpZ2h0IDIwMTcgR29vZ2xlIExMQwovLwovLyBMaWNlbnNlZCB1bmRlciB0aGUgQXBhY2hlIExpY2Vuc2UsIFZlcnNpb24gMi4wICh0aGUgIkxpY2Vuc2UiKTsKLy8geW91IG1heSBub3QgdXNlIHRoaXMgZmlsZSBleGNlcHQgaW4gY29tcGxpYW5jZSB3aXRoIHRoZSBMaWNlbnNlLgovLyBZb3UgbWF5IG9idGFpbiBhIGNvcHkgb2YgdGhlIExpY2Vuc2UgYXQKLy8KLy8gICAgICBodHRwOi8vd3d3LmFwYWNoZS5vcmcvbGljZW5zZXMvTElDRU5TRS0yLjAKLy8KLy8gVW5sZXNzIHJlcXVpcmVkIGJ5IGFwcGxpY2FibGUgbGF3IG9yIGFncmVlZCB0byBpbiB3cml0aW5nLCBzb2Z0d2FyZQovLyBkaXN0cmlidXRlZCB1bmRlciB0aGUgTGljZW5zZSBpcyBkaXN0cmlidXRlZCBvbiBhbiAiQVMgSVMiIEJBU0lTLAovLyBXSVRIT1VUIFdBUlJBTlRJRVMgT1IgQ09ORElUSU9OUyBPRiBBTlkgS0lORCwgZWl0aGVyIGV4cHJlc3Mgb3IgaW1wbGllZC4KLy8gU2VlIHRoZSBMaWNlbnNlIGZvciB0aGUgc3BlY2lmaWMgbGFuZ3VhZ2UgZ292ZXJuaW5nIHBlcm1pc3Npb25zIGFuZAovLyBsaW1pdGF0aW9ucyB1bmRlciB0aGUgTGljZW5zZS4KCi8qKgogKiBAZmlsZW92ZXJ2aWV3IEhlbHBlcnMgZm9yIGdvb2dsZS5jb2xhYiBQeXRob24gbW9kdWxlLgogKi8KKGZ1bmN0aW9uKHNjb3BlKSB7CmZ1bmN0aW9uIHNwYW4odGV4dCwgc3R5bGVBdHRyaWJ1dGVzID0ge30pIHsKICBjb25zdCBlbGVtZW50ID0gZG9jdW1lbnQuY3JlYXRlRWxlbWVudCgnc3BhbicpOwogIGVsZW1lbnQudGV4dENvbnRlbnQgPSB0ZXh0OwogIGZvciAoY29uc3Qga2V5IG9mIE9iamVjdC5rZXlzKHN0eWxlQXR0cmlidXRlcykpIHsKICAgIGVsZW1lbnQuc3R5bGVba2V5XSA9IHN0eWxlQXR0cmlidXRlc1trZXldOwogIH0KICByZXR1cm4gZWxlbWVudDsKfQoKLy8gTWF4IG51bWJlciBvZiBieXRlcyB3aGljaCB3aWxsIGJlIHVwbG9hZGVkIGF0IGEgdGltZS4KY29uc3QgTUFYX1BBWUxPQURfU0laRSA9IDEwMCAqIDEwMjQ7CgpmdW5jdGlvbiBfdXBsb2FkRmlsZXMoaW5wdXRJZCwgb3V0cHV0SWQpIHsKICBjb25zdCBzdGVwcyA9IHVwbG9hZEZpbGVzU3RlcChpbnB1dElkLCBvdXRwdXRJZCk7CiAgY29uc3Qgb3V0cHV0RWxlbWVudCA9IGRvY3VtZW50LmdldEVsZW1lbnRCeUlkKG91dHB1dElkKTsKICAvLyBDYWNoZSBzdGVwcyBvbiB0aGUgb3V0cHV0RWxlbWVudCB0byBtYWtlIGl0IGF2YWlsYWJsZSBmb3IgdGhlIG5leHQgY2FsbAogIC8vIHRvIHVwbG9hZEZpbGVzQ29udGludWUgZnJvbSBQeXRob24uCiAgb3V0cHV0RWxlbWVudC5zdGVwcyA9IHN0ZXBzOwoKICByZXR1cm4gX3VwbG9hZEZpbGVzQ29udGludWUob3V0cHV0SWQpOwp9CgovLyBUaGlzIGlzIHJvdWdobHkgYW4gYXN5bmMgZ2VuZXJhdG9yIChub3Qgc3VwcG9ydGVkIGluIHRoZSBicm93c2VyIHlldCksCi8vIHdoZXJlIHRoZXJlIGFyZSBtdWx0aXBsZSBhc3luY2hyb25vdXMgc3RlcHMgYW5kIHRoZSBQeXRob24gc2lkZSBpcyBnb2luZwovLyB0byBwb2xsIGZvciBjb21wbGV0aW9uIG9mIGVhY2ggc3RlcC4KLy8gVGhpcyB1c2VzIGEgUHJvbWlzZSB0byBibG9jayB0aGUgcHl0aG9uIHNpZGUgb24gY29tcGxldGlvbiBvZiBlYWNoIHN0ZXAsCi8vIHRoZW4gcGFzc2VzIHRoZSByZXN1bHQgb2YgdGhlIHByZXZpb3VzIHN0ZXAgYXMgdGhlIGlucHV0IHRvIHRoZSBuZXh0IHN0ZXAuCmZ1bmN0aW9uIF91cGxvYWRGaWxlc0NvbnRpbnVlKG91dHB1dElkKSB7CiAgY29uc3Qgb3V0cHV0RWxlbWVudCA9IGRvY3VtZW50LmdldEVsZW1lbnRCeUlkKG91dHB1dElkKTsKICBjb25zdCBzdGVwcyA9IG91dHB1dEVsZW1lbnQuc3RlcHM7CgogIGNvbnN0IG5leHQgPSBzdGVwcy5uZXh0KG91dHB1dEVsZW1lbnQubGFzdFByb21pc2VWYWx1ZSk7CiAgcmV0dXJuIFByb21pc2UucmVzb2x2ZShuZXh0LnZhbHVlLnByb21pc2UpLnRoZW4oKHZhbHVlKSA9PiB7CiAgICAvLyBDYWNoZSB0aGUgbGFzdCBwcm9taXNlIHZhbHVlIHRvIG1ha2UgaXQgYXZhaWxhYmxlIHRvIHRoZSBuZXh0CiAgICAvLyBzdGVwIG9mIHRoZSBnZW5lcmF0b3IuCiAgICBvdXRwdXRFbGVtZW50Lmxhc3RQcm9taXNlVmFsdWUgPSB2YWx1ZTsKICAgIHJldHVybiBuZXh0LnZhbHVlLnJlc3BvbnNlOwogIH0pOwp9CgovKioKICogR2VuZXJhdG9yIGZ1bmN0aW9uIHdoaWNoIGlzIGNhbGxlZCBiZXR3ZWVuIGVhY2ggYXN5bmMgc3RlcCBvZiB0aGUgdXBsb2FkCiAqIHByb2Nlc3MuCiAqIEBwYXJhbSB7c3RyaW5nfSBpbnB1dElkIEVsZW1lbnQgSUQgb2YgdGhlIGlucHV0IGZpbGUgcGlja2VyIGVsZW1lbnQuCiAqIEBwYXJhbSB7c3RyaW5nfSBvdXRwdXRJZCBFbGVtZW50IElEIG9mIHRoZSBvdXRwdXQgZGlzcGxheS4KICogQHJldHVybiB7IUl0ZXJhYmxlPCFPYmplY3Q+fSBJdGVyYWJsZSBvZiBuZXh0IHN0ZXBzLgogKi8KZnVuY3Rpb24qIHVwbG9hZEZpbGVzU3RlcChpbnB1dElkLCBvdXRwdXRJZCkgewogIGNvbnN0IGlucHV0RWxlbWVudCA9IGRvY3VtZW50LmdldEVsZW1lbnRCeUlkKGlucHV0SWQpOwogIGlucHV0RWxlbWVudC5kaXNhYmxlZCA9IGZhbHNlOwoKICBjb25zdCBvdXRwdXRFbGVtZW50ID0gZG9jdW1lbnQuZ2V0RWxlbWVudEJ5SWQob3V0cHV0SWQpOwogIG91dHB1dEVsZW1lbnQuaW5uZXJIVE1MID0gJyc7CgogIGNvbnN0IHBpY2tlZFByb21pc2UgPSBuZXcgUHJvbWlzZSgocmVzb2x2ZSkgPT4gewogICAgaW5wdXRFbGVtZW50LmFkZEV2ZW50TGlzdGVuZXIoJ2NoYW5nZScsIChlKSA9PiB7CiAgICAgIHJlc29sdmUoZS50YXJnZXQuZmlsZXMpOwogICAgfSk7CiAgfSk7CgogIGNvbnN0IGNhbmNlbCA9IGRvY3VtZW50LmNyZWF0ZUVsZW1lbnQoJ2J1dHRvbicpOwogIGlucHV0RWxlbWVudC5wYXJlbnRFbGVtZW50LmFwcGVuZENoaWxkKGNhbmNlbCk7CiAgY2FuY2VsLnRleHRDb250ZW50ID0gJ0NhbmNlbCB1cGxvYWQnOwogIGNvbnN0IGNhbmNlbFByb21pc2UgPSBuZXcgUHJvbWlzZSgocmVzb2x2ZSkgPT4gewogICAgY2FuY2VsLm9uY2xpY2sgPSAoKSA9PiB7CiAgICAgIHJlc29sdmUobnVsbCk7CiAgICB9OwogIH0pOwoKICAvLyBXYWl0IGZvciB0aGUgdXNlciB0byBwaWNrIHRoZSBmaWxlcy4KICBjb25zdCBmaWxlcyA9IHlpZWxkIHsKICAgIHByb21pc2U6IFByb21pc2UucmFjZShbcGlja2VkUHJvbWlzZSwgY2FuY2VsUHJvbWlzZV0pLAogICAgcmVzcG9uc2U6IHsKICAgICAgYWN0aW9uOiAnc3RhcnRpbmcnLAogICAgfQogIH07CgogIGNhbmNlbC5yZW1vdmUoKTsKCiAgLy8gRGlzYWJsZSB0aGUgaW5wdXQgZWxlbWVudCBzaW5jZSBmdXJ0aGVyIHBpY2tzIGFyZSBub3QgYWxsb3dlZC4KICBpbnB1dEVsZW1lbnQuZGlzYWJsZWQgPSB0cnVlOwoKICBpZiAoIWZpbGVzKSB7CiAgICByZXR1cm4gewogICAgICByZXNwb25zZTogewogICAgICAgIGFjdGlvbjogJ2NvbXBsZXRlJywKICAgICAgfQogICAgfTsKICB9CgogIGZvciAoY29uc3QgZmlsZSBvZiBmaWxlcykgewogICAgY29uc3QgbGkgPSBkb2N1bWVudC5jcmVhdGVFbGVtZW50KCdsaScpOwogICAgbGkuYXBwZW5kKHNwYW4oZmlsZS5uYW1lLCB7Zm9udFdlaWdodDogJ2JvbGQnfSkpOwogICAgbGkuYXBwZW5kKHNwYW4oCiAgICAgICAgYCgke2ZpbGUudHlwZSB8fCAnbi9hJ30pIC0gJHtmaWxlLnNpemV9IGJ5dGVzLCBgICsKICAgICAgICBgbGFzdCBtb2RpZmllZDogJHsKICAgICAgICAgICAgZmlsZS5sYXN0TW9kaWZpZWREYXRlID8gZmlsZS5sYXN0TW9kaWZpZWREYXRlLnRvTG9jYWxlRGF0ZVN0cmluZygpIDoKICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgJ24vYSd9IC0gYCkpOwogICAgY29uc3QgcGVyY2VudCA9IHNwYW4oJzAlIGRvbmUnKTsKICAgIGxpLmFwcGVuZENoaWxkKHBlcmNlbnQpOwoKICAgIG91dHB1dEVsZW1lbnQuYXBwZW5kQ2hpbGQobGkpOwoKICAgIGNvbnN0IGZpbGVEYXRhUHJvbWlzZSA9IG5ldyBQcm9taXNlKChyZXNvbHZlKSA9PiB7CiAgICAgIGNvbnN0IHJlYWRlciA9IG5ldyBGaWxlUmVhZGVyKCk7CiAgICAgIHJlYWRlci5vbmxvYWQgPSAoZSkgPT4gewogICAgICAgIHJlc29sdmUoZS50YXJnZXQucmVzdWx0KTsKICAgICAgfTsKICAgICAgcmVhZGVyLnJlYWRBc0FycmF5QnVmZmVyKGZpbGUpOwogICAgfSk7CiAgICAvLyBXYWl0IGZvciB0aGUgZGF0YSB0byBiZSByZWFkeS4KICAgIGxldCBmaWxlRGF0YSA9IHlpZWxkIHsKICAgICAgcHJvbWlzZTogZmlsZURhdGFQcm9taXNlLAogICAgICByZXNwb25zZTogewogICAgICAgIGFjdGlvbjogJ2NvbnRpbnVlJywKICAgICAgfQogICAgfTsKCiAgICAvLyBVc2UgYSBjaHVua2VkIHNlbmRpbmcgdG8gYXZvaWQgbWVzc2FnZSBzaXplIGxpbWl0cy4gU2VlIGIvNjIxMTU2NjAuCiAgICBsZXQgcG9zaXRpb24gPSAwOwogICAgd2hpbGUgKHBvc2l0aW9uIDwgZmlsZURhdGEuYnl0ZUxlbmd0aCkgewogICAgICBjb25zdCBsZW5ndGggPSBNYXRoLm1pbihmaWxlRGF0YS5ieXRlTGVuZ3RoIC0gcG9zaXRpb24sIE1BWF9QQVlMT0FEX1NJWkUpOwogICAgICBjb25zdCBjaHVuayA9IG5ldyBVaW50OEFycmF5KGZpbGVEYXRhLCBwb3NpdGlvbiwgbGVuZ3RoKTsKICAgICAgcG9zaXRpb24gKz0gbGVuZ3RoOwoKICAgICAgY29uc3QgYmFzZTY0ID0gYnRvYShTdHJpbmcuZnJvbUNoYXJDb2RlLmFwcGx5KG51bGwsIGNodW5rKSk7CiAgICAgIHlpZWxkIHsKICAgICAgICByZXNwb25zZTogewogICAgICAgICAgYWN0aW9uOiAnYXBwZW5kJywKICAgICAgICAgIGZpbGU6IGZpbGUubmFtZSwKICAgICAgICAgIGRhdGE6IGJhc2U2NCwKICAgICAgICB9LAogICAgICB9OwogICAgICBwZXJjZW50LnRleHRDb250ZW50ID0KICAgICAgICAgIGAke01hdGgucm91bmQoKHBvc2l0aW9uIC8gZmlsZURhdGEuYnl0ZUxlbmd0aCkgKiAxMDApfSUgZG9uZWA7CiAgICB9CiAgfQoKICAvLyBBbGwgZG9uZS4KICB5aWVsZCB7CiAgICByZXNwb25zZTogewogICAgICBhY3Rpb246ICdjb21wbGV0ZScsCiAgICB9CiAgfTsKfQoKc2NvcGUuZ29vZ2xlID0gc2NvcGUuZ29vZ2xlIHx8IHt9OwpzY29wZS5nb29nbGUuY29sYWIgPSBzY29wZS5nb29nbGUuY29sYWIgfHwge307CnNjb3BlLmdvb2dsZS5jb2xhYi5fZmlsZXMgPSB7CiAgX3VwbG9hZEZpbGVzLAogIF91cGxvYWRGaWxlc0NvbnRpbnVlLAp9Owp9KShzZWxmKTsK", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 76} executionInfo={"status": "ok", "timestamp": 1619757138889, "user_tz": -540, "elapsed": 67596, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="6ad488e0-0215-4eff-b774-35191b704932"
from google.colab import files
uploaded = files.upload()
# + id="-qLCNbV9XMrc" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619757162851, "user_tz": -540, "elapsed": 679, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="2db05aee-d1b0-4c87-99f4-560911f68489"
# !ls '/content/drive/My Drive/MUNCH.zip'
# + colab={"base_uri": "https://localhost:8080/"} id="u09225Mao0cb" executionInfo={"status": "ok", "timestamp": 1619757170177, "user_tz": -540, "elapsed": 602, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="17f0d522-243f-4c60-d056-dafe39632190"
# !pwd
# + id="5zNKwafgzHtw" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619757226610, "user_tz": -540, "elapsed": 817, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="e6614521-e771-4100-dbc9-f8a08f48ec31"
# !unzip 'MUNCH.zip' -d /content/colab-sg2-ada-pytorch/stylegan2-ada-pytorch
# + id="qd_bYsurYxTM"
# !ls m3
# + id="r2NmSQzRjoXU" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619757191489, "user_tz": -540, "elapsed": 837, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="48518e5e-7a6a-4bb4-f037-eb73ffb72136"
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="Dy-FFW8FpLho" executionInfo={"status": "ok", "timestamp": 1619757245136, "user_tz": -540, "elapsed": 2366, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="0aa309c4-adef-40a2-9b36-5c2f7996fce3"
# !python dataset_tool.py --source=m3 --dest=m3.zip --width=512 --height=512
# + [markdown] id="QwcuKnoJjy6N"
# transfer learning 기존 네트워크사용해서 !!!
# + colab={"base_uri": "https://localhost:8080/"} id="f7xPM9qLjsqq" outputId="91e1cecb-23ba-4873-a454-7f29f859e35a"
# !python train.py --outdir=training_runs --data=m3.zip --resume=https://nvlabs-fi-cdn.nvidia.com/stylegan2-ada/pretrained/afhqwild.pkl --gpus=1 --mirror=1
# + id="ImzBAMGfj6qt" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619756858816, "user_tz": -540, "elapsed": 945, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjK5icpssdxim4CNM813_k0CxYcl_syRlrFhRSc=s64", "userId": "07447657114463313189"}} outputId="b2703b10-802a-40be-8e4d-8dee6e684918"
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="v-a6FELFIgmx" executionInfo={"status": "ok", "timestamp": 1619741854078, "user_tz": -540, "elapsed": 1867, "user": {"displayName": "SHOT", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gh5vvPwI0wxV86goecWvaz89UBqVzkx1WnnD5ovSU8=s64", "userId": "16892596591524153853"}} outputId="599c2deb-b2b3-417f-80c6-dd352f8bd9aa"
# !python dataset_tool.py --source=m3 --dest=m3.zip --width=256 --height=512
# + colab={"base_uri": "https://localhost:8080/"} id="S4kG4rFcJtps" executionInfo={"status": "ok", "timestamp": 1619750672721, "user_tz": -540, "elapsed": 292, "user": {"displayName": "Pado\ud30c\ub3c4", "photoUrl": "https://<KEY>", "userId": "07447657114463313189"}} outputId="4ca0aab2-acf8-4791-c75f-b51ec5117cec"
# !python train.py --outdir=training_runs --data=m3.zip --gpus=1
# + [markdown] id="maP-aJUNkth-"
# train 훈련 ~~~~
#
# + id="XLghfnCsjrc9"
dataset = 'degas' #@param {type:"string"}
resume_pkl = 'https://drive.google.com/file/d/1yyEUSPwaGqudWldiWTMu1nkSRNbEush7/view?usp=sharing' #@param {type:"string"}
lod_kimg = 30 #@param {type:"integer"}
# %run train.py --data $dataset --resume $resume_pkl --lod_kimg $lod_kimg
# + [markdown] id="cZkcJ58P97Ls"
# ## Dataset Preparation
#
# Upload a .zip of square images to the `datasets` folder. Previously you had to convert your model to .tfrecords. That’s no longer needed :)
# + [markdown] id="5B-h6FpB9FaK"
# ## Train model
# + [markdown] id="bNc-3wTO-MUd"
# Below are a series of variables you need to set to run the training. You probably won’t need to touch most of them.
#
# * `dataset_path`: this is the path to your .zip file
# * `resume_from`: if you’re starting a new dataset I recommend `'ffhq1024'` or `'./pretrained/wikiart.pkl'`
# * `mirror_x` and `mirror_y`: Allow the dataset to use horizontal or vertical mirroring.
# + id="EL-M7WnnfMDI"
# !python train.py --gpus=1 --cfg=$config --metrics=None --outdir=./results --data=$dataset_path --snap=$snapshot_count --resume=$resume_from --augpipe=$augs --initstrength=$aug_strength --gamma=$gamma_value --mirror=$mirror_x --mirrory=False --nkimg=$train_count
# + [markdown] id="RgvSvfyi_R_-"
# ### Resume Training
#
# Once Colab has shutdown, you’ll need to resume your training. Reset the variables above, particularly the `resume_from` and `aug_strength` settings.
#
# 1. Point `resume_from` to the last .pkl you trained (you’ll find these in the `results` folder)
# 2. Update `aug_strength` to match the augment value of the last pkl file. Often you’ll see this in the console, but you may need to look at the `log.txt`. Updating this makes sure training stays as stable as possible.
# 3. You may want to update `train_count` to keep track of your training progress.
#
# Once all of this has been reset, run that variable cell and the training command cell after it.
# + [markdown] id="VznRirOE5ENI"
# ## Convert Legacy Model
#
# If you have an older version of a model (Tensorflow based StyleGAN, or Runway downloaded .pkl file) you’ll need to convert to the newest version. If you’ve trained in this notebook you do **not** need to use this cell.
#
# `--source`: path to model that you want to convert
#
# `--dest`: path and file name to convert to.
# + id="CzkP-Rww5Np9"
# !python legacy.py --source=/content/drive/MyDrive/runway.pkl --dest=/content/drive/MyDrive/colab-sg2-ada-pytorch/stylegan2-ada-pytorch/runway.pkl
# + [markdown] id="L6EtrPqL9ILk"
# ## Testing/Inference
#
# Also known as "Inference", "Evaluation" or "Testing" the model. This is the process of usinng your trained model to generate new material, usually images or videos.
# + [markdown] id="mYdyfH0O8In_"
# ### Generate Single Images
#
# `--network`: Make sure the `--network` argument points to your .pkl file. (My preferred method is to right click on the file in the Files pane to your left and choose `Copy Path`, then paste that into the argument after the `=` sign).
#
# `--seeds`: This allows you to choose random seeds from the model. Remember that our input to StyleGAN is a 512-dimensional array. These seeds will generate those 512 values. Each seed will generate a different, random array. The same seed value will also always generate the same random array, so we can later use it for other purposes like interpolation.
#
# `--truncation`: Truncation, well, truncates the latent space. This can have a subtle or dramatic affect on your images depending on the value you use. The smaller the number the more realistic your images should appear, but this will also affect diversity. Most people choose between 0.5 and 1.0, but technically it's infinite.
#
# + id="VYRXenMoZSHf"
# !python generate.py --outdir=/content/out/images/ --trunc=0.8 --seeds=0-499 --network=/content/drive/MyDrive/network-snapshot-008720.pkl
# + [markdown] id="VjOTCWVonoVL"
# ### Truncation Traversal
#
# Below you can take one seed and look at the changes to it across any truncation amount. -1 to 1 will be pretty realistic images, but the further out you get the weirder it gets.
#
# #### Options
# `--network`: Again, this should be the path to your .pkl file.
#
# `--seeds`: Pass this only one seed. Pick a favorite from your generated images.
#
# `--start`: Starting truncation value.
#
# `--stop`: Stopping truncation value. This should be larger than the start value. (Will probably break if its not).
#
# `--increment`: How much each frame should increment the truncation value. Make this really small if you want a long, slow interpolation. (stop-start/increment=total frames)
#
# + id="nyzdGr7OnrMG"
# !python generate.py --process="truncation" --outdir=/content/out/trunc-trav-3/ --start=-0.8 --stop=2.8 --increment=0.02 --seeds=470 --network=/content/drive/MyDrive/stylegan2-transfer-models/mixed6k-network-snapshot-016470.pkl
# + [markdown] id="OSzj0igO8Lfu"
# ### Interpolations
#
# Interpolation is the process of generating very small changes to a vector in order to make it appear animated from frame to frame.
#
# We’ll look at different examples of interpolation below.
#
# #### Options
#
# `--network`: path to your .pkl file
#
# `--interpolation`: Walk type defines the type of interpolation you want. In some cases it can also specify whether you want the z space or the w space.
#
# `--frames`: How many frames you want to produce. Use this to manage the length of your video.
#
# `--trunc`: truncation value
# + [markdown] id="OJSqafIzNwhx"
# #### Linear Interpolation
# + id="sqkiskly8S5_"
# !python generate.py --outdir=/content/out/video1-w-0.5/ --space="z" --trunc=0.5 --process="interpolation" --seeds=463,470 --network=/content/drive/MyDrive/stylegan2-transfer-models/mixed6k-network-snapshot-016470.pkl --frames=48
# + id="DCUEV3aO8s_X"
# !python generate.py --outdir=out/video1-w/ --space="w" --trunc=1 --process="interpolation" --seeds=85,265,297,849 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
# + id="pmKbwZDD8gjM"
# !zip -r vid1.zip /content/out/video1-w-0.5
# + [markdown] id="Yi3d7xzpN2Uj"
# #### Slerp Interpolation
#
# This gets a little heady, but technically linear interpolations are not the best in high-dimensional GANs. [This github link](https://github.com/soumith/dcgan.torch/issues/14) is one of the more popular explanations ad discussions.
#
# In reality I do not find a huge difference between linear and spherical interpolations (the difference in z- and w-space is enough in many cases), but I’ve implemented slerp here for anyone interested.
#
# Note: Slerp in w space currently isn’t supported. I’m working on it.
# + id="I0-cUd3fB_kJ"
# !python generate.py --outdir=out/video1/ --trunc=1 --process="interpolation" --interpolation="slerp" --seeds=85,265,297,849 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
# + [markdown] id="uP1HsU_CPcF5"
# #### Noise Loop
#
# If you want to just make a random but fun interpolation of your model the noise loop is the way to go. It creates a random path thru the z space to show you a diverse set of images.
#
# `--interpolation="noiseloop"`: set this to use the noise loop funtion
#
# `--diameter`: This controls how "wide" the loop is. Make it smaller to show a less diverse range of samples. Make it larger to cover a lot of samples. This plus `--frames` can help determine how fast the video feels.
#
# `--random_seed`: this allows you to change your starting place in the z space. Note: this value has nothing to do with the seeds you use to generate images. It just allows you to randomize your start point (and if you want to return to it you can use the same seed multiple times).
#
# Noise loops currently only work in z space.
# + id="gfR6DhfvN8b_"
# !python generate.py --outdir=out/video-noiseloop-0.9d/ --trunc=0.8 --process="interpolation" --interpolation="noiseloop" --diameter=0.9 --random_seed=100 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
# + [markdown] id="PkKFb-4CedOq"
# #### Circular Loop
#
# The noise loop is, well, noisy. This circular loop will feel much more even, while still providing a random loop.
#
# I recommend using a higher `--diameter` value than you do with noise loops. Something between `50.0` and `500.0` alongside `--frames` can help control speed and diversity.
# + id="Ao62za9_QfOF"
# !python generate.py --outdir=out/video-circularloop/ --trunc=1 --process="interpolation" --interpolation="circularloop" --diameter=800.00 --frames=720 --random_seed=90 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
# + id="T8Ld_ozNJCOn"
# + [markdown] id="qz-fVtzyAHg1"
# ## Projection
# + [markdown] id="ez7tXSpCA_zh"
#
#
# * `--target`: this is a path to the image file that you want to "find" in your model. This image must be the exact same size as your model.
# * `--num-steps`: how many iterations the projctor should run for. Lower will mean less steps and less likelihood of a good projection. Higher will take longer but will likely produce better images.
#
#
# + id="p84CtZUGAKnR"
# !python projector.py --help
# + id="80YTcjIQARWh"
# !python projector.py --network=/content/drive/MyDrive/colab-sg2-ada-pytorch/stylegan2-ada-pytorch/results/00023-chin-morris-mirror-11gb-gpu-gamma50-bg-resumecustom/network-snapshot-000304.pkl --outdir=/content/projector/ --target=/content/img005421_0.png --num-steps=200 --seed=0
# + [markdown] id="hAxADbdpHHib"
# ### <NAME>’ Projector
# + id="iwS_ey9QF-nk"
# !python /content/stylegan2-ada-pytorch/pbaylies_projector.py --help
# + id="-yj06MAABoLe"
# !python /content/stylegan2-ada-pytorch/pbaylies_projector.py --network=/content/drive/MyDrive/colab-sg2-ada-pytorch/stylegan2-ada-pytorch/results/00023-chin-morris-mirror-11gb-gpu-gamma50-bg-resumecustom/network-snapshot-000304.pkl --outdir=/content/projector-no-clip/ --target-image=/content/img005421_0.png --num-steps=200 --use-clip=False --seed=0
# + [markdown] id="qywlaS5pgzyH"
# ## Combine NPZ files together
# + id="M2VooqrNfIpw"
# !python combine_npz.py --outdir=/content/npz --npzs='/content/projector/projected_w.npz,/content/projector-no-clip/projected_w.npz'
# + id="uIqgl5nIHwpp"
# !python generate.py --help
# + id="4cgezYN8Dsyh"
# !python generate.py --process=interpolation --interpolation=linear --space=w --network=/content/drive/MyDrive/colab-sg2-ada-pytorch/stylegan2-ada-pytorch/results/00023-chin-morris-mirror-11gb-gpu-gamma50-bg-resumecustom/network-snapshot-000304.pkl --outdir=/content/test/ --projected-w=/content/npz/combined.npz
# + [markdown] id="lF7RCnSAsWrq"
# ## Feature Extraction using Closed Form Factorization
#
# Feature Extraction is the process of finding “human readable” vectors in a StyleGAN model. For example, let’s say you wanted to find a vector that could open or close a mouth in a face model.
#
# The feature extractor tries to automate the procss of finding important vectors in your model.
#
# `--ckpt`: This is the path to your .pkl file. In other places its called `--network` (It’s a long story for why its name changed here)
# `--out`: path to save your output feature vector file. The file name must end in `.pt`!
# + id="1Hek6TFZCKD-"
# !python closed_form_factorization.py --out=/content/ladies-black-cff.pt --ckpt=/content/drive/MyDrive/network-snapshot-008720.pkl
# + [markdown] id="WxLgeNeJRqFh"
# Once this cell is finished you’ll want to save that `.pt` file somewhere for reuse.
#
# This process just created the vctor values, but we need to test it on some seed values to determine what each vector actually changes. The `apply_factor.py` script does this.
#
# Arguments to try:
#
#
# * `-i`: This stands for index. By default, the cell above will produce 512 vectors, so `-i` can be any value from 0 to 511. I recommend starting with a higher value.
# * `-d`: This stands for degrees. This means how much change you want to see along th vector. I recommend a value between 5 and 10 to start with.
# * `--seeds`: You know what these are by now right? :)
# * `--ckpt`: path to your .pkl file
# * `--video`: adding this to your argument will produce a video that animates your seeds along the vector path. I find it much easier to figure out what’s changing with an animation.
# * `--output`: where to save the images/video
#
# Lastly you need to add the path to the `.pt` file you made in th above cell. It’s weird, but you don’t need to add any arguments bfore it, just make sure its after `apply_factor.pt`
#
#
# + id="dEDSl2VpCSJL"
# !python apply_factor.py -i 0 -d 10 --seeds 5,10 --ckpt /content/drive/MyDrive/network-snapshot-008720.pkl /content/ladies-black-cff.pt --output /content/cff-vid/ --video
# + [markdown] id="mzwhrjGlTMZ3"
# That just produced images or video for a single vector, but there are 511 more! To generate every vector, you can uuse the cell below. Update any arguments you want, but don’t touch the `-i {i}` part.
#
# **Warning:** This takes a long time, especially if you have more than one seed value (pro tip: don’t usee more than one seed value)! Also, this will take up a good amount of space in Google Drive. You’ve been warned!
# + id="6aFj6mcKDmqk"
for i in range(512):
# !python apply_factor.py -i {i} -d 10 --seeds 177 --ckpt /content/drive/MyDrive/network-snapshot-008720.pkl /content/ladies-black-cff.pt --output /content/drive/MyDrive/ladiesblack-cff-17/ --video #--out_prefix 'ladiesblack-factor-{i}'
# + id="UVfmNV5JEcdp"
|
SG6_T.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: diagnosis
# language: python
# name: diagnosis
# ---
# +
from __future__ import division
import numpy as np
from catboost import CatBoostClassifier, Pool
import pickle
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedKFold
from tqdm import tqdm
import gc
import copy
import warnings
import random
warnings.filterwarnings('ignore')
# -
age_test = pd.read_csv("../data/age_test.csv", header = None)
age_train = pd.read_csv("../data/age_train.csv", header = None, names=['uId','age'])
age_train['age'].value_counts().plot.bar()
user_basic_info_data = pd.read_csv( r'../data/user_basic_info.csv',header=None,names=['uId','gender','city','prodName','ramCapacity','ramLeftRation',
'romCapacity','romLeftRation','color','fontSize','ct','carrier','os'])
bisic_data = pd.merge(age_train,user_basic_info_data,how='inner',on='uId')
bisic_data
bisic_data.plot.bar(x='age', y='fontSize')
bisic_data['color'].value_counts().plot.bar()
|
src/Data_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.style as style
from matplotlib import collections as mc
import seaborn as sns
import pandas as pd
import scipy.sparse as sps
import scipy.sparse.linalg
style.use('ggplot')
def laplacian_fd(h):
"""Poisson on a 2x2 square, with neumann
boundary conditions f'(x) = 0 on boundary"""
N = int(2/h) + 1
# Vector of x values
x = np.tile(np.linspace(-1, 1, N), N)
# Vector of y values
y = np.repeat(np.linspace(-1, 1, N), N)
# Build LHS
main = -4*np.ones(N**2)
side = np.ones(N**2-1)
side[np.arange(1,N**2)%N==0] = 0
side[np.arange(0,N**2 - 1)%N==0] = 2
up_down = np.ones(N**2-N)
up_down[np.arange(0, N)] = 2
diagonals = [main, np.flip(side), side, np.flip(up_down), up_down]
laplacian = sps.diags(diagonals, [0, -1, 1, -N, N], format="csr")
# Build RHS
rhs = np.cos(np.pi*x)*np.sin(np.pi*y)
return x, y, sps.linalg.spsolve((1/h**2)*laplacian, rhs)
def plot_heatmap(x, y, sol, title):
data = {'x': x, 'y': y, 'solution': sol}
df = pd.DataFrame(data=data)
pivot = df.pivot(index='y', columns='x', values='solution')
ax = sns.heatmap(pivot)
ax.invert_yaxis()
ax = plt.title(title)
# # Test Problem 1
# Laplacian over Square Domain with Neumann Boundary conditions
#
# Consider the equation
#
# $$ u_{xx} + u_{yy} = -cos(\pi x) sin(\pi y)$$
#
# over the square domain $[-1, 1]\times[-1,1]$
# with the Neumann boundary condition
#
# $$ \frac{\partial u(x, y)}{\partial n} = 0 $$
#
# on the boundary.
file = np.loadtxt("square_laplace.txt")
x = file[:, 0]
y = file[:, 1]
sol = file[:, 2]
plot_heatmap(x, y, sol, "Solution Using EB Code")
x, y, solution = laplacian_fd(.125)
plot_heatmap(x, y, solution, "Solution Using Finite Differences")
# # Test Problem 2
# ## a) Neumann Boundary Conditions Over a Circle
#
# $$\nabla^2 \phi = x^2 + y^2 $$
#
# with the Neumann boundary condition, $ \phi(x, y) = 1/4$ on the boundary $x^2 + y^2 = 1$.
#
# The solution is
#
# $$\phi(x, y) = \frac{1}{16}(x^2 + y^2)^2$$
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle2/laplace-%d.txt" % i)
# Interior Cells
inside_cells = data.loc[data['Covered ID'] == 1]
inside_x = inside_cells['CenterX']
inside_y = inside_cells['CenterY']
inside_laplace = inside_cells['Laplacian']
inside_analytic = inside_x**2 + inside_y**2
err = inside_cells['Volume Fraction']*inside_laplace - inside_cells['Volume Fraction']*inside_analytic
norm.append(np.max(err))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Interior Cells, Slope = %f" % slope)
plt.show()
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle2/laplace-%d.txt" % i)
# exterior Cells
exterior_cells = data.loc[data['Covered ID'] == 2]
exterior_x = exterior_cells['CenterX']
exterior_y = exterior_cells['CenterY']
exterior_laplace = exterior_cells['Laplacian']
exterior_analytic = exterior_x**2 + exterior_y**2
err = exterior_cells['Volume Fraction']*(exterior_laplace - exterior_analytic)
norm.append(np.abs(np.max(err)))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Boundary Cells, Slope = %f" % slope)
plt.show()
# ## b) Neuman Boundary Conditions over Circle of Radius $\sqrt{2}$
data = pd.read_csv("laplace_mesh_refine/circle_2rad2/laplace-4.txt")
x = data['CenterX']
y = data['CenterY']
plot_heatmap(x, y, data['Laplacian'], "Circle Radius Root 2 Laplacian")
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle_2rad2/laplace-%d.txt" % i)
exterior_cells = data.loc[data['Covered ID'] == 2]
exterior_x = exterior_cells['CenterX']
exterior_y = exterior_cells['CenterY']
exterior_laplace = exterior_cells['Laplacian']
exterior_analytic = exterior_x**2 + exterior_y**2
err = exterior_cells['Volume Fraction']*(exterior_laplace - exterior_analytic)
norm.append(np.abs(np.max(err)))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Boundary Cells, Slope = %f" % slope)
plt.show()
norm
# ## c) Neumann Boundary Conditions Over Circle Shifted
# ### Volume Moments
data = pd.read_csv("laplace_mesh_refine/circle_origin2/laplace-3.txt")
data = data.loc[data['Covered ID'] > 0]
x = data['CenterX']
y = data['CenterY']
plot_heatmap(x, y, data['Laplacian'] - (x**2 + y**2), "Centered Circle Volume Moments")
def phi(x, y):
return 1/16*(x**2 + y**2)**2
data = pd.read_csv("../laplace_out.txt")
data = data.loc[data['Covered ID'] > 0]
x = data['CenterX']
y = data['CenterY']
#data
plot_heatmap(x, y, data['Laplacian'], "Realigned Volume Moments")
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle_shifted2/laplace-%d.txt" % i)
x = data['CenterX']
y = data['CenterY']
volume = data['Volume Fraction']*data['Cell Size']**2
err = np.abs(np.pi - volume.sum())
norm.append(err)
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Volume Moments, Slope = %f" % slope)
plt.show()
norm
# ### Laplacian
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle_shifted2/laplace-%d.txt" % i)
exterior_cells = data.loc[data['Covered ID'] == 1]
exterior_x = exterior_cells['CenterX']
exterior_y = exterior_cells['CenterY']
exterior_laplace = exterior_cells['Laplacian']
exterior_analytic = exterior_x**2 + exterior_y**2
err = exterior_cells['Volume Fraction']*(exterior_laplace - exterior_analytic)
norm.append(np.abs(np.max(err)))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Boundary Cells, Slope = %f" % slope)
plt.show()
norm
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle_shifted2/laplace-%d.txt" % i)
exterior_cells = data.loc[data['Covered ID'] == 2]
exterior_x = exterior_cells['CenterX']
exterior_y = exterior_cells['CenterY']
exterior_laplace = exterior_cells['Laplacian']
exterior_analytic = exterior_x**2 + exterior_y**2
err = exterior_cells['Volume Fraction']*(exterior_laplace - exterior_analytic)
norm.append(np.abs(np.max(err)))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Boundary Cells, Slope = %f" % slope)
plt.show()
norm
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle_shifted2/laplace-%d.txt" % i)
circumference = data['Boundary Length'].sum()
err = 2*np.pi - circumference
# Throw Out Cells Not In Domain
norm.append(np.abs(err))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Circle Circumference, Slope = %f" % slope)
plt.show()
norm
# ## d) Neumann Boundary Conditions Over an Ellipse
#
# $$\nabla^2 \phi = x^2 + y^2 $$
#
# with the Neumann boundary condition, $ \phi(x, y) = 1/4$ on the boundary $x^2 + 2y^2 = 1$.
#
# The solution is
#
# $$\phi(x, y) = \frac{1}{16}(x^2 + y^2)^2$$
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/ellipse2/laplace-%d.txt" % i)
# Interior Cells
inside_cells = data.loc[data['Covered ID'] == 1]
inside_x = inside_cells['CenterX']
inside_y = inside_cells['CenterY']
inside_laplace = inside_cells['Laplacian']
inside_analytic = inside_x**2 + inside_y**2
err = inside_cells['Volume Fraction']*inside_laplace - inside_cells['Volume Fraction']*inside_analytic
norm.append(np.max(err))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Interior Cells, Slope = %f" % slope)
plt.show()
norm = []
h = []
for i in range(3, 8):
data = pd.read_csv("laplace_mesh_refine/ellipse2/laplace-%d.txt" % i)
exterior_cells = data.loc[data['Covered ID'] == 2]
exterior_x = exterior_cells['CenterX']
exterior_y = exterior_cells['CenterY']
exterior_laplace = exterior_cells['Laplacian']
exterior_analytic = exterior_x**2 + exterior_y**2
err = exterior_cells['Volume Fraction']*(exterior_laplace - exterior_analytic)
norm.append(np.max(np.abs(err)))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Boundary Cells, Slope = %f" % slope)
plt.show()
norm
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/ellipse2/laplace-%d.txt" % i)
volume = (data['Volume Fraction']*data['Cell Size']**2).sum()
err = (1/np.sqrt(2)*np.pi) - volume
# Throw Out Cells Not In Domain
norm.append(np.abs(err))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Ellipse Area, Slope = %f" % slope)
plt.show()
norm
# ## e) Ellipse Flipped
#
# $$\nabla^2 \phi = x^2 + y^2 $$
#
# with the Neumann boundary condition, $ \phi(x, y) = 1/4$ on the boundary $2x^2 + y^2 = 1$.
#
# The solution is
#
# $$\phi(x, y) = \frac{1}{16}(x^2 + y^2)^2$$
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/ellipseflip2/laplace-%d.txt" % i)
# Interior Cells
inside_cells = data.loc[data['Covered ID'] == 1]
inside_x = inside_cells['CenterX']
inside_y = inside_cells['CenterY']
inside_laplace = inside_cells['Laplacian']
inside_analytic = inside_x**2 + inside_y**2
err = inside_cells['Volume Fraction']*inside_laplace - inside_cells['Volume Fraction']*inside_analytic
norm.append(np.max(err))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Interior Cells, Slope = %f" % slope)
plt.show()
norm = []
h = []
for i in range(3, 7):
data = pd.read_csv("laplace_mesh_refine/ellipseflip2/laplace-%d.txt" % i)
exterior_cells = data.loc[data['Covered ID'] == 2]
exterior_x = exterior_cells['CenterX']
exterior_y = exterior_cells['CenterY']
exterior_laplace = exterior_cells['Laplacian']
exterior_analytic = exterior_x**2 + exterior_y**2
err = exterior_cells['Volume Fraction']*(exterior_laplace - exterior_analytic)
norm.append(np.max(np.abs(err)))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Boundary Cells, Slope = %f" % slope)
plt.show()
norm
# # Test Problem 3
# ## Area of a circle
#
# For the circle $x^2 + y^2 = 1$,
#
# $err = |\pi - \text{sum of EB volumes} | $$
1/np.sqrt(2)
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle4/laplace-%d.txt" % i)
volume = (data['Volume Fraction']*data['Cell Size']**2).sum()
err = np.pi - volume
# Throw Out Cells Not In Domain
norm.append(np.abs(err))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Circle Area, Slope = %f" % slope)
plt.show()
# A comparison of circumference
#
# $$err = |2\pi - \textit{sum of EB boundary faces}|$$
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle2/laplace-%d.txt" % i)
circumference = data['Boundary Length'].sum()
err = 2*np.pi - circumference
# Throw Out Cells Not In Domain
norm.append(np.abs(err))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Circle Circumference, Slope = %f" % slope)
plt.show()
norm
# # Test Problem 4
# ## Neumann Boundary Condition On a Circle
#
# Consider the Laplacian over the unit circle
#
# $$ \frac{\partial^2 u}{\partial r^2} + \frac{1}{r}\frac{\partial u}{\partial r} + \frac{1}{r^2}\frac{\partial^2 u}{\partial \theta^2} = 0$$
#
# with boundary conditions $u_r(1, \theta) = sin(\theta)$.
#
# The solution is
#
# $$u(r, \theta) = r sin(\theta)$$
#
# or in cartesian coordinates
#
# $$u(x, y) = y$$
norm = []
h = []
for i in range(2, 7):
data = pd.read_csv("laplace_mesh_refine/circle4/laplace-%d.txt" % i)
# Interior Cells
inside_cells = data.loc[data['Covered ID'] == 1]
inside_x = inside_cells['CenterX']
inside_y = inside_cells['CenterY']
inside_laplace = inside_cells['Laplacian']
inside_analytic = inside_y
err = inside_laplace
norm.append(np.max(err))
h.append(1/2**i)
plt.loglog(h, norm)
plt.ylabel('Error Max')
plt.xlabel('h')
slope = (np.log(norm[-1])-np.log(norm[0]))/(np.log(h[-1])-np.log(h[0]))
plt.title("Convergence of Interior Cells, Slope = %f" % slope)
plt.show()
# # Test Problem X
# ## Dirichlet Boundary Conditions Over a Circle
#
# $$\nabla^2 \phi = x^2 + y^2 $$
#
# with the Dirichlet boundary condition, $ \phi(x, y) = 0$ on the boundary $x^2 + y^2 = 1$.
#
# The solution is
#
# $$\phi(x, y) = \frac{1}{16}(x^2 + y^2)^2 - \frac{1}{16}$$
|
outputs/solver_analysis/LaplacianConvergence.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import scipy as sp
from profit.sur.backend.gp import (kern_sqexp, gp_nll, gp_matrix,
gp_matrix_train, gp_optimize)
from profit.sur.backend.gpytorch import GPyTorchSurrogate
def rosenbrock(x, y, a, b):
return (a - x)**2 + b * (y - x**2)**2
def f(r, u, v):
return rosenbrock((r - 0.5) + u - 5, 1 + 3 * (v - 0.6), a=1, b=3)/2
# +
u = np.linspace(4.7, 5.3, 40)
v = np.linspace(0.55, 0.6, 40)
y = np.fromiter((f(0.25, uk, vk) for vk in v for uk in u), float)
[U,V] = np.meshgrid(u, v)
Y = y.reshape(U.shape)
plt.figure()
plt.contour(U,V,Y)
plt.colorbar()
plt.show()
# +
#%% Generate training data
utrain = u[::5]
vtrain = v[::5]
xtrain = np.array([[uk, vk] for vk in vtrain for uk in utrain])
ytrain = np.fromiter((f(0.25, uk, vk) for vk in vtrain for uk in utrain), float)
ntrain = len(ytrain)
#sigma_meas = 1e-10
#sigma_meas = 1e-5
sigma_meas = 1e-2*(np.max(ytrain)-np.min(ytrain))
#%% Plot and optimize hyperparameters
# hypaplot = np.linspace(0.1,2,100)
# nlls = np.fromiter(
# (gp_nll(hyp, xtrain, ytrain, sigma_meas) for hyp in hypaplot), float)
# plt.figure()
# plt.title('Negative log likelihood in kernel hyperparameters')
# plt.plot(hypaplot, nlls)
# #plt.ylim([-80,-60])
# plt.show()
# + tags=[]
sur = GPyTorchSurrogate()
sur.train(xtrain, ytrain)
xtest = np.array([[uk, vtrain[1]] for uk in u])
ytest = np.fromiter((f(0.25, xk[0], xk[1]) for xk in xtest), float)
ftest = sur.predict(xtest)
plt.figure()
plt.errorbar(xtrain[8:16,0], ytrain[8:16], sigma_meas*1.96, capsize=2, fmt='.')
plt.plot(xtest[:,0], ytest)
plt.plot(xtest[:,0], ftest)
plt.show()
# -
sur.predict(xtrain)
(ytrain-np.mean(ytrain))/np.std(ytrain)
|
draft/mockup_scripts/mockup_sur_gpytorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# imports
import sqlite3
import pandas as pd
# create function to create SQL connection to database
def create_connection(db_file, verbose=False):
conn = None
try:
conn = sqlite3.connect(db_file)
if verbose:
print(f'Using SQLite version: {sqlite3.version}')
print(f'Creating Connection to {db_file}...')
return conn
except sqlite3.Error as e:
print(e)
# create function to query a database
def select_all_query(db_file, query, verbose=False):
conn = create_connection(db_file, verbose)
cur = conn.cursor()
if not query.startswith('SELECT'):
raise ValueError('Query should begin with `SELECT`')
cur.execute(query)
rows = cur.fetchall()
if verbose:
for row in rows:
print(row)
return rows
# How many total Characters are there?
tot_char = select_all_query('rpg_db.sqlite3', 'SELECT COUNT(*) FROM charactercreator_character')
print('Total Characters:', tot_char[0][0])
# +
# How many [characters] of each specific subclass?
# Total clerics
sub_char_cler = select_all_query('rpg_db.sqlite3',
'SELECT COUNT(*) FROM charactercreator_character as cc INNER JOIN \
charactercreator_cleric as cleric on cc.character_id = cleric.character_ptr_id')
print('Total Clerics:', sub_char_cler[0][0])
# -
# Total fighters
sub_char_fight = select_all_query('rpg_db.sqlite3',
'SELECT COUNT(*) FROM charactercreator_character as cc INNER JOIN \
charactercreator_fighter as fighter on cc.character_id = fighter.character_ptr_id')
print('Total Fighters:', sub_char_fight[0][0])
# Total mages (including necromancers)
sub_char_mage = select_all_query('rpg_db.sqlite3',
'SELECT COUNT(*) FROM charactercreator_character as cc INNER JOIN \
charactercreator_mage as mage on cc.character_id = mage.character_ptr_id')
print('Total Mages (includes Necromancers):', sub_char_mage[0][0])
# Total thieves
sub_char_thief = select_all_query('rpg_db.sqlite3',
'SELECT COUNT(*) FROM charactercreator_character as cc INNER JOIN \
charactercreator_thief as thief on cc.character_id = thief.character_ptr_id')
print('Total Thieves:', sub_char_thief[0][0])
# How many total Items?
tot_items = select_all_query('rpg_db.sqlite3',
'SELECT COUNT(*) FROM armory_item')
print('Total Items:', tot_items[0][0])
# +
# How many of the Items are weapons? How many are not?
# Total weapons
tot_weapons = select_all_query('rpg_db.sqlite3',
'SELECT COUNT(*) FROM armory_item as item INNER JOIN armory_weapon \
as weapon on item.item_id = weapon.item_ptr_id')
print('Total Weapons:', tot_weapons[0][0])
# Total non-weapons
tot_non_weapons = select_all_query('rpg_db.sqlite3', 'SELECT (SELECT COUNT(*) FROM armory_item) - \
(SELECT COUNT(*) FROM armory_item as item INNER JOIN armory_weapon as weapon on item.item_id = weapon.item_ptr_id)')
print('Total Non-Weapons:', tot_non_weapons[0][0])
# +
# How many Items does each character have? (Return first 20 rows)
# create query to pull 20 characters with their corresponding item count from database
query = '''SELECT character_id as `Character Id`, COUNT(item_id) as `Item Count`
FROM charactercreator_character_inventory
GROUP BY character_id LIMIT 20;'''
# create conn variable to access database
conn = create_connection('rpg_db.sqlite3')
# create dataframe using pandas read_sql functionality
df = pd.read_sql(query, conn)
df.head()
# +
# How many Weapons does each character have? (Return first 20 rows)
# create query to pull 20 characters with their corresponding item count from database
query = '''SELECT cci.character_id as `Character Id`, COUNT(aw.item_ptr_id) as `Weapon Count`
FROM charactercreator_character_inventory as cci
INNER JOIN armory_item as ai ON cci.item_id = ai.item_id
INNER JOIN armory_weapon as aw ON ai.item_id = aw.item_ptr_id
GROUP BY cci.character_id
LIMIT 20;'''
# create conn variable to access database
conn = create_connection('rpg_db.sqlite3')
# create dataframe using pandas read_sql functionality
df = pd.read_sql(query, conn)
df.head()
# +
# On average, how many Items does each Character have?
# create query to find average items per character
query = '''SELECT AVG(c)
FROM(
SELECT character_id, COUNT(item_id) as c
FROM charactercreator_character_inventory
GROUP BY character_id
)
'''
# connect to db
conn = create_connection('rpg_db.sqlite3')
# create dataframe
df = pd.read_sql(query, conn)
df
# +
# On average, how many Weapons does each character have?
# create query to find average weapons per character
query = '''SELECT AVG(wc)
FROM (SELECT cci.character_id as `Character Id`, COUNT(aw.item_ptr_id) as wc
FROM charactercreator_character_inventory as cci
INNER JOIN armory_item as ai ON cci.item_id = ai.item_id
LEFT JOIN armory_weapon as aw ON ai.item_id = aw.item_ptr_id
GROUP BY cci.character_id)
'''
# connect to db
conn = create_connection('rpg_db.sqlite3')
# create db
df = pd.read_sql(query, conn)
df
# -
# Create database file if it doesn't exist
with sqlite3.connect('buddymove_holidayiq.sqlite3') as conn:
# 1. Read csv file
df = pd.read_csv('buddymove_holidayiq.csv')
# 2. DROP TABLE review IF EXISTS
drop_query = 'DROP TABLE IF EXISTS review'
conn.cursor().execute(drop_query)
# 3. INSERT TABLE review
df.to_sql('review', conn, index=False)
query = 'SELECT * FROM review'
df = pd.read_sql(query, conn)
df.head()
# Count how many rows you have - it should be 249!
df.shape
# +
# How many users who reviewed at least 100 `Nature` in the category also
# reviewed at least 100 in the `Shopping` category? - 78
# query
query = '''SELECT *
FROM review
WHERE `Nature` >= 100 AND `Shopping` >= 100
'''
# connect
conn = create_connection('buddymove_holidayiq.sqlite3')
# create dataframe, run df.describe() to obtain count
df = pd.read_sql(query, conn)
df.describe()
|
module1-introduction-to-sql/assignment1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: DESI 19.12
# language: python
# name: desi-19.12
# ---
# # Simulating DESI Spectra
#
# The goal of this notebook is to demonstrate how to generate some simple DESI spectra using the `quickspectra` utility. For simplicity we will only generate 1D spectra and skip the more computationally intensive (yet still instructive!) step of extracting 1D spectra from simulated 2D spectra (*i.e.*, so-called "pixel-level simulations"). In this tutorial we will:
#
# * generate 100 random QSO spectra
# * simulate them under dark time conditions
# * plot the truth and the noisy simulated spectra
# * run redshift fitting
# * re-simulate when the moon is quite bright
# * re-run redshift fitting
# * compare redshift performance with and without moon
#
# The heart of `quickspectra` is the `SpecSim` package, which you can read about here:
# http://specsim.readthedocs.io/en/stable
#
# If you identify any errors or have requests for additional functionality please create a new issue on
# https://github.com/desihub/desisim/issues
# or send a note to <<EMAIL>>.
# ## Getting started.
#
# See https://desi.lbl.gov/trac/wiki/Computing/JupyterAtNERSC to configure a jupyter server at NERSC with pre-installed DESI code. This notebook was tested with the "DESI 19.12" kernel.
#
# Alternately, see https://desi.lbl.gov/trac/wiki/Pipeline/GettingStarted/Laptop for instructions to install code locally.
#
# First, import all the package dependencies.
# +
import os
import numpy as np
from astropy.io import fits
from astropy.table import Table
# -
import desisim.templates
import desispec.io
# This import of `geomask` is a temporary hack to deal with an issue with the matplotlib backend in the 0.28.0 version of `desitarget`.
from desitarget import geomask
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Simulate with quickspectra
#
# The simplest way to simulate spectra is using the `quickspectra` script. We'll generate a set of noiseless template spectra, save them to a file, and then run `quickspectra` to simulate noise and write out a file that can be used as input for redshift fitting.
# ### Start by simulating some QSO spectra
qso_maker = desisim.templates.SIMQSO()
# %time flux, wave, meta, objmeta = qso_maker.make_templates(nmodel=100)
# What are the outputs?
# * `flux[nspec, nwave]` 2D array of flux [1e-17 erg/s/cm2/A]
# * `wave[nwave]` 1D array of observed-frame (vacuum) wavelengths corresponding to `flux`
# * `meta` table of basic metadata about the targets that's independent of the target type (e.g., redshift).
# * `objmeta` table of target-specific metadata (e.g., QSO emission-line flux strengths).
print('flux.shape', flux.shape)
print('wave.shape', wave.shape)
print('meta.colnames', meta.colnames)
print('objmeta.colnames', objmeta.colnames)
# Note that the (unique) `TARGETID` column can be used to sync up the `meta` and `objmeta` columns when simulating a mixture of target types.
# +
plt.figure(figsize=(9,4))
plt.subplot(121)
plt.hist(meta['REDSHIFT'], 20, (0,5))
plt.xlabel('redshift')
plt.subplot(122)
mag_g = 22.5 - 2.5 * np.log10(meta['FLUX_G'])
plt.hist(mag_g, 20, (15, 25))
plt.xlabel('g magnitude')
# -
# ### Write those to a file and run quickspectra
simdir = os.path.join(os.environ['SCRATCH'], 'desi', 'simspec')
os.makedirs(simdir, exist_ok=True)
infile = os.path.join(simdir, 'qso-input-spectra.fits')
hdr = fits.Header()
hdr['EXTNAME'] = 'WAVELENGTH'
hdr['BUNIT'] = 'Angstrom'
fits.writeto(infile, wave, header=hdr, overwrite=True)
hdr['EXTNAME'] = 'FLUX'
hdr['BUNIT'] = '10^-17 erg/(s*cm^2*Angstrom)' # Satisifes FITS standard AND Astropy-compatible.
fits.append(infile, flux, header=hdr)
specoutfile = os.path.join(simdir, 'qso-observed-spectra.fits')
cmd = 'quickspectra -i {} -o {}'.format(infile, specoutfile)
print(cmd)
# !$cmd
# ### Let's see what we got
spectra = desispec.io.read_spectra(specoutfile)
# +
from scipy.signal import medfilt
def plotspec(spectra, i, truewave=None, trueflux=None, nfilter=11):
plt.plot(spectra.wave['b'], medfilt(spectra.flux['b'][i], nfilter), 'b', alpha=0.5)
plt.plot(spectra.wave['r'], medfilt(spectra.flux['r'][i], nfilter), 'r', alpha=0.5)
plt.plot(spectra.wave['z'], medfilt(spectra.flux['z'][i], nfilter), 'k', alpha=0.5)
if truewave is not None and trueflux is not None:
plt.plot(truewave, trueflux[i], 'k-')
plt.axhline(0, color='k', alpha=0.2)
ymin = ymax = 0.0
for x in ['b', 'r', 'z']:
tmpmin, tmpmax = np.percentile(spectra.flux[x][i], [1, 99])
ymin = min(tmpmin, ymin)
ymax = max(tmpmax, ymax)
plt.ylim(ymin, ymax)
plt.ylabel('flux [1e-17 erg/s/cm2/A]')
plt.xlabel('wavelength [A]')
# plotspec(spectra, 0, wave, flux)
# -
plt.figure(figsize=(12, 9))
for i in range(9):
plt.subplot(3, 3, i+1)
plotspec(spectra, i, wave, flux)
# ## Fit redshifts
#
# Next we'll run the redrock redshift fitter (`rrdesi`) on these spectra.
#
# If at NERSC, run this via an interactive batch node so that we don't abuse the single jupyter server node.
#
# **Note**: if this step doesn't work, check your .bashrc.ext, .bash_profile.ext, or .tcshrc.ext files to see if you are defining
# an incompatible python / desi version that could be overriding the
# environment of this notebook after the job is launched.
zoutfile = os.path.join(simdir, 'qso-zbest.fits')
cmd = 'rrdesi {} --zbest {}'.format(specoutfile, zoutfile)
if 'NERSC_HOST' in os.environ:
print('Running on a batch node:')
print(cmd)
print()
srun = 'srun -A desi -N 1 -t 00:10:00 -C haswell --qos interactive'
cmd = '{srun} {cmd} --mp 32'.format(srun=srun, cmd=cmd)
# !$cmd
zbest = Table.read(zoutfile, 'ZBEST')
plt.plot(meta['REDSHIFT'], zbest['Z'], '.')
plt.xlabel('true redshift'); plt.ylabel('fitted redshift')
# ### Re-simulate with the moon up and at a higher airmass
specoutfile_moon = os.path.join(simdir, 'qso-moon-spectra.fits')
cmd = 'quickspectra -i {} -o {} --moonfrac 0.9 --moonalt 70 --moonsep 20 --airmass 1.3'.format(
infile, specoutfile_moon)
print(cmd)
# !$cmd
zoutfile_moon = os.path.join(simdir, 'qso-zbest-moon.fits')
cmd = 'rrdesi {} --zbest {}'.format(specoutfile_moon, zoutfile_moon)
if 'NERSC_HOST' in os.environ:
print('Running on a batch node:')
print(cmd)
print()
srun = 'srun -A desi -N 1 -t 00:10:00 -C haswell --qos interactive'
cmd = '{srun} {cmd} --mp 32'.format(srun=srun, cmd=cmd)
print(cmd)
# !$cmd
zbest_moon = Table.read(zoutfile_moon, 'ZBEST')
# +
plt.figure(figsize=(9,9))
plt.subplot(221)
plt.plot(meta['REDSHIFT'], zbest['Z'], '.')
plt.ylabel('fitted redshift')
plt.title('no moon')
plt.subplot(222)
plt.plot(meta['REDSHIFT'], zbest_moon['Z'], '.')
plt.title('with moon')
plt.subplot(223)
dv = 3e5*(zbest['Z'] - meta['REDSHIFT'])/(1+meta['REDSHIFT'])
plt.plot(meta['REDSHIFT'], dv, '.')
plt.ylim(-1000, 1000)
plt.ylabel('dv [km/s]')
plt.xlabel('true redshift')
plt.subplot(224)
dv = 3e5*(zbest_moon['Z'] - meta['REDSHIFT'])/(1+meta['REDSHIFT'])
plt.plot(meta['REDSHIFT'], dv, '.')
plt.ylim(-1000, 1000)
plt.xlabel('true redshift')
# -
# Unsurprisingly, it is harder to fit a redshift on a spectrum polluted with a lot of moonlight
# ## Exercises
# 1. Run `help(qso_maker.make_templates)` to see what other options
# are available for generating QSO templates. Try adjusting the magnitude
# or redshift ranges and resimulating
#
# 2. This tutorial used `desisim.templates.SIMQSO()` to generate QSO templates. There are also template generators for `ELG`, `LRG`, `BGS`, `STD`, `MWS_STAR`, `STAR`, `WD`; run `help(desisim.templates)` for details. Try generating other template classes and studying their redshift efficiency.
#
# 3. Simulate more QSOs and study their efficiency vs. S/N or g-band magnitude.
# ## Appendix: Code versions
from desitutorials import print_code_versions
print("This tutorial last ran successfully to completion using the following versions of the following modules:")
print_code_versions()
# ## Appendix: other spectro simulators
#
# This tutorial focused on quickspectra, which simulates spectra outside of the context
# of the full spectroscopic pipeline. Under the hood of this script is [specsim](http://specsim.readthedocs.io/en/stable), which has many more options, e.g. for adjusting input fiberloss fractions based upon object sizes. See the [specsim tutorials](https://github.com/desihub/specsim/tree/master/docs/nb) for details.
#
# Note: the [minitest notebook](https://github.com/desihub/desitest/blob/master/mini/minitest.ipynb) in the [desitest](https://github.com/desihub/desitest) has instructions for the full end-to-end chain covering survey simulations, mocks, fiber assignment, spectral simulation, running the DESI spectro pipeline, and ending with a redshift catalog. But that takes ~2 hours to run and consumes ~1500 MPP hours at NERSC, so it is primarily used for reference and integration testing.
|
simulating-desi-spectra.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importing the standard libraries
# +
import matplotlib.pyplot as plt
from imutils import paths
import numpy as np
import pathlib
import imutils
import pickle
import cv2
import os
# %matplotlib inline
# -
# Above, we have the matplotlib and OpenCV (cv2) libraries for image processing, Numpy library will be used to work with arrays and vectors because the images will be treated as numbers, corresponding to their pixel values. The libraries Imutils, Pathlib and OS will be essential to make the verification of directories.
BASE_PATH = 'dataset'
# The directory named BASE_PATH has the name of the folder that contains the other two folders with the images that will be used and the .csv files of the bounding boxes of each image. In the same folder where "dataset" is located we have this .ipynb file.
IMAGE_PATH = os.path.sep.join([BASE_PATH, 'images'])
ANNOTS_PATH = os.path.sep.join([BASE_PATH, 'annotations'])
# So, using the OS library method, we were able to put together the paths to the folder for the images and the .csv files by associating the IMAGE_PATH and ANNOTHS_PATH variables with the directories to the required files.
# # Importing the framework and his tools
# +
import tensorflow as tf
from tensorflow.keras.applications import VGG16
from tensorflow.keras.layers import Flatten, Dropout, Dense, Input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.preprocessing.image import load_img
from tensorflow.keras.utils import to_categorical
from sklearn.preprocessing import LabelBinarizer
from sklearn.model_selection import train_test_split
# -
# The Machine Learning platform we will use is TensorFlow and along with it the Keras API. We will test the use of VGG 16 in our prototype, and we will download it from the framework, and we should also import the functions that we will use in building our neural network:
#
# 👉🏼 Flatten to turn it into a 1-dimensional layer,
#
# 👉🏼 Dropout to speed up the learning processing,
#
# 👉🏼 Dense to create the "Fully-Connected Layers",
#
# 👉🏼 Input to define the inputs in our VGG 16.
# We also have other functions for image processing like img_to_array and load_img, and also the ADAM optimizer.
#
# Finally we will use sklearn to split our data into training and testing, and to categorize the classes in one-hot form.
data = []
labels = []
bboxes = []
imagePaths = []
# Here we initialize empty lists that will hold some important information in the course of the prototype:
#
#
# 👉🏼 data: Images,
#
# 👉🏼 labels: Categories,
#
# 👉🏼 bboxes: Bounding Box coordinates (x, y),
#
# 👉🏼 imagePaths: The directory of the images.
# Now we will go through our .csv files that correspond to the images we are using:
#
# 🔸 First column: Image name,
#
# 🔸 Second column: Category,
#
# 🔸 Third column: Bounding box (bbox) x initial coordinate,
#
# 🔸 Fourth column: initial bbox y coordinate,
#
# 🔸 Fifth column: end bbox x coordinate,
#
# 🔸 Sixth column: final bbox y coordinate.
for csvPath in paths.list_files(ANNOTS_PATH, validExts=(".csv")):
rows = open(csvPath).read().strip().split("\n")
for row in rows:
row = row.split(",")
(filename, label, startX, startY, endX, endY) = row
imagePath = os.path.sep.join([IMAGE_PATH, label, filename])
image = cv2.imread(imagePath)
(h, w) = image.shape[:2]
startX = float(startX) / w
startY = float(startY) / h
endX = float(endX) / w
endY = float(endY) / h
image = load_img(imagePath, target_size=(224, 224))
image = img_to_array(image)
data.append(image)
labels.append(label)
bboxes.append((startX, startY, endX, endY))
imagePaths.append(imagePath)
# Above, the variable 'row' will store the values taken from the .csv that was performed externally, an example of each of the categories used can be seen below:
#
#
# 🔸 ['145_0054_jpg.rf.5d469949fa7bbacd2a2d996583abede5.jpg', 'Motorbikes', '11', '32', '184', '191']
#
#
# 🔸 ['159_0009_jpg.rf.0016fb51718b8b1b6a997c70fb920aa6.jpg', 'People', '115', '17', '199', '158']
#
#
# 🔸 ['246_0012_jpg.rf.298ce90186d7705adb835b93f2f21903.jpg', 'Wine_Bottle', '11', '5', '92', '63']
#
#
# Above, the imagePath variable will have stored the location for us to access the images, an example of each of the categories used can be seen below:
#
#
# 🔸 dataset\images\Motorbikes\145_0054_jpg.rf.5d469949fa7bbacd2a2d996583abede5.jpg
#
#
# 🔸 dataset\images\People\159_0009_jpg.rf.0016fb51718b8b1b6a997c70fb920aa6.jpg
#
#
# 🔸 dataset\images\Wine_Bottle\246_0012_jpg.rf.298ce90186d7705adb835b93f2f21903.jpg
#
#
# So, having the directories, we will "read" them with OpenCV and store them in the variable "image", from where we will be able to extract the characteristics we are interested in as the size of the image and transform the image into an array.
#
# Finally, all the features that are extracted from each image are added to the previously empty lists.
#
# Remember that we will make a change in the size of the images to use VGG 16 CNN.
# # Data Processing
# +
data = np.array(data, dtype="float32") / 255.0
labels = np.array(labels)
bboxes = np.array(bboxes, dtype="float32")
imagePaths = np.array(imagePaths)
lb = LabelBinarizer()
labels = lb.fit_transform(labels)
# -
# Here, for each list of values we will work in a different way. For the list that contains the images, we will have to divide the arrays by 255 for the normalization of the data, because the pixel values go from 0 to 255, and leaving 0 to 1 will speed up the learning process.
#
# In the labels list we have the categories, which when we perform the fit_transform method with the LabelBinarizer function from Scikit-learn we will have the one-hot encoding. An example for the category Motorbikes is: [1, 0, 0], for People it is: [0, 1, 0], and for Wine_Bottle is: [0 , 0, 1].
#
# In the list bboxes we have the coordinates of the bounding boxes of the images to be used in the prototype.
#
# And finally, in the list imagePaths we have the directories of the images.
# +
split = train_test_split(data, labels, bboxes, imagePaths, test_size = 0.20, random_state = 42)
(trainImages, testImages) = split[:2]
(trainLabels, testLabels) = split[2:4]
(trainBBoxes, testBBoxes) = split[4:6]
(trainPaths, testPaths) = split[6:]
# -
# In the variable "split", we will have stored the outputs from the train_test_split method of Scikit-learn, for splitting our data.
# # Model Building
# Below, we will see that two variables have been set that contain, respectively, the number of learning "epochs" and the size of the training batch, the latter usually being a power of 2.
#
#
# In the following lines we are loading the classic VGG 16 neural net with the pre-trained weights and parameters from the ImageNet database, leaving out the Fully Connected layers with the command "include_top = False", so that we can build new 1-dimensional layers responsible for finding the categories we will work in our database.
#
#
# Right afterwards we set our VGG 16, because we will not train it to update its parameters during learning.
#
#
# We will build the Fully Connected layers for the Bounding Box coordinate predictions. And then below that the Fully Connected layers for category predictions.
# First, bboxHead is responsible for predicting the bounding box (x, y)-coordinates of the object to be categorized in the image. We have a Fuly Connected layer consisting of 128, 64, 32, and 4 nodes, respectively.
#
#
#
# The most important part for the bounding box predictions is the final layer:
#
#
#
# 👉🏼 The 4 nodes corresponding to the coordinates for the top left and top right of the Bounding Box.
#
#
# 👉🏼 We use a sigmoid function to ensure that our predicted output values are in the range [0,1].
#
#
# Then, softmaxHead, is responsible for predicting the category of the detected object.
# +
import warnings
warnings.filterwarnings('ignore')
NUM_EPOCHS = 20
BATCH_SIZE = 32
vgg = VGG16(weights = "imagenet", include_top = False, input_tensor = Input(shape = (224, 224, 3)))
vgg.trainable = False
flatten = vgg.output
flatten = Flatten()(flatten)
# Fully-connected layer - bounding box
bboxHead = Dense(128, activation = "relu")(flatten)
bboxHead = Dense(64, activation = "relu")(bboxHead)
bboxHead = Dense(32, activation = "relu")(bboxHead)
bboxHead = Dense(4, activation = "sigmoid", name = "bounding_box")(bboxHead)
# Fully-connected layer - categorias
softmaxHead = Dense(512, activation = "relu")(flatten)
softmaxHead = Dropout(0.5)(softmaxHead)
softmaxHead = Dense(512, activation = "relu")(softmaxHead)
softmaxHead = Dropout(0.5)(softmaxHead)
softmaxHead = Dense(len(lb.classes_), activation = "softmax", name = "class_label")(softmaxHead)
model = Model(inputs=vgg.input, outputs = (bboxHead, softmaxHead))
losses = {"class_label": "categorical_crossentropy", "bounding_box": "mean_squared_error",}
lossWeights = {"class_label": 1.0, "bounding_box": 1.0}
opt = Adam(lr = 0.0001)
# -
# Furthermore, we have to create some dictionaries that will store important information from our model, especially with respect to the loss. So, the "loss" for the categories, given that we have one-hot coding is "categororial_crossentropy", while the "loss" for the bounding boxes is "mean_squared_error".
#
# We have a "lossWeights" dictionary that will tell the framework how to "weight" each of the layers during training, at an equal rate for both categories and bounding boxes.
#
# We will initialize the ADAM optimizer with the Learning Rate equal to 0.0001
#
# With the optimizer initialized, we compile the model and check the built neural network with model.summary().
model.compile(loss = losses, optimizer = opt, metrics = ["accuracy"], loss_weights = lossWeights)
print(model.summary())
# We will also build two more dictionaries to store the training and the test.
# +
trainTargets = {"class_label": trainLabels,"bounding_box": trainBBoxes}
testTargets = {"class_label": testLabels,"bounding_box": testBBoxes}
# -
# # Fit the Model
H = model.fit(trainImages, trainTargets, validation_data = (testImages, testTargets),
batch_size = BATCH_SIZE, epochs = NUM_EPOCHS, verbose = 1)
# After we trained the model, we will make graphs for performance analysis and save the model.
model.save('Modelo Detecção de Objetos v_01', save_format = 'h5')
# +
lossNames = ["loss", "class_label_loss", "bounding_box_loss"]
N = np.arange(0, NUM_EPOCHS)
plt.style.use("ggplot")
(fig, ax) = plt.subplots(3, 1, figsize = (13, 13))
for (i, l) in enumerate(lossNames):
title = "Loss for {}".format(l) if l != "loss" else "Total loss"
ax[i].set_title(title)
ax[i].set_xlabel("Epoch #")
ax[i].set_ylabel("Loss")
ax[i].plot(N, H.history[l], label=l)
ax[i].plot(N, H.history["val_" + l], label = "val_" + l)
ax[i].legend()
# -
plt.style.use("ggplot")
plt.figure(figsize = (13, 13))
plt.plot(N, H.history["class_label_acc"], label = "class_label_train_acc")
plt.plot(N, H.history["val_class_label_acc"], label = "val_class_label_acc")
plt.title("Class Label Accuracy")
plt.xlabel("Epoch #")
plt.ylabel("Accuracy")
plt.legend(loc = "lower left")
# # Data Analysis
# To start predictions analysis of our model, let's look at how many test examples we have:
len(testImages)
# What we can do is plot some test images to visualize their predictions. For this we created a function that uses a dictionary for the categories according to the value taken from "testLabels".
SAMPLE = 40
# +
classes = {0: 'Motorbike',
1: 'People',
2: 'Wine Bottle'}
def plt_sample(X, y, index):
plt.figure(figsize = (8, 6))
plt.imshow(X[index])
class_sample = np.argmax(y[index])
plt.xlabel(classes[class_sample])
# -
plt_sample(testImages, testLabels, SAMPLE)
# We then see that the 41st image in testImages corresponds to a Motorbike, remembering that the label shown on the axis is the original label.
# We will now make our model's predictions on the test images.
(boxPreds, labelPreds) = model.predict(testImages)
# Above we see the method responsible for making the predictions, in our case we have to predict the class to which the object to be detected belongs, as well as the position of the bounding box.
# +
classPreds = [np.argmax(element) for element in labelPreds]
print('The predicted category is: ', classPreds[SAMPLE])
label_test = classes[classPreds[SAMPLE]]
print('It corresponds to: {}'.format(label_test))
# -
# So, we just need to change the value of the variable SAMPLE to a range between 0 and 136, which are our test images, so we can check if the category predictions were correct with respect to the image we are analyzing.
#
# For the Bounding box we have to take a list that will contain the predicted coordinates of the model.
(startX, startY, endX, endY) = boxPreds[SAMPLE]
# Uma forma de melhorar a precisão da bounding box, faremos um resize na imagem.
# +
Img = testImages[SAMPLE]
Img = imutils.resize(Img, width=600)
plt.imshow(Img)
# -
# Extracting the values for the Bounding Box calculations, we have:
(h, w) = Img.shape[:2]
startX = int(startX * w)
startY = int(startY * h)
endX = int(endX * w)
endY = int(endY * h)
# +
y = startY - 10 if startY - 10 > 10 else startY + 10
cv2.putText(Img, label_test, (startX, y), cv2.FONT_HERSHEY_SIMPLEX, 0.65, (0, 255, 0), 2)
cv2.rectangle(Img, (startX, startY), (endX, endY),(0, 255, 0), 2)
plt.figure(figsize = (12,10))
plt.imshow(Img)
# -
# # One more example
# +
SAMPLE = 1
classes = {0: 'Motorbike',
1: 'People',
2: 'Wine Bottle'}
def plt_sample(X, y, index):
plt.figure(figsize = (8, 6))
plt.imshow(X[index])
class_sample = np.argmax(y[index])
plt.xlabel(classes[class_sample])
# -
plt_sample(testImages, testLabels, SAMPLE)
# +
classPreds = [np.argmax(element) for element in labelPreds]
print('The predicted category is: ', classPreds[SAMPLE])
label_test = classes[classPreds[SAMPLE]]
print('It corresponds to: {}'.format(label_test))
# +
(startX, startY, endX, endY) = boxPreds[SAMPLE]
Img = testImages[SAMPLE]
Img = imutils.resize(Img, width=600)
(h, w) = Img.shape[:2]
startX = int(startX * w)
startY = int(startY * h)
endX = int(endX * w)
endY = int(endY * h)
y = startY - 10 if startY - 10 > 10 else startY + 10
cv2.putText(Img, label_test, (startX, y), cv2.FONT_HERSHEY_SIMPLEX, 0.65, (0, 255, 0), 2)
cv2.rectangle(Img, (startX, startY), (endX, endY),(0, 255, 0), 2)
plt.figure(figsize = (12,10))
plt.imshow(Img)
|
Project/Multi-class object detection and bounding box regression with Keras, TensorFlow, and Deep Learning V_02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Abstract
#
# A Census official would like to use a given number of factors to predict salary of more than 50K. He hires consultants who would create a model that can predict salary and that has a good accuracy score.
#
#
# # Introduction
#
# As the consultants we have data from the Census department. The dataset has categorical data so we hope to use a classification model to predict the salary given a set of features. This a binary problem, where salary is classified as either equal to or less than 50K, or classified as greater than 50K.
#
#
# ### Problem
# For the given Census data can we predict whether a person will earn <=50K or >50?
#
# ### Solution
# Create a classification regression that predicts salary.
#
# ## Goal
# Train a classification model than performs better than 0.75 accuracy.
#
#
# # Research Questions
# - Which model has the best accuracy test score?
# - What are some of the ways we can reduce bias?
# - How do we test our model?
# ## DataSet
#
# +
# import numpy, pandas, matplotlib, seaborn and sklearn
import numpy as np
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn; seaborn.set()
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
# -
# ### Open data file
#
# This data file is a .data there we put a delimiter to open it. It also does not have a column header.
# Using information on the attributes from the source website we make a column header.
# +
# Open data file
# Since it is a .data file it has a , delimiter
# Need to add columns names as it does not have column names
df=pd.read_csv("adult.data", delimiter=",", names=['age','workclass','fnlwgt','education','edu_num','marital_status','occupation','relationship','race','sex','capital_gain','capital_loss','hours_per_week','native_country','salary'])
df.head(5)
# -
df.shape
# +
# This is the Target variable
# Check the ratio of salary
df["salary"].value_counts()
# +
# Model accuracy rate
24720/(24720+7841)
# -
# #### Define the X and Y variables.
# +
# Define the X and Y variables
y = df.loc[:, "salary"]
X = df.loc[:, ['age','workclass','fnlwgt','education','edu_num','marital_status','occupation','relationship','race','sex','capital_gain','capital_loss','hours_per_week','native_country']]
X.head(2)
# -
y.value_counts()
# # Exploratory Data Analysis
# %matplotlib inline
import matplotlib.pyplot as plt
X.hist(bins=50, figsize=(20,15))
plt.show()
# In the graphs above, the age graph is skewed towards the right, which is expected as this is data of a working population. It also has some spikes, however they follow the general shape of the curve.
#
# The graphs for the capital gain and capital loss both indicate that majority of the people have no capital gain or loss.
#
# The education numbers indicate the level of education, these show that most peole have a high school education, followed by some college education.
#
# Most people work 40 hours a week which is expected for most full time jobs.
#
# The fnlwgt is a reference number so that will be dropped.
X.groupby('education')[['edu_num']].count()
# +
# Make a graph for y
plt.hist(y)
plt.title("Distribution of Salary")
plt.ylabel("Numbers")
# -
# From the data it is evident that there are way more people who earn equal to or less than 50K.
seaborn.barplot(x="sex",y="hours_per_week", hue="salary", data=df)
plt.ylabel('Hours per week')
plt.title('Hours worked per week')
# This graphs shows that men tend to earn alittle more money than women and also work afew more hours per week.
seaborn.barplot(x="salary",y="hours_per_week", hue="sex", data=df)
plt.ylabel('Hours per week')
plt.title('Hours worked per week')
# People who earn more than 50K tend to work afew more hours per week.
# # Data Cleaning
# ### Check for missing data
# +
# Check for missing values
df.info()
# -
# The data above indicates that there is no missing value.
# +
# Check for outliers
df.edu_num.plot(kind = 'box')
# -
# This shows the lower level of education has outliers, however it is possible in a population to find afew people who have a very low level of education probably because of a disability or other disadvantages.
# # Feature Engineering
# +
# Ratio of <=50K and >50K
24720/(24720+7841)
# -
# ### Create Feature
#
# To get the net gain it is important to subtract capital_gain from capital_loss this will give us the capital net gain. So we create a column capital_net.
# +
X['capital_net'] = X['capital_gain']-X['capital_loss']
X.head(5)
# -
# ### Drop columns
#
# From our exploratory analysis we see that we will need to drop the `fnlwgt` column. This is because this are just reference numbers.We will also drop the `relationship` column because it is very similar to the `marital_status`, therefore it is redundant. The `education` column will be dropped, because it is the same as the `edu-num` column, which represents the education levels with a number. Since, we have generated the column `capital_net`, we can now drop `capital_gain` and `capital_loss`.
# +
# Drop columns
X.drop(['fnlwgt','education','relationship','capital_gain','capital_loss'], axis=1, inplace=True)
X.head(5)
# -
# ### Transform Data
# ### Transform Y
#
# Y has categorical data <=50K and >50K, we need to change the y data to numerical values. We use the LabelEncoder
# +
# Convert target data into numerical values
le = LabelEncoder()
# +
# Fit label encoder to y
le.fit(y)
# -
# Transform y
le.transform(y)
# +
# Class names
le.classes_
# -
# ### OneHot Encoder
#
# The X data is converted from categorical data in X to numeric using the OneHot Encoder from sklearn. The OneHot Encoder ensures that the categorical features do not become ordered or ranked when transformed from strings to numerical format.
# The columns `workclass`,`occupation`,`race`,`sex`,`native_country`, are all strings and need to be changed to numerical values, in order to perform a classification regression. The mentioned columns are in the X target variables.
# +
# import OneHotEncoder
##
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
# +
# instantiate
## encoder = OneHotEncoder(drop = None, categories = 'auto')
##
columnTransformer = ColumnTransformer([OneHotEncoder(drop = None, categories = 'auto')])
# +
##X=np.array(columnTransformer.fit_transform(X), dtype = np.str)
encoder = OneHotEncoder()
encoder.fit(X)
# +
X = encoder.transform(X)
# -
# Looking at X we see that it is now transformed to numpy array by OneHotEncoder.
# Looking at X
X
# ### Splitting Data
# +
# Use train-test-split already imported at the beginning
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.2, random_state = 1, stratify = y)
# -
X_train.shape
X_test.shape
np.unique(y_train, return_counts = True)
19775/(19775+6273)
np.unique(y_test, return_counts = True)
# # Modeling
# ## Logistic Regression
# import Logistic Regression from sklearn
from sklearn.linear_model import LogisticRegression
# +
# Already imported Logistic Regression from sklearn
# Instantiate LR with no penalty
lr = LogisticRegression(penalty = 'none', max_iter = 5000)
# +
# fit model
lr.fit(X_train, y_train)
# +
# Learned the coefficients
lr.coef_
# +
# Check the intercept
lr.intercept_
# +
# Score ## Accuracy
lr.score(X_train, y_train)
# +
# Score ## Accuracy
score=lr.score(X_test, y_test)
print(score)
# -
# ### Confusion Matrix for Logistic Regression
# +
# from sklearn import confusion matrix
from sklearn.metrics import confusion_matrix
# +
# Create a confusion metrics
# -
y_pred = lr.predict(X_test)
# +
# Check prediction
y_pred = lr.predict(X_test[1:10])
print(y_pred)
# -
cm = confusion_matrix(y_test, y_pred)
cm
# +
# Plot the confusion matrix
seaborn.heatmap(cm, annot=True, fmt=".2f", cmap=plt.cm.Reds)
plt.title('Logistic regression \n Accuracy Score {0}'.format(score), fontsize=14)
plt.xlabel(('Predicted'), fontsize=14)
plt.ylabel(('Actual'), fontsize=14)
# -
# ### Cross Validation for Logistic Regression
# +
# from sklearn.model_selection import cross_validate
from sklearn.model_selection import cross_validate
# +
# call cross-validate with return_train_score, return estimator, cv = 5
# -
estimator = LogisticRegression(penalty = 'none', max_iter = 10000)
cv_fivefolds = cross_validate(estimator = estimator,
X = X_train,
y = y_train,
cv = 5,
return_estimator = True,
return_train_score = True, verbose = 2)
# +
# investigate cv results
cv_fivefolds['train_score']
# -
cv_fivefolds['test_score']
# +
# Find the mean and standard deviation for cv
lrvalidation_mean = cv_fivefolds['test_score'].mean()
lrvalidation_std = cv_fivefolds['test_score'].std()
# -
print('Logistic Regression 5-fold cv results %.3f =/- %.3f'%(lrvalidation_mean, lrvalidation_std))
# ## Decision Trees
# +
# from sklearn import Decision Tree classifier
from sklearn.tree import DecisionTreeClassifier
# +
# instantiate
dt_clf = DecisionTreeClassifier()
# +
# fit the model
dt_clf.fit(X_train, y_train)
# +
# Score ## accuracy
dt_clf.score(X_train, y_train)
# +
# If we check test score
treescore = dt_clf.score(X_test, y_test)
print(treescore)
# -
# ### Confusion Matrix for Decision Tree
# +
# Create a confusion matrix
# -
y_pred_tree = dt_clf.predict(X_test)
cm_tree = confusion_matrix(y_test, y_pred_tree)
cm_tree
# +
# Plot the confusion matrix
seaborn.heatmap(cm_tree, annot=True, fmt=".2f", cmap=plt.cm.copper)
plt.title('Decision Tree \n Accuracy Score {0}'.format(treescore), fontsize=14)
plt.xlabel(('Predicted'), fontsize=14)
plt.ylabel(('Actual'), fontsize=14)
# -
# ### Cross Validation of DecisionTrees
# +
# Cross Validate
# -
cv_fivefold = cross_validate(estimator = dt_clf,
X = X_train,
y = y_train,
cv = 5,
return_train_score = True,
return_estimator = True,
verbose = 2)
# +
# investigate cv results for Decision Tree
cv_fivefold['train_score']
# -
cv_fivefold['test_score']
# +
# Find the mean and standard for cv
dtvalidation_mean = cv_fivefold['test_score'].mean()
dtvalidation_std = cv_fivefold['test_score'].std()
# -
# Print results
print('Decision Tree 5-fold cv results %.3f =/- %.3f'%(validation_mean, validation_std))
# ### Evaluating Model Performance
#
#
# Which model performed best? We put the score from the 5-fold cross validate into a table to answer the question.
#
# +
model_performance = pd.DataFrame({
"Model":["Logistic Regression","Decision Tree"],
"Validation Mean":[lrvalidation_mean, dtvalidation_mean],
"Validation Standard deviation":[lrvalidation_std, dtvalidation_std]
})
model_performance.sort_values(by = "Validation Mean", ascending = False)
# -
# # Conclusion
# ### Goal
#
#
# Our goal was to train a classification model that performs better than 0.75 accuracy. We were able to achieve this goal for both the Logistic Regression and Decision Tree. Using the cross-validation the Logistic regression had a mean validation test score of 0.87 and the Decision Tree has a validation test score mean of 0.83.
#
# Using the confusion matrix, the Logistic regression was able to predict correctly 5664 of 6513 of the test data. The Decision tree on the other hand predicted 5475 of the 6513 of the test data. Therefore, the Logistic Regression predict more correctly.
#
# ### Research question
#
# - Which model has the best accuracy score?
#
# On the training data the Logistic Regression had an accuracy score of 0.877, while the Decision Tree had a score of 0.978. However on the test score, using the cross-validation table, the Logistic Regression had a train score of about 0.871 with a standard deviation of +/- 0.002. On the other hand, Decision Tree had a test score of 0.832 on the cross-validation with a standard deviation of +/- 0.003. Therefore, the Logistic Regression model performed better.
#
# - What are some of the ways we can reduce bias?
#
# To reduce bias in our model we used the one-hot encoding method instead of the label encoding. Label encoding gives the categories an integer value, so if there are 4 categories it assigns a number 1, 2, 3, or 4 to the categories, however this can create bias because the machine recognizes 4 as greater than 1 which not the case in categorical data. The onehot encoder assigns a different identifier using 0s and 1s, for each category in a feature and is not hierarchial.
#
# - How do we test our model?
#
# To test our model, we divided our data set into training and test data. We used used the training dataset to train our model, then we used the test dataset to test our model.
# ## Limitations
#
# This project only tested two classifier models.
# There are other classifier models that could probably have had a better score.
# ## References
# - <NAME>, et.al, An Introduction to Statistical Learning with Application in R, Springer, 8th edition 2017
#
# - [Kaggle](https://www.kaggle.com/samsonqian/titanic-guide-with-sklearn-and-eda)
#
# - [pydata](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.drop.html)
#
# - [PyData](https://www.youtube.com/watch?v=ioXKxulmwVQ)
#
# - [sklearn](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html?highlight=onehot%20encoding)
#
# - <NAME> and <NAME>, 1996. [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Census+Income)
# Data Mining and Visualization
# Silicon Graphics.
#
# - [adult data](https://www.youtube.com/watch?v=RdggP4yuIHY)
#
# - [Confusion matrix](https://www.youtube.com/watch?v=87Zebzxzh-A)
#
# - [Stackoverflow](https://stackoverflow.com/questions/31797013/how-to-open-a-data-file-extension)
|
Midterm Technical Report.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from argparse import Namespace
import os
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
from torch.optim.lr_scheduler import StepLR
print('GPU :', torch.cuda.is_available())
print('CUDA:', torch.version.cuda)
# +
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 32, 3, 1)
self.conv2 = nn.Conv2d(32, 64, 3, 1)
self.dropout1 = nn.Dropout2d(0.25)
self.dropout2 = nn.Dropout2d(0.5)
self.fc1 = nn.Linear(9216, 128)
self.fc2 = nn.Linear(128, 10)
def forward(self, x):
x = self.conv1(x)
x = F.relu(x)
x = self.conv2(x)
x = F.max_pool2d(x, 2)
x = self.dropout1(x)
x = torch.flatten(x, 1)
x = self.fc1(x)
x = F.relu(x)
x = self.dropout2(x)
x = self.fc2(x)
output = F.log_softmax(x, dim=1)
return output
def train(args, model, device, train_loader, optimizer, epoch):
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
output = model(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
if batch_idx % args.log_interval == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
def test(args, model, device, test_loader):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
test_loss += F.nll_loss(output, target, reduction='sum').item() # sum up batch loss
pred = output.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
# +
args = Namespace()
args.batch_size = 512
args.test_batch_size = 1000
args.epochs = 3
args.lr = 0.01
args.gamma = 0.7
args.seed = 31337
args.log_interval = 50
args.cuda = True
use_cuda = args.cuda and torch.cuda.is_available()
if use_cuda:
print('using CUDA')
else:
print('using CPU')
torch.manual_seed(args.seed)
device = torch.device("cuda" if use_cuda else "cpu")
kwargs = {'num_workers': 1, 'pin_memory': True} if use_cuda else {}
train_loader = torch.utils.data.DataLoader(
datasets.MNIST(os.path.join(os.environ['HOME'], 'workspace/ml-data'),
train=True,
download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=args.batch_size, shuffle=True, **kwargs)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST(os.path.join(os.environ['HOME'], 'workspace/ml-data'),
train=False,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=args.test_batch_size, shuffle=True, **kwargs)
model = Net().to(device)
optimizer = optim.Adam(model.parameters(), lr=args.lr)
scheduler = StepLR(optimizer, step_size=1, gamma=args.gamma)
for epoch in range(1, args.epochs + 1):
train(args, model, device, train_loader, optimizer, epoch)
test(args, model, device, test_loader)
scheduler.step()
# torch.save(model.state_dict(), "mnist_cnn.pt")
|
src/mnist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="SimCWmcWi0s5"
record = { 11602259 : {"name": "<NAME>" , "Pno": 1234, "cgpa" : 7.5},
11602260 : {"name": "<NAME>", "Pno": 5678, "cgpa" : 8},
11602261 : {"name": "<NAME>" , "Pno": 8984, "cgpa" : 7.8}}
# + colab={"base_uri": "https://localhost:8080/"} id="s2VuzKe6Vz0_" outputId="59f55095-b4ce-438d-9352-2e7129da64a2"
reg = 11602260
print(record[reg]['name'])
print(record[reg]['Pno'])
print(record[reg]['cgpa'])
# + id="oj3OmNRYkfEQ"
record[11602260]['Pno'] = 999
# + colab={"base_uri": "https://localhost:8080/"} id="_fzlbrWvkfRG" outputId="049917ea-df45-40d1-a2d9-a846ee2832d1"
record
# + id="DQbyOIOOYD-Z"
record = { 11602259 : {"name": "<NAME>" , "Pno": 1234, "cgpa" : 7.5},
11602260 : {"name": "<NAME>", "Pno": 5678, "cgpa" : 8.2},
11602261 : {"name": "<NAME>" , "Pno": 8984, "cgpa" : 7.8}}
# + id="GnZedQXvZncV"
import json
# + id="B50KFXNhZuCH"
js = json.dumps(record)
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="ip8ZcEIRZzEq" outputId="d78c29c4-f584-4d28-a952-46bdbfa2fe72"
js
# + colab={"base_uri": "https://localhost:8080/"} id="4gf1WCRLaXfa" outputId="59000a2d-9061-4960-94cb-9a558c1bb5df"
record
# + colab={"base_uri": "https://localhost:8080/"} id="3lSt3ACGaYQ_" outputId="b0f80a12-41d7-46d0-b7b7-a575139f7f62"
type(js)
# + colab={"base_uri": "https://localhost:8080/"} id="4s1t69dtacak" outputId="27bdbdd2-80da-4cc7-ef24-6e73470c626e"
type(record)
# + id="1zShzE_uad9A"
fd = open("record.txt",'w')
fd.write(js)
fd.close()
# + id="XPO-SFEMa6e9"
fd = open("record.txt",'r')
txt = fd.read()
fd.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="beJ863Wwo9RX" outputId="22f75014-7d4c-46de-ed8f-1f57285e1e5f"
txt
# + id="iHg0qnn7bBWQ"
record = json.loads(txt)
# + colab={"base_uri": "https://localhost:8080/"} id="T1wd9FbqbDWv" outputId="b693f4b1-6697-4098-c305-56dceb85c32c"
record
# + colab={"base_uri": "https://localhost:8080/"} id="LYPqrTgtbPWL" outputId="cb001b72-9dd4-4c1f-fe17-d17ee3324ef7"
record['11602259']
# + id="vhH1wMYDbU16"
import time
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="Rmr2z1AKcAum" outputId="6852095a-c20a-4870-c22e-5e38a57a7a7f"
time.ctime()
# + colab={"base_uri": "https://localhost:8080/"} id="2oxCQoa-cCZI" outputId="fd1709a1-27cb-44ca-9ab8-d6d4e5611d77"
record.values()
|
Working with JSON Assignment/JSON_based_UMS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import load_boston
x = np.linspace(-3,5,20)
y = 2 * x + 3
y_noise = np.random.normal(0,2,20)
y += y_noise
plt.scatter(x,y)
plt.show()
for y_guess in [3*x + 8,4 * x + 3, -2 * x]:
plt.scatter(x,y_noise)
plt.plot(x,y_guess)
plt.show()
def calculate_loss_function(x_input,y_input,a,b):
assert len(x_input) == len(y_input)
y_predicted = a*x + b
d = np.power(y - y_predicted,2)
return np.sum(d)/len(x)
calculate_loss_function(x,y,3,8)
calculate_loss_function(x,y,2,3)
def compute_gradients(x, y, a, b):
a_gradient = -2 / len(x) * np.sum(x * (y - (a * x + b)))
b_gradient = -2 / len(y) * np.sum(y - (a * x + b))
return (a_gradient, b_gradient)
np.array(compute_gradients(x,y,-2,0)) * 0.001
a,b = -10,20
def perform_gradient_descent(x, y, a, b, learning_rate):
a_gradient = -2 / len(x) * np.sum(x * (y - (a * x + b)))
b_gradient = -2 / len(y) * np.sum(y - (a * x + b))
new_a = a - a_gradient * learning_rate
new_b = b - b_gradient * learning_rate
return (new_a, new_b)
alpha = 0.01 # Learning rate
for step in range(3041):
a, b = perform_gradient_descent(x, y, a, b, alpha)
if step % 100 == 0:
error = calculate_loss_function(x, y, a, b)
print("Step {}: a = {}, b = {}, J = {}".format(step, a, b, error))
print("Final line: {} * x + {}".format(a, b))
print(a,b)
y_guessed = a * x + b
plt.scatter(x,y)
plt.plot(x,y_guessed)
plt.show()
boston_data = load_boston()
print(boston_data.DESCR)
|
06-Regression-Models/practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import util.preprocessing as preprocessing
import util.detection_util as detection_util
import sys
sys.path.append('../')
import pandas as pd
import numpy as np
import cv2
import os
import xlrd
import openpyxl
import matplotlib.pyplot as plt
import util.config as config
from matplotlib import pyplot as plt
np.warnings.filterwarnings('ignore', category=np.VisibleDeprecationWarning)
plt.rcParams['figure.figsize'] = [12, 6]
# -
df = pd.read_excel(r'D:\Users\avatar\PycharmProjects\pig-face-recognition\sample\detection-metrics.xlsx')
print(df.head().to_string())
df = df.astype({'image_name': 'str', 'iou': 'float', 'mAP': 'int'})
df.info()
# +
threshold = 0.75
ax = df.plot(x='image_name', y=[1], kind='bar',color='#607c8e', figsize=(9,6))
ax.plot([0.,11], [threshold, threshold], "k--")
# df['sharpness'].hist(bins='auto',color='#607c8e',alpha=0.7, rwidth=0.85)
plt.title('Intersection over Union')
# plt.xlabel('ImageId')
plt.ylabel('IoU')
plt.show()
# -
|
util/detection-metrics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
df = pd.read_csv("datasets/titanic_data.csv")
# #### Cross Validation
from sklearn.model_selection import ShuffleSplit
rs = ShuffleSplit(n_splits=5, test_size=.25, random_state=0)
for train_index, test_index in rs.split(df):
print(len(train_index), len(test_index))
train_fold = df.iloc[train_index]
test_fold = df.iloc[test_index]
from sklearn.model_selection import StratifiedKFold
from sklearn.base import clone
# #### Train-Test Split
# * Remove a subset for testing
# * Cross-validate on training set
# +
from sklearn.model_selection import train_test_split
X = df.drop("Survived", axis=1)
y = df[["Survived"]]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# -
print(X_train.shape)
print(X_test.shape)
# +
skfolds = StratifiedKFold(n_splits=5, random_state=42)
for train_index, val_index in skfolds.split(X_train, y_train):
print(len(train_index), len(test_index))
train_fold = df.iloc[train_index]
val_fold = df.iloc[val_index]
# +
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import SGDClassifier
# sgd_clf = SGDClassifier(random_state=42)
# cross_val_score(sgd_clf, X_train, y_train, cv=3, scoring="accuracy")
|
training/cross_validation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.1 64-bit (''.venv'': venv)'
# language: python
# name: python3
# ---
from azureml.core import Workspace
ws = Workspace.from_config()
# +
import urllib.request
from azureml.core.model import Model
# Download model
from pathlib import Path
my_file = Path("./model.onnx")
if not my_file.exists():
urllib.request.urlretrieve("https://aka.ms/bidaf-9-model", "./model.onnx")
# +
# Register model
model = Model.register(ws, model_name="bidaf_onnx", model_path="./model.onnx")
# +
from azureml.core import Environment
from azureml.core.model import InferenceConfig
env = Environment(name="AzureML-onnxruntime-1.6-ubuntu18.04-py37-cpu-inference")
dummy_inference_config = InferenceConfig(
environment=env,
source_directory="./source_dir",
entry_script="./echo_score.py",
)
# +
from azureml.core.webservice import LocalWebservice
deployment_config = LocalWebservice.deploy_configuration(port=6789)
# -
service = Model.deploy(
ws,
"myservice",
[model],
dummy_inference_config,
deployment_config,
overwrite=True,
)
service.wait_for_deployment(show_output=True)
print(service.get_logs())
|
module3/after/model_deploy_locally/deploy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import urllib.request, json , time, os, difflib, itertools
import pandas as pd
from multiprocessing.dummy import Pool
from datetime import datetime
# +
try:
import httplib
except:
import http.client as httplib
def check_internet():
conn = httplib.HTTPConnection("www.google.com", timeout=5)
try:
conn.request("HEAD", "/")
conn.close()
# print("True")
return True
except:
conn.close()
# print("False")
return False
# -
check_internet()
def get_historic_price(query_url,json_path,csv_path):
stock_id=query_url.split("&period")[0].split("symbol=")[1]
#стираем файлы если уже есть
if os.path.exists(csv_path+stock_id+'.csv') and os.stat(csv_path+stock_id+'.csv').st_size != 0:
os.remove(csv_path+stock_id+'.csv')
while not check_internet():
print("Could not connect, trying again in 5 seconds...")
time.sleep(5)
try:
with urllib.request.urlopen(query_url) as url:
parsed = json.loads(url.read().decode())
except:
print("||| Historical data of "+stock_id+" doesn't exist")
return
else:
if os.path.exists(json_path+stock_id+'.json') and os.stat(json_path+stock_id+'.json').st_size != 0:
os.remove(json_path+stock_id+'.json')
with open(json_path+stock_id+'.json', 'w') as outfile:
json.dump(parsed, outfile, indent=4)
try:
Date=[]
for i in parsed['chart']['result'][0]['timestamp']:
Date.append(datetime.utcfromtimestamp(int(i)).strftime('%Y-%m-%d'))
#Low=parsed['chart']['result'][0]['indicators']['quote'][0]['low']
#Open=parsed['chart']['result'][0]['indicators']['quote'][0]['open']
#Volume=parsed['chart']['result'][0]['indicators']['quote'][0]['volume']
#High=parsed['chart']['result'][0]['indicators']['quote'][0]['high']
Close=parsed['chart']['result'][0]['indicators']['quote'][0]['close']
#Adjusted_Close=parsed['chart']['result'][0]['indicators']['adjclose'][0]['adjclose']
df=pd.DataFrame(list(zip(Date,Close)),columns =['Date',stock_id[1:]])
if os.path.exists(csv_path+stock_id+'.csv'):
os.remove(csv_path+stock_id+'.csv')
df.to_csv(csv_path+stock_id+'.csv', sep=',', index=None)
print(">>> Historical data of "+stock_id+" saved")
except:
print(">>> Historical data of "+stock_id+" could not be saved")
return
json_path = "data/json/"
csv_path = "data/csv/"
if not os.path.isdir(json_path):
os.makedirs(json_path)
if not os.path.isdir(csv_path):
os.makedirs(csv_path)
from tickers import tickers_dict
# Период с 2016-06-30 по 2021-06-30, интервал 1 неделя (допустмые значения [1m, 2m, 5m, 15m, 30m, 60m, 90m, 1h, 1d, 5d, 1wk, 1mo, 3mo])
timestamp_start = str(int(datetime.strptime('2018-06-30', "%Y-%m-%d").timestamp()))
timestamp_end = str(int(datetime.strptime('2019-06-30', "%Y-%m-%d").timestamp()))
interval = '1d'
timestamp_start
# +
query_urls=[]
for ticker in tickers_dict.values():
query_urls.append("https://query1.finance.yahoo.com/v8/finance/chart/"+ticker[0]+"?symbol="+ticker[0]+"&period1="+timestamp_start+"&period2="+timestamp_end+"&interval="+interval)
# -
query_urls
# %%time
with Pool(processes=10) as pool:
pool.starmap(get_historic_price, zip(query_urls, itertools.repeat(json_path), itertools.repeat(csv_path)))
|
data_scraping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# ### Observations:
# * Cities around the equator have the highest temperature
# * There is no strong relationship between latitude and humidity
# * Most cities have wind speed of less than 15 miles per hour.
#
#
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
import openweathermapy.core as ow
# Import API key
from api_keys import api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(low=-90.000, high=90.000, size=1500)
lngs = np.random.uniform(low=-180.000, high=180.000, size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
# Create settings dictionary with information we're interested in
settings = {"units": "imperial", "appid": api_key}
# Get current weather
weather_data = []
city_number=1
print("Beginning Data Retrieval")
print("---------------------------------------")
for city in cities:
try:
weather_data.append(ow.get_current(city, **settings))
print(f"Processing Record {city_number} | {city}")
city_number+=1
except:
print("City not found. Skipping...")
