code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_braket
# language: python
# name: conda_braket
# ---
# # Basic Demo for D-Wave on Braket: The Minimum Vertex Cover Problem
# This tutorial discusses the ```BraketSampler``` and ```BraketDWaveSampler```. In essence, they are both doing the same thing; each one just accepts different parameter names. Specifically, the ```BraketDWaveSampler``` allows users familiar with D-Wave to use D-Wave parameter names, e.g., ```answer_mode```, whereas the ```BraketSampler``` parameter names are consistent with the rest of the Braket experience.
# __Minimum Vertex Cover problem__: Here we consider a well-known combinatorial optimization problem on graphs called the Minimum Vertex Cover problem. Given an undirected graph with a vertex set $V$ and an edge set $E$, a vertex cover is a subset of the vertices (nodes) such that each edge in the graph is incident to at least one vertex in the subset. The Minimum Vertex Cover problem seeks to find a cover with a minimum number of vertices in the subset.
import json
from braket.aws import AwsDevice
from braket.ocean_plugin import BraketSampler, BraketDWaveSampler
import matplotlib.pyplot as plt
import networkx as nx
import dwave_networkx as dnx
from dimod.binary_quadratic_model import BinaryQuadraticModel
from dwave.system.composites import EmbeddingComposite
# __NOTE__: Enter your S3 bucket and key below.
# Enter the S3 bucket you created during onboarding in the code below
my_bucket = "amazon-braket-Your-Bucket-Name" # the name of the bucket
my_prefix = "Your-Folder-Name" # the name of the folder in the bucket
s3_folder = (my_bucket, my_prefix)
# +
# session and device
# use the DW_2000Q device
device = AwsDevice("arn:aws:braket:::device/qpu/d-wave/DW_2000Q_6")
print('Device:', device)
# or use the Advantage_system4 device with over 5000 qubits to solve larger problems
device = AwsDevice("arn:aws:braket:::device/qpu/d-wave/Advantage_system4")
print('Device:', device)
# -
# ## HELPER FUNCTIONS
# helper function to get colors for every node
def get_colors(result, n):
"""
return list of colors=[0, 0, 1, 0, 1, 1, ...] for graph
"""
# Obtain colors of each vertex
colors = [0 for _ in range(n)]
for ii in range(n):
if ii in result:
colors[ii] = 1
return colors
# helper function to plot graph
def get_graph(graph, pos):
"""
plot graph with labels
"""
# positions for all nodes
# pos = nx.spring_layout(graph)
# nodes
nx.draw_networkx_nodes(graph, pos, node_size=400)
# edges
nx.draw_networkx_edges(graph, pos)
# labels
nx.draw_networkx_labels(graph, pos, font_weight='bold', font_color='w')
# plot the graph
plt.axis('off')
#plt.savefig("figures/random_graph.png") # save as png
plt.show();
# helper function to plot graph
def get_colored_graph(graph, pos, colors):
"""
plot colored graph for given solution
"""
# positions for all nodes
# pos = nx.spring_layout(graph)
colorlist = ['#377eb8', '#e41a1c']
nx.draw_networkx(graph, pos, node_color=[colorlist[colors[int(node)]] for node in graph.nodes],
node_size=400, font_weight='bold', font_color='w')
# plot the graph
plt.axis('off');
# plt.savefig("./figures/weighted_graph.png") # save as png
# ## BRAKET SAMPLER: Minimum Vertex Cover Problem
# A ```sampler```, as defined [here](https://docs.ocean.dwavesys.com/en/latest/glossary.html#term-sampler) and [here](https://docs.ocean.dwavesys.com/projects/system/en/stable/reference/samplers.html), accepts a binary quadratic model (```BQM```) and returns variable assignments. Samplers generally try to find minimizing values but can also sample from distributions defined by the ```BQM```: Samplers are processes that sample from low energy states of a problem’s objective function, which is a mathematical expression of the energy of a system. A binary quadratic model (BQM) sampler samples from low energy states in models such as those defined by an Ising equation or a QUBO problem and returns an iterable of samples, in order of increasing energy.
# ```BraketSampler``` is a structured sampler that uses Braket-formatted parameters and properties. For example, instead of ```answer_mode```, which is used for D-Wave QPU samplers, Braket uses ```resultFormat``` instead. Below is a sample example of solving the minimum vertex cover problem using ```BraketSampler```.
# +
# set sampler
sampler = BraketSampler(s3_folder, 'arn:aws:braket:::device/qpu/d-wave/DW_2000Q_6')
# EmbeddingComposite automatically maps the problem to the structure of the solver.
embedded_sampler = EmbeddingComposite(sampler)
# -
# ### Minimum Vertex Cover Problem: Star Graph
# +
# set graph
n = 6
graph = nx.star_graph(n-1)
# positions for all nodes
pos = nx.spring_layout(graph)
# plot graph with labels
get_graph(graph, pos)
# +
# run problem on D-Wave
result = dnx.min_vertex_cover(graph, embedded_sampler, resultFormat="HISTOGRAM")
print('Result to MVC problem:', result)
print('Size of the vertex cover:', len(result))
# get coloring
colors = get_colors(result, n)
# plot result
get_colored_graph(graph, pos, colors)
# -
# __Discussion__: For this specific star graph we just need one single vertex, the one at the center, to cover the entire set of edges. Clearly, this solution is the minimal vertex cover.
# ### Minimum Vertex Cover Problem: Erdos Renyi graph
# Let us consider a more complicated graph, from the family of random Erdoes-Renyi graphs. Such a graph can be readily generated using the ```networkx``` library. As input we set the desired number of vertices and edges connecting pairs of vertices.
# +
# setup Erdos Renyi graph
n = 10 # 10 nodes
m = 20 # 20 edges
# set graph
graph = nx.gnm_random_graph(n, m, seed=42)
# positions for all nodes
pos = nx.spring_layout(graph)
# plot graph with labels
get_graph(graph, pos)
# +
# run problem on D-Wave
result = dnx.min_vertex_cover(graph, embedded_sampler, resultFormat="HISTOGRAM")
print('Result to MVC problem:', result)
print('Size of the vertex cover:', len(result))
# get coloring
colors = get_colors(result, n)
# plot result
get_colored_graph(graph, pos, colors)
# -
# __Discussion__: By inspection, we can check that with the subset of blue-colored vertices we can reach every edge in the graph. This vertex cover is a subset of the vertices such that each edge in the graph is incident to at least one vertex in the subset. We have used the ```BraketSampler``` so far. Alternatively, we can use the ```BraketDWaveSampler```; this is just a matter of syntactic preferences.
# ## BRAKET D-WAVE SAMPLER: Minimum Vertex Cover Problem
# ```BraketDWaveSampler``` is a structured sampler that uses D-Wave-formatted parameters and properties. It is interchangeable with D-Wave's ```DWaveSampler```. Only the parameter inputs to the solver need to be changed to be D-Wave formatted (e.g. ```answer_mode``` instead of ```resultFormat```).
#
# Below is the same example as above of solving the minimum vertex cover problem. We now consider a larger problem with more nodes and edges, and solve the problem with the DWave advantage system.
# +
# setup Erdos Renyi graph
n = 100 # 100 nodes
m = 400 # 400 edges
# set graph
graph = nx.gnm_random_graph(n, m, seed=42)
# positions for all nodes
pos = nx.spring_layout(graph)
# plot graph with labels
get_graph(graph, pos)
# set sampler
sampler = BraketDWaveSampler(s3_folder,'arn:aws:braket:::device/qpu/d-wave/Advantage_system4')
# EmbeddingComposite automatically maps the problem to the structure of the solver.
embedded_sampler = EmbeddingComposite(sampler)
# +
# run problem on D-Wave: note replacement of 'resultFormat' with 'answer_mode'
result = dnx.min_vertex_cover(graph, embedded_sampler, answer_mode="histogram")
print('Result to MVC problem:', result)
print('Size of the vertex cover:', len(result))
# get coloring
colors = get_colors(result, n)
# plot result
get_colored_graph(graph, pos, colors)
# -
# ---
# ## APPENDIX
# +
# set sampler
sampler = BraketSampler(s3_folder,'arn:aws:braket:::device/qpu/d-wave/DW_2000Q_6')
# EmbeddingComposite automatically maps the problem to the structure of the solver.
embedded_sampler = EmbeddingComposite(sampler)
# setup Erdos Renyi graph
n = 15 # 10 nodes
m = 30 # 20 edges
# set graph
graph = nx.gnm_random_graph(n, m, seed=42)
# positions for all nodes
pos = nx.spring_layout(graph)
# plot graph with labels
get_graph(graph, pos)
# The below result should be 0 because node 0 is connected to the 4 other nodes in a star graph
result = dnx.min_vertex_cover(graph, embedded_sampler, resultFormat="HISTOGRAM")
print('Result to MVC problem:', result)
print('Size of the vertex cover:', len(result))
# +
# Obtain colors of each vertex
colors = [0 for _ in range(n)]
for ii in range(n):
if ii in result:
colors[ii] = 1
print(colors)
# -
# plot result
get_colored_graph(graph, pos, colors)
print(graph.edges)
colorlist = ['#e41a1c', '#377eb8', '#4daf4a', '#984ea3', '#ff7f00', '#ffff33', '#a65628', '#f781bf']
nx.draw_networkx(graph, pos, node_color=[colorlist[colors[int(node)]] for node in graph.nodes],
node_size=400, font_weight='bold', font_color='w')
# plot the graph
plt.axis('off');
# plt.savefig("./figures/weighted_graph.png") # save as png
# plt.show();
# ### APPENDIX: Packages
# Display our environmnet used for this demo.
# ! pip freeze
| examples/quantum_annealing/Dwave_MinimumVertexCoverProblem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: Binning process with sklearn Pipeline
# This example shows how to use a binning process as a transformation within a Scikit-learn Pipeline. A pipeline generally comprises the application of one or more transforms and a final estimator.
# +
from optbinning import BinningProcess
from sklearn.datasets import load_boston
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
# -
# To get us started, let’s load a well-known dataset from the UCI repository
# +
data = load_boston()
variable_names = data.feature_names
X = data.data
y = data.target
# -
variable_names
categorical_variables = ['CHAS']
# Instantiate a ``BinningProcess`` object class with variable names and the list of numerical variables to be considered categorical. Create pipeline object by providing two steps: a binning process transformer and a linear regression estimator.
binning_process = BinningProcess(variable_names,
categorical_variables=categorical_variables)
lr = Pipeline(steps=[('binning_process', binning_process),
('regressor', LinearRegression())])
# Split dataset into train and test Fit pipeline with training data.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
lr.fit(X_train, y_train)
# +
y_test_predict = lr.predict(X_test)
print("MSE: {:.3f}".format(mean_squared_error(y_test, y_test_predict)))
print("R2 score: {:.3f}".format(r2_score(y_test, y_test_predict)))
# -
# In this case, the performance metrics show that the binning process transformation is effective in improving predictions.
# +
lr2 = LinearRegression()
lr2.fit(X_train, y_train)
y_test_predict = lr2.predict(X_test)
print("MSE: {:.3f}".format(mean_squared_error(y_test, y_test_predict)))
print("R2 score: {:.3f}".format(r2_score(y_test, y_test_predict)))
# -
# #### Binning process statistics
# The binning process of the pipeline can be retrieved to show information about the problem and timing statistics.
binning_process.information(print_level=1)
# The ``summary`` method returns basic statistics for each binned variable.
binning_process.summary()
| doc/source/tutorials/tutorial_binning_process_sklearn_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="O5-sFs4SMVx5"
# # How to Create an Interactive Scatterplot with Altair
# > Rendering Vega visualization grammar in Jupyter Notebooks
#
# - toc: false
# - badges: true
# - comments: true
# - categories: [altair, jupyter, python]
# - author: <NAME>
# - image: images/altair_scatterplot.jpg
# - search_exclude: false
# - permalink: /altair-test
# + [markdown] colab_type="text" id="8Y1hUKSjMVx6"
# > Tip: Here are the required libraries and dataset.
# + colab={} colab_type="code" id="lE8kEcV7MVx7"
#collapse-hide
import pandas as pd
import altair as alt
movies = 'https://vega.github.io/vega-datasets/data/movies.json'
# + [markdown] colab_type="text" id="mrjZklKNMVx-"
# ## Example: Interactive Dropdown and Tooltips
#
# Chart taken from [this repo](https://github.com/uwdata/visualization-curriculum), specifically [this notebook](https://github.com/uwdata/visualization-curriculum/blob/master/altair_interaction.ipynb).
# + colab={"base_uri": "https://localhost:8080/", "height": 407} colab_type="code" id="aSzu9ou2MVx_" outputId="92a79829-d88d-4fdf-ce96-2a8068be861c"
#collapse-hide
df = pd.read_json(movies) # load movies data
genres = df['Major_Genre'].unique() # get unique field values
genres = list(filter(lambda d: d is not None, genres)) # filter out null values
genres.sort() # sort alphabetically
mpaa = ['G', 'PG', 'PG-13', 'R', 'NC-17', 'Not Rated']
# single-value selection over [Major_Genre, MPAA_Rating] pairs
# use specific hard-wired values as the initial selected values
selection = alt.selection_single(
name='Select',
fields=['Major_Genre', 'MPAA_Rating'],
init={'Major_Genre': 'Drama', 'MPAA_Rating': 'R'},
bind={'Major_Genre': alt.binding_select(options=genres), 'MPAA_Rating': alt.binding_radio(options=mpaa)}
)
# scatter plot, modify opacity based on selection
alt.Chart(movies).mark_circle().add_selection(
selection
).encode(
x='Rotten_Tomatoes_Rating:Q',
y='IMDB_Rating:Q',
tooltip=['Title:N', 'Release_Date:N', 'IMDB_Rating:Q', 'Rotten_Tomatoes_Rating:Q'],
opacity=alt.condition(selection, alt.value(0.75), alt.value(0.05))
)
# + colab={"base_uri": "https://localhost:8080/", "height": 407} colab_type="code" id="pCqjdgUIMVyB" outputId="89847177-c083-4946-fe31-68cbe1f6aa5f"
#hide
alt.Chart(movies).mark_circle().add_selection(
selection
).encode(
x='Rotten_Tomatoes_Rating:Q',
y='IMDB_Rating:Q',
tooltip=['Title:N', 'Release_Date:N', 'IMDB_Rating:Q', 'Rotten_Tomatoes_Rating:Q'],
opacity=alt.condition(selection, alt.value(0.75), alt.value(0.05))
)
| _notebooks/2020-04-28-altair-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Foreground-background separation using robust PCA
# %load_ext autoreload
# %autoreload 2
# To install PIL, look for the Pillow package on pip
from PIL import Image, ImageDraw, ImageFont
import numpy as np
import random as rand
from sklearn.decomposition import TruncatedSVD
# +
# Draw a kindergarten worthy background complete with a token sun and a house
im = Image.new( "RGB", (800, 600), '#666666' ) # the sky
ImageDraw.Draw( im ).rectangle( (0, 500, 800, 600), "#888888" ) # the ground
ImageDraw.Draw( im ).rectangle( (100, 400, 250, 500), "#bbbbbb" ) # the house
ImageDraw.Draw( im ).rectangle( (150, 450, 200, 500), "#444444" ) # the door
ImageDraw.Draw( im ).rectangle( (110, 420, 140, 450), "#666666" ) # the left window
ImageDraw.Draw( im ).rectangle( (210, 420, 240, 450), "#666666" ) # the right window
ImageDraw.Draw( im ).polygon( (80, 400, 270, 400, 175, 350), "#555555" ) # the roof
ImageDraw.Draw( im ).ellipse( (650, 50, 750, 150), "#bbbbbb" ) # the sun
# Create animated foreground with a ball bouncing around and a flying piece of text
# Warning: the method does not handle slow moving elements very well since they confuse
# the method into thinking that slow moving elements are background elements. To see this,
# decrease the speed of either the ball or the text by reducing the absolute value of
# the increments to posx, posy, posr and you may find residual (static) ghosts in the background
# The above is not very surprising though -- if a person is standing (absolutely) still in a
# video, would we call that person a part of the background or the foreground?
frames = []
posx = 850
posy = 300
posr = 0
numFrames = 20
for n in range( numFrames ):
frame = im.copy()
draw = ImageDraw.Draw( frame )
draw.ellipse( (posx, posy, posx + 50, posy + 50), "white" )
draw.multiline_text( (posr, posr), "CS771, IITK", "white", ImageFont.truetype("ariblk.ttf", 50) )
frames.append( frame )
posx -= 25
posy += 20
posr += 15
for n in range( numFrames ):
frame = im.copy()
draw = ImageDraw.Draw( frame )
draw.ellipse( (posx, posy, posx + 50, posy + 50), "white" )
draw.multiline_text( (posr, posr), "CS771, IITK", "white", ImageFont.truetype("ariblk.ttf", 50) )
frames.append( frame )
posx -= 25
posy -= 20
posr += 15
for n in range( numFrames ):
frame = im.copy()
draw = ImageDraw.Draw( frame )
draw.ellipse( (posx, posy, posx + 50, posy + 50), "white" )
draw.multiline_text( (posr, posr), "CS771, IITK", "white", ImageFont.truetype("ariblk.ttf", 50) )
frames.append( frame )
posx += 25
posy -= 20
posr += 15
for n in range( numFrames ):
frame = im.copy()
draw = ImageDraw.Draw( frame )
draw.ellipse( (posx, posy, posx + 50, posy + 50), "white" )
draw.multiline_text( (posr, posr), "CS771, IITK", "white", ImageFont.truetype("ariblk.ttf", 50) )
frames.append( frame )
posx += 25
posy += 20
posr += 15
frames[0].save( "bounce_cs771.gif", save_all = True, append_images = frames[1:], duration = 40, loop = 0 )
# -
# <img src = "http://www.cse.iitk.ac.in/users/purushot/tmp/cs771/bounce_cs771.gif" width = 300px align = "left">
# +
# Handling full color images is a pain since we would have to do foreground/background
# extraction separately for all three channels i.e. R, G, B. Easier to convert the GIF
# to a grayscale format so that there is only one channel to worry about
def convertToGrayscale( filename, newFilename ):
im = Image.open( filename )
newFrames = []
try:
while True:
newFrame = im.convert( mode = "L" )
newFrames.append( newFrame )
im.seek( im.tell() + 1 )
except EOFError:
pass
newFrames[0].save( newFilename, save_all = True, append_images = newFrames[1:], duration = im.info["duration"], loop = 0 )
# Dont forget to close files to prevent handle/memory leaks
im.close()
# Find out how many frames are there in this GIF animation video
def getLength( filename ):
im = Image.open( filename )
numFrames = 0
try:
while True:
numFrames += 1
im.seek( im.tell() + 1 )
except EOFError:
pass
# Dont forget to close files to prevent handle/memory leaks
im.close()
return numFrames
# Extract the frames from this GIF file and return all frames stacked in a matrix
def getData( filename ):
numFrames = getLength( filename )
im = Image.open( filename )
data = np.zeros( (numFrames, np.prod( np.array( im.size ) )) )
try:
while True:
frame = im.convert( mode = "L" )
data[im.tell()] = np.array( list( frame.getdata() ) )
im.seek( im.tell() + 1 )
except EOFError:
pass
# Dont forget to close files to prevent handle/memory leaks
im.close()
return data, numFrames
# +
# Retain only the top few entries in a matrix (by magnitude)
# and set everything else to zero
def applyHardThresholding( X, numPixels ):
(n, d) = X.shape
arr = X.reshape(-1)
idx = np.argsort( np.abs( X.reshape(-1) ) )[::-1]
XThresh = np.zeros( idx.shape )
XThresh[idx[:numPixels * n]] = arr[idx[:numPixels * n]]
return XThresh.reshape( (n,d) ), idx.reshape( (n,d) )
# Do "robust" PCA i.e. extract leading components even when the
# matrix may have corruptions (foreground elements)
# Warning: this is a highly simplified version of the algorithm
# For a more effective algorithm (also more involved), please see
# Netrapalli et al, Non-convex Robust PCA, NIPS 2014
# X: a matrix that we suspect is low rank but for some corruptions
# i.e. X = L + S where L is a low-rank matrix and S is a sparse matrix
# (sparse because there is not too much foreground in any frame)
# r: the rank we anticipate for L. For us r = 1 since background is static
# niterAltOpt: for how many iteration should we run alternating optimization?
# numFGPixels: a (rough) upper bound on the number of foreground pixels in any frame
def doRPCA( X, r = 1, niterAltOpt = 0, numFGPixels = 0 ):
(n, d) = X.shape
svd = TruncatedSVD( n_components = 1, n_iter = 10 )
# Extract the leading right singular vector of X
# If life were simple (for example if there had been no
# foreground at all), this would have been the background image
svd.fit( X )
L = svd.inverse_transform( svd.transform( X ) )
# Life is not so simple and the foreground elements will somewhat corrupt
# the background so extract the foreground by looking at pixels that differ
# the most from what we think is currently the background iamge
S, idx = applyHardThresholding( X - L, numFGPixels )
# Given an estimate of the foreground, do PCA to extract the background
# Then given the background, extract the background by hard thresholding
for t in range( niterAltOpt ):
svd = TruncatedSVD( n_components = r, n_iter = 10 )
clean = X - S
# May also perform SVD on a random set of data points to improve speed
# e.g. svd.fit( clean[np.random.choice(n, nSamples, replace = False)] )
svd.fit( clean )
L = svd.inverse_transform( svd.transform( clean ) )
(S, idx) = applyHardThresholding( X - L, numFGPixels )
return S, X - S
filename = "bounce_cs771.gif"
data, numFrames = getData( filename )
# Discard pixels that never vary in the entire video -- they are clearly background
# In the example we have taken, over 50% of the pixels never vary - discarding them
# from further processing will speed up our method
var = np.var( data, axis = 0 )
idxBackground = (var == 0)
goldBackground = np.mean( data[:, idxBackground], axis = 0 )
mixedData = data[:, ~idxBackground]
# Normalizing pixel values avoids very large values in computations
dataNorm = mixedData/256
# Alternating optimization is actually beneficial
# Try setting niterAltOpt = 0 and you will find ghost images in the background
foreground = np.zeros( data.shape )
background = np.zeros( data.shape )
# Put back in the background pixels we had removed
background[:,idxBackground] = goldBackground/256
(foreground[:, ~idxBackground], background[:, ~idxBackground]) = doRPCA( dataNorm, r = 1, niterAltOpt = 1, numFGPixels = 13000 )
# +
# Un-normalize the matrix entries and make them integers
# so that they make sense as pixel values
def cleanup( X ):
X = np.around( X * 256 )
X[X < 0] = 0
X[X > 255] = 255
X = X.astype(int)
return X
foreground = cleanup( foreground )
background = cleanup( background )
# +
# Save the extracted foreground and background images as GIF files
# so that we can enjoy the foreground and background as animations
newFrames = []
for i in range( numFrames ):
newFrame = Image.new( "L", im.size )
newFrame.putdata( foreground[i] )
newFrames.append( newFrame )
newFrames[0].save( 'foreground.gif', save_all = True, append_images = newFrames[1:], duration = 40, loop = 0 )
newFrames = []
for i in range( numFrames ):
newFrame = Image.new( "L", im.size )
newFrame.putdata( background[i] )
newFrames.append( newFrame )
newFrames[0].save( 'background.gif', save_all = True, append_images = newFrames[1:], duration = 40, loop = 0 )
# -
# <h3> Original </h3>
# <img src = "http://www.cse.iitk.ac.in/users/purushot/tmp/cs771/bounce_cs771.gif" width = 300px align = "left">
# <h3> Extracted Foreground and Background </h3>
# <img src = "http://www.cse.iitk.ac.in/users/purushot/tmp/cs771/foreground.gif" width = 300px align = "left"><img src = "http://www.cse.iitk.ac.in/users/purushot/tmp/cs771/background.gif" width = 300px align = "left">
| lecture_code/17_Principal Component Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:hodemulator]
# language: python
# name: conda-env-hodemulator-py
# ---
# I'm running a few sets of chains that are fitting the redMagic HOD to a shuffled SHAM, as well as SHAMs with assembly bias to see what happens. I want all the tests I want to do to be in one place. This notebook will do the following:
# * Load the chain and plot the contours
# * Calculate the MAP values
# * Load the original HOD and clustering for the SHAM sample of interest.
# * Populate the halo catalog and calculate its clustering.
# * Plot the best fit clustering against the clustering in the fit
# * Plot the best fit HOD to the true HOD of the sample
from matplotlib import pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set()
import numpy as np
from chainconsumer import ChainConsumer
from astropy.table import Table
from halotools.utils.table_utils import compute_conditional_percentiles
from halotools.mock_observables import hod_from_mock, wp, tpcf
from pearce.mocks import cat_dict
from pearce.mocks.customHODModels import *
from math import ceil
#fname = '/u/ki/swmclau2/des/PearceMCMC/100_walkers_1000_steps_chain_shuffled_sham_3.npy'
fname = '/u/ki/swmclau2/des/PearceMCMC/200_walkers_5000_steps_chain_shuffled_sham2.npy'
#fname = '/u/ki/swmclau2/des/PearceMCMC/200_walkers_5000_steps_chain_vpeak_sham_no_ab.npy'
#fname = '/u/ki/swmclau2/des/PearceMCMC/200_walkers_5000_steps_chain_vpeak_sham_no_ab_2.npy'
#fname = '/u/ki/swmclau2/des/PearceMCMC/200_walkers_5000_steps_chain_vpeak_sham_fscab.npy'
chain = np.loadtxt(fname )
chain.shape
n_walkers = 200
n_burn = 500
n_params = chain.shape[1]
# + active=""
# c = chain.reshape((n_walkers, -1, n_params), ord)
# chain = c[:,n_burn:, :].reshape((-1, n_params))
# -
chain = chain[n_burn*n_walkers:, :]
chain.shape
param_names = [r'$\log{M_{min}}$', r'$\log{M_0}$',r'$\sigma_{log{M}}$', r'$\log{M_1}$', r'$\alpha$']
#param_names = [r'$\log{M_{min}}$',r'$\mathcal{A}_{cen}$', r'$\log{M_0}$',r'$\sigma_{log{M}}$',\
# r'$\mathcal{B}_{sat}$', r'$f_{cen}$', r'$f_{sat}$',r'$\mathcal{A}_{sat}$', \
# r'$\mathcal{B}_{cen}$', r'$\log{M_1}$', r'$\alpha$']
#param_names = [r'$f_{sat}$',r'$\mathcal{A}_{sat}$',r'$f_{cen}$', r'$\mathcal{A}_{cen}$']
#param_names = [r'$\mathcal{A}_{sat}$',r'$\mathcal{A}_{cen}$']
c = ChainConsumer()
c.add_chain(chain, parameters=param_names, walkers=n_walkers)
c.configure(statistics='max')
gelman_rubin_converged = c.diagnostic.gelman_rubin()
print gelman_rubin_converged
# + active=""
# emulation_point = [('mean_occupation_centrals_assembias_param1',0.6),\
# ('mean_occupation_satellites_assembias_param1',-0.7),
# ('mean_occupation_centrals_assembias_split1',0.8),
# ('mean_occupation_satellites_assembias_split1',0.5)]
#
# em_params = dict(emulation_point)
# print em_params.keys()
# -
best_fit_vals = np.array([ 12.87364502, 12.23898854,0.53433088, 13.97462748, 1.04479171])
fig = c.plotter.plot(figsize=(8,8), parameters=param_names, truth = best_fit_vals)
fig.show()
fig = c.plotter.plot_distributions(figsize=(10,10) )
fig.show()
summary = c.analysis.get_summary()
MAP = np.array([summary[p][1] for p in param_names])
print MAP
print param_names
# +
PMASS = 591421440.0000001 #chinchilla 400/ 2048
halo_catalog = Table.read('/u/ki/swmclau2/des/AB_tests/abmatched_halos.hdf5', format = 'hdf5')
# +
shuffle_type = 'sh_shuffled'
mag_type = 'vpeak'
mag_key = 'halo_%s_mag'%(mag_type)
upid_key = 'halo_upid'
#mag_key = 'halo%s_%s_mag'%(shuffle_type, mag_type)
#upid_key = 'halo%s_upid'%shuffle_type
# +
cosmo_params = {'simname':'chinchilla', 'Lbox':400.0, 'scale_factors':[0.658, 1.0]}
cat = cat_dict[cosmo_params['simname']](**cosmo_params)#construct the specified catalog!
cat.load_catalog(1.0)
#cat.h = 1.0
# +
mag_cut = -21
min_ptcl = 200
halo_catalog = halo_catalog[halo_catalog['halo_mvir'] > min_ptcl*cat.pmass] #mass cut
galaxy_catalog = halo_catalog[ halo_catalog[mag_key] < mag_cut ] # mag cut
# -
def compute_mass_bins(prim_haloprop, dlog10_prim_haloprop=0.1):
lg10_min_prim_haloprop = np.log10(np.min(prim_haloprop))-0.001
lg10_max_prim_haloprop = np.log10(np.max(prim_haloprop))+0.001
num_prim_haloprop_bins = (lg10_max_prim_haloprop-lg10_min_prim_haloprop)/dlog10_prim_haloprop
return np.logspace(
lg10_min_prim_haloprop, lg10_max_prim_haloprop,
num=int(ceil(num_prim_haloprop_bins)))
mass_bins = compute_mass_bins(halo_catalog['halo_mvir'], 0.2)
mass_bin_centers = (mass_bins[1:]+mass_bins[:-1])/2.0
#mass_bin_centers = 10**((np.log10(mass_bins[1:])+np.log10(mass_bins[:-1]))/2.0)
# +
cen_mask = galaxy_catalog['halo_upid']==-1
mass_key = 'halo_mvir_host_halo'#
#mass_key = 'halo_%s_host_mvir'%shuffle_type
cen_hod_sham, _ = hod_from_mock(galaxy_catalog[cen_mask][mass_key],\
halo_catalog['halo_mvir'],\
mass_bins)
sat_hod_sham, _ = hod_from_mock(galaxy_catalog[~cen_mask][mass_key],\
halo_catalog['halo_mvir'],\
mass_bins)
# -
np.savetxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/cen_hod.npy', cen_hod_sham)
np.savetxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/sat_hod.npy', sat_hod_sham)
np.savetxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/mbc.npy', mass_bin_centers)
training_dir = '/u/ki/swmclau2/des/PearceLHC_wp_z_sham_free_split_no_rsd/'
# +
#rp_bins = np.loadtxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/rp_bins.npy')
#rp_bins = np.loadtxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/rpoints.npy')
rp_bins = np.loadtxt(training_dir+'a_1.00000/global_file.npy')
#rp_bins = np.logspace(-1.1, 1.6, 16)
#rpoints = np.loadtxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/rpoints.npy')
rpoints = (rp_bins[1:]+rp_bins[:-1])/2.0
#wp_sham = np.log10(np.loadtxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/sham_%s%s_wp.npy'%(mag_type, shuffle_type)))
# +
#np.savetxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/rp_bins.npy', rp_bins)
# -
cat.load_model(1.0, HOD='fscabRedMagic')
#cat.load_model(1.0, HOD=(FSAssembiasTabulatedCens, FSAssembiasTabulatedSats), hod_kwargs = {'prim_haloprop_vals': mass_bin_centers,
#cat.load_model(1.0, HOD=(HSAssembiasTabulatedCens, HSAssembiasTabulatedSats), hod_kwargs = {'prim_haloprop_vals': mass_bin_centers,
#'sec_haloprop_key': 'halo_%s'%(mag_type),
# 'cen_hod_vals':cen_hod_sham,
# 'sat_hod_vals':sat_hod_sham})
cat.model.param_dict
#summary = c.analysis.get_summary()
MAP = np.array([summary[p][1] for p in param_names])
#MAP = np.zeros((len(param_names),))
#MAP = chain[np.random.randint(chain.shape[0]), :]
print MAP
param_names
#names = ['mean_occupation_satellites_assembias_split1','mean_occupation_satellites_assembias_param1',
# 'mean_occupation_centrals_assembias_split1', 'mean_occupation_centrals_assembias_param1']
names = ['logMmin','logM0','sigma_logM', 'logM1', 'alpha']
#names = ['mean_occupation_satellites_assembias_param1', 'mean_occupation_centrals_assembias_param1']
params = dict(zip(names, MAP))
params['f_c'] = 1.0
cat.populated_once = False
cat.populate(params)
cat.model.param_dict
# +
cen_mask = cat.model.mock.galaxy_table['gal_type']=='centrals'
cen_hod_mock, _ = hod_from_mock(cat.model.mock.galaxy_table[cen_mask]['halo_mvir'],\
halo_catalog['halo_mvir'],\
mass_bins)
sat_hod_mock, _ = hod_from_mock(cat.model.mock.galaxy_table[~cen_mask]['halo_mvir'],\
halo_catalog['halo_mvir'],\
mass_bins)
# -
wp_hod = cat.calc_wp(rp_bins, pi_max = 40, do_jackknife=False, RSD=False)
print wp_hod
#wp_hod = cat.calc_xi(rp_bins, do_jackknife=False)
from halotools.mock_observables import return_xyz_formatted_array
# +
sham_pos = np.c_[galaxy_catalog['halo_x'],\
galaxy_catalog['halo_y'],\
galaxy_catalog['halo_z']]
wp_sham = wp(sham_pos*cat.h, rp_bins, 40.0*cat.h, period=cat.Lbox*cat.h, num_threads=1)
print wp_sham
#wp_sham = tpcf(sham_pos*h, rp_bins, period=400.0*h, num_threads=1)
#wp_sham = np.log10(wp(sham_pos, rp_bins, 40.0, period=400.0, num_threads=1))
# +
#np.savetxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/sham_%s_%s_wp.npy'%(mag_type, shuffle_type), wp_sham)
#np.savetxt('/nfs/slac/g/ki/ki18/des/swmclau2/AB_tests/sham_vpeak_shuffled_nd.npy',np.array([cat.calc_analytic_nd()]) )
# -
plt.plot(mass_bin_centers, cen_hod_sham+sat_hod_sham, label = 'SHAM')
plt.plot(mass_bin_centers, cen_hod_mock+sat_hod_mock, label = 'HOD Fit')
plt.legend(loc='best')
plt.xlabel(r"$\log{M_{vir}}$")
plt.ylabel(r"$\log{<N|M>}$")
plt.loglog()
plt.show();
# +
plt.plot(mass_bin_centers, sat_hod_sham, label = 'SHAM')
plt.plot(mass_bin_centers, sat_hod_mock, label = 'HOD')
plt.legend(loc='best')
plt.vlines(PMASS*min_ptcl,0, 100)
plt.loglog()
plt.show();
# +
plt.plot(mass_bin_centers, cen_hod_sham, label = 'SHAM')
plt.plot(mass_bin_centers, cen_hod_mock, label = 'HOD')
plt.legend(loc='best')
plt.vlines(PMASS*min_ptcl,0, 100)
plt.loglog()
plt.show();
# -
sham_nd = len(galaxy_catalog)/(cat.Lbox*0.7)**3
hod_nd = cat.calc_analytic_nd()
print sham_nd/hod_nd
plt.plot(rpoints, wp_sham, label = 'SHAM')
plt.plot(rpoints, wp_hod, label = 'HOD Fit')
plt.loglog()
#plt.xscale('log')
#plt.ylim([-0.2, 3.5])
plt.legend(loc='best')
plt.xlabel(r"$r_p$")
plt.ylabel(r"$w_p(r_p)$")
plt.show()
#plt.plot(rpoints, wp_sham, label = 'SHAM')
plt.plot(rpoints, wp_sham/wp_hod, label = 'HOD Fit')
plt.xscale('log')
#plt.xscale('log')
#plt.ylim([-0.2, 3.5])
plt.legend(loc='best')
plt.xlabel(r"$r_p$")
plt.ylabel(r"$w_p(r_p)$")
plt.show()
wp_hod
# +
plt.figure(figsize=(10,8))
#plt.plot(rpoints, wp_sham/wp_sham, label = 'SHAM')
plt.plot(rpoints, wp_hod/wp_sham, label = 'HOD Fit')
plt.xscale('log')
plt.legend(loc='best',fontsize = 15)
plt.xlim([1e-1, 30e0]);
#plt.ylim([1,15000])
plt.xlabel(r'$r$',fontsize = 15)
plt.ylabel(r'$\xi(r)$',fontsize = 15)
plt.show()
# -
print wp_sham/wp_hod
print wp_sham
print wp_hod
from pearce.emulator import OriginalRecipe, ExtraCrispy
# +
training_dir = '/u/ki/swmclau2/des/PearceLHC_wp_z_sham_emulator_no_rsd/'
em_method = 'gp'
split_method = 'random'
load_fixed_params = {'z':0.0}
emu = ExtraCrispy(training_dir,10, 2, split_method, method=em_method, fixed_params=load_fixed_params)
# -
for n in names:
print n, emu.get_param_bounds(n)
| notebooks/Compare Zheng-SHAM Fit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Benchmarking against the Wisconsin Diagnostic Breast Cancer (WDBC) Dataset
import os
import sys
sys.path.append(os.path.join(os.getcwd(), '../svm2plus/'))
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.svm import SVC
from svm2plus import SVC2Plus
from sklearn.datasets import load_breast_cancer
from sklearn.metrics import precision_recall_fscore_support, accuracy_score
data = load_breast_cancer()
X, y = load_breast_cancer(return_X_y=True)
# +
# Original parameter grid as proposed by Vapnik
param_grid_svc = [{'C': np.exp2(np.linspace(-5, 5, 21)),
'gamma': np.exp2(np.linspace(-6, 6, 25)),
'kernel': ['rbf']}]
# lmbda is a regularization parameter, just like C. So we CV it in the same way.
param_grid_svc2p = [{'C': np.exp2(np.linspace(-5, 5, 21)),
'lmbda': np.exp2(np.linspace(-5, 5, 21)),
'gamma': np.exp2(np.linspace(-6, 6, 25)),
'decision_kernel': ['rbf'],
'correcting_kernel': ['rbf']}]
# -
# ## Privileged Features = `mean`
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1618)
Z_train = X_train[:, :10]
X_train = X_train[:, 10:]
X_test = X_test[:, 10:]
# -
# Train SVC with grid searching
svc = GridSearchCV(SVC(), param_grid_svc, scoring='accuracy', return_train_score=True)
svc.fit(X_train, y_train);
precision_recall_fscore_support(y_test, svc.predict(X_test))
accuracy_score(y_test, svc.predict(X_test))
# +
# Train SVC2+ with grid searching
svc2p = GridSearchCV(SVC2Plus(), param_grid_svc2p, scoring='accuracy', return_train_score=True)
svc2p.fit(X=X_train, y=y_train, Z=Z_train)
preds = svc2p.predict(X_test)
precision_recall_fscore_support(y_test, preds)
# -
accuracy_score(y_test, preds)
# ## Privileged Features = `standard error`
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1618)
Z_train = X_train[:, 10:20]
X_train = np.hstack([X_train[:, :10], X_train[:, 20:]])
X_test = np.hstack([X_test[:, :10], X_test[:, 20:]])
# -
# Train SVC with grid searching
svc = GridSearchCV(SVC(), param_grid_svc, scoring='accuracy', return_train_score=True)
svc.fit(X_train, y_train);
precision_recall_fscore_support(y_test, svc.predict(X_test))
accuracy_score(y_test, svc.predict(X_test))
# +
# Train SVC2+ with grid searching
svc2p = GridSearchCV(SVC2Plus(), param_grid_svc2p, scoring='accuracy', return_train_score=True)
svc2p.fit(X=X_train, y=y_train, Z=Z_train)
preds = svc2p.predict(X_test)
precision_recall_fscore_support(y_test, preds)
# -
accuracy_score(y_test, preds)
# ## Privileged Features = `worst`
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1618)
Z_train = X_train[:, 20:]
X_train = X_train[:, :20]
X_test = X_test[:, :20]
# -
# Train SVC with grid searching
svc = GridSearchCV(SVC(), param_grid_svc, scoring='accuracy', return_train_score=True)
svc.fit(X_train, y_train);
precision_recall_fscore_support(y_test, svc.predict(X_test))
accuracy_score(y_test, svc.predict(X_test))
# +
# Train SVC2+ with grid searching
svc2p = GridSearchCV(SVC2Plus(), param_grid_svc2p, scoring='accuracy', return_train_score=True)
svc2p.fit(X=X_train, y=y_train, Z=Z_train)
preds = svc2p.predict(X_test)
precision_recall_fscore_support(y_test, preds)
# -
accuracy_score(y_test, preds)
| benchmark/benchmark_wdbc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Additive Regression
# $$f_c \sim \cal{GP}(0,k_c),\; \forall c \in [1..C]$$
# $$y^{(n)}|f_1...f_C,x^{(n)} = y^{(n)}|\sum_c f_c(x^{(n)}_c)$$
#
# Functions $f_c$ are all functions of separate covariates $x_c$
#
#
# ### Mean Field Posterior
#
# $$ p(f_1,...f_C|X,Y)=\prod_c p(f_c|u_c)q(u_c)$$
import tensorflow as tf
import os
import numpy as np
from tqdm import tqdm
from matplotlib import pyplot as plt
from matplotlib import cm
tf.logging.set_verbosity(tf.logging.ERROR)
from tensorflow.contrib.opt import ScipyOptimizerInterface as soi
np.random.seed(10)
# ## Simulating synthetic data
# +
lik = 'Poisson'
lik = 'Gaussian'
assert lik in ['Poisson','Gaussian']
#---------------------------------------------------
# Declaring additive model parameters
N =500
D = 3 # number of covariates
R = 1 # number of trials
fs = [lambda x:np.sin(x)**3, lambda x:np.cos(3.*x), lambda x:x]
f_indices = [[d] for d in range(D)]
C = len(f_indices) # number of additive terms
#---------------------------------------------------
# Simulating data
np_link,tf_link = np.exp, tf.exp
xmin,xmax=-3,3
X_np = np.random.uniform(xmin,xmax,(N,D))
F_np = np.hstack([fs[d](X_np[:,f_indices[d]]) for d in range(D)])
pred_np = np.sum(F_np,axis=1,keepdims=True)
Y_np = pred_np + np.random.randn(N,R)*.1
if lik == 'Gaussian':
Y_np = pred_np + np.random.randn(N,R)*.5
elif lik=='Poisson':
link = np.exp
rate = np.tile(link(pred_np),[1,R])
Y_np = np.random.poisson(rate,size=(N,R))
# -
colors_c = plt.cm.winter(np.linspace(0,1,C))
fig,ax = plt.subplots(1,C,figsize=(C*4,4))
for c in range(C):
i = f_indices[c]
if len(f_indices[c])==1:
o = np.argsort(X_np[:,f_indices[c]],0)
ax[c].plot(X_np[o,i],F_np[o,c],'-',color=colors_c[c])
ax[c].set_xlabel('$x_%d$'%i[0],fontsize=20)
ax[c].set_ylabel('$f_%d(x_%d)$'%(i[0],i[0]),fontsize=20)
elif len(f_indices[c])==2:
ax[c].scatter(X_np[:,i[0]],
X_np[:,i[1]],
c=F_np[:,c],linewidth=0)
ax[c].set_xlabel('$x_%d$'%i[0],fontsize=20)
ax[c].set_ylabel('$x_%d$'%i[1],fontsize=20)
ax[c].set_title('$f(x_%d,x_%d)$'%(i[0],i[1]),fontsize=20)
plt.suptitle('True underlying functions',y=1.05,fontsize=20)
fig.tight_layout()
plt.show()
# ## Constructing tensorflow model
# +
import sys
sys.path.append('../SVGPs')
from kernels import RBF
from likelihoods import Gaussian, Poisson, Gaussian_with_link
from settings import np_float_type,int_type
from model import SVAGP_DS
#---------------------------------------------------
# Constructing tensorflow model
X = tf.placeholder(tf.float32,[N,D])
Y = tf.placeholder(tf.float32,[N,R])
ks,Zs = [],[]
ks =[]
with tf.variable_scope("kernels") as scope:
for c in range(C):
with tf.variable_scope("kernel%d"%c) as scope:
input_dim = len(f_indices[c])
ks.append( RBF(input_dim,lengthscales=.5*np.ones(input_dim), variance=1.))
with tf.variable_scope("likelihood") as scope:
if lik=='Gaussian':
likelihood = Gaussian(variance=1)
elif lik == 'Poisson':
likelihood = Poisson()
with tf.variable_scope("ind_points") as scope:
for c in range(C):
with tf.variable_scope("ind_points%d"%c) as scope:
input_dim = len(f_indices[c])
Z_ = np.random.uniform(xmin,xmax,[10,input_dim]).astype(np_float_type)
Zs.append( tf.Variable(Z_,tf.float32,name='Z') )
with tf.variable_scope("model") as scope:
m= SVAGP_DS(X,Y,ks,likelihood,Zs,q_diag=True,f_indices=f_indices,n_samp=10)
# -
# ## Running inference and learning
# +
#---------------------------------------------------
sess = tf.Session()
sess.run(tf.global_variables_initializer()) # reset values to wrong
# declare loss
loss = -m.build_likelihood()
opt_global = tf.train.AdamOptimizer(1e-3).minimize(loss)
init = tf.global_variables_initializer()
sess.run(init) # reset values to wrong
feed_dic = {Y:Y_np, X:X_np}
#---------------------------------------------------
print('Running Optimization...')
nit = 8000
loss_array = np.zeros((nit,))
for it in tqdm(range( nit)):
sess.run(opt_global,feed_dict=feed_dic)
loss_array[it]= float(sess.run(loss, feed_dic))
# declare which optimization to perform
Fs_mean,Fs_var = sess.run(m.build_predict_fs(X), feed_dic)
pred_samples = m.sample_predictor(X)
pred_mean= sess.run(tf.reduce_mean(pred_samples,axis=-1) ,feed_dic)
Zs = sess.run(m.Zs, feed_dic)
sess.close()
print('Done')
# +
fig,axarr = plt.subplots(1,2,figsize=(8,4))
ax=axarr[0]
ax.plot(loss_array[:it], linewidth=3, color='blue')
ax.set_xlabel('iterations',fontsize=20)
ax.set_ylabel('Variational Objective',fontsize=20)
ax=axarr[1]
ax.plot(pred_mean,pred_np,'.',color='blue')
ax.plot([pred_mean.min(),pred_mean.max()],
[pred_mean.min(),pred_mean.max()],
'--',linewidth=2,color='grey')
ax.set_xlabel('True predictor',fontsize=20)
ax.set_ylabel('Predicted predictor',fontsize=20)
fig.tight_layout()
plt.show()
plt.close()
fig,ax = plt.subplots(1,C,figsize=(C*5,5))
for c in range(C):
i = f_indices[c]
if len(i)==1:
o = np.argsort(X_np[:,i],0)
f,s = Fs_mean[c,:,0],np.sqrt(Fs_var[c,:,0])
ax[c].vlines(Zs[c],ymin=f.min(),ymax=f.max(),alpha=.5,color=colors_c[c])
ax[c].plot(X_np[o,i],f[o],color=colors_c[c])
ax[c].fill_between(X_np[o,i].flatten(),
(f-s)[o].flatten(),
y2=(f+s)[o].flatten(),
alpha=.3,facecolor=colors_c[c])
ax[c].plot(X_np[o,i],F_np[o,c],'--',color=colors_c[c])
ax[c].set_xlabel('$x_%d$'%i[0],fontsize=20)
ax[c].set_ylabel('$f_%d(x_%d)$'%(i[0],i[0]),fontsize=20)
elif len(f_indices[c])==2:
ax[c].scatter(X_np[:,i[0]],
X_np[:,i[1]],
c=Fs_mean[c,:,0],linewidth=0)
ax[c].scatter(Zs[c][:,0],Zs[c][:,1],
c='r', marker=(5, 1))
ax[c].set_xlabel('$x_%d$'%i[0],fontsize=20)
ax[c].set_ylabel('$x_%d$'%i[1],fontsize=20)
ax[c].set_title('$f(x_%d,x_%d)$'%(i[0],i[1]),fontsize=20)
plt.suptitle('Inferred underlying functions',y=1.05,fontsize=20)
fig.tight_layout()
plt.show()
# -
| notebooks/demo_SVAGP-1d-MF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mbarbetti/unifi-physics-lab3/blob/main/toymc_lifetime.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="q826KbPaUCgY"
# Toy Monte Carlo per verificare il p-valore del test del $\chi^2$ di Pearson per una distribuzione esponenziale
# + id="CEyDnMmD6Jz8"
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
from scipy.stats import poisson
from scipy.stats import chi2
plt.rcParams['figure.figsize'] = [10, 5]
# + id="-BYLoS9VCw9c"
def chi2_calc(x_values, ncounts, ntot, mean):
# print (x_values, ncounts)
# print (ntot, mean)
### Calculate chi2 and print fit results
# delta = x_values[1:] - x_values[:-1]
# nexpect = ntot*delta*np.exp(-x_values[:-1]/mean)/mean
nexpect = ntot*(np.exp(-x_values[:-1]/mean)-np.exp(-x_values[1:]/mean))
# print (nexpect)
residuals = ncounts - nexpect
# print (residuals)
squares = np.square(residuals)/nexpect
# print (squares)
chi2fit = squares.sum()
return chi2fit
# + id="OggLyIPc6K33"
ntoy = 10000
true_mean = 2000
nevents = 5000
nbins = 10
max_time = 10000
chi2dist = np.zeros(ntoy)
for i in range(ntoy):
sample = np.random.exponential(true_mean, np.random.poisson(nevents))
histo = np.histogram(sample, bins = nbins, range = (0, max_time))
chi2mc = chi2_calc(histo[1], histo[0], nevents, true_mean)
if (i==0):
plt.xlabel('Time [ns]')
plt.ylabel('Number of events')
plt.hist ( sample, bins = nbins, range = (0, max_time), label='Simulated decay time')
plt.yscale("linear") #linear or log
plt.show()
# print(histo[1][:-1])
# print(histo[0])
# print (chi2mc)
chi2dist[i] = chi2mc
nchi2bins = 100
x_max = 2*nbins
ndf = nbins
binwidth = x_max/nchi2bins
x_values = np.arange (0, x_max, binwidth)
y_values = ntoy*binwidth*chi2.pdf(x_values,ndf)
#print (binwidth, x_values, y_values)
plt.plot(x_values, y_values, label=f'chi2 with ndf={ndf}', color = "red", zorder = 1, lw = 2)
plt.hist(chi2dist, bins = nchi2bins, range = (0,x_max), label='Toy mc (true mean)')
plt.legend()
plt.plot()
print ("valore medio del chi2 (true mean)= ", np.mean(chi2dist))
# Adesso calcolando la media dal campione
next_plot = True
if (next_plot):
chi2dist = np.zeros(ntoy)
for i in range(ntoy):
sample = np.random.exponential(true_mean, np.random.poisson(nevents))
histo = np.histogram(sample, bins = nbins, range = (0, max_time))
chi2mc = chi2_calc(histo[1], histo[0], nevents, np.mean(sample))
chi2dist[i] = chi2mc
plt.hist(chi2dist, bins = nchi2bins, range = (0,x_max), label='Toy mc (data mean)', alpha = 0.5)
plt.legend()
plt.plot()
print ("valore medio del chi2 (data mean)= ", np.mean(chi2dist))
| toymc_lifetime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ***
# ### Q1
# ***
# 1000 ~ 9999 까지(1000과 9999도 계산에 포함)의 네 자리 숫자 가운데에 '10'을 포함하는 숫자의 갯수는?
# #### Answer1
# ***
# ### Q2
# ***
# 10 ~ 99999 까지(10과 99999도 계산에 포함)의 숫자 가운데에 20의 배수이며 '080'을 포함하는 숫자의 갯수는?
# #### Answer2
# ***
# ### Q3
# ***
# d = {'Hospital':0, 'PostOffice':1, 'Phamacy':2, 'School':3, 'Home':4, 'Convenience':5, 'DepartmentStore':6, 'BeautySalon':7, 'Lotteria':8}는 사전 자료형이며,
# 각 element의 key는 건물 이름을 의미하고 value는 아래 'map'에서 건물의 위치를 의미한다.
# '철수'는 매일 집에서 09:00에 나와서 정확히 30분마다 인접한 건물로 이동한다고 했을 때, 18:00에 'Hospital'에 있을 확률 p(0.0 <= p <= 1.0)는 얼마인가?
# '철수'는 30분마다 꼭 인접한 건물로 이동해야하며(같은 건물에 30분을 초과하여 체류할 수 없음) 대각선에 위치한 건물로는 이동할 수 없다고 가정한다.
# | |map| |
# |:--:|:--:|:--:|
# |0 |1 |2 |
# |3 |4 |5 |
# |6 |7 |8 |
d = {'Hospital':0, 'PostOffice':1, 'Phamacy':2, 'School':3, 'Home':4, 'Convenience':5, 'DepartmentStore':6, 'BeautySalon':7, 'Lotteria':8}
print(d)
# #### Answer3
# 2020.09.10 Q1 ~ Q3 by <NAME>
| python3.6/Assignment01.ipynb |
% -*- coding: utf-8 -*-
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Octave
% language: octave
% name: octave
% ---
% # Einschrittverfahren (Blatt 1, Aufgabe 4)
%
% ## Einführung
% Die Lösung des Anfangswertproblems
% $$
% \dot{u}(t)=\cos(\frac{5}{3}t+1)+\frac{1}{2}u(t)+\frac{1}{2},\quad x(0)=-1/2
% $$
% soll mit numerischen Methoden bestimmt werden. Die rechte Seite der DGL sowie den Anfangwert definieren wir hier:
function udot=myrhs(u, t)
udot = cos(5/3*t+1) +0.5 * u + 0.5;
endfunction
u0 = [-0.5];
% Nun soll die Lösung des Anfangswertproblems dargestellt werden. Dazuz benutzen wir ´´lsode'
% +
% plot gnuplot
% addpath("../../basics")
% Berechne Referenzloesung (mit lsode)
tref = linspace(0.0, 5.0, 50);
uref = lsode(@myrhs, u0, tref);
% Darstellung des Richtungsfeld (25x25)
newplot()
hold on
title("Numerische Loesung von \dot u = f(u,t)")
xlabel("t")
ylabel("u(t)")
grid on
plot(tref, uref(:,1), 'color', 'black')
plot_richtungsfeld_1d(@myrhs, (t=linspace(0, 5, 25)), u=linspace(-3, 3, 25))
% -
% ## Definition der Einschrittverfahren
% Hier beginnt jetzt für Sie die Arbeit. Bitte implemtieren Sie die nachfolgenden Funktionen:
% + magic_args="Das explizite Eulerverfahren ist bereits vorgegeben ;-). "
function ueulerT=euler_explicit(f, u0, tvec)
% Initialisierung.
ueulerT=zeros(length(tvec), size(u0));
ueulerT(1,:) = u0';
% Zeitschritte.
for i=2:length(tvec)
# Zeitschrittweite.
tau = tvec(i) - tvec(i-1);
% Letzter Zeitpunkt und Wert.
tlast = tvec(i-1);
ulast = ueulerT(i-1,:)(:); % Spaltenvektor
% Wert im naechsten Zeitpunkt.
ueulerT(i,:) = ulast + tau*f(ulast, tlast);
end
endfunction
% + magic_args="TODO: Die nachfolgenden Verfahren sind zu implementieren. Bitte fuegen Sie hier eigenen Code ein."
% Heun-Verfahren (explizit)
function uheunT=heun_explicit(f, u0, tvec)
% Initialisierung.
uheunT=zeros(length(tvec), size(u0));
uheunT(1,:) = u0';
% Zeitschritte.
for i=2:length(tvec)
% TODO: ...
end
endfunction
% Runge-Kutta 4 (explizit)
function urk4T=rk4_explicit(f, u0, tvec)
% TODO: ...
endfunction
% Euler (implizit).
function ueulerT=euler_implicit(f, u0, tvec)
% TODO: ...
% ACHTUNG: Diese Funktion ist schwierig, da ein Gleichungssystem geloest werden muss.
% Verwenden Sie hierzu z.B. die Funktion "fsolve" (s. help fsolve). Diese findet Nullstellen einer gegebenen Funktion.
% Um "fsolve" zu verwenden, muessen Sie daher ggf. innerhalb von "euler_implicit" eine weitere Funktion mit der Defektgleichung implementieren.
endfunction
% -
% ## Test 1: Sind die Funktionen vorhanden?
% Ja, klar!
% +
% addpath("../../basics")
exist("euler_explicit")
exist("euler_implicit")
exist("heun_explicit")
exist("rk4_explicit")
% -
% ## Test 2: Erzeuge numerische Lösungen
% A) Numerische Loesungen
NSteps = 5
tdiskret = linspace(0.0, 5.0, NSteps + 1)
% ### a) Expliziter Euler
# TODO: Entfernen Sie das Semikolon, um Ausgaben sichtbar zu machen.
yeuler = euler_explicit(@myrhs, u0, tdiskret);
% ### b) Impliziter Euler
# TODO: Entfernen Sie das Semikolon, um Ausgaben sichtbar zu machen.
yeuler2 = euler_implicit(@myrhs, u0, tdiskret);
% ### c) Heun
# TODO: Entfernen Sie das Semikolon, um Ausgaben sichtbar zu machen.
yheun = heun_explicit(@myrhs, u0, tdiskret);
% ### d) RK4 explizit
# TODO: Entfernen Sie das Semikolon, um Ausgaben sichtbar zu machen.
yrk4 = rk4_explicit(@myrhs, u0, tdiskret);
% ## Test 3: Gemeinsamer Plot aller Lösungen
% +
%plot inline
% Plot von Richtungsfeld und Referenzlösung
newplot()
hold on
xlabel('t')
ylabel('u(t)')
grid on
plot_richtungsfeld_1d(@myrhs, (t=linspace(0, 5, 25)), u=linspace(-3, 3, 31))
plot(tref, uref(:,1), 'color', 'black')
% Eigene Loesungen
% TODO: Bitte ggf. ein-/auskommentieren!
plot(tdiskret, yeuler(:,1), 'color', 'blue')
plot(tdiskret(1:3), yeuler2(1:3,1), 'color', 'cyan')
plot(tdiskret, yheun(:,1), 'color', 'green')
plot(tdiskret, yrk4, 'color', 'red')
% -
help fsolve
| jupyter/templates/U01-einschrittverfahren/U01-einschrittverfahren.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Rock, Paper, Scissors
#
# In this week's exercise you will be working with TFDS and the rock-paper-scissors dataset. You'll do a few tasks such as exploring the info of the dataset in order to figure out the name of the splits. You'll also write code to see if the dataset supports the new S3 API before creating your own versions of the dataset.
# ## Setup
# + colab={} colab_type="code" id="TTBSvHcSLBzc"
# Use all imports
from os import getcwd
import tensorflow as tf
import tensorflow_datasets as tfds
# -
# ## Extract the Rock, Paper, Scissors Dataset
#
# In the cell below, you will extract the `rock_paper_scissors` dataset and then print its info. Take note of the splits, what they're called, and their size.
# + colab={} colab_type="code" id="KGsVrzy84WI2"
# EXERCISE: Use tfds.load to extract the rock_paper_scissors dataset.
filePath = f"{getcwd()}/../tmp2"
data, info = tfds.load("rock_paper_scissors",data_dir=filePath, with_info=True)
print(info)
# + colab={} colab_type="code" id="epPGTUqE5Z2E"
# EXERCISE: In the space below, write code that iterates through the splits
# without hardcoding any keys. The code should extract 'test' and 'train' as
# the keys, and then print out the number of items in the dataset for each key.
# HINT: num_examples property is very useful here.
for set in data:
print(set,":",info.splits[set].num_examples)
# EXPECTED OUTPUT
# test:372
# train:2520
# -
# ## Use the New S3 API
#
# Before using the new S3 API, you must first find out whether the `rock_paper_scissors` dataset implements the new S3 API. In the cell below you should use version `3.*.*` of the `rock_paper_scissors` dataset.
# + colab={} colab_type="code" id="Ms5ld5Ov6_OP"
# EXERCISE: In the space below, use the tfds.builder to fetch the
# rock_paper_scissors dataset and check to see if it supports the
# new S3 API.
# HINT: The builder should 'implement' something
rps_builder = tfds.builder("rock_paper_scissors:3.*.*",data_dir=filePath) # YOUR CODE HERE (Include the following arguments in your code: "rock_paper_scissors:3.*.*", data_dir=filePath)
print(rps_builder.version.implements(tfds.core.Experiment.S3))
# Expected output:
# True
# -
# ## Create New Datasets with the S3 API
#
# Sometimes datasets are too big for prototyping. In the cell below, you will create a smaller dataset, where instead of using all of the training data and all of the test data, you instead have a `small_train` and `small_test` each of which are comprised of the first 10% of the records in their respective datasets.
# + colab={} colab_type="code" id="QMGkJW6j7Ldl"
# EXERCISE: In the space below, create two small datasets, `small_train`
# and `small_test`, each of which are comprised of the first 10% of the
# records in their respective datasets.
small_train = tfds.load("rock_paper_scissors:3.*.*", split='train[:10%]',data_dir=filePath) # YOUR CODE HERE (Include the following arguments in your code: "rock_paper_scissors:3.*.*", data_dir=filePath)
small_test = tfds.load("rock_paper_scissors:3.*.*", split='test[:10%]',data_dir=filePath) # YOUR CODE HERE (Include the following arguments in your code: "rock_paper_scissors:3.*.*", data_dir=filePath)
# No expected output yet, that's in the next cell
# + colab={} colab_type="code" id="SOm99-zO_nAe"
# EXERCISE: Print out the size (length) of the small versions of the datasets.
print(len(list(small_train)))
print(len(list(small_test)))
# Expected output
# 252
# 37
# -
# The original dataset doesn't have a validation set, just training and testing sets. In the cell below, you will use TFDS to create new datasets according to these rules:
#
# * `new_train`: The new training set should be the first 90% of the original training set.
#
#
# * `new_test`: The new test set should be the first 90% of the original test set.
#
#
# * `validation`: The new validation set should be the last 10% of the original training set + the last 10% of the original test set.
# + colab={} colab_type="code" id="jL7KXYi17s_1"
# EXERCISE: In the space below, create 3 new datasets according to
# the rules indicated above.
new_train = tfds.load("rock_paper_scissors:3.*.*", split='train[:90%]',data_dir=filePath) # YOUR CODE HERE (Include the following arguments in your code: "rock_paper_scissors:3.*.*", data_dir=filePath)
print(len(list(new_train)))
new_test = tfds.load("rock_paper_scissors:3.*.*", split='test[:90%]',data_dir=filePath) # YOUR CODE HERE (Include the following arguments in your code: "rock_paper_scissors:3.*.*", data_dir=filePath)
print(len(list(new_test)))
validation = tfds.load("rock_paper_scissors:3.*.*", split='train[90%:] + test[90%:]',data_dir=filePath) # YOUR CODE HERE (Include the following arguments in your code: "rock_paper_scissors:3.*.*", data_dir=filePath)
print(len(list(validation)))
# Expected output
# 2268
# 335
# 289
# -
# # Submission Instructions
# +
# Now click the 'Submit Assignment' button above.
# -
# # When you're done or would like to take a break, please run the two cells below to save your work and close the Notebook. This frees up resources for your fellow learners.
# + language="javascript"
# <!-- Save the notebook -->
# IPython.notebook.save_checkpoint();
# + language="javascript"
# <!-- Shutdown and close the notebook -->
# window.onbeforeunload = null
# window.close();
# IPython.notebook.session.delete();
| Data Pipelines with TensorFlow Data Services/Week 1/Exercises/TFDS_Week1_Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib notebook
from colicoords import load, CellPlot, CellListPlot, IterCellPlot, iter_subplots, save
import matplotlib.pyplot as plt
import numpy as np
import os
from collections import namedtuple
from tqdm.auto import tqdm
from scipy.signal import medfilt
from skimage.feature import peak_local_max
c41_02 = load('c41_02_binary_opt.hdf5')
c41_03 = load('c41_03_binary_opt.hdf5')
# +
clp_c41_02 = CellListPlot(c41_02)
clp_c41_03 = CellListPlot(c41_03)
fig, ax = plt.subplots()
clp_c41_02.hist_intensity(ax=ax, data_name='g500', linewidth=0, label='c41_2')
clp_c41_03.hist_intensity(ax=ax, data_name='g500', linewidth=0, label='c41_3', alpha=0.75)
plt.savefig('intensity comparison.png')
plt.legend()
# -
storm_dtype = [('x', float), ('y', float), ('intensity', float), ('frame', int)]
def add_peakfind(cell, med=9, thd=7500, min_dst=5):
img = cell.data.data_dict['g500']
mf = medfilt(img, 9)
img_bg = img - mf
cell.data.add_data(img_bg, 'fluorescence', 'flu_mf')
peaks = peak_local_max(img_bg, min_distance=min_dst, threshold_abs=thd)
y, x = peaks.T
new_storm = np.empty(len(x), dtype=storm_dtype)
new_storm['x'] = x
new_storm['y'] = y
new_storm['intensity'] = np.ones_like(x)
new_storm['frame'] = np.ones_like(x)
cell.data.add_data(new_storm, 'storm', 'storm_thd_{}'.format(thd))
len(c41_02), len(c41_03)
# +
c41_02_new = c41_02.copy()
[add_peakfind(c) for c in tqdm(c41_02_new)]
c41_03_new = c41_03.copy()
[add_peakfind(c) for c in tqdm(c41_03_new)]
''
# +
icp = IterCellPlot(c41_02_new)
fig, axes = iter_subplots(2, 1, figsize=(8,6))
icp.imshow('g500', ax=axes[0])
icp.plot_storm(data_name='storm_thd_7500', ax=axes[0])
icp.imshow('flu_mf', ax=axes[1])
icp.plot_storm(data_name='storm_thd_7500', ax=axes[1])
plt.tight_layout()
fig.display()
# +
icp = IterCellPlot(c41_03_new)
fig, axes = iter_subplots(2, 1, figsize=(8,6))
icp.imshow('g500', ax=axes[0])
icp.plot_storm(data_name='storm_thd_7500', ax=axes[0])
icp.imshow('flu_mf', ax=axes[1])
icp.plot_storm(data_name='storm_thd_7500', ax=axes[1])
plt.tight_layout()
fig.display()
# +
labels = ['c41_02', 'c41_03']
fig, ax = plt.subplots()
nums = []
for cells in [c41_02_new, c41_03_new]:
num = [len(c.data.data_dict['storm_thd_7500']) for c in cells]
nums.append(num)
ax.hist(nums, bins = np.arange(15), label=labels, density=True)
ax.legend()
#fig.text(0.04, 0.5, 'Number of spots', va='center', rotation='vertical')
plt.ylabel('Fraction of cells')
plt.xlabel('Number of spots')
plt.savefig('spots per cell_c41 epec escc.png')
# -
save('c41_02_with_spots.hdf5', c41_02_new)
save('c41_03_with_spots.hdf5', c41_03_new)
| src_data/20191004_c41_eyfp-escv_AHT40_repeats/05_number_of_peaks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pymongo import MongoClient
from bson.objectid import ObjectId
import altair as alt
import pandas as pd
import math
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('seaborn-darkgrid')
# -
client = MongoClient('192.168.178.25', 27017)
db = client.TFE
collection = db.results
# # Analyse des résultats de tout les modèles en faisant varier le number de features de TF-IDF
# - experiment_id 24: sans SMOTE
# - experiment_id 25 : avec SMOTE
# # Analysis of results without SMOTE
results = pd.DataFrame(columns = ['model', 'max_features', 'type', 'recall', 'precision', 'f1'])
for result in collection.find({'experiment_id' : 24}, {'report' : 1, '_id' : 0, 'max_features' : 1, 'smote' : 1, 'downsampling' : 1}):
for model in result['report']:
results = results.append({'model' : model['model'], 'type' : 'fake', 'recall' : model['classification_report']['fake']['recall'], 'precision' : model['classification_report']['fake']['precision'], 'max_features' : result['max_features'], 'f1' : model['classification_report']['weighted avg']['f1-score']}, ignore_index=True)
results = results.append({'model' : model['model'], 'type' : 'reliable', 'recall' : model['classification_report']['reliable']['recall'], 'precision' : model['classification_report']['reliable']['precision'], 'max_features' : result['max_features'], 'f1' : model['classification_report']['weighted avg']['f1-score']}, ignore_index=True)
results
fp = open('tab.tex', 'w')
results.to_latex(fp)
fp.close()
# +
#results['max_features'] = results['max_features'].apply(lambda x: math.log10(x))
# +
fig, axes = plt.subplots(nrows = 1, ncols = 2, figsize=(15,5))
for key, grp in results[results['type'] == 'fake'].groupby(['model']):
axes[0].set_xscale("log", nonposx='clip')
axes[0].plot(grp['max_features'], grp['recall'], label=key, marker='o')
axes[0].set_title('Recall for positive class : fake')
#axes[0].xscale('log')
axes[0].legend()
for key, grp in results[results['type'] == 'reliable'].groupby(['model']):
axes[1].set_xscale("log", nonposx='clip')
axes[1].plot(grp['max_features'], grp['recall'], label=key, marker='o')
axes[1].set_title('Recall for positive class : reliable')
#axes[1].xscale('log')
axes[1].legend()
plt.savefig('output/ML_fake_recall.pdf')
# +
fig, axes = plt.subplots(nrows = 1, ncols = 2, figsize=(15,5))
for key, grp in results[results['type'] == 'fake'].groupby(['model']):
axes[0].plot(grp['max_features'], grp['precision'], label=key, marker='o')
axes[0].set_title('Precision for positive class : fake')
#axes[0].xscale('log')
axes[0].legend()
for key, grp in results[results['type'] == 'reliable'].groupby(['model']):
axes[1].plot(grp['max_features'], grp['precision'], label=key, marker='o')
axes[1].set_title('Precision for positive class : reliable')
#axes[1].xscale('log')
axes[1].legend()
plt.savefig('output/ML_fake_precision.pdf')
# -
results = pd.DataFrame(columns = ['model', 'recall', 'precision', 'max_features', 'f1'])
for result in collection.find({'experiment_id' : 24}, {'report' : 1, '_id' : 0, 'max_features' : 1, 'smote' : 1, 'downsampling' : 1}):
for model in result['report']:
results = results.append({'model' : model['model'], 'recall' : model['classification_report']['weighted avg']['recall'], 'precision' : model['classification_report']['weighted avg']['precision'], 'max_features' : result['max_features'], 'f1' : model['classification_report']['weighted avg']['f1-score']}, ignore_index=True)
results
fp = open('tab.tex', 'w')
results.to_latex(fp)
fp.close()
fig, axes = plt.subplots(nrows = 1, ncols = 3, figsize=(15,5))
for key, grp in results.groupby(['model']):
axes[0].plot(grp['max_features'], grp['f1'], label=key, marker='o')
axes[0].set_title('Weighted average F1-score')
#axes[0].xscale('log')
axes[0].legend()
axes[1].plot(grp['max_features'], grp['precision'], label=key, marker='o')
axes[1].set_title('Weighted average precision')
#axes[0].xscale('log')
axes[1].legend()
axes[2].plot(grp['max_features'], grp['recall'], label=key, marker='o')
axes[2].set_title('Weighted average recall')
#axes[0].xscale('log')
axes[2].legend()
plt.tight_layout()
plt.savefig('output/ML_fake_average.pdf')
# # Analysis of the results with SMOTE
results = pd.DataFrame(columns = ['model', 'type', 'recall', 'precision', 'max_features', 'f1'])
for result in collection.find({'experiment_id' : 25}, {'report' : 1, '_id' : 0, 'max_features' : 1, 'smote' : 1, 'downsampling' : 1}):
for model in result['report']:
results = results.append({'model' : model['model'], 'type' : 'fake', 'recall' : model['classification_report']['fake']['recall'], 'precision' : model['classification_report']['fake']['precision'], 'max_features' : result['max_features'], 'f1' : model['classification_report']['weighted avg']['f1-score']}, ignore_index=True)
results = results.append({'model' : model['model'], 'type' : 'reliable', 'recall' : model['classification_report']['reliable']['recall'], 'precision' : model['classification_report']['reliable']['precision'], 'max_features' : result['max_features'], 'f1' : model['classification_report']['weighted avg']['f1-score']}, ignore_index=True)
# +
fig, axes = plt.subplots(nrows = 1, ncols = 2, figsize=(15,5))
for key, grp in results[results['type'] == 'fake'].groupby(['model']):
axes[0].plot(grp['max_features'], grp['recall'], label=key, marker='o')
axes[0].set_title('Recall for positive class : fake')
#axes[0].xscale('log')
axes[0].legend()
for key, grp in results[results['type'] == 'reliable'].groupby(['model']):
axes[1].plot(grp['max_features'], grp['recall'], label=key, marker='o')
axes[1].set_title('Recall for positive class : reliable')
#axes[1].xscale('log')
axes[1].legend()
plt.savefig('output/ML_SMOTE_fake_recall.pdf')
# +
fig, axes = plt.subplots(nrows = 1, ncols = 2, figsize=(15,5))
for key, grp in results[results['type'] == 'fake'].groupby(['model']):
axes[0].plot(grp['max_features'], grp['precision'], label=key, marker='o')
axes[0].set_title('Precision for positive class : fake')
#axes[0].xscale('log')
axes[0].legend()
for key, grp in results[results['type'] == 'reliable'].groupby(['model']):
axes[1].plot(grp['max_features'], grp['precision'], label=key, marker='o')
axes[1].set_title('Precision for positive class : reliable')
#axes[1].xscale('log')
axes[1].legend()
plt.savefig('output/ML_SMOTE_fake_precision.pdf')
# -
results = pd.DataFrame(columns = ['model', 'recall', 'precision', 'max_features', 'f1'])
for result in collection.find({'experiment_id' : 25}, {'report' : 1, '_id' : 0, 'max_features' : 1, 'smote' : 1, 'downsampling' : 1}):
for model in result['report']:
results = results.append({'model' : model['model'], 'recall' : model['classification_report']['weighted avg']['recall'], 'precision' : model['classification_report']['weighted avg']['precision'], 'max_features' : result['max_features'], 'f1' : model['classification_report']['weighted avg']['f1-score']}, ignore_index=True)
fig, axes = plt.subplots(nrows = 1, ncols = 3, figsize=(15,5))
for key, grp in results.groupby(['model']):
axes[0].plot(grp['max_features'], grp['f1'], label=key, marker='o')
axes[0].set_title('Weighted average F1-score')
#axes[0].xscale('log')
axes[0].legend()
axes[1].plot(grp['max_features'], grp['precision'], label=key, marker='o')
axes[1].set_title('Weighted average precision')
#axes[0].xscale('log')
axes[1].legend()
axes[2].plot(grp['max_features'], grp['recall'], label=key, marker='o')
axes[2].set_title('Weighted average recall')
#axes[0].xscale('log')
axes[2].legend()
plt.tight_layout()
plt.savefig('output/ML_SMOTE_fake_average.pdf')
| Code/Analysis/analysis1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# Inpirations
# 1. https://www.kaggle.com/lextoumbourou/humpback-whale-id-data-and-aug-exploration
# 2. https://www.kaggle.com/sunnybeta322/what-am-i-whale-let-me-tell-you
# 3. https://www.kaggle.com/mmrosenb/whales-some-image-processing
# 4. https://www.kaggle.com/orangutan/keras-vgg19-starter
# 5. https://www.kaggle.com/gimunu/data-augmentation-with-keras-into-cnn/notebook
# Datasets
# You msut download the datasets from https://www.kaggle.com/c/whale-categorization-playground/data
# And locate them under "inputs" folder
import os
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import keras
from keras.models import Sequential
from keras.layers import Dense, Conv2D, MaxPooling2D, Flatten, Dropout
from keras.losses import categorical_crossentropy
from keras.optimizers import Adam
from sklearn.model_selection import train_test_split
import os
from tqdm import tqdm
from sklearn import preprocessing
import cv2
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
from subprocess import check_output
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# -
# ## Initiating Hyper Params
# +
pool_size = (2,2)
learning_rate = 0.07
num_of_epochs = 10
im_size = 128
training_dir = '../inputs/train/'
testing_dir = '../inputs/test/'
train_csv_path = '../inputs/train.csv'
batch_size = 64 # Reduce the batch size if you run out of GPU memory
epoch = 10
k_size = (4,4)
drop_probability = 0.5
hidden_size = 256
batch_size = 64
input_shape = (im_size, im_size)
X_train_check = './X_train.npy'
y_train_check = './y_train.npy'
X_test_check = './X_test.npy'
y_train_onehot_check = './y_train_onehot.npy'
# Utils
def process_image(file_dir, name):
fname = '{fdir}{fname}'.format(fdir=file_dir, fname=name)
x = cv2.imread(fname)
resized_x = cv2.resize(x, (im_size, im_size))
return resized_x / 255
# -
# ## Code
# +
# Placeholders
X_train = []
y_train = []
X_test = []
try:
X_train = np.load(X_train_check)
y_train = np.load(y_train_check)
X_test = np.load(X_test_check)
except:
print('Loading data from scratch')
df_train = pd.read_csv(train_csv_path)
# Loading training and label data
for file, label in tqdm(df_train.values):
img = process_image(training_dir, file)
X_train.append(img)
y_train.append(label)
# Loading testing data
test_files = []
for root, dirs, files in os.walk(testing_dir):
for filename in files:
test_files.append(filename)
for file in tqdm(test_files):
img = process_image(testing_dir, file)
X_test.append(img)
X_train = np.array(X_train)
y_train = np.array(y_train)
X_test = np.array(X_test)
np.save(X_train_check, X_train)
np.save(y_train_check, y_train)
np.save(X_test_check, X_test)
# -
# Checking shapes
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
try:
y_train = np.load(y_train_onehot_check)
except:
# Label one hot encoder
one_hot = pd.get_dummies(y_train, sparse = True)
one_hot_labels = np.asarray(one_hot)
np.save(y_train_onehot_check, one_hot_labels)
y_train = one_hot_labels
print(y_train.shape)
print(y_train[:5])
y_train_raw = np.array(y_train, np.uint8)
x_train_raw = np.array(X_train, np.float32)
x_test = np.array(X_test, np.float32)
print(y_train_raw.shape)
print(x_train_raw.shape)
print(x_test.shape)
num_classes = y_train_raw.shape[1]
print(num_classes)
X_train, X_valid, Y_train, Y_valid = train_test_split(x_train_raw, y_train_raw, test_size=0.3, random_state=1)
# +
model = Sequential()
input_shape = X_train_raw.shape[1:]
print('checking input_shape ', input_shape)
model.add(Convolution2D(32, kernel_size=k_size, activation="relu", input_shape=X_train_raw.shape[1:]))
model.add(MaxPooling2D(pool_size=pool_size, strides=(2,2)))
model.add(Convolution2D(64, kernel_size=k_size, activation="relu"))
model.add(MaxPooling2D(pool_size=pool_size, strides=(1,1)))
model.add(Convolution2D(512, kernel_size=k_size, activation="relu"))
model.add(MaxPooling2D(pool_size=pool_size, strides=(2,2)))
model.add(Flatten())
model.add(Dense(1024, activation="relu"))
model.add(Dropout(0.25))
model.add(Dense(512, activation="relu"))
model.add(Dense(32, activation="relu"))
model.add(Dropout(0.25))
model.add(Dense(num_classes, activation="softmax"))
# COST AND OPTIMIZER
model.compile(loss=categorical_crossentropy,
optimizer=Adam(lr=0.01),
metrics=['accuracy'])
model.summary()
# -
model.fit(
X_train,
Y_train,
validation_data=(X_valid, Y_valid),
batch_size=batch_size,
epochs=num_of_epochs,
verbose=1)
# Evaluate
scores = model.evaluate(X_valid, Y_valid, verbose=1)
print('Accuracy: {}'.format(scores[1] * 100))
| 009-cnn/challenge-humpback-whale-classification/src/Humpback Whale classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. 挖掘数据定义
# 遍历每一列, 阅读没类的描述确保理解这些数据表示的侧面, 并使用统计和绘图遍历列
# 可以查看其它data scientist的工作来理解每一列
# #### 2. 选择一个metric
# #### 3. Data Exploration and Data Cleaning
# 1. 此阶段要处理异常值和缺失值, 之后才能建模
# 2. 先查看数据标签的分布, 看数据是否平衡
# 如果变迁季度不平衡, 就要注意在交叉验证时使用分层抽样
# 3. 不同的数据类型表示不同的统计类型
# 1. float可能是连续值
# 2. int可能是boolean型或离散的有序变量
# 可通过查看int型列的值个数确定; train.select_dtypes('object').count_values()
# 3. object/categorical可能是标签
# train.select_dtypes('object').head()
# 4. 缺失值处理
# 计算每一列缺失值比例
# ```python
# import pandas as pd
# # Number of missing in each column
# missing = pd.DataFrame(data.isnull().sum()).rename(columns = {0: 'total'})
# # Create a percentage missing
# missing['percent'] = missing['total'] / len(data)
# ```
#
# #### 4. Feature Engineering
# 分为手动特征工程和自动特征工程(featuretools,十分之一的时间却有更好的效果)
# 手动特征工程:
# 1. 首先, 根据id查看每一列的统计属性
# ```python
# # Aggregate individual data for each household
# id_agg = ind.groupby('id').agg(['min', 'max', 'mean', 'sum'])
# 这会多出来4倍的统计属性, 而且多数存在线性相关性 , 因此需要做特征选择
# ```
# 2. 其次, 可以利用领域知识构建新特征
# ```python
# # No toilet, no electricity, no floor, no water service, no ceiling
# house['warning'] = 1 * (house['sanitario1'] +
# (house['elec'] == 0) +
# house['pisonotiene'] +
# house['abastaguano'] +
# (house['cielorazo'] == 0))
# ```
# 最好的方法是手动增加新特征后,再使用feature tools进行自动特征
#
# #### 5. Feature Selection
# 一种方法是查看两个特征之间的相关性, 相关系数高的叫做高线性相关
# ```python
# import numpy as np
#
# threshold = 0.95
#
# # Create correlation matrix
# corr_matrix = data.corr()
#
# # Select upper triangle of correlation matrix
# upper = corr_matrix.where(np.triu(np.ones(corr_matrix.shape), k=1).astype(np.bool))
#
# # Find index of feature columns with correlation greater than 0.95
# to_drop = [column for column in upper.columns if any(abs(upper[column]) > threshold)]
#
# data = data.drop(columns = to_drop)
#
# 还可以通过模型, 如随机森林来选择特征
# ```
| blog/Feature Engineer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Dynamically change surface plot data
#
# Note that in this case each time the dataset is transferred.
# (opposite to marching cubes plot)
# +
import k3d
import numpy as np
from ipywidgets import interact, interactive, fixed
import ipywidgets as widgets
import time
plot = k3d.plot()
Nx = 264
Ny = 264
xmin,xmax = -3,3
ymin,ymax = 0,3
x = np.linspace(xmin,xmax,Nx)
y = np.linspace(ymin,ymax,Ny)
x,y = np.meshgrid(x,y,indexing='ij')
surface = k3d.surface(np.sin(x**2+y**2),xmin=xmin,xmax=xmax,ymin=ymin,ymax=ymax,\
color=int(np.random.randint(0, 0xFFFFFF, 1)[0]))
plot += surface
plot.camera_auto_fit = False
plot.grid_auto_fit = False
# -
surface.heights = (np.sin(x**2)+np.cos(y**3))
surface.color=0xff0000
@interact(phi=widgets.FloatSlider(value=0.,min=0,max=2*np.pi,step=0.1))
def g(phi):
f = np.sin(x**2+y**2-phi)
surface.heights = f
plot.display()
# %time data = (np.sin(x**2)+np.cos(y**3))
# %time surface.heights = data+2
| examples/SageDays74/surface_interact.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from qutip import *
from scipy.optimize import minimize
from scipy.sparse.linalg import expm
# +
Hdrift = sigmaz().full()
Hctrls = [sigmax().full(),sigmay().full()]
init = fock(2,0).full()
target = fock(2,1).full()
dt = 1
lengthOfControl = 20
numberOfControls = len(Hctrls)
initControlAmplitudes = np.random.rand(numberOfControls, lengthOfControl)
def cost(controlAmplitudes):
"""Computes the cost function from the control signals.
Args:
controlAmplitudes (array):
An array with shape [numberOfControls, lengthOfControl].
It will be flattened by `scipy.optimize.minimize` to shape
(numberOfControls*lengthOfControl,) automatically.
Returns:
The cost (float). For example:
0.01
"""
U = qeye(2)
for index1 in range(lengthOfControl):
Ht = Hdrift
for index0 in range(numberOfControls):
index = index0 * lengthOfControl + index1
Ht += Hctrls[index0] * controlAmplitudes[index]
Ut = expm(-1j * Ht * dt)
U = np.matmul(Ut, U)
final = np.matmul(U, init)
cost = np.sum(np.abs(final - target))
return cost
minimizeResult = minimize(cost, initControlAmplitudes, method='L-BFGS-B')
# +
import matplotlib.pyplot as plt
x = minimizeResult.x
for index0 in range(numberOfControls):
plt.plot(x)
plt.show()
# -
| .ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf13
# language: python
# name: tf13
# ---
# # 实现过程中遇到的一些问题
# ## 已解决问题
# ### 训练的时候,各项损失都存在,但是正则项损失为 NaN
#
# 可能的原因有:
# 1. 检查学习率步长、正则项损失项的权重系数;
# 1. 网络中用到BatchNorm层,但是在训练一开始时,没有设置为训练阶段 keras.backend.set_learning_phase(True);
# 1. 检查图片:数据如果进行了预处理,检查是否有可能导致输入为NaN,例如空白图在除以方差进行归一化,则有可能NaN;
# 1. 检查标签数据:本来应该为正的标签,出现了负数(如bounding box标注点反了,导致宽/高为负数)。
#
# ### model.fit函数打印的总损失,不等于损失计算过程中记录的各项损失之和
#
# 我在`yolov3/yolov3_loss.py`的`YOLOv3Loss`类中定义了6个`tf.Variable(shape=(3,))`,分别对应了各个head的损失函数中 **前期校正损失项(rectified_coord_loss), 中心点坐标损失项(coord_loss_xy), 预测宽高损失项(coord_loss_wh), 背景损失项(noobj_iou_loss), 前景损失项(obj_iou_loss), 类别损失项(class_loss)**。然后在`utils/logger_callback.py`中调用了`self.model.losses`得到了模型的L2正则项。
#
# 如果日志打印中,总的损失函数值并不等于这些项的加和,有时大了,有时小了,因为总损失项做了一个历史平均的操作,而我打印的是当前迭代的子损失项的值。
# ## 遗留问题
# ### 加入条件分支获取校正损失项后,出现奇怪的错误
#
# 一开始实现前期计算anchor校正损失项时,我使用的是以下的实现代码逻辑(用迭代次数为条件判断是否需要计算校正损失项,如下代码`# 1`),但是不知道为什么一直报错。而如果把`# 1`的代码改为`# 2`处的代码,则不会报错。具体为什么我也没细究。
#
# ```python
# def loss(self, targets, predicts):
# # 0. train-from-scratch 前期(12800 pic)坐标回归时,将预测wh回归为anchor,预测xy回归为grid的中心点
# def get_rectified_coord_loss():
# nonlocal predicts
# with tf.control_dependencies([keras.backend.update_add(self.current_num, tf.shape(predicts)[0])]):
# predicts = tf.reshape(predicts, shape=[-1, self.height, self.width, self.box_num, self.box_len])
# rectified_loss = tf.reduce_sum(tf.square(predicts[:, :, :, :, 0:4]), axis=[1, 2, 3, 4])
# rectified_loss = tf.multiply(self.rectified_loss_weight, rectified_loss)
# rectified_loss = tf.reduce_mean(rectified_loss, keepdims=True)
# return rectified_loss
# # 前期矫正的图片数小于预定义的坐标校正图片数,则继续加坐标校正损失
# rectified_coord_loss = tf.cond(self.current_num <= self.rectified_coord_num,
# lambda: get_rectified_coord_loss(),
# lambda: tf.zeros(shape=(1,), dtype=tf.float32)) # 1
# # rectified_coord_loss = get_rectified_coord_loss() # 2
# # 1. 解码网络的输出
# decode_predicts, predicts_boxes = self.decoder.decode(predicts)
# # 2. 解码标签:(N, obj_num, gt_label_len)的标签矩阵, (N, obj_num, 4)的bounding boxes坐标
# targets, targets_boxes = self._decode_target(targets)
# # 3. 逐张图片计算损失函数,(N, 4),按样本维度遍历
# yolov2_loss = tf.map_fn(lambda inp: self._single_image_loss(inp[0], inp[1], inp[2], inp[3]),
# (targets, targets_boxes, decode_predicts, predicts_boxes), dtype=tf.float32,
# parallel_iterations=1)
# yolov2_loss = tf.reduce_mean(yolov2_loss, axis=0)
# # 4. 汇总并记录所有损失 (6,)
# update_op = [
# self.coord_loss_xy.assign(yolov2_loss[0]),
# self.coord_loss_wh.assign(yolov2_loss[1]),
# self.noobj_iou_loss.assign(yolov2_loss[2]),
# self.obj_iou_loss.assign(yolov2_loss[3]),
# self.class_loss.assign(yolov2_loss[4]),
# self.rectified_coord_loss.assign(rectified_coord_loss)
# ]
# # 前期矫正的图片数小于预定义的坐标校正图片数,则继续加坐标校正损失
# total_loss = tf.concat([yolov2_loss, rectified_coord_loss], axis=-1)
# # 4. 汇总所有损失
# with tf.control_dependencies(update_op):
# total_loss = tf.reduce_sum(total_loss)
# return total_loss
# ```
#
# 后面我直接把条件判断转移到`4 损失汇总`去了,然后也不会再报错:
#
# ```python
# # 前期矫正的图片数小于预定义的坐标校正图片数,则继续加坐标校正损失
# # [rectified_coord_loss, coord_loss_xy, coord_loss_wh, noobj_iou_loss, obj_iou_loss, class_loss]
# total_loss = tf.cond(self.current_num <= self.rectified_coord_num,
# lambda: tf.concat([yolov2_loss, get_rectified_coord_loss()], axis=-1),
# lambda: yolov2_loss)
# ```
| 1_learning_note/.ipynb_checkpoints/Question-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cleaning Sequences
# There are multiple ways of sequencing the steps in your data cleaning process. We've used one so far, once in Lesson 1 and once in the **Data Cleaning Process** example video in this lesson. The **Define**, **Code**, and **Test** Markdown headers were used once in this sequence, with multiple definitions, cleaning operations, and tests under each header, respectively.
#
# It looked like this:
# ______
# ## Gather
import pandas as pd
df = pd.read_csv('animals.csv')
# ## Assess
df.head()
# #### Quality
# - bb before every animal name
# - ! instead of . for decimal in body weight and brain weight
# ## Clean
df_clean = df.copy()
# #### Define
# - Remove 'bb' before every animal name using string slicing
# - Replace ! with . in body weight and brain weight columns
# #### Code
# Remove 'bb' before every animal name using string slicing
df_clean['Animal'] = df_clean['Animal'].str[2:]
# Replace ! with . in body weight and brain weight columns
df_clean['Body weight (kg)'] = df_clean['Body weight (kg)'].str.replace('!', '.')
df_clean['Brain weight (g)'] = df_clean['Brain weight (g)'].str.replace('!', '.')
# #### Test
df_clean.head()
# ________
# But you can also use multiple **Define**, **Code**, and **Test** headers, one for each data quality and tidiness issue (or group of data quality and tidiness issues). Effectively, you are defining then coding then testing immediately. This sequence is helpful when you have a lot of quality and tidiness issues to clean. Since that is the case in this lesson, this sequence will be used.
#
# Pasting each assessment above the **Define** header as its own header can also be helpful.
#
# Here's what this sequence looks like using the *animals.csv* dataset (and reusing the above *Gather* and *Assess* steps):
# Reload df_clean with dirty animals.csv
df_clean = df.copy()
# _______
# ## Clean
# #### bb before every animal name
# ##### Define
# Remove 'bb' before every animal name using string slicing.
# ##### Code
df_clean['Animal'] = df_clean['Animal'].str[2:]
# ##### Test
df_clean.Animal.head()
# # #### ! instead of . for decimal in body weight and brain weight
# ##### Define
# Replace ! with . in body weight and brain weight columns
# ##### Code
df_clean['Body weight (kg)'] = df_clean['Body weight (kg)'].str.replace('!', '.')
df_clean['Brain weight (g)'] = df_clean['Brain weight (g)'].str.replace('!', '.')
# ##### Test
df_clean.head()
| Part 4 Data Wrangling/study material/cleaning-sequences.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import folium
import json
import os
os.getcwd()
os.chdir("../")
print(os.getcwd())
survey = pd.read_stata("data/zuobiao_raw_frame.dta")
cn_province = pd.read_csv("data/province_cn.csv")
geodata= pd.read_json("data/cn_province.json")
# +
province = list()
for i in range(len(geodata['features'])):
province.append(geodata['features'][i]['properties']["NAME_1"])
geodata["province"] = province
geodata.head(5)
# -
cn_province.head(5)
# ## Data Wragling
# Drop oversea respondents and merge the dataset
survey = survey[survey.overseas == 0].reset_index().drop(columns="index")
data = pd.merge(survey, cn_province, left_on="provgb", right_on="GB")
data.head(5)
# +
# Create a dummy variable for having a college degree
data["college"] = np.where(data["educ"]>=3, 1, 0)
# Aggregate count, gender, and age
d = data[["GB", "id2"]].groupby("GB").count().merge(
cn_province, on="GB").drop(columns=["city", "country"])
d = data[["GB", "age"]].groupby("GB").mean().merge(d, on="GB")
d = data[["GB", "gender"]].groupby("GB").mean().merge(d, on="GB")
d = data[["GB", "college"]].groupby("GB").mean().merge(d, on="GB")
# rename the columns and display the data
d.columns = ["GB", "education", "gender", "age", "count", "lat", "lng", "province"]
d.head(5)
# -
# ## Drawing Map
# +
# Initiate the map
sv_map = folium.Map(
location=[d.lat.mean(), d.lng.mean()], tiles="cartodbpositron", zoom_start=4)
# Create a feature group
sv = folium.map.FeatureGroup(name="Survey Statistics")
# Draw boundaries
for segment in geodata['features']:
folium.GeoJson(data=segment['geometry'],
style_function=lambda x: {"color": "#000000", "weight": 1, 'fillOpacity': 0}).add_to(sv)
sv_map.add_child(sv)
# Draw circle on map representing the respondent sizes
for i in range(0, len(d)):
# Set up popup window
iframe = (folium.IFrame("<p style='color:blue;font-family:verdana;font-weight:bold'>" + str(d.iloc[i]['province'] + " Respondent Statistics") + '<p/>' +
"<ul style='font-family:verdana'>" +
'<li> Number of Responses: ' +
str(d.iloc[i]['count']) + '</li>'
'<li> Average Age of Respondents: ' +
str(np.round(d.iloc[i]['age'], 1)) + '</li>' +
'<li> Percentage of Male Respondants: ' +
str(np.round(d.iloc[i]['gender']* 100, 1)) + '</li>' +
'<li> Percentage of College Degree or Above: ' +
str(np.round(d.iloc[i]['education']* 100, 1)) + '</li> </ul>',
width=440, height=160))
popup = folium.Popup(iframe, max_width=400)
folium.CircleMarker(
location=[d.iloc[i]['lat'], d.iloc[i]['lng']],
popup=popup,
radius=np.sqrt(d.iloc[i]["count"])/5,
color='green',
alpha=0.4,
fill=True,
fill_color="green",
fill_opacity=0.4).add_to(sv)
sv_map.add_child(sv)
folium.LayerControl().add_to(sv_map)
# -
sv_map.save("survey_statistics.html")
| code/SurveyMap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # 机器学习简介
# -
# ## 1 如何分类
#
# 按任务类型
#
# - 回归模型:预测某个无法枚举的数值
# - 分类模型:将样本分为两类或多类
# - 结构化学习模型:输出不是向量而是其他结构,如给定长文本聚集成短的总结
#
# 按学习理论划分
#
# - 有监督学习:训练样本带有标签
# - 半监督学习:训练样本部分有标签;与使用所有标签数据相比,使用训练集的训练模型在训练时可以更为准确,而且训练成本更低
# - 无监督学习:训练样本全都没有标签,如聚类。根据样本间的相似性对样本集进行聚类试图使类内差距最小化,类间差距最大化
# - 强化学习:智能体(agent)通过与环境进行交互获得的奖赏来指导自己的行为,最终目标是使智能体获得最大的奖赏
# - 迁移学习:运用已存有的知识或者数据对不同担忧关联的领域问题进行求解;目的是通过迁移已有知识或者数据来解决目标领域中有标签样本数据比较少甚至没有的学习问题
#
# 判别式模型和生成式模型
# - 判别方法:由数据直接学习决策函数 $Y=f(X)$,或者由条件分布概率 $P(Y|X)$ 作为预测模型为判别模型
# - 常见判别模型:线性回归、boosting、SVM、决策树、感知机、线性判别分析 LDA、逻辑斯蒂回归等
# - 生成方法:由数据学习 $x$ 和 $y$ 的联合概率密度分布函数 $P(Y,X)$,然后通过贝叶斯公式求出条件概率分布 $P(Y|X)$ 作为预测的模型为生成模型
# - 常见生成模型:朴素贝叶斯、隐马尔科夫模型、高斯混合模型、文档主题生成模型 LDA 等
# ## 2 性能度量
#
# 衡量模型泛化能力的评价标准就是性能度量
#
# 对比不同模型的能力时,使用不同的性能度量往往会导致不同的判别结果
# ### 2.1 回归问题常用的性能度量指标
#
# 均方误差:$MSE=\displaystyle\frac{1}{n}\sum_{i=0}^{n}(f(x_i)-y_i)^2$
#
# 均方根误差(RMSE),观测值与真值偏差的平方和与观测次数 $m$ 比值的平方根,用于衡量观测值同真值之间的偏差:$RMSE=\sqrt{MSE}$
#
# 和方误差:$SSE=\displaystyle\sum_{i=0}^{n}(f(x_i)-y_i)^2$
#
# 平均绝对误差:$MAE=\displaystyle\frac{1}{n}\sum_{i=0}^{n}|f(x_i)-y_i|$
#
# 平均绝对百分比误差:$MPAE=\displaystyle\frac{1}{n}\sum_{i=0}^{n}\frac{|f(x_i)-y_i|}{y_i}$
#
# 平均平方百分比误差:$MASE=\displaystyle\frac{1}{n}\sum_{i=0}^{n}\left(\frac{|f(x_i)-y_i|}{y_i}\right)^2$
#
# 决定系数:$R^2=1-\displaystyle\frac{SSE}{SST}=1-\frac{\sum_{i=0}^{n}(f(x_i)-y_i)^2}{\sum_{i=0}^{n}(y_i-\bar{y})^2}$
# ### 2.2 分类问题常用的性能度量指标
#
# - TP,FN,FP,TN
# - TP,true positive,真正例,正样本被预测为正样本
# - FN,false negative,假负例,正样本被预测为负样本
# - FP,false positive,假正例,负样本被预测为正样本
# - TN,true negative,真负例,负样本被预测为负样本
# - 精确率(查准率) $Precision=\displaystyle\frac{TP}{TP+FP}$
# - 召回率(查全率) $Recall=\displaystyle\frac{TP}{TP+FN}$
# - 真正例率,正例被判断为正例的概率 $TPR=\displaystyle\frac{TP}{TP+FN}$
# - 假正例率,反例被判断为正例的概率 $FPR=\displaystyle\frac{FP}{TM+FP}$
# - $F_1$ 是精准率与召回率的调和平均值 $\displaystyle\frac{1}{F_1}=\frac{1}{2}\times\frac{1}{Precision}+\frac{1}{2}\times\frac{1}{Recall}$
# - 错误率与准确率
# - 错误率 $e=\displaystyle\sum_{i=1}^{n}I(f(x_i\neq y_i))$
# - 准确率 $acc=1-e$
# - AUC 与 ROC 曲线
# ## 3 特征工程
#
# 数据和特征决定了及学习的上限,而模型和算法只是逼近了这个上限而已
# ### 3.1 数据预处理
# #### 3.1.1 无量纲化
#
# 解决数据的量纲不同的问题,使不同的数据转换到同一规格
#
# (1)标准化
#
# 假设前提时特征值服从正态分布,标准化后将其转换为标准正态分布
#
# 在深度学习里,将数据标准化到一个特定的范围能够在反向传播中保证更好的收敛
#
# 如果不进行数据标准化,有些特征(值很大)将会对损失函数影响更大,使得其他值比较小的特征重要性降低
#
# 数据标准化可以使得每个特征的重要性更加均衡
#
# $$
# x'=\displaystyle\frac{x-\mu}{\sigma}
# $$
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from sklearn.preprocessing import StandardScaler
x1 = StandardScaler().fit_transform(x)
# + [markdown] pycharm={"name": "#%% md\n"}
# (2)归一化/区间缩放法
#
# 利用边界值信息,将特征的取值区间缩放到某个特点的范围,如 $[0,1]$ 等
#
# 适用于数据量较小的工程,利用两个最值进行缩放
#
# $$
# x'=\displaystyle\frac{x-\min}{\max - \min}
# $$
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from sklearn.preprocessing import MinMaxScaler
x1 = MinMaxScaler().fit_transform(x)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### 3.1.2 哑编码与独热编码
#
# 将离散特征的取值(如中国、美国、德国)扩展到了欧氏空间
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from sklearn.preprocessing import OneHotEncoder
# 独热编码
OneHotEncoder().fit_transform(x)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### 3.1.3 缺失值补充
#
# 常用方法:均值、就近补齐、$K-$最近距离填充等
#
# 有时候缺失值也是一种特征,可以补充一列将数据缺失与否赋值为0、1
#
# 缺失值太多,可以直接舍弃该列特征,否则可能会带来较大噪声
#
# 如果缺失值较少时(如少于$10\%$),可以考虑对缺失值进行填充
#
# 填充策略:
#
# 1. 用一个异常值填充(如0或-999)并将缺失值作为一个特征处理
# 2. 用均值或者条件均值填充
# 1. 数据是不平衡的,那么应该使用条件均值填充
# 2. 条件均值指与缺失值所属标签相同的所有数据的均值
# 3. 用相邻数据填充
# 4. 利用插值算法
# 5. 数据拟合
# 1. 将缺失值作为一个预测问题来处理
# 2. 将数据分为正常数据和缺失数据,对有值的数据采用随机森林等方法拟合,然后对有缺失值的数据用预测的值来填充
# -
# ### 3.2 特征选择
#
# 选入大量的特征不仅会降低模型效果,也会耗费大量的计算时间
# #### 3.2.1 方差选择法
#
# 若某列特征数值变化一直平缓,说明这个特征对结果的影响较小
#
# 可以计算各个特征的方差,选择方差大于自设阈值的特征
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from sklearn.feature_selection import VarianceThreshold
VarianceThreshold(threshold=10).fit_transform(x)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### 3.2.2 相关系数、统计检验
#
# pearson 相关系数,应用于连续变量
#
# 卡方检验,应用于离散变量
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from numpy import array
from sklearn.feature_selection import SelectKBest
# person 相关系数
from scipy.stats import pearsonr
SelectKBest(lambda X, Y: array(map(lambda x:pearsonr(x, Y), X.T)).T, k=5).fit_transform(x, y)
# 卡方检验
from sklearn.feature_selection import chi2
SelectKBest(chi2, k=5).fit_transform(x, y)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### 3.2.3 互信息法
#
# 评价自变量对因变量的相关性
#
# $$
# I(X;Y)=\displaystyle\sum_{x\in X}\sum_{y\in Y}p(x,y)\log\frac{p(x,y)}{p(x)p(y)}
# $$
# -
# #### 3.2.4 基于机器学习的特征选择法
#
# 针对特征和响应变量建立预测模型,例如用基于树的方法(决策树、随机森林、GBDT),或者扩展的线性模型
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from sklearn.feature_selection import SelectFromModel
from sklearn.ensemble import GradientBoostingClassifier
SelectFromModel(GradientBoostingClassifier()).fit_transform(x, y)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### 3.3 降维特征
#
# 若特征矩阵过大,会导致训练时间过长,需要降低特征矩阵维度
#
# 降维是通过保留重要的特征,减少数据特征的维度
#
# 特征的重要性取决于该特征能够表达多少数据集的信息,也取决于使用什么方法进行降维
#
# 降维的好处由节省存储空间,加快计算速度,避免模型过拟合等
# -
# #### 3.3.1 主成分分析法 PCA
#
# 将数据变换到一个新的坐标系统中的线性变换
#
# 主要目的是为让映射后得到的向量具有最大的不相关性
#
# PCA 追求的是在降维之后能够最大化保持数据的内在信息,并通过衡量在投影方向上的数据方差的大小来衡量该方向的重要性
#
# (1)计算相关系数矩阵
#
# $$
# R=
# \begin{bmatrix}
# r_{11} & ... & r_{1p} \\
# . & & . \\
# r_{p1} & ... & r_{pp} \\
# \end{bmatrix}
# $$
#
# $r_{ij}(i,j=1,2,...,p)$为原变量$x_i$与$y_j$的相关系数,其计算公式为
#
# $$
# r_{ij}=\displaystyle\frac{\sum_{k=1}^n(x_{ki}-\bar{x_i})(x_{kj}-\bar{x_j})}{(x_{ki}-\bar{x_i})^2(x_{kj}-\bar{x_j})^2}
# $$
#
# (2)计算特征值与特征向量
#
# 解特征方程 $|\lambda I-R|=0$,用雅可比法求出特征值,并使其按大小顺序排列 $\lambda_1\geq\lambda_2\geq ... \geq\lambda_p\geq 0$
#
# 特征值 $\lambda_i$ 对应的特征向量为 $e_i$,且 $||e||=1$
#
# (3)计算主成分贡献率及累计贡献率
#
# 对应的单位特征向量 $e_i$ 就是主成分 $z_i$ 的关于原变量的系数,即 $z_i=xe_{i}^{T}$
#
# 贡献率:$\alpha_i=\displaystyle\frac{\lambda_i}{\sum_{k=1}^p\lambda_k},i=1,2,...,p$
#
# 累计贡献率:$\displaystyle\frac{\sum_{k=1}^i\lambda_k}{\sum_{k=1}^p\lambda_k},i=1,2,...,p$
#
# 一般取累计贡献率达 $85\%\sim 95\%$ 的特征值 $\lambda_1,\lambda_2,...,\lambda_m$ 所对应的第1、2、...、第 $m(m\leq p)$ 个主成分 $z_1,z_2,...,z_m$
#
# (4)计算主成分载荷
#
# 主成分载荷是反映主成分 $z_i$ 与原变量 $x_j$ 之间的相互关联程度
#
# $$
# l_{ij}=p(z_i,x_j)=\sqrt{\lambda_i}e_{ij}(i,j=1,2,...,p)
# $$
#
# 将变量标准化后再计算其协方差矩阵
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from sklearn.decomposition import PCA
# k 为主成分的数目
PCA(n_components=k).fit_transform(x)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### 3.3.2 线性判别分析法 LDA
#
# -
# #### 3.3.3 局部线性嵌入 LLE
# ## 4. 过拟合、欠拟合与正则化
#
# 过拟合的表现是训练数据误差逐渐减小,验证集上误差增大
#
# 训练初期由于训练不足,学习器的拟合能力不够强,偏差比较大
# ### 4.1 过拟合与欠拟合的区别,什么是正则化
#
# 欠拟合,模型不能在训练集上获得足够低的训练误差,由于特征维度过少,导致拟合的函数无法满足训练集,导致误差较大
#
# 过拟合,模型的训练误差与测试误差(泛化误差)之间的差距过大
#
# 正则化,减少测试误差的策略,代价可能增大训练误差
# ### 4.2 解决欠拟合的方法
#
# 1. 加入新的特征
# 1. 深度学习,利用因子分解机、子编码器等
# 2. 增加模型复杂度
# 1. 线性模型增加高次项
# 2. 深度学习增加网络层数、神经元个数
# 3. 减少正则化项的系数
# ### 4.3 防止过拟合的方法
# #### 4.3.1 正则化
#
# $L_1$ 正则化,减少参数的绝对值总和,$||x||_1=\sum_{i}|x_i|$
#
# $L_2$ 正则化,减少参数平方的总和,$||x||_2=\sum_{i}x_i^2$
#
# 混合 $L_1$ 与 $L_2$ 正则化,$\alpha||x||_1+\displaystyle\frac{1-\alpha}{2}||x||_2$
# #### 4.3.2 Batch Normalization
#
# 作用是缓解梯度消失,加速网络的训练,防止过拟合,降低了参数初始化的要求
#
# 由于训练数据和测试数据的分布不同回降低模型的泛化能力,因此应该在开始训练前对所有数据做归一化处理
#
# Batch Normalization 会针对每一批数据在网络的每一层输入之前增加归一化处理,目的是为了使输入均值为 0,标准差为 1
#
# 利用公式表示就是针对每层的第 $k$ 个神经元,计算这一批数据在第 $k$ 个神经元的均值与标准差,然后将归一化后的值作为该神经元的激活值
#
# $$
# \hat{x_k}=\displaystyle\frac{x_k-E[x_k]}{\sqrt{var[x_k+\epsilon]}}
# $$
#
# Batch Normalization 通过对数据分布进行额外约束来增强模型的泛化能力,同时由于破坏了之前学到的特征分布,从而也降低了模型的拟合能力
#
# 为了恢复数据的原始分布,Batch Normalization 引入了一个重构变换来还原最优的输入数据分布
#
# $$
# y_k=BK(x_k)=\gamma\hat{x_k}+\beta
# $$
# #### 4.3.3 Dropout
#
# 对神经网络进行随即删减,降低网络复杂度
#
# 具体过程
#
# 1. 第一次迭代时随机删除一部分的隐藏单元,保持输入输出层不变,更新神经网络中的权值
# 2. 第二次迭代时随机删除掉一部分,和上一次删除的不一样
# 3. 第三次以及之后都是这样,直至训练结束
#
# 运用 Dropout 相当于训练了非常多个仅有部分隐层单元的神经网络,每个这种神经网络都能够给出一个分类结果,这些结果有的是正确的,有的是错误的
#
# 随着训练的进行,大部分网络都能够给出正确的分类结果
# #### 4.3.4 迭代截断
#
# 模型对训练数据集迭代收敛之前停止迭代防止过拟合
# #### 4.3.5 交叉验证
#
# $k-$fold 交叉验证是把训练样例分成 $k$ 份,然后进行 $k$ 次交叉验证过程,每次使用不同的一份样本作为验证集合,其余 $k-1$ 份样本合并作为训练集合
# ## 5 偏差与方差
#
# 偏差度量了学习算法的期望预测与真是结果的偏离程度
#
# 偏差用于描述模型的拟合能力,方差用于描述模型的稳定性
# ### 5.1 泛化误差、偏差、方差与噪声之间的关系
#
# $D$:训练集
#
# $y$:$x$ 真实的标记
#
# $y_D$:$x$ 在训练集 $D$ 中的标记
#
# $F$:通过训练集 $D$ 学到的模型
#
# $f(x;D)$:由训练集 $D$ 学到的模型 $f$ 对 $x$ 的预测输出
#
# $\bar{f}(x)$:模型 $f$ 对 $x$ 的期望预测输出
#
# 学习器在训练集上的误差称为“训练误差”或“经验误差”,在新样本上的误差被称为“泛化误差”
#
# 对于泛化误差,以回归任务为例,期望泛化误差为:$Err(x)=E[(y_D-f(x;D))^2]$
#
# 方差公式:$var(x)=E[(f(x;D)-\bar{f}(x))^2]$
#
# 噪声即为真实标记与数据集中的实际标记间的偏差,公式为:$\epsilon=E[(y_D-y)^2]$
#
# 假定噪声期望为 0,即 $E(y_D-y)=0$
#
# 偏差即为期望预测与真实标记的误差,偏差平方的公式为:$bias^2(x)=(\bar{f}(x)-y)^2$
#
# 则 $Err(x)=bias^2(x)+\epsilon+var(x)$,泛化误差可分解为偏差、方差和噪声之和
# ### 5.2 导致偏差和方差的原因
#
# 偏差是由于模型的复杂度不够或者对学习算法做了错误的假设
#
# 训练误差主要由偏差造成的
#
# 方差通常是由于模型的复杂度过高导致的
#
# 由方差引起的误差通常体现在测试误差相对训练误差的变化上
| ML/Notes/01-base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [[source]](../api/alibi.explainers.counterfactual.rst)
# # Counterfactual Instances
# <div class="alert alert-info">
# Note
#
# To enable support for counterfactual Instances, you may need to run
#
# ```bash
# pip install alibi[tensorflow]
# ```
# </div>
# ## Overview
# A counterfactual explanation of an outcome or a situation $Y$ takes the form "If $X$ had not occured, $Y$ would not have occured" ([Interpretable Machine Learning](https://christophm.github.io/interpretable-ml-book/counterfactual.html)). In the context of a machine learning classifier $X$ would be an instance of interest and $Y$ would be the label predicted by the model. The task of finding a counterfactual explanation is then to find some $X^\prime$ that is in some way related to the original instance $X$ but leading to a different prediction $Y^\prime$. Reasoning in counterfactual terms is very natural for humans, e.g. asking what should have been done differently to achieve a different result. As a consequence counterfactual instances for machine learning predictions is a promising method for human-interpretable explanations.
#
# The counterfactual method described here is the most basic way of defining the problem of finding such $X^\prime$. Our algorithm loosely follows Wachter et al. (2017): [Counterfactual Explanations without Opening the Black Box: Automated Decisions and the GDPR](https://arxiv.org/abs/1711.00399). For an extension to the basic method which provides ways of finding higher quality counterfactual instances $X^\prime$ in a quicker time, please refer to [Counterfactuals Guided by Prototypes](CFProto.ipynb).
#
# We can reason that the most basic requirements for a counterfactual $X^\prime$ are as follows:
#
# - The predicted class of $X^\prime$ is different from the predicted class of $X$
# - The difference between $X$ and $X^\prime$ should be human-interpretable.
#
# While the first condition is straight-forward, the second condition does not immediately lend itself to a condition as we need to first define "interpretability" in a mathematical sense. For this method we restrict ourselves to a particular definition by asserting that $X^\prime$ should be as close as possible to $X$ without violating the first condition. The main issue with this definition of "interpretability" is that the difference between $X^\prime$ and $X$ required to change the model prediciton might be so small as to be un-interpretable to the human eye in which case [we need a more sophisticated approach](CFProto.ipynb).
#
# That being said, we can now cast the search for $X^\prime$ as a simple optimization problem with the following loss:
#
# $$L = L_{\text{pred}} + \lambda L_{\text{dist}},$$
#
# where the first loss term $L_{\text{pred}}$ guides the search towards points $X^\prime$ which would change the model prediction and the second term $\lambda L_{\text{dist}}$ ensures that $X^\prime$ is close to $X$. This form of loss has a single hyperparameter $\lambda$ weighing the contributions of the two competing terms.
#
# The specific loss in our implementation is as follows:
#
# $$L(X^\prime\vert X) = (f_t(X^\prime) - p_t)^2 + \lambda L_1(X^\prime, X).$$
#
# Here $t$ is the desired target class for $X^\prime$ which can either be specified in advance or left up to the optimization algorithm to find, $p_t$ is the target probability of this class (typically $p_t=1$), $f_t$ is the model prediction on class $t$ and $L_1$ is the distance between the proposed counterfactual instance $X^\prime$ and the instance to be explained $X$. The use of the $L_1$ distance should ensure that the $X^\prime$ is a sparse counterfactual - minimizing the number of features to be changed in order to change the prediction.
#
# The optimal value of the hyperparameter $\lambda$ will vary from dataset to dataset and even within a dataset for each instance to be explained and the desired target class. As such it is difficult to set and we learn it as part of the optimization algorithm, i.e. we want to optimize
#
# $$\min_{X^{\prime}}\max_{\lambda}L(X^\prime\vert X)$$
#
# subject to
#
# $$\vert f_t(X^\prime)-p_t\vert\leq\epsilon \text{ (counterfactual constraint)},$$
#
# where $\epsilon$ is a tolerance parameter. In practice this is done in two steps, on the first pass we sweep a broad range of $\lambda$, e.g. $\lambda\in(10^{-1},\dots,10^{-10}$) to find lower and upper bounds $\lambda_{\text{lb}}, \lambda_{\text{ub}}$ where counterfactuals exist. Then we use bisection to find the maximum $\lambda\in[\lambda_{\text{lb}}, \lambda_{\text{ub}}]$ such that the counterfactual constraint still holds. The result is a set of counterfactual instances $X^\prime$ with varying distance from the test instance $X$.
# ## Usage
# ### Initialization
# The counterfactual (CF) explainer method works on fully black-box models, meaning they can work with arbitrary functions that take arrays and return arrays. However, if the user has access to a full TensorFlow (TF) or Keras model, this can be passed in as well to take advantage of the automatic differentiation in TF to speed up the search. This section describes the initialization for a TF/Keras model, for fully black-box models refer to [numerical gradients](#Numerical-Gradients).
#
# First we load the TF/Keras model:
#
# ```python
# model = load_model('my_model.h5')
# ```
#
# Then we can initialize the counterfactual object:
#
# ```python
# shape = (1,) + x_train.shape[1:]
# cf = Counterfactual(model, shape, distance_fn='l1', target_proba=1.0,
# target_class='other', max_iter=1000, early_stop=50, lam_init=1e-1,
# max_lam_steps=10, tol=0.05, learning_rate_init=0.1,
# feature_range=(-1e10, 1e10), eps=0.01, init='identity',
# decay=True, write_dir=None, debug=False)
# ```
#
# Besides passing the model, we set a number of **hyperparameters** ...
#
# ... **general**:
#
# * `shape`: shape of the instance to be explained, starting with batch dimension. Currently only single explanations are supported, so the batch dimension should be equal to 1.
#
# * `feature_range`: global or feature-wise min and max values for the perturbed instance.
#
# * `write_dir`: write directory for Tensorboard logging of the loss terms. It can be helpful when tuning the hyperparameters for your use case. It makes it easy to verify that e.g. not 1 loss term dominates the optimization, that the number of iterations is OK etc. You can access Tensorboard by running `tensorboard --logdir {write_dir}` in the terminal.
#
# * `debug`: flag to enable/disable writing to Tensorboard.
#
# ... related to the **optimizer**:
#
# * `max_iterations`: number of loss optimization steps for each value of $\lambda$; the multiplier of the distance loss term.
#
# * `learning_rate_init`: initial learning rate, follows linear decay.
#
# * `decay`: flag to disable learning rate decay if desired
#
# * `early_stop`: early stopping criterion for the search. If no counterfactuals are found for this many steps or if this many counterfactuals are found in a row we change $\lambda$ accordingly and continue the search.
# * `init`: how to initialize the search, currently only `"identity"` is supported meaning the search starts from the original instance.
#
#
# ... related to the **objective function**:
#
# * `distance_fn`: distance function between the test instance $X$ and the proposed counterfactual $X^\prime$, currently only `"l1"` is supported.
#
# * `target_proba`: desired target probability for the returned counterfactual instance. Defaults to `1.0`, but it could be useful to reduce it to allow a looser definition of a counterfactual instance.
#
# * `tol`: the tolerance within the `target_proba`, this works in tandem with `target_proba` to specify a range of acceptable predicted probability values for the counterfactual.
#
# * `target_class`: desired target class for the returned counterfactual instance. Can be either an integer denoting the specific class membership or the string `other` which will find a counterfactual instance whose predicted class is anything other than the class of the test instance.
#
# * `lam_init`: initial value of the hyperparameter $\lambda$. This is set to a high value $\lambda=1e^{-1}$ and annealed during the search to find good bounds for $\lambda$ and for most applications should be fine to leave as default.
#
# * `max_lam_steps`: the number of steps (outer loops) to search for with a different value of $\lambda$.
#
#
#
#
# While the default values for the loss term coefficients worked well for the simple examples provided in the notebooks, it is recommended to test their robustness for your own applications.
# <div class="alert alert-warning">
# Warning
#
# Once a `Counterfactual` instance is initialized, the parameters of it are frozen even if creating a new instance. This is due to TensorFlow behaviour which holds on to some global state. In order to change parameters of the explainer in the same session (e.g. for explaining different models), you will need to reset the TensorFlow graph manually:
#
# ```python
# import tensorflow as tf
# tf.keras.backend.clear_session()
# ```
# You may need to reload your model after this. Then you can create a new `Counterfactual` instance with new parameters.
#
# </div>
# ### Fit
#
# The method is purely unsupervised so no fit method is necessary.
# ### Explanation
#
# We can now explain the instance $X$:
#
# ```python
# explanation = cf.explain(X)
# ```
#
# The ```explain``` method returns an `Explanation` object with the following attributes:
#
# * *cf*: dictionary containing the counterfactual instance found with the smallest distance to the test instance, it has the following keys:
#
# * *X*: the counterfactual instance
# * *distance*: distance to the original instance
# * *lambda*: value of $\lambda$ corresponding to the counterfactual
# * *index*: the step in the search procedure when the counterfactual was found
# * *class*: predicted class of the counterfactual
# * *proba*: predicted class probabilities of the counterfactual
# * *loss*: counterfactual loss
#
# * *orig_class*: predicted class of original instance
#
# * *orig_proba*: predicted class probabilites of the original instance
#
# * *all*: dictionary of all instances encountered during the search that satisfy the counterfactual constraint but have higher distance to the original instance than the returned counterfactual. This is organized by levels of $\lambda$, i.e. ```explanation['all'][0]``` will be a list of dictionaries corresponding to instances satisfying the counterfactual condition found in the first iteration over $\lambda$ during bisection.
