code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 数据预处理sklearn.preprocessing 的几个模块
#
# 方法来自[这里](https://blog.csdn.net/weixin_40807247/article/details/82793220)
# +
import pandas as pd
from path import Path
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
root = Path('F:/datasets/kaggle/House')
df_train = pd.read_csv(root/'train.csv')
print(df_train.columns)
y_np = df_train['SalePrice'].values#return np(1460,) order 1 tensor
y_np_reshaped = y_np.reshape(-1,1)# 貌似所有PreProcessing都得用[n,1]的shape,order2 tensor
# -
# ## 标准化
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(y_np_reshaped)
y_scale_1 = scaler.transform(y_np_reshaped)# 可以顺便记录变换的过程(scaler)
y_inversed = scaler.inverse_transform(y_scale_1)#基本就是y_np_reshaped
fig, (ax1, ax2) = plt.subplots(2,1, figsize=(10,5))
sns.distplot(y_scale_1,ax=ax1)
sns.distplot(y_inversed,ax=ax2)
print(pd.Series(y_scale_1.squeeze(1)).describe())
print(pd.Series(y_inversed.squeeze(1)).describe())#对numpy->series
# -
# ## 常规方法标准化
# +
from sklearn.preprocessing import scale
y_scale_0 = scale(y_np_reshaped)# 最朴素的, 直接一个函数变换
y_scale_2 = StandardScaler().fit_transform(y_np_reshaped)# 效果和0号一样
y_scaled = pd.Series(y_scale_0.squeeze(1))#(1460,1)->(1460,)
#sale_price = preprocessing.StandardScaler().fit_transform(sale_price.values)
print(pd.Series(y_scaled).describe())
# +
## 正则化
from sklearn.preprocessing import normalize
y_norm = normalize(y_np_reshaped,norm='l2')
sns.distplot(y_norm)
pd.Series(y_norm.squeeze(1)).describe()
# -
# ## 独热编码
# +
from sklearn.preprocessing import OneHotEncoder
enc = OneHotEncoder()
enc.fit([[0, 0, 3],
[1, 1, 0],
[0, 2, 1],
[1, 0, 2]]) # fit来学习编码
enc.transform([[0, 0, 0]]).toarray()# 进行编码
enc.fit([['B',1,'MALE'],
['B',0,'FEMALE'],
['C',2,'FEMALE'],
['A',0,'MALE']])
# +
print(enc.transform([['A', 0, 'MALE']]).toarray())#独热编码还是从首位ascll码值开始排列的,reutrun a tuple
y1 = enc.transform([['A', 0, 'MALE']]).toarray()[:,:enc.n_values_[0]+1]# 进行编码
y2 = enc.transform([['A', 0, 'MALE']]).toarray()[:,enc.n_values_[0]:enc.n_values_[1]+1]# 进行编码
y3 = enc.transform([['A', 0, 'MALE']]).toarray()[:,enc.n_values_[1]:enc.n_values_[2]+1]# 进行编码
print(y1,y2,y3)
#print(enc.categories_)# 各个特征的变量空间
#print(enc.n_values_)# 各个特征的变量空间大小
# -
# ## 空值补齐
# +
# 5 弥补缺失数据
# 在scikit-learn的模型中都是假设输入的数据是数值型的,并且都是有意义的,如果有缺失数据是通过NAN,或者空值表示的话,就无法识别与计算了。
# 要弥补缺失值,可以使用均值,中位数,众数等等。Imputer这个类可以实现。请看:
import numpy as np
from sklearn.preprocessing import Imputer
imp = Imputer(missing_values='NaN', strategy='mean', axis=0)#通过均值补齐axis=0 的Nan值
data = [[1, 2],
[np.nan, 3],
[7, 6]]
imp.fit(data)
# -
x = [[np.nan, 2], [6, np.nan], [7, 6]]
x_ = imp.transform(x)
from sklearn.datasets import load_digits
import matplotlib.pyplot as plt
X,y = load_digits(return_X_y=True)
# +
X.shape()
#plt.gray()
#plt.matshow(X.images[0])
#plt.show()
| House/sklearn.preprocessing_using.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # With SentencePiece tokenizer
# ## Initial setup
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from fastai import *
from fastai.text import *
bs = 512
data_path = Config.data_path()
lang = 'nl'
name = f'{lang}wiki_sp' # Use a different directory.
path = data_path/name
path.mkdir(exist_ok=True, parents=True)
lm_fns = [f'{lang}_wt', f'{lang}_wt_vocab']
# ## Download wikipedia data
# +
# from nlputils import split_wiki,get_wiki
# +
# get_wiki(path, lang)
# +
# path.ls()
# -
# ### Split in separate files
dest = path/'docs_small'
dest.ls()[:5]
# ## Create databunch for language model
# This takes about 45 minutes:
data = (TextList.from_folder(dest, processor=[OpenFileProcessor(), SPProcessor()])
.split_by_rand_pct(0.1, seed=42)
.label_for_lm()
.databunch(bs=bs, num_workers=1, bptt=70))
data.save(f'{lang}_databunch_sp') # Different databunch
len(data.vocab.itos),len(data.train_ds)
# data.train_ds[:5]
data.vocab.itos[:1000]
data.show_batch()
# ## Train language model
data = load_data(dest, f'{lang}_databunch_sp', bs=bs, num_workers=1)
# +
# data.train_ds[:1]
# -
config = dict(emb_sz=400, n_hid=1152, n_layers=3, pad_token=1, qrnn=False, bidir=False, output_p=0.1,
hidden_p=0.15, input_p=0.25, embed_p=0.02, weight_p=0.2, tie_weights=True, out_bias=True)
learn = language_model_learner(data, AWD_LSTM, config=config, drop_mult=1.0, pretrained=False)
# learn = language_model_learner(data, AWD_LSTM, config=config, drop_mult=1.5, pretrained=False)
# learn = language_model_learner(data, AWD_LSTM, drop_mult=1.0, pretrained=False)
learn.unfreeze()
learn.lr_find()
learn.recorder.plot()
# lr = 3e-3
lr = 1e-2
# lr = 2e-2
# +
# learn.fit_one_cycle(1, lr, moms=(0.8, 0.7))
# +
# Previous run (lr = 5e-3)
# learn.fit_one_cycle(5, lr, moms=(0.8, 0.7))
# -
learn.fit_one_cycle(10, lr, moms=(0.8, 0.7))
# +
# learn.fit_one_cycle(1, lr, moms=(0.8, 0.7))
# -
mdl_path = path/'models'
mdl_path.mkdir(exist_ok=True)
# learn.to_fp32().save(mdl_path/lm_fns[0], with_opt=False)
learn.save(mdl_path/lm_fns[0], with_opt=False)
learn.data.vocab.save(mdl_path/(lm_fns[1] + '.pkl'))
# +
TEXT = '''Het beleg van Utrecht
Het beleg van Utrecht'''
N_WORDS = 200
N_SENTENCES = 2
print("\n".join(learn.predict(TEXT, N_WORDS, temperature=0.85) for _ in range(N_SENTENCES)))
# -
# learn = language_model_learner(data, AWD_LSTM, drop_mult=1.,
# path = path,
# pretrained_fnames=lm_fns)
learn.export()
# +
TEXT = '''<NAME>
<NAME>'''
N_WORDS = 500
N_SENTENCES = 1
print("\n".join(learn.predict(TEXT, N_WORDS, temperature=0.65) for _ in range(N_SENTENCES)))
# -
| 0-initial-language-model-with-sp-small.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import base64
import string
import re
from collections import Counter
from nltk.corpus import stopwords
stopwords = stopwords.words('english')
df = pd.read_csv('../data/train.csv')
df.head()
victorian_stopwords = ["'s", "one", "may", "seem", "yet", "could", "thus"]
stopwords.append("'s")
stopwords.append("one")
type(stopwords)
df.isnull().sum()
# The plot below shows the distribution of words among each represented author.
# +
fig = plt.figure(figsize=(8,4))
sns.set_theme(palette = 'rocket')
sns.barplot(x = df['author'].unique(), y= df['author'].value_counts())
plt.grid(color='w', linestyle='-', linewidth=1, zorder = 0)
plt.xlabel('Author')
plt.ylabel('Number of Lines')
plt.title('Comparison of Lines by Author')
plt.show()
# -
from sklearn.model_selection import train_test_split
train, test = train_test_split(df, test_size=0.33, random_state=42)
print('Text sample:', train['text'].iloc[0])
print('Author of this text:', train['author'].iloc[0])
print('Training Data Shape:', train.shape)
print('Testing Data Shape:', test.shape)
import spacy
nlp = spacy.load('en_core_web_sm')
punctuations = string.punctuation
def cleanup_text(docs, logging=True):
texts = []
counter = 1
for doc in docs:
if counter % 1000 == 0 and logging:
print("Processed %d out of %d documents." % (counter, len(docs)))
counter += 1
doc = nlp(doc, disable=['parser', 'ner'])
tokens = [tok.lemma_.lower().strip() for tok in doc if tok.lemma_ != '-PRON-']
tokens = [tok for tok in tokens if tok not in stopwords and tok not in punctuations]
tokens = ' '.join(tokens)
texts.append(tokens)
return pd.Series(texts)
EAP_text = [text for text in train[train['author'] == 'EAP']['text']]
HPL_text = [text for text in train[train['author'] == 'HPL']['text']]
MWS_text = [text for text in train[train['author'] == 'MWS']['text']]
EAP_clean = cleanup_text(EAP_text)
EAP_clean = ' '.join(EAP_clean).split()
HPL_clean = cleanup_text(HPL_text)
HPL_clean = ' '.join(HPL_clean).split()
MWS_clean = cleanup_text(MWS_text)
MWS_clean = ' '.join(MWS_clean).split()
EAP_counts = Counter(EAP_clean)
HPL_counts = Counter(HPL_clean)
MWS_counts = Counter(MWS_clean)
EAP_common_words = [word[0] for word in EAP_counts.most_common(20)]
EAP_common_counts = [word[1] for word in EAP_counts.most_common(20)]
# +
fig = plt.figure(figsize=(18,6))
sns.barplot(x=EAP_common_words, y=EAP_common_counts, palette = 'rocket')
plt.title('Most Common Words used in short stories written by <NAME>')
plt.show()
# -
HPL_common_words = [word[0] for word in HPL_counts.most_common(20)]
HPL_common_counts = [word[1] for word in HPL_counts.most_common(20)]
fig = plt.figure(figsize=(18,6))
sns.barplot(x=HPL_common_words, y=HPL_common_counts, palette='rocket')
plt.title('Most Common Words used in the short stories of H.P. Lovecraft')
plt.show()
#plt.savefig(f'images/{author}_words.png')
MWS_common_words = [word[0] for word in MWS_counts.most_common(20)]
MWS_common_counts = [word[1] for word in MWS_counts.most_common(20)]
fig = plt.figure(figsize=(18,6))
sns.barplot(x=MWS_common_words, y=MWS_common_counts, palette = 'rocket')
plt.title('Most Common Words used in the short stories of Mary Wallstonecraft Shelley')
plt.show()
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.base import TransformerMixin
from sklearn.pipeline import Pipeline
from sklearn.svm import LinearSVC
from sklearn.feature_extraction.stop_words import ENGLISH_STOP_WORDS
from sklearn.metrics import accuracy_score
from nltk.corpus import stopwords
import string
import re
import spacy
spacy.load('en')
from spacy.lang.en import English
parser = English()
# +
STOPLIST = set(stopwords.words('english') + list(ENGLISH_STOP_WORDS))
SYMBOLS = " ".join(string.punctuation).split(" ") + ["-", "...", "”", "”"]
class CleanTextTransformer(TransformerMixin):
def transform(self, X, **transform_params):
return [cleanText(text) for text in X]
def fit(self, X, y=None, **fit_params):
return self
def get_params(self, deep=True):
return {}
def cleanText(text):
text = text.strip().replace("\n", " ").replace("\r", " ")
text = text.lower()
return text
def tokenizeText(sample):
tokens = parser(sample)
lemmas = []
for tok in tokens:
lemmas.append(tok.lemma_.lower().strip() if tok.lemma_ != "-PRON-" else tok.lower_)
tokens = lemmas
tokens = [tok for tok in tokens if tok not in STOPLIST]
tokens = [tok for tok in tokens if tok not in SYMBOLS]
return tokens
# -
from sklearn.naive_bayes import MultinomialNB
def printNMostInformative(vectorizer, clf, N):
feature_names = vectorizer.get_feature_names()
feature_with_fns = sorted(zip(clf.feature_log_prob_[0], feature_names))
topClass1 = coefs_with_fns[:N]
topClass2 = coefs_with_fns[:-(N + 1):-1]
topClass3 =
print("Class 1 best: ")
for feat in topClass1:
print(feat)
print("Class 2 best: ")
for feat in topClass2:
print(feat)
print( "Class 3 best: ")
for feat in topClass3:
print(feat)
vectorizer = CountVectorizer(tokenizer=tokenizeText, ngram_range=(1,1))
clf = MultinomialNB()
# +
pipe = Pipeline([('cleanText', CleanTextTransformer()), ('vectorizer', vectorizer), ('clf', clf)])
# data
train1 = train['text'].tolist()
labelsTrain1 = train['author'].tolist()
test1 = test['text'].tolist()
labelsTest1 = test['author'].tolist()
# train
pipe.fit(train1, labelsTrain1)
# test
preds = pipe.predict(test1)
print("accuracy:", accuracy_score(labelsTest1, preds))
print("Top 20 features used to predict: ")
printNMostInformative(vectorizer, clf, 20)
pipe = Pipeline([('cleanText', CleanTextTransformer()), ('vectorizer', vectorizer)])
transform = pipe.fit_transform(train1, labelsTrain1)
vocab = vectorizer.get_feature_names()
for i in range(len(train1)):
s = ""
indexIntoVocab = transform.indices[transform.indptr[i]:transform.indptr[i+1]]
numOccurences = transform.data[transform.indptr[i]:transform.indptr[i+1]]
for idx, num in zip(indexIntoVocab, numOccurences):
s += str((vocab[idx], num))
# -
from sklearn import metrics
print(metrics.classification_report(labelsTest1, preds,
target_names=df['author'].unique()))
| notebooks/drafts_and_sketches/spacy_training-NB version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="beeMNE6NupHF"
# - Suppose $f$ is convex and $X$ is martingale, prove that $g(t) = \mathbb E[f(X_t)]$ is increasing.
# + [markdown] colab_type="text" id="skUVOXnhvJdK"
# - __Proof__ (your proof here)
# + [markdown] colab_type="text" id="lw-yQOBaupHI"
# - Let $t \mapsto e^{-rt}S_t$ be a martingale,
# then prove that $$C(t) = \mathbb E[e^{-rt}(S_t - K)^+]$$ is increasing.
# + [markdown] colab_type="text" id="oOQi7_7fvVn2"
# - __Proof__ (your proof here)
# + [markdown] colab_type="text" id="E09uh6uDupHJ"
# - Suppose $r = 0$ and $S$ is martingale, prove that
# $P(t) = \mathbb E [(S_t - K)^-]$ is increasing.
# + [markdown] colab_type="text" id="sVD2bl4uvpBy"
# - __Proof__ (here)
# + [markdown] colab={} colab_type="code" id="F2yYY1Q8upHK"
# - Prove or disprove the following statement: Let $r>0$ and $S$ is a martingale. Then $P(t) = \mathbb E [(S_t - K)^-]$ is increasing.
# -
# - __Proof__ (here)
| src/20montone_revised.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Amazon Augmented AI (Amazon A2I)
# Amazon Augmented AI (Amazon A2I) makes it easy to build the workflows required for human review of ML predictions. Amazon A2I brings human review to all developers, removing the undifferentiated heavy lifting associated with building human review systems or managing large numbers of human reviewers.
#
# You can create your own workflows for ML models built on Amazon SageMaker or any other tools. Using Amazon A2I, you can allow human reviewers to step in when a model is unable to make a high confidence prediction or to audit its predictions on an on-going basis.
#
# Learn more here: https://aws.amazon.com/augmented-ai/
# # Integrate human reviewers in Amazon Comprehend Text Classification
# To incorporate Amazon A2I into your human review workflows, you need three resources:
#
# * **Worker Task Template** to create a Human Task UI for the worker. The worker UI displays your input data, such as documents or images, and instructions to workers. It also provides interactive tools that the worker uses to complete your tasks. For more information, see https://docs.aws.amazon.com/sagemaker/latest/dg/a2i-instructions-overview.html
#
# * **Flow Definition** to create a Human Review Workflow.You use the flow definition to configure your human workforce and provide information about how to accomplish the human review task. You can create a flow definition in the Amazon Augmented AI console or with Amazon A2I APIs. To learn more about both of these options, see https://docs.aws.amazon.com/sagemaker/latest/dg/a2i-create-flow-definition.html
#
# * **Human Loop** starts your human review workflow. When you use one of the built-in task types, the corresponding AWS service creates and starts a human loop on your behalf when the conditions specified in your flow definition are met or for each object if no conditions were specified. When a human loop is triggered, human review tasks are sent to the workers as specified in the flow definition.
#
# When using a custom task type, as this notebook will show, you start a human loop using the Amazon Augmented AI Runtime API. When you call StartHumanLoop in your custom application, a task is sent to human reviewers.
# ### Install Latest SDK
# First, let's get the latest installations of our dependencies
# !pip install -qU pip
# !pip install -qU boto3
# !pip install -qU botocore
# ## Setup
# We need to set up the following data:
# * `region` - Region to call A2I
# * `bucket` - A S3 bucket accessible by the given role
# * Used to store the sample images & output results
# * Must be within the same region A2I is called from
# * `role` - The IAM role used as part of StartHumanLoop. By default, this notebook will use the execution role
# * `workteam` - Subsets or Group of people within your workorce to send the work to
# ### Role and Permissions
#
# The AWS IAM Role used to execute the notebook needs to have the following permissions:
#
# * ComprehendFullAccess
# * SagemakerFullAccess
# * S3 Read/Write Access to the BUCKET listed above
# * AmazonSageMakerMechanicalTurkAccess (if using MechanicalTurk as your Workforce)
# #### Setup Bucket and Paths
# +
import boto3
import sagemaker
import pandas as pd
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
# -
output_path = f's3://{bucket}/a2i-results'
print(output_path)
# ### Workteam or Workforce
# A workforce is the group of workers that you have selected to label your dataset. You can choose either the Amazon Mechanical Turk workforce, a vendor-managed workforce, or you can create your own private workforce for human reviews. Whichever workforce type you choose, Amazon Augmented AI takes care of sending tasks to workers.
#
# When you use a private workforce, you also create work teams, a group of workers from your workforce that are assigned to Amazon Augmented AI human review tasks. You can have multiple work teams and can assign one or more work teams to each job.
# To create your Workteam, navigate here: https://console.aws.amazon.com/sagemaker/groundtruth#/labeling-workforces/create
#
#
# After you have created your workteam, replace YOUR_WORKTEAM_ARN below
WORKTEAM_ARN = 'arn:aws:sagemaker:us-east-1:835319576252:workteam/private-crowd/dsoaws'
# Visit: https://docs.aws.amazon.com/sagemaker/latest/dg/a2i-permissions-security.html to add the necessary permissions to your role
# ## Client Setup
# Here we are going to setup the rest of our clients.
# +
import io
import json
import uuid
import time
import boto3
import botocore
# Amazon SageMaker client
sagemaker = boto3.client('sagemaker', region)
# Amazon Comprehend client
comprehend = boto3.client('comprehend', region)
# Amazon Augment AI (A2I) client
a2i = boto3.client('sagemaker-a2i-runtime')
# Amazon S3 client
s3 = boto3.client('s3', region)
# -
# ## Create Control Plane Resources
# ### Create the Human Task UI using a Worker Task Template
#
# Create a human task UI resource, giving a UI template. This template will be rendered to the human workers whenever human interaction is required.
#
# Below we've provided a simple demo template that is compatible with AWS Comprehend's Detect Sentiment API.
#
# For other pre-built UIs (70+), check: https://github.com/aws-samples/amazon-a2i-sample-task-uis
# +
template = r"""
<script src="https://assets.crowd.aws/crowd-html-elements.js"></script>
<crowd-form>
<crowd-classifier
name="sentiment"
categories="['Positive', 'Negative', 'Neutral', 'Mixed']"
initial-value="{{ task.input.initialValue }}"
header="What sentiment does this text convey?"
>
<classification-target>
{{ task.input.taskObject }}
</classification-target>
<full-instructions header="Text Classification Instructions">
<p><strong>Positive</strong> sentiment include: joy, excitement, delight</p>
<p><strong>Negative</strong> sentiment include: anger, sarcasm, anxiety</p>
<p><strong>Neutral</strong>: neither positive or negative, such as stating a fact</p>
<p><strong>Mixed</strong>: when the sentiment is mixed</p>
</full-instructions>
<short-instructions>
Choose the primary sentiment that is expressed by the text.
</short-instructions>
</crowd-classifier>
</crowd-form>
"""
def create_task_ui():
'''
Creates a Human Task UI resource.
Returns:
struct: HumanTaskUiArn
'''
response = sagemaker.create_human_task_ui(
HumanTaskUiName=taskUIName,
UiTemplate={'Content': template})
return response
# +
# Task UI name - this value is unique per account and region. You can also provide your own value here.
taskUIName = 'ui-comprehend-' + str(uuid.uuid4())
# Create task UI
humanTaskUiResponse = create_task_ui()
humanTaskUiArn = humanTaskUiResponse['HumanTaskUiArn']
print(humanTaskUiArn)
# -
# ### Creating the Flow Definition
# In this section, we're going to create a flow definition definition. Flow Definitions allow us to specify:
#
# * The workforce that your tasks will be sent to.
# * The instructions that your workforce will receive. This is called a worker task template.
# * The configuration of your worker tasks, including the number of workers that receive a task and time limits to complete tasks.
# * Where your output data will be stored.
#
# This demo is going to use the API, but you can optionally create this workflow definition in the console as well.
#
# For more details and instructions, see: https://docs.aws.amazon.com/sagemaker/latest/dg/a2i-create-flow-definition.html.
# +
# Flow definition name - this value is unique per account and region. You can also provide your own value here.
flowDefinitionName = 'fd-dsoaws-comprehend-' + str(uuid.uuid4())
create_workflow_definition_response = sagemaker.create_flow_definition(
FlowDefinitionName= flowDefinitionName,
RoleArn= role,
HumanLoopConfig= {
"WorkteamArn": WORKTEAM_ARN,
"HumanTaskUiArn": humanTaskUiArn,
"TaskCount": 1,
"TaskDescription": "Identify the sentiment of the provided text",
"TaskTitle": "Classify Sentiment of Text"
},
OutputConfig={
"S3OutputPath" : output_path
}
)
flowDefinitionArn = create_workflow_definition_response['FlowDefinitionArn']
# -
# Describe flow definition - status should be active
for x in range(60):
describeFlowDefinitionResponse = sagemaker.describe_flow_definition(FlowDefinitionName=flowDefinitionName)
print(describeFlowDefinitionResponse['FlowDefinitionStatus'])
if (describeFlowDefinitionResponse['FlowDefinitionStatus'] == 'Active'):
print("Flow Definition is active")
break
time.sleep(2)
# ## Human Loops
# ### Detect Sentiment with AWS Comprehend
# Now that we have setup our Flow Definition, we are ready to call AWS Comprehend and start our human loops. In this tutorial, we are interested in starting a HumanLoop only if the SentimentScore returned by AWS Comprehend is less than 99%.
#
# So, with a bit of logic, we can check the response for each call to Detect Sentiment, and if the SentimentScore is less than 99%, we will kick off a HumanLoop to engage our workforce for a human review.
# # Sample Data
sample_reviews = ['I enjoy this product',
'I am unhappy with this product',
'It is okay',
'sometimes it works']
# # Comprehend helper method
# Will help us parse Detect Sentiment API responses
def capsToCamel(all_caps_string):
if all_caps_string == 'POSITIVE':
return 'Positive'
elif all_caps_string == 'NEGATIVE':
return 'Negative'
elif all_caps_string == 'NEUTRAL':
return 'Neutral'
# +
human_loops_started = []
SENTIMENT_SCORE_THRESHOLD = 0.99
for sample_review in sample_reviews:
# Call AWS Comprehend's Detect Sentiment API
response = comprehend.detect_sentiment(Text=sample_review, LanguageCode='en')
sentiment = response['Sentiment']
print(f'Processing sample_review: \"{sample_review}\"')
# Our condition for when we want to engage a human for review
if (response['SentimentScore'][capsToCamel(sentiment)]< SENTIMENT_SCORE_THRESHOLD):
humanLoopName = str(uuid.uuid4())
inputContent = {
"initialValue": sentiment.title(),
"taskObject": sample_review
}
start_loop_response = a2i.start_human_loop(
HumanLoopName=humanLoopName,
FlowDefinitionArn=flowDefinitionArn,
HumanLoopInput={
"InputContent": json.dumps(inputContent)
}
)
human_loops_started.append(humanLoopName)
print(f'SentimentScore of {response["SentimentScore"][capsToCamel(sentiment)]} is less than the threshold of {SENTIMENT_SCORE_THRESHOLD}')
print(f'Starting human loop with name: {humanLoopName} \n')
else:
print(f'SentimentScore of {response["SentimentScore"][capsToCamel(sentiment)]} is above threshold of {SENTIMENT_SCORE_THRESHOLD}')
print('No human loop created. \n')
# -
# ### Check Status of Human Loop
completed_human_loops = []
for human_loop_name in human_loops_started:
resp = a2i.describe_human_loop(HumanLoopName=human_loop_name)
print(f'HumanLoop Name: {human_loop_name}')
print(f'HumanLoop Status: {resp["HumanLoopStatus"]}')
print(f'HumanLoop Output Destination: {resp["HumanLoopOutput"]}')
print('\n')
if resp["HumanLoopStatus"] == "Completed":
completed_human_loops.append(resp)
# # Wait For Human Workers to Complete Their Tasks
# Navigate to the link below and login with your email and password that you used when you setup the Private Workforce.
workteamName = WORKTEAM_ARN[WORKTEAM_ARN.rfind('/') + 1:]
print("Navigate to the private worker portal and do the tasks. Make sure you've invited yourself to your workteam!")
print('https://' + sagemaker.describe_workteam(WorkteamName=workteamName)['Workteam']['SubDomain'])
# ### Check Status of Human Loop Again
completed_human_loops = []
for human_loop_name in human_loops_started:
resp = a2i.describe_human_loop(HumanLoopName=human_loop_name)
print(f'HumanLoop Name: {human_loop_name}')
print(f'HumanLoop Status: {resp["HumanLoopStatus"]}')
print(f'HumanLoop Output Destination: {resp["HumanLoopOutput"]}')
print('\n')
if resp["HumanLoopStatus"] == "Completed":
completed_human_loops.append(resp)
# ### View Task Results
# Once work is completed, Amazon A2I stores results in your S3 bucket and sends a Cloudwatch event. Your results should be available in the S3 OUTPUT_PATH when all work is completed.
# +
import re
import pprint
pp = pprint.PrettyPrinter(indent=4)
for resp in completed_human_loops:
splitted_string = re.split('s3://' + bucket + '/', resp['HumanLoopOutput']['OutputS3Uri'])
output_bucket_key = splitted_string[1]
response = s3.get_object(Bucket=bucket, Key=output_bucket_key)
content = response["Body"].read()
json_output = json.loads(content)
pp.pprint(json_output)
print('\n')
# -
# ### The End!
| 10_pipeline/human_in_the_loop/wip/06_Augmented_AI_With_Comprehend_Built_In_Sentiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys, os
sys.path.insert(0, os.path.join("..", ".."))
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.collections
import numpy as np
import open_cp.network
import open_cp.network_hotspot
import open_cp.sources.chicago
import open_cp.geometry
# -
# # Geometry reduction
#
# This notebook is some musings on [combinatorial explosion](https://en.wikipedia.org/wiki/Combinatorial_explosion) problems I am having.
#
# A problem we run into is that our graph algorithms are:
# - Slow. Massively not helped by a pure Python implementation
# - We end up considering huge numbers of paths.
#
# Let's look more closely at the graphs. Suppose we "topologically reduce" the graph: delete any vertex which has just two neighbours. This does not change the search we need to perform when assigning risk, but it does decrease the number of edges and vertices in the graph. **On second thoughts** this didn't seem to help that much.
with open("Case study Chicago/input.graph", "rb") as f:
graph = open_cp.network.PlanarGraph.from_bytes(f.read())
reduced = open_cp.network.simple_reduce_graph(graph)
graph.number_edges, reduced.number_edges
# +
fig, axes = plt.subplots(ncols=2, figsize=(18,8))
for ax in axes:
lc = matplotlib.collections.LineCollection(graph.as_lines(), color="black", linewidth=0.5)
ax.add_collection(lc)
xcs, ycs = [], []
for k in reduced.vertices:
xcs.append(graph.vertices[k][0])
ycs.append(graph.vertices[k][1])
for ax in axes:
ax.scatter(xcs, ycs)
axes[0].set(xlim=[355000, 365000], ylim=[565000, 575000])
axes[1].set(xlim=[358000, 360000], ylim=[567000, 569000])
None
# -
# # Just too many paths!
#
# The underlying problem seems to be that there is just a combinatorial explosion in the number of paths to consider.
#
# - We find a couple of nearby points in a dense maze of streets and avenues.
# - Then find how many paths there are from one starting point, of various lengths.
# - This is needed for the KDE algorithm-- there is just an explosion in the number of paths to consider as the maximum length increases.
xmin, xmax = 359000, 359200
ymin, ymax = 568500, 568750
[k for k in reduced.vertices if graph.vertices[k][0] >= xmin and graph.vertices[k][0] <= xmax
and graph.vertices[k][1] >= ymin and graph.vertices[k][1] <= ymax]
# +
verts = [6743, 17940]
fig, ax = plt.subplots(figsize=(8,8))
lc = matplotlib.collections.LineCollection(graph.as_lines(), color="black", linewidth=0.5)
ax.add_collection(lc)
xcs, ycs = [], []
for k in verts:
xcs.append(graph.vertices[k][0])
ycs.append(graph.vertices[k][1])
ax.scatter(xcs, ycs)
ax.set(xlim=[358000, 360000], ylim=[567500, 569500])
None
# -
# Not too many paths between these two vertices...
len(list(graph.paths_between(6743, 17940, 1000)))
# +
# Explosion in the number of paths to consider for the KDE method...
lengths = [10,20,50,100,200,500,1000,1100,1200,1300]
num_paths = []
for length in lengths:
nup = len(list(graph.walk_with_degrees(6743, None, length, 10000000000)))
num_paths.append(nup)
plt.plot(lengths, num_paths)
# -
# # Some hope
#
# Let's look at an example: pick a nearby edge and see which paths end up crossing it (these are the paths we end up summing in the KDE method to get the final risk estimate).
#
# - We then find that the "cumulative degree" rapidly gets very large
# - As we divide by this in the KDE method, we might conjecture that most of these paths are irrelevant.
# - This indeed turns out to be the case: most of the summands add very little to the log likelihood.
# +
print(graph.neighbourhood_edges(6743))
print(graph.edges[25201])
print(graph.neighbourhood_edges(13452))
edge = 25202
out = [ path for path in graph.walk_with_degrees(6743, None, 1300, 100000000)
if path[0] == edge ]
len(out)
# -
out.sort(key = lambda x : x[3])
out[:5], out[-5:]
# Example kernel
kernel = open_cp.network_hotspot.TriangleKernel(1500)
out1 = [ kernel.integrate(start, end) / div
for _, start, end, div in out ]
cumulative_sum = [0]
for x in out1:
cumulative_sum.append(cumulative_sum[-1] + x)
cumulative_sum = np.asarray(cumulative_sum)
#plt.plot(cummulative_sum)
plt.plot(np.log(cumulative_sum[1:]))
out[25], out[50]
# # Limiting the maximum degree
#
# If we limit the maximum degree to 20000 then the number of paths to consider begins to tail off, which is hopeful...
# +
lengths = list(range(100,2100,100))
num_paths = []
for length in lengths:
nup = len(list(graph.walk_with_degrees(6743, None, length, 20000)))
num_paths.append(nup)
plt.plot(lengths, num_paths)
# +
import pickle, lzma
with lzma.open("Case study Chicago/input.pic.xz", "rb") as f:
timed_points = pickle.load(f)
trainer = open_cp.network_hotspot.Trainer()
trainer.graph = graph
trainer.maximum_edge_length = 20
trainer.data = timed_points
predictor = trainer.compile()
# -
predictor.graph.number_edges, graph.number_edges
graph.vertices[6743], predictor.graph.vertices[6743]
# +
lengths = list(range(100,2100,100))
num_paths = []
for length in lengths:
nup = len(list(predictor.graph.walk_with_degrees(6743, None, length, 20000)))
num_paths.append(nup)
plt.plot(lengths, num_paths)
# -
# # Another approach to caching
#
# On the basis that [Almost all programming can be viewed as an exercise in caching](http://plasmasturm.org/log/542/) lets think a bit more.
#
# - Let's assume that maximum edge length is small compared to spatial scale.
# - So there should be little loss in accuracy by assuming all events occur at the mid-point of the edge they lie on
# - And we can approximate the "integral" of the kernel by taking it's value at the midpoint and multiply by the length of the edge.
#
# However, we are still left considering _all_ paths between edges, as the way to combine the length of the path, and the product of degrees, depends heavily on the (space) kernel is use.
#
# - So we either cache it all (which is a vast amount of data, probably too much for memory and time).
# - Or cache results which are only valid for one kernel. This doesn't seem like much (or any?) improvement over the caching system I am currently using.
# # Approximations
# +
fig, axes = plt.subplots(ncols=2, figsize=(19,8))
for ax in axes:
lc = matplotlib.collections.LineCollection(predictor.graph.as_lines(), color="black", linewidth=0.5)
ax.add_collection(lc)
xcs, ycs = [], []
xcs.append(graph.vertices[6743][0])
ycs.append(graph.vertices[6743][1])
ax.scatter(xcs, ycs)
(x1,y1),(x2,y2) = predictor.graph.as_lines()[8076]
ax.plot([x1,x2], [y1,y2], color="blue", linewidth=2)
axes[0].set(xlim=[358000, 360000], ylim=[567500, 569500])
axes[1].set(xlim=[358800, 359200], ylim=[568600, 569000])
None
# -
predictor.graph.edges[8076]
predictor.kernel = open_cp.network_hotspot.TriangleKernel(500)
risks = np.zeros(len(predictor.graph.edges))
predictor.add(risks, 8076, -1, 0.5, 1)
predictor.add(risks, 8076, 1, 0.5, 1)
# Remember to normalise!
risks /= predictor.graph.lengths
# +
fig, ax = plt.subplots(figsize=(8,8))
lines = np.asarray(predictor.graph.as_lines())
mask = risks > 0
ri, li = risks[mask], lines[mask]
lc = matplotlib.collections.LineCollection(li, color="black", linewidth=5)
lc.set_array(ri)
lc.set(cmap="Blues")
ax.add_collection(lc)
x, y = graph.vertices[6743]
d = 700
ax.set(xlim=[x-d, x+d], ylim=[y-d, y+d])
None
# -
# Look at the street edge 8076 is on in more detail.
# - The following code finds the edges in the street, stopping at the intersection
# - Then we plot the risk, which decays pretty linearly, as hoped.
edges = []
edge = 8076
while True:
if edge in edges:
break
edges.append(edge)
for k in predictor.graph.edges[edge]:
nhood = list(predictor.graph.neighbourhood_edges(k))
nhood.remove(edge)
if len(nhood) == 1:
edge = nhood[0]
break
edges
# +
r = [risks[i] for i in edges]
fig, axes = plt.subplots(ncols=2, figsize=(16,5))
ax = axes[0]
ax.plot(r)
ax.set(title="Risk against edge")
ax = axes[1]
x = [0]
for i in edges:
x.append(x[-1] + predictor.graph.length(i))
x = np.asarray(x)
ax.plot((x[:-1]+x[1:])/2, r)
for i in range(len(x)-1):
ax.plot([x[i], x[i+1]], [r[i], r[i]], color="black")
ax.set(title="Risk against distance", xlabel="meters")
None
# -
# ## Compare to distance between edges
#
# - Form the [Derived graph](https://en.wikipedia.org/wiki/Line_graph) and compute distances between the edges
# - Compare this to the "risk" our KDE method found above.
#
# **Conclusions:**
# - Not a good match
# - Especially for larger values of risk, we get multiple correlation lines, as compared to distance.
# - This is to be expected, as the KDE methods explicitly takes account of vertex degree.
dgraph = open_cp.network.to_derived_graph(predictor.graph, use_edge_indicies=True)
paths, prevs = open_cp.network.shortest_paths(dgraph, 8076)
assert min(paths.keys()) == 0
assert max(paths.keys()) == len(paths) - 1
paths = [paths[i] for i in range(len(paths))]
paths = np.asarray(paths)
disconnected = paths == -1
assert np.all(risks[disconnected] == 0)
risks, paths = risks[~disconnected], paths[~disconnected]
mask = risks > 0
np.max(paths[mask]), np.min(paths[~mask])
fig, axes = plt.subplots(ncols=2, figsize=(19,8))
ax = axes[0]
ax.scatter(risks[mask], paths[mask], marker="x", linewidth=1, color="black")
ax.set(xlabel="Risk", ylabel="Distance", xlim=[-0.0001, 0.0021])
ax = axes[1]
ax.scatter(np.log(risks[mask]), paths[mask], marker="x", linewidth=1, color="black")
ax.set(xlabel="Log risk", ylabel="Distance")
None
# ## Take account of vertex degree
#
# An obvious tactic is to compute the shortest path distance, but taking account of the vertex degree in the same way the KDE method does. Unfortunately, performing this calculation in the derived graph is wrong (draw a few example graphs...)
# +
risks = np.zeros(len(predictor.graph.edges))
predictor.add(risks, 8076, -1, 0.5, 1)
predictor.add(risks, 8076, 1, 0.5, 1)
risks /= predictor.graph.lengths
k1, k2 = predictor.graph.edges[8076]
paths1, prevs1 = open_cp.network.shortest_paths(predictor.graph, k1)
paths2, prevs2 = open_cp.network.shortest_paths(predictor.graph, k2)
paths1 = np.asarray([paths1[i] for i in range(len(paths1))])
paths2 = np.asarray([paths2[i] for i in range(len(paths2))])
paths, prevs = open_cp.network.shortest_paths(dgraph, 8076)
paths = np.asarray([paths[i] for i in range(len(paths))])
epaths, eprevs = open_cp.network.shortest_edge_paths(predictor.graph, 8076)
# +
def shortest(edge_index):
k1, k2 = predictor.graph.edges[edge_index]
halves = (predictor.graph.length(edge_index) + predictor.graph.length(8076)) / 2
choices = [paths1[k1], paths1[k2], paths2[k1], paths2[k2]]
le = min(choices)
index = choices.index(le)
return le + halves, index
edge = 8010
k1, k2 = predictor.graph.edges[edge]
le = predictor.graph.length(edge) / 2
shortest(edge), paths[edge], min(epaths[k1]+le, epaths[k2]+le)
# +
def degree_of_shortest_path(edge_index):
k1, k2 = predictor.graph.edges[edge_index]
kk1, kk2 = predictor.graph.edges[8076]
_, choice = shortest(edge_index)
if choice == 0:
vertex, pa = k1, prevs1
elif choice == 1:
vertex, pa = k2, prevs1
elif choice == 2:
vertex, pa = k1, prevs2
elif choice == 3:
vertex, pa = k2, prevs2
else:
raise AssertionError
v = vertex
cum_deg = max(1, predictor.graph.degree(v) - 1)
while True:
vv = pa[v]
if v == vv:
break
v = vv
d = max(1, predictor.graph.degree(v) - 1)
cum_deg *= d
return cum_deg
degree_of_shortest_path(8010)
# +
adjusted_dists = np.empty_like(risks)
for i, r in enumerate(risks):
if r == 0:
continue
adjusted_dists[i], _ = shortest(i)
adjusted_dists[i] = predictor.kernel(adjusted_dists[i])
adjusted_dists[i] = adjusted_dists[i] / degree_of_shortest_path(i)
mask = (risks > 0)
r = risks[mask]
adjusted_dists = adjusted_dists[mask]
# +
fig, axes = plt.subplots(ncols=2, figsize=(16,8))
ax = axes[0]
ax.scatter(r, adjusted_dists)
ax.set(xlabel="Risk", ylabel="From distance")
xmin, xmax = 0, max(r)
xd = (xmax - xmin) / 10
ymin, ymax = 0, max(adjusted_dists)
yd = (ymax - ymin) / 10
ax.set(xlim=[xmin-xd, xmax+xd], ylim=[ymin-yd, ymax+yd])
x = np.linspace(0, 0.002, 2)
ax.plot(x, x, color="red")
ax = axes[1]
mask = (r>0) & (adjusted_dists>0)
ax.scatter(np.log(r[mask]), np.log(adjusted_dists[mask]))
ax.set(xlabel="Log risk", ylabel="Log from distance")
None
# -
pred = open_cp.network_hotspot.ApproxPredictor(predictor)
risks_approx = np.zeros(len(predictor.graph.edges))
pred.add_edge(risks_approx, 8076, 0.5, 1)
risks_approx /= predictor.graph.lengths
# +
fig, axes = plt.subplots(ncols=2, figsize=(16,8))
ax = axes[0]
ax.scatter(risks, risks_approx)
ax.set(xlabel="Risk", ylabel="Approx risk")
xmin, xmax = 0, max(r)
xd = (xmax - xmin) / 10
ymin, ymax = 0, max(adjusted_dists)
yd = (ymax - ymin) / 10
ax.set(xlim=[xmin-xd, xmax+xd], ylim=[ymin-yd, ymax+yd])
x = np.linspace(0, 0.002, 2)
ax.plot(x, x, color="red")
ax = axes[1]
mask = (risks>0) & (risks_approx>0)
ax.scatter(np.log(risks[mask]), np.log(risks_approx[mask]))
ax.set(xlabel="Log risk", ylabel="Log from distance")
None
# -
| examples/Networks/Geometry reduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.metrics import confusion_matrix
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
import seaborn as sns
sns.set()
df = pd.read_csv('Real_fake_news.csv')
df.head()
df.tail()
df.shape
df.info()
df.describe()
# Dropping the unnecessary columns
df.drop(['Unnamed: 0','Unnamed: 0.1'], axis=1, inplace=True)
df.head()
# Dropping the rows with null values
df.dropna()
df.shape
#FInding rows that have duplicate values
df[df.duplicated(keep = 'last')]
#Removing the duplicates
df = df.drop_duplicates(subset = None, keep ='first')
df.head()
df.BinaryNumTarget.value_counts()
# +
# Defining numerical and categorical variables
num_atr=[]
cat_atr=['author' , 'statement' , 'source']
# -
# Correlation Matrix
df.corr()
df.dtypes
df.columns
df.head()
# +
# Random Forest Classification Algorithm
# -
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import export_graphviz
from sklearn.datasets import load_iris
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import mean_squared_error,r2_score
from sklearn.preprocessing import OneHotEncoder as onehot
from sklearn.preprocessing import LabelEncoder
# +
X= df.drop(['BinaryNumTarget', 'BinaryTarget', 'target'], axis=1)
y= df['BinaryNumTarget']
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.3,random_state=1)
# -
y.shape
X.shape
le = LabelEncoder()
X_train[cat_atr] = X_train[cat_atr].apply(le.fit_transform)
X_train[cat_atr].head()
Xtr = X_train[cat_atr]
Xtr.shape
# +
le = LabelEncoder()
X_test[cat_atr] = X_test[cat_atr].apply(le.fit_transform)
Xtr1 = X_test[cat_atr]
Xtr1.shape
# +
from sklearn.ensemble import RandomForestClassifier
clf=RandomForestClassifier(n_estimators=100)
clf.fit(Xtr,y_train)
y_pred=clf.predict(Xtr1)
# -
from sklearn import metrics
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
metrics.confusion_matrix(y_test, y_pred)
print(classification_report(y_test, y_pred))
from sklearn.ensemble import RandomForestClassifier
clf=RandomForestClassifier(n_estimators=100)
clf.fit(X_train,y_train)
df.columns
df.feature_names = df[['author', 'statement', 'source']]
df.target_names = df['BinaryNumTarget']
import pandas as pd
feature_imp = pd.Series(clf.feature_importances_).sort_values(ascending=False)
feature_imp
list(zip(df.columns, feature_imp))
#visualizing feature importance results
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# Creating a bar plot
sns.barplot(x=feature_imp, y=feature_imp.index)
# Add labels to your graph
plt.xlabel('Feature Importance Score')
plt.ylabel('Features')
plt.title("Visualizing Important Features")
plt.legend()
plt.show()
from sklearn.ensemble import RandomForestClassifier
clf=RandomForestClassifier(n_estimators=100)
clf.fit(Xtr,y_train)
y_pred=clf.predict(Xtr1)
from sklearn import metrics
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
| Real Fake News - Random Forest Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aubricot/computer_vision_with_eol_images/blob/master/classification_for_image_tagging/flower_fruit/classify_images.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_TcYNLBrWC0C"
# # Run images through flower/fruit classification pipeline
# ---
# *Last Updated 29 September 2020*
# 1) Run images through Model 7 and 11 and save results to tsv in batches of 5,000 images at a time.
# 2) Post-process results from image batches to filter predictions using confidence values chosen in [det_conf_threshold.ipynb](https://colab.research.google.com/github/aubricot/computer_vision_with_eol_images/blob/master/classification_for_image_tagging/flower_fruit/det_conf_threshold.ipynb) and save results to tsv.
# 3) Display filtered classification results on images and adjust confidence thresholds as needed.
# + [markdown] id="ZYW4W2aqdnTN"
# ### Imports
# ---
# + id="6k81-h_UV_ny"
# Mount google drive to import/export files
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + id="1AGFM4fSWhbT"
# For working with data and plotting graphs
import itertools
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# For image classification and training
import tensorflow as tf
# + [markdown] id="u2PEaR_a_0QH"
# ### Run images through model(s) for classification of flowers/fruits
# ---
# Use model(s) and confidence threshold(s) selected in det_conf_threshold.ipynb
# + [markdown] id="b0u9Dd5OmWAO"
# #### Define functions & variables
# + id="N0dRSsDAk-GW" cellView="both"
import csv
# Load trained model from path
TRAIN_SESS_NUM = "07"
saved_model_path = '/content/drive/My Drive/summer20/classification/flower_fruit/saved_models/' + TRAIN_SESS_NUM
flower_model = tf.keras.models.load_model(saved_model_path)
TRAIN_SESS_NUM = "11"
saved_model_path = '/content/drive/My Drive/summer20/classification/flower_fruit/saved_models/' + TRAIN_SESS_NUM
null_model = tf.keras.models.load_model(saved_model_path)
label_names = ['Flower', 'Fruit', 'Null']
# Load in image from URL
# Modified from https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/saved_model.ipynb#scrollTo=JhVecdzJTsKE
def image_from_url(url, fn):
file = tf.keras.utils.get_file(fn, url) # Filename doesn't matter
disp_img = tf.keras.preprocessing.image.load_img(file)
img = tf.keras.preprocessing.image.load_img(file, target_size=[224, 224])
x = tf.keras.preprocessing.image.img_to_array(img)
x = tf.keras.applications.mobilenet_v2.preprocess_input(
x[tf.newaxis,...])
return x, disp_img
# Read in EOL image bundle dataframe
# TO DO: Type in image bundle address using form field to right
bundle = 'https://editors.eol.org/other_files/bundle_images/files/images_for_Angiosperms_20K_breakdown_000031.txt' #@param {type:"string"}
df = pd.read_csv(bundle, sep='\t', header=0)
df.head()
# + [markdown] id="tEgxXYbTmY1P"
# #### Run 20K image bundle through classification pipeline
# + id="IL0DthODQw45"
# Write header row of output crops file
# TO DO: Change file name for each bundle/run abcd if doing 4 batches using dropdown form to right
tags_file = "angiosperm_tags_flowfru_20k_d" #@param ["angiosperm_tags_flowfru_20k_a", "angiosperm_tags_flowfru_20k_b", "angiosperm_tags_flowfru_20k_c", "angiosperm_tags_flowfru_20k_d"]
tags_fpath = "/content/drive/My Drive/summer20/classification/flower_fruit/results/" + tags_file + ".tsv"
with open(tags_fpath, 'a') as out_file:
tsv_writer = csv.writer(out_file, delimiter='\t')
tsv_writer.writerow(["eolMediaURL", "identifier", \
"dataObjectVersionID", "ancestry", \
"tag7", "tag7_conf", "tag11", "tag11_conf"])
# + id="0ZXo6iVvBF0G"
# Set number of seconds to timeout if image url taking too long to open
import socket
socket.setdefaulttimeout(10)
import time
# TO DO: Set start and end rows to run inference for from EOL image bundle using form field to right
# If running in 4 batches of 5000 images, use values in dropdown menu
start = 15000 #@param ["0", "5000", "10000", "15000"] {type:"raw"}
end = 20000 #@param ["5000", "10000", "15000", "20000"] {type:"raw"}
# Loop through EOL image bundle to classify images and generate tags
for i, row in df.iloc[start:end].iterrows():
try:
# Get url from image bundle
url = df['eolMediaURL'][i]
# Read in image from url
fn = str(i) + '.jpg'
img, disp_img = image_from_url(url, fn)
# Record inference time
start_time = time.time()
# Detection and draw boxes on image
# For flowers/fruits (reproductive structures)
predictions = flower_model.predict(img, batch_size=1)
label_num = np.argmax(predictions)
f_conf = predictions[0][label_num]
f_class = label_names[label_num]
# For null (no reproductive structures)
predictions = null_model.predict(img, batch_size=1)
label_num = np.argmax(predictions)
n_conf = predictions[0][label_num]
n_class = label_names[label_num]
end_time = time.time()
# Display progress message after each image
print('Inference complete for {} of {} images'.format(i, (end-start)))
# Optional: Show classification results for images
# Only use to view predictions on <50 images at a time
#_, ax = plt.subplots(figsize=(10, 10))
#ax.imshow(disp_img)
#plt.axis('off')
#plt.title("{}) Mod 7 Prediction: {}, Confidence: {}%, \
#\n Mod 11 Prediction: {}, Confidence: {}%, Inference Time: {}".format(i, \
#f_class, f_conf, n_class, n_conf,format(end_time-start_time, '.3f')))
# Export tagging results to tsv
# Define variables for export
identifier = df['identifier'][i]
dataObjectVersionID = df['dataObjectVersionID'][i]
ancestry = df['ancestry'][i]
with open(tags_fpath, 'a') as out_file:
tsv_writer = csv.writer(out_file, delimiter='\t')
tsv_writer.writerow([url, identifier, dataObjectVersionID, ancestry, \
f_class, f_conf, n_class, n_conf])
except:
print('Check if URL from {} is valid'.format(url))
# + [markdown] id="ZRDQAbG8OziR"
# ### Post-process classification predictions using confidence threshold values for models 7 and 11 chosen in det_conf_threshold.ipynb
# ---
# + id="gzb2pJI0O-ru"
# Combine exported model predictions and confidence values from above to one dataframe
base = '/content/drive/My Drive/summer20/classification/flower_fruit/results/angiosperm_tags_flowfru_20k_'
exts = ['a.tsv', 'b.tsv', 'c.tsv', 'd.tsv']
all_filenames = [base + e for e in exts]
df = pd.concat([pd.read_csv(f, sep='\t', header=0, na_filter = False) for f in all_filenames], ignore_index=True)
# Filter predictions using determined confidence value thresholds
# Make column for "reproductive structures present?" tag
df['reprod'] = np.nan
# Adjust final tag based on Model 7 and 11 predictions and confidence values
for i, row in df.iterrows():
# If Model 7 predicts flower with >1.6 confidence
if df['tag7'][i]=="Flower" and df['tag7_conf'][i]>1.6:
# And Model 11 does not predict null with >= 1.5 confidence
if df['tag11'][i]=="Null" and df['tag11_conf'][i]>=1.5:
# Reproductive structures present -> YES
df['reprod'][i] = "Y"
# And Model 11 predicts null with >= 1.5 confidence
elif df['tag11'][i]=="Null" and df['tag11_conf'][i]<1.5:
# Reproductive structures present -> NO
df['reprod'][i] = "N"
# And Model 11 predicts fruit or flower with any confidence
else:
# Reproductive structures present -> NO
df['reprod'][i] = "Y"
# If Model 7 predicts flower with <= 1.6 confidence
elif df['tag7'][i]=="Flower" and df['tag7_conf'][i]<=1.6:
# Reproductive structures present -> Maybe
df['reprod'][i] = "M"
# If Model 7 predicts fruit or null with any confidence
else:
# Reproductive structures present -> NO
df['reprod'][i] = "N"
# Make all tags for grasses -> N (Poaceae, especially bamboo had bad classification results on manual inspection)
taxon = "Poaceae"
df['reprod'].loc[df.ancestry.str.contains(taxon, case=False, na=False)] = "N"
# Write results to tsv
df.to_csv("/content/drive/My Drive/summer20/classification/flower_fruit/results/angiosperm_tags_flowfru_20k_finaltags.tsv", sep='\t', index=False)
# + id="_iwVwlLKNWSY"
# Inspect results
print(df.head(10))
print("Number of positive identified reproductive structures: {}".format(len(df[df['reprod']=="Y"])))
print("Number of possible identified reproductive structures: {}".format(len(df[df['reprod']=="M"])))
print("Number of negative identified reproductive structures: {}".format(len(df[df['reprod']=="N"])))
# + [markdown] id="tDnCXzDGVa6t"
# ### Display final classification results on images
# ---
# + id="TxurlbjZJd9q"
# Set number of seconds to timeout if image url taking too long to open
import socket
socket.setdefaulttimeout(10)
# TO DO: Update file path to finaltags.tsv file
path = "/content/drive/My Drive/summer20/classification/flower_fruit/results/"
f = "angiosperm_tags_flowfru_20k_finaltags.tsv" #@param
fpath = path + f
df = pd.read_csv(fpath, sep='\t', header=0, na_filter = False)
# Function to load in image from URL
# Modified from https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/saved_model.ipynb#scrollTo=JhVecdzJTsKE
def image_from_url(url, fn):
file = tf.keras.utils.get_file(fn, url) # Filename doesn't matter
disp_img = tf.keras.preprocessing.image.load_img(file)
img = tf.keras.preprocessing.image.load_img(file, target_size=[224, 224])
x = tf.keras.preprocessing.image.img_to_array(img)
x = tf.keras.applications.mobilenet_v2.preprocess_input(
x[tf.newaxis,...])
return x, disp_img
# TO DO: Set start and end rows to run inference for from EOL image bundle using form field to right
# If running in 4 batches of 5000 images, use values in dropdown menu
start = 0#@param {type:"raw"}
end = 50 #@param {type:"raw"}
# Loop through EOL image bundle to classify images and generate tags
for i, row in df.iloc[start:end].iterrows():
try:
# Get url from image bundle
url = df['eolMediaURL'][i]
# Read in image from url
fn = str(i) + '.jpg'
img, disp_img = image_from_url(url, fn)
# Record inference time
pred = df['reprod'][i]
# Display progress message after each image is loaded
print('Successfully loaded {} of {} images'.format(i+1, (end-start)))
# Show classification results for images
# Only use to view predictions on <50 images at a time
_, ax = plt.subplots(figsize=(10, 10))
ax.imshow(disp_img)
plt.axis('off')
plt.title("{}) Combined Mod 7 & 11 Prediction: {}".format(i+1, pred))
except:
print('Check if URL from {} is valid'.format(url))
| classification_for_image_tagging/flower_fruit/classify_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import plotly.express as px
import plotly.graph_objects as go
from plotly.subplots import make_subplots
df = pd.read_csv('model_stats.csv')
df.drop(columns=['Unnamed: 0', 'Unnamed: 0.1', 'Unnamed: 0.1.1', 'Unnamed: 0.1.1.1', 'Unnamed: 0.1.1.1.1', 'Unnamed: 0.1.1.1.1.1'], inplace=True)
px.bar(df, x='model_datetime', y=['mean_mse', 'median_mse'], barmode='group', title='Median and Mean MSE values (lower is better)')
df.mean_mse.describe()
px.box(df, x='mean_mse')
df.median_mse.describe()
px.box(df, x='median_mse')
| model_analysis_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv
# language: python
# name: venv
# ---
# ## Membership inference on text
# ### Stanford Sentiment Treebank (SST) movie review dataset for sentiment analysis
# +
import torch
from torchtext import data
from torchtext import datasets
import sys
import seaborn as sns
from sklearn.metrics import roc_curve, auc
sys.path.insert(0, '../../../Utils/')
import matplotlib.pyplot as plt
# %matplotlib inline
import models
from train import *
from metrics import *
print("Python: %s" % sys.version)
print("Pytorch: %s" % torch.__version__)
# -
# ### Load SST using Torchtext
# +
# To fix the following error: OSError: [E050] Can't find model 'en'. It doesn't seem to be a shortcut link, a Python package or a valid path to a data directory.
# Run:
# python -m spacy download en
TEXT = data.Field(tokenize='spacy')
LABEL = data.LabelField(tensor_type=torch.LongTensor)
target, val, shadow = datasets.SST.splits(TEXT, LABEL, root='../../../Datasets/SST_data', fine_grained=True)
target_in, target_out = target.split(split_ratio=0.5)
shadow_in, shadow_out = shadow.split(split_ratio=0.5)
TEXT.build_vocab(target, max_size=25000, vectors="glove.6B.100d", vectors_cache='../../../Datasets/SST_data/vector_cache')
LABEL.build_vocab(target)
BATCH_SIZE = 32
shadow_in_itr, shadow_out_itr, target_in_itr, target_out_itr, val_itr = data.BucketIterator.splits(
(shadow_in, shadow_out, target_in, target_out, val),
batch_size = BATCH_SIZE,
sort_key= lambda x: len(x.text),
repeat=False
)
# -
# ### Create bidirectional LSTM model for sentiment analysis
# +
vocab_size = len(TEXT.vocab)
embedding_size = 100
hidden_size = 256
output_size = 5
target_model = models.RNN(vocab_size, embedding_size, hidden_size, output_size)
shadow_model = models.RNN(vocab_size, embedding_size, hidden_size, output_size)
pretrained_embeddings = TEXT.vocab.vectors
target_model.embedding.weight.data.copy_(pretrained_embeddings)
shadow_model.embedding.weight.data.copy_(pretrained_embeddings)
print("")
target_optimizer = torch.optim.Adam(target_model.parameters())
shadow_optimizer = torch.optim.Adam(shadow_model.parameters())
criterion = torch.nn.CrossEntropyLoss()
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
target_model = target_model.to(device)
shadow_model = shadow_model.to(device)
criterion = criterion.to(device)
# -
# ### Utility functions
# +
def classification_accuracy(preds, y):
correct = (preds == y).float() #convert into float for division
acc = correct.sum()/len(correct)
return acc
def binary_accuracy(preds, y):
rounded_preds = torch.round(preds)
correct = (rounded_preds == y).float() #convert into float for division
acc = correct.sum()/len(correct)
return acc
# -
# ### Evaluation functions
# +
def evaluate(model, iterator, criterion):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
acc = classification_accuracy(predictions.argmax(dim=1), batch.label)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
def evaluate_inference(target_model, attack_model, in_iterator, out_iterator, criterion, k):
epoch_loss = 0
epoch_acc = 0
shadow_model.eval()
attack_model.eval()
predictions = np.array([])
labels = np.array([])
with torch.no_grad():
for in_batch, out_batch in zip(in_iterator, out_iterator):
in_size = len(in_batch.label)
out_size = len(out_batch.label)
in_lbl = torch.ones(in_size).to(device)
out_lbl = torch.zeros(out_size).to(device)
in_predictions = torch.nn.functional.softmax(target_model(in_batch.text.detach()), dim=1).detach()
out_predictions = torch.nn.functional.softmax(target_model(out_batch.text.detach()), dim=1).detach()
in_sort, _ = torch.sort(in_predictions, descending=True)
in_top_k = in_sort[:,:k].clone().to(device)
out_sort, _ = torch.sort(out_predictions, descending=True)
out_top_k = out_sort[:,:k].clone().to(device)
in_inference = attack_model(in_top_k).squeeze(1)
out_inference = attack_model(out_top_k).squeeze(1)
in_probability = torch.nn.functional.sigmoid(in_inference).detach().cpu().numpy()
out_probability = torch.nn.functional.sigmoid(out_inference).detach().cpu().numpy()
loss = (criterion(in_inference, in_lbl) + criterion(out_inference, out_lbl)) / 2
acc = (binary_accuracy(in_inference, in_lbl) + binary_accuracy(out_inference, out_lbl)) / 2
predictions = np.concatenate((predictions, in_probability), axis=0)
labels = np.concatenate((labels, np.ones(in_size)), axis=0)
predictions = np.concatenate((predictions, out_probability), axis=0)
labels = np.concatenate((labels, np.zeros(out_size)), axis=0)
epoch_loss += loss.item()
epoch_acc += acc.item()
fpr, tpr, thresholds = roc_curve(labels, predictions, pos_label=1)
roc_auc = auc(fpr, tpr)
return epoch_loss / len(in_iterator), epoch_acc / len(in_iterator), fpr, tpr, roc_auc
# -
# ### Train functions
# +
def train(model, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
optimizer.zero_grad()
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
acc = classification_accuracy(predictions.argmax(dim=1), batch.label)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
def train_nlp_attack(shadow_model, attack_model, in_iterator, out_iterator, optimizer, criterion, k):
epoch_loss = 0
epoch_acc = 0
shadow_model.eval()
attack_model.train()
in_input = np.empty((0,2))
out_input = np.empty((0,2))
for in_batch, out_batch in zip(in_iterator, out_iterator):
optimizer.zero_grad()
in_predictions = torch.nn.functional.softmax(shadow_model(in_batch.text.detach()), dim=1).detach()
out_predictions = torch.nn.functional.softmax(shadow_model(out_batch.text.detach()), dim=1).detach()
in_lbl = torch.ones(len(in_batch.label)).to(device)
out_lbl = torch.zeros(len(out_batch.label)).to(device)
in_sort, _ = torch.sort(in_predictions, descending=True)
in_top_k = in_sort[:,:k].clone().to(device)
out_sort, _ = torch.sort(out_predictions, descending=True)
out_top_k = out_sort[:,:k].clone().to(device)
in_inference = attack_model(in_top_k).squeeze(1)
out_inference = attack_model(out_top_k).squeeze(1)
in_input = np.vstack((in_input, torch.cat((torch.max(in_predictions, dim=1, keepdim=True)[0], in_batch.label.view(-1,1).type(torch.cuda.FloatTensor)), dim=1).cpu().numpy() ))
out_input = np.vstack((out_input, torch.cat((torch.max(out_predictions, dim=1, keepdim=True)[0], out_batch.label.view(-1,1).type(torch.cuda.FloatTensor)), dim=1).cpu().numpy() ))
loss = (criterion(in_inference, in_lbl) + criterion(out_inference, out_lbl)) / 2
acc = (binary_accuracy(in_inference, in_lbl) + binary_accuracy(out_inference, out_lbl)) / 2
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(in_iterator), epoch_acc / len(in_iterator), in_input, out_input
# -
# ### Train target model
# +
n_epochs_classification = 30
for epoch in range(n_epochs_classification):
train_loss, train_acc = train(target_model, target_in_itr, target_optimizer, criterion)
valid_loss, valid_acc = evaluate(target_model, val_itr, criterion)
print('Epoch: %02d, Train Loss: %.3f, Train Acc: %.2f%%, Val. Loss: %.3f, Val. Acc: %.2f%%' % (epoch+1, train_loss, train_acc*100, valid_loss, valid_acc*100))
# -
# ### Train shadow model
for epoch in range(n_epochs_classification):
train_loss, train_acc = train(shadow_model, shadow_in_itr, shadow_optimizer, criterion)
valid_loss, valid_acc = evaluate(shadow_model, val_itr, criterion)
print('Epoch: %02d, Train Loss: %.3f, Train Acc: %.2f%%, Val. Loss: %.3f, Val. Acc: %.2f%%' % (epoch+1, train_loss, train_acc*100, valid_loss, valid_acc*100))
# ### Evaluate models on test set
# +
target_test_loss, target_test_acc = evaluate(target_model, val_itr, criterion)
shadow_test_loss, shadow_test_acc = evaluate(shadow_model, val_itr, criterion)
print('Target net test accuracy: %.2f , Shadow net test accuracy: %.2f' % (target_test_acc, shadow_test_acc))
# -
# ### Create ML-leaks adversary 1 model (multi layer perceptron)
# +
attack_model = models.mlp(3,1,32).to(device)
attack_loss = torch.nn.BCELoss()
attack_optim = torch.optim.Adam(attack_model.parameters(), lr=0.01)
# -
# ### Train attack model
# +
n_epochs_attack = 30
for epoch in range(n_epochs_attack):
train_loss, train_acc, in_input, out_input = train_nlp_attack(shadow_model, attack_model, shadow_in_itr, shadow_out_itr, attack_optim, attack_loss, 3)
valid_loss, valid_acc, fpr, tpr, roc_auc = evaluate_inference(target_model, attack_model, target_in_itr, target_out_itr, attack_loss, 3)
print('Epoch: %02d, Train Loss: %.3f, Train Acc: %.2f%%, Val. Loss: %.3f, Val. Acc: %.2f%%' % (epoch+1, train_loss, train_acc*100, valid_loss, valid_acc*100))
# -
# ### Attack Results
# +
lw = 2
plt.figure()
plt.plot(fpr, tpr, lw=lw,
label='ROC curve (auc = %.2f)' % (roc_auc))
plt.plot([0,1],[0,1], color='navy', lw=lw, linestyle='--')
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.xlim([0.0,1.0])
plt.ylim([0.0, 1.05])
plt.title('SST Membership Inference ROC curve')
plt.legend(loc="lower right")
plt.show()
import pickle
results = (fpr, tpr, roc_auc)
pickle.dump(results, open("sst_results", "wb"))
# +
# in_positive = in_input[in_input[:,1] == 1][:,0]
# in_negative = in_input[in_input[:,1] == 0][:,0]
# out_positive = out_input[out_input[:,1] == 1][:,0]
# out_negative = out_input[out_input[:,1] == 0][:,0]
# plt.figure()
# sns.distplot(in_positive[:100],label='in positive', kde=True, hist=True, norm_hist=True)
# sns.distplot(out_positive[:100],label='out positive', kde=True, hist=True, norm_hist=True)
# plt.legend()
# plt.xlim([0.4, 1.05])
# plt.ylim([0.0, 20])
# plt.title("Positive predictions")
# plt.show()
# plt.figure()
# sns.distplot(in_negative,label='in negative')
# sns.distplot(out_negative,label='out negative')
# plt.legend()
# plt.title("Negative predictions")
# plt.show()
| Attack_baselines/Membership_Inference/ml_leaks/ml-leaks_adversary_1-SST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Rf2qwVcKOLh6"
# Time series data structure
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from statsmodels.tsa.stattools import adfuller
from numpy import log
import math
# + id="ILN5RdKQQg46"
df=pd.read_csv('/content/airline-passengers.csv', header=0, index_col=0, squeeze=True)
# + colab={"base_uri": "https://localhost:8080/"} id="ycOAJboJQz4Y" outputId="c36bb96e-815d-4746-bc81-975c90960d84"
#put alll values in x variable
X=df.values
#app addfuller test on the data (x) and stores it in Result
result= adfuller(X)
result
# + colab={"base_uri": "https://localhost:8080/"} id="yLObRf9YXEt3" outputId="56e89d8f-1bb9-44ee-f342-f2340bd4f425"
X_log = log(X)
result_log= adfuller(X_log)
result_log
# + colab={"base_uri": "https://localhost:8080/"} id="KBORyy3eSBGH" outputId="4caf4cfd-85ce-4a8c-bdb5-2a431c84596d"
print('ADF Statistics: %f' %result[0])
print('p_values: %f' %result[1])
print('Critical Values:')
for key, value in result[4].items():
print('\t%s: %.3f' %(key, value))
# + colab={"base_uri": "https://localhost:8080/"} id="c10COVVrSyGd" outputId="ae71eb2f-32c5-4113-f956-d410e1add248"
if result[0]<result[4]['5%']:
print('Rejected Null Hypothesis- so the time series is non stationary')
else:
print('Fail to Reject Null Hypothesis- so the time series is stationary')
# + id="qmZZerjjVDtL"
| Time_Series_Analysis_ADFTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from gridworld import GridworldEnv
env = GridworldEnv()
def value_iteration(env, theta=0.0001, discount_factor=1.0):
"""
Value Iteration Algorithm.
Args:
env: OpenAI env. env.P represents the transition probabilities of the environment.
env.P[s][a] is a list of transition tuples (prob, next_state, reward, done).
env.nS is a number of states in the environment.
env.nA is a number of actions in the environment.
theta: We stop evaluation once our value function change is less than theta for all states.
discount_factor: Gamma discount factor.
Returns:
A tuple (policy, V) of the optimal policy and the optimal value function.
"""
V = np.zeros(env.nS)
policy = np.zeros([env.nS, env.nA])
#helper function returning best action,value pair for particular state
def best_action_value(env,V,s,discount_factor=discount_factor):
A = dict()
for a in range(env.nA):
A[a] = 0
for (prob, next_state, reward, done) in env.P[s][a]:
A[a] += prob*( reward + discount_factor*V[next_state] ) # bellman equation
best_a = 0
best_v = float('-inf')
for a,v in A.items():
if best_v < v:
best_v = v
best_a = a
return best_a, best_v
#optimize value function
while True:
biggest_change = 0
for s in range(env.nS):
old_v = V[s]
_, new_v = best_action_value(env,V,s,discount_factor)
V[s] = new_v
if biggest_change < abs(old_v-new_v):
biggest_change = abs(old_v-new_v)
if biggest_change < theta:
break
#find best policy
for s in range(env.nS):
best_a,_ = best_action_value(env,V,s,discount_factor)
policy[s] = np.eye(env.nA)[best_a]
return policy, V
# +
policy, v = value_iteration(env)
print("Policy Probability Distribution:")
print(policy)
print("")
print("Reshaped Grid Policy (0=up, 1=right, 2=down, 3=left):")
print(np.reshape(np.argmax(policy, axis=1), env.shape))
print("")
print("Value Function:")
print(v)
print("")
print("Reshaped Grid Value Function:")
print(v.reshape(env.shape))
print("")
# -
# Test the value function
expected_v = np.array([ 0, -1, -2, -3, -1, -2, -3, -2, -2, -3, -2, -1, -3, -2, -1, 0])
np.testing.assert_array_almost_equal(v, expected_v, decimal=2)
| Classroom-Codes/Policy Optimization Methods/Gridworld - Value Iteration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ve_sales_prediction
# language: python
# name: ve_sales_prediction
# ---
# # <font color='#002726'> Data Science em Produção </font>
#
# =-=- ROSSMANN - STORE SALES PREDICTION -=-=
# + [markdown] heading_collapsed=true
# # <font color='#3F0094'> 0. Imports </font>
# + hidden=true
# general use
import numpy as np
import pandas as pd
# helper function
import inflection
# feature engineering and data analysis
import seaborn as sns
from matplotlib import gridspec, pyplot as plt
from IPython.display import Image
from scipy.stats import ranksums, chi2_contingency
from datetime import datetime, timedelta
from statsmodels.stats.weightstats import DescrStatsW, CompareMeans
# data preparation
from sklearn.preprocessing import RobustScaler, MinMaxScaler, LabelEncoder
# feature selection
from boruta import BorutaPy
# machine learning
import pickle
import xgboost as xgb
from sklearn.linear_model import LinearRegression, Lasso
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error, mean_absolute_percentage_error, mean_squared_error
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 0.1. Helper Functions </font>
# + code_folding=[] hidden=true
# Notebook Setups
sns.set_style('darkgrid')
sns.set_context('talk')
sns.set_palette('Set2')
# Functions
def bootstrap(attribute, estimate='mean', n_repeat=100, n_sample=100, ci=95):
"""Bootstrap"""
results = []
if estimate == 'mean':
for n in range(n_repeat):
sample = np.random.choice(a=attribute, size=n_sample)
results.append(np.mean(sample))
elif estimate == 'median':
for n in range(n_repeat):
sample = np.random.choice(a=attribute, size=n_sample)
results.append(np.median(sample))
else:
results = [0]
ci_bottom = (100 - ci) / 2
ci_top = ci + (100 - ci) / 2
statistic_mean = np.mean(results)
statistic_std_error = np.std(results)
lower_percentile = np.percentile(results, q=ci_bottom)
upper_percentile = np.percentile(results, q=ci_top)
return [statistic_mean, statistic_std_error, lower_percentile, upper_percentile]
def cramer_v(x, y):
"""Cramér's V formula to measure the association between two nominal variables"""
# confusion matrix - getting the values only (as matrix)
cm = pd.crosstab(x, y).values
# chi2_contingency from scipy returns a list, the first value is the statistic test result
chi2 = chi2_contingency(cm)[0]
# n is the grand total of observations
n = cm.sum()
# number of rows and columns
r, k = cm.shape
# bias correction
phi_cor = max(0, chi2/n - (k-1)*(r-1)/(n-1))
k_cor = k - (k-1)**2/(n-1)
r_cor = r - (r-1)**2/(n-1)
return np.sqrt((phi_cor) / (min(k_cor-1, r_cor-1)))
def ml_error(model_name, y, yhat):
"""Tests machine learning model prediction error."""
# mean absolute error (MAE)
mae = mean_absolute_error(y , yhat)
# mean absolute percentage error (MAPE)
mape = mean_absolute_percentage_error(y, yhat)
# root-mean-square error (RMSE)
rmse = np.sqrt(mean_squared_error(y, yhat))
return pd.DataFrame({'Model Name': model_name,
'MAE': mae,
'MAPE': mape,
'RMSE': rmse}, index=[0])
def cross_validation(X_train, kfold, model_name, ml_model, verbose=False):
# lists to keep the error results
mae_list, mape_list, rmse_list = [], [], []
# cross validation folds
for k in range(kfold, 0, -1):
# checking if verbose is true
if verbose:
print(f'\nKFold Number: {k}')
# start date and end date of validation
validation_start_date = X_train['date'].max() - timedelta(days=k*6*7)
validation_end_date = X_train['date'].max() - timedelta(days=(k-1)*6*7)
# splitting into traning and validation
training = X_train[X_train['date'] < validation_start_date]
validation = X_train[(X_train['date'] >= validation_start_date) & (X_train['date'] <= validation_end_date)]
# preparing training and validation datasets - removing response subsets
# training
y_training = training['sales']
x_training_ml = training.drop(['date', 'sales'], axis=1)
# validation
y_validation = validation['sales']
x_validation_ml = validation.drop(['date', 'sales'], axis=1)
# model
model_fit = ml_model.fit(x_training_ml, y_training)
# predicition
yhat = model_fit.predict(x_validation_ml)
# performance
result = ml_error(model_name, np.expm1(y_validation), np.expm1(yhat))
# appending error values to the lists in each iteration of KFold
mae_list.append(result['MAE'][0])
mape_list.append(result['MAPE'][0])
rmse_list.append(result['RMSE'][0])
# returning a dataframe with mean and std of each error measure
return pd.DataFrame({
'Model Name': model_name,
'MAE CV': np.round(np.mean(mae_list), 2).astype(str) + ' +/- ' + np.round((np.std(mae_list)), 2).astype(str),
'MAPE CV': np.round(np.mean(mape_list), 2).astype(str) + ' +/- ' + np.round((np.std(mape_list)), 2).astype(str),
'RMSE CV': np.round(np.mean(rmse_list), 2).astype(str) + ' +/- ' + np.round((np.std(rmse_list)), 2).astype(str),
}, index=[0])
# + [markdown] hidden=true
# ## <font color='#200CF'> 0.2. Loading Data </font>
# + hidden=true
# loading historical data - including Sales
df_sales_raw = pd.read_csv('../raw_data/train.csv', low_memory=False)
# loading information about the stores
df_store_raw = pd.read_csv('../raw_data/store.csv', low_memory=False)
# merging dataframes
df_raw = pd.merge(df_sales_raw, df_store_raw, how='left', on='Store')
# + [markdown] hidden=true
# ### <font color='#F37126'> Data Fields </font>
# + [markdown] hidden=true
# **Most of the fields are self-explanatory. The following are descriptions for those that aren't.**
#
# - **Id** - an Id that represents a (Store, Date) duple within the test set;
# - **Store** - a unique Id for each store;
# - **Sales** - the turnover for any given day (this is what you are predicting);
# - **Customers** - the number of customers on a given day;
# - **Open** - an indicator for whether the store was open: 0 = closed, 1 = open;
# - **StateHoliday** - indicates a state holiday. Normally all stores, with few exceptions, are closed on state holidays. Note that all schools are closed on public holidays and weekends. a = public holiday, b = Easter holiday, c = Christmas, 0 = None;
# - **SchoolHoliday** - indicates if the (Store, Date) was affected by the closure of public schools;
# - **StoreType** - differentiates between 4 different store models: a, b, c, d;
# - **Assortment** - describes an assortment level: a = basic, b = extra, c = extended;
# - **CompetitionDistance** - distance in meters to the nearest competitor store;
# - **CompetitionOpenSince[Month/Year]** - gives the approximate year and month of the time the nearest competitor was opened;
# - **Promo** - indicates whether a store is running a promo on that day;
# - **Promo2** - Promo2 is a continuing and consecutive promotion for some stores: 0 = store is not participating, 1 = store is participating;
# - **Promo2Since[Year/Week]** - describes the year and calendar week when the store started participating in Promo2;
# - **PromoInterval** - describes the consecutive intervals Promo2 is started, naming the months the promotion is started anew. E.g. "Feb,May,Aug,Nov" means each round starts in February, May, August, November of any given year for that store.
# + [markdown] heading_collapsed=true
# # <font color='#3F0094'> 1. Descriptive Data Analysis </font>
# + [markdown] hidden=true
# ## <font color='#200CF'> 1.0. Dataframe in Progress Backup </font>
# + hidden=true
df1 = df_raw.copy()
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 1.1. Column Renaming </font>
# + hidden=true
df1.columns
# + hidden=true
# renaming df1 column names
snake_case = lambda x: inflection.underscore(x)
df1.columns = list(map(snake_case, df1.columns))
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 1.2. Data Dimension </font>
# + hidden=true
print(f'Store Dataframe - Number of Rows: {df1.shape[0]}. \nStore Dataframe - Number of Columns: {df1.shape[1]}.')
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 1.3. Data Types </font>
# + hidden=true
# dataframe data types
df1.dtypes
# + hidden=true
# setting date column as datetime type
df1['date'] = pd.to_datetime(df1['date'])
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 1.4. NA Check </font>
# + hidden=true
# checking NA - All NA values came from store.csv
df1.isna().sum()
# + hidden=true
# checking NA using info()
df1.info()
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 1.5. Filling in Missing/Null Values </font>
# + [markdown] hidden=true
# **Number of NA Values**
#
# competition_distance 2642
#
# competition_open_since_month 323348
# competition_open_since_year 323348
#
# promo2_since_week 508031
# promo2_since_year 508031
# promo_interval 508031
# + hidden=true
# competition_distance
# maximun distance x 2
max_dist_x_2 = df1['competition_distance'].max() * 2
# assuming competitors are twice as far away as the greatest distance found
df1['competition_distance'] = df1['competition_distance'].apply(lambda x: max_dist_x_2 if np.isnan(x) else x)
# competition_open_since_year
# frequency per year of existing competition_open_since_year data
frequency = df1['competition_open_since_year'].value_counts(
normalize=True).reset_index().rename(
columns={'index': 'year',
'competition_open_since_year': 'percent'})
# True/False missing/Null Series
missing = df1['competition_open_since_year'].isna()
# Using Numpy's random.choice to fill out missing data based on the frequency of existing info
df1.loc[missing,'competition_open_since_year'] = np.random.choice(frequency.year,
size=len(df1[missing]),
p=frequency.percent)
# competition_open_since_month
# frequency per month of existing competition_open_since_month data
frequency = df1['competition_open_since_month'].value_counts(
normalize=True).reset_index().rename(
columns={'index': 'month',
'competition_open_since_month': 'percent'})
# True/False missing/Null Series
missing = df1['competition_open_since_month'].isna()
# Using Numpy's random.choice to fill out missing data based on the frequency of existing info
df1.loc[missing,'competition_open_since_month'] = np.random.choice(frequency.month,
size=len(df1[missing]),
p=frequency.percent)
# promo2_since_week AND promo2_since_year
# the same date of sale will be used as a reference to fill in the NA values
# then a new timedelta column will be created (promo2_duration)
#promo2_since_week
df1['promo2_since_week'] = df1[['date', 'promo2_since_week']].apply(lambda x: x['date'].week if np.isnan(x['promo2_since_week']) else x['promo2_since_week'], axis=1)
# promo2_since_year
df1['promo2_since_year'] = df1[['date', 'promo2_since_year']].apply(lambda x: x['date'].year if np.isnan(x['promo2_since_year']) else x['promo2_since_year'], axis=1)
# promo_interval
# filling in NA with 'none'
df1['promo_interval'].fillna(value='none', inplace=True)
# creating a column with current month
df1['curr_month'] = df1['date'].dt.strftime('%b')
# creating a column to indicate whether promo2 is active
df1['promo2_active'] = df1.apply(lambda x: 1 if ((
x['curr_month'] in x['promo_interval'].split(',')) and (
x['date'] >= datetime.fromisocalendar(int(x['promo2_since_year']), int(x['promo2_since_week']), 1)) and (
x['promo'] == 1)) else 0, axis=1)
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 1.6. Changing Data Types </font>
# + hidden=true
df1.dtypes
# + hidden=true
# Changing DTypes from float to integer
df1['competition_distance'] = df1['competition_distance'].astype(int)
df1['competition_open_since_month'] = df1['competition_open_since_month'].astype(int)
df1['competition_open_since_year'] = df1['competition_open_since_year'].astype(int)
df1['promo2_since_week'] = df1['promo2_since_week'].astype(int)
df1['promo2_since_year'] = df1['promo2_since_year'].astype(int)
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 1.7. Descriptive Statistics </font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 1.7.0. Numeric vs Categorical - Attributes Split </font>
# + hidden=true
# dataframe - numeric attributes
df_numeric = df1.select_dtypes(include=['int64', 'float64'])
# dataframe - categorical attributes
df_categorical = df1.select_dtypes(exclude=['int64', 'float64', 'datetime64[ns]'])
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 1.7.1. Numeric Attributes </font>
# + hidden=true
# using DF describe() method
df1.describe().T
# + hidden=true
# central tendency metrics - mean, median
ct_mean = df_numeric.apply(np.mean)
ct_median = df_numeric.apply(np.median)
# dispersion metrics - std, min, max, range, skew, kurtosis
d_std = df_numeric.apply(np.std)
d_min = df_numeric.apply(min)
d_max = df_numeric.apply(max)
d_range = df_numeric.apply(lambda x: x.max() - x.min())
d_skew = df_numeric.apply(lambda x: x.skew())
d_kurtosis = df_numeric.apply(lambda x: x.kurtosis())
metrics = pd.DataFrame({
'min': d_min, 'max': d_max, 'range': d_range, 'mean': ct_mean,
'median': ct_median, 'std': d_std, 'skew': d_skew, 'kurtosis': d_kurtosis
})
metrics
# + [markdown] hidden=true
# **competition_distance**
# - Skew: highly skewed data, high positive value means that the right-hand tail is much longer than the left-hand tail.
# - Kurtosis: increases as the tails become heavier, the high positive value indicates a very peaked curve.
#
# **competition_open_since_year**
# - Skew: highly skewed data, high negative value means that the left-hand tail is longer than the right-hand tail.
# - Kurtosis: increases as the tails become heavier, the high positive value indicates a very peaked curve.
#
# **sales**
# - Skewness is close to zero, indicating that the data is not too skewed
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 1.7.2. Categorical Attributes </font>
# + hidden=true
# verifying unique valuesfor each categorical attribute
df_categorical.apply(lambda x: len(x.unique()))
# + [markdown] hidden=true
# **BOXPLOT OF CATEGORICAL ATTRIBUTES**
# + hidden=true hide_input=true
# Boxplot - Categorical Attributes
# not considering when: sales = 0
aux = df1[df1['sales'] > 0]
plt.figure(figsize=(24,10))
plt.subplot(1, 3, 1)
ax1 = sns.boxplot(x='state_holiday', y='sales', data=aux)
ax1.set_title('Boxplot - state_holiday', fontsize=18, pad=10)
ax1.set_xticklabels(labels=['None', 'Public', 'Easter', 'Christmas'])
plt.subplot(1, 3, 2)
ax2 = sns.boxplot(x='store_type', y='sales', data=aux)
ax2.set_title('Boxplot - store_type', fontsize=18, pad=10)
plt.subplot(1, 3, 3)
ax3 = sns.boxplot(x='assortment', y='sales', data=aux)
ax3.set_title('Boxplot - assortment', fontsize=18, pad=10)
plt.show()
# + [markdown] hidden=true
# **BOXPLOT OF BINARY CATEGORICAL ATTRIBUTES**
# + hidden=true hide_input=true
# Boxplot - Binary ategorical Attributes
plt.figure(figsize=(24,10))
plt.subplot(1, 3, 1)
ax1 = sns.boxplot(x='promo', y='sales', data=df1)
ax1.set_title('Boxplot - promo', fontsize=18, pad=10)
plt.subplot(1, 3, 2)
ax2 = sns.boxplot(x='promo2_active', y='sales', data=df1)
ax2.set_title('Boxplot - promo2_active', fontsize=18, pad=10)
plt.subplot(1, 3, 3)
ax3 = sns.boxplot(x='school_holiday', y='sales', data=df1)
ax3.set_title('Boxplot - school_holiday', fontsize=18, pad=10)
plt.show()
# + [markdown] heading_collapsed=true
# # <font color='#3F0094'> 2. Feature Egineering </font>
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 2.0. Dataframe in Progress Backup </font>
# + hidden=true
df2 = df1.copy()
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 2.1. Mind Map </font>
# + hidden=true
# made on coggle.it
Image('../img/mind_map01.png')
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 2.2. Hypothesis </font>
# + [markdown] hidden=true
# **Based on Descriptive Statistics and on Mind Map**
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 2.2.1. Store-related Hypothesis </font>
# + [markdown] hidden=true
#
# **H1.** The larger the assortment the greater the global sales of the stores should be.
#
# **H2.** The median sales of stores with the largest assortment should be the highest.
#
# **H3.** The volume of sales varies according to the type of the store.
#
# **H4.** The average value of sales for a specific type of store is higher than the average value for other types (store types: a, b, c, d).
#
# **H5.** The sales revenue of stores are lower the closer the competitors are located.
#
# **H6.** The average sales value of stores with competitors whose distance is less than 1000 meters is lower than or equal to the average value of other stores.
#
# **H7.** The total sales revenue of stores with competitors for a longer time should be higher.
#
# **H8.** The average sales values of stores whose competitors opened less than 18 months ago is lower than the average values of other stores.
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 2.2.2. Product-related Hypothesis </font>
# + [markdown] hidden=true
# **H9.** The sales revenue should be greater when running a promotion (promo) than when not running a promo.
#
# **H10.** The average sales value of stores should be greater when running a promotion (promo) than when not running a promo.
#
# **H11.** The average sales value of stores with continuing and consecutive promotion (promo2) should be greater than those that do not have extended promotion.
#
# **H12.** The sales revenue of stores running promo2 should grow over time.
#
# **H13.** The median sales value of stores running promo2 for a longer period of time (more than 12 months) should be higher than stores running promo2 for a shorter period of time.
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 2.2.3. Time-related Hypothesis </font>
# + [markdown] hidden=true
# **H14.** The average ticket per customer should be lower during holiday periods.
#
# **H15.** Sales during the Christmas holiday are expected to be the biggest of the year. .
#
# **H16.** Stores affected by the closure of public schools on school holidays should sell less, on average.
#
# **H17.** The revenue in the last quarter of the year should be higher than in the other quarters.
#
# **H18.** Sales behavior is not constant throughout the month, it should be higher in the first 7 days and decreases in the following weeks.
#
# **H19.** Average sales during the weekend should be lower than during the rest of the week.
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 2.3. Feature Engineering </font>
# + hidden=true
# year
df2['year'] = df2['date'].dt.year
# month
df2['month'] = df2['date'].dt.month
# day
df2['day'] = df2['date'].dt.day
# week_of_year
df2['week_of_year'] = df2['date'].dt.isocalendar().week.astype('int64')
# year_week
df2['year_week'] = df2['date'].dt.strftime('%Y-%V')
# competition_months_old
# calculating the competition period, extracting the days and dividing by 30 to get the period in months
df2['competition_months_old'] = df2.apply(lambda x: (
x['date'] - datetime(year=x['competition_open_since_year'],
month=x['competition_open_since_month'],
day=1)).days / 30, axis=1).astype(int)
# assigning zero to negative values of competition_months_old
# in this case it makes no sense to work with the time that is left for the competitor to open
df2.loc[df2['competition_months_old'] < 0, 'competition_months_old'] = 0
# promo2_months_old
# calculation method: zero(0) if promo2 is zero(0) else (actual_date - promo2_starting_date) >> timedelta format
# >> then use .days and divide by 30 to extract the number of months >> as integer
df2['promo2_months_old'] = df2.apply(lambda x: 0 if x['promo2'] == 0 else (
x['date'] - datetime.fromisocalendar(x['promo2_since_year'],
x['promo2_since_week'],
1)).days / 30, axis=1).astype(int)
# assigning zero to negative values of promo2_months_old
# since the store is not yet participating (but will in the future)
df2.loc[df2['promo2_months_old'] < 0, 'promo2_months_old'] = 0
# assortment
df2['assortment'] = df2['assortment'].map({'a': 'basic', 'b': 'extra', 'c': 'extended'})
# state_holiday
df2['state_holiday'] = df2['state_holiday'].map({'0': 'none', 'a': 'public', 'b': 'easter', 'c': 'christmas'})
# =-=-=-=- WARNING: EDA USE ONLY -=-=-=-=
# customer_avg_ticket
df2['customers_avg_ticket'] = (df2['sales'] / df2['customers'])
df2['customers_avg_ticket'].fillna(value=0, inplace=True)
# + [markdown] heading_collapsed=true
# # <font color='#3F0094'> 3. Feature Filtering </font>
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 3.0. Dataframe in Progress Backup </font>
# + hidden=true
df3 = df2.copy()
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 3.1. Filtering Rows </font>
# + hidden=true
# eliminating all records where stores are closed and sales are zero
df3 = df3[(df3['open'] != 0) & (df3['sales'] > 0)]
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 3.2. Filtering Columns </font>
# + [markdown] hidden=true
# **customers:** the number of customers will not be available to be used in the model prediction, as it is an unknown and variable value in the future.