# Timer
time.sleep(1.2)
print("---------------------------------------")
print("Retrieval Complete")
#weather_data
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
# +
# Create an "extracts" object to get the temperature, latitude,longitude, cloudiness, wind speed,
#humidity,data and country for each city
summary = ["name","clouds.all","sys.country","dt","main.humidity","coord.lat", "coord.lon","main.temp_max","wind.speed"]
column_names = ["City","Cloudiness","Country","Date ","Humidity", "Lat","Lng", "Max Temp", "Wind Speed"]
# Create a Pandas DataFrame with the results
data = [response(*summary) for response in weather_data]
weather_data_df = pd.DataFrame(data,columns=column_names)
city_weather_df=weather_data_df.set_index("City")
city_weather_count=city_weather_df.count()
print(city_weather_count)
# -
city_weather_df.head()
# +
#output to csv
city_weather_df.to_csv(output_data_file)
# -
# ### Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# #### Latitude vs. Temperature Plot
# +
plt.scatter(city_weather_df["Lat"], city_weather_df["Max Temp"], marker="o",
facecolors="teal", edgecolors="black")
# Incorporate the other graph properties
plt.title("City Latitude vs. Max Temperature (8/23/2019)")
plt.ylabel("Max Temperature (F)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("CityLatvsMaxTemp.png",bbox_inches="tight", dpi = 300)
# Show plot
plt.show()
# -
# #### Latitude vs. Humidity Plot
# +
plt.scatter(city_weather_df["Lat"],city_weather_df["Humidity"], marker="o",
facecolors="teal", edgecolors="black")
# Incorporate the other graph properties
plt.title("City Latitude vs. Humidity (8/23/2019)")
plt.ylabel("Humidity (%)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("CityLatvsHumidity.png",bbox_inches="tight", dpi = 300)
# Show plot
plt.show()
# -
# #### Latitude vs. Cloudiness Plot
# +
plt.scatter(city_weather_df["Lat"],city_weather_df["Cloudiness"], marker="o",
facecolors="teal", edgecolors="black")
# Incorporate the other graph properties
plt.title("City Latitude vs. Cloudiness (8/23/2019)")
plt.ylabel("Cloudiness (%)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("CityLatvsCloudiness.png",bbox_inches="tight", dpi = 300)
# Show plot
plt.show()
# -
# #### Latitude vs. Wind Speed Plot
# +
plt.scatter(city_weather_df["Lat"],city_weather_df["Wind Speed"], marker="o",
facecolors="teal", edgecolors="black")
# Incorporate the other graph properties
plt.title("City Latitude vs. Wind Speed (8/23/2019)")
plt.ylabel("Wind Speed (mph)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("CityLatvsWindSpeed.png",bbox_inches="tight", dpi = 300)
# Show plot
plt.show()
# -
|
resources/WeatherPy_Diana_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DoWhy: Different estimation methods for causal inference
# This is a quick introduction to the DoWhy causal inference library.
# We will load in a sample dataset and use different methods for estimating the causal effect of a (pre-specified)treatment variable on a (pre-specified) outcome variable.
#
# First, let us add the required path for Python to find the DoWhy code and load all required packages
import os, sys
sys.path.append(os.path.abspath("../../"))
# +
import numpy as np
import pandas as pd
import logging
import dowhy
from dowhy.do_why import CausalModel
import dowhy.datasets
# -
# Now, let us load a dataset. For simplicity, we simulate a dataset with linear relationships between common causes and treatment, and common causes and outcome.
#
# Beta is the true causal effect.
data = dowhy.datasets.linear_dataset(beta=10,
num_common_causes=5,
num_instruments = 2,
num_samples=10000,
treatment_is_binary=True)
df = data["df"]
# Note that we are using a pandas dataframe to load the data.
# ## Identifying the causal estimand
# We now input a causal graph in the DOT graph format.
# With graph
model=CausalModel(
data = df,
treatment=data["treatment_name"],
outcome=data["outcome_name"],
graph=data["gml_graph"],
instruments=data["instrument_names"],
logging_level = logging.INFO
)
model.view_model()
from IPython.display import Image, display
display(Image(filename="causal_model.png"))
# We get a causal graph. Now identification and estimation is done.
identified_estimand = model.identify_effect()
print(identified_estimand)
# ## Method 1: Regression
#
# Use linear regression.
causal_estimate_reg = model.estimate_effect(identified_estimand,
method_name="backdoor.linear_regression",
test_significance=True)
print(causal_estimate_reg)
print("Causal Estimate is " + str(causal_estimate_reg.value))
# ## Method 2: Stratification
#
# We will be using propensity scores to stratify units in the data.
causal_estimate_strat = model.estimate_effect(identified_estimand,
method_name="backdoor.propensity_score_stratification")
print(causal_estimate_strat)
print("Causal Estimate is " + str(causal_estimate_strat.value))
# ## Method 3: Matching
#
# We will be using propensity scores to match units in the data.
causal_estimate_match = model.estimate_effect(identified_estimand,
method_name="backdoor.propensity_score_matching")
print(causal_estimate_match)
print("Causal Estimate is " + str(causal_estimate_match.value))
# ## Method 4: Weighting
#
# We will be using (inverse) propensity scores to assign weights to units in the data.
causal_estimate_ipw = model.estimate_effect(identified_estimand,
method_name="backdoor.propensity_score_weighting")
print(causal_estimate_ipw)
print("Causal Estimate is " + str(causal_estimate_ipw.value))
# ## Method 5: Instrumental Variable
#
# We will be using the Wald estimator for the provided instrumental variable.
causal_estimate_iv = model.estimate_effect(identified_estimand,
method_name="iv.instrumental_variable", method_params={'iv_instrument_name':'Z1'})
print(causal_estimate_iv)
print("Causal Estimate is " + str(causal_estimate_iv.value))
# ## Method 6: Regression Discontinuity
#
# We will be internally converting this to an equivalent instrumental variables problem.
causal_estimate_regdist = model.estimate_effect(identified_estimand,
method_name="iv.regression_discontinuity",
method_params={'rd_variable_name':'Z1',
'rd_threshold_value':0.5,
'rd_bandwidth': 0.1})
print(causal_estimate_regdist)
print("Causal Estimate is " + str(causal_estimate_regdist.value))
|
docs/source/dowhy_estimation_methods.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Insert HTML in a NoteBook
# > How to write HTML inside a Notebook for fastpages
#
# - toc: false
# - badges: false
# - comments: true
# - author: <NAME>
# - categories: [HTML, fastpages]
# The fastpages blog engine is very useful to write a notebook. But I wanted to be able to write HTML directly in my notebook to implement a template.
#
# For the moment, I'm not going to do like that, I'll keep my template in a classic notebook, but I'll leave you here the technique to write HTML from a notebook which will be interpreted by fastpages in HTML.
# + active=""
# <h3>This is HTML written in the notebook in a plain text cell (not Markdown), with h3 tags</h3>
# -
#
# If I try to write html with print in a code cell, it fails.
age = 45
name = "Paul"
gap = 10
print(f"<h3>In {gap} year(s), {name} will be {age + gap} year(s) old.</h3>")
#
# With this formula, a new plain text cell is created, so that's good!
get_ipython().run_cell_magic(u'HTML', u'',f"<h3>In {gap} year(s), {name} will be {age + gap} year(s) old.</h3>")
|
_notebooks/2021-04-18-TestHTML.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Instacart dataset EDA 프로젝트
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="instacartl2.png" alt="Drawing" style="width: 1000px"/>
# <img src="instacart1.jpg" alt="Drawing" style="width: 500px"/>
#
# + slideshow={"slide_type": "skip"}
from IPython.display import Image
# + [markdown] slideshow={"slide_type": "slide"}
# # 회사 개요
# - **"식품계의 우버"라 불리는 미국의 신선식품 배달 서비스 스타트업. (ex : 한국의 B마트)**
# - **월마트, 세이프웨이, 코스트코등 대형마트 부터 지역 슈퍼마켓**
# - **설립 2년 만에 유니콘**
# - **8년간 16억달러(약 1조8800억원) 투자유치**
# + [markdown] slideshow={"slide_type": "slide"}
# # Dataset 개요
# > - **총 유저수 : 206,209(약 20만 명)**
# > - **판매된 총 제품수 : 33,819,106(약 3380만 건)**
# > - **품목 수 : 49,688(약 5만 종)**
# > - **총 주문량 : 3,421,083(약 340만 건)**
# > - **품목당 평균 판매 횟수 : 약 680건**
# > - **유저 1명당 평균 주문 횟수 : 16.5회**
# > - **재구매율 : 약 59%**
# + [markdown] slideshow={"slide_type": "slide"}
# ## 선택한 이유 :
# > - **데이터의 양**
# > - **실제 생활과 밀접**
# > - **예측 모델**
# + [markdown] slideshow={"slide_type": "slide"}
# ## 데이터 로드
# + slideshow={"slide_type": "-"}
df_orders = pd.read_csv("origin/orders.csv")
df_order_products = pd.read_csv("origin/merged_order_products.csv")
df_ptoducts = pd.read_csv("origin/products.csv")
df_aisles = pd.read_csv("origin/aisles.csv")
df_departments = pd.read_csv("origin/departments.csv")
product_plus_aisle = pd.merge(df_ptoducts, df_aisles)
product_plus_aisle_plus_departments = pd.merge(
product_plus_aisle, df_departments)
product_plus_aisle_plus_departments_plus_order_products = pd.merge(
product_plus_aisle_plus_departments, df_order_products)
raw_data = pd.merge(
product_plus_aisle_plus_departments_plus_order_products, df_orders)
# + [markdown] slideshow={"slide_type": "slide"}
# ## raw_data 컬럼
#
# ### 시간관련
# - order_dow: 주문한 요일
# - order_hour_of_day: 주문한 시간
#
#
# ### 제품관련
# - product_id: 제품 ID
# - product_name: 제품 이름
# - aisle_id: 소분류 ID
# - aisle: 소분류 이름
# - department_id: 대분류 ID
# - department: 대분류 이름
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### 주문관련
# - user_id: 고객 ID
# - order_id: 주문 ID
# - add_to_cart_order: 각 제품이 장바구니에 추가 된 순서
# - reordered: 사용자가 해당 제품을 과거에 주문한 경우 1, 그렇지 않으면 0
#
# ### etc
# - days_since_prior: 마지막 주문 이후 일 수, 30 일 제한 (NA는 order_number1) 재주문 기간의 텀
# - aisle_id : foreign key
# - department_id: foreign key
# - eval_set: 이 순서가 속하는 평가 세트
# - order_number: 이 사용자의 주문 순서 번호 (1 = 첫 번째, n = n 번째)
# + [markdown] slideshow={"slide_type": "slide"}
# # 그래서 뭐가 궁금한가?
# + [markdown] slideshow={"slide_type": "slide"}
# # 이용자들은 어떤 물건을 재구매할 확률이 높은 걸까?
# + [markdown] slideshow={"slide_type": "slide"}
# ## 가설 1 : 시간 및 요일등에 따라 품목과 재구매 정도의 차이가 있을것이다.
# ## 가설 2 : 판매량이 높은 제품이 재구매도 많을 것이다.
# ## 가설 3 : 장바구니의 첫번째 담은 물건의 재구매의 확률이 높을 것이다.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 시간 및 요일별 구매 및 재구매 횟수
# + code_folding=[] slideshow={"slide_type": "slide"}
# 시간,요일 별 구매 횟수
df = pd.pivot_table(raw_data, index=raw_data['order_dow'],
columns=raw_data['order_hour_of_day'], aggfunc='count', values='user_id')
df = df / 1000
df = df.astype("int")
# 시간,주 별 재구매 횟수
columns = [ 'reordered', 'order_dow', 'order_hour_of_day']
data_time = raw_data[columns]
time_table = data_time.pivot_table(
index='order_dow', columns='order_hour_of_day',
values='reordered', aggfunc=sum)
time_table = time_table / 1000
time_table = time_table.astype("int")
# + code_folding=[11] slideshow={"slide_type": "slide"}
plt.figure(figsize=(15, 10))
plt.subplot(121)
ax = plt.axes()
ax.set_title('시간 요일별 주문 횟수 (횟수/1000)')
sns.heatmap(df, cmap="BrBG", center=3, annot=True,
fmt="d", square=True, cbar=False, ax=ax)
plt.tight_layout()
plt.figure(figsize=(15, 10))
plt.subplot(122)
ax = plt.axes()
ax.set_title('시간 요일별 재주문 횟수 (횟수/1000)')
sns.heatmap(time_table, cmap="BrBG", center=3, annot=True,
fmt="d", square=True, cbar=False, ax=ax)
plt.tight_layout()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# - ### 09시 ~ 17시 사이에 주문이 집중
# - ### 토,일요일에 주문이 집중(0,1)
# - ### 시간, 요일에 따른 구매횟수와 재주문 횟수는 비슷한 빈도를 보임
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### 구매가 가장 많이 일어나는 시간대에 무슨 품목이 많이 팔리는가?
# + slideshow={"slide_type": "slide"}
# 요일 별 구매 상품 순위
daily_product = pd.pivot_table(
raw_data, index=raw_data['product_name'], columns=raw_data['order_dow'], aggfunc='count', values='user_id')
result = []
for i in range(0, 2):
a = daily_product[i].sort_values(ascending=False)[:10].index
result.append(a)
daily_product = pd.DataFrame(result)
daily_product['order_dow'] = [0, 1]
daily_product.set_index('order_dow')
# + slideshow={"slide_type": "slide"}
# 시간대 별 구매 상품 순위
df = pd.pivot_table(raw_data, index=raw_data['product_name'],
columns=raw_data['order_hour_of_day'], aggfunc='count', values='user_id')
result = []
for i in range(9, 17):
a = df[i].sort_values(ascending=False)[:10].index
result.append(a)
hourly_product = pd.DataFrame(result)
hourly_product['hour'] = [9, 10, 11, 12, 13, 14, 15, 16]
hourly_product.set_index('hour')
hourly_product
# + [markdown] slideshow={"slide_type": "slide"}
# # 가설 1의 결론:
#
# - ### 토,일요일 낮시간대 (09~17시) 에 바나나(유기농), 딸기, 시금치, 아보카도 등이 많이 팔린다
# - ### 요일, 시간별 구매횟수와 재구매 횟수는 비슷한 분포를 보인다.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 가설 2 : 판매량이 높은 제품이 재구매도 많을 것이다.
# + [markdown] slideshow={"slide_type": "slide"}
# ## product
# + slideshow={"slide_type": "-"}
# 분석용 데이터 생성
columns = ['product_id', 'reordered', 'product_name']
data_product = raw_data[columns]
# 피봇테이블
product_table = data_product.pivot_table(
index='product_name', columns='reordered', aggfunc='count', fill_value=0)
columns = ['first_order', 'reorder']
product_table.columns = columns
# 테이블 정리
product_table['product_total'] = product_table['first_order'] + \
product_table['reorder']
product_table['product_reorder_rate'] = product_table['reorder'] / \
product_table['product_total'] * 100
product_table.sort_values(by='product_total', ascending=False, inplace=True)
product_table.head()
# + code_folding=[1] slideshow={"slide_type": "slide"}
# 몇개 이상의 제품이 팔려야 많이 팔린것인가?
def numoforder(n):
return product_table[product_table['product_total'] >= n]
n = 1000
print("{}개 이상 팔린 제품의 총 개수:".format(n),numoforder(n)['product_total'].sum())
print("총 팔린 제품의 개수:",product_table['product_total'].sum())
print(f'{n}개 이상 팔리면 전체 품목에서',round(numoforder(n)['product_total'].sum() / product_table['product_total'].sum() * 100,2),'%를 차지합니다.' )
# + slideshow={"slide_type": "-"}
product_table = product_table[product_table['product_total'] >= 1000]
product_table.sort_values(by='product_name', ascending=True, inplace=True)
# + [markdown] slideshow={"slide_type": "slide"}
# ## 두 데이터의 단위의 차이가 크기 때문에 스피어만 상관계수를 이용
# + slideshow={"slide_type": "-"}
# 스피어만 상관계수
print("스피어만 상관계수", sp.stats.spearmanr(
product_table['product_total'], product_table['product_reorder_rate'])[0])
# 피어슨 상관계수
print("피어슨 상관계수", np.corrcoef(
product_table['product_total'], product_table['product_reorder_rate'])[0, 1])
# + code_folding=[] slideshow={"slide_type": "slide"}
# 산점도
from sklearn import preprocessing
plt.plot(np.log(product_table['product_reorder_rate']),
np.log(preprocessing.minmax_scale(product_table['product_total'])),
linestyle='none',
marker='o',
markersize=3,
color='blue',
alpha=0.5)
plt.title("재구매 비율의 산점도")
plt.xticks()
plt.yticks()
plt.tight_layout()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Aisle
# + code_folding=[] slideshow={"slide_type": "-"}
# 분석용 데이터 프레임 생성
columns = ['aisle_id', 'reordered', 'aisle']
data_aisle = raw_data[columns]
# 피봇테이블 생성
aisle_table = data_aisle.pivot_table(
index='aisle', columns='reordered', aggfunc='count')
columns = ['first_order', 'reorder']
aisle_table.columns = columns
# 피봇테이블 정리
aisle_table['aisle_total'] = aisle_table['first_order'] + \
aisle_table['reorder']
aisle_table['aisle_reorder_rate'] = aisle_table['reorder'] / \
aisle_table['aisle_total'] * 100
aisle_table.sort_values(by='aisle', ascending=False, inplace=True)
aisle_table.head()
# + code_folding=[] slideshow={"slide_type": "slide"}
# 스피어만 상관계수
print("스피어만 상관계수", sp.stats.spearmanr(
aisle_table['aisle_total'], aisle_table['aisle_reorder_rate'])[0])
# 피어슨 상관계수
print("피어슨 상관계수", np.corrcoef(
aisle_table['aisle_total'], aisle_table['aisle_reorder_rate'])[0, 1])
# + code_folding=[0] slideshow={"slide_type": "slide"}
# 산점도
plt.subplot(121)
plt.plot(np.log(aisle_table['aisle_reorder_rate']),
np.log(aisle_table['aisle_total']),
linestyle='none',
marker='o',
markersize=3,
color='blue',
alpha=0.5)
plt.title("재구매 비율의 산점도(aisel별)")
plt.tight_layout()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## 가설 2 결론 :
# - ### 양의 상관계를 보이는것으로 보아, 주문량이 증가하면 재구매율도 같이 높다.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 가설 3 : 장바구니의 첫번째 담은 물건의 재구매의 확률이 높을 것이다.
# + slideshow={"slide_type": "slide"}
# 분석용 데이터셋 생성
columns = ['order_id', 'product_id', 'add_to_cart_order',
'reordered', 'days_since_prior_order']
data_cart = raw_data[columns]
data_cart.head(2)
# + code_folding=[] slideshow={"slide_type": "-"}
# 총 주문갯수가 1개인 데이터의 갯수
a = data_cart['order_id'].value_counts() == 1
print(a.sum(), '개')
# + slideshow={"slide_type": "-"}
# 전체 주문이 1인 데이터 갯수 전처리
# 전체 주문이 1인 데이터를 데이터 프레임 및 True = 1, False = 0 으로 변환
a = pd.DataFrame(a)
a.replace(False, 0, inplace=True)
# 컬럼명 변환
a.columns = ['first']
# merge용 index 생성
a.reset_index(inplace=True)
# + slideshow={"slide_type": "-"}
# 분석용 데이터 셋과, 전체 주문이 1개인 데이터 merge
data_cart_two_more = data_cart.merge(a, left_on='order_id', right_on='index')
# + slideshow={"slide_type": "slide"}
# 전체 주문이 1개인 데이터 삭제
data_cart_two_more[data_cart_two_more['first'] != 1]
# + slideshow={"slide_type": "slide"}
# 분석에 필요한 컬럼만 재 정렬
data_cart_two_more = data_cart_two_more[['order_id', 'product_id',
'add_to_cart_order', 'reordered', 'days_since_prior_order']]
# + slideshow={"slide_type": "-"}
# 장바구니에 1번째로 담긴 품목만 확인
data_cart_two_more = data_cart_two_more[data_cart_two_more['add_to_cart_order'] == 1]
data_cart_two_more
# + [markdown] slideshow={"slide_type": "-"}
# ## 각 주문에서 장바구니에 제일 처음 담은 물품만 남기고 필터링
# ## 3,346,083 (약 330만)
# + slideshow={"slide_type": "skip"}
# 피벗테이블 생성
table_cart = data_cart_two_more.pivot_table(
index='product_id', columns='reordered', values='add_to_cart_order', aggfunc='count', fill_value=0)
table_cart.reset_index(inplace=True)
table_cart.columns = ['product_id', 'first_order', 'reorder']
# proudct_name을 알기위해 df_ptoducts merge
table_cart_product = table_cart.merge(
df_ptoducts, left_on='product_id', right_on='product_id')
# + slideshow={"slide_type": "skip"}
# 피벗테이블의 컬럼명 정리
table_cart_product = table_cart_product[[
'product_id', 'product_name', 'first_order', 'reorder']]
# 첫 주문과, 재주문의 총합 (1번째로 장바구니에 추가된 제품들의)
table_cart_product['total'] = table_cart_product['first_order'] + \
table_cart_product['reorder']
# 1번째로 장바구니에 추가된 제품의 재구매 비율
table_cart_product['reorder_rate'] = table_cart_product['reorder'] / \
table_cart_product['total'] * 100
# + [markdown] slideshow={"slide_type": "slide"}
# ## 장바구니에 처음 담긴 물품의 재구매율 상위 5위 비교
# + slideshow={"slide_type": "-"}
# 주문량이 많지 않음
table_cart_product.sort_values(by='reorder_rate', ascending=False).head()
# + [markdown] slideshow={"slide_type": "slide"}
# ## 판매된 제품 수량별 재구매율
# + slideshow={"slide_type": "-"}
# 대표성 확인을 위한 함수 설정
def numofproduct(n):
return table_cart_product[table_cart_product['total'] >= n].sort_values(by='reorder_rate', ascending=False)
# + code_folding=[0] slideshow={"slide_type": "slide"}
# 100개 이상 팔린 제품의 reorder_rate
n = 100
print(f'{n}개 이상 팔린 제품은', numofproduct(n)['total'].count(), '가지 입니다.')
print(f'{n}개 이상 팔린 제품수의 총합은', numofproduct(n)['total'].sum(), '개 입니다.')
print('장바구니에 1번째로 선택된 제품의 총합은', table_cart_product['total'].sum(), '개 입니다.')
print(f'{n}개 이상 팔린 제품은 장바구니에 1번째로 선택된 제품 총합의', round(numofproduct(n)[
'total'].sum() / table_cart_product['total'].sum() * 100, 2), '% 입니다')
# + code_folding=[0] slideshow={"slide_type": "-"}
# 1000개 이상 팔린 제품의 reorder_rate
n = 1000
print(f'{n}개 이상 팔린 제품은', numofproduct(n)['total'].count(), '가지 입니다.')
print(f'{n}개 이상 팔린 제품은', numofproduct(n)['total'].sum(), '개 입니다.')
print('장바구니에 1번째로 선택된 제품의 총합은', table_cart_product['total'].sum(), '개 입니다.')
print(f'{n}개 이상 팔린 제품은 장바구니에 1번째로 선택된 제품 총합의', round(numofproduct(n)[
'total'].sum() / table_cart_product['total'].sum() * 100, 2), '% 입니다')
# + code_folding=[0] slideshow={"slide_type": "-"}
# 10000개 이상 팔린 제품의 reorder_rate
n = 10000
print(f'{n}개 이상 팔린 제품은', numofproduct(n)['total'].count(), '가지 입니다.')
print(f'{n}개 이상 팔린 제품은', numofproduct(n)['total'].sum(), '개 입니다.')
print('장바구니에 1번째로 선택된 제품의 총합은', table_cart_product['total'].sum(), '개 입니다.')
print(f'{n}개 이상 팔린 제품은 장바구니에 1번째로 선택된 제품 총합의', round(numofproduct(n)[
'total'].sum() / table_cart_product['total'].sum() * 100, 2), '% 입니다')
# + slideshow={"slide_type": "slide"}
numofproduct(100).head(3)
# + slideshow={"slide_type": "-"}
numofproduct(1000).head(3)
# + slideshow={"slide_type": "-"}
numofproduct(10000).head(3)
# + [markdown] slideshow={"slide_type": "slide"}
# ## 가설 3 결론 : 장바구니에 첫번째로 들어간 제품은 재구매 한 재품일 확률이 높다.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 가설 1 결론
# - ### 요일, 시간별 구매횟수와 재구매 횟수는 비슷한 분포를 보인다.
#
# ## 가설 2 결론
# - ### 양의 상관계를 보이는것으로 보아, 주문량이 증가하면 재구매율도 같이 높다.
#
# ## 가설 3 결론
# - ### 장바구니에 첫번째로 들어간 제품은 재구매 한 재품일 확률이 높다.
# + [markdown] slideshow={"slide_type": "slide"}
# # 그래서 이용자들은 어떤 물건을 재구매할 확률이 높은 걸까?
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### 재구매율은 시간 및 요일보다, 제품의 주문량이 높고, 장바구니에 첫번째로 들어간 제품일수록 높아진다.
# + [markdown] slideshow={"slide_type": "slide"}
# ## So what?
# > **사업이 발전하려면, 재구매율을 높여야 한다.**
#
# > **이를 위해, 주문량이 높고, 장바구니에 처음들어간 제품을 우선으로 프로모션하고,
# 유사한 제품을 취급하는 채널들과의 연계를 발전시켜야 한다.**
# + [markdown] slideshow={"slide_type": "slide"}
# # 그리고...
#
# - #### 처음 구매한 주문이 약 40%를 차지 한다는건, 취급하는 상점이 증가되거나, 신규유저가 지속적으로 유입되서 새로운 주문이 증가했다고 볼수 있다.
# - #### 즉 이 데이터셋이 수집된 기간 동안 사업이 지속적으로 성장했다고도 볼수 있다.
#
# + slideshow={"slide_type": "-"}
# 처음 구매한 비율
1 - (raw_data["reordered"].sum() / len(raw_data["reordered"]))
# + [markdown] slideshow={"slide_type": "slide"}
# # 아쉬운점
#
# - #### 예측 모델을 만들어 봤으면..
# - #### barplot 같은 좀더 직관적인 그래프 였으면..
# - #### 날짜 데이터가 있었다면..
|
EDAproject/EDA_project.ipynb
|
% -*- coding: utf-8 -*-
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Matlab
% language: matlab
% name: matlab
% ---
% EE4C03
% ===
% Statistical Digital Signal Processing and Modeling
% ============
%
% # Session N.01
%
% Note: Interact with Octave/Matlab in Notebook. All commands are interpreted by Octave/Matlab. Help on commands is available using the `%help` magic or using `?` with a command.
% ## 1. Projections
%
% If $\bf v$ is a vector, then a projection onto $\bf v$ is the matrix
%
% $$
% {\bf P}= \frac{1}{{\bf v}^H{\bf v}}{\bf v}{\bf v}^{H}
% $$
% ##### a. Generate a random $5 \times 1$ vector
% modify code here
v = randn(1,1)
% ##### b. Construct the Projection matrix $\bf P$
% modify code here
% hint: v^H = v'
% hint: notice that v'*v is a scalar
% hint: x./y does element-wise division
% advice: For Matlab/Octave for defining symmetric matrices is
% encouraged to use paranthesis, e.g., (x*x')
P = eye(1)
% +
% caution:!!
% P2 = v*inv(v'*v)*v'; might work! but...Matlab/Octave do...shady things
% extra: what could be the "numerical" problem that you might encounter later?
% -
% Check the properties of an (orthogonal) projection matrix, i.e.,
%
% $$
% {\bf P P = P}\\
% {\bf P}^H = \bf P
% $$
% +
% add code here
% [code]
% -
% Also checks that it leaves $\bf v$ intact: $\bf Pv = v$.
% +
% add code here
% [code]
% -
% ##### c. Do an eigenvalue decomposition of $\bf P$, i.e., ${\bf P = U\Lambda U}^H$
% +
% add code here
% hint: use 'help eig' to see the sintaxis
% [U,Lambda] = [code]
% -
% Look at $\bf \Lambda$. What is the rank of $\bf P$?
% +
% check your answer using:
% 'rank' function, type 'help rank' to know more about the command
% or by plotting the eigenvalues, e.g., 'plot(diag(E))'
% -
% Note that the eigenvalues are not necessarily sorted (because generally they may be complex). We can find the sorting order and correct for it:
% +
% uncomment code below to sort (remove '%' in front of code)
% Warning: Check that the name of the variable match the ones you used!!
%[~,index] = sort(diag(Lambda),'descend')
%U = U(:, index); Lambda = Lambda(index,index)
% -
% Check that you still have the same $\bf P$!
% +
% construct Pnew using the sorted eigenvectors and eigenvalues, i.e., U and Lambda
%Pnew = [code]
%disp(Pnew)
%disp(P)
% -
% ##### d. Split ${\bf U} = [{\bf u}_1, {\bf U}_2]$ where ${\bf u}_2$ is the first column of $\bf U$, and ${\bf U}_2$ are the remaining columns. How does ${\bf u}_1$ relate to ${\bf v}$? And ${\bf U}_2$?
% +
% add code
% u1 = U([code])
% U2 = U([code])
% -
% Check that ${\bf u}_1 = \alpha{\bf v}$, and ${\bf U}_2^H{\bf v} = \bf 0$. Also check that ${\bf PU}_2 = \bf 0$. Why do these properties hold?
% +
% add code
% [code] to check scaling property of u_1 and v
% [code] to check orthogonality property of U_2 with v
% [code] to check orthogonality property of P and U_2
% -
% ##### e. Define the projection on the orthogonal complement of $\bf v$,
%
% $$
% {\bf P}^{\perp} = \bf I - P.
% $$
% +
% add code
% Pperp = [code]
% -
% Check that ${\bf P}^{\perp} = {\bf U}_2{\bf U}_2^H$. Why is that?
% +
% add code
% [code] to check property
% -
% Why is ${\bf U}_2{\bf U}_2^H$ a projection? Why don't we have to write ${\bf U}_2({\bf U}_2^H{\bf U}_2)^{-1}{\bf U}_2^H$?
% +
% hint: check your answer trying U2'*U2
% [code]
% -
% ##### f. Generate a random vector $\bf x$. Split $\bf x$ into ${\bf x}_{\rm par}:=\bf Px$ and ${\bf x}_{\rm perp} := {\bf P}^\perp{\bf x}$.
% +
% add code
% x = [code]
% xpar = [code]
% xperp = [code]
% -
% Verify hat ${\bf x = Px + P}^{\perp}{\bf x}$
% +
% add code
% [code]
% -
% What is the geometric picture that goes with this?
% *Extra:* If you want, you can generalize this exercise to a matrix $\bf V$ consiting of $2$ (or more) random columns. This works as long as $\bf V$ is a 'tall' matrix (more rows than columns).
% ## 2. Singular value decomposition
% The singular value decomposition (SVD) is closely related to an eigenvalue decomposition, but is more general: it exists for any matrix (can be rectangular). It will be often used in future
% courses in the MSc (you might see it referred to as PCA as well). A short intro follows in Appendix A of the PDF.
% ##### a. Create a matlab function to construct a (complex) vector ${\bf a}(\theta)$:
%
% ` a = @(theta) [code]` (this is what is called inline function)
% where $\theta$ is an angle (in radians or degrees) and $M$ is the dimension of $\bf a$
%
% $$
% {\bf a}(\theta) = \begin{bmatrix}
% 1 \\ e^{-j\phi} \\ e^{-j2\phi} \\ \vdots \\ e^{-j(M-1)\phi}
% \end{bmatrix}, \;\; \phi = \pi\sin(\theta),\; j =\sqrt{-1}
% $$
%
% This vector is called the direction vector in array signal processing (see the ET4147 course later this year). The entries are phases corresponding to propagation delays experienced by a plane wave signals hitting an antenna array, and it occurs in communication (antenna arrays), radar, radio astronomy, ultrasound, and MRI.
% hint: an inline function, for example, to compute pi*sin(theta) is
phi = @(theta) pi*sin(theta);
% phi should be 'zero' for theta = pi
disp(phi(pi))
% +
% use the above inline function to make an inline function for a(theta)
% a = @(theta) [code]
% -
% Let $A = [{\bf a}(\theta_1), {\bf a}(\theta_2)]$ where $\theta_1 = 0^o$, $\theta_2 = 30^o$ (convert this to radians if needed). Let $\bf S$ be a random matrix with $2$ rows and $N = 20$ samples,
% +
% add code
% generate A matrix
% a_1 = a([code])
% a_2 = a([code])
% A = [code]
% generate random matrix
% hint: use 'randn' function
% S = [code]
% -
% Then generate a data matrix
%
% $$
% {\bf X = AS}
% $$
% +
% add code
% X = [code]
% -
% What do you think is the rank of $\bf X$?
% ##### b. Construct ${\bf R = XX}^H$. What is the rank of $\bf R$?
% +
% add code
% R = [code]
% -
% In the course we will see this matrix very often (correlation matrix).
% ##### c. Compute the SVD of ${\bf X = U\Sigma V}^H$. Also compute the eigenvalue decomposition of ${\bf R = Q\Lambda Q}^H$.
% +
% add code
% hint: use 'help svd' to see the sintaxis of SVD
% [U,Sigma, V] = [code]
% [Q,Lambda] = [code]
% -
% ##### d. Compare $\bf \Sigma$ to $\bf \Lambda$, verify ${\bf \Sigma}^2 = \bf \Lambda$, up to _sorting_. What is the rank of $\bf X$, as judged from $\bf\Sigma$? Very that $\bf R$ is a positive matrix: its eigenvalues are _non-negative_.
% +
% try something if needed to corroborate your conclusions.
% add code
% -
% Compare $\bf U$ to $\bf Q$: show that they are the same, up to a permutation of the columns and a (complex) scaling of the columns. You can do that by checking ${\bf U}^H\bf Q$.
% +
% add code
% [code]
% -
% ##### e. Suppose we compute an SVD of $\bf R$.
% +
% add code
% [U1,Sigma1,V1] = [code]
% -
% Does that give the same results as the eigenvalue decomposition of $\bf R$? What is the relation between ${\bf U}_1$ and ${\bf V}_1$?
% +
% add code
% try, for example, U_1'*V1 to corroborate your conclusions
% -
% ##### f. Plot the singular values:
% +
% uncomment the following commands
% plot(diag(Sigma1),'+');
% hold on
% -
% ##### g. Now, take $\theta_2 = 5^o$. Regenerate, $\bf X$, recompute the SVD, and plot the new singular values (use a different color)
% +
% add code here
% X = [code]
% R = [code]
% [U, Sigma2, V] = [code]
% [plot Sigma1]
% hold on
% [plot Sigma2]
% -
% ##### h. Now, add a bit of (complex) noise to $\bf X$, i.e., write
% `X1 = X + 0.01*( randn(size(X)) + 1i*randn(size(X)) );`
% +
% add code
% X1 = [code]
% -
% Compute the singular values of ${\bf X}_1$, and plot them in the same plot (use a different color). What do you observe?
% +
% add code
% [U1, Sigma1, V1] = [code]
% -
% Compare ${\bf U}$ to ${\bf U}_1$ using ${\bf U}^H{\bf U}_1$. We would want to conclude that the first two singular vectors have hardly changed (and span the same space), while the other $3$ columns might be quite different, but still span the same space. How can you conclude that by looking at ${\bf U}^H{\bf U}_1$?
% +
% add code
% inspect U'*U1
% -
% We will see applications of this when we discuss the MUSIC algorithm at the end of the course.
% ##### i. Note the size of $\bf \Sigma$ and of $\bf V$. In fact, $\bf X$ and $\bf \Sigma$ have the same size, and $\bf \Sigma$ contains many columns with just zeros: not very efficient!
% Alternatively, we almost always compute the ’economy-size SVD’,
% `[U,Sigma,V] = svd(X,'econ')`
% +
% add code for economy size SVD
% [U, Sigma, V] = [code]
% -
% Check the size of the resulting matrices. Check that still ${\bf X} = {\bf U\Sigma V}^H$. Check that ${\bf V}^H{\bf V = I}$.
% +
% add code
% hint: to check size use command 'size()'
% [code]
% [code] to check X
% [code] to check self inner product of V
% -
% Now $\bf\Sigma$ is square, and ${\bf V}$ is _rectangular_. It _cannot_ be a _unitary_ matrix anymore. We
% usually are not interested in the columns that were dropped.
% If $\bf X$ was tall, then the economy-size SVD will result in $\bf U$ being truncated to the size of ${\bf X}$ (with
% ${\bf U}^H{\bf U = I})$, while now $\bf V$ remains square. We’ll see an example in the next section.
% ## 3. Convolution and equalization
% The matrix equation corresponding to a convolution $y[n] = x[n] \ast h[n] = \displaystyle\sum_{k=0}^{L-1} h[k] x[n-k]$ is
% $$
% \bf y = \bf H \bf x \quad \Leftrightarrow \quad
% \begin{bmatrix}
% \fbox{$y[0]$} \\
% y[1] \\
% y[2] \\
% \vdots \\
% \vdots \\
% \vdots \\
% \vdots \\
% y[N_y-1] \end{bmatrix}
% =
% \begin{bmatrix}
% \fbox{$h[0]$} & & & {\textbf 0}\\
% h[1] & h[0] \\
% h[2] & h[1] & \ddots & \\
% \vdots & h[2] & \ddots & h[0] \\
% h[L-1] & \vdots & \ddots & h[1] \\
% & h[L-1] & \ddots & h[2] \\
% & & \ddots & \vdots \\
% {\textbf 0} & & & h[L-1]
% \end{bmatrix}
% \begin{bmatrix}
% \fbox{$x[0]$} \\ \vdots \\ x[N_x-1] \end{bmatrix}
% \label{eq:conv} \tag{1}
% $$
%
% where the ``box'' indicates the location of time-index 0,
% $L$ is the channel length (assuming an FIR channel),
% $N_x$ is the length of the input sequence (subsequent samples are
% supposed to be zero), and $N_y$ is the length of the output sequence
% (ignoring the other samples).
% Note that $\bf H$ has size $N_x+L-1 \times N_x$ so that $N_y = N_x + L-1$.
% $\bf H$ is always tall.
%
% $\bf H$ has a Toeplitz structure: constant along diagonals. That structure
% always appears when we have shift invariance--we will see it often
% during the course.
% ##### a. Take a simple channel, `h = [1 2 3]'`, and input signal, `x = randn(4,1)`. Generate `y[n]` using `y = filter(h,1,x)`.
% +
% add code here
% [code]
% -
% Also create the matrices in equation (1) and check that $\bf y = \bf H \bf x$. In matlab, you can use the function `toeplitz`:
% +
% add code here
% hint: H = toeplitz([h; 0; 0; 0], [h(1) 0 0 0])
% [code]
% -
% ##### b. Check $\bf H^H \bf H$ and $\bf H \bf H^H$. Are these Toeplitz matrices?
% +
% add code here
% [code]
% -
% Suppose we observe $\bf y$ and know the channel $\bf h$.
% The input signal can be estimated by taking a left inverse $\bf H^\dagger$ of $\bf H$, such that $\bf H^\dagger \bf H = \bf I$. We can usually take
%
% $$
% \bf H^\dagger = (\bf H^H \bf H)^{-1} \bf H^H
% $$
% where, for now, we assume that $\bf H^H \bf H$ is invertible. This results in
%
% $$
% \hat{\bf x} = \bf H^\dagger \bf y = (\bf H^H \bf H)^{-1} \bf H^H \bf y \,.
% \label{eq:loc:sourceestim1}
% $$
% ##### c. Construct $\bf H^\dagger$. Verify that $\bf H^\dagger \bf H = \bf I$ and that $\bf H \bf H^\dagger$ is a projection. Explain why this is the case.
% +
% add code here
% [code]
% -
% ##### d. Compute the singular values of $\bf H$ and of $\bf H^\dagger$. What do you notice?
% +
% add code here
% [code]
% -
% ##### e. Verify in matlab that $\hat{\bf x} = \bf H^\dagger \bf y$ gives back the original signal (noiseless case).
% +
% add code here
% [code]
% -
% ## 4. Equalization: a rank-deficient case
% Suppose that, in equation (1), we start to measure the
% output only after the input signal has stopped (e.g., in communication,
% we can listen only after we stopped transmitting).
% Also, suppose that the channel is generated by an
% auto-regressive (AR) model:
% $$
% H(z) = \frac{1}{1-a z^{-1}}
% \qquad\Leftrightarrow\qquad
% \mathbf{h} = [1,\; a,\; a^2,\; \cdots]^T \,.
% $$
%
% Since the impulse response is infinitely long, we obtain
%
% $$
% \mathbf{y} = \mathbf{H} \mathbf{x} \quad \Leftrightarrow \quad
% \begin{bmatrix}
% y[N_x-1] \\ y[N_x] \\ y[N_x+1] \\ \vdots \\ \vdots \\ y[N_y-1] \end{bmatrix} =
% \begin{bmatrix} h[N_x-1] & \cdots & h[1] & h[0] \\ h[N_x] & \cdots & h[2] & h[1] \\\vdots & \vdots & h[3] & h[2] \\ \vdots & \vdots & \vdots & h[3] \\ \vdots & \vdots & \vdots & \vdots \\ h[N_y-1] & \vdots & \vdots & \vdots \end{bmatrix}
% \begin{bmatrix}
% x[0] \\ \vdots \\ x[N_x-1] \end{bmatrix}
% \label{eq:conv2}
% $$
%
% This is similar to (1), but now the top triangle part is clipped off.
% ##### a. Take the AR parameter $a =0.8$, take $N_x = 3$, $N_y = 20$, and generate the above data model in matlab.
% +
% add code here
% [code]
% -
% ##### b. Using the SVD, what is the rank of $\bf H$?
% +
% add code here
% [code]
% -
%
% If the (economy-size) SVD of $\bf H$ is $\bf H = \bf U \bf \Sigma \bf V^H$, then the
% (economy-size) SVD of
% $\bf H^\dagger$ is $\bf H^\dagger = \bf V \bf \Sigma^{-1} \bf U^H$.
% ##### c.Why do we refer to the economy-size SVD here?
% +
% type your answer here
% -
% ##### d. Using the SVD properties, explain why $\bf H^\dagger \bf H = \bf I$ and $\bf H \bf H^\dagger$ is a projection.
% +
% type your answer here
% -
% If $\bf \Sigma$ has entries that are (nearly) zero, then $\bf \Sigma^{-1}$ has
% entries that go to infinity. We define the pseudo-inverse of
% $\bf \Sigma$ as
%
% $$
% \bf \Sigma = \begin{bmatrix}
% \sigma_1 \\
% & \sigma_2 \\
% && 0 \\
% &&& 0 \end{bmatrix}
% \qquad \Rightarrow \qquad
% \bf \Sigma^\dagger = \begin{bmatrix}
% 1/\sigma_1 \\
% & 1/\sigma_2 \\
% && 0 \\
% &&& 0 \end{bmatrix}
% $$
%
% So, the nonzero entries are inverted, and the zero entries are kept zero. (In practice, we specify a tolerance $\epsilon$ and do not invert entries that are smaller than $\epsilon$.)
%
% The (Moore-Penrose) pseudo-inverse of a matrix $\bf X$ is then
%
% $$
% \bf X = \bf U \bf \Sigma \bf V^H
% \qquad \Rightarrow \qquad
% \bf X^\dagger = \bf V \bf \Sigma^\dagger \bf U^H
% $$
%
% This generalizes the left inverse that we saw before. it satisfies 4
% properties:
%
% $$
% \bf X \bf X^\dagger \bf X = \bf X \,,\qquad
% \bf X^\dagger \bf X \bf X^\dagger = \bf X^\dagger\,,\qquad
% \bf X \bf X^\dagger = \bf P_c \,,\qquad
% \bf X^\dagger \bf X = \bf P_r \,,\qquad
% $$
%
% where $\bf P_c$ is a projector onto the column span of $\bf X$, and $\bf P_r$
% a projector onto its row span.
%
% In matlab, you say `Xi = pinv(X);`
% and you can specify a tolerance $\epsilon$ as well.
%
% ##### e. Compute the pseudo-inverse of $\bf H$ in matlab.
% +
% add code here
% [code]
% -
% ##### f. Compute $\hat{\bf x} = \bf H^\dagger \bf y$. Do you get back the original $\bf x$?
% +
% add code here
% [code]
% -
% ## 5. Sinusoids
% ##### a. Generate a time domain sequence $x[n] = e^{j \omega n}$, for $\omega = 0.2 \pi$ and $n = 1, \cdots, N$. Take $N=20$.
% ##### b. Create a data matrix (Hankel matrix)
%
% $$
% \bf X = \begin{bmatrix}
% x[1] & x[2] & \cdots & x[N-M+1]\\
% x[2] & x[3] & \vdots & \vdots\\
% x[3] & x[4] & \vdots & \vdots \\
% \vdots & & & \vdots \\
% x[M] & x[M+1] & \cdots & x[N]
% \end{bmatrix}
% $$
%
% Take $M=5$. You can use the Matlab function `hankel`.
% +
% add code here
% [code]
% -
% Hankel matrices are constant along anti-diagonals. They are similar
% to Toeplitz matrices (by permuting columns or rows), so they also
% appear often in the context of shift-invariant systems.
% ##### c. What is the rank of $\bf X$? (Use `svd` for this.)
% +
% add code here
% [code]
% -
% ##### d. Can you theoretically explain the rank?
% Look at the shift-invariance structure:
% $$
% \begin{bmatrix}
% x[2] \\
% x[3] \\
% x[4] \\
% \vdots \\
% x[M+1]
% \end{bmatrix}
% =
% \begin{bmatrix}
% x[1] \\
% x[2] \\
% x[3] \\
% \vdots \\
% x[M]
% \end{bmatrix}
% e^{j\omega}
% $$
%
% This shows that the 2nd column is the same as the first, except for a
% scaling. Extend this argument to the full matrix.
% ##### e. Now, repeat: generate a time domain sequence $x[n] = \sin(\omega n)$, for $\omega = 0.2 \pi$ and $n = 1, \cdots, N$. Take $N=20$. Construct the same matrix $\bf X$, check the rank, explain.
% +
% add code here
% [code]
% -
% ##### f. Let $y[n] = x[n] + e[n]$, where $e[n]$ is a small noise disturbance, e.g. `e = 0.1 * randn(1,20)`. Construct a Hankel matrix $\bf Y$ from $y[n]$, and compute the SVD.
% +
% add code here
% [code]
% -
% Compare the singular values to those of $\bf X$.
% +
% add code here
% [code]
% -
% ##### g. To remove the noise, compute the SVD $\bf Y = \bf U \bf \Sigma\bf V^H$, and set all small singular values of $\bf Y$ equal to zero. This gives an approximate matrix $\bf{\hat{\Sigma}}$. Compute $\bf{\hat{Y}} = \bf U \bf{\hat{\Sigma}}\bf V^H$. This is called the Truncated SVD.
%
% +
% add code here
% [code]
% -
% From the first column and bottom row, regenerate a time-domain signal $\hat{y}[n]$. In a plot, compare $x[n]$, $y[n]$ and $\hat{y}[n]$. How well did you remove the noise?
% +
% add code here
% [code]
|
index.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prepare All Data
# ### Import necessary modules
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pickle
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# -
# ### Processing component information
# ### Load training and test datasets
# +
train = pd.read_csv('input/train_set.csv', parse_dates=[2,])
# y_train = train ["cost"].values
# train_df = train_df.drop(['cost'], axis = 1)
# y is converted to log(1+y), RMSLE is then coverted into RMSE for new y
# y_train = np.log1p(y_train)
# generate fake negative id for training data
train['id'] = -1 * np.arange(0, len(train))
train = train.set_index('id')
test = pd.read_csv('input/test_set.csv', parse_dates=[3,], index_col = 'id')
test["cost"] = 0
# merge training data X and test X together
# so the training data and test data can be preprocessed in the same way
all_data = pd.concat([train, test])
# -
# ### Merge tube, specs, and end form data
# +
all_data = all_data.reset_index()
# merge tube data
tube = pd.read_csv('input/tube.csv', na_values = ['NONE', 9999], true_values = 'Y', false_values = 'N')
all_data = pd.merge(all_data, tube, on='tube_assembly_id', how='left')
# merge bom data, bom = bill of material
# bom = pd.read_csv('input/bill_of_materials.csv')
# all_data = pd.merge(all_data, bom, on='tube_assembly_id', how='left')
# specs of the tubes
specs = pd.read_csv('input/specs.csv')
specs[specs.notnull()] = 1
specs = specs.fillna(0)
all_data['spec_num'] = specs[['spec' + str(x) for x in range(1, 10)]].sum(axis = 1)
# merge end forming data
end_form = pd.read_csv('input/tube_end_form.csv')
# all_data.loc[all_data['end_a'] == "NONE", 'end_a_forming'] = -999
# all_data.loc[all_data['end_x'] == "NONE", 'end_x_forming'] = -999
for idx,row in end_form.iterrows():
if row['forming'] == 'Yes':
end_forming_value = 1
if row['forming'] == 'No':
end_forming_value = 0
all_data.loc[all_data['end_a'] == row['end_form_id'], 'end_a_forming'] = end_forming_value
all_data.loc[all_data['end_x'] == row['end_form_id'], 'end_x_forming'] = end_forming_value
bom_comp = pickle.load(open('bom_comp.pkl', 'rb'))
all_data = pd.merge(all_data, bom_comp, on='tube_assembly_id', how='left')
all_data = all_data.set_index('id', drop = True)
float_column = list(all_data.select_dtypes(include=['float64']).columns)
all_data[float_column] = all_data[float_column].fillna(0)
# year and month are treated as feature
all_data['year'] = all_data['quote_date'].dt.year
all_data['month'] = all_data['quote_date'].dt.month
all_data = all_data.drop('quote_date', axis = 1)
# there is soft leak in the assembly id
# assembly id is treated as a feature
all_data['tube_assembly_id'] = all_data['tube_assembly_id'].str[3: ]
all_data['tube_assembly_id'] = all_data['tube_assembly_id'].astype('int64')
all_data['cross_section'] = all_data['diameter'] ** 2 - (
all_data['diameter'] - all_data['wall']) ** 2
# price = fixed price + variable price / quantity
all_data['quantity'] = 1 / all_data['quantity']
# -
#Dump the merged data to use in modeling
pickle.dump(all_data, open('all_data.pkl', 'wb'))
all_data.info()
all_data.head()
all_data.tail()
all_data.head()
|
Caterpillar Tube Pricing/Preprocessing_All_Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import numpy as np
# +
health_info = r"C:\Users\david\Desktop\Davids Branch\Row-2-Group-Project\Health Insurance Coverage by State CSV.csv"
health_info = pd.read_csv(health_info)
health_info
cancer_info = r"C:\Users\david\Desktop\Davids Branch\Row-2-Group-Project\Rupesh Cancer Data.csv"
cancer_info = pd.read_csv(cancer_info)
# -
obesity_info = r"C:\Users\david\Desktop\Davids Branch\Row-2-Group-Project\Obesity Rates by State.csv"
obesity_info = pd.read_csv(obesity_info)
obesity_info
obesity_info_no_Hawaii = obesity_info[obesity_info["State"]!= "Hawaii"]
obesity_info_no_Hawaii
# +
plt.scatter(obesity_info_no_Hawaii["Obesity Prevalence"], cancer_death_rate)
plt.xlabel("Obesity Prevalence")
plt.ylabel("Cancer Death Rate per 100,000")
x_axis= obesity_info_no_Hawaii["Obesity Prevalence"]
y_axis= cancer_death_rate
correlation = st.pearsonr(x_axis,y_axis)
print(f"The pearson correlation between both factors is {round(correlation[0],2)}")
# -
health_info
uninsured_rates = health_info["Uninsured Percentage (2016)"]
uninsured_rates
obesity_rates = obesity_info["Obesity Prevalence"]
obesity_rates
# +
# data1 = obesity_rates
# data2 = uninsured_rates
# fig, ax1 = plt.subplots()
# color = 'tab:red'
# ax1.set_xlabel('States')
# ax1.set_ylabel('Obesity Rates', color=color)
# ax1.scatter(obesity_info['State'], data1, color=color)
# ax1.tick_params(axis='y', labelcolor=color)
# ax2 = ax1.twinx() # instantiate a second axes that shares the same x-axis
# color = 'tab:blue'
# ax2.set_ylabel('Unisured Rates', color=color) # we already handled the x-label with ax1
# ax2.scatter(obesity_info['State'], data2, color=color)
# ax2.tick_params(axis='y', labelcolor=color)
# fig.tight_layout() # otherwise the right y-label is slightly clipped
# plt.xticks(rotation=45)
# plt.show()
plt.scatter(uninsured_rates, obesity_rates)
plt.xlabel("Percentage of Population Uninsured")
plt.ylabel("Obesity Prevalence")
plt.show
x_axis= uninsured_rates
y_axis= obesity_rates
correlation = st.pearsonr(x_axis,y_axis)
print(f"The pearson correlation between both factors is {round(correlation[0],2)}")
# -
cancer_info
# +
cancer_useful_info = cancer_info[["Incidence Rate", "Death Rate"]]
cancer_useful_info
cancer_incidence_rate = cancer_info["Incidence Rate"]
cancer_death_rate = cancer_info[cancer_info["Death Rate", "State"]
cancer_death_per_hundred = cancer_info["Cancer Death_per_hundred_cancer_patient"]
# +
#drop Hawaii
list(uninsured_rates)
uninsured_rates_no_Hawaii = health_info[health_info["State"]!= "Hawaii"]
uninsured_rates_no_Hawaii
plt.scatter(uninsured_rates_no_Hawaii['Uninsured Percentage (2016)'], cancer_death_rate)
plt.xlabel("Percentage of Population Uninsured ")
plt.ylabel("Cancer Death Rate per 100,000")
x_axis= uninsured_rates_no_Hawaii['Uninsured Percentage (2016)']
y_axis= cancer_death_rate
correlation = st.pearsonr(x_axis,y_axis)
print(f"The pearson correlation between both factors is {round(correlation[0],2)}")
# -
# +
plt.scatter(uninsured_rates_no_Hawaii['Uninsured Percentage (2016)'], cancer_incidence_rate)
correlation = st.pearsonr(uninsured_rates_no_Hawaii['Uninsured Percentage (2016)'],cancer_incidence_rate)
print(f"The pearson correlation between both factors is {round(correlation[0],2)}")
plt.xlabel("Percentage of Population Uninsured ")
plt.ylabel("Cancer Incidence Rate per 100,000")
# -
# +
plt.scatter(uninsured_rates_no_Hawaii['Uninsured Percentage (2016)'], cancer_death_per_hundred)
correlation = st.pearsonr(uninsured_rates_no_Hawaii['Uninsured Percentage (2016)'], cancer_death_per_hundred)
print(f"The pearson correlation between both factors is {round(correlation[0],2)}")
plt.xlabel("Percentage of Population Uninsured ")
plt.ylabel("Cancer Death Rate per 100 Incidences")
# -
# +
#max values
health_insurance_extremes = uninsured_rates_no_Hawaii.sort_values("Uninsured Percentage (2016)")
health_insurance_extremes.head(10)
health_insurance_extremes.tail(10)
health_insurance_extremes_final = pd.concat([health_insurance_extremes.head(10),health_insurance_extremes.tail(10)])
health_insurance_extremes_final
# cx= bx.sort_values("Cancer Death_per_hundred_cancer_patient")
# cx.head(10)
# cx.tail(10)
# dx = pd.concat([cx.head(10),cx.tail(10)])
# dx
# -
# +
#joined_health_insurance_extremes = pd.merge(health_insurance_extremes_final, cancer_info[["Uninsured Percentage (2016)", "Incidence Rate", "State" ]], on = "State")
joined_health_insurance_extremes = pd.merge(health_insurance_extremes_final, cancer_info, on="State")
joined_health_insurance_extremes
# -
plt.scatter(joined_health_insurance_extremes["Uninsured Percentage (2016)"], joined_health_insurance_extremes["Death Rate"])
# +
plt.scatter(health_insurance_extremes.head(10)["Uninsured Percentage (2016)"], joined_health_insurance_extremes.head(10)["Death Rate"])
x_axis= health_insurance_extremes.head(10)["Uninsured Percentage (2016)"]
y_axis= joined_health_insurance_extremes.head(10)["Death Rate"]
correlation = st.pearsonr(x_axis,y_axis)
plt.xlabel("Percentage of Population Uninsured ")
plt.ylabel("Cancer Death Rate per 100,000")
print(f"The pearson correlation between both factors is {round(correlation[0],2)}")
# -
cancer_info
|
Cancer Comparisons Final.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Fix Alteration Filing Metadata
# ### Environment Set up
# %run /workspaces/lear/tests/data/default-bcr-business-setup-TEST.ipynb
import dpath
import json
from contextlib import suppress
from legal_api import db
from legal_api.models import Filing
from sqlalchemy import desc
from sqlalchemy.orm.attributes import flag_modified
# ### Get Completed Filings by type
def get_completed_filings_by_type(filing_type: str):
"""Return the filings of a particular type."""
filings = db.session.query(Filing). \
filter(Filing._filing_type == filing_type). \
filter(Filing._status == Filing.Status.COMPLETED). \
order_by(desc(Filing.filing_date)). \
all()
return filings
# ### Update Alteration Filing Metadata
def update_alteration_metadata():
completed_alteration_filings = get_completed_filings_by_type('alteration')
for filing in completed_alteration_filings:
try:
filing_meta_data = filing._meta_data
filing_meta_data['legalFilings'] = ['alteration']
print(filing_meta_data)
# Update the from and to legal type always.
alteration_meta = {}
from_legal_type = dpath.util.get(filing._filing_json, '/filing/business/legalType')
to_legal_type = dpath.util.get(filing._filing_json, '/filing/alteration/business/legalType')
if from_legal_type and to_legal_type:
alteration_meta = {**alteration_meta, **{'fromLegalType': from_legal_type,
'toLegalType': to_legal_type}}
# Update the fromLegalName and toLegalName if there is a name change
with suppress(IndexError, KeyError, TypeError):
from_legal_name = dpath.util.get(filing._filing_json, '/filing/business/legalName')
identifier = dpath.util.get(filing._filing_json, '/filing/business/identifier')
name_request_json = dpath.util.get(filing._filing_json, '/filing/alteration/nameRequest')
to_legal_name = name_request_json.get('legalName', identifier[2:] + ' B.C. LTD.')
if from_legal_name != to_legal_name:
alteration_meta = {**alteration_meta, **{'fromLegalName': from_legal_name,
'toLegalName': to_legal_name}}
filing_meta_data['alteration'] = alteration_meta
filing._meta_data = filing_meta_data
# For ORM to help identify that the json has changed.
flag_modified(filing, "_meta_data")
db.session.add(filing)
db.session.commit()
print( f'\033[92m Updated {filing.id} - {filing._meta_data}')
except Exception as e:
print(f'\033[91mUpdate failed for filing Id - {filing.id}')
update_alteration_metadata()
|
tests/data/Alterations_update_metadata/fix_alteration_filing_metadata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="DGPlYumZnO1t"
# # Introduction to Linear Regression
#
#
#
# ## Learning Objectives
#
# 1. Analyze a Pandas Dataframe
# 2. Create Seaborn plots for Exporatory Data Analysis
# 2. Train a Linear Regression Model using Scikit-Learn
#
#
# ## Introduction
# This lab is in introduction to linear regression using Python and Scikit-Learn. This lab serves as a foundation for more complex algortithms and machine learning models that you will encounter in the course. We will train a linear regression model to predict housing price.
#
# Each learning objective will correspond to a __#TODO__ in the [student lab notebook](https://github.com/GoogleCloudPlatform/training-data-analyst/blob/master/courses/machine_learning/deepdive2/ml_on_gcloud_v2/labs/02_intro_linear_regression.ipynb) -- try to complete that notebook first before reviewing this solution notebook.
# + [markdown] colab_type="text" id="AsHg6SD2nO1v"
# ### Import Libraries
# + colab={} colab_type="code" id="gEXV-RxPnO1w"
import os
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import seaborn as sns # Seaborn is a Python data visualization library based on matplotlib.
# %matplotlib inline
# + [markdown] colab_type="text" id="dr2TkzKRnO1z"
# ### Load the Dataset
#
# We will use the [USA housing prices](https://www.kaggle.com/kanths028/usa-housing) dataset found on Kaggle. The data contains the following columns:
#
# * 'Avg. Area Income': Avg. Income of residents of the city house is located in.
# * 'Avg. Area House Age': Avg Age of Houses in same city
# * 'Avg. Area Number of Rooms': Avg Number of Rooms for Houses in same city
# * 'Avg. Area Number of Bedrooms': Avg Number of Bedrooms for Houses in same city
# * 'Area Population': Population of city house is located in
# * 'Price': Price that the house sold at
# * 'Address': Address for the house
# -
# Here, we create a directory called usahousing. This directory will hold the dataset that we copy from Google Cloud Storage.
if not os.path.isdir("../data/usahousing"):
os.makedirs("../data/usahousing")
# Next, we copy the Usahousing dataset from Google Cloud Storage.
# !gsutil cp gs://feat_eng/data/USA_Housing.csv ../data/usahousing
# Then we use the "ls" command to list files in the directory. This ensures that the dataset was copied.
# !ls -l ../data/usahousing
# Next, we read the dataset into a Pandas dataframe.
# + colab={} colab_type="code" id="CzrXJI8VnO10"
df_USAhousing = pd.read_csv('../data/usahousing/USA_Housing.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="Y6VJQ1tdnO12" outputId="7a1d4eed-3e83-44a8-f495-a9b74444d3ec"
# Show the first five row.
df_USAhousing.head()
# -
# Let's check for any null values.
df_USAhousing.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 297} colab_type="code" id="nRTsvSzqnO17" outputId="f44ad14e-5fb4-4c70-e71c-9d149bca4869"
df_USAhousing.describe()
# -
df_USAhousing.info()
# Let's take a peek at the first and last five rows of the data for all columns.
print(df_USAhousing,5)
# + [markdown] colab_type="text" id="QWVdsrmgnO1_"
# ## Exploratory Data Analysis (EDA)
#
# Let's create some simple plots to check out the data!
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="ESLg7Y0tnO1_" outputId="69b971f3-142b-4e3b-efe3-f759dc532819"
sns.pairplot(df_USAhousing)
# + colab={"base_uri": "https://localhost:8080/", "height": 296} colab_type="code" id="SOsTLClWnO2B" outputId="b8a78674-5ddb-4706-90b4-37d7d83e8092"
sns.distplot(df_USAhousing['Price'])
# + colab={"base_uri": "https://localhost:8080/", "height": 434} colab_type="code" id="NFnb70lhnO2D" outputId="5a6d8960-94c4-4b36-e53f-6b4be076b571"
sns.heatmap(df_USAhousing.corr())
# + [markdown] colab_type="text" id="OIPKB4hanO2F"
# ## Training a Linear Regression Model
#
# Regression is a supervised machine learning process. It is similar to classification, but rather than predicting a label, we try to predict a continuous value. Linear regression defines the relationship between a target variable (y) and a set of predictive features (x). Simply stated, If you need to predict a number, then use regression.
#
# Let's now begin to train our regression model! We will need to first split up our data into an X array that contains the features to train on, and a y array with the target variable, in this case the Price column. We will toss out the Address column because it only has text info that the linear regression model can't use.
# -
# ### X and y arrays
#
# Next, let's define the features and label. Briefly, feature is input; label is output. This applies to both classification and regression problems.
# + colab={} colab_type="code" id="ZEEGuBAnnO2F"
X = df_USAhousing[['Avg. Area Income', 'Avg. Area House Age', 'Avg. Area Number of Rooms',
'Avg. Area Number of Bedrooms', 'Area Population']]
y = df_USAhousing['Price']
# + [markdown] colab_type="text" id="X97FWdDOnO2H"
# ## Train - Test - Split
#
# Now let's split the data into a training set and a testing set. We will train out model on the training set and then use the test set to evaluate the model. Note that we are using 40% of the data for testing.
#
# #### What is Random State?
# If an integer for random state is not specified in the code, then every time the code is executed, a new random value is generated and the train and test datasets will have different values each time. However, if a fixed value is assigned -- like random_state = 0 or 1 or 101 or any other integer, then no matter how many times you execute your code the result would be the same, e.g. the same values will be in the train and test datasets. Thus, the random state that you provide is used as a seed to the random number generator. This ensures that the random numbers are generated in the same order.
# + colab={} colab_type="code" id="fS99Llq8nO2J"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=101)
# + [markdown] colab_type="text" id="uh_3f6dgnO2K"
# ## Creating and Training the Model
# + colab={} colab_type="code" id="4E_FGrFEnO2L"
from sklearn.linear_model import LinearRegression
# + colab={} colab_type="code" id="dwh-yr1VnO2M"
lm = LinearRegression()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="NAoKCQtpnO2O" outputId="889705d3-f50e-41a0-a800-72d08717a150"
lm.fit(X_train,y_train)
# + [markdown] colab_type="text" id="hQi2T_gbnO2P"
# ## Model Evaluation
#
# Let's evaluate the model by checking out it's coefficients and how we can interpret them.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="IpunBLdtnO2Q" outputId="e5745749-fd73-435f-8ebe-0edcbf03e489"
# print the intercept
print(lm.intercept_)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="DjmfEks3nO2T" outputId="5fea28c6-404c-4f67-b96a-07a39e8b37e4"
coeff_df = pd.DataFrame(lm.coef_,X.columns,columns=['Coefficient'])
coeff_df
# + [markdown] colab_type="text" id="3bw5d5bvnO2V"
# Interpreting the coefficients:
#
# - Holding all other features fixed, a 1 unit increase in **Avg. Area Income** is associated with an **increase of \$21.52 **.
# - Holding all other features fixed, a 1 unit increase in **Avg. Area House Age** is associated with an **increase of \$164883.28 **.
# - Holding all other features fixed, a 1 unit increase in **Avg. Area Number of Rooms** is associated with an **increase of \$122368.67 **.
# - Holding all other features fixed, a 1 unit increase in **Avg. Area Number of Bedrooms** is associated with an **increase of \$2233.80 **.
# - Holding all other features fixed, a 1 unit increase in **Area Population** is associated with an **increase of \$15.15 **.
#
#
# + [markdown] colab_type="text" id="c_NExEFynO2X"
# ## Predictions from our Model
#
# Let's grab predictions off our test set and see how well it did!
# + colab={} colab_type="code" id="xmzvk_5OnO2Y"
predictions = lm.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="bQgBCRAnnO2a" outputId="75c72025-2a9b-4e31-bc72-0ebae691edb0"
plt.scatter(y_test,predictions)
# + [markdown] colab_type="text" id="aKW2IiSynO2b"
# **Residual Histogram**
# + colab={"base_uri": "https://localhost:8080/", "height": 279} colab_type="code" id="s_daF-wunO2b" outputId="f98e6963-ba34-412d-dedd-19b5f1f973d8"
sns.distplot((y_test-predictions),bins=50);
# + [markdown] colab_type="text" id="Znh-A9YrnO2d"
# ## Regression Evaluation Metrics
#
#
# Here are three common evaluation metrics for regression problems:
#
# **Mean Absolute Error** (MAE) is the mean of the absolute value of the errors:
#
# $$\frac 1n\sum_{i=1}^n|y_i-\hat{y}_i|$$
#
# **Mean Squared Error** (MSE) is the mean of the squared errors:
#
# $$\frac 1n\sum_{i=1}^n(y_i-\hat{y}_i)^2$$
#
# **Root Mean Squared Error** (RMSE) is the square root of the mean of the squared errors:
#
# $$\sqrt{\frac 1n\sum_{i=1}^n(y_i-\hat{y}_i)^2}$$
#
# Comparing these metrics:
#
# - **MAE** is the easiest to understand, because it's the average error.
# - **MSE** is more popular than MAE, because MSE "punishes" larger errors, which tends to be useful in the real world.
# - **RMSE** is even more popular than MSE, because RMSE is interpretable in the "y" units.
#
# All of these are **loss functions**, because we want to minimize them.
# + colab={} colab_type="code" id="ePMew8WdnO2d"
from sklearn import metrics
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="Ev152sRanO2f" outputId="aa8f66e7-7f1b-4c0e-c7a3-cd753ebad170"
print('MAE:', metrics.mean_absolute_error(y_test, predictions))
print('MSE:', metrics.mean_squared_error(y_test, predictions))
print('RMSE:', np.sqrt(metrics.mean_squared_error(y_test, predictions)))
# + [markdown] colab_type="text" id="0s8Veb58nO2g"
# Copyright 2020 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
|
courses/machine_learning/deepdive2/launching_into_ml/solutions/intro_linear_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Norod/my-colab-experiments/blob/master/Norod78_hebrew_gpt_neo_small.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Vp3XPuaTu9jl"
#
# # How to generate text: using different decoding methods for language generation with Transformers
# + [markdown] id="KxLvv6UaPa33"
# ### **Introduction**
#
# In recent years, there has been an increasing interest in open-ended language generation thanks to the rise of large transformer-based language models trained on millions of webpages, such as OpenAI's famous [GPT2 model](https://openai.com/blog/better-language-models/). The results on conditioned open-ended language generation are impressive, e.g. [GPT2 on unicorns](https://openai.com/blog/better-language-models/#samples), [XLNet](https://medium.com/@amanrusia/xlnet-speaks-comparison-to-gpt-2-ea1a4e9ba39e), [Controlled language with CTRL](https://blog.einstein.ai/introducing-a-conditional-transformer-language-model-for-controllable-generation/). Besides the improved transformer architecture and massive unsupervised training data, **better decoding methods** have also played an important role.
#
# This blog post gives a brief overview of different decoding strategies and more importantly shows how *you* can implement them with very little effort using the popular `transformers` library!
#
# All of the following functionalities can be used for **auto-regressive** language generation ([here](http://jalammar.github.io/illustrated-gpt2/) a refresher). In short, *auto-regressive* language generation is based on the assumption that the probability distribution of a word sequence can be decomposed into the product of conditional next word distributions:
# $$ P(w_{1:T} | W_0 ) = \prod_{t=1}^T P(w_{t} | w_{1: t-1}, W_0) \text{ ,with } w_{1: 0} = \emptyset, $$
#
# and $W_0$ being the initial *context* word sequence. The length $T$ of the word sequence is usually determined *on-the-fly* and corresponds to the timestep $t=T$ the EOS token is generated from $P(w_{t} | w_{1: t-1}, W_{0})$.
#
#
# Auto-regressive language generation is now available for `GPT2`, `XLNet`, `OpenAi-GPT`, `CTRL`, `TransfoXL`, `XLM`, `Bart`, `T5` in both PyTorch and Tensorflow >= 2.0!
#
# We will give a tour of the currently most prominent decoding methods, mainly *Greedy search*, *Beam search*, *Top-K sampling* and *Top-p sampling*.
#
# + [markdown] id="Si4GyYhOQMzi"
# Let's quickly install transformers and load the model.
# + id="XbzZ_IVTtoQe" colab={"base_uri": "https://localhost:8080/"} outputId="14e2ee95-e7e9-44e8-d970-67fa25fb2d73"
# !pip install tokenizers==0.10.2 transformers==4.5.1
# + id="ue2kOQhXTAMU" colab={"base_uri": "https://localhost:8080/"} outputId="618e0902-60d0-4dbf-db01-d80cb6dff4ff"
import tensorflow as tf
from transformers import AutoTokenizer, AutoModelForCausalLM
tokenizer = AutoTokenizer.from_pretrained("Norod78/hebrew-gpt_neo-small")
model = AutoModelForCausalLM.from_pretrained("Norod78/hebrew-gpt_neo-small", pad_token_id=tokenizer.eos_token_id)
# + id="xYX5szSH3dN7"
prompt_text = "אני נהנה לטייל עם הכלב החמוד שלי"
max_len = 128
# + colab={"base_uri": "https://localhost:8080/"} id="DY6B3YWd3O4X" outputId="59f07a36-b9fa-48ab-ca0f-caa0e7e9d717"
import numpy as np
import torch
seed = 1000
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
n_gpu = 0 if torch.cuda.is_available()==False else torch.cuda.device_count()
print(f"device: {device}, n_gpu: {n_gpu}")
np.random.seed(seed)
torch.manual_seed(seed)
if n_gpu > 0:
torch.cuda.manual_seed_all(seed)
model.to(device)
encoded_prompt = tokenizer.encode(
prompt_text, add_special_tokens=False, return_tensors="pt")
encoded_prompt = encoded_prompt.to(device)
if encoded_prompt.size()[-1] == 0:
input_ids = None
else:
input_ids = encoded_prompt
print(input_ids)
# + [markdown] id="a8Y7cgu9ohXP"
# ### **Greedy Search**
#
# Greedy search simply selects the word with the highest probability as its next word: $w_t = argmax_{w}P(w | w_{1:t-1})$ at each timestep $t$. The following sketch shows greedy search.
#
# 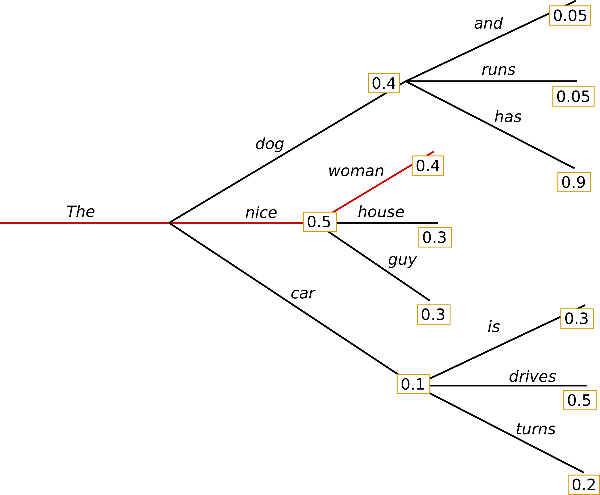
#
# Starting from the word $\text{"The"}$, the algorithm
# greedily chooses the next word of highest probability $\text{"nice"}$ and so on, so that the final generated word sequence is $\text{"The", "nice", "woman"}$ having an overall probability of $0.5 \times 0.4 = 0.2$.
#
# In the following we will generate word sequences using GPT2 on the context $(\text{"I", "enjoy", "walking", "with", "my", "cute", "dog"})$. Let's see how greedy search can be used in `transformers` as follows:
# + id="OWLd_J6lXz_t" colab={"base_uri": "https://localhost:8080/"} outputId="8055da89-69c7-4fc0-976a-a97eb1ebc0f2"
# generate text until the output length (which includes the context length) reaches max_len
greedy_output = model.generate(input_ids, max_length=max_len)
print("Output:\n" + 100 * '-')
print(tokenizer.decode(greedy_output[0], skip_special_tokens=True))
# + [markdown] id="BBn1ePmJvhrl"
# Alright! We have generated our first short text with GPT2 😊. The generated words following the context are reasonable, but the model quickly starts repeating itself! This is a very common problem in language generation in general and seems to be even more so in greedy and beam search - check out [Vijayakumar et al., 2016](https://arxiv.org/abs/1610.02424) and [Shao et al., 2017](https://arxiv.org/abs/1701.03185).
#
# The major drawback of greedy search though is that it misses high probability words hidden behind a low probability word as can be seen in our sketch above:
#
# The word $\text{"has"}$ with its high conditional probability of $0.9$ is hidden behind the word $\text{"dog"}$, which has only the second-highest conditional probability, so that greedy search misses the word sequence $\text{"The"}, \text{"dog"}, \text{"has"}$.
#
# Thankfully, we have beam search to alleviate this problem!
#
# + [markdown] id="g8DnXZ1WiuNd"
# ### **Beam search**
#
# Beam search reduces the risk of missing hidden high probability word sequences by keeping the most likely `num_beams` of hypotheses at each time step and eventually choosing the hypothesis that has the overall highest probability. Let's illustrate with `num_beams=2`:
#
# 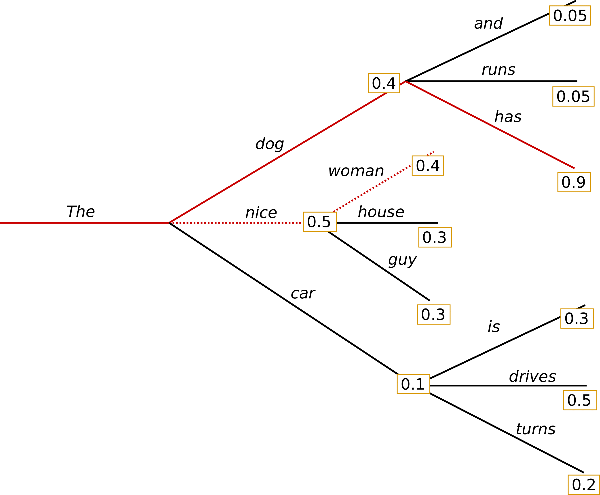
#
# At time step $1$, besides the most likely hypothesis $\text{"The", "woman"}$, beam search also keeps track of the second most likely one $\text{"The", "dog"}$. At time step $2$, beam search finds that the word sequence $\text{"The", "dog", "has"}$ has with $0.36$ a higher probability than $\text{"The", "nice", "woman"}$, which has $0.2$. Great, it has found the most likely word sequence in our toy example!
#
# Beam search will always find an output sequence with higher probability than greedy search, but is not guaranteed to find the most likely output.
#
# Let's see how beam search can be used in `transformers`. We set `num_beams > 1` and `early_stopping=True` so that generation is finished when all beam hypotheses reached the EOS token.
# + id="R1R5kx30Ynej" colab={"base_uri": "https://localhost:8080/"} outputId="d094dff7-8144-4db1-b4bc-db9ec12faac7"
# activate beam search and early_stopping
beam_output = model.generate(
input_ids,
max_length=50,
num_beams=5,
early_stopping=True
)
print("Output:\n" + 100 * '-')
print(tokenizer.decode(beam_output[0], skip_special_tokens=True))
# + [markdown] id="AZ6xs-KLi9jT"
# While the result is arguably more fluent, the output still includes repetitions of the same word sequences.
# A simple remedy is to introduce *n-grams* (*a.k.a* word sequences of $n$ words) penalties as introduced by [Paulus et al. (2017)](https://arxiv.org/abs/1705.04304) and [Klein et al. (2017)](https://arxiv.org/abs/1701.02810). The most common *n-grams* penalty makes sure that no *n-gram* appears twice by manually setting the probability of next words that could create an already seen *n-gram* to $0$.
#
# Let's try it out by setting `no_repeat_ngram_size=2` so that no *2-gram* appears twice:
# + id="jy3iVJgfnkMi" colab={"base_uri": "https://localhost:8080/"} outputId="7c0c4ff3-ac4b-4bad-e860-a8ab1116dde8"
# set no_repeat_ngram_size to 2
beam_output = model.generate(
input_ids,
max_length=max_len,
num_beams=5,
no_repeat_ngram_size=2,
early_stopping=True
)
print("Output:\n" + 100 * '-')
print(tokenizer.decode(beam_output[0], skip_special_tokens=True))
# + [markdown] id="nxsksOGDpmA0"
# Nice, that looks much better! We can see that the repetition does not appear anymore. Nevertheless, *n-gram* penalties have to be used with care. An article generated about the city *New York* should not use a *2-gram* penalty or otherwise, the name of the city would only appear once in the whole text!
#
# Another important feature about beam search is that we can compare the top beams after generation and choose the generated beam that fits our purpose best.
#
# In `transformers`, we simply set the parameter `num_return_sequences` to the number of highest scoring beams that should be returned. Make sure though that `num_return_sequences <= num_beams`!
# + id="5ClO3VphqGp6" colab={"base_uri": "https://localhost:8080/"} outputId="2f0d7262-d0ce-4c45-a1f6-790a73306ac5"
# set return_num_sequences > 1
beam_outputs = model.generate(
input_ids,
max_length=50,
num_beams=5,
no_repeat_ngram_size=2,
num_return_sequences=5,
early_stopping=True
)
# now we have 3 output sequences
print("Output:\n" + 100 * '-')
for i, beam_output in enumerate(beam_outputs):
print("{}: {}".format(i, tokenizer.decode(beam_output, skip_special_tokens=True)))
# + [markdown] id="HhLKyfdbsjXc"
# As can be seen, the five beam hypotheses are only marginally different to each other - which should not be too surprising when using only 5 beams.
#
# In open-ended generation, a couple of reasons have recently been brought forward why beam search might not be the best possible option:
#
# - Beam search can work very well in tasks where the length of the desired generation is more or less predictable as in machine translation or summarization - see [Murray et al. (2018)](https://arxiv.org/abs/1808.10006) and [Yang et al. (2018)](https://arxiv.org/abs/1808.09582). But this is not the case for open-ended generation where the desired output length can vary greatly, e.g. dialog and story generation.
#
# - We have seen that beam search heavily suffers from repetitive generation. This is especially hard to control with *n-gram*- or other penalties in story generation since finding a good trade-off between forced "no-repetition" and repeating cycles of identical *n-grams* requires a lot of finetuning.
#
# - As argued in [<NAME> et al. (2019)](https://arxiv.org/abs/1904.09751), high quality human language does not follow a distribution of high probability next words. In other words, as humans, we want generated text to surprise us and not to be boring/predictable. The authors show this nicely by plotting the probability, a model would give to human text vs. what beam search does.
#
# 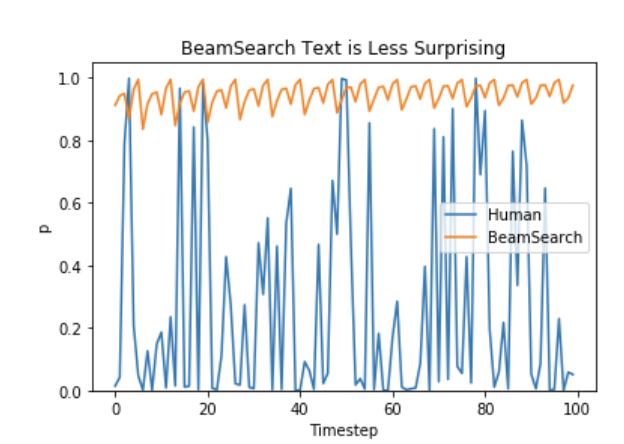
#
#
# So let's stop being boring and introduce some randomness 🤪.
# + [markdown] id="XbbIyK84wHq6"
# ### **Sampling**
#
# In its most basic form, sampling means randomly picking the next word $w_t$ according to its conditional probability distribution:
#
# $$w_t \sim P(w|w_{1:t-1})$$
#
# Taking the example from above, the following graphic visualizes language generation when sampling.
#
# 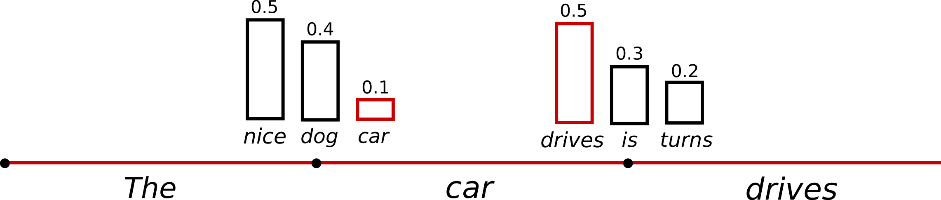
#
# It becomes obvious that language generation using sampling is not *deterministic* anymore. The word
# $\text{"car"}$ is sampled from the conditioned probability distribution $P(w | \text{"The"})$, followed by sampling $\text{"drives"}$ from $P(w | \text{"The"}, \text{"car"})$.
#
# In `transformers`, we set `do_sample=True` and deactivate *Top-K* sampling (more on this later) via `top_k=0`. In the following, we will fix `random_seed=0` for illustration purposes. Feel free to change the `random_seed` to play around with the model.
#
# + id="aRAz4D-Ks0_4" colab={"base_uri": "https://localhost:8080/"} outputId="7460d383-433b-419e-dd87-6b37d34c72f1"
# activate sampling and deactivate top_k by setting top_k sampling to 0
sample_output = model.generate(
input_ids,
do_sample=True,
max_length=max_len,
top_k=0
)
print("Output:\n" + 100 * '-')
print(tokenizer.decode(sample_output[0], skip_special_tokens=True))
# + [markdown] id="mQHuo911wfT-"
# Interesting! The text seems alright - but when taking a closer look, it is not very coherent. the *3-grams* *new hand sense* and *local batte harness* are very weird and don't sound like they were written by a human. That is the big problem when sampling word sequences: The models often generate incoherent gibberish, *cf.* [Ari Holtzman et al. (2019)](https://arxiv.org/abs/1904.09751).
#
# A trick is to make the distribution $P(w|w_{1:t-1})$ sharper (increasing the likelihood of high probability words and decreasing the likelihood of low probability words) by lowering the so-called `temperature` of the [softmax](https://en.wikipedia.org/wiki/Softmax_function#Smooth_arg_max).
#
# An illustration of applying temperature to our example from above could look as follows.
#
# 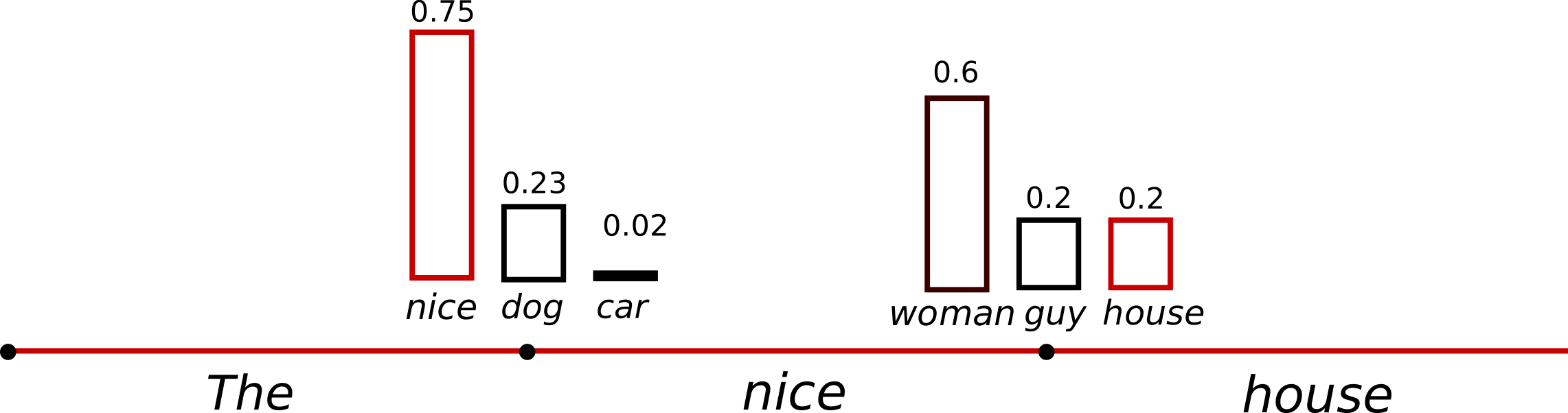
#
# The conditional next word distribution of step $t=1$ becomes much sharper leaving almost no chance for word $\text{"car"}$ to be selected.
#
#
# Let's see how we can cool down the distribution in the library by setting `temperature=0.7`:
# + id="WgJredc-0j0Z" colab={"base_uri": "https://localhost:8080/"} outputId="e28a5e9a-5814-405d-8038-b4cc09254712"
# use temperature to decrease the sensitivity to low probability candidates
sample_output = model.generate(
input_ids,
do_sample=True,
max_length=50,
top_k=50,
temperature=0.7
)
print("Output:\n" + 100 * '-')
print(tokenizer.decode(sample_output[0], skip_special_tokens=True))
# + [markdown] id="kzGuu24hZZnq"
# OK. There are less weird n-grams and the output is a bit more coherent now! While applying temperature can make a distribution less random, in its limit, when setting `temperature` $ \to 0$, temperature scaled sampling becomes equal to greedy decoding and will suffer from the same problems as before.
#
#
# + [markdown] id="binNTroyzQBu"
# ### **Top-K Sampling**
#
# [Fan et. al (2018)](https://arxiv.org/pdf/1805.04833.pdf) introduced a simple, but very powerful sampling scheme, called ***Top-K*** sampling. In *Top-K* sampling, the *K* most likely next words are filtered and the probability mass is redistributed among only those *K* next words.
# GPT2 adopted this sampling scheme, which was one of the reasons for its success in story generation.
#
# We extend the range of words used for both sampling steps in the example above from 3 words to 10 words to better illustrate *Top-K* sampling.
#
# 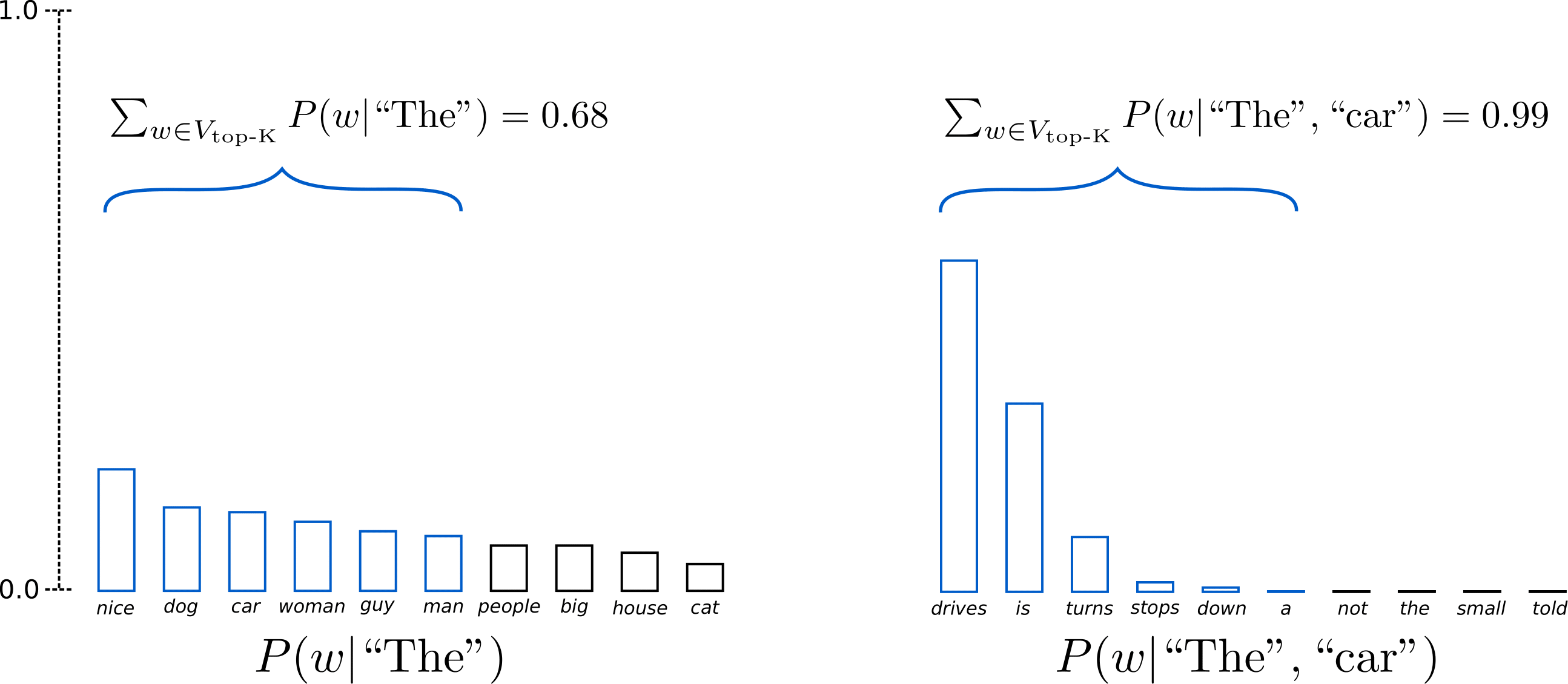
#
# Having set $K = 6$, in both sampling steps we limit our sampling pool to 6 words. While the 6 most likely words, defined as $V_{\text{top-K}}$ encompass only *ca.* two-thirds of the whole probability mass in the first step, it includes almost all of the probability mass in the second step. Nevertheless, we see that it successfully eliminates the rather weird candidates $\text{"not", "the", "small", "told"}$
# in the second sampling step.
#
#
# Let's see how *Top-K* can be used in the library by setting `top_k=50`:
# + id="HBtDOdD0wx3l" colab={"base_uri": "https://localhost:8080/"} outputId="3529ee04-dd1e-4013-a9d3-7e205a6e48d8"
# set top_k to 50
sample_output = model.generate(
input_ids,
do_sample=True,
max_length=max_len,
top_k=50
)
print("Output:\n" + 100 * '-')
print(tokenizer.decode(sample_output[0], skip_special_tokens=True))
# + [markdown] id="Y77H5m4ZmhEX"
# Not bad at all! The text is arguably the most *human-sounding* text so far.
# One concern though with *Top-K* sampling is that it does not dynamically adapt the number of words that are filtered from the next word probability distribution $P(w|w_{1:t-1})$.