# ### Numerical Gradients
# So far, the whole optimization problem could be defined within the TF graph, making automatic differentiation possible. It is however possible that we do not have access to the model architecture and weights, and are only provided with a ```predict``` function returning probabilities for each class. The counterfactual can then be initialized in the same way as before, but using a prediction function:
#
# ```python
# # define model
# model = load_model('mnist_cnn.h5')
# predict_fn = lambda x: cnn.predict(x)
#
# # initialize explainer
# shape = (1,) + x_train.shape[1:]
# cf = Counterfactual(predict_fn, shape, distance_fn='l1', target_proba=1.0,
# target_class='other', max_iter=1000, early_stop=50, lam_init=1e-1,
# max_lam_steps=10, tol=0.05, learning_rate_init=0.1,
# feature_range=(-1e10, 1e10), eps=0.01, init
# ```
#
#
# In this case, we need to evaluate the gradients of the loss function with respect to the input features $X$ numerically:
#
# $$\frac{\partial L_{\text{pred}}}{\partial X} = \frac{\partial L_\text{pred}}{\partial p} \frac{\partial p}{\partial X}
# $$
#
# where $L_\text{pred}$ is the predict function loss term, $p$ the predict function and $x$ the input features to optimize. There is now an additional hyperparameter to consider:
#
# * `eps`: a float or an array of floats to define the perturbation size used to compute the numerical gradients of $^{\delta p}/_{\delta X}$. If a single float, the same perturbation size is used for all features, if the array dimension is *(1 x nb of features)*, then a separate perturbation value can be used for each feature. For the Iris dataset, `eps` could look as follows:
#
# ```python
# eps = np.array([[1e-2, 1e-2, 1e-2, 1e-2]]) # 4 features, also equivalent to eps=1e-2
# ```
# ## Examples
# [Counterfactual instances on MNIST](../examples/cf_mnist.ipynb)
| doc/source/methods/CF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# +
import vcf
# %matplotlib inline
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import matplotlib.ticker as ticker
from pylab import plot, show, savefig, xlim, figure, hold, ylim, legend, boxplot, setp, axes
from itertools import compress
from pylab import MaxNLocator
import seaborn as sns; sns.set()
from matplotlib.colors import LogNorm
from matplotlib import gridspec
import ast
import itertools
import seaborn as sns
from sklearn.preprocessing import StandardScaler
import fastcluster
from sklearn import cluster, datasets
import scipy.cluster.hierarchy as hier
from sklearn.cluster import KMeans
import time
import sys
import math
import Bio
from Bio.Alphabet import IUPAC
from Bio.Blast.Applications import NcbiblastnCommandline
from Bio.Blast import NCBIXML
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
from Bio.SeqFeature import SeqFeature, FeatureLocation
from Bio import pairwise2
from Bio import SeqIO
from Bio.Graphics import GenomeDiagram
from Bio.SeqUtils import GC
from Bio.Align.Applications import MuscleCommandline
from StringIO import StringIO
from Bio import AlignIO
from Bio.Align import AlignInfo
from Bio.Seq import MutableSeq
import itertools
import networkx as nx
import scipy
import datetime as dt
import statsmodels.api as sm
import scipy.stats as stats
import decimal
#for exporting to Adobe Illustrator
mpl.rcParams['pdf.fonttype'] = 42
mpl.rcParams['ps.fonttype'] = 42
# -
# #### Import Sample Annotation file for filtered *longitudinal* isolates pairs
sample_annotation = pd.read_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/sample_annotation_files/Longitudinal_fastq_path_names_and_JankyPipe_tags_filtered_final.csv' , sep = ',').set_index('patient_id')
sample_annotation.head()
# #################################################################################################################################################################################################################################
# ## [1] Collect all AR nSNPs with $\Delta AF \ge 5\%$ (in genes & intergenic regions associated with antibiotic resistance)
# #################################################################################################################################################################################################################################
# #### Load in filtered Base Calls from longitudinal Base Call analysis (pairs of Base Calls that are different subjects, filtered for low coverage have a difference in Alternate Allele Frequency $\ge 5\%$)
# The DataFrame below will consist of two rows per in-host SNP, corresponding to the 1st and 2nd alternate allele frequencies.
within_host_Base_Call_variants_all_patients = pd.read_pickle('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/pickled_files/variant_calling/longitudinal_SNPs/longitudinal_SNP_variants_05_delta_in_alt_AF.pkl')
within_host_Base_Call_variants_all_patients.head(n=10)
np.shape(within_host_Base_Call_variants_all_patients)
# ### *Filter 1*: Filter out paired Base Calls if (minor allele has less than 5 reads supporting it) and ($\Delta AF$ < 20%)
# +
#list that stores the DataFrame indices of all SNPs that don't pass Delta AF threshold
BaseCalls_to_Drop = []
#dictionary for ordering of bases in Base Counts field
base_order_dict = {'A':0 , 'C':1 , 'G':2 , 'T':3}
delta_AF_threshold = 0.20
#for each pair of Base Calls,
for Base_Call_A_index , Base_Call_B_index in zip( range(0 , np.shape(within_host_Base_Call_variants_all_patients)[0] , 2) , range(1 , np.shape(within_host_Base_Call_variants_all_patients)[0] , 2) ):
############################################################################################################################################
#get relevant info for the Base Calls corresponding to both isolates
Base_Call_isolate_A_data = within_host_Base_Call_variants_all_patients.loc[Base_Call_A_index , :]
Base_Call_isolate_B_data = within_host_Base_Call_variants_all_patients.loc[Base_Call_B_index , :]
#pull BC 'field' (base count in pileups at for each isolates in the pair)
Base_Call_isolate_A_base_counts = Base_Call_isolate_A_data['INFO']['BC']
Base_Call_isolate_B_base_counts = Base_Call_isolate_B_data['INFO']['BC']
#pull QP 'field' (quality-percentage in pileups supporting each base; for each isolate in the pair)
Base_Call_isolate_A_QP = Base_Call_isolate_A_data['INFO']['QP']
Base_Call_isolate_B_QP = Base_Call_isolate_B_data['INFO']['QP']
#get reference and alternate alleles for both isolates
#reference alleles
isolate_A_ref_allele = Base_Call_isolate_A_data['ref_base']
isolate_B_ref_allele = Base_Call_isolate_B_data['ref_base']
#alternate alleles (if there are any)
isolate_A_alt_allele = Base_Call_isolate_A_data['alt_base']
isolate_B_alt_allele = Base_Call_isolate_B_data['alt_base']
#get the RAW NUMBER OF READS supporting each allele (ref alleles for isolates A & B and possibly alt alleles for isolates A & B)
#reference allele counts
isolate_A_ref_allele_read_counts = Base_Call_isolate_A_base_counts[base_order_dict[isolate_A_ref_allele]]
isolate_B_ref_allele_read_counts = Base_Call_isolate_B_base_counts[base_order_dict[isolate_B_ref_allele]]
#alternate allele counts
if isolate_A_alt_allele != 'Z':
isolate_A_alt_allele_read_counts = Base_Call_isolate_A_base_counts[base_order_dict[isolate_A_alt_allele]]
elif isolate_A_alt_allele == 'Z':
isolate_A_alt_allele_read_counts = 0
if isolate_B_alt_allele != 'Z':
isolate_B_alt_allele_read_counts = Base_Call_isolate_B_base_counts[base_order_dict[isolate_B_alt_allele]]
elif isolate_B_alt_allele == 'Z':
isolate_B_alt_allele_read_counts = 0
#get the WEIGHTED PERCENTAGE OF READS supporting each allele (ref alleles for isolates A & B and possibly alt alleles for isolates A & B)
#reference allele weighted %
isolate_A_ref_allele_QP = Base_Call_isolate_A_QP[base_order_dict[isolate_A_ref_allele]]
isolate_B_ref_allele_QP = Base_Call_isolate_B_QP[base_order_dict[isolate_B_ref_allele]]
#alternate allele weighted %
if isolate_A_alt_allele != 'Z':
isolate_A_alt_allele_QP = Base_Call_isolate_A_QP[base_order_dict[isolate_A_alt_allele]]
elif isolate_A_alt_allele == 'Z':
isolate_A_alt_allele_QP = 0
if isolate_B_alt_allele != 'Z':
isolate_B_alt_allele_QP = Base_Call_isolate_B_QP[base_order_dict[isolate_B_alt_allele]]
elif isolate_B_alt_allele == 'Z':
isolate_B_alt_allele_QP = 0
############################################################################################################################################
#pull Alterante AF for the base calls corresponding to each isolates in a serial pair
Base_Call_isolate_A_alt_AF = within_host_Base_Call_variants_all_patients.loc[Base_Call_A_index , 'alt_AF']
Base_Call_isolate_B_alt_AF = within_host_Base_Call_variants_all_patients.loc[Base_Call_B_index , 'alt_AF']
#calculate the difference of the Alternate Allele Frequencies of both serial isolates
delta_AF = abs(Base_Call_isolate_B_alt_AF - Base_Call_isolate_A_alt_AF)
############################################################################################################################################
#make sure at least 5 reads supports each allele if the call is mixed (that is some reads support the reference and some support the alternate)
## if one allele has 0 reads supporting it, whether its the reference or alternate, the other allele should have at least 25 reads supporting it (min depth)
#print str(isolate_A_ref_allele_QP) + ' ' + str(isolate_A_ref_allele_read_counts) + ' ' + str(isolate_A_alt_allele_QP) + ' ' + str(isolate_A_alt_allele_read_counts)
#if either allele has QP > 0, then it should have at least 5 reads supporting that percentage
#if reads support allele for at least 1%, then there should be at least 5 reads supporting that allele
if ( ( (isolate_A_ref_allele_QP > 0) and (isolate_A_ref_allele_read_counts < 5) ) or ( (isolate_A_alt_allele_QP > 0) and (isolate_A_alt_allele_read_counts < 5) ) ) and (delta_AF < delta_AF_threshold):
#minor allele in mixed call had less than 5 reads supporting, drop base calls corresponding to BOTH isolates
BaseCalls_to_Drop.append(Base_Call_A_index)
BaseCalls_to_Drop.append(Base_Call_B_index)
elif ( ( (isolate_B_ref_allele_QP > 0) and (isolate_B_ref_allele_read_counts < 5) ) or ( (isolate_B_alt_allele_QP > 0) and (isolate_B_alt_allele_read_counts < 5) ) ) and (delta_AF < delta_AF_threshold):
#minor allele in mixed call had less than 5 reads supporting, drop base calls corresponding to BOTH isolates
BaseCalls_to_Drop.append(Base_Call_A_index)
BaseCalls_to_Drop.append(Base_Call_B_index)
within_host_Base_Call_variants_all_patients.drop(labels = BaseCalls_to_Drop , axis = 0 , inplace = True)
#reset index
within_host_Base_Call_variants_all_patients.reset_index(drop = True, inplace = True)
# -
np.shape(within_host_Base_Call_variants_all_patients)
within_host_Base_Call_variants_all_patients.head()
# ### *Filter 2*: Drop Synonymous SNPs (drops sSNPs, retains nSNPs & iSNPs)
# +
#list that stores the DataFrame indices of all SNPs that don't pass Delta AF threshold
BaseCalls_to_Drop = []
#for each pair of Base Calls, store (alternate Allele Frequency of isolate_1) and (alternate Allele Frequency of isolate_2 - alternate Allele Frequency of isolate_1)
for Base_Call_A_index , Base_Call_B_index in zip( range(0 , np.shape(within_host_Base_Call_variants_all_patients)[0] , 2) , range(1 , np.shape(within_host_Base_Call_variants_all_patients)[0] , 2) ):
#pull SNP type
Base_Call_isolate_A_SNP_type = within_host_Base_Call_variants_all_patients.loc[Base_Call_A_index , 'SNP_ftype']
Base_Call_isolate_B_SNP_type = within_host_Base_Call_variants_all_patients.loc[Base_Call_B_index , 'SNP_ftype']
#put SNP types (for both Base Calls) in a list
SNP_type_list = [Base_Call_isolate_A_SNP_type , Base_Call_isolate_B_SNP_type]
#if Synonymous SNP, then drop
if 'S' in SNP_type_list:
BaseCalls_to_Drop.append(Base_Call_A_index)
BaseCalls_to_Drop.append(Base_Call_B_index)
within_host_Base_Call_variants_all_patients.drop(labels = BaseCalls_to_Drop , axis = 0 , inplace = True)
#reset index
within_host_Base_Call_variants_all_patients.reset_index(drop = True, inplace = True)
# -
within_host_Base_Call_variants_all_patients.head(n=10)
np.shape(within_host_Base_Call_variants_all_patients)
# ### *Filter 3*: Subset to loci known to be associated with resistance to antibiotics
# #### Loci associated with Antibiotic Resistance (N = 28 genes) + (N = 6 intergenic regions) + (N = 1 rRNA)
gene_categories = pd.read_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/gene_categories/gene_categories.csv').set_index('name')
antibiotic_resistance_genes = gene_categories[gene_categories.Gene_Category == 'Antibiotic Resistance']
antibiotic_resistance_genes = list(antibiotic_resistance_genes.gene_id)
# +
reference_genome_annotation = pd.read_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/H37Rv/h37rv_genome_summary.txt', '\t').set_index('name')
#list that holds all reference positions corresponding to SNPs we will scan for
H37Rv_positions_of_interest = []
genic_regions_of_interest = pd.DataFrame(columns = ['H37Rv_start' , 'H37Rv_end' , 'type' , 'gene_after' , 'gene_before' , 'symbol' , 'description'])
gene_i = 0
for gene_id in antibiotic_resistance_genes:
#get reference positions tht corresond to each gene
gene_id_info = reference_genome_annotation.loc[gene_id, :]
gene_id = gene_id_info.name
chrom_start = gene_id_info.chromStart
chrom_end = gene_id_info.chromEnd
#find the position of the first base relative to H37Rv in 5' -> 3'
H37Rv_start = min(chrom_start , chrom_end)
H37Rv_end = max(chrom_start , chrom_end)
#store relevant gene info in DataFrame
genic_regions_of_interest.loc[gene_i , :] = [H37Rv_start , H37Rv_end , 'gene' , gene_id , gene_id , gene_id_info.symbol , gene_id_info.description]
gene_i += 1
#store all corresponding H37Rv Reference Positions to gene_id in list of reference positions to scan
H37Rv_positions_of_interest = H37Rv_positions_of_interest + range(H37Rv_start+1 , H37Rv_end+1)
#get rid of redundant positions & sort
H37Rv_positions_of_interest = list(set(H37Rv_positions_of_interest))
H37Rv_positions_of_interest = list(np.sort(np.array(H37Rv_positions_of_interest)))
# -
len( H37Rv_positions_of_interest )
genic_regions_of_interest.head()
# #### Intergenic Regions associated with Antibiotic Resistance
# +
#create DataFrame to store coordinates taken from tuberculist
intergenic_regions_of_interest = pd.DataFrame(columns = ['H37Rv_start' , 'H37Rv_end' , 'type' , 'gene_after' , 'gene_before' , 'symbol' , 'description'])
intergenic_regions_of_interest.loc[0 , :] = [1673299 , 1673439 , 'promoter' , 'fabG1' , 'Rv1482c' , np.nan , 'promoter_fabG1-inhA'] #promoter for inhA
intergenic_regions_of_interest.loc[1 , :] = [2713784 , 2714123 , 'intergenic' , 'eis' , 'Rv2415c' , np.nan , np.nan]
intergenic_regions_of_interest.loc[2 , :] = [2726087 , 2726192 , 'promoter' , 'ahpC' , 'oxyR\'' , np.nan , 'promoter_ahpC']
intergenic_regions_of_interest.loc[3 , :] = [2289241 , 2289281 , 'promoter' , 'Rv2044c' , 'pncA' , np.nan , 'promoter_pncA'] #promoter for pncA
intergenic_regions_of_interest.loc[4 , :] = [4243147 , 4243232 , 'promoter' , 'embA' , 'embC' , np.nan , 'promoter_embA-embB']
intergenic_regions_of_interest.loc[5 , :] = [3067945 , 3068188 , 'intergenic' , 'hsdS.1' , 'thyX' , np.nan , np.nan]
intergenic_regions_of_interest.loc[6 , :] = [2715332 , 2715471 , 'promoter' , 'Rv2417c' , 'eis' , np.nan , 'promoter_eis'] #promoter for eis
#add rrs coordinates as well
intergenic_regions_of_interest.loc[7 , :] = [1471845 , 1473382 , 'rRNA' , 'rrl' , 'mcr3' , np.nan , 'ribosomal_RNA_16S'] #rRNA
# -
intergenic_regions_of_interest
# +
for intergenic_region_i in list(intergenic_regions_of_interest.index):
#get reference positions tht corresond to each intergenic region
intergenic_region_info = intergenic_regions_of_interest.loc[intergenic_region_i, :]
#find the position of the first base relative to H37Rv in 5' -> 3'
H37Rv_start = intergenic_region_info.H37Rv_start
H37Rv_end = intergenic_region_info.H37Rv_end
#store all corresponding H37Rv Reference Positions to gene_id in list of reference positions to scan
H37Rv_positions_of_interest = H37Rv_positions_of_interest + range(H37Rv_start+1 , H37Rv_end+1)
#get rid of redundant positions & sort
H37Rv_positions_of_interest = list(set(H37Rv_positions_of_interest))
H37Rv_positions_of_interest = list(np.sort(np.array(H37Rv_positions_of_interest)))
# -
len( H37Rv_positions_of_interest )
# #### output CSV file of regions that we will search for mutations within
H37Rv_regions_of_interest = genic_regions_of_interest.append(intergenic_regions_of_interest)
H37Rv_regions_of_interest.reset_index(inplace = True , drop = True)
H37Rv_regions_of_interest.head()
H37Rv_regions_of_interest.tail()
H37Rv_regions_of_interest
H37Rv_regions_of_interest.to_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/gene_categories/H37Rv_AR_regions_of_interest.csv' , sep = ',')
# ### *Filter* out Base Calls that do not occur in our H37Rv positions of interest
# +
#convert list of H37Rv positions to check SNPs for into a set which is much faster to lookup than a list
H37Rv_positions_of_interest = set(H37Rv_positions_of_interest)
#list that stores the DataFrame indices of all SNPs that don't pass Delta AF threshold
BaseCalls_to_Drop = []
#for each pair of Base Calls, store (alternate Allele Frequency of isolate_1) and (alternate Allele Frequency of isolate_2 - alternate Allele Frequency of isolate_1)
for Base_Call_A_index , Base_Call_B_index in zip( range(0 , np.shape(within_host_Base_Call_variants_all_patients)[0] , 2) , range(1 , np.shape(within_host_Base_Call_variants_all_patients)[0] , 2) ):
#pull reference positions (only need the H37Rv ref position for 1 of the pair of base calls)
Base_Call_H37Rv_Ref_Pos = within_host_Base_Call_variants_all_patients.loc[Base_Call_A_index , 'ref_position']
#if Reference Position is not associated with Antibiotic Resistance, then drop
if Base_Call_H37Rv_Ref_Pos not in H37Rv_positions_of_interest:
BaseCalls_to_Drop.append(Base_Call_A_index)
BaseCalls_to_Drop.append(Base_Call_B_index)
within_host_Base_Call_variants_all_patients.drop(labels = BaseCalls_to_Drop , axis = 0 , inplace = True)
#reset index
within_host_Base_Call_variants_all_patients.reset_index(drop = True, inplace = True)
# -
within_host_Base_Call_variants_all_patients.head(n=10)
np.shape(within_host_Base_Call_variants_all_patients)
# output Filtered DataFrame (containing only Ref Positions that we care about) for downstream analysis
within_host_Base_Call_variants_all_patients.to_pickle('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/pickled_files/variant_calling/longitudinal_SNPs/longitudinal_SNP_variants_05_delta_in_alt_AF_antibiotic_resistance.pkl')
# ouput Filtered DataFrame as CSV for publication
within_host_Base_Call_variants_all_patients.to_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/variant_calling/longitudinal_SNPs/longitudinal_SNP_variants_05_delta_in_alt_AF_antibiotic_resistance.csv')
# #################################################################################################################################################################################################################################
# ## [2] Analyze Antibiotic Resistance Associated SNPs
# #################################################################################################################################################################################################################################
# Load in DataFrame of filtered variants (from above)
within_host_Base_Call_variants_all_patients = pd.read_pickle('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/pickled_files/variant_calling/longitudinal_SNPs/longitudinal_SNP_variants_05_delta_in_alt_AF_antibiotic_resistance.pkl')
# ### Re-format: DataFrame to contain 1 row per SNP
# +
all_SNPs_within_patients = pd.DataFrame()
#common information to both Base Calls (can just look at isolate A)
population = {}
patient_id = {}
ref_position = {}
genomic_coord = {}
ref_allele = {}
alt_allele = {}
gene_id = {}
Gene_Category = {}
symbol = {}
#look at info for both Base Calls
alt_AF_diff = {}
SNP_type = {}
AA_change = {}
SNP_index = 0
#iterate through indices for isolate A (store common information for patient isolate A & B came from and Base Call), calculate different in Alternate Allele Frequencies, store Syn or NSyn info
for even_index in range(0 , np.shape(within_host_Base_Call_variants_all_patients)[0] , 2):
#Base Call info for isolate A
Base_Call_info_isolate_A = within_host_Base_Call_variants_all_patients.loc[even_index , :]
#Base Call info for isolate B
Base_Call_info_isolate_B = within_host_Base_Call_variants_all_patients.loc[even_index+1 , :]
population[SNP_index] = Base_Call_info_isolate_A.population
patient_id[SNP_index] = Base_Call_info_isolate_A.patient_id
ref_position[SNP_index] = Base_Call_info_isolate_A.ref_position
genomic_coord[SNP_index] = Base_Call_info_isolate_A.gene_coord
ref_allele[SNP_index] = Base_Call_info_isolate_A.ref_base
gene_id[SNP_index] = Base_Call_info_isolate_A.gene_id
Gene_Category[SNP_index] = Base_Call_info_isolate_A.gene_category
symbol[SNP_index] = Base_Call_info_isolate_A.gene_symbol
#look for alternate allele between both base calls, ignore dummy 'Z' holder
for alt_base_i in list( set([Base_Call_info_isolate_A.alt_base , Base_Call_info_isolate_B.alt_base]) ):
if alt_base_i != 'Z':
alt_allele[SNP_index] = alt_base_i
break
#get difference in Alternate Allele Frequencies
alt_AF_diff[SNP_index] = abs(Base_Call_info_isolate_A.alt_AF - Base_Call_info_isolate_B.alt_AF)
#get type of SNP
if 'I' in Base_Call_info_isolate_A.SNP_ftype + '_' + Base_Call_info_isolate_B.SNP_ftype:
SNP_type[SNP_index] = 'I'
elif 'N' in Base_Call_info_isolate_A.SNP_ftype + '_' + Base_Call_info_isolate_B.SNP_ftype:
SNP_type[SNP_index] = 'N'
#get AA change
AA_change_calls = [Base_Call_info_isolate_A.AA_change , Base_Call_info_isolate_B.AA_change]
try:
AA_change_calls.remove('None')
except ValueError:
pass
AA_change[SNP_index] = AA_change_calls[0]
SNP_index += 1
#convert dictionaries into series
population = pd.Series(population)
patient_id = pd.Series(patient_id)
ref_position = pd.Series(ref_position)
genomic_coord = pd.Series(genomic_coord)
ref_allele = pd.Series(ref_allele)
alt_allele = pd.Series(alt_allele)
gene_id = pd.Series(gene_id)
Gene_Category = pd.Series(Gene_Category)
symbol = pd.Series(symbol)
alt_AF_diff = pd.Series(alt_AF_diff)
SNP_type = pd.Series(SNP_type)
AA_change = pd.Series(AA_change)
#create DataFrame
all_SNPs_within_patients['population'] = population
all_SNPs_within_patients['patient_id'] = patient_id
all_SNPs_within_patients['ref_position'] = ref_position
all_SNPs_within_patients['genomic_coord'] = genomic_coord
all_SNPs_within_patients['ref_allele'] = ref_allele
all_SNPs_within_patients['alt_allele'] = alt_allele
all_SNPs_within_patients['gene_id'] = gene_id
all_SNPs_within_patients['Gene_Category'] = Gene_Category
all_SNPs_within_patients['symbol'] = symbol
all_SNPs_within_patients['alt_AF_difference'] = alt_AF_diff
all_SNPs_within_patients['SNP_type'] = SNP_type
all_SNPs_within_patients['AA_change'] = AA_change
# -
all_SNPs_within_patients.head(n=6)
np.shape(all_SNPs_within_patients)
# Store as a CSV file
all_SNPs_within_patients.to_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/variant_calling/longitudinal_SNPs/SNPs_between_isolates_delta_05_antibiotic_resistance.csv' , sep = ',')
# ### Re-format: Starting Alternate Allele Frequency vs. Ending Alternate Allele Frequency
within_host_Base_Call_variants_all_patients_sample_order = pd.merge( within_host_Base_Call_variants_all_patients , sample_annotation.loc[: , ['sample_order' , 'tag']] , how = 'left' , on = 'tag')
within_host_Base_Call_variants_all_patients_sample_order.head()
np.shape(within_host_Base_Call_variants_all_patients_sample_order)
# +
beginning_alternate_AF = []
ending_alternate_AF = []
change_in_alternate_AF = []
Gene_Category = []
Gene_IDs = []
SNP_types = []
#for each pair of Base Calls, store (alternate Allele Frequency of isolate_1) and (alternate Allele Frequency of isolate_2 - alternate Allele Frequency of isolate_1)
for Base_Call_A_index , Base_Call_B_index in zip( range(0 , np.shape(within_host_Base_Call_variants_all_patients_sample_order)[0] , 2) , range(1 , np.shape(within_host_Base_Call_variants_all_patients_sample_order)[0] , 2) ):
#find the Base Call index that corresponds to the earlier isolate
if within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'sample_order'] == 1:
Base_Call_isolate_1 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , :]
Base_Call_isolate_2 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_B_index , :]
elif within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_B_index , 'sample_order'] == 1:
Base_Call_isolate_1 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_B_index , :]
Base_Call_isolate_2 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , :]
#store the Alterante Allele Frequency of the first serial isolate
beginning_alternate_AF.append(Base_Call_isolate_1.alt_AF)
#stotre the Alternate Allele Frequency of the second serial isolate
ending_alternate_AF.append(Base_Call_isolate_2.alt_AF)
#store the difference of the Alternate Allele Frequencies of both serial isolates
change_in_alternate_AF.append( abs(Base_Call_isolate_2.alt_AF - Base_Call_isolate_1.alt_AF) )
#store the functional impact of mutation
SNP_type_for_isolates = [Base_Call_isolate_1.SNP_ftype , Base_Call_isolate_2.SNP_ftype]
if 'N' in SNP_type_for_isolates:
SNP_types.append('N')
elif 'I' in SNP_type_for_isolates:
SNP_types.append('I')
#store Gene Category & gene_id of the gene that Reference Position is located on
Gene_Category.append(Base_Call_isolate_1.gene_category)
Gene_IDs.append(Base_Call_isolate_1.gene_id)
#create DataFrame with all Information
change_in_AF_DF = pd.DataFrame()
change_in_AF_DF['beginning_alt_AF'] = beginning_alternate_AF
change_in_AF_DF['ending_alt_AF'] = ending_alternate_AF
change_in_AF_DF['change_alt_AF'] = change_in_alternate_AF
change_in_AF_DF['Gene_Category'] = Gene_Category
change_in_AF_DF['gene_id'] = Gene_IDs
change_in_AF_DF['SNP_type'] = SNP_types
# -
change_in_AF_DF.head()
np.shape(change_in_AF_DF)
# #### nSNPs in genes associated with Antibiotic Resistance
np.shape( change_in_AF_DF[change_in_AF_DF.Gene_Category == 'Antibiotic Resistance'] )
# #### SNPs in rrs
np.shape( change_in_AF_DF[change_in_AF_DF.gene_id == 'Rvnr01'] )
# #### SNPs in intergenic regions
np.shape( change_in_AF_DF[change_in_AF_DF.SNP_type == 'I'] )
# #################################################################################################################################################################################################################################
# ## [3] Analysis 1
# #################################################################################################################################################################################################################################
# #### Full DF of AR SNPs with: $\Delta AF \ge 5 \%$ , across all 200 subjects, and in an AR-loci
within_host_Base_Call_variants_all_patients_sample_order.head()
np.shape(within_host_Base_Call_variants_all_patients_sample_order)
# #### Collect the alternate AF's for isolate 1 and isolate 2 for each SNP
# +
alt_AF_1_array = []
alt_AF_2_array = []
#for each pair of Base Calls, store (alternate Allele Frequency of isolate_1) and (alternate Allele Frequency of isolate_2 - alternate Allele Frequency of isolate_1)
for Base_Call_A_index , Base_Call_B_index in zip( range(0 , np.shape(within_host_Base_Call_variants_all_patients_sample_order)[0] , 2) , range(1 , np.shape(within_host_Base_Call_variants_all_patients_sample_order)[0] , 2) ):
#pull alternate AFs
if within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'sample_order'] == 1:
alt_AF_1 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'alt_AF']
alt_AF_2 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_B_index , 'alt_AF']
elif within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'sample_order'] == 2:
alt_AF_2 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'alt_AF']
alt_AF_1 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_B_index , 'alt_AF']
alt_AF_1_array.append(alt_AF_1)
alt_AF_2_array.append(alt_AF_2)
alt_AF_1_array = np.array(alt_AF_1_array)
alt_AF_2_array = np.array(alt_AF_2_array)
# -
plt.style.use('ggplot')
plt.rcParams['lines.linewidth']=0
plt.rcParams['axes.facecolor']='1.0'
plt.rcParams['xtick.color']='black'
plt.rcParams['axes.grid']=False
plt.rcParams['axes.edgecolor']='black'
plt.rcParams['grid.color']= '1.0'
plt.rcParams.update({'font.size': 14})
# +
#store the values in a dependent and independent vector
Y = alt_AF_2_array
X = alt_AF_1_array
X = sm.add_constant(X)
#run the OLS with statsmodels
model = sm.OLS(Y , X)
results = model.fit()
#store the fitted parameters from the regression
constant_from_OLS = results.params[0]
coeff_from_OLS = results.params[1]
#create a vector for the line using the output parameters from the OLS regression
X_values_line_best_fit = np.arange( min(alt_AF_1_array) , max(alt_AF_1_array) , 0.01)
Y_values_line_best_fit = coeff_from_OLS * X_values_line_best_fit + constant_from_OLS
# -
results.summary()
# +
fig , ax = plt.subplots()
epsilon = 0.02
plt.scatter( alt_AF_1_array , alt_AF_2_array , color = '0.0' , edgecolor = 'white' , linewidth = 1.5 , alpha = 0.5 , s = 35)
#plot Line of Best Fit from OLS regression
#plt.plot(X_values_line_best_fit , Y_values_line_best_fit , color = 'xkcd:bright red' , linestyle = '-' , linewidth = 1.5 , zorder=0 , alpha = 1.0)
#plt.plot(X_values_line_best_fit , X_values_line_best_fit , color = 'k' , linestyle = '--' , linewidth = 1.0 , zorder=0 , alpha = 1.0)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.tick_params(labelcolor = 'k')
plt.ylabel('Alternate Allele Frequency at Timepoint 2' , fontweight = 'bold', fontsize = 11 , color = 'k')
plt.xlabel('Alternate Allele Frequency at Timepoint 1' , fontweight = 'bold', fontsize = 11 , color = 'k')
plt.title( 'Trajectories of 1939 SNPs in AR-associated Regions with $\Delta AF >= 5 \%$\nacross all Subjects' , fontsize = 11 , color = 'k')
plt.xlim(0.0-epsilon , np.max(alt_AF_1_array)+epsilon)
plt.ylim(0.0-epsilon , np.max(alt_AF_2_array)+epsilon+0.02)
fig = plt.gcf()
fig.set_size_inches(7.0, 7.0)
fig.tight_layout()
file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/figures/antibiotic_resistance_analysis/alt1_vs_alt2_for_AR_SNPs_all_subjects.pdf'
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = True)
plt.show()
# -
# #### Heatmap version of plot above
plt.style.use('ggplot')
plt.rcParams['lines.linewidth']=0
plt.rcParams['axes.facecolor']='1.0'
plt.rcParams['xtick.color']='black'
plt.rcParams['axes.grid']=False
plt.rcParams['axes.edgecolor']='black'
plt.rcParams['grid.color']= '1.0'
plt.rcParams.update({'font.size': 14})
# +
#define colormap
def truncate_colormap(cmap, minval=0.0, maxval=1.0, n=100):
new_cmap = mpl.colors.LinearSegmentedColormap.from_list('trunc({n},{a:.2f},{b:.2f})'.format(n=cmap.name, a=minval, b=maxval), cmap(np.linspace(minval, maxval, n)))
return new_cmap
def heatmap_AF1_vs_AF2(X , Y , color_map , file_name):
############# create AF_1 vs AF_2 scatter plot #############
fig = plt.figure(figsize=(7, 7.7), dpi=100)
from matplotlib import gridspec
gs = gridspec.GridSpec(2, 1,
width_ratios = [1],
height_ratios = [50,1.5],
wspace=0.00,
hspace=0.19)
cmap = plt.get_cmap(color_map)
trunc_cmap = truncate_colormap(cmap, 0.07, 1.0)
#HEATMAP
axmatrix = fig.add_subplot(gs[0])
# turn off the frame
axmatrix.set_frame_on(False)
X = X + 0.0001 #add psuedo-count to push into bin next-over (to the right)
Y = Y + 0.0001 #add psuedo-count to push into bin next-over (to the right)
heatmap_array , xedges , yedges = np.histogram2d(X , Y , bins = 101)
norm = mpl.colors.PowerNorm(gamma=1./5.3)
heatmap = plt.pcolor(heatmap_array.T , cmap = trunc_cmap , norm = norm , edgecolors = '1.0', linewidth = 0.35)
#X-TICKS
#------------------------------------------------------------------------------------------------------------------------
axmatrix.set_xticks(np.arange(0 , 101 , 10) + 0.5, minor=False)
axmatrix.set_xticklabels( [str(float(decimal.Decimal(str(tick_i)))) for tick_i in np.arange(0.0 , 1.01 , 0.10) ] , rotation='horizontal', fontsize = 7)
axmatrix.tick_params(labelsize=8 , pad = 1.5 , axis = 'x')
#Y-TICKS
#------------------------------------------------------------------------------------------------------------------------
axmatrix.set_yticks(np.arange(0 , 101 , 10) + 0.5, minor=False)
axmatrix.set_yticklabels( [str(float(decimal.Decimal(str(tick_i)))) for tick_i in np.arange(0.0 , 1.01 , 0.10) ] , rotation='horizontal', fontsize = 7)
axmatrix.tick_params(labelsize=8 , pad = 1.5 , axis = 'y')
#COLORBAR
#------------------------------------------------------------------------------------------------------------------------
color_bar_ax = fig.add_subplot(gs[1])
cb = plt.colorbar( heatmap , cax = color_bar_ax , orientation = 'horizontal' , ticks = [0] + list( np.ceil( np.logspace( math.log(1 , 5) , math.log(int(np.max( heatmap_array )) , 5) , num = 9 , base = 5)).astype(int) ) + [int(np.max( heatmap_array ))] )
cb.outline.set_linewidth(0.0)
color_bar_ax.tick_params(labelsize=7 , pad = 3)
plt.tick_params(
axis='x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
labelbottom=True) # labels along the bottom edge are off
#------------------------------------------------------------------------------------------------------------------------
axmatrix.set_ylabel('Alternate Allele Frequency at Timepoint 2' , color = 'k' , fontsize = 10)
axmatrix.set_xlabel('Alternate Allele Frequency at Timepoint 1' , color = 'k' , fontsize = 10)
axmatrix.set_title( 'Trajectories of 1939 SNPs in AR-associated Regions with $\Delta AF >= 5 \%$\nacross all Subjects' , fontsize = 10 , color = 'k')
axmatrix.tick_params(labelcolor = 'k')
#output for Adobe Illustrator
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = True)
plt.show()
##################################################################
# -
#color within-host Base Calls w/ a difference in Alternate Allele Frequencies >= 1% in Green (SNPs that we ARE taking into account)
X = alt_AF_1_array
Y = alt_AF_2_array
color_map = 'gist_gray_r'
file_name = file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/figures/antibiotic_resistance_analysis/alt1_vs_alt2_for_AR_SNPs_failure_relapse_subjects_heatmap.pdf'
heatmap_AF1_vs_AF2(X , Y , color_map , file_name)
len(alt_AF_1_array)
len(alt_AF_2_array)
# #### Calculate TPR & FPR (using an AR fixation threshold of 75%)
# +
AF_fixation_threshold = 0.75
###########
#DISCARD SNPs if AF1 >= AF_fixation_threshold and AF2 >= AF_fixation_threshold
###########
SNPs_fixed_both_timepoints = [ (AF1 >= AF_fixation_threshold) and (AF2 >= AF_fixation_threshold) for AF1 , AF2 in zip( alt_AF_1_array , alt_AF_2_array ) ]
SNPs_not_fixed_both_timespoints = np.array( [not fixed_both_timepoints for fixed_both_timepoints in SNPs_fixed_both_timepoints] )
alt_AF_1_array = alt_AF_1_array[SNPs_not_fixed_both_timespoints]
alt_AF_2_array = alt_AF_2_array[SNPs_not_fixed_both_timespoints]
###########
# -
len(alt_AF_1_array)
len(alt_AF_2_array)
# #### How many of the 1,919 nSNPs had
#
# 1. $AF_1 = 0\%$ and $AF_2 > 75\%$
#
# 2. $AF_1 \ge 1\%$ and $AF_2 > 75\%$
#
# these would be the SNPs undetectable in the first timepoint that swept and fixed
np.sum((alt_AF_1_array == 0) & (alt_AF_2_array > 0.75))
np.sum((alt_AF_1_array >= 0.01) & (alt_AF_2_array > 0.75))
# +
#take arrays for AF1 , AF2 and join into 1 array
alt_AF_array = np.vstack((alt_AF_1_array , alt_AF_2_array))
alt_AF_array = alt_AF_array.T
#rates of True Positives for every alt_AF1 classfier
TP_dict = {}
#rates of False Positives for every alt_AF1 classfier
FP_dict = {}
num_SNPs = float(len(alt_AF_1_array))
for alt_AF in np.arange(0.0 , 1.01 , 0.01):
#boolean for SNPs with alt AF1 >= alt_AF varying classifier
SNPs_classified_as_fixed_bool = alt_AF_array[: , 0] >= alt_AF
#boolean for SNPs with alt AF2 >= fixation threshold
SNPs_actually_fixed_bool = alt_AF_array[: , 1] >= AF_fixation_threshold
#True Positives: SNPs classified as going to fix (alt AF1 >= varying freq) and actually fixed ( alt AF2 >= fixation threshold )
TP_bool = [(classified_fixed) and (fixed) for classified_fixed , fixed in zip(SNPs_classified_as_fixed_bool , SNPs_actually_fixed_bool)]
#False Positives: SNPs classfifed as going to fix (alt AF1 >= varying freq) and NOT fixed (alt AF2 < fixation threshold)
FP_bool = [(classified_fixed) and (not fixed) for classified_fixed , fixed in zip(SNPs_classified_as_fixed_bool , SNPs_actually_fixed_bool)]
#True Negatives: SNPs not classified as going to fix (alt AF1 < varying freq) and NOT fixed (alt AF2 < fixation threshold)
TN_bool = [(not classified_fixed) and (not fixed) for classified_fixed , fixed in zip(SNPs_classified_as_fixed_bool , SNPs_actually_fixed_bool)]
#False Negatives : SNPs not classified as going to fix (alt AF1 < varying freq) and actually fixed ( alt AF2 >= fixation threshold )
FN_bool = [(not classified_fixed) and (fixed) for classified_fixed , fixed in zip(SNPs_classified_as_fixed_bool , SNPs_actually_fixed_bool)]
#calculate True & False positive rates
TP_rate = float(sum(TP_bool)) / float(sum(TP_bool) + sum(FN_bool))
FP_rate = float(sum(FP_bool)) / float(sum(FP_bool) + sum(TN_bool))
TP_dict[alt_AF] = TP_rate
FP_dict[alt_AF] = FP_rate
#convert dicts -> series
TP_rates = pd.Series(TP_dict)
FP_rates = pd.Series(FP_dict)
# -
plt.style.use('ggplot')
plt.rcParams['lines.linewidth']=0
plt.rcParams['axes.facecolor']='1.0'
plt.rcParams['xtick.color']='black'
plt.rcParams['axes.grid']=False
plt.rcParams['axes.edgecolor']='black'
plt.rcParams['grid.color']= '1.0'
plt.rcParams.update({'font.size': 14})
# +
fig , ax = plt.subplots()
epsilon = 0.02
#use this to change size of points as AF1 classifier increases
AF1_classifying_thresholds = np.arange(0.0 , 1.01 , 0.01)
plt.scatter( AF1_classifying_thresholds , TP_rates , color = '1.0' , marker = 'o' , edgecolor = 'blue' , linewidth = 1.25 , alpha = 0.80 , s = 50)
plt.scatter( AF1_classifying_thresholds , FP_rates , color = '1.0' , marker = 'o' , edgecolor = 'red' , linewidth = 1.25 , alpha = 0.80 , s = 50)
plt.scatter( 0.19 , TP_rates[0.19] , color = 'blue' , marker = 'o')
plt.scatter( 0.19 , FP_rates[0.19] , color = 'red' , marker = 'o')
plt.plot([0.0 , 1.0] , [0.05 , 0.05] , c = '0.5', linestyle = '-' , linewidth = 1.25)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.tick_params(labelcolor = 'k')
plt.ylabel('True Positive Rate (blue) \ False Positive Rate (red)' , fontweight = 'bold', fontsize = 11 , color = 'k')
plt.xlabel('AF1 Threshold' , fontweight = 'bold', fontsize = 11 , color = 'k')
plt.title( 'TPR/FPR analysis of 1919 SNPs in AR-associated Regions with\n$\Delta AF >= 5 \%$ across all Subjects', fontsize = 11 , color = 'k')
plt.xlim(0.0 - epsilon , 1.0 + epsilon)
plt.ylim(0.0 - epsilon , 1.0 + epsilon)
fig = plt.gcf()
fig.set_size_inches(7.0, 7.0)
fig.tight_layout()
file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/figures/antibiotic_resistance_analysis/TPR_FPR_vs_AF1_threshold_for_AR_SNPs_all_subjects_fixation_thresh_75.pdf'
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = True)
plt.show()
# -
FP_rates[FP_rates <= 0.05].head()
# Sensitivity
TP_rates[0.19]
# Specificity
1 - FP_rates[0.19]
TP = sum( [ (AF1 >= 0.19) and (AF2 >= AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] )
print TP
FP = sum( [ (AF1 >= 0.19) and (AF2 < AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] )
print FP
TN = sum( [ (AF1 < 0.19) and (AF2 < AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] )
print TN
FN = sum( [ (AF1 < 0.19) and (AF2 >= AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] )
print FN
Sensitivity = float(TP) / (float(TP) + float(FN))
print Sensitivity
Specificity = 1 - (float(FP) / (float(FP) + float(TN)))
print Specificity
# Get the number of isolates that had $19\% \le \text{AF}_1 <75\%$ and $\text{AF}_2 \ge 75\%$
bool_filter = np.array( [ (AF1 >= 0.19) and (AF2 >= AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] )
alt_AF_array[: , 0][bool_filter]
alt_AF_array[: , 1][bool_filter]
print np.mean( alt_AF_array[: , 1][bool_filter] - alt_AF_array[: , 0][bool_filter] )
# #### Breakdown of SNPs that fixed
# $\text{AF}_1 \ge 19\%$ and $\text{AF}_2 \ge 75\%$
sum( [ (AF1 >= 0.19) and (AF2 >= AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] ) #TP
# $0\% < \text{AF}_1 < 19\%$ and $\text{AF}_2 \ge 75\%$
sum( [ (0 < AF1 < 0.19) and (AF2 >= AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] ) #FN
# $\text{AF}_1 = 0\%$ and $\text{AF}_2 \ge 75\%$
sum( [ (AF1 == 0) and (AF2 >= AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] ) #FN
# #### Breakdown of SNPs that didn't fix
# $\text{AF}_1 \ge 19\%$ and $\text{AF}_2 < 75\%$
sum( [ (AF1 >= 0.19) and (AF2 < AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] )
# $0\% < \text{AF}_1 < 19\%$ and $\text{AF}_2 < 75\%$
sum( [ (0 < AF1 < 0.19) and (AF2 < AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] )
# $\text{AF}_1 = 0\%$ and $\text{AF}_2 < 75\%$
sum( [ (AF1 == 0) and (AF2 < AF_fixation_threshold) for AF1 , AF2 in zip( list(alt_AF_array[: , 0]) , list(alt_AF_array[: , 1]) ) ] )
# #### In how many isolate/subjects did the above SNPs appear within?
np.shape(within_host_Base_Call_variants_all_patients_sample_order)
within_host_Base_Call_variants_all_patients_sample_order.head(n=2)
# +
Base_Call_indices_to_keep = []
#for each pair of Base Calls, store (alternate Allele Frequency of isolate_1) and (alternate Allele Frequency of isolate_2 - alternate Allele Frequency of isolate_1)
for Base_Call_A_index , Base_Call_B_index in zip( range(0 , np.shape(within_host_Base_Call_variants_all_patients_sample_order)[0] , 2) , range(1 , np.shape(within_host_Base_Call_variants_all_patients_sample_order)[0] , 2) ):
#pull alternate AFs
if within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'sample_order'] == 1:
alt_AF_1 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'alt_AF']
alt_AF_2 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_B_index , 'alt_AF']
if not( (alt_AF_1 >= AF_fixation_threshold) and (alt_AF_2 >= AF_fixation_threshold) ):
if (alt_AF_1 >= 0.19) and (alt_AF_2 >= AF_fixation_threshold):
Base_Call_indices_to_keep.append(Base_Call_A_index)
Base_Call_indices_to_keep.append(Base_Call_B_index)
elif within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'sample_order'] == 2:
alt_AF_2 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_A_index , 'alt_AF']
alt_AF_1 = within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_B_index , 'alt_AF']
if not( (alt_AF_1 >= AF_fixation_threshold) and (alt_AF_2 >= AF_fixation_threshold) ):
if (alt_AF_1 >= 0.19) and (alt_AF_2 >= AF_fixation_threshold):
Base_Call_indices_to_keep.append(Base_Call_A_index)
Base_Call_indices_to_keep.append(Base_Call_B_index)
# -
print(float(len(Base_Call_indices_to_keep)) / 2) #number of mutations that fit criteria
within_host_Base_Call_variants_all_patients_sample_order.loc[Base_Call_indices_to_keep , :]
# #################################################################################################################################################################################################################################
# ## [4] Analysis 2 - Logistic Regression
# #################################################################################################################################################################################################################################
# #### Subset to 'fixed' AR SNPs with $\Delta AF >= 40\%$
all_SNPs_within_patients_AR_fSNPs = all_SNPs_within_patients[all_SNPs_within_patients.alt_AF_difference >= 0.40]
np.shape(all_SNPs_within_patients_AR_fSNPs)
all_SNPs_within_patients_AR_fSNPs.head()
# #### Get timing between isolate collection for each subject
time_between_serial_isolate_collection = pd.read_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/sample_annotation_files/Longitudinal_sample_annotation_for_temporal_analysis_with_dates.csv').set_index('population')
time_between_serial_isolate_collection.head()
np.shape(time_between_serial_isolate_collection)
# #### Retain only patients that for which we have date Information for both samples
# +
#get dates for each sample in pair
dates_for_first_sample = list( time_between_serial_isolate_collection.iloc[range(0 , np.shape(time_between_serial_isolate_collection)[0] , 2) , 7] )
dates_for_second_sample = list( time_between_serial_isolate_collection.iloc[range(1 , np.shape(time_between_serial_isolate_collection)[0] , 2) , 7] )
#get a boolean to check that there are dates for both samples
dates_for_both_samples = [ ( (isinstance(date_1, basestring)) and (isinstance(date_2, basestring)) ) for date_1 , date_2 in zip(dates_for_first_sample , dates_for_second_sample) ]
#get list of patients to include that have dates for sample retreival for both samples
patients_to_keep = list( time_between_serial_isolate_collection[time_between_serial_isolate_collection.sample_order == 1][dates_for_both_samples].patient_id )
#filter out patients with 1 or 2 missing dates
time_between_serial_isolate_collection_filter = [patient in patients_to_keep for patient in list(time_between_serial_isolate_collection.patient_id)]
time_between_serial_isolate_dropped_filter = [not dates_both_isolates for dates_both_isolates in time_between_serial_isolate_collection_filter]
time_between_serial_isolate_collection_dropped = time_between_serial_isolate_collection[time_between_serial_isolate_dropped_filter]
time_between_serial_isolate_collection = time_between_serial_isolate_collection[time_between_serial_isolate_collection_filter]
# -
np.shape(time_between_serial_isolate_collection)
np.shape(time_between_serial_isolate_collection_dropped )
# #### Dropped 2 patients from Witney et. al. dataset that didn't have date information both both isolates - WITNEY:27 & WITNEY:28
time_between_serial_isolate_collection_dropped
# #### Drop samples that have the same date as collection date for both isolates
# +
#get dates for each sample in pair
dates_for_first_sample = list( time_between_serial_isolate_collection.iloc[range(0 , np.shape(time_between_serial_isolate_collection)[0] , 2) , 7] )
dates_for_second_sample = list( time_between_serial_isolate_collection.iloc[range(1 , np.shape(time_between_serial_isolate_collection)[0] , 2) , 7] )
#check to make sure that the dates of collection differ between both samples
dates_differ_for_both_samples = [date_1 != date_2 for date_1 , date_2 in zip(dates_for_first_sample , dates_for_second_sample) ]
#get list of patients to include that have dates for sample retreival for both samples
patients_to_keep = list( time_between_serial_isolate_collection[time_between_serial_isolate_collection.sample_order == 1][dates_differ_for_both_samples].patient_id )
#filter out patients with 1 or 2 missing dates
time_between_serial_isolate_collection_filter = [patient in patients_to_keep for patient in list(time_between_serial_isolate_collection.patient_id)]
time_between_serial_isolate_dropped_filter = [not dates_both_isolates for dates_both_isolates in time_between_serial_isolate_collection_filter]
time_between_serial_isolate_collection_dropped = time_between_serial_isolate_collection[time_between_serial_isolate_dropped_filter]
time_between_serial_isolate_collection = time_between_serial_isolate_collection[time_between_serial_isolate_collection_filter]
# -
np.shape(time_between_serial_isolate_collection)
np.shape(time_between_serial_isolate_collection_dropped )
# #### Dropped 3 patients from Walker et. al. dataset that had same date information both both isolates - WALKER:P000059, WALKER:P000259 & WALKER:P000267
time_between_serial_isolate_collection_dropped
num_patients = np.shape(time_between_serial_isolate_collection)[0] / 2
print num_patients
# #### Get Pairwise distances for AR fSNPs for all subjects
# +
pairwise_variant_distances = pd.DataFrame(index = patients_to_keep , columns = ['population' , 'pairwise_AR_fSNP_distance' , 'time_between_collection'])
for patient_id in patients_to_keep:
population = sample_annotation.loc[patient_id , 'population'].values[0]
#pairwise distance between isolates
SNP_distance = np.sum(all_SNPs_within_patients_AR_fSNPs.patient_id == patient_id)
#append info to dataframe
pairwise_variant_distances.loc[patient_id , 'population'] = population
pairwise_variant_distances.loc[patient_id , 'pairwise_AR_fSNP_distance'] = SNP_distance
# -
pairwise_variant_distances.head()
np.shape(pairwise_variant_distances)
# #### Find distance between all pairs of times
for patient_id in pairwise_variant_distances.index:
population = pairwise_variant_distances.loc[patient_id , 'population']
patient_id = patient_id
if population == 'BRYANT':
bryant_pair = time_between_serial_isolate_collection[time_between_serial_isolate_collection.patient_id == patient_id]
first_date = bryant_pair[bryant_pair.sample_order == 1].date_information.values[0]
second_date = bryant_pair[bryant_pair.sample_order == 2].date_information.values[0]
num_days_between = (int(second_date) - int(first_date))*7 #dates are in # of weeks apart
if population == 'CASALI':
casali_pair = time_between_serial_isolate_collection[time_between_serial_isolate_collection.patient_id == patient_id]
first_date = casali_pair[casali_pair.sample_order == 1].date_information.values[0]
day = first_date.split('/')[0]
month = first_date.split('/')[1]
year = '20' + first_date.split('/')[2]
first_date = pd.to_datetime(pd.DataFrame({'year':[year] , 'month':[month] , 'day':[day]}))
second_date = casali_pair[casali_pair.sample_order == 2].date_information.values[0]
day = second_date.split('/')[0]
month = second_date.split('/')[1]
year = '20' + second_date.split('/')[2]
second_date = pd.to_datetime(pd.DataFrame({'year':[year] , 'month':[month] , 'day':[day]}))
num_days_between = (second_date - first_date).dt.days[0]
elif (population == 'CETR') or (population == 'GUERRA') or (population == 'WITNEY') or (population == 'XU'):
serial_pair = time_between_serial_isolate_collection[time_between_serial_isolate_collection.patient_id == patient_id]
first_date = serial_pair[serial_pair.sample_order == 1].date_information.values[0]
day = first_date.split('/')[1]
month = first_date.split('/')[0]
if int(first_date.split('/')[2]) in range(50,100): #some dates in 1990's others in 2000's
year = '19' + first_date.split('/')[2]
else:
year = '20' + first_date.split('/')[2]
first_date = pd.to_datetime(pd.DataFrame({'year':[year] , 'month':[month] , 'day':[day]}))
second_date = serial_pair[serial_pair.sample_order == 2].date_information.values[0]
day = second_date.split('/')[1]
month = second_date.split('/')[0]
if int(second_date.split('/')[2]) in range(50,100):
year = '19' + second_date.split('/')[2]
else:
year = '20' + second_date.split('/')[2]
second_date = pd.to_datetime(pd.DataFrame({'year':[year] , 'month':[month] , 'day':[day]}))
num_days_between = (second_date - first_date).dt.days[0]
elif (population == 'TRAUNER') or (population == 'BRYANT'):
serial_pair = time_between_serial_isolate_collection[time_between_serial_isolate_collection.patient_id == patient_id]
first_date = serial_pair[serial_pair.sample_order == 1].date_information.values[0]
second_date = serial_pair[serial_pair.sample_order == 2].date_information.values[0]
num_days_between = (int(second_date) - int(first_date))*7 #dates are in # of weeks apart
elif population == 'WALKER':
walker_pair = time_between_serial_isolate_collection[time_between_serial_isolate_collection.patient_id == patient_id]
first_date = walker_pair[walker_pair.sample_order == 1].date_information.values[0]
day = '1'
month = '1'
year = first_date
first_date = pd.to_datetime(pd.DataFrame({'year':[year] , 'month':[month] , 'day':[day]}))
second_date = walker_pair[walker_pair.sample_order == 2].date_information.values[0]
day = '1'
month = '1'
year = second_date
second_date = pd.to_datetime(pd.DataFrame({'year':[year] , 'month':[month] , 'day':[day]}))
num_days_between = (second_date - first_date).dt.days[0]
#append distance between isolate collection
pairwise_variant_distances.loc[patient_id , 'time_between_collection'] = num_days_between
pairwise_variant_distances.head()
np.shape(pairwise_variant_distances) #195 subjects with date information for both isolates
pairwise_variant_distances.pairwise_AR_fSNP_distance.sum()
# #### convert time between isolate collection to months (30 days)
pairwise_variant_distances.loc[: , 'time_between_collection'] = pairwise_variant_distances.time_between_collection.astype(float) / 30.0
pairwise_variant_distances.loc[: , 'time_between_collection'] = [round(x, 2) for x in pairwise_variant_distances.time_between_collection] #round the nearest 2nd decimal place
pairwise_variant_distances.head()
# ### Logistic Regression
# Define *Independent* and *Dependent* variables
# +
x = np.array(pairwise_variant_distances.time_between_collection).reshape(-1, 1)
x = x.astype(float)
x = sm.add_constant(x)
y = np.array([int(i) for i in list(pairwise_variant_distances.pairwise_AR_fSNP_distance > 0)])
y = y.astype(float)
# -
# Create a Model and Train It
# +
model = sm.Logit(y, x)
result = model.fit(method='newton')
# -
result.summary()
result.params
result.pvalues
result.conf_int()
coeff_x = result.params[1]
coeff_x_95_CF = result.conf_int()[1,:]
# Calculate Odds Ratio
OR_x = np.exp(coeff_x)
print OR_x
OR_x_95_CF = np.exp(coeff_x_95_CF)
print OR_x_95_CF
# Odds Ratio OR (95% CI) = 1.0229 (1.0016 , 1.0446)
#
# p-val = 0.035
# #################################################################################################################################################################################################################################
# ## [5] Analysis 3 - Clonal Interference Patterns
# #################################################################################################################################################################################################################################
within_host_Base_Call_variants_all_patients_sample_order.head()
np.shape(within_host_Base_Call_variants_all_patients_sample_order)
plt.style.use('ggplot')
plt.rcParams['lines.linewidth']=0
plt.rcParams['axes.facecolor']='1.0'
plt.rcParams['xtick.color']='black'
plt.rcParams['axes.grid']=False
plt.rcParams['axes.edgecolor']='black'
plt.rcParams['grid.color']= '1.0'
plt.rcParams.update({'font.size': 14})
# +
fig , ax = plt.subplots()
n , bins , patches = plt.hist(list(within_host_Base_Call_variants_all_patients_sample_order.alt_AF) , rwidth = 1.0 , edgecolor='white', bins = 75 , linewidth = 1.5 , color = 'black')
plt.plot([.4 , .4] , [0 , max(n)] , 'r--' , linewidth = 1.5)
plt.xlabel('Alternate Allele Frequency', fontweight = 'bold' , fontsize = 12, color = 'black')
plt.ylabel('Number of Base Calls\n(N = ' + str(np.shape(within_host_Base_Call_variants_all_patients_sample_order)[0]) + ')', fontweight = 'bold' , fontsize = 12, color = 'black')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.tick_params(labelcolor = 'k')
ax.set_yscale("log", nonposy='clip')
ax.tick_params(direction='out', length=3, width=1.25, colors='k') #make tick marks darker
ax.set_xlim(-0.01 , 1.01)
fig = plt.gcf()
fig.set_size_inches(10.0, 5.5)
fig.tight_layout()
file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/figures/antibiotic_resistance_analysis/clonal_interference_examples/alt_AF_distribution_all_AR_SNPs.pdf'
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = True)
plt.show()
# -
within_host_Base_Call_variants_all_patients_sample_order.head()
# #### List of subjects that have at least 1 SNP within-host
subject_ids = list( set( list( within_host_Base_Call_variants_all_patients_sample_order.patient_id ) ) )
# +
clonal_interference_SNP_candidates = pd.DataFrame(columns = within_host_Base_Call_variants_all_patients_sample_order.columns)
for subject_id in subject_ids:
within_host_Base_Call_variants_for_patient = within_host_Base_Call_variants_all_patients_sample_order[within_host_Base_Call_variants_all_patients_sample_order.patient_id == subject_id]
for gene_id in list(set(list(within_host_Base_Call_variants_for_patient.gene_id))):
within_host_Base_Call_variants_for_patient_and_geneID = within_host_Base_Call_variants_for_patient[within_host_Base_Call_variants_for_patient.gene_id == gene_id]
#1st AFs
sample_1_AFs = []
#2nd AFs
sample_2_AFs = []
#for each pair of Base Calls, store (alternate Allele Frequency of isolate_1) and (alternate Allele Frequency of isolate_2 - alternate Allele Frequency of isolate_1)
for Base_Call_A_index , Base_Call_B_index in zip( list( within_host_Base_Call_variants_for_patient_and_geneID.iloc[ range(0 , np.shape(within_host_Base_Call_variants_for_patient_and_geneID)[0] , 2) ].index ) , list( within_host_Base_Call_variants_for_patient_and_geneID.iloc[ range(1 , np.shape(within_host_Base_Call_variants_for_patient_and_geneID)[0] , 2) ].index ) ):
#pull alternate AF & sample number for each isolate in pair
if within_host_Base_Call_variants_for_patient_and_geneID.loc[Base_Call_A_index , 'sample_order'] == 1:
Base_Call_H37Rv_AF_1 = within_host_Base_Call_variants_for_patient_and_geneID.loc[Base_Call_A_index , 'alt_AF']
Base_Call_H37Rv_AF_2 = within_host_Base_Call_variants_for_patient_and_geneID.loc[Base_Call_B_index , 'alt_AF']
elif within_host_Base_Call_variants_for_patient_and_geneID.loc[Base_Call_A_index , 'sample_order'] == 2:
Base_Call_H37Rv_AF_2 = within_host_Base_Call_variants_for_patient_and_geneID.loc[Base_Call_A_index , 'alt_AF']
Base_Call_H37Rv_AF_1 = within_host_Base_Call_variants_for_patient_and_geneID.loc[Base_Call_B_index , 'alt_AF']
#store 1st & 2nd AFs in lists
sample_1_AFs.append(Base_Call_H37Rv_AF_1)
sample_2_AFs.append(Base_Call_H37Rv_AF_2)
#see if any AFs > 0.40
if sum(np.array(sample_1_AFs + sample_2_AFs) >= 0.40 ) > 0:
#check to see if AFs for any SNPs within the same subject & gene 'move in' opposite directions
if ( sum( np.array( sample_1_AFs ) < np.array(sample_2_AFs) ) == len(sample_1_AFs) ) or ( sum( np.array( sample_1_AFs ) > np.array(sample_2_AFs) ) == len(sample_1_AFs) ):
pass
else:
clonal_interference_SNP_candidates = clonal_interference_SNP_candidates.append(within_host_Base_Call_variants_for_patient_and_geneID)
#reset index
clonal_interference_SNP_candidates.reset_index(inplace = True , drop = True)
# -
# #### Check for evidence of clonal interference in Intergenic regions
clonal_interference_SNP_candidates[clonal_interference_SNP_candidates.SNP_ftype == 'I']
# No intergenic regions have SNPs that fit the criteria for clonal interference pattern
clonal_interference_SNP_candidates.head()
np.shape(clonal_interference_SNP_candidates)
# Find all gene - subject pairs that show evidence of clonal interference
gene_subject_combos = list( set( [gene_id + ':' + subject_id for gene_id,subject_id in zip( list(clonal_interference_SNP_candidates.gene_id) , list(clonal_interference_SNP_candidates.patient_id) ) ] ) )
for gene_subject in gene_subject_combos:
gene_id = gene_subject.split(':')[0]
subject_id = gene_subject.split(':')[1]
clonal_interference_SNP_candidates_for_gene = clonal_interference_SNP_candidates[clonal_interference_SNP_candidates.gene_id == gene_id]
clonal_interference_SNP_candidates_for_gene_and_subject = clonal_interference_SNP_candidates_for_gene[clonal_interference_SNP_candidates_for_gene.patient_id == subject_id]
#get AFs of first and second sample
AFs_sample_1 = list( clonal_interference_SNP_candidates_for_gene_and_subject[clonal_interference_SNP_candidates_for_gene_and_subject.sample_order == 1].alt_AF )
AFs_sample_2 = list( clonal_interference_SNP_candidates_for_gene_and_subject[clonal_interference_SNP_candidates_for_gene_and_subject.sample_order == 2].alt_AF )
#get time between samples
time_btwn_isolate_collection = pairwise_variant_distances.loc[subject_id , 'time_between_collection']
gene_symbol = list(clonal_interference_SNP_candidates_for_gene_and_subject.gene_symbol)[0]
#get the AA change for each SNP
AA_change_list = []
for sample_1_i , sample_2_i in zip(np.arange(0 , np.shape(clonal_interference_SNP_candidates_for_gene_and_subject)[0] , 2) , np.arange(1 , np.shape(clonal_interference_SNP_candidates_for_gene_and_subject)[0] , 2)):
#get AA change
AA_change_calls = list(clonal_interference_SNP_candidates_for_gene_and_subject.iloc[[sample_1_i , sample_2_i]].AA_change)
try:
AA_change_calls.remove('None')
except ValueError:
pass
AA_change_list.append(AA_change_calls[0])
#get a list of each AA change in decreasing order of AF at 2nd time point (SNP that swept is listed first)
AA_change_and_AF2 = pd.Series(dict(zip(AA_change_list , AFs_sample_2))).sort_values(ascending = False)
AA_change_ordered_by_AF2 = list(AA_change_and_AF2.index)
#label for AA changes
AA_i = 0
AA_change_label = ''
for AA_change in AA_change_ordered_by_AF2:
if AA_i == 0:
AA_change_label = AA_change
AA_i += 1
else:
AA_change_label = AA_change_label + ', ' + AA_change
AA_i += 1
if AA_i % 3 == 0:
AA_change_label = AA_change_label + '\n'
print subject_id + ' : ' + gene_symbol
print AA_change_and_AF2
print '-----------------------'
####################################################################################################################################################################################
fig , ax = plt.subplots()
for SNP_i_AF1 , SNP_i_AF2 in zip(AFs_sample_1 , AFs_sample_2):
plt.scatter([0 , 1] , [SNP_i_AF1 , SNP_i_AF2] , linewidth = 2.25 , edgecolor = 'k' , color = '0.8' , s = 35 , alpha = 0.90)
plt.plot([0 , 1] , [SNP_i_AF1 , SNP_i_AF2] , linewidth = 0.5 , color = '0.0' , linestyle = '-')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.set_xticks([0 , 1]) #xtick positions
ax.set_xticklabels(['0' , str(time_btwn_isolate_collection)]) #xtick labels
#set y limit to between 0 & 1
ax.set_ylim(-0.02, 1.02)
ax.set_yticks([0, 0.2, 0.4, 0.6, 0.8, 1.0]) #ytick positions
ax.set_yticklabels([0, 0.2, 0.4, 0.6, 0.8, 1.0])
ax.tick_params(labelcolor = 'k')
plt.xlabel('Months Between Collection\n' + AA_change_label, fontsize = 12 , color = 'k')
plt.ylabel('Alternate Allele Frequency' , fontweight = 'bold', fontsize = 12 , color = 'k')
plt.title(subject_id + ' : ' + gene_symbol, fontweight = 'bold' , fontsize = 14 , color = 'k')
fig = plt.gcf()
fig.set_size_inches(3.0, 6.5)
fig.tight_layout()
file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/figures/antibiotic_resistance_analysis/clonal_interference_examples/clonal_intererence_' + gene_subject + '.pdf'
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = True)
plt.show()
# #################################################################################################################################################################################################################################
# ## [6] Analysis 4 - Treatment Failure Subjects Analysis
# #################################################################################################################################################################################################################################
# ### *Subset* to treatment failure group: $>60$ days / 2 months between isolate collection
pairwise_variant_distances_treatment_failure = pairwise_variant_distances[pairwise_variant_distances.time_between_collection >= 1.7]
np.shape(pairwise_variant_distances_treatment_failure)
# #### Find proportion of isolates that had an AR fSNP
float( np.shape(pairwise_variant_distances_treatment_failure[pairwise_variant_distances_treatment_failure.pairwise_AR_fSNP_distance > 0])[0] )
float(np.shape(pairwise_variant_distances_treatment_failure)[0])
# List of patient IDs that had at least 1 **AR fSNP**
print(list(pairwise_variant_distances_treatment_failure[pairwise_variant_distances_treatment_failure.pairwise_AR_fSNP_distance > 0].index))
# proportion of subjects with $> 2$ months isolate collection that had an AR SNP sweep between samples
float( np.shape( pairwise_variant_distances_treatment_failure[pairwise_variant_distances_treatment_failure.pairwise_AR_fSNP_distance > 0] )[0] ) / float(np.shape(pairwise_variant_distances_treatment_failure)[0])
# #### Get all positions that are associated with AR from Farhat et. al. 2016 (minimal set of 238 AR variants)
# +
filepath = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/predictive_AR_mutations/Table_S17_238_DR_mutations.txt'
AR_variants_DF = pd.DataFrame(columns = ['variant_type' , 'region_type' , 'ref_position'])
with open(filepath) as fp:
for cnt, line in enumerate(fp):
#print("Line {}: {}".format(cnt, line))
AR_variants_DF.loc[cnt , :] = [line.split('_')[0] , line.split('_')[1] , int(line.split('_')[2])]
#sort values by Reference Position
AR_variants_DF.sort_values(by = 'ref_position' , ascending = True , inplace = True)
#drop duplicate reference positions
duplicated_variants = list( AR_variants_DF[AR_variants_DF.duplicated(subset = 'ref_position' , keep = 'first')].index )
AR_variants_DF.drop(duplicated_variants , axis = 0 , inplace = True)
#drop any variant that isn't a SNP and re-index
AR_variants_DF = AR_variants_DF[AR_variants_DF.variant_type == 'SNP']
AR_variants_DF.reset_index(drop = True , inplace = True)
# -
np.shape(AR_variants_DF)
AR_variants_DF.head()
H37Rv_positions_of_interest = list( AR_variants_DF.ref_position )
# #### Also load variants for *Rifampicin* and *Isoniazid* resistance from Farhat et. al. 2016
# SNPs associated with *RIF* resistance
# +
filepath = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/predictive_AR_mutations/Table_S17_238_DR_mutations_RIF.txt'
AR_RIF_variants_DF = pd.DataFrame(columns = ['variant_type' , 'region_type' , 'ref_position' , 'ref_allele' , 'alt_allele'])
with open(filepath) as fp:
for cnt, line in enumerate(fp):
#print("Line {}: {}".format(cnt, line))
AR_RIF_variants_DF.loc[cnt , :] = [line.split('_')[0] , line.split('_')[1] , int(line.split('_')[2]) , line.split('_')[3][0] , line.split('_')[3][1]]
#sort values by Reference Position
AR_RIF_variants_DF.sort_values(by = 'ref_position' , ascending = True , inplace = True)
#drop any variant that isn't a SNP and re-index
AR_RIF_variants_DF = AR_RIF_variants_DF[AR_RIF_variants_DF.variant_type == 'SNP']
AR_RIF_variants_DF.reset_index(drop = True , inplace = True)
# -
AR_RIF_variants_DF.head()
np.shape(AR_RIF_variants_DF)
# SNPs associated with *INH* resistance
# +
filepath = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/predictive_AR_mutations/Table_S17_238_DR_mutations_INH.txt'
AR_INH_variants_DF = pd.DataFrame(columns = ['variant_type' , 'region_type' , 'ref_position' , 'ref_allele' , 'alt_allele'])
with open(filepath) as fp:
for cnt, line in enumerate(fp):
#print("Line {}: {}".format(cnt, line))
AR_INH_variants_DF.loc[cnt , :] = [line.split('_')[0] , line.split('_')[1] , int(line.split('_')[2]) , line.split('_')[3][0] , line.split('_')[3][1]]
#sort values by Reference Position
AR_INH_variants_DF.sort_values(by = 'ref_position' , ascending = True , inplace = True)
#drop any variant that isn't a SNP and re-index
AR_INH_variants_DF = AR_INH_variants_DF[AR_INH_variants_DF.variant_type == 'SNP']
AR_INH_variants_DF.reset_index(drop = True , inplace = True)
# -
AR_INH_variants_DF.head()
np.shape(AR_INH_variants_DF)
# Create list of Rifampicin & Isonazid Resistance mutations from DataFrames
# +
#list of RIF mutations
AR_RIF_variants = [str(ref_pos) + '_' + alt_allele for ref_pos, alt_allele in zip(AR_RIF_variants_DF.ref_position , AR_RIF_variants_DF.alt_allele)]
#list of INH mutations
AR_INH_variants = [str(ref_pos) + '_' + alt_allele for ref_pos, alt_allele in zip(AR_INH_variants_DF.ref_position , AR_INH_variants_DF.alt_allele)]
# -
# ### *Function* to Extract SNPs from VCF file
def SNPs_from_VCF(VCF_file):
vcf_reader = vcf.Reader(open(VCF_file , 'r'))
#create dictionaries to store information for each call
ref_bases = {}
qry_bases = {}
ref_positions = {}
INFO_for_call = {}
#indexer for dataframe containing SNPs
index = 0
#iterate through each Variant Call
for record in vcf_reader:
#check to see if the call is a PASS by Pilon
if record.FILTER == []:
#check to see if SNP is located in a region associated with Antibiotic Resistance
if record.POS in H37Rv_positions_of_interest:
#check to see if variant is SNP; length of reference allele is 1, there is only 1 alternate allele, length of alternate allele is 1
if (len(record.REF) == 1) and (len(record.ALT) == 1) and (len(str( record.ALT[0] )) == 1):
##### Retrieve Relevant information for filtering quality of Base Call #####
# Mean Base Quality @ locus
BQ = record.INFO['BQ']
# Mean Mapping Quality @ locus
MQ = record.INFO['MQ']
# Number of Reads w/ Deletion
DC = record.INFO['DC']
# Number of Reads w/ Insertion
IC = record.INFO['IC']
# Depth of Valid Reads in Pileup
VD = record.INFO['DP']
### Filtering Criteria
#---> Mean Base Quality > 20
#---> Mean Mapping Quality > 30
#---> No Reads Supporting Insertions
#---> No Reads Supporting Deletions
#---> Number of High Quality Reads >= 25
if (BQ > 20) and (MQ > 30) and (DC == 0) and (IC == 0) and (VD >= 25): #SNP passed all filtering criteria!
# Filtering Criteria for mutant allele frequency calculation (range: 0.75 - 1.0)
ref_allele = str(record.REF)
alt_allele = str(record.ALT[0])
#After extensive filtering and categorization, store all of the pertinent information about the Base Call
ref_bases[index] = ref_allele
qry_bases[index] = alt_allele
ref_positions[index] = record.POS
INFO_for_call[index] = record.INFO
index += 1
#convert dictionaries to series
ref_bases = pd.Series(ref_bases)
qry_bases = pd.Series(qry_bases)
ref_positions = pd.Series(ref_positions)
INFO_for_call = pd.Series(INFO_for_call)
#create DataFrame to hold all base calls for a given sample
Variant_Call_DF = pd.DataFrame()
Variant_Call_DF['ref_base'] = ref_bases
Variant_Call_DF['alt_base'] = qry_bases
Variant_Call_DF['ref_position'] = ref_positions
Variant_Call_DF['INFO'] = INFO_for_call
return Variant_Call_DF #DataFrame for base calls for a single isolate
# ### *Functions* to Annotate SNPs
# #### Cell to annotate SNPs
# +
# Important Packages
################################################################################################################################################################################################
import os
import pandas as pd
import numpy as np
import sys
import pickle
import Bio
from Bio.Alphabet import IUPAC
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
from Bio.SeqFeature import SeqFeature, FeatureLocation
from Bio import SeqIO
from StringIO import StringIO
from Bio import AlignIO
from Bio.Align import AlignInfo
from Bio.Seq import MutableSeq
################################################################################################################################################################################################
# Relevant Information for H37Rv sequence SNP functional annotation
################################################################################################################################################################################################
####### Collect all DNA and Amino Acid sequences corresponding to genes on H37Rv #######
#load reference genome and reference annotation
reference_genome = '/n/data1/hms/dbmi/farhat/bin/work-horse/bin/h37rv.fasta'
for reference_genome in SeqIO.parse(reference_genome, "fasta"):
reference_genome.seq.alphabet = IUPAC.unambiguous_dna
reference_genome_annotation = pd.read_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/H37Rv/h37rv_genome_summary.txt', '\t').set_index('name')
####### Function to translate coding DNA sequences #######
def translate(gene_id, sequence):
#find which strand the gene is located on and translate
strand = reference_genome_annotation.loc[gene_id, 'strand']
if strand == '+':
protein_sequence = sequence.translate(table="Bacterial", cds=False)
elif strand == '-':
protein_sequence = sequence.reverse_complement().translate(table="Bacterial", cds=False)
return protein_sequence
####### Load in dictionaries for SNP annotation #######
with open('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/pickled_files/dicts_for_SNP_annotation/H37Rv_gene_seq_records.pickle', 'rb') as handle:
ref_gene_sequences_records = pickle.load(handle)
with open('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/pickled_files/dicts_for_SNP_annotation/H37Rv_protein_seq_records.pickle', 'rb') as handle:
ref_protein_sequences_records = pickle.load(handle)
with open('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/pickled_files/dicts_for_SNP_annotation/H37Rv_coord_gene_mapping.pickle', 'rb') as handle:
ReferencePosition_Gene_mapping = pickle.load(handle)
####### get Gene Categories #######
gene_categories = pd.read_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/gene_categories/gene_categories.csv').set_index('name')
gene_categories_dict = dict([gene_id , gene_category] for gene_id, gene_category in zip(list(gene_categories.gene_id) , list(gene_categories.Gene_Category)))
####### get Gene Symbols #######
gene_symbol_dict = dict([gene_id , gene_symbol] for gene_id, gene_symbol in zip(list(reference_genome_annotation.symbol.index) , list( reference_genome_annotation.symbol )))
################################################################################################################################################################################################
# Function to annotate Intergenic SNPs
################################################################################################################################################################################################
def find_flanking_genes_for_intergenic_region(intergenic_ref_pos):
#this function finds the genes flagging an intergenic region given a reference position
#find gene immediately in the 5' direction
for i in range(0 , 100000):
#move toward 5' direction
if ReferencePosition_Gene_mapping[intergenic_ref_pos - i] != []:
gene_to_left = ReferencePosition_Gene_mapping[intergenic_ref_pos - i][0]
break
#find gene immediately in the 3' direction
for i in range(0 , 100000):
#move toward 3' direction
try:
if ReferencePosition_Gene_mapping[intergenic_ref_pos + i] != []:
gene_to_right = ReferencePosition_Gene_mapping[intergenic_ref_pos + i][0]
break
#KeyError means we have hit the 'end' of the chromosome, the intergenic region at then end of H37Rv in 5' > 3' orientation
#since TB chromosome is circular the gene to the 'right' is Rv0001
except KeyError:
gene_to_right = 'Rv0001'
break
return gene_to_left + '_' + gene_to_right
################################################################################################################################################################################################
# Function to determine whether SNPs are Synonymous or Non-Synonymous; Returns gene coordinate, codon position, AA changes, Gene Category & Symbol
################################################################################################################################################################################################
def SNP_annotate(ref_seq_position , alt_allele_i):
'''
This function takes as input a reference position on H37Rv located within a
gene and an alternate allele and returns whether the base change
would correspond to a different Amino Acid sequence that results
from translating the DNA sequence into an AA sequence.
'''
gene_intergenic_id_list = []
genomic_coord_list = []
gene_category_list = []
gene_symbol_list = []
Syn_NSyn_list = []
AA_change_list = []
#get the Reference Allele from the complete H37Rv reference genome, indexing starts from 0
ref_allele_i = reference_genome.seq[int(ref_seq_position) - 1]
#find the gene that SNP occurs on; check list corresponding to H37Rv coordinate to see if there are any genes associated with RefPosition
if len(ReferencePosition_Gene_mapping[ref_seq_position]) > 0:
#iterate through all genes that ReferencePosition is mapped to (i.e. SNP might correspond to 2 genes)
for gene_intergenic_id in ReferencePosition_Gene_mapping[ref_seq_position]:
#find genomic coordinate of SNP relative to gene (subtract 1 since reference seq starts counting at 1)
gene_relative_coord = (ref_seq_position - 1) - min( reference_genome_annotation.loc[gene_intergenic_id , 'chromStart'] , reference_genome_annotation.loc[gene_intergenic_id , 'chromEnd'] )
#find the genomic coordinate (relative to the gene, in the 5' to 3' direction)
strand = reference_genome_annotation.loc[gene_intergenic_id, 'strand']
if strand == '+':
genomic_5_to_3_coord = (ref_seq_position) - reference_genome_annotation.loc[gene_intergenic_id , 'chromStart']
elif strand == '-':
genomic_5_to_3_coord = (reference_genome_annotation.loc[gene_intergenic_id , 'chromEnd']) - (ref_seq_position-1)
#find gene category (if one exists)
try:
gene_category_i = gene_categories_dict[gene_intergenic_id]
except KeyError:
gene_category_i = 'None'
#find gene symbol (if one exists)
try:
gene_symbol_i = gene_symbol_dict[gene_intergenic_id]
except KeyError:
gene_symbol_i = 'None'
#alternate allele is an actual base
if alt_allele_i in ['A','C','G','T']:
#translate into protein sequence with the SNP in place if not InDel or intergenic region
SNP_change = alt_allele_i
#ALTERNATE allele (is it Syn or NSyn?)
#get sequence from dictionary of sequences (and convert to mutable object)
test_gene_sequence = ref_gene_sequences_records[gene_intergenic_id].seq.tomutable()
#change reference gene sequence by the SNP in the query sequence
test_gene_sequence[int(gene_relative_coord)] = SNP_change
#convert back immutable object
test_gene_sequence = test_gene_sequence.toseq()
#translate sequence into amino acid seq
test_protein_sequence = translate(gene_intergenic_id , test_gene_sequence)
#store the H37Rv AA seq to compare against
H37Rv_AA_sequence = ref_protein_sequences_records[gene_intergenic_id].seq
#get the codon number where the SNP occurs within
## take the genomic coordinate (relative to the gene, in the 5' to 3' direction), divide by 3, then take the ceiling of this number (will be fraction if SNP occurs in 1st or 2nd position on codon)
strand = reference_genome_annotation.loc[gene_intergenic_id, 'strand']
if strand == '+':
genomic_5_to_3_coord = (ref_seq_position) - reference_genome_annotation.loc[gene_intergenic_id , 'chromStart']
elif strand == '-':
genomic_5_to_3_coord = (reference_genome_annotation.loc[gene_intergenic_id , 'chromEnd']) - (ref_seq_position-1)
codon_coord = int(np.ceil( float( genomic_5_to_3_coord) / 3.0 ))
#compare to AA seq of original gene
if test_protein_sequence == H37Rv_AA_sequence:
SNP_type = 'S'
#get the AA before & after
AA_change = H37Rv_AA_sequence[codon_coord-1] + str(codon_coord) + test_protein_sequence[codon_coord-1]
else:
SNP_type = 'N'
#get the AA before & after
AA_change = H37Rv_AA_sequence[codon_coord-1] + str(codon_coord) + test_protein_sequence[codon_coord-1]
#alternate allele is a dummy (Base Call completely supports the Reference Allele)
else:
SNP_type = 'None'
AA_change = 'None'
#store relevant info in lists
gene_intergenic_id_list.append(gene_intergenic_id)
genomic_coord_list.append(genomic_5_to_3_coord)
gene_category_list.append(gene_category_i)
gene_symbol_list.append(gene_symbol_i)
Syn_NSyn_list.append(SNP_type)
AA_change_list.append(AA_change)
#if no gene in H37Rv corresponds to the Reference Position for SNP, then SNP must be intergenic
else:
gene_intergenic_id = find_flanking_genes_for_intergenic_region(ref_seq_position)
genomic_5_to_3_coord = 'None'
gene_category_i = 'None'
gene_symbol_i = 'None'
SNP_type = 'I'
AA_change = 'None'
#store relevant info in lists
gene_intergenic_id_list.append(gene_intergenic_id)
genomic_coord_list.append(genomic_5_to_3_coord)
gene_category_list.append(gene_category_i)
gene_symbol_list.append(gene_symbol_i)
Syn_NSyn_list.append(SNP_type)
AA_change_list.append(AA_change)
#if there is only a single gene associated with this SNP, just return the individual elememts
if len(gene_intergenic_id_list) == 1:
return [ref_allele_i , gene_intergenic_id , genomic_5_to_3_coord , gene_category_i , gene_symbol_i , SNP_type , AA_change]
#else if there are two genes associated with this SNP, return elements for each SNP annotation in a list
elif len(gene_intergenic_id_list) > 1:
return [ref_allele_i , gene_intergenic_id_list , genomic_coord_list , gene_category_list , gene_symbol_list , Syn_NSyn_list , AA_change_list]
################################################################################################################################################################################################
# -
# ### Get the isolate tags for the first isolate collected for each subject for group with $>60$ days / $2$ months between isolate collection
patients_to_keep = list(pairwise_variant_distances_treatment_failure.index)
len( patients_to_keep )
isolate_tags_for_first_sample = sample_annotation[sample_annotation.sample_order == 1].loc[patients_to_keep , :]
np.shape(isolate_tags_for_first_sample)
isolate_tags_for_first_sample.head()
# ### Iterate through Reduced VCF corresponding to the 1st sample from each subject and collect all AR SNPs if present
# +
#create a dictionary to hold the AR SNPs detected in the first isolate for each subject
AR_SNPs_1st_sample_dict = {}
all_AR_SNPS_1st_sample_df = pd.DataFrame()
isolate_i = 0
for isolate_tag in list(isolate_tags_for_first_sample.tag):
#path to Reduced VCF file
Reduced_VCF_file = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/JankyPipe/reduced_VCF_files_for_AR_SNP_detection_longitudinal_isolates/' + isolate_tag + '/' + isolate_tag + '_REDUCED.vcf'
#pull SNPs from VCF file
SNPs_from_isolate = SNPs_from_VCF(Reduced_VCF_file)
################################################################################
### Annotate SNPs
################################################################################
gene_id_list = []
gene_coord_list = []
gene_category_list = []
gene_symbol_list = []
SNP_ftype_list = []
AA_change_list = []
#Annotate Filtered Base Calls (make sure there is at least 1 SNP)
if np.shape(SNPs_from_isolate)[0] > 0:
for ref_position_i , alt_base_i in zip(list(SNPs_from_isolate.ref_position) , list(SNPs_from_isolate.alt_base)):
#annotate SNP
gene_id_i , gene_coord_i , gene_category_i , gene_symbol_i , SNP_ftype_i , AA_change_i = SNP_annotate(ref_position_i , alt_base_i)[1:]
gene_id_list.append(gene_id_i)
gene_coord_list.append(gene_coord_i)
gene_category_list.append(gene_category_i)
gene_symbol_list.append(gene_symbol_i)
SNP_ftype_list.append(SNP_ftype_i)
AA_change_list.append(AA_change_i)
#create columns to store SNP annotation info
SNPs_from_isolate['gene_id'] = gene_id_list
SNPs_from_isolate['gene_coord'] = gene_coord_list
SNPs_from_isolate['gene_category'] = gene_category_list
SNPs_from_isolate['gene_symbol'] = gene_symbol_list
SNPs_from_isolate['SNP_ftype'] = SNP_ftype_list
SNPs_from_isolate['AA_change'] = AA_change_list
#No predictive AR SNPs detected from this isolate (empty DataFrame)
else:
SNPs_from_isolate['gene_id'] = ""
SNPs_from_isolate['gene_coord'] = ""
SNPs_from_isolate['gene_category'] = ""
SNPs_from_isolate['gene_symbol'] = ""
SNPs_from_isolate['SNP_ftype'] = ""
SNPs_from_isolate['AA_change'] = ""
#drop synonymous SNPs & re-index
SNPs_from_isolate = SNPs_from_isolate[SNPs_from_isolate.SNP_ftype != 'S']
SNPs_from_isolate.reset_index(inplace = True , drop = True)
#add column to patient_id & isolate tag
patient_id = sample_annotation[sample_annotation.tag == isolate_tag].index[0]
isolate_tag = sample_annotation[sample_annotation.tag == isolate_tag].tag[0]
SNPs_from_isolate['patient_id'] = [patient_id]*np.shape(SNPs_from_isolate)[0]
SNPs_from_isolate['isolate_tag'] = [isolate_tag]*np.shape(SNPs_from_isolate)[0]
#store AR SNPs in dict
AR_SNPs_1st_sample_dict[patient_id] = SNPs_from_isolate
#create a DataFrame that stores all AR SNPs detected in the 1st sample of each serial pair
all_AR_SNPS_1st_sample_df = all_AR_SNPS_1st_sample_df.append(SNPs_from_isolate)
isolate_i += 1
if isolate_i % 20 == 0:
print isolate_i
#reset index for DataFrame containing all AR SNPs in first clinical isoaltes for each serial pair
all_AR_SNPS_1st_sample_df.reset_index(inplace = True , drop = True)
# -
all_AR_SNPS_1st_sample_df.head()
np.shape(all_AR_SNPS_1st_sample_df)
sum( all_AR_SNPS_1st_sample_df.SNP_ftype == 'S' ) #check to make sure no Synonymous SNPs
# #### Filter out any *gid* E92D mutations since these are likely lineage markers
# +
non_gid_E92D_SNPs_filter = [not ((all_AR_SNPS_1st_sample_df.loc[SNP_i, :].AA_change == 'E92D') and (all_AR_SNPS_1st_sample_df.loc[SNP_i, :].gene_id == 'Rv3919c')) for SNP_i in all_AR_SNPS_1st_sample_df.index]
all_AR_SNPS_1st_sample_df = all_AR_SNPS_1st_sample_df[non_gid_E92D_SNPs_filter]
#reset index
all_AR_SNPS_1st_sample_df.reset_index(inplace = True , drop = True)
# -
np.shape(all_AR_SNPS_1st_sample_df)
# Output as a CSV file
all_AR_SNPS_1st_sample_df.to_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/antibiotic_resistance_longitudinal_SNPs/AR_SNPs_1st_isolate_failure_relapse_subjects.csv', sep = ',')
# #### Count the number of AR SNPs in first collected isolate for each subject (with at least 1 AR SNP in first isolate collected)
from collections import Counter
AR_SNPs_in_1st_isolate_per_subject = pd.Series( Counter(list(all_AR_SNPS_1st_sample_df.patient_id)) )
len(AR_SNPs_in_1st_isolate_per_subject)
AR_SNPs_in_1st_isolate_per_subject.head()
# #### Add in 'zero' count for subjects with no AR SNPs in first collected isolate
for subject_id in patients_to_keep:
if subject_id not in list(AR_SNPs_in_1st_isolate_per_subject.index):
AR_SNPs_in_1st_isolate_per_subject[subject_id] = 0
len(AR_SNPs_in_1st_isolate_per_subject)
min(AR_SNPs_in_1st_isolate_per_subject)
max(AR_SNPs_in_1st_isolate_per_subject)
# +
fig , ax = plt.subplots()
n , bins , patches = plt.hist(AR_SNPs_in_1st_isolate_per_subject , rwidth = 1.0 , edgecolor='white', bins = range(0 , max(AR_SNPs_in_1st_isolate_per_subject) + 2) , linewidth = 2.0 , color = 'k' , align = 'left')
plt.ylabel('Number of Subjects', fontweight = 'bold' , fontsize = 12, color = 'k')
plt.xlabel('Number of AR SNPs in 1st isolate', fontweight = 'bold' , fontsize = 12, color = 'k')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.set_xticks(range(0 , max(AR_SNPs_in_1st_isolate_per_subject) + 1 , 1)) #xtick positions
ax.set_xticklabels([str(x) for x in range(0 , max(AR_SNPs_in_1st_isolate_per_subject) + 1 , 1)]) #xtick labels
ax.tick_params(labelcolor = 'k')
ax.tick_params(direction='out', length=3, width=1.25, colors='k') #make tick marks darker
ax.set_xlim(-0.6 , max(AR_SNPs_in_1st_isolate_per_subject) + 0.6)
fig = plt.gcf()
fig.set_size_inches(10.0, 5.5)
fig.tight_layout()
file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/figures/antibiotic_resistance_analysis/number_AR_SNPs_in_1st_isolate_failure_relapse_subjects.pdf'
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = True)
plt.show()
# -
# #### SNPs most represented in 1st collected isolate
pd.Series( Counter( all_AR_SNPS_1st_sample_df.ref_position ) ).sort_values(ascending = False).head(n = 20)
# #### Find out how many subjects had MDR TB for the 1st collected isolate
# +
patients_with_MDR = []
for patient_id in patients_to_keep:
RIF_res = False
INH_res = False
#subset to AR SNPs detected in 1st collected isolate
all_AR_SNPS_1st_sample_df_for_patient = all_AR_SNPS_1st_sample_df[all_AR_SNPS_1st_sample_df.patient_id == patient_id]
#create list of Ref Positions + Alternate Allele
all_AR_SNPS_1st_sample_for_patient = [str(int(ref_pos)) + '_' + alt_allele for ref_pos, alt_allele in zip(all_AR_SNPS_1st_sample_df_for_patient.ref_position , all_AR_SNPS_1st_sample_df_for_patient.alt_base)]
#check for SNP assoc. with RIF & INH resistance
for AR_SNP in all_AR_SNPS_1st_sample_for_patient:
if AR_SNP in AR_RIF_variants:
RIF_res = True
elif AR_SNP in AR_INH_variants:
INH_res = True
#store patient in list of the patient has at least 1 RIF SNP and 1 INH SNP in 1st collected isolate
if ((RIF_res == True) and (INH_res == True)):
patients_with_MDR.append(patient_id)
# -
print patients_with_MDR
len(patients_with_MDR)
len(patients_to_keep)
print 'Proportion of Subjects with MDR TB = ' + str(float(len(patients_with_MDR)) / float(len(patients_to_keep)))
# ### Get the number of AR SNPs that cropped up between longitudinal isolates (with $AF \ge 40\%$) from subjects with treatment failure
AR_SNPs_between_isolates_per_subject = pd.Series()
for subject_id in list(AR_SNPs_in_1st_isolate_per_subject.index):
AR_SNPs_between_isolates_per_subject[subject_id] = np.sum(all_SNPs_within_patients_AR_fSNPs.patient_id == subject_id)
AR_SNPs_between_isolates_per_subject.head()
len(AR_SNPs_between_isolates_per_subject)
# ### Test for association between pre-existing AR & AR SNP acquisition in *failure* subjects
# #### Split subjects into 2 groups, subjects with no AR SNPs in 1st isolate (Group B1) & subjects with at least 1 AR SNP in 1st isolate (Group B2)
group_B1_subjects = list( AR_SNPs_in_1st_isolate_per_subject[AR_SNPs_in_1st_isolate_per_subject == 0].index )
group_B2_subjects = list( AR_SNPs_in_1st_isolate_per_subject[AR_SNPs_in_1st_isolate_per_subject > 0].index )
len(group_B1_subjects)
len(group_B2_subjects)
# #### Group B1
# Subjects with no AR SNPs in 1st collected longitudinal sample who also had no AR SNPs crop up between isolate collection
sum( AR_SNPs_between_isolates_per_subject[group_B1_subjects] == 0 )
# Subjects with no AR SNPs in 1st collected longitudinal sample who also had at least 1 AR SNP crop up between isolate collection
sum( AR_SNPs_between_isolates_per_subject[group_B1_subjects] > 0 )
float(7) / float(105)
# #### Group B2
# Subjects who had at least 1st collected longitudinal sample who also had no AR SNPs crop up between isolate collection
sum( AR_SNPs_between_isolates_per_subject[group_B2_subjects] == 0 )
# Subjects who had at least 1st collected longitudinal sample who also had at least 1 AR SNP crop up between isolate collection
sum( AR_SNPs_between_isolates_per_subject[group_B2_subjects] > 0 )
float(20) / float(73)
# #### Fisher exact test to test independence
AR_SNP_cont_table = np.array([[98,7] , [53,20]])
AR_SNP_cont_table
oddsratio, pvalue = stats.fisher_exact(AR_SNP_cont_table) #rows: (1) subjects with no AR SNPs in 1st isolate (2) subjects with at least 1 AR SNP in 1st isolate ; columns: (1) no AR SNPs between isolate collection (2) at least 1 AR SNP between isolate collection
oddsratio
pvalue
# ### Further analysis on *B2* group that acquired an AR SNP between sample collection
# - $\frac{27}{178} \approx 15\%$ treatment failure subjects had an AR fSNP between sampling
#
# - $\frac{20}{27} \approx 74\%$ of cases of AR fSNP acquisition (e.g. *Resistance Amplification*) occurred in the **B2** group (isolates genotypically resistant at the outset).
#
# - Of the $\frac{20}{27} \approx 74\%$ subjects, how many developed an AR fSNP that resulted in a resistance mutation to an **additional** drug, the acqusition of an AR fSNP that incurs resistance to a drug for which no resistance mutations were detected at the outset?
# Get the subject IDs for subjects that:
# 1. Had sample collections time $\ge 2$ months (classified as treatment failure)
# 2. Were genotypically resistant at the outset ($\ge 1$ pAR SNP detected in 1st isolate)
# 3. Developed Resistance Amplification ($\ge 1$ AR fSNP between sample collection)
subjects_treatment_failure_AND_genotypically_resistant_outset_AND_AR_fSNP_acquisition = AR_SNPs_between_isolates_per_subject[group_B2_subjects][AR_SNPs_between_isolates_per_subject[group_B2_subjects] > 0]
len(subjects_treatment_failure_AND_genotypically_resistant_outset_AND_AR_fSNP_acquisition)
# Annotation for Antibiotic Resistance Loci
H37Rv_regions_of_interest = pd.read_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/gene_categories/H37Rv_AR_regions_of_interest.csv' , sep = ',').set_index('Unnamed: 0')
H37Rv_regions_of_interest.index.rename('' , inplace = True)
H37Rv_regions_of_interest.head()
# #### Collect all H37Rv locus tags with (AR SNPs at the outset) or (AR SNPs acquired between sampling)
# +
gene_locus_tag_list = []
for subject_id in subjects_treatment_failure_AND_genotypically_resistant_outset_AND_AR_fSNP_acquisition.index:
#collect genes with SNPs for this subject
genes_with_resistance_SNPs_outset_subject_i = list(set(all_AR_SNPS_1st_sample_df[all_AR_SNPS_1st_sample_df.patient_id == subject_id].gene_id)) #genes with SNPs at outset
genes_with_resistance_SNPs_acquired_subject_i = list(set(all_SNPs_within_patients_AR_fSNPs[all_SNPs_within_patients_AR_fSNPs.patient_id == subject_id].gene_id)) #genes with SNPs acquired between samples
#get genes with AR SNPs at outset across subset of subjects
for gene_locus_tag in genes_with_resistance_SNPs_outset_subject_i:
gene_locus_tag_list.append(gene_locus_tag)
#get genes with AR SNPs acquired across subset of subjects
for gene_locus_tag in genes_with_resistance_SNPs_acquired_subject_i:
gene_locus_tag_list.append(gene_locus_tag)
gene_locus_tag_list = list(set(gene_locus_tag_list))
# -
print gene_locus_tag_list
# Create a dictionary that contains the symbol for each locus tag
gene_locus_tag_symbol_dict = {
'Rv0006':'gyrA',
'Rv0341':'iniB',
'Rv0005':'gyrB',
'Rv3793_Rv3794':'promoter_embA-embB',
'Rv0682':'rpsL',
'Rv3854c':'ethA',
'Rv2043c':'pncA',
'Rv3806c':'ubiA',
'Rv3795':'embB',
'Rv3794':'embA',
'Rv3793':'embC',
'Rv0668':'rpoC',
'Rv1694':'tlyA',
'Rv1908c':'katG',
'Rv1484':'inhA',
'Rv2447c':'folC',
'Rv3919c':'gid',
'Rv2245':'kasA',
'Rv0667':'rpoB',
'Rv1482c_Rv1483':'promoter_fabG1-inhA'
}
# #### Collect all locus symbols with (AR SNPs at the outset) or (AR SNPs acquired between sampling) for each subject
# +
genes_with_resistance_SNPs_outset = []
genes_with_resistance_SNPs_acquired = []
for subject_id in subjects_treatment_failure_AND_genotypically_resistant_outset_AND_AR_fSNP_acquisition.index:
#collect genes with SNPs for this subject
genes_with_resistance_SNPs_outset_subject_i = list(set(all_AR_SNPS_1st_sample_df[all_AR_SNPS_1st_sample_df.patient_id == subject_id].gene_id)) #genes with SNPs at outset
genes_with_resistance_SNPs_acquired_subject_i = list(set(all_SNPs_within_patients_AR_fSNPs[all_SNPs_within_patients_AR_fSNPs.patient_id == subject_id].gene_id)) #genes with SNPs acquired between samples
#get symbols for genes
gene_symbols_with_resistance_SNPs_outset_subject_i = []
for gene_locus_tag in genes_with_resistance_SNPs_outset_subject_i:
gene_symbols_with_resistance_SNPs_outset_subject_i.append(gene_locus_tag_symbol_dict[gene_locus_tag])
gene_symbols_with_resistance_SNPs_acquired_subject_i = []
for gene_locus_tag in genes_with_resistance_SNPs_acquired_subject_i:
gene_symbols_with_resistance_SNPs_acquired_subject_i.append(gene_locus_tag_symbol_dict[gene_locus_tag])
#store gene symbols for genes with pre-existing AR SNPs & acquired AR SNPs
genes_with_resistance_SNPs_outset.append(gene_symbols_with_resistance_SNPs_outset_subject_i)
genes_with_resistance_SNPs_acquired.append(gene_symbols_with_resistance_SNPs_acquired_subject_i)
#create DataFrame
treatment_failure_AR_pSNP_fSNP_subjects_genes = pd.DataFrame(index = subjects_treatment_failure_AND_genotypically_resistant_outset_AND_AR_fSNP_acquisition.index)
treatment_failure_AR_pSNP_fSNP_subjects_genes['genes with AR pSNPs'] = genes_with_resistance_SNPs_outset
treatment_failure_AR_pSNP_fSNP_subjects_genes['genes with AR fSNPs'] = genes_with_resistance_SNPs_acquired
# -
treatment_failure_AR_pSNP_fSNP_subjects_genes
for subject_id in treatment_failure_AR_pSNP_fSNP_subjects_genes.index:
print subject_id
print treatment_failure_AR_pSNP_fSNP_subjects_genes.loc[subject_id , 'genes with AR pSNPs']
print treatment_failure_AR_pSNP_fSNP_subjects_genes.loc[subject_id , 'genes with AR fSNPs']
print ''
# #### Subjects above that developed resistance to an *additional* antibiotic
# **1700** -
# - pAR SNPs in genes = embB, pncA, rpoB, inhA
# - AR fSNP in genes = katG
#
# **1846** +
# - pAR SNPs in genes = pncA, embB, embA, embC, kasA, rpoB
# - AR fSNP in genes = gyrA
#
# **1960** +
# - pAR SNPs in genes = iniB, katG, gid, rpoB, embB
# - AR fSNP in genes = gyrA
#
# **1972** +
# - pAR SNPs in genes = pncA, embB, embC, katG, promoter_fabG1-inhA, kasA, rpoB
# - AR fSNP in genes = gyrA, tlyA
#
# **2307** +
# - pAR SNPs in genes = gyrA, katG, embB, rpoB, pncA
# - AR fSNP in genes = promoter_fabG1-inhA, ethA
#
# **2511** +
# - pAR SNPs in genes = katG
# - AR fSNP in genes = rpoB
#
# **2523** +
# - pAR SNPs in genes = ethA, pncA, embB, inhA, kasA, rpoB
# - AR fSNP in genes = gyrA
#
# **2688** -
# - pAR SNPs in genes = pncA, embB, embC, inhA, gid, kasA, rpoB, promoter_fabG1-inhA
# - AR fSNP in genes = rpoC
#
# **2968** -
# - pAR SNPs in genes = embB, rpoB
# - AR fSNP in genes = ubiA
#
# **3451** +
# - pAR SNPs in genes = embB, pncA, rpoB, inhA
# - AR fSNP in genes = gyrA, pncA
#
# **3673** -
# - pAR SNPs in genes = pncA, embC, inhA, gid, kasA, rpoB, promoter_fabG1-inhA
# - AR fSNP in genes = embB
#
# **D** +
# - pAR SNPs in genes = gyrA, embB, rpsL, pncA, katG, rpoB
# - AR fSNP in genes = folC
#
# **I0000024-5** +
# - pAR SNPs in genes = pncA, embB, embA, embC, katG, kasA, rpoB
# - AR fSNP in genes = gyrB
#
# **KPS_5** +
# - pAR SNPs in genes = katG
# - AR fSNP in genes = embB
#
# **P000071** +
# - pAR SNPs in genes = rpoB
# - AR fSNP in genes = promoter_embA-embB, gid
#
# **P000155** +
# - pAR SNPs in genes = katG, rpoB, embB
# - AR fSNP in genes = gyrA
#
# **P000226** +
# - pAR SNPs in genes = gyrA, embB, rpoB
# - AR fSNP in genes = katG
#
# **P052** +
# - pAR SNPs in genes = promoter_fabG1-inhA
# - AR fSNP in genes = rpoC, rpoB
#
# **P10** +
# - pAR SNPs in genes = rpsL, embB, rpoB, promoter_fabG1-inhA
# - AR fSNP in genes = gyrA
#
# **P251** +
# - pAR SNPs in genes = promoter_fabG1-inhA
# - AR fSNP in genes = rpoB
# #### Of the $\frac{27}{178} \approx 15\%$ of subjects that developed an AR fSNP between sampling, $\frac{16}{27} \approx 59\%$ occurred among subjects that had at least 1pAR SNP and resulted in acquisition of an AR fSNP to an addtional antiobiotic (for which there were not AR mutations in the first isolate).
# ### Test for association between pre-existing MDR & AR SNP acquisition in *failure* subjects
# #### Split subjects into 2 groups, subjects without MDR in 1st isolate (Group C1) & subjects with MDR in 1st isolate (Group C2)
group_C1_subjects = set(patients_to_keep) - set(patients_with_MDR)
group_C2_subjects = patients_with_MDR
len(group_C1_subjects)
len(group_C2_subjects)
# #### Group C1
# Subjects without MDR in 1st collected longitudinal sample who also had no AR SNPs crop up between isolate collection
sum( AR_SNPs_between_isolates_per_subject[group_C1_subjects] == 0 )
# Subjects without MDR in 1st collected longitudinal sample who also had at least 1 AR SNP crop up between isolate collection
sum( AR_SNPs_between_isolates_per_subject[group_C1_subjects] > 0 )
float(13) / float(131)
# #### Group C2
# Subjects who had MDR in 1st collected longitudinal sample who also had no AR SNPs crop up between isolate collection
sum( AR_SNPs_between_isolates_per_subject[group_C2_subjects] == 0 )
# Subjects who had MDR in 1st collected longitudinal sample who also had at least 1 AR SNP crop up between isolate collection
sum( AR_SNPs_between_isolates_per_subject[group_C2_subjects] > 0 )
float(14) / float(47)
# #### Fisher exact test to test independence
AR_SNP_cont_table = np.array([[118,13] , [33,14]])
AR_SNP_cont_table
oddsratio, pvalue = stats.fisher_exact(AR_SNP_cont_table) #rows: (1) subjects without MDR in 1st isolate (2) subjects with MDR in 1st isolate ; columns: (1) no AR SNPs between isolate collection (2) at least 1 AR SNP between isolate collection
oddsratio
pvalue
| Longitudinal Isolates/(H) Antibiotic Resistance Allele Dynamics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import sys
sys.path.insert(0, '..')
import asyncio
import json
import collections
import math
# %matplotlib notebook
import numpy as nps
import matplotlib.pyplot as plt
from matplotlib.animation import FuncAnimation
from matplotlib.colors import LinearSegmentedColormap
plt.style.use('seaborn')
plt.rcParams['figure.figsize'] = [14, 8]
plt.rcParams['figure.dpi'] = 70 # 200 e.g. is really fine, but slower
from jupyterplot import ProgressPlot
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import pandas as pd
from pandas_connector import connector
# +
ip = "127.0.0.1"
port = 9090
connector.setup(ip=ip, port=port)
# -
task = connector.connect(200)
connector.dataframe.tail(5)
selection_fixation = float(connector.constants["selectionFixation"].replace(',', '.'))
selection_tolerance = float(connector.constants["selectionTolerance"].replace(',', '.'))
selection_target_acceleration = float(connector.constants["selectionTargetAcceleration"].replace(',', '.'))
# +
# Plotting
fig, ax = plt.subplots(2, 2, figsize=(12, 10))
line1, = ax[0][0].plot([], [], linestyle='-')
ax[0][0].axhline(selection_tolerance, color='blue', lw=2, alpha=0.25)
ax[0][0].axhline(-selection_tolerance, color='blue', lw=2, alpha=0.25)
line2, = ax[0][1].plot([], [], label="X Acceleration", linestyle='-')
ax[0][1].axhline(selection_target_acceleration - selection_tolerance, color='blue', lw=2, alpha=0.25)
ax[0][1].axhline(-selection_target_acceleration + selection_tolerance*2, color='blue', lw=2, alpha=0.25)
line3, = ax[1][0].plot([], [], label="Fixation", linestyle='-')
ax[1][0].axhline(selection_fixation, color='blue', lw=2, alpha=0.25)
line4, = ax[1][1].plot([], [], label="Selection", linestyle='-')
ax[0][0].set_title("EGHG Distance X Acceleration Means")
ax[0][1].set_title("EGHG Distance X Acceleration")
ax[1][0].set_title("Fixation")
ax[1][1].set_title("Selection")
def init():
line1.set_data([], [])
#line2.set_data([], [])
line3.set_data([], [])
#line4.set_data([], [])
ax[0][0].set_xlim(0, 200)
ax[0][0].set_ylim(-500, 500)
ax[0][1].set_xlim(0, 200)
ax[0][1].set_ylim(-2000, 2000)
ax[1][0].set_xlim(0, 200)
ax[1][0].set_ylim(-0.5, 1.5)
ax[1][1].set_xlim(0, 200)
ax[1][1].set_ylim(-0.5, 1.5)
ax[1][0].set_xlabel("Iteration (1/60s)")
ax[1][1].set_xlabel("Iteration (1/60s)")
def update(frame):
try:
line1.set_data(connector.dataframe.index, connector.dataframe["eyeGazeHeadGaze.distance.x_acceleration.means"])
line2.set_data(connector.dataframe.index, connector.dataframe["eyeGazeHeadGaze.cartesian.acceleration.x"])
line3.set_data(connector.dataframe.index, connector.dataframe["eyeGazeHeadGaze.fixation"])
line4.set_data(connector.dataframe.index, connector.dataframe["eyeGazeHeadGaze.selection"].astype(int))
return line1, line2, line3, line4,
except:
pass
animation = FuncAnimation(fig, update, init_func=init, interval=200, blit=True)
plt.show()
# -
connector.disconnect(task)
connector.dataframe["eyeGazeHeadGaze.selection"].str.astype(int)
| notebook/presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import unittest
import isolation
import game_agent
from importlib import reload
class IsolationTest(unittest.TestCase):
"""Unit tests for isolation agents"""
def setUp(self):
reload(game_agent)
self.player1 = "Player1"
self.player2 = "Player2"
self.game = isolation.Board(self.player1, self.player2)
def test_minimax(self):
# TODO: All methods must start with "test_"
return minimax(self.game,10)
if __name__ == '__main__':
unittest.main()
# +
"""Finish all TODO items in this file to complete the isolation project, then
test your agent's strength against a set of known agents using tournament.py
and include the results in your report.
"""
import random
class SearchTimeout(Exception):
"""Subclass base exception for code clarity. """
pass
def custom_score(game, player):
"""Calculate the heuristic value of a game state from the point of view
of the given player.
This should be the best heuristic function for your project submission.
Note: this function should be called from within a Player instance as
`self.score()` -- you should not need to call this function directly.
Parameters
----------
game : `isolation.Board`
An instance of `isolation.Board` encoding the current state of the
game (e.g., player locations and blocked cells).
player : object
A player instance in the current game (i.e., an object corresponding to
one of the player objects `game.__player_1__` or `game.__player_2__`.)
Returns
-------
float
The heuristic value of the current game state to the specified player.
"""
# TODO: finish this function!
raise NotImplementedError
def custom_score_2(game, player):
"""Calculate the heuristic value of a game state from the point of view
of the given player.
Note: this function should be called from within a Player instance as
`self.score()` -- you should not need to call this function directly.
Parameters
----------
game : `isolation.Board`
An instance of `isolation.Board` encoding the current state of the
game (e.g., player locations and blocked cells).
player : object
A player instance in the current game (i.e., an object corresponding to
one of the player objects `game.__player_1__` or `game.__player_2__`.)
Returns
-------
float
The heuristic value of the current game state to the specified player.
"""
# TODO: finish this function!
raise NotImplementedError
def custom_score_3(game, player):
"""Calculate the heuristic value of a game state from the point of view
of the given player.
Note: this function should be called from within a Player instance as
`self.score()` -- you should not need to call this function directly.
Parameters
----------
game : `isolation.Board`
An instance of `isolation.Board` encoding the current state of the
game (e.g., player locations and blocked cells).
player : object
A player instance in the current game (i.e., an object corresponding to
one of the player objects `game.__player_1__` or `game.__player_2__`.)
Returns
-------
float
The heuristic value of the current game state to the specified player.
"""
# TODO: finish this function!
raise NotImplementedError
class IsolationPlayer:
"""Base class for minimax and alphabeta agents -- this class is never
constructed or tested directly.
******************** DO NOT MODIFY THIS CLASS ********************
Parameters
----------
search_depth : int (optional)
A strictly positive integer (i.e., 1, 2, 3,...) for the number of
layers in the game tree to explore for fixed-depth search. (i.e., a
depth of one (1) would only explore the immediate sucessors of the
current state.)
score_fn : callable (optional)
A function to use for heuristic evaluation of game states.
timeout : float (optional)
Time remaining (in milliseconds) when search is aborted. Should be a
positive value large enough to allow the function to return before the
timer expires.
"""
def __init__(self, search_depth=3, score_fn=custom_score, timeout=10.):
self.search_depth = search_depth
self.score = score_fn
self.time_left = None
self.TIMER_THRESHOLD = timeout
class MinimaxPlayer(IsolationPlayer):
"""Game-playing agent that chooses a move using depth-limited minimax
search. You must finish and test this player to make sure it properly uses
minimax to return a good move before the search time limit expires.
"""
def get_move(self, game, time_left):
"""Search for the best move from the available legal moves and return a
result before the time limit expires.
************** YOU DO NOT NEED TO MODIFY THIS FUNCTION *************
For fixed-depth search, this function simply wraps the call to the
minimax method, but this method provides a common interface for all
Isolation agents, and you will replace it in the AlphaBetaPlayer with
iterative deepening search.
Parameters
----------
game : `isolation.Board`
An instance of `isolation.Board` encoding the current state of the
game (e.g., player locations and blocked cells).
time_left : callable
A function that returns the number of milliseconds left in the
current turn. Returning with any less than 0 ms remaining forfeits
the game.
Returns
-------
(int, int)
Board coordinates corresponding to a legal move; may return
(-1, -1) if there are no available legal moves.
"""
self.time_left = time_left
# Initialize the best move so that this function returns something
# in case the search fails due to timeout
best_move = (-1, -1)
try:
# The try/except block will automatically catch the exception
# raised when the timer is about to expire.
return self.minimax(game, self.search_depth)
except SearchTimeout:
pass # Handle any actions required after timeout as needed
# Return the best move from the last completed search iteration
return best_move
def terminal_test(game):
g = game.get_legal_moves()
if g == [(-1,-1)]:
return True
else:
return False
def minimax(self, game, depth):
"""Implement depth-limited minimax search algorithm as described in
the lectures.
This should be a modified version of MINIMAX-DECISION in the AIMA text.
https://github.com/aimacode/aima-pseudocode/blob/master/md/Minimax-Decision.md
**********************************************************************
You MAY add additional methods to this class, or define helper
functions to implement the required functionality.
**********************************************************************
Parameters
----------
game : isolation.Board
An instance of the Isolation game `Board` class representing the
current game state
depth : int
Depth is an integer representing the maximum number of plies to
search in the game tree before aborting
Returns
-------
(int, int)
The board coordinates of the best move found in the current search;
(-1, -1) if there are no legal moves
Notes
-----
(1) You MUST use the `self.score()` method for board evaluation
to pass the project tests; you cannot call any other evaluation
function directly.
(2) If you use any helper functions (e.g., as shown in the AIMA
pseudocode) then you must copy the timer check into the top of
each helper function or else your agent will timeout during
testing.
"""
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
# TODO: finish this function!
best_score = float("-inf")
best_move = None
for m in game.get_legal_moves():
# call has been updated with a depth limit
v = min_value(gameState.forecast_move(m), depth - 1)
if v > best_score:
best_score = v
best_move = m
return best_move
def min_value(game,depth):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if terminal_test(game):
return 1 # by Assumption 2
while depth <0:
v = float("inf")
for m in game.get_legal_moves():
v = min(v, max_value(gameState.forecast_move(m),depth-1))
depth -= 1
return v
else: return 0
def max_value(game,depth):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if terminal_test(game):
return -1 # by assumption 2
while depth <0:
v = float("-inf")
for m in game.get_legal_moves():
v = max(v, min_value(game.forecast_move(m),depth-1))
depth -= 1
return v
else: return 0
class AlphaBetaPlayer(IsolationPlayer):
"""Game-playing agent that chooses a move using iterative deepening minimax
search with alpha-beta pruning. You must finish and test this player to
make sure it returns a good move before the search time limit expires.
"""
def get_move(self, game, time_left):
"""Search for the best move from the available legal moves and return a
result before the time limit expires.
Modify the get_move() method from the MinimaxPlayer class to implement
iterative deepening search instead of fixed-depth search.
**********************************************************************
NOTE: If time_left() < 0 when this function returns, the agent will
forfeit the game due to timeout. You must return _before_ the
timer reaches 0.
**********************************************************************
Parameters
----------
game : `isolation.Board`
An instance of `isolation.Board` encoding the current state of the
game (e.g., player locations and blocked cells).
time_left : callable
A function that returns the number of milliseconds left in the
current turn. Returning with any less than 0 ms remaining forfeits
the game.
Returns
-------
(int, int)
Board coordinates corresponding to a legal move; may return
(-1, -1) if there are no available legal moves.
"""
self.time_left = time_left
# TODO: finish this function!
raise NotImplementedError
def alphabeta(self, game, depth, alpha=float("-inf"), beta=float("inf")):
"""Implement depth-limited minimax search with alpha-beta pruning as
described in the lectures.
This should be a modified version of ALPHA-BETA-SEARCH in the AIMA text
https://github.com/aimacode/aima-pseudocode/blob/master/md/Alpha-Beta-Search.md
**********************************************************************
You MAY add additional methods to this class, or define helper
functions to implement the required functionality.
**********************************************************************
Parameters
----------
game : isolation.Board
An instance of the Isolation game `Board` class representing the
current game state
depth : int
Depth is an integer representing the maximum number of plies to
search in the game tree before aborting
alpha : float
Alpha limits the lower bound of search on minimizing layers
beta : float
Beta limits the upper bound of search on maximizing layers
Returns
-------
(int, int)
The board coordinates of the best move found in the current search;
(-1, -1) if there are no legal moves
Notes
-----
(1) You MUST use the `self.score()` method for board evaluation
to pass the project tests; you cannot call any other evaluation
function directly.
(2) If you use any helper functions (e.g., as shown in the AIMA
pseudocode) then you must copy the timer check into the top of
each helper function or else your agent will timeout during
testing.
"""
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
# TODO: finish this function!
raise NotImplementedError
# -
def minimax(self, game, depth):
"""Implement depth-limited minimax search algorithm as described in
the lectures.
This should be a modified version of MINIMAX-DECISION in the AIMA text.
https://github.com/aimacode/aima-pseudocode/blob/master/md/Minimax-Decision.md
**********************************************************************
You MAY add additional methods to this class, or define helper
functions to implement the required functionality.
**********************************************************************
Parameters
----------
game : isolation.Board
An instance of the Isolation game `Board` class representing the
current game state
depth : int
Depth is an integer representing the maximum number of plies to
search in the game tree before aborting
Returns
-------
(int, int)
The board coordinates of the best move found in the current search;
(-1, -1) if there are no legal moves
Notes
-----
(1) You MUST use the `self.score()` method for board evaluation
to pass the project tests; you cannot call any other evaluation
function directly.
(2) If you use any helper functions (e.g., as shown in the AIMA
pseudocode) then you must copy the timer check into the top of
each helper function or else your agent will timeout during
testing.
"""
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
def terminal_test(self,game):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
return not bool(game.get_legal_moves())
def min_value(self,game,depth):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if self.terminal_test(game):
return 1 # by Assumption 2
while depth <0:
v = float("inf")
for m in game.get_legal_moves():
v = min(v, self.max_value(game.forecast_move(m),depth-1))
depth -= 1
return v
else: return self.score(game,self)
def max_value(self,game,depth):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if self.terminal_test(game):
return -1 # by assumption 2
while depth <0:
v = float("-inf")
for m in game.get_legal_moves():
v = max(v, self.min_value(game.forecast_move(m),depth-1))
depth -= 1
return v
else: return self.score(game,self)
# TODO: finish this function!
best_score = float("-inf")
best_move = None
for m in game.get_legal_moves():
v = self.min_value(game.forecast_move(m), depth - 1)
if v > best_score:
best_score = v
best_move = m
return best_move
def alphabeta(self, game, depth, alpha=float("-inf"), beta=float("inf")):
"""Implement depth-limited minimax search with alpha-beta pruning as
described in the lectures.
This should be a modified version of ALPHA-BETA-SEARCH in the AIMA text
https://github.com/aimacode/aima-pseudocode/blob/master/md/Alpha-Beta-Search.md
**********************************************************************
You MAY add additional methods to this class, or define helper
functions to implement the required functionality.
**********************************************************************
Parameters
----------
game : isolation.Board
An instance of the Isolation game `Board` class representing the
current game state
depth : int
Depth is an integer representing the maximum number of plies to
search in the game tree before aborting
alpha : float
Alpha limits the lower bound of search on minimizing layers
beta : float
Beta limits the upper bound of search on maximizing layers
Returns
-------
(int, int)
The board coordinates of the best move found in the current search;
(-1, -1) if there are no legal moves
Notes
-----
(1) You MUST use the `self.score()` method for board evaluation
to pass the project tests; you cannot call any other evaluation
function directly.
(2) If you use any helper functions (e.g., as shown in the AIMA
pseudocode) then you must copy the timer check into the top of
each helper function or else your agent will timeout during
testing.