#
# **open:** column has record 1 only.
#
# **promo_interval, curr_month:** auxiliary columns already used in the feature engineering step.
#
# **Important Warning:** column **customers_avg_ticket** will only be used during EDA and will be discarded later.
# + hidden=true
# list of columns to be droped
cols_drop = ['customers', 'open', 'promo_interval', 'curr_month']
df3.drop(cols_drop, axis=1, inplace=True)
# + hidden=true
df3.shape
# + [markdown] heading_collapsed=true
# # <font color='#3F0094'> 4. Exploratory Data Analysis </font>
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 4.0. Dataframe in Progress Backup </font>
# + hidden=true
# dataframe copy
df4 = df3.copy()
# + code_folding=[] hidden=true
# dataframe - numeric attributes - binary attributes droped
df_numeric = df4.select_dtypes(include=['int64', 'float64'])
# dataframe - categorical attributes
df_categorical = df4.select_dtypes(exclude=['int64', 'float64', 'datetime64[ns]']).drop('year_week', axis=1)
# dataframe - categorical attributes + binary variables
df_cat_n_bin = df_categorical.join(df4[['promo', 'promo2', 'school_holiday']], how='left')
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 4.1. Univariate Analysis </font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 4.1.1. Response Variable </font>
# + hidden=true hide_input=false
# sales histogram
plt.figure()
ax = sns.histplot(data=df4, x='sales', stat='proportion', bins=100, kde=True)
y_min, y_max = ax.get_ylim()
ax.figure.set_size_inches(17, 7)
ax.set_title('Sales Histogram', fontsize=20, pad=10)
median = np.median(df_numeric['sales'])
ax.vlines(x=median, ymin=0, ymax=y_max*0.9, linestyles='dashed', label='median', colors='firebrick')
ax.annotate(f'median = {median}', xy=(median*1.15, y_max*0.8), fontsize=14, color='firebrick')
plt.savefig('../img/univar_analysis/sales_histogram.png')
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 4.1.2. Numeric Variable </font>
# + hidden=true
df_numeric.shape
# + [markdown] hidden=true
# **NUMERIC VARIABLES HISTOGRAMS**
# + hidden=true hide_input=true
# ploting numeric attributes histograms
axes = list()
n_bins = 50
n=0
fig, axes = plt.subplots(nrows=4, ncols=4)
fig.set_size_inches(25, 25)
for i in range(4):
for j in range(4):
if n < 15:
axes[i][j].hist(df_numeric.iloc[:, n], bins=n_bins)
axes[i][j].set_title(df_numeric.iloc[:, n].name)
n += 1
# plt.savefig('../img/univar_analysis/numeric_attr_histograms.png')
fig;
# + code_folding=[] hidden=true hide_input=true
# competition_distance
plt.figure(figsize=(20,10))
plt.suptitle('Competitor Distance Analysis', fontsize=22)
plt.subplot(1, 2, 1)
ax1 = sns.histplot(data=df4, x='competition_distance', bins=100)
ax1.set_title("Histogram", fontsize=18, pad=10)
# cumulative counts as bins increase.
plt.subplot(1, 2, 2)
ax2 = sns.histplot(data=df4, x='competition_distance', bins=100, cumulative=True)
ax2.set_title("Cumulative Histogram", fontsize=18, pad=10)
# plt.savefig('../img/univar_analysis/competitor_distance.png')
plt.show()
# + code_folding=[] hidden=true hide_input=true
# competition_open_since_year -- competition_months_old
plt.figure(figsize=(20,20))
plt.suptitle('Competition Over Time', fontsize=22)
# analysing values between 1985 and present day (30 years)
plt.subplot(2, 2, 1)
ax1 = sns.histplot(data=df4.query("competition_open_since_year > 1985"), x='competition_open_since_year', bins=30)
ax1.set_title("Histogram of years when competitors opened", fontsize=18, pad=10)
ax1.set_xlabel("")
plt.subplot(2, 2, 2)
ax1 = sns.histplot(data=df4.query("competition_open_since_year > 1985"), x='competition_open_since_year', bins=30, cumulative=True)
ax1.set_title("Histogram of years when competitors opened\nCumulative", fontsize=18, pad=10)
ax1.set_xlabel("")
# analysing values greater than 0 and lower than 360 (30 years)
plt.subplot(2, 2, 3)
ax2 = sns.histplot(data=df4.query("competition_months_old > 0 and competition_months_old < 360"), x='competition_months_old', bins=30)
ax2.set_title("Histogram of time elapsed since \ncompetitors' inauguration (in months)", fontsize=18, pad=10)
ax2.set_xlabel("")
plt.subplot(2, 2, 4)
ax2 = sns.histplot(data=df4.query("competition_months_old > 0 and competition_months_old < 360"), x='competition_months_old', bins=30, cumulative=True)
ax2.set_title("Histogram of time elapsed since competitors' \ninauguration (in months) - Cumulative", fontsize=18, pad=10)
ax2.set_xlabel("")
# plt.savefig('../img/univar_analysis/competition_time.png')
plt.show()
# + code_folding=[] hidden=true hide_input=true
# promo2_since_year -- promo2_months_old
plt.figure(figsize=(20,20))
plt.suptitle('Extended Promotion Analysis', fontsize=22)
#
plt.subplot(2, 2, 1)
ax1 = sns.histplot(data=df4, x='promo2_since_year', bins = 7)
ax1.set_title("Histogram of years when extended promo started", fontsize=18, pad=10)
ax1.set_xlabel("")
plt.subplot(2, 2, 2)
ax1 = sns.histplot(data=df4, x='promo2_since_year', bins=50, cumulative=True)
ax1.set_title("Histogram of years when extended promo started \nCumulative", fontsize=18, pad=10)
ax1.set_xlabel("")
# analysing values greater than zero
plt.subplot(2, 2, 3)
ax2 = sns.histplot(data=df4.query("promo2_months_old > 0"), x='promo2_months_old', bins=14)
ax2.set_title("Histogram of time elapsed since \nextended promo started (in months)", fontsize=18, pad=10)
ax2.set_xlabel("")
ax2.set_xticks(ticks=np.arange(0, 72, 6))
plt.subplot(2, 2, 4)
ax2 = sns.histplot(data=df4.query("promo2_months_old > 0"), x='promo2_months_old', bins=14, cumulative=True)
ax2.set_title("Histogram of time elapsed since extended \npromo started (in months) - Cumulative", fontsize=18, pad=10)
ax2.set_xlabel("")
ax2.set_xticks(ticks=np.arange(0, 72, 6))
# plt.savefig('../img/univar_analysis/promo2_time.png')
plt.show()
# + code_folding=[] hidden=true hide_input=true
# histograms - customers_avg_ticket AND sales
plt.figure(figsize=(20, 16))
plt.subplot(2, 1, 1)
ax1 = sns.histplot(data=df4, x='customers_avg_ticket', stat='proportion', bins=100, kde=True)
ax1.set_title('Customer Average Ticket Histogram', fontsize=20, pad=15)
ax1.set_xlabel('')
ax1.set_xlim(left=0)
median1 = np.median(df4['customers_avg_ticket'])
_, y1_max = ax1.get_ylim()
ax1.vlines(x=median1, ymin=0, ymax=y1_max*0.9, linestyles='dashed', label='median', colors='firebrick')
ax1.annotate(f'median = $ {median1} / customer', xy=(median1*1.15, y1_max*0.8), fontsize=14, color='firebrick')
plt.subplot(2, 1, 2)
ax2 = sns.histplot(data=df4, x='sales', stat='proportion', bins=100, kde=True)
ax2.set_title('Sales Histogram', fontsize=20, pad=10)
ax2.set_xlim(left=0)
median2 = np.median(df4['sales'])
_, y2_max = ax2.get_ylim()
ax2.vlines(x=median2, ymin=0, ymax=y2_max*0.9, linestyles='dashed', label='median', colors='firebrick')
ax2.annotate(f'median = {median2}', xy=(median2*1.15, y2_max*0.8), fontsize=14, color='firebrick')
# plt.savefig('../img/univar_analysis/customer_ticket_histogram.png')
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 4.1.3. Categorical Variable </font>
# + [markdown] hidden=true
# **STATE HOLIDAY**
# + hidden=true hide_input=true
df4.query("state_holiday != 'none'").value_counts(subset='state_holiday')
# + hidden=true hide_input=true
# state_holiday
# not considering regular day -> state_holiday == 'none'
plt.figure(figsize=(20,10))
plt.subplot(1, 2, 1)
ax1 = sns.countplot(x='state_holiday', data=df4.query("state_holiday != 'none'"))
# ax.figure.set_size_inches(10, 10)
ax1.set_title('Countplot: State Holiday', fontsize=20, pad=10)
ax1.set_xlabel('')
plt.subplot(1, 2, 2)
ax2 = sns.histplot(x='sales', data=df4.query("state_holiday != 'none'"), hue='state_holiday', kde=True)
ax2.set_title('Sales Histogram \nAccording to State Holiday', fontsize=20, pad=10)
ax2.set_xlabel('')
plt.show()
# + [markdown] hidden=true
# **STORE TYPES**
# + hidden=true hide_input=true
df4.value_counts(subset='store_type')
# + hidden=true hide_input=true
# store_type
plt.figure(figsize=(20,10))
plt.subplot(1, 2, 1)
ax1 = sns.countplot(x='store_type', data=df4, order=['a','b','c','d'])
# ax.figure.set_size_inches(10, 10)
ax1.set_title('Countplot: Store Types', fontsize=20, pad=10)
ax1.set_xlabel('')
plt.subplot(1, 2, 2)
ax2 = sns.histplot(x='sales', data=df4, hue='store_type', stat='percent', bins=50, hue_order=['a','b','c','d'], kde=True)
ax2.set_title('Sales Histogram \nAccording to Store Types', fontsize=20, pad=10)
ax2.set_xlabel('')
plt.show()
# + [markdown] hidden=true
# **ASSORTMENT**
# + hidden=true hide_input=true
df4['assortment'].value_counts()
# + hidden=true hide_input=true
# assortment
plt.figure(figsize=(20,10))
plt.subplot(1, 2, 1)
ax1 = sns.countplot(x='assortment', data=df4, order=['basic','extended','extra'])
# ax.figure.set_size_inches(10, 10)
ax1.set_title('Countplot: Assortment Level', fontsize=20, pad=10)
ax1.set_xlabel('')
plt.subplot(1, 2, 2)
ax2 = sns.histplot(x='sales', data=df4, hue='assortment', stat='percent', bins=50, hue_order=['basic','extended','extra'], kde=True)
ax2.set_title('Sales Histogram \nAccording to Assortment Level', fontsize=20, pad=10)
ax2.set_xlabel('')
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 4.2. Bivariate Analysis </font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H1. </font>
#
# **The larger the assortment the greater the global sales of the stores should be.**
# + hidden=true
# group by assortment then sum the sales
aux1 = df4[['assortment', 'sales']].groupby('assortment').sum().reset_index()
# group by year-weak an by assortment then sum the sales
aux2 = df4[['year_week', 'assortment', 'sales']].groupby(['year_week', 'assortment']).sum().reset_index()
# pivoting - each year-week in a row and differents assortments in the columns
aux2 = aux2.pivot(index='year_week', columns='assortment', values='sales')
plt.figure(figsize=(22, 18))
plt.suptitle('Global Sales Analysis by Assortment')
plt.subplot(2, 2, 1)
sns.barplot(x='assortment', y='sales', data=aux1)
plt.xlabel('')
plt.ylabel('Sales Revenue')
plt.subplot(2, 2, 2)
sns.lineplot(data=aux2)
plt.xticks(ticks=[10,34,58,82,106,130], fontsize=12)
plt.xlabel('Year-Week', fontsize=15)
plt.subplot(2, 2, 3)
sns.lineplot(data=aux2[['basic', 'extended']])
plt.xticks(ticks=[10,34,58,82,106,130], fontsize=12)
plt.xlabel('Year-Week', fontsize=15)
plt.ylabel('Sales Revenue')
plt.subplot(2, 2, 4)
sns.lineplot(data=aux2[['extra']])
plt.xticks(ticks=[10,34,58,82,106,130], fontsize=12)
plt.xlabel('Year-Week', fontsize=15)
# plt.savefig('../img/bivar_analysis/assortment_global_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**The number of stores with 'basic' and 'extended' assortment level is much higher (roughly fifty times greater) than the number of stores with 'extra' assortment level, so the sales volume of 'extra' assortment level stores is much smaller when compared to the other types of stores.**</font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H2. </font>
#
# **The median sales of stores with the largest assortment should be the highest.**
# + hidden=true
aux1 = df4[['assortment', 'sales']].groupby('assortment').aggregate(func=['count', 'sum', 'median']).droplevel(level=0, axis='columns')
aux1
# + hidden=true
# median sales by assortment - bar plot
aux1 = df4[['assortment', 'sales']].groupby('assortment').aggregate(func=['count', 'sum', 'median']).droplevel(level=0, axis='columns')
plt.figure(figsize=(18, 9))
plt.title('Medain Value of Sales by Assortment', fontsize=22)
sns.barplot(x=aux1.index, y='median', data=aux1)
plt.xlabel('')
plt.ylabel('Median Sales Value', fontsize=16)
# plt.savefig('../img/bivar_analysis/assortment_median_sales.png')
plt.show()
# + [markdown] hidden=true
# **Although the total number of sales of stores with the 'extra' assortment is much smaller, the median sales value of these stores is higher than the median sales value of the other stores.**
# + hidden=true
# The bootstrap (sampling with replacement from a data set) is a powerful
# tool for assessing the variability of a sample statistic.
# using to calculate the confidence interval for the median sales value,
# according to the store assortment level, with a confidence level of 95%.
# selecting all sales revenue according to the assortment level
sales_basic_assort = df4.loc[df4['assortment'] == 'basic', 'sales']
sales_extended_assort = df4.loc[df4['assortment'] == 'extended', 'sales']
sales_extra_assort = df4.loc[df4['assortment'] == 'extra', 'sales']
# bootstrapp each series of values, take a sample of 500 values,
# caluculate its median and repeat the process 500 times
boot_basic = bootstrap(sales_basic_assort, estimate = 'median', n_repeat=500, n_sample=500, ci=99)
boot_extended = bootstrap(sales_extended_assort, estimate = 'median', n_repeat=500, n_sample=500, ci=99)
boot_extra = bootstrap(sales_extra_assort, estimate = 'median', n_repeat=500, n_sample=500, ci=99)
assortment_bootstrap_statistics = pd.DataFrame([boot_basic, boot_extended, boot_extra],
columns = ['statistic_mean', 'standard_error', 'lower_ci', 'upper_ci'],
index = ['basic', 'extended', 'extra'])
assortment_bootstrap_statistics
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H3. </font>
#
# **The volume of sales varies according to the type of the store.**
# + hidden=true
# group by assortment then sum the sales
aux1 = df4[['store_type', 'sales']].groupby('store_type').sum().reset_index()
aux1['sales_share'] = aux1['sales'] / aux1['sales'].sum()
aux1
# + hidden=true
# group by assortment then sum the sales
aux1 = df4[['store_type', 'sales']].groupby('store_type').sum().reset_index()
# group by year-weak an by assortment then sum the sales
aux2 = df4[['year_week', 'store_type', 'sales']].groupby(['year_week', 'store_type']).sum().reset_index()
# pivoting - each year-week in a row and differents assortments in the columns
aux2 = aux2.pivot(index='year_week', columns='store_type', values='sales')
plt.figure(figsize=(22, 18))
plt.suptitle('Global Sales Analysis by Store Type')
plt.subplot(2, 2, 1)
sns.barplot(x='store_type', y='sales', data=aux1)
plt.xlabel('')
plt.ylabel('Sales Revenue')
plt.subplot(2, 2, 2)
sns.lineplot(data=aux2)
plt.xticks(ticks=[10,34,58,82,106,130], fontsize=12)
plt.xlabel('Year-Week', fontsize=15)
plt.subplot(2, 2, 3)
sns.lineplot(data=aux2[['a', 'd']])
plt.xticks(ticks=[10,34,58,82,106,130], fontsize=12)
plt.xlabel('Year-Week', fontsize=15)
plt.ylabel('Sales Revenue')
plt.subplot(2, 2, 4)
sns.lineplot(data=aux2[['b', 'c']])
plt.xticks(ticks=[10,34,58,82,106,130], fontsize=12)
plt.xlabel('Year-Week', fontsize=15)
# plt.savefig('../img/bivar_analysis/store_type_global_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**Approximately 54% of sales come from type A stores, followed by type 3 stores with 30%, 13% come from type C stores and less than 3% from type B stores.**</font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H4. </font>
#
# **The average value of sales for a specific type of store is higher than the average value for other types (store types: a, b, c, d).**
# + hidden=true
df4[['store_type', 'sales']].groupby('store_type').aggregate(func=['count', 'sum', 'mean']).reset_index()
# + hidden=true
# store-types / assortment - bar plot
# A bar plot represents an estimate of Central Tendency (MEAN) for a numeric variable with the height of
# each rectangle and provides some indication of the uncertainty around that estimate using error bars.
plt.figure(figsize=(18, 9))
plt.title('Average Sales by Store Types', fontsize=22)
# ci -> confidence interval of 95%
sns.barplot(x='store_type', y='sales', order=['a', 'b', 'c', 'd'], data=df4, ci=95)
plt.xlabel('')
plt.ylabel('Average Sales Value')
# plt.savefig('../img/bivar_analysis/store_type_avg_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='red'> **The average sales value of type B stores seems to be considerably greater than the average sales value of the other types of stores.** </font>
#
# <b> Performing a Statistic Test </b>
# + [markdown] hidden=true
# **Hipothesis**: $H_0$ e $H_1$
#
# The null hypothesis always contains an equality claim: equal to; less than or equal to; greater than or equal to. So:
#
# $\mu_1 \Rightarrow$ Average Sales Value of Type B Stores.
#
# $\mu_2 \Rightarrow$ Average Sales Value of Typea A, C or D Stores.
#
# $
# \begin{cases}
# H_0: \mu_1 \leq \mu_2\\
# H_1: \mu_1 > \mu_2
# \end{cases}
# $
# + hidden=true hide_input=false
# using DescrStatsW and CompareMeans from statsmodels
# getting 2000 random sample of sales values for each type of store type
sales_store_type_a = df4.loc[df4['store_type'] == 'a', 'sales'].sample(n=2000)
sales_store_type_b = df4.loc[df4['store_type'] == 'b', 'sales'].sample(n=2000)
sales_store_type_c = df4.loc[df4['store_type'] == 'c', 'sales'].sample(n=2000)
sales_store_type_d = df4.loc[df4['store_type'] == 'd', 'sales'].sample(n=2000)
# calculating statistics with DescrStatsW
stats_a = DescrStatsW(sales_store_type_a)
stats_b = DescrStatsW(sales_store_type_b)
stats_c = DescrStatsW(sales_store_type_c)
stats_d = DescrStatsW(sales_store_type_d)
# using CompareMeans
test_b_a = CompareMeans(stats_b, stats_a)
test_b_c = CompareMeans(stats_b, stats_c)
test_b_d = CompareMeans(stats_b, stats_d)
# performing ztest_ind
# H_null: Average Sales Value of Type B Stores is less than or equal to Types (A, C, D) Stores
z_b_a, pvalue_b_a = test_b_a.ztest_ind(alternative='larger', value=0)
z_b_c, pvalue_b_c = test_b_c.ztest_ind(alternative='larger', value=0)
z_b_d, pvalue_b_d = test_b_d.ztest_ind(alternative='larger', value=0)
pd.DataFrame({
'z': [z_b_a, z_b_c, z_b_d],
'p_value': [round(pvalue_b_a, 6), round(pvalue_b_c, 6), round(pvalue_b_d, 6)],
'H_null_rejected': [pvalue_b_a < 0.05, pvalue_b_c < 0.05, pvalue_b_d < 0.05]},
index=['b_a', 'b_c', 'b_d'])
# + [markdown] hidden=true
# <font color='black'><b> Store_Type per Assortment -vs- Sales </b></font>
# + hidden=true
df4[['store_type', 'assortment', 'sales']].groupby(['store_type', 'assortment']).aggregate(func=['count', 'sum', 'mean']).reset_index()
# + hidden=true
# store-types / assortment - bar plot
# A bar plot represents an estimate of Central Tendency (MEAN) for a numeric variable with the height of
# each rectangle and provides some indication of the uncertainty around that estimate using error bars.
# ci -> confidence interval of 95%
ax = sns.barplot(x='store_type', y='sales', hue='assortment', order=['a', 'b', 'c', 'd'],
hue_order=['basic','extended','extra'], data=df4, ci=95)
ax.figure.set_size_inches(18, 9)
ax.set_title('Average Sales by Store Types and Assortment Level', fontsize=20, pad=10)
ax.set_xlabel('')
ax.set_ylabel('Average Sales Value')
# ax.get_figure().savefig('../img/bivar_analysis/storetype_hue_assortment_avg_sales.png')
ax;
# + [markdown] hidden=true
# **IMPORTANT:** The average sales value of Type B stores stands out even more when the types of stores are separated by assortment types, as can be seen with the average sales of Type B stores with extended assortment levels, it is still more expressive.
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H5. </font>
#
# **The sales revenue of stores are lower the closer the competitors are located.**
# + hidden=true
aux1 = df4[['competition_distance', 'sales']].groupby('competition_distance').sum().reset_index()
bins = list(np.arange(0, 25000, 1000)) + [30000, 40000, 50000, 160000]
aux1['competition_distance_binned'] = pd.cut(x=aux1['competition_distance'], bins=bins)
aux2 = aux1[['competition_distance_binned', 'sales']].groupby('competition_distance_binned').sum().reset_index()
grid = gridspec.GridSpec(2, 2)
plt.figure(figsize=(20,18))
plt.suptitle('Sales Revenue by Competition Distance', fontsize=22)
plt.subplot(grid[0,:])
sns.barplot(x='competition_distance_binned', y='sales', data=aux2)
plt.xlabel('')
plt.ylabel('Sales Revenue')
plt.xticks(fontsize=9, rotation=30)
plt.subplot(grid[1,0])
sns.scatterplot(x='competition_distance', y='sales', data=aux1)
plt.ylabel('Sales Revenue')
plt.xlabel('Distance in Meters')
plt.xticks(fontsize=12)
plt.xlim(-2000, 160000)
plt.subplot(grid[1,1])
sns.heatmap(aux1.corr(method='pearson'), annot=True)
# plt.savefig('../img/bivar_analysis/competition_distance_global_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**In fact, the sum of sales of stores with closer competitors is considerably higher than the sum of sales of stores with more distant competitors, especially for distances above 3000 meters.** </font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H6. </font>
#
# **Regarding the stores with competitors whose distance is less than 1000 meters their average sales value is lower than the average sales value of the other stores.**
# + hidden=true
print(f"The average sales value of stores whose distance is less than 1000 meters: ${(df4.loc[df4['competition_distance'] < 1000, 'sales'].mean()):.2f}.", end='\n\n')
print(f"The average sales value of stores whose distance is greater than 1000 meters: ${(df4.loc[df4['competition_distance'] >= 1000, 'sales'].mean()):.2f}.")
# + [markdown] hidden=true
# <font color='firebrick'><b>In fact, the data shows that the average sales value of stores with competitors that are located less than 1000 meters away is higher than the average sales value of other stores. </b></font>
# + hidden=true
# competition distance avg sales
aux1 = df4[['competition_distance', 'sales']]
aux2 = pd.cut(x=aux1['competition_distance'], bins=[0, 1000, 160000])
aux2.name = 'competition_distance_binned'
aux1 = aux1.join(aux2, how='left')
plt.figure(figsize=(19,8))
plt.title('Average Store Sales by Competition Distance', fontsize=22)
sns.barplot(x='competition_distance_binned', y='sales', data=aux1)
plt.xlabel('Distance in Meters')
plt.ylabel('Average Sales Value')
# plt.savefig('../img/bivar_analysis/competition_distance_avg_sales.png')
plt.show()
# + [markdown] hidden=true
# **STATISTICAL TESTS TO VERIFY IF THE SETS ARE FROM THE SAME DISTRIBUTUION**
#
# **The Wilcoxon rank-sum test tests the null hypothesis that two sets of measurements are drawn from the same distribution.**
#
# **The alternative hypothesis is that values in one sample are more likely to be larger than the values in the other sample.**
#
# If p_value greater than significance level (usually 5%) then the null hypothesis cannot be rejected.
# + [markdown] hidden=true
# **Hipothesis**: $H_0$ e $H_1$
#
# $\mu_1 \Rightarrow$ Average Sales Value of Stores whose competitors distance is less than 1000 meters.
#
# $\mu_2 \Rightarrow$ Average Sales Value of Stores whose competitors distance is grater than 1000 meters.
#
# The null hypothesis always contains an equality claim. So:
# $
# \begin{cases}
# H_0: \mu_1 \leq \mu_2\\
# H_1: \mu_1 > \mu_2
# \end{cases}
# $
# + hidden=true
# unsing ranksums from scipy.stats
# getting 10000 random sample of sales values for each distance
sales_less_1k = df4.loc[df4['competition_distance'] < 1000, 'sales'].sample(10000)
sales_greater_1k = df4.loc[df4['competition_distance'] >= 1000, 'sales'].sample(10000)
statistic, p_value = ranksums(sales_less_1k, sales_greater_1k, alternative='greater')
print(f'p_value: {p_value:.6f}')
# + [markdown] hidden=true
# <font color='firebrick'><b>Once p_value is less than 5% (significance level) it can be said that the sales values of stores with competitors whose distance is less than 1000 meters is, in fact, higher than the sales values of the other stores, just the opposite of the initial assumption set out in Hypotheses f.</b></font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H7. </font>
#
# **The total sales revenue of stores with competitors for a longer time should be higher.**
# + hidden=true
aux1 = df4[(df4['competition_months_old'] > 0) & (df4['competition_months_old'] < 120)][['competition_months_old', 'sales']].groupby('competition_months_old').sum().reset_index()
plt.figure(figsize=(20, 18))
plt.suptitle('Global Sales Analisys by Long Time Competition', fontsize=22)
plt.subplot(2, 1, 1)
sns.barplot(x='competition_months_old', y='sales', data=aux1)
plt.xlabel('Months', fontsize=13)
plt.ylabel('Sales Revenue')
plt.xticks(fontsize=10, rotation=90)
plt.subplot(2, 1, 2)
sns.heatmap(aux1.corr(method='pearson'), annot=True)
# plt.savefig('../img/bivar_analysis/competition_months_global_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**Stores with more recent competitors haver higher total sales values than stores with older competitors.**
#
# **However, it is important to emphasize that there has been a great increase in the opening of competitors in recent years, so more stores have started to have competitors nearby.**
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H8. </font>
#
# ***The average sales values of stores whose competitors opened less than 18 months ago are lower than the average values of other stores.***
# + hidden=true
# competition_months_old
sales_competition_18_less = df4[(df4['competition_months_old'] < 18) & (df4['competition_months_old'] > 0)]['sales']
print(f"The average sales value of stores whose competitors opened less than 18 months ago: ${sales_competition_18_less.mean():.2f}.", end='\n\n')
sales_competition_18_more = df4[df4['competition_months_old'] > 18]['sales']
print(f"The average sales value of stores whose competitors opened more than 18 months ago: ${sales_competition_18_more.mean():.2f}.")
# + hidden=true
# competition_months_old average sales bar plot
aux1 = df4.loc[df4['competition_months_old'] > 0, ['competition_months_old', 'sales']]
aux2 = pd.cut(x=aux1['competition_months_old'], bins=[0, 18, 1410])
aux2.name = 'competition_months_binned'
aux1 = aux1.join(aux2, how='left')
plt.figure(figsize=(19,8))
plt.title('Average Store Sales by Long Time Competition', fontsize=22)
sns.barplot(x='competition_months_binned', y='sales', data=aux1)
plt.xlabel('Time in Months')
plt.ylabel('Average Sales Value')
# plt.savefig('../img/bivar_analysis/competition_months_avg_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**The difference between the averages is less than 3% and the statistical test results state that there is no statistically significant difference between sales of stores with more or less than 18 months of competition.**</font>
#
# **Performing a Boostrap and calculating the confidence interval.**
# + hidden=true
# selecting all sales revenue according to the competition time - greater or less than 18 months
# less than 18 months but greater than zero
sales_competition_18_less = df4[(df4['competition_months_old'] < 18) & (df4['competition_months_old'] > 0)]['sales']
sales_competition_18_more = df4[df4['competition_months_old'] > 18]['sales']
boot_less_18 = bootstrap(sales_competition_18_less, estimate='mean', n_repeat=500, n_sample=1000, ci=95)
boot_more_18 = bootstrap(sales_competition_18_more, estimate='mean', n_repeat=500, n_sample=1000, ci=95)
competition_months_bootstrap_statistics = pd.DataFrame([boot_less_18, boot_more_18],
columns=['statistic_mean', 'standard_error', 'lower_ci', 'upper_ci'],
index=['less_than_18', 'more_than_18'])
competition_months_bootstrap_statistics
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H9. </font>
#
# **The sales revenue should be greater when running a promotion (promo) than when not running a promo.**
# + hidden=true
# total sales by promo
aux1 = df4[['promo', 'sales']].groupby('promo').sum().reset_index()
plt.figure(figsize=(20,10))
plt.title('Global Revenue by Sales Period')
sns.barplot(x='promo', y='sales', data=aux1)
plt.xlabel('')
plt.ylabel('Sales Revenue')
plt.xticks(ticks=[0, 1], labels=['Off Promo', 'On Promo'])
# plt.savefig('../img/bivar_analysis/promo_global_sales.png')
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H10. </font>
#
# **The average sales value of stores should be greater when running a promotion (promo) than when not running a promo.**
# + hidden=true
df4[['promo', 'sales']].groupby('promo').aggregate(func=['count', 'sum', 'mean']).reset_index()
# + [markdown] hidden=true
# <font color='firebrick'>**The average sales value of stores during the period they are on promotion (promo) is considerably higher than the average sales value outside the promotion period.** </font>
# + hidden=true
# promo - bar plot
# A bar plot represents an estimate of Central Tendency (MEAN) for a numeric variable with the height of
# each rectangle and provides some indication of the uncertainty around that estimate using error bars.
# ci -> confidence interval of 95%
ax = sns.barplot(x='promo', y='sales', data=df4, ci=95)
ax.figure.set_size_inches(16, 8)
ax.set_title('Average Sales Value by Sales Period \nPromo', fontsize=20, pad=10)
ax.set_xlabel('')
ax.set_ylabel('Average Sales Value')
ax.set_xticklabels(['Off Promo', 'On Promo'])
# ax.get_figure().savefig('../img/bivar_analysis/promo_avg_sales.png')
ax;
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H11. </font>
#
# **The average sales value of stores with continuing and consecutive promotion (promo2) should be greater than those that do not have extended promotion.**
# + hidden=true
df4[['promo2', 'promo', 'sales']].groupby(['promo2', 'promo']).aggregate(func=['count', 'sum', 'mean']).reset_index()
# + [markdown] hidden=true
# <b>The average sales value of stores that are participating in the extended promotion period is lower than the average sales value of stores that are not participating, whether they have active promotion or not.
#
# It is necessary to identify possible causes for poor sales performance or reassess the marketing strategy for those stores specifically.
#
# However, it should be noted that the average sales value of stores with extended promotion is higher during the promotion period than outside this period.</b>
# + code_folding=[] hidden=true
# promo2 - bar plot
# A bar plot represents an estimate of Central Tendency (MEAN) for a numeric variable with the height of
# each rectangle and provides some indication of the uncertainty around that estimate using error bars.
# ci -> confidence interval of 95%
ax = sns.barplot(x='promo2', y='sales', hue='promo', data=df4, ci=95)
ax.figure.set_size_inches(16, 8)
ax.set_title('Comparison of sales of stores that are participating \n vs. not participating in the extended promotion', fontsize=20, pad=10)
ax.set_xlabel('')
ax.set_ylabel('average sales')
ax.set_xticklabels(['store is not participating', 'store is participating'])
# ax.get_figure().savefig('../img/bivar_analysis/promo2_comparison_avg_sales.png')
ax;
# + hidden=true
# analysing the average sales of promo2 stores only
# comparing the results: promo2-on vs promo2-off
df4.query("promo2 == 1")[['promo2_active', 'sales']].groupby('promo2_active').aggregate(func=['count', 'sum', 'mean']).reset_index()
# + hidden=true
# promo2_active - bar plot
# Analysing stores that participate in Promo2
# comparing the results: promo2-off vs promo2-on
ax = sns.barplot(x='promo2_active', y='sales', data=df4.query("promo2 == 1"), ci=95)
ax.figure.set_size_inches(16, 8)
ax.set_title('Bar Plot: Promo2 \nOff vs On', fontsize=20, pad=10)
ax.set_xlabel('')
ax.set_xticklabels(['not_active', 'active'])
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H12. </font>
#
# **The sales revenue of stores running promo2 should grow over time.**
# + hidden=true
# sales revenue over promo2 time
aux1 = df4.loc[(df4['promo2'] == 1) & (df4['promo2_months_old'] > 12), ['promo2_months_old', 'sales']].groupby('promo2_months_old').sum().reset_index()
plt.figure(figsize=(22,10))
plt.suptitle('Sales Revenue over Promo2 Time', fontsize=22)
plt.subplot(1, 2, 1)
sns.barplot(x='promo2_months_old', y='sales', data=aux1)
plt.xlabel('')
plt.ylabel('Sales Revenue')
plt.xticks(fontsize=9)
plt.subplot(1, 2, 2)
sns.heatmap(data=aux1.corr(method='pearson'), annot=True)
# plt.savefig('../img/bivar_analysis/promo2_global_sales.png')
plt.show()
# + [markdown] hidden=true
#
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H13. </font>
#
# **The median sales value of stores running promo2 for a longer period of time (more than 12 months) should be higher than stores running promo2 for a shorter period of time.**
# + hidden=true
# stores participating in promo 2 for over 12 months
median_sales_promo2_over_12 = df4.loc[(df4['promo2'] == 1) & (df4['promo2_months_old'] > 12), 'sales'].median()
# stores participating in promo 2 for less than 12 months
median_sales_promo2_less_12 = df4.loc[(df4['promo2'] == 1) & (df4['promo2_months_old'] <= 12) & (df4['promo2_months_old'] > 0), 'sales'].median()
print(f'Median sales of stores participating in promo 2 for over 12 months: $ {median_sales_promo2_over_12:.2f}.', end='\n\n')
print(f'Median sales of stores participating in promo 2 for less than 12 months: $ {median_sales_promo2_less_12:.2f}.')
# + hidden=true
aux1 = df4.loc[(df4['promo2'] == 1) & (df4['promo2_months_old'] > 0), ['promo2_months_old', 'sales']]
aux2 = pd.cut(x=aux1['promo2_months_old'], bins=[0, 12, 75])
aux2.name = 'promo2_months_binned'
aux1 = aux1.join(aux2, how='left')
plt.figure(figsize=(20,9))
plt.title('Average Sales Value over Promo2 Long Time', fontsize=20)
sns.barplot(x='promo2_months_binned', y='sales', data=aux1)
plt.xlabel('')
plt.ylabel('Average Sales Value')
plt.xticks(ticks=[0, 1], labels=['Less than 12 months', "Over 12 months"])
# plt.savefig('../img/bivar_analysis/promo2_avg_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**Despite being similar values, the median sales value of stores that have been participating in the promo2 for over 12 months is higher.**</font>
#
#
# **Performing a Boostrap and calculating the confidence interval.**
# + hidden=true
# selecting all sales of stores participating in promo 2 and splitting into greater than or less than 12 months old
sales_promo2_over_12 = df4.loc[(df4['promo2'] == 1) & (df4['promo2_months_old'] > 12), 'sales']
# less than 12 months but greater than zero
sales_promo2_less_12 = df4.loc[(df4['promo2'] == 1) & (df4['promo2_months_old'] <= 12) & (df4['promo2_months_old'] > 0), 'sales']
boot_over_12 = bootstrap(sales_promo2_over_12, estimate='median', n_repeat=500, n_sample=1000, ci=95)
boot_less_12 = bootstrap(sales_promo2_less_12, estimate='median', n_repeat=500, n_sample=1000, ci=95)
promo2_months_bootstrap_statistics = pd.DataFrame([boot_over_12, boot_less_12],
columns=['statistic_mean', 'standard_error', 'lower_ci', 'upper_ci'],
index=['over_12', 'less_than_12'])
promo2_months_bootstrap_statistics
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H14. </font>
#
# **The average ticket per customer should be lower during holiday periods.**
# + hidden=true
# customer average ticket by state holiday
plt.figure(figsize=(19,10))
plt.title('Customer Average Ticket by Holiday/Regular Day', fontsize=20)
sns.barplot(x='state_holiday', y='customers_avg_ticket', data=df4)
plt.xlabel('')
plt.ylabel('Customer Average Ticket')
# plt.savefig('../img/bivar_analysis/customer_avg_ticket_holiday.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**The customer average ticket price is considerably higher in a regular day than during any state holiday.** </font>
# + hidden=true hide_input=true
aux1 = df4[['state_holiday', 'customers_avg_ticket']].groupby('state_holiday').mean().reset_index()
aux1
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H15. </font>
#
# **Sales during the Christmas holiday are expected to be the biggest of the year. .**
# + hidden=true
# sales during holidays
aux1 = df4.loc[df4['state_holiday'] != 'none', ['year', 'state_holiday', 'sales']].groupby(['year', 'state_holiday']).sum().reset_index()
plt.figure(figsize=(20,10))
plt.title('Sales Revenue during State Holidays per Year', fontsize=20)
sns.barplot(x='year', y='sales', hue='state_holiday', data=aux1)
plt.xlabel('')
plt.ylabel('Sales Revenue')
# plt.savefig('../img/bivar_analysis/state_holiday_global_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**Sales during Christmas are lower than during the other State Holidays.**</font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H16. </font>
#
# **Stores affected by the closure of public schools on school holidays should sell less.**
# + hidden=true
# sales vs school holidays
aux1 = df4[['month', 'school_holiday', 'sales']].groupby(['month', 'school_holiday']).sum().reset_index()
plt.figure(figsize=(20,8))
plt.suptitle('How Stores Sales Are \nAfected By School Holiday', fontsize=20)
plt.subplot(1, 2, 1)
ax1 = sns.barplot(x='month', y='sales', hue='school_holiday', data=aux1)
ax1.set_title('Total Sales vs Month', fontsize=16)
ax1.set_xlabel('Month')
ax1.set_ylabel('Sales Revenue')
plt.subplot(1, 2, 2)
ax2 = sns.barplot(x='school_holiday', y='sales', data=df4)
ax2.set_title('Influence of the school holiday on average sales', fontsize=15)
ax2.set_xlabel('')
ax2.set_ylabel('Average Sales')
ax2.set_xticklabels(['regular day', 'school holiday'])
# plt.savefig('../img/bivar_analysis/school_holiday_sales.png')
plt.show()
# + hidden=true hide_input=false
df4[['school_holiday', 'sales']].groupby('school_holiday').mean()
# + [markdown] hidden=true
# <font color='firebrick'>**The difference between the average sales values of the stores is less than 5%.**</font>
#
# **Performing a Boostrap and calculating the confidence interval.**
# + hidden=true
# splitting sales into during school holiday and off school holidays
on_school_holiday = df4.loc[df4['school_holiday'] == 1, 'sales']
# less than 12 months but greater than zero
off_school_holiday = df4.loc[df4['school_holiday'] == 0, 'sales']
boot_on = bootstrap(on_school_holiday, estimate='mean', n_repeat=500, n_sample=1000, ci=95)
boot_off = bootstrap(off_school_holiday, estimate='mean', n_repeat=500, n_sample=1000, ci=95)
school_holiday_bootstrap_statistics = pd.DataFrame([boot_on, boot_off],
columns=['statistic_mean', 'standard_error', 'lower_ci', 'upper_ci'],
index=['school_holiday', 'not_school_holiday'])
school_holiday_bootstrap_statistics
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H17. </font>
#
# **The revenue in the last quarter of the year should be higher than in the other quarters.**
# + hidden=true
# sales revenue over the quarters of the years
# mapping the quarters
quarter_map = {1:1, 2:1, 3:1,
4:2, 5:2, 6:2,
7:3, 8:3, 9:3,
10:4, 11:4, 12:4}
# the sales data of 2015 stops in july - considering 2013 and 2014 only
aux1 = df4.query("year != 2015")[['month', 'sales']].groupby('month').sum().reset_index()
aux1['quarter'] = aux1['month'].map(quarter_map)
aux2 = aux1[['quarter', 'sales']].groupby('quarter').sum().reset_index()
plt.figure(figsize=(20,10))
plt.suptitle('Sales Revenue vs Quarters')
plt.subplot(2, 2, 1)
ax1 = sns.barplot(x='quarter', y='sales', data=aux2)
ax1.set_xlabel('')
#ax3.set_xticklabels(ticks=[0,1,2,3], labels=['1st', '2nd', '3rd', '4th'])
plt.subplot(2, 2, 2)
ax2 = sns.regplot(x='quarter', y='sales', data=aux2)
ax2.set_xlabel('')
plt.subplot(2, 2, 3)
ax3 = sns.barplot(x='month', y='sales', data=aux1)
ax3.set_xlabel('')
plt.subplot(2, 2, 4)
ax4 = sns.heatmap(aux2.corr(method='pearson'), annot=True)
ax4.set_xlabel('')
# plt.savefig('../img/bivar_analysis/quarters_global_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**There is an increase in sales in the last quarter of the year, but the difference is not significant in relation to the other quarters**</font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H18. </font>
#
# **Sales behavior is not constant throughout the month, it should be higher in the first 7 days and decreases in the following weeks.**
# + hidden=true
# Sales Revenue vs Days of Month
aux1 = df4[['day', 'sales']].groupby('day').sum().reset_index()
grids = gridspec.GridSpec(nrows=2, ncols=2)
plt.figure(figsize=(20,16))
plt.suptitle('Sales Revenue vs Days of Month')
plt.subplot(grid[0, 0:])
sns.barplot(x='day', y='sales', data=aux1)
plt.xlabel('')
plt.subplot(grid[1, 0])
sns.regplot(x='day', y='sales', data=aux1)
plt.xlabel('')
plt.subplot(grid[1, 1])
sns.heatmap(aux1.corr(method='pearson'), annot=True)
# plt.savefig('../img/bivar_analysis/day_global_sales.png')
plt.show()
# + [markdown] hidden=true
# <font color='firebrick'>**There is a drop in sales throughout the month.**</font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> Hypothesis H19. </font>
#
# **Average sales in the weekend days should be lower than in the other days of the week.**
# + hidden=true
df4[['day_of_week', 'sales']].groupby('day_of_week').aggregate(func=['count', 'sum', 'mean'])
# + hidden=true
aux1 = df4[['day_of_week', 'sales']].groupby('day_of_week').mean().reset_index()
plt.figure(figsize=(22,9))
plt.suptitle('Average Sales by Weekday/Weekend', fontsize=20)
plt.subplot(1, 3, 1)
sns.barplot(x='day_of_week', y='sales', data=aux1)
plt.xlabel('')
plt.xticks(ticks=[0,1,2,3,4,5,6],
labels=['mon', 'tue', 'wed', 'thu', 'fri', 'sat', 'sun'])
plt.subplot(1, 3, 2)
sns.regplot(x='day_of_week', y='sales', data=aux1)
plt.xlabel('')
plt.subplot(1, 3, 3)
sns.heatmap(aux1.corr(method='pearson'), annot=True)
# plt.savefig('../img/bivar_analysis/weekend_avg_sales.png')
plt.show()
# + hidden=true
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 4.3. Multivariate Analysis </font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 4.3.1. Numeric Attributes </font>
# + hidden=true
correlation = df_numeric.corr(method='pearson')
plt.figure(figsize=(26,18))
plt.title('Numeric Attributes Multivariate Analysis', fontsize=22, pad=10)
sns.heatmap(correlation, annot=True)
# plt.savefig('../img/multivar_analysis/numeric_attributes_multivar_analysis.png')
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 4.3.2. Categorical Attributes </font>
# + [markdown] hidden=true
# https://en.wikipedia.org/wiki/Cram%C3%A9r%27s_V
# + code_folding=[0] hidden=true
# calculating Cramér's V
a1 = cramer_v(df_cat_n_bin['state_holiday'], df_cat_n_bin['state_holiday'])
a2 = cramer_v(df_cat_n_bin['state_holiday'], df_cat_n_bin['store_type'])
a3 = cramer_v(df_cat_n_bin['state_holiday'], df_cat_n_bin['assortment'])
a4 = cramer_v(df_cat_n_bin['state_holiday'], df_cat_n_bin['promo'])
a5 = cramer_v(df_cat_n_bin['state_holiday'], df_cat_n_bin['promo2'])
a6 = cramer_v(df_cat_n_bin['state_holiday'], df_cat_n_bin['school_holiday'])
b1 = cramer_v(df_cat_n_bin['store_type'], df_cat_n_bin['state_holiday'])
b2 = cramer_v(df_cat_n_bin['store_type'], df_cat_n_bin['store_type'])
b3 = cramer_v(df_cat_n_bin['store_type'], df_cat_n_bin['assortment'])
b4 = cramer_v(df_cat_n_bin['store_type'], df_cat_n_bin['promo'])
b5 = cramer_v(df_cat_n_bin['store_type'], df_cat_n_bin['promo2'])
b6 = cramer_v(df_cat_n_bin['store_type'], df_cat_n_bin['school_holiday'])
c1 = cramer_v(df_cat_n_bin['assortment'], df_cat_n_bin['state_holiday'])
c2 = cramer_v(df_cat_n_bin['assortment'], df_cat_n_bin['store_type'])
c3 = cramer_v(df_cat_n_bin['assortment'], df_cat_n_bin['assortment'])
c4 = cramer_v(df_cat_n_bin['assortment'], df_cat_n_bin['promo'])
c5 = cramer_v(df_cat_n_bin['assortment'], df_cat_n_bin['promo2'])
c6 = cramer_v(df_cat_n_bin['assortment'], df_cat_n_bin['school_holiday'])
d1 = cramer_v(df_cat_n_bin['promo'], df_cat_n_bin['state_holiday'])
d2 = cramer_v(df_cat_n_bin['promo'], df_cat_n_bin['store_type'])
d3 = cramer_v(df_cat_n_bin['promo'], df_cat_n_bin['assortment'])
d4 = cramer_v(df_cat_n_bin['promo'], df_cat_n_bin['promo'])
d5 = cramer_v(df_cat_n_bin['promo'], df_cat_n_bin['promo2'])
d6 = cramer_v(df_cat_n_bin['promo'], df_cat_n_bin['school_holiday'])
e1 = cramer_v(df_cat_n_bin['promo2'], df_cat_n_bin['state_holiday'])
e2 = cramer_v(df_cat_n_bin['promo2'], df_cat_n_bin['store_type'])
e3 = cramer_v(df_cat_n_bin['promo2'], df_cat_n_bin['assortment'])
e4 = cramer_v(df_cat_n_bin['promo2'], df_cat_n_bin['promo'])
e5 = cramer_v(df_cat_n_bin['promo2'], df_cat_n_bin['promo2'])
e6 = cramer_v(df_cat_n_bin['promo2'], df_cat_n_bin['school_holiday'])
f1 = cramer_v(df_cat_n_bin['school_holiday'], df_cat_n_bin['state_holiday'])
f2 = cramer_v(df_cat_n_bin['school_holiday'], df_cat_n_bin['store_type'])
f3 = cramer_v(df_cat_n_bin['school_holiday'], df_cat_n_bin['assortment'])
f4 = cramer_v(df_cat_n_bin['school_holiday'], df_cat_n_bin['promo'])
f5 = cramer_v(df_cat_n_bin['school_holiday'], df_cat_n_bin['promo2'])
f6 = cramer_v(df_cat_n_bin['school_holiday'], df_cat_n_bin['school_holiday'])
# creating dataframe with Cramer's V results
df_cramer_v = pd.DataFrame({'state_holiday': [a1, a2, a3, a4, a5, a6],
'store_type': [b1, b2, b3, b4, b5, b6],
'assortment': [c1, c2, c3, c4, c5, c6],
'promo': [d1, d2, d3, d4, d5, d6],
'promo2': [e1, e2, e3, e4, e5, e6],
'school_holiday': [f1, f2, f3, f4, f5, f6]})
# using columns names to set the indexes names
df_cramer_v.set_index(keys=df_cramer_v.columns, drop=False, inplace=True)
# + hidden=true hide_input=true
# heatmap
plt.figure(figsize=(19, 8))
plt.title('Categorical Attributes Heatmap', fontsize=21, pad=10)
sns.heatmap(df_cramer_v, annot=True)
# plt.savefig('../img/multivar_analysis/categorical_attributes_multivar_analysis.png')
plt.show()
# + [markdown] heading_collapsed=true
# # <font color='#3F0094'> 5. Data Preparation </font>
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 5.0. Dataframe Copy for Data Preparation </font>
# + hidden=true
# copying dataframe before filling in null values
# and before feature engineering
df5 = df_raw.copy()
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 5.1. Feature Engineering for ML Models </font>
# + [markdown] hidden=true
# <font color='firebrick'><b>Some features will receive different treatments from those used for Exploratory Data Analysis (EDA) and some feature engineering are not necessary for machine learning models.</b></font>
#
# **Features with different treatment:** competition_open_since_year, competition_open_since_month
#
# **Features not created / engineered:** customers_avg_ticket, year_week / assortment
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 5.1.1. Data Cleaning </font>
# + hidden=true
# renaming df5 column names
snake_case = lambda x: inflection.underscore(x)
df5.columns = list(map(snake_case, df5.columns))
# setting date column as datetime type
df5['date'] = pd.to_datetime(df5['date'])
## =-= Filling in Missing/Null Values =-= ##
## competition_distance - using maximum distance x 2
# maximun distance x 2
max_dist_x_2 = df5['competition_distance'].max() * 2
# assuming competitors are twice as far away as the greatest distance found
df5['competition_distance'] = df5['competition_distance'].apply(lambda x: max_dist_x_2 if np.isnan(x) else x)
## competition_open_since_year
# assign the year of the latest date if NA
df5.loc[df5['competition_open_since_year'].isna(), 'competition_open_since_year'] = df5['date'].max().year
## competition_open_since_month
# assign the month of the latest date if NA
df5.loc[df5['competition_open_since_month'].isna(), 'competition_open_since_month'] = df5['date'].max().month
# promo2_since_week AND promo2_since_year
# in case of NA values the date of sale will be used -- the difference between these dates will be used later
## promo2_since_week
df5['promo2_since_week'] = df5[['date', 'promo2_since_week']].apply(lambda x: x['date'].week if np.isnan(x['promo2_since_week']) else x['promo2_since_week'], axis=1)
## promo2_since_year
df5['promo2_since_year'] = df5[['date', 'promo2_since_year']].apply(lambda x: x['date'].year if np.isnan(x['promo2_since_year']) else x['promo2_since_year'], axis=1)
## promo_interval: used to create a new column -> promo2_active
# filling in NA with 'none'
df5['promo_interval'].fillna(value='none', inplace=True)
# creating a column with current month
df5['curr_month'] = df5['date'].dt.strftime('%b')
## creating a column to indicate whether promo2 is active
df5['promo2_active'] = df5.apply(lambda x: 1 if ((
x['curr_month'] in x['promo_interval'].split(',')) and (
x['date'] >= datetime.fromisocalendar(int(x['promo2_since_year']), int(x['promo2_since_week']), 1)) and (
x['promo'] == 1)) else 0, axis=1)
## =-= Changing Data Types =-= ##
# Changing DTypes from float to integer
df5['competition_distance'] = df5['competition_distance'].astype(int)
df5['competition_open_since_month'] = df5['competition_open_since_month'].astype(int)
df5['competition_open_since_year'] = df5['competition_open_since_year'].astype(int)
df5['promo2_since_week'] = df5['promo2_since_week'].astype(int)
df5['promo2_since_year'] = df5['promo2_since_year'].astype(int)
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 5.1.2. Feature Engineering </font>
# + hidden=true
## =-= Dates =-= ##
# year
df5['year'] = df5['date'].dt.year
# month
df5['month'] = df5['date'].dt.month
# day
df5['day'] = df5['date'].dt.day
# week_of_year
df5['week_of_year'] = df5['date'].dt.isocalendar().week.astype('int64')
# competition_months_old
# calculating the competition period, extracting the days and dividing by 30 to get the period in months
df5['competition_months_old'] = df5.apply(lambda x: (
x['date'] - datetime(year=x['competition_open_since_year'],
month=x['competition_open_since_month'],
day=1)).days / 30, axis=1).astype(int)
# assigning zero to negative values of competition_months_old
# in this case it makes no sense to work with the time that is left for the competitor to open
df5.loc[df5['competition_months_old'] < 0, 'competition_months_old'] = 0
# promo2_months_old
# calculation method: zero(0) if promo2 is zero(0) else (actual_date - promo2_starting_date) >> timedelta format
# >> then use .days and divide by 30 to extract the number of months >> as integer
df5['promo2_months_old'] = df5.apply(lambda x: 0 if x['promo2'] == 0 else (
x['date'] - datetime.fromisocalendar(x['promo2_since_year'],
x['promo2_since_week'],
1)).days / 30, axis=1).astype(int)
# assigning zero to negative values of promo2_months_old
# since the store is not yet participating (but will in the future)
df5.loc[df5['promo2_months_old'] < 0, 'promo2_months_old'] = 0
## =-= Filtering Features =-= ##
# eliminating all records where stores are closed
df5 = df5[(df5['open'] != 0) & (df5['sales'] > 0)]
# list of columns to be droped
cols_drop = ['customers', 'open', 'promo_interval', 'curr_month']
df5.drop(cols_drop, axis=1, inplace=True)
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 5.2. Feature Scaling - Standardization </font>
# + [markdown] hidden=true
# **Also called Z-score normalization. Standardization typically means rescales data to have a mean of 0 and a standard deviation of 1 (unit variance).**
# + [markdown] hidden=true
# <font color='firebrick'>**None of the features behavior is close to a Gaussian (normal) distribution, so standardization is not recommended.**</font>
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 5.3. Feature Scaling - Normalization </font>
# + [markdown] hidden=true
# **Normalization typically means rescales the values into a range of [0, 1] .**
# + [markdown] hidden=true
# **ROBUST SCALER**
#
# Its use is indicated for cases where data have outliers. To overcome this, the median and interquartile range can be used to rescale numeric variables.
# + hidden=true
# rescaling with Robust Scaler
rs = RobustScaler()
# competition_distance
df5['competition_distance'] = rs.fit_transform(df5[['competition_distance']].values)
# pickle.dump(rs, open('../parameters/competition_distance_scaler.pkl', 'wb'))
# competition_months_old
df5['competition_months_old'] = rs.fit_transform(df5[['competition_months_old']].values)
# pickle.dump(rs, open('../parameters/competition_months_old_scaler.pkl', 'wb'))
# + [markdown] hidden=true
# **MIN-MAX SCALER**
# + hidden=true
# rescaling with Min-Max Scaler
mms = MinMaxScaler()
# promo2_months_old
df5['promo2_months_old'] = mms.fit_transform(df5[['promo2_months_old']].values)
# pickle.dump(mms, open('../parameters/promo2_months_old_scaler.pkl', 'wb'))
# year
df5['year'] = mms.fit_transform(df5[['year']].values)
# pickle.dump(mms, open('../parameters/year_scaler.pkl', 'wb'))
# + [markdown] heading_collapsed=true hidden=true
# ## <font color='#200CF'> 5.4. Feature Transformation </font>
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 5.3.1. Encoding </font>
#
# **Enconding: Transforming Categorical Features Into Numeric Features**
# + [markdown] hidden=true
# **ONE HOT ENCODING -- ORDINAL ENCODING -- LABEL ENCODING**
# + hidden=true
# state_holiday - One Hot Encoding
df5 = pd.get_dummies(df5, prefix=['st_hol'], columns=['state_holiday'])
# assortment - Ordinal Encoding
assortment_dict = {'a': 1, 'b': 2, 'c': 3}
df5['assortment'] = df5['assortment'].map(assortment_dict)
# store_type - Label Encoding
le = LabelEncoder()
df5['store_type'] = le.fit_transform(df5['store_type'])
# pickle.dump(le, open('../parameters/store_type_scaler.pkl', 'wb'))
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 5.3.2. Nature Transformation </font>
# + hidden=true
# month
df5['month_sin'] = df5['month'].apply(lambda x: np.sin(x * (2. * np.pi / 12)))
df5['month_cos'] = df5['month'].apply(lambda x: np.cos(x * (2. * np.pi / 12)))
# day
df5['day_sin'] = df5['day'].apply(lambda x: np.sin(x * (2. * np.pi / 30)))
df5['day_cos'] = df5['day'].apply(lambda x: np.cos(x * (2. * np.pi / 30)))
# day_of_week
df5['day_of_week_sin'] = df5['day_of_week'].apply(lambda x: np.sin(x * (2. * np.pi / 7)))
df5['day_of_week_cos'] = df5['day_of_week'].apply(lambda x: np.cos(x * (2. * np.pi / 7)))
# week_of_year
df5['week_of_year_sin'] = df5['week_of_year'].apply(lambda x: np.sin(x * (2. * np.pi / 52)))
df5['week_of_year_cos'] = df5['week_of_year'].apply(lambda x: np.cos(x * (2. * np.pi / 52)))
# + [markdown] heading_collapsed=true hidden=true
# ### <font color='#2365FF'> 5.3.3. Response Variable Tranformation - Log Transform </font>
# + hidden=true
df5['sales'] = np.log1p(df5['sales'])
# -
# # <font color='#3F0094'> 6. Feature Selection </font>
# ## <font color='#200CF'> 6.0. Dataframe in Progress Backup </font>
df6 = df5.copy()
# ## <font color='#200CF'> 6.1. Dataframe Split into Training and Test Dataset </font>
# droping irrelevant and variables that were derived
cols_drop = ['month', 'day', 'day_of_week', 'week_of_year']
df6.drop(labels=cols_drop, axis=1, inplace=True)
# +
# selecting the last 7 months as test dataset and all previous dates as train dataset
X_train = df6[df6['date'] < '2015-01-01']
y_train = X_train['sales']
X_test = df6[df6['date'] >= '2015-01-01']
y_test = X_test['sales']
# -
# ## <font color='#200CF'> 6.2. Boruta as Feature Selector </font>
# +
# train and test dataset for boruta
X_train_n = X_train.drop(labels=['date', 'sales'], axis=1).values
y_train_n = y_train.values.ravel()
# defining RandomForestRegressor
rf = RandomForestRegressor(n_jobs=-1)
# defining BorutaPy
boruta = BorutaPy(rf, n_estimators='auto', verbose=2, random_state=42).fit(X_train_n, y_train_n)
# -
# ### <font color='#2365FF'> 6.2.1. Best Features from Boruta </font>
# +
# all features except date and sales
X_train_fs = X_train.head(1).drop(['date', 'sales'], axis=1)
# features selected by boruta
cols_selected = boruta.support_.tolist()
cols_selected_names = X_train_fs.iloc[:, cols_selected].columns.tolist()
print(f"List of columns selected by Boruta:\n{', '.join(cols_selected_names)}.")
# features not selected by boruta
cols_rejected_boruta = list(np.setdiff1d(X_train_fs.columns, cols_selected_names))
print(f"\nList of columns rejected by Boruta:\n{', '.join(cols_rejected_boruta)}.")
# -
# ## <font color='#200CF'> 6.3. Feature Selection - Final Decision </font>
# +
# using boruta feature selection + adding month_sin
selected_features = [
'store', 'promo', 'store_type', 'assortment', 'competition_distance', 'competition_open_since_month',
'competition_open_since_year', 'promo2', 'promo2_since_week', 'promo2_since_year', 'competition_months_old',
'promo2_months_old', 'month_sin', 'month_cos', 'day_sin', 'day_cos', 'day_of_week_sin', 'day_of_week_cos',
'week_of_year_sin', 'week_of_year_cos']
# inserting 'date' and 'sales' back to the features list
selected_features.extend(['date', 'sales'])
| notebooks/stg06_feature_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: text_data
# language: python
# name: text_data
# ---
# https://towardsdatascience.com/end-to-end-topic-modeling-in-python-latent-dirichlet-allocation-lda-35ce4ed6b3e0
# https://www.machinelearningplus.com/nlp/topic-modeling-python-sklearn-examples/
# 1) Describe how topic models define a document. Why is this a useful framework? What substantive questions might it answer? Why isn’t it useful and which questions might it be ill-suited to?
#
# LDA assumes, like humans assume, that words carry strong semantic information and similar documents will use similar words. Secondly, documents are pobability distributions over latent topics and topics are probability distributions over words.
#
# An important distinction to other models is that LDA works with probability distributions not strict word-frequencies.
#
# Topic models define a document as a mixture of a small number of topics and its words atrributed to one of the topics. This framework is useful as it allows, for example, genitists to study the genome in an applied way. Another example, engineers, may classify documents and approximate their relation to other topics.
#
# With plate notation, let's describe the below image:
# - $K$ number of topics
# - $M$ number of documents
# - $N$ number of words in a given document
# - β parameter of Dirichlet prior on per-topic word distro
# - $\Phi$ word distribution for topic $K$ (sums to 1)
# - $a$ parameter of Dirichlet prior on per-document topic distro
# - $\Theta$ topic distro for document
# - $z$ topic word in document
# - $w$ specific word
#
# "What substantive questions might it answer?"
# Topic models are able to derive 'topics' from uncategorized text, without a human having to read it!
#
# "Why isn't it useful and which questions might it be ill-suit to?"
# If your corpus is constantly changing, the sensitive hyper-parameters will need maintenance. If not maintained unintented drift will occur and the derived topics will not be useful.
# <img src="../images/lda_k.png">
# Source: https://en.wikipedia.org/wiki/Latent_Dirichlet_allocation
# # Setup
# The below notebook was built surrounding this tutorial:
# https://www.machinelearningplus.com/nlp/topic-modeling-python-sklearn-examples
# +
# import packages
import numpy as np
import pandas as pd
import seaborn as sns
import re, nltk, spacy, gensim
# sklearn
from sklearn.cluster import KMeans
from sklearn.model_selection import GridSearchCV
from sklearn.metrics.pairwise import euclidean_distances
from sklearn.decomposition import LatentDirichletAllocation, TruncatedSVD
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
# misc
from pprint import pprint
# plotting tools
import pyLDAvis
import pyLDAvis.sklearn
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# # Load Data
# load data from processed dir
df_game_reviews = pd.read_csv(r'../data/processed/game_reviews_processed.csv')
# +
# clean
# remove all data colums aside from recommendationid, review, and timestamp_created.
def prune_cols(df):
# keep only needed cols and subset english only
global df_game_reviews
df_game_reviews = df_game_reviews.loc[df_game_reviews['language'] == 'english'] # subsets for english
df_game_reviews = df_game_reviews[["recommendationid", "review", "timestamp_created"]] # dops unnecessary cols
return df_game_reviews
prune_cols(df_game_reviews)
# -
# send review col to list
reviews = df_game_reviews.review.tolist()
# +
# clean
# remove new line delimeter
def remove_line(reviews):
temp_lst = []
for item in reviews:
item = str(item)
item = item.replace('\n', ' ')
temp_lst.append(item)
reviews = temp_lst
temp_lst = None
df_game_reviews = None
return reviews
remove_line(reviews); # note: output hidden, because it's a nastly-long block of text.
# -
# # Exploratory Analysis
# +
# remove punctuation and make all words lowercase
def rem_punct_and_lower(df):
temp_lst = ['.','\n','!','@','#','$','%','^','&','*','(',')','<', '>', '?', ':']
for item in temp_lst:
#parse through temp_lst
df['review'] = df['review'].str.replace(item, '')
df["review"] = df["review"].str.lower() # covert words to lowercase
df['review'] = df['review'].astype(str) # convert erraythang to str
return df
rem_punct_and_lower(df_game_reviews)
def plot_50_most_common_words(count_data, count_vectorizer):
import matplotlib.pyplot as plt
words = count_vectorizer.get_feature_names()
total_counts = np.zeros(len(words))
for t in count_data:
total_counts+=t.toarray()[0]
count_dict = (zip(words, total_counts))
count_dict = sorted(count_dict, key=lambda x:x[1], reverse=True)[0:50]
words = [w[0] for w in count_dict]
counts = [w[1] for w in count_dict]
x_pos = np.arange(len(words))
plt.figure(2, figsize=(15, 15/1.6180))
plt.subplot(title='50 most common words')
sns.set_context("notebook", font_scale=1.25, rc={"lines.linewidth": 2.5})
sns.barplot(x_pos, counts, palette='husl')
plt.xticks(x_pos, words, rotation=90)
plt.xlabel('words')
plt.ylabel('counts')
plt.show()
# Initialise the count vectorizer with the English stop words
count_vectorizer = CountVectorizer(stop_words='english')
# Fit and transform the processed reviews
count_data = count_vectorizer.fit_transform(df_game_reviews['review'])
# Visualise the 50 most common words
plot_50_most_common_words(count_data, count_vectorizer)
# -
# # Tokenize
# +
def sent_to_words(sentences):
for sentence in sentences:
yield(gensim.utils.simple_preprocess(str(sentence), deacc=True)) # deacc = True removes punctuations
data_words = list(sent_to_words(reviews))
print(data_words[:1])
# -
# # Lemmatization
# +
def lemmatization(texts, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV']):
"""https://spacy.io/api/annotation"""
texts_out = []
for sent in texts:
doc = nlp(" ".join(sent))
texts_out.append(" ".join([token.lemma_ if token.lemma_ not in ['-PRON-'] else '' for token in doc if token.pos_ in allowed_postags]))
return texts_out
# Initialize spacy 'en' model, keeping only tagger component (for efficiency)
nlp = spacy.load('en', disable=['parser', 'ner'])
# Do lemmatization keeping only Noun, Adj, Verb, Adverb
data_lemmatized = lemmatization(data_words, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV'])
print(data_lemmatized[:2])
# -
# # Create Document-Word matrix
# +
vectorizer = CountVectorizer(analyzer='word',
min_df=10, # minimum reqd occurences of a word
stop_words='english', # remove stop words
lowercase=True, # convert all words to lowercase
token_pattern='[a-zA-Z0-9]{3,}', # num chars > 3
# max_features=50000, # max number of uniq words
)
data_vectorized = vectorizer.fit_transform(data_lemmatized)
# -
# # Check Sparsicity
# +
# Materialize the sparse data
data_dense = data_vectorized.todense()
# Compute Sparsicity = Percentage of Non-Zero cells
print("Sparsicity: ", ((data_dense > 0).sum()/data_dense.size)*100, "%")
# -
# # Build LDA model and Diagnose performance with perplexit and log-lielihood
# (A model with higher log-likelihood and lower perplexity (exp(-1. * log-likelihood per word)) is considered to be good. Let’s check for our model.)
# ### Find 'Best' model parameters
# (# Fit LDA with a few different values for K aka n_components. How does the value of K seem to change your results?)
# +
# define Search Param
search_params = {'n_components': [1, 2, 3, 4, 5, 6, 7, 8, 9, 10,
11,12,13,14,15,16,17,18,19,20,
21,22,23,24,25,26,27,28,29,30,
31,32,33,34,35,36,37,38,39,40,
41,42,43,44,45,46,47,48,49,50,
51,52,53,54,55,56,57,58,59,60,
61,62,63,64,65,66,67,68,69,70,
71,72,73,74,75,76,77,78,79,80,
81,82,83,84,85,86,87,88,89,90,
91,92,93,94,95,96,97,98,99,100] #number of topics
}
# init the Model
lda = LatentDirichletAllocation(doc_topic_prior=None,
evaluate_every=-1,
learning_decay=0.7,
learning_offset=10.0,
max_doc_update_iter=100,
max_iter=10,
mean_change_tol=0.001,
n_components=5,
n_jobs=-1,
perp_tol=0.1,
random_state=42,
topic_word_prior=None,
total_samples=1000000.0,
verbose=0
)
# init Grid Search Class
model = GridSearchCV(cv = None,
estimator=lda,
param_grid=search_params,
pre_dispatch='2*n_jobs',
refit = True,
scoring = None,
verbose = 0,
n_jobs = 1,
return_train_score = 'warn'
)
# do the Grid Search
model.fit(data_vectorized)
# +
# Best Model
best_lda_model = model.best_estimator_
# Model Parameters
print("Best Model's Params: ", model.best_params_)
# Log Likelihood Score
print("Best Log Likelihood Score: ", model.best_score_)
# Perplexity
print("Model Perplexity: ", best_lda_model.perplexity(data_vectorized))
#previous runs:
#Best Model's Params: {'n_components': 1}
#Best Log Likelihood Score: -849166.1577512659
#Model Perplexity: 698.6306022156406
#Best Model's Params: {'learning_decay': 0.5, 'n_components': 1}
#Best Log Likelihood Score: -849166.1577512659
#Model Perplexity: 698.6306022156406
# -
# # Fit LDA with a few different values for K. How does the value of K seem to change your results?
# (the increase of K's value, decreased the model's strength)
# +
#plot of learning optimal params
def plot_scores(model):
temp_params = []
temp_mean_test_score = []
n_topics = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10,
11,12,13,14,15,16,17,18,19,20,
21,22,23,24,25,26,27,28,29,30,
31,32,33,34,35,36,37,38,39,40,
41,42,43,44,45,46,47,48,49,50,
51,52,53,54,55,56,57,58,59,60,
61,62,63,64,65,66,67,68,69,70,
71,72,73,74,75,76,77,78,79,80,
81,82,83,84,85,86,87,88,89,90,
91,92,93,94,95,96,97,98,99,100] # K
for item in model.cv_results_['mean_test_score']:
temp_mean_test_score.append(item)
plt.figure(figsize=(12, 8))
plt.plot(n_topics, temp_mean_test_score)
plt.title("Choosing Optimal LDA Model")
plt.xlabel("Num Topics (K)")
plt.ylabel("Log Likelyhood Scores")
#plt.legend(title='Learning decay', loc='best')
plt.show()
plot_scores(model)
# -
# # Dominant Topic in each Document
# (note: because there is only 1 topic, this section becomes irrelevant. I'm leaving it in here because I might change topic count)
# +
# Create Document - Topic Matrix
lda_output = best_lda_model.transform(data_vectorized)
# column names
topicnames = ["Topic" + str(i) for i in range(best_lda_model.n_components)]
# index names
docnames = ["Doc" + str(i) for i in range(len(reviews))]
# Make the pandas dataframe
df_document_topic = pd.DataFrame(np.round(lda_output, 2), columns=topicnames, index=docnames)
# Get dominant topic for each document
dominant_topic = np.argmax(df_document_topic.values, axis=1)
df_document_topic['dominant_topic'] = dominant_topic
# Styling
def color_green(val):
color = 'green' if val > .1 else 'black'
return 'color: {col}'.format(col=color)
def make_bold(val):
weight = 700 if val > .1 else 400
return 'font-weight: {weight}'.format(weight=weight)
# Apply Style
df_document_topics = df_document_topic.head(15).style.applymap(color_green).applymap(make_bold)
df_document_topics
# -
# # Visualize LDA Model
# (this section is also not used due to topic count)
# Here's an example of the graphic: https://user-images.githubusercontent.com/20510239/28708365-9d274b3a-73ae-11e7-805f-6c65adbd4907.png
# +
#pyLDAvis.enable_notebook()
#panel = pyLDAvis.sklearn.prepare(best_lda_model, data_vectorized, vectorizer, mds='tsne')
#panel
# -
# # Using your knowledge of the corpus, choose the best value for K and justify this result substantively. Fit a topic model with this value, interpret it substantively as it relates to your research question, and write these results up:
# # Inspect Topic Keywords
# +
# Topic-Keyword Matrix
df_topic_keywords = pd.DataFrame(best_lda_model.components_)
# Assign Column and Index
df_topic_keywords.columns = vectorizer.get_feature_names()
df_topic_keywords.index = topicnames
# View
df_topic_keywords.head()
# +
#top 15 keywords
# Show top n keywords for each topic
def show_topics(vectorizer=vectorizer, lda_model=best_lda_model, n_words=20):
keywords = np.array(vectorizer.get_feature_names())
topic_keywords = []
for topic_weights in lda_model.components_:
top_keyword_locs = (-topic_weights).argsort()[:n_words]
topic_keywords.append(keywords.take(top_keyword_locs))
return topic_keywords
topic_keywords = show_topics(vectorizer=vectorizer, lda_model=best_lda_model, n_words=15)
# Topic - Keywords Dataframe
df_topic_keywords = pd.DataFrame(topic_keywords)
df_topic_keywords.columns = ['Word '+str(i) for i in range(df_topic_keywords.shape[1])]
df_topic_keywords.index = ['Topic '+str(i) for i in range(df_topic_keywords.shape[0])]
df_topic_keywords
# -
# # Cluster Documents that share similar plot.
# (with just one topic, this section is useless but I'm leaving it in, in the event that I change topic count in the future)
# +
# Construct the k-means clusters
#from sklearn.cluster import KMeans
#clusters = KMeans(n_clusters=15, random_state=100).fit_predict(lda_output)
# Build the Singular Value Decomposition(SVD) model
#svd_model = TruncatedSVD(n_components=2) # 2 components
#lda_output_svd = svd_model.fit_transform(lda_output)
# X and Y axes of the plot using SVD decomposition
#x = lda_output_svd[:, 0]
#y = lda_output_svd[:, 1]
# Weights for the 15 columns of lda_output, for each component
#print("Component's weights: \n", np.round(svd_model.components_, 2))
# Percentage of total information in 'lda_output' explained by the two components
#print("Perc of Variance Explained: \n", np.round(svd_model.explained_variance_ratio_, 2))
# -
# # Return Simiar Docs for test review
def predict_topic(text, nlp=nlp):
global sent_to_words
global lemmatization
# Step 1: Clean with simple_preprocess
mytext_2 = list(sent_to_words(text))
# Step 2: Lemmatize
mytext_3 = lemmatization(mytext_2, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV'])
# Step 3: Vectorize transform
mytext_4 = vectorizer.transform(mytext_3)
# Step 4: LDA Transform
topic_probability_scores = best_lda_model.transform(mytext_4)
topic = df_topic_keywords.iloc[np.argmax(topic_probability_scores), :].values.tolist()
return topic, topic_probability_scores
# +
nlp = spacy.load('en', disable=['parser', 'ner'])
def similar_documents(text, doc_topic_probs, documents = reviews, nlp=nlp, top_n=5, verbose=False):
topic, x = predict_topic(text)
dists = euclidean_distances(x.reshape(1, -1), doc_topic_probs)[0]
doc_ids = np.argsort(dists)[:top_n]
if verbose:
print("Topic KeyWords: ", topic)
print("Topic Prob Scores of text: ", np.round(x, 1))
print("Most Similar Doc's Probs: ", np.round(doc_topic_probs[doc_ids], 1))
return doc_ids, np.take(documents, doc_ids)
# -
# Get similar documents
mytext = ["I love this game, it's awesome."]
doc_ids, docs = similar_documents(text=mytext, doc_topic_probs=lda_output, documents = reviews, top_n=1, verbose=True)
print('\n', docs[0][:500])
# # Closing Thoughts:
# I am not happy with my topic count selection of one.
# In retrospect I wonder if I should have removed the word 'game' from the corpus. I might have observed more detail in my automated topic selection, given the overwhelming prevalence of the word in each document.
#
# Also, unrelated, at around 80 topics my graph shows some variability. I would have liked to explore that.
| notebooks/TAD_Week_6_Broker_Carl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from pandas import DataFrame
# + pycharm={"name": "#%%\n"}
df: DataFrame = pd.read_csv('../../data/interim/2_further_exploration.csv')
df[['year', 'month', 'day']] = df[['year', 'month', 'day']].astype(int)
holidays_df = pd.read_csv('../../data/external/us_holidays.csv')
holidays_df.head()
# + pycharm={"name": "#%%\n"}
is_holiday = []
def pad_number(n, l):
return str(n.astype(int)).zfill(l)
for i, row in df.iterrows():
date = f"{pad_number(row['year'], 4)}-{pad_number(row['month'], 2)}-{pad_number(row['day'],2)}"
is_holiday.append(date in holidays_df['Date'].values)
df['is_holiday'] = is_holiday
# + pycharm={"name": "#%%\n"}
df[['is_holiday']].value_counts()
# + pycharm={"name": "#%%\n"}
"2004-07-04" in holidays_df['Date'].values
# + pycharm={"name": "#%%\n"}
| notebooks/0.1_data_processing_tests/0.3-mmykhaylov-us-holidays-processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Buscas supervisionadas
# ## Imports
# +
# imports necessarios
from search import *
from notebook import psource, heatmap, gaussian_kernel, show_map, final_path_colors, display_visual, plot_NQueens
import networkx as nx
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import MultipleLocator
import time
from statistics import mean, stdev
from math import sqrt
from memory_profiler import memory_usage
# Needed to hide warnings in the matplotlib sections
import warnings
warnings.filterwarnings("ignore")
# -
# ## Criação do mapa e do grafo
# +
# make the dict where the key is associated with his neighbors
mapa = {}
for i in range(0,60):
for j in range(0,60):
mapa[(i,j)] = {(i+1,j):1, (i-1,j):1, (i,j+1):1, (i,j-1):1}
grafo = UndirectedGraph(mapa)
# -
# ## Modelagem da classe problema
class RobotProblem(Problem):
"""Problema para encontrar o goal saindo de uma posicao (x,y) com um robo."""
def __init__(self, initial, goal, mapa, graph):
Problem.__init__(self, initial, goal)
self.mapa = mapa
self.graph = graph
def actions(self, actual_pos):
"""The actions at a graph node are just its neighbors."""
neighbors = list(self.graph.get(actual_pos).keys())
valid_actions = []
for act in neighbors:
if act[0] == 0 or act[0] == 60 or act[1] == 0 or act[1] == 60:
i = 1
elif (act[0] == 20 and (0<= act[1] <= 40)):
i = 2
elif (act[0] == 40 and (20<= act[1] <= 60)):
i = 3
else:
valid_actions.append(act)
return valid_actions
def result(self, state, action):
"""The result of going to a neighbor is just that neighbor."""
return action
def path_cost(self, cost_so_far, state1, action, state2):
return cost_so_far + 1
def goal_test(self, state):
if state[0] == self.goal[0] and state[1] == self.goal[1]:
return True
else:
return False
def heuristic_1(self, node):
"""h function is straight-line distance from a node's state to goal."""
locs = getattr(self.graph, 'locations', None)
if locs:
if type(node) is str:
return int(distance(locs[node], locs[self.goal]))
return int(distance(locs[node.state], locs[self.goal]))
else:
return infinity
def heuristic_2(self,node):
""" Manhattan Heuristic Function """
x1,y1 = node.state[0], node.state[1]
x2,y2 = self.goal[0], self.goal[1]
return abs(x2 - x1) + abs(y2 - y1)
# ## Busca supervisionada A*: Heuristica 1
# ### Calculo do consumo de memoria
def calc_memory_a_h1():
init_pos = (10,10)
goal_pos = (50,50)
robot_problem = RobotProblem(init_pos, goal_pos, mapa, grafo)
node = astar_search(robot_problem, h=robot_problem.heuristic_1)
mem_usage = memory_usage(calc_memory_a_h1)
print('Memória usada (em intervalos de .1 segundos): %s' % mem_usage)
print('Maximo de memoria usada: %s' % max(mem_usage))
# ### Calculo do custo da busca e o caminho percorrido
# +
init_pos = (10,10)
goal_pos = (50,50)
robot_problem = RobotProblem(init_pos, goal_pos, mapa, grafo)
node = astar_search(robot_problem, h=robot_problem.heuristic_1)
print("Custo da busca A* com a primeira heuristica: " + str(node.path_cost))
# -
list_nodes = []
for n in node.path():
list_nodes.append(n.state)
x = []
y = []
for nod in list_nodes:
x.append(nod[0])
y.append(nod[1])
# +
fig = plt.figure()
plt.xlim(0,60)
plt.ylim(0,60)
plt.title('Caminho percorrido pelo robo na busca A* com a primeira heuristica')
plt.annotate("",
xy=(0,0), xycoords='data',
xytext=(0, 60), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(0,0), xycoords='data',
xytext=(60, 0), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(60,0), xycoords='data',
xytext=(60, 60), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(0,60), xycoords='data',
xytext=(60, 60), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(40,20), xycoords='data',
xytext=(40, 60), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(20,0), xycoords='data',
xytext=(20, 40), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.scatter(x,y)
plt.scatter(10,10,color='r')
plt.scatter(50,50,color='r')
plt.show()
# -
# ### Calculo do tempo gasto pelo A* com inicio em (10,10) e fim em (50,50) usando a heuristica 1
# +
init_pos = (10,10)
goal_pos = (50,50)
robot_problem = RobotProblem(init_pos, goal_pos, mapa, grafo)
times = []
for i in range(0,1000):
start = time.time()
node = astar_search(robot_problem, h=robot_problem.heuristic_1)
end = time.time()
times.append(end - start)
# -
media_a_1 = mean(times)
desvio_a_1 = stdev(times)
intervalo_conf = '(' + str( media_a_1 - 1.96 * (desvio_a_1 / (len(times)) ** (1/2)) ) + ',' + str( media_a_1 + 1.96 * (desvio_a_1 / (len(times)) ** (1/2)) ) + ')'
print("Media do tempo gasto para a busca A* com a primeira heuristica: " + str(media_a_1))
print("Desvio padrao do tempo gasto para a busca A* com a primeira heuristica: " + str(desvio_a_1))
print("Intervalo de confiança para a busca A* com a primeira heuristica: " + intervalo_conf)
fig = plt.figure()
plt.hist(times,bins=50)
plt.title('Histograma para o tempo de execucao do A* com a primeira heuristica')
plt.show()
# ### Projecao da relacao entre distancia em linha reta e tempo para o A* com a primeira heuristica
goal_pos = (50,50)
x = []
y = []
for i in range(5,50):
for j in range(5,50):
if i != 20 and i != 40:
init_pos = (i,i)
distancia_linha_reta = sqrt( (goal_pos[0] - init_pos[0]) ** 2 + (goal_pos[1] - init_pos[1]) ** 2)
robot_problem = RobotProblem(init_pos, goal_pos, mapa, grafo)
start = time.time()
node = astar_search(robot_problem, h=robot_problem.heuristic_1)
end = time.time()
x.append(distancia_linha_reta)
y.append(end - start)
import pandas as pd
data = {'x':[], 'y':[]}
df = pd.DataFrame(data)
df['x'] = x
df['y'] = y
df
fig = plt.figure()
plt.scatter(x,y)
plt.ylim(0.2, 1)
plt.title("Distancia em linha reta x Tempo A*-heuristica1")
plt.xlabel("Distancia em linha reta entre os pontos inicial e final")
plt.ylabel("Tempo da busca A* com a primeira heuristica")
plt.show()
# ## Busca supervisionada A*: Heuristica 2
# ### Calculo do consumo de memoria
def calc_memory_a_h2():
init_pos = (10,10)
goal_pos = (50,50)
robot_problem = RobotProblem(init_pos, goal_pos, mapa, grafo)
node = astar_search(robot_problem, h=robot_problem.heuristic_2)
mem_usage = memory_usage(calc_memory_a_h2)
print('Memória usada (em intervalos de .1 segundos): %s' % mem_usage)
print('Maximo de memoria usada: %s' % max(mem_usage))
# ### Calculo do custo da busca e o caminho percorrido
# +
init_pos = (10,10)
goal_pos = (50,50)
robot_problem = RobotProblem(init_pos, goal_pos, mapa, grafo)
node = astar_search(robot_problem, h=robot_problem.heuristic_2)
print("Custo da busca A* com a segunda heuristica: " + str(node.path_cost))
# -
list_nodes = []
for n in node.path():
list_nodes.append(n.state)
x = []
y = []
for nod in list_nodes:
x.append(nod[0])
y.append(nod[1])
# +
fig = plt.figure()
plt.xlim(0,60)
plt.ylim(0,60)
plt.title('Caminho percorrido pelo robo na busca A* com a segunda heuristica')
plt.annotate("",
xy=(0,0), xycoords='data',
xytext=(0, 60), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(0,0), xycoords='data',
xytext=(60, 0), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(60,0), xycoords='data',
xytext=(60, 60), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(0,60), xycoords='data',
xytext=(60, 60), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(40,20), xycoords='data',
xytext=(40, 60), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.annotate("",
xy=(20,0), xycoords='data',
xytext=(20, 40), textcoords='data',
arrowprops=dict(arrowstyle="-",
edgecolor = "black",
linewidth=5,
alpha=0.65,
connectionstyle="arc3,rad=0."),
)
plt.scatter(x,y)
plt.scatter(10,10,color='r')
plt.scatter(50,50,color='r')
plt.show()
# -
# ### Calculo do tempo gasto pelo A* com inicio em (10,10) e fim em (50,50) usando a heuristica 2
# +
init_pos = (10,10)
goal_pos = (50,50)
robot_problem = RobotProblem(init_pos, goal_pos, mapa, grafo)
times = []
for i in range(0,1000):
start = time.time()
node = astar_search(robot_problem, h=robot_problem.heuristic_2)
end = time.time()
times.append(end - start)
# -
media_a_2 = mean(times)
desvio_a_2 = stdev(times)
intervalo_conf = '(' + str( media_a_2 - 1.96 * (desvio_a_2 / (len(times)) ** (1/2)) ) + ',' + str( media_a_2 + 1.96 * (desvio_a_2 / (len(times)) ** (1/2)) ) + ')'
print("Media do tempo gasto para a busca A* com a segunda heuristica: " + str(media_a_2))
print("Desvio padrao do tempo gasto para a busca A* com a segunda heuristica: " + str(desvio_a_2))
print("Intervalo de confiança para a busca A* com a segunda heuristica: " + intervalo_conf)
fig = plt.figure()
plt.hist(times,bins=50)
plt.title('Histograma para o tempo de execucao do A* com a segunda heuristica')
plt.show()
# ### Projecao da relacao entre distancia em linha reta e tempo para o A* com a segunda heuristica
goal_pos = (50,50)
x = []
y = []
for i in range(5,50):
for j in range(5,50):
if i != 20 and i != 40:
init_pos = (i,i)
distancia_linha_reta = sqrt( (goal_pos[0] - init_pos[0]) ** 2 + (goal_pos[1] - init_pos[1]) ** 2)
robot_problem = RobotProblem(init_pos, goal_pos, mapa, grafo)
start = time.time()
node = astar_search(robot_problem, h=robot_problem.heuristic_2)
end = time.time()
x.append(distancia_linha_reta)
y.append(end - start)
import pandas as pd
data = {'x':[], 'y':[]}
df = pd.DataFrame(data)
df['x'] = x
df['y'] = y
df
fig = plt.figure()
plt.scatter(x,y)
plt.ylim(-0.05, 0.45)
plt.title("Distancia em linha reta x Tempo A*-heuristica2")
plt.xlabel("Distancia em linha reta entre os pontos inicial e final")
plt.ylabel("Tempo da busca A* com a segunda heuristica")
plt.show()
| projeto1/proj1-busca-supervisionada.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # External catolog queries:
#
# This notbook will demonstrate how to query catalogs using the CatalogQuery class. Each instance of this class
# has to be connected to (and search in) a specific catalog and collection. Once the instance is created it provides methods to query for:
#
# - all the sources with a certain distance.
# - closest source at a given position.
# - binary search: return yes/no if anything is around the positon.
# - user defined queries.
#
# In all cases the results of the queries will be return an astropy.table.Table object (this can be dsiabled in the last case, the user-defined query). The first item on the above list (search within a given radius) provides the basic block for the other two types of position-matching queries (the closest match and the binary search). It is implemented in three different ways:
#
# - using healpix: if the catalog sources have been assigned an healpix index, the search first returns the sources in the 9 healpix close to the target position (the central one + 8 neighbours)
# - using geoJSON/legacy coordinates: if the catalog documents have the position arranged in one of these two formats, the query is based on the `geoWithin` and `centerSphere` mongo filters.
# - raw: this method uses the `where` keyword to evaluate a javascript function on each document. This function computes the distance of the point to the target and compares it to the search radius. This method does not require any additional field to be added to the catalog.
#
# These core functions are defined in the catquery_utils module.
#
# Here below the different queries will be run using a set points uniformly distributed on a sphere. The performances of the different methods will be compared.
# +
from extcats import CatalogQuery
# initialize the CatalogQuery object pointing it to an existsing database
mqc_query = CatalogQuery.CatalogQuery(
cat_name = 'milliquas', # name of the database
coll_name = 'srcs', # name of the collection with the sources
ra_key = 'ra', dec_key = 'dec', # name of catalog fields for the coordinates
dbclient = None)
# specify target position (same format as the 'ra_key' and
# 'dec_key specified at initilization) and serach radius
target_ra, target_dec, rs = 5.458082, 16.035756, 100.
target_ra, target_dec, rs = 321.6639722, -89.48325, 100.
# -
# ## example queries: all sources within a given radius from target
#
# In the following cell we illustrate the three search methods implemented in CatalogQuery and how they
# are applied to the basic search pattern: retrieve all the sources within a given radius from the specified position.
#
# **IMPORTANT:** when using the HEALPIX method, the natural query does not return results in a circle, but rather in a square-like pattern of HEALpixels that covers the search radius. By default the matches are then skimmed for sources outside the circular search radius. Use pass circular = False to the query functions to disable this behaviour.
# +
# the 'raw' method does not require any pre-formatting of the catalog.
# It first selects points within a box of radius 'box_scale' times larger than the
# search radius using $gte and $lte operators, then uses the $where expression
# to compute the angular distance of the sources in the box from the target.
out_raw = mqc_query.findwithin(target_ra, target_dec, rs, method = 'raw', box_scale = 2.5)
if not out_raw is None:
print ("%d sources found around target position using the 'raw' method."%len(out_raw))
# the '2dsphere' method uses instead the use mongodb searches in
# spherical geometry using "$geoWithin" and "$centerSphere" operators.
# it requires the catalog documents to have been assigned a geoJSON
# or 'legacy pair' field of type 'Point' (see insert_example notebook).
out_2dsphere = mqc_query.findwithin(target_ra, target_dec, rs, method = '2dsphere')
if not out_2dsphere is None:
print ("%d sources found around target position using the '2dsphere' method."%len(out_2dsphere))
# finally, the healpix method can be used to speed up queries using a
# spatial prepartinioning of the data based on a HEALPix grid. In this
# case, the sources in the catalog should be assigned a field containing
# the ID of the healpix that contains it.
out_healpix = mqc_query.findwithin(target_ra, target_dec, rs, method = 'healpix')
if not out_healpix is None:
print ("%d sources found around target position using the 'healpix' method."%len(out_healpix))
out_healpix_square = mqc_query.findwithin(target_ra, target_dec, rs, method = 'healpix', circular = False)
if not out_healpix_square is None:
print ("%d sources found around target position using the 'healpix' (square) method."%len(out_healpix_square))
# ======================================== #
# make a plot with the query results #
# ======================================== #
# %matplotlib notebook
import matplotlib.pyplot as plt
# get a random sample from the catalog
cat_pos=[[o['ra'], o['dec']] for o in
mqc_query.src_coll.aggregate([{ '$sample': { 'size': 5000 }}])]
cat_ra, cat_dec = zip(*cat_pos)
fig=plt.figure()
ax=fig.add_subplot(111)#, projection="aitoff")
ax.scatter(cat_ra, cat_dec, label="random sample", c="k", s=50, marker="o", zorder=1)
ax.scatter(out_raw['ra'], out_raw['dec'], label="matches (RAW)", c="r", s=100, marker="+")
ax.scatter(out_2dsphere['ra'], out_2dsphere['dec'], label="matches (2D sphere)", c="b", s=100, marker="x")
ax.scatter(out_healpix['ra'], out_healpix['dec'], label="matches (HEALPix)", c="m", s=100, marker="v")
ax.scatter(
out_healpix_square['ra'], out_healpix_square['dec'], label="matches (HEALPix square)", c="g", s=50, marker="v")
ax.scatter(target_ra, target_dec, label='target', s=200, c='y', marker='*', zorder=0)
ax.set_xlim(target_ra-2, target_ra+2)
ax.set_ylim(target_dec-3, target_dec+3)
ax.legend(loc='best')
fig.show()
# -
# ## example queries: closest source to target
#
# For this use case we want to find the closest source to target position if within a given maximum distance. The method _findclosest_ use calls internally the _findwithin*_ methods show above, then use astropy's SkyCoord functionalities to compute the cource-target distance and return the closest match.
#
# **NOTE:** The method _findclosest_ returns the closest source and its distance to the target in arcseconds.
#
# +
rawcp, rawcp_dist = mqc_query.findclosest(target_ra, target_dec, rs, method = 'raw')
s2dcp, s2d_dist = mqc_query.findclosest(target_ra, target_dec, rs, method = '2dsphere')
hpcp, hpcp_dist = mqc_query.findclosest(target_ra, target_dec, rs, method = 'healpix')
# here we verify that all the counterparts are actually the same
print (' Database ID | cp-dist ["]')
print ("------------------------------------------")
print (rawcp['_id'], "|", rawcp_dist)
print (s2dcp['_id'], "|", s2d_dist)
print (hpcp['_id'], "|", hpcp_dist)
# -
# ## example queries: is there anything close to the target?
#
#
# The last functionality provided by the CatalogQuery class is that of the _binarysearch_ whose return value is a boolean: True if some counterpart is found within the search radius, False otherwise.
# +
raw_bool = mqc_query.binaryserach(target_ra, target_dec, rs, method = 'raw')
s2d_bool = mqc_query.binaryserach(target_ra, target_dec, rs, method = '2dsphere')
hp_bool = mqc_query.binaryserach(target_ra, target_dec, rs, method = 'healpix')
# here we verify that all the counterparts are actually the same
print (raw_bool, s2d_bool, hp_bool)
# -
# ## test query performances
#
#
# The CatalogQuery provides a convenience method to test queryes using a set of randomly distributed points on a sphere. This allows the user to measure the query time, and if needed, refine the indexing and querying strategy.