# This can be problematic as some words might be sampled from a very sharp distribution (distribution on the right in the graph above), whereas others from a much more flat distribution (distribution on the left in the graph above).
#
# In step $t=1$, *Top-K* eliminates the possibility to
# sample $\text{"people", "big", "house", "cat"}$, which seem like reasonable candidates. On the other hand, in step $t=2$ the method includes the arguably ill-fitted words $\text{"down", "a"}$ in the sample pool of words. Thus, limiting the sample pool to a fixed size *K* could endanger the model to produce gibberish for sharp distributions and limit the model's creativity for flat distribution.
# This intuition led [<NAME> et al. (2019)](https://arxiv.org/abs/1904.09751) to create ***Top-p***- or ***nucleus***-sampling.
#
#
# + [markdown] id="ki9LAaexzV3H"
# ### **Top-p (nucleus) sampling**
#
# Instead of sampling only from the most likely *K* words, in *Top-p* sampling chooses from the smallest possible set of words whose cumulative probability exceeds the probability *p*. The probability mass is then redistributed among this set of words. This way, the size of the set of words (*a.k.a* the number of words in the set) can dynamically increase and decrease according to the next word's probability distribution. Ok, that was very wordy, let's visualize.
#
# 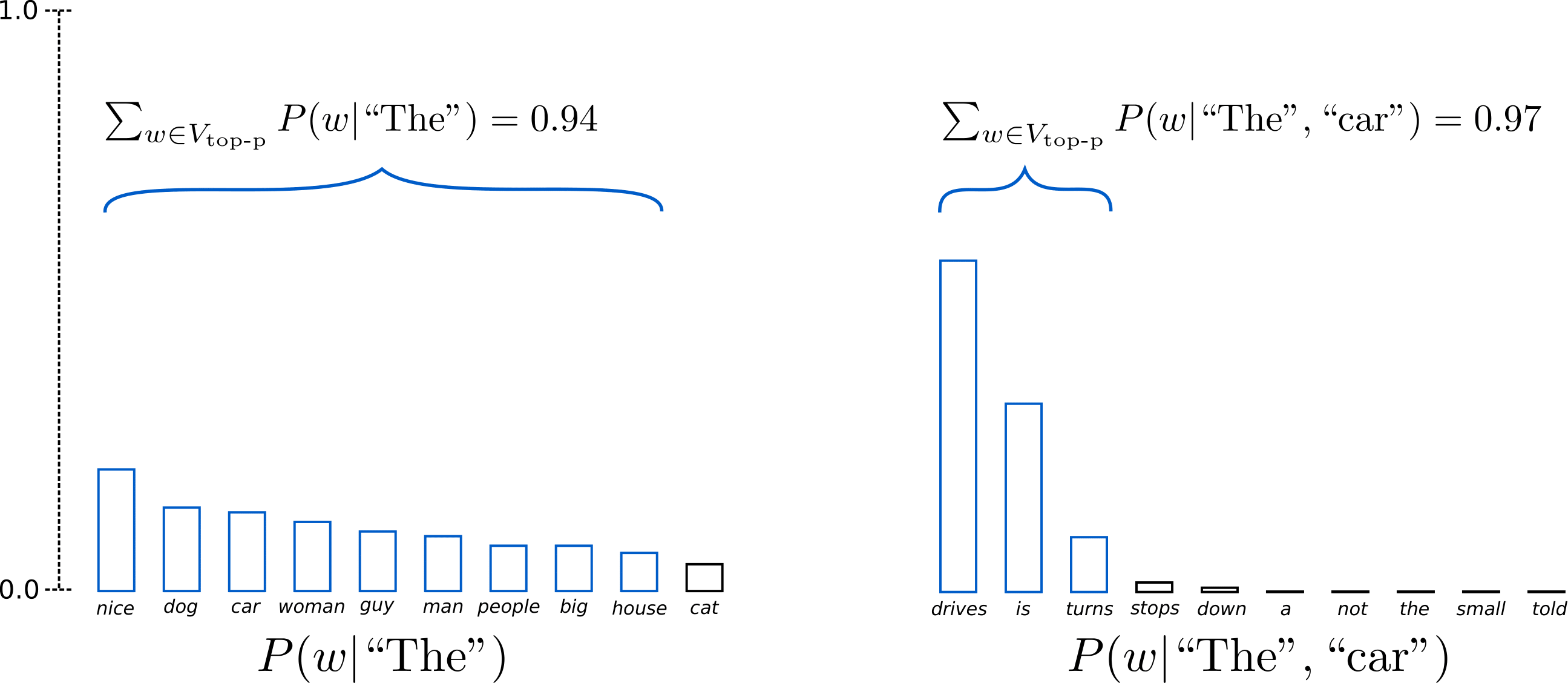
#
# Having set $p=0.92$, *Top-p* sampling picks the *minimum* number of words to exceed together $p=92\%$ of the probability mass, defined as $V_{\text{top-p}}$. In the first example, this included the 9 most likely words, whereas it only has to pick the top 3 words in the second example to exceed 92%. Quite simple actually! It can be seen that it keeps a wide range of words where the next word is arguably less predictable, *e.g.* $P(w | \text{"The"})$, and only a few words when the next word seems more predictable, *e.g.* $P(w | \text{"The", "car"})$.
#
# Alright, time to check it out in `transformers`!
# We activate *Top-p* sampling by setting `0 < top_p < 1`:
# + id="EvwIc7YAx77F" colab={"base_uri": "https://localhost:8080/"} outputId="2b4a4148-7382-4ebd-8300-b7adba4f9b36"
# deactivate top_k sampling and sample only from 92% most likely words
sample_output = model.generate(
input_ids,
do_sample=True,
max_length=max_len,
top_p=0.92,
top_k=0
)
print("Output:\n" + 100 * '-')
print(tokenizer.decode(sample_output[0], skip_special_tokens=True))
# + [markdown] id="tn-8gLaR4lat"
# Great, that sounds like it could have been written by a human. Well, maybe not quite yet.
#
# While in theory, *Top-p* seems more elegant than *Top-K*, both methods work well in practice. *Top-p* can also be used in combination with *Top-K*, which can avoid very low ranked words while allowing for some dynamic selection.
#
# Finally, to get multiple independently sampled outputs, we can *again* set the parameter `num_return_sequences > 1`:
# + id="3kY8P9VG8Gi9" colab={"base_uri": "https://localhost:8080/"} outputId="d19ef270-77d2-4d8b-d6d6-7cecfb3b7603"
# set top_k = 50 and set top_p = 0.95 and num_return_sequences = 3
sample_outputs = model.generate(
input_ids,
do_sample=True,
max_length=max_len,
top_k=50,
top_p=0.95,
num_return_sequences=3
)
print("Output:\n" + 100 * '-')
for i, sample_output in enumerate(sample_outputs):
print("{}: {}".format(i, tokenizer.decode(sample_output, skip_special_tokens=True)))
# + [markdown] id="-vRPfMl88rk0"
# Cool, now you should have all the tools to let your model write your stories with `transformers`!
# + [markdown] id="NsWd7e98Vcs3"
# ### **Conclusion**
#
# As *ad-hoc* decoding methods, *top-p* and *top-K* sampling seem to produce more fluent text than traditional *greedy* - and *beam* search on open-ended language generation.
# Recently, there has been more evidence though that the apparent flaws of *greedy* and *beam* search - mainly generating repetitive word sequences - are caused by the model (especially the way the model is trained), rather than the decoding method, *cf.* [Welleck et al. (2019)](https://arxiv.org/pdf/1908.04319.pdf). Also, as demonstrated in [Welleck et al. (2020)](https://arxiv.org/abs/2002.02492), it looks as *top-K* and *top-p* sampling also suffer from generating repetitive word sequences.
#
# In [Welleck et al. (2019)](https://arxiv.org/pdf/1908.04319.pdf), the authors show that according to human evaluations, *beam* search can generate more fluent text than *Top-p* sampling, when adapting the model's training objective.
#
# Open-ended language generation is a rapidly evolving field of research and as it is often the case there is no one-size-fits-all method here, so one has to see what works best in one's specific use case.
#
# Good thing, that *you* can try out all the different decoding methods in `transfomers` 🤗.
#
# That was a short introduction on how to use different decoding methods in `transformers` and recent trends in open-ended language generation.
#
# Feedback and questions are very welcome on the [Github repository](https://github.com/huggingface/transformers).
#
# For more fun generating stories, please take a look at [Writing with Transformers](https://transformer.huggingface.co).
#
# Thanks to everybody, who has contributed to the blog post: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME> and <NAME>.
#
# + [markdown] id="w4CYi91h11yd"
# ### **Appendix**
#
# There are a couple of additional parameters for the `generate` method that were not mentioned above. We will explain them here briefly!
#
# - `min_length` can be used to force the model to not produce an EOS token (= not finish the sentence) before `min_length` is reached. This is used quite frequently in summarization, but can be useful in general if the user wants to have longer outputs.
# - `repetition_penalty` can be used to penalize words that were already generated or belong to the context. It was first introduced by [Kesker et al. (2019)](https://arxiv.org/abs/1909.05858) and is also used in the training objective in [Welleck et al. (2019)](https://arxiv.org/pdf/1908.04319.pdf). It can be quite effective at preventing repetitions, but seems to be very sensitive to different models and use cases, *e.g.* see this [discussion](https://github.com/huggingface/transformers/pull/2303) on Github.
#
# - `attention_mask` can be used to mask padded tokens
# - `pad_token_id`, `bos_token_id`, `eos_token_id`: If the model does not have those tokens by default, the user can manually choose other token ids to represent them.
#
# For more information please also look into the `generate` function [docstring](https://huggingface.co/transformers/main_classes/model.html?highlight=generate#transformers.TFPreTrainedModel.generate).
|
Norod78_hebrew_gpt_neo_small.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scene Classification
# ## 1. Preprocess
# - Import pkg
# - Extract zip file
# - Preview "scene_classes.csv"
# - Preview "scene_{0}_annotations_20170904.json"
# - Test the image and pickle function
# - Split data into serval pickle file
# This part need jupyter notebook start with "jupyter notebook --NotebookApp.iopub_data_rate_limit=1000000000" (https://github.com/jupyter/notebook/issues/2287)
#
# Reference:
# - https://challenger.ai/competitions
# - https://github.com/jupyter/notebook/issues/2287
# ### Import pkg
import numpy as np
import pandas as pd
# import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import seaborn as sns
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, BatchNormalization
from keras.optimizers import Adam
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import LearningRateScheduler, TensorBoard
# import zipfile
import os
import zipfile
import math
from time import time
from IPython.display import display
import pdb
import json
from PIL import Image
import glob
import pickle
# ### Extract zip file
# +
input_path = './input'
datasetName = 'train'
date = '20170904'
zip_path = input_path + '/ai_challenger_scene_{0}_{1}.zip'.format(datasetName, date)
extract_path = input_path + '/ai_challenger_scene_{0}_{1}'.format(datasetName, date)
image_path = extract_path + '/scene_{0}_images_{1}'.format(datasetName, date)
scene_classes_path = extract_path + '/scene_classes.csv'
scene_annotations_path = extract_path + '/scene_{0}_annotations_{1}.json'.format(datasetName, date)
print(input_path)
print(zip_path)
print(extract_path)
print(image_path)
print(scene_classes_path)
print(scene_annotations_path)
# -
if not os.path.isdir(extract_path):
with zipfile.ZipFile(zip_path) as file:
for name in file.namelist():
file.extract(name, input_path)
# ### Preview "scene_classes.csv"
scene_classes = pd.read_csv(scene_classes_path, header=None)
display(scene_classes.head())
# ### Preview "scene_{0}_annotations_20170904.json"
#
# **This part need jupyter notebook start with "jupyter notebook --NotebookApp.iopub_data_rate_limit=1000000000"**
# https://github.com/jupyter/notebook/issues/2287
with open(scene_annotations_path, 'r', encoding='utf-8') as file:
content = ''
for line in file:
content = content + line
scene_annotations = json.loads(content)
#We get a list
print('scene_{0}_annotations.type: %{1}'.format(datasetName, type(scene_annotations)))
print('scene_{0}_annotations.shape: %{1}'.format(datasetName, len(scene_annotations)))
order = 0
print(scene_annotations[order])
print('label_id[{0}]:\t{1}'.format(order, scene_annotations[order]['label_id']))
print('image_id[{0}]:\t{1}'.format(order, scene_annotations[order]['image_id']))
# ### Test the image and pickle function
# +
# length = len(scene_annotations)
length = 2
box = (224, 224)
x_shape = [length, box[0], box[1], 3]
x_data = np.zeros(x_shape)
y_data = np.zeros(length)
fig, ax = plt.subplots(3, length, figsize=(12, 12))
for i in range(length):
y_data[i] = scene_annotations[i]['label_id']
path = image_path + '/' + scene_annotations[i]['image_id']
img = Image.open(path)
img1 = img.resize(box, Image.ANTIALIAS) # resizes image in-place
imgData = np.asarray(img1)
ax[0][i].imshow(imgData)
imgData = imgData.astype("float32")
imgData = imgData/255.0
ax[1][i].imshow(imgData)
x_data[i] = imgData
ax[2][i].imshow(x_data[i])
print('Data save into pickle file:')
print(y_data)
print(x_data.shape)
# print(x_data[0])
pickleFolder = 'pickle_{0}'.format(datasetName)
pickle_path = input_path + '/' + pickleFolder
if not os.path.isdir(pickle_path):
os.mkdir(pickle_path)
x_data_path = pickle_path + '/x_data_sample.p'
y_data_path = pickle_path + '/y_data_sample.p'
pickle.dump(x_data, open(x_data_path, 'wb'))
pickle.dump(y_data, open(y_data_path, 'wb'))
x_data = pickle.load(open(x_data_path, mode='rb'))
y_data = pickle.load(open(y_data_path, mode='rb'))
print('Load data from pickle file:')
print(y_data_path)
print(x_data_path)
print(y_data)
print(x_data.shape)
# -
# ### Split data into serval pickle file
def convert_and_save_data(partNum, partLen, thisPartLen):
x_shape = [partLen, box[0], box[1], 3]
x_data = np.zeros(x_shape)
y_data = np.zeros(partLen)
# fig, ax = plt.subplots(3, length, figsize=(12, 12))
for i in range(partLen):
y_data[i] = scene_annotations[i]['label_id']
path = image_path + '/' + scene_annotations[i]['image_id']
img = Image.open(path)
img1 = img.resize(box, Image.ANTIALIAS) # resizes image in-place
imgData = np.asarray(img1)
# ax[0][i].imshow(imgData)
imgData = imgData.astype("float32")
imgData = imgData/255.0
# ax[1][i].imshow(imgData)
x_data[i] = imgData
# ax[2][i].imshow(x_data[i])
print('Data save into pickle file:')
print(y_data.shape)
print(x_data.shape)
# print(x_data[0])
pickleFolder = 'pickle_{0}'.format(datasetName)
pickle_path = input_path + '/' + pickleFolder
if not os.path.isdir(pickle_path):
os.mkdir(pickle_path)
x_data_path = pickle_path + '/x_data' + str(partNum) + '.p'
y_data_path = pickle_path + '/y_data' + str(partNum) + '.p'
pickle.dump(x_data, open(x_data_path, 'wb'))
pickle.dump(y_data, open(y_data_path, 'wb'))
x_data = pickle.load(open(x_data_path, mode='rb'))
y_data = pickle.load(open(y_data_path, mode='rb'))
print('Load data from pickle file:')
print(y_data.shape)
print(x_data.shape)
# +
length = len(scene_annotations)
# length = 25
partLen = 1000
partAmount = math.ceil(length/partLen)
print('data length:\t%s' %length)
print('partLen:\t%s' %partLen)
print('partAmount:\t%s' %partAmount)
box = (224, 224)
print('image box:\t%sx%s' %box)
for i in range(partAmount):
remainder = length - i*partLen
if remainder < partLen:
thisPartLen = remainder
else:
thisPartLen = partLen
# x_shape = [thisPartLen, box[0], box[1], 3]
# x_data = np.zeros(x_shape)
# y_data = np.zeros(thisPartLen)
print('thisPartLen {0}:\t{1}\t' %(i, thisPartLen))
convert_and_save_data(i, partLen, thisPartLen)
# -
print('Done!')
|
SceneClassification2017/backup/doc20170923/1. Preprocess.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ##I am splitting v3_Clean_model_add_Pis_feat.ipynb into 2 notebooks
#
# -this one on modelling and
#
# -another one on creating stats data set (just Pisch for now)
# v1_stats_tools_Pish.ipynb
# +
##part 1
##use models with default
##use data set with H/A +1, -1
##do full window for now
##next:
##check if just excluding first 10 days helps (chaotic)
##check if different windows help
##next
## can try tuning (for loops by hand, or ... use grid_search (use ML mastery code))
##-I think tuning will be faster ... just do by hand ... loop over the possible things
##-ONE for loop over i = (a,b,c,d)... for each model i[0]
##Orrr can try adding features ... here we have to worry about:
##-adding basic features eg pp, and correct fo%
##-scaling numericals
##-dummy vars for categoricals (are there any?) besides H/A
##-num_windows and which lengths for moving avgs
##-filtering the features for increasing complexity inteligently
##-There is a dicotemy:
##(a)use H/A + numerics or ... here I think it can be made more like time-series
##(b) just use mumerics (moving avg) ... here I think the order of the games is not important (note Leung did this, and random train)
# +
import pandas as pd
import numpy as np
from sklearn.metrics import accuracy_score, precision_score, recall_score, roc_auc_score, confusion_matrix
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score, f1_score
import matplotlib.pyplot as plt
import seaborn as sns
##couple evaluation functions ##removed model_name as variable
def evaluate_binary_classification(y_test, y_pred, y_proba=None, graph = False):
accuracy = accuracy_score(y_test, y_pred)
precision = precision_score(y_test, y_pred)
recall = recall_score(y_test, y_pred)
f1 = f1_score(y_test, y_pred)
#try:
if y_proba != None:
rocauc_score = roc_auc_score(y_test, y_proba)
else:
rocauc_score = "no roc"
#except:
# pass
cm = confusion_matrix(y_test, y_pred)
if graph == True:
sns.heatmap(cm, annot=True)
plt.tight_layout()
plt.title(f'{model_name}', y=1.1)
plt.ylabel('Actual label')
plt.xlabel('Predicted label')
plt.show()
print("accuracy: ", accuracy)
print("precision: ", precision)
print("recall: ", recall)
print("f1 score: ", f1)
print("rocauc: ", rocauc_score)
print(cm)
#return accuracy, precision, recall, f1, rocauc_score
def evaluate_regression(y_test, y_pred):
mae = mean_absolute_error(y_test, y_pred)
mse = mean_squared_error(y_test, y_pred)
r2 = r2_score(y_test, y_pred)
print("mae", mae)
print("mse", mse)
print('r2', r2)
##display null values
def perc_null(X):
total = X.isnull().sum().sort_values(ascending=False)
data_types = X.dtypes
percent = (X.isnull().sum()/X.isnull().count()).sort_values(ascending=False)
missing_data = pd.concat([total, data_types, percent], axis=1, keys=['Total','Type' ,'Percent'])
return missing_data
# +
#this takes the odds eg -200 is the favorite, 140 is underdog and says fav wins
def fav_win(x):
if x <=0:
return 1
if x>0:
return 0
v_fav_win = np.vectorize(fav_win)
def make_win(x):
if x <= 0:
return 0
if x >0:
return 1
v_make_win = np.vectorize(make_win)
# +
def regr_model_results(model, model_name, X, dates, step, window_size, prediction_size, drop_first_k_days = 0): #X = data
results_dic['model_name'] = []
results_dic['date'] = []
results_dic['mae'] = []
results_dic['mse'] = []
results_dic['r2'] = []
#drop first k days from dates and X
dates = dates[drop_first_k_days :]
X = X.loc[X['full_date'].isin(dates), :].copy()
for i in range(step, len(dates), step): ##eg step =10, so 17 rounds
model.fit(X.loc[X['full_date'].isin(dates[max(i-window_size ,0):i]), :],y.loc[y['full_date'].isin(dates[max(i-window_size,0):i]),'goal_difference' ])
y_pred = model.predict(X.loc[X['full_date'].isin(dates[i:i+prediction_size]), :])
y_test = y.loc[y['full_date'].isin(dates[i:i+prediction_size]),'goal_difference' ]
mae = mean_absolute_error(y_test, y_pred)
mse = mean_squared_error(y_test, y_pred)
r2 = r2_score(y_test, y_pred)
results_dic['model_name'].append(model_name)
results_dic['date'].append(dates[i])
results_dic['mae'].append(mae)
results_dic['mse'].append(mse)
results_dic['r2'].append(r2)
return results_dic #!
# +
def class_model_results(model, model_name, X, dates, step, window_size, prediction_size, drop_first_k_days = 0): #X = data
results_dic ={}
results_dic['model_name'] = []
results_dic['date'] = []
results_dic['accuracy'] = []
results_dic['f1_score'] = []
#results_dic['precision'] = []
# results_dic['recall'] = []
#drop first k days from dates and X
dates = dates[drop_first_k_days :]
X = X.loc[X['full_date'].isin(dates), :].copy()
for i in range(step, len(dates), step): ##eg step =10, so 17 rounds
model.fit(X.loc[X['full_date'].isin(dates[max(i-window_size ,0):i]), :],y.loc[y['full_date'].isin(dates[max(i-window_size,0):i]),'won' ])
y_pred = model.predict(X.loc[X['full_date'].isin(dates[i:i+prediction_size]), :])
y_test = y.loc[y['full_date'].isin(dates[i:i+prediction_size]),'won' ]
accuracy = accuracy_score(y_test, y_pred)
#recision = precision_score(y_test, y_pred, zero_division = 0)
#recall = recall_score(y_test, y_pred)
f1 = f1_score(y_test, y_pred) #, average = None)
results_dic['model_name'].append(model_name) #append same model name every iter so same length as others
results_dic['date'].append(dates[i])
results_dic['accuracy'].append(accuracy)
results_dic['f1_score'].append(f1)
#results_dic['precision'].append(precision)
#results_dic['recall'].append(recall)
results_dic['model_name'].append('model_name'+'_avg') #append same model name every iter so same length as others
results_dic['date'].append('average')
results_dic['accuracy'].append(round(np.mean(np.array(results_dic['accuracy'])), 2) )
results_dic['f1_score'].append(round(np.mean(np.array(results_dic['f1_score'])), 2) )
return results_dic #!
# +
##note KNN or other clusters might be helpful group the teams in smart way ... but not now.
#models
##regression
from sklearn.linear_model import Ridge
from sklearn.ensemble import RandomForestRegressor
#classifiers (non-tree)
from sklearn.linear_model import RidgeClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression, SGDRegressor, SGDClassifier
from sklearn.svm import SVC
#tree-based classifiers
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import BaggingClassifier
from sklearn.ensemble import GradientBoostingClassifier
from xgboost import XGBClassifier
from xgboost import XGBRegressor
##regression models
lr = Ridge(alpha=0.001)
rfr = RandomForestRegressor(max_depth=3, random_state=0)
xgbr = XGBRegressor()
##classifier models
lrc = RidgeClassifier()
gnb = GaussianNB()
lgr = LogisticRegression(random_state = 0)
svc = SVC()
#tree-based classifiers
rfc = RandomForestClassifier(max_depth=3, random_state=0)
bc = BaggingClassifier()
gbc = GradientBoostingClassifier()
xgbc = XGBClassifier()
# -
# ##TUNING INFO
#
#
# ##hyper_parameters from here
# ##https://machinelearningmastery.com/hyperparameters-for-classification-machine-learning-algorithms/
# ##for xgboost from here
# ##https://machinelearningmastery.com/extreme-gradient-boosting-ensemble-in-python/
#
# #xgb
#
# trees = [10, 50, 100, 500, 1000, 5000] #100 #num of trees
# max_depth = range(1,11) ##3-5
# rates = [0.0001, 0.001, 0.01, 0.1, 1.0] #0.1
# subsample in arange(0.1, 1.1, 0.1): #0.4, 0.5 ##this is 0.1, 0.2 ... 1.0 # % of features to sample
#
#
# #svc
# kernels in [‘linear’, ‘poly’, ‘rbf’, ‘sigmoid’] #if you use poly, then adjust degree
# C in [100, 10, 1.0, 0.1, 0.001]
#
# #gb
#
# learning_rate in [0.001, 0.01, 0.1]
# n_estimators [10, 100, 1000]
# subsample in [0.5, 0.7, 1.0]
# max_depth in [3, 7, 9]
#
#
# #rfc
# max_features [1 to 20] #key
# max_features in [‘sqrt’, ‘log2’]
# n_estimators in [10, 100, 1000]
#
# #bc
# n_estimators in [10, 100, 1000]
#
# svm_dic = {'kernels':[‘linear’, ‘poly’, ‘rbf’, ‘sigmoid’]}
# lrc_dic = {'alpha': [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0]}
# lgr_hp_dic = {'solver': [‘newton-cg’, ‘lbfgs’, ‘liblinear’, ‘sag’, ‘saga’], 'penalty' : [‘none’, ‘l1’, ‘l2’, ‘elasticnet’],
# 'C' :[100, 10, 1.0, 0.1, 0.01]}
# +
##classifier models
lrc = RidgeClassifier()
gnb = GaussianNB()
lgr = LogisticRegression(random_state = 0)
svc = SVC(kernel = 'rbf')
xgbr = XGBRegressor()
#tree-based classifiers
rfc = RandomForestClassifier(max_depth=5, random_state=0)
bc = BaggingClassifier()
gbc = GradientBoostingClassifier()
xgbc = XGBClassifier()
# +
data_dic = {}
file_path_12 = '/Users/joejohns/data_bootcamp/GitHub/final_project_nhl_prediction/Note_books/Explore_Models/data_dummies_Pis_v2_20122013.csv'
data_dic[20122013] = pd.read_csv(file_path_12)
for season in [20152016, 20162017, 20172018, 20182019]:
file_path_seas = '/Users/joejohns/data_bootcamp/GitHub/final_project_nhl_prediction/Note_books/Explore_Models/'+'data_dummies_Pis_xg_Corsi_v3_'+str(season)+'.csv'
data_dic[season] = pd.read_csv(file_path_seas)
# data_bootcamp/GitHub/final_project_nhl_prediction/Note_books/Explore_Models/data_dummies_Pis_v2_20122013.csv
#data_bootcamp/GitHub/final_project_nhl_prediction/Note_books/Explore_Models/data_dummies_Pis_xg_Corsi_v3_20152016.csv
# +
k =5
for season in [20122013,20152016, 20162017, 20172018, 20182019]:
filter = (data_dic[season]['full_date'] >= data_dic[season]['full_date'][0]+k).copy() #removes first k = 5 days of season where there are nan values
data_dic[season]= data_dic[season].loc[filter, :].copy()
# -
data_12 = data_dic[20122013].copy()
data_15 = data_dic[20152016].copy()
data_16 = data_dic[20162017].copy()
data_17 = data_dic[20172018].copy()
data_18 = data_dic[20182019].copy()
data_15_17 = pd.concat([data_15,data_16])
data_17_19 = pd.concat([data_17,data_18])
#Note Bene
data_12.rename(columns ={'win%':'win%_cumul'}, inplace = True)
data_12.rename(columns ={'last_10_games_win%' :'win%_last_10_games'}, inplace = True)
#(1230, 50)
#(1230, 50)
#(1271, 51) Vegas, baby
#(1271, 51)'win%_last_10_games'
# +
#perc_null(data_18) takes 5 days to get rid of NaN ... also good to get rid of randomness in first 5 days (maybe even 20 days ...)
# +
#data_12.iloc[:10,7:]
##columns are all safe ...
#win%_cumul HAS to be previous day (NOT including day of ... o/w model can inspect
#which teams win% went up and which ... actually kinda tough bec it's difference )
##anyway I checked in v1_stats 'SJS' ... that win% = win%_cumul is *strictly* the previous days
# -
columns_to_scale = list(data_15.iloc[:5, 38:].columns)
data_15.iloc[:5, 38:].columns
columns_target = list(data_15.iloc[:5, :8].columns)
data_15.iloc[:5, :8].columns
# +
X_12 = data_12.iloc[:,7:].copy()
X_15 = data_15.drop(columns = columns_target ).copy()
X_16 = data_16.drop(columns = columns_target ).copy()
X_17 = data_17.drop(columns = columns_target ).copy()
X_18 = data_18.drop(columns = columns_target ).copy()
y_12 = data_12.iloc[:,:7].copy()
y_15 = data_15.loc[:, columns_target ].copy()
y_16 = data_16.loc[:, columns_target ].copy()
y_17 = data_17.loc[:, columns_target ].copy()
y_18 = data_18.loc[:, columns_target ].copy()
list_X = [X_12, X_15,X_16, X_17, X_18]
list_y = [y_12, y_15, y_16,y_17, y_18 ]
list_Xy = zip(list_X, list_y)
list_data = [data_12, data_15, data_16, data_17, data_18]
# -
# End of baselines ... on to modelling ...
# +
##regression models
lr = Ridge(alpha=0.001)
rfr = RandomForestRegressor(max_depth=3, random_state=0)
xgbr = XGBRegressor()
sgdr = SGDRegressor() #has partial_fit()
##classifier models
lrc = RidgeClassifier()
gnb = GaussianNB() #has partial_fit()
lgr = LogisticRegression(random_state = 0, max_iter = 10**5)
svc = SVC()
sgdc = SGDClassifier() #has partial_fit()
#tree-based classifiers
rfc = RandomForestClassifier(max_depth=3, random_state=0)
bc = BaggingClassifier()
gbc = GradientBoostingClassifier()
xgbc = XGBClassifier(use_label_encoder=False)
##have partial_fit()
#['BernoulliNB', 'GaussianNB', 'MiniBatchKMeans', 'MultinomialNB', 'PassiveAggressiveClassifier', PassiveAggressiveRegressor', 'Perceptron', 'SGDClassifier', 'SGDRegressor']
# +
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler, MinMaxScaler
std_scal = StandardScaler()
mm_scal = MinMaxScaler()
# -
#X_15 as test case
X_15.shape
columns_to_scale = list(data_15.iloc[:5, 38:].columns)
data_15.iloc[:5, 38:].columns
columns_target = list(data_15.iloc[:5, :8].columns)
data_15.iloc[:5, :8].columns
# +
#experimenting with std_scaler ...
#NOte Bene ... you must assign the numpy object std_scal.fit_transform(Y.loc[:,columns_to_scale]).copy()
#to the df ... if you assign pd.DataFrame(same) it will give NaNs (why?)
Y = X_12.iloc[:300, :].copy()
#Y.loc[:,columns_to_scale] = pd.DataFrame(std_scal.fit_transform(Y.loc[:,columns_to_scale])).copy()
Y.loc[:,columns_to_scale] = std_scal.fit_transform(Y.loc[:,columns_to_scale]).copy()
Y.loc[:,columns_to_scale]
Y2 = X_12.iloc[300:, :].copy()
Y2.loc[:,columns_to_scale] = std_scal.transform(Y2.loc[:,columns_to_scale])
#X_12.iloc[300:, :]
#are these two scalars independent? yes ... but this doesn't work if you do std_scal = Sta ... and use that ... bec std_scal is one instance
scal = StandardScaler()
scal_T = StandardScaler()
Y = X_12.iloc[:300, :].copy()
Y_T = X_12.iloc[300:, :].copy()
#Y.loc[:,columns_to_scale] = pd.DataFrame(std_scal.fit_transform(Y.loc[:,columns_to_scale])).copy()
#scal.fitY.loc[:,columns_to_scale])
scal.fit(Y.loc[:,columns_to_scale])
scal_T.fit(Y_T.loc[:,columns_to_scale])
scal.transform(Y.loc[:,columns_to_scale])
# + active=""
# str(lrc)
# +
T = 800 #train until this game in season (out of 1200)
d = 100 #predict this many games
scal = std_scal
#X_15 has about 1200 games
#set the data frame and target
X = X_15.copy()
y = y_15.loc[:, 'won'].copy()
#how does it predict on next season?? Terrible! lol ... train around 80% test 47%
#W = X_16.copy()
#z = y_16.loc[:, 'won'].copy()
for model in [lrc, gnb, lgr, svc, rfc, bc, gbc, xgbc]:
y_train= y.iloc[:T].copy()
y_test = y.iloc[T:T+d].copy()
#y_test = z.iloc[:100].copy()
X_train = X.iloc[:T, :].copy()
X_test= X.iloc[T:T+d, :].copy()
#X_test = W.iloc[:100].copy()
#do standard/minmax scaling on X_train numeric columns ... better to do pipeline?
#X_train.loc[:, columns_to_scale] = scal.fit_transform(X_train.loc[:, columns_to_scale]).copy()
#fit the scaler from train portion to the test portion
#X_test.loc[:, columns_to_scale] = scal.transform(X_test.loc[:, columns_to_scale]).copy()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
acc = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test,y_pred)
print(model, ' TEST error ', acc,)# f1)
y_pred_train_err = model.predict(X_train) #! careful with this code
f1_train_err = f1_score(y_train,y_pred_train_err)
acc_train_err = accuracy_score(y_train, y_pred_train_err)
#print(' training error ', model, acc_train_err, ) #f1_train_err)
#I am following this stackexchange
#from sklearn.preprocessing import MinMaxScaler
#In [93]: mms = MinMaxScaler()
#In [94]: df[['x','z']] = mms.fit_transform(df[['x','z']])
#the one with check mark does pipe line tho :-) https://stackoverflow.com/questions/43834242
#ugghhh! terrible ...
# +
#ok pretty pretty good start ... logistic and ridgec 0.5835. (they are identical .. )
# -
X.shape
# +
#T = 800 #train until this game in season (out of 1200)
d = 20 #predict this many games
scal = StandardScaler()
#X_15 has about 1200 games
#set the data frame and target
X = X_15.copy()
y = y_15.loc[:, 'won'].copy()
model = gnb
#model = sgdc
##quick checks
counter = 0
acc_sum_train = 0
acc_sum_test = 0
#for model in [gnb, sgdc]: #partial_fit
for T in range(d, 800,d):
y_train= y.iloc[:T].copy()
y_test = y.iloc[T:T+d].copy()
#y_test = z.iloc[:100].copy()
X_train = X.iloc[:T, :].copy()
X_test= X.iloc[T:T+d, :].copy()
#X_test = W.iloc[:100].copy()
#do standard/minmax scaling on X_train numeric columns ... better to do pipeline?
X_train.loc[:, columns_to_scale] = scal.fit_transform(X_train.loc[:, columns_to_scale]).copy()
#fit the scaler from train portion to the test portion
X_test.loc[:, columns_to_scale] = scal.transform(X_test.loc[:, columns_to_scale]).copy()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
acc = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test,y_pred)
y_pred_train_err = model.predict(X_train) #! careful with this code
f1_train_err = f1_score(y_train,y_pred_train_err)
acc_train_err = accuracy_score(y_train, y_pred_train_err)
#print(' training error ', model, acc_train_err, ) #f1_train_err)
#print(model, ' train error ', acc_train_err, ' f1_train ', f1_train_err, ' TEST ERROR ', acc, ' f1 ', f1)
acc_sum_test += acc
acc_sum_train += acc_train_err
counter +=1
avg_acc = acc_sum_test/counter
avg_acc_train = acc_sum_train/counter
print(model, 'NOO partial_fit' ' avg train error ', avg_acc_train, ' AVERAGE TEST ERROR ', avg_acc,)
# -
dic = {}
# +
####
D = 1020 #train until this game in season (out of 1200)
d = 20 #predict this many games
scal = StandardScaler()
#X_15 has about 1200 games
#set the data frame and target
X = X_15.copy()
y = y_15.loc[:, 'won'].copy()
model = sgdc
#model = sgdc
##quick checks
counter = 0
acc_sum_train_np = 0
acc_sum_test_np = 0
#for model in [gnb, sgdc]: #partial_fit
for T in range(d, D,d):
y_train= y.iloc[:T].copy()
y_test = y.iloc[T:T+d].copy()
#y_test = z.iloc[:100].copy()
X_train = X.iloc[:T, :].copy()
X_test= X.iloc[T:T+d, :].copy()
#X_test = W.iloc[:100].copy()
#do standard/minmax scaling on X_train numeric columns ... better to do pipeline?
X_train.loc[:, columns_to_scale] = scal.fit_transform(X_train.loc[:, columns_to_scale]).copy()
#fit the scaler from train portion to the test portion
X_test.loc[:, columns_to_scale] = scal.transform(X_test.loc[:, columns_to_scale]).copy()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
acc = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test,y_pred)
y_pred_train_err = model.predict(X_train) #! careful with this code
f1_train_err = f1_score(y_train,y_pred_train_err)
acc_train_err = accuracy_score(y_train, y_pred_train_err)
#print(' training error ', model, acc_train_err, ) #f1_train_err)
#print(model, ' train error ', acc_train_err, ' f1_train ', f1_train_err, ' TEST ERROR ', acc, ' f1 ', f1)
acc_sum_test_np += acc
acc_sum_train_np += acc_train_err
counter +=1
avg_acc_np = acc_sum_test_np/counter
avg_acc_train_np = acc_sum_train_np/counter
print(model, 'NOO partial_fit' ' avg train error ', avg_acc_train_np, ' AVERAGE TEST ERROR ', avg_acc_np,)
#T = 800 #train until this game in season (out of 1200)
#d = 20 #predict this many games
#two independent scalers, one for <=T, one for T-d, T
scal = StandardScaler()
scal_T = StandardScaler()
#X_15 has about 1200 games
#set the data frame and target
X = X_15.copy()
y = y_15.loc[:, 'won'].copy()
#model = gnb
#model = sgdc
##quick checks
#for model in [gnb, sgdc]: #partial_fit
counter = 0
acc_sum_train = 0
acc_sum_test = 0
acc_sum_test_T = 0
for T in range(d, D ,d):
y_train= y.iloc[T-d:T].copy()
y_test = y.iloc[T:T+d].copy()
#y_test = z.iloc[:100].copy()
X_train = X.iloc[T-d:T, :].copy()
X_scaling_T = X.iloc[:T, :].copy() #use this to scale the test data ... if we use T-d to T scaler will fluctuate a lot.
X_test= X.iloc[T:T+d, :].copy()
X_test_T= X.iloc[T:T+d, :].copy()
#X_test = W.iloc[:100].copy()
#do standard/minmax scaling on X_train numeric columns ... better to do pipeline?
X_train.loc[:, columns_to_scale] = scal.fit_transform(X_train.loc[:, columns_to_scale]).copy()
#fit the scaler on all < = T and use this to transform [T to T+d] (and try not as well)
scal_T.fit(X_scaling_T.loc[:, columns_to_scale])
X_test_T.loc[:, columns_to_scale] = scal_T.transform(X_test_T.loc[:, columns_to_scale]).copy()
X_test.loc[:, columns_to_scale] = scal.transform(X_test.loc[:, columns_to_scale]).copy()
model.partial_fit(X_train, y_train)
y_pred_T = model.predict(X_test_T)
y_pred = model.predict(X_test)
acc_T = accuracy_score(y_test, y_pred_T)
acc= accuracy_score(y_test, y_pred)
f1 = f1_score(y_test,y_pred)
y_pred_train_err = model.predict(X_train) #! careful with this code
f1_train_err = f1_score(y_train,y_pred_train_err)
acc_train_err = accuracy_score(y_train, y_pred_train_err)
#print(' training error ', model, acc_train_err, ) #f1_train_err)
#print(model, 'with partial_fit ', ' train error ', acc_train_err, ' f1_train ', f1_train_err, ' TEST ERROR ', acc, ' f1 ', f1)
acc_sum_test += acc
acc_sum_test_T += acc_T
acc_sum_train += acc_train_err
counter +=1
avg_acc = acc_sum_test/counter
avg_acc_T = acc_sum_test_T/counter
avg_acc_train = acc_sum_train/counter
dic[D] = [D, avg_acc_train, avg_acc, avg_acc_T, avg_acc_np]
#print(dic[D])
print(model, 'with partial_fit ', ' avg train accuracy ', avg_acc_train, ' AVERAGE TEST accuracy (acc, acc_T) ', avg_acc, avg_acc_T)
# -
dic
# +
model = gnb
##quick checks
#for model in [gnb, sgdc]: #partial_fit
for d in range(20,660,20):
model.fit(X.iloc[:d :], y_win[:d])
model.fit(X.iloc[:d :], y_win[:d])
#y_pred1 = pipe_lgr.predict(X.iloc[d:, :])
y_pred = model.predict(X.iloc[d:, :])
y_test = y_win[d:].copy()
acc = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test,y_pred)
acc = accuracy_score(y_test, y_pred)
print(d, model, acc, f1)
#print(d, pipe_lgr, acc1, f11)
#print(d, lgr, acc2, f12)
# -
# +
#T = 800 #train until this game in season (out of 1200)
d = 20 #predict this many games
scal = std_scal
#X_15 has about 1200 games
#set the data frame and target
X = X_15.copy()
y = y_15.loc[:, 'won'].copy()
model = sgdc
##quick checks
#for model in [gnb, sgdc]: #partial_fit
for T in range(900, 1100,20):
y_train= y.iloc[:T].copy()
y_test = y.iloc[T:T+d].copy()
#y_test = z.iloc[:100].copy()
X_train = X.iloc[:T, :].copy()
X_test= X.iloc[T:T+d, :].copy()
#X_test = W.iloc[:100].copy()
#do standard/minmax scaling on X_train numeric columns ... better to do pipeline?
X_train.loc[:, columns_to_scale] = scal.fit_transform(X_train.loc[:, columns_to_scale]).copy()
#fit the scaler from train portion to the test portion
X_test.loc[:, columns_to_scale] = scal.transform(X_test.loc[:, columns_to_scale]).copy()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
acc = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test,y_pred)
y_pred_train_err = model.predict(X_train) #! careful with this code
f1_train_err = f1_score(y_train,y_pred_train_err)
acc_train_err = accuracy_score(y_train, y_pred_train_err)
#print(' training error ', model, acc_train_err, ) #f1_train_err)
print(model, ' train error ', acc_train_err, ' f1_train ', f1_train_err, ' TEST ERROR ', acc, ' f1 ', f1)
# +
#T = 800 #train until this game in season (out of 1200)
d = 20 #predict this many games
scal = std_scal
#X_15 has about 1200 games
#set the data frame and target
X = X_15.copy()
y = y_15.loc[:, 'won'].copy()
model = gnb
##quick checks
#for model in [gnb, sgdc]: #partial_fit
for T in range(d, 500 ,d):
y_train= y.iloc[T-d:T].copy()
y_test = y.iloc[T:T+d].copy()
#y_test = z.iloc[:100].copy()
X_train = X.iloc[T-d:T, :].copy()
X_test= X.iloc[T:T+d, :].copy()
#X_test = W.iloc[:100].copy()
#do standard/minmax scaling on X_train numeric columns ... better to do pipeline?
X_train.loc[:, columns_to_scale] = scal.fit_transform(X_train.loc[:, columns_to_scale]).copy()
#fit the scaler from train portion to the test portion
X_test.loc[:, columns_to_scale] = scal.transform(X_test.loc[:, columns_to_scale]).copy()
model.partial_fit(X_train, y_train)
y_pred = model.predict(X_test)
acc = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test,y_pred)
y_pred_train_err = model.predict(X_train) #! careful with this code
f1_train_err = f1_score(y_train,y_pred_train_err)
acc_train_err = accuracy_score(y_train, y_pred_train_err)
#print(' training error ', model, acc_train_err, ) #f1_train_err)
print(model, 'with partial_fit ', ' train error ', acc_train_err, ' f1_train ', f1_train_err, ' TEST ERROR ', acc, ' f1 ', f1)
# +
model = gnb
##quick checks
#for model in [gnb, sgdc]: #partial_fit
for d in range(20,660,20):
model.fit(X.iloc[:d :], y_win[:d])
model.fit(X.iloc[:d :], y_win[:d])
#y_pred1 = pipe_lgr.predict(X.iloc[d:, :])
y_pred = model.predict(X.iloc[d:, :])
y_test = y_win[d:].copy()
acc = accuracy_score(y_test, y_pred)
f1 = f1_score(y_test,y_pred)
acc = accuracy_score(y_test, y_pred)
print(d, model, acc, f1)
#print(d, pipe_lgr, acc1, f11)
#print(d, lgr, acc2, f12)
# +
##example from stack overflow how to do multiple variable line graphs ...
num_rows = 20
years = list(range(1990, 1990 + num_rows))
data_preproc = pd.DataFrame({
'Year': years,
'A': np.random.randn(num_rows).cumsum(),
'B': np.random.randn(num_rows).cumsum(),
'C': np.random.randn(num_rows).cumsum(),
'D': np.random.randn(num_rows).cumsum()})
# -
data_preproc[0:3]
pd.melt(data_preproc, ['Year'])[0:3] ##seaborn WHY? would you do that
import pandas as pd
import seaborn as sns
sns.set()
sns.set_theme(style='darkgrid', context='talk')
sns.lineplot(x='Year', y='value', hue='variable',
data=pd.melt(data_preproc, ['Year']))
class_model_results(lgr, model_name ='logistic', X, dates, step, window_size, prediction_size, drop_first_k_days = 0):
|
Note_books/Explore_Models/.ipynb_checkpoints/v2_Model2_Pisch_Eval_more_seasons_corsi_xg-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # P21 Graphing Live Twitter Sentiment Analysis
#
# Now that we have live data coming in from the Twitter streaming API, why not also have a live graph that shows the sentiment trend? To do this, we're going to combine this tutorial with the live matplotlib graphing tutorial.
#
# If you want to know more about how the code works, see that tutorial. Otherwise:
# +
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from matplotlib import style
import time
style.use("ggplot")
fig = plt.figure()
ax1 = fig.add_subplot(1,1,1)
def animate(i):
pullData = open("twitter-out.txt","r").read()
lines = pullData.split('\n')
xar = []
yar = []
x = 0
y = 0
for l in lines[-200:]:
x += 1
if "pos" in l:
y += 1
elif "neg" in l:
y -= 1
xar.append(x)
yar.append(y)
ax1.clear()
ax1.plot(xar,yar)
ani = animation.FuncAnimation(fig, animate, interval=1000)
plt.show()
# -
|
lei/P21 Graphing Live Twitter Sentiment Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os, sys
import scipy
import numpy as np
from astropy.io import fits
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook
from astropy.table import Table
from scipy.ndimage import minimum_filter1d
from scipy.ndimage.filters import percentile_filter
plt.rcParams['font.size'] = 5
# -
d = 23
master_log = master_log = Table.read('/Users/arcticfox/Documents/youngStars/veloce/master_log.tab', format='ascii')
date = '2020-11-{0}'.format(d)
directory = '2011{0}'.format(d)
table = master_log[master_log['ObsDate']==date]
fileformat = '{0}nov3{1:04d}.fits'
table[table['Frame']==110]
# +
files = np.sort([i for i in os.listdir(directory) if i.endswith('.fits')])
science_frames, bias_frames, dark_frames, flat_frames = [], [], [], []
for i in range(len(table)):
if 'TIC' in table['ObjType'][i]:
science_frames.append(fileformat.format(d, table['Frame'][i]))
elif 'BiasFrame' == table['ObjType'][i]:
bias_frames.append(fileformat.format(d, table['Frame'][i]))
elif 'DarkFrame' in table['ObjType'][i]:
dark_frames.append(fileformat.format(d, table['Frame'][i]))
elif 'FlatField' in table['ObjType'][i]:
flat_frames.append(fileformat.format(d, table['Frame'][i]))
else:
continue
#dark_inds = dark_inds[np.argwhere(np.diff(dark_inds)>10)[0][0]+1:]
#bias_inds = bias_inds[4:]
#flat_inds = flat_inds[2:]
#science_frames = np.unique(np.sort([os.path.join(directory, i) for i in science_frames]))
bias_frames = np.unique(np.sort([os.path.join(directory, i) for i in bias_frames]))
dark_frames = np.unique(np.sort([os.path.join(directory, i) for i in dark_frames]))[6:46]#[27:-5]
flat_frames = np.unique(np.sort([os.path.join(directory, i) for i in flat_frames]))[20:]#[23:]
# -
len(bias_frames), len(dark_frames), len(flat_frames)
# ## Creating master frames
def master_file(files, output_fn, fntype='dark'):
arrs = []
for fn in tqdm_notebook(files):
hdu = fits.open(fn)
if hdu[0].data.shape == (4112, 4202):
arrs.append(hdu[0].data)
hdu.close()
arrs = np.array(arrs)
if fntype == 'bias' or fntype == 'dark':
masked = np.copy(arrs) + 0.0
for i in range(len(arrs)):
rows, cols = np.where(arrs[i]>1000)
masked[i][rows,cols] = np.nan
masked = np.array(masked)
med = np.nanmedian(masked, axis=0)
else:
med = np.nanmedian(arrs, axis=0)
np.save(output_fn, med)
return med
# +
if 'dark_med.npy' not in os.listdir(directory):
DARK_MED = master_file(dark_frames,
os.path.join(directory, 'dark_med.npy'),
fntype='dark')
else:
DARK_MED = np.load(os.path.join(directory, 'dark_med.npy'))
if 'bias_med.npy' not in os.listdir(directory):
BIAS_MED = master_file(bias_frames,
os.path.join(directory, 'bias_med.npy'),
fntype='bias')
else:
BIAS_MED = np.load(os.path.join(directory, 'bias_med.npy'))
if 'flat_med.npy' not in os.listdir(directory):
FLAT_MED = master_file(flat_frames,
os.path.join(directory, 'flat_med.npy'),
fntype='flat')
else:
FLAT_MED = np.load(os.path.join(directory, 'flat_med.npy'))
# -
# ## Science Frames
def extract_science(files):
outputfns = []
arrs = []
for fn in tqdm_notebook(files):
hdu = fits.open(os.path.join(directory,fn))
np.save(fn[:-5]+'.npy', hdu[0].data)
outputfns.append(fn[:-5]+'.npy')
arrs.append(hdu[0].data)
return np.array(arrs), outputfns
science_arrs, science_files = extract_science(science_frames)
directory
# # Creating the dot models
# +
# #%matplotlib inline
def get_outliers(x_value, sigma=0.8, plot=False):
arr = science_arrs[0][x_value:x_value+1][0] + 0.0
x = np.arange(0,len(arr),1,dtype=int)
outliers = np.where(arr >= np.nanmedian(arr) + sigma*np.nanstd(arr))[0]
if plot:
plt.figure(figsize=(1,1))
plt.plot(x, arr, 'k')
plt.ylim(800,1800)
plt.plot(x[outliers], arr[outliers], 'o')
plt.show()
return outliers
def group_inds(values, sep):
results = []
for i, v in enumerate(values):
if i == 0:
mini = maxi = v
temp = [v]
else:
# SETS 4 CADENCE LIMIT
if (np.abs(v-maxi) <= sep):
temp.append(v)
if v > maxi:
maxi = v
if v < mini:
mini = v
else:
results.append(int(np.nanmin(temp)))
mini = maxi = v
temp = [v]
# GETS THE LAST GROUP
if i == len(values)-1:
results.append(int(np.nanmin(temp)))
return np.array(results)
# +
rows, cols = np.where(science_arrs[0] > 1100)
mask = np.zeros(science_arrs[0].shape)
mask[rows,cols] = 1
# -
plt.imshow(DARK_MED[3000:3500,3000:3500], vmin=300, vmax=2000)
# %matplotlib notebook
plt.figure(figsize=(1,1))
plt.imshow(mask, cmap='Greys_r', vmin=0, vmax=1)#, alpha=0.9)
plt.plot(cols, rows, '.')
plt.colorbar()
plt.show()
# +
plt.figure(figsize=(1,1))
sargs = np.argsort(cols)
cols, rows = cols[sargs]+0, rows[sargs]+0
starts = np.where((rows<=795) & (rows>=770))[0]
ends = np.where((rows<=3330) & (rows>=3290))[0]
starts = group_inds(starts, sep=100)
ends = group_inds(ends, sep=100)
plt.plot(cols, rows, 'k.', ms=1)
plt.plot(cols[ends], rows[ends], '.', ms=1)
plt.plot(cols[starts], rows[starts], 'r.', ms=1)
# -
starts = np.delete(starts, [0, 35])
ends = np.delete(ends, [17, 39, 40])
len(starts), len(ends)
# +
mid_ends = np.where((rows>=2892) & (rows<=2908))[0]
mid_starts = np.where((rows>=1160) & (rows<=1180))[0]
mid = np.where((rows>=1995) & (rows<=2010))[0]
mid_starts = group_inds(mid_starts, sep=100)
mid_ends = group_inds(mid_ends, sep=100)
mid = group_inds(mid, sep=100)
plt.figure(figsize=(1,1))
ms = 3
plt.plot(cols, rows, 'k.', ms=1)
plt.plot(cols[mid_ends], rows[mid_ends], 'b.', ms=ms)
plt.plot(cols[starts], rows[starts], 'r.', ms=ms)
for i in range(len(mid_starts)):
plt.plot(cols[mid_starts[i]], rows[mid_starts[i]], '.', ms=ms)
plt.plot(cols[ends], rows[ends], 'g.', ms=ms)
plt.plot(cols[mid], rows[mid], 'y.', ms=ms)
# -
len(starts), len(mid_starts), len(mid), len(mid_ends), len(ends)
#starts = np.delete(starts, [23])
mid_starts = np.delete(mid_starts, [27, 29, 33, 35, 38, 42, 43])
mid = np.delete(mid, [17, 38, 39])
mid_ends = np.delete(mid_ends, [24, 30, 37, 40, 41, 42])
#ends = np.delete(ends, [-1])
len(starts), len(mid_starts), len(mid), len(mid_ends), len(ends)
plt.figure(figsize=(1,1))
plt.plot(cols, rows, 'k.', ms=1)
plt.plot(cols[ends], rows[ends], 'b.', ms=1)
plt.plot(cols[starts], rows[starts], 'r.', ms=1)
plt.plot(cols[mid_starts], rows[mid_starts], '.', c='darkorange', ms=1)
plt.plot(cols[mid_ends], rows[mid_ends], 'g.', ms=1)
plt.plot(cols[mid], rows[mid], 'y.', ms=1)
dot_array = np.array([starts, mid_starts, mid, mid_ends, ends])
fit_x = np.arange(300, 4000, 1)
plt.figure(figsize=(1,1))
plt.plot(rows, cols, 'k.', ms=1)
models = np.zeros((len(mid), len(fit_x)))
for i in range(len(mid)):
plt.plot(rows[dot_array[:,i]], cols[dot_array[:,i]], '.', ms=1)
fit = np.polyfit(rows[dot_array[:,i]], cols[dot_array[:,i]], deg=2)
model = np.poly1d(fit)
plt.plot(fit_x, model(fit_x), lw=1)
models[i] = model(fit_x)
np.save('./{0}/models.npy'.format(directory), models)
# ## Discretize model gap fits
discrete = np.zeros(models.shape, dtype=int)
for i in range(len(models)):
discrete[i] = np.round(models[i])
np.save('./{0}/discrete_models.npy'.format(directory), discrete)
|
feinstein_notebooks/create_order_models_201123.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Face Generation
# In this project, you'll use generative adversarial networks to generate new images of faces.
# ### Get the Data
# You'll be using two datasets in this project:
# - MNIST
# - CelebA
#
# Since the celebA dataset is complex and you're doing GANs in a project for the first time, we want you to test your neural network on MNIST before CelebA. Running the GANs on MNIST will allow you to see how well your model trains sooner.
#
# If you're using [FloydHub](https://www.floydhub.com/), set `data_dir` to "/input" and use the [FloydHub data ID](http://docs.floydhub.com/home/using_datasets/) "R5KrjnANiKVhLWAkpXhNBe".
# +
# # !conda install -y tqdm pillow matplotlib
# +
data_dir = './data'
# FloydHub - Use with data ID "R5KrjnANiKVhLWAkpXhNBe"
#data_dir = '/input'
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import helper
helper.download_extract('mnist', data_dir)
helper.download_extract('celeba', data_dir)
# -
# ## Explore the Data
# ### MNIST
# As you're aware, the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset contains images of handwritten digits. You can view the first number of examples by changing `show_n_images`.
# +
show_n_images = 25
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# %matplotlib inline
import os
from glob import glob
from matplotlib import pyplot
mnist_images = helper.get_batch(glob(os.path.join(data_dir, 'mnist/*.jpg'))[:show_n_images], 28, 28, 'L')
pyplot.imshow(helper.images_square_grid(mnist_images, 'L'), cmap='gray')
# -
# ### CelebA
# The [CelebFaces Attributes Dataset (CelebA)](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) dataset contains over 200,000 celebrity images with annotations. Since you're going to be generating faces, you won't need the annotations. You can view the first number of examples by changing `show_n_images`.
# +
show_n_images = 25
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
mnist_images = helper.get_batch(glob(os.path.join(data_dir, 'img_align_celeba/*.jpg'))[:show_n_images], 28, 28, 'RGB')
pyplot.imshow(helper.images_square_grid(mnist_images, 'RGB'))
# -
# ## Preprocess the Data
# Since the project's main focus is on building the GANs, we'll preprocess the data for you. The values of the MNIST and CelebA dataset will be in the range of -0.5 to 0.5 of 28x28 dimensional images. The CelebA images will be cropped to remove parts of the image that don't include a face, then resized down to 28x28.
#
# The MNIST images are black and white images with a single [color channel](https://en.wikipedia.org/wiki/Channel_(digital_image%29) while the CelebA images have [3 color channels (RGB color channel)](https://en.wikipedia.org/wiki/Channel_(digital_image%29#RGB_Images).
# ## Build the Neural Network
# You'll build the components necessary to build a GANs by implementing the following functions below:
# - `model_inputs`
# - `discriminator`
# - `generator`
# - `model_loss`
# - `model_opt`
# - `train`
#
# ### Check the Version of TensorFlow and Access to GPU
# This will check to make sure you have the correct version of TensorFlow and access to a GPU
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
from distutils.version import LooseVersion
import warnings
import tensorflow as tf
# Check TensorFlow Version
assert LooseVersion(tf.__version__) >= LooseVersion('1.0'), 'Please use TensorFlow version 1.0 or newer. You are using {}'.format(tf.__version__)
print('TensorFlow Version: {}'.format(tf.__version__))
# Check for a GPU
if not tf.test.gpu_device_name():
warnings.warn('No GPU found. Please use a GPU to train your neural network.')
else:
print('Default GPU Device: {}'.format(tf.test.gpu_device_name()))
# -
# ### Input
# Implement the `model_inputs` function to create TF Placeholders for the Neural Network. It should create the following placeholders:
# - Real input images placeholder with rank 4 using `image_width`, `image_height`, and `image_channels`.
# - Z input placeholder with rank 2 using `z_dim`.
# - Learning rate placeholder with rank 0.
#
# Return the placeholders in the following the tuple (tensor of real input images, tensor of z data)
# +
import problem_unittests as tests
def model_inputs(image_width, image_height, image_channels, z_dim):
"""
Create the model inputs
:param image_width: The input image width
:param image_height: The input image height
:param image_channels: The number of image channels
:param z_dim: The dimension of Z
:return: Tuple of (tensor of real input images, tensor of z data, learning rate)
"""
# TODO: Implement Function
image_input = tf.placeholder(tf.float32, (None, image_width, image_height, image_channels), name='image_input')
z_input = tf.placeholder(tf.float32, (None, z_dim), name='z_input')
lr = tf.placeholder(tf.float32, name='learning_rate')
return image_input, z_input, lr
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_model_inputs(model_inputs)
# -
# ### Discriminator
# Implement `discriminator` to create a discriminator neural network that discriminates on `images`. This function should be able to reuse the variables in the neural network. Use [`tf.variable_scope`](https://www.tensorflow.org/api_docs/python/tf/variable_scope) with a scope name of "discriminator" to allow the variables to be reused. The function should return a tuple of (tensor output of the discriminator, tensor logits of the discriminator).
# +
def discriminator(images, reuse=False):
"""
Create the discriminator network
:param images: Tensor of input image(s)
:param reuse: Boolean if the weights should be reused
:return: Tuple of (tensor output of the discriminator, tensor logits of the discriminator)
"""
# TODO: Implement Function
kernel_size = 5
alpha = 0.2
with tf.variable_scope('discriminator', reuse=reuse):
# (28, 82, 3) -> (14,14, 128) -> (7, 7, 256) -> Flatten -> FC
# input: (None, 28, 28, 3)
x = images
# Conv2D(stride=2) -> output: (None, 14, 14, 128), note: we don't use BatchNorm for the 1st layer
x = tf.layers.conv2d(x, 128, kernel_size, strides=2, padding='same')
x = tf.maximum(alpha*x, x)
# Conv2D(stride=2) -> output(None, 7, 7, 256)
x = tf.layers.conv2d(x, 256, kernel_size, strides=2, padding='same')
x = tf.layers.batch_normalization(x)
x = tf.maximum(alpha*x, x)
# Flatten
x = tf.reshape(x, (-1, 7 * 7 * 256))
# FC/Logits
logits = tf.layers.dense(x, 1, activation=None)
# Sigmoid
output = tf.sigmoid(logits)
return output, logits
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_discriminator(discriminator, tf)
# -
# ### Generator
# Implement `generator` to generate an image using `z`. This function should be able to reuse the variables in the neural network. Use [`tf.variable_scope`](https://www.tensorflow.org/api_docs/python/tf/variable_scope) with a scope name of "generator" to allow the variables to be reused. The function should return the generated 28 x 28 x `out_channel_dim` images.
# +
def generator(z, out_channel_dim, is_train=True):
"""
Create the generator network
:param z: Input z
:param out_channel_dim: The number of channels in the output image
:param is_train: Boolean if generator is being used for training
:return: The tensor output of the generator
"""
# TODO: Implement Function
alpha = 0.2
kernel_size = 5
# When training, we want reuse the vars, but
with tf.variable_scope('generator', reuse=(not is_train)):
# FC -> Reshape_to_Cube -> Conv2D_T -> Conv2D_T -> Tanh
# FC /wo activation
x = z
x = tf.layers.dense(x, 7*7*256, activation=None)
# Reshape to the cube
x = tf.reshape(x, (-1, 7, 7, 256))
x = tf.layers.batch_normalization(x, training=is_train)
x = tf.maximum(alpha*x, x)
# Conv2D_T -> (None, 14, 14, 128)
x = tf.layers.conv2d_transpose(x, 128, kernel_size, strides=2, padding='same', activation=None)
x = tf.layers.batch_normalization(x, training=is_train)
x = tf.maximum(alpha*x, x)
# Conv2D_T -> (None, 28, 28, out_channel_dim=1 or 3)
x = tf.layers.conv2d_transpose(x, out_channel_dim, kernel_size, strides=2, padding='same', activation=None)
#x = tf.layers.batch_normalization(x, training=is_train)
#x = tf.maximum(alpha*x, x)
# Tanh
output = tf.tanh(x)
return output
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_generator(generator, tf)
# -
# ### Loss
# Implement `model_loss` to build the GANs for training and calculate the loss. The function should return a tuple of (discriminator loss, generator loss). Use the following functions you implemented:
# - `discriminator(images, reuse=False)`
# - `generator(z, out_channel_dim, is_train=True)`
# +
def model_loss(input_real, input_z, out_channel_dim):
"""
Get the loss for the discriminator and generator
:param input_real: Images from the real dataset
:param input_z: Z input
:param out_channel_dim: The number of channels in the output image
:return: A tuple of (discriminator loss, generator loss)
"""
# TODO: Implement Function
g_model = generator(input_z, out_channel_dim, is_train=True)
d_model_real, d_logits_real = discriminator(input_real, reuse=False)
d_model_fake, d_logits_fake = discriminator(g_model, reuse=True)
sigmoid_ce_with_logits = tf.nn.sigmoid_cross_entropy_with_logits
# Label smoothing for Discriminator
# - https://github.com/soumith/ganhacks
# - https://arxiv.org/pdf/1606.03498.pdf
d_loss_real = tf.reduce_mean(sigmoid_ce_with_logits(logits=d_logits_real, labels=tf.ones_like(d_model_real) * 0.9))
#d_loss_fake = tf.reduce_mean(sigmoid_ce_with_logits(logits=d_logits_fake, labels=tf.zeros_like(d_model_fake)))
d_loss_fake = tf.reduce_mean(sigmoid_ce_with_logits(logits=d_logits_fake, labels=tf.ones_like(d_model_fake) * 0.1))
d_loss = d_loss_real + d_loss_fake
g_loss = tf.reduce_mean(sigmoid_ce_with_logits(logits=d_logits_fake, labels=tf.ones_like(d_model_fake)))
return d_loss, g_loss
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_model_loss(model_loss)
# -
# ### Optimization
# Implement `model_opt` to create the optimization operations for the GANs. Use [`tf.trainable_variables`](https://www.tensorflow.org/api_docs/python/tf/trainable_variables) to get all the trainable variables. Filter the variables with names that are in the discriminator and generator scope names. The function should return a tuple of (discriminator training operation, generator training operation).
# +
def model_opt(d_loss, g_loss, learning_rate, beta1):
"""
Get optimization operations
:param d_loss: Discriminator loss Tensor
:param g_loss: Generator loss Tensor
:param learning_rate: Learning Rate Placeholder
:param beta1: The exponential decay rate for the 1st moment in the optimizer
:return: A tuple of (discriminator training operation, generator training operation)
"""
# TODO: Implement Function
t_vars = tf.trainable_variables()
d_vars = [var for var in t_vars if var.name.startswith('discriminator')]
g_vars = [var for var in t_vars if var.name.startswith('generator')]
with tf.control_dependencies(tf.get_collection(tf.GraphKeys.UPDATE_OPS)):
d_train_opt = tf.train.AdamOptimizer(learning_rate, beta1=beta1).minimize(d_loss, var_list=d_vars)
g_train_opt = tf.train.AdamOptimizer(learning_rate, beta1=beta1).minimize(g_loss, var_list=g_vars)
return d_train_opt, g_train_opt
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_model_opt(model_opt, tf)
# -
# ## Neural Network Training
# ### Show Output
# Use this function to show the current output of the generator during training. It will help you determine how well the GANs is training.
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
def show_generator_output(sess, n_images, input_z, out_channel_dim, image_mode):
"""
Show example output for the generator
:param sess: TensorFlow session
:param n_images: Number of Images to display
:param input_z: Input Z Tensor
:param out_channel_dim: The number of channels in the output image
:param image_mode: The mode to use for images ("RGB" or "L")
"""
cmap = None if image_mode == 'RGB' else 'gray'
z_dim = input_z.get_shape().as_list()[-1]
example_z = np.random.uniform(-1, 1, size=[n_images, z_dim])
samples = sess.run(
generator(input_z, out_channel_dim, False),
feed_dict={input_z: example_z})
images_grid = helper.images_square_grid(samples, image_mode)
pyplot.imshow(images_grid, cmap=cmap)
pyplot.show()
# -
# ### Train
# Implement `train` to build and train the GANs. Use the following functions you implemented:
# - `model_inputs(image_width, image_height, image_channels, z_dim)`
# - `model_loss(input_real, input_z, out_channel_dim)`
# - `model_opt(d_loss, g_loss, learning_rate, beta1)`
#
# Use the `show_generator_output` to show `generator` output while you train. Running `show_generator_output` for every batch will drastically increase training time and increase the size of the notebook. It's recommended to print the `generator` output every 100 batches.
# +
def train(epoch_count, batch_size, z_dim, learning_rate, beta1, get_batches, data_shape, data_image_mode):
"""
Train the GAN
:param epoch_count: Number of epochs
:param batch_size: Batch Size
:param z_dim: Z dimension
:param learning_rate: Learning Rate
:param beta1: The exponential decay rate for the 1st moment in the optimizer
:param get_batches: Function to get batches
:param data_shape: Shape of the data
:param data_image_mode: The image mode to use for images ("RGB" or "L")
"""
# TODO: Build Model
image_width, image_height, image_channels = data_shape[1:] # data_shape=(n_batches, w, h, ch)
input_real, input_z, lr = model_inputs(image_width, image_height, image_channels, z_dim)
print('input data_shape: ', data_shape)
print('input_real: {}, input_z: {}, lr: {}'.format(input_real.get_shape(), input_z.get_shape(), learning_rate))
d_loss, g_loss = model_loss(input_real, input_z, image_channels)
d_train_opt, g_train_opt = model_opt(d_loss, g_loss, lr, beta1)
batch_step = 0
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch_i in range(epoch_count):
# batch_images: (None, W, H, Ch)
for batch_images in get_batches(batch_size):
# TODO: Train Model
batch_step += 1
# As Generator output is -1.0~1.0, amplify real image to -1.0~1.0 (as they were -0.5 to 0.5)
batch_images *= 2.0
batch_z = np.random.uniform(-1, 1, size=(batch_size, z_dim))
# Run optimizers
_ = sess.run(d_train_opt, feed_dict={input_real: batch_images, input_z: batch_z, lr: learning_rate})
_ = sess.run(g_train_opt, feed_dict={input_z: batch_z, lr: learning_rate})
# Show training status and images
if batch_step % 100 == 0:
# Train status
train_loss_d = d_loss.eval({input_real: batch_images, input_z: batch_z })
train_loss_g = g_loss.eval({input_z: batch_z})
print("Epoch {}/{}, Batch Step:{}, Proc Imgs: {}k ...".format(epoch_i+1, epoch_count, batch_step, batch_step*batch_size//1000),
"Discriminator Loss: {:.4f}...".format(train_loss_d),
"Generator Loss: {:.4f}".format(train_loss_g))
if batch_step % 500 == 0:
show_generator_output(sess, 5**2, input_z, image_channels, data_image_mode)
# last
show_generator_output(sess, 5**2, input_z, image_channels, data_image_mode)
# -
# ### MNIST
# Test your GANs architecture on MNIST. After 2 epochs, the GANs should be able to generate images that look like handwritten digits. Make sure the loss of the generator is lower than the loss of the discriminator or close to 0.
# +
batch_size = 100
z_dim = 100
learning_rate = 2e-4
beta1 = 0.3
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
epochs = 2
mnist_dataset = helper.Dataset('mnist', glob(os.path.join(data_dir, 'mnist/*.jpg')))
with tf.Graph().as_default():
train(epochs, batch_size, z_dim, learning_rate, beta1, mnist_dataset.get_batches,
mnist_dataset.shape, mnist_dataset.image_mode)
# -
# ### CelebA
# Run your GANs on CelebA. It will take around 20 minutes on the average GPU to run one epoch. You can run the whole epoch or stop when it starts to generate realistic faces.
# +
batch_size = 100
z_dim = 100
learning_rate = 2e-4
beta1 = 0.3
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
epochs = 1
celeba_dataset = helper.Dataset('celeba', glob(os.path.join(data_dir, 'img_align_celeba/*.jpg')))
with tf.Graph().as_default():
train(epochs, batch_size, z_dim, learning_rate, beta1, celeba_dataset.get_batches,
celeba_dataset.shape, celeba_dataset.image_mode)
# -
# ### Submitting This Project
# When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_face_generation.ipynb" and save it as a HTML file under "File" -> "Download as". Include the "helper.py" and "problem_unittests.py" files in your submission.
|
face_generation/try_and_errors/dlnd_face_generation-b100.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install -r ../requirements.txt
from em_examples.DCWidgetResLayer2_5D import *
from IPython.display import display
# %matplotlib inline
from matplotlib import rcParams
rcParams['font.size'] = 16
# # Effects of a highly resisitive surface layer
# # Purpose
#
# For a direct current resistivity (DCR) survey, currents are injected to the earth, and flow.
# Depending upon the conductivity contrast current flow in the earth will be distorted, and these changes
# can be measurable on the sufurface electrodes.
# Here, we focus on a cylinder target embedded in a halfspace below a highly resistive surface layer, and investigate what are happening in the earth when static currents are injected. Different from a sphere case, which is a finite target, the resistive layer will also impact the illumination of the target (conductor or resistor).
# By investigating changes in currents, electric fields, potential, and charges upon different geometry, Tx and Rx location, we understand geometric effects of the resistive layer for DCR survey.
# # Setup
#
# <img src=https://github.com/geoscixyz/em_apps/blob/master/images/DC_ResLayer_Setup.png?raw=true>
# # Question
#
# - How does the cylinder affect the apparent resistivity without the resistive layer?
# - How does the resistive layer affect the apparent resistivity? Is there a difference if you add or remove the cylinder target?
# ## Plate model
# - **survey**: Type of survey
# - **A**: Electrode A (+) location
# - **B**: Electrode B (-) location
# - **M**: Electrode A (+) location
# - **N**: Electrode B (-) location
# - **$dz_{layer}$**: thickness of the resistive layer
# - **$zc_{ayer}$**: z location of the resistive layer
# - **xc**: x location of cylinder center
# - **zc**: z location of cylinder center
# - **$\rho_{1}$**: Resistivity of the half-space
# - **$\rho_{2}$**: Resistivity of the layer
# - **$\rho_{3}$**: Resistivity of the cylinder
# - **Field**: Field to visualize
# - **Type**: which part of the field
# - **Scale**: Linear or Log Scale visualization
#
# ### **Do not forget to hit Run Interact to update the figure after you made modifications**
app = ResLayer_app()
|
LabExercise_1/DC_Layer_Cylinder_2_5D.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('food')
# language: python
# name: python3
# ---
# +
from aiogram import Bot, Dispatcher, executor, types
from aiogram.types import ContentType
from aiogram.dispatcher.filters.state import State, StatesGroup
from aiogram.types.message import ContentTypes
from aiogram.dispatcher import FSMContext
from aiogram.contrib.fsm_storage.memory import MemoryStorage
from sqlalchemy import update
API_TOKEN = "<KEY>"
# -
import asyncio
from food.paths import *
from food.search import *
import pandas as pd
from food.psql import *
import pytz
timezones = pytz.all_timezones
import requests
from requests.structures import CaseInsensitiveDict
import urllib
from tzwhere import tzwhere
import nest_asyncio
nest_asyncio.apply()
def geocode(q):
geocoding_key = '<KEY>'
url = "https://api.geoapify.com/v1/geocode/search?"
params = {"apiKey":geocoding_key,
"text":q}
resp = requests.get(url + urllib.parse.urlencode(params)).json()
return pd.json_normalize(resp['features']).sort_values('properties.rank.importance',ascending = False)[['properties.lat','properties.lon']].iloc[0].to_list()
def get_tz(q):
lat,lon = geocode(q)
return tzwhere.tzwhere().tzNameAt(lat,lon)
async def async_get_tz(q):
return get_tz(q)
async def async_search_image(url, env='prod'):
return search_image(url,env)
async def async_geocode(q):
return geocode(q)
async def async_insert_on_conflict(*args, **qwargs):
return insert_on_conflict(*args, **qwargs)
async def add_sender(message):
sender = message['from'].to_python()
sender = pd.DataFrame(sender,index=[0]).drop(columns =['is_bot'])
await async_insert_on_conflict(sender,'users',unique_cols=['id'])
# +
# Initialize bot and dispatcher
bot = Bot(token=API_TOKEN)
storage = MemoryStorage()
dp = Dispatcher(bot, storage=storage)
dishes_table = Dishes.__table__
# -
ml_version = 0.2
"""0.2 - """
class CState(StatesGroup):
init = State()
photo_taken = State()
measured = State()
set_timezone = State()
change_last = State()
change_weight = State()
@dp.message_handler(commands=['start'])
async def send_welcome(message: types.Message):
await CState.init.set()
await message.reply("""Counting calories as easy as taking pictures. Just capture everything before you eat it\n
Now send a photo of your meal to try""")
@dp.message_handler(commands=['set_timezone'])
async def send_welcome(message: types.Message, state: FSMContext):
await CState.set_timezone.set()
await message.reply(f"please search your town to set timezone")
@dp.message_handler(commands=['test'])
async def send_welcome(message: types.Message):
reply_msg = types.message.Message(message_id=1931,from_user=message['from'])
await reply_msg.reply("""test""")
@dp.message_handler(commands=['change_last_item'])
async def change_last(message: types.Message, state: FSMContext):
print('change_last_item')
global m
m = message
message_id = engine.execute(f"""select message_id from food.dishes
where user_id={message['from']['id']} and
grams >0
order by id desc limit 1""").first()[0]
async with state.proxy() as data: data['message_id'] = message_id
reply_msg = types.message.Message(message_id = message_id,
from_user = types.user.User(id = message['from']['id']),
chat = types.chat.Chat(id = message['from']['id']))
await CState.change_last.set()
btns_text = tuple('change weight','remove','cancel')
keyboard_markup = types.ReplyKeyboardMarkup(row_width=2)
keyboard_markup.add(*(types.KeyboardButton(text) for text in btns_text))
await reply_msg.reply("do you want to cansel mesurment for this item ?", reply_markup=keyboard_markup)
# +
@dp.message_handler(state=CState.change_last)
async def change_last_remove(message: types.Message, state: FSMContext):
if message.text == 'remove':
async with state.proxy() as data: message_id = data['message_id']
stmt = (
dishes_table.update()
.where(dishes_table.c.message_id == message_id)
.values(grams=0)
.returning(dishes_table.c.id)
)
session.execute(stmt)
session.commit()
await message.reply(""" your last item measurment has been removed""")
elif message.text == 'change_weight':
await CState.change_weight.set()
btns_text = tuple([(str(p)) for p in range(10,510,10)])
keyboard_markup = types.ReplyKeyboardMarkup(row_width=2)
keyboard_markup.add(*(types.KeyboardButton(text) for text in btns_text))
await message.reply("set weight for the dish", reply_markup=keyboard_markup)
# -
@dp.message_handler(lambda message: message.text.isdigit(),state=CState.change_weight)
async def change_last_change(message: types.Message, state: FSMContext):
if message.text == 'change weight':
async with state.proxy() as data: message_id = data['message_id']
stmt = (
dishes_table.update()
.where(dishes_table.c.message_id == message_id)
.values(grams=0)
.returning(dishes_table.c.id)
)
session.execute(stmt)
session.commit()
await message.reply(""" your last item measurment has been removed""")
@dp.message_handler(state=CState.set_timezone)
async def set_timezone(message: types.Message, state: FSMContext):
await types.ChatActions.typing()
await add_sender(message)
tz = await async_get_tz(message.text)
df = pd.DataFrame([[message['from']['id'],'tz',tz,pd.Timestamp.utcnow()]],columns = ['user_id','property','value','timestamp'])
df.to_sql('user_properties',schema = schema,con = engine,if_exists = 'append',index = False)
await message.reply(f"your tz is set to {tz}")
@dp.message_handler(content_types=ContentType.PHOTO)
async def process_photo(message: types.Message, state: FSMContext):
global m
m = message
await types.ChatActions.typing()
await add_sender(message)
photo = message['photo'][-1]
await photo.download(reference_images_path/photo['file_id'])
image_url = await photo.get_url()
dish = await async_search_image(url=image_url, env='prod')
dish['photo_id'] = photo['file_id']
dish['message_id'] = message['message_id']
sender = message['from'].to_python()
dish['user_id'] = sender['id']
dish['ml_version'] = ml_version
dish['timestamp']=pd.Timestamp.utcnow()
# async with state.proxy() as data: data['dish'] = dish.to_dict(orient = 'records')[0]
dish.to_sql('dishes',schema = schema,if_exists = 'append',index = False,con=engine)
await CState.photo_taken.set()
async with state.proxy() as data: data['photo_id'] = photo['file_id']
btns_text = tuple([(str(p)) for p in range(10,510,10)])
keyboard_markup = types.ReplyKeyboardMarkup(row_width=2)
keyboard_markup.add(*(types.KeyboardButton(text) for text in btns_text))
await message.reply("set weight of the dish you are going to eat", reply_markup=keyboard_markup)
@dp.message_handler(lambda message: message.text.isdigit(), state=CState.photo_taken)
async def measure(message: types.Message, state: FSMContext):
grams = int(message.text)
last_photo_id,energy = engine.execute(f"""select photo_id,energy from {schema}.dishes
where user_id={message['from']['id']}
order by id desc limit 1""").first()
# async with state.proxy() as data: last_photo_id = data['photo_id']
stmt = (
dishes_table.update()
.where(dishes_table.c.photo_id == last_photo_id)
.values(grams=grams)
.returning(dishes_table.c.id)
)
session.execute(stmt)
session.commit()
# async with state.proxy() as data: dish = data['dish']
energy = energy/100*grams
today_consumed = pd.read_sql(f"""select energy,grams,timestamp from {schema}.dishes
where user_id = {message['from']['id']} and timestamp > now() - interval '24 hours'
and grams is not null;""",engine).set_index("timestamp")
today_consumed= today_consumed['energy']/100*today_consumed['grams']
user_tz = engine.execute(f"""select value from food.user_properties
where user_id={message['from']['id']} and
property='tz'
order by id desc limit 1""").first()
user_tz = user_tz[0] if user_tz else 'UTC'
today_consumed = today_consumed.tz_convert(user_tz)
now = pd.Timestamp.now(tz = user_tz)
today_consumed = today_consumed.reset_index()
this_morning = pd.Timestamp(year = now.year,month = now.month,day = now.day,hour = 3,tz = user_tz)
today_consumed = today_consumed[today_consumed['timestamp'] > pd.Timestamp(this_morning)][0].sum()
today_consumed
await message.reply(f"""you have consumed {energy} ccal with whis dish \n
today consumed {today_consumed}""")
if __name__ == '__main__':
executor.start_polling(dp)
m
|
aiogram_bot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 第3部 Pythonによるデータ分析|Pythonで学ぶ統計学入門
#
# ## 10章 分割表の検定
# ### 実装:p値の計算
# +
# 数値計算に使うライブラリ
import numpy as np
import pandas as pd
import scipy as sp
from scipy import stats
# グラフを描画するライブラリ
from matplotlib import pyplot as plt
import seaborn as sns
sns.set()
# 表示桁数の指定
# %precision 3
# グラフをjupyter Notebook内に表示させるための指定
# %matplotlib inline
# -
# p値を求める
1 - sp.stats.chi2.cdf(x = 6.667, df = 1)
# ### 実装:分割表の検定
# データの読み込み
click_data = pd.read_csv("3-10-1-click_data.csv")
print(click_data)
# 分割表形式に変換
cross = pd.pivot_table(
data = click_data,
values = "freq",
aggfunc = "sum",
index = "color",
columns = "click"
)
print(cross)
# 検定の実行
sp.stats.chi2_contingency(cross, correction = False)
|
stats-newtextbook-python/samples/3-10-分割表の検定.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow35]
# language: python
# name: conda-env-tensorflow35-py
# ---
from keras import datasets
import numpy as np
import matplotlib.pyplot as plt
import keras
from keras import models
from keras.layers import Dense, Activation, Conv2D, Flatten
from keras.optimizers import Adam
import math
import cv2
# Mnist data load
(x_train, y_train), (x_test, y_test) = datasets.mnist.load_data()
y_train = keras.utils.to_categorical(y_train)
y_test = keras.utils.to_categorical(y_test)
x_train.shape, y_train.shape
# +
x_train = x_train.reshape(-1, 28, 28, 1)
x_test = x_test.reshape(-1, 28, 28, 1)
x_train = x_train / 255.0
x_test = x_test / 255.0
# -
x_train.shape, y_train.shape
# Simple CNN Network build
def build_net():
model = models.Sequential()
model.add(Conv2D(16, (3,3), padding = 'same', input_shape = (28, 28, 1)))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(10))
model.add(Activation('softmax'))
model.compile(loss = 'categorical_crossentropy', optimizer = Adam(lr = 0.001)
,metrics = ['accuracy'])
return model
mnist_net = build_net()
class DataGenerator:
def __init__(self, x_set, y_set, x_shape, y_shape, batch_size, do_shuffle, do_augment):
self.x_set = x_set
self.y_set = y_set
self.total_batch = x_shape[0]
self.img_h = x_shape[1]
self.img_w = x_shape[2]
self.img_c = x_shape[3]
self.class_num = y_shape[1]
self.batch_size = batch_size
self.do_shuffle = do_shuffle
self.do_augment = do_augment
def change_brightness(self, img, brightness_range = np.random.uniform(0, 0.2)):
# convert BGR to HSV colorspace
# randomly change the brightness of image
if self.img_c < 3:
img = np.resize(img, (self.img_h, self.img_w, 3))
img = np.asarray(img, dtype=np.float32)
hsv = cv2.cvtColor(img, cv2.COLOR_BGR2HSV)
rand = brightness_range
hsv[:, :, 2] = rand*hsv[:, :, 2]
new_img = cv2.cvtColor(hsv, cv2.COLOR_HSV2BGR)
# new_img = cv2.cvtColor(new_img, cv2.COLOR_BGR2GRAY)
new_img = np.resize(new_img, (self.img_h, self.img_w, self.img_c))
return new_img
def zoom(self, img, zoom_range = np.random.randint(0, 10)):
# image zoom
zoom_pix = zoom_range
zoom_factor = 1 + (2*zoom_pix)/self.img_h
image = cv2.resize(img, None, fx=zoom_factor,
fy=zoom_factor, interpolation=cv2.INTER_LINEAR)
top_crop = (image.shape[0] - self.img_h)//2
left_crop = (image.shape[1] - self.img_w)//2
new_img = image[top_crop: top_crop+self.img_h,
left_crop: left_crop+self.img_w].reshape([self.img_h, self.img_w, self.img_c])
return new_img
def get_data(self, index):
if self.do_augment == True:
do_zoom = np.random.randint(0,2,1)
if do_zoom == 1:
X, Y = self.zoom(self.x_set[index]),self.y_set[index]
else:
X, Y = self.change_brightness(self.x_set[index]),self.y_set[index]
else:
X, Y = self.x_set[index],self.y_set[index]
return X, Y
def generator(self):
while True:
if self.do_shuffle == True:
idx_arr = np.random.permutation(self.total_batch)
else:
idx_arr = np.arange(self.total_batch)
for batch in range(0, len(idx_arr), self.batch_size):
l_bound = batch
r_bound = batch + self.batch_size
if r_bound > len(idx_arr):
r_bound = len(idx_arr)
l_bound = r_bound - self.batch_size
current_batch = idx_arr[l_bound:r_bound]
x_data = np.empty([self.batch_size, self.img_h, self.img_w, self.img_c], dtype = np.float32)
y_data = np.empty([self.batch_size, self.class_num], dtype = np.int32)
for i, v in enumerate(current_batch):
x_data[i], y_data[i] = self.get_data(v)
yield (x_data, y_data)
params = {'x_set' : x_train, 'y_set' : y_train, 'x_shape' : x_train.shape,
'y_shape' : y_train.shape, 'batch_size' : 128, 'do_shuffle' : False, 'do_augment' : True}
datagen = DataGenerator(**params)
mnist_net.fit_generator(datagen.generator(), steps_per_epoch = params['x_shape'][0] / params['batch_size'], epochs = 2)
result = mnist_net.evaluate(x_test,y_test)
print('Test Loss : ', result[0])
print('Test Accuracy : ', result[1]*100, '%')
datax,datay = next(datagen.generator())
datax.shape
check_num = 1
# different range
plt.imshow(np.squeeze(datax[check_num]), cmap = 'gray')
plt.colorbar()
plt.imshow(np.squeeze(x_train[check_num]), cmap = 'gray')
plt.colorbar()
|
keras_custom_generator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SotaYoshida/Lecture_DataScience/blob/2021/notebooks/Python_chapter2_ListLoop.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="9f041YDhyHKb"
#
# ## Pythonの基本 その2:
#
#
# [この章の目的]
# プログラミングで非常に重要な概念である配列や繰り返し処理について学ぶ。
#
# + [markdown] id="FxEjmIHu3StG"
# ### リスト
#
# 実際にいろんなデータを扱う際には、 値や変数をまとめて処理したくなります。
# こうしたときに必要になるのが以下で扱う[リスト]型の変数です。
#
# リスト型は、値や変数などを括弧[ ]で括り、カンマで区切ることで作ることができます。
#
# ```[1.0, 2.0, 3.0]```
# また好きな名前の変数にリストを代入する(リストに名前を割り振る)こともできます
# + id="dxpkCjfvjsDD" colab={"base_uri": "https://localhost:8080/"} outputId="05c4508c-8a0a-4cf9-899a-773c4d94bd69"
a=[178.0, 180.0, 153.0]
print(a)
print("変数aの型(type)は", type(a)) # 変数aの型をprint
# + [markdown] id="CI7AqILkzWYY"
# リストの要素にできるのは数値だけではなく、文字列のリストも作ることができます。
# + id="a7c5bw5O36CC"
b = [ "Aさん", "Bさん", "宇大太郎さん"]
# + [markdown] id="SkfATN0Jj_Oo"
# リストに入っている要素の数は、len関数(lengthの略)で見ることができます。
# + id="nexyb9lAkJC0" colab={"base_uri": "https://localhost:8080/"} outputId="fb6fda33-3e49-4c65-fc78-5204f0963c80"
print("リストaは", a)
print("長さは", len(a))
# + [markdown] id="7NWIj0tWzxGa"
# 当然、長さに名前をつけて適当な変数に突っ込むこともできます。
#
# + id="yQWDGLFCmszW" colab={"base_uri": "https://localhost:8080/"} outputId="77c74baa-c1d8-4243-8cf2-b471b4723389"
ln_a = len(a)
print("リストaの長さは", len(a), "で、型は", type(ln_a))
# + [markdown] id="gjwXE2cJz05q"
# 文字列と値を組み合わせたようなリストも作ることができます。
# (例:名前、身長、体重)
# ```["Aさん", 178, 66]```
#
# また、これを拡張して、入れ子に(リストのリストを作成)することもできます。
#
# ``` [ ["Aさん", 178,66], ["Bさん", 180, 70] ]```
# + [markdown] id="8kVQe42tkK-D"
# リストの中の要素にアクセスするときは、半角括弧を使って[整数]といった形で"番地"を指定します。
# このときの番地(あるいは座標といっても良いですが)を指定する整数のことをインデックス(index)と呼びます。
# Pythonでは、要素にアクセスするためのインデックスは1からではなく、0からカウントします。
#
#
# + id="kRMlRuwhj2Rc" colab={"base_uri": "https://localhost:8080/"} outputId="dabbeb45-5e5f-4bbe-ad9e-ab8f2456d056"
a = ["Aさん", 178, 66]
print(a[0])
print(a[1])
print(a[2])
# + [markdown] id="Ci65y6MBoSvU"
# ですので、a[3]にアクセスしようとすると、
#
# + id="mkON3qP1occT" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="82aba27a-4b87-460c-8da0-d2c78005d42f"
print(a[3])
# + [markdown] id="zHlSjTaJn9UU"
# list index out of range (リストのインデックスが用意された範囲(range)を逸脱している)というエラーが出ます。
#
#
# 慣れないうちは0からカウントするのを変に思うかもしれませんが、
# これはプログラミング言語の仕様によるもので、Pythonの他にもC/C++なども0からカウントします。
#
# (ちなみにFORTRAN, Juliaなどは1からカウントします。)
#
# ググると、こうした0-based indexingと呼ばれる言語の利点がいくつか出てきます。
# (正直私の好みは1-based indexingです)
# 代表的なものは負のインデックスが自然に使えることでしょうか?
# + id="D_hxwmblo4GK" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="10de7c9e-f89e-4c15-cb77-aba93985fa27"
print(a)
print(a[-1])
print(a[-2])
# + [markdown] id="zGabsr9X4CcI"
# **(編集注:10/25の続きはここから)**
# + [markdown] id="--1OHCHKo9An"
# [-1]でアクセスすれば末尾の要素に、[-2]番目の要素は後ろから二番目といった具合です。
#
# 長いリストを作って「えっとこれ前から何番目の要素だったっけ...?」というときに、
# 後ろから3番目の要素をサクッと取得できるのが便利なときもあります。
#
# 入れ子にしたリストの要素を見たり、使いたいときには、少々慣れが必要です。
# ```a = [ [[1,2], [3,4]], 5, 6]```
# このような場合では、一番外側の括弧から数えて、
# 何番目の階層になっているかを考えることが必要になります。
# 練習してみましょう。
#
# + [markdown] id="_yXPKxkk1XYh"
# **練習1**
#
#
# 1. [+コード]でセルを以下に追加し、下のリストを作成する
# ```tmp = [ ["Aさん", 178,66], ["Bさん", 180, 70] ]```
# 2. ```tmp[i][j]```のi,jに可能な整数を入れて、Aさんの身長、Bさんの体重などをprint関数で表示してみましょう。
#
# 3. 2.と同じことを、負のインデックスを駆使してやってみましょう。
#
# 4. ```a = [ [[1,2], [3,4]], 5, 6]```を作成して、
# len(a[0]) #0番目の要素の数をprintしてみましょう。
# 5. ```print(a[0][0][1])```を実行してみましょう。
#
# 6. 4.5.の挙動から推測して、aのリストの中から4を取り出したい場合どうすればいいのかprint関数を使いながら考えてみましょう。
#
#
#
#
#
# + id="8Nd1FeZynI9W"
a = [ [[1,2], [3,4]], 5, 6]
# + [markdown] id="ZB9epIx_VAQ6"
#
# ---
#
# #### リスト同士の演算や要素の追加
#
# 2つのリストを結合することもできます。
# + id="YUsuP81s3_BG"
a=[1,3] ; b=[2,4]
a+b
# + [markdown] id="hfEDbcvd6bFE"
#
# 上のリストを"座標"だと思えば、要素ごとの和になってほしいような気もしますが、
# リストの足し算は要素ごとの和ではなく、リストの結合を意味します。
#
# > (注) 要素ごとの和のような、数学などで必要な演算は、
# 以降で扱うnumpy.array型と呼ばれるものにすれば簡単に実行できます
#
# + [markdown] id="CDhimmq_0x-q"
# 長さや階層の異なる2つのリストでも可能です.
# + id="b3DFZtE602hP"
c=[1,2,4]; d=[[5,6],[7,8]]
c+d
# + [markdown] id="s568X9r-4c6J"
# リストにあとから要素を加えたくなるときもあります.
# そんなときは ```append```関数 か ```+=``` を使います。
# + id="b_1nWHpd4ptJ"
A = [ "Aさん", 178,66]
A.append("O型")
print(A)
B = [ "Bさん", 180,70]
B += ["A型"]
print(B)
# + [markdown] id="qpkIcDzA8S4X"
# 厳密には両者は違うのですが、この授業では見た目がスッキリするので後者を使います。
#
# 入れ子のリストを作るときは
# + id="lqfalNz788Rx"
data = [ ]
data += [ ["Aさん", 178,66] ]
data += [ [ "Bさん",180,70] ]
print("data", data)
# + [markdown] id="wT8k9-LB9Y74"
# とします。
# # += でリストに要素を追加するときは
# 「一番外側の[ ]に囲まれたものを左辺の変数の最上位の階層に加える」
# と覚えておけば、リストを追加したいときに[ ]が二重に必要な理由がわかるかと思います。
# + id="GRI40dgS1U-u"
data2 = [ ]
data2 += ["Aさん", 178,66] # 1重カッコ これだと、リストの"要素"をdata2に突っ込むのであって
data2 += [ "Bさん",180,70] # リストをdata2に突っ込むのではない点に注意!
print(data2)
print("data2", data2)
# + [markdown] id="u3Q7Qi8mnE4l"
# 上のコードだと、人ごとにデータが区切られていないので扱うのに不便ですね。
#
# リストに格納する情報に血液型を加えたりなんかして、
# 同じ様なことを手元でも是非やってみましょう。
# どんどん試すのが一番です.
# + id="_l9GJWZ98srB"
### 名前,身長,体重,血液型,住んでる市区町村
a = [ "Aさん", 178, 66, "A型", "宇都宮市"]
# + [markdown] id="cdCiouyW2oqJ"
# リストの要素は後から更新することもできます.
#
#
# + id="XIATihMK9Lh6"
data = [ ["Aさん", 178,66],["Bさん",180,70] ]
# + [markdown] id="vdRIu-li9Oqe"
# というリストがあったとして、Aさんの体重を修正したければ、
# + id="fZ0mjwjn2uXd"
data[0][2] = 58 #Aさんの体重を更新
print(data) #リストを表示
# + [markdown] id="BsLWAvmb3JDd"
# ### リスト操作の注意点
#
# 授業で扱う程度の内容のプログラミングに関する疑問は、
# ググれば自分でだいたい解決することができます。
# ただしこの項目で述べることは(とくに初学者にとって)
# 「明らかにバグがある(何かがオカシイ,意図しない振る舞いをする)のだけど、
# どこがおかしいのか分からないので**そもそもどうググっていいかが分からない**」という点で、
# 少し事情が異なります。
#
# 例を見せるために、以下の2種類のリストを用意します.
#
# + id="IUJ1-lAY3T0n"
data1 = [[ "Aさん", 178,66], [ "Bさん",180,70] ]
tmp = ["Aさん", 178,66]
data2 =[ tmp, tmp]
print("data1", data1)
print("data2", data2)
# + [markdown] id="t2FS4d0U3-rq"
# data2のようにまず雛形のリストtmpを作って人数分の長さを持つリストを作ってから中身を編集しようと考えた場合、
# + id="w-uX1Alb4OOi"
data2[1][0]="Bさん"
data2[1][1]=180
data2[1][2]=70
# + [markdown] id="iQwPT_WN4WG_"
# という操作を思いつきます。
# data2の2番目(0から数えて1番目)の要素だけを編集したつもりでもdata2をprintすると
#
# + id="rkK0uOpz4huX"
print(data2)
# + [markdown] id="rVVjGsPu4jhj"
# data2の最初の要素(Aさんのままであってほしいリスト)まで上書きされてしまっています!!
#
# これは直感に反しているという点で、初学者が陥りやすい落とし穴です。
# 「変数は値を格納する場所ではなく、値の保管される場所(id)を格納する場所だ」
# と1章の注で書きました。
#
# + id="xt1Q_B0f5QYL"
tmp = ["Aさん", 178,66]
data2 =[ tmp, tmp]
print(id(data2[0]), id(data2[1])) #それぞれのidを調べてprint
print("idが等しいか", id(data2[0])== id(data2[1])) #id()は変数のidを確認する関数
# + [markdown] id="HktuZ7f85_FY"
# 今の場合、data2を最初に作ったときには、0番目と1番目の要素(リスト)は同一のidを持つtmpというリストです。
# したがって、tmpの中身を書き換える操作(data2[1][0]="Bさん")は、tmpの更新を通してdata2の要素いずれもに影響します。
#
#
# + [markdown] id="LU95twWnrFGG"
# このように、(特に)リストを入れ子にする際には、注意が必要です。
# なんかへんだな?と思ったときは要素のidに気を配ってみるのも重要です.
#
# 上のコードで、意図したものと違う挙動を起こした原因は、
# リストtmpを参照する形でdata2を作ったことでした。
# これはcopyモジュール内のcopy関数を用いて配列のコピーを作成することで回避できます。
#
# (モジュールについては4章で説明します)
# + id="Bor6WGZzrIXO"
import copy #copyというモジュールをインポートする
tmp=["Aさん",178,66]
data2 = [ copy.copy(tmp), copy.copy(tmp)]
print(id(data2[0]) == id(data2[1])) # ← data2の0番目と1番目のidが同じ(参照元が同じ)だと困るのでFalseであってほしい
# + [markdown] id="EZQPPyoaryQS"
# また、リストのリスト(ネストされたリストといったりします)それ自体をcopyしたいときは、
# ```copy.deepcopy()```を使います。
# + id="WkRQ4_klncVY"
import copy
data = [[ "Aさん", 178,66], ["Bさん",180,70] ]
copydata = copy.copy(data)
deepcopydata = copy.deepcopy(data)
print(id(data), id(copydata),id(deepcopydata))
print(id(data[0]), id(copydata[0]), id(deepcopydata[0]))
# + [markdown] id="di-HvKQhpV4c"
# となり、```data```というリストと```copydata```という"リスト同士"が異なるidを持っていても、それぞれの0番目の要素のidを見てみると、同じものを参照していることがわかります。
#
# このように、ネストされたリストをコピーして別々に扱いたい場合は特に注意が必要です。
#
# (私も初めてPythonを使ったときに、この点に気が付かずに時間をかなり溶かしました)
#
#
# + [markdown] id="a0gQkWzeUqgj"
# ### インデックスの取得
# + [markdown] id="V_MaDugdUs2T"
#
# ```index```関数を使ってリスト内の、[興味のある要素]のインデックスを取得することができます。
# + id="DsvZFrmmU4iz"
tlist = [ "いちご", "りんご", "ぶどう"]
tlist.index("りんご")
# + [markdown] id="ZIyt7579U7bR"
# 重複する要素がある場合、初めにヒットしたインデックスを返します。
# + id="WD1qkBNvU8fH"
tlist2 = [ "いちご", "りんご", "ぶどう","メロン","りんご"]
tlist2.index("りんご")
# + [markdown] id="97rfD_PvVKU2"
# 複雑なデータを扱う際に「あれ、あの要素ってどこの番地にあるんだっけ?」といった状況に便利な関数です。
# + [markdown] id="rYMz3ClOhp9D"
# ### スライスを用いた部分リストの取得
# + [markdown] id="GSRzB_IPiPCO"
# 以下の```a```のようなリストがあったとき、
# + id="mkyfjA4ZiPtu"
a= [ "years", 1990, 1995, 2000, 2005,2010, 2015,2020]
# + [markdown] id="0WmC8_qwiZPi"
# 始点,コロン,終点でインデックスの範囲を指定して、部分的に取り出すことが出来ます。
# + id="z2fish5zixSf" colab={"base_uri": "https://localhost:8080/"} outputId="86967ee5-6486-4775-c7c6-dddb943062d1"
a[2:4]
# + [markdown] id="LWya_HtMjKXw"
# 負のインデックスを使用することもできます。
# + id="MKJ5-cJPi_xy" colab={"base_uri": "https://localhost:8080/"} outputId="0a7d52f9-7536-4355-80e2-6c06859b5d0a"
a[1:-1]
# + [markdown] id="COx3lOQZjCFo"
# 終点のインデックスに相当する要素は取り出されないことに注意しましょう。
# + id="XasfLIqRjHbe"
a[1:]
# + [markdown] id="sG4YvJTQjIE0"
# とすると、最後の要素まで含まれます。
#
# + [markdown] id="1ecruqL3pHA4"
# ### for文(ループ処理)
#
# for文(ループ処理とも呼ぶ)は、プログラミングの中で**最も重要な概念の一つ**です。
# 理由は、for文は"作業"を繰り返すための処理の一つで、
# プログラミングが最も効力を発揮する点だからです。
# (詳しくはノートのエクセルの処理などの話を参照)
#
# まず以下のコードを表示してみましょう:
# + id="VgjXia0upIya"
for i in range(5):
print(i)
# + [markdown] id="vLdv2seGpj7Z"
# 上のコードは、iを0から4まで変化させながら、
# iをprintする作業を繰り返しなさいというものです。
# range関数は、range型のオブジェクトを生成する関数で、
# range(5)は0から整数を5つ(0,1,2,3,4)を生成する関数です。
# やはり0始まりで、5が含まれないことに注意です。
#
#
# また、range関数の引数をrange(始点,終点,間隔)と3つにすると、
# 様々な数のリストのようなものを作ることができます。
# in 関数は、iにrange()で指定した数値の範囲(リストみたいなもの)を
# 順番に突っ込んでいく関数と理解しておけばひとまずOKです。
#
# 以下のような偶数(奇数)のときだけ調べたいなどの操作にも使えて便利です。
#
#
# + id="HJ_Q30y-qNZv"
for i in range(0,6,2):
print(i) #やはり6は含まれないことに注意
# + [markdown] id="YMq8UkMY1UVt"
# 次に、リストの中身を順番に表示させてみましょう。
# + id="DosIH6zO1aeW"
kudamono = ["いちご", "りんご","ぶどう","メロン"]
for tmp in kudamono:
print(tmp)
# + [markdown] id="Mi49kwOX1iJ0"
# 上のコードでは、tmpという変数にkudamonoというリストの中身が順番に当てはめられている事がわかります。
#
# ループを用いてリストの中身にアクセスする方法は主に2通りで、
#
# 1. インデックスのループを回す
# 2. 要素に順番にアクセスする
#
# で、上のコードは2.に相当します。
#
# 上の例で、1.の方法を採る場合は
# + id="SnMf5IBXAtuY"
for i in range(len(kudamono)):
print(kudamono[i])
# + [markdown] id="QbiNRP3tAwgD"
# とすればよいです。
#
# インデックスと要素を同時に取得して使いたいときには
# + id="1sBjeAU3A8CJ"
for i, tmp in enumerate(kudamono):
print(i, tmp)
# + [markdown] id="_iaSPpKgBBha"
# とします。
# これによって、「"いちご"の2個あとの要素を取得する」といった作業も実装できます。
# + id="3tNkMDDMBLi0"
for i, tmp in enumerate(kudamono):
if tmp == "いちご":
print("いちごの2個後ろの要素は", kudamono[i+2])
# + [markdown] id="s_IqtFV6rvKv"
# ##### **(重要!)Pythonでは操作をするブロックはインデントで指定される**
# + [markdown] id="FE1LiUrVqrbU"
# # さて、上のfor文のコードには、print関数の前に半角スペースが4つあるのに気がついたでしょうか?
# Pythonでは、for,whileなどの繰り返しや、if,tryなどの条件分岐・例外処理の一連の作業"ブロック"を、半角スペース4つをつかってインデントを下げることで表現します。
#
# 下のように、for文を使っているにも関わらず正しくインデントされていない場合はエラーが出ます。
# + id="ItpayRhWyBIZ"
for i in range(2):
print(i)
# + [markdown] id="z1T_hDMgyCHI"
# さて、インデントは初めての人には少々分かりづらいかもしれませんので、簡単な例で挙動を確認しておきましょう。
#
# **練習問題**
#
# 以下のコードを実行すると、
# 何回数字がprintされるでしょうか?
# コードを実行する前に、三択で答えてみてください。
# A. 8回 B. 11回 C. 13回
# + id="qM5QYdcJs5O-"
for i in range(2):
print(i)
for j in range(5):
print(i,j)
print(i,j)
# + [markdown] id="tZxftTZBszWq"
#
#
# i に関する繰り返し(ループと呼びます)は、i=0,1の2回で、
# インデントが1ブロック下げられた操作を実行しますので、print(i)を2回と、
# jに関するループを2回繰り返します。
# jはj=0,1,2,3,4をとり、jのループでは```print(i,j)```は合計2×5=10回呼ばれます。
# 最後の行にある```print(i,j)```はどのループの中にも入っていませんから、呼ばれるのは1回だけです。
#
# ということで、答えはC.の13回でした。
#
# + [markdown] id="llWzeLsx2kDq"
# 慣れないうちは
# + id="VAgzCZ6K2lUI"
for i in range(2):
print(i)
for j in range(5):
print(i,j)
## End j loop
## End i loop
# + [markdown] id="1geoDWVC2sjM"
# などのように、どこでループを閉じるのかをコメントしておくと良いかもしれません。
# コードをイジっていくうちに
# + id="USOd9Zdp2zaU"
for i in range(2):
print(i)
for j in range(5):
print(i,j)
# + [markdown] id="jWjLB9Dy243B"
# と言ったように意図しないインデントになってしまって、
# 正しい答えを与えない(バグを作ってしまう)可能性があります。
#
# > 細かい注) Pythonでは通常インデント幅は4つの半角スペースで指定されます。
# Google Colab. では、 [ツール]→[設定]→[エディタ]から変更できますので、
# 幅2が使いやすければ変更してください。
# また、プログラムファイルを編集する際、Tabキーを使ってインデントを指定するのと、
# スペースキーを用いてインデントを指定するのとで、どちらを好むかは、
# [タブ/スペース論争]として知られています。
# + [markdown] id="gPHPdQWAvKxc"
# forループを用いて、作業を繰り返すことができると言いました。
# たとえば、リスト内包表記と呼ばれる書き方で、
# 要素がたくさんのリストを一気につくることができます。
# + id="oNiE_0IHwpUm"
a = [ i for i in range(1,1000,2)] ## i を1から999まで2ずつ変えていったときの値を詰めたリストを作成
print(a) #表示してみましょう
# + [markdown] id="_r-xOslCuYQt"
# **練習**
#
# リスト
# ```data = [ ["Aさん", 178,66] , ["Bさん",180,70], ["Cさん", 165,55]]```を用意し、
# [体重を二乗して身長にかけ合わせた量]の総和を計算するコードを作ってみましょう。
#
# ヒント1: 身長は入れ子になっているリストの[1]番目,体重は[2]番目
# ヒント2: total=0.0を定義して、体重の二乗×身長をfor文を使ってどんどん足していく。
# + id="FJpla_3Fvbjw" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="64b8215a-beae-4946-d188-1e0c0b860172"
###以下は、不完全なコードです。これにあと3行ほど書き足してみましょう。
###ちなみに答えは 2156493です.
data = [ ["Aさん", 178,66] , ["Bさん",180,70], ["Cさん", 165,55]]
total = 0.0
for tmp in data :
total += tmp[1] * tmp[2]**2
print(total)
# + id="3BWzSAT0mYoO"
total = 0.0
for tmp in data:
total += tmp[1] * tmp[2]**2
# + [markdown] id="Uf275kXcK-JN"
# さて、ループ(for文)の使い方がなんとなく分かったでしょうか?
# どんどん使ってみて、なれておいてください。
#
# **第1回のレポートでも多用することになるはずです**
# + [markdown] id="qucZen5vJ9PE"
# 他にも、たとえば、 1, 10, 100, 1000, 10000,...というループを回したい時、
# + id="o7dH6aHKKP4i" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="407e88e9-5ba1-4ccf-d28e-677e195a060f"
for i in [1,10,100,1000,10000]:
print(i)
# + [markdown] id="JTg8PfD3KW5U"
# と書くのではなく、べき乗に対するループを回す
# + id="Lwz3s8taKcvF" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="d33812aa-b5bd-49e8-bba6-e9a234aa2ea0"
for p in range(6):
print(10**p)
# + [markdown] id="8qLiZx-VKhh-"
# など、考えたい値を直接ループで扱うのではなく、
# 間接的な値(今の場合[10のp乗]のp)を扱う、
# などの考え方も、コードをスッキリさせる上で重要です。
# + [markdown] id="66r8pzgrfwOi"
# ### 条件分岐
#
# `if`文では、書かれた条件が成立した場合、後に続くブロックを実行します。
#
# if文を使って命題の真偽を判定することで、条件分岐を表現することができます。
#
#
# + id="JOz7cyvQf2NX"
a=3.0; b =-0.3
if a > b:
print("aはbよりも大きい")
if b > a:
print("bはaよりも大きい") ##これは呼び出されない
# + [markdown] id="QF6tLC7agluM"
# 条件を満たすときはA,満たさないときはBをしたい、という場合には```else```を使う。
# + id="SacEU4f0gsmv"
if a< b:
print("aはbよりも小さい")
else:
print("a>=b")
# + [markdown] id="t0pZV6Ghg31K"
# ```elif```を用いれば、もう少し複雑な条件を表現することができる。
#
# たとえば、もし条件1が満たされればAを実行、満たされない場合、条件2が満たされればBを、さらに1も2も満たされない場合はCを実行する、という場合、
# + id="a68DTqL-hak5"
if a < b: #条件1
print("a<b")
elif a ==b: #条件2
print("a=b")
else:
print("a>b")
# + [markdown] id="fOvfCVFUiL1d"
# if文は入れ子構造にすることもできる。
# その際はインデントを使ってブロックを表現する。
#
# たとえば、 aが偶数の場合は値をそのまま表示して、
# aが奇数の場合、3の倍数かそうでないかで処理を変える事を考えると、以下のようになる
# + id="ioNZB5iAiM_d"
if a % 2 == 0 :
print(a)
else:
if a % 3 == 0 :
print(str(a)+"は3の倍数です")
else:
print(str(a)+"は3の倍数ではありません")
# + [markdown] id="eRVHarmaiiw7"
# そういえば昔「3の倍数のときだけアホになる」芸人がいましたね。皆さんの世代は覚えてないかもしれません。
# + [markdown] id="5qWPQPGMii74"
# if文を使って条件分岐を作るときは、
# **条件分岐にモレがないか**注意が必要です。
# 例えば、変数`a`の値によって処理を行う場合
# + [markdown] id="Tv3MJC6ukVIM"
# ```py
# if a > 0:
# ## なんらかの処理1
# if a < 0:
# ## なんらかの処理2
# ```
#
# と書いてしまうと、`a=0`の場合、if文を2つともすり抜けてしまい、バグの原因になることがあります。
# 少々面倒でも```else```を使って、意図しないすり抜けがないかチェックするのが良いでしょう。
# + id="2oZ3vaYb3s1t"
a = 0 #aをいろんな値に変えて実行してみてください
if a > 0: #aが0より大きい場合
print("処理1:a+2=", a+2)
elif a<0:
print("処理2:a*2=", a*2)
else :
print("ゼロだよ?なんにもしなくていいの?")