"""
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
# TODO: finish this function!
def terminal_state(game):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
return not bool(game.get_legal_moves())
def min_value(game,depth,alpha,beta):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if terminal_state(game):
return 1
if depth ==0: return self.score(game,self)
v = float('inf')
for m in game.get_legal_moves():
v = min(v, max_value(game.forecast_move(m),depth-1,alpha,beta))
if v <= alpha:
return v
beta = min(beta,v)
return v
def max_value(game,depth,alpha,beta):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if terminal_state(game):
return -1
if depth ==0: return self.score(game,self)
v = float('-inf')
for m in game.get_legal_moves():
v = max(v, min_value(game.forecast_move(m),depth-1,alpha,beta))
if v >= beta:
return v
alpha = max(alpha,v)
return v
best_score = float('-inf')
best_move = None
for m in game.get_legal_moves():
v = min_value(game.forecast_move(m),depth-1,alpha,beta)
if v > best_score:
best_score = v
best_move = m
return best_move
self.time_left = time_left
best_move = (-1,-1)
self.iter_depth = 1
legal_moves = game.get_legal_moves()
while self.time_left() > self.TIMER_THRESHOLD:
if not legal_moves:
return (-1,-1)
best_move = self.alphabeta(game,self.iter_depth)
self.iter_depth += 1
return best_move
else:
raise SearchTimeout()
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
# TODO: finish this function!
def terminal_state(game):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
return not bool(game.get_legal_moves())
def min_value(game,depth,alpha,beta):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if terminal_state(game):
return 1
if depth ==0: return self.score(game,self)
v = float('inf')
for m in game.get_legal_moves():
v = min(v, max_value(game.forecast_move(m),depth-1,alpha,beta))
if v <= alpha:
return v
beta = min(beta,v)
return v
def max_value(game,depth,alpha,beta):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if terminal_state(game):
return -1
if depth ==0: return self.score(game,self)
v = float('-inf')
for m in game.get_legal_moves():
v = max(v, min_value(game.forecast_move(m),depth-1,alpha,beta))
if v >= beta:
return v
alpha = max(alpha,v)
return v
best_score = float('-inf')
best_move = None
for m in game.get_legal_moves():
v = min_value(game.forecast_move(m),depth-1,alpha,beta)
if v > best_score:
best_score = v
best_move = m
alpha = max(alpha,best_score)
return best_move
def get_move(self, game, time_left):
"""Search for the best move from the available legal moves and return a
result before the time limit expires.
Modify the get_move() method from the MinimaxPlayer class to implement
iterative deepening search instead of fixed-depth search.
**********************************************************************
NOTE: If time_left() < 0 when this function returns, the agent will
forfeit the game due to timeout. You must return _before_ the
timer reaches 0.
**********************************************************************
Parameters
----------
game : `isolation.Board`
An instance of `isolation.Board` encoding the current state of the
game (e.g., player locations and blocked cells).
time_left : callable
A function that returns the number of milliseconds left in the
current turn. Returning with any less than 0 ms remaining forfeits
the game.
Returns
-------
(int, int)
Board coordinates corresponding to a legal move; may return
(-1, -1) if there are no available legal moves.
"""
self.time_left = time_left
best_move = (-1,-1)
if game.get_legal_moves():
best_move = game.get_legal_moves()[0]
self.iter_depth = 1
legal_moves = game.get_legal_moves()
try:
while True:
best_move = self.alphabeta(game,self.iter_depth)
self.iter_depth += 1
except SearchTimeout:
pass
return best_move
def terminal_state(game):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
return not bool(game.get_legal_moves())
def min_value(game,depth,alpha,beta):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if self.terminal_state(game):
return 1
if depth ==0: return self.score(game,self)
v = float('inf')
for m in game.get_legal_moves():
v = min(v, self.max_value(game.forecast_move(m),depth-1,alpha,beta))
if v <= alpha:
return v
beta = min(beta,v)
return v
def max_value(game,depth,alpha,beta):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if self.terminal_state(game):
return -1
if depth ==0: return self.score(game,self)
v = float('-inf')
for m in game.get_legal_moves():
v = max(v, self.min_value(game.forecast_move(m),depth-1,alpha,beta))
if v >= beta:
return v
alpha = max(alpha,v)
return v
def alphabeta(self, game, depth, alpha=float("-inf"), beta=float("inf")):
"""Implement depth-limited minimax search with alpha-beta pruning as
described in the lectures.
This should be a modified version of ALPHA-BETA-SEARCH in the AIMA text
https://github.com/aimacode/aima-pseudocode/blob/master/md/Alpha-Beta-Search.md
**********************************************************************
You MAY add additional methods to this class, or define helper
functions to implement the required functionality.
**********************************************************************
Parameters
----------
game : isolation.Board
An instance of the Isolation game `Board` class representing the
current game state
depth : int
Depth is an integer representing the maximum number of plies to
search in the game tree before aborting
alpha : float
Alpha limits the lower bound of search on minimizing layers
beta : float
Beta limits the upper bound of search on maximizing layers
Returns
-------
(int, int)
The board coordinates of the best move found in the current search;
(-1, -1) if there are no legal moves
Notes
-----
(1) You MUST use the `self.score()` method for board evaluation
to pass the project tests; you cannot call any other evaluation
function directly.
(2) If you use any helper functions (e.g., as shown in the AIMA
pseudocode) then you must copy the timer check into the top of
each helper function or else your agent will timeout during
testing.
"""
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
# TODO: finish this function!
best_score = float('-inf')
best_move = None
for m in game.get_legal_moves():
v = self.min_value(game.forecast_move(m),depth-1,alpha,beta)
if v > best_score:
best_score = v
best_move = m
alpha = max(alpha,best_score)
return best_move
self.time_left = time_left
self.iter_depth = 1
if not game.get_legal_moves():
best_move = (-1,-1)
best_move = game.get_legal_moves()[0]
try:
while True:
best_move = self.alphabeta(game,self.iter_depth)
self.iter_depth += 1
except SearchTimeout:
pass
return best_move
self.time_left = time_left
self.iter_depth = 1
if not game.get_legal_moves():
best_move = (-1,-1)
best_move = game.get_legal_moves()[0]
try:
while True:
best_move = self.alphabeta(game,self.iter_depth)
self.iter_depth += 1
except SearchTimeout:
pass
return best_move
if game.is_loser(player):
return float('-inf')
if game.is_winner(player):
return float('inf')
own_moves = len(game.get_legal_moves(player))
opp_moves = len(game.get_legal_moves(game.get_opponent(player)))
return float(own_moves - 1.5*opp_moves)
if game.is_loser(player):
return float('-inf')
if game.is_winner(player):
return float('inf')
w, h = game.width/2., game.height/2.
own_moves = game.get_legal_moves(player)
opp_moves = game.get(game.get_opponent(player))
own_moves_count = len(own_moves)
oppn_moves_count = len(opp_moves)
oppn_weight, own_weight = weight,1
for m in own_moves:
if w == m[0] and h == m[1]:
own_weight *= center_weight
for m in opp_moves:
if w == m[0] and h == m[1]:
oppn_weight *= center_weight
return float(own_moves_count*own_weight - oppn_moves_count*oppn_weight)
own_moves = game.get_legal_moves(player)
opp_moves = game.get_legal_moves(game.get_opponent(player))
overlap = [m for m in own_moves if m in opp_moves]
score = 0
if player == game.active_player:
if len(overlap)>0:
score += 2
else:
if len(overlap) > 0:
score -= 2
return float(score)
if not game.get_legal_moves():
return (-1,-1)
best_score = float('-inf')
best_move = game.get_legal_moves()[0]
for m in game.get_legal_moves():
v = min_value(game.forecast_move(m),depth-1,alpha,beta)
if v > best_score:
best_score = v
best_move = m
alpha = max(alpha,best_score)
return best_move
oppn_moves_count = len(opp_moves)
score = 0
if player == game.active_player:
if own_moves_count > oppn_moves_count:
score += 4
elif own_moves_count < oppn_moves_count:
score -= 2
else:
if own_moves_count > oppn_moves_count:
score += 2
elif own_moves_count < oppn_moves_count:
score -= 4
return float(score)
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Micle5858/mit-deep-learning/blob/master/Reinforcement_Learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-ADWvu7NKN2r" colab_type="text"
# ##Reinforcement Learning
# The next and final topic in this course covers *Reinforcement Learning*. This technique is different than many of the other machine learning techniques we have seen earlier and has many applications in training agents (an AI) to interact with enviornments like games. Rather than feeding our machine learning model millions of examples we let our model come up with its own examples by exploring an enviornemt. The concept is simple. Humans learn by exploring and learning from mistakes and past experiences so let's have our computer do the same.
#
#
# + [markdown] id="HGCR3JWQLaQb" colab_type="text"
# ###Terminology
# Before we dive into explaining reinforcement learning we need to define a few key peices of terminology.
#
# **Enviornemt** In reinforcement learning tasks we have a notion of the enviornment. This is what our *agent* will explore. An example of an enviornment in the case of training an AI to play say a game of mario would be the level we are training the agent on.
#
# **Agent** an agent is an entity that is exploring the enviornment. Our agent will interact and take different actions within the enviornment. In our mario example the mario character within the game would be our agent.
#
# **State** always our agent will be in what we call a *state*. The state simply tells us about the status of the agent. The most common example of a state is the location of the agent within the enviornment. Moving locations would change the agents state.
#
# **Action** any interaction between the agent and enviornment would be considered an action. For example, moving to the left or jumping would be an action. An action may or may not change the current *state* of the agent. In fact, the act of doing nothing is an action as well! The action of say not pressing a key if we are using our mario example.
#
# **Reward** every action that our agent takes will result in a reward of some magnitude (positive or negative). The goal of our agent will be to maximize its reward in an enviornment. Sometimes the reward will be clear, for example if an agent performs an action which increases their score in the enviornment we could say they've recieved a positive reward. If the agent were to perform an action which results in them losing score or possibly dying in the enviornment then they would recieve a negative reward.
#
# The most important part of reinforcement learning is determing how to reward the agent. After all, the goal of the agent is to maximize its rewards. This means we should reward the agent appropiatly such that it reaches the desired goal.
#
#
# + [markdown] id="AoOJy9s4ZJJt" colab_type="text"
# ###Q-Learning
# Now that we have a vague idea of how reinforcement learning works it's time to talk about a specific technique in reinforcement learning called *Q-Learning*.
#
# Q-Learning is a simple yet quite powerful technique in machine learning that involves learning a matrix of action-reward values. This matrix is often reffered to as a Q-Table or Q-Matrix. The matrix is in shape (number of possible states, number of possible actions) where each value at matrix[n, m] represents the agents expected reward given they are in state n and take action m. The Q-learning algorithm defines the way we update the values in the matrix and decide what action to take at each state. The idea is that after a succesful training/learning of this Q-Table/matrix we can determine the action an agent should take in any state by looking at that states row in the matrix and taking the maximium value column as the action.
#
# **Consider this example.**
#
# Let's say A1-A4 are the possible actions and we have 3 states represented by each row (state 1 - state 3).
#
# | A1 | A2 | A3 | A4 |
# |:--: |:--: |:--: |:--: |
# | 0 | 0 | 10 | 5 |
# | 5 | 10 | 0 | 0 |
# | 10 | 5 | 0 | 0 |
#
# If that was our Q-Table/matrix then the following would be the preffered actions in each state.
#
# > State 1: A3
#
# > State 2: A2
#
# > State 3: A1
#
# We can see that this is because the values in each of those columns are the highest for those states!
#
#
# + [markdown] id="u5uLpN1yemTx" colab_type="text"
# ###Learning the Q-Table
# So that's simple, right? Now how do we create this table and find those values. Well this is where we will dicuss how the Q-Learning algorithm updates the values in our Q-Table.
#
# I'll start by noting that our Q-Table starts of with all 0 values. This is because the agent has yet to learn anything about the enviornment.
#
# Our agent learns by exploring the enviornment and observing the outcome/reward from each action it takes in each state. But how does it know what action to take in each state? There are two ways that our agent can decide on which action to take.
# 1. Randomly picking a valid action
# 2. Using the current Q-Table to find the best action.
#
# Near the beginning of our agents learning it will mostly take random actions in order to explore the enviornment and enter many different states. As it starts to explore more of the enviornment it will start to gradually rely more on it's learned values (Q-Table) to take actions. This means that as our agent explores more of the enviornment it will develop a better understanding and start to take "correct" or better actions more often. It's important that the agent has a good balance of taking random actions and using learned values to ensure it does get trapped in a local maximum.
#
# After each new action our agent wil record the new state (if any) that it has entered and the reward that it recieved from taking that action. These values will be used to update the Q-Table. The agent will stop taking new actions only once a certain time limit is reached or it has acheived the goal or reached the end of the enviornment.
#
# ####Updating Q-Values
# The formula for updating the Q-Table after each action is as follows:
# > $ Q[state, action] = Q[state, action] + \alpha * (reward + \gamma * max(Q[newState, :]) - Q[state, action]) $
#
# - $\alpha$ stands for the **Learning Rate**
#
# - $\gamma$ stands for the **Discount Factor**
#
# ####Learning Rate $\alpha$
# The learning rate $\alpha$ is a numeric constant that defines how much change is permitted on each QTable update. A high learning rate means that each update will introduce a large change to the current state-action value. A small learning rate means that each update has a more subtle change. Modifying the learning rate will change how the agent explores the enviornment and how quickly it determines the final values in the QTable.
#
# ####Discount Factor $\gamma$
# Discount factor also know as gamma ($\gamma$) is used to balance how much focus is put on the current and future reward. A high discount factor means that future rewards will be considered more heavily.
#
# <br/>
# <p>To perform updates on this table we will let the agent explpore the enviornment for a certain period of time and use each of its actions to make an update. Slowly we should start to notice the agent learning and choosing better actions. </p>
#
#
# + [markdown] id="rwIl0sJgmu4D" colab_type="text"
# ##Q-Learning Example
# For this example we will use the Q-Learning algorithm to train an agent to navigate a popular enviornment from the [Open AI Gym](https://gym.openai.com/). The Open AI Gym was developed so programmers could practice machine learning using unique enviornments. Intersting fact, <NAME> is one of the founders of OpenAI!
#
# Let's start by looking at what Open AI Gym is.
# + id="rSETF0zqokYr" colab_type="code" colab={}
import gym # all you have to do to import and use open ai gym!
# + [markdown] id="8cH3AmCzotO1" colab_type="text"
# Once you import gym you can load an enviornment using the line ```gym.make("enviornment")```.
# + id="UKN1ScBco3dp" colab_type="code" colab={}
env = gym.make('FrozenLake-v0') # we are going to use the FrozenLake enviornment
# + [markdown] id="3SvSlmVwo8cY" colab_type="text"
# There are a few other commands that can be used to interact and get information about the enviornment.
# + id="FF3icIeapFct" colab_type="code" colab={}
print(env.observation_space.n) # get number of states
print(env.action_space.n) # get number of actions
# + id="lc9cwp03pQVn" colab_type="code" colab={}
env.reset() # reset enviornment to default state
# + id="sngyjPDapUt7" colab_type="code" colab={}
action = env.action_space.sample() # get a random action
# + id="HeEfi8xypXya" colab_type="code" colab={}
new_state, reward, done, info = env.step(action) # take action, notice it returns information about the action
# + id="_1W3D81ipdaS" colab_type="code" colab={}
env.render() # render the GUI for the enviornment
# + [markdown] id="vmW6HAbQp01f" colab_type="text"
# ###Frozen Lake Enviornment
# Now that we have a basic understanding of how the gym enviornment works it's time to discuss the specific problem we will be solving.
#
# The enviornment we loaded above ```FrozenLake-v0``` is one of the simplest enviornments in Open AI Gym. The goal of the agent is to navigate a frozen lake and find the Goal without falling through the ice (render the enviornment above to see an example).
#
# There are:
# - 16 states (one for each square)
# - 4 possible actions (LEFT, RIGHT, DOWN, UP)
# - 4 different types of blocks (F: frozen, H: hole, S: start, G: goal)
#
#
# + [markdown] id="YlWoK75ZrK2b" colab_type="text"
# ###Building the Q-Table
# The first thing we need to do is build an empty Q-Table that we can use to store and update our values.
# + id="r767K4s0rR2p" colab_type="code" colab={}
import gym
import numpy as np
import time
env = gym.make('FrozenLake-v0')
STATES = env.observation_space.n
ACTIONS = env.action_space.n
# + id="UAzMWGatrVIk" colab_type="code" colab={}
Q = np.zeros((STATES, ACTIONS)) # create a matrix with all 0 values
Q
# + [markdown] id="vc_h8tLSrpmc" colab_type="text"
# ###Constants
# As we discussed we need to define some constants that will be used to update our Q-Table and tell our agent when to stop training.
# + id="-FQapdnnr6P1" colab_type="code" colab={}
EPISODES = 2000 # how many times to run the enviornment from the beginning
MAX_STEPS = 100 # max number of steps allowed for each run of enviornment
LEARNING_RATE = 0.81 # learning rate
GAMMA = 0.96
# + [markdown] id="NxrAj91rsMfm" colab_type="text"
# ###Picking an Action
# Remember that we can pick an action using one of two methods:
# 1. Randomly picking a valid action
# 2. Using the current Q-Table to find the best action.
#
# Here we will define a new value $\epsilon$ that will tell us the probabillity of selecting a random action. This value will start off very high and slowly decrease as the agent learns more about the enviornment.
# + id="YUAQVyX0sWDb" colab_type="code" colab={}
epsilon = 0.9 # start with a 90% chance of picking a random action
# code to pick action
if np.random.uniform(0, 1) < epsilon: # we will check if a randomly selected value is less than epsilon.
action = env.action_space.sample() # take random action
else:
action = np.argmax(Q[state, :]) # use Q table to pick best action based on current values
# + [markdown] id="5n-i0B7Atige" colab_type="text"
# ###Updating Q Values
# The code below implements the formula discussed above.
# + id="9r7R1W6Qtnh8" colab_type="code" colab={}
Q[state, action] = Q[state, action] + LEARNING_RATE * (reward + GAMMA * np.max(Q[new_state, :]) - Q[state, action])
# + [markdown] id="__afaD62uh8G" colab_type="text"
# ###Putting it Together
# Now that we know how to do some basic things we can combine these together to create our Q-Learning algorithm,
# + id="AGiYCiNuutHz" colab_type="code" colab={}
import gym
import numpy as np
import time
env = gym.make('FrozenLake-v0')
STATES = env.observation_space.n
ACTIONS = env.action_space.n
Q = np.zeros((STATES, ACTIONS))
EPISODES = 1500 # how many times to run the enviornment from the beginning
MAX_STEPS = 100 # max number of steps allowed for each run of enviornment
LEARNING_RATE = 0.81 # learning rate
GAMMA = 0.96
RENDER = False # if you want to see training set to true
epsilon = 0.9
# + id="jFRtn5dUu5ZI" colab_type="code" colab={}
rewards = []
for episode in range(EPISODES):
state = env.reset()
for _ in range(MAX_STEPS):
if RENDER:
env.render()
if np.random.uniform(0, 1) < epsilon:
action = env.action_space.sample()
else:
action = np.argmax(Q[state, :])
next_state, reward, done, _ = env.step(action)
Q[state, action] = Q[state, action] + LEARNING_RATE * (reward + GAMMA * np.max(Q[next_state, :]) - Q[state, action])
state = next_state
if done:
rewards.append(reward)
epsilon -= 0.001
break # reached goal
print(Q)
print(f"Average reward: {sum(rewards)/len(rewards)}:")
# and now we can see our Q values!
# + id="Zo-tNznd65US" colab_type="code" colab={}
# we can plot the training progress and see how the agent improved
import matplotlib.pyplot as plt
def get_average(values):
return sum(values)/len(values)
avg_rewards = []
for i in range(0, len(rewards), 100):
avg_rewards.append(get_average(rewards[i:i+100]))
plt.plot(avg_rewards)
plt.ylabel('average reward')
plt.xlabel('episodes (100\'s)')
plt.show()
# + [markdown] id="gy4YH2m9s1ww" colab_type="text"
# ##Sources
# 1. <NAME>. “Simple Reinforcement Learning: Q-Learning.” Medium, Towards Data Science, 1 July 2019, https://towardsdatascience.com/simple-reinforcement-learning-q-learning-fcddc4b6fe56.
#
# 2. Openai. “Openai/Gym.” GitHub, https://github.com/openai/gym/wiki/FrozenLake-v0.
| Reinforcement_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object-Oriented Programming 2
# In the previous workbook we touched on the basics of OOP in python. This should be enough to get you going with programming in an object-oriented manner, however objects in python can be MUCH more powerful than demonstrated there. Using various bits of magic that are either general properties of OOP, or specific to the python implementation, we can do a lot!
# ## "Dunder" methods
#
# ### Making our classes more usable
# Let's start with the chair class we created in the last session:
class chair:
def __init__(self, legs, height, colour):
self.legs = legs
self.height = height
self.colour = colour
def paint(self, newcolour):
self.colour = newcolour
my_chair = chair(6, 0.8, "black")
# Now usually for an object, if we want to know what is in it, we can print it, like so:
testlist = [1,2,3]
print(testlist)
# Let's see what happens if we print our chair instance.
print(my_chair)
# Obviously this is not very useful. It tells us that the object is of the class `chair` and the memory address at which it decides. Wouldn't it be nice if instead it told us about the chair itself? We can do that by implementing another special method called `__str__()`.
# +
class chair:
def __init__(self, legs, height, colour):
self.legs = legs
self.height = height
self.colour = colour
def __str__(self):
return "A {} legged, {}m tall, {} chair.".format(self.legs, self.height, self.colour)
def paint(self, newcolour):
self.colour = newcolour
my_chair = chair(6, 0.8, "black")
# -
# Now let's see what happens if we print this new chair:
print(my_chair)
# Cool! Now whenever we want to get an idea of what chairs we have, we can just print them!
#
# Note that the `__str__()` method *must* return a string, though it can do other things in the function before it returns that.
#
# These magical methods that do strange things are known as **dunder** methods, as they start and end with a double underscore. There are many of these [(see here for more)](https://docs.python.org/3/reference/datamodel.html#special-method-names) and they do a wide variety of different things.
# ### `__repr__()`
# Sometimes it can be nice to have a way to get the code which can be used to generate an object. This is where the `__repr__()` dunder method comes in. It allows you to create a representation of an object in its current state. Let's implement `__repr__()` for our chair class
class chair:
def __init__(self, legs, height, colour):
self.legs = legs
self.height = height
self.colour = colour
def __repr__(self):
return 'chair({}, {}, "{}")'.format(self.legs, self.height, self.colour)
def __str__(self):
return "A {} legged, {}m tall, {} chair.".format(self.legs, self.height, self.colour)
def paint(self, newcolour):
self.colour = newcolour
# Now let's reinstantiate `my_chair` with the new class definition
my_chair = chair(6, 0.8, "black")
# We can use the `repr()` function to check the representation of this object:
repr(my_chair)
# As you can see it returns the code that we wrote to generate the object in the first place!
#
# Something else important to note is that if you have not implemented the `__str__()` dunder method, when you call `print()` on an object it will fall back on `__repr__()` if that exists, and then only as a last resort will it print the unhelpful `<__main__.chair object at 0x7f70903cc550>` that we got earlier.
#
# It is generally good practice if you're going to be using a class for any serious work, to implement these methods, especially `__repr__()`.
# ### Making an iterable
# One major use of classes is as a container for data. You can think of these as structures that hold data and metadata in a standard form, kind of like a list or dictionary, but with **MORE FUNCTIONALITY**.
# Let's make a simple class to hold a stack of chairs:
class chairstack:
def __init__(self, chairs, location="lab"):
self.chairs = chairs
self.location = location
def __repr__(self):
return "chairstack({})".format(self.chairs)
def __str__(self):
return "Chair stack containing {} chair/s.".format(len(self.chairs))
# +
import random
chair_stack = []
for x in range(10):
# Make 10 green chairs with random numbers of legs and random heights
chair_stack.append(chair(random.randint(1, 10), random.random()*2, "green"))
stack = chairstack(chair_stack)
# -
print(stack)
repr(stack)
# Now, what if we wanted to get height of the second chair in the stack? We can retrieve it as a property of the stack as follows...
print(stack.chairs[1].height)
# However that feels a bit ungainly. When we are talking about the stack, really we want to be able to just say `stack[1].height` as when we're indexing the stack we know we just want to be getting an individual chair, i.e. when we say `stack[1]` we will always want the entry at `stack.chairs[1]`.
#
# Luckily there are extra methods that we can write which will allow this behaviour.
# +
class chairstack:
def __init__(self, chairs, location="lab"):
self.chairs = chairs
self.location = location
def __repr__(self):
return "chairstack({})".format(self.chairs)
def __str__(self):
return "Chair stack containing {} chair/s.".format(len(self.chairs))
def __len__(self):
return len(self.chairs)
def __getitem__(self, chair_num):
return self.chairs[chair_num]
stack = chairstack(chair_stack)
# -
for x in stack:
print(x)
print(stack[1])
# Here, the `__len__()` method is called when you request `len(object)`, something you could either do just from the terminal, or this is also used internally when defining the parameters of an iteration.
#
# The `__getitem__()` method allows you to return an entry at a selected offset.
#
# There are also another two functions \[`__iter__()` and `__next__()`\] that allow more efficient iteration of these sorts of classes, however they don't improve usability as such, just make things quicker.
# ### There's so much more
# Dunder methods are widely varied, and defining them allows us to customise nearly every aspect of how a class operates in the wider context of a python program. It is definitely worthwhile at least becoming familiar with the fact that they exist, because you can end up making your life a whole chunk easier.
# ## Subclassing
# Another major feature of classes in python is the idea of subclassing.
#
# A chair (as we have been talking about the whole time so far) is a type of seat. There are also other types of seat such as stools and beanbags.
#
# Wouldn't it be nice to have a different class for each of these types of seat? However it's also a lot of work to implement 3 classes with almost exactly the same code inside.
#
# We can solve this using the joint ideas of subclassing and inheritance. A subclass *inherits* the methods and attributes of the parent class (also called the superclass).
# +
class Seat:
def __init__(self, legs, height, colour):
self.legs = legs
self.height = height
self.colour = colour
def __repr__(self):
return 'Seat({}, {}, "{}")'.format(self.legs, self.height, self.colour)
def __str__(self):
return "A {} legged, {}m tall, {} seat.".format(self.legs, self.height, self.colour)
def paint(self, newcolour):
self.colour = newcolour
class Stool(Seat):
pass
# -
my_stool = Stool(3, 0.9, "Black")
print(my_stool)
# Now you'll notice that the `Stool` class here acts like a `Seat` in all ways. However let's implement a beanbag.
# +
class Beanbag(Seat):
def __init__(self, legs, height, colour, filling):
self.filling = filling
super().__init__(legs, height, colour)
my_beanbag = Beanbag(0, 0.4, "Blue", "Polyester")
print(my_beanbag)
# -
# Now notice here we added an extra attribute that only applies to Beanbags. Stools are not usually filled with anything for instance.
#
# A subclass has all the attributes and methods of its superclass, and of itself. If there is a conflict in name between the two, the methods/properties of the subclass take priority. This is called method overloading or overriding.
#
# *Note: There is actually a subtle difference between the two. Overloading is replacing the method with another one that has the same name, but different arguments. Overriding uses the same name and same arguments.*
#
# The Beanbag here is also unpaintable, as it is a flexible fabric and doesn't take paint very well. Thus we can override this method to raise an error as follows.
class Beanbag(Seat):
def __init__(self, legs, height, colour, filling):
self.filling = filling
super().__init__(legs, height, colour)
def paint(self, newcolour):
raise TypeError("Cannot paint a Beanbag.")
# +
my_beanbag = Beanbag(0, 0.4, "Blue", "Polyester")
my_beanbag.paint("Red")
# -
# ## Custom exceptions
# We use exceptions a lot when coding in python. They tell us when we screwed something up, and we can use them to control the path of execution.
#
# The exceptions defined in Python as standard are very useful, but they don't necessarily cover exactly what you need. Sometimes you might want an exception that only handles certain elements. This can be done using the wonderful power of **subclassing**!
#
# Lets take a trivial function that makes animals say things.
# +
def say_woof(subject):
subject = subject.lower()
if subject == "dog":
print("Woof")
else:
raise AttributeError("Only dogs can go 'woof', {}s cannot go 'woof'".format(subject))
say_woof("cat")
# -
# This did the job very well, however it is quite broad. Anything that triggers an attribute error looks the same here, so if you try to set the subject to `1` then we will also get an attribute error.
say_woof(1)
# We may want to catch when the error is expected, but fail when it is unexpected, and at present we cannot differentiate between one of our own raised errors and an error that is triggered from some other operation failing. This is where a custom exception comes in. Let's define a new one:
# +
class WoofError(Exception): # Hey look, a subclass!!
pass
def say_woof(subject):
subject = subject.lower()
if subject == "dog":
print("Woof")
else:
raise WoofError("Only dogs can go 'woof', {0}s cannot go 'woof', they are {0}s...".format(subject))
say_woof("chair")
# -
say_woof(1)
# Now we can catch `WoofError` exceptions without accidentally catching any exceptions that are not WoofErrors.
#
# Your custom exceptions can do whatever you want them to do, and as such are very useful when writing larger scripts and packages.
| content/notebooks/test/OOP2.ipynb |
# ---
# layout: post
# title: "1st 함께하는 딥러닝 컨퍼런스"
# author: 김태영
# date: 2018-06-28 10:00:00
# categories: seminar
# comments: true
# image: http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Conference_All_Together_DLCAT_title1.png
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# 이제 딥러닝 기술은 무서운 속도로 각 분야에 퍼져가고 있습니다. 그에 따라 활용 사례도 늘어나고 있고, 실전 적용에 막히는 여러가지 문제도 해결하고자 많은 연구가 활발히 이루어지고 있습니다. 딥러닝을 공부하시는 분이라면 딥러닝 코어 및 알고리즘은 물론 타 분야의 활용사례, 최근에 유행하고 있는 GAN과 강화학습까지 관심을 가지고 계실겁니다. 다양한 주제로 즐겁게 소통하고자 첫번째 '함께하는 딥러닝 컨퍼런스'를 대전에서 개최합니다. 대전은 정부출연연구원 및 정부청사, 우수한 대학교, 최첨단 기술 중심의 벤처회사들이 밀집된 지역인 만큼 다른 지역과는 또 다른 느낌의 소통이 이뤄질 것 같아 기대되네요~
#
# 
#
# 1. 일시 : 2018년 6월 28일 (10시~18시)
# 2. 장소 : (우)34113 대전광역시 유성구 가정로 217 과학기술연합대학원대학교
# * 세션 A: 강당 300명
# * 세션 B: 사이언스홀 60명
# * 세션 C: 대회의실 30명
#
# 3. 주최 : (주)인스페이스, 케라스 코리아, 대딥사, 한국원자력연구원
# 4. 후원 : UST 과학기술연합대학원대학교, NIPA 정보통신산업진흥원
# 5. 대상 : 인공지능 및 딥러닝에 관심있거나 관련 연구를 수행하시는 분들 (약 500명)
#
# 
# ---
#
# 발표는 아래 세가지 방식으로 진행됩니다.
# * 세미나(50분) : 어떤 주제에 대해 깊이 있는 지식이나 전반적인 내용을 다룹니다.
# * 라이트닝톡(20분) : 자신의 연구 및 기관에 대한 소개, 자유로운 토론 위주로 합니다.
# * 핸즈온랩 : 직접 따라하면서 실습을 할 수 있는 시간을 가집니다.
# ### 딥러닝 몰라도 간(GAN)부터 보기, 김태영 - 입문 (세미나)
# ---
# 
#
# #### 연사소개
# (주)인스페이스 기술이사, 케라스 코리아, 대딥사 운영자이며, 경희대학교 우주탐사학과 박사과정에 진학 중입니다. 다양한 분야에 딥러닝 모델을 심을 수 있도록 대중화에 힘쓰고 있습니다.
#
# #### 발표소개
# 요즘은 인공지능이 그림도 그리고, 진짜같은 사람 목소리도 만들어내는 세상입니다. 이러한 것들을 생성모델이라고 하고, 그 근간에는 GAN이라는 것이 있습니다. 딥러닝을 잘 모르시는 분들이라도 레고 사람을 조립해보면서 기본적은 GAN의 개념을 익혀보겠습니다. CGAN, LSGAN, InfoGAN뿐만아니라 Pix2Pix, CycleGAN 등도 레고 사람으로 조립해보면서 확실히 간(GAN) 보도록 하겠습니다.
#
# 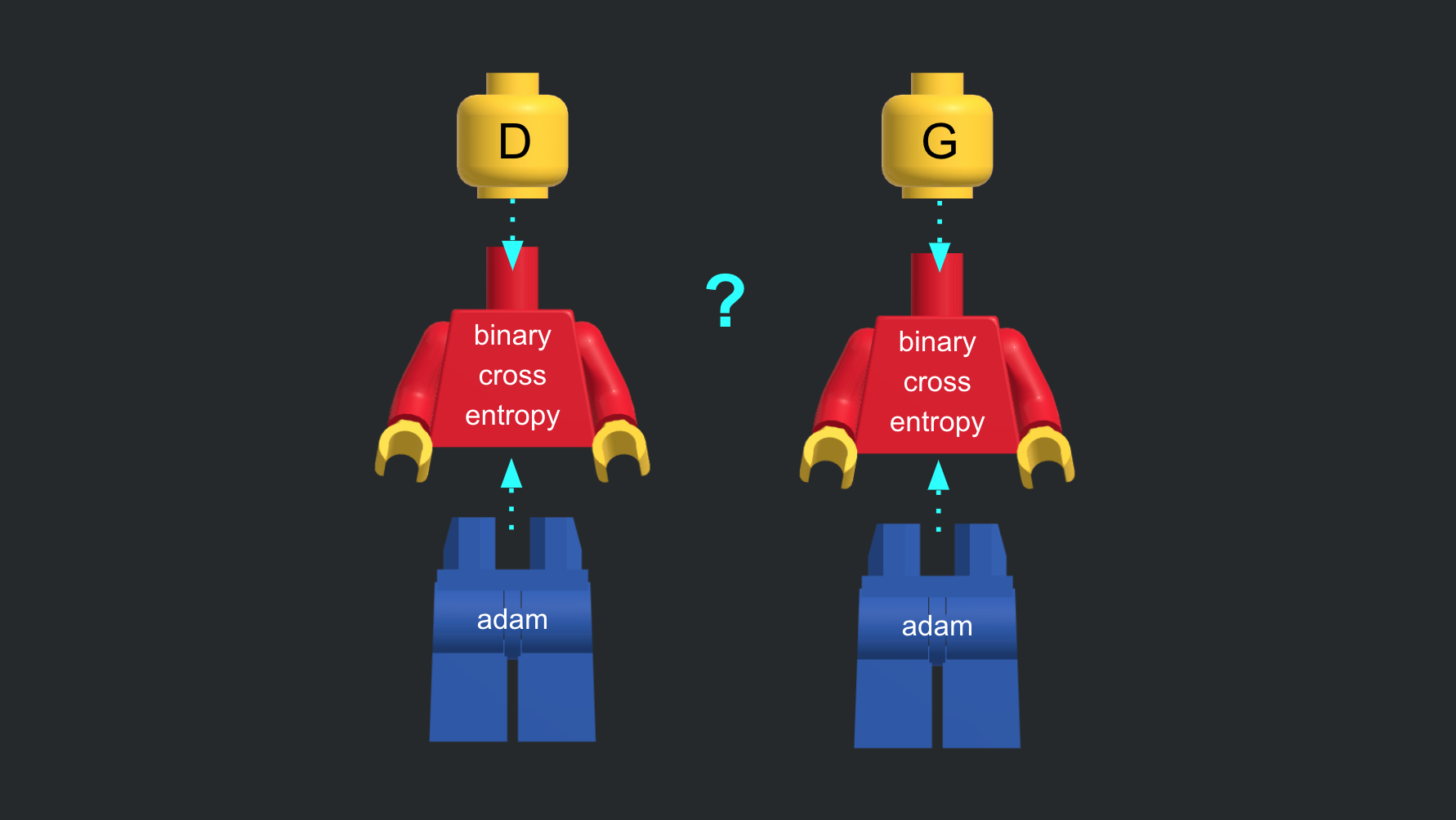
#
# 딥러닝 모델은 네트워크, 목표함수(objective function), 최적화기(optimizer)으로 구성되고, 레고 사람으로 비유하면 머리, 상반신, 하반신으로 나눌 수 있습니다. GAN에 관심을 가지신 분들은 Generator (생성자)와 Discriminator(판별자)가 있다는 것을 아실겁니다. 그럼 GAN에서는 두 개의 모델이 있는 데, 이걸 어떻게 조립해야할까요? 그림에 있는 화살표처럼 조립하면 될까요? 그렇지 않습니다. 조금은 잔인한 방법으로 조립됩니다. 좀 더 확장된 GAN들은 좀 더 잔인합니다. 노약자나 심신이 약하신 분들은 주의하시길 바랍니다.
#
# #### 참고자료
# * 블로그 : [안녕? 레고사람, 안녕? 딥러닝 모델](https://tykimos.github.io/2018/05/04/Making_Keras_Model_using_Lego_Minifigures/)
#
# #### 발표자료
# * 구글 프리젠테이션 : [모두의 손에 딥러닝 툴을...](https://docs.google.com/presentation/d/1dCyZmxGQgICmUp4t_ora4K3q2J52oxPw5CWFIrC0J-k/edit)
#
# #### 블로그 및 깃헙
#
# * [김태영의 케라스 블로그](https://tykimos.github.io) : 케라스와 딥러닝에 관련된 강좌 및 세미나 자료가 정리되어 있습니다. 딥러닝 입문에 추천드립니다.
#
# #### 저서
#
# * 블록과 함께하는 파이썬 딥러닝 케라스 : 케라스 사용에 필요한 딥러닝 개념 설명과 실습 가능한 파이썬 소스코드를 담고 있으며 직관적으로 딥러닝 모델 구성을 이해할 수 있도록 블록쌓기 예제가 포함되어 있습니다. [교보문고 바로가기](http://www.kyobobook.co.kr/product/detailViewKor.laf?ejkGb=KOR&mallGb=KOR&barcode=9788960882133&orderClick=LAG&Kc=#N&OV_REFFER=http://click.linkprice.com/click.php?m=kbbook&a=A100532541&l=9999&l_cd1=0&u_id=jhqg2qh8q101t9ww02yqe&l_cd2=0&tu=http%3A%2F%2Fwww.kyobobook.co.kr%2Fproduct%2FdetailViewKor.laf%3FejkGb%3DKOR%26mallGb%3DKOR%26barcode%3D9788960882133%26orderClick%3DLAG%26Kc%3D%23N)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/tykimo](https://www.facebook.com/tykimo)
#
# 
# ### 딥러닝 보물상자 '캐글' 알아보기 , 이유한 - 입문 (세미나)
# ---
# 
#
# #### 연사소개
# 현재 KAIST 생명화학공학과에서 박사과정 재학중이며, 전공은 분자 시뮬레이션입니다. 데이터 사이언스, AI를 어디서 배울지 고민하다가 캐글을 알게 된 후, 캐글에 푹 빠져버린 대학원생입니다. 저에게 큰 도움을 준 캐글을 다른 사람들과 함께 하고 싶어 발표자로 참여하게 되었습니다. 현재 [캐글 코리아](https://www.facebook.com/groups/230717130993727/)을 운영 중에 있습니다.
#
# #### 발표소개
# 전세계 데이터를 사랑하는 사람들에게 데이터를 제공하고, 함께 즐길 수 있도록 해주는 캐글을 소개하려고 합니다. 캐글 내 유능한 데이터분석가들의 귀하디 귀한 노하우를 거저!!! 얻을 수 있는 방법을 소개해드릴 겁니다. 또한 자신의 노하우도 거저!!! 나눠주는 방법도 소개해드릴 겁니다. 제 세션에 참석하시는 모든 분들이 캐글러가 되는 게 제 발표 목표입니다!
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_lyh_file.pdf)
#
# #### 블로그 및 깃헙
# * 블로그 : 캐글코리아 - [http://kaggle-kr.tistory.com/](http://kaggle-kr.tistory.com/)
# * 유투브 : [https://www.youtube.com/channel/UC--LgKcZVgffjsxudoXg5pQ/?reload=9](https://www.youtube.com/channel/UC--LgKcZVgffjsxudoXg5pQ/?reload=9)
#
# #### 연락처
# * 이메일 : <EMAIL> 또는 <EMAIL>
# * 페이스북 : [https://www.facebook.com/youhan.lee.33](https://www.facebook.com/youhan.lee.33)
#
# 
# ### 타이타닉 생존자 예측을 해보자, 이유한 - 입문 (핸즈온랩)
# ---
#
# #### 핸즈온랩소개
# 핸즈온랩에서는 타이타닉 생존자 예측하는 문제를 머신러닝으로 풀어보는 시간을 가질 예정입니다. 실무에 인공지능을 접목하실려는 분들께서는 생생한 꿀팁들을 가지고 가질 수가 있습니다. 실제 가입 및 실습을 진행합니다.
#
# #### 준비물
# 1. 노트북
# ### Graph Neural Network, 류성옥 - 알고리즘 (세미나)
# ---
# 
#
# #### 연사소개
# 석박사통합과정 5년차이며 KAIST 화학과 전공입니다. 양자역학을 연구하다가, AI가 세상을 바꿀 기술이라 생각되어 딥러닝을 공부하는 대학원생입니다. AI를 활용하여 Smart Drug Discovery를 연구중이며, Graph Neural Network 을 이용한 네트워크 분석에 관심이 많습니다.
#
# #### 발표소개
# 격화된 공간에서 표현되는 이미지와 다르게, 네트워크/물리/화학 등 시스템을 구성하는 객체들띠리 복잡한 관계를 형성하는 문제들은 Graph로 문제가 정의되곤 합니다. 이러한 문제를 data-driven으로 해결하기 위한 Graph Neural Network에 대해서 소개하고, 실제로 분자들의 구조-물성 관계(Quantitative Structure Activity Relationship; QSAR)을 딥러닝 모델을 개발하여 연구한 사례를 소개합니다.
# 뱀다리) 당일 4주훈련을 마치고 퇴소하여 까까머리로 발표할 예정입니다^.
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_rsy_file.pdf)
#
# #### 참고자료
# * 논문 : [Deeply learning molecular structure-property relationships using graph attention neural network](https://arxiv.org/abs/1805.10988)
#
# #### 블로그 및 깃헙
# * 블로그 : [http://suma_maple.blog.me](http://suma_maple.blog.me)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/ryu.seongok.1](https://www.facebook.com/ryu.seongok.1)
#
# 
# ### Deep convolutional framelets: application to diffuse optical tomography, 유재준 - 알고리즘 (세미나)
# ---
# 
#
# #### 연사소개
# 올해 2월 KAIST 바이오및뇌공학과 박사 졸업 후 네이버 클로바에서 일하고 있습니다. GANs, VAE, Autoregressive model 등 generative model과 interpretable model에 대한 연구에 관심이 있습니다.
#
# #### 발표소개
# Super-resolution, image denoising, reconstruction, segmentation 등의 다양한 inverse problem에서 deep learning이 매우 좋은 결과를 보여주고 있습니다. 이미 기존 방식들이 어느 정도 잘 하고 있던 분야에서 성능을 더 잘 내고 있는 것은 분명한데, 그렇다면 더 복잡한 물리적 현상도 deep learning을 이용하여 학습하는 것이 가능할까요? 이번 발표에서는 Inverse scattering problem이라는 좀 더 어려운 문제를 deep learning으로 풀어보고자 노력한 제 박사학위 연구(밑바닥부터 졸업까지 삽질기)를 소개하고자 합니다. 덧붙여서 시간이 허락한다면, deep learning에서 특정 구조가 잘 동작하는 이유를 분석하고, 어떤 식으로 구조를 디자인해야 할 지에 대해 신호처리 관점에서 풀어낸 deep convolutional framelets 이론을 같이 소개하겠습니다.
#
# #### 발표자료
# * PDF : [Deep convolutional framelets](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_yjj_file.pdf)
#
# #### 참고자료
# * [Deep Learning Can Reverse Photon Migration for Diffuse Optical Tomography](https://arxiv.org/abs/1712.00912)
# * [A Mathematical Framework for Deep Learning in Elastic Source Imaging](https://arxiv.org/abs/1802.10055)
# * [Deep Convolutional Framelets: A General Deep Learning Framework for Inverse Problems](https://arxiv.org/abs/1707.00372)
#
# #### 블로그 및 깃헙
# * 블로그 : [http://jaejunyoo.blogspot.com](http://jaejunyoo.blogspot.com)
# * 깃헙 : [https://github.com/jaejun-yoo/](https://github.com/jaejun-yoo/)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/thinking.factory](https://www.facebook.com/thinking.factory)
#
# 
# ### TensorFlow Eager Execution, 전태균 - 코어 (라이트닝톡)
# ---
# 
#
# #### 연사소개
# (주)쎄트렉아이 : TensorFlow KR 운영자, ML Google Developer Experts
#
# #### 발표소개
# 구글이 올해 3월 정식으로 공개한 Eager Execution에 대해서 소개합니다. Eager Execution은 TensorFlow를 대화형 명령 스타일기로 프로그래밍 할 수 있도록 해주는 것입니다. MNIST, linear regression, GAN, RNN, CNN(ResNet) 의 예제를 활용법을 소개합니다.
# 코어 및 알고리즘
#
# #### 발표자료
# * 구글 프리젠테이션 : [보기](http://bit.ly/TFE_DDC)
#
# #### 참고자료
# * [https://github.com/tgjeon/TF-Eager-Execution-Guide-KR](https://github.com/tgjeon/TF-Eager-Execution-Guide-KR)
#
# #### 블로그 및 깃헙
# 깃헙 : [gitHub.com/tgjeon](gitHub.com/tgjeon)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/taegyun.jeon](https://www.facebook.com/taegyun.jeon)
# ### 우리는 왜 강화학습을 해야하는가, 차금강 - 강화학습 (세미나)
# ---
# 
#
# #### 발표소개
# 비전문가들에게 강화학습은 알파고로 대변되고 있습니다. 강화학습이란 무엇인가, 일반 딥러닝과 무엇이 다른것인가, 강화학습을 하기 위해서는 어떤 것들이 필요한가를 짚은 후 여러 적용사례들을 소개드리겠습니다.(적용사례: 벽돌피하기, CartPole(장대세우기), 스타크래프트2)
#
# 
#
# 강화학습이라는 것이 주로 공학에서는 로봇에 많이 사용되고 있고 주로 손로봇에서는 pick and place가 가장 기본적인 문제로 거론되는데 그에 대한 문제를 강화학습으로 충분히 풀 수 있다 라는 것을 의미하고 있습니다
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_cgg_file.pdf)
#
# #### 블로그 및 깃헙
# * 블로그 : 캐글코리아 - [http://kaggle-kr.tistory.com/](http://kaggle-kr.tistory.com/)
# * 깃헙 : [https://github.com/chagmgang](https://github.com/chagmgang)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/profile.php?id=100002147815509](https://www.facebook.com/profile.php?id=100002147815509)
# ### 달려라! 인공지능, 정원석 - 강화학습 (세미나)
# ---
# 
#
# #### 연사소개
# 프리랜서 연구원으로 스마트팩토리에 딥러닝과 강화학습을 적용하는 프로젝트에 참여하고 있으며, 특히 강화학습 연구에 몰두하고 있습니다. 지식과 경험을 나누는 것이 가장 큰 가치라고 생각하여 발표자로 참여하게 되었습니다.
#
# #### 발표소개
# 현재 강화학습은 큰 환경에 적용할때 한계를 보이고 있습니다. 이 한계는 강화학습을 게임에서 적용할때 뿐만 아니라, 미래에 “보편적인 인공지능”을 만들기 위해 우리가 풀어야할 숙제입니다. 이 한계를 해결하기 위해 Atari, Super Mario, Sonic의 환경에 강화학습을 적용하여, 목표를 달성하는 에이전트를 만들기 위해 시도한 과정들을 다음의 순서로 공유하고자 합니다.
#
# 1. 강화학습의 탄생
# 2. 큰 환경에서의 한계점
# 3. To the Rainbow
# 4. 보편적 인공지능의 시작
#
# 
#
# 2013년 인공지능회사 Deepmind가 강화학습에 딥러닝을 적용하여, Atari 게임에서 사람보다 게임을 잘하는 인공지능 에이전트를 만들었습니다. 하지만 화면이 계속 변화며 에이전트가 선택할수 있는 행동의 수가 많아지는 환경에서, 이 학습법의 성능은 매우 좋지 않습니다. 우리가 살고 있는 실제 환경은, 선택할수 있는 행동의 조합이 Atari 게임과 비교할수 없을정도로 많으며, 화면 또한 끊임없이 변하고 있습니다. 또한, 인류는 사물, 사람, 동물 들과 커뮤니케이션하며 공유하며 살기에 서로 밀접한 관계를 맺고 있습니다. 강화학습을 실제 환경에 적용하기 위해 Atari, Super mario, Sonic, Unity ml agent 환경으로 연구한 The Rise of Reinforcement Learning by Wonseok Jung에 대해 공유하고자 합니다.
#
# 강화학습으로 슈퍼마리오 에이전트를 만들수 있는 튜토리얼 또한 제공됩니다.
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_jws_file.pdf)
#
# #### 블로그 및 깃헙
# * 블로그 : [https://wonseokjung.github.io](https://wonseokjung.github.io)
# * 깃헙 : [https://github.com/wonseokjung](https://github.com/wonseokjung)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/ws.jung.798](https://www.facebook.com/ws.jung.798)
# ### 밑바닥부터 만들어보는 핸드메이드 인공지능, 정원석 - 강화학습 (핸즈온랩)
# ---
#
# #### 핸드온랩소개
# 강화학습이 처음이신분들을 위한 시간입니다. Atari 환경에서 DQN 알고리즘을 적용하여 똑똑한 강화학습 에이전트를 만듭니다.
#
# #### 준비물
# 1. 노트북
# 2. Python 3.5 : [https://www.python.org/downloads/](https://www.python.org/downloads/)
# 3. Pytorch : [https://pytorch.org/](https://pytorch.org/)
# 4. OpenAI gym : [https://github.com/openai/gym](https://github.com/openai/gym)
# 5. Jupyter notebook : [http://jupyter.org/](http://jupyter.org/)
# 6. Numpy
# 7. Matplotlib : [https://matplotlib.org/](https://matplotlib.org/)
# ### 알파오목이랑 게임하기, 김정대 - 강화학습 (세미나)
# ---
# 
#
# #### 연사소개
# 알파고를 사랑한 수학강사. 셀프 플레이 강화학습을 실제 삶에 적용하는 사람. 퇴근 후에 두뇌가 풀가동되는 신인류.
#
# #### 발표소개
# 알파고 제로에 대한 기본적인 원리에 대해서 알아보고, 이 알고리즘을 이용하여 알파오목을 구현한 경험을 공유합니다. 사람과 알파오목 대전을 보실 수도(?) 있습니다.
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_kjd_file.pdf)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/kekmodel](https://www.facebook.com/kekmodel)
# ### 안.전.제.일. 강화학습!, 이동민 - 강화학습 (세미나)
# ---
# 
#
# #### 연사소개
# 강화학습을 사랑하고 공부하고 있는 학부생입니다. 강화학습은 저의 연구분야이며, 취미이며, 일상입니다.
#
# #### 발표소개
# 본격적인 내용에 앞서 우리가 왜 강화학습을 공부해야 하는지, 강화학습이란 무엇인지에 대해서 알아보겠습니다. 그리고 ‘어떻게 하면 강화학습을 더 안전하고 빠르게 학습할 수 있을까?’에 대해서 소개하고자 합니다.
#
# 1. What is Reinforcement Learning?
# 2. Artificial General Intelligence
# 3. Planning & Learning
# 4. Safe Reinforcement Learning
#
# 
#
# #### 발표자료
# * 슬라이드쉐어 : [보기](https://www.slideshare.net/DongMinLee32/ss-103395612)
#
# #### 참고자료
# * 슬라이드쉐어 : [강화학습의 개요](https://www.slideshare.net/DongMinLee32/ss-91521646)
# * 슬라이드쉐어 : [강화학습 알고리즘의 흐름도 Part 1](https://www.slideshare.net/DongMinLee32/part-1-91522059)
# * 슬라이드쉐어 : [강화학습 알고리즘의 흐름도 Part 2](https://www.slideshare.net/DongMinLee32/part-2-91522217)
# * 슬라이드쉐어 : [Planning and Learning with Tabular Methods](https://www.slideshare.net/DongMinLee32/planning-and-learning-with-tabular-methods)
#
# #### 블로그 및 깃헙
# * 블로그 : [https://dongminlee.tistory.com](https://dongminlee.tistory.com)
# * 깃헙 : [https://github.com/dongminleeai](https://github.com/dongminleeai)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/dongminleeai](https://www.facebook.com/dongminleeai)
# ### GTA로 자율주행, 김준태 - 강화학습 (세미나)
# ---
# 
#
# #### 연사소개
# 대전대학교 전자정보통신공학과 학부생이고 대학원 진학 예정입니다.
#
# #### 발표소개
# GTA란 게임을 이용하여 어떻게 주행 데이터를 모으고 학습을 한 후 자율주행을 하는지 알아보고, GTA를 강화학습을 하기 위해 밑바닥부터 환경 구축을 하고 있는 제 이야기를 듣고 같이 이야기 해봐요!!
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_kjt_file.pdf)
#
# #### 블로그 및 깃헙
# * 블로그 : 캐글코리아 - [http://kaggle-kr.tistory.com/](http://kaggle-kr.tistory.com/)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/kjt7889](https://www.facebook.com/kjt7889)
# ### Introduction to GAN, 이지민 - 의료 (세미나)
# ---
# 
#
# #### 연사소개
# 서울대학교 방사선의학물리연구실 / TensorFlowKR 운영진 : 늘 행복하고 싶은 대학원생이며, 딥러닝을 이용하여 의료영상을 분석하는 연구를 하고 있습니다. :)
#
# #### 발표소개
# 생성 모델(Generative Model) 인 Auto-Regressive model, Variational Auto-Encoder, GAN (Generative Adversarial Networks)에 대해 간단히 소개하고, 그 중 가장 주목받고 있는 GAN 의 심화 개념과 응용 사례도 함께 다루어보고자 합니다.
#
# #### 발표자료
# * 슬라이드 쉐어 : [Introduction to GAN](https://www.slideshare.net/JiminLee36/introduction-to-gan)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/jimin.lee.5623](https://www.facebook.com/jimin.lee.5623)
# ### TensorFlow Object Detection API를 활용한 대장 종양 검출, 김영하 - 의료 (라이트닝톡)
# ---
# 
#
# #### 연사소개
# 데이터를 사랑하는 사람들과 디플러스에서 즐겁게 연구하는 연구원. 재미있는 주제를 찾아 연구하고 같이 재미있는 것을 만들어보고자 합니다.
#
# #### 발표소개
# TensorFlow에서 Object Detection API를 제공하고 있으며, COCO, Kitti, Open Images 데이터셋으로 학습이 된 모델들도 제공하고 있습니다. 이 모델에서 제공하는 클래스이외에 의학영상 및 동영상 (대장 내시경)에서 종양을 검출할 수 있는지에 대한 연구를 하고 있습니다.
#
# 
#
# 자신만의 Object Detection을 하기 위한 데이터셋 준비 > 종양을 검출하기 위해 데이터셋으로 기존 모델에 학습 > 학습된 결과로 종양 검출 확인
#
# #### 참고자료
# * [https://github.com/tensorflow/models/tree/master/research/object_detection](https://github.com/tensorflow/models/tree/master/research/object_detection)
#
# #### 블로그 및 깃헙
# * 블로그 : [https://brunch.co.kr/@fermat39](https://brunch.co.kr/@fermat39)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/profile.php?id=100001647473870](https://www.facebook.com/profile.php?id=100001647473870)
# ### MRI 를 이용한 치매 질환 예측, 박세진 - 의료 (세미나)
# ---
# 
#
# #### 연사소개
# 컴퓨터 비전을 좋아하는 아들바보입니다. 관심분야는 의료영상, 자율주행.
#
# #### 발표소개
# 객체인식 기법과 머신러닝을 사용하여 MRI 영상에서 관심 뇌영역을 분할하고 뇌위축을 인식하여 치매(알츠하이머) 위험도를 예측하는 기법을 소개합니다.
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_psj_file.pdf)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/profile.php?id=100002023963861](https://www.facebook.com/profile.php?id=100002023963861)
# ### 나의 감성을 알아주는 너, 송규예 - 감성 (세미나)
# ---
# 
#
# #### 연사소개
# 감성기술로 행복한 세상을 만들어가고 있는 orbis.ai(오르비스에이아이) 기업의 공동창업자이자, 데이터과학자인 송규예 입니다. 감성인식과 감성표현을 위한 딥러닝 모델개발을 하고 있습니다. 개인적으로는 사람의 세포부터 무의식까지, 사람에 대한 모든 것을 파헤치고 있으며 사람, 자연 구조와 닮은 기술구현에 관심이 많습니다.
#
# #### 발표소개
# "딥러닝 X 심리" . 어울릴 듯, 어울리지않는 두 분야의 콜라보레이션 ☆사람의 감정인식을 위한 딥러닝 기술과 감정표현 측면에서 활용할 수 있는 딥러닝 기술에는 무엇이 있을지 다뤄보며, 저희 회사, 오르비스에이아이에서 진행하고 있는 감정인식 기술에 대해서 설명합니다.
#
# 1. intro : 딥러닝과 심리의 운명적 만남 (부제 : 나는 네가 필요해)
# 2. 감정인식 및 감정표현 딥러닝 기술 소개
# 3. 오르비스에이아이에서 진행하는 감정인식 기술 소개
#
# 
#
# 따뜻한 "공감"과 행동유도의 핵심 "동기부여"를 통해 잠재되어 있던 무의식을 의식상태로 끌어올려 행복을 도모합니다. 이 과정에서 딥러닝 기술이 어떻게 적용되는지 집중탐구합니다.
#
# #### 발표자료
# * 구글 드라이브 : [보기](https://docs.google.com/presentation/d/1Fl82bA_-oBttNUs5gSE2R1Ta4ZRhqy-WmsqJ66oY2vc/edit?usp=sharing)
#
# #### 블로그 및 깃헙
# * 깃헙 : [https://github.com/Kyuye](https://github.com/Kyuye)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://facebook.com/ivyheart2](https://facebook.com/ivyheart2)
# ### 근전도 생체신호 데이터로 손 모양 생성하기, 박상민 - 임베디드 (세미나)
# ---
# 
#
# #### 연사소개
# 개발자를 꿈꾸는 한국의 어느 평범한(?) 고3 입니다. 대덕SW마이스터고등학교 임베디드과를 재학중이며, 임베디드SW와 딥러닝에 관심이 많아 주로 공부하고 있습니다. 이론적으로, 전문적으로 한 분야를 깊게 알지는 못하지만, 능력있는 개발자가 되기위해 열심히 달려가고 있습니다. 전문가분들의, 개발자분들의 따끔한 충고와 조언은 언제든지 환영입니다~
#
# #### 발표소개
# 사람의 팔에는 EMG라는 '근전도 생체 신호'가 나옵니다. 이를 이용하여 손 모양을 생성해 내고자 합니다. 근전도 생체신호와 딥러닝의 이미지 생성모델(GAN)로 손의 모양을 생성해내는 프로젝트를 소개해드리겠습니다. 처음에는 혼자 시작했지만 현재는 능력있고, 멋진 팀원들과 함께 팀을이루어 프로젝트를 진행하고 있습니다. 프로젝트를 진행하면서 어떤 문제들이 있었고, 어떻게 해결하려고 했는지, 또 어떤 삽질들을 했는지 '고등학생의 딥러닝 프로젝트 삽질기'를 소개해 드리고자 합니다.
#
# 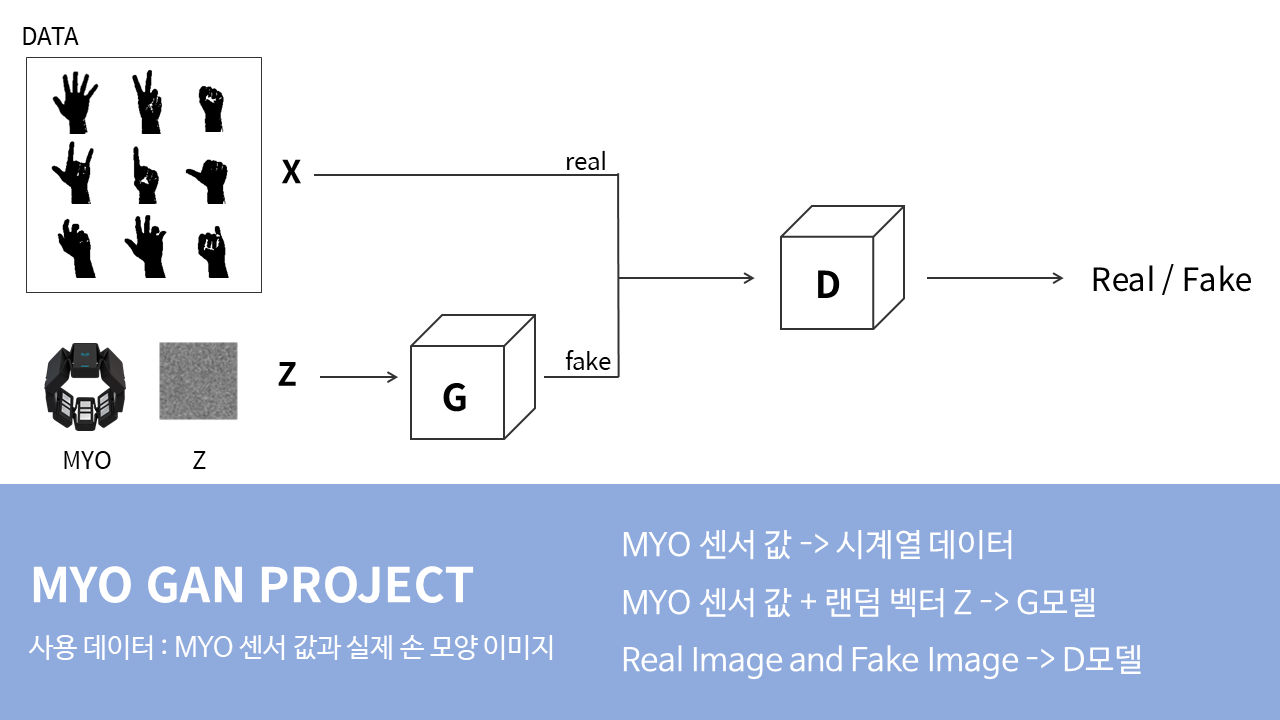
#
# MYO의 센서값과 실제 손 모양 이미지를 이용해 GAN모델을 학습시킨다면 이 모델은 MYO의 센서값만으로도 손 모양 이미지를 생성해 낼 수 있을 것 입니다. 이를 이용한다면 많은 분야에 적용하여 사용할 수 있지 않을까요?
#
# #### 발표자료
# * 구글 드라이브 : [보기](https://drive.google.com/file/d/1CBiAEaz04P2yzzECUkLpVzhwuct22Bli/view?usp=sharing)
#
# #### 블로그 및 깃헙
# * 깃헙 : https://github.com/jigeria
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/jigeria114](https://www.facebook.com/jigeria114)
# ### 딥러닝과 함께하는 최적설계와 시뮬레이션, 원자력분야의 적용사례, 유용균 - 공학 (세미나)
# ---
# 
#
# #### 연사소개
# 원자력연구원 선임연구원. 딥러닝 덕질 중 연구원내 인공지능 연구 전담조직을 목표로 삽질 중.
#
# #### 발표소개
# 본 세미나에서는 전문적인 딥러닝 지식보다 그 동안 알려지지 않았던 원자력 및 기계공학 분야의 응용사례에 대해서 소개
#
# 1. 계산과학 분야의 머신러닝 적용
# 2. 최적설계와 딥러닝의 융합
# 3. 딥러닝과 최적설계를 융합한 뼈 CT 사진 고해상화
# 4. 원자력 분야의 머신러닝 적용 사례
#
# 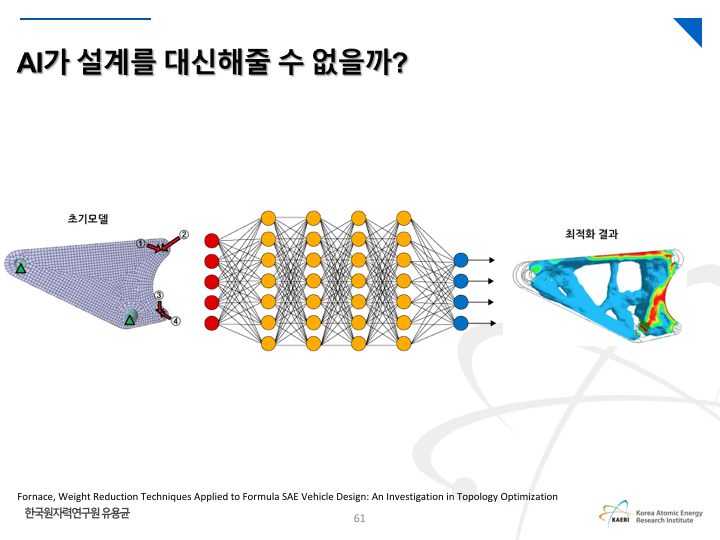
#
# 기존 최적설계방법론은 반복적인 해석이 필요하기 때문에 결과를 얻기위하여 많은 시간이 요구됩니다. 합니다. 고전적인 최적설계 방법론 대신 인공신경망을 이용하면 최적의 구조를 빠르게 찾을 수 있지 않을까요?
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_yyk_file.pdf)
#
# #### 연락처
# * 이메일 : <EMAIL>, <EMAIL>
# * 페이스북 : [https://www.facebook.com/yoyogo96](https://www.facebook.com/yoyogo96)
# ### 항공우주분야에서의 머신러닝기술, 김홍배 - 공학 (라이트닝톡)
# ---
# 
#
# #### 연사소개
# 한국항공우주연구원 인공지능연구실: 구조진동 전공. 시험평가 전문가. 우주 광학 전문가. 컴퓨터 비젼과 머신 러닝분야에 관심이 많고 현재는 머신러닝기술의 항공우주분야 응용에 집중하고 있습니다.
#
# #### 발표소개
# 머신러닝기술이 항공우주분야에 어떻게 적용되고 있는지 간단하게 소개하고 참가자들과 토론하고자 합니다.
#
#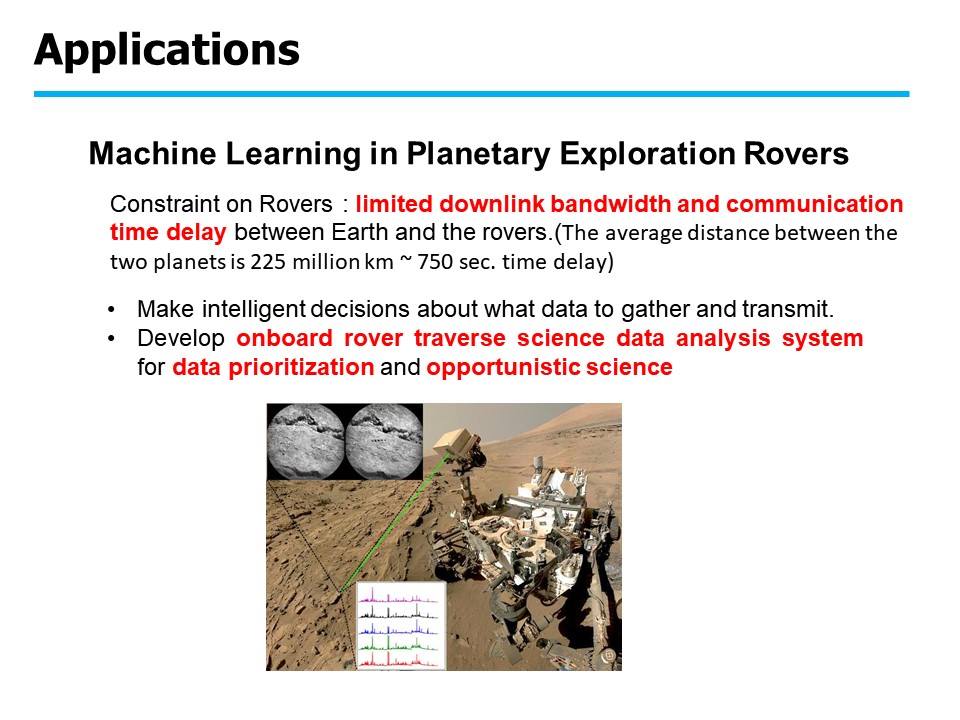 
#
# Autonomous Exploration for Gathering Increased Science(AEGIS) 화성이나 달 탐사용 Rover의 자율탐사시스템으로 데이터의 전송 지연등에 의한 탐사지연을 최소화 하기 위하여 NASA에서 개발 중인 인공지능 기술. 미리 지정된 목표인지, 처음 탐지된 대상인지 등을 인공지능 기술을 이용하여 결정하고 추가적인 탐사 여부와 지구로의 데이터 전송등을 효율적으로 관리합니다.
#
# #### 발표자료
# * 슬라이드쉐어 : [보기](https://www.slideshare.net/ssuser06e0c5/machine-learning-applications-in-aerospace-domain)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/profile.php?id=100001173798815](https://www.facebook.com/profile.php?id=100001173798815)
# ### 딥러닝을 활용한 테스트 자동화 연구, 곽병권 - 품질 (라이트닝톡)
# ---
# 
#
# #### 연사소개
# NGLE이라는 QA 전문 회사에서 QA-Test 자동화에 인공지능 적용을 연구
#
# #### 발표소개
# 모바일 게임 환경에서의 블랙박스 테스트 상황에서의 딥러닝 적용 방안 연구, Imitation Learning, Behavioral-Cloning등의 적용등을 소개합니다.
#
# 
#
# 저희 회사에서 주로 하는 게임 품질관리와 테스팅을 설명 한 책입니다. 게임 테스팅이 예상처럼 쉽지 않고 효율적인 테스팅을 위해서는 지치지 않고 누락없는 반복 테스팅이 필요한데, 이를 위해 인공지능을 활용하는 방법을 연구하고 있습니다.
#
# #### 발표자료
# * PDF : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_kbg_file.pdf)
#
# #### 블로그 및 깃헙
# * 깃헙 : [https://github.com/Steven-A3](https://github.com/Steven-A3)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/byeongkwon.kwak](https://www.facebook.com/byeongkwon.kwak)
# ### 우주과학 이미지 자료 변환하기, 신경인, 박은수, 이하림 - 태양 분야 적용 사례 (세미나)
# ---
# 
#
# #### 연사소개
# 경희대학교 우주탐사학과 태양 & 우주기상 연구실 박은수, 이하림, 신경인 연구원. 딥러닝 모델을 설계하고 우주기상분야에 적용하면서 과학기술 분야와 딥러닝 사이의 연결고리가 되고 싶은 대학원생들입니다.
#
# #### 발표소개
# 우주기상 분야에서의 딥러닝 적용 사례 (박은수)
#
# 
#
# 많은 관측 위성이 다양한 파장대에서 태양 영상을 관측하고 있습니다. 태양 영상을 다른 종류의 영상으로 바꾸어 보았으며, 이를 이용하여 데이터 결손을 복원하거나, 두 데이터간의 물리적 연관성을 찾고자 하는 시도를 하고 있습니다.
#
# 태양 흑점 스케치로 자기장 지도 생성, 400년전 기록을 복원! (이하림)
#
# 
#
# 갈릴레오는 1600년대 초 최초로 태양 흑점을 관측하였습니다. 이 자료와 딥러닝을 사용하여 최초의 태양 자기장 지도를 생성한 결과에 대하여 발표하겠습니다.
#
# 갓 pix2pix 변신은 무죄 (신경인)
#
# 
#
# Pix2Pix를 개선한 모델로 생성한 이미지와 실제 이미지의 비교입니다. 왼쪽 이미지가 모델이 생성한 이미지, 오른쪽 이미지가 실제 이미지입니다. 확대된 부분을 비교해보았을 때, 세밀한 구조까지 복원할 수 있음을 알 수 있습니다.
#
# #### 발표자료
# * 박은수 발표자료 : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_pes_file.pdf)
# * 이하림 발표자료 : [다운](http://tykimos.github.io/warehouse/2018-6-28-ISS_1st_Deep_Learning_Together_lhl_file.pdf)
#
# #### 블로그 및 깃헙
# * 신경인 깃헙 : [https://github.com/GyunginShin](https://github.com/GyunginShin)
#
# #### 연락처
# * 박은수 이메일 : <EMAIL>
# * 이하림 이메일 : <EMAIL>
# * 신경인 이메일 : <EMAIL>
# ### 내 손 위의 딥러닝(모바일에 딥러닝 심기), 전미정 - 모바일 (세미나)
# ---
# 
#
# #### 연사소개
# 안녕하세요, iOS 개발자 미정입니다. ☕️를 마시며 하루를 시작하는 걸 좋아합니다. 가끔📱개발을 하며 주로 🐈와 놀거나 잡니다. 최근에는 딥러닝에 관심이 생겨 모바일에 딥러닝을 심기 위한 다양한 방법을 공부하고있습니다 🙂
#
# #### 발표소개
# 딥러닝이 컴퓨터를 떠나 모바일에 들어오기 시작했습니다. 서버 통신을 하지 않고 모바일에서 구동되는 딥러닝이 어떤 의미가 있으며, 어떻게 구현할 수 있는지, 어떤 모델들을 심을 수 있는지 개발자 입장에서 정리하고 직접 구현해봤습니다. 실제로 내 손위에서 어떻게 딥러닝이 작동하는지 소개해드리겠습니다.
#
# 
#
# 외부에서 훈련된 딥러닝 모델을 모바일 기기에 심어 모바일 딥러닝을 구현해보았습니다. (쉿, 이건 비밀인데 모바일 자체에서도 훈련이 되기 시작했습니다.)
#
# #### 발표자료
# * 슬라이드 쉐어 : [보기](https://www.slideshare.net/MijeongJeon1/ios-103594100)
#
# #### 참고자료
# * [https://github.com/MijeongJeon/ShapeDetector_Keras_CoreML](https://github.com/MijeongJeon/ShapeDetector_Keras_CoreML)
#
# #### 블로그 및 깃헙
# * 블로그 : https://mijeongjeon.github.io
# * 깃헙 : https://github.com/mijeongjeon
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/Ninevincent](https://www.facebook.com/Ninevincent)
# ### TensorFlow Object Detection API를 활용한 대장 종양 검출, 김영하 - 의료 (세미나)
#
# ---
# 
#
# #### 연사소개
# 데이터를 사랑하는 사람들과 디플러스에서 즐겁게 연구하는 연구원. 재미있는 주제를 찾아 연구하고 같이 재미있는 것을 만들어보고자 합니다.
#
# #### 발표소개
# Tensorflow에서 Object Detection API를 제공하고 있으며, COCO, Kitti, Open Images 데이터셋으로 학습이 된 모델들도 제공하고 있습니다. 이 모델에서 제공하는 클래스이외에 의학영상 및 동영상 (대장 내시경)에서 종양을 검출할 수 있는지에 대한 연구를 하고 있습니다.
#
# 
#
# 자신만의 Object Detection을 하기 위한 데이터셋 준비 > 종양을 검출하기 위해 데이터셋으로 기존 모델에 학습 > 학습된 결과로 종양 검출 확인
#
# #### 참고자료
# * [https://github.com/tensorflow/models/tree/master/research/object_detection](https://github.com/tensorflow/models/tree/master/research/object_detection)
#
# #### 블로그 및 깃헙
# * 블로그 : [https://brunch.co.kr/@fermat39](https://brunch.co.kr/@fermat39)
#
# #### 번역서 및 저서
# * Splunk 앱 제작과 대시보드 개발(에이콘출판사) : 빅데이터 플랫폼 Splunk 플랫폼의 앱 개발 설명 [바로가기](http://www.kyobobook.co.kr/product/detailViewKor.laf?ejkGb=KOR&mallGb=KOR&barcode=9788960778207&orderClick=LAG&Kc=)
# * 파이썬 웹 스크래핑(에이콘출판사) : scrapy, beautifulsoup를 활용한 웹 스크래핑 [바로가기](http://www.kyobobook.co.kr/product/detailViewKor.laf?ejkGb=KOR&mallGb=KOR&barcode=9788960779594&orderClick=LAH&Kc=)
# * 뷰티플 자바스크립트(비제이퍼블릭) : 자바스크립트 중급서 [바로가기](http://www.kyobobook.co.kr/product/detailViewKor.laf?mallGb=KOR&ejkGb=KOR&orderClick=LEB&barcode=9791186697184)
# * 누구나 쉽게 배우는 스몰베이직(비제이퍼블릭) : 스몰베이직 초급서 [바로가기](http://www.kyobobook.co.kr/product/detailViewKor.laf?ejkGb=KOR&mallGb=KOR&barcode=9791186697375&orderClick=LAH&Kc=)
# * 머신러닝을 이용한 이미지 분석(비제이퍼블릭) : 텐서플로를 이용해서 자신만의 데이터셋 제작 및 이미지 학습 및 분석 (집필중)
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/profile.php?id=100001647473870](https://www.facebook.com/profile.php?id=100001647473870)
# ### 서포터즈 - 이상훈
# ---
# 
#
# 삼성생명 DA Lab 근무중, 케라스 코리아, 스파크 사용자 모임 운영자입니다. 현재 이미지 인식과 텍스트 처리 관련된 업무를 하고 있고 분산환경에서의 hyperparameter 튜닝에 관심이 많습니다.
#
# #### 저서, 역서
# * 빅데이터 실무 기술 가이드 공동 저자
# * 실시간 분석의 모든것 역자
#
# #### 연락처
# * 이메일 : <EMAIL>
# * 페이스북 : [https://www.facebook.com/phoenixlee.sh](https://www.facebook.com/phoenixlee.sh)
# ### 서포터즈 - 황준오
# ---
# 
#
# 부경대학교 컴퓨터공학과에 재학중인 4학년 학생입니다. 원활한 컨퍼런스 진행을 위해 열심히 서포트 하겠습니다!
#
# #### 프로필
# * 이메일 : <EMAIL>
# * 깃헙 : [https://github.com/bactoria](https://github.com/bactoria)
# ### 서포터즈 - 홍정훈
# ---
# 
#
# 미시간 대학교에서 데이터사이언스를 공부하고 있습니다.
#
# #### 프로필
# * 이메일 : <EMAIL>
# ### 서포터즈 - 조수현
# ---
# 
#
# 숭실대학교에서 하이브리드 기계학습 기반 예측문제에 관한 연구를 주로 했습니다. 현재 오스템임플란트 e-commerce 연구실에서 서버개발자로 일하고 있습니다. RL과 GANs에 관심이 많습니다. 다양한 방식으로 지식과 도움을 나누는 사람이 되고자 합니다. 고기, 회 그리고 커피를 좋아합니다. 소박한 영역에서 역할을 다하여 원활한 컨퍼런스가 되도록 하겠습니다.
#
# #### 프로필
# * 이메일 : <EMAIL>
# ### 서포터즈 - 신채원
# ---
# 
#
# 한밭대학교 산업경영공학과 4학년입니다.
#
# #### 프로필
# * 이메일 : <EMAIL>
# ### 서포터즈 - 이병호
# ---
# 
#
# 카이스트와 퍼듀에서 기하학을 전공하였습니다. 주로 자연어처리와 음악 생성, 그리고 딥러닝의 수학적 측면에 관심을 갖고 있습니다.
#
# #### 프로필
# * 이메일 : <EMAIL>@kaist.ac.kr
# * 페이스북: [https://www.facebook.com/arthematist](https://www.facebook.com/arthematist)
# ### 서포터즈 - 강천성
# ---
# 
#
# 충남대학교 컴퓨터공학과 4학년 재학중, 캐글을 시작으로 데이터 사이언스 분야에 발을 들여놓았습니다. 현재 음성 변조 관련 졸업 프로젝트를 진행중이며, 머신러닝 및 딥러닝 관련 기술에 관심이 많습니다.
#
# #### 프로필
# * 이메일 : <EMAIL>
# * 블로그 : 캐글코리아 - [http://kaggle-kr.tistory.com/](http://kaggle-kr.tistory.com/)
# ### 서포터즈 - 김은희
# ---
# 
#
# KAIST에서 기계학습 기반 사용자 이용내역 데이터 기반 토픽모델링 확장 추천알고리즘으로 박사학위를 수여했으며, 딥러닝 기반의 운전자 프로파이링 알고리즘, ECG이용 졸음 운전 감지등의 프로젝트를 수행해 왔습니다. 다양한 데이터로 재미있는 결과물들을 얻어내는 프로젝트들에 관심이 있고, 또 이를 잘 하시는 기업과 연사들에게 관심이 있습니다. 현재 충남대학교 SW중심대학의 사업의 하나로 학생들의 프로젝트를 완성도있게 도와주실 수 있는 기업체들과 다양한 산업 주제들을 제안해주고 싶으신 함께하실 연사 분들을 멘토로 모시고자 딥러닝 컨퍼런스 서포터즈로 활동하려고 합니다. 좋은 인연 많이 만들게 되길 기대합니다~
#
# #### 프로필
# * 이메일 : <EMAIL>
# ---
# ### 상세 프로그램
#
# 아래 스케줄표는 변동될 수 있습니다.
#
# |구분|강당|사이언스홀|대회의실|
# |-|-|-|-|
# |09:50~10:10(20분)|발표자소개|발표자소개|발표자소개|
# |10:10~10:55(45분)|박은수,이하림,신경인<br>우주과학 이미지 자료 변환하기|이지민<br>Introduction to GAN|이유한<br>타이타닉 생존자 예측을 해보자|
# |10:55~11:15(20분)|전태균<br>TensorFlow Eager Execution|김홍배<br>항공우주분야에서의 머신러닝기술|이유한<br>타이타닉 생존자 예측을 해보자|
# |11:15~12:00(45분)|유재준<br>Deep convolutional framelets: application to diffuse optical tomography|이유한<br>딥러닝 보물상자 ‘캐글’ 알아보기|-|
# |12:00~13:00(60분)|점심|점심|점심|
# |13:00~13:45(45분)|차금강<br>우리는 왜 강화학습을 해야하는가|송규예<br>나의 감성을 알아주는 너|-|
# |13:45~14:30(45분)|이동민<br>안.전.제.일. 강화학습!|박세진<br>MRI 를 이용한 치매 질환 예측|-|
# |14:30~15:15(45분)|박상민<br>근전도 생체신호 데이터로 손 모양 생성하기|정원석<br>달려라! 인공지능|-|
# |15:15~15:30(15분)|휴식|휴식|휴식|
# |15:30~16:15(45분)|유용균<br>딥러닝과 함께하는 최적설계와 시뮬레이션|김준태<br>GTA로 자율주행|정원석<br>밑바닥부터 만들어보는 핸드메이드 인공지능|
# |16:15~16:35(20분)|곽병권<br>딥러닝을 활용한 테스트 자동화 연구|김영하<br>TensorFlow Object Detection API를 활용한 대장 종양 검출|정원석<br>밑바닥부터 만들어보는 핸드메이드 인공지능|
# |16:35~17:05(30분)|김태영<br>딥러닝 몰라도 간(GAN)부터 보기|전미정<br>내 손 위의 딥러닝(모바일에 딥러닝 심기)|-|
# |17:05~17:50(45분)|류성옥<br>Graph Neural Network|김정대<br>알파오목이랑 게임하기|-|
# ---
#
# ### 참가신청 - 마감되었습니다!
#
# 댓글로 참가신청을 받습니다. 댓글 양식은 아래와 같으며 '한 줄'로 작성 부탁드리겠습니다.
#
# * 이름, 기관, 이메일, 분야, 참석계기
# * 예) 김태영, 인스페이스, <EMAIL>, 우주, 위성 운영 효율화를 위해 강화학습을 적용해보고자 합니다.
#
# 댓글을 달아도 스팸처리되어서 바로 표시 안될 수도 있습니다. 제가 다 스팸아님으로 처리하고 있으니, 크게 신경 안 쓰셔도 됩니다. 그리고 혹시 참석신청하셨으나 부득이한 이유로 참석이 힘드신 분은 미리 알려주세요~ 다른 분들에게 참석 기회를 드리고자 합니다.
# ### 후기
#
# 연사분들 및 서포터즈분들입니다. 고생많으셨습니다! 내년에도 또 뵈어요~
#
# 
# ---
#
# ### 같이 보기
#
# * [다른 세미나 보기](https://tykimos.github.io/seminar/)
# * [케라스 기초 강좌](https://tykimos.github.io/lecture/)
# * [케라스 코리아](https://www.facebook.com/groups/KerasKorea/)
| _writing/2018-6-28-ISS_1st_Deep_Learning_Conference_All_Together.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import matplotlib.pyplot as plt
import numpy as np
import pickle
import os
# -
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
GPU = "0"
os.environ["CUDA_VISIBLE_DEVICES"]=GPU
from frm_dataset_loader import load_dataset
from conf_dataset_1 import *
# +
fname = 'datasets/valid_1.dat'
(comb_valid,carrier_valid,clean_valid,fading_valid,_,
freq_valid,timing_offNum_valid,timing_step_valid,
_,mod_valid,snr_valid) = load_dataset(fname,max_sps,len(mod_list))
# +
fname = 'datasets/test_1.dat'
(comb_test,carrier_test,clean_test,fading_test,raw_test,
freq_test,timing_offNum_test,timing_step_test,
coeff_test,mod_test,snr_test) = load_dataset(fname,max_sps,len(mod_list))
# +
from frm_nn_zoo_01 import create_dualPath
pkt_size_net = pkt_size
nn_all = create_dualPath(pkt_size = pkt_size_net,n_mods=len(mod_list))
nn_all.summary()
# +
from frm_train_generator import train_generator
gen = train_generator(100,64,pkt_size,max_sps,mod_list,
sps_rng,pulse_ebw_list,timing_offset_rng,fading_spread_rng,freq_err_rng,phase_err_rng,snr_rng)
len(gen.__getitem__(0)[1])
# +
# nn_all.load_weights('../models/033.h5')
import keras
patience = 10
n_epochs = 100
decay_rate = None #0.8
decay_step = None #25
def step_decay(epoch,lr):
decay_rate = decay_rate
decay_step = decay_step
if epoch % decay_step == 0 and epoch:
return lr * decay_rate
return lr
filepath = 'tmp/tmp_'+GPU
import time
c=[keras.callbacks.EarlyStopping(monitor='loss', min_delta=0.0, patience=patience, verbose=1, mode='auto'),
keras.callbacks.ModelCheckpoint(filepath, monitor='val_loss', verbose=1, save_best_only=True,
save_weights_only=True),]
#keras.callbacks.LearningRateScheduler(step_decay)]
indx_valid = slice(None,None)
batch_size = 200
samples_per_epoch = 800000
gen = train_generator(samples_per_epoch,batch_size,pkt_size,max_sps,mod_list,
sps_rng,pulse_ebw_list,timing_offset_rng,fading_spread_rng,freq_err_rng,phase_err_rng,snr_rng)
history = nn_all.fit_generator(gen,
epochs = n_epochs,callbacks=c,
workers=10, use_multiprocessing=True,
validation_data =
(comb_valid[indx_valid],[freq_valid[indx_valid],fading_valid[indx_valid],fading_valid[indx_valid],
clean_valid[indx_valid],timing_step_valid[indx_valid],
timing_offNum_valid[indx_valid],mod_valid[indx_valid]]))
# +
plt.plot(history.history['loss'][1:])
plt.plot(history.history['val_loss'][1:])
plt.title('Model Loss')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Valid'], loc='upper left')
plt.show()
print(history.history)
# -
filepath = 'tmp/tmp_'+GPU
nn_all.load_weights(filepath)
op_eval = nn_all.evaluate(comb_test,[freq_test,fading_test,fading_test,clean_test,timing_step_test,timing_offNum_test,mod_test])
print(op_eval)
# +
import datetime
outputs = {}
outputs['dataset_params'] = [pkt_size,max_sps,mod_list,sps_rng,pulse_ebw_list,timing_offset_rng,fading_spread_rng,freq_err_rng,phase_err_rng,snr_rng]
outputs['date'] = f'{datetime.datetime.now():%Y-%m-%d %H:%M:%S%z}'
outputs['history'] = history.history
outputs['train_params'] = [patience,n_epochs,decay_rate,decay_step]
outputs['op_eval'] = op_eval
# -
FNAME = '001'
import pickle
with open(f'outputs/{FNAME}.pkl','wb') as f:
pickle.dump(outputs,f)
# +
nn_all.save_weights(f'models/{FNAME}.h5')
# nn_demod.save('models/{Di}_sig_data.h5'.format(",".join(mod_list)))
# -
| 001_d1_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf2
# language: python
# name: tf2
# ---
# # Abalone
#
# In this example, we'll demonstrate how to use [dataduit](https://github.com/JackBurdick/dataduit) to create tensorflow datasets from a pandas dataframe by specifying a config file.
#
# We'll then demonstrate how to use yeahml to create/build/evaluate a model on the created data.
#
# #### Note:
# > The model for this project likely doesn't make sense. I am not personally familiar with the dataset/problem, I was interested showing an example
# %load_ext autoreload
# %autoreload 2
# %load_ext nb_black
import pandas as pd
import tensorflow as tf
import dataduit as dd
import yeahml as yml
import matplotlib.pyplot as plt
# ## Create Datasets
# Reading a file from online
# more information can be found here:
# > https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/
h = [
"sex",
"length",
"diameter",
"height",
"whole_weight",
"shucked_weight",
"viscera_weight",
"shell_weight",
"rings",
]
df = pd.read_csv(
"https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.data",
names=h,
)
# only use 2 of the features
dd_dict = {
"meta": {
"name": "abalone",
"logging": {"log_stream_level": "INFO"},
"in": {"from": "memory", "type": "pandas"},
},
"read": {
"split_percents": [75, 15, 10],
"split_names": ["train", "val", "test"],
"iterate": {
"return_type": "tuple",
"schema": {
"x": {
"length": {
"indicator": "length",
"datatype": {
"in": {"options": {"dtype": "float64", "shape": 1}},
"out": {},
},
"special": "decode",
},
"diameter": {
"indicator": "diameter",
"datatype": {
"in": {"options": {"dtype": "float64", "shape": 1}},
"out": {},
},
"special": "decode",
},
},
"y": {
"rings": {
"datatype": {
"in": {"options": {"dtype": "int64", "shape": 1}},
"out": {},
}
}
},
},
},
},
}
# create the datasets based on the above defined names/splits/specifed data
ds_dict_raw = dd.read(dd_dict, df)
ds_dict = {"abalone": ds_dict_raw[0]}
# `ds_dict` is a dictionary containing the tensorflow datasets (as specified above). which can be accessed like this:
#
# ```python
# ds_val = ds_dict["val"]
# ```
# ## Specify the Model
# +
# # %load_ext autoreload
# # %autoreload 2
# import yeahml as yml
# import tensorflow as tf
# -
example = "./main_config.yml"
yml_dict = yml.create_configs(example)
yml_dict["callbacks"]
# ## Build the model
# If you receive an error:
# AttributeError: 'google.protobuf.pyext._message.RepeatedCompositeCo' object has no attribute 'append'
# I personally used `pip install -U protobuf=3.8.0` to resolve
# per https://github.com/tensorflow/tensorflow/issues/33348
model = yml.build_model(yml_dict)
model.summary()
# ## Train the Model
#
# Notice here that we're using the created training and validation sets from `ds_dict`
# +
# a = ds_dict["abalone"]['train']
# print(a)
# +
# import pprint
# pprint.pprint(yml_dict['optimize'])
# print("--")
# pprint.pprint(yml_dict['performance'])
# ds_dict = {
# "abalone": {
# "train": tf.data.Dataset.from_tensor_slices([1, 2, 3]),
# "val": tf.data.Dataset.from_tensor_slices([1, 2, 3]),
# }
# }
# -
ds_dict
train_dict = yml.train_model(model, yml_dict, ds_dict)
yml.basic_plot_tracker(
train_dict["tracker"],
skip=8,
metrics=True,
local=False,
training=True,
validation=False,
loss=True,
size=(16, 8),
)
# ## Evaluate the Model
eval_dict = yml.eval_model(
model, yml_dict, datasets=ds_dict, eval_split="test", pred_dict={"write": None}
)
# print(eval_dict)
# TODO: will need to revist this problem and build a 'real' model at some point
in_hash = list(eval_dict["abalone"].keys())[0]
plt.scatter(
eval_dict["abalone"][in_hash]["out"]["pred"],
eval_dict["abalone"][in_hash]["out"]["target"],
)
# ## Inspect model in Tensorflow
#
# In the command line you can navigate to the `albalone` directory and run: (provided tensorboard is installed in your environment)
#
# ```bash
# tensorboard --logdir model_a/
# ```
| examples/abalone/abalone.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # This notebook shows an example on how to calibrate parameteres using goal seeking (Targer/instruments)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why specify a model in Latex?
#
# Sometime the **implementation** of a model in software don't match the **specification** of the model in
# the text in which the model is presented. It can be a chalange to make sure that the specification is
# updated in order to reflect changes made in the implementation.
#
# By extracting the model from a Latex script which describes and specify the model a one can always be sure that simulations reflect the model as described in the paper.
#
# Also the author is forced to make a complete specification of the model, else it won't run.
# + [markdown] slideshow={"slide_type": "slide"}
# ## The Economic Credit Loss model
# This jupyter notebook is inspired by IMF working paper (WP/20/111) The Expected Credit Loss Modeling from a Top-Down Stress Testing Perspective by <NAME>, <NAME>, <NAME>, <NAME>. The working paper and the associated material is located https://www.imf.org/en/Publications/WP/Issues/2020/07/03/Expected-Credit-Loss-Modeling-from-a-Top-Down-Stress-Testing-Perspective-49545
#
# from the abstract of the paper:
# > The objective of this paper is to present an integrated tool suite for IFRS 9- and CECL compatible
# estimation in top-down solvency stress tests. The tool suite serves as an
# illustration for institutions wishing to include accounting-based approaches for credit risk
# modeling in top-down stress tests.
#
# This is a jupyter notebook build with the purpose of illustrating the conversion of a model from Latex to ModelFlow. The purpose is testing so take very much care.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Import libraries
# + slideshow={"slide_type": "fragment"}
# %load_ext autoreload
# %autoreload 2
import pandas as pd
from IPython.core.display import HTML,Markdown,Latex
from modelclass import model
import modeljupytermagic
# some useful stuf
model.widescreen()
pd.set_option('display.max_rows', None, 'display.max_columns', 10, 'precision', 2)
sortdf = lambda df: df[sorted([c for c in df.columns])]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Fetch the precooked model and baseline
# The model and data has been specified in another Jupyter notebook:
#
# [file/Model for economic credit loss with Z score transition nynamic - setup]
# + slideshow={"slide_type": "fragment"}
ecl,baseline = model.modelload('ecl.pcim')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Input values for the adverse scenario
# + slideshow={"slide_type": "subslide"}
# %%dataframe projection_adverse nshow
LOAN_GROWTH M_S1 M_S2 WRO_S3 Z
2021 1.00e-02 0.04 0.03 0.06 0.00
2022 -1.00e-02 0.04 0.03 0.06 -0.65
2023 -8.00e-03 0.04 0.03 0.06 -0.84
2024 -6.00e-03 0.04 0.03 0.06 -0.99
2025 -4.00e-03 0.04 0.03 0.06 -0.69
2026 -2.00e-03 0.04 0.03 0.06 -0.39
2027 -0.00e+00 0.04 0.03 0.06 -0.24
# + [markdown] slideshow={"slide_type": "slide"}
# ## Create an adverse Dataframe
# + slideshow={"slide_type": "fragment"}
adverse = baseline.copy()
adverse.update(projection_adverse)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Run Model
# For the Baseline and the adverse scenario
# + slideshow={"slide_type": "fragment"}
base_result = ecl(baseline,'2022','2026',keep='Baseline',silent=1)
adverse_result = ecl(adverse,keep = 'Adverse')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Find the Z consistent with an increase in Probability of default
# Here the .invert (goal seeking) method. It can be used for multible goals and targets, but here only for one of each
# + slideshow={"slide_type": "fragment"}
targets = adverse.loc['2022':,['PD_PIT']]+0.005 # set the target (goal)
instruments = ['Z'] # Set the instrumt
res = ecl.invert(baseline,targets = targets, instruments = instruments) # Now run
ecl.keep_solutions['PD_PIT target'] = res
# + slideshow={"slide_type": "fragment"}
import datetime
ecl.smpl('2021','2026')
figs = ecl.keep_plot('z loan_S3',showtype='level',legend=False,
vline=[('2022',' Projections')]);
# -
figs = ecl.keep_plot('tr_*_S3',showtype='level',legend=False);
| Examples/Economic Credit Loss/ECL Z - goal seeking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1. Setup and Import libraries
# django path
mysite_path = "C:\\Data\\UCL\\@MSc Project\\DB\\mysite\\"
# +
# standard packages
import os
import sys
import re
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from datetime import datetime
import django
from set_django_db import set_django_db
from asgiref.sync import sync_to_async
from IPython.core.display import HTML
# %matplotlib inline
# +
# set django models
set_django_db(mysite_path)
from tables_daniel.models import Company, Review
# specifically for Jupyter notebooks
os.environ["DJANGO_ALLOW_ASYNC_UNSAFE"] = "true"
# -
# center plots
HTML("""
<style>
.output_png {
display: table-cell;
text-align: center;
vertical_align: middle;
}
</style>
""")
# ## 2. Load, merge and filter the datasets
#
# <hr>
#
# **Content**
#
# 2.1 Load companies
# 2.2 Load reviews
# 2.3 Some useful merges/adds
# 2.4 Filter the data from the monitored period between 2018-07-01 and 2020-06-30
# 2.5 Filter only the reviews for the companies with at least 10 reviews
# 2.6 Add a column concatenating pros & cons
# 2.7 Update employee relatioship
# ### 2.1 Companies
# +
companies = pd.DataFrame(
list(
Company
.objects
.values('id', 'Company', 'Symbol', 'Sector', 'ListedOn')
.all()
)
)
companies_id = list(companies.id)
# -
# ### 2.2 Reviews
# +
reviews = list(
Review
.objects
.values(
'id', 'Company_id', 'ReviewTitle', 'Rating',
'JobTitle', 'EmployeeRelationship',
'Contract', 'Pros', 'Cons',
'Year', 'Month', 'Day'
)
.all()
.filter(Company_id = company_id) for company_id in companies_id
)
reviews_df = pd.DataFrame(
sum([list(reviews_i) for reviews_i in reviews],[])
).drop_duplicates()
# -
"""
for i in range(reviews_df.shape[0]):
row = dict(reviews_df.iloc[i,:])
review = (
Review
.objects
.values('id', 'JobTitle' ,'EmployeeRelationship')
.get(id=row['id'])
)
if review['JobTitle'] in ['Former Employee', 'Current Employee']:
new_jobTitle = review['EmployeeRelationship']
new_relationship = review['JobTitle']
(Review
.objects
.filter(id=row['id'])
.update(
JobTitle = new_jobTitle,
EmployeeRelationship = new_relationship
)
)
else:
pass
if (i+1)%100==0:
print(i+1)
"""
# ### 2.3 Some useful merges/adds
# +
# add sector and company name
reviews_df = reviews_df.merge(
companies[['id', 'Company', 'Sector', 'ListedOn']].rename(columns={'id': 'Company_id'}),
on='Company_id'
)
# add date column used for filtering
reviews_df['Date'] = reviews_df.apply(lambda x: '-'.join(
[str(x['Year']), str(x['Month']), str(x['Day'])]
), axis=1
)
reviews_df
# +
def string_to_date(date_str):
try:
return datetime.strptime(date_str, '%Y-%m-%d')
except:
return datetime.strptime('1800-1-1', '%Y-%m-%d')
def string_to_YM(date_str):
try:
return datetime.strptime(date_str, '%Y-%m')
except:
return datetime.strptime('1800-1-1', '%Y-%m-%d')
reviews_df['Date'] = reviews_df['Date'].apply(lambda x: string_to_date(x))
reviews_df['Year-Month'] = reviews_df.apply(lambda x: string_to_YM('-'.join([str(x['Year']), str(x['Month'])])), axis=1)
# -
# ### 2.4 Filter the data from the monitored period between
# +
# further analysis focusing only on the companies with at least 10 reviews in the monitored period
min_date = datetime.strptime('2018-7-1', '%Y-%m-%d')
max_date = datetime.strptime('2020-6-30', '%Y-%m-%d')
reviews_df = pd.DataFrame(
reviews_df[(reviews_df.Date >= min_date) & (reviews_df.Date <= max_date)]
)
reviews_df
# -
# ### 2.5 Filter only the reviews for companies with at least 10 reviews
# +
# count reviews
reviews_count = (
reviews_df
.groupby('Company')
.Rating
.count()
)
# filter companies
companies_filtered = list(reviews_count[reviews_count>10].index)
# +
reviews_df = reviews_df[reviews_df.Company.isin(companies_filtered)]
print(
f"There are {reviews_df.shape[0]:.0f} reviews in total."
)
# -
# ### 2.6 Add a column of concatenated pros & cons + their length
# +
reviews_df['Review'] = reviews_df['Pros'] + ' ' + reviews_df['Cons']
reviews_df['ReviewLength'] = reviews_df['Review'].apply(lambda x: len(x))
# -
reviews_df.head()
# ### 2.7 Update employee relationship
# +
def update_EmployeeRelationship(x):
if x not in ['Current Employee', 'Former Employee']:
return 'Not specified'
else:
return x
reviews_df['EmployeeRelationship'] = [update_EmployeeRelationship(reviews_df.loc[row, 'EmployeeRelationship']) for row in reviews_df.index]
# -
# ### 2.8 Add columns with total reviews to companies_df
# 1. get number of reviews per company
reviewsPerCompany = (
reviews_df
.groupby('Company')
.Rating
.count()
)
# +
# 1. get number of reviews per company
reviewsPerCompany = (
reviews_df
.groupby('Company')
.Rating
.count()
)
# 2. assign these values to company_df
companies_filtered_df = companies[companies.Company.isin(companies_filtered)]
companies_filtered_df['TotalReviews'] = 0
companies_filtered_df['TotalReviews'] = [reviewsPerCompany.loc[company] for company in companies_filtered_df.Company]
# -
## save the filtered companies as Company name, symbol
companies_filtered_df[['Company', 'Symbol', 'ListedOn']].to_excel('C:\\Data\\UCL\\companies_bonds.xlsx')
# ## 3. Summary statistics
#
# <hr>
#
# **Content**
#
# 3.1 Distribution of ratings/reviews over former/current and part/full-time employees
# 3.2 Mean, median, standard deviation and quantiles of total reviews per company
# 3.3 Mean, meadin, std and quantiles per sectors
# +
# helper quantile/quartile functions
def q1(x):
return x.quantile(.25)
def q3(x):
return x.quantile(.75)
def q10(x):
return x.quantile(.1)
def q90(x):
return x.quantile(.9)
# +
# Overall rating stats
print(
reviews_df
.Rating
.agg(['mean', 'std', q1, 'median', q3])
)
# +
# Overall reviews stats
print(
reviews_df
.ReviewLength
.agg(['mean', 'std', q1, 'median', q3])
)
# -
# ### 3.1 Distribution of ratings/reviews over former/current and part/full-time employees
print(
reviews_df
.groupby('Contract')
.Rating
.agg(['count', 'mean', 'std', q1, 'median', q3])
)
print(
reviews_df
.groupby('EmployeeRelationship')
.Rating
.agg(['count', 'mean', 'std', q1, 'median', q3])
)
print(
reviews_df
.groupby('Contract')
.ReviewLength
.agg(['count', 'mean', 'std', q1, 'median', q3])
)
print(
reviews_df
.groupby('EmployeeRelationship')
.ReviewLength
.agg(['count', 'mean', 'std', q1, 'median', q3])
)
# ### 3.2 Mean, median, standard deviation and quantiles of total reviews per company
print(
reviews_df
.groupby('Company')
.Rating
.count()
.agg(['mean', 'std', q10, q1, 'median', q3, q90, 'max'])
)
# **Global**
# +
file_path = r'C:\Data\UCL\@MSc Project - Data and sources\Images\histogram01.png'
plt.figure(figsize=(7,5))
ax = (reviews_df
.groupby('Company')
.Rating
.count()
.plot
.hist()
)
# set label, title, font size etc.
ax.set_xlabel('Number of reviews', fontsize=12)
ax.set_ylabel('Frequency', fontsize=12)
ax.set_title('Number of reviews per company', fontsize=14)
ax.set_xlim((0,companies_filtered_df.TotalReviews.max()))
plt.tight_layout()
plt.savefig(fname=file_path, dpi=300)
# -
# **Focused on bottom 90th companies**
# +
file_path = r'C:\Data\UCL\@MSc Project - Data and sources\Images\histogram01_zoomed.png'
bins = list(range(0, 1500, 100))
bins.append(companies_filtered_df.TotalReviews.max())
plt.figure(figsize=(7,5))
ax = (reviews_df
.groupby('Company')
.Rating
.count()
.plot
.hist(bins=bins)
)
# set label, title, font size etc.
ax.set_xlabel('Number of reviews', fontsize=12)
ax.set_ylabel('Frequency', fontsize=12)
ax.set_title('Number of reviews per company (zoom on the bottom 90th)', fontsize=14)
ax.set_xlim((0, 1400))
plt.tight_layout()
plt.savefig(fname=file_path, dpi=300)
# -
# **Combo**
# +
# setup
filepath = r'C:\Data\UCL\@MSc Project - Data and sources\Images\histogram01_combo.png'
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2, figsize=(14,5))
# global
ax1=(reviews_df
.groupby('Company')
.Rating
.count()
.hist(ax=ax1, grid=False)
)
ax1.set_xlabel('Number of reviews', fontsize=12)
ax1.set_ylabel('Frequency', fontsize=12)
ax1.set_title('Number of reviews per company', fontsize=14)
ax1.set_xlim((0,companies_filtered_df.TotalReviews.max()))
# zoom
bins = list(range(0, 1500, 100))
bins.append(companies_filtered_df.TotalReviews.max())
ax2=(reviews_df
.groupby('Company')
.Rating
.count()
.hist(ax=ax2, bins=bins,grid=False)
)
ax2.set_xlabel('Number of reviews', fontsize=12)
ax2.set_ylabel('Frequency', fontsize=12)
ax2.set_title('Number of reviews per company (zoom on the bottom 90th)', fontsize=14)
ax2.set_xlim((0, 1400))
# tight_layout and save
plt.tight_layout()
plt.savefig(fname=filepath, dpi=300)
# -
# ### 3.3 Mean, meadin, std and quantiles per market and sector
# #### 3.3.1 Review length
print(
companies_filtered_df
.TotalReviews
.agg(['mean', 'std', q10, q1, 'median', q3, q90, 'max'])
)
# +
filepath = r'C:/Data/UCL/@MSc Project - Data and sources/Exploration csv/StockMarket_hist.csv'
(companies_filtered_df
.groupby('ListedOn')
.TotalReviews
.agg(['count', 'mean', 'std', q10, q1, 'median', q3, q90, 'max'])
.round(2)
).to_csv(filepath)
print(
companies_filtered_df
.groupby('ListedOn')
.TotalReviews
.agg(['count', 'sum', 'mean', 'std', q10, q1, 'median', q3, q90, 'max'])
.round(2)
)
# +
filepath = r'C:/Data/UCL/@MSc Project - Data and sources/Exploration csv/Sector_hist.csv'
(companies_filtered_df
.groupby('Sector')
.TotalReviews
.agg(['count', 'mean', 'std', q10, q1, 'median', q3, q90, 'max'])
.round(2)
).to_csv(filepath)
print(
companies_filtered_df
.groupby('Sector')
.TotalReviews
.agg(['count', 'sum', 'mean', 'std', q10, q1, 'median', q3, q90, 'max'])
.round(2)
)
# +
filepath = r'C:/Data/UCL/@MSc Project - Data and sources/Exploration csv/Reviews_IndexSector.xlsx'
(companies_filtered_df
.groupby(['ListedOn', 'Sector'])
.TotalReviews
.agg(['mean', 'std', q10, q1, 'median', q3, q90, 'max'])
.round(2)
).to_excel(filepath)
print(
companies_filtered_df
.groupby(['ListedOn', 'Sector'])
.TotalReviews
.agg(['sum', 'mean', 'std', q10, q1, 'median', q3, q90, 'max'])
.round(2)
)
# -
(reviews_df
.groupby('ListedOn')
.ReviewLength
.agg(['mean', 'std', q1, 'median', q3])
)
(reviews_df
.groupby('Sector')
.ReviewLength
.agg(['mean', 'std', q1, 'median', q3])
)
# #### 3.3.2 Ratings
# +
filepath = r'C:/Data/UCL/@MSc Project - Data and sources/Exploration csv/Ratings - descriptive stats.csv'
(reviews_df
.groupby('ListedOn')
.Rating
.agg(['mean', 'std', q1, 'median', q3])
.round(2)
).to_csv(filepath)
print(
reviews_df
.groupby('Sector')
.Rating
.agg(['mean', 'std', q1, 'median', q3])
.round(2)
)
# -
print(
reviews_df
.groupby('ListedOn')
.Rating
.agg(['mean', 'std', q1, 'median', q3])
)
# +
filepath = r'C:/Data/UCL/@MSc Project - Data and sources/Exploration csv/RatingsIndexSector.xlsx'
(reviews_df
.groupby(['ListedOn', 'Sector'])
.Rating
.agg(['mean', 'std', q1, 'median', q3])
.round(2)
).to_excel(filepath)
print(
reviews_df
.groupby(['ListedOn', 'Sector'])
.Rating
.agg(['mean', 'std', q1, 'median', q3])
.round(2)
)
# -
# ### 3.4 Monthly mean rating per sector (+ 3M moving average)
# +
reviews_MonthSector = pd.DataFrame(
reviews_df
.groupby(['Sector', 'Year-Month'])
.agg(['count', 'mean'])
)
reviews_MonthSector = pd.DataFrame(
reviews_MonthSector.to_records()
)[['Sector', 'Year-Month', "('Rating', 'mean')", "('Rating', 'count')"]]
reviews_MonthSector.columns = ['Sector', 'Year-Month', 'Rating', 'Count']
reviews_MonthSector.head()
# +
sectors = reviews_MonthSector.Sector.unique()
# add 3-month rating average to the DF
reviews_MonthSector['3M_Average']=0
i=0
for sector in sectors:
avg = reviews_MonthSector[reviews_MonthSector.Sector==sector].Rating.rolling(window=3).mean()
start = i
end = i+avg.shape[0]
reviews_MonthSector.iloc[start:end, -1] = avg
i+=avg.shape[0]
# -
reviews_MonthSector
# ### 3.5 Weighted 3M average
# +
s = '1| 3 | 5|7'
[int(x.strip()) for x in s.split('|')]
# -
| Exploration/Summary Statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 4
#
# Before working on this assignment please read these instructions fully. In the submission area, you will notice that you can click the link to **Preview the Grading** for each step of the assignment. This is the criteria that will be used for peer grading. Please familiarize yourself with the criteria before beginning the assignment.
#
# This assignment requires that you to find **at least** two datasets on the web which are related, and that you visualize these datasets to answer a question with the broad topic of **sports or athletics** (see below) for the region of **Fort Lauderdale, Florida, United States**, or **United States** more broadly.
#
# You can merge these datasets with data from different regions if you like! For instance, you might want to compare **Fort Lauderdale, Florida, United States** to Ann Arbor, USA. In that case at least one source file must be about **Fort Lauderdale, Florida, United States**.
#
# You are welcome to choose datasets at your discretion, but keep in mind **they will be shared with your peers**, so choose appropriate datasets. Sensitive, confidential, illicit, and proprietary materials are not good choices for datasets for this assignment. You are welcome to upload datasets of your own as well, and link to them using a third party repository such as github, bitbucket, pastebin, etc. Please be aware of the Coursera terms of service with respect to intellectual property.
#
# Also, you are welcome to preserve data in its original language, but for the purposes of grading you should provide english translations. You are welcome to provide multiple visuals in different languages if you would like!
#
# As this assignment is for the whole course, you must incorporate principles discussed in the first week, such as having as high data-ink ratio (Tufte) and aligning with Cairo’s principles of truth, beauty, function, and insight.
#
# Here are the assignment instructions:
#
# * State the region and the domain category that your data sets are about (e.g., **Fort Lauderdale, Florida, United States** and **sports or athletics**).
# * You must state a question about the domain category and region that you identified as being interesting.
# * You must provide at least two links to available datasets. These could be links to files such as CSV or Excel files, or links to websites which might have data in tabular form, such as Wikipedia pages.
# * You must upload an image which addresses the research question you stated. In addition to addressing the question, this visual should follow Cairo's principles of truthfulness, functionality, beauty, and insightfulness.
# * You must contribute a short (1-2 paragraph) written justification of how your visualization addresses your stated research question.
#
# What do we mean by **sports or athletics**? For this category we are interested in sporting events or athletics broadly, please feel free to creatively interpret the category when building your research question!
#
# ## Tips
# * Wikipedia is an excellent source of data, and I strongly encourage you to explore it for new data sources.
# * Many governments run open data initiatives at the city, region, and country levels, and these are wonderful resources for localized data sources.
# * Several international agencies, such as the [United Nations](http://data.un.org/), the [World Bank](http://data.worldbank.org/), the [Global Open Data Index](http://index.okfn.org/place/) are other great places to look for data.
# * This assignment requires you to convert and clean datafiles. Check out the discussion forums for tips on how to do this from various sources, and share your successes with your fellow students!
#
# ## Example
# Looking for an example? Here's what our course assistant put together for the **Ann Arbor, MI, USA** area using **sports and athletics** as the topic. [Example Solution File](./readonly/Assignment4_example.pdf)
# ## What this is
# My assigned region and the domain category are Fort Lauderdale, Florida, United States and sports or athletics. I don't know any sports team in FLL and doubt that there would be any data publicly available if such team exists, so had to look for data from somewhere else. I had always foundd Gold State Warrior interesting, how a team of unknown players became a dominating even invincible force in the league, so I chose to analyze a controversial player, <NAME> and his role in the warrior danasty (2015~2018). A lots of NBA fans don't like Draymond Green's flamboyant personality and think he is overated and warrior's sucess have little to do with him.
#
# This assingment requirements would indicate a research on one dataset about bay area and another dataset of golden state warrior. I'm not really interested in how warriors are affectting bay area, traffic, skyrocketing ticket price... Instead I'm merging four data sets from four years and extracting Draymond Green and Warror as two datasets of interests
#
# I'm trying to answer a simple question, does warrior net points correlate with Draymond's minutes or points? Only the most obviously relevant metrics are looked at, namely DrayG's points, minuntes played and game outcome (by how many pts warriors won or lost the games he played). This is really just an homework exercise of basic data query, cleanning, processing and visuallization. It's by no means a complete evaluation on how much warriors need Draymond. A true money ball type of analysis would require a lot more sophisticated efforts.
#
# The regular seasons game stats data used are scrapped from https://www.basketball-reference.com/ with beautifulsoup, this part is not included in code cells since BS isn't available on course console.
#
#
#
#
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import re
# %matplotlib notebook
# ## Function for reading and cleaning data
# Game# is the nth Game Draymond Green played in the regular season, it's used to remove the game he didn't play due to injury,rest, ban, etc.
# Results is whether warriors won or lost the game (parsed to 'WorL') and by how many points (parsed to 'PointDiff').
# PointDiff is a good indicator how warrors did as a team.
def cleantable (filename):
df=pd.read_csv(filename,header=None,usecols=[0,6,8,26],names =['Game#','Result','MinutePlay','Points'])
df.dropna(subset = ['Game#'],inplace=True)
df['PointDiff']=df['Result'].apply(lambda x: int(re.search('\(([^)]+)', x).group(1)))
df['WorL']=df['Result'].apply(lambda x: x[0])
df['MinutePlay']=df['MinutePlay'].apply(lambda x: int(x.split(':')[0])+float(x.split(':')[1])/60)
df.drop(['Game#','Result'],axis=1,inplace=True)
return df
# ## Import and concatenate data
DrayG=pd.DataFrame()
for year in [2015,2016,2017,2018]:
filename='DraymondGreen'+str(year)+'.csv'
DrayG=DrayG.append(cleantable(filename))
# ## Draymond's Minutes vs Points
#
# A quick look susggests his points is somewhat "correlatable" to minutes on the court, other than the bottom section of data pts from games of low efficiency (high mins yet low score).
#
# Also noticable is that some of the best games warrior played (won by the largest margin, bright yellow points) are when Draymond played moderate mins and scored minimal points
#
DrayG.plot.scatter('MinutePlay', 'Points', c='PointDiff', colormap='viridis',title='Corr(Minutes,Points)={}'.format(DrayG['MinutePlay'].corr(DrayG['Points'])))
# ## Are Draymond's minutes or points correlated to how well warriors played? Not really.
pd.tools.plotting.scatter_matrix(DrayG);
# ## Zooming in, it' even clearer
sns.jointplot(DrayG['MinutePlay'], DrayG['PointDiff'],kind='kde', space=0);
sns.jointplot(DrayG['Points'], DrayG['PointDiff'], kind='kde', space=0);
# ## Looking at it from a different way, same thing
plt.figure()
plt.subplot(121)
sns.swarmplot('WorL', 'Points', data=DrayG);
plt.subplot(122)
sns.swarmplot('WorL', 'MinutePlay', data=DrayG);
| Assignment4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Qiskit Comparison
# This notebook highlights how my simulator is a nice little subset of qiskit's simulator.
# ## Qiskit Circuit
# To validate my implementation, first we create a simple circuit in qiskit and run simluation on it.
# +
from qiskit import QuantumCircuit
qc = QuantumCircuit(5, 5)
qc.h(0)
qc.h(1)
qc.cz(0, 4)
qc.unitary([[0.5 + 0.5j, 0.5 - 0.5j], [0.5 - 0.5j, 0.5 + 0.5j]], 1)
out = qc.measure([0, 1, 2, 3, 4], [0, 1, 2, 3, 4])
qc.draw()
# +
from qiskit import Aer, execute
import time
def simulate(circuit, num_shots):
backend_sim = Aer.get_backend('qasm_simulator')
job_sim = execute(circuit, backend_sim, shots=num_shots)
result_sim = job_sim.result()
return result_sim.get_counts(circuit)
start = time.time()
qiskit_counts = simulate(qc, num_shots=10000)
qiskit_diff = time.time() - start
# -
# ## JQiskit Circuit
# Here is the equivalent simulator in `jqiskit`. Notice how the API is pretty much the same, the main difference is that the `qc.measure()` function does both the simulation and state generation.
# +
from jqiskit.api import QuantumCircuit as JQuantumCircuit
qc = JQuantumCircuit(5)
qc.h(0)
qc.h(1)
qc.cz(0, 4)
qc.sqrtnot(1)
start = time.time()
jqiskit_counts, state = qc.measure(num_shots=10000)
jqiskit_diff = time.time() - start
# -
# ## Timing Comparison
#
# This clearly isn't a fair fight, as qiskit is probably doing a bunch of stuff under the hood for more complex features that I'm not doing for this simple simulator. But for what it's worth the simple version is significantly faster for the circuit described above. This may indicate a need for a more light-weight framework in the future for faster hyper parameter searches and stuff like that. For example, you can imagine a world where variational optimizations are first done on a faster, more ideal solver and then fine-tuned on a slower more accuate one.
print(f'jqiskit_speed: {jqiskit_diff * 1000:.2f} ms')
print(f' qiskit_speed: {qiskit_diff * 1000:.2f} ms')
# ## Validation
#
# I then compared the two by looking at the simulation output states. Note how qiskit is the opposite endian-ness as my implementation, so I had to flip the state to do a proper comparison.
# +
assert len(jqiskit_counts) == len(qiskit_counts), "Number of states don't match!"
for state_str in jqiskit_counts:
print(f'state: {state_str}, qiskit: {qiskit_counts[state_str[::-1]]}; jqiskit: {jqiskit_counts[state_str]}')
| notebooks/QiskitComparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [stats]
# language: python
# name: stats
# ---
# ### Working on tumour data
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
data = pd.read_csv('data.csv')
data.head()
data.describe()
data.shape
data.corr
plt.hist(data['smoothness_mean'])
f,(ax_box, ax_hist) = plt.subplots(2, gridspec_kw={'height_ratios':(.15, .85)})
sns.boxplot(data['texture_worst'], ax=ax_box)
sns.distplot(data['texture_worst'], ax=ax_hist)
sns.countplot(data['diagnosis'])
plt.show()
sns.pairplot(data=data, hue='diagnosis', vars=['radius_mean', 'texture_mean', 'perimeter_mean', 'area_mean', 'smoothness_mean'])
plt.show()
plt.figure(figsize=(20, 10))
sns.heatmap(data.corr(), annot=False) # annot is for values over the map
plt.show()
y = data['diagnosis']
y
data['diagnosis'].value_counts()
| Session_3_Tumour_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="MpkYHwCqk7W-"
# # <div align="center">**`dm_control` tutorial**</div>
# # <div align="center">[](https://colab.research.google.com/github/deepmind/dm_control/blob/master/tutorial.ipynb)</div>
#
#
#
#
#
#
#
# + [markdown] id="_UbO9uhtBSX5"
# > <p><small><small>Copyright 2020 The dm_control Authors.</small></p>
# > <p><small><small>Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at <a href="http://www.apache.org/licenses/LICENSE-2.0">http://www.apache.org/licenses/LICENSE-2.0</a>.</small></small></p>
# > <p><small><small>Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.</small></small></p>
# + [markdown] id="aThGKGp0cD76"
# This notebook provides an overview tutorial of DeepMind's `dm_control` package, hosted at the [deepmind/dm_control](https://github.com/deepmind/dm_control) repository on GitHub.