#
# +
# test te three main types of queries with the healpix method
mqc_query.test_queries(query_type = 'within', method = 'healpix', rs_arcsec = 3, npoints=1e4)
# here we don't seed the rng, to avoid mongo using some cached results
mqc_query.test_queries(query_type = 'within', method = 'healpix', rs_arcsec = 3, npoints=1e4, rnd_seed = None)
mqc_query.test_queries(query_type = 'closest', method = 'healpix', rs_arcsec = 3, npoints=1e4)
mqc_query.test_queries(query_type = 'binary', method = 'healpix', rs_arcsec = 3, npoints=1e4)
# and the other query methods as well (they are much slower, since there are not indexes to support them)
mqc_query.test_queries(query_type = 'closest', method ='raw', rs_arcsec = 3, npoints=10)
mqc_query.test_queries(query_type = 'closest', method ='2dsphere', rs_arcsec = 3, npoints=100)
# -
| notebooks/query_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import Imputer
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn import metrics
import sklearn.preprocessing
# +
train_file = "../data/train.csv"
test_file = "../data/test.csv"
train_data_raw = pd.read_csv(train_file)
test_data_raw = pd.read_csv(test_file)
# -
train_data = clean_func(train_data_raw)
# +
### CLEAN DATA FUNC
def clean_func(train_data):
## DO IMPUTATION
# FARE
imp_fare = Imputer(missing_values="NaN", strategy="mean")
imp_fare.fit(train_data[["Fare"]])
train_data[["Fare"]]=imp_fare.transform(train_data[["Fare"]]).ravel()
# Age
imp=Imputer(missing_values="NaN", strategy="mean")
imp.fit(train_data[["Age"]])
train_data[["Age"]]=imp.transform(train_data[["Age"]]).ravel()
# Filna
train_data["Cabin"] = train_data["Cabin"].fillna("")
# one hot encoding
sex_features = pd.get_dummies(train_data["Sex"])
embarked_features = pd.get_dummies(train_data["Embarked"])
# rename embarked features
embarked_features = embarked_features.rename(columns={'C': 'embarked_cobh'
, 'Q': 'embark_queenstown'
, 'S': 'embark_southampton'})
# Concat new features
train_data_extras = pd.concat([train_data,sex_features,embarked_features],axis=1)
# HACK - REMOVE T WHICH IS NOT IN TEST LIKELY ERRROR
cabin_letters = pd.get_dummies(train_data['Cabin'].map(lambda x: "empty" if len(x)==0 or x[0]=="T" else x[0]))
# cabin_letters = pd.get_dummies(train_data['Cabin'].map(lambda x: "empty" if len(x)==0 else x[0]))
cabin_letters.columns = ["Cabin_letter_"+i for i in cabin_letters.columns]
train_data_extras = pd.concat([train_data_extras,cabin_letters],axis=1)
train_data_extras["Cabin_number"] = train_data['Cabin'].map(lambda x: -99 if len(x)==0 else x.split(" ")[0][1:])
return train_data_extras
# -
# +
### NULL HANDLE
imp=Imputer(missing_values="NaN", strategy="mean")
imp.fit(train_data[["Age"]])
train_data[["Age"]]=imp.transform(train_data[["Age"]]).ravel() # what is ravel???
train_data["Cabin"] = train_data["Cabin"].fillna("")
# -
train_data.head(3)
# +
# LOOK AT UNIQUE VALUES
text_col = ["Sex","Ticket","Cabin","Embarked"]
for i in text_col:
print(i,train_data[i].unique()[0:20])
## Sex, Embarked can be 1 hot encoded
# -
# one hot encoding
sex_features = pd.get_dummies(train_data["Sex"])
embarked_features = pd.get_dummies(train_data["Embarked"])
train_data_extras = pd.concat([train_data,sex_features,embarked_features],axis=1)
# +
## Advanced string processing
# Name, Cabin, ticket
## Remove all digits
#tmp = train_data['Cabin'].map(lambda x: "".join([i for i in x if not i.isdigit()]))
cabin_letters = pd.get_dummies(train_data['Cabin'].map(lambda x: "empty" if len(x)==0 else x[0]))
cabin_letters.columns = ["Cabin_letter_"+i for i in cabin_letters.columns]
train_data_extras = pd.concat([train_data_extras,cabin_letters],axis=1)
train_data_extras["Cabin_number"] = train_data['Cabin'].map(lambda x: -99 if len(x)==0 else x.split(" ")[0][1:])
# train_data_extras.head()
#["Cabin"].unique()
# -
# Null handling
train_data.isnull().sum()
# +
target = "Survived"
a = train_data.dtypes
b = a[(a=="int64") | (a=="float64") | (a=="uint8")]
numerics = [i for i in b.index if i not in target]
print(numerics)
print(target)
# -
train_data[numerics].head(3)
X_train,X_test,Y_train,Y_test = train_test_split(train_data[numerics]
,train_data[target].values
,test_size=0.3
,random_state=42)
# # Models
# - logreg
# - random forest
# +
log_reg = LogisticRegression(penalty="l2", dual=False, tol=0.0001, C=1.0
, fit_intercept=True, intercept_scaling=1
, class_weight=None, random_state=None
, solver="liblinear", max_iter=100
, multi_class="ovr", verbose=0
, warm_start=False, n_jobs=1)
log_reg.fit(X_train,Y_train)
# -
Y_pred = log_reg.predict(X_test)
metrics.accuracy_score(Y_test,Y_pred)
# ### random forest naive
# +
model_rf = RandomForestClassifier(
n_estimators=100
)
model_rf.fit(train_data[numerics], train_data[target])
# +
# Cross Validation RF
scores = cross_val_score(model_rf, train_data[numerics], train_data[target], cv=10)
print(scores)
# -
pred_rf = model_rf.predict(X_test)
metrics.accuracy_score(Y_test,pred_rf)
# ### Random Forest Grid Search
# +
model_rf_gs = RandomForestClassifier()
# -
# parmeter dict
param_grid = dict(
n_estimators=np.arange(60,101,20)
, min_samples_leaf=np.arange(2,4,1)
, criterion = ["gini","entropy"]
, max_features = np.arange(0.1,0.5,0.1)
)
print(param_grid)
grid = GridSearchCV(model_rf_gs,param_grid=param_grid,scoring = "accuracy", cv = 5)
grid.fit(train_data[numerics], train_data[target])
""
# model_rf.fit(train_data[numerics], train_data[target])
# +
#print(grid)
# for i in ['params',"mean_train_score","mean_test_score"]:
# print(i)
# print(grid.cv_results_[i])
#grid.cv_results_
# -
print(grid.best_params_)
print(grid.best_score_)
# +
model_rf_gs = RandomForestClassifier(**grid.best_params_)
model_rf_gs.fit(train_data[numerics],train_data[target])
""
#print(**grid.best_params_)
# +
# get parameters
coef = list(log_reg.coef_.ravel())
intercept = log_reg.intercept_
# print them
print intercept
for id, i in enumerate(coef):
print(numerics[id],i)
# -
# # PREDICT AND STORE OUTPUT
# +
### HACK TO COMPUTE TEST RESULT
test_data = clean_func(test_data_raw)
#test_data[["Age"]]=imp.transform(test_data[["Age"]]).ravel()
# +
## DO IMPUTATION ON FARE
# imp_fare = Imputer(missing_values="NaN", strategy="mean")
# imp_fare.fit(train_data[["Fare"]])
# test_data[["Fare"]]=imp_fare.transform(test_data[["Fare"]]).ravel() # what is ravel???
# -
test_data.isnull().sum()
# +
#test_data_y = log_reg.predict(test_data)
test_data_y = log_reg.predict(test_data[numerics])
#train_data[numerics].head(3)
# -
output = pd.DataFrame(zip(list(test_data["PassengerId"]),list(test_data_y)))
output.columns = ["PassengerId","Survived"]
output.to_csv(index=False, path_or_buf= "../data/output.csv")
def output(data,file_name):
output = pd.DataFrame(zip(list(test_data["PassengerId"]),list(data)))
output.columns = ["PassengerId","Survived"]
output.to_csv(index=False, path_or_buf= "../data/{file_name}.csv".format(file_name=file_name))
# +
model_rf_data_y = model_rf.predict(test_data[numerics])
output(model_rf_data_y,"predict_rf_1")
# -
model_rf_gs_data_y =model_rf_gs.predict(test_data[numerics])
output(model_rf_gs_data_y,"predict_rf_gs_1")
| 00_kaggle_titanic/notebooks/titanic_explore1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# # Linear Mixed Effects Models
#
# With linear mixed effects models, we wish to model a linear
# relationship for data points with inputs of varying type, categorized
# into subgroups, and associated to a real-valued output.
#
# We demonstrate with an example in Edward. A webpage version is available
# [here](http://edwardlib.org/tutorials/linear-mixed-effects-models).
# + ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# %matplotlib inline
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import edward as ed
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
from edward.models import Normal
plt.style.use('ggplot')
ed.set_seed(42)
# + [markdown] ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# ## Data
#
# We use the `InstEval` data set from the popular
# [lme4 R package](http://lme4.r-forge.r-project.org) (Bates, Mächler, Bolker, & Walker, 2015).
# It is a data set of instructor evaluation ratings, where the inputs
# (covariates) include categories such as `students` and
# `departments`, and our response variable of interest is the instructor
# evaluation rating.
# + ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# s - students - 1:2972
# d - instructors - codes that need to be remapped
# dept also needs to be remapped
data = pd.read_csv('data/insteval.csv')
data['dcodes'] = data['d'].astype('category').cat.codes
data['deptcodes'] = data['dept'].astype('category').cat.codes
data['s'] = data['s'] - 1
train = data.sample(frac=0.8)
test = data.drop(train.index)
train.head()
# + [markdown] ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# In the code, we denote:
# # + `students` as `s`
# # + `instructors` as `d`
# # + `departments` as `dept`
# # + `service` as `service`
# + ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
s_train = train['s'].values.astype(int)
d_train = train['dcodes'].values.astype(int)
dept_train = train['deptcodes'].values.astype(int)
y_train = train['y'].values.astype(float)
service_train = train['service'].values.astype(int)
n_obs_train = train.shape[0]
s_test = test['s'].values.astype(int)
d_test = test['dcodes'].values.astype(int)
dept_test = test['deptcodes'].values.astype(int)
y_test = test['y'].values.astype(float)
service_test = test['service'].values.astype(int)
n_obs_test = test.shape[0]
# + ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
n_s = 2972 # number of students
n_d = 1128 # number of instructors
n_dept = 14 # number of departments
n_obs = train.shape[0] # number of observations
# + [markdown] ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# ## Model
#
# With linear regression, one makes an independence assumption where
# each data point regresses with a constant slope among
# each other. In our setting, the observations come from
# groups which may have varying slopes and intercepts. Thus we'd like to
# build a model that can capture this behavior (<NAME>, 2006).
#
# For examples of this phenomena:
# # + The observations from a single student are not independent of
# each other. Rather, some students may systematically give low (or
# high) lecture ratings.
# # + The observations from a single teacher are not independent of
# each other. We expect good teachers to get generally good ratings and
# bad teachers to get generally bad ratings.
# # + The observations from a single department are not independent of
# each other. One department may generally have dry material and thus be
# rated lower than others.
#
#
# Typical linear regression takes the form
#
# \begin{equation*}
# \mathbf{y} = \mathbf{X}\beta + \epsilon,
# \end{equation*}
#
# where $\mathbf{X}$ corresponds to fixed effects with coefficients
# $\beta$ and $\epsilon$ corresponds to random noise,
# $\epsilon\sim\mathcal{N}(\mathbf{0}, \mathbf{I})$.
#
# In a linear mixed effects model, we add an additional term
# $\mathbf{Z}\eta$, where $\mathbf{Z}$ corresponds to random effects
# with coefficients $\eta$. The model takes the form
#
# \begin{align*}
# \eta &\sim \mathcal{N}(\mathbf{0}, \sigma^2 \mathbf{I}), \\
# \mathbf{y} &= \mathbf{X}\beta + \mathbf{Z}\eta + \epsilon.
# \end{align*}
#
# Given data, the goal is to infer $\beta$, $\eta$, and $\sigma^2$,
# where $\beta$ are model parameters ("fixed effects"), $\eta$ are
# latent variables ("random effects"), and $\sigma^2$ is a variance
# component parameter.
#
# Because the random effects have mean 0, the data's mean is captured by
# $\mathbf{X}\beta$. The random effects component $\mathbf{Z}\eta$
# captures variations in the data (e.g. Instructor \#54 is rated 1.4
# points higher than the mean).
#
# A natural question is the difference between fixed and random effects.
# A fixed effect is an effect that is constant for a given population. A
# random effect is an effect that varies for a given population (i.e.,
# it may be constant within subpopulations but varies within the overall
# population). We illustrate below in our example:
#
# # + Select `service` as the fixed effect. It is a binary covariate
# corresponding to whether the lecture belongs to the lecturer's main
# department. No matter how much additional data we collect, it
# can only take on the values in $0$ and $1$.
# # + Select the categorical values of `students`, `teachers`,
# and `departments` as the random effects. Given more
# observations from the population of instructor evaluation ratings, we
# may be looking at new students, teachers, or departments.
#
# In the syntax of R's lme4 package (Bates et al., 2015), the model
# can be summarized as
#
# ```
# y ~ 1 + (1|students) + (1|instructor) + (1|dept) + service
# ```
# where `1` denotes an intercept term,`(1|x)` denotes a
# random effect for `x`, and `x` denotes a fixed effect.
# + ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# Set up placeholders for the data inputs.
s_ph = tf.placeholder(tf.int32, [None])
d_ph = tf.placeholder(tf.int32, [None])
dept_ph = tf.placeholder(tf.int32, [None])
service_ph = tf.placeholder(tf.float32, [None])
# Set up fixed effects.
mu = tf.Variable(tf.random_normal([]))
service = tf.Variable(tf.random_normal([]))
sigma_s = tf.sqrt(tf.exp(tf.Variable(tf.random_normal([]))))
sigma_d = tf.sqrt(tf.exp(tf.Variable(tf.random_normal([]))))
sigma_dept = tf.sqrt(tf.exp(tf.Variable(tf.random_normal([]))))
# Set up random effects.
eta_s = Normal(loc=tf.zeros(n_s), scale=sigma_s * tf.ones(n_s))
eta_d = Normal(loc=tf.zeros(n_d), scale=sigma_d * tf.ones(n_d))
eta_dept = Normal(loc=tf.zeros(n_dept), scale=sigma_dept * tf.ones(n_dept))
yhat = (tf.gather(eta_s, s_ph) +
tf.gather(eta_d, d_ph) +
tf.gather(eta_dept, dept_ph) +
mu + service * service_ph)
y = Normal(loc=yhat, scale=tf.ones(n_obs))
# + [markdown] ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# ## Inference
#
# Given data, we aim to infer the model's fixed and random effects.
# In this analysis, we use variational inference with the
# $\text{KL}(q\|p)$ divergence measure. We specify fully factorized
# normal approximations for the random effects and pass in all training
# data for inference. Under the algorithm, the fixed effects will be
# estimated under a variational EM scheme.
# + ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
q_eta_s = Normal(
loc=tf.Variable(tf.random_normal([n_s])),
scale=tf.nn.softplus(tf.Variable(tf.random_normal([n_s]))))
q_eta_d = Normal(
loc=tf.Variable(tf.random_normal([n_d])),
scale=tf.nn.softplus(tf.Variable(tf.random_normal([n_d]))))
q_eta_dept = Normal(
loc=tf.Variable(tf.random_normal([n_dept])),
scale=tf.nn.softplus(tf.Variable(tf.random_normal([n_dept]))))
latent_vars = {
eta_s: q_eta_s,
eta_d: q_eta_d,
eta_dept: q_eta_dept}
data = {
y: y_train,
s_ph: s_train,
d_ph: d_train,
dept_ph: dept_train,
service_ph: service_train}
inference = ed.KLqp(latent_vars, data)
# + [markdown] ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# One way to critique the fitted model is a residual plot, i.e., a
# plot of the difference between the predicted value and the observed
# value for each data point. Below we manually run inference,
# initializing the algorithm and performing individual updates within a
# loop. We form residual plots as the algorithm progresses. This helps
# us examine how the algorithm proceeds to infer the random and fixed
# effects from data.
#
# To form residuals, we first make predictions on test data. We do this
# by copying `yhat` defined in the model and replacing its
# dependence on random effects with their inferred means. During the
# algorithm, we evaluate the predictions, feeding in test inputs.
#
# We have also fit the same model (`y ~ service + (1|dept) + (1|s) + (1|d)`,
# fit on the entire `InstEval` dataset, specifically) in `lme4`. We
# have saved the random effect estimates and will compare them to our
# learned parameters.
# + ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
yhat_test = ed.copy(yhat, {
eta_s: q_eta_s.mean(),
eta_d: q_eta_d.mean(),
eta_dept: q_eta_dept.mean()})
# + ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
inference.initialize(n_print=2000, n_iter=10000)
tf.global_variables_initializer().run()
for _ in range(inference.n_iter):
# Update and print progress of algorithm.
info_dict = inference.update()
inference.print_progress(info_dict)
t = info_dict['t']
if t == 1 or t % inference.n_print == 0:
# Make predictions on test data.
yhat_vals = yhat_test.eval(feed_dict={
s_ph: s_test,
d_ph: d_test,
dept_ph: dept_test,
service_ph: service_test})
# Form residual plot.
plt.title("Residuals for Predicted Ratings on Test Set")
plt.xlim(-4, 4)
plt.ylim(0, 800)
plt.hist(yhat_vals - y_test, 75)
plt.show()
# + [markdown] ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# ## Criticism
#
# Above, we described a method for diagnosing the fit of the model via
# residual plots. See the residual plot at the end of the algorithm.
#
# The residuals appear normally distributed with mean 0. This is a good
# sanity check for the model.
#
# We can also compare our learned parameters to those estimated by R's
# `lme4`.
# -
student_effects_lme4 = pd.read_csv('data/insteval_student_ranefs_r.csv')
instructor_effects_lme4 = pd.read_csv('data/insteval_instructor_ranefs_r.csv')
dept_effects_lme4 = pd.read_csv('data/insteval_dept_ranefs_r.csv')
student_effects_edward = q_eta_s.mean().eval()
instructor_effects_edward = q_eta_d.mean().eval()
dept_effects_edward = q_eta_dept.mean().eval()
plt.title("Student Effects Comparison")
plt.xlim(-1, 1)
plt.ylim(-1, 1)
plt.xlabel("Student Effects from lme4")
plt.ylabel("Student Effects from edward")
plt.scatter(student_effects_lme4["(Intercept)"],
student_effects_edward,
alpha=0.25)
plt.show()
plt.title("Instructor Effects Comparison")
plt.xlim(-1.5, 1.5)
plt.ylim(-1.5, 1.5)
plt.xlabel("Instructor Effects from lme4")
plt.ylabel("Instructor Effects from edward")
plt.scatter(instructor_effects_lme4["(Intercept)"],
instructor_effects_edward,
alpha=0.25)
plt.show()
# Great! Our estimates for both student and instructor effects seem to
# match those from `lme4` closely. We have set up a slightly different
# model here (for example, our overall mean is regularized, as are our
# variances for student, department, and instructor effects, which is not
# true of `lme4`s model), and we have a different inference method, so we
# should not expect to find exactly the same parameters as `lme4`. But
# it is reassuring that they match up closely!
# Add in the intercept from R and edward
dept_effects_and_intercept_lme4 = 3.28259 + dept_effects_lme4["(Intercept)"]
dept_effects_and_intercept_edward = mu.eval() + dept_effects_edward
plt.title("Departmental Effects Comparison")
plt.xlim(3.0, 3.5)
plt.ylim(3.0, 3.5)
plt.xlabel("Department Effects from lme4")
plt.ylabel("Department Effects from edward")
plt.scatter(dept_effects_and_intercept_lme4,
dept_effects_and_intercept_edward,
s=0.01 * train.dept.value_counts())
plt.show()
# Our department effects do not match up nearly as well with those from `lme4`.
# There are likely several reasons for this:
# * We regularize the overal mean, while `lme4` doesn't, which causes the
# edward model to put some of the intercept into the department effects,
# which are allowed to vary more widely since we learn a variance
# * We are using 80% of the data to train the edward model, while our `lme4`
# estimate uses the whole `InstEval` data set
# * The department effects are the weakest in the model and difficult to
# estimate.
# + [markdown] ein.tags=["worksheet-0"] slideshow={"slide_type": "-"}
# ## Acknowledgments
#
# We thank <NAME> for writing the initial version of this
# tutorial.
| notebooks/linear_mixed_effects_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Session 1: Introduction to Tensorflow
# <p class='lead'>
# Creative Applications of Deep Learning with Tensorflow<br />
# <NAME><br />
# Kadenze, Inc.<br />
# </p>
#
# <a name="learning-goals"></a>
# # Learning Goals
#
# * Learn the basic idea behind machine learning: learning from data and discovering representations
# * Learn how to preprocess a dataset using its mean and standard deviation
# * Learn the basic components of a Tensorflow Graph
#
# # Table of Contents
# <!-- MarkdownTOC autolink=true autoanchor=true bracket=round -->
#
# - [Introduction](#introduction)
# - [Promo](#promo)
# - [Session Overview](#session-overview)
# - [Learning From Data](#learning-from-data)
# - [Deep Learning vs. Machine Learning](#deep-learning-vs-machine-learning)
# - [Invariances](#invariances)
# - [Scope of Learning](#scope-of-learning)
# - [Existing datasets](#existing-datasets)
# - [Preprocessing Data](#preprocessing-data)
# - [Understanding Image Shapes](#understanding-image-shapes)
# - [The Batch Dimension](#the-batch-dimension)
# - [Mean/Deviation of Images](#meandeviation-of-images)
# - [Dataset Preprocessing](#dataset-preprocessing)
# - [Histograms](#histograms)
# - [Histogram Equalization](#histogram-equalization)
# - [Tensorflow Basics](#tensorflow-basics)
# - [Variables](#variables)
# - [Tensors](#tensors)
# - [Graphs](#graphs)
# - [Operations](#operations)
# - [Tensor](#tensor)
# - [Sessions](#sessions)
# - [Tensor Shapes](#tensor-shapes)
# - [Many Operations](#many-operations)
# - [Convolution](#convolution)
# - [Creating a 2-D Gaussian Kernel](#creating-a-2-d-gaussian-kernel)
# - [Convolving an Image with a Gaussian](#convolving-an-image-with-a-gaussian)
# - [Convolve/Filter an image using a Gaussian Kernel](#convolvefilter-an-image-using-a-gaussian-kernel)
# - [Modulating the Gaussian with a Sine Wave to create Gabor Kernel](#modulating-the-gaussian-with-a-sine-wave-to-create-gabor-kernel)
# - [Manipulating an image with this Gabor](#manipulating-an-image-with-this-gabor)
# - [Homework](#homework)
# - [Next Session](#next-session)
# - [Reading Material](#reading-material)
#
# <!-- /MarkdownTOC -->
#
# <a name="introduction"></a>
# # Introduction
#
# This course introduces you to deep learning: the state-of-the-art approach to building artificial intelligence algorithms. We cover the basic components of deep learning, what it means, how it works, and develop code necessary to build various algorithms such as deep convolutional networks, variational autoencoders, generative adversarial networks, and recurrent neural networks. A major focus of this course will be to not only understand how to build the necessary components of these algorithms, but also how to apply them for exploring creative applications. We'll see how to train a computer to recognize objects in an image and use this knowledge to drive new and interesting behaviors, from understanding the similarities and differences in large datasets and using them to self-organize, to understanding how to infinitely generate entirely new content or match the aesthetics or contents of another image. Deep learning offers enormous potential for creative applications and in this course we interrogate what's possible. Through practical applications and guided homework assignments, you'll be expected to create datasets, develop and train neural networks, explore your own media collections using existing state-of-the-art deep nets, synthesize new content from generative algorithms, and understand deep learning's potential for creating entirely new aesthetics and new ways of interacting with large amounts of data.
#
# <a name="promo"></a>
# ## Promo
#
# Deep learning has emerged at the forefront of nearly every major computational breakthrough in the last 4 years. It is no wonder that it is already in many of the products we use today, from netflix or amazon's personalized recommendations; to the filters that block our spam; to ways that we interact with personal assistants like Apple's Siri or Microsoft Cortana, even to the very ways our personal health is monitored. And sure deep learning algorithms are capable of some amazing things. But it's not just science applications that are benefiting from this research.
#
# Artists too are starting to explore how Deep Learning can be used in their own practice. Photographers are starting to explore different ways of exploring visual media. Generative artists are writing algorithms to create entirely new aesthetics. Filmmakers are exploring virtual worlds ripe with potential for procedural content.
#
# In this course, we're going straight to the state of the art. And we're going to learn it all. We'll see how to make an algorithm paint an image, or hallucinate objects in a photograph. We'll see how to train a computer to recognize objects in an image and use this knowledge to drive new and interesting behaviors, from understanding the similarities and differences in large datasets to using them to self organize, to understanding how to infinitely generate entirely new content or match the aesthetics or contents of other images. We'll even see how to teach a computer to read and synthesize new phrases.
#
# But we won't just be using other peoples code to do all of this. We're going to develop everything ourselves using Tensorflow and I'm going to show you how to do it. This course isn't just for artists nor is it just for programmers. It's for people that want to learn more about how to apply deep learning with a hands on approach, straight into the python console, and learn what it all means through creative thinking and interaction.
#
# I'm <NAME>, artist, researcher and Director of Machine Intelligence at Kadenze. For the last 10 years, I've been exploring creative uses of computational models making use of machine and deep learning, film datasets, eye-tracking, EEG, and fMRI recordings exploring applications such as generative film experiences, augmented reality hallucinations, and expressive control of large audiovisual corpora.
#
# But this course isn't just about me. It's about bringing all of you together. It's about bringing together different backgrounds, different practices, and sticking all of you in the same virtual room, giving you access to state of the art methods in deep learning, some really amazing stuff, and then letting you go wild on the Kadenze platform. We've been working very hard to build a platform for learning that rivals anything else out there for learning this stuff.
#
# You'll be able to share your content, upload videos, comment and exchange code and ideas, all led by the course I've developed for us. But before we get there we're going to have to cover a lot of groundwork. The basics that we'll use to develop state of the art algorithms in deep learning. And that's really so we can better interrogate what's possible, ask the bigger questions, and be able to explore just where all this is heading in more depth. With all of that in mind, Let's get started>
#
# Join me as we learn all about Creative Applications of Deep Learning with Tensorflow.
#
# <a name="session-overview"></a>
# ## Session Overview
#
# We're first going to talk about Deep Learning, what it is, and how it relates to other branches of learning. We'll then talk about the major components of Deep Learning, the importance of datasets, and the nature of representation, which is at the heart of deep learning.
#
# If you've never used Python before, we'll be jumping straight into using libraries like numpy, matplotlib, and scipy. Before starting this session, please check the resources section for a notebook introducing some fundamentals of python programming. When you feel comfortable with loading images from a directory, resizing, cropping, how to change an image datatype from unsigned int to float32, and what the range of each data type should be, then come back here and pick up where you left off. We'll then get our hands dirty with Tensorflow, Google's library for machine intelligence. We'll learn the basic components of creating a computational graph with Tensorflow, including how to convolve an image to detect interesting features at different scales. This groundwork will finally lead us towards automatically learning our handcrafted features/algorithms.
#
# <a name="learning-from-data"></a>
# # Learning From Data
#
# <a name="deep-learning-vs-machine-learning"></a>
# ## Deep Learning vs. Machine Learning
#
# So what is this word I keep using, Deep Learning. And how is it different to Machine Learning? Well Deep Learning is a *type* of Machine Learning algorithm that uses Neural Networks to learn. The type of learning is "Deep" because it is composed of many layers of Neural Networks. In this course we're really going to focus on supervised and unsupervised Deep Learning. But there are many other incredibly valuable branches of Machine Learning such as Reinforcement Learning, Dictionary Learning, Probabilistic Graphical Models and Bayesian Methods (Bishop), or Genetic and Evolutionary Algorithms. And any of these branches could certainly even be combined with each other or with Deep Networks as well. We won't really be able to get into these other branches of learning in this course. Instead, we'll focus more on building "networks", short for neural networks, and how they can do some really amazing things. Before we can get into all that, we're going to need to understand a bit more about data and its importance in deep learning.
#
# <a name="invariances"></a>
# ## Invariances
#
# Deep Learning requires data. A lot of it. It's really one of the major reasons as to why Deep Learning has been so successful. Having many examples of the thing we are trying to learn is the first thing you'll need before even thinking about Deep Learning. Often, it is the biggest blocker to learning about something in the world. Even as a child, we need a lot of experience with something before we begin to understand it. I find I spend most of my time just finding the right data for a network to learn. Getting it from various sources, making sure it all looks right and is labeled. That is a lot of work. The rest of it is easy as we'll see by the end of this course.
#
# Let's say we would like build a network that is capable of looking at an image and saying what object is in the image. There are so many possible ways that an object could be manifested in an image. It's rare to ever see just a single object in isolation. In order to teach a computer about an object, we would have to be able to give it an image of an object in every possible way that it could exist.
#
# We generally call these ways of existing "invariances". That just means we are trying not to vary based on some factor. We are invariant to it. For instance, an object could appear to one side of an image, or another. We call that translation invariance. Or it could be from one angle or another. That's called rotation invariance. Or it could be closer to the camera, or farther. and That would be scale invariance. There are plenty of other types of invariances, such as perspective or brightness or exposure to give a few more examples for photographic images.
#
# <a name="scope-of-learning"></a>
# ## Scope of Learning
#
# With Deep Learning, you will always need a dataset that will teach the algorithm about the world. But you aren't really teaching it everything. You are only teaching it what is in your dataset! That is a very important distinction. If I show my algorithm only faces of people which are always placed in the center of an image, it will not be able to understand anything about faces that are not in the center of the image! Well at least that's mostly true.
#
# That's not to say that a network is incapable of transfering what it has learned to learn new concepts more easily. Or to learn things that might be necessary for it to learn other representations. For instance, a network that has been trained to learn about birds, probably knows a good bit about trees, branches, and other bird-like hangouts, depending on the dataset. But, in general, we are limited to learning what our dataset has access to.
#
# So if you're thinking about creating a dataset, you're going to have to think about what it is that you want to teach your network. What sort of images will it see? What representations do you think your network could learn given the data you've shown it?
#
# One of the major contributions to the success of Deep Learning algorithms is the amount of data out there. Datasets have grown from orders of hundreds to thousands to many millions. The more data you have, the more capable your network will be at determining whatever its objective is.
#
# <a name="existing-datasets"></a>
# ## Existing datasets
#
# With that in mind, let's try to find a dataset that we can work with. There are a ton of datasets out there that current machine learning researchers use. For instance if I do a quick Google search for Deep Learning Datasets, i can see for instance a link on deeplearning.net, listing a few interesting ones e.g. http://deeplearning.net/datasets/, including MNIST, CalTech, CelebNet, LFW, CIFAR, MS Coco, Illustration2Vec, and there are ton more. And these are primarily image based. But if you are interested in finding more, just do a quick search or drop a quick message on the forums if you're looking for something in particular.
#
# * MNIST
# * CalTech
# * CelebNet
# * ImageNet: http://www.image-net.org/
# * LFW
# * CIFAR10
# * CIFAR100
# * MS Coco: http://mscoco.org/home/
# * WLFDB: http://wlfdb.stevenhoi.com/
# * Flickr 8k: http://nlp.cs.illinois.edu/HockenmaierGroup/Framing_Image_Description/KCCA.html
# * Flickr 30k
#
# <a name="preprocessing-data"></a>
# # Preprocessing Data
#
# In this section, we're going to learn a bit about working with an image based dataset. We'll see how image dimensions are formatted as a single image and how they're represented as a collection using a 4-d array. We'll then look at how we can perform dataset normalization. If you're comfortable with all of this, please feel free to skip to the next video.
#
# We're first going to load some libraries that we'll be making use of.
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# I'll be using a popular image dataset for faces called the CelebFaces dataset. I've provided some helper functions which you can find on the resources page, which will just help us with manipulating images and loading this dataset.
from libs import utils
# utils.<tab>
files = utils.get_celeb_files()
# Let's get the 50th image in this list of files, and then read the file at that location as an image, setting the result to a variable, `img`, and inspect a bit further what's going on:
img = plt.imread(files[50])
# img.<tab>
print(img)
# When I print out this image, I can see all the numbers that represent this image. We can use the function `imshow` to see this:
# If nothing is drawn and you are using notebook, try uncommenting the next line:
# #%matplotlib inline
plt.imshow(img)
# <a name="understanding-image-shapes"></a>
# ## Understanding Image Shapes
#
# Let's break this data down a bit more. We can see the dimensions of the data using the `shape` accessor:
img.shape
# (218, 178, 3)
# This means that the image has 218 rows, 178 columns, and 3 color channels corresponding to the Red, Green, and Blue channels of the image, or RGB. Let's try looking at just one of the color channels.
plt.imshow(img[:, :, 0], cmap='gray')
#plt.imshow(img[:, :, 1], cmap='gray')
#plt.imshow(img[:, :, 2], cmap='gray')
# We use the special colon operator to say take every value in this dimension. This is saying, give me every row, every column, and the 0th dimension of the color channels. What we're seeing is the amount of Red, Green, or Blue contributing to the overall color image.
#
# Let's use another helper function which will load every image file in the celeb dataset rather than just give us the filenames like before. By default, this will just return the first 100 images because loading the entire dataset is a bit cumbersome. In one of the later sessions, I'll show you how tensorflow can handle loading images using a pipeline so we can load this same dataset. For now, let's stick with this:
imgs = utils.get_celeb_imgs()
# We now have a list containing our images. Each index of the `imgs` list is another image which we can access using the square brackets:
plt.imshow(imgs[0])
# <a name="the-batch-dimension"></a>
# ## The Batch Dimension
#
# Remember that an image has a shape describing the height, width, channels:
imgs[0].shape
# It turns out we'll often use another convention for storing many images in an array using a new dimension called the batch dimension. The resulting image shape will be exactly the same, except we'll stick on a new dimension on the beginning... giving us number of images x the height x the width x the number of color channels.
#
# N x H x W x C
#
# A Color image should have 3 color channels, RGB.
#
# We can combine all of our images to have these 4 dimensions by telling numpy to give us an array of all the images.
data = np.array(imgs)
data.shape
# This will only work if every image in our list is exactly the same size. So if you have a wide image, short image, long image, forget about it. You'll need them all to be the same size. If you are unsure of how to get all of your images into the same size, then please please refer to the online resources for the notebook I've provided which shows you exactly how to take a bunch of images of different sizes, and crop and resize them the best we can to make them all the same size.
#
# <a name="meandeviation-of-images"></a>
# ## Mean/Deviation of Images
#
# Now that we have our data in a single numpy variable, we can do alot of cool stuff. Let's look at the mean of the batch channel:
mean_img = np.mean(data, axis=0)
plt.imshow(mean_img.astype(np.uint8))
data.shape
# This is the first step towards building our robot overlords. We've reduced down our entire dataset to a single representation which describes what most of our dataset looks like. There is one other very useful statistic which we can look at very easily:
std_img = np.std(data, axis=0)
plt.imshow(std_img.astype(np.uint8))
std_img[0,0]
std_img.shape
#
# So this is incredibly cool. We've just shown where changes are likely to be in our dataset of images. Or put another way, we're showing where and how much variance there is in our previous mean image representation.
#
# We're looking at this per color channel. So we'll see variance for each color channel represented separately, and then combined as a color image. We can try to look at the average variance over all color channels by taking their mean:
plt.imshow(np.mean(std_img, axis=2).astype(np.uint8))
# This is showing us on average, how every color channel will vary as a heatmap. The more red, the more likely that our mean image is not the best representation. The more blue, the less likely that our mean image is far off from any other possible image.
#
# <a name="dataset-preprocessing"></a>
# ## Dataset Preprocessing
#
# Think back to when I described what we're trying to accomplish when we build a model for machine learning? We're trying to build a model that understands invariances. We need our model to be able to express *all* of the things that can possibly change in our data. Well, this is the first step in understanding what can change. If we are looking to use deep learning to learn something complex about our data, it will often start by modeling both the mean and standard deviation of our dataset. We can help speed things up by "preprocessing" our dataset by removing the mean and standard deviation. What does this mean? Subtracting the mean, and dividing by the standard deviation. Another word for that is "normalization".
#
# <a name="histograms"></a>
# ## Histograms
#
# Let's have a look at our dataset another way to see why this might be a useful thing to do. We're first going to convert our `batch` x `height` x `width` x `channels` array into a 1 dimensional array. Instead of having 4 dimensions, we'll now just have 1 dimension of every pixel value stretched out in a long vector, or 1 dimensional array.
flattened = data.ravel()
print(data[:1])
print(flattened[:10])
# We first convert our N x H x W x C dimensional array into a 1 dimensional array. The values of this array will be based on the last dimensions order. So we'll have: [<font color='red'>251</font>, <font color='green'>238</font>, <font color='blue'>205</font>, <font color='red'>251</font>, <font color='green'>238</font>, <font color='blue'>206</font>, <font color='red'>253</font>, <font color='green'>240</font>, <font color='blue'>207</font>, ...]
#
# We can visualize what the "distribution", or range and frequency of possible values are. This is a very useful thing to know. It tells us whether our data is predictable or not.
plt.hist(flattened.ravel(), 255)
# The last line is saying give me a histogram of every value in the vector, and use 255 bins. Each bin is grouping a range of values. The bars of each bin describe the frequency, or how many times anything within that range of values appears.In other words, it is telling us if there is something that seems to happen more than anything else. If there is, it is likely that a neural network will take advantage of that.
#
#
# <a name="histogram-equalization"></a>
# ## Histogram Equalization
#
# The mean of our dataset looks like this:
plt.hist(mean_img.ravel(), 255)
# When we subtract an image by our mean image, we remove all of this information from it. And that means that the rest of the information is really what is important for describing what is unique about it.
#
# Let's try and compare the histogram before and after "normalizing our data":
bins = 20
fig, axs = plt.subplots(1, 3, figsize=(12, 6), sharey=True, sharex=True)
axs[0].hist((data[0]).ravel(), bins)
axs[0].set_title('img distribution')
axs[1].hist((mean_img).ravel(), bins)
axs[1].set_title('mean distribution')
axs[2].hist((data[0] - mean_img).ravel(), bins)
axs[2].set_title('(img - mean) distribution')
# What we can see from the histograms is the original image's distribution of values from 0 - 255. The mean image's data distribution is mostly centered around the value 100. When we look at the difference of the original image and the mean image as a histogram, we can see that the distribution is now centered around 0. What we are seeing is the distribution of values that were above the mean image's intensity, and which were below it. Let's take it one step further and complete the normalization by dividing by the standard deviation of our dataset:
fig, axs = plt.subplots(1, 3, figsize=(12, 6), sharey=True, sharex=True)
axs[0].hist((data[0] - mean_img).ravel(), bins)
axs[0].set_title('(img - mean) distribution')
axs[1].hist((std_img).ravel(), bins)
axs[1].set_title('std deviation distribution')
axs[2].hist(((data[0] - mean_img) / std_img).ravel(), bins)
axs[2].set_title('((img - mean) / std_dev) distribution')
# Now our data has been squished into a peak! We'll have to look at it on a different scale to see what's going on:
axs[2].set_xlim([-150, 150])
axs[2].set_xlim([-100, 100])
axs[2].set_xlim([-50, 50])
axs[2].set_xlim([-10, 10])
axs[2].set_xlim([-5, 5])
# What we can see is that the data is in the range of -3 to 3, with the bulk of the data centered around -1 to 1. This is the effect of normalizing our data: most of the data will be around 0, where some deviations of it will follow between -3 to 3.
#
# If our data does not end up looking like this, then we should either (1): get much more data to calculate our mean/std deviation, or (2): either try another method of normalization, such as scaling the values between 0 to 1, or -1 to 1, or possibly not bother with normalization at all. There are other options that one could explore, including different types of normalization such as local contrast normalization for images or PCA based normalization but we won't have time to get into those in this course.
#
# <a name="tensorflow-basics"></a>
# # Tensorflow Basics
#
# Let's now switch gears and start working with Google's Library for Numerical Computation, TensorFlow. This library can do most of the things we've done so far. However, it has a very different approach for doing so. And it can do a whole lot more cool stuff which we'll eventually get into. The major difference to take away from the remainder of this session is that instead of computing things immediately, we first define things that we want to compute later using what's called a `Graph`. Everything in Tensorflow takes place in a computational graph and running and evaluating anything in the graph requires a `Session`. Let's take a look at how these both work and then we'll get into the benefits of why this is useful:
#
# <a name="variables"></a>
# ## Variables
#
# We're first going to import the tensorflow library:
import tensorflow as tf
# Let's take a look at how we might create a range of numbers. Using numpy, we could for instance use the linear space function:
# +
x = np.linspace(-3.0, 3.0, 100)
# Immediately, the result is given to us. An array of 100 numbers equally spaced from -3.0 to 3.0.
print(x)
# We know from numpy arrays that they have a `shape`, in this case a 1-dimensional array of 100 values
print(x.shape)
# and a `dtype`, in this case float64, or 64 bit floating point values.
print(x.dtype)
# -
# <a name="tensors"></a>
# ## Tensors
#
# In tensorflow, we could try to do the same thing using their linear space function:
x = tf.linspace(-3.0, 3.0, 100)
print(x)
# Instead of a `numpy.array`, we are returned a `tf.Tensor`. The name of it is "LinSpace:0". Wherever we see this colon 0, that just means the output of. So the name of this Tensor is saying, the output of LinSpace.
#
# Think of `tf.Tensor`s the same way as you would the `numpy.array`. It is described by its `shape`, in this case, only 1 dimension of 100 values. And it has a `dtype`, in this case, `float32`. But *unlike* the `numpy.array`, there are no values printed here! That's because it actually hasn't computed its values yet. Instead, it just refers to the output of a `tf.Operation` which has been already been added to Tensorflow's default computational graph. The result of that operation is the tensor that we are returned.
#
# <a name="graphs"></a>
# ## Graphs
#
# Let's try and inspect the underlying graph. We can request the "default" graph where all of our operations have been added:
g = tf.get_default_graph()
# <a name="operations"></a>
# ## Operations
#
# And from this graph, we can get a list of all the operations that have been added, and print out their names:
[op.name for op in g.get_operations()]
# So Tensorflow has named each of our operations to generally reflect what they are doing. There are a few parameters that are all prefixed by LinSpace, and then the last one which is the operation which takes all of the parameters and creates an output for the linspace.
#
# <a name="tensor"></a>
# ## Tensor
#
# We can request the output of any operation, which is a tensor, by asking the graph for the tensor's name:
g.get_tensor_by_name('LinSpace' + ':0')
# What I've done is asked for the `tf.Tensor` that comes from the operation "LinSpace". So remember, the result of a `tf.Operation` is a `tf.Tensor`. Remember that was the same name as the tensor `x` we created before.
#
# <a name="sessions"></a>
# ## Sessions
#
# In order to actually compute anything in tensorflow, we need to create a `tf.Session`. The session is responsible for evaluating the `tf.Graph`. Let's see how this works:
# +
# We're first going to create a session:
sess = tf.Session()
# Now we tell our session to compute anything we've created in the tensorflow graph.
computed_x = sess.run(x)
print(computed_x)
# Alternatively, we could tell the previous Tensor to evaluate itself using this session:
computed_x = x.eval(session=sess)
print(computed_x)
# We can close the session after we're done like so:
sess.close()
# -
# We could also explicitly tell the session which graph we want to manage:
sess = tf.Session(graph=g)
sess.close()
# By default, it grabs the default graph. But we could have created a new graph like so:
g2 = tf.Graph()
# And then used this graph only in our session.
#
# To simplify things, since we'll be working in iPython's interactive console, we can create an `tf.InteractiveSession`:
sess = tf.InteractiveSession()
x.eval()
# Now we didn't have to explicitly tell the `eval` function about our session. We'll leave this session open for the rest of the lecture.
#
# <a name="tensor-shapes"></a>
# ## Tensor Shapes
# +
# We can find out the shape of a tensor like so:
print(x.get_shape())
# %% Or in a more friendly format
print(x.get_shape().as_list())
# -
# <a name="many-operations"></a>
# ## Many Operations
#
# Lets try a set of operations now. We'll try to create a Gaussian curve. This should resemble a normalized histogram where most of the data is centered around the mean of 0. It's also sometimes refered to by the bell curve or normal curve.
# +
# The 1 dimensional gaussian takes two parameters, the mean value, and the standard deviation, which is commonly denoted by the name sigma.
mean = 0.0
sigma = 1.0
# Don't worry about trying to learn or remember this formula. I always have to refer to textbooks or check online for the exact formula.
z = (tf.exp(tf.negative(tf.pow(x - mean, 2.0) /
(2.0 * tf.pow(sigma, 2.0)))) *
(1.0 / (sigma * tf.sqrt(2.0 * 3.1415))))
# -
# Just like before, amazingly, we haven't actually computed anything. We *have just added a bunch of operations to Tensorflow's graph. Whenever we want the value or output of this operation, we'll have to explicitly ask for the part of the graph we're interested in before we can see its result. Since we've created an interactive session, we should just be able to say the name of the Tensor that we're interested in, and call the `eval` function:
res = z.eval()
plt.plot(res)
# if nothing is drawn, and you are using ipython notebook, uncomment the next two lines:
# #%matplotlib inline
#plt.plot(res)
# <a name="convolution"></a>
# # Convolution
#
# <a name="creating-a-2-d-gaussian-kernel"></a>
# ## Creating a 2-D Gaussian Kernel
#
# Let's try creating a 2-dimensional Gaussian. This can be done by multiplying a vector by its transpose. If you aren't familiar with matrix math, I'll review a few important concepts. This is about 98% of what neural networks do so if you're unfamiliar with this, then please stick with me through this and it'll be smooth sailing. First, to multiply two matrices, their inner dimensions must agree, and the resulting matrix will have the shape of the outer dimensions.
#
# So let's say we have two matrices, X and Y. In order for us to multiply them, X's columns must match Y's rows. I try to remember it like so:
# <pre>
# (X_rows, X_cols) x (Y_rows, Y_cols)
# | | | |
# | |___________| |
# | ^ |
# | inner dimensions |
# | must match |
# | |
# |__________________________|
# ^
# resulting dimensions
# of matrix multiplication
# </pre>
# But our matrix is actually a vector, or a 1 dimensional matrix. That means its dimensions are N x 1. So to multiply them, we'd have:
# <pre>
# (N, 1) x (1, N)
# | | | |
# | |___________| |
# | ^ |
# | inner dimensions |
# | must match |
# | |
# |__________________________|
# ^
# resulting dimensions
# of matrix multiplication
# </pre>
# +
# Let's store the number of values in our Gaussian curve.
ksize = z.get_shape().as_list()[0]
# Let's multiply the two to get a 2d gaussian
z_2d = tf.matmul(tf.reshape(z, [ksize, 1]), tf.reshape(z, [1, ksize]))
# Execute the graph
plt.imshow(z_2d.eval())
# -
ksize
# <a name="convolving-an-image-with-a-gaussian"></a>
# ## Convolving an Image with a Gaussian
#
# A very common operation that we'll come across with Deep Learning is convolution. We're going to explore what this means using our new gaussian kernel that we've just created. For now, just think of it as a way of filtering information. We're going to effectively filter our image using this Gaussian function, as if the gaussian function is the lens through which we'll see our image data. What it will do is at every location we tell it to filter, it will average the image values around it based on what the kernel's values are. The Gaussian's kernel is basically saying, take a lot the center, a then decesasingly less as you go farther away from the center. The effect of convolving the image with this type of kernel is that the entire image will be blurred. If you would like an interactive exploratin of convolution, this website is great:
#
# http://setosa.io/ev/image-kernels/
# Let's first load an image. We're going to need a grayscale image to begin with. skimage has some images we can play with. If you do not have the skimage module, you can load your own image, or get skimage by pip installing "scikit-image".
from skimage import data
img = data.camera().astype(np.float32)
plt.imshow(img, cmap='gray')
print(img.shape)
# Notice our img shape is 2-dimensional. For image convolution in Tensorflow, we need our images to be 4 dimensional. Remember that when we load many iamges and combine them in a single numpy array, the resulting shape has the number of images first.
#
# N x H x W x C
#
# In order to perform 2d convolution with tensorflow, we'll need the same dimensions for our image. With just 1 grayscale image, this means the shape will be:
#
# 1 x H x W x 1
# +
# We could use the numpy reshape function to reshape our numpy array
img_4d = img.reshape([1, img.shape[0], img.shape[1], 1])
print(img_4d.shape)
# but since we'll be using tensorflow, we can use the tensorflow reshape function:
img_4d = tf.reshape(img, [1, img.shape[0], img.shape[1], 1])
print(img_4d)
# -
# Instead of getting a numpy array back, we get a tensorflow tensor. This means we can't access the `shape` parameter like we did with the numpy array. But instead, we can use `get_shape()`, and `get_shape().as_list()`:
print(img_4d.get_shape())
print(img_4d.get_shape().as_list())
# The H x W image is now part of a 4 dimensional array, where the other dimensions of N and C are 1. So there is only 1 image and only 1 channel.
#
# We'll also have to reshape our Gaussian Kernel to be 4-dimensional as well. The dimensions for kernels are slightly different! Remember that the image is:
#
# Number of Images x Image Height x Image Width x Number of Channels
#
# we have:
#
# Kernel Height x Kernel Width x Number of Input Channels x Number of Output Channels
#
# Our Kernel already has a height and width of `ksize` so we'll stick with that for now. The number of input channels should match the number of channels on the image we want to convolve. And for now, we just keep the same number of output channels as the input channels, but we'll later see how this comes into play.
# Reshape the 2d kernel to tensorflow's required 4d format: H x W x I x O
z_4d = tf.reshape(z_2d, [ksize, ksize, 1, 1])
print(z_4d.get_shape().as_list())
# <a name="convolvefilter-an-image-using-a-gaussian-kernel"></a>
# ## Convolve/Filter an image using a Gaussian Kernel
#
# We can now use our previous Gaussian Kernel to convolve our image:
convolved = tf.nn.conv2d(img_4d, z_4d, strides=[1, 1, 1, 1], padding='SAME')
res = convolved.eval()
print(res.shape)
# There are two new parameters here: `strides`, and `padding`. Strides says how to move our kernel across the image. Basically, we'll only ever use it for one of two sets of parameters:
#
# [1, 1, 1, 1], which means, we are going to convolve every single image, every pixel, and every color channel by whatever the kernel is.
#
# and the second option:
#
# [1, 2, 2, 1], which means, we are going to convolve every single image, but every other pixel, in every single color channel.
#
# Padding says what to do at the borders. If we say "SAME", that means we want the same dimensions going in as we do going out. In order to do this, zeros must be padded around the image. If we say "VALID", that means no padding is used, and the image dimensions will actually change.
# +
# Matplotlib cannot handle plotting 4D images! We'll have to convert this back to the original shape. There are a few ways we could do this. We could plot by "squeezing" the singleton dimensions.
plt.imshow(np.squeeze(res), cmap='gray')
# Or we could specify the exact dimensions we want to visualize:
plt.imshow(res[0, :, :, 0], cmap='gray')
# -
# <a name="modulating-the-gaussian-with-a-sine-wave-to-create-gabor-kernel"></a>
# ## Modulating the Gaussian with a Sine Wave to create Gabor Kernel
#
# We've now seen how to use tensorflow to create a set of operations which create a 2-dimensional Gaussian kernel, and how to use that kernel to filter or convolve another image. Let's create another interesting convolution kernel called a Gabor. This is a lot like the Gaussian kernel, except we use a sine wave to modulate that.
#
# <graphic: draw 1d gaussian wave, 1d sine, show modulation as multiplication and resulting gabor.>
#
# We first use linspace to get a set of values the same range as our gaussian, which should be from -3 standard deviations to +3 standard deviations.
xs = tf.linspace(-3.0, 3.0, ksize)
# We then calculate the sine of these values, which should give us a nice wave
ys = tf.sin(xs)
plt.figure()
plt.plot(ys.eval())
# And for multiplication, we'll need to convert this 1-dimensional vector to a matrix: N x 1
ys = tf.reshape(ys, [ksize, 1])
# We then repeat this wave across the matrix by using a multiplication of ones:
ones = tf.ones((1, ksize))
wave = tf.matmul(ys, ones)
plt.imshow(wave.eval(), cmap='gray')
# We can directly multiply our old Gaussian kernel by this wave and get a gabor kernel:
gabor = tf.multiply(wave, z_2d)
plt.imshow(gabor.eval(), cmap='gray')
# <a name="manipulating-an-image-with-this-gabor"></a>
# ## Manipulating an image with this Gabor
#
# We've already gone through the work of convolving an image. The only thing that has changed is the kernel that we want to convolve with. We could have made life easier by specifying in our graph which elements we wanted to be specified later. Tensorflow calls these "placeholders", meaning, we're not sure what these are yet, but we know they'll fit in the graph like so, generally the input and output of the network.
#
# Let's rewrite our convolution operation using a placeholder for the image and the kernel and then see how the same operation could have been done. We're going to set the image dimensions to `None` x `None`. This is something special for placeholders which tells tensorflow "let this dimension be any possible value". 1, 5, 100, 1000, it doesn't matter.
# +
# This is a placeholder which will become part of the tensorflow graph, but
# which we have to later explicitly define whenever we run/evaluate the graph.
# Pretty much everything you do in tensorflow can have a name. If we don't
# specify the name, tensorflow will give a default one, like "Placeholder_0".
# Let's use a more useful name to help us understand what's happening.
img = tf.placeholder(tf.float32, shape=[None, None], name='img')
# We'll reshape the 2d image to a 3-d tensor just like before:
# Except now we'll make use of another tensorflow function, expand dims, which adds a singleton dimension at the axis we specify.
# We use it to reshape our H x W image to include a channel dimension of 1
# our new dimensions will end up being: H x W x 1
img_3d = tf.expand_dims(img, 2)
dims = img_3d.get_shape()
print(dims)
# And again to get: 1 x H x W x 1
img_4d = tf.expand_dims(img_3d, 0)
print(img_4d.get_shape().as_list())
# Let's create another set of placeholders for our Gabor's parameters:
mean = tf.placeholder(tf.float32, name='mean')
sigma = tf.placeholder(tf.float32, name='sigma')
ksize = tf.placeholder(tf.int32, name='ksize')
# Then finally redo the entire set of operations we've done to convolve our
# image, except with our placeholders
x = tf.linspace(-3.0, 3.0, ksize)
z = (tf.exp(tf.negative(tf.pow(x - mean, 2.0) /
(2.0 * tf.pow(sigma, 2.0)))) *
(1.0 / (sigma * tf.sqrt(2.0 * 3.1415))))
z_2d = tf.matmul(
tf.reshape(z, tf.stack([ksize, 1])),
tf.reshape(z, tf.stack([1, ksize])))
ys = tf.sin(x)
ys = tf.reshape(ys, tf.stack([ksize, 1]))
ones = tf.ones(tf.stack([1, ksize]))
wave = tf.matmul(ys, ones)
gabor = tf.multiply(wave, z_2d)
gabor_4d = tf.reshape(gabor, tf.stack([ksize, ksize, 1, 1]))
# And finally, convolve the two:
convolved = tf.nn.conv2d(img_4d, gabor_4d, strides=[1, 1, 1, 1], padding='SAME', name='convolved')
convolved_img = convolved[0, :, :, 0]
# -
# What we've done is create an entire graph from our placeholders which is capable of convolving an image with a gabor kernel. In order to compute it, we have to specify all of the placeholders required for its computation.
#
# If we try to evaluate it without specifying placeholders beforehand, we will get an error `InvalidArgumentError: You must feed a value for placeholder tensor 'img' with dtype float and shape [512,512]`:
convolved_img.eval()
# It's saying that we didn't specify our placeholder for `img`. In order to "feed a value", we use the `feed_dict` parameter like so:
convolved_img.eval(feed_dict={img: data.camera()})
# But that's not the only placeholder in our graph! We also have placeholders for `mean`, `sigma`, and `ksize`. Once we specify all of them, we'll have our result:
res = convolved_img.eval(feed_dict={
img: data.camera(), mean:0.0, sigma:1.0, ksize:100})
plt.imshow(res, cmap='gray')
# Now, instead of having to rewrite the entire graph, we can just specify the different placeholders.
res = convolved_img.eval(feed_dict={
img: data.camera(),
mean: 0.0,
sigma: 0.5,
ksize: 32
})
plt.imshow(res, cmap='gray')
# <a name="homework"></a>
# # Homework
#
# For your first assignment, we'll work on creating our own dataset. You'll need to find at least 100 images and work through the [notebook](session-1.ipynb).
# <a name="next-session"></a>
# # Next Session
#
# In the next session, we'll create our first Neural Network and see how it can be used to paint an image.
#
# <a name="reading-material"></a>
# # Reading Material
#
# <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., … <NAME>. (2015). TensorFlow : Large-Scale Machine Learning on Heterogeneous Distributed Systems.
# https://arxiv.org/abs/1603.04467
#
# <NAME>, <NAME>, <NAME>. Representation Learning: A Review and New Perspectives. 24 Jun 2012.
# https://arxiv.org/abs/1206.5538
#
# <NAME>. Deep Learning in Neural Networks: An Overview. Neural Networks, 61, p 85-117, 2015.
# https://arxiv.org/abs/1404.7828
#
# LeCun, Yann, <NAME>, and <NAME>. “Deep learning.” Nature 521, no. 7553 (2015): 436-444.
#
# <NAME>ellow <NAME> and <NAME>. Deep Learning. 2016.
# http://www.deeplearningbook.org/
| session-1/lecture-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#from customplot import *
#import sqlite3
# %autosave 0
import itertools
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import reciprocal, uniform
from sklearn.cluster import KMeans
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import VotingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import SGDClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import log_loss
from sklearn.metrics import precision_score
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import recall_score
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_val_predict
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import RandomizedSearchCV
from sklearn.neighbors import KNeighborsClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import PolynomialFeatures
from sklearn.svm import SVC, LinearSVC
from sklearn.tree import DecisionTreeClassifier
# -
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
#print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
#Setando configurações de visualização
pd.options.display.max_rows=350
pd.options.display.max_columns=60
df=pd.read_csv('baseProjeto_entradaModelo.csv', index_col=0)
df
df.columns
X=df[['ATRIB_MED1', 'ATRIB_MAX1',
'ATRIB_DIST1', 'DIFP', 'MGP1', 'MGP2', 'MGP3', 'MGP4', 'MGP5', 'MGP6',
'MGP7', 'MGP8', 'MGP9', 'MGP10', 'MGP11', 'MGP12', 'MGP13', 'MGP14']]
X
X.info()
'''
#cat=['MGP1_sim', 'MGP2_sim', 'MGP3_sim', 'MGP4_sim',
'MGP5_sim', 'MGP6_sim', 'MGP7_sim', 'MGP8_sim', 'MGP9_sim', 'MGP10_sim',
'MGP11_sim', 'MGP12_sim', 'MGP13_sim', 'MGP14_sim',]
#X[cat] = X[cat].astype('category')
'''
X.info()
y = df['Perda30']
#X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.20, random_state=21)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.20, random_state=42, stratify=y)
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.fit_transform(X_test)
# # Comparing the Classifier
random_forest_clf = RandomForestClassifier(bootstrap=False, class_weight=None,
criterion='entropy', max_depth=7, max_features='auto',
max_leaf_nodes=None, min_impurity_decrease=0.0,
min_impurity_split=None, min_samples_leaf=1,
min_samples_split=2, min_weight_fraction_leaf=0.0,
n_estimators=525, n_jobs=-1, oob_score=False, random_state=42,
verbose=0, warm_start=True)
svc_clf = SVC(C=3.0, cache_size=200, class_weight=None, coef0=0.0,
decision_function_shape='ovr', degree=6, gamma=0.1,
kernel='poly', max_iter=-1, probability=True, random_state=None,
shrinking=True, tol=0.001, verbose=False)
mlp_clf = MLPClassifier(activation='relu', alpha=1.8, batch_size='auto',
beta_1=0.9, beta_2=0.999, early_stopping=False,
epsilon=1e-08, hidden_layer_sizes=(50, 100),
learning_rate='constant', learning_rate_init=0.001,
max_iter=1000, momentum=0.9, nesterovs_momentum=True,
power_t=0.5, random_state=42, shuffle=True, solver='lbfgs',
tol=0.0001, validation_fraction=0.1, verbose=False,
warm_start=False)
estimators = [random_forest_clf, svc_clf, mlp_clf]
for estimator in estimators:
print("Training the", estimator)
estimator.fit(X_train_scaled, y_train)
[estimator.score(X_test_scaled, y_test) for estimator in estimators]
# # Voting Classifier
named_estimators = [
("random_forest_clf", random_forest_clf),
("svc_clf", svc_clf),
("mlp_clf", mlp_clf),
]
voting_clf = VotingClassifier(named_estimators, n_jobs=-1)
print(voting_clf.voting)
voting_clf.fit(X_train_scaled, y_train)
voting_clf.score(X_test_scaled, y_test)
[estimator.score(X_test_scaled, y_test) for estimator in voting_clf.estimators_]
voting_clf.set_params(random_forest_clf=None)
voting_clf.estimators
# +
#del voting_clf.estimators_[0]
# -
voting_clf.score(X_test_scaled, y_test)
[estimator.score(X_test_scaled, y_test) for estimator in voting_clf.estimators_]
voting_clf.voting = "soft"
print(voting_clf.voting)
voting_clf.score(X_test_scaled, y_test)
[estimator.score(X_test_scaled, y_test) for estimator in voting_clf.estimators_]
# ## Evaluating the Essemble With Cross-Validation
# y_pred_prob = voting_clf.predict_proba(X_test_scaled)[:,1]
y_scores = cross_val_predict(voting_clf, X_train_scaled, y_train, cv=3, method='predict_proba')
y_train_pred = cross_val_predict(voting_clf, X_train_scaled, y_train, cv=3)
# hack to work around issue #9589 in Scikit-Learn 0.19.0
if y_scores.ndim == 2:
y_scores = y_scores[:, 1]
precisions, recalls, thresholds = precision_recall_curve(y_train, y_scores)
def plot_precision_recall_vs_threshold(precisions, recalls, thresholds):
plt.plot(thresholds, precisions[:-1], "b--", label="Precision")
plt.plot(thresholds, recalls[:-1], "g-", label="Recall")
plt.xlabel("Threshold")
plt.legend(loc="upper left")
plt.ylim([0, 1])
plot_precision_recall_vs_threshold(precisions, recalls, thresholds)
plt.show()
# +
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_train, y_scores)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_train, y_scores)
# +
#print(confusion_matrix(y_test,y_pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_train, y_train_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['Sem Perda','Perda'],
title='Confusion matrix, without normalization')
# -
print(classification_report(y_test, y_pred))
# ## Predicting the Classes in Test Set
y_pred = voting_clf.predict(X_test_scaled)
# +
y_pred_prob = voting_clf.predict_proba(X_test_scaled)[:,1]
# Generate ROC curve values: fpr, tpr, thresholds
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
roc_auc_score(y_test, y_pred_prob)
# +
#print(confusion_matrix(y_test,y_pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['Sem Perda','Perda'],
title='Confusion matrix, without normalization')
# -
print(classification_report(y_test, y_pred))
| Model-Study/mlModelsEssemble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 64-bit
# name: python37364bitc6f8272de35d4e508f36e0915614a098
# ---
# +
import random
class Player():
def __init__(self, player):
self.name = player.get("name", "Dywane")
self.choice = player.get("choice", "Scissors")
choices = ["Rock", "Paper", "Scissors"]
def validate_choice(option):
if option == "0" or option == "1" or option == "2":
return choices[int(option)]
else:
raise Exception("Invalid Option", option)
morris = {"name": "Morris", "choice": random.choice(choices) }
me = {"name": input("what is your name?"), "choice": validate_choice(input("what is your choice? \ 0 = Rock \ 1 = Paper \ 2 = Scissors"))}
player1 = Player(morris)
player2 = Player(me)
def judge_game(player1, player2):
player1_wins = (player1.choice == choices[0] and player2.choice == choices[2]
or player1.choice == choices[1] and player2.choice == choices[0]
or player1.choice == choices[2] and player2.choice == choices[1])
if player1_wins:
return player1.name + " wins with " + player1.choice
return player2.name + " wins with " + player2.choice
def bao_jin_dub(player1, player2):
if player1.choice == player2.choice:
return "Draw"
else:
return judge_game(player1, player2)
bao_jin_dub(player1, player2)
| 2020-01-beginner/scripts/rock.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/facebookresearch/vissl/blob/v0.1.6/tutorials/Installation_V0_1_6.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Zl6yrjD75Fx7"
# Copyright (c) Facebook, Inc. and its affiliates. All Rights Reserved
# + [markdown] id="6XzxTZfKwFNo"
# # Installation
#
# VISSL installation is simple: we provide pre-built binaries (pip, conda) and also instructions for building from source (pip, conda).
#
# + [markdown] id="imQmMr-626YJ"
# ## Requirements
#
# At a high level, project requires following system dependencies.
#
# - Linux
# - Python>=3.6.2 and <3.9
# - PyTorch>=1.4
# - torchvision (matching PyTorch install)
# - CUDA (must be a version supported by the pytorch version)
# - OpenCV
#
# Note our circle-ci runs tests on Pytorch versions 1.6.0 and 1.9.1 and these are our preferred versions. If you are using FSDP using the [fairscale](https://github.com/facebookresearch/fairscale) library, they run their unit tests on Pytorch versions 1.6.0, 1.7.1, and 1.8.1.
#
#
# + [markdown] id="SP8rIA5U3D-1"
# ## Installing VISSL from pre-built binaries
#
# VISSL provides conda and pip binaries pre-built.
#
#
# + [markdown] id="D7hyDr_F3MQw"
# ### Install VISSL conda package
#
# This assumes you have conda 10.2.
#
# ```bash
# conda create -n vissl python=3.8
# conda activate vissl
# conda install -c pytorch pytorch=1.7.1 torchvision cudatoolkit=10.2
# conda install -c vissl -c iopath -c conda-forge -c pytorch -c defaults apex vissl
# ```
#
# For other versions of PyTorch, Python, CUDA, please modify the above instructions with the
# desired version. VISSL provides Apex packages for all combinations of pytorch, python and compatible cuda.
#
# + [markdown] id="PRMtGG1c3QrY"
#
# ### Install VISSL pip package
#
# This example is with pytorch 1.5.1 and cuda 10.1. Please modify the PyTorch version, cuda version and accordingly apex version below for the desired settings.
#
# - We use `PyTorch=1.5.1` with `CUDA 10.1` in the following instruction (user can chose their desired version).
# - There are several ways to install opencv, one possibility is as follows.
# - For APEX, we provide pre-built binary built with optimized C++/CUDA extensions provided by APEX.
# Note that, for the APEX install, you need to get the versions of CUDA, PyTorch, and Python correct in the URL. We provide APEX versions for all possible combinations of Python, PyTorch, and CUDA. Select the right APEX Wheels if you want a different combination.
#
# + id="N2q2fEUp3sqP"
# Install: PyTorch (we assume 1.5.1 but VISSL works with all PyTorch versions >=1.4)
# !pip install torch==1.5.1+cu101 torchvision==0.6.1+cu101 -f https://download.pytorch.org/whl/torch_stable.html
# install opencv
# !pip install opencv-python
# install apex by checking system settings: cuda version, pytorch version, python version
import sys
import torch
version_str="".join([
f"py3{sys.version_info.minor}_cu",
torch.version.cuda.replace(".",""),
f"_pyt{torch.__version__[0:5:2]}"
])
print(version_str)
# install apex (pre-compiled with optimizer C++ extensions and CUDA kernels)
# !pip install apex -f https://dl.fbaipublicfiles.com/vissl/packaging/apexwheels/{version_str}/download.html
# install VISSL
# !pip install vissl
# + [markdown] id="JPhZPxPa3UhL"
# ## Installing VISSL from source
# The following instructions assume that you have a CUDA version installed and working.
#
# + [markdown] id="naGJKTvM3Yon"
# ### Install from source in PIP environment
#
# **Step 1: Create Virtual environment (pip)**
# ```bash
# python3 -m venv ~/venv
# . ~/venv/bin/activate
# ```
#
# **Step 2: Install PyTorch (pip)**
#
# ```bash
# pip install torch==1.7.1+cu101 torchvision==0.8.2+cu101 -f https://download.pytorch.org/whl/torch_stable.html
# ```
#
# **Step 3: Install APEX (pip)**
#
# ```bash
# pip install apex -f https://dl.fbaipublicfiles.com/vissl/packaging/apexwheels/py37_cu101_pyt171/download.html
# ```
#
# **Step 4: Install VISSL**
#
# ```bash
# # clone vissl repository
# # # cd $HOME && git clone --recursive https://github.com/facebookresearch/vissl.git && cd $HOME/vissl/
# # install vissl dependencies
# pip install --progress-bar off -r requirements.txt
# pip install opencv-python
# # update classy vision install to current main.
# pip uninstall -y classy_vision
# pip install classy-vision@https://github.com/facebookresearch/ClassyVision/tarball/main
# # install vissl dev mode (e stands for editable)
# pip install -e .[dev]
# # verify installation
# python -c 'import vissl, apex, cv2'
# ```
#
# + [markdown] id="cdrzFg163bp5"
# ### Install from source in Conda environment
#
# **Step 1: Create Conda environment**
#
# If you don't have anaconda, [run this bash script to install conda](https://github.com/facebookresearch/vissl/blob/master/docker/common/install_conda.sh).
#
# ```bash
# conda create -n vissl_env python=3.7
# source activate vissl_env
# ```
#
# **Step 2: Install PyTorch (conda)**
#
# ```bash
# conda install pytorch torchvision cudatoolkit=10.1 -c pytorch
# ```
#
# **Step 3: Install APEX (conda)**
#
# ```bash
# conda install -c vissl apex
# ```
#
# **Step 4: Install VISSL**
# Follow [step4 instructions from the PIP installation](#step-4-install-vissl-from-source)
#
# That's it! You are now ready to use VISSL.
#
# + [markdown] id="u6Fxe3MWxqsI"
# VISSL should be successfuly installed by now and all the dependencies should be available.
# + id="Np6atgoOTPrA"
import vissl
import tensorboard
import apex
import torch
| tutorials/Installation_V0_1_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# modules required for handling dataframes
import numpy as np
import os
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import matplotlib
import itertools
import seaborn as sns
from decimal import Decimal
# +
#count pivot table of dataframe with taxonomic columns:
#rows - rank names, columns - barcode, sorted by desired class
def generate_ncbi_taxonomy_pivot_k2(tax_df, rank, bcs, num):
"""From tax_df, generate a pivot table listing num rank counts, sorted by bcs"""
pivot_table = tax_df.pivot_table(values='seq_length_k2',
index=rank,
columns='superkingdom_k2',
aggfunc='sum',
fill_value=0)
pivot_table.columns.name = None
pivot_table = pivot_table.sort_values(bcs, axis=0, ascending=False).head(n=num)
return pivot_table
#count pivot table of dataframe with taxonomic columns:
#rows - rank names, columns - barcode, sorted by desired class
def generate_ncbi_taxonomy_pivot_minimap(tax_df, rank, bcs, num):
"""From tax_df, generate a pivot table listing num rank counts, sorted by bcs"""
pivot_table = tax_df.pivot_table(values='seqlen',
index=rank,
columns='kingdom',
aggfunc='count',
fill_value=0)
pivot_table.columns.name = None
pivot_table = pivot_table.sort_values(bcs, axis=0, ascending=False).head(n=num)
return pivot_table
def add_df_headers(df, header_list):
df.columns = header_list
def trim_df_columns(df,column_list):
df.drop(columns=column_list, inplace=True)
# -
# Define some global variables.
sourcedir = '/home/yiheng/MinION_data/' # the directory where all the documents of each sequencing run are stored.
genera_in_mock = ['Aspergillus','Blastobotrys','Candida','Diutina', 'Nakaseomyces', 'Clavispora','Cryptococcus','Cyberlindnera',
'Debaryomyces','Geotrichum','Kluyveromyces','Kodamaea','Lomentospora','Magnusiomyces','Meyerozyma','Pichia',
'Rhodotorula','Scedosporium','Trichophyton', 'Trichosporon', 'Wickerhamomyces','Yarrowia','Zygoascus', 'Purpureocillium']
fungal_phylums = ['Blastocladiomycota', 'Chytridiomycota', 'Glomeromycota', 'Microsporidia',
'Neocallimastigomycota', 'Ascomycota', 'Basidiomycota', 'Mucoromycota']
def get_tab(seq_tech, sample, db, algorithm):
if sample == 'IE':
barcode = '05'
elif sample == 'MC':
barcode = '06'
else:
print('wrong sample name.')
if seq_tech == 'nanopore':
blast_tab_path = os.path.join(sourcedir, 'barcode%s' % barcode, 'barcode%s.%sdb_%s.tab' % (barcode, db, algorithm))
blast_tab_df = pd.read_csv(blast_tab_path, sep='\t', index_col=0, header=0)
elif seq_tech == 'illumina':
blast_tab_path = os.path.join(sourcedir, 'mock_com/illumina', '%s1' % sample, 'finaldf.%s1.%s.%s.tab' % (sample, db, algorithm))
blast_tab_df = pd.read_csv(blast_tab_path, sep='\t', index_col=0, header=0)
else:
print('''Check again your barcode, db or algorithm.
sample must be chosen from 'IE' or 'MC'.
db must be from 'nt' or 'refseq_fungi_updated'.
algorithm must be from 'blast' or 'k2'.''')
return blast_tab_df
# The next step is to calculate the precision and recall rates for each analysis
# For kraken data, the finaldf included all the unclassified reads/contig, so we could calculate recall rates straightaway.
# For blast data, the finaldf only included the classified reads, so we need to call the corresponding kraken finaldf to get the total length of each sample (from reads with Q>7)
# There are quite few if conditions in the function. This is because the final dataframe (tab file) are not consistantly generated.
# This function set an example of how to deal with inconsistant dataframe.
# I could clean up the final tab file from each analysis because I generated those files,
# but this kind of conditions can be useful if the tab file were generated by others.
# Nevertheless, this is a bad function.
def calculate_precision_recall_rates(seq_tech, sample, db, algorithm):
tab_df = get_tab(seq_tech, sample, db, algorithm)
if algorithm == 'k2':
if seq_tech == 'nanopore':
tab_df_classified = tab_df[tab_df.taxid_k2 != 0]
tab_df_classified_true = tab_df_classified[tab_df_classified.genus_k2.isin(genera_in_mock)]
recall_rate = tab_df_classified.seq_length_k2.sum()/tab_df.seq_length_k2.sum()
precision_rate = tab_df_classified_true.seq_length_k2.sum()/tab_df_classified.seq_length_k2.sum()
elif seq_tech == 'illumina':
tab_df_classified = tab_df[tab_df.taxid != 0]
tab_df_classified_true = tab_df_classified[tab_df_classified.genus.isin(genera_in_mock)]
recall_rate = tab_df_classified.total_cov.sum()/tab_df.total_cov.sum()
precision_rate = tab_df_classified_true.total_cov.sum()/tab_df_classified.total_cov.sum()
else:
print('check the sequencing tech you typed in.')
elif algorithm == 'blast':
# this k2 df is used for capture the sequencing length or total coverage.
if db == 'refseq_fungi_updated':
tab_df_k2 = get_tab(seq_tech, sample, 'fungi', 'k2')
else:
tab_df_k2 = get_tab(seq_tech, sample, db, 'k2')
if seq_tech == 'nanopore':
tab_df_classified = tab_df[(tab_df.superkingdom_blast != 'Unclassified') & (~tab_df.superkingdom_blast.isna())]
tab_df_classified_true = tab_df_classified[tab_df_classified.genus_blast.isin(genera_in_mock)]
subset_tab_df_k2 = tab_df_k2[tab_df_k2.read_id_k2.isin(tab_df_classified.qseqid_blast)]
subset_tab_df_k2_true = tab_df_k2[tab_df_k2.read_id_k2.isin(tab_df_classified_true.qseqid_blast)]
recall_rate = subset_tab_df_k2.seq_length_k2.sum()/tab_df_k2.seq_length_k2.sum()
precision_rate = subset_tab_df_k2_true.seq_length_k2.sum()/subset_tab_df_k2.seq_length_k2.sum()
elif seq_tech == 'illumina':
tab_df_classified = tab_df[(tab_df.superkingdom != 'Unclassified') & (~tab_df.superkingdom.isna())]
tab_df_classified_true = tab_df_classified[tab_df_classified.genus.isin(genera_in_mock)]
subset_tab_df_k2 = tab_df_k2[tab_df_k2.contig.isin(tab_df_classified.contig)]
subset_tab_df_k2_true = tab_df_k2[tab_df_k2.contig.isin(tab_df_classified_true.contig)]
recall_rate = subset_tab_df_k2.total_cov.sum()/tab_df_k2.total_cov.sum()
precision_rate = subset_tab_df_k2_true.total_cov.sum()/subset_tab_df_k2.total_cov.sum()
else:
print('check the sequencing tech you typed in.')
else:
print('check the algorithm you typed in.')
return (precision_rate, recall_rate)
MC_precision_recall = pd.DataFrame(columns=['Precision', 'Recall'])
MC_precision_recall.loc[0] = calculate_precision_recall_rates('illumina', 'MC', 'refseq_fungi_updated', 'blast')
MC_precision_recall.loc[1] = calculate_precision_recall_rates('illumina', 'MC', 'nt', 'blast')
MC_precision_recall.loc[2] = calculate_precision_recall_rates('nanopore', 'MC', 'refseq_fungi_updated', 'blast')
MC_precision_recall.loc[3] = calculate_precision_recall_rates('nanopore', 'MC', 'nt', 'blast')
MC_precision_recall.loc[4] = calculate_precision_recall_rates('illumina', 'MC', 'fungi', 'k2')
MC_precision_recall.loc[5] = calculate_precision_recall_rates('illumina', 'MC', 'nt', 'k2')
MC_precision_recall.loc[6] = calculate_precision_recall_rates('nanopore', 'MC', 'fungi', 'k2')
MC_precision_recall.loc[7] = calculate_precision_recall_rates('nanopore', 'MC', 'nt', 'k2')
IE_precision_recall = pd.DataFrame(columns=['Precision', 'Recall'])
IE_precision_recall.loc[0] = calculate_precision_recall_rates('illumina', 'IE', 'refseq_fungi_updated', 'blast')
IE_precision_recall.loc[1] = calculate_precision_recall_rates('illumina', 'IE', 'nt', 'blast')
IE_precision_recall.loc[2] = calculate_precision_recall_rates('nanopore', 'IE', 'refseq_fungi_updated', 'blast')
IE_precision_recall.loc[3] = calculate_precision_recall_rates('nanopore', 'IE', 'nt', 'blast')
IE_precision_recall.loc[4] = calculate_precision_recall_rates('illumina', 'IE', 'fungi', 'k2')
IE_precision_recall.loc[5] = calculate_precision_recall_rates('illumina', 'IE', 'nt', 'k2')
IE_precision_recall.loc[6] = calculate_precision_recall_rates('nanopore', 'IE', 'fungi', 'k2')
IE_precision_recall.loc[7] = calculate_precision_recall_rates('nanopore', 'IE', 'nt', 'k2')
IE_precision_recall['category'] = ['illumina+blastn+fungi', 'illumina+blastn+nt', 'nanopore+blastn+fungi', 'nanopore+blastn+nt',
'illumina+kraken2+fungi', 'illumina+kraken2+nt', 'nanopore+kraken2+fungi', 'nanopore+kraken2+nt']
MC_precision_recall['category'] = ['illumina+blastn+fungi', 'illumina+blastn+nt', 'nanopore+blastn+fungi', 'nanopore+blastn+nt',
'illumina+kraken2+fungi', 'illumina+kraken2+nt', 'nanopore+kraken2+fungi', 'nanopore+kraken2+nt']
markers = ["s", "s", "o", "o", "s", "s", "o", "o"]
edgecolors = ['tab:green', 'tab:green', 'tab:green', 'tab:green', 'tab:red', 'tab:red', 'tab:red', 'tab:red']
facecolors = ['tab:green', 'none', 'tab:green', 'none', 'tab:red', 'none', 'tab:red', 'none']
for i in range(8): #for each of the 7 features
mi = markers[i] #marker for ith feature
xi = IE_precision_recall.Precision[i]*100 #x array for ith feature .. here is where you would generalize different x for every feature
yi = IE_precision_recall.Recall[i]*100 #y array for ith feature
ci = facecolors[i] #color for ith feature
di = edgecolors[i]
ei = IE_precision_recall.category[i]
plt.scatter(xi,yi,marker=mi, facecolors=ci, edgecolors=di, label=ei)
# plt.legend(loc='center left', fontsize=12, bbox_to_anchor=(1, 0.5))
plt.xlabel('Genus level precision (%)', fontsize=14)
plt.ylabel('Genus level recall (%)', fontsize=14)
plt.xlim([65, 100])
plt.ylim([65, 100])
ax = plt.gca()
ax.xaxis.grid(linestyle='dotted')
ax.yaxis.grid(linestyle='dotted')
ax.set_aspect('equal', adjustable='box')
ax.text(95.5, 97, 'PD', fontweight=300, fontsize=18)
ax.text(58, 102, 'B', fontweight=300, fontsize=24)
plt.savefig(os.path.join(sourcedir, 'figures', 'Figure1B.png'), bbox_inches='tight', dpi=600)
plt.show()
for i in range(8): #for each of the 7 features
mi = markers[i] #marker for ith feature
xi = MC_precision_recall.Precision[i]*100 #x array for ith feature .. here is where you would generalize different x for every feature
yi = MC_precision_recall.Recall[i]*100 #y array for ith feature
ci = facecolors[i] #color for ith feature
di = edgecolors[i]
ei = IE_precision_recall.category[i]
plt.scatter(xi,yi,marker=mi, facecolors=ci, edgecolors=di, label=ei)
plt.legend(loc='center left', fontsize=12, bbox_to_anchor=(1, 0.5))
plt.xlabel('Genus level precision (%)', fontsize=14)
plt.ylabel('Genus level recall (%)', fontsize=14)
plt.xlim([65, 100])
plt.ylim([65, 100])
ax = plt.gca()
ax.xaxis.grid(linestyle='dotted')
ax.yaxis.grid(linestyle='dotted')
ax.set_aspect('equal', adjustable='box')
ax.text(95.5, 97, 'PB', fontweight=300, fontsize=18)
ax.text(58, 102, 'C', fontweight=300, fontsize=24)
plt.savefig(os.path.join(sourcedir, 'figures', 'Figure1C.png'), bbox_inches='tight', dpi=600)
def calculate_concordance_algorithmfixed(seq_tech, sample, algorithm):
if algorithm == 'blast':
fungi_df = get_tab(seq_tech, sample, 'refseq_fungi_updated', algorithm)
elif algorithm == 'k2':
fungi_df = get_tab(seq_tech, sample, 'fungi', algorithm)
else:
print('Wrong db name.')
nt_df = get_tab(seq_tech, sample, 'nt', algorithm)
if seq_tech == 'illumina':
fungi_df.columns = ['%s_fungi' % x for x in fungi_df.columns]
nt_df.columns = ['%s_nt' % x for x in nt_df.columns]
merged_df = pd.merge(fungi_df, nt_df, how="outer", left_on='contig_fungi', right_on='contig_nt')
# This line is only for the k2 algorithm,
# As the blast of illumina data has already removed the unmapped contigs.
merged_df_trimed = merged_df[~merged_df.total_cov_fungi.isna()]
# For simplicity, it doesn't need to drop any colomns
# As we have already dropped NaN rows in the total_cov column so we can quickly fill the rest of columns with na
merged_df_trimed = merged_df_trimed.fillna(value='unclassified')
# This is the column that is used for sum up total length
column_to_sum = 'total_cov_fungi'
elif seq_tech == 'nanopore':
if algorithm == 'blast':
fungi_df.columns = [x[:-6] for x in fungi_df.columns]
nt_df.columns = [x[:-6] for x in nt_df.columns]
fungi_df.columns = ['%s_fungi' % x for x in fungi_df.columns]
nt_df.columns = ['%s_nt' % x for x in nt_df.columns]
merged_df = pd.merge(fungi_df, nt_df, how="outer", left_on='qseqid_fungi', right_on='qseqid_nt')
# This is to fill the one column with the value from another
merged_df['sequence_length_template_fungi'] = merged_df['sequence_length_template_fungi'\
].T.fillna(merged_df['sequence_length_template_nt']).T
merged_df['qseqid_fungi'] = merged_df['qseqid_fungi'].T.fillna(merged_df['qseqid_nt']).T
column_to_sum = 'sequence_length_template_fungi'
merged_df_trimed = merged_df#.fillna(value='unclassified')
# merged_df_trimed = merged_df_trimed[(merged_df_trimed.superkingdom_fungi != 'Unclassified')
# | (merged_df_trimed.superkingdom_nt != 'Unclassified')]
elif algorithm == 'k2':
fungi_df.columns = [x[:-3] for x in fungi_df.columns]
nt_df.columns = [x[:-3] for x in nt_df.columns]
fungi_df.columns = ['%s_fungi' % x for x in fungi_df.columns]
nt_df.columns = ['%s_nt' % x for x in nt_df.columns]
merged_df = pd.merge(fungi_df, nt_df, how="outer", left_on='read_id_fungi', right_on='read_id_nt')
column_to_sum = 'seq_length_fungi'
merged_df_trimed = merged_df#.fillna(value='unclassified')
merged_df_trimed = merged_df_trimed[(merged_df_trimed.classification_fungi == 'C')
| (merged_df_trimed.classification_nt == 'C')]
else:
print('Wrong algorithm.')
else:
print('Wrong sequencing tech.')
# This is to get the reads/contigs that classified into the same genus
merged_df_trimed_agreed_classification = merged_df_trimed[merged_df_trimed.genus_fungi ==
merged_df_trimed.genus_nt]
# This is to calculate the overlapped percentage of different algorithms against the same database
concordance = merged_df_trimed_agreed_classification[column_to_sum].sum()/merged_df_trimed[column_to_sum].sum()
return concordance
def trim_merged_df_mixed_algorithm(merged_df, seq_tech):
trimed_merged_df = merged_df[(~merged_df.superkingdom_blast.isna())
| (merged_df.classification_k2 == 'C')]
if seq_tech == 'illumina':
trimed_merged_df = trimed_merged_df.drop(columns=['len_blast', 'ave_cov_blast', 'total_cov_blast', 'evalue_blast',
'pident_blast', 'pmatch_blast', 'len_k2', 'ave_cov_k2', 'classification_k2',
'taxid_k2', 'seqlen_k2', 'kmer_profile_k2'])
# all the contigs that has no coverage means they are false assembly. They won't contribute to the analysis so we get rid of them
trimed_merged_df = trimed_merged_df[~trimed_merged_df.total_cov_k2.isna()]
elif seq_tech == 'nanopore':
trimed_merged_df = trimed_merged_df.drop(columns=['batch_id_blast', 'sequence_length_template_blast', 'scaling_median_template_blast',
'scaling_mad_template_blast', 'pident_blast', 'nident_blast', 'staxids_blast', 'sscinames_blast',
'log_evalue_blast', 'pmatch_blast', 'classification_k2', 'taxid_k2', 'kmer_profile_k2'])
else:
print('Data type not recognized. Can only recognize illumina or nanopore data.')
trimed_merged_df = trimed_merged_df.reset_index(drop=True)
return trimed_merged_df
def calculate_concordance_dbfixed(seq_tech, sample, db):
if db == 'fungi':
blast_df = get_tab(seq_tech, sample, 'refseq_fungi_updated', 'blast')
elif db == 'nt':
blast_df = get_tab(seq_tech, sample, db, 'blast')
else:
print('Wrong db name.')
k2_df = get_tab(seq_tech, sample, db, 'k2')
if seq_tech == 'illumina':
blast_df.columns = ['%s_blast' % x for x in blast_df.columns]
k2_df.columns = ['%s_k2' % x for x in k2_df.columns]
merged_df = pd.merge(blast_df, k2_df, how="outer", left_on='contig_blast', right_on='contig_k2')
# This is the column that is used for sum up total length
column_to_sum = 'total_cov_k2'
elif seq_tech == 'nanopore':
merged_df = pd.merge(blast_df, k2_df, how="outer", left_on='qseqid_blast', right_on='read_id_k2')
column_to_sum = 'seq_length_k2'
else:
print('Wrong sequencing tech.')
# This is to first get the reads/contigs that were both classified by the algorithms
# It's in the triming step.
merged_df_trimed = trim_merged_df_mixed_algorithm(merged_df, seq_tech)
# This is to get the reads/contigs that classified into the same genus
merged_df_trimed_agreed_classification = merged_df_trimed[merged_df_trimed.genus_blast ==
merged_df_trimed.genus_k2]
# This is to calculate the overlapped percentage of different algorithms against the same database
concordance = merged_df_trimed_agreed_classification[column_to_sum].sum()/merged_df_trimed[column_to_sum].sum()
return concordance
Classification_strategy_list = ['Same database different algorithm', 'Same database different algorithm', 'Same algorithm different database', 'Same algorithm different database']
illumina_IE_df = pd.DataFrame()
illumina_IE_df['Concordance'] = [calculate_concordance_algorithmfixed('illumina', 'IE', 'blast'),
calculate_concordance_algorithmfixed('illumina', 'IE', 'k2'),
calculate_concordance_dbfixed('illumina', 'IE', 'fungi'),
calculate_concordance_dbfixed('illumina', 'IE', 'nt')]
illumina_IE_df['Classification strategy'] = Classification_strategy_list
illumina_IE_df['Sequencing strategy'] = 'PD-Illumina'
nanopore_IE_df = pd.DataFrame()
nanopore_IE_df['Concordance'] = [calculate_concordance_algorithmfixed('nanopore', 'IE', 'blast'),
calculate_concordance_algorithmfixed('nanopore', 'IE', 'k2'),
calculate_concordance_dbfixed('nanopore', 'IE', 'fungi'),
calculate_concordance_dbfixed('nanopore', 'IE', 'nt')]
nanopore_IE_df['Classification strategy'] = Classification_strategy_list
nanopore_IE_df['Sequencing strategy'] = 'PD-Nanopore'
illumina_MC_df = pd.DataFrame()
illumina_MC_df['Concordance'] = [calculate_concordance_algorithmfixed('illumina', 'MC', 'blast'),
calculate_concordance_algorithmfixed('illumina', 'MC', 'k2'),
calculate_concordance_dbfixed('illumina', 'MC', 'fungi'),
calculate_concordance_dbfixed('illumina', 'MC', 'nt')]
illumina_MC_df['Classification strategy'] = Classification_strategy_list
illumina_MC_df['Sequencing strategy'] = ' PB-Illumina '
nanopore_MC_df = pd.DataFrame()
nanopore_MC_df['Concordance'] = [calculate_concordance_algorithmfixed('nanopore', 'MC', 'blast'),
calculate_concordance_algorithmfixed('nanopore', 'MC', 'k2'),
calculate_concordance_dbfixed('nanopore', 'MC', 'fungi'),
calculate_concordance_dbfixed('nanopore', 'MC', 'nt')]
nanopore_MC_df['Classification strategy'] = Classification_strategy_list
nanopore_MC_df['Sequencing strategy'] = ' PB-Nanopore '
concordance_df_combined = pd.concat([illumina_IE_df, nanopore_IE_df, illumina_MC_df, nanopore_MC_df], ignore_index=True)
concordance_df_combined.Concordance = concordance_df_combined.Concordance*100
mean_confordance_illumina_IE_algorithmfixed = (calculate_concordance_algorithmfixed('illumina', 'IE', 'blast') +
calculate_concordance_algorithmfixed('illumina', 'IE', 'k2'))/0.02
mean_confordance_illumina_IE_dbfixed = (calculate_concordance_dbfixed('illumina', 'IE', 'fungi') +
calculate_concordance_dbfixed('illumina', 'IE', 'nt'))/0.02
mean_confordance_nanopore_IE_algorithmfixed = (calculate_concordance_algorithmfixed('nanopore', 'IE', 'blast') +
calculate_concordance_algorithmfixed('nanopore', 'IE', 'k2'))/0.02
mean_confordance_nanopore_IE_dbfixed = (calculate_concordance_dbfixed('nanopore', 'IE', 'fungi') +
calculate_concordance_dbfixed('nanopore', 'IE', 'nt'))/0.02
mean_confordance_illumina_MC_algorithmfixed = (calculate_concordance_algorithmfixed('illumina', 'MC', 'blast') +
calculate_concordance_algorithmfixed('illumina', 'MC', 'k2'))/0.02
mean_confordance_illumina_MC_dbfixed = (calculate_concordance_dbfixed('illumina', 'MC', 'fungi') +
calculate_concordance_dbfixed('illumina', 'MC', 'nt'))/0.02
mean_confordance_nanopore_MC_algorithmfixed = (calculate_concordance_algorithmfixed('nanopore', 'MC', 'blast') +
calculate_concordance_algorithmfixed('nanopore', 'MC', 'k2'))/0.02
mean_confordance_nanopore_MC_dbfixed = (calculate_concordance_dbfixed('nanopore', 'MC', 'fungi') +
calculate_concordance_dbfixed('nanopore', 'MC', 'nt'))/0.02
concordance_df_combined
# +
sns.set(style="ticks")
x_labels = ['PD-Illumina', 'PD-Nanopore', ' PB-Illumina ', ' PB-Nanopore ']
x_pos = np.arange(len(x_labels))
ax = sns.swarmplot(x="Sequencing strategy", y="Concordance", hue='Classification strategy', edgecolor='1',
data=concordance_df_combined, palette="tab10", dodge=True)
ax.set_xlabel(' ', fontsize=14)
ax.yaxis.grid(linestyle='dotted')
handles, labels = ax.get_legend_handles_labels()
ax.legend(handles=handles[2:], labels=labels[2:])
ax.set_xticklabels(x_labels, fontsize=14)
plt.xticks(rotation=-15)
ax.yaxis.set_label_text('Genus level concordance (%)', fontsize=14)
plt.ylim(60, 100)
# This is to plot the average/mean lines for each sample
ax.plot([-0.275,-0.125], [mean_confordance_illumina_IE_algorithmfixed,mean_confordance_illumina_IE_algorithmfixed], lw=2, color='k')
ax.plot([0.125,0.275], [mean_confordance_illumina_IE_dbfixed,mean_confordance_illumina_IE_dbfixed], lw=2, color='k')
ax.plot([0.725,0.875], [mean_confordance_nanopore_IE_algorithmfixed,mean_confordance_nanopore_IE_algorithmfixed], lw=2, color='k')
ax.plot([1.125,1.275], [mean_confordance_nanopore_IE_dbfixed,mean_confordance_nanopore_IE_dbfixed], lw=2, color='k')
ax.plot([1.725,1.875], [mean_confordance_illumina_MC_algorithmfixed,mean_confordance_illumina_MC_algorithmfixed], lw=2, color='k')
ax.plot([2.125,2.275], [mean_confordance_illumina_MC_dbfixed,mean_confordance_illumina_MC_dbfixed], lw=2, color='k')
ax.plot([2.725,2.875], [mean_confordance_nanopore_MC_algorithmfixed,mean_confordance_nanopore_MC_algorithmfixed], lw=2, color='k')
ax.plot([3.125,3.275], [mean_confordance_nanopore_MC_dbfixed,mean_confordance_nanopore_MC_dbfixed], lw=2, color='k')
ax.text(-1.05, 102, 'A', fontweight=300, fontsize=24)
#plt.show()
plt.savefig(os.path.join(sourcedir, 'figures', 'Figure1A_all_reads.png'), bbox_inches="tight", dpi=1000)
# -
| Data_analysis/Figure_plotting/Plot_Figure_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: VPython
# language: python
# name: vpython
# ---
# +
import ipywidgets as wd
from vpython import *
# This version uses Jupyter notebook slider
# See Color-RGB-HSV2 for a version that uses VPython slider
scene.userzoom = False
scene.userspin = False
scene.width = 400
scene.height = 200
scene.range = 1
box(pos=vector(10,0,0)) # Force creation of canvas; box is not seen because it is outside the canvas
scene.caption = 'You can Ctrl-C copy the RGB and HSV values shown above.'
C = ['Red', 'Green', 'Blue', 'Hue', 'Saturation', 'Value']
sliders = []
lastvalues = []
adjusting = False
def set_background(change):
global adjusting
if adjusting: return # in the process of resetting various slider values
s = []
N = -1
for i in range(6): # Get values for all 6 sliders
val = sliders[i].value
s.append(val)
if val != lastvalues[i]:
N = i
lastvalues[i] = val
adjusting = True # prevent circular updating of slider values
if N < 3:
rgb = vector(s[0],s[1],s[2])
hsv = color.rgb_to_hsv(rgb)
sliders[3].value = lastvalues[3] = int(1000*hsv.x)/1000 # reset HSV slider positions; display 3 figures
sliders[4].value = lastvalues[4] = int(1000*hsv.y)/1000
sliders[5].value = lastvalues[5] = int(1000*hsv.z)/1000
else:
hsv = vector(s[3],s[4],s[5])
rgb = color.hsv_to_rgb(hsv)
sliders[0].value = lastvalues[0] = int(1000*rgb.x)/1000 # reset RGB slider positions; display 3 figures
sliders[1].value = lastvalues[1] = int(1000*rgb.y)/1000
sliders[2].value = lastvalues[2] = int(1000*rgb.z)/1000
scene.background = rgb
# For readability, limit precision of display of quantities to 3 figures
f = "RGB = <{:1.3f}, {:1.3f}, {:1.3f}>, HSV = <{:1.3f}, {:1.3f}, {:1.3f}>"
scene.title = f.format(rgb.x, rgb.y, rgb.z, hsv.x, hsv.y, hsv.z)
adjusting = False
for i in range(6):
sliders.append(wd.FloatSlider(description=C[i], min=0, max=1, step=0.001, value=0))
lastvalues.append(0)
sliders[i].observe(set_background, names='value') # watch for changes in slider.value
display(wd.VBox(children=sliders)) # stack (default horizontal) sliders vertically
sliders[0].value = 1 # make the background red
# -
| Demos/Color-RGB-HSV1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Generative models
#
# Most **classification algorithms** fall into one of two categories:
# - discriminative classifiers
# - generative classifiers
#
# **Discriminative classifiers** model the target variable, y, as a direct function of the predictor variables, x. Example: logistic regression uses the following model, where 𝜷 is a length-D vector of coefficients and x is a length-D vector of predictors:
# 
#
# **Generative classifiers** instead view the predictors as being generated according to their class — i.e., they see x as a function of y, rather than the other way around. They then use Bayes’ rule to get from p(x|y = k) to P(y = k|x), as explained below.
#
# Generative models can be broken down into the three following steps. Suppose we have a classification task with K unordered classes, represented by k = 1, 2, …, K.
#
# - Estimate the prior probability that a target belongs to any given class. I.e., estimate P(y = k) for k = 1, 2, …, K.
# - Estimate the density of the predictors conditional on the target belonging to each class. I.e., estimate p(x|y = k) for k = 1, 2, …, K.
# - Calculate the posterior probability that the target belongs to any given class. I.e., calculate P(y = k|x), which is proportional to p(x|y = k)P(y = k) by Bayes’ rule.
#
# We then classify an observation as belonging to the class k for which the following expression is greatest:
#
# 
# ### Class Priors estimation for Generative classifiers
#
# Let I_nk be an indicator which equals 1 if y_n = k and 0 otherwise.
#
# 
#
# Our estimate of P(y = k) is just the sample fraction of the observations from class k.
#
# 
# ### Data Likelihood
#
# The next step is to model the conditional distribution of x given y so that we can estimate this distribution’s parameters. This of course depends on the family of distributions we choose to model x. Three common approaches are detailed below.sns.