# + [markdown] id="5JH8ZSsU2lL8"
# ### タプル(tuple)型, 丸括弧
#
# リストに似た型としてタプルと辞書があります。
#
# タプルは"immutable"(要素が変更不可)なリストと覚えておけばよいでしょう。
# リストは要素を[]で囲むことで作ることができました。
# タプルは丸括弧()で囲むことで作ることができます
# + id="4CNAqWljyjS1"
a = (1.0, 2.0, 3.0)
print(a, type(a))
print(a[0]) ##要素にアクセスするときはタプルのときでもやはり角括弧を使う
# + [markdown] id="I1IILsgUyk4T"
# 値をリストのように格納しておきたい、という状況下かつ、リストの値を後で変更することがないなら
# タプルを使うのも一つの手です.
#
# タブルを使うメリットとしては以下の通りです:
# * (予期せず)値を更新しようとするとエラーを吐いて教えてくれる
# * (場合によりけりですが)リストよりも早く処理が実行される(ことがある)
#
# + id="d-wmV-24zIa9"
##たとえば...
a = (1.0, 2.0, 3.0)
##たとえば...2番目の2.0を4.0に変更しようとすると....
a[1] = 4.0
# + [markdown] id="Xp-zrOGUzclF"
# ↑エラーがでたかと思います。
#
# 次に中身が同じ(1から5000までの整数)リストとタプルを用意して、
# 1万回要素の和を計算するという計算をしてみましょう.
# (計算自体に意味はありません)
# timeというライブラリを使って2つの作業に係る時間を調べてみると...
# + id="RNRVJHBvzrRR" colab={"base_uri": "https://localhost:8080/"} outputId="c5a6efd2-4afc-41cf-971f-564cf9d7b6b6"
import time #時間を計測するためのライブラリをインポート
itnum=10000 #繰り返す回数を設定
#リストを使った計算
t0= time.time()
a = [ i for i in range(1,5001) ] #リストを定義
for i in range(itnum):
sum(a)
t1 = time.time()
#タプルを使った計算
t2= time.time()
b = ( i for i in range(1,5001)) #タプルを定義
for i in range(itnum):
sum(b)
t3 = time.time()
print("リストの処理にかかった時間", t1-t0)
print("タプルの処理にかかった時間", t3-t2)
# + [markdown] id="_DFBdeB61zTN"
# タプルの方が実行時間が短い事がわかります.
# 今の例では差は人間にとっては気にならない程度の差ですが、
# 複雑な処理になってコードがなかなか計算を完了しないときには、
# リストをタプルにするなど、コードのパフォーマンスを改善する作業が必要となります。
# + [markdown] id="X69TUPNt2nfh"
# ### 辞書型, 波括弧
#
# 辞書型は、キーと要素の2つをあわせ持つ型です.
# リストにいれたものをいっぺんに扱うときに、
# 毎回番地を指定したりループで要素を回して、望むものを持ってくるのは面倒です。
#
# たとえば以下の名前と年齢のリスト
# ```
# a=[[ "Aさん",25],["Bさん",21],["Cさん",18]]
# ```
#
# があったとき、これまで習った方法だけを駆使して
# Bさんの年齢を取得するには以下のようなコードになります
# + id="oWV6iYr520v5"
a=[[ "Aさん",25],["Bさん",21],["Cさん",18]]
for tmp in a:
if tmp[0] == "Bさん" :
print("Bさんの年齢=", tmp[1])
# + [markdown] id="QfC_TsDD3mYh"
# このような使い方をしたい場合には、予め名前と年齢という2つの関係のある量を辞書として定義してしまえばよいのです。
#
# 辞書型は波括弧{}で囲むことで構成できます.
#
#
#
# + id="GgnqzQYq4XbS"
Dict_age = {'Aさん' : 25, 'Bさん': 21, 'Cさん': 18}
# + [markdown] id="n25YgOic5Gnp"
# Bさんの値(今は年齢)を知りたければ以下のように一行で取得可能です.
# + id="bHh_DVUZ5KTH"
Dict_age["Bさん"]
# + [markdown] id="HP-pNfHg4Y5e"
# 辞書を作る際には、 要素を取り出すためのキー(key)と値(value)の組み合わせで指定します.
# その際
# ```
# {"key" : value}
# ```
# とkeyとvalueをコロン:をつかって区切り、複数要素を入れる際はカンマで区切ります.
# keyは文字列や数字(たとえば小中学校の出席番号とか)を使用することができ、
# valueは様々な型が使えます。
#
# 値(value)として、リストを保持することもできます.
# 次のように年齢と出身県のリストを値にもつ辞書にしてみましょう.
#
# + id="PWzZQh4p5lqC"
Dict = {'Aさん' : [25,"栃木県"], 'Bさん': [21,"茨城県"], 'Cさん': [18,"群馬県"]}
# + [markdown] id="cDz_Xk0I6D1k"
# Cさんの個人情報にアクセスする際は
# + id="gqsBmc4o6Rkn"
Dict["Cさん"]
# + [markdown] id="E9fJ-WT76ROd"
# とすればいいことがわかります.
#
# 慣れないうちはタプルや辞書を使わずリストだけ覚えておけば問題ないのですが
# (僕も結構面倒だからリストでやっちゃう)
# 複雑な処理になるとパフォーマンス差が顕著になったり、
# コードの可読性が低くなったり、 ミスの原因になるので
# タプルや辞書型をうまく組み合わせながら使いましょう。
#
# また複数の情報を一つのオブジェクトにまとめる際はDataFrameというライブラリがよく使われます。
# 辞書の上位互換のようなイメージです. そのうち以降の章で扱うかもしれません.
# + [markdown] id="EGUKtqjeeyk9"
# ### While文
#
# for文に似た概念としてWhile文が存在します
# + [markdown] id="snmJfYawx-7u"
# プログラムを書いていくうちに、繰り返しの数が前もってわからないケースに遭遇するかと思います.
#
# たとえば
# * [連続で6が5回でるまでサイコロをふる]
# * [利益がある値を超えるまで株の売買をくりかえす]
#
# といったイメージです。
# この様な場合は、何回処理を繰り返せば良いか予め知ることはほとんど不可能です。
#
# サイコロの例の場合だと、たとえば5回ふっただけで6が連続で出る奇跡的な状況も有りえますし、
# 1000回ふっても100万回降っても連続で5回は出ないかもしれません。
# (某漫画の地下チンチロ編のようなイカサマサイコロを使用するとグッと確率はあがります)
#
# このような処理を実装したい場合にはwhile文を使います.
#
# 以下のような状況を考えてみましょう
#
# > Aさんは100万円を手にカジノにやってきて、
# 掛け金が20万かつ50%の確率で勝ったり負けたりするギャンブルに目をつけました。
# 「手元のお金が150万以上か50万以下になったら帰ろう...」と心に決めて挑みます.
#
#
# + id="IDmdI51Mx-Tp"
import random #ランダムに勝ち負けを決めるためrandomというライブラリを使います
money = 1000000
while 500000 < money < 1500000: #手元に50万以上-150万未満ある限り賭け続ける
if random.choice([True,False]): ## choice([True,False])でTrue(勝ち)とFalse(負け)をランダムに生成
money += 200000 #true 勝ちの場合
print("勝った!!")
else :
money -= 200000#負けの場合 20万失う
print("負けた..")
### 500000 < money < 1500000がfalse つまりお金が50万以下か150万以上になったら
### while文を脱出して以下のprint文が読まれる
print("最終的な所持金は..."+str(money)+"円だ")
# + [markdown] id="b9q7gVgWATh0"
# といった具合です. 使用例が思い浮かばなければ[while文なんてのがある]とだけ覚えておけば当面はOKです.
#
# 上のコードはrandomモジュールの(疑似)乱数を用いているので、実行ごとに買ったり負けたり結果が変わります。
# 何度か実行して見てください。
#
# [余談] これを応用すると↓のようなものを作って遊ぶこともできます(作者は私ではありません)。
# 十亀vs松田シミュレータ https://mattz.xii.jp/yakiu/yakiu.html
# (十亀選手(ライオンズ)と松田選手(ホークス)の驚異的な相性を元に作られた対戦シミュレータ)
# + [markdown] id="xYWOCDT8M0Uj"
# ### ループ(for, while)内での特殊な操作
# + [markdown] id="2NxxShOxNFCH"
# #### break ループ処理を途中で抜けたい場合
# breakはfor文やwhile文を途中で抜け出すのに使います。
# 用途としては[目的を果たしたのでもうループを繰り返す必要がないとき]や
# [予期せぬ事が起きた場合にループ処理を終わらせてプログラムを終了するとき]などに使います.
#
# 先程のカジノの例で、 [最大10回ゲームをやるが1回でも負ければ即座に賭けをやめたい]場合、
# 以下のようなコードになります.
# + id="HPC5NF3FNUo3"
import random
money = 1000000
for i in range(10):
if random.choice([True,False]):
money += 200000
print("勝った!!")
else :
money -= 200000
print("負けたので帰ります")
break
print("最終的な所持金は..."+str(money)+"円だ")
# + [markdown] id="iwv7BiM8NG47"
# #### continue
# continue文は、forやwhile文の中で[以降の処理を無視する]のに使います.
# 具体的な用途としては[特定の条件を満たす場合にのみ適用する処理を書きたい]場合などがあります.
# 再びカジノの例で考えてみましょう.
#
# まず、Aさんに勝負のたびに所持金を叫ばせるコードは
# + id="eWE2Z2jVObxz"
import random
money = 1000000
while 500000 < money < 1500000:
if random.choice([True,False]):
money += 200000
else :
money -= 200000
print("今の所持金は..."+str(money)+"円だ")
print("最終的な所持金は..."+str(money)+"円だ")
# + [markdown] id="eMPfQSziPEOP"
# となります。これに少しずつ、より複雑な条件を加えていくことにしましょう.
# 最初の掛け金を一旦5万円にすることにして
# [連続で勝った場合、コインを投げて表が出たら掛け金を2倍にする]という条件をいれてみましょう。
# これをプログラムで表現するのには以下のような実装が考えられます.
#
# + id="M1F8_JYdPJFI"
import random
money = 1000000
hit = 0 #連続勝数を記録する変数hitを定義
bet = 50000 #掛け金をセット
while 500000 < money < 1500000:
if random.choice([True,False]):#勝ち負け50%
money += bet
hit +=1 #勝ったら連勝数をプラス1する
else :
money -= bet
hit = 0 #負けたら連続勝ち数を0にリセット
print("今の所持金は..."+str(money)+"円だ")
if hit > 2 and random.choice([True,False]) : #ここでのrandom.choiceはコインの表裏に相当
print("掛け金をレイズだ!!","次は"+str(bet)+"円を賭けるぜ")
bet = bet * 2
print("最終的な所持金は..."+str(money)+"円だ")
# + [markdown] id="A8XHg35QQtD0"
# さらに[掛け金をレイズするかどうか悩んで、コインを投げて決めるのは、所持金が80万以上のときだけ]
# という条件を実装するのにcontinueを使ってみましょう
# + id="5AgDgp8fQ3He"
import random
money = 1000000
hit = 0 #連続勝数を記録する変数hitを定義
bet = 50000 #掛け金をセット
while 500000 < money < 1500000:
if random.choice([True,False]):#勝ち負け50%
money += bet
hit +=1 #勝ったら連勝数をプラス1する
else :
money -= bet
hit = 0 #負けたら連続勝ち数を0にリセット
print("今の所持金は..."+str(money)+"円だ")
if money < 800000:
continue # もし所持金が80万未満ならcontinue(以下の処理は実行しない)
if hit > 2 and random.choice([True,False]) : #ここでのrandom.choiceはコインの表裏に相当
bet = bet * 2
print("掛け金をレイズだ!!","次は"+str(bet)+"円を賭けるぞ")
print("最終的な所持金は..."+str(money)+"円だ")
# + [markdown] id="p_ZvXiuc2ece"
# ### 例外処理
# + [markdown] id="6hnBZUQ1XpUW"
# 以下の処理は[ある値から都度10を引いていって平方根を取った値を表示する]というコードです
#
# + id="dWDJaf9G2hPi"
import math
#import mathの代わりに import numpy as np でも良い
s = 124
for i in range(20):
s -= 10
print(math.sqrt(s)) ##numpyライブラリを(npという名前で)importすればnp.sqrt()とも書けます
# + [markdown] id="ZX5FTHyOYdcF"
# しかし、あるところでsの値が負になってしまい、sqrtが計算できなくなってしまいます
# (虚数を導入すれば定義できますが、mathのsqrt関数は非負の引数に対して定義されていますのでエラーが出ます)
#
# エラーが出る=予期しないことが起こる とプログラムがそこで止まってしまいます。
# 通常はそれで良いのですが、複雑な状況になると[エラーを無視してとにかくプログラムを最後まで実行させたい][エラーが起こるときにエラーを回避するような仕組みをプログラム自体に実装したい]といった状況が起こります.
#
# 今の例でいうと[sの値が正なら平方根を表示して、負の場合はエラーメッセージだけを表示してエラーが起きた回数をカウントする]という作業が必要な場合は以下のように
# try: 試行したい処理のブロック
# except: 例外(エラー)が起こった場合の処理のブロック
# を駆使することで、最後までプログラムを実行させることができます.
# + id="7nhzIqnHZDRJ"
import math
#import mathの代わりに import numpy でも良い
s = 124
hit = 0
for i in range(20):
s -= 10
try:
print(math.sqrt(s)) ##numpyライブラリを(npという名前で)importすればnp.sqrt()とも書けます
except :
print("sの値が"+str(s)+"になったのでsqrtが計算できません")
hit += 1
print(str(hit)+"回 sqrtの計算でエラーがありました")
# + [markdown] id="vcWhB2ld2EjT"
# このノートブックでは説明しませんがexceptの後に具体的な例外を指定して
# 例外の種類に応じた操作を行うことも出来ます。
# https://docs.python.org/ja/3/library/exceptions.html
# + [markdown] id="dPK_KIGcyuod"
# # LICENSE
# + [markdown] id="q943wB7Z4DYK"
#
# Copyright (C) 2021 <NAME>
#
# [ライセンス:クリエイティブ・コモンズ 4.0 表示 (CC-BY 4.0)](https://creativecommons.org/licenses/by/4.0/deed.ja)
|
notebooks/Python_chapter2_ListLoop.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
# <h2>Explore data</h2>
df = pd.read_csv('china_gdp.csv')
df.head(9)
plt.figure(figsize=(8,5))
x_data, y_data = (df["Year"].values, df["Value"].values)
plt.plot(x_data, y_data, 'ro')
plt.ylabel('GDP')
plt.xlabel('Year')
plt.show()
# +
X = np.arange(-5.0, 5.0, 0.1)
Y = 1.0 / (1.0 + np.exp(-X))
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# <h2>Model</h2>
def sigmoid(x, Beta_1, Beta_2):
y = 1 / (1 + np.exp(-Beta_1*(x-Beta_2)))
return y
# +
beta_1 = 0.10
beta_2 = 1990.0
#logistic function
Y_pred = sigmoid(x_data, beta_1 , beta_2)
#plot initial prediction against datapoints
plt.plot(x_data, Y_pred*15000000000000.)
plt.plot(x_data, y_data, 'ro')
# -
# Normalize data
xdata =x_data/max(x_data)
ydata =y_data/max(y_data)
from scipy.optimize import curve_fit
popt, pcov = curve_fit(sigmoid, xdata, ydata)
#print the final parameters
print(" beta_1 = %f, beta_2 = %f" % (popt[0], popt[1]))
x = np.linspace(1960, 2015, 55)
x = x/max(x)
plt.figure(figsize=(8,5))
y = sigmoid(x, popt[0], popt[1])
plt.plot(xdata, ydata, 'ro', label='data')
plt.plot(x,y, linewidth=3.0, label='fit')
plt.legend(loc='best')
plt.ylabel('GDP')
plt.xlabel('Year')
plt.show()
# <h2>Evaluation</h2>
msk = np.random.rand(len(df)) < 0.8
train_x = xdata[msk]
test_x = xdata[~msk]
train_y = ydata[msk]
test_y = ydata[~msk]
popt, pcov = curve_fit(sigmoid,train_x,train_y)
y_hat = sigmoid(test_x,*popt)
print("Mean absolute error: %.2f" % np.mean(np.absolute(y_hat - test_y)))
print("Residual sum of squares (MSE): %.2f" % np.mean((y_hat - test_y) ** 2))
from sklearn.metrics import r2_score
print("R2-score: %.2f" % r2_score(y_hat , test_y) )
|
linearRegression/non-linear-regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="j1k6Y9afpxL6"
# # Intro
#
# [PyTorch](https://pytorch.org/) is a very powerful machine learning framework. Central to PyTorch are [tensors](https://pytorch.org/docs/stable/tensors.html), a generalization of matrices to higher ranks. One intuitive example of a tensor is an image with three color channels: A 3-channel (red, green, blue) image which is 64 pixels wide and 64 pixels tall is a $3\times64\times64$ tensor. You can access the PyTorch framework by writing `import torch` near the top of your code, along with all of your other import statements.
#
# This guide will help introduce you to the functionality of PyTorch, but don't worry too much about memorizing it: the assignments will link to relevant documentation where necessary.
# + colab={} colab_type="code" id="fwp6T5ZMteDC"
import torch
# + [markdown] colab_type="text" id="IvXp0rlPBqdQ"
# # Why PyTorch?
#
# One important question worth asking is, why is PyTorch being used for this course? There is a great breakdown by [the Gradient](https://thegradient.pub/state-of-ml-frameworks-2019-pytorch-dominates-research-tensorflow-dominates-industry/) looking at the state of machine learning frameworks today. In part, as highlighted by the article, PyTorch is generally more pythonic than alternative frameworks, easier to debug, and is the most-used language in machine learning research by a large and growing margin. While PyTorch's primary alternative, Tensorflow, has attempted to integrate many of PyTorch's features, Tensorflow's implementations come with some inherent limitations highlighted in the article.
#
# Notably, while PyTorch's industry usage has grown, Tensorflow is still (for now) a slight favorite in industry. In practice, the features that make PyTorch attractive for research also make it attractive for education, and the general trend of machine learning research and practice to PyTorch makes it the more proactive choice.
# + [markdown] colab_type="text" id="MCgwdP20r1yX"
# # Tensor Properties
# One way to create tensors from a list or an array is to use `torch.Tensor`. It'll be used to set up examples in this notebook, but you'll never need to use it in the course - in fact, if you find yourself needing it, that's probably not the correct answer.
# + colab={} colab_type="code" id="B0hgYekGsxlB"
example_tensor = torch.Tensor(
[
[[1, 2], [3, 4]],
[[5, 6], [7, 8]],
[[9, 0], [1, 2]]
]
)
# + [markdown] colab_type="text" id="9dO4C2oft7zq"
# You can view the tensor in the notebook by simple printing it out (though some larger tensors will be cut off)
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="U2FKEzeYuEOX" outputId="dfa12ff7-afd1-4737-a669-54f36b4209dd"
example_tensor
# + [markdown] colab_type="text" id="VUwlmUngw-VR"
# ## Tensor Properties: Device
#
# One important property is the device of the tensor - throughout this notebook you'll be sticking to tensors which are on the CPU. However, throughout the course you'll also be using tensors on GPU (that is, a graphics card which will be provided for you to use for the course). To view the device of the tensor, all you need to write is `example_tensor.device`. To move a tensor to a new device, you can write `new_tensor = example_tensor.to(device)` where device will be either `cpu` or `cuda`.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="R7SF44_Vw9h0" outputId="57f90e38-f9e1-4115-8f27-ebe651d5b2fa"
example_tensor.device
# + [markdown] colab_type="text" id="FkfySyFduHQi"
# ## Tensor Properties: Shape
#
# And you can get the number of elements in each dimension by printing out the tensor's shape, using `example_tensor.shape`, something you're likely familiar with if you've used numpy. For example, this tensor is a $3\times2\times2$ tensor, since it has 3 elements, each of which are $2\times2$.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="DKmfzpOBun0t" outputId="883009b6-7300-4329-f9ec-df99cc36d846"
example_tensor.shape
# + [markdown] colab_type="text" id="aL954xmAuq4b"
# You can also get the size of a particular dimension $n$ using `example_tensor.shape[n]` or equivalently `example_tensor.size(n)`
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="7IKy3BB8uqBo" outputId="7fac1275-132f-4d2b-bf63-73065a2aea6a"
print("shape[0] =", example_tensor.shape[0])
print("size(1) =", example_tensor.size(1))
# + [markdown] colab_type="text" id="3pzzG8bav5rl"
# Finally, it is sometimes useful to get the number of dimensions (rank) or the number of elements, which you can do as follows
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="l_j9qTwyv41-" outputId="5921cbd1-19a2-4543-9488-3f72c0cb4970"
print("Rank =", len(example_tensor.shape))
print("Number of elements =", example_tensor.numel())
# + [markdown] colab_type="text" id="gibyKQJQzLkm"
# # Indexing Tensors
#
# As with numpy, you can access specific elements or subsets of elements of a tensor. To access the $n$-th element, you can simply write `example_tensor[n]` - as with Python in general, these dimensions are 0-indexed.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="F87bFA5SzNz7" outputId="1b0a8381-6fd8-40b4-a5c8-88cc80029f8e"
example_tensor[1]
# + [markdown] colab_type="text" id="1CegGw5wzpGa"
# In addition, if you want to access the $j$-th dimension of the $i$-th example, you can write `example_tensor[i, j]`
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="bl1JSZcRz0xn" outputId="7f98e47b-66cb-4927-b784-7e4bcb9eb687"
example_tensor[1, 1, 0]
# + [markdown] colab_type="text" id="dyQRCRIa4NaY"
# Note that if you'd like to get a Python scalar value from a tensor, you can use `example_scalar.item()`
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="e56KSJOq4YOE" outputId="29e1fd13-32df-40c5-e558-3193fa5da629"
example_tensor[1, 1, 0].item()
# + [markdown] colab_type="text" id="wZdMEQfu0A7h"
# In addition, you can index into the ith element of a column by using `x[:, i]`. For example, if you want the top-left element of each element in `example_tensor`, which is the `0, 0` element of each matrix, you can write:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="x2cFxJx50eGH" outputId="e66eade9-4b4b-4c7a-ea99-a83195d10541"
example_tensor[:, 0, 0]
# + [markdown] colab_type="text" id="w-rTBP-1whd2"
# # Initializing Tensors
#
# There are many ways to create new tensors in PyTorch, but in this course, the most important ones are:
#
# [`torch.ones_like`](https://pytorch.org/docs/master/generated/torch.ones_like.html): creates a tensor of all ones with the same shape and device as `example_tensor`.
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="g7gbs4AnwlIo" outputId="b0c67ed9-e33f-47d6-d95c-e53bc4f90dec"
torch.ones_like(example_tensor)
# + [markdown] colab_type="text" id="_aIbSlaJy9Z0"
# [`torch.zeros_like`](https://pytorch.org/docs/master/generated/torch.zeros_like.html): creates a tensor of all zeros with the same shape and device as `example_tensor`
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="X4cWQduzzCd8" outputId="dbc8a5fa-8db1-4f6d-e38e-d1deb982ff36"
torch.zeros_like(example_tensor)
# + [markdown] colab_type="text" id="wsOmgS1izDS_"
# [`torch.randn_like`](https://pytorch.org/docs/stable/generated/torch.randn_like.html): creates a tensor with every element sampled from a [Normal (or Gaussian) distribution](https://en.wikipedia.org/wiki/Normal_distribution) with the same shape and device as `example_tensor`
#
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="2hto51IazDow" outputId="cb62a68a-6171-4d1e-eb9b-f31784464aac"
torch.randn_like(example_tensor)
# + [markdown] colab_type="text" id="HXp0i5Cf6AGj"
# Sometimes (though less often than you'd expect), you might need to initialize a tensor knowing only the shape and device, without a tensor for reference for `ones_like` or `randn_like`. In this case, you can create a $2x2$ tensor as follows:
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="RZRqt3-S6cUZ" outputId="7bef97cc-a303-4200-c0f8-ef9bf3cb4996"
torch.randn(2, 2, device='cpu') # Alternatively, for a GPU tensor, you'd use device='cuda'
# + [markdown] colab_type="text" id="JTkmDwVsrM6R"
# # Basic Functions
#
# There are a number of basic functions that you should know to use PyTorch - if you're familiar with numpy, all commonly-used functions exist in PyTorch, usually with the same name. You can perform element-wise multiplication / division by a scalar $c$ by simply writing `c * example_tensor`, and element-wise addition / subtraction by a scalar by writing `example_tensor + c`
#
# Note that most operations are not in-place in PyTorch, which means that they don't change the original variable's data (However, you can reassign the same variable name to the changed data if you'd like, such as `example_tensor = example_tensor + 1`)
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="FpfwOUdopsF_" outputId="32347400-2e6a-40c6-e6f1-21e6aacde795"
(example_tensor - 5) * 2
# + [markdown] colab_type="text" id="uciZnx4b3UjX"
# You can calculate the mean or standard deviation of a tensor using [`example_tensor.mean()`](https://pytorch.org/docs/stable/generated/torch.mean.html) or [`example_tensor.std()`](https://pytorch.org/docs/stable/generated/torch.std.html).
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="0ELXUKG7329z" outputId="720dd190-7dd4-43f1-e53c-cba4263eb2be"
print("Mean:", example_tensor.mean())
print("Stdev:", example_tensor.std())
# + [markdown] colab_type="text" id="_QsyTRym32SX"
# You might also want to find the mean or standard deviation along a particular dimension. To do this you can simple pass the number corresponding to that dimension to the function. For example, if you want to get the average $2\times2$ matrix of the $3\times2\times2$ `example_tensor` you can write:
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="eCJl3Im25B9k" outputId="4bd9decd-579e-462c-bde1-ee8d9d1b2061"
example_tensor.mean(0)
# Equivalently, you could also write:
# example_tensor.mean(dim=0)
# example_tensor.mean(axis=0)
# torch.mean(example_tensor, 0)
# torch.mean(example_tensor, dim=0)
# torch.mean(example_tensor, axis=0)
# + [markdown] colab_type="text" id="Vb-_5ubc8t97"
# PyTorch has many other powerful functions but these should be all of PyTorch functions you need for this course outside of its neural network module (`torch.nn`).
# + [markdown] colab_type="text" id="RtWjExD69JEs"
# # PyTorch Neural Network Module (`torch.nn`)
#
# PyTorch has a lot of powerful classes in its `torch.nn` module (Usually, imported as simply `nn`). These classes allow you to create a new function which transforms a tensor in specific way, often retaining information when called multiple times.
# + colab={} colab_type="code" id="UYrgloYo_slC"
import torch.nn as nn
# + [markdown] colab_type="text" id="uyCPVmTD_kkl"
# ## `nn.Linear`
#
# To create a linear layer, you need to pass it the number of input dimensions and the number of output dimensions. The linear object initialized as `nn.Linear(10, 2)` will take in a $n\times10$ matrix and return an $n\times2$ matrix, where all $n$ elements have had the same linear transformation performed. For example, you can initialize a linear layer which performs the operation $Ax + b$, where $A$ and $b$ are initialized randomly when you generate the [`nn.Linear()`](https://pytorch.org/docs/stable/generated/torch.nn.Linear.html) object.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="pNPaHPo89VrN" outputId="c14dc316-ae68-49d3-a8eb-8ad6e1464f01"
linear = nn.Linear(10, 2)
example_input = torch.randn(3, 10)
example_output = linear(example_input)
example_output
# + [markdown] colab_type="text" id="YGNULkJR_mzn"
# ## `nn.ReLU`
#
# [`nn.ReLU()`](https://pytorch.org/docs/stable/generated/torch.nn.ReLU.html) will create an object that, when receiving a tensor, will perform a ReLU activation function. This will be reviewed further in lecture, but in essence, a ReLU non-linearity sets all negative numbers in a tensor to zero. In general, the simplest neural networks are composed of series of linear transformations, each followed by activation functions.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="nGxVFS3nBASc" outputId="d5f57584-1bad-4803-ba8c-b69881db4a1f"
relu = nn.ReLU()
relu_output = relu(example_output)
relu_output
# + [markdown] colab_type="text" id="KzfOEZ03AJzA"
# ## `nn.BatchNorm1d`
#
# [`nn.BatchNorm1d`](https://pytorch.org/docs/stable/generated/torch.nn.BatchNorm1d.html) is a normalization technique that will rescale a batch of $n$ inputs to have a consistent mean and standard deviation between batches.
#
# As indicated by the `1d` in its name, this is for situations where you expects a set of inputs, where each of them is a flat list of numbers. In other words, each input is a vector, not a matrix or higher-dimensional tensor. For a set of images, each of which is a higher-dimensional tensor, you'd use [`nn.BatchNorm2d`](https://pytorch.org/docs/stable/generated/torch.nn.BatchNorm2d.html), discussed later on this page.
#
# `nn.BatchNorm1d` takes an argument of the number of input dimensions of each object in the batch (the size of each example vector).
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="O4tYsi9-G9vM" outputId="ba61d37c-a8af-4663-fcc2-1691c6d241de"
batchnorm = nn.BatchNorm1d(2)
batchnorm_output = batchnorm(relu_output)
batchnorm_output
# + [markdown] colab_type="text" id="EMZewDz9Idr1"
# ## `nn.Sequential`
#
# [`nn.Sequential`](https://pytorch.org/docs/stable/generated/torch.nn.Sequential.html) creates a single operation that performs a sequence of operations. For example, you can write a neural network layer with a batch normalization as
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="R3GhASjyJt3N" outputId="3ef779ca-a17b-42fd-f2e5-fbb5fdc60b13"
mlp_layer = nn.Sequential(
nn.Linear(5, 2),
nn.BatchNorm1d(2),
nn.ReLU()
)
test_example = torch.randn(5,5) + 1
print("input: ")
print(test_example)
print("output: ")
print(mlp_layer(test_example))
# + [markdown] colab_type="text" id="SToQiSv5K5Yb"
# # Optimization
#
# One of the most important aspects of essentially any machine learning framework is its automatic differentiation library.
# + [markdown] colab_type="text" id="r4GZFCZ0QqI1"
# ## Optimizers
#
# To create an optimizer in PyTorch, you'll need to use the `torch.optim` module, often imported as `optim`. [`optim.Adam`](https://pytorch.org/docs/stable/optim.html#torch.optim.Adam) corresponds to the Adam optimizer. To create an optimizer object, you'll need to pass it the parameters to be optimized and the learning rate, `lr`, as well as any other parameters specific to the optimizer.
#
# For all `nn` objects, you can access their parameters as a list using their `parameters()` method, as follows:
# + colab={} colab_type="code" id="AIcCbs35K4wY"
import torch.optim as optim
adam_opt = optim.Adam(mlp_layer.parameters(), lr=1e-1)
# + [markdown] colab_type="text" id="-BsPFZu2M0Xx"
# ## Training Loop
#
# A (basic) training step in PyTorch consists of four basic parts:
#
#
# 1. Set all of the gradients to zero using `opt.zero_grad()`
# 2. Calculate the loss, `loss`
# 3. Calculate the gradients with respect to the loss using `loss.backward()`
# 4. Update the parameters being optimized using `opt.step()`
#
# That might look like the following code (and you'll notice that if you run it several times, the loss goes down):
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="zm6lPx4sOJht" outputId="c21672bd-a306-42ab-face-9a299511a059"
train_example = torch.randn(100,5) + 1
adam_opt.zero_grad()
# We'll use a simple loss function of mean distance from 1
# torch.abs takes the absolute value of a tensor
cur_loss = torch.abs(1 - mlp_layer(train_example)).mean()
cur_loss.backward()
adam_opt.step()
print(cur_loss)
# + [markdown] colab_type="text" id="wDjhZBCeTc6o"
# ## `requires_grad_()`
#
# You can also tell PyTorch that it needs to calculate the gradient with respect to a tensor that you created by saying `example_tensor.requires_grad_()`, which will change it in-place. This means that even if PyTorch wouldn't normally store a grad for that particular tensor, it will for that specified tensor.
# + [markdown] colab_type="text" id="mB22ovHyUEvH"
# ## `with torch.no_grad():`
#
# PyTorch will usually calculate the gradients as it proceeds through a set of operations on tensors. This can often take up unnecessary computations and memory, especially if you're performing an evaluation. However, you can wrap a piece of code with `with torch.no_grad()` to prevent the gradients from being calculated in a piece of code.
# + [markdown] colab_type="text" id="kowb1M425CE_"
#
# ## `detach():`
#
# Sometimes, you want to calculate and use a tensor's value without calculating its gradients. For example, if you have two models, A and B, and you want to directly optimize the parameters of A with respect to the output of B, without calculating the gradients through B, then you could feed the detached output of B to A. There are many reasons you might want to do this, including efficiency or cyclical dependencies (i.e. A depends on B depends on A).
# + [markdown] colab_type="text" id="-9HY2wgKLOr-"
# # New `nn` Classes
#
# You can also create new classes which extend the `nn` module. For these classes, all class attributes, as in `self.layer` or `self.param` will automatically treated as parameters if they are themselves `nn` objects or if they are tensors wrapped in `nn.Parameter` which are initialized with the class.
#
# The `__init__` function defines what will happen when the object is created. The first line of the init function of a class, for example, `WellNamedClass`, needs to be `super(WellNamedClass, self).__init__()`.
#
# The `forward` function defines what runs if you create that object `model` and pass it a tensor `x`, as in `model(x)`. If you choose the function signature, `(self, x)`, then each call of the forward function, gets two pieces of information: `self`, which is a reference to the object with which you can access all of its parameters, and `x`, which is the current tensor for which you'd like to return `y`.
#
# One class might look like the following:
# + colab={} colab_type="code" id="WOip473tQs-d"
class ExampleModule(nn.Module):
def __init__(self, input_dims, output_dims):
super(ExampleModule, self).__init__()
self.linear = nn.Linear(input_dims, output_dims)
self.exponent = nn.Parameter(torch.tensor(1.))
def forward(self, x):
x = self.linear(x)
# This is the notation for element-wise exponentiation,
# which matches python in general
x = x ** self.exponent
return x
# + [markdown] colab_type="text" id="x4CUFH_GS5UY"
# And you can view its parameters as follows
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="YuelIiE4S3KR" outputId="27a52620-ca40-4dc8-dff5-4f3a56ba0e5b"
example_model = ExampleModule(10, 2)
list(example_model.parameters())
# + [markdown] colab_type="text" id="1F7E1wKN5tez"
# And you can print out their names too, as follows:
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="dYTuTDsQ5pnY" outputId="6635a493-7318-4688-bd18-bfba41d43e9d"
list(example_model.named_parameters())
# + [markdown] colab_type="text" id="iWPoIqX2UsaH"
# And here's an example of the class in action:
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="7NXwbg5tUroC" outputId="0836e447-7c37-464e-b196-048ae0a0cc73"
input = torch.randn(2, 10)
example_model(input)
# + [markdown] colab_type="text" id="6Ocol8DABScy"
# # 2D Operations
#
# You won't need these for the first lesson, and the theory behind each of these will be reviewed more in later lectures, but here is a quick reference:
#
#
# * 2D convolutions: [`nn.Conv2d`](https://pytorch.org/docs/master/generated/torch.nn.Conv2d.html) requires the number of input and output channels, as well as the kernel size.
# * 2D transposed convolutions (aka deconvolutions): [`nn.ConvTranspose2d`](https://pytorch.org/docs/master/generated/torch.nn.ConvTranspose2d.html) also requires the number of input and output channels, as well as the kernel size
# * 2D batch normalization: [`nn.BatchNorm2d`](https://pytorch.org/docs/stable/generated/torch.nn.BatchNorm2d.html) requires the number of input dimensions
# * Resizing images: [`nn.Upsample`](https://pytorch.org/docs/master/generated/torch.nn.Upsample.html) requires the final size or a scale factor. Alternatively, [`nn.functional.interpolate`](https://pytorch.org/docs/stable/nn.functional.html#torch.nn.functional.interpolate) takes the same arguments.
#
#
#
#
|
C1 - Build Basic Generative Adversarial Networks/Week 1/Intro_to_PyTorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Use scikit-learn and custom library to predict temperature with `ibm-watson-machine-learning`
# This notebook contains steps and code to train a Scikit-Learn model that uses a custom defined transformer and use it with Watson Machine Learning service. Once the model is trained, this notebook contains steps to persist the model and custom defined transformer to Watson Machine Learning Repository, deploy and score it using Watson Machine Learning python client.
#
# In this notebook, we use GNFUV dataset that contains mobile sensor readings data about humidity and temperature from Unmanned Surface Vehicles in a test-bed in Athens, to train a Scikit-Learn model for predicting the temperature.
#
# Some familiarity with Python is helpful. This notebook uses Python-3.8 and scikit-learn.
# ## Learning goals
#
# The learning goals of this notebook are:
#
# - Train a model with custom defined transformer
# - Persist the custom defined transformer and the model in Watson Machine Learning repository.
# - Deploy the model using Watson Machine Learning Service
# - Perform predictions using the deployed model
#
# ## Contents
# 1. [Set up the environment](#setup)
# 2. [Install python library containing custom transformer implementation](#install_lib)
# 3. [Prepare training data](#load)
# 4. [Train the scikit-learn model](#train)
# 5. [Save the model and library to WML Repository](#upload)
# 6. [Deploy and score data](#deploy)
# 7. [Clean up](#cleanup)
# 8. [Summary and next steps](#summary)
#
# <a id="setup"></a>
# ## 1. Set up the environment
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
#
# - Contact with your Cloud Pack for Data administrator and ask him for your account credentials
# ### Connection to WML
#
# Authenticate the Watson Machine Learning service on IBM Cloud Pack for Data. You need to provide platform `url`, your `username` and `api_key`.
username = 'PASTE YOUR USERNAME HERE'
api_key = 'PASTE YOUR API_KEY HERE'
url = 'PASTE THE PLATFORM URL HERE'
wml_credentials = {
"username": username,
"apikey": api_key,
"url": url,
"instance_id": 'openshift',
"version": '4.0'
}
# Alternatively you can use `username` and `password` to authenticate WML services.
#
# ```
# wml_credentials = {
# "username": ***,
# "password": ***,
# "url": ***,
# "instance_id": 'openshift',
# "version": '4.0'
# }
#
# ```
# ### Install and import the `ibm-watson-machine-learning` package
# **Note:** `ibm-watson-machine-learning` documentation can be found <a href="http://ibm-wml-api-pyclient.mybluemix.net/" target="_blank" rel="noopener no referrer">here</a>.
# !pip install -U ibm-watson-machine-learning
# +
from ibm_watson_machine_learning import APIClient
client = APIClient(wml_credentials)
# -
# ### Working with spaces
#
# First of all, you need to create a space that will be used for your work. If you do not have space already created, you can use `{PLATFORM_URL}/ml-runtime/spaces?context=icp4data` to create one.
#
# - Click New Deployment Space
# - Create an empty space
# - Go to space `Settings` tab
# - Copy `space_id` and paste it below
#
# **Tip**: You can also use SDK to prepare the space for your work. More information can be found [here](https://github.com/IBM/watson-machine-learning-samples/blob/master/cpd4.0/notebooks/python_sdk/instance-management/Space%20management.ipynb).
#
# **Action**: Assign space ID below
space_id = 'PASTE YOUR SPACE ID HERE'
# You can use `list` method to print all existing spaces.
client.spaces.list(limit=10)
# To be able to interact with all resources available in Watson Machine Learning, you need to set **space** which you will be using.
client.set.default_space(space_id)
# <a id="install_lib"></a>
#
# ## 2. Install the library containing custom transformer
# Library - `linalgnorm-0.1` is a python distributable package that contains the implementation of a user defined Scikit-Learn transformer - `LNormalizer` . <br>
# Any 3rd party libraries that are required for the custom transformer must be defined as the dependency for the corresponding library that contains implementation of the transformer.
#
#
# In this section, we will create the library and install it in the current notebook environment.
# !mkdir -p linalgnorm-0.1/linalg_norm
# Define a custom scikit transformer.
# +
# %%writefile linalgnorm-0.1/linalg_norm/sklearn_transformers.py
from sklearn.base import BaseEstimator, TransformerMixin
import numpy as np
class LNormalizer(BaseEstimator, TransformerMixin):
def __init__(self, norm_ord=2):
self.norm_ord = norm_ord
self.row_norm_vals = None
def fit(self, X, y=None):
self.row_norm_vals = np.linalg.norm(X, ord=self.norm_ord, axis=0)
def transform(self, X, y=None):
return X / self.row_norm_vals
def fit_transform(self, X, y=None):
self.fit(X, y)
return self.transform(X, y)
def get_norm_vals(self):
return self.row_norm_vals
# -
# Wrap created code into Python source distribution package.
# +
# %%writefile linalgnorm-0.1/linalg_norm/__init__.py
__version__ = "0.1"
# +
# %%writefile linalgnorm-0.1/README.md
A simple library containing a simple custom scikit estimator.
# +
# %%writefile linalgnorm-0.1/setup.py
from setuptools import setup
VERSION='0.1'
setup(name='linalgnorm',
version=VERSION,
url='https://github.ibm.com/NGP-TWC/repository/',
author='IBM',
author_email='<EMAIL>',
license='IBM',
packages=[
'linalg_norm'
],
zip_safe=False
)
# + language="bash"
#
# cd linalgnorm-0.1
# python setup.py sdist --formats=zip
# cd ..
# mv linalgnorm-0.1/dist/linalgnorm-0.1.zip .
# rm -rf linalgnorm-0.1
# -
# Install the downloaded library using `pip` command
# !pip install linalgnorm-0.1.zip
# <a id="load"></a>
#
# ## 3. Download training dataset and prepare training data
# Download the data from UCI repository - https://archive.ics.uci.edu/ml/machine-learning-databases/00452/GNFUV%20USV%20Dataset.zip
# !rm -rf dataset
# !mkdir dataset
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/00452/GNFUV%20USV%20Dataset.zip --output-document=dataset/gnfuv_dataset.zip
# !unzip dataset/gnfuv_dataset.zip -d dataset
# Create pandas datafame based on the downloaded dataset
import json
import pandas as pd
import numpy as np
import os
from datetime import datetime
from json import JSONDecodeError
# +
home_dir = './dataset'
pi_dirs = os.listdir(home_dir)
data_list = []
base_time = None
columns = None
for pi_dir in pi_dirs:
if 'pi' not in pi_dir:
continue
curr_dir = os.path.join(home_dir, pi_dir)
data_file = os.path.join(curr_dir, os.listdir(curr_dir)[0])
with open(data_file, 'r') as f:
line = f.readline().strip().replace("'", '"')
while line != '':
try:
input_json = json.loads(line)
sensor_datetime = datetime.fromtimestamp(input_json['time'])
if base_time is None:
base_time = datetime(sensor_datetime.year, sensor_datetime.month, sensor_datetime.day, 0, 0, 0, 0)
input_json['time'] = (sensor_datetime - base_time).seconds
data_list.append(list(input_json.values()))
if columns is None:
columns = list(input_json.keys())
except JSONDecodeError as je:
pass
line = f.readline().strip().replace("'", '"')
data_df = pd.DataFrame(data_list, columns=columns)
# -
data_df.head()
# Create training and test datasets from the downloaded GNFUV-USV dataset.
# +
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
Y = data_df['temperature']
X = data_df.drop('temperature', axis=1)
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.25, random_state=143)
# -
# <a id="train"></a>
#
# ## 4. Train a model
#
# In this section, you will use the custom transformer as a stage in the Scikit-Learn `Pipeline` and train a model.
# #### Import the custom transformer
# Here, import the custom transformer that has been defined in `linalgnorm-0.1.zip` and create an instance of it that will inturn be used as stage in `sklearn.Pipeline`
from linalg_norm.sklearn_transformers import LNormalizer
lnorm_transf = LNormalizer()
# Import other objects required to train a model
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LinearRegression
# Now, you can create a `Pipeline` with user defined transformer as one of the stages and train the model
skl_pipeline = Pipeline(steps=[('normalizer', lnorm_transf), ('regression_estimator', LinearRegression())])
skl_pipeline.fit(X_train.loc[:, ['time', 'humidity']].values, y_train)
y_pred = skl_pipeline.predict(X_test.loc[:, ['time', 'humidity']].values)
rmse = np.mean((np.round(y_pred) - y_test.values)**2)**0.5
print('RMSE: {}'.format(rmse))
# <a id="upload"></a>
#
# ## 5. Persist the model and custom library
#
# In this section, using `ibm-watson_machine_learning` SDK, you will ...
# - save the library `linalgnorm-0.1.zip` in WML Repository by creating a package extension resource
# - create a Software Specification resource and bind the package resource to it. This Software Specification resource will be used to configure the online deployment runtime environment for a model
# - bind Software Specification resource to the model and save the model to WML Repository
# ### Create package extension
# Define the meta data required to create package extension resource. <br>
#
# The value for `file_path` in `client.package_extensions.LibraryMetaNames.store()` contains the library file name that must be uploaded to the WML.
#
# **Note:** You can also use conda environment configuration file `yaml` as package extension input. In such case set the `TYPE` to `conda_yml` and `file_path` to yaml file.
# ```
# client.package_extensions.ConfigurationMetaNames.TYPE = "conda_yml"
# ```
# +
meta_prop_pkg_extn = {
client.package_extensions.ConfigurationMetaNames.NAME: "K_Linag_norm_skl",
client.package_extensions.ConfigurationMetaNames.DESCRIPTION: "Pkg extension for custom lib",
client.package_extensions.ConfigurationMetaNames.TYPE: "pip_zip"
}
pkg_extn_details = client.package_extensions.store(meta_props=meta_prop_pkg_extn, file_path="linalgnorm-0.1.zip")
pkg_extn_uid = client.package_extensions.get_uid(pkg_extn_details)
pkg_extn_url = client.package_extensions.get_href(pkg_extn_details)
# -
# Display the details of the package extension resource that was created in the above cell.
details = client.package_extensions.get_details(pkg_extn_uid)
# ### Create software specification and add custom library
# Define the meta data required to create software spec resource and bind the package. This software spec resource will be used to configure the online deployment runtime environment for a model.
client.software_specifications.ConfigurationMetaNames.show()
# #### List base software specifications
client.software_specifications.list()
# #### Select base software specification to extend
base_sw_spec_uid = client.software_specifications.get_uid_by_name("default_py3.8")
# #### Define new software specification based on base one and custom library
# +
meta_prop_sw_spec = {
client.software_specifications.ConfigurationMetaNames.NAME: "linalgnorm-0.1",
client.software_specifications.ConfigurationMetaNames.DESCRIPTION: "Software specification for linalgnorm-0.1",
client.software_specifications.ConfigurationMetaNames.BASE_SOFTWARE_SPECIFICATION: {"guid": base_sw_spec_uid}
}
sw_spec_details = client.software_specifications.store(meta_props=meta_prop_sw_spec)
sw_spec_uid = client.software_specifications.get_uid(sw_spec_details)
client.software_specifications.add_package_extension(sw_spec_uid, pkg_extn_uid)
# -
# ### Save the model
# Define the metadata to save the trained model to WML Repository along with the information about the software spec resource required for the model.
#
# The `client.repository.ModelMetaNames.SOFTWARE_SPEC_UID` metadata property is used to specify the GUID of the software spec resource that needs to be associated with the model.
# +
model_props = {
client.repository.ModelMetaNames.NAME: "Temp prediction model with custom lib",
client.repository.ModelMetaNames.TYPE: 'scikit-learn_0.23',
client.repository.ModelMetaNames.SOFTWARE_SPEC_UID: sw_spec_uid
}
# -
# Save the model to the WML Repository and display its saved metadata.
published_model = client.repository.store_model(model=skl_pipeline, meta_props=model_props)
published_model_uid = client.repository.get_model_uid(published_model)
model_details = client.repository.get_details(published_model_uid)
print(json.dumps(model_details, indent=2))
# <a id="deploy"></a>
#
# ## 6 Deploy and Score
#
# In this section, you will deploy the saved model that uses the custom transformer and perform predictions. You will use WML client to perform these tasks.
# ### Deploy the model
# +
metadata = {
client.deployments.ConfigurationMetaNames.NAME: "Deployment of custom lib model",
client.deployments.ConfigurationMetaNames.ONLINE: {}
}
created_deployment = client.deployments.create(published_model_uid, meta_props=metadata)
# -
# <a id="score"></a>
# ### Predict using the deployed model
# **Note**: Here we use deployment `uid` saved in published_model object. In next section, we show how to retrive deployment url from Watson Machine Learning instance.
deployment_uid = client.deployments.get_uid(created_deployment)
# Now you can print an online scoring endpoint.
scoring_endpoint = client.deployments.get_scoring_href(created_deployment)
print(scoring_endpoint)
# Prepare the payload for prediction. The payload contains the input records for which predictions has to be performed.
scoring_payload = {
"input_data": [{
'fields': ["time", "humidity"],
'values': [[79863, 47]]}]
}
# Execute the method to perform online predictions and display the prediction results
predictions = client.deployments.score(deployment_uid, scoring_payload)
print(json.dumps(predictions, indent=2))
# <a id="cleanup"></a>
# ## 7. Clean up
# If you want to clean up all created assets:
# - experiments
# - trainings
# - pipelines
# - model definitions
# - models
# - functions
# - deployments
#
# please follow up this sample [notebook](https://github.com/IBM/watson-machine-learning-samples/blob/master/cpd4.0/notebooks/python_sdk/instance-management/Machine%20Learning%20artifacts%20management.ipynb).
# <a id="summary"></a>
#
# ## 8. Summary
#
# You successfully completed this notebook!
#
# You learned how to use a scikit-learn model with custom transformer in Watson Machine Learning service to deploy and score.
#
# Check out our [Online Documentation](https://dataplatform.cloud.ibm.com/docs/content/analyze-data/wml-setup.html) for more samples, tutorials, documentation, how-tos, and blog posts.
# ## Author
#
# **<NAME>**, is a senior technical lead in IBM Watson Machine Learning team. Krishna works on developing cloud services that caters to different stages of machine learning and deep learning modeling life cycle.
#
# **<NAME>**, PhD, is a Software Architect and Data Scientist at IBM.
# Copyright © 2020, 2021, 2022 IBM. This notebook and its source code are released under the terms of the MIT License.
|
cpd4.0/notebooks/python_sdk/deployments/custom_library/Use scikit-learn and custom library to predict temperature.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Xg0MysBbFf0X" outputId="c914049d-6006-47a9-bbd7-d7cabf3b9fd7"
# !pip install pyLDAvis
# + colab={"base_uri": "https://localhost:8080/"} id="BtxmA63EXVjW" outputId="cca06c41-c078-4ee4-a6d6-97f9d7e9a89d"
from google.colab import drive
drive.mount('/content/gdrive')
# + colab={"base_uri": "https://localhost:8080/"} id="KZemYpfPYb2x" outputId="19251319-881c-4257-a395-f3fc22c34467"
# !ls "/content/gdrive/My Drive/MLOps Hackathon"
# + colab={"base_uri": "https://localhost:8080/"} id="5FbBLDmBh4LD" outputId="75bf045d-14dc-4778-fc43-859e403f6af8"
# !pip install -U nltk
# + id="8lLMo3VqYoo7"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# + id="G-mA0f5naPNu"
from os import path
from PIL import Image
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
# + id="dSyvx0v1YrxE"
#Load the dataset
data = pd.read_csv("/content/gdrive/My Drive/MLOps Hackathon/dm2_data.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="WJTfsZ_vYsRO" outputId="13458476-5ebe-46c6-d158-0bd2a0b54984"
data.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 137} id="VT_xscdcbbBi" outputId="05fee652-d1dc-447c-9f6b-43faeab74b02"
' '.join(data['paragraphs'][0].split(' '))
# + colab={"base_uri": "https://localhost:8080/"} id="ufdfH9YHZngl" outputId="c43f7478-e706-4a11-ea8f-c3f91d28f1f1"
data.describe
# + id="kJpocEgDaBx2"
# #?WordCloud
# + id="KVWFbKfRZk8f"
text = ','.join(list(data['paragraphs'].values))
# + id="Lf0xrm6LcasH"
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(text)
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="1-SUPGj1cgDw" outputId="2d925243-b6c9-46d2-86c9-f9666b7b36c3"
# Display the generated image:
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="3LGhkPuriNBL" outputId="62323c50-cc99-43f2-9c61-ee8158d3e267"
import nltk
nltk.download('punkt')
# + id="TnZR06rQhprF"
from nltk.tokenize import word_tokenize
# + id="DeznUCK1hwPr"
tokens = word_tokenize(text)
# convert to lower case
tokens = [w.lower() for w in tokens]
# + colab={"base_uri": "https://localhost:8080/"} id="OXOUezNwiQqh" outputId="62d45cdb-7e88-45aa-b0da-32cedcebf6b0"
tokens[:10]
# + id="ki5A9EmXhuMC"
# remove punctuation from each word
import string
table = str.maketrans('', '', string.punctuation)
stripped = [w.translate(table) for w in tokens]
# remove remaining tokens that are not alphabetic
words = [word for word in stripped if word.isalpha()]
# + colab={"base_uri": "https://localhost:8080/"} id="DwYopRlXibtL" outputId="11d2dfe1-be5b-4c79-ee3b-02a500978ad0"
nltk.download('stopwords')
# + id="7GvoA9YeiWJq"
from nltk.corpus import stopwords
# + colab={"base_uri": "https://localhost:8080/"} id="RbSiYAwuiYty" outputId="eae52105-f542-4633-8b21-97a91e4427f6"
stop_words = set(stopwords.words('english'))
stop_words.update(['type','increased','patient','study','patients','et','al','https','use'])
words = [w for w in words if not w in stop_words]
print(words[:100])
# + id="3N6GB5sviqvJ"
word_string = ' '.join(words)
# + id="seDIOOMDimje"
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(word_string)
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="_BEEdUmXi0iJ" outputId="b7a9c061-04b4-4b44-c968-37f79fff4b8d"
# Display the generated image:
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="MTBwPvwA9FrM" outputId="a1168e8e-0327-4682-a167-4796476e7167"
import gensim
from gensim.utils import simple_preprocess
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
stop_words.extend(['from', 'subject', 're', 'edu', 'use','https','www','doi','org','et','al','xa','dm','com','dpp','ref'])
def sent_to_words(sentences):
for sentence in sentences:
# deacc=True removes punctuations
yield(gensim.utils.simple_preprocess(str(sentence), deacc=True))
def remove_stopwords(texts):
return [[word for word in simple_preprocess(str(doc))
if word not in stop_words] for doc in texts]
data = data.paragraphs.values.tolist()
data_words = list(sent_to_words(data))
# remove stop words
data_words = remove_stopwords(data_words)
print(data_words[:1][0][:30])
# + colab={"base_uri": "https://localhost:8080/"} id="RWeaYp_uEp0N" outputId="aeba933b-cf57-4bca-d916-b6f078fadeed"
import gensim.corpora as corpora
# Create Dictionary
id2word = corpora.Dictionary(data_words)
# Create Corpus
texts = data_words
# Term Document Frequency
corpus = [id2word.doc2bow(text) for text in texts]
# View
print(corpus[:1][0][:30])
# + colab={"base_uri": "https://localhost:8080/"} id="leWRz4OwEswK" outputId="d0ebc9a0-f8a9-4396-bcc0-9bbde7e416a8"
from pprint import pprint
# number of topics
num_topics = 10
# Build LDA model
lda_model = gensim.models.LdaMulticore(corpus=corpus,
id2word=id2word,
num_topics=num_topics)
# Print the Keyword in the 10 topics
pprint(lda_model.print_topics())
doc_lda = lda_model[corpus]
# + colab={"base_uri": "https://localhost:8080/", "height": 915} id="0m9WCAyIFBy9" outputId="de23a004-4218-4961-982b-7ab333f76f57"
import pyLDAvis
import pyLDAvis.gensim_models as gensimvis
pyLDAvis.enable_notebook()
vis = gensimvis.prepare(lda_model, corpus, id2word)
vis
|
nlp/nlpanalysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Light Actors {#light_actors_example}
# ============
#
# Positional lights in PyVista have customizable beam shapes, see the
# `ref_light_beam_shape_example`{.interpreted-text role="ref"} example.
# Spotlights are special in the sense that they are unidirectional lights
# with a finite position, so they can be visualized using a cone.
#
# This is exactly the purpose of a `vtk.vtkLightActor`, the functionality
# of which can be enabled for spotlights:
#
# +
import numpy as np
import pyvista as pv
from pyvista import examples
cow = examples.download_cow()
cow.rotate_x(90, inplace=True)
plotter = pv.Plotter(lighting='none', window_size=(1000, 1000))
plotter.add_mesh(cow, color='white')
floor = pv.Plane(center=(*cow.center[:2], cow.bounds[-2]), i_size=30, j_size=25)
plotter.add_mesh(floor, color='green')
UFO = pv.Light(position=(0, 0, 10), focal_point=(0, 0, 0), color='white')
UFO.positional = True
UFO.cone_angle = 40
UFO.exponent = 10
UFO.intensity = 3
UFO.show_actor()
plotter.add_light(UFO)
# enable shadows to better demonstrate lighting
plotter.enable_shadows()
plotter.camera_position = [(28, 30, 22), (0.77, 0, -0.44), (0, 0, 1)]
plotter.show()
# -
# Light actors can be very useful when designing complex scenes where
# spotlights are involved in lighting.
#
# +
plotter = pv.Plotter(lighting='none')
plane = pv.Plane(i_size=4, j_size=4)
plotter.add_mesh(plane, color='white')
rot120 = np.array([[-0.5, -np.sqrt(3) / 2, 0], [np.sqrt(3) / 2, -0.5, 0], [0, 0, 1]])
position = (-1.5, -1.5, 3)
focus = (-0.5, -0.5, 0)
colors = ['red', 'lime', 'blue']
for color in colors:
position = rot120 @ position
focus = rot120 @ focus
light = pv.Light(position=position, focal_point=focus, color=color)
light.positional = True
light.cone_angle = 15
light.show_actor()
plotter.add_light(light)
plotter.show()
# -
# One thing to watch out for is that the light actors are represented such
# that their cone has a fixed height. This implies that for very large
# cone angles we typically end up with enormous light actors, in which
# case setting a manual camera position before rendering is usually a good
# idea. Increasing the first example\'s cone angle and omitting the manual
# camera positioning exemplifies the problem:
#
# +
plotter = pv.Plotter(lighting='none')
plotter.add_mesh(cow, color='white')
floor = pv.Plane(center=(*cow.center[:2], cow.bounds[-2]), i_size=30, j_size=25)
plotter.add_mesh(floor, color='green')
UFO = pv.Light(position=(0, 0, 10), focal_point=(0, 0, 0), color='white')
UFO.positional = True
UFO.cone_angle = 89
UFO.exponent = 10
UFO.intensity = 3
UFO.show_actor()
plotter.add_light(UFO)
plotter.show()
|
examples/04-lights/actors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Last Mile Delivery Scheduling Problem
# We will build an optimization model to solve this problem. Let's first write out the math formulation.
#
# <br>
#
# <b>Sets: </b>
#
# Employees: $i \in I=\{1,...,19 \}$
#
# Jobs: $j, k \in \Theta =\{1,...,49 \}$
#
# <br>
#
# <b>Parameters:</b>
#
# $JS_j$: job $j$ starttime
#
# $JE_j$: job $j$ endtime
#
# $SS_i$: time of employee $i$ starts shift
#
# $SE_i$: time of employee $i$ ends shift
#
# <br>
#
# <b>Decision variables:</b>
#
# $
# x_{ij} = \left\{
# \begin{array}\\
# 1 & \text{if job } j \text{ is assigned to employee } i \\
# 0 & otherwise \\
# \end{array}
# \right.
# $
#
# $
# z_{ijk} = \left\{
# \begin{array}\\
# 1 & \text{ job } j \text{ starts before job } k,\text{ done by employee }i \\
# 0 & \text{ job } j \text{ starts after job } k,\text{ done by employee }i \\
# \end{array}
# \right.
# $
#
# The problem can be solved as the following MIP.
#
# \begin{align}
# \text{Maximize:} \quad & \sum_{i \in I} \sum_{j \in \Theta} x_{ij} \\
# \text{Subject to:} \quad & \sum_{j \in \Theta} x_{ij} \leq 1, \quad\qquad\qquad\qquad\qquad\quad \forall j \\
# & x_{ij} \cdot (JS_j - SS_i) \geq 0, \qquad\qquad\qquad \forall i,j \\
# & x_{ij} \cdot (SE_i - JE_j) \geq 0, \qquad\qquad\qquad \forall i,j \\
# & x_{ij} \cdot JS_j \geq x_{ik} \cdot JE_k - M \cdot z_{ijk}, \quad\qquad \forall i,j, k \ and\ j <k\\
# & x_{ik} \cdot JS_k \geq x_{ij} \cdot JE_j - M \cdot (1-z_{ijk}), \quad \forall i,j, k \ and\ j <k\\
# \end{align}
# +
# This routine will need Gurobi to be installed
# conda config --add channels https://conda.anaconda.org/gurobi
# conda install gurobi
# -
import pandas as pd
import numpy as np
import gurobipy as gp
from gurobipy import GRB
import datetime
import matplotlib.pyplot as plt
ev = pd.read_excel('input/data_carvana.xlsx',sheet_name='events')
dur = pd.read_excel('input/data_carvana.xlsx',sheet_name='duration')
res = pd.read_excel('input/data_carvana.xlsx',sheet_name='resources')
res.head()
dur.columns
# Events to be scheduled
ev = ev.drop(['Unnamed: 4', 'Unnamed: 5', 'Unnamed: 6', 'Unnamed: 7'],axis=1)
# Durations
dur = dur.drop(['Unnamed: 3', 'Unnamed: 4', 'Unnamed: 5', 'Unnamed: 6', 'Unnamed: 7', 'Unnamed: 8'], axis=1)
ev.head()
dur.columns = ['OriginAddressId', 'AddressId', 'TripDuration']
# Merge "Events" and "Durations" to be "jobs"
jobs = pd.merge(ev, dur, on='AddressId')
jobs = jobs.drop('OriginAddressId',axis=1)
jobs.head()
# Convert datetime.time format to seconds
def time_to_seconds(time):
return (time.hour * 60 + time.minute) * 60 + time.second
# resource = {resourceID: [ShiftStart, ShiftEnd], ... }
resource = {}
for i in range(len(res.index)):
resource[res['ResourceId'][i]] = [time_to_seconds(res['StartTime'][i]), time_to_seconds(res['EndTime'][i]) ]
# events = {eventID: [AppStart, AppEnd], ... }
events = {}
for i in range(len(ev.index)):
events[jobs['EventId'][i]] = [time_to_seconds(jobs['Appointment Starttime'][i]),
time_to_seconds(jobs['Appointment Endtime'][i]) ]
# Pre-processing data for solving MIP in Gurobi
emp_num, shift_start, shift_end = gp.multidict(resource)
job_num, job_start, job_end = gp.multidict(events)
availability=[]
for i in emp_num:
for j in job_num:
availability.append((i,j))
bigM = 100000 # 1 day = 86400 seconds.
# Model
m = gp.Model("carvana_schedule")
# Decision var: x[emp_num, job_num]
x = m.addVars(availability, vtype = GRB.BINARY, name="x")
# Decision var: z[emp_num, job_num, job_num]
z = m.addVars(emp_num, job_num, job_num, vtype = GRB.BINARY, name="z")
# Objective
m.setObjective(gp.quicksum(x[i] for i in availability), GRB.MAXIMIZE)
# contranit (1): Each job can only be done by no more than ONE employee
m.addConstrs( (x.sum('*',s) <= 1 for s in job_num), "eachjob")
# contranit (2)(3): Each scheduled job must be within that employee's shift hours.
for i in emp_num:
for j in job_num:
m.addConstr( x[i, j] * (job_start[j] - shift_start[i]) >= 0 , "workhour1[%s, %s]"%(i, j) )
m.addConstr( x[i, j] * (job_end[j] - shift_end[i]) <= 0 , "workhour2[%s, %s]"%(i, j) )
# contranit (4)(5): Disjunctive constraints: for employee i, scheduled job j and k cannot overlap.
for i in emp_num:
for j in job_num:
for k in job_num:
if j<k:
#print(i,j,k)
m.addConstr( x[i,j]*job_start[j] >= x[i,k]*job_end[k] - bigM*z[i,j,k] , "disjunctive1[%s, %s, %s]"%(i, j, k) )
m.addConstr( x[i,k]*job_start[k] >= x[i,j]*job_end[j] - bigM*(1-z[i,j,k]) , "disjunctive2[%s, %s, %s]"%(i, j, k) )
# Solve
m.optimize()
# +
# print results
solution = m.getAttr('x',x)
print('Total numbers of events scheduled: %g' % m.objVal)
schedule = pd.DataFrame(columns=['ResourceId','EventId','Appointment Starttime','Appointment Endtime'])
for i in emp_num:
for j in job_num:
if solution[i,j] == 1:
schedule.loc[len(schedule.index)] = [i,j, ev.at[ev[ev['EventId'] == j].index[0], 'Appointment Starttime'], ev.at[ev[ev['EventId'] == j].index[0], 'Appointment Endtime']]
#print('Employee %s takes event %s' % (i, j))
schedule.to_excel("LastMileDeliverySchedule_XH.xlsx")
schedule.head()
# -
employees = sorted(list(schedule['ResourceId'].unique()))
# Dictionary: emp_sch = {ResourceId1: [(AppStart1, AppDur1), (AppStart2, AppDur2), (AppStart3, AppDur3)... ],
# ResourceId2: [(), ()],
# ... }
emp_sch = {}
for i in employees:
emp_sch[i] = []
for st in list( schedule[schedule['ResourceId'] == i]['Appointment Starttime']):
emp_sch[i].append( (time_to_seconds(st), time_to_seconds(schedule.at[schedule[schedule['Appointment Starttime'] == st].index[0], 'Appointment Endtime'])- time_to_seconds(st) ) )
# +
fig, gnt = plt.subplots()
fig.set_size_inches(18.5, 10.5)
gnt.set_ylim(0, 10+10*len(employees))
gnt.set_xlim(30000, 70000)
plt.title("Carvana Last Mile Delivery Schedule")
gnt.set_xlabel('Time')
gnt.set_ylabel('EmployeeID')
# Setting x axis
gnt.set_xticks( list(range(25200,79200,3600)) ) # 7:00 - 22:00
gnt.set_xticklabels(['7:00', '8:00', '9:00', '10:00', '11:00', '12:00', '13:00', '14:00',
'15:00', '16:00', '17:00', '18:00', '19:00', '20:00', '21:00'])
# Setting y axis
gnt.set_yticks( list(range(0, 10*len(employees)+20, 10 )) )
emp_labels = employees.copy()
emp_labels.insert(0,'')
emp_labels.append('')
gnt.set_yticklabels(emp_labels)
gnt.grid(True, color = 'tab:olive', linestyle = ':', linewidth = 0.5)
for h, i in enumerate(emp_sch):
gnt.broken_barh(emp_sch[i], (5+10*h, 9), facecolors =('green'))
plt.savefig("CarvanaSchedule.png")
# -
|
LMD_V0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="H0k6MdAyNNYG" outputId="546e85ba-0400-40eb-fd5d-2b44d2f38fa2"
import tensorflow as tf
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import os
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem import SnowballStemmer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
import re
# + id="WwlS13ULNtCp"
data = pd.read_csv('/content/drive/MyDrive/Do_an-NCKH/Dataset/spam_ham_dataset.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="1dzrxapLN6TC" outputId="262710b9-d3e2-48a3-8c8d-17e175d485fc"
data
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="t2X_mFU3N7lg" outputId="6b2cf958-a9f6-47d0-df8c-fcbb24891587"
data = data.drop(['Unnamed: 0', 'label'], axis=1)
data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="g9eFqfVMOAgG" outputId="b79b668e-5b12-4043-dd84-39be2615678a"
val_count = data.label_num.value_counts()
plt.figure(figsize=(8,4))
plt.bar(val_count.index, val_count.values)
plt.title("Spam/ham Data Distribution")
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="qGeJIzDwOEqD" outputId="00956ab7-b518-4faf-a4dd-804c91e38a7a"
stop_words = stopwords.words('english')
stemmer = SnowballStemmer('english')
text_cleaning_re = "@\S+|https?:\S+|http?:\S+|[^A-Za-z0-9]:\S+|subject:\S+|nbsp"
data.head()
# + id="soefeHw6OHW5"
def preprocess(text, stem=False):
text = re.sub(text_cleaning_re, ' ', str(text).lower()).strip()
tokens = []
for token in text.split():
if token not in stop_words:
if stem:
tokens.append(stemmer.stem(token))
else:
tokens.append(token)
return " ".join(tokens)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="gI7RNBIJONaM" outputId="b8438b29-43aa-41cb-d6ec-b8d0ca595157"
data.text = data.text.apply(lambda x: preprocess(x))
data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="z-rfJbEVORjP" outputId="83af6973-2044-4daf-c53b-a5d44b572500"
TRAIN_SIZE = 0.8
MAX_NB_WORDS = 100000
MAX_SEQUENCE_LENGTH = 50
x = data['text']
y = data['label_num']
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=1-TRAIN_SIZE,
random_state=7) # Splits Dataset into Training and Testing set
print("Train Data size:", len(x_train))
print("Test Data size", len(x_test))
# + id="gZly6A67OYrK"
# + [markdown] id="ibNa09gGOd5l"
# # TOKENIZER
# + colab={"base_uri": "https://localhost:8080/"} id="BRN8UIA1Oems" outputId="4ad0b380-81a0-4637-a111-5790d13e08df"
from keras.preprocessing.text import Tokenizer
tokenizer = Tokenizer()
tokenizer.fit_on_texts(x_train)
word_index = tokenizer.word_index
vocab_size = len(tokenizer.word_index) + 1000
print("Vocabulary Size :", vocab_size)
# + colab={"base_uri": "https://localhost:8080/"} id="f8KZstxdOgVj" outputId="686bdac9-add2-41b4-eb75-c023f228a8e4"
from keras.preprocessing.sequence import pad_sequences
x_train = pad_sequences(tokenizer.texts_to_sequences(x_train),
maxlen = MAX_SEQUENCE_LENGTH)
x_test = pad_sequences(tokenizer.texts_to_sequences(x_test),
maxlen = MAX_SEQUENCE_LENGTH)
print("Training X Shape:",x_train.shape)
print("Testing X Shape:",x_test.shape)
# + id="E_JIABG4OiEZ"
# + [markdown] id="1WJmEdAWOl40"
# # LSTM
# + id="sBkquL77OmWO"
#LSTM hyperparameters
n_lstm = 200
drop_lstm =0.2
# + id="fX807V3DOqsM"
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Embedding, GlobalAveragePooling1D, Dense, Dropout, LSTM, Bidirectional
# + id="TpV-AJkBOrOA"
embeding_dim = 16
# + id="fUSczIiwOxkX"
model2 = Sequential()
model2.add(Embedding(vocab_size, embeding_dim, input_length=MAX_SEQUENCE_LENGTH))
model2.add(Bidirectional(LSTM(n_lstm, dropout=drop_lstm, return_sequences=True)))
model2.add(Dense(1, activation='sigmoid'))
model2.compile(loss = 'binary_crossentropy', optimizer = 'adam', metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="Rc8LFQ9aO3WL" outputId="460340de-498e-404c-e2b5-e946b3d98c7f"
# Training
num_epochs = 30
early_stop = EarlyStopping(monitor='val_loss', patience=2)
history = model2.fit(x_train, y_train, epochs=num_epochs,
validation_data=(x_test, y_test),callbacks =[early_stop], verbose=2)
# + colab={"base_uri": "https://localhost:8080/", "height": 573} id="5qnpkmWPO6H2" outputId="71053187-2bcd-4ebd-95f6-dbf10dfa204f"
# Create a dataframe
metrics = pd.DataFrame(history.history)
# Rename column
metrics.rename(columns = {'loss': 'Training_Loss', 'accuracy': 'Training_Accuracy',
'val_loss': 'Validation_Loss', 'val_accuracy': 'Validation_Accuracy'}, inplace = True)
def plot_graphs1(var1, var2, string):
metrics[[var1, var2]].plot()
plt.title('BiLSTM Model: Training and Validation ' + string)
plt.xlabel ('Number of epochs')
plt.ylabel(string)
plt.legend([var1, var2])
# Plot
plot_graphs1('Training_Loss', 'Validation_Loss', 'loss')
plot_graphs1('Training_Accuracy', 'Validation_Accuracy', 'accuracy')
# + id="c7dacICIO973"
# + [markdown] id="S7Tw8D6gPLDd"
# # PPDL
# + id="ZNi8n3BUPMdy"
model1 = Sequential()
model1.add(model2.layers[0])
# + id="SbcIzaADPQY0"
x_input = x_test[0:5]
y_model = model1.predict(x_input) # Dùng để so sánh kết quả
# + [markdown] id="b0Myv5gQPVJv"
# ## Server
# + id="o51fwiEUPSwD"
model = Sequential()
for layer in model2.layers[1:]:
model.add(layer)
model.build(input_shape = model2.layers[0].output_shape)
# + colab={"base_uri": "https://localhost:8080/"} id="7KyapxGVPc35" outputId="6468a815-1190-4df9-cfc3-1b874c30f6d6"
lstW = model2.layers[0].get_weights()[0] #Lấy weight của layer 0
lstW.shape
# + id="SJXlYtLRPfmY"
k = np.random.rand(2,2).astype(np.float32)
kd = np.linalg.inv(k)
# + id="H4GahNcsPllR"
lk = []
for i in range(lstW.shape[0]):
wi = lstW[i,:]
w_plus = np.random.uniform(-1, 1, len(wi)).astype(np.float32)
w_ = np.array([wi, w_plus])
lk.append(k.dot(w_))
#Tạo nhiễu và nhân với khóa K để khi gửi về Client không bị lộ Weight
LW = tf.convert_to_tensor(lk)
# + colab={"base_uri": "https://localhost:8080/"} id="ykjEudKEPmYm" outputId="968d6157-40ff-4e52-a915-0739071f486c"
LW.shape
# + [markdown] id="kVDZ97EjPqu6"
# ## Client
# + id="XZf_rdTKPoJb"
X = np.zeros((x_input.shape[0], x_input.shape[1], 2))
for i in range(X.shape[0]):
X_ = []
for j in x_input[i]:
x_ = np.random.randint(0, LW.shape[0])
xx = [j, x_]
X_.append(xx)
j += 1
X[i] = X_
#Tạo nhiễu cho Input
# + id="RBhuAEM7P7RC"
X_one_hot = tf.one_hot(X, LW.shape[0])
Y0 = X_one_hot@LW[:,0]
Y1 = X_one_hot@LW[:,1]
# + id="z30-SgLpQBhA"
Y = np.zeros_like(Y0)
for i in range(Y0.shape[0]):
for j in range(Y0.shape[1]):
Y[i,j,0] = Y0[i,j,0]
Y[i,j,1] = Y1[i,j,0]
Y = tf.convert_to_tensor(Y)
# + colab={"base_uri": "https://localhost:8080/"} id="UnS3o0jcQEw0" outputId="c8d9206e-829c-4850-bd95-6a1464de7c2b"
Y.shape
# + [markdown] id="ahPvKYZBQJDz"
# ## Serer
# + id="-ftTRVodQGha"
lstY = []
for i in range(Y.shape[0]):
lst = []
for j in range(Y.shape[1]):
y_ = kd@Y[i,j]
lst.append(y_[0])
lstY.append(lst)
lstY = tf.convert_to_tensor(lstY)
# + colab={"base_uri": "https://localhost:8080/"} id="S84KcRaDQO3X" outputId="7de4f7a0-9b16-4a85-e0f2-9b1a8f57193f"
model.predict(lstY)[0]
# + colab={"base_uri": "https://localhost:8080/"} id="pdCSoI4PQUL2" outputId="68f8bd2e-0404-41f0-dd52-6c4566420768"
model2.predict(x_input)[0]
# + id="ZiOS78q9QXjp"
#Check xong, 2 kết qủa dự đoán đều giông nhau.
# + id="fUF5Fy5PQbMG"
|
PPDL_NLP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HzSorCPNMpPX"
# **Note:** *Change to GPU runtime for faster execution*
# + [markdown] id="ozZOFLgRM-AM"
# # First, mount the drive to get access to dataset
# + colab={"base_uri": "https://localhost:8080/"} id="PQ-j4b23PLqs" executionInfo={"status": "ok", "timestamp": 1615953723787, "user_tz": 360, "elapsed": 302, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="0912c6d2-c897-41a3-9660-d6a60e346114"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="i9evyTXrM_qJ"
# # Load the dataset from drive
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="HzL3mY2RPUub" executionInfo={"status": "ok", "timestamp": 1615953725362, "user_tz": 360, "elapsed": 1867, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="51e712d2-36da-4821-b796-6045cccfa728"
import os
import pandas as pd
# Specify the path to dataset and the file name
DATASET_DIR = os.path.join(os.getcwd(), r'drive/MyDrive/NLP Final Project/datasets/sentiment140_kaggle')
FILE = os.path.join(DATASET_DIR, r'Sentiment140.annotated.resampled.100000.csv')
# Open the csv file as a pandas DataFrame with specified columns
tweets_df = pd.read_csv(FILE, sep=',', encoding='utf-8')
tweets_df
# + [markdown] id="SVban3lZNCPT"
# # Cleaning the raw tweets
# + colab={"base_uri": "https://localhost:8080/"} id="m3IJE08rPZV6" executionInfo={"status": "ok", "timestamp": 1615953725890, "user_tz": 360, "elapsed": 2388, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="6c093d80-10ed-4503-ee55-ee5253125183"
import re
import nltk
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
nltk.download('punkt')
nltk.download('stopwords')
# + [markdown] id="0Zql-pjLNQsl"
# > Remove the word 'not' from stopwords as it specifies a negative emotion
# + id="oaD6O8yHPgNQ" executionInfo={"status": "ok", "timestamp": 1615953725890, "user_tz": 360, "elapsed": 2383, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
all_stopwords = stopwords.words('english')
all_stopwords.remove('not')
def clean(tweet):
# Remove @usermentions (for a tweet)
tweet = re.sub('@[^\s]+',' ',tweet)
# Remove urls (for a tweet)
tweet = re.sub('((www\.[^\s]+)|(https?://[^\s]+))',' ',tweet)
# Replace #word with word (for a tweet)
tweet = re.sub(r'#([^\s]+)', r'\1', tweet)
# Remove all remaining punctuations
tweet = re.sub(r'[^a-zA-Z]', ' ', tweet)
# Convert tweet to lowercase
tweet = tweet.lower()
# Remove stop words
tweet = tweet.split()
tweet = ' '.join([word for word in tweet if not word in set(all_stopwords)])
return tweet
# + [markdown] id="kLXZ9l9sNT_P"
# > Apply to the `clean` function to DataFrame
# + colab={"base_uri": "https://localhost:8080/"} id="GamYbgQTPqBh" executionInfo={"status": "ok", "timestamp": 1615953771716, "user_tz": 360, "elapsed": 48204, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="63da2c46-e6b1-4e55-8498-f276b7162a17"
# %%time
tweets_df['cleaned_tweet'] = tweets_df['tweet'].apply(clean)
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="2g7z3nlXPtZ9" executionInfo={"status": "ok", "timestamp": 1615953771717, "user_tz": 360, "elapsed": 48198, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="962f5ac6-6b13-442f-f908-94110764e6d2"
tweets_df
# + [markdown] id="6eB56ryuNWDT"
# # Split the data into train set and test set
# + id="0qqH2hfUPx47" executionInfo={"status": "ok", "timestamp": 1615953772102, "user_tz": 360, "elapsed": 48577, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
from sklearn.model_selection import train_test_split
X = tweets_df.cleaned_tweet.values
y = tweets_df.label.values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# + [markdown] id="HyckSuumNhZJ"
# # Specify the hyper parameters
# + id="ERdBtEcHP1N_" executionInfo={"status": "ok", "timestamp": 1615953772103, "user_tz": 360, "elapsed": 48574, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
MAX_VOCAB_SIZE = 20000
MAX_LEN = 1000
EMBEDDING_DIM = int(round(MAX_VOCAB_SIZE ** 0.25))
# + [markdown] id="4pDVFpuqNv6A"
# # Prepare the data for training
# + [markdown] id="WGh9EPtYN2ZO"
# > **Tokenize**
# + colab={"base_uri": "https://localhost:8080/"} id="JGidjpW6QIys" executionInfo={"status": "ok", "timestamp": 1615953782770, "user_tz": 360, "elapsed": 59234, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="31f69a27-e923-48ab-e60e-fc03967381c3"
# %%time
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
tokenizer = Tokenizer(num_words=MAX_VOCAB_SIZE)
tokenizer.fit_on_texts(X_train)
# + [markdown] id="FpL45yPEN628"
# > **Convert to sequences**
# + colab={"base_uri": "https://localhost:8080/"} id="GIYNJVhKQPAS" executionInfo={"status": "ok", "timestamp": 1615953793325, "user_tz": 360, "elapsed": 69783, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="c0b810ab-88d3-40c7-bd4e-888a4327a959"
# %%time
X_train = tokenizer.texts_to_sequences(X_train)
X_test = tokenizer.texts_to_sequences(X_test)
# + [markdown] id="p6m2Jb18OL6s"
# > **Pad the sequences to convert all inputs to same shape**
# + colab={"base_uri": "https://localhost:8080/"} id="-5DmT-lbQUhy" executionInfo={"status": "ok", "timestamp": 1615953797920, "user_tz": 360, "elapsed": 74371, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="96b55392-68b8-4d70-d7e2-ec864062a837"
# %%time
from tensorflow.keras.preprocessing.sequence import pad_sequences
X_train = pad_sequences(X_train, maxlen=MAX_LEN)
X_test = pad_sequences(X_test, maxlen=MAX_LEN)
# + [markdown] id="DU4AgyDSOjIe"
# > **Label encode the 10 classes and convert them to one-hot encodings**
# + id="MjNJxO68QZWz" executionInfo={"status": "ok", "timestamp": 1615953798237, "user_tz": 360, "elapsed": 74683, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
from sklearn.preprocessing import LabelEncoder
from keras.utils.np_utils import to_categorical
encoder = LabelEncoder()
transformed_labels = encoder.fit_transform(tweets_df.label.values)
y_train = encoder.transform(y_train)
y_test = encoder.transform(y_test)
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
# + [markdown] id="wseVYX6UOv_c"
# # CNN model
# + id="ZbcHadOlQqQ5" executionInfo={"status": "ok", "timestamp": 1615953798239, "user_tz": 360, "elapsed": 74680, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Flatten
from tensorflow.keras.layers import Embedding
from tensorflow.keras.layers import Conv1D, MaxPooling1D
# + [markdown] id="9j1AcJrfQxTl"
# > **Specify model parameters**
# + id="nmDDekQQQ2Zk" executionInfo={"status": "ok", "timestamp": 1615953798239, "user_tz": 360, "elapsed": 74676, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
FILTERS = 32
KERNEL_SIZE = 3
EPOCHS = 5
BATCH_SIZE = 32
# + [markdown] id="keKa_caiO2oE"
# > **Create a model**
# + colab={"base_uri": "https://localhost:8080/"} id="tvw13qb2QuM4" executionInfo={"status": "ok", "timestamp": 1615953798897, "user_tz": 360, "elapsed": 75329, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="fd739bdd-602f-44d8-dfbd-c516862f6114"
model = Sequential([
Embedding(MAX_VOCAB_SIZE, EMBEDDING_DIM, input_length=MAX_LEN),
Conv1D(FILTERS, KERNEL_SIZE, padding='valid', activation='relu'),
MaxPooling1D(),
Flatten(),
Dense(512, activation='relu'),
Dense(128, activation='relu'),
Dense(10, activation='softmax'),
])
model.summary()
# + [markdown] id="I1_tK2C7O5V7"
# > **Compile the model**
# + id="FxqDsBdKRUyJ" executionInfo={"status": "ok", "timestamp": 1615953798898, "user_tz": 360, "elapsed": 75325, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# + [markdown] id="AkpiUxWPTfXn"
# > **Implement callback to stop training on 95% accuracy**
# + id="wXKVMPtoTdO2" executionInfo={"status": "ok", "timestamp": 1615953798898, "user_tz": 360, "elapsed": 75320, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
# Implement callback function to stop training
# when accuracy reaches ACCURACY_THRESHOLD
ACCURACY_THRESHOLD = 0.95
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if(logs.get('accuracy') is not None and logs.get('accuracy') >= ACCURACY_THRESHOLD):
print("\nReached %2.2f%% accuracy, so stopping training!!" %(ACCURACY_THRESHOLD*100))
self.model.stop_training = True
# Instantiate a callback object
callbacks = myCallback()
# + [markdown] id="5h8X93PTO7Lj"
# > **Fit the model**
# + colab={"base_uri": "https://localhost:8080/"} id="j4Z6-NWpRYE3" executionInfo={"status": "ok", "timestamp": 1615954984713, "user_tz": 360, "elapsed": 1261125, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="64600a68-3752-4176-bdbb-5c195eeb65cf"
model.fit(X_train, y_train, batch_size=BATCH_SIZE, epochs=EPOCHS, callbacks=[callbacks])
# + [markdown] id="5UUkqmCdRxGO"
# > **Predict on test data**
# + id="pHYDPD5ARf-f" executionInfo={"status": "ok", "timestamp": 1615954995001, "user_tz": 360, "elapsed": 1271408, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}}
y_pred = model.predict(X_test)
# + [markdown] id="qPdI1w0dSQtb"
# # Classification report
# + colab={"base_uri": "https://localhost:8080/"} id="R2S93dNgSScS" executionInfo={"status": "ok", "timestamp": 1615955409159, "user_tz": 360, "elapsed": 611, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="22cf1154-87c4-4c07-91f2-c6beb2e19cf8"
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score
import numpy as np
print("Report:\n")
average = 'macro'
print(f"Accuracy score : {accuracy_score(np.argmax(y_test, axis=1), np.argmax(y_pred, axis=1))}")
print(f"Precision score : {precision_score(np.argmax(y_test, axis=1), np.argmax(y_pred, axis=1), average=average)}")
print(f"Recall score : {recall_score(np.argmax(y_test, axis=1), np.argmax(y_pred, axis=1), average=average)}")
print(f"F1 score : {f1_score(np.argmax(y_test, axis=1), np.argmax(y_pred, axis=1), average=average)}")
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="ykQXVsxkSkCp" executionInfo={"status": "ok", "timestamp": 1615955449874, "user_tz": 360, "elapsed": 1136, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16523504641951076127"}} outputId="2bba2766-896e-4e05-f676-f3ca91dbf1fa"
from sklearn.metrics import confusion_matrix
import seaborn as sns
import matplotlib.pyplot as plt
cm = confusion_matrix(np.argmax(y_test, axis=1), np.argmax(y_pred, axis=1))
plt.figure(figsize=(20, 5))
sns.heatmap(cm, fmt="d", annot=True, cmap='Blues', xticklabels=list(set(y)), yticklabels=list(set(y)))
|
Simple CNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Keras: Tabular Classify Binary
# 
# Reference this great blog for machine learning cookbooks: [MachineLearningMastery.com "Binary Classification"](https://machinelearningmastery.com/binary-classification-tutorial-with-the-keras-deep-learning-library/).