#
# It is adjunct to this [tech report](http://arxiv.org/abs/2006.12983).
#
# **A Colab runtime with GPU acceleration is required.** If you're using a CPU-only runtime, you can switch using the menu "Runtime > Change runtime type".
# + [markdown] id="YkBQUjm6gbGF"
# <!-- Internal installation instructions. -->
# + [markdown] id="YvyGCsgSCxHQ"
# ### Installing `dm_control` on Colab
# + id="IbZxYDxzoz5R"
#@title Run to install MuJoCo and `dm_control`
import distutils.util
import subprocess
if subprocess.run('nvidia-smi').returncode:
raise RuntimeError(
'Cannot communicate with GPU. '
'Make sure you are using a GPU Colab runtime. '
'Go to the Runtime menu and select Choose runtime type.')
mujoco_dir = "$HOME/.mujoco"
print('Installing OpenGL dependencies...')
# !apt-get update -qq
# !apt-get install -qq -y --no-install-recommends libglew2.0 > /dev/null
print('Downloading MuJoCo...')
MUJOCO_VERSION = 210
MUJOCO_ARCHIVE = (
f'mujoco{MUJOCO_VERSION}-{distutils.util.get_platform()}.tar.gz')
# !wget -q "https://mujoco.org/download/{MUJOCO_ARCHIVE}"
# !wget -q "https://mujoco.org/download/{MUJOCO_ARCHIVE}.sha256"
# check_result = !shasum -c "{MUJOCO_ARCHIVE}.sha256"
if _exit_code:
raise RuntimeError(
'Downloaded MuJoCo archive is corrupted (checksum mismatch)')
print('Unpacking MuJoCo...')
MUJOCO_DIR = '$HOME/.mujoco'
# !mkdir -p "{MUJOCO_DIR}"
# !tar -zxf {MUJOCO_ARCHIVE} -C "{MUJOCO_DIR}"
# Configure dm_control to use the EGL rendering backend (requires GPU)
# %env MUJOCO_GL=egl
print('Installing dm_control...')
# !pip install -q dm_control
print('Checking that the dm_control installation succeeded...')
try:
from dm_control import suite
env = suite.load('cartpole', 'swingup')
pixels = env.physics.render()
except Exception as e:
raise e from RuntimeError(
'Something went wrong during installation. Check the shell output above '
'for more information.\n'
'If using a hosted Colab runtime, make sure you enable GPU acceleration '
'by going to the Runtime menu and selecting "Choose runtime type".')
else:
del suite, env, pixels
# + [markdown] id="wtDN43hIJh2C"
# # Imports
#
# Run both of these cells:
# + cellView="form" id="T5f4w3Kq2X14"
#@title All `dm_control` imports required for this tutorial
# The basic mujoco wrapper.
from dm_control import mujoco
# Access to enums and MuJoCo library functions.
from dm_control.mujoco.wrapper.mjbindings import enums
from dm_control.mujoco.wrapper.mjbindings import mjlib
# PyMJCF
from dm_control import mjcf
# Composer high level imports
from dm_control import composer
from dm_control.composer.observation import observable
from dm_control.composer import variation
# Imports for Composer tutorial example
from dm_control.composer.variation import distributions
from dm_control.composer.variation import noises
from dm_control.locomotion.arenas import floors
# Control Suite
from dm_control import suite
# Run through corridor example
from dm_control.locomotion.walkers import cmu_humanoid
from dm_control.locomotion.arenas import corridors as corridor_arenas
from dm_control.locomotion.tasks import corridors as corridor_tasks
# Soccer
from dm_control.locomotion import soccer
# Manipulation
from dm_control import manipulation
# + id="gKc1FNhKiVJX"
#@title Other imports and helper functions
# General
import copy
import os
import itertools
from IPython.display import clear_output
import numpy as np
# Graphics-related
import matplotlib
import matplotlib.animation as animation
import matplotlib.pyplot as plt
from IPython.display import HTML
import PIL.Image
# Internal loading of video libraries.
# Use svg backend for figure rendering
# %config InlineBackend.figure_format = 'svg'
# Font sizes
SMALL_SIZE = 8
MEDIUM_SIZE = 10
BIGGER_SIZE = 12
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
# Inline video helper function
if os.environ.get('COLAB_NOTEBOOK_TEST', False):
# We skip video generation during tests, as it is quite expensive.
display_video = lambda *args, **kwargs: None
else:
def display_video(frames, framerate=30):
height, width, _ = frames[0].shape
dpi = 70
orig_backend = matplotlib.get_backend()
matplotlib.use('Agg') # Switch to headless 'Agg' to inhibit figure rendering.
fig, ax = plt.subplots(1, 1, figsize=(width / dpi, height / dpi), dpi=dpi)
matplotlib.use(orig_backend) # Switch back to the original backend.
ax.set_axis_off()
ax.set_aspect('equal')
ax.set_position([0, 0, 1, 1])
im = ax.imshow(frames[0])
def update(frame):
im.set_data(frame)
return [im]
interval = 1000/framerate
anim = animation.FuncAnimation(fig=fig, func=update, frames=frames,
interval=interval, blit=True, repeat=False)
return HTML(anim.to_html5_video())
# Seed numpy's global RNG so that cell outputs are deterministic. We also try to
# use RandomState instances that are local to a single cell wherever possible.
np.random.seed(42)
# + [markdown] id="jZXz9rPYGA-Y"
# # Model definition, compilation and rendering
#
#
# + [markdown] id="MRBaZsf1d7Gb"
# We begin by describing some basic concepts of the [MuJoCo](http://mujoco.org/) physics simulation library, but recommend the [official documentation](http://mujoco.org/book/) for details.
#
# Let's define a simple model with two geoms and a light.
# + id="ZS2utl59ZTsr"
#@title A static model {vertical-output: true}
static_model = """
<mujoco>
<worldbody>
<light name="top" pos="0 0 1"/>
<geom name="red_box" type="box" size=".2 .2 .2" rgba="1 0 0 1"/>
<geom name="green_sphere" pos=".2 .2 .2" size=".1" rgba="0 1 0 1"/>
</worldbody>
</mujoco>
"""
physics = mujoco.Physics.from_xml_string(static_model)
pixels = physics.render()
PIL.Image.fromarray(pixels)
# + [markdown] id="p4vPllljTJh8"
# `static_model` is written in MuJoCo's XML-based [MJCF](http://www.mujoco.org/book/modeling.html) modeling language. The `from_xml_string()` method invokes the model compiler, which instantiates the library's internal data structures. These can be accessed via the `physics` object, see below.
# + [markdown] id="MdUF2UYmR4TA"
# ## Adding DOFs and simulating, advanced rendering
# This is a perfectly legitimate model, but if we simulate it, nothing will happen except for time advancing. This is because this model has no degrees of freedom (DOFs). We add DOFs by adding **joints** to bodies, specifying how they can move with respect to their parents. Let us add a hinge joint and re-render, visualizing the joint axis.
# + id="R7zokzd_yeEg"
#@title A child body with a joint { vertical-output: true }
swinging_body = """
<mujoco>
<worldbody>
<light name="top" pos="0 0 1"/>
<body name="box_and_sphere" euler="0 0 -30">
<joint name="swing" type="hinge" axis="1 -1 0" pos="-.2 -.2 -.2"/>
<geom name="red_box" type="box" size=".2 .2 .2" rgba="1 0 0 1"/>
<geom name="green_sphere" pos=".2 .2 .2" size=".1" rgba="0 1 0 1"/>
</body>
</worldbody>
</mujoco>
"""
physics = mujoco.Physics.from_xml_string(swinging_body)
# Visualize the joint axis.
scene_option = mujoco.wrapper.core.MjvOption()
scene_option.flags[enums.mjtVisFlag.mjVIS_JOINT] = True
pixels = physics.render(scene_option=scene_option)
PIL.Image.fromarray(pixels)
# + [markdown] id="INOGGV0PTQus"
# The things that move (and which have inertia) are called *bodies*. The body's child `joint` specifies how that body can move with respect to its parent, in this case `box_and_sphere` w.r.t the `worldbody`.
#
# Note that the body's frame is **rotated** with an `euler` directive, and its children, the geoms and the joint, rotate with it. This is to emphasize the local-to-parent-frame nature of position and orientation directives in MJCF.
#
# Let's make a video, to get a sense of the dynamics and to see the body swinging under gravity.
# + id="Z_57VMUDpGrj"
#@title Making a video {vertical-output: true}
duration = 2 # (seconds)
framerate = 30 # (Hz)
# Visualize the joint axis
scene_option = mujoco.wrapper.core.MjvOption()
scene_option.flags[enums.mjtVisFlag.mjVIS_JOINT] = True
# Simulate and display video.
frames = []
physics.reset() # Reset state and time
while physics.data.time < duration:
physics.step()
if len(frames) < physics.data.time * framerate:
pixels = physics.render(scene_option=scene_option)
frames.append(pixels)
display_video(frames, framerate)
# + [markdown] id="yYvS1UaciMX_"
# Note how we collect the video frames. Because physics simulation timesteps are generally much smaller than framerates (the default timestep is 2ms), we don't render after each step.
# + [markdown] id="nQ8XOnRQx7T1"
# ## Rendering options
#
# Like joint visualisation, additional rendering options are exposed as parameters to the `render` method.
# + id="AQITZiIgx7T2"
#@title Enable transparency and frame visualization {vertical-output: true}
scene_option = mujoco.wrapper.core.MjvOption()
scene_option.frame = enums.mjtFrame.mjFRAME_GEOM
scene_option.flags[enums.mjtVisFlag.mjVIS_TRANSPARENT] = True
pixels = physics.render(scene_option=scene_option)
PIL.Image.fromarray(pixels)
# + id="PDDgY48vx7T6"
#@title Depth rendering {vertical-output: true}
# depth is a float array, in meters.
depth = physics.render(depth=True)
# Shift nearest values to the origin.
depth -= depth.min()
# Scale by 2 mean distances of near rays.
depth /= 2*depth[depth <= 1].mean()
# Scale to [0, 255]
pixels = 255*np.clip(depth, 0, 1)
PIL.Image.fromarray(pixels.astype(np.uint8))
# + id="PNwiIrgpx7T8"
#@title Segmentation rendering {vertical-output: true}
seg = physics.render(segmentation=True)
# Display the contents of the first channel, which contains object
# IDs. The second channel, seg[:, :, 1], contains object types.
geom_ids = seg[:, :, 0]
# Infinity is mapped to -1
geom_ids = geom_ids.astype(np.float64) + 1
# Scale to [0, 1]
geom_ids = geom_ids / geom_ids.max()
pixels = 255*geom_ids
PIL.Image.fromarray(pixels.astype(np.uint8))
# + id="uCJQlv3cQcJQ"
#@title Projecting from world to camera coordinates {vertical-output: true}
# Get the world coordinates of the box corners
box_pos = physics.named.data.geom_xpos['red_box']
box_mat = physics.named.data.geom_xmat['red_box'].reshape(3, 3)
box_size = physics.named.model.geom_size['red_box']
offsets = np.array([-1, 1]) * box_size[:, None]
xyz_local = np.stack(itertools.product(*offsets)).T
xyz_global = box_pos[:, None] + box_mat @ xyz_local
# Camera matrices multiply homogenous [x, y, z, 1] vectors.
corners_homogeneous = np.ones((4, xyz_global.shape[1]), dtype=float)
corners_homogeneous[:3, :] = xyz_global
# Get the camera matrix.
camera = mujoco.Camera(physics)
camera_matrix = camera.matrix
# Project world coordinates into pixel space. See:
# https://en.wikipedia.org/wiki/3D_projection#Mathematical_formula
xs, ys, s = camera_matrix @ corners_homogeneous
# x and y are in the pixel coordinate system.
x = xs / s
y = ys / s
# Render the camera view and overlay the projected corner coordinates.
pixels = camera.render()
fig, ax = plt.subplots(1, 1)
ax.imshow(pixels)
ax.plot(x, y, '+', c='w')
ax.set_axis_off()
# + [markdown] id="gf9h_wi9weet"
# # MuJoCo basics and named indexing
# + [markdown] id="NCcZxrDDB1Cj"
# ## `mjModel`
# MuJoCo's `mjModel`, encapsulated in `physics.model`, contains the *model description*, including the default initial state and other fixed quantities which are not a function of the state, e.g. the positions of geoms in the frame of their parent body. The (x, y, z) offsets of the `box` and `sphere` geoms, relative their parent body `box_and_sphere` are given by `model.geom_pos`:
# + id="wx8NANvOF8g1"
physics.model.geom_pos
# + [markdown] id="v9O1VNdmHn_K"
# Docstrings of attributes provide short descriptions.
# + id="8_0MwaeYHn_N"
help(type(physics.model).geom_pos)
# + [markdown] id="Wee5ATLtIQn_"
# The `model.opt` structure contains global quantities like
# + id="BhzbZIfDIU2-"
print('timestep', physics.model.opt.timestep)
print('gravity', physics.model.opt.gravity)
# + [markdown] id="t5hY0fyXFLcf"
# ## `mjData`
# `mjData`, encapsulated in `physics.data`, contains the *state* and quantities that depend on it. The state is made up of time, generalized positions and generalised velocities. These are respectively `data.time`, `data.qpos` and `data.qvel`.
#
# Let's print the state of the swinging body where we left it:
# + id="acwZtDwp9mQU"
print(physics.data.time, physics.data.qpos, physics.data.qvel)
# + [markdown] id="7YlmcLcA-WQu"
# `physics.data` also contains functions of the state, for example the cartesian positions of objects in the world frame. The (x, y, z) positions of our two geoms are in `data.geom_xpos`:
# + id="CPwDcAQ0-uUE"
print(physics.data.geom_xpos)
# + [markdown] id="Z0UodCxS_v49"
# ## Named indexing
#
# The semantics of the above arrays are made clearer using the `named` wrapper, which assigns names to rows and type names to columns.
# + id="cLARcaK6-xCU"
print(physics.named.data.geom_xpos)
# + [markdown] id="wgXOUZNZHIx6"
# Note how `model.geom_pos` and `data.geom_xpos` have similar semantics but very different meanings.
# + id="-cW61ClRHS8a"
print(physics.named.model.geom_pos)
# + [markdown] id="-lQ0AChVASMv"
# Name strings can be used to index **into** the relevant quantities, making code much more readable and robust.
# + id="Rj4ad9fQAnFZ"
physics.named.data.geom_xpos['green_sphere', 'z']
# + [markdown] id="axr_p6APAzFn"
# Joint names can be used to index into quantities in configuration space (beginning with the letter `q`):
# + id="hluF9aDG9O1W"
physics.named.data.qpos['swing']
# + [markdown] id="3IhfyD2Q1pjv"
# We can mix NumPy slicing operations with named indexing. As an example, we can set the color of the box using its name (`"red_box"`) as an index into the rows of the `geom_rgba` array.
# + id="f5vVUullUvWH"
#@title Changing colors using named indexing{vertical-output: true}
random_rgb = np.random.rand(3)
physics.named.model.geom_rgba['red_box', :3] = random_rgb
pixels = physics.render()
PIL.Image.fromarray(pixels)
# + [markdown] id="elzPPdq-KhLI"
# Note that while `physics.model` quantities will not be changed by the engine, we can change them ourselves between steps. This however is generally not recommended, the preferred approach being to modify the model at the XML level using the PyMJCF library, see below.
# + [markdown] id="22ENjtVuhwsm"
# ## Setting the state with `reset_context()`
#
# In order for `data` quantities that are functions of the state to be in sync with the state, MuJoCo's `mj_step1()` needs to be called. This is facilitated by the `reset_context()` context, please see in-depth discussion in Section 2.1 of the [tech report](https://arxiv.org/abs/2006.12983).
# + id="WBPprCtWgXFN"
physics.named.data.qpos['swing'] = np.pi
print('Without reset_context, spatial positions are not updated:',
physics.named.data.geom_xpos['green_sphere', ['z']])
with physics.reset_context():
physics.named.data.qpos['swing'] = np.pi
print('After reset_context, positions are up-to-date:',
physics.named.data.geom_xpos['green_sphere', ['z']])
# + [markdown] id="SHppAOjvSupc"
# ## Free bodies: the self-inverting "tippe-top"
#
# A free body is a body with a `free` joint, with 6 movement DOFs: 3 translations and 3 rotations. We could give our `box_and_sphere` body a free joint and watch it fall, but let's look at something more interesting. A "tippe top" is a spinning toy which flips itself on its head ([Wikipedia](https://en.wikipedia.org/wiki/Tippe_top)). We model it as follows:
# + id="xasXQpVMjIwA"
#@title The "tippe-top" model{vertical-output: true}
tippe_top = """
<mujoco model="tippe top">
<option integrator="RK4"/>
<asset>
<texture name="grid" type="2d" builtin="checker" rgb1=".1 .2 .3"
rgb2=".2 .3 .4" width="300" height="300"/>
<material name="grid" texture="grid" texrepeat="8 8" reflectance=".2"/>
</asset>
<worldbody>
<geom size=".2 .2 .01" type="plane" material="grid"/>
<light pos="0 0 .6"/>
<camera name="closeup" pos="0 -.1 .07" xyaxes="1 0 0 0 1 2"/>
<body name="top" pos="0 0 .02">
<freejoint/>
<geom name="ball" type="sphere" size=".02" />
<geom name="stem" type="cylinder" pos="0 0 .02" size="0.004 .008"/>
<geom name="ballast" type="box" size=".023 .023 0.005" pos="0 0 -.015"
contype="0" conaffinity="0" group="3"/>
</body>
</worldbody>
<keyframe>
<key name="spinning" qpos="0 0 0.02 1 0 0 0" qvel="0 0 0 0 1 200" />
</keyframe>
</mujoco>
"""
physics = mujoco.Physics.from_xml_string(tippe_top)
PIL.Image.fromarray(physics.render(camera_id='closeup'))
# + [markdown] id="bvHlr6maJYIG"
# Note several new features of this model definition:
# 0. The free joint is added with the `<freejoint/>` clause, which is similar to `<joint type="free"/>`, but prohibits unphysical attributes like friction or stiffness.
# 1. We use the `<option/>` clause to set the integrator to the more accurate Runge Kutta 4th order.
# 2. We define the floor's grid material inside the `<asset/>` clause and reference it in the floor geom.
# 3. We use an invisible and non-colliding box geom called `ballast` to move the top's center-of-mass lower. Having a low center of mass is (counter-intuitively) required for the flipping behaviour to occur.
# 4. We save our initial spinning state as a keyframe. It has a high rotational velocity around the z-axis, but is not perfectly oriented with the world.
# 5. We define a `<camera>` in our model, and then render from it using the `camera_id` argument to `render()`.
# Let us examine the state:
# + id="o4S9nYhHOKmb"
print('positions', physics.data.qpos)
print('velocities', physics.data.qvel)
# + [markdown] id="71UgzBAqWdtZ"
# The velocities are easy to interpret, 6 zeros, one for each DOF. What about the length-7 positions? We can see the initial 2cm height of the body; the subsequent four numbers are the 3D orientation, defined by a *unit quaternion*. These normalized four-vectors, which preserve the topology of the orientation group, are the reason that `data.qpos` can be bigger than `data.qvel`: 3D orientations are represented with **4** numbers while angular velocities are **3** numbers.
# + id="5P4HkhKNGQvs"
#@title Video of the tippe-top {vertical-output: true}
#@test {"timeout": 600}
duration = 7 # (seconds)
framerate = 60 # (Hz)
# Simulate and display video.
frames = []
physics.reset(0) # Reset to keyframe 0 (load a saved state).
while physics.data.time < duration:
physics.step()
if len(frames) < (physics.data.time) * framerate:
pixels = physics.render(camera_id='closeup')
frames.append(pixels)
display_video(frames, framerate)
# + [markdown] id="rRuFKD2ubPgu"
# ### Measuring values from `physics.data`
# The `physics.data` structure contains all of the dynamic variables and intermediate results produced by the simulation. These are expected to change on each timestep.
#
# Below we simulate for 2000 timesteps and plot the state and height of the sphere as a function of time.
# + id="1XXB6asJoZ2N"
#@title Measuring values {vertical-output: true}
timevals = []
angular_velocity = []
stem_height = []
# Simulate and save data
physics.reset(0)
while physics.data.time < duration:
physics.step()
timevals.append(physics.data.time)
angular_velocity.append(physics.data.qvel[3:6].copy())
stem_height.append(physics.named.data.geom_xpos['stem', 'z'])
dpi = 100
width = 480
height = 640
figsize = (width / dpi, height / dpi)
_, ax = plt.subplots(2, 1, figsize=figsize, dpi=dpi, sharex=True)
ax[0].plot(timevals, angular_velocity)
ax[0].set_title('angular velocity')
ax[0].set_ylabel('radians / second')
ax[1].plot(timevals, stem_height)
ax[1].set_xlabel('time (seconds)')
ax[1].set_ylabel('meters')
_ = ax[1].set_title('stem height')
# + [markdown] id="UAMItwu8e1WR"
# # PyMJCF tutorial
#
#
#
# + [markdown] id="hPiY8m3MssKM"
# This library provides a Python object model for MuJoCo's XML-based
# [MJCF](http://www.mujoco.org/book/modeling.html) physics modeling language. The
# goal of the library is to allow users to easily interact with and modify MJCF
# models in Python, similarly to what the JavaScript DOM does for HTML.
#
# A key feature of this library is the ability to easily compose multiple separate
# MJCF models into a larger one. Disambiguation of duplicated names from different
# models, or multiple instances of the same model, is handled automatically.
#
# One typical use case is when we want robots with a variable number of joints. This is a fundamental change to the kinematics, requiring a new XML descriptor and new binary model to be compiled.
#
# The following snippets realise this scenario and provide a quick example of this library's use case.
# + id="gKny5EJ4uVzu"
class Leg(object):
"""A 2-DoF leg with position actuators."""
def __init__(self, length, rgba):
self.model = mjcf.RootElement()
# Defaults:
self.model.default.joint.damping = 2
self.model.default.joint.type = 'hinge'
self.model.default.geom.type = 'capsule'
self.model.default.geom.rgba = rgba # Continued below...
# Thigh:
self.thigh = self.model.worldbody.add('body')
self.hip = self.thigh.add('joint', axis=[0, 0, 1])
self.thigh.add('geom', fromto=[0, 0, 0, length, 0, 0], size=[length/4])
# Hip:
self.shin = self.thigh.add('body', pos=[length, 0, 0])
self.knee = self.shin.add('joint', axis=[0, 1, 0])
self.shin.add('geom', fromto=[0, 0, 0, 0, 0, -length], size=[length/5])
# Position actuators:
self.model.actuator.add('position', joint=self.hip, kp=10)
self.model.actuator.add('position', joint=self.knee, kp=10)
# + [markdown] id="cFJerI--UtTy"
# The `Leg` class describes an abstract articulated leg, with two joints and corresponding proportional-derivative actuators.
#
# Note that:
#
# - MJCF attributes correspond directly to arguments of the `add()` method.
# - When referencing elements, e.g when specifying the joint to which an actuator is attached, the MJCF element itself is used, rather than the name string.
# + cellView="both" id="SESlL_TidKHx"
BODY_RADIUS = 0.1
BODY_SIZE = (BODY_RADIUS, BODY_RADIUS, BODY_RADIUS / 2)
random_state = np.random.RandomState(42)
def make_creature(num_legs):
"""Constructs a creature with `num_legs` legs."""
rgba = random_state.uniform([0, 0, 0, 1], [1, 1, 1, 1])
model = mjcf.RootElement()
model.compiler.angle = 'radian' # Use radians.
# Make the torso geom.
model.worldbody.add(
'geom', name='torso', type='ellipsoid', size=BODY_SIZE, rgba=rgba)
# Attach legs to equidistant sites on the circumference.
for i in range(num_legs):
theta = 2 * i * np.pi / num_legs
hip_pos = BODY_RADIUS * np.array([np.cos(theta), np.sin(theta), 0])
hip_site = model.worldbody.add('site', pos=hip_pos, euler=[0, 0, theta])
leg = Leg(length=BODY_RADIUS, rgba=rgba)
hip_site.attach(leg.model)
return model
# + [markdown] id="elyuJiI3U3kM"
# The `make_creature` function uses PyMJCF's `attach()` method to procedurally attach legs to the torso. Note that at this stage both the torso and hip attachment sites are children of the `worldbody`, since their parent body has yet to be instantiated. We'll now make an arena with a chequered floor and two lights, and place our creatures in a grid.
# + id="F7_Tx9P9U_VJ"
#@title Six Creatures on a floor.{vertical-output: true}
arena = mjcf.RootElement()
chequered = arena.asset.add('texture', type='2d', builtin='checker', width=300,
height=300, rgb1=[.2, .3, .4], rgb2=[.3, .4, .5])
grid = arena.asset.add('material', name='grid', texture=chequered,
texrepeat=[5, 5], reflectance=.2)
arena.worldbody.add('geom', type='plane', size=[2, 2, .1], material=grid)
for x in [-2, 2]:
arena.worldbody.add('light', pos=[x, -1, 3], dir=[-x, 1, -2])
# Instantiate 6 creatures with 3 to 8 legs.
creatures = [make_creature(num_legs=num_legs) for num_legs in range(3, 9)]
# Place them on a grid in the arena.
height = .15
grid = 5 * BODY_RADIUS
xpos, ypos, zpos = np.meshgrid([-grid, 0, grid], [0, grid], [height])
for i, model in enumerate(creatures):
# Place spawn sites on a grid.
spawn_pos = (xpos.flat[i], ypos.flat[i], zpos.flat[i])
spawn_site = arena.worldbody.add('site', pos=spawn_pos, group=3)
# Attach to the arena at the spawn sites, with a free joint.
spawn_site.attach(model).add('freejoint')
# Instantiate the physics and render.
physics = mjcf.Physics.from_mjcf_model(arena)
PIL.Image.fromarray(physics.render())
# + [markdown] id="cMfDaD7PfuoI"
# Multi-legged creatures, ready to roam! Let's inject some controls and watch them move. We'll generate a sinusoidal open-loop control signal of fixed frequency and random phase, recording both video frames and the horizontal positions of the torso geoms, in order to plot the movement trajectories.
# + id="8Gx39DMEUZDt"
#@title Video of the movement{vertical-output: true}
#@test {"timeout": 600}
duration = 10 # (Seconds)
framerate = 30 # (Hz)
video = []
pos_x = []
pos_y = []
torsos = [] # List of torso geom elements.
actuators = [] # List of actuator elements.
for creature in creatures:
torsos.append(creature.find('geom', 'torso'))
actuators.extend(creature.find_all('actuator'))
# Control signal frequency, phase, amplitude.
freq = 5
phase = 2 * np.pi * random_state.rand(len(actuators))
amp = 0.9
# Simulate, saving video frames and torso locations.
physics.reset()
while physics.data.time < duration:
# Inject controls and step the physics.
physics.bind(actuators).ctrl = amp * np.sin(freq * physics.data.time + phase)
physics.step()
# Save torso horizontal positions using bind().
pos_x.append(physics.bind(torsos).xpos[:, 0].copy())
pos_y.append(physics.bind(torsos).xpos[:, 1].copy())
# Save video frames.
if len(video) < physics.data.time * framerate:
pixels = physics.render()
video.append(pixels.copy())
display_video(video, framerate)
# + id="u09JfenWYLZu"
#@title Movement trajectories{vertical-output: true}
creature_colors = physics.bind(torsos).rgba[:, :3]
fig, ax = plt.subplots(figsize=(4, 4))
ax.set_prop_cycle(color=creature_colors)
_ = ax.plot(pos_x, pos_y, linewidth=4)
# + [markdown] id="kggQyvNpf_Y9"
# The plot above shows the corresponding movement trajectories of creature positions. Note how `physics.bind(torsos)` was used to access both `xpos` and `rgba` values. Once the `Physics` had been instantiated by `from_mjcf_model()`, the `bind()` method will expose both the associated `mjData` and `mjModel` fields of an `mjcf` element, providing unified access to all quantities in the simulation.
# + [markdown] id="wcRX_wu_8q8u"
# # Composer tutorial
# + [markdown] id="1DMhNPE5tSdw"
# In this tutorial we will create a task requiring our "creature" above to press a colour-changing button on the floor with a prescribed force. We begin by implementing our creature as a `composer.Entity`:
# + id="WwfzIqgNuFKt"
#@title The `Creature` class
class Creature(composer.Entity):
"""A multi-legged creature derived from `composer.Entity`."""
def _build(self, num_legs):
self._model = make_creature(num_legs)
def _build_observables(self):
return CreatureObservables(self)
@property
def mjcf_model(self):
return self._model
@property
def actuators(self):
return tuple(self._model.find_all('actuator'))
# Add simple observable features for joint angles and velocities.
class CreatureObservables(composer.Observables):
@composer.observable
def joint_positions(self):
all_joints = self._entity.mjcf_model.find_all('joint')
return observable.MJCFFeature('qpos', all_joints)
@composer.observable
def joint_velocities(self):
all_joints = self._entity.mjcf_model.find_all('joint')
return observable.MJCFFeature('qvel', all_joints)
# + [markdown] id="CXZOBK6RkjxH"
# The `Creature` Entity includes generic Observables for joint angles and velocities. Because `find_all()` is called on the `Creature`'s MJCF model, it will only return the creature's leg joints, and not the "free" joint with which it will be attached to the world. Note that Composer Entities should override the `_build` and `_build_observables` methods rather than `__init__`. The implementation of `__init__` in the base class calls `_build` and `_build_observables`, in that order, to ensure that the entity's MJCF model is created before its observables. This was a design choice which allows the user to refer to an observable as an attribute (`entity.observables.foo`) while still making it clear which attributes are observables. The stateful `Button` class derives from `composer.Entity` and implements the `initialize_episode` and `after_substep` callbacks.
# + id="BE9VU2EOvR-u"
#@title The `Button` class
NUM_SUBSTEPS = 25 # The number of physics substeps per control timestep.
class Button(composer.Entity):
"""A button Entity which changes colour when pressed with certain force."""
def _build(self, target_force_range=(5, 10)):
self._min_force, self._max_force = target_force_range
self._mjcf_model = mjcf.RootElement()
self._geom = self._mjcf_model.worldbody.add(
'geom', type='cylinder', size=[0.25, 0.02], rgba=[1, 0, 0, 1])
self._site = self._mjcf_model.worldbody.add(
'site', type='cylinder', size=self._geom.size*1.01, rgba=[1, 0, 0, 0])
self._sensor = self._mjcf_model.sensor.add('touch', site=self._site)
self._num_activated_steps = 0
def _build_observables(self):
return ButtonObservables(self)
@property
def mjcf_model(self):
return self._mjcf_model
# Update the activation (and colour) if the desired force is applied.
def _update_activation(self, physics):
current_force = physics.bind(self.touch_sensor).sensordata[0]
self._is_activated = (current_force >= self._min_force and
current_force <= self._max_force)
physics.bind(self._geom).rgba = (
[0, 1, 0, 1] if self._is_activated else [1, 0, 0, 1])
self._num_activated_steps += int(self._is_activated)
def initialize_episode(self, physics, random_state):
self._reward = 0.0
self._num_activated_steps = 0
self._update_activation(physics)
def after_substep(self, physics, random_state):
self._update_activation(physics)
@property
def touch_sensor(self):
return self._sensor
@property
def num_activated_steps(self):
return self._num_activated_steps
class ButtonObservables(composer.Observables):
"""A touch sensor which averages contact force over physics substeps."""
@composer.observable
def touch_force(self):
return observable.MJCFFeature('sensordata', self._entity.touch_sensor,
buffer_size=NUM_SUBSTEPS, aggregator='mean')
# + [markdown] id="D9vB5nCwkyIW"
# Note how the Button counts the number of sub-steps during which it is pressed with the desired force. It also exposes an `Observable` of the force being applied to the button, whose value is an average of the readings over the physics time-steps.
#
# We import some `variation` modules and an arena factory:
# + id="aDTTQMHtVawM"
#@title Random initialiser using `composer.variation`
class UniformCircle(variation.Variation):
"""A uniformly sampled horizontal point on a circle of radius `distance`."""
def __init__(self, distance):
self._distance = distance
self._heading = distributions.Uniform(0, 2*np.pi)
def __call__(self, initial_value=None, current_value=None, random_state=None):
distance, heading = variation.evaluate(
(self._distance, self._heading), random_state=random_state)
return (distance*np.cos(heading), distance*np.sin(heading), 0)
# + id="dgZwP-pvxJdt"
#@title The `PressWithSpecificForce` task
class PressWithSpecificForce(composer.Task):
def __init__(self, creature):
self._creature = creature
self._arena = floors.Floor()
self._arena.add_free_entity(self._creature)
self._arena.mjcf_model.worldbody.add('light', pos=(0, 0, 4))
self._button = Button()
self._arena.attach(self._button)
# Configure initial poses
self._creature_initial_pose = (0, 0, 0.15)
button_distance = distributions.Uniform(0.5, .75)
self._button_initial_pose = UniformCircle(button_distance)
# Configure variators
self._mjcf_variator = variation.MJCFVariator()
self._physics_variator = variation.PhysicsVariator()
# Configure and enable observables
pos_corrptor = noises.Additive(distributions.Normal(scale=0.01))
self._creature.observables.joint_positions.corruptor = pos_corrptor
self._creature.observables.joint_positions.enabled = True
vel_corruptor = noises.Multiplicative(distributions.LogNormal(sigma=0.01))
self._creature.observables.joint_velocities.corruptor = vel_corruptor
self._creature.observables.joint_velocities.enabled = True
self._button.observables.touch_force.enabled = True
def to_button(physics):
button_pos, _ = self._button.get_pose(physics)
return self._creature.global_vector_to_local_frame(physics, button_pos)
self._task_observables = {}
self._task_observables['button_position'] = observable.Generic(to_button)
for obs in self._task_observables.values():
obs.enabled = True
self.control_timestep = NUM_SUBSTEPS * self.physics_timestep
@property
def root_entity(self):
return self._arena
@property
def task_observables(self):
return self._task_observables
def initialize_episode_mjcf(self, random_state):
self._mjcf_variator.apply_variations(random_state)
def initialize_episode(self, physics, random_state):
self._physics_variator.apply_variations(physics, random_state)
creature_pose, button_pose = variation.evaluate(
(self._creature_initial_pose, self._button_initial_pose),
random_state=random_state)
self._creature.set_pose(physics, position=creature_pose)
self._button.set_pose(physics, position=button_pose)
def get_reward(self, physics):
return self._button.num_activated_steps / NUM_SUBSTEPS
# + id="dRuuZdLpthbv"
#@title Instantiating an environment{vertical-output: true}
creature = Creature(num_legs=4)
task = PressWithSpecificForce(creature)
env = composer.Environment(task, random_state=np.random.RandomState(42))
env.reset()
PIL.Image.fromarray(env.physics.render())
# + [markdown] id="giTL_6euZFlw"
# # The *Control Suite*
# + [markdown] id="zfIcrDECtdB2"
# The **Control Suite** is a set of stable, well-tested tasks designed to serve as a benchmark for continuous control learning agents. Tasks are written using the basic MuJoCo wrapper interface. Standardised action, observation and reward structures make suite-wide benchmarking simple and learning curves easy to interpret. Control Suite domains are not meant to be modified, in order to facilitate benchmarking. For full details regarding benchmarking, please refer to our original [publication](https://arxiv.org/abs/1801.00690).
#
# A video of solved benchmark tasks is [available here](https://www.youtube.com/watch?v=rAai4QzcYbs&feature=youtu.be).
#
# The suite come with convenient module level tuples for iterating over tasks:
# + id="a_whTJG8uTp1"
#@title Iterating over tasks{vertical-output: true}
max_len = max(len(d) for d, _ in suite.BENCHMARKING)
for domain, task in suite.BENCHMARKING:
print(f'{domain:<{max_len}} {task}')
# + id="qN8y3etfZFly"
#@title Loading and simulating a `suite` task{vertical-output: true}
# Load the environment
random_state = np.random.RandomState(42)
env = suite.load('hopper', 'stand', task_kwargs={'random': random_state})
# Simulate episode with random actions
duration = 4 # Seconds
frames = []
ticks = []
rewards = []
observations = []
spec = env.action_spec()
time_step = env.reset()
while env.physics.data.time < duration:
action = random_state.uniform(spec.minimum, spec.maximum, spec.shape)
time_step = env.step(action)
camera0 = env.physics.render(camera_id=0, height=200, width=200)
camera1 = env.physics.render(camera_id=1, height=200, width=200)
frames.append(np.hstack((camera0, camera1)))
rewards.append(time_step.reward)
observations.append(copy.deepcopy(time_step.observation))
ticks.append(env.physics.data.time)
html_video = display_video(frames, framerate=1./env.control_timestep())
# Show video and plot reward and observations
num_sensors = len(time_step.observation)
_, ax = plt.subplots(1 + num_sensors, 1, sharex=True, figsize=(4, 8))
ax[0].plot(ticks, rewards)
ax[0].set_ylabel('reward')
ax[-1].set_xlabel('time')
for i, key in enumerate(time_step.observation):
data = np.asarray([observations[j][key] for j in range(len(observations))])
ax[i+1].plot(ticks, data, label=key)
ax[i+1].set_ylabel(key)
html_video
# + id="ggVbQr5hZFl5"
#@title Visualizing an initial state of one task per domain in the Control Suite
domains_tasks = {domain: task for domain, task in suite.ALL_TASKS}
random_state = np.random.RandomState(42)
num_domains = len(domains_tasks)
n_col = num_domains // int(np.sqrt(num_domains))
n_row = num_domains // n_col + int(0 < num_domains % n_col)
_, ax = plt.subplots(n_row, n_col, figsize=(12, 12))
for a in ax.flat:
a.axis('off')
a.grid(False)
print(f'Iterating over all {num_domains} domains in the Suite:')
for j, [domain, task] in enumerate(domains_tasks.items()):
print(domain, task)
env = suite.load(domain, task, task_kwargs={'random': random_state})
timestep = env.reset()
pixels = env.physics.render(height=200, width=200, camera_id=0)
ax.flat[j].imshow(pixels)
ax.flat[j].set_title(domain + ': ' + task)
clear_output()
# + [markdown] id="JHSvxHiaopDb"
# # Locomotion
# + [markdown] id="yTn3C03dpHzL"
# ## Humanoid running along corridor with obstacles
#
# As an illustrative example of using the Locomotion infrastructure to build an RL environment, consider placing a humanoid in a corridor with walls, and a task specifying that the humanoid will be rewarded for running along this corridor, navigating around the wall obstacles using vision. We instantiate the environment as a composition of the Walker, Arena, and Task as follows. First, we build a position-controlled CMU humanoid walker.
# + id="gE8rrB7PpN9X"
#@title A position controlled `cmu_humanoid`
walker = cmu_humanoid.CMUHumanoidPositionControlledV2020(
observable_options={'egocentric_camera': dict(enabled=True)})
# + [markdown] id="3fYbaDflBrgE"
# Next, we construct a corridor-shaped arena that is obstructed by walls.
# + id="t-O17Fnm3E6R"
#@title A corridor arena with wall obstacles
arena = corridor_arenas.WallsCorridor(
wall_gap=3.,
wall_width=distributions.Uniform(2., 3.),
wall_height=distributions.Uniform(2.5, 3.5),
corridor_width=4.,
corridor_length=30.,
)
# + [markdown] id="970nN38eBx-R"
# The task constructor places the walker in the arena.
# + id="dz4Jy2UGpQ4Z"
#@title A task to navigate the arena
task = corridor_tasks.RunThroughCorridor(
walker=walker,
arena=arena,
walker_spawn_position=(0.5, 0, 0),
target_velocity=3.0,
physics_timestep=0.005,
control_timestep=0.03,
)
# + [markdown] id="r-Oy-qTSB4HW"
# Finally, a task that rewards the agent for running down the corridor at a specific velocity is instantiated as a `composer.Environment`.
# + id="sQXlaEZk3ytl"
#@title The `RunThroughCorridor` environment
env = composer.Environment(
task=task,
time_limit=10,
random_state=np.random.RandomState(42),
strip_singleton_obs_buffer_dim=True,
)
env.reset()
pixels = []
for camera_id in range(3):
pixels.append(env.physics.render(camera_id=camera_id, width=240))
PIL.Image.fromarray(np.hstack(pixels))
# + [markdown] id="HuuQLm8YopDe"
# ## Multi-Agent Soccer
# + [markdown] id="OPNshDEEopDf"
# Building on Composer and Locomotion libraries, the Multi-agent soccer environments, introduced in [this paper](https://arxiv.org/abs/1902.07151), follow a consistent task structure of Walkers, Arena, and Task where instead of a single walker, we inject multiple walkers that can interact with each other physically in the same scene. The code snippet below shows how to instantiate a 2-vs-2 Multi-agent Soccer environment with the simple, 5 degree-of-freedom `BoxHead` walker type.
# + id="zAb3je0DAeQo"
#@title 2-v-2 `Boxhead` soccer
random_state = np.random.RandomState(42)
env = soccer.load(
team_size=2,
time_limit=45.,
random_state=random_state,
disable_walker_contacts=False,
walker_type=soccer.WalkerType.BOXHEAD,
)
env.reset()
pixels = []
# Select a random subset of 6 cameras (soccer envs have lots of cameras)
cameras = random_state.choice(env.physics.model.ncam, 6, replace=False)
for camera_id in cameras:
pixels.append(env.physics.render(camera_id=camera_id, width=240))
image = np.vstack((np.hstack(pixels[:3]), np.hstack(pixels[3:])))
PIL.Image.fromarray(image)
# + [markdown] id="J_5C2k0NGvxE"
# It can trivially be replaced by e.g. the `WalkerType.ANT` walker:
# + id="WDIGodhBG-Mn"
#@title 3-v-3 `Ant` soccer
random_state = np.random.RandomState(42)
env = soccer.load(
team_size=3,
time_limit=45.,
random_state=random_state,
disable_walker_contacts=False,
walker_type=soccer.WalkerType.ANT,
)
env.reset()
pixels = []
cameras = random_state.choice(env.physics.model.ncam, 6, replace=False)
for camera_id in cameras:
pixels.append(env.physics.render(camera_id=camera_id, width=240))
image = np.vstack((np.hstack(pixels[:3]), np.hstack(pixels[3:])))
PIL.Image.fromarray(image)
# + [markdown] id="MvK9BW4A5c9p"
# # Manipulation
# + [markdown] id="jPt27n2Dch_m"
# The `manipulation` module provides a robotic arm, a set of simple objects, and tools for building reward functions for manipulation tasks.
# + id="cZxmJoovahCA"
#@title Listing all `manipulation` tasks{vertical-output: true}
# `ALL` is a tuple containing the names of all of the environments in the suite.
print('\n'.join(manipulation.ALL))
# + id="oj0cJFlR5nTS"
#@title Listing `manipulation` tasks that use vision{vertical-output: true}
print('\n'.join(manipulation.get_environments_by_tag('vision')))
# + id="e_6q4FqFIKxy"
#@title Loading and simulating a `manipulation` task{vertical-output: true}
#@test {"timeout": 180}
env = manipulation.load('stack_2_of_3_bricks_random_order_vision', seed=42)
action_spec = env.action_spec()
def sample_random_action():
return env.random_state.uniform(
low=action_spec.minimum,
high=action_spec.maximum,
).astype(action_spec.dtype, copy=False)
# Step the environment through a full episode using random actions and record
# the camera observations.
frames = []
timestep = env.reset()
frames.append(timestep.observation['front_close'])
while not timestep.last():
timestep = env.step(sample_random_action())
frames.append(timestep.observation['front_close'])
all_frames = np.concatenate(frames, axis=0)
display_video(all_frames, 30)
| tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
labels = 'Frogs', 'Hogs', 'Dogs', 'Logs'
sizes = [15, 30, 45, 10]
explode = (0, 0.1, 0, 0)
#这个explode是指将饼状图的部分与其他分割的大小,这里之分割出hogs,度数为0.1,这个数字表示和主图分割的距离
fig1, ax1 = plt.subplots()
ax1.pie(sizes, explode=explode, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
ax1.axis('equal')
#这个equal是让整个饼图为一个圆形,如果不加,则是椭圆形
plt.show()
# +
import numpy as np
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
# vc=[1,2,39,0,8]
# vb=[1,2,38,0,8]
# image = np.corrcoef(vc, vb)
np.random.seed(0)
image = np.random.uniform(size=(6, 6))
# print (image)
ax.imshow(image, cmap=plt.cm.gray, interpolation='nearest',origin='upper')
ax.set_title('dropped spines')
# Move left and bottom spines outward by 10 points
ax.spines['left'].set_position(('outward', 10))
ax.spines['bottom'].set_position(('outward', 10))
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
plt.show()
#image是一个6*6的矩阵,此类型的图可以显示矩阵数字的大小,颜色越浅,说明这个位置的数组越大
#举个栗子
print(image[2][1])
print(image[2][2])
#是不是很神奇 [滑稽]
# +
import pandas as pd
from numpy import mean, multiply, cov, corrcoef, std
df = pd.read_csv('matplotlib3.csv')
df.rename(columns={'Unnamed: 0':'date'},inplace=True)
df.head()
# -
b=[1,3,5,6]
print (np.cov(b))
print (sum((np.multiply(b,b))-np.mean(b)*np.mean(b))/3)
vc=[1,2,39,0,8]
vb=[1,2,38,0,8]
print (mean(multiply((vc-mean(vc)),(vb-mean(vb))))/(std(vb)*std(vc)))
#corrcoef得到相关系数矩阵(向量的相似程度)
print (corrcoef(vc,vb))
# +
l = []
for x in df.columns[1:]:
l.append(df[x].values)
matrix = np.mat(l)
image = corrcoef(matrix)
fig, ax = plt.subplots(figsize=(8, 8))
ax.imshow(image, cmap=plt.cm.gray, interpolation='nearest',origin='lowwer')
# ax.imshow?
ax.set_title('dropped spines')
# Move left and bottom spines outward by 10 points
ax.spines['left'].set_position(('outward', 10))
ax.spines['bottom'].set_position(('outward', 10))
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
plt.xticks([0, 1, 2, 3, 4, 5],df.columns[1:])
plt.yticks([0, 1, 2, 3, 4, 5],df.columns[1:])
plt.show()
# -
fig, ax = plt.subplots(figsize=(10, 6))
plt.plot(df.index, df['601398.XSHG'], label='601398.XSHG')
plt.plot(df.index, df['601939.XSHG'], label='601939.XSHG')
plt.plot(df.index, df['601939.XSHG']-df['601398.XSHG'], label='dif')
plt.legend(loc='upper left')
ax.spines['left'].set_position(('outward', 10))
ax.spines['bottom'].set_position(('outward', 10))
plt.show()
| jupyter/jupyters/matplotlib_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/6/6c/Star_Wars_Logo.svg/2000px-Star_Wars_Logo.svg.png" width="30%" height="30%" align="left">
# # SWAPI (Star Wars API)
# The Star Wars API, or "swapi" (Swah-pee) is the world's first quantified and programmatically-accessible data source for all the data from the Star Wars canon universe!
import swapi
# ### Sort All Planets in Star Wars (将所有星球按大小排列) :
for planet in swapi.get_all('planets').order_by('diameter'):
print(planet.name)
# ## Tatooine
# <img src="https://upload.wikimedia.org/wikipedia/en/7/7f/Tatooine.jpg" width="25%" height="25%" align="left">
# <img src="http://cdn0.dailydot.com/uploaded/images/original/2015/3/24/tatooine.jpg" width="56%" height="56%">
# Tatooine was a desert planet located in the Outer Rim Territories. The planet was part of a binary star system. Due to the lack of surface water, many residents of the planet instead drew water from the atmosphere via moisture farms.
# Tatooine was the homeworld of Anakin Skywalker, the Chosen One, and his son <NAME>. Anakin's former master, <NAME>, also resided on the planet after the Clone Wars ended, going by the name <NAME>. The Hutt Clan maintained a presence on Tatooine as well, with Jabba the Hutt owning a palace in the desert.
# _______________
# ## Naboo
# <img src="http://vignette4.wikia.nocookie.net/es.starwars/images/5/50/Naboo.jpg/revision/latest?cb=20060318190228" width="33%" height="33%" align="left">
# <img src="http://vignette2.wikia.nocookie.net/starwars/images/0/01/Theed-MoaTM.png/revision/latest?cb=20121002233339" width="55%" height="55%">
# An idyllic world close to the border of the Outer Rim Territories, Naboo is inhabited by peaceful humans known as the Naboo, and an indigenous species of intelligent amphibians called the Gungans. Naboo's surface consists of swampy lakes, rolling plains and green hills. Its population centers are beautiful -- Naboo's river cities are filled with classical architecture and greenery, while the underwater Gungan settlements are a beautiful display of exotic hydrostatic bubble technology.
# _______________
# ## Kamino
# <img src="http://vignette3.wikia.nocookie.net/starwars/images/5/57/Kamino.jpg/revision/latest?cb=20071007054356" width="20%" height="20%" align="left">
# <img src="http://img.lum.dolimg.com/v1/images/databank_kamino_01_169_f9027822.jpeg?region=0%2C0%2C1560%2C878&width=768" width="35%" height="35%">
# Kamino (pronounced /kə'minoʊ/) is an ocean planet. It was here that the Army of the Republic was generated. It is inhabited by a race of tall, elegant, long-necked creatures, called Kaminoans, who keep to themselves and are known for their cloning technology. 10 years before the Clone Wars, <NAME>, using Jedi Master Sifo-Dyas secretly funded the production of an army for the Republic. In Star Wars: Episode II, Obi-Wan Kenobi was directed here to follow a lead on the whereabouts of Senator Amidala's would-be assassin through a single poison dart. During Obi-Wan's stay on Kamino at Tipoca City, he discovered the source of the Clone Army's genetic code, one <NAME> and his "son" <NAME>.
# ___________
# ## Coruscant
# <img src="http://vignette3.wikia.nocookie.net/starwars/images/1/17/Coruscant-AoTCW.jpg/revision/latest?cb=20100115192302" width="22%" height="22%" align="left">
# <img src="http://vignette2.wikia.nocookie.net/starwars/images/0/06/Coruscant_apartment_view.png/revision/latest?cb=20130118200812" width="50%" height="50%">
# Coruscant (pronounced /'kɔɹəsɑnt/), also known as Imperial Center during the reign of the Galactic Empire, was a planet that served as the galactic capital and seat of government of the Galactic Republic and the Galactic Empire, and was the homeworld of the human species. Coruscant was notable for its vast urban sprawl covering the entire planet called Galactic City that was constantly awake as airspeeders and starships moved in the flow of Coruscant's skylane traffic.
# _______________________
# ## Geonosis
# <img src="http://vignette2.wikia.nocookie.net/starwars/images/6/6d/Geonosis_AotC.png/revision/latest?cb=20121231120327" width="40%" height="40%" align="left">
# <img src="http://cdn.obsidianportal.com/assets/132814/Paisaje_Geonosis.jpg" width="50%" height="50%">
# A harsh rocky world less than a parsec away from Tatooine, Geonosis is a ringed planet beyond the borders of the Galactic Republic. Its uninviting surface is made up of harsh, rocky desert, and the creatures that evolved on Geonosis are well equipped to survive in the brutal environment. The most advanced lifeform are the Geonosians, sentient insectoids that inhabit towering spire-hives. The Geonosians maintain large factories for the production of droids and weapons for the Separatist cause in the Clone Wars. After the Clone Wars, the Empire chose Geonosis as the construction site of the first Death Star, restricting travel to the system and sterilizing the planet’s surface.
# _________________________
# ## Utapau
# <img src="http://vignette4.wikia.nocookie.net/starwars/images/c/ce/UtapauRotS.png/revision/latest?cb=20160118063015" height="22%" width="22%" align="left">
# <img src="http://vignette2.wikia.nocookie.net/starwars/images/f/f6/Tenth_Level.png/revision/latest?cb=20130215052019" height="56%" width="56%" >
# Ringed by numerous moons, the remote sanctuary of Utapau is a planet pitted with enormous sinkholes. Its windswept surface is dry and desert-like, but deep in the porous crust, at the bottom of the sinkholes are pools of life-giving water. Many of the sinkholes on one of the planet's small continents are lined with intricate cities that stretch deep into caves and crevices beneath the planet's surface, as well as mining operations that seek to uncover valuable minerals from the planet's depths.
# ## Mustafar
# <img src="http://vignette2.wikia.nocookie.net/starwars/images/a/af/Mustafar_DB.png/revision/latest?cb=20160118061913" height="23%" width="23%" align="left">
# <img src="http://img.lum.dolimg.com/v1/images/databank_mustafar_01_169_5b470758.jpeg?region=0%2C0%2C1560%2C878&width=768" height="40%" width="40%">
#
# A tiny, fiery planet in the Outer Rim, Mustafar maintains an erratic orbit between two gas giants. Mustafar is rich in unique and valuable minerals which have long been mined by the Tech Union. Its lava pits and rivers make this planet a dangerous habitat; its natives have burly, tough bodies that can withstand extreme heat. The planet became the backdrop for the fateful duel between Obi-Wan Kenobi and Anakin Skywalker. After the rise of the Empire, captured Jedi were brought to the volcanic world for interrogation and execution.
# _____________________
# ## Jakku
# <img src="http://www.starwarsnewsnet.com/wp-content/uploads/2015/06/Jakku.jpg" height="22%" width="22%" align="left">
# <img src="http://vignette1.wikia.nocookie.net/banthapedia/images/e/ef/Jakku.jpg/revision/latest?cb=20160105162421" height="40%" width="40%">
# A harsh desert world, Jakku is a lawless place that’s home to thieves, outlaws and refugees. During the final years of the Galactic Civil War, Jakku was the site of a secret Imperial research base and served as a jumping-off point for warships heading into the Unknown Regions. The Empire made a last stand above the frontier planet, leading to a pitched battle between rebel and Imperial fleets. Jakku’s sandy wastes are littered with old wrecks, picked over by desperate scavengers searching for parts that can be salvaged and sold.
# ___________________
# ### Query Persons Who Drove More Than 1 Starships (查看哪些人开过1艘以上的飞船):
for people in swapi.get_all("people").iter():
if len(people.starships) > 1:
print(people.name)
# ### Index Whether Jar Jar Binks Appeared in Episode 1 (检索Jar Jar Binks是否在电影中出现):
pm = swapi.get_film(4) #Here Episode 1 is the 4th movie in Star Wars
jj = swapi.get_person(36)
for c in pm.get_characters().iter():
if c.name == jj.name:
print('Jar Jar Binks, strange character!')
# <img src="http://img.lum.dolimg.com/v1/images/databank_jarjarbinks_01_169_c70767ab.jpeg" width="50%" height="50%" align="left">
# # All Episodes (所有电影):
films = swapi.get_all('films')
for film in films.iter():
print(film)
# <img src="http://img.lum.dolimg.com/v1/images/Star-Wars-New-Hope-IV-Poster_c217085b.jpeg?region=49%2C43%2C580%2C914&width=480" width="20%" height="20%" align="left">
# <img src="http://img.lum.dolimg.com/v1/images/Star-Wars-Empire-Strikes-Back-V-Poster_878f7fce.jpeg?region=25%2C22%2C612%2C953&width=480" width="20%" height="20%" align="left">
# <img src="http://img.lum.dolimg.com/v1/images/Star-Wars-Return-Jedi-VI-Poster_a10501d2.jpeg?region=12%2C9%2C618%2C982&width=480" width="20%" height="20%" align="left">
# <img src="http://img.lum.dolimg.com/v1/images/Star-Wars-Phantom-Menace-I-Poster_3c1ff9eb.jpeg?region=15%2C9%2C651%2C979&width=480" width="20%" height="20%" align="left">
# <img src="http://img.lum.dolimg.com/v1/images/Star-Wars-Attack-Clones-II-Poster_53baa2e7.jpeg?region=18%2C0%2C660%2C1000&width=480" width="20%" height="20%" align="left">
# <img src="http://img.lum.dolimg.com/v1/images/Star-Wars-Revenge-Sith-III-Poster_646108ce.jpeg?region=0%2C0%2C736%2C1090&width=480" width="20%" height="20%" align="left">
# <img src="http://a.dilcdn.com/bl/wp-content/uploads/sites/6/2015/10/tfa_poster_wide_header-1536x864-959818851016.jpg" width="50%" height="50%" align="left">
# # All Characters (所有角色):
persons = swapi.get_all('people')
for person in persons.iter():
print(person)
# # All Starships (所有飞船):
starships = swapi.get_all('starships')
for starship in starships.iter():
print(starship)
# ### Millennium Falcon (千年鹰号):
# <img src="http://vignette1.wikia.nocookie.net/starwars/images/4/43/MillenniumFalconTFA-Fathead.png/revision/latest?cb=20151013004955" width="40%" height="40%" align="left">
# A legendary starship despite its humble origins and deceptively shabby exterior, the Millennium Falcon has played a role in some of the Rebel Alliance's greatest victories over the Empire. On the surface, the Falcon looks like a worn down junker.
# _______________
# ### TIE Advanced x1
# <img src="http://vignette1.wikia.nocookie.net/starwars/images/9/92/TIEfighter2-Fathead.png/revision/latest?cb=20150309055216" width="25%" height="25%" align="left">
# The TIE Advanced x1 was an advanced prototype starfighter that was part of the TIE line manufactered by Sienar Fleet Systems. An update of the TIE Advanced v1 prototype, the TIE Advanced x1 featured a highly sophisticated target tracking system, making them useful against deflector shield generators. Several years before the Battle of Yavin, the Sith Lord Darth Vader piloted a TIE Advanced without any backup into the midst of a rebel fleet in a daring attack that resulted in the destruction of the rebels' flagship, Phoenix Home. Later, during the Battle of Yavin, Vader again piloted a TIE Advanced x1 to pursue and eliminate the Rebel fighters attacking the first Death Star. In this, he was almost entirely successful, until his fighter was unexpectedly knocked out of the battle when a TIE/LN starfighter collided with it after a surprise attack by the Millennium Falcon. After the collision, Vader regained control of the starfighter and escaped the destruction of the Death Star.
# ___________________
# ### X-Wing
# <img src="http://vignette4.wikia.nocookie.net/starwars/images/8/87/X-wing_SWB.png/revision/latest?cb=20151124042943" width="40%" height="40%" align="left">
# X-wing starfighters were a type of starfighter marked by their distinctive S-foils that resembled an 'X' in attack formation. During the Galactic Civil War, the Rebel Alliance used T-65B X-wing starfighters in a number of battles, including the Battle of Yavin and the Battle of Endor.
# ________________
# ### Naboo Royal Starship
# <img src="http://vignette2.wikia.nocookie.net/starwars/images/9/9e/Naboo_Royal_Starship.png/revision/latest?cb=20130619200757" width="40%" height="40%" align="left">
# The Naboo Royal Starship boasts a strikingly beautiful design that embodies the craftsmanship that prevailed in the peaceful years of the Republic. The polished, streamlined J-type 327 Nubian vessel lacks any offensive weaponry, but does feature strong shields and a competent hyperdrive. During the Trade Federation invasion of Naboo, Queen Amidala and her retinue escaped aboard the Royal Starship. Piloted by <NAME>, the vessel narrowly avoided the Trade Federation blockade, though its hyperdrive was damaged. The ship limped to Tatooine, where a replacement T-14 hyperdrive generator was procured.
# _______________________________
# ### Republic Cruiser
# <img src="http://img.lum.dolimg.com/v1/images/databank_republicattackcruiser_01_169_812f153d.jpeg?region=0%2C0%2C1560%2C878&width=768" width="50%" height="50%" align="left">
# The Republic attack cruiser was not primarily a cargo transport, troop carrier or other replenishment or supply vessel; it was developed and honed for ship-to-ship combat. Its huge armored shape had an aggressive, dagger-like profile, studded with heavy weapons emplacements capable of punching through powerful shields and sturdy armor. The front half of the ship's spine contained a massive flight deck capable of scrambling the latest in Republic starfighters. The huge hangar doors opened to allow ships to land or takeoff from the deck; the vessel also had a smaller docking bay entrance on its lower side.
# ________________
# ### AA-9 Coruscant freighter
# <img src="http://vignette3.wikia.nocookie.net/starwars/images/c/c7/Aa9coruscantfreighter.jpg/revision/latest?cb=20091201131352" width="35%" height="35%" align="left">
# In the days leading up to the Clone Wars, these ships were transports favored by refugees trying to escape Coruscant since travelers had no need to register themselves. It was for that reason that Jedi Master <NAME> suggested this method of transportation to then-padawan Anakin Skywalker when he was escorting Senator Amidala back to Naboo after an attempt on her life. The ships were outfitted with everything needed to carry a massive amount of people of various species across the galaxy, including cafeterias and various servant droids.
# ____________
# ### Death Star
# <img src="http://vignette3.wikia.nocookie.net/starwars/images/7/72/DeathStar1-SWE.png/revision/latest?cb=20150121020639" width="35%" height="35%" align="left">
# The Death Star, also known as the Death Star I, First Death Star, DS-1 platform[6] and previously known as the Ultimate Weapon and Sentinel Base, was a moon-sized deep space mobile battle station constructed by the Galactic Empire after the defeat of the Separatists in the Clone Wars. It boasted a primary weapon with sufficient firepower to destroy an entire planet with a single shot.
# _______________________
# # All Planets (所有星球):
planets = swapi.get_all('planets')
for planet in planets.order_by('diameter'):
print(planet)
# +
person = swapi.get_person(1)
planet = swapi.get_planet(1)
ship = swapi.get_starship(5)
film = swapi.get_film(4)
print(people)
print(planet)
print(ship)
print(film)
# -
# <img src="http://vignette2.wikia.nocookie.net/mec/images/9/9c/0000000000.jpg/revision/latest?cb=20120401033106" width="20%" height="20%" align="left">
# <img src="http://vignette4.wikia.nocookie.net/starwars/images/7/7b/Clone_trooper_armor.png/revision/latest?cb=20141223035439" width="26%" height="26%" align="left">
# <img src="http://vignette3.wikia.nocookie.net/theclonewiki/images/b/b1/Codyrun2.png/revision/latest?cb=20130308221228" width="25%" height="25%" align="left">
# <img src="http://vignette2.wikia.nocookie.net/starwars/images/2/21/Shock_troops_SWE.png/revision/latest?cb=20140619091751" width="25%" height="25%" align="left">
dir(person)
dir(planet)
dir(ship)
dir(film)
| StarWarsAPI/StarWarsAPI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import os
import matplotlib.pyplot as plt
# ## Non Uniform Data Generation:
# #### This notebook contains the code for generating the synthetic data used in the paper.
def generate_nonuniform_data(n, d, eta, kappa, index):
# The dimension of the relevant Region
d1 = np.int(d ** eta)
nt = 10000
exponent = (eta + kappa) / 2.0
r = d ** exponent
print(r'd_1 is %d'%(d1))
print(r'Kappa is %f'%(kappa))
print(r'The radius $\sqrt{d}$ is %f'%(np.sqrt(d)))
print('The radius r is %f'%(r))
# Making the features
np.random.seed(145)
# Train Data
X = np.random.normal(size=(n, d))
X = X.astype(np.float32)
for i in range(n):
X[i, :d1] = X[i, :d1] / np.linalg.norm(X[i, :d1]) * r
X[i, d1:] = X[i, d1:] / np.linalg.norm(X[i, d1:]) * np.sqrt(d)
# Test Data
np.random.seed(2)
XT = np.random.normal(size=(nt, d))
XT = XT.astype(np.float32)
for i in range(nt):
XT[i, :d1] = XT[i, :d1] / np.linalg.norm(XT[i, :d1]) * r
XT[i, d1:] = XT[i, d1:] / np.linalg.norm(XT[i, d1:]) * np.sqrt(d)
directory = './datasets/synthetic/'
np.save(directory + 'X_train_anisotropic_%d_%d_%d.npy'%(d, d1, index), X)
np.save(directory + 'X_test_anisotropic_%d_%d_%d.npy'%(d, d1, index), XT)
X0 = X[:, :d1]
X1 = XT[:, :d1]
del X, XT
# Make the labels
np.random.seed(14)
f = []
# The function has no linear component
beta2 = np.random.exponential(scale=1.0, size=(d1 - 1, 1))
beta3 = np.random.exponential(scale=1.0, size=(d1 - 2, 1))
beta4 = np.random.exponential(scale=1.0, size=(d1 - 3, 1))
Z = np.multiply(X0[:, :-1], X0[:, 1:])
temp = np.dot(Z, beta2)
f.append(temp)
Z = np.multiply(X0[:, :-2], X0[:, 1:-1])
Z = np.multiply(Z, X0[:, 2:])
temp = np.dot(Z, beta3)
f.append(temp)
Z = np.multiply(X0[:, :-3], X0[:, 1:-2])
Z = np.multiply(Z, X0[:, 2:-1])
Z = np.multiply(Z, X0[:, 3:])
temp = np.dot(Z, beta4)
f.append(temp)
normalization = [np.sqrt(np.mean(t ** 2)) for t in f]
for i in range(len(f)):
f[i] = f[i] / normalization[i]
totalf = f[0] + f[1] + f[2]
totalf = totalf.astype(np.float32)
g = []
Z = np.multiply(X1[:, :-1], X1[:, 1:])
temp = np.dot(Z, beta2)
g.append(temp)
Z = np.multiply(X1[:, :-2], X1[:, 1:-1])
Z = np.multiply(Z, X1[:, 2:])
temp = np.dot(Z, beta3)
g.append(temp)
Z = np.multiply(X1[:, :-3], X1[:, 1:-2])
Z = np.multiply(Z, X1[:, 2:-1])
Z = np.multiply(Z, X1[:, 3:])
temp = np.dot(Z, beta4)
g.append(temp)
for i in range(len(g)):
g[i] = g[i] / normalization[i]
totalg = g[0] + g[1] + g[2]
totalg = totalg.astype(np.float32)
np.save(directory + 'y_train_anisotropic_%d_%d_%d.npy'%(d, d1, index), totalf)
np.save(directory + 'y_test_anisotropic_%d_%d_%d.npy'%(d, d1, index), totalg)
d = 1024
eta = 2.0 / 5.0
n = 1024 * 1024
kappa_mat = np.linspace(0, 1, num=10, endpoint=False)
kappa_mat
for index in range(10):
generate_nonuniform_data(n, d, eta, kappa_mat[index], index)
| data generation code/Data Generation-Anisotropic-Shared.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # softmax回归的从零开始实现
#
# 这一节我们来动手实现softmax回归。首先导入本节实现所需的包或模块。
# + attributes={"classes": [], "id": "", "n": "1"}
# %matplotlib inline
import d2lzh as d2l
from mxnet import autograd, nd
# -
# ## 读取数据集
#
# 我们将使用Fashion-MNIST数据集,并设置批量大小为256。
# + attributes={"classes": [], "id": "", "n": "2"}
batch_size = 256
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size)
# -
# ## 初始化模型参数
#
# 跟线性回归中的例子一样,我们将使用向量表示每个样本。已知每个样本输入是高和宽均为28像素的图像。模型的输入向量的长度是$28 \times 28 = 784$:该向量的每个元素对应图像中每个像素。由于图像有10个类别,单层神经网络输出层的输出个数为10,因此softmax回归的权重和偏差参数分别为$784 \times 10$和$1 \times 10$的矩阵。
# + attributes={"classes": [], "id": "", "n": "3"}
# 将输入 24*24 进行展开
num_inputs = 784
# 因为有10类,所以输出为10
num_outputs = 10
W = nd.random.normal(scale=0.01, shape=(num_inputs, num_outputs))
b = nd.zeros(num_outputs)
# -
# 同之前一样,我们要为模型参数附上梯度。
# + attributes={"classes": [], "id": "", "n": "4"}
W.attach_grad()
b.attach_grad()
# -
# ## 实现softmax运算
#
# 在介绍如何定义softmax回归之前,我们先描述一下对如何对多维`NDArray`按维度操作。在下面的例子中,给定一个`NDArray`矩阵`X`。我们可以只对其中同一列(`axis=0`)或同一行(`axis=1`)的元素求和,并在结果中保留行和列这两个维度(`keepdims=True`)。
# + attributes={"classes": [], "id": "", "n": "5"}
X = nd.array([[1, 2, 3], [4, 5, 6]])
X.sum(axis=0, keepdims=True), X.sum(axis=1, keepdims=True)
# -
# 下面我们就可以定义前面小节里介绍的softmax运算了。在下面的函数中,矩阵`X`的行数是样本数,列数是输出个数。为了表达样本预测各个输出的概率,softmax运算会先通过`exp`函数对每个元素做指数运算,再对`exp`矩阵同行元素求和,最后令矩阵每行各元素与该行元素之和相除。这样一来,最终得到的矩阵每行元素和为1且非负。因此,该矩阵每行都是合法的概率分布。softmax运算的输出矩阵中的任意一行元素代表了一个样本在各个输出类别上的预测概率。
# + attributes={"classes": [], "id": "", "n": "6"}
def softmax(X):
X_exp = X.exp()
partition = X_exp.sum(axis=1, keepdims=True)
return X_exp / partition # 这里应用了广播机制
# -
# 可以看到,对于随机输入,我们将每个元素变成了非负数,且每一行和为1。
# + attributes={"classes": [], "id": "", "n": "7"}
X = nd.random.normal(shape=(2, 5))
X_prob = softmax(X)
X_prob, X_prob.sum(axis=1)
# -
# ## 定义模型
#
# 有了softmax运算,我们可以定义上节描述的softmax回归模型了。这里通过`reshape`函数将每张原始图像改成长度为`num_inputs`的向量。
# + attributes={"classes": [], "id": "", "n": "8"}
def net(X):
# 通过 reshape将三维的变成二维
return softmax(nd.dot(X.reshape((-1, num_inputs)), W) + b)
# -
# ## 定义损失函数
# 用于概率的损失函数
#
# 上一节中,我们介绍了softmax回归使用的交叉熵损失函数。为了得到标签的预测概率,我们可以使用`pick`函数。在下面的例子中,变量`y_hat`是2个样本在3个类别的预测概率,变量`y`是这2个样本的标签类别。通过使用`pick`函数,我们得到了2个样本的标签的预测概率。与[“softmax回归”](softmax-regression.ipynb)一节数学表述中标签类别离散值从1开始逐一递增不同,在代码中,标签类别的离散值是从0开始逐一递增的。
# + attributes={"classes": [], "id": "", "n": "9"}
y_hat = nd.array([[0.1, 0.3, 0.6], [0.3, 0.2, 0.5]])
y = nd.array([0, 2], dtype='int32')
nd.pick(y_hat, y)
# -
# 下面实现了[“softmax回归”](softmax-regression.ipynb)一节中介绍的交叉熵损失函数。
# + attributes={"classes": [], "id": "", "n": "10"}
def cross_entropy(y_hat, y):
return -nd.pick(y_hat, y).log()
# -
# ## 计算分类准确率
#
# 给定一个类别的预测概率分布`y_hat`,我们把预测概率最大的类别作为输出类别。如果它与真实类别`y`一致,说明这次预测是正确的。分类准确率即正确预测数量与总预测数量之比。
#
# 为了演示准确率的计算,下面定义准确率`accuracy`函数。其中`y_hat.argmax(axis=1)`返回矩阵`y_hat`每行中最大元素的索引,且返回结果与变量`y`形状相同。我们在[“数据操作”](../chapter_prerequisite/ndarray.ipynb)一节介绍过,相等条件判别式`(y_hat.argmax(axis=1) == y)`是一个值为0(相等为假)或1(相等为真)的`NDArray`。由于标签类型为整数,我们先将变量`y`变换为浮点数再进行相等条件判断。
# + attributes={"classes": [], "id": "", "n": "11"}
def accuracy(y_hat, y):
return (y_hat.argmax(axis=1) == y.astype('float32')).mean().asscalar()
# -
# 让我们继续使用在演示`pick`函数时定义的变量`y_hat`和`y`,并将它们分别作为预测概率分布和标签。可以看到,第一个样本预测类别为2(该行最大元素0.6在本行的索引为2),与真实标签0不一致;第二个样本预测类别为2(该行最大元素0.5在本行的索引为2),与真实标签2一致。因此,这两个样本上的分类准确率为0.5。
# + attributes={"classes": [], "id": "", "n": "12"}
accuracy(y_hat, y)
# -
# 类似地,我们可以评价模型`net`在数据集`data_iter`上的准确率。
# + attributes={"classes": [], "id": "", "n": "13"}
# 本函数已保存在d2lzh包中方便以后使用。该函数将被逐步改进:它的完整实现将在“图像增广”一节中
# 描述
def evaluate_accuracy(data_iter, net):
acc_sum, n = 0.0, 0
for X, y in data_iter:
y = y.astype('float32')
acc_sum += (net(X).argmax(axis=1) == y).sum().asscalar()
n += y.size
return acc_sum / n
# -
# 因为我们随机初始化了模型`net`,所以这个随机模型的准确率应该接近于类别个数10的倒数0.1。
# + attributes={"classes": [], "id": "", "n": "14"}
evaluate_accuracy(test_iter, net)
# -
# ## 训练模型
#
# 训练softmax回归的实现与[“线性回归的从零开始实现”](linear-regression-scratch.ipynb)一节介绍的线性回归中的实现非常相似。我们同样使用小批量随机梯度下降来优化模型的损失函数。在训练模型时,迭代周期数`num_epochs`和学习率`lr`都是可以调的超参数。改变它们的值可能会得到分类更准确的模型。
# + attributes={"classes": [], "id": "", "n": "21"}
num_epochs, lr = 5, 0.1
# 本函数已保存在d2lzh包中方便以后使用
def train_ch3(net, train_iter, test_iter, loss, num_epochs, batch_size,
params=None, lr=None, trainer=None):
for epoch in range(num_epochs):
train_l_sum, train_acc_sum, n = 0.0, 0.0, 0
for X, y in train_iter:
with autograd.record():
y_hat = net(X)
l = loss(y_hat, y).sum()
l.backward()
if trainer is None:
d2l.sgd(params, lr, batch_size)
else:
trainer.step(batch_size) # “softmax回归的简洁实现”一节将用到
y = y.astype('float32')
train_l_sum += l.asscalar()
train_acc_sum += (y_hat.argmax(axis=1) == y).sum().asscalar()
n += y.size
test_acc = evaluate_accuracy(test_iter, net)
print('epoch %d, loss %.4f, train acc %.3f, test acc %.3f'
% (epoch + 1, train_l_sum / n, train_acc_sum / n, test_acc))
train_ch3(net, train_iter, test_iter, cross_entropy, num_epochs, batch_size,
[W, b], lr)
# -
# ## 预测
#
# 训练完成后,现在就可以演示如何对图像进行分类了。给定一系列图像(第三行图像输出),我们比较一下它们的真实标签(第一行文本输出)和模型预测结果(第二行文本输出)。
# +
for X, y in test_iter:
break
true_labels = d2l.get_fashion_mnist_labels(y.asnumpy())
pred_labels = d2l.get_fashion_mnist_labels(net(X).argmax(axis=1).asnumpy())
titles = [true + '\n' + pred for true, pred in zip(true_labels, pred_labels)]
d2l.show_fashion_mnist(X[0:9], titles[0:9])
# -
# ## 小结
#
# * 可以使用softmax回归做多类别分类。与训练线性回归相比,你会发现训练softmax回归的步骤和它非常相似:获取并读取数据、定义模型和损失函数并使用优化算法训练模型。事实上,绝大多数深度学习模型的训练都有着类似的步骤。
#
# ## 练习
#
# * 在本节中,我们直接按照softmax运算的数学定义来实现softmax函数。这可能会造成什么问题?(提示:试一试计算$\exp(50)$的大小。)
# * 本节中的`cross_entropy`函数是按照[“softmax回归”](softmax-regression.ipynb)一节中的交叉熵损失函数的数学定义实现的。这样的实现方式可能有什么问题?(提示:思考一下对数函数的定义域。)
# * 你能想到哪些办法来解决上面的两个问题?
#
#
#
# ## 扫码直达[讨论区](https://discuss.gluon.ai/t/topic/741)
#
# 
| 深度学习/d2l-zh-1.1/2_chapter_deep-learning-basics/softmax-regression-scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + cellView="form" colab={} colab_type="code" id="k2QMetc6kava"
#@title ##### License
# Copyright 2018 The GraphNets Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ============================================================================
# + [markdown] colab_type="text" id="c5CPvyHM2CnU"
# # Physical dynamics of a mass-spring system
# This notebook and the accompanying code demonstrates how to use the Graph Nets library to learn to predict the motion of a set of masses connected by springs.
#
# The network is trained to predict the behaviour of a chain of five masses, connected by identical springs. The first and last masses are fixed; the others are subject to gravity.
#
# After training, the network's prediction ability is illustrated by comparing its output to the true behaviour of the structure. Then the network's ability to generalise is tested, by using it to predict the behaviour of a similar but more complicated mass/spring structure.
# + cellView="form" colab={} colab_type="code" id="Ss54UNGvkz5M"
#@title ### Install the Graph Nets library on this Colaboratory runtime { form-width: "60%", run: "auto"}
#@markdown <br>1. Connect to a local or hosted Colaboratory runtime by clicking the **Connect** button at the top-right.<br>2. Choose "Yes" below to install the Graph Nets library on the runtime machine with the correct dependencies. Note, this works both with local and hosted Colaboratory runtimes.
install_graph_nets_library = "No" #@param ["Yes", "No"]
if install_graph_nets_library.lower() == "yes":
print("Installing Graph Nets library and dependencies:")
print("Output message from command:\n")
# !pip install graph_nets "dm-sonnet<2" "tensorflow_probability<0.9"
else:
print("Skipping installation of Graph Nets library")
# + [markdown] colab_type="text" id="7E4elJkXFR4a"
# ### Install dependencies locally
#
# If you are running this notebook locally (i.e., not through Colaboratory), you will also need to install a few more dependencies. Run the following on the command line to install the graph networks library, as well as a few other dependencies:
#
# ```
# pip install graph_nets matplotlib scipy "tensorflow>=1.15,<2" "dm-sonnet<2" "tensorflow_probability<0.9"
# ```
# + [markdown] colab_type="text" id="hyVaNA-bGug3"
# # Code
# + cellView="form" colab={} colab_type="code" id="RzKvtoAgRA8e"
#@title Imports { form-width: "30%" }
# The demo dependencies are not installed with the library, but you can install
# them with:
#
# $ pip install jupyter matplotlib scipy
#
# Run the demo with:
#
# $ jupyter notebook <path>/<to>/<demos>/shortest_path.ipynb
# #%tensorflow_version 1.x # For Google Colab only.
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import time
from graph_nets import blocks
from graph_nets import utils_tf
from graph_nets.demos import models
from matplotlib import pyplot as plt
import numpy as np
import sonnet as snt
import tensorflow as tf
try:
import seaborn as sns
except ImportError:
pass
else:
sns.reset_orig()
SEED = 1
np.random.seed(SEED)
tf.set_random_seed(SEED)
# + cellView="form" colab={} colab_type="code" id="toCQhJIM93en"
#@title Helper functions { form-width: "30%" }
# pylint: disable=redefined-outer-name
def to_matrix(l, n):
return [l[i:i+n] for i in xrange(0, len(l), n)]
def base_graph(n, d):
"""Define a basic mass-spring system graph structure.
These are n masses (1kg) connected by springs in a chain-like structure. The
first and last masses are fixed. The masses are vertically aligned at the
start and are d meters apart; this is also the rest length for the springs
connecting them. Springs have spring constant 50 N/m and gravity is 10 N in
the negative y-direction.
Args:
n: number of masses
d: distance between masses (as well as springs' rest length)
Returns:
data_dict: dictionary with globals, nodes, edges, receivers and senders
to represent a structure like the one above.
"""
# Nodes
# Generate initial position and velocity for all masses.
# The left-most mass has is at position (0, 0); other masses (ordered left to
# right) have x-coordinate d meters apart from their left neighbor, and
# y-coordinate 0. All masses have initial velocity 0m/s.
numlines = 2 #the number of springs #200821 jsban
#nodes = np.zeros((n, 5), dtype=np.float32)
nodes = np.zeros((numlines, n, 5), dtype = np.float32)
half_width = d * n / 2.0
#nodes[:, 0] = np.linspace(-half_width, half_width, num=n, endpoint=False, dtype=np.float32)
# indicate that the first and last masses are fixed
#nodes[(0, -1)] = 1.#nodes[(0, -1), -1] = 1.
for i in range(numlines):
nodes[i,:][:, 0] = np.linspace(-half_width, half_width, num=n, endpoint=False, dtype=np.float32)
# indicate that the first and last masses are fixed
nodes[i,:][(0, -1), -1] = 1.
# Edges.
edges, senders, receivers = [], [], []
for j in range(numlines):
for i in range(n - 1):
left_node = i
right_node = i + 1
# The 'if' statements prevent incoming edges to fixed ends of the string.
if right_node < n - 1:
# Left incoming edge.
edges.append([50., d])
senders.append(left_node)
receivers.append(right_node)
if left_node > 0:
# Right incoming edge.
edges.append([50., d])
senders.append(right_node)
receivers.append(left_node)
senders = to_matrix(senders, int(len(senders)/2))
receivers = to_matrix(receivers, int(len(receivers)/2))
edges = to_matrix(edges, int(len(edges)/2))
return {
"globals": [0., -10.],
"nodes": nodes,
"edges": edges,
"receivers": receivers,
"senders": senders
}
def hookes_law(receiver_nodes, sender_nodes, k, x_rest):
"""Applies Hooke's law to springs connecting some nodes.
Args:
receiver_nodes: Ex5 tf.Tensor of [x, y, v_x, v_y, is_fixed] features for the
receiver node of each edge.
sender_nodes: Ex5 tf.Tensor of [x, y, v_x, v_y, is_fixed] features for the
sender node of each edge.
k: Spring constant for each edge.
x_rest: Rest length of each edge.
Returns:
Nx2 Tensor of the force [f_x, f_y] acting on each edge.
"""
diff = receiver_nodes[..., 0:2] - sender_nodes[..., 0:2]
x = tf.norm(diff, axis=-1, keepdims=True)
force_magnitude = -1 * tf.multiply(k, (x - x_rest) / x)
force = force_magnitude * diff
return force
def euler_integration(nodes, force_per_node, step_size):
"""Applies one step of Euler integration.
Args:
nodes: Ex5 tf.Tensor of [x, y, v_x, v_y, is_fixed] features for each node.
force_per_node: Ex2 tf.Tensor of the force [f_x, f_y] acting on each edge.
step_size: Scalar.
Returns:
A tf.Tensor of the same shape as `nodes` but with positions and velocities
updated.
"""
is_fixed = nodes[..., 4:5]
# set forces to zero for fixed nodes
force_per_node *= 1 - is_fixed
new_vel = nodes[..., 2:4] + force_per_node * step_size
return new_vel
class SpringMassSimulator(snt.AbstractModule):
"""Implements a basic Physics Simulator using the blocks library."""
def __init__(self, step_size, name="SpringMassSimulator"):
super(SpringMassSimulator, self).__init__(name=name)
self._step_size = step_size
with self._enter_variable_scope():
self._aggregator = blocks.ReceivedEdgesToNodesAggregator(
reducer=tf.unsorted_segment_sum)
def _build(self, graph):
"""Builds a SpringMassSimulator.
Args:
graph: A graphs.GraphsTuple having, for some integers N, E, G:
- edges: Nx2 tf.Tensor of [spring_constant, rest_length] for each
edge.
- nodes: Ex5 tf.Tensor of [x, y, v_x, v_y, is_fixed] features for each
node.
- globals: Gx2 tf.Tensor containing the gravitational constant.
Returns:
A graphs.GraphsTuple of the same shape as `graph`, but where:
- edges: Holds the force [f_x, f_y] acting on each edge.
- nodes: Holds positions and velocities after applying one step of
Euler integration.
"""
receiver_nodes = blocks.broadcast_receiver_nodes_to_edges(graph)
sender_nodes = blocks.broadcast_sender_nodes_to_edges(graph)
spring_force_per_edge = hookes_law(receiver_nodes, sender_nodes,
graph.edges[..., 0:1],
graph.edges[..., 1:2])
graph = graph.replace(edges=spring_force_per_edge)
spring_force_per_node = self._aggregator(graph)
gravity = blocks.broadcast_globals_to_nodes(graph)
updated_velocities = euler_integration(
graph.nodes, spring_force_per_node + gravity, self._step_size)
graph = graph.replace(nodes=updated_velocities)
return graph
def prediction_to_next_state(input_graph, predicted_graph, step_size):
# manually integrate velocities to compute new positions
new_pos = input_graph.nodes[..., :2] + predicted_graph.nodes * step_size
new_nodes = tf.concat(
[new_pos, predicted_graph.nodes, input_graph.nodes[..., 4:5]], axis=-1)
return input_graph.replace(nodes=new_nodes)
def roll_out_physics(simulator, graph, steps, step_size):
"""Apply some number of steps of physical laws to an interaction network.
Args:
simulator: A SpringMassSimulator, or some module or callable with the same
signature.
graph: A graphs.GraphsTuple having, for some integers N, E, G:
- edges: Nx2 tf.Tensor of [spring_constant, rest_length] for each edge.
- nodes: Ex5 tf.Tensor of [x, y, v_x, v_y, is_fixed] features for each
node.
- globals: Gx2 tf.Tensor containing the gravitational constant.
steps: An integer.
step_size: Scalar.
Returns:
A pair of:
- The graph, updated after `steps` steps of simulation;
- A `steps+1`xNx5 tf.Tensor of the node features at each step.
"""
def body(t, graph, nodes_per_step):
predicted_graph = simulator(graph)
if isinstance(predicted_graph, list):
predicted_graph = predicted_graph[-1]
graph = prediction_to_next_state(graph, predicted_graph, step_size)
return t + 1, graph, nodes_per_step.write(t, graph.nodes)
nodes_per_step = tf.TensorArray(
dtype=graph.nodes.dtype, size=steps + 1, element_shape=graph.nodes.shape)
nodes_per_step = nodes_per_step.write(0, graph.nodes)
_, g, nodes_per_step = tf.while_loop(
lambda t, *unused_args: t <= steps,
body,
loop_vars=[1, graph, nodes_per_step])
return g, nodes_per_step.stack()
def apply_noise(graph, node_noise_level, edge_noise_level, global_noise_level):
"""Applies uniformly-distributed noise to a graph of a physical system.
Noise is applied to:
- the x and y coordinates (independently) of the nodes;
- the spring constants of the edges;
- the y coordinate of the global gravitational constant.
Args:
graph: a graphs.GraphsTuple having, for some integers N, E, G:
- nodes: Nx5 Tensor of [x, y, _, _, _] for each node.
- edges: Ex2 Tensor of [spring_constant, _] for each edge.
- globals: Gx2 tf.Tensor containing the gravitational constant.
node_noise_level: Maximum distance to perturb nodes' x and y coordinates.
edge_noise_level: Maximum amount to perturb edge spring constants.
global_noise_level: Maximum amount to perturb the Y component of gravity.
Returns:
The input graph, but with noise applied.
"""
node_position_noise = tf.random_uniform(
[graph.nodes.shape[0].value, 2],
minval=-node_noise_level,
maxval=node_noise_level)
edge_spring_constant_noise = tf.random_uniform(
[graph.edges.shape[0].value, 1],
minval=-edge_noise_level,
maxval=edge_noise_level)
global_gravity_y_noise = tf.random_uniform(
[graph.globals.shape[0].value, 1],
minval=-global_noise_level,
maxval=global_noise_level)
return graph.replace(
nodes=tf.concat(
[graph.nodes[..., :2] + node_position_noise, graph.nodes[..., 2:]],
axis=-1),
edges=tf.concat(
[
graph.edges[..., :1] + edge_spring_constant_noise,
graph.edges[..., 1:]
],
axis=-1),
globals=tf.concat(
[
graph.globals[..., :1],
graph.globals[..., 1:] + global_gravity_y_noise
],
axis=-1))
def set_rest_lengths(graph):
"""Computes and sets rest lengths for the springs in a physical system.
The rest length is taken to be the distance between each edge's nodes.
Args:
graph: a graphs.GraphsTuple having, for some integers N, E:
- nodes: Nx5 Tensor of [x, y, _, _, _] for each node.
- edges: Ex2 Tensor of [spring_constant, _] for each edge.
Returns:
The input graph, but with [spring_constant, rest_length] for each edge.
"""
receiver_nodes = blocks.broadcast_receiver_nodes_to_edges(graph)
sender_nodes = blocks.broadcast_sender_nodes_to_edges(graph)
rest_length = tf.norm(
receiver_nodes[..., :2] - sender_nodes[..., :2], axis=-1, keepdims=True)
return graph.replace(
edges=tf.concat([graph.edges[..., :1], rest_length], axis=-1))
def generate_trajectory(simulator, graph, steps, step_size, node_noise_level,
edge_noise_level, global_noise_level):
"""Applies noise and then simulates a physical system for a number of steps.
Args:
simulator: A SpringMassSimulator, or some module or callable with the same
signature.
graph: a graphs.GraphsTuple having, for some integers N, E, G:
- nodes: Nx5 Tensor of [x, y, v_x, v_y, is_fixed] for each node.
- edges: Ex2 Tensor of [spring_constant, _] for each edge.
- globals: Gx2 tf.Tensor containing the gravitational constant.
steps: Integer; the length of trajectory to generate.
step_size: Scalar.
node_noise_level: Maximum distance to perturb nodes' x and y coordinates.
edge_noise_level: Maximum amount to perturb edge spring constants.
global_noise_level: Maximum amount to perturb the Y component of gravity.
Returns:
A pair of:
- The input graph, but with rest lengths computed and noise applied.
- A `steps+1`xNx5 tf.Tensor of the node features at each step.
"""
graph = apply_noise(graph, node_noise_level, edge_noise_level,
global_noise_level)
graph = set_rest_lengths(graph)
_, n = roll_out_physics(simulator, graph, steps, step_size)
return graph, n
def create_loss_ops(target_op, output_ops):
"""Create supervised loss operations from targets and outputs.
Args:
target_op: The target velocity tf.Tensor.
output_ops: The list of output graphs from the model.
Returns:
A list of loss values (tf.Tensor), one per output op.
"""
loss_ops = [
tf.reduce_mean(
tf.reduce_sum((output_op.nodes - target_op[..., 2:4])**2, axis=-1))
for output_op in output_ops
]
return loss_ops
def make_all_runnable_in_session(*args):
"""Apply make_runnable_in_session to an iterable of graphs."""
return [utils_tf.make_runnable_in_session(a) for a in args]
# pylint: enable=redefined-outer-name
# + cellView="form" colab={} colab_type="code" id="pf7u0zuN_ktd"
#@title Set up model training and evaluation { form-width: "30%" }
# The model we explore includes three components:
# - An "Encoder" graph net, which independently encodes the edge, node, and
# global attributes (does not compute relations etc.).
# - A "Core" graph net, which performs N rounds of processing (message-passing)
# steps. The input to the Core is the concatenation of the Encoder's output
# and the previous output of the Core (labeled "Hidden(t)" below, where "t" is
# the processing step).
# - A "Decoder" graph net, which independently decodes the edge, node, and
# global attributes (does not compute relations etc.), on each
# message-passing step.
#
# Hidden(t) Hidden(t+1)
# | ^
# *---------* | *------* | *---------*
# | | | | | | | |
# Input --->| Encoder | *->| Core |--*->| Decoder |---> Output(t)
# | |---->| | | |
# *---------* *------* *---------*
#
# The model is trained by supervised learning. Input mass-spring systems are
# procedurally generated, where the nodes represent the positions, velocities,
# and indicators of whether the mass is fixed in space or free to move, the
# edges represent the spring constant and spring rest length, and the global
# attribute represents the variable coefficient of gravitational acceleration.
# The outputs/targets have the same structure, with the nodes representing the
# masses' next-step states.
#
# The training loss is computed on the output of each processing step. The
# reason for this is to encourage the model to try to solve the problem in as
# few steps as possible. It also helps make the output of intermediate steps
# more interpretable.
#
# There's no need for a separate evaluate dataset because the inputs are
# never repeated, so the training loss is the measure of performance on graphs
# from the input distribution.
#
# We also evaluate how well the models generalize to systems which are one mass
# larger, and smaller, than those from the training distribution. The loss is
# computed as the mean over a 50-step rollout, where each step's input is the
# the previous step's output.
#
# Variables with the suffix _tr are training parameters, and variables with the
# suffix _ge are test/generalization parameters.
#
# After around 10000-20000 training iterations the model reaches good
# performance on mass-spring systems with 5-8 masses.
tf.reset_default_graph()
rand = np.random.RandomState(SEED)
# Model parameters.
num_processing_steps_tr = 1
num_processing_steps_ge = 1
# Data / training parameters.
num_training_iterations = 100000
batch_size_tr = 256
batch_size_ge = 100
num_time_steps = 50
step_size = 0.1
num_masses_min_max_tr = (5, 9)
dist_between_masses_min_max_tr = (0.2, 1.0)
# Create the model.
model = models.EncodeProcessDecode(node_output_size=2)
# Data.
# Base graphs for training.
num_masses_tr = rand.randint(*num_masses_min_max_tr, size=batch_size_tr)
dist_between_masses_tr = rand.uniform(
*dist_between_masses_min_max_tr, size=batch_size_tr)
static_graph_tr = [
base_graph(n, d) for n, d in zip(num_masses_tr, dist_between_masses_tr)
]
base_graph_tr = utils_tf.data_dicts_to_graphs_tuple(static_graph_tr)
# Base graphs for testing.
# 4 masses 1m apart in a chain like structure.
base_graph_4_ge = utils_tf.data_dicts_to_graphs_tuple(
[base_graph(4, 1)] * batch_size_ge)
# 9 masses 0.5m apart in a chain like structure.
base_graph_9_ge = utils_tf.data_dicts_to_graphs_tuple(
[base_graph(9, 0.5)] * batch_size_ge)
# True physics simulator for data generation.
simulator = SpringMassSimulator(step_size=step_size)
# Training.
# Generate a training trajectory by adding noise to initial
# position, spring constants and gravity
initial_conditions_tr, true_trajectory_tr = generate_trajectory(
simulator,
base_graph_tr,
num_time_steps,
step_size,
node_noise_level=0.04,
edge_noise_level=5.0,
global_noise_level=1.0)
# Random start step.
t = tf.random_uniform([], minval=0, maxval=num_time_steps - 1, dtype=tf.int32)
input_graph_tr = initial_conditions_tr.replace(nodes=true_trajectory_tr[t])
target_nodes_tr = true_trajectory_tr[t + 1]
output_ops_tr = model(input_graph_tr, num_processing_steps_tr)
# Test data: 4-mass string.
initial_conditions_4_ge, true_trajectory_4_ge = generate_trajectory(
lambda x: model(x, num_processing_steps_ge),
base_graph_4_ge,
num_time_steps,
step_size,
node_noise_level=0.04,
edge_noise_level=5.0,
global_noise_level=1.0)
_, true_nodes_rollout_4_ge = roll_out_physics(
simulator, initial_conditions_4_ge, num_time_steps, step_size)
_, predicted_nodes_rollout_4_ge = roll_out_physics(
lambda x: model(x, num_processing_steps_ge), initial_conditions_4_ge,
num_time_steps, step_size)
# Test data: 9-mass string.
initial_conditions_9_ge, true_trajectory_9_ge = generate_trajectory(
lambda x: model(x, num_processing_steps_ge),
base_graph_9_ge,
num_time_steps,
step_size,
node_noise_level=0.04,
edge_noise_level=5.0,
global_noise_level=1.0)
_, true_nodes_rollout_9_ge = roll_out_physics(
simulator, initial_conditions_9_ge, num_time_steps, step_size)
_, predicted_nodes_rollout_9_ge = roll_out_physics(
lambda x: model(x, num_processing_steps_ge), initial_conditions_9_ge,
num_time_steps, step_size)
# Training loss.
loss_ops_tr = create_loss_ops(target_nodes_tr, output_ops_tr)
# Training loss across processing steps.
loss_op_tr = sum(loss_ops_tr) / num_processing_steps_tr
# Test/generalization loss: 4-mass.
loss_op_4_ge = tf.reduce_mean(
tf.reduce_sum(
(predicted_nodes_rollout_4_ge[..., 2:4] -
true_nodes_rollout_4_ge[..., 2:4])**2,
axis=-1))
# Test/generalization loss: 9-mass string.
loss_op_9_ge = tf.reduce_mean(
tf.reduce_sum(
(predicted_nodes_rollout_9_ge[..., 2:4] -
true_nodes_rollout_9_ge[..., 2:4])**2,
axis=-1))
# Optimizer.
learning_rate = 1e-3
optimizer = tf.train.AdamOptimizer(learning_rate)
step_op = optimizer.minimize(loss_op_tr)
input_graph_tr = make_all_runnable_in_session(input_graph_tr)
initial_conditions_4_ge = make_all_runnable_in_session(initial_conditions_4_ge)
initial_conditions_9_ge = make_all_runnable_in_session(initial_conditions_9_ge)
# + cellView="form" colab={} colab_type="code" id="TpABTYk0Ap-V"
#@title Reset session { form-width: "30%" }
# This cell resets the Tensorflow session, but keeps the same computational
# graph.
try:
sess.close()
except NameError:
pass
sess = tf.Session()
sess.run(tf.global_variables_initializer())
last_iteration = 0
logged_iterations = []
losses_tr = []
losses_4_ge = []
losses_9_ge = []
# + cellView="form" colab={} colab_type="code" id="4PCPvXiHA7q7"
#@title Run training { form-width: "30%" }
# You can interrupt this cell's training loop at any time, and visualize the
# intermediate results by running the next cell (below). You can then resume
# training by simply executing this cell again.
# How much time between logging and printing the current results.
log_every_seconds = 20
print("# (iteration number), T (elapsed seconds), "
"Ltr (training 1-step loss), "
"Lge4 (test/generalization rollout loss for 4-mass strings), "
"Lge9 (test/generalization rollout loss for 9-mass strings)")
start_time = time.time()
last_log_time = start_time
for iteration in range(last_iteration, num_training_iterations):
last_iteration = iteration
train_values = sess.run({
"step": step_op,
"loss": loss_op_tr,
"input_graph": input_graph_tr,
"target_nodes": target_nodes_tr,
"outputs": output_ops_tr
})
the_time = time.time()
elapsed_since_last_log = the_time - last_log_time
if elapsed_since_last_log > log_every_seconds:
last_log_time = the_time
test_values = sess.run({
"loss_4": loss_op_4_ge,
"true_rollout_4": true_nodes_rollout_4_ge,
"predicted_rollout_4": predicted_nodes_rollout_4_ge,
"loss_9": loss_op_9_ge,
"true_rollout_9": true_nodes_rollout_9_ge,
"predicted_rollout_9": predicted_nodes_rollout_9_ge
})
elapsed = time.time() - start_time
losses_tr.append(train_values["loss"])
losses_4_ge.append(test_values["loss_4"])
losses_9_ge.append(test_values["loss_9"])
logged_iterations.append(iteration)
print("# {:05d}, T {:.1f}, Ltr {:.4f}, Lge4 {:.4f}, Lge9 {:.4f}".format(
iteration, elapsed, train_values["loss"], test_values["loss_4"],
test_values["loss_9"]))
# + cellView="form" colab={} colab_type="code" id="j1FgiIm-pmRq"
#@title Visualize loss curves { form-width: "30%" }
# This cell visualizes the results of training. You can visualize the
# intermediate results by interrupting execution of the cell above, and running
# this cell. You can then resume training by simply executing the above cell
# again.
def get_node_trajectories(rollout_array, batch_size): # pylint: disable=redefined-outer-name
return np.split(rollout_array[..., :2], batch_size, axis=1)
fig = plt.figure(1, figsize=(18, 3))
fig.clf()
x = np.array(logged_iterations)
# Next-step Loss.
y = losses_tr
ax = fig.add_subplot(1, 3, 1)
ax.plot(x, y, "k")
ax.set_title("Next step loss")
# Rollout 5 loss.
y = losses_4_ge
ax = fig.add_subplot(1, 3, 2)
ax.plot(x, y, "k")
ax.set_title("Rollout loss: 5-mass string")
# Rollout 9 loss.
y = losses_9_ge
ax = fig.add_subplot(1, 3, 3)
ax.plot(x, y, "k")
ax.set_title("Rollout loss: 9-mass string")
# Visualize trajectories.
true_rollouts_4 = get_node_trajectories(test_values["true_rollout_4"],
batch_size_ge)
predicted_rollouts_4 = get_node_trajectories(test_values["predicted_rollout_4"],
batch_size_ge)
true_rollouts_9 = get_node_trajectories(test_values["true_rollout_9"],
batch_size_ge)
predicted_rollouts_9 = get_node_trajectories(test_values["predicted_rollout_9"],
batch_size_ge)
true_rollouts = true_rollouts_4
predicted_rollouts = predicted_rollouts_4
true_rollouts = true_rollouts_9
predicted_rollouts = predicted_rollouts_9
num_graphs = len(true_rollouts)
num_time_steps = true_rollouts[0].shape[0]
# Plot state sequences.
max_graphs_to_plot = 1
num_graphs_to_plot = min(num_graphs, max_graphs_to_plot)
num_steps_to_plot = 24
max_time_step = num_time_steps - 1
step_indices = np.floor(np.linspace(0, max_time_step,
num_steps_to_plot)).astype(int).tolist()
w = 6
h = int(np.ceil(num_steps_to_plot / w))
fig = plt.figure(101, figsize=(18, 8))
fig.clf()
for i, (true_rollout, predicted_rollout) in enumerate(
zip(true_rollouts, predicted_rollouts)):
xys = np.hstack([predicted_rollout, true_rollout]).reshape([-1, 2])
xs = xys[:, 0]
ys = xys[:, 1]
b = 0.05
xmin = xs.min() - b * xs.ptp()
xmax = xs.max() + b * xs.ptp()
ymin = ys.min() - b * ys.ptp()
ymax = ys.max() + b * ys.ptp()
if i >= num_graphs_to_plot:
break
for j, step_index in enumerate(step_indices):
iax = i * w + j + 1
ax = fig.add_subplot(h, w, iax)
ax.plot(
true_rollout[step_index, :, 0],
true_rollout[step_index, :, 1],
"k",
label="True")
ax.plot(
predicted_rollout[step_index, :, 0],
predicted_rollout[step_index, :, 1],
"r",
label="Predicted")
ax.set_title("Example {:02d}: frame {:03d}".format(i, step_index))
ax.set_xlim(xmin, xmax)
ax.set_ylim(ymin, ymax)
ax.set_xticks([])
ax.set_yticks([])
if j == 0:
ax.legend(loc=3)
# Plot x and y trajectories over time.
max_graphs_to_plot = 3
num_graphs_to_plot = min(len(true_rollouts), max_graphs_to_plot)
w = 2
h = num_graphs_to_plot
fig = plt.figure(102, figsize=(18, 12))
fig.clf()
for i, (true_rollout, predicted_rollout) in enumerate(
zip(true_rollouts, predicted_rollouts)):
if i >= num_graphs_to_plot:
break
t = np.arange(num_time_steps)
for j in range(2):
coord_string = "x" if j == 0 else "y"
iax = i * 2 + j + 1
ax = fig.add_subplot(h, w, iax)
ax.plot(t, true_rollout[..., j], "k", label="True")
ax.plot(t, predicted_rollout[..., j], "r", label="Predicted")
ax.set_xlabel("Time")
ax.set_ylabel("{} coordinate".format(coord_string))
ax.set_title("Example {:02d}: Predicted vs actual coords over time".format(
i))
ax.set_frame_on(False)
if i == 0 and j == 1:
handles, labels = ax.get_legend_handles_labels()
unique_labels = []
unique_handles = []
for i, (handle, label) in enumerate(zip(handles, labels)): # pylint: disable=redefined-outer-name
if label not in unique_labels:
unique_labels.append(label)
unique_handles.append(handle)
ax.legend(unique_handles, unique_labels, loc=3)
# -
| graph_nets/demos/physics-Copy2-changescene.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 资产组合构建
# ## 本段代码利用quantOS系统进行资产分析及组合构建,并通过比较沪深300指数波动率及其成分
# ## 股波动率中位数,说明组合的作用
# ## 1. 根据资产性质组合中债指数,沪深300全收益指数以及阿尔法指数
# ## 2. 比较沪深300指数波动率及成份股波动率中位数
# ## 系统设置
# +
# encoding: utf-8
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import datetime
import seaborn as sns
import matplotlib.mlab as mlab
import scipy.stats as stats
sns.set_style('darkgrid')
sns.set_context('poster')
# %matplotlib inline
# +
from jaqs.data import RemoteDataService
import jaqs.util as jutil
from __future__ import print_function, unicode_literals, division, absolute_import
from jaqs.data import RemoteDataService, DataView
# +
dataview_dir_path = '.'
backtest_result_dir_path = '.'
import os
phone = os.environ.get('QUANTOS_USER')
token = os.environ.get('QUANTOS_TOKEN')
data_config = {
"remote.data.address": "tcp://data.quantos.org:8910",
"remote.data.username": phone,
"timeout": 3600,
"remote.data.password": <PASSWORD>
}
ds = RemoteDataService()
ds.init_from_config(data_config)
# -
# ## 1. 根据资产性质进行大类资产配置
# ## 导入收益率数据文件
df = pd.read_csv('data.csv')
df = df.set_index('date')
df.index = [pd.Timestamp(str(x)) for x in df.index]
print('三个标的年化波动率为(%):')
print(np.std(df[['bond', 'alpha', '300']]) * np.sqrt(242) * 100)
# ## 设置权重,并计算每日组合收益率
annual_std = np.std(df[['bond', 'alpha', '300']]) * np.sqrt(242) * 100
weight = np.array(1.0/np.sum(1.0/annual_std)*(1.0/annual_std))
weight = np.array([0.6, 0.2,0.2])
df['Combined'] = df.multiply(weight, axis = 1).sum(axis = 1)
# ## 计算每个组合累积收益率
df['cum_bond'] = (1 + df['bond']).cumprod()
df['cum_alpha'] =(1 + df['alpha']).cumprod()
df['cum_hs300'] = (1 + df['300']).cumprod()
df['cum_weighted'] = (1 + df['Combined']).cumprod()
# df *= 100
fig, ax = plt.subplots(figsize = (16,8))
plt.plot(df.index, df.cum_bond, lw = 1, color = 'red', label = 'Bond')
plt.plot(df.index, df.cum_alpha, lw = 1, color = 'blue', label = 'Alpha')
plt.plot(df.index, df.cum_hs300, lw = 1, color = 'green', label = 'Hs300')
plt.legend()
fig, ax = plt.subplots(figsize = (16,8))
plt.plot(df.index, df.cum_weighted, lw = 1, color = 'purple', label = 'Combined')
plt.legend()
# ## 计算各标的收益率指标
print('各标的年化收益率为(%):')
print(np.mean(df[['bond', 'alpha', '300', 'Combined']]) * 242 * 100)
print('各个标的年化波动率为(%):')
print(np.std(df[['bond', 'alpha', '300', 'Combined']]) * np.sqrt(242) * 100)
risk_free_rate = 0.03 / 242
print('各个标的年化Sharpe Ratio为(%):')
print(np.mean(df[['bond', 'alpha', '300', 'Combined']] - risk_free_rate)/np.std(df[['bond', 'alpha', '300', 'Combined']]) * np.sqrt(242))
# ## 计算correlation
df[['bond', 'alpha', '300', 'Combined']].corr()
# # 2. 沪深300成份股年化波动率
START_DATE, END_DATE = 20160101, 20180328
UNIVERSE = '000300.SH'
# +
dataview_props = {# Start and end date of back-test
'start_date': START_DATE, 'end_date': END_DATE,
# Investment universe and performance benchmark
'universe': UNIVERSE, 'benchmark': UNIVERSE,
# Data fields that we need
'fields': 'free_share,total_share,float_share,float_mv,sw1',
# freq = 1 means we use daily data. Please do not change this.
'freq': 1}
# DataView utilizes RemoteDataService to get various data and store them
dv = DataView()
dv.init_from_config(dataview_props, ds)
dv.prepare_data()
# -
# ## 计算个股日收益率及相应波动率
dv.add_formula('daily_ret', 'Return(close_adj, 1)', is_quarterly = False)
daily_ret = dv.get_ts('daily_ret')
daily_std = np.std(daily_ret, axis = 0) * np.sqrt(242)
daily_std = daily_std.dropna()
print('沪深300成份股年化波动率中位数为: %.3f' % np.median(daily_std))
print('沪深300成份股年化波动率0.25分位数为: %.3f' % np.percentile(daily_std, 25))
print('沪深300成份股年化波动率0.75分位数为: %.3f' % np.percentile(daily_std, 75))
# ## 计算指数日收益率及相应波动率
benchmark_ret = (dv.data_benchmark.diff()/dv.data_benchmark.shift(1)).dropna().values
benchmark_std = np.std(benchmark_ret) * np.sqrt(242)
print('沪深300指数年化波动率为: %.3f' % benchmark_std)
# +
fig = plt.figure(figsize=(18,18))
v = daily_std.values
avgRet = np.median(v)
pct_25 = np.percentile(daily_std, 25)
pct_75 = np.percentile(daily_std, 75)
stdRet = np.std(v)
x = np.linspace(avgRet - 3*stdRet, avgRet + 3*stdRet, 100)
y = mlab.normpdf(x, avgRet, stdRet)
kde = stats.gaussian_kde(v)
# plot the histogram
plt.subplot(211)
plt.hist(v, 50, weights = np.ones(len(v))/len(v), alpha = 0.4)
plt.axvline(x = benchmark_std, color = 'red', linestyle = '--', linewidth = 0.8, label = '300 Index Std')
plt.axvline(x = avgRet, color = 'grey', linestyle = '--', linewidth = 0.8, label = 'Median Stock Std')
plt.axvline(x = pct_25, color = 'blue', linestyle = '--', linewidth = 0.8, label = '25% Std')
plt.axvline(x = pct_75, color = 'blue', linestyle = '--', linewidth = 0.8, label = '75% Std')
plt.ylabel('Percentage', fontsize = 10)
plt.legend(fontsize = 12)
# -
| example/WallStreetLectures/ipython/lecture14_portfolio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from nltk.classify import NaiveBayesClassifier
from nltk.sentiment import SentimentAnalyzer
from nltk.sentiment.util import *
from nltk.sentiment.vader import SentimentIntensityAnalyzer
from nltk import tokenize
import pandas as pd
# -
DATASET_PATH = '../datasets/CHUNK_RC_2018-02-28'
SAVE_CLASSIFIER = '../datasets/nltk_sentiment_NBClassifier'
df = pd.read_json(DATASET_PATH, lines=True, chunksize=1e4).read()
df = df[(df.body != '[deleted]') & (df.body != '[removed]')]
# +
all_comments = [(tokenize.word_tokenize(comment.body)+[str(comment.score)],
comment.controversiality) for comment in df.itertuples()]
no_contro = [comment for comment in all_comments if not comment[1]]
contro = [comment for comment in all_comments if comment[1]]
print(len(no_contro), len(contro))
# -
training, testing = split_train_test(all_comments)
sentim_analyzer = SentimentAnalyzer()
all_words_neg = sentim_analyzer.all_words([mark_negation(doc) for doc in training])
unigram_feats = sentim_analyzer.unigram_word_feats(all_words_neg, min_freq=4)
sentim_analyzer.add_feat_extractor(extract_unigram_feats, unigrams=unigram_feats)
# +
def vader_feats(document, analyzer):
# document is list of words, where last element is score
vs = analyzer.polarity_scores(' '.join(document[:-1]))
return {k+'_VAD': v for k, v in vs.items()}
sentim_analyzer.add_feat_extractor(vader_feats, analyzer=SentimentIntensityAnalyzer())
# +
def score_feat(document):
return {'_score': document[-1]}
sentim_analyzer.add_feat_extractor(score_feat)
# -
training_set = sentim_analyzer.apply_features(training)
test_set = sentim_analyzer.apply_features(testing)
trainer = NaiveBayesClassifier.train
classifier = sentim_analyzer.train(trainer, training_set)
evaluate = sentim_analyzer.evaluate(test_set, verbose=True)
# +
from sklearn.utils import resample
# downsampling to fix imbalance
no_contro_down = resample(no_contro, replace=False, n_samples=len(contro), random_state=123456)
train_down, test_down = split_train_test(no_contro_down + contro)
sa_down = SentimentAnalyzer()
allneg_down = sa_down.all_words([mark_negation(doc) for doc in train_down])
unifeats_down = sa_down.unigram_word_feats(allneg_down, min_freq=4)
sa_down.add_feat_extractor(extract_unigram_feats, unigrams=unifeats_down)
sa_down.add_feat_extractor(vader_feats, analyzer=SentimentIntensityAnalyzer())
sa_down.add_feat_extractor(score_feat)
trainset_down = sa_down.apply_features(train_down)
testset_down = sa_down.apply_features(test_down)
classifier_down = sa_down.train(trainer, trainset_down, save_classifier=SAVE_CLASSIFIER)
evaluate_down = sa_down.evaluate(testset_down, verbose=True)
| notebooks/nltk_sentiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from fredapi import Fred
import numpy as np
import requests
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
# -
fred = Fred(api_key='6f2f81fce97aa422c54836a13d08ebe2')
series = ['SP500','GDP']
data = [fred.get_series(x) for x in series]
for i, df in enumerate(data):
print(df.count())
df.plot(title=series[i])
plt.show()
# +
#endpoint = '/repos/:owner/:repo/actions/secrets/public-key'
#requests.get(endpoint)
fred = Fred(api_key='6f2f81fce97aa422c54836a13d08ebe2')#,response_type='df')
series = ['SP500','FEDFUNDS','FGEXPND']
data = [fred.get_series(x) for x in series]
df = pd.DataFrame()
for i, data_series in enumerate(data):
#if series[i] = 'FGEXPEND':
info_list = fred.search(series[i])[['title','units_short']]
print(info_list)
#print(data_series.title)
df[series[i]+' ('+str(info_list['units_short'][0])+')'] = data_series
#print(df.head())
df_filt = df.fillna(method="ffill")
df_filt.rename(columns={'SP500 (Index)':'SP500 ($ /share)'}, inplace=True)
print(df_filt.dropna(how='any').count())
df_filt.dropna(how='any').head()
df_filt.plot()
# +
ax = df_filt.plot(secondary_y='FEDFUNDS (%)')
ax2 = ax.twinx()
#ax2.set_yscale('log')
plt.show()
# -
fred.search('corporate tax rate')#['id','']
# +
df = pd.DataFrame({'x': [1,2,0,4], 'y': [1e1, 5e3, 3e2, 3e4]})
ax = df_filt.plot(secondary_y='FEDFUNDS (%)')
ax2 = ax.twinx()
#ax2.set_yscale('log')
plt.show()
# -
info_list['units_short'][1]
| notebooks/0.1.0-whs-FRED_API_Testt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 小批量随机梯度下降
# :label:`sec_minibatch_sgd`
#
# 到目前为止,我们在基于梯度的学习方法中遇到了两个极端情况:
# :numref:`sec_gd`中使用完整数据集来计算梯度并更新参数,
# :numref:`sec_sgd`中一次处理一个训练样本来取得进展。
# 二者各有利弊:每当数据非常相似时,梯度下降并不是非常“数据高效”。
# 而由于CPU和GPU无法充分利用向量化,随机梯度下降并不特别“计算高效”。
# 这暗示了两者之间可能有折中方案,这便涉及到*小批量随机梯度下降*(minibatch gradient descent)。
#
# ## 向量化和缓存
#
# 使用小批量的决策的核心是计算效率。
# 当考虑与多个GPU和多台服务器并行处理时,这一点最容易被理解。在这种情况下,我们需要向每个GPU发送至少一张图像。
# 有了每台服务器8个GPU和16台服务器,我们就能得到大小为128的小批量。
#
# 当涉及到单个GPU甚至CPU时,事情会更微妙一些:
# 这些设备有多种类型的内存、通常情况下多种类型的计算单元以及在它们之间不同的带宽限制。
# 例如,一个CPU有少量寄存器(register),L1和L2缓存,以及L3缓存(在不同的处理器内核之间共享)。
# 随着缓存的大小的增加,它们的延迟也在增加,同时带宽在减少。
# 可以说,处理器能够执行的操作远比主内存接口所能提供的多得多。
#
# 首先,具有16个内核和AVX-512向量化的2GHz CPU每秒可处理高达$2 \cdot 10^9 \cdot 16 \cdot 32 = 10^{12}$个字节。
# 同时,GPU的性能很容易超过该数字100倍。
# 而另一方面,中端服务器处理器的带宽可能不超过100Gb/s,即不到处理器满负荷所需的十分之一。
# 更糟糕的是,并非所有的内存入口都是相等的:内存接口通常为64位或更宽(例如,在最多384位的GPU上)。
# 因此读取单个字节会导致由于更宽的存取而产生的代价。
#
# 其次,第一次存取的额外开销很大,而按序存取(sequential access)或突发读取(burst read)相对开销较小。
# 有关更深入的讨论,请参阅此[维基百科文章](https://en.wikipedia.org/wiki/Cache_hierarchy)。
#
# 减轻这些限制的方法是使用足够快的CPU缓存层次结构来为处理器提供数据。
# 这是深度学习中批量处理背后的推动力。
# 举一个简单的例子:矩阵-矩阵乘法。
# 比如$\mathbf{A} = \mathbf{B}\mathbf{C}$,我们有很多方法来计算$\mathbf{A}$。例如,我们可以尝试以下方法:
#
# 1. 我们可以计算$\mathbf{A}_{ij} = \mathbf{B}_{i,:} \mathbf{C}_{:,j}^\top$,也就是说,我们可以通过点积进行逐元素计算。
# 1. 我们可以计算$\mathbf{A}_{:,j} = \mathbf{B} \mathbf{C}_{:,j}^\top$,也就是说,我们可以一次计算一列。同样,我们可以一次计算$\mathbf{A}$一行$\mathbf{A}_{i,:}$。
# 1. 我们可以简单地计算$\mathbf{A} = \mathbf{B} \mathbf{C}$。
# 1. 我们可以将$\mathbf{B}$和$\mathbf{C}$分成较小的区块矩阵,然后一次计算$\mathbf{A}$的一个区块。
#
# 如果我们使用第一个选择,每次我们计算一个元素$\mathbf{A}_{ij}$时,都需要将一行和一列向量复制到CPU中。
# 更糟糕的是,由于矩阵元素是按顺序对齐的,因此当从内存中读取它们时,我们需要访问两个向量中许多不相交的位置。
# 第二种选择相对更有利:我们能够在遍历$\mathbf{B}$的同时,将列向量$\mathbf{C}_{:,j}$保留在CPU缓存中。
# 它将内存带宽需求减半,相应地提高了访问速度。
# 第三种选择表面上是最可取的,然而大多数矩阵可能不能完全放入缓存中。
# 第四种选择提供了一个实践上很有用的方案:我们可以将矩阵的区块移到缓存中然后在本地将它们相乘。
# 让我们来看看这些操作在实践中的效率如何。
#
# 除了计算效率之外,Python和深度学习框架本身带来的额外开销也是相当大的。
# 回想一下,每次我们执行代码时,Python解释器都会向深度学习框架发送一个命令,要求将其插入到计算图中并在调度过程中处理它。
# 这样的额外开销可能是非常不利的。
# 总而言之,我们最好用向量化(和矩阵)。
#
# + origin_pos=2 tab=["pytorch"]
# %matplotlib inline
import numpy as np
import torch
from torch import nn
from d2l import torch as d2l
timer = d2l.Timer()
A = torch.zeros(256, 256)
B = torch.randn(256, 256)
C = torch.randn(256, 256)
# + [markdown] origin_pos=4
# 按元素分配只需遍历分别为$\mathbf{B}$和$\mathbf{C}$的所有行和列,即可将该值分配给$\mathbf{A}$。
#
# + origin_pos=6 tab=["pytorch"]
# 逐元素计算A=BC
timer.start()
for i in range(256):
for j in range(256):
A[i, j] = torch.dot(B[i, :], C[:, j])
timer.stop()
# + [markdown] origin_pos=8
# 更快的策略是执行按列分配。
#
# + origin_pos=10 tab=["pytorch"]
# 逐列计算A=BC
timer.start()
for j in range(256):
A[:, j] = torch.mv(B, C[:, j])
timer.stop()
# + [markdown] origin_pos=12
# 最有效的方法是在一个区块中执行整个操作。让我们看看它们各自的操作速度是多少。
#
# + origin_pos=14 tab=["pytorch"]
# 一次性计算A=BC
timer.start()
A = torch.mm(B, C)
timer.stop()
# 乘法和加法作为单独的操作(在实践中融合)
gigaflops = [2/i for i in timer.times]
print(f'performance in Gigaflops: element {gigaflops[0]:.3f}, '
f'column {gigaflops[1]:.3f}, full {gigaflops[2]:.3f}')
# + [markdown] origin_pos=16
# ## 小批量
#
# :label:`sec_minibatches`
#
# 之前我们会理所当然地读取数据的*小批量*,而不是观测单个数据来更新参数,现在简要解释一下原因。
# 处理单个观测值需要我们执行许多单一矩阵-矢量(甚至矢量-矢量)乘法,这耗费相当大,而且对应深度学习框架也要巨大的开销。
# 这既适用于计算梯度以更新参数时,也适用于用神经网络预测。
# 也就是说,每当我们执行$\mathbf{w} \leftarrow \mathbf{w} - \eta_t \mathbf{g}_t$时,消耗巨大。其中
#
# $$\mathbf{g}_t = \partial_{\mathbf{w}} f(\mathbf{x}_{t}, \mathbf{w}).$$
#
# 我们可以通过将其应用于一个小批量观测值来提高此操作的*计算*效率。
# 也就是说,我们将梯度$\mathbf{g}_t$替换为一个小批量而不是单个观测值
#
# $$\mathbf{g}_t = \partial_{\mathbf{w}} \frac{1}{|\mathcal{B}_t|} \sum_{i \in \mathcal{B}_t} f(\mathbf{x}_{i}, \mathbf{w}).$$
#
# 让我们看看这对$\mathbf{g}_t$的统计属性有什么影响:由于$\mathbf{x}_t$和小批量$\mathcal{B}_t$的所有元素都是从训练集中随机抽出的,因此梯度的期望保持不变。
# 另一方面,方差显著降低。
# 由于小批量梯度由正在被平均计算的$b := |\mathcal{B}_t|$个独立梯度组成,其标准差降低了$b^{-\frac{1}{2}}$。
# 这本身就是一件好事,因为这意味着更新与完整的梯度更接近了。
#
# 直观来说,这表明选择大型的小批量$\mathcal{B}_t$将是普遍可行的。
# 然而,经过一段时间后,与计算代价的线性增长相比,标准差的额外减少是微乎其微的。
# 在实践中我们选择一个足够大的小批量,它可以提供良好的计算效率同时仍适合GPU的内存。
# 下面,我们来看看这些高效的代码。
# 在里面我们执行相同的矩阵-矩阵乘法,但是这次我们将其一次性分为64列的“小批量”。
#
# + origin_pos=18 tab=["pytorch"]
timer.start()
for j in range(0, 256, 64):
A[:, j:j+64] = torch.mm(B, C[:, j:j+64])
timer.stop()
print(f'performance in Gigaflops: block {2 / timer.times[3]:.3f}')
# + [markdown] origin_pos=20
# 显而易见,小批量上的计算基本上与完整矩阵一样有效。
# 需要注意的是,在 :numref:`sec_batch_norm`中,我们使用了一种在很大程度上取决于小批量中的方差的正则化。
# 随着后者增加,方差会减少,随之而来的是批量规范化带来的噪声注入的好处。
# 关于实例,请参阅 :cite:`Ioffe.2017`,了解有关如何重新缩放并计算适当项目。
#
# ## 读取数据集
#
# 让我们来看看如何从数据中有效地生成小批量。
# 下面我们使用NASA开发的测试机翼的数据集[不同飞行器产生的噪声](https://archive.ics.uci.edu/ml/datasets/Airfoil+Self-Noise)来比较这些优化算法。
# 为方便起见,我们只使用前$1,500$样本。
# 数据已作预处理:我们移除了均值并将方差重新缩放到每个坐标为$1$。
#
# + origin_pos=22 tab=["pytorch"]
#@save
d2l.DATA_HUB['airfoil'] = (d2l.DATA_URL + 'airfoil_self_noise.dat',
'76e5be1548fd8222e5074cf0faae75edff8cf93f')
#@save
def get_data_ch11(batch_size=10, n=1500):
data = np.genfromtxt(d2l.download('airfoil'),
dtype=np.float32, delimiter='\t')
data = torch.from_numpy((data - data.mean(axis=0)) / data.std(axis=0))
data_iter = d2l.load_array((data[:n, :-1], data[:n, -1]),
batch_size, is_train=True)
return data_iter, data.shape[1]-1
# + [markdown] origin_pos=24
# ## 从零开始实现
#
# :numref:`sec_linear_scratch`一节中已经实现过小批量随机梯度下降算法。
# 我们在这里将它的输入参数变得更加通用,主要是为了方便本章后面介绍的其他优化算法也可以使用同样的输入。
# 具体来说,我们添加了一个状态输入`states`并将超参数放在字典`hyperparams`中。
# 此外,我们将在训练函数里对各个小批量样本的损失求平均,因此优化算法中的梯度不需要除以批量大小。
#
# + origin_pos=26 tab=["pytorch"]
def sgd(params, states, hyperparams):
for p in params:
p.data.sub_(hyperparams['lr'] * p.grad)
p.grad.data.zero_()
# + [markdown] origin_pos=28
# 下面实现一个通用的训练函数,以方便本章后面介绍的其他优化算法使用。
# 它初始化了一个线性回归模型,然后可以使用小批量随机梯度下降以及后续小节介绍的其他算法来训练模型。
#
# + origin_pos=30 tab=["pytorch"]
#@save
def train_ch11(trainer_fn, states, hyperparams, data_iter,
feature_dim, num_epochs=2):
# 初始化模型
w = torch.normal(mean=0.0, std=0.01, size=(feature_dim, 1),
requires_grad=True)
b = torch.zeros((1), requires_grad=True)
net, loss = lambda X: d2l.linreg(X, w, b), d2l.squared_loss
# 训练模型
animator = d2l.Animator(xlabel='epoch', ylabel='loss',
xlim=[0, num_epochs], ylim=[0.22, 0.35])
n, timer = 0, d2l.Timer()
for _ in range(num_epochs):
for X, y in data_iter:
l = loss(net(X), y).mean()
l.backward()
trainer_fn([w, b], states, hyperparams)
n += X.shape[0]
if n % 200 == 0:
timer.stop()
animator.add(n/X.shape[0]/len(data_iter),
(d2l.evaluate_loss(net, data_iter, loss),))
timer.start()
print(f'loss: {animator.Y[0][-1]:.3f}, {timer.avg():.3f} sec/epoch')
return timer.cumsum(), animator.Y[0]
# + [markdown] origin_pos=32
# 让我们来看看批量梯度下降的优化是如何进行的。
# 这可以通过将小批量设置为1500(即样本总数)来实现。
# 因此,模型参数每个迭代轮数只迭代一次。
#
# + origin_pos=33 tab=["pytorch"]
def train_sgd(lr, batch_size, num_epochs=2):
data_iter, feature_dim = get_data_ch11(batch_size)
return train_ch11(
sgd, None, {'lr': lr}, data_iter, feature_dim, num_epochs)
gd_res = train_sgd(1, 1500, 10)
# + [markdown] origin_pos=34
# 当批量大小为1时,优化使用的是随机梯度下降。
# 为了简化实现,我们选择了很小的学习率。
# 在随机梯度下降的实验中,每当一个样本被处理,模型参数都会更新。
# 在这个例子中,这相当于每个迭代轮数有1500次更新。
# 可以看到,目标函数值的下降在1个迭代轮数后就变得较为平缓。
# 尽管两个例子在一个迭代轮数内都处理了1500个样本,但实验中随机梯度下降的一个迭代轮数耗时更多。
# 这是因为随机梯度下降更频繁地更新了参数,而且一次处理单个观测值效率较低。
#
# + origin_pos=35 tab=["pytorch"]
sgd_res = train_sgd(0.005, 1)
# + [markdown] origin_pos=36
# 最后,当批量大小等于100时,我们使用小批量随机梯度下降进行优化。
# 每个迭代轮数所需的时间比随机梯度下降和批量梯度下降所需的时间短。
#
# + origin_pos=37 tab=["pytorch"]
mini1_res = train_sgd(.4, 100)
# + [markdown] origin_pos=38
# 将批量大小减少到10,每个迭代轮数的时间都会增加,因为每批工作负载的执行效率变得更低。
#
# + origin_pos=39 tab=["pytorch"]
mini2_res = train_sgd(.05, 10)
# + [markdown] origin_pos=40
# 现在我们可以比较前四个实验的时间与损失。
# 可以看出,尽管在处理的样本数方面,随机梯度下降的收敛速度快于梯度下降,但与梯度下降相比,它需要更多的时间来达到同样的损失,因为逐个样本来计算梯度并不那么有效。
# 小批量随机梯度下降能够平衡收敛速度和计算效率。
# 大小为10的小批量比随机梯度下降更有效;
# 大小为100的小批量在运行时间上甚至优于梯度下降。
#
# + origin_pos=41 tab=["pytorch"]
d2l.set_figsize([6, 3])
d2l.plot(*list(map(list, zip(gd_res, sgd_res, mini1_res, mini2_res))),
'time (sec)', 'loss', xlim=[1e-2, 10],
legend=['gd', 'sgd', 'batch size=100', 'batch size=10'])
d2l.plt.gca().set_xscale('log')
# + [markdown] origin_pos=42
# ## 简洁实现
#
# 下面用深度学习框架自带算法实现一个通用的训练函数,我们将在本章中其它小节使用它。
#
# + origin_pos=44 tab=["pytorch"]
#@save
def train_concise_ch11(trainer_fn, hyperparams, data_iter, num_epochs=4):
# 初始化模型
net = nn.Sequential(nn.Linear(5, 1))
def init_weights(m):
if type(m) == nn.Linear:
torch.nn.init.normal_(m.weight, std=0.01)
net.apply(init_weights)
optimizer = trainer_fn(net.parameters(), **hyperparams)
loss = nn.MSELoss(reduction='none')
animator = d2l.Animator(xlabel='epoch', ylabel='loss',
xlim=[0, num_epochs], ylim=[0.22, 0.35])
n, timer = 0, d2l.Timer()
for _ in range(num_epochs):
for X, y in data_iter:
optimizer.zero_grad()
out = net(X)
y = y.reshape(out.shape)
l = loss(out, y)
l.mean().backward()
optimizer.step()
n += X.shape[0]
if n % 200 == 0:
timer.stop()
# MSELoss计算平方误差时不带系数1/2
animator.add(n/X.shape[0]/len(data_iter),
(d2l.evaluate_loss(net, data_iter, loss) / 2,))
timer.start()
print(f'loss: {animator.Y[0][-1]:.3f}, {timer.avg():.3f} sec/epoch')
# + [markdown] origin_pos=46
# 下面使用这个训练函数,复现之前的实验。
#
# + origin_pos=48 tab=["pytorch"]
data_iter, _ = get_data_ch11(10)
trainer = torch.optim.SGD
train_concise_ch11(trainer, {'lr': 0.01}, data_iter)
# + [markdown] origin_pos=50
# ## 小结
#
# * 由于减少了深度学习框架的额外开销,使用更好的内存定位以及CPU和GPU上的缓存,向量化使代码更加高效。
# * 随机梯度下降的“统计效率”与大批量一次处理数据的“计算效率”之间存在权衡。小批量随机梯度下降提供了两全其美的答案:计算和统计效率。
# * 在小批量随机梯度下降中,我们处理通过训练数据的随机排列获得的批量数据(即每个观测值只处理一次,但按随机顺序)。
# * 在训练期间降低学习率有助于训练。
# * 一般来说,小批量随机梯度下降比随机梯度下降和梯度下降的速度快,收敛风险较小。
#
# ## 练习
#
# 1. 修改批量大小和学习率,并观察目标函数值的下降率以及每个迭代轮数消耗的时间。
# 1. 将小批量随机梯度下降与实际从训练集中*取样替换*的变体进行比较。会看出什么?
# 1. 一个邪恶的精灵在没通知你的情况下复制了你的数据集(即每个观测发生两次,你的数据集增加到原始大小的两倍,但没有人告诉你)。随机梯度下降、小批量随机梯度下降和梯度下降的表现将如何变化?
#
# + [markdown] origin_pos=52 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/4325)
#
| pytorch/chapter_optimization/minibatch-sgd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="kkAvSg_-4dPz"
# <div align="center">
# <h1><img width="30" src="https://madewithml.com/static/images/rounded_logo.png"> <a href="https://madewithml.com/">Made With ML</a></h1>
# Applied ML · MLOps · Production
# <br>
# Join 20K+ developers in learning how to responsibly <a href="https://madewithml.com/about/">deliver value</a> with ML.
# </div>
#
# <br>
#
# <div align="center">
# <a target="_blank" href="https://newsletter.madewithml.com"><img src="https://img.shields.io/badge/Subscribe-20K-brightgreen"></a>
# <a target="_blank" href="https://github.com/GokuMohandas/madewithml"><img src="https://img.shields.io/github/stars/GokuMohandas/madewithml.svg?style=social&label=Star"></a>
# <a target="_blank" href="https://www.linkedin.com/in/goku"><img src="https://img.shields.io/badge/style--5eba00.svg?label=LinkedIn&logo=linkedin&style=social"></a>
# <a target="_blank" href="https://twitter.com/GokuMohandas"><img src="https://img.shields.io/twitter/follow/GokuMohandas.svg?label=Follow&style=social"></a>
# <p>🔥 Among the <a href="https://github.com/topics/deep-learning" target="_blank">top ML</a> repositories on GitHub</p>
# </div>
#
# <br>
# <hr>
# + [markdown] id="eTdCMVl9YAXw"
# # Convolutional Neural Networks (CNN)
#
# In this lesson we will explore the basics of Convolutional Neural Networks (CNNs) applied to text for natural language processing (NLP) tasks.