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_iris
data = load_iris()
X, y = data.data, data.target
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=0.1)
# -
# ### Linear Discriminative Analysis (LDA)
#
# In LDA, we assume the following distribution for x
#
# 
#
# for k = 1, 2, …, K. Note that each class has the same covariance matrix but a unique mean vector.
# +
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
clf = LinearDiscriminantAnalysis()
clf.fit(X_train, Y_train)
y_pred = clf.predict(X_train)
print("Train Accuracy ", np.mean(y_pred == Y_train))
colors = {0: 'r',1: 'g',2: 'b'}
fig, ax = plt.subplots(1, 2)
for x, y in zip(X_train, y_pred):
ax[0].scatter(x[0],x[1],c=colors[y])
ax[0].set_xlabel("predicted")
for x, y in zip(X_train, Y_train):
ax[1].scatter(x[0],x[1],c=colors[y])
ax[1].set_xlabel("real")
plt.show()
# -
# ### Quadratic Discriminant Analysis (QDA)
#
# QDA looks very similar to LDA but assumes each class has its own covariance matrix. I.e.,
#
# 
#
#
# +
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
clf = QuadraticDiscriminantAnalysis()
clf.fit(X_train, Y_train)
y_pred = clf.predict(X_train)
print("Train Accuracy ", np.mean(y_pred == Y_train))
colors = {0: 'r',1: 'g',2: 'b'}
fig, ax = plt.subplots(1, 2)
for x, y in zip(X_train, y_pred):
ax[0].scatter(x[0],x[1],c=colors[y])
ax[0].set_xlabel("predicted")
for x, y in zip(X_train, Y_train):
ax[1].scatter(x[0],x[1],c=colors[y])
ax[1].set_xlabel("real")
plt.show()
# -
# ### Naive Bayes
#
# Naive Bayes assumes the random variables within x are independent conditional on the class of the observation. That is, if x is D-dimensional,
#
# 
# +
from sklearn.naive_bayes import GaussianNB
clf = GaussianNB()
clf.fit(X_train, Y_train)
y_pred = clf.predict(X_train)
print("Train Accuracy ", np.mean(y_pred == Y_train))
colors = {0: 'r',1: 'g',2: 'b'}
fig, ax = plt.subplots(1, 2)
for x, y in zip(X_train, y_pred):
ax[0].scatter(x[0],x[1],c=colors[y])
ax[0].set_xlabel("predicted")
for x, y in zip(X_train, Y_train):
ax[1].scatter(x[0],x[1],c=colors[y])
ax[1].set_xlabel("real")
plt.show()
# -
# ### References
#
# 1. https://www.section.io/engineering-education/linear-discriminant-analysis/
# 2. https://towardsdatascience.com/generative-classification-algorithms-from-scratch-d6bf0a81dcf7
# 3.
| notebooks/02-GenerativeModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.2
# language: julia
# name: julia-0.6
# ---
include("conv_2d.jl")
using Images,TestImages
using NNlib
using BenchmarkTools
img2tensor(img)=convert(Array{Float64},img)
tensor2img(tensor)=convert(Array{ColorTypes.Gray{FixedPointNumbers.Normed{UInt8,8}},2},max.(0.,min.(tensor,1.)))
img =testimage("cameraman")
fil=[-1. -2. -1.; 0. 0. 0.; 1. 2. 1.]'
tensor=img2tensor(img)
dest=zeros(512,512)
apply_filter_on_img!(tensor,fil,dest,(1,1),(1,1))
tensor2img(dest)
@benchmark apply_filter_on_img!($tensor,$fil,$dest,(1,1),(1,1))
imker=centered(fil)
dest2 = imfilter(tensor, imker, Fill(0.))
tensor2img(dest2)
@benchmark imfilter(tensor, imker, Fill(0.))
img_lena =testimage("lena")
ch_lena=channelview(img_lena)
tensor_lena=zeros(256,256,3,2)
for i=1:2
for j=1:3
tensor_lena[:,:,j,i]=ch_lena[j,:,:]
end
end
fil_lena=rand(3,3,3,4)
dest_lena=zeros(256,256,4,2)
conv_2d!(tensor_lena,fil_lena,dest_lena,(1,1),(1,1))
@benchmark conv_2d!($tensor_lena,$fil_lena,$dest_lena,(1,1),(1,1))
NNlib.conv2d!(dest_lena,tensor_lena,fil_lena,padding=1,stride=1);
@benchmark NNlib.conv2d!($dest_lena,$tensor_lena,$fil_lena,padding=1,stride=1)
using ProfileView
Profile.clear()
@profile conv_2d!(tensor_lena,fil_lena,dest_lena,(1,1),(1,1))
ProfileView.view()
cld(1,1)
A=rand(100,100)
@benchmark for i=1:100 for j=1:100 A[i,j] end end
eachindex(A)
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# Again, this code is coming mostly from the book "Grokking Machine Learning"
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import numpy as np
import random
import turicreate as tc
data = pd.DataFrame({
"tjilp":[1,0,2,1,1,2,3,2,4,2,3,4,4,3],
"mwah": [0,2,1,2,3,2,2,3,4,4,1,2,0,3],
"mood": ["Sad", "Sad", "Sad", "Sad", "Happy", "Happy", "Sad", "Happy", "Happy", "Happy", "Sad", "Happy", "Happy","Sad"]
})
data["label"]=data["mood"].apply(lambda x: 1 if x == "Happy" else 0)
happy_sentence = data[data["mood"] == "Happy"]
sad_sentence = data[data["mood"] == "Sad"]
data
# +
def sigmoid(x):
return np.exp(x)/(1+np.exp(x))
def score(weights, bias, features):
return np.dot(weights, features) + bias
def prediction(weights, bias, features):
return sigmoid(score(weights, bias, features))
def log_loss(weights, bias, features, label):
pred = prediction(weights, bias, features)
return -label*np.log(pred) - (1-label)*np.log(1-pred)
def total_log_loss(weights, bias, features, labels):
total_error = 0
for i in range(len(features)):
total_error += log_loss(weights, bias, features[i], labels[i])
return total_error
def logistic_trick(weights, bias, features, label, learning_rate = 0.01):
pred = prediction(weights, bias, features)
for i in range(len(weights)):
weights[i] += (label-pred)*features[i]*learning_rate
bias += (label-pred)*learning_rate
return weights, bias
def logistic_regression_algorithm(features, labels, learning_rate = 0.01, epochs = 1000):
weights = [1 for i in range(len(features[0]))]
bias = 0.0
errors = []
for i in range(epochs):
errors.append(total_log_loss(weights, bias, features, labels))
j = random.randint(0, len(features)-1)
weights, bias = logistic_trick(weights, bias, features[j], labels[j])
return weights, bias
# -
features = data[["tjilp","mwah"]].to_numpy()
labels = data["label"].to_numpy()
found_weights, found_bias = logistic_regression_algorithm(features, labels)
# +
def plot_sentiment(happy_data, sad_data, line = []):
tick_spacing = 1
fig, ax = plt.subplots(1,1)
ax.scatter(happy_data["tjilp"], happy_data["mwah"], c='g',marker='o', label='Happy')
ax.scatter(sad_data["tjilp"], sad_data["mwah"], c='r',marker='x', label='Sad')
if line and len(line) > 1:
ax.plot(line[0],line[1])
ax.xaxis.set_major_locator(ticker.MultipleLocator(tick_spacing))
plt.title('Happy or Sad sentence')
plt.ylabel('Mwah')
plt.xlabel('Tjilp')
plt.rcParams["figure.figsize"] = (8,6)
plt.legend()
plt.grid()
plt.show()
def calculate_x2 (x1, weights, bias):
return (-1*weights[0] * x1 - bias)/weights[1]
# -
x_2_4 = calculate_x2(4,found_weights, found_bias)
x_2_0 = calculate_x2(0, found_weights, found_bias)
plot_sentiment(happy_sentence, sad_sentence, [[0, 4],[x_2_0, x_2_4]])
| 4_logistic_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lustraka/puzzles/blob/main/AoC2021/Optimization.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="dDZWBB4Ll10C"
# # Advent of Code Puzzles
# [Advent of Code 2021](https://adventofcode.com/2021) | [reddit/adventofcode](https://www.reddit.com/r/adventofcode/)
# + id="ri3O_1i3l2mo"
import requests
import pandas as pd
import numpy as np
path = 'https://raw.githubusercontent.com/lustraka/puzzles/main/AoC2021/data/'
# + [markdown] id="TnFAd8h1l8NI"
# ## Day 7: The Treachery of Whales
# - [The Gaussian hare and the Laplacian tortoise: computability of squared-error versus absolute-error estimators](https://projecteuclid.org/journals/statistical-science/volume-12/issue-4/The-Gaussian-hare-and-the-Laplacian-tortoise--computability-of/10.1214/ss/1030037960.full)
# - [L1 regression estimates median whereas L2 regression estimates mean?](https://stats.stackexchange.com/questions/34613/l1-regression-estimates-median-whereas-l2-regression-estimates-mean)
# + id="HZXVSS9bl736" outputId="a4610eb4-86b1-487c-8c62-f3a91c6aca6b" colab={"base_uri": "https://localhost:8080/"}
def parse(data):
print(data[-10:])
return np.array(data.split(',')).astype(int)
example = """16,1,2,0,4,2,7,1,2,14"""
data = parse(example)
data
# + [markdown] id="M02h9rbdmDnJ"
# ### Linear algebra ([numpy.linalg](https://numpy.org/doc/stable/reference/routines.linalg.html?highlight=linalg#module-numpy.linalg))
# The NumPy linear algebra functions rely on BLAS and LAPACK to provide efficient low level implementations of standard linear algebra algorithms.
#
# `linalg.norm(x, ord=None, axis=None, keepdims=False)` computes up to 8 matrix or vector norms (see `ord` param notes). I am not sure if computing some norm of a vector helps to solve this puzzle.
# + id="pTEs_AiymGZg" outputId="351ac9ab-9146-4407-e824-8839a94fdb54" colab={"base_uri": "https://localhost:8080/"}
from numpy import linalg as LA
# Calculate norm of a vector
for ord in [None, np.inf, -np.inf, 0, 1, -1, 2, -2]:
print(ord, '\t:', LA.norm(data, ord))
# + [markdown] id="wN8Z34nNpFKP"
# ### Logistic Regression
# + id="cIv0u3Mhma-J" outputId="7d6e221b-1d2f-4cfd-b8e9-8dc123f54388" colab={"base_uri": "https://localhost:8080/"}
from sklearn.linear_model import LogisticRegression
# Fit a logit classifier
clf = LogisticRegression(penalty='l1', solver='liblinear')
clf.fit(data.reshape(-1,1), range(len(data)))
print(clf.get_params())
print(clf.coef_)
# + id="rONmDtFEpUgp"
| AoC2021/Optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/priya-saw-1999/letsupgrade-python-b7/blob/master/Day2assignmentb7.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="wUfBcV0mrpCR" colab_type="code" colab={}
#ASSIGNMENT1
# + [markdown] id="wUHkkQ9HNSWU" colab_type="text"
# LIST
# + id="j7-sQwqgwKQD" colab_type="code" colab={}
lst=[100,99,123456,20.567]
# + id="FcOqdJ9IDJV-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dfe595f6-6749-4736-a1a3-c96a9f21ecd9"
lst
# + id="d4hKg3UgFbQy" colab_type="code" colab={}
lst.append("priya")
# + id="q0-XCHLMFoZC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cbd5c5bd-8c89-4a1d-8f4b-bc77b3d0dcac"
lst
# + id="-X9AQhdPG7om" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3db7cafd-b9b9-44be-d668-09ed894e2fab"
lst.index("priya")
# + id="B9JKefTxHULw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cc4a9be3-1ca2-4737-dc53-c4916060a8b8"
lst.count("123456")
# + id="foDKX7kWsRKG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2522ed51-00fc-46f5-c44d-ec1be78d5d2d"
lst
# + id="hAvTXotmtwCp" colab_type="code" colab={}
lst.clear()
# + id="xfJgt7Dkt_pn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="487db118-cd87-4462-ce69-90cea3d843ef"
lst
# + id="BA5R6KgTubLL" colab_type="code" colab={}
lst=[12,35,96,56,23,67,8]
# + id="nC7PXZxBu0dB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1a568647-f4f1-488d-bc93-d8e48f267f1f"
lst
# + id="4NlMm2q6u6TE" colab_type="code" colab={}
lst.sort()
# + id="naAyswcVvCS2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ff039862-e0e1-4b34-8396-49f61dfa9974"
lst
# + id="WimWBpV7vv_1" colab_type="code" colab={}
lst.insert(5,60)
# + id="v_Hwm-QAwAK0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="67656281-262e-475e-815e-5d83ef5a8680"
lst
# + [markdown] id="WH4D50v5PAz8" colab_type="text"
# DICTIONARY
# + id="JhR5ftUl7Kfy" colab_type="code" colab={}
Dict = {"Englishmarks":85, "computer": "90", "PCM": [79, 80, 95], "COLLEGE": "STJOSEPH"}
# + id="QPgu0iuU77jk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="e963f758-04ed-448f-a391-4098955a1741"
Dict
# + id="aZjhme9A82An" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="322b0362-3af7-4f94-9857-ee864b9994a4"
Dict["COLLEGE",]
# + id="V2X0JERd9Etm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="358089fb-04af-418d-bd3b-089049f69f4c"
Dict.keys()
# + id="AzS7zuCZ9aWo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="a7f9a076-3ebb-4ae7-b3cf-a3ace8c94b08"
Dict.values()
# + id="VQ8C4FGE9fC5" colab_type="code" colab={}
Dict['electronic']='99'
# + id="YDjad1Ob9_wZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="4a2a1ff8-fd3f-4672-dd93-84f966b06950"
Dict
# + id="vbTPhdKLAGGO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9c6a79b2-651f-4388-8628-67b4f1f964fa"
Dict['PCM']
# + id="FNgE-4vDAcO9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aeccbe0b-cbbb-42d7-a5c3-551154dcc783"
'Englishmarks' in dict
# + [markdown] id="Rd1HK4_oBaEK" colab_type="text"
# SETS
# + id="IkJTlMZYBbXk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="2177e50a-5324-4147-b2b9-efb2ba7e31a7"
set1 = {"pop", "rock", "soul", "hard rock", "rock", "R&B", "rock", "disco"}
set1
# + id="DSR_UbBFB94n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="bafedf34-c945-494a-b879-a9726dc399ee"
Days={"Monday", "Friday", "Wednesday", "Saturday", "Tuesday","Monday","Friday"}
Days
# + id="3WCLcFFlCauc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="46a8eaea-b35e-4294-ad6e-e2a13cf1b902"
Days.add("Sunday")
Days
# + id="tU4rld0NDI_O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="34f65aec-83ca-4953-dc4d-ff48232284f9"
Days.remove("Monday")
Days
# + id="R2hPAuKqDSp7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="50dc0c42-bad7-42be-f8ac-2a20e53fe87e"
"Monday" in Days
# + id="Ugk6aYSkDd69" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="0b2580d8-f343-414e-9945-2c91d5b20145"
Days={"sunday","monday","tuesday"}
Months={"january","February","march"}
Days,Months
# + id="Uc0g9xrjENzX" colab_type="code" colab={}
Set1={1,5,7,9,6,4,5,9,6,4,3,5,6}
Set2={6,4,7,9,5,4,9,3,2,0,9,7,5}
# + id="wmBEoci2FS1H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="98fa4099-8679-49d9-b78d-4dfa3cc7af56"
Set1.union(Set2)
# + id="9g3VDuV-F3KC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9bf6f5b2-fb50-4ab3-a5cf-df7bfc382645"
Set1.intersection(Set2)
# + id="nH570nXMF-EK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="857577b6-dcfb-4fad-b171-bf4073865e28"
Set2.intersection(Set1)
# + id="bmwehsXwGGPZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d7b8a50f-cf30-4137-da27-d1f8ecdc75fa"
set(Set1).issuperset(Set2)
# + id="gusYoGwKGeyP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f96be6da-6729-4f4c-d3c3-59cf7dcb3409"
set(Set1).issubset(Set2)
# + [markdown] id="ECajyrGUG25e" colab_type="text"
# TUPLE
# + id="vQw0MWf4G77n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="23b229b0-8455-40c0-e2af-dc96278a0bfd"
tup1=(9,6,5,7,4,3,2,1,10,25,14,13,12,19,18,17,16,15)
tup1
# + id="9gYNev3qIuH-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="4bf1f5b9-685b-4570-f1c2-a46bd2933408"
tup1.sort()
tup1
# + [markdown] id="p-E1XXXIKQRc" colab_type="text"
# STRINGS
# + id="D-h7yScPKPj2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 476} outputId="57228576-08d0-4cd4-f58f-84a5d6b27444"
str="q","w","e","r","t","y","u","i","o","p","a","s","d","f","g","h" "i","j","j","k","l","z","x","c","v","b","n","m"
str
# + id="wBnVcCihQckL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1fe0e4d2-e3aa-477f-ba80-f6fc1d9a7685"
print(len(str))
| Day2assignmentb7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
salaries = pd.read_csv('data/baseballdatabank-master/core/Salaries.csv')
salaries[salaries['yearID'] == 2002].sort_values('salary',ascending=False).head(20)
salaries[salaries['yearID'] == 2002].sort_values('salary',ascending=False).head(23)['playerID'].to_list()
salaries[(salaries['teamID'].str.contains('OAK') == True) & (salaries['yearID'] == 2002)]
salaries[(salaries['playerID'].str.contains('hatte') == True) & (salaries['yearID'] == 2002)]
| Salary_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # An Introduction to the Amazon SageMaker IP Insights Algorithm
# #### Unsupervised anomaly detection for susicipous IP addresses
# -------
# 1. [Introduction](#Introduction)
# 2. [Setup](#Setup)
# 3. [Training](#Training)
# 4. [Inference](#Inference)
# 5. [Epilogue](#Epilogue)
#
# ## Introduction
# -------
#
# The Amazon SageMaker IP Insights algorithm uses statistical modeling and neural networks to capture associations between online resources (such as account IDs or hostnames) and IPv4 addresses. Under the hood, it learns vector representations for online resources and IP addresses. This essentially means that if the vector representing an IP address and an online resource are close together, then it is likey for that IP address to access that online resource, even if it has never accessed it before.
#
# In this notebook, we use the Amazon SageMaker IP Insights algorithm to train a model on synthetic data. We then use this model to perform inference on the data and show how to discover anomalies. After running this notebook, you should be able to:
#
# - obtain, transform, and store data for use in Amazon SageMaker,
# - create an AWS SageMaker training job to produce an IP Insights model,
# - use the model to perform inference with an Amazon SageMaker endpoint.
#
# If you would like to know more, please check out the [SageMaker IP Inisghts Documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/ip-insights.html).
#
# ## Setup
# ------
# *This notebook was created and tested on a ml.m4.xlarge notebook instance.*
#
# Our first step is to setup our AWS credentials so that AWS SageMaker can store and access training data and model artifacts.
#
# ### Select Amazon S3 Bucket
# We first need to specify the locations where we will store our training data and trained model artifacts. ***This is the only cell of this notebook that you will need to edit.*** In particular, we need the following data:
#
# - `bucket` - An S3 bucket accessible by this account.
# - `prefix` - The location in the bucket where this notebook's input and output data will be stored. (The default value is sufficient.)
# +
import boto3
import botocore
import os
import sagemaker
bucket = sagemaker.Session().default_bucket()
prefix = 'sagemaker/ipinsights-tutorial'
execution_role = sagemaker.get_execution_role()
# check if the bucket exists
try:
boto3.Session().client('s3').head_bucket(Bucket=bucket)
except botocore.exceptions.ParamValidationError as e:
print('Hey! You either forgot to specify your S3 bucket'
' or you gave your bucket an invalid name!')
except botocore.exceptions.ClientError as e:
if e.response['Error']['Code'] == '403':
print("Hey! You don't have permission to access the bucket, {}.".format(bucket))
elif e.response['Error']['Code'] == '404':
print("Hey! Your bucket, {}, doesn't exist!".format(bucket))
else:
raise
else:
print('Training input/output will be stored in: s3://{}/{}'.format(bucket, prefix))
# -
# ### Dataset
#
# Apache Web Server ("httpd") is the most popular web server used on the internet. And luckily for us, it logs all requests processed by the server - by default. If a web page requires HTTP authentication, the Apache Web Server will log the IP address and authenticated user name for each requested resource.
#
# The [access logs](https://httpd.apache.org/docs/2.4/logs.html) are typically on the server under the file `/var/log/httpd/access_log`. From the example log output below, we see which IP addresses each user has connected with:
#
# ```
# 192.168.1.100 - user1 [15/Oct/2018:18:58:32 +0000] "GET /login_success?userId=1 HTTP/1.1" 200 476 "-" "Mozilla/5.0 (Macintosh; Intel Mac OS X 10_12_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/69.0.3497.100 Safari/537.36"
# 192.168.1.102 - user2 [15/Oct/2018:18:58:35 +0000] "GET /login_success?userId=2 HTTP/1.1" 200 - "-" "Mozilla/5.0 (Macintosh; Intel Mac OS X 10_12_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/69.0.3497.100 Safari/537.36"
# ...
# ```
#
# If we want to train an algorithm to detect suspicious activity, this dataset is ideal for SageMaker IP Insights.
#
# First, we determine the resource we want to be analyzing (such as a login page or access to a protected file). Then, we construct a dataset containing the history of all past user interactions with the resource. We extract out each 'access event' from the log and store the corresponding user name and IP address in a headerless CSV file with two columns. The first column will contain the user identifier string, and the second will contain the IPv4 address in decimal-dot notation.
#
# ```
# user1, 192.168.1.100
# user2, 192.168.127.12
# ...
# ```
#
# As a side note, the dataset should include all access events. That means some `<user_name, ip_address>` pairs will be repeated.
#
# #### User Activity Simulation
# For this example, we are going to simulate our own web-traffic logs. We mock up a toy website example and simulate users logging into the website from mobile devices.
#
# The details of the simulation are explained in the script [here](./generate_data.py).
#
#
# Install dependency
# !pip install tqdm
# +
from generate_data import generate_dataset
# We simulate traffic for 10,000 users. This should yield about 3 million log lines (~700 MB).
NUM_USERS = 10000
log_file = 'ipinsights_web_traffic.log'
generate_dataset(NUM_USERS, log_file)
# Visualize a few log lines
# !head $log_file
# -
# ### Prepare the dataset
# Now that we have our logs, we need to transform them into a format that IP Insights can use. As we mentioned above, we need to:
# 1. Choose the resource which we want to analyze users' history for
# 2. Extract our users' usage history of IP addresses
# 3. In addition, we want to separate our dataset into a training and test set. This will allow us to check for overfitting by evaluating our model on 'unseen' login events.
#
# For the rest of the notebook, we assume that the Apache Access Logs are in the Common Log Format as defined by the [Apache documentation](https://httpd.apache.org/docs/2.4/logs.html#accesslog). We start with reading the logs into a Pandas DataFrame for easy data exploration and pre-processing.
# +
import pandas as pd
df = pd.read_csv(
log_file,
sep=" ",
na_values='-',
header=None,
names=['ip_address','rcf_id','user','timestamp','time_zone','request', 'status', 'size', 'referer', 'user_agent']
)
df.head()
# -
# We convert the log timestamp strings into Python datetimes so that we can sort and compare the data more easily.
# Convert time stamps to DateTime objects
df['timestamp'] = pd.to_datetime(df['timestamp'], format='[%d/%b/%Y:%H:%M:%S')
# We also verify the time zones of all of the time stamps. If the log contains more than one time zone, we would need to standardize the timestamps.
# Check if they are all in the same timezone
num_time_zones = len(df['time_zone'].unique())
num_time_zones
# As we see above, there is only one value in the entire `time_zone` column. Therefore, all of the timestamps are in the same time zone, and we do not need to standardize them. We can skip the next cell and go to [1. Selecting a Resource](#1.-Select-Resource).
#
# If there is more than one time_zone in your dataset, then we parse the timezone offset and update the corresponding datetime object.
#
# **Note:** The next cell takes about 5-10 minutes to run.
# +
from datetime import datetime
import pytz
def apply_timezone(row):
tz = row[1]
tz_offset = int(tz[:3]) * 60 # Hour offset
tz_offset += int(tz[3:5]) # Minutes offset
return row[0].replace(tzinfo=pytz.FixedOffset(tz_offset))
if num_time_zones > 1:
df['timestamp'] = df[['timestamp','time_zone']].apply(apply_timezone, axis=1)
# -
# #### 1. Select Resource
# Our goal is to train an IP Insights algorithm to analyze the history of user logins such that we can predict how suspicious a login event is.
#
# In our simulated web server, the server logs a `GET` request to the `/login_success` page everytime a user successfully logs in. We filter our Apache logs for `GET` requests for `/login_success`. We also filter for requests that have a `status_code == 200`, to ensure that the page request was well formed.
#
# **Note:** every web server handles logins differently. For your dataset, determine which resource you will need to be analyzing to correctly frame this problem. Depending on your usecase, you may need to do more data exploration and preprocessing.
df = df[(df['request'].str.startswith('GET /login_success')) & (df['status'] == 200)]
# #### 2. Extract Users and IP address
# Now that our DataFrame only includes log events for the resource we want to analyze, we extract the relevant fields to construct a IP Insights dataset.
#
# IP Insights takes in a headerless CSV file with two columns: an entity (username) ID string and the IPv4 address in decimal-dot notation. Fortunately, the Apache Web Server Access Logs output IP addresses and authentcated usernames in their own columns.
#
# **Note:** Each website handles user authentication differently. If the Access Log does not output an authenticated user, you could explore the website's query strings or work with your website developers on another solution.
df = df[['user', 'ip_address', 'timestamp']]
# #### 3. Create training and test dataset
# As part of training a model, we want to evaluate how it generalizes to data it has never seen before.
#
# Typically, you create a test set by reserving a random percentage of your dataset and evaluating the model after training. However, for machine learning models that make future predictions on historical data, we want to use out-of-time testing. Instead of randomly sampling our dataset, we split our dataset into two contiguous time windows. The first window is the training set, and the second is the test set.
#
# We first look at the time range of our dataset to select a date to use as the partition between the training and test set.
df['timestamp'].describe()
# We have login events for 10 days. Let's take the first week (7 days) of data as training and then use the last 3 days for the test set.
# +
time_partition = datetime(2018, 11, 11, tzinfo=pytz.FixedOffset(0)) if num_time_zones > 1 else datetime(2018, 11, 11)
train_df = df[df['timestamp'] <= time_partition]
test_df = df[df['timestamp'] > time_partition]
# -
# Now that we have our training dataset, we shuffle it.
#
# Shuffling improves the model's performance since SageMaker IP Insights uses stochastic gradient descent. This ensures that login events for the same user are less likely to occur in the same mini batch. This allows the model to improve its performance in between predictions of the same user, which will improve training convergence.
# Shuffle train data
train_df = train_df.sample(frac=1)
train_df.head()
# ### Store Data on S3
# Now that we have simulated (or scraped) our datasets, we have to prepare and upload it to S3.
#
# We will be doing local inference, therefore we don't need to upload our test dataset.
# Output dataset as headerless CSV
train_data = train_df.to_csv(index=False, header=False, columns=['user', 'ip_address'])
# +
# Upload data to S3 key
train_data_file = 'train.csv'
key = os.path.join(prefix, 'train', train_data_file)
s3_train_data = 's3://{}/{}'.format(bucket, key)
print('Uploading data to: {}'.format(s3_train_data))
boto3.resource('s3').Bucket(bucket).Object(key).put(Body=train_data)
# Configure SageMaker IP Insights Input Channels
input_data = {
'train': sagemaker.session.s3_input(s3_train_data, distribution='FullyReplicated', content_type='text/csv')
}
# -
# ## Training
# ---
# Once the data is preprocessed and available in the necessary format, the next step is to train our model on the data. There are number of parameters required by the SageMaker IP Insights algorithm to configure the model and define the computational environment in which training will take place. The first of these is to point to a container image which holds the algorithms training and hosting code:
# +
from sagemaker.amazon.amazon_estimator import get_image_uri
image = get_image_uri(boto3.Session().region_name, 'ipinsights')
# -
# Then, we need to determine the training cluster to use. The IP Insights algorithm supports both CPU and GPU training. We recommend using GPU machines as they will train faster. However, when the size of your dataset increases, it can become more economical to use multiple CPU machines running with distributed training. See [Recommended Instance Types](https://docs.aws.amazon.com/sagemaker/latest/dg/ip-insights.html#ip-insights-instances) for more details.
#
# ### Training Job Configuration
# - **train_instance_type**: the instance type to train on. We recommend `p3.2xlarge` for single GPU, `p3.8xlarge` for multi-GPU, and `m5.2xlarge` if using distributed training with CPU;
# - **train_instance_count**: the number of worker nodes in the training cluster.
#
# We need to also configure SageMaker IP Insights-specific hypeparameters:
#
# ### Model Hyperparameters
# - **num_entity_vectors**: the total number of embeddings to train. We use an internal hashing mechanism to map the entity ID strings to an embedding index; therefore, using an embedding size larger than the total number of possible values helps reduce the number of hash collisions. We recommend this value to be 2x the total number of unique entites (i.e. user names) in your dataset;
# - **vector_dim**: the size of the entity and IP embedding vectors. The larger the value, the more information can be encoded using these representations but using too large vector representations may cause the model to overfit, especially for small training data sets;
# - **num_ip_encoder_layers**: the number of layers in the IP encoder network. The larger the number of layers, the higher the model capacity to capture patterns among IP addresses. However, large number of layers increases the chance of overfitting. `num_ip_encoder_layers=1` is a good value to start experimenting with;
# - **random_negative_sampling_rate**: the number of randomly generated negative samples to produce per 1 positive sample; `random_negative_sampling_rate=1` is a good value to start experimenting with;
# - Random negative samples are produced by drawing each octet from a uniform distributed of [0, 255];
# - **shuffled_negative_sampling_rate**: the number of shuffled negative samples to produce per 1 positive sample; `shuffled_negative_sampling_rate=1` is a good value to start experimenting with;
# - Shuffled negative samples are produced by shuffling the accounts within a batch;
#
# ### Training Hyperparameters
# - **epochs**: the number of epochs to train. Increase this value if you continue to see the accuracy and cross entropy improving over the last few epochs;
# - **mini_batch_size**: how many examples in each mini_batch. A smaller number improves convergence with stochastic gradient descent. But a larger number is necessary if using shuffled_negative_sampling to avoid sampling a wrong account for a negative sample;
# - **learning_rate**: the learning rate for the Adam optimizer (try ranges in [0.001, 0.1]). Too large learning rate may cause the model to diverge since the training would be likely to overshoot minima. On the other hand, too small learning rate slows down the convergence;
# - **weight_decay**: L2 regularization coefficient. Regularization is required to prevent the model from overfitting the training data. Too large of a value will prevent the model from learning anything;
#
# For more details, see [Amazon SageMaker IP Insights (Hyperparameters)](https://docs.aws.amazon.com/sagemaker/latest/dg/ip-insights-hyperparameters.html). Additionally, most of these hyperparameters can be found using SageMaker Automatic Model Tuning; see [Amazon SageMaker IP Insights (Model Tuning)](https://docs.aws.amazon.com/sagemaker/latest/dg/ip-insights-tuning.html) for more details.
# +
# Set up the estimator with training job configuration
ip_insights = sagemaker.estimator.Estimator(
image,
execution_role,
train_instance_count=1,
train_instance_type='ml.p3.2xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sagemaker.Session())
# Configure algorithm-specific hyperparameters
ip_insights.set_hyperparameters(
num_entity_vectors='20000',
random_negative_sampling_rate='5',
vector_dim='128',
mini_batch_size='1000',
epochs='5',
learning_rate='0.01',
)
# Start the training job (should take about ~1.5 minute / epoch to complete)
ip_insights.fit(input_data)
# -
# If you see the message
#
# > Completed - Training job completed
#
# at the bottom of the output logs then that means training successfully completed and the output of the SageMaker IP Insights model was stored in the specified output path. You can also view information about and the status of a training job using the AWS SageMaker console. Just click on the "Jobs" tab and select training job matching the training job name, below:
print('Training job name: {}'.format(ip_insights.latest_training_job.job_name))
# ## Inference
# -----
#
# Now that we have trained a SageMaker IP Insights model, we can deploy the model to an endpoint to start performing inference on data. In this case, that means providing it a `<user, IP address>` pair and predicting their compatability scores.
#
# We can create an inference endpoint using the SageMaker Python SDK `deploy()`function from the job we defined above. We specify the instance type where inference will be performed, as well as the initial number of instnaces to spin up. We recommend using the `ml.m5` instance as it provides the most memory at the lowest cost. Verify how large your model is in S3 and pick the instance type with the appropriate amount of memory.
predictor = ip_insights.deploy(
initial_instance_count=1,
instance_type='ml.m5.xlarge'
)
# Congratulations, you now have a SageMaker IP Insights inference endpoint! You could start integrating this endpoint with your production services to start querying incoming requests for abnormal behavior.
#
# You can confirm the endpoint configuration and status by navigating to the "Endpoints" tab in the AWS SageMaker console and selecting the endpoint matching the endpoint name below:
print('Endpoint name: {}'.format(predictor.endpoint))
# ### Data Serialization/Deserialization
# We can pass data in a variety of formats to our inference endpoint. In this example, we will pass CSV-formmated data. Other available formats are JSON-formated and JSON Lines-formatted. We make use of the SageMaker Python SDK utilities: `csv_serializer` and `json_deserializer` when configuring the inference endpoint
# +
from sagemaker.predictor import csv_serializer, json_deserializer
predictor.content_type = 'text/csv'
predictor.serializer = csv_serializer
predictor.accept = 'application/json'
predictor.deserializer = json_deserializer
# -
# Now that the predictor is configured, it is as easy as passing in a matrix of inference data.
# We can take a few samples from the simulated dataset above, so we can see what the output looks like.
inference_data = [(data[0], data[1]) for data in train_df[:5].values]
predictor.predict(inference_data)
# By default, the predictor will only output the `dot_product` between the learned IP address and the online resource (in this case, the user ID). The dot product summarizes the compatibility between the IP address and online resource. The larger the value, the more the algorithm thinks the IP address is likely to be used by the user. This compatability score is sufficient for most applications, as we can define a threshold for what we constitute as an anomalous score.
#
# However, more advanced users may want to inspect the learned embeddings and use them in further applications. We can configure the predictor to provide the learned embeddings by specifing the `verbose=True` parameter to the Accept heading. You should see that each 'prediction' object contains three keys: `ip_embedding`, `entity_embedding`, and `dot_product`.
predictor.accept = 'application/json; verbose=True'
predictor.predict(inference_data)
# ## Compute Anomaly Scores
# ----
# The `dot_product` output of the model provides a good measure of how compatible an IP address and online resource are. However, the range of the dot_product is unbounded. This means to be able to consider an event as anomolous we need to define a threshold. Such that when we score an event, if the dot_product is above the threshold we can flag the behavior as anomolous.However, picking a threshold can be more of an art, and a good threshold depends on the specifics of your problem and dataset.
#
# In the following section, we show how to pick a simple threshold by comparing the score distributions between known normal and malicious traffic:
# 1. We construct a test set of 'Normal' traffic;
# 2. Inject 'Malicious' traffic into the dataset;
# 3. Plot the distribution of dot_product scores for the model on 'Normal' trafic and the 'Malicious' traffic.
# 3. Select a threshold value which separates the normal distribution from the malicious traffic threshold. This value is based on your false-positive tolerance.
#
# ### 1. Construct 'Normal' Traffic Dataset
#
# We previously [created a test set](#3.-Create-training-and-test-dataset) from our simulated Apache access logs dataset. We use this test dataset as the 'Normal' traffic in the test case.
test_df.head()
# ### 2. Inject Malicious Traffic
# If we had a dataset with enough real malicious activity, we would use that to determine a good threshold. Those are hard to come by. So instead, we simulate malicious web traffic that mimics a realistic attack scenario.
#
# We take a set of user accounts from the test set and randomly generate IP addresses. The users should not have used these IP addresses during training. This simulates an attacker logging in to a user account without knowledge of their IP history.
# +
import numpy as np
from generate_data import draw_ip
# We only need the dot product. Let's reset the predictor output type.
predictor.accept = 'application/json; verbose=False'
def score_ip_insights(predictor, df):
def get_score(result):
"""Return the negative to the dot product of the predictions from the model."""
return [-prediction["dot_product"] for prediction in result["predictions"]]
df = df[['user', 'ip_address']]
result = predictor.predict(df.values)
return get_score(result)
def create_test_case(train_df, test_df, num_samples, attack_freq):
"""Creates a test case from provided train and test data frames.
This generates test case for accounts that are both in training and testing data sets.
:param train_df: (panda.DataFrame with columns ['user', 'ip_address']) training DataFrame
:param test_df: (panda.DataFrame with columns ['user', 'ip_address']) testing DataFrame
:param num_samples: (int) number of test samples to use
:param attack_freq: (float) the ratio of negative_samples:positive_samples to generate for test case
:return: DataFrame with both good and bad traffic, with labels
"""
# Get all possible accounts. The IP Insights model can only make predictions on users it has seen in training
# Therefore, filter the test dataset for unseen accounts, as their results will not mean anything.
valid_accounts = set(train_df['user'])
valid_test_df = test_df[test_df['user'].isin(valid_accounts)]
good_traffic = valid_test_df.sample(num_samples, replace=False)
good_traffic = good_traffic[['user', 'ip_address']]
good_traffic['label'] = 0
# Generate malicious traffic
num_bad_traffic = int(num_samples * attack_freq)
bad_traffic_accounts = np.random.choice(list(valid_accounts), size=num_bad_traffic, replace=True)
bad_traffic_ips = [draw_ip() for i in range(num_bad_traffic)]
bad_traffic = pd.DataFrame({'user': bad_traffic_accounts, 'ip_address': bad_traffic_ips})
bad_traffic['label'] = 1
# All traffic labels are: 0 for good traffic; 1 for bad traffic.
all_traffic = good_traffic.append(bad_traffic)
return all_traffic
# -
NUM_SAMPLES = 100000
test_case = create_test_case(train_df, test_df, num_samples=NUM_SAMPLES, attack_freq=1)
test_case.head()
test_case_scores = score_ip_insights(predictor, test_case)
# ### 3. Plot Distribution
#
# Now, we plot the distribution of scores. Looking at this distribution will inform us on where we can set a good threshold, based on our risk tolerance.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
n, x = np.histogram(test_case_scores[:NUM_SAMPLES], bins=100, density=True)
plt.plot(x[1:], n)
n, x = np.histogram(test_case_scores[NUM_SAMPLES:], bins=100, density=True)
plt.plot(x[1:], n)
plt.legend(["Normal", "Random IP"])
plt.xlabel("IP Insights Score")
plt.ylabel("Frequency")
plt.figure()
# -
# ### 4. Selecting a Good Threshold
#
# As we see in the figure above, there is a clear separation between normal traffic and random traffic.
# We could select a threshold depending on the application.
#
# - If we were working with low impact decisions, such as whether to ask for another factor or authentication during login, we could use a `threshold = 0.0`. This would result in catching more true-positives, at the cost of more false-positives.
#
# - If our decision system were more sensitive to false positives, we could choose a larger threshold, such as `threshold = 10.0`. That way if we were sending the flagged cases to manual investigation, we would have a higher confidence that the acitivty was suspicious.
# +
threshold = 0.0
flagged_cases = test_case[np.array(test_case_scores) > threshold]
num_flagged_cases = len(flagged_cases)
num_true_positives = len(flagged_cases[flagged_cases['label'] == 1])
num_false_positives = len(flagged_cases[flagged_cases['label'] == 0])
num_all_positives = len(test_case.loc[test_case['label'] == 1])
print("When threshold is set to: {}".format(threshold))
print("Total of {} flagged cases".format(num_flagged_cases))
print("Total of {} flagged cases are true positives".format(num_true_positives))
print("True Positive Rate: {}".format(num_true_positives/float(num_flagged_cases)))
print("Recall: {}".format(num_true_positives/float(num_all_positives)))
print("Precision: {}".format(num_true_positives/float(num_flagged_cases)))
# -
# ## Epilogue
# ----
#
# In this notebook, we have showed how to configure the basic training, deployment, and usage of the Amazon SageMaker IP Insights algorithm. All SageMaker algorithms come with support for two additional services that make optimizing and using the algorithm that much easier: Automatic Model Tuning and Batch Transform service.
#
#
# ### Amazon SageMaker Automatic Model Tuning
# The results above were based on using the default hyperparameters of the SageMaker IP Insights algorithm. If we wanted to improve the model's performance even more, we can use [Amazon SageMaker Automatic Model Tuning](https://docs.aws.amazon.com/sagemaker/latest/dg/automatic-model-tuning.html) to automate the process of finding the hyperparameters.
#
# #### Validation Dataset
# Previously, we separated our dataset into a training and test set to validate the performance of a single IP Insights model. However, when we do model tuning, we train many IP Insights models in parallel. If we were to use the same test dataset to select the best model, we bias our model selection such that we don't know if we selected the best model in general, or just the best model for that particular dateaset.
#
# Therefore, we need to separate our test set into a validation dataset and a test dataset. The validation dataset is used for model selection. Then once we pick the model with the best performance, we evaluate it the winner on a test set just as before.
#
# #### Validation Metrics
# For SageMaker Automatic Model Tuning to work, we need an objective metric which determines the performance of the model we want to optimize. Because SageMaker IP Insights is an usupervised algorithm, we do not have a clearly defined metric for performance (such as percentage of fraudulent events discovered).
#
# We allow the user to provide a validation set of sample data (same format as training data bove) through the `validation` channel. We then fix the negative sampling strategy to use `random_negative_sampling_rate=1` and `shuffled_negative_sampling_rate=0` and generate a validation dataset by assigning corresponding labels to the real and simulated data. We then calculate the model's `descriminator_auc` metric. We do this by taking the model's predicted labels and the 'true' simulated labels and compute the Area Under ROC Curve (AUC) on the model's performance.
#
# We set up the `HyperParameterTuner` to maximize the `discriminator_auc` on the validation dataset. We also need to set the search space for the hyperparameters. We give recommended ranges for the hyperparmaeters in the [Amazon SageMaker IP Insights (Hyperparameters)](https://docs.aws.amazon.com/sagemaker/latest/dg/ip-insights-hyperparameters.html) documentation.
#
test_df['timestamp'].describe()
# The test set we constructed above spans 3 days. We reserve the first day as the validation set and the subsequent two days for the test set.
# +
time_partition = datetime(2018, 11, 13, tzinfo=pytz.FixedOffset(0)) if num_time_zones > 1 else datetime(2018, 11, 13)
validation_df = test_df[test_df['timestamp'] < time_partition]
test_df = test_df[test_df['timestamp'] >= time_partition]
valid_data = validation_df.to_csv(index=False, header=False, columns=['user', 'ip_address'])
# -
# We then upload the validation data to S3 and specify it as the validation channel.
# +
# Upload data to S3 key
validation_data_file = 'valid.csv'
key = os.path.join(prefix, 'validation', validation_data_file)
boto3.resource('s3').Bucket(bucket).Object(key).put(Body=valid_data)
s3_valid_data = 's3://{}/{}'.format(bucket, key)
print('Validation data has been uploaded to: {}'.format(s3_valid_data))
# Configure SageMaker IP Insights Input Channels
input_data = {
'train': s3_train_data,
'validation': s3_valid_data
}
# +
from sagemaker.tuner import HyperparameterTuner, IntegerParameter
# Configure HyperparameterTuner
ip_insights_tuner = HyperparameterTuner(
estimator=ip_insights, # previously-configured Estimator object
objective_metric_name='validation:discriminator_auc',
hyperparameter_ranges={'vector_dim': IntegerParameter(64, 1024)},
max_jobs=4,
max_parallel_jobs=2)
# Start hyperparameter tuning job
ip_insights_tuner.fit(input_data, include_cls_metadata=False)
# +
# Wait for all the jobs to finish
ip_insights_tuner.wait()
# Visualize training job results
ip_insights_tuner.analytics().dataframe()
# -
# Deploy best model
tuned_predictor = ip_insights_tuner.deploy(
initial_instance_count=1,
instance_type='ml.m4.xlarge',
content_type='text/csv',
serializer=csv_serializer,
accept='application/json',
deserializer=json_deserializer
)
# Make a prediction against the SageMaker endpoint
tuned_predictor.predict(inference_data)
# We should have the best performing model from the training job! Now we can determine thresholds and make predictions just like we did with the inference endpoint [above](#Inference).
# ### Batch Transform
# Let's say we want to score all of the login events at the end of the day and aggregate flagged cases for investigators to look at in the morning. If we store the daily login events in S3, we can use IP Insights with [Amazon SageMaker Batch Transform](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-batch.html) to run inference and store the IP Insights scores back in S3 for future analysis.
#
# Below, we take the training job from before and evaluate it on the validation data we put in S3.
# +
transformer = ip_insights.transformer(
instance_count=1,
instance_type='ml.m4.xlarge',
)
transformer.transform(
s3_valid_data,
content_type='text/csv',
split_type='Line'
)
# -
# Wait for Transform Job to finish
transformer.wait()
print("Batch Transform output is at: {}".format(transformer.output_path))
# ### Stop and Delete the Endpoint
# If you are done with this model, then we should delete the endpoint before we close the notebook. Or else you will continue to pay for the endpoint while it is running.
#
# To do so execute the cell below. Alternately, you can navigate to the "Endpoints" tab in the SageMaker console, select the endpoint with the name stored in the variable endpoint_name, and select "Delete" from the "Actions" dropdown menu.
ip_insights_tuner.delete_endpoint()
sagemaker.Session().delete_endpoint(predictor.endpoint)
| introduction_to_amazon_algorithms/ipinsights_login/ipinsights-tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
from sklearn import datasets
dataset = datasets.load_iris()
x = dataset.data
y = dataset.target
# + pycharm={"name": "#%%\n"}
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
knn.fit(x, y)
# + pycharm={"name": "#%%\n"}
from joblib import dump, load
dump(knn, 'iris_model_knn.joblib')
| Flask-Project/Iris-Pred-Flutter/model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Objectives:
# 1)To visualize and analyze the time series data.
# 2)To develop a predictive model using Support Vector Machines.
#
#Importing all relevant libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.colors as mcolors
import random
import math
import time
from sklearn.model_selection import RandomizedSearchCV, train_test_split
from sklearn.svm import SVR
from sklearn.metrics import mean_squared_error, mean_absolute_error
import datetime
import operator
plt.style.use('seaborn')
#reading all relevant datasets
confirmedcases=pd.read_csv(r'C:\Users\Administrator\Desktop\time_series_covid_19_confirmed.csv')
recoveredcases=pd.read_csv(r'C:\Users\Administrator\Desktop\time_series_covid_19_recovered.csv')
expiredcases=pd.read_csv(r'C:\Users\Administrator\Desktop\time_series_covid_19_deaths.csv')
# +
#Data wrangling to make sure dataset is in required shape
confirmed_cases=confirmedcases.drop(columns=['Lat', 'Long'])
recovered_cases=recoveredcases.drop(columns=['Lat', 'Long'])
expired_cases=expiredcases.drop(columns=['Lat', 'Long'])
confirmed_cases["Province/State"].fillna("For all provinces combined", inplace = True)
recovered_cases["Province/State"].fillna("For all provinces combined", inplace = True)
expired_cases["Province/State"].fillna("For all provinces combined", inplace = True)
columns=confirmed_cases.keys()
confirmed=confirmed_cases.loc[:,columns[2]:columns[-1]]
recovered=recovered_cases.loc[:,columns[2]:columns[-1]]
expired=expired_cases.loc[:,columns[2]:columns[-1]]
dates=confirmed.keys()
totalcases=[]
totalrecoveries=[]
totaldeaths=[]
for i in dates:
confirmedcount=confirmed[i].sum()
totalcases.append(confirmedcount)
recoveredcount=recovered[i].sum()
totalrecoveries.append(recoveredcount)
expiredcount=expired[i].sum()
totaldeaths.append(expiredcount)
# +
#total cases till 17th May 2020 globally:
confirmedcount
# -
#total recoveries till 17th May 2020 globally:
recoveredcount
#total deaths till 17th May 2020 globally:
expiredcount
# +
#Converting all the appended data to numpy arrays
days_since_1_22_2020=np.array([i for i in range(len(dates))]).reshape(-1,1)
totalcases=np.array(totalcases).reshape(-1,1)
totalrecoveries=np.array(totalrecoveries).reshape(-1,1)
totaldeaths=np.array(totaldeaths).reshape(-1,1)
#days_since_1_22_2020=np.ravel(days_since_1_22_2020,order='C').reshape(-1,1)
#totalcases=np.ravel(totalcases,order='C').reshape(-1,1)
# +
#building dates array of forecast for next 60 days
daysinfuture=60
futureforecast=np.array([i for i in range (len(dates)+daysinfuture)]).reshape(-1,1)
adjusted_dates=futureforecast[:-60]
firstdate='5/17/2020'
#converting into datetime format from int type
start_date=datetime.datetime.strptime(firstdate,'%m/%d/%Y')
futureforecastdates=[]
for i in range(len(futureforecast)):
futureforecastdates.append((start_date+datetime.timedelta(days=i)).strftime('%m/%d/%Y'))
# +
#for visualizations of data till 17th May 2020 we tinker with our dataset to compute certain statistics in order to understand it better
latest_confirmed_cases=confirmedcases[dates[-1]]
latest_recovered_cases=recoveredcases[dates[-1]]
latest_expired_cases=expiredcases[dates[-1]]
#----confirmed positive cases for each country----
countries=list(confirmedcases['Country/Region'].unique())
countrywise_confirmed_cases=[]
zerocases=[]
for i in countries:
cases = latest_confirmed_cases[confirmedcases['Country/Region']==i].sum()
if cases > 0:
countrywise_confirmed_cases.append(cases)
else:
zerocases.append(i)
for i in zerocases:
countries.remove(i)
countries=[k for k, v in sorted(zip(countries,countrywise_confirmed_cases), key=operator.itemgetter(1),reverse=True)] #sorts in accordance to magnitude
for i in range(len(countries)):
countrywise_confirmed_cases[i] = latest_confirmed_cases[confirmedcases['Country/Region']==countries[i]].sum()
print("Confirmed cases by Country are:")
for i in range(len(countries)):
print(f'{countries[i]}:{countrywise_confirmed_cases[i]} case(s)')
# +
#----recovered cases for each country----
countries=list(recoveredcases['Country/Region'].unique())
countrywise_recovered_cases=[]
zerocases=[]
for i in countries:
cases = latest_recovered_cases[recoveredcases['Country/Region']==i].sum()
if cases > 0:
countrywise_recovered_cases.append(cases)
else:
zerocases.append(i)
for i in zerocases:
countries.remove(i)
for i in range(len(countries)):
countrywise_recovered_cases[i] = latest_recovered_cases[recovered_cases['Country/Region']==countries[i]].sum()
print("Recovered cases by Country are:")
for i in range(len(countries)):
print(f'{countries[i]}:{countrywise_recovered_cases[i]} case(s)')
# +
#----death cases for each country----
countries=list(expiredcases['Country/Region'].unique())
countrywise_expired_cases=[]
zerocases=[]
for i in countries:
cases = latest_expired_cases[expiredcases['Country/Region']==i].sum()
if cases > 0:
countrywise_expired_cases.append(cases)
else:
zerocases.append(i)
for i in zerocases:
countries.remove(i)
for i in range(len(countries)):
countrywise_expired_cases[i] = latest_expired_cases[expiredcases['Country/Region']==countries[i]].sum()
print("Death cases by Country are:")
for i in range(len(countries)):
print(f'{countries[i]}:{countrywise_expired_cases[i]} case(s)')
# +
#plotting the no of positive cases per country's bar graph against the total no of cases
countries=list(confirmedcases['Country/Region'].unique())
for i in range(len(countries)):
countrywise_confirmed_cases[i] = latest_confirmed_cases[confirmedcases['Country/Region']==countries[i]].sum()
plt.figure(figsize=(10,32))
plt.barh(countries,countrywise_confirmed_cases)
plt.title("Number of confirmed cases countrywise",size=15)
plt.xlabel("Number of confirmed cases")
plt.show()
# +
#Visualizations for top 25 countries with most positive confirmed cases using Bar graph
visual_unique_countries=[]
visual_confirmed_cases=[]
latest_confirmed_cases=confirmedcases[dates[-1]]
latest_recovered_cases=recoveredcases[dates[-1]]
latest_expired_cases=expiredcases[dates[-1]]
#----confirmed positive cases for each country----
countries=list(confirmedcases['Country/Region'].unique())
countrywise_confirmed_cases=[]
zerocases=[]
for i in countries:
cases = latest_confirmed_cases[confirmedcases['Country/Region']==i].sum()
if cases > 0:
countrywise_confirmed_cases.append(cases)
else:
zerocases.append(i)
for i in zerocases:
countries.remove(i)
countries=[k for k, v in sorted(zip(countries,countrywise_confirmed_cases), key=operator.itemgetter(1),reverse=True)] #sorts in accordance to magnitude
for i in range(len(countries)):
countrywise_confirmed_cases[i] = latest_confirmed_cases[confirmedcases['Country/Region']==countries[i]].sum()
others=np.sum(countrywise_confirmed_cases[25:]) #the countries that are not in top 25
for i in range(len(countrywise_confirmed_cases[:25])):
visual_unique_countries.append(countries[i])
visual_confirmed_cases.append(countrywise_confirmed_cases[i])
plt.figure(figsize=(10,32))
plt.barh(visual_unique_countries,visual_confirmed_cases)
plt.title("Number of confirmed cases countrywise",size=15)
plt.xlabel("Number of confirmed cases")
plt.show()
# -
#Visualization of top 25 countries with respect to total confirmed positive cases via Pie chart
plt.figure(figsize=(10,32))
c=random.choices(list(mcolors.CSS4_COLORS.values()),k= len(countries))
plt.pie(visual_confirmed_cases,colors=c)
plt.title("Positive confirmed cases countrywise for top 25 nations")
plt.legend(visual_unique_countries[0:],loc='best')
plt.show()
# +
#Implementing a SVM model for predictive analysis:
kernel = ['poly','sigmoid','rbf']
c = [0.01, 0.1, 1, 10]
gamma = [0.01, 0.1, 1]
epsilon = [0.01, 0.1 , 1]
shrinking = [True, False]
svm_grid = {'kernel':kernel, 'C':c, 'gamma':gamma, 'epsilon':epsilon, 'shrinking':shrinking}
svm = SVR()
svm_search = RandomizedSearchCV(svm,svm_grid,scoring='neg_mean_squared_error',cv=3,return_train_score=True,n_jobs=-1,n_iter=40,verbose=1)
X_train_confirmed, X_test_confirmed, y_train_confirmed, y_test_confirmed = train_test_split(days_since_1_22_2020, totalcases, test_size=0.20, shuffle=False)
svm_search.fit(X_train_confirmed,y_train_confirmed)
# -
svm_search.best_params_
svm_confirmed = svm_search.best_estimator_
svm_confirmed
svm_pred = svm_confirmed.predict(futureforecast)
svm_pred
# +
#Checking against test data
svm_test_pred = svm_confirmed.predict(X_test_confirmed)
plt.plot(svm_test_pred)
plt.plot(y_test_confirmed)
#printing the Mean absolute and Mean squared errors:
print('MAE:',mean_absolute_error(svm_test_pred,y_test_confirmed))
print('MSE:',mean_squared_error(svm_test_pred,y_test_confirmed))
# +
#Total number of cases over time:
plt.figure(figsize=(10,32))
plt.plot(adjusted_dates,totalcases)
plt.title("Number of cases over time",size=30)
plt.xlabel("Days since 17/05/2020",size=30)
plt.ylabel("Number of cases",size=30)
plt.xticks(size=15)
plt.yticks(size=15)
plt.show()
# +
#Confirmed vs Predicted cases:
plt.figure(figsize=(10,32))
plt.plot(adjusted_dates,totalcases)
plt.plot(futureforecast,svm_pred, linestyle='dashed', color='red')
plt.title("Number of cases over time",size=30)
plt.xlabel("Days since 17/05/2020",size=30)
plt.ylabel("Number of cases",size=30)
plt.legend(['Confirmed Cases','SVM Predictions'])
plt.xticks(size=15)
plt.yticks(size=15)
plt.show()
# +
#Forecast of confirmed cases over the next 60 days using SVM:
print("SVM's next 6 months prediction is as follows:")
set(zip(futureforecastdates[:-10],svm_pred[:-10]))
# -
# Key Insights/Takeaways:
# 1)
# 2)
# 3)
#
#
| COVID-19 Outspread Predictive Model using SVM (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!pip install seaborn
import sympy as sym
from sympy import apart
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
sns.set()
sns.set_style("whitegrid", {'grid.linestyle': '--'})
sym.init_printing()
# %matplotlib inline
# -
# # Respuesta dinámica
#
# Ya se ha establecido que el primer paso para analizar un sistema es obtener un modelo.
#
# En el área de la **ingeniería de control**, se configura una base de comparación del comportamiento de sistemas controlados contra señales de prueba particulares.
# ## Señales de prueba
#
# Las señales de prueba más comunes corresponden a funciones impulso, escalón, rampa, parábola, etc.
#
# Como ya se ha estudiado, la respuesta ante la señal impulso permite caracterizar completamente al sistema, sin embargo, no es posible obtener esta señal en laboratorio.
#
# La señal de prueba más apropiada para un sistema debe determinarse por la forma más frecuente de las excitaciones en su operación normal.
#
# ### Impulso
# Entendido como la derivada del escalón.
# \begin{equation}
# \delta(t) = \frac{d\epsilon}{dt}
# \end{equation}
#
# Aplicando la transformada de Laplace se obtiene:
# \begin{equation}
# \mathcal{L} \{ \delta(t) \} = 1
# \end{equation}
#
# Determina completamente el comportamiento del sistema.
#
# ### Escalón
# \begin{equation}
# \epsilon(t) = \begin{cases} 0 & t<0 \\ 1 & t > 0 \end{cases}
# \end{equation}
#
# Aplicando la transformada de Laplace se obtiene:
# \begin{equation}
# \mathcal{L} \{ \epsilon(t) \} = \frac{1}{s}
# \end{equation}
#
# Apropiada para visualizar el comportamiento de sistemas que buscan mantener variables fijas.
#
# ### Rampa
# Entendida como la integral del escalón.
#
# \begin{equation}
# rampa(t) = \int_{-\infty}^{t} \epsilon(\tau) d\tau = t \cdot \epsilon(t)
# \end{equation}
#
# Aplicando la transformada de Laplace se obtiene:
# \begin{equation}
# \mathcal{L} \{ rampa(t) \} = \frac{1}{s^2}
# \end{equation}
#
# Apropiada para visualizar el comportamiento de sistemas que buscan variables que cambian con velocidad constante.
#
# ### Parábola
# Entendida como la integral de la rampa.
#
# \begin{equation}
# parabola(t) = \int_{-\infty}^{t} rampa(\tau) d\tau = \frac{t^2}{2} \cdot \epsilon(t)
# \end{equation}
#
# Aplicando la transformada de Laplace se obtiene:
# \begin{equation}
# \mathcal{L} \{ parabola(t) \} = \frac{1}{s^3}
# \end{equation}
#
# Apropiada para visualizar el comportamiento de sistemas que buscan variables que cambian con aceleración constante.
#
# ## Respuesta transitoria y respuesta estacionaria
#
# Cuando un [sistema estable](https://en.wikipedia.org/wiki/BIBO_stability) es excitado en el instante $t=0$, este responde de manera continua (respuesta transitoria) desde un estado inicial relacionado con las condiciones iniciales hasta que logra equilibrio (respuesta estacionaria).
#
#
# 
# ## Sistemas de primer orden
#
# Considere la siguiente función de transferencia:
#
# 
#
# \begin{equation}
# G(s) = \frac{C(s)}{R(s)} = \frac{a}{b s + c}
# \end{equation}
#
# Este sistema tiene $3$ parámetros ($a$, $b$ y $c$), sin embargo, para facilitar el análisis de estos sistemas puede transformarse la función a su forma canónica al dividir numerador y denominador por $c$.
#
# \begin{equation}
# G(s) = \frac{C(s)}{R(s)} = \frac{\frac{a}{c}}{\frac{b}{c} s + \frac{c}{c}}= \frac{k}{\tau s + 1}
# \end{equation}
#
#
# En la forma canónica, se definen los parámetro como:
# - $k$: ganancia en estado estacionario.
# - $\tau$: constante de tiempo.
# ### Respuesta al escalón
#
# Si el sistema es excitado con un escalón unitario, la señal de salida en el dominio de la frecuencia es:
#
# \begin{equation}
# C(s) = G(s) \cdot R(s) = \frac{k}{\tau s + 1} \cdot \frac{1}{s}
# \end{equation}
#
# Observe que esta expresión puede reescrbirse como:
#
# \begin{equation}
# C(s)= k \cdot \left ( \frac{1}{s} - \frac{\tau}{\tau s + 1} \right )
# \end{equation}
#
# La transformada inversa de Laplace permite obtener una expresión para la señal de salida en el dominio del tiempo $c(t)$.
#
# \begin{equation}
# c(t)= k \left ( 1 - e^{-t/\tau} \right ) \epsilon(t)
# \end{equation}
#
# Observe que:
#
# - $c(t<0)=0$ (condción inicial).
# - $\lim_{t\rightarrow \infty} c(t)=k$ (respuesta estacionaria).
# - La respuesta transitoria es dominada por $e^{-t/\tau}$.
# - $c(\tau) = 0.632k$, el sistema alcanza el $63.2\%$ del valor final después de un tiempo $\tau$.
# - $c(2\tau) = 0.865k$, el sistema alcanza el $86.5\%$ del valor final después de un tiempo $2\tau$.
# - $c(3\tau) = 0.95k$, el sistema alcanza el $95\%$ del valor final después de un tiempo $3\tau$.
# - $\frac{dc}{dt}\big\vert_{t = 0^+} = \frac{d}{dt} \left ( k \left ( 1 - e^{-t/\tau} \right ) \right ) \big\vert_{t = 0^+} = \frac{k}{\tau} e^{-t/\tau} \big\vert_{t = 0^+} =\frac{k}{\tau}$
# - El sistema tiene un polo ubicado en $s=\frac{-1}{\tau}$
#
# +
k, tau = sym.symbols('k, tau',real=True, positive=True)
t = sym.symbols('t',real=True, positive=True)
s = sym.Symbol('s')
def L(f):
return sym.laplace_transform(f, t, s, noconds=True)
def invL(F):
return sym.inverse_laplace_transform(F, s, t)
# -
# la entrada r(t) es escalon unitario
r = sym.Heaviside(t)
r
# la entrada R(s) es la transformada de Laplace de r(t)
R = L(r)
R
# Se define la función de transferencia del sistema G(s)
G = k/(tau*s + 1)
g =invL(G)
G
# la salida se calcula como C(s) = G(s)R(s)
C = G*R
C
# la salida c(t) es la transformada inversa de Laplace de C(s)
c = invL(C)
c
# Hasta este punto se ha encontrado la respuesta general de un sistema de primer orden ante una entrada escalón. Se recomienda variar los parámetros del sistema para identificar cómo varía su comportamiento.
# Para valores particulares de k y tau
K = 1
T = 1
kt = {k: K, tau: T}
ckt = c.subs(kt)
ckt
plt.rcParams['figure.figsize'] = 9, 3
g0 = sym.plot(r,(t,0,10),show = False, line_color='green');
g1 = sym.plot(ckt, (t,0,10),show = False, line_color='blue')
g2 = sym.plot(t*K/T,(t,0,T),show = False, line_color='red');
g0.extend(g1)
g0.extend(g2)
g0.show()
kt
# ### Respesta ante la rampa
#
# Si el sistema es excitado con una rampa unitaria, la señal de salida en el dominio de la frecuencia es:
#
# \begin{equation}
# C(s) = G(s) \cdot R(s) = \frac{k}{\tau s + 1} \cdot \frac{1}{s^2}
# \end{equation}
#
# Observe que esta expresión puede reescrbirse como:
#
# \begin{equation}
# C(s)= k \cdot \left ( \frac{1}{s^2} - \frac{\tau}{s} + \frac{\tau^2}{\tau s + 1} \right )
# \end{equation}
#
# La transformada inversa de Laplace permite obtener una expresión para la señal de salida en el dominio del tiempo $c(t)$.
#
# \begin{equation}
# c(t)= k \left ( t - \tau + \tau e^{-t/\tau} \right ) \epsilon(t)
# \end{equation}
#
# Observe que:
#
# - $c(t)$ se compone de una rampa ($t$), una constante ($\tau$) y una exponencial ($\tau e^{-t/\tau}$).
# - $c(t<0)=0$ (condción inicial).
# - A medida que $t\rightarrow \infty$, $\tau e^{-t/\tau} \rightarrow 0$, por lo que $\lim_{t\rightarrow \infty} c(t)=k \left ( t - \tau \right )$, es decir hace un seguimiento de la rampa con una desviación que depende de $k$ y $\tau$.
#
# - $\lim_{t\rightarrow \infty} \frac{dc}{dt} = \lim_{t\rightarrow \infty} k \left (1 - e^{-t/\tau} \right ) = k$
#
#
# la salida se calcula como C(s) = G(s)R(s)
C_ramp = G*(1/s**2)
C_ramp
# la salida c_ramp(t) es la transformada inversa de Laplace de C(s)
c_ramp = invL(C_ramp)
c_ramp
# Para valores particulares de k y tau
K = 1.3
T = 2
kt = {k: K, tau: T}
c_rampkt = c_ramp.subs(kt)
c_rampkt
plt.rcParams['figure.figsize'] = 9, 3
g0 = sym.plot(t,(t,0,10),show = False, line_color='green');
g1 = sym.plot(c_rampkt, (t,0,10),show = False, line_color='blue')
g2 = sym.plot(t*K,(t,0,10),show = False, line_color='red');
g0.extend(g1)
g0.extend(g2)
g0.show()
kt
# +
from ipywidgets import interact
evalfimpulse = sym.lambdify((k, tau, t), g , 'numpy')
evalfstep = sym.lambdify((k, tau, t), c, 'numpy')
evalframp = sym.lambdify((k, tau, t), c_ramp, 'numpy')
ts = np.linspace(0, 10)
def orden1(tau_in, K_in):
plt.figure(figsize=(7, 5))
ax_impulse = plt.subplot2grid((3, 2), (0, 0))
ax_step = plt.subplot2grid((3, 2), (1, 0))
ax_ramp = plt.subplot2grid((3, 2), (2, 0))
ax_complex = plt.subplot2grid((2, 2), (0, 1), rowspan=2)
ax_impulse.plot(ts, evalfimpulse(K_in, tau_in, ts))
ax_impulse.set_title('Resp. al impulso')
ax_impulse.set_ylim(0, 10)
tau_height = 1 - np.exp(-1)
ax_step.set_title('Resp. al escalón')
ax_step.plot(ts, evalfstep(K_in, tau_in, ts),ts,K_in*ts**0)
ax_step.set_ylim(0, 10)
ax_ramp.set_title('Resp. a la rampa')
ax_ramp.plot(ts, evalframp(K_in, tau_in, ts),ts,K_in*ts)
ax_ramp.set_ylim(0, 100)
ax_complex.set_title('Polos y ceros')
ax_complex.scatter(-1/tau_in, 0, marker='x', s=30)
ax_complex.axhline(0, color='black')
ax_complex.axvline(0, color='black')
ax_complex.axis([-10, 10, -10, 10])
interact(orden1, tau_in=(0.1, 5), K_in=(0.1, 10.));
# -
# Una vez analizadas las simulaciones anteriores, comparta sus ideas para obtener modelos de primer orden de forma experimental.
# ## Sistemas de segundo orden
#
# Considere la siguiente función de transferencia:
#
# 
#
# \begin{equation}
# G(s) = \frac{C(s)}{R(s)} = \frac{a}{b s^2 + c s + d}
# \end{equation}
#
# Este sistema tiene $4$ parámetros ($a$, $b$, $c$ y $d$), sin embargo, para facilitar el análisis de estos sistemas, puede transformarse la función a su forma canónica al dividir numerador y denominador por $b$.
#
# \begin{equation}
# G(s) = \frac{C(s)}{R(s)} = \frac{\frac{a}{b}}{\frac{b}{b} s^2 + \frac{c}{b} s + \frac{d}{b}} = \frac{k\omega_0^2}{s^2 + 2 \zeta\omega_0s + \omega_0^2}
# \end{equation}
#
# En la forma canónica, se definen los parámetro como:
# - $k$: ganancia en estado estacionario.
# - $\omega_0$: frecuencia natural.
# - $\zeta$: coeficiente de amortiguación.
# Considerando que el sistema es estable, puede observarse que:
# - Este sistema no tiene **ceros**, pues no existe un valor de $s$ que haga que el numerador $k\omega_0^2$ sea igual a cero.
# - El sistema tiene dos **polos** ubicados en
#
# $$s_1,s_2 =-\zeta \omega_0 \pm \omega_0\sqrt{\zeta^2 - 1}$$
#
# Así, deben considerarse 3 casos:
# 1. $\zeta < 1$, por lo cual $\sqrt{\zeta^2 - 1}$ arroja un parte imaginaria y el sistema tiene dos polos complejos conjugados. En este caso, el sistema es **subamortiguado**.
#
# 2. $\zeta = 1$, por lo cual $\sqrt{\zeta^2 - 1} = 0$ y el sistema tendría dos polos reales repetidos. En este caso, el sistema es **críticamente amortiguado**.
#
# 3. $\zeta > 1$, por lo cual $\sqrt{\zeta^2 - 1}$ arroja un parte real y el sistema tiene dos polos reales diferentes. En este caso, el sistema es **sobreamortiguado**.
# ### Caso subamortiguado
#
# La función de transferencia del sistema es:
#
# \begin{equation}
# G(s) = \frac{C(s)}{R(s)} = \frac{k\omega_0^2}{s^2 + 2 \zeta\omega_0s + \omega_0^2}
# \end{equation}
#
# En este caso se considera que $\zeta < 1$, por lo cual $\sqrt{\zeta^2 - 1}$ arroja un parte imaginaria y el sistema tiene dos polos complejos conjugados.
#
# #### Respuesta al escalón
#
# Si el sistema es excitado con un escalón unitario, la señal de salida en el dominio de la frecuencia es:
#
# \begin{equation}
# C(s) = G(s) \cdot R(s) = \frac{k\omega_0^2}{s^2 + 2 \zeta\omega_0s + \omega_0^2} \cdot \frac{1}{s}
# \end{equation}
#
# Observe que esta expresión puede reescrbirse como:
#
# \begin{equation}
# C(s)= k \cdot \left ( \frac{1}{s} - \frac{s + 2\zeta\omega_0}{s^2 + 2\zeta\omega_0s + \omega_0^2} \right )
# \end{equation}
#
# Suponiendo que la parte imaginaria de los polos es $\omega_d = \omega_0\sqrt{1-\zeta^2}$, la respuesta se puede reescribir como:
#
# \begin{equation}
# C(s)= k \cdot \left ( \frac{1}{s} - \frac{s + \zeta\omega_0}{(s+\zeta\omega_0)^2 + \omega_d^2} - \frac{\zeta\omega_0}{(s+\zeta\omega_0)^2 + \omega_d^2} \right )
# \end{equation}
#
# La transformada inversa de Laplace permite obtener una expresión para la señal de salida en el dominio del tiempo $c(t)$.
#
# \begin{align}
# c(t) &= k \left ( 1 - e^{-\zeta\omega_0 t} \cos{\omega_d t} - \frac{\zeta\omega_0}{\omega_d}e^{-\zeta\omega_0 t} \sin{\omega_d t}\right ) \epsilon(t) \\
# &= k \left ( 1 - e^{-\zeta\omega_0 t} \cos{\omega_d t} - \frac{\zeta}{\sqrt{1-\zeta^2}}e^{-\zeta\omega_0 t} \sin{\omega_d t}\right ) \epsilon(t) \\
# &= k \left ( 1 - \frac{e^{-\zeta\omega_0 t}}{\sqrt{1-\zeta^2}} \sin \left (\omega_d t + \tan^{-1} \frac{\sqrt{1-\zeta^2}}{\zeta} \right )\right ) \epsilon(t) \\
# &= k \left ( 1 - \frac{e^{-\zeta\omega_0 t}}{\sqrt{1-\zeta^2}} \sin \left (\omega_0 \sqrt{1-\zeta^2} t + \tan^{-1} \frac{\sqrt{1-\zeta^2}}{\zeta} \right )\right ) \epsilon(t)
# \end{align}
#
# Observe que:
#
# - $c(t<0)=0$ (condción inicial).
# - $\lim_{t\rightarrow \infty} c(t)=k$ (respuesta estacionaria).
# - La respuesta transitoria es dominada por $e^{-\zeta\omega_0 t}$ siendo modulada por una oscilación cuya frecuencia es $\omega_d = \omega_0 \sqrt{1-\zeta^2}$.
# - Si $\zeta = 0 $, el sistema oscilaría eternamente.
# - El sistema tiene dos polos ubicado en $s_1,s_2 =-\zeta \omega_0 \pm j \omega_d$
# +
k, omega0, zeta, omegad = sym.symbols('k, omega_0, zeta, omega_d',real=True, positive=True)
t = sym.symbols('t',real=True, positive=True)
wd = {omegad : omega0*sym.sqrt(1-zeta**2)}
s = sym.Symbol('s')
G2_sub = k*omega0**2/((s + zeta*omega0 + omegad*1j)*(s + zeta*omega0 - omegad*1j))
G2_sub # La función de transferencia
# -
G2_sub_step = G2_sub*(1/s) # La respuesta ante el escalón
G2_sub_step = G2_sub_step.apart(s) # Descompone larespuesta al escalón en fracciones parciales
G2_sub_step
g2_sub_step = invL(G2_sub_step).subs(wd).factor().simplify()
g2_sub_step
parametros = {k: -5,omega0: 8, zeta: 0.25}
sym.plot(g2_sub_step.subs(parametros),(t,0,10))
# ### Caso críticamente amortiguado
#
# En este caso se considera que $\zeta = 1$, por lo cual $\sqrt{\zeta^2 - 1}$ arroja un parte imaginaria igual a cero polos reales repetidos. La función de transferencia del sistema queda:
#
# \begin{equation}
# G(s) = \frac{C(s)}{R(s)} = \frac{k\omega_0^2}{s^2 + 2 \omega_0s + \omega_0^2} = \frac{k\omega_0^2}{\left (s + \omega_0 \right )^2}
# \end{equation}
#
#
# #### Respuesta al escalón
#
# Si el sistema es excitado con un escalón unitario, la señal de salida en el dominio de la frecuencia es:
#
# \begin{equation}
# C(s) = G(s) \cdot R(s) = \frac{k\omega_0^2}{\left (s + \omega_0 \right )^2} \cdot \frac{1}{s}
# \end{equation}
#
# Observe que esta expresión puede reescrbirse como:
#
# \begin{equation}
# C(s)= k \cdot \left ( \frac{1}{s} - \frac{\omega_0}{\left ( s + \omega_0 \right )^2} - \frac{k}{s + \omega_0} \right )
# \end{equation}
#
# Aplicando la transformada inversa se obtiene:
#
# \begin{align}
# c(t) &= k \left ( 1 - \omega_0 t e^{-\omega_0 t} - e^{-\omega_0 t} \right ) \epsilon(t) \\
# &= k \left ( 1 - (1 + \omega_0 t )e^{-\omega_0 t} \right ) \epsilon(t)
# \end{align}
#
#
# - $c(t<0)=0$ (condción inicial).
# - $\lim_{t\rightarrow \infty} c(t)=k$ (respuesta estacionaria).
# - La respuesta transitoria es dominada por $(1 + \omega_0 t )e^{-\omega_0 t}$.
# - El sistema tiene dos polos ubicados en $s_1,s_2 =-\omega_0$
#
# +
k, omega0 = sym.symbols('k, omega_0',real=True, positive=True)
t = sym.symbols('t',real=True, positive=True)
s = sym.Symbol('s')
G2_cri = k*omega0**2/(s + omega0)**2
G2_cri # La función de transferencia
# -
G2_cri_step = G2_cri*(1/s) # La respuesta ante el escalón
G2_cri_step = G2_cri_step.apart(s) # Descompone larespuesta al escalón en fracciones parciales
G2_cri_step
g2_cri_step = invL(G2_cri_step).factor().simplify()
g2_cri_step
parametros = {k: 5,omega0: 6}
sym.plot(g2_cri_step.subs(parametros),(t,0,2))
# ### Caso sobreamortiguado
#
# En este caso se considera que $\zeta > 1$, por lo cual $\sqrt{\zeta^2 - 1}$ arroja un parte imaginaria igual a cero y polos reales diferentes en $s_1,s_2 = \omega_0 (-\zeta \pm \sqrt{\zeta^2 - 1})$. La función de transferencia del sistema queda:
#
# \begin{equation}
# G(s) = \frac{C(s)}{R(s)} = \frac{k\omega_0^2}{s^2 + 2 \zeta \omega_0s + \omega_0^2} = \frac{k s_1 s_2}{\left (-s + s_1 \right ) \left (-s + s_2 \right )}
# \end{equation}
#
#
# #### Respuesta al escalón
#
# Si el sistema es excitado con un escalón unitario, la señal de salida en el dominio de la frecuencia es:
#
# \begin{equation}
# C(s) = G(s) \cdot R(s) = \frac{k s_1 s_2}{\left (-s + s_1 \right ) \left (-s + s_2 \right )} \cdot \frac{1}{s}
# \end{equation}
#
# Observe que esta expresión puede reescrbirse como:
#
# \begin{equation}
# C(s)= k \cdot \left ( \frac{1}{s} - \frac{s_1}{s_1 - s_2}\frac{1}{s+s_2} + \frac{s_2}{s_1 - s_2}\frac{1}{s+s_1} \right )
# \end{equation}
#
# \begin{equation}
# C(s)= k \cdot \left ( \frac{1}{s} + \frac{1}{s_1 - s_2} \left ( \frac{s_1}{s+s_2} + \frac{s_2}{s+s_1} \right ) \right )
# \end{equation}
#
# Aplicando la transformada inversa se obtiene:
#
# \begin{equation}
# c(t)= k \cdot \left ( 1 + \frac{1}{s_1 - s_2} \left (s_1 e^{-s_2 t} + s_2 e^{-s_1 t} \right ) \right )
# \end{equation}
#
#
# - $c(t<0)=0$ (condción inicial).
# - $\lim_{t\rightarrow \infty} c(t)=k$ (respuesta estacionaria).
# - La respuesta transitoria es dominada por dos exponenciales reales.
# - El sistema tiene dos polos reales ubicados en $s_1,s_2 = \omega_0 (-\zeta \pm \sqrt{\zeta^2 - 1})$
#
# +
k, s1,s2,zeta,omega0 = sym.symbols('k, s_1, s_2,\zeta,\omega_0',real=True, positive=True)
t = sym.symbols('t',real=True, positive=True)
s = sym.Symbol('s')
G2_sob = k*s1*s2/((-s + s1)*(-s+s2))
G2_sob # La función de transferencia
# -
G2_sob_step = G2_sob*(1/s) # La respuesta ante el escalón
G2_sob_step = G2_sob_step.apart(s) # Descompone larespuesta al escalón en fracciones parciales
G2_sob_step
g2_sob_step = invL(G2_sob_step).factor().simplify()
g2_sob_step
s1s2 = {s1: omega0*(-zeta + sym.sqrt(zeta**2-1)) ,s2: omega0*(-zeta - sym.sqrt(zeta**2-1))}
# \omega_0 (-\zeta \pm \sqrt{\zeta^2 - 1})
g2_sob_step = g2_sob_step.subs(s1s2).simplify()
g2_sob_step
# +
parametros1 = {k: 1.0,omega0: 2.0, zeta: 1.1}
parametros2 = {k: 1.0,omega0: 2.0, zeta: 2.5}
parametros3 = {k: 1.0,omega0: 2.0, zeta: 5}
g1 = sym.plot(g2_sob_step.subs(parametros1),
(t,0,10),show=False,line_color='green')
g2 = sym.plot(g2_sob_step.subs(parametros2),
(t,0,10),show=False, line_color='blue')
g3 = sym.plot(g2_sob_step.subs(parametros3),
(t,0,10),show=False, line_color='red')
g1.extend(g2)
g1.extend(g3)
g1.show()
# -
# **Pregunta**
# ¿Qué esperaría que ocurra si el coeficiente de amoriguamiento de un sistema de segundo orden es "muy grande"?
| RespuestaDinamica.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sympy import *
init_printing(use_unicode=True)
# +
# A Matrix object is constructed by providing a list
# of row vectors that make up the matrix
# -
A = Matrix([1,2,3])
print(A)
# For pretty printing
A
AA = Matrix([[1,2],[3,4]])
AA
print(AA)
## Matrix transpose
B = AA.T
B
# Matrix multiplication
M = Matrix([[1,2,3],[4,5,6]])
M
N = M.T
N
MN = M*N
MN
# shape of the metrix. Number of rows and columns.
M.shape
MN.shape
# accessing entries
M = Matrix([[1,2,3],[4,5,6]])
M
M.row(0)
M.col(1)
# last column
M.col(-1)
M[:,1]
M[0,:]
# insert row/col after a particular row/col
M = M.row_insert(1,Matrix([[0,4,9]]))
M
N = 2*M
N
N**2
N**-1
NNinv = N.inv()
NNinv
# determinant of a matrix
Ndet = N.det()
Ndet
NNinv.det()
# matrix constructors
I = eye(4)
I
Z = zeros(3,4)
Z
ones(2,3)
A = Matrix([[1,-1,0],[-1,2,-1],[0,-1,1]])
A
A.eigenvals()
A.eigenvects()
# diagonalisation A = P*D*Pinv
P, D = A.diagonalize()
D
P
P*D*P**-1 == A
# characteristic polynomials
lamda = symbols('lamda')
p = A.charpoly(lamda)
p
factor(p)
| 5-matrices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Linear Regression as a model of Emotional Appraisal
#
# The first example we discuss involves building a simple linear regression model in Pyro. We consider a case that is relevant to affective computing --- how should we build a model to reason about someone's emotions? There are lots of emotion theories that one can draw from, and probabilistic programming offers an elegant way to specify (and test!) these theories.
#
#
# Before we jump in, we want to introduce the dataset that we'll be working with throughout the rest of the tutorial. This dataset is of a managable size (so training shouldn't take too long), but still complex enough that we can demonstrate some of the nice features of applying probabilistic programming to model real-life, AI-relevant examples (in this case, relevant to affective computing).
# ### Dataset
#
# We will be using data from a published paper (Ong, Zaki, & Goodman, 2015; Experiment 3), which is available [here](https://github.com/desmond-ong/affCog) but we have also reproduced the data in the current repository. In this experiment, we showed human participants an agent playing a gamble; he spins a wheel with three possible outcomes, and wins the amount on the wheel.
#
# <div style="width: 500px; margin: auto;"></div>
# (Figure modified from Ong et al, 2015, Figure 9).
#
# On some trials, participants see the outcome that the agent won ((i) above). On other trials ((ii) above), participants were not shown the outcome, but instead were shown what ostensibly was the agent's facial expression after seeing the outcome. And on the last third of trials ((iii) above), participants were shown both the outcome and the agent's facial expression.
# Following these, participants were asked to rate how they thought the agent felt, on 8 emotions, using a 9 point Likert scale.
#
# Thus, the dataset consists of some "outcome only" trials where participants saw outcomes and rated the agent's emotions, "facial expression only" trials where participants attributed emotions to a facial expression, and trials where they saw both and had to integrate the information from both the outcome and the facial expression to make a judgment.
# ### Appraising the outcome
#
# Let us first consider the "outcome only" trials. Many established emotion theories and affective computing theories hold that people experiencing events (e.g., winning the lottery, missing the bus) will evaluate the situation according to a set of important features. Was the outcome desirable? Was the outcome surprising? Was the outcome controllable? This evaluation is known as **appraisal**.
#
# Put another way, the number of situations that people encounter in daily life vary immensely along a large number of dimensions, some important (the amount that one wins in the lottery) and some not so important (the color of the lottery ticket). **Appraisal** is computationally necessary to reduce the complexities of everyday situations into a low dimensional set of emotion-relevant dimensions.
#
# ### A Linear Regression model with Appraisal
#
# Thus, we already have the few basic ingreidents of our theory. We have an observable variable (the outcome). We have an appraisal process that converts the outcome into a small number of relevant features. And we have the emotion ratings that people produce. Let's construct a basic regression model:
#
# <div style="width: 300px; margin: auto;"></div>
# Here's how to read the model above, which uses graphical model notation. Shaded circles represent observed variables, while unshaded represents latent, or unobserved variables. Small rectangles represent parameters (to be fitted). We have $N$ i.i.d. pairs of (*Outcome*, *Emotion Ratings*), where $N$ is the size of the dataset, and so these are represented in the large rectangle (called a "plate"), to indicate that they are repeated $N$ times. And between these *Outcome* and *Rating* pairs, we have an appraisal transformation.
#
# In a linear regression, we have $K+1$ regression weights that map the appraisal to the emotion ratings (the $+1$ is for the bias term). If the appraisal variables are given by $\{1, a_1, a_2, \ldots, a_K\} = \vec{a}$, we can write a regression equation:
#
# *Rating* = $\vec{\beta} \cdot \vec{a}$ = $\beta_0$ + $\beta_1 a_1$ + $\beta_2 a_2$ + $\ldots$ + $\beta_K a_K$ + $\epsilon$
#
# where the $\beta_i$'s represent the regression weights for the $i$-th appraisal variable, $\beta_0$ is the bias term and $\epsilon$ is an error term. Notice that the $\beta_i$'s should remain the same across all $N$ observations: thus, it is left out of the "plate" in the model diagram above.
#
# We assume that each $\beta_i$ is drawn from a Normal distribution parameterized by a mean (location parameter) $\mu_i$ and a standard deviation (or scale parameter) $\sigma_i$, i.e., $\beta_i \sim N(\mu_i, \sigma_i)$. We wish to learn these parameters $\mu_i, \sigma_i$ from the data.
#
#
# (Note: *Appraisal* in this model is a strange creature. We could represent it as a latent variable (so like *outcome*, but unshaded). Here, to reflect the fact that *appraisal* is a modular function that can be tested scientifically against data, we chose to represent it more like a fittable parameter.).
# Let's write some Pyro!
# ##### Preamble
#
# This first chunk of code imports the necessary python packages and functions that we will use
# +
from __future__ import division, print_function, absolute_import
# %matplotlib inline
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import pyro
import pyro.distributions as dist
from pyro.distributions import Normal
from pyro.infer import SVI, Trace_ELBO
from pyro.optim import Adam
# -
# This next chunk defines some variable and names that are specific to this dataset, as well as a function to read in the data.
#
# The data is stored in `outcome_emotion_dataset`, which is a torch Tensor of size (1541, 17), indicating that there are N=1,541 observations of 17 variables. The first 9 are the parameterization of the outcome (the 3 payoffs on the wheel and their probabilities, which outcome they won and that probability, and the angle within the sector that the wheel landed on), and the next 8 are the emotion variables. All the variables are scaled so that they lie within [0,1].
# +
# data location
dataset_path = os.path.join(os.path.abspath('..'), "CognitionData", "data_wheelOnly.csv")
OUTCOME_VAR_NAMES = ["payoff1", "payoff2", "payoff3", "prob1", "prob2", "prob3", "win", "winProb", "angleProp"]
EMOTION_VAR_NAMES = ["happy", "sad", "anger", "surprise", "disgust", "fear", "content", "disapp"]
OUTCOME_VAR_DIM = len(OUTCOME_VAR_NAMES)
EMOTION_VAR_DIM = len(EMOTION_VAR_NAMES)
def load_outcome_emotion_dataset(csv_file, normalize_values=True, preview_datafile=False):
data_readin = pd.read_csv(csv_file)
outcome_data = data_readin.loc[:,OUTCOME_VAR_NAMES]
if normalize_values:
####
## payoff1, payoff2, payoff3 and win are between 0 and 100
## need to normalize to [0,1] to match the rest of the variables,
## by dividing payoff1, payoff2, payoff3 and win by 100
####
outcome_data.loc[:,"payoff1"] = outcome_data.loc[:,"payoff1"]/100
outcome_data.loc[:,"payoff2"] = outcome_data.loc[:,"payoff2"]/100
outcome_data.loc[:,"payoff3"] = outcome_data.loc[:,"payoff3"]/100
outcome_data.loc[:,"win"] = outcome_data.loc[:,"win"]/100
outcome_data_tensor = torch.tensor(outcome_data.values).type(torch.Tensor)
# the actual data has 8 emotions, but for illustration we just use 1 emotion, happy
# the rest of the functions below assume a 1-D "y" variable
emotion_data = data_readin.loc[:,EMOTION_VAR_NAMES]
#emotion_data = data_readin.loc[:, "happy"]
if normalize_values:
## note that emotions are transformed from a 9 point Likert to [0,1] via emo <- (emo-1)/8
emotion_data = (emotion_data-1)/8
#emotion_data = emotion_data.values.reshape( emotion_data.shape[0] , 1)
#emotion_data = torch.tensor(emotion_data).type(torch.Tensor)
emotion_data_tensor = torch.tensor(emotion_data.values).type(torch.Tensor)
if preview_datafile:
print("Preview of first 3 rows:")
print(outcome_data.loc[0:2,:])
print(emotion_data.loc[0:2,:])
data = torch.cat((outcome_data_tensor, emotion_data_tensor), 1)
return data
# reads in datafile.
print("Reading in dataset...")
outcome_emotion_dataset = load_outcome_emotion_dataset(csv_file=dataset_path, preview_datafile=True)
N_samples = outcome_emotion_dataset.shape[0]
print("Shape of dataset: ", outcome_emotion_dataset.shape)
# -
# `compute_appraisal()` is a function that takes in an outcome vector, and returns a vector of appraisal values. The example below reproduces the appraisal function used in Ong et al (2015). But more generally, this is a modular function that can be substituted out to test other possible operationalizations of appraisal theories.
def compute_appraisal(outcome_data):
# We have a simple hard-coded these appraisals, for illustration
# This is following Ong, Zaki, & Goodman (2015)
# the outcome data columns are, in order:
# ["payoff1", "payoff2", "payoff3", "prob1", "prob2", "prob3", "win", "winProb", "angleProp"]
# the 3 appraisal variables are:
# amount won ("win"),
# Prediction Error PE = win - EV, where EV = prob1*payoff1 + prob2*payoff2 + prob3*payoff3
# absolute value of PE
# if outcome_data only has 1 observation, reshape so vectorization works
if(len(outcome_data.shape)==1):
outcome_data = outcome_data.view(1,9)
print(outcome_data.shape)
# initializing appraisalVals
appraisalVals = torch.zeros(size=(outcome_data.shape[0],3))
appraisalVals[:,0] = outcome_data[:,6] # amount won
# Expected value
EV = outcome_data[:,0] * outcome_data[:,3] + \
outcome_data[:,1] * outcome_data[:,4] + \
outcome_data[:,2] * outcome_data[:,5]
# prediction error and absolute PE
appraisalVals[:,1] = appraisalVals[:,0] - EV
appraisalVals[:,2] = abs(appraisalVals[:,1])
return(appraisalVals)
# ##### Model
#
# Next, we have the model. Let's break down what goes on.
#
# First, the model samples some $\beta$ coefficients from a Normal with some priors over the mean and the scale (in this case, mean of 0 and scale of 1).
# This is achieved using the `pyro.sample()` function.
# For example,
#
# `b_0 = pyro.sample("b_0", Normal(coeff_mean_prior, coeff_scale_prior))`
#
#
# Note that the `sample()` function takes in a variable name, which allows Pyro to uniquely identify that variable in its variable store. (As such, the variable names are unique, and you can only have one `sample()` function with a particular variable name in this function).
#
#
#
# Next, the function will loop over the observed data, using `pyro.iarange()`. This function defines a special Pyro environment with a unique name (`"map"`), within which Pyro understands that each iteration of the "loop" is conditionally independent. Thus, the computation on each data-point is conditionally independent from the computation on other data-points. (This reflects the plate-notation in the model above; each datapoint is independent, BUT the $\beta$ coefficients are the same across all of them, that's why they were defined before the `pyro.iarange()` loop)
#
# Within this loop, we take the `outcome_data`, run it through `compute_appraisal()` to get a small 3-dimensional `appraisal_vars`. We manually compute the regression equation:
#
# `prediction = b_0 + b_1 * appraisal_vars[:,0] + b_2 * appraisal_vars[:,1] + b_3 * appraisal_vars[:,2]`
#
# Thus, `prediction` is the mean of the Normal distribution that the linear regression model predicts.
# Finally, we condition on the observed data:
#
# `pyro.sample("obs", Normal(prediction, 1), obs = emotion_data)`
#
# Notice we use `pyro.sample()` again. We draw a sample from a Normal with mean `prediction` and scale 1, but this time, we condition that this sample is equal to the observed `emotion_data`, using the argument `obs = ...`.
#
#
# And that's basically it for this function. Pyro's `irange()` and `iarange()` [functions](http://pyro.ai/examples/svi_part_ii.html#iarange) allow a flexible way to perform computations on individual datapoints while taking care of conditional independencies (the `i` in `irange()` and `iarange()`). One difference is that `iarange()` is vectorized, so we can perform the calculations on the entire data Tensor instead of individual observations.
#
#
def fit_regression_model(data):
# define the parameters that control the gaussian prior over the regression coeffs.
# mean = 0, scale = 1
coeff_mean_prior = torch.tensor(0.0)
coeff_scale_prior = torch.tensor(1.0)
# sample b_0 (intercept) and b_1 to b_3 (regression coeffs)
b_0 = pyro.sample("b_0", Normal(coeff_mean_prior, coeff_scale_prior))
b_1 = pyro.sample("b_1", Normal(coeff_mean_prior, coeff_scale_prior))
b_2 = pyro.sample("b_2", Normal(coeff_mean_prior, coeff_scale_prior))
b_3 = pyro.sample("b_3", Normal(coeff_mean_prior, coeff_scale_prior))
# loop over observed data
with pyro.iarange("map", data.shape[0]):
outcome_data = data[:, :(OUTCOME_VAR_DIM)]
# Here, for simplification, we are only taking one emotion variable (happy)
# instead of all 8 emotions
emotion_data = data[:, OUTCOME_VAR_DIM]
appraisal_vars = compute_appraisal(outcome_data)
# run the regression forward
prediction = b_0 + b_1 * appraisal_vars[:,0] + b_2 * appraisal_vars[:,1] + b_3 * appraisal_vars[:,2]
# condition on the observed data
pyro.sample("obs", Normal(prediction, 1), obs = emotion_data)
# ##### Guide
#
# We will use stochastic variational inference to do inference (e.g., see [here](http://pyro.ai/examples/svi_part_i.html) and [here](http://pyro.ai/examples/svi_part_ii.html), or next example for more description). SVI involves using a parameterized distribution to help approximate the posterior distribution that we want to infer. Practically this means defining a function that runs 'parallel' to the `model()` that helps to guide the `model()` in making better sampling choices. Hence, this function is sometimes called the `guide()`.
#
# Specifically, we define the `guide()` such that:
#
# - it contains variational parameters via `pyro.param()`. In this case, these parameters are the $\mu_i, \sigma_i$'s that parameterize the regression parameters $b_i \sim N(\mu_i, \sigma_i)$.
# - it has the same unconditional `pyro.sample()` call signature as `model()`. In this case, `pyro.sample("b_0",...)` through `"b_4"`
# - it does not have any conditioned `pyro.sample()` calls.
def fit_regression_guide(data):
mean_b0_param = pyro.param("guide_mean_b0", torch.tensor(0.0))
scale_b0_param = pyro.param("guide_scale_b0", torch.tensor(1.0))
mean_b1_param = pyro.param("guide_mean_b1", torch.tensor(0.0))
scale_b1_param = pyro.param("guide_scale_b1", torch.tensor(1.0))
mean_b2_param = pyro.param("guide_mean_b2", torch.tensor(0.0))
scale_b2_param = pyro.param("guide_scale_b2", torch.tensor(1.0))
mean_b3_param = pyro.param("guide_mean_b3", torch.tensor(0.0))
scale_b3_param = pyro.param("guide_scale_b3", torch.tensor(1.0))
# sample coefficients from Normal(mean, scale)
pyro.sample("b_0", Normal(mean_b0_param, scale_b0_param))
pyro.sample("b_1", Normal(mean_b1_param, scale_b1_param))
pyro.sample("b_2", Normal(mean_b2_param, scale_b2_param))
pyro.sample("b_3", Normal(mean_b3_param, scale_b3_param))
# ##### Fitting the model
#
# Next, we can proceed to actually fit the model. The first step is to refresh the parameter store using `pyro.clear_param_store()`.
#
# We will use Stochastic Variational Inference `SVI()` which takes in the model and guide that we wrote above, as well as an optimization algorithm (here we use `torch.optim.Adam()`) and a loss function (here we use `Trace_ELBO()`). When `svi.step(data)` is called, it runs SVI over the `data`. Thus, here we simply define a loop that runs over the entire dataset `num_iterations` times. (We can easily modify this to do mini-batching, for example, for large datasets.)
# +
pyro.clear_param_store()
num_iterations = 1000
# setup the optimizer with some learning rate
optimizer = Adam({"lr": 0.005})
# setup the inference algorithm
svi = SVI(fit_regression_model, fit_regression_guide, optimizer, loss=Trace_ELBO())
# do gradient steps
losses = []
for thisIteration in range(num_iterations):
# calculate the loss and take a gradient step
thisLoss = svi.step(outcome_emotion_dataset)
losses.append(thisLoss)
if thisIteration % 100 == 0:
print("[iteration %04d] loss: %.4f" % (thisIteration + 1, thisLoss / float(N_samples)))
plt.plot(losses)
plt.title("ELBO")
plt.xlabel("step")
plt.ylabel("loss")
# -
# output the learned variational parameters
print("b0 ~ Normal(%.4f, %.4f)" % (pyro.param("guide_mean_b0").item(), pyro.param("guide_scale_b0").item()))
print("b1 ~ Normal(%.4f, %.4f)" % (pyro.param("guide_mean_b1").item(), pyro.param("guide_scale_b1").item()))
print("b2 ~ Normal(%.4f, %.4f)" % (pyro.param("guide_mean_b2").item(), pyro.param("guide_scale_b2").item()))
print("b3 ~ Normal(%.4f, %.4f)" % (pyro.param("guide_mean_b3").item(), pyro.param("guide_scale_b3").item()))
# ### Coda
#
# Hopefully the example above illustrates several appealing features of probabilistic programming:
#
# - We can use PPLs to specify theory, and importantly, uncertainty in theory and random processes.
# - Theory is represented as modular chunks of code. (As in our `compute_appraisal()` function, or in the linear regression model.) This allows us to substitute out different parts of the model, which is handy for optimization or theory testing!
# - Want to try a different appraisal representation? No problem!
# - Want to try using a feed-forward neural network instead of a linear regression? Sure (and in fact we will cover that in the next example!)
# - Inference and learning is orthogonal to model specification. Thus, the modeler can focus on specifying the model, while Pyro does most of the heavy lifting (by leveraging PyTorch modules).
#
#
# In the next example, we show how to incorporate a feed-forward neural network in order to build a deep generative model.
#
# ------
# ##### Extra: Some further extensions to the linear regression model
#
# In the following code we modify the earlier model to handle an arbitrary number of regression parameters, and introduce the notion of "lifting" models to get distributions over *models* that we can sample from.
#
# Instead of using a low-dimensional `compute_appraisal()` function, we demonstrate a standard linear regression by defining an `appraisalRegressionModule()` that inherits from PyTorch's `nn.module`. (Note that a linear regression can be thought of as a feed-forward neural network with 0 hidden layers and no non-linearities, such that the output layer is just a linear combination of the input units). This is achieved using `nn.Linear()`:
# +
class appraisalRegressionModule(nn.Module):
def __init__(self, num_features):
super(appraisalRegressionModule, self).__init__()
self.linear = nn.Linear(num_features, 1)
def forward(self, outcome):
return self.linear(outcome)
regression_model = appraisalRegressionModule(OUTCOME_VAR_DIM)
# -
# Next, we write the model. The main difference now is that we can define separate location and scale priors for each regressor, given by `weights_loc` and `weights_scale` below, which are each Tensors of Size(1,`OUTCOME_VAR_DIM`) (or any arbitrary length given by an appraisal transformation).