# +
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.callbacks import History
from sklearn.preprocessing import LabelBinarizer, PowerTransformer
import aiqc
from aiqc import datum
# -
# ---
# ## Example Data
# Reference [Example Datasets](example_datasets.ipynb) for more information.
df = datum.to_pandas('sonar.csv')
df.head()
# ---
# ## a) High-Level API
# Reference [High-Level API Docs](api_high_level.ipynb) for more information including how to work with non-tabular data.
splitset = aiqc.Pipeline.Tabular.make(
df_or_path = df
, dtype = None
, feature_cols_excluded = 'object'
, feature_interpolaters = None
, feature_window = None
, feature_encoders = dict(
sklearn_preprocess = PowerTransformer(method='yeo-johnson', copy=False)
, dtypes = ['float64']
)
, feature_reshape_indices = None
, label_column = 'object'
, label_interpolater = None
, label_encoder = dict(sklearn_preprocess = LabelBinarizer(sparse_output=False))
, size_test = 0.12
, size_validation = 0.22
, fold_count = None
, bin_count = None
)
def fn_build(features_shape, label_shape, **hp):
model = Sequential(name='Sonar')
model.add(Dense(hp['neuron_count'], activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(0.30))
model.add(Dense(hp['neuron_count'], activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(0.30))
model.add(Dense(hp['neuron_count'], activation='relu', kernel_initializer='he_uniform'))
model.add(Dense(units=label_shape[0], activation='sigmoid', kernel_initializer='glorot_uniform'))
return model
def fn_train(model, loser, optimizer, samples_train, samples_evaluate, **hp):
model.compile(
loss=loser
, optimizer=optimizer
, metrics=['accuracy']
)
model.fit(
samples_train['features'], samples_train['labels']
, validation_data = (samples_evaluate['features'], samples_evaluate['labels'])
, verbose = 0
, batch_size = 3
, epochs = hp['epochs']
, callbacks = [History()]
)
return model
hyperparameters = {
"neuron_count": [25, 50]
, "epochs": [75, 150]
}
queue = aiqc.Experiment.make(
library = "keras"
, analysis_type = "classification_binary"
, fn_build = fn_build
, fn_train = fn_train
, splitset_id = splitset.id
, repeat_count = 2
, hide_test = False
, hyperparameters = hyperparameters
, fn_lose = None #automated
, fn_optimize = None #automated
, fn_predict = None #automated
, foldset_id = None
)
queue.run_jobs()
# For more information on visualization of performance metrics, reference the [Visualization & Metrics](visualization.html) documentation.
# ---
# ## b) Low-Level API
# Reference [Low-Level API Docs](api_low_level.ipynb) for more information including how to work with non-tabular data and defining optimizers.
dataset = aiqc.Dataset.Tabular.from_pandas(df)
label_column = 'object'
label = dataset.make_label(columns=[label_column])
labelcoder = label.make_labelcoder(sklearn_preprocess = LabelBinarizer(sparse_output=False))
feature = dataset.make_feature(exclude_columns=[label_column])
encoderset = feature.make_encoderset()
featurecoder_0 = encoderset.make_featurecoder(
sklearn_preprocess = PowerTransformer(method='yeo-johnson', copy=False)
, dtypes = ['float64']
)
splitset = aiqc.Splitset.make(
feature_ids = [feature.id]
, label_id = label.id
, size_test = 0.22
, size_validation = 0.12
)
def fn_build(features_shape, label_shape, **hp):
model = Sequential(name='Sonar')
model.add(Dense(hp['neuron_count'], activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(0.30))
model.add(Dense(hp['neuron_count'], activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(0.30))
model.add(Dense(hp['neuron_count'], activation='relu', kernel_initializer='he_uniform'))
model.add(Dense(units=label_shape[0], activation='sigmoid', kernel_initializer='glorot_uniform'))
return model
def fn_train(model, loser, optimizer, samples_train, samples_evaluate, **hp):
model.compile(
loss=loser
, optimizer=optimizer
, metrics=['accuracy']
)
model.fit(
samples_train['features'], samples_train['labels']
, validation_data = (samples_evaluate['features'], samples_evaluate['labels'])
, verbose = 0
, batch_size = 3
, epochs = hp['epochs']
, callbacks = [History()]
)
return model
algorithm = aiqc.Algorithm.make(
library = "keras"
, analysis_type = "classification_binary"
, fn_build = fn_build
, fn_train = fn_train
)
hyperparameters = {
"neuron_count": [25, 50]
, "epochs": [75, 150]
}
hyperparamset = algorithm.make_hyperparamset(
hyperparameters = hyperparameters
)
queue = algorithm.make_queue(
splitset_id = splitset.id
, hyperparamset_id = hyperparamset.id
, repeat_count = 2
)
queue.run_jobs()
# For more information on visualization of performance metrics, reference the [Visualization & Metrics](visualization.html) documentation.
|
docs/_build/html/notebooks/keras_binary_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# File to Load
city_data_load = "data/city_data.csv"
ride_data_load = "data/ride_data.csv"
# Read the City and Ride Data
city_pd = pd.read_csv(city_data_load)
ride_pd = pd.read_csv(ride_data_load)
# Combine the data into a single dataset
merge_table = pd.merge(ride_pd, city_pd, on="city", how="left")
# Display the data table for preview
merge_table.head()
# -
# ## Bubble Plot of Ride Sharing Data
# +
# Obtain the x and y coordinates for each of the three city types
scatterplot_data = pd.DataFrame({
"x_axis": merge_table.groupby("city")["ride_id"].count(),
"y_axis": merge_table.groupby("city")["fare"].mean(),
"bubble_size": (10 * merge_table.groupby("city")["driver_count"].mean())
})
scatterplot_data = pd.merge(scatterplot_data, city_pd, on="city", how="left")
scatterplot_data["color"] = scatterplot_data["type"].replace({
'Urban': 'lightcoral', 'Suburban': 'lightskyblue', 'Rural': 'gold'})
scatter_urban = scatterplot_data.loc[scatterplot_data["type"] == "Urban", :]
scatter_suburban = scatterplot_data.loc[scatterplot_data["type"] == "Suburban", :]
scatter_rural = scatterplot_data.loc[scatterplot_data["type"] == "Rural", :]
# Build the scatter plots for each city types
# urban
plt.scatter(scatter_urban["x_axis"], scatter_urban["y_axis"], marker="o",
facecolors=scatter_urban["color"], edgecolors="black",
s=scatter_urban["bubble_size"], alpha=0.6)
# suburban
plt.scatter(scatter_suburban["x_axis"], scatter_suburban["y_axis"], marker="o",
facecolors=scatter_suburban["color"], edgecolors="black",
s=scatter_suburban["bubble_size"], alpha=0.6)
# rural
plt.scatter(scatter_rural["x_axis"], scatter_rural["y_axis"], marker="o",
facecolors=scatter_rural["color"], edgecolors="black",
s=scatter_rural["bubble_size"], alpha=0.6)
# Incorporate the other graph properties
plt.title("Pyber Ride Sharing Data (2016)", fontsize=14)
plt.xlabel("Total Number of Rides (Per City)", fontsize=11)
plt.ylabel("Average Fare ($)", fontsize=11)
plt.grid()
# Create a legend
city_types = scatterplot_data["type"].unique()
lgnd = plt.legend(city_types, title='City Types')
lgnd.legendHandles[0]._sizes=[40]
lgnd.legendHandles[1]._sizes=[40]
lgnd.legendHandles[2]._sizes=[40]
# Incorporate a text label regarding circle size
text_note = "Note:\nCircle size correlates with driver count per city."
plt.figtext(0.925, 0.6, text_note, fontsize=11)
# Save Figure
plt.savefig("pyber_ride_sharing_data_2016.png", bbox_inches="tight")
# Show plot
plt.show()
# -
# ## Total Fares by City Type
# +
# Calculate Type Percents
total_fares = pd.DataFrame({"fares_by_type": merge_table.groupby("type")["fare"].sum()})
total_fares["colors"] = ["gold", "lightskyblue", "lightcoral"]
# Build Pie Chart
x_axis = np.arange(0, len(city_types))
explode = (0, 0, 0.1)
plt.pie(total_fares["fares_by_type"], explode=explode, labels=total_fares.index,
colors=total_fares["colors"], autopct="%1.1f%%", shadow=True, startangle=155)
plt.title("% of Total Fares by City Type", fontsize=14)
# Save Figure
plt.savefig("total_fares_by_city_type.png")
# Show Figure
plt.show()
# -
# ## Total Rides by City Type
# +
# Calculate Ride Percents
total_rides = pd.DataFrame({"rides_by_type": merge_table.groupby("type")["ride_id"].count()})
total_rides["colors"] = ["gold", "lightskyblue", "lightcoral"]
total_rides
# Build Pie Chart
x_axis = np.arange(0, len(city_types))
explode = (0, 0, 0.1)
plt.pie(total_rides["rides_by_type"], explode=explode, labels=total_rides.index,
colors=total_rides["colors"], autopct="%1.1f%%", shadow=True, startangle=155)
plt.title("% of Total Rides by City Type", fontsize=14)
# Save Figure
plt.savefig("total_rides_by_city_type.png")
# Show Figure
plt.show()
# -
# ## Total Drivers by City Type
# +
# Calculate Driver Percents
total_drivers = pd.DataFrame({"drivers_by_type": city_pd.groupby("type")["driver_count"].sum()})
total_drivers["colors"] = ["gold", "lightskyblue", "lightcoral"]
total_drivers
# Build Pie Charts
x_axis = np.arange(0, len(city_types))
explode = (0, 0, 0.1)
plt.pie(total_drivers["drivers_by_type"], explode=explode, labels=total_drivers.index,
colors=total_drivers["colors"], autopct="%1.1f%%", shadow=True, startangle=160)
plt.title("% of Total Drivers by City Type", fontsize=14)
# Save Figure
plt.savefig("total_drivers_by_city_type.png")
# Show Figure
plt.show()
# -
# ## Written Analysis
# #### - <NAME>
#
# * Fares are inversely proportional to city density. However there is not enough customers in rural areas for this to be a significant benefit for the company. On the other hand, suburbans areas should not be overlooked, making up 30.5% of total fares while being 26.3% of total rides/transactions. If Pyber markets to and gets more usage in suburban cities, this increased market presence can have a good impact on the bottom line as the company makes more money per suburban ride (than urban rides, if fares can stay consistent).
# * This ride sharing app has a huge presense in urban areas. There's more rides and more drivers. By sheer numbers alone (since a company is about making money and urban areas account for 62.7% of fares) this is the best market to focus on even though the average fare is lower than other city types.
# * Since drivers are paid per ride (typically a percentage of the fare plus bonuses for meeting thresholds), the average urban driver takes home less money compared to their rural and suburban counterparts. To maintain market presence in urban areas, perhaps urban drivers should be slightly better incentivized, which can translate to both driver and rider satisfaction / retention.
|
pyber_final_wendychau.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Concentration of the norm for random vectors
# Based on section 3.1 of Vershynin's 'High Dimensional Probability'
#
# <NAME> (<EMAIL>) 2020
#
# Thanks to <NAME> for suggesting some of the projects in section 3.1 of this notebook.
# Imports
import numpy as np
from matplotlib import pyplot as plt
import pandas as pd
from sklearn.preprocessing import StandardScaler
# ## 1. Making random vectors
#
# Recall that random vectors were first discussed in Theorem 0.0.2 of the Appetizer, which was an approximate form of Caratheodory's theorem. An $n$-dimensional random vector $X=(X_1,\dots,X_n)$ is a vector whose components $X_1,\dots,X_n$ are each random variables. Often we consider random vectors whose components are independent and identically distributed (i.i.d.).
#
# In order to work with random vectors we're going to need to create examples of them.
#
# We first do this by using the distributions implemented in numpy.
#
# Later we will use real-world data to produce examples.
# +
print('We first create a random number generator with numpy.\n')
rng = np.random.default_rng()
print('We can use this random number generator to make a random vector with 13 components coming from the standard normal distribution with mean 0 and variance 1.\n')
x = rng.standard_normal(13)
print('We display the components of this vector.')
print(x)
print()
print('We can also make a random vector of dimension 7 whose components are sampled from the normal distribution with mean 2.3 and variance 1.1.\n')
y = rng.normal(2.3,1.1,7)
print('We display the components of this vector.')
print(y)
print()
print('We can sample components for our vectors from other distributions, as well.\n')
print('Here we create and display a random vector of dimension 5 whose components are sampled from the Wald, or Inverse Gaussian, distribution with mean 2.4.')
z = rng.wald(2.4,1,5)
print(z)
# -
# ### 1.1 Projects, exercises, and comments on making random vectors
# Exercise 1.1.1: Create a random vector with 21 components coming from the standard normal distribution. Display the components of your vector to make sure you have indeed done this.
# +
# Your code for exercise 1.1.1.
# -
# Exercise 1.1.2: Create and display a random vector with 11 components coming from the normal distribution with mean -2 and variance 1.3.
# +
# Your code for exercise 1.1.2.
# -
# Exercise 1.1.3: Find out what the second parameter to the Wald distribution in the previous example is by using the documentation.
#
# Documentation of the distributions available in numpy can be found [here](https://numpy.org/doc/stable/reference/random/generator.html).
# +
# Your code for exercise 1.1.3.
# -
# Exercise 1.1.4: Create and display a vector whose components are generated according to another distribution available in numpy.
# +
# Your code for exercise 1.1.4.
# -
# Exercise 1.1.5: Create and display a vector $X=(X_1,\dots,X_{20})$ of dimension 20 whose $i^{\text{th}}$ component is sampled from the normal distribution of mean $i$ with variance $\frac{1}{i}$.
# +
# Your code for exercise 1.1.5.
# -
# ## 2. A class for random vectors
#
# So far our random vectors have been numpy arrays. Let's create a class which represents a random vector and carries some of the methods we would like to use on random vectors.
#
# We create a class which will represent our conception of a random vector.
#
# There is nothing stopping us from defining a laundry-list of functions which operate on numpy vectors and going from there, but using the object-oriented approach helps keep us organized and distinguish what we add from what already exists in numpy.
class RandomVector:
"""
A random vector.
Attributes:
components (numpy.ndarray): The components of the vector.
dim (int): The dimension in which the vector lies.
"""
def __init__(self,dim,distribution=lambda dim: np.random.default_rng().standard_normal(dim)):
"""
Args:
dim (int): The dimension in which the vector lies.
distribution (function): The distribution(s) according to which the vector is generated.
This function should create a numpy array with `dim` many entries.
The list of available distributions in numpy can be found here:
https://numpy.org/doc/stable/reference/random/generator.html
These distributions can be passed in as pure (lambda) functions.
See examples below.
"""
# Demand that `dim` is a natural number.
assert dim>0 and type(dim) is int
self.components = distribution(dim)
# Let the vector keep track of its dimension. (This is the same as its length.)
self.dim = dim
def __repr__(self):
"""
Make it so that printing the vector returns basic information about it.
In order to see the components of a random vector `x` use `print(x.components)`.
Returns:
str: The basic information on the vector.
"""
return 'a {}-dimensional random vector (id: {})'.format(self.dim,id(self))
def mean(self):
"""
Compute the mean of the random vector by taking the mean of its components.
Returns:
numpy.float64: The computed mean of the vector.
"""
return self.components.mean()
def norm_squared(self):
"""
Compute the Euclidean norm squared of the vector, which is the sum of the squares of the entries.
Returns:
numpy.float64: The square of the norm of the vector.
"""
return sum(self.components**2)
# +
print('Examples of the `RandomVector` class.\n')
print('Create a random vector `x` in 10 dimensions.')
x = RandomVector(10)
print()
print('Have `x` give us information about itself.')
print(x)
print()
print('Check the dimension of `x`.')
print(x.dim)
print()
print('View the components of `x`.')
print(x.components)
print()
print('Note that the dimension is the same as the length of the component array.')
print(x.dim)
print(len(x.components))
print()
print('Check the mean of `x` as computed from its components.')
print(x.mean())
print()
print('Create another random vector in 10 dimensions.')
y = RandomVector(10)
print()
print('Note that `x` and `y` are distinct.')
print(x)
print(y)
print()
print('We can also specify the mean and variance of our random vector.')
print('Create a 4-dimensional random vector of mean 7 and variance 1.2.')
z = RandomVector(4,lambda dim: rng.normal(7,1.2,dim))
print()
print('See some basic information on `z`.')
print(z)
print()
print('Show the computed mean of `z`.')
print('Note that this is in general distinct from the specified mean.')
print(z.mean())
print()
print('Compute the norm squared of `z`.')
print(z.norm_squared())
# -
# ### 2.1 Projects, exercises, and comments on a class for random vectors
#
# Now that we have our fancy new class for creating random vectors we can build on it for future projects.
#
#
#
#
#
#
# Exercise 2.1.1: Add a method to RandomVector which computes the variance of the vector directly from its components. (Hint: You may have already written similar code before, which you can reuse.)
# +
# Your code for exercise 2.1.1.
# You can experiment here, but be sure to add your new method to the class defined above and rerun the corresponding cell.
# -
# Exercise 2.1.2: Add a method to RandomVector which computes the $L_p$ norm of the vector for any $p$.
# +
# Your code for exercise 2.1.2.
# -
# Project 2.1.3: In analogy with RandomVector create a RandomMatrix class whose components can either come from inputted data or be sampled from a distribution. Make use of existing methods for matrix algebra in numpy to multiply your RandomMatrices and RandomVectors.
# +
# Your code for project 2.1.3.
# You can also create your own notebook in case things get unwieldy here.
# -
# ## 3. Concentration of the norm
#
# Given a random vector $X$ of dimension $n$ whose entries are independent random variables with zero means and unit variances we are told in section 3.1 of the text that $$\mathbb{E}\|X\|_2^2=n.$$ We write a function to experimentally verify this and see what happens with other types of random vectors.
def square_norm_expectation(m,dim,distribution=lambda dim: np.random.default_rng().standard_normal(dim)):
"""
Find the computed mean of the norm squared for a random vector.
Args:
m (int): The number of vectors we should use in our test.
dim (int): The dimension of the ambient real vector space.
distribution (function): The distribution according to which the vector is generated.
Returns:
numpy.float64: The approximate expectation of the norm squared of such a random vector.
"""
# Create an immutable set of `m` random vectors in the appropriate space.
vectors = frozenset(RandomVector(dim,distribution) for i in range(m))
# Make a tuple out of the norms squared of these vectors.
norms_squared = tuple(x.norm_squared() for x in vectors)
# Compute the average (counting measure expectation) and return it.
return sum(norms_squared)/m
# +
print('Examples of the `square_norm_expectation` function.\n')
print('Compute the approximate expectation of the norm squared of a random vector with zero means and unit variances from the normal distribution.')
print('In this case we use 10000 samples in a 17 dimensional space.')
print(square_norm_expectation(1000,17))
print()
print('We plot the computed expectation obtained from 1000 samples for various choices of `dim` from 1 to 30.')
x = np.arange(1,31)
y = np.array(tuple(square_norm_expectation(1000,dim) for dim in range(1,31)))
plt.title("Computed expectation for various dimensions (normal distribution)")
plt.xlabel("Dimension")
plt.ylabel("Computed expectation")
plt.plot(x,y)
plt.show()
print('We can also use a different mean, say 10 rather than 0.')
x = np.arange(1,31)
y = np.array(tuple(square_norm_expectation(1000,dim,lambda dim: rng.normal(10,size=dim)) for dim in range(1,31)))
plt.title("Computed expectation for various dimensions (normal distribution, mean=10)")
plt.xlabel("Dimension")
plt.ylabel("Computed expectation")
plt.plot(x,y)
plt.show()
print('Other distributions also give different results.')
x = np.arange(1,31)
y = np.array(tuple(square_norm_expectation(1000,dim,lambda dim: rng.wald(1,1,dim)) for dim in range(1,31)))
plt.title("Computed expectation for various dimensions (Wald distribution)")
plt.xlabel("Dimension")
plt.ylabel("Computed expectation")
plt.plot(x,y)
plt.show()
# -
# ### 3.1 Projects, exercises, and comments on concentration of the norm.
# Exercise 3.1.1: Compute the expectation of the norm squared $$\|X\|^2=X_1^2+\cdots+X_n^2$$ of a random vector $X=(X_1,\dots,X_n)$ in $n$ dimensions whose components are independent random variables with mean $\mu$ and variance $\sigma$. Explain how this formula agrees with our experimental plots above.
# +
# Your code for exercise 3.1.1.
# This exercise asks you to perform a calculation and explain how the result fits with our experiments, but you can also implement it as a Python function if you'd like.
# -
# Project 3.1.2: Examine the variance of $\|X\|^2$ for a random vector $X$ whose components are i.i.d. random variables. Why is the line in the second example so much straighter?
# +
# Your code for project 3.1.2.
# -
# Project 3.1.3: Plot random vectors in two dimensions normalized by their length. Look at the distribution of the distances between successive points on the circle.
# +
# Your code for project 3.1.3.
# -
# ## 4 Real-world data and RandomVector
# As alluded to previously, we can also create a RandomVector from real-world data. We can then use our RandomVector class to study this data just as we did with the vectors we got from the random number generator in numpy.
#
# Let's take a look at the quantity of various nutrients present in the Mystic River in Massachusetts, as recorded by the [Massachusetts Water Resources Authority](http://www.mwra.state.ma.us/index.html). This organization's publicly available data can be found [here](http://www.mwra.state.ma.us/harbor/html/wq_data.htm), but please be sure to let them know if you use it for anything substantial so the government researchers who collected it can be apprised.
# Use the `read_excel` method in pandas to create a dataframe from the Mystic River nutrient data.
# This is a little different from the earthquake example we did on Sunday because the data is in an Excel file.
# Remember not to load data like this too many times automatically, since it will act like a denial of service attack on the data provider.
# We need to specify the sheet within the Excel file that we want pandas to look at with `sheet_name`.
# We also need to tell pandas to skip the first few rows, which contain other information.
mystic_data = pd.read_excel('http://www.mwra.state.ma.us/harbor/graphic/mr_nutrients.xlsx',sheet_name='Nutrients-Mystic all yrs',skiprows=range(4))
# Examine the resulting dataframe.
mystic_data
# As is often the case, we need to preprocess this raw data a little bit.
#
# Let's only look at the mouth of the Mystic River and use only those measurements of the levels of ammonium, nitrate or nitrite, phosphate, chlorophyll A, and phaeophytin which were taken at the bottom of the river.
# Create a new dataframe consisting only of those rows for measurements at the bottom and mouth of the river.
mystic_df = mystic_data.loc[(mystic_data['Subregion']=='MYSTIC MOUTH') & (mystic_data['Surface or Bottom']=='B')]
# Restrict this dataframe to only those columns pertaining to the five nutrients previously indicated.
mystic_df = mystic_df[['Ammonium (uM)','Nitrate+nitrite (uM)','Phosphate (uM)','Chlorophyll a (ug/L)','Phaeophytin (ug/L)']]
# Throw out any rows where some of those five nutrients were not measured.
mystic_df = mystic_df.dropna()
# Display the resulting dataframe.
mystic_df
# We want to treat each column in this dataframe as a (sample of a) random vector in $\mathbb{R}^5$.
#
# In order to do this we want to write a function which takes a dataframe as input and outputs a collection of RandomVectors. This will be made simpler if we have a clean way of making a RandomVector from a specified array in numpy. That is, by default a RandomVector wants to be created by telling to what its dimension should be and how it should generate its components. If those components are just specified in advance, the resulting function we pass in is a constant, creating an unnecessarily messy-looking construct. We now sweep this under the rug using a subclass.
class DataVector(RandomVector):
"""
A random vector generated by given data rather than a Python function.
Attributes:
components (numpy.ndarray): The components of the vector.
dim (int): The dimension in which the vector lies.
"""
def __init__(self,components):
"""
Args:
components (iterable): The components of the vector.
"""
# Cast `components` to a numpy array if it isn't already one.
components = np.array(components)
# Make `self` into a RandomVector with the appropriate dimension and components.
RandomVector.__init__(self,len(components),lambda dim: components)
# +
print('Examples of the `DataVector` class.\n')
print('Create a vector `x` with components [2,1,4] and print its basic information and components.')
x = DataVector([2,1,4])
print(x)
print(x.components)
print()
print('Note that `x` inherits all the methods of RandomVector.')
print(x.mean())
print(x.norm_squared())
# -
# Now we are ready to write a function which turns a dataframe into a collection of RandomVectors.
def make_vectors_from_dataframe(df):
"""
Create a collection of random vectors from a given dataframe.
Each row of the dataframe becomes the array of components for a RandomVector.
The supplied dataframe should contain only numerical entries.
Args:
df (pandas.core.frame.DataFrame): The dataframe to process.
Returns:
frozenset of DataVector: The set of vectors so generated.
"""
return frozenset(DataVector(df.loc[key].array) for key in df.index)
# Create a collection of RandomVectors from our processed Mystic River dataframe.
mystic_vectors = make_vectors_from_dataframe(mystic_df)
# Compute the average norm squared of these vectors.
print(sum(x.norm_squared() for x in mystic_vectors)/len(mystic_vectors))
# We can use the StandardScalar preprocessing function from [scikit-learn](https://scikit-learn.org/stable/index.html) in order to subtract of means and normalize by variances in each column of our dataset.
#
# Taking the average norm squared of the resulting vectors gives us a quantity very close to 5, agreeing with our earlier calculations.
# Make a new dataframe by using StandardScalar to give each column zero mean and unit variance.
mystic_normalized = pd.DataFrame(StandardScaler().fit_transform(mystic_df))
# Make this dataframe into a collection of RandomVectors.
mystic_normalized_vectors = make_vectors_from_dataframe(mystic_normalized)
# Compute the average norm squared of these vectors.
print(sum(x.norm_squared() for x in mystic_normalized_vectors)/len(mystic_normalized_vectors))
# ### 4.1 Projects, exercises, and comments on real-world data and RandomVector.
# Exercise 4.1.1: If the components of these nutrient data vectors were sampled from distribution with identical means and variances we would expect our formula from exercise 3.1.1 to hold. Compute the means and variances of the components of these vectors to determine whether or not this is the case. (Hint: A single line of code will answer this.)
# +
# Your code for exercise 4.1.1.
# -
# Exercise 4.1.2: Use numpy to compute the covariance and correlation matrices of the nutrient data vectors. Can you explain those correlations which are strongest? (Or do you have a (bio)chemist friend who can?) How does using the normalized rather than unnormalized vectors change the resulting values?
# +
# Your code for exercise 4.1.2.
# -
# Exercise 4.1.3: Create a histogram of the norm squared for the nutrient data vectors and a histogram of the norm squared for a collection of vectors generated with independent components. Compare the results and explain the similarities or differences.
# +
# Your code for exercise 4.1.3.
# -
# Project 4.1.4: Choose your own data set and produce a collection of DataVectors from it. See if you can modify the DataVector class so that it doesn't create a new object when it is given an invalid argument for its components.
# +
# Your code for project 4.1.4.
# -
# Copyright (c) 2020 TRIPODS/GradStemForAll 2020 Team
|
Other Notebooks/norm_concentration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.0
# language: julia
# name: julia-1.0
# ---
# # Advanced Problem Answers
#
# ## Metaprogramming Problem
macro myevalpoly(z,a...)
isempty(a) && error("You forgot to pass coefficients!")
ex = :($(a[length(a)]))
for i in 1:length(a)-1
ex = :($ex * $(z) + $(a[length(a)-i]) )
end
println(ex)
ex
end
@myevalpoly 7 2 3 4 5
@evalpoly 7 2 3 4 5
# ## Plot the roots of Wilkinson's polynomial with perturbation
#
# First, we need to construct coefficients $a_k$. For the polynomial $\prod_{i=1}^4 (x-z_i)$, we have the coefficients $$\left(
# \begin{array}{c}
# z_1 z_2 z_3 z_4 \\
# -z_1 z_2 z_3-z_1 z_4 z_3-z_2 z_4 z_3-z_1 z_2 z_4 \\
# z_1 z_2+z_3 z_2+z_4 z_2+z_1 z_3+z_1 z_4+z_3 z_4 \\
# -z_1-z_2-z_3-z_4 \\
# 1 \\
# \end{array}
# \right),$$ thus we can exploit the structure and write a double `for` loop to calculate the coefficients. A more general formula is
# $$
# \begin{cases}
# 1 = a_{n}\\
# x_{1}+x_{2}+\dots +x_{n-1}+x_{n}=-a_{n-1}\\
# (x_{1}x_{2}+x_{1}x_{3}+\cdots +x_{1}x_{n})+(x_{2}x_{3}+x_{2}x_{4}+\cdots +x_{2}x_{n})+\cdots +x_{n-1}x_{n}=a_{n-2}\\
# \quad \vdots \\
# x_{1}x_{2}\dots x_{n}=(-1)^{n}a_{0}.\end{cases}
# $$
# Checkout [Vieta's formulas](https://en.wikipedia.org/wiki/Vieta%27s_formulas) for more information.
function root2coeff(z::AbstractVector{T}) where T
N = length(z)
co = zeros(T, N+1)
# The last coefficient is always one
co[end] = 1
# The outer loop adds one root at a time
for j in 1:N, i in j:-1:1
co[end-i] -= z[j]*co[end-i+1]
end
co
end
@show typemax(Int), typemax(Int128)
root2coeff(1:20)
# Those numbers are close to `typemax(Int)`, so integer overflows may occur, lets use `Int128` instead.
root2coeff(Int128(1):20)
# Next, we need to construct a [companion matrix](https://en.wikipedia.org/wiki/Companion_matrix) and solve for roots.
# A companion matrix is in the form of
# $$
# \begin{bmatrix}0&0&\dots &0&-z_{1}\\1&0&\dots &0&-z_{2}\\0&1&\dots &0&-z_{3}\\\vdots &\vdots &\ddots &\vdots &\vdots \\0&0&\dots &1&-z_{n-1}\end{bmatrix}.
# $$
function poly_roots(z)
len = length(z)
# construct the ones part
mat = diagm(ones(len-2), -1)
# insert coefficients
mat[:, end] = -z[1:end-1]
eigvals(mat)
end
# We have everything ready now. We just need to calculate all the roots and plot it.
using Random
Random.seed!(1)
function wilkinson_poly_roots(n=100)
# original coefficients
coeff = root2coeff(Int128(1):20)
rts = Vector{Complex{Float64}}[]
# add perturbation
for i in 1:n
pert_coeff = coeff.*(1+rand(21)*1e-10)
push!(rts, poly_roots(pert_coeff))
end
rts
end
using Plots; gr()
function plt_wilkinson_roots(rts)
# plot roots without perturbation
plt = scatter(1:20, zeros(20), color = :green, markersize = 5, legend=false)
for i in eachindex(rts)
# plot roots with perturbation
scatter!(plt, real.(rts[i]), imag.(rts[i]), color = :red, markersize = .5)
end
plt
end
wilkinson_poly_roots() |> plt_wilkinson_roots
|
Notebooks/.ipynb_checkpoints/AdvancedProblemAnswers-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Deploying and Interacting with Campaigns <a class="anchor" id="top"></a>
#
# In this notebook, you will deploy and interact with campaigns in Amazon Personalize.
#
# 1. [Introduction](#intro)
# 1. [Create campaigns](#create)
# 1. [Interact with campaigns](#interact)
# 1. [Batch recommendations](#batch)
# 1. [Wrap up](#wrapup)
#
# ## Introduction <a class="anchor" id="intro"></a>
# [Back to top](#top)
#
# At this point, you should have several solutions and at least one solution version for each. Once a solution version is created, it is possible to get recommendations from them, and to get a feel for their overall behavior.
#
# This notebook starts off by deploying each of the solution versions from the previous notebook into individual campaigns. Once they are active, there are resources for querying the recommendations, and helper functions to digest the output into something more human-readable.
#
# As you with your customer on Amazon Personalize, you can modify the helper functions to fit the structure of their data input files to keep the additional rendering working.
#
# To get started, once again, we need to import libraries, load values from previous notebooks, and load the SDK.
# +
import time
from time import sleep
import json
from datetime import datetime
import uuid
import boto3
import pandas as pd
# -
# %store -r
# +
personalize = boto3.client('personalize')
personalize_runtime = boto3.client('personalize-runtime')
# Establish a connection to Personalize's event streaming
personalize_events = boto3.client(service_name='personalize-events')
# -
# ## Create campaigns <a class="anchor" id="create"></a>
# [Back to top](#top)
#
# A campaign is a hosted solution version; an endpoint which you can query for recommendations. Pricing is set by estimating throughput capacity (requests from users for personalization per second). When deploying a campaign, you set a minimum throughput per second (TPS) value. This service, like many within AWS, will automatically scale based on demand, but if latency is critical, you may want to provision ahead for larger demand. For this POC and demo, all minimum throughput thresholds are set to 1. For more information, see the [pricing page](https://aws.amazon.com/personalize/pricing/).
#
# Let's start deploying the campaigns.
# ### HRNN
#
# Deploy a campaign for your HRNN solution version. It can take around 10 minutes to deploy a campaign. Normally, we would use a while loop to poll until the task is completed. However the task would block other cells from executing, and the goal here is to create multiple campaigns. So we will set up the while loop for all of the campaigns further down in the notebook. There, you will also find instructions for viewing the progress in the AWS console.
# +
hrnn_create_campaign_response = personalize.create_campaign(
name = "personalize-poc-hrnn",
solutionVersionArn = hrnn_solution_version_arn,
minProvisionedTPS = 1
)
hrnn_campaign_arn = hrnn_create_campaign_response['campaignArn']
print(json.dumps(hrnn_create_campaign_response, indent=2))
# -
# ### SIMS
#
# Deploy a campaign for your SIMS solution version. It can take around 10 minutes to deploy a campaign. Normally, we would use a while loop to poll until the task is completed. However the task would block other cells from executing, and the goal here is to create multiple campaigns. So we will set up the while loop for all of the campaigns further down in the notebook. There, you will also find instructions for viewing the progress in the AWS console.
# +
sims_create_campaign_response = personalize.create_campaign(
name = "personalize-poc-SIMS",
solutionVersionArn = sims_solution_version_arn,
minProvisionedTPS = 1
)
sims_campaign_arn = sims_create_campaign_response['campaignArn']
print(json.dumps(sims_create_campaign_response, indent=2))
# -
# ### Personalized Ranking
#
# Deploy a campaign for your personalized ranking solution version. It can take around 10 minutes to deploy a campaign. Normally, we would use a while loop to poll until the task is completed. However the task would block other cells from executing, and the goal here is to create multiple campaigns. So we will set up the while loop for all of the campaigns further down in the notebook. There, you will also find instructions for viewing the progress in the AWS console.
# +
rerank_create_campaign_response = personalize.create_campaign(
name = "personalize-poc-rerank",
solutionVersionArn = rerank_solution_version_arn,
minProvisionedTPS = 1
)
rerank_campaign_arn = rerank_create_campaign_response['campaignArn']
print(json.dumps(rerank_create_campaign_response, indent=2))
# -
# ### View campaign creation status
#
# As promised, how to view the status updates in the console:
#
# * In another browser tab you should already have the AWS Console up from opening this notebook instance.
# * Switch to that tab and search at the top for the service `Personalize`, then go to that service page.
# * Click `View dataset groups`.
# * Click the name of your dataset group, most likely something with POC in the name.
# * Click `Campaigns`.
# * You will now see a list of all of the campaigns you created above, including a column with the status of the campaign. Once it is `Active`, your campaign is ready to be queried.
#
# Or simply run the cell below to keep track of the campaign creation status.
# +
in_progress_campaigns = [
hrnn_campaign_arn,
sims_campaign_arn,
rerank_campaign_arn
]
max_time = time.time() + 3*60*60 # 3 hours
while time.time() < max_time:
for campaign_arn in in_progress_campaigns:
version_response = personalize.describe_campaign(
campaignArn = campaign_arn
)
status = version_response["campaign"]["status"]
if status == "ACTIVE":
print("Build succeeded for {}".format(campaign_arn))
in_progress_campaigns.remove(campaign_arn)
elif status == "CREATE FAILED":
print("Build failed for {}".format(campaign_arn))
in_progress_campaigns.remove(campaign_arn)
if len(in_progress_campaigns) <= 0:
break
else:
print("At least one campaign build is still in progress")
time.sleep(60)
# -
# ## Interact with campaigns <a class="anchor" id="interact"></a>
# [Back to top](#top)
#
# Now that all campaigns are deployed and active, we can start to get recommendations via an API call. Each of the campaigns is based on a different recipe, which behave in slightly different ways because they serve different use cases. We will cover each campaign in a different order than used in previous notebooks, in order to deal with the possible complexities in ascending order (i.e. simplest first).
#
# First, let's create a supporting function to help make sense of the results returned by a Personalize campaign. Personalize returns only an `item_id`. This is great for keeping data compact, but it means you need to query a database or lookup table to get a human-readable result for the notebooks. We will create a helper function to return a human-readable result from the LastFM dataset.
#
# Start by loading in the dataset which we can use for our lookup table.
# +
# Create a dataframe for the items by reading in the correct source CSV
items_df = pd.read_csv(data_dir + '/artists.dat', delimiter='\t', index_col=0)
# Render some sample data
items_df.head(5)
# -
# By defining the ID column as the index column it is trivial to return an artist by just querying the ID.
item_id_example = 987
artist = items_df.loc[item_id_example]['name']
print(artist)
# That isn't terrible, but it would get messy to repeat this everywhere in our code, so the function below will clean that up.
def get_artist_by_id(artist_id, artist_df=items_df):
"""
This takes in an artist_id from Personalize so it will be a string,
converts it to an int, and then does a lookup in a default or specified
dataframe.
A really broad try/except clause was added in case anything goes wrong.
Feel free to add more debugging or filtering here to improve results if
you hit an error.
"""
try:
return artist_df.loc[int(artist_id)]['name']
except:
return "Error obtaining artist"
# Now let's test a few simple values to check our error catching.
# A known good id
print(get_artist_by_id(artist_id="987"))
# A bad type of value
print(get_artist_by_id(artist_id="987.9393939"))
# Really bad values
print(get_artist_by_id(artist_id="Steve"))
# Great! Now we have a way of rendering results.
# ### SIMS
#
# SIMS requires just an item as input, and it will return items which users interact with in similar ways to their interaction with the input item. In this particular case the item is an artist.
#
# So, let's sample some data from our dataset to test our SIMS campaign. Grab 5 random artists from our dataframe.
samples = items_df.sample(5)
samples
# The cells below will handle getting recommendations from SIMS and rendering the results. Let's see what the recommendations are for the first item we looked at earlier in this notebook (Earth, Wind & Fire).
get_recommendations_response = personalize_runtime.get_recommendations(
campaignArn = sims_campaign_arn,
itemId = str(987),
)
item_list = get_recommendations_response['itemList']
for item in item_list:
print(get_artist_by_id(artist_id=item['itemId']))
# Congrats, this is your first list of recommendations! This list is fine, but it would be better to see the recommendations for our sample collection of artists render in a nice dataframe. Again, let's create a helper function to achieve this.
# +
# Update DF rendering
pd.set_option('display.max_rows', 30)
def get_new_recommendations_df(recommendations_df, artist_ID):
# Get the artist name
artist_name = get_artist_by_id(artist_ID)
# Get the recommendations
get_recommendations_response = personalize_runtime.get_recommendations(
campaignArn = sims_campaign_arn,
itemId = str(artist_ID),
)
# Build a new dataframe of recommendations
item_list = get_recommendations_response['itemList']
recommendation_list = []
for item in item_list:
artist = get_artist_by_id(item['itemId'])
recommendation_list.append(artist)
new_rec_DF = pd.DataFrame(recommendation_list, columns = [artist_name])
# Add this dataframe to the old one
recommendations_df = pd.concat([recommendations_df, new_rec_DF], axis=1)
return recommendations_df
# -
# Now, let's test the helper function with the sample of artists we chose before.
# +
sims_recommendations_df = pd.DataFrame()
artists = samples.index.tolist()
for artist in artists:
sims_recommendations_df = get_new_recommendations_df(sims_recommendations_df, artist)
sims_recommendations_df
# -
# You may notice that a lot of the items look the same, hopefully not all of them do. This shows that the evaluation metrics should not be the only thing you rely on when evaluating your solution version. So when this happens, what can you do to improve the results?
#
# This is a good time to think about the hyperparameters of the Personalize recipes. The SIMS recipe has a `popularity_discount_factor` hyperparameter (see [documentation](https://docs.aws.amazon.com/personalize/latest/dg/native-recipe-sims.html)). Leveraging this hyperparameter allows you to control the nuance you see in the results. This parameter and its behavior will be unique to every dataset you encounter, and depends on the goals of the business. You can iterate on the value of this hyperparameter until you are satisfied with the results, or you can start by leveraging Personalize's hyperparameter optimization (HPO) feature. For more information on hyperparameters and HPO tuning, see the [documentation](https://docs.aws.amazon.com/personalize/latest/dg/customizing-solution-config-hpo.html).
# ### HRNN
#
# HRNN is one of the more advanced algorithms provided by Amazon Personalize. It supports personalization of the items for a specific user based on their past behavior and can intake real time events in order to alter recommendations for a user without retraining.
#
# Since HRNN relies on having a sampling of users, let's load the data we need for that and select 3 random users.
users_df = pd.read_csv(data_dir + '/user_artists.dat', delimiter='\t', index_col=0)
# Render some sample data
users_df.head(5)
users = users_df.sample(3).index.tolist()
users
# Now we render the recommendations for our 3 random users from above. After that, we will explore real-time interactions before moving on to Personalized Ranking.
#
# Again, we create a helper function to render the results in a nice dataframe.
#
# #### API call results
# +
# Update DF rendering
pd.set_option('display.max_rows', 30)
def get_new_recommendations_df_users(recommendations_df, user_id):
# Get the artist name
#artist_name = get_artist_by_id(artist_ID)
# Get the recommendations
get_recommendations_response = personalize_runtime.get_recommendations(
campaignArn = hrnn_campaign_arn,
userId = str(user_id),
)
# Build a new dataframe of recommendations
item_list = get_recommendations_response['itemList']
recommendation_list = []
for item in item_list:
artist = get_artist_by_id(item['itemId'])
recommendation_list.append(artist)
#print(recommendation_list)
new_rec_DF = pd.DataFrame(recommendation_list, columns = [user_id])
# Add this dataframe to the old one
recommendations_df = pd.concat([recommendations_df, new_rec_DF], axis=1)
return recommendations_df
# +
recommendations_df_users = pd.DataFrame()
users = users_df.sample(3).index.tolist()
print(users)
for user in users:
recommendations_df_users = get_new_recommendations_df_users(recommendations_df_users, user)
recommendations_df_users
# -
# Here we clearly see that the recommendations for each user are different. If you were to need a cache for these results, you could start by running the API calls through all your users and store the results, or you could use a batch export, which will be covered later in this notebook.
#
# The next topic is real-time events. Personalize has the ability to listen to events from your application in order to update the recommendations shown to the user. This is especially useful in media workloads, like video-on-demand, where a customer's intent may differ based on if they are watching with their children or on their own.
#
# Additionally the events that are recorded via this system are stored until a delete call from you is issued, and they are used as historical data alongside the other interaction data you provided when you train your next models.
#
# #### Real time events
#
# Start by creating an event tracker that is attached to the campaign.
response = personalize.create_event_tracker(
name='ArtistTracker',
datasetGroupArn=dataset_group_arn
)
print(response['eventTrackerArn'])
print(response['trackingId'])
TRACKING_ID = response['trackingId']
event_tracker_arn = response['eventTrackerArn']
# We will create some code that simulates a user interacting with a particular item. After running this code, you will get recommendations that differ from the results above.
#
# We start by creating some methods for the simulation of real time events.
# +
session_dict = {}
def send_artist_click(USER_ID, ITEM_ID):
"""
Simulates a click as an envent
to send an event to Amazon Personalize's Event Tracker
"""
# Configure Session
try:
session_ID = session_dict[str(USER_ID)]
except:
session_dict[str(USER_ID)] = str(uuid.uuid1())
session_ID = session_dict[str(USER_ID)]
# Configure Properties:
event = {
"itemId": str(ITEM_ID),
}
event_json = json.dumps(event)
# Make Call
personalize_events.put_events(
trackingId = TRACKING_ID,
userId= str(USER_ID),
sessionId = session_ID,
eventList = [{
'sentAt': int(time.time()),
'eventType': 'EVENT_TYPE',
'properties': event_json
}]
)
def get_new_recommendations_df_users_real_time(recommendations_df, user_id, item_id):
# Get the artist name (header of column)
artist_name = get_artist_by_id(item_id)
# Interact with the artist
send_artist_click(USER_ID=user_id, ITEM_ID=item_id)
# Get the recommendations (note you should have a base recommendation DF created before)
get_recommendations_response = personalize_runtime.get_recommendations(
campaignArn = hrnn_campaign_arn,
userId = str(user_id),
)
# Build a new dataframe of recommendations
item_list = get_recommendations_response['itemList']
recommendation_list = []
for item in item_list:
artist = get_artist_by_id(item['itemId'])
recommendation_list.append(artist)
new_rec_DF = pd.DataFrame(recommendation_list, columns = [artist_name])
# Add this dataframe to the old one
#recommendations_df = recommendations_df.join(new_rec_DF)
recommendations_df = pd.concat([recommendations_df, new_rec_DF], axis=1)
return recommendations_df
# -
# At this point, we haven't generated any real-time events yet; we have only set up the code. To compare the recommendations before and after the real-time events, let's pick one user and generate the original recommendations for them.
# +
# First pick a user
user_id = users_df.sample(1).index.tolist()[0]
# Get recommendations for the user
get_recommendations_response = personalize_runtime.get_recommendations(
campaignArn = hrnn_campaign_arn,
userId = str(user_id),
)
# Build a new dataframe for the recommendations
item_list = get_recommendations_response['itemList']
recommendation_list = []
for item in item_list:
artist = get_artist_by_id(item['itemId'])
recommendation_list.append(artist)
user_recommendations_df = pd.DataFrame(recommendation_list, columns = [user_id])
user_recommendations_df
# -
# Ok, so now we have a list of recommendations for this user before we have applied any real-time events. Now let's pick 3 random artists which we will simulate our user interacting with, and then see how this changes the recommendations.
# Next generate 3 random artists
artists = items_df.sample(3).index.tolist()
# Note this will take about 15 seconds to complete due to the sleeps
for artist in artists:
user_recommendations_df = get_new_recommendations_df_users_real_time(user_recommendations_df, user_id, artist)
time.sleep(5)
user_recommendations_df
# In the cell above, the first column after the index is the user's default recommendations from HRNN, and each column after that has a header of the artist that they interacted with via a real time event, and the recommendations after this event occurred.
#
# The behavior may not shift very much; this is due to the relatively limited nature of this dataset. If you wanted to better understand this, try simulating clicking random artists of random genres, and you should see a more pronounced impact.
# ### Personalized Ranking
#
# The core use case for personalized ranking is to take a collection of items and to render them in priority or probable order of interest for a user. To demonstrate this, we will need a random user and a random collection of 25 items.
rerank_user = users_df.sample(1).index.tolist()[0]
rerank_items = items_df.sample(25).index.tolist()
# Now build a nice dataframe that shows the input data.
rerank_list = []
for item in rerank_items:
artist = get_artist_by_id(item)
rerank_list.append(artist)
rerank_df = pd.DataFrame(rerank_list, columns = [rerank_user])
rerank_df
# Then make the personalized ranking API call.
# +
# Convert user to string:
user_id = str(rerank_user)
rerank_item_list = []
for item in rerank_items:
rerank_item_list.append(str(item))
# Get recommended reranking
get_recommendations_response_rerank = personalize_runtime.get_personalized_ranking(
campaignArn = rerank_campaign_arn,
userId = user_id,
inputList = rerank_item_list
)
get_recommendations_response_rerank
# -
# Now add the reranked items as a second column to the original dataframe, for a side-by-side comparison.
ranked_list = []
item_list = get_recommendations_response_rerank['personalizedRanking']
for item in item_list:
artist = get_artist_by_id(item['itemId'])
ranked_list.append(artist)
ranked_df = pd.DataFrame(ranked_list, columns = ['Re-Ranked'])
rerank_df = pd.concat([rerank_df, ranked_df], axis=1)
rerank_df
# You can see above how each entry was re-ordered based on the model's understanding of the user. This is a popular task when you have a collection of items to surface a user, a list of promotions for example, or if you are filtering on a category and want to show the most likely good items.
# ## Batch recommendations <a class="anchor" id="batch"></a>
# [Back to top](#top)
#
# There are many cases where you may want to have a larger dataset of exported recommendations. Recently, Amazon Personalize launched batch recommendations as a way to export a collection of recommendations to S3. In this example, we will walk through how to do this for the HRNN solution. For more information about batch recommendations, please see the [documentation](https://docs.aws.amazon.com/personalize/latest/dg/getting-recommendations.html#recommendations-batch). This feature applies to all recipes, but the output format will vary.
#
# A simple implementation looks like this:
#
# ```python
# import boto3
#
# personalize_rec = boto3.client(service_name='personalize')
#
# personalize_rec.create_batch_inference_job (
# solutionVersionArn = "Solution version ARN",
# jobName = "Batch job name",
# roleArn = "IAM role ARN",
# jobInput =
# {"s3DataSource": {"path": S3 input path}},
# jobOutput =
# {"s3DataDestination": {"path":S3 output path"}}
# )
# ```
#
# The SDK import, the solution version arn, and role arns have all been determined. This just leaves an input, an output, and a job name to be defined.
#
# Starting with the input for HRNN, it looks like:
#
#
# ```JSON
# {"userId": "4638"}
# {"userId": "663"}
# {"userId": "3384"}
# ```
#
# This should yield an output that looks like this:
#
# ```JSON
# {"input":{"userId":"4638"}, "output": {"recommendedItems": ["296", "1", "260", "318"]}}
# {"input":{"userId":"663"}, "output": {"recommendedItems": ["1393", "3793", "2701", "3826"]}}
# {"input":{"userId":"3384"}, "output": {"recommendedItems": ["8368", "5989", "40815", "48780"]}}
# ```
#
# The output is a JSON Lines file. It consists of individual JSON objects, one per line. So we will need to put in more work later to digest the results in this format.
# ### Building the input file
#
# When you are using the batch feature, you specify the users that you'd like to receive recommendations for when the job has completed. The cell below will again select a few random users and will then build the file and save it to disk. From there, you will upload it to S3 to use in the API call later.
# +
# Get the user list
batch_users = users_df.sample(3).index.tolist()
# Write the file to disk
json_input_filename = "json_input.json"
with open(data_dir + "/" + json_input_filename, 'w') as json_input:
for user_id in batch_users:
json_input.write('{"userId": "' + str(user_id) + '"}\n')
# -
# Showcase the input file:
# !cat $data_dir"/"$json_input_filename
# Upload the file to S3 and save the path as a variable for later.
# Upload files to S3
boto3.Session().resource('s3').Bucket(bucket_name).Object(json_input_filename).upload_file(data_dir+"/"+json_input_filename)
s3_input_path = "s3://" + bucket_name + "/" + json_input_filename
print(s3_input_path)
# Batch recommendations read the input from the file we've uploaded to S3. Similarly, batch recommendations will save the output to file in S3. So we define the output path where the results should be saved.
# Define the output path
s3_output_path = "s3://" + bucket_name + "/"
print(s3_output_path)
# Now just make the call to kick off the batch export process.
batchInferenceJobArn = personalize.create_batch_inference_job (
solutionVersionArn = hrnn_solution_version_arn,
jobName = "POC-Batch-Inference-Job-HRNN",
roleArn = role_arn,
jobInput =
{"s3DataSource": {"path": s3_input_path}},
jobOutput =
{"s3DataDestination":{"path": s3_output_path}}
)
batchInferenceJobArn = batchInferenceJobArn['batchInferenceJobArn']
# Run the while loop below to track the status of the batch recommendation call. This can take around 25 minutes to complete, because Personalize needs to stand up the infrastructure to perform the task. We are testing the feature with a dataset of only 3 users, which is not an efficient use of this mechanism. Normally, you would only use this feature for bulk processing, in which case the efficiencies will become clear.
# +
current_time = datetime.now()
print("Import Started on: ", current_time.strftime("%I:%M:%S %p"))
max_time = time.time() + 3*60*60 # 3 hours
while time.time() < max_time:
describe_dataset_inference_job_response = personalize.describe_batch_inference_job(
batchInferenceJobArn = batchInferenceJobArn
)
status = describe_dataset_inference_job_response["batchInferenceJob"]['status']
print("DatasetInferenceJob: {}".format(status))
if status == "ACTIVE" or status == "CREATE FAILED":
break
time.sleep(60)
current_time = datetime.now()
print("Import Completed on: ", current_time.strftime("%I:%M:%S %p"))
# -
# Once the batch recommendations job has finished processing, we can grab the output uploaded to S3 and parse it.
# +
s3 = boto3.client('s3')
export_name = json_input_filename + ".out"
s3.download_file(bucket_name, export_name, data_dir+"/"+export_name)
# Update DF rendering
pd.set_option('display.max_rows', 30)
with open(data_dir+"/"+export_name) as json_file:
# Get the first line and parse it
line = json.loads(json_file.readline())
# Do the same for the other lines
while line:
# extract the user ID
col_header = "User: " + line['input']['userId']
# Create a list for all the artists
recommendation_list = []
# Add all the entries
for item in line['output']['recommendedItems']:
artist = get_artist_by_id(item)
recommendation_list.append(artist)
if 'bulk_recommendations_df' in locals():
new_rec_DF = pd.DataFrame(recommendation_list, columns = [col_header])
bulk_recommendations_df = bulk_recommendations_df.join(new_rec_DF)
else:
bulk_recommendations_df = pd.DataFrame(recommendation_list, columns=[col_header])
try:
line = json.loads(json_file.readline())
except:
line = None
bulk_recommendations_df
# -
# ## Wrap up <a class="anchor" id="wrapup"></a>
# [Back to top](#top)
#
# With that you now have a fully working collection of models to tackle various recommendation and personalization scenarios, as well as the skills to manipulate customer data to better integrate with the service, and a knowledge of how to do all this over APIs and by leveraging open source data science tools.
#
# Use these notebooks as a guide to getting started with your customers for POCs. As you find missing components, or discover new approaches, cut a pull request and provide any additional helpful components that may be missing from this collection.
#
# You'll want to make sure that you clean up all of the resources deployed during this POC. We have provided a separate notebook which shows you how to identify and delete the resources in `04_Clean_Up_Resources.ipynb`.
|
workshops/POC_in_a_box/completed/03_Deploying_Campaigns_and_Interacting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"autoexec": {"startup": false, "wait_interval": 0.0}, "base_uri": "https://localhost:8080/", "height": 192.0} colab_type="code" executionInfo={"elapsed": 49348.0, "status": "ok", "timestamp": 1529348255559.0, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-PiJFrbeyvzs/AAAAAAAAAAI/AAAAAAAAFzs/UKFF-SotCVo/s50-c-k-no/photo.jpg", "userId": "112993143985468015422"}, "user_tz": 240.0} id="oloy5NpyUxmL" outputId="3c4de2d8-e39c-4bcb-d0b2-8cdacfd28121"
import os
import skimage.io
import matplotlib.pyplot as plt
# %matplotlib inline
# Import Mask RCNN
from mrcnn.config import Config
import mrcnn.model as modellib
from mrcnn import utils
# Root directory of the project
ROOT_DIR = os.path.abspath("../")
print('Project Directory: {}'.format(ROOT_DIR))
# Root directory of the dataset
DATA_DIR = os.path.join(ROOT_DIR, "dataset/wad")
print('Data Directory: {}'.format(DATA_DIR))
# Directory to save logs and trained model
LOGS_DIR = os.path.join(ROOT_DIR, "logs")
print('Logs and Model Directory: {}'.format(LOGS_DIR))
# Local path to trained coco weights file
COCO_MODEL_PATH = os.path.join(LOGS_DIR, 'mask_rcnn_coco.h5')
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
# + [markdown] colab_type="text" id="ZrDSKKPUTJII"
# ## Configuration
# + colab={"autoexec": {"startup": false, "wait_interval": 0.0}, "base_uri": "https://localhost:8080/", "height": 892.0} colab_type="code" executionInfo={"elapsed": 184.0, "status": "ok", "timestamp": 1529348394269.0, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-PiJFrbeyvzs/AAAAAAAAAAI/AAAAAAAAFzs/UKFF-SotCVo/s50-c-k-no/photo.jpg", "userId": "112993143985468015422"}, "user_tz": 240.0} id="OTT9HIDCTC8i" outputId="79f542b0-7ee6-437f-fd7a-effa5b9f932c"
from project import wad_data
cfg = wad_data.WADConfig()
cfg.display()
# + [markdown] colab_type="text" id="Unxlaax_TSfv"
# ## Dataset
#
# load_mask has been tested. green light .. green light
# + colab={"autoexec": {"startup": false, "wait_interval": 0.0}} colab_type="code" id="dEyP-1tATFRb"
dataset_train = wad_data.WADDataset()
dataset_val = dataset_train.load_data(DATA_DIR, "train", val_size=0.2)
dataset_train.prepare()
dataset_val.prepare()
dataset_val.save_data_to_file(os.path.join(LOGS_DIR, "last_run_validation.pkl"))
# + [markdown] colab_type="text" id="UUHgP9PsvYIb"
# ## Training
# + colab={"autoexec": {"startup": false, "wait_interval": 0.0}, "base_uri": "https://localhost:8080/", "height": 684.0} colab_type="code" id="PK_mhBMzvW7D" outputId="2548bddd-ff94-4b8d-af4f-63131aab93aa"
STARTING_WEIGHTS = os.path.join(LOGS_DIR, 'mask_rcnn_cooc.h5')
# Create model in training mode
model = modellib.MaskRCNN(mode="training", config=cfg, model_dir=LOGS_DIR)
model.load_weights(STARTING_WEIGHTS, by_name=True, exclude=[
"mrcnn_class_logits", "mrcnn_bbox_fc",
"mrcnn_bbox", "mrcnn_mask"])
model.train(dataset_train, dataset_val,
learning_rate=cfg.LEARNING_RATE,
epochs=1,
layers='heads')
|
tests/wad_data_training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Students Turn Activity 1 JSON Traversal Review
#
# This activity is an opportunity to practice loading and parsing JSON in Python.
#
# ## Instructions
#
# * Load the provided JSON
#
# * Retrieve the video's title
#
# * Retrieve the video's rating
#
# * Retrieve the link to the video's thumbnail
#
# * Retrieve the number of views this video has
# +
# Dependencies
import json
import requests
import os
# Load JSON
filepath = os.path.join("Resources", "youtube_response.json")
with open(filepath) as jsonfile:
video_json = json.load(jsonfile)
# -
# Isolate "data items" for easy reading
# CODE HERE
data_items = video_json["data"]["items"]
print(data_items)
# Retrieve the video's title
# CODE HERE
print(data_items[0]["title"])
# Retrieve the video's rating
# CODE HERE
print(data_items[0]["rating"])
# +
# Retrieve the link to the video's default thumbnail
# CODE HERE
print(data_items[0]["thumbnail"]["default"])
# +
# Retrieve the number of views this video has
# CODE HERE
print(data_items[0]["viewCount"])
# -
# # Students Turn Activity 2 Requests & Responses
#
# This activity provides practice making API calls, converting the response to JSON, and then manipulating the result with Python.
#
# ## Instructions
#
# * Make a request to the following endpoint (http://nyt-mongo-scraper.herokuapp.com/api/headlines), and store the response.
#
# * JSON-ify the response.
#
# * Print the JSON representations of the first and last posts.
#
# * Print number of posts received.
# Dependencies
import json
import requests
# +
# Specify the URL
# CODE HERE
url = "http://nyt-mongo-scraper.herokuapp.com/api/headlines"
# Make request and store response
# CODE HERE
variable = requests.get(url).json()
variable
# -
# Print first and last articles
# CODE HERE
print(variable[0])
print(variable[-1]) #!!!!!!!!!!!!!! index of -1 does the last!!
#Print the number of responses received.
# CODE HERE
print(len(variable))
# # Instructor Turn Activity 3 Open Weather Request
# Dependencies
import json
import requests
from config import api_key
# +
# Save config information
url = "http://api.openweathermap.org/data/2.5/weather?"
city = "London"
# Build query URL
query_url = url + "appid=" + api_key + "&q=" + city
# -
# Get weather data
weather_response = requests.get(query_url)
weather_json = weather_response.json()
# Get the temperature from the response
print(f"The weather API responded with: {json.dumps(weather_json, indent=2)}.")
# # Students Turn Activity 4 Weather in Bujumbura
#
# This activity gives students practice with making API calls and handling responses.
#
# ## Instructions
#
# * Save all of your "config" information—i.e., your API key; the base URL; etc.—before moving on.
#
# * Build your query URL. Check the documentation to figure out how to request temperatures in Celsius.
#
# * Make your request, and save the API's response.
#
# * Retrieve the current temperature in Bujumbura from the JSON response.
#
# * Print the temperature to the console.
#
# ## Bonus
#
# * Augment your code to report the temperature in both Fahrenheit _and_ Celsius.
# +
# Dependencies
import requests
from config import api_key
# Save config information.
url = "http://api.openweathermap.org/data/2.5/weather?"
city = "Bujumbura"
units = "metric"
# -
# Build query URL and request your results in Celsius
# CODE HERE
query = url + "appid=" + api_key + "&q=" + city + "&units=metric"
# Get weather data
# CODE HERE
weather_bermuda = requests.get(query).json()
weather_bermuda
# Get temperature from JSON response
# CODE HERE
temp = weather_bermuda["main"]["temp"]
# Report temperature
# CODE HERE
print(f"The temperature is {temp} degrees Celsius")
# BONUS
# # Instructor Turn Activity 5 Open Weather DataFrame
# Dependencies
import csv
import matplotlib.pyplot as plt
import requests
import pandas as pd
from config import api_key
# +
# Save config information.
url = "http://api.openweathermap.org/data/2.5/weather?"
units = "metric"
# Build partial query URL
query_url = f"{url}appid={api_key}&units={units}&q="
# +
cities = ["Paris", "London", "Oslo", "Beijing"]
# set up lists to hold reponse info
lat = []
temp = []
# Loop through the list of cities and perform a request for data on each
for city in cities:
response = requests.get(query_url + city).json()
lat.append(response['coord']['lat'])
temp.append(response['main']['temp'])
print(f"The latitude information received is: {lat}")
print(f"The temperature information received is: {temp}")
# -
# create a data frame from cities, lat, and temp
weather_dict = {
"city": cities,
"lat": lat,
"temp": temp
}
weather_data = pd.DataFrame(weather_dict)
weather_data.head()
# +
# Build a scatter plot for each data type
plt.scatter(weather_data["lat"], weather_data["temp"], marker="o")
# Incorporate the other graph properties
plt.title("Temperature in World Cities")
plt.ylabel("Temperature (Celsius)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("Images/TemperatureInWorldCities.png")
# Show plot
plt.show()
# -
# ### Students Turn Activity 6 TV Ratings
#
# In this activity, you will create an application that reads in a list of TV shows, makes multiple requests from an API to retrieve rating information, creates a pandas dataframe, and a visually displays the data.
#
# ## Instructions:
#
# * You may use the list provided in the starter file or create your own.
#
# * Request information from the TVmaze API's Show Search endpoint (https://www.tvmaze.com/api#show-search) on each show and store the name and rating information into lists.
#
# * Put this data into a dictionary, and load that dict into a Pandas DataFrame.
#
# * Use matplotlib to create a bar chart comparing the ratings of each show.
#Dependencies
import requests
import json
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# +
#list of tv show titles to query
tv_shows = ["Altered Carbon", "Grey's Anatomy", "This is Us", "The Flash", "Vikings", "Shameless", "Arrow", "Peaky Blinders", "Dirk Gently"]
# make iterative requests to TVmaze search endpoint
# tv maze show search base url
base_url = "http://api.tvmaze.com/search/shows?q="
name = []
rating = []
for show in tv_shows:
request = base_url + show
tvmaze = requests.get(request).json()
name.append(tvmaze[0]["show"]["name"])
rating.append(tvmaze[0]["show"]["rating"]["average"])
# set up lists to hold response data for name and rating
# loop through tv show titles, make requests and parse
print(name)
print(rating)
# -
# create dataframe
# CODE HERE
shows_dict = {
"name": name,
"rating": rating,
}
shows_data = pd.DataFrame(shows_dict)
shows_data.head()
# +
# use matplotlib to create a bar chart from the dataframe
#CODE HERE
shows_data.plot(kind="bar")
# create a list of numbers for x values
plt.xticks = (len(shows_data["name"]), shows_data["name"])
# create bar chart and set the values of xticks
shows_data["name"]
# -
# # Instructor Turn Activity 7 Exception Handling
# +
students = {
# Name : Age
"James": 27,
"Sarah": 19,
"Jocelyn": 28
}
print(students["Jezebel"])
print("This line will never print.")
# +
students = {
# Name : Age
"James": 27,
"Sarah": 19,
"Jocelyn": 28
}
# Try to access key that doesn't exist
try:
students["Jezebel"]
except KeyError:
print("Oops, that key doesn't exist.")
# "Catching" the error lets the rest of our code execute
print("...But the program doesn't die early!")
# -
# # Student Turn Activity
# ## Making Exceptions
#
# ### Instructions:
#
# * Without removing any of the lines from the starter code provided, create `try` and `except` blocks that will allow the application to run without terminating.
# Your assignment is to get the last line to print without changing any
# of the code below. Instead, wrap each line that throws an error in a
# try/except block.
try:
print("Infinity looks like + " + str(10 / 0) + ".")
except ZeroDivisionError:
print("error")
try:
print("I think her name was + " + name + "?")
except TypeError:
print("error")
try:
print("Your name is a nonsense number. Look: " + int("Gabriel"))
except ValueError:
print("error")
print("You made it through the gauntlet--the message survived!")
# Your assignment is to get the last line to print without changing any
# of the code below. Instead, wrap each line that throws an error in a
# try/exept block.
# CODE HERE
# # Instructor Turn Activity 9 Open Weather Wrapper
# !pip install openweathermapy
# +
# Dependencies
import openweathermapy.core as owm
#config
from config import api_key
# -
# Create settings dictionary with information we're interested in
settings = {"units": "metric", "appid": api_key}
# Get current weather
current_weather_paris = owm.get_current("Paris", **settings)
print(f"Current weather object for Paris: {current_weather_paris}.")
# +
summary = ["name", "main.temp"]
data = current_weather_paris(*summary)
print(f"The current weather summary for Paris is: {data}.")
# -
# # Students Turn Activity 10 Map Wrap
#
# This activity demonstrates the additional ease of use afforded by API wrappers.
#
# ## Instructions
#
# * Install the openweathermapy API wrapper.
#
# * Create a settings object with your API key and preferred units of measurement.
#
# * Get data for each city that is listed within `cities.csv`.
#
# * Create a list to get the temperature, latitude, and longitude in each city
#
# * Create a Pandas DataFrame with the results.
#
# * Print your summaries to verify that everything went smoothly.
#
# Hint: Don't forget to utilize the openweathermapy documentation where needed: http://openweathermapy.readthedocs.io/en/latest/
#
# ## Bonus:
#
# * If you finish early, read about and experiment with the `*` syntax.
#
# * Pass a `columns` keyword argument to `pd.DataFrame`, and provide labels for the temperature and coordinate data.
# +
# Dependencies
import csv
import matplotlib.pyplot as plt
import openweathermapy as ow
import pandas as pd
# import api_key from config file
from config import api_key
# +
# Create a settings object with your API key and preferred units
settings = {"units": "metric", "appid": api_key}
# +
# Get data for each city in cities.csv
city_data = pd.read_csv("Resources/cities.csv", header=None)
city_names = city_data[0].tolist()
city_names
city_temp = []
for city in city_names:
current_weather = owm.get_current(city, **settings)
#print(f"Current weather object for {city}: {current_weather}.")
city_temp.append(current_weather)
print(city_temp)
# -
# Create an "extracts" object to get the temperature, latitude,
# and longitude in each city
summary = ["name", "main.temp", "coord.lat"]
for city in city_temp:
data = city_temp[0](*summary)
print(f"The current weather summary for Paris is: {data}.")
# Create a Pandas DataFrame with the results
# BONUS:
# # Instructor Turn Activity 11 World Bank
# +
# Dependencies
import requests
url = "http://api.worldbank.org/v2/"
format = "json"
# Get country information in JSON format
countries_response = requests.get(f"{url}countries?format={format}").json()
print(countries_response)
# First element is general information, second is countries themselves
countries = countries_response[1]
# -
# Report the names
for country in countries:
print(country["name"])
# # Students Turn Activity 11 Lending Types
#
# This activity provides an opportunity to practice making two API calls in sequence in which the second API call depends on the response of the first.
#
# ## Instructions
#
# * Retrieve a list of the lending types the world bank keeps track of, and extract the ID key from each of them.
#
# * Next, determine how many countries are categorized under each lending type. Use a dict to store this information.
#
# * This data is stored as the first element of the response array.
#
# * Finally, print the number of countries of each lending type.
# +
# Dependencies
import requests
url = "http://api.worldbank.org/v2/"
format = "json"
# +
# Get the list of lending types the world bank has
# CODE HERE
lending_types = requests.get(f"{url}lendingTypes?format={format}").json()
types = lending_types[1]
for type in types:
print(type["value"])
# -
# Next, determine how many countries fall into each lending type.
# Hint: Look at the first element of the response array.
# CODE HERE
IBRD = []
Blend = []
IDA = []
Not_classified = []
#print(countries)
for country in countries:
if country["lendingType"]["value"] == "IBRD":
IBRD.append(country["name"])
elif country["lendingType"]["value"] == "Blend":
Blend.append(country["name"])
elif country["lendingType"]["value"] == "IDA":
IDA.append(country["name"])
else:
Not_classified.append(country["name"])
print(IBRD, Blend, IDA, Not_classified)
# Print the number of countries of each lending type
# CODE HERE
# # Instructor Turn 12 Activity CityPy
# !pip install openweathermapy
# Dependencies
from citipy import citipy
# Some random coordinates
coordinates = [(200, 200), (23, 200), (42, 100)]
cities = []
for coordinate_pair in coordinates:
lat, lon = coordinate_pair
cities.append(citipy.nearest_city(lat, lon))
for city in cities:
country_code = city.country_code
name = city.city_name
print(f"The country code of {name} is '{country_code}'.")
|
Activities Week 6 (API)/Python_Api_Part2/Day2/Day2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] nbgrader={}
# # Numpy Exercise 3
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
# + nbgrader={}
import antipackage
import github.ellisonbg.misc.vizarray as va
# + [markdown] nbgrader={}
# ## Geometric Brownian motion
# + [markdown] nbgrader={}
# Here is a function that produces standard Brownian motion using NumPy. This is also known as a [Wiener Process](http://en.wikipedia.org/wiki/Wiener_process).
# + nbgrader={}
def brownian(maxt, n):
"""Return one realization of a Brownian (Wiener) process with n steps and a max time of t."""
t = np.linspace(0.0,maxt,n)
h = t[1]-t[0]
Z = np.random.normal(0.0,1.0,n-1)
dW = np.sqrt(h)*Z
W = np.zeros(n)
W[1:] = dW.cumsum()
return t, W
# + [markdown] nbgrader={}
# Call the `brownian` function to simulate a Wiener process with `1000` steps and max time of `1.0`. Save the results as two arrays `t` and `W`.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "grade": false, "grade_id": "numpyex03a", "points": 2, "solution": true}
t = brownian(1.0, 1000)[0]
W = brownian(1.0, 1000)[1]
# + deletable=false nbgrader={"checksum": "b671a523fd8cb7621c2445244189d5a4", "grade": true, "grade_id": "numpyex03a", "points": 2}
assert isinstance(t, np.ndarray)
assert isinstance(W, np.ndarray)
assert t.dtype==np.dtype(float)
assert W.dtype==np.dtype(float)
assert len(t)==len(W)==1000
# + [markdown] nbgrader={}
# Visualize the process using `plt.plot` with `t` on the x-axis and `W(t)` on the y-axis. Label your x and y axes.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "grade": false, "grade_id": "numpyex03b", "points": 2, "solution": true}
plt.plot(t, W)
plt.xlabel('t')
plt.ylabel('W(t)')
# + deletable=false nbgrader={"checksum": "1a35840ca7eaf864f9201ee4e0d947e0", "grade": true, "grade_id": "numpyex03b", "points": 2}
assert True # this is for grading
# + [markdown] nbgrader={}
# Use `np.diff` to compute the changes at each step of the motion, `dW`, and then compute the mean and standard deviation of those differences.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "grade": false, "grade_id": "numpyex03c", "points": 2, "solution": true}
dW = np.diff(W)
print(dW.mean(), dW.std())
# + deletable=false nbgrader={"checksum": "b2236af662ecc138c4b78af673b476c1", "grade": true, "grade_id": "numpyex03c", "points": 2}
assert len(dW)==len(W)-1
assert dW.dtype==np.dtype(float)
# + [markdown] nbgrader={}
# Write a function that takes $W(t)$ and converts it to geometric Brownian motion using the equation:
#
# $$
# X(t) = X_0 e^{((\mu - \sigma^2/2)t + \sigma W(t))}
# $$
#
# Use Numpy ufuncs and no loops in your function.
# + nbgrader={"checksum": "2b05883af2c87bc938fc4f7fe7e35f66", "grade": false, "grade_id": "numpyex03d", "points": 2, "solution": true}
def geo_brownian(t, W, X0, mu, sigma):
"Return X(t) for geometric brownian motion with drift mu, volatility sigma."""
X = X0 * np.exp(((mu-sigma**2)/2)*t + sigma*W)
return X
# + deletable=false nbgrader={"checksum": "401ffd490410ab0a18612d641e24c02f", "grade": true, "grade_id": "numpyex03d", "points": 2}
assert True # leave this for grading
# + [markdown] nbgrader={}
# Use your function to simulate geometric brownian motion, $X(t)$ for $X_0=1.0$, $\mu=0.5$ and $\sigma=0.3$ with the Wiener process you computed above.
#
# Visualize the process using `plt.plot` with `t` on the x-axis and `X(t)` on the y-axis. Label your x and y axes.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "grade": false, "grade_id": "numpyex03f", "points": 2, "solution": true}
X = geo_brownian(t, W, 1.0, 0.5, 0.3)
plt.plot(t, X)
plt.xlabel('t')
plt.ylabel('X(t)')
# + deletable=false nbgrader={"checksum": "00e3fda54f3eba73d67842cf7f02777a", "grade": true, "grade_id": "numpyex03e", "points": 2}
assert True # leave this for grading
|
assignments/assignment03/NumpyEx03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Network visualization
#
# This notebook constructs a network visualization connecting bacterial species to KEGG pathways.
# Parameters
d_col = 'C(Advanced_Stage_label)[T.Local]' # node color
ew = "rank" # edge weight
taxa_level = 'Rank6' # taxonomy level
# +
# Preliminaries
# %matplotlib inline
import numpy as np
import matplotlib.mlab as mlab
import matplotlib.pyplot as plt
import seaborn as sns
from IPython.display import display, HTML
def widen_notebook():
display(HTML("<style>.container { width:100% !important; }</style>"))
widen_notebook()
# -
# data files
# !ls ../data/edges/lung-cancer
edges_txt = "../data/edges/lung-cancer/Cancer_related_path1_edges.txt"
kegg_txt = "../data/edges/lung-cancer/LC_KO_metadata.txt"
microbes_txt = "../data/edges/lung-cancer/microbe-metadata.txt"
full_microbes_biom = "../data/edges/lung-cancer/Microbiome_with_paired_RNA_otu_table.a1.biom.txt"
filtered_microbes_biom = "../data/edges/lung-cancer/microbes.txt"
# +
# Read data files into lists of dictionaries
def split_commas(line):
return line.strip().split("\t")
def CSVtodicts(filename):
f = open(filename)
result = []
headers = split_commas(f.readline())
for line in f.readlines():
values = split_commas(line)
dictionary = dict(zip(headers, values))
result.append(dictionary)
return result
edges = CSVtodicts(edges_txt)
keggs = CSVtodicts(kegg_txt)
microbes = CSVtodicts(microbes_txt)
len(edges), len(keggs), len(microbes)
# +
import pandas as pd
microbes = pd.read_table(microbes_txt)
keggs = pd.read_table(kegg_txt)
edges = pd.read_table(edges_txt)
full_microbe_counts = pd.read_table(full_microbes_biom, skiprows=1, index_col=0)
filtered_microbe_counts = pd.read_table(filtered_microbes_biom, skiprows=1, index_col=0)
# scrub dataframe
featureid = '#SampleID'
# -
microbes['abbv_name'] = microbes.apply(lambda x: '%s%d' % (x[taxa_level], x['#SampleID']), axis=1)
taxa = full_microbe_counts.iloc[:, -1]
microbe_counts = full_microbe_counts.iloc[:, :-1]
microbe_counts.shape, filtered_microbe_counts.shape
microbe_counts = microbe_counts.loc[:, filtered_microbe_counts.columns]
sns.distplot(microbe_counts.sum(axis=0))
microbe_props = microbe_counts.apply(lambda x: x / x.sum(), axis=0)
microbe_props = microbe_props.loc[microbes['#SampleID']].T
mean_microbe_abundance = np.log(microbe_props.mean(axis=0))
norm_microbe_abundance = (mean_microbe_abundance - mean_microbe_abundance.min()) / (mean_microbe_abundance.max() - mean_microbe_abundance.min())
# +
fontmin = 8
fontmax = 30
fontsize = norm_microbe_abundance * (fontmax - fontmin) + fontmin
sns.distplot(norm_microbe_abundance)
# -
select_microbes = list(set(edges.src.values))
select_kegg = list(set(edges.dest.values))
microbes.head()
def abbreviate(x):
return x.split('|')[-1]
abbreviate('k__Viruses|p__Viruses_noname|c__Viruses_noname|o__Viruses_noname')
# +
edges['src'] = edges.src.apply(lambda x: int(x.replace('\"', '')))
edges['dest'] = edges.dest.apply(lambda x: x.replace('\"', ''))
microbe_dicts = microbes.T.to_dict().values()
kegg_dicts = keggs.T.to_dict().values()
edge_dicts = edges.T.to_dict().values()
# -
microbe_metadata = microbes.set_index('#SampleID')
# scrub edges, because of R ...
edges['src_abbv'] = edges.src.apply(lambda x: microbe_metadata.loc[x, 'abbv_name'])
# +
# name abbreviation mappings.
def abbreviate(d):
return '%s%d' % (d[taxa_level], d[featureid])
def microbe_name_dict(dicts):
return dict([abbreviate(d), d] for d in dicts)
name2microbe = microbe_name_dict(microbe_dicts)
name2microbe.items()[15]
# -
def kegg_name_dict(dicts):
return dict([d['#OTUID'], d] for d in dicts)
name2kegg = kegg_name_dict(kegg_dicts)
name2kegg.items()[32]
# Construct the network graph from the edges.
from jp_gene_viz import dGraph
G = dGraph.WGraph()
for e in edge_dicts:
name = microbe_metadata.loc[e['src'], 'abbv_name']
G.add_edge(name, e["dest"], e[ew], e)
# Construct the network widget from the graph
from jp_gene_viz import dNetwork
dNetwork.load_javascript_support()
N = dNetwork.NetworkDisplay()
N.load_data(G)
import matplotlib.colors as colors
class MidpointNormalize(colors.Normalize):
def __init__(self, vmin=None, vmax=None, vcenter=None, clip=False):
self.vcenter = vcenter
colors.Normalize.__init__(self, vmin, vmax, clip)
def __call__(self, value, clip=None):
# I'm ignoring masked values and all kinds of edge cases to make a
# simple example...
x, y = [self.vmin, self.vcenter, self.vmax], [0, 0.5, 1]
return np.ma.masked_array(np.interp(value, x, y))
import matplotlib as mpl
from matplotlib.colors import rgb2hex
# TODO: make parameter
# https://matplotlib.org/3.1.0/tutorials/colors/colormaps.html
#cmap = plt.get_cmap('RdYlGn')
cmap = plt.get_cmap('PiYG')
edge_cmap = plt.get_cmap('Greys')
microbe_norm = MidpointNormalize(vmin=-2., vcenter=0, vmax=1.5)
#microbe_norm = mpl.colors.Normalize(vmin=0, vmax=1)
kegg_norm = mpl.colors.Normalize(vmin=-2, vmax=2)
edges.head()
edge_lookup = edges.set_index(['src_abbv', 'dest'])
# +
# Configure and display the network
# TODO: remove congested labels
N.labels_button.value = True
N.size_slider.value = 1000
# TODO: add node size / font size are variable features
# TODO: swap circles with squares
# TODO: light grey dashed lines for low probability edges
# TODO: add edge weight size
# TODO: remove labels programmatically
# TODO: allow for edges to be colored on a gradient (i.e. greys)
# main goals
# 1. focus on pathways of interest
# 2. advance vs local
# 3. weighted by relative abundance
# colorize the nodes based on weights (hacky, sorry)
dg = N.display_graph
for node_name in dg.node_weights:
svg_name = dg.node_name(node_name)
if node_name in name2microbe:
d = name2microbe[node_name]
value = name2microbe[node_name][d_col]
if np.isnan(value):
value = 0
node_color = rgb2hex(cmap(microbe_norm(value))[:3])
value = norm_microbe_abundance.loc[d['#SampleID']]
#node_color = rgb2hex(cmap(microbe_norm(value))[:3])
N.override_node(node_name, color=node_color, radius=value*10, shape='circle')
N.override_label(node_name, hide=value < 0.3, font_size=value*18)
else:
N.override_node(node_name, shape='rect', radius=5, color='#6193F7')
N.override_label(node_name, font_size=18)
for src, dest in dg.edge_weights:
m = np.log(edges['rank']).min()
p = edge_lookup.loc[(src, dest), 'rank']
width = np.log(p - m)/10
N.override_edge(src, dest, color=rgb2hex(edge_cmap(p+0.1)), stroke_width=width)
# show labels
N.labels_button.value = True
# rerun the layout
N.layout_click()
# draw the network with the new colors and sizes
N.draw()
# show the network
N.show()
# -
sns.distplot(edges['rank'])
cmap(microbe_norm(value))
cmap(microbe_norm(-2))
|
notebooks/cancer-pathway1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="XQbgW2ifjSkB" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
# + id="fhXEgJCf1pNi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 180} outputId="637e2232-9f09-466f-d545-98cfecf69a9c" executionInfo={"status": "error", "timestamp": 1581619027648, "user_tz": -60, "elapsed": 478, "user": {"displayName": "<NAME>17bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
# + id="GIR4snOSj-DA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a62cc3b7-9ac1-45d6-8ff5-16ac53616a52" executionInfo={"status": "ok", "timestamp": 1581616878378, "user_tz": -60, "elapsed": 737, "user": {"displayName": "<NAME>17bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
# cd "/content/drive/My Drive/Colab Notebooks/dw-matrix"
# + id="4908m8RpkR61" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2d60b414-9234-40c2-a2eb-ac55a7441c87" executionInfo={"status": "ok", "timestamp": 1581616881915, "user_tz": -60, "elapsed": 2190, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
# ls data
# + id="5_wWupjVkVzd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ebeaa40e-3a26-480e-e7c9-0d435ff6f517" executionInfo={"status": "ok", "timestamp": 1581616894655, "user_tz": -60, "elapsed": 2296, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
df = pd.read_csv("data/men_shoes.csv", low_memory=False)
df.shape
# + id="vaHRS015kmlk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="cdc2fd2d-9d03-4ccf-8c6d-a60dc5215a2e" executionInfo={"status": "ok", "timestamp": 1581616896878, "user_tz": -60, "elapsed": 493, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
df.columns
# + id="vYD9vmRplLQ9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c7483e60-9e57-453c-e6d9-5639a3cd2088" executionInfo={"status": "ok", "timestamp": 1581616918176, "user_tz": -60, "elapsed": 475, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
mean_price = np.mean( df['prices_amountmin'] )
mean_price
# + id="PlEsz_ySn6cJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e116bfbd-9d48-44f5-e910-b47f53b81af6" executionInfo={"status": "ok", "timestamp": 1581617257301, "user_tz": -60, "elapsed": 540, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
y_true = df['prices_amountmin'] # to jest nasza wartość prawidłowa
y_pred = [mean_price] * y_true.shape[0] # y_pred to wartośc którą chcemy sprognozować
mean_absolute_error(y_true, y_pred) # tutaj sprawdzamy jak radzi sobie nasz model - o ile $ się myli - im mniej tym lepiej
# + id="UGmyyUgAueS6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="7aec93b6-81d7-4c7d-8a35-0c490b6a26e9" executionInfo={"status": "ok", "timestamp": 1581617487489, "user_tz": -60, "elapsed": 934, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
df['prices_amountmin'].hist(bins=100)
# + id="LrDtjKgyv5oB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="1d831720-1310-4b29-80a5-8a5c1e871ccf" executionInfo={"status": "ok", "timestamp": 1581617630381, "user_tz": -60, "elapsed": 723, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
np.log1p( df['prices_amountmin'] ).hist(bins=100) # zmieniamy wykres na funkcję logarytmiczną
# + id="ZdPXCPIUwPx5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="69d3aaf4-5568-4862-de75-e3e20bf78b8a" executionInfo={"status": "ok", "timestamp": 1581617748353, "user_tz": -60, "elapsed": 505, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
y_true = df['prices_amountmin'] # to jest nasza wartość prawidłowa
y_pred = [np.median(y_true)] * y_true.shape[0] # y_pred to wartośc którą chcemy sprognozować
mean_absolute_error(y_true, y_pred)
# + id="G6M238Zow5aY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7d47f322-e81f-4adc-a523-fc4e3cf29c0d" executionInfo={"status": "ok", "timestamp": 1581617911502, "user_tz": -60, "elapsed": 1804, "user": {"displayName": "<NAME>017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
[np.median(y_true)]
# + id="2SmxrUD2xg72" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2797b021-3de1-411f-b9e4-c2dd2111fd1d" executionInfo={"status": "ok", "timestamp": 1581618368357, "user_tz": -60, "elapsed": 643, "user": {"displayName": "<NAME>17bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
# teraz robimy transformację logarytmiczną
y_true = df['prices_amountmin'] # to jest nasza wartość prawidłowa
price_log_mean = np.expm1( np.mean(np.log1p(y_true) ) ) # najpierw nasz y_true przepuszczamy przez funkcję logarytmiczną,
# a następnie przez funkcję wartość średnia i mamy średnią wartość wartości zlogarytmizowanej
y_pred = [price_log_mean] * y_true.shape[0] # y_pred to wartośc którą chcemy sprognozować
mean_absolute_error(y_true, y_pred)
# + id="He6BH-L3zQtf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="978f4556-0034-422c-cb34-89da8990d477" executionInfo={"status": "ok", "timestamp": 1581618463639, "user_tz": -60, "elapsed": 625, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
df.columns
# + id="bKWk4BFzzn-O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="b92e6836-06e1-4164-cb5c-e6b8d66592eb" executionInfo={"status": "ok", "timestamp": 1581618501655, "user_tz": -60, "elapsed": 654, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
df.brand.value_counts()
# + id="1u6SN4tgzuy4" colab_type="code" colab={}
# przypisujemy unikalne id do wartości tekstowych
df['brand_cat'] = df['brand'].factorize()[0] # mamy dwa tuple, ale nam jest potrzebny tylko jeden, dlatego wybieram peirwsyz z listy [0]
# + id="xX2xTNZD0QRt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f970ac7e-97c9-449e-d72e-1debe1ef08e8" executionInfo={"status": "ok", "timestamp": 1581619241353, "user_tz": -60, "elapsed": 553, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/<KEY>_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
# X to macierz którą będziemy przekazywać, składa się z cech (kolumn) - tutaj jedna cecha czyli brand i wierszy
feats = ['brand_cat'] # tutaj mamy listę cech
X = df[ feats]
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, scoring='neg_mean_absolute_error')
np.mean(scores), np.std(scores)
# + id="kgHNfnEJ1TVO" colab_type="code" colab={}
# tutaj tworzymy funkcję run_model, żeby za każdym razem nie musiec kopiować kodu
def run_model(feats):
X = df[ feats]
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="7jVN4s5z2-g7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6a00ed35-5a01-4aea-8470-fa2db1fedce6" executionInfo={"status": "ok", "timestamp": 1581619421046, "user_tz": -60, "elapsed": 598, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
run_model (['brand_cat'])
# + id="zWqh5dXw3RxW" colab_type="code" colab={}
# manufacturer
df['manufacturer_cat'] = df['manufacturer'].factorize()[0]
# + id="Z9mBdkDx3zKP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a8403998-5fc6-4d6b-e325-e8cca5ba07af" executionInfo={"status": "ok", "timestamp": 1581619576483, "user_tz": -60, "elapsed": 760, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
run_model (['manufacturer_cat'])
# + id="__VMa20Y33rN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f9a29f68-a9d7-46a0-b236-8bae597bdab4" executionInfo={"status": "ok", "timestamp": 1581619620449, "user_tz": -60, "elapsed": 586, "user": {"displayName": "Leszek \u017bukowski", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCd_cyF11i8J2zx_Xa8Vis0E3EwhFXmS28Gwq2y=s64", "userId": "12512059599918919065"}}
run_model (['brand_cat', 'manufacturer_cat'])
# + id="LaaqQy_k4CdF" colab_type="code" colab={}
|
matrix_one/day4.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
//
// # Load PyTorch model
//
// In this tutorial, you learn how to load an existing PyTorch model and use it to run a prediction task.
//
// We will run the inference in DJL way with [example](https://pytorch.org/hub/pytorch_vision_resnet/) on the pytorch official website.
//
//
// ## Preparation
//
// This tutorial requires the installation of Java Kernel. For more information on installing the Java Kernel, see the [README](https://github.com/awslabs/djl/blob/master/jupyter/README.md).
// +
// %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
// %maven ai.djl:api:0.6.0-SNAPSHOT
// %maven ai.djl.pytorch:pytorch-engine:0.6.0-SNAPSHOT
// %maven org.slf4j:slf4j-api:1.7.26
// %maven org.slf4j:slf4j-simple:1.7.26
// %maven net.java.dev.jna:jna:5.3.0
// See https://github.com/awslabs/djl/blob/master/pytorch/pytorch-engine/README.md
// for more PyTorch library selection options
// %maven ai.djl.pytorch:pytorch-native-auto:1.5.0
// -
import java.awt.image.*;
import ai.djl.*;
import ai.djl.inference.*;
import ai.djl.modality.*;
import ai.djl.modality.cv.*;
import ai.djl.modality.cv.util.*;
import ai.djl.modality.cv.transform.*;
import ai.djl.modality.cv.translator.*;
import ai.djl.repository.zoo.*;
import ai.djl.translate.*;
import ai.djl.training.util.*;
// ## Step 1: Prepare your model
//
// This tutorial assumes that you have a TorchScript model.
// DJL only supports the TorchScript format for loading models from PyTorch, so other models will need to be [converted](https://github.com/awslabs/djl/blob/master/docs/pytorch/how_to_convert_your_model_to_torchscript.md).
// A TorchScript model includes the model structure and all of the parameters.
//
// We will be using a pre-trained `resnet18` model. First, use the `DownloadUtils` to download the model files and save them in the `build/pytorch_models` folder
DownloadUtils.download("https://djl-ai.s3.amazonaws.com/mlrepo/model/cv/image_classification/ai/djl/pytorch/resnet/0.0.1/traced_resnet18.pt.gz", "build/pytorch_models/resnet18/resnet18.pt", new ProgressBar());
// In order to do image classification, you will also need the synset.txt which stores the classification class labels. We will need the synset containing the Imagenet labels with which resnet18 was originally trained.
DownloadUtils.download("https://djl-ai.s3.amazonaws.com/mlrepo/model/cv/image_classification/ai/djl/pytorch/synset.txt", "build/pytorch_models/resnet18/synset.txt", new ProgressBar());
// ## Step 2: Create a Translator
//
// We will create a transformation pipeline which maps the transforms shown in the [PyTorch example](https://pytorch.org/hub/pytorch_vision_resnet/).
// ```python
// ...
// preprocess = transforms.Compose([
// transforms.Resize(256),
// transforms.CenterCrop(224),
// transforms.ToTensor(),
// transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]),
// ])
// ...
// ```
//
// Then, we will use this pipeline to create the [`Translator`](https://javadoc.io/static/ai.djl/api/0.5.0/index.html?ai/djl/translate/Translator.html)
// +
Pipeline pipeline = new Pipeline();
pipeline.add(new Resize(256))
.add(new CenterCrop(224, 224))
.add(new ToTensor())
.add(new Normalize(
new float[] {0.485f, 0.456f, 0.406f},
new float[] {0.229f, 0.224f, 0.225f}));
Translator<Image, Classifications> translator = ImageClassificationTranslator.builder()
.setPipeline(pipeline)
.optApplySoftmax(true)
.build();
// -
// ## Step 3: Load your model
//
// Next, we will set the model zoo location to the `build/pytorch_models` directory we saved the model to. You can also create your own [`Repository`](https://javadoc.io/static/ai.djl/repository/0.5.0/index.html?ai/djl/repository/Repository.html) to avoid manually managing files.
//
// Next, we add some search criteria to find the resnet18 model and load it.
// +
// Search for models in the build/pytorch_models folder
System.setProperty("ai.djl.repository.zoo.location", "build/pytorch_models/resnet18");
Criteria<Image, Classifications> criteria = Criteria.builder()
.setTypes(Image.class, Classifications.class)
// only search the model in local directory
// "ai.djl.localmodelzoo:{name of the model}"
.optArtifactId("ai.djl.localmodelzoo:resnet18")
.optTranslator(translator)
.optProgress(new ProgressBar()).build();
ZooModel model = ModelZoo.loadModel(criteria);
// -
// ## Step 4: Load image for classification
//
// We will use a sample dog image to run our prediction on.
var img = ImageFactory.getInstance().fromUrl("https://github.com/pytorch/hub/raw/master/dog.jpg");
img.getWrappedImage()
// ## Step 5: Run inference
//
// Lastly, we will need to create a predictor using our model and translator. Once we have a predictor, we simply need to call the predict method on our test image.
// +
Predictor<Image, Classifications> predictor = model.newPredictor();
Classifications classifications = predictor.predict(img);
classifications
// -
// ## Summary
//
// Now, you can load any TorchScript model and run inference using it.
//
// You might also want to check out [load_mxnet_model.ipynb](https://github.com/awslabs/djl/blob/master/jupyter/load_mxnet_model.ipynb) which demonstrates loading a local model directly instead of through the Model Zoo API.
// To optimize inference performance, you might check out [how_to_optimize_inference_performance](https://github.com/awslabs/djl/blob/master/docs/pytorch/how_to_optimize_inference_performance.md).
|
jupyter/load_pytorch_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install plotly
# !pip install chart_studio
# ## Select stock symbol and date range
symbol = 'NETE'
start_date = "2020-12-11"
end_date = "2020-12-12"
anchored_vwaps_start = ['2020-12-11 09:30:00', '2020-12-11 14:20:00']
# #### Imports
# silence warnings
import warnings
import numpy as np
import pandas as pd
import pytz
import plotly.graph_objects as go
import chart_studio.plotly as py
from datetime import datetime
import alpaca_trade_api as tradeapi
from liualgotrader.common.market_data import get_symbol_data
from liualgotrader.fincalcs.vwap import anchored_vwap
from liualgotrader.fincalcs.resample import resample, ResampleRangeType
# %matplotlib inline
warnings.filterwarnings("ignore")
est = pytz.timezone("US/Eastern")
# #### Load symbol data
start_date = est.localize(datetime.strptime(start_date, "%Y-%m-%d"))
end_date = est.localize(datetime.strptime(end_date, "%Y-%m-%d"))
ohlc_data = get_symbol_data(symbol, start_date, end_date)
ohlc_data
if ohlc_data is None or ohlc_data.empty:
assert False, "No data loaded"
# ## Visuals
# +
anchored_vwap_starts = [
datetime.strptime(anchored_vwap_start, "%Y-%m-%d %H:%M:%S").replace(tzinfo=est)
for anchored_vwap_start in anchored_vwaps_start
]
org_ohlc_data = ohlc_data
for sample_rate in (
ResampleRangeType.min_1,
ResampleRangeType.min_5,
ResampleRangeType.min_15,
):
ohlc_data = resample(org_ohlc_data, sample_rate)
anchored_vwap_indicators = [
anchored_vwap(ohlc_data, anchored_vwap_start)
for anchored_vwap_start in anchored_vwap_starts
]
trace1 = {
"x": ohlc_data.index,
"open": ohlc_data.open,
"high": ohlc_data.high,
"low": ohlc_data.low,
"close": ohlc_data.close,
"type": "candlestick",
"name": f"{symbol} {sample_rate} bars",
"yaxis": "y2",
"showlegend": True,
}
trace2 = [
{
"x": anchored_vwap_indicator.index,
"y": anchored_vwap_indicator,
"type": "scatter",
"mode": "lines",
"line": {"width": 2, "color": "black"},
"yaxis": "y2",
"name": f"VWAP-{indx+1}",
"showlegend": True,
}
for indx, anchored_vwap_indicator in enumerate(anchored_vwap_indicators)
]
fig = dict(data=[trace1], layout=dict())
fig["layout"]["plot_bgcolor"] = "rgb(200, 200, 200)"
fig["layout"]["xaxis"] = dict(rangeselector=dict(visible=True))
fig["layout"]["yaxis"] = dict(domain=[0, 0.2], showticklabels=False)
fig["layout"]["yaxis2"] = dict(domain=[0.2, 0.8])
fig["layout"]["legend"] = dict(
orientation="h",
y=0.95,
x=0.3,
yanchor="bottom",
)
fig["layout"]["margin"] = dict(t=40, b=40, r=40, l=40)
rangeselector = dict(
# visibe = True,
x=0,
y=0.9,
bgcolor="rgba(150, 200, 250, 0.4)",
font=dict(size=13),
buttons=list(
[
dict(count=1, label="1 yr", step="year"),
dict(count=3, label="3 mo", step="month", stepmode="backward"),
dict(count=1, label="1 mo", step="month", stepmode="backward"),
dict(count=7, label="1 wk", step="day", stepmode="backward"),
dict(step="all"),
]
),
)
fig["layout"]["xaxis"]["rangeselector"] = rangeselector
fig["data"] += trace2
colors = []
for i in range(len(ohlc_data.close)):
if i != 0:
if ohlc_data.close[i] > ohlc_data.close[i - 1]:
colors.append("green")
else:
colors.append("red")
else:
colors.append("red")
fig["data"].append(
dict(
x=ohlc_data.index,
y=ohlc_data.volume,
marker=dict(color=colors),
type="bar",
yaxis="y",
name="Volume",
showlegend=False,
)
)
f = go.Figure(data=fig["data"], layout=fig["layout"])
f.show()
|
analysis/notebooks/Labs/anchored-vwap-lab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Tutorial 5: Model
#
# ## Overview
#
# In this tutorial we will cover:
#
# * [Instantiating and Compiling a Model](#t05compile)
# * [The Model Function](#t05model)
# * [Custom Models](#t05custom)
# * [FastEstimator Models](#t05fe)
# * [Pre-Trained Models](#t05trained)
# * [The Optimizer Function](#t05optimizer)
# * [Loading Model Weights](#t05weights)
# * [Specifying a Model Name](#t05name)
# * [Related Apphub Examples](#t05apphub)
# <a id='t05compile'></a>
# ## Instantiating and Compiling a model
#
# We need to specify two things to instantiate and compile a model:
# * model_fn
# * optimizer_fn
#
# Model definitions can be implemented in Tensorflow or Pytorch and instantiated by calling **`fe.build`** which constructs a model instance and associates it with the specified optimizer.