# + [markdown] id="xuabAj4PYj57"
# <div align="left">
# <a target="_blank" href="https://madewithml.com/courses/ml-foundations/convolutional-neural-networks/"><img src="https://img.shields.io/badge/📖 Read-blog post-9cf"></a>
# <a href="https://github.com/GokuMohandas/madewithml/blob/main/notebooks/11_Convolutional_Neural_Networks.ipynb" role="button"><img src="https://img.shields.io/static/v1?label=&message=View%20On%20GitHub&color=586069&logo=github&labelColor=2f363d"></a>
# <a href="https://colab.research.google.com/github/GokuMohandas/madewithml/blob/main/notebooks/11_Convolutional_Neural_Networks.ipynb"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"></a>
# </div>
# + [markdown] id="Xz15aMLj7gPX"
# # Overview
# + [markdown] id="Ym2pswjn5GNw"
# At the core of CNNs are filters (aka weights, kernels, etc.) which convolve (slide) across our input to extract relevant features. The filters are initialized randomly but learn to act as feature extractors via parameter sharing.
# + [markdown] id="TPGoKi5k4j5y"
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/madewithml/main/images/ml-foundations/cnn/convolution.gif" width="500">
# </div>
# + [markdown] id="JqxyljU18hvt"
# * **Objective:** Extract meaningful spatial substructure from encoded data.
# * **Advantages:**
# * Small number of weights (shared)
# * Parallelizable
# * Detects spatial substrcutures (feature extractors)
# * [Interpretability](https://arxiv.org/abs/1312.6034) via filters
# * Can be used for processing in images, text, time-series, etc.
# * **Disadvantages:**
# * Many hyperparameters (kernel size, strides, etc.) to tune.
# * **Miscellaneous:**
# * Lot's of deep CNN architectures constantly updated for SOTA performance.
# * Very popular feature extractor that's usually prepended onto other architectures.
#
# + [markdown] id="6GLD2nXvo-r4"
# # Set up
# + id="y3qKSoEe57na"
import numpy as np
import pandas as pd
import random
import torch
import torch.nn as nn
# + id="ZMLLoU5htnxT"
SEED = 1234
# + id="xK1YEDDotQ3V"
def set_seeds(seed=1234):
"""Set seeds for reproducability."""
np.random.seed(seed)
random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.cuda.manual_seed_all(seed) # multi-GPU
# + id="XU1TUOBitpaR"
# Set seeds for reproducability
set_seeds(seed=SEED)
# + colab={"base_uri": "https://localhost:8080/"} id="86uoTigW0ylA" executionInfo={"status": "ok", "timestamp": 1608329398628, "user_tz": 420, "elapsed": 3911, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="bd4c1448-851c-4415-c8ac-09d76183fc54"
# Set device
cuda = True
device = torch.device('cuda' if (
torch.cuda.is_available() and cuda) else 'cpu')
torch.set_default_tensor_type('torch.FloatTensor')
if device.type == 'cuda':
torch.set_default_tensor_type('torch.cuda.FloatTensor')
print (device)
# + [markdown] id="c69z9wpJ56nE"
# ## Load data
# + [markdown] id="2V_nEp5G58M0"
# We will download the [AG News dataset](http://www.di.unipi.it/~gulli/AG_corpus_of_news_articles.html), which consists of 120K text samples from 4 unique classes (`Business`, `Sci/Tech`, `Sports`, `World`)
# + id="cdjdvnOGrsZP" colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"status": "ok", "timestamp": 1608329399051, "user_tz": 420, "elapsed": 4304, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="53c06734-a2a9-4928-f080-49b00e9f150c"
# Load data
url = "https://raw.githubusercontent.com/GokuMohandas/madewithml/main/datasets/news.csv"
df = pd.read_csv(url, header=0) # load
df = df.sample(frac=1).reset_index(drop=True) # shuffle
df.head()
# + [markdown] id="RQUDEgwloxhF"
# ## Preprocessing
# + [markdown] id="2QKp1TyPpBKG"
# We're going to clean up our input data first by doing operations such as lower text, removing stop (filler) words, filters using regular expressions, etc.
# + id="S-Mv_g0cowkR"
import nltk
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
import re
# + colab={"base_uri": "https://localhost:8080/"} id="K0DwdEzxownP" executionInfo={"status": "ok", "timestamp": 1608329400344, "user_tz": 420, "elapsed": 5539, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="078c0da6-c18e-416f-900f-01336eb531d1"
nltk.download('stopwords')
STOPWORDS = stopwords.words('english')
print (STOPWORDS[:5])
porter = PorterStemmer()
# + id="w1Yyrsp0owqk"
def preprocess(text, stopwords=STOPWORDS):
"""Conditional preprocessing on our text unique to our task."""
# Lower
text = text.lower()
# Remove stopwords
pattern = re.compile(r'\b(' + r'|'.join(stopwords) + r')\b\s*')
text = pattern.sub('', text)
# Remove words in paranthesis
text = re.sub(r'\([^)]*\)', '', text)
# Spacing and filters
text = re.sub(r"([-;;.,!?<=>])", r" \1 ", text)
text = re.sub('[^A-Za-z0-9]+', ' ', text) # remove non alphanumeric chars
text = re.sub(' +', ' ', text) # remove multiple spaces
text = text.strip()
return text
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="zvcSy1wSowtC" executionInfo={"status": "ok", "timestamp": 1608329400349, "user_tz": 420, "elapsed": 5482, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="9d8a1d6a-bd53-4824-b0b4-7d029a83baa2"
# Sample
text = "Great week for the NYSE!"
preprocess(text=text)
# + colab={"base_uri": "https://localhost:8080/"} id="czjQ3lrrowwh" executionInfo={"status": "ok", "timestamp": 1608329402722, "user_tz": 420, "elapsed": 7810, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="6f554009-55ba-4763-fdda-f942bc72e7f6"
# Apply to dataframe
preprocessed_df = df.copy()
preprocessed_df.title = preprocessed_df.title.apply(preprocess)
print (f"{df.title.values[0]}\n\n{preprocessed_df.title.values[0]}")
# + [markdown] id="h5ZPPgPApLL1"
# > If you have preprocessing steps like standardization, etc. that are calculated, you need to separate the training and test set first before applying those operations. This is because we cannot apply any knowledge gained from the test set accidentally (data leak) during preprocessing/training. However for global preprocessing steps like the function above where we aren't learning anything from the data itself, we can perform before splitting the data.
# + [markdown] id="zgStr_fDpMU4"
# ## Split data
# + id="287RCymQowyV"
import collections
from sklearn.model_selection import train_test_split
# + id="BU--xNveow11"
TRAIN_SIZE = 0.7
VAL_SIZE = 0.15
TEST_SIZE = 0.15
# + id="OyQJ6x0vpP7c"
def train_val_test_split(X, y, train_size):
"""Split dataset into data splits."""
X_train, X_, y_train, y_ = train_test_split(X, y, train_size=TRAIN_SIZE, stratify=y)
X_val, X_test, y_val, y_test = train_test_split(X_, y_, train_size=0.5, stratify=y_)
return X_train, X_val, X_test, y_train, y_val, y_test
# + id="KKvtQWxepP-C"
# Data
X = preprocessed_df["title"].values
y = preprocessed_df["category"].values
# + colab={"base_uri": "https://localhost:8080/"} id="bnjaba3CpQAm" executionInfo={"status": "ok", "timestamp": 1608329402732, "user_tz": 420, "elapsed": 7720, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="3ce08f5e-6be3-40cb-ab9c-1cbf28d37f05"
# Create data splits
X_train, X_val, X_test, y_train, y_val, y_test = train_val_test_split(
X=X, y=y, train_size=TRAIN_SIZE)
print (f"X_train: {X_train.shape}, y_train: {y_train.shape}")
print (f"X_val: {X_val.shape}, y_val: {y_val.shape}")
print (f"X_test: {X_test.shape}, y_test: {y_test.shape}")
print (f"Sample point: {X_train[0]} → {y_train[0]}")
# + [markdown] id="6JJBLk0rpVrX"
# ## LabelEncoder
# + [markdown] id="gxCbJ294pYfd"
# Next we'll define a `LabelEncoder` to encode our text labels into unique indices
# + id="_voCWJ41pQIH"
import itertools
# + id="p9Ya97mMp0uR"
class LabelEncoder(object):
"""Label encoder for tag labels."""
def __init__(self, class_to_index={}):
self.class_to_index = class_to_index
self.index_to_class = {v: k for k, v in self.class_to_index.items()}
self.classes = list(self.class_to_index.keys())
def __len__(self):
return len(self.class_to_index)
def __str__(self):
return f"<LabelEncoder(num_classes={len(self)})>"
def fit(self, y):
classes = np.unique(y)
for i, class_ in enumerate(classes):
self.class_to_index[class_] = i
self.index_to_class = {v: k for k, v in self.class_to_index.items()}
self.classes = list(self.class_to_index.keys())
return self
def encode(self, y):
encoded = np.zeros((len(y)), dtype=int)
for i, item in enumerate(y):
encoded[i] = self.class_to_index[item]
return encoded
def decode(self, y):
classes = []
for i, item in enumerate(y):
classes.append(self.index_to_class[item])
return classes
def save(self, fp):
with open(fp, 'w') as fp:
contents = {'class_to_index': self.class_to_index}
json.dump(contents, fp, indent=4, sort_keys=False)
@classmethod
def load(cls, fp):
with open(fp, 'r') as fp:
kwargs = json.load(fp=fp)
return cls(**kwargs)
# + colab={"base_uri": "https://localhost:8080/"} id="WIFZYOLzp0ws" executionInfo={"status": "ok", "timestamp": 1608329402988, "user_tz": 420, "elapsed": 7917, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="29255ac5-cec4-43a2-debf-a71e84cefabe"
# Encode
label_encoder = LabelEncoder()
label_encoder.fit(y_train)
NUM_CLASSES = len(label_encoder)
label_encoder.class_to_index
# + colab={"base_uri": "https://localhost:8080/"} id="MUMaapBCp0zq" executionInfo={"status": "ok", "timestamp": 1608329402990, "user_tz": 420, "elapsed": 7888, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="d3aa414a-477f-448e-e0e5-18ae41c1327b"
# Convert labels to tokens
print (f"y_train[0]: {y_train[0]}")
y_train = label_encoder.encode(y_train)
y_val = label_encoder.encode(y_val)
y_test = label_encoder.encode(y_test)
print (f"y_train[0]: {y_train[0]}")
# + colab={"base_uri": "https://localhost:8080/"} id="U2jYqr5Yp015" executionInfo={"status": "ok", "timestamp": 1608329402991, "user_tz": 420, "elapsed": 7866, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="535ccae3-a0ed-42f1-f795-7b71bdf47071"
# Class weights
counts = np.bincount(y_train)
class_weights = {i: 1.0/count for i, count in enumerate(counts)}
print (f"counts: {counts}\nweights: {class_weights}")
# + [markdown] id="8ULHakYnp7fA"
# # Tokenizer
# + [markdown] id="FXEyU6evp-4P"
# Our input data is text and we can't feed it directly to our models. So, we'll define a `Tokenizer` to convert our text input data into token indices. This means that every token (we can decide what a token is char, word, sub-word, etc.) is mapped to a unique index which allows us to represent our text as an array of indices.
# + id="DxSzCrYJpQKq"
import json
from collections import Counter
from more_itertools import take
# + id="ymD8m7Lfow4J"
class Tokenizer(object):
def __init__(self, char_level, num_tokens=None,
pad_token='<PAD>', oov_token='<UNK>',
token_to_index=None):
self.char_level = char_level
self.separator = '' if self.char_level else ' '
if num_tokens: num_tokens -= 2 # pad + unk tokens
self.num_tokens = num_tokens
self.pad_token = pad_token
self.oov_token = oov_token
if not token_to_index:
token_to_index = {pad_token: 0, oov_token: 1}
self.token_to_index = token_to_index
self.index_to_token = {v: k for k, v in self.token_to_index.items()}
def __len__(self):
return len(self.token_to_index)
def __str__(self):
return f"<Tokenizer(num_tokens={len(self)})>"
def fit_on_texts(self, texts):
if not self.char_level:
texts = [text.split(" ") for text in texts]
all_tokens = [token for text in texts for token in text]
counts = Counter(all_tokens).most_common(self.num_tokens)
self.min_token_freq = counts[-1][1]
for token, count in counts:
index = len(self)
self.token_to_index[token] = index
self.index_to_token[index] = token
return self
def texts_to_sequences(self, texts):
sequences = []
for text in texts:
if not self.char_level:
text = text.split(' ')
sequence = []
for token in text:
sequence.append(self.token_to_index.get(
token, self.token_to_index[self.oov_token]))
sequences.append(np.asarray(sequence))
return sequences
def sequences_to_texts(self, sequences):
texts = []
for sequence in sequences:
text = []
for index in sequence:
text.append(self.index_to_token.get(index, self.oov_token))
texts.append(self.separator.join([token for token in text]))
return texts
def save(self, fp):
with open(fp, 'w') as fp:
contents = {
'char_level': self.char_level,
'oov_token': self.oov_token,
'token_to_index': self.token_to_index
}
json.dump(contents, fp, indent=4, sort_keys=False)
@classmethod
def load(cls, fp):
with open(fp, 'r') as fp:
kwargs = json.load(fp=fp)
return cls(**kwargs)
# + [markdown] id="UUNjbGizqb1v"
# We're going to restrict the number of tokens in our `Tokenizer` to the top 500 most frequent tokens (stop words already removed) because the full vocabulary size (~30K) is too large to run on Google Colab notebooks.
#
# > It's important that we only fit using our train data split because during inference, our model will not always know every token so it's important to replicate that scenario with our validation and test splits as well.
# + colab={"base_uri": "https://localhost:8080/"} id="2HWkNc94qVH8" executionInfo={"status": "ok", "timestamp": 1608329403510, "user_tz": 420, "elapsed": 8326, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="b9064379-200b-477d-b617-61ae91eee41a"
# Tokenize
tokenizer = Tokenizer(char_level=False, num_tokens=500)
tokenizer.fit_on_texts(texts=X_train)
VOCAB_SIZE = len(tokenizer)
print (tokenizer)
# + colab={"base_uri": "https://localhost:8080/"} id="9x_RK5xBqVKX" executionInfo={"status": "ok", "timestamp": 1608329403512, "user_tz": 420, "elapsed": 8300, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="00d035cd-4f54-4860-fd38-431ef8f55099"
# Sample of tokens
print (take(5, tokenizer.token_to_index.items()))
print (f"least freq token's freq: {tokenizer.min_token_freq}") # use this to adjust num_tokens
# + colab={"base_uri": "https://localhost:8080/"} id="7CQYu3DFqVNR" executionInfo={"status": "ok", "timestamp": 1608329403709, "user_tz": 420, "elapsed": 8471, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="d6c6baee-dbeb-4b82-85b9-682e65eb4764"
# Convert texts to sequences of indices
X_train = tokenizer.texts_to_sequences(X_train)
X_val = tokenizer.texts_to_sequences(X_val)
X_test = tokenizer.texts_to_sequences(X_test)
preprocessed_text = tokenizer.sequences_to_texts([X_train[0]])[0]
print ("Text to indices:\n"
f" (preprocessed) → {preprocessed_text}\n"
f" (tokenized) → {X_train[0]}")
# + [markdown] id="X_BtTeojq4UU"
# # One-hot encoding
# + [markdown] id="XgB24lwpq7aA"
# One-hot encoding creates a binary column for each unique value for the feature we're trying to map. All of the values in each token's array will be 0 except at the index that this specific token is represented by.
#
# There are 5 words in the vocabulary:
# ```json
# {
# "a": 0,
# "e": 1,
# "i": 2,
# "o": 3,
# "u": 4
# }
# ```
#
# Then the text `aou` would be represented by:
# ```python
# [[1. 0. 0. 0. 0.]
# [0. 0. 0. 1. 0.]
# [0. 0. 0. 0. 1.]]
# ```
#
# One-hot encoding allows us to represent our data in a way that our models can process the data and isn't biased by the actual value of the token (ex. if your labels were actual numbers).
#
# > We have already applied one-hot encoding in the previous lessons when we encoded our labels. Each label was represented by a unique index but when determining loss, we effectively use it's one hot representation and compared it to the predicted probability distribution. We never explicitly wrote this out since all of our previous tasks were multi-class which means every input had just one output class, so the 0s didn't affect the loss (though it did matter during back propagation).
# + id="tYqX76Lpq4aK"
def to_categorical(seq, num_classes):
"""One-hot encode a sequence of tokens."""
one_hot = np.zeros((len(seq), num_classes))
for i, item in enumerate(seq):
one_hot[i, item] = 1.
return one_hot
# + colab={"base_uri": "https://localhost:8080/"} id="Nze7Wq-sq4cv" executionInfo={"status": "ok", "timestamp": 1608329403713, "user_tz": 420, "elapsed": 8433, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="cbb0ee0a-a054-4f01-d11a-f3d8b19e541b"
# One-hot encoding
print (X_train[0])
print (len(X_train[0]))
cat = to_categorical(seq=X_train[0], num_classes=len(tokenizer))
print (cat)
print (cat.shape)
# + id="FHAIYQLMq4fq"
# Convert tokens to one-hot
vocab_size = len(tokenizer)
X_train = [to_categorical(seq, num_classes=vocab_size) for seq in X_train]
X_val = [to_categorical(seq, num_classes=vocab_size) for seq in X_val]
X_test = [to_categorical(seq, num_classes=vocab_size) for seq in X_test]
# + [markdown] id="AvdiT_-6rnYF"
# # Padding
# + [markdown] id="M384IKVPrnpI"
# Our inputs are all of varying length but we need each batch to be uniformly shaped. Therefore, we will use padding to make all the inputs in the batch the same length. Our padding index will be 0 (note that this is consistent with the `<PAD>` token defined in our `Tokenizer`).
#
# > One-hot encoding creates a batch of shape (`N`, `max_seq_len`, `vocab_size`) so we'll need to be able to pad 3D sequences.
# + id="440sFfwBrnu8"
def pad_sequences(sequences, max_seq_len=0):
"""Pad sequences to max length in sequence."""
max_seq_len = max(max_seq_len, max(len(sequence) for sequence in sequences))
num_classes = sequences[0].shape[-1]
padded_sequences = np.zeros((len(sequences), max_seq_len, num_classes))
for i, sequence in enumerate(sequences):
padded_sequences[i][:len(sequence)] = sequence
return padded_sequences
# + colab={"base_uri": "https://localhost:8080/"} id="NeQoMsyUrnyD" executionInfo={"status": "ok", "timestamp": 1608329405580, "user_tz": 420, "elapsed": 10247, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="8aac87eb-f021-4774-8fa3-0f93747d881a"
# 3D sequences
print (X_train[0].shape, X_train[1].shape, X_train[2].shape)
padded = pad_sequences(X_train[0:3])
print (padded.shape)
# + [markdown] id="ii6RhseQsMKD"
# # Dataset
# + [markdown] id="Kd8PKjN8soc5"
# We're going to place our data into a [`Dataset`](https://pytorch.org/docs/stable/data.html#torch.utils.data.Dataset) and use a [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) to efficiently create batches for training and evaluation.
# + id="hNOtsJAgwv_4"
FILTER_SIZE = 1 # unigram
# + id="VMS8vwTqrn3O"
class Dataset(torch.utils.data.Dataset):
def __init__(self, X, y, max_filter_size):
self.X = X
self.y = y
self.max_filter_size = max_filter_size
def __len__(self):
return len(self.y)
def __str__(self):
return f"<Dataset(N={len(self)})>"
def __getitem__(self, index):
X = self.X[index]
y = self.y[index]
return [X, y]
def collate_fn(self, batch):
"""Processing on a batch."""
# Get inputs
batch = np.array(batch, dtype=object)
X = batch[:, 0]
y = np.stack(batch[:, 1], axis=0)
# Pad sequences
X = pad_sequences(X, max_seq_len=self.max_filter_size)
# Cast
X = torch.FloatTensor(X.astype(np.int32))
y = torch.LongTensor(y.astype(np.int32))
return X, y
def create_dataloader(self, batch_size, shuffle=False, drop_last=False):
return torch.utils.data.DataLoader(
dataset=self, batch_size=batch_size, collate_fn=self.collate_fn,
shuffle=shuffle, drop_last=drop_last, pin_memory=True)
# + colab={"base_uri": "https://localhost:8080/"} id="tXHORNGWrn6L" executionInfo={"status": "ok", "timestamp": 1608329405586, "user_tz": 420, "elapsed": 10188, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="ae6d9ea0-7992-4bb3-e5ee-eeb1184f27af"
# Create datasets for embedding
train_dataset = Dataset(X=X_train, y=y_train, max_filter_size=FILTER_SIZE)
val_dataset = Dataset(X=X_val, y=y_val, max_filter_size=FILTER_SIZE)
test_dataset = Dataset(X=X_test, y=y_test, max_filter_size=FILTER_SIZE)
print ("Datasets:\n"
f" Train dataset:{train_dataset.__str__()}\n"
f" Val dataset: {val_dataset.__str__()}\n"
f" Test dataset: {test_dataset.__str__()}\n"
"Sample point:\n"
f" X: {test_dataset[0][0]}\n"
f" y: {test_dataset[0][1]}")
# + colab={"base_uri": "https://localhost:8080/"} id="77K7kzitqwdX" executionInfo={"status": "ok", "timestamp": 1608329416454, "user_tz": 420, "elapsed": 21026, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="fab54836-13df-49ac-d93f-dbd04ee37245"
# Create dataloaders
batch_size = 64
train_dataloader = train_dataset.create_dataloader(batch_size=batch_size)
val_dataloader = val_dataset.create_dataloader(batch_size=batch_size)
test_dataloader = test_dataset.create_dataloader(batch_size=batch_size)
batch_X, batch_y = next(iter(test_dataloader))
print ("Sample batch:\n"
f" X: {list(batch_X.size())}\n"
f" y: {list(batch_y.size())}\n"
"Sample point:\n"
f" X: {batch_X[0]}\n"
f" y: {batch_y[0]}")
# + [markdown] id="Y5FMz_VkzvvG"
# # CNN
# + [markdown] id="I6Cs6GYInnD8"
# ## Inputs
# + [markdown] id="sWf6pSvunpoC"
# We're going to learn about CNNs by applying them on 1D text data. In the dummy example below, our inputs are composed of character tokens that are one-hot encoded. We have a batch of N samples, where each sample has 8 characters and each character is represented by an array of 10 values (`vocab size=10`). This gives our inputs the size `(N, 8, 10)`.
#
# > With PyTorch, when dealing with convolution, our inputs (X) need to have the channels as the second dimension, so our inputs will be `(N, 10, 8)`.
# + id="c4AW9_QGpBqS"
import math
import torch
import torch.nn as nn
import torch.nn.functional as F
# + id="bCQeJEUVnnrR" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416461, "user_tz": 420, "elapsed": 20989, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="4c09110b-fb0e-45cd-ea0d-c93786a17e98"
# Assume all our inputs are padded to have the same # of words
batch_size = 64
max_seq_len = 8 # words per input
vocab_size = 10 # one hot size
x = torch.randn(batch_size, max_seq_len, vocab_size)
print(f"X: {x.shape}")
x = x.transpose(1, 2)
print(f"X: {x.shape}")
# + [markdown] id="UAJMhb7DsFgV"
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/madewithml/main/images/ml-foundations/cnn/inputs.png" width="500">
# </div>
#
# This diagram above is for char-level tokens but extends to any level of tokenization (word-level in our case).
# + [markdown] id="3uLslqxFl_au"
# ## Filters
# + [markdown] id="JhhTVijAl-Yp"
# At the core of CNNs are filters (aka weights, kernels, etc.) which convolve (slide) across our input to extract relevant features. The filters are initialized randomly but learn to pick up meaningful features from the input that aid in optimizing for the objective. The intuition here is that each filter represents a feature and we will use this filter on other inputs to capture the same feature (feature extraction via parameter sharing).
#
# We can see convolution in the diagram below where we simplified the filters and inputs to be 2D for ease of visualization. Also note that the values are 0/1s but in reality they can be any floating point value.
# + [markdown] id="du4gRM5htR9W"
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/madewithml/main/images/ml-foundations/cnn/convolution.gif" width="500">
# </div>
# + [markdown] id="96PfAWzYsOEI"
# Now let's return to our actual inputs `x`, which is of shape (8, 10) [`max_seq_len`, `vocab_size`] and we want to convolve on this input using filters. We will use 50 filters that are of size (1, 3) and has the same depth as the number of channels (`num_channels` = `vocab_size` = `one_hot_size` = 10). This gives our filter a shape of (3, 10, 50) [`kernel_size`, `vocab_size`, `num_filters`]
#
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/madewithml/main/images/ml-foundations/cnn/filters.png" width="500">
# </div>
# + [markdown] id="26nX0cVX-Bwl"
# * **stride**: amount the filters move from one convolution operation to the next.
# * **padding**: values (typically zero) padded to the input, typically to create a volume with whole number dimensions.
# + [markdown] id="q5DH85gdEdR_"
# So far we've used a `stride` of 1 and `VALID` padding (no padding) but let's look at an example with a higher stride and difference between different padding approaches.
#
# Padding types:
# * **VALID**: no padding, the filters only use the "valid" values in the input. If the filter cannot reach all the input values (filters go left to right), the extra values on the right are dropped.
# * **SAME**: adds padding evenly to the right (preferred) and left sides of the input so that all values in the input are processed.
#
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/madewithml/main/images/ml-foundations/cnn/padding.png" width="500">
# </div>
# + [markdown] id="wN7QCnCWuwiG"
# We're going to use the [Conv1d](https://pytorch.org/docs/stable/generated/torch.nn.Conv1d.html#torch.nn.Conv1d) layer to process our inputs.
# + id="0bwK2BE6diB3" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416464, "user_tz": 420, "elapsed": 20967, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="02f074d8-95bb-415d-9e8b-43bce2a84e4a"
# Convolutional filters (VALID padding)
vocab_size = 10 # one hot size
num_filters = 50 # num filters
filter_size = 3 # filters are 3X3
stride = 1
padding = 0 # valid padding (no padding)
conv1 = nn.Conv1d(in_channels=vocab_size, out_channels=num_filters,
kernel_size=filter_size, stride=stride,
padding=padding, padding_mode='zeros')
print("conv: {}".format(conv1.weight.shape))
# + id="HYxn8MejfGi0" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416467, "user_tz": 420, "elapsed": 20947, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="28db5639-ac9c-4882-840f-be851a066f94"
# Forward pass
z = conv1(x)
print (f"z: {z.shape}")
# + [markdown] id="wFD9qBslxs4A"
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/madewithml/main/images/ml-foundations/cnn/conv.png" width="700">
# </div>
# + [markdown] id="tcBbTPW6tZtr"
# When we apply these filter on our inputs, we receive an output of shape (N, 6, 50). We get 50 for the output channel dim because we used 50 filters and 6 for the conv outputs because:
#
# $W_1 = \frac{W_2 - F + 2P}{S} + 1 = \frac{8 - 3 + 2(0)}{1} + 1 = 6$
#
# $H_1 = \frac{H_2 - F + 2P}{S} + 1 = \frac{1 - 1 + 2(0)}{1} + 1 = 1$
#
# $D_2 = D_1 $
#
# where:
# * `W`: width of each input = 8
# * `H`: height of each input = 1
# * `D`: depth (# channels)
# * `F`: filter size = 3
# * `P`: padding = 0
# * `S`: stride = 1
# + [markdown] id="NSE9tWhUKHPb"
# Now we'll add padding so that the convolutional outputs are the same shape as our inputs. The amount of padding for the `SAME` padding can be determined using the same equation. We want out output to have the same width as our input, so we solve for P:
#
# $ \frac{W-F+2P}{S} + 1 = W $
#
# $ P = \frac{S(W-1) - W + F}{2} $
#
# If $P$ is not a whole number, we round up (using `math.ceil`) and place the extra padding on the right side.
# + id="uXB9FwR6EkeA" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416470, "user_tz": 420, "elapsed": 20927, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="f7af9beb-2084-4857-f340-1d1ae8d3139b"
# Convolutional filters (SAME padding)
vocab_size = 10 # one hot size
num_filters = 50 # num filters
filter_size = 3 # filters are 3X3
stride = 1
conv = nn.Conv1d(in_channels=vocab_size, out_channels=num_filters,
kernel_size=filter_size, stride=stride)
print("conv: {}".format(conv.weight.shape))
# + id="HQLr0RG9uIgK" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416472, "user_tz": 420, "elapsed": 20905, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="0574101c-1413-491b-dd80-6994378ab32a"
# `SAME` padding
padding_left = int((conv.stride[0]*(max_seq_len-1) - max_seq_len + filter_size)/2)
padding_right = int(math.ceil((conv.stride[0]*(max_seq_len-1) - max_seq_len + filter_size)/2))
print (f"padding: {(padding_left, padding_right)}")
# + id="KCxWRzxAEmJW" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416475, "user_tz": 420, "elapsed": 20879, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="26fe7b05-2059-4599-98ad-0f666556a1d4"
# Forward pass
z = conv(F.pad(x, (padding_left, padding_right)))
print (f"z: {z.shape}")
# + [markdown] id="FRD72rMHvWHN"
# > We will explore larger dimensional convolution layers in subsequent lessons. For example, [Conv2D](https://pytorch.org/docs/stable/generated/torch.nn.Conv2d.html#torch.nn.Conv2d) is used with 3D inputs (images, char-level text, etc.) and [Conv3D](https://pytorch.org/docs/stable/generated/torch.nn.Conv3d.html#torch.nn.Conv3d) is used for 4D inputs (videos, time-series, etc.).
# + [markdown] id="HpB8HSJNwLp-"
# ## Pooling
# + [markdown] id="WxRnbPy6wLuc"
# The result of convolving filters on an input is a feature map. Due to the nature of convolution and overlaps, our feature map will have lots of redundant information. Pooling is a way to summarize a high-dimensional feature map into a lower dimensional one for simplified downstream computation. The pooling operation can be the max value, average, etc. in a certain receptive field. Below is an example of pooling where the outputs from a conv layer are `4X4` and we're going to apply max pool filters of size `2X2`.
#
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/madewithml/main/images/ml-foundations/cnn/pooling.png" width="500">
# </div>
# + [markdown] id="ok0EBECKc2QU"
# $W_2 = \frac{W_1 - F}{S} + 1 = \frac{4 - 2}{2} + 1 = 2$
#
# $H_2 = \frac{H_1 - F}{S} + 1 = \frac{4 - 2}{2} + 1 = 2$
#
# $ D_2 = D_1 $
#
# where:
# * `W`: width of each input = 4
# * `H`: height of each input = 4
# * `D`: depth (# channels)
# * `F`: filter size = 2
# * `S`: stride = 2
# + [markdown] id="5ijJtky9QHeX"
# In our use case, we want to just take the one max value so we will use the [MaxPool1D](https://pytorch.org/docs/stable/generated/torch.nn.MaxPool1d.html#torch.nn.MaxPool1d) layer, so our max-pool filter size will be max_seq_len.
#
# + id="niptcsv2wUPA" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416477, "user_tz": 420, "elapsed": 20856, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="40ff5268-8688-4536-8c20-9a2355a009fa"
# Max pooling
pool_output = F.max_pool1d(z, z.size(2))
print("Size: {}".format(pool_output.shape))
# + [markdown] id="ccelFfH-s3ZY"
# ## Batch Normalization
# + [markdown] id="9f67F4o1HHQp"
# The last topic we'll cover before constructing our model is [batch normalization](https://arxiv.org/abs/1502.03167). It's an operation that will standardize (mean=0, std=1) the activations from the previous layer. Recall that we used to standardize our inputs in previous notebooks so our model can optimize quickly with larger learning rates. It's the same concept here but we continue to maintain standardized values throughout the forward pass to further aid optimization.
# + id="owtCbYoZs82g" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416479, "user_tz": 420, "elapsed": 20834, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="c2399f3a-a21a-45a8-8ddf-873ed6d6163b"
# Batch normalization
batch_norm = nn.BatchNorm1d(num_features=num_filters)
z = batch_norm(conv(x)) # applied to activations (after conv layer & before pooling)
print (f"z: {z.shape}")
# + id="3ABUSm-vyaTG" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416483, "user_tz": 420, "elapsed": 20812, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="405e600d-e99f-46f1-89ad-cb713332270b"
# Mean and std before batchnorm
print (f"mean: {torch.mean(conv1(x)):.2f}, std: {torch.std(conv(x)):.2f}")
# + id="fNzIKpJUyQqk" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416486, "user_tz": 420, "elapsed": 20789, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="fca1b7e5-a699-4e12-b3c3-3e288bac2afb"
# Mean and std after batchnorm
print (f"mean: {torch.mean(z):.2f}, std: {torch.std(z):.2f}")
# + [markdown] id="Hpo4QXTOtGV6"
# # Modeling
# + [markdown] id="pfhjWZRD94hK"
# ## Model
# + [markdown] id="zVmJGm8m-KIz"
# Let's visualize the model's forward pass.
#
# 1. We'll first tokenize our inputs (`batch_size`, `max_seq_len`).
# 2. Then we'll one-hot encode our tokenized inputs (`batch_size`, `max_seq_len`, `vocab_size`).
# 3. We'll apply convolution via filters (`filter_size`, `vocab_size`, `num_filters`) followed by batch normalization. Our filters act as character level n-gram detectors.
# 4. We'll apply 1D global max pooling which will extract the most relevant information from the feature maps for making the decision.
# 5. We feed the pool outputs to a fully-connected (FC) layer (with dropout).
# 6. We use one more FC layer with softmax to derive class probabilities.
# + [markdown] id="3ilKr0yTgl6o"
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/madewithml/main/images/ml-foundations/cnn/model.png" width="1000">
# </div>
# + id="jKXRELvlwQHe"
NUM_FILTERS = 50
HIDDEN_DIM = 100
DROPOUT_P = 0.1
# + id="UPP5ROd69mXC"
class CNN(nn.Module):
def __init__(self, vocab_size, num_filters, filter_size,
hidden_dim, dropout_p, num_classes):
super(CNN, self).__init__()
# Convolutional filters
self.filter_size = filter_size
self.conv = nn.Conv1d(
in_channels=vocab_size, out_channels=num_filters,
kernel_size=filter_size, stride=1, padding=0, padding_mode='zeros')
self.batch_norm = nn.BatchNorm1d(num_features=num_filters)
# FC layers
self.fc1 = nn.Linear(num_filters, hidden_dim)
self.dropout = nn.Dropout(dropout_p)
self.fc2 = nn.Linear(hidden_dim, num_classes)
def forward(self, inputs, channel_first=False, apply_softmax=False):
# Rearrange input so num_channels is in dim 1 (N, C, L)
x_in, = inputs
if not channel_first:
x_in = x_in.transpose(1, 2)
# Padding for `SAME` padding
max_seq_len = x_in.shape[2]
padding_left = int((self.conv.stride[0]*(max_seq_len-1) - max_seq_len + self.filter_size)/2)
padding_right = int(math.ceil((self.conv.stride[0]*(max_seq_len-1) - max_seq_len + self.filter_size)/2))
# Conv outputs
z = self.conv(F.pad(x_in, (padding_left, padding_right)))
z = F.max_pool1d(z, z.size(2)).squeeze(2)
# FC layer
z = self.fc1(z)
z = self.dropout(z)
y_pred = self.fc2(z)
if apply_softmax:
y_pred = F.softmax(y_pred, dim=1)
return y_pred
# + id="wD4sRUS5_lwq" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608329416494, "user_tz": 420, "elapsed": 20741, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="a22a4bc0-7c3a-4228-b6ba-9e8852a0aa2d"
# Initialize model
model = CNN(vocab_size=VOCAB_SIZE, num_filters=NUM_FILTERS, filter_size=FILTER_SIZE,
hidden_dim=HIDDEN_DIM, dropout_p=DROPOUT_P, num_classes=NUM_CLASSES)
model = model.to(device) # set device
print (model.named_parameters)
# + [markdown] id="s1L7-vxbgCup"
# > We used `SAME` padding (w/ stride=1) which means that the conv outputs will have the same width (`max_seq_len`) as our inputs. The amount of padding differs for each batch based on the `max_seq_len` but you can calculate it by solving for P in the equation below.
#
# $ \frac{W_1 - F + 2P}{S} + 1 = W_2 $
#
# $ \frac{\text{max_seq_len } - \text{ filter_size } + 2P}{\text{stride}} + 1 = \text{max_seq_len} $
#
# $ P = \frac{\text{stride}(\text{max_seq_len}-1) - \text{max_seq_len} + \text{filter_size}}{2} $
#
# If $P$ is not a whole number, we round up (using `math.ceil`) and place the extra padding on the right side.
# + [markdown] id="tzZTZI-y2Tzy"
# ## Training
# + [markdown] id="SYw0JY9k2VPk"
# Let's create the `Trainer` class that we'll use to facilitate training for our experiments. Notice that we're now moving the `train` function inside this class.
# + id="2Cd0KoTq3MhV"
from torch.optim import Adam
# + id="NPUmSaoR3PQc"
LEARNING_RATE = 1e-3
PATIENCE = 5
NUM_EPOCHS = 10
# + id="H3EqfumC2Ud4"
class Trainer(object):
def __init__(self, model, device, loss_fn=None, optimizer=None, scheduler=None):
# Set params
self.model = model
self.device = device
self.loss_fn = loss_fn
self.optimizer = optimizer
self.scheduler = scheduler
def train_step(self, dataloader):
"""Train step."""
# Set model to train mode
self.model.train()
loss = 0.0
# Iterate over train batches
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, targets = batch[:-1], batch[-1]
self.optimizer.zero_grad() # Reset gradients
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, targets) # Define loss
J.backward() # Backward pass
self.optimizer.step() # Update weights
# Cumulative Metrics
loss += (J.detach().item() - loss) / (i + 1)
return loss
def eval_step(self, dataloader):
"""Validation or test step."""
# Set model to eval mode
self.model.eval()
loss = 0.0
y_trues, y_probs = [], []
# Iterate over val batches
with torch.no_grad():
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, y_true = batch[:-1], batch[-1]
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, y_true).item()
# Cumulative Metrics
loss += (J - loss) / (i + 1)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
y_trues.extend(y_true.cpu().numpy())
return loss, np.vstack(y_trues), np.vstack(y_probs)
def predict_step(self, dataloader):
"""Prediction step."""
# Set model to eval mode
self.model.eval()
y_probs = []
# Iterate over val batches
with torch.no_grad():
for i, batch in enumerate(dataloader):
# Forward pass w/ inputs
inputs, targets = batch[:-1], batch[-1]
y_prob = self.model(inputs, apply_softmax=True)
# Store outputs
y_probs.extend(y_prob)
return np.vstack(y_probs)
def train(self, num_epochs, patience, train_dataloader, val_dataloader):
best_val_loss = np.inf
for epoch in range(num_epochs):
# Steps
train_loss = self.train_step(dataloader=train_dataloader)
val_loss, _, _ = self.eval_step(dataloader=val_dataloader)
self.scheduler.step(val_loss)
# Early stopping
if val_loss < best_val_loss:
best_val_loss = val_loss
best_model = self.model
_patience = patience # reset _patience
else:
_patience -= 1
if not _patience: # 0
print("Stopping early!")
break
# Logging
print(
f"Epoch: {epoch+1} | "
f"train_loss: {train_loss:.5f}, "
f"val_loss: {val_loss:.5f}, "
f"lr: {self.optimizer.param_groups[0]['lr']:.2E}, "
f"_patience: {_patience}"
)
return best_model
# + id="SMfTLWuB290z"
# Define Loss
class_weights_tensor = torch.Tensor(list(class_weights.values())).to(device)
loss = nn.CrossEntropyLoss(weight=class_weights_tensor)
# + id="wAjAGYpX3FG5"
# Define optimizer & scheduler
optimizer = Adam(model.parameters(), lr=LEARNING_RATE)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(
optimizer, mode='min', factor=0.1, patience=3)
# + id="GoA55gCt3FJk"
# Trainer module
trainer = Trainer(
model=model, device=device, loss_fn=loss_fn,
optimizer=optimizer, scheduler=scheduler)
# + colab={"base_uri": "https://localhost:8080/"} id="ub6AB1qB3JCm" executionInfo={"status": "ok", "timestamp": 1608329481433, "user_tz": 420, "elapsed": 85574, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="7763483e-d152-4530-ccf3-199b05bb54a0"
# Train
best_model = trainer.train(
NUM_EPOCHS, PATIENCE, train_dataloader, val_dataloader)
# + [markdown] id="fG6ejmWj4DhH"
# ## Evaluation
# + id="8k2WdDeH3S6P"
import json
from pathlib import Path
from sklearn.metrics import precision_recall_fscore_support
# + id="Sne12LQZ4HTx"
def get_performance(y_true, y_pred, classes):
"""Per-class performance metrics."""
# Performance
performance = {"overall": {}, "class": {}}
# Overall performance
metrics = precision_recall_fscore_support(y_true, y_pred, average="weighted")
performance["overall"]["precision"] = metrics[0]
performance["overall"]["recall"] = metrics[1]
performance["overall"]["f1"] = metrics[2]
performance["overall"]["num_samples"] = np.float64(len(y_true))
# Per-class performance
metrics = precision_recall_fscore_support(y_true, y_pred, average=None)
for i in range(len(classes)):
performance["class"][classes[i]] = {
"precision": metrics[0][i],
"recall": metrics[1][i],
"f1": metrics[2][i],
"num_samples": np.float64(metrics[3][i]),
}
return performance
# + id="TXRbjnaz4I1H"
# Get predictions
test_loss, y_true, y_prob = trainer.eval_step(dataloader=test_dataloader)
y_pred = np.argmax(y_prob, axis=1)
# + colab={"base_uri": "https://localhost:8080/"} id="b3fVoGmZ5WRR" executionInfo={"status": "ok", "timestamp": 1608329482415, "user_tz": 420, "elapsed": 86486, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="925933ec-16e1-4dec-914c-4ce5307a1cab"
# Determine performance
performance = get_performance(
y_true=y_test, y_pred=y_pred, classes=label_encoder.classes)
print (json.dumps(performance['overall'], indent=2))
# + id="wIgO8aQd46p1"
# Save artifacts
dir = Path("cnn")
dir.mkdir(parents=True, exist_ok=True)
label_encoder.save(fp=Path(dir, 'label_encoder.json'))
tokenizer.save(fp=Path(dir, 'tokenizer.json'))
torch.save(best_model.state_dict(), Path(dir, 'model.pt'))
with open(Path(dir, 'performance.json'), "w") as fp:
json.dump(performance, indent=2, sort_keys=False, fp=fp)
# + [markdown] id="9SYtJtBB42jA"
# ## Inference
# + id="u0pGgxCN9W9I"
def get_probability_distribution(y_prob, classes):
"""Create a dict of class probabilities from an array."""
results = {}
for i, class_ in enumerate(classes):
results[class_] = np.float64(y_prob[i])
sorted_results = {k: v for k, v in sorted(
results.items(), key=lambda item: item[1], reverse=True)}
return sorted_results
# + colab={"base_uri": "https://localhost:8080/"} id="WciQjoMf4KZL" executionInfo={"status": "ok", "timestamp": 1608329482420, "user_tz": 420, "elapsed": 86434, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="48942e17-41b4-47f9-989e-a100c28c4a8d"
# Load artifacts
device = torch.device("cpu")
label_encoder = LabelEncoder.load(fp=Path(dir, 'label_encoder.json'))
tokenizer = Tokenizer.load(fp=Path(dir, 'tokenizer.json'))
model = CNN(
vocab_size=VOCAB_SIZE, num_filters=NUM_FILTERS, filter_size=FILTER_SIZE,
hidden_dim=HIDDEN_DIM, dropout_p=DROPOUT_P, num_classes=NUM_CLASSES)
model.load_state_dict(torch.load(Path(dir, 'model.pt'), map_location=device))
model.to(device)
# + id="5uioiNWd5akm"
# Initialize trainer
trainer = Trainer(model=model, device=device)
# + colab={"base_uri": "https://localhost:8080/"} id="rKHHNuw65dPx" executionInfo={"status": "ok", "timestamp": 1608329482425, "user_tz": 420, "elapsed": 86395, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="96ea1657-6aab-405f-f085-70d80cf7a92d"
# Dataloader
text = "What a day for the new york stock market to go bust!"
sequences = tokenizer.texts_to_sequences([preprocess(text)])
print (tokenizer.sequences_to_texts(sequences))
X = [to_categorical(seq, num_classes=len(tokenizer)) for seq in sequences]
y_filler = label_encoder.encode([label_encoder.classes[0]]*len(X))
dataset = Dataset(X=X, y=y_filler, max_filter_size=FILTER_SIZE)
dataloader = dataset.create_dataloader(batch_size=batch_size)
# + colab={"base_uri": "https://localhost:8080/"} id="QcZnAxrm5dSY" executionInfo={"status": "ok", "timestamp": 1608329482426, "user_tz": 420, "elapsed": 86367, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="0be1317f-1065-4340-d8cf-f020075bd7a9"
# Inference
y_prob = trainer.predict_step(dataloader)
y_pred = np.argmax(y_prob, axis=1)
label_encoder.decode(y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="WhjciE436mYQ" executionInfo={"status": "ok", "timestamp": 1608329482427, "user_tz": 420, "elapsed": 86340, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="04a9bc66-9692-4fac-ecc0-991a84686c55"
# Class distributions
prob_dist = get_probability_distribution(y_prob=y_prob[0], classes=label_encoder.classes)
print (json.dumps(prob_dist, indent=2))
# + [markdown] id="1TsxDKyRArrQ"
# # Interpretability
# + [markdown] id="wqa6OxyeAtkj"
# We went through all the trouble of padding our inputs before convolution to result is outputs of the same shape as our inputs so we can try to get some interpretability. Since every token is mapped to a convolutional output on whcih we apply max pooling, we can see which token's output was most influential towards the prediction. We first need to get the conv outputs from our model:
# + id="T-TywFzL54LS"
import collections
import seaborn as sns
# + id="-I6gSfZ9BSOD"
class InterpretableCNN(nn.Module):
def __init__(self, vocab_size, num_filters, filter_size,
hidden_dim, dropout_p, num_classes):
super(InterpretableCNN, self).__init__()
# Convolutional filters
self.filter_size = filter_size
self.conv = nn.Conv1d(
in_channels=vocab_size, out_channels=num_filters,
kernel_size=filter_size, stride=1, padding=0, padding_mode='zeros')
self.batch_norm = nn.BatchNorm1d(num_features=num_filters)
# FC layers
self.fc1 = nn.Linear(num_filters, hidden_dim)
self.dropout = nn.Dropout(dropout_p)
self.fc2 = nn.Linear(hidden_dim, num_classes)
def forward(self, inputs, channel_first=False, apply_softmax=False):
# Rearrange input so num_channels is in dim 1 (N, C, L)
x_in, = inputs
if not channel_first:
x_in = x_in.transpose(1, 2)
# Padding for `SAME` padding
max_seq_len = x_in.shape[2]
padding_left = int((self.conv.stride[0]*(max_seq_len-1) - max_seq_len + self.filter_size)/2)
padding_right = int(math.ceil((self.conv.stride[0]*(max_seq_len-1) - max_seq_len + self.filter_size)/2))
# Conv outputs
z = self.conv(F.pad(x_in, (padding_left, padding_right)))
return z
# + id="kdgTQI3qBwVe"
# Initialize
interpretable_model = InterpretableCNN(
vocab_size=len(tokenizer), num_filters=NUM_FILTERS, filter_size=FILTER_SIZE,
hidden_dim=HIDDEN_DIM, dropout_p=DROPOUT_P, num_classes=NUM_CLASSES)
# + colab={"base_uri": "https://localhost:8080/"} id="iF8ckOT8BSQ5" executionInfo={"status": "ok", "timestamp": 1608329482431, "user_tz": 420, "elapsed": 86287, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="cab4e23e-4634-4081-acc3-15cb3a9402ea"
# Load weights (same architecture)
interpretable_model.load_state_dict(torch.load(Path(dir, 'model.pt'), map_location=device))
interpretable_model.to(device)
# + id="kFDPbdHLCJpa"
# Initialize trainer
interpretable_trainer = Trainer(model=interpretable_model, device=device)
# + colab={"base_uri": "https://localhost:8080/"} id="defuFVDYBSTn" executionInfo={"status": "ok", "timestamp": 1608329513133, "user_tz": 420, "elapsed": 609, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="03d0a31e-d853-42ff-bd7b-fe42282f7d12"
# Get conv outputs
conv_outputs = interpretable_trainer.predict_step(dataloader)
print (conv_outputs.shape) # (num_filters, max_seq_len)
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="S8BeyVdk6Mzd" executionInfo={"status": "ok", "timestamp": 1608329546685, "user_tz": 420, "elapsed": 685, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} outputId="cca94fd3-d355-4ac1-fdc6-0a573bbf90ec"
# Visualize a bi-gram filter's outputs
tokens = tokenizer.sequences_to_texts(sequences)[0].split(' ')
sns.heatmap(conv_outputs, xticklabels=tokens)
# + [markdown] id="NWakIG-C6Ybh"
# The filters have high values for the words `stock` and `market` which influenced the `Business` category classification.
# + [markdown] id="3UsWNPkaHGUZ"
# > This is a crude technique (maxpool doesn't strictly behave this way on a batch) loosely based off of more elaborate [interpretability](https://arxiv.org/abs/1312.6034) methods.
| notebooks/11_Convolutional_Neural_Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Machine Translation with Seq2Seq and Transformers
# In this exercise you will implement a [Sequence to Sequence(Seq2Seq)](https://arxiv.org/abs/1703.03906) and a [Transformer](https://arxiv.org/pdf/1706.03762.pdf) model and use them to perform machine translation.
#
# **A quick note: if you receive the following TypeError "super(type, obj): obj must be an instance or subtype of type", try re-importing that part or restarting your kernel and re-running all cells.** Once you have finished making changes to`the model constuctor, you can avoid this issue by commenting out all of the model instantiations after the first (e.g. lines starting with "model = TransformerTranslator(*args, **kwargs)").
# +
import numpy as np
import csv
import torch
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# -
# # ** 1: Introduction**
#
# ## Multi30K: Multilingual English-German Image Descriptions
#
# [Multi30K](https://github.com/multi30k/dataset) is a dataset for machine translation tasks. It is a multilingual corpus containing English sentences and their German translation. In total it contains 31014 sentences(29000 for training, 1014 for validation, and 1000 for testing).
# As one example:
#
# En: `Two young, White males are outside near many bushes.`
#
# De: `Zwei junge weiße Männer sind im Freien in der Nähe vieler Büsche.`
#
# You can read more info about the dataset [here](https://arxiv.org/abs/1605.00459). The following parts of this assignment will be based on this dataset.
#
# ## TorchText: A PyTorch Toolkit for Text Dataset and NLP Tasks
# [TorchText](https://github.com/pytorch/text) is a PyTorch package that consists of data processing utilities and popular datasets for natural language. The key idea of TorchText is that datasets can be organized in *Field*, *TranslationDataset*, and *BucketIterator* classes. They serve to help with data splitting and loading, token encoding, sequence padding, etc. You don't need to know about how TorchText works in detail, but you might want to know about why those classes are needed and what operations are necessary for machine translation. This knowledge can be migrated to all sequential data modeling. In the following parts, we will provide you with some code to help you understand.
#
# You can refer to torchtext's documentation(v0.9.0) [here](https://pytorch.org/text/).
#
# ## Spacy
# Spacy is package designed for tokenization in many languages. Tokenization is a process of splitting raw text data into lists of tokens that can be further processed. Since TorchText only provides tokenizer for English, we will be using Spacy for our assignment.
#
#
# **Notice: For the following assignment, we strongly recommend you to work in a virtual python environment. We recommend Anaconda, a powerful environment control tool. You can download it [here](https://www.anaconda.com/products/individual)**.
# ## ** 1.1: Prerequisites**
# Before you start this assignment, you need to have all required packages installed either on the terminal you are using, or in the virtual environment. Please make sure you have the following package installed:
#
# `PyTorch, TorchText, Spacy, Tqdm, Numpy`
#
# You can first check using either `pip freeze` in terminal or `conda list` in conda environment. Then run the following code block to make sure they can be imported.
# + tags=[]
# Just run this block. Please do not modify the following code.
import math
import time
# Pytorch package
import torch
import torch.nn as nn
import torch.optim as optim
# Torchtest package
from torchtext.datasets import Multi30k
from torchtext.data import Field, BucketIterator
# Tqdm progress bar
from tqdm import tqdm_notebook, tqdm
# Code provide to you for training and evaluation
from utils import train, evaluate, set_seed_nb, unit_test_values
# -
# Once you properly import the above packages, you can proceed to download Spacy English and German tokenizers by running the following command in your **terminal**. They will take some time.
#
# `python -m spacy download en`
#
# `python -m spacy download de`
# Now lets check your GPU availability and load some sanity checkers. By default you should be using your gpu for this assignment if you have one available.
# + tags=[]
# Check device availability
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print("You are using device: %s" % device)
# -
# load checkers
d1 = torch.load('./data/d1.pt')
d2 = torch.load('./data/d2.pt')
d3 = torch.load('./data/d3.pt')
d4 = torch.load('./data/d4.pt')
# ## **1.2: Preprocess Data**
# With TorchText and Spacy tokenizers ready, you can now prepare the data using *TorchText* objects. Just run the following code blocks. Read the comment and try to understand what they are for.
# + tags=[]
# You don't need to modify any code in this block
# Define the maximum length of the sentence. Shorter sentences will be padded to that length and longer sentences will be croped. Given that the average length of the sentence in the corpus is around 13, we can set it to 20
MAX_LEN = 20
# Define the source and target language
SRC = Field(tokenize = "spacy",
tokenizer_language="de",
init_token = '<sos>',
eos_token = '<eos>',
fix_length = MAX_LEN,
lower = True)
TRG = Field(tokenize = "spacy",
tokenizer_language="en",
init_token = '<sos>',
eos_token = '<eos>',
fix_length = MAX_LEN,
lower = True)
# Download and split the data. It should take some time
train_data, valid_data, test_data = Multi30k.splits(exts = ('.de', '.en'),
fields = (SRC, TRG))
# +
# Define Batchsize
BATCH_SIZE = 128
# Build the vocabulary associated with each language
SRC.build_vocab(train_data, min_freq = 2)
TRG.build_vocab(train_data, min_freq = 2)
# Get the padding index to be ignored later in loss calculation
PAD_IDX = TRG.vocab.stoi['<pad>']
# Get data-loaders using BucketIterator
train_loader, valid_loader, test_loader = BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE, device = device)
# Get the input and the output sizes for model
input_size = len(SRC.vocab)
output_size = len(TRG.vocab)
# -
# # **2: Implement Vanilla RNN and LSTM**
# In this section, you will need to implement a Vanilla RNN and an LSTM unit using PyTorch Linear layers and nn.Parameter. This is designed to help you to understand how they work behind the scene. The code you will be working with is in *LSTM.py* and *RNN.py* under *naive* folder. Please refer to instructions among this notebook and those files.
#
# ## **2.1: Implement an RNN Unit**
# In this section you will be using PyTorch Linear layers and activations to implement a vanilla RNN unit. You are only implementing an RNN cell unit over one time step! The test case reflects this by having only one sequence. Please refer to the following structure and complete the code in RNN.py:
# <center><img src="imgs/RNN.png" width="300" align="center"/></center>
# Run the following block to check your implementation
# + tags=[]
from models.naive.RNN import VanillaRNN
set_seed_nb()
x1,x2 = (1,4), (-1,2)
h1,h2 = (-1,2,0,4), (0,1,3,-1)
batch = 4
x = torch.FloatTensor(np.linspace(x1,x2,batch))
h = torch.FloatTensor(np.linspace(h1,h2,batch))
rnn = VanillaRNN(x.shape[-1], h.shape[-1], 3)
out, hidden = rnn.forward(x,h)
expected_out, expected_hidden = unit_test_values('rnn')
if out is not None:
print('Close to out: ', expected_out.allclose(out, atol=1e-4))
print('Close to hidden: ', expected_hidden.allclose(hidden, atol=1e-4))
print('hidden: ',hidden)
print('expected_hidden: ', expected_hidden)
print('out: ', out)
print('expected_out', expected_out)
else:
print("NOT IMPLEMENTED")
# -
# ## **2.2: Implement an LSTM Unit**
# In this section you will be using PyTorch nn.Parameter and activations to implement an LSTM unit. You can simply translate the following equations using nn.Parameter and PyTorch activation functions to build an LSTM from scratch:
# \begin{array}{ll} \\
# i_t = \sigma(W_{ii} x_t + b_{ii} + W_{hi} h_{t-1} + b_{hi}) \\
# f_t = \sigma(W_{if} x_t + b_{if} + W_{hf} h_{t-1} + b_{hf}) \\
# g_t = \tanh(W_{ig} x_t + b_{ig} + W_{hg} h_{t-1} + b_{hg}) \\
# o_t = \sigma(W_{io} x_t + b_{io} + W_{ho} h_{t-1} + b_{ho}) \\
# c_t = f_t \odot c_{t-1} + i_t \odot g_t \\
# h_t = o_t \odot \tanh(c_t) \\
# \end{array}
#
# Here's a great visualization of the above equation from [Colah's blog](https://colah.github.io/posts/2015-08-Understanding-LSTMs/) to help you understand LSTM unit. You can also read more about it from that blog.
# <center><img src="imgs/lstm.png" width="400" height="300" align="center"/></center>
#
# If you want to see nn.Parameter in example, check out this [tutorial](https://pytorch.org/tutorials/beginner/nn_tutorial.html) from PyTorch. Run the following block to check your implementation.
#
# Note that in this case we are implementing a full loop with LSTM, iterating over each time step. The test cases reflect this as there are multiple sequences.
# + tags=[]
from models.naive.LSTM import LSTM
set_seed_nb()
x1,x2 = np.mgrid[-1:3:3j, -1:4:2j]
h1,h2 = np.mgrid[-2:2:3j, 1:3:4j]
batch = 4
x = torch.FloatTensor(np.linspace(x1,x2,batch))
h = torch.FloatTensor(np.linspace(h1,h2,batch))
lstm = LSTM(x.shape[-1], h.shape[-1])
h_t, c_t = lstm.forward(x)
expected_ht, expected_ct = unit_test_values('lstm')
if h_t is not None:
print('Close to h_t: ', expected_ht.allclose(h_t, atol=1e-4))
print('Close to c_t; ', expected_ct.allclose(c_t, atol=1e-4))
else:
print("NOT IMPLEMENTED")
# -
# # **3: Train a Seq2Seq Model**
# In this section, you will be working on implementing a simple Seq2Seq model. You will first implement an Encoder and a Decoder, and then join them together with a Seq2Seq architecture. You will need to complete the code in *Decoder.py*, *Encoder.py*, and *Seq2Seq.py* under *seq2seq* folder. Please refer to the instructions in those files.
# ## **3.1: Implement the Encoder**
# In this section you will be implementing an RNN/LSTM based encoder to model English texts. Please refer to the instructions in *seq2seq/Encoder.py*. Run the following block to check your implementation.
# +
from models.seq2seq.Encoder import Encoder
set_seed_nb()
i, n, h = 10, 4, 2
encoder = Encoder(i, n, h, h)
x_array = np.random.rand(5,1) * 10
x = torch.LongTensor(x_array)
out, hidden = encoder.forward(x)
expected_out, expected_hidden = unit_test_values('encoder')
print('Close to out: ', expected_out.allclose(out, atol=1e-4))
print('Close to hidden: ', expected_hidden.allclose(hidden, atol=1e-4))
# -
# ## **3.2: Implement the Decoder**
# In this section you will be implementing an RNN/LSTM based decoder to model German texts. Please refer to the instructions in *seq2seq/Decoder.py*. Run the following block to check your implementation.
# +
from models.seq2seq.Decoder import Decoder
set_seed_nb()
i, n, h = 10, 2, 2
decoder = Decoder(h, n, n, i)
x_array = np.random.rand(5, 1) * 10
x = torch.LongTensor(x_array)
_, enc_hidden = unit_test_values('encoder')
out, hidden = decoder.forward(x,enc_hidden)
expected_out, expected_hidden = unit_test_values('decoder')
print('Close to out: ', expected_out.allclose(out, atol=1e-4))
print('Close to hidden: ', expected_hidden.allclose(hidden, atol=1e-4))
# -
# ## **3.3: Implement the Seq2Seq**
# In this section you will be implementing the Seq2Seq model that utilizes the Encoder and Decoder you implemented. Please refer to the instructions in *seq2seq/Seq2Seq.py*. Run the following block to check your implementation.
# +
from models.seq2seq.Seq2Seq import Seq2Seq
set_seed_nb()
embedding_size = 32
hidden_size = 32
input_size = 8
output_size = 8
batch, seq = 1, 2
encoder = Encoder(input_size, embedding_size, hidden_size, hidden_size)
decoder = Decoder(embedding_size, hidden_size, hidden_size, output_size)
seq2seq = Seq2Seq(encoder, decoder, 'cpu')
x_array = np.random.rand(batch, seq) * 10
x = torch.LongTensor(x_array)
out = seq2seq.forward(x)
expected_out = unit_test_values('seq2seq')
print('Close to out: ', expected_out.allclose(out, atol=1e-4))
# -
# ## **3.4: Train your Seq2Seq model**
# Now it's time to combine what we have and train a Seq2Seq translator. We provided you with some training code and you can simply run them to see how your translator works. If you implemented everything correctly, you should see some meaningful translation in the output. You can modify the hyperparameters to improve the results. You can also tune the BATCH_SIZE in section 1.2.
# +
# Hyperparameters. You are welcome to modify these
encoder_emb_size = 32
encoder_hidden_size = 64
encoder_dropout = 0.2
decoder_emb_size = 32
decoder_hidden_size = 64
decoder_dropout = 0.2
learning_rate = 1e-3
model_type = "RNN"
EPOCHS = 10
#input size and output size
input_size = len(SRC.vocab)
output_size = len(TRG.vocab)
# +
# Declare models, optimizer, and loss function
encoder = Encoder(input_size, encoder_emb_size, encoder_hidden_size, decoder_hidden_size, dropout = encoder_dropout, model_type = model_type)
decoder = Decoder(decoder_emb_size, encoder_hidden_size, encoder_hidden_size, output_size, dropout = decoder_dropout, model_type = model_type)
seq2seq_model = Seq2Seq(encoder, decoder, device)
optimizer = optim.Adam(seq2seq_model.parameters(), lr = learning_rate)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(optimizer)
criterion = nn.CrossEntropyLoss(ignore_index=PAD_IDX)
# -
for epoch_idx in range(EPOCHS):
print("-----------------------------------")
print("Epoch %d" % (epoch_idx+1))
print("-----------------------------------")
train_loss, avg_train_loss = train(seq2seq_model, train_loader, optimizer, criterion)
scheduler.step(train_loss)
val_loss, avg_val_loss = evaluate(seq2seq_model, valid_loader, criterion)
avg_train_loss = avg_train_loss.item()
avg_val_loss = avg_val_loss.item()
print("Training Loss: %.4f. Validation Loss: %.4f. " % (avg_train_loss, avg_val_loss))
print("Training Perplexity: %.4f. Validation Perplexity: %.4f. " % (np.exp(avg_train_loss), np.exp(avg_val_loss)))
# # **4: Train a Transformer**
#
# We will be implementing a one-layer Transformer **encoder** which, similar to an RNN, can encode a sequence of inputs and produce a final output of possibility of tokens in target language. This is the architecture:
#
# <center><img src="imgs/encoder.png" align="center"/></center>
#
#
# You can refer to the [original paper](https://arxiv.org/pdf/1706.03762.pdf) for more details.
#
# ## The Corpus of Linguistic Acceptability (CoLA)
#
# The Corpus of Linguistic Acceptability ([CoLA](https://nyu-mll.github.io/CoLA/)) in its full form consists of 10657 sentences from 23 linguistics publications, expertly annotated for acceptability (grammaticality) by their original authors. Native English speakers consistently report a sharp contrast in acceptability between pairs of sentences.
# Some examples include:
#
# `What did Betsy paint a picture of?` (Correct)
#
# `What was a picture of painted by Betsy?` (Incorrect)
#
# You can read more info about the dataset [here](https://arxiv.org/pdf/1805.12471.pdf). This is a binary classification task (predict 1 for correct grammar and 0 otherwise).
#
# We will be using this dataset as a sanity checker for the forward pass of the Transformer architecture discussed in class. The general intuitive notion is that we will _encode_ the sequence of tokens in the sentence, and then predict a binary output based on the final state that is the output of the model.
# ## Load the preprocessed data
#
# We've appended a "CLS" token to the beginning of each sequence, which can be used to make predictions. The benefit of appending this token to the beginning of the sequence (rather than the end) is that we can extract it quite easily (we don't need to remove paddings and figure out the length of each individual sequence in the batch). We'll come back to this.
#
# We've additionally already constructed a vocabulary and converted all of the strings of tokens into integers which can be used for vocabulary lookup for you. Feel free to explore the data here.
# + tags=[]
train_inxs = np.load('./data/train_inxs.npy')
val_inxs = np.load('./data/val_inxs.npy')
train_labels = np.load('./data/train_labels.npy')
val_labels = np.load('./data/val_labels.npy')
# load dictionary
word_to_ix = {}
with open("./data/word_to_ix.csv", "r") as f:
reader = csv.reader(f)
for line in reader:
word_to_ix[line[0]] = line[1]
print("Vocabulary Size:", len(word_to_ix))
print(train_inxs.shape) # 7000 training instances, of (maximum/padded) length 43 words.
print(val_inxs.shape) # 1551 validation instances, of (maximum/padded) length 43 words.
print(train_labels.shape)
print(val_labels.shape)
d1 = torch.load('./data/d1.pt')
d2 = torch.load('./data/d2.pt')
d3 = torch.load('./data/d3.pt')
d4 = torch.load('./data/d4.pt')
# -
# Instead of using numpy for this model, we will be using Pytorch to implement the forward pass. You will not need to implement the backward pass for the various layers in this assigment.
#
# The file `models/Transformer.py` contains the model class and methods for each layer. This is where you will write your implementations.
# ## **4.1: Embeddings**
#
# We will format our input embeddings similarly to how they are constructed in [BERT (source of figure)](https://arxiv.org/pdf/1810.04805.pdf). Recall from lecture that unlike a RNN, a Transformer does not include any positional information about the order in which the words in the sentence occur. Because of this, we need to append a positional encoding token at each position. (We will ignore the segment embeddings and [SEP] token here, since we are only encoding one sentence at a time). We have already appended the [CLS] token for you in the previous step.
# <center><img src="imgs/embedding.png" align="center"/></center>
#
# Your first task is to implement the embedding lookup, including the addition of positional encodings. Open the file `Transformer.py` and complete all code parts for `Deliverable 1`.
# +
from models.Transformer import TransformerTranslator
inputs = train_inxs[0:2]
inputs = torch.LongTensor(inputs)
model = TransformerTranslator(input_size=len(word_to_ix), output_size=2, device=device, hidden_dim=128, num_heads=2, dim_feedforward=2048, dim_k=96, dim_v=96, dim_q=96, max_length=train_inxs.shape[1])
embeds = model.embed(inputs)
try:
print("Difference:", torch.sum(torch.pairwise_distance(embeds, d1)).item()) # should be very small (<0.01)
except:
print("NOT IMPLEMENTED")
# -
# ## **4.2: Multi-head Self-Attention**
#
# Attention can be computed in matrix-form using the following formula:
# <center><img src="imgs/attn.png" align="center"/></center>
#
# We want to have multiple self-attention operations, computed in parallel. Each of these is called a *head*. We concatenate the heads and multiply them with the matrix `attention_head_projection` to produce the output of this layer.
#
# After every multi-head self-attention and feedforward layer, there is a residual connection + layer normalization. Make sure to implement this, using the following formula:
# <center><img src="imgs/layer_norm.png" align="center"/></center>
#
#
# Open the file `models/transformer.py` and implement the `multihead_attention` function.
# We have already initialized all of the layers you will need in the constructor.
# +
hidden_states = model.multi_head_attention(embeds)
try:
print("Difference:", torch.sum(torch.pairwise_distance(hidden_states, d2)).item()) # should be very small (<0.01)
except:
print("NOT IMPLEMENTED")
# -
# ## **4.3: Element-Wise Feed-forward Layer**
#
# Open the file `models/transformer.py` and complete code for `Deliverable 3`: the element-wise feed-forward layer consisting of two linear transformers with a ReLU layer in between.
#
# <center><img src="imgs/ffn.png" align="center"/></center>
#
# +
outputs = model.feedforward_layer(hidden_states)
try:
print("Difference:", torch.sum(torch.pairwise_distance(outputs, d3)).item()) # should be very small (<0.01)
except:
print("NOT IMPLEMENTED")
# -
# ## **4.4: Final Layer**
#
# Open the file `models/transformer.py` and complete code for `Deliverable 4`, to produce probability scores for all tokens in target language.
# +
scores = model.final_layer(outputs)
try:
print("Difference:", torch.sum(torch.pairwise_distance(scores, d4)).item()) # should be very small (<3e-5)
except:
print("NOT IMPLEMENTED")
# -
# ## **4.5: Putting it all together**
#
# Open the file `models/transformer.py` and complete the method `forward`, by putting together all of the methods you have developed in the right order to perform a full forward pass.
#
# +
inputs = train_inxs[0:2]
inputs = torch.LongTensor(inputs)
outputs = model.forward(inputs)
try:
print("Difference:", torch.sum(torch.pairwise_distance(outputs, scores)).item()) # should be very small (<3e-5)
except:
print("NOT IMPLEMENTED")
# -
# Great! We've just implemented a Transformer forward pass for translation. One of the big perks of using PyTorch is that with a simple training loop, we can rely on automatic differentation ([autograd](https://pytorch.org/tutorials/beginner/blitz/autograd_tutorial.html)) to do the work of the backward pass for us. This is not required for this assignment, but you can explore this on your own.
# ## **4.6: Train the Transformer**
# Now you can start training the Transformer translator. We provided you with some training code and you can simply run them to see how your translator works. If you implemented everything correctly, you should see some meaningful translation in the output. Compare the results from the Seq2Seq model, which one is better? You can modify the hyperparameters to improve the results. You can also tune the BATCH_SIZE in section 1.2.
# +
# Hyperparameters
learning_rate = 1e-3
EPOCHS = 10
# Model
trans_model = TransformerTranslator(input_size, output_size, device, max_length = MAX_LEN).to(device)
# optimizer = optim.Adam(model.parameters(), lr = learning_rate)
optimizer = torch.optim.Adam(trans_model.parameters(), lr=learning_rate)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(optimizer)
criterion = nn.CrossEntropyLoss(ignore_index=PAD_IDX)
# -
for epoch_idx in range(EPOCHS):
print("-----------------------------------")
print("Epoch %d" % (epoch_idx+1))
print("-----------------------------------")
train_loss, avg_train_loss = train(trans_model, train_loader, optimizer, criterion)
scheduler.step(train_loss)
val_loss, avg_val_loss = evaluate(trans_model, valid_loader, criterion)
avg_train_loss = avg_train_loss.item()
avg_val_loss = avg_val_loss.item()
print("Training Loss: %.4f. Validation Loss: %.4f. " % (avg_train_loss, avg_val_loss))
print("Training Perplexity: %.4f. Validation Perplexity: %.4f. " % (np.exp(avg_train_loss), np.exp(avg_val_loss)))
# **Translations**
#
# Run the code below to see some of your translations. Modify to your liking.
def translate(model, dataloader):
model.eval()
with torch.no_grad():
# Get the progress bar
progress_bar = tqdm(dataloader, ascii = True)
for batch_idx, data in enumerate(progress_bar):
source = data.src.transpose(1,0)
target = data.trg.transpose(1,0)
translation = model(source)
return target, translation
# Select Transformer or Seq2Seq model
model = trans_model
# model = seq2seq_model
#Set model equal to trans_model or seq2seq_model
target, translation = translate(model, valid_loader)
raw = np.array([list(map(lambda x: TRG.vocab.itos[x], target[i])) for i in range(target.shape[0])])
raw
token_trans = np.argmax(translation.cpu().numpy(), axis = 2)
translated = np.array([list(map(lambda x: TRG.vocab.itos[x], token_trans[i])) for i in range(token_trans.shape[0])])
translated
| assignment4/Machine_Translation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# nbdev: hide this cell and export functions to `model_analysis` by default
# hide
# default_exp model_fitting
# -
# # Likelihood-Based Model Fitting
# We'll demo a way to fit models to datasets with item repetitions based on the likelihood of each recall event given the model and a specified parameter configuration. The big difference from more traditional datasets is the variable item presentation orders across trials. This requires extra model simulation, since study phases can't be simulated just once. To ensure code is performant, we use a few tricks to keep simulations to a minimum.
# +
# export
import numpy as np
from numba import njit
from repfr.models import DefaultCMR
@njit(fastmath=True, nogil=True)
def cmr_rep_likelihood(
trials, presentations, list_types, list_length, encoding_drift_rate, start_drift_rate,
recall_drift_rate, shared_support, item_support, learning_rate,
primacy_scale, primacy_decay, stop_probability_scale,
stop_probability_growth, choice_sensitivity):
"""
Generalized cost function for fitting the InstanceCMR model optimized
using the numba library.
Output scales inversely with the likelihood that the model and specified
parameters would generate the specified trials. For model fitting, is
usually wrapped in another function that fixes and frees parameters for
optimization.
**Arguments**:
- data_to_fit: typed list of int64-arrays where rows identify a unique
trial of responses and columns corresponds to a unique recall index.
- A configuration for each parameter of `InstanceCMR` as delineated in
`Formal Specification`.
**Returns** the negative sum of log-likelihoods across specified trials
conditional on the specified parameters and the mechanisms of InstanceCMR.
"""
likelihood = np.ones((len(trials), list_length))
# we can use the same model for list types 1 and 2
stable_models = [DefaultCMR(
list_length, list_length, encoding_drift_rate, start_drift_rate,
recall_drift_rate, shared_support, item_support, learning_rate,
primacy_scale, primacy_decay, stop_probability_scale,
stop_probability_growth, choice_sensitivity),
DefaultCMR(
int(list_length/2), list_length, encoding_drift_rate, start_drift_rate,
recall_drift_rate, shared_support, item_support, learning_rate,
primacy_scale, primacy_decay, stop_probability_scale,
stop_probability_growth, choice_sensitivity)]
stable_models[0].experience(np.eye(list_length, list_length))
stable_models[1].experience(np.eye(int(list_length/2), int(list_length/2))[np.repeat(np.arange(int(list_length/2)), 2)])
for trial_index in range(len(trials)):
item_count = np.max(presentations[trial_index])+1
if list_types[trial_index] > 2:
model = DefaultCMR(
item_count, list_length, encoding_drift_rate, start_drift_rate,
recall_drift_rate, shared_support, item_support, learning_rate,
primacy_scale, primacy_decay, stop_probability_scale,
stop_probability_growth, choice_sensitivity)
model.experience(np.eye(item_count, item_count)[presentations[trial_index]])
else:
model = stable_models[list_types[trial_index]-1]
trial = trials[trial_index]
model.force_recall()
for recall_index in range(len(trial) + 1):
# identify index of item recalled; if zero then recall is over
if recall_index == len(trial) and len(trial) < item_count:
recall = 0
elif trial[recall_index] == 0:
recall = 0
else:
recall = presentations[trial_index][trial[recall_index]-1] + 1
# store probability of and simulate recalling item with this index
likelihood[trial_index, recall_index] = \
model.outcome_probabilities(model.context)[recall]
if recall == 0:
break
model.force_recall(recall)
# reset model to its pre-retrieval (but post-encoding) state
model.force_recall(0)
return -np.sum(np.log(likelihood))
# -
# For model fitting, we have to prepare a wrapper function that calls `cmr_rep_likelihood` using static parameters for some values:
# +
# export
def cmr_rep_objective_function(data_to_fit, presentations, list_types, list_length, fixed_parameters, free_parameters):
"""
Generates and returns an objective function for input to support search
through parameter space for ICMR model fit using an optimization function.
Arguments:
- fixed_parameters: dictionary mapping parameter names to values they'll
be fixed to during search, overloaded by free_parameters if overlap
- free_parameters: list of strings naming parameters for fit during search
- data_to_fit: array where rows identify a unique trial of responses and
columns corresponds to a unique recall index
Returns a function that accepts a vector x specifying arbitrary values for
free parameters and returns evaluation of icmr_likelihood using the model
class, all parameters, and provided data.
"""
return lambda x: cmr_rep_likelihood(data_to_fit, presentations, list_types, list_length, **{**fixed_parameters, **{
free_parameters[i]:x[i] for i in range(len(x))}})
# -
# ## Single-Subject Demo
# +
from repfr.datasets import prepare_repdata
trials, events, list_length, presentations, list_types, rep_data, subjects = prepare_repdata(
'data/repFR.mat')
events.head()
# +
import numpy as np
lb = np.finfo(float).eps
hand_fit_parameters = {
'encoding_drift_rate': .8,
'start_drift_rate': .7,
'recall_drift_rate': .8,
'shared_support': 0.01,
'item_support': 1.0,
'learning_rate': .3,
'primacy_scale': 1,
'primacy_decay': 1,
'stop_probability_scale': 0.01,
'stop_probability_growth': 0.3,
'choice_sensitivity': 2
}
cmr_rep_likelihood(trials[:48], presentations[:48], list_types[:48], list_length, **hand_fit_parameters)
# -
# %%timeit
cmr_rep_likelihood(trials[:48], presentations[:48], list_types[:48], list_length, **hand_fit_parameters)
# Now we perform the single-subject fitting...
# +
from scipy.optimize import differential_evolution
import numpy as np
free_parameters = [
'encoding_drift_rate',
'start_drift_rate',
'recall_drift_rate',
'shared_support',
'item_support',
'learning_rate',
'primacy_scale',
'primacy_decay',
'stop_probability_scale',
'stop_probability_growth',
'choice_sensitivity']
lb = np.finfo(float).eps
ub = 1-np.finfo(float).eps
bounds = [
(lb, ub),
(lb, ub),
(lb, ub),
(lb, ub),
(lb, ub),
(lb, ub),
(lb, 100),
(lb, 100),
(lb, ub),
(lb, 10),
(lb, 10)
]
# cost function to be minimized
# ours scales inversely with the probability that the data could have been
# generated using the specified parameters and our model
cost_function = cmr_rep_objective_function(
trials[:48], presentations[:48], list_types[:48], list_length, {}, free_parameters)
result = differential_evolution(cost_function, bounds, disp=True)
print(result)
# -
# ```
# fun: 1867.69002758535
# jac: array([-0.67825567, -0.0390628 , 1.13855094, -0.69376256, -0.76609013,
# 0.09645191, 0.21655069, -0.12521468, 2.94776328, -0.05979928,
# -0.26766429])
# message: 'Optimization terminated successfully.'
# nfev: 16944
# nit: 75
# success: True
# x: array([8.23734393e-01, 6.87377340e-01, 9.63817756e-01, 7.92149143e-02,
# 2.95856638e-01, 2.50030299e-01, 5.94400358e+00, 1.61904883e-01,
# 1.54430590e-03, 2.56455105e-01, 1.61047189e+00])
# ```
# +
from repfr.model_analysis import sim_recall_probability_by_lag
stored_result = [8.23734393e-01, 6.87377340e-01, 9.63817756e-01, 7.92149143e-02,
2.95856638e-01, 2.50030299e-01, 5.94400358e+00, 1.61904883e-01,
1.54430590e-03, 2.56455105e-01, 1.61047189e+00]
parameters = {
'presentation_count': list_length,
}
cmr_result = sim_recall_probability_by_lag(
DefaultCMR, {**parameters, **{free_parameters[i]:stored_result[i] for i in range(len(stored_result))}},
presentations[list_types==4], experiment_count=100)
# +
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_theme(style="white")
tips = sns.load_dataset("tips")
ax = sns.barplot(x=['N/A', '0', '1-2', '3-5', '6-8'],
y=cmr_result[-1])
plt.xlabel('Number of Intervening Items Between Repetitions')
plt.ylabel('Recall Probability')
plt.title('CMR Simulations After Fitting Across Conditions')
plt.show()
| 05_Fitting_By_Likelihood.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: book_env
# language: python
# name: book_env
# ---
#
import pandas as pd
import numpy as np
data = pd.read_csv('parsed.csv')
data.columns
data.title.head()
data.isin(['Ring of Fire']).any()
x = data['title'].str.extract(r'([^,]*)$')
y = data['title'].str.extract(r'( - .*)$')
data['place'].sort_values().unique()
# 1. Find the 95th percentile of earthquake magnitude in Japan using the
# magType of 'mb'.
data.loc[data.magType == 'mb'].mag.quantile(0.95)
# 2. Find the percentage of earthquakes in Indonesia that were coupled with
# tsunamis.
indonesia = data.loc[(data.place == 'Indonesia') & (data.type == 'earthquake'), ['type', 'tsunami', 'place']]
indonesia_tsunami = indonesia.loc[indonesia.tsunami != 0]
(indonesia_tsunami.tsunami.count()/indonesia.tsunami.count())*100
# 3. Get summary statistics for earthquakes in Nevada.
data.loc[(data.place.str.contains(r'NV|Nevada', na=False) & (data.type == 'earthquake'))].describe()
# 4. Add a column to the dataframe indicating whether or not the earthquake
# happened in a country or US state that is on the Ring of Fire. Use:
# Bolivia, Chile, Ecuador, Peru, Costa Rica, Guatemala, Mexico (be careful not to
# select New Mexico), Japan, Philippines, Indonesia, New Zealand,
# Antarctica (look for Antarctic), Canada, Fiji, Alaska, Washington,
# California, Russia, Taiwan, Tonga, and Kermadec Islands.
#adding Ring of Fire location boolean mask
data['rof_loc'] = data['place'].isin(
['Bolivia','Chile','Ecuador','Peru','Costa Rica','Guatemala','Mexico',
'Japan','Philippines','Indonesia','New Zealand','Antarctic','Canada',
'Fiji', 'Alaska','Washington','California','Russia','Taiwan','Tonga','Kermadec Islands'])
data['rof_earthquake'] = (data['type'] == 'earthquake') & (data['rof_loc'] == 1)
data[['type','title','place','rof_loc','rof_earthquake']].describe()
# 5. Calculate the number of earthquakes in the Ring of Fire locations and the
# number outside them.
#number of earthquakes in the Ring of Fire locations
data.loc[(data['rof_loc'] == 1) & (data['type']== 'earthquake'),'type'].count()
#number of earthquakes outside Ring of Fire locations
data.loc[(data['rof_loc'] == 0) & (data['type']== 'earthquake'),'type'].count()
# 6. Find the tsunami count along the Ring of Fire.
data.loc[(data['rof_loc'] == 1) & (data['tsunami']== 1), 'tsunami'].count()
| ch_02/data/exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ML Pipeline Preparation
# Follow the instructions below to help you create your ML pipeline.
# ### 1. Import libraries and load data from database.
# - Import Python libraries
# - Load dataset from database with [`read_sql_table`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_sql_table.html)
# - Define feature and target variables X and Y
# +
# import libraries
from sqlalchemy import create_engine
import numpy as np
import pandas as pd
from sklearn.metrics import confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.multioutput import MultiOutputClassifier
from sklearn.metrics import classification_report,confusion_matrix, precision_score,\
recall_score,accuracy_score, f1_score, make_scorer
from sklearn.base import BaseEstimator, TransformerMixin
import nltk
from nltk import word_tokenize
import pickle
# -
def load_data():
# load data from database
#engine = create_engine('sqlite:///DisasterResponse_new.db')
#df = pd.read_sql("SELECT * FROM DisasterResponse_new", engine)
df = pd.read_csv("DisasterResponse_new.csv")
X = df.message
y = df.loc[:,"related":"direct_report"]
category_names=y.columns
return X, y,category_names
# Use the first five messages as a sample to take a look at the data
X,y,category_names=load_data()
print(X[:5])
y.head(5)
# ### 2. Normalize text data
# A sequence of functions used to clean up HTML markups, expand contractions, stem and lemmatize, remove special characters, get rid of stop words, and remove accents from characters, etc. is defined in the notebook called Text_Normalization_Function. Run the notebook and the functions will be available in this notebook
# %run ./Text_Normalization_Function.ipynb
# The normalize_corpus can be used as a customized preprocessor in CountVectorizer.\
# **preprocessor** should be a callable, default=None. Override the preprocessing (strip_accents and lowercase) stage while preserving the tokenizing and n-grams generation steps. It should return a text **(not a series or list)**.
#
# However, if a function is used to normalize the corpus before feeding to CountVectorizer, the function should return a series or list.
# Use the first five messages as a sample to take a look at result after CountVectorizer
bow_vectorizer = CountVectorizer(preprocessor=normalize_corpus)
NORM_corpus_train_bow = bow_vectorizer.fit_transform(X[:5])
NORM_corpus_train_bow_table= pd.DataFrame(data = NORM_corpus_train_bow.todense(),
columns = bow_vectorizer.get_feature_names())
NORM_corpus_train_bow_table.head()
# ### 3. Add other features besides the TF-IDF
# Other characteristics of the text, such as length, may also affect the results. I defined a function to count the number of tokens contained in the text
class Text_Length_Extractor(BaseEstimator, TransformerMixin):
def get_length(self, text):
length=len(word_tokenize(text))
return length
def __init__(self):
pass
def fit(self, X, y=None):
return self
def transform(self, X):
X_length = pd.Series(X).apply(self.get_length)
# In order to use FeatureUnion to combine the Text_Length_Extractor with the text_pipeline,
# We must convert X_length into a dataframe. Otherwise, ValueError: blocks[0,:] has incompatible row dimensions.
return pd.DataFrame(X_length)
# ### 4. Build a machine learning pipeline
# This machine pipeline should take in the `message` column as input and output classification results on the other 36 categories in the dataset. You may find the [MultiOutputClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.multioutput.MultiOutputClassifier.html) helpful for predicting multiple target variables.
pipeline = Pipeline([
('features', FeatureUnion([
('text_pipeline',Pipeline([
('vect', CountVectorizer(preprocessor=normalize_corpus)),
('tfidf', TfidfTransformer())
])),
('text_length',Text_Length_Extractor())
])),
('clf', MultiOutputClassifier(estimator=RandomForestClassifier(random_state=42)))
])
# ### 5. Train pipeline
# - Split data into train and test sets
# - Train pipeline
X_train, X_test, y_train, y_test = train_test_split(X, y)
pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_test)
# ### 6. Test your model
# Report the f1 score, precision and recall for each output category of the dataset.
# The y_pred is a numpy array with a shape of (6554, 36), so we have to access it by referring to its index number.
print(y_pred)
print(y_pred[:,0])
# The y_test is a pd dataframe, if we want to access it by referring to its column number, we can use df.iloc, integer-location based indexing
y_test.head()
# Parameter average: required for multiclass/multilabel targets.
#
# Binary:Only report results for the class specified by pos_label (default is 1).
#
# Macro average (averaging the unweighted mean per label), weighted average (averaging the support-weighted mean per label).
#
# Take F1 score as an example:
#
# Macro F1 calculates the F1 separated by class but not using weights for the aggregation: F1class1+F1class2+⋅⋅⋅+F1classN, which resuls in a bigger penalisation when the model does not perform well with the minority classes(when there is imbalance)
#
# Weighted F1 score calculates the F1 score for each class independently but when it adds them together uses a weight that depends on the number of true labels of each class: F1class1∗W1+F1class2∗W2+⋅⋅⋅+F1classN∗WN.Therefore favouring the majority class
print(classification_report(y_test.iloc[:,0], y_pred[:,0]))
# In this project, I use the default average parameter, binary. The recall and precision for some small categories such as offer and child alone are almost zero. The classifier classified almost everything as 0 due to an imbalance in the training data
#
# Unlike the common problem with only one column of y, this project has 36 columns of y. In order to evaluate the prediction of each column, I use for loop
metrics_list_all=[]
for col in range(y_test.shape[1]):
accuracy = accuracy_score(y_test.iloc[:,col], y_pred[:,col])
precision=precision_score(y_test.iloc[:,col], y_pred[:,col])
recall = recall_score(y_test.iloc[:,col], y_pred[:,col])
f_1 = f1_score(y_test.iloc[:,col], y_pred[:,col])
metrics_list=[accuracy,precision,recall,f_1]
metrics_list_all.append(metrics_list)
metrics_df=pd.DataFrame(metrics_list_all,index=category_names,columns=["Accuracy","Precision","Recall","F_1"])
print(metrics_df)
# If I calculate the accuracy score directly, it will give back very weird result
accuracy_score(y_test.values, y_pred),pipeline.score(X_test,y_test)
# However, if use reshape to flatten the data from having 36 different columns to 1 column (appending data of each column one after the other), the result will be the same as using for loop to calculate the accuracy score of each column and then calculate the average
#
# numpy.reshape(a, newshape, order='C') gives a new shape to an array without changing its data.
accuracy_score(y_test.values.reshape(-1,1), y_pred.reshape(-1,1))
print(("The average accuracy score among all categories is {:.4f},\nthe average precision score score among all categories is {:.4f},\nthe average recall score among all categories is {:.4f},\nthe average F 1 score among all categories is {:.4f}").format(metrics_df.mean()["Accuracy"],metrics_df.mean()["Precision"],metrics_df.mean()["Recall"],metrics_df.mean()["F_1"]))
# ### 7. Improve your model
# Use grid search to find better parameters.
# Define a score used in scoring parameter
def avg_accuracy(y_test, y_pred):
"""
This is the score_func used in make_scorer, which would be used in in GridSearchCV
"""
avg_accuracy=accuracy_score(y_test.values.reshape(-1,1), y_pred.reshape(-1,1))
return avg_accuracy
avg_accuracy_cv = make_scorer(avg_accuracy)
# Take a look at what parameters are available to be tuned
list(pipeline.get_params())
# +
parameters = parameters = {
#'features__text_pipeline__vect__ngram_range': ((1, 1), (1, 2)),
'clf__estimator__max_depth': [15, 30],
'clf__estimator__n_estimators': [100, 250]}
cv = GridSearchCV(
pipeline,
param_grid=parameters,
cv=3,
scoring=avg_accuracy_cv,
verbose=3)
cv.fit(X_train, y_train)
# -
# ### 8. Test your model
# Show the accuracy, precision, and recall of the tuned model.
#
# Since this project focuses on code quality, process, and pipelines, there is no minimum performance metric needed to pass. However, make sure to fine tune your models for accuracy, precision and recall to make your project stand out - especially for your portfolio!
def evaluate_model(model, X_test, y_test,category_names):
"""
The evaluate_model function will return the accuracy, precision, and recall, and f1 scores for each output category of the dataset.
INPUTS:
model- a trained model for evaluation
X_test - a panda data frame or Numpy array, contains the untouched values of features.
y_pred - a Numpy array, contains predicted category values of the messages.
OUTPUT:
metrics_df, a panda dataframe that contains accuracy, precision, and recall, and f1 scores for each output category of the dataset.
"""
y_pred=model.predict(X_test)
metrics_list_all=[]
for col in range(y_test.shape[1]):
accuracy = accuracy_score(y_test.iloc[:,col], y_pred[:,col])
precision=precision_score(y_test.iloc[:,col], y_pred[:,col])
recall = recall_score(y_test.iloc[:,col], y_pred[:,col])
f_1 = f1_score(y_test.iloc[:,col], y_pred[:,col])
metrics_list=[accuracy,precision,recall,f_1]
metrics_list_all.append(metrics_list)
metrics_df=pd.DataFrame(metrics_list_all,index=category_names,columns=["Accuracy","Precision","Recall","F_1"])
print(metrics_df)
print("----------------------------------------------------------------------")
print(("The average accuracy score among all categories is {:.4f},\nthe average precision score score among all categories is {:.4f},\nthe average recall score among all categories is {:.4f},\nthe average F 1 score among all categories is {:.4f}").format(metrics_df.mean()["Accuracy"],metrics_df.mean()["Precision"],metrics_df.mean()["Recall"],metrics_df.mean()["F_1"]))
return None
# Get the best model and store it as best_randomforest
best_randomforest=cv.best_estimator_
evaluate_model(best_randomforest, X_test, y_test,category_names)
# ### 9. Export your model as a pickle file
# **Pickle** is the standard way of serializing objects in Python.You can use the pickle operation to serialize your machine learning algorithms and save the serialized format to a file.
#
# Later you can load this file to deserialize your model and use it to make new predictions.
filename = 'best_randomforest.pkl'
pickle.dump(pipeline, open(filename, 'wb'))
# ### 10. Use this notebook to complete `train.py`
# Use the template file attached in the Resources folder to write a script that runs the steps above to create a database and export a model based on a new dataset specified by the user.
| ML Pipeline Preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# *<NAME>*
# last modified: 03/31/2014
# <hr>
# I am really looking forward to your comments and suggestions to improve and extend this tutorial! Just send me a quick note
# via Twitter: [@rasbt](https://twitter.com/rasbt)
# or Email: [<EMAIL>](mailto:<EMAIL>)
# <hr>
# ### Problem Category
# - Statistical Pattern Recognition
# - Supervised Learning
# - Parametric Learning
# - Bayes Decision Theory
# - Univariate data
# - 2-class problem
# - equal variances
# - equal priors
# - Gaussian model (2 parameters)
# - No Risk function
# <hr>
# <p><a name="sections"></a>
# <br></p>
#
# # Sections
#
#
# <p>• <a href="#given">Given information</a><br>
# • <a href="#deriving_db">Deriving the decision boundary</a><br>
# • <a href="#plotting_db">Plotting the class conditional densities, posterior probabilities, and decision boundary</a><br>
# • <a href="#classify_rand">Classifying some random example data</a><br>
# • <a href="#emp_err">Calculating the empirical error rate</a><br>
#
#
#
#
#
#
#
#
# <hr>
# <p><a name="given"></a>
# <br></p>
#
# ## Given information:
#
# [<a href="#sections">back to top</a>] <br>
#
#
# ####model: continuous univariate normal (Gaussian) model for the class-conditional densities
#
#
# $ p(x | \omega_j) \sim N(\mu|\sigma^2) $
#
# $ p(x | \omega_j) \sim \frac{1}{\sqrt{2\pi\sigma^2}} \exp{ \bigg[-\frac{1}{2}\bigg( \frac{x-\mu}{\sigma}\bigg)^2 \bigg] } $
#
#
# ####Prior probabilities:
#
# $ P(\omega_1) = P(\omega_1) = 0.5 $
#
# #### Variances of the sample distributions
#
# $ \sigma_1^2 = \sigma_2^2 = 1 $
#
# #### Means of the sample distributions
#
# $ \mu_1 = 4, \quad \mu_2 = 10 $
#
#
# <br>
# <p><a name="deriving_db"></a>
# <br></p>
#
# ## Deriving the decision boundary
# [<a href="#sections">back to top</a>] <br>
# ### Bayes' Rule:
#
#
# $ P(\omega_j|x) = \frac{p(x|\omega_j) * P(\omega_j)}{p(x)} $
#
# ###Bayes' Decision Rule:
#
# Decide $ \omega_1 $ if $ P(\omega_1|x) > P(\omega_2|x) $ else decide $ \omega_2 $.
# <br>
#
#
# \begin{equation}
# \Rightarrow \frac{p(x|\omega_1) * P(\omega_1)}{p(x)} > \frac{p(x|\omega_2) * P(\omega_2)}{p(x)}
# \end{equation}
#
# We can drop $ p(x) $ since it is just a scale factor.
#
#
# $ \Rightarrow P(x|\omega_1) * P(\omega_1) > p(x|\omega_2) * P(\omega_2) $
#
# $ \Rightarrow \frac{p(x|\omega_1)}{p(x|\omega_2)} > \frac{P(\omega_2)}{P(\omega_1)} $
#
# $ \Rightarrow \frac{p(x|\omega_1)}{p(x|\omega_2)} > \frac{0.5}{0.5} $
#
# $ \Rightarrow \frac{p(x|\omega_1)}{p(x|\omega_2)} > 1 $
#
# $ \Rightarrow \frac{1}{\sqrt{2\pi\sigma_1^2}} \exp{ \bigg[-\frac{1}{2}\bigg( \frac{x-\mu_1}{\sigma_1}\bigg)^2 \bigg] } > \frac{1}{\sqrt{2\pi\sigma_2^2}} \exp{ \bigg[-\frac{1}{2}\bigg( \frac{x-\mu_2}{\sigma_2}\bigg)^2 \bigg] } $
#
#
# Since we have equal variances, we can drop the first term completely.
#
#
#
#
# $ Rightarrow \exp{ \bigg[-\frac{1}{2}\bigg( \frac{x-\mu_1}{\sigma_1}\bigg)^2 \bigg] } > \exp{ \bigg[-\frac{1}{2}\bigg( \frac{x-\mu_2}{\sigma_2}\bigg)^2 \bigg] } \quad\quad \bigg| \;ln, \quad \mu_1 = 4, \quad \mu_2 = 10, \quad \sigma=1 $
#
# $ \Rightarrow -\frac{1}{2} (x-4)^2 > -\frac{1}{2} (x-10)^2 \quad \bigg| \; \times(-2) $
#
# $ \Rightarrow (x-4)^2 < (x-10)^2 $
#
# $ \Rightarrow x^2 - 8x + 16 < x^2 - 20x + 100 $
#
# $ \Rightarrow 12x < 84 $
#
# $ \Rightarrow x < 7 $
#
# <p><a name="plotting_db"></a>
# <br></p>
#
# ## Plotting the class conditional densities, posterior probabilities, and decision boundary
#
# [<a href="#sections">back to top</a>] <br>
# +
# %pylab inline
import numpy as np
from matplotlib import pyplot as plt
def pdf(x, mu, sigma):
"""
Calculates the normal distribution's probability density
function (PDF).
"""
term1 = 1.0 / ( math.sqrt(2*np.pi) * sigma )
term2 = np.exp( -0.5 * ( (x-mu)/sigma )**2 )
return term1 * term2
# generating some sample data
x = np.arange(0, 100, 0.05)
# probability density functions
pdf1 = pdf(x, mu=4, sigma=1)
pdf2 = pdf(x, mu=10, sigma=1)
# Class conditional densities (likelihoods)
plt.plot(x, pdf1)
plt.plot(x, pdf2)
plt.title('Class conditional densities (likelihoods)')
plt.ylabel('p(x)')
plt.xlabel('random variable x')
plt.legend(['p(x|w_1) ~ N(4,1)', 'p(x|w_2) ~ N(10,1)'], loc='upper right')
plt.ylim([0,0.5])
plt.xlim([0,20])
plt.show()
# +
def posterior(likelihood, prior):
"""
Calculates the posterior probability (after Bayes Rule) without
the scale factor p(x) (=evidence).