#
# Rather than typing out `OUTCOME_VAR_DIM` `pyro.sample()` statements, a fancier way of sampling using `weights_loc, weights_scale` is to "lift" the regression model using `pyro.random_module()`. This takes a model and a prior, and returns a *distribution over possible models*. In other words, we can think of this step as sampling models from a "model-distribution" (which under the hood, Pyro does by sampling the regression weights from their priors). Note that we do not see any `pyro.sample()` calls in the code below, as compared to the earlier code: these are taken care of by Pyro and will be executed when sampling from the distribution returned by `random_module()`.
#
# The rest of the code is the same as above.
def bayesianRegressionModel(data):
# Create unit normal priors over the parameters
weights_loc = torch.zeros(size=(torch.Size((1, OUTCOME_VAR_DIM))))
weights_scale = torch.ones(size=(torch.Size((1, OUTCOME_VAR_DIM))))
weights_prior = Normal(weights_loc, weights_scale).independent(1)
# location and scale prior for the bias
bias_loc = torch.zeros(size=(torch.Size((1, ))))
bias_scale = torch.ones(size=(torch.Size((1, ))))
bias_prior = Normal(bias_loc, bias_scale).independent(1)
priors = {'linear.weight': weights_prior, 'linear.bias': bias_prior}
# lift module parameters to random variables sampled from the priors
lifted_module = pyro.random_module("module", regression_model, priors)
# sample a model (which also samples from weights_prior and bias_prior)
sampled_regression_model = lifted_module()
with pyro.iarange("map", data.shape[0]):
outcome_data = data[:, :(OUTCOME_VAR_DIM)]
# Here, for simplification, we are only taking one emotion variable (happy)
# instead of all 8 emotions
emotion_data = data[:, OUTCOME_VAR_DIM]
# run the regressor forward conditioned on data
prediction = sampled_regression_model(outcome_data).squeeze(-1)
# condition on the observed data
pyro.sample("obs", Normal(prediction, 1), obs = emotion_data)
# We write the guide function in a parallel manner to the model function. Note that we register the variational parameters in Pyro's parameter store using the `pyro.param()` function, just like above. We do the same trick of lifting the regression model using a `pyro.random_module()` call and sample from it.
def bayesianRegressionGuide(data):
# define our variational parameters
weights_loc = torch.randn(1, OUTCOME_VAR_DIM)
# Note that the scale has to be non-negative. Thus, we use exp() to get a non-negative number.
# we also use a narrower scale (exp(-1) ~ 0.35 instead of exp(0) = 1)
weights_scale = torch.exp(-1.0 * torch.ones(1, OUTCOME_VAR_DIM) + 0.05 * torch.randn(1, OUTCOME_VAR_DIM))
bias_loc = torch.randn(1)
bias_scale = torch.exp(-1.0 * torch.ones(1) + 0.05 * torch.randn(1))
# using pyro.param() to register the variational parameters
weight_loc_param = pyro.param("guide_loc_weight", weights_loc)
weight_scale_param = pyro.param("guide_scale_weight", weights_scale)
bias_loc_param = pyro.param("guide_loc_bias", bias_loc)
bias_scale_param = pyro.param("guide_scale_bias", bias_scale)
# guide distributions for w and b
weight_dist = Normal(weight_loc_param, weight_scale_param).independent(1)
bias_dist = Normal(bias_loc_param, bias_scale_param).independent(1)
dists = {'linear.weight': weight_dist, 'linear.bias': bias_dist}
# lift the module and sample from that distribution
lifted_module = pyro.random_module("module", regression_model, dists)
return lifted_module()
# Next we fit this regression model using SVI; this is identical to the code earlier.
# +
pyro.clear_param_store()
num_iterations = 1000
# setup the optimizer with some learning rate
optimizer = Adam({"lr": 0.005})
# setup the inference algorithm
bayesianRegressionSVI = SVI(bayesianRegressionModel, bayesianRegressionGuide, optimizer, loss=Trace_ELBO())
# do gradient steps
losses = []
for thisIteration in range(num_iterations):
# calculate the loss and take a gradient step
thisLoss = bayesianRegressionSVI.step(outcome_emotion_dataset)
losses.append(thisLoss)
if thisIteration % 100 == 0:
print("[iteration %04d] loss: %.4f" % (thisIteration + 1, thisLoss / float(N_samples)))
plt.plot(losses)
plt.title("ELBO")
plt.xlabel("step")
plt.ylabel("loss")
# +
#for name in pyro.get_param_store().get_all_param_names():
# print(name, pyro.param(name).data.numpy())
guide_loc_weight = pyro.param("guide_loc_weight")[0]
guide_scale_weight = pyro.param("guide_scale_weight")[0]
print("b0 ~ Normal(%.4f, %.4f)" % (pyro.param("guide_loc_bias").item(), pyro.param("guide_scale_bias").item()))
for j in xrange(len(guide_loc_weight)):
print("b%1d" % (j+1), "~ Normal(%.4f, %.4f)" % (guide_loc_weight[j], guide_scale_weight[j]))
# -
# -----
#
# Written by: <NAME> (<EMAIL>)
#
# References:
#
# Pyro [Bayesian Regression tutorial](http://pyro.ai/examples/bayesian_regression.html)
#
#
# Data from https://github.com/desmond-ong/affCog, from the following paper:
#
# <NAME>., <NAME>., & <NAME>. (2015). Affective Cognition: Exploring lay theories of emotion. *Cognition*, 143, 141-162.
| code/LinearRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Signals and Systems
#
# __Signal__ : Any physical quantity that changes with time or space or both is a signal
# > Imagine a linear array describing some property(like amplitude) of a quantity(physical quantity such as sound) wrt time or space or both.
#
# Processing Options
#
# - Addition of two or more signals
# - Subtraction of signals
# - Multiplication
# - Division
# - Differentiation
# - Integration
# - Filtering
# ## Transducers
#
# * Comvert one form of signal into another form of signal
# ## __Classification of signals__
#
# 1. Prdicatable Signal
# 2. Unpredicatable Signal
#
#
# _Predicatable signals_ are functions that can be plotted as a function of time.
# for example
# +
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
A=np.linspace(-10,10,1000)
B=np.sin(A)
plt.figure(figsize=(16,4))
plt.plot(B,"r-")
plt.show()
# -
# _Unpredicatable signals_ are functions that cannot be plotted as function as space or time.
# Examples are
#
# 1. Information carrying signal
# 2. Noise signal
# ### Frequency Domain Description
#
# ???
#
# ### Sepctrum of periodic and non-periodic signals
# # Notation
#
# * x(t) denotes a signal
# * x denotes the instantaneous amplitude
# * t denotes time
# * Nature of variables
# Continuous
# Discrete
# ## Analog Signal
#
# - Both x and t continuous variables (Continuous Amplitude, Continuous time CA/CT) : Analog signal
#
#
# ## Digital Signal
#
# - DA/DT
# - Most transducers give analog signal
# Microphone output
# Thermocouple
# * Analog to Digital converters
# Convert analog signal into a digital signal
#
# ___Sampling___ : Converts CA/CT signal into CA/DT signals
#
# How to?
#
# - __Multiplier__ : One input is the signal and the other input is a periodic trains of narrow pulses
#
# 
# _Quantization_ : This converts continuos amplitude to discrete amplitude
#
# _Encoding_ : Converts quantized amplitudes to binary values
#
# About (DA/CT)
#
# About Basic Elements of a DSP System
#
# DSP vs ASP
# ## Discrete Time Signal
# ___Discrete time signals___ are “the signals or quantities that can be defined and represented at certain time instants of the sequence.” These are finite or countable sets of number sequences. They are also called digitalized signals.
a=np.linspace(-5,5,10)
# plt.figure(figsize=(16,8))
plt.stem(a,linefmt="-.",use_line_collection=True,label="DT")
plt.plot(a,"r-",label="CT")
plt.legend()
plt.show()
# plt.hist(a)
# - Continuous Time signal
# x(t) : t is a continuous variable
# Can be described by an equation
# Graph
# - Discrete Time signal
# x(n) : n is an integer
# Can be described by an equation
# By a line plot
# By a sequence of values within a bracket
#
# __Sequencd__ :An infinite sequence
#
# x(n) = { ---- , x(-3), x(-2), x(-1), x(0),x(1),x(2), ----}
#
#
# The arrow denotes the value at n = 0
# i.e.
#
# x(n) = { ----, -2,1.3 , 2 -3.4, -1 , -2, 1, ----}
# ___Finite length signal___ is defined only for a finite time
# interval
#
#
# ___Zero Padding___
#
# * Signal length can be increased by adding zero valued samples outside the range
# - x(n) = { 3, -2, 1, 0, 2, -3, -1, 0, 2, 1 }
# * The length is 10. This can be increased
# - x(n) = { 0,0,3, -2, 1, 0, 2, -3, -1, 0, 2, 1,0,0 } Now N = 14
#
# ### Basic Operations on discrete time signals
# * Signal addition
# - think of it as adding two linear arrays of same size where each array contains data representing data describing amplitude of a signal
# - additing of matrix of size 1xN(where DT has N samples)
# * Scalar Addition
# - think of it as adding a scalar to an array of some size where array contains data representing data describing amplitude of a signal
#
# * Signal Mulitplication
# - - think of it as multiplying two linear arrays of same size where each array contains data representing data describing amplitude of a signal
#
# * Scalar Multiplication
# - think of it as multiplying a scalar to an array of some size where array contains data representing data describing amplitude of a signal
# +
x=np.linspace(-5,5,20)
y=np.linspace(-4,9,20)
z=x+y
x=np.sin(x)
y=np.cos(y)
z=x+y
plt.figure(figsize=(10,7))
plt.title("Example of Signal Addition")
plt.stem(range(len(x)),x,"r",markerfmt="ro",label="sinx",use_line_collection=True)
plt.stem(range(len(y)),y,"b",markerfmt="bo",label="siny",use_line_collection=True)
plt.stem(range(len(z)),z,"g",markerfmt="go",label="sinx+siny",use_line_collection=True)
plt.plot(x,"r-.",label="x-")
plt.plot(y,"b-.",label="y-")
plt.plot(z,"g--",label="x+y -")
plt.legend()
plt.plot()
# -
# ## Transformation of independent variable
#
# - Time delay
# - Time shifting
# - delay
# - advance
# - Time Scaling
# - Decimation
# - Decimation refers to the process of reducing signal length by discarding signal samples
# - y(n) = x(Mn)
#
# M is an integer.
#
# Every Mth sample of x(n) is kept and discarding
# (M-1) samples in between.
#
# - Interpolation
# - Interpolation refers to the process of increasing the signal length by inserting zeros between signal samples.
# - y(n)=x(n/L) n=0,+-L,+-2L
# - y(n)=0
# - The process consists of inserting L-1 zero valued samples between each two consecutive samples of
# x(n)
#
#
# __Time Reversal__
#
# __Combined Operations__
# ### Some basic Discrete time signals
#
# __Unit Step__
A=np.zeros(11)
A[5:]=1
plt.stem(np.arange(-5,6),A,"c",markerfmt="c.",use_line_collection=True)
# __Unit Impulse__
A=np.zeros(11)
A[5]=1
plt.stem(np.arange(-5,6),A,"c",markerfmt="c.",use_line_collection=True)
# __Unit Ramp signal__
A=np.zeros(30)
A[15:]=np.linspace(0,1.,15)
plt.stem(np.linspace(-5,5,30),A,"c",markerfmt="co",use_line_collection=True)
# #### Discrete time real exponential signals
# Search
#
# ### Discrete time plot for sine wave
a=np.linspace(-6.24,6.24,20)
b=np.sin(a)
plt.stem(np.arange(20),b,"g",markerfmt="go",use_line_collection=True)
| SS1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda]
# language: python
# name: conda-env-anaconda-py
# ---
import pandas as pd
import numpy as np
import networkx as nx
import matplotlib.pyplot as plt
# ## Basic Graph Setup
# +
# Creating a Graph
G = nx.Graph() # Right now G is empty
# Add a node
G.add_node(1)
G.add_nodes_from([2,3]) # You can also add a list of nodes by passing a list argument
# Add edges
G.add_edge(1,2)
e = (2,3)
G.add_edge(*e) # * unpacks the tuple
G.add_edges_from([(1,2), (1,3)]) # Just like nodes we can add edges from a list
# -
G.nodes()
G.edges()
G[1]
G[1][2]
nx.draw(G)
# ## Anaylsis of a dataset
data = pd.read_csv('Graph Theory-airlines.csv')
data.shape
data.dtypes
# Convert values and fix na...
# +
# converting sched_dep_time to 'std' - Scheduled time of departure
data['std'] = data.sched_dep_time.astype(str).str.replace('(\d{2}$)', '') + ':' + data.sched_dep_time.astype(str).str.extract('(\d{2}$)', expand=False) + ':00'
# converting sched_arr_time to 'sta' - Scheduled time of arrival
data['sta'] = data.sched_arr_time.astype(str).str.replace('(\d{2}$)', '') + ':' + data.sched_arr_time.astype(str).str.extract('(\d{2}$)', expand=False) + ':00'
# converting dep_time to 'atd' - Actual time of departure
data['atd'] = data.dep_time.fillna(0).astype(np.int64).astype(str).str.replace('(\d{2}$)', '') + ':' + data.dep_time.fillna(0).astype(np.int64).astype(str).str.extract('(\d{2}$)', expand=False) + ':00'
# converting arr_time to 'ata' - Actual time of arrival
data['ata'] = data.arr_time.fillna(0).astype(np.int64).astype(str).str.replace('(\d{2}$)', '') + ':' + data.arr_time.fillna(0).astype(np.int64).astype(str).str.extract('(\d{2}$)', expand=False) + ':00'
data['date'] = pd.to_datetime(data[['year', 'month', 'day']])
# finally we drop the columns we don't need
data = data.drop(columns = ['year', 'month', 'day'])
# -
# Create graph using origin and dest as nodes. THe other values will be attributes of the edges.
airlines_graph = nx.from_pandas_edgelist(data, source='origin', target='dest', edge_attr=True,)
airlines_graph.nodes()
airlines_graph.edges()
nx.draw_networkx(airlines_graph, with_labels=True) # Quick view of the Graph. As expected we see 3 very busy airports
nx.algorithms.degree_centrality(airlines_graph) # Notice the 3 airports from which all of our 100 rows of data originates
nx.density(airlines_graph) # Average edge density of the Graphs
nx.average_shortest_path_length(airlines_graph) # Average shortest path length for ALL paths in the Graph
nx.average_degree_connectivity(airlines_graph) # For a node of degree k - What is the average of its neighbours' degree?
# Let us find all the paths available
for path in nx.all_simple_paths(airlines_graph, source='JAX', target='DFW'):
print(path)
# Let us find the dijkstra path from JAX to DFW.
# You can read more in-depth on how dijkstra works from this resource - https://courses.csail.mit.edu/6.006/fall11/lectures/lecture16.pdf
dijpath = nx.dijkstra_path(airlines_graph, source='JAX', target='DFW')
dijpath
# Let us try to find the dijkstra path weighted by airtime (approximate case)
shortpath = nx.dijkstra_path(airlines_graph, source='JAX', target='DFW', weight='air_time')
shortpath
| MachineLearning/Graph Theory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ensembles of classifiers
# In this case, if we want to use a keras NN in our Voting Ensemble, we cannot use the native sklearn function. We need to build the ensemble by hand.
# %load_ext autoreload
# %autoreload 2
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import os
import scipy
import sklearn
# plt.style.use('fivethirtyeight')
sns.set_style("whitegrid")
sns.set_context("notebook")
DATA_PATH = '../data/'
VAL_SPLITS = 4
# +
# Seed value
# Apparently you may use different seed values at each stage
seed_value= 0
# 1. Set the `PYTHONHASHSEED` environment variable at a fixed value
import os
os.environ['PYTHONHASHSEED']=str(seed_value)
# 2. Set the `python` built-in pseudo-random generator at a fixed value
import random
random.seed(seed_value)
# 3. Set the `numpy` pseudo-random generator at a fixed value
import numpy as np
np.random.seed(seed_value)
# 4. Set the `tensorflow` pseudo-random generator at a fixed value
import tensorflow as tf
tf.set_random_seed(seed_value)
# 5. Configure a new global `tensorflow` session
from keras import backend as K
session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
sess = tf.Session(graph=tf.get_default_graph(), config=session_conf)
K.set_session(sess)
# -
from plot_utils import plot_confusion_matrix
from cv_utils import run_cv_f1
from cv_utils import plot_cv_roc
from cv_utils import plot_cv_roc_prc
# +
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import f1_score
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.neural_network import MLPClassifier
# Experimental: Based on LightGMB https://github.com/Microsoft/LightGBM
from sklearn.experimental import enable_hist_gradient_boosting
from sklearn.ensemble import HistGradientBoostingClassifier
# Pipelines
from sklearn.pipeline import Pipeline
import xgboost as xgb
from sklearn.metrics import f1_score, accuracy_score, precision_score
from sklearn_utils import FeatureSelectorDic
# -
# For this part of the project, we will only work with the training set, that we will split again into train and validation to perform the hyperparameter tuning.
#
# We will save the test set for the final part, when we have already tuned our hyperparameters.
# +
df = pd.read_csv(os.path.join(DATA_PATH,'df_train.csv'))
df.drop(columns= df.columns[0:2],inplace=True)
idx_to_feat = dict(enumerate([feat for feat in df.columns if feat is not 'Class']))
feat_to_idx = {feat : idx for idx,feat in idx_to_feat.items()}
cv = StratifiedShuffleSplit(n_splits=VAL_SPLITS,test_size=0.15,random_state=0)
X = df.drop(columns='Class').to_numpy()
y = df['Class'].to_numpy()
df.head()
# -
# ## Ensemble by hand (Hard voting)
def hard_vote_predict(estimators, X, weights=None):
"""
Combine a dictionary of estimators to create a hard voting ensemble.
Parameters
----------
estimators : dict
Dictionary with name (str): model entries with predict method.
If the method predict returns probabilities, then the name should
end with 'prob'.
X : np.array
Input.
weights : list, tuple or np.array, default=None
List of weights for each estimator. If None, then it is uniform.
"""
if weights is None:
weights = np.ones(len(estimators))
else:
assert len(weights) == len(
estimators), 'Number of estimators should be the same as number of weights'
weights = np.array(weights)
weights = weights.reshape((-1, 1))
y_preds = []
for name, clf in estimators.items():
y_pred = clf.predict(X)
if name.endswith('prob'):
y_pred = (1 * (y_pred > 0.5)).reshape((-1))
y_preds.append(y_pred)
y_preds = np.array(y_preds)
y_final = 1 * (np.mean(weights * y_preds, axis=0) > 0.5)
return y_final
# +
from keras.layers import Input, Dense
from keras.models import Model
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.layers import LeakyReLU
def create_clf(input_dim):
clf1 = Sequential([
Dense(8, input_shape=(input_dim,)),
LeakyReLU(),
Dense(4),
LeakyReLU(),
Dense(1, activation='sigmoid')
], name='clf')
return clf1
# +
INPUT_DIM = X.shape[1]
clf1 = create_clf(INPUT_DIM)
clf1.compile(optimizer='adam',
loss='binary_crossentropy')
# clf2 = RandomForestClassifier(n_estimators=100,
# max_depth=6,
# random_state=0,n_jobs=-1, max_features=6)
clf2 = KNeighborsClassifier(n_neighbors=3, n_jobs=-1)
clf3 = xgb.sklearn.XGBClassifier(n_jobs=-1,max_depth=5, random_state=0)
# clf3 = LogisticRegression(n_jobs=-1)
sklearn_clfs = [clf2,clf3]
clfs = [clf1]+sklearn_clfs
# +
metrics = []
accuracy = []
precision = []
metrics_train = []
accuracy_train = []
precision_train = []
for i, (idx_t, idx_v) in enumerate(cv.split(X,y)):
X_train = X[idx_t]
y_train = y[idx_t]
X_val = X[idx_v]
y_val = y[idx_v]
#Devuelve cuatro vectrores de dos elementos, el primero con los indices de train y el segundo con
#los de validacion
clf1.fit(X_train,y_train,batch_size=512,epochs=50,verbose=0)
for clf_ in sklearn_clfs:
clf_.fit(X_train,y_train)
estimators = dict(zip(['nn_prob','rf','knn'],clfs))
y_pred = hard_vote_predict(estimators,X_val)
acc_va = accuracy_score(y_val, y_pred)
pre_va = precision_score(y_val, y_pred)
# error_va = mean_squared_error(y_val, y_pred)
f1_va = f1_score(y_val, y_pred)
#print('Recall:', acc)
#print('Precision:', pre)
#print('Error cuadratico medio:', error)
y_pred_train = hard_vote_predict(estimators,X_train)
acc_train = accuracy_score(y_train, y_pred_train)
pre_train = precision_score(y_train, y_pred_train)
# error_train = mean_squared_error(y_train, y_pred_train)
f1_train = f1_score(y_train, y_pred_train)
metrics.append(f1_va)
accuracy.append(acc_va)
precision.append(pre_va)
metrics_train.append(f1_train)
accuracy_train.append(acc_train)
precision_train.append(pre_train)
print('Fold {} has ended!'.format(i+1))
metric_mean = np.mean(metrics)
metric_std = np.std(metrics, ddof = 1)
print('Metric value validation(va): {:.2f} +- {:.2f}'.format(metric_mean,metric_std))
#print('Mean validation: recall {:.4f} precision {:.4f}'.format(np.mean(accuracy), np.mean(precision)))
metric_train_mean = np.mean(metrics_train)
metric_train_std = np.std(metrics_train, ddof = 1)
print('Metric value train: {:.2f} +- {:.2f}'.format(metric_train_mean,metric_train_std))
# -
| notebooks/8_Ensembles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ZszpXDZ1oTqm"
# Notebook by <NAME>
# + colab={} colab_type="code" id="0G7bnbPmH5A6"
from fastai.vision import *
# + [markdown] colab_type="text" id="YknwvbQjC62F"
# # Dataset:
#
# Our dataset today will be ImageWoof. [Link](https://github.com/fastai/imagenette)
#
# Goal: Using no pre-trained weights, see how well of accuracy we can get in x epochs
#
# This dataset is generally harder than imagenette, both are a subset of ImageNet.
#
# Models are leaning more towards being faster, more effecient
#
# + colab={} colab_type="code" id="zHZANDslHpGq"
def get_data(size, woof, bs, workers=None):
if size<=128: path = "https://s3.amazonaws.com/fast-ai-imageclas/imagewoof2.tgz" if woof else URLs.IMAGENETTE
elif size<=224: path = URLs.IMAGEWOOF_320 if woof else URLs.IMAGENETTE_320
else : path = URLs.IMAGEWOOF if woof else URLs.IMAGENETTE
path = untar_data(path)
n_gpus = num_distrib() or 1
if workers is None: workers = min(8, num_cpus()//n_gpus)
return (ImageList.from_folder(path).split_by_folder(valid='val')
.label_from_folder().transform(([flip_lr(p=0.5)], []), size=size)
.databunch(bs=bs, num_workers=workers)
.presize(size, scale=(0.35,1))
.normalize(imagenet_stats))
# + colab={} colab_type="code" id="yImxYNeJH-AN"
data = get_data(128, True, 64)
# + [markdown] colab_type="text" id="0Xpkh8REIct4"
# We will be following a progression that started on the fastai forums [here](https://forums.fast.ai/t/meet-mish-new-activation-function-possible-successor-to-relu/53299/) on August 26th of this year.
#
# In this "competition" included:
# * [Less](https://forums.fast.ai/u/lessw2020)
# * [Seb](https://forums.fast.ai/u/seb)
# * [<NAME>](https://forums.fast.ai/u/grankin)
# * [<NAME>](https://forums.fast.ai/u/redknight)
# * [<NAME>](https://forums.fast.ai/u/oguiza)
#
# + [markdown] colab_type="text" id="Ehv8AcGK65bK"
# # The Competition:
#
# * Lasted roughly 3 days
# * We explored a variety of papers and combining various ideas to see what *together* could work the best
# + [markdown] colab_type="text" id="KrMPAy167OAr"
# ## Papers Referenced:
#
# * [Bag of Tricks for Resnet (aka the birth of xResNet)](https://arxiv.org/abs/1812.01187)
# * [Large Batch Optimization for Deep Learning, LAMB](https://arxiv.org/abs/1904.00962)
# * [Large Batch Training of Convolutional Networks, LARS](https://arxiv.org/pdf/1708.03888.pdf)
# * [Lookahead Optimizer: k steps forward, 1 step back](https://arxiv.org/abs/1907.08610)
# * [Mish: A Self Regularized Non-Monotonic Neural Activation Function](https://arxiv.org/abs/1908.08681v1)
# * [On the Variance of the Adaptive Learning Rate and Beyond, RAdam](https://arxiv.org/abs/1908.03265)
# * [Self-Attention Generative Adversarial Networks](https://arxiv.org/abs/1805.08318)
# * [Stochastic Gradient Methods with Layer-wise
# Adaptive Moments for Training of Deep Networks, Novograd](https://arxiv.org/pdf/1905.11286.pdf)
#
#
# ## Other Equally as Important Noteables:
# * Flatten + Anneal Scheduling - <NAME>
# * Simple Self Attention - Seb
# + [markdown] colab_type="text" id="55fuRIRsIqxs"
# One trend you will notice throughout this exercise is we (everyone mentioned above and myself) all tried combining a variety of these tools and papers together before Seb eventually came up with the winning solution. For a bit of context, here is the pre-competition State of the Art for ImageWoof:
# 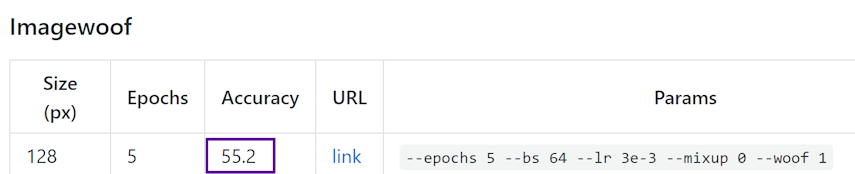
# + [markdown] colab_type="text" id="ciDEdyPUJwf3"
# And here was the winning results:
#
# 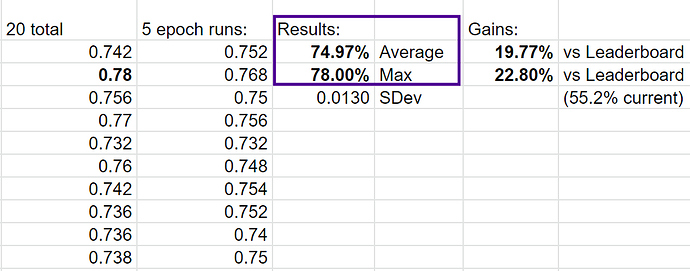
# + [markdown] colab_type="text" id="2s9t11BkKGQ_"
# As a general rule of thumb, we always want to make sure our results are reproducable, hence the multiple runs and reports of the Standard Deviation, Mean, and the Maximum found. For today, we will just do one run of five for time. Following no particular order, here is a list of what was tested, and what we will be testing today:
#
# * Baseline (Adam + xResnet50) + OneCycle
# * Ranger (RAdam + LookAhead) + OneCycle
# * Ranger + Flatten Anneal
# * Ranger + MXResnet (xResnet50 + Mish) + Flatten Anneal
# * RangerLars (Ralamb + LARS + Ranger) + Flatten Anneal
# * RangerLars + xResnet50 + Flatten Anneal
# * Ranger + SimpleSelfAttention + MXResnet + Flatten Anneal
#
# The last of which did achieve the best score overall.
# + [markdown] colab_type="text" id="br_RmMD6QpAT"
# ## Functions:
#
# For the sake of simplicity, we will borrow from Seb's gitub repository.
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="dLwNYYepRAAH" outputId="18da618b-0afa-4264-d7c4-3a353904ef76"
# !git clone https://github.com/sdoria/mish
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="e3ou9mwDRbgk" outputId="37849893-97af-4be3-f96c-033dc5b21dc9"
# %cd mish
from rangerlars import *
from mish import *
from mxresnet import *
from ranger import *
# + [markdown] colab_type="text" id="0l1-KBsiRlbb"
# # Running the tests
#
# For our tests, we will use the overall accuracy as well as the top_k, as this is what was used in Jeremy's example. Do note that top_k is not quite as relevent here as we only have 10 classes
# + [markdown] colab_type="text" id="YoUy5mNWRrDU"
# ## Baseline
# + colab={} colab_type="code" id="vpDu6BIDR_2s"
opt_func = partial(optim.Adam, betas=(0.9,0.99), eps=1e-6)
# + colab={} colab_type="code" id="G4qPOSmFRsGs"
learn = Learner(data, models.xresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
bn_wd=False, true_wd=True, loss_func=LabelSmoothingCrossEntropy(),
metrics=[accuracy, top_k_accuracy])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="3oaAeTgbSVOu" outputId="1cf83d01-4929-4596-bca7-673395c742ed"
learn.fit_one_cycle(5, 3e-3, div_factor=10, pct_start=0.3)
# + [markdown] colab_type="text" id="maPDM39xWSJX"
# ## Ranger + OneCycle
# + colab={} colab_type="code" id="S_JKRL4RWWVI"
opt_func = partial(Ranger, betas=(0.9,0.99), eps=1e-6)
# + colab={} colab_type="code" id="nMJS945OWciJ"
learn = Learner(data, models.xresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
bn_wd=False, true_wd=True, loss_func=LabelSmoothingCrossEntropy(),
metrics=[accuracy, top_k_accuracy])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="v5zn1DygWfIo" outputId="3cfe337e-eda6-42b6-a98f-66d9d29615f2"
learn.fit_one_cycle(5, 3e-3, div_factor=10, pct_start=0.3)
# + [markdown] colab_type="text" id="LG3ETMSVYh0c"
# ## Ranger + Flatten Anneal
# + colab={} colab_type="code" id="d-hQoUBkZBjS"
from fastai.callbacks import *
# + colab={} colab_type="code" id="Da-Nsip8YkyG"
def flattenAnneal(learn:Learner, lr:float, n_epochs:int, start_pct:float):
n = len(learn.data.train_dl)
anneal_start = int(n*n_epochs*start_pct)
anneal_end = int(n*n_epochs) - anneal_start
phases = [TrainingPhase(anneal_start).schedule_hp('lr', lr),
TrainingPhase(anneal_end).schedule_hp('lr', lr, anneal=annealing_cos)]
sched = GeneralScheduler(learn, phases)
learn.callbacks.append(sched)
learn.fit(n_epochs)
# + colab={} colab_type="code" id="rtllGYw1Znca"
opt_func = partial(Ranger, betas=(0.9,0.99), eps=1e-6)
learn = Learner(data, models.xresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
bn_wd=False, true_wd=True, loss_func=LabelSmoothingCrossEntropy(),
metrics=[accuracy, top_k_accuracy])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="GYr2ZXxcZjz7" outputId="6679f3f6-de3e-4f07-dfd4-1b223fb9a550"
flattenAnneal(learn, 3e-3, 5, 0.7)
# + [markdown] colab_type="text" id="LkNMMZTtctc4"
# ## Ranger + MXResnet + Flatten Anneal
# + colab={} colab_type="code" id="ghwm7gIYcy_E"
opt_func = partial(Ranger, betas=(0.9,0.99), eps=1e-6)
learn = Learner(data, mxresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
bn_wd=False, true_wd=True, loss_func=LabelSmoothingCrossEntropy(),
metrics=[accuracy, top_k_accuracy])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="IBw2Sz1Ec8zU" outputId="59793100-e8b9-4e72-b441-6c9e94187309"
flattenAnneal(learn, 4e-3, 5, 0.7)
# + [markdown] colab_type="text" id="xePStNUtfIvx"
# ## RangerLars + MXResnet + Flatten Anneal
# + colab={} colab_type="code" id="53D4GKcefNn-"
opt_func = partial(RangerLars, betas=(0.9,0.99), eps=1e-6)
learn = Learner(data, mxresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
bn_wd=False, true_wd=True, loss_func=LabelSmoothingCrossEntropy(),
metrics=[accuracy, top_k_accuracy])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="LbHTrlnhfR0e" outputId="f751b735-b335-4cc3-d964-72897e80e0dd"
flattenAnneal(learn, 4e-3, 5, 0.72)
# + [markdown] colab_type="text" id="Q9vO4lTZhL-S"
# ## RangerLars + xResnet50 + Flatten Anneal
# + colab={} colab_type="code" id="bkXiMbRshSnG"
opt_func = partial(RangerLars, betas=(0.9,0.99), eps=1e-6)
learn = Learner(data, models.xresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
bn_wd=False, true_wd=True, loss_func=LabelSmoothingCrossEntropy(),
metrics=[accuracy, top_k_accuracy])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="oS6GGZU0hZvl" outputId="0a80bd3e-6c42-4794-880e-fe972d1a7031"
flattenAnneal(learn, 4e-3, 5, 0.72)
# + [markdown] colab_type="text" id="ryhJNYQViFPA"
# ## Ranger + SimpleSelfAttention + MXResnet + Flatten Anneal
# + colab={} colab_type="code" id="_nRLZJmdiJVJ"
opt_func = partial(Ranger, betas=(0.95,0.99), eps=1e-6)
learn = Learner(data, mxresnet50(c_out=10, sa=True), wd=1e-2, opt_func=opt_func,
bn_wd=False, true_wd=True, loss_func=LabelSmoothingCrossEntropy(),
metrics=[accuracy, top_k_accuracy])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="RiWGHmgZldOa" outputId="bd3ef8e1-a4e2-4f68-c908-c60cb63bc712"
flattenAnneal(learn, 4e-3, 5, 0.72)
# + [markdown] colab_type="text" id="wBM72p6cnHtn"
# As we can see, 74.6 is what we got. The highest recorded is 78%.
#
# From here:
#
# I encourage you all to try out some of the combinations seen here today and apply a bit more to it. For instance, are we using the best hyperparameters? What about Cut-Out? MixUp? Plenty more to explore!
# + [markdown] colab_type="text" id="jIjkr4zmn3Fl"
# Thanks to everyone mentioned above for their hard work and determination to getting to where we are now. The fastai forum is an amazing place to bounce ideas and try new things. Also thank you to Jeremy for making *all* of this possible!
| ImageWoofChampionship.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow_gpu] *
# language: python
# name: conda-env-tensorflow_gpu-py
# ---
import torch
from scipy.io import loadmat
import matplotlib.pyplot as plt
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
from sklearn.decomposition import PCA
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
from torch.utils.data import WeightedRandomSampler
from sklearn.preprocessing import StandardScaler, MaxAbsScaler, MinMaxScaler
import numpy as np
from tqdm import tqdm
import warnings
warnings.filterwarnings('ignore')
torch.manual_seed(31415926)
# +
train_data_list = []
train_label_list = []
test_data_list = []
test_label_list = []
# scaler = StandardScaler()
scaler = MinMaxScaler()
# scaler = MaxAbsScaler()
for i in range(1, 11):
mat_data = loadmat("data/train/"+str(i)+".mat")
train_data_list.append(scaler.fit_transform(mat_data['de_feature']))
train_label_list.append(mat_data['label'])
for i in range(11, 14):
mat_data = loadmat("data/test/"+str(i)+".mat")
test_data_list.append(scaler.fit_transform(mat_data['de_feature']))
test_label_list.append(mat_data['label'])
train_datas = np.concatenate(train_data_list)
train_labels = np.concatenate(train_label_list)
test_datas = np.concatenate(test_data_list)
test_labels = np.concatenate(test_label_list)
# pca = PCA(n_components=2)
# train_datas = pca.fit_transform(train_datas)
# test_data_list = [pca.fit_transform(x) for x in test_data_list]
# -
train_data_list[1].shape
class sentimentDataset(Dataset):
def __init__(self, data, labels=None):
self.data = data
self.labels = labels
self.len = data.shape[0]
def __getitem__(self, idx):
data_tensor = torch.tensor(self.data[idx], dtype=torch.float32)
if self.labels is not None:
label_tensor = torch.tensor(self.labels[idx], dtype=torch.long)
return data_tensor, label_tensor
def __len__(self):
return self.len
# +
EPOCHS = 25
DEV_NUM = 0
IN_FEATURE_DIM = 310
class baseline(nn.Module):
def __init__(self):
super().__init__()
self.fc = nn.Sequential(nn.Linear(IN_FEATURE_DIM, 256),
nn.ReLU(True),
nn.Linear(256, 64),
nn.ReLU(True),
nn.Linear(64, 4))
def forward(self, datas, labels=None):
logits = self.fc(datas)
# logit = F.softmax(logits)
outputs = (logits,)
if labels is not None:
loss_fnt = nn.CrossEntropyLoss()
loss = loss_fnt(logits.view(-1, 4), labels.view(-1))
outputs = (loss,) + outputs
return outputs
class cnn_baseline(nn.Module):
def __init__(self, out_channels=256):
super().__init__()
self.conv1 = nn.Conv2d(1, out_channels, kernel_size=[1, 62])
self.fc1 = nn.Sequential(nn.Linear(out_channels, 64),
nn.ReLU(True),
nn.Linear(64, 4))
def forward(self, datas, labels=None):
datas = datas.view(-1, 5, 62) # (batch_size, 5, 62)
expand_datas = datas.unsqueeze(1) # (batch_size, 1, 5, 62)
conved = self.conv1(expand_datas).squeeze() # (batch_size, out_channels, 5)
pooled = F.max_pool1d(conved, kernel_size=5).squeeze() # (batch_size, out_channels)
logits = self.fc1(pooled)
outputs = (logits,)
if labels is not None:
loss_fnt = nn.CrossEntropyLoss()
loss = loss_fnt(logits.view(-1, 4), labels.view(-1))
outputs = (loss,) + outputs
return outputs
# +
def get_predictions(model, dataloader, compute_acc=False):
if torch.cuda.is_available():
model.to("cuda")
model.eval()
predictions = None
correct = 0
total = 0
with torch.no_grad():
for sample in dataloader:
datas = sample[0]
if torch.cuda.is_available():
datas = datas.to("cuda")
outputs = model(datas)
logits = F.softmax(outputs[0], dim=1)
_, pred = torch.max(logits.data, dim=1)
if compute_acc:
labels = sample[1]
if torch.cuda.is_available():
labels = labels.to("cuda")
total += labels.shape[0]
correct += (pred == labels.squeeze()).sum().item()
if predictions is None:
predictions = pred
else:
predictions = torch.cat((predictions, pred))
model.train()
if compute_acc:
acc = correct / total
return predictions, acc
else:
return predictions
def train_model(model, trainset, validloaders: list):
trainloader = DataLoader(trainset, batch_size=64, shuffle=True)
device = torch.device("cuda:"+str(DEV_NUM) if torch.cuda.is_available() else "cpu")
optimizer = torch.optim.Adam(model.parameters(), lr = 1e-4)
model = model.to(device)
model.train()
best_acc = 0.0
for epoch in range(EPOCHS):
running_loss = 0.0
for datas, labels in trainloader:
datas = datas.to(device)
labels = labels.to(device)
optimizer.zero_grad()
outputs = model(datas, labels)
loss = outputs[0]
loss.backward()
optimizer.step()
running_loss += loss.item()
_, train_acc = get_predictions(model, trainloader, compute_acc=True)
pres_and_accs = [get_predictions(model, validloader, compute_acc=True) for validloader in validloaders]
accs = np.array([x[1] for x in pres_and_accs])
print("In epoch %d, running_loss: %.3f, train_acc: %.3f, valid_avg_acc: %.3f," %(epoch, running_loss, train_acc, accs.mean())\
+ " accs: " + str(accs))
print("Training done...")
# -
model = baseline()
trainset = sentimentDataset(train_datas, train_labels)
testsets = [sentimentDataset(test_data_list[i], test_label_list[i]) for i in range(3)]
testloaders = [DataLoader(testset, batch_size=64) for testset in testsets]
train_model(model, trainset, testloaders)
| baselines/NN_baseline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ML Pipeline Preparation
# Follow the instructions below to help you create your ML pipeline.
# ### 1. Import libraries and load data from database.
# - Import Python libraries
# - Load dataset from database with [`read_sql_table`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_sql_table.html)
# - Define feature and target variables X and Y
# +
# import libraries
from sqlalchemy import create_engine
import numpy as np
import pandas as pd
from sklearn.metrics import confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.multioutput import MultiOutputClassifier
from sklearn.metrics import classification_report,confusion_matrix, precision_score,\
recall_score,accuracy_score, f1_score, make_scorer
from sklearn.base import BaseEstimator, TransformerMixin
import nltk
from nltk import word_tokenize
import pickle
# +
# import libraries
import nltk
nltk.download(['punkt', 'wordnet'])
import pandas as pd
import numpy as np
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
from sqlalchemy import create_engine
import sqlite3
from sklearn.pipeline import Pipeline
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.multioutput import MultiOutputClassifier
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
import pickle
# -
# load daata from database
conn = sqlite3.connect('Clean_Messages.db')
df = pd.read_sql('SELECT * FROM Clean_Messages', conn)
df = df.dropna()
X = df["message"]
Y = df.drop("message",1)
# ### 2. Write a tokenization function to process your text data
def tokenize(text):
"""
Takes a Python string object and returns a list of processed words
of the text.
INPUT:
- text - Python str object - A raw text data
OUTPUT:
- stem_words - Python list object - A list of processed words from the input `text`.
"""
tokens = word_tokenize(text)
lemmatizer = WordNetLemmatizer()
clean_tokens = []
for tok in tokens:
clean_tok = lemmatizer.lemmatize(tok).lower().strip()
clean_tokens.append(clean_tok)
return clean_tokens
#remove all non numeric columns from the Y set
Y = Y.drop("id",1)
Y = Y.drop("genre",1)
# ### 3. Build a machine learning pipeline
# This machine pipeline should take in the `message` column as input and output classification results on the other 36 categories in the dataset. You may find the [MultiOutputClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.multioutput.MultiOutputClassifier.html) helpful for predicting multiple target variables.
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf',MultiOutputClassifier(RandomForestClassifier(n_estimators=1000, random_state=0)))
])
# ### 4. Train pipeline
# - Split data into train and test sets
# - Train pipeline
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.33, random_state=42)
pipeline.fit(X_train, y_train)
# ### 5. Test your model
# Report the f1 score, precision and recall for each output category of the dataset. You can do this by iterating through the columns and calling sklearn's `classification_report` on each.
y_pred = pipeline.predict(X_test)
y_pred
y_pred.shape
category_names=Y.columns
y_test
x=0
for column in y_test.columns:
print(classification_report(y_test[column], y_pred[:,x]))
x=x+1
metrics_list_all=[]
for col in range(y_test.shape[1]):
accuracy = accuracy_score(y_test.iloc[:,col], y_pred[:,col])
precision=precision_score(y_test.iloc[:,col], y_pred[:,col],average='micro')
recall = recall_score(y_test.iloc[:,col], y_pred[:,col],average='micro')
f_1 = f1_score(y_test.iloc[:,col], y_pred[:,col],average='micro')
metrics_list=[accuracy,precision,recall,f_1]
metrics_list_all.append(metrics_list)
metrics_df=pd.DataFrame(metrics_list_all,index=category_names,columns=["Accuracy","Precision","Recall","F_1"])
print(metrics_df)
def avg_accuracy_score(y_true, y_pred):
"""
Assumes that the numpy arrays `y_true` and `y_pred` ararys
are of the same shape and returns the average of the
accuracy score computed columnwise.
y_true - Numpy array - An (m x n) matrix
y_pred - Numpy array - An (m x n) matrix
avg_accuracy - Numpy float64 object - Average of accuracy score
"""
# initialise an empty list
accuracy_results = []
# for each column index in either y_true or y_pred
for idx in range(y_true.shape[-1]):
# Get the accuracy score of the idx-th column of y_true and y_pred
accuracy = accuracy_score(y_true[:,idx], y_pred[:,idx])
# Update accuracy_results with accuracy
accuracy_results.append(accuracy)
# Take the mean of accuracy_results
avg_accuracy = np.mean(accuracy_results)
return avg_accuracy
average_accuracy_score =make_scorer(avg_accuracy_score)
list(pipeline.get_params())
# ### 6. Improve your model
# Use grid search to find better parameters.
# +
parameters = [
{
'clf__estimator__max_leaf_nodes': [50, 100, 200],
'clf__estimator__min_samples_split': [2, 3, 4],
}
]
cv = GridSearchCV(pipeline, param_grid=parameters,
scoring=average_accuracy_score,
verbose=10,
return_train_score=True
)
cv.fit(X_train, y_train)
# -
# ### 7. Test your model
# Show the accuracy, precision, and recall of the tuned model.
#
# Since this project focuses on code quality, process, and pipelines, there is no minimum performance metric needed to pass. However, make sure to fine tune your models for accuracy, precision and recall to make your project stand out - especially for your portfolio!
metrics_list_all=[]
for col in range(y_test.shape[1]):
accuracy = accuracy_score(y_test.iloc[:,col], y_pred[:,col])
precision=precision_score(y_test.iloc[:,col], y_pred[:,col],average='micro')
recall = recall_score(y_test.iloc[:,col], y_pred[:,col],average='micro')
f_1 = f1_score(y_test.iloc[:,col], y_pred[:,col],average='micro')
metrics_list=[accuracy,precision,recall,f_1]
metrics_list_all.append(metrics_list)
metrics_df=pd.DataFrame(metrics_list_all,index=category_names,columns=["Accuracy","Precision","Recall","F_1"])
print(metrics_df)
# ### 8. Try improving your model further. Here are a few ideas:
# * try other machine learning algorithms
# * add other features besides the TF-IDF
# ### 9. Export your model as a pickle file
# ### 10. Use this notebook to complete `train.py`
# Use the template file attached in the Resources folder to write a script that runs the steps above to create a database and export a model based on a new dataset specified by the user.
Y.
np.sum(Y.isnull())
| ML Pipeline Preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
#import graphlab
import graphlab
#load data
iris_data = graphlab.SFrame.read_csv("Desktop/Q1 Course/FP/MachineLearningSamples/extra-data/iris.csv")
#remove label column and turn this problem to a unsupervised machine learning problem
iris_data = iris_data.remove_column('species')
iris_data.head()
iris_data.tail()
#train a kmeans model
model = graphlab.kmeans.create(iris_data, num_clusters = 3)
model.summary()
graphlab.canvas.set_target("ipynb")
model.show()
cluster_result = model.cluster_id
#list model result
cluster_result
| Graphlab/Kmeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:analysis3-20.04]
# language: python
# name: conda-env-analysis3-20.04-py
# ---
# # Age at the Bottom of the Ocean
#
# This notebook shows a simple example of plotting ocean Ideal Age. Ideal Age is a fictitious tracer which is set to zero in the surface grid-cell every timestep, and is aged by 1 year per year otherwise. It is a useful proxy for nutrients, such as carbon or oxygen (but not an exact analogue).
#
# One of the interesting aspects of age is that we can use it to show pathways of the densest water in the ocean by plotting a map of age in the lowest gridcell. This plot requires a couple of tricks to extract information from the lowest cell.
#
# **Requirements:** COSIMA Cookbook, preferably installed via the `analysis3-20.04` conda installation on NCI.
#
# Firstly, get all the standard preliminaries out of the way.
# +
# %matplotlib inline
import cosima_cookbook as cc
import matplotlib.pyplot as plt
import numpy as np
import netCDF4 as nc
import xarray as xr
import cartopy.crs as ccrs
import cmocean as cm
from collections import OrderedDict
import cftime
import glob
import logging
logging.captureWarnings(True)
logging.getLogger('py.warnings').setLevel(logging.ERROR)
from dask.distributed import Client
# -
client = Client(n_workers=4)
client
# Add a database session - choose one appropriate for the experiment you would like to interrogate.
session = cc.database.create_session('/g/data/ik11/databases/ryf9091.db')
# Now, let's set the experiment and time interval, and average ideal age over a year.
expt='01deg_jra55v13_ryf9091'
variable = 'age_global'
start_time='2099-01-01'
end_time = '2099-12-31'
age = cc.querying.getvar(expt,variable,session,ncfile='ocean.nc',start_time=start_time, end_time=end_time).sel(time=slice(start_time,end_time)).mean('time').load()
# Here we grab the `kmt` variable out of `ocean_grid.nc`. Note that this is a static variable, so we just look for the last file. This variable tells us the lowest cell which is active at each (x,y) location.
variable='kmt'
kmt = cc.querying.getvar(expt,variable,session,ncfile='ocean_grid.nc', n=-1).fillna(1.0).astype(int) - 1
kmt.load()
# Provided that `kmt` is loaded, `xarray` is smart enough to figure out what this line means, and extracts a 2-D field of bottom age for us.
bottom_age = age.isel(st_ocean=kmt)
# Load some stuff to help with plotting
import cartopy.feature as cft
land_50m = cft.NaturalEarthFeature('physical', 'land', '50m',
edgecolor='face',
facecolor=cft.COLORS['land'])
geolon_t = xr.open_dataset('/g/data/ik11/grids/ocean_grid_01.nc').geolon_t
geolat_t = xr.open_dataset('/g/data/ik11/grids/ocean_grid_01.nc').geolat_t
# And here is the plot:
# +
fig = plt.figure(figsize=(10,6))
ax = plt.axes(projection=ccrs.Robinson(central_longitude=-100))
ax.coastlines(resolution='50m')
ax.add_feature(land_50m,color='gray')
gl = ax.gridlines(draw_labels=False)
p1 = ax.pcolormesh(geolon_t, geolat_t, bottom_age, transform=ccrs.PlateCarree(),
cmap=cm.cm.matter, vmin=60, vmax=200)
plt.title('Ocean Bottom Age')
ax_cb = plt.axes([0.92, 0.25, 0.015, 0.5])
cb = plt.colorbar(p1, cax=ax_cb, orientation='vertical')
cb.ax.set_ylabel('Age (yrs)')
# -
# A few things to note here:
# * The continental shelves are all young - this is just because they are shallow.
# * The North Atlantic is also relatively young, due to formation of NADW. Note that both the Deep Western Boundary Currents and the Mid-Atlantic Ridge both sustain southward transport of this young water.
# * A signal following AABW pathways (northwards at the western boundaries) shows slightly younger water in these regions, but it has mixed somewhat with older water above.
# * Even after 200 years, the water in the NE Pacific has not experienced any ventilation...
| DocumentedExamples/Age_at_the_Bottom.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Installation
# Just pip install:
#
# ```pip install omegaconf```
#
# If you want to try this notebook after checking out the repository be sure to run
# ```python setup.py develop``` at the repository root before running this code.
# # Creating OmegaConf objects
# ### Empty
# + pycharm={"name": "#%%\n"}
from omegaconf import OmegaConf
conf = OmegaConf.create()
print(conf)
# -
# ### From a dictionary
# + pycharm={"name": "#%%\n"}
conf = OmegaConf.create(dict(k='v',list=[1,dict(a='1',b='2')]))
print(OmegaConf.to_yaml(conf))
# -
# ### From a list
# + pycharm={"name": "#%%\n"}
conf = OmegaConf.create([1, dict(a=10, b=dict(a=10))])
print(OmegaConf.to_yaml(conf))
# -
# ### From a yaml file
# + pycharm={"name": "#%%\n"}
conf = OmegaConf.load('../source/example.yaml')
print(OmegaConf.to_yaml(conf))
# -
# ### From a yaml string
# + pycharm={"name": "#%%\n"}
yaml = """
a: b
b: c
list:
- item1
- item2
"""
conf = OmegaConf.create(yaml)
print(OmegaConf.to_yaml(conf))
# -
# ### From a dot-list
# + pycharm={"name": "#%%\n"}
dot_list = ["a.aa.aaa=1", "a.aa.bbb=2", "a.bb.aaa=3", "a.bb.bbb=4"]
conf = OmegaConf.from_dotlist(dot_list)
print(OmegaConf.to_yaml(conf))
# -
# ### From command line arguments
#
# To parse the content of sys.arg:
# + pycharm={"name": "#%%\n"}
# Simulating command line arguments
import sys
sys.argv = ['your-program.py', 'server.port=82', 'log.file=log2.txt']
conf = OmegaConf.from_cli()
print(OmegaConf.to_yaml(conf))
# -
# # Access and manipulation
# Input yaml file:
# + pycharm={"name": "#%%\n"}
conf = OmegaConf.load('../source/example.yaml')
print(OmegaConf.to_yaml(conf))
# -
# #### Object style access:
# + pycharm={"name": "#%%\n"}
conf.server.port
# -
# #### dictionary style access
# + pycharm={"name": "#%%\n"}
conf['log']['rotation']
# -
# #### items in list
# + pycharm={"name": "#%%\n"}
conf.users[0]
# -
# #### Changing existing keys
# + pycharm={"name": "#%%\n"}
conf.server.port = 81
# -
# #### Adding new keys
# + pycharm={"name": "#%%\n"}
conf.server.hostname = "localhost"
# -
# #### Adding a new dictionary
# + pycharm={"name": "#%%\n"}
conf.database = {'hostname': 'database01', 'port': 3306}
# -
# #### providing default values
# + pycharm={"name": "#%%\n"}
conf.get('missing_key', 'a default value')
# -
# #### Accessing mandatory values
# Accessing fields with the value *???* will cause a MissingMandatoryValue exception.
# Use this to indicate that the value must be set before accessing.
# + pycharm={"name": "#%%\n"}
from omegaconf import MissingMandatoryValue
try:
conf.log.file
except MissingMandatoryValue as exc:
print(exc)
# -
# # Variable interpolation
#
# OmegaConf support variable interpolation, Interpolations are evaluated lazily on access.
#
# ## Config node interpolation
# The interpolated variable can be the path to another node in the configuration, and in that case the value will be the value of that node.
# This path may use either dot-notation (``foo.1``), brackets (``[foo][1]``) or a mix of both (``foo[1]``, ``[foo].1``).
#
# Interpolations are absolute by default. Relative interpolation are prefixed by one or more dots: The first dot denotes the level of the node itself and additional dots are going up the parent hierarchy. e.g. **${..foo}** points to the **foo** sibling of the parent of the current node.
# + pycharm={"name": "#%%\n"}
conf = OmegaConf.load('../source/config_interpolation.yaml')
print(OmegaConf.to_yaml(conf))
# -
# Primitive interpolation types are inherited from the referenced value
print("conf.client.server_port: ", conf.client.server_port, type(conf.client.server_port).__name__)
# Composite interpolation types are always string
print("conf.client.url: ", conf.client.url, type(conf.client.url).__name__)
# `to_yaml()` will resolve interpolations if `resolve=True` is passed
print(OmegaConf.to_yaml(conf, resolve=True))
# Interpolations may be nested, enabling more advanced behavior like dynamically selecting a sub-config:
cfg = OmegaConf.create(
{
"plans": {"A": "plan A", "B": "plan B"},
"selected_plan": "A",
"plan": "${plans[${selected_plan}]}",
}
)
print(f"Default: cfg.plan = {cfg.plan}")
cfg.selected_plan = "B"
print(f"After selecting plan B: cfg.plan = {cfg.plan}")
# Interpolated nodes can be any node in the config, not just leaf nodes:
cfg = OmegaConf.create(
{
"john": {"height": 180, "weight": 75},
"player": "${john}",
}
)
(cfg.player.height, cfg.player.weight)
# ## Environment variable interpolation
#
# Access to environment variables is supported using ``oc.env``.
# Let's set up the environment first (only needed for this demonstration)
import os
os.environ['USER'] = 'omry'
# Here is an example config file interpolates with the USER environment variable:
# + pycharm={"name": "#%%\n"}
conf = OmegaConf.load('../source/env_interpolation.yaml')
print(OmegaConf.to_yaml(conf))
# + pycharm={"name": "#%%\n"}
conf = OmegaConf.load('../source/env_interpolation.yaml')
print(OmegaConf.to_yaml(conf, resolve=True))
# -
# You can specify a default value to use in case the environment variable is not set.
# In such a case, the default value is converted to a string using ``str(default)``, unless it is ``null`` (representing Python ``None``) - in which case ``None`` is returned.
#
# The following example falls back to default passwords when ``DB_PASSWORD`` is not defined:
cfg = OmegaConf.create(
{
"database": {
"password1": "${oc.env:DB_PASSWORD,password}",
"password2": "${oc.env:DB_PASSWORD,12345}",
"password3": "${oc.env:DB_PASSWORD,null}",
},
}
)
print(repr(cfg.database.password1))
print(repr(cfg.database.password2))
print(repr(cfg.database.password3))
# ## Decoding strings with interpolations
# With ``oc.decode``, strings can be converted into their corresponding data types using the OmegaConf grammar.
# This grammar recognizes typical data types like ``bool``, ``int``, ``float``, ``dict`` and ``list``,
# e.g. ``"true"``, ``"1"``, ``"1e-3"``, ``"{a: b}"``, ``"[a, b, c]"``.
# It will also resolve interpolations like ``"${foo}"``, returning the corresponding value of the node.
#
# Note that:
#
# - When providing as input to ``oc.decode`` a string that is meant to be decoded into another string, in general
# the input string should be quoted (since only a subset of characters are allowed by the grammar in unquoted
# strings). For instance, a proper string interpolation could be: ``"'Hi! My name is: ${name}'"`` (with extra quotes).
# - ``None`` (written as ``null`` in the grammar) is the only valid non-string input to ``oc.decode`` (returning ``None`` in that case)
#
# This resolver can be useful for instance to parse environment variables:
# +
cfg = OmegaConf.create(
{
"database": {
"port": "${oc.decode:${oc.env:DB_PORT}}",
"nodes": "${oc.decode:${oc.env:DB_NODES}}",
"timeout": "${oc.decode:${oc.env:DB_TIMEOUT,null}}",
}
}
)
os.environ["DB_PORT"] = "3308" # integer
os.environ["DB_NODES"] = "[host1, host2, host3]" # list
os.environ.pop("DB_TIMEOUT", None) # unset variable
print("port (int):", repr(cfg.database.port))
print("nodes (list):", repr(cfg.database.nodes))
print("timeout (missing variable):", repr(cfg.database.timeout))
os.environ["DB_TIMEOUT"] = "${.port}"
print("timeout (interpolation):", repr(cfg.database.timeout))
# -
# ## Custom interpolations
# You can add additional interpolation types using custom resolvers.
# The example below creates a resolver that adds 10 to the given value.
# + pycharm={"name": "#%%\n"}
OmegaConf.register_new_resolver("plus_10", lambda x: x + 10)
conf = OmegaConf.create({'key': '${plus_10:990}'})
conf.key
# -
# You can take advantage of nested interpolations to perform custom operations over variables:
OmegaConf.register_new_resolver("plus", lambda x, y: x + y)
conf = OmegaConf.create({"a": 1, "b": 2, "a_plus_b": "${plus:${a},${b}}"})
conf.a_plus_b
# By default a custom resolver is called on every access, but it is possible to cache its output
# by registering it with ``use_cache=True``.
# This may be useful either for performance reasons or to ensure the same value is always returned.
# Note that the cache is based on the string literals representing the resolver's inputs, and not
# the inputs themselves:
# +
import random
random.seed(1234)
OmegaConf.register_new_resolver("cached", random.randint, use_cache=True)
OmegaConf.register_new_resolver("uncached", random.randint)
cfg = OmegaConf.create(
{
"uncached": "${uncached:0,10000}",
"cached_1": "${cached:0,10000}",
"cached_2": "${cached:0, 10000}",
"cached_3": "${cached:0,${uncached}}",
}
)
# not the same since the cache is disabled by default
print("Without cache:", cfg.uncached, "!=", cfg.uncached)
# same value on repeated access thanks to the cache
print("With cache:", cfg.cached_1, "==", cfg.cached_1)
# same value as `cached_1` since the input is the same
print("With cache (same input):", cfg.cached_2, "==", cfg.cached_1)
# same value even if `uncached` changes, because the cache is based
# on the string literal "${uncached}" that remains the same
print("With cache (interpolation):", cfg.cached_3, "==", cfg.cached_3)
# -
# # Merging configurations
# Merging configurations enables the creation of reusable configuration files for each logical component instead of a single config file for each variation of your task.
#
# Machine learning experiment example:
# ```python
# conf = OmegaConf.merge(base_cfg, model_cfg, optimizer_cfg, dataset_cfg)
# ```
#
# Web server configuration example:
#
# ```python
# conf = OmegaConf.merge(server_cfg, plugin1_cfg, site1_cfg, site2_cfg)
# ```
#
# The following example creates two configs from files, and one from the cli. It then combines them into a single object. Note how the port changes to 82, and how the users lists are combined.
# + pycharm={"name": "#%%\n"}
base_conf = OmegaConf.load('../source/example2.yaml')
print(OmegaConf.to_yaml(base_conf))
# + pycharm={"name": "#%%\n"}
second_conf = OmegaConf.load('../source/example3.yaml')
print(OmegaConf.to_yaml(second_conf))
# + pycharm={"name": "#%%\n"}
from omegaconf import OmegaConf
import sys
# Merge configs:
conf = OmegaConf.merge(base_conf, second_conf)
# Simulate command line arguments
sys.argv = ['program.py', 'server.port=82']
# Merge with cli arguments
conf.merge_with_cli()
print(OmegaConf.to_yaml(conf))
| docs/notebook/Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import torch
import torch.nn as nn
import torch.nn.functional as F
# -
data = pd.read_csv("../../Data/Churn_Modelling.csv")
data.shape
data.head()
data.isnull().any()
data.nunique()
data = data.drop(['RowNumber', 'CustomerId', 'Surname'], axis=1)
data.dtypes
# ### EDA
data.Exited.value_counts().plot(kind='pie', autopct='%1.0f%%',explode=(0.05, 0.05))
sns.countplot(x='Geography', data=data)
sns.countplot(x='Exited',hue='Geography', data=data)
# ### Pre-processing
data.columns
numerical_columns = ['CreditScore', 'Age', 'Tenure', 'Balance', 'NumOfProducts', 'EstimatedSalary']
categorical_columns = ['Geography', 'Gender', 'HasCrCard', 'IsActiveMember']
output = ['Exited']
for cat in categorical_columns:
data[cat] = data[cat].astype('category')
data.dtypes
# +
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
for col in categorical_columns:
data[col] = encoder.fit_transform(data[col])
# -
data.head()
# ### Normalise
# +
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
for col in [numerical_columns,categorical_columns, output]:
data[col] = scaler.fit_transform(data[col])
# -
data.head()
x = data.loc[:, :"EstimatedSalary"]
y = data['Exited']
x.head()
y.head()
print(x.shape, y.shape)
# +
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x,y,test_size=0.3)
# -
print("x_train dim:",x_train.shape, "\ty_train dim:", y_train.shape)
print("x_test dim:",x_test.shape, "\ty_test dim:", y_test.shape)
# ### Convert to tensor
x_train_tensor = torch.tensor(x_train.values, dtype=torch.float)
y_train_tensor = torch.tensor(y_train.values, dtype=torch.long)
x_test_tensor = torch.tensor(x_test.values, dtype=torch.float)
y_test_tensor = torch.tensor(y_test.values, dtype=torch.long)
print("x_train dim:",x_train_tensor.shape, "\ty_train dim:", y_train_tensor.shape)
print("x_test dim:",x_test_tensor.shape, "\ty_test dim:", y_test_tensor.shape)
# ### Model
class Network(nn.Module):
def __init__(self, n_input, h, n_output):
super().__init__()
self.layer = nn.Linear(n_input, h)
self.output = nn.Linear(h, n_output)
def forward(self, x):
x = self.layer(x)
x = self.output(x)
x = torch.sigmoid(x)
return x
n_input, n_output = x_train_tensor.shape[1], 2
h = 100
# +
model = Network(n_input, h, n_output)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
# -
losses = []
epochs = 500
for e in range(1, epochs+1):
y_pred = model(x_train_tensor)
loss = criterion(y_pred, y_train_tensor)
losses.append(loss)
if e%50 ==0:
print(f"epochs: {e} ===> loss:{loss}")
if torch.isnan(loss):
break
optimizer.zero_grad()
loss.backward()
optimizer.step()
plt.plot(range(len(losses)), losses)
plt.xlabel("# epochs")
plt.ylabel("loss")
plt.show()
with torch.no_grad():
y_val = model(x_test_tensor)
loss = criterion(y_val, y_test_tensor)
print("Test loss: ", loss)
y_val = np.argmax(y_val, axis=1)
y_val
# +
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
print(confusion_matrix(y_test_tensor, y_val))
print(classification_report(y_test_tensor, y_val))
accuracy = accuracy_score(y_test_tensor, y_val)*100
print(f'Accuracy: {accuracy:.2f}')
| Classification/Classification-pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Microgrid simulator
# Object-oriented light framework to run custom microgrid resources in lockstep and plot/aggregate results.
# All units are s|W|Ws if not stated otherwise, more about conventions in the glossary.
#
#
# # Table of contents
# 1. [Parameters](#si3Eitei)
# 2. [Model toolset](#eid8ieWi)
# 3. [Tests](#Dei4Uezu)
# 3. [Example](#arg4Uezu)
# 4. [Appendix](#aeZae4ai)
# 4.0. [Glossary](#aec2ieTe)
# 4.1. [Notebook version](#aeDae4ai)
# 4.2. [License](#aeZGe4ai)
#
# ### Model Toolset <a name="eid8ieWi"></a>
# Helper functions to build the model
# +
import math
import random
import time
import os
from collections import OrderedDict
from collections.abc import Iterable
import pickle
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from IPython.core.debugger import set_trace#for ipdb
from IPython.core.display import display, HTML, clear_output
# -
display(HTML("<style>.container { width:98% !important; }</style>"))#this cell width
# %matplotlib inline
np.seterr(all='raise')
plt.rcParams["figure.figsize"] = (15,2.5)
VERBOSE = 0
# control verbose level of simulation_load/store/init/run()
# 0:= silent
# 1:= stringify each resource each step
# 2:= 1 with repr() instead of str()
# #### Parameters
# #### Helper functions and resources
def to_kWh(E_in):
"""takes [Ws]"""
return E_in /1000 /3600
def to_Ws(E_in):
"""takes [kWh]"""
return E_in *1000 *3600
# +
def get_params(param, ct_grp='', j=None):
"""Calculates given parameter p for each timestep k. It implements given formulas
takes param which is the parameter of interest, str
takes ct_grp which specifies the collection from the magg, defaults to all in R, str
takes j specifying the entity in the collection, defaults to all in ct_grp, int
returns param_k in 1,K-ndarray
no-side effects;
"""
param = param.upper()
if ct_grp == '' and j is not None:
raise ValueError(f'no ct_grp selected yet j={j} given')
if param == 'SDR':
if ct_grp == '':#lookup mrss
P_k_prod = get_power(ct_rs=None, ct_flow='prod')
P_k_cons = get_power(ct_rs=None, ct_flow='cons')
if P_k_prod is None:
P_k_prod = np.zeros( (Resource.K) )
if P_k_cons is None:
P_k_cons = np.zeros( (Resource.K) )
E_k_loss_grid = 0
SDR_k = np.ones( (Resource.K) )
dividend = (-1 * P_k_cons + E_k_loss_grid)
np.divide(P_k_prod, dividend, out=SDR_k, where=dividend!=0)#to go around ZeroDivisionError
return SDR_k
else:
P_k_prod_ = get_power_grp(ct_grp, j=j, ct_rs=None, ct_flow='prod')#j=(j or None), indicated by _
P_k_cons_ = get_power_grp(ct_grp, j=j, ct_rs=None, ct_flow='cons')
SDR_k_ = np.ones( (Resource.K) )
dividend = (-1 * P_k_cons_)
np.divide(P_k_prod_, dividend, out=SDR_k_, where=dividend!=0)
return SDR_k_
elif param == 'SSR':
SDR_k_ = get_params('SDR', ct_grp, j)
return np.where(SDR_k_ <= 1, SDR_k_, 1)
elif param == 'SCR':
SDR_k_ = get_params('SDR', ct_grp, j)
return np.where(SDR_k_ >= 1, 1/SDR_k_, 1)
elif param == 'RCR':
if ct_grp == '':
P_k_prod = get_power(ct_rs=None, ct_flow='prod')
P_k_cons = get_power(ct_rs=None, ct_flow='cons')
if P_k_prod is None:
P_k_prod = np.zeros( (Resource.K) )
if P_k_cons is None:
P_k_cons = np.zeros( (Resource.K) )
P_k_res = np.absolute(P_k_prod + P_k_cons)
P_k_res_abs = np.absolute(P_k_prod) + np.absolute(P_k_cons)
out = np.zeros( (Resource.K) )
np.divide(P_k_res, P_k_res_abs, out=out, where=P_k_res_abs!=0)
return 1 - out
else:
P_k_prod_ = get_power_grp(ct_grp, j=j, ct_rs=None, ct_flow='prod')
P_k_cons_ = get_power_grp(ct_grp, j=j, ct_rs=None, ct_flow='cons')
P_k_res_ = np.absolute(P_k_prod_ + P_k_cons_)
P_k_res_abs_ = np.absolute(P_k_prod_) + np.absolute(P_k_cons_)
out = np.zeros( (Resource.K) )
np.divide(P_k_res_, P_k_res_abs_, out=out, where=P_k_res_abs_!=0)
return 1 - out
else:
raise NotImplementedError(f'{param}')
def get_param(param, ct_grp='', j=None):
"""Calculates given parameter p for the simulated time; implements given formulas
takes param which is the parameter of interest, str
takes ct_grp which specifies the collection from the magg, defaults to all in R, str
takes j specifying the entity in the collection, defaults to all in ct_grp, int
returns param value
side effects: reads from current mdata
"""
param = param.upper()
if ct_grp == '' and j is not None:
raise ValueError(f'no ct_grp selected yet j={j} not None')
if param == 'SDR':
return np.mean(get_params(param, ct_grp, j))
elif param == 'SSR':
if ct_grp == '':
P_k_res = get_power()
P_k_res_cons = np.where(P_k_res < 0, P_k_res, 0)
P_k_cons = get_power(ct_flow='cons')
if np.sum(P_k_cons) != 0:
return 1 - np.sum(P_k_res_cons) / np.sum(P_k_cons)
else:#calc is equal to: (pv+dch)/(ld+ch+inv_loss), es loss is to be excl.
P_k_res_ = get_power_grp(ct_grp, j=j)
P_k_res_cons_ = np.where(P_k_res_ < 0, P_k_res_, 0)
P_k_cons_ = get_power_grp(ct_grp, j=j, ct_flow='cons')
if np.sum(P_k_cons_) != 0:
return 1 - np.sum(P_k_res_cons_) / np.sum(P_k_cons_)
return 1#no cons means no res_cons
elif param == 'SCR':
if ct_grp == '':
P_k_res = get_power()
P_k_res_prod = np.where(P_k_res > 0, P_k_res, 0)
P_k_prod = get_power(ct_flow='prod')
if np.sum(P_k_prod) != 0:
return 1 - np.sum(P_k_res_prod) / np.sum(P_k_prod)
else:
P_k_res_ = get_power_grp(ct_grp, j=j)
P_k_res_prod_ = np.where(P_k_res_ > 0, P_k_res_, 0)
P_k_prod_ = get_power_grp(ct_grp, j=j, ct_flow='prod')
if np.sum(P_k_prod_) != 0:
return 1 - np.sum(P_k_res_prod_) / np.sum(P_k_prod_)
return 1#no prod means no res_prod
elif param == 'RCR':
return np.mean(get_params(param, ct_grp, j))
elif param == 'GUR':
if ct_grp == '':
raise NotImplementedError(f'{param}: choose ct_grp')
else:
P_k_J_res = np.zeros( (Resource.K) )
P_k_RJ_res = np.zeros( (Resource.K) )
if j is None:#all j
j_ = range(len(magg[ct_grp]))
else:#one j
j_ = [j]
for j__ in j_:
P_k_J_res += np.absolute(get_power_grp(ct_grp, j=j__))
P_k_RJ_res += get_power_grp(ct_grp, j=j__, ct_flow='prod')
P_k_RJ_res += get_power_grp(ct_grp, j=j__, ct_flow='cons')*-1
if P_k_RJ_res.sum() != 0:
return P_k_J_res.sum() / P_k_RJ_res.sum()
return 1#no RJ_res means no J_res, since |x|+|y| is 0, so is |x+y|
else:
raise NotImplementedError(f'{param}')
# +
def arc_to_deg(arc):
"""convert spherical arc length [m] to great circle distance [deg]"""
return float(arc)/6371/1000 * 180/math.pi
def deg_to_arc(deg):
"""convert great circle distance [deg] to spherical arc length [m]"""
return float(deg)*6371*1000 * math.pi/180
def latlon_to_xyz(lat,lon):
"""Convert angluar to cartesian coordiantes
latitude is the 90deg - zenith angle in range [-90;90]
lonitude is the azimuthal angle in range [-180;180]
"""
r = 6371 # https://en.wikipedia.org/wiki/Earth_radius
theta = math.pi/2 - math.radians(lat)
phi = math.radians(lon)
x = r * math.sin(theta) * math.cos(phi) # bronstein (3.381a)
y = r * math.sin(theta) * math.sin(phi)
z = r * math.cos(theta)
return [x,y,z]
def xyz_to_latlon (x,y,z):
"""Convert cartesian to angular lat/lon coordiantes"""
r = math.sqrt(x**2 + y**2 + z**2)
theta = math.asin(z/r) # https://stackoverflow.com/a/1185413/4933053
phi = math.atan2(y,x)
lat = math.degrees(theta)
lon = math.degrees(phi)
return [lat,lon]
def gen_coords(seed, N, center=(500,500), edge=1000, is_latlon=False):
"""Generates batch of coords inside a square
with specified edge length [m] and its center
takes seed to reproduce batch, int
takes N is batch size of tupels
takes center of square, [m]
takes edge length of square
takes is_latlon bool flag
returns N,2 ndarray
"""
half_edge = (float(edge) / 2)
if is_latlon:
half_edge /= (6371 * 1000 * math.pi/180)#to deg
if abs(center[0])+ half_edge > 90:#reject unhandled corner-cases
raise ValueError('bad lat center due to cornercase')
if abs(center[1])+ half_edge > 180:
raise ValueError('bad lon center due to cornercase')
np.random.seed(seed)#to reproduce batch
batch = np.random.random(2 * N).reshape(2, N)#[0;1[
batch[0] = (center[0] - half_edge) + batch[0] * half_edge * 2
batch[1] = (center[1] - half_edge) + batch[1] * half_edge * 2
return batch.transpose()
def center_point(resources):
"""calculate the coordinate center of rss batch
takes resources list
each dimension is averaged on arithmetic mean
returns the center (x,y,z)
(if: for latlon see geopy.distance.great_circle or geopy.geocoders)
"""
center = [0,0,0]
for rs in resources:
xyz = latlon_to_xyz(rs.cord2, rs.cord1)
center[0] += xyz[0]
center[1] += xyz[1]
center[2] += xyz[2]
center[:] = [axis / len(resources) for axis in center]
return center
# +
def sample_sinus(steps, period=2*math.pi, phase=0, amp=1, offset=0):
"""Sample specified sinus-wave into array
takes period and phase in radians;
takes amplitude and offset to scale and move up/down
returns flat ndarray of steps length
"""
assert steps != 0
samples = np.zeros( (steps) , dtype=np.float64)#default
step_size = period / steps
for i in range(steps):
samples[i] = amp * math.sin(phase + step_size * i) + offset
return samples
def sample_recs(recs, repeat=1):
"""Builds rectangular pattern
takes recs in .,3 ndarray used to build the pattern
start,length are non-negative integers
amp is integer
eg [[start,length,amp],[0,4,2],[2,1,-1]]
takes repeat to concat the period repeat times
returns flat ndarray with specified pulse
"""
size = recs[:,:2].sum(1).max()#number of elements
pattern = np.zeros( (size) )
for rec in recs:
start = rec[0]
end = rec[0] + rec[1]
amp = rec[2]
pattern[start:end] += amp
return np.tile(pattern, repeat)#clone and concat
# +
def get_rss(ct_grp, j=None, ct_rs=None):
"""Access helper to return resources by j
takes ct_grp which is a key from magg, str
takes j the positional index the entity has in the magg, None is all, int
takes ct_rs to include only resources of this category, None is all, str
eg get_rss('hh', 1, 'es')
gets prosumed power es of second entity in magg['hh']
returns list of resources of j/all and [] if no such resource(s)
side-effect: reads from mrss,magg,mmap
"""
ret_val = []
if j is None:#get rss of all j
allr_of_js = magg[ct_grp]
else:#get rss of j
allr_of_js = [magg[ct_grp][j]]
for allr_of_j in allr_of_js:
for r in allr_of_j:
if r not in mmap:
raise KeyError(f'magg[{j}] has r={r}, which is not in mrss')
location = mmap[r]
rs = mrss[location[0]][location[1]]
if ct_rs is not None and ct_rs != rs.ct:
continue
ret_val.append(rs)
return ret_val
def get_power(ct_rs=None, ct_flow=None):
"""Access helper to aggregate resource logs
takes ct_rs to include only resources of this category, None is all, str
if ct_rs does not exist, power returned 0 and no raise
takes ct_flow to include only 'prod' or 'cons', None is both, str
eg get_power('es')
gets prosumed power of all es
returns power in 1,K ndarray, and 1,K zeros if no such resource(s)
side-effect: reads from mrss
"""
ret_val = np.zeros( (Resource.K) )
for key, value in mrss.items():
for rs in value:
if ct_rs is not None and ct_rs != rs.ct:#wrong rs type
continue
if rs.log_index_P is None:
#print('log_index_P',rs)#prints eg inverters
continue
tmp = rs.view[rs.log_index_P]#select power
if ct_flow is None:
ret_val += tmp
elif ct_flow == 'prod':
ret_val += np.where(tmp > 0, tmp, 0)
elif ct_flow == 'cons':
ret_val += np.where(tmp < 0, tmp, 0)
else:
raise ValueError(f"no such flow '{ct_flow}'")
return ret_val
def get_power_grp(ct_grp, j=None, ct_rs=None, ct_flow=None):
"""Access helper to aggregate resource logs by j
takes ct_grp which is a key from magg, str
takes j the positional index the entity has in the magg, None is all, int
takes ct_rs to include only resources of this category, None is all, str
takes ct_flow to include only 'prod' or 'cons', None is both, str
eg get_power_grp('hh', 1, 'es')
gets prosumed power es of second hh in magg['hh']
returns power in 1,K ndarray, and 1,K zeros if no such resource(s)
side-effect: reads from mrss,magg,mmap
"""
if ct_grp not in magg:
raise ValueError(f"ct_grp '{ct_grp}' not in magg")
ret_val = np.zeros( (Resource.K) )
if j is None:#get rss of all j
allr_of_js = magg[ct_grp]
else:#get rss of j
allr_of_js = [magg[ct_grp][j]]
for allr_of_j in allr_of_js:
for r in allr_of_j:
if r not in mmap:
raise KeyError(f'magg[{j}] has r={r}, which is not in mrss')
location = mmap[r]
rs = mrss[location[0]][location[1]]
if ct_rs is not None and ct_rs != rs.ct:
continue
if rs.log_index_P is None:
# print('log_index_P',rs)
continue
tmp = rs.view[rs.log_index_P]#select power
if ct_flow is None:
ret_val += tmp
elif ct_flow == 'prod':
ret_val += np.where(tmp > 0, tmp, 0)
elif ct_flow == 'cons':
ret_val += np.where(tmp < 0, tmp, 0)
else:
raise ValueError(f"no such flow '{ct_flow}'")
return ret_val
# +
def plot_cords(rsss, colors=None, is_latlon=False):
"""plots resources by their geospatial coordinates
takes rsss list of resource lists, eg [[pv0,pv1,..],[ld]]
takes colors list, strings from matplotlib eg ['red','green',..]
takes is_latlon flag, bool
each rss list is offset to prevent overpainting dots
"""
fig, axs = plt.subplots(nrows=1, ncols=1, figsize=(4,4))
for rss,m in zip(rsss, range(len(rsss))):
cords = np.zeros( (len(rss), 2) )
for i in range(len(rss)):
cords[i] = [rss[i].cord1+m*3, rss[i].cord2+m*3]
if colors is None:
plt.scatter(cords[:,0], cords[:,1])
else:
if len(colors) != len(rsss):
raise ValueError(f'number of colors dont match number of rss')
plt.scatter(cords[:,0], cords[:,1], color=colors[m])
axs.set_aspect('equal')
axs.grid(True)
axs.set_title('resource locations [cord1, cord2]')
if is_latlon == True:
xticks = ax.get_xticks()
yticks = ax.get_yticks()
print((f'x_resolution {deg_to_arc(xticks[2] - xticks[1]):.0f}m ' +
f'y_resolution {deg_to_arc(yticks[2] - yticks[1]):.0f}m'))
plt.show()
def plot_logs(rss=[], nums=[], titles=[], use_area=True):
"""Plots one log per given resource. wraps plot_bar
takes rss resources, list
takes nums to pick one log of the resource at the same rss position, list
takes titles for log, optional, list of strings
takse use_area for fast area plot instead of bars, bool
"""
if type(rss) != list:
raise TypeError(f'rss bad type {type(rss)}')
if type(nums) != list:
raise TypeError(f'nums bad type {type(nums)}')
if len(rss) != len(nums):
raise ValueError(f'length not equal rss:{len(rss)} nums:{len(nums)}')
if len(titles) < len(rss):
titles.extend(['']*(len(rss)-len(titles)))
for rs, num in zip(rss, nums):
if num >= len(rs.log_titles):
print(f'num {num} out of bounds for {rs}')
logs = [None] * len(rss)
for i,rs,idc,title in zip(range(len(rss)), rss, nums, titles):#pull logs out of rss
logs[i] = rs.view[idc]
titles[i] = rs.log_titles[idc] + ' ' + title
plot_bar(logs, titles, use_area)
def plot_bar(ys=[[]], titles=[''], use_area=False):
"""Plots multiple data series. wraps pyplot bar
takes list of y lists, eg [[1,2], [3,4]]
takes titles which get appended to each plot title; list of str
takes use_area to use area instead of bars, boolean
"""
if not isinstance(ys[0], Iterable):
raise TypeError(f"'ys' not a list of lists, wrap even one y, eg [[1,2],], have {type(ys[0])}")
if not isinstance(titles[0], Iterable):
raise TypeError(f"'titles' not a list of lists, wrap even on one y, eg [[1,2],], have {type(titles[0])}")
if len(titles) < len(ys):#extend for zip
titles.extend('-' * (len(ys)-len(titles)))
fig, axs = plt.subplots(len(ys),1, figsize=(15,len(ys)*2.5), sharex=True)
if not isinstance(axs,np.ndarray):
axs = np.array( [axs] )
for ax,y,title in zip(axs,ys,titles):
ax.set_title(title)
x = np.arange(len(y))
if use_area:
ax.fill_between(x, y, color='blue', alpha=0.3)
else:
ax.bar(x, y, width=1, align='edge', color='ghostwhite', edgecolor='grey')
plt.show()
# +
def print_(keys_values):
"""verbose helper"""
line = ''
for key,value in keys_values:
if VERBOSE == 1:
line += f'{key}={value}, '
elif VERBOSE == 2:
line += f'{repr(key)}={repr(value)}, '
if line:
print(line)
def _views_to_mrss(mdata):
"""used internally, see function simulation_load"""
r_set = set()
for key, value in mrss.items():#set data
for i in range(len(value)):
view = mdata[key][i]#get slice of cuboid
mrss[key][i].set_view(view)#set to resource
if value[i].r in r_set:#check that r is unique
raise ValueError(f'r of {value[i]} already in use')
else:
r_set.add(value[i].r)
print_([[key, mrss[key][i]]])#ct_grp_rs
def simulation_load(name):
"""push views of mdata into each resource
takes name of binary file in load_store folder to load logs from.
ensure to have the same mrss/rs structure in place
from when it got saved, else mdata mismatches and errors.
rs internal states, except number of logs are irrelevant.
same mmap,magg not needed except when calling helpers, eg get_power
side-effect: writes to mrss,mdata
"""
global mdata
with open('load_store/' + name,"rb") as f:
mdata = pickle.load(f)
_views_to_mrss(mdata)
def simulation_store(name):
"""saves mdata to file
takes name to create file in load_store folder to binary dump.
overwrites existing file
"""
with open('load_store/' + name,"wb") as f:
pickle.dump(mdata,f)
def simulation_init():
"""Generates references alias numpy-views from the data model
and saves them to each resource in the resource model.
eg mrss['hh_ld'][0].view[1] --views--> mdata['hh_ld'][0][1]
Also generates lookup table mmap used internally for 'no-search'
in helper functions when associating resources to collections in magg.
returns mdata, mmap
side-effect: reads mdata, writes mrss
"""
mdata = {}
for key, value in mrss.items():#allocate data
if len(value) == 0:
print(f'please remove empty key {key} from mrss')
cuboid = np.zeros( (len(mrss[key]), len(mrss[key][0].log_titles), Resource.K) )
mdata[key] = cuboid
_views_to_mrss(mdata)
mmap = {}
for key, value in mrss.items():#access map
for i in range(len(value)):
location = (key, i)
mmap[value[i].r] = location
return mdata, mmap
def jump_back(k):
"""Calls jump() on each rs,
errors if at least one rs did not overwrite base class
"""
for key, value in mrss.items():
for rs in value:
print_([['k',k], [key, rs]])
rs.jump(Control.jump_to_k)
return Control.jump_to_k, None#reset flag
def simulation_run(up_to_k, clear_output_=True):
"""Runs the simulation for simulated_time = K * kclock
takes up_to_k to run the simulation up to but excluding this value, int
side-effect write/read on mdata/mrss
"""
if 'ctrl' in mrss and next(iter(mrss.keys())) != 'ctrl':
raise KeyError(f"key 'ctrl' in dict not first")
k = 0#global counter
time_start = time.time()
for key, value in mrss.items():#init round
for rs in value:
rs.init()
print_([['init','-'], [key, rs]])
while k < up_to_k:
for key, value in mrss.items():#rs step ahead of global count
for rs in value:
print_([['k',k], [key, rs]])
rs.step(k)#local step
if clear_output_:
clear_output(wait=True)
time_yet = (time.time() - time_start) / (k+1) * (up_to_k-k-1)#[s] elapsed time per step * steps_yet
print(f'progress k {k} {k+1}/{up_to_k} {(k+1)/up_to_k*100:.0f}% rem {time_yet/60:.2f}m')
k += 1#progress global time
if Control.jump_to_k is not None:#jump requested
k,Control.jump_to_k = jump_back(k)
for key, value in mrss.items():#ended round, k == up_to_k
for rs in value:
rs.end(k)
print_([['k',k], [key, rs]])
# +
class Resource:
"""Base class to build grid components.
Each resource has a unique number r and is at simulation step _k.
The time-log aggregates simulation results for later analysis.
"""
K = None
kclock = None#[s] see notation
def __init__(self, r, ct, cord1=0., cord2=0.):
"""takes r to identify resource, unique, int
takes ct to specify category of this resource, str
takes coordinates, float, optional;
cord1/cord2 for x/y or lon/lat
"""
assert Resource.K is not None, 'set K first'
assert Resource.kclock is not None, 'set kclock first'
self.r = r
self.ct = ct
self.cord1 = cord1
self.cord2 = cord2
self._k = 0
self.view = None#simulation_init
def __repr__(self):
return (f'Resource(r={self.r}, ct={self.ct}, ' +
f'cord1={self.cord1}, cord2={self.cord2}, K={self.K}, _k={self._k}, kclock={self.kclock})')
def __str__(self):
return (f'Resource(r={self.r}, ct={self.ct}, cord1={self.cord2}, cord2={self.cord1})')
def set_view(self, view):
"""Hands in data each resource operates on; Done by simulation_init
"""
self.view = view
def jump(self, k):
"""Set the state to k; to be overwritten by subclasses
takes k the timepoint to set the state to, k >= 0 and k <= current k
Called after each resource concluded the last step and gets
reset to the state at jump_to_k ktime.
Ctrl is the first to be rolled back.
"""
raise NotImplementedError(f'on {self}')
def init(self):
"""Called before simulation starts stepping
"""
def end(self,K):
"""Called once after simulation ended, eg to assert etc
"""
def get_k(self):
"""Returns local k
"""
return self._k
def set_k(self, k):
"""Returns local k
"""
self._k = k
def step(self, k):
"""Perform state transition and run local time behavior.
Progress time on this resource to run step of global counter k
which runs it to k+1 ktime moving it 1 kspan forward
function to be sub-class @extended and called right at start
"""
if k != self._k:
raise ValueError((f'out-of-step self._k={self._k} != '+
f'k={k}, please re-initialize resource model mrss'))
else:
self._k += 1
class TimeSeries(Resource):
"""Predefined power specialized resource
"""
log_titles = (#number of logs read/written during simulation
'P_pros [W]',#0 := produced/consumed power
'E_pros [Ws]'#1 := produced/consumed energy
)
log_index_P = 0#logs index for power prosumed, else None
def __init__(self, *args, **kwargs):
super(TimeSeries, self).__init__(*args, **kwargs)
def __repr__(self):#Optional
return (f'TimeSeries({super(TimeSeries, self).__repr__()}), ' +
f'log_index_P={self.log_index_P}, logs)')
def __str__(self):#Optional
return (f'TimeSeries({super(TimeSeries, self).__repr__()})')
def jump(self, k):#Optional
"""see base class"""
local_k = self.get_k()
if k > local_k:
raise ValueError(f'k {k} greater than local k {local_k}')
if k < local_k:#rolling logs/variables back to k
self.view[1,k:] = 0
self.set_k(k)
def step(self,k):
Resource.step(self, k)
E_prod = self.view[0,k] * Resource.kclock
self.view[1,k] = E_prod
class Control(Resource):
"""Implements microgrid control logic, blueprint
"""
log_titles = ('P_foo [W]',
)
log_index_P = None#index in log_titles that logs power prosumed
jump_to_k = None#set simulation time back to/including k.
#eg current k is 10 and back to 5 makes all rss finish 10 and jump to 5
#useful for eg back-tracking; can be called multiple times; call anywhere in step()
def __init__(self, **kwargs):
super(Control, self).__init__(**kwargs)
def __repr__(self):
return (f'Control({super(Control, self).__repr__()})')
def __str__(self):
return (f'Control({super(Control, self).__repr__()})')
def jump(self, k):
"""see base class
"""
local_k = self.get_k()
if k > local_k:
raise ValueError(f'unable to roll to future, k {k} greater than local k {local_k}')
if k == 0:
self.view[:,k:] = 0#jump all logs
elif k < local_k:
self.view[:,k:] = 0
self.set_k(k)
def init(self):
print(f'simulation about to start')
def end(self, K):
pass
def step(self, k):
Resource.step(self, k)
"""place logic here, make sure to have energy conservation in==out
"""
P_bal = 0#energy conservation
#... place logic here
assert P_bal == 0, 'power inflow != outflow'
# -
# ### Examples <a name="arg4Uezu"></a>
# ### Scenario1:
# The energy residual of one household is logged over one day.
# +----------+
# | GRID |
# | ^ |
# | | |
# | v |
# | LD |
# +----------+
#
class Control(Resource):
"""Implements microgrid control logic, blueprint
"""
log_titles = ('E_residual [Ws]',
)
log_index_P = None#index in log_titles that logs power prosumed
jump_to_k = None
def __init__(self, **kwargs):
super(Control, self).__init__(**kwargs)
def step(self, k):
Resource.step(self, k)
"""place logic here, make sure to have energy conservation in==out
"""
P_bal = 0#energy conservation
P_bal += get_power_grp('hh', j=0, ct_rs='ld')[k]
self.view[0,k] = P_bal*Resource.kclock
P_bal -= P_bal#have it logged, now subtract from balance
assert P_bal == 0, 'power inflow != outflow'
# %%time
Resource.K = 1440# one day on 60 seconds resolution
Resource.kclock = 60
mrss = OrderedDict({
'ctrl': [Control(r=0,ct='ctrl',cord1=0,cord2=0), ],
'hh_ld': [TimeSeries(r=1,ct='ld'),],#load resource
})
magg = {
'hh': [[0,1],],
}
mdata, mmap = simulation_init()
mdata['hh_ld'][0] = np.genfromtxt('time_series/lpg/000.csv', delimiter=",")[:1440]*-1
simulation_run(up_to_k=Resource.K, clear_output_=True)
rs = get_rss('hh', j=0, ct_rs='ctrl')[0]
plot_logs(rss=[rs], nums=[0], titles=[''])
# ### Scenario2:
# One household self-charges his battery to achieve maximum
# self-consumption on its PV system. Wiring:
# +--------------------+
# | GRID |
# | | |
# |PV+-->INV+--->x<->ES|
# | v |
# | LD |
# +--------------------+
#
class Inverter(Resource):
"""Used to model inverter loss DC/AC conversion, true power;
Device not source nor sink, so in/outflow have the same sign.
Choose P_set sign arbitrayly but consistent, eg >0: DC/AC <0:AC/DC,
so to indicate flow direction alias >0:inverter <0:rectifier,
behaviour for +/- on P_set the same;
[1] doi 10.3390/en8064853 model and parameters
[2] https://www.researchgate.net/publication/319351878
"""
log_titles = (#number of logs read/written during simulation
'P_set [W]',#0 := power inflow,
'P_loss [W]'#1 := power loss reducing power outflow
)
log_index_P = 1#logs index for power prosumed, inverter is a 'consumer'
def __init__(self, P_nom, CC, *args, **kwargs):
"""takes P_nom nominal power, float, + or -"""
super(Inverter, self).__init__(*args, **kwargs)
self.P_nom_abs = abs(P_nom)#soft/hard upper bound
self.CC_A = CC[0]
self.CC_B = CC[1]
self.CC_C = CC[2]
self.P_loss = None
def __repr__(self):
return (f'Inverter({super(Inverter, self).__repr__()}), ' +
f'log_index_P={self.log_index_P}, logs)')
def __str__(self):
return (f'Inverter({super(Inverter, self).__repr__()})')
def get_eff(self,k):
"""returns efficiency [0;1] for P_set;
do set P_set at k ktime before calling
"""
P_set = self.view[0,k]
if P_set == 0:
return 1e-3
else:
return (abs(P_set) + self.P_loss) / abs(P_set)
def rollback(self, k):
"""see base class"""
local_k = self.get_k()
if k > local_k:
raise ValueError(f'unable to roll to future, k {k} greater than local k {local_k}')
if k < local_k:#rolling logs/variables back to k
self.view[:,k:] = 0
self.set_k(k)
@staticmethod
def _lookup_eff(P_out):
"""returns eff for given abs(P_out) inverter outflow
lookup tables are precalculated for given inverter
CC_i in simulation are all the same, so one lookup suff.
side-effect: read from lookup_P_out and lookup_eff
"""
idx = (np.abs(lookup_P_out - P_out)).argmin()#nearest
eff = lookup_eff[idx]
if eff == 0:
eff = 1e-3#cvent div0, even on low P_out since lookup nearest
return eff
def _get_eff(self, P_set):
"""calculate eff, corresponds to the backwards lookup"""
eff = self.CC_A * P_set / (self.CC_B - P_set) + self.CC_C * P_set
return eff
def set_P_by_out(self,k, P_out):
"""Set inflow for given outflow
eff from external lookup table (for now)
returns self
"""
P_out_abs = abs(P_out)
eff = Inverter._lookup_eff(P_out_abs)
self.P_loss = (1/eff - 1) * P_out_abs *-1#loss convention
P_set = self.P_loss*-1*math.copysign(1,P_out) + P_out
if abs(P_set) > self.P_nom_abs and False:#and TRUE to not allow P_nom exceed
raise ValueError(f'Inverter P_set={P_set} exceeds P_nom_abs={self.P_nom_abs}')
self.view[0,k] = P_set
self.view[1,k] = self.P_loss
return self
def set_P(self,k, P_set):
"""Sets the power inflow
returns self
"""
if abs(P_set) > self.P_nom_abs and False:#s.a.
raise ValueError(f'Inverter P_set={P_set} exceeds P_nom_abs={self.P_nom_abs}')
if P_set == 0:
eff = 1
else:
eff = self._get_eff(abs(P_set))
self.P_loss = (1 - eff) * abs(P_set) *-1#loss convention
self.view[0,k] = P_set
self.view[1,k] = self.P_loss
return self
def step(self,k):
Resource.step(self, k)
#loss precalculated when set
class BatteryRPL(Resource):
"""Prosumage specialized resource with three states charge/discharge/idle
BatteryRPL rewrites the loss of BatteryR from constant to a percentage of
the current soc. This models the effect of increased loss on high socs
Furthermore it adds idle losses to charge,discharge powers equal or
below pivot_P_idle. This models the effect of idle loss when 'about' idle.