# <a id='t05model'></a>
# ## Model Function
#
# `model_fn` should be a function/lambda function which returns either a `tf.keras.Model` or `torch.nn.Module`. FastEstimator provides several ways to specify the model architecture:
#
# * Custom model architecture
# * Importing a pre-built model architecture from FastEstimator
# * Importing pre-trained models/architectures from PyTorch or TensorFlow
# <a id='t05custom'></a>
# ### Custom model architecture
# Let's create a custom model in TensorFlow and PyTorch for demonstration.
# +
# Some preliminary imports
import tensorflow as tf
# Since we will be mixing TF and Torch in the tutorial, we need to stop TF from taking all of the GPU memory.
# Normally you would pick either TF or Torch, so you don't need to worry about this.
physical_devices = tf.config.list_physical_devices('GPU')
for device in physical_devices:
try:
tf.config.experimental.set_memory_growth(device, True)
except:
pass
import torch
import torch.nn as nn
import fastestimator as fe
# -
# #### tf.keras.Model
# +
def my_model_tf(input_shape=(30, ), num_classes=2):
model = tf.keras.Sequential()
model.add(tf.keras.layers.Dense(32, activation="relu", input_shape=input_shape))
model.add(tf.keras.layers.Dense(8, activation="relu"))
model.add(tf.keras.layers.Dense(num_classes, activation="softmax"))
return model
model_tf = fe.build(model_fn=my_model_tf, optimizer_fn="adam")
# -
# #### torch.nn.Module
# +
class my_model_torch(nn.Module):
def __init__(self, num_inputs=30, num_classes=2):
super().__init__()
self.layers = nn.Sequential(nn.Linear(num_inputs, 32),
nn.ReLU(inplace=True),
nn.Linear(32, 8),
nn.ReLU(inplace=True),
nn.Linear(8, num_classes))
def forward(self, x):
x = self.layers(x)
x_label = torch.softmax(x, dim=-1)
return x_label
model_torch = fe.build(model_fn=my_model_torch, optimizer_fn="adam")
# -
# <a id='t05fe'></a>
# ### Importing model architecture from FastEstimator
#
# Below we import a PyTorch LeNet architecture from FastEstimator. See our [Architectures](../../fastestimator/architecture) folder for a full list of the architectures provided by FastEstimator.
# +
from fastestimator.architecture.pytorch import LeNet
# from fastestimator.architecture.tensorflow import LeNet # One can also use a TensorFlow model
model = fe.build(model_fn=LeNet, optimizer_fn="adam")
# -
# <a id='t05trained'></a>
# ### Importing pre-trained models/architectures from PyTorch or TensorFlow
#
# Below we show how to define a model function using a pre-trained resnet model provided by TensorFlow and PyTorch respectively. We load the pre-trained models using a lambda function.
# #### Pre-trained model from tf.keras.applications
resnet50_tf = fe.build(model_fn=lambda: tf.keras.applications.ResNet50(weights='imagenet'), optimizer_fn="adam")
# #### Pre-trained model from torchvision
# +
from torchvision import models
resnet50_torch = fe.build(model_fn=lambda: models.resnet50(pretrained=True), optimizer_fn="adam")
# -
# <a id='t05optimizer'></a>
# ## Optimizer function
#
# `optimizer_fn` can be a string or lambda function.
#
# ### Optimizer from String
# Specifying a string for the `optimizer_fn` loads the optimizer with default parameters. The optimizer strings accepted by FastEstimator are as follows:
# - Adadelta: 'adadelta'
# - Adagrad: 'adagrad'
# - Adam: 'adam'
# - Adamax: 'adamax'
# - RMSprop: 'rmsprop'
# - SGD: 'sgd'
# ### Optimizer from Function
#
# To specify specific values for the optimizer learning rate or other parameters, we need to pass a lambda function to the `optimizer_fn`.
# +
# TensorFlow
model_tf = fe.build(model_fn=my_model_tf, optimizer_fn=lambda: tf.optimizers.Adam(1e-4))
# PyTorch
model_torch = fe.build(model_fn=my_model_torch, optimizer_fn=lambda x: torch.optim.Adam(params=x, lr=1e-4))
# -
# If a model function returns multiple models, a list of optimizers can be provided. See the **[pggan apphub](../../apphub/image_generation/pggan/pggan.ipynb)** for an example with multiple models and optimizers.
# <a id='t05weights'></a>
# ## Loading model weights
#
# We often need to load the weights of a saved model. Model weights can be loaded by specifying the path of the saved weights using the `weights_path` parameter. Let's use the resnet models created earlier to showcase this.
# #### Saving model weights
# Here, we create a temporary directory and use FastEstimator backend to save the weights of our previously created resnet50 models:
# +
import os
import tempfile
model_dir = tempfile.mkdtemp()
# TensorFlow
fe.backend.save_model(resnet50_tf, save_dir=model_dir, model_name= "resnet50_tf")
# PyTorch
fe.backend.save_model(resnet50_torch, save_dir=model_dir, model_name= "resnet50_torch")
# -
# #### Loading weights for TensorFlow and PyTorch models
# TensorFlow
resnet50_tf = fe.build(model_fn=lambda: tf.keras.applications.ResNet50(weights=None),
optimizer_fn="adam",
weights_path=os.path.join(model_dir, "resnet50_tf.h5"))
# PyTorch
resnet50_torch = fe.build(model_fn=lambda: models.resnet50(pretrained=False),
optimizer_fn="adam",
weights_path=os.path.join(model_dir, "resnet50_torch.pt"))
# <a id='t05name'></a>
# ## Specifying a Model Name
#
# The name of a model can be specified using the `model_name` parameter. The name of the model is helpful in distinguishing models when multiple are present.
model = fe.build(model_fn=LeNet, optimizer_fn="adam", model_name="LeNet")
print("Model Name: ", model.model_name)
# If a model function returns multiple models, a list of model_names can be given. See the **[pggan apphub](../../apphub/image_generation/pggan/pggan.ipynb)** for an illustration with multiple models and model names.
# <a id='t05apphub'></a>
# ## Apphub Examples
# You can find some practical examples of the concepts described here in the following FastEstimator Apphubs:
#
# * [PG-GAN](../../apphub/image_generation/pggan/pggan.ipynb)
# * [Uncertainty Weighted Loss](../../apphub/multi_task_learning/uncertainty_weighted_loss/uncertainty_loss.ipynb)
|
tutorial/beginner/t05_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (fastai)
# language: python
# name: fastai
# ---
# # Train a 'DESCRIPTION' classifier
#
# Instead of building from a MIMIC trained language model, or a Wikitext-103 model, start with blank AWD-LSTM model
from fastai.text import *
from sklearn.model_selection import train_test_split
import glob
import gc
# Setup filenames and paths
# +
# pandas doesn't understand ~, so provide full path
base_path = Path.home() / 'mimic'
# files used during processing - all aggregated here
admissions_file = base_path/'ADMISSIONS.csv'
notes_file = base_path/'NOTEEVENTS.csv'
class_file = 'cl_data.pickle'
notes_pickle_file = base_path/'noteevents.pickle'
init_model_file = base_path/'cl_head'
cycles_file = base_path/'cl_num_iterations.pickle'
enc_file = 'cl_enc'
descr_ft_file = 'cl_fine_tuned_'
training_history_file = 'no_ft_cl_history'
transformer_training_history_file = 'no_ft_cl_tfr_history'
transformerxl_training_history_file = 'no_ft_cl_trfxl_history'
# -
# Setup parameters for models
# original data set too large to work with in reasonable time due to limted GPU resources
pct_data_sample = 0.1
# how much to hold out for validation
valid_pct = 0.2
# for repeatability - different seed than used with language model
seed = 1776
### FOR AWD_LSTM
### changing batch size affects learning rate
# batch size of 64 GPU uses about 16GB RAM (seems to work, but next run fails)
# batch size of 48 GPU uses 16GB RAM at peak
### FOR TRANSFORMER
# batch size of 48 requires more than 16GB RAM
# batch size of 32 requires more than 16GB RAM
# batch size of 4 requires about 15 GB RAM
### FOR TRANSFORMERXML
# batch size of 8 requires more than 16GB RAM
# batch size of 4 requires about 11 GB RAM
bs=4
# if this doesn't free memory, can restart Python kernel.
# if that still doesn't work, try OS items mentioned here: https://docs.fast.ai/dev/gpu.html
def release_mem():
gc.collect()
torch.cuda.empty_cache()
release_mem()
orig_df = pd.DataFrame()
if os.path.isfile(notes_pickle_file):
print('Loading noteevent pickle file')
orig_df = pd.read_pickle(notes_pickle_file)
print(orig_df.shape)
else:
print('Could not find noteevent pickle file; creating it')
# run this the first time to covert CSV to Pickle file
orig_df = pd.read_csv(notes_file, low_memory=False, memory_map=True)
orig_df.to_pickle(notes_pickle_file)
df = orig_df.sample(frac=pct_data_sample, random_state=seed)
df.head()
print('Unique Categories:', len(df.CATEGORY.unique()))
print('Unique Descriptions:', len(df.DESCRIPTION.unique()))
# #### This is a very CPU and RAM intensive process - no GPU involved
#
# Also, since there are a wide range of descriptions, not all descriptions present in the test set are in the validation set, so cannot learn all of them.
filename = base_path/class_file
if os.path.isfile(filename):
data_cl = load_data(base_path, class_file, bs=bs)
print('loaded existing data')
else:
# do I need a vocab here? test with and without...
data_cl = (TextList.from_df(df, base_path, cols='TEXT')
#df has several columns; actual text is in column TEXT
.split_by_rand_pct(valid_pct=valid_pct, seed=seed)
#We randomly split and keep 20% for validation, set see for repeatability
.label_from_df(cols='DESCRIPTION')
#building classifier to automatically determine DESCRIPTION
.databunch(bs=bs))
data_cl.save(filename)
print('created new data bunch')
learn = text_classifier_learner(data_cl, AWD_LSTM, drop_mult=0.5, pretrained=False, metrics=[accuracy, FBeta(average='weighted', beta=1)])
learn.lr_find()
release_mem()
# Change learning rate based on results from the above plot
learn.recorder.plot()
# ### AWD_LSTM training
# First unfrozen training with `learn.fit_one_cycle(1, 5e-2, moms=(0.8,0.7))` results in
#
# Total time: 22:36
#
# epoch train_loss valid_loss accuracy time
# 0 0.967378 0.638532 0.870705 22:36
#
# First frozen training with `pretrained=False` and bs of 64
#
# Total time: 42:07
#
# epoch train_loss valid_loss accuracy time
# 0 2.440479 2.399600 0.545564 42:07
#
# Unfrozen run with `learn.fit_one_cycle(1, 1e-1, moms=(0.8,0.7))` and `pretrained=False` and `bs=48`
#
# Total time: 56:26
#
# epoch train_loss valid_loss accuracy time
# 0 2.530828 2.415014 0.545564 56:26
if os.path.isfile(str(init_model_file) + '.pth'):
learn.load(init_model_file)
print('loaded initial learner')
else:
print('Training new initial learner')
learn.fit_one_cycle(1, 5e-2, moms=(0.8,0.7),
callbacks=[
callbacks.CSVLogger(learn, filename=training_history_file, append=True)
])
print('Saving new learner')
learn.save(init_model_file)
print('Finished generating new learner')
learn.unfreeze()
learn.fit_one_cycle(1, 1e-1, moms=(0.8,0.7),
callbacks=[
callbacks.CSVLogger(learn, filename=training_history_file, append=True)
])
release_mem()
# ### Try Transformer instead of AWD_LSTM
#
# This architecture requires a very small batch size (4) to fit in GPU memory, is very slow, and has poor accuracy. Also of note is the very low training loss and corresponding very high validation loss.
#
# Total time: 4:02:13
#
# epoch train_loss valid_loss accuracy time
# 0 2.913743 144113.546875 0.000024 4:02:13
#
# Perhaps I picked the wrong learning rate, or other hyper parameters?
learn = text_classifier_learner(data_cl, Transformer, drop_mult=0.5, pretrained=False)
learn.unfreeze()
learn.lr_find()
learn.recorder.plot()
release_mem()
learn.unfreeze()
learn.fit_one_cycle(1, 1e-1, moms=(0.8,0.7),
callbacks=[
callbacks.CSVLogger(learn, filename=transformer_training_history_file, append=True)
])
release_mem()
# ### Try TransformerXL
#
# Total time: 4:11:31
#
# epoch train_loss valid_loss accuracy time
# 0 2.446510 11688.685547 0.000289 4:11:31
#
learn = text_classifier_learner(data_cl, TransformerXL, drop_mult=0.5, pretrained=False)
learn.unfreeze()
learn.lr_find()
learn.recorder.plot()
release_mem()
learn.unfreeze()
learn.fit_one_cycle(1, 1e-1, moms=(0.8,0.7),
callbacks=[
callbacks.CSVLogger(learn, filename=transformerxl_training_history_file, append=True)
])
release_mem()
|
sourcecode/descr_classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Exercise set 5**
# ==============
#
#
# >The goal of this exercise is to perform and analyze experimental
# >designs. After this exercise you should have gained familiarity
# >with both full and fractional factorial designs and concepts
# >such as **effects**, **confounding**, **defining contrast**,
# >**generators** and **resolution**.
#
#
# **Exercise 5.1**
#
# The growth rate of a certain bacterium specie depends
# on the concentration of nutrients such as phosphate,
# sucrose, and nitrate. We have conducted a set of
# experiments where we have investigated how the
# growth is influenced by the concentration of
# phosphate ($P$), the concentration of sucrose ($S$) and
# the concentration of nitrate ($N$). The design
# matrix and the measured growth rate are given in table 1.
#
#
# |$P$ | $S$ | $N$ | **Growth rate** |
# |:---:|:---:|:---:|:---:|
# |$+$ | $-$ | $-$ | $7$ |
# |$-$ | $+$ | $-$ | $10$ |
# |$+$ | $-$ | $+$ | $8$ |
# |$-$ | $+$ | $+$ | $11$ |
# |$-$ | $-$ | $-$ | $11$ |
# |$+$ | $+$ | $+$ | $12$ |
# |$+$ | $+$ | $-$ | $7$ |
# |$-$ | $-$ | $+$ | $7$ |
#
# | |
# |:---|
# |**Table 1:** *Experimental design matrix for the growth rate of the investigated bacteria. The factors are the concentration of phosphate ($P$), the concentration of sucrose ($S$), and the concentration of nitrate ($N$).*|
#
#
# **(a)** Compute all the main effects.
#
#
# +
# Your code here
# -
# **Your answer to question 5.1(a):** *Double click here*
# **(b)** Extend the design matrix with the possible $2$-factor and $3$-factor
# interaction effects. Compute these interaction effects.
#
#
# +
# Your code here
# -
# **Your answer to question 5.1(b):** *Double click here*
# **(c)** What factors and interactions seem
# to increase the growth rate?
#
#
# +
# Your code here
# -
# **Your answer to question 5.1(c):** *Double click here*
# **(d)** Make two least-squares models of the data
# given in table 1:
#
# * (i) Model 1, which only includes the main effects.
#
# * (ii) Model 2, which includes the main effects and
# the interactions.
#
#
# When making the least-squares models, convert "$+$" to $1$ and
# "$-$" to $-1$.
#
# Compare the two models with the effects you have calculated, and the
# conclusions you made in point **(c)**.
#
# +
# Your code here
# -
# **Your answer to question 5.1(d):** *Double click here*
#
# **Exercise 5.2**
#
# You have recently started a new job as the lead
# experimental chemist in a company that makes chocolate bars.
# A new, and supposedly tasty, chocolate is being developed, and the
# main ingredients that you can vary are:
#
#
# * The amount of cocoa ($A$).
#
# * The number of pecan nuts ($B$).
#
# * The amount of caramel ($C$).
#
# * The amount of milk powder ($D$).
#
# * The amount of sugar ($E$).
#
# * The amount of vanilla ($F$).
#
# You are tasked with carrying out a maximum of $16$ experiments (limited due to cost
# and time constraints) in which the best mixture of the main ingredients ($A$–$F$)
# is found. ("Best" is here determined by a tasting panel made up of $30$ people.)
# For this task you decide on making a two-level fractional factorial design.
#
#
# **(a)** How many experiments would you have to carry out
# if you were to perform a full factorial design?
#
#
# +
# Your code here
# -
# **Your answer to question 5.2(a):** *Double click here*
# **(b)** As stated, you can only carry out $16$ experiments.
# Explain what confounding is and why the set up with
# $16$ experiments will lead to confounding.
#
#
# +
# Your code here
# -
# **Your answer to question 5.2(b):** *Double click here*
# **(c)** After talking with the chocolate design team, you
# decide on the following generators:
#
# * $E = ABC$.
#
# * $F = BCD$.
#
# What is a defining contrast, and what are the
# defining contrasts in this case?
#
#
# +
# Your code here
# -
# **Your answer to question 5.2(c):** *Double click here*
# **(d)** Are any of the main effects confounded with $2$-factor
# interactions in this case?
#
#
# +
# Your code here
# -
# **Your answer to question 5.2(d):** *Double click here*
# **(e)** What is the resolution for this design? Write out
# the short-hand representation of this design on the
# form $2^{N-p}_R$, where $N$ is the number of factors,
# $p$ is the number of generators, and $R$ the resolution.
#
#
# +
# Your code here
# -
# **Your answer to question 5.2(e):** *Double click here*
# **(f)** Construct the design matrix for the current design, but show
# only the columns for the main effects.
#
#
# +
# Your code here
# -
# **Your answer to question 5.2(f):** *Double click here*
# **(g)** Another member of your team suggests doing just $8$ experiments
# as this will cut time and cost.
# Do you think this is a good idea? Why/why not? What would
# the design matrix look like in this case?
#
# +
# Your code here
# -
# **Your answer to question 5.2(g):** *Double click here*
#
#
|
exercises/05_Exercise_Set_5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumTopics: RTLWiki
# +
import gc
import pickle
import time
from topicnet.cooking_machine.dataset import Dataset
from topicnet.cooking_machine.models import TopicModel
from topicnet.cooking_machine.model_constructor import init_simple_default_model
# -
import matplotlib.pyplot as plt
# %matplotlib inline
# +
import sys
sys.path.insert(0, '..')
from run_search import (
_optimize_scores,
)
from topnum.data.vowpal_wabbit_text_collection import VowpalWabbitTextCollection
from topnum.scores import (
PerplexityScore,
EntropyScore,
DiversityScore
)
from topnum.search_methods.optimize_scores_method import OptimizeScoresMethod
# +
# -
from topnum.model_constructor import init_bcg_sparse_model, init_decorrelated_PLSA, init_LDA, init_PLSA
# ## Data
# +
PATH = '/home/vbulatov/Projects/tmp_notebooks/MKB10.csv'
BATCHES = './MKB_batches'
dataset = Dataset(PATH, batch_vectorizer_path=BATCHES)
dataset.get_possible_modalities()
# -
modalities = {
"@text": 1.0,
}
# +
vw_file_path = "MKB_vw.txt"
dataset.write_vw(vw_file_path)
# +
# TODO: instead of modalities let's use predefined model families
# TODO: output_file_path
# -
# ## Experiments
main_modality_name = next(iter(modalities.keys()))
# +
main_modality_name, modalities = main_modality_name, modalities
modality_names = list(modalities.keys())
# vw_file_path = args.vw_file_path
output_file_path = "output.json"
min_num_topics = 5
max_num_topics = 30
num_topics_interval = 5
#num_topics_interval = 1
num_fit_iterations = 10
num_restarts = 1
# +
scores = [
EntropyScore('res', threshold_factor=1, class_ids=modality_names),
EntropyScore('res2', threshold_factor=2, class_ids=modality_names),
EntropyScore('res3', threshold_factor=3, class_ids=modality_names),
DiversityScore('ds_l2'),
DiversityScore('ds_cosine', metric="cosine"),
DiversityScore('ds_js', metric="jensenshannon"),
DiversityScore('ds_h', metric="hellinger"),
]
# +
text_collection = VowpalWabbitTextCollection(
vw_file_path,
main_modality=main_modality_name,
modalities=modalities
)
optimizer = OptimizeScoresMethod(
model_family="PLSA",
scores=scores,
min_num_topics=min_num_topics,
max_num_topics=max_num_topics,
num_topics_interval=num_topics_interval,
num_fit_iterations=num_fit_iterations,
num_restarts=num_restarts,
experiment_name="__TEST3__"
)
# -
t_start
# +
t_start = time.time()
optimizer.search_for_optimum(text_collection)
# -
# # ! ls 'num_topics_experiments/__TEST2___-1/##20h14m04s_02d03m2020y###/model'
# tm = TopicModel.load('num_topics_experiments/__TEST2___-1/##20h14m04s_02d03m2020y###')
# tm2 = TopicModel.load('num_topics_experiments/__TEST2___-1/##19h55m00s_02d03m2020y###')
#
# import pickle
#
# with open('num_topics_experiments/__TEST2___-1/##20h14m04s_02d03m2020y###/ds_js.p', "rb") as f:
# js = pickle.load(f)
#
# js.call(tm), js.call(tm2)
# tm.scores["ds_js"]
# tm2.scores["ds_js"]
# import sys
# del sys.modules['topnum.search_methods.optimize_scores_method']
# from topnum.search_methods.optimize_scores_method import _summarize_models, restore_failed_experiment
# result, detailed_result = restore_failed_experiment('num_topics_experiments', '__TEST2__')
# detailed_result['ds_h']
# +
t_end = time.time()
t_end - t_start
# -
detailed_result = optimizer._detailed_result
optimizer._detailed_result
optimizer._result['score_results'].keys()
optimizer._result['score_results']['ds_h']
# The lower the entropy, the better is supposed to be the result model.
# On X axis is the number of topics, on Y axis — the score.
# +
plt.plot(detailed_result['Topic<EMAIL>_<EMAIL>'].T)
plt.show()
# +
plt.plot(detailed_result['Topic<EMAIL>.average_purity'].T)
plt.show()
# +
plt.plot(detailed_result['TopicKernel<EMAIL>.average_size'].T)
plt.show()
# +
plt.plot(detailed_result['res'].T)
plt.show()
# +
plt.plot(detailed_result['ds_js'].T)
plt.show()
# -
plt.plot(detailed_result['res2'].T.mean(axis=1))
plt.show()
plt.plot(detailed_result['ds_h'].T)
plt.show()
plt.plot(detailed_result['ds_l2'].T.mean(axis=1))
plt.show()
with open("detailed_result_rtl.p", "wb") as f:
pickle.dump(detailed_result, f)
# All models are saved and can be restored
tm = TopicModel.load(
"./num_topics_experiments/e68cc1ff_experiment_-1/##13h32m50s_20d02m2020y###"
)
df.sum().sum() /
df.mean().mean() * 25/24
T = 25
2/(T * (T - 1)) * sum(condensed_distances)
phi.shape
|
demos/NumTopics_RTLWiki_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# +
train_path = '../data/tiantic/train.csv'
test_path = '../data/tiantic/test.csv'
data = pd.read_csv(train_path)
test_data = pd.read_csv(test_path)
# -
def plot_confusion_matrix(matrix, classes, title="Confusion Matrix", cmap=plt.cm.Blues):
plt.imshow(matrix, interpolation='nearest', cmap=cmap) #interpolation: 把某块显示成一种颜色
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
thresh = matrix.max() / 2.
#plt.xticks : 第一个数组参数是标示的位置,第二个数组是标示的文字
plt.xticks(tick_marks, classes)
plt.yticks(tick_marks, classes)
for y in range(len(matrix)):
for x in range(len(matrix[0])):
plt.text(x, y, matrix[y,x]
,horizontalalignment="center"
,color="white" if matrix[y,x] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# ## 数据预处理
def preprocess(data):
# 填充空数据
data['Age'] = data['Age'].fillna(data['Age'].median())
data['Embarked'] = data['Embarked'].fillna('S')
data['Fare'] = data['Fare'].fillna(data['Fare'].median())
# 将年龄转换字符型为数字型
data.loc[data['Sex'] == 'male', 'Sex'] = 0
data.loc[data['Sex'] == 'female', 'Sex'] = 0
data.loc[data['Embarked'] == 'S', 'Embarked'] = 0
data.loc[data['Embarked'] == 'Q', 'Embarked'] = 1
data.loc[data['Embarked'] == 'C', 'Embarked'] = 2
preprocess(data)
preprocess(test_data)
# ### 线性回归与逻辑回归模型预测
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.model_selection import KFold
from sklearn.metrics import accuracy_score, confusion_matrix
# +
lr_features = ['Pclass', 'Sex', 'Age', 'SibSp', 'Parch', 'Fare', 'Embarked']
x_data = data.loc[:, lr_features]
y_data = data['Survived']
# 划分训练集与测试集
lr_x_train, lr_x_test, lr_y_train, lr_y_test = train_test_split(lr_x_data, lr_y_data, test_size=0.2)
# 交叉验证
kf = KFold(n_splits=5, random_state=42)
lr_accuracies = []
lgr_accuracies = []
lr = LinearRegression()
lgr = LogisticRegression(C=10, penalty='l2', solver='liblinear')
for train, test in kf.split(lr_x_train):
lr.fit(lr_x_train.iloc[train, :], lr_y_train.iloc[train])
lgr.fit(lr_x_train.iloc[train, :], lr_y_train.iloc[train])
lr_predict = lr.predict(lr_x_train.iloc[test, :])
lgr_predict = lgr.predict(lr_x_train.iloc[test, :])
lr_predictions = np.zeros(len(lr_predict))
lr_predictions[lr_predict > 0.5] = 1
lr_accuracy = accuracy_score(lr_y_train.iloc[test].values, lr_predictions)
lgr_accuracy = accuracy_score(lr_y_train.iloc[test].values, lgr_predict)
lr_accuracies.append(lr_accuracy)
lgr_accuracies.append(lgr_accuracy)
print('线性回归算法精确度:%.3f' % np.mean(lr_accuracies))
print('逻辑回归算法精确度:%.3f' % np.mean(lgr_accuracies))
# +
lr.fit(lr_x_train, lr_y_train)
lgr.fit(lr_x_train, lr_y_train)
lr_predictions_proba = lr.predict(lr_x_test)
lr_predictions = np.zeros(len(lr_predictions_proba))
lr_predictions[lr_predictions_proba > 0.5] = 1
lgr_predictions = lgr.predict(lr_x_test)
lr_accuracy = accuracy_score(lr_y_test, lr_predictions)
lgr_accuracy = accuracy_score(lr_y_test, lgr_predictions)
print('线性回归算法精确度:%.4f' % np.mean(lr_accuracy))
print('逻辑回归算法精确度:%.4f' % np.mean(lgr_accuracy))
# -
# ### 随机森林
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
# **min_samples_leaf**:这个值限制了叶子节点最少的样本数,如果某叶子节点数目小于样本数,则会和兄弟节点一起被剪枝。 默认是1,可以输入最少的样本数的整数,或者最少样本数占样本总数的百分比。如果样本量不大,不需要管这个值。如果样本量数量级非常大,则推荐增大这个值。之前的10万样本项目使用min_samples_leaf的值为5,仅供参考。
#
# **min_samples_split**:这个值限制了子树继续划分的条件,如果某节点的样本数少于min_samples_split,则不会继续再尝试选择最优特征来进行划分。 默认是2.如果样本量不大,不需要管这个值。如果样本量数量级非常大,则推荐增大这个值。我之前的一个项目例子,有大概10万样本,建立决策树时,我选择了min_samples_split=10。可以作为参考。
# +
rfc = RandomForestClassifier(random_state=1, n_estimators=55, min_samples_leaf=2, min_samples_split=2)
kf = KFold(5)
course = cross_val_score(rfc, x_data, y_data, cv=kf)
print(np.mean(course))
rfc.fit(x_data, y_data)
predict = rfc.predict(x_data)
score = accuracy_score(y_data.values.ravel(), predict)
cm = confusion_matrix(y_data.values.ravel(), predict)
plot_confusion_matrix(cm, classes=[0,1])
print("精度:", score)
# -
# ### 自动特征选择
from sklearn.feature_selection import SelectKBest, f_classif
selector = SelectKBest(f_classif, k='all')
selector.fit(x_data, y_data)
selector.scores_
len(lr_features)
lr_features
|
practice/Tiantic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3
# ---
# <center>
# <img src="https://gitlab.com/ibm/skills-network/courses/placeholder101/-/raw/master/labs/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
# </center>
#
# # **Space X Falcon 9 First Stage Landing Prediction**
#
# ## Lab 2: Data wrangling
#
# Estimated time needed: **60** minutes
#
# In this lab, we will perform some Exploratory Data Analysis (EDA) to find some patterns in the data and determine what would be the label for training supervised models.
#
# In the data set, there are several different cases where the booster did not land successfully. Sometimes a landing was attempted but failed due to an accident; for example, <code>True Ocean</code> means the mission outcome was successfully landed to a specific region of the ocean while <code>False Ocean</code> means the mission outcome was unsuccessfully landed to a specific region of the ocean. <code>True RTLS</code> means the mission outcome was successfully landed to a ground pad <code>False RTLS</code> means the mission outcome was unsuccessfully landed to a ground pad.<code>True ASDS</code> means the mission outcome was successfully landed on a drone ship <code>False ASDS</code> means the mission outcome was unsuccessfully landed on a drone ship.
#
# In this lab we will mainly convert those outcomes into Training Labels with `1` means the booster successfully landed `0` means it was unsuccessful.
#
# Falcon 9 first stage will land successfully
#
# 
#
# Several examples of an unsuccessful landing are shown here:
#
# 
#
#
# ## Objectives
#
# Perform exploratory Data Analysis and determine Training Labels
#
# * Exploratory Data Analysis
# * Determine Training Labels
#
# ***
#
# ## Import Libraries and Define Auxiliary Functions
#
# We will import the following libraries.
#
# Pandas is a software library written for the Python programming language for data manipulation and analysis.
import pandas as pd
#NumPy is a library for the Python programming language, adding support for large, multi-dimensional arrays and matrices, along with a large collection of high-level mathematical functions to operate on these arrays
import numpy as np
# ### Data Analysis
#
# Load Space X dataset, from last section.
#
df=pd.read_csv("https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/datasets/dataset_part_1.csv")
df.head(10)
# Identify and calculate the percentage of the missing values in each attribute
#
df.isnull().sum()/df.count()*100
# Identify which columns are numerical and categorical:
#
df.dtypes
# ### TASK 1: Calculate the number of launches on each site
#
# The data contains several Space X launch facilities: <a href='https://en.wikipedia.org/wiki/List_of_Cape_Canaveral_and_Merritt_Island_launch_sites?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01'>Cape Canaveral Space</a> Launch Complex 40 <b>VAFB SLC 4E </b> , Vandenberg Air Force Base Space Launch Complex 4E <b>(SLC-4E)</b>, Kennedy Space Center Launch Complex 39A <b>KSC LC 39A </b>.The location of each Launch Is placed in the column <code>LaunchSite</code>
#
# Next, let's see the number of launches for each site.
#
# Use the method <code>value_counts()</code> on the column <code>LaunchSite</code> to determine the number of launches on each site:
#
# Apply value_counts() on column LaunchSite
df.loc[:, 'LaunchSite'].value_counts()
# Each launch aims to an dedicated orbit, and here are some common orbit types:
#
# * <b>LEO</b>: Low Earth orbit (LEO)is an Earth-centred orbit with an altitude of 2,000 km (1,200 mi) or less (approximately one-third of the radius of Earth),\[1] or with at least 11.25 periods per day (an orbital period of 128 minutes or less) and an eccentricity less than 0.25.\[2] Most of the manmade objects in outer space are in LEO <a href='https://en.wikipedia.org/wiki/Low_Earth_orbit?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01'>\[1]</a>.
#
# * <b>VLEO</b>: Very Low Earth Orbits (VLEO) can be defined as the orbits with a mean altitude below 450 km. Operating in these orbits can provide a number of benefits to Earth observation spacecraft as the spacecraft operates closer to the observation<a href='https://www.researchgate.net/publication/271499606_Very_Low_Earth_Orbit_mission_concepts_for_Earth_Observation_Benefits_and_challenges?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01'>\[2]</a>.
#
# * <b>GTO</b> A geosynchronous orbit is a high Earth orbit that allows satellites to match Earth's rotation. Located at 22,236 miles (35,786 kilometers) above Earth's equator, this position is a valuable spot for monitoring weather, communications and surveillance. Because the satellite orbits at the same speed that the Earth is turning, the satellite seems to stay in place over a single longitude, though it may drift north to south,” NASA wrote on its Earth Observatory website <a href="https://www.space.com/29222-geosynchronous-orbit.html?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01" >\[3] </a>.
#
# * <b>SSO (or SO)</b>: It is a Sun-synchronous orbit also called a heliosynchronous orbit is a nearly polar orbit around a planet, in which the satellite passes over any given point of the planet's surface at the same local mean solar time <a href="https://en.wikipedia.org/wiki/Sun-synchronous_orbit?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">\[4] <a>.
#
# * <b>ES-L1 </b>:At the Lagrange points the gravitational forces of the two large bodies cancel out in such a way that a small object placed in orbit there is in equilibrium relative to the center of mass of the large bodies. L1 is one such point between the sun and the earth <a href="https://en.wikipedia.org/wiki/Lagrange_point?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01#L1_point">\[5]</a> .
#
# * <b>HEO</b> A highly elliptical orbit, is an elliptic orbit with high eccentricity, usually referring to one around Earth <a href="https://en.wikipedia.org/wiki/Highly_elliptical_orbit?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">\[6]</a>.
#
# * <b> ISS </b> A modular space station (habitable artificial satellite) in low Earth orbit. It is a multinational collaborative project between five participating space agencies: NASA (United States), Roscosmos (Russia), JAXA (Japan), ESA (Europe), and CSA (Canada)<a href="https://en.wikipedia.org/wiki/International_Space_Station?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01"> \[7] </a>
#
# * <b> MEO </b> Geocentric orbits ranging in altitude from 2,000 km (1,200 mi) to just below geosynchronous orbit at 35,786 kilometers (22,236 mi). Also known as an intermediate circular orbit. These are "most commonly at 20,200 kilometers (12,600 mi), or 20,650 kilometers (12,830 mi), with an orbital period of 12 hours <a href="https://en.wikipedia.org/wiki/List_of_orbits?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01"> \[8] </a>
#
# * <b> HEO </b> Geocentric orbits above the altitude of geosynchronous orbit (35,786 km or 22,236 mi) <a href="https://en.wikipedia.org/wiki/List_of_orbits?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01"> \[9] </a>
#
# * <b> GEO </b> It is a circular geosynchronous orbit 35,786 kilometres (22,236 miles) above Earth's equator and following the direction of Earth's rotation <a href="https://en.wikipedia.org/wiki/Geostationary_orbit?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01"> \[10] </a>
#
# * <b> PO </b> It is one type of satellites in which a satellite passes above or nearly above both poles of the body being orbited (usually a planet such as the Earth <a href="https://en.wikipedia.org/wiki/Polar_orbit?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01"> \[11] </a>
#
# some are shown in the following plot:
#
# 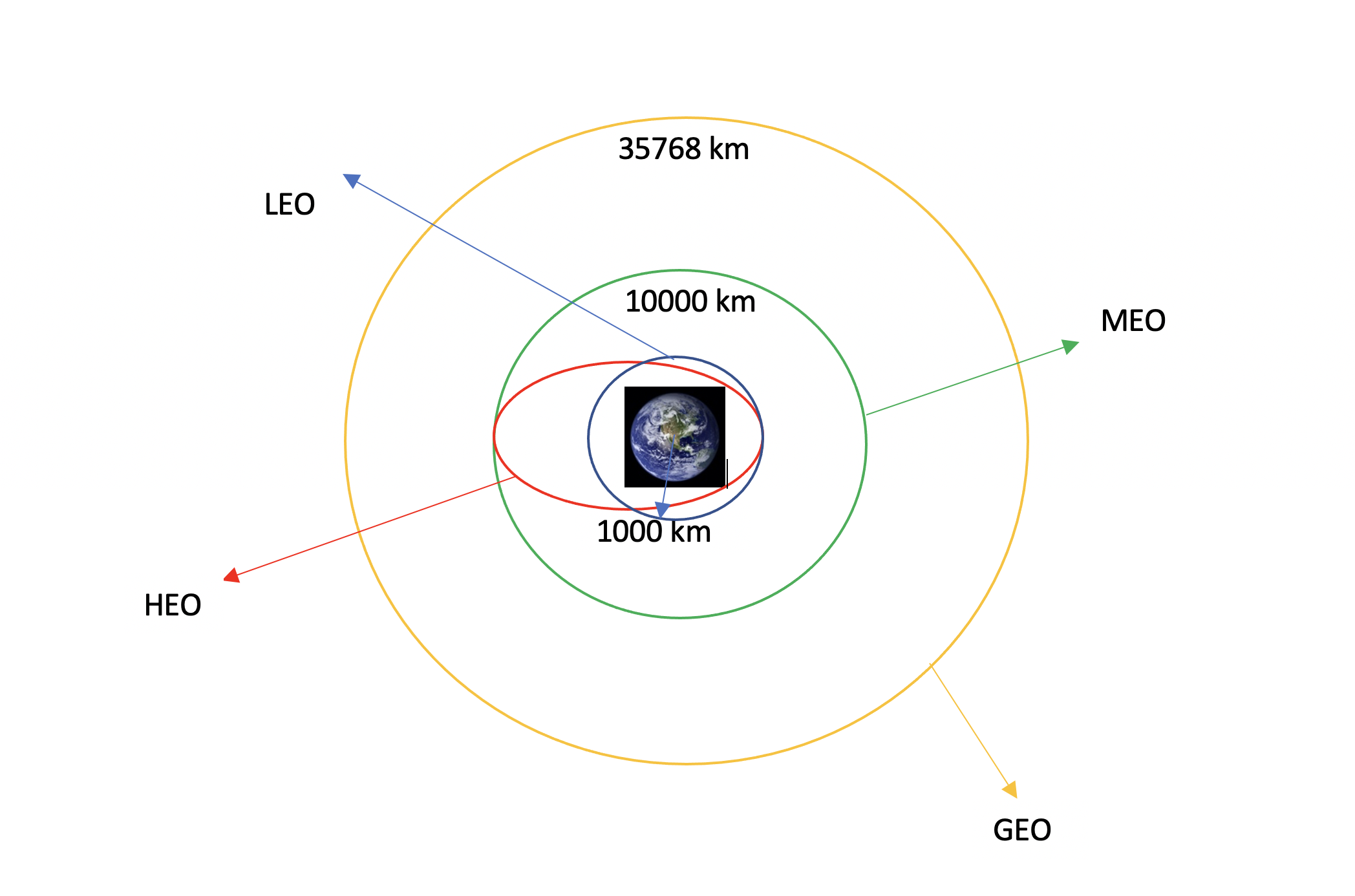
#
# ### TASK 2: Calculate the number and occurrence of each orbit
#
# Use the method <code>.value_counts()</code> to determine the number and occurrence of each orbit in the column <code>Orbit</code>
#
# Apply value_counts on Orbit column
df.loc[:, 'Orbit'].value_counts()
# ### TASK 3: Calculate the number and occurence of mission outcome per orbit type
#
# Use the method <code>.value_counts()</code> on the column <code>Outcome</code> to determine the number of <code>landing_outcomes</code>.Then assign it to a variable landing_outcomes.
#
# landing_outcomes = values on Outcome column
landing_outcomes = df.loc[:, 'Outcome'].value_counts()
landing_outcomes
# <code>True Ocean</code> means the mission outcome was successfully landed to a specific region of the ocean while <code>False Ocean</code> means the mission outcome was unsuccessfully landed to a specific region of the ocean. <code>True RTLS</code> means the mission outcome was successfully landed to a ground pad <code>False RTLS</code> means the mission outcome was unsuccessfully landed to a ground pad.<code>True ASDS</code> means the mission outcome was successfully landed to a drone ship <code>False ASDS</code> means the mission outcome was unsuccessfully landed to a drone ship. <code>None ASDS</code> and <code>None None</code> these represent a failure to land.
#
for i,outcome in enumerate(landing_outcomes.keys()):
print(i,outcome)
# We create a set of outcomes where the second stage did not land successfully:
#
bad_outcomes=set(landing_outcomes.keys()[[1,3,5,6,7]])
bad_outcomes
# ### TASK 4: Create a landing outcome label from Outcome column
#
# Using the <code>Outcome</code>, create a list where the element is zero if the corresponding row in <code>Outcome</code> is in the set <code>bad_outcome</code>; otherwise, it's one. Then assign it to the variable <code>landing_class</code>:
#
# landing_class = 0 if bad_outcome
# landing_class = 1 otherwise
landing_class = df.loc[:, 'Outcome'].map(lambda x: int(not x in bad_outcomes))
landing_class
# This variable will represent the classification variable that represents the outcome of each launch. If the value is zero, the first stage did not land successfully; one means the first stage landed Successfully
#
df['Class']=landing_class
df[['Class']].head(8)
df.head(5)
# We can use the following line of code to determine the success rate:
#
df["Class"].mean()
# We can now export it to a CSV for the next section,but to make the answers consistent, in the next lab we will provide data in a pre-selected date range.
#
# <code>df.to_csv("dataset_part\_2.csv", index=False)</code>
#
# ## Authors
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01"><NAME></a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
#
# <a href="https://www.linkedin.com/in/nayefaboutayoun/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01"><NAME></a> is a Data Scientist at IBM and pursuing a Master of Management in Artificial intelligence degree at Queen's University.
#
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ------------- | ----------------------- |
# | 2021-08-31 | 1.1 | <NAME> | Changed Markdown |
# | 2020-09-20 | 1.0 | Joseph | Modified Multiple Areas |
# | 2020-11-04 | 1.1. | Nayef | updating the input data |
# | 2021-05-026 | 1.1. | Joseph | updating the input data |
#
# Copyright © 2021 IBM Corporation. All rights reserved.
#
|
labs-jupyter-spacex-Data wrangling.ipynb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <font style="font-size:96px; font-weight:bolder; color:#0040a0"><img src="http://montage.ipac.caltech.edu/docs/M51_logo.png" alt="M" style="float: left; padding: 25px 30px 25px 0px;" /></font>
#
# <i><b>Montage</b> Montage is an astronomical image toolkit with components for reprojection, background matching, coaddition and visualization of FITS files. It can be used as a set of command-line tools (Linux, OS X and Windows), C library calls (Linux and OS X) and as Python binary extension modules.
#
# The Montage source is written in ANSI-C and code can be downloaded from GitHub ( https://github.com/Caltech-IPAC/Montage ). The Python package can be installed from PyPI ("</i>pip install MontagePy<i>"). The package has no external dependencies. See http://montage.ipac.caltech.edu/ for details on the design and applications of Montage.
#
# # MontagePy.main modules: mMakeHdr
#
# Much of the Montage processing is based on the specification of an output image as captured in an ASCII file version of the output FITS header. Sometimes such a header is cloned from an existing FITS file and sometimes created from scratch using an editor or the utility mHdr.
#
# Another common approach is the create a header is to base it on a table of source locations or image metadata by drawing a bounding box around the table contents. mMakeHdr reads through a table and determines such a bounding box using a spherical geometry variant of the rotating calipers technique from computational geometry. The box can either be of overall minimum size or can include the constraint that North be oriented upward in the image.
# +
from MontagePy.main import mMakeHdr, mViewer
help(mMakeHdr)
# -
# ## mMakeHdr Example
#
# The principle parameters for mMakeHdr are the table file to be fit (either just coordinates or image metadata) and the output header file name. There are optional settings for specifying the coordinate system to use, the pixel scale (the default is derived from the table), and whether the image should be "North-up".
#
# We have a set of images retrieved from the 2MASS archive that overlap with a 1-degree box around M17. Using the metadata from those image and mMakeHdr, we will derive a header file which is somewhat larger, since it will bound the entirity of those images:
#
# +
rtn = mMakeHdr('M17/remote.tbl', 'work/M17/bounding.hdr')
print(rtn)
# -
# ### Header
#
# Here is the generated header:
with open('work/M17/bounding.hdr', 'r') as fin:
print(fin.read(), end='')
# ## Error Handling
#
# If mMakeHdr encounters an error, the return structure will just have two elements: a status of 1 ("error") and a message string that tries to diagnose the reason for the error.
#
# For instance, if the user asks for a non-existent input table:
# +
rtn = mMakeHdr('M17/unknown.tbl', "work/M17/bounding.hdr")
print(rtn)
# -
#
#
# ## Classic Montage: mMakeHdr as a Stand-Alone Program
#
# ### mMakeHdr Unix/Windows Command-line Arguments
#
# <p>mMakeHdr can also be run as a command-line tool in Linux, OS X, and Windows:</p>
#
# <p><tt>
# <b>Usage:</b> mMakeHdr [-d level] [-s statusfile] [-p(ixel-scale) cdelt | -P maxpixel] [-e edgepixels] [-n] images.tbl template.hdr [system [equinox]] (where system = EQUJ|EQUB|ECLJ|ECLB|GAL|SGAL)
# </tt></p>
# <p> </p>
# <p>If you are writing in C/C++, mMakeHdr can be accessed as a library function:</p>
#
# <pre>
# /*-***********************************************************************/
# /* */
# /* mMakeHdr */
# /* */
# /* Create the best header 'bounding' a table or set of tables (each */
# /* with image metadata or point sources). */
# /* */
# /* char *tblfile Input image metadata table or source table */
# /* or table of tables */
# /* */
# /* char *template Output image header template */
# /* */
# /* char *csys Coordinate system (e.g. 'EquJ', 'Galactic'). */
# /* Fairly forgiving */
# /* */
# /* double equinox Coordinate system equinox (e.g. 2000.0) */
# /* */
# /* double pixelScale Pixel scale in degrees */
# /* */
# /* int northAligned Defaults to minimum bounding box around */
# /* input images. This forces template to be */
# /* north-aligned */
# /* */
# /* double pad Optional extra padding around output template */
# /* */
# /* int isPercentage Pad is in pixels by default. This changes */
# /* that to a percentage of the image size */
# /* */
# /* int maxPixel Setting the pixel scale can result in really */
# /* big images. This forces a maximum number */
# /* of pixels in NAXIS1, NAXIS2 */
# /* */
# /* int debug Debugging output level */
# /* */
# /*************************************************************************/
#
# struct mMakeHdrReturn *mMakeHdr(char *tblfile, char *template, char *csysin, double equinox, double pixelScale,
# int northAligned, double pad, int isPercentage, int maxPixel, int debugin)
# </pre>
# <p><b>Return Structure</b></p>
# <pre>
# struct mMakeHdrReturn
# {
# int status; // Return status (0: OK, 1:ERROR)
# char msg [1024]; // Return message (for error return)
# char json[4096]; // Return parameters as JSON string
# char note[1024]; // Cautionary message (only there if needed).
# int count; // Number of images in metadata table.
# int ncube; // Number of images that have 3/4 dimensions.
# int naxis1; // X axis pixel count in output template.
# int naxis2; // Y axis pixel count in output template.
# double clon; // Center longitude for template.
# double clat; // Center latitude for template.
# double lonsize; // Template dimensions in X.
# double latsize; // Template dimensions in Y.
# double posang; // Rotation angle of template.
# double lon1; // Image corners (lon of first corner).
# double lat1; // Image corners (lat of first corner).
# double lon2; // Image corners (lon of second corner).
# double lat2; // Image corners (lat of second corner).
# double lon3; // Image corners (lon of third corner).
# double lat3; // Image corners (lat of third corner).
# double lon4; // Image corners (lon of fourth corner).
# double lat4; // Image corners (lat of fourth corner).
# };
# </pre>
|
mMakeHdr.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# #### 통계
# - 표본 분포
# - 통계적 추론
# - 전수조사 불가능
# - 표본 조사를 통해 모집단 해석
# - 표본 조사는 오차 발생
# - 적절한 표본 추출 방법
# - 표본과 모집단의 관계 이해
# - 무작위 난수
#
# - 표본 평균의 분포
# - 표본 평균은 표본의 선택에 따라 달라짐
# - 표본 평균은 확률 변수
#
# - 표본 평균 : 모평균을 알아내는 통계량
#
# - 중심 극한 정리
# - n이 충분히 큰 경우 n >= 30
# - 근사적으로 정규분포를 따름
#
# - 추정
# - 모 평균의 추정
# - 점 추정
# - 구간 추정
#
# - 표본 평균 추정
#
# - 구간 추정
# - 표본의 크기가 클때 중심 극한 정리 사용
#
#
# - 고1 남학생 평균키 추정
# - 36명 표본 평균 173.6
# - 표본 표준 편차 3.6
# - 평균키에 대한 95% 신뢰구간
#
# 36명이기때문에 30명 이상임 중심극한정리 사용
#
# alpha = 0.05
# z0.025 = 1.96
# 중심 극한 정리 공식에 대입
#
#
# 계란 30개의 표본이 무게
#
# w = ~ 30개 있을때
#
# 무게에 대한 95% 신뢰구간
#
# 1) 표본평균 구함 np.mean(w)
# 2) 표준편차 구함 np.std(w, ddof = 1)
# 3) alpha = 0.05
# 4) zalpha = scipy.stats.norm.ppf(1-alpha*0.5)
# ppf 누적함수
#
# - 모비율의 추정
# - 점 추정
# - 특정 속성을 갖는 표본의 개수
#
# - 구간추정
# - n이 충분히 클때
# - np > 5
# - n( 1-p ) > 5
# - X~N(np, np(1-p)) 정규분포
#
# - 확률 변수 X의 표준화
#
# 신뢰 구간의 의미를 잘 알아두고 공식들은 그때 찾아서 하는 것
#
# - 검정
# - 통계적 가설검정
# - 가설 검정의 원리
#
# 예) 고1 평균 170.5
# 1학년 랜덤 30명 평균 171.3
#
# 올해 신입생 평균키가 170.5 보다
# 크다고 할 수 있는가?
#
#
# 귀무가설
# - 명제
# 대립가설
# - 명제와 반대되는 혹은 다른 명제
#
#
# - 모평균의 검정
# - 검증 자체 전부 동일함
#
#
#
#
#
#
# #### 엔트로피
# - 자기정보
# - i(A)
# - A : 사건
# - logb(1/P(A)) == -logbP(A)
# - 확률이 높은 사건:
# - 확률이 높은 사건이 일어나면 정보가 많지 않음
# - 확률이 낮은 사건이 일어나야 정보가 많음
# - 낮은 사건에 정보가 더 많다는 말
# - i(AB) = i(A) + i(B)
#
# - P(H) = 1/8, P(T) = 7/8
# - i(H) = 3 , i(T) = 0.19
#
# - [엔트로피](https://ko.wikipedia.org/wiki/%EC%97%94%ED%8A%B8%EB%A1%9C%ED%94%BC#%ED%86%B5%EA%B3%84%EC%97%AD%ED%95%99%EC%A0%81_%EC%A0%95%EC%9D%98)
# - 자기 정보의 평균
# - 교차 엔트로피
# - H(P,Q)
# - P,Q가 얼마나 비슷한지 표현하는 수치
# - 분류 문제에서 손실함수
# - 주어진 대상이 A인지 아닌지 판단
# - 주어진 대상이 A,B,C 어느것 인지 판단
#
# 교체 엔트로피가 인공지능에서 손실함수로 자주사용 함
|
_notebooks/TIL9.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/zacksnyder-lsds/Unit_1_build_week/blob/master/Unit_1_build_week.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="cl7TmhMP2nDY" colab_type="text"
# #Reading in the data and initial cleaning
# + id="f3qUVbdPyOuC" colab_type="code" colab={}
#Imports for the data reading and initial cleaning section
import pandas as pd
# + id="GdWH6EiLk6_f" colab_type="code" colab={}
#reading in the datasets I split in a seperate notebook so I could upload to github
url1 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva1%20(1).csv'
url2 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva2.csv'
url3 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva3.csv'
url4 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva4.csv'
url5 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva5.csv'
url6 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva6.csv'
url7 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva7.csv'
url8 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva8.csv'
url9 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva9.csv'
url10 = 'https://raw.githubusercontent.com/zacksnyder-lsds/Unit_1_build_week/master/kiva10.csv'
# + id="beS-HYNOxzZK" colab_type="code" colab={}
#converting the csv files back to dataframes for use
kiva1 = pd.read_csv(url1)
kiva2 = pd.read_csv(url2)
kiva3 = pd.read_csv(url3)
kiva4 = pd.read_csv(url4)
kiva5 = pd.read_csv(url5)
kiva6 = pd.read_csv(url6)
kiva7 = pd.read_csv(url7)
kiva8 = pd.read_csv(url8)
kiva9 = pd.read_csv(url9)
kiva10 = pd.read_csv(url10)
# + id="sOP39Z0jyNb5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 753} outputId="5dafbc22-77f2-41cc-e3f1-4611e6b8bd39"
#checking to make sure all the data read in smoothly
[print(name.shape) for name in [kiva1,kiva2,kiva3,kiva4,kiva5,kiva6,kiva7,kiva8,kiva9,kiva10]]
kiva2.head()
# + id="zrqkF-9By11d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 702} outputId="d7da566d-8ff0-41af-8e2c-d9fa8532ff79"
#concatinating the dataframes back together to the original from Kaggle
kiva = pd.concat([kiva1,kiva2,kiva3,kiva4,kiva5,kiva6,kiva7,kiva8,kiva9,kiva10])
kiva.reset_index()
print(kiva.shape)
kiva.sample(5)
# + id="p3Mzohv2zcpI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 585} outputId="d98b33b4-ed05-47b3-a8c6-d1773e018259"
#getting rid of the previous index column renamed unanamed 0
kiva.drop('Unnamed: 0', axis=1, inplace=True)
kiva.sample(5)
# + [markdown] id="hSWaAqj53xOS" colab_type="text"
# #Deeper Cleaning
# + [markdown] id="Moczewml339H" colab_type="text"
# ###grabbing just the top 25 countries
# + id="rtctJMxe13vv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="a81ba21f-0c69-42c1-df44-8d4c4ec53e00"
#finding the top 25 countries that Kiva serves
top_code = kiva['country_code'].value_counts()[:26].index
top_code
# + id="MP3gTKv3_UIM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="47c15992-44ae-4c02-c52c-827acb7cfbc4"
#finding out what percentage of the loans by Kiva fall in the top 25 countries
a = kiva['country_code'].value_counts()[:26].sum()
b = kiva['country_code'].value_counts().sum()
percent_described = a/b
print(percent_described*100, '% of the loans occur in the top 25 countries')
# + id="fpe-dorN2WdE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="37a23485-257e-48de-ff8a-d0e0bef68a23"
#creating a column for top 25 countries
kiva['top_25_countries'] = 'Other'
kiva['top_25_countries'].sample(5)
# + id="4IFXnOyQ5IXe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 343} outputId="8fc5513f-8524-412c-cd7f-6a2ca63203fa"
#locking just the top 25 countries
for code in top_code:
kiva.loc[kiva['country_code'] == code, 'top_25_countries'] = code
kiva[['top_25_countries','country_code']].sample(10)
# + id="vn82_bSN96ni" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="ee2e3aa2-c494-4af2-af42-9e578dac0619"
#getting rid of the unneeded column to keep things tidy
kiva.drop('country_code', axis=1, inplace=True)
kiva.sample(3)
# + [markdown] id="9CcpzzfP9xws" colab_type="text"
# ###grabbing just single borrowers
# + id="emTtgv0Z6M2j" colab_type="code" colab={}
#creating a column for single borrowers
kiva['single_gender'] = 'More than 1 borrower'
# + id="xSek3RYs6M0T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 343} outputId="fb004ac3-ee9c-4500-8a29-bfed9af655f7"
#locking the single borrowers to thier gender
kiva.loc[kiva['borrower_genders'] == 'male', 'single_gender'] = 'male'
kiva.loc[kiva['borrower_genders'] == 'female', 'single_gender'] = 'female'
kiva[['single_gender', 'borrower_genders']].sample(10)
# + id="-16i_v5N6Mw1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 568} outputId="759dc0d9-85d2-4972-8078-d9a719b0764a"
#getting rid of the unneeded column
kiva.drop('borrower_genders', axis=1, inplace=True)
kiva.head()
# + [markdown] id="MurlOAeXKC9q" colab_type="text"
# ###Trimming the fat
# + id="iwUq4v-Q6Mtc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="d3b1d4b9-88ae-4771-cd5c-42cb5cd47d22"
#determining what columns will be useful for my analysis
list(kiva.columns)
# + [markdown] id="xc4M3tJgK83A" colab_type="text"
# I want to isolate loan_amount, sector, use, activity, top 25 countries and single gender
# + id="qIdGzYuL6Mpu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="3b4bb62d-f529-43d7-87fb-95e057b049ce"
#isolating the columns
clean_columns = ['loan_amount', 'sector', 'use', 'activity', 'top_25_countries', 'single_gender','country']
kiva_clean = pd.concat([pd.DataFrame(kiva[column]) for column in clean_columns], axis=1)
kiva_clean.sample(5)
# + id="cSrEQeQOSBQZ" colab_type="code" colab={}
#defining a fucntion to drop the unwanted parts of the column
def p90x(data, column, unwanted):
data.set_index(data[column], inplace=True)
data.drop(unwanted, inplace=True)
data.drop(column, axis=1,inplace=True)
data.reset_index(inplace=True)
# + id="LWlXisem_BEM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="921d6b9c-1d97-4b21-d459-588a69566d70"
#running the function on top_25_countries and single_gender
p90x(kiva_clean, 'top_25_countries', 'Other')
p90x(kiva_clean, 'single_gender', 'More than 1 borrower')
kiva_clean.sample(10)
# + id="oYT95tFoOkdE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="e6831ca7-898f-4c7b-a24c-ada1471ee9d9"
kiva_clean['sector'].nunique()
# + [markdown] id="iKiyAGEe0EkJ" colab_type="text"
# #Work on Graph One
# + [markdown] id="mxl8NTc0d_B6" colab_type="text"
# ###doing some cleaning
# + id="sQ1Wd_NceKkv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="64cd5b40-c4b9-4a80-d7a4-8b78b0ec38a1"
#looking at the unique countries in my cleaned data so I can match them with thier continent
kiva_clean['country'].unique()
# + id="afySo7mWf8MC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="a570b37f-61a7-426c-ad71-bbcd5b2c2f0d"
#creating a new continent column
kiva_clean['cont'] = 'error'
kiva_clean.head()
# + id="rWxDVkMHe7dg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="323e3709-9270-407e-ef1a-87bacc493b08"
#creating a variable that matches countries to their continent
country_cont = [('Pakistan', 'Asia'), ('India', 'Asia'), ('Kenya', 'Africa'),
('Nicaragua', 'Americas'), ('El Salvador', 'Americas'),
('Philippines', 'Asia'), ('Peru', 'Americas'), ('Cambodia', 'Asia'),
('Honduras', 'Americas'), ('Palestine', 'Asia'), ('United States', 'Americas'),
('Colombia', 'Americas'), ('Tajikistan', 'Asia'), ('Ecuador', 'Americas'),
('Bolivia', 'Americas'), ('Uganda', 'Africa'), ('Indonesia', 'Asia'),
('Guatemala', 'Americas'), ('Mali', 'Africa'), ('Vietnam', 'Asia'),
('Armenia', 'Asia'), ('Paraguay', 'Americas'), ('Lebanon', 'Asia'),
('Samoa', 'Australia'), ('Rwanda', 'Africa'), ('Nigeria', 'Africa')]
for country, cont in country_cont:
kiva_clean.loc[kiva_clean['country'] == country, 'cont'] = cont
kiva_clean[['country', 'cont']].sample(10)
# + id="gOkwUS9B-0Y0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="b01cd33e-d714-43b6-b601-5bd854f71b3f"
#checking that I didn't miss anything
kiva_clean['cont'].unique()
# + [markdown] id="P06Z3PIJeGlf" colab_type="text"
# ###Plotly sunburst graph
# + id="I-3MCcnVbz5o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="3eaa088d-c809-4cd8-f984-d2f5bf3ed9d5"
# !pip install plotly==4.5.2
# + id="RLM0bpFN_BBc" colab_type="code" colab={}
#defining a function to run plotly in colab cells
def enable_plotly_in_cell():
import IPython
from plotly.offline import init_notebook_mode
display(IPython.core.display.HTML('''<script src="/static/components/requirejs/require.js"></script>'''))
init_notebook_mode(connected=False)
# + id="X6KxS4Z0Qmrq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="5d4d50aa-9918-43ba-9565-1ab5da30cb59"
loan_amount = kiva_clean.groupby(['cont', 'country', 'sector', 'single_gender'])['loan_amount'].sum()
loan_amount
# + id="wyIBpr2U_A-O" colab_type="code" colab={}
enable_plotly_in_cell()
#making the basics of the sunburst graph
import plotly.express as px
import numpy as np
fig = px.sunburst(kiva_clean, path=['cont', 'country', 'sector', 'single_gender'], values = 'loan_amount', color='cont',
color_discrete_map={'Asia': '#FF4F58FF', 'Africa': '#A89C94FF', 'Americas': '#669DB3FF', 'Australia' : '#F0F6F7FF'},
maxdepth=2,
)
fig.update_layout(extendsunburstcolors=True,
title = 'Who Gets the Biggest Bite of the Kiva Microlending Pie?',
xaxis_title = 'Please click to explore further!'
)
fig.show()
# + [markdown] id="bmG5N12ZNwRp" colab_type="text"
# ###pushing my plot to plotly (will be omited from github repository to protect my API key)
# + id="DyDoP04B_A6r" colab_type="code" colab={}
# !pip install chart_studio
import chart_studio
# + id="WwgFqUux_A3a" colab_type="code" colab={}
#username = 'zacksnyder1998'
#api_key = add_api_key_here
#chart_studio.tools.set_credentials_file(username=username, api_key=api_key)
# + id="TPnyCaB8_A0B" colab_type="code" colab={}
#import chart_studio.plotly as py
#py.plot(fig, filename = 'Who Gets the Biggest Bite of the Kiva Microlending Pie?', auto_open=True)
# + [markdown] id="CoWQv0xdVuzR" colab_type="text"
# #Graph 2: stacked bar male vs female per country
#
# + [markdown] id="zBVdE_drWdqc" colab_type="text"
# ###getting the needed info into variables
# + id="Vgua0P-D_Aqj" colab_type="code" colab={}
#creating a dataset with just males and just females
male_kiva = kiva_clean[kiva_clean['single_gender'] == 'male']
female_kiva = kiva_clean[kiva_clean['single_gender'] == 'female']
male_kiva.sample(5)
# + id="E5hGflqhqzEt" colab_type="code" colab={}
average_male = male_kiva.groupby('country')['loan_amount'].mean()
average_male
# + id="AzzXLZIHY-yO" colab_type="code" colab={}
female_kiva.sample(5)
# + id="qIi219_MrKow" colab_type="code" colab={}
average_female = female_kiva.groupby('country')['loan_amount'].mean()
average_female
# + id="mm0da1faZCfn" colab_type="code" colab={}
#finding the total loan amount by country for males and females
male_loan = male_kiva.groupby('country')['loan_amount'].sum()
female_loan = female_kiva.groupby('country')['loan_amount'].sum()
male_loan
# + [markdown] id="34bjlGj1aMuy" colab_type="text"
# ###Creating the plotly bar plot
# + id="3Bx2sSPPZudE" colab_type="code" colab={}
#running my import
import plotly.graph_objects as go
# + id="OVKyWzUQaMS5" colab_type="code" colab={}
enable_plotly_in_cell()
fig2 = go.Figure(data=[
go.Bar(name='Males', x=male_loan.index, y=male_loan.values),
go.Bar(name='Females', x=female_loan.index, y=female_loan.values)
])
fig2.update_layout(barmode='stack',
title = 'Who gets the money?: Males Vs. Females in the Kiva Dataset',
xaxis_title = 'Top 25 Countries',
yaxis_title= 'Total Amount Loaned')
fig2.show()
# + [markdown] id="K7PHRx4UfRKw" colab_type="text"
# ###pushing my plot to plotly (will be omited from github repository to protect my API key)
# + id="YztA10NNaMP0" colab_type="code" colab={}
#creating the credentials file
# username = 'zacksnyder1998'
# api_key = Insert_Api_key_here
# chart_studio.tools.set_credentials_file(username=username, api_key=api_key)
# import chart_studio.plotly as py
# py.plot(fig2, filename = 'Who gets the money?: Males Vs. Females in the Kiva Dataset', auto_open=True)
# + [markdown] id="aVxAxSo3lIxd" colab_type="text"
# #graph 3: loan amount by sector
#
# + [markdown] id="g5UC2_KZnylL" colab_type="text"
# ###Grabbing what I need
# + id="0GHeT9EEaMM_" colab_type="code" colab={}
#creating the data to draw from
sector_amounts = kiva_clean.groupby('sector')['loan_amount'].mean()
sector_amounts
# + id="dnbHpCWlaMKT" colab_type="code" colab={}
sector_numbers = kiva_clean['sector'].value_counts()
sector_numbers.sort_index(inplace=True)
sector_numbers
# + [markdown] id="W2rMY3y_n4k_" colab_type="text"
# ###Creating the visual
# + id="4np6Rk6JaMGq" colab_type="code" colab={}
enable_plotly_in_cell()
fig3 = px.bar([sector_amounts, sector_numbers], x=sector_numbers.index, y = sector_numbers.values,
color = sector_amounts.values, labels = {'color': 'Average loan Amount (USD)', 'x':'Sector', 'y': 'Total Loans Made'})
fig3.update_layout(title='Who gets the money?: Loans made and average loan amount per sector by Kiva')
fig3.show()
# + [markdown] id="0tvnJTeAFf9Q" colab_type="text"
# ###Pushing the visual to chart studio
# + id="A44c-2mKaMDs" colab_type="code" colab={}
# username = 'zacksnyder1998'
# api_key = api_key_here
# chart_studio.tools.set_credentials_file(username=username, api_key=api_key)
# import chart_studio.plotly as py
# py.plot(fig3, filename = 'Who gets the money?: Loans made and average loan amount per sector by Kiva', auto_open=True)
# + [markdown] id="xp0ZGbxaMbON" colab_type="text"
# #Graph 3: Average loan amount and number of loans per country
# + [markdown] id="1hK_OtZaM6WU" colab_type="text"
# ###Grabbing the needed data
# + id="0pAcRFagaMAb" colab_type="code" colab={}
#finding the average loan amounts per country
country_avg_loan = kiva_clean.groupby('country')['loan_amount'].mean()
country_avg_loan
# + id="OTK03EUfaL9H" colab_type="code" colab={}
#finding number of loans provided
count_amount = kiva_clean['country'].value_counts()
count_amount.sort_index(inplace=True)
count_amount
# + [markdown] id="xn78K62fQP9T" colab_type="text"
# ###creating the graph
# + id="AMEMURvEaL50" colab_type="code" colab={}
#creating a bar plot with color based on average loan amount
enable_plotly_in_cell()
fig4 = px.bar([country_avg_loan, count_amount], x=count_amount.index, y = count_amount.values,
color = country_avg_loan.values, labels = {'color': 'Average loan Amount (USD)', 'x':'Country', 'y': 'Total Loans Made'})
fig4.update_layout(title='Who gets the money?: Loans made and average loan amount per country by Kiva')
fig4.show()
# + [markdown] id="ctfbJ58gTmZK" colab_type="text"
# ###pushing this to chart_studio
#
# + id="6-kafeAiaL2i" colab_type="code" colab={}
#creating the credentials file
# username = 'zacksnyder1998'
# api_key = Api_key_here
# chart_studio.tools.set_credentials_file(username=username, api_key=api_key)
# import chart_studio.plotly as py
# py.plot(fig4, filename = 'Who gets the money?: Loans made and average loan amount per country by Kiva', auto_open=True)
# + id="79Sx6JI6Os-A" colab_type="code" colab={}
#creating an embedable link for the visualization
|
Unit_1_build_week.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://raw.githubusercontent.com/Qiskit/qiskit-tutorials/master/images/qiskit-heading.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# ## _*Entanglement*_
#
#
# The latest version of this notebook is available on https://github.com/qiskit/qiskit-tutorial.
#
# ***
# ### Contributors
# <NAME>, <NAME>, <NAME>, <NAME>
#
#
# ### Qiskit Package Versions
import qiskit
qiskit.__qiskit_version__
# ## Introduction
# Many people tend to think quantum physics is hard math, but this is not actually true. Quantum concepts are very similar to those seen in the linear algebra classes you may have taken as a freshman in college, or even in high school. The challenge of quantum physics is the necessity to accept counter-intuitive ideas, and its lack of a simple underlying theory. We believe that if you can grasp the following two Principles, you will have a good start:
# 1. A physical system in a definite state can still behave randomly.
# 2. Two systems that are too far apart to influence each other can nevertheless behave in ways that, though individually random, are somehow strongly correlated.
#
# In this tutorial, we will be discussing the second of these Principles, the first is discussed in [this other tutorial](superposition.ipynb).
# +
# useful additional packages
import matplotlib.pyplot as plt
# %matplotlib inline
# importing Qiskit
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister
from qiskit import BasicAer, IBMQ, execute
# import basic plot tools
from qiskit.tools.visualization import plot_histogram
# +
backend = BasicAer.get_backend('qasm_simulator') # run on local simulator by default
# Uncomment the following lines to run on a real device
# IBMQ.load_accounts()
# from qiskit.providers.ibmq import least_busy
# backend = least_busy(IBMQ.backends(operational=True, simulator=False))
# print("the best backend is " + backend.name())
# -
# ## Entanglement<a id='section2'></a>
#
# The core idea behind the second Principle is *entanglement*. Upon reading the Principle, one might be inclined to think that entanglement is simply strong correlation between two entitities -- but entanglement goes well beyond mere perfect (classical) correlation. If you and I read the same paper, we will have learned the same information. If a third person comes along and reads the same paper they <i>also</i> will have learned this information. All three persons in this case are perfectly correlated, and they will remain correlated even if they are separated from each other.
#
# The situation with quantum entanglement is a bit more subtle. In the quantum world, you and I could read the same quantum paper, and yet we will not learn what information is actually contained in the paper until we get together and share our information. However, when we are together, we find that we can unlock more information from the paper than we initially thought possible. Thus, quantum entanglement goes much further than perfect correlation.
#
# To demonstrate this, we will define the controlled-NOT (CNOT) gate and the composition of two systems. The convention we use Qiskit is to label states by writing the first qubit's name in the rightmost position, thereby allowing us to easily convert from binary to decimal. As a result, we define the tensor product between operators $q_0$ and $q_1$ by $q_1\otimes q_0$.
#
# Taking $q_0$ as the control and $q_1$ as the target, the CNOT with this representation is given by
#
# $$ CNOT =\begin{pmatrix} 1 & 0 & 0 & 0\\ 0 & 0 & 0 & 1\\0& 0& 1 & 0\\0 & 1 & 0 & 0 \end{pmatrix},$$
#
# which is non-standard in the quantum community, but more easily connects to classical computing, where the least significant bit (LSB) is typically on the right. An entangled state of the two qubits can be made via an $H$ gate on the control qubit, followed by the CNOT gate. This generates a particular maximally entangled two-qubit state known as a Bell state, named after <NAME> ([learn more about Bell and his contributions to quantum physics and entanglement](https://en.wikipedia.org/wiki/John_Stewart_Bell)).
#
# To explore this, we can prepare an entangled state of two qubits, and then ask questions about the qubit states. The questions we can ask are:
# * What is the state of the first qubit in the standard basis?
# * What is the state of the first qubit in the superposition basis?
# * What is the state of the second qubit in the standard basis?
# * What is the state of the second qubit in the superposition basis?
# * What is the state of both qubits in the standard basis?
# * what is the state of both qubits in the superposition basis?
#
# Below is a program with six such circuits for these six questions.
# +
# Creating registers
q2 = QuantumRegister(2)
c1 = ClassicalRegister(1)
c2 = ClassicalRegister(2)
# quantum circuit to make an entangled bell state
bell = QuantumCircuit(q2)
bell.h(q2[0])
bell.cx(q2[0], q2[1])
# quantum circuit to measure q0 in the standard basis
measureIZ = QuantumCircuit(q2, c1)
measureIZ.measure(q2[0], c1[0])
bellIZ = bell+measureIZ
# quantum circuit to measure q0 in the superposition basis
measureIX = QuantumCircuit(q2, c1)
measureIX.h(q2[0])
measureIX.measure(q2[0], c1[0])
bellIX = bell+measureIX
# quantum circuit to measure q1 in the standard basis
measureZI = QuantumCircuit(q2, c1)
measureZI.measure(q2[1], c1[0])
bellZI = bell+measureZI
# quantum circuit to measure q1 in the superposition basis
measureXI = QuantumCircuit(q2, c1)
measureXI.h(q2[1])
measureXI.measure(q2[1], c1[0])
bellXI = bell+measureXI
# quantum circuit to measure q in the standard basis
measureZZ = QuantumCircuit(q2, c2)
measureZZ.measure(q2[0], c2[0])
measureZZ.measure(q2[1], c2[1])
bellZZ = bell+measureZZ
# quantum circuit to measure q in the superposition basis
measureXX = QuantumCircuit(q2, c2)
measureXX.h(q2[0])
measureXX.h(q2[1])
measureXX.measure(q2[0], c2[0])
measureXX.measure(q2[1], c2[1])
bellXX = bell+measureXX
# -
bellIZ.draw(output='mpl')
bellIX.draw(output='mpl')
bellZI.draw(output='mpl')
bellXI.draw(output='mpl')
bellZZ.draw(output='mpl')
bellXX.draw(output='mpl')
# Let's begin by running just the first two questions, looking at the results of the first qubit ($q_0$) using a computational and then a superposition measurement.
# +
circuits = [bellIZ,bellIX,bellZI,bellXI,bellZZ,bellXX]
job = execute(circuits, backend)
result = job.result()
plot_histogram(result.get_counts(bellIZ))
# -
result.get_counts(bellIZ)
# We find that the result is random. Half the time $q_0$ is in $|0\rangle$, and the other half it is in the $|1\rangle$ state. You may wonder whether this is like the superposition from earlier in the tutorial. Maybe the qubit has a perfectly definite state, and we are simply measuring in another basis. What would you expect if you did the experiment and measured in the superposition basis? Recall we do this by adding an $H$ gate before the measurement...which is exactly what we have checked with the second question.
plot_histogram(result.get_counts(bellIX))
# In this case, we see that the result is still random, regardless of whether we measure in the computational or the superposition basis. This tells us that we actually know nothing about the first qubit. What about the second qubit, $q_1$? The next lines will run experiments measuring the second qubit in both the computational and superposition bases.
plot_histogram(result.get_counts(bellZI))
plot_histogram(result.get_counts(bellXI))
# Once again, all the experiments give random outcomes. It seems we know nothing about either qubit in our system! In our previous analogy, this is equivalent to two readers separately reading a quantum paper and extracting no information whatsoever from it on their own.
#
# What do you expect, however, when the readers get together? Below we will measure both in the joint computational basis.
plot_histogram(result.get_counts(bellZZ))
# Here we see that with high probability, if $q_0$ is in state 0, $q_1$ will be in 0 as well; the same goes if $q_0$ is in state 1. They are perfectly correlated.
#
# What about if we measure both in the superposition basis?
plot_histogram(result.get_counts(bellXX))
# Here we see that the system **also** has perfect correlations (accounting for experimental noise). Therefore, if $q_0$ is measured in state $|0\rangle$, we know $q_1$ is in this state as well; likewise, if $q_0$ is measured in state $|+\rangle$, we know $q_1$ is also in this state. These correlations have led to much confusion in science, because any attempt to relate the unusual behavior of quantum entanglement to our everyday experiences is a fruitless endeavor.
#
# Finally, we need to point out that having correlated outcomes does not necessarily imply that what we are observing is an entangled state. What would we observe, for example, if we prepared half of our shots in the $|00\rangle$ state and half of the shots in the $|11\rangle$ state? Let's have a look
# +
# quantum circuit to make a mixed state
mixed1 = QuantumCircuit(q2, c2)
mixed2 = QuantumCircuit(q2, c2)
mixed2.x(q2)
mixed1.measure(q2[0], c2[0])
mixed1.measure(q2[1], c2[1])
mixed2.measure(q2[0], c2[0])
mixed2.measure(q2[1], c2[1])
mixed1.draw(output='mpl')
# -
mixed2.draw(output='mpl')
# +
mixed_state = [mixed1,mixed2]
job = execute(mixed_state, backend)
result = job.result()
counts1 = result.get_counts(mixed_state[0])
counts2 = result.get_counts(mixed_state[1])
from collections import Counter
ground = Counter(counts1)
excited = Counter(counts2)
plot_histogram(ground+excited)
# -
# We do see the same kind of correlation indeed as we observed in the "bell_measureZZ" circuit. But we know this is not an entangled state! All we have done is leave the qubits in their ground state for some of the shots and flip both qubits for some of the shots. This is called a mixed state and it is a classical state. Now, would we observe a similar outcome if we measured this mixed state in the superposition basis? We will leave this for the reader to try.
#
# This is just a taste of what happens in the quantum world with multi-qubit states. Please continue to [Testing Entanglement](entanglement_testing.ipynb) to explore further!
|
terra/qis_intro/entanglement_introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/RohitKeshari/PyTorch-Tutorial/blob/master/CIFAR10_cnn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="lMdD_a2DUlhM" colab_type="code" colab={}
# it's specific to colab
# http://pytorch.org/
from os.path import exists
from wheel.pep425tags import get_abbr_impl, get_impl_ver, get_abi_tag
platform = '{}{}-{}'.format(get_abbr_impl(), get_impl_ver(), get_abi_tag())
# cuda_output = !ldconfig -p|grep cudart.so|sed -e 's/.*\.\([0-9]*\)\.\([0-9]*\)$/cu\1\2/'
accelerator = cuda_output[0] if exists('/dev/nvidia0') else 'cpu'
# !pip install -q http://download.pytorch.org/whl/{accelerator}/torch-0.4.1-{platform}-linux_x86_64.whl torchvision
import torch
# + [markdown] id="iFp_urAMV_lc" colab_type="text"
# **Loading and normalizing CIFAR10**
# + id="AxphNDjXU3ml" colab_type="code" colab={}
import torch
import torchvision
import torchvision.transforms as transforms
# + [markdown] id="WEyRrFDeWiIl" colab_type="text"
# *The output of torchvision datasets are PILImage images of range [0, 1]. We transform them to Tensors of normalized range [-1, 1]*
# + id="adbWRsNpVDH_" colab_type="code" outputId="2463d461-f6d9-4e67-daa9-d24f892d6dcd" colab={"base_uri": "https://localhost:8080/", "height": 51}
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=4,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=4,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# + [markdown] id="RLgqYDdfWth2" colab_type="text"
# **Let us show some of the training images, for fun**
# + id="29BLpo_aVTPw" colab_type="code" outputId="8c4d7ef9-95de-4126-8166-82bbc041b672" colab={"base_uri": "https://localhost:8080/", "height": 184}
import matplotlib.pyplot as plt
import numpy as np
# functions to show an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
# get some random training images
dataiter = iter(trainloader)
images, labels = dataiter.next()
# show images
imshow(torchvision.utils.make_grid(images))
# print labels
print(' '.join('%5s' % classes[labels[j]] for j in range(4)))
# + [markdown] id="AFV9_OLkXOnt" colab_type="text"
# Define a Convolutional Neural Network (**CNN**)
# + id="yXT9x6qBVePh" colab_type="code" colab={}
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = Net()
# + [markdown] id="2pk1KA4QVzNo" colab_type="text"
# **Define a Loss function and optimizer**
# + id="q5f-gPkWVkpm" colab_type="code" colab={}
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
# + [markdown] id="24hK-JfaXU2g" colab_type="text"
# **Train the network**
# + id="7IY8K8QzWZ_g" colab_type="code" outputId="b7fff608-b89b-4237-b296-1619e53f39a4" colab={"base_uri": "https://localhost:8080/", "height": 238}
for epoch in range(2): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
print('Finished Training')
# + [markdown] id="oZRUlR23Xis4" colab_type="text"
# **Test the network on the test data**
# + id="aosEga3UXkWR" colab_type="code" outputId="9fd58e5a-6f58-4825-d5ff-5c5be39bf484" colab={"base_uri": "https://localhost:8080/", "height": 629}
dataiter = iter(testloader)
images, labels = dataiter.next()
# print images
imshow(torchvision.utils.make_grid(images))
print('GroundTruth: ', ' '.join('%5s' % classes[labels[j]] for j in range(4)))
# + [markdown] id="iMxH2R2tXvgy" colab_type="text"
# Okay, now let us see **what the neural network thinks** these examples above are:
# + id="O6lJxJhgXqSw" colab_type="code" colab={}
outputs = net(images)
# + [markdown] id="lOrqOV6WX7cJ" colab_type="text"
# The outputs are energies for the 10 classes. The higher the energy for a class, the more the network thinks that the image is of the particular class. So,** let’s get the index of the highest energy**:
# + id="aRqMK6i-X8ng" colab_type="code" outputId="4a8b0888-245c-47b1-99d0-6d6bca0b4fdb" colab={"base_uri": "https://localhost:8080/", "height": 34}
_, predicted = torch.max(outputs, 1)
print('Predicted: ', ' '.join('%5s' % classes[predicted[j]]
for j in range(4)))
# + [markdown] id="e9lvkQuBYO4O" colab_type="text"
# The results seem pretty good.
#
# Let us look at how **the network performs on the whole dataset**.
# + id="G8oze9N1YQAx" colab_type="code" outputId="ec11a03d-1b74-4b5b-e131-93645f21b84e" colab={"base_uri": "https://localhost:8080/", "height": 34}
correct = 0
total = 0
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
# + [markdown] id="yWh3oy1qYb2e" colab_type="text"
# **That looks waaay better** than chance, which is 10% accuracy (randomly picking a class out of 10 classes). Seems like **the network learnt something**.
#
# Hmmm, what are the **classes that performed well**, and the classes that did not perform well:
# + id="jjooR7JxYcuc" colab_type="code" outputId="9967d3d7-2173-4bbb-f7fb-231cc36a6bfe" colab={"base_uri": "https://localhost:8080/", "height": 187}
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predicted = torch.max(outputs, 1)
c = (predicted == labels).squeeze()
for i in range(4):
label = labels[i]
class_correct[label] += c[i].item()
class_total[label] += 1
for i in range(10):
print('Accuracy of %5s : %2d %%' % (
classes[i], 100 * class_correct[i] / class_total[i]))
# + [markdown] id="PlATrqFOYtLK" colab_type="text"
# Training on GPU
# + [markdown] id="KNbc6SmSYw-V" colab_type="text"
# Just like how you transfer a Tensor onto the GPU, you transfer the neural net onto the GPU.
#
# Let’s first define our device as the first visible cuda device if we have CUDA available:
# + id="OqAhOFqwYuH7" colab_type="code" outputId="1a0b5771-d243-4b6f-fd73-40134acff8c3" colab={"base_uri": "https://localhost:8080/", "height": 34}
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Assuming that we are on a CUDA machine, this should print a CUDA device:
print(device)
# + id="QIeFs2jaQOQQ" colab_type="code" outputId="98ed0d64-a9f1-4b36-86a8-54882937d631" colab={"base_uri": "https://localhost:8080/", "height": 34}
torch.cuda.current_device()
# + id="pxL9Z4W3QUpe" colab_type="code" outputId="83292c3a-480b-4bcc-ac24-cbb1d605ca02" colab={"base_uri": "https://localhost:8080/", "height": 34}
torch.cuda.device(0)
# + id="3Enf_QjDQaC7" colab_type="code" outputId="cee313b3-425c-485f-9a87-337be0b29f55" colab={"base_uri": "https://localhost:8080/", "height": 34}
torch.cuda.device_count()
# + id="o8CEs2kFQe71" colab_type="code" outputId="d56b1406-5bd7-4227-90e3-5d6382c26010" colab={"base_uri": "https://localhost:8080/", "height": 34}
torch.cuda.get_device_name(0)
# + [markdown] id="rRC1DFAdY8Fo" colab_type="text"
# The rest of this section assumes that device is a CUDA device.
#
# Then these methods will recursively go over all modules and convert their parameters and buffers to CUDA tensors
# + id="p4nVN2qEY8_N" colab_type="code" outputId="b5cabfca-41d8-4e33-8007-e37349deeaf2" colab={"base_uri": "https://localhost:8080/", "height": 153}
net.to(device)
# + [markdown] id="Yi631xvVZFNE" colab_type="text"
# Remember that you will have to send the inputs and targets at every step to the GPU too:
# + id="uvc9RY9tZg0P" colab_type="code" outputId="84c506a3-b954-478f-db97-9a12c3c6bcd6" colab={"base_uri": "https://localhost:8080/", "height": 238}
#net=net.cuda()
criterion = nn.CrossEntropyLoss().cuda()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
for epoch in range(2): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.cuda(), labels.cuda()
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
print('Finished Training')
# + [markdown] id="-YnBbkAVWpIJ" colab_type="text"
# Why dont I notice MASSIVE speedup compared to CPU? Because your **network is realllly small**.
# + [markdown] id="WeeESGuzW5xh" colab_type="text"
# **Exercise:** Try increasing the width of your network (argument 2 of the first nn.Conv2d, and argument 1 of the second nn.Conv2d – they need to be the same number), see what kind of speedup you get.
|
CIFAR10_cnn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
tf.set_random_seed(1)
np.random.seed(1)
# fake data
x = np.linspace(-1, 1, 100)[:, np.newaxis] # shape (100, 1)
noise = np.random.normal(0, 0.1, size=x.shape)
y = np.power(x, 2) + noise # shape (100, 1) + some noise
# plot data
plt.scatter(x, y)
plt.show()
tf_x = tf.placeholder(tf.float32, x.shape) # input x
tf_y = tf.placeholder(tf.float32, y.shape) # input y
# neural network layers
l1 = tf.layers.dense(tf_x, 10, tf.nn.relu) # hidden layer
output = tf.layers.dense(l1, 1) # output layer
loss = tf.losses.mean_squared_error(tf_y, output) # compute cost
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.5)
train_op = optimizer.minimize(loss)
sess = tf.Session() # control training and others
sess.run(tf.global_variables_initializer()) # initialize var in graph
plt.ion() # something about plotting
for step in range(100):
# train and net output
_, l, pred = sess.run([train_op, loss, output], {tf_x: x, tf_y: y})
if step % 5 == 0:
# plot and show learning process
plt.cla()
plt.scatter(x, y)
plt.plot(x, pred, 'r-', lw=5)
plt.text(0.5, 0, 'Loss=%.4f' % l, fontdict={'size': 20, 'color': 'red'})
plt.pause(0.1)
plt.ioff()
plt.show()
|
TensorflowTUT2/301_simple_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
import copy
import logging
import sys
import os
import sys
import importlib
import numpy as np
from collections import defaultdict
sys.path.insert(0, '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc')
from tools_pattern import get_eucledean_dist
import compress_pickle
import my_plot
from my_plot import MyPlotData, my_box_plot
import seaborn as sns
script_n = 'plot_210614_1ba_signal_vs_noise_10_ref_delta'
data_script = 'batch_210608_stability_vs_redundancy_1ba'
db_path = '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/dimensionality_sim2/' \
f'{data_script}/'
n_mfs = 488
n_grcs = 1459
pattern_type = 'binary'
db = {}
direction = '10'
noise = '0.2'
model = 'shuffle'
db[model] = compress_pickle.load(
db_path+f'{data_script}_{model}_{pattern_type}_{n_grcs}_{n_mfs}_dir_{direction}_noise_{noise}_0.3_256_40.gz')
model = 'global_random'
db[model] = compress_pickle.load(
db_path+f'{data_script}_{model}_{pattern_type}_{n_grcs}_{n_mfs}_dir_{direction}_noise_{noise}_0.3_256_40.gz')
def get_average_signal_strength(hist_sum):
return sum(hist_sum)/len(hist_sum)
def get_signal_variance(hist_sum):
return np.std(hist_sum, ddof=1)
def get_low_signal_val(hist_sum, pct=.025):
return sorted(hist_sum, reverse=False)[int(len(hist_sum)*pct)]
def get_signal_variance_width(hist_sum):
hist_sum = sorted(hist_sum)
return hist_sum[int(.95*len(hist_sum))] - hist_sum[int(.05*len(hist_sum))]
def get_signal_loss(hist_sum, ref_sum):
hist_sum = sorted(hist_sum)
return ref_sum- hist_sum[int(.5*len(hist_sum))]
# avg_grc_dim_list = defaultdict(list)
# for ress in db['random']:
# ress_tries = ress
# for ress in ress_tries:
# # print(ress)
# for noise in ress:
# res = ress[noise]
# grc_dim = res['grc_dim']
# avg_grc_dim_list[noise].append(grc_dim)
# avg_grc_dim = {}
# for noise in avg_grc_dim_list:
# avg_grc_dim[noise] = sum(avg_grc_dim_list[noise])/len(avg_grc_dim_list[noise])
# +
name_map = {
'scaleup4': "Observed",
'global_random': "Global Random",
'random': "Global Random",
# 'naive_random_17': "Local Random",
'shuffle': "Shuffle",
}
palette = {
name_map['scaleup4']: sns.color_palette()[0],
name_map['global_random']: sns.color_palette()[1],
name_map['random']: sns.color_palette()[1],
name_map['shuffle']: sns.color_palette()[2],
# name_map['naive_random_21']: sns.color_palette()[2],
}
mpd = MyPlotData()
ress_ref = db['shuffle'][0][0]
resss_ref2 = db['shuffle'][0]
for model_name in [
'shuffle',
'global_random',
]:
ress = db[model_name]
ress_tries = ress[0] # get the first element in tuple
for n_try, ress in enumerate(ress_tries):
if n_try >= len(resss_ref2):
print(n_try)
continue
ress_ref2 = resss_ref2[n_try]
for noise in ress:
res = ress[noise]
res_ref2 = ress_ref2[noise]
mpd.add_data_point(
model=name_map[model_name],
avg_signal=get_average_signal_strength(res['hist_sum']),
ref_delta=res['ref_delta'],
variance=get_signal_variance(res['hist_sum']),
low_signal=get_low_signal_val(res['hist_sum']),
variance_width=get_signal_variance_width(res['hist_sum']),
signal_loss=get_signal_loss(res['hist_sum'], res['ref_sum1']),
noise=noise,
)
# +
importlib.reload(my_plot); my_plot.my_relplot(
mpd,
x='noise',
y='ref_delta',
hue='model',
context='paper',
palette=palette,
linewidth=1,
width=10,
# ylim=[0, None],
# xlim=[0, 150],
y_axis_label='Dim. Norm. ($x$)',
x_axis_label='GrC count (%)',
save_filename=f'{script_n}.svg',
show=True,
)
|
analysis/dimensionalty_sim/plot_210614_1ba_signal_vs_noise_10_ref_delta.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Conv2D, MaxPooling2D
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# +
AUTOTUNE = tf.data.experimental.AUTOTUNE
train_ds = train_ds.cache().shuffle(1000).prefetch(buffer_size=AUTOTUNE)
val_ds = val_ds.cache().prefetch(buffer_size=AUTOTUNE)
# +
num_classes = 5
model = Sequential([
layers.experimental.preprocessing.Rescaling(1./255, input_shape=(img_height, img_width, 3)),
layers.Conv2D(16, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(32, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(64, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dense(num_classes)
])
# +
img_width, img_height = 150, 150
data_dir = 'data/cat_dog_train'
validation_data_dir = 'data/cat_dog_val'
nb_train_samples = 2000
nb_validation_samples = 800
epochs = 50
batch_size = 16
if K.image_data_format() == 'channels_first':
input_shape = (3, img_width, img_height)
else:
input_shape = (img_width, img_height, 3)
model = Sequential()
model.add(Conv2D(32, (3, 3), input_shape=input_shape))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(32, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(64))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(1))
model.add(Activation('sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
# -
def define_model(nb_filters, kernel_size, input_shape, pool_size):
model = Sequential() # model is a linear stack of layers (don't change)
# note: the convolutional layers and dense layers require an activation function
# see https://keras.io/activations/
# and https://en.wikipedia.org/wiki/Activation_function
# options: 'linear', 'sigmoid', 'tanh', 'relu', 'softplus', 'softsign'
model.add(Conv2D(nb_filters,
(kernel_size[0], kernel_size[1]),
padding='valid',
input_shape=input_shape)) # first conv. layer KEEP
model.add(Activation('relu')) # Activation specification necessary for Conv2D and Dense layers
model.add(Conv2D(nb_filters,
(kernel_size[0], kernel_size[1]),
padding='valid')) # 2nd conv. layer KEEP
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=pool_size)) # decreases size, helps prevent overfitting
model.add(Dropout(0.5)) # zeros out some fraction of inputs, helps prevent overfitting
model.add(Flatten()) # necessary to flatten before going into conventional dense layer KEEP
print('Model flattened out to ', model.output_shape)
# now start a typical neural network
model.add(Dense(32)) # (only) 32 neurons in this layer, really? KEEP
model.add(Activation('tanh'))
model.add(Dropout(0.5)) # zeros out some fraction of inputs, helps prevent overfitting
model.add(Dense(nb_classes)) # 10 final nodes (one for each class) KEEP
model.add(Activation('softmax')) # softmax at end to pick between classes 0-9 KEEP
# many optimizers available, see https://keras.io/optimizers/#usage-of-optimizers
# suggest you KEEP loss at 'categorical_crossentropy' for this multiclass problem,
# and KEEP metrics at 'accuracy'
# suggest limiting optimizers to one of these: 'adam', 'adadelta', 'sgd'
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
return model
batch_size = 2000 # number of training samples used at a time to update the weights
nb_classes = 10 # number of output possibilities: [0 - 9] KEEP
nb_epoch = 10 # number of passes through the entire train dataset before weights "final"
img_rows, img_cols = 28, 28 # the size of the MNIST images KEEP
input_shape = (img_rows, img_cols, 1) # 1 channel image input (grayscale) KEEP
nb_filters = 50 # number of convolutional filters to use
pool_size = (2, 2) # pooling decreases image size, reduces computation, adds translational invariance
kernel_size = (4, 4) # convolutional kernel size, slides over image to learn features
plt.imshow(img_hsv[:,:,0], cmap='gray');
X_train, X_test, Y_train, Y_test = load_and_featurize_data()
model = define_model(nb_filters, kernel_size, input_shape, pool_size)
# +
# during fit process watch train and test error simultaneously
model.fit(X_train, Y_train, batch_size=batch_size, epochs=nb_epoch,
verbose=1, validation_data=(X_test, Y_test))
score = model.evaluate(X_test, Y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1]) # this is the one we care about
|
notebooks/archive/run_models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from face_recognition import load_image_file, face_encodings
# +
# load img to numpy array
img_array = load_image_file("images/merkel.jpg")
# Generates a list of all faces in the image where each face is represented as a list of 128 face encodings that are found by a pretrainend CNN. Note that it will only find faces where its landmarkpoints are visible.
encodings = face_encodings(img_array)
n_faces = len(encodings)
print("Number of faces:", n_faces)
# -
if n_faces > 0:
print(encodings[0])
|
3_face_encoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# Any results you write to the current directory are saved as output.