"""
return likelihood * prior
# probability density functions
posterior1 = posterior(pdf(x, mu=4, sigma=1), 0.5)
posterior2 = posterior(pdf(x, mu=10, sigma=1), 0.5)
# Class conditional densities (likelihoods)
plt.plot(x, posterior1)
plt.plot(x, posterior2)
plt.title('Posterior Probabilities w. Decision Boundary')
plt.ylabel('P(w)')
plt.xlabel('random variable x')
plt.legend(['P(w_1|x)', 'p(w_2|X)'], loc='upper right')
plt.ylim([0,0.5])
plt.xlim([0,20])
plt.axvline(7, color='r', alpha=0.8, linestyle=':', linewidth=2)
plt.annotate('R1', xy=(4, 0.3), xytext=(4, 0.3))
plt.annotate('R2', xy=(10, 0.3), xytext=(10, 0.3))
plt.show()
# -
# <p><a name="classify_rand"></a>
# <br></p>
#
# ## Classifying some random example data
#
# [<a href="#sections">back to top</a>] <br>
# #### Note on generating univariate random data from a Normal Distribution
#
# We can generate random samples drawn from a Normal distribution via the `np.random.randn()` function. Its default is a standard Normal distribution with $ \mu = 0 $ and $ \sigma^2 = 1 $. In order to draw random data from $ N(\mu, \sigma^2) $, we use
# `sigma * np.random.randn(...) + mu`
# +
# Parameters
mu_1 = 4
mu_2 = 10
sigma_1_sqr = 1
sigma_2_sqr = 1
# Generating 10 random samples drawn from a Normal Distribution for class 1 & 2
x1_samples = sigma_1_sqr**0.5 * np.random.randn(10) + mu_1
x2_samples = sigma_1_sqr**0.5 * np.random.randn(10) + mu_2
y = [0 for i in range(10)]
# Plotting sample data with a decision boundary
plt.scatter(x1_samples, y, marker='o', color='green', s=40, alpha=0.5)
plt.scatter(x2_samples, y, marker='^', color='blue', s=40, alpha=0.5)
plt.title('Classifying random example data from 2 classes')
plt.ylabel('P(x)')
plt.xlabel('random variable x')
plt.legend(['w_1', 'w_2'], loc='upper right')
plt.ylim([-0.1,0.1])
plt.xlim([0,20])
plt.axvline(7, color='r', alpha=0.8, linestyle=':', linewidth=2)
plt.annotate('R1', xy=(4, 0.05), xytext=(4, 0.05))
plt.annotate('R2', xy=(10, 0.05), xytext=(10, 0.05))
plt.show()
# -
# <p><a name="emp_err"></a>
# <br></p>
#
# ## Calculating the empirical error rate
#
# [<a href="#sections">back to top</a>] <br>
# +
w1_as_w2, w2_as_w1 = 0, 0
for x1,x2 in zip(x1_samples, x2_samples):
if x1 >= 7:
w1_as_w2 += 1
if x2 < 7:
w2_as_w1 += 1
emp_err = (w1_as_w2 + w2_as_w1) / float(len(x1_samples) + len(x2_samples))
print('Empirical Error: {}%'.format(emp_err * 100))
# -
test complete; Gopal
| tests/others/1_stat_superv_parametric.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import random
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import tensorflow_datasets as tfds
# +
# https://keras.io/examples/vision/reptile/
# +
learning_rate = 0.003
meta_step_size = 0.25
inner_batch_size = 25
eval_batch_size = 25
meta_iters = 2000
eval_iters = 5
inner_iters = 4
eval_interval = 1
train_shots = 20
shots = 5
classes = 5
# +
class Dataset:
# This class will facilitate the creation of a few-shot dataset
# from the Omniglot dataset that can be sampled from quickly while also
# allowing to create new labels at the same time.
def __init__(self, dataset_dir, split_type):
# Download the tfrecord files containing the omniglot data and convert to a
# dataset.
## split = "train" if training else "test"
## ds = tfds.load("omniglot", split=split, as_supervised=True, shuffle_files=False)
# IMPORTING CUSTOM DATASET: https://www.tensorflow.org/datasets/api_docs/python/tfds/folder_dataset/ImageFolder
split = split_type if split_type in ["train", "test", "val"] else None
builder = tfds.ImageFolder(dataset_dir)
ds = builder.as_dataset(split=split_type, as_supervised=True, shuffle_files=False)
# Iterate over the dataset to get each individual image and its class,
# and put that data into a dictionary.
self.data = {}
def extraction(image, label):
# This function will shrink the Omniglot images to the desired size,
# scale pixel values and convert the RGB image to grayscale
image = tf.image.convert_image_dtype(image, tf.float32)
image = tf.image.rgb_to_grayscale(image)
image = tf.image.resize(image, [28, 28])
return image, label
for image, label in ds.map(extraction):
image = image.numpy()
label = str(label.numpy())
if label not in self.data:
self.data[label] = []
self.data[label].append(image)
self.labels = list(self.data.keys())
def get_mini_dataset(
self, batch_size, repetitions, shots, num_classes, split=False
):
temp_labels = np.zeros(shape=(num_classes * shots))
temp_images = np.zeros(shape=(num_classes * shots, 28, 28, 1))
if split:
test_labels = np.zeros(shape=(num_classes))
test_images = np.zeros(shape=(num_classes, 28, 28, 1))
# Get a random subset of labels from the entire label set.
label_subset = random.choices(self.labels, k=num_classes)
for class_idx, class_obj in enumerate(label_subset):
# Use enumerated index value as a temporary label for mini-batch in
# few shot learning.
temp_labels[class_idx * shots : (class_idx + 1) * shots] = class_idx
# If creating a split dataset for testing, select an extra sample from each
# label to create the test dataset.
if split:
test_labels[class_idx] = class_idx
images_to_split = random.choices(
self.data[label_subset[class_idx]], k=shots + 1
)
test_images[class_idx] = images_to_split[-1]
temp_images[
class_idx * shots : (class_idx + 1) * shots
] = images_to_split[:-1]
else:
# For each index in the randomly selected label_subset, sample the
# necessary number of images.
temp_images[
class_idx * shots : (class_idx + 1) * shots
] = random.choices(self.data[label_subset[class_idx]], k=shots)
dataset = tf.data.Dataset.from_tensor_slices(
(temp_images.astype(np.float32), temp_labels.astype(np.int32))
)
dataset = dataset.shuffle(100).batch(batch_size).repeat(repetitions)
if split:
return dataset, test_images, test_labels
return dataset
import urllib3
urllib3.disable_warnings() # Disable SSL warnings that may happen during download.
path_to_dataset = "../dataset/AWEDataset/awe-train-test/"
train_dataset = Dataset(path_to_dataset, split_type="train")
test_dataset = Dataset(path_to_dataset, split_type="val")
# -
# +
_, axarr = plt.subplots(nrows=5, ncols=5, figsize=(20, 20))
sample_keys = list(train_dataset.data.keys())
for a in range(5):
for b in range(5):
temp_image = train_dataset.data[sample_keys[a]][b]
temp_image = np.stack((temp_image[:, :, 0],) * 3, axis=2)
temp_image *= 255
temp_image = np.clip(temp_image, 0, 255).astype("uint8")
if b == 2:
axarr[a, b].set_title("Class : " + sample_keys[a])
axarr[a, b].imshow(temp_image, cmap="gray")
axarr[a, b].xaxis.set_visible(False)
axarr[a, b].yaxis.set_visible(False)
plt.show()
# -
# +
def conv_bn(x):
x = layers.Conv2D(filters=64, kernel_size=3, strides=2, padding="same")(x)
x = layers.BatchNormalization()(x)
return layers.ReLU()(x)
inputs = layers.Input(shape=(28, 28, 1))
x = conv_bn(inputs)
x = conv_bn(x)
x = conv_bn(x)
x = conv_bn(x)
x = layers.Flatten()(x)
outputs = layers.Dense(classes, activation="softmax")(x)
model = keras.Model(inputs=inputs, outputs=outputs)
model.compile()
optimizer = keras.optimizers.SGD(learning_rate=learning_rate)
# -
training = []
testing = []
for meta_iter in range(meta_iters):
frac_done = meta_iter / meta_iters
cur_meta_step_size = (1 - frac_done) * meta_step_size
# Temporarily save the weights from the model.
old_vars = model.get_weights()
# Get a sample from the full dataset.
mini_dataset = train_dataset.get_mini_dataset(
inner_batch_size, inner_iters, train_shots, classes
)
for images, labels in mini_dataset:
with tf.GradientTape() as tape:
preds = model(images)
loss = keras.losses.sparse_categorical_crossentropy(labels, preds)
grads = tape.gradient(loss, model.trainable_weights)
optimizer.apply_gradients(zip(grads, model.trainable_weights))
new_vars = model.get_weights()
# Perform SGD for the meta step.
for var in range(len(new_vars)):
new_vars[var] = old_vars[var] + (
(new_vars[var] - old_vars[var]) * cur_meta_step_size
)
# After the meta-learning step, reload the newly-trained weights into the model.
model.set_weights(new_vars)
# Evaluation loop
if meta_iter % eval_interval == 0:
accuracies = []
for dataset in (train_dataset, test_dataset):
# Sample a mini dataset from the full dataset.
train_set, test_images, test_labels = dataset.get_mini_dataset(
eval_batch_size, eval_iters, shots, classes, split=True
)
old_vars = model.get_weights()
# Train on the samples and get the resulting accuracies.
for images, labels in train_set:
with tf.GradientTape() as tape:
preds = model(images)
loss = keras.losses.sparse_categorical_crossentropy(labels, preds)
grads = tape.gradient(loss, model.trainable_weights)
optimizer.apply_gradients(zip(grads, model.trainable_weights))
test_preds = model.predict(test_images)
test_preds = tf.argmax(test_preds).numpy()
num_correct = (test_preds == test_labels).sum()
# Reset the weights after getting the evaluation accuracies.
model.set_weights(old_vars)
accuracies.append(num_correct / classes)
training.append(accuracies[0])
testing.append(accuracies[1])
if meta_iter % 100 == 0:
print(
"batch %d: train=%f test=%f" % (meta_iter, accuracies[0], accuracies[1])
)
# +
# First, some preprocessing to smooth the training and testing arrays for display.
window_length = 100
train_s = np.r_[
training[window_length - 1 : 0 : -1], training, training[-1:-window_length:-1]
]
test_s = np.r_[
testing[window_length - 1 : 0 : -1], testing, testing[-1:-window_length:-1]
]
w = np.hamming(window_length)
train_y = np.convolve(w / w.sum(), train_s, mode="valid")
test_y = np.convolve(w / w.sum(), test_s, mode="valid")
# Display the training accuracies.
x = np.arange(0, len(test_y), 1)
plt.plot(x, test_y, x, train_y)
plt.legend(["test", "train"])
plt.grid()
train_set, test_images, test_labels = dataset.get_mini_dataset(
eval_batch_size, eval_iters, shots, classes, split=True
)
for images, labels in train_set:
with tf.GradientTape() as tape:
preds = model(images)
loss = keras.losses.sparse_categorical_crossentropy(labels, preds)
grads = tape.gradient(loss, model.trainable_weights)
optimizer.apply_gradients(zip(grads, model.trainable_weights))
test_preds = model.predict(test_images)
test_preds = tf.argmax(test_preds).numpy()
_, axarr = plt.subplots(nrows=1, ncols=5, figsize=(20, 20))
sample_keys = list(train_dataset.data.keys())
for i, ax in zip(range(5), axarr):
temp_image = np.stack((test_images[i, :, :, 0],) * 3, axis=2)
temp_image *= 255
temp_image = np.clip(temp_image, 0, 255).astype("uint8")
ax.set_title(
"Label : {}, Prediction : {}".format(int(test_labels[i]), test_preds[i])
)
ax.imshow(temp_image, cmap="gray")
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
plt.show()
# -
| code/notebooks/Keras-OneShot-5-ear-recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/iamgeekyaseem/Learning_Tree/blob/main/T_Learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="GGucOtxhiDVF"
# Ob represent the binary and rest 11001 represent number 25 in binary
# + id="aYEEqSIlUa_6" colab={"base_uri": "https://localhost:8080/"} outputId="19f76f70-7838-4fc5-f8b5-e744ed43ee3b"
x=bin(25)
print(x[3:])
# + [markdown] id="Vw7wVA5riV_a"
# import maths function
# + id="XKbXMcOAiCdV"
#import math
#x = math.sqrt(25)
#print(math.floor(2.9)) == 2
#print(math.ceil(2.9)) == 3
#m=math.pow(2,3) #gives float number
#print(int(m))
#-----------------------------------------------
#import math as m #"as" function defiunes the math module as m for short form use
##that is also called alies(not sure about the spelling lol)
#x=m.sqrt(25)
#or
#m=math.sqrt(25)
#print(x,m)
#------------------------------------------------
#importing only specific mathematical functions for ex: sqrt and power
#from math import sqrt
#math.sqrt(25)
#math.pow(2,2)
help('math')
# + id="08fBAKIOB2DC"
import sys
x=int(sys.argv[1]) #HERE THE FIRST INPUT WILL BE INDEXED AS ZERO
y=int(sys.argv[2])
z=x**y
print(z) #i yhink it will work in command prompt only https://www.youtube.com/watch?v=4OX49nLNPEE&list=PLsyeobzWxl7poL9JTVyndKe62ieoN-MZ3&index=22
# + [markdown] id="IcD_M2RnKMPt"
# 1.print numbers from 1 to 100 except number divisible by 3 and 5
# 2. print hashtag squre pattern
# + id="soTTw71IDC3o"
#1-----------------------------------------------
#for i in range(1,100):
# if i%3==0 or i%5==0:
# continue
#else:
# print(i)
# + id="QjOMhQiMN1tA"
#perfect sqr num in 1 to 100
for i in range(1,101):
z=i*i
if z>=100:
break
print(z)
# + id="boZuDRJUE8hK"
#fibonacchi sequence 0 1 1 2 3 5 8 13 21...
num=int(input("Numbers of terms that you want to print: "))
first=0
second=1
print("The fibonacchi sequence is 0 1 ",end="")
for i in range(2,num):
third=first+second
first=second
second=third
print(third, end=" ")
# + id="XQ-qt9EK6EA2" colab={"base_uri": "https://localhost:8080/", "height": 135} outputId="80c3a578-06f3-45b2-86cd-fbd23f1e5960"
for i in range(5):
for a in range(4-i):
print(a+1+i,end='')
print(
a,b="ABCD","PQR"
for i in range(4):
print(a[:i+1]+b[i:])
# + id="jsQdt2Dy-uME"
#sort an array in ascenting order
from array import * #'*' means all the functions of array
#help('array')
nums = array('i', [2,1,3,6,2,9,5,0])
#nums.reverse()
cums=nums.tolist() #convert array to list
cums.sort() #sorts the list
print("Ascending order")
for i in cums:
print(i,end=" ")
print()
#for descending order----------------------------
cums.reverse()
print("Descending order")
for i in cums:
print(i, end=" ")
#for i in nums: #or.... for i in range(len(nums)):
# + id="1EYMq_lnUtp4"
#Taking input from user in an array
#first we need to specify the length of the array
from array import *
n= int(input("Length of array: "))
arr=array('i',[])
for i in range(n):
arr.append(int(input("Enter the next number: ")))
print(arr)
#now we search for the index number of one of the input number
#manual method
val=int(input("The number you want to know the index of: "))
k=0
for e in arr:
if e==val:
print(k)
break
k+=1
#by array function
print(arr.index(val))
# + id="7iK0qM_E_VDK"
#prime number or not check
num=int(input("Enter number: "))
for i in range(2,num//2):
if num%i==0:
print("Not Prime")
break
else:
print("Prime")
# + id="13lLlPOQPMWp"
#factorial of a number eg: 8=1.2.4
num=int(input("Enter number: "))
print("Factorials are: ")
for i in range(1,num//2+1):
if num%i==0:
print(i,end=" ")
# + id="S6eYaLPokRMe"
#prime number b/w 1 to 100
for num in range(2, 101):
for i in range(2, num):
if (num % i) == 0:
break
else:
print(num,end=" ")
# + id="G2qRNKrMH3Vx"
#print the triangle pattern like arrow
for i in range(1,6):
for j in range(1,i+1):
print("*", end=" ")
print()
for i in range(5,1,-1):
for j in range(1,i):
print("*", end=" ")
print()
# + [markdown] id="ShCypYfnKRQA"
# Print the value of 1/1 + 2^2/2 + 3^3/3 + 4^4/4 +......
# + id="f54uSQrzKLi5"
#assignmemt 01
n=int(input("Enter some num: "))
sum,x=0,1
while x<=n:
sum=sum+x**x
x=x+1
print(sum)
# + [markdown] id="5TrBKiIaK4Tc"
# Print sum of 1-x+x^2-x^3+x^4-x^5....
# + id="4qgrZocRKnY-"
#assignmemt 02
#value of m(here m=2) in series
m=int(input("Enter number: "))
n=int(input("Upto what nth term you wanna print: "))
sum,x=1,1
while x<=n:
sum=sum+(-m)**x
x=x+1
print(sum)
# + id="oJMEIF28N-IT"
#reversing digit of array
from array import *
#n= int(input("Length of array: "))
arr=array('i',[1,2,3,4,5,6])
#for i in range(n):
#arr.append(int(input("Enter the next number: ")))
for i in arr:
m=(arr[::-i])
break
print(m)
# + id="Ql9Ty9wmFxfl" colab={"base_uri": "https://localhost:8080/"} outputId="65e140ab-f886-4d47-d5ee-b94e4b04be88"
# !pip install numpy
# + id="2BboMJYwGGaZ"
from numpy import *
print("-------------------------------------------------------------------------")
arr = array([1,2,3,4,5])
print(arr)
print("-------------------------------------------------------------------------")
arr1 = array([1,2,3,4,5.0]) #explicit conversion
print(arr1)
print("-------------------------------------------------------------------------")
arr2 = array([1,2,3,4,5],float)
print(arr2)
print("-------------------------------------------------------------------------")
arr3 = array([1,2,3,4,5])
print(arr3.dtype)
print("-------------------------------------------------------------------------")
arr4 = linspace(1,5,10) #5 included.... 10 is number of divisions
print(arr4)
print("-------------------------------------------------------------------------")
arr4 = arange(1,50,2) # just like in range
print(arr4)
print("-------------------------------------------------------------------------")
arr5 = logspace(1,5,10) #gap with 1 to the power of 10 to 5 to the power of 10
print(arr5)
print("-------------------------------------------------------------------------")
print('%.2f' %arr5[0]) #2f denotes upto two digit of float number at index number 0
arr6 = zeros(5)#5 denotes the sixe of array and by default it is of float
print(arr6)
print("-------------------------------------------------------------------------")
arr7 = ones(5,int)
print(arr7)
# + id="voQbrZ77dkCY"
from numpy import *
#numerical operation in numpy
arr1= array([1,2,3,4,5])
arr2= array([1,4,6,8,2])
arr1 = arr1 + 1
print(arr1)
print(arr2)
arr3 = arr1 + arr2
print(arr3)
print('Joining two array we use "concatenate" function')
print(concatenate([arr1,arr2]))
# + id="W-Uyo3ZXf4MO"
#functions in numpy array
arr1= array([1,2,3,4,5,5,5,90,30])
print(sin(arr1)) #'''sin/cos/log/sqrt/sum(sum of all element of the same array/
#min/max/sort/unique
print(sort(arr1))
print(unique(arr1))
# + id="XRQgwYMJiBj8"
#copying an array
# + id="9AUcOU39zCuq"
#functions
def add(a,b): #a,b are arguments (1. Formal Arguments)
c=a+b
print(c)
add(5,6) #5,6 are 2. Actual Arguments
# + id="QuJYcCyXzt-7"
'''lets say i want to add with multiple arguments i'll use '*' in arguments
which while passing the arguments will convert it into tuples'''
def add(*a):
c=0
for i in a:
c=c+i
return c
add(5,6,8,581)
# + id="4nJidrzT2vjk"
''' there is no pass by value or pass by reference in python'''
def update(x):
x=8
print(id(x))
print("x", x)
update(10)
a=11
print(id(a))
update(a)
print(id(a))
# + id="X0VuC1Z034WK"
''' lets say i don't know the sequence in the def '''
def person(name,age=18):
print("name = ",name)
print("age = ",age)
person("aseem", 19)
person(19, "aseem") #sequence is wrong here
'''if i dont know the sequence then i can use keywords to
exactly define to a particular argument'''
person(name="Aseem", age=19)
#lets say i didnt passed the 2nd argument i.e age it
#will automatically take that by default if its written in defs arguments
person("aseem")
# + id="-ERB6Emr6Xzj"
#making 1d 2d 3d matrices or determinants in array
from numpy import *
arr1= array([
[1,2,3,4,5,6],
[2,3,5,6,3,5]
])
arr2=arr1.flatten()#2d to 1d array
arr3=arr1.reshape(3,4)#changes the 2d array row and column
arr4=arr1.reshape(2,2,3) #2 2d array and each 2d array having 2 rows and 3 columns
m=matrix(arr1) #gives matrix
print(arr1)
print(arr1.dtype)#type of array
print(arr1.ndim)# no. of dimention/ ranking
print(arr1.shape)#shape of 2d array
print(arr1.size)#row x column
print(arr2)
print(arr3)
print(arr4)
print(m)# now its matrix (loks same as 2d array) we can perform various matrix operations
# + id="tkwJVqjnnc20"
#concept of global and local variable
a = 10
def something():
#global a #makes
a=15
print("in func", a)
something()
print('outer', a)
# + id="B3GCoN4bSU-f"
lst=[]
if __name__ == '__main__':
for _ in range(int(input())):
name = input()
score = float(input())
lst.append([name,score])
print(max(lst))
# + id="l05IuGiphxMU"
#OOPs
class Employee:
no_of_leaves = 8
pass
harry = Employee()
rohan = Employee()
harry.name = "Harry"
harry.salary = 455
harry.role = "Instructor"
rohan.name = "Rohan"
rohan.salary = 4554
rohan.role = "Student"
print(Employee.no_of_leaves)
print(Employee.__dict__)
Employee.no_of_leaves = 9
print(Employee.__dict__)
print(Employee.no_of_leaves)
# + colab={"base_uri": "https://localhost:8080/", "height": 208} id="221q275cuCrr" outputId="09d5375c-77d3-47f6-dbe4-7fec0d7506b6"
#Decorator
def dec1(func1):
def nowexec():
print('Before executing the func1')
func1()
print("After executing the func1")
return nowexec
@dec1
def aseem():
print("I am Aseem")
aseem()
# + colab={"base_uri": "https://localhost:8080/"} id="fLimq8jwLyu9" outputId="dc79dee9-c945-4346-f1a5-0724d7e5a3d0"
def aseem():
print("I am Aseem")
aseem()
# + id="Ib6SLBvQIqhj"
class Employee:
def __init__(self, aname, asalary, arole): #"Constructor" that helps to give arguments to Employee in harry.Employee(*arguments*)
self.name = aname
self.salary = asalary
self.role = arole
def printDetails(self):
return f"Name is {self.name}. Salaray is {self.salary}. Role is {self.role}"
harry = Employee("Harry", 455, "Instructor")
larry = Employee("Larry", 125, "Manager")
print(harry.salary)
# + colab={"base_uri": "https://localhost:8080/", "height": 226} id="0bDeDiYkL6p3" outputId="1ff35362-8045-4012-9d27-10eab88402a3"
class Employee:
leaves=10
def __init__(self, aname, asalary, arole):
self.name=aname
self.salary=asalary
self.role=arole
def printDetails(self):
return f"Name {self.name}. Salary {self.salary}. Role {self.role}"
@classmethod
def change_leaves(cls, newleaves):
cls.leaves=newleaves
harry=Employee("Harry", 9999, "Instructor")
#print(harry.printDetails())
harry.change_leaves(30)
print(harry.leaves())
# + id="zfXftS9HSJrc"
class Employee:
no_of_leaves = 8
def __init__(self, aname, asalary, arole):
self.name = aname
self.salary = asalary
self.role = arole
def printdetails(self):
return f"The Name is {self.name}. Salary is {self.salary} and role is {self.role}"
@classmethod
def change_leaves(cls, newleaves):
cls.no_of_leaves = newleaves
| T_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import arcpy, os, datetime
from arcpy import env
env.workspace = r"T:\MPO\RTP\FY20 2045 Update\Data and Resources\Network_Analysis\Network_Analysis.gdb"
env.overwriteOutput = True
arcpy.management.Dissolve("SAWalkingTransit", "SAWalkingTransit_Dissolve", None, None, "MULTI_PART", "DISSOLVE_LINES")
arcpy.management.Dissolve("SAWalkingHighFreqTransit", "SAWalkingHighFreqTransit_Dis", None, None,
"MULTI_PART", "DISSOLVE_LINES")
selectedParcel = arcpy.management.SelectLayerByLocation("parcels_FeatureToPoint", "COMPLETELY_WITHIN",
"SAWalkingTransit_Dissolve",
None, "SUBSET_SELECTION", "NOT_INVERT")
fieldsum = arcpy.da.TableToNumPyArray(selectedParcel, "hh", skip_nulls=True)
val = fieldsum["hh"].sum()
val
| analysis/spatial_analysis/households_transit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/granularus/Fibonacci/blob/main/Fibonacci_and_the_Golden_Ratio.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UuIdHUVgJuiG"
# First we'll caculate some terms in the Fibonacci sequence, each term is the sum of the two that precede it. This code is in the official python tutorial.
# + id="ybtQC8kHJoXZ" colab={"base_uri": "https://localhost:8080/"} outputId="7c5851f3-f30e-4064-cb50-767f5fda60c9"
a, b=0, 1 # the first two terms, each subequent term is the sum of the two previous terms
while a<10:
print(a)
a, b =b, a+b
# + [markdown] id="vCiB2hGEKEB8"
# Here is a version with the output cleaned up, I've used the command end to replace the newline usually generated by a comma.
# + id="fC0Knh1GKLHj" colab={"base_uri": "https://localhost:8080/"} outputId="8d2255b1-0a48-4cd7-9919-9b33cb0afb56"
a, b=0, 1 # the first two terms, each subequent term is the sum of the two previous terms
while a<10:
print(a, end=',')
a, b =b, a+b
# + [markdown] id="ddYmR52lMQRF"
# Here is a quick calculation of the Golden Ratio that someone asked about yesterday. For those who don't know what the Golden Ratio is, I'll return to it if I have time.
# + colab={"base_uri": "https://localhost:8080/"} id="hc6eO63RMn2B" outputId="57373fa4-835d-434f-b488-cc68f2bee72d"
a, b=0, 1 # the first two terms, each subequent term is the sum of the two previous terms
while a<10000:
print(a)
if a>0:
print("Golden Ratio", b/a)
a, b =b, a+b
# + [markdown] id="86B1bzs_Jto_"
# Yesterday, someone asked about the golden ratio, we can easily investigate this, but to make it clearer we need a version of the code that saves the results of prior iterations. We can do this. In the text below, I use a new method called range. range will return an object that produces a sequence of integers from start (inclusive) to stop (exclusive) by step. range(i, j) produces i, i+1, i+2, ..., j-1. Start defaults to 0 when it is omitted! range(4) produces 0, 1, 2, 3. This time I'm using a list that is initialized, but it is the same basic method that I used in the calendar problem.
# + colab={"base_uri": "https://localhost:8080/"} id="wEnxYHKoLfmk" outputId="cee238b5-a915-4fff-87fc-2c86cbe3a59f"
# Fibonacci series using an iterative method, with values saved in a list
fiblist = [0,1]
for i in range(10):
fiblist.append(fiblist[i] + fiblist[i+1])
print (fiblist)
# + [markdown] id="r_PhStT6ONXl"
# Now what is the Golden Ratio. Mathematically, two quantities are in the golden ratio if their ratio is the same as the ratio of their sum to the larger of the two quantities. Expressed algebraically, for quantities a and b with a > b > 0,
# this means $\phi$=$\frac{a}{b}=\frac{(a+b)}{a}=1.618033...$. (See Wikipedia if you want to know more). The ratio of successive terms in the Fibonacci sequence tends to the Golden Ratio. We will verify this.
# + colab={"base_uri": "https://localhost:8080/"} id="e_HGnEsFP_Uk" outputId="0c514977-0d6d-4ccb-d628-3a22a1e55c61"
# Fibonacci series using an iterative method, with values saved in a list
#Including a calculation of the Golden Ratio
fiblist = [0,1]
for i in range(1000):
fiblist.append(fiblist[i] + fiblist[i+1])
gratio=[fiblist[i] / fiblist[i-1] for i in range(2,len(fiblist))]
print (fiblist)
print (gratio)
# + [markdown] id="d6wcsOquXgFg"
# I found a version of this code written in Python 2 and rewrote it in Python 3, the major changes were the print statements, and that I was able to omit some float statements because divison always yields floating point numbers in Python 3. Modifying existing code is a great way to learn. You might notice that this uses the methods that I used in the equinox problem. The original code is here:
# http://pi3.sites.sheffield.ac.uk/tutorials/week-1-fibonacci
#
| Fibonacci_and_the_Golden_Ratio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="dvN2-COJ9w4k"
# # Introduction
#
# In this notebook, we train [YOLOv4 tiny](https://github.com/AlexeyAB/darknet/issues/6067) on custom data. We will convert this to a TensorFlow representation and finally TensorFlow Lite file to use on device.
#
# We also recommend reading our blog post on [How To Train YOLOv4 And Convert It To TensorFlow (And TensorFlow Lite!)](https://blog.roboflow.ai/how-to-train-yolov4-and-convert-it-to-tensorflow) side by side.
#
# We will take the following steps to get YOLOv4 from training on custom data to a TensorFlow (and TensorFlow Lite) representation:
#
#
# 1. Set up the Custom Dataset
# 2. Train the Model with Darknet
# 3. Convert the weights to TensorFlow's .pb representation
# 4. Convert the weights to TensorFlow Lite's .tflite representation
#
#
# When you are done you will have a custom detector that you can use. It will make inference like this:
#
# #### 
#
# ### **Reach out for support**
#
# If you run into any hurdles on your own data set or just want to share some cool results in your own domain, [reach out!](https://roboflow.ai)
#
#
#
# #### 
# + [markdown] id="ef1_2DwjF9Cy"
# #1. Set up the Custom Dataset
#
#
# + [markdown] id="zbniFj-eSimL"
# We'll use Roboflow to convert our dataset from any format to the YOLO Darknet format.
#
# 1. To do so, create a free [Roboflow account](https://app.roboflow.ai).
# 2. Upload your images and their annotations (in any format: VOC XML, COCO JSON, TensorFlow CSV, etc).
# 3. Apply preprocessing and augmentation steps you may like. We recommend at least `auto-orient` and a `resize` to 416x416. Generate your dataset.
# 4. Export your dataset in the **YOLO Darknet format**.
# 5. Copy your download link, and paste it below.
#
# See our [blog post](https://blog.roboflow.ai/training-yolov4-on-a-custom-dataset/) for greater detail.
#
# In this example, I used the open source [BCCD Dataset](https://public.roboflow.ai/object-detection/bccd). (You can `fork` it to your Roboflow account to follow along.)
# + id="Cdj4tmT5Cmdl" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134704785, "user_tz": -180, "elapsed": 3494, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="e2910e80-0234-4285-b8fb-8b5770d1bc13"
#if you already have YOLO darknet format, you can skip this step
#otherwise we recommend formatting in Roboflow
# %cd /content
# %mkdir dataset
# %cd ./dataset
# !curl -L "https://app.roboflow.com/ds/iVF4a6uj2L?key=<KEY>" > roboflow.zip; unzip roboflow.zip; rm roboflow.zip
# + [markdown] id="nKlSotFT9Zv-"
# #2. Train a Custom Model on DarkNet
# + [markdown] id="ZefjbxBqAZ10"
# ***Since we already have a [notebook](https://colab.research.google.com/drive/1PWOwg038EOGNddf6SXDG5AsC8PIcAe-G#scrollTo=NjKzw2TvZrOQ) on how to train YOLOv4 with Darknet, we have simply included the contents here as well.***
# + [markdown] id="GNVU7eu9CQj3"
# ## Introduction
#
#
# In this notebook, we implement the tiny version of [YOLOv4](https://arxiv.org/pdf/2004.10934.pdf) for training on your own dataset, [YOLOv4 tiny](https://github.com/AlexeyAB/darknet/issues/6067).
#
# We also recommend reading our blog post on [Training YOLOv4 on custom data](https://blog.roboflow.ai/training-yolov4-on-a-custom-dataset/) side by side.
#
# We will take the following steps to implement YOLOv4 on our custom data:
# * Configure our GPU environment on Google Colab
# * Install the Darknet YOLOv4 training environment
# * Download our custom dataset for YOLOv4 and set up directories
# * Configure a custom YOLOv4 training config file for Darknet
# * Train our custom YOLOv4 object detector
# * Reload YOLOv4 trained weights and make inference on test images
#
# When you are done you will have a custom detector that you can use. It will make inference like this:
#
# #### 
#
# ### **Reach out for support**
#
# If you run into any hurdles on your own data set or just want to share some cool results in your own domain, [reach out!](https://roboflow.ai)
#
#
#
# #### 
# + [markdown] id="mDTvGt2zt7cm"
# ## Configuring CUDA on Colab for YOLOv4
#
#
# + id="u-bguKWgtxSx" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134704786, "user_tz": -180, "elapsed": 106, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="a7dad940-caff-410e-aa8f-a8488cf8f726"
# CUDA: Let's check that Nvidia CUDA drivers are already pre-installed and which version is it. This can be helpful for debugging.
# !/usr/local/cuda/bin/nvcc --version
# + [markdown] id="id9hqvGjGdFf"
# **IMPORTANT!** If you're not training on a Tesla P100 GPU, we will need to tweak our Darknet configuration later based on what type of GPU we have. Let's set that now while we're inspecting the GPU.
# + id="o6BRAVo182G5" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134704786, "user_tz": -180, "elapsed": 21, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="42a5eab8-ef45-41a4-9921-ec6b73d02343"
#take a look at the kind of GPU we have
# !nvidia-smi
# + id="z5FkXxGmGK4Q" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134710724, "user_tz": -180, "elapsed": 530, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="a01ccd9f-22c1-483f-98a0-fe0d9b9416e5"
# Change the number depending on what GPU is listed above, under NVIDIA-SMI > Name.
# Tesla K80: 30
# Tesla P100: 60
# Tesla T4: 75
# %env compute_capability=75
# + [markdown] id="16pvdFMa1FEe"
# ## Installing Darknet for YOLOv4 on Colab
#
#
#
# + id="K9uY-38P93oz" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134711448, "user_tz": -180, "elapsed": 416, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="f8e58d2d-d4eb-4b52-fe2a-405ed93b41ce"
# %cd /content/
# %rm -rf darknet
# + id="HQEktcfj9y9O" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134721967, "user_tz": -180, "elapsed": 5328, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="33a79e04-74fb-4d95-8e01-5163909d18ed"
#we clone the fork of darknet maintained by roboflow
#small changes have been made to configure darknet for training
# !git clone https://github.com/roboflow-ai/darknet.git
# + [markdown] id="7FS9Fd4-Yi8-"
# **IMPORTANT! If you're not using a Tesla P100 GPU**, then uncomment the sed command and replace the arch and code with that matching your GPU. A list can be found [here](http://arnon.dk/matching-sm-architectures-arch-and-gencode-for-various-nvidia-cards/). You can check with the command nvidia-smi (should be run above).
# + id="QyMBDkaL-Aep" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134768973, "user_tz": -180, "elapsed": 47016, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="3b04c1db-2a3e-442c-cb02-49e1c066c84f"
#install environment from the Makefile. Changes to mitigate CUDA error.
# %cd darknet/
# !sed -i 's/OPENCV=0/OPENCV=1/g' Makefile
# !sed -i 's/GPU=0/GPU=1/g' Makefile
# !sed -i 's/CUDNN=0/CUDNN=1/g' Makefile
# !sed -i "s/ARCH= -gencode arch=compute_60,code=sm_60/ARCH= -gencode arch=compute_${compute_capability},code=sm_${compute_capability}/g" Makefile
# !make
# + id="LGPDEjfAALrQ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134770399, "user_tz": -180, "elapsed": 1446, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="903c3b0c-1294-4bc8-dbb8-b6a909c080d1"
#download the newly released yolov4-tiny weights
# %cd /content/darknet
# !wget https://github.com/AlexeyAB/darknet/releases/download/darknet_yolo_v4_pre/yolov4-tiny.weights
# !wget https://github.com/AlexeyAB/darknet/releases/download/darknet_yolo_v4_pre/yolov4-tiny.conv.29
# + [markdown] id="oh-vu9eWSyYw"
# ## Configure from Custom Dataset
# + id="CFciBw3iSwqt" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134771686, "user_tz": -180, "elapsed": 1291, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="25951968-cd79-42db-c9b2-78dae5e254b4"
#Copy dataset
# %cp -r /content/dataset/. /content/darknet/
#Set up training file directories for custom dataset
# %cd /content/darknet/
# %cp train/_darknet.labels data/obj.names
# %mkdir data/obj
# #copy image and labels
# %cp train/*.jpg data/obj/
# %cp valid/*.jpg data/obj/
# %cp train/*.txt data/obj/
# %cp valid/*.txt data/obj/
with open('data/obj.data', 'w') as out:
out.write('classes = 3\n')
out.write('train = data/train.txt\n')
out.write('valid = data/valid.txt\n')
out.write('names = data/obj.names\n')
out.write('backup = backup/')
#write train file (just the image list)
import os
with open('data/train.txt', 'w') as out:
for img in [f for f in os.listdir('train') if f.endswith('jpg')]:
out.write('data/obj/' + img + '\n')
#write the valid file (just the image list)
import os
with open('data/valid.txt', 'w') as out:
for img in [f for f in os.listdir('valid') if f.endswith('jpg')]:
out.write('data/obj/' + img + '\n')
# + [markdown] id="5HtRqO3QvjkP"
# ## Write Custom Training Config for YOLOv4
# + id="U_WJcqHhpeVr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627134771687, "user_tz": -180, "elapsed": 10, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="b2d58dda-15b0-4d4c-e790-3ba2c16a3851"
#we build config dynamically based on number of classes
#we build iteratively from base config files. This is the same file shape as cfg/yolo-obj.cfg
def file_len(fname):
with open(fname) as f:
for i, l in enumerate(f):
pass
return i + 1
num_classes = file_len('train/_darknet.labels')
max_batches = num_classes*2000
steps1 = .8 * max_batches
steps2 = .9 * max_batches
steps_str = str(steps1)+','+str(steps2)
num_filters = (num_classes + 5) * 3
print("writing config for a custom YOLOv4 detector detecting number of classes: " + str(num_classes))
#Instructions from the darknet repo
#change line max_batches to (classes*2000 but not less than number of training images, and not less than 6000), f.e. max_batches=6000 if you train for 3 classes
#change line steps to 80% and 90% of max_batches, f.e. steps=4800,5400
if os.path.exists('./cfg/custom-yolov4-tiny-detector.cfg'): os.remove('./cfg/custom-yolov4-tiny-detector.cfg')
#customize iPython writefile so we can write variables
from IPython.core.magic import register_line_cell_magic
@register_line_cell_magic
def writetemplate(line, cell):
with open(line, 'w') as f:
f.write(cell.format(**globals()))
# + id="03VuD4NHnxFx" executionInfo={"status": "ok", "timestamp": 1627134771688, "user_tz": -180, "elapsed": 7, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}}
# %%writetemplate ./cfg/custom-yolov4-tiny-detector.cfg
[net]
# Testing
#batch=1
#subdivisions=1
# Training
batch=64
subdivisions=16
width=416
height=416
channels=3
momentum=0.9
decay=0.0005
angle=0
saturation = 1.5
exposure = 1.5
hue=.1
learning_rate=0.00261
burn_in=1000
max_batches = {max_batches}
policy=steps
steps={steps_str}
scales=.1,.1
[convolutional]
batch_normalize=1
filters=32
size=3
stride=2
pad=1
activation=leaky
[convolutional]
batch_normalize=1
filters=64
size=3
stride=2
pad=1
activation=leaky
[convolutional]
batch_normalize=1
filters=64
size=3
stride=1
pad=1
activation=leaky
[route]
layers=-1
groups=2
group_id=1
[convolutional]
batch_normalize=1
filters=32
size=3
stride=1
pad=1
activation=leaky
[convolutional]
batch_normalize=1
filters=32
size=3
stride=1
pad=1
activation=leaky
[route]
layers = -1,-2
[convolutional]
batch_normalize=1
filters=64
size=1
stride=1
pad=1
activation=leaky
[route]
layers = -6,-1
[maxpool]
size=2
stride=2
[convolutional]
batch_normalize=1
filters=128
size=3
stride=1
pad=1
activation=leaky
[route]
layers=-1
groups=2
group_id=1
[convolutional]
batch_normalize=1
filters=64
size=3
stride=1
pad=1
activation=leaky
[convolutional]
batch_normalize=1
filters=64
size=3
stride=1
pad=1
activation=leaky
[route]
layers = -1,-2
[convolutional]
batch_normalize=1
filters=128
size=1
stride=1
pad=1
activation=leaky
[route]
layers = -6,-1
[maxpool]
size=2
stride=2
[convolutional]
batch_normalize=1
filters=256
size=3
stride=1
pad=1
activation=leaky
[route]
layers=-1
groups=2
group_id=1
[convolutional]
batch_normalize=1
filters=128
size=3
stride=1
pad=1
activation=leaky
[convolutional]
batch_normalize=1
filters=128
size=3
stride=1
pad=1
activation=leaky
[route]
layers = -1,-2
[convolutional]
batch_normalize=1
filters=256
size=1
stride=1
pad=1
activation=leaky
[route]
layers = -6,-1
[maxpool]
size=2
stride=2
[convolutional]
batch_normalize=1
filters=512
size=3
stride=1
pad=1
activation=leaky
##################################
[convolutional]
batch_normalize=1
filters=256
size=1
stride=1
pad=1
activation=leaky
[convolutional]
batch_normalize=1
filters=512
size=3
stride=1
pad=1
activation=leaky
[convolutional]
size=1
stride=1
pad=1
filters={num_filters}
activation=linear
[yolo]
mask = 3,4,5
anchors = 10,14, 23,27, 37,58, 81,82, 135,169, 344,319
classes={num_classes}
num=6
jitter=.3
scale_x_y = 1.05
cls_normalizer=1.0
iou_normalizer=0.07
iou_loss=ciou
ignore_thresh = .7
truth_thresh = 1
random=0
nms_kind=greedynms
beta_nms=0.6
[route]
layers = -4
[convolutional]
batch_normalize=1
filters=128
size=1
stride=1
pad=1
activation=leaky
[upsample]
stride=2
[route]
layers = -1, 23
[convolutional]
batch_normalize=1
filters=256
size=3
stride=1
pad=1
activation=leaky
[convolutional]
size=1
stride=1
pad=1
filters={num_filters}
activation=linear
[yolo]
mask = 1,2,3
anchors = 10,14, 23,27, 37,58, 81,82, 135,169, 344,319
classes={num_classes}
num=6
jitter=.3
scale_x_y = 1.05
cls_normalizer=1.0
iou_normalizer=0.07
iou_loss=ciou
ignore_thresh = .7
truth_thresh = 1
random=0
nms_kind=greedynms
beta_nms=0.6
# + id="u2LAciMh4Cut" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627131109049, "user_tz": -180, "elapsed": 15, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="7536eab5-68fb-421d-bdb8-d40780972431"
#here is the file that was just written.
#you may consider adjusting certain things
#like the number of subdivisions 64 runs faster but Colab GPU may not be big enough
#if Colab GPU memory is too small, you will need to adjust subdivisions to 16
# %cat cfg/custom-yolov4-tiny-detector.cfg
# + [markdown] id="vWrG9EGamSpH"
# ## Train Custom YOLOv4 Detector
# + id="6miYFbvExqMd" colab={"base_uri": "https://localhost:8080/"} outputId="5a4db500-18f3-4874-966f-d2e98cfbf29b"
# !./darknet detector train data/obj.data cfg/custom-yolov4-tiny-detector.cfg yolov4-tiny.conv.29 -dont_show -map
#If you get CUDA out of memory adjust subdivisions above!
#adjust max batches down for shorter training above
# + [markdown] id="nBnwpBV5ZXxQ"
# ## Infer Custom Objects with Saved YOLOv4 Weights
# + id="FzoJQQw8Zdco"
#define utility function
def imShow(path):
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
image = cv2.imread(path)
height, width = image.shape[:2]
resized_image = cv2.resize(image,(3*width, 3*height), interpolation = cv2.INTER_CUBIC)
fig = plt.gcf()
fig.set_size_inches(18, 10)
plt.axis("off")
#plt.rcParams['figure.figsize'] = [10, 5]
plt.imshow(cv2.cvtColor(resized_image, cv2.COLOR_BGR2RGB))
plt.show()
# + id="e3dJB6NZv4kh"
#check if weigths have saved yet
#backup houses the last weights for our detector
#(file yolo-obj_last.weights will be saved to the build\darknet\x64\backup\ for each 100 iterations)
#(file yolo-obj_xxxx.weights will be saved to the build\darknet\x64\backup\ for each 1000 iterations)
#After training is complete - get result yolo-obj_final.weights from path build\darknet\x64\bac
# !ls backup
#if it is empty you haven't trained for long enough yet, you need to train for at least 100 iterations
# + id="0gwfboR8Es-W" colab={"base_uri": "https://localhost:8080/", "height": 518} executionInfo={"status": "error", "timestamp": 1625578777255, "user_tz": -180, "elapsed": 10723, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="7b357827-3b0a-4e10-e75e-a4fc517faa5c"
#save final weights to google drive
from google.colab import drive
drive.mount('/content/drive')
# + id="FV7y6ZIP7aSx"
# Darknet Weights
# !cp /content/darknet/backup/custom-yolov4-tiny-detector_final.weights "/content/drive/My Drive"
# + id="x-_E3O5Mf4Mf"
#coco.names is hardcoded somewhere in the detector
# %cp data/obj.names data/coco.names
# + id="NjKzw2TvZrOQ"
#/test has images that we can test our detector on
test_images = [f for f in os.listdir('test') if f.endswith('.jpg')]
import random
img_path = "test/" + random.choice(test_images);
#test out our detector!
# !./darknet detect cfg/custom-yolov4-tiny-detector.cfg backup/custom-yolov4-tiny-detector_final.weights {img_path} -dont-show
imShow('predictions.jpg')
# + [markdown] id="d5m2lgRUHBWj"
# #3. Convert the weights to TensorFlow's .pb representation
# + [markdown] id="cHHTvG8bHLiG"
# Darknet produces a .weights file specific to Darknet. If we want to use the YOLOv4 model in TensorFlow, we'll need to convert it.
#
# To do this, we'll use the following tool: https://github.com/hunglc007/tensorflow-yolov4-tflite.
# + [markdown] id="rJvP691HHrOm"
# ## Install and Configure
# + [markdown] id="r3GUvX21Hyoe"
# First, we'll clone the repository.
# + id="GX277FMyH1Sz"
# %cd /content
# %rm -rf tensorflow-yolov4-tflite
# !git clone https://github.com/hunglc007/tensorflow-yolov4-tflite.git
# %cd /content/tensorflow-yolov4-tflite
# + [markdown] id="PMRigsA_FC74"
# Then, we'll change the labels from the default COCO to our own custom ones.
# + id="ntIqg1-x9i3a"
# !cp /content/darknet/data/obj.names /content/tensorflow-yolov4-tflite/data/classes/
# !ls /content/tensorflow-yolov4-tflite/data/classes/
# + id="59XOp9cA-G0S"
# !sed -i "s/coco.names/obj.names/g" /content/tensorflow-yolov4-tflite/core/config.py
# + [markdown] id="aWIyKzuYLeC0"
# ## Convert
# + [markdown] id="ncsA58syFMVf"
# Time to convert! We'll convert to both a regular TensorFlow SavedModel and to TensorFlow Lite. For TensorFlow Lite, we'll convert to a different TensorFlow SavedModel beforehand.
# + id="wturTKvcLgix"
# %cd /content/tensorflow-yolov4-tflite
# Regular TensorFlow SavedModel
# !python save_model.py \
# --weights /content/darknet/backup/custom-yolov4-tiny-detector_final.weights \
# --output ./checkpoints/yolov4-tiny-416 \
# --input_size 416 \
# --model yolov4 \
# --tiny \
#
# SavedModel to convert to TFLite
# !python save_model.py \
# --weights /content/darknet/backup/custom-yolov4-tiny-detector_final.weights \
# --output ./checkpoints/yolov4-tiny-pretflite-416 \
# --input_size 416 \
# --model yolov4 \
# --tiny \
# --framework tflite
# + [markdown] id="3Jdcq5seHD22"
# #4. Convert the TensorFlow weights to TensorFlow Lite
# + [markdown] id="i5BApg9dOwpM"
# From the generated TensorFlow SavedModel, we will convert to .tflite
# + id="uggKjSzoOrMK"
# %cd /content/tensorflow-yolov4-tflite
# !python convert_tflite.py --weights ./checkpoints/yolov4-tiny-pretflite-416 --output ./checkpoints/yolov4-tiny-416.tflite
# + id="jOJz36iOGS0G"
# !ls /content/darknet/test
# + id="vUPDhisfF-b5"
# Verify
# %cd /content/tensorflow-yolov4-tflite
# !python detect.py --weights ./checkpoints/yolov4-tiny-416 --size 416 --model yolov4 \
# --image /content/darknet/test/fcbbda4f0678bfcf_jpg.rf.732c1c2a76d23be01587721f43b0e9bf.jpg \
# # --framework tflite
# + id="L3PL0QTpIZwe"
# %cd /content/tensorflow-yolov4-tflite/
# !ls ./checkpoints
# !cp -a ./checkpoints/* /content/drive/MyDrive/tf_models/vehicle_recognition
from IPython.display import Image
Image('/content/tensorflow-yolov4-tflite/result.png')
# + [markdown] id="HaJjTg2HPEX3"
# # Save your Model
# + [markdown] id="Fv9Zc5loPbL5"
# You can save your model to your Google Drive for further use.
# + colab={"base_uri": "https://localhost:8080/"} id="LXHfcjmrZOc7" executionInfo={"status": "ok", "timestamp": 1624613353566, "user_tz": -180, "elapsed": 1637, "user": {"displayName": "\u05e8\u05d5\u05e2\u05d9 \u05d1\u05df \u05d9\u05d5\u05e1\u05e3", "photoUrl": "", "userId": "01149447703223654261"}} outputId="85e82454-4f5c-49bc-9468-0a208b312a43"
# !ls ./checkpoints
# !cd /content
# !cp -r /content/tensorflow-yolov4-tflite/* /content/drive/MyDrive/vehicle_detection_project/ver2
# + id="he7-AJfJPh-K"
# Choose what to copy
# TensorFlow SavedModel
# !cp -r /content/tensorflow-yolov4-tflite/checkpoints/yolov4-tiny-416/ "/content/drive/My Drive"
# TensorFlow Lite
# !cp /content/tensorflow-yolov4-tflite/checkpoints/yolov4-tiny-416.tflite "/content/drive/My Drive"
| colab_notebook_for_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sampath11/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/Sam_Kumar_LS_DS_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="11OzdxWTM7UR" colab_type="text"
# ## Assignment - Build a confidence interval
#
# A confidence interval refers to a neighborhood around some point estimate, the size of which is determined by the desired p-value. For instance, we might say that 52% of Americans prefer tacos to burritos, with a 95% confidence interval of +/- 5%.
#
# 52% (0.52) is the point estimate, and +/- 5% (the interval $[0.47, 0.57]$) is the confidence interval. "95% confidence" means a p-value $\leq 1 - 0.95 = 0.05$.
#
# In this case, the confidence interval includes $0.5$ - which is the natural null hypothesis (that half of Americans prefer tacos and half burritos, thus there is no clear favorite). So in this case, we could use the confidence interval to report that we've failed to reject the null hypothesis.
#
# But providing the full analysis with a confidence interval, including a graphical representation of it, can be a helpful and powerful way to tell your story. Done well, it is also more intuitive to a layperson than simply saying "fail to reject the null hypothesis" - it shows that in fact the data does *not* give a single clear result (the point estimate) but a whole range of possibilities.
#
# How is a confidence interval built, and how should it be interpreted? It does *not* mean that 95% of the data lies in that interval - instead, the frequentist interpretation is "if we were to repeat this experiment 100 times, we would expect the average result to lie in this interval ~95 times."
#
# For a 95% confidence interval and a normal(-ish) distribution, you can simply remember that +/-2 standard deviations contains 95% of the probability mass, and so the 95% confidence interval based on a given sample is centered at the mean (point estimate) and has a range of +/- 2 (or technically 1.96) standard deviations.
#
# Different distributions/assumptions (90% confidence, 99% confidence) will require different math, but the overall process and interpretation (with a frequentist approach) will be the same.
#
# Your assignment - using the data from the prior module ([congressional voting records](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records)):
#
#
# ### Confidence Intervals:
# 1. Generate and numerically represent a confidence interval
# 2. Graphically (with a plot) represent the confidence interval
# 3. Interpret the confidence interval - what does it tell you about the data and its distribution?
#
# ### Chi-squared tests:
# 4. Take a dataset that we have used in the past in class that has **categorical** variables. Pick two of those categorical variables and run a chi-squared tests on that data
# - By hand using Numpy
# - In a single line using Scipy
#
# + id="GUk60wty-u0g" colab_type="code" colab={}
import pandas as pd
import numpy as np
from scipy.stats import ttest_ind, ttest_ind_from_stats, ttest_rel
import matplotlib.pyplot as plt
from scipy import stats
# + id="Ckcr4A4FM7cs" colab_type="code" colab={}
# TODO - your code!
from scipy import stats
def confidence_interval(data, confidence=0.95):
"""
Calculate a confidence interval around a sample mean for given data.
Using t-distribution and two-tailed test, default 95% confidence.
Arguments:
data - iterable (list or numpy array) of sample observations
confidence - level of confidence for the interval
Returns:
tuple of (mean, lower bound, upper bound)
"""
data = np.array(data)
mean = np.mean(data)
n = len(data)
stderr = stats.sem(data)
t = stats.t.ppf((1 + confidence) / 2.0, n - 1)
interval = stderr * t
return (mean, mean - interval, mean + interval)
# + id="jdjCBJ_8_X9j" colab_type="code" outputId="bb2ca9a2-3f6e-42e4-c754-9259f3e03e83" colab={"base_uri": "https://localhost:8080/", "height": 202}
# !wget "https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data"
# + id="9p3_dnLp_Dcc" colab_type="code" outputId="7678b99e-27ff-436a-caf4-15144a51bc29" colab={"base_uri": "https://localhost:8080/", "height": 266}
## Read the file
header_names = ["Party",
"handicapped-infants",
"water-project-cost-sharing",
"adoption-of-the-budget-resolutionAlcalinity of ash",
"physician-fee-freeze",
"el-salvador-aid",
"religious-groups-in-schools",
"anti-satellite-test-ban",
"aid-to-nicaraguan-contras",
"mx-missile",
"immigration",
"synfuels-corporation-cutback",
"education-spending"
"superfund-right-to-sue",
"crime",
"duty-free-exports",
"export-administration-act-south-africa"
]
df = pd.read_csv ("house-votes-84.data", names=header_names, header=None, index_col=False, na_values='?')
df.head()
# + id="iUyoF7KJ_evd" colab_type="code" colab={}
## Replace n, y
df = df.replace({"n":0, "y":int('1')})
# + id="b-237DDz_gmQ" colab_type="code" outputId="628682b1-f473-452b-980b-f5ac028fb184" colab={"base_uri": "https://localhost:8080/", "height": 266}
df.head()
# + id="sMJIRkEvgXrv" colab_type="code" colab={}
## remove na from handicapped infants column
df['handicapped-infants'] = df['handicapped-infants'].dropna()
# + id="6SH8ISOfAwdy" colab_type="code" outputId="e145e85b-9bbe-410b-8820-eb91af72e6cf" colab={"base_uri": "https://localhost:8080/", "height": 266}
#filter Dems party
df_dems = df[(df.Party =="democrat")]
df_dems.head()
# + id="gyKIjFesA1AZ" colab_type="code" outputId="ed3c4784-bbd7-410a-a7b1-e2f0ac83f877" colab={"base_uri": "https://localhost:8080/", "height": 266}
df_rep = df[df.Party == "republican"]
df_rep.head()
# + id="6e-JV79bB2NB" colab_type="code" colab={}
## remove NaN
arr_dem = df_dems['physician-fee-freeze'].dropna()
# + id="k2HXcAXHNrds" colab_type="code" colab={}
## remove NaN from rep as well
arr_rep = df_rep['physician-fee-freeze'].dropna()
# + id="uh-AtCi8N0eI" colab_type="code" outputId="f629d123-d535-4c70-c023-e1d34dd556e3" colab={"base_uri": "https://localhost:8080/", "height": 1000}
arr_rep
# + id="nLO0qDP1DPPF" colab_type="code" outputId="9fd669cd-0dba-4e95-c063-8d26da40c4d9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
arr_dem
# + id="0fMoN9UEA-1e" colab_type="code" outputId="cfef1632-7244-4a7d-bb38-4d9f21386bc3" colab={"base_uri": "https://localhost:8080/", "height": 34}
## We looked at this issue - physician-fee-freeze in earlier assignment
print ( confidence_interval (arr_dem, 0.95) )
# + id="DEF4hcS-FpPj" colab_type="code" outputId="ae845b92-2691-4739-f371-d489243e46d0" colab={"base_uri": "https://localhost:8080/", "height": 319}
# plot it
plt.hist (arr_dem, bins=10)
# + [markdown] id="ekZ9PnkGDmza" colab_type="text"
# ## As we can see from the above we get the mean of 0.054 (54%), with range of 0.026 (26%) - 0.08 (80%) with confidence interval of 95%
# + id="0J78KaykgIDP" colab_type="code" colab={}
## Another alternative way to calculate .... get into one array
df_handicap = df['handicapped-infants'].dropna()
# + id="z534MC6dhGvV" colab_type="code" outputId="75ad35f0-2c2f-46ed-c909-951a68e52105" colab={"base_uri": "https://localhost:8080/", "height": 118}
df_handicap.head()
# + id="Dmxdn1tRldTA" colab_type="code" outputId="be5ee114-5e26-4444-a255-29e376d8b1ca" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df_handicap
# + id="uxdDcdM0hMbK" colab_type="code" outputId="7aaf1b3c-7e79-4b28-c8d3-313a4051227c" colab={"base_uri": "https://localhost:8080/", "height": 286}
df_handicap.hist()
# + id="cuvNEC1wlLiS" colab_type="code" outputId="e61493ce-b83c-4cc7-8c06-6516c7285de0" colab={"base_uri": "https://localhost:8080/", "height": 34}
handicap_size = df_handicap.size
print (handicap_size)
# + id="9GV9uWgGhhMz" colab_type="code" outputId="6ed71823-ef50-4ddc-9a64-2b308f227207" colab={"base_uri": "https://localhost:8080/", "height": 34}
handicap_mean = df_handicap.mean()
handicap_std = np.std(df_handicap, ddof=1)
print(handicap_mean, handicap_std)
# + id="naPZhAndlEXR" colab_type="code" outputId="a77a36a9-f693-467f-c08f-d212a07ff736" colab={"base_uri": "https://localhost:8080/", "height": 34}
handicap_err = handicap_std/np.sqrt(handicap_size)
handicap_err
# + id="mTmugOcjlq9n" colab_type="code" outputId="d1b84b36-14d5-4faa-f9d7-970ca82f9481" colab={"base_uri": "https://localhost:8080/", "height": 34}
t = 1.984 # 95% confidence
(handicap_mean, handicap_mean - t*handicap_err, handicap_mean + t*handicap_err)
# + id="m8WWLfIJmAYT" colab_type="code" outputId="64fa41f2-d80d-4ca7-a2d6-791047b1ff7c" colab={"base_uri": "https://localhost:8080/", "height": 34}
confidence_interval(df_handicap, confidence=0.95)
# + [markdown] id="4DdrTMNgmRmJ" colab_type="text"
# ## 95% confidence interval for handicapped_infants issue is 0.39 and 0.49
# + id="gzaBApgWoEtP" colab_type="code" colab={}
## Lets get another issue and draw graph
df_water = df['water-project-cost-sharing'].dropna()
# + id="AGukI_KVpUsd" colab_type="code" outputId="7df854ae-7c31-4a75-e262-b65c1720115e" colab={"base_uri": "https://localhost:8080/", "height": 118}
df_water.head()
# + id="26w0vDthpXVM" colab_type="code" outputId="64dd6aae-120a-4e53-de9f-1536a3cb6db7" colab={"base_uri": "https://localhost:8080/", "height": 286}
df_water.hist()
# + id="YTGnATCVpaTr" colab_type="code" outputId="acfe03e6-238f-43d9-bb32-8b775e6478ac" colab={"base_uri": "https://localhost:8080/", "height": 34}
water_size = df_water.size
print (water_size)
# + id="t9s_kh6IpoLl" colab_type="code" outputId="92f7718b-924c-47e9-f03d-af4930a33b8a" colab={"base_uri": "https://localhost:8080/", "height": 34}
water_mean = df_water.mean()
water_std = np.std(df_water, ddof=1)
water_newstd = df_water.std()
print(water_mean, water_std, water_newstd)
# + id="j_FZ0cCbp2G-" colab_type="code" outputId="f7700c10-90ae-41a6-f403-aa24abad03fd" colab={"base_uri": "https://localhost:8080/", "height": 34}
water_err = water_std/np.sqrt(water_size)
water_err
# + id="gdp_3IfLqJyO" colab_type="code" outputId="6d823a10-e616-4793-e91a-9ab0234e8208" colab={"base_uri": "https://localhost:8080/", "height": 34}
t = 1.984 # 95% confidence
(water_mean, water_mean - t*water_err, water_mean + t*water_err)
# + id="MnLcpY9kqYeJ" colab_type="code" outputId="6b8198fb-8453-45b4-a1e4-20f55818033d" colab={"base_uri": "https://localhost:8080/", "height": 34}
confidence_interval(df_water, confidence=0.95)
# + id="yWWhosXfqyO1" colab_type="code" outputId="e25f1f8b-6407-4f67-f656-f6d7cf6573cc" colab={"base_uri": "https://localhost:8080/", "height": 269}
# build the bar plot showing means
import matplotlib.pyplot as plt
plt.bar(['handicap', 'water'],[handicap_mean, water_mean])
plt.show()
# + [markdown] id="98Jz2smzqnyT" colab_type="text"
# ## 95% Confidence interval is between 0.45 - 0.55
# + id="-nRRec6Eq7Oi" colab_type="code" outputId="2cd92e4a-6240-4b55-d5c8-262f30425eab" colab={"base_uri": "https://localhost:8080/", "height": 295}
plt.bar(['handicap', 'water'],[handicap_mean, water_mean], yerr=[0.5*handicap_err, 0.5*water_err])
plt.xlabel('Voting Issue')
plt.ylabel('Means)')
plt.title('Means of handicapped and water project yes/no votes with 95% CI')
plt.show()
# + id="su-txyXxH9Ql" colab_type="code" outputId="14e86e25-e414-4f67-f9f8-1d286bc928ae" colab={"base_uri": "https://localhost:8080/", "height": 1000}
help(stats.chi2_contingency)
# + id="s9CiuEw-PN2u" colab_type="code" outputId="d228d8b6-cd79-4016-a45e-07abbd10ccd5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
print (arr_dem)
# + id="PZaJvbbXQWz5" colab_type="code" outputId="20683d68-43eb-47a7-9041-f2b936112b0a" colab={"base_uri": "https://localhost:8080/", "height": 118}
arr_dem.head()
# + id="o2iH0V9yQ-s8" colab_type="code" outputId="7bdcbdc1-8428-4bce-cebe-f5d26dd14a78" colab={"base_uri": "https://localhost:8080/", "height": 118}
arr_rep.head()
# + id="dm_GUI3NRHl5" colab_type="code" colab={}
# + [markdown] id="4ohsJhQUmEuS" colab_type="text"
# ## Stretch goals:
#
# 1. Write a summary of your findings, mixing prose and math/code/results. *Note* - yes, this is by definition a political topic. It is challenging but important to keep your writing voice *neutral* and stick to the facts of the data. Data science often involves considering controversial issues, so it's important to be sensitive about them (especially if you want to publish).
# 2. Apply the techniques you learned today to your project data or other data of your choice, and write/discuss your findings here.
# 3. Refactor your code so it is elegant, readable, and can be easily run for all issues.
# + [markdown] id="nyJ3ySr7R2k9" colab_type="text"
# ## Resources
#
# - [Interactive visualize the Chi-Squared test](https://homepage.divms.uiowa.edu/~mbognar/applets/chisq.html)
# - [Calculation of Chi-Squared test statistic](https://en.wikipedia.org/wiki/Pearson%27s_chi-squared_test)
# - [Visualization of a confidence interval generated by R code](https://commons.wikimedia.org/wiki/File:Confidence-interval.svg)
# - [Expected value of a squared standard normal](https://math.stackexchange.com/questions/264061/expected-value-calculation-for-squared-normal-distribution) (it's 1 - which is why the expected value of a Chi-Squared with $n$ degrees of freedom is $n$, as it's the sum of $n$ squared standard normals)
| Sam_Kumar_LS_DS_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # A simple harmonic oscillator
#
# This demo implements a simple harmonic oscillator
# in a 2D neural population.
# The oscillator is more visually interesting on its own
# than the integrator, but the principle at work is the same.
# Here, instead of having the recurrent input just integrate
# (i.e. feeding the full input value back to the population),
# we have two dimensions which interact.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import nengo
from nengo.processes import Piecewise
# -
# ## Step 1: Create the Model
#
# The model consists of a single neural ensemble that we will call `Neurons`.
# Create the model object
model = nengo.Network(label="Oscillator")
with model:
# Create the ensemble for the oscillator
neurons = nengo.Ensemble(200, dimensions=2)
# ## Step 2: Provide Input to the Model
#
# A brief input signal is provided
# to trigger the oscillatory behavior of the neural representation.
with model:
# Create an input signal
input = nengo.Node(Piecewise({0: [1, 0], 0.1: [0, 0]}))
# Connect the input signal to the neural ensemble
nengo.Connection(input, neurons)
# Create the feedback connection
nengo.Connection(neurons, neurons, transform=[[1, 1], [-1, 1]], synapse=0.1)
# ## Step 3: Add Probes
#
# These probes will collect data from the input signal and the neural ensemble.
with model:
input_probe = nengo.Probe(input, "output")
neuron_probe = nengo.Probe(neurons, "decoded_output", synapse=0.1)
# ## Step 4: Run the Model
# Create the simulator
with nengo.Simulator(model) as sim:
# Run it for 5 seconds
sim.run(5)
# ## Step 5: Plot the Results
plt.figure()
plt.plot(sim.trange(), sim.data[neuron_probe])
plt.xlabel("Time (s)", fontsize="large")
plt.legend(["$x_0$", "$x_1$"])
data = sim.data[neuron_probe]
plt.figure()
plt.plot(data[:, 0], data[:, 1], label="Decoded Output")
plt.xlabel("$x_0$", fontsize=20)
plt.ylabel("$x_1$", fontsize=20)
plt.legend()
| docs/examples/dynamics/oscillator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
plt.style.use('ggplot')
customers=['ABC', 'DEF', 'GHI', 'JKL', 'MNO']
customers_index=range(len(customers))
sale_amounts = [127,90,201,111,232]
fig = plt.figure()
ax1 = fig.add_subplot(1,1,1)
ax1.bar(customers_index, sale_amounts, align='center', color='darkblue')
ax1.xaxis.set_ticks_position('bottom')
ax1.yaxis.set_ticks_position('left')
plt.xticks(customers_index, customers, rotation=0, fontsize='small')
plt.xlabel('Cusotmer Name')
plt.ylabel('Sale Amount')
plt.title('Sale Amount per Customer')
plt.savefig('bar_plot.png', dpi=400, bbox_inches='tight')
plt.show()
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
mu1,mu2,sigma=100,130,15
x1=mu1+sigma*np.random.randn(10000)
x2=mu2+sigma*np.random.randn(10000)
fig=plt.figure()
ax1=fig.add_subplot(1,1,1)
n,bins,patches=ax1.hist(x1, bins=50, normed=False, color='darkgreen')
n,bins,patches=ax1.hist(x2, bins=50, normed=False, color='orange', alpha=0.5)
ax1.xaxis.set_ticks_position('bottom')
ax1.yaxis.set_ticks_position('left')
plt.xlabel('Bins')
plt.ylabel('Number of Values in Bin')
fig.suptitle('Histograms', fontsize=14, fontweight='bold')
ax1.set_title('Two Frequency Distributions')
plt.savefig('histogram.png', dpi=400, bbox_inches='tight')
plt.show()
# +
import seaborn as sns
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
sns.set(color_codes=True)
#히스토그램
x=np.random.normal(size=1000)
sns.distplot(x, bins=20, kde=False, rug=True, label='Histogram w/o Density')
sns.utils.axlabel('Value', 'Frequency')
plt.title('Histogram of a Random Sample from a Normal Distribution')
plt.legend()
plt.show()
# +
import seaborn as sns
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
sns.set(color_codes=True)
#회귀선과 일변량 히스토그램을 포함한 산점도
mean, cov=[5,10],[(1,.5),(.5,1)]
data=np.random.multivariate_normal(mean, cov, 200)
data_frame=pd.DataFrame(data, columns=['x', 'y'])
sns.jointplot(x='x', y='y', data=data_frame, kind='reg').set_axis_labels('x', 'y')
plt.suptitle("Joint Plot of Two Variables with Bivariate and Univariate Graphs")
plt.show()
# -
#쌍별 이변량 산점도
iris=sns.load_dataset('iris')
sns.pairplot(iris)
plt.show()
# +
tips = sns.load_dataset("tips")
#여러 변수에 대한 상자그림
sns.factorplot(x='time', y='total_bill', hue='smoker', col='day', data=tips, kind='box', size=4, aspect=.5)
plt.show()
# -
#부스트트랩 신뢰구간을 포함한 선형회귀모형
sns.lmplot(x='total_bill', y='tip', data=tips)
plt.show()
#부트스트랩 신뢰구간을 포함한 로지스틱 회귀모형
tips['big_tip']=(tips.tip/tips.total_bill) > .15
sns.lmplot(x='total_bill', y='big_tip', data=tips, logistic=True, y_jitter=.03).set_axis_labels('Total Bill', 'Big Tip')
plt.title("Logistic Regression of Big Tip vs. Total Bill")
plt.show()
| 3_basic_bar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pyspark.sql import SparkSession # main entry point for DataFrame and SQL functionality
from pyspark.sql.functions import col # for returning a column based on a given column name
from pyspark.sql.functions import lit # for adding a new column to PySpark DataFrame
from pyspark.ml.classification import LogisticRegression # for classification model
from pyspark.ml.feature import OneHotEncoder, StringIndexer, VectorAssembler # for preparing data for classification
from pyspark.ml.evaluation import MulticlassClassificationEvaluator # for evaluating classification models
from pyspark.ml import Pipeline
import pandas as pd # for data frames
import numpy as np # for arrays
import time # for timing cells
import matplotlib.pyplot as plt # plotting graphs
# +
# Decision Tree
from pyspark.ml.classification import DecisionTreeClassifier
spark = SparkSession.builder.appName('Decision_Tree').getOrCreate()
spark
# +
# Use file created in the earlier HW to model Decision Tree.
ratings_df = spark.read.csv('ratings.csv', header=True, inferSchema=True)
ratings_df.count()
# -
ratings_columns = ratings_df.columns
pd.DataFrame(ratings_df.take(6000), columns=ratings_columns).groupby('ground_truth').count()
ratings_df.printSchema()
# +
feature_columns = ['album_score', 'artist_score', 'genre_score']
stages = []
assembler_inputs = feature_columns
assembler = VectorAssembler(inputCols=assembler_inputs, outputCol='features') # merges multiple columns into a vector column
stages += [assembler]
label_column = 'ground_truth'
label_string_idx = StringIndexer(inputCol=label_column, outputCol='label')
stages += [label_string_idx]
pipeline = Pipeline(stages=stages) # initialize the pipeline
pipeline_model = pipeline.fit(ratings_df) # fit the pipeline model
train_df = pipeline_model.transform(ratings_df) # transform the input DF with the pipeline model
# -
selected_columns = ['label', 'features'] + ratings_columns
train_df = train_df.select(selected_columns)
train_df.printSchema()
pd.DataFrame(train_df.take(5), columns=train_df.columns).transpose()
train_df, test_df = train_df.randomSplit([0.7, 0.3], seed=2021)
print(f'Training Dataset Count: {train_df.count()}')
print(f'Test Dataset Count: {test_df.count()}')
# +
# Load and prepare prediction file
prediction_df = spark.read.csv('genre_test.txt', sep='|', inferSchema=True) # inferSchema scans csv twice to get datatype of each column
prediction_df = prediction_df.withColumnRenamed("_c0", "userID").withColumnRenamed("_c1", "trackID").withColumnRenamed("_c2", "albumScore").withColumnRenamed("_c3", "artistScore").withColumnRenamed("_c4", "genreScore")
prediction_df.count()
# -
prediction_columns = prediction_df.columns
prediction_columns
prediction_df = prediction_df.withColumn('prediction', lit('0'))
pd.DataFrame(prediction_df.take(5), columns=prediction_df.columns).transpose()
prediction_df.printSchema()
feature_columns = ['albumScore', 'artistScore', 'genreScore']
stages = []
assembler_inputs = feature_columns
assembler = VectorAssembler(inputCols=assembler_inputs, outputCol='features') # merges multiple columns into a vector column
stages += [assembler]
# +
label_column = 'prediction'
label_string_idx = StringIndexer(inputCol=label_column, outputCol='label')
stages += [label_string_idx]
prediction_pipeline = Pipeline(stages=stages) # initialize the pipeline
prediction_pipeline_model = prediction_pipeline.fit(prediction_df) # fit the pipeline model
prediction_df = prediction_pipeline_model.transform(prediction_df) # transform the input DF with the pipeline model
selected_columns = ['label', 'features'] + prediction_columns
prediction_df = prediction_df.select(selected_columns)
prediction_df.printSchema()
# -
pd.DataFrame(prediction_df.take(5), columns=prediction_df.columns).transpose()
# +
# Decision Tree
start_time = time.time()
dt = DecisionTreeClassifier(featuresCol='features', labelCol='label', maxDepth=20)
dt_model = dt.fit(train_df)
end_time = time.time()
elapsed_time = end_time - start_time
print(f'Done! Time elapsed - {elapsed_time:.2f} seconds.')
# -
predictions_dt = dt_model.transform(test_df)
evaluator = MulticlassClassificationEvaluator(labelCol='label', predictionCol='prediction', metricName='accuracy') # initialize an Evaluator for Multiclass Classification
accuracy = evaluator.evaluate(predictions_dt) # evaluate decision tree model on predictions
print(f'Test Error = {1.0 - accuracy:.2%}')
sort_predictions_dt = predictions_dt.select('userID', 'trackID', 'label', 'probability', 'rawPrediction', 'prediction').sort(col('userID').asc(), col('probability').desc())
sort_predictions_dt.show(6)
dt_predictions = dt_model.transform(prediction_df) # transform prediction_df with decision tree model
dt_predictions.select('userID', 'trackID', 'probability', 'rawPrediction', 'prediction').show(12)
# col('userID').asc() sort the user ascending
# col('probability').desc() sort the probability descending (from large to small)
sort_dt_predictions = dt_predictions.select('userID', 'trackID', 'probability', 'rawPrediction', 'prediction').sort(col('userID').asc(), col('probability').desc())
sort_dt_predictions.show(6)
pd_sort_dt_predictions = sort_dt_predictions.toPandas().fillna(0.0) # create pandas df
pd_sort_dt_predictions
columns_to_write = ['userID', 'trackID']
pd_sort_dt_predictions.to_csv('decision_tree_predictions.csv', index=False, header=None, columns=columns_to_write) # write to csv (without header)
f_dt_predictions = open('decision_tree_predictions.csv')
f_dt_final_predictions = open('decision_tree_final_predictions.csv', 'w')
# Write header
f_dt_final_predictions.write('TrackID,Predictor\n')
# +
# Initialize some values
last_user_id = -1
track_id_out_vec = [0] * 6
start_time = time.time()
# Go through each line of the predictions file
for line in f_dt_predictions:
arr_out = line.strip().split(',') # remove any spaces/new lines and create list
user_id_out = arr_out[0] # set user
track_id_out = arr_out[1] # set track
if user_id_out != last_user_id: # if new user reached
i = 0 # reset i
track_id_out_vec[i] = track_id_out # add trackID to trackID array
i = i + 1 # increment i
last_user_id = user_id_out # set last_user_id as current userID
if i == 6: # if last entry for current user reached
# Here we set the predictions
predictions = np.ones(shape=(6)) # initialize numpy array for predictions
for index in range(0, 3):
predictions[index] = 0 # set first 3 values in array to 0 (other 3 are 1)
# Here we write to the final predictions file for the 6 track predictions for the current user
for ii in range(0, 6):
out_str = str(user_id_out) + '_' + str(track_id_out_vec[ii]) + ',' + str(int(predictions[ii]))
f_dt_final_predictions.write(out_str + '\n')
end_time = time.time()
elapsed_time = end_time - start_time
print(f'Done! Time elapsed - {elapsed_time:.2f} seconds.')
f_dt_predictions.close()
f_dt_final_predictions.close()
# +
# Random Forest
from pyspark.ml.classification import RandomForestClassifier
start_time = time.time()
rf = RandomForestClassifier(featuresCol='features', labelCol='label')
rf_model = rf.fit(train_df)
end_time = time.time()
elapsed_time = end_time - start_time
print(f'Done! Time elapsed - {elapsed_time:.2f} seconds.')
predictions_rf = rf_model.transform(test_df)
evaluator = MulticlassClassificationEvaluator(labelCol='label', predictionCol='prediction', metricName='accuracy') # initialize an Evaluator for Multiclass Classification
accuracy = evaluator.evaluate(predictions_rf) # evaluate random forest model on predictions
print(f'Test Error = {1.0 - accuracy:.2%}')
# -
sort_predictions_rf = predictions_rf.select('userID', 'trackID', 'label', 'probability', 'rawPrediction', 'prediction').sort(col('userID').asc(), col('probability').desc())
sort_predictions_rf.show(6)
rf_predictions = dt_model.transform(prediction_df) # transform prediction_df with random forest model
rf_predictions.select('userID', 'trackID', 'probability', 'rawPrediction', 'prediction').show(12)
# col('userID').asc() sort the user ascending
# col('probability').desc() sort the probability descending (from large to small)
sort_rf_predictions = rf_predictions.select('userID', 'trackID', 'probability', 'rawPrediction', 'prediction').sort(col('userID').asc(), col('probability').desc())
sort_rf_predictions.show(6)
pd_sort_rf_predictions = sort_rf_predictions.toPandas().fillna(0.0) # create pandas df
pd_sort_rf_predictions
columns_to_write = ['userID', 'trackID']
pd_sort_rf_predictions.to_csv('random_forest_predictions.csv', index=False, header=None, columns=columns_to_write) # write to csv (without headers)
f_rf_predictions = open('random_forest_predictions.csv')
f_rf_final_predictions = open('random_forest_final_predictions.csv', 'w')
# Write header
f_rf_final_predictions.write('TrackID,Predictor\n')
# +
# Initialize some values
last_user_id = -1
track_id_out_vec = [0] * 6
start_time = time.time()
# Go through each line of the predictions file
for line in f_rf_predictions:
arr_out = line.strip().split(',') # remove any spaces/new lines and create list
user_id_out = arr_out[0] # set user
track_id_out = arr_out[1] # set track
if user_id_out != last_user_id: # if new user reached
i = 0 # reset i
track_id_out_vec[i] = track_id_out # add trackID to trackID array
i = i + 1 # increment i
last_user_id = user_id_out # set last_user_id as current userID
if i == 6: # if last entry for current user reached
# Here we set the predictions
predictions = np.ones(shape=(6)) # initialize numpy array for predictions
for index in range(0, 3):
predictions[index] = 0 # set first 3 values in array to 0 (other 3 are 1)
# Here we write to the final predictions file for the 6 track predictions for the current user
for ii in range(0, 6):
out_str = str(user_id_out) + '_' + str(track_id_out_vec[ii]) + ',' + str(int(predictions[ii]))
f_rf_final_predictions.write(out_str + '\n')
end_time = time.time()
elapsed_time = end_time - start_time
print(f'Done! Time elapsed - {elapsed_time:.2f} seconds.')
f_rf_predictions.close()
f_rf_final_predictions.close()
# +
# Gradient Boosted Tree
from pyspark.ml.classification import GBTClassifier
start_time = time.time()
gbt = GBTClassifier(maxIter=100)
gbt_model = gbt.fit(train_df)
end_time = time.time()
elapsed_time = end_time - start_time
print(f'Done! Time elapsed - {elapsed_time:.2f} seconds.')
predictions_gbt = gbt_model.transform(test_df)
evaluator = MulticlassClassificationEvaluator(labelCol='label', predictionCol='prediction', metricName='accuracy') # initialize an Evaluator for Multiclass Classification
accuracy = evaluator.evaluate(predictions_gbt) # evaluate random forest model on predictions
print(f'Test Error = {1.0 - accuracy:.2%}')
# +
sort_predictions_gbt = predictions_gbt.select('userID', 'trackID', 'label', 'probability', 'rawPrediction', 'prediction').sort(col('userID').asc(), col('probability').desc())
sort_predictions_gbt.show(6)
gbt_predictions = gbt_model.transform(prediction_df) # transform prediction_df with gradient-boosted tree model
gbt_predictions.select('userID', 'trackID', 'probability', 'rawPrediction', 'prediction').show(12)
# -
# col('userID').asc() sort the user ascending
# col('probability').desc() sort the probability descending (from large to small)
sort_gbt_predictions = gbt_predictions.select('userID', 'trackID', 'probability', 'rawPrediction', 'prediction').sort(col('userID').asc(), col('probability').desc())
sort_gbt_predictions.show(6)
pd_sort_gbt_predictions = sort_gbt_predictions.toPandas().fillna(0.0) # create pandas df
pd_sort_gbt_predictions
columns_to_write = ['userID', 'trackID']
pd_sort_gbt_predictions.to_csv('grad_boost_predictions.csv', index=False, header=None, columns=columns_to_write) # write to csv (without headers)
f_gbt_predictions = open('grad_boost_predictions.csv')
f_gbt_final_predictions = open('grad_boost_final_predictions.csv', 'w')
# Write header
f_gbt_final_predictions.write('TrackID,Predictor\n')
# +
# Initialize some values
last_user_id = -1
track_id_out_vec = [0] * 6
start_time = time.time()
# Go through each line of the predictions file
for line in f_gbt_predictions:
arr_out = line.strip().split(',') # remove any spaces/new lines and create list
user_id_out = arr_out[0] # set user
track_id_out = arr_out[1] # set track
if user_id_out != last_user_id: # if new user reached
i = 0 # reset i
track_id_out_vec[i] = track_id_out # add trackID to trackID array
i = i + 1 # increment i
last_user_id = user_id_out # set last_user_id as current userID
if i == 6: # if last entry for current user reached
# Here we set the predictions
predictions = np.ones(shape=(6)) # initialize numpy array for predictions
for index in range(0, 3):
predictions[index] = 0 # set first 3 values in array to 0 (other 3 are 1)
# Here we write to the final predictions file for the 6 track predictions for the current user
for ii in range(0, 6):
out_str = str(user_id_out) + '_' + str(track_id_out_vec[ii]) + ',' + str(int(predictions[ii]))
f_gbt_final_predictions.write(out_str + '\n')
end_time = time.time()
elapsed_time = end_time - start_time
print(f'Done! Time elapsed - {elapsed_time:.2f} seconds.')
f_gbt_predictions.close()
f_gbt_final_predictions.close()
# -
| Music Recommendation System Project/Source Code/Grad_Boost_DT_RF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Quantum Gates
# Quantum gates are like the lies we tell our parents: they are models of reality that we use to solve a problem. These gates are unit operations used to alter the characteristics of what is being processed. Although in many cases we are not certain of the mechanisms taking place in the quantum realm, we are still able to measure and predict the outcomes, so these gates are placed to allow us manipulate the quantum phenomena and make the outcome useful.
# The Hadamah Gate uses concepts of mathematical matrixes to entangle quantum particles.
| 2020-11-12-secondPost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import libraries
import json
import pandas as pd
from pandas import DataFrame
from pandas.io.json import json_normalize #package for flattening json in pandas df
# +
file_path = './test_data/weather_obs-1.json'
#load json object
with open(file_path) as f:
d = json.load(f)
# Parse JSON file into a DataFrame
raw_data = json_normalize(d['SiteRep'])
raw_data
# +
# Parse the DV columns from JSON file
location = json_normalize(data=d['SiteRep']['DV'],
record_path=['Location'],
meta=['dataDate', 'type'])
# new = DataFrame(pd.json.loads(location.head(1).to_json()))
a = DataFrame(location.iloc[18])
# location.head(2)
# isinstance(location.iloc[19]['Period'], list)
len(location)
# +
a = DataFrame(location.iloc[18])
# location.head(2)
a.T
# -
b = json_normalize(data=a['Period'], record_path=['Rep'],
meta=['type', 'value'])
b
# +
# COPY META COLUMNS TO PERIOD RESULTS DATAFRAME
# a: 1-row DataFrame containing Period and meta columns
# b: results DataFrame
def copy_meta_to_period(a, b):
headers = list(a)
for i in range(1,len(headers)):
b[str(headers[i])] = a.iloc[0][i]
return b
def handle_lists(x, i):
a = DataFrame(x.iloc[i].to_dict())
b = json_normalize(data=a['Period'], record_path=['Rep'], meta=['type', 'value'])
return a, b
def handle_objects(x, i):
a = DataFrame(x.iloc[i])
b = json_normalize(data=a.T['Period'], record_path=['Rep'], meta=['type', 'value'])
return a, b
# Loop over all rows of location
result = DataFrame()
y = DataFrame()
z = DataFrame()
for i in range (0, len(location)):
if (isinstance(location.iloc[i]['Period'], list)): # Handle lists
[a, b] = handle_lists(location, i)
else: # Handle objects
[a, b] = handle_objects(location, i)
if (len(result) == 0):
result = copy_meta_to_period(a, b)
else:
result.append(copy_meta_to_period(a, b), ignore_index=True)
# print('#' + str(i) + ' ' + 'done')
result
# +
# period = json_normalize(data=location.head(1)['Period'], record_path=['Rep'])
# period
def normalize_period(df):
return json_normalize(data=df['Period'].to_dict(), record_path=[0, 'Rep'],
meta=[['0', 'type'], ['0', 'value']])
location.head(1).apply(normalize_period)
# +
period = json_normalize(data=d['SiteRep']['DV'], record_path=['Location','Period', 'Rep'],
meta=['dataDate',
['Location', 'continent'],
['Location', 'country'],
['Location', 'name'],
['Location', 'elevation'],
['Location', 'lat'],
['Location', 'lon'],
['Location', 'i'],
['Location', 'Period', 'type'],
['Location', 'Period', 'value'],
'type'])
period
| new_services/notebooks/.ipynb_checkpoints/weather_json2csv-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tensorflow.examples.tutorials.mnist import input_data
import tensorflow as tf
mnist = input_data.read_data_sets("MNIST_data", one_hot=True)
sess = tf.InteractiveSession()
# +
def weight_variable(shape):
initial = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(initial)
def bias_variable(shape):
initial = tf.constant(0.1, shape=shape)
return tf.Variable(initial)
# -
def conv2d(x, W):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME')
def max_pool_2x2(x):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1],
padding="SAME")
x = tf.placeholder(tf.float32, [None, 784])
y_ = tf.placeholder(tf.float32, [None, 10])
x_image = tf.reshape(x, [-1, 28, 28, 1])
W_conv1 = weight_variable([5, 5, 1, 32])
b_conv1 = bias_variable([32])
h_conv1 = tf.nn.relu(conv2d(x_image, W_conv1) + b_conv1)
h_pool1 = max_pool_2x2(h_conv1)
W_conv2 = weight_variable([5, 5, 32, 64])
b_conv2 = bias_variable([64])
h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2) + b_conv2)
h_pool2 = max_pool_2x2(h_conv2)
W_fc1 = weight_variable([7*7*64, 1024])
b_fc1 = bias_variable([1024])
h_pool2_flat = tf.reshape(h_pool2, [-1, 7*7*64])
h_fc1 = tf.nn.relu(tf.matmul(h_pool2_flat, W_fc1) + b_fc1)
keep_prob = tf.placeholder(tf.float32)
h_fc1_drop = tf.nn.dropout(h_fc1, keep_prob)
W_fc2 = weight_variable([1024, 10])
b_fc2 = bias_variable([10])
y_conv = tf.nn.softmax(tf.matmul(h_fc1_drop, W_fc2) + b_fc2)
cross_entropy = tf.reduce_mean(-tf.reduce_sum(y_*tf.log(y_conv), reduction_indices=[1]))
train_step = tf.train.AdamOptimizer(1e-4).minimize(cross_entropy)
correct_prediction = tf.equal(tf.argmax(y_conv, 1), tf.argmax(y_, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
tf.global_variables_initializer().run()
for i in range(2000):
batch = mnist.train.next_batch(100)
if i%100 ==0:
train_accuracy = accuracy.eval(feed_dict={x:batch[0], y_:batch[1],
keep_prob:1.0})
print("step %d, training accuracy %g"%(i, train_accuracy))
train_step.run({x:batch[0], y_:batch[1], keep_prob:0.75})
| src/CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## This note book contains code for pulling and aggregating the bi-daily scapes from 4chan into one dataframe which can be easily used for further analysis
#
# If you don't have access to s3, here's an excellent guide on obtaining access. Keys can be obtained by asking moderaters in the slack channel for far-right https://github.com/Data4Democracy/tutorials/blob/master/aws/AWS_Boto3_s3_intro.ipynb
#
# From this notebook anyone should be able to work on some more analysis, and the text of the messages from these scrapes looks very clean.
#
# If you only want to pull a certain set of dates, just adjust the regex in match_string, or add some more
# conditionals to the loop which grabs the list of files to be read.
#
# You can also read fewer files (if you have a slow connection or not that much memory) by shortening the files list before the second for loop.
import boto
import boto3
import pandas as pd
import re
from IPython.display import clear_output
session = boto3.Session(profile_name='default')
s3 = session.resource('s3')
bucket = s3.Bucket("far-right")
session.available_profiles
# +
base_url = 's3:far-right/'
match_string = "info-source/daily/[0-9]+/fourchan/fourchan"
files = []
print("Getting bucket and files info")
for obj in bucket.objects.all():
if bool(re.search(match_string, obj.key)):
files.append(obj.key)
df = pd.DataFrame()
for i, file in enumerate(files):
clear_output()
print("Loading file: " + str(i + 1) + " out of " + str(len(files)))
if df.empty:
df = pd.read_json(base_url + file)
else:
df = pd.concat([df, pd.read_json(base_url + file)])
clear_output()
print("Completed Loading Files")
# -
df.shape
df.head()
| exploratory_notebooks/Aggregating_bidaily_4chan_scrapes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mirsazzathossain/CSE317-Lab/blob/main/Lab_8_Building_Least_Square_Binary_Classifier.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="4lcxLlsWZB9u"
# #### **1. Import necessary packages:**
# + id="VLQvwLcFZBqn"
# Write appropriate code
# + [markdown] id="XuidPMi2ZFcr"
# #### **2. Upload and load dataset:**
# At first we have to upload the dataset to google colab to start working with it. Please download the **"Admission_classifier.csv"** dataset from piazza resourse or [click here](https://piazza.com/class_profile/get_resource/ku1fdd7zhev3r2/kwv4ddrap491yz) to download it. Then click on files form sidebar, drag and drop your file to side bar to upload the dataset.
#
# Now, use `data = pd.read_csv("Admission_classifier.csv")` to load the data.
# + id="h4uFtFXFYr6B"
# Write appropriate code
# + [markdown] id="blie1N5zbScV"
# #### **3. Preprocess the Data:**
# * To visualize the loaded data use `print(data.head())`.
# * Now, after visualizing the data did you observe we have an extra column named `Serial No.`?
# * This certainly is not a feature, so we will drop this column. Use `data.drop('Serial No.', axis=1, inplace=True)` to drop the column.
# * See the column `'Label'` is also not a feature rather it is our target. $y_i = 1$ means admitted. $y_i = -1$ means not admitted for $i^{th}$ student.
# * We will store it in a seperate variable `Y` using `Y = data['Label']`.
# * Convert `Y` to numpy array using `Y = Y.values`
# * Dorp the column from `data` using `data.drop('Label', axis=1, inplace=True)`
# * In `data` we are left with two features gre scores & toefl scores. Covert it to numpy array and store in a new variable `X` using `X = data.values`. So, `X` is the matrix of feature columns, each column in `X` will be the feature vectors. Assume the admission acceptance is decided based on these two features. So for every $i^{th}$ student has features $x_1$, and $x_2$
# + id="cIRwbvSFbTyO"
# Write appropriate code
# + [markdown] id="My89w3uAdd7X"
# #### **4. Add a ones column vector to X & partition the data and labels:**
# Add a new column cosisting ones as $0^{th}$ column to X. Saw the [numpy documentation](https://numpy.org/doc/stable/reference/generated/numpy.c_.html) for more details. Devide data X and Y into X_train, X_test, Y_train and Y_test. Train dataset will contains 300 datapoints and test dataset will contains 100 datapoint.
# + id="1ty_IXl8ebt-"
# Write appropriate code
# + [markdown] id="U28akWEDeoQp"
# #### **5. Solve the system of equation:**
# This time we design the classifier as follows. The classifier will give the appropriate admission label for an student. If the value $c_0 +c_1 x_1 +c_2x_2$ is positive, sign function outputs $1$. Otherwise it outputs $-1$.
# $$y_p = np.sign(c_0 +c_1x_1+c_2x_2)$$
# Now, calculate the coefficients $c = (c_0, c_1, c_2)$ by finding the least square solution for $(Xc = Y)$. Follow the exact procedure followed by earlier lab. So you will find $ĉ$ using the pseudo inverse of X. Use `X_train` and `X_train` as dataset.
# + id="7rL-VKk0huQ5"
# Write appropriate code
# + [markdown] id="WlqRl9FPiomB"
# #### **6. Find predicted labels:**
# Find the predicted labels `Y_test_pred`, by using the formula `Y_test_pred = np.sign(X_test * c_hat)`.
# + id="Y7Cw0JdejaB1"
# Write appropriate code
# + [markdown] id="DQ4HbwgtkVg8"
# #### **7. Calculate the accuracy:**
# Apparently `Y_test` are the ground truths for the test data. Now you have built a classifier that is defined by `c_hat`. Calculate the accuracy of your predicted label: `Y_test_pred` and groundtruth labels: `Y_test`.
# + id="8eST1-00kwxj"
# Write appropriate code
# + [markdown] id="904KzezAljx6"
# #### **8. Seperate the data into positive and negative:**
# Finally, separate `X_train` into `X_pos`, `X_neg` using the concept of boolean array indexing and `Y_train`. So that all `X_pos` will have labes `1`. And `X_neg` will have `-1` labels.
# + id="QGyNtxVImQnD"
# Write appropriate code
# + [markdown] id="hc2jqZi_n_Y2"
# #### **9. Plot 2D figure with X_pos & X_neg:**
# Plot 2D figure with `X_pos` and `X_neg` so that for each data point has feature $x_1$ will be on `x-axis` and $x_2$ will be on `y-axis`. Use marker ‘o’ with red color for `X_neg` and marker ‘*’ with blue color for `X_pos`.
#
# **Note: Previously we added one vector in $0^{th}$ column of `X`. Hence, feature $x_1=X[1]$ and feature $x_2=X[2]$
# + id="XZn-i-mapgoV"
# Write appropriate code
# + [markdown] id="XFSs_f7Yqyuc"
# #### **10. Plot the discriminating line:**
# Plot the discriminating line produced by the equation $c_0 + c_1 x_1 + c_2x_2 = 0$ by finding:
# $$x_2 = -( c_0 + c_1x_1) / c_2$$
# for $x_1 = 290:350$. Now, plot previous plots along with a new line plot using $x_1, x_2$ to see the boundary
# + id="FL-4ct5Mrn9z"
# Write appropriate code
| Lab_8_Building_Least_Square_Binary_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import stats
data = pd.read_csv("movies.csv")
data.head()
data[['binary','budget','domgross_2013$', 'intgross_2013$']].groupby('binary').mean()
data[['binary','budget','domgross_2013$', 'intgross_2013$']].groupby('binary').mean().plot.bar()
data.loc[data['binary'] == 'PASS'][['year','budget']].head()
data.loc[data['binary'] == 'PASS'][['year','budget']].groupby('year').mean().plot.line()
data.loc[data['binary'] == 'FAIL'][['year','budget']].groupby('year').mean().plot.line()
x_fail = sorted(data.year.unique())[1:]
y_fail = data.loc[data['binary'] == 'FAIL'][['year','budget']].groupby('year').mean()
x_pass = sorted(data.year.unique())
y_pass = data.loc[data['binary'] == 'PASS'][['year','budget']].groupby('year').mean()
f, ax = plt.subplots(1)
ax.plot(x_fail, y_fail)
del x_pass[1]
del x_pass[4]
#x_pass
#y_pass.loc[1971] = [0.0]
#y_pass.loc[1975] = [0.0]
#y_pass
ax.plot(x_pass, y_pass)
data[['year','binary','budget','domgross_2013$', 'intgross_2013$']].pivot_table(columns=[data.binary])
data.pivot_table(columns=data.binary)
data[['binary','budget','domgross_2013$', 'intgross_2013$']].groupby('binary').mean()
A = data.loc[data['binary'] == 'PASS']
B = data.loc[data['binary'] == 'FAIL']
A.columns = pd.MultiIndex.from_product([['A'], A.columns])
B.columns = pd.MultiIndex.from_product([['B'], B.columns])
pd.concat([A, B], axis = 1)
| trial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/WittmannF/rnn-tutorial-rnnlm/blob/master/RNNLM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="JgDGxaEJ1YmP" colab_type="text"
# # Recurrent Neural Networks Tutorial, Part 2 – Implementing a Language Model RNN with Python, Numpy and Theano
# + id="j5W3YjRI1_EE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c384a5e7-41ad-4ff9-bf29-f1fc72284501"
# !git clone https://github.com/WittmannF/rnn-tutorial-rnnlm.git
# + id="Sm0Rzg5w2BXX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="84e6e1d6-2c40-4426-c2ed-d24a5461bfb5"
# cd rnn-tutorial-rnnlm
# + id="QCfyOlkW1YmR" colab_type="code" colab={}
import csv
import itertools
import operator
import numpy as np
import nltk
import sys
from datetime import datetime
from utils import *
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# + id="w0JgkkU51YmW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="79144738-3aee-4092-f777-ebf9e6d29139"
# Download NLTK model data (you need to do this once)
nltk.download("book")
# + [markdown] id="6vsv4vJ11Ymb" colab_type="text"
# #### This the second part of the Recurrent Neural Network Tutorial. [The first part is here](http://www.wildml.com/2015/09/recurrent-neural-networks-tutorial-part-1-introduction-to-rnns/).
#
# In this part we will implement a full Recurrent Neural Network from scratch using Python and optimize our implementation using [Theano](http://deeplearning.net/software/theano/), a library to perform operations on a GPU. **[The full code is available on Github](https://github.com/dennybritz/rnn-tutorial-rnnlm/)**. I will skip over some boilerplate code that is not essential to understanding Recurrent Neural Networks, but all of that is also [on Github](https://github.com/dennybritz/rnn-tutorial-rnnlm).
# + [markdown] id="WSpL6N4D1Ymc" colab_type="text"
# ### Language Modeling
#
# Our goal is to build a [Language Model](https://en.wikipedia.org/wiki/Language_model) using a Recurrent Neural Network. Here's what that means. Let's say we have sentence of $m$ words. Language Model allows us to predict the probability of observing the sentence (in a given dataset) as:
#
# $
# \begin{aligned}
# P(w_1,...,w_m) = \prod_{i=1}^{m}P(w_i \mid w_1,..., w_{i-1})
# \end{aligned}
# $
#
# In words, the probability of a sentence is the product of probabilities of each word given the words that came before it. So, the probability of the sentence "He went to buy some chocolate" would be the probability of "chocolate" given "He went to buy some", multiplied by the probability of "some" given "He went to buy", and so on.
#
# Why is that useful? Why would we want to assign a probability to observing a sentence?
#
# First, such a model can be used as a scoring mechanism. For example, a Machine Translation system typically generates multiple candidates for an input sentence. You could use a language model to pick the most probable sentence. Intuitively, the most probable sentence is likely to be grammatically correct. Similar scoring happens in speech recognition systems.
#
# But solving the Language Modeling problem also has a cool side effect. Because we can predict the probability of a word given the preceding words, we are able to generate new text. It's a *generative model*. Given an existing sequence of words we sample a next word from the predicted probabilities, and repeat the process until we have a full sentence. <NAME> [has a great post](http://karpathy.github.io/2015/05/21/rnn-effectiveness/) that demonstrates what language models are capable of. His models are trained on single characters as opposed to full words, and can generate anything from Shakespeare to Linux Code.
#
# Note that in the above equation the probability of each word is conditioned on **all** previous words. In practice, many models have a hard time representing such long-term dependencies due to computational or memory constraints. They are typically limited to looking at only a few of the previous words. RNNs can, in theory, capture such long-term dependencies, but in practice it's a bit more complex. We'll explore that in a later post.
# + [markdown] id="WCTH6ixJ1Yme" colab_type="text"
# ### Training Data and Preprocessing
#
# To train our language model we need text to learn from. Fortunately we don't need any labels to train a language model, just raw text. I downloaded 15,000 longish reddit comments from a [dataset available on Google's BigQuery](https://bigquery.cloud.google.com/table/fh-bigquery:reddit_comments.2015_08). Text generated by our model will sound like reddit commenters (hopefully)! But as with most Machine Learning projects we first need to do some pre-processing to get our data into the right format.
#
# #### 1. Tokenize Text
#
# We have raw text, but we want to make predictions on a per-word basis. This means we must *tokenize* our comments into sentences, and sentences into words. We could just split each of the comments by spaces, but that wouldn't handle punctuation properly. The sentence "He left!" should be 3 tokens: "He", "left", "!". We'll use [NLTK's](http://www.nltk.org/) `word_tokenize` and `sent_tokenize` methods, which do most of the hard work for us.
#
# #### 2. Remove infrequent words
#
# Most words in our text will only appear one or two times. It's a good idea to remove these infrequent words. Having a huge vocabulary will make our model slow to train (we'll talk about why that is later), and because we don't have a lot of contextual examples for such words we wouldn't be able to learn how to use them correctly anyway. That's quite similar to how humans learn. To really understand how to appropriately use a word you need to have seen it in different contexts.
#
# In our code we limit our vocabulary to the `vocabulary_size` most common words (which I set to 8000, but feel free to change it). We replace all words not included in our vocabulary by `UNKNOWN_TOKEN`. For example, if we don't include the word "nonlinearities" in our vocabulary, the sentence "nonlineraties are important in neural networks" becomes "UNKNOWN_TOKEN are important in Neural Networks". The word `UNKNOWN_TOKEN` will become part of our vocabulary and we will predict it just like any other word. When we generate new text we can replace `UNKNOWN_TOKEN` again, for example by taking a randomly sampled word not in our vocabulary, or we could just generate sentences until we get one that doesn't contain an unknown token.
#
# #### 3. Prepend special start and end tokens
#
# We also want to learn which words tend start and end a sentence. To do this we prepend a special `SENTENCE_START` token, and append a special `SENTENCE_END` token to each sentence. This allows us to ask: Given that the first token is `SENTENCE_START`, what is the likely next word (the actual first word of the sentence)?
#
#
# #### 4. Build training data matrices
#
# The input to our Recurrent Neural Networks are vectors, not strings. So we create a mapping between words and indices, `index_to_word`, and `word_to_index`. For example, the word "friendly" may be at index 2001. A training example $x$ may look like `[0, 179, 341, 416]`, where 0 corresponds to `SENTENCE_START`. The corresponding label $y$ would be `[179, 341, 416, 1]`. Remember that our goal is to predict the next word, so y is just the x vector shifted by one position with the last element being the `SENTENCE_END` token. In other words, the correct prediction for word `179` above would be `341`, the actual next word.
# + id="cdYPayyP8Vw4" colab_type="code" colab={}
vocabulary_size = 8000
unknown_token = "UNKNOWN_TOKEN"
sentence_start_token = "SENTENCE_START"
sentence_end_token = "SENTENCE_END"
# + id="QWtrSKXH8WU0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="603f669b-d2f7-435b-a931-6081ec1dc506"
print("Reading CSV file...")
# + id="OnYkY-nz8Vt5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="f54c9871-f47f-42de-dc53-3e31224b9aa8"
df = pd.read_csv('data/reddit-comments-2015-08.csv'); df.head()
# + id="zz-v4zh6BYnI" colab_type="code" colab={}
#sentences = itertools.chain(*[nltk.sent_tokenize(x[0].decode('utf-8').lower()) for x in reader])
sentences = []
for x in df['body'].values:
try:
s = nltk.sent_tokenize(x.lower())
except:
s = nltk.sent_tokenize(x.decode('utf-8').lower())
sentences.extend(s)
# + id="NXakStX2Hqyu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 205} outputId="54444068-8e3e-480a-a04f-7e64dad7270d"
sentences[:10]
# + id="hTYylvayFKd2" colab_type="code" colab={}
sentences = ["%s %s %s" % (sentence_start_token, x, sentence_end_token) for x in sentences]
# + id="kliwJ1yXBsBu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 205} outputId="6a9deade-cb7b-4289-d1bd-65c7cb994af8"
sentences[:10]
# + id="Umdg29DcAeIH" colab_type="code" colab={}
# Tokenize the sentences into words
tokenized_sentences = [nltk.word_tokenize(sent) for sent in sentences]
# + id="KoF2zlZiFR8z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="306adb8b-bd9b-459b-e891-b16049e39416"
# Count the word frequencies
word_freq = nltk.FreqDist(itertools.chain(*tokenized_sentences))
print("Found %d unique words tokens." % len(list(word_freq.items())))
# + id="q-uOPu-_FuKp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cf7f21fb-36a1-480c-cab0-e33c682ec943"
word_freq['cat']
# + id="2V8WQpX2F2Et" colab_type="code" colab={}
# Get the most common words and build index_to_word and word_to_index vectors
vocab = word_freq.most_common(vocabulary_size-1)
index_to_word = [x[0] for x in vocab]
index_to_word.append(unknown_token)
word_to_index = dict([(w,i) for i,w in enumerate(index_to_word)])
# + id="MYdjo5jVF2M5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="de316767-3e63-4ab8-8bdf-e5163d4785b1"
print("Using vocabulary size %d." % vocabulary_size)
print("The least frequent word in our vocabulary is '%s' and appeared %d times." % (vocab[-1][0], vocab[-1][1]))
# + id="01z8XEcAGGBh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="f5422bf9-86dc-4027-8cd2-04abfcd5d0bc"
# Replace all words not in our vocabulary with the unknown token
for i, sent in enumerate(tokenized_sentences):
tokenized_sentences[i] = [w if w in word_to_index else unknown_token for w in sent]
print("\nExample sentence: '%s'" % sentences[0])
print("\nExample sentence after Pre-processing: '%s'" % tokenized_sentences[0])
# + id="JJhjtGWU1Ymm" colab_type="code" colab={}
# Create the training data
X_train = np.asarray([[word_to_index[w] for w in sent[:-1]] for sent in tokenized_sentences])
y_train = np.asarray([[word_to_index[w] for w in sent[1:]] for sent in tokenized_sentences])
# + [markdown] id="LKTKWIkX1Ymr" colab_type="text"
# Here's an actual training example from our text:
# + id="bQRyRvUe1Ymr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="08a4114f-fdb1-409a-fd62-9e0aad834f5e"
# Print an training data example
x_example, y_example = X_train[0], y_train[0]
print("x:\n%s\n%s" % (" ".join([index_to_word[x] for x in x_example]), x_example))
print("\ny:\n%s\n%s" % (" ".join([index_to_word[x] for x in y_example]), y_example))
# + [markdown] id="yG1fpj5u1Ymy" colab_type="text"
# #### Building the RNN
#
# For a general overview of RNNs take a look at [first part of the tutorial](http://www.wildml.com/2015/09/recurrent-neural-networks-tutorial-part-1-introduction-to-rnns/).
#
# 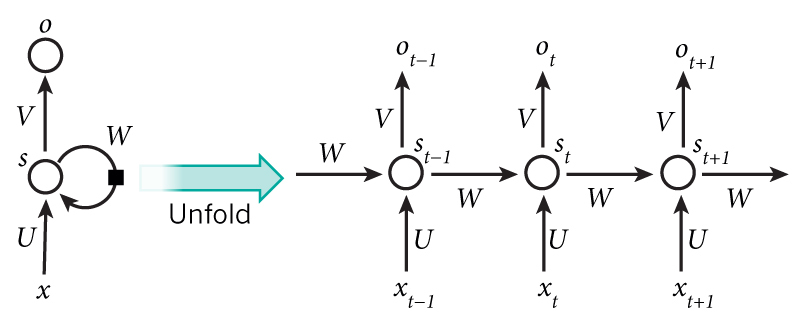
#
# Let's get concrete and see what the RNN for our language model looks like. The input $x$ will be a sequence of words (just like the example printed above) and each $x_t$ is a single word. But there's one more thing: Because of how matrix multiplication works we can't simply use a word index (like 36) as an input. Instead, we represent each word as a *one-hot vector* of size `vocabulary_size`. For example, the word with index 36 would be the vector of all 0's and a 1 at position 36. So, each $x_t$ will become a vector, and $x$ will be a matrix, with each row representing a word. We'll perform this transformation in our Neural Network code instead of doing it in the pre-processing. The output of our network $o$ has a similar format. Each $o_t$ is a vector of `vocabulary_size` elements, and each element represents the probability of that word being the next word in the sentence.
#
# Let's recap the equations for the RNN from the first part of the tutorial:
#
# $
# \begin{aligned}
# s_t &= \tanh(Ux_t + Ws_{t-1}) \\
# o_t &= \mathrm{softmax}(Vs_t)
# \end{aligned}
# $
#
# I always find it useful to write down the dimensions of the matrices and vectors. Let's assume we pick a vocabulary size $C = 8000$ and a hidden layer size $H = 100$. You can think of the hidden layer size as the "memory" of our network. Making it bigger allows us to learn more complex patterns, but also results in additional computation. Then we have:
#
# $
# \begin{aligned}
# x_t & \in \mathbb{R}^{8000} \\
# o_t & \in \mathbb{R}^{8000} \\
# s_t & \in \mathbb{R}^{100} \\
# U & \in \mathbb{R}^{100 \times 8000} \\
# V & \in \mathbb{R}^{8000 \times 100} \\
# W & \in \mathbb{R}^{100 \times 100} \\
# \end{aligned}
# $
#
# This is valuable information. Remember that $U,V$ and $W$ are the parameters of our network we want to learn from data. Thus, we need to learn a total of $2HC + H^2$ parameters. In the case of $C=8000$ and $H=100$ that's 1,610,000. The dimensions also tell us the bottleneck of our model. Note that because $x_t$ is a one-hot vector, multiplying it with $U$ is essentially the same as selecting a column of U, so we don't need to perform the full multiplication. Then, the biggest matrix multiplication in our network is $Vs_t$. That's why we want to keep our vocabulary size small if possible.
#
# Armed with this, it's time to start our implementation.
# + [markdown] id="QnjXzaME1Ymz" colab_type="text"
# #### Initialization
#
# We start by declaring a RNN class an initializing our parameters. I'm calling this class `RNNNumpy` because we will implement a Theano version later. Initializing the parameters $U,V$ and $W$ is a bit tricky. We can't just initialize them to 0's because that would result in symmetric calculations in all our layers. We must initialize them randomly. Because proper initialization seems to have an impact on training results there has been lot of research in this area. It turns out that the best initialization depends on the activation function ($\tanh$ in our case) and one [recommended](http://jmlr.org/proceedings/papers/v9/glorot10a/glorot10a.pdf) approach is to initialize the weights randomly in the interval from $\left[-\frac{1}{\sqrt{n}}, \frac{1}{\sqrt{n}}\right]$ where $n$ is the number of incoming connections from the previous layer. This may sound overly complicated, but don't worry too much about it. As long as you initialize your parameters to small random values it typically works out fine.
# + id="ZIfWWGcD1Ym0" colab_type="code" colab={}
class RNNNumpy:
def __init__(self, word_dim, hidden_dim=100, bptt_truncate=4):
# Assign instance variables
self.word_dim = word_dim
self.hidden_dim = hidden_dim
self.bptt_truncate = bptt_truncate
# Randomly initialize the network parameters
self.U = np.random.uniform(-np.sqrt(1./word_dim), np.sqrt(1./word_dim), (hidden_dim, word_dim))
self.V = np.random.uniform(-np.sqrt(1./hidden_dim), np.sqrt(1./hidden_dim), (word_dim, hidden_dim))
self.W = np.random.uniform(-np.sqrt(1./hidden_dim), np.sqrt(1./hidden_dim), (hidden_dim, hidden_dim))
# + [markdown] id="KY6zy5Zo1Ym5" colab_type="text"
# Above, `word_dim` is the size of our vocabulary, and `hidden_dim` is the size of our hidden layer (we can pick it). Don't worry about the `bptt_truncate` parameter for now, we'll explain what that is later.
# + [markdown] id="hv6vd-ec1Ym6" colab_type="text"
# #### Forward Propagation
#
# Next, let's implement the forward propagation (predicting word probabilities) defined by our equations above:
# + id="QrwSoWnR1Ym7" colab_type="code" colab={}
def forward_propagation(self, x):
# The total number of time steps
T = len(x)
# During forward propagation we save all hidden states in s because need them later.
# We add one additional element for the initial hidden, which we set to 0
s = np.zeros((T + 1, self.hidden_dim))
s[-1] = np.zeros(self.hidden_dim)
# The outputs at each time step. Again, we save them for later.
o = np.zeros((T, self.word_dim))
# For each time step...
for t in np.arange(T):
# Note that we are indxing U by x[t]. This is the same as multiplying U with a one-hot vector.
s[t] = np.tanh(self.U[:,x[t]] + self.W.dot(s[t-1]))
o[t] = softmax(self.V.dot(s[t]))
return [o, s]
RNNNumpy.forward_propagation = forward_propagation
# + [markdown] id="lbjIYmE-1Ym_" colab_type="text"
# We not only return the calculated outputs, but also the hidden states. We will use them later to calculate the gradients, and by returning them here we avoid duplicate computation. Each $o_t$ is a vector of probabilities representing the words in our vocabulary, but sometimes, for example when evaluating our model, all we want is the next word with the highest probability. We call this function `predict`:
# + id="0NM4uK0B1YnA" colab_type="code" colab={}
def predict(self, x):
# Perform forward propagation and return index of the highest score
o, s = self.forward_propagation(x)
return np.argmax(o, axis=1)
RNNNumpy.predict = predict
# + [markdown] id="D6e7qKcZ1YnK" colab_type="text"
# Let's try our newly implemented methods and see an example output:
# + id="6Wn76vKq1YnL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 151} outputId="170e5402-75c8-4c32-8318-690a47946014"
np.random.seed(10)
model = RNNNumpy(vocabulary_size)
o, s = model.forward_propagation(X_train[10])
print(o.shape)
print(o)
# + [markdown] id="XMji2ZjD1YnV" colab_type="text"
# For each word in the sentence (45 above), our model made 8000 predictions representing probabilities of the next word. Note that because we initialized $U,V,W$ to random values these predictions are completely random right now. The following gives the indices of the highest probability predictions for each word:
# + id="fxJXrNGT1YnW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="5fab3fee-c8ec-4035-c3c3-c301ed02604b"
predictions = model.predict(X_train[10])
print(predictions.shape)
print(predictions)
# + [markdown] id="yQz4EFq91Ynd" colab_type="text"
# #### Calculating the Loss
#
# To train our network we need a way to measure the errors it makes. We call this the loss function $L$, and our goal is find the parameters $U,V$ and $W$ that minimize the loss function for our training data. A common choice for the loss function is the [cross-entropy loss](https://en.wikipedia.org/wiki/Cross_entropy#Cross-entropy_error_function_and_logistic_regression). If we have $N$ training examples (words in our text) and $C$ classes (the size of our vocabulary) then the loss with respect to our predictions $o$ and the true labels $y$ is given by:
#
# $
# \begin{aligned}
# L(y,o) = - \frac{1}{N} \sum_{n \in N} y_{n} \log o_{n}
# \end{aligned}
# $
#
# The formula looks a bit complicated, but all it really does is sum over our training examples and add to the loss based on how off our prediction are. The further away $y$ (the correct words) and $o$ (our predictions), the greater the loss will be. We implement the function `calculate_loss`:
# + id="oK_BUDNg1Yne" colab_type="code" colab={}
def calculate_total_loss(self, x, y):
L = 0
# For each sentence...
for i in np.arange(len(y)):
o, s = self.forward_propagation(x[i])
# We only care about our prediction of the "correct" words
correct_word_predictions = o[np.arange(len(y[i])), y[i]]
# Add to the loss based on how off we were
L += -1 * np.sum(np.log(correct_word_predictions))
return L
def calculate_loss(self, x, y):
# Divide the total loss by the number of training examples
N = np.sum((len(y_i) for y_i in y))
return self.calculate_total_loss(x,y)/N
RNNNumpy.calculate_total_loss = calculate_total_loss
RNNNumpy.calculate_loss = calculate_loss
# + [markdown] id="xgsPci971Ynm" colab_type="text"
# Let's take a step back and think about what the loss should be for random predictions. That will give us a baseline and make sure our implementation is correct. We have $C$ words in our vocabulary, so each word should be (on average) predicted with probability $1/C$, which would yield a loss of $L = -\frac{1}{N} N \log\frac{1}{C} = \log C$:
# + id="foTd4kN81Ynn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="6ece0b2b-2904-4be1-fe76-65c21d8f06c0"
# Limit to 1000 examples to save time
print("Expected Loss for random predictions: %f" % np.log(vocabulary_size))
print("Actual loss: %f" % model.calculate_loss(X_train[:1000], y_train[:1000]))
# + [markdown] id="EcGbAX0V1Ynq" colab_type="text"
# Pretty close! Keep in mind that evaluating the loss on the full dataset is an expensive operation and can take hours if you have a lot of data!
# + [markdown] id="qH7ldtZ41Ynr" colab_type="text"
# #### Training the RNN with SGD and Backpropagation Through Time (BPTT)
#
# Remember that we want to find the parameters $U,V$ and $W$ that minimize the total loss on the training data. The most common way to do this is SGD, Stochastic Gradient Descent. The idea behind SGD is pretty simple. We iterate over all our training examples and during each iteration we nudge the parameters into a direction that reduces the error. These directions are given by the gradients on the loss: $\frac{\partial L}{\partial U}, \frac{\partial L}{\partial V}, \frac{\partial L}{\partial W}$. SGD also needs a *learning rate*, which defines how big of a step we want to make in each iteration. SGD is the most popular optimization method not only for Neural Networks, but also for many other Machine Learning algorithms. As such there has been a lot of research on how to optimize SGD using batching, parallelism and adaptive learning rates. Even though the basic idea is simple, implementing SGD in a really efficient way can become very complex. If you want to learn more about SGD [this](http://cs231n.github.io/optimization-1/) is a good place to start. Due to its popularity there are a wealth of tutorials floating around the web, and I don't want to duplicate them here. I'll implement a simple version of SGD that should be understandable even without a background in optimization.
#
# But how do we calculate those gradients we mentioned above? In a [traditional Neural Network](http://www.wildml.com/2015/09/implementing-a-neural-network-from-scratch/) we do this through the backpropagation algorithm. In RNNs we use a slightly modified version of the this algorithm called Backpropagation Through Time (BPTT). Because the parameters are shared by all time steps in the network, the gradient at each output depends not only on the calculations of the current time step, but also the previous time steps. If you know calculus, it really is just applying the chain rule. The next part of the tutorial will be all about BPTT, so I won't go into detailed derivation here. For a general introduction to backpropagation check out [this](http://colah.github.io/posts/2015-08-Backprop/) and this [post](http://cs231n.github.io/optimization-2/). For now you can treat BPTT as a black box. It takes as input a training example $(x,y)$ and returns the gradients $\frac{\partial L}{\partial U}, \frac{\partial L}{\partial V}, \frac{\partial L}{\partial W}$.
# + id="i_X0HS_E1Ynr" colab_type="code" colab={}
def bptt(self, x, y):
T = len(y)
# Perform forward propagation
o, s = self.forward_propagation(x)
# We accumulate the gradients in these variables
dLdU = np.zeros(self.U.shape)
dLdV = np.zeros(self.V.shape)
dLdW = np.zeros(self.W.shape)
delta_o = o
delta_o[np.arange(len(y)), y] -= 1.
# For each output backwards...
for t in np.arange(T)[::-1]:
dLdV += np.outer(delta_o[t], s[t].T)
# Initial delta calculation
delta_t = self.V.T.dot(delta_o[t]) * (1 - (s[t] ** 2))
# Backpropagation through time (for at most self.bptt_truncate steps)
for bptt_step in np.arange(max(0, t-self.bptt_truncate), t+1)[::-1]:
# print "Backpropagation step t=%d bptt step=%d " % (t, bptt_step)
dLdW += np.outer(delta_t, s[bptt_step-1])
dLdU[:,x[bptt_step]] += delta_t
# Update delta for next step
delta_t = self.W.T.dot(delta_t) * (1 - s[bptt_step-1] ** 2)
return [dLdU, dLdV, dLdW]
RNNNumpy.bptt = bptt
# + [markdown] id="RSF6NViw1Ynz" colab_type="text"
# #### Gradient Checking
#
# Whenever you implement backpropagation it is good idea to also implement *gradient checking*, which is a way of verifying that your implementation is correct. The idea behind gradient checking is that derivative of a parameter is equal to the slope at the point, which we can approximate by slightly changing the parameter and then dividing by the change:
#
# $
# \begin{aligned}
# \frac{\partial L}{\partial \theta} \approx \lim_{h \to 0} \frac{J(\theta + h) - J(\theta -h)}{2h}
# \end{aligned}
# $
#
# We then compare the gradient we calculated using backpropagation to the gradient we estimated with the method above. If there's no large difference we are good. The approximation needs to calculate the total loss for *every* parameter, so that gradient checking is very expensive (remember, we had more than a million parameters in the example above). So it's a good idea to perform it on a model with a smaller vocabulary.
# + id="HP09rY4U1Yn0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 154} outputId="3d28b0b7-1a08-408e-c86e-7cb34778ad08"
def gradient_check(self, x, y, h=0.001, error_threshold=0.01):
# Calculate the gradients using backpropagation. We want to checker if these are correct.
bptt_gradients = model.bptt(x, y)
# List of all parameters we want to check.
model_parameters = ['U', 'V', 'W']
# Gradient check for each parameter
for pidx, pname in enumerate(model_parameters):
# Get the actual parameter value from the mode, e.g. model.W
parameter = operator.attrgetter(pname)(self)
print("Performing gradient check for parameter %s with size %d." % (pname, np.prod(parameter.shape)))
# Iterate over each element of the parameter matrix, e.g. (0,0), (0,1), ...
it = np.nditer(parameter, flags=['multi_index'], op_flags=['readwrite'])
while not it.finished:
ix = it.multi_index
# Save the original value so we can reset it later
original_value = parameter[ix]
# Estimate the gradient using (f(x+h) - f(x-h))/(2*h)
parameter[ix] = original_value + h
gradplus = model.calculate_total_loss([x],[y])
parameter[ix] = original_value - h
gradminus = model.calculate_total_loss([x],[y])
estimated_gradient = (gradplus - gradminus)/(2*h)
# Reset parameter to original value
parameter[ix] = original_value
# The gradient for this parameter calculated using backpropagation
backprop_gradient = bptt_gradients[pidx][ix]
# calculate The relative error: (|x - y|/(|x| + |y|))
relative_error = np.abs(backprop_gradient - estimated_gradient)/(np.abs(backprop_gradient) + np.abs(estimated_gradient))
# If the error is to large fail the gradient check
if relative_error > error_threshold:
print("Gradient Check ERROR: parameter=%s ix=%s" % (pname, ix))
print("+h Loss: %f" % gradplus)
print("-h Loss: %f" % gradminus)
print("Estimated_gradient: %f" % estimated_gradient)
print("Backpropagation gradient: %f" % backprop_gradient)
print("Relative Error: %f" % relative_error)
return
it.iternext()
print("Gradient check for parameter %s passed." % (pname))
RNNNumpy.gradient_check = gradient_check
# To avoid performing millions of expensive calculations we use a smaller vocabulary size for checking.
grad_check_vocab_size = 100
np.random.seed(10)
model = RNNNumpy(grad_check_vocab_size, 10, bptt_truncate=1000)
model.gradient_check([0,1,2,3], [1,2,3,4])
# + [markdown] id="1lPGRAIz1Yn7" colab_type="text"
# #### SGD Implementation
#
# Now that we are able to calculate the gradients for our parameters we can implement SGD. I like to do this in two steps: 1. A function `sdg_step` that calculates the gradients and performs the updates for one batch. 2. An outer loop that iterates through the training set and adjusts the learning rate.
# + id="Fi_IN8mG1Yn8" colab_type="code" colab={}
# Performs one step of SGD.
def numpy_sdg_step(self, x, y, learning_rate):
# Calculate the gradients
dLdU, dLdV, dLdW = self.bptt(x, y)
# Change parameters according to gradients and learning rate
self.U -= learning_rate * dLdU
self.V -= learning_rate * dLdV
self.W -= learning_rate * dLdW
RNNNumpy.sgd_step = numpy_sdg_step
# + id="LHOaeeZe1YoD" colab_type="code" colab={}
# Outer SGD Loop
# - model: The RNN model instance
# - X_train: The training data set
# - y_train: The training data labels
# - learning_rate: Initial learning rate for SGD
# - nepoch: Number of times to iterate through the complete dataset
# - evaluate_loss_after: Evaluate the loss after this many epochs
def train_with_sgd(model, X_train, y_train, learning_rate=0.005, nepoch=100, evaluate_loss_after=5):
# We keep track of the losses so we can plot them later
losses = []
num_examples_seen = 0
for epoch in range(nepoch):
# Optionally evaluate the loss
if (epoch % evaluate_loss_after == 0):
loss = model.calculate_loss(X_train, y_train)
losses.append((num_examples_seen, loss))
time = datetime.now().strftime('%Y-%m-%d %H:%M:%S')
print("%s: Loss after num_examples_seen=%d epoch=%d: %f" % (time, num_examples_seen, epoch, loss))
# Adjust the learning rate if loss increases
if (len(losses) > 1 and losses[-1][1] > losses[-2][1]):
learning_rate = learning_rate * 0.5
print("Setting learning rate to %f" % learning_rate)
sys.stdout.flush()
# For each training example...
for i in range(len(y_train)):
# One SGD step
model.sgd_step(X_train[i], y_train[i], learning_rate)
num_examples_seen += 1
# + [markdown] id="LsY7dZTX1YoK" colab_type="text"
# Done! Let's try to get a sense of how long it would take to train our network:
# + id="BBLscBgt1YoL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cd9a5a6f-d365-4ec6-a19c-6635a86a37c9"
np.random.seed(10)
model = RNNNumpy(vocabulary_size)
# %timeit model.sgd_step(X_train[10], y_train[10], 0.005)
# + [markdown] id="XX8aDcVX1YoR" colab_type="text"
# Uh-oh, bad news. One step of SGD takes approximately 350 milliseconds on my laptop. We have about 80,000 examples in our training data, so one epoch (iteration over the whole data set) would take several hours. Multiple epochs would take days, or even weeks! And we're still working with a small dataset compared to what's being used by many of the companies and researchers out there. What now?
#
# Fortunately there are many ways to speed up our code. We could stick with the same model and make our code run faster, or we could modify our model to be less computationally expensive, or both. Researchers have identified many ways to make models less computationally expensive, for example by using a hierarchical softmax or adding projection layers to avoid the large matrix multiplications (see also [here](http://arxiv.org/pdf/1301.3781.pdf) or [here](http://www.fit.vutbr.cz/research/groups/speech/publi/2011/mikolov_icassp2011_5528.pdf)). But I want to keep our model simple and go the first route: Make our implementation run faster using a GPU. Before doing that though, let's just try to run SGD with a small dataset and check if the loss actually decreases:
# + id="ZX5O1EhA1YoS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="5974eaae-b519-44ec-bdf0-1fb9623936b3"
np.random.seed(10)
# Train on a small subset of the data to see what happens
model = RNNNumpy(vocabulary_size)
losses = train_with_sgd(model, X_train[:100], y_train[:100], nepoch=10, evaluate_loss_after=1)
# + id="-6eQJ7kb6vIN" colab_type="code" colab={}
# Now that we have our model we can ask it to generate new text for us!
# Let's implement a helper function to generate new sentences:
def generate_sentence(model):
# We start the sentence with the start token
new_sentence = [word_to_index[sentence_start_token]]
# Repeat until we get an end token
while not new_sentence[-1] == word_to_index[sentence_end_token]:
next_word_probs = model.forward_propagation(new_sentence)
sampled_word = word_to_index[unknown_token]
# We don't want to sample unknown words
while sampled_word == word_to_index[unknown_token]:
samples = np.random.multinomial(1, next_word_probs[-1])
sampled_word = np.argmax(samples)
new_sentence.append(sampled_word)
sentence_str = [index_to_word[x] for x in new_sentence[1:-1]]
return sentence_str
# + id="ygtTmb4Z7Mob" colab_type="code" colab={}
num_sentences = 10
senten_min_length = 7
i = 0 #for i in range(num_sentences):
# + id="ek110Llh7NYk" colab_type="code" colab={}
sent = []
# + id="zOefDNJU7RlY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="82647aa0-5d2a-4ac3-d064-c887f5c96f75"
print(f'While len(sent): {len(sent)} < senten_min_length: {senten_min_length} ')
# + id="nvkGVfh17RrB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="42dcd98e-33d6-4467-f64a-1cb617c8ded7"
# generate_sentence(model)
model
# + id="TtDvU_7xLbad" colab_type="code" colab={}
# We start the sentence with the start token
new_sentence = [word_to_index[sentence_start_token]]
# + id="jkz5orOKLi5n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6480faac-4899-4146-9163-823db65e8c58"
# Repeat until we get an end token
print(f"while not new_sentence[-1]: {new_sentence[-1]} == word_to_index[sentence_end_token] {word_to_index[sentence_end_token]}")
# + id="O_l4UydkLi9L" colab_type="code" colab={}
next_word_probs = model.forward_propagation(new_sentence)
# + id="EkGsXQT_M_Oh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9a44c186-abf2-4691-a9f5-99053d9c46ea"
len(next_word_probs)
# + id="kkwvnCmOMMrJ" colab_type="code" colab={}
sampled_word = word_to_index[unknown_token]
# + id="LuYZwLpUMOqk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="57215103-25a5-4eb1-b4be-e9b89be3321f"
sampled_word
# + id="o4BY7aBlMYdV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e734a967-df52-43bf-bcdb-7bceae9a14e4"
# We don't want to sample unknown words
print(f"while sampled_word: {sampled_word} == word_to_index[unknown_token] {word_to_index[unknown_token]}")
# + id="YCUiyokPMFyz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9b64ea55-7911-4cb2-982b-d7caef41d521"
#samples = np.random.multinomial(1, next_word_probs[-1])
len(next_word_probs[-1])
# + id="QiWiFjdVMF2z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 162} outputId="c4095cca-f777-4b66-a098-9220145459df"
sampled_word = np.argmax(samples)
# + id="TPOmO9wyMFwI" colab_type="code" colab={}
new_sentence.append(sampled_word)
# + id="JAnG8-koLi3S" colab_type="code" colab={}
sentence_str = [index_to_word[x] for x in new_sentence[1:-1]]
# + id="twmVBT8lLbYL" colab_type="code" colab={}
# + id="cDw8ZBVL6w5B" colab_type="code" colab={}
num_sentences = 10
senten_min_length = 7
for i in range(num_sentences):
sent = []
# We want long sentences, not sentences with one or two words
while len(sent) < senten_min_length:
sent = generate_sentence(model)
print(" ".join(sent))
# + id="5qKjPELj63uu" colab_type="code" colab={}
| RNNLM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic core
# This module contains all the basic functions we need in other modules of the fastai library (split with [`torch_core`](/torch_core.html#torch_core) that contains the ones requiring pytorch). Its documentation can easily be skipped at a first read, unless you want to know what a given function does.