Since the model/parameters are highly depended on the batt/circuit/env,
abstraction prevents misconfig, yet models relevant effects for scheduling.
Maximum charge/discharge power = function(soc)
Idle loss = percent * current_soc
Efficiency = constant
[0] http://science.sciencemag.org/content/334/6058/928
[1] http://dx.doi.org/10.1016/j.jpowsour.2012.10.060
[2] https://doi.org/10.1016/j.electacta.2017.10.153
[3] https://www.researchgate.net/publication/319351878
[4] https://doi.org/10.1007/s11465-018-0516-8
"""
log_titles = ('P_ext [W]',#1 := P_ext produced/consumed power
'E_ext [Ws]',#2 := E_ext produced/consumed energy
'E_loss [Ws]',#3 := E_loss energy loss, be it charge/discharge/idle
'E_soc [Ws]',#4 := E_soc total energy stored at k ktime
)
log_index_P = 0#index in log_titles that logs power prosumed
def __init__(self, E_soc, E_soc_max, P_ch_const, P_dch_const, pct_dch_idle, eff_rt, pivot_soc_dch=0, *args, **kwargs):
"""constructor for initial state
takes E_soc and E_soc_max for initial and maximum state of charge, <=0
takes P_ch_const and P_dch_const for desired charge/discharge
these are eventually undercut but not exceeded
takes pct_dch_idle percent of current soc before the step to assign to loss
takes eff_rt round-trip efficiency to simulate loss, [0;1]
"""
assert E_soc <= 0 and E_soc >= E_soc_max, 'bad args'
assert P_ch_const <= 0 and P_dch_const >= 0, 'bad args'
assert eff_rt <= 1 and eff_rt >= 0, 'bad args'
super(BatteryRPL, self).__init__(*args, **kwargs)
self.E_soc = float(E_soc)
self.E_soc_max = float(E_soc_max)
self.P_ext = None
self.E_ext = None
self.E_loss = None
self.P_ch_const = float(P_ch_const)
self.P_dch_const = float(P_dch_const)
self.P_set = 0#fix, use set_P
"""Charge and discharge power are limited for operational safety
Charge power is constant, but linearly decreases if pivot_soc_ch exceeded.
Discharge power is constant, but instant 0 if pivot_soc_dch undercut.
power ch |---linear increase--|---------const--------|
soc soc_max pivot_soc_ch* 0
power dch |------------const----------|-------0-------|
soc soc_max pivot_soc_dch* 0
"""
self.pivot_soc_ch = 0.9#[0;1]
self.pivot_soc_dch = pivot_soc_dch#[0;1]
self.pivot_P_ch = 1/3#[0;1] of P_ch_const to charge at soc_max
self.P_ch_max = np.zeros( (1,2) )#P_dch_max for [eff_pivot, eff_rt]
self.P_dch_max = np.zeros( (1,2) )
"""Idle losses are a fraction of the current soc, positive float"""
self.pct_dch_idle = abs(float(pct_dch_idle))
"""charge and discharge efficiency instant-drops on low power rate
Charge/discharge are subject to loss of eff_pivot or sqrt(eff_rt)
eff |---------sqrt(eff_rt)--------|-----+idle-----|
power P_ch/dch_const pivot_P_idle 0
"""
self.pivot_P_idle = abs(10.)#inspired by [3]
self.eff_rt = float(eff_rt)
self.update_max_ch_dch_power()
def __repr__(self):#verbose
return (f'BatteryRPL({super(BatteryRPL, self).__repr__()}, ' +
f'E_soc={self.E_soc}, E_soc_max={self.E_soc_max}, '+
f'E_ext={self.E_ext}, E_loss={self.E_loss}, '+
f'P_ch_const={self.P_ch_const}, P_dch_const={self.P_dch_const}, '+
f'P_set={self.P_set}, pivot_soc_ch={self.pivot_soc_ch}, '+
f'pivot_soc_dch={self.pivot_soc_dch}, pivot_P_ch={self.pivot_P_ch}, '+
f'P_ch_max={self.P_ch_max}, P_dch_max={self.P_dch_max}, '+
f'pct_dch_idle={self.pct_dch_idle}, pivot_P_idle={self.pivot_P_idle}, '+
f'eff={self.eff_rt}, log_index_P={self.log_index_P}, logs)')
def __str__(self):
return f'BatteryRPL({super(BatteryRPL, self).__repr__()})'
def rollback(self, k):
"""see base class"""
local_k = self.get_k()
if k > local_k:
raise ValueError(f'unable to roll to future, k {k} greater than local k {local_k}')
if k == 0:
self.P_set = 0
self.P_ext = None
self.E_ext = None
self.E_loss = None
self.E_soc = self.view[3,0]#from start
elif k < local_k:#rolling logs/variables back to k
self.P_set = self.view[0,k-1]#same as P_ext
self.P_ext = self.view[0,k-1]
self.E_ext = self.view[1,k-1]
self.E_loss = self.view[2,k-1]
self.E_soc = self.view[3,k]#from k
self.view[:,k:] = 0
self.set_k(k)
self.update_max_ch_dch_power()
def set_P(self, k, P_set):
"""charge/discharge/idle power -/+/0 respectively, int/float
|---P_ch----|---P_dch---|
P_ch_max -0+ P_dch_max
"""
P_set = float(P_set)
self.view[0, k] = P_set#expose right after set but before step for get_power(_grp)
if P_set >= self.P_ch_max and P_set <= self.P_dch_max:
self.P_set = P_set
else:
raise ValueError((f'limits exceeded ' +
f'ch<=have<=dch {self.P_ch_max}<={P_set}<={self.P_dch_max}'))
def update_max_ch_dch_power(self):
#charge limits pysical
if self.E_soc == self.E_soc_max:#full limit
self.P_ch_max = 0
elif self.E_soc <= self.pivot_soc_ch * self.E_soc_max:#pivot reached
"""linear function interpolating charging power between two points
(E_soc_max,pivot_P_ch*P_ch_const) (pivot_soc_ch*E_soc_max,P_ch_const)
"""
m = ((self.P_ch_const - self.pivot_P_ch * self.P_ch_const) /
(self.pivot_soc_ch * self.E_soc_max - self.E_soc_max))
t = (self.P_ch_const - m * self.pivot_soc_ch * self.E_soc_max)
self.P_ch_max = m * self.E_soc + t
else:
self.P_ch_max = self.P_ch_const
#charge limits discretization
P_ch_max_suff = (self.E_soc_max - self.E_soc) / math.sqrt(self.eff_rt) / Resource.kclock
#charge limits, necessary and sufficient
self.P_ch_max = max(self.P_ch_max, P_ch_max_suff)*(1-1e-9)#prevent flip soc>0
#discharge limits pysical
if self.E_soc >= self.pivot_soc_dch * self.E_soc_max:#pivot reached
self.P_dch_max = 0
else:#necessary
self.P_dch_max = self.P_dch_const
#discharge limits discretization, sufficient
P_dch_max_suff = -1 * self.E_soc * math.sqrt(self.eff_rt) / Resource.kclock
#discharge limits, necessary and sufficient
self.P_dch_max = min(self.P_dch_max, P_dch_max_suff)*(1-1e-9)
if self.P_dch_max < 25:
self.P_dch_max = 0
def step(self,k):
Resource.step(self, k)
E_ext = self.P_set * Resource.kclock#externally prosumed
self.view[3,k] = self.E_soc#at k ktime
eff = math.sqrt(self.eff_rt)
if self.P_set > 0:#discharge
self.P_ext = self.P_set
self.E_ext = E_ext
self.E_loss = (E_ext / eff - E_ext) * -1#loss sign convention
self.E_soc += E_ext / eff
elif self.P_set < 0:#charge
self.P_ext = self.P_set
self.E_ext = E_ext
self.E_loss = E_ext * (1 - eff)
self.E_soc += E_ext * eff
elif self.P_set == 0:#idle
self.P_ext = 0
self.E_ext = 0
self.E_loss = self.pct_dch_idle * self.E_soc
self.E_soc += self.pct_dch_idle * self.E_soc * -1
if self.P_set != 0 and abs(self.P_set) <= self.pivot_P_idle:#add idle loss on low ch/dch P
self.E_loss += self.pct_dch_idle * self.E_soc
self.E_soc += self.pct_dch_idle * self.E_soc * -1
assert self.E_soc <= 0, 'soc flipped'#probably prec. err
assert abs(self.E_soc - (self.view[3,k] + self.E_ext + self.E_loss*-1))<1e-6#energy conservation
if self.E_soc > -1e-9:#classify as loss
self.E_loss += self.E_soc
self.E_soc = 0
#log to data store, at k kspan
self.view[0,k] = self.P_ext
self.view[1,k] = self.E_ext
self.view[2,k] = self.E_loss
#transition done, forward simulation time on device
self.update_max_ch_dch_power()
# curve-fit inverter
from scipy.optimize import curve_fit
def func(x, a, b, c):
return a*x / (b-x) + c*x
xdata = np.array([0, 10, 100, 1000, 2000, 5000, 1e4])
ydata = np.array([0, 0.1, 0.45, 0.97, 0.94, 0.92, 0.8])
popt, pcov = curve_fit(func, xdata, ydata) #inverter coeff ABC
# idle for battery
pct_dch_idle = 1-(95/100)**(1/(30*24*60))# 5% idle loss over the month to 1min kclock
soc = -10
for k in range(30*24*60):#verify
soc = soc * (1 - pct_dch_idle)
print(f'start {-10}kWh, end month {soc:.3}kWh, {1-soc/-10:.2}%, pct_dch_idle {pct_dch_idle*1e2:.3}%/min')
class Control(Resource):
"""Implements microgrid control logic, blueprint
"""
log_titles = ('P_j_inv_out [W]',
'P_j_res [W]',
)
log_index_P = None#index in log_titles that logs power prosumed
jump_to_k = None
def __init__(self, **kwargs):
super(Control, self).__init__(**kwargs)
def step(self, k):
Resource.step(self, k)
"""place logic here, make sure to have energy conservation in==out
"""
P_bal = 0#energy conservation
P_j_pv = get_power_grp('hh', j=0, ct_rs='pv')[k]
P_j_ld = get_power_grp('hh', j=0, ct_rs='ld')[k]
rs_inv = get_rss('hh', j=0, ct_rs='inv')[0]#unpack the only inv
rs_es = get_rss('hh', j=0, ct_rs='es')[0]#unpack the only es
P_j_pv_inv = rs_inv.set_P(k, P_j_pv).get_eff(k) * P_j_pv#reduced by inv. loss
P_j_res = P_j_pv_inv + P_j_ld
if P_j_res > 0:#oversupply
if P_j_res > rs_es.P_ch_max*-1:#if unable to consume oversupply
rs_es.set_P(k, rs_es.P_ch_max)
self.view[1,k] = P_j_res + rs_es.P_ch_max#left overs flow to extern
else:
rs_es.set_P(k, P_j_res*-1)#residual zeroed
else:#undersupply
if P_j_res*-1 > rs_es.P_dch_max:
rs_es.set_P(k, rs_es.P_dch_max)
self.view[1,k] = P_j_res + rs_es.P_dch_max#inflow from extern
else:
rs_es.set_P(k, P_j_res*-1)#residual zeroed
self.view[0,k] = P_j_pv_inv#log inverter output
assert P_bal == 0, 'power inflow != outflow'
# %%time
Resource.K = 1440# one day on 60 seconds resolution
Resource.kclock = 60
mrss = OrderedDict({
'ctrl': [Control(r=0,ct='ctrl',cord1=0,cord2=0), ],
'hh_ld': [TimeSeries(r=1,ct='ld'),],#load resource
'hh_pv': [TimeSeries(r=2,ct='pv')],
'hh_inv': [Inverter(r=3,ct='inv',P_nom=3500, CC=popt)],
'hh_es': [BatteryRPL(r=4,ct='es',E_soc=-to_Ws(1),\
E_soc_max=-to_Ws(5), P_ch_const=-4500,P_dch_const=5000,\
pct_dch_idle=pct_dch_idle, eff_rt=0.9), ],
})
magg = {
'hh': [[0,1,2,3,4],],
}
mdata, mmap = simulation_init()
mdata['hh_ld'][0] = np.genfromtxt('time_series/lpg/000.csv', delimiter=",")[:1440]*-1 #!negative
mdata['hh_pv'][0] = np.genfromtxt('time_series/crest/pv_1qm_clearness_index_varies.csv', delimiter=",").transpose()[0,:1440]*10#10qm
simulation_run(up_to_k=Resource.K, clear_output_=True)
# +
ctrl = get_rss('hh', j=0, ct_rs='ctrl')[0]
es = get_rss('hh', j=0, ct_rs='es')[0]
plot_logs(rss=[ctrl,ctrl,es], nums=[0,1,3], titles=['','', ''])
# -
# ### Scenario3:
# Two households balance their load on one shared battery. They try to balance out supply and demand for maximum SSR and SCR
# +-----------------------------------------+
# | |
# | +----------------+GRID |
# | | | |
# | PV+-->INV+----+-+ PV+-->INV+--->x<->ES |
# | v v |
# | LD LD |
# +-----------------------------------------+
class Control(Resource):
"""Implements microgrid control logic, blueprint
"""
log_titles = ('P_J_inv_out [W]',
'P_J_res [W]',
)
log_index_P = None#index in log_titles that logs power prosumed
jump_to_k = None
def __init__(self, **kwargs):
super(Control, self).__init__(**kwargs)
def step(self, k):
Resource.step(self, k)
"""place logic here, make sure to have energy conservation in==out
"""
P_bal = 0#energy conservation
P_j0_pv = get_power_grp('hh', j=0, ct_rs='pv')[k]
P_j1_pv = get_power_grp('hh', j=1, ct_rs='pv')[k]
P_J_ld = get_power_grp('hh', j=None, ct_rs='ld')[k]
rs_j0_inv = get_rss('hh', j=0, ct_rs='inv')[0]
rs_j1_inv = get_rss('hh', j=1, ct_rs='inv')[0]
rs_es = get_rss('hh', j=1, ct_rs='es')[0]
P_j0_pv_inv = rs_j0_inv.set_P(k, P_j0_pv).get_eff(k) * P_j0_pv
P_j1_pv_inv = rs_j1_inv.set_P(k, P_j1_pv).get_eff(k) * P_j1_pv
P_J_res = P_j0_pv_inv + P_j1_pv_inv + P_J_ld
if P_J_res > 0:#oversupply
if P_J_res > rs_es.P_ch_max*-1:#if unable to consume oversupply
rs_es.set_P(k, rs_es.P_ch_max)
self.view[1,k] = P_J_res + rs_es.P_ch_max#left overs flow to extern
else:
rs_es.set_P(k, P_J_res*-1)#residual zeroed
else:#undersupply
if P_J_res*-1 > rs_es.P_dch_max:
rs_es.set_P(k, rs_es.P_dch_max)
self.view[1,k] = P_J_res + rs_es.P_dch_max#inflow from extern
else:
rs_es.set_P(k, P_J_res*-1)#residual zeroed
self.view[0,k] = P_j0_pv_inv + P_j1_pv_inv#log total inverter output
assert P_bal == 0, 'power inflow != outflow'
# %%time
Resource.K = 1440# one day on 60 seconds resolution
Resource.kclock = 60
mrss = OrderedDict({
'ctrl': [Control(r=0,ct='ctrl',cord1=0,cord2=0), ],
'hh_ld': [TimeSeries(r=1,ct='ld'),TimeSeries(r=5,ct='ld')],#load resource
'hh_pv': [TimeSeries(r=2,ct='pv'),TimeSeries(r=6,ct='pv')],
'hh_inv': [Inverter(r=3,ct='inv',P_nom=3500, CC=popt), Inverter(r=7,ct='inv',P_nom=3500, CC=popt)],
'hh_es': [BatteryRPL(r=4,ct='es',E_soc=-to_Ws(0),\
E_soc_max=-to_Ws(5), P_ch_const=-4500,P_dch_const=5000,\
pct_dch_idle=pct_dch_idle, eff_rt=0.9), ],
})
magg = {
'hh': [[5,6,7], [0,1,2,3,4]],
}
mdata, mmap = simulation_init()
mdata['hh_ld'][0] = np.genfromtxt('time_series/lpg/000.csv', delimiter=",")[:1440]*-1
mdata['hh_ld'][1] = np.genfromtxt('time_series/lpg/001.csv', delimiter=",")[:1440]*-1
mdata['hh_pv'][0] = np.genfromtxt('time_series/crest/pv_1qm_clearness_index_varies.csv', delimiter=",").transpose()[0,:1440]*10#10qm
mdata['hh_pv'][1] = np.genfromtxt('time_series/crest/pv_1qm_clearness_index_varies.csv', delimiter=",").transpose()[0,:1440]*20#15qm
simulation_run(up_to_k=Resource.K, clear_output_=True)
# +
ctrl = get_rss('hh', j=1, ct_rs='ctrl')[0]
es = get_rss('hh', j=1, ct_rs='es')[0]
plot_logs(rss=[ctrl,ctrl,es], nums=[0,1,3], titles=['','', ''], use_area=True)
# -
plot_bar([get_power_grp('hh', ct_rs='ld')[700:1000]], ['total load [W] k700:k1000'])
# #### Powerful queries allow verifying the community SSR right away:
f"The SSR of the community is {get_param(param='SSR', ct_grp='hh')} of total consumed {to_kWh(get_power_grp('hh', ct_flow='cons')*Resource.kclock).sum():.6f}kWh incl. loss"
res = get_power_grp('hh')
E_J_res = to_kWh(res[res<0].sum() * Resource.kclock)
E_J_cons = to_kWh((get_power_grp('hh', ct_flow='cons').sum()) * Resource.kclock)
1-E_J_res/E_J_cons#same as above, fin
to_kWh((get_power_grp('hh', ct_flow='prod').sum()) * Resource.kclock),\
to_kWh((get_power_grp('hh', ct_rs='pv').sum()) * Resource.kclock),\
to_kWh((get_power_grp('hh', ct_rs='es', ct_flow='prod').sum()) * Resource.kclock)#prod = pv + es_dch
# ### Appendix <a name="aeZae4ai"></a>
# #### Glossary <a name="aec2ieTe"></a>
# abbreviations
# pv photovoltaic system
# es energy storage system
# soc state of energy charge
# inv inverter
# hh household
# dof degree of freedom
# jpn jupyter notebook
# xfmr transformer
#
# E_ energy [Ws], ∈ |R float
# P_ power [W], ∈ |R float
# C_ cost, <0 negative cost is revenue, >0 is cost
# ∈ |R float denoted in [curr]
# c_ cost per unit,
# ∈ |R+ float, denoted in [curr/Ws]
# sub-idices
# res residual
# prod produced
# cons consumed
# r resource index
# r=foo selector, only resources of type foo
# R number of resources
# const constant
# bal balance
# ctrl control
# ch charge
# dch discharge
# J collection size of ct_grp in aggregator model
# j collection/peer index, ∈ [0;J-1]
# <type> specifier eg load|pv|es|res|...
#
# time scale
# k simulation time counter, ∈ [0;K-1]
# increments each step to advance time
# K number of simulation steps
#
# kclock step time, maps simulation time to clock-time, ∈ |R or |N, [s]
# describes time passed each step,
# modifying the value is ok and changes granularity of the data
#
# modifying its unit requires resource adjustments
# imported resource models with different step value/unit must
# be adjusted, eg 2[ms] model must step 500* per k if kclock 1[s]
# common units are [s|m|h|d]
# units are ktime, kspan, ctime, cspan for concrete k
# at k ktime is a timepoint
# at k kspan is one timespan and starts at k ktime
# for k kspan is a timespan of length k with unspecified start
#
# kr used to specify relative timespans, see examples
# ekE divides k axis into epochs of length E ∈ |N
# e = int(k/E) converts k to epoch e
# k = e*E converts epoch e to k
#
# sign convention
# <0 := outflow alias consume or export
# >0 := inflow alias produce or import
# from system point of view consumer counting arrow system
# from resource pov called producer counting arrow system
# the system is a virtual construct only for energy conservation
# the pysical grid would be a loss afflicted resource itself
# loss is always negative, since it reduces the system's energy amount
# +------------------+
# | +--------+ |
# | |resource|<-cons |<-imp
# | +--------+->prod |->exp,
# | system |
# +------------------+
#
# examples
# P_k_res of any k
# P_K_res, P_res for all k summarized
# P_j_res of any j
# P_J_res for all j summarized, since there are multiple J's it remains;
# k2 simulation time after two steps 0-1-now
# in text called 2 ktime or 4 ctime with kclock=2
# K * kclock total simulated time in ctime
# P_k10,j0,load power load of peer 0 at ktime 10
# P_j0,k1,pv photovoltaic power of peer 0 at 1 ktime
# P_j,k1,pv photovoltaic power of unspecified peer at 1 ktime
# P_j0:9,res total residual power of peers [0;8]
# P_J,res total residual power of all peers
# time irrelevant/omitted or specified elsewhere
# P_J,pv total photovoltaic power of all peers
# E_K,res total residual energy over simulated time K kspan
# P_r0,k1 power of resource with index 0 at 1 ktime
# P_R total power of all devices
# if peers hold all resources, == P_J,res
# E_k10:20 energy sum over 10 kspan from k [10;19]
# E_kr0:10 energy sum over 10 kspan; 'when' unkown
# E_2k15 energy sum over 15 kspan from k [30;44]
#
# + language="latex"
# \begin{align}
# E_{k} & = \intop_{k}^{k+kclock} P(t) dt = P_{k} * kclock\\
# E & = (P) * kclock = (\sum_{k=1}^{K} P_{k}) * kclock = \sum_{k=1}^{K} (P_{k} * kclock) \\
# \end{align}
# -
# energy power relation
# due to discretization it is constant during kclock simplifying the integral.
# energy/power calculations can be done distributive a*(b+c)= a*b+a*c
# + language="latex"
# \begin{align}
# P_{k,r} & = \begin{cases}
# P_{k,r,prod} \quad & \text{if } P_{k,r} \geq 0 \text{ producer}\\
# P_{k,r,cons} \quad & \text{if } P_{k,r} < 0 \text{ consumer}\\
# \end{cases}\\
# P_{k,r,res} & = P_{k,r,prod} + P_{k,r,cons} \\
# \end{align}
# -
# resource sign conventions, outflow positive, inflow negative
# addon: P_k,r,res makes only sense if one resource has two ports.
# #### Power
# + language="latex"
# \begin{align}
# P_{k,j,prod} & = \sum_{i=1}^{R \land j} P_{k,j,i,prod} \quad \text{and}\quad P_{k,j,cons} = \sum_{i=1}^{R \land j} P_{k,j,i,cons}\\
# P_{k,j,res} & = P_{k,j,prod} + P_{k,j,cons} = \sum_{i=1}^{R \land j} P_{k,j,i} \\
# P_{J,res} & = \sum_{j=1}^{J} P_{j,res} \quad \text{and}\quad
# P_{R,prod} = \sum_{r=1}^{R} P_{r,prod}\\
# \end{align}
# -
# power produced and consumed of peer j at k ktime over its own resource set
# power residual of peer j at k ktime over its own resource set
# total power of all peers at k ktime
# total power of all resources at k ktime
# ### Parameters<a name="si3Eitei"></a>
# + language="latex"
# \begin{align}
# SSR_{k,j} & = 1 - \frac{P_{k,j,res,cons}}{P_{k,j,cons}}\\
# SSR_{j} & = 1 - \frac{P_{j,res,cons}}{P_{j,cons}}\\
# SCR_{k,j} & = 1 - \frac{P_{k,j,res,prod}}{P_{k,j,prod}}\\
# SCR_{j} & = 1 - \frac{P_{j,res,prod}}{P_{j,prod}}\\
# \end{align}
# -
# self-sufficiency-rate of peer j at k ktime
# self-sufficiency-rate of peer j for simulated time
# analog the self-consumption rate
# + language="latex"
# \begin{align}
# SDR_{k,j} & = \dfrac{ P_{k,j,prod} }{ -P_{k,j,cons} } \quad \text{if } P_{k,j,cons} \text{ not 0 else 1}\\
# SDR_{j} & = \dfrac{ P_{j,prod} }{ -P_{j,cons} }\\
# SDR_{k} & = \frac{ \sum_{r=1}^{R} P_{k,r,prod} }{ -\sum_{i=1}^{R} P_{k,r,cons} }\\
# SDR & = \frac{ \sum_{r=1}^{R} P_{r,prod} }{ -\sum_{i=1}^{R} P_{r,cons} }\\
# \end{align}
# -
# supply-demand-ratio of peer j at k ktime
# supply-demand-ratio of peer j
# the second two formulas remove the peer and use the r index to make a general statement
# + language="latex"
# \begin{align}
# RCR_{k} & = 1 - \frac{ |\sum_{j=1}^{J} P_{k,j,res}| }{ \sum_{j=1}^{J} |P_{k,j,res}| }\\
# RCR_{k,j} & = 1 - \frac{ |\sum_{i=1}^{R \land j} P_{k,r}| }{ \sum_{i=1}^{R \land j} |P_{k,r}| }\\
# \end{align}
# -
# residual-cancellation-rate at k ktime
# describes anticyclic relation of residual load, [0;1]
# the closer to 1 the better peers complement each other in residual power;
# describes destructive interference; if P_kJ_res = 0 the RCR_k is 1
# example for RCR = 0 and RCR = 1
# +---+ +---+
# |j0 | |j0 |
# +-------+-> +--------->
# |j1 | k |j1 | k
# +---+ +---+
# residual-cancellation-rate of peer j at k ktime
# if P_k_Rofj = 0 the RCR_kj is 1
# + language="latex"
# \begin{align}
# GUR_{J} & = \frac{ \sum_{k=1}^{K} \sum_{j=1}^{J} |P_{k,j,res}| }
# { \sum_{k=1}^{K} \sum_{i=1}^{R \land J} |P_{k,r}| }\\
# \end{align}
# -
# grid-utilization rate
# measure for the percentage of energy flow over smart-meters from J subsystems in/out of the system; or
# total absolute residual power of all peers (P_J_res) divided by total absolute power of peer rss; or
# of all energyflow how much went over the grid
# #### Loss
# + language="latex"
# \begin{align}
# E_{k,r,loss} & = \begin{cases}
# E_{k,r,loss,ch} \quad & \text{if } P_{k,r,set} < 0 \text{ charge loss and ct_rs='es'}\\
# E_{k,r,loss,idle} \quad & \text{if } P_{k,r,set} = 0 \text{ idle loss and ct_rs='es'}\\
# E_{k,r,loss,dch} \quad & \text{if } P_{k,r,set} > 0 \text{ discharge loss and ct_rs='es'}\\
# \end{cases}\\
#
# E_{k,loss,es} & = \sum_{i=1}^{|es|} E_{k,i,loss,es}\\
#
# E_{k,loss,es,idle} & = \sum_{i=1}^{|es|} E_{k,i,loss,es,idle}\\
#
# E_{k,j,loss,es} & = \sum_{i=1}^{|es|\land j} E_{k,j,i,loss,es}\\
#
# E_{k,R,loss} & = \sum_{r=1}^{R} E_{k,r,loss}\\
#
# E_{k,loss} & = E_{k,R,loss} + E_{k,loss,grid}\\
# \end{align}
# -
# energy loss of resource r of category 'es' at k kspan
# since the es is only in one state, non-state losses are considered 0
# total energy loss of all es at k kspan
# total idle energy loss of all es at k kspan
# total energy loss of peer j and its es at k kspan
# total energy loss of all resources at k kspan
# total energy loss of the system at k kspan
# #### Notebook version <a name="aeDae4ai"></a>
#
from IPython.utils.sysinfo import get_sys_info
get_sys_info()
# #### License notice <a name="aeZGe4ai"></a>
# MIT License
#
# Copyright (c) 2019
# -----BEGIN PUBLIC KEY-----
# MIIBIjANBgkqhkiG9w0BAQEFAAOCAQ8AMIIBCgKCAQEAtAgeP1hhQHBHPICTc2ho
# vJFNXA2qtf0HjuXXV7i+imaN7RI4hUPQMo4nNCYjeiD3vzAdBTtWRQrI2ONmiFTk
# ntAuD0Mg03q+mj/88aawnZbtXBF4QM5sYClInIuW23uhSq17SseWCXtEhmHtz155
# 4LllN4FBC11/R0shrAvFH4dAn2sM8PBg+FGze2wUaJbEl2rLe+qoek10krbSrpUP
# VXCsyVyicR1IaOhldH4I8zpvB6CSPzOkzhQhbxRhxvKwN7kaVlzVGg2u3ccgffHP
# dldIk2D14rz0hJ0Ix1qheAQW/+2haBP/lbwW2iLtiyC47sVeDbCpd66Zi9lKDUe4
# nwIDAQAB
# -----END PUBLIC KEY-----
#
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all
# copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
# SOFTWARE.
# https://tldrlegal.com/license/mit-license
| template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Important classes of Spark SQL and DataFrames:
#
# - :class:`pyspark.sql.SQLContext`
# Main entry point for :class:`DataFrame` and SQL functionality.
# - :class:`pyspark.sql.DataFrame`
# A distributed collection of data grouped into named columns.
# - :class:`pyspark.sql.Column`
# A column expression in a :class:`DataFrame`.
# - :class:`pyspark.sql.Row`
# A row of data in a :class:`DataFrame`.
# - :class:`pyspark.sql.HiveContext`
# Main entry point for accessing data stored in Apache Hive.
# - :class:`pyspark.sql.GroupedData`
# Aggregation methods, returned by :func:`DataFrame.groupBy`.
# - :class:`pyspark.sql.DataFrameNaFunctions`
# Methods for handling missing data (null values).
# - :class:`pyspark.sql.DataFrameStatFunctions`
# Methods for statistics functionality.
# - :class:`pyspark.sql.functions`
# List of built-in functions available for :class:`DataFrame`.
# - :class:`pyspark.sql.types`
# List of data types available.
# - :class:`pyspark.sql.Window`
# For working with window functions.
# +
from pyspark import SparkContext
#sc.stop()
sc = SparkContext(master="local[3]")
from pyspark import SparkContext
from pyspark.sql import *
sqlContext = SQLContext(sc)
# -
# ## DataframeStatFunctions
#
# Methods for statistics functionality. [documented here](http://takwatanabe.me/pyspark/generated/generated/pyspark.sql.DataFrameStatFunctions.html)
#
# * **approxQuantile(col, probabilities, relativeError)** Calculates the approximate quantiles of a numerical column of a DataFrame.
# * **corr(col1, col2[, method])** Calculates the correlation of two columns of a DataFrame as a double value.
# * **cov(col1, col2)** Calculate the sample covariance for the given columns, specified by their names, as a double value.
# * **crosstab(col1, col2)** Computes a pair-wise frequency table of the given columns.
# * **freqItems(cols[, support])** Finding frequent items for columns, possibly with false positives.
# * **sampleBy(col, fractions[, seed])** Returns a stratified sample without replacement based on the fraction given on each stratum.
# +
# DataFrameStatFunctions.corr?
# -
| Sections/Section1-Spark-Basics/2.SparkSQL/4.More on spark.sql.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2020년 9월 1일 화요일
# ### leetCode - Self Dividing Numbers (Python)
# ### 문제 : https://leetcode.com/problems/self-dividing-numbers/
# ### 블로그 : https://somjang.tistory.com/entry/leetCode-728-Self-Dividing-Numbers-Python
# ### 첫번째 시도
class Solution:
def selfDividingNumbers(self, left: int, right: int) -> List[int]:
answer = [num for num in range(left, right+1) if '0' not in str(num) and all([num % int(n) == 0 for n in str(num)])]
return answer
| DAY 101 ~ 200/DAY195_[leetCode] Self Dividing Numbers (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
my_list = [1,2,3,4,5,6,7]
for number in my_list:
print(number)
# ## range
range(20)
list(range(20))
for number in list(range(20)):
print(number * 5 )
for num in list(range(5,21,4)):
print(num)
# ## enumerate
index = 0
for number in list(range(5,15)):
print(f"no: {number} ix: {index}")
index += 1
for element in enumerate(list(range(5,15))):
print(element)
for (index,number) in enumerate(list(range(5,15))):
print(index)
print(number)
# ## random
from random import randint
randint(0,1000)
randint(0,1000)
my_list_2 = list(range(0,10))
my_list_2
from random import shuffle
shuffle(my_list_2)
my_list_2
# ## zip
sport_list = ["run","swim","basketball"]
calories_list = [100,200,300]
day_list = ["monday","tuesday","wednesday"]
new_list = list(zip(sport_list,calories_list,day_list))
new_list
for element in new_list:
print(element)
# ## list advanced
# +
new_list = []
my_string = "metallica"
for element in my_string:
new_list.append(element)
# -
new_list
new_list = [element for element in my_string]
new_list
new_list_2 = [number**5 for number in list(range(0,10))]
new_list_2
| PythonCourse-master/PythonCourse-master/16-UsefulMethods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import scipy.io as sio
import numpy as np
import matplotlib.pyplot as plt
from numpy import matmul as mm
import math
data = sio.loadmat('practice.mat')
# -
pose = data['pose']; ranges = data['ranges']
scanAngles = data['scanAngles']; t = data['t']
tmp1 = ranges[:,0].reshape(-1,1)*np.cos(scanAngles)
tmp2 = -ranges[:,0].reshape(-1,1)*np.sin(scanAngles)
lidar_local = np.hstack((tmp1,tmp2))
plt.figure(figsize=(20,10))
plt.plot(0,0,'rs')
plt.plot(lidar_local[:,0],lidar_local[:,1],'.-')
plt.axis('equal')
plt.gca().invert_yaxis()
plt.xlabel('x'); plt.ylabel('y')
plt.grid(True)
plt.title('Lidar measurement in the body frame')
| Robotics/EstimationAndLearning/Week3/.ipynb_checkpoints/example_lidar-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Prestack seismic
#
# **[Smaller single gather file on S3/agilegeo (3.8GB)](https://s3.amazonaws.com/agilegeo/3D_gathers_pstm_nmo_X1001.sgy)**
#
# **[Larger gathers files on Open Seismic Repository (ca. 10GB)](https://opendtect.org/osr/pmwiki.php/Main/PENOBSCOT3DSABLEISLAND)**
#
# For now we'll satisfy ourselves with reading some prestack seismic data from disk, and looking at it.
#
# Eventually we can:
#
# - Extract prestack attributes from the gathers.
# - Examine a well from one of the gather locations.
# - Model the AVO behaviour at the well and compare to the prestack data.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Read the SEGY file
#
# This is a large file — and this is only 1 of 30 or so files in this ca. 90GB dataset – so we will use `headonly=True` to only parse the headers with `_read_segy()`, then we can index into the stream as before. The difference is that this time, the data stays on disk until we do that read.
filename = '../data/3D_gathers_pstm_nmo_X1001.sgy'
# +
from obspy.io.segy.segy import _read_segy
# Only read the headers, otherwise you will get memory issues
stream = _read_segy(filename, headonly=True)
# -
stream
x = np.array(list(stream.textual_file_header.decode()))
print('\n'.join(''.join(row) for row in x.reshape((40, 80))))
# ## Organize the data
# This is where we get the data from disk. We'll just grab a bit. There are about 22 traces per gather, so we'll go for 100-ish gathers.
#
# The tricky thing with this dataset is that there are a variable number of traces per gather, so we can't just read it like a regular 3D.
traces_to_read = 2200
data = np.vstack([t.data for t in stream.traces[:traces_to_read]])
gno = np.array([t.header.trace_number_within_the_ensemble for t in stream.traces[:traces_to_read]])
# We'll need the number of time samples.
_, t = data.shape
# Find the max number of traces in a gather.
values, counts = np.unique(gno, return_counts=True)
# Collect all the traces according to the trace header no.
gathers = np.vstack([data[gno==i] for i in values])
# +
# Make a mask, False where there's no trace for that gather.
mask = np.arange(np.amax(counts)) < counts[:, None]
# We have to mask with a 1D array so make the out one row per trace.
out = np.zeros((mask.size, t))
out[np.ravel(mask)] = gathers
# +
lines, traces = mask.shape
# Reshape back to the 3D geometry.
g3 = out.reshape(lines, traces, t)
# -
plt.figure(figsize=(3,14))
plt.imshow(g3[68].T, cmap='Greys', aspect='auto')
plt.show()
# The stack is formed by averaging these traces, usually a swath of angles. So fewer traces contribute near the top; more at the bottom. The complete stack would look like this:
# +
idx = np.arange(0, g3.shape[-1])
perc = np.percentile(g3, 95)
plt.figure(figsize=(8,20))
plt.imshow(g3[67].T, cmap='Greys', aspect=.1)
plt.plot(np.mean(g3[67], axis=0)/perc, idx)
plt.ylim(400, 0)
plt.show()
# -
# ## Make an angle stack
#
# We'll pretend there's a constant velocity of 2000 m/s to make life easier. Then 1 ms corresponds to 1 m, so with a sample interval of 4 ms, we're looking at 4 m samples in depth.
#
# In the header, it says this:
#
# output offset pannels 175m - 3175m with bin size 50m
#
# I don't totally understand the bin size remark. We have 22 traces, we'll assume the near trace is 175 m offset, and the far is 3175 m.
#
# $$ \tan \theta = \frac{x}{d} \ \ \mathrm{so} \ \ d = \frac{x}{\tan \theta} $$
# +
theta = np.radians(30)
d = 4*idx / np.tan(theta)
# We need the index.
incr = 3175 / 22
d_i = 1 + (d // incr).astype(int)
d_i[d_i >= 22] = 22
# -
d_i[:40]
plt.figure(figsize=(6,14))
plt.imshow(g3[68].T, cmap='Greys', interpolation='none', aspect='auto')
plt.plot(d_i, idx)
plt.ylim(1000, 0)
plt.show()
# We'll apply that as a mute. We'll make `NaN`s so we can easily drop them out of the mean.
# Bah, there has to be a more elegant way...
for gather in g3:
for row, i in zip(gather.T, d_i):
row[i:] = np.nan
plt.figure(figsize=(6,14))
plt.imshow(g3[68].T, cmap='Greys', interpolation='none', aspect='auto')
plt.plot(d_i, idx)
plt.ylim(1000, 0)
plt.show()
# ## Make a stacked section
plt.figure(figsize=(16,16))
plt.imshow(np.nanmean(g3.T, axis=1), cmap='Greys', vmin=-perc, vmax=perc, interpolation='none', aspect='auto')
plt.show()
# ## The power of stack
# Remember, one of the points of stacking is noise reduction. Let's look at a really noisy trace and see what happens when we stack it. We'll use the mean trace from before:
tr = np.nanmean(g3[68], axis=0)/perc
panel = np.repeat(tr, 50).reshape(tr.size, 50)
panel += np.random.random(panel.shape) * np.ptp(panel) - np.ptp(panel)/2
plt.figure(figsize=(16,8))
plt.imshow(panel, cmap="Greys", interpolation='none', aspect='auto')
plt.show()
plt.figure(figsize=(16,2))
plt.plot(tr)
plt.plot(np.nanmean(panel, axis=1))
plt.show()
plt.figure(figsize=(16,2))
plt.plot(tr[600:800])
plt.plot(np.nanmean(panel, axis=1)[600:800])
plt.show()
# <hr />
#
# <div>
# <img src="https://avatars1.githubusercontent.com/u/1692321?s=50"><p style="text-align:center">© Agile Geoscience 2016</p>
# </div>
| notebooks/Prestack_seismic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
pd.set_option('max_columns', None)
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
# +
mission = "../../missions.csv"
mission_df = pd.read_csv(mission, index_col='ID').drop(columns=['Unnamed: 0'])
astronauts = "../../astronauts.csv"
astronauts_df = pd.read_csv(astronauts).drop(columns=['Unnamed: 0', 'id', 'original_name'])
# -
astronauts_df.head(5)
#### astronauts_df['occupation'] = astronauts_df['occupation'].str.lower()
astronauts_df.loc[astronauts_df['occupation'] == 'space tourist', 'occupation'] = 'other (space tourist)'
# Check for the number of null
astronauts_df.isnull().sum()
def impute_shuttles(row) -> pd.Series:
if 'Soyuz' in row['in_orbit']:
row['ascend_shuttle'] = row['in_orbit']
row['descend_shuttle'] = row['in_orbit']
return row
astronauts_df.loc[astronauts_df['in_orbit'].str.contains("Soyuz")]
missing_shuttles = astronauts_df.loc[astronauts_df['ascend_shuttle'].isna()]
missing_shuttles = missing_shuttles.apply(impute_shuttles, axis=1)
astronauts_df.loc[astronauts_df['ascend_shuttle'].isna()] = missing_shuttles
# astronauts_df.loc[astronauts_df['mission_title'].isna()]
astronauts_df.loc[(astronauts_df['year_of_mission'] == 2019) & (astronauts_df['ascend_shuttle'].str.contains('Soyuz 13'))]
missing_mission_titles = astronauts_df.loc[astronauts_df['mission_title'].isna()]
missing_mission_titles['mission_title'] = 60
astronauts_df.loc[astronauts_df['mission_title'].isna()] = missing_mission_titles
# +
# Select all Selection process in US
astronauts_df.loc[(astronauts_df['nationality'] == "U.S.") & (astronauts_df['military_civilian'] == "civilian")]['selection'].unique()
astronauts_df.loc[astronauts_df['nationality'] == "U.K./U.S."]
astronauts_df.loc[astronauts_df['selection'] == "MirCorp"]
# -
missing_selection = astronauts_df.loc[astronauts_df['selection'].isna()]
missing_selection['selection'] = 'MirCorp'
astronauts_df.loc[astronauts_df['selection'].isna()] = missing_selection
print("Total missing values in the entire astronauts dataset:", astronauts_df.isnull().sum().sum())
astronauts_df.info()
mission_df.info()
mission_df.sample(5)
mission_df['Cost'] = mission_df['Cost'].str.replace(',', '')
mission_df['Cost'] = pd.to_numeric(mission_df['Cost'])
mission_df['Cost'].describe()
mission_df.loc[mission_df['Company Name'] == "Arm??e de l'Air", ['Company Name']] = "Arme de l'Air"
mission_df['Country'] = mission_df['Location'].str.extract(r'^.*?([^\t,]*)$')
mission_df['Country'] = mission_df['Country'].str.strip()
mission_df[['Vehicle', 'Rocket']] = mission_df['Detail'].str.split('|', expand=True)
mission_df.groupby('Country').size()
mission_df[~(mission_df['Date'].str.match(pat=".*(?=\d{2}:\d{2} [A-Z]{3})"))].sample(10)
usa_missions = mission_df.loc[(mission_df['Country'] == "India") & (mission_df['Cost'].notnull())]
usa_missions.groupby('Company Name').agg(min_cost=('Cost', 'min'),
max_cost=('Cost', 'max'),
median_cost=('Cost', np.median),
count=('Cost','count'))
null_usa_missions = mission_df.loc[(mission_df['Country'] == "India") & (mission_df['Cost'].isnull())]
null_usa_missions.groupby('Company Name').size()
mission_df[(mission_df['Country'] == "India") & mission_df['Cost'].notnull()]
mission_df.loc[mission_df['Vehicle'].str.contains("PSLV")]
| src/notebooks/version_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: spline_dist(tf)
# language: python
# name: tf
# ---
# +
from __future__ import print_function, unicode_literals, absolute_import, division
import sys
import numpy as np
import matplotlib
matplotlib.rcParams["image.interpolation"] = None
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
from glob import glob
from tqdm import tqdm
from tifffile import imread
from csbdeep.utils import Path, normalize
from splinedist import fill_label_holes, random_label_cmap, calculate_extents, gputools_available
from splinedist.matching import matching, matching_dataset
from splinedist.models import Config2D, SplineDist2D, SplineDistData2D
np.random.seed(42)
lbl_cmap = random_label_cmap()
import splinegenerator as sg
from splinedist.utils import phi_generator, grid_generator, get_contoursize_max
import os
os.environ['CUDA_VISIBLE_DEVICES']='0'
# -
# # Data
#
# We assume that data has already been downloaded via notebook [1_data.ipynb](1_data.ipynb).
#
# <div class="alert alert-block alert-info">
# Training data (for input `X` with associated label masks `Y`) can be provided via lists of numpy arrays, where each image can have a different size. Alternatively, a single numpy array can also be used if all images have the same size. Label images need to be integer-valued.
# </div>
X = sorted(glob('data/dsb2018/train/images/*.tif'))
Y = sorted(glob('data/dsb2018/train/masks/*.tif'))
assert all(Path(x).name==Path(y).name for x,y in zip(X,Y))
X = list(map(imread,X))
Y = list(map(imread,Y))
n_channel = 1 if X[0].ndim == 2 else X[0].shape[-1]
# Normalize images and fill small label holes.
print(n_channel)
# +
axis_norm = (0,1) # normalize channels independently
# axis_norm = (0,1,2) # normalize channels jointly
if n_channel > 1:
print("Normalizing image channels %s." % ('jointly' if axis_norm is None or 2 in axis_norm else 'independently'))
sys.stdout.flush()
X = [normalize(x,1,99.8,axis=axis_norm) for x in tqdm(X)]
Y = [fill_label_holes(y) for y in tqdm(Y)]
# -
# Split into train and validation datasets.
assert len(X) > 1, "not enough training data"
rng = np.random.RandomState(42)
ind = rng.permutation(len(X))
n_val = max(1, int(round(0.15 * len(ind))))
ind_train, ind_val = ind[:-n_val], ind[-n_val:]
X_val, Y_val = [X[i] for i in ind_val] , [Y[i] for i in ind_val]
X_trn, Y_trn = [X[i] for i in ind_train], [Y[i] for i in ind_train]
print('number of images: %3d' % len(X))
print('- training: %3d' % len(X_trn))
print('- validation: %3d' % len(X_val))
# Training data consists of pairs of input image and label instances.
def plot_img_label(img, lbl, img_title="image", lbl_title="label", **kwargs):
fig, (ai,al) = plt.subplots(1,2, figsize=(12,5), gridspec_kw=dict(width_ratios=(1.25,1)))
im = ai.imshow(img, cmap='gray', clim=(0,1))
ai.set_title(img_title)
fig.colorbar(im, ax=ai)
al.imshow(lbl, cmap=lbl_cmap)
al.set_title(lbl_title)
plt.tight_layout()
i = min(9, len(X)-1)
img, lbl = X[i], Y[i]
assert img.ndim in (2,3)
img = img if (img.ndim==2 or img.shape[-1]==3) else img[...,0]
plot_img_label(img,lbl)
None;
# # Configuration
#
# A `SplineDist2D` model is specified via a `Config2D` object.
np_load_new = lambda *a,**k: np.load(*a, allow_pickle=True, **k)
Y_trn = np_load_new('Y_train.npy')
X_trn = np_load_new('Y_train.npy')
X_val = np_load_new('X_val.npy')
Y_val = np_load_new('Y_val.npy')
print(Config2D.__doc__)
print(Y_trn[0].shape)
# + code_folding=[]
# choose the number of control points (M)
M = 8
n_params = 2 * M
n_channel = 1
# Predict on subsampled grid for increased efficiency and larger field of view
grid = (2,2)
# compute the size of the largest contour present in the image-set
contoursize_max = get_contoursize_max(Y_trn)
conf = Config2D (
n_params = n_params,
grid = grid,
n_channel_in = n_channel,
contoursize_max = contoursize_max,
)
# -
phi_generator(M, conf.contoursize_max)
grid_generator(M, conf.train_patch_size, conf.grid)
model = SplineDist2D(conf, name='splinedist', basedir='models')
# Check if the neural network has a large enough field of view to see up to the boundary of most objects.
# +
# median_size = calculate_extents(list(Y), np.median)
# fov = np.array(model._axes_tile_overlap('YX'))
# print(f"median object size: {median_size}")
# print(f"network field of view : {fov}")
# if any(median_size > fov):
# print("WARNING: median object size larger than field of view of the neural network.")
# -
# # Data Augmentation
# You can define a function/callable that applies augmentation to each batch of the data generator.
# We here use an `augmenter` that applies random rotations, flips, and intensity changes, which are typically sensible for (2D) microscopy images (but you can disable augmentation by setting `augmenter = None`).
# +
def random_fliprot(img, mask):
assert img.ndim >= mask.ndim
axes = tuple(range(mask.ndim))
perm = tuple(np.random.permutation(axes))
img = img.transpose(perm + tuple(range(mask.ndim, img.ndim)))
mask = mask.transpose(perm)
for ax in axes:
if np.random.rand() > 0.5:
img = np.flip(img, axis=ax)
mask = np.flip(mask, axis=ax)
return img, mask
def random_intensity_change(img):
img = img*np.random.uniform(0.6,2) + np.random.uniform(-0.2,0.2)
return img
def augmenter(x, y):
"""Augmentation of a single input/label image pair.
x is an input image
y is the corresponding ground-truth label image
"""
x, y = random_fliprot(x, y)
x = random_intensity_change(x)
# add some gaussian noise
sig = 0.02*np.random.uniform(0,1)
x = x + sig*np.random.normal(0,1,x.shape)
return x, y
# -
# plot some augmented examples
img, lbl = X[0],Y[0]
plot_img_label(img, lbl)
for _ in range(3):
img_aug, lbl_aug = augmenter(img,lbl)
plot_img_label(img_aug, lbl_aug, img_title="image augmented", lbl_title="label augmented")
# # Training
model.train(X_trn, Y_trn, validation_data=(X_val,Y_val), augmenter=augmenter, epochs = 300)
# # Visualization
# First predict the labels for all validation images:
Y_val_pred = [model.predict_instances(x, n_tiles=model._guess_n_tiles(x), show_tile_progress=False)[0]
for x in tqdm(X_val)]
# Plot a GT/prediction example
plot_img_label(X_val[0],Y_val[0], lbl_title="label GT")
plot_img_label(X_val[0],Y_val_pred[0], lbl_title="label Pred")
| splinedist/training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dependencies
import pandas as pd
from splinter import Browser
from bs4 import BeautifulSoup as bs
import pymongo
import requests
from webdriver_manager.chrome import ChromeDriverManager
# Set up of Chrome webdriver
executable_path = {"executable_path": ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
# Scraping preparation and store data in dictionary
get_mars_data = {}
url = 'https://mars.nasa.gov/news/'
response= requests.get(url)
soup = bs(response.text, 'html.parser')
# # NASA Mars News:
# Scrape the NASA Mars News Site and collect the latest News Title and Paragraph Text. Assign the text to variables that you can reference later.
# +
news_title = soup.find('div', class_= 'content_title').text
news_paragraph = soup.find('div', class_= 'rollover_description_inner').text
print('Most Recent Nasa News Article...')
print(f'Title: {news_title}')
print(f'Substance: {news_paragraph}')
# Push values to mars dictionary
get_mars_data['recent_news'] = news_title
get_mars_data['recent_news_substance'] = news_paragraph
# -
# # JPL Mars Space Images - Featured Image
# Visit the url for JPL Featured Space Image here. Use splinter to navigate the site and find the image url for the current Featured Mars Image and assign the url string to a variable called featured_image_url.
# +
# Url we will be scraping images from
url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
base_url = 'https://www.jpl.nasa.gov'
response = requests.get(url)
soup = bs(response.text, 'html.parser')
splint_url = base_url + soup.find('a', class_="button fancybox")["data-fancybox-href"]
print(f"URL to Featured Nasa Image: {splint_url}")
# -
# # Mars Facts
# Visit the Mars Facts webpage here and use Pandas to scrape the table containing facts about the planet including Diameter, Mass, etc.Use Pandas to convert the data to a HTML table string.
# +
url = 'https://space-facts.com/mars/'
# Read table data from url
facts_table = pd.read_html(url)
# Convert to dataframe
mars_facts_df = facts_table[0]
mars_facts_df.columns = ['Type', 'Measurement']
mars_facts_df
# -
#create HTML table
html_table = mars_facts_df.to_html(border=3)
#Remove enter characters
get_mars_data['mars_facts_html'] = html_table.replace('\n', '')
print(get_mars_data['mars_facts_html'])
# # Mars Hemispheres
# Visit the Mars Facts webpage here and use Pandas to scrape the table containing facts about the planet including Diameter, Mass, etc.Use Pandas to convert the data to a HTML table string.
# +
url = 'https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
base_url = "https://astrogeology.usgs.gov"
# Obtain the webpage
response = requests.get(url)
soup = bs(response.text, 'html.parser')
# Grab all image urls and append to list
results = soup.find_all('a', class_="itemLink product-item")
full_res_img_url = []
for result in results:
# Combine link and base url
full_res_img_url.append(base_url + result['href'])
print(full_res_img_url)
# +
#create a empty list for diction
hem_img_urls = []
base_url = 'https://astrogeology.usgs.gov'
for url in full_res_img_url:
# Obtain webpage from diff website
response = requests.get(url)
soup = bs(response.text, 'html.parser')
#Retrieve url to full resolution image
image_url = soup.find('div', class_="downloads").find('ul').find('li').find('a')['href']
#Retrieve the subject
title = soup.find('h2', class_="title").text
#initial diction and put into list
res_dict = { "title":title,"img_url": image_url }
hem_img_urls.append(res_dict)
print(title)
print(image_url)
print(hem_img_urls)
get_mars_data['hemisphere_image_urls'] = hem_img_urls
#print all data from diction
print(get_mars_data)
# -
# Print all data from mars dictionary
print(get_mars_data)
| mission_to_mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="J4vCjuSlDx-_"
# # "RH Analytics e Machine Learning"
# > "Análise exploratória e preditiva de um dataset da IBM sobre rotatividade de funcionários - Post sobre o projeto publicado no Medium"
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Data Science, RH Analytics, Modelos de classificação, Tree-based models, Dados não balanceados, Projeto]
# + [markdown] id="SSY93U0MNowR"
# 
# + [markdown] id="VBdCmjPxMEjq"
# > Link: https://fmarcelneves.medium.com/rh-analytics-e-machine-learning-e584635d7d87?source=friends_link&sk=7720272d4edf972c1131d51c18a457cf
| _notebooks/2021-02-20-RH-Analytics-e-Machine-Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="fKeQUmclBWXn"
# ## Import libraries
# + id="Ns6AoQi17lD_" executionInfo={"status": "ok", "timestamp": 1643918476601, "user_tz": -60, "elapsed": 30, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
from google.colab import drive
from pathlib import Path
from matplotlib import pyplot as plt
import pandas as pd
import numpy as np
import time
import os
import csv
import concurrent.futures
# + [markdown] id="EHH7RkSyBXvi"
# ## Utility functions
# + [markdown] id="F9yqiodWBaLc"
# ### Create annot and load descriptors
# + id="EFyndPYP7p4l" executionInfo={"status": "ok", "timestamp": 1643918476605, "user_tz": -60, "elapsed": 30, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def create_annot(path):
image_list = list(Path(path).glob('*/*.jpg'))
# the identity name is in the path (the name of the parent directory)
names_list = [i.parent.name for i in image_list] # get the identity of each image
# keep info in a pandas DataFrame
annot = pd.DataFrame({'identity': names_list, 'image_path': image_list})
return annot
def concatenate_annots(list_of_paths):
concat_annot = pd.DataFrame()
with concurrent.futures.ThreadPoolExecutor() as executor:
annots = [executor.submit(create_annot, path) for path in list_of_paths]
for annot in annots:
new_annot = annot.result()
concat_annot = concat_annot.append(new_annot, ignore_index = True)
return concat_annot
# + id="QWcji5EX_CGz" executionInfo={"status": "ok", "timestamp": 1643918476607, "user_tz": -60, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def load_descriptors(path):
with open(path, 'rb') as file:
return np.load(file)
def concatenate_descriptors(list_of_paths):
concat_descriptors = None
with concurrent.futures.ThreadPoolExecutor() as executor:
descriptors = [executor.submit(load_descriptors, path) for path in list_of_paths]
for descriptor in descriptors:
new_descriptor = descriptor.result()
if concat_descriptors is None:
concat_descriptors = new_descriptor
else:
concat_descriptors = np.concatenate([concat_descriptors, new_descriptor])
return concat_descriptors
# + [markdown] id="TCmdeNDNBfUg"
# ### Create pivots
# + id="ZG4HAdh4Bp9D" executionInfo={"status": "ok", "timestamp": 1643918476609, "user_tz": -60, "elapsed": 31, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def generate_pivots(descriptors, n, strategy="rnd"):
if strategy == "kMED":
kmedoids = sklearn_extra.cluster.KMedoids(n_clusters=n).fit(descriptors)
return kmedoids.cluster_centers_
if strategy != "rnd":
print(strategy, "was not implemented. Random pivots were returned")
pivots_id = np.random.choice(np.arange(len(descriptors)), size=n)
return descriptors[pivots_id]
def generate_list_of_pivots(descriptors, t, n, strategy="rnd"):
list_of_pivots = []
with concurrent.futures.ThreadPoolExecutor() as executor:
pivots = [executor.submit(generate_pivots, descriptors, n, strategy) for i in range(t)]
for pivot in concurrent.futures.as_completed(pivots):
new_pivot = pivot.result()
list_of_pivots.append(new_pivot)
return list_of_pivots
# + [markdown] id="IMBq1OIeyQ6a"
# ### Save test results
# + id="rklTlhWEyUzX" executionInfo={"status": "ok", "timestamp": 1643918476612, "user_tz": -60, "elapsed": 30, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def save_results(dir, file_name, results):
with open(os.path.join(dir, file_name +".csv"), 'w') as f:
writer = csv.writer(f)
# write the header
writer.writerow(["CLASS", "AP", "QUERY TIME"])
# write the data
for r in results:
writer.writerow(r)
# + [markdown] id="jHeezazzBiPh"
# ## Test Performance
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 22335, "status": "ok", "timestamp": 1643918501702, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}, "user_tz": -60} id="NnDcoPCV5ZCB" outputId="18600499-f80b-4814-e142-0f7c98767118"
drive.mount('/content/drive', force_remount=True)
# + [markdown] id="kGeWc472KezN"
# ### Create annot and load descriptors for the database
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="2LuXwWaQgffp" executionInfo={"status": "ok", "timestamp": 1643918659062, "user_tz": -60, "elapsed": 31688, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="1ae40ae5-639f-4c43-91fe-7b857afb4f57"
db_annot = concatenate_annots(['/content/drive/MyDrive/CV_Birds/train', '/content/drive/MyDrive/CV_Birds/mirflickr25k'])
db_annot
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643918660911, "user_tz": -60, "elapsed": 1866, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="71ec2ff6-dde7-473f-82c0-80e438586fd6" id="7L5mbCaDRA-6"
db_descriptors = concatenate_descriptors(['/content/drive/MyDrive/CV_Birds/features/training/AutoEncoder/512to128withPace64_feature_extraction.npy','/content/drive/MyDrive/CV_Birds/features/distractor/AutoEncoder/512to128withPace64_feature_extraction.npy'])
db_descriptors.shape
# + [markdown] id="CHoguzpHKm7F"
# ### Create annot and load descriptors for the test set
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="dMINWcaLGzMR" executionInfo={"status": "ok", "timestamp": 1643918678796, "user_tz": -60, "elapsed": 17896, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="3e7069e1-936b-4bd8-b868-55ad2f0f912e"
query_annot = create_annot('/content/drive/MyDrive/CV_Birds/test')
query_annot
# + colab={"base_uri": "https://localhost:8080/"} id="FewaIymhHA-y" executionInfo={"status": "ok", "timestamp": 1643918678798, "user_tz": -60, "elapsed": 32, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="2c1a9f0f-350d-426b-f195-ed9b69fce632"
query_descriptors = load_descriptors('/content/drive/MyDrive/CV_Birds/features/test/AutoEncoder/512to128withPace64_feature_extraction.npy')
query_descriptors.shape
# + [markdown] id="gzVn-4qOKrpY"
# To run our tests we select only the first image of each species within the test set. Please note that within the test set we have 5 images per species.
# + id="echKP4-dKsOV" executionInfo={"status": "ok", "timestamp": 1643918678799, "user_tz": -60, "elapsed": 23, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
queries_indexes = [x for x in range(325*5) if x%5 == 0]
# + [markdown] id="J1Di9pSXKsvz"
# ### Create PP-Index
# + id="PCf1g6xx9IcJ" executionInfo={"status": "ok", "timestamp": 1643918678800, "user_tz": -60, "elapsed": 19, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def get_descriptor_from_id(id_object):
return db_descriptors[id_object]
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2440, "status": "ok", "timestamp": 1643918681224, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}, "user_tz": -60} id="da0Jm02g6Ia4" outputId="2e140be8-81c7-4c29-f108-f96ed1302350"
# %cd "/content/drive/MyDrive/CV_Birds/Notebooks/PP-Index"
# %run PPIndex.ipynb
# + id="MYC5TDVOTqam" executionInfo={"status": "ok", "timestamp": 1643918681224, "user_tz": -60, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
# generate pivots
pivots = generate_pivots(db_descriptors, 40, "rnd")
# + id="iOgkJ5lgn_0-" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643918793931, "user_tz": -60, "elapsed": 79975, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="48d616d5-11bd-4107-d2e2-008716313dcc"
# cosine tree
cosine_tree = PrefixTree(pivots, length=3, distance_metric='cosine', base_directory="/content/cosine", tree_file='tree_structure')
if cosine_tree.is_empty():
cosine_tree.insert_objects_into_tree(range(len(db_descriptors)))
cosine_tree.save()
# + id="r_OvrPCaMjx1" executionInfo={"status": "ok", "timestamp": 1643920109926, "user_tz": -60, "elapsed": 103870, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
# !cp /content/cosine/tree* /content/drive/MyDrive/CV_Birds/indexes/feature_extraction/tree/cosine/
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643918851593, "user_tz": -60, "elapsed": 57684, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="53d22b1f-68f1-43a3-d3a7-bba731695bba" id="1kPIk2FASafI"
# euclidean tree
euclidean_tree = PrefixTree(pivots, length=3, distance_metric='euclidean', base_directory="/content/euclidean", tree_file='tree_structure')
if euclidean_tree.is_empty():
euclidean_tree.insert_objects_into_tree(range(len(db_descriptors)))
euclidean_tree.save()
# + id="vyD4ewAzM0io" executionInfo={"status": "ok", "timestamp": 1643921252893, "user_tz": -60, "elapsed": 80578, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
# !cp /content/euclidean/tree* /content/drive/MyDrive/CV_Birds/indexes/feature_extraction/tree/euclidean/
# + [markdown] id="cof_BX1EPvuM"
# ### Compute mAP
# + id="lB2JFYSZkey6" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643738205201, "user_tz": -60, "elapsed": 540, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "02615844185003710625"}} outputId="022c5ace-b8df-4710-a1ce-a45f3eddd22c"
birds_db = db_annot.loc[db_annot['identity'] != 'mirflickr']
counts = birds_db.groupby('identity').count()
print("Minimum number of images per species:", int(counts.min()))
print("Maximum number of images per species:", int(counts.max()))
print("Average number of images:", float(counts.sum()/325))
# + [markdown] id="do4DjuqJQPnG"
# Since at most we have 249 images per species, we use $n=250$.
# + id="XJhUa-cBQOXv" executionInfo={"status": "ok", "timestamp": 1643918851595, "user_tz": -60, "elapsed": 42, "user": {"displayName": "Federica Baldi", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
n = 250
# + [markdown] id="5MK_5J-dUhyB"
# The formula for Average Precision is the following:
#
# > $AP@n=\frac{1}{GTP}\sum_{k=1}^{n}P@k×rel@k$
#
# where $GTP$ refers to the total number of ground truth positives, $n$ refers to the total number of images we are interested in, $P@k$ refers to the precision@k and $rel@k$ is a relevance function.
#
# The relevance function is an indicator function which equals 1 if the document at rank $k$ is relevant and equals to 0 otherwise.
# + id="3ZXyZ9EPgwaY" executionInfo={"status": "ok", "timestamp": 1643918851596, "user_tz": -60, "elapsed": 41, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def compute_ap(query_index, retrieved_ids):
query_identity = query_annot['identity'][query_index]
print(query_index//5, query_identity)
GTP = len(db_annot.loc[db_annot['identity'] == query_identity])
relevant = 0
precision_summation = 0
for k, id in enumerate(retrieved_ids):
if db_annot['identity'][id] == query_identity: # relevant result
relevant = relevant + 1
precision_at_k = relevant/(k+1)
precision_summation = precision_summation + precision_at_k
return (query_identity, precision_summation/GTP)
# + [markdown] id="TBKg0G9aWuyH"
# For each query, $Q$, we can calculate a corresponding $AP$. Then, the $mAP$ is simply the mean of all the queries that were made.
# > $mAP = \frac{1}{N}\sum_{i=1}^{N}AP_i$
#
# In our case, $N=325$ (one query per species)
# + [markdown] id="H7uS1x7wYmqO"
# #### Simple tree
# + [markdown] id="ChtTD0irZhq0"
# ##### Cosine
# + id="2BZqRtvtiEnR" executionInfo={"status": "ok", "timestamp": 1643918851596, "user_tz": -60, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def cosine_tree_queries(query_index, n):
start_time = time.time()
ids, distances = cosine_tree.find_nearest_neighbors(query_descriptors[query_index], n)
end_time = time.time()
ids = ids.tolist()
return compute_ap(query_index, ids) + (end_time - start_time,)
# + id="o8bgk06_h_nV" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643919040240, "user_tz": -60, "elapsed": 188681, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="18cf8cf1-fb82-4247-a0e0-3800971ef760"
aps = []
for query_index in queries_indexes:
aps.append(cosine_tree_queries(query_index, n))
# + id="lHPcHDmlwSyr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643919040241, "user_tz": -60, "elapsed": 35, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="0fd2c930-34ad-44f0-aafa-a13592650dcf"
aps
# + id="8ALJRETfxOR1" executionInfo={"status": "ok", "timestamp": 1643919040242, "user_tz": -60, "elapsed": 26, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
ap_at_n = np.array([ap[1] for ap in aps])
query_time = np.array(([ap[2] for ap in aps]))
# + id="Yj6BWFaSxSj2" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643919040242, "user_tz": -60, "elapsed": 24, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="f43cac8d-41c5-4b9f-d733-cfce0795b0cc"
mAP_at_n = np.mean(ap_at_n, axis=0)
avg_query_time = np.mean(query_time, axis=0)
print("mAP:", mAP_at_n)
print("avg. query time: ", avg_query_time)
# + id="nU2Uhd37wayi" executionInfo={"status": "ok", "timestamp": 1643919041223, "user_tz": -60, "elapsed": 990, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
save_results('/content/drive/MyDrive/CV_Birds/performance/fine_tuning/index/AutoEncoder', 'AE_FE_tree_cosine_results', aps)
# + [markdown] id="bcy223BIZl2A"
# ##### Euclidean
# + id="FFi7wbguaHhf" executionInfo={"status": "ok", "timestamp": 1643919041227, "user_tz": -60, "elapsed": 23, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def euclidean_tree_queries(query_index, n):
start_time = time.time()
ids, distances = euclidean_tree.find_nearest_neighbors(query_descriptors[query_index], n)
end_time = time.time()
ids = ids.tolist()
return compute_ap(query_index, ids) + (end_time - start_time,)
# + colab={"base_uri": "https://localhost:8080/"} outputId="03577486-70c2-49fb-fb22-252cbe6edc1c" id="L9N7GK7mYzcG" executionInfo={"status": "ok", "timestamp": 1643919215709, "user_tz": -60, "elapsed": 174502, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
aps = []
for query_index in queries_indexes:
aps.append(euclidean_tree_queries(query_index, n))
# + colab={"base_uri": "https://localhost:8080/"} id="Rq8aEJNPZV1L" executionInfo={"status": "ok", "timestamp": 1643919215710, "user_tz": -60, "elapsed": 33, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="1c677de1-0b64-4ea0-d1a2-5037e91e0640"
aps
# + id="sS4uH1GrZZFi" executionInfo={"status": "ok", "timestamp": 1643919215711, "user_tz": -60, "elapsed": 28, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
ap_at_n = np.array([ap[1] for ap in aps])
query_time = np.array(([ap[2] for ap in aps]))
# + colab={"base_uri": "https://localhost:8080/"} id="gavW2smpZbFC" executionInfo={"status": "ok", "timestamp": 1643919215712, "user_tz": -60, "elapsed": 28, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="41fcca1d-5af7-4112-f347-ef02e6391174"
mAP_at_n = np.mean(ap_at_n, axis=0)
avg_query_time = np.mean(query_time, axis=0)
print("mAP:", mAP_at_n)
print("avg. query time: ", avg_query_time)
# + id="xS4uWooKZdsq" executionInfo={"status": "ok", "timestamp": 1643919215713, "user_tz": -60, "elapsed": 15, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
save_results('/content/drive/MyDrive/CV_Birds/performance/fine_tuning/index/AutoEncoder', 'AE_FE_tree_euclidean_results', aps)
# + [markdown] id="6iVsNzVuZ9hA"
# #### Tree with query perturbation
# + [markdown] id="w-aT-baNaCCK"
# ##### Cosine
# + id="BG2WAq1qaCCL" executionInfo={"status": "ok", "timestamp": 1643919217000, "user_tz": -60, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def cosine_pert_tree_queries(query_index, n):
start_time = time.time()
ids, distances = cosine_tree.find_nearest_neighbors_with_query_perturbation(query_descriptors[query_index], n, perturbations=3)
end_time = time.time()
ids = ids.tolist()
return compute_ap(query_index, ids) + (end_time - start_time,)
# + colab={"base_uri": "https://localhost:8080/"} outputId="94439998-3fcc-4a6e-887e-723db09e9457" id="1Shgyjw3aCCL" executionInfo={"status": "ok", "timestamp": 1643919615262, "user_tz": -60, "elapsed": 398270, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
aps = []
for query_index in queries_indexes:
aps.append(cosine_pert_tree_queries(query_index, n))
# + colab={"base_uri": "https://localhost:8080/"} id="_0jxgnl5aCCL" executionInfo={"status": "ok", "timestamp": 1643919615264, "user_tz": -60, "elapsed": 57, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="137511ec-d1c0-479a-d3e4-f35987d522a5"
aps
# + id="YCEPTeC1aCCM" executionInfo={"status": "ok", "timestamp": 1643919615265, "user_tz": -60, "elapsed": 46, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
ap_at_n = np.array([ap[1] for ap in aps])
query_time = np.array(([ap[2] for ap in aps]))
# + colab={"base_uri": "https://localhost:8080/"} id="Xqf8hvv5aCCM" executionInfo={"status": "ok", "timestamp": 1643919615267, "user_tz": -60, "elapsed": 46, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="039682df-c5e4-409e-80b7-8d4ddba58e72"
mAP_at_n = np.mean(ap_at_n, axis=0)
avg_query_time = np.mean(query_time, axis=0)
print("mAP:", mAP_at_n)
print("avg. query time: ", avg_query_time)
# + id="yDFA3FDlaCCM" executionInfo={"status": "ok", "timestamp": 1643919615268, "user_tz": -60, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
save_results('/content/drive/MyDrive/CV_Birds/performance/fine_tuning/index/AutoEncoder', 'AE_FE_pert_tree_cosine_results', aps)
# + [markdown] id="wQCJ9k0Ea7Gn"
# ##### Euclidean
# + id="FNRO6uePa7Gp" executionInfo={"status": "ok", "timestamp": 1643919615269, "user_tz": -60, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
def euclidean_pert_tree_queries(query_index, n):
start_time = time.time()
ids, distances = euclidean_tree.find_nearest_neighbors_with_query_perturbation(query_descriptors[query_index], n, perturbations=3)
end_time = time.time()
ids = ids.tolist()
return compute_ap(query_index, ids) + (end_time - start_time,)
# + colab={"base_uri": "https://localhost:8080/"} outputId="ef0e63d4-2a6a-4e9a-c1ec-64be6533cff0" id="9IQIAusfa7Gp" executionInfo={"status": "ok", "timestamp": 1643920006067, "user_tz": -60, "elapsed": 390818, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
aps = []
for query_index in queries_indexes:
aps.append(euclidean_pert_tree_queries(query_index, n))
# + id="ANdbSaWZa7Gq" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643920006068, "user_tz": -60, "elapsed": 31, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="9ad41d7b-95b6-48b3-8dd5-bfc72b7b5276"
aps
# + id="ptVx9KwXa7Gr" executionInfo={"status": "ok", "timestamp": 1643920006071, "user_tz": -60, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
ap_at_n = np.array([ap[1] for ap in aps])
query_time = np.array(([ap[2] for ap in aps]))
# + id="MStqeGtZa7Gr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643920006072, "user_tz": -60, "elapsed": 28, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}} outputId="d06b5b29-c0d2-4fb6-8f75-2a8c2aea9bef"
mAP_at_n = np.mean(ap_at_n, axis=0)
avg_query_time = np.mean(query_time, axis=0)
print("mAP:", mAP_at_n)
print("avg. query time: ", avg_query_time)
# + id="7lLXLauja7Gs" executionInfo={"status": "ok", "timestamp": 1643920006073, "user_tz": -60, "elapsed": 18, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0eALl9wD-GJPtSTMAFK0DvDlcGWh5ZTpdGFtwCIo=s64", "userId": "15726835528723494588"}}
save_results('/content/drive/MyDrive/CV_Birds/performance/fine_tuning/index/AutoEncoder', 'AE_FE_pert_tree_euclidean_results', aps)
| Notebooks/Performance-Evaluation/FE_AutoEncoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.insert(1, '../preprocessing/')
import numpy as np
import pickle
import scipy.stats as spstats
import matplotlib.pyplot as plt
#import seaborn as sns
#import pandas_profiling
from sklearn.utils import Bunch
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import cross_val_score, StratifiedShuffleSplit, StratifiedKFold
from sklearn.metrics import classification_report, f1_score, accuracy_score, make_scorer
import re
import pandas as pd
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
pd.set_option('display.max_colwidth', None)
from ordered_set import OrderedSet
from func_def import *
# %matplotlib inline
# +
# data from variable guide
data_variable_cat = {}
with open("data_variable_cat.pkl", "rb") as f:
data_variable_cat = pickle.load(f)
len(data_variable_cat)
# -
df_data_comp = pd.read_pickle(orginal_data_path+'Optima_Data_Report_Cases_9584_filled_pickle')
df_data_comp.sort_values(by=['GLOBAL_PATIENT_DB_ID', 'EPISODE_DATE'], inplace=True)
df_data_comp.head(1)
# +
# Analysis Recall Objects
# Rename these columns
df_data_comp.rename(columns={'COGNITIVE EXAM 120-161: (161) RECALLS OBJECTS':'COGNITIVE EXAM 120-161: (161) RECALLS OBJECTS_3',
'COGNITIVE EXAM 120-161: (146) RECALLS OBJECTS':'COGNITIVE EXAM 120-161: (146) RECALLS OBJECTS_6'}, inplace=True)
df_data_comp[['COGNITIVE EXAM 120-161: (161) RECALLS OBJECTS_3', 'COGNITIVE EXAM 120-161: (146) RECALLS OBJECTS_6']].hist()
# -
df_data_comp['durations(years)'] = df_data_comp.groupby(by='GLOBAL_PATIENT_DB_ID')['EPISODE_DATE'].transform(lambda x: (x - x.iloc[0])/(np.timedelta64(1, 'D')*365.25))
df_data_comp['MINI MENTAL SCORE PRE'] = df_data_comp.groupby(by='GLOBAL_PATIENT_DB_ID')['CAMDEX SCORES: MINI MENTAL SCORE'].transform(lambda x: x.shift(+1))
misdiagnosed_df = pd.read_csv(data_path+'misdiagnosed.csv')
display(misdiagnosed_df.head(5))
misdiagnosed_df['EPISODE_DATE'] = pd.to_datetime(misdiagnosed_df['EPISODE_DATE'])
# Rename GLOBAL_PATIENT_DB_ID to patient_id
df_data_comp.rename(columns={"GLOBAL_PATIENT_DB_ID": "patient_id"}, inplace=True)
# Merge With Misdiagnosed patients
df_data_comp= df_data_comp.merge(misdiagnosed_df[['patient_id', 'EPISODE_DATE', 'Misdiagnosed','Misdiagnosed1']], how='left', on=['patient_id', 'EPISODE_DATE'])
print (df_data_comp.shape)
display(df_data_comp.head(1))
df_data_comp['Misdiagnosed1'] = df_data_comp['Misdiagnosed1'].replace(['NO', 'YES', 'UNKNOWN'],[0, 1, 2])
df_data_comp['Misdiagnosed'] = df_data_comp['Misdiagnosed'].replace(['NO', 'YES', 'UNKNOWN'],[0, 1, 2])
# +
for i, j in zip(df_data_comp, df_data_comp.dtypes):
if not (j == "float64" or j == "int64" or j == 'uint8' or j == 'datetime64[ns]'):
print(i, j)
df_data_comp[i] = pd.to_numeric(df_data_comp[i], errors='coerce')
df_data_comp.shape
# -
df_data_comp = df_data_comp.replace([-1], [np.nan])
df_data_comp = df_data_comp[df_data_comp['Misdiagnosed1']<2]
df_data_comp = df_data_comp.astype({col: str('float64') for col, dtype in zip (df_data_comp.columns.tolist(), df_data_comp.dtypes.tolist()) if 'int' in str(dtype) or str(dtype)=='object'})
categorical_columns = [col for col in df_data_comp.columns if col in data_variable_cat.keys()]
for column in categorical_columns:
def replace_numerical_category(column, x):
if x in data_variable_cat[column]:
x = data_variable_cat[column][x]
else:
x = np.nan
return x
df_data_comp[column]=df_data_comp[column].apply(lambda x : replace_numerical_category(column, x))
# replace with Unlnown
df_data_comp[categorical_columns] = df_data_comp[categorical_columns].replace([np.nan], ['Unknown'])
df_data_comp[categorical_columns] = df_data_comp[categorical_columns].replace(['Not asked'], ['Unknown'])
df_data_comp[categorical_columns] = df_data_comp[categorical_columns].replace(['Not known'], ['Unknown'])
# +
def find_mixed_type_list(l):
for i in range(0,len(l)-1):
if type(l[i])!=type(l[i+1]):
return True
return False
list_corrupted_columns = []
for col in categorical_columns:
if find_mixed_type_list(df_data_comp[col].unique().tolist()):
list_corrupted_columns.append(col)
print (col,': ',df_data_comp[col].unique().tolist())
print(len(list_corrupted_columns))
# -
for col in list_corrupted_columns:
print (prepared_dataset.groupby(col)[col].count())
df_data_comp[categorical_columns] = df_data_comp[categorical_columns].replace(['Unknown'], [np.nan])
df_data_comp.shape
df_data_comp = df_data_comp.drop(columns=['patient_id', 'EPISODE_DATE', 'CAMDEX SCORES: MINI MENTAL SCORE', 'OPTIMA DIAGNOSES V 2010: PETERSEN MCI',
'Misdiagnosed', 'MINI MENTAL SCORE PRE', 'durations(years)', 'EPISODE'])
df_data_comp.shape
# Drop all features except MMSE features
columns_mmse = [col for col in df_data_comp.columns if 'COGNITIVE EXAM ' in col]
print (columns_mmse)
df_data_comp = df_data_comp[columns_mmse + ['Misdiagnosed1']]
print (df_data_comp.shape)
print (df_data_comp.columns)
df_data_comp_save = df_data_comp
df_data_comp = df_data_comp_save
# +
# Take only columns which are filled for 133 misdiagnosed patients almost
df_data_comp_X_misdiag = df_data_comp[df_data_comp['Misdiagnosed1']==1]
df_data_comp_X_misdiag = drop_missing_columns(df_data_comp_X_misdiag[df_data_comp_X_misdiag.isna().sum(axis=1)<25], 0.98) # thresold to decide about missing values 1506 in this case
print (df_data_comp_X_misdiag.shape)
df_data_comp = df_data_comp[df_data_comp_X_misdiag.columns]
df_data_comp.shape
# -
df_data_comp_save = df_data_comp
df_data_comp = df_data_comp_save
# +
df_data_comp = drop_missing_columns(df_data_comp[df_data_comp.isna().sum(axis=1)<4], 0.99)
print (df_data_comp[df_data_comp['Misdiagnosed1']==1].shape, df_data_comp[df_data_comp['Misdiagnosed1']==0].shape)
# +
# # feature transforamtion - one-hot encoding
prepared_dataset_exp = df_data_comp
# select categorical data columns
categorical_columns_final_exp = [col for col in prepared_dataset_exp.columns if col in categorical_columns]
new_prepared_data = prepared_dataset_exp.drop(categorical_columns_final_exp, axis=1)
for i in categorical_columns_final_exp:
x = pd.get_dummies(prepared_dataset_exp[i]).add_prefix(i+'::')
new_prepared_data = pd.concat([new_prepared_data, x], axis=1)
df_data_comp = new_prepared_data
print (df_data_comp.shape)
print(df_data_comp.columns.tolist())
# +
# rename NotAsked columns
# notasked_dict = {col:col.replace('::No', '::NO') for col in df_data_comp.columns if 'No' in col.split(sep='::')}
# print (notasked_dict)
# df_data_comp.rename(columns=notasked_dict, inplace=True)
# +
# drop Nagative Features # if there is only two values in columns only
# let it do later # for binary categroies
s1 = set([col.replace('::Incorrect', '') for col in df_data_comp.columns if 'Incorrect' in col.split('::')])-set([col.replace('::Correct', '') for col in df_data_comp.columns if 'Correct' in col.split('::')])
s2 = set([col.replace('::Yes', '') for col in df_data_comp.columns if 'Yes' in col.split('::')])-set([col.replace('::No', '') for col in df_data_comp.columns if 'No' in col.split('::')])
s3 = set([col.replace('::Correct', '') for col in df_data_comp.columns if 'Correct' in col.split('::')])-set([col.replace('::Incorrect', '') for col in df_data_comp.columns if 'Incorrect' in col.split('::')])
s4 = set([col.replace('::No', '') for col in df_data_comp.columns if 'No' in col.split('::')])-set([col.replace('::Yes', '') for col in df_data_comp.columns if 'Yes' in col.split('::')])
s = s1.union(s2).union(s3).union(s4)
s_list = list(s)
print (len(s_list))
# save df of s_list
exp_columns = [col for col in df_data_comp.columns if re.sub('::.*', '', col) in s_list and ('::No' in col or '::Incorrect' in col)]
print (exp_columns)
print (s_list)
# -
# drop Nagative Features # if there is only two values in columns only
df_data_comp = df_data_comp.drop(columns=[col for col in df_data_comp.columns if (('::Incorrect' in col or '::No' in col)) & (col not in exp_columns)])
print (df_data_comp.shape, df_data_comp.columns.tolist())
print (df_data_comp.shape)
df_data_comp = df_data_comp.dropna()
df_data_comp.shape
# drop duplicates
df_data_comp.drop_duplicates(inplace=True)
df_data_comp.shape
df_data_comp[df_data_comp['Misdiagnosed1']==0].shape, df_data_comp[df_data_comp['Misdiagnosed1']==1].shape
# +
# outlier detection
from sklearn.ensemble import IsolationForest
X = df_data_comp[df_data_comp['Misdiagnosed1']==0].drop(columns=['Misdiagnosed1'])
clf = IsolationForest(random_state=0).fit(X)
outlier_no_label = clf.predict(X)
from sklearn.ensemble import IsolationForest
X = df_data_comp[df_data_comp['Misdiagnosed1']==1].drop(columns=['Misdiagnosed1'])
clf = IsolationForest(random_state=0).fit(X)
outlier_yes_label = clf.predict(X)
# Original Size
print (sum(outlier_no_label)+ (len(outlier_no_label)-sum(outlier_no_label))/2)
print (sum(outlier_yes_label)+ (len(outlier_yes_label)-sum(outlier_yes_label))/2)
# -
df_data_comp['outlier_label'] = 0.0
df_data_comp.loc[df_data_comp['Misdiagnosed1']==0, 'outlier_label']=outlier_no_label
df_data_comp.loc[df_data_comp['Misdiagnosed1']==1, 'outlier_label']=outlier_yes_label
print (sum(df_data_comp['outlier_label']))
sum(df_data_comp[df_data_comp['Misdiagnosed1']==0]['outlier_label']), sum(df_data_comp[df_data_comp['Misdiagnosed1']==1]['outlier_label'])
# dron No outliers
df_X_y = df_data_comp[(df_data_comp['outlier_label']==1) | (df_data_comp['Misdiagnosed1']==1)]
df_X = df_X_y.drop(columns=['Misdiagnosed1', 'outlier_label'])
df_y = df_X_y['Misdiagnosed1']
print (df_X.shape, df_y.shape)
# +
X_full_imput, y_full_imput = df_X.values, df_y.values #X_full.values, y_full.values
# model training
rf_estimator = RandomForestClassifier(random_state=0)
cv = StratifiedShuffleSplit(n_splits=5, test_size=0.2, random_state=123)
important_features = set()
important_features_size = 40
for i, (train, test) in enumerate(cv.split(X_full_imput, y_full_imput)):
rf_estimator.fit(X_full_imput[train], y_full_imput[train])
y_predicted = rf_estimator.predict(X_full_imput[test])
print (classification_report(y_full_imput[test], y_predicted))
# print important features
# model important feature
fea_importance = rf_estimator.feature_importances_
indices = np.argsort(fea_importance)[::-1]
for f in range(important_features_size):
# print("%d. feature: %s (%f)" % (f + 1, X_full.columns.values[indices[f]], fea_importance[indices[f]]))
important_features.add(df_X.columns.values[indices[f]])
#lime interpretability
'''explainer = lime.lime_tabular.LimeTabularExplainer(np.array(X_full_imput[train]),
feature_names=[change_feature_names(fea) for fea in X_full.columns.values],
class_names= ['No Dementia', 'Dementia'],#rf_estimator.classes_,
discretize_continuous=True, random_state=123)
exp = explainer.explain_instance(X_full_imput[test][5], rf_estimator.predict_proba, num_features=10)
#exp.show_in_notebook(show_table=True, show_all=False)
exp.save_to_file('model_1DT_'+str(i)+'.html')'''
#print (exp.as_list())
#fig = exp.as_pyplot_figure()
#plt.show()
# shap interpretability
#important feature list
print ('important_features: ', list(important_features))
# -
df_X, df_y = df_X[list(important_features)], df_y
# +
# Random Forest Classfier
from sklearn.ensemble import RandomForestClassifier
from sklearn import svm, datasets
from sklearn.model_selection import cross_val_score, cross_validate, cross_val_predict
from sklearn.metrics import classification_report
import graphviz
from sklearn import tree
from six import StringIO
from IPython.display import Image
from sklearn.tree import export_graphviz
import pydotplus, joblib
from svglib.svglib import svg2rlg
from reportlab.graphics import renderPDF, renderPM
from sklearn.model_selection import train_test_split
import re
from dtreeviz.trees import *
# patient_df_X_fill_data[patient_df_y_cat==0]
X, y = df_X, df_y
clf = RandomForestClassifier(n_estimators=100)
print (cross_validate(clf, X, y, scoring=['recall_macro', 'precision_macro', 'f1_macro', 'accuracy'], cv=5) )
# y_pred = cross_val_predict(clf,X, y, cv=5 )
# print(classification_report(y, y_pred, target_names=['NO','YES']))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42, stratify=y)
feature_names = df_X.columns
clf = tree.DecisionTreeClassifier(max_depth=3, random_state=0).fit(X_train, y_train)
clf.score(X_test, y_test)
y_pred = clf.predict(X_test)
print (classification_report(y_test, y_pred))
'''dot_data = StringIO()
export_graphviz(clf, out_file=dot_data,
filled=True, rounded=True,
special_characters=True, feature_names=feature_names,
class_names=['NO', 'YES'])
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())'''
def change_feature_names(feature_name):
feature_name = feature_name.replace('_',' ')
p1 = '\w.*\d.*-\d.*:\s\(\d.*\w\)\s'
p2 = '\w.*:\s'
feature_name = re.sub(p1, '', feature_name)
# feature_name = re.sub(p2, '', feature_name)
for key, value in score_dict.items():
if feature_name in key:
feature_name = feature_name+'{}'.format(value)
return feature_name
bool_feature_names_DT = df_X.select_dtypes(include='uint8').columns
feature_names_DT = [change_feature_names(i) for i in feature_names]
bool_feature_names_DT = [change_feature_names(i) for i in bool_feature_names_DT] # Important 0: NO and 1: YES
bool_feature_names_true_DT = [i for i in bool_feature_names_DT if '::' in i] #('IDENTIFIES' in i or 'RECALL' in i) and '_1.0' in i ]
bool_feature_names_false_DT = [i for i in bool_feature_names_DT if '::' in i] #('IDENTIFIES' in i or 'RECALL' in i) and '_0.0' in i ]
feature_names_for_split_DT = [i for i in feature_names_DT if ' SCORE' in i]
viz = dtreeviz(clf,
x_data=X_train,
y_data=y_train,
target_name='class',
feature_names=feature_names_DT,
bool_feature_names_true=bool_feature_names_true_DT,
bool_feature_names_false=bool_feature_names_false_DT,
feature_names_for_split=feature_names_for_split_DT,
class_names=['misdiagnosed-No', 'misdiagnosed-Yes'],
fancy=False, label_fontsize=40, ticks_fontsize=2)
viz.save('original_dataset.svg')
drawing = svg2rlg("./original_dataset.svg".format(i))
renderPDF.drawToFile(drawing, "./original_dataset.pdf".format(i))
# +
from imblearn.over_sampling import SMOTE
smote = SMOTE(sampling_strategy='auto')
data_p_s, target_p_s = smote.fit_resample(df_X, df_y)
print (data_p_s.shape, target_p_s.shape)
# patient_df_X_fill_data[patient_df_y_cat==0]
X, y = data_p_s, target_p_s
clf = RandomForestClassifier(n_estimators=100)
print (cross_validate(clf, X, y, scoring=['recall_macro', 'precision_macro', 'f1_macro', 'accuracy'], cv=5) )
# y_pred = cross_val_predict(clf,X, y, cv=5 )
# print(classification_report(y, y_pred, target_names=['NO','YES']))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42, stratify=y)
feature_names = df_X.columns
clf = tree.DecisionTreeClassifier(max_depth=3, random_state=0).fit(X_train, y_train)
clf.score(X_test, y_test)
y_pred = clf.predict(X_test)
print (classification_report(y_test, y_pred))
'''dot_data = StringIO()
export_graphviz(clf, out_file=dot_data,
filled=True, rounded=True,
special_characters=True, feature_names=feature_names,
class_names=['NO', 'YES'])
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())'''
bool_feature_names_DT = df_X.select_dtypes(include='uint8').columns
feature_names_DT = [change_feature_names(i) for i in feature_names]
bool_feature_names_DT = [change_feature_names(i) for i in bool_feature_names_DT] # Important 0: NO and 1: YES
bool_feature_names_true_DT = [i for i in bool_feature_names_DT if '::' in i] #('IDENTIFIES' in i or 'RECALL' in i) and '_1.0' in i ]
bool_feature_names_false_DT = [i for i in bool_feature_names_DT if '::' in i] #('IDENTIFIES' in i or 'RECALL' in i) and '_0.0' in i ]
feature_names_for_split_DT = [i for i in feature_names_DT if ' SCORE' in i]
viz = dtreeviz(clf,
x_data=X_train,
y_data=y_train,
target_name='class',
feature_names=feature_names_DT,
bool_feature_names_true=bool_feature_names_true_DT,
bool_feature_names_false=bool_feature_names_false_DT,
feature_names_for_split=feature_names_for_split_DT,
class_names=['misdiagnosed-No', 'misdiagnosed-Yes'],
fancy=False, label_fontsize=40, ticks_fontsize=2)
viz.save('oversampled_smote.svg')
drawing = svg2rlg("./oversampled_smote.svg".format(i))
renderPDF.drawToFile(drawing, "./oversampled_smote.pdf".format(i))
# +
from collections import Counter
from imblearn.under_sampling import ClusterCentroids
cc = ClusterCentroids(random_state=0)
X_resampled, y_resampled = cc.fit_resample(df_X, df_y)
print(sorted(Counter(y_resampled).items()))
X, y = X_resampled, y_resampled
clf = RandomForestClassifier(n_estimators=100)
print (cross_validate(clf, X, y, scoring=['recall_macro', 'precision_macro', 'f1_macro', 'accuracy'], cv=5) )
# y_pred = cross_val_predict(clf,X, y, cv=5 )
# print(classification_report(y, y_pred, target_names=['NO','YES']))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42, stratify=y)
feature_names = df_X.columns
clf = tree.DecisionTreeClassifier(max_depth=3, random_state=0).fit(X_train, y_train)
clf.score(X_test, y_test)
y_pred = clf.predict(X_test)
print (classification_report(y_test, y_pred))
'''dot_data = StringIO()
export_graphviz(clf, out_file=dot_data,
filled=True, rounded=True,
special_characters=True, feature_names=feature_names,
class_names=['NO', 'YES'])
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())'''
bool_feature_names_DT = df_X.select_dtypes(include='uint8').columns
feature_names_DT = [change_feature_names(i) for i in feature_names]
bool_feature_names_DT = [change_feature_names(i) for i in bool_feature_names_DT] # Important 0: NO and 1: YES
bool_feature_names_true_DT = [i for i in bool_feature_names_DT if '::' in i] #('IDENTIFIES' in i or 'RECALL' in i) and '_1.0' in i ]
bool_feature_names_false_DT = [i for i in bool_feature_names_DT if '::' in i] #('IDENTIFIES' in i or 'RECALL' in i) and '_0.0' in i ]
feature_names_for_split_DT = [i for i in feature_names_DT if ' SCORE' in i]
viz = dtreeviz(clf,
x_data=X_train,
y_data=y_train,
target_name='class',
feature_names=feature_names_DT,
bool_feature_names_true=bool_feature_names_true_DT,
bool_feature_names_false=bool_feature_names_false_DT,
feature_names_for_split=feature_names_for_split_DT,
class_names=['misdiagnosed-No', 'misdiagnosed-Yes'],
fancy=False, label_fontsize=40, ticks_fontsize=2)
viz.save('undersampled_clustercentroid.svg')
drawing = svg2rlg("./undersampled_clustercentroid.svg".format(i))
renderPDF.drawToFile(drawing, "./undersampled_clustercentroid.pdf".format(i))
# +
from imblearn.under_sampling import RandomUnderSampler
rus = RandomUnderSampler(random_state=0)
X, y = rus.fit_resample(df_X, df_y)
clf = RandomForestClassifier(n_estimators=100)
print (cross_validate(clf, X, y, scoring=['recall_macro', 'precision_macro', 'f1_macro', 'accuracy'], cv=5) )
# y_pred = cross_val_predict(clf,X, y, cv=5 )
# print(classification_report(y, y_pred, target_names=['NO','YES']))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42, stratify=y)
feature_names = df_X.columns
clf = tree.DecisionTreeClassifier(max_depth=3, random_state=0).fit(X_train, y_train)
clf.score(X_test, y_test)
y_pred = clf.predict(X_test)
print (classification_report(y_test, y_pred))
'''dot_data = StringIO()
export_graphviz(clf, out_file=dot_data,
filled=True, rounded=True,
special_characters=True, feature_names=feature_names,
class_names=['NO', 'YES'])
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())'''
bool_feature_names_DT = df_X.select_dtypes(include='uint8').columns
feature_names_DT = [change_feature_names(i) for i in feature_names]
bool_feature_names_DT = [change_feature_names(i) for i in bool_feature_names_DT] # Important 0: NO and 1: YES
bool_feature_names_true_DT = [i for i in bool_feature_names_DT if '::' in i] #('IDENTIFIES' in i or 'RECALL' in i) and '_1.0' in i ]
bool_feature_names_false_DT = [i for i in bool_feature_names_DT if '::' in i] #('IDENTIFIES' in i or 'RECALL' in i) and '_0.0' in i ]
feature_names_for_split_DT = [i for i in feature_names_DT if ' SCORE' in i]
viz = dtreeviz(clf,
x_data=X_train,
y_data=y_train,
target_name='class',
feature_names=feature_names_DT,
bool_feature_names_true=bool_feature_names_true_DT,
bool_feature_names_false=bool_feature_names_false_DT,
feature_names_for_split=feature_names_for_split_DT,
class_names=['misdiagnosed-No', 'misdiagnosed-Yes'],
fancy=False, label_fontsize=40, ticks_fontsize=2)
viz.save('undersampled_random.svg')
drawing = svg2rlg("./undersampled_random.svg".format(i))
renderPDF.drawToFile(drawing, "./undersampled_random.pdf".format(i))
# -
| dementia_optima/diagnosed_pat_classification/mmse_variables_with_outliers/.ipynb_checkpoints/mmse_variable_with_outliers-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from typing import List
def any_index_equal_to_element(arr: List[int]) -> bool:
len_arr = len(arr)
if len_arr == 0:
return False
if len_arr == 1:
return arr[0] == 0
left_i = 0
right_i = len_arr - 1
while left_i <= right_i:
i = (left_i + right_i) // 2
element = arr[i]
if element != i:
left_i = i + 1
else:
return True
return False
# +
import unittest
class TestAreIndexEqualToElement(unittest.TestCase):
def test_empty(self):
self.assertFalse(any_index_equal_to_element([]))
def test_false(self):
test_cases = [
[1],
[1, 2, 3],
[4, 5, 6, 7]
]
for case in test_cases:
self.assertFalse(any_index_equal_to_element(case))
def test_true(self):
test_cases = [
[0],
[0, 1],
[1, 1, 1],
[0, 0, 2, 5],
[1, 2, 3, 3]
]
for case in test_cases:
self.assertTrue(any_index_equal_to_element(case))
def main():
test = TestAreIndexEqualToElement()
test.test_empty()
test.test_false()
test.test_true()
print("are_index_equal_to_element OK")
if __name__ == "__main__":
main()
# -
| chapter_3/problem_3_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 5.1 Data Prep
#
# ##### Description
#
# Prepping the data for modeling. Data will be split into train and test subsets.
#
# ##### Notebook Steps
#
# 1. Connect Spark
# 1. Input data
# 1. Basic data review
# 1. Visualize relationships
# ## 1. Connect Spark
# %load_ext sparkmagic.magics
# %manage_spark
# ## 2. Input Data
# + language="spark"
# df = spark.read.csv("s3://jolfr-capstone3/clean/features.csv", header=True, inferSchema=True)
# -
# ## 3. Split Data
# + language="spark"
# from pyspark.sql.functions import percent_rank
# from pyspark.sql import Window
# df = df.withColumn("rank", percent_rank().over(Window.partitionBy().orderBy("time")))
# + language="spark"
# train_df = df.where("rank <= .8").drop("rank")
# print((train_df.count(), len(train_df.columns)))
# + language="spark"
# test_df = df.where("rank > .8").drop("rank")
# print((test_df.count(), len(test_df.columns)))
# -
# ## Downsample Data
# Downsample data to speed in training.
# + language="spark"
# train_df, drop = train_df.randomSplit(weights = [0.05, 0.95], seed = 42)
# + language="spark"
# test_df, drop = test_df.randomSplit(weights = [0.05, 0.95], seed = 42)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 4. Drop Unnecessary Columns
# + language="spark"
# train_df = train_df.drop("msno").drop("time")
# + pycharm={"name": "#%%\n"} language="spark"
# test_df = test_df.drop("msno").drop("time")
# + language="spark"
# from pyspark.sql.types import FloatType
# from pyspark.sql.functions import col
# train_df = train_df.withColumn("label", col("label").cast(FloatType()))
# test_df = test_df.withColumn("label", col("label").cast(FloatType()))
# -
# ## 5. Output Data
# + pycharm={"name": "#%%\n"} language="spark"
# train_df.write.format("com.databricks.spark.csv").option("header", "true").mode('overwrite').save('s3://jolfr-capstone3/training/train.csv')
# + pycharm={"name": "#%%\n"} language="spark"
# test_df.write.format("com.databricks.spark.csv").option("header", "true").mode('overwrite').save('s3://jolfr-capstone3/validation/validate.csv')
# -
| notebooks/5-data_prep/5.1-data_prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ##### Copyright 2018 The TensorFlow Constrained Optimization Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
#
# > http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# ## Overview
#
# In this notebook, we explore the problem of classification with fairness on the [[UCI Adult dataset]](https://archive.ics.uci.edu/ml/datasets/adult). We show how to set up a classification problem with data-dependent fairness constraints using the TensorFlow Constrained Optimization library and then subsequently train to optimize fairness.