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set()
# -
# ---
# # Load datasets
# +
train = pd.read_csv('../input/titanic/train.csv')
test = pd.read_csv('../input/titanic/test.csv')
dataset = [train, test]
# -
# ### Understand the loaded data and think about how to do feature engineering or data cleansing.
train.shape, test.shape
train.head()
# 1. Need to extract information as categorical variables.
# 2. Convert to a numerical value.
# 3. Remove the missing value.
train.isnull().sum()
# #### There are 177 missing values in Age, 687 missing values in Cabin, and 2 missing values in Embarked.
test.isnull().sum()
# #### There are 86 missing values in Age, 1 missing value in Fare, and 327 missing values in Cabin
# ---
# # Visualization
def visualization(feature):
O = train[train['Survived'] == 1][feature].value_counts()
X = train[train['Survived'] == 0][feature].value_counts()
visual_df = pd.DataFrame([O, X])
visual_df.index = ['Survived', 'Dead']
visual_df.plot(kind = 'bar', stacked=True, figsize=(12, 5), title=feature)
visualization('Pclass')
# #### If Pclass is 1, likely to survive, and if it is 3, likely to die.
visualization('Sex')
# #### If Sex is female, likely to survive, and if it is male, likely to die.
visualization('Embarked')
# #### If Embarked is S, likely to die, if it is C, likely to survive, and if it is Q, likely to die.
# ---
# # Feature Engineering
# #### Merge SibSP and Parch, add 1, and create a new feature called Family.
# +
for vector in dataset:
vector['Family'] = vector['SibSp'] + vector['Parch'] + 1
train[['Family', 'Survived']].groupby(['Family'], as_index=False).mean()
# -
# #### Create a feature called Alone to check if boarding with family affects survival.
# +
for vector in dataset:
vector['Alone'] = 0 # boarded with family
vector.loc[vector['Family'] == 1, 'Alone'] = 1 # boarded alone
train[['Alone', 'Survived']].groupby(['Alone'], as_index=False).mean()
# -
# #### Boarded alone shows a survival rate of 50% and boarded with a family shows a survival rate of 30%.
# #### SibSP, Parch, and Family features used to create the Alone feature are now unnecessary, so delete them.
train = train.drop(['SibSp', 'Parch', 'Family'], axis=1)
test = test.drop(['SibSp', 'Parch', 'Family'], axis=1)
# #### Replace 2 missing values of Embarked with S, the largest amount of data.
train['Embarked'] = train['Embarked'].fillna('S')
test['Embarked'] = test['Embarked'].fillna('S')
# #### Fare's missing values are grouped into Pclass and replaced with the median of 1, 2, 3.
# #### Visualize to see the distribution of the fare.
# +
facet = sns.FacetGrid(train, aspect=4)
facet.map(sns.kdeplot, 'Fare', shade=True)
facet.set(xlim = (0, train['Fare'].max()))
facet.add_legend()
plt.show()
# -
# #### To utilize the Fare data, create 4 groups and put similar amounts of data in each group.
# +
train['Fare_Division'] = pd.qcut(train['Fare'], 4)
test['Fare_Division'] = pd.qcut(test['Fare'],4 )
train[['Fare_Division', 'Survived']].groupby(['Fare_Division'], as_index=False).mean()
# -
# #### Extract Title data from Name data.
# +
train['Title'] = train['Name'].str.extract('([A-Za-z]+)\.', expand=False)
test['Title'] = test['Name'].str.extract('([A-Za-z]+)\.', expand=False)
train = train.drop(['Name'], axis=1)
test = test.drop(['Name'], axis=1)
# -
# #### Create a group for each Title and replace Age's missing value with the average of the Title.
train['Title'].value_counts()
test['Title'].value_counts()
train['Age'].fillna(train.groupby('Title')['Age'].transform('median'), inplace=True)
test['Age'].fillna(test.groupby('Title')['Age'].transform('median'), inplace=True)
train['Age'].isnull().sum(), test['Age'].isnull().sum()
# #### 1 missing value remains in test data.
test[test['Age'].isnull()]
# #### Because only one data has Title MS in the test data, the missing value is not replaced.
# #### The data is replaced with the average value of the train data.
train[train['Title'] == 'Ms']['Age'].value_counts()
test.loc[test['Age'].isnull(), 'Age'] = 28
# #### Visualize the distribution of age data and divide it into 5 groups.
# +
facet = sns.FacetGrid(train, aspect=4)
facet.map(sns.kdeplot, 'Age', shade=True)
facet.set(xlim = (0, train['Age'].max()))
facet.add_legend()
plt.show()
# +
train['Age_Division'] = pd.qcut(train['Age'], 5)
test['Age_Division'] = pd.qcut(test['Age'], 5)
train[['Age_Division', 'Survived']].groupby(['Age_Division'], as_index=False).mean()
# -
# #### Cabin data is excluded because the percentage of missing values is 77%.
# ---
# # Data Cleansing
train.head()
sexMap = {'male':0, 'female':1}
train['Sex'] = train['Sex'].map(sexMap)
test['Sex'] = test['Sex'].map(sexMap)
embarkedMap = {'S':0, 'C':1, 'Q':2}
train['Embarked'] = train['Embarked'].map(embarkedMap)
test['Embarked'] = test['Embarked'].map(embarkedMap)
titleMap = {'Mr':0, 'Miss':1, 'Mrs':2, 'Marster':3, 'Dr':3, 'Rev':3, 'Mlle':3, 'Col':3, 'Major':3,
'Ms':3, 'Lady':3, 'Jonkheer':3, 'Mme':3, 'Capt':3, 'Sir':3, 'Don':3, 'Countess':3}
train['Title'] = train['Title'].map(titleMap)
test['Title'] = test['Title'].map(titleMap)
# +
train.loc[train['Fare'] <= 7.91, 'Fare'] = 0
train.loc[(7.91 < train['Fare']) & (train['Fare'] <= 14.454), 'Fare'] = 1
train.loc[(14.454 < train['Fare']) & (train['Fare'] <= 31), 'Fare'] = 2
train.loc[31 < train['Fare'], 'Fare'] = 3
test.loc[test['Fare'] <= 7.91, 'Fare'] = 0
test.loc[(7.91 < test['Fare']) & (test['Fare'] <= 14.454), 'Fare'] = 1
test.loc[(14.454 < test['Fare']) & (test['Fare'] <= 31), 'Fare'] = 2
test.loc[31 < test['Fare'], 'Fare'] = 3
# +
train.loc[train['Age'] <= 20, 'Age'] = 0
train.loc[(20 < train['Age']) & (train['Age'] <= 26), 'Age'] = 1
train.loc[(26 < train['Age']) & (train['Age'] <= 30), 'Age'] = 2
train.loc[(30 < train['Age']) & (train['Age'] <= 38), 'Age'] = 3
train.loc[38 < train['Age'], 'Age'] = 4
test.loc[test['Age'] <= 20, 'Age'] = 0
test.loc[(20 < test['Age']) & (test['Age'] <= 26), 'Age'] = 1
test.loc[(26 < test['Age']) & (test['Age'] <= 30), 'Age'] = 2
test.loc[(30 < test['Age']) & (test['Age'] <= 38), 'Age'] = 3
test.loc[38 < test['Age'], 'Age'] = 4
# -
train = train.drop(['PassengerId', 'Ticket', 'Cabin', 'Fare_Division', 'Age_Division'], axis=1)
test = test.drop(['Ticket', 'Cabin', 'Fare_Division', 'Age_Division'], axis=1)
train.head()
# ---
# # Gradient Descent Algorithm & Logistic Regression
# +
train['tmp'] = 1
test['tmp'] = 1
train_df = pd.DataFrame(train, columns=['tmp', 'Pclass', 'Sex', 'Age', 'Fare', 'Alone', 'Title'])
test_df = pd.DataFrame(test, columns=['tmp', 'Pclass', 'Sex', 'Age', 'Fare', 'Alone', 'Title'])
target = train['Survived']
train_list = train_df.values.tolist()
train_list = np.array(train_list)
test_list = test_df.values.tolist()
test_list = np.array(test_list)
target_list = target.values.tolist()
target_list = np.array(target_list)
# -
# #### tmp is an added value for later use in partial derivatives to compute the loss function.
# +
import random
w_list = np.zeros(7)
for i in range(0, 7):
w_list[i] = random.random() * 2 - 1
for i in range(0, 7):
print(w_list[i])
a = 0.0001
# +
cnt = 0
while(cnt < 300000):
Zi = train_list[:].dot(w_list) # Zi = w0 + w1 * x1 + w2 * x2 + ... wn * xn
Hi = 1 / (1 + np.exp(-Zi)) # Hi = 1 / (1 + e^(-Zi))
Hi_y = Hi - target_list
for i in range(0, 7):
w_list[i] = w_list[i] - a * np.sum(train_list[:, i] * Hi_y) / 891
cnt = cnt + 1
# -
answer_list = []
for i in range(0, 418):
Zx = test_list[i].dot(w_list)
if(Zx >= 0):
answer_list.append(1)
else:
answer_list.append(0)
submission_df = pd.DataFrame({
"PassengerId":test["PassengerId"],
"Survived":answer_list
})
submission_df.to_csv('submission.csv', index=False)
|
titanic/titanic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="cPWEkZgEXNuH" colab_type="code" colab={}
"""
This work is inspired by this tutorial
https://www.youtube.com/watch?v=ws-ZbiFV1Ms&t=1116s
https://github.com/Hvass-Labs/TensorFlow-Tutorials/blob/master/14_DeepDream.ipynb
"""
# + id="sNu1oYHI7yOj" colab_type="code" colab={}
import numpy as np
import tensorflow as tf
import pandas as pd
import math
from PIL import Image
from IPython.display import Image as imshow
from scipy.ndimage.filters import gaussian_filter
# + id="uXVUQ7fsEupG" colab_type="code" outputId="2f5b5a0f-f87f-4401-dd08-361dd95204f9" colab={"base_uri": "https://localhost:8080/", "height": 406}
# downloading files for inception network
# !wget https://raw.githubusercontent.com/ElephantHunters/Deep-Dream-using-Tensorflow/master/download.py
# !wget https://raw.githubusercontent.com/ElephantHunters/Deep-Dream-using-Tensorflow/master/inception5h.py
# + id="4dKYxMK0JOVy" colab_type="code" outputId="dfccaa24-d13c-4a3c-8e28-891186f9e2f7" colab={"base_uri": "https://localhost:8080/", "height": 84}
import inception5h
inception5h.maybe_download()
# + id="8HnPfDQcJ1Di" colab_type="code" outputId="5e812453-ac81-4de4-f523-99c373cae655" colab={"base_uri": "https://localhost:8080/", "height": 87}
# importing model
model = inception5h.Inception5h()
# + id="EYyDtCwoKCUM" colab_type="code" colab={}
# functions for image processing
def load_img(loc):
"""
function to load images
loc: location of the image on the disk
"""
return np.float32(Image.open(loc))
def save_img(img, name):
"""
img: np array of the image
name: save name
functions saves the imageon disk
"""
# Ensure the pixel-values are between 0 and 255.
image = np.clip(img, 0.0, 255.0)
# Convert to bytes.
image = image.astype(np.uint8)
# Write the image-file in jpeg-format.
with open(name, 'wb') as file:
Image.fromarray(image).save(file, 'jpeg')
def show_img(img):
"""
img: path of image on disk
function to display images stored on disk
"""
return imshow(img)
# + id="ybGQt9r0QNEX" colab_type="code" colab={}
def img_gradient(gradient, img):
"""
gradient: gradient of the image
img: actual input image
function to calculate the gradient of the image
"""
# make the feed_dict of the image
feed_input = model.create_feed_dict(image = img)
grad = session.run(gradient, feed_dict=feed_input)
# normalizing the gradients
grad /= (np.std(grad) + 1e-8)
return grad
# + id="9xTl925z5C1w" colab_type="code" colab={}
def optimize_image(layer_tensor, image, epochs=10, learning_rate=3.0, show_gradient=False):
"""
Use gradient ascent to optimize an image so it maximizes the
mean value of the given layer_tensor.
Parameters:
layer_tensor: Reference to a tensor that will be maximized.
image: Input image used as the starting point.
show_gradient: Plot the gradient in each iteration.
"""
# making a copy of image
img = image.copy()
# get the gradient function w.r.t. image
gradient = model.get_gradient(layer_tensor)
# training loop
for i in range(epochs):
grad = img_gradient(gradient, img)
# applying gaussian blur to the image several times to make the image smooth
sigma = (i * 4.0) / epochs + 0.5 ## yes i know i took it from the tutorial!
grad_gauss_1 = gaussian_filter(grad, sigma=sigma)
grad_gauss_2 = gaussian_filter(grad, sigma=sigma*0.5)
grad_gauss_3 = gaussian_filter(grad, sigma=sigma*2.0)
# adding the blurred gradients together
grad = (grad_gauss_1 + grad_gauss_2 + grad_gauss_3)
# reshaping gradient according to image dimensions
grad = grad.reshape([img.shape[0], img.shape[1], img.shape[2]])
# updating the image by adding the gradient to it
img += grad*learning_rate
if i%5 == 0:
print(" >> Iteration " , i, " complete!")
print(" >> Training complete!")
return img
# + id="vdV-UrRy_8e5" colab_type="code" colab={}
# running tensorflow session
session = tf.InteractiveSession(graph=model.graph)
# + id="kxyxa16PCHp1" colab_type="code" outputId="ce32eb6d-d14e-486b-ebdf-f214f049aa4d" colab={"base_uri": "https://localhost:8080/", "height": 290}
# input image
input_image = load_img("subject_img.jpg")
input_image1 = load_img("subject_img1.jpg")
show_img("subject_img.jpg")
# + id="JYrYlNl3OvNC" colab_type="code" outputId="13dc7f53-767a-44aa-af79-ea69a1fc95f5" colab={"base_uri": "https://localhost:8080/", "height": 867}
show_img("subject_img1.jpg")
# + id="xiVJdiF3EQcE" colab_type="code" colab={}
# choosing a hidden convolutional layer from the inception model
layer_tensor = model.layer_tensors[6]
# + id="_y-BoLBnEweG" colab_type="code" outputId="5da5dc4b-9b11-4179-a3d5-f65dbbb00bac" colab={"base_uri": "https://localhost:8080/", "height": 252}
result = optimize_image(layer_tensor, input_image, epochs=30, learning_rate=7.0)
result1 = optimize_image(layer_tensor, input_image1, epochs=30, learning_rate=7.0)
# + id="hLFp4my7GgyT" colab_type="code" colab={}
# saving result image to disk
save_img(result, "result.jpg")
save_img(result1, "result1.jpg")
# + id="l-pk1pjkHS1b" colab_type="code" outputId="58342653-4ea1-47f7-921b-cf5db9f0dfca" colab={"base_uri": "https://localhost:8080/", "height": 290}
show_img("result.jpg")
# + id="Q_gaCoajO4W8" colab_type="code" outputId="917eecfb-8a50-4aaa-99e9-c9faf0ff3044" colab={"base_uri": "https://localhost:8080/", "height": 867}
show_img("result1.jpg")
|
Deep Dream/Deep_Dream.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
#
# ___
# # Ecommerce Purchases Exercise
#
# In this Exercise you will be given some Fake Data about some purchases done through Amazon! Just go ahead and follow the directions and try your best to answer the questions and complete the tasks. Feel free to reference the solutions. Most of the tasks can be solved in different ways. For the most part, the questions get progressively harder.
#
# Please excuse anything that doesn't make "Real-World" sense in the dataframe, all the data is fake and made-up.
#
# Also note that all of these questions can be answered with one line of code.
# ____
# **Import pandas and read in the Ecommerce Purchases csv file and set it to a DataFrame called ecom.**
import pandas as pd
ecom = pd.read_csv('Ecommerce Purchases.csv')
# **Check the head of the DataFrame.**
ecom.head()
# **How many rows and columns are there?**
ecom.info()
# **What is the average Purchase Price?**
ecom['Purchase Price'].mean()
# **What were the highest and lowest purchase prices?**
max_price = ecom['Purchase Price'].max()
min_price = ecom['Purchase Price'].min()
print("Max price was : ",max_price)
print("Min price was : ",min_price)
# **How many people have English 'en' as their Language of choice on the website?**
ecom[ecom['Language']=='en'].count()
# **How many people have the job title of "Lawyer" ?**
#
ecom[ecom['Job']=='Lawyer'].count()
# **How many people made the purchase during the AM and how many people made the purchase during PM ?**
#
# **(Hint: Check out [value_counts()](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.value_counts.html) )**
am_purchase = ecom[ecom['AM or PM']=='AM'].count()['AM or PM']
pm_purchase = ecom[ecom['AM or PM']=='PM'].count()['AM or PM']
print("Purchases Made during AM : ",am_purchase)
print("Purchases Made during PM : ",pm_purchase)
# Alternate way using the recommended method would be :
ecom['AM or PM'].value_counts()
# **What are the 5 most common Job Titles?**
ecom['Job'].value_counts().head(5)
# **Someone made a purchase that came from Lot: "90 WT" , what was the Purchase Price for this transaction?**
ecom[ecom['Lot']=="90 WT"]['Purchase Price']
# **What is the Email of the person with the following Credit Card Number: 4926535242672853**
ecom[ecom['Credit Card']==4926535242672853]['Email']
# **How many people have American Express as their Credit Card Provider *and* made a purchase above $95 ?**
ecom[((ecom['CC Provider']=='American Express')&(ecom['Purchase Price']>95))].count()
# **Hard: How many people have a credit card that expires in 2025?**
sum(ecom['CC Exp Date'].apply(lambda x: x[3:]) == '25')
# **Hard: What are the top 5 most popular email providers/hosts (e.g. gmail.com, yahoo.com, etc...)**
ecom['Email'].apply(lambda x: x.split('@')[1]).value_counts().head(5)
# # Great Job!
|
07. Pandas Exercises/7.2 ecommerce_purchases_exercise .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D5_DimensionalityReduction/W1D5_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text"
# # Tutorial 1: Geometric view of data
# **Week 1, Day 5: Dimensionality Reduction**
#
# **By Neuromatch Academy**
#
# __Content creators:__ <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# + [markdown] colab_type="text"
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# + [markdown] colab_type="text"
# ---
# # Tutorial Objectives
#
# In this notebook we'll explore how multivariate data can be represented in different orthonormal bases. This will help us build intuition that will be helpful in understanding PCA in the following tutorial.
#
# Overview:
# - Generate correlated multivariate data.
# - Define an arbitrary orthonormal basis.
# - Project the data onto the new basis.
# + cellView="form"
# @title Video 1: Geometric view of data
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="THu9yHnpq9I", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# + [markdown] colab_type="text"
# ---
# # Setup
# + colab={} colab_type="code"
# Import
import numpy as np
import matplotlib.pyplot as plt
# + cellView="form" colab={} colab_type="code"
# @title Figure Settings
import ipywidgets as widgets # interactive display
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# + cellView="form" colab={} colab_type="code"
# @title Helper functions
def get_data(cov_matrix):
"""
Returns a matrix of 1000 samples from a bivariate, zero-mean Gaussian.
Note that samples are sorted in ascending order for the first random variable
Args:
cov_matrix (numpy array of floats): desired covariance matrix
Returns:
(numpy array of floats) : samples from the bivariate Gaussian, with each
column corresponding to a different random
variable
"""
mean = np.array([0, 0])
X = np.random.multivariate_normal(mean, cov_matrix, size=1000)
indices_for_sorting = np.argsort(X[:, 0])
X = X[indices_for_sorting, :]
return X
def plot_data(X):
"""
Plots bivariate data. Includes a plot of each random variable, and a scatter
plot of their joint activity. The title indicates the sample correlation
calculated from the data.
Args:
X (numpy array of floats) : Data matrix each column corresponds to a
different random variable
Returns:
Nothing.
"""
fig = plt.figure(figsize=[8, 4])
gs = fig.add_gridspec(2, 2)
ax1 = fig.add_subplot(gs[0, 0])
ax1.plot(X[:, 0], color='k')
plt.ylabel('Neuron 1')
plt.title('Sample var 1: {:.1f}'.format(np.var(X[:, 0])))
ax1.set_xticklabels([])
ax2 = fig.add_subplot(gs[1, 0])
ax2.plot(X[:, 1], color='k')
plt.xlabel('Sample Number')
plt.ylabel('Neuron 2')
plt.title('Sample var 2: {:.1f}'.format(np.var(X[:, 1])))
ax3 = fig.add_subplot(gs[:, 1])
ax3.plot(X[:, 0], X[:, 1], '.', markerfacecolor=[.5, .5, .5],
markeredgewidth=0)
ax3.axis('equal')
plt.xlabel('Neuron 1 activity')
plt.ylabel('Neuron 2 activity')
plt.title('Sample corr: {:.1f}'.format(np.corrcoef(X[:, 0], X[:, 1])[0, 1]))
plt.show()
def plot_basis_vectors(X, W):
"""
Plots bivariate data as well as new basis vectors.
Args:
X (numpy array of floats) : Data matrix each column corresponds to a
different random variable
W (numpy array of floats) : Square matrix representing new orthonormal
basis each column represents a basis vector
Returns:
Nothing.
"""
plt.figure(figsize=[4, 4])
plt.plot(X[:, 0], X[:, 1], '.', color=[.5, .5, .5], label='Data')
plt.axis('equal')
plt.xlabel('Neuron 1 activity')
plt.ylabel('Neuron 2 activity')
plt.plot([0, W[0, 0]], [0, W[1, 0]], color='r', linewidth=3,
label='Basis vector 1')
plt.plot([0, W[0, 1]], [0, W[1, 1]], color='b', linewidth=3,
label='Basis vector 2')
plt.legend()
plt.show()
def plot_data_new_basis(Y):
"""
Plots bivariate data after transformation to new bases.
Similar to plot_data but with colors corresponding to projections onto
basis 1 (red) and basis 2 (blue). The title indicates the sample correlation
calculated from the data.
Note that samples are re-sorted in ascending order for the first
random variable.
Args:
Y (numpy array of floats): Data matrix in new basis each column
corresponds to a different random variable
Returns:
Nothing.
"""
fig = plt.figure(figsize=[8, 4])
gs = fig.add_gridspec(2, 2)
ax1 = fig.add_subplot(gs[0, 0])
ax1.plot(Y[:, 0], 'r')
plt.xlabel
plt.ylabel('Projection \n basis vector 1')
plt.title('Sample var 1: {:.1f}'.format(np.var(Y[:, 0])))
ax1.set_xticklabels([])
ax2 = fig.add_subplot(gs[1, 0])
ax2.plot(Y[:, 1], 'b')
plt.xlabel('Sample number')
plt.ylabel('Projection \n basis vector 2')
plt.title('Sample var 2: {:.1f}'.format(np.var(Y[:, 1])))
ax3 = fig.add_subplot(gs[:, 1])
ax3.plot(Y[:, 0], Y[:, 1], '.', color=[.5, .5, .5])
ax3.axis('equal')
plt.xlabel('Projection basis vector 1')
plt.ylabel('Projection basis vector 2')
plt.title('Sample corr: {:.1f}'.format(np.corrcoef(Y[:, 0], Y[:, 1])[0, 1]))
plt.show()
# + [markdown] colab_type="text"
# ---
# # Section 1: Generate correlated multivariate data
# + cellView="form"
# @title Video 2: Multivariate data
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="jcTq2PgU5Vw", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# + [markdown] colab_type="text"
# To gain intuition, we will first use a simple model to generate multivariate data. Specifically, we will draw random samples from a *bivariate normal distribution*. This is an extension of the one-dimensional normal distribution to two dimensions, in which each $x_i$ is marginally normal with mean $\mu_i$ and variance $\sigma_i^2$:
#
# \begin{align}
# x_i \sim \mathcal{N}(\mu_i,\sigma_i^2).
# \end{align}
#
# Additionally, the joint distribution for $x_1$ and $x_2$ has a specified correlation coefficient $\rho$. Recall that the correlation coefficient is a normalized version of the covariance, and ranges between -1 and +1:
#
# \begin{align}
# \rho = \frac{\text{cov}(x_1,x_2)}{\sqrt{\sigma_1^2 \sigma_2^2}}.
# \end{align}
#
# For simplicity, we will assume that the mean of each variable has already been subtracted, so that $\mu_i=0$. The remaining parameters can be summarized in the covariance matrix, which for two dimensions has the following form:
#
# \begin{equation*}
# {\bf \Sigma} =
# \begin{pmatrix}
# \text{var}(x_1) & \text{cov}(x_1,x_2) \\
# \text{cov}(x_1,x_2) &\text{var}(x_2)
# \end{pmatrix}.
# \end{equation*}
#
# In general, $\bf \Sigma$ is a symmetric matrix with the variances $\text{var}(x_i) = \sigma_i^2$ on the diagonal, and the covariances on the off-diagonal. Later, we will see that the covariance matrix plays a key role in PCA.
#
#
# + [markdown] colab_type="text"
#
# ## Exercise 1: Draw samples from a distribution
#
# We have provided code to draw random samples from a zero-mean bivariate normal distribution. Throughout this tutorial, we'll imagine these samples represent the activity (firing rates) of two recorded neurons on different trials. Fill in the function below to calculate the covariance matrix given the desired variances and correlation coefficient. The covariance can be found by rearranging the equation above:
#
# \begin{align}
# \text{cov}(x_1,x_2) = \rho \sqrt{\sigma_1^2 \sigma_2^2}.
# \end{align}
#
# Use these functions to generate and plot data while varying the parameters. You should get a feel for how changing the correlation coefficient affects the geometry of the simulated data.
#
# **Steps**
# * Fill in the function `calculate_cov_matrix` to calculate the desired covariance.
# * Generate and plot the data for $\sigma_1^2 =1$, $\sigma_1^2 =1$, and $\rho = .8$. Try plotting the data for different values of the correlation coefficent: $\rho = -1, -.5, 0, .5, 1$.
# + colab={"base_uri": "https://localhost:8080/", "height": 514} colab_type="code" outputId="62dc3b47-bdec-445a-e8b8-2335f2793419"
help(plot_data)
help(get_data)
# + colab={} colab_type="code"
def calculate_cov_matrix(var_1, var_2, corr_coef):
"""
Calculates the covariance matrix based on the variances and correlation
coefficient.
Args:
var_1 (scalar) : variance of the first random variable
var_2 (scalar) : variance of the second random variable
corr_coef (scalar) : correlation coefficient
Returns:
(numpy array of floats) : covariance matrix
"""
#################################################
## TODO for students: calculate the covariance matrix
# Fill out function and remove
raise NotImplementedError("Student excercise: calculate the covariance matrix!")
#################################################
# Calculate the covariance from the variances and correlation
cov = ...
cov_matrix = np.array([[var_1, cov], [cov, var_2]])
return cov_matrix
###################################################################
## TO DO for students: generate and plot bivariate Gaussian data with variances of 1
## and a correlation coefficients of: 0.8
## repeat while varying the correlation coefficient from -1 to 1
###################################################################
np.random.seed(2020) # set random seed
variance_1 = 1
variance_2 = 1
corr_coef = 0.8
# Uncomment to test your code and plot
# cov_matrix = calculate_cov_matrix(variance_1, variance_2, corr_coef)
# X = get_data(cov_matrix)
# plot_data(X)
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" outputId="87407e56-4bd4-4cf0-d040-57381772daad"
# to_remove solution
def calculate_cov_matrix(var_1, var_2, corr_coef):
"""
Calculates the covariance matrix based on the variances and correlation
coefficient.
Args:
var_1 (scalar) : variance of the first random variable
var_2 (scalar) : variance of the second random variable
corr_coef (scalar) : correlation coefficient
Returns:
(numpy array of floats) : covariance matrix
"""
# Calculate the covariance from the variances and correlation
cov = corr_coef * np.sqrt(var_1 * var_2)
cov_matrix = np.array([[var_1, cov], [cov, var_2]])
return cov_matrix
np.random.seed(2020) # set random seed
variance_1 = 1
variance_2 = 1
corr_coef = 0.8
# Uncomment to test your code and plot
cov_matrix = calculate_cov_matrix(variance_1, variance_2, corr_coef)
X = get_data(cov_matrix)
with plt.xkcd():
plot_data(X)
# + [markdown] colab_type="text"
# ---
# # Section 2: Define a new orthonormal basis
#
# + cellView="form"
# @title Video 3: Orthonormal bases
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="PC1RZELnrIg", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# + [markdown] colab_type="text"
# Next, we will define a new orthonormal basis of vectors ${\bf u} = [u_1,u_2]$ and ${\bf w} = [w_1,w_2]$. As we learned in the video, two vectors are orthonormal if:
#
# 1. They are orthogonal (i.e., their dot product is zero):
# \begin{equation}
# {\bf u\cdot w} = u_1 w_1 + u_2 w_2 = 0
# \end{equation}
# 2. They have unit length:
# \begin{equation}
# ||{\bf u} || = ||{\bf w} || = 1
# \end{equation}
#
# In two dimensions, it is easy to make an arbitrary orthonormal basis. All we need is a random vector ${\bf u}$, which we have normalized. If we now define the second basis vector to be ${\bf w} = [-u_2,u_1]$, we can check that both conditions are satisfied:
# \begin{equation}
# {\bf u\cdot w} = - u_1 u_2 + u_2 u_1 = 0
# \end{equation}
# and
# \begin{equation}
# {|| {\bf w} ||} = \sqrt{(-u_2)^2 + u_1^2} = \sqrt{u_1^2 + u_2^2} = 1,
# \end{equation}
# where we used the fact that ${\bf u}$ is normalized. So, with an arbitrary input vector, we can define an orthonormal basis, which we will write in matrix by stacking the basis vectors horizontally:
#
# \begin{equation}
# {{\bf W} } =
# \begin{pmatrix}
# u_1 & w_1 \\
# u_2 & w_2
# \end{pmatrix}.
# \end{equation}
#
#
# + [markdown] colab_type="text"
# ## Exercise 2: Find an orthonormal basis
#
# In this exercise you will fill in the function below to define an orthonormal basis, given a single arbitrary 2-dimensional vector as an input.
#
# **Steps**
# * Modify the function `define_orthonormal_basis` to first normalize the first basis vector $\bf u$.
# * Then complete the function by finding a basis vector $\bf w$ that is orthogonal to $\bf u$.
# * Test the function using initial basis vector ${\bf u} = [3,1]$. Plot the resulting basis vectors on top of the data scatter plot using the function `plot_basis_vectors`. (For the data, use $\sigma_1^2 =1$, $\sigma_2^2 =1$, and $\rho = .8$).
# + colab={"base_uri": "https://localhost:8080/", "height": 257} colab_type="code" outputId="875de345-7eb6-4a82-a05f-eedb5926e91f"
help(plot_basis_vectors)
# + colab={} colab_type="code"
def define_orthonormal_basis(u):
"""
Calculates an orthonormal basis given an arbitrary vector u.
Args:
u (numpy array of floats) : arbitrary 2-dimensional vector used for new
basis
Returns:
(numpy array of floats) : new orthonormal basis
columns correspond to basis vectors
"""
#################################################
## TODO for students: calculate the orthonormal basis
# Fill out function and remove
raise NotImplementedError("Student excercise: implement the orthonormal basis function")
#################################################
# normalize vector u
u = ...
# calculate vector w that is orthogonal to w
w = ...
W = np.column_stack([u, w])
return W
np.random.seed(2020) # set random seed
variance_1 = 1
variance_2 = 1
corr_coef = 0.8
cov_matrix = calculate_cov_matrix(variance_1, variance_2, corr_coef)
X = get_data(cov_matrix)
u = np.array([3, 1])
# Uncomment and run below to plot the basis vectors
# W = define_orthonormal_basis(u)
# plot_basis_vectors(X, W)
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" outputId="692ac8a7-45b5-4a00-ad38-4522f4c0ecdd"
# to_remove solution
def define_orthonormal_basis(u):
"""
Calculates an orthonormal basis given an arbitrary vector u.
Args:
u (numpy array of floats) : arbitrary 2-dimensional vector used for new
basis
Returns:
(numpy array of floats) : new orthonormal basis
columns correspond to basis vectors
"""
# normalize vector u
u = u / np.sqrt(u[0] ** 2 + u[1] ** 2)
# calculate vector w that is orthogonal to w
w = np.array([-u[1], u[0]])
W = np.column_stack([u, w])
return W
np.random.seed(2020) # set random seed
variance_1 = 1
variance_2 = 1
corr_coef = 0.8
cov_matrix = calculate_cov_matrix(variance_1, variance_2, corr_coef)
X = get_data(cov_matrix)
u = np.array([3, 1])
# Uncomment and run below to plot the basis vectors
W = define_orthonormal_basis(u)
with plt.xkcd():
plot_basis_vectors(X, W)
# + [markdown] colab_type="text"
# ---
# # Section 3: Project data onto new basis
# + cellView="form"
# @title Video 4: Change of basis
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="Mj6BRQPKKUc", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# + [markdown] colab_type="text"
#
#
# Finally, we will express our data in the new basis that we have just found. Since $\bf W$ is orthonormal, we can project the data into our new basis using simple matrix multiplication :
#
# \begin{equation}
# {\bf Y = X W}.
# \end{equation}
#
# We will explore the geometry of the transformed data $\bf Y$ as we vary the choice of basis.
# + [markdown] colab_type="text"
# ## Exercise 3: Define an orthonormal basis
# In this exercise you will fill in the function below to define an orthonormal basis, given a single arbitrary vector as an input.
#
# **Steps**
# * Complete the function `change_of_basis` to project the data onto the new basis.
# * Plot the projected data using the function `plot_data_new_basis`.
# * What happens to the correlation coefficient in the new basis? Does it increase or decrease?
# * What happens to variance?
#
#
# + colab={} colab_type="code"
def change_of_basis(X, W):
"""
Projects data onto new basis W.
Args:
X (numpy array of floats) : Data matrix each column corresponding to a
different random variable
W (numpy array of floats) : new orthonormal basis columns correspond to
basis vectors
Returns:
(numpy array of floats) : Data matrix expressed in new basis
"""
#################################################
## TODO for students: project the data onto o new basis W
# Fill out function and remove
raise NotImplementedError("Student excercise: implement change of basis")
#################################################
# project data onto new basis described by W
Y = ...
return Y
# Unomment below to transform the data by projecting it into the new basis
# Y = change_of_basis(X, W)
# plot_data_new_basis(Y)
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" outputId="6ae7008b-0f5c-4d43-eb19-45787a995387"
# to_remove solution
def change_of_basis(X, W):
"""
Projects data onto new basis W.
Args:
X (numpy array of floats) : Data matrix each column corresponding to a
different random variable
W (numpy array of floats) : new orthonormal basis columns correspond to
basis vectors
Returns:
(numpy array of floats) : Data matrix expressed in new basis
"""
# project data onto new basis described by W
Y = np.matmul(X, W)
return Y
# Unomment below to transform the data by projecting it into the new basis
Y = change_of_basis(X, W)
with plt.xkcd():
plot_data_new_basis(Y)
# + [markdown] colab_type="text"
# ## Interactive Demo: Play with the basis vectors
# To see what happens to the correlation as we change the basis vectors, run the cell below. The parameter $\theta$ controls the angle of $\bf u$ in degrees. Use the slider to rotate the basis vectors.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 588, "referenced_widgets": ["103182f1916b4aab81d4444e690d15a3", "8770c3fad5af4ae98f6b19c1cd15f6b6", "f3ae30ce068f4e37823bc82cea26ca32", "724721c3b71842fa863c727fa8ab2d9f", "f231860ae93845129fc2280eecd09aeb", "80a7274133094fcab8370a06f03b0ab5", "8b23c192ec1c49468b0949104c242257"]} colab_type="code" outputId="657b6946-a7aa-4a4d-b57c-ee4eba1eac0c"
# @title
# @markdown Make sure you execute this cell to enable the widget!
def refresh(theta=0):
u = [1, np.tan(theta * np.pi / 180)]
W = define_orthonormal_basis(u)
Y = change_of_basis(X, W)
plot_basis_vectors(X, W)
plot_data_new_basis(Y)
_ = widgets.interact(refresh, theta=(0, 90, 5))
# + [markdown] colab_type="text"
# ## Questions
#
# * What happens to the projected data as you rotate the basis?
# * How does the correlation coefficient change? How does the variance of the projection onto each basis vector change?
# * Are you able to find a basis in which the projected data is **uncorrelated**?
# + [markdown] colab_type="text"
# ---
# # Summary
#
# - In this tutorial, we learned that multivariate data can be visualized as a cloud of points in a high-dimensional vector space. The geometry of this cloud is shaped by the covariance matrix.
#
# - Multivariate data can be represented in a new orthonormal basis using the dot product. These new basis vectors correspond to specific mixtures of the original variables - for example, in neuroscience, they could represent different ratios of activation across a population of neurons.
#
# - The projected data (after transforming into the new basis) will generally have a different geometry from the original data. In particular, taking basis vectors that are aligned with the spread of cloud of points decorrelates the data.
#
# * These concepts - covariance, projections, and orthonormal bases - are key for understanding PCA, which we be our focus in the next tutorial.
|
tutorials/W1D5_DimensionalityReduction/W1D5_Tutorial1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit (conda)
# language: python
# name: python3
# ---
# # Relatório de Análise VIII
# ## Identificando e Removendo Outliers
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
plt.rc('figure', figsize = (14, 6))
dados = pd.read_csv('../data/aluguel/aluguel_residencial.csv', sep = ';')
dados.boxplot(['Valor'])
dados[dados['Valor'] >= 500000]
valor = dados['Valor']
Q1 = valor.quantile(.25)
Q3 = valor.quantile(.75)
IIQ = Q3 - Q1
limite_inferior = Q1 - 1.5 * IIQ
limite_superior = Q3 + 1.5 * IIQ
selecao = (valor >= limite_inferior) & (valor <= limite_superior)
dados_new = dados[selecao]
dados_new.boxplot(['Valor'])
dados.hist(['Valor'])
dados_new.hist(['Valor'])
# ## Identificando e removendo Outliers (Continuação)
dados.boxplot(['Valor'], by = ['Tipo'])
grupo_tipo = dados.groupby('Tipo')['Valor']
type(grupo_tipo)
grupo_tipo.groups
q1 = grupo_tipo.quantile(.25)
q3 = grupo_tipo.quantile(.75)
iiq = q3 - q1
limite_inferior = q1 - 1.5 * iiq
limite_superior = q3 + 1.5 * iiq
q1
q3
iiq
limite_inferior
limite_superior
limite_superior['Casa']
dados_new = pd.DataFrame()
for tipo in grupo_tipo.groups.keys():
eh_tipo = dados['Tipo'] == tipo
eh_dentro_limite = (dados['Valor'] >= limite_inferior[tipo]) & (dados['Valor'] <= limite_superior[tipo])
selecao = eh_tipo & eh_dentro_limite
dados_selecao = dados[selecao]
dados_new = pd.concat([dados_new, dados_selecao])
dados_new.boxplot(['Valor'], by = ['Tipo'])
dados_new.to_csv('../data/aluguel/aluguel_residencial_sem_outliers.csv', sep =';', index = False)
# ## Exercicios
import pandas as pd
dados = pd.read_csv('../data/aluguel/aluguel_amostra.csv', sep = ';')
dados.head(10)
valor = dados['Valor m2']
q1 = valor.quantile(.25)
q3 = valor.quantile(.75)
iiq = q3 - q1
limite_inferior = q1 - 1.5 * iiq
limite_superior = q3 + 1.5 * iiq
print(f'1° quartil: {q1}')
print(f'3° quartil: {q3}')
print(f'Intervalo interquartil: {iiq:.2f}')
print(f'Limite inferior: {limite_inferior:.2f}')
print(f'Limite superior: {limite_superior:.2f}')
|
notebook/identificando-e-removendo-outliers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
from scipy.special import expit, logit
x = np.linspace(0, 1, 121)
x1 = x-.5
x1 *= 50
y = expit(x1)
plt.plot(x, y)
plt.grid()
plt.xlabel('x')
plt.title('expit(x)')
plt.show()
# +
from scipy.stats import norm
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1)
x = np.linspace(norm.ppf(0.01),
norm.ppf(0.99), 100)
rv = norm()
ax.plot(x, rv.pdf(x), 'k-', lw=5, label='frozen pdf')
fig.patch.set_visible(False)
ax.axis('off')
plt.tight_layout()
plt.savefig('normal_dist_1x.svg')
# -
fig, ax = plt.subplots(1, 1)
rv = norm()
ax.plot(x, rv.pdf(x*2), 'k-', lw=5, label='frozen pdf')
fig.patch.set_visible(False)
ax.axis('off')
plt.tight_layout()
plt.savefig('normal_dist_2x.svg')
|
analysis/mf_grc_analysis/test_sigmoid.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from utilsADCN import dataLoader
from ADCNbasic import ADCN
from ADCNmainloop import ADCNmain
from model import simpleMPL
import numpy as np
import pdb
import torch
import random
from torchvision import datasets, transforms
# random seed control
np.random.seed(0)
torch.manual_seed(0)
random.seed(0)
dataStream = dataLoader('./data/creditcarddefault.mat', maxMinNorm = True)
dataStream.labeledData.shape
device = torch.device('cuda:0')
nHidNodeExtractor = dataStream.nInput*4
nExtractedFeature = dataStream.nInput*4
nFeaturClustering = dataStream.nInput*2
allMetrics = []
n_trials = 5
for i_trial in range(0, n_trials):
print('Trial: ', i_trial)
ADCNnet = ADCN(dataStream.nOutput, nInput = nExtractedFeature, nHiddenNode = nFeaturClustering)
ADCNnet.ADCNcnn = simpleMPL(dataStream.nInput, nNodes = nHidNodeExtractor, nOutput = nExtractedFeature)
ADCNnet.desiredLabels = [0,1]
ADCNnet, performanceHistory, allPerformance = ADCNmain(ADCNnet, dataStream, device = device)
allMetrics.append(allPerformance)
# +
# all results
# 0: accuracy
# 1: ARI
# 2: NMI
# 3: f1_score
# 4: precision_score
# 5: recall_score
# 6: training_time
# 7: testingTime
# 8: nHiddenLayer
# 9: nHiddenNode
# 10: nCluster
meanResults = np.round_(np.mean(allMetrics,0), decimals=2)
stdResults = np.round_(np.std(allMetrics,0), decimals=2)
print('\n')
print('========== Performance SEA ==========')
print('Preq Accuracy: ', meanResults[0].item(), '(+/-)',stdResults[0].item())
print('ARI: ', meanResults[1].item(), '(+/-)',stdResults[1].item())
print('NMI: ', meanResults[2].item(), '(+/-)',stdResults[2].item())
print('F1 score: ', meanResults[3].item(), '(+/-)',stdResults[3].item())
print('Precision: ', meanResults[4].item(), '(+/-)',stdResults[4].item())
print('Recall: ', meanResults[5].item(), '(+/-)',stdResults[5].item())
print('Training time: ', meanResults[6].item(), '(+/-)',stdResults[6].item())
print('Testing time: ', meanResults[7].item(), '(+/-)',stdResults[7].item())
print('\n')
print('========== Network ==========')
print('Number of hidden layers: ', meanResults[8].item(), '(+/-)',stdResults[8].item())
print('Number of features: ', meanResults[9].item(), '(+/-)',stdResults[9].item())
print('Number of clusters: ', meanResults[10].item(), '(+/-)',stdResults[10].item())
# -
|
ADCN-creditcard.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model with Metrics
#
# ## Dependencies
#
# ```pip install seldon-core```
#
# ## Summary of Custom Metrics
#
# Example testing a model with custom metrics.
#
# Metrics can be
#
# * A ```COUNTER``` : the returned value will increment the current value
# * A ```GAUGE``` : the returned value will overwrite the current value
# * A ```TIMER``` : a number of millisecs. Prometheus SUM and COUNT metrics will be created.
#
# You need to provide a list of dictionaries each with the following:
#
# * a ```type``` : COUNTER, GAUGE, or TIMER
# * a ```key``` : a user defined key
# * a ```value``` : a float value
#
# See example code below:
#
# !pygmentize ModelWithMetrics.py
# ## REST
# !s2i build -E environment_rest . seldonio/seldon-core-s2i-python3:0.15 model-with-metrics-rest:0.1
# !docker run --name "model-with-metrics" -d --rm -p 5000:5000 model-with-metrics-rest:0.1
# ### Test predict
# !seldon-core-tester contract.json 0.0.0.0 5000 -p
# !docker rm model-with-metrics --force
# ## gRPC
# !s2i build -E environment_grpc . seldonio/seldon-core-s2i-python3:0.15 model-with-metrics-grpc:0.1
# !docker run --name "model-with-metrics" -d --rm -p 5000:5000 model-with-metrics-grpc:0.1
# ### Test predict
# !seldon-core-tester contract.json 0.0.0.0 5000 -p --grpc
# !docker rm model-with-metrics --force
# ## Test using Minikube
#
# **Due to a [minikube/s2i issue](https://github.com/SeldonIO/seldon-core/issues/253) you will need [s2i >= 1.1.13](https://github.com/openshift/source-to-image/releases/tag/v1.1.13)**
# !minikube start --memory 4096
# ## Setup Seldon Core
#
# Use the setup notebook to [Setup Cluster](../../seldon_core_setup.ipynb#Setup-Cluster) with [Ambassador Ingress](../../seldon_core_setup.ipynb#Ambassador) and [Install Seldon Core](../../seldon_core_setup.ipynb#Install-Seldon-Core). Instructions [also online](./seldon_core_setup.html).
# * Port forward the dashboard when running
# ```
# kubectl port-forward $(kubectl get pods -n default -l app=grafana-prom-server -o jsonpath='{.items[0].metadata.name}') -n default 3000:3000
# ```
# * Visit http://localhost:3000/dashboard/db/prediction-analytics?refresh=5s&orgId=1 and login using "admin" and the password you set above when launching with helm.
# ## REST
# !eval $(minikube docker-env) && s2i build -E environment_rest . seldonio/seldon-core-s2i-python3:0.15 model-with-metrics-rest:0.1
# !kubectl create -f deployment-rest.json
# !kubectl rollout status deploy/mymodel-mymodel-b79af31
# ### Test predict
# !seldon-core-api-tester contract.json `minikube ip` `kubectl get svc ambassador -o jsonpath='{.spec.ports[0].nodePort}'` \
# mymodel --namespace seldon -p
# !kubectl delete -f deployment-rest.json
# ## gRPC
# !eval $(minikube docker-env) && s2i build -E environment_grpc . seldonio/seldon-core-s2i-python3:0.15 model-with-metrics-grpc:0.1
# !kubectl create -f deployment-grpc.json
# !kubectl rollout status deploy/mymodel-mymodel-5818788
# ### Validate on Grafana
#
# To check the metrics have appeared on Prometheus and are available in Grafana you could create a new graph in a dashboard and use the query:
#
# ```
# mycounter_total
# ```
#
# ### Test predict
# !seldon-core-api-tester contract.json `minikube ip` `kubectl get svc ambassador -o jsonpath='{.spec.ports[0].nodePort}'` \
# mymodel --namespace seldon -p --grpc
# !kubectl delete -f deployment-grpc.json
# !minikube delete
|
examples/models/template_model_with_metrics/modelWithMetrics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The chaos game and the Sierpinski triangle
#
# Source: https://www.johndcook.com/blog/2017/07/08/the-chaos-game-and-the-sierpinski-triangle/
#
# TODO: https://twitter.com/franssoa/status/1102223543897636865
from scipy import sqrt, zeros
import matplotlib.pyplot as plt
from random import random, randint
def midpoint(p, q):
return (0.5*(p[0] + q[0]), 0.5*(p[1] + q[1]))
# +
# Three corners of an equilateral triangle
corner = [(0, 0), (0.5, sqrt(3)/2), (1, 0)]
N = 1000
x = zeros(N)
y = zeros(N)
x[0] = random()
y[0] = random()
for i in range(1, N):
k = randint(0, 2) # random triangle vertex
x[i], y[i] = midpoint( corner[k], (x[i-1], y[i-1]) )
# -
plt.scatter(x, y)
plt.show()
|
nb_sci_maths/maths_chaos_sierpinski_triangle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import logging
logging.basicConfig(level=logging.ERROR)
# +
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
X, y = load_iris(return_X_y=True, as_frame=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
X[:3]
# -
y[:3]
# +
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
X, y = load_iris(return_X_y=True, as_frame=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
from hypergbm.estimators import XGBoostEstimator
from hypergbm.pipeline import Pipeline
from hypergbm.sklearn.transformers import FeatureGenerationTransformer
from hypernets.core.ops import ModuleChoice, HyperInput
from hypernets.core.search_space import HyperSpace
from hypernets.tabular.column_selector import column_exclude_datetime
def search_space(task=None):
space = HyperSpace()
with space.as_default():
input = HyperInput(name='input1')
feature_gen = FeatureGenerationTransformer(task=task,
trans_primitives=["add_numeric", "subtract_numeric", "divide_numeric", "multiply_numeric"]) # Add feature generation to search space
full_pipeline = Pipeline([feature_gen], name=f'feature_gen_and_preprocess', columns=column_exclude_datetime)(input)
xgb_est = XGBoostEstimator(fit_kwargs={})
ModuleChoice([xgb_est], name='estimator_options')(full_pipeline)
space.set_inputs(input)
return space
# +
from hypergbm import HyperGBM
from hypernets.searchers.evolution_searcher import EvolutionSearcher
rs = EvolutionSearcher(search_space, 200, 100, optimize_direction='max')
hk = HyperGBM(rs, task='multiclass', reward_metric='accuracy', callbacks=[])
hk.search(X_train, y_train, X_eval=X_test, y_eval=y_test)
# +
estimator = hk.load_estimator(hk.get_best_trial().model_file)
y_pred = estimator.predict(X_test)
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_pred)
|
hypergbm/examples/misc/feature_generation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # REINFORCE
#
# ---
#
# In this notebook, we will train REINFORCE with OpenAI Gym's Cartpole environment.
# ### 1. Import the Necessary Packages
# +
import gym
gym.logger.set_level(40) # suppress warnings (please remove if gives error)
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
# %matplotlib inline
import torch
torch.manual_seed(0) # set random seed
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.distributions import Categorical
# -
# ### 2. Define the Architecture of the Policy
# +
env = gym.make('Acrobot-v1')
env.seed(0)
print('observation space:', env.observation_space)
print('action space:', env.action_space)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
class Policy(nn.Module):
def __init__(self, s_size=4, h_size=16, a_size=2):
super(Policy, self).__init__()
self.fc1 = nn.Linear(s_size, h_size).to(device)
self.fc2 = nn.Linear(h_size, h_size).to(device)
self.fc3 = nn.Linear(h_size, a_size).to(device)
def forward(self, x):
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return F.softmax(x, dim=-1)
def act(self, state):
state = torch.from_numpy(state).float().unsqueeze(0).to(device)
probs = self.forward(state).cpu()
m = Categorical(probs)
action = m.sample()
return action.item(), m.log_prob(action)
# -
state = env.reset()
state = torch.from_numpy(state).float().to(device)
policy = Policy()
print("state", state)
prob = policy.forward(state)
print("prob",prob)
g = Categorical(prob)
print("categorical ",g)
print(g.sample())
a = [x for x in range(1,10)]
b = [x for x in range(20,30)]
a = [torch.from_numpy(np.array(x)) for x in a]
a = torch.from_numpy(np.array(a))
# ### 3. Train the Agent with REINFORCE
# +
policy = Policy(6,16,3).to(device)
optimizer = optim.Adam(policy.parameters(), lr=1e-2)
def reinforce(n_episodes=4000, max_t=1000, gamma=1.0, print_every=100):
scores_deque = deque(maxlen=100)
scores = []
for i_episode in range(1, n_episodes+1):
saved_log_probs = []
rewards = []
state = env.reset()
for t in range(max_t):
action, log_prob = policy.act(state)
saved_log_probs.append(log_prob)
state, reward, done, _ = env.step(action)
rewards.append(reward)
if done:
break
scores_deque.append(sum(rewards))
scores.append(sum(rewards))
discounts = [gamma**i for i in range(len(rewards)+1)]
R = sum([a*b for a,b in zip(discounts, rewards)])
policy_loss = []
for log_prob in saved_log_probs:
policy_loss.append(-log_prob * R)
policy_loss = torch.cat(policy_loss).sum()
optimizer.zero_grad()
policy_loss.backward()
optimizer.step()
if i_episode % print_every == 0:
print('Episode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_deque)))
if np.mean(scores_deque)>=195.0:
print('Environment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_deque)))
break
return scores
scores = reinforce()
# -
# ### 4. Plot the Scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# ### 5. Watch a Smart Agent!
# +
env = gym.make('CartPole-v0')
state = env.reset()
for t in range(1000):
action, logprob = policy.act(state)
print('\r action {} with log prob {}'.format(action, logprob), end = ' ')
env.render()
state, reward, done, _ = env.step(action)
if done:
break
env.close()
# -
|
reinforce/REINFORCE-Copy1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# # Greybox Fuzzing with Grammars
#
# <!--
# Previously, we have learned about [mutational fuzzing](GreyboxFuzzer.ipynb), which generates new inputs by mutating seed inputs. Most mutational fuzzers represent inputs as a sequence of bytes and apply byte-level mutations to this byte sequence. Such byte-level mutations work great for compact file formats with a small number of structural constraints. However, most file formats impose a high-level structure on these byte sequences.
#
# Common components of a regular file are file header, data chunks, checksums, data fields, and meta data. Only if this file structure is correctly reflected will the file be accepted by the parser. Otherwise, the file is quickly rejected before reaching interesting parts in the program. It is not easy to generate valid files by [random fuzzing](Fuzzer.ipynb). For instance, only a tiniest proportion of random strings are valid PDF files or valid JPEG image files.
# -->
#
# <!--
# Maybe we can start with a valid file and generate new valid files by small mutations applied to the original file? Indeed, this is the main insight of ([blackbox](MutationFuzzer.ipynb) and [greybox](GreyboxFuzzer.ipynb)) mutational fuzzing. However, many file formats are so complex that even small modifications lead to invalid inputs that are quickly rejected by the parser.
# -->
#
# In this chapter, we introduce two important extensions to our syntactic fuzzing techniques:
#
# 1. We show how to combine [parsing](Parser.ipynb) and [fuzzing](GrammarFuzzer.ipynb) with grammars. This allows to _mutate_ existing inputs while preserving syntactical correctness, and to _reuse_ fragments from existing inputs while generating new ones. The combination of parsing and fuzzing, as demonstrated in this chapter, has been highly successful in practice: The _LangFuzz_ fuzzer for JavaScript has found more than 2,600 bugs in JavaScript interpreters this way.
#
# 2. In the previous chapters, we have used grammars in a _black-box_ manner – that is, we have used them to generate inputs regardless of the program being tested. In this chapter, we introduce mutational _greybox fuzzing with grammars_: Techniques that make use of _feedback from the program under test_ to guide test generations towards specific goals. As in [lexical greybox fuzzing](GreyboxFuzzer.ipynb), this feedback is mostly _coverage_, allowing us to direct grammar-based testing towards uncovered code parts.
#
#
# <!--
# In this chapter, we encode file formats as [grammars](Grammars.ipynb) and make the mutational fuzzer input-structure-aware. We investigate opportunities to inform the fuzzer about the validity of the generated inputs. Specifically, we explore dictionaries, grammars, structural mutators, and validity-based power schedules
# -->
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# **Prerequisites**
#
# * We build on several concepts from [the chapter on greybox fuzzing (without grammars)](GreyboxFuzzer.ipynb).
# * As the title suggests, you should know how to fuzz with grammars [from the chapter on grammars](Grammars.ipynb).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
# First, we [recall](GreyboxFuzzer.ipynb#Ingredients-for-Greybox-Fuzzing) a few basic ingredients for mutational fuzzers.
# * **Seed**. A _seed_ is an input that is used by the fuzzer to generate new inputs by applying a sequence of mutations.
# * **Mutator**. A _mutator_ implements a set of mutation operators that applied to an input produce a slightly modified input.
# * **PowerSchedule**. A _power schedule_ assigns _energy_ to a seed. A seed with higher energy is fuzzed more often throughout the fuzzing campaign.
# * **MutationFuzzer**. Our _mutational blackbox fuzzer_ generates inputs by mutating seeds in an initial population of inputs.
# * **GreyboxFuzzer**. Our _greybox fuzzer_ dynamically adds inputs to the population of seeds that increased coverage.
# * **FunctionCoverageRunner**. Our _function coverage runner_ collects coverage information for the execution of a given Python function.
#
# Let's try to get a feeling for these concepts.
# + slideshow={"slide_type": "skip"}
import bookutils
# + slideshow={"slide_type": "skip"}
from GreyboxFuzzer import Mutator, Seed, PowerSchedule, MutationFuzzer, GreyboxFuzzer
from MutationFuzzer import FunctionCoverageRunner
# + [markdown] slideshow={"slide_type": "subslide"}
# The following command applies a mutation to the input "Hello World".
# + slideshow={"slide_type": "fragment"}
Mutator().mutate("Hello World")
# + [markdown] slideshow={"slide_type": "fragment"}
# The default power schedule assigns energy uniformly across all seeds. Let's check whether this works.
#
# We choose 10k times from a population of three seeds. As we see in the `hits` counter, each seed is chosen about a third of the time.
# + slideshow={"slide_type": "subslide"}
population = [Seed("A"), Seed("B"), Seed("C")]
schedule = PowerSchedule()
hits = {
"A" : 0,
"B" : 0,
"C" : 0
}
for i in range(10000):
seed = schedule.choose(population)
hits[seed.data] += 1
hits
# + [markdown] slideshow={"slide_type": "subslide"}
# Before explaining the function coverage runner, lets import Python's HTML parser as example...
# + slideshow={"slide_type": "skip"}
from html.parser import HTMLParser
# + [markdown] slideshow={"slide_type": "fragment"}
# ... and create a _wrapper function_ that passes each input into a new parser object.
# + slideshow={"slide_type": "fragment"}
def my_parser(inp):
parser = HTMLParser()
parser.feed(inp)
# + [markdown] slideshow={"slide_type": "fragment"}
# The `FunctionCoverageRunner` constructor takes a Python `function` to execute. The function `run()` takes an input, passes it on to the Python `function`, and collects the coverage information for this execution. The function `coverage()` returns a list of tuples `(function name, line number)` for each statement that has been covered in the Python `function`.
# + slideshow={"slide_type": "subslide"}
runner = FunctionCoverageRunner(my_parser)
runner.run("Hello World")
cov = runner.coverage()
list(cov)[:5] # Print 5 statements covered in HTMLParser
# + [markdown] slideshow={"slide_type": "fragment"}
# Our greybox fuzzer takes a seed population, mutator, and power schedule. Let's generate 5000 fuzz inputs starting with an "empty" seed corpus.
# + slideshow={"slide_type": "skip"}
import time
import random
# + slideshow={"slide_type": "subslide"}
n = 5000
seed_input = " " # empty seed
runner = FunctionCoverageRunner(my_parser)
fuzzer = GreyboxFuzzer([seed_input], Mutator(), PowerSchedule())
start = time.time()
fuzzer.runs(runner, trials=n)
end = time.time()
"It took the fuzzer %0.2f seconds to generate and execute %d inputs." % (end - start, n)
# + slideshow={"slide_type": "fragment"}
"During this fuzzing campaign, we covered %d statements." % len(runner.coverage())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Building a Keyword Dictionary
#
# To fuzz our HTML parser, it may be useful to inform a mutational fuzzer about important keywords in the input – that is, important HTML keywords. To this end, we extend our mutator to consider keywords from a _dictionary_.
# + slideshow={"slide_type": "subslide"}
class DictMutator(Mutator):
def __init__(self, dictionary):
super().__init__()
self.dictionary = dictionary
self.mutators.append(self.insert_from_dictionary)
def insert_from_dictionary(self,s):
"""Returns s with a keyword from the dictionary inserted"""
pos = random.randint(0, len(s))
random_keyword = random.choice(self.dictionary)
return s[:pos] + random_keyword + s[pos:]
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's try to add a few HTML tags and attributes and see whether the coverage with `DictMutator` increases.
# + slideshow={"slide_type": "subslide"}
runner = FunctionCoverageRunner(my_parser)
dict_mutator = DictMutator(["<a>","</a>","<a/>", "='a'"])
dict_fuzzer = GreyboxFuzzer([seed_input], dict_mutator, PowerSchedule())
start = time.time()
dict_fuzzer.runs(runner, trials = n)
end = time.time()
"It took the fuzzer %0.2f seconds to generate and execute %d inputs." % (end - start, n)
# + [markdown] slideshow={"slide_type": "fragment"}
# Clearly, it takes longer. In our experience, this means more code is covered:
# + slideshow={"slide_type": "fragment"}
"During this fuzzing campaign, we covered %d statements." % len(runner.coverage())
# + [markdown] slideshow={"slide_type": "fragment"}
# How do the fuzzers compare in terms of coverage over time?
# + slideshow={"slide_type": "skip"}
from Coverage import population_coverage
# + slideshow={"slide_type": "skip"}
import matplotlib.pyplot as plt
# + slideshow={"slide_type": "subslide"}
_, dict_cov = population_coverage(dict_fuzzer.inputs, my_parser)
_, fuzz_cov = population_coverage(fuzzer.inputs, my_parser)
line_dict, = plt.plot(dict_cov, label="With Dictionary")
line_fuzz, = plt.plot(fuzz_cov, label="Without Dictionary")
plt.legend(handles=[line_dict, line_fuzz])
plt.xlim(0,n)
plt.title('Coverage over time')
plt.xlabel('# of inputs')
plt.ylabel('lines covered');
# + [markdown] slideshow={"slide_type": "subslide"}
# <!-- \todo{Andreas: Section on mining keywords using parser-directed fuzzing or AUTOGRAM?} -->
#
# ***Summary.*** Informing the fuzzer about important keywords already goes a long way towards achieving lots of coverage quickly.
#
# ***Try it.*** Open this chapter as Jupyter notebook and add other HTML-related keywords to the dictionary in order to see whether the difference in coverage actually increases (given the same budget of 5k generated test inputs).
#
# ***Read up.*** <NAME>, author of AFL, wrote several great blog posts on [making up grammars with a dictionary in hand](https://lcamtuf.blogspot.com/2015/01/afl-fuzz-making-up-grammar-with.html) and [pulling JPEGs out of thin air](https://lcamtuf.blogspot.com/2014/11/pulling-jpegs-out-of-thin-air.html)!
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=false
# ## Fuzzing with Input Fragments
#
# While dictionaries are helpful to inject important keywords into seed inputs, they do not allow to maintain the structural integrity of the generated inputs. Instead, we need to make the fuzzer aware of the _input structure_. We can do this using [grammars](Grammars.ipynb). Our first approach
#
# 1. [parses](Parser.ipynb) the seed inputs,
# 2. disassembles them into input fragments, and
# 3. generates new files by reassembling these fragments according to the rules of the grammar.
#
# This combination of _parsing_ and _fuzzing_ can be very powerful, as we will see in an instant
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Parsing and Recombining JavaScript, or How to Make 50,000 USD in Four Weeks
#
# In "Fuzzing with Code Fragments" \cite{Holler2012}, Holler, Herzig, and Zeller apply these steps to fuzz a JavaScript interpreter. They use a JavaScript grammar to
#
# 1. _parse_ (valid) JavaScript inputs into parse trees,
# 2. _disassemble_ them into fragments (subtrees),
# 3. _recombine_ these fragments into valid JavaScript programs again, and
# 4. _feed_ these programs into a JavaScript interpreter for execution.
# + [markdown] slideshow={"slide_type": "subslide"}
# As in most fuzzing scenarios, the aim is to cause the JavaScript interpreter to crash. Here is an example of LangFuzz-generated JavaScript code (from \cite{Holler2012}) that caused a crash in the Mozilla JavaScript interpreter:
#
# ```javascript
# var haystack = "foo";
# var re_text = "^foo";
# haystack += "x";
# re_text += "(x)";
# var re = new RegExp(re_text);
# re.test(haystack);
# RegExp.input = Number();
# print(RegExp.$1);
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# From a crash of the JavaScript interpreter, it is frequently possible to construct an *exploit* that will not only crash the interpreter, but instead have it execute code under the attacker's control. Therefore, such crashes are serious flaws, which is why you get a bug bounty if you report them.
# + [markdown] slideshow={"slide_type": "subslide"}
# In the first four weeks of running his _LangFuzz_ tool, <NAME>, first author of that paper, netted _more than USD 50,000 in bug bounties_. To date, LangFuzz has found more than 2,600 bugs in the JavaScript browsers of Mozilla Firefox, Google Chrome, and Microsoft Edge. If you use any of these browsers (say, on your Android phone), the combination of parsing and fuzzing has contributed significantly in making your browsing session secure.
#
# (Note that these are the same Holler and Zeller who are co-authors of this book. If you ever wondered why we devote a couple of chapters on grammar-based fuzzing, that's because we have had some great experience with it.)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Parsing and Recombining HTML
#
# In this book, let us stay with HTML input for a while. To generate valid HTML inputs for our Python `HTMLParser`, we should first define a simple grammar. It allows to define HTML tags with attributes. Our context-free grammar does not require that opening and closing tags must match. However, we will see that such context-sensitive features can be maintained in the derived input fragments, and thus in the generated inputs.
# + slideshow={"slide_type": "skip"}
import string
# + slideshow={"slide_type": "skip"}
from Grammars import is_valid_grammar, srange
# + slideshow={"slide_type": "subslide"}
XML_TOKENS = {"<id>","<text>"}
XML_GRAMMAR = {
"<start>": ["<xml-tree>"],
"<xml-tree>": ["<text>",
"<xml-open-tag><xml-tree><xml-close-tag>",
"<xml-openclose-tag>",
"<xml-tree><xml-tree>"],
"<xml-open-tag>": ["<<id>>", "<<id> <xml-attribute>>"],
"<xml-openclose-tag>": ["<<id>/>", "<<id> <xml-attribute>/>"],
"<xml-close-tag>": ["</<id>>"],
"<xml-attribute>" : ["<id>=<id>", "<xml-attribute> <xml-attribute>"],
"<id>": ["<letter>", "<id><letter>"],
"<text>" : ["<text><letter_space>","<letter_space>"],
"<letter>": srange(string.ascii_letters + string.digits +"\""+"'"+"."),
"<letter_space>": srange(string.ascii_letters + string.digits +"\""+"'"+" "+"\t"),
}
# + slideshow={"slide_type": "subslide"}
assert is_valid_grammar(XML_GRAMMAR)
# + [markdown] slideshow={"slide_type": "fragment"}
# In order to parse an input into a derivation tree, we use the [Earley parser](Parser.ipynb#Parsing-Context-Free-Grammars).
# + slideshow={"slide_type": "skip"}
from Parser import EarleyParser
from GrammarFuzzer import display_tree
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's run the parser on a simple HTML input and display all possible parse trees. A *parse tree* represents the input structure according to the given grammar.
# + slideshow={"slide_type": "fragment"}
parser = EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS)
for tree in parser.parse("<html>Text</html>"):
display_tree(tree)
# + [markdown] slideshow={"slide_type": "fragment"}
# As we can see, the input starts with an opening tag, contains some text, and ends with a closing tag. Excellent. This is a structure that we can work with.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Building the Fragment Pool
# We are now ready to implement our first input-structure-aware mutator. Let's initialize the mutator with the dictionary `fragments` representing the empty fragment pool. It contains a key for each symbol in the grammar (and the empty set as value).
# + slideshow={"slide_type": "fragment"}
class FragmentMutator(Mutator):
def __init__(self, parser):
"""Initialize empty fragment pool and add parser"""
self.parser = parser
self.fragments = {k: [] for k in self.parser.cgrammar}
super().__init__()
# + [markdown] slideshow={"slide_type": "subslide"}
# The `FragmentMutator` adds fragments recursively. A *fragment* is a subtree in the parse tree and consists of the symbol of the current node and child nodes (i.e., descendant fragments). We can exclude fragments starting with symbols that are tokens, terminals, or not part of the grammar.
# + slideshow={"slide_type": "skip"}
from Parser import terminals
# + slideshow={"slide_type": "subslide"}
class FragmentMutator(FragmentMutator):
def add_fragment(self, fragment):
"""Recursively adds fragments to the fragment pool"""
(symbol, children) = fragment
if not self.is_excluded(symbol):
self.fragments[symbol].append(fragment)
for subfragment in children:
self.add_fragment(subfragment)
def is_excluded(self, symbol):
"""Returns true if a fragment starting with a specific
symbol and all its decendents can be excluded"""
return ((not symbol in self.parser.grammar()) or
symbol in self.parser.tokens or
symbol in terminals(self.parser.grammar()))
# + [markdown] slideshow={"slide_type": "subslide"}
# Parsing can take a long time, particularly if there is too much ambiguity during the parsing. In order to maintain the efficiency of mutational fuzzing, we will limit the parsing time to 200ms.
# + slideshow={"slide_type": "skip"}
import signal
# + slideshow={"slide_type": "fragment"}
class Timeout(Exception): pass
def timeout(signum, frame):
raise Timeout()
# Register timeout() as handler for signal 'SIGALRM'"
signal.signal(signal.SIGALRM, timeout);
# + [markdown] slideshow={"slide_type": "subslide"}
# The function `add_to_fragment_pool()` parses a seed (no longer than 200ms) and adds all its fragments to the fragment pool. If the parsing of the `seed` was successful, the attribute `seed.has_structure` is set to `True`. Otherwise, it is set to `False`.
#
# <!-- \todo{Convert this to `ExpectTimeout` (or make ExpectTimeout more efficient)} -->
# + slideshow={"slide_type": "subslide"}
class FragmentMutator(FragmentMutator):
def add_to_fragment_pool(self, seed):
"""Adds all fragments of a seed to the fragment pool"""
try: # only allow quick parsing of 200ms max
signal.setitimer(signal.ITIMER_REAL, 0.2)
seed.structure = next(self.parser.parse(seed.data))
signal.setitimer(signal.ITIMER_REAL, 0)
self.add_fragment(seed.structure)
seed.has_structure = True
except (SyntaxError, Timeout):
seed.has_structure = False
signal.setitimer(signal.ITIMER_REAL, 0)
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's see how `FragmentMutator` fills the fragment pool for a simple HTML seed input. We initialize mutator with the `EarleyParser` which itself is initialized with our `XML_GRAMMAR`.
# + slideshow={"slide_type": "skip"}
from GrammarFuzzer import tree_to_string
# + slideshow={"slide_type": "subslide"}
valid_seed = Seed("<html><header><title>Hello</title></header><body>World<br/></body></html>")
fragment_mutator = FragmentMutator(EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS))
fragment_mutator.add_to_fragment_pool(valid_seed)
for key in fragment_mutator.fragments:
print(key)
for f in fragment_mutator.fragments[key]:
print("|-%s" % tree_to_string(f))
# + [markdown] slideshow={"slide_type": "subslide"}
# For many symbols in the grammar, we have collected a number of fragments. There are several open and closing tags and several interesting fragments starting with the `xml-tree` symbol.
#
# ***Summary***. For each interesting symbol in the grammar, the `FragmentMutator` has a set of fragments. These fragments are extracted by first parsing the inputs to be mutated.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Fragment-Based Mutation
#
# We can use the fragments in the fragment pool to generate new inputs. Every seed that is being mutated is disassembled into fragments, and memoized – i.e., disassembled only the first time around.
# + slideshow={"slide_type": "subslide"}
class FragmentMutator(FragmentMutator):
def __init__(self, parser):
"""Initialize mutators"""
super().__init__(parser)
self.seen_seeds = []
def mutate(self, seed):
"""Implement structure-aware mutation. Memoize seeds."""
if not seed in self.seen_seeds:
self.seen_seeds.append(seed)
self.add_to_fragment_pool(seed)
return super().mutate(seed)
# + [markdown] slideshow={"slide_type": "subslide"}
# Our first structural mutation operator is `swap_fragments()`, which choses a random fragment in the given seed and substitutes it with a random fragment from the pool. We make sure that both fragments start with the same symbol. For instance, we may swap a closing tag in the seed HTML by another closing tag from the fragment pool.
#
# In order to choose a random fragment, the mutator counts all fragments (`n_count`) below the root fragment associated with the start-symbol.
# + slideshow={"slide_type": "subslide"}
class FragmentMutator(FragmentMutator):
def count_nodes(self, fragment):
"""Returns the number of nodes in the fragment"""
symbol, children = fragment
if self.is_excluded(symbol):
return 0
return 1 + sum(map(self.count_nodes, children))
# + [markdown] slideshow={"slide_type": "fragment"}
# In order to swap the chosen fragment – identified using the "global" variable `self.to_swap` – the seed's parse tree is traversed recursively.
# + slideshow={"slide_type": "subslide"}
class FragmentMutator(FragmentMutator):
def recursive_swap(self, fragment):
"""Recursively finds the fragment to swap."""
symbol, children = fragment
if self.is_excluded(symbol):
return symbol, children
self.to_swap -= 1
if self.to_swap == 0:
return random.choice(list(self.fragments[symbol]))
return symbol, list(map(self.recursive_swap, children))
# + [markdown] slideshow={"slide_type": "fragment"}
# Our structural mutator chooses a random number between 2 (i.e., excluding the `start` symbol) and the total number of fragments (`n_count`) and uses the recursive swapping to generate the new fragment. The new fragment is serialized as string and returned as new seed.
# + slideshow={"slide_type": "subslide"}
class FragmentMutator(FragmentMutator):
def __init__(self, parser):
super().__init__(parser)
self.mutators = [self.swap_fragment]
def swap_fragment(self, seed):
"""Substitutes a random fragment with another with the same symbol"""
if seed.has_structure:
n_nodes = self.count_nodes(seed.structure)
self.to_swap = random.randint(2, n_nodes)
new_structure = self.recursive_swap(seed.structure)
new_seed = Seed(tree_to_string(new_structure))
new_seed.has_structure = True
new_seed.structure = new_structure
return new_seed
return seed
# + slideshow={"slide_type": "subslide"}
valid_seed = Seed("<html><header><title>Hello</title></header><body>World<br/></body></html>")
lf_mutator = FragmentMutator(parser)
print(valid_seed)
lf_mutator.mutate(valid_seed)
# + [markdown] slideshow={"slide_type": "fragment"}
# As we can see, one fragment has been substituted by another.
#
# We can use a similar recursive traversal to *remove* a random fragment.
# + slideshow={"slide_type": "subslide"}
class FragmentMutator(FragmentMutator):
def recursive_delete(self, fragment):
"""Recursively finds the fragment to delete"""
symbol, children = fragment
if self.is_excluded(symbol):
return symbol, children
self.to_delete -= 1
if self.to_delete == 0:
return symbol, []
return symbol, list(map(self.recursive_delete, children))
# + [markdown] slideshow={"slide_type": "fragment"}
# We should also define the corresponding structural deletion operator, as well.
# + slideshow={"slide_type": "subslide"}
class FragmentMutator(FragmentMutator):
def __init__(self, parser):
super().__init__(parser)
self.mutators.append(self.delete_fragment)
def delete_fragment(self, seed):
"""Deletes a random fragment"""
if seed.has_structure:
n_nodes = self.count_nodes(seed.structure)
self.to_delete = random.randint(2, n_nodes)
new_structure = self.recursive_delete(seed.structure)
new_seed = Seed(tree_to_string(new_structure))
new_seed.has_structure = True
new_seed.structure = new_structure
# do not return an empty new_seed
if not new_seed.data: return seed
else: return new_seed
return seed
# + [markdown] slideshow={"slide_type": "subslide"}
# ***Summary***. We now have all ingredients for structure-aware fuzzing. Our mutator disassembles all seeds into fragments, which are then added to the fragment pool. Our mutator swaps random fragments in a given seed with fragments of the same type. And our mutator deletes random fragments in a given seed. This allows to maintain a high degree of validity for the generated inputs w.r.t. the given grammar.
#
# ***Try it***. Try adding other structural mutation operators. How would an *add-operator* know the position in a given seed file, where it is okay to add a fragment starting with a certain symbol?
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Fragment-Based Fuzzing
#
# We can now define a input-structure aware fuzzer as pioneered in LangFuzzer. To implement LangFuzz, we modify our [blackbox mutational fuzzer](GreyboxFuzzer.ipynb#Blackbox-Mutation-based-Fuzzer) to stack up to four structural mutations.
# + slideshow={"slide_type": "fragment"}
class LangFuzzer(MutationFuzzer):
def create_candidate(self):
"""Returns an input generated by fuzzing a seed in the population"""
candidate = self.schedule.choose(self.population)
trials = random.randint(1,4)
for i in range(trials):
candidate = self.mutator.mutate(candidate)
return candidate
# + [markdown] slideshow={"slide_type": "subslide"}
# Okay, let's take our first input-structure aware fuzzer for a spin. Being careful, we set n=300 for now.
# + slideshow={"slide_type": "fragment"}
n = 300
runner = FunctionCoverageRunner(my_parser)
mutator = FragmentMutator(EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS))
schedule = PowerSchedule()
langFuzzer = LangFuzzer([valid_seed.data], mutator, schedule)
start = time.time()
langFuzzer.runs(runner, trials = n)
end = time.time()
"It took LangFuzzer %0.2f seconds to generate and execute %d inputs." % (end - start, n)
# + [markdown] slideshow={"slide_type": "subslide"}
# We observe that structural mutation is *sooo very slow*. This is despite our time budget of 200ms for parsing. In contrast, our blackbox fuzzer alone can generate about 10k inputs per second!
# + slideshow={"slide_type": "fragment"}
runner = FunctionCoverageRunner(my_parser)
mutator = Mutator()
schedule = PowerSchedule()
blackFuzzer = MutationFuzzer([valid_seed.data], mutator, schedule)
start = time.time()
blackFuzzer.runs(runner, trials = n)
end = time.time()
"It took a blackbox fuzzer %0.2f seconds to generate and execute %d inputs." % (end - start, n)
# + [markdown] slideshow={"slide_type": "subslide"}
# Indeed, our blackbox fuzzer is done in the blink of an eye.
#
# ***Try it***. We can deal with this overhead using [deferred parsing](https://arxiv.org/abs/1811.09447). Instead of wasting time in the beginning of the fuzzing campaign when a byte-level mutator would make efficient progress, deferred parsing suggests to invest time in structural mutation only later in the fuzzing campaign when it becomes viable.
# + slideshow={"slide_type": "fragment"}
blackbox_coverage = len(runner.coverage())
"During this fuzzing campaign, the blackbox fuzzer covered %d statements." % blackbox_coverage
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's print some stats for our fuzzing campaigns. Since we'll need to print stats more often later, we should wrap this into a function. In order to measure coverage, we import the [population_coverage](Coverage.ipynb#Coverage-of-Basic-Fuzzing) function. It takes a set of inputs and a Python function, executes the inputs on that function and collects coverage information. Specifically, it returns a tuple `(all_coverage, cumulative_coverage)` where `all_coverage` is the set of statements covered by all inputs, and `cumulative_coverage` is the number of statements covered as the number of executed inputs increases. We are just interested in the latter to plot coverage over time.
# + slideshow={"slide_type": "skip"}
from Coverage import population_coverage
# + slideshow={"slide_type": "subslide"}
def print_stats(fuzzer, parser):
coverage, _ = population_coverage(fuzzer.inputs, my_parser)
has_structure = 0
for seed in fuzzer.inputs:
# reuse memoized information
if hasattr(seed, "has_structure"):
if seed.has_structure:
has_structure += 1
else:
if isinstance(seed, str):
seed = Seed(seed)
try:
signal.setitimer(signal.ITIMER_REAL, 0.2)
next(parser.parse(seed.data))
signal.setitimer(signal.ITIMER_REAL, 0)
has_structure += 1
except (SyntaxError, Timeout):
signal.setitimer(signal.ITIMER_REAL, 0)
print("From the %d generated inputs, %d (%0.2f%%) can be parsed.\n"
"In total, %d statements are covered." % (
len(fuzzer.inputs),
has_structure,
100 * has_structure / len(fuzzer.inputs),
len(coverage)))
# + [markdown] slideshow={"slide_type": "subslide"}
# For LangFuzzer, let's see how many of the inputs generated by LangFuzz are valid (i.e., parsable) and how many statements were covered.
# + slideshow={"slide_type": "fragment"}
print_stats(langFuzzer, EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS))
# + [markdown] slideshow={"slide_type": "fragment"}
# What are the stats for the mutational fuzzer that uses only byte-level mutation (and no grammars)?
# + slideshow={"slide_type": "fragment"}
print_stats(blackFuzzer, EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS))
# + [markdown] slideshow={"slide_type": "subslide"}
# ***Summary***. Our fragment-level blackbox fuzzer (LangFuzzer) generates *more valid inputs* but achieves *less code coverage* than a fuzzer with our byte-level fuzzer. So, there is some value in generating inputs that do not stick to the provided grammar.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Integration with Greybox Fuzzing
#
# In the following we integrate fragment-level blackbox fuzzing (LangFuzz-style) with [byte-level greybox fuzzing](GreyboxFuzzer.ipynb#Greybox-Mutation-based-Fuzzer) (AFL-style). The additional coverage-feedback might allow us to increase code coverage more quickly.
#
# A [greybox fuzzer](GreyboxFuzzer.ipynb#Greybox-Mutation-based-Fuzzer) adds to the seed population all generated inputs which increase code coverage. Inputs are generated in two stages, stacking up to four structural mutations and up to 32 byte-level mutations.
# + slideshow={"slide_type": "subslide"}
class GreyboxGrammarFuzzer(GreyboxFuzzer):
def __init__(self, seeds, byte_mutator, tree_mutator, schedule):
super().__init__(seeds, byte_mutator, schedule)
self.tree_mutator = tree_mutator
def create_candidate(self):
"""Returns an input generated by structural mutation of a seed in the population"""
seed = self.schedule.choose(self.population)
# Structural mutation
trials = random.randint(0,4)
for i in range(trials):
seed = self.tree_mutator.mutate(seed)
# Byte-level mutation
candidate = seed.data
if trials == 0 or not seed.has_structure or 1 == random.randint(0, 1):
dumb_trials = min(len(seed.data), 1 << random.randint(1,5))
for i in range(dumb_trials):
candidate = self.mutator.mutate(candidate)
return candidate
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's run our integrated fuzzer with the [standard byte-level mutator](GreyboxFuzzer.ipynb#Mutator-and-Seed) and our [fragment-based structural mutator](#Fragment-based-Mutation) that was introduced above.
# + slideshow={"slide_type": "subslide"}
runner = FunctionCoverageRunner(my_parser)
byte_mutator = Mutator()
tree_mutator = FragmentMutator(EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS))
schedule = PowerSchedule()
gg_fuzzer = GreyboxGrammarFuzzer([valid_seed.data], byte_mutator, tree_mutator, schedule)
start = time.time()
gg_fuzzer.runs(runner, trials = n)
end = time.time()
"It took the greybox grammar fuzzer %0.2f seconds to generate and execute %d inputs." % (end - start, n)
# + slideshow={"slide_type": "subslide"}
print_stats(gg_fuzzer, EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS))
# + [markdown] slideshow={"slide_type": "subslide"}
# ***Summary***. Our structural greybox fuzzer
# * runs faster than the fragment-based LangFuzzer,
# * achieves more coverage than both the fragment-based LangFuzzer and the vanilla blackbox mutational fuzzer, and
# * generates fewer valid inputs than even the vanilla blackbox mutational fuzzer.
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=false
# ## Mutating Invalid Seeds
#
# In the previous section, we have seen that most inputs that are added as seeds are *invalid* w.r.t. our given grammar. Yet, in order to apply our fragment-based mutators, we need it to parse the seed successfully. Otherwise, the entire fragment-based approach becomes useless. The question arises: *How can we derive structure from (invalid) seeds that cannot be parsed successfully?*
#
# To this end, we introduce the idea of _region-based mutation_, first explored with the [AFLSmart](https://github.com/aflsmart/aflsmart) structural greybox fuzzer \cite{Pham2018aflsmart}. AFLSmart implements byte-level, fragment-based, and region-based mutation as well as validity-based power schedules. We define *region-based mutators*, where a *region* is a consecutive sequence of bytes in the input that can be associated with a symbol in the grammar.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Determining Symbol Regions
# The function `chart_parse` of the [Earley parser](Parser.ipynb#The-Parsing-Algorithm) produces a parse table for a string. For each letter in the string, this table gives the potential symbol and a *region* of neighboring letters that might belong to the same symbol.
# + code_folding=[] slideshow={"slide_type": "subslide"}
invalid_seed = Seed("<html><body><i>World</i><br/>>/body></html>")
parser = EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS)
table = parser.chart_parse(invalid_seed.data, parser.start_symbol())
for column in table:
print(column)
print("---")
# + [markdown] slideshow={"slide_type": "subslide"}
# The number of columns in this table that are associated with potential symbols correspond to the number of letters that could be parsed successfully. In other words, we can use this table to compute the longest parsable substring.
# + slideshow={"slide_type": "fragment"}
cols = [col for col in table if col.states]
parsable = invalid_seed.data[:len(cols)-1]
print("'%s'" % invalid_seed)
parsable
# + [markdown] slideshow={"slide_type": "fragment"}
# From this, we can compute the *degree of validity* for an input.
# + slideshow={"slide_type": "subslide"}
validity = 100 * len(parsable) / len(invalid_seed.data)
"%0.1f%% of the string can be parsed successfully." % validity
# + [markdown] slideshow={"slide_type": "subslide"}
# ***Summary***. Unlike input fragments, input regions can be derived even if the parser fails to generate the entire parse tree.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Region-based Mutation
# To fuzz invalid seeds, the region-based mutator associates symbols from the grammar with regions (i.e., indexed substrings) in the seed. The [overridden](#Building-the-Fragment-Pool) method `add_to_fragment_pool()` first tries to mine the fragments from the seed. If this fails, the region mutator uses [Earley parser](Parser.ipynb#The-Parsing-Algorithm) to derive the parse table. For each column (i.e., letter), it extracts the symbols and corresponding regions. This allows the mutator to store the set of regions with each symbol.
# + slideshow={"slide_type": "subslide"}
class RegionMutator(FragmentMutator):
def add_to_fragment_pool(self, seed):
"""Mark fragments and regions in a seed file"""
super().add_to_fragment_pool(seed)
if not seed.has_structure:
try:
signal.setitimer(signal.ITIMER_REAL, 0.2) # set 200ms timeout
seed.regions = {k: set() for k in self.parser.cgrammar}
for column in self.parser.chart_parse(seed.data, self.parser.start_symbol()):
for state in column.states:
if (not self.is_excluded(state.name) and
state.e_col.index - state.s_col.index > 1 and
state.finished()):
seed.regions[state.name].add((state.s_col.index, state.e_col.index))
signal.setitimer(signal.ITIMER_REAL, 0) # cancel timeout
seed.has_regions = True
except Timeout:
seed.has_regions = False
else:
seed.has_regions = False
# + [markdown] slideshow={"slide_type": "subslide"}
# This is how these regions look like for our invalid seed. A region consists of a start and end index in the seed string.
# + slideshow={"slide_type": "subslide"}
mutator = RegionMutator(parser)
mutator.add_to_fragment_pool(invalid_seed)
for symbol in invalid_seed.regions:
print(symbol)
for (s, e) in invalid_seed.regions[symbol]:
print("|-(%d,%d) : %s" % (s, e, invalid_seed.data[s:e]))
# + [markdown] slideshow={"slide_type": "subslide"}
# Now that we know which regions in the seed belong to which symbol, we can define region-based swap and delete operators.
# + slideshow={"slide_type": "subslide"}
class RegionMutator(RegionMutator):
def swap_fragment(self, seed):
"""Chooses a random region and swaps it with a fragment
that starts with the same symbol"""
if not seed.has_structure and seed.has_regions:
regions = [r for r in seed.regions
if (len(seed.regions[r]) > 0 and
len(self.fragments[r]) > 0)]
if len(regions) == 0: return seed
key = random.choice(list(regions))
s, e = random.choice(list(seed.regions[key]))
swap_structure = random.choice(self.fragments[key])
swap_string = tree_to_string(swap_structure)
new_seed = Seed(seed.data[:s] + swap_string + seed.data[e:])
new_seed.has_structure = False
new_seed.has_regions = False
return new_seed
else:
return super().swap_fragment(seed)
# + slideshow={"slide_type": "subslide"}
class RegionMutator(RegionMutator):
def delete_fragment(self, seed):
"""Deletes a random region"""
if not seed.has_structure and seed.has_regions:
regions = [r for r in seed.regions
if len(seed.regions[r]) > 0]
if len(regions) == 0: return seed
key = random.choice(list(regions))
s, e = (0, 0)
while (e - s < 2):
s, e = random.choice(list(seed.regions[key]))
new_seed = Seed(seed.data[:s] + seed.data[e:])
new_seed.has_structure = False
new_seed.has_regions = False
return new_seed
else:
return super().delete_fragment(seed)
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's try our new region-based mutator. We add a simple, valid seed to the fragment pool and attempt to mutate the invalid seed.
# + slideshow={"slide_type": "fragment"}
simple_seed = Seed("<b>Text</b>")
mutator = RegionMutator(parser)
mutator.add_to_fragment_pool(simple_seed)
print(invalid_seed)
mutator.mutate(invalid_seed)
# + [markdown] slideshow={"slide_type": "subslide"}
# ***Summary***. We can use the Earley parser to generate a parse table and assign regions in the input to symbols in the grammar. Our region mutators can substitute these region with fragments from the fragment pool that start with the same symbol, or delete these regions entirely.
#
# ***Try it***. Implement a region pool (similar to the fragment pool) and a `swap_region()` mutator.
# You can execute your own code by opening this chapter as Jupyter notebook.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Region-Based Fuzzing
#
# Let's try our shiny new region mutator by integrating it with our [structure-aware greybox fuzzer](#Integration-with-Greybox-Fuzzing).
# + slideshow={"slide_type": "subslide"}
runner = FunctionCoverageRunner(my_parser)
byte_mutator = Mutator()
tree_mutator = RegionMutator(EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS))
schedule = PowerSchedule()
regionFuzzer = GreyboxGrammarFuzzer([valid_seed.data], byte_mutator, tree_mutator, schedule)
start = time.time()
regionFuzzer.runs(runner, trials = n)
end = time.time()
"It took the structural greybox fuzzer with region mutator\
%0.2f seconds to generate and execute %d inputs." % (end - start, n)
# + [markdown] slideshow={"slide_type": "subslide"}
# We can see that the structural greybox fuzzer with region-based mutator is slower than the [fragment-based mutator alone](#Fragment-based-Fuzzing). This is because region-based structural mutation is applicable for *all seeds*. In contrast, fragment-based mutators were applicable only for tiny number of parsable seeds. Otherwise, only (very efficient) byte-level mutators were applied.
#
# Let's also print the average degree of validity for the seeds in the population.
# + slideshow={"slide_type": "subslide"}
def print_more_stats(fuzzer, parser):
print_stats(fuzzer, parser)
validity = 0
total = 0
for seed in fuzzer.population:
if not seed.data: continue
table = parser.chart_parse(seed.data, parser.start_symbol())
cols = [col for col in table if col.states]
parsable = invalid_seed.data[:len(cols)-1]
validity += len(parsable) / len(seed.data)
total += 1
print("On average, %0.1f%% of a seed in the population can be successfully parsed." % (100 * validity / total))
# + slideshow={"slide_type": "subslide"}
print_more_stats(regionFuzzer, parser)
# + [markdown] slideshow={"slide_type": "subslide"}
# ***Summary***. Compared to fragment-based mutation, a greybox fuzzer with region-based mutation achieves *higher coverage* but generates a *smaller number of valid inputs*. The higher coverage is explained by leveraging at least *some* structure for seeds that cannot be parsed successfully.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Focusing on Valid Seeds
#
# In the previous section, we have a problem: The low (degree of) validity. To address this problem, a _validity-based power schedule_ assigns more [energy](GreyboxFuzzer.ipynb#Power-Schedules) to seeds that have a higher degree of validity. In other words, the fuzzer _spends more time fuzzing seeds that are more valid_.
# + slideshow={"slide_type": "skip"}
import math
# + slideshow={"slide_type": "subslide"}
class AFLSmartSchedule(PowerSchedule):
def __init__(self, parser, exponent):
self.parser = parser
self.exponent = exponent
def parsable(self, seed):
"""Returns the substring that is parsable"""
table = self.parser.chart_parse(seed.data, self.parser.start_symbol())
cols = [col for col in table if col.states]
return seed.data[:len(cols)-1]
def degree_of_validity(self, seed):
"""Returns the proportion of a seed that is parsable"""
if hasattr(seed, "validity"): return seed.validity
seed.validity = (len(self.parsable(seed)) / len(seed.data)
if len(seed.data) > 0 else 0)
return seed.validity
def assignEnergy(self, population):
"""Assign exponential energy proportional to degree of validity"""
for seed in population:
seed.energy = ((self.degree_of_validity(seed) / math.log(len(seed.data))) ** self.exponent
if len(seed.data) > 1 else 0)
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's play with the degree of validity by passing in a valid seed ...
# + slideshow={"slide_type": "fragment"}
smart_schedule = AFLSmartSchedule(parser, 1)
print("%11s: %s" % ("Entire seed", simple_seed))
print("%11s: %s" % ("Parsable", smart_schedule.parsable(simple_seed)))
"Degree of validity: %0.2f%%" % (100 * smart_schedule.degree_of_validity(simple_seed))
# + [markdown] slideshow={"slide_type": "fragment"}
# ... and an invalid seed.
# + slideshow={"slide_type": "subslide"}
print("%11s: %s" % ("Entire seed", invalid_seed))
print("%11s: %s" % ("Parsable", smart_schedule.parsable(invalid_seed)))
"Degree of validity: %0.2f%%" % (100 * smart_schedule.degree_of_validity(invalid_seed))
# + [markdown] slideshow={"slide_type": "fragment"}
# Excellent. We can compute the degree of validity as the proportion of the string that can be parsed.
#
# Let's plug the validity-based power schedule into the structure-aware greybox fuzzer.
# + slideshow={"slide_type": "subslide"}
runner = FunctionCoverageRunner(my_parser)
byte_mutator = Mutator()
tree_mutator = RegionMutator(EarleyParser(XML_GRAMMAR, tokens=XML_TOKENS))
schedule = AFLSmartSchedule(parser, 1)
aflsmart = GreyboxGrammarFuzzer([valid_seed.data], byte_mutator, tree_mutator, schedule)
start = time.time()
aflsmart.runs(runner, trials = n)
end = time.time()
"It took AFLSmart %0.2f seconds to generate and execute %d inputs." % (end - start, n)
# + slideshow={"slide_type": "subslide"}
print_more_stats(aflsmart, parser)
# + [markdown] slideshow={"slide_type": "subslide"}
# ***Summary***. Indeed, by spending more time fuzzing seeds with a higher degree of validity, we also generate inputs with a higher degree of validity. More inputs are entirely valid w.r.t. the given grammar.
#
# ***Read up***. Learn more about region-based fuzzing, deferred parsing, and validity-based schedules in the original AFLSmart paper: "[Smart Greybox Fuzzing](https://arxiv.org/abs/1811.09447)" by Pham et al.. Download and improve AFLSmart: [https://github.com/aflsmart/aflsmart](https://github.com/aflsmart/aflsmart).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Mining Seeds
#
# By now, it should have become clear that the _choice of seeds_ can very much influence the success of fuzzing. One aspect is _variability_ – our seeds should cover as many different features as possible in order to increase coverage. Another aspect, however, is the _likelihood of a seed to induce errors_ – that is, if a seed was involved in causing a failure before, then a mutation of this very seed may be likely to induce failures again. This is because fixes for past failures typically are successful in letting the concrete failure no longer occur, but sometimes may fail to capture all conditions under which a failure may occur. Hence, even if the original failure is fixed, the likelihood of an error in the _surroundings_ of the original failure-inducing input is still higher. It thus pays off to use as seeds _inputs that are known to have caused failures before_.
#
# To put things in context, Holler's _LangFuzz_ fuzzer used as seeds JavaScript inputs from CVE reports. These were published as failure-inducing inputs at a time when the error already had been fixed; thus they could do no harm anymore. Yet, by using such inputs as seeds, LangFuzz would create plenty of mutations and recombinations of all their features, many of which would (and do) find errors again and again.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Lessons Learned
#
# * A **dictionary** is useful to inject important keywords into the generated inputs.
#
# * **Fragment-based mutation** first disassembles seeds into fragments, and reassembles these fragments to generate new inputs. A *fragment* is a subtree in the seed's parse tree. However, fragment-based mutation requires that the seeds can be parsed successfully, which may not be true for seeds discovered by a coverage-based greybox fuzzer.
#
# * **Region-based mutation** marks regions in the input as belonging to a certain symbol in the grammar. For instance, it may identify a substring '</a>' as closing tag. These regions can then be deleted or substituted by fragments or regions belonging to the same symbol. Unlike fragment-based mutation, region-based mutation is applicable to *all* seeds - even those that can be parsed only partially. However, the degree of validity is still quite low for the generated inputs.
#
# * A **validity-based power schedule** invests more energy into seeds with a higher degree of validity. The inputs that are generated also have a higher degree of validity.
#
# * **Mining seeds** from repositories of previous failure-inducing inputs results in input fragments associated with past failures, raising the likelihood to find more failures in the vicinity.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Next Steps
#
# This chapter closes our discussion of syntactic fuzzing techniques.
#
# * In the [next chapter](Reducer.ipynb), we discuss how to _reduce failure-inducing inputs_ after a failure, keeping only those portions of the input that are necessary for reproducing the failure.
# * The [next part](04_Semantical_Fuzzing.ipynb) will go from syntactical to _semantical_ fuzzing, considering code semantics for targeted test generation.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
#
# This chapter builds on the following two works:
#
# * The _LangFuzz_ fuzzer \cite{Holler2012} is an efficient (and effective!) grammar-based fuzzer for (mostly) JavaScript. It uses the grammar for parsing seeds and recombining their inputs with generated parts and found 2,600 bugs in JavaScript interpreters to date.
#
# * Smart greybox fuzzing ([AFLSmart](https://github.com/aflsmart/aflsmart)) brings together coverage-based fuzzing and grammar-based (structural) fuzzing, as described in \cite{Pham2018aflsmart}. The resulting AFLSMART tool has discovered 42 zero-day vulnerabilities in widely-used, well-tested tools and libraries; so far 17 CVEs were assigned.
#
# Recent fuzzing work also brings together grammar-based fuzzing and coverage.
#
# * _Superion_ \cite{Wang2019superion} is equivalent to our section "Integration with Greybox Fuzzing", as above – that is, a combination of LangFuzz and Greybox Fuzzing, but no AFL-style byte-level mutation. Superion can improve the code coverage (i.e., 16.7% and 8.8% in line and function coverage) and bug-finding capability over AFL and jsfunfuzz. According to the authors, they found 30 new bugs, among which they discovered 21 new vulnerabilities with 16 CVEs assigned and 3.2K USD bug bounty rewards received.
#
# * _Nautilus_ \cite{Aschermann2019nautilus} also combines grammar-based fuzzing with coverage feedback. It maintains the parse tree for all seeds and generated inputs. To allow AFL-style byte-level mutations, it "collapses" subtrees back to byte-level representations. This has the advantage of not having to re-parse generated seeds; however, over time, Nautilus de-generates to structure-unaware greybox fuzzing because it does not re-parse collapsed subtrees to reconstitute input structure for later seeds where most of the parse tree is collapsed. Nautilus identified bugs in mruby, PHP, ChakraCore, and in Lua; reporting these bugs was awarded with a sum of 2600 USD and 6 CVEs were assigned.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Exercises
#
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# ### Exercise 1: The Big Greybox Fuzzer Shoot-Out
#
# Use our implementations of greybox techniques and evaluate them on a benchmark. Which technique (and which sub-technique) has which impact and why? Also take into account the specific approaches of Superion \cite{Wang2019superion} and Nautilus \cite{Aschermann2019nautilus}, possibly even on the benchmarks used by these approaches.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** To be added by Summer 2019.
|
docs/notebooks/GreyboxGrammarFuzzer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # <NAME> het hele jaar door
# Bespreking API backend - Python AWS Lambda CloudFunctions.
# ## Één router en vijf API's
#
# 1. lambda_handler, 2. sessionmake, 3. sessionread, 4. userread, 5. userwrite, 6. usersread
#
# De code van deze zes functies wordt hieronder in code blokken getoond maar kan niet gedraaid worden. Daarvoor ontbreekt de AWS Environment. Deze AWS environment levert requests af aan de lambda_handler en stuurt responses terug. Ook is Cloud Storage efficiënt benaderbaar met de boto3 module.
#
# ### Router - lambda_handler
#
# De router kijkt welke variabelen er in de request worden aangeleverd en "beslist" op basis daarvan welke API functie er aangeroepen moet worden. De response de API functie wordt dan weer doorgestuurd naar de client. Leuk feitje: de Pythons requests functie levert base64 versleutelde requests. De Web Fetch levert "ruwe" utf-8 data. De handler moet een decoder bevatten.