# + hide_input=true
from fastai.gen_doc.nbdoc import *
from fastai.core import *
# -
# ## Global constants
# `default_cpus = min(16, num_cpus())` <div style="text-align: right"><a href="https://github.com/fastai/fastai/blob/master/fastai/core.py#L45">[source]</a></div>
# ## Check functions
# + hide_input=true
show_doc(has_arg)
# -
# Examples for two [`fastai.core`](/core.html#core) functions. Docstring shown before calling [`has_arg`](/core.html#has_arg) for reference
#
has_arg(download_url,'url')
has_arg(index_row,'x')
has_arg(index_row,'a')
# + hide_input=true
show_doc(ifnone)
# -
param,alt_param = None,5
ifnone(param,alt_param)
param,alt_param = None,[1,2,3]
ifnone(param,alt_param)
# + hide_input=true
show_doc(is1d)
# -
two_d_array = np.arange(12).reshape(6,2)
print( two_d_array )
print( is1d(two_d_array) )
is1d(two_d_array.flatten())
# + hide_input=true
show_doc(is_listy)
# -
# Check if `x` is a `Collection`. `Tuple` or `List` qualify
some_data = [1,2,3]
is_listy(some_data)
some_data = (1,2,3)
is_listy(some_data)
some_data = 1024
print( is_listy(some_data) )
print( is_listy( [some_data] ) )
some_data = dict([('a',1),('b',2),('c',3)])
print( some_data )
print( some_data.keys() )
print( is_listy(some_data) )
print( is_listy(some_data.keys()) )
print( is_listy(list(some_data.keys())) )
# + hide_input=true
show_doc(is_tuple)
# -
# Check if `x` is a `tuple`.
print( is_tuple( [1,2,3] ) )
print( is_tuple( (1,2,3) ) )
# ## Collection related functions
# + hide_input=true
show_doc(arange_of)
# -
arange_of([5,6,7])
type(arange_of([5,6,7]))
# + hide_input=true
show_doc(array)
# -
array([1,2,3])
# Note that after we call the generator, we do not reset. So the [`array`](/core.html#array) call has 5 less entries than it would if we ran from the start of the generator.
# +
def data_gen():
i = 100.01
while i<200:
yield i
i += 1.
ex_data_gen = data_gen()
for _ in range(5):
print(next(ex_data_gen))
# -
array(ex_data_gen)
# +
ex_data_gen_int = data_gen()
array(ex_data_gen_int,dtype=int) #Cast output to int array
# + hide_input=true
show_doc(arrays_split)
# +
data_a = np.arange(15)
data_b = np.arange(15)[::-1]
mask_a = (data_a > 10)
print(data_a)
print(data_b)
print(mask_a)
# -
arrays_split(mask_a,data_a)
np.vstack([data_a,data_b]).transpose().shape
arrays_split(mask_a,np.vstack([data_a,data_b]).transpose()) #must match on dimension 0
# + hide_input=true
show_doc(chunks)
# -
# You can transform a `Collection` into an `Iterable` of 'n' sized chunks by calling [`chunks`](/core.html#chunks):
data = [0,1,2,3,4,5,6,7,8,9]
for chunk in chunks(data, 2):
print(chunk)
for chunk in chunks(data, 3):
print(chunk)
# + hide_input=true
show_doc(df_names_to_idx)
# -
ex_df = pd.DataFrame.from_dict({"a":[1,1,1],"b":[2,2,2]})
print(ex_df)
df_names_to_idx('b',ex_df)
# + hide_input=true
show_doc(extract_kwargs)
# -
key_word_args = {"a":2,"some_list":[1,2,3],"param":'mean'}
key_word_args
(extracted_val,remainder) = extract_kwargs(['param'],key_word_args)
print( extracted_val,remainder )
# + hide_input=true
show_doc(idx_dict)
# -
idx_dict(['a','b','c'])
# + hide_input=true
show_doc(index_row)
# -
data = [0,1,2,3,4,5,6,7,8,9]
index_row(data,4)
index_row(pd.Series(data),7)
data_df = pd.DataFrame([data[::-1],data]).transpose()
data_df
index_row(data_df,7)
# + hide_input=true
show_doc(listify)
# -
to_match = np.arange(12)
listify('a',to_match)
listify('a',5)
listify(77.1,3)
listify( (1,2,3) )
listify((1,2,3),('a','b','c'))
# + hide_input=true
show_doc(random_split)
# -
# Splitting is done here with `random.uniform()` so you may not get the exact split percentage for small data sets
data = np.arange(20).reshape(10,2)
data.tolist()
random_split(0.20,data.tolist())
random_split(0.20,pd.DataFrame(data))
# + hide_input=true
show_doc(range_of)
# -
range_of([5,4,3])
range_of(np.arange(10)[::-1])
# + hide_input=true
show_doc(series2cat)
# -
data_df = pd.DataFrame.from_dict({"a":[1,1,1,2,2,2],"b":['f','e','f','g','g','g']})
data_df
data_df['b']
series2cat(data_df,'b')
data_df['b']
series2cat(data_df,'a')
data_df['a']
# + hide_input=true
show_doc(split_kwargs_by_func)
# -
key_word_args = {'url':'http://fast.ai','dest':'./','new_var':[1,2,3],'testvalue':42}
split_kwargs_by_func(key_word_args,download_url)
# + hide_input=true
show_doc(to_int)
# -
to_int(3.1415)
data = [1.2,3.4,7.25]
to_int(data)
# + hide_input=true
show_doc(uniqueify)
# -
uniqueify( pd.Series(data=['a','a','b','b','f','g']) )
# ## Files management and downloads
# + hide_input=true
show_doc(download_url)
# + hide_input=true
show_doc(find_classes)
# + hide_input=true
show_doc(join_path)
# + hide_input=true
show_doc(join_paths)
# + hide_input=true
show_doc(loadtxt_str)
# + hide_input=true
show_doc(save_texts)
# -
# ## Multiprocessing
# + hide_input=true
show_doc(num_cpus)
# + hide_input=true
show_doc(parallel)
# + hide_input=true
show_doc(partition)
# + hide_input=true
show_doc(partition_by_cores)
# -
# ## Data block API
# + hide_input=true
show_doc(ItemBase, title_level=3)
# -
# All items used in fastai should subclass this. Must have a [`data`](/tabular.data.html#tabular.data) field that will be used when collating in mini-batches.
# + hide_input=true
show_doc(ItemBase.apply_tfms)
# + hide_input=true
show_doc(ItemBase.show)
# -
# The default behavior is to set the string representation of this object as title of `ax`.
# + hide_input=true
show_doc(Category, title_level=3)
# -
# Create a [`Category`](/core.html#Category) with an `obj` of index [`data`](/tabular.data.html#tabular.data) in a certain classes list.
# + hide_input=true
show_doc(EmptyLabel, title_level=3)
# + hide_input=true
show_doc(MultiCategory, title_level=3)
# -
# Create a [`MultiCategory`](/core.html#MultiCategory) with an `obj` that is a collection of labels. [`data`](/tabular.data.html#tabular.data) corresponds to the one-hot encoded labels and `raw` is a list of associated string.
# + hide_input=true
show_doc(FloatItem)
# -
# ## Others
# + hide_input=true
show_doc(camel2snake)
# -
camel2snake('DeviceDataLoader')
# + hide_input=true
show_doc(even_mults)
# + hide_input=true
show_doc(func_args)
# + hide_input=true
show_doc(noop)
# -
# Return `x`.
# + hide_input=true
show_doc(one_hot)
# + hide_input=true
show_doc(show_some)
# + hide_input=true
show_doc(subplots)
# + hide_input=true
show_doc(text2html_table)
# -
# ## Undocumented Methods - Methods moved below this line will intentionally be hidden
# ## New Methods - Please document or move to the undocumented section
| docs_src/core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (geog)
# language: python
# name: geog
# ---
# # Calculate zonal nighttime lights and animated nighttime lights
#
# This notebook calculates zonal statistics on VIIRS nighttime lights for a series of industrial parks. For each of those parks, we calculate a series of buffers, and summarize nighttime lights.
#
# Finally, and animation of nighttime lights is created for a 10km buffer around the IP.
# +
import sys, os, importlib
import rasterio, boto3
import pandas as pd
import geopandas as gpd
from shapely.geometry import Point, box
sys.path.append("../../../../gostrocks/src")
import GOSTRocks.rasterMisc as rMisc
import GOSTRocks
from GOSTRocks.misc import tPrint
# +
ip_locations = "Data/IP_Locations.csv"
out_folder = "/home/wb411133/temp/ETH_NTL/"
viirs_folder = os.path.join(out_folder, "Data/VIIRS")
zonal_res = os.path.join(out_folder, "Data/IP_Locations_NTL_ZONAL.csv")
if not os.path.exists(viirs_folder):
os.makedirs(viirs_folder)
# Read in csv, re project to UTM, and write to file
inD = pd.read_csv(ip_locations)
inD_geom = [Point(x['Lon'], x['Lat']) for idx, x in inD.iterrows()]
inD = gpd.GeoDataFrame(inD, geometry=inD_geom, crs="epsg:4326")
inD = inD.to_crs("EPSG:20138")
inD.to_file(ip_locations.replace(".csv", ".geojson"), driver="GeoJSON")
# -
inD['Industrial Park'].unique()
# # Zonal Stats
# +
buffers = list(range(5, 51, 5))
buffers.append(2)
buffers.append(3)
try:
del final
except:
pass
for buf in buffers:
curD = inD.copy()
curD['geometry'] = curD.buffer(buf * 1000)
curD['BUFFER_KM'] = buf
try:
final = final.append(curD)
except:
final = curD
final = final.sort_values(["Industrial Park", "BUFFER_KM"])
inD = final
inD = inD.to_crs("epsg:4326")
inD.to_file(ip_locations.replace(".csv", "_BUFFERED.geojson"), driver="GeoJSON")
# +
def get_all_s3_keys(s3, bucket, prefix=''):
"""Get a list of all keys in an S3 bucket."""
keys = []
kwargs = {'Bucket': bucket, 'Prefix': prefix}
while True:
resp = s3.list_objects_v2(**kwargs)
for obj in resp['Contents']:
keys.append(obj['Key'])
try:
kwargs['ContinuationToken'] = resp['NextContinuationToken']
except KeyError:
break
return keys
# Get a list of the VIIRS images in S3. This example leverages the GOST teams S3 bucket
s3_base = 'wbgdecinternal-ntl'
s3 = boto3.client('s3')
all_objects = get_all_s3_keys(s3, s3_base, "NTL/VIIRS_UNZIP")
# -
sel_images = []
for obj in all_objects:
s3_path = f's3://{s3_base}/{obj}'
try:
curR = rasterio.open(s3_path)
cur_extent = box(*curR.bounds)
col_name = obj.split("/")[-1].replace(".tif", '')
if cur_extent.intersects(inD.unary_union) and (not col_name in inD.columns) \
and (not "cf_cvg" in obj) and (not "n_cf" in obj) :
tPrint(obj)
res = rMisc.zonalStats(inD, curR, minVal=0.05)
res = pd.DataFrame(res,columns=['SUM','MIN','MAX','MEAN'])
inD[col_name] = res['SUM'].values
except:
tPrint("Error Processing %s" % obj)
origD = inD.copy()
# +
# drop columns that contain counts ("n_cf")
inD = inD.loc[:,[not 'n_cf' in x for x in inD.columns]]
# rename
def fix_col_names(x):
if 'DNB' in x:
return(x.split("_")[2].split("-")[0])
else:
return(x)
new_cols = [fix_col_names(x) for x in inD.columns]
inD.columns = new_cols
# -
pd.DataFrame(inD.drop(['geometry'], axis=1)).to_csv(os.path.join(out_folder, "NTL_2020_summaries.csv"))
# # Generate animated maps
# If you want to clip out the raster data for the below mapping, run this block
sel_D = inD.loc[inD['BUFFER_KM'] == 10]
for idx, row in sel_D.iterrows():
tPrint(out_folder)
out_folder = os.path.join(viirs_folder, row['Industrial Park'])
out_map_folder = os.path.join(out_folder, "MAPS")
if not os.path.exists(out_map_folder):
os.makedirs(out_map_folder)
viirs_files = []
for cur_tif in all_files:
file = f'{cur_tif.split("/")[5]}.tif'
out_file = os.path.join(os.path.join(out_folder, "%s" % file))
viirs_files.append(out_file)
if not os.path.exists(out_file):
row['geometry'] = box(*row.geometry.bounds)
rMisc.clipRaster(rasterio.open(cur_tif), gpd.GeoDataFrame(pd.DataFrame(sel_D.loc[idx]).transpose(), geometry="geometry", crs=inD.crs), out_file)
viirs_files.sort()
out_files = []
for f in viirs_files:
out_file = os.path.join(out_map_folder, os.path.basename(f))
out_files.append(out_file)
rMisc.map_viirs(f, out_file, text_x = 0, text_y = 6)
kwargs = {'duration':0.3}
images=[]
for filename in out_files:
images.append(imageio.imread(filename))
imageio.mimsave("%s_timelapse.gif" % out_folder, images, **kwargs)
importlib.reload(rMisc)
in_tif = f
rMisc.map_viirs(f, out_file, text_x = 0, text_y = 6)
rasterio.open(in_tif).read()
importlib.reload(rMisc)
for idx, row in sel_D.iterrows():
out_folder = os.path.join(viirs_folder, row['Industrial Park'])
viirs_files = [os.path.join(out_folder, x) for x in os.listdir(out_folder) if x.endswith("tif")]
# +
import imageio
for filename in out_files:
images.append(imageio.imread(filename))
imageio.mimsave("%s_timelapse.gif" % out_folder, images, **kwargs)
# -
"%s_timelapse.gif" % out_folder
| Implementations/FY21/ZON_ETH_IPNighttimeLights/ZON_ETH_NighttimeLights.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Question 85
#
# > **_By using list comprehension, please write a program to print the list after removing the 0th,4th,5th numbers in [12,24,35,70,88,120,155]._**
#
# ---
#
# ### Hints
#
# > **_Use list comprehension to delete a bunch of element from a list.Use enumerate() to get (index, value) tuple._**
#
l = [12,24,35,70,88,120,155]
f = [v for i, v in enumerate(l) if i not in (0, 4, 5)]
print(f)
# # Question 86
#
# > **_By using list comprehension, please write a program to print the list after removing the value 24 in [12,24,35,24,88,120,155]._**
#
# ---
#
# ### Hints
#
# > **_Use list's remove method to delete a value._**
l = [12,24,35,24,88,120,155]
nl = [x for x in l if x!=24]
print(nl)
# # Question 87
#
# > **_With two given lists [1,3,6,78,35,55] and [12,24,35,24,88,120,155], write a program to make a list whose elements are intersection of the above given lists._**
#
# ---
#
# ### Hints
#
# > **_Use set() and "&=" to do set intersection operation._**
#
l1 = [1,3,6,78,35,55]
l2 = [12,24,35,24,88,120,155]
nl = list(set(l1).intersection(set(l2)))
print(nl)
# # Question 88
#
# > **_With a given list [12,24,35,24,88,120,155,88,120,155], write a program to print this list after removing all duplicate values with original order reserved._**
#
# ---
#
# ### Hints
#
# > **_Use set() to store a number of values without duplicate._**
#
li = [12,24,35,24,88,120,155,88,120,155]
lir = sorted(list(set(li[::-1])), reverse = True)
print(lir)
# # Question 89
#
# > **_Define a class Person and its two child classes: Male and Female. All classes have a method "getGender" which can print "Male" for Male class and "Female" for Female class._**
#
# ---
#
# ### Hints
#
# > **_Use Subclass(Parentclass) to define a child class._**
#
# +
class Person():
def __init__(self):
self.gender = 'Unknown'
def getGender(self):
print(self.gender)
class Male(Person):
def __init__(self):
self.gender = 'Male'
class Female(Person):
def __init__(self):
self.gender = 'Female'
san = Male()
pre = Female()
san.getGender()
pre.getGender()
# -
# # Question 90
#
# > **_Please write a program which count and print the numbers of each character in a string input by console._**
#
# > **_Example:
# > If the following string is given as input to the program:_**
#
# ```
# abcdefgabc
# ```
#
# > **_Then, the output of the program should be:_**
#
# ```
# a,2
# c,2
# b,2
# e,1
# d,1
# g,1
# f,1
# ```
#
# ### Hints
#
# > **_Use dict to store key/value pairs.
# > Use dict.get() method to lookup a key with default value._**
#
s = input('Enter a String: ')
d = dict()
for c in s:
n = s.count(c)
if c not in d.keys():
d[c] = n
for k, v in d.items():
print(f'{k},{v}')
# # Question 91
#
# > **_Please write a program which accepts a string from console and print it in reverse order._**
#
# > **Example:
# > If the following string is given as input to the program:\***
#
# ```
# rise to vote sir
# ```
#
# > **_Then, the output of the program should be:_**
#
# ```
# ris etov ot esir
# ```
#
# ### Hints
#
# > **_Use list[::-1] to iterate a list in a reverse order._**
#
s = input('Enter a String: ')
print(s[::-1])
# # Question 92
#
# > **_Please write a program which accepts a string from console and print the characters that have even indexes._**
#
# > **_Example:
# > If the following string is given as input to the program:_**
#
# ```
# H1e2l3l4o5w6o7r8l9d
# ```
#
# > **_Then, the output of the program should be:_**
#
# ```
# Helloworld
# ```
#
# ### Hints
#
# > **_Use list[::2] to iterate a list by step 2._**
s = input('Enter a String: ')
l = [v for i,v in enumerate(s) if i%2 == 0]
print(''.join(l))
# # Question 93
#
# > **_Please write a program which prints all permutations of [1,2,3]_**
#
# ---
#
# ### Hints
#
# > **_Use itertools.permutations() to get permutations of list._**
#
import itertools as i
l = [1,2,3]
nl = list()
for x in range(1, len(l)+1):
for t in list(i.permutations(l, x)):
nl.append(t)
print(nl)
# # Question 94
#
# > **_Write a program to solve a classic ancient Chinese puzzle:
# > We count 35 heads and 94 legs among the chickens and rabbits in a farm. How many rabbits and how many chickens do we have?_**
#
# ---
#
# ### Hints
#
# > **_Use for loop to iterate all possible solutions._**
#
# +
h = int(input('Heads: '))
l = int(input('Legs: '))
total = h
combos = list()
legs_of_animals = {'Rabbit': 4, 'Chicken': 2}
for i in range(1, total+1):
rabbit_legs = legs_of_animals['Rabbit']*i
chicken_legs = legs_of_animals['Chicken']*(total - i)
total_legs = rabbit_legs+chicken_legs
if total_legs == l:
combos.append((i, total-i))
for r, c in combos:
print(f'Rabbits: {r}, Chicken: {c}')
# -
# # Question 95
#
# > **_Given the participants' score sheet for your University Sports Day, you are required to find the runner-up score. You are given scores. Store them in a list and find the score of the runner-up._**
#
# > **_If the following string is given as input to the program:_**
# >
# > ```
# > 5
# > 2 3 6 6 5
# > ```
# >
# > **_Then, the output of the program should be:_**
# >
# > ```
# > 5
# > ```
#
# ### Hints
#
# > **_Make the scores unique and then find 2nd best number_**
#
n = int(input('N: '))
l = [int(i) for i in input('Enter the values: ').split()]
rs = sorted(list(set(l)))[-2]
print(rs)
# # Question 96
#
# > **_You are given a string S and width W.
# > Your task is to wrap the string into a paragraph of width._**
#
# > **_If the following string is given as input to the program:_**
# >
# > ```
# > ABCDEFGHIJKLIMNOQRSTUVWXYZ
# > 4
# > ```
# >
# > **_Then, the output of the program should be:_**
# >
# > ```
# > ABCD
# > EFGH
# > IJKL
# > IMNO
# > QRST
# > UVWX
# > YZ
# > ```
#
# ### Hints
#
# > **_Use wrap function of textwrap module_**
#
import textwrap as t
s = input('S: ')
w = int(input('W: '))
paras= t.wrap(s, w)
for p in paras:
print(p)
# # Question 97
#
# > **_You are given an integer, N. Your task is to print an alphabet rangoli of size N. (Rangoli is a form of Indian folk art based on creation of patterns.)_**
#
# > **_Different sizes of alphabet rangoli are shown below:_**
# >
# > ```
# > #size 3
# >
# > ----c----
# > --c-b-c--
# > c-b-a-b-c
# > --c-b-c--
# > ----c----
# >
# > #size 5
# >
# > --------e--------
# > ------e-d-e------
# > ----e-d-c-d-e----
# > --e-d-c-b-c-d-e--
# > e-d-c-b-a-b-c-d-e
# > --e-d-c-b-c-d-e--
# > ----e-d-c-d-e----
# > ------e-d-e------
# > --------e--------
# > ```
#
# ### Hints
#
# > **_First print the half of the Rangoli in the given way and save each line in a list. Then print the list in reverse order to get the rest._**
#
# +
import string
n = int(input('Size of Pyramid: '))
line_nos = (n*2) - 1
line_size = (line_nos)*2 - 1
chars = string.ascii_lowercase
ans = []
for i in range(n):
left = '-'.join(chars[n - i - 1:n])
mid = left[-1:0:-1] + left
final = mid.center(line_size, '-')
ans.append(final)
if len(ans) > 1:
for i in ans[n - 2::-1]:
ans.append(i)
ans = '\n'.join(ans)
print(ans)
# -
# # Question 98
#
# > **_You are given a date. Your task is to find what the day is on that date._**
#
# **Input**
#
# > **_A single line of input containing the space separated month, day and year, respectively, in MM DD YYYY format._**
# >
# > ```
# > 08 05 2015
# > ```
#
# **Output**
#
# > **_Output the correct day in capital letters._**
# >
# > ```
# > WEDNESDAY
# > ```
#
# ---
#
# ### Hints
#
# > **_Use weekday function of calendar module_**
#
import calendar as c
week_days = {0: 'MONDAY',
1: 'TUESDAY',
2: 'WEDNESDAY',
3: 'THURSDAY',
4: 'FRIDAY',
5: 'SATURDAY',
6: 'SUNDAY'}
dt = input().split()
print(dt)
wk = c.weekday(int(dt[2]), int(dt[0]), int(dt[1]))
print(week_days[wk])
# # Question 99
#
# > **_Given 2 sets of integers, M and N, print their symmetric difference in ascending order. The term symmetric difference indicates those values that exist in either M or N but do not exist in both._**
#
# **Input**
#
# > **_The first line of input contains an integer, M.The second line contains M space-separated integers.The third line contains an integer, N.The fourth line contains N space-separated integers._**
# >
# > ```
# > 4
# > 2 4 5 9
# > 4
# > 2 4 11 12
# > ```
#
# **Output**
#
# > **_Output the symmetric difference integers in ascending order, one per line._**
# >
# > ```
# > 5
# > 9
# > 11
# > 12
# > ```
#
# ---
#
# ### Hints
#
# > **_Use \'^\' to make symmetric difference operation._**
#
# +
len1 = int(input('Enter S11 len: '))
s1 = set([int(s) for s in input('Enter S1: ').split()])
len2 = int(input('Enter S2 len: '))
s2 = set([int(s) for s in input('Enter S2: ').split()])
s = set.symmetric_difference(s1, s2)
for c in s:
print(c)
# -
# # Question 100
#
# > **_You are given words. Some words may repeat. For each word, output its number of occurrences. The output order should correspond with the input order of appearance of the word. See the sample input/output for clarification._**
#
# > **_If the following string is given as input to the program:_**
# >
# > ```
# > 4
# > bcdef
# > abcdefg
# > bcde
# > bcdef
# > ```
# >
# > **_Then, the output of the program should be:_**
# >
# > ```
# > 3
# > 2 1 1
# > ```
#
# ### Hints
#
# > **_Make a list to get the input order and a dictionary to count the word frequency_**
#
# +
n = int(input())
wl = list()
wd = dict()
for i in range(n):
w = input()
if w not in wd:
wl.append(w)
wd[w] = wd.get(w, 0) + 1
print(len(wl))
for w in wl:
print(wd[w], end=' ')
# -
# # Question 101
#
# > **_You are given a string.Your task is to count the frequency of letters of the string and print the letters in descending order of frequency._**
#
# > **_If the following string is given as input to the program:_**
# >
# > ```
# > aabbbccde
# > ```
# >
# > **_Then, the output of the program should be:_**
# >
# > ```
# > b 3
# > a 2
# > c 2
# > d 1
# > e 1
# > ```
#
# ### Hints
#
# > **_Count frequency with dictionary and sort by Value from dictionary Items_**
#
s = input('S: ')
d = dict()
d = {c: s.count(c) for c in s if c not in d.keys()}
dl = list(d.items())
sorted_dl = sorted(dl, key = lambda x: x[1], reverse=True)
for t in sorted_dl:
print(t[0], t[1])
# # Question 102
#
# > **_Write a Python program that accepts a string and calculate the number of digits and letters._**
#
# **Input**
#
# > ```
# > Hello321Bye360
# > ```
#
# **Output**
#
# > ```
# > Digit - 6
# > Letter - 8
# > ```
#
# ---
#
# ### Hints
#
# > **_Use isdigit() and isalpha() function_**
#
s = input('S: ')
l = 0
d = 0
for c in s:
if c.isalpha():
l+=1
elif c.isdigit():
d+=1
else:
pass
print(f'Digit - {d}')
print(f'Letter - {l}')
# # Question 103
#
# > **_Given a number N.Find Sum of 1 to N Using Recursion_**
#
# **Input**
#
# > ```
# > 5
# > ```
#
# **Output**
#
# > ```
# > 15
# > ```
#
# ---
#
# ### Hints
#
# > **_Make a recursive function to get the sum_**
#
# +
def recursive_sum (n):
if n <=1:
return n
else:
return n + recursive_sum(n-1)
n = int(input('N: '))
print(recursive_sum(n))
| NoteBooks/Day 6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(tidytext)
library(stringr)
library(tidyverse)
library(tibble)
library(ggplot2)
library(lubridate)
library(SnowballC)
library(RSQLite)
system('head -n 1 ../../data/facebook3March18to1Sep18/0_sorted.csv', intern = TRUE)
system('tail -n 1 ../../data/facebook3March18to1Sep18/0_sorted.csv', intern = TRUE)
con = dbConnect(SQLite(), dbname="../../data/twitter.db")
tweets_df = dbGetQuery(con, 'select * from facebook where DateTime > "2018-03-03" and DateTime < "2018-05-03" ')
#tweets_df <- read.csv('../../data/facebook3March18to1Sep18/0_sorted.csv')
head(tweets_df)
# +
remove_reg <- "&|<|>"
#unnest_reg <- "([^A-Za-z_\\d#@']|'(?![A-Za-z_\\d#@]))"
tidy_tweets <- tweets_df %>%
filter(!str_detect(TweetText, "^RT")) %>%
mutate(text = str_remove_all(TweetText, remove_reg)) %>%
unnest_tokens(word, text, token = "tweets") %>%
mutate(word=wordStem(word, language = 'english')) %>%
filter(!word %in% stop_words$word,
!word %in% str_remove_all(stop_words$word, "'"),
str_detect(word, "[a-z]"))
# -
tidy_tweets$DateTime <- tidy_tweets$DateTime %>% as.Date(format = '%Y-%m-%d')
str(tidy_tweets)
summary(tidy_tweets$DateTime)
tidy_tweets %>% count(DateTime) %>%
ggplot() + aes(x=DateTime, y=n) + geom_line() +geom_point() + ylab('No of Tweets')
# Calculate frequency of words against each month
tidy_tweets_freq <- tidy_tweets %>% mutate(week = week(DateTime)) %>% group_by(week) %>% count(word, sort=TRUE) %>%
left_join(tidy_tweets %>% mutate(week = week(DateTime)) %>% group_by(week) %>% summarize(total=n())) %>%
mutate(freq = n/total) %>% mutate(logn = round(log(n))) %>% select(week, word, freq, n, logn, total)
tidy_tweets_orig <- tidy_tweets_freq
n=50
tidy_tweets_freq <- tidy_tweets_orig %>% arrange(desc(n)) %>% group_by(week) %>% top_n(freq, n=n)
tidy_tweets_freq %>% arrange(week)
# Let's convert words to columns (dimensions)
tidy_tweets_plot <- tidy_tweets_freq %>% select(-n,-total, -logn) %>% spread(week, freq, fill = 0)
tidy_tweets_ca <- tidy_tweets_freq %>% select(-n, -freq,-total) %>% spread(week, logn, fill = 0)
saveRDS(tidy_tweets_ca, file = paste0('../../data/tidy_tweets_per_week_log_facebook_',n,'.rds'))
tidy_tweets_ca
#saveRDS(tidy_tweets_plot, file = paste0('../../data/tidy_tweets_per_week_log_facebook_',n,'.rds'))
tidy_tweets_plot
# +
#tidy_tweets_plot %>% top_n(n = 1000) %>% ggplot() + aes('29','30') + geom_jitter() + geom_text(aes(label = word), check_overlap = TRUE, vjust = 0)
# -
| code/rlang/tidytext_facebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://githubtocolab.com/giswqs/leafmap/blob/master/examples/notebooks/28_publish_map.ipynb)
# [](https://gishub.org/leafmap-binder)
#
# Uncomment the following line to install [leafmap](https://leafmap.org) if needed.
# +
# # !pip install leafmap
# -
# To follow this tutorial, you will need to [sign up](https://datapane.com/accounts/signup/) for an account with <https://datapane.com>, then install and authenticate the `datapane` Python package. More information can be found [here](https://docs.datapane.com/tutorials/tut-getting-started).
#
# - `pip install datapane`
# - `datapane login`
# - `datapane ping`
import leafmap.foliumap as leafmap
# If you are using a recently implemented leafmap feature that has not yet been released to PyPI or conda-forge, you can uncomment the following line to install the development version from GitHub.
# +
# leafmap.update_package()
# -
# Create an elevation map of North America.
m = leafmap.Map()
m.add_basemap('USGS 3DEP Elevation')
colors = ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']
vmin = 0
vmax = 4000
m.add_colorbar(colors=colors, vmin=vmin, vmax=vmax)
m
# Publish the map to [datapane.com](https://datapane.com)
m.publish(name="Elevation Map of North America")
# Create a land use and land cover map.
m = leafmap.Map()
m.add_basemap("NLCD 2016 CONUS Land Cover")
m.add_legend(builtin_legend='NLCD')
m
# Publish the map to [datapane.com](https://datapane.com).
m.publish(name="National Land Cover Database (NLCD) 2016")
# Create a world population heat map.
m = leafmap.Map()
in_csv = "https://raw.githubusercontent.com/giswqs/leafmap/master/examples/data/world_cities.csv"
m.add_heatmap(in_csv, latitude="latitude", longitude='longitude', value="pop_max", name="Heat map", radius=20)
colors = ['blue', 'lime', 'red']
vmin = 0
vmax = 10000
m.add_colorbar(colors=colors, vmin=vmin, vmax=vmax)
m
# Publish the map to [datapane.com](https://datapane.com).
m.publish(name="World Population Heat Map")
| examples/notebooks/28_publish_map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:HIDA] *
# language: python
# name: conda-env-HIDA-py
# ---
from netCDF4 import Dataset
import cartopy.crs as ccrs
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# # EDA
VUL = Dataset('data/Volc_Forc_AOD_1st_mill.nc', 'r')['AOD'][:, 0, 0]
SUN = Dataset('data/Solar_forcing_1st_mill.nc', 'r')['TSI'][:, 0, 0]
# ## Target Data
fig, axes = plt.subplots(1, 2, figsize=(15,5))
axes[0].plot(SUN)
axes[0].set_title('Sun activity')
axes[1].plot(VUL, '-o')
axes[1].set_title('Volcanic activity')
# ## Temperature Data
# unpacking variables
R1 = Dataset('data/T2m_R1_ym_1stMill.nc', 'r')
temperature = R1.variables['T2m'][:]
lat = R1.variables['lat'][:]
lon = R1.variables['lon'][:]
global_temp = np.mean(temperature)
std_temp = np.std(temperature)
temperature_norm = (temperature - global_temp) / std_temp
# +
fig, axes = plt.subplots(1, 2, figsize=(15, 5), subplot_kw={'projection': ccrs.Miller()})
for kk in [0, 1]:
axes[kk].set_global()
axes[kk].coastlines()
axes[0].contourf(lon, lat, temperature_norm[0], transform=ccrs.PlateCarree(), cmap='coolwarm') # didn't use transform, but looks ok...
axes[0].set_title('Year 0')
axes[1].contourf(lon, lat, temperature_norm[-1], transform=ccrs.PlateCarree(), cmap='coolwarm') # didn't use transform, but looks ok...
axes[1].set_title('Year 999')
plt.show()
# -
# ## Find Vulcanic peak events
# +
fig, axes = plt.subplots(1, 2, figsize=(15, 5), subplot_kw={'projection': ccrs.EckertIII()})
for kk in [0, 1]:
axes[kk].set_global()
axes[kk].coastlines()
axes[0].contourf(lon, lat, temperature_norm[np.argmax(VUL)],transform=ccrs.PlateCarree(), cmap='coolwarm') # didn't use transform, but looks ok...
axes[0].set_title(f'Max Volcanic, year {np.argmax(VUL)}')
axes[1].contourf(lon, lat, temperature_norm[np.argmax(SUN)],transform=ccrs.PlateCarree(), cmap='coolwarm') # didn't use transform, but looks ok...
axes[1].set_title(f'Max Sun, year {np.argmax(SUN)}')
plt.show()
# +
fig, axes = plt.subplots(1, 2, figsize=(15, 5), subplot_kw={'projection': ccrs.PlateCarree()})
for kk in [0, 1]:
axes[kk].set_global()
axes[kk].coastlines()
axes[0].contourf(lon, lat, temperature_norm[260], cmap='coolwarm') # didn't use transform, but looks ok...
axes[0].set_title(f'Max Volcanic, year {260}')
axes[1].contourf(lon, lat, temperature_norm[811], cmap='coolwarm') # didn't use transform, but looks ok...
axes[1].set_title(f'Max Sun, year {811}')
plt.show()
# -
| netCDF_simon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Allow system to discover tf folder in import
import sys
sys.path.append('../')
# +
# Bring in helloWorld from models folder
from models import helloWorld
# Run its sample function
helloWorld.runSample()
# -
| tf/notebooks/.ipynb_checkpoints/Run Hello World-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# from mpl_toolkits.basemap import Basemap
from google.cloud import bigquery
import matplotlib.pyplot as plt
import matplotlib as mpl
import seaborn as sns
import pandas as pd
import numpy as np
import shutil
# ### Get the closest weather stations in Montreal
client = bigquery.Client()
stations = client.query("""
SELECT
name, id,
state,
latitude,
longitude,
ST_DISTANCE(
ST_GEOGPOINT(-73.573570, 45.522490),
ST_GEOGPOINT(longitude, latitude)
) AS dist_ms
FROM
`bigquery-public-data.ghcn_d.ghcnd_stations`
ORDER BY
dist_ms ASC
LIMIT
20
""")
results = stations.result()
for row in results:
print("{} : {} : {} : {} : {} : {}".format(row.name, row.id, row.state, row.latitude, row.longitude, row.dist_ms))
# ### Get the weather features from the weather stations
#
# - Time period: 2020, don't have 2017
# - prcp: rain precipitation in millimeters
# - tmin: minimum temperature in degree celsius
# - tmax: maximum temperature in degree celsius
# - hasws: if there was significant weather events or not, such as fog, hail, rain
def get_weather(year):
weather = client.query("""
SELECT
date,
MAX(prcp) AS prcp,
MAX(tmin) AS tmin,
MAX(tmax) AS tmax,
IF(MAX(haswx) = 'True', 'True', 'False') AS haswx
FROM (
SELECT
date,
IF (element = 'PRCP', value/10, NULL) AS prcp,
IF (element = 'TMIN', value/10, NULL) AS tmin,
IF (element = 'TMAX', value/10, NULL) AS tmax,
IF (SUBSTR(element, 0, 2) = 'WT', 'True', NULL) AS haswx
FROM
`bigquery-public-data.ghcn_d.ghcnd_{}`
WHERE
id = 'CA007022250'
AND qflag IS NULL)
GROUP BY
date
ORDER BY
date ASC
""".format(year))
results = weather.result()
return results
# +
data = {'date' : [], 'prcp' : [], 'tmin' : [], 'tmax' : [], 'haswx' : []}
for y in [2018, 2019, 2020]:
res = get_weather(y)
for r in res:
data["date"].append(r.date)
data["prcp"].append(r.prcp)
data["tmin"].append(r.tmin)
data["tmax"].append(r.tmax)
data["haswx"].append(r.haswx)
df = pd.DataFrame(data=data)
df['haswx'].value_counts()
# -
gs_uri = 'gs://videotron-ai-bucket/'
dataset_path = gs_uri + 'dataset/'
df.to_csv(dataset_path + 'daily_weather_montreal.csv', index=False)
| notebooks/weather.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 给定一个有 N 个节点的二叉树,每个节点都有一个不同于其他节点且处于 {1, ..., N} 中的值。
#
# 通过交换节点的左子节点和右子节点,可以翻转该二叉树中的节点。
#
# 考虑从根节点开始的先序遍历报告的 N 值序列。将这一 N 值序列称为树的行程。
#
# (回想一下,节点的先序遍历意味着我们报告当前节点的值,然后先序遍历左子节点,再先序遍历右子节点。)
#
# 我们的目标是翻转最少的树中节点,以便树的行程与给定的行程 voyage 相匹配。
#
# 如果可以,则返回翻转的所有节点的值的列表。你可以按任何顺序返回答案。
#
# 如果不能,则返回列表 [-1]。
# -
# <img src='https://assets.leetcode-cn.com/aliyun-lc-upload/uploads/2019/01/05/1219-01.png' width=200> <img src='https://assets.leetcode-cn.com/aliyun-lc-upload/uploads/2019/01/05/1219-02.png' width=200> <img src='https://assets.leetcode-cn.com/aliyun-lc-upload/uploads/2019/01/05/1219-02.png' width=200>
# + active=""
# 例1:
# 输入:root = [1,2], voyage = [2,1]
# 输出:[-1]
#
# 示例 2:
# 输入:root = [1,2,3], voyage = [1,3,2]
# 输出:[1]
#
# 示例 3:
# 输入:root = [1,2,3], voyage = [1,2,3]
# 输出:[]
#
# 提示:
# 1 <= N <= 100
# -
| DFS/1119/971. Flip Binary Tree To Match Preorder Traversal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 1: The Basics with Simple Calcium
# The system of calcium's ground state excited to its first excited state with an on-resonance laser is one of the simplest systems to model as there is only a ground S-state and an excited P-state with no hyperfine structure. We can verify results already published by [Murray 2003](https://iopscience.iop.org/article/10.1088/0953-4075/36/24/011/meta) by using the `LASED` package. In this paper the transition between the ground S-state and the first excited singlet P-state is modelled with the laser on-resonance and 200 MHz & 500 MHz detuned from resonance.
# First, we'll set up the states. A diagram of the system we are going to model is seen below. The ground S-state is represented by one sub-state lebelled as $|1\rangle$ and the excited P-state is represented by three sub-states labelled as $|2\rangle$, $|3\rangle$, and $|4\rangle$.
#
# The numbers indicate the labelling I have given them. It is convention to label the sub-states in your system by labelling the lowest in energy state first. The sub-state with the lowest $m$ value is labelled as $|1\rangle$ and then the 2nd lowest $m$ value as $|2\rangle$ etc. until you run out of sub-states. Then, the next highest in energy state is labelled.
#
# The wavelength of the difference in energy between these two states is also shown as 422.8 nm.
#
# 
# Now, we'll import the `LASED` library and a plotting library. I use `Plotly` to plot all the figures here but any plotting library can be used.
import LASED as las
from IPython.display import Image
import plotly.graph_objects as go # For plotting
import time # NOT NEEDED: For seeing how quickly the time evolution is calculated
import numpy as np
# ## Setting up the System
#
# With LASED you can declare atomic sub-states using the `State` object. These states are used to declare a `LaserAtomSystem` which can then be used to calculate the time evolution of the system.
#
# First, we must declare the system's variables: wavelength of the transition, lifetime of the excited state, isospin, etc.
#
# **Note**: LASED has a timescale of nanoseconds so all times will be input in nanoseconds. If I want a lifetime of 4.6e-9 s then I have to input 4.6 into my `LaserAtomSystem` object.
wavelength_ca = 422.8e-9 # wavelength of Ca transition in metres
tau_ca = 4.6 # lifetime in nanoseconds
I_ca = 0 # Isospin of calcium
# Create the `State` objects by providing the `label` of the sub-state with the convention as above. **The system may not be modelled correctly if you do not stick to this labelling convention**.
#
# Each sub-state must have a relative angular frequency `w` associated with it. This angular frequency is related to the energy as usual $E = \hbar w$. All frequencies should be in gigaradians per second. The energies are _relative_ so you just have to set a zero point and then all other sub-states have energies relative to this point. I have set the zero-point as the energy of the ground sub-state.
#
# Each sub-state must be labelled with its corresponding quantum numbers: orbital angular momentum `L`, spin `S`, projection of total angular momentum `m`. The isospin `I` is assumed to be zero if not specified. The total angular momentum (without isospin) can be specified with keyword `J` and is calculated as `J=L+S` if not specified. The total angular momentum with isospin can be specified with keyword `F` and if not specifed is calculated as `F=J+I`.
#
# To create the ground or excited state you must insert each sub-state in an ordered list starting with the smallest labelled state to the highest labelled state.
# +
# Calculate angular frequency of the transition
w_e = las.angularFreq(wavelength_ca) # Converted to angular frequency in Grad/s
# Create states
s1 = las.State(label = 1, w = 0, m = 0, L = 0, S = 0)
s2 = las.State(label = 2, w = w_e, m = -1, L = 1, S = 0)
s3 = las.State(label = 3, w = w_e, m = 0, L = 1, S = 0)
s4 = las.State(label = 4, w = w_e, m = 1, L = 1, S = 0)
print(s3)
# Create ground and excited states list
G_ca = [s1]
E_ca = [s2, s3, s4]
# -
# Declare the laser parameters. The intensity of the laser `laser_intensity` must be in units of mW/mm$^2$.
#
# The polarisation of the laser is defined by keyword `Q` and is either right-hand circular ($\sigma^+$) with a +1, left-hand circular ($\sigma^-$) with a -1, and linear ($\pi$) polarisation with the polarisation axis defined along the axis with a 0 (angle of polarisation of zero degrees). `Q` is defined as a list of any of these three values. If `Q` is 0 then it is defined that the linear polarisation is aligned with the x-axis (if the z-axis is the quantisation axis). This is known as the collision frame in scattering experiments.
#
# The `detuning` of the laser away from resonance can be specified as well. If no detuning is given then the laser is assumed to be on-resonance i.e. `detuning = 0`
#
# **Note on using more than one polarisation**: If more than one value is in the list then the laser is defined as having simulataneous polarisations of the same laser acting upon the atom. This can be possible if working in the natural frame when the laser is travelling along the direction of the quantisation axis and the polarisation is linear in the collision frame. When working in this natural frame `Q = [-1, 1]`. You must normalise the Rabi frequencies if you do this by a normalisation constant by giving the `LaserAtomSystem` an attribute called `rabi_scaling`. In this case of two simultaneous Rabi frequencies, the correct time evolution would be scaled by `1/np.sqrt(2)`. If `n` simultaneous polarisations are used then it must be scaled by `1/np.sqrt(n)`. You must also apply the `rabi_factors` attribute as the simultaneous combination of LHC and RHC Rabi frequencies is only equal to a linear exciation if the total Rabi frequency is $1/\sqrt(2)(\Omega^{-1}-\Omega^{+1})$ so the RHC Rabi frequency must be multiplied by -1. This is achieved by setting the `rabi_factors` attribute to `rabi_factors = [1, -1]` if using a `Q` as stated above. Each element of `Q` is multiplied by the corresponding element in `rabi_factors`.
intensity_ca = 100 # mW/mm^-2
Q_ca = [0]
detuning = 0.2*2*np.pi # detuning here is 200 MHz in Grad/s
detuning2 = 0.5*2*np.pi # detuning here is 500 MHz in Grad/s
# The `time` over which the simulation run must be sepcified. It must be specified with a list with every discrete time step (in nanoseconds) in it. Numpy's `linspace` is handy for this task.
# Simulation parameters
start_time = 0
stop_time = 50 # in ns
time_steps = 501
time_ca = np.linspace(start_time, stop_time, time_steps)
# Create a `LaserAtomSystem` object by using the variables stated above. Three system's are created here for different detunings.
calcium_system = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_intensity = intensity_ca)
calcium_system200MHzdetuned = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_intensity = intensity_ca)
calcium_system500MHzdetuned = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_intensity = intensity_ca)
# ## Time Evolution of the System
# Perform a `timeEvolution` of each system. The `pretty_print_eq` keyword is used here to print out the system's equations of motion using `Sympy`. There are many other keywords which can be used with `timeEvolution` including averaging over the doppler profile of the atoms, averaging the Gaussian laser beam profile, and numerically printing the equations.
#
# I have just timed this piece of code to see how long it takes.
#
# **Note**: An initial condition density matrix can be specified at t = 0 to evolve. If none is stated then **all ground sub-states are populated equally with no coherence between sub-states**.
tic = time.perf_counter()
calcium_system.timeEvolution(time_ca,
pretty_print_eq = True)
calcium_system200MHzdetuned.timeEvolution(time_ca,
detuning = detuning)
calcium_system500MHzdetuned.timeEvolution(time_ca,
detuning = detuning2)
toc = time.perf_counter()
print(f"The code finished in {toc-tic:0.4f} seconds")
# ## Saving and Plotting
#
# We can save the data to a .csv file and now plot the data generated to see the time evolution.
#
# To save to csv use the `saveToCSV("filename")` function on your `LaserAtomSystem` object.
calcium_system.saveToCSV("SavedData/SimpleCalciumNoDetuning.csv")
calcium_system200MHzdetuned.saveToCSV("SavedData/SimpleCalcium200MHzDetuning.csv")
calcium_system500MHzdetuned.saveToCSV("SavedData/SimpleCalcium200MHzDetuning.csv")
# Now, we can plot the evolution of Calcium with no detuning, 200 MHz detuning, and 500 MHz detuning.
#
# To access the density matrix elements over the time evolution use the function `Rho_t(e, g)` on the `LaserAtomSystem`. This gives a list of the element $\rho_{eg}$ for each interval in `time`. Each element of the density matrix is `complex` so the real part is taken by using the `abs()` function.
# +
rho_33 = [abs(rho) for rho in calcium_system.Rho_t(s3, s3)]
rho_33_200MHzdetuned = [abs(rho) for rho in calcium_system200MHzdetuned.Rho_t(s3, s3)]
rho_33_500MHzdetuned = [abs(rho) for rho in calcium_system500MHzdetuned.Rho_t(s3, s3)]
fig_ca = go.Figure()
fig_ca.add_trace(go.Scatter(x = time_ca,
y = rho_33,
mode = 'lines',
name = "Rho_33 (δ = 0 MHz)",
marker = dict(
color = 'red',
symbol = 'circle',
)))
fig_ca.add_trace(go.Scatter(x = time_ca,
y = rho_33_200MHzdetuned,
mode = 'lines',
name = "Rho_33 (δ = 200 MHz)",
marker = dict(
color = 'blue',
symbol = 'circle',
)))
fig_ca.add_trace(go.Scatter(x = time_ca,
y = rho_33_500MHzdetuned,
mode = 'lines',
name = "Rho_33 (δ = 500 MHz)",
marker = dict(
color = 'green',
symbol = 'circle',
)))
fig_ca.update_layout(title = "Calcium: Time Evolution of Upper State Population with Linear Excitation: I = 100 mW",
xaxis_title = "Time (ns)",
yaxis_title = "Population",
font = dict(
size = 11))
fig_ca.write_image("SavedPlots/tutorial1-ca.png")
Image("SavedPlots/tutorial1-ca.png")
# -
# ## Elliptical Polarisation
#
# To excite an atomic system with ellipticaly polarised light use the relation that elliptically polarisaed light can be composed of right-hand circular (RHC) $\sigma^+$ and left-hand circular (LHC) $\sigma^-$ light in varying weights. So, elliptical light can be described as:
#
# $$
# \epsilon = \frac{1}{\sqrt{L^2+R^2}}(L\sigma^-+R\sigma^+)
# $$
#
# where $R$ and $L$ are weights denoting how elliptically polarised the light is in either right or left-handed direction. If $R = L$ then the light is just linearly polarised. The ratio of L to R determines how elliptically polarised the light is.
#
# The Rabi frequency for elliptically polarisation follows from the expression above and is the superposition of the Rabi frequency for RHC and LHC light with differing weights:
#
# $$
# \Omega^{\epsilon}=\frac{1}{\sqrt{L^2+R^2}}(L\Omega^{-1}-R\Omega^{+1})
# $$
#
# To encode this in `LASED` the polarisation key word `Q = [1, -1]` with `rabi_factors = [L, R]` and `rabi_scaling = 1/np.sqrt(L*L+R*R)` for normalisation. With this description of the `LaserAtomSystem` the laser beam's direction of travel is down the quantisation axis. As an exmaple, the simple calcium system will be modelled with different weights: one system with more a more LHC ellipse and the other with a more RHC ellipse.
#
# For the first system, use values of L = 0.75 and R = 0.25.
# +
# Declare polarisation and Rabi parameters
Q_ellipse = [1, -1]
rabi_factors_lhc = [0.75, 0.25]
rabi_scaling_lhc = 1/np.sqrt(0.75*0.75+0.25*0.25)
# Create laser-atom system
calcium_system_lhc = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ellipse, wavelength_ca,
laser_intensity = intensity_ca, rabi_scaling = rabi_scaling_lhc,
rabi_factors = rabi_factors_lhc)
# Time eveolve system
calcium_system_lhc.timeEvolution(time_ca)
# Plot all excited states
rho_e = [[abs(rho) for rho in calcium_system_lhc.Rho_t(s, s)] for s in E_ca]
fig_ca_lhc = go.Figure()
for i, rho in enumerate(rho_e):
fig_ca_lhc.add_trace(go.Scatter(x = time_ca,
y = rho,
name = f"m_F = {E_ca[i].m}",
mode = 'lines'))
fig_ca_lhc.update_layout(title = "Calcium: Time Evolution of Upper States with Elliptical Polarisation (L, R) = (0.75, 0.25)",
xaxis_title = "Time (ns)",
yaxis_title = "Population",
font = dict(
size = 11))
fig_ca_lhc.write_image(f"SavedPlots/tutorial1-ellipselhc.png")
Image(f"SavedPlots/tutorial1-ellipselhc.png")
# -
# Now, model a system with a slightly more RHC elliptcal polarisation.
# +
# Rabi parameters
rabi_factors_rhc = [0.45, 0.55]
rabi_scaling_rhc = 1/np.sqrt(0.45*0.45+0.55*0.55)
# Create laser-atom system
calcium_system_rhc = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ellipse, wavelength_ca,
laser_intensity = intensity_ca, rabi_scaling = rabi_scaling_rhc,
rabi_factors = rabi_factors_rhc)
# Time eveolve system
calcium_system_rhc.timeEvolution(time_ca)
# Plot all excited states
rho_e = [[abs(rho) for rho in calcium_system_rhc.Rho_t(s, s)] for s in E_ca]
fig_ca_rhc = go.Figure()
for i, rho in enumerate(rho_e):
fig_ca_rhc.add_trace(go.Scatter(x = time_ca,
y = rho,
name = f"m_F = {E_ca[i].m}",
mode = 'lines'))
fig_ca_rhc.update_layout(title = "Calcium: Time Evolution of Upper States with Elliptical Polarisation (L, R) = (0.45, 0.55)",
xaxis_title = "Time (ns)",
yaxis_title = "Population",
font = dict(
size = 11))
fig_ca_rhc.write_image(f"SavedPlots/tutorial1-ellipserhc.png")
Image(f"SavedPlots/tutorial1-ellipserhc.png")
# -
# If L = R then this would be the same as a linear polarisation in the reference frame where the laser beam is travelling down the quantisation axis. This is called the natural frame. To put this in the collision frame, where the polarisation vector is along the quantisation axis, then rotations must be used.
# ## Setting the Initial Conditions
#
# There are two ways to specify the initial condition $\rho_0$ in `LASED`: one way is to set the entire density matrix for the system in one function using the object's variable `rho_0` directly and the other way is to set the initial condition for each sub-state equally using the function `setRho_0(s1,s2, value)` where `s1` and `s2` are state objects specifying the density matrix element which is initialised to `value`. In the example below calcium will be initialised with two different conditions.
#
# The sum of the populations of the density matrix must be equal to 1 and the coherences must be set so that $\rho_{eg} = \rho_{ge}^*$.
# +
calcium_system_init1 = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_intensity = intensity_ca)
# Set the populations of the ground state 1 and excited state 3 to 0.5 each with no coherences between them
calcium_system_init1.setRho_0(s1, s1, 0.2)
calcium_system_init1.setRho_0(s3, s3, 0.8)
# Perform time evolution
calcium_system_init1.timeEvolution(time_ca)
# Plot the excited population
rho_33_init1 = [abs(rho) for rho in calcium_system_init1.Rho_t(s3, s3)]
# -
# Now, we are going to build a flattened density matrix `rho_0` and input this straight into the system. To build the initial density matrix we must input the density matrix elements in the correct position. To do this, use the `index(e, g, n)` function to find the index which the density matrix element will sit in a flattened density matrix.
# +
calcium_system_init2 = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_intensity = intensity_ca)
# Set out the density matrix elements to be initialised
rho_11_t0 = 0.2
rho_33_t0 = 0.8
rho_13_t0 = 0.5+0.5j # These coherences can be complex!
rho_31_t0 = 0.5-0.5j
# Now build a flattened density matrix with these conditions
n = 4 # nunber of sub-states in the Calcium system
rho_0 = np.zeros((n*n, 1), dtype = complex) # Make an empty 2D array with only one column
rho_0[las.index(s1, s1, n), 0] = rho_11_t0
rho_0[las.index(s3, s3, n), 0] = rho_33_t0
rho_0[las.index(s1, s3, n), 0] = rho_13_t0
rho_0[las.index(s3, s1, n), 0] = rho_31_t0
print(rho_0)
# -
# Above is the correct form of `rho_0` we want to input. We can directly input this into the `LaserAtomSystem` object by using the key identifier `LaserAtomSystem.rho_0`.
#
# **Note**: be careful here as there is no safety conditions which have to be met to input this variable. Make sure that this is the correct size and format of the flattened density matrix.
# +
calcium_system_init2.rho_0 = rho_0 # Set the variable directly
# Perform time evolution
calcium_system_init2.timeEvolution(time_ca)
# Plot the excited population
rho_33_init2 = [abs(rho) for rho in calcium_system_init2.Rho_t(s3, s3)]
fig_ca_init = go.Figure(go.Scatter(x = time_ca,
y = rho_33_init1,
mode = 'lines',
name = "ρ<sub>33</sub> No Coherences at t=0",
marker = dict(
color = 'blue',
symbol = 'cross'
)))
fig_ca_init.add_trace(go.Scatter(x = time_ca,
y = rho_33_init2,
mode = 'lines',
name = "ρ<sub>33</sub> Coherences\n at t=0",
marker = dict(
color = 'red',
symbol = 'circle'
)))
fig_ca_init.update_layout(title = f"Calcium: Time Evolution with Differnt Initial Conditions",
xaxis_title = "Time (ns)",
yaxis_title = "Population",
font = dict(
size = 11))
fig_ca_init.write_image("SavedPlots/tutorial1-ca-init.png")
Image("SavedPlots/tutorial1-ca-init.png")
# -
# ## Gaussian Laser Beam Profiles
#
# A laser beam usually does not have a flat beam profile (known as a "top-hat" distribution) in intensity. As the beam has spatial variation in intensity the atoms being excited experience a non-uniform time evolution. To model the effects of the beam profile the beam can be split up into regions of approximate uniform intensity and each spatial portion of the beam is used to time-evolve a part of the system being illuminated. Then, each part of the system is summed together and normalised which results in the entire system being modelled.
#
# `LASED` supports the modelling of a Gaussian TEM$_{00}$ laser beam profile. The 2D standard deviation of the Gaussian must be declared with keyword `r_sigma` when performing the `timeEvolution()` of the `LaserAtomSystem`. The number of portions which the beam is split into must be chosen as well. This is declared with the keyword `n_beam_averaging` when using `timeEvolution()`. The Gaussian averaging is applied to the time evolution of the system only if it is declared that `beam_profile_averaging = True` inside the `timeEvolution()` function. If these are left out then a "top-hat" distribution of laser intensity is assumed. Also, to use the Gaussian avergaing over the beam profile, the keyword `laser_power` must be defined in the `LaserAtomSystem`. This is the total power which the laser delivers as opposed to the intensity over a mm$^2$.
#
# Below, the laser parameters are declared for this system and the simple calcium system is modelled using these parameters.
#
# **Note**: If using this averaging the model will loop over the time evolution with the number defined in `n_intensity` so the model will be much slower if a larger number is input. The larger number also results in a more accurate representation of the beam profile. Usually, a `n_intensity` of around 50 is enough for most cases.
# +
# Laser parameters
laser_power = 100 # laser intensity in mW
r_sigma = 0.75 # radial distance to 2D standard deviation in mm
n_intensity = 20
calcium_system_gaussian = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_power = laser_power)
calcium_system_gaussian.timeEvolution(time_ca,
r_sigma = r_sigma,
n_beam_averaging = n_intensity,
beam_profile_averaging = True)
# Plot the excited population
rho_33_gaussian = [abs(rho) for rho in calcium_system_gaussian.Rho_t(s3, s3)]
# -
# Now, we can plot the result and compare to the time evolution when excited with a "top-hat" beam profile.
fig_ca_gauss = go.Figure()
fig_ca_gauss.add_trace(go.Scatter(x = time_ca,
y = rho_33,
mode = 'lines',
name = "ρ<sub>33</sub> Top Hat Beam Profile",
marker = dict(
color = 'red',
symbol = 'circle'
)))
fig_ca_gauss.add_trace(go.Scatter(x = time_ca,
y = rho_33_gaussian,
mode = "lines",
name = "ρ<sub>33</sub> Gaussian Beam Profile",
marker = dict(
color = "blue",
symbol = "x"
)))
fig_ca_gauss.update_layout(title = f"Calcium: Time Evolution with Differnt Beam Profiles",
xaxis_title = "Time (ns)",
yaxis_title = "Population",
font = dict(
size = 11))
fig_ca_gauss.write_image("SavedPlots/tutorial1-ca-gaussian.png")
Image("SavedPlots/tutorial1-ca-gaussian.png")
# ### Doppler Detuning from the Atomic Velocity Profile
#
# When using `LASED` the atoms being excited are usually defined as being stationary unless specified. If the atoms are not stationary and have some velocity with respect to the laser beam then the frequency of the laser is detuned from resonance due to the fixed velocity. In experiments an atomic beam is sometimes used to provide the atoms to some interaction region where the laser-excitation takes place. If a velocity component is in (or opposite to) the direction of the laser beam then detuning occurs. The velocity component can be specified using the `atomic_velocity` keyword in the `timeEvolution()`. This is specified in units of m/s in the direction of the laser beam. If the direction is opposite to this then the `atomic_velocity` is negative.
#
# Detuning can also occur due to the Maxwell-Boltzmann distribution of atomic velocities. This results in a Gaussian detuning profile. This can be modelled by splitting the detuning due to the velocity distribution of atoms into uniform sections and time-evolving the system with these uniform detunings and then summing up the time evolution for each detuning and normalising. The detuning due to this Doppler broadening can be modelled in `LASED` by defining a `doppler_width` in Grad/s in `timeEvolution()` and a list with all the detunings to be used for the averaging process called `doppler_detunings`. The more elements in `doppler_detunings` the more the time evolution of the system is calculated and the more time it will take to model the system.
# Model the atomic velocity introducing a Doppler shift
atomic_velocity = 50 # Velocity component of atoms in direction of laser beam in m/s
# Set up the system
calcium_system_atomic_velocity = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_intensity = intensity_ca)
# Perform time evolution
calcium_system_atomic_velocity.timeEvolution(time_ca, atomic_velocity = atomic_velocity)
# Plot the excited state population
rho_33_atomic_velocity = [abs(rho) for rho in calcium_system_atomic_velocity.Rho_t(s3, s3)]
# Detuning can also occur due to the Maxwell-Boltzmann distribution of atomic velocities. This results in a Gaussian detuning profile. This can be modelled by splitting the detuning due to the velocity distribution of atoms into uniform sections and time-evolving the system with these uniform detunings and then summing up the time evolution for each detuning and normalising. The detuning due to this Doppler broadening can be modelled in `LASED` by defining a `doppler_width` in Grad/s in `timeEvolution()` and a list with all the detunings to be used for the averaging process called `doppler_detunings`. The more elements in `doppler_detunings` the more the time evolution of the system is calculated and the more time it will take to model the system. Then, use the statement `doppler_averaging = True` in the `timeEvolution()` function.
# Declare the Doppler profile parameters
doppler_width = 0.3*2*np.pi # doppler width here is 300 MHz but have to convert it into Grad/s so multiply by 2*PI and scale
delta_upper = 3*doppler_width
delta_lower = -3*doppler_width
doppler_steps = 30
doppler_detunings = np.linspace(delta_lower, delta_upper, doppler_steps)
# Set up the system
calcium_system_doppler = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_intensity = intensity_ca)
# Perform time evolution
calcium_system_doppler.timeEvolution(time_ca,
doppler_width = doppler_width,
doppler_detunings = doppler_detunings,
doppler_averaging = True)
# Plot the excited state population
rho_33_doppler = [abs(rho) for rho in calcium_system_doppler.Rho_t(s3, s3)]
# Plot the results to compare.
fig_ca_doppler = go.Figure()
fig_ca_doppler.add_trace(go.Scatter(x = time_ca,
y = rho_33,
mode = 'lines',
name = "ρ<sub>33</sub> No Doppler",
marker = dict(
color = 'red',
symbol = 'circle'
)))
fig_ca_doppler.add_trace(go.Scatter(x = time_ca,
y = rho_33_atomic_velocity,
mode = "lines",
name = "ρ<sub>33</sub> 50 m/s Atomic Velocity",
marker = dict(
color = "blue",
symbol = "x"
)))
fig_ca_doppler.add_trace(go.Scatter(x = time_ca,
y = rho_33_doppler,
mode = "lines",
name = "ρ<sub>33</sub> 300 MHz Doppler Profile",
marker = dict(
color = "green",
symbol = "x"
)))
fig_ca_doppler.update_layout(title = f"Calcium: Time Evolution with Doppler Detunings",
xaxis_title = "Time (ns)",
yaxis_title = "Population",
font = dict(
size = 11))
fig_ca_doppler.write_image("SavedPlots/tutorial1-ca-doppler.png")
Image("SavedPlots/tutorial1-ca-doppler.png")
# ## Doppler Detuning and Gaussian Beam Profile
#
# In `LASED` Doppler profile and Gaussian beam averaging can be modelled in the same system.
#
# **Note**: When using _both_ Doppler and Gaussian beam averaging the number of times the system is time evolved will be `n_intensity` multiplied by the number of elements in `doppler_detunings`.
# Set up system
calcium_system_gauss_and_dopp = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_power = laser_power)
# Time evolve the system
calcium_system_gauss_and_dopp.timeEvolution(time_ca,
r_sigma = r_sigma,
n_beam_averaging = n_intensity,
doppler_width = doppler_width,
doppler_detunings = doppler_detunings,
doppler_averaging = True,
beam_profile_averaging = True)
# Plot the excited state population
rho_33_gauss_and_dopp = [abs(rho) for rho in calcium_system_gauss_and_dopp.Rho_t(s3, s3)]
# Now, plot the results.
fig_ca_gauss_and_dopp = go.Figure()
fig_ca_gauss_and_dopp.add_trace(go.Scatter(x = time_ca,
y = rho_33,
mode = 'lines',
name = "ρ<sub>33</sub> No Doppler or Gaussian",
marker = dict(
color = 'red',
symbol = 'circle'
)))
fig_ca_gauss_and_dopp.add_trace(go.Scatter(x = time_ca,
y = rho_33_gaussian,
mode = "lines",
name = "ρ<sub>33</sub> Gaussian Beam Profile",
marker = dict(
color = "blue",
symbol = "x"
)))
fig_ca_gauss_and_dopp.add_trace(go.Scatter(x = time_ca,
y = rho_33_gauss_and_dopp,
mode = "lines",
name = "ρ<sub>33</sub> 300 MHz Doppler+Gaussian",
marker = dict(
color = "green",
symbol = "x"
)))
fig_ca_gauss_and_dopp.update_layout(title = f"Calcium: Time Evolution with Doppler and Gaussian Averaging",
xaxis_title = "Time (ns)",
yaxis_title = "Population",
font = dict(
size = 11))
fig_ca_gauss_and_dopp.write_image("SavedPlots/tutorial1-ca-gauss-and-dopp.png")
Image("SavedPlots/tutorial1-ca-gauss-and-dopp.png")
# ## Exporting the Equations of Motion
# `LASED` can print the equations of motion and/or export the equations to a .tex and a .pdf file. Note, that you must have a filename included if you want to export to a .tex file or .pdf file. Use the key idenifier `pretty_print_eq_filename` in `timeEvolution()`.
#
# **Note**: You must have `pdflatex` installed on your system to generate a .pdf file. This is used to convert the .tex file produced to a .pdf file. You can do this on Windows or Mac by installing MiKTeX on your system.
calcium_system_to_print = las.LaserAtomSystem(E_ca, G_ca, tau_ca, Q_ca, wavelength_ca,
laser_intensity = intensity_ca)
calcium_system_to_print.timeEvolution(time_ca,
pretty_print_eq = True,
pretty_print_eq_tex = True,
pretty_print_eq_pdf = True,
pretty_print_eq_filename = "CalciumSystemEquations")
| docs/source/tutorials/Tutorial1-SimpleCalcium.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Source: https://realpython.com/python-pep8/
# How to Write Beautiful Python Code With PEP 8
#
# by <NAME>
# # Why We Need PEP 8
# > “Readability counts.”
# >
# > — The Zen of Python
#
# > “Code is read much more often than it is written.”
# >
# > — <NAME>
#
# > “Any fool can write code that a computer can understand,
# >
# > good programmers write code that humans can understand.”
# >
# > — <NAME>
#
#
# - Open source collaboration
import this
# # Naming Conventions
# > “Explicit is better than implicit.”
# >
# > — The Zen of Python
#
# Note: Never use l, O, or I single letter names as these can be mistaken for 1 and 0, depending on typeface:
#
# ```python
# O = 2 # This may look like you're trying to reassign 2 to zero
# ```
# ## Naming Styles
# Type | Naming Convention | Examples
# -----|-------------------|---------
# Function | Use a lowercase word or words. Separate words by underscores to improve readability. | function, my_function
# Variable | Use a lowercase single letter, word, or words. Separate words with underscores to improve readability. | x, var, my_variable
# Class | Start each word with a capital letter. Do not separate words with underscores. This style is called camel case. | Model, MyClass
# Method | Use a lowercase word or words. Separate words with underscores to improve readability. | class_method, method
# Constant | Use an uppercase single letter, word, or words. Separate words with underscores to improve readability. | CONSTANT, MY_CONSTANT, MY_LONG_CONSTANT
# Module | Use a short, lowercase word or words. Separate words with underscores to improve readability. | module.py, my_module.py
# Package | Use a short, lowercase word or words. Do not separate words with underscores. | package, mypackage
# ## How to Choose Names
# ### Variable
# ```python
# >>> # Not recommended
# >>> x = '<NAME>'
# >>> y, z = x.split()
# >>> print(z, y, sep=', ')
# '<NAME>'
# ```
#
# ```python
# >>> # Recommended
# >>> name = '<NAME>'
# >>> first_name, last_name = name.split()
# >>> print(last_name, first_name, sep=', ')
# '<NAME>'
# ```
#
# ### Method
# ```python
# # Not recommended
# def db(x):
# return x * 2
# ```
#
# ```python
# # Recommended
# def multiply_by_two(x):
# return x * 2
# ```
# # Code Layout
# > “Beautiful is better than ugly.”
# >
# > — The Zen of Python
# ## Blank Lines
# - Surround method definitions inside classes with a single blank line.
#
# ```python
# class MyClass:
# def first_method(self):
# return None
#
# def second_method(self):
# return None
# ```
#
#
# - Use blank lines sparingly inside functions to show clear steps.
#
# ```python
# def calculate_variance(number_list):
# sum_list = 0
# for number in number_list:
# sum_list = sum_list + number
# mean = sum_list / len(number_list)
#
# sum_squares = 0
# for number in number_list:
# sum_squares = sum_squares + number**2
# mean_squares = sum_squares / len(number_list)
#
# return mean_squares - mean**2
# ```
# ## Maximum Line Length and Line Breaking
#
# Max Line Length: 79 characters
#
# ```python
# def function(arg_one, arg_two,
# arg_three, arg_four):
# return arg_one
# ```
#
# Line Breaking
#
# ```python
# from mypkg import example1, \
# example2, example3
# ```
#
# ```python
# # Recommended
# total = (first_variable
# + second_variable
# - third_variable)
# ```
#
# ```python
# # Not Recommended
# total = (first_variable +
# second_variable -
# third_variable)
# ```
# # Indentation
# > “There should be one—and preferably only one—obvious way to do it.”
# >
# > — The Zen of Python
#
# Indentation: 4 characters
#
# ```python
# x = 3
# if x > 5:
# print('x is larger than 5')
# ```
#
# - Use **4 consecutive spaces** to indicate indentation.
# - Prefer **spaces** over tabs.
# ## Indentation Following Line Breaks
# ```python
# x = 5
# if (x > 3 and
# x < 10):
# print(x)
# ```
#
# - Add a comment after the final condition. Due to syntax highlighting in most editors, this will separate the conditions from the nested code:
#
# ```python
# x = 5
# if (x > 3 and
# x < 10):
# # Both conditions satisfied
# print(x)
# ```
#
# - Add extra indentation on the line continuation:
#
# ```python
# x = 5
# if (x > 3 and
# x < 10):
# print(x)
# ```
# - hanging indent
#
# ```python
# var = function(
# arg_one, arg_two,
# arg_three, arg_four)
# ```
#
# ```python
# # Not Recommended
# var = function(arg_one, arg_two,
# arg_three, arg_four)
# ```
#
#
# ```python
# # Not Recommended
# def function(
# arg_one, arg_two,
# arg_three, arg_four):
# return arg_one
# ```
#
#
# ```python
# # Better
# def function(
# arg_one, arg_two,
# arg_three, arg_four):
# return arg_one
# ```
# ## Where to Put the Closing Brace
# - Line up the closing brace with the first non-whitespace character of the previous line:
#
# ```python
# list_of_numbers = [
# 1, 2, 3,
# 4, 5, 6,
# 7, 8, 9
# ]
# ```
#
# - Line up the closing brace with the first character of the line that starts the construct:
#
# ```python
# list_of_numbers = [
# 1, 2, 3,
# 4, 5, 6,
# 7, 8, 9
# ]
# ```
#
# You are free to chose which option you use. But, as always, **consistency is key**, so try to stick to one of the above methods.
# # Comments
# > “If the implementation is hard to explain, it’s a bad idea.”
# >
# > — The Zen of Python
#
# - Limit the line length of comments and docstrings to 72 characters.
# - Use complete sentences, starting with a capital letter.
# - Make sure to update comments if you change your code.
# ## Block Comments
# - Indent block comments to the same level as the code they describe.
# - Start each line with a # followed by a single space.
# - Separate paragraphs by a line containing a single #.
#
# ```python
# x = '<NAME>' # Student Name
# ```
#
# ```python
# student_name = '<NAME>'
# ```
#
# ```python
# empty_list = [] # Initialize empty list
#
# x = 5
# x = x * 5 # Multiply x by 5
# ```
# ## Documentation Strings
# - Surround docstrings with three double quotes on either side, as in `"""This is a docstring"""`.
#
# - Write them for all public modules, functions, classes, and methods.
#
# - Put the """ that ends a multiline docstring on a line by itself:
#
# ```python
# def quadratic(a, b, c, x):
# """Solve quadratic equation via the quadratic formula.
#
# A quadratic equation has the following form:
# ax**2 + bx + c = 0
#
# There always two solutions to a quadratic equation: x_1 & x_2.
# """
# x_1 = (- b+(b**2-4*a*c)**(1/2)) / (2*a)
# x_2 = (- b-(b**2-4*a*c)**(1/2)) / (2*a)
#
# return x_1, x_2
# ```
#
# - For one-line docstrings, keep the """ on the same line:
#
# ```python
# def quadratic(a, b, c, x):
# """Use the quadratic formula"""
# x_1 = (- b+(b**2-4*a*c)**(1/2)) / (2*a)
# x_2 = (- b-(b**2-4*a*c)**(1/2)) / (2*a)
#
# return x_1, x_2
# ```
# # Whitespace in Expressions and Statements
# > “Sparse is better than dense.”
# >
# > — The Zen of Python
#
# ```python
# # Recommended
# def function(default_parameter=5):
# # ...
#
#
# # Not recommended
# def function(default_parameter = 5):
# # ...
# ```
# ```python
# # Recommended
# y = x**2 + 5
# z = (x+y) * (x-y)
#
# # Not Recommended
# y = x ** 2 + 5
# z = (x + y) * (x - y)
# ```
# ```python
# # Not recommended
# if x > 5 and x % 2 == 0:
# print('x is larger than 5 and divisible by 2!')
# ```
#
# ```python
# # Recommended
# if x>5 and x%2==0:
# print('x is larger than 5 and divisible by 2!')
# ```
# ## When to Avoid Adding Whitespace
#
# - Immediately inside parentheses, brackets, or braces:
#
# ```python
# # Recommended
# my_list = [1, 2, 3]
#
# # Not recommended
# my_list = [ 1, 2, 3, ]
# ```
# - Before a comma, semicolon, or colon:
#
# ```python
# x = 5
# y = 6
#
# # Recommended
# print(x, y)
#
# # Not recommended
# print(x , y)
# ```
# - Before the open parenthesis that starts the argument list of a function call:
#
# ```python
# def double(x):
# return x * 2
#
# # Recommended
# double(3)
#
# # Not recommended
# double (3)
# ```
# - Before the open bracket that starts an index or slice:
#
# ```python
# # Recommended
# list[3]
#
# # Not recommended
# list [3]
# ```
# - Between a trailing comma and a closing parenthesis:
#
# ```python
# # Recommended
# tuple = (1,)
#
# # Not recommended
# tuple = (1, )
# ```
# - To align assignment operators:
#
# ```python
# # Recommended
# var1 = 5
# var2 = 6
# some_long_var = 7
#
# # Not recommended
# var1 = 5
# var2 = 6
# some_long_var = 7
# ```
# # Programming Recommendations
# > “Simple is better than complex.”
# >
# > — The Zen of Python
# Don’t compare boolean values to `True` or `False` using the equivalence operator.
# ```python
# # Not recommended
# my_bool = 6 > 5
# if my_bool == True:
# return '6 is bigger than 5'
# ```
#
#
# ```python
# # Recommended
# if my_bool:
# return '6 is bigger than 5'
# ```
# Use the fact that empty sequences are falsy in `if` statements.
#
# ```python
# # Not recommended
# my_list = []
# if not len(my_list):
# print('List is empty!')
# ```
#
#
# ```python
# # Recommended
# my_list = []
# if not my_list:
# print('List is empty!')
# ```
# Use `is not` rather than `not ... is` in `if` statements.
#
# ```python
# # Recommended
# if x is not None:
# return 'x exists!'
# ```
#
#
# ```python
# # Not recommended
# if not x is None:
# return 'x exists!'
# ```
# Don’t use `if x:` when you mean`if x is not None:`
#
# ```python
# # Not Recommended
# if arg:
# # Do something with arg...
# ```
#
#
# ```python
# # Recommended
# if arg is not None:
# # Do something with arg...
# ```
# Use `.startswith()` and `.endswith()` instead of slicing
#
# ```python
# # Not recommended
# if word[:3] == 'cat':
# print('The word starts with "cat"')
# ```
#
#
# ```python
# # Recommended
# if word.startswith('cat'):
# print('The word starts with "cat"')
# ```
# ```python
# # Not recommended
# if file_name[-3:] == 'jpg':
# print('The file is a JPEG')
# ```
#
#
# ```python
# # Recommended
# if file_name.endswith('jpg'):
# print('The file is a JPEG')
# ```
#
# - Readability & Simplicity
# # When to Ignore PEP 8
# - If complying with PEP 8 would break compatibility with existing software
# - If code surrounding what you’re working on is inconsistent with PEP 8
# - If code needs to remain compatible with older versions of Python
# # More info
# - https://realpython.com/python-pep8/
| Real-Python-PEP-8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="iOQxSDQrumWb"
# # Causal inference for decision-making in growth hacking and upselling in Python
#
# > "In this article we discuss differences between experimental and observational data and pitfalls in using the latter for data-driven decision-making."
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - categories: [python, dowhy, causal inference]
# + [markdown] id="6Oo6ETETld4t"
#
# ## Introduction
#
# Wow, growth hacking *and* upselling all in the same article? Also Python.
#
# Okay, let's start at the beginning. Imagine the following scenario: You're responsible for increasing the amount of money users spend on your e-commerce platform.
#
# You and your team come up with different measures you could implement to achieve your goal. Two of these measures could be:
#
# - Provide a discount on your best-selling items,
# - Implement a rewards program that incentivices repeat purchases.
#
# Both of these measures are fairly complex with each incurring a certain, probably known, amount of cost and an unknown effect on your customers' spending behaviour.
#
# To decide which of these two possible measures is worth both the effort and incurred cost you need to estimate their effect on customer spend.
#
# A natural way of estimating this effect is computing the following:
#
# $\textrm{avg}(\textrm{spend} | \textrm{treatment} = 1) - \textrm{avg}(\textrm{spend} | \textrm{treatment} = 0) = \textrm{ATE}$.
#
# Essentially you would compute the average spend of users who received the treatment (received a discount or signed up for rewards) and subtract from that the average spend of users who didn't receive the treatment.
#
# Without discussing the details of the underlying potential outcomes framework, the above expression is called the average treatment effect (ATE).
#
# ## Let's estimate the average treatment effect and make a decision!
#
# So now we'll just analyze our e-commerce data of treated and untreated customers and compute the average treatment effect (ATE) for each proposed measure, right? Right?
#
# Before you rush ahead with your ATE computations - now is a good time to take a step back and contemplate how your data was generated in the first place(data-generating process).
#
# ## References and further material ...
#
# Before we continue: My example here is based on a tutorial by the authors of the excellent DoWhy library. You can find the original tutorial here:
#
# https://github.com/microsoft/dowhy/blob/master/docs/source/example_notebooks/dowhy_example_effect_of_memberrewards_program.ipynb
#
# And more on DoWhy here: https://microsoft.github.io/dowhy/
# + [markdown] id="axpOMHkmSMxB"
# ## Install and load libraries
# + id="dbEmWDDng9Xm"
# !pip install dowhy --quiet
# + id="NtKCnld4-mAE"
import random
import pandas as pd
import numpy as np
np.random.seed(42)
random.seed(42)
# + [markdown] id="zdJwuUU8it_G"
# ## Randomized controlled trial / experimental data
#
# So where were we ... ah right! Where does our e-commerece data come from?
#
# Since we don't actually run an e-commerce operation here we will have to simulate our data (remember: these ideas are based on the above DoWhy tutorial).
#
# Imagine we observe the monthly spend of each of our 10,000 users over the course of a year. Each user will spend with a certain distribution (here, a Poisson distribution) and there are both high and low spenders with different mean spends.
#
# Over the course of the year, each user can sign up to our rewards program in any month and once they have signed up their spend goes up by 50% relative to what they would've spent without signing up.
#
# So far so mundane: Different customers show different spending behaviour and signing up to our rewards program increases their spend.
#
# Now the big question is: How are treatment assignment (rewards program signup) and outcome (spending behaviour) related?
#
# If treatment and outcome, interpreted as random variables, are independent of one another then according to the potential outcome framework we can compute the ATE as easily as shown above:
#
# $\textrm{ATE} = \textrm{avg}(\textrm{spend} | \textrm{treatment} = 1) - \textrm{avg}(\textrm{spend} | \textrm{treatment} = 0)$
#
# When are treatment and outcome independent? The gold standard for achieving their independence in a data set is the randomized controlled trial (RCT).
#
# In our scenario what an RCT would look like is randomly signing up our users to our rewards program - indepndent of their spending behaviour or any other characteristic.
#
# So we would go through our list of 10,000 users and flip a coin for each of them, sign them up to our program in a random month of the year based on our coin, and send them on their merry way to continue buying stuff in our online shop.
#
# Let's put all of this into a bit of code that simulates the spending behaviour of our users according to our thought experiment:
# + id="mlw4vvkie8pz"
# Creating some simulated data for our example
num_users = 10000
num_months = 12
df = pd.DataFrame({
'user_id': np.repeat(np.arange(num_users), num_months),
'month': np.tile(np.arange(1, num_months+1), num_users), # months are from 1 to 12
'high_spender': np.repeat(np.random.randint(0, 2, size=num_users), num_months),
})
df['spend'] = None
df.loc[df['high_spender'] == 0, 'spend'] = np.random.poisson(250, df.loc[df['high_spender'] == 0].shape[0])
df.loc[df['high_spender'] == 1, 'spend'] = np.random.poisson(750, df.loc[df['high_spender'] == 1].shape[0])
df["spend"] = df["spend"] - df["month"] * 10
signup_months = np.random.choice(
np.arange(1, num_months),
num_users
) * np.random.randint(0, 2, size=num_users) # signup_months == 0 means customer did not sign up
df['signup_month'] = np.repeat(signup_months, num_months)
# A customer is in the treatment group if and only if they signed up
df["treatment"] = df["signup_month"] > 0
# Simulating a simple treatment effect of 50%
after_signup = (df["signup_month"] < df["month"]) & (df["treatment"])
df.loc[after_signup, "spend"] = df[after_signup]["spend"] * 1.5
# + [markdown] id="XrY6roZSZjCj"
# Let's look at user `0` and their treatment assignment as well as spend (since we're sampling random variables here you'll see something different from me):
# + colab={"base_uri": "https://localhost:8080/", "height": 421} id="hlwbC0lLZiGw" outputId="240c8090-2994-46a0-945a-a4277b3fcf39"
df.loc[df['user_id'] == 0]
# + [markdown] id="Q7SPSlRTe8p1"
# ## Average treatment effect on post-signup spend for experimental data
#
# The effect we're interested in is the impact of rewards signup on spending behaviour - i.e. the effect on post-signup spend.
#
# Since customers can sign up any month of the year, we'll choose one month at random and compute the effect with respect to that one month.
#
# So let's create a new table from our time series where we collect post-signup spend for those customers that signed up in `month = 6` alongside the spend of customers who never signed up.
# + colab={"base_uri": "https://localhost:8080/"} id="_HxHA4GXe8p3" outputId="39dc76c8-dbcb-4193-a370-b81c1ad85dd2"
month = 6
post_signup_spend = (
df[df.signup_month.isin([0, month])]
.groupby(["user_id", "signup_month", "treatment"])
.apply(
lambda x: pd.Series(
{
"post_spend": x.loc[x.month > month, "spend"].mean(),
}
)
)
.reset_index()
)
print(post_signup_spend)
# + [markdown] id="meQD6PzFq-ML"
# To get the average treatment effect (ATE) of our rewards signup treatment we now compute the average post-signup spend of the customers who signed up and subtract from that the average spend of users who didn't sign up:
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="mispkMh35rVu" outputId="ebe193cf-a539-4275-b75f-f0b0be235626"
post_spend = post_signup_spend\
.groupby('treatment')\
.agg({'post_spend': 'mean'})
post_spend
# + [markdown] id="wOJ2rZ25rvKE"
# So the ATE of rewards signup on post-signup spend is:
# + colab={"base_uri": "https://localhost:8080/"} id="5vAxLCIoUTVT" outputId="17a2496e-a140-46df-99b2-2dd2cc04d0bc"
post_spend.loc[True, 'post_spend'] - post_spend.loc[False, 'post_spend']
# + [markdown] id="OKEJdagJr1IO"
# Since we simulated the treatment effect ourselves (50% post-signup spend increase) let's see if we can recover this effect from our data:
# + colab={"base_uri": "https://localhost:8080/"} id="Ia_DKBM6rmS5" outputId="3af5a20b-51e0-4c71-8e85-16530609d89a"
post_spend.loc[True, 'post_spend'] / post_spend.loc[False, 'post_spend']
# + [markdown] id="urSbFWzysIHV"
# The post-signup spend for treated customers is roughly 50% greater than the spend for untreated customers - exactly the treatment effect we simulated!
#
# Remember, however, that we are dealing with clean experimental data from a randomized controlled trial (RCT) here! The potential outcome framework tells us that for data from an RCT the simple ATE formula we used here yields the correct treatment effect due to independence of treatment assignment and outcome.
#
# So the fact that we recovered the actual (simulated) treatment effect is nice to see but not surprising.
# + [markdown] id="Nfg1xDo7zABH"
# ## The issue with randomized controlled trials and observational data
#
# Our above thought experiment where we randomly assigned our customers to our rewards program isn't very realistic.
#
# Randomly signing up paying customers to rewards programs without their consent may upset some and may not even be permissible. The same issue with randomized treatment assignment pops up everywhere - clean randomized controlled trials are oftentimes too expensive, infeasible to implement, unethical, or not permitted.
#
# But since we still need to experiment with our shop to drive spending behaviour we'll still go ahead and implement our rewards program. Only that this time we'll place a regular signup page in our shop where our customers can decide for themselves if they want to sign up or not.
#
# Activating our signup page and simply observing how users and their spend behaves gives us **observational data**.
#
# We usually call "observational data" just "data" without giving much thought to where they came from. I mean we've all dealt with lots of different kinds of data (marketing data, R&D measurements, HR data, etc.) and all these data were simply "observed" and didn't come out of a carefully set up experiment.
#
# Simulating our observational data we've got the same 10,000 customers over a span of a year. We still have the same high and low spenders.
#
# Only that now our high spenders are far more likely to sign up to our rewards program than our low spenders. My reasoning for this is that customers who spend more are also more likely to show greater brand loyalty towards us and our rewards program. Further, they visit our shop more frequently hence are more likely to notice our new rewards program and the signup page. We could also add this behaviour as random variables to our simulation below but just take a shortcut and give low spenders a 5% chance of signing up and high spenders a 95% chance.
# + id="vFQ5Bgf2zzKQ" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="2eef91a7-247a-45c1-fbbf-780900964a51"
num_users = 10000
num_months = 12
df = pd.DataFrame({
'user_id': np.repeat(np.arange(num_users), num_months),
'month': np.tile(np.arange(1, num_months+1), num_users), # months are from 1 to 12
'high_spender': np.repeat(np.random.randint(0, 2, size=num_users), num_months),
})
df['spend'] = None
df.loc[df['high_spender'] == 0, 'spend'] = np.random.poisson(250, df.loc[df['high_spender'] == 0].shape[0])
df.loc[df['high_spender'] == 1, 'spend'] = np.random.poisson(750, df.loc[df['high_spender'] == 1].shape[0])
signup_months = df[['user_id', 'high_spender']].drop_duplicates().copy()
signup_months['signup_month'] = None
signup_months.loc[signup_months['high_spender'] == 0, 'signup_month'] = np.random.choice(
np.arange(1, num_months),
(signup_months['high_spender'] == 0).sum()
) * np.random.binomial(1, .05, size=(signup_months['high_spender'] == 0).sum())
signup_months.loc[signup_months['high_spender'] == 1, 'signup_month'] = np.random.choice(
np.arange(1, num_months),
(signup_months['high_spender'] == 1).sum()
) * np.random.binomial(1, .95, size=(signup_months['high_spender'] == 1).sum())
df = df.merge(signup_months)
df["treatment"] = df["signup_month"] > 0
after_signup = (df["signup_month"] < df["month"]) & (df["treatment"])
df.loc[after_signup, "spend"] = df[after_signup]["spend"] * 1.5
df
# + [markdown] id="rpP-8x-VYFAa"
# Now imagine you weren't aware of causality, confounders, high / low spenders, and all that. You simply published your rewards signup page and observed your customers' spending behaviour over a span of a year. Chances are you'll compute the average treatment effect the exact same way we did above for our randomized controlled trial:
# + colab={"base_uri": "https://localhost:8080/"} id="MuQ0udCR9qYK" outputId="ec2a770f-3736-4345-87b7-f20fc34854ca"
month = 6
post_signup_spend = (
df[df.signup_month.isin([0, month])]
.groupby(["user_id", "signup_month", "treatment"])
.apply(
lambda x: pd.Series(
{
"post_spend": x.loc[x.month > month, "spend"].mean(),
"pre_spend": x.loc[x.month < month, "spend"].mean(),
}
)
)
.reset_index()
)
print(post_signup_spend)
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="yoJqX6g4Tjtw" outputId="5e17a814-b50e-4978-ffbd-8ce065e141a6"
post_spend = post_signup_spend\
.groupby('treatment')\
.agg({'post_spend': 'mean'})
post_spend
# + colab={"base_uri": "https://localhost:8080/"} id="7D74LF4kUAkB" outputId="6d8d05f5-b555-4343-e7d0-111122159cef"
post_spend.loc[True, 'post_spend'] - post_spend.loc[False, 'post_spend']
# + colab={"base_uri": "https://localhost:8080/"} id="NbeU9ZmlZGIZ" outputId="217c9554-bbae-467e-8405-20dafc6c3098"
post_spend.loc[True, 'post_spend'] / post_spend.loc[False, 'post_spend']
# + [markdown] id="W308IJJ4ZKBv"
# Performing the exact same computation as above, now we're estimating an average treatment effect of almost 400% instead of the actual 50%!
#
# So what went wrong here?
#
# Observational data got us!
#
# Realize that in our observational data the outcome (spend) is not indepndent of treatment assignmnet (rewards program signup): High spenders are far more likely to sign up hence are overrepresented in our treatment group while low spenders are overrepresented in our control group (users that didn't sign up).
#
# So when we compute the above difference or ratio we don't just see the average treatment effect of rewards signup we also see the inherent difference in spending between high and low spenders.
#
# So if we ignore how our observational data are generated we'll overestimate the effect our rewards program has and likely make decisions that seem to be supported by data but in reality aren't.
#
# Also notice that we often make this same mistake when training machine learning algorithms on observational data. Chances are someone will ask you to train a regression model to predict the effectiveness of the rewards program and your model will end up with the same inflated estimate as above.
# + [markdown] id="-wv-YTihe8p4"
# So how do we fix this? And how can we estimate the true treatment effect from our observational data?
#
# Generally, we know from experience in e-commerece that people who tend to spend more are more likely to sign up to our rewards program. So we could segment our users into spend buckets and compute the treatment effect within each bucket to try and breeak this confounding link in our observational data.
#
# Notice that in practice we won't have a `high spender` flag for our customers so we'll have to go by our customers' observed spending behaviour.
#
# The causal inference framework offers an established approach here: Relying on our domain knowledge, we define a causal model that describes how we believe our observational data were generated.
#
# Let's draw this as a graph with nodes and edges:
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="Y51lPko52klm" outputId="f02cc228-c836-41e1-a008-6bbb485ec8d4"
import os, sys
sys.path.append(os.path.abspath("../../../"))
import dowhy
causal_graph = """digraph {
treatment[label="Program Signup in month i"];
pre_spend;
post_spend;
U[label="Unobserved Confounders"];
pre_spend -> treatment;
pre_spend -> post_spend;
treatment->post_spend;
U->treatment; U->pre_spend; U->post_spend;
}"""
model = dowhy.CausalModel(
data=post_signup_spend,
graph=causal_graph.replace("\n", " "),
treatment="treatment",
outcome="post_spend"
)
model.view_model()
# + [markdown] id="QxJ6MxYXmvhD"
# Our causal model states what we described above: Pre-signup spend influences both rewards signup (treatment assignment) and post-signup spend. This is the story about our high and low spenders.
#
# Treatment (rewards signup) influences post-signup spending behaviour - this is the effect we're actually interested in.
#
# We also added a node `U` to signify possible other confounding factors that may exist in reality but weren't observed as part of our data.
# + [markdown] id="nkek0w7Re8p5"
# ## Identification / identifying the causal effect
#
# We will now apply do-calculus to our causal model from above to figure out ways to cleanly estimate the treatment effect we're after:
# + colab={"base_uri": "https://localhost:8080/"} id="j1smWlfve8p5" outputId="7d06e053-e073-4354-ac65-50c63ef8fa7f"
identified_estimand = model.identify_effect(proceed_when_unidentifiable=True)
print(identified_estimand)
# + [markdown] id="RZKrj9Qne8p6"
# Very broadly and sloppily stated there a three ways to segment (or slice and dice) our observational data to get to subsets of our data within which we can cleanly compute the average treatment effect:
#
# - Backdoor adjustment,
# - Frontdoor adjustment, and
# - Instrumental variables.
#
# The above printout tells us that based on both the causal model we constructed and our observational data there is a backdoor-adjusted estimator for our desired treatment effect.
#
# This backdoor adjustment actually follows closely what we already said above: We'll compute the post-spend given pre-spend (segment our customers based on their spending behaviour).
# + [markdown] id="KwNeZccJe8p6"
# ## Estimating the treatment effect
#
# There are various ways to perform the backdoor adjustment that DoWhy identified for us above. One of them is called propensity score matching:
# + colab={"base_uri": "https://localhost:8080/"} id="QXRu-N1ie8p7" outputId="ce197642-0ac6-4d68-c0b3-a1df51fd1f1e"
estimate = model.estimate_effect(
identified_estimand,
method_name="backdoor1.propensity_score_matching",
target_units="ate"
)
print(estimate)
# + [markdown] id="RhH_KpINxzih"
# DoWhy provides us with an estimated ATE for our observational data that is pretty close to the ATE we computed for our experimental data from our randomized controlled trial.
#
# Even if the ATE estimate DoWhy provides doesn't match exactly our experimental ATE we're now in a much better position to take a decision regarding our rewards program based on our observational data.
#
# So next time we want to base decisions on observational data it'll be worthwhile defining a causal model of the underlying data-generating process and using a library such as DoWhy that helps us identify and apply adjustment strategies.
| _notebooks/2021-08-18-Causal_inference_for_decision_making_in_growth_hacking_and_upselling_in_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["remove_input"]
from datascience import *
# %matplotlib inline
path_data = '../../data/'
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
import math
import numpy as np
from scipy import stats
# + tags=["remove_input"]
colors = Table.read_table(path_data + 'roulette_wheel.csv').column('Color')
pockets = make_array('0','00')
for i in np.arange(1, 37):
pockets = np.append(pockets, str(i))
wheel = Table().with_columns(
'Pocket', pockets,
'Color', colors
)
# -
# ### The Central Limit Theorem ###
# Very few of the data histograms that we have seen in this course have been bell shaped. When we have come across a bell shaped distribution, it has almost invariably been an empirical histogram of a statistic based on a random sample.
#
# The examples below show two very different situations in which an approximate bell shape appears in such histograms.
# ### Net Gain in Roulette ###
# In an earlier section, the bell appeared as the rough shape of the total amount of money we would make if we placed the same bet repeatedly on different spins of a roulette wheel.
wheel
# Recall that the bet on red pays even money, 1 to 1. We defined the function `red_winnings` that returns the net winnings on one \$1 bet on red. Specifically, the function takes a color as its argument and returns 1 if the color is red. For all other colors it returns -1.
def red_winnings(color):
if color == 'red':
return 1
else:
return -1
# The table `red` shows each pocket's winnings on red.
red = wheel.with_column(
'Winnings: Red', wheel.apply(red_winnings, 'Color')
)
red
# Your net gain on one bet is one random draw from the `Winnings: Red` column. There is an 18/38 chance making \$1, and a 20/38 chance of making -\$1. This probability distribution is shown in the histogram below.
red.select('Winnings: Red').hist(bins=np.arange(-1.5, 1.6, 1))
# Now suppose you bet many times on red. Your net winnings will be the sum of many draws made at random with replacement from the distribution above.
#
# It will take a bit of math to list all the possible values of your net winnings along with all of their chances. We won't do that; instead, we will approximate the probability distribution by simulation, as we have done all along in this course.
#
# The code below simulates your net gain if you bet \$1 on red on 400 different spins of the roulette wheel.
# +
num_bets = 400
repetitions = 10000
net_gain_red = make_array()
for i in np.arange(repetitions):
spins = red.sample(num_bets)
new_net_gain_red = spins.column('Winnings: Red').sum()
net_gain_red = np.append(net_gain_red, new_net_gain_red)
results = Table().with_column(
'Net Gain on Red', net_gain_red
)
# -
results.hist(bins=np.arange(-80, 50, 6))
# That's a roughly bell shaped histogram, even though the distribution we are drawing from is nowhere near bell shaped.
#
# **Center.** The distribution is centered near -20 dollars, roughly. To see why, note that your winnings will be \$1 on about 18/38 of the bets, and -$1 on the remaining 20/38. So your average winnings per dollar bet will be roughly -5.26 cents:
average_per_bet = 1*(18/38) + (-1)*(20/38)
average_per_bet
# So in 400 bets you expect that your net gain will be about -\$21:
400 * average_per_bet
# For confirmation, we can compute the mean of the 10,000 simulated net gains:
np.mean(results.column(0))
# **Spread.** Run your eye along the curve starting at the center and notice that the point of inflection is near 0. On a bell shaped curve, the SD is the distance from the center to a point of inflection. The center is roughly -\$20, which means that the SD of the distribution is around \$20.
#
# In the next section we will see where the \$20 comes from. For now, let's confirm our observation by simply calculating the SD of the 10,000 simulated net gains:
np.std(results.column(0))
# **Summary.** The net gain in 400 bets is the sum of the 400 amounts won on each individual bet. The probability distribution of that sum is approximately normal, with an average and an SD that we can approximate.
# ### Average Flight Delay ###
# The table `united` contains data on departure delays of 13,825 United Airlines domestic flights out of San Francisco airport in the summer of 2015. As we have seen before, the distribution of delays has a long right-hand tail.
united = Table.read_table(path_data + 'united_summer2015.csv')
united.select('Delay').hist(bins=np.arange(-20, 300, 10))
# The mean delay was about 16.6 minutes and the SD was about 39.5 minutes. Notice how large the SD is, compared to the mean. Those large deviations on the right have an effect, even though they are a very small proportion of the data.
# +
mean_delay = np.mean(united.column('Delay'))
sd_delay = np.std(united.column('Delay'))
mean_delay, sd_delay
# -
# Now suppose we sampled 400 delays at random with replacement. You could sample without replacement if you like, but the results would be very similar to with-replacement sampling. If you sample a few hundred out of 13,825 without replacement, you hardly change the population each time you pull out a value.
#
# In the sample, what could the average delay be? We expect it to be around 16 or 17, because that's the population average; but it is likely to be somewhat off. Let's see what we get by sampling. We'll work with the table `delay` that only contains the column of delays.
delay = united.select('Delay')
np.mean(delay.sample(400).column('Delay'))
# The sample average varies according to how the sample comes out, so we will simulate the sampling process repeatedly and draw the empirical histogram of the sample average. That will be an approximation to the probability histogram of the sample average.
# +
sample_size = 400
repetitions = 10000
means = make_array()
for i in np.arange(repetitions):
sample = delay.sample(sample_size)
new_mean = np.mean(sample.column('Delay'))
means = np.append(means, new_mean)
results = Table().with_column(
'Sample Mean', means
)
# -
results.hist(bins=np.arange(10, 25, 0.5))
# Once again, we see a rough bell shape, even though we are drawing from a very skewed distribution. The bell is centered somewhere between 16 ad 17, as we expect.
# ### Central Limit Theorem ###
#
# The reason why the bell shape appears in such settings is a remarkable result of probability theory called the **Central Limit Theorem**.
#
# **The Central Limit Theorem says that the probability distribution of the sum or average of a large random sample drawn with replacement will be roughly normal, *regardless of the distribution of the population from which the sample is drawn*.**
#
# As we noted when we were studying Chebychev's bounds, results that can be applied to random samples *regardless of the distribution of the population* are very powerful, because in data science we rarely know the distribution of the population.
#
# The Central Limit Theorem makes it possible to make inferences with very little knowledge about the population, provided we have a large random sample. That is why it is central to the field of statistical inference.
# ### Proportion of Purple Flowers ###
# Recall Mendel's probability model for the colors of the flowers of a species of pea plant. The model says that the flower colors of the plants are like draws made at random with replacement from {Purple, Purple, Purple, White}.
#
# In a large sample of plants, about what proportion will have purple flowers? We would expect the answer to be about 0.75, the proportion purple in the model. And, because proportions are means, the Central Limit Theorem says that the distribution of the sample proportion of purple plants is roughly normal.
#
# We can confirm this by simulation. Let's simulate the proportion of purple-flowered plants in a sample of 200 plants.
# +
colors = make_array('Purple', 'Purple', 'Purple', 'White')
model = Table().with_column('Color', colors)
model
# +
props = make_array()
num_plants = 200
repetitions = 10000
for i in np.arange(repetitions):
sample = model.sample(num_plants)
new_prop = np.count_nonzero(sample.column('Color') == 'Purple')/num_plants
props = np.append(props, new_prop)
results = Table().with_column('Sample Proportion: 200', props)
# -
results.hist(bins=np.arange(0.65, 0.85, 0.01))
# There's that normal curve again, as predicted by the Central Limit Theorem, centered at around 0.75 just as you would expect.
#
# How would this distribution change if we increased the sample size? Let's run the code again with a sample size of 800, and collect the results of simulations in the same table in which we collected simulations based on a sample size of 200. We will keep the number of `repetitions` the same as before so that the two columns have the same length.
# +
props2 = make_array()
num_plants = 800
for i in np.arange(repetitions):
sample = model.sample(num_plants)
new_prop = np.count_nonzero(sample.column('Color') == 'Purple')/num_plants
props2 = np.append(props2, new_prop)
results = results.with_column('Sample Proportion: 800', props2)
# -
results.hist(bins=np.arange(0.65, 0.85, 0.01))
# Both distributions are approximately normal but one is narrower than the other. The proportions based on a sample size of 800 are more tightly clustered around 0.75 than those from a sample size of 200. Increasing the sample size has decreased the variability in the sample proportion.
#
# This should not be surprising. We have leaned many times on the intuition that a larger sample size generally reduces the variability of a statistic. However, in the case of a sample average, we can *quantify* the relationship between sample size and variability.
#
# Exactly how does the sample size affect the variability of a sample average or proportion? That is the question we will examine in the next section.
| interactivecontent/understand-the-normal-curve/central-limit-theorem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from matplotlib import pyplot as plt
import cv2
import numpy as np
# +
from IPython.display import display, Javascript
from google.colab.output import eval_js
from base64 import b64decode
def take_photo(filename='photo.jpg', quality=0.8):
js = Javascript('
async function takePhoto(quality){
const div = document.createElement('div');
const capture = document.createElement('button');
capture.textContent = 'Capture';
div.appendChild(capture);
const video = document.createElement('video');
video.style.displpay = 'block';
const stream = await navigator.mediaDevices.getUserMedia({videp: True});
document.body.appendChild(div);
div.appendChild(video);
video.srObject = stream;
await video.play();
#Resize the output to fit the video element
google.colab.output.setIframeHeight(document.documentElement.scrollHeight, True);
#Wait for capture to be clicked.
await new promisse((resolve) => capture.onclick = resolve);
const canvas = document.createElement('canvas')
canvas.width = video.videoWidth;
canvas.height = video.videoHeight;
#show the image which was just taken
display(Image(filename))
except Exception as err:
#Errors will be thrown if the user does not have a webcam or if they do not
#grant the page permission to acess it
print(str(err))
# -
image = cv2.imread('photo.jpg')
ss.switchToSelectiveSearchFast()
retangulos = ss.process()
retangulos.shape
# +
#vamos pegar k retangulos aleatorios e plotar
k = 100
retangulos_aleatorios = retangulos[np.random.randint(low = 0, high = retangulos.shape[0], size = k),]
output = image.copy()
for (a, b, w, h) in retangulos[0:100,]:
color = [np.random.randint(0, 255) for j in range(0,3)]
cv2.rectangle(output, (a, b), (a + w, b + h), color, 2)
plt.imshow(output)
# -
ss.switchToSelectiveSearchQuality()
retangulos = ss.process()
retangulos.shape
# +
#vamos pegar k retangulos aleatorios e plotar
k = 100
retangulos_aleatorios = retangulos[np.random.randint(low = 0, high = retangulos.shape[0], size = k),]
output = imagecopy()
for (a, b, w, h) in retangulos[0:100]:
color = [np.random.randint(0,255) for j in range(0,3)]
cv2.rectangle(output, (a, b), (a + w, b + h),color, 2)
plt.imshow(output)
# -
| Selective Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Yet another look at Telco customer churn problem
# ## TL;DR
# ### This notebook aims at showing the difference between Kaggle world and the real-world using telco churn dataset. An example, how you could look at the problem and build applicable solution, that might be acceptable by the business.
# ### How to do some of the feature engineering and how to create training dataset that makes sense
# ### How to build a baseline model - I am not focusing on tuning the model or building multiple models, you all know how to do that better than me.
# ### I am trying to focus on consequences of the model and how important it is to put modelling into right business context and be able to show model value.
# - <a href='#1'>1. EDA</a>
# - <a href='#1.1'>1.1. Exploratory data analysis</a>
# - <a href='#1.2'>1.2. Missing Values exploration</a>
# - <a href='#1.3'>1.3. Churn breakdown</a>
# - <a href='#1.4'>1.4. Data distributions against Churn flag</a>
# - <a href='#2'>2. Feature preparation</a>
# - <a href='#2.1'>2.1. Feature engineering</a>
# - <a href='#3'>3. Modelling</a>
# - <a href='#3.1'>3.1. Analysing and transforming results</a>
# - <a href='#3.2'>3.2. Threshold selection</a>
# - <a href='#4'>4. The consequences of our model</a>
# - <a href='#4.1'>4.1. Feature Importance</a>
# - <a href='#4.2'>4.2. Unconfusing confusion matrices</a>
# - <a href='#4.3'>4.3. Campaigning the model</a>
# - <a href='#4.4'>4.4. Customer Lifetime Value</a>
# - <a href='#5'>5. Conclusions</a>
# %matplotlib inline
import pandas as pd
import numpy as np
import itertools
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelBinarizer, LabelEncoder
from sklearn.model_selection import train_test_split
import lightgbm as lgbm
from sklearn.metrics import auc, f1_score, roc_auc_score, roc_curve, confusion_matrix, accuracy_score, precision_score
import seaborn as sns
import time
pd.set_option('display.max_rows', 90)
pd.set_option('display.max_columns', 200)
pd.set_option('display.width', 1000)
# ### <a id='1'>1. EDA</a>
in_file = 'data\cell2celltrain.csv'
# ### <a id='1.1'>1.1. Exploratory data analysis</a>
df = pd.read_csv(in_file)
df.head()
df.shape
# ### Class sizes
df.Churn.value_counts()
# ### Ratios
# We take a note of class imbalance
df.Churn.value_counts()/df.shape[0]
# ### <a id='1.2'>1.2. Missing Values exploration</a>
# ### The usual suspects missing values check
# number of missing values in dataset
df.isnull().sum().values.sum()
# ### Boom! We have missing vals but why? Let's roll up sleeves and check.
# #### Despite number of techniques available, we will use common sense most of all, to determine whether or not missing value is there for a reason or missing at random
missing = list()
for x in df.columns:
if df[x].isnull().sum() != 0:
print(x, df[x].isnull().sum())
missing.append(x)
# #### From above list we can remove straight away: MonthlyRevenue, MonthlyMinutes, TotalRecurringCharge, RoamingCalls, PercChangeMinutes, PercChangeRevenues, Handsets, HandsetModels, CurrentEquipmentDays, OverageMinutes.
# #### As, above features simply could have zero value or be not present at all. We'll set them to zero 'safely'. We'll investigate the remaining lot!
# First of, what on earthe is this feature name? is the Director assisting calls?
df.DirectorAssistedCalls.describe()
# #### Stats are showing that the most values are zero, so this could be calls to call centre, where customer said
# ### 'I want to speak to your MANAGER!'
plt.figure(figsize=(7,7))
plt.grid(True)
sns.distplot(df.DirectorAssistedCalls.fillna(0))
plt.xlim(right=25)
# does it have an impact on churn?
df.Churn[df.DirectorAssistedCalls != 0].value_counts()
# ... hmmmm interesting, recall from above that our churn population in data set is 14711, and with this feature we're capturing a lot of churners. However, way more non-churnes to be fair.
# ratios against all customer population
df.Churn[df.DirectorAssistedCalls != 0].value_counts()/df.shape[0]
# ### I think we can zero-out this feature
# ### We've left with missing vals in Age let's look at stats first
df.AgeHH1.describe()
df.AgeHH2.describe()
# ### How many records are impacted by 0 age
# This could be due to many reasons:
# > customers given option not to report age
#
# > 'Clerical' error
#
# > recent change in system
#
# > ETL process crashed
#
df.Churn[(df.AgeHH1.fillna(0) == 0)&(df.AgeHH2.fillna(0) == 0)].value_counts()
# ### Some missing values conclusions.
# In my opinion is safe to 0 substitute all missing variables in this data set.
# As we showing above, there could be some data missing at random, however vast majority is due to the processes like data gathering, data entering, choice for customer that results in missing information.
df = df.fillna(0)
# ### <a id='1.3'>1.3. Churn breakdown</a>
# ## Let's take a look at churn and how it's reflected in some of the features
#get a list of categoricals
categoricals = list()
for x in df.columns:
if df[x].dtype == 'object':
categoricals.append(x)
df[categoricals].nunique()
# ### Most of categoricals are binary so let's see how the churn is reflected in some other variables
def plot_val_counts(df, col=''):
plt.figure(figsize=(5,5))
plt.grid(True)
plt.bar(df[col][df.Churn=='Yes'].value_counts().index,
df[col][df.Churn=='Yes'].value_counts().values)
plt.title(f'{col}')
plt.xticks(rotation=-90)
plot_val_counts(df, col='HandsetPrice')
plot_val_counts(df, col='CreditRating')
plot_val_counts(df, col='Occupation')
plot_val_counts(df, col='PrizmCode')
# ### <a id='1.4'>1.4. Data distributions</a>
# ### Data distributions
def plot_distro(df, col = '', y_limit=None, x_limit_r=None, x_limit_l = None):
plt.figure(figsize=(10,10))
plt.grid(True)
sns.distplot(df[col][df.Churn == 'Yes'])
sns.distplot(df[col][df.Churn == 'No'])
plt.legend(['churn_flag_yes', 'churn_flag_no'])
if y_limit:
plt.ylim(top=y_limit)
if x_limit_r:
plt.xlim(right=x_limit_r)
if x_limit_l:
plt.xlim(left=x_limit_l)
plot_distro(df, col='PercChangeMinutes', x_limit_r=1200, x_limit_l=-1200)
plot_distro(df, col='TotalRecurringCharge', x_limit_r=180)
plot_distro(df, col='DirectorAssistedCalls', y_limit=.3, x_limit_r=10)
plot_distro(df, col='MonthlyRevenue', x_limit_r=200)
plt.figure(figsize=(10,10))
plt.grid(True)
sns.boxplot(x=df.Occupation[df.Churn == 'Yes'], y=df.MonthlyRevenue[df.Churn == 'Yes'])
#sns.boxplot(x=df.Occupation[df.Churn == 'No'], y=df.MonthlyRevenue[df.Churn == 'No'])
# ### Not much of a difference visible from this perspective, let's zoom in
plt.figure(figsize=(10,10))
plt.grid(True)
sns.boxplot(x=df.Occupation, y=df.MonthlyRevenue, hue=df.Churn)
plt.ylim(top=100)
plt.figure(figsize=(10,10))
plt.grid(True)
sns.boxplot(x=df.ChildrenInHH, y=df.MonthlyRevenue, hue=df.Churn)
#sns.boxplot(x=df.ChildrenInHH[df.Churn == 'No'], y=df.MonthlyRevenue[df.Churn == 'No'])
plt.ylim(top=150)
# ### Let's note that differences are just subtle, there are no obvious relationships between Occupation and churn
# ### Customer tenure and churn
df.MonthsInService.describe()
tenure_churn = df.MonthsInService[df.Churn == 'Yes'].value_counts()
tenure_no_churn = df.MonthsInService[df.Churn == 'No'].value_counts()
tenure = pd.merge(tenure_churn.reset_index(), tenure_no_churn.reset_index(), on='index')
tenure = tenure.sort_values(by='index')
tenure = tenure.reset_index().drop(columns='level_0')
tenure.columns
plt.figure(figsize=(10,10))
plt.grid(True)
sns.pointplot(x=tenure.index, y=tenure.MonthsInService_x, color='red')
sns.pointplot(x=tenure.index, y=tenure.MonthsInService_y, color='green')
plt.xticks(rotation=90)
plt.title('When the churn picks')
# ### <a id='2.1'>2.1. Feature preparation</a>
# ### Let's look at type of features and how we could categorize them
def get_lists_of_dtypes(df):
"""
Helper function to create list of features by type and by number of unique
values they consist of.
"""
strings = list()
integers = list()
floats = list()
# Checking for partial string match to append accordingly value type
# As here we might have different type of ints and floats
# Note that strings we're returning as dictionary, to have number of unique vals for each feature
for x in df.columns[2:]:
if str(df[x].dtype)[:3] in 'obj':
strings.append({x:len(df[x].unique())})
elif str(df[x].dtype)[:3] in 'int':
integers.append(x)
elif str(df[x].dtype)[:3] in 'flo':
floats.append(x)
else:
continue
return strings,integers, floats
s, i, f = get_lists_of_dtypes(df)
# ## Let's investigate strings and number of unique values
# ### Below we can see that features with 2 unique values are already encoded
# ### as binary type [0,1] - so we can leave them as they are
# ### remaining features we will one hot encode including ServiceArea despite having 748 unique values
# ### as this feature might have an impact on model, due to larger population density
# ### therefore, might have higher/lower churn propensity
s
# ### <a id='2.1'>2.1. Feature engineering</a>
def prep_categorical_features(s):
"""
helper function to return features that we want to one hot encode
"""
one_hot = list()
binary = list()
for x in s:
for k, v in x.items():
if v > 2:
one_hot.append(k)
else:
binary.append(k)
return one_hot, binary
one_hot, binary = prep_categorical_features(s)
# ### Prep numerical pairwise features
# ### as in many feature engineering techniques we will use some of the well known tricks
# > pairwise sum, diff, ratio
#
# > min, max, mean
#
# ### Let's write a function to do just that
def pairwise(col_1, col_2):
"""
calculates pairwise features
for given two dataframe columns
"""
tot = col_1 + col_2
diff = col_1 - col_2
ratio = col_1/col_2
return tot, diff, ratio
def stats(col):
"""
calculates stats for given
dataframe column
"""
mini = col.min()
maxi = col.max()
avg = col.mean()
return mini, maxi, avg
def feature_engine_numericals(dff, i, f):
"""
Expands dataframe based on current lists of
numerical features (int, floats)
"""
numericals = i + f
df = dff.copy()
for x in numericals:
df[f'{x}_min'], df[f'{x}_max'], df[f'{x}_mean'] = stats(df[x])
for e in numericals:
if e==x:
pass
else:
df[f'sum_{x}_{e}'], df[f'diff_{x}_{e}'], df[f'ratio_{x}_{e}'] = pairwise(df[x], df[e])
return df
# %%time
pair_df = feature_engine_numericals(df, i, f)
# ### Feature engineering on categoricals transformations
def feature_engine_categoricals(dff, binary, one_hot):
"""
Function to expand dataframe by one-hot encoding
categorical variables also, changes datatype to float
"""
df = dff.copy()
lb = LabelBinarizer()
for b in binary:
df[f'{b}_tr'] = lb.fit_transform(df[b]).astype(np.float64)
df = df.drop(columns=b)
df = pd.get_dummies(df, columns=one_hot, dtype=float)
return df
# %%time
pair_df = feature_engine_categoricals(pair_df, binary, one_hot)
pair_df.shape, df.shape
pair_df.head()
# ### <a id='3'>3. Modelling</a>
# ### Encode labels and separate features and labels
le = LabelEncoder()
lab = le.fit_transform(pair_df.Churn).astype(np.float64)
l = pd.DataFrame({'lbls':pair_df.Churn, 'l_tr':lab})
l.head()
# > 1 - customers who have churned
#
# > 0 - customers retained
feats = pair_df.iloc[:,2:]
# ### Split for train test
x_train, x_test, y_train, y_test = train_test_split(feats, lab, test_size = .25, random_state = 7)
# ### Split train for train and eval
x_tr, x_ev, y_tr, y_ev = train_test_split(x_train, y_train, test_size = .05, random_state = 7)
x_tr.shape, x_ev.shape, y_tr.dtype, y_ev.dtype
train_data = lgbm.Dataset(data=x_tr, label=y_tr)
val_data = lgbm.Dataset(data=x_ev, label=y_ev)
# ### Model params
# +
# tuning copied from https://www.kaggle.com/avanwyk/a-lightgbm-overview
# Note that there is no param search here, as this is meant to be a base line model.
advanced_params = {
'boosting_type': 'gbdt',
'objective': 'binary',
'metric': 'auc',
'learning_rate': 0.1,
'num_leaves': 141, # more leaves increases accuracy, but may lead to overfitting.
'max_depth': 7, # the maximum tree depth. Shallower trees reduce overfitting.
'min_split_gain': 0, # minimal loss gain to perform a split
'min_child_samples': 21, # or min_data_in_leaf: specifies the minimum samples per leaf node.
'min_child_weight': 5, # minimal sum hessian in one leaf. Controls overfitting.
'lambda_l1': 0.5, # L1 regularization
'lambda_l2': 0.5, # L2 regularization
'feature_fraction': 0.7, # randomly select a fraction of the features before building each tree.
# Speeds up training and controls overfitting.
'bagging_fraction': 0.5, # allows for bagging or subsampling of data to speed up training.
'bagging_freq': 0, # perform bagging on every Kth iteration, disabled if 0.
'scale_pos_weight': 99, # add a weight to the positive class examples (compensates for imbalance).
'subsample_for_bin': 200000, # amount of data to sample to determine histogram bins
'max_bin': 1000, # the maximum number of bins to bucket feature values in.
# LightGBM autocompresses memory based on this value. Larger bins improves accuracy.
'nthread': 4, # number of threads to use for LightGBM, best set to number of actual cores.
}
# -
# ### Model train
# train function from https://www.kaggle.com/avanwyk/a-lightgbm-overview
def train_gbm(params, training_set, validation_set, init_gbm=None, boost_rounds=100, early_stopping_rounds=0, metric='auc'):
evals_result = {}
gbm = lgbm.train(params, # parameter dict to use
training_set,
init_model=init_gbm, # initial model to use, for continuous training.
num_boost_round=boost_rounds, # the boosting rounds or number of iterations.
early_stopping_rounds=early_stopping_rounds, # early stopping iterations.
# stop training if *no* metric improves on *any* validation data.
valid_sets=validation_set,
evals_result=evals_result, # dict to store evaluation results in.
verbose_eval=True) # print evaluations during training.
return gbm, evals_result
gbm, evals_result = train_gbm(advanced_params, training_set=train_data, validation_set=val_data,
boost_rounds=1000, early_stopping_rounds=50)
y_hat = gbm.predict(x_test)
# ### <a id='3.1'>3.1. Analysing and transforming results</a>
test_res = pd.DataFrame({'y_true':y_test, 'y_hat':y_hat})
test_res.y_hat[test_res.y_true == 0].shape, test_res.y_hat[test_res.y_true == 1].shape
roc_auc_score(test_res.y_true, test_res.y_hat)
test_res.y_hat[test_res.y_true == 0].describe()
test_res.y_hat[test_res.y_true == 1].describe()
def plot_distro(df, col = '', fiter_col = '', y_limit=None, x_limit_r=None, x_limit_l = None):
plt.figure(figsize=(10,10))
plt.grid(True)
sns.distplot(df[col][df[fiter_col] == 1])
sns.distplot(df[col][df[fiter_col] == 0])
plt.legend(['churn_flag_yes', 'churn_flag_no'])
if y_limit:
plt.ylim(top=y_limit)
if x_limit_r:
plt.xlim(right=x_limit_r)
if x_limit_l:
plt.xlim(left=x_limit_l)
plot_distro(test_res, col = 'y_hat', fiter_col = 'y_true', y_limit=None, x_limit_r=None, x_limit_l = .8)
# ### Let's transform the results to separate two distributions as far as possible
# ### <a id='3.2'>3.2. Threshold selection</a>
plt.figure(figsize=(10,10))
plt.grid(True)
sns.distplot(np.exp(test_res.y_hat[test_res.y_true == 0]), color='green')
sns.distplot(np.exp(test_res.y_hat[test_res.y_true == 1]), color='red')
plt.title('Distribution of the results, for two classes')
plt.legend(['no_churn', 'churn'])
# +
plt.figure(figsize=(12,12))
plt.grid(True)
sns.distplot(1/np.log(test_res.y_hat[test_res.y_true == 0]), color='green')
sns.distplot(1/np.log(test_res.y_hat[test_res.y_true == 1]), color='red')
plt.plot([-36.3, -36.3], [0, 0.024], 'bo--', linewidth=2.5)
plt.plot([-45, -45], [0, 0.024], 'go--', linewidth=2.5)
plt.title('Distribution of the results, for two classes with upper thresholds')
plt.legend(['best_auc_threshold','threshold_business', 'no_churn', 'churn'])
# -
# ### Now we can investigate how to select the threshold for setting churn flag = 1
# ### Above plot implies that there's no perfect separation, as this is always the case in real life
# ### Therefore, we have to look for so called 'swee tspot' where we maximise impact of the model
# ### In the industry treshold is usually determined by business who uses model
# ### In Kaggle we're looking for best AUC threshold, if that's the metric on leader board
# ### I will focus on business case here. You all know how to do it the other way ;)
# #### from below descriptive stats we could start at -39
1/np.log(test_res['y_hat'][test_res['y_true'] == 1]).describe()
# ### Transform results accordingly
test_res['y_transformed'] = 1/np.log(test_res['y_hat'])
def plot_roc_curve(test_res, threshold = -39):
ns_probs = [0 for _ in range(len(test_res))]
fpr, tpr, threshold = roc_curve(test_res.y_true, np.where(test_res.y_transformed < threshold, 1, 0))
_fpr_, _tpr_, _threshold_ = roc_curve(test_res.y_true, ns_probs)
roc_auc = auc(fpr, tpr)
plt.figure(figsize=(10,10))
plt.grid(True)
plt.title("ROC Curve. Area under Curve: {:.3f}".format(roc_auc))
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
_ = plt.plot(fpr, tpr, 'r')
__ = plt.plot(_fpr_, _tpr_, 'b', ls = '--' )
plot_roc_curve(test_res, -36.3)
plot_roc_curve(test_res, -45)
# ### <a id='4'>4. The consequences of our model</a>
# ### Each model deployed in production or considered for deployment has its consequences. Rarely spoken in ML literature in these terms. As we, Data Scientists, tend to talk in True Positive, True Negatives, False Positives, False Negatives terms. We need to bring these terms to business in their jargon, especially to be able to explain the impact and put Dollar figure on top of the results. In my experience the simpler these terms are the more successful you will be with productionizing the model, that's what we aim for, right?