# +
import sys
sys.path.insert(0,'/Users/neelguha/Dropbox/NeelResearch/fairness/code/tensorflow_constrained_optimization/')
import math
import random
import numpy as np
import pandas as pd
import warnings
from six.moves import xrange
import tensorflow as tf
import tensorflow_constrained_optimization as tfco
import matplotlib.pyplot as plt
warnings.filterwarnings('ignore')
# %matplotlib inline
# -
# ### Reading and processing dataset.
#
# We load the [[UCI Adult dataset]](https://archive.ics.uci.edu/ml/datasets/adult) and do some pre-processing. The dataset is based on census data and the goal is to predict whether someone's income is over 50k. We construct four protected groups, two based on gender (Male and Female) and two based on race (White and Black).
#
# We preprocess the features as done in works such as [[ZafarEtAl15]](https://arxiv.org/abs/1507.05259) and [[GohEtAl16]](https://arxiv.org/abs/1606.07558). We transform the categorical features into binary ones and transform the continuous feature into buckets based on each feature's 5 quantiles values in training.
#
# The fairness goal is that of equal opportunity. That is, we would like the true positive rates of our classifier on the protected groups to match that of the overall dataset.
# +
CATEGORICAL_COLUMNS = [
'workclass', 'education', 'marital_status', 'occupation', 'relationship',
'race', 'gender', 'native_country'
]
CONTINUOUS_COLUMNS = [
'age', 'capital_gain', 'capital_loss', 'hours_per_week', 'education_num'
]
COLUMNS = [
'age', 'workclass', 'fnlwgt', 'education', 'education_num',
'marital_status', 'occupation', 'relationship', 'race', 'gender',
'capital_gain', 'capital_loss', 'hours_per_week', 'native_country',
'income_bracket'
]
LABEL_COLUMN = 'label'
PROTECTED_COLUMNS = [
'gender_Female', 'gender_Male', 'race_White', 'race_Black'
]
def get_data():
train_df_raw = pd.read_csv("https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data", names=COLUMNS, skipinitialspace=True)
test_df_raw = pd.read_csv("https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.test", names=COLUMNS, skipinitialspace=True, skiprows=1)
train_df_raw[LABEL_COLUMN] = (train_df_raw['income_bracket'].apply(lambda x: '>50K' in x)).astype(int)
test_df_raw[LABEL_COLUMN] = (test_df_raw['income_bracket'].apply(lambda x: '>50K' in x)).astype(int)
# Preprocessing Features
pd.options.mode.chained_assignment = None # default='warn'
# Functions for preprocessing categorical and continuous columns.
def binarize_categorical_columns(input_train_df, input_test_df, categorical_columns=[]):
def fix_columns(input_train_df, input_test_df):
test_df_missing_cols = set(input_train_df.columns) - set(input_test_df.columns)
for c in test_df_missing_cols:
input_test_df[c] = 0
train_df_missing_cols = set(input_test_df.columns) - set(input_train_df.columns)
for c in train_df_missing_cols:
input_train_df[c] = 0
input_train_df = input_train_df[input_test_df.columns]
return input_train_df, input_test_df
# Binarize categorical columns.
binarized_train_df = pd.get_dummies(input_train_df, columns=categorical_columns)
binarized_test_df = pd.get_dummies(input_test_df, columns=categorical_columns)
# Make sure the train and test dataframes have the same binarized columns.
fixed_train_df, fixed_test_df = fix_columns(binarized_train_df, binarized_test_df)
return fixed_train_df, fixed_test_df
def bucketize_continuous_column(input_train_df,
input_test_df,
continuous_column_name,
num_quantiles=None,
bins=None):
assert (num_quantiles is None or bins is None)
if num_quantiles is not None:
train_quantized, bins_quantized = pd.qcut(
input_train_df[continuous_column_name],
num_quantiles,
retbins=True,
labels=False)
input_train_df[continuous_column_name] = pd.cut(
input_train_df[continuous_column_name], bins_quantized, labels=False)
input_test_df[continuous_column_name] = pd.cut(
input_test_df[continuous_column_name], bins_quantized, labels=False)
elif bins is not None:
input_train_df[continuous_column_name] = pd.cut(
input_train_df[continuous_column_name], bins, labels=False)
input_test_df[continuous_column_name] = pd.cut(
input_test_df[continuous_column_name], bins, labels=False)
# Filter out all columns except the ones specified.
train_df = train_df_raw[CATEGORICAL_COLUMNS + CONTINUOUS_COLUMNS + [LABEL_COLUMN]]
test_df = test_df_raw[CATEGORICAL_COLUMNS + CONTINUOUS_COLUMNS + [LABEL_COLUMN]]
# Bucketize continuous columns.
bucketize_continuous_column(train_df, test_df, 'age', num_quantiles=4)
bucketize_continuous_column(train_df, test_df, 'capital_gain', bins=[-1, 1, 4000, 10000, 100000])
bucketize_continuous_column(train_df, test_df, 'capital_loss', bins=[-1, 1, 1800, 1950, 4500])
bucketize_continuous_column(train_df, test_df, 'hours_per_week', bins=[0, 39, 41, 50, 100])
bucketize_continuous_column(train_df, test_df, 'education_num', bins=[0, 8, 9, 11, 16])
train_df, test_df = binarize_categorical_columns(train_df, test_df, categorical_columns=CATEGORICAL_COLUMNS + CONTINUOUS_COLUMNS)
feature_names = list(train_df.keys())
feature_names.remove(LABEL_COLUMN)
num_features = len(feature_names)
return train_df, test_df, feature_names
train_df, test_df, FEATURE_NAMES = get_data()
# -
train_df.head()
# ### Model.
#
# We use a linear model and predict positively or negatively based on threshold at 0.
#
# In the following code, we initialize the placeholders and model. In build_train_op, we set up the constrained optimization problem. We create a rate context for the entire dataset, and compute the overall false positive rate as the positive prediction rate on the negatively labeled subset. We then construct a constraint for each of the protected groups based on the difference between the true positive rates of the protected group and that of the overall dataset. We then construct a minimization problem using RateMinimizationProblem and use the ProxyLagrangianOptimizer as the solver. build_train_op initializes a training operation which will later be used to actually train the model.
class Model(object):
def __init__(self,
tpr_max_diff=0):
tf.random.set_random_seed(123)
self.tpr_max_diff = tpr_max_diff
num_features = len(FEATURE_NAMES)
self.features_placeholder = tf.placeholder(
tf.float32, shape=(None, num_features), name='features_placeholder')
self.labels_placeholder = tf.placeholder(
tf.float32, shape=(None, 1), name='labels_placeholder')
self.protected_placeholders = [tf.placeholder(tf.float32, shape=(None, 1), name=attribute+"_placeholder") for attribute in PROTECTED_COLUMNS]
# We use a linear model.
self.predictions_tensor = tf.layers.dense(inputs=self.features_placeholder, units=1, activation=None)
def build_train_op(self,
learning_rate,
unconstrained=False):
ctx = tfco.rate_context(self.predictions_tensor, self.labels_placeholder)
positive_slice = ctx.subset(self.labels_placeholder > 0)
overall_tpr = tfco.positive_prediction_rate(positive_slice)
constraints = []
if not unconstrained:
for placeholder in self.protected_placeholders:
slice_tpr = tfco.positive_prediction_rate(ctx.subset((placeholder > 0) & (self.labels_placeholder > 0)))
constraints.append(slice_tpr >= overall_tpr - self.tpr_max_diff)
mp = tfco.RateMinimizationProblem(tfco.error_rate(ctx), constraints)
opt = tfco.ProxyLagrangianOptimizer(tf.train.AdamOptimizer(learning_rate))
self.train_op = opt.minimize(minimization_problem=mp)
return self.train_op
def feed_dict_helper(self, dataframe):
feed_dict = {self.features_placeholder:
dataframe[FEATURE_NAMES],
self.labels_placeholder:
dataframe[[LABEL_COLUMN]],}
for i, protected_attribute in enumerate(PROTECTED_COLUMNS):
feed_dict[self.protected_placeholders[i]] = dataframe[[protected_attribute]]
return feed_dict
# ### Training.
#
# Below is the function which performs the training of our constrained optimization problem. Each call to the function does one epoch through the dataset and then yields the training and testing predictions.
def training_generator(model,
train_df,
test_df,
minibatch_size,
num_iterations_per_loop=1,
num_loops=1):
random.seed(31337)
num_rows = train_df.shape[0]
minibatch_size = min(minibatch_size, num_rows)
permutation = list(range(train_df.shape[0]))
random.shuffle(permutation)
session = tf.Session()
session.run((tf.global_variables_initializer(),
tf.local_variables_initializer()))
minibatch_start_index = 0
for n in xrange(num_loops):
for _ in xrange(num_iterations_per_loop):
minibatch_indices = []
while len(minibatch_indices) < minibatch_size:
minibatch_end_index = (
minibatch_start_index + minibatch_size - len(minibatch_indices))
if minibatch_end_index >= num_rows:
minibatch_indices += range(minibatch_start_index, num_rows)
minibatch_start_index = 0
else:
minibatch_indices += range(minibatch_start_index, minibatch_end_index)
minibatch_start_index = minibatch_end_index
session.run(
model.train_op,
feed_dict=model.feed_dict_helper(
train_df.iloc[[permutation[ii] for ii in minibatch_indices]]))
train_predictions = session.run(
model.predictions_tensor,
feed_dict=model.feed_dict_helper(train_df))
session.run(
model.predictions_tensor,
feed_dict=model.feed_dict_helper(train_df))
session.run(
model.predictions_tensor,
feed_dict=model.feed_dict_helper(train_df))
test_predictions = session.run(
model.predictions_tensor,
feed_dict=model.feed_dict_helper(test_df))
yield (train_predictions, test_predictions)
# ### Computing accuracy and fairness metrics.
# +
def error_rate(predictions, labels):
signed_labels = (
(labels > 0).astype(np.float32) - (labels <= 0).astype(np.float32))
numerator = (np.multiply(signed_labels, predictions) <= 0).sum()
denominator = predictions.shape[0]
return float(numerator) / float(denominator)
def positive_prediction_rate(predictions, subset):
numerator = np.multiply((predictions > 0).astype(np.float32),
(subset > 0).astype(np.float32)).sum()
denominator = (subset > 0).sum()
return float(numerator) / float(denominator)
def tpr(df):
"""Measure the true positive rate."""
fp = sum((df['predictions'] >= 0.0) & (df[LABEL_COLUMN] > 0.5))
ln = sum(df[LABEL_COLUMN] > 0.5)
return float(fp) / float(ln)
def _get_error_rate_and_constraints(df, tpr_max_diff):
"""Computes the error and fairness violations."""
error_rate_local = error_rate(df[['predictions']], df[[LABEL_COLUMN]])
overall_tpr = tpr(df)
return error_rate_local, [(overall_tpr - tpr_max_diff) - tpr(df[df[protected_attribute] > 0.5]) for protected_attribute in PROTECTED_COLUMNS]
def _get_exp_error_rate_constraints(cand_dist, error_rates_vector, constraints_matrix):
"""Computes the expected error and fairness violations on a randomized solution."""
expected_error_rate = np.dot(cand_dist, error_rates_vector)
expected_constraints = np.matmul(cand_dist, constraints_matrix)
return expected_error_rate, expected_constraints
def training_helper(model,
train_df,
test_df,
minibatch_size,
num_iterations_per_loop=1,
num_loops=1):
train_error_rate_vector = []
train_constraints_matrix = []
test_error_rate_vector = []
test_constraints_matrix = []
for train, test in training_generator(
model, train_df, test_df, minibatch_size, num_iterations_per_loop,
num_loops):
train_df['predictions'] = train
test_df['predictions'] = test
train_error_rate, train_constraints = _get_error_rate_and_constraints(
train_df, model.tpr_max_diff)
train_error_rate_vector.append(train_error_rate)
train_constraints_matrix.append(train_constraints)
test_error_rate, test_constraints = _get_error_rate_and_constraints(
test_df, model.tpr_max_diff)
test_error_rate_vector.append(test_error_rate)
test_constraints_matrix.append(test_constraints)
return (train_error_rate_vector, train_constraints_matrix, test_error_rate_vector, test_constraints_matrix)
def get_tpr_subset(df, subsets):
filtered = df
for subset in subsets:
filtered = filtered[filtered[subset] > 0]
return tpr(filtered)
def get_acc_subset(df, subsets):
filtered = df
for subset in subsets:
filtered = filtered[filtered[subset] > 0]
predictions = filtered['predictions']
labels = filtered['label']
return np.mean(np.array(predictions > 0.0) == np.array(labels > 0.0))
# -
# ### Baseline without constraints.
#
# We now declare the model, build the training op, and then perform the training. We use a linear classifier, and train using the ADAM optimizer with learning rate 0.01, with minibatch size of 100 over 40 epochs. We first train without fairness constraints to show the baseline performance. We see that without training fair fairness, we obtain a high fairness violation.
#
# +
model = Model(tpr_max_diff=0.05)
model.build_train_op(0.01, unconstrained=True)
# training_helper returns the list of errors and violations over each epoch.
train_errors, train_violations, test_errors, test_violations = training_helper(
model,
train_df,
test_df,
100,
num_iterations_per_loop=326,
num_loops=40)
# -
print("Train Error", train_errors[-1])
print("Train Violation", max(train_violations[-1]))
print()
print("Test Error", test_errors[-1])
print("Test Violation", max(test_violations[-1]))
print("Baseline overall accuracy: %f" % (1 - error_rate(test_df['predictions'], test_df['label'])))
print("Baseline overall TPR: %f" % tpr(test_df))
subsets = [
['gender_Female'], ['gender_Male'], ['race_White'], ['race_Black'],
['gender_Female', 'race_White'],
['gender_Female', 'race_Black'],
['gender_Male', 'race_White'],
['gender_Male', 'race_Black']
]
for subset in subsets:
acc = get_acc_subset(test_df, subset)
print(subset, "Accuracy:", acc)
print()
for subset in subsets:
tpr_val = get_tpr_subset(test_df, subset)
print(subset, "TPR:", tpr_val)
# ### Training with fairness constraints.
#
# We now show train with the constraints using the procedure of [[CoJiSr19]](https://arxiv.org/abs/1804.06500) and returning the last solution found. We see that the fairness violation improves.
#
# We allow an additive fairness slack of 0.05. That is, when training and evaluating the fairness constraints, the true positive rate difference between protected group has to be at least that of the overall dataset up to a slack of at most 0.05. Thus, the fairness constraints would be of the form TPR_p >= TPR - 0.05, where TPR_p and TPR denotes the true positive rates of the protected group and the overall dataset, respectively.
#
# +
model = Model(tpr_max_diff=0.01)
model.build_train_op(0.01, unconstrained=False)
# training_helper returns the list of errors and violations over each epoch.
train_errors, train_violations, test_errors, test_violations = training_helper(
model,
train_df,
test_df,
100,
num_iterations_per_loop=326,
num_loops=40)
# -
print("Train Error", train_errors[-1])
print("Train Violation", max(train_violations[-1]))
print()
print("Test Error", test_errors[-1])
print("Test Violation", max(test_violations[-1]))
print("Baseline overall accuracy: %f" % (1 - error_rate(test_df['predictions'], test_df['label'])))
print("Baseline overall TPR: %f" % tpr(test_df))
subsets = [
['gender_Female'], ['gender_Male'], ['race_White'], ['race_Black'],
['gender_Female', 'race_White'],
['gender_Female', 'race_Black'],
['gender_Male', 'race_White'],
['gender_Male', 'race_Black']
]
for subset in subsets:
acc = get_acc_subset(test_df, subset)
print(subset, "Accuracy:", acc)
print()
for subset in subsets:
tpr_val = get_tpr_subset(test_df, subset)
print(subset, "TPR:", tpr_val)
# ### Improving using Best Iterate instead of Last Iterate.
#
# As discussed in [[CotterEtAl18b]](https://arxiv.org/abs/1809.04198), the last iterate may not be the best choice and suggests a simple heuristic to choose the best iterate out of the ones found after each epoch. The heuristic proceeds by ranking each of the solutions based on accuracy and fairness separately with respect to the training data. Any solutions which satisfy the constraints are equally ranked top in terms fairness. Each solution thus has two ranks. Then, the chosen solution is the one with the smallest maximum of the two ranks. We see that this improves the fairness and can find a better accuracy / fairness trade-off on the training data.
#
# This solution can be calculated using find_best_candidate_index given the list of training errors and violations associated with each of the epochs.
# +
best_cand_index = tfco.find_best_candidate_index(train_errors, train_violations)
print("Train Error", train_errors[best_cand_index])
print("Train Violation", max(train_violations[best_cand_index]))
print()
print("Test Error", test_errors[best_cand_index])
print("Test Violation", max(test_violations[best_cand_index]))
# -
# ### Using stochastic solutions.
#
# As discussed in [[CoJiSr19]](https://arxiv.org/abs/1804.06500), neither the best nor last iterate will come with theoretical guarantees. One can instead use randomized solutions, which come with theoretical guarantees. However, as discussed in [[CotterEtAl18b]](https://arxiv.org/abs/1809.04198), there may not always be a clear practical benefits. We show how to use these solutions here for sake of completeness.
# #### T-stochastic solution.
# The first and simplest randomized solution suggested is the T-stochastic, which simply takes the average of all of the iterates found at each epoch.
print("Train Error", np.mean(train_errors))
print("Train Violation", max(np.mean(train_violations, axis=0)))
print()
print("Test Error", np.mean(test_errors))
print("Test Violation", max(np.mean(test_violations, axis=0)))
# #### m-stochastic solution.
# [[CoJiSr19]](https://arxiv.org/abs/1804.06500) presents a method which shrinks down the T-stochastic solution down to one that is supported on at most (m+1) points where m is the number of constraints and is guaranteed to be at least as good as the T-stochastic solution. Here we see that indeed there is benefit in performing the shrinking.
#
# This solution can be computed using find_best_candidate_distribution by passing in the training errors and violations found at each epoch and returns the weight of each constituent. We see that indeed, it is sparse.
cand_dist = tfco.find_best_candidate_distribution(train_errors, train_violations)
print(cand_dist)
# +
m_stoch_error_train, m_stoch_violations_train = _get_exp_error_rate_constraints(cand_dist, train_errors, train_violations)
m_stoch_error_test, m_stoch_violations_test = _get_exp_error_rate_constraints(cand_dist, test_errors, test_violations)
print("Train Error", m_stoch_error_train)
print("Train Violation", max(m_stoch_violations_train))
print()
print("Test Error", m_stoch_error_test)
print("Test Violation", max(m_stoch_violations_test))
| examples/jupyter_notebooks/Fairness_ipums.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import modules
# +
#camera modules
import picamera as pc
from picamera.array import PiRGBArray
#thread management
from multiprocessing import Queue
from threading import Thread,Lock
from collections import deque
#image manipulation
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
#general modules
import time,datetime
import pickle
import sys
#saving images and vision analysis
import io
import os
from PIL import Image
#uploading to google cloud storage
from gcloud import storage
from oauth2client.service_account import ServiceAccountCredentials
# -
# # Initialize Picamera and Capture Frames from Video Stream
class Video_Camera(Thread):
""" Initializes PiCamera and captures frames from the video stream.
Functions:
__init__(self,fps,width,height,vflip,hflip,mins): Initializes
the camera and video stream using parameters including
fps (int): The amount of Frames captured Per Second by camera
width (int): Width of image to be captured in pixels
height (int): Height of image to be captured in pixels
vflip (boolean): Used to flip the image vertically
hflip (boolean): Used to flip the image horizontally
mins (int): Number of minutes of video to store in memory.
Used to minimize memory footprint.
initialize_camera(self): Starts camera class
initialize_video_stream(self): Starts color (RGB) video stream
apply_camera_day_settings(self): Alters camera properties to day settings
apply_camera_night_settings(self): Alters camera properties to night settings
run(self): Starts thread, Initializes PiCamera and captures frames
from the video stream
Args:
msg (Thread): A thread is an execution context, which is the
information a CPU needs to execute a stream of instructions.
"""
def __init__(self,fps,width,height,vflip,hflip,mins):
self.fps=fps
self.width=width
self.height=height
self.vflip=vflip
self.hflip=hflip
self.mins=mins
#Deque storage data structures
#Set max length of X minutes to prevent memory errors
self.input_deque=deque(maxlen=fps*mins*60)
#start the thread, deamon and kill switch
super(Video_Camera, self).__init__()
self.daemon = True
self.kill_all_threads= False
print (self, 'created')
#Initialize camera and video stream
self.initialize_camera()
self.initialize_video_stream()
self.person_present=False
print ('Camera and video stream initialized')
def initialize_camera(self):
self.camera = pc.PiCamera(
resolution=(self.width,self.height),
framerate=int(self.fps))
#Set camera properties
self.camera.vflip = self.vflip
self.camera.hflip = self.hflip
self.apply_camera_day_settings()
def initialize_video_stream(self):
self.rawCapture = pc.array.PiRGBArray(self.camera, size=self.camera.resolution)
self.stream = self.camera.capture_continuous(self.rawCapture,
format="bgr",
use_video_port=True)
def apply_camera_day_settings(self):
self.camera.exposure_mode = 'auto'
self.camera.contrast=0
self.camera.brightness=50
self.camera.exposure_compensation=0
def apply_camera_night_settings(self):
self.camera.exposure_mode = 'auto'
self.camera.contrast=20
self.camera.brightness=90
self.camera.exposure_compensation=6
def run(self):
#This method is run when the command start() is given to the thread
print ('Video stream is now being captured')
for f in self.stream:
#add frame with timestamp to input queue
self.input_deque.append({
'time':time.time(),
'frame_raw':f.array})
#remove the frame from the stream
self.rawCapture.truncate(0)
if self.kill_all_threads==True:
print (self, 'terminated')
break
# ### FPS must be over 1, and an integer
#Initialize Video_Camera Thread
video_camera = Video_Camera(fps=2,
width=384,
height=224,
vflip=True,
hflip=False,
mins=1)
#Begin capturing raw video and store data in the input_queue
video_camera.start()
# +
class Frame_Writer(Thread):
"""
Running process to write to cloud storage
"""
def __init__(self, Video_Camera, project, bucket_name, base_filename, local_filename):
self.Video_Camera=Video_Camera
self.project=project
self.bucket_name=bucket_name
self.base_filename=base_filename
self.local_filename=local_filename
# for writing to could storage
client = storage.Client(self.project)
self.bucket = client.get_bucket(self.bucket_name)
# thread init
super(Frame_Writer, self).__init__()
self.daemon = True
self.kill_all_threads= False
print (self, 'created')
def write_to_storage(self,output_name,processed_image):
# declare end name in the bucket
blob = self.bucket.blob(self.base_filename+output_name)
# upload from the image itself
im = Image.fromarray(processed_image)
imageBuffer = io.BytesIO()
im.save(imageBuffer, 'JPEG')
imageBuffer.seek(0)
blob.upload_from_file(file_obj=imageBuffer,size=imageBuffer.getbuffer().nbytes)
# close out this memory
imageBuffer.close()
def run(self):
while self.kill_all_threads!=True:
#This method is run when the command start() is given to the thread
raw_image = None
count = 0
# sample from the left (oldest) side of the deque by removing it
try:
oldest_deque_entry = self.Video_Camera.input_deque.popleft()
raw_frame = oldest_deque_entry['frame_raw']
processed_image = np.flipud(raw_frame[:, :, ::-1])
# push into cloud storage
output_name=time.strftime('%Y_%m_%d-%H_%M_%S',time.localtime(oldest_deque_entry['time']))
self.write_to_storage(output_name,processed_image)
print ('Processed image sent to storage: %S',output_name)
print ('Current time: %s',time.strftime('%Y_%m_%d-%H_%M_%S',time.localtime(time.time())))
except IndexError:
print ('No images available to consume from the queue')
#sleep if there is nothing to pull
time.sleep((self.Video_Camera.fps)/2.0)
# -
#Initialize Frame_Writer Thread
frame_writer = Frame_Writer(
Video_Camera=video_camera,
project='loyal-order-204316',
bucket_name='pi_images_iotworldhackathon',
base_filename='pi_test_v0_',
local_filename='temp_image.jpg')
#Begin capturing raw video and store data in the input_queue
frame_writer.start()
# ### Debugging Functions
def visualize_recent_frame(video_camera):
try:
# sample from the right side of the deque by removing it
newest_deque_entry=video_camera.input_deque.pop()
# add this frame back to the deque
video_camera.input_deque.append(newest_deque_entry)
#process the frame
raw_frame=newest_deque_entry['frame_raw']
processed_frame=(np.flipud(raw_frame[:, :, ::-1]))
plt.imshow(processed_frame)
except IndexError:
print("Cannot pop from an empty deque - let deque fill first")
# ### Visualize most recent frame (If available on input queue)
visualize_recent_frame(video_camera)
| notebooks/PiCamera - Video Processing To Cloud Storage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <p style="text-align: center;">Grouping Similar Problems</p>
# In this example, we are going to group customer support problems into problem groups based on their attributes. Then we will analyze the groups to see similarities and differences.
# ## Loading the Dataset
# +
# %matplotlib inline
from pandas import Series, DataFrame
import pandas as pd
import numpy as np
import os
import matplotlib.pylab as plt
from sklearn.model_selection import train_test_split
from sklearn.cluster import KMeans
import sklearn.metrics
raw_data = pd.read_csv("issues.csv")
raw_data.dtypes
# -
# The dataset contains one record for each unique problem type. It has metrics for each type like count, average calls to resolve, average resolution time etc.
raw_data.head()
# ## Group Data into similar clusters
#
# Now, we will use K-Means clustering to group data based on their attribute. First, we need to determine the optimal number of groups. For that we conduct the knee test to see where the knee happens.
# +
clust_data = raw_data.drop("PROBLEM_TYPE",axis=1)
#Finding optimal no. of clusters
from scipy.spatial.distance import cdist
clusters=range(1,10)
meanDistortions=[]
for k in clusters:
model=KMeans(n_clusters=k)
model.fit(clust_data)
prediction=model.predict(clust_data)
meanDistortions.append(sum(np.min(cdist(clust_data, model.cluster_centers_, 'euclidean'), axis=1)) / clust_data.shape[0])
#plt.cla()
plt.plot(clusters, meanDistortions, 'bx-')
plt.xlabel('k')
plt.ylabel('Average distortion')
plt.title('Selecting k with the Elbow Method')
# -
# Looking at the plot, we see that the knee happens at cluster=3. That is the ideal number of clusters. We now perform the actual clustering for 3. Then we add the cluster ID to the original dataset.
# +
#Optimal clusters is 3
final_model=KMeans(3)
final_model.fit(clust_data)
prediction=final_model.predict(clust_data)
#Join predicted clusters back to raw data
raw_data["GROUP"] = prediction
print("Groups Assigned : \n")
raw_data[["GROUP","PROBLEM_TYPE"]]
# -
# ## Analyze the groups
#
# We now do a set of boxplots to see how the groups differ for various feature attributes.
#
# We start off with Count.
plt.cla()
plt.boxplot([[raw_data["COUNT"][raw_data.GROUP==0]],
[raw_data["COUNT"][raw_data.GROUP==1]] ,
[raw_data["COUNT"][raw_data.GROUP==2]] ],
labels=('GROUP 1','GROUP 2','GROUP 3'))
# +
We can see that the count of incidents range differently for different groups.
Next we see avg. calls to resolve.
# -
#Now for Avg. Calls to resolve
plt.cla()
plt.boxplot([[raw_data["AVG_CALLS_TO_RESOLVE"][raw_data.GROUP==0]],
[raw_data["AVG_CALLS_TO_RESOLVE"][raw_data.GROUP==1]] ,
[raw_data["AVG_CALLS_TO_RESOLVE"][raw_data.GROUP==2]] ],
labels=('GROUP 1','GROUP 2','GROUP 3'))
# Group 2 has hardly any time needed to resolve. This points to problems that are simple and straight forward. The business need to look at these incidents and provide a self-service path (product help, online help) for the customer instead of wasting agent's time
#
# Next we see Reoccurance Rate.
plt.cla()
plt.boxplot([[raw_data["REOCCUR_RATE"][raw_data.GROUP==0]],
[raw_data["REOCCUR_RATE"][raw_data.GROUP==1]] ,
[raw_data["REOCCUR_RATE"][raw_data.GROUP==2]] ],
labels=('GROUP 1','GROUP 2','GROUP 3'))
# Group 2 has really high reoccurance rate. This set of incidents need to be analyze to see how the product quality can be improved to prevent these from happening.
plt.cla()
plt.boxplot([[raw_data["REPLACEMENT_RATE"][raw_data.GROUP==0]],
[raw_data["REPLACEMENT_RATE"][raw_data.GROUP==1]] ,
[raw_data["REPLACEMENT_RATE"][raw_data.GROUP==2]] ],
labels=('GROUP 1','GROUP 2','GROUP 3'))
# Replacement rates vary widely for Group 1. It does not provide any significant pattern to act upon.
| Files_Predictive_Customer_Analytics/Notebooks/05_05/.ipynb_checkpoints/grouping-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Data Science Process
# In this lesson, you’ll learn about the process used for many data science problems. This will serve you as a useful guide on how to approach future data science problems as well as your first project. If some of this material feels like review, feel free to zip ahead to only the required parts. If you want a deeper understanding of any of the areas introduced in this lesson, sit tight because we’ll be doing just that later on on this program.
#
# ## CRISP-DM
# Given the large number of industries and problems that fall under the umbrella of Data Science, there are a lot of different tools and individual nuances of a particular company of industry for finding solutions.
#
# However, there’s actually a common process used to find many solutions in Data Science. This is known as the Cross Industry Standard Process for Data Mining of CRISP-DM. This process has been an industry standard for analyzing data for years and it has six major phases. First, developing business understanding. Second, developing data understanding. Third, prepping your data to be analyzed. Fourth, modeling the data. Fifth, evaluating the results to answer your questions of interest and finally, deploying changes based on the results of your analysis.
#
# We will look at each of these phases a bit closer in upcoming lessons.
#
# The Data Science Process - Business & Data Understanding
# In the previous lesson, you received a quick look at the six steps of the CRISP-DM process. Let’s take a look at the first two steps of this process in a bit more detail.
#
# First, CRISP-DM says your need business understanding, but I think of data science as being more general than just business context, you need to understanding the problem. Are you interested in acquiring new customers? Are you interested in assessing of a new cancer treatment outperforms existing treatments? Are you interested in finding a better way to communicate, or travel, or retain information? Each of these questions falls under business understanding.
#
# The second step of the CRISP-DM process is data understanding. This means you need to gain an understanding of the data necessary to answer your question. Sometimes you might have a mountain of data at your disposal that you need to dig through to find insights. Other times you may need to collect data, which means you will have to understand what kind of data we’ll be able to provide you with the insights you need. This is often difficult to know ahead of time, which is why businesses tend to collect all the data they can first, so they can later identify which data they need to use to find their insights.
#
# The Data Science Process - Business & Data Understanding Example
# For this lesson, you will get hands-on practice with a dataset from Stack Overflow developer survey results, from 2017. The data for this example is available in the Resources tab for you to follow along on your machine, as well in the workspaces in the classroom. The data was collected by providing a survey to developers on Stack Overflow’s website. A quick look at the columns in the dataset provides a number of questions that we might ask.
#
# This dataset, can provide some insight and to developers around the world, and get you an idea of their experiences. Anything from their advice to other developers, to how they learn new skills, to where they live, or what programming languages they use, can all be understood from this dataset. You will use the workspace and quizzes in the following sections to familiarize yourself with the data, and we will answer a few questions that I found interesting in my analysis and explore questions that you are most interested in.
#
# ## The Data Science Process - Business & Data Understanding Example
#
#
# Working with Categorical Varibles
# One of the most common methods for encoding categorical variables is with ones and zeros. There are advantages of this method. With linear models, you have the ability to easily interpret the weights on each of these values, and it provides a lot of flexibility in determining how each level of the categorical variable influences the response.
#
# On the downside, when a categorical variable has lots of levels, creating dummy variables encoded in this way, can add a lot of new columns into your dataset. In this dataset, we’re working with 42,00 additional columns. If the number of columns were to exceed the number of rows, many machine learning algorithms will be unable to optimize for a solution at all.
#
# There are some additional techniques shown in the post and the links below
#
# Let’s try implementing this technique to see how it works out.
#
# ### A Look at the Data
#
# In order to get a better understanding of the data we will be looking at throughout this lesson, let's take a look at some of the characteristics of the dataset.
#
# First, let's read in the data and necessary libraries.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.core.display import display, HTML
# %matplotlib inline
df = pd.read_csv('./survey_results_public.csv')
df.head()
# -
# As you work through the notebook(s) in this and future parts of this program, you will see some consistency in how to test your solutions to assure they match what we achieved! In every environment, there is a solution file and a test file. There will be checks for each solution built into each notebook, but if you get stuck, you may also open the solution notebook to see how we find any of the solutions. Let's take a look at an example.
#
# ### Question 1
#
# **1.** Provide the number of rows and columns in this dataset.
# +
#This is how you see the number of rows and columns in a dataset.
shape_touple = df.shape
print(f'shape_touple => {shape_touple}')
number_of_rows = shape_touple[0]
print(f'number_of_rows => {number_of_rows:,.0F}')
number_of_columns = shape_touple[1]
print(f'number_of_columns => {number_of_columns:,.0F}')
# -
# ### Question 2
#
# **2.** Which columns had no missing values? Provide a set of column names that have no missing values.
# +
columns_wo_missing_values = set(df.columns[df.isnull().mean() == 0])
print('The columns that had no missing values are:')
display(HTML('<ol>'))
for col_name in columns_wo_missing_values:
display(HTML(f'<li>{col_name}</li>'))
display(HTML('</ol>'))
# -
# ### Question 3
#
# **3.** Which columns have the most missing values? Provide a set of column names that have more than 75% if their values missing.
# +
columns_missing_over_sevetyfive_percent = set(df.columns[df.isnull().mean() > .75])
print('The columns that have more than 75% of their values missing:')
display(HTML('<ol>'))
for col_name in columns_missing_over_sevetyfive_percent:
display(HTML(f'<li>{col_name}</li>'))
display(HTML('</ol>'))
# -
# ### Question 4
#
# **4.** Provide a pandas series of the different **Professional** status values in the dataset along with the count of the number of individuals with each status. Store this pandas series in **status_vals**. If you are correct, you should see a bar chart of the proportion of individuals in each status.
# +
status_vals = df['Professional'].value_counts()
print(status_vals)
(status_vals/df.shape[0]).plot(kind="bar")
plt.title("What kind of developer are you?")
# -
# ### Question 5
#
# **5.** Provide a pandas series of the different **FormalEducation** status values in the dataset along with the count of how many individuals received that formal education. Store this pandas series in **ed_vals**. If you are correct, you should see a bar chart of the proportion of individuals in each status.
# +
ed_vals = df['FormalEducation'].value_counts()
print(ed_vals)
(ed_vals/df.shape[0]).plot(kind='bar')
plt.title('What kind of Formal Education You Have?')
# -
# ### Question 6
#
# **6.** Provide a pandas series of the different **Country** values in the dataset along with the count of how many individuals are from each country. Store this pandas series in **count_vals**. If you are correct, you should see a bar chart of the proportion of individuals in each country.
# +
count_vals = df['Country'].value_counts()
print(count_vals)
(count_vals[:20]/df.shape[0]).plot(kind='bar')
plt.title('What Country are You From?')
# -
| lessons/CRISP_DM/TheDataScienceProcess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Bounding-Box Detection
# In this tutorial, we show you how to start with an image bounding box detection project. Following are the steps in this tutorial; feel free to jump ahead some steps if you've already done some previous tutorials:
#
# 1. [What is object detection?](#intro)
# 2. [Connection to Kili](#connect)
# 3. [Creating the project and setting up the interface](#project)
# 4. [Importing data](#data)
# 5. [Labeling](#labeling)
# 6. [Exporting labels](#export)
# 7. [Quality management](#quality)
# 8. [More advanced concepts](#concepts)
# # What is object detection<a id='intro'></a>
# The task of object detection consists of detecting instances of objects in an asset. In this tutorial, we're interested in image object detection. In concrete terms, this means detecting the presence of objects of different categories within images. Each category is composed of objects that all have common features. For example, we’ll consider the task of image object detection for the categories *Circle* and *Square*. We'll then try to identify these objects in our images, knowing that circles are round and squares have four edges.
#
# In image object detection, our goal is not only to identify the presence of given objects, but to identify their positions and dimensions. This can be achieved in an approximate way by using bounding boxes. A bounding box is defined as the smallest rectangle that completely covers an object in a given image. By using bounding boxes, we can get information about the position of an object (the center of the box) as well as its dimensions. The main advantage of using bounding boxes is that their representation is very simple (just the position of four lines).
#
# In the image below, you'll find an example of Kili's interface for image object detection for an image of a car. We use a single bounding box to represent a car.
#
# 
#
# Using image object detection can be very helpful in domains such as facial recognition, medicine, etc. It's especially interesting to be able to automate this process; however, to achieve good performance, it's generally necessary to have vast amounts of labeled data. A great way to label your data efficiently is using Kili. The next steps of this tutorial will walk you through the creation of your first image object detection project at Kili.
# # Connecting to Kili <a id='connect'></a>
# The first step is to be able to connect to the platform.
#
# If you use the SaaS version of Kili (see [here](https://cloud.kili-technology.com/docs/hosting/saas/)), you use by default the Auth0 login identification, or your company's authentication if it has been implemented.
#
# <img src="../img/auth0.png" width="400" />
#
# If you use Kili on-premise (see [here](https://cloud.kili-technology.com/docs/hosting/on-premise-entreprise/)), you will probably use our own authentication :
#
# <img src="../img/noauth0.png" width="400" />
#
# You need your organization admin to create your profile, and depending on the authentication implementation, you can sign up and set your password, or use the temporary one provided to you by the admin.
#
# If everything succeeds, you should arrive at the Projects page shown in the beginning of the next section.
# # Creating the project <a id='project'></a>
# ## List of projects
# You'll arrive on a list of projects.
#
# 
#
# You can refer to this [document](https://cloud.kili-technology.com/docs/concepts/definitions/) to find the definitions of key concepts at Kili. One of them is a project, which is a combination of:
# - a dataset (a list of assets)
# - members (project users; each can have different roles)
# - an interface (describing the annotation plan)
# ## Create the project
# You can either create a project [from the interface](https://cloud.kili-technology.com/docs/projects/new-project/#docsNav) or [from the API](https://github.com/kili-technology/kili-playground/blob/master/recipes/create_project.ipynb).
#
# To create a project from the interface, select `Create New` from the list of projects. and a description, then select `Image Object Detection (bounding box)`. Finally, clic `Save` as shown below:
#
# 
# <details>
# <summary style="display: list-item;"> Follow these instructions to create a project from the API </summary>
#
# From the API, you can create a project with a single call, which allows you to store and share project interfaces:
# - First, [connect to Kili](https://github.com/kili-technology/kili-playground/blob/master/README.md#get-started)
#
#
# ```python
# # Authentication
# import os
#
# # # !pip install kili # uncomment if you don't have kili installed already
# from kili.client import Kili
#
# api_key = os.getenv('KILI_USER_API_KEY')
# api_endpoint = os.getenv('KILI_API_ENDPOINT') # If you use Kili SaaS, use the url 'https://cloud.kili-technology.com/api/label/v2/graphql'
#
# kili = Kili(api_key=api_key, api_endpoint=api_endpoint)
# ```
#
# - Then call the method `create_project`: <a id='command'></a>
# ```python
# kili.create_project(
# title='Project Title',
# description='Project Description',
# input_type='IMAGE',
# json_interface=interface
# )
# ```
#
# with `interface` such as:
#
#
# ```python
# interface = {
# "jobRendererWidth": 0.17,
# "jobs": {
# "JOB_0": {
# "mlTask": "OBJECT_DETECTION",
# "tools": [
# "rectangle"
# ],
# "instruction": "Categories",
# "required": 1,
# "isChild": False,
# "isVisible": True,
# "content": {
# "categories": {
# "OBJECT_A": {
# "name": "Object A",
# "children": [],
# "color": "#0755FF"
# },
# "OBJECT_B": {
# "name": "Object B",
# "children": [],
# "color": "#EEBA00"
# }
# },
# "input": "radio"
# }
# }
# }
# }
# ```
#
#
# ```python
# result = kili.create_project(
# title='Project Title',
# description='Project Description',
# input_type='IMAGE',
# json_interface=interface
# )
# print(result)
# ```
#
# ```python
# Out: {'id': 'ckm4pmqmk0000d49k6ewu2um5'}
# ```
# </details>
# ## Access your project
# This creates a project with a simple interface, a radio button, and two categories: `Object A` and `Object B`. Once logged in, you can see your project in the list of projects:
#
# 
#
# If you want to modify or view the interface, go to the Settings tab. First, click on the Settings button in the sidebar.
#
# <img src="../img/sidebar_settings.png" width=100/>
#
# You can find both the form and the JSON versions of the interface:
#
# 
#
# [Find out how to modify the interface dynamically](https://cloud.kili-technology.com/docs/projects/customize-interface/#docsNav).
#
# If you want to go back to the list of projects, you can either click on `Kili Technology` in the top bar, or on the list of projects in the sidebar:
#
# <img src="../img/sidebar_listprojects.png" width=100>
# <details class="mydetails">
# <summary style="display: list-item;"> Follow these instructions to create a project from the API </summary>
#
# When you run the [command](#command) to create a project, it outputs a unique identifier of the project. This identifier is used to recognize, access and modify the project from the API.
#
# <a id="command"></a>
# ```python
# kili.create_project(
# title='Project Title',
# description='Project Description',
# input_type='IMAGE',
# json_interface=interface
# )
# ```
#
# Example of such an output:
#
# ```python
# {'id': 'ckkpj7stx1bxc0jvk1gn9cu5v'}
# ```
#
# Another way to get this project identifier is to look at the URL you're in:
#
# 
# </details>
# # Importing data <a id='data'></a>
# The next step is to import data.
#
# You can import data either [from the interface](https://cloud.kili-technology.com/docs/data-ingestion/data-ingestion-made-easy/) or [from the API](https://cloud.kili-technology.com/docs/python-graphql-api/recipes/import_assets/#kili-tutorial-importing-assets).
#
#
# To import data from the interface, go to the `Dataset` tab in your project, then click on `Add New`. There you'll have two tabs. From the first tab called `Upload Local Data` you'll be able to select files from your local computer to upload. From the second table called `Connect Cloud Data` you should provide a `.csv` file containing the URLs to your data stored in the cloud. These steps are shown below:
#
# 
# <details>
# <summary style="display: list-item;"> Follow these instructions to import data from the API </summary>
#
# Next, simply call [this function](https://cloud.kili-technology.com/docs/python-graphql-api/python-api/#append_many_to_dataset) :
#
# ```python
# kili.append_many_to_dataset(
# project_id="ckkpj7stx1bxc0jvk1gn9cu5v",
# content_array=["path-to-local-image OR url-to-image"],
# external_id_array=["your-identifier-of-the-image"]
# )
# ```
#
#
# ```python
# # Example
#
# project_id = result['id']
#
# kili.append_many_to_dataset(
# project_id=project_id,
# content_array=["https://raw.githubusercontent.com/kili-technology/kili-playground/master/recipes/img/car_2.jpg"],
# external_id_array=["car_2.jpg"]
# )
# ```
#
# ```python
# Out: {'id': 'ckm4pmqmk0000d49k6ewu2um5'}
# ```
# </details>
# # Labeling <a id='labeling'></a>
# When you create a project, you automatically become an admin of the project. This means that you can directly label. If you want to add members to the project, follow [](https://cloud.kili-technology.com/docs/projects/settings/#manage-project-members).
# ## Label a specific asset
# To annotate a specific asset, you can go to the dataset tab (in the side panel):
#
# <img src="../img/sidebar_dataset.png" width=100>
#
# 
#
# On the assets table, simply click on the line/asset (i.e., image here) you want to annotate.
# ## Label the first asset in the queue
# Otherwise, you can start to annotate right away with the *Start Labeling* button.
# ## How to label ?
# Suppose that you've created a simple project as described in [Creating the project](#project). Now, to annotate, go to *Overview* and click on *Start Labeling*. You should arrive at the image of a car. Now just select a class (*Object A* or *Object B*) and drag-and-drop your bounding box. You should use the red lines as guides while annotating.
#
# 
#
# Select the category you want by clicking on the right radio button, or by pressing the key underlined in the class name "o" for *Object A* and "b" for *Object B*.
#
# Then, click `Submit` to send the label.
# <details>
# <summary style="display: list-item;"> Follow these instructions to add a label from the API </summary>
#
# For this task, you need to know the identifier of the asset (image)—either from the URL when you are on an asset:
#
# 
#
# or from the API, retrieving the assets of the project:
#
#
# ```python
# assets = kili.assets(
# project_id=project_id,
# fields=['id']
# )
# asset_id = assets[0]['id']
# print(asset_id)
# ```
#
# 100%|██████████| 1/1 [00:00<00:00, 27.40it/s]
#
# ckm4pmuy30006d49kh0q64i0g
#
#
#
#
#
#
# ```python
# kili.append_to_labels(
# json_response=json_response,
# label_asset_id=asset_id,
# project_id=project_id
# )
# ```
#
# Output: {'id': 'ckm4pmzlj0009d49k1avaeubv'}
#
# with `json_response` as:
#
# ```python
# json_response = {
# "JOB_0": {
# "annotations": [
# {
# "boundingPoly": [
# {
# "normalizedVertices": [
# {"x": 0.1583940754900832, "y": 0.8310155382604694},
# {"x": 0.1583940754900832, "y": 0.3193681402157794},
# {"x": 0.8192660632614974, "y": 0.3193681402157794},
# {"x": 0.8192660632614974, "y": 0.8310155382604694}
# ]
# }
# ],
# "categories": [{"confidence": 100, "name": "OBJECT_A"}],
# "mid": "2021042719075237-60379",
# "score": None,
# "type": "rectangle"
# }
# ]
# }
# }
# ```
#
# </details>
# # Exporting labels <a id='export'></a>
# ## Through the interface
# In the "Dataset" tab, you can export your labels.
#
# 
#
# 1. Choose your format and click on “Download.” An asynchronous job is triggered, preparing your data.
# 2. Next, you get a notification. Click on it, and click on the “Download” button to download your data.
#
# Notification appears | Notification list
# :--:|:--:
#  | <img src="../img/notification_opened.png" width=400>
# If you choose Kili's classic API format, you get this file:
#
# ```json
# [
# {
# "content": "https://cloud.kili-technology.com/api/label/v2/files?id=f436f198-cede-4380-a119-f5d827f8a8fa",
# "externalId": "car_2.jpg",
# "id": "ckm0ligy900uuc49k1idydxsk",
# "jsonMetadata": {},
# "labels": [
# {
# "author": {
# "email": "email of the author of the label",
# "id": "id of the author of the label",
# "name": "Admin"
# },
# "createdAt": "2021-03-08T14:32:09.063Z",
# "isLatestLabelForUser": true,
# "jsonResponse": {
# "JOB_0": {
# "annotations": [
# {
# "boundingPoly": [
# {
# "normalizedVertices": [
# { "x": 0.1583940754900832, "y": 0.8310155382604694 },
# { "x": 0.1583940754900832, "y": 0.3193681402157794 },
# { "x": 0.8192660632614974, "y": 0.3193681402157794 },
# { "x": 0.8192660632614974, "y": 0.8310155382604694 }
# ]
# }
# ],
# "categories": [{ "confidence": 100, "name": "OBJECT_A" }],
# "mid": "2021042719075237-60379",
# "score": null,
# "type": "rectangle"
# }
# ]
# }
# },
# "labelType": "DEFAULT",
# "modelName": null,
# "skipped": false
# }
# ]
# }
# ]
# ```
#
# [For details on the data export, click here](https://cloud.kili-technology.com/docs/data-export/data-export/#docsNav).
# <details>
# <summary style="display: list-item;"> Follow these instructions to export labels from the API </summary>
#
# ```python
# labels = kili.labels(
# project_id=project_id
# )
#
# def hide_sensitive(label):
# label['author'] = {
# 'email': 'email of the author of the label',
# 'id': 'identifier of the author of the label',
# 'name': 'name of the author of the label'
# }
# return label
#
# result_hidden = [hide_sensitive(label) for label in labels]
# result_hidden
# ```
#
#
#
#
# [{'author': {'email': 'email of the author of the label',
# 'id': 'identifier of the author of the label',
# 'name': 'name of the author of the label'},
# 'id': 'ckm4pmzlj0009d49k1avaeubv',
# 'jsonResponse': {
# "JOB_0": {
# "annotations": [
# {
# "boundingPoly": [
# {
# "normalizedVertices": [
# { "x": 0.1583940754900832, "y": 0.8310155382604694 },
# { "x": 0.1583940754900832, "y": 0.3193681402157794 },
# { "x": 0.8192660632614974, "y": 0.3193681402157794 },
# { "x": 0.8192660632614974, "y": 0.8310155382604694 }
# ]
# }
# ],
# "categories": [{ "confidence": 100, "name": "OBJECT_A" }],
# "mid": "2021042719075237-60379",
# "score": null,
# "type": "rectangle"
# }
# ]
# }
# },
# 'labelType': 'DEFAULT',
# 'secondsToLabel': 0,
# 'skipped': False}]
#
#
#
# Our API uses GraphQL. You can simply choose the fields you want to fetch by specifying a list:
#
#
# ```python
# labels = kili.labels(
# project_id=project_id,
# fields=['id', 'createdAt', 'labelOf.externalId']
# )
# assert len(labels) > 0
# labels
# ```
#
#
#
#
# [{'labelOf': {'externalId': 'car_2.jpg'},
# 'id': 'ckm4pmzlj0009d49k1avaeubv',
# 'createdAt': '2021-03-11T10:10:20.984Z'}]
#
#
#
# Of course, you have plenty more options/filters:
#
#
# ```python
# help(kili.labels)
# ```
#
# Help on method labels in module kili.queries.label:
#
# labels(asset_id: str = None, asset_status_in: List[str] = None, asset_external_id_in: List[str] = None, author_in: List[str] = None, created_at: str = None, created_at_gte: str = None, created_at_lte: str = None, fields: list = ['author.email', 'author.id', 'author.name', 'id', 'jsonResponse', 'labelType', 'secondsToLabel', 'skipped'], first: int = None, honeypot_mark_gte: float = None, honeypot_mark_lte: float = None, id_contains: List[str] = None, json_response_contains: List[str] = None, label_id: str = None, project_id: str = None, skip: int = 0, skipped: bool = None, type_in: List[str] = None, user_id: str = None) method of kili.playground.Playground instance
# Get an array of labels from a project given a set of criteria
#
# Parameters
# ----------
# - asset_id : str, optional (default = None)
# Identifier of the asset.
# - asset_status_in : list of str, optional (default = None)
# Returned labels should have a status that belongs to that list, if given.
# Possible choices : {'TODO', 'ONGOING', 'LABELED', 'REVIEWED'}
# - asset_external_id_in : list of str, optional (default = None)
# Returned labels should have an external id that belongs to that list, if given.
# - author_in : list of str, optional (default = None)
# Returned labels should have a label whose status belongs to that list, if given.
# - created_at : string, optional (default = None)
# Returned labels should have a label whose creation date is equal to this date.
# Formatted string should have format : "YYYY-MM-DD"
# - created_at_gt : string, optional (default = None)
# Returned labels should have a label whose creation date is greater than this date.
# Formatted string should have format : "YYYY-MM-DD"
# - created_at_lt : string, optional (default = None)
# Returned labels should have a label whose creation date is lower than this date.
# Formatted string should have format : "YYYY-MM-DD"
# - fields : list of string, optional (default = ['author.email', 'author.id','author.name', 'id', 'jsonResponse', 'labelType', 'secondsToLabel', 'skipped'])
# All the fields to request among the possible fields for the labels.
# See [the documentation](https://cloud.kili-technology.com/docs/python-graphql-api/graphql-api/#label) for all possible fields.
# - first : int, optional (default = None)
# Maximum number of labels to return. Can only be between 0 and 100.
# - honeypot_mark_gt : float, optional (default = None)
# Returned labels should have a label whose honeypot is greater than this number.
# - honeypot_mark_lt : float, optional (default = None)
# Returned labels should have a label whose honeypot is lower than this number.
# - id_contains : list of str, optional (default = None)
# Filters out labels not belonging to that list. If empty, no filtering is applied.
# - json_response_contains : list of str, optional (default = None)
# Returned labels should have a substring of the jsonResponse that belongs to that list, if given.
# - label_id : str
# Identifier of the label.
# - project_id : str
# Identifier of the project.
# - skip : int, optional (default = None)
# Number of labels to skip (they are ordered by their date of creation, first to last).
# - skipped : bool, optional (default = None)
# Returned labels should have a label which is skipped
# - type_in : list of str, optional (default = None)
# Returned labels should have a label whose type belongs to that list, if given.
# - user_id : str
# Identifier of the user.
#
#
# Returns
# -------
# - a result object which contains the query if it was successful, or an error message else.
#
# Examples
# -------
# >>> # List all labels of a project and their assets external ID
# >>> playground.labels(project_id=project_id, fields=['jsonResponse', 'labelOf.externalId'])
#
# </details>
# # Quality Management<a id='quality'></a>
# To ensure that your model performs well, it's essential that your annotations are good quality. Using Kili, you have two main ways to measure the quality of the annotations: consensus and honeypot. Consensus basically is the measure of agreement between annotations from different annotators. Honeypot is measured by comparing the annotations of your annotators to a gold standard that you should provide beforehand.
#
# To access the quality management tab, go to ”Settings” (gear icon), then “Quality Management” as shown below:
#
# 
#
# Please follow the links below for detailed information:
#
# - [Quality management](https://cloud.kili-technology.com/docs/quality/quality-management/#docsNav)
# - Settings up quality metrics : [Consensus](https://cloud.kili-technology.com/docs/quality/consensus/#docsNav) and [Honeypot](https://cloud.kili-technology.com/docs/quality/honeypot/)
# # More advanced concepts <a id='concepts'></a>
# Here we list some of the more advanced features:
#
# - [Importing predictions](https://cloud.kili-technology.com/docs/python-graphql-api/recipes/import_predictions/#docsNav)
# - [Reviewing the labels](https://cloud.kili-technology.com/docs/quality/review-process/#docsNav)
# - [Issue/Question system](https://cloud.kili-technology.com/docs/quality/question-issue/#docsNav)
#
# [The full API definition can be found here](https://cloud.kili-technology.com/docs/python-graphql-api/python-api/#docsNav).
| recipes/getting-started/getting_started-image_bounding_box_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src='imgs/ooipy_banner.png' align="center">
#
# # OOIPY Demo
#
# This Jupyter Notebook walks through the basic functionality of OOIpy
#
# ## In this demo you will learn:
# - How to download broadband (Fs=64kHz) and low frequency (Fs = 200) Hz from OOI server
# - How to compute spectrograms and PSDs from the data
# - How to save spectrograms and PSDs to pickle files
# - How to plot hydrophone data, spectrograms and PSDs
# - How to save Hydrophone Data to .wav file
# ## Environment Setup
# + tags=[]
import ooipy
import datetime
from matplotlib import pyplot as plt
# Some Jupyter magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# -
# ## Download Data from OOI
#
# The functions ooipy.request.hydrophone_request.get_acoustic_data and ooipy.request.hydrophone_request.get_acoustic_data_LF require node string that specifies the OOI node to get data from. The available nodes are listed below
#
# ### Broadband Hydrophones
# * [Oregon Shelf Base Seafloor (Fs = 64 kHz)](https://ooinet.oceanobservatories.org/data_access/?search=CE02SHBP-LJ01D-11-HYDBBA106)
# * 'LJ01D'
# * [Oregon Slope Base Seafloor (Fs = 64 kHz)](https://ooinet.oceanobservatories.org/data_access/?search=RS01SLBS-LJ01A-09-HYDBBA102)
# * 'LJ01A'
# * [Slope Base Shallow (Fs = 64 kHz)](https://ooinet.oceanobservatories.org/data_access/?search=RS01SBPS-PC01A-08-HYDBBA103)
# * 'PC01A'
# * [Axial Base Shallow Profiler (Fs = 64 kHz)](https://ooinet.oceanobservatories.org/data_access/?search=RS03AXPS-PC03A-08-HYDBBA303)
# * 'PC03A'
# * [Offshore Base Seafloor (Fs = 64 kHz)](https://ooinet.oceanobservatories.org/data_access/?search=CE04OSBP-LJ01C-11-HYDBBA105)
# * 'LJ01C'
# * [Axial Base Seafloor (Fs = 64 kHz)](https://ooinet.oceanobservatories.org/data_access/?search=RS03AXBS-LJ03A-09-HYDBBA302)
# * 'LJ03A'
#
# ### Low Frequency Hydrophones
# * [Axial Base Seaflor (Fs = 200 Hz)](https://ooinet.oceanobservatories.org/data_access/?search=RS03AXBS-MJ03A-05-HYDLFA301)
# * 'Axial_Base'
# * 'AXABA1'
# * [Central Caldera (Fs = 200 Hz)](https://ooinet.oceanobservatories.org/data_access/?search=RS03CCAL-MJ03F-06-HYDLFA305)
# * 'Central_Caldera'
# * 'AXCC1'
# * [Eastern Caldera (Fs = 200 Hz)](https://ooinet.oceanobservatories.org/data_access/?search=RS03ECAL-MJ03E-09-HYDLFA304)
# * 'Eastern_Caldera'
# * 'AXEC2'
# * [Southern Hydrate (Fs = 200 Hz)](https://ooinet.oceanobservatories.org/data_access/?search=RS01SUM1-LJ01B-05-HYDLFA104)
# * 'Southern_Hydrate'
# * 'HYS14'
# * ['Oregon Slope Base Seafloor (Fs = 200 Hz)](https://ooinet.oceanobservatories.org/data_access/?search=RS01SLBS-MJ01A-05-HYDLFA101)
# * 'Slope_Base'
# * 'HYSB1'
#
# For more detailed information about hydrophones nodes see https://ooinet.oceanobservatories.org/
#
# For a map of hydrophones supported by OOIpy, see https://www.google.com/maps/d/u/1/viewer?mid=1_QKOPTxX2m5CTwgKR5fAGLO0lmbBgT7w&ll=45.16765319565428%2C-127.15744999999998&z=7
# +
# Specify start time, end time, and node for data download (1 minutes of data)
start_time = datetime.datetime(2017,7,1,0,0,0)
end_time = datetime.datetime(2017,7,1,0,1,0)
node1 = 'LJ01D'
node2 = 'Eastern_Caldera'
# Download Broadband data
print('Downloading Broadband Data:')
hdata_broadband = ooipy.get_acoustic_data(start_time, end_time, node1, verbose=True)
print('')
print('Downloading Low Frequency Data:')
hdata_lowfreq = ooipy.get_acoustic_data_LF(start_time, end_time, node2, verbose=True, zero_mean=True)
# -
# The returned data type is the ooipy.hydrophone.basic.HydrophoneData object.
# https://ooipy.readthedocs.io/en/latest/hydrophone.html#ooipy.hydrophone.basic.HydrophoneData
#
# To see the data stats, do the following:
print(hdata_broadband.stats)
print(hdata_lowfreq.stats)
# ### Plot Time Series Data
ooipy.plot(hdata_broadband)
ooipy.plot(hdata_lowfreq)
# ### Computing Spectrograms and PSDs
# +
spec1 = hdata_broadband.compute_spectrogram()
spec2 = hdata_lowfreq.compute_spectrogram(avg_time=1, overlap=0)
psd1 = hdata_broadband.compute_psd_welch()
psd2 = hdata_lowfreq.compute_psd_welch()
# -
ooipy.plot(spec1, fmin=0, fmax=28000, res_reduction_time=100, xlabel_rot=30)
ooipy.plot(spec2, fmin=0, fmax=100, xlabel_rot=30)
ooipy.plot(psd1, fmin=10, fmax=32000)
ooipy.plot(psd2, fmin=0.1,fmax=100)
# To save a spectrogram or PSD to pickle file, use the save() method of the Spectrogram or PSD class
psd2.save(filename='psd2.pkl')
spec2.save(filename='spec2.pkl')
# ### Save Hydrophone Data to Wav File
# to save the data to a wav file, use the wav_write method of the HydrophoneData class.
#
# Options include normalizing waveform (for media player playback), and changing the sample rate to desired sample rate.
# Note, normalize option only works if data is already zero mean
hdata_broadband.wav_write(filename='raw_data.wav')
hdata_lowfreq.wav_write(filename='audio_for_playback.wav', norm=True, new_sample_rate=44100)
# ### Different PSD estimators
# +
# power spectral density estimate of noise data using Welch's method
fig, ax = plt.subplots(figsize=(22,14), dpi=100)
# 1. using median averaging (default)
hdata_broadband.compute_psd_welch()
f = hdata_broadband.psd.freq / 1000
plt.plot(f, hdata_broadband.psd.values, label='Welch median',color='r')
# 2. using mean averaging
hdata_broadband.compute_psd_welch(avg_method='mean')
plt.plot(f, hdata_broadband.psd.values, label='Welch mean',color='b')
plt.xlabel('frequency [kHz]')
plt.ylabel('SDF [dB re µPa**2/Hz]')
plt.xlim(1,25)
plt.ylim(25,70)
plt.legend()
# -
| hydrophone_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Grove ADC Example
#
# This example shows how to use the Grove ADC.
#
# A Grove I2C ADC (v1.2) and PYNQ Grove Adapter are required. An analog input is also required. In this example, the Grove slide potentiometer was used.
#
# In the example, the ADC is initialized, a test read is done, and then the sensor is set to log a reading every 100 milliseconds. The ADC can be connected to any Grove peripheral that provides an analog voltage.
# ## 1. Using Pmod to Grove Adapter
# This example uses the PYNQ Pmod to Grove adapter. The adapter is connected to PMODA, and the grove ADC is connected to group `G4` on adapter.
# ### 1. Simple ADC read()
from pynq.overlays.base import BaseOverlay
base = BaseOverlay("base.bit")
# +
from pynq.lib.pmod import Grove_ADC
from pynq.lib.pmod import PMOD_GROVE_G4
grove_adc = Grove_ADC(base.PMODA,PMOD_GROVE_G4)
print("{} V".format(round(grove_adc.read(),4)))
# -
# ### 2. Starting logging once every 100 milliseconds
grove_adc.set_log_interval_ms(100)
grove_adc.start_log()
# ### 3. Try to change the input signal during the logging.
#
# For example, if using the Grove slide potentiometer, move the slider back and forth (slowly).
#
# Stop the logging whenever done trying to change sensor's value.
log = grove_adc.get_log()
# ### 4. Plot values over time
# The voltage values can be logged and displayed.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
plt.plot(range(len(log)), log, 'ro')
plt.title('Grove ADC Voltage Log')
plt.axis([0, len(log), min(log), max(log)])
plt.show()
# -
# ## 2. Using Arduino Shield
# This example uses the PYNQ Arduino shield. The grove ADC can be connected to any of the `I2C` groups on the shield.
# ### 1. Instantiation and read a single value
from pynq.lib.arduino import Grove_ADC
from pynq.lib.arduino import ARDUINO_GROVE_I2C
grove_adc = Grove_ADC(base.ARDUINO,ARDUINO_GROVE_I2C)
print("{} V".format(round(grove_adc.read(),4)))
# ### 2. Starting logging once every 100 milliseconds
grove_adc.set_log_interval_ms(100)
grove_adc.start_log()
# ### 3. Try to change the input signal during the logging.
#
# For example, if using the Grove slide potentiometer, move the slider back and forth (slowly).
#
# Stop the logging whenever done trying to change sensor's value.
log = grove_adc.get_log()
# ### 4. Plot values over time
# The voltage values can be logged and displayed.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
plt.plot(range(len(log)), log, 'ro')
plt.title('Grove ADC Voltage Log')
plt.axis([0, len(log), min(log), max(log)])
plt.show()
| boards/Pynq-Z2/base/notebooks/pmod/pmod_grove_adc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dilip1906/dsmp-pre-work/blob/master/Python_Getting_Started_Code_Along_.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="s9UNWnCNcydR" colab_type="text"
# # IPL Dataset Analysis
#
# ## Problem Statement
# We want to know as to what happens during an IPL match which raises several questions in our mind with our limited knowledge about the game called cricket on which it is based. This analysis is done to know as which factors led one of the team to win and how does it matter.
# + [markdown] id="AcXDZ50BcydT" colab_type="text"
# ## About the Dataset :
# The Indian Premier League (IPL) is a professional T20 cricket league in India contested during April-May of every year by teams representing Indian cities. It is the most-attended cricket league in the world and ranks sixth among all the sports leagues. It has teams with players from around the world and is very competitive and entertaining with a lot of close matches between teams.
#
# The IPL and other cricket related datasets are available at [cricsheet.org](https://cricsheet.org/%c2%a0(data). Feel free to visit the website and explore the data by yourself as exploring new sources of data is one of the interesting activities a data scientist gets to do.
# + [markdown] id="FCz9YiTocydW" colab_type="text"
# ### Analysing data with basic python operation
# + [markdown] id="VlyIHiL2cydZ" colab_type="text"
# ## Read the data of the format .yaml type
# + id="B8RfqR3Ccyda" colab_type="code" colab={}
import yaml
# + id="SnnFWf4Xcydi" colab_type="code" colab={}
import yaml
# using with open command to read the file
with open('/content/ipl_match.yaml') as f:
data = yaml.load(f)
# + id="K1GjMGEIcydn" colab_type="code" outputId="dcdaf062-898c-4c2a-f190-ce293ac319a4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
data
# + [markdown] id="9KuZ3hiQcydv" colab_type="text"
# Now let's find answers to some prilminary questions such as
# + [markdown] id="NQaI0d3vcydx" colab_type="text"
# ### Can you guess the data type with which your working on ?
# + id="D9C2-JKYcydy" colab_type="code" outputId="da8d2390-33c3-4b19-9b14-69cde66a60a0" colab={"base_uri": "https://localhost:8080/", "height": 34}
type(data)
# + id="ZgkwOXIGeyIu" colab_type="code" outputId="4a391002-1e1c-4dcf-fd59-15bce0f624a8" colab={"base_uri": "https://localhost:8080/", "height": 241}
data.get('info')
# + [markdown] id="ekR3vc1gcyd3" colab_type="text"
# ### In which city the match was played and where was it played ?
# + id="k3Ot6h6scyd4" colab_type="code" outputId="ab3bb787-e1a8-423a-adde-a9d88a3b8dba" colab={"base_uri": "https://localhost:8080/", "height": 51}
print('The Match was played at: ',data.get('info').get('city'))
print('The venue for the match was: ', data.get('info').get('venue'))
# + [markdown] id="GICfVwxwcyd_" colab_type="text"
# ### Which are all the teams that played in the tournament ? How many teams participated in total?
# + id="vINa9mIOcyeA" colab_type="code" outputId="8dfe2fa0-bbe7-4f48-9f99-6a4b1fd74ffd" colab={"base_uri": "https://localhost:8080/", "height": 71}
x = data.get('info').get('teams')
for i in range(len(x)):
print('The teams participated are: ', x[i], end = " ")
i+=1
print('\n The number of teams played are: ', len(x))
# + [markdown] id="TqXCPgZjcyeD" colab_type="text"
# ### Which team won the toss and what was the decision of toss winner ?
# + id="R6h1CdqvcyeE" colab_type="code" colab={}
y = data.get('info').get('toss')
# + id="ZPE_LspncyeI" colab_type="code" outputId="e3ff39ff-67c6-4e1b-9163-3ff7958e91a5" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(y['winner'], 'won the toss')
print(y['winner'], 'decides to', y['decision'])
# + [markdown] id="528y--2qcyeK" colab_type="text"
# ### Find the first bowler who played the first ball of the first inning. Also the first batsman who faced first delivery ?
# + id="icSMfmIIcyeL" colab_type="code" outputId="5831a679-f73d-426e-a733-1193d60da528" colab={"base_uri": "https://localhost:8080/", "height": 51}
x = data['innings'][0]
y = x['1st innings']
z = y['deliveries']
#print(z)
#print(type(z))
a = z[0]
#print(a)
b = a[0.1]
#print(b)
c = b['bowler']
print('The first ball was bowled by: ', c)
print('The first batsman who faced the delivery : ', b['batsman'])
# + id="rQqDY8BlpFRa" colab_type="code" colab={}
# + [markdown] id="CQgaycI3cyeN" colab_type="text"
# ### How many deliveries were delivered in first inning ?
# + id="K3fw9Nr1cyeO" colab_type="code" outputId="c5158877-c4b6-4d11-e44c-88440218a366" colab={"base_uri": "https://localhost:8080/", "height": 34}
y = data.get('innings')
#print(y)
a = y[0]
#print(a)
b = a.get('1st innings')
#print(b)
c = b.get('deliveries')
print("There were {} deliveries bowled in 1 st innings ".format(len(c)))
# + [markdown] id="YmFnWKVvcyeP" colab_type="text"
# ### How many deliveries were delivered in second inning ?
# + id="eCoofLrbcyeQ" colab_type="code" outputId="59286a5a-bde6-4e8f-fed4-2d802bb258e8" colab={"base_uri": "https://localhost:8080/", "height": 34}
#print(y[1])
z = y[1]
#print(z.get('2nd innings'))
d = z.get('2nd innings')
e = len(d.get('deliveries'))
print('There were {} ball were bowled in second innings'.format(e))
# + [markdown] id="8hTLjewJcyeR" colab_type="text"
# ### Which team won and how ?
#
# + id="__U5hNimcyeS" colab_type="code" outputId="2b6483cd-b6a0-43ce-e651-2da1c6348ba9" colab={"base_uri": "https://localhost:8080/", "height": 34}
win = data.get('info').get('outcome')
#print(win)
print('{} won the game by {} runs'.format(win.get('winner'), win.get('by').get('runs')))
# + id="ZcH01Ge3voZO" colab_type="code" colab={}
| Python_Getting_Started_Code_Along_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # D1 - 03 - Mini Projects
#
# ## Content
# - Scalar product of two lists/tuples
# - Arithmetic mean of a sequence
# - Linear regression
# - Numerical differentiation
# - Taylor series expansion
#
# ## Remember jupyter notebooks
# - To run the currently highlighted cell, hold <kbd>⇧ Shift</kbd> and press <kbd>⏎ Enter</kbd>.
# - To get help for a specific function, place the cursor within the function's brackets, hold <kbd>⇧ Shift</kbd>, and press <kbd>⇥ Tab</kbd>.
#
# ## Scalar product
# Implement a function
#
# ```Python
# def scalar_product(a, b):
# pass
# ```
#
# which implements the scalar product
# $$\left\langle \mathbf{a},\mathbf{b} \right\rangle = \sum\limits_{n=0}^{N-1} a_n b_n$$
# where $N$ is the number of elements in each $\mathbf{a}$ and $\mathbf{b}$. Both variables `a` and `b` can be `list`s or `tuple`s, and their elements should be numerical (`float` or `int`).
#
# **Bonus**: the function should not return a numerical result if both variables have different lengths or contain non-numerical elements.
def scalar_product(a, b):
pass
assert scalar_product([0] * 100, [1] * 100) == 0
assert scalar_product([1] * 100, [1, -1] * 50) == 0
assert scalar_product([1] * 100, range(100)) == 99 * 50
# ## Arithmetic mean
# Implement a function
# ```Python
# def mean(a):
# pass
# ```
# which computes the arithmetic mean of a sequence:
# $$\bar{a} = \frac{\sum_{n=0}^{N-1} a_n}{N}$$
# where $N$ is the number of elements $a_0,\dots,a_{N-1}$. The parameter `a` may be any type of `iterable` with only numerical elements.
#
# **Bonus**: for a sequence of length 0, e.g., an empty list, the function should return 0.
def mean(a):
pass
assert mean(range(100)) == 99 * 0.5
assert mean([]) == 0
assert mean([1] * 1000) == 1
# ## Linear regression
# Implement a function
# ```Python
# def linear_regression(x, y):
# slope = None
# const = None
# return slope, const
# ```
# which performs a simple linear regression
# $$\begin{eqnarray*}
# \textrm{slope} & = & \frac{\sum_{n=0}^{N-1} \left( x_n - \bar{x} \middle) \middle( y_n - \bar{y} \right)}{\sum_{n=0}^{N-1} \left( x_n - \bar{x} \right)^2} \\
# \textrm{const} & = & \bar{y} - \textrm{slope } \bar{x}
# \end{eqnarray*}$$
#
# for value pairs $(x_0, y_0),\dots,(x_{N-1},y_{N-1})$. The parameters `x` and `y` may be any type of `iterable` with only numerical elements; both must have the same length.
def linear_regression(x_values, y_values):
pass
x = [10, 14, 16, 15, 16, 20]
y = [ 1, 3, 5, 6, 5, 11]
slope, const = linear_regression(x, y)
assert 0.97 < slope < 0.99
assert -9.72 < const < -9.70
# ## Numerical differentiation
# Implement a function
# ```Python
# def differentiate(func, x, dx):
# pass
# ```
# where `func` is a reference to some function `f(x)`, `x` is the position where `func` shall be differentiated, and `dx` is the denominator of the differential quotient:
# $$f^\prime(x) \approx \frac{f(x + \frac{1}{2} \text{d}x) - f(x - \frac{1}{2} \text{d}x)}{\text{d}x}.$$
def differentiate(func, x, dx):
pass
# +
def f(x):
return x**2 + 1
assert -0.1 < differentiate(f, 0, 0.01) < 0.1
assert 1.9 < differentiate(f, 1, 0.01) < 2.1
assert 3.9 < differentiate(f, 2, 0.01) < 4.1
# -
# ## Taylor series expansion
#
# The Taylor series for $\sin(x)$ is:
#
# $$\sin(x)=\sum_{k=0}^\infty \frac{(-1)^k x^{1+2 k}}{(1+2 k)!}$$
#
# Implement a function
#
# ```Python
# def taylor_sin(x, n):
# pass
# ```
#
# where `x` is the position where the $\sin$ should be evaluated, and `n` is the degree of the approximating polynomial. It would be helpful to implement a factorial function that returns the factorial of a given integer n.
# +
def factorial(n):
pass
def taylor_sin(x,n):
pass
# +
from math import pi
order =
assert taylor_sin(0, order) = 0
assert taylor_sin(pi / 2, order) = 1
assert taylor_sin(pi, order) = 0
assert taylor_sin(-pi / 2, order) = -1
assert taylor_sin(2 * pi, order) = 0
| D1-03-mini-projects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # !pip install ray[tune]
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from sklearn.metrics import mean_squared_error
from hyperopt import hp
from ray import tune
from hyperopt import fmin, tpe, hp,Trials, space_eval
import scipy.stats
df = pd.read_csv("../../Data/Raw/flightLogData.csv")
plt.figure(figsize=(20, 10))
plt.plot(df.Time, df['Altitude'], linewidth=2, color="r", label="Altitude")
plt.plot(df.Time, df['Vertical_velocity'], linewidth=2, color="y", label="Vertical_velocity")
plt.plot(df.Time, df['Vertical_acceleration'], linewidth=2, color="b", label="Vertical_acceleration")
plt.legend()
plt.show()
temp_df = df[['Altitude', "Vertical_velocity", "Vertical_acceleration"]]
noise = np.random.normal(2, 5, temp_df.shape)
noisy_df = temp_df + noise
noisy_df['Time'] = df['Time']
plt.figure(figsize=(20, 10))
plt.plot(noisy_df.Time, noisy_df['Altitude'], linewidth=2, color="r", label="Altitude")
plt.plot(noisy_df.Time, noisy_df['Vertical_velocity'], linewidth=2, color="y", label="Vertical_velocity")
plt.plot(noisy_df.Time, noisy_df['Vertical_acceleration'], linewidth=2, color="b", label="Vertical_acceleration")
plt.legend()
plt.show()
# ## Altitude
q = 0.001
A = np.array([[1.0, 0.1, 0.005], [0, 1.0, 0.1], [0, 0, 1]])
H = np.array([[1.0, 0.0, 0.0],[ 0.0, 0.0, 1.0]])
P = np.array([[1.0, 0.0, 0.0], [0.0, 1.0, 0.0], [0.0, 0.0, 1.0]])
# R = np.array([[0.5, 0.0], [0.0, 0.0012]])
# Q = np.array([[q, 0.0, 0.0], [0.0, q, 0.0], [0.0, 0.0, q]])
I = np.identity(3)
x_hat = np.array([[0.0], [0.0], [0.0]])
Y = np.array([[0.0], [0.0]])
def kalman_update(param):
r1, r2, q1 = param['r1'], param['r2'], param['q1']
R = np.array([[r1, 0.0], [0.0, r2]])
Q = np.array([[q1, 0.0, 0.0], [0.0, q1, 0.0], [0.0, 0.0, q1]])
A = np.array([[1.0, 0.05, 0.00125], [0, 1.0, 0.05], [0, 0, 1]])
H = np.array([[1.0, 0.0, 0.0],[ 0.0, 0.0, 1.0]])
P = np.array([[1.0, 0.0, 0.0], [0.0, 1.0, 0.0], [0.0, 0.0, 1.0]])
I = np.identity(3)
x_hat = np.array([[0.0], [0.0], [0.0]])
Y = np.array([[0.0], [0.0]])
new_altitude = []
new_acceleration = []
new_velocity = []
for altitude, az in zip(noisy_df['Altitude'], noisy_df['Vertical_acceleration']):
Z = np.array([[altitude], [az]])
x_hat_minus = np.dot(A, x_hat)
P_minus = np.dot(np.dot(A, P), np.transpose(A)) + Q
K = np.dot(np.dot(P_minus, np.transpose(H)), np.linalg.inv((np.dot(np.dot(H, P_minus), np.transpose(H)) + R)))
Y = Z - np.dot(H, x_hat_minus)
x_hat = x_hat_minus + np.dot(K, Y)
P = np.dot((I - np.dot(K, H)), P_minus)
Y = Z - np.dot(H, x_hat_minus)
new_altitude.append(float(x_hat[0]))
new_velocity.append(float(x_hat[1]))
new_acceleration.append(float(x_hat[2]))
return new_altitude
def objective_function(param):
r1, r2, q1 = param['r1'], param['r2'], param['q1']
R = np.array([[r1, 0.0], [0.0, r2]])
Q = np.array([[q1, 0.0, 0.0], [0.0, q1, 0.0], [0.0, 0.0, q1]])
A = np.array([[1.0, 0.05, 0.00125], [0, 1.0, 0.05], [0, 0, 1]])
H = np.array([[1.0, 0.0, 0.0],[ 0.0, 0.0, 1.0]])
P = np.array([[1.0, 0.0, 0.0], [0.0, 1.0, 0.0], [0.0, 0.0, 1.0]])
I = np.identity(3)
x_hat = np.array([[0.0], [0.0], [0.0]])
Y = np.array([[0.0], [0.0]])
new_altitude = []
new_acceleration = []
new_velocity = []
for altitude, az in zip(noisy_df['Altitude'], noisy_df['Vertical_acceleration']):
Z = np.array([[altitude], [az]])
x_hat_minus = np.dot(A, x_hat)
P_minus = np.dot(np.dot(A, P), np.transpose(A)) + Q
K = np.dot(np.dot(P_minus, np.transpose(H)), np.linalg.inv((np.dot(np.dot(H, P_minus), np.transpose(H)) + R)))
Y = Z - np.dot(H, x_hat_minus)
x_hat = x_hat_minus + np.dot(K, Y)
P = np.dot((I - np.dot(K, H)), P_minus)
Y = Z - np.dot(H, x_hat_minus)
new_altitude.append(float(x_hat[0]))
new_velocity.append(float(x_hat[1]))
new_acceleration.append(float(x_hat[2]))
return mean_squared_error(df['Altitude'], new_altitude)
# +
# space = {
# "r1": hp.choice("r1", np.arange(0.01, 90, 0.005)),
# "r2": hp.choice("r2", np.arange(0.01, 90, 0.005)),
# "q1": hp.choice("q1", np.arange(0.0001, 0.0009, 0.0001))
# }
# + tags=[]
len(np.arange(0.00001, 0.09, 0.00001))
# + tags=[]
space = {
"r1": hp.choice("r1", np.arange(0.001, 90, 0.001)),
"r2": hp.choice("r2", np.arange(0.001, 90, 0.001)),
"q1": hp.choice("q1", np.arange(0.00001, 0.09, 0.00001))
}
# +
# Initialize trials object
trials = Trials()
best = fmin(fn=objective_function, space = space, algo=tpe.suggest, max_evals=100, trials=trials )
# -
print(best)
# -> {'a': 1, 'c2': 0.01420615366247227}
print(space_eval(space, best))
# -> ('case 2', 0.01420615366247227}
d1 = space_eval(space, best)
objective_function(d1)
# %%timeit
objective_function({'q1': 0.0013, 'r1': 0.25, 'r2': 0.65})
objective_function({'q1': 0.0013, 'r1': 0.25, 'r2': 0.65})
y = kalman_update(d1)
current = kalman_update({'q1': 0.0013, 'r1': 0.25, 'r2': 0.65})
plt.figure(figsize=(20, 10))
plt.plot(noisy_df.Time, df['Altitude'], linewidth=2, color="r", label="Actual")
plt.plot(noisy_df.Time, current, linewidth=2, color="g", label="ESP32")
plt.plot(noisy_df.Time, noisy_df['Altitude'], linewidth=2, color="y", label="Noisy")
plt.plot(noisy_df.Time, y, linewidth=2, color="b", label="Predicted")
plt.legend()
plt.show()
def kalman_update_return_velocity(param):
r1, r2, q1 = param['r1'], param['r2'], param['q1']
R = np.array([[r1, 0.0], [0.0, r2]])
Q = np.array([[q1, 0.0, 0.0], [0.0, q1, 0.0], [0.0, 0.0, q1]])
A = np.array([[1.0, 0.05, 0.00125], [0, 1.0, 0.05], [0, 0, 1]])
H = np.array([[1.0, 0.0, 0.0],[ 0.0, 0.0, 1.0]])
P = np.array([[1.0, 0.0, 0.0], [0.0, 1.0, 0.0], [0.0, 0.0, 1.0]])
I = np.identity(3)
x_hat = np.array([[0.0], [0.0], [0.0]])
Y = np.array([[0.0], [0.0]])
new_altitude = []
new_acceleration = []
new_velocity = []
for altitude, az in zip(noisy_df['Altitude'], noisy_df['Vertical_acceleration']):
Z = np.array([[altitude], [az]])
x_hat_minus = np.dot(A, x_hat)
P_minus = np.dot(np.dot(A, P), np.transpose(A)) + Q
K = np.dot(np.dot(P_minus, np.transpose(H)), np.linalg.inv((np.dot(np.dot(H, P_minus), np.transpose(H)) + R)))
Y = Z - np.dot(H, x_hat_minus)
x_hat = x_hat_minus + np.dot(K, Y)
P = np.dot((I - np.dot(K, H)), P_minus)
Y = Z - np.dot(H, x_hat_minus)
new_altitude.append(float(x_hat[0]))
new_velocity.append(float(x_hat[1]))
new_acceleration.append(float(x_hat[2]))
return new_velocity
def objective_function(param):
r1, r2, q1 = param['r1'], param['r2'], param['q1']
R = np.array([[r1, 0.0], [0.0, r2]])
Q = np.array([[q1, 0.0, 0.0], [0.0, q1, 0.0], [0.0, 0.0, q1]])
A = np.array([[1.0, 0.05, 0.00125], [0, 1.0, 0.05], [0, 0, 1]])
H = np.array([[1.0, 0.0, 0.0],[ 0.0, 0.0, 1.0]])
P = np.array([[1.0, 0.0, 0.0], [0.0, 1.0, 0.0], [0.0, 0.0, 1.0]])
I = np.identity(3)
x_hat = np.array([[0.0], [0.0], [0.0]])
Y = np.array([[0.0], [0.0]])
new_altitude = []
new_acceleration = []
new_velocity = []
for altitude, az in zip(noisy_df['Altitude'], noisy_df['Vertical_acceleration']):
Z = np.array([[altitude], [az]])
x_hat_minus = np.dot(A, x_hat)
P_minus = np.dot(np.dot(A, P), np.transpose(A)) + Q
K = np.dot(np.dot(P_minus, np.transpose(H)), np.linalg.inv((np.dot(np.dot(H, P_minus), np.transpose(H)) + R)))
Y = Z - np.dot(H, x_hat_minus)
x_hat = x_hat_minus + np.dot(K, Y)
P = np.dot((I - np.dot(K, H)), P_minus)
Y = Z - np.dot(H, x_hat_minus)
new_altitude.append(float(x_hat[0]))
new_velocity.append(float(x_hat[1]))
new_acceleration.append(float(x_hat[2]))
return mean_squared_error(df['Vertical_velocity'], new_velocity)
space = {
"r1": hp.choice("r1", np.arange(0.001, 90, 0.001)),
"r2": hp.choice("r2", np.arange(0.001, 90, 0.001)),
"q1": hp.choice("q1", np.arange(0.00001, 0.09, 0.00001))
}
# +
# Initialize trials object
trials = Trials()
best = fmin(fn=objective_function, space = space, algo=tpe.suggest, max_evals=100, trials=trials )
# -
print(best)
print(space_eval(space, best))
d2 = space_eval(space, best)
objective_function(d2)
y = kalman_update_return_velocity(d2)
current = kalman_update_return_velocity({'q1': 0.0013, 'r1': 0.25, 'r2': 0.65})
previous = kalman_update_return_velocity({'q1': 0.08519, 'r1': 4.719, 'r2': 56.443})
plt.figure(figsize=(20, 10))
plt.plot(noisy_df.Time, df['Vertical_velocity'], linewidth=2, color="r", label="Actual")
plt.plot(noisy_df.Time, current, linewidth=2, color="g", label="ESP32")
plt.plot(noisy_df.Time, previous, linewidth=2, color="c", label="With previous data")
plt.plot(noisy_df.Time, noisy_df['Vertical_velocity'], linewidth=2, color="y", label="Noisy")
plt.plot(noisy_df.Time, y, linewidth=2, color="b", label="Predicted")
plt.legend()
plt.show()
| Src/Notebooks/oprimizeValues.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Spam Filtering Using The [Enron Dataset][1]
# [1]: http://www.aueb.gr/users/ion/data/enron-spam/
from pymldb import Connection
mldb = Connection('http://localhost/')
# Let's start by loading the dataset. We have already merged the different email files in a sensible manner into a .csv file, which we've made available online. Since this dataset is actually made up of six different datasets, we'll restrict ourself to the first one for simplicity, using a "where" clause.
print mldb.post('/v1/procedures', {
'type': 'import.text',
'params': {
'dataFileUrl': 'http://public.mldb.ai/datasets/enron.csv.gz',
'outputDataset': 'enron_data',
'named': "'enron_' + dataset + '_mail_' + index",
'where': 'dataset = 1',
'runOnCreation': True
}
})
# This is what the dataset looks like.
#
# *index*: order in which the emails arrived in the user's inbox
# *msg*: actual content of the email
# *label*: was the email legitimate (*ham*) or not (*spam*)
mldb.query('select index, msg, label from enron_data order by index limit 10')
# Let's create a *sql.expression* that will simply tokenize the emails into a bag of words. Those will be our features on which we will train a classifier.
print mldb.put('/v1/functions/bow', {
'type': 'sql.expression',
'params': {
'expression': """
tokenize(msg, {splitChars: ' :.-!?''"()[],', quoteChar: ''}) as bow
"""
}
})
# Then we can generate the features for the whole dataset, and write them into a new dataset, using the *transform* procedure.
print mldb.post('/v1/procedures', {
'type': 'transform',
'params': {
'inputData': """
select bow({msg})[bow] as *, label = 'spam' as message_is_spam
from enron_data
""",
'outputDataset': 'enron_features',
'runOnCreation': True
}
})
# Here is a snapshot of the sparse feature matrix:
mldb.query('select * from enron_features limit 10')
# Finally, let's train a very simple classifier, by training on half of the messages, and testing on the other half. This classifier will give a score to every email, and we can then choose a threshold where everything above the threshold is classified as spam, and every thing below as ham.
# +
res = mldb.post('/v1/procedures', {
'type': 'classifier.experiment',
'params': {
'experimentName': 'enron_experiment1',
'inputData': '''
select
{* excluding(message_is_spam)} as features,
message_is_spam as label
from enron_features''',
'modelFileUrlPattern': 'file://enron_model_$runid.cls',
'algorithm': 'bbdt',
'runOnCreation': True
}
})
print 'AUC =', res.json()['status']['firstRun']['status']['folds'][0]['resultsTest']['auc']
# -
# This is an impressive-looking AUC!
#
# But [the AUC score of a classifier is only a very generic measure of performance][1]. When having a specific problem like spam filtering, we're better off using a performance metric that truly matches our intuition about what a good spam filter ought to be. Namely, a good spam filtering algorithm should almost never flag as spam a legitime email, while keeping your inbox as spam-free as possible. This is what should be used to choose the threshold for the classifier, and then to measure its performance.
#
# So instead of the AUC (that doesn't pick a specific threshold but uses all of them), let's use as our performance metric the best [$F_{0.05}$ score][2], which gives 20 times more importance to precision than recall. In other words, this metric represents the fact that classifying as spam **only** what is really spam is 20 times more important than finding all the spam.
#
# Let's see how we are doing with that metric.
# [1]: http://mldb.ai/blog/posts/2016/01/ml-meets-economics/
# [2]: https://en.wikipedia.org/wiki/F1_score
print mldb.put('/v1/functions/enron_score', {
'type': 'sql.expression',
'params': {
'expression': """
(1 + pow(ratio, 2)) * (precision * recall) / (precision * pow(ratio, 2) + recall) as enron_score
"""
}
})
mldb.query("""
select
"falseNegatives" as spam_in_inbox,
"trueNegatives" as ham_in_inbox,
"falsePositives" as ham_in_junkmail,
"truePositives" as spam_in_junkmail,
enron_score({precision, recall, ratio:0.05}) as *
named 'best_score'
from enron_experiment1_results_0
order by enron_score desc
limit 1
""")
# As you can see, in order to maximize our score (i.e. to get very few ham messages in the junkmail folder) we have to accept a very high proportion of spam in our inbox, meaning that even though we have a very impressive-looking AUC score, our spam filter isn't actually very good!
#
# You can read more about the dangers of relying too much on AUC and the benefits of using a problem-specific measure in our [Machine Learning Meets Economics](http://blog.mldb.ai/blog/posts/2016/01/ml-meets-economics/) series of blog posts.
# ## Where to next?
#
# Check out the other [Tutorials and Demos](../../../../doc/#builtin/Demos.md.html).
| container_files/demos/Enron Spam Filtering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "slide"}
import networkx as nx
import matplotlib.pyplot as plt
from collections import Counter
from custom import load_data as cf
from custom import ecdf
import warnings
warnings.filterwarnings('ignore')
from nxviz import CircosPlot
import numpy as np
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# + [markdown] slideshow={"slide_type": "slide"}
# ## Load Data
#
# We will load the [sociopatterns network](http://konect.uni-koblenz.de/networks/sociopatterns-infectious) data for this notebook. From the Konect website:
#
# > This network describes the face-to-face behavior of people during the exhibition INFECTIOUS: STAY AWAY in 2009 at the Science Gallery in Dublin. Nodes represent exhibition visitors; edges represent face-to-face contacts that were active for at least 20 seconds. Multiple edges between two nodes are possible and denote multiple contacts. The network contains the data from the day with the most interactions.
# + slideshow={"slide_type": "fragment"}
# Load the sociopatterns network data.
G = cf.load_sociopatterns_network()
# -
# How many nodes and edges are present?
len(G.nodes()), len(G.edges())
# + [markdown] slideshow={"slide_type": "slide"}
# # Hubs: How do we evaluate the importance of some individuals in a network?
#
# Within a social network, there will be certain individuals which perform certain important functions. For example, there may be hyper-connected individuals who are connected to many, many more people. They would be of use in the spreading of information. Alternatively, if this were a disease contact network, identifying them would be useful in stopping the spread of diseases. How would one identify these people?