# +
def lambda_handler(event, context):
# print("event", event)
req = {}
if 'body' in list( event.keys()):
req = event['body']
if 'isBase64Encoded' in list( event.keys()) and event["isBase64Encoded"]:
req = base64.b64decode(event['body']).decode('utf-8')
req = json.loads( req )
req = req["body"]
elif type(req) is not dict:
req = json.loads( event['body'] )
print("req", req)
reqkeys = list( req.keys() )
if "loginname" in reqkeys:
if req["loginname"]=="session":
if "wwdwsession" in reqkeys:
resjson = usermethods.sessionread(req["wwdwsession"] )
else:
resjson = {}
else:
resjson = usermethods.sessionmake(req["loginname"] )
elif "article" in reqkeys:
resjson = usermethods.userwrite( req )
elif "documenthash" in reqkeys:
resjson = usermethods.userread( req["documenthash"] )
elif "usersmake" in reqkeys:
resjson = usersmethods.usersmake()
else:
resjson = usersmethods.usersread()
return {
"statusCode": 200,
"headers": {
"Content-Type": "application/json"
},
"body": json.dumps( resjson )
}
# -
# ### Session make
#
# De sessionmake API genereert een sessionid en koppelt dit aan een userid. Als een user bijvoorbeeld zijn gegevens wil wijzigen dient deze het sessionid en de te wijzigen data aan te leveren. De wijziging wordt dan doorgevoerd op het userid dat in het bestand sessions.json is gekoppeld aan de een sessionid. Als er nog geen user bestaat voor de loginnaam dan maakt sessionmake tevens een nieuwe (lege) user aan.
# +
def sessionmake(loginname):
nowdtm = datetime.today().strftime('%Y%m%d')
# allways a new session
wwdwsession = nowdtm + "".join((random.choice("abcdefghij0123456789") for i in range(8)))
object_content = s3.Object("wwdw", "users.json")
file_content = object_content.get()['Body'].read().decode('utf-8')
users = json.loads(file_content)
user = list(filter(lambda u: u['loginname'] == loginname, users))
if(len(user)==1):
user = user[0]
wwdwid = user["wwdwid"]
uservoornemens = userread(wwdwid)
else:
# write new wwdwuid
wwdwid = "wwdwid" + nowdtm + "".join((random.choice("abcdefghij0123456789") for i in range(4)))
uservoornemens = {
"username": "",
"loginname": loginname,
"wwdwid": wwdwid,
"voornemen": [{
"text": "",
"step":[{ "text": "" }],
"thought":[{ "text": ""}]
}]
}
s3object = s3.Object('wwdw', wwdwid+'.json')
s3object.put( Body = ( bytes( json.dumps( uservoornemens ).encode('UTF-8') ) ) )
# append new wwdwuid to users
del uservoornemens["voornemen"]
users.append( uservoornemens )
s3object = s3.Object('wwdw', 'users.json')
s3object.put( Body = ( bytes( json.dumps( users ).encode('UTF-8') ) ) )
uservoornemens["wwdwsession"] = wwdwsession
# upsert sessions {}
object_content = s3.Object("wwdw", "sessions.json")
file_content = object_content.get()['Body'].read().decode('utf-8')
sessions = json.loads(file_content)
sessions[wwdwsession] = {"wwdwid": wwdwid}
s3object = s3.Object('wwdw', 'sessions.json')
s3object.put( Body = ( bytes( json.dumps( sessions ).encode('UTF-8') ) ) )
print("session for", loginname, wwdwid, uservoornemens["wwdwsession"] )
del uservoornemens["loginname"]
return uservoornemens
# -
# ### Session read
#
# Op de client wordt het sessionid opgeslagen in een eeuwig durend cookie. Dit cookie wordt meegestuurd met elk request en indien nodig gebruikt. Als de [live demo](https://jhmj-io.github.io/ba-wk2201-wwdw/) opnieuw wordt aangeroepen wordt bij de onload het sessionid uit het cookie op de server geverifieerd. Als het bestaat dan is de user ingelogd.
def sessionread(wwdwsession):
object_content = s3.Object("wwdw", "sessions.json")
file_content = object_content.get()['Body'].read().decode('utf-8')
sessions = json.loads(file_content)
if wwdwsession in list(sessions.keys()):
print("sessionread", wwdwsession, sessions[wwdwsession])
uservoornemens = userread( sessions[wwdwsession]["wwdwid"] )
del uservoornemens["loginname"]
return uservoornemens
else:
return {}
# ### User read
#
# Opvragen van de voornemens data van een user aan de hand van zijn userid.
#
# +
def userread( wwdwid ):
object_content = s3.Object("wwdw", wwdwid + ".json" )
file_content = object_content.get()['Body'].read().decode('utf-8')
usermapped = json.loads(file_content)
usermapped["wwdwid"] = wwdwid # should already be in dict
return usermapped
# -
# Bovenstaande functie levert de response op onderstaand request. Voor de goede orde: het request wordt normaal gesproken uitgevoerd door Web Fetch -in een browser- en de response komt dus van de userread die draait in de AWS Lambda Cloud Function achter: https://8lgmayxgl6.execute-api.eu-central-1.amazonaws.com/default/wwdw
# +
import json
import requests
cloudfunction = 'https://8lgmayxgl6.execute-api.eu-central-1.amazonaws.com/default/wwdw'
req = {
"body": {
"documenthash": "wwdwid202201072841"
}
}
vdata = requests.post(cloudfunction, data=json.dumps(req))
vdata.json()
# -
# ### User write
#
# Opslaan van al dan niet gewijzgde data in het voor elke gebruiker unieke userid.json bestand in Cloud Storage. De data wordt alleen opgeslagen als er een bestaand sessionid wordt meegeleverd. Hackers opgelet: je kan een sessionid vinden in het eeuwig durende cookie en dit dan tevens inzetten op andere computers. Je kan weliswaar alleen de data van de gebruiker gekoppeld aan die session er mee wijzigen.
def userwrite( req ):
wwdwsession = req["wwdwsession"]
article = req["article"]
object_content = s3.Object("wwdw", "sessions.json")
file_content = object_content.get()['Body'].read().decode('utf-8')
sessions = json.loads(file_content)
if wwdwsession not in list(sessions.keys()):
return {"articlewrite": "fail" }
print("uservoornemens article", article )
uservoornemens = userread( sessions[wwdwsession]["wwdwid"] )
print("uservoornemens s3", uservoornemens )
uservoornemens["username"] = article["username"]
uservoornemens["voornemen"] = article["voornemen"]
wwdwid = sessions[wwdwsession]["wwdwid"]
s3object = s3.Object('wwdw', wwdwid+'.json')
s3object.put( Body = ( bytes( json.dumps( uservoornemens ).encode('UTF-8') ) ) )
# update username in users
object_content = s3.Object("wwdw", "users.json")
file_content = object_content.get()['Body'].read().decode('utf-8')
users = json.loads(file_content)
useri = [i for i, d in enumerate(users) if d["wwdwid"]==wwdwid ][0]
users[useri]["username"] = article["username"]
s3object = s3.Object('wwdw', 'users.json')
s3object.put( Body = ( bytes( json.dumps( users ).encode('UTF-8') ) ) )
return {"articlewrite": "OK"}
# Onderstaand een write request dat door bovenstaande functie wordt uitgevoerd.
# +
cloudfunction = 'https://8lgmayxgl6.execute-api.eu-central-1.amazonaws.com/default/wwdw'
req = {
"body": {
"wwdwsession": "bla-die-bla",
"article": {
"username": "Sander!",
"voornemen": [
{
"text": "Meer muziek",
"step": [{"text": "Kan ook luisten"}],
"thought": [{"text": "Gaat samen met Bit Academy"}]
}
]
}
}
}
vdata = requests.post(cloudfunction, data=json.dumps(req))
vdata.json()
# -
# ### Users read
#
# Op de home page wordt een lijst gegeven van de waaghalzen die op de [live demo](https://jhmj-io.github.io/ba-wk2201-wwdw/) hun goede voornemens bespreken.
def usersread():
object_content = s3.Object("wwdw", "users.json")
file_content = object_content.get()['Body'].read().decode('utf-8')
json_content = json.loads(file_content)
usersmapped = list( map(lambda u: {"username": u["username"], "wwdwid": u["wwdwid"]}, json_content) )
return usersmapped
# Onderstaand request vraagt om de user lijst.
# +
cloudfunction = 'https://8lgmayxgl6.execute-api.eu-central-1.amazonaws.com/default/wwdw'
req = {
"body": {
}
}
vdata = requests.post(cloudfunction, data=json.dumps(req))
vdata.json()
|
wwdw.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# The notebook comes alive with the interactive widgets
# ## Speeding up the bottleneck in the REPL
#
# <img src="Flow.svg"></img>
9*9
def f(x):
print(x * x)
f(9)
from ipywidgets import *
interact(f, x=(0, 100));
# # Interactive Jupyter widgets
#
# A Python widget is an object that represents a control on the front end, like a slider. A single control can be displayed multiple times - they all represent the same python object.
# +
slider = FloatSlider(
value=7.5,
min=5.0,
max=10.0,
step=0.1,
description='Input:',
)
slider
# -
slider
# The control attributes, like its value, are automatically synced between the frontend and the kernel.
slider.value
slider.value = 9
# You can trigger actions in the kernel when a control value changes by "observing" the value. Here we set a global variable when the slider value changes.
square = slider.value * slider.value
def handle_change(change):
global square
square = change.new * change.new
slider.observe(handle_change, 'value')
square
square
# You can link control attributes and lay them out together.
text = FloatText(description='Value')
link((slider, 'value'), (text, 'value'))
VBox([slider, text])
# # Jupyter widgets as a framework
#
# Jupyter widgets forms a framework for representing python objects interactively. Some large open-source interactive controls based on Jupyter widgets include:
#
# - bqplot - 2d plotting library
# - pythreejs - low-level 3d graphics library
# - ipyvolume - 3d plotting and volume rendering
# - ipyleaflet - interactive maps
# - ipywebrtc - video streaming
# - ipysheet - interactive spreadsheets
# - ...
|
jupyter_interactive_widgets/notebooks/00.00-introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 投影
# ### PCA
import numpy as np
m = 60
np.random.seed(4)
w1, w2 = 0.1, 0.3
noise = 0.1
angles = np.random.rand(m) * 3 * np.pi / 2 - 0.5
X = np.empty((m, 3))
X[:, 0] = np.cos(angles) + np.sin(angles)/2 + noise * np.random.randn(m) / 2
X[:, 1] = np.sin(angles) * 0.7 + noise * np.random.randn(m) / 2
X[:, 2] = X[:, 0] * w1 + X[:, 1] * w2 + noise * np.random.randn(m)
# +
# 使用主成分分析,返回一个经过主成分变换的实例数组
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
X2D = pca.fit_transform(X)
# -
# pca属性访问
pca.explained_variance_ratio_
# 打印所有主成分,每一个主成分为一个水平向量
pca.components_
# +
pca = PCA()
pca.fit(X)
cumsum = np.cumsum(pca.explained_variance_ratio_)
# 找出所有累计解释率大于0.95的所有组合,取索引
d = np.argmax(cumsum>=0.95) + 1
# -
pca = PCA(n_components=0.95)
X_reduced = pca.fit_transform(X)
pca.components_
# ## 增量PCA
from sklearn.decomposition import IncrementalPCA
n_batches =100
for X_batch in np.array_split(X)
x=np.random.rand(19)
np.split(x, 4)
np.array_split(x, 4)
|
data_science/7_dimensionality_reduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ### 导包
import numpy as np
import matplotlib.pyplot as plt
from keras.models import Sequential
from keras.layers import LSTM, TimeDistributed, Dense
from keras.optimizers import Adam
# ### 自定义函数
# +
BATCH_START = 0
TIME_STEPS = 20
BATCH_SIZE = 50
INPUT_SIZE = 1
OUTPUT_SIZE = 1
CELL_SIZE = 20
def get_batch():
global BATCH_START, TIME_STEPS #定义全局变量
# xs shape (50batch, 20steps)
xs = np.arange(BATCH_START, BATCH_START+TIME_STEPS*BATCH_SIZE).reshape((BATCH_SIZE, TIME_STEPS)) / (10*np.pi)
seq = np.sin(xs)
res = np.cos(xs)
BATCH_START += TIME_STEPS
# plt.plot(xs[0, :], res[0, :], 'r', xs[0, :], seq[0, :], 'b--')
# plt.show()
return [seq[:, :, np.newaxis], res[:, :, np.newaxis], xs]
#np.newaxis的功能是在行或列上插入新维度
# -
bat = get_batch()
print(bat[0].shape)
print(bat[1].shape)
print(bat[2].shape)
# +
a = np.arange(5)
print(a,a.shape)
a = a[:,np.newaxis] #在列上增加维度
print(a,a.shape)
a = a[np.newaxis,:] #在行上增加维度
print(a,a.shape)
b = np.array([[1,2,3],[4,5,6]])
print(b)
print(b[1:])
# -
# ### 构建模型
model = Sequential()
# build a LSTM RNN
model.add(LSTM(
batch_input_shape=(BATCH_SIZE, TIME_STEPS, INPUT_SIZE), # Or: input_dim=INPUT_SIZE, input_length=TIME_STEPS,
units=CELL_SIZE, #LSTM层输出维度
return_sequences=True, # True: output at all steps. False: output as last step.
stateful=True, # True: the final state of batch1 is feed into the initial state of batch2
#If a RNN is stateful, it needs to know its batch size.
))
# add output layer
model.add(TimeDistributed(Dense(OUTPUT_SIZE)))
# ### 编译模型
LR = 0.006
adam = Adam(LR)
model.compile(optimizer=adam,loss='mse',)
# ### 训练模型
print('Training ------------')
for step in range(501):
# data shape = (batch_num, steps, inputs/outputs)
X_batch, Y_batch, xs = get_batch()
cost = model.train_on_batch(X_batch, Y_batch)
pred = model.predict(X_batch, BATCH_SIZE) #用正弦拟合余弦
plt.plot(xs[0, :], Y_batch[0].flatten(), 'r', xs[0, :], pred.flatten()[:TIME_STEPS], 'b--')
plt.ylim((-1.5, 1.5))
plt.draw()
plt.pause(0.1)
if step % 10 == 0:
print('train cost: ', cost)
plt.show()
|
keras_rnn_LSTM(regressor).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="93fdd79c-3f56-47c6-9497-f0c55a672b49" _kg_hide-input=false _kg_hide-output=false _uuid="822f4627a6a1a96be570de15c34efbbc7df7940a"
# Simple trading analysis methods are listed.
# + _cell_guid="4ccc9463-00bb-4ec3-b0ef-3d129adf9cbc" _uuid="b25269396b81f3991953c302e434486834d91d75"
import pandas as pd
import matplotlib.pyplot as plt
def stock_graph(symbol,title):
df = pd.read_csv("../input/Data/Stocks/{}.us.txt".format(symbol))
df[['Close']].plot()
plt.title(title)
plt.show()
stock_graph("aapl","Apple Stock")
# + _cell_guid="e7924feb-e3f5-4730-b4f6-f2a48ce74047" _uuid="96946e065504fac9ad7d6107e57a919f007fd24d"
# Get stock data for multiple stocks for given symbols and dates and graph it
def stocks_data(symbols, dates):
df = pd.DataFrame(index=dates)
for symbol in symbols:
df_temp = pd.read_csv("../input/Data/Stocks/{}.us.txt".format(symbol), index_col='Date',
parse_dates=True, usecols=['Date', 'Close'], na_values=['nan'])
df_temp = df_temp.rename(columns={'Close': symbol})
df = df.join(df_temp)
return df
dates = pd.date_range('2016-01-02','2016-12-31',freq='B')
symbols = ['goog','ibm','aapl']
df = stocks_data(symbols, dates)
df.fillna(method='pad')
#print(df)
df.interpolate().plot()
plt.show()
# + _cell_guid="049dcda8-03ca-49de-b241-6638e8384620" _uuid="6a18769e40467c1e02371803344efe8d3718d636"
# Normalized Stocks - base value from 2016-01-04
print(df.iloc[1,:])
df = df / df.iloc[1,:]
df.interpolate().plot()
plt.show()
# + _cell_guid="ae1dd296-6bd2-450e-b83b-d35b1253be2d" _uuid="4a5bcd1c086c6f2679922ed5312a4182994b8e29"
# Daily Returns for a symbol with date range
def daily_return(df):
dr = df.copy()
dr = dr[:-1].values / dr[1:] - 1
return dr
dates = pd.date_range('2016-01-01','2016-12-31',freq='B')
symbols = ['aapl']
df = stocks_data(symbols, dates)
dr = daily_return(df)
dr = dr.interpolate()
dr.interpolate().plot()
plt.title('Apple Daily Returns')
plt.show()
dr.hist(bins=20)
plt.show()
# + _cell_guid="ccf1503e-0cc3-48b0-acb9-dd9fa71055f4" _uuid="d6b5a11f3545e3d6028dba5d186345a078e04be2"
#Cumulative Returns
def cum_return(df):
dr = df.copy()
dr.cumsum()
return dr
dates = pd.date_range('2016-01-01','2016-12-31',freq='B')
symbols = ['aapl']
df = stocks_data(symbols, dates)
dr = cum_return(df)
dr.plot()
plt.title('Apple Cumulative Returns')
plt.show()
dr.hist()
# + _cell_guid="f924db65-16f1-4992-a531-7b3a00393be7" _uuid="563af54acf5abd8ecc4054b47a517263beafdb52"
# Scatterplot between GOOG vs AAPL
dates = pd.date_range('2016-01-01','2016-12-31',freq='B')
symbols = ['goog','aapl']
df = stocks_data(symbols, dates)
dr = daily_return(df)
dr.plot(kind='scatter',x='goog', y='aapl')
plt.show()
# + _cell_guid="47ebc05b-d080-4a75-82f8-af580ab51971" _uuid="e679fca2af64d2f58b1f5813e4991a748ed5fc6e"
# Technical Indicators
# Bollinger Bands
def get_bbands(df, ndays):
db = df.copy()
dm = df.rolling(ndays).mean()
ds = df.rolling(ndays).std()
db['upperBB'] = dm + 2 * ds
db['lowerBB'] = dm - 2 * ds
return db
# Simple Moving Average
def get_SMA(df, ndays):
dm = df.copy()
dm.rolling(ndays).mean()
return dm
# Expontential Moving Average
def get_EMA(df, ndays):
dm = df.ewm( span = ndays, min_periods = ndays - 1).mean()
return dm
# Rate of Change
def get_ROC(df, ndays):
dn = df.diff(ndays)
dd = df.shift(ndays)
dr = dn/dd
return dr
dates = pd.date_range('2016-01-01','2016-12-31',freq='B')
symbols = ['aapl']
df = stocks_data(symbols, dates)
dm = get_SMA(df, 10)
dm.plot()
plt.title('Simple Moving Average')
plt.show()
dm = get_EMA(df, 10)
dm.plot()
plt.title('Expontential Moving Average')
plt.show()
dr = get_ROC(df, 1)
dr.plot()
plt.title('Rate of Change')
plt.show()
|
stochastic-study/3.simple-stock-analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tensorflow import keras
model_enged = keras.models.load_model('Models/engagement86.h5')
model_bored = keras.models.load_model('Models/boredom.h5')
model_conf = keras.models.load_model('Models/confusion.h5')
model_frus = keras.models.load_model('Models/Frustration.h5')
model_enged._name = 'engagement'
model_conf._name = 'confusion'
model_bored._name = 'boredom'
model_frus._name = 'frustration'
input = keras.layers.Input(shape = [48,48,1])
y_enged = model_enged(input)
y_conf = model_conf(input)
y_bored = model_bored(input)
y_frust = model_frus(input)
model = keras.Model(inputs=input, outputs=[y_enged, y_conf,y_bored, y_frust])
keras.utils.plot_model(
model,
to_file="parallel_model.png",
show_shapes=True,
expand_nested=True,
dpi=150,
)
model.save('parallel_model.h5')
|
Training/parallel_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="bV_8eZ_-ffaD"
# #Install Dependencies
# + colab={"base_uri": "https://localhost:8080/"} id="-lWvHG1-e6YU" outputId="0eae445c-4963-402d-cf96-b887584836d0"
# !pip install nltk
# !git clone https://github.com/ContentSide/lingx.git
import pandas as pd
from nltk import agreement
# + [markdown] id="m_aemCQ-fl0q"
# #Provide path to CSV file
# + id="YllztayHfBE8"
path_to_csv_file = "/content/lingx/resources/TPRDB/EN-ZH_IMBst18/HumanEvaluations/errors_for_cal_kappa.csv"
# + [markdown] id="WNB_R6GyfqV7"
# #Read and Normalize the dataframe
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="SMzILuvUe2oc" outputId="5163ca13-23f2-42ac-d498-3f4cc2db414b"
df = pd.read_csv(path_to_csv_file)
df_numeral = df[['Any', 'Accuracy', 'Fluency',
'Style', 'Critical','Minor']]
normalized_df=(df_numeral-df_numeral.min())/(df_numeral.max()-df_numeral.min())
normalized_df = pd.concat([
df[['SessionSeg', 'Annotator']],
normalized_df,
],
axis=1)
normalized_df
# + [markdown] id="LngQ-Au_fv3c"
# # Create two functions to feed `nltk.agreement.AnnotationTask` class
# + colab={"base_uri": "https://localhost:8080/"} id="nD6GE89ffPgL" outputId="dfc18c6c-38a4-40aa-b4c2-0b0710f22abb"
def get_annotation_task(
coder_column = "Annotator",
item_column = "SessionSeg",
label_column = "Any"):
annotation_task = []
for index, item in normalized_df.iterrows():
annotation_task.append([item[coder_column], item[item_column], item[label_column]])
return annotation_task
def distance_function(x,y):
delta=x-y
return abs(delta)
# to test the function
annotation_task = get_annotation_task(coder_column = "Annotator", item_column = "SessionSeg", label_column = "Any")
print("Sample of annotation task data:\n")
print(annotation_task[0:2])
# + [markdown] id="VCW1STUDgHXC"
# # Calculating Inter-Coder Agreements
# + colab={"base_uri": "https://localhost:8080/"} id="f4Ehd5k7fQgT" outputId="b9abe28b-d4ba-463f-8f0d-38e73aeedac9"
label_column = ['Any', 'Accuracy', 'Fluency',
'Style', 'Critical','Minor']
def distance_function(x,y):
delta=x-y
return abs(delta)
for item in label_column:
annotation_task = get_annotation_task(coder_column = "Annotator", item_column = "SessionSeg", label_column = item)
rating_task = agreement.AnnotationTask(data=annotation_task, distance=distance_function)
print(f"Label : {item}")
print("kappa " +str(rating_task.kappa()))
print("fleiss " + str(rating_task.multi_kappa()))
print("alpha " +str(rating_task.alpha()))
print("scotts " + str(rating_task.pi()))
print()
# + id="GnXTlILohGOO"
|
resources/IHCI2021/Inter_Coder_Agreement.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="-nW-8StPOhv0" outputId="7b002823-b72b-4e05-a67e-315d1e11bb9f"
# !git clone https://github.com/omidrk/RPcovidActiveLearning.git
# + id="O5uug9qIRopL"
# !pip install captum
# + id="oVMd29y4PLyn"
import os
os.chdir('/content/RPcovidActiveLearning')
# + [markdown] id="_1CUk7-7Pxx5"
# First model is trained in passive mode. To achive this we defined 5 diffrent setting. Each setting, code have been changed menually so running the code wont reproduce the same result.
# + [markdown] id="5eriVjAaQavq"
# First setting: train for 1 epoch and 20% of all dataset.
# Train with 18000 data, batch 100,180 iteration
# + colab={"base_uri": "https://localhost:8080/"} id="VzBwZ0UrPQHW" outputId="dd854a42-dcc4-4ce5-bd5f-38092b0777c1"
# !python main.py
# + [markdown] id="EoWKvybFSeKX"
# setting 2: epoch 2, batch 100
# + colab={"base_uri": "https://localhost:8080/"} id="eDBm7GF7PVt_" outputId="f9611637-cb2e-4b1e-a63e-61e7a71cfdce"
# !python main.py
# + [markdown] id="_D1fp9ZqTAsP"
# Setting 3: epoch 5, batch 100
# + colab={"base_uri": "https://localhost:8080/"} id="DXo8optITFNu" outputId="e70d1a36-5c08-4f1a-a0d8-8e756ef9f941"
# !python main.py
|
Experiments/PassiveSetting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="CCQY7jpBfMur"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="z6X9omPnfO_h" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="F1xIRPtY0E1w"
# # Обзор Keras
# + [markdown] colab_type="text" id="VyOjQZHhZxaA"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/keras/overview"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Смотрите на TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ru/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запустите в Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/ru/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучайте код на GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/ru/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Скачайте ноутбук</a>
# </td>
# </table>
# + [markdown] id="fj66ZXAzrJC2" colab_type="text"
# Note: Вся информация в этом разделе переведена с помощью русскоговорящего Tensorflow сообщества на общественных началах. Поскольку этот перевод не является официальным, мы не гарантируем что он на 100% аккуратен и соответствует [официальной документации на английском языке](https://www.tensorflow.org/?hl=en). Если у вас есть предложение как исправить этот перевод, мы будем очень рады увидеть pull request в [tensorflow/docs](https://github.com/tensorflow/docs) репозиторий GitHub. Если вы хотите помочь сделать документацию по Tensorflow лучше (сделать сам перевод или проверить перевод подготовленный кем-то другим), напишите нам на [<EMAIL> list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ru).
# + [markdown] colab_type="text" id="VUJTep_x5-R8"
# Это руководство даст вам основы для начала работы с Keras. Чтение займет 10 минут.
# + [markdown] colab_type="text" id="IsK5aF2xZ-40"
# ## Импортируйте tf.keras
#
# `tf.keras` является реализацией TensorFlow
# [спецификации Keras API](https://keras.io). Это высокоуровневый
# API для построения и обучения моделей включающий первоклассную поддержку для
# TensorFlow-специфичной функциональности, такой как [eager execution](../eager.ipynb),
# конвейеры `tf.data`, и [Estimators](../estimator.ipynb).
# `tf.keras` делает использование TensorFlow проще не жертвуя при этом гибкостью и
# производительностью.
#
# Для начала, импортируйте `tf.keras` как часть установки вашей TensorFlow:
# + colab_type="code" id="TgPcBFru0E1z" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# # %tensorflow_version существуют только в Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
from tensorflow import keras
# + [markdown] colab_type="text" id="lj03RamP0E13"
# `tf.keras` может выполнять любой Keras-совместимый код, но имейте ввиду:
#
# * Версия `tf.keras` в последнем релизе TensorFlow может отличаться от
# последней версии `keras` в PyPI. Проверьте `tf.keras.__version__`.
# * При [сохранении весоа моделей](./save_and_serialize.ipynb), `tf.keras` делает это по умолчанию
# [в формате checkpoint](../checkpoint.ipynb). Передайте параметр `save_format='h5'` для
# использования HDF5 (или добавьте к имени файла расширение `.h5`).
# + [markdown] colab_type="text" id="7e1LPcXx0gR6"
# ## Постройте простую модель
#
# ### Последовательная модель
#
# В Keras, вы собираете *слои (layers)* для построения *моделей (models)*. Модель это (обычно) граф
# слоев. Наиболее распространенным видом модели является стек слоев:
# модель `tf.keras.Sequential`.
#
# Построение простой полносвязной сети (т.е. многослойного перцептрона):
# + colab_type="code" id="WM-DUVQB0E14" colab={}
from tensorflow.keras import layers
model = tf.keras.Sequential()
# Добавим полносвязный слой с 64 узлами к модели:
model.add(layers.Dense(64, activation='relu'))
# Добавим другой слой:
model.add(layers.Dense(64, activation='relu'))
# Добавим слой softmax с 10 выходами:
model.add(layers.Dense(10, activation='softmax'))
# + [markdown] colab_type="text" id="I2oH0-cxH7YA"
# Вы можете найти короткий, но полный пример того, как использовать последовательные (Sequential) модели [здесь](https://www.tensorflow.org/tutorials/quickstart/beginner).
#
# Чтобы узнать о построении более сложных чем последовательные (Sequential), см:
# - [Руководство по Keras Functional API](./functional.ipynb)
# - [Руководство по написанию слоев и моделей с сабклассингом с нуля](./custom_layers_and_models.ipynb)
# + [markdown] colab_type="text" id="-ztyTipu0E18"
# ### Настройте слои
#
# Доступно много разновидностей слоев `tf.keras.layers`. Большинство из них используют общий конструктор
# аргументов:
#
# * `activation`: Установка функции активации для слоя. В этом параметре
# указывается имя встроенной функции или вызываемый объект. У параметра
# нет значения по умолчанию.
# * `kernel_initializer` И `bias_initializer`: Схемы инициализации
# создающие веса слоя (ядро и сдвиг). В этом параметре может быть имя
# или вызываемый объект. По умолчанию используется инициализатор `"Glorot uniform"`.
# * `kernel_regularizer` и `bias_regularizer`: Схемы регуляризации
# добавляемые к весам слоя (ядро и сдвиг), такие как L1 или L2
# регуляризации. По умолчанию регуляризация не устанавливается.
#
# Следующие примеры слоев `tf.keras.layers.Dense` используют
# аргументы конструктора:
# + colab_type="code" id="MlL7PBtp0E19" colab={}
# Создать слой с сигмоидой:
layers.Dense(64, activation='sigmoid')
# Или:
layers.Dense(64, activation=tf.keras.activations.sigmoid)
# Линейный слой с регуляризацией L1 с коэфициентом 0.01 примененной к матрице ядра:
layers.Dense(64, kernel_regularizer=tf.keras.regularizers.l1(0.01))
# Линейный слой с регуляризацией L2 с коэффициентом 0.01 примененной к вектору сдвига:
layers.Dense(64, bias_regularizer=tf.keras.regularizers.l2(0.01))
# Линейный слой с ядром инициализированным случайной ортогональной матрицей:
layers.Dense(64, kernel_initializer='orthogonal')
# Линейный слой с вектором сдвига инициализированным значениями 2.0:
layers.Dense(64, bias_initializer=tf.keras.initializers.Constant(2.0))
# + [markdown] colab_type="text" id="9NR6reyk0E2A"
# ## Обучение и оценка
#
# ### Настройка обучения
#
# После того как модель сконструирована, настройте процесс ее обучения вызовом
# метода `compile`:
# + colab_type="code" id="sJ4AOn090E2A" colab={}
model = tf.keras.Sequential([
# Добавляем полносвязный слой с 64 узлами к модели:
layers.Dense(64, activation='relu', input_shape=(32,)),
# Добавляем другой слой:
layers.Dense(64, activation='relu'),
# Добавляем слой softmax с 10 выходами:
layers.Dense(10, activation='softmax')])
model.compile(optimizer=tf.keras.optimizers.Adam(0.01),
loss='categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] colab_type="text" id="HG-RAa9F0E2D"
# `tf.keras.Model.compile` принимает три важных аргумента:
#
# * `optimizer`: Этот объект определяет процедуру обучения. Передайте в него экземпляры
# оптимизатора из модуля `tf.keras.optimizers`, такие как
# `tf.keras.optimizers.Adam` или
# `tf.keras.optimizers.SGD`. Если вы просто хотите использовать значения по умолчанию, вы также можете указать оптимизаторы ключевыми словами, такими как `'adam'` или `'sgd'`.
# * `loss`: Это функция которая минимизируется в процессе обучения. Среди распространенных вариантов
# mean square error (`mse`), `categorical_crossentropy`, и
# `binary_crossentropy`. Функции потерь указываются по имени или по
# передаче вызываемого объекта из модуля `tf.keras.losses`.
# * `metrics`: Используются для мониторинга обучения. Это строковые имена или вызываемые объекты из
# модуля `tf.keras.metrics`.
# * Кроме того, чтобы быть уверенным, что модель обучается и оценивается eagerly, проверьте что вы передали компилятору параметр `run_eagerly=True`
#
#
# Далее посмотрим несколько примеров конфигурации модели для обучения:
# + colab_type="code" id="St4Mgdar0E2E" colab={}
# Сконфигурируем модель для регрессии со среднеквадратичной ошибкой.
model.compile(optimizer=tf.keras.optimizers.Adam(0.01),
loss='mse', # срееднеквадратичная ошибка
metrics=['mae']) # средняя абсолютная ошибка
# Сконфигурируем модель для категориальной классификации.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.01),
loss=tf.keras.losses.CategoricalCrossentropy(),
metrics=[tf.keras.metrics.CategoricalAccuracy()])
# + [markdown] colab_type="text" id="yjI5rbi80E2G"
# ### Обучение на данных NumPy
#
# Для небольших датасетов используйте помещающиеся в память массивы [NumPy](https://www.numpy.org/)
# для обучения и оценки модели. Модель "обучается" на тренировочных даннных
# используя метод `fit`:
# + colab_type="code" id="3CvP6L-m0E2I" colab={}
import numpy as np
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
model.fit(data, labels, epochs=10, batch_size=32)
# + [markdown] colab_type="text" id="N-pnVaFe0E2N"
# `tf.keras.Model.fit` принимает три важных аргумента:
#
# * `epochs`: Обучение разбито на *эпохи*. Эпоха это одна итерация
# по всем входным данным (это делается небольшими партиями).
# * `batch_size`: При передаче данных NumPy, модель разбивает данные на меньшие
# блоки (batches) и итерирует по этим блокам во время обучения. Это число
# указывает размер каждого блока данных. Помните, что последний блок может быть меньшего
# размера если общее число записей не делится на размер партии.
# * `validation_data`: При прототипировании модели вы хотите легко отслеживать её
# производительность на валидационных данных. Передача с этим аргументом кортежа входных данных
# и меток позволяет модели отопражать значения функции потерь и метрики в режиме вывода
# для передаваемых данных в конце каждой эпохи.
#
# Вот пример использования `validation_data`:
# + colab_type="code" id="gFcXzVQa0E2N" colab={}
import numpy as np
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
val_data = np.random.random((100, 32))
val_labels = np.random.random((100, 10))
model.fit(data, labels, epochs=10, batch_size=32,
validation_data=(val_data, val_labels))
# + [markdown] colab_type="text" id="-6ImyXzz0E2Q"
# ### Обучение с использованием наборов данных tf.data
#
# Используйте [Datasets API](../data.ipynb) для масштабирования больших баз данных
# или обучения на нескольких устройствах. Передайте экземпляр `tf.data.Dataset` в метод
# `fit`:
# + colab_type="code" id="OziqhpIj0E2R" colab={}
# Создает экземпляр учебного датасета:
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
model.fit(dataset, epochs=10)
# + [markdown] colab_type="text" id="I7BcMHkB0E2U"
# Поскольку `Dataset` выдает данные пакетами, этот кусок кода не требует аргумента `batch_size`.
#
# Датасеты могут быть также использованы для валидации:
# + colab_type="code" id="YPMb3A0N0E2V" colab={}
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
val_dataset = tf.data.Dataset.from_tensor_slices((val_data, val_labels))
val_dataset = val_dataset.batch(32)
model.fit(dataset, epochs=10,
validation_data=val_dataset)
# + [markdown] colab_type="text" id="IgGdlXso0E2X"
# ### Оценка и предсказание
#
# Методы `tf.keras.Model.evaluate` и `tf.keras.Model.predict` могут использовать данные
# NumPy и `tf.data.Dataset`.
#
# Вот так можно *оценить* потери в режиме вывода и метрики для предоставленных данных:
# + colab_type="code" id="mhDbOHEK0E2Y" colab={}
# С массивом Numpy
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
model.evaluate(data, labels, batch_size=32)
# С датасетом
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
model.evaluate(dataset)
# + [markdown] colab_type="text" id="UXUTmDfb0E2b"
# А вот как *предсказать* вывод последнего уровня в режиме вывода для предоставленных данных,
# в виде массива NumPy:
# + colab_type="code" id="9e3JsSoQ0E2c" colab={}
result = model.predict(data, batch_size=32)
print(result.shape)
# + [markdown] colab_type="text" id="GuTb71gYILLG"
# Полное руководство по обучению и оценке модели, включая описание написания пользовательских циклов обучения с нуля, см. в [руководстве по обучению и оценке] (./ train_and_evaluate.ipynb).
# + [markdown] colab_type="text" id="fzEOW4Cn0E2h"
# ## Построение сложных моделей
#
# ### The Functional API
#
# Модель `tf.keras.Sequential` это простой стек слоев с помощью которого
# нельзя представить произвольную модель. Используйте
# [Keras functional API](./functional.ipynb)
# для построения сложных топологий моделей, таких как:
#
# * Модели с несколькими входами,
# * Модели с несколькими выходами,
# * Модели с общими слоями (один и тот же слой вызывается несколько раз),
# * Модели с непоследовательными потоками данных (напр. остаточные связи).
#
# Построение модели с functional API работает следующим образом:
#
# 1. Экземпляр слоя является вызываемым и возвращает тензор.
# 2. Входные и выходные тензоры используются для определения экземпляра
# `tf.keras.Model`
# 3. Эта модель обучается точно так же как и `Sequential` модель.
#
# Следующий пример использует functional API для построения простой, полносвязной
# сети:
# + colab_type="code" id="mROj832r0E2i" colab={}
inputs = tf.keras.Input(shape=(32,)) # Возвращает входной плейсхолдер
# Экземпляр слоя вызывается на тензор и возвращает тензор.
x = layers.Dense(64, activation='relu')(inputs)
x = layers.Dense(64, activation='relu')(x)
predictions = layers.Dense(10, activation='softmax')(x)
# + [markdown] colab_type="text" id="AFmspHeG1_W7"
# Создайте экземпляр модели с данными входами и выходами.
# + colab_type="code" id="5k5uzlyu16HM" colab={}
model = tf.keras.Model(inputs=inputs, outputs=predictions)
# Шаг компиляции определяет конфигурацию обучения.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Обучение за 5 эпох
model.fit(data, labels, batch_size=32, epochs=5)
# + [markdown] colab_type="text" id="EcKSLH3i0E2k"
# ### Сабклассинг моделей
#
# Создайте полностью настраиваемую модель с помощью сабклассинга `tf.keras.Model` и определения
# вашего собственного прямого распространения. Создайте слои в методе `__init__` и установите их как
# атрибуты экземпляра класса. Определите прямое распространение в методе `call`.
#
# Сабклассинг модели особенно полезен когда включен
# [eager execution](../eager.ipynb), поскольку он позволяет написать
# прямое распространение императивно.
#
# Примечание: если вам нужно чтобы ваша модель *всегда* выполнялась императивно, вы можете установить `dynamic=True` когда вызываете конструктор `super`.
#
# > Ключевой момент: Используйте правильный API для работы. Хоть сабклассинг модели обеспечивает
# гибкость, за нее приходится платить большей сложностью и большими возможностями для
# пользовательских ошибок. Если это возможно выбирайте functional API.
#
# Следующий пример показывает сабклассированную `tf.keras.Model` использующую пользовательское прямое
# распространение, которое не обязательно выполнять императивно:
# + colab_type="code" id="KLiHWzcn2Fzk" colab={}
class MyModel(tf.keras.Model):
def __init__(self, num_classes=10):
super(MyModel, self).__init__(name='my_model')
self.num_classes = num_classes
# Определим свои слои тут.
self.dense_1 = layers.Dense(32, activation='relu')
self.dense_2 = layers.Dense(num_classes, activation='sigmoid')
def call(self, inputs):
# Определим тут свое прямое распространение,
# с использованием ранее определенных слоев (в `__init__`).
x = self.dense_1(inputs)
return self.dense_2(x)
# + [markdown] colab_type="text" id="ShDD4fv72KGc"
# Создайте экземпляр класса новой модели:
# + colab_type="code" id="42C-qQHm0E2l" colab={}
model = MyModel(num_classes=10)
# Шаг компиляции определяет конфигурацию обучения.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Обучение за 5 эпох.
model.fit(data, labels, batch_size=32, epochs=5)
# + [markdown] colab_type="text" id="yqRQiKj20E2o"
# ### Пользовательские слои
#
# Создайте пользовательский слой сабклассингом `tf.keras.layers.Layer` и реализацией
# следующих методов:
#
# * `__init__`: Опционально определите подслои которые будут использоваться в этом слое.
# * `build`: Создайте веса слоя. Добавьте веса при помощи метода
# `add_weight`.
# * `call`: Определите прямое распространение.
# * Опционально, слой может быть сериализован реализацией метода `get_config`
# и метода класса `from_config`.
#
# Ниже пример пользовательского слоя который осуществляет умножение матрицы (`matmul`) поданной на вход
# с матрицей ядра:
# + colab_type="code" id="l7BFnIHr2WNc" colab={}
class MyLayer(layers.Layer):
def __init__(self, output_dim, **kwargs):
self.output_dim = output_dim
super(MyLayer, self).__init__(**kwargs)
def build(self, input_shape):
# Создадим обучаемую весовую переменную для этого слоя.
self.kernel = self.add_weight(name='kernel',
shape=(input_shape[1], self.output_dim),
initializer='uniform',
trainable=True)
def call(self, inputs):
return tf.matmul(inputs, self.kernel)
def get_config(self):
base_config = super(MyLayer, self).get_config()
base_config['output_dim'] = self.output_dim
return base_config
@classmethod
def from_config(cls, config):
return cls(**config)
# + [markdown] colab_type="text" id="8wXDRgXV2ZrF"
# Создайте модель с использованием вашего пользовательского слоя:
# + colab_type="code" id="uqH-cY0h0E2p" colab={}
model = tf.keras.Sequential([
MyLayer(10),
layers.Activation('softmax')])
# Шаг компиляции определяет конфигурацию обучения
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Обучение за 5 эпох.
model.fit(data, labels, batch_size=32, epochs=5)
# + [markdown] colab_type="text" id="llipvR5wIl_t"
# Узнайте больше о создании новых слоев и моделей с нуля с помощью сабклассинга в [Руководстве написания слоев и моделей с нуля](./custom_layers_and_models.ipynb).
# + [markdown] colab_type="text" id="Lu8cc3AJ0E2v"
# ## Колбеки
#
# Колбек это объект переданный модели чтобы кастомизировать и расширить ее поведение
# во время обучения. Вы можете написать свой пользовательский колбек или использовать встроенный
# `tf.keras.callbacks` который включает:
#
# * `tf.keras.callbacks.ModelCheckpoint`: Сохранение контрольных точек модели за
# регулярные интервалы.
# * `tf.keras.callbacks.LearningRateScheduler`: Динамичное изменение шага
# обучения.
# * `tf.keras.callbacks.EarlyStopping`: Остановка обучения в том случае когда
# результат на валидации перестает улучшаться.
# * `tf.keras.callbacks.TensorBoard`: Мониторинг поведения модели с помощью
# [TensorBoard](https://tensorflow.org/tensorboard).
#
# Для использования `tf.keras.callbacks.Callback`, передайте ее методу модели `fit`:
# + colab_type="code" id="rdYwzSYV0E2v" colab={}
callbacks = [
# Остановить обучение если `val_loss` перестанет улучшаться в течение 2 эпох
tf.keras.callbacks.EarlyStopping(patience=2, monitor='val_loss'),
# Записать логи TensorBoard в каталог `./logs`
tf.keras.callbacks.TensorBoard(log_dir='./logs')
]
model.fit(data, labels, batch_size=32, epochs=5, callbacks=callbacks,
validation_data=(val_data, val_labels))
# + [markdown] colab_type="text" id="ghhaGfX62abv"
# <a name='save_and_restore'></a>
# ## Сохранение и восстановление
# + [markdown] colab_type="text" id="qnl7K-aI0E2z"
# <a name="weights_only"></a>
# ### Сохранение только значений весов
#
# Сохраните и загрузите веса модели с помощью `tf.keras.Model.save_weights`:
# + colab_type="code" id="uQIANjB94fLB" colab={}
model = tf.keras.Sequential([
layers.Dense(64, activation='relu', input_shape=(32,)),
layers.Dense(10, activation='softmax')])
model.compile(optimizer=tf.keras.optimizers.Adam(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# + colab_type="code" id="4eoHJ-ny0E21" colab={}
# Сохраним веса в файл TensorFlow Checkpoint
model.save_weights('./weights/my_model')
# Восстановим состояние модели,
# для этого необходима модель с такой же архитектурой.
model.load_weights('./weights/my_model')
# + [markdown] colab_type="text" id="u25Id3xe0E25"
# По умолчанию веса модели сохраняются в формате
# [TensorFlow checkpoint](../checkpoint.ipynb). Веса могут быть также
# сохранены в формате Keras HDF5 (значение по умолчанию для универсальной
# реализации Keras):
# + colab_type="code" id="JSAYoFEd0E26" colab={}
# Сохранение весов в файл HDF5
model.save_weights('my_model.h5', save_format='h5')
# Восстановление состояния модели
model.load_weights('my_model.h5')
# + [markdown] colab_type="text" id="mje_yKL10E29"
# ### Сохранение только конфигурации модели
#
# Конфигурация модели может быть сохранена - это сериализует архитектуру модели
# без всяких весов. Сохраненная конфигурация может восстановить и инициализировать ту же
# модель, даже без кода определяющего исходную модель. Keras поддерживает
# форматы сериализации JSON и YAML:
# + colab_type="code" id="EbET0oJTzGkq" colab={}
# Сериализация модели в формат JSON
json_string = model.to_json()
json_string
# + colab_type="code" id="pX_badhH3yWV" colab={}
import json
import pprint
pprint.pprint(json.loads(json_string))
# + [markdown] colab_type="text" id="Q7CIa05r4yTb"
# Восстановление модели (заново инициализированной) из JSON:
# + colab_type="code" id="J9UFv9k00E2_" colab={}
fresh_model = tf.keras.models.model_from_json(json_string)
# + [markdown] colab_type="text" id="t5NHtICh4uHK"
# Сериализация модели в формат YAML требует установки `pyyaml` *перед тем как импортировать TensorFlow*:
# + colab_type="code" id="aj24KB3Z36S4" colab={}
yaml_string = model.to_yaml()
print(yaml_string)
# + [markdown] colab_type="text" id="O53Kerfl43v7"
# Восстановление модели из YAML:
# + colab_type="code" id="77yRuwg03_MG" colab={}
fresh_model = tf.keras.models.model_from_yaml(yaml_string)
# + [markdown] colab_type="text" id="xPvOSSzM0E3B"
# Внимание: сабклассированные модели не сериализуемы, потому что их архитектура
# определяется кодом Python в теле метода `call`.
# + [markdown] colab_type="text" id="iu8qMwld4-71"
#
# ### Сохранение всей модели в один файл
#
# Вся модель может быть сохранена в файл содержащий значения весов, конфигурацию
# модели, и даже конфигурацию оптимизатора. Это позволит вам
# установить контрольную точку модели и продолжить обучение позже с точно того же положения
# даже без доступа к исходному коду.
# + colab_type="code" id="45oNY34Z0E3C" colab={}
# Создадим простую модель
model = tf.keras.Sequential([
layers.Dense(10, activation='softmax', input_shape=(32,)),
layers.Dense(10, activation='softmax')
])
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(data, labels, batch_size=32, epochs=5)
# Сохраним всю модель в файл HDF5
model.save('my_model.h5')
# Пересоздадим в точности эту модель включая веса и оптимизатор.
model = tf.keras.models.load_model('my_model.h5')
# + [markdown] colab_type="text" id="wGVBURDtI_I6"
# Узнайте больше о сохранении и сериализации моделей Keras в руководстве по [сохранению и сериализации моделей](./save_and_serialize.ipynb).
# + [markdown] colab_type="text" id="PMOWhDOB0E3E"
# <a name="eager_execution"></a>
# ## Eager execution
#
# [Eager execution](../eager.ipynb) это императивное программирование
# среда которая выполняет операции немедленно. Это не требуется для
# Keras, но поддерживается `tf.keras` и полезно для проверки вашей программы и
# отладки.
#
# Все строящие модели API `tf.keras` совместимы eager execution.
# И хотя могут быть использованы `Sequential` и functional API, eager execution
# особенно полезно при *сабклассировании модели* и построении *пользовательских слоев* — эти API
# требуют от вас написание прямого распространения в виде кода (вместо API которые
# создают модели путем сборки существующих слоев).
#
# См [руководство eager execution](../eager.ipynb) для
# примеров использования моделей Keras с пользовательскими циклами обучения и `tf.GradientTape`.
# Вы можете также найти полный коротки пример [тут](https://www.tensorflow.org/tutorials/quickstart/advanced).
# + [markdown] colab_type="text" id="2wG3NVco5B5V"
# ## Распределение
#
# + [markdown] colab_type="text" id="6PJZ6e9J5JHF"
# ### Множественные GPU
#
# `tf.keras` модели можно запускать на множестве GPU с использованием
# `tf.distribute.Strategy`. Этот API обеспечивает распределенное
# обучение на нескольких GPU практически без изменений в существующем коде.
#
# На данный момент, `tf.distribute.MirroredStrategy` единственная поддерживаемая
# стратегия распределения. `MirroredStrategy` выполняет репликацию в графах с
# синхронным обучением используя all-reduce на одной машине. Для использования
# `distribute.Strategy` , вложите инсталляцию оптимизатора, конструкцию и компиляцию модели в `Strategy`'s `.scope()`, затем
# обучите модель.
#
# Следующий пример распределяет `tf.keras.Model` между множеством GPU на
# одной машине.
#
# Сперва определим модель внутри области распределенной стратегии:
# + colab_type="code" id="sbaRr7g-0E3I" colab={}
strategy = tf.distribute.MirroredStrategy()
with strategy.scope():
model = tf.keras.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10,)))
model.add(layers.Dense(1, activation='sigmoid'))
optimizer = tf.keras.optimizers.SGD(0.2)
model.compile(loss='binary_crossentropy', optimizer=optimizer)
model.summary()
# + [markdown] colab_type="text" id="rO9MiL6X0E3O"
# Затем обучим модель на данных как обычно:
# + colab_type="code" id="BEwFq4PM0E3P" colab={}
x = np.random.random((1024, 10))
y = np.random.randint(2, size=(1024, 1))
x = tf.cast(x, tf.float32)
dataset = tf.data.Dataset.from_tensor_slices((x, y))
dataset = dataset.shuffle(buffer_size=1024).batch(32)
model.fit(dataset, epochs=1)
# + [markdown] colab_type="text" id="N6BXU5F90E3U"
# Для дополнительной информации см. [полное руководство по распределенному обучению наа TensorFlow](../distributed_training.ipynb).
|
site/ru/guide/keras/overview.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cloud forest demo
#
# This notebook demonstrates how to work with contemporary cloud-based spatial analysis and indexing tools. It accompanies [this blog post](https://salo.ai/blog/2021/01/cfo-cloud). We'll work with:
#
# - Cloud-Optimized Geotiffs (COG)
# - Google Earth Engine
# - SpatioTemporal Asset Catalogs (STAC)
#
# We'll also describe integrations between these tools and the CFO API.
#
# If you have conda installed, you can run the following to load this notebook:
#
# ```bash
# git clone https://github.com/forestobservatory/cfo-api.git
# # cd cfo-api
# conda env update
# conda activate cfo
# jupyter notebook
# ```
# ## Section 0: loading packages, authenticating resources, defining functions.
# +
# %matplotlib notebook
import os
import cfo
import ee
import folium
import numpy as np
import pystac
import matplotlib.pyplot as plt
from osgeo import gdal, ogr, osr
from time import time
gdal.UseExceptions()
# -
# we'll write a couple of files to your local machine, so be sure to change this path!
outdir = "/home/cba/cfo"
# if this is your first time running earth engine from a notebook, run this code block.
ee.Authenticate()
# +
# while you don't need to authenticate each time, you do need to initialize earth engine each time
ee.Initialize()
# same with the cfo api
forest = cfo.api()
forest.authenticate()
# -
# this code block will verify you're connected to earth engine by printing the elevation of Mount Everest
dem = ee.Image('USGS/SRTMGL1_003')
peak = [86.9250, 27.9881]
xy = ee.Geometry.Point(peak)
elev = dem.sample(xy, 30).first().get('elevation').getInfo()
print('Mount Everest elevation (m):', elev)
# Below are some functions and parameters we'll use later. You don't have to read closely; just run the block.
# +
# color palettes
cfo_green = ["#f9fae5", "#ccd682", "#a6bd34", "#72b416", "#325900"]
cfo_change = ["#E34649", "#FDC591", "#E8E84C", "#9AD94C", "#2093BD"]
# add earth engine tiles to folium
def addLayer(self, eeImage, visParams={}, name="Layer"):
"""
Adds an earth engine image object to a folium map.
source: https://colab.research.google.com/github/google/earthengine-api/blob/master/python/examples/ipynb/ee-api-colab-setup.ipynb
"""
map_id_dict = ee.Image(eeImage).getMapId(visParams)
folium.raster_layers.TileLayer(
tiles=map_id_dict["tile_fetcher"].url_format,
attr="Map Data © <a href='https://earthengine.google.com/'>Google Earth Engine</a>",
name=name,
overlay=True,
control=True,
).add_to(self)
# add the map function to the map object
folium.Map.addLayer = addLayer
# coordinate to pixel conversion
def pixel_project(raster_path: str, xy: list, epsg: int = 4326, wkt: str = None):
"""
Converts geographic coordinates to pixel units for an input raster file.
"""
# read the raster data properties
ref = gdal.Open(raster_path, gdal.GA_ReadOnly)
nx = ref.RasterXSize
ny = ref.RasterYSize
geo = ref.GetGeoTransform()
inv_geo = gdal.InvGeoTransform(geo)
prj = ref.GetProjection()
raster_srs = osr.SpatialReference()
raster_srs.ImportFromWkt(prj)
ref = None
# create the inverse transformation
xy_srs = osr.SpatialReference()
if wkt is None:
xy_srs.ImportFromEPSG(epsg)
else:
xy_srs.ImportFromWkt(wkt)
xy_srs.SetAxisMappingStrategy(osr.OAMS_TRADITIONAL_GIS_ORDER)
raster_srs.SetAxisMappingStrategy(osr.OAMS_TRADITIONAL_GIS_ORDER)
transformer = osr.CoordinateTransformation(xy_srs, raster_srs)
# re-project the point and compute raster coordinates
point = ogr.Geometry(ogr.wkbPoint)
point.AddPoint(xy[0], xy[1])
point.Transform(transformer)
x_px, y_px = [int(p) for p in gdal.ApplyGeoTransform(inv_geo, point.GetX(), point.GetY())]
# warn if these values are outside the raster boundaries
if x_px < 0 or y_px < 0 or x_px > nx or y_px > ny:
raise Exception("Coordinates are outside raster extent")
return x_px, y_px
# -
# # Section 1. Reading and clipping COG data with gdal
#
# One of the key advantages of cloud-optimized geotiffs is that you can rapidly read arbitrary subsets of data. We'll work through an example reading the height of a single tree at a point location and an example of clipping to the extent of a recent wildfire.
#
# In the api searches below, we set `geography="California"` to return the full statewide datasets. For help with the options you can set for the parameters below, try `forest.list_metrics()`, `forest.list_geography_types()`, `forest.list_geographies()`, etc.
# +
# use the cfo api to get the 2020 high res canopy height asset ID
ch_ids = forest.search(
geography="California",
metric="CanopyHeight",
year=2020,
resolution=3,
)
# this should return a 1-element list, and we'll need a string, so extract it
ch_id = ch_ids.pop()
print(f"CFO 2020 3m Asset ID: {ch_id}")
# then get the virtual file path
ch_path = forest.fetch(ch_id, gdal=True)
# -
# In this first block, we'll use Forest Observatory data to get the height of a neat Monterey Pine at entrance to the San Francisco Botanic Gardens in Golden Gate Park.
# +
# set the lat/lon coordinates of the tree (from google maps)
pine_coords = [-122.46776, 37.7673]
# time the cell
start = time()
# since these are in lat/lon, we'll first convert to pixel values. This command will fail outside of California.
x, y = pixel_project(ch_path, pine_coords, epsg=4326)
# then read the value at that location
ch_ref = gdal.Open(ch_path, gdal.GA_ReadOnly)
height = ch_ref.ReadAsArray(x, y, 1, 1)
print(f"Height of the Monterey Pine at the SF Botanic Garden Entrance: {int(height)} meters tall")
# close the file reference and report duration
ch_ref = None
duration = time() - start
print(f"Time elapsed: {duration:0.3f} seconds")
# -
# The first request takes a few seconds, but if you update the coordinates and make additional requests, it typically takes less than 1/10th of a second! Much faster than climbing the tree. But, less fun.
#
# Next, we'll use `gdal` to clip a map of canopy height to the extent of the CZU fire. We'll use a file provided in this repository that contains the geometry of the fire's extent.
# +
# set the output file path
ch_split = ch_id.split('-')
ch_split[0] = "CZU"
ch_filename = "-".join(ch_split) + ".tif"
ch_output = os.path.join(outdir, ch_filename)
# report starting
print(f"Clipping CFO asset ID {ch_id} to file: {ch_output}")
# use the geojson file in this directory
vector_path = 'czu-perimeter.geojson'
# time the cell
start = time()
# set the file creation options
options = gdal.WarpOptions(
creationOptions = ["COMPRESS=DEFLATE", "TILED=YES", "BIGTIFF=YES", "NUM_THREADS=ALL_CPUS"],
cutlineDSName = vector_path,
cropToCutline = True,
)
# and run the command
warp = gdal.Warp(ch_output, ch_path, options=options)
warp.FlushCache()
del warp
# report duration
duration = time() - start
print(f"Time elapsed: {duration:0.3f} seconds")
# -
# Pretty good timing, considering we're pulling 3 meter resolution data from the cloud and clipping it to the extent of a major wildfire.
#
# Let's plot it for reference
# +
# read and mask the data
ch_ref = gdal.Open(ch_output, gdal.GA_ReadOnly)
band = ch_ref.GetRasterBand(1)
height = ch_ref.ReadAsArray().astype(float)
height[height == band.GetNoDataValue()] = np.nan
ch_ref = None
# get the range to show based on a 2% stretch
vmin = np.nanpercentile(height, 1)
vmax = np.nanpercentile(height, 99)
# and plot
plt.figure(figsize=(5,5), dpi=100)
cover_map = plt.imshow(
height,
vmin=vmin,
vmax=vmax,
cmap=plt.cm.viridis,
)
plt.title("Pre-fire canopy height in the\nperimeter of the CZU fire")
colorbar = plt.colorbar(cover_map)
colorbar.set_label("Canopy Height (%)")
plt.tight_layout()
# -
# # Section 2: Google Earth Engine
#
# Next, we'll demonstrate how to read and interact with these data in Google Earth Engine. We'll map canopy cover in 2016 and 2020, then compute the change between years with simple band math.
#
# The change map will show areas of canopy loss in warm colors (reds and yellows) and canopy gain in cool colors (blues and greens). These correspond to areas where wildfires and timber harvest removed trees, and to areas where regeneration restored forests.
#
# We only have 3 meter resolution data available for these two years, so we won't specify years in the API search request.
# +
# first we'll get the asset IDs
cc_ids = forest.search(
geography="California",
metric="CanopyCover",
resolution=3,
)
cc_ids.sort()
print(cc_ids)
# get the bucket paths for where each file is stored
cc_bucket_paths = [forest.fetch(cc_id, bucket=True) for cc_id in cc_ids]
# +
# read them as earth engine images
cc_2016 = ee.Image.loadGeoTIFF(cc_bucket_paths[0])
cc_2020 = ee.Image.loadGeoTIFF(cc_bucket_paths[1])
# compute the difference between years
change = cc_2020.subtract(cc_2016)
# mask changes below a change detection threshold. We selected 15% because the canopy cover RMSE is ~11%
thresh = 15
mask = change.gt(thresh).Or(change.lt(-thresh))
change = change.mask(mask)
# -
# Now that we've loaded the data into Earth Engine and computed the difference between years, we'll create an interactive map to show these data.
# +
# set up visualization parameters
coverViz = {
"min": 0,
"max": 90,
"palette": cfo_green,
}
changeViz = {
"min": -30,
"max": 30,
"palette": cfo_change,
}
# create the notebook map object
mapCenter = [41.2, -123]
m = folium.Map(
location=mapCenter,
zoom_start=9,
tiles="Stamen Toner",
)
# add each layer
m.addLayer(cc_2016, coverViz, "Canopy Cover 2016")
m.addLayer(cc_2020, coverViz, "Canopy Cover 2020")
m.addLayer(change, changeViz, "Change")
# add a color ramp
change_ramp = folium.branca.colormap.LinearColormap(
cfo_change,
vmin = -30,
vmax = 30,
caption = "% Canopy Cover Change, 2016-2020"
)
change_ramp.add_to(m)
# and add some map controls
m.add_child(folium.LayerControl())
display(m)
# -
# I recommend turning the 2020 layer on and off to visualize the change between each years, and to look at change alone on map background.
#
# Don't forget to zoom in--you'll find all sorts of detail.
#
# The final Earth Engine task will be to get the canopy cover pixel value for the tree from the Botanic Garden shown earlier.
# it'll be a simple point lookup
xy = ee.Geometry.Point(pine_coords)
pine_cover = cc_2020.sample(xy, 3).first().get('B0').getInfo()
print(f'Monterey Pine Canopy Cover: {pine_cover}%')
# # Section 3: STAC search
#
# Our final demo will be brief, and relates to querying the Forest Observatory datasets via an emerging geospatial data indexing standard, STAC.
#
# We'll just run through a couple of example functions for working with the Forest Observatory STAC catalog.
# +
# set and read the catalog
catalog_url = "https://storage.googleapis.com/cfo-public/catalog.json"
catalog = pystac.Catalog.from_file(catalog_url)
# what's in this catalog?
print(catalog.description)
# iteratively retrieve the items within the catalog
print("\nCatalog contents:")
print(catalog.describe())
# -
# There's only one colletion within this catalog now, which you see at the top of the `.describe()` output. We can use the `id` of this collection to retrieve it, then query the collection for more information
# +
veg = catalog.get_child('vegetation')
items = veg.get_all_items()
print(veg.description)
print(f"\nThere are {len(list(items))} items in this collection:")
for item in veg.get_all_items():
metric = item.properties['metric']
units = item.properties['units']
resolution = item.properties['gsd']
date = item.get_datetime()
print(f"{date.year} {metric} - {units} - {resolution}m gsd")
# -
# Each item in this dictionary stores information on what spatial extent it covers, the time and date of collection, as well as links to related objects.
# each item contains a lot of information, which you can easily retrieve
item.to_dict()
# Within each of these items can be a nested set of assets, which contain paths to the sources of the geospatial data.
# I've found this to be a rather circuitous route to getting asset data..
assets = item.get_assets()
asset_keys = assets.keys()
for key in asset_keys:
asset = assets[key]
print(asset)
# # Conclusion
#
# Thanks for following along with us. We're working hard to make sure our data are easy to access, use & understand, all in service of trying to support California's conservation and climate change mitigation goals.
#
# Please visit the Forest Observatory [user forums](https://groups.google.com/a/forestobservatory.com/g/community?pli=1) or [get in touch](mailto:<EMAIL>) if you have any questions.
|
demos/cloud-forest-demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="HfnpnWU2rUmM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2249} outputId="364de1af-1c38-413a-bd2f-3727108a1fec"
# #!git clone https://github.com/zhan006/TensorFlow-Tutorials.git
import os
os.chdir("/content/TensorFlow-Tutorials/")
import tensorflow as tf
from mnist import MNIST
import mnist
import matplotlib.pyplot as plt
import numpy as np
data=MNIST()
imgshape=data.img_shape
x=tf.placeholder('float',[None,784],name="origin")
y_=tf.placeholder('float',[None,10],name="truelabel")
loss = -tf.reduce_mean(y_ * tf.log(y_pred))
x_image=tf.reshape(x,[-1,28,28,1])
cov1_weight=init_weight([5,5,1,32])
cov1_bias=init_bias([32])
cov1_c=conv2(x_image,cov1_weight)+cov1_bias
cov1_a=tf.nn.relu(cov1_c)
cov1_max=maxpooling(cov1_a)
cov2_weight=init_weight([3,3,32,64])
cov2_bias=init_bias([64])
cov2_c=conv2(cov1_max,cov2_weight)+cov2_bias
cov2_a=tf.nn.relu(cov2_c)
cov_max=maxpooling(cov2_a)
fc1_input=tf.reshape(cov_max,[-1,7*7*64])
weight1=init_weight([7*7*64,1024])
bias1=init_bias([1024])
fc1_logit=tf.matmul(fc1_input,weight1)+bias1
fc1_ac=tf.nn.relu(fc1_logit)
weight2=init_weight([1024,10])
bias2=init_bias([10])
fc2_logit=tf.matmul(fc1_ac,weight2)+bias2
y_pred=tf.nn.softmax(fc2_logit)
y_pred_cls=tf.argmax(y_pred,1)
x_test=data.x_test
y_test=data.y_test
y_cls=data.y_test_cls
#print(y_cls)
sess=tf.Session()
optimizer=tf.train.GradientDescentOptimizer(0.1).minimize(loss)
sess.run(tf.global_variables_initializer())
y_true=tf.placeholder(tf.int64,[None])
accuracy=tf.equal(y_true,y_pred_cls)
acc=tf.reduce_mean(tf.cast(accuracy,tf.float32))
def plotnumber(arr,label,pre_label=None):
fig,axe=plt.subplots(3,3)
fig.subplots_adjust(hspace=1,wspace=0.3)
for i,ax in enumerate(axe.flat):
ax.imshow(arr[i].reshape(imgshape))
ax.set_xticks([])
ax.set_yticks([])
if pre_label is None:
xlabel=label[i]
ax.set_xlabel('true: '+str(xlabel))
else:
xlabel=label[i]
tlabel=pre_label[i]
ax.set_xlabel('true:{} pre:{}'.format(xlabel,tlabel))
plt.show()
def training(number):
for i in range(number):
xbat,ybat,_=data.random_batch(50)
print(xbat.shape,ybat.shape)
feeddict={x:xbat,y_:ybat}
sess.run(optimizer,feed_dict=feeddict)
def acce():
feed={x:x_test,y_true:y_cls}
a=sess.run(acc,feed_dict=feed)
print(a)
def prediction():
feed={x:x_test}
pr=sess.run(y_pred_cls,feed_dict=feed)
return pr
def pltweights():
w=sess.run(weights)
fig,axe=plt.subplots(3,4)
wmax=np.max(w)
wmin=np.min(w)
for i,axe in enumerate(axe.flat):
if i<10:
image=w[:,i].reshape(imgshape)
axe.imshow(image,vmax=wmax,vmin=wmin,cmap='seismic')
axe.set_xticks([])
axe.set_yticks([])
plt.show()
def init_weight(shape):
init=tf.truncated_normal(shape,stddev=0.1)
return tf.Variable(init)
def init_bias(shape):
init=tf.constant(0.1,shape=shape)
return tf.Variable(init)
def conv2(x,W):
return tf.nn.conv2d(x,W,strides=[1,1,1,1],padding="SAME")
def maxpooling(x):
return tf.nn.max_pool(x,ksize=[1,2,2,1],strides=[1,2,2,1],padding="SAME")
training(5000)
acce()
pr=prediction()
plotnumber(x_test[5:14],y_cls[5:14],pr[5:14])
#pltweights()
|
prc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# download & unzip files (for windows user you have to manually downlad and extract files)
# make sure the extracted folder img_test & img_train contain list of images only
# ! wget -q https://github.com/CISC-372/Notebook/releases/download/a3/test.zip -O test.zip
# ! wget -q https://github.com/CISC-372/Notebook/releases/download/a3/train.zip -O train.zip
# ! wget -q https://github.com/CISC-372/Notebook/releases/download/a3/y_train.csv -O y_train.csv
# ! unzip -q test.zip
# ! unzip -q train.zip
# +
from tqdm import tqdm
from PIL import Image
import pandas as pd
from tqdm.notebook import tqdm
import os
import numpy as np
def load_data(folder):
images = []
for file in tqdm(os.listdir(folder)):
file_id = file.replace('.png', '')
image = Image.open(
os.path.join(folder, file)
).convert('RGBA').resize((256, 256))
arr = np.array(image)
images.append(
(int(file_id), arr)
)
images.sort(key=lambda i: i[0])
return np.array([v for _id, v in images])
x_train = load_data('train')
y_train = pd.read_csv('y_train.csv')['infection']
# -
# check image loading
import matplotlib.pyplot as plt
plt.imshow(x_train[5])
# +
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, Flatten, Dense, Dropout, Input
def build():
img_in = Input(shape=(256, 256, 4))
flattened = Flatten()(img_in)
fc1 = Dense(64)(flattened)
#fc1 = Dropout(0.3)(fc1)
fc2 = Dense(32)(fc1)
#fc2 = Dropout(0.3)(fc2)
output = Dense(1, activation = 'sigmoid')(fc2)
model = tf.keras.Model(inputs=img_in, outputs=output)
return model
model = build()
model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss='binary_crossentropy',
metrics=['BinaryAccuracy', 'AUC']
)
model.summary()
# +
epochs = 30
batch_size = 64
history = model.fit(x = x_train,
y = y_train,
batch_size = batch_size,
validation_split=0.3,
epochs=epochs
)
# +
x_test = load_data('test')
y_test = model.predict(x_test)
y_test_df = pd.DataFrame()
y_test_df['id'] = np.arange(len(y_test))
y_test_df['infection'] = y_test.astype(float)
y_test_df.to_csv('submission.csv', index=False)
# -
|
a3_2021_template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="P6xk2_jLsvFF"
# # [作業目標]
#
# 1. [簡答題] 請問下列程式碼,運算結果分別為何?
#
# ```
# a = np.array( [20,30,40,50] )
# b = np.array( [1,2,3,4] )
# c = 1
# d = np.array( [1] )
# e = np.array( [1,2] )
# ```
#
# 2. 如何在不用迴圈的情況下計算 (A+B)*(-A/2) ?那用迴圈怎麼做?
#
# 3. 請問如何計算「1x6 的單位矩陣」和「6x1 的單位矩陣」的內積和外積?
# + [markdown] colab_type="text" id="uXGll28asvFS"
# # 作業
# -
# ### 1. [簡答題] 請問下列程式碼,運算結果分別為何?
#
# ```
# a = np.array( [20,30,40,50] )
# b = np.array( [1,2,3,4] )
# c = 1
# d = np.array( [1] )
# e = np.array( [1,2] )
# ```
#
# + colab={} colab_type="code" id="5QrYoyNWsvFS"
import numpy as np
a = np.array( [20,30,40,50] )
b = np.array( [1,2,3,4] )
c = 1
d = np.array( [1] )
e = np.array( [1,2] )
print(a + a)
print(a + b)
print(a + c)
print(a + d)
print(a + e) #error 形狀不同無法廣播
# -
# ### 2. 如何在不用迴圈的情況下計算 (A+B)*(-A/2) ?那用迴圈怎麼做?
#
# +
# 記得先 Import 正確的套件
import numpy as np
# +
A = np.ones(3)*1
B = np.ones(3)*2
print ((A+B)*(-A/2))
# -
# ### 3. 請問如何計算「1x6 的單位矩陣」和「6x1 的單位矩陣」的內積和外積?
#
# + colab={} colab_type="code" id="e1gVI0tvsvFY"
A = np.ones((1,6))
B = np.ones((6,1))
print(A*B)
print(A@B)
# -
|
homework/04 Homework_Melissa.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from matplotlib.image import imread
import numpy as np
import matplotlib.pyplot as plt
import os
plt.rcParams['figure.figsize'] = [12, 8]
plt.rcParams.update({'font.size': 18})
A = imread(os.path.join('..','DATA','dog.jpg'))
B = np.mean(A, -1); # Convert RGB to grayscale
Bt = np.fft.fft2(B)
Btsort = np.sort(np.abs(Bt.reshape(-1))) # sort by magnitude
# Zero out all small coefficients and inverse transform
for keep in (0.1, 0.05, 0.01, 0.002):
thresh = Btsort[int(np.floor((1-keep)*len(Btsort)))]
ind = np.abs(Bt)>thresh # Find small indices
Atlow = Bt * ind # Threshold small indices
Alow = np.fft.ifft2(Atlow).real # Compressed image
plt.figure()
plt.imshow(Alow,cmap='gray')
plt.axis('off')
plt.title('Compressed image: keep = ' + str(keep))
# -
|
code/CH02/CH02_SEC06_2_Compress.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="cvXwyS263AMk" outputId="57646cdd-58c1-4ddb-8805-9178cb0a2048"
"""
You can run either this notebook locally (if you have all the dependencies and a GPU) or on Google Colab.
Instructions for setting up Colab are as follows:
1. Open a new Python 3 notebook.
2. Import this notebook from GitHub (File -> Upload Notebook -> "GITHUB" tab -> copy/paste GitHub URL)
3. Connect to an instance with a GPU (Runtime -> Change runtime type -> select "GPU" for hardware accelerator)
4. Run this cell to set up dependencies.
"""
# If you're using Google Colab and not running locally, run this cell.
## Install dependencies
# !pip install wget
# !apt-get install sox libsndfile1 ffmpeg
# !pip install unidecode
# ## Install NeMo
BRANCH = 'main'
# !python -m pip install git+https://github.com/NVIDIA/NeMo.git@$BRANCH#egg=nemo_toolkit[asr]
## Install TorchAudio
# !pip install torchaudio>=0.6.0 -f https://download.pytorch.org/whl/torch_stable.html
## Grab the config we'll use in this example
# !mkdir configs
# + [markdown] colab_type="text" id="Kqg4Rwki4jBX"
# # Introduction
#
# Data augmentation is a useful method to improve the performance of models which is applicable across multiple domains. Certain augmentations can also substantially improve robustness of models to noisy samples.
#
# In this notebook, we describe how to construct an augmentation pipeline inside [Neural Modules (NeMo)](https://github.com/NVIDIA/NeMo), enable augmented training of a [MatchboxNet model](https://arxiv.org/abs/2004.08531 ) ( based on QuartzNet, from the paper ["QuartzNet: Deep Automatic Speech Recognition with 1D Time-Channel Separable Convolutions"](https://arxiv.org/abs/1910.10261)) and finally how to construct custom augmentations to add to NeMo.
#
# The notebook will follow the steps below:
#
# - Dataset preparation: Preparing a noise dataset using an example file.
#
# - Construct a data augmentation pipeline.
#
# - Construct a custom augmentation and register it for use in NeMo.
# + [markdown] colab_type="text" id="5XieMEo84pJ-"
# ## Note
# Data augmentation is valuable for many datasets, but it comes at the cost of increased training time if samples are augmented during training time. Certain augmentations are particularly costly, in terms of how much time they take to process a single sample. A few examples of slow augmentations available in NeMo are :
#
# - Speed Perturbation
# - Time Stretch Perturbation (Sample level)
# - Noise Perturbation
# - Impulse Perturbation
# - Time Stretch Augmentation (Batch level, Neural Module)
#
# For such augmentations, it is advisable to pre-process the dataset offline for a one time preprocessing cost and then train the dataset on this augmented training set.
# + [markdown] colab_type="text" id="Tgc_ZHDl4sMy"
# ## Taking a Look at Our Data (AN4)
#
# The AN4 dataset, also known as the Alphanumeric dataset, was collected and published by Carnegie Mellon University. It consists of recordings of people spelling out addresses, names, telephone numbers, etc., one letter or number at a time, as well as their corresponding transcripts. We choose to use AN4 for this tutorial because it is relatively small, with 948 training and 130 test utterances, and so it trains quickly.
#
# Before we get started, let's download and prepare the dataset. The utterances are available as `.sph` files, so we will need to convert them to `.wav` for later processing. Please make sure you have [Sox](http://sox.sourceforge.net/) installed for this step (instructions to setup your environment are available at the beginning of this notebook).
# + colab={} colab_type="code" id="DtLm_XuQ3pmk"
# This is where the an4/ directory will be placed.
# Change this if you don't want the data to be extracted in the current directory.
data_dir = '.'
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="HjfLhUtH4wNc" outputId="f0a9cd46-6709-49dd-9103-1e0ef61de745"
import glob
import os
import subprocess
import tarfile
import wget
# Download the dataset. This will take a few moments...
print("******")
if not os.path.exists(data_dir + '/an4_sphere.tar.gz'):
an4_url = 'http://www.speech.cs.cmu.edu/databases/an4/an4_sphere.tar.gz'
an4_path = wget.download(an4_url, data_dir)
print(f"Dataset downloaded at: {an4_path}")
else:
print("Tarfile already exists.")
an4_path = data_dir + '/an4_sphere.tar.gz'
if not os.path.exists(data_dir + '/an4/'):
# Untar and convert .sph to .wav (using sox)
tar = tarfile.open(an4_path)
tar.extractall(path=data_dir)
print("Converting .sph to .wav...")
sph_list = glob.glob(data_dir + '/an4/**/*.sph', recursive=True)
for sph_path in sph_list:
wav_path = sph_path[:-4] + '.wav'
cmd = ["sox", sph_path, wav_path]
subprocess.run(cmd)
print("Finished conversion.\n******")
# + [markdown] colab_type="text" id="HqJmf4WB5P1x"
# You should now have a folder called `an4` that contains `etc/an4_train.transcription`, `etc/an4_test.transcription`, audio files in `wav/an4_clstk` and `wav/an4test_clstk`, along with some other files we will not need.
#
# We now build a few manifest files which will be used later:
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="AmR6CH025C8E" outputId="0cd776ea-078f-4ab8-8a79-eed3e1c05839"
# --- Building Manifest Files --- #
import json
import librosa
# Function to build a manifest
def build_manifest(transcripts_path, manifest_path, wav_path):
with open(transcripts_path, 'r') as fin:
with open(manifest_path, 'w') as fout:
for line in fin:
# Lines look like this:
# <s> transcript </s> (fileID)
transcript = line[: line.find('(')-1].lower()
transcript = transcript.replace('<s>', '').replace('</s>', '')
transcript = transcript.strip()
file_id = line[line.find('(')+1 : -2] # e.g. "cen4-fash-b"
audio_path = os.path.join(
data_dir, wav_path,
file_id[file_id.find('-')+1 : file_id.rfind('-')],
file_id + '.wav')
duration = librosa.core.get_duration(filename=audio_path)
# Write the metadata to the manifest
metadata = {
"audio_filepath": audio_path,
"duration": duration,
"text": transcript
}
json.dump(metadata, fout)
fout.write('\n')
# Building Manifests
print("******")
train_transcripts = data_dir + '/an4/etc/an4_train.transcription'
train_manifest = data_dir + '/an4/train_manifest.json'
build_manifest(train_transcripts, train_manifest, 'an4/wav/an4_clstk')
print("Training manifest created.")
test_transcripts = data_dir + '/an4/etc/an4_test.transcription'
test_manifest = data_dir + '/an4/test_manifest.json'
build_manifest(test_transcripts, test_manifest, 'an4/wav/an4test_clstk')
print("Test manifest created.")
print("******")
# + [markdown] colab_type="text" id="EQsXzh7x5zIQ"
# ## Prepare the path to manifest files
# + colab={} colab_type="code" id="vmOa0IRC5eW4"
dataset_basedir = os.path.join(data_dir, 'an4')
train_dataset = os.path.join(dataset_basedir, 'train_manifest.json')
test_dataset = os.path.join(dataset_basedir, 'test_manifest.json')
# + [markdown] colab_type="text" id="pz9LC3yZ6J1Q"
# ## Read a few rows of the manifest file
#
# Manifest files are the data structure used by NeMo to declare a few important details about the data :
#
# 1) `audio_filepath`: Refers to the path to the raw audio file <br>
# 2) `text`: The text transcript of this sample <br>
# 3) `duration`: The length of the audio file, in seconds.
# + colab={} colab_type="code" id="3OzZQiX751iz"
# !head -n 5 {train_dataset}
# + [markdown] colab_type="text" id="pD9bprV66Oai"
# # Data Augmentation Pipeline
#
# Constructing a data augmentation pipeline in NeMo is as simple as composing a nested dictionary that describes two things :
#
# 1) The probability of that augmentation occurring - using the `prob` keyword <br>
# 2) The keyword arguments required by that augmentation class
#
# Below, we show a few samples of these augmentations. Note, in order to distinguish between the original sample and the perturbed sample, we exaggerate the perturbation strength significantly.
# + colab={} colab_type="code" id="l5bc7gYO6MHG"
import torch
import IPython.display as ipd
# + [markdown] colab_type="text" id="L8Bd8s3e6TeK"
# ## Audio file preparation
# + colab={} colab_type="code" id="g7f9riZz6Qnj"
# Import the data augmentation component from ASR collection
from nemo.collections.asr.parts.preprocessing import perturb, segment
# + colab={} colab_type="code" id="wK8uwpt16d6I"
# Lets see the available perturbations
perturb.perturbation_types
# + [markdown] colab_type="text" id="IP1VpkOA6nE-"
# ### Obtain a baseline audio file
# + colab={} colab_type="code" id="sj4DNMmZ6ktm"
filepath = librosa.util.example_audio_file()
sample, sr = librosa.core.load(filepath)
ipd.Audio(sample, rate=sr)
# + [markdown] colab_type="text" id="M9mZNm296tNf"
# ### Convert to WAV format
# + colab={} colab_type="code" id="QDjlgLc-6vtq"
import soundfile as sf
# lets convert this ogg file into a wave to be compatible with NeMo
if not os.path.exists('./media'):
os.makedirs('./media/')
filename = 'Kevin_MacLeod_-_Vibe_Ace.wav'
filepath = os.path.join('media', filename)
sf.write(filepath, sample, samplerate=sr)
# + colab={} colab_type="code" id="FEkV-ikT6xgB"
sample, sr = librosa.core.load(filepath)
ipd.Audio(sample, rate=sr)
# + colab={} colab_type="code" id="gmuwEwIQ6zK3"
# NeMo has its own support class for loading wav files
def load_audio() -> segment.AudioSegment:
filename = 'Kevin_MacLeod_-_Vibe_Ace.wav'
filepath = os.path.join('media', filename)
sample_segment = segment.AudioSegment.from_file(filepath, target_sr=sr)
return sample_segment
sample_segment = load_audio()
ipd.Audio(sample_segment.samples, rate=sr)
# + [markdown] colab_type="text" id="hTnf1g1y63wZ"
# ## White Noise Perturbation
#
# White Noise perturbation is performed by the following steps : <br>
# 1) Randomly sample the amplitude of the noise from a uniformly distributed range (defined in dB) <br>
# 2) Sample gaussian noise (mean = 0, std = 1) with same length as audio signal <br>
# 3) Scale this gaussian noise by the amplitude (in dB scale) <br>
# 4) Add this noise vector to the original sample
#
# Notably, the original signal should not have a "hissing sound" constantly present in the perturbed version.
# + colab={} colab_type="code" id="2jaPQyUY65ij"
white_noise = perturb.WhiteNoisePerturbation(min_level=-50, max_level=-30)
# Perturb the audio file
sample_segment = load_audio()
white_noise.perturb(sample_segment)
ipd.Audio(sample_segment.samples, rate=sr)
# + [markdown] colab_type="text" id="2dfwesJU7DhN"
# ## Shift Perturbation
#
# Shift perturbation is performed by the following steps : <br>
# 1) Randomly sample the shift factor of the signal from a uniformly distributed range (defined in milliseconds) <br>
# 2) Depending on the sign of the shift, we shift the original signal to the left or the right. <br>
# 3) The boundary locations are filled with zeros after the shift of the signal <br>
#
# Notably, the perturbed signal below skips the first 25 to 50 seconds of the original audio below, and the remainder of the time is simply silence.
# + colab={} colab_type="code" id="2ONq8dBI7BZf"
shift = perturb.ShiftPerturbation(min_shift_ms=25000.0, max_shift_ms=50000.0)
# Perturb the audio file
sample_segment = load_audio()
shift.perturb(sample_segment)
ipd.Audio(sample_segment.samples, rate=sr)
# + [markdown] colab_type="text" id="kywA3h4T7G_S"
# ## Data Dependent Perturbations
#
# Some perturbations require an external data source in order to perturb the original sample. Noise Perturbation is a perfect example of one such augmentation that requires an external noise source dataset in order to perturb the original data.
# + [markdown] colab_type="text" id="eYm2DgGQ7KPe"
# ### Preparing a manifest of "noise" samples
# + colab={} colab_type="code" id="RXZ1o85E7FLT"
# Lets prepare a manifest file using the baseline file itself, cut into 1 second segments
def write_manifest(filepath, data_dir='./media/', manifest_name='noise_manifest', duration_max=None, duration_stride=1.0, filter_long=False, duration_limit=10.0):
if duration_max is None:
duration_max = 1e9
with open(os.path.join(data_dir, manifest_name + '.json'), 'w') as fout:
try:
x, _sr = librosa.load(filepath)
duration = librosa.get_duration(x, sr=_sr)
except Exception:
print(f"\n>>>>>>>>> WARNING: Librosa failed to load file {filepath}. Skipping this file !\n")
return
if filter_long and duration > duration_limit:
print(f"Skipping sound sample {filepath}, exceeds duration limit of {duration_limit}")
return
offsets = []
durations = []
if duration > duration_max:
current_offset = 0.0
while current_offset < duration:
difference = duration - current_offset
segment_duration = min(duration_max, difference)
offsets.append(current_offset)
durations.append(segment_duration)
current_offset += duration_stride
else:
offsets.append(0.0)
durations.append(duration)
for duration, offset in zip(durations, offsets):
metadata = {
'audio_filepath': filepath,
'duration': duration,
'label': 'noise',
'text': '_', # for compatibility with ASRAudioText collection
'offset': offset,
}
json.dump(metadata, fout)
fout.write('\n')
fout.flush()
print(f"Wrote {len(durations)} segments for filename {filename}")
print("Finished preparing manifest !")
# + colab={} colab_type="code" id="wLTT8jlP7NdU"
filename = 'Kevin_MacLeod_-_Vibe_Ace.wav'
filepath = os.path.join('media', filename)
# Write a "noise" manifest file
write_manifest(filepath, manifest_name='noise_1s', duration_max=1.0, duration_stride=1.0)
# + colab={} colab_type="code" id="izbdrSmd7PY5"
# Lets read this noise manifest file
noise_manifest_path = os.path.join('media', 'noise_1s.json')
# !head -n 5 {noise_manifest_path}
# + colab={} colab_type="code" id="82yq0TOV7Q_4"
# Lets create a helper method to load the first file in the train dataset of AN4
# Load the first sample in the manifest
def load_gsc_sample() -> segment.AudioSegment:
with open(train_dataset, 'r') as f:
line = f.readline()
line = json.loads(line)
gsc_filepath = line['audio_filepath']
sample_segment = segment.AudioSegment.from_file(gsc_filepath)
return sample_segment
gsc_sample_segment = load_gsc_sample()
ipd.Audio(gsc_sample_segment.samples, rate=16000)
# + [markdown] colab_type="text" id="zV9ypBqz7V9a"
# ## Noise Augmentation
#
# Noise perturbation is performed by the following steps : <br>
# 1) Randomly sample the amplitude scale of the noise sample from a uniformly distributed range (defined in dB) <br>
# 2) Randomly choose an audio clip from the set of noise audio samples available <br>
# 3) Compute the gain (in dB) required for the noise clip as compared to the original sample and scale the noise by this factor <br>
# 4) If the noise snippet is of shorter duration than the original audio, then randomly select an index in time from the original sample, where the noise snippet will be added <br>
# 5) If instead the noise snippet is longer than the duration of the original audio, then randomly subsegment the noise snippet and add the full snippet to the original audio <br>
#
# Notably, the noise perturbed sample should sound as if there are two sounds playing at the same time (overlapping audio) as compared to the original signal. The magnitude of the noise will be dependent on step (3) and the location where the noise is added will depend on steps (4) and (5).
# + colab={} colab_type="code" id="cjSXci1v7Tlg"
import random
rng = random.Random(0)
noise = perturb.NoisePerturbation(manifest_path=noise_manifest_path,
min_snr_db=-10, max_snr_db=-10,
max_gain_db=300.0, rng=rng)
# Perturb the audio file
sample_segment = load_gsc_sample()
noise.perturb(sample_segment)
ipd.Audio(sample_segment.samples, rate=16000)
# -
# ## RIR and Noise Perturbation
# RIR augmentation with additive foreground and background noise.
# In this implementation audio data is augmented by first convolving the audio with a Room Impulse Response
# and then adding foreground noise and background noise at various SNRs. RIR, foreground and background noises
# should either be supplied with a manifest file or as tarred audio files (faster).
# ### Prepare rir data and manifest
# This is where the rir data will be downloaded.
# Change this if you don't want the data to be extracted in the current directory.
rir_data_path = '.'
# !python ../../scripts/dataset_processing/get_openslr_rir_data.py --data_root {rir_data_path}
rir_manifest_path = os.path.join(rir_data_path, 'processed', 'rir.json')
# !head -n 3 {rir_manifest_path}
# ### Create RIR instance
rir = perturb.RirAndNoisePerturbation(rir_manifest_path=rir_manifest_path,
rir_prob=1,
noise_manifest_paths=[noise_manifest_path], # use noise_manifest_path from previous step
bg_noise_manifest_paths=[noise_manifest_path],
min_snr_db=[20], # foreground noise snr
max_snr_db=[20],
bg_min_snr_db=[20], # background noise snr
bg_max_snr_db=[20],
noise_tar_filepaths=[None], # `[None]` to indicates that noise audio files are not tar.
bg_noise_tar_filepaths=[None])
# ### Perturb the audio
sample_segment = load_gsc_sample()
rir.perturb(sample_segment)
ipd.Audio(sample_segment.samples, rate=16000)
# + [markdown] colab_type="text" id="kJjUkGJu7ern"
# ## Speed Perturbation
#
# Speed perturbation changes the speed of the speech, but does not preserve pitch of the sound. Try a few random augmentations to see how the pitch changes with change in duration of the audio file.
#
# **Note**: This is a very slow augmentation and is not advised to perform online augmentation for large datasets as it can dramatically increase training time.
# + colab={} colab_type="code" id="Ic-ziInU7ZKC"
resample_type = 'kaiser_best' # Can be ['kaiser_best', 'kaiser_fast', 'fft', 'scipy']
speed = perturb.SpeedPerturbation(sr, resample_type, min_speed_rate=0.5, max_speed_rate=2.0, num_rates=-1)
# Perturb the audio file
sample_segment = load_gsc_sample()
speed.perturb(sample_segment)
ipd.Audio(sample_segment.samples, rate=16000)
# + [markdown] colab_type="text" id="bhHX3dyh7jPq"
# ## Time Stretch Perturbation
#
# Time Stretch perturbation changes the speed of the speech, and also preserve pitch of the sound.
# Try a few random augmentations to see how the pitch remains close to the same with change in duration of the audio file.
# + [markdown] colab_type="text" id="8_kNSfcK7lfP"
# ### Note about speed optimizations
#
# Time stretch is a costly augmentation, and can easily cause training time to increase drastically. It is suggested that one installs the `numba` library using conda to use a more optimized augmentation kernel.
#
# ```python
# conda install numba
# ```
# + colab={} colab_type="code" id="Dpeb0QUZ7g3l"
time_stretch = perturb.TimeStretchPerturbation(min_speed_rate=0.5, max_speed_rate=2.0, num_rates=3)
# Perturb the audio file
sample_segment = load_gsc_sample()
time_stretch.perturb(sample_segment)
ipd.Audio(sample_segment.samples, rate=16000)
# + [markdown] colab_type="text" id="vhH1-Ga87rCX"
# # Augmentation Pipeline
#
# The augmentation pipeline can be constructed in multiple ways, either explicitly by instantiating the objects of these perturbations or implicitly by providing the arguments to these augmentations as a nested dictionary.
#
# We will show both approaches in the following sections
# + [markdown] colab_type="text" id="RC8_NOD97tlW"
# ## Explicit definition
# + [markdown] colab_type="text" id="UwWE7swo72WP"
# ### Instantiate the perturbations
# + colab={} colab_type="code" id="GdLYn0hx7pRU"
perturbations = [
perturb.WhiteNoisePerturbation(min_level=-90, max_level=-46),
perturb.GainPerturbation(min_gain_dbfs=0, max_gain_dbfs=50),
perturb.NoisePerturbation(manifest_path=noise_manifest_path,
min_snr_db=0, max_snr_db=50, max_gain_db=300.0)
]
# + [markdown] colab_type="text" id="CDSSbZ8w7zzR"
# ### Select chance of perturbations being applied
# + colab={} colab_type="code" id="NmoxfLSL7xPJ"
probas = [1.0, 1.0, 0.5]
# + [markdown] colab_type="text" id="wl0tnrMq79Jh"
# ### Prepare the audio augmentation object
# + colab={} colab_type="code" id="nO6T4U4f767o"
augmentations = list(zip(probas, perturbations))
audio_augmentations = perturb.AudioAugmentor(augmentations)
audio_augmentations._pipeline
# + [markdown] colab_type="text" id="9cgI9yUx8Cyv"
# ## Implicit definition
#
# Implicit definitions are preferred since they can be prepared in the actual configuration object.
# + colab={} colab_type="code" id="tiqrKFTM7_mH"
perturb.perturbation_types # Available perturbations
# + [markdown] colab_type="text" id="dbeXwLdw8VEc"
# ### Prepare the nested dictionary
# + colab={} colab_type="code" id="mbE0qEA98TRI"
audio_augmentations = dict(
white_noise = dict(
prob=1.0,
min_level=-90,
max_level=-46
),
gain = dict(
prob=1.0,
min_gain_dbfs=0,
max_gain_dbfs=50
),
noise = dict(
prob=0.5,
manifest_path=noise_manifest_path,
min_snr_db=0,
max_snr_db=50,
max_gain_db=300.0
)
)
audio_augmentations
# + [markdown] colab_type="text" id="tcsoCe9-8ZM9"
# ### Supply `augmentor` as an argument to the `model.train_ds` config
#
# Most of the common datasets used by ASR models support the keyword `augmentor` - which can include a nested dictionary defining the implicit definition of an augmentation pipeline.
#
# Note, all ASR models support implicit declaration of augmentations. This includes -
#
# 1) Speech To Label Models <br>
# 2) Speech To Text Models <br>
# 3) Speech To Text Models with BPE/WPE Support<br>
# + [markdown] colab_type="text" id="0WOJC0fdBL5J"
# # Training - Application of augmentations
#
# We will be describing the data loaders for a MatchboxNet model from the paper "[MatchboxNet: 1D Time-Channel Separable Convolutional Neural Network Architecture for Speech Commands Recognition](https://arxiv.org/abs/2004.08531)". The benefit of MatchboxNet over JASPER models is that they use Separable Convolutions, which greatly reduce the number of parameters required to get good model accuracy.
#
# <ins>Care must be taken not to apply augmentations to the test set!</ins>
#
# + colab={} colab_type="code" id="7iDWiIrzBzUA"
from omegaconf import OmegaConf
# + colab={} colab_type="code" id="yv3KWNjcAUnQ"
# We will download the MatchboxNet configuration file for either v1 or v2 dataset here
DATASET_VER = 1
if DATASET_VER == 1:
MODEL_CONFIG = "matchboxnet_3x1x64_v1.yaml"
else:
MODEL_CONFIG = "matchboxnet_3x1x64_v2.yaml"
if not os.path.exists(f"configs/{MODEL_CONFIG}"):
# !wget -P configs/ "https://raw.githubusercontent.com/NVIDIA/NeMo/$BRANCH/examples/asr/conf/matchboxnet/{MODEL_CONFIG}"
# + colab={} colab_type="code" id="vOcv0ri3BkmA"
# This line will load the entire config of the MatchboxNet model
config_path = f"configs/{MODEL_CONFIG}"
config = OmegaConf.load(config_path)
# + [markdown] colab_type="text" id="mLsyceMSCIHV"
# ### Augmentation in train set only
#
# Note how the train dataset config supports the `augmentor` implicit definition, however the test config does not.
#
# This is essential to avoid mistakenly performing Test Time Augmentation.
# + colab={} colab_type="code" id="VgUVm7lGB8Cz"
# Has `augmentor`
print(OmegaConf.to_yaml(config.model.train_ds))
# + colab={} colab_type="code" id="gURwQ2eyCE7o"
# Does not have `augmentor`
print(OmegaConf.to_yaml(config.model.test_ds))
# + [markdown] colab_type="text" id="_UV74AVlCo_m"
# # Custom Perturbations
#
# We can define and use custom perturbations as required simply by extending the `Perturbation` class.
#
# Let's look at how we can build a custom Noise Perturbation that we can use to evaluate the effect of noise at inference time, in order to measure the model's robustness to noise
#
# In evaluation mode, we want to set an explicit value for the `snr_db` parameter instead of uniformly sample it from a range. This allows us to control the signal to noise ratio without relying on randomness from the training implementation of `NoisePerturbation`.
#
# Further, we force a random seed in order to produce reproduceable results on the evaluation set.
#
# With this combination, we can easily evaluate each sample in the test set `S` times (`S` being the number of random seeds), and can evaluate each of these samples at `D` levels of Signal to Noise Ratio (in dB).
# + colab={} colab_type="code" id="Q9YBmBiZCbAX"
# We use a NeMo utility to parse the manifest file for us
from nemo.collections.common.parts.preprocessing import collections, parsers
class NoisePerturbationEval(perturb.Perturbation):
def __init__(
self, manifest_path=None, snr_db=40, max_gain_db=300.0, seed=None,
):
seed = seed if seed is not None else 0
self._manifest = collections.ASRAudioText(manifest_path, parser=parsers.make_parser([]))
self._snr_db = snr_db
self._max_gain_db = max_gain_db
self._rng = random.Random(seed)
# This is mostly obtained from the original NoisePerturbation class itself
def perturb(self, data):
snr_db = self._snr_db
noise_record = self._rng.sample(self._manifest.data, 1)[0]
noise = AudioSegment.from_file(noise_record.audio_file, target_sr=data.sample_rate)
noise_gain_db = min(data.rms_db - noise.rms_db - snr_db, self._max_gain_db)
# calculate noise segment to use
start_time = 0.0
if noise.duration > (start_time + data.duration):
noise.subsegment(start_time=start_time, end_time=start_time + data.duration)
# adjust gain for snr purposes and superimpose
noise.gain_db(noise_gain_db)
if noise._samples.shape[0] < data._samples.shape[0]:
noise_idx = data._samples.shape[0] // 2 # midpoint of audio
while (noise_idx + noise._samples.shape[0]) > data._samples.shape[0]:
noise_idx = noise_idx // 2 # half the initial starting point
data._samples[noise_idx: noise_idx + noise._samples.shape[0]] += noise._samples
else:
data._samples += noise._samples
# + [markdown] colab_type="text" id="qR8qiwSkC1eE"
# ## Registering augmentations
#
# We can use either approach to submit this test time augmentation to the Data Loaders.
#
# In order to obtain the convenience of the implicit method, we must register this augmentation into NeMo's directory of available augmentations. This can be done as follows -
# + colab={} colab_type="code" id="40Z4Fm88CxWA"
perturb.register_perturbation(name='noise_eval', perturbation=NoisePerturbationEval)
# + colab={} colab_type="code" id="jVVbRxb-C4hB"
# Lets check the registry of allowed perturbations !
perturb.perturbation_types
# + [markdown] colab_type="text" id="2fiHz6CdC-B1"
# ## Overriding pre-existing augmentations
#
# **Note**: It is not allowed to overwrite already registered perturbations using the `perturb.register_perturbation` method. It will raise a `ValueError` in order to prevent overwriting the pre-existing perturbation types
|
tutorials/asr/05_Online_Noise_Augmentation.ipynb
|