# ### So, here we will focus a bit on how business see this problem and what we Data Scientist might do to maximise chance of success
test_res.y_transformed
# ### Here we'll look at confusion matrices, where we can see how change in the threshold impacts the results. Having the results in this form we have one foot in the door. But still we need more explanation.
#
# ### So, here is a little bit of industry context. Most of the companies these days have models that have customer retention as a main goal. These models are run by Marketing teams or CVM teams (Customer Value Management) or perhaps some other function. As Data Scientist you need to gain their trust. They won't buy in to Blackbox solution right away i.e. Deep Learning model, they will have to understand what's going on, under the hood. This we can achieve in multiple ways, some examples:
# > Feature importance that make sense to the business, tree-based models are good for that, but sometimes results are funny
#
# > Confusion matrix explanation and business impact of the model
# ### <a id='4.1'>4.1 Feature Importance</a>
lgbm.plot_importance(gbm, figsize=(10,12), max_num_features=15,importance_type='split' )
lgbm.plot_importance(gbm, figsize=(10,12), max_num_features=15,importance_type='gain' )
# ### From above feature importance plots, we see that almost all most important features are engineered.
# ### <a id='4.2'>4.2 Unconfusing confusion matrices</a>
def plot_conf_mat(cm):
"""
Helper function to plot confusion matrix.
With text centerred.
"""
plt.figure(figsize=(8,8))
ax = sns.heatmap(cm, annot=True,fmt="d",annot_kws={"size": 16})
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
# ### Here we visualise matrices.
# > First from auc view point
#
# > Second what I call business view point
# AUC
cm_auc = confusion_matrix(test_res.y_true, np.where(test_res.y_transformed < -36.3, 1, 0), labels=[0, 1])
plot_conf_mat(cm_auc)
# Business
cm_bus = confusion_matrix(test_res.y_true, np.where(test_res.y_transformed < -43, 1, 0), labels=[0, 1])
plot_conf_mat(cm_bus)
# Our testing population
test_res.y_true.value_counts()
# ### <a id='4.3'>4.3 Campaigning the model</a>
# ### So now, let's see how the model would be campaigned, and what would be an impact of the model
test_res.head()
# first threshold to explore is AUC oriented -36.3
auc_based = test_res[test_res.y_transformed <= -36.3]
auc_based.y_true.value_counts()
auc_based.y_true.value_counts()/auc_based.shape[0]
# ### So, our model would target over 5k customers in a campaign 60% of which wouldn't churn, 40% with higher churn propensity. Here the model would offer incentives to 3k customers that don't need them and 2k that might need them. The question here is, how many of 2k customers we would retain and how many would churn anyway?
#
# ### Unfortunately, there's no ideal solution we'd have to run campaign and study the results with A/B tests
# second threshold to explore would be Business oriented -43
bus_based = test_res[test_res.y_transformed <= -43]
bus_based.y_true.value_counts()
bus_based.y_true.value_counts()/bus_based.shape[0]
# ### <a id='4.4'>4.4 Customer Lifetime Value</a>
# ### We know now that business-oriented campaign will be smaller targeting 3.5k customers of which 55% wouldn't churn and 45% with higher churn propensity.
# ### One of the ways to look at this problem from business point of view is consider Customer Lifetime Value (CLV).
#
# ### Where CLV = Avg. revenue per period x [ Retention Rate / (1 - Retention Rate) ] as per http://www.customer-lifetimevalue.com/
#
# ### Here we will make the following assumptions. period 12 months, retention rate 0.6
df.MonthlyRevenue.describe()
# average per month and retention rate
avg_pm = 58.5
ret_rate = .6
def clv(avg_pm, ret_rate):
"""
Example calculation of CLV per year
with assumed retention rate of customers.
"""
clv = 12 * avg_pm / (ret_rate/(1-ret_rate))
return clv
cust_lv = clv(avg_pm, ret_rate)
cust_lv
# ### If we assume that retention campaign would offer and extra 10% discount per month, we can see the impact right away for both models auc and business one.
# ### To view that let's assume that campaign retains 50% of customers with high churn propensity
# so we will have 4000 customers with auc model
# and 2650 from business model
# retention rate would be the same
new_avg = 58.5 * .9
new_avg
campaign_clv = clv(new_avg, ret_rate)
campaign_clv
auc_campaing = 4000 * campaign_clv
bus_campaign = 2650 * campaign_clv
auc_campaing, bus_campaign
# ### AUC campaign discounts 3k customers at 10%, BUS campaign discounts 1.9k customers at 10%. So, during the next retention cycle, customers would expect another discount for loyalty. In my opinion is better to have campaigns optimized for precision
pr = precision_score(test_res.y_true, np.where(test_res.y_transformed < -43, 1, 0))
pr
acc = accuracy_score(test_res.y_true, np.where(test_res.y_transformed < -43, 1, 0))
acc
precision_score(test_res.y_true, np.where(test_res.y_transformed < -36.3, 1, 0))
accuracy_score(test_res.y_true, np.where(test_res.y_transformed < -36.3, 1, 0))
f1_score(test_res.y_true, np.where(test_res.y_transformed < -43, 1, 0))
f1_score(test_res.y_true, np.where(test_res.y_transformed < -36.3, 1, 0))
# ### <a id='5'>5. Conclusions:</a>
# ### In this example we looked at the churn problem with somewhat in-depth analysis. Moreover, this example should show you the journey that most of the models built in industry are going through. Before you educate business to the terms that Data Scientists are using, you might want to simplify your solution to one that you can fully explain. Very few business units would sing-up for solution they don’t trust and understand. Building trustworthy solution is rather lengthy process, so gear-up some patience and keep going it will work eventually.
# ### I would like to stress, that this is not a full solution. I got inspired by this excellent example of analysis, visualisation and multiple models in this kernel https://www.kaggle.com/pavanraj159/telecom-customer-churn-prediction.
#
# ## Next steps:
# ### Build multiple models to improve precision or accuracy, this you can always get from the business, they know their costs, once you explain the upside and downside of each solution. Setup A/B test to see how well campaigns are doing against treatment and control groups. Consider lift modelling to optimise specific KPI or business target. You also could like my github repo 😊 and upvote this kernel 😊 if you found it useful.
| churn_analysis_md.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <img src="images/csdms_logo.jpg">
# # Example 1
#
# Read and explore the output from Example 1 -- a vector parameter study that evaluates an objective function over the Rosenbrock function.
# Use `pylab` magic:
# %pylab inline
# Read the Dakota tabular data file.
dat_file = '../examples/1-rosenbrock/dakota.dat'
data = numpy.loadtxt(dat_file, skiprows=1, unpack=True, usecols=[0,2,3,4])
data
# Plot the path taken in the vector parameter study.
plot(data[1,], data[2,], 'ro')
xlim((-2, 2))
ylim((-2, 2))
xlabel('$x_1$')
ylabel('$x_2$')
title('Planview of parameter study locations')
# Plot the values of the Rosenbrock function at the study locations.
plot(data[-1,], 'bo')
xlabel('index')
ylabel('Rosenbrock fuction value')
title('Rosenbrock function values at study locations')
# What's the minimum value of the function over the study locations?
min(data[-1,:])
| notebooks/1-rosenbrock.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
var1 = "This is a module"
var1
# Area of a triangle
def triangleArea(base, height):
return (1/2) * (base * height)
# Area of a Circle
def circleArea(r):
return 3.14 * (r**2)
# Area of a square
def squareArea(length):
return length * length
| myFunctions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('../Tools/')
import DSM as dsmx
def exampleCCS():
#pimmler eppinger example 1994
## http://web.mit.edu/eppinger/www/pdf/Pimmler_DTM1994.pdf
## required = 2
## desired = 1
## indifferent = 0 (default value)
## undesired = -1
## detrimental = -2
##
## create a dict of format [ S E
## I M ]
###########################################################
ccs=dsmx.DSM(name='Climate Control System', dsmType='interactions', directed='no')
'''
ccs.addComponent('Radiator A')
ccs.addComponent('Engine Fan B')
ccs.addComponent('Heater Core C')
ccs.addComponent('Heater Hoses D')
ccs.addComponent('Condenser E')
ccs.addComponent('Compressor F')
ccs.addComponent('Evaporator Case G')
ccs.addComponent('Evaporator Core H')
ccs.addComponent('Accumulator I')
ccs.addComponent('Refrigeration Controls J')
ccs.addComponent('Air Controls K')
ccs.addComponent('Sensors L')
ccs.addComponent('Command Distribution M')
ccs.addComponent('Actuators N')
ccs.addComponent('Blower Controller O')
ccs.addComponent('Blower Motor P')
'''
#input all components (not required)
ccs.addComponent(['A','B','C','D','E','F','G','H','I','J','K','L','M','N','O','P'])
#input interactions
ccs.addRelation(['A'],['B'], [{'s':2, 'e':0, 'i':0, 'm':2}])
ccs.addRelation(['A'],['E'], [{'s':2, 'e':-2, 'i':0, 'm':0}])
ccs.addRelation(['B'],['E'], [{'s':2, 'e':0, 'i':0, 'm':2}])
ccs.addRelation(['B'],['M'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['C'],['D'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['C'],['G'], [{'s':2, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['C'],['H'], [{'s':-1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['C'],['P'], [{'s':0, 'e':0, 'i':0, 'm':2}])
ccs.addRelation(['D'],['I'], [{'s':-1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['E'],['F'], [{'s':0, 'e':2, 'i':0, 'm':2}])
ccs.addRelation(['E'],['H'], [{'s':-2, 'e':2, 'i':0, 'm':2}])
ccs.addRelation(['F'],['H'], [{'s':0, 'e':2, 'i':0, 'm':2}])
ccs.addRelation(['F'],['I'], [{'s':1, 'e':0, 'i':0, 'm':2}])
ccs.addRelation(['F'],['J'], [{'s':0, 'e':0, 'i':2, 'm':0}])
ccs.addRelation(['F'],['K'], [{'s':0, 'e':0, 'i':2, 'm':0}])
ccs.addRelation(['F'],['M'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['G'],['H'], [{'s':2, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['G'],['N'], [{'s':2, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['G'],['O'], [{'s':2, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['G'],['P'], [{'s':2, 'e':0, 'i':0, 'm':2}])
ccs.addRelation(['H'],['I'], [{'s':1, 'e':0, 'i':0, 'm':2}])
ccs.addRelation(['H'],['P'], [{'s':0, 'e':0, 'i':0, 'm':2}])
ccs.addRelation(['I'],['J'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['J'],['K'], [{'s':0, 'e':0, 'i':2, 'm':0}])
ccs.addRelation(['J'],['M'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['K'],['L'], [{'s':0, 'e':0, 'i':2, 'm':0}])
ccs.addRelation(['K'],['M'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['K'],['N'], [{'s':0, 'e':0, 'i':2, 'm':0}])
ccs.addRelation(['K'],['O'], [{'s':0, 'e':0, 'i':2, 'm':0}])
ccs.addRelation(['L'],['M'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['M'],['N'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['M'],['O'], [{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['M'],['P'],[{'s':1, 'e':0, 'i':0, 'm':0}])
ccs.addRelation(['O'],['P'], [{'s':2, 'e':0, 'i':0, 'm':2}])
return ccs
# -
ccs=exampleCCS()
ccs.display()
#ccs.to2Dinteractions({'s':0.45, 'e':0.2, 'i':0.1, 'm':0.25})
| Notebooks/Climate Control System.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %cd ..
# +
import numpy as np
import torch
import torch.nn as nn
import torchaudio
from torchaudio import transforms
import librosa
import matplotlib.pyplot as plt
from src.data.midi import parse_midi
from src.data.data_modules import MAPSDataModule
# -
sample_rate = 16000
n_mels = 229
hop_length = 512
n_fft = 2048
# +
audio_path = "data/processed/MAPS_MUS/AkPnBsdf/MAPS_MUS-alb_se3_AkPnBsdf.wav"
wav, sr = torchaudio.load(audio_path)
wav = wav.mean(dim=0)
wav_16k = transforms.Resample(sr, new_freq=16000)(wav)
mel = transforms.MelSpectrogram(n_mels=229, hop_length=512, n_fft=2048)(wav_16k)
mel.shape
# -
transforms.MelSpectrogram(n_mels=229, hop_length=512, n_fft=2048).hop_length
plt.figure(figsize=(20,20))
plt.imshow(mel.log2()[1,:,:1000].numpy())
# quantize labels from MIDI file
midi_path = "data/raw/MAPS/ENSTDkAm2/ENSTDkAm/MUS/MAPS_MUS-bk_xmas1_ENSTDkAm.mid"
note_df = parse_midi(midi_path)
note_df.head()
# +
onset_length_in_ms = 32
onset_length_in_samples = sample_rate * onset_length_in_ms // 1000
hops_in_onset = onset_length_in_samples // hop_length
offset_length_in_ms = 32
offset_length_in_samples = sample_rate * offset_length_in_ms // 1000
hops_in_offset = offset_length_in_samples // hop_length
MIDI_MIN_PITCH = 21
MIDI_MAX_PITCH = 108
n_pitches = MIDI_MAX_PITCH - MIDI_MIN_PITCH + 1
# +
# quantize continous note labels into discrete hops
n_hops = mel.shape[1]
frame_labels = torch.zeros(n_hops, n_pitches, dtype=torch.uint8)
velocity_arr = torch.zeros(n_hops, n_pitches, dtype=torch.uint8)
for _, (onset, offset, pitch, velocity) in note_df.iterrows():
onset_start = int(round(onset * sample_rate / hop_length))
onset_end = min(n_hops, onset_start + hops_in_onset)
frame_end = int((round(offset * sample_rate / hop_length)))
frame_end = min(n_hops, frame_end)
offset_end = min(n_hops, frame_end + hops_in_offset)
p = int(pitch) - MIDI_MIN_PITCH
frame_labels[onset_start:onset_end, p] = 3
frame_labels[onset_end:frame_end, p] = 2
frame_labels[frame_end:offset_end, p] = 1
velocity_arr[onset_start:offset_end, p] = velocity
# +
# test pytorch dataset
from src.data.datasets import MAPSDataset
max_steps = int((5 * 16000) / 512)
ds = MAPSDataset(data_dir="data/processed/MAPS_MUS/", subsets=["AkPnBcht", "AkPnBsdf"],
max_steps=max_steps,
audio_transform=transforms.MelSpectrogram(n_mels=229, hop_length=512, n_fft=2048))
# -
from src.data.data_modules import MAPSDataModule
dm = MAPSDataModule(batch_size=4)
dm.setup()
dm.size()
loader = dm.train_dataloader()
sample = next(iter(loader))
sample["audio"].shape, sample["onsets"].shape
| notebooks/1.1-duc-dataset-prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
#
# ## NRPyPN: Validated Post-Newtonian Expressions for Input into Wolfram Mathematica, SymPy, or Highly-Optimized C Codes
#
# ### Lead author: [<NAME>](http://astro.phys.wvu.edu/zetienne/) $\leftarrow$ Please feel free to email contributions, comments, revisions, or errata!
#
# ### Introduction
#
# [Post-Newtonian theory](https://en.wikipedia.org/wiki/Post-Newtonian_expansion) results in some of the longest and most complex mathematical expressions ever derived by humanity.
#
# These expressions form the core of most gravitational wave data analysis codes, but generally the expressions are written in a format that is either inaccessible by others (i.e., closed-source) or written directly in e.g., C code and thus incompatible with symbolic algebra packages like [Wolfram Mathematica](https://www.wolfram.com/mathematica/) or the Python-based [SymPy](https://www.sympy.org/).
#
# Once in a symbolic algebra package, these expressions could be manipulated, extended, and output as *more* optimized C codes (e.g., using SymPy/NRPy+, thus speeding up gravitational wave data analysis).
#
# This repository aims to provide a trusted source for validated Post-Newtonian expressions useful for gravitational wave astronomy, using the open-source [SymPy](https://www.sympy.org/) computer algebra system, so that expressions can be output into [Wolfram Mathematica](https://www.wolfram.com/mathematica/) or highly optimized C codes using the SymPy-based [NRPy+](http://nrpyplus.net/).
#
# ***If you are unfamiliar with using Jupyter Notebooks, first review the official [Jupyter Notebook Basics Guide](https://nbviewer.jupyter.org/github/jupyter/notebook/blob/master/docs/source/examples/Notebook/Notebook%20Basics.ipynb).***
#
# ### PART 0: Basic Functions for Expediting Equation Imports
#
# + [NRPyPN_shortcuts](NRPyPN_shortcuts.ipynb)
#
# ### PART 1: Post-Newtonian Hamiltonian for Binary Black Holes
#
# The Post-Newtonian Hamiltonian $H$ can be written as a sum of contributions:
# \begin{array}
# \ H = H_{\rm Orb, NS} + H_{\rm SO} + H_{\rm SS} + H_{\rm SSS},
# \end{array}
# where
# * $H_{\rm Orb, NS}$ is the non-spinning, purely orbital contribution
# * $H_{\rm SO}$ accounts for spin-orbit coupling
# * $H_{\rm SS}$ accounts for spin-spin contributions
# * $H_{\rm SSS}$ accounts for spin-spin-spin contributions
#
# Click on any term below to open a notebook implementing that Hamiltonian component.
# #### $H_{\rm Orb, NS}$, up to and including third post-Newtonian order (3PN)
# + [$H_{\rm Orb, NS}$](PN-Hamiltonian-Nonspinning.ipynb), as summarized in [Buonanno, Chen, and Damour (2006)](https://arxiv.org/abs/gr-qc/0508067) (see references therein for sources)
#
# #### $H_{\rm SO}$, up to and including 3.5 post-Newtonian order (3.5PN)
# + [The $H_{\rm SO}$ notebook](PN-Hamiltonian-Spin-Orbit.ipynb) contains all spin-orbit coupling terms up to and including 3.5PN order:
# * 1.5PN order (i.e., $H_{\rm SO, 1.5PN}$), as summarized in [Buonanno, Chen, and Damour (2006)](https://arxiv.org/abs/gr-qc/0508067) (see references therein for sources)
# * 2.5PN order (i.e., $H_{\rm SO, 2.5PN}$), as derived in [Damour, Jaranowski, and Schäfer (2008)](https://arxiv.org/abs/0711.1048)
# * 3.5PN order (i.e., $H_{\rm SO, 3.5PN}$), as derived in [Hartung and Steinhoff (2011)](https://arxiv.org/abs/1104.3079)
#
# #### $H_{\rm SS}$, up to and including third post-Newtonian order (3PN)
# + [The $H_{\rm SS}$ notebook](PN-Hamiltonian-Spin-Spin.ipynb) contains all spin-orbit coupling terms up to and including 3PN order
# + 2PN order (i.e., $H_{S_1,S_2,{\rm 2PN}}+H_{S_1^2,{\rm 2PN}}+H_{S_2^2,{\rm 2PN}}$), as summarized in [Buonanno, Chen, and Damour (2006)](https://arxiv.org/abs/gr-qc/0508067) (see references therein for sources)
# + $S_1,S_2$ term at 3PN order (i.e., $H_{S_1,S_2,{\rm 3PN}}$), as derived in [Steinhoff, Hergt, and Schäfer (2008a)](https://arxiv.org/abs/0712.1716)
# + $S_1^2$ term at 3PN order (i.e., $H_{S_1^2,{\rm 3PN}}$), as derived in [Steinhoff, Hergt, and Schäfer (2008b)](https://arxiv.org/abs/0809.2200)
#
# #### $H_{\rm SSS}$, up to and including third post-Newtonian order (3PN)
# + [$H_{SSS,{\rm 3PN}}$](PN-Hamiltonian-SSS.ipynb), as derived in [Levi and Steinhoff (2015)](https://arxiv.org/abs/1410.2601)
#
# ### Part 2: $\frac{dE_{\rm GW}}{dt}$, the gravitational wave flux, and $\frac{dM}{dt}$, the tidal energy injected into the black holes
#
# + [Gravitational-wave and mass fluxes $\frac{dE_{\rm GW}}{dt}$ and $\frac{dM}{dt}$](PN-dE_GW_dt_and_dM_dt.ipynb), as reviewed by
# * [Blanchet (2014)](https://link.springer.com/content/pdf/10.12942/lrr-2014-2.pdf), for the nonspinning terms
# * [Ossokine *et al* (2015)](https://arxiv.org/abs/1502.01747), including precessing spin terms
# * [Ajith *et al* (2007)](https://arxiv.org/abs/0709.0093), including tidal-heating injected energy into the black holes [Alvi (2001)](https://arxiv.org/abs/gr-qc/0107080)
#
# ### Part 3: Quasicircular Orbital Parameters $p_t$ and $p_r$
#
# + [Tangential component of momentum $p_t$](PN-p_t.ipynb), as derived by
# * [Ramos-Buades, Husa, and Pratten (2018)](https://arxiv.org/abs/1810.00036) up to and including 3.5PN order
# * ... and validated against [Healy, Lousto, Nakano, and Zlochower (2017)](https://arxiv.org/abs/1702.00872), who derive the expression up to and including 3PN order
# + [Orbital angular frequency $M\Omega$](PN-MOmega.ipynb), as derived by
# * [<NAME>, and Pratten (2018)](https://arxiv.org/abs/1810.00036) up to and including 3.5PN order
# * ... and validated against [Healy, Lousto, Nakano, and Zlochower (2017)](https://arxiv.org/abs/1702.00872), who derive the expression up to and including 3PN order
# + [Radial component of momentum $p_r$](PN-p_r.ipynb), as derived by
# * [Healy, Lousto, Nakano, and Zlochower (2017)](https://arxiv.org/abs/1702.00872) up to and including 3PN order
# * [<NAME>, and Pratten (2018)](https://arxiv.org/abs/1810.00036) up to and including 3.5PN order
# ## Fast $p_t$ and $p_r$ for binary black hole initial data
# +
#### INPUT PARAMETERS:
qmassratio = 4.0 # m2/m1; by convention must be >= 1
nr = 13.0 # Orbital separation
# Dimensionless spin parameters of each black hole
nchi1x = +0. # chi_{1x}, x-component of spin vector for black hole 1
nchi1y = +0.1
nchi1z = +0.
nchi2x = -0.3535
nchi2y = +0.3535
nchi2z = +0.5
#### DON'T TOUCH; see output after running this cell.
from NRPyPN_shortcuts import * # NRPyPN: shortcuts for e.g., vector operations
# Compute p_t, the radial component of momentum
import PN_p_t as pt
pt.f_p_t(m1,m2, chi1U,chi2U, r)
# Compute p_r, the radial component of momentum
import PN_p_r as pr
pr.f_p_r(m1,m2, n12U,n21U, chi1U,chi2U, S1U,S2U, p1U,p2U, r)
nPt = num_eval(pt.p_t,
qmassratio=qmassratio, nr=nr,
nchi1x=nchi1x,nchi1y=nchi1y,nchi1z=nchi1z,
nchi2x=nchi2x,nchi2y=nchi2y,nchi2z=nchi2z)
np_r = num_eval(pr.p_r.subs(gamma_EulerMascheroni,0.5772156649015328606065120900824024310421),
qmassratio = qmassratio, nr=nr,
nchi1x=nchi1x, nchi1y=nchi1y, nchi1z=nchi1z,
nchi2x=nchi2x, nchi2y=nchi2y, nchi2z=nchi2z, nPt=nPt)
print("P_t = ",nPt)
print("P_r = ",np_r)
| NRPyPN/NRPyPN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''algotrading'': conda)'
# name: python3
# ---
# # Historical Stock Prices
# Import libraries
import yfinance as yf
from yahoofinancials import YahooFinancials
import pandas as pd
# +
# Input one or several stock tickers to obtain historical stock prices
ticker = "BRK-B"
start= "2015-08-07"
end= "2021-07-20"
# for multi ticker
#multi_stock_history = yf.download(ticker, start=start, end=end, progress=False)
#multi_stock_history.isnull().sum(axis=1)
# for single ticker
stock_history = yf.download(ticker, start=start, end=end, progress=False)
stock_history.isnull().sum(axis=1)
# +
# Dataframe of historical stock prices
#multi_stock_history.tail()
# for single ticker
stock_history["Ticker"] = ticker
stock_history.tail()
# +
# Read in Adj Close prices
#multi_stock_history_close = multi_stock_history["Adj Close"].ffill()
#multi_stock_history_close
# for single ticker
stock_history_close = stock_history["Adj Close"].ffill()
stock_history_close
# +
# Print csv files for either single or multi stock prices
#multi_stock_history_close.to_csv('../Resources/multi_stock_prices.csv')
# for single ticker
stock_history_close.to_csv('../Resources/stock_historical_prices.csv')
# -
| stock_price_historical/yahoo_historical.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load and process molecules with `rdkit`
# This notebook does the following:
# - Molecules downloaded in the [previous notebook](./ 1_Get_Molecular_libraries.ipynb) are processed using `rdkit`.
#
# The output of this notebook is a the file `rdKit_db_molecules.obj`, which is a pandas data frame containing the rdkit object of each molecule.
import pandas as pd
import numpy as np
from glob import glob
from pathlib import Path
from rdkit import Chem
from rdkit.Chem import AllChem
from rdkit import RDLogger
RDLogger.DisableLog('rdApp.*')
import sys
sys.path.append('../..')
from helper_modules.analyze_molecules_with_rdkit import *
# ## DEKOIS Molecules
# +
sdf_input_path = './datasets/DEKOIS2/sdf/'
list_sdf_files = get_files_list(sdf_input_path, actives_name='ligand')
df_DEKOIS = get_mol_dataframe(load_molecules_from_dir(list_sdf_files))
df_DEKOIS.head()
# -
df_DEKOIS.loc['ligand_4', 'mol_rdk']
# ## DUD 2006
# +
sdf_input_path = './datasets/DUD/sdf/'
list_sdf_files = get_files_list(sdf_input_path, actives_name='ligand')
df_DUD = get_mol_dataframe(load_molecules_from_dir(list_sdf_files))
df_DUD.head()
# -
df_DUD.loc['ligand_4', 'mol_rdk']
# ## Cocrystalized molecules
# +
# Path to SDF Files
sdf_input_path = './datasets/COCRYS/sdf/'
# Function to sort the molecules by their PDB id
sort_function = lambda x: x.split('/')[-1].split('_')[0]
# Get the list of files
list_sdf_files = get_files_list(sdf_input_path,
actives_name='LIG',
sufix='',
sort_func = sort_function
)
# Compute a dataframe with the molecules as rdkit objects
df_pdi_lig = load_cocrys_molecules_from_dir(list_sdf_files)
# Update the dataframe
df_COCRYS = df_pdi_lig[['Lig', 'mol_rdk']]
df_COCRYS['Activity'] = 'active'
df_COCRYS['sanitized'] = [True if i != 'v3' else False for i in df_pdi_lig.validation]
df_COCRYS = df_COCRYS[['Lig', 'Activity', 'mol_rdk', 'sanitized']]
df_COCRYS = df_COCRYS.drop_duplicates('Lig').set_index('Lig')
df_COCRYS.sanitized.value_counts()
print('Shape', df_COCRYS.shape)
df_COCRYS.head()
# + tags=[]
df_COCRYS.loc['AQ4', 'mol_rdk']
# -
# ## Merge all dataframes
# +
list_dfs = [df_COCRYS, df_DEKOIS, df_DUD]
list_dfs_names = ['COCRYS', 'DEKOIS2', 'DUD']
# Create the final dataframe
df_all_libraries = pd.concat(list_dfs, keys = list_dfs_names)
df_all_libraries['Activity'] = df_all_libraries['Activity']\
.replace({'active': 1, 'inactive': 0})
df_all_libraries
# -
# Save the dataframe
output_file = './rdKit_db_molecules.obj'
if not Path(output_file).exists():
df_all_libraries.to_pickle(output_file)
| egfr/2_Molecular_libraries/2_Loading_molecules_from_db_with_rdkit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.011446, "end_time": "2022-02-05T16:28:23.517894", "exception": false, "start_time": "2022-02-05T16:28:23.506448", "status": "completed"} tags=[]
# #### What are you trying to do in this notebook?
# In this competition we will be ranking comments in order of severity of toxicity. We are given a list of comments, and each comment should be scored according to their relative toxicity. Comments with a higher degree of toxicity should receive a higher numerical value compared to comments with a lower degree of toxicity. In order to avoid leaks, the same text needs to be put into same Folds. For a single document this is easy, but for a pair of documents to both be in same folds is a bit tricky. This simple notebook tracks pairs of text recursively to group them and try to create a leak-free Fold split.
#
# #### Why are you trying it?
# The focus in this competition is on ranking the severity of comment toxicity from innocuous to outrageous.
#
# In Jigsaw's fourth Kaggle competition, we return to the Wikipedia Talk page comments featured in our first Kaggle competition. When we ask human judges to look at individual comments, without any context, to decide which ones are toxic and which ones are innocuous, it is rarely an easy task. In addition, each individual may have their own bar for toxicity. We've tried to work around this by aggregating the decisions with a majority vote. But many researchers have rightly pointed out that this discards meaningful information.
# + papermill={"duration": 0.066705, "end_time": "2022-02-05T16:28:23.596933", "exception": false, "start_time": "2022-02-05T16:28:23.530228", "status": "completed"} tags=[]
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + papermill={"duration": 0.94784, "end_time": "2022-02-05T16:28:24.555732", "exception": false, "start_time": "2022-02-05T16:28:23.607892", "status": "completed"} tags=[]
import pandas as pd
import numpy as np
import sklearn as sk
import matplotlib.pyplot as plt
import seaborn as sb
# + papermill={"duration": 66.124349, "end_time": "2022-02-05T16:29:30.690971", "exception": false, "start_time": "2022-02-05T16:28:24.566622", "status": "completed"} tags=[]
import pandas as pd
import numpy as np
from sklearn.linear_model import Ridge
from sklearn.feature_extraction.text import TfidfVectorizer
from scipy.stats import rankdata
jr = pd.read_csv("../input/jigsaw-regression-based-data/train_data_version2.csv")
jr.shape
df = jr[['text', 'y']]
vec = TfidfVectorizer(analyzer='char_wb', max_df=0.8, min_df=1, ngram_range=(2, 5) )
X = vec.fit_transform(df['text'])
z = df["y"].values
y=np.around ( z ,decimals = 2)
model1=Ridge(alpha=0.5)
model1.fit(X, y)
df_test = pd.read_csv("../input/jigsaw-toxic-severity-rating/comments_to_score.csv")
test=vec.transform(df_test['text'])
jr_preds=model1.predict(test)
df_test['score1']=rankdata( jr_preds, method='ordinal')
rud_df = pd.read_csv("../input/ruddit-jigsaw-dataset/Dataset/ruddit_with_text.csv")
rud_df['y'] = rud_df["offensiveness_score"]
df = rud_df[['txt', 'y']].rename(columns={'txt': 'text'})
vec = TfidfVectorizer(analyzer='char_wb', max_df=0.7, min_df=3, ngram_range=(3, 4) )
X = vec.fit_transform(df['text'])
z = df["y"].values
y=np.around ( z ,decimals = 1)
model1=Ridge(alpha=0.5)
model1.fit(X, y)
test=vec.transform(df_test['text'])
rud_preds=model1.predict(test)
df_test['score2']=rankdata( rud_preds, method='ordinal')
df_test['score']=df_test['score1']+df_test['score2']
df_test['score']=rankdata( df_test['score'], method='ordinal')
df_test[['comment_id', 'score']].to_csv("submission1.csv", index=False)
# + papermill={"duration": 41.464847, "end_time": "2022-02-05T16:30:12.166792", "exception": false, "start_time": "2022-02-05T16:29:30.701945", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
import nltk
import re
from bs4 import BeautifulSoup
from tqdm.auto import tqdm
TRAIN_DATA_PATH = "/kaggle/input/jigsaw-toxic-comment-classification-challenge/train.csv"
VALID_DATA_PATH = "/kaggle/input/jigsaw-toxic-severity-rating/validation_data.csv"
TEST_DATA_PATH = "/kaggle/input/jigsaw-toxic-severity-rating/comments_to_score.csv"
df_train2 = pd.read_csv(TRAIN_DATA_PATH)
df_valid2 = pd.read_csv(VALID_DATA_PATH)
df_test2 = pd.read_csv(TEST_DATA_PATH)
cat_mtpl = {'obscene': 0.16, 'toxic': 0.32, 'threat': 1.5,
'insult': 0.64, 'severe_toxic': 1.5, 'identity_hate': 1.5}
for category in cat_mtpl:
df_train2[category] = df_train2[category] * cat_mtpl[category]
df_train2['score'] = df_train2.loc[:, 'toxic':'identity_hate'].mean(axis=1)
df_train2['y'] = df_train2['score']
min_len = (df_train2['y'] > 0).sum()
df_y0_undersample = df_train2[df_train2['y'] == 0].sample(n=min_len, random_state=41)
df_train_new = pd.concat([df_train2[df_train2['y'] > 0], df_y0_undersample])
from tokenizers import (
decoders,
models,
normalizers,
pre_tokenizers,
processors,
trainers,
Tokenizer,
)
raw_tokenizer = Tokenizer(models.WordPiece(unk_token="[UNK]"))
raw_tokenizer.normalizer = normalizers.BertNormalizer(lowercase=True)
raw_tokenizer.pre_tokenizer = pre_tokenizers.BertPreTokenizer()
special_tokens = ["[UNK]", "[PAD]", "[CLS]", "[SEP]", "[MASK]"]
trainer = trainers.WordPieceTrainer(vocab_size=25000, special_tokens=special_tokens)
from datasets import Dataset
dataset = Dataset.from_pandas(df_train_new[['comment_text']])
def get_training_corpus():
for i in range(0, len(dataset), 1000):
yield dataset[i : i + 1000]["comment_text"]
raw_tokenizer.train_from_iterator(get_training_corpus(), trainer=trainer)
from transformers import PreTrainedTokenizerFast
tokenizer = PreTrainedTokenizerFast(
tokenizer_object=raw_tokenizer,
unk_token="[UNK]",
pad_token="[PAD]",
cls_token="[CLS]",
sep_token="[SEP]",
mask_token="[MASK]",
)
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import Ridge
def dummy_fun(doc):
return doc
labels = df_train_new['y']
comments = df_train_new['comment_text']
tokenized_comments = tokenizer(comments.to_list())['input_ids']
vectorizer = TfidfVectorizer(
analyzer = 'word',
tokenizer = dummy_fun,
preprocessor = dummy_fun,
token_pattern = None)
comments_tr = vectorizer.fit_transform(tokenized_comments)
regressor = Ridge(random_state=42, alpha=0.8)
regressor.fit(comments_tr, labels)
less_toxic_comments = df_valid2['less_toxic']
more_toxic_comments = df_valid2['more_toxic']
less_toxic_comments = tokenizer(less_toxic_comments.to_list())['input_ids']
more_toxic_comments = tokenizer(more_toxic_comments.to_list())['input_ids']
less_toxic = vectorizer.transform(less_toxic_comments)
more_toxic = vectorizer.transform(more_toxic_comments)
y_pred_less = regressor.predict(less_toxic)
y_pred_more = regressor.predict(more_toxic)
print(f'val : {(y_pred_less < y_pred_more).mean()}')
texts = df_test2['text']
texts = tokenizer(texts.to_list())['input_ids']
texts = vectorizer.transform(texts)
df_test2['prediction'] = regressor.predict(texts)
df_test2 = df_test2[['comment_id','prediction']]
df_test2['score'] = df_test2['prediction']
df_test2 = df_test2[['comment_id','score']]
df_test2.to_csv('./submission2.csv', index=False)
# + papermill={"duration": 181.716675, "end_time": "2022-02-05T16:33:13.895347", "exception": false, "start_time": "2022-02-05T16:30:12.178672", "status": "completed"} tags=[]
import pandas as pd
import numpy as np
from tqdm.auto import tqdm
from bs4 import BeautifulSoup
from collections import defaultdict
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
import re
import scipy
from scipy import sparse
from IPython.display import display
from pprint import pprint
from matplotlib import pyplot as plt
import time
import scipy.optimize as optimize
import warnings
warnings.filterwarnings("ignore")
pd.options.display.max_colwidth=300
pd.options.display.max_columns = 100
from sklearn.model_selection import train_test_split
from nltk.tokenize import word_tokenize
from sklearn.linear_model import Ridge, Lasso, BayesianRidge
from sklearn.svm import SVR
df_train = pd.read_csv("../input/jigsaw-toxic-comment-classification-challenge/train.csv")
df_sub = pd.read_csv("../input/jigsaw-toxic-severity-rating/comments_to_score.csv")
cat_mtpl = {'obscene': 0.16, 'toxic': 0.32, 'threat': 1.5,
'insult': 0.64, 'severe_toxic': 1.5, 'identity_hate': 1.5}
for category in cat_mtpl:
df_train[category] = df_train[category] * cat_mtpl[category]
df_train['score'] = df_train.loc[:, 'toxic':'identity_hate'].sum(axis=1)
df_train['y'] = df_train['score']
min_len = (df_train['y'] > 0).sum()
df_y0_undersample = df_train[df_train['y'] == 0].sample(n=min_len, random_state=201)
df_train_new = pd.concat([df_train[df_train['y'] > 0], df_y0_undersample])
df_train = df_train.rename(columns={'comment_text':'text'})
def text_cleaning(text):
'''
Cleans text into a basic form for NLP. Operations include the following:-
1. Remove special charecters like &, #, etc
2. Removes extra spaces
3. Removes embedded URL links
4. Removes HTML tags
5. Removes emojis
text - Text piece to be cleaned.
'''
template = re.compile(r'https?://\S+|www\.\S+')
text = template.sub(r'', text)
soup = BeautifulSoup(text, 'lxml')
only_text = soup.get_text()
text = only_text
emoji_pattern = re.compile("["
u"\U0001F600-\U0001F64F"
u"\U0001F300-\U0001F5FF"
u"\U0001F680-\U0001F6FF"
u"\U0001F1E0-\U0001F1FF"
u"\U00002702-\U000027B0"
u"\U000024C2-\U0001F251"
"]+", flags=re.UNICODE)
text = emoji_pattern.sub(r'', text)
text = re.sub(r"[^a-zA-Z\d]", " ", text)
text = re.sub(' +', ' ', text)
text = text.strip()
return text
tqdm.pandas()
df_train['text'] = df_train['text'].progress_apply(text_cleaning)
df = df_train.copy()
df['y'].value_counts(normalize=True)
min_len = (df['y'] >= 0.1).sum()
df_y0_undersample = df[df['y'] == 0].sample(n=min_len * 2, random_state=42)
df = pd.concat([df[df['y'] >= 0.1], df_y0_undersample])
vec = TfidfVectorizer(min_df= 3, max_df=0.8, analyzer = 'char_wb', ngram_range = (3,5))
X = vec.fit_transform(df['text'])
model = Ridge(alpha=0.5)
model.fit(X, df['y'])
l_model = Ridge(alpha=1.)
l_model.fit(X, df['y'])
s_model = Ridge(alpha=2.)
s_model.fit(X, df['y'])
df_val = pd.read_csv("../input/jigsaw-toxic-severity-rating/validation_data.csv")
tqdm.pandas()
df_val['less_toxic'] = df_val['less_toxic'].progress_apply(text_cleaning)
df_val['more_toxic'] = df_val['more_toxic'].progress_apply(text_cleaning)
X_less_toxic = vec.transform(df_val['less_toxic'])
X_more_toxic = vec.transform(df_val['more_toxic'])
p1 = model.predict(X_less_toxic)
p2 = model.predict(X_more_toxic)
# Validation Accuracy
print(f'val : {(p1 < p2).mean()}')
df_sub = pd.read_csv("../input/jigsaw-toxic-severity-rating/comments_to_score.csv")
tqdm.pandas()
df_sub['text'] = df_sub['text'].progress_apply(text_cleaning)
X_test = vec.transform(df_sub['text'])
p3 = model.predict(X_test)
p4 = l_model.predict(X_test)
p5 = s_model.predict(X_test)
df_sub['score'] = (p3 + p4 + p5) / 3.
df_sub['score'] = df_sub['score']
df_sub[['comment_id', 'score']].to_csv("submission3.csv", index=False)
# + papermill={"duration": 74.818758, "end_time": "2022-02-05T16:34:28.728182", "exception": false, "start_time": "2022-02-05T16:33:13.909424", "status": "completed"} tags=[]
from sklearn.linear_model import LinearRegression
from sklearn.feature_extraction.text import TfidfVectorizer
test_df = pd.read_csv("/kaggle/input/jigsaw-toxic-severity-rating/comments_to_score.csv")
valid_df = pd.read_csv("/kaggle/input/jigsaw-toxic-severity-rating/validation_data.csv")
train_df=pd.read_csv("../input/ruddit-jigsaw-dataset/Dataset/ruddit_with_text.csv")
train = train_df[["txt", "offensiveness_score"]]
tfvec = TfidfVectorizer(analyzer = 'char_wb', ngram_range = (3,5))
tfv = tfvec.fit_transform(train["txt"])
X=tfv
Y=train['offensiveness_score']
reg = LinearRegression().fit(X,Y)
print(reg.score(X,Y))
tfv_comments = tfvec.transform(test_df["text"])
pred1 = reg.predict(tfv_comments)
data2 = pd.read_csv("../input/jigsaw-regression-based-data/train_data_version2.csv")
df2 = data2[['text', 'y']]
vec = TfidfVectorizer(analyzer='char_wb', ngram_range=(2, 5))
X = vec.fit_transform(df2['text'])
w = df2["y"].values
y = np.around (w ,decimals = 2)
from sklearn.linear_model import Ridge
reg2=Ridge(alpha=0.3)
reg2.fit(X, y)
reg2.score(X,y)
test=vec.transform(test_df['text'])
pred2=reg2.predict(test)
sub = pd.DataFrame()
sub["comment_id"] = test_df["comment_id"]
sub["score"] = pred1 + pred2
sub.to_csv('submission4.csv',index=False)
# + papermill={"duration": 0.049999, "end_time": "2022-02-05T16:34:28.794391", "exception": false, "start_time": "2022-02-05T16:34:28.744392", "status": "completed"} tags=[]
data = pd.read_csv("./submission1.csv",index_col="comment_id")
data["score1"] = data["score"]
data["score2"] = pd.read_csv("./submission2.csv",index_col="comment_id")["score"]
data["score2"] = rankdata( data["score2"], method='ordinal')
data["score3"] = pd.read_csv("./submission3.csv",index_col="comment_id")["score"]
data["score3"] = rankdata( data["score3"], method='ordinal')
data["score4"] = pd.read_csv("./submission4.csv",index_col="comment_id")["score"]
data["score4"] = rankdata( data["score4"], method='ordinal')
# + papermill={"duration": 1.955948, "end_time": "2022-02-05T16:34:30.764100", "exception": false, "start_time": "2022-02-05T16:34:28.808152", "status": "completed"} tags=[]
for f in ['score1','score2','score3','score4']:
for i in range(0, 500):
data[f].iloc[i] = data[f].iloc[i] * 1.35
for i in range(801, 1300):
data[f].iloc[i] = data[f].iloc[i] * 1.45
for i in range(1601, 2200):
data[f].iloc[i] = data[f].iloc[i] * 0.81
for i in range(2501, 2980):
data[f].iloc[i] = data[f].iloc[i] * 0.85
for i in range(3001, 4000):
data[f].iloc[i] = data[f].iloc[i] * 1.42
for i in range(4001, 4500):
data[f].iloc[i] = data[f].iloc[i] * 1.45
for i in range(4501, 4940):
data[f].iloc[i] = data[f].iloc[i] * 0.86
for i in range(5501, 5980):
data[f].iloc[i] = data[f].iloc[i] * 0.83
for i in range(6201, 6700):
data[f].iloc[i] = data[f].iloc[i] * 1.45
for i in range(7001, 7536):
data[f].iloc[i] = data[f].iloc[i] * 1.42
# + papermill={"duration": 0.036157, "end_time": "2022-02-05T16:34:30.814756", "exception": false, "start_time": "2022-02-05T16:34:30.778599", "status": "completed"} tags=[]
data["score"] = .88*data["score1"] + .88*data["score2"] + data["score4"]*0.88
data["score"] = rankdata( data["score"], method='ordinal')
data.head()
# + papermill={"duration": 0.516584, "end_time": "2022-02-05T16:34:31.345653", "exception": false, "start_time": "2022-02-05T16:34:30.829069", "status": "completed"} tags=[]
df_test = data
for i in range(0, 500):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 1.47
for i in range(801, 1300):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 1.45
for i in range(1601, 2200):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 0.85
for i in range(2501, 2980):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 0.83
for i in range(3001, 4000):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 1.42
for i in range(4001, 4500):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 1.45
for i in range(4501, 4940):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 0.86
for i in range(5501, 5980):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 0.83
for i in range(6201, 6700):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 1.45
for i in range(7001, 7536):
df_test['score'].iloc[i] = df_test['score'].iloc[i] * 1.45
# + papermill={"duration": 0.040315, "end_time": "2022-02-05T16:34:31.400344", "exception": false, "start_time": "2022-02-05T16:34:31.360029", "status": "completed"} tags=[]
df_test["score"] = rankdata( df_test["score"], method='ordinal')
df_test["score"].to_csv('./submission.csv')
# + papermill={"duration": 0.029684, "end_time": "2022-02-05T16:34:31.444591", "exception": false, "start_time": "2022-02-05T16:34:31.414907", "status": "completed"} tags=[]
pd.read_csv("./submission.csv")
# + [markdown] papermill={"duration": 0.014477, "end_time": "2022-02-05T16:34:31.473499", "exception": false, "start_time": "2022-02-05T16:34:31.459022", "status": "completed"} tags=[]
# #### Did it work?
# There is no training data for this competition. We can refer to previous Jigsaw competitions for data that might be useful to train models. But note that the task of previous competitions has been to predict the probability that a comment was toxic, rather than the degree or severity of a comment's toxicity.
#
# #### What did you not understand about this process?
# Well, everything provides in the competition data page. I've no problem while working on it. If you guys don't understand the thing that I'll do in this notebook then please comment on this notebook.
#
# #### What else do you think you can try as part of this approach?
# While we don't include training data, we do provide a set of paired toxicity rankings that can be used to validate models.
| toxicity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_pytorch_p38)
# language: python
# name: conda_pytorch_p38
# ---
# +
import os
from os import listdir
import time
# Torch libs
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from torch.utils.data.sampler import WeightedRandomSampler
import torchvision.transforms as T
from torchvision.utils import make_grid
from torchvision.models import resnet50
import torchvision.transforms as transforms
from torchvision.io import read_image
from sklearn.model_selection import train_test_split
# Data libs
import pandas as pd
import numpy as np
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## EDA:
# Check if the image has a dog or not<br>
# If yes, count number of dogs/faces
# ### Read a doggo
# !pwd
# #### Chihuahua
doggo_1 = cv2.imread('/home/ec2-user/doggo_vision/dogs/Images/n02085620-Chihuahua/n02085620_199.jpg')
plt.imshow(doggo_1)
# #### Beagle
doggo_2 = cv2.imread('/home/ec2-user/doggo_vision/dogs/Images/n02088364-beagle/n02088364_2019.jpg')
plt.imshow(doggo_2)
# ## Cascade Classifier
# (something new?)
# ### 1. Haar Cascade
# Haar cascade function is trained from a lot of positive (faces) and negative (non-faces) human images
#
# https://docs.opencv.org/3.4/db/d28/tutorial_cascade_classifier.html
#
# Download XML file from this GitHub repo:<br>
# https://raw.githubusercontent.com/opencv/opencv/master/data/haarcascades/haarcascade_frontalface_default.xml
face_cascade = cv2.CascadeClassifier('haarcascade_frontalface_default.xml')
# #### 2. Count # of faces
faces = face_cascade.detectMultiScale(doggo_1, 1.1, 4)
print("# of faces in doggo 1: ", len(faces))
# ### Object detection ("General" rule)
#
# **No occurence:** If there is no face in the image, then the object is not present in the image
#
# **> 1 occurence**: If there are more than one occurences of the object, we process each individual using a looping construct
if(len(faces)!=0):
for (x, y, w, h) in faces:
cv2.rectangle(doggo_1, (x, y), (x+w, y+h), (255, 0, 0), 2)
plt.imshow(doggo_1)
# **NOTE:** The dataset has images that has a human face(s) along with dog face(s)
# #### Detect images with dog + human faces
def detect_hooman(path):
img = cv2.imread(path)
# Convert to Grayscale
gray_img = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
hooman_faces = face_cascade.detectMultiScale(gray_img)
if(len(hooman_faces) > 0):
return True
return False
# ### 2. Doggo Cascade
# This cascade function is trained from a lot of positive (faces) and negative (non-faces) human images
#
# https://github.com/kskd1804/dog_face_haar_cascade
doggo_cascade = cv2.CascadeClassifier('dog_cascade.xml')
# ^this is still a work in progress lol <br>
# hopefully, someone creates a working XML doggo image to make it ezpz
# ## Neural Nets
# (the OG way)
# ### Annotations
# (what's this folder doing here? :P)
# Annotation file is typically a XML file which looks quite similar to how HTML tags would look and what this file does is that it stores the metadata for respective images. <br>
#
# The advantage of using the annotations file is that we don't need to manually count the # of images or calculate image dimensions and stuff
# **Sample annotations file**
# +
import xml.etree.ElementTree as ET
from pathlib import Path
# Chihuahua
path = '/home/ec2-user/doggo_vision/dogs/Annotation/n02085620-Chihuahua/n02085620_10074'
with open(path) as annot_file:
print(''.join(annot_file.readlines()))
# +
import xml.etree.ElementTree as ET
tree = ET.parse('/home/ec2-user/doggo_vision/dogs/Annotation/n02085620-Chihuahua/n02085620_10074')
root = tree.getroot()
# +
img_info = []
for node in root.iter('object'):
name = node.find('name').text
pose = node.find('pose').text
print("Name: ", name)
print("Pose: ", pose)
for node in root.iter('size'):
width = int(node.find('width').text)
height = int(node.find('height').text)
depth = int(node.find('depth').text)
print("Width: ", width)
print("Height: ", height)
print("Depth: ", depth)
# -
# **What's the info in `<bndbox>`?**
# +
sample_annotations = []
for node in root.iter('bndbox'):
xmin = int(node.find('xmin').text)
ymin = int(node.find('ymin').text)
xmax = int(node.find('xmax').text)
ymax = int(node.find('ymax').text)
sample_annotations.append([xmin, ymin, xmax, ymax])
print(sample_annotations)
# +
from PIL import Image, ImageDraw
sample_image = Image.open('/home/ec2-user/doggo_vision/dogs/Images/n02085620-Chihuahua/n02085620_10074.jpg')
sample_image
# -
# **Tadaaaa!** annotations give us the detected object directly! <br>
# This is actally pre-computed info which helps us skip the step of **Object Detection** while performing EDA
# +
sample_image_annotated = sample_image.copy()
img_bbox = ImageDraw.Draw(sample_image_annotated)
for bbox in sample_annotations:
print(bbox)
img_bbox.rectangle(bbox, outline="green")
sample_image_annotated
# -
# #### Create labels for each dog using the Annotations file
def get_doggo_info(file_path):
for node in root.iter('object'):
name = node.find('name').text
pose = node.find('pose').text
for node in root.iter('size'):
width = int(node.find('width').text)
height = int(node.find('height').text)
depth = int(node.find('depth').text)
for node in root.iter('bndbox'):
xmin = int(node.find('xmin').text)
ymin = int(node.find('ymin').text)
xmax = int(node.find('xmax').text)
ymax = int(node.find('ymax').text)
# ### 1. Using pre-trained image using `ImageNet` dataset
# #### Check if CUDA is available to use
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
device
# #### Create `labels` from Felix's code
# +
os.chdir('/home/ec2-user/doggo_vision/dogs/Images')
labels = ['Chihuahua']
image_path = []
label_num = []
label_count = 0
for root, dirs, files in os.walk("."):
for name in files:
# print(os.path.join(root, name))
# print(root.split("-",1)[1])
labels.append(root.split("-",1)[1])
image_path.append(os.path.join(root, name))
if labels[-1] != labels[-2]:
label_count += 1
label_num.append(label_count)
df = pd.DataFrame({'labels':labels[1:],'image_path':image_path, 'label_num':label_num})
# Create label dict for later use
breeds = pd.Series(df.labels.values,index=df.label_num).to_dict()
# saving the dataframe
df.to_csv('/home/ec2-user/doggo_vision/experiments/chins/labels.csv')
display(df.head())
display(df.tail())
# +
# Test annotations
doggo_labels = pd.read_csv("/home/ec2-user/doggo_vision/experiments/chins/labels.csv")
index = 400
print("Index: ", doggo_labels.iloc[index, 0], "\nLabel: ", doggo_labels.iloc[index, 1], "\nImage directory: ", doggo_labels.iloc[index, 2], "\nLabel number: ", doggo_labels.iloc[index, 3])
print("\nTotal number of samples: ", doggo_labels.shape[0], "\nDog breeds/ unique labels: ", len(pd.unique(doggo_labels['labels'])))
# print("\nBreed value counts:", doggo_labels['labels'].value_counts())
print("\nNumber images per breed \nMax: ", max(doggo_labels['labels'].value_counts()), "\nMin:", min(doggo_labels['labels'].value_counts()))
# +
# Create train, validate and test with proportional classes
from sklearn.model_selection import train_test_split
train_ratio = 0.75
validation_ratio = 0.15
test_ratio = 0.10
label_num = np.array(doggo_labels.iloc[:, 3])
# labels
dataY = label_num
# features
dataX_dummy = range(len(label_num))
# train is now 75% of the entire data set
index_train, index_test, y_train, y_test = train_test_split(dataX_dummy, dataY, test_size=1 - train_ratio, stratify=dataY)
# test is now 10% of the initial data set
# validation is now 15% of the initial data set
index_val, index_test, y_val, y_test = train_test_split(index_test, y_test, test_size=test_ratio/(test_ratio + validation_ratio), stratify=y_test)
# -
# Check indices work
img_annotations = pd.read_csv("../../experiments/chins/labels.csv").iloc[index_train]
display(img_annotations.head())
print(img_annotations.iloc[0,2])
print(img_annotations.iloc[1,2])
# #### Using same Dataset class and config as Felix
# #### Different ways to optimize
# 1. Optimal way to crop the image
# 2. Changing number of channels - Grayscale, RGB, CMYK
# 3. Normalize the tensor - either with 0s/1s or with mean/variance
# **TODO:** Some images have channels > 4 which will cause issues while using the pre-trained ResNet50 model which is trained on images with 3 channels.<br>
#
# _Ways to rectify:_<br>
# 1. Convert channel 4 to Grayscale (2)<br>
# 2. Convert Grayscale (2) to RBG (3)<br>
class DoggoDataset(Dataset):
def __init__(self, indices, transform=None):
self.img_annotations = pd.read_csv("../../experiments/chins/labels.csv").iloc[indices]
# Convert to Grayscale and crop it to 120x120
self.transform = transforms.Compose([
transforms.Resize((120,120)),
# transforms.Grayscale(),
transforms.ToTensor()
# transforms.Normalize((0, 0, 0),(1, 1, 1))
])
def __len__(self):
return len(self.img_annotations)
def __getitem__(self, idx):
img_path = self.img_annotations.iloc[idx, 2]
image = Image.open(img_path)
label = self.img_annotations.iloc[idx, 3]
if self.transform:
image = self.transform(image)
return image, label
# ### `DataLoader()`
# #### Adding an extra parameter `num_workers`
#
# `num_workers` can be decided based off of # of cores on the EC2 instance<br>
# I'm currently using **p2.xlarge** instance which has 4 CPU cores
# Doubling the `batch_size` to 128 since we're using CUDA on a GPU instance
# +
train_set = DoggoDataset(index_train)
val_set = DoggoDataset(index_val)
test_set = DoggoDataset(index_test)
# num_workers=4
train_dataloader = DataLoader(train_set, batch_size=128, num_workers=4, shuffle=True)
val_dataloader = DataLoader(val_set, batch_size=128, num_workers=4, shuffle=True)
test_dataloader = DataLoader(test_set, batch_size=128, num_workers=4, shuffle=True)
# -
# ### Custom functions
# to make stuff modular
# #### 1. Accuracy
def get_accuracy(pred, true):
# Converting pred to 0 or 1
pred = [1 if pred[i] >= 0.5 else 0 for i in range(len(pred))]
# Calculating accuracy by comparing predictions with true labels
acc = [1 if pred[i] == true[i] else 0 for i in range(len(pred))]
# Compute accuracy
acc = np.sum(acc) / len(pred)
return (acc * 100)
# #### 2. Train the model over one epoch
def train_one_epoch(train_dataloader):
epoch_loss = []
epoch_acc = []
start_time = time.time()
for images, labels in train_dataloader:
# ResNet is trained on images with only 3 channels
if(images.shape[1] == 3):
# Load images and labels to device - in our case GPU!
images = images.to(device)
# print(images.shape)
labels = labels.to(device)
# print(labels.shape)
labels = labels.reshape((labels.shape[0], 1)) # [N, 1] - to match with preds shape
labels = labels.to(torch.float32)
# Reseting Gradients
optimizer.zero_grad()
# Forward
preds = model(images)
# Calculating Loss
_loss = criterion(preds, labels)
loss = _loss.item()
epoch_loss.append(loss)
# Calculating Accuracy
acc = get_accuracy(preds, labels)
epoch_acc.append(acc)
# Backward
_loss.backward()
optimizer.step()
# Overall Epoch Results
end_time = time.time()
total_time = end_time - start_time
# Acc and Loss
epoch_loss = np.mean(epoch_loss)
epoch_acc = np.mean(epoch_acc)
# Log the results
train_logs["loss"].append(epoch_loss)
train_logs["accuracy"].append(epoch_acc)
train_logs["time"].append(total_time)
return epoch_loss, epoch_acc, total_time
# #### 3. Validate the model over one epoch
def val_one_epoch(val_dataloader, best_val_acc):
epoch_loss = []
epoch_acc = []
start_time = time.time()
for images, labels in val_dataloader:
# ResNet is trained on images with only 3 channels
if(images.shape[1] == 3):
# Load images and labels to device - again GPU!
images = images.to(device)
labels = labels.to(device)
labels = labels.reshape((labels.shape[0], 1)) # [N, 1] - to match with preds shape
labels = labels.to(torch.float32)
# Forward
preds = model(images)
# Calculating Loss
_loss = criterion(preds, labels)
loss = _loss.item()
epoch_loss.append(loss)
# Calculating Accuracy
acc = get_accuracy(preds, labels)
epoch_acc.append(acc)
# Overall Epoch Results
end_time = time.time()
total_time = end_time - start_time
# Acc and Loss
epoch_loss = np.mean(epoch_loss)
epoch_acc = np.mean(epoch_acc)
# Log the results
val_logs["loss"].append(epoch_loss)
val_logs["accuracy"].append(epoch_acc)
val_logs["time"].append(total_time)
# Save the best model
if epoch_acc > best_val_acc:
best_val_acc = epoch_acc
torch.save(model.state_dict(),"resnet50_best.pth")
return epoch_loss, epoch_acc, total_time, best_val_acc
# ## `ResNet50`
# Let's gooooooo
# +
model = resnet50(pretrained = True)
model.fc = nn.Sequential(
nn.Linear(2048, 1, bias = True),
nn.Sigmoid()
)
# -
model
# #### Model stuff
# +
# Optimizer
optimizer = torch.optim.Adam(model.parameters(), lr = 0.0001)
# Learning Rate Scheduler
lr_scheduler = torch.optim.lr_scheduler.StepLR(optimizer, step_size = 5, gamma = 0.5)
# Loss Function
criterion = nn.BCELoss()
# Logs
train_logs = {"loss" : [], "accuracy" : [], "time" : []}
val_logs = {"loss" : [], "accuracy" : [], "time" : []}
# Loading model to device
model.to(device)
# No of epochs
epochs = 10
# +
best_val_acc = 0 # this will be computed in the validation step
for epoch in range(epochs):
# Training
loss, acc, _time = train_one_epoch(train_dataloader)
print("\nTraining")
print("Epoch {}".format(epoch+1))
print("Loss : {}".format(round(loss, 4)))
print("Acc : {}".format(round(acc, 4)))
print("Time : {}".format(round(_time, 4)))
# Validation
loss, acc, _time, best_val_acc = val_one_epoch(val_dataloader, best_val_acc)
print("\nValidating")
print("Epoch {}".format(epoch+1))
print("Loss : {}".format(round(loss, 4)))
print("Acc : {}".format(round(acc, 4)))
print("Time : {}".format(round(_time, 4)))
# -
# ### TODO:
#
# 1. Dockerize the entire codebase
# 2. Use Tensorflow serving to fetch the latest trained model instead of training every time
# 3. Compare the performance of ResNet34, VGG-16 and a few more architectures with ResNet
| experiments/chins/Ollie.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Static & Transient DataFrames in PyNastran
#
# The Jupyter notebook for this demo can be found in:
# - docs\quick_start\demo\op2_pandas_multi_case.ipynb
# - https://github.com/SteveDoyle2/pyNastran/tree/master/docs/quick_start/demo/op2_pandas_multi_case.ipynb
# +
import os
import pandas as pd
import pyNastran
from pyNastran.op2.op2 import read_op2
pkg_path = pyNastran.__path__[0]
model_path = os.path.join(pkg_path, '..', 'models')
# -
# ## Solid Bending
# Let's show off ``combine=True/False``. We'll talk about the keys soon.
solid_bending_op2 = os.path.join(model_path, 'solid_bending', 'solid_bending.op2')
solid_bending = read_op2(solid_bending_op2, combine=False, debug=False)
print(solid_bending.displacements.keys())
solid_bending_op2 = os.path.join(model_path, 'solid_bending', 'solid_bending.op2')
solid_bending2 = read_op2(solid_bending_op2, combine=True, debug=False)
print(solid_bending2.displacements.keys())
# ## Single Subcase Buckling Example
# The keys cannot be "combined" despite us telling the program that it was OK.
# We'll get the following values that we need to handle.
# #### isubcase, analysis_code, sort_method, count, subtitle
# * isubcase -> the same key that you're used to accessing
# * sort_method -> 1 (SORT1), 2 (SORT2)
# * count -> the optimization count
# * subtitle -> the analysis subtitle (changes for superlements)
# * analysis code -> the "type" of solution
# ### Partial code for calculating analysis code:
#
# if trans_word == 'LOAD STEP': # nonlinear statics
# analysis_code = 10
# elif trans_word in ['TIME', 'TIME STEP']: # TODO check name
# analysis_code = 6
# elif trans_word == 'EIGENVALUE': # normal modes
# analysis_code = 2
# elif trans_word == 'FREQ': # TODO check name
# analysis_code = 5
# elif trans_word == 'FREQUENCY':
# analysis_code = 5
# elif trans_word == 'COMPLEX EIGENVALUE':
# analysis_code = 9
# else:
# raise NotImplementedError('transient_word=%r is not supported...' % trans_word)
#
#
# ### Let's look at an odd case:
#
# You can do buckling as one subcase or two subcases (makes parsing it a lot easier!).
#
# However, you **have** to do this once you start messing around with superelements or multi-step optimization.
#
# For optimization, sometimes Nastran will downselect elements and do an optimization on that and print out a subset of the elements.
# At the end, it will rerun an analysis to double check the constraints are satisfied.
# It does not always do multi-step optimization.
op2_filename = os.path.join(model_path, 'sol_101_elements', 'buckling_solid_shell_bar.op2')
model = read_op2(op2_filename, combine=True, debug=False, build_dataframe=True)
# +
stress_keys = model.cquad4_stress.keys()
print (stress_keys)
# old: subcase, analysis_code, sort_method, count, ogs, subtitle
#key0 = (1, 1, 1, 0, '')
#key1 = (1, 8, 1, 0, '')
# new: subcase, analysis_code, sort_method, count, isuperelmemnt_adaptivity_index, pval_step
key0 = (1, 1, 1, 0, 0, '', '')
key1 = (1, 8, 1, 0, 0, '', '')
# -
# Keys:
# * key0 is the "static" key
# * key1 is the "buckling" key
#
# Similarly:
# * Transient solutions can have preload
# * Frequency solutions can have loadsets (???)
# ## Moving onto the data frames
# * The static case is the initial deflection state
# * The buckling case is "transient", where the modes (called load steps or lsdvmn here) represent the "times"
#
# pyNastran reads these tables differently and handles them differently internally. They look very similar though.
# +
stress_static = model.cquad4_stress[key0].data_frame
stress_transient = model.cquad4_stress[key1].data_frame
# The final calculated factor:
# Is it a None or not?
# This defines if it's static or transient
print('stress_static.nonlinear_factor = %s' % model.cquad4_stress[key0].nonlinear_factor)
print('stress_transient.nonlinear_factor = %s' % model.cquad4_stress[key1].nonlinear_factor)
print('data_names = %s' % model.cquad4_stress[key1].data_names)
print('loadsteps = %s' % model.cquad4_stress[key1].lsdvmns)
print('eigenvalues = %s' % model.cquad4_stress[key1].eigrs)
# -
# ## Static Table
# +
# Sets default precision of real numbers for pandas output\n"
pd.set_option('precision', 2)
stress_static.head(20)
# -
# ## Transient Table
# +
# Sets default precision of real numbers for pandas output\n"
pd.set_option('precision', 3)
#import numpy as np
#np.set_printoptions(formatter={'all':lambda x: '%g'})
stress_transient.head(20)
| docs/quick_start/demo/op2_pandas_multi_case.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Вступление
#
# Всем привет! На сегодняшнем семинаре мы познакомимся с библиотекой **pytorch**. Он очень похож на numpy, с одним лишь отличием (на самом деле их больше, но сейчас мы поговорим про самое главное) -- pytorch может считать градиенты за вас. Таким образом вам не надо будет руками писать обратный проход в нейросетях.
#
# #### Семинар построен следующим образом:
#
# 0. Повторение лекции + а зачем все это?
# 1. Вспоминаем numpy и сравниваем операции в pytorch
# 2. Создаем тензоры в pytorch
# 3. Работаем с градиентами руками
# 4. Моя первая нейросеть
# #### 0. Повторение лекции + а зачем все это?
# 
# 
# 
# 
# ### 1. Вспоминаем numpy и сравниваем операции в pytorch
#
# Мы можем создавать матрицы, перемножать их, складывать, транспонировать и в целом совершать любые матричные операции
# +
import numpy as np
import torch
import torchvision
import matplotlib.pyplot as plt
import torch.nn as nn
import torch.nn.functional as F
from sklearn.datasets import load_boston
from tqdm.notebook import tqdm
# %matplotlib inline
# -
a = np.random.rand(5, 3) # создали случайную матрицу
a
print("Проверили размеры : %s\n" % (a.shape,))
print("Добавили 5 :\n%s\n" % (a + 5))
print("X*X^T :\n%s\n" % np.dot(a, a.T))
print("Среднее по колонкам :\n%s\n" % (a.mean(axis=-1)))
print("Изменили размеры :\n%s\n" % (a.reshape(3, 5).shape,))
# #### Разминка.
#
# При помощи numpy посчитайте сумму квадратов натуральных чисел от 1 до 10000.
# +
# YOUR CODE
# -
# Аналогичные операции в **pytorch** выглядят следующим образом, синтаксис отличается, но совсем немного:
x = torch.rand(5, 3)
x
print("Проверили размеры : %s\n" % (x.shape,))
print("Добавили 5 :\n%s\n" % (x + 5))
print("X*X^T (1):\n%s\n" % (torch.matmul(x, x.transpose(1, 0))))
print("X*X^T (2):\n%s\n" % (x.mm(x.t())))
print("Среднее по колонкам :\n%s\n" % (x.mean(dim=-1)))
print("Изменили размеры :\n%s\n" % (x.view([3, 5]).shape,))
print("Изменили размеры :\n%s\n" % (x.view_as(x.t()).shape,))
# Небольшой пример того, как меняются операции:
#
# * `x.reshape([1,2,8]) -> x.view(1,2,8)`
#
# * `x.sum(axis=-1) -> x.sum(dim=-1)`
#
# * `x.astype('int64') -> x.type(torch.LongTensor)`
#
# Для помощи вам есть [таблица](https://github.com/torch/torch7/wiki/Torch-for-Numpy-users), которая поможет вам найти аналог операции в numpy
#
# #### Разминка на pytorch
#
# При помощи pytorch посчитайте сумму квадратов натуральных чисел от 1 до 10000.
# +
# YOUR CODE
# -
# ### 2. Создаем тензоры в pytorch и снова изучаем базовые операции
x = torch.empty(5, 3) # пустой тензор
print(x)
x = torch.rand(5, 3) # тензор со случайными числами
print(x)
x = torch.zeros(5, 3, dtype=torch.long) # тензор с нулями и указанием типов чисел
print(x)
x = torch.tensor([5.5, 3]) # конструируем тензор из питоновского листа
print(x)
x = x.new_ones(5, 3, dtype=torch.double) # используем уже созданный тензор для создания тензора из единичек
print(x, x.size())
x = torch.randn_like(x, dtype=torch.float) # создаем матрицу с размерами как у x
print(x, x.size())
y = torch.rand(5, 3)
print(x + y) # операция сложение
z = torch.add(x, y) # очередная операция сложения
print(z)
torch.add(x, y, out=z) # и наконец последний вид
print(z)
print(x * y) # поэлементное умножение
print(x @ y.t()) # матричное умножение
print(x.mm(y.t())) # и опять матричное умножение
print(x.unsqueeze(0).shape) # добавили измерение в начало, аналог броадкастинга
print(x.unsqueeze(0).squeeze(0).shape) # убрали измерение в начале, аналог броадкастинга
# Мы также можем делать обычные срезы и переводить матрицы назад в numpy:
a = np.ones((3, 5))
x = torch.ones((3, 5))
print(np.allclose(x.numpy(), a))
print(np.allclose(x.numpy()[:, 1], a[:, 1]))
# ### 3. Работаем с градиентами руками
boston = load_boston()
plt.scatter(boston.data[:, -1], boston.target)
# В pytorch есть возможность при создании тензора указывать нужно ли считать по нему градиент или нет, с помощью параметра `requires_grad`. Когда `requires_grad=True` мы сообщаем фреймворку, о том, что мы хотим следить за всеми тензорами, которые получаются из созданного. Иными словами, у любого тензора, у которого указан данный параметр, будет доступ к цепочке операций и преобразований совершенными с ними. Если эти функции дифференцируемые, то у тензора появляется параметр `.grad`, в котором хранится значение градиента.
# 
# Если к результирующему тензору применить метод `.backward()`, то фреймворк посчитает по цепочке градиенту для всех тензоров, у которых `requires_grad=True`.
# +
w = torch.rand(1, requires_grad=True)
b = torch.rand(1, requires_grad=True)
x = torch.tensor(boston.data[:, -1] / boston.data[:, -1].max(), dtype=torch.float32)
y = torch.tensor(boston.target, dtype=torch.float32)
assert w.grad is None # только создали тензоры и в них нет градиентов
assert b.grad is None
# -
y_pred = w * x + b # и опять совершаем операции с тензорами
loss = torch.mean((y_pred - y)**2) # совершаем операции с тензорами
loss.backward() # считаем градиенты
# +
assert w.grad is not None # сделали операции и посчитали градиенты, значение должно было появится
assert b.grad is not None
print("dL/dw = \n", w.grad)
print("dL/db = \n", b.grad)
# -
# __Ремарка__. Для доступа к значениям в тензоре используйте атрибут `.data`:
w.data
# +
from IPython.display import clear_output
for i in range(100):
y_pred = w * x + b
# попробуйте сделать полиномиальную регрессию в данном предсказании и посчитать градиенты после
loss = torch.mean((y_pred - y)**2)
loss.backward()
# делаем шаг градиентного спуска с lr = .05
w.data -= # YOUR CODE
b.data -= # YOUR CODE
# обнуляем градиенты, чтобы на следующем шаге опять посчитать и не аккумулировать их
w.grad.data.zero_()
b.grad.data.zero_()
# рисуем картинки
if (i+1) % 5 == 0:
clear_output(True)
plt.scatter(x.data.numpy(), y.data.numpy())
plt.scatter(x.data.numpy(), y_pred.data.numpy(),
color='orange', linewidth=5)
plt.show()
print("loss = ", loss.data.numpy())
if loss.data.numpy() < 0.5:
print("Done!")
break
# -
# ### 4. Моя первая нейросеть
# Для того, чтобы разобраться как обучать нейросите в pytorch, нужно освоить три вещи:
#
# 1. Как формировать батчи и пихать их в сетку
# 2. Как сделать сетку
# 3. Как написать цикл обучения и отслеживать метрики
# #### Как формировать батчи и пихать их в сетку
#
# Чтобы в данном фреймворке иметь возможность итерироваться по данным и применять к ним преобразования, например, аугментации, о которых вы узнаете позже -- нужно создать свой класс унаследованный от `torch.utils.data.Dataset`.
#
# Вот пример из документации:
#
# ```
# class FaceLandmarksDataset(torch.utils.data.Dataset):
# """Face Landmarks dataset."""
#
# def __init__(self, csv_file, root_dir, transform=None):
# """
# Args:
# csv_file (string): Path to the csv file with annotations.
# root_dir (string): Directory with all the images.
# transform (callable, optional): Optional transform to be applied
# on a sample.
# """
# self.landmarks_frame = pd.read_csv(csv_file)
# self.root_dir = root_dir
# self.transform = transform
#
# def __len__(self):
# return len(self.landmarks_frame)
#
# def __getitem__(self, idx):
# if torch.is_tensor(idx):
# idx = idx.tolist()
#
# img_name = os.path.join(self.root_dir,
# self.landmarks_frame.iloc[idx, 0])
# image = io.imread(img_name)
# landmarks = self.landmarks_frame.iloc[idx, 1:]
# landmarks = np.array([landmarks])
# landmarks = landmarks.astype('float').reshape(-1, 2)
# sample = {'image': image, 'landmarks': landmarks}
#
# if self.transform:
# sample = self.transform(sample)
#
# return sample
# ```
#
# Как вы видите, у такого класса **должно быть** два метода:
#
# * `__len__` -- возвращает информацию о том, сколько объектов у нас в датасете
# * `__getitem__` -- возвращает семпл и таргет к нему
#
#
# Теперь давайте напишем такой сами, в качестве датасета сгенерируем рандомные данные.
class RandomDataset(torch.utils.data.Dataset):
"""Our random dataset"""
def __init__(self, x, y):
self.x=x
self.y=y
def __len__(self):
return len(self.x)
def __getitem__(self, idx):
return {'sample': torch.tensor(x[idx, :], dtype=torch.float), 'target': y[idx]}
x = np.random.rand(1000, 5)
y = np.random.rand(1000)
our_dataset = RandomDataset(x, y)
our_dataset.__getitem__(1)
# Для того, чтобы из данных получать батчи в pytorch используется такая сущность как даталоадер, который принимает на вход класс унаследованный от `torch.utils.data.Dataset`. Сейчас посмотрим на пример:
dataloader = torch.utils.data.DataLoader(our_dataset, batch_size=4)
# Работают с ним следующим образом:
for batch in dataloader:
batch_x = batch['sample']
batch_y = batch['target']
break
print('Sample:', batch_x)
print('Target:', batch_y)
# #### Как сделать сетку
# Для того, чтобы в high-level pytorch создавать нейросети используется модуль `nn`. Нейросеть должна быть унаследована от класса `nn.Module`. Пример как это может выглядеть:
#
# ```
# class Model(nn.Module):
# def __init__(self):
# super(Model, self).__init__()
# self.conv1 = nn.Conv2d(1, 20, 5)
# self.conv2 = nn.Conv2d(20, 20, 5)
#
# def forward(self, x):
# x = F.relu(self.conv1(x))
# return F.relu(self.conv2(x))
# ```
#
# Как мы видим на данном примере, у данного класса должно быть метод `forward`, который определяет прямой проход нейросети. Также из класса выше видно, что модуль `nn` содержит в себе реализацию большинства слоев, а модуль `nn.functional` -- функций активаций.
#
# Есть еще один способ создать нейросеть и давайте его разберем на практике:
model = nn.Sequential() # создаем пустую модель, в которую будем добавлять слои
model.add_module('l1', nn.Linear(5, 3)) # добавили слой с 5-ю нейронами на вход и 3-мя на выход
model.add_module('l2', nn.ReLU()) # добавили функцию активации
model.add_module('l3', nn.Linear(3, 1)) # добавили слой с 3-мя нейронами на вход и 5-ю на выход
y_pred = model(batch_x) # получили предсказания модели
# #### Как написать цикл обучения и отслеживать метрики
#
# Давайте теперь соберем теперь загрузку данных, создание модели и обучим на уже созданном для нас датасете MNIST
# +
mnist_train = torchvision.datasets.MNIST(
'./mnist/', train=True, download=True,
transform=torchvision.transforms.ToTensor()
) # используем готовый класс от торча для загрузки данных для тренировки
mnist_val = torchvision.datasets.MNIST(
'./mnist/', train=False, download=True,
transform=torchvision.transforms.ToTensor()
) # используем готовый класс от торча для загрузки данных для валидации
train_dataloader = torch.utils.data.DataLoader(
mnist_train, batch_size=4, shuffle=True, num_workers=1
) # так как это уже унаследованный от Dataset класс, его можно сразу пихать в даталоадер
val_dataloader = torch.utils.data.DataLoader(
mnist_val, batch_size=4, shuffle=True, num_workers=1
) # так как это уже унаследованный от Dataset класс, его можно сразу пихать в даталоадер
# -
for i in [0, 1]:
plt.subplot(1, 2, i + 1)
plt.imshow(mnist_train[i][0].squeeze(0).numpy().reshape([28, 28]))
plt.title(str(mnist_train[i][1]))
plt.show()
# +
model = nn.Sequential(
nn.Flatten(), # превращаем картинку 28х28 в вектор размером 784
nn.Linear(784, 128), # входной слой размером 784 нейронов с выходом в 128 нейронов
nn.ReLU(), # функция активации релу
nn.Linear(128, 10), # последний слой размером 128 нейронов с выходом 10 нейронов
nn.Softmax(dim=-1) # софтмакс для получения вероятностного распределения над метками класса
)
optimizer = torch.optim.SGD(model.parameters(), lr=0.05) # создаем оптимизатор и передаем туда параметры модели
# -
# Веса моделей хранятся в виде матриц и выглядят так:
[x for x in model.named_parameters()]
# _Красиво_ трекать метрики в полуавтоматическом режиме мы будем в [wandb](https://wandb.ai). Для этого регистрируемся на сайте, устанавливаем и логинимся(это того стоит):
# +
# !pip install wandb --upgrade --quiet
import wandb
wandb.login()
# -
wandb.init(project="pytorch-demo") # инициализируем проект
wandb.watch(model) # сохраняем параметры сетки в wandb + просим следить за градиентами сетки
print()
# Можно перейти по ссылке и следить за нашей моделью прямо во время обучения!
for epoch in range(11): # всего у нас будет 10 эпох (10 раз подряд пройдемся по всем батчам из трейна)
for x_train, y_train in tqdm(train_dataloader): # берем батч из трейн лоадера
y_pred = model(x_train) # делаем предсказания
loss = F.cross_entropy(y_pred, y_train) # считаем лосс
loss.backward() # считаем градиенты обратным проходом
optimizer.step() # обновляем параметры сети
optimizer.zero_grad() # обнуляем посчитанные градиенты параметров
if epoch % 2 == 0:
val_loss = [] # сюда будем складывать **средний по бачу** лосс
val_accuracy = []
with torch.no_grad(): # мы считаем качество, поэтому мы запрещаем фреймворку считать градиенты по параметрам
for x_val, y_val in tqdm(val_dataloader): # берем батч из вал лоадера
y_pred = model(x_val) # делаем предсказания
loss = F.cross_entropy(y_pred, y_val) # считаем лосс
val_loss.append(loss.numpy()) # добавляем в массив
val_accuracy.extend((torch.argmax(y_pred, dim=-1) == y_val).numpy().tolist())
# скидываем метрики на wandb и автоматом смотрим на графики
wandb.log({"mean val loss": np.mean(val_loss),
"mean val accuracy": np.mean(val_accuracy)})
# печатаем метрики
print('Epoch: {epoch}, loss: {loss}, accuracy: {accuracy}'.format(
epoch=epoch, loss=np.mean(val_loss), accuracy=np.mean(val_accuracy)
))
# ### Дополнительные материалы:
#
# * [PyTroch на простейшем примере с пояснениями](https://github.com/Kaixhin/grokking-pytorch)
# * [Хорошая книга про pytorch](https://pytorch.org/assets/deep-learning/Deep-Learning-with-PyTorch.pdf)
# * [Использование pytorch на GPU](https://pytorch.org/docs/master/notes/cuda.html)
# * [Pytorch за 60 минут](http://pytorch.org/tutorials/beginner/deep_learning_60min_blitz.html)
# * [Как устроено автоматическое дифференцирование в pytorch](http://videolectures.net/site/normal_dl/tag=1129745/deeplearning2017_johnson_automatic_differentiation_01.pdf)
| Courses/IadMl/IntroToDeepLearning/seminars/sem01/sem01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
" Import the libraries "
import os
import sys
import math
import copy
import numpy as np
import pandas as pd
from sklearn.neural_network import MLPRegressor
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
# +
" Import the scripts of SD for Explaining "
absFilePath = os.path.dirname(os.path.dirname(os.getcwd()))
newPath = os.path.join(absFilePath, 'SplitSD4X')
sys.path.append(newPath)
from fill_missing_values import *
from missing_values_table import *
from neighbors_generation import *
from patterns_extraction import *
from performances import *
from subgroups_discovery import *
from sp_lime import *
from features_importance import *
# +
" Import matplotlib and sns to display graphs and figures "
import matplotlib
from matplotlib import pyplot as plt
from matplotlib import transforms
from matplotlib.lines import Line2D
import seaborn as sns
# %matplotlib inline
# -
" Import the plot functions"
from plotFunctions.plotMSE import *
from plotFunctions.plotScore import *
# +
" Define the functions to save and load data "
import pickle
def save_obj(obj, name):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def load_obj(name):
with open(name + '.pkl', 'rb') as f:
return pickle.load(f)
# +
" LOAD THE DATA "
path = './saved_data/'
data_train = load_obj(path + 'data_train')
target_train = load_obj(path + 'target_train')
data_test = load_obj(path + 'data_test')
target_test = load_obj(path + 'target_test')
list_neigh = load_obj(path + 'list_neighbors')
" Number of instances to explain "
n = np.size(data_test,0)
" Number of neighbors "
nb_neighs = np.size(list_neigh[0],0)
# -
# ## Multi Layer Perceptron Model
# +
" using Sklearn MLP regressor as black box: "
mlp = make_pipeline(StandardScaler(),
MLPRegressor(hidden_layer_sizes=(50, 50),
tol=1e-2,
max_iter=1000,
random_state=0))
model_nt = mlp.fit(data_train, target_train)
target_pred_nt = model_nt.predict(data_test)
print("The score of the Multi-Layer-Perceptron Regressor model is ",round(model_nt.score(data_test, target_test),4))
# -
# ## Experiments
list_subgroups = load_obj(path + 'list_subgroups')
patterns = load_obj(path + 'patterns')
att_names = load_obj(path + 'att_names')
path = './data_to_plot/'
# ### Are subgroup models faithful to the black box ?
# #### 1. MSE Error
f_mse = open(path + "mse.txt","w")
# * **Global white box**
# +
sse_wb = loss_global_wb (data_test,n,list_neigh,model_nt)
mse_wb = sse_wb / (n * nb_neighs)
print('The MSE Error of the global white box approach is : ','{:.2e}'.format(mse_wb))
"Store the results in the mse file"
f_mse.write("global\n")
f_mse.write('{:.2e}'.format(mse_wb)+'\n')
f_mse.close()
# -
# * **Submodular Pick LIME (SP LIME)**
subrgoups_splime = sb_splime (data_test,list_neigh,model_nt)
sse_splime = loss_sd (subrgoups_splime,data_test,list_neigh,model_nt)
mse_splime = sse_splime / (n * nb_neighs)
print('The MSE Error of the SPLIME approach is : ','{:.2e}'.format(mse_splime))
# * **Local Models**
# +
sse_local_m = loss_local_models(n,list_neigh,model_nt)
mse_local_m = sse_local_m / (n * nb_neighs)
print('The MSE Error of the local models approach is : ','{:.2e}'.format(mse_local_m))
"Store the results in the mse file"
f_mse = open(path + "mse.txt","a")
f_mse.write("local\n")
f_mse.write('{:.2e}'.format(mse_local_m)+'\n')
f_mse.close()
# -
# * **SPLIT SD4X (Our Approach)**
# +
nb_models = [10, 20, 50, 100]
for j in nb_models :
S = list_subgroups[j-2]
sse_S = loss_sd (S,data_test,list_neigh,model_nt)
mse_S = sse_S / (n * nb_neighs)
print(f"{'The MSE Error of the SD4X approach ['+str(j)+' models]':<50}{'is : {:.2e}'.format(mse_S)}")
"Store the results in the mse file"
nb_models_max = 100
f_mse = open(path + "mse.txt","a")
f_mse.write("sd\n")
f_mse.write('{:.2e}'.format(mse_wb)+'\n')
for i in range (0,nb_models_max-1) :
S = list_subgroups[i]
sse_S = loss_sd (S,data_test,list_neigh,model_nt)
mse_S = sse_S / (n * nb_neighs)
f_mse.write('{:.2e}'.format(mse_S)+'\n')
f_mse.close()
# -
# #### *MSE Error Graph*
plotMSE('gpu_performances',path)
# #### 2. R squared
f_rsqr = open(path + "rsquared.txt","w")
# * **Global white box**
# +
rsqr_wb = rsquared_global_wb(data_test,n,list_neigh,model_nt)
print('The Rsquared of the global white box model is : ',round(rsqr_wb,4))
"Store the results in the rsquared file"
f_rsqr.write("A:")
f_rsqr.write(str(round(rsqr_wb,4))+'\n')
f_rsqr.close()
# -
# * **Submodular Pick LIME (SP LIME)**
# +
rsqr_splime = rsquared_splime(subrgoups_splime,data_test,n,list_neigh,model_nt)
print('The Rsquared given by the splime approach is : ',round(rsqr_splime,4))
"Store the results in the rsquared file"
f_rsqr = open(path + "rsquared.txt","a")
f_rsqr.write("B:")
f_rsqr.write(str(round(rsqr_splime,4))+'\n')
f_rsqr.close()
# -
# * **SPLIT SD4X (Our Approach)**
S = list_subgroups[nb_models_max-2]
print("The number of subgroups is",len(S))
# +
nb_models = [10, 20, 50, 100]
" Store the results in the rsquared file "
f_rsqr = open(path + "rsquared.txt","a")
char = 99
for j in nb_models :
S = list_subgroups[j-2]
" Generation of the white box models for each subgroup"
W = lin_models(S,data_test,list_neigh,model_nt) #W is dictionnary
rsqr_sd = rsquared_sd (S,W,n)
print(f"{'The Rsquared given by the SD4X approach ['+str(j)+' models]':<55}{'is : {}'.format(round(rsqr_sd,4))}")
f_rsqr.write(chr(char).capitalize()+':')
f_rsqr.write(str(round(rsqr_sd,4))+'\n')
char += 1
f_rsqr.close()
# -
# * **Local Models**
# +
rsqr_local_m = rsquared_local_models (n,list_neigh,model_nt)
print('The Rsquared given by the local models approach is : ',round(rsqr_local_m,4))
" Store the results in the rsquared file "
f_rsqr = open(path + "rsquared.txt","a")
f_rsqr.write("G:")
f_rsqr.write(str(round(rsqr_local_m,4))+'\n')
f_rsqr.close()
# -
# #### *R squared Graph*
print('A : Global White Box')
print('B : SPLIME')
print('C : SD4X [10 models]')
print('D : SD4X [20 models]')
print('E : SD4X [50 models]')
print('F : SD4X [100 models]')
print('G : Local white box models')
plotOneScore('gpu_performances',path)
# ### Are subgroup models different from each other ?
# #### 1. Silimarities between the models (cosinus distance)
" We generate the the white box models "
S = list_subgroups[nb_models_max-2]
W_ = lin_models_for_sim(S,data_test,list_neigh,model_nt)
" We calculate the cosinus distance between the models"
dist = similarity (W_)
" The average of the non similarity between the models"
# we define a treshold so that if the dist between two models is less than the thereshold, they are considered unsimilars
treshold = 0.5
average_non_sim = avg_non_similar (dist,treshold)
print('The average of the non similarity between the models is',round(average_non_sim,2))
# #### *Heatmap of cosinus similarity*
# +
f, ax = plt.subplots(figsize=(20, 15))
" we set the window "
window = 15
ax = sns.heatmap(dist[:window,:window],linewidths=.5,yticklabels=False,xticklabels=False,
cmap ='RdYlGn',vmin=-1, vmax=1,square=True)
plt.title("gpu")
plt.savefig('FIGURES/heatmap_gpu_performances.png')
# -
# ### Are subgroup models human interpretable?
# #### 1. Features Importance
# +
" We plot the the features importance of the First K subgroups which covers the most instances"
K_first = 10
l_best_subgroups = sort_subgroups_support(S,K_first)
" We chose the best K features "
K_best_features = 5
plot_explanations ('gpu_performances',W_,l_best_subgroups,att_names,patterns,K_best_features)
# -
# ### Comparaison with SPLIME
# #### 1. Overlapping
# +
" We plot the matrix of overlapping "
mat_overlap = matrix_overlaps(subrgoups_splime,data_test)
print("The average of overlapping in SPLIME is", round(avg_overlaps (mat_overlap),4),'\n')
print("Red == overlapping ")
print("Green == no overlapping ")
f, ax = plt.subplots(figsize=(20, 16))
ax = sns.heatmap(mat_overlap,linewidths=.5,yticklabels=False,xticklabels=False,cmap ='RdYlGn',vmin=-1, vmax=1)
# -
" The average on instances covers by more than one subgroup "
subrgoups_splime = sb_splime (data_test,list_neigh,model_nt)
print(" The average on instances covers by more than one subgroup is :",round(cover_more (subrgoups_splime,data_test,n),4))
# #### 2. Precision of SPLIME subgroups
" The average of precisions "
print("The average of precisions for SPLIME subgroups is :",round(sb_precision (subrgoups_splime,data_test,n),4))
| tabular data/regression/Benchmarks/4. gpu performances/gpu_performs_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (tensorflow)
# language: python
# name: rga
# ---
# # T81-558: Applications of Deep Neural Networks
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), School of Engineering and Applied Science, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
#
# **Module 6 Assignment: Image Processing**
#
# **Student Name: <NAME>**
# # Assignment Instructions
#
# For this assignment you will use two images:
#
# * [Dog House](https://github.com/jeffheaton/t81_558_deep_learning/raw/master/photos/hickory_home.jpg)
# * [Land Scape](https://github.com/jeffheaton/t81_558_deep_learning/raw/master/photos/landscape.jpg)
#
#
# Your code should work with any image; however, these are the two that the **submit** function is expecting. The goal is to convert both images into square-sized. In this module we saw how to convert to a square by cropping. This time we will convert to a square by adding space. If an image is [landscape orientation](https://en.wikipedia.org/wiki/Page_orientation) you will need to add space at the top and bottom. Similarly for portrait (taller than wide) you will add space at the sides. Make sure that the image is centered between the space.
#
# The following diagram illustrates this.
#
# 
#
# To calculate the color to add to the space, take the average of all RGB values. Essentially sum all the red values, green, and blue and divide by total number of pixels. Notice how the darker landscape picture above has a darker color added to the above/below space? This is due to this averaging. Make sure you convert your average RGB to integer, RGB does not have fractional values.
#
# The submit function will check to see if your height and width match my solution. If your height and width are non-square or do not match my dimensions, you likely have a problem with your assignment.
#
# The submit function also takes three pixels and tests them. Pixels 1 and 3 are the upper left and lower-right, these are the average color and should match my solution exactly. You might see a difference in pixel 2, which is in the center, if you center the image differently than I do. If you want to match my solution, make sure to round to integer after any divisions.
#
# # Assignment Submit Function
#
# You will submit the 10 programming assignments electronically. The following submit function can be used to do this. My server will perform a basic check of each assignment and let you know if it sees any basic problems.
#
# **It is unlikely that should need to modify this function.**
# +
import base64
import os
import numpy as np
import pandas as pd
import requests
# This function submits an assignment. You can submit an assignment as much as you like, only the final
# submission counts. The paramaters are as follows:
# data - Pandas dataframe output.
# key - Your student key that was emailed to you.
# no - The assignment class number, should be 1 through 1.
# source_file - The full path to your Python or IPYNB file. This must have "_class1" as part of its name.
# . The number must match your assignment number. For example "_class2" for class assignment #2.
def submit(data,key,no,source_file=None):
if source_file is None and '__file__' not in globals(): raise Exception('Must specify a filename when a Jupyter notebook.')
if source_file is None: source_file = __file__
suffix = '_class{}'.format(no)
if suffix not in source_file: raise Exception('{} must be part of the filename.'.format(suffix))
with open(source_file, "rb") as image_file:
encoded_python = base64.b64encode(image_file.read()).decode('ascii')
ext = os.path.splitext(source_file)[-1].lower()
if ext not in ['.ipynb','.py']: raise Exception("Source file is {} must be .py or .ipynb".format(ext))
r = requests.post("https://api.heatonresearch.com/assignment-submit",
headers={'x-api-key':key}, json={'csv':base64.b64encode(data.to_csv(index=False).encode('ascii')).decode("ascii"),
'assignment': no, 'ext':ext, 'py':encoded_python})
if r.status_code == 200:
print("Success: {}".format(r.text))
else: print("Failure: {}".format(r.text))
# These functions are provided to build a submission dataframe from the two images that you must
# generate for this assignment. It is unlikely that you would need to modify these.
def scan_pixel(d,img_array,img_num,pix_num,x,y):
d[f'img{img_num}-p{pix_num}-rgb0'] = [img_array[y,x,0]]
d[f'img{img_num}-p{pix_num}-rgb1'] = [img_array[y,x,1]]
d[f'img{img_num}-p{pix_num}-rgb2'] = [img_array[y,x,2]]
def scan_image(d,img_num,img):
img_array = np.asarray(img)
rows = img_array.shape[0]
cols = img_array.shape[1]
d[f'img{img_num}-height'] = [rows]
d[f'img{img_num}-width'] = [cols]
scan_pixel(d,img_array,img_num,0,0,0)
scan_pixel(d,img_array,img_num,1,int(cols/2),int(rows/2))
scan_pixel(d,img_array,img_num,2,cols-1,rows-1)
def build_submit(submit_img1, submit_img2):
d = {}
scan_image(d,1,submit_img1)
scan_image(d,2,submit_img2)
return pd.DataFrame(d)
# -
# # Assignment #6 Sample Code
#
# The following code provides a starting point for this assignment.
# +
# %matplotlib inline
import os
import pandas as pd
import io
import requests
import numpy as np
from scipy.stats import zscore
from PIL import Image, ImageFile
from matplotlib.pyplot import imshow
import requests
from io import BytesIO
import numpy as np
# This is your student key that I emailed to you at the beginnning of the semester.
key = "<KEY>" # This is an example key and will not work.
# You must also identify your source file. (modify for your local setup)
# file='/resources/t81_558_deep_learning/assignment_yourname_class6.ipynb' # IBM Data Science Workbench
# file='C:\\Users\\jeffh\\projects\\t81_558_deep_learning\\t81_558_class6_intro_python.ipynb' # Windows
file='/Users/jheaton/projects/t81_558_deep_learning/assignments/assignment_yourname_class6.ipynb' # Mac/Linux
def fill_square_image(img):
# ************* Add your solution here*********
# You should not need to modify the other code.
# The return statement should be replaced with your own.
return img
# Handle first image
url = "https://github.com/jeffheaton/t81_558_deep_learning/raw/master/photos/hickory_home.jpg"
response = requests.get(url)
img = Image.open(BytesIO(response.content))
img.load()
submit_img1 = fill_square_image(img)
display(submit_img1)
# Handle second image
url = "https://github.com/jeffheaton/t81_558_deep_learning/raw/master/photos/landscape.jpg"
response = requests.get(url)
img = Image.open(BytesIO(response.content))
img.load()
submit_img2 = fill_square_image(img)
display(submit_img2)
# -
# Submit
submit_df = build_submit(submit_img1, submit_img2)
submit(source_file=file,data=submit_df,key=key,no=6)
| assignments/assignment_yourname_class6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # zipline MeanReversion Backtest
# In the chapter 04, we introduced `zipline` to simulate the computation of alpha factors from trailing cross-sectional market, fundamental, and alternative data.
#
# Now we will exploit the alpha factors to derive and act on buy and sell signals using the custom MeanReversion factor developed in the last chapter.
# Run the following from the command line to create a `conda` environment with `zipline` and `pyfolio`:
# ```
# conda env create -f environment.yml
# ```
# This assumes you have miniconda3 installed.
# ## Imports
# +
import sys
from pathlib import Path
import pandas as pd
from pytz import UTC
from zipline import run_algorithm
from zipline.api import (attach_pipeline, date_rules, time_rules, order_target_percent,
pipeline_output, record, schedule_function, get_open_orders, calendars,
set_commission, set_slippage)
from zipline.finance import commission, slippage
from zipline.pipeline import Pipeline, CustomFactor
from zipline.pipeline.factors import Returns, AverageDollarVolume
import logbook
import matplotlib.pyplot as plt
import seaborn
from pyfolio.utils import extract_rets_pos_txn_from_zipline
# -
sns.set_style('darkgrid')
# ## Logging Setup
# setup stdout logging
zipline_logging = logbook.NestedSetup([
logbook.NullHandler(level=logbook.DEBUG),
logbook.StreamHandler(sys.stdout, level=logbook.INFO),
logbook.StreamHandler(sys.stderr, level=logbook.ERROR),
])
zipline_logging.push_application()
# ## Algo Settings
# Settings
MONTH = 21
YEAR = 12 * MONTH
N_LONGS = 200
N_SHORTS = 0
VOL_SCREEN = 1000
start = pd.Timestamp('2010-01-01', tz=UTC)
end = pd.Timestamp('2018-01-01', tz=UTC)
capital_base = 1e7
# ## Mean Reversion Factor
class MeanReversion(CustomFactor):
"""Compute ratio of latest monthly return to 12m average,
normalized by std dev of monthly returns"""
inputs = [Returns(window_length=MONTH)]
window_length = YEAR
def compute(self, today, assets, out, monthly_returns):
df = pd.DataFrame(monthly_returns)
out[:] = df.iloc[-1].sub(df.mean()).div(df.std())
# ## Create Pipeline
# The Pipeline created by the `compute_factors()` method returns a table with a long and a short column for the 25 stocks with the largest negative and positive deviations of their last monthly return from its annual average, normalized by the standard deviation. It also limited the universe to the 500 stocks with the highest average trading volume over the last 30 trading days.
def compute_factors():
"""Create factor pipeline incl. mean reversion,
filtered by 30d Dollar Volume; capture factor ranks"""
mean_reversion = MeanReversion()
dollar_volume = AverageDollarVolume(window_length=30)
return Pipeline(columns={'longs' : mean_reversion.bottom(N_LONGS),
'shorts' : mean_reversion.top(N_SHORTS),
'ranking': mean_reversion.rank(ascending=False)},
screen=dollar_volume.top(VOL_SCREEN))
# Before_trading_start() ensures the daily execution of the pipeline and the recording of the results, including the current prices.
def before_trading_start(context, data):
"""Run factor pipeline"""
context.factor_data = pipeline_output('factor_pipeline')
record(factor_data=context.factor_data.ranking)
assets = context.factor_data.index
record(prices=data.current(assets, 'price'))
# ## Set up Rebalancing
# The new rebalance() method submits trade orders to the exec_trades() method for the assets flagged for long and short positions by the pipeline with equal positive and negative weights. It also divests any current holdings that are no longer included in the factor signals:
def rebalance(context, data):
"""Compute long, short and obsolete holdings; place trade orders"""
factor_data = context.factor_data
assets = factor_data.index
longs = assets[factor_data.longs]
shorts = assets[factor_data.shorts]
divest = context.portfolio.positions.keys() - longs.union(shorts)
exec_trades(data, assets=divest, target_percent=0)
exec_trades(data, assets=longs, target_percent=1 / N_LONGS if N_LONGS else 0)
exec_trades(data, assets=shorts, target_percent=-1 / N_SHORTS if N_SHORTS else 0)
def exec_trades(data, assets, target_percent):
"""Place orders for assets using target portfolio percentage"""
for asset in assets:
if data.can_trade(asset) and not get_open_orders(asset):
order_target_percent(asset, target_percent)
# ## Initialize Backtest
# The `rebalance()` method runs according to `date_rules` and `time_rules` set by the `schedule_function()` utility at the beginning of the week, right after market_open as stipulated by the built-in US_EQUITIES calendar (see docs for details on rules).
#
# You can also specify a trade commission both in relative terms and as a minimum amount. There is also an option to define slippage, which is the cost of an adverse change in price between trade decision and execution
def initialize(context):
"""Setup: register pipeline, schedule rebalancing,
and set trading params"""
attach_pipeline(compute_factors(), 'factor_pipeline')
schedule_function(rebalance,
date_rules.week_start(),
time_rules.market_open(),
calendar=calendars.US_EQUITIES)
set_commission(us_equities=commission.PerShare(cost=0.00075, min_trade_cost=.01))
set_slippage(us_equities=slippage.VolumeShareSlippage(volume_limit=0.0025, price_impact=0.01))
# ## Run Algorithm
# The algorithm executes upon calling the run_algorithm() function and returns the backtest performance DataFrame.
backtest = run_algorithm(start=start,
end=end,
initialize=initialize,
before_trading_start=before_trading_start,
capital_base=capital_base)
# ## Extract pyfolio Inputs
# The `extract_rets_pos_txn_from_zipline` utility provided by `pyfolio` extracts the data used to compute performance metrics.
returns, positions, transactions = extract_rets_pos_txn_from_zipline(backtest)
# ## Persist Results for use with `pyfolio`
with pd.HDFStore('backtests.h5') as store:
store.put('backtest', backtest)
store.put('returns', returns)
store.put('positions', positions)
store.put('transactions', transactions)
# ## Plot Results
fig, axes= plt.subplots(nrows=2, figsize=(14,6))
returns.add(1).cumprod().sub(1).plot(ax=axes[0], title='Cumulative Returns')
transactions.groupby(transactions.dt.dt.day).txn_dollars.sum().cumsum().plot(ax=axes[1], title='Cumulative Transactions')
fig.tight_layout();
positions.info()
transactions.describe()
| Chapter05/01_trading_zipline/alpha_factor_zipline_with_trades.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
from pycqed.instrument_drivers.physical_instruments.ZurichInstruments import ZI_HDAWG8
AWG8_8003 = ZI_HDAWG8.ZI_HDAWG8('AWG8_8003', device='dev8003')
# +
# Run this to create the parameter JSON files.
# AWG8_8003.create_parameter_files()
# -
# Getting all parameters currently takes several mins, this will be improved in the future
AWG8_8003.print_readable_snapshot(update=True)
# +
# Set the outputs to max (what units?)
AWG8_8003.sigouts_0_amplitudes_0(1)
AWG8_8003.sigouts_0_amplitudes_1(1)
# +
# Configure one of the AWGs with a simple program
program_a = '''
wave w = -0.5*join(ones(128), blackman(128, 1, 0.2), -ones(128));
while (1) {
playWave(-ones(128));
playWave(ones(128));
playWave(zeros(128));
playWave(2, -ones(128));
playWave(2, ones(128));
playWave(2, zeros(128));
}'''
program_b = '''
wave w = -0.5*join(ones(128), blackman(128, 1, 0.2), -ones(128));
while (1) {
playWave(w, w);
}'''
AWG8_8003.configure_awg_from_string(program_a) # To be made a public method
#AWG8_8003.configure_awg_from_string(program_b) # To be made a public method
# -
# Start the AWG and turn on signal outputs 0 and 1
for i in range(8):
AWG8_8003.set('sigouts_{}_enables_0'.format(i), 0)
AWG8_8003.set('sigouts_{}_enables_1'.format(i), 0)
AWG8_8003.awgs_0_enable(1)
AWG8_8003.sigouts_0_on(1)
AWG8_8003.sigouts_1_on(1)
# +
AWG8_8003.raw_sigouts_0_mode(1)
AWG8_8003.raw_sigouts_1_mode(1)
# -
# The offset functionality is currently broken.
AWG8_8003._dev.setd('sigouts/0/offset', 1)
AWG8_8003._dev.geti('sigouts/0/offset')
for i in range(8):
AWG8_8003.set('sigouts_{}_on'.format(i), 1)
| examples/AWG-8 driver example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tensor Flow Object detection
# +
# !pip install object_detection
import numpy as np
import os
import six.moves.urllib as urllib
import sys
import tarfile
import tensorflow as tf
import zipfile
from distutils.version import StrictVersion
from collections import defaultdict
from io import StringIO
from matplotlib import pyplot as plt
from PIL import Image
# This is needed since the notebook is stored in the object_detection folder.
sys.path.append("..")
from object_detection.utils import ops as utils_ops
if StrictVersion(tf.__version__) < StrictVersion('1.9.0'):
raise ImportError('Please upgrade your TensorFlow installation to v1.9.* or later!')
# -
| 00-BasicSyntax/object_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # AI-LAB LESSON 3: Constraint Optimization
#
# In the third session, we will work on the Constraint Optimization Problems (COPs). A real-life problem frequently involves both hard and soft constraints, when we formalize problems that have both types of constraints we get a constraint network augmented with a global cost function over all the variables. COPs find a complete assignment for all the variables, satisfying the hard constraints and optimizing the cost function.
from bucket_elimination import BucketElimination
from bucket import Bucket
# The algorithm you will be asked to implement make use of two class **Bucket** and **BucketElimination** and require a basic knowledge of the python data strucutre **Dictionary**.
# ### Python Dictionaries
# A dictionary is a basic data structure implemented in python (in this lesson we use only the basic feature of this structure).
# Following some **hints:**
# +
dic = { "key_0":5, "key_1":8 }
print( "Dictionary:", dic )
dic["key_0"] = 0
dic["key_1"] = 1
dic["key_2"] = 2
print( "Dictionary:", dic )
print( "\nIterate over keys:" )
for key in dic.keys(): print( "\t", key, "\t", dic[key] )
print( "\nIterate over values:" )
for val in dic.values(): print( "\t", val )
# -
# #### Python Unpacking
# Python implements the operator **"*"** for the unpacking of a list of variables. This operator can be useful to pass an array of parameters to a function (from a python list), avoiding the explicit extraction of each parameter from the list. This operator could also be useful when the number of parameters is unknown (or parametric). Following some **hints:**
# +
def custom_function_1( var_1, var_2, var_3 ):
custom_sum = var_1 + var_2 + var_3
return custom_sum
def custom_function_2( var_1, var_2, var_3, var_4 ):
custom_sum = var_1 + var_2 + var_3 + var_4
return custom_sum
variable_list = [[1, 1, 1], [3, 4, 5, 7]]
print( "Explicit extraction (f1):", custom_function_1(variable_list[0][0], variable_list[0][1], variable_list[0][2]) )
print( "Python unpacking (f1):", custom_function_1(*variable_list[0]) )
print( "Python unpacking (f2):", custom_function_2(*variable_list[1]) )
# -
# ### Bucket Class
#
# The class **Bucket** implements the data structure necessary for the bucket elimination and accepts the following arguments:
# * *variable (str)* - a string that represent the variable of the bucket (literals)
# * *soft_cnst (list)* - the soft contraints, a list of lists, each list is built with the function name for the first element, followed by the intereseted variables.
# * *ineq_cnst (list)* - the hard contraints (only inequality constraints), a list of lists, each list represent the variable interested in the inequality contraints
#
# The class **Bucket** also implements the static method *plot_table (table)*, which prints the given table in a human-readable form.
# Example in the code snippet of the bucket elimination class below.
# +
bucket_a = Bucket( variable='a', soft_cnst=[], ineq_cnst=[['a', 'b']] )
bucket_b = Bucket( variable='b', soft_cnst=[], ineq_cnst=[] )
print( bucket_a )
print( bucket_b )
# -
# ### Bucket Elimination Class
# The class **BucketElimination** implements the basic methods for the bucket elimination in a tabular form and accepts the following argument:
# * *domain (str)* - the domain of all the variables for the problem, a dictionary with the variable name as key and a list of strings for the correspondin discrete domain.
#
# The following methods are also pre-implemented:
# * *add( bucket )* - method that add an object of the class bucket to the problem.
# * *bucket_processing()* - process all the buckets in the given order (following the add chain)
# * *value_propagation()* - propagate the value based on the bucket elimination procedure to obtain the global maximum of the given problem and the corresponding assignment for the variables.
# * *plot_assignment_as_graph( assignment, soft_eval )* - plot the colored graph following the assignment for the variables.
# * *get_tables()* - get method that returns the list of the generated tables
#
# The variable assignment, returned by the method *value_propagation()*, is a python dictionary where the **key** is name of the variable and the **value** is the assigned value from the given domain.
# +
domains = { 'a':['R', 'G', 'B'], 'b':['R', 'B', 'Y'] }
bucket_elimination = BucketElimination( domains )
bucket_elimination.add( bucket_a )
bucket_elimination.add( bucket_b )
print( "Print tables BEFORE the bucket processing:" )
for table in bucket_elimination.get_tables():
print()
Bucket.plot_table( table )
bucket_elimination.bucket_processing()
print( "\nPrint tables AFTER the bucket processing:" )
for table in bucket_elimination.get_tables():
print()
Bucket.plot_table( table )
assignment, global_maximum = bucket_elimination.value_propagation()
print( "\nVariable Assignment:", assignment )
print( "\nGlobal Maximum:", global_maximum )
print( "\nPlot the assignment in a graphical form:\n" )
bucket_elimination.plot_assignment_as_graph( assignment, soft_eval=[] )
# -
# ## Assignment: Bucket Elimination
#
# Your assignment is to implement (or complete) all the necessary functions for the bucket elimination algorithm. In particular you must implement the following functions:
# * **constraint_partitioning( bucket_elimination, variable_order, soft_constraints, hard_constraints )** - in this function you have to implement the logic behind the bucket elimination constraints partitioning, given all the soft constraints and the hard constraints, this function generates all the bucket (in the given order following the algorithm) and add all the bucket to the given bucket elimination class.
# * **main_bucket_elimination( problem_name, problem_definition )** - in this function you have to implement the logic behind the bucket elimination process to correctly compute the final_tables, assignment and global maximum (here you should exploit the Bucket and the BucketElimination class and methods).
# * **get_max_table_size( final_tables )** - this function must return the maximum number of elements that appear in one of the tables generated during the process, i.e. the number of elements (rows*columns) that appear in one of the tables in the entire process.
# * **evaluate_soft_constraints( assignment, soft_constraints )** - this function must returns a list with the results of the evaluation of the soft constraints given the variables assignment.
# +
def constraint_partitioning( bucket_elimination: BucketElimination, variable_order, soft_constraints, hard_constraints ):
"""
Generate the bucket with the corresponding constraints in the correct order (inverse of the given), and add all the buckets to the bucket_elimination object that represent the problem.
Parameters
----------
bucket_elimination : BucketElimination
the object of the class BucketElimination that represent the current problem (empty).
variable_order : list
the variables that appear in the problem in the given order.
soft_constraints : list
the soft contraints, a list of lists, each list is built with the function name for the first element, followed by the intereseted variables.
hard_constraints : list
the hard contraints (only inequality constraints), a list of lists, each list represent the variable interested in the inequality contraints.
Returns:
--------
bucket_elimination : BucketElimination
the object of the class BucketElimination that represents the current problem (with the bucket filled).
"""
variable_order.reverse()
for var in variable_order:
constraints = []
for cnstr in soft_constraints:
bucket_elimination.
print(var)
return bucket_elimination
main_bucket_elimination( "Problem Graph Coloring", PROBLEM_GC )
# +
def main_bucket_elimination( problem_name, problem_definition ):
"""
Main script of the bucket elimination, given the problem definition compute the global_maximum,
the correct assignment and the memory cost of the process.
Parameters
----------
problem_name : str
the name of the problem, for visualization purpose.
problem_definition : list
complete definition of the problem, a list that contain (in order):
problem_domains, variable_order, problem_soft_constraints and problem_hard_constraints.
"""
# Extract the problem constant from the parameter "problem_definition"
problem_domains, variable_order, problem_soft_constraints, problem_hard_constraints = problem_definition
assignment, global_maximum, max_table_size = None, None, None
bucket_elimination = BucketElimination(problem_domains)
constraint_partitioning(bucket_elimination, variable_order, problem_soft_constraints, problem_hard_constraints)
#
# YOUR CODE HERE ...
#
evaluations = []
# Plot all the computed results
print( f"\nBucket Elimination for the: {problem_name}:" )
print( f"\tVariable Assignment: {assignment}" )
print( f"\tGlobal Maximum Found: {global_maximum}" )
print( f"\tMaximum Table Size (with the order {variable_order}): {max_table_size}" )
print( "\tGraphical Visualization:" )
bucket_elimination.plot_assignment_as_graph( assignment, evaluations )
# -
def get_max_table_size( final_tables ):
"""
Compute the maximum number of elements that appear in one of the table generated inside the main process.
Parameters
----------
final_tables : list
list of the tables generated inside the loop for each bucket.
Returns:
--------
max_table_size : int
the number of elements inside the largest table (i.e., number of row multiplied by the number of columns).
"""
# Variable initialization
max_table_size = 0
#
# YOUR CODE HERE ...
#
return max_table_size
def evaluate_soft_constraints( assignment, soft_constraints ):
"""
Compute the value of the soft constraints, evaluating them on the given the variables assignment.
Parameters
----------
assignment : list
the assignment for each variable to obtain the maximum (the key is the literal and the value is the assigned value).
soft_constraints : list
the soft contraints, a list of lists, each list is built with the function name for the first element, followed by the intereseted variables.
Returns:
--------
evaluations : list
a list with the results of the evaluation of the soft constraints given the variables assignment.
"""
# Variable initialization
evaluations = []
#
# YOUR CODE HERE ...
#
return evaluations
# ### Problem Definitions:
#
# The following initializations provide the structure for the 3 problems of this lesson:
# +
def F_1( x_i, x_j ):
if x_i != x_j: return 0
elif x_i == 'R' and x_j == 'R': return -1
elif x_i == 'B' and x_j == 'B': return -2
else: raise ValueError("Invalid Value for F")
def F_2( x_i, x_j ):
if x_i != x_j: return 0
elif x_i == 'R' and x_j == 'R': return 2
elif x_i == 'B' and x_j == 'B': return 1
else: raise ValueError("Invalid Value for F")
PROBLEM_GC = [
{ 'X1':['R', 'B', 'Y'], 'X2':['R', 'B', 'Y'], 'X3':['R', 'B', 'Y'], 'X4':['R', 'B', 'Y'], 'X5':['R', 'B', 'Y'] }, # PROBLEM DOMAINS
['X5', 'X4', 'X3', 'X2', 'X1'], # PROBLEM ORDER
[], # PROBLEM SOFT CONSTRAINTS
[['X1', 'X2'], ['X2', 'X3'], ['X3', 'X4'], ['X2', 'X4'], ['X1', 'X4'], ['X2', 'X5'], ['X3', 'X5'], ['X1', 'X5']] # PROBLEM HARD CONSTRAINTS
]
PROBLEM_2 = [
{ 'X1':['R', 'B'], 'X2':['R', 'B'], 'X3':['R', 'B'], 'X4':['R', 'B'] }, # PROBLEM DOMAINS
['X1', 'X2', 'X3', 'X4'], # PROBLEM ORDER
[[F_2, 'X1', 'X2'], [F_2, 'X2', 'X3'], [F_2, 'X2', 'X4']], # PROBLEM SOFT CONSTRAINTS
[['X1', 'X3'], ['X3', 'X4']] # PROBLEM HARD CONSTRAINTS
]
PROBLEM_1 = [
{ 'X1':['R', 'B'], 'X2':['R', 'B'], 'X3':['R', 'B'], 'X4':['R', 'B'] }, # PROBLEM DOMAINS
['X4', 'X3', 'X2', 'X1'], # PROBLEM ORDER
[[F_1, 'X1', 'X2'], [F_1, 'X1', 'X4'], [F_1, 'X2', 'X4'], [F_1, 'X3', 'X4']], # PROBLEM SOFT CONSTRAINTS
[] # PROBLEM HARD CONSTRAINTS
]
# -
# ## Exercise: Graph Coloring
#
# The following code calls your *get_bucket_elimination_tables* and *get_bucket_elimination_assignment* to the graph coloring problem:
main_bucket_elimination( "Problem Graph Coloring", PROBLEM_GC )
# Correct results can be found [here](lesson_3_results.txt) and with the resulting graph below:
#
# <img src="images/graph_coloring.png" width="250">
#
# ### Exercise: Partial Tests 15/05/2013
#
# The following code calls your *get_bucket_elimination_tables* and *get_bucket_elimination_assignment* to compute and solve a complete bucket elimination problem and prints the results (tabular and graphical form). The problems are extracted from the partial test of *15/05/2013* and *04/05/2016*
main_bucket_elimination( "Partial Test 15/05/2013", PROBLEM_1 )
# Correct results can be found [here](lesson_3_results.txt) and with the resulting graph below:
#
# <img src="images/partial_test_a.png" width="250">
#
# ## Exercise: Partial Tests 04/05/2016
#
# The following code calls your *get_bucket_elimination_tables* and *get_bucket_elimination_assignment* to compute and solve a complete bucket elimination problem and prints the results (tabular and graphical form). The problems are extracted from the partial test of *15/05/2013* and *04/05/2016*
main_bucket_elimination( "Partial Test 04/05/2016", PROBLEM_2 )
# Correct results can be found [here](lesson_3_results.txt) and with the resulting graph below:
#
# <img src="images/partial_test_b.png" width="250">
#
# ## Analysis: Variables Order
#
# Now that you have correctly implemented the bucket elimination algorithm, what can you say about the solutions they compute? Changing the order of the variables, does the result change? And the memory cost?
| lesson_3/lesson_3_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (gpytorch)
# language: python
# name: gpytorch
# ---
# import sys
# sys.path.append('/Users/guanchun/Auto_Sci')
# print(sys.path)
import typing
import numpy as np
from typing import Union, Tuple
#from auto_sci import LotkaVolterra
from auto_sci.experiments.simulation import *
def greeting(name: str) -> str:
return 'Hello ' + name
greeting('Tom')
xs = [2, 3, 4]
print(xs)
print(type(xs))
xs.append('Tom')
print(xs)
xs.pop()
print(xs)
len(xs)
xy = np.array([5, 6, 7])
print(type(xy))
print(xy)
u = Union()
def test_var_arg(farg, *args):
print('formal argument is ' + farg)
for arg in args:
print ('other arguments are', arg )
test_var_arg('H', 'NI', 45)
def test_var_kwarg(farg, **kwargs):
print('formal arg is ', farg)
for key in kwargs:
print(('other keywords argument is %s : %s')%(key, kwargs[key]))
test_var_kwarg(farg = 'He', time = '2019', place = 'Zurich')
a = (3, 4, 5)
print(type(a))
pwd
test.__name__
# cd ..
pwd
import foo
# +
lotka_volterra_simulator = LotkaVolterra(true_param=(2.0, 1.0, 4.0, 1.0),
noise_variance=0.1**2)
system_obs, t_obs = lotka_volterra_simulator.observe(initial_state=(5.0, 3.0),
initial_time=0.0,
final_time=2.0,
t_delta_integration=0.01,
t_delta_observation=0.1)
n_states, n_points = system_obs.shape
# -
print(n_states)
print(system_obs)
pwd
| auto_sci/tests/test.ipynb |