# + [markdown] slideshow={"slide_type": "slide"}
# ## Approach 1: Neighbors
#
# One way we could compute this is to find out the number of people an individual is conencted to. NetworkX let's us do this by giving us a `G.neighbors(node)` function.
# + slideshow={"slide_type": "fragment"}
# Let's find out the number of neighbors that individual #7 has.
len(list(G.neighbors(7)))
# -
# **API Note:** As of NetworkX 2.0, `G.neighbors(node)` now returns a `dict_keyiterator`, which means we have to cast them as a `list` first in order to compute its length.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise
#
# Can you create a ranked list of the importance of each individual, based on the number of neighbors they have? (3 min.)
#
# Hint: One suggested output would be a list of tuples, where the first element in each tuple is the node ID (an integer number), and the second element is the number of neighbors that it has.
#
# Hint: Python's `sorted(iterable, key=lambda x:...., reverse=True)` function may be of help here.
# + slideshow={"slide_type": "fragment"}
sorted([______________], key=lambda x: __________, reverse=True)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Approach 2: Degree Centrality
#
# The number of other nodes that one node is connected to is a measure of its centrality. NetworkX implements a **degree centrality**, which is defined as the number of neighbors that a node has normalized to the number of individuals it could be connected to in the entire graph. This is accessed by using `nx.degree_centrality(G)`
# + slideshow={"slide_type": "fragment"}
nx.degree_centrality(G)
# Uncomment the next line to show a truncated version.
# list(nx.degree_centrality(G).items())[0:5]
# + [markdown] slideshow={"slide_type": "subslide"}
# If you inspect the dictionary closely, you will find that node 51 is the one that has the highest degree centrality, just as we had measured by counting the number of neighbors.
#
# There are other measures of centrality, namely **betweenness centrality**, **flow centrality** and **load centrality**. You can take a look at their definitions on the NetworkX API docs and their cited references. You can also define your own measures if those don't fit your needs, but that is an advanced topic that won't be dealt with here.
#
# The NetworkX API docs that document the centrality measures are here: http://networkx.readthedocs.io/en/networkx-1.11/reference/algorithms.centrality.html?highlight=centrality#module-networkx.algorithms.centrality
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercises
#
# The following exercises are designed to get you familiar with the concept of "distribution of metrics" on a graph.
#
# 1. Can you create an ECDF of the distribution of degree centralities?
# 2. Can you create an ECDF of the distribution of number of neighbors?
# 3. Can you create a scatterplot of the degree centralities against number of neighbors?
# 4. If I have `n` nodes, then how many possible edges are there in total, assuming self-edges are allowed? What if self-edges are not allowed?
#
# Exercise Time: 8 minutes.
#
# Here is what an ECDF is (https://en.wikipedia.org/wiki/Empirical_distribution_function).
# + [markdown] slideshow={"slide_type": "subslide"}
# Hint: You may want to use:
#
# ecdf(list_of_values)
#
# to get the empirical CDF x- and y-values for plotting, and
#
# plt.scatter(x_values, y_values)
#
# Hint: You can access the dictionary `.keys()` and `.values()` and cast them as a list.
#
# If you know the Matplotlib API, feel free to get fancy :).
# + slideshow={"slide_type": "subslide"}
# Possible Answers:
fig = plt.figure(0)
# Get a list of degree centrality scores for all of the nodes.
degree_centralities = list(____________)
x, y = ecdf(___________)
# Plot the histogram of degree centralities.
plt.scatter(____________)
# Set the plot title.
plt.title('Degree Centralities')
# + slideshow={"slide_type": "subslide"}
fig = plt.figure(1)
neighbors = [__________]
x, y = ecdf(_________)
plt.scatter(_________)
# plt.yscale('log')
plt.title('Number of Neighbors')
# + slideshow={"slide_type": "subslide"}
fig = plt.figure(2)
plt.scatter(_____________, ____________, alpha=0.1)
plt.xlabel('Degree Centralities')
plt.ylabel('Number of Neighbors')
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise
#
# Before we move on to paths in a network, see if you can use the Circos plot to visualize the network. Order and color the nodes according to the `order` keyword. (2 min.)
#
# The CircosPlot API needs documentation written; for now, I am providing the following "on-the-spot" docs for you.
# -
# To instantiate and draw a CircosPlot:
#
# ```python
# c = CircosPlot(G, node_order='node_key', node_color='node_key')
# c.draw()
# plt.show() # or plt.savefig(...)
# ```
#
# Notes:
#
# - `'node_key'` is a key in the node metadata dictionary that the CircosPlot constructor uses for determining the colour, grouping, and ordering of the nodes.
# - In the following exercise, you may want to use `order`, which is already encoded on each node in the graph.
# + slideshow={"slide_type": "subslide"}
c = CircosPlot(__________)
c._______()
plt.savefig('images/sociopatterns.png', dpi=300)
# + [markdown] slideshow={"slide_type": "subslide"}
# What can you deduce about the structure of the network, based on this visualization?
# + [markdown] slideshow={"slide_type": "fragment"}
# Place your own notes here :)
# + [markdown] slideshow={"slide_type": "slide"}
# # Paths in a Network
#
# Graph traversal is akin to walking along the graph, node by node, restricted by the edges that connect the nodes. Graph traversal is particularly useful for understanding the local structure (e.g. connectivity, retrieving the exact relationships) of certain portions of the graph and for finding paths that connect two nodes in the network.
#
# Using the synthetic social network, we will figure out how to answer the following questions:
#
# 1. How long will it take for a message to spread through this group of friends? (making some assumptions, of course)
# 2. How do we find the shortest path to get from individual A to individual B?
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Shortest Path
#
# Let's say we wanted to find the shortest path between two nodes. How would we approach this? One approach is what one would call a **breadth-first search** (http://en.wikipedia.org/wiki/Breadth-first_search). While not necessarily the fastest, it is the easiest to conceptualize.
# + [markdown] slideshow={"slide_type": "subslide"}
# The approach is essentially as such:
#
# 1. Begin with a queue of the starting node.
# 2. Add the neighbors of that node to the queue.
# 1. If destination node is present in the queue, end.
# 2. If destination node is not present, proceed.
# 3. For each node in the queue:
# 2. Add neighbors of the node to the queue. Check if destination node is present or not.
# 3. If destination node is present, end. <!--Credit: @cavaunpeu for finding bug in pseudocode.-->
# 4. If destination node is not present, continue.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise
#
# Try implementing this algorithm in a function called `path_exists(node1, node2, G)`. (15 min.)
#
# The function should take in two nodes, `node1` and `node2`, and the graph `G` that they belong to, and return a Boolean that indicates whether a path exists between those two nodes or not. For convenience, also print out whether a path exists or not between the two nodes.
# + slideshow={"slide_type": "subslide"}
def path_exists(node1, node2, G):
"""
This function checks whether a path exists between two nodes (node1, node2)
in graph G.
"""
visited_nodes = set()
queue = [node1]
# Fill in code below
for node in queue:
neighbors = G.neighbors(_______)
if ______ in neighbors:
print('Path exists between nodes {0} and {1}'.format(node1, node2))
return True
else:
_________.add(______)
_______.extend(________)
print('Path does not exist between nodes {0} and {1}'.format(node1, node2))
return False
# + slideshow={"slide_type": "subslide"}
# Test your answer below
def test_path_exists():
assert path_exists(18, 10, G)
assert path_exists(22, 51, G)
test_path_exists()
# + [markdown] slideshow={"slide_type": "subslide"}
# If you write an algorithm that runs breadth-first, the recursion pattern is likely to follow what we have done above. If you do a depth-first search (i.e. DFS), the recursion pattern is likely to look a bit different. Take it as a challenge exercise to figure out how a DFS looks like.
#
# Meanwhile... thankfully, NetworkX has a function for us to use, titled `has_path`, so we don't have to implement this on our own. :-) Check it out [here](https://networkx.github.io/documentation/stable/reference/algorithms/generated/networkx.algorithms.shortest_paths.generic.shortest_path.html#networkx.algorithms.shortest_paths.generic.shortest_path).
# + slideshow={"slide_type": "fragment"}
nx.has_path(G, 400, 1)
# + [markdown] slideshow={"slide_type": "subslide"}
# NetworkX also has [other shortest](https://networkx.github.io/documentation/stable/reference/algorithms/shortest_paths.html) path algorithms implemented.
#
# We can build upon these to build our own graph query functions. Let's see if we can trace the shortest path from one node to another.
# + [markdown] slideshow={"slide_type": "subslide"}
# `nx.shortest_path(G, source, target)` gives us a list of nodes that exist within one of the shortest paths between the two nodes. (Not all paths are guaranteed to be found.)
# + slideshow={"slide_type": "fragment"}
nx.shortest_path(G, 4, 400)
# + [markdown] slideshow={"slide_type": "fragment"}
# Incidentally, the node list is in order as well.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise
#
# Write a function that extracts the edges in the shortest path between two nodes and puts them into a new graph, and draws it to the screen. It should also return an error if there is no path between the two nodes. (5 min.)
#
# Hint: You may want to use `G.subgraph(iterable_of_nodes)` to extract just the nodes and edges of interest from the graph `G`. You might want to use the following lines of code somewhere:
#
# newG = G.subgraph(nodes_of_interest)
# nx.draw(newG)
#
# newG will be comprised of the nodes of interest and the edges that connect them.
# + slideshow={"slide_type": "subslide"}
# Possible Answer:
def extract_path_edges(G, source, target):
# Check to make sure that a path does exists between source and target.
if _______________________:
________ = nx._____________(__________)
newG = G.subgraph(________)
return newG
else:
raise Exception('Path does not exist between nodes {0} and {1}.'.format(source, target))
newG = extract_path_edges(G, 4, 400)
nx.draw(newG, with_labels=True)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Challenge Exercise (at home)
#
# These exercises below are designed to let you become more familiar with manipulating and visualizing subsets of a graph's nodes.
#
# Write a function that extracts only node, its neighbors, and the edges between that node and its neighbors as a new graph. Then, draw the new graph to screen.
# + slideshow={"slide_type": "skip"}
def extract_neighbor_edges(G, node):
return newG
fig = plt.figure(0)
newG = extract_neighbor_edges(G, 19)
nx.draw(newG, with_labels=True)
# + slideshow={"slide_type": "skip"}
def extract_neighbor_edges2(G, node):
return newG
fig = plt.figure(1)
newG = extract_neighbor_edges2(G, 19)
nx.draw(newG, with_labels=True)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Challenge Exercises (at home)
#
# Let's try some other problems that build on the NetworkX API. Refer to the following for the relevant functions:
#
# http://networkx.readthedocs.io/en/networkx-1.11/reference/algorithms.shortest_paths.html
#
# 1. If we want a message to go from one person to another person, and we assume that the message takes 1 day for the initial step and 1 additional day per step in the transmission chain (i.e. the first step takes 1 day, the second step takes 2 days etc.), how long will the message take to spread from any two given individuals? Write a function to compute this.
# 2. What is the distribution of message spread times from person to person? What about chain lengths?
# + slideshow={"slide_type": "skip"}
# Possible answer to Question 1:
# All we need here is the length of the path.
def compute_transmission_time(G, source, target):
"""
Fill in code below.
"""
return __________
compute_transmission_time(G, 14, 4)
# + slideshow={"slide_type": "skip"}
# Possible answer to Question 2:
# We need to know the length of every single shortest path between every pair of nodes.
# If we don't put a source and target into the nx.shortest_path_length(G) function call, then
# we get a dictionary of dictionaries, where all source-->target-->lengths are shown.
lengths = []
times = []
## Fill in code below ##
plt.figure(0)
plt.bar(Counter(lengths).keys(), Counter(lengths).values())
plt.figure(1)
plt.bar(Counter(times).keys(), Counter(times).values())
# + [markdown] slideshow={"slide_type": "slide"}
# # Hubs Revisited
#
# If a message has to be passed through the network in the shortest time possible, there may be "bottleneck" nodes through which information must always flow through. Such a node has a high **betweenness centrality**. This is implemented as one of NetworkX's centrality algorithms. Check out the Wikipedia page for a further description.
#
# http://en.wikipedia.org/wiki/Betweenness_centrality
# + slideshow={"slide_type": "subslide"}
btws = nx.betweenness_centrality(G, normalized=False)
plt.bar(list(btws.keys()), list(btws.values()))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise
#
# Plot betweeness centrality against degree centrality for the network data. (5 min.)
# + slideshow={"slide_type": "subslide"}
plt.scatter(__________, ____________)
plt.xlabel('degree')
plt.ylabel('betweeness')
plt.title('centrality scatterplot')
# + [markdown] slideshow={"slide_type": "subslide"}
# **Think about it...**
#
# From the scatter plot, we can see that the dots don't all fall on the same line. Degree centrality and betweenness centrality don't necessarily correlate. Can you think of scenarios where this is true?
# + [markdown] slideshow={"slide_type": "fragment"}
# What would be the degree centrality and betweenness centrality of the middle connecting node in the **barbell graph** below?
# + slideshow={"slide_type": "fragment"}
nx.draw(nx.barbell_graph(5, 1))
# -
| archive/3-hubs-and-paths-student.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SET 13 WEEK-3
# # Structured Programming Using Python
# **DONE BY:
# <br><NAME>
# <br>RA1911026010029
# <br>CSE-AIML**
# # 1.
# **Write the python code to calculate the gross pay of an employee. Create a class name called employee and get the following inputs from the user.
# <br>• Name of the employee,
# <br>• Age,
# <br>• DOB,
# <br>• DOJ,
# <br>• Basic pay
# <br>• HRA
# <br>• Deductions
# <br>Print the gross salary of each employee.**
# +
class Employee:
_name=""
_age=0
_dob=""
_doj=""
_basic_pay=0
_hra=0
_deduc=0
# constructor to set data
def __init__(self,name,age,dob,doj,basic_pay,hra,deduc):
self._name=name
self._age=age
self._dob=dob
self._doj=doj
self._basic_pay=basic_pay
self._hra=hra
self._deduc=deduc
# function to calculate gross salary
def calc0029(self) :
gross_sal=self._basic_pay+self._hra
net_sal=gross_sal-self._deduc
print(f"\033[1mThe gross salary is {gross_sal} & The net salary is {net_sal}\033[0m")
n0029=int(input('enter the no.of employees :'))
for i in range(n0029):
print(f"\033[1m please enter the employee id{i+1} details :\033[0m")
name29=input(f"please enter the name :")
age29=int(input("please enter the age :"))
dob29=input("please enter the date of birth :")
doj29=input("please enter the date of joining :")
basic_pay29=int(input("please enter the basic pay :"))
hra29=int(input("please enter the home rent allowance :"))
deduc29=int(input("please enter the deduction amount :"))
obj0029=Employee(name29,age29,dob29,doj29,basic_pay29,hra29,deduc29)
#obj.setData(name,age,dob,doj,basic_pay,hra,deduc)
obj0029.calc0029()
# -
# # 2.
# **Write a Python class named Rectangle constructed by a length and width and a method which will compute the area of a rectangle.**
# +
class Rectangle():
def __init__(self, l, w):
self.length = l
self.width = w
def rectangle_area(self):
return self.length*self.width
a0029,b0029=map(int,input("Enter the length and breadth of rectangle (in cm) : ").split())
obj0029 = Rectangle(a0029,b0029)
print("\033[1mArea of rectangle is \033[1m"+str(obj0029.rectangle_area())+" cm²")
# -
# # 3.
# **Write a Python class to find a pair of elements (indices of the two numbers) from a given
# array whose sum equals a specific target number.
# <br>Input: numbers= [10,20,10,40,50,60,70], target=50
# <br>Output: 3, 4**
class sol0029:
def Sum0029(self, nums, target):
lookup = {}
for i, num in enumerate(nums,1):
if target - num in lookup:
return (lookup[target - num], i )
lookup[num] = i
print("\033[1m %d,%d"% sol0029().Sum0029([10,20,10,40,50,60,70],50))
# # 4.
# **Write a Python class to find the three elements that sum to zero from a set of n real numbers.
# <br>Input array : [-25, -10, -7, -3, 2, 4, 8, 10]
# <br>Output : [[-10, 2, 8], [-7, -3, 10]]**
# +
class sol0029:
def zerosum0029(self, nums0029):
nums0029, result0029, i = sorted(nums0029), [], 0
while i < len(nums0029) - 2:
j, k = i + 1, len(nums0029) - 1
while j < k:
if nums0029[i] + nums0029[j] + nums0029[k] < 0:
j += 1
elif nums0029[i] + nums0029[j] + nums0029[k] > 0:
k -= 1
else:
result0029.append([nums0029[i], nums0029[j], nums0029[k]])
j, k = j + 1, k - 1
while j < k and nums0029[j] == nums0029[j - 1]:
j += 1
while j < k and nums0029[k] == nums0029[k + 1]:
k -= 1
i += 1
while i < len(nums0029) - 2 and nums0029[i] == nums0029[i - 1]:
i += 1
return result0029
print(sol0029().zerosum0029([-25, -10, -7, -3, 2, 4, 8, 10]))
# -
# # 5.
# **Write a Python class to reverse a string word by word.
# <br>Input string : 'hello .py'
# <br>Expected Output : '.py hello'**
# +
class sol0029:
def reverse_it0029(self, s):
return ' '.join(reversed(s.split()))
print("\033[1mInput : \033[0m")
s=input()
print("\033[1mOutput : \033[0m")
print(sol0029().reverse_it0029(s))
# -
# # Thank you
| WEEK-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: milo_py37
# language: python
# name: milo_py37
# ---
# ## Plotting ploidy of focal population clones
from matplotlib import pyplot as pl
import matplotlib.patches as patches
import numpy as np
import pandas as pd
from glob import glob
# this was installed like: pip install FlowCytometryTools
import FlowCytometryTools as fct
import seaborn as sns
sns.set_style('white')
# %matplotlib inline
cbs = sns.color_palette('colorblind')
def get_plate_data(dir_base, rows=8, cols=12, gated=False, lsr2=False):
td = dict()
c = 1
for let in [chr(i+65) for i in range(rows)]:
for col in range(1, cols+1):
try:
well = let + str(col).zfill(2)
if lsr2:
ending = '_' + str(c).zfill(3) + '.fcs'
else:
ending = '.fcs'
if gated:
fname = dir_base + let + str(col) + '_' + well + gated + ending
else:
fname = dir_base + let + str(col) + '_' + well + ending
samp = fct.FCMeasurement(ID=well, datafile=fname)
c += 1
td[well] = samp
except FileNotFoundError:
print(fname, 'not found!')
return td
p1 = pd.read_csv('../accessory_files/P1_Foc_B1_rearray.csv')
rearray = {r['rearray_well'][0] + str(int(r['rearray_well'][1:])-8).zfill(2): r['plate']+r['well'] for j, r in p1[p1['Generation']==10150].iterrows()}
p2 = pd.read_csv('../accessory_files/P2_Foc_B1_rearray.csv')
rearray.update({r['rearray_well'][0] + str(int(r['rearray_well'][1:])-4).zfill(2): r['plate']+r['well'] for j, r in p2[p2['Generation']==10150].iterrows()})
p3 = pd.read_csv('../accessory_files/P3_Foc_B1_rearray.csv')
rearray.update({r['rearray_well'][0] + str(int(r['rearray_well'][1:])).zfill(2): r['plate']+r['well'] for j, r in p3[p3['Generation']==10150].iterrows()})
well_info = pd.read_csv('../accessory_files/VLTE_by_well_info.csv')
rearray_back = {rearray[i]: i for i in rearray}
well_info['FC_well'] = well_info['platewell'].apply(lambda pw: rearray_back.get(pw, None))
wells_use = well_info[pd.notnull(well_info['FC_well'])][['platewell', 'FC_well', 'strain', 'plate']]
print(len(wells_use))
wells_use.iloc[:3]
clone_plates = ['FCA', 'FCB']
datas = {cp: get_plate_data('../../Data/Ploidy/Clone_plates_10K_8_25_20/' + cp + '_B1_10K/Specimen_001_') for cp in clone_plates}
redo_dat = get_plate_data('../../Data/Ploidy/FC_redos_9_3_20/Specimen_001_', cols=2)
# renames to the FC well (from the redo plate well).
# Those that were repeated as internal checks are marked as _extra here and are not plotted below
redo_renamer = {
'B01': 'B01', 'B02': 'A02',
'C01': 'C01', 'C02': 'A03',
'D01': 'D01', 'D02': 'A04',
'E01': 'E01_extra', 'E02': 'C09',
'F01': 'F01_extra', 'F02': 'C09_2',
'G01': 'B02', 'G02': 'C11',
'H01': 'B04_extra', 'H02': 'C04_extra',
}
redo_back = {redo_renamer[i]: i for i in redo_renamer}
plate2env = {'P1': r'YPD 30$\degree$C', 'P2': r'SC 30$\degree$C', 'P3': r'SC 37$\degree$C'}
strains_for_print = {'a': '$MATa$', 'diploid': 'Diploid', 'alpha': r'$MAT\alpha$'}
mybins = [i*3000 for i in range(100)]
for strain in ['a', 'alpha', 'diploid']:
nrows = {'a': 6, 'diploid': 6, 'alpha': 3}[strain]
fig, subps = pl.subplots(nrows, 6, figsize=(7.5, nrows*1), dpi=200, sharex=True, sharey=True)
subs = [subps[i][j] for i in range(nrows) for j in range(6)]
td = wells_use[wells_use['strain']==strain]
c = 0
for cp in ['FCB', 'FCA']:
s = 0
for j, row in td.iterrows():
well = row['FC_well']
sub = subs[s]
if well in redo_back:
if cp=='FCA':
sub.hist(redo_dat[redo_back[well]]['FITC-A'], bins=mybins, histtype=u'step', edgecolor=cbs[3], lw=0.8, alpha=0.7, density=True, zorder=1)
elif well+'_2' in redo_back: # only relevant for C9, which had two clones in the redos
sub.hist(redo_dat[redo_back[well+'_2']]['FITC-A'], bins=mybins, histtype=u'step', edgecolor=cbs[2], lw=0.8, alpha=0.7, density=True, zorder=1)
if s % 6 == 0:
sub.set_ylabel(plate2env[row['plate']], rotation='horizontal', ha='right')
s += 1
if len(datas[cp][well]['FITC-A']) > 1000:
sub.hist(datas[cp][well]['FITC-A'], bins=mybins, histtype=u'step', edgecolor=cbs[c+2], lw=0.8, alpha=0.7, density=True, zorder=1)
else:
pass # old (from when I was checking which ones I needed to redo)
#print(rearray[well], cp, len(datas[cp][well]['FITC-A']))
sub.set_xlim([0,250000])
sub.set_ylim([0, 0.00005])
sub.add_patch(patches.Rectangle((0,0),50000,1,facecolor='#DDDDDD', zorder=-1))
sub.set_yticks([])
sub.set_xticks([])
sub.set_title(rearray[well], fontsize=6, y=0.6, x=0.7)
c += 1
sns.despine()
fig.savefig('../../Output/Figs/supp_figs/ploidy/'+strain+'_clones_10K_ploidy.svg', background='transparent', bbox_inches='tight', pad_inches=0.1)
odd = ['P1B03', 'P1B04', 'P1B11', 'P1H11', 'P3F11']
nrows = 15
fig, subps = pl.subplots(nrows, 6, figsize=(7.5, 10), dpi=200, sharex=True, sharey=True)
subs = [subps[i][j] for i in range(nrows) for j in range(6)]
s = 0
for strain in ['a', 'alpha', 'diploid']:
td = wells_use[wells_use['strain']==strain]
for j, row in td.iterrows():
sub = subs[s]
if s % 6 == 0:
sub.set_ylabel(plate2env[row['plate']]+'\n'+strains_for_print[strain], rotation='horizontal', ha='right', va='center')
s += 1
c = 0
for cp in ['FCB', 'FCA']:
well = row['FC_well']
if well in redo_back:
if cp=='FCA':
sub.hist(redo_dat[redo_back[well]]['FITC-A'], bins=mybins, histtype=u'step', edgecolor=cbs[3], lw=0.8, alpha=0.7, density=True, zorder=1)
elif well+'_2' in redo_back: # only relevant for C9, which had two clones in the redos
sub.hist(redo_dat[redo_back[well+'_2']]['FITC-A'], bins=mybins, histtype=u'step', edgecolor=cbs[2], lw=0.8, alpha=0.7, density=True, zorder=1)
if len(datas[cp][well]['FITC-A']) > 1000:
sub.hist(datas[cp][well]['FITC-A'], bins=mybins, histtype=u'step', edgecolor=cbs[c+2], lw=0.8, alpha=0.7, density=True, zorder=1)
sub.set_xlim([0,250000])
sub.set_ylim([0, 0.00005])
sub.add_patch(patches.Rectangle((0,0),50000,1,facecolor='#DDDDDD', zorder=-1))
sub.set_yticks([])
sub.set_xticks([])
if rearray[well] in odd:
sub.annotate('* ' + rearray[well], fontsize=6, xy=(0.7, 0.6), xycoords='axes fraction', ha='center')
else:
sub.annotate(rearray[well], fontsize=6, xy=(0.7, 0.6), xycoords='axes fraction', ha='center')
c += 1
sns.despine()
fig.savefig('../../FINAL_FIGURES/supp/Fig8Supp1.pdf', background='transparent', bbox_inches='tight', pad_inches=0.1)
| Other_and_plotting/.ipynb_checkpoints/PLOT_ploidy-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 21.2 Classification
#
#
# ### Class Objectives
#
# * Will understand how to calculate and apply the fundamental classification algorithms: logistic regression, SVM, KNN, decision trees, and random forests.
#
# * Will understand how to quantify and validate classification models including calculating a classification report.
#
# * Will understand how to apply `GridSearchCV` to hyper tune model parameters.
#
# # Instructor Turn Activity 1 Logistic Regression
#
# Logistic Regression is a statistical method for predicting binary outcomes from data.
#
# Examples of this are "yes" vs "no" or "young" vs "old".
#
# These are categories that translate to probability of being a 0 or a 1
# We can calculate logistic regression by adding an activation function as the final step to our linear model.
#
# This converts the linear regression output to a probability.
#
# * Logistic Regression is a statistical method for predicting binary outcomes from data. With linear regression, our linear model may provide a numerical output such as age. With logistic regression, the numerical value for age could be translated to a probability between 0 and 1. This discrete output could then be labeled as "young" vs "old".
#
# 
#
# * Logistic regression is calculated by applying an activation function as the final step to our linear model. This transforms a numerical range to a bounded probability between 0 and 1.
#
# 
#
# * We can use logistic regression to predict which category or class a new data point should have.
#
# 
# 
# 
# 
#
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
# Generate some data
# * The `make_blobs` function to generate two different groups (classes) of data. We can then apply logistic regression to determine if new data points belong to the purple group or the yellow group.
#
# 
#
# * We create our model using the `LogisticRegression` class from Sklearn.
#
# 
#
# * Then we fit (train) the model using our training data.
#
# 
#
# * And validate it using the test data.
#
# 
#
# * And finally, we can make predictions.
#
# 
#
# 
# +
# make_blobs Generate isotropic Gaussian blobs for clustering.
from sklearn.datasets import make_blobs
X, y = make_blobs(centers=2, random_state=42)
print(f"Labels: {y[:10]}")
print(f"Data: {X[:10]}")
# -
# Visualizing both classes
plt.scatter(X[:, 0], X[:, 1], c=y)
# Split our data into training and testing
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1, stratify=y)
# -
# Create a Logistic Regression Model
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier
# Fit (train) or model using the training data
classifier.fit(X_train, y_train)
# Validate the model using the test data
print(f"Training Data Score: {classifier.score(X_train, y_train)}")
print(f"Testing Data Score: {classifier.score(X_test, y_test)}")
# Make predictions
# Generate a new data point (the red circle)
import numpy as np
new_data = np.array([[-0, 6]])
plt.scatter(X[:, 0], X[:, 1], c=y)
plt.scatter(new_data[0, 0], new_data[0, 1], c="r", marker="o", s=100)
# Predict the class (purple or yellow) of the new data point
predictions = classifier.predict(new_data)
print("Classes are either 0 (purple) or 1 (yellow)")
print(f"The new point was classified as: {predictions}")
predictions = classifier.predict(X_test)
pd.DataFrame({"Prediction": predictions, "Actual": y_test})
# # Students Turn Activity 2 Voice Gender Recognition
#
# * In this activity, you will apply logistic regression to predict the gender of a voice using acoustic properties of the voice and speech.
#
# ## Instructions
#
# * Split your data into training and testing data.
#
# * Create a logistic regression model with sklearn.
#
# * Fit the model to the training data.
#
# * Make 10 predictions and compare those to the testing data labels.
#
# * Compute the R2 score for the training and testing data separately.
#
# - - -
#
# # Voice Gender
# Gender Recognition by Voice and Speech Analysis
#
# This database was created to identify a voice as male or female, based upon acoustic properties of the voice and speech. The dataset consists of 3,168 recorded voice samples, collected from male and female speakers. The voice samples are pre-processed by acoustic analysis in R using the seewave and tuneR packages, with an analyzed frequency range of 0hz-280hz (human vocal range).
#
# ## The Dataset
# The following acoustic properties of each voice are measured and included within the CSV:
#
# * meanfreq: mean frequency (in kHz)
# * sd: standard deviation of frequency
# * median: median frequency (in kHz)
# * Q25: first quantile (in kHz)
# * Q75: third quantile (in kHz)
# * IQR: interquantile range (in kHz)
# * skew: skewness (see note in specprop description)
# * kurt: kurtosis (see note in specprop description)
# * sp.ent: spectral entropy
# * sfm: spectral flatness
# * mode: mode frequency
# * centroid: frequency centroid (see specprop)
# * peakf: peak frequency (frequency with highest energy)
# * meanfun: average of fundamental frequency measured across acoustic signal
# * minfun: minimum fundamental frequency measured across acoustic signal
# * maxfun: maximum fundamental frequency measured across acoustic signal
# * meandom: average of dominant frequency measured across acoustic signal
# * mindom: minimum of dominant frequency measured across acoustic signal
# * maxdom: maximum of dominant frequency measured across acoustic signal
# * dfrange: range of dominant frequency measured across acoustic signal
# * modindx: modulation index. Calculated as the accumulated absolute difference between adjacent measurements of fundamental frequencies divided by the frequency range
# * label: male or female
# * Logistic regression is used to predict categories or labels.
#
#
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import os
voice = pd.read_csv(os.path.join('Resources', 'voice.csv'))
voice.head()
# +
# Assign X (data) and y (target)
X = voice.drop("label", axis=1)
y = voice["label"]
print(X.shape, y.shape)
print(X)
print(y)
# -
Split our data into training and testing
# +
# Split the data using train_test_split
# YOUR CODE HERE
# -
Create a Logistic Regression Model
# +
# Create a logistic regression model
# YOUR CODE HERE
# -
Fit (train) or model using the training data
# +
# Fit the model to the data
# YOUR CODE HERE
# -
Validate the model using the test data
# +
# Print the r2 score for the test data
# YOUR CODE HERE
# -
Make predictions
# +
# Make predictions using the X_test and y_test data
# Print at least 10 predictions vs their actual labels
# YOUR CODE HERE
# -
# # Explanations:
# * The `stratify` parameter in `train_test_split` to obtain a representative sample of each category in our test data.
# 
#
# * We will perform logistic regression to our dataset in order to predict the label `male` or `female`.
#
# 
# # Instructor Turn Activity 3 Trees
# * Decision Trees encode a series of True/False questions that can be interpreted as if-else statements
#
# 
#
# 
#
# * Decision trees have a depth: the number of `if-else` statements encountered before making a decision.
#
# * Decision trees can become very complex and very deep, depending on how many questions have to be answered. Deep and complex trees tend to overfit to the data and do not generalize well.
#
# 
#
# * Random Forests:
#
# * Instead of one large, complex tree, you use many small and simple decision trees and average their outputs.
#
# * These simple trees are created by randomly sampling the data and creating a decision tree for only that small portion of data. This is known as a **weak classifier** because it is only trained on a small piece of the original data and by itself is only slightly better than a random guess. However, many "slightly better than average" small decision trees can be combined to create a **strong classifier**, which has much better decision making power.
#
# * Another benefit to this algorithm is that it is robust against overfitting. This is because all of those weak classifiers are trained on different pieces of the data.
#
# 
#
# * Each node in the tree attempts to split the data based on some criteria of the input data. The top of the tree will be the decision point that makes the biggest split. Each sub-node makes a finer and finer grain decision as the depth increases.
#
# 
#
# * Point out that the training phase of the decision tree algorithm learns which features best split the data.
#
# * Explain a byproduct of the Random Forest algorithm is a ranking of feature importance (i.e. which features have the most impact on the decision).
#
# * Scikit-Learn provides an attribute called `feature_importances_`, where you can see which features were the most significant.
#
# ```python
# sorted(zip(rf.feature_importances_, iris.feature_names), reverse=True)
# ```
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import load_iris
# Load the Iris Dataset
iris = load_iris()
print(iris.DESCR)
# Create a random forest classifier
rf = RandomForestClassifier(n_estimators=200)
rf = rf.fit(iris.data, iris.target)
rf.score(iris.data, iris.target)
# Random Forests in sklearn will automatically calculate feature importance
importances = rf.feature_importances_
importances
# We can sort the features by their importance
sorted(zip(rf.feature_importances_, iris.feature_names), reverse=True)
# # Students Turn Activity # Trees
#
# * In this activity, you will compare the performance of a decision tree to a random forest classifier using the Pima Diabetes DataSet.
#
# ## Instructions
#
# * Use the Pima Diabetes DataSet and train a decision tree classifier to predict the diabetes label (positive or negative). Print the score for the trained model using the test data.
#
# * Repeat the exercise using a Random Forest Classifier with SciKit-Learn. You will need to investigate the SciKit-Learn documentation to determine how to build and train this model.
#
# * Experiment with different numbers of estimators in your random forest model. Try different values between 100 and 1000 and compare the scores.
#
# - - -
#
#
from sklearn import tree
import pandas as pd
import os
df = pd.read_csv(os.path.join("Resources", "diabetes.csv"))
df.head()
target = df["Outcome"]
target_names = ["negative", "positive"]
data = df.drop("Outcome", axis=1)
feature_names = data.columns
data.head()
# +
# Split the data using train_test_split
# YOUR CODE HERE
# +
# Create a Decision Tree Classifier
# YOUR CODE HERE
# +
# Fit the classifier to the data
# YOUR CODE HERE
# +
# Calculate the R2 score for the test data
# YOUR CODE HERE
# +
# Bonus
# Create, fit, and score a Random Forest Classifier
# YOUR CODE HERE
# -
# * The accuracy improves slightly when using a random forest classifier. Change the number of estimators in the random forest model and re-compute the score to show how it changes.
#
# 
#
# # Instructor Turn Activity KNN
# * The KNN algorithm is a simple, yet robust machine learning algorithm. It can be used for both regression and classification. However, it is typically used for classification.
#
# * Walk through the examples provided and show how K changes the classification. we use odd numbers for k so that there isn't a tie between neighboring points.
#
# 
#
# 
#
# 
#
# 
#
# * Finally, the `k` for KNN is often calculated computationally with a loop.
#
# * Point out that the best `k` value for this dataset is where the score is both accurate and has started to stabilize.
#
# 
#
# 
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
iris = load_iris()
print(iris.DESCR)
X = iris.data
y = iris.target
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42, stratify=y)
# +
from sklearn.preprocessing import StandardScaler
# Create a StandardScater model and fit it to the training data
X_scaler = StandardScaler().fit(X_train.reshape(-1, 1))
# +
# Transform the training and testing data using the X_scaler and y_scaler models
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# -
# # K Nearest Neighbors
# +
# Loop through different k values to see which has the highest accuracy
# Note: We only use odd numbers because we don't want any ties
train_scores = []
test_scores = []
for k in range(1, 20, 2):
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X_train_scaled, y_train)
train_score = knn.score(X_train_scaled, y_train)
test_score = knn.score(X_test_scaled, y_test)
train_scores.append(train_score)
test_scores.append(test_score)
print(f"k: {k}, Train/Test Score: {train_score:.3f}/{test_score:.3f}")
plt.plot(range(1, 20, 2), train_scores, marker='o', label='training')
plt.plot(range(1, 20, 2), test_scores, marker="x", label='testing')
plt.legend()
plt.xlabel("k neighbors")
plt.ylabel("Testing accuracy Score")
plt.show()
# -
# Note that k: 9 provides the best accuracy where the classifier starts to stablize
knn = KNeighborsClassifier(n_neighbors=9)
knn.fit(X_train, y_train)
print('k=9 Test Acc: %.3f' % knn.score(X_test, y_test))
new_iris_data = [[4.3, 3.2, 1.3, 0.2]]
predicted_class = knn.predict(new_iris_data)
print(predicted_class)
# # Students Turn KNN Activity
#
# * In this activity, you will determine the best `k` value in KNN to predict diabetes for the Pima Diabetes DataSet.
#
# ## Instructions
#
# * Calculate the Train and Test scores for `k` ranging from 1 to 20. Use only odd numbers for the k values.
#
# * Plot the `k` values for both the train and test data to determine where the best combination of scores occur. This point will be the optimal `k` value for your model.
#
# * Re-train your model using the `k` value that you found to have the best scores. Print the score for this value.
#
# - - -
#
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
import pandas as pd
import os
df = pd.read_csv(os.path.join("Resources", "diabetes 2.csv"))
df.head()
target = df["Outcome"]
target_names = ["negative", "positive"]
data = df.drop("Outcome", axis=1)
feature_names = data.columns
data.head()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data, target, random_state=42)
# +
# Loop through different k values to see which has the highest accuracy
# Note: We only use odd numbers because we don't want any ties
# YOUR CODE HERE
plt.plot(range(1, 20, 2), train_scores, marker='o')
plt.plot(range(1, 20, 2), test_scores, marker="x")
plt.xlabel("k neighbors")
plt.ylabel("Testing accuracy Score")
plt.show()
# +
# Choose the best k from above and re-fit the KNN Classifier using that k value.
# print the score for the test data
# YOUR CODE HERE
# -
# * For this activity, `K=13` seems to be the best combination of both the train and test scores.
#
# 
#
# * Ask students for any additional questions before moving on.
#
# # Instructor Turn Activity 7
#
# ### 12. Instructor Do: SVM (0:10)
#
# * The goal of a linear classifier is to find a line that separates two groups of classes. However, there may be many options for choosing this line and each boundary could result in misclassification of new data.
#
# 
#
# 
#
# * SVM try to find a hyperplane that maximizes the boundaries between groups. This is like building a virtual wall between groups where you want the wall to be as thick as possible.
#
# 
#
#
# * There are different kernels available for the SVM model in SciKit-Learn, but we are going to use the linear model in this example.
#
# 
#
# * The decision boundaries for the trained model. The algorithm shows how it maximized the boundaries between the two groups.
#
# 
#
# * Next, show an example of "real" data where the boundaries are overlapping. In this case, the svm algorithm will "soften" the margins and allow some of the data points to cross over the margin boundaries in order to obtain a fit.
#
# 
#
# * Generate a classification report to quantify and validate the model performance.
#
# 
#
# * precision and recall to deep dive into the meaning behind each score.
#
# 
# 
# 
# 
# 
# 
# 
# 
# 
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
style.use("ggplot")
# from matplotlib import rcParams
# rcParams['figure.figsize'] = 10, 8
# -
from sklearn.datasets.samples_generator import make_blobs
X, y = make_blobs(n_samples=40, centers=2, random_state=42, cluster_std=1.25)
plt.scatter(X[:, 0], X[:, 1], c=y, s=100, cmap="bwr");
plt.show()
# Support vector machine linear classifier
from sklearn.svm import SVC
model = SVC(kernel='linear')
model.fit(X, y)
# +
# WARNING! BOILERPLATE CODE HERE!
# Plot the decision boundaries
x_min = X[:, 0].min()
x_max = X[:, 0].max()
y_min = X[:, 1].min()
y_max = X[:, 1].max()
XX, YY = np.mgrid[x_min:x_max, y_min:y_max]
Z = model.decision_function(np.c_[XX.ravel(), YY.ravel()])
# Put the result into a color plot
Z = Z.reshape(XX.shape)
# plt.pcolormesh(XX, YY, Z > 0, cmap=plt.cm.Paired)
plt.contour(XX, YY, Z, colors=['k', 'k', 'k'],
linestyles=['--', '-', '--'], levels=[-.5, 0, .5])
plt.scatter(X[:, 0], X[:, 1], c=y, cmap='bwr', edgecolor='k', s=100)
plt.show()
# -
# # Validation
X, y = make_blobs(n_samples=100, centers=2, random_state=0, cluster_std=.95)
plt.scatter(X[:, 0], X[:, 1], c=y, s=100, cmap="bwr");
plt.show()
# Split data into training and testing
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Fit to the training data and validate with the test data
model = SVC(kernel='linear')
model.fit(X_train, y_train)
predictions = model.predict(X_test)
# +
# Plot the decision boundaries
x_min = X[:, 0].min()
x_max = X[:, 0].max()
y_min = X[:, 1].min()
y_max = X[:, 1].max()
XX, YY = np.mgrid[x_min:x_max, y_min:y_max]
Z = model.decision_function(np.c_[XX.ravel(), YY.ravel()])
# Put the result into a color plot
Z = Z.reshape(XX.shape)
# plt.pcolormesh(XX, YY, Z > 0, cmap=plt.cm.Paired)
plt.contour(XX, YY, Z, colors=['k', 'k', 'k'],
linestyles=['--', '-', '--'], levels=[-.5, 0, .5])
plt.scatter(X[:, 0], X[:, 1], c=y, cmap='bwr', edgecolor='k', s=100)
plt.show()
# -
# Calculate classification report
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions,
target_names=["blue", "red"]))
# # Students Turn Activity 8 SVM
#
# * In this activity, apply a support vector machine classifier predict diabetes for the Pima Diabetes DataSet.
#
# ## Instructions
#
# * Import a support vector machine linear classifier and fit the model to the data.
#
# * Compute the classification report for this model using the test data.
#
# - - -
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
import pandas as pd
import numpy as np
import os
df = pd.read_csv(os.path.join("Resources", "diabetes.csv"))
df.head()
target = df["Outcome"]
target_names = ["negative", "positive"]
data = df.drop("Outcome", axis=1)
feature_names = data.columns
data.head()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data, target, random_state=42)
# +
# Create a support vector machine linear classifer and fit it to the training data
# YOUR CODE HERE
# +
# Print the model score using the test data
# YOUR CODE HERE
# +
# Calculate the classification report
# YOUR CODE HERE
# -
# * The F1 scores indicate that this model is slightly more accurate at predicting negative cases of diabetes than positive cases.
#
# 
# # Instructor Turn Activity 9 GridSearch
# * The code for hyperparameter tuning with `GridSearchCV`.
#
# * The SVM model to highlight the different features available for the model. Each of these features can be adjusted and tweaked to improve model performance.
#
# 
#
# * In machine learning, there are few if any general rules on how to adjust these parameters. Instead, machine learning practitioners often use a brute force approach where they try different combinations of values to see which has the best performance. This is known as `hyperparameter tuning`
#
# * To simplify the hyperparameter tuning process, SciKit-Learn provides a tool called `GridSearchCV`. This class is known as a `meta-estimator`. That is, it takes a model and a dictionary of parameter settings and tests all combinations of parameter settings to see which settings have the best performance.
#
# 
#
# 
#
# 
# 
# 
#
# * Once the model has been trained, the best parameters can be accessed through the `best_params_` attribute.
#
# 
#
# * Similarly, the best score can be accessed through the `best_score_` attribute.
#
# 
#
# * The grid meta-estimator basically wraps the original model, so you can access the model functions like `predict`.
#
# 
#
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
style.use("ggplot")
# from matplotlib import rcParams
# rcParams['figure.figsize'] = 10, 8
# -
from sklearn.datasets.samples_generator import make_blobs
X, y = make_blobs(n_samples=100, centers=2, random_state=0, cluster_std=.95)
plt.scatter(X[:, 0], X[:, 1], c=y, s=100, cmap="bwr");
plt.show()
# Split data into training and testing
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Create the SVC Model
from sklearn.svm import SVC
model = SVC(kernel='linear')
model
# Create the GridSearch estimator along with a parameter object containing the values to adjust
from sklearn.model_selection import GridSearchCV
param_grid = {'C': [1, 5, 10, 50],
'gamma': [0.0001, 0.0005, 0.001, 0.005]}
grid = GridSearchCV(model, param_grid, verbose=3)
# Fit the model using the grid search estimator.
# This will take the SVC model and try each combination of parameters
grid.fit(X_train, y_train)
# List the best parameters for this dataset
print(grid.best_params_)
# List the best score
print(grid)
print(grid.best_score_)
# Make predictions with the hypertuned model
predictions = grid.predict(X_test)
# Calculate classification report
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions,
target_names=["blue", "red"]))
# # Student Turn Activity 10 Grid Search and Hyper-Parameter Tuning
#
# * In this activity, you will revisit the SVM activity for the Pima Diabetes DataSet and apply `GridSearchCV` to tune the model parameters.
#
# ## Instructions
#
# * Use the starter code provided and apply `GridSearchCV` to the model. Change the `C` and `gamma` parameters.
#
# * For `C`, use the following list of settings: `[1, 5, 10]`.
#
# * For `gamma`, use the following list of settings: `[0.0001, 0.001, 0.01]`.
#
# * Print the best parameters and best score for the tuned model.
#
# * Calculate predictions using the `X_test` data and print the classification report.
#
# - - -
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
import pandas as pd
import numpy as np
import os
df = pd.read_csv(os.path.join("Resources", "diabetes.csv"))
df.head()
target = df["Outcome"]
target_names = ["negative", "positive"]
data = df.drop("Outcome", axis=1)
feature_names = data.columns
data.head()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data, target, random_state=42)
# Support vector machine linear classifier
from sklearn.svm import SVC
model = SVC(kernel='linear')
# +
# Create the GridSearch estimator along with a parameter object containing the values to adjust
# Try adjusting `C` with values of 1, 5, and 10. Adjust `gamma` using .0001, 0.001, and 0.01
# YOUR CODE HERE
# +
# Fit the model using the grid search estimator.
# This will take the SVC model and try each combination of parameters
# YOUR CODE HERE
# +
# List the best parameters for this dataset
# YOUR CODE HERE
# +
# List the best score
# YOUR CODE HERE
# +
# Make predictions with the hypertuned model
# YOUR CODE HERE
# +
# Calculate classification report
# YOUR CODE HERE
# -
# * The Grid Search tested our model with 27 different combinations of parameters and data.
# * Applying GridSearch saves us considerable time vs manually changing these values ourselves.
#
# * Knowing which parameters to tune and which values to use comes from both experience and Sklearn's documentation for their algorithms.
#
# * By simply tuning two of our hyperparameters, the model score increased from 0.729 to an accuracy score of 0.774!
#
# 
#
# - - -
| ML Samples/Day2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ox)
# language: python
# name: ox
# ---
# # Save/load street network models to/from disk
#
# Author: [<NAME>](https://geoffboeing.com/)
#
# - [Overview of OSMnx](http://geoffboeing.com/2016/11/osmnx-python-street-networks/)
# - [GitHub repo](https://github.com/gboeing/osmnx)
# - [Examples, demos, tutorials](https://github.com/gboeing/osmnx-examples)
# - [Documentation](https://osmnx.readthedocs.io/en/stable/)
# - [Journal article/citation](http://geoffboeing.com/publications/osmnx-complex-street-networks/)
#
# This notebook demonstrates how to save networks to disk as shapefiles, geopackages, graphml, and xml, and how to load an OSMnx-created network from a graphml file.
import osmnx as ox
# %matplotlib inline
ox.config(log_console=True)
ox.__version__
# get a network
place = 'Piedmont, California, USA'
G = ox.graph_from_place(place, network_type='drive')
# ## Shapefiles and GeoPackages for GIS
# save graph as a shapefile
ox.save_graph_shapefile(G, filepath='./data/piedmont')
# save graph as a geopackage
ox.save_graph_geopackage(G, filepath='./data/piedmont.gpkg')
# ## GraphML files for saving network and preserving topological detail
# save/load graph as a graphml file: this is the best way to save your model
# for subsequent work later
filepath = './data/piedmont.graphml'
ox.save_graphml(G, filepath)
G = ox.load_graphml(filepath)
# if you want to work with your model in gephi, use gephi compatibility mode
ox.save_graphml(G, filepath=filepath, gephi=True)
# ## SVG file to work with in Adobe Illustrator
# save street network as SVG
fig, ax = ox.plot_graph(G, show=False, save=True, close=True, filepath='./images/piedmont.svg')
# ## Save points of interest or building footprints
# get all "amenities" and save as a geopackage via geopandas
gdf = ox.geometries_from_place(place, tags={'amenity':True})
gdf = gdf.apply(lambda c: c.astype(str) if c.name != 'geometry' else c, axis=0)
gdf.to_file('./data/pois.gpkg', driver='GPKG')
# get all building footprints and save as a geopackage via geopandas
gdf = ox.geometries_from_place(place, tags={'building':True})
gdf = gdf.apply(lambda c: c.astype(str) if c.name != 'geometry' else c, axis=0)
gdf.to_file('./data/building_footprints.gpkg', driver='GPKG')
# ## Save .osm XML files
#
# To save your graph to disk as a .osm formatted XML file, ensure that you created the graph with `ox.settings.all_oneway=True` for `save_graph_xml` to work properly. See docstring for details.
#
# To save/load full-featured OSMnx graphs to/from disk for later use, use the `save_graphml` and `load_graphml` functions instead.
# save graph to disk as .osm xml file
ox.config(all_oneway=True, log_console=True, use_cache=True)
G = ox.graph_from_place('Piedmont, California, USA', network_type='drive')
ox.save_graph_xml(G, filepath='./data/piedmont.osm')
| notebooks/05-save-load-networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
# # Load Data and Convert to Parquet
# Load csv 1
df1 = pd.read_csv('../test-data/weather.20160201.csv')
df1.info()
# Load csv 2
df2 = pd.read_csv('../test-data/weather.20160301.csv')
df2.info()
# Union both
df = df1.append(df2)
df.info()
# Save to Parquet
df.to_parquet('../result/weather.parquet')
# Read from Parquet (just for example)
df = pd.read_parquet('../result/weather.parquet')
df.info()
# # Hottest Day
# Get Highest Temperature
max_temperature = df['ScreenTemperature'].max()
print(f'Max Temperature: {max_temperature}')
# Get record with highest temperature
df_hottest = df[ df['ScreenTemperature'] == max_temperature ]
# Hottest Day
hottest_date = df_hottest['ObservationDate'].to_numpy()[0][:10]
print(f'Hottest Day: {hottest_date}')
# Region & Country
region = df_hottest['Region'].to_numpy()[0]
country = df_hottest['Country'].to_numpy()[0]
print(f'Region: {region}')
print(f'Country: {country}')
# ### Questions
print('Q: Which date was the hottest day?')
print(f'R: {hottest_date}')
print()
print('Q: What was the temperature on that day?')
print(f'R: {max_temperature}')
print()
print('Q: In which region was the hottest day?')
print(f'R: {region}')
| notebook/weather-with-pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model Prediction using SAVIME and PYSAVIME
#
# __The goal of this notebook is to introduce the model execution and prediction feature, using the Savime system and the PySavime access API. The models and data used will be the ones created in the first notebook: Part-01.
#
# Check the variables `savime_host` and `savime_port`, which point to the host and port where Savime is listening to, respectively. We assume Savime is initialized, and Tfx is listening to the port 8501.
# +
# %load_ext autoreload
# %reload_ext autoreload
# %autoreload 2
import os
import sys
if not 'notebooks' in os.listdir('.'):
current_dir = os.path.abspath(os.getcwd())
parent_dir = os.path.dirname(current_dir)
os.chdir(parent_dir)
# We define the data file path : a json storing information about
# the x and y partitions used in part-01.
data_file = 'saved_models_elastic_net/data.json'
# Configuring host and port where Savime is listening to
savime_host = '127.0.0.1'
savime_port = 65000
# -
# Abaixo é realizada a inclusão das módulos necessários para execução deste notebook. Atente para a inclusão da biblioteca pysame.
# +
import json
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
# Savime imports
import pysavime
from pysavime.util.converter import DataVariableBlockConverter
from pysavime.util.data_variable import DataVariableBlockOps
# Importing Python prediction client for Tfx
from src.predictor_consumer import PredictionConsumer
from src.util import read_numpy_array_from_disk, export_numpy_array_to_c_array
# Ommiting tensorflow warnings
import tensorflow as tf
tf.get_logger().setLevel('ERROR')
# -
# Here we load the data generated on part-01.
# +
# Reading input json data file
with open(data_file, 'r') as _in:
data = json.load(_in)
# Reading x and y arrays
output_dir = data['output_dir']
# Directories Definition
x_file_path = os.path.join(output_dir, data['x_file_name'])
y_file_path = os.path.join(output_dir, data['y_file_name'])
x_c_file_path = os.path.join(output_dir, 'x_data')
y_c_file_path = os.path.join(output_dir, 'y_data')
# Converting generated data to format compatible with Savime
x_array = read_numpy_array_from_disk(x_file_path)
y_array = read_numpy_array_from_disk(y_file_path)
export_numpy_array_to_c_array(x_array, 'float64', x_c_file_path)
export_numpy_array_to_c_array(y_array, 'float64', y_c_file_path)
print('X values:\n', x_array)
print('Y values:\n', y_array)
# -
# Now we define the queries which we will run in Savime to build the necessary structures to create and load our datasets: CREATE_DATASET, CREATE_TAR and LOAD_SUBTAR.
# +
# Definition of the dataset to be used:
num_observations = len(x_array)
num_features = x_array.shape[1]
y_num_columns = y_array.shape[1] if len(y_array.shape) == 2 else 1
x_dataset = pysavime.define.file_dataset('x', x_c_file_path, 'double', length=num_features)
y_dataset = pysavime.define.file_dataset('y', y_c_file_path, 'double', length=y_num_columns)
# Tar Definition
index = pysavime.define.implicit_tar_dimension('index', 'int32', 1, num_observations)
x = pysavime.define.tar_attribute('x', 'double', num_features)
y = pysavime.define.tar_attribute('y', 'double', y_num_columns)
tar = pysavime.define.tar('target-function', [index], [x, y])
# Definition of subtar loading commands
subtar_index = pysavime.define.ordered_subtar_dimension(index, 1, num_observations)
subtar_x = pysavime.define.subtar_attribute(x, x_dataset)
subtar_y = pysavime.define.subtar_attribute(y, y_dataset)
subtar = pysavime.define.subtar(tar, [subtar_index], [subtar_x, subtar_y])
# The defined commands are:
print(x_dataset.create_query_str(), y_dataset.create_query_str(), sep='\n')
print(tar.create_query_str())
print(subtar.load_query_str())
# -
# Finally, we run the previously defined commands on savime
#
# 1. We open and close the connection using Savime ('with' context)
# 2. Creation of a command execution object, attached to the opened conection
# 3.
# 1. Dataset Creation
# 2. Subtar Creation
# 3. Loading the datasets into the subtar
with pysavime.Client(host=savime_host, port=savime_port) as client:
client.execute(pysavime.operator.create(x_dataset))
client.execute(pysavime.operator.create(y_dataset))
client.execute(pysavime.operator.create(tar))
client.execute(pysavime.operator.load(subtar))
# Next, for each saved model, we get the prediction's mean squared error on the data domain. To do so, we
# execute the following steps:
#
# 1. Register the model on the system: `pysavime.operator.register_model`
# 2. Execute the predictive query: `pysavime.operator.predict`
# 3. We calculate the squared difference between the query output and the true y value:
# `pysavime.operator.derive`
# 4. From this value, we calculate the mean squared error: `pysavime.operator.aggregate`
# +
mse = {}
registered_models = data['iid']
with pysavime.Client(host=savime_host, port=savime_port) as client:
# dim_spec specifies the size of the predictive query window.
# It's a list of pairs, in which the first element specifies the dimension,
# and the second element specifies the number of observations
dim_spec = [(index.name, num_observations)]
for model_name, i in registered_models.items():
# A model is registered in Savime, i.e., we associate it with a Tar, identify what is the input attribute
# and the format of the multidimensional input array. In this case, we are sending the complete
# observations array, but it's also possible to predict only a section of it
# Register the model that we will use
register_cmd = pysavime.operator.register_model(model_name=model_name, model_tar=tar.name, input_attribute=x.name,
dim_specification=dim_spec)
client.execute(register_cmd)
# Calculate the mean squared error
predict_cmd = pysavime.operator.predict(tar=tar.name, model_name=model_name, input_attribute=x.name)
derive_cmd = pysavime.operator.derive(predict_cmd, 'squared_difference', '(op_result - y)^2')
aggregate_cmd = pysavime.operator.aggregate(derive_cmd, 'avg', 'squared_difference', 'mse')
print(aggregate_cmd)
mse[model_name] = client.execute(aggregate_cmd)
# -
# Next, we register each model's error and we build a data frame.
# +
print(mse[model_name])
d = {key: value[0].attrs['mse'][0][0] for key, value in mse.items()}
df = pd.DataFrame.from_dict(d, orient='index')
print(d)
# -
# Finally, we display each model's mean squared error.
# Note that model 25 exhibits the best results, since it was trained in points from
# all of the partitions of the domain.
# +
# Ordering the dataframe
df['index'] = df.index
df['index'] = df['index'].apply(lambda x: int(x.split('_')[-1]))
df = df.sort_values('index')
# Graph display
fig, ax = plt.subplots()
fig.set_size_inches(10, 6)
sns.barplot(x='index', y=0, data=df, ax=ax, color='darkblue')
# Adjusting the label
plt.xticks(rotation=90)
plt.xlabel('Models')
_ = plt.ylabel('MSE')
# -
| notebooks/.ipynb_checkpoints/pysavime-part-02-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
import pickle
import numpy as np
with open('../work_dirs/pspnet_icip_all/results.pkl', 'rb') as f:
res = pickle.load(f)
res[0]
| playground/export_res.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Air Quality Tensor
# * `<date> <location> <air pollutants> (measurement)`
# * Beijing Air Quality
# * 2,454,305 out of 2,524,536 (35,063 * 12 * 6)
# * Korea Air Quality
# * 11,270,028 out of 18,368,364 (9,478 * 323 * 6)
# * Madrid Air Quality
# * 8,036,759 out of 21,587,328 (64,248 * 24 * 14)
import csv
import time
import numpy as np
import pandas as pd
beijing_df = pd.read_csv('../Data/air_quality/BeijingAirQuality/beijing.tensor', delimiter='\t', header=None)
korea_df = pd.read_csv('../Data/air_quality/KoreaAirQuality/korea_airquality.tensor', delimiter='\t', header=None)
madrid_df = pd.read_csv('../Data/air_quality/MadridAirQuality/1hour_madrid.tensor', delimiter='\t', header=None)
def get_tensor(df):
start = time.time()
dims = df[[0,1,2]].max()+1
tensor = np.empty(dims) * np.nan
tensor.shape
for i, row in df.iterrows():
indices = [[index] for index in np.int64(np.asarray(row[:-1]))]
tensor[tuple(indices)] = np.double(row[3])
avg = []
for i in range(tensor.shape[2]):
avg.append(np.nanmean(tensor[:,:,i]))
inds = np.where(np.isnan(tensor))
for ind in zip(inds[0], inds[1], inds[2]):
tensor[ind] = avg[ind[-1]]
print(time.time() - start)
return tensor
beijing_tensor = get_tensor(beijing_df)
korea_tensor = get_tensor(korea_df)
madrid_tensor = get_tensor(madrid_df)
np.where(np.isnan(beijing_tensor))
np.where(np.isnan(korea_tensor))
np.where(np.isnan(madrid_tensor))
print(beijing_tensor.shape)
print(korea_tensor.shape)
print(madrid_tensor.shape)
# ### Hall from OLSTEC
# * `hall1-200.mat`: 144 * 176 * 200
# * `hall_144x100_frame2900-3899_pan.mat`: 144 * 100 * 1000
# +
# for sample video
from cv2 import VideoWriter, VideoWriter_fourcc, imshow
def make_video(tensor, filename, isColor=True):
start = time.time()
height = tensor.shape[1]
width = tensor.shape[2]
FPS = 24
fourcc = VideoWriter_fourcc(*'MP42')
video = VideoWriter(filename, fourcc, float(FPS), (width, height), isColor)
for frame in tensor:
video.write(np.uint8(frame))
video.release()
print('created', filename, time.time()-start)
# +
from scipy.io import loadmat
hall1 = loadmat('../Data/hall/hall1-200.mat')['XO']
hall1 = np.moveaxis(hall1, -1, 0)
hall1 = hall1.reshape(200, 144, 176, order='F')
# hall1.shape
make_video(hall1, 'hall1.avi', False)
# -
import h5py
with h5py.File('../Data/hall/hall_144x100_frame2900-3899_pan.mat', 'r') as f:
hall2 = np.array(f['X0'])
hall2 = hall2.reshape(1000, 144, 100, order='F')
make_video(hall2, 'hall2.avi', False)
| Data/0_data_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gym
import numpy as np
import matplotlib.pyplot as plt
from tqdm import tqdm, trange
import pandas as pd
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.autograd import Variable
from torch.distributions import Categorical
import warnings
warnings.simplefilter("ignore")
# %matplotlib inline
print(torch.__version__)
env = gym.make('CartPole-v1')
env.seed(1); torch.manual_seed(1);
# Hyperparameters for model
l_rate = 0.01
gamma_value = 0.99
# +
class PolicyGradient(nn.Module):
def __init__(self):
super(PolicyGradient, self).__init__()
# Define the action space and state space
self.action_space = env.action_space.n
self.state_space = env.observation_space.shape[0]
self.l1 = nn.Linear(self.state_space, 128, bias=False)
self.l2 = nn.Linear(128, self.action_space, bias=False)
self.gamma_value = gamma_value
# Episode policy and reward history
self.history_policy = Variable(torch.Tensor())
self.reward_episode = []
# Overall reward and loss history
self.history_reward = []
self.history_loss = []
def forward(self, x):
model = torch.nn.Sequential(
self.l1,
nn.Dropout(p=0.5),
nn.ReLU(),
self.l2,
nn.Softmax(dim=-1)
)
return model(x)
policy = PolicyGradient()
optimizer = optim.Adam(policy.parameters(), lr=l_rate)
# +
def choose_action(state):
# Run the policy model and choose an action based on the probabilities in state
state = torch.from_numpy(state).type(torch.FloatTensor)
state = policy(Variable(state))
c = Categorical(state)
action = c.sample()
if policy.history_policy.dim() != 0:
try:
policy.history_policy = torch.cat([policy.history_policy, c.log_prob(action)])
except:
policy.history_policy = (c.log_prob(action))
else:
policy.history_policy = (c.log_prob(action))
return action
def update_policy():
R = 0
rewards = []
# Discount future rewards back to the present using gamma
for r in policy.reward_episode[::-1]:
R = r + policy.gamma_value * R
rewards.insert(0,R)
# Scale rewards
rewards = torch.FloatTensor(rewards)
x = np.finfo(np.float32).eps
x = np.array(x)
x = torch.from_numpy(x)
rewards = (rewards - rewards.mean()) / (rewards.std() + x)
# Calculate the loss loss
loss = (torch.sum(torch.mul(policy.history_policy, Variable(rewards)).mul(-1), -1))
# Update the weights of the network
optimizer.zero_grad()
loss.backward()
optimizer.step()
#Save and intialize episode history counters
policy.history_loss.append(loss.data[0])
policy.history_reward.append(np.sum(policy.reward_episode))
policy.history_policy = Variable(torch.Tensor())
policy.reward_episode= []
def main_function(episodes):
running_total_reward = 10
for e in range(episodes):
# Reset the environment and record the starting state
state = env.reset()
done = False
for time in range(1000):
action = choose_action(state)
# Step through environment using chosen action
state, reward, done, _ = env.step(action.data.item())
# Save reward
policy.reward_episode.append(reward)
if done:
break
# Used to determine when the environment is solved.
running_total_reward = (running_total_reward * 0.99) + (time * 0.01)
update_policy()
if e % 50 == 0:
print('Episode number {}, Last length: {:5d}, Average length: {:.2f}'.format(e, time, running_total_reward))
if running_total_reward > env.spec.reward_threshold:
print("Solved! Running reward is now {} and the last episode runs to {} time steps!".format(running_total_reward, time))
break
episodes = 1000
main_function(episodes)
# -
| Chapter09/Policy Gradient Reinforcement Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Train, tune, and deploy a custom ML model using Financial Transactions Fraud Detection Algorithm from AWS Marketplace
#
#
# This solution is a deep learning based algorithm capable of training on transactional data, to identify spurious transactions.
#
#
#
# This sample notebook shows you how to train a custom ML model using Financial Transactions Fraud Detection from AWS Marketplace.
#
# > **Note**: This is a reference notebook and it cannot run unless you make changes suggested in the notebook.
#
# #### Pre-requisites:
# 1. **Note**: This notebook contains elements which render correctly in Jupyter interface. Open this notebook from an Amazon SageMaker Notebook Instance or Amazon SageMaker Studio.
# 1. Ensure that IAM role used has **AmazonSageMakerFullAccess**
# 1. Some hands-on experience using [Amazon SageMaker](https://aws.amazon.com/sagemaker/).
# 1. To use this algorithm successfully, ensure that:
# 1. Either your IAM role has these three permissions and you have authority to make AWS Marketplace subscriptions in the AWS account used:
# 1. **aws-marketplace:ViewSubscriptions**
# 1. **aws-marketplace:Unsubscribe**
# 1. **aws-marketplace:Subscribe**
# 2. or your AWS account has a subscription to For Seller to update: Financial Transactions Fraud Detection.
#
# #### Contents:
# 1. [Subscribe to the algorithm](#1.-Subscribe-to-the-algorithm)
# 1. [Prepare dataset](#2.-Prepare-dataset)
# 1. [Dataset format expected by the algorithm](#A.-Dataset-format-expected-by-the-algorithm)
# 1. [Configure and visualize train and test dataset](#B.-Configure-and-visualize-train-and-test-dataset)
# 1. [Upload datasets to Amazon S3](#C.-Upload-datasets-to-Amazon-S3)
# 1. [Train a machine learning model](#3:-Train-a-machine-learning-model)
# 1. [Set up environment](#3.1-Set-up-environment)
# 1. [Train a model](#3.2-Train-a-model)
# 1. [Deploy model and verify results](#4:-Deploy-model-and-verify-results)
# 1. [Deploy trained model](#A.-Deploy-trained-model)
# 1. [Create input payload](#B.-Create-input-payload)
# 1. [Perform real-time inference](#C.-Perform-real-time-inference)
# 1. [Visualize output](#D.-Visualize-output)
# 1. [Calculate relevant metrics](#E.-Calculate-relevant-metrics)
# 1. [Delete the endpoint](#F.-Delete-the-endpoint)
# 1. [Tune your model! (optional)](#5:-Tune-your-model!-(optional))
# 1. [Tuning Guidelines](#A.-Tuning-Guidelines)
# 1. [Define Tuning configuration](#B.-Define-Tuning-configuration)
# 1. [Run a model tuning job](#C.-Run-a-model-tuning-job)
# 1. [Perform Batch inference](#6.-Perform-Batch-inference)
# 1. [Clean-up](#7.-Clean-up)
# 1. [Delete the model](#A.-Delete-the-model)
# 1. [Unsubscribe to the listing (optional)](#B.-Unsubscribe-to-the-listing-(optional))
#
#
# #### Usage instructions
# You can run this notebook one cell at a time (By using Shift+Enter for running a cell).
# ### 1. Subscribe to the algorithm
# To subscribe to the algorithm:
# 1. Open the algorithm listing page Financial Transactions Fraud Detection
# 1. On the AWS Marketplace listing, click on **Continue to subscribe** button.
# 1. On the **Subscribe to this software** page, review and click on **"Accept Offer"** if you agree with EULA, pricing, and support terms.
# 1. Once you click on **Continue to configuration button** and then choose a **region**, you will see a **Product Arn**. This is the algorithm ARN that you need to specify while training a custom ML model. Copy the ARN corresponding to your region and specify the same in the following cell.
algo_arn='arn:aws:sagemaker:us-east-2:786796469737:algorithm/fraud-fin-tensor-1'
# ### 2. Prepare dataset
import base64
import json
import uuid
from sagemaker import ModelPackage
import sagemaker as sage
from sagemaker import get_execution_role
from sagemaker import ModelPackage
from urllib.parse import urlparse
import boto3
from IPython.display import Image
from PIL import Image as ImageEdit
import urllib.request
import numpy as np
# #### A. Dataset format expected by the algorithm
# The algorithm requires data in the format as decribed for best results
#
# 1. Input data should be in binarized format to train and learn the patterns
#
# 1. Build the binary variables judiciously and develop binary encoded columns, rather generating one-hot binary variables
#
# 1. Keep the ratio of fraudulent data to non-fraudulent more then 1:1000
#
# 1. Try to incorporate as much patterns from non-fraudulent data as possible to increase out of sample accuracy
# #### B. Configure and visualize train and test dataset
training_dataset='train.csv'
test_dataset='test.csv'
import pandas as pd
df = pd.read_csv(training_dataset)
df.head()
# #### C. Upload datasets to Amazon S3
sagemaker_session = sage.Session()
bucket=sagemaker_session.default_bucket()
bucket
training_data=sagemaker_session.upload_data(training_dataset, bucket=bucket, key_prefix='financial_fraud')
test_data=sagemaker_session.upload_data(test_dataset, bucket=bucket, key_prefix='financial_fraud')
# ## 3: Train a machine learning model
# Now that dataset is available in an accessible Amazon S3 bucket, we are ready to train a machine learning model.
# ### 3.1 Set up environment
role = get_execution_role()
output_location = 's3://{}/financial_fraud/{}'.format(bucket, 'output')
# ### 3.2 Train a model
# You can also find more information about hyperparametes in **Hyperparameters** section of Financial Transactions Fraud Detection
#Define hyperparameters
hyperparameters={"epoches":100,"batch_size":10}
# For information on creating an `Estimator` object, see [documentation](https://sagemaker.readthedocs.io/en/stable/api/training/estimators.html)
#Create an estimator object for running a training job
estimator = sage.algorithm.AlgorithmEstimator(
algorithm_arn=algo_arn,
base_job_name="financial-fraud-training",
role=role,
train_instance_count=1,
train_instance_type='ml.m5.large',
input_mode="File",
output_path=output_location,
sagemaker_session=sagemaker_session,
hyperparameters=hyperparameters,
instance_count=1,
instance_type='ml.m5.large'
)
#Run the training job.
estimator.fit({"training": training_data,"test":test_data})
# See this [blog-post](https://aws.amazon.com/blogs/machine-learning/easily-monitor-and-visualize-metrics-while-training-models-on-amazon-sagemaker/) for more information how to visualize metrics during the process. You can also open the training job from [Amazon SageMaker console](https://console.aws.amazon.com/sagemaker/home?#/jobs/) and monitor the metrics/logs in **Monitor** section.
# ### 4: Deploy model and verify results
# Now you can deploy the model for performing real-time inference.
# +
model_name='financial_fraud_inference'
content_type='text/csv'
real_time_inference_instance_type='ml.m5.large'
batch_transform_inference_instance_type='ml.m5.large'
# -
# #### A. Deploy trained model
from sagemaker.predictor import csv_serializer
predictor = estimator.deploy(1, real_time_inference_instance_type, serializer=csv_serializer)
# Once endpoint is created, you can perform real-time inference.
# #### B. Create input payload
df = pd.read_csv("infrence.csv")
print(predictor.predict(df.values))
# #### C. Perform real-time inference
file_name = "infrence.csv"
output_file_name = "result.txt"
# !aws sagemaker-runtime invoke-endpoint \
# --endpoint-name $predictor.endpoint \
# --body fileb://$file_name \
# --content-type $content_type \
# --region $sagemaker_session.boto_region_name \
# $output_file_name
# #### D. Visualize output
with open("result.txt","r") as file:
print(file.read())
# #### F. Delete the endpoint
# Now that you have successfully performed a real-time inference, you do not need the endpoint any more. you can terminate the same to avoid being charged.
predictor.delete_endpoint(delete_endpoint_config=True)
# Since this is an experiment, you do not need to run a hyperparameter tuning job. However, if you would like to see how to tune a model trained using a third-party algorithm with Amazon SageMaker's hyperparameter tuning functionality, you can run the optional tuning step.
# ### 5. Perform Batch inference
# In this section, you will perform batch inference using multiple input payloads together.
#upload the batch-transform job input files to S3
transform_input_folder = "infrence.csv"
transform_input = sagemaker_session.upload_data(transform_input_folder, key_prefix=model_name)
print("Transform input uploaded to " + transform_input)
#Run the batch-transform job
transformer = estimator.transformer(1, batch_transform_inference_instance_type)
transformer.transform(transform_input, content_type=content_type)
transformer.wait()
#output is available on following path
transformer.output_path
# ### 7. Clean-up
# #### A. Delete the model
estimator.delete_endpoint()
# #### B. Unsubscribe to the listing (optional)
# If you would like to unsubscribe to the algorithm, follow these steps. Before you cancel the subscription, ensure that you do not have any [deployable model](https://console.aws.amazon.com/sagemaker/home#/models) created from the model package or using the algorithm. Note - You can find this information by looking at the container name associated with the model.
#
# **Steps to unsubscribe to product from AWS Marketplace**:
# 1. Navigate to __Machine Learning__ tab on [__Your Software subscriptions page__](https://aws.amazon.com/marketplace/ai/library?productType=ml&ref_=mlmp_gitdemo_indust)
# 2. Locate the listing that you want to cancel the subscription for, and then choose __Cancel Subscription__ to cancel the subscription.
#
#
| Financial Transactions Fraud Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/diaboloshogunate/GoogleColabML/blob/main/CIFAR-10-3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="IsrCBkyjvqlW"
# imports
import itertools
from keras.datasets import cifar10
from keras.models import Sequential
import keras.layers as layers
from keras.preprocessing.image import ImageDataGenerator
from matplotlib import pyplot
from tensorflow import keras
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.optimizers import SGD
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 186} id="jru81UyTMddC" outputId="0ad5adb2-db70-4420-d8af-8c8ca134199b"
# load data
(trainX, trainY), (testX, testY) = cifar10.load_data()
trainY = to_categorical(trainY)
testY = to_categorical(testY)
trainNormX = trainX.astype('float32')
testNormX = testX.astype('float32')
trainNormX = trainNormX / 255.0
testNormX = testNormX / 255.0
print('Train: X=%s, y=%s' % (trainX.shape, trainY.shape))
print('Test: X=%s, y=%s' % (testX.shape, testY.shape))
for i in range(3):
pyplot.subplot(131 + i)
pyplot.imshow(trainX[i])
pyplot.show()
# + id="Q0utu1LnNsyc" colab={"base_uri": "https://localhost:8080/"} outputId="56f23c6a-d033-4cda-93c1-2ff8039f071d"
# define parameters
learningRates = [0.01]
activationMethods = ["tanh"]
lossFunctions = ["categorical_crossentropy"]
epochs = [40]
experiments = list(itertools.product(learningRates, activationMethods, lossFunctions, epochs))
experiments
# + id="J5Q68UD5utRF" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="955c0c75-9b94-4c17-9441-2af643183621"
# experiment models
selectedModel = None
selectedModelLoss = None
for experiment, parameters in enumerate(experiments):
learningRate, activation, loss, epochs = parameters
# display info
print(f"Experiment {experiment+1}")
print(f"Learning Rate: {learningRate}, Activation: {activation}, Loss: {loss}, Epochs: {epochs}")
print()
# build model
opt = SGD(learning_rate=learningRate, momentum=0)
model = keras.Sequential()
model.add(layers.Conv2D(filters=6, kernel_size=(3, 3), activation=activation, padding="same", input_shape=(32,32,3)))
model.add(layers.AveragePooling2D(pool_size=(2,2), strides=(1,1)))
model.add(layers.Conv2D(filters=12, kernel_size=(3, 3), activation=activation, padding="same", input_shape=(32,32,3)))
model.add(layers.AveragePooling2D(pool_size=(2,2), strides=(1,1)))
model.add(layers.Conv2D(filters=18, kernel_size=(3, 3), activation=activation, padding="same", input_shape=(32,32,3)))
model.add(layers.AveragePooling2D(pool_size=(2,2), strides=(1,1)))
model.add(layers.Conv2D(filters=24, kernel_size=(3, 3), activation=activation, padding="same", input_shape=(32,32,3)))
model.add(layers.AveragePooling2D(pool_size=(2,2), strides=(1,1)))
model.add(layers.Conv2D(filters=30, kernel_size=(3, 3), activation=activation, padding="same", input_shape=(32,32,3)))
model.add(layers.AveragePooling2D(pool_size=(2,2), strides=(1,1)))
model.add(layers.Flatten())
model.add(layers.Dense(units=120, activation=activation))
model.add(layers.Dense(units=84, activation=activation))
model.add(layers.Dense(units=10, activation = 'softmax'))
model.compile(optimizer=opt, loss=loss, metrics=['accuracy'])
history = model.fit(x=trainNormX, y=trainY, epochs=epochs, batch_size=200, validation_data=(testNormX, testY), verbose=1);
lossValue = history.history["loss"][-1]
if(selectedModel == None or selectedModelLoss > lossValue):
selectedModel = model
selectedModelLoss = lossValue
# plot accuracy
pd.DataFrame(history.history).plot(figsize=(8,5))
pyplot.legend(["Train Loss", "Train Accuracy", "Test Loss", "Test Accuracy"])
pyplot.show()
| CIFAR-10-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "slide"}
########################################################################
# File : CZI_ZARR.ipynb
# Version : 0.1
# Author : czsrh
# Date : 12.11.2019
# Insitution : Carl Zeiss Microscopy GmbH
#
# Disclaimer: Just for testing - Use at your own risk.
# Feedback or Improvements are welcome.
########################################################################
# + slideshow={"slide_type": "slide"}
# this can be used to switch on/off warnings
import warnings
warnings.filterwarnings('ignore')
warnings.simplefilter('ignore')
# import the libraries mentioned above
from apeer_ometiff_library import io, processing, omexmlClass
import czifile as zis
import xmltodict
import os
import numpy as np
import ipywidgets as widgets
import napari
import imgfileutils as imf
import xml.etree.ElementTree as ET
import zarr
# + slideshow={"slide_type": "slide"}
# define your testfiles here
#testfolder = r'C:\Users\m1srh\Documents\GitHub\ipy_notebooks\Read_OMETIFF_CZI\testdata'
testfolder = r'/datadisk1/tuxedo/Github/ipy_notebooks/Read_OMETIFF_CZI/testdata'
imgdict = {1: os.path.join(testfolder, r'CellDivision_T=10_Z=15_CH=2_DCV_small_green.ome.tiff'),
2: os.path.join(testfolder, r'CellDivision_T=10_Z=15_CH=2_DCV_small_red.ome.tiff'),
3: os.path.join(testfolder, r'CellDivision_T=10_Z=15_CH=2_DCV_small.ome.tiff'),
4: os.path.join(testfolder, r'CellDivision_T=10_Z=15_CH=2_DCV_small_green.czi'),
5: os.path.join(testfolder, r'CellDivision_T=10_Z=15_CH=2_DCV_small_red.czi'),
6: os.path.join(testfolder, r'CellDivision_T=10_Z=15_CH=2_DCV_small.czi')
}
#filename = imgdict[6]
#filename = r'/datadisk1/tuxedo/testpictures/Testdata_Zeiss/celldivison/CellDivision_T=15_Z=20_CH=2_DCV.czi'
#filename = r'L:\Data\Testdata_Zeiss\Castor\Castor_Beta\Castor_Beta2\20150610\Test4_4Ch_Hoechst_FITC_A568_MitoDR_20X0.95_1.0X_02_WellD3.czi'
filename = r"C:\Users\m1srh\OneDrive - Carl Zeiss AG\Testdata_Zeiss\LatticeLightSheet\LS_Mitosis_T=150-300.czi"
# + slideshow={"slide_type": "slide"}
# read metadata and array differently for OME-TIFF or CZI data
if filename.lower().endswith('.ome.tiff') or filename.lower().endswith('.ome.tif'):
# Return value is an array of order (T, Z, C, X, Y)
(array, omexml) = io.read_ometiff(filename)
metadata, add_metadata = imf.get_metadata(filename, series=0)
if filename.lower().endswith('.czi'):
# get the array and the metadata
array, metadata, add_metadata = imf.get_array_czi(filename, return_addmd=False)
# -
# show all the metadata
for key, value in metadata.items():
# print all key-value pairs for the dictionary
print(key, ' : ', value)
# + slideshow={"slide_type": "slide"}
# outout the shape of the returned numpy array
# shape of numpy array
print('Array Shape: ', array.shape)
# dimension order from metadata
print('Dimension Order (BioFormats) : ', metadata['DimOrder BF Array'])
# shape and dimension entry from CZI file as returned by czifile.py
print('CZI Array Shape : ', metadata['Shape'])
print('CZI Dimension Entry : ', metadata['Axes'])
# + slideshow={"slide_type": "slide"}
z = zarr.array(array, chunks=(1, 1, 5, 7964, 7164), dtype='uint16')
zarr_info = z.info
print(type(zarr_info))
if type(z) is np.ndarray:
print('NumPy Array')
elif type(z) is zarr.core.Array:
print('ZARR')
# + slideshow={"slide_type": "slide"}
# display data using ipy widgets
if metadata['Extension'] == 'ome.tiff':
ui, out = imf.create_ipyviewer_ome_tiff(array, metadata)
if metadata['Extension'] == 'czi':
#ui, out = imf.create_ipyviewer_czi(array, metadata)
ui, out = imf.create_ipyviewer_czi(z[:], metadata)
display(ui, out)
# + slideshow={"slide_type": "slide"}
# try to configure napari automatiaclly based on metadata
#imf.show_napari(array, metadata)
imf.show_napari(z[:], metadata)
# + slideshow={"slide_type": "slide"}
# configure napari viewer manually - check array shape and dimensions order carefully
# get the scalefactors
scalefactors = imf.get_scalefactor(metadata)
print(scalefactors)
array = np.squeeze(array, axis=(0, 1))
viewer = napari.Viewer()
# add every channel as a single layer
for ch in range(metadata['SizeC']):
chname = metadata['Channels'][ch]
viewer.add_image(array[ch, :, :, :], name=chname, scale=(1, scalefactors['zx'], 1, 1))
# -
# jupyter nbconvert CZI-ZARR.ipynb --to slides --post serve
| CZI-ZARR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cross-Resonance Gate
#
# *Copyright (c) 2021 Institute for Quantum Computing, Baidu Inc. All Rights Reserved.*
# ## Outline
#
# This tutorial introduces how to generate optimized pulses for Cross-Resonance (CR) gate using Quanlse. Unlike the iSWAP and CZ gate implementation in previous tutorials, CR gate is implemented using an all-microwave drive. The outline of this tutorial is as follows:
# - Introduction
# - Preparation
# - Construct Hamiltonian
# - Generate and optimize pulses via Quanlse Cloud Service
# - Summary
# ## Introduction
#
# **Fundamentals**
#
# Unlike some of the other gates we have seen before, the Cross-Resonance (CR) gate only uses microwaves to implement the two-qubit interaction such that we could avoid noise due to magnetic flux. The physical realization of the CR gate includes two coupled qubits with fixed frequencies. This can be done by driving the control qubit at the frequency of the target qubit. This is shown in the figure below:
#
#
# 
#
#
#
# We will first look at the effective Hamiltonian of the system (for details, please refer to Ref. \[1\] ). In the doubly rotating frame, the effective Hamiltonian for cross-resonance effect in terms of the drive strength $A$, detuning $\Delta$, drive phase $\phi_0$, and coupling strength $g_{01}$ is given \[1\] (for simplicity, we choose $\hbar = 1$) :
#
#
# $$
# \hat{H}_{\rm eff} = \frac{A}{4\Delta}g_{01}(\cos{\phi_0}\hat{\sigma}_0^z\hat{\sigma}_1^x+\sin{\phi_0}\hat{\sigma}_0^z\hat{\sigma}_1^y).
# $$
#
# When $\phi_0=0$, the cross-resonance effect allows for effective coupling of $\hat{\sigma}^z_0\otimes\hat{\sigma}_1^x$. We can thus derive the time evolution matrix from the effective Hamiltonian above:
#
# $$
# U_{\rm CR}(\theta)=e^{-i\frac{\theta}{2}\hat{\sigma}^z_0\otimes\hat{\sigma}^x_1},
# $$
#
# where $\theta = \Omega_0 gt/(2\Delta)$ ($t$ is the gate time). We can see that the cross-resonance effect enables a conditional rotation on qubit 1 (target qubit) depending on the state of qubit 0 (control qubit).
#
#
# Following the derivation above, the matrix form of the CR gate is (refer to \[2\] for more details):
# $$
# U_{\rm CR}(\theta) = \begin{bmatrix}
# \cos{\frac{\theta}{2}} & -i\sin{\frac{\theta}{2}} & 0 & 0 \\
# -i\sin{\frac{\theta}{2}} & \cos{\frac{\theta}{2}} & 0 & 0 \\
# 0 & 0 & \cos{\frac{\theta}{2}} & i\sin{\frac{\theta}{2}} \\
# 0 & 0 & i\sin{\frac{\theta}{2}} & \cos{\frac{\theta}{2}}
# \end{bmatrix}.
# $$
#
#
# In particular, the matrix representation of a CR gate with $\theta = -\frac{\pi}{2}$ is:
#
# $$
# U_{\rm CR}(-\pi/2) = \frac{\sqrt{2}}{2}
# \begin{bmatrix}
# 1 & i & 0 & 0 \\
# i & 1 & 0 & 0 \\
# 0 & 0 & 1 & -i \\
# 0 & 0 & -i & 1
# \end{bmatrix}.
# $$
#
# **Application**
#
# Having analyzed some of the fundamentals of the CR gate, we now switch our focus to the applications of the CR gate in quantum computing - one of which is the implementation of a CNOT gate with a CR gate and two additional single-qubit gates.
#
# 
#
# In this tutorial, we will model the system consisting of two three-level qubits and apply the drive pulse to the control qubit (qubit $q_0$) at the frequency of the target qubit (qubit $q_1$). By performing a rotating wave approximation (RWA), the Hamiltonian can be expressed as (refer to \[1\] for more details):
#
# $$
# \hat{H}_{\rm sys} = (\omega_{\rm q0}-\omega_{\rm d})\hat{a}_{0}^{\dagger }\hat{a}_0 + (\omega_{\rm q1}-\omega_{\rm d})\hat{a}_1^\dagger \hat{a}_1 + \frac{\alpha_0}{2} \hat{a}^{\dagger2}_0\hat{a}^2_0 + \frac{\alpha_1}{2} \hat{a}^{\dagger2}_1\hat{a}^2_1+\frac{g}{2}(\hat{a}_0\hat{a}_1^\dagger + \hat{a}_0^\dagger\hat{a}_1) + \Omega_0^x(t)\frac{\hat{a}^\dagger_0+\hat{a}_0}{2}.
# $$
#
# Please refer to the following chart for symbols' definitions:
#
# | Notation | Definition |
# |:--------:|:----------:|
# |$\omega_{\rm qi}$ | qubit $q_i$'s frequency |
# |$\omega_{\rm d}$|drive frequency|
# |$\hat{a}_i^{\dagger}$ | creation operator |
# |$\hat{a}_i$ | annihilation operator |
# |$\alpha_i$| qubit $q_i$'s anharmonicity |
# | $g$ | coupling strength |
# | $\Omega_0^x$(t) | pulse on the x channel |
# ## Preparation
#
# After you have successfully installed Quanlse, you could run the Quanlse program below following this tutorial. To run this particular tutorial, you would need to import the following packages from Quanlse and other commonly-used Python libraries:
# +
# Import Hamiltonian-related module
from Quanlse.Utils import Hamiltonian as qham
from Quanlse.Utils.Operator import driveX, number, duff
# Import optimizer for the cross-resonance gate
from Quanlse.remoteOptimizer import remoteOptimizeCr
# Import tools to analyze the result
from Quanlse.Utils.Tools import project, unitaryInfidelity
# Import numpy and math
from numpy import round
from math import pi
# -
# ## Construct Hamiltonian
#
# Now, we need to construct the above Hamiltonian using Quanlse. In Quanlse, all information regarding a Hamiltonian is stored in a dictionary. We start by defining some of the basic parameters needed for constructing a Hamiltonian dictionary: the sampling period, the number of qubits in the system, and the system's energy levels to consider. To initialize our Hamiltonian dictionary, we call the function `createHam()` from the module `Hamiltonian`.
# +
# Sampling period
dt = 2.0
# Number of qubits
qubits = 2
# System energy level
level = 3
# Initialize the Hamiltonian
ham = qham.createHam(title='cr-gate', dt=dt, qubitNum=qubits, sysLevel=level)
# -
# Now we can start constructing our Hamiltonian. Before we start, we would need to define a few constants to pass in as the function's arguments:
# Parameters setting
g = 0.0038 * (2 * pi) # Coupling strength 1
wq0 = 4.914 * (2 * pi) # Transition frequency for qubit 0, GHz
wq1 = 4.714 * (2 * pi) # Transition frequency for qubit 1, GHz
wd = wq1 # Drive frequency is the frequency for qubit 1, GHz
anharm0 = -0.33 * (2 * pi) # Anharmonicity for qubit 0, GHz
anharm1 = -0.33 * (2 * pi) # Anharmonicity for qubit 1, GHz
detuning0 = wq0 - wd
detuning1 = wq1 - wd
# Now we need to add the following terms to the Hamiltonian dictionary we initilized earlier:
# $$
# \hat{H}_{\rm drift} = (\omega_{\rm q0}-\omega_{\rm d}) \hat{a}_0^\dagger \hat{a}_0 + (\omega_{\rm q1}-\omega_{\rm d}) \hat{a}_1^\dagger \hat{a}_1 + \frac{\alpha_0}{2} \hat{a}_0^{\dagger}\hat{a}_0^{\dagger}\hat{a}_0 \hat{a}_0 + \frac{\alpha_1}{2} \hat{a}_1^{\dagger}\hat{a}_1^{\dagger}\hat{a}_1 \hat{a}_1,
# $$
# $$
# \hat{H}_{\rm coup} = \frac{g_{01}}{2}(\hat{a}_0 \hat{a}_1^\dagger+\hat{a}^\dagger_0 \hat{a}_1).
# $$
#
# In Quanlse's `Operator` module, we have provided tools that would allow the users to construct the commonly used operators quickly. The detuning term $(\omega_{\rm q}-\omega_{\rm d})\hat{a}^\dagger \hat{a}$ and the anharmonicity term $\frac{\alpha}{2}\hat{a}^\dagger\hat{a}^\dagger \hat{a} \hat{a} $ can be respectively generated using `number(n)` and `duff(n)` from the `Operator` module: the two functions `number(n)` and `duff(n)` return the $n \times n$ matrices for number operators and duffing operators. The coupling term, which takes the form, $\frac{g}{2}(\hat{a}_i^\dagger\hat{a}_j+\hat{a}_i\hat{a}_j^\dagger$), can be directly added to the Hamiltonian using function `addCoupling()`.
# +
# Add the detuning terms
qham.addDrift(ham, name='detuning0', onQubits=0, amp=detuning0, matrices=number(level))
qham.addDrift(ham, name='detuning1', onQubits=1, amp=detuning1, matrices=number(level))
# Add the anharmonicity terms
qham.addDrift(ham, name='anharm0', onQubits=0, amp=0.5 * anharm0, matrices=duff(level))
qham.addDrift(ham, name='anharm1', onQubits=1, amp=0.5 * anharm1, matrices=duff(level))
# Add the coupling term
qham.addCoupling(ham, 'coupling', [0, 1], g=0.5 * g)
# -
# Finally, we add the control term to the system Hamiltonian. The matrix for the control term can be defined by `driveX(level)` in the `Operator` module.
# $$
# \hat{H}_{\rm ctrl} = \Omega_0^x(t)\frac{\hat{a}^\dagger_0+\hat{a}_0}{2}.
# $$
# Add the control term
qham.addControl(ham, name='q0-ctrlx', onQubits=0, matrices=driveX(level))
# With the system Hamiltonian built, we can now move on to the optimization.
# ## Generate and optimize pulse via Quanlse Cloud Service
# The optimization process usually takes a long time to process on local devices; however, we provide a cloud service that could speed up this process significantly. To use the Quanlse Cloud Service, the users need to acquire a token from http://quantum-hub.baidu.com.
# +
# Import tools to get access to cloud service
from Quanlse import Define
# To use remoteOptimizerCr on cloud, paste your token (a string) here
Define.hubToken = ''
# -
# To find the optimized pulse for CR gate, we use the function `remoteOptimizeCr()`, which takes the Hamiltonian we had previously defined, amplitude's bound, gate time, maximum iterations, and target infidelity. By calling `remoteOptimizeCr()`, the user can submit the optimization task to the Quanlse's server. If the user wants to further mitigate the infidelity, we encourage trying an increased gate time `tg` (the duration of a CR gate is around 200 to 400 nanoseconds). Users can also try increasing the search space by setting larger `aBound` and `maxIter`.
#
# The gate infidelity for performance assessment throughout this tutorial is defined as ${\rm infid} = 1 - \frac{1}{d}\left|{\rm Tr}[U^\dagger_{\rm goal}P(U)]\right|$, where $U_{\rm goal}$ is exactly the target unitary transformation $U_{\rm CR}(-\pi/2)$; $d$ is the dimension of $U_{\rm goal}$; and $U$ is the unitary evolution of the three-level system defined previously. Note that $P(U)$ in particular describes the evolution projected to the computational subspace.
# +
# Set amplitude bound
aBound = (1.0, 3.0)
# Run the optimization
ham, infidelity = remoteOptimizeCr(ham, aBound=aBound, tg=200, maxIter=5, targetInfidelity=0.005)
# -
# We can visualize the generated pulse using `plotWaves()`. (details regarding `plotWaves()` are covered in [single-qubit-gate.ipynb](https://quanlse.baidu.com/#/doc/tutorial-single-qubit))
# Print waves and the infidelity
qham.plotWaves(ham, ['q0-ctrlx'])
print(f'infidelity: {infidelity}')
# The users can also print the the projected evolution $P(U)$ using the following lines:
# Print the projected evolution
result = qham.simulate(ham)
process2d = project(result["unitary"], qubits, level, 2)
print("The projected evolution P(U):\n", round(process2d, 2))
# Moreover, for those interested in acquiring the numerical data of the generated pulse for each `dt`, use function `getPulseSequences()`, which takes a Hamiltonian dictionary and channels' names as parameters.
qham.getPulseSequences(ham, 'q0-ctrlx')
# ## Summary
#
# From constructing the system Hamiltonian to generating an optimized pulse on Quanlse Cloud Service, we have successfully devised a pulse to implement a cross-resonace gate with high fidelity. The users are encouraged to try parameter values different from this tutorial to obtain the optimal result.
# + [markdown] pycharm={"name": "#%%\n"}
# ## References
#
# \[1\] [<NAME>, and <NAME>. "Fully microwave-tunable universal gates in superconducting qubits with linear couplings and fixed transition frequencies." *Physical Review B* 81.13 (2010): 134507.](https://qulab.eng.yale.edu/documents/papers/Rigetti,%20Devoret%20-%20Fully%20Microwave-Tunable%20Universal%20Gates%20in%20Superconducting%20Qubits%20with%20Linear%20Couplings%20and%20Fixed%20Transition%20Frequencies.pdf)
#
# \[2\] Nielsen, <NAME>., and <NAME>. Quantum Computation and Quantum Information: 10th Anniversary Edition. Cambridge University Press, 2010.
| Tutorial/EN/tutorial-cr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import syft as sy
# # Part 1: Launch a Duet Server and upload data
duet = sy.launch_duet(loopback=True)
import torch as th
import numpy as np
# +
data = th.FloatTensor(np.array([5, 15, 25, 35, 45, 55]).reshape(-1, 1))
data = data.tag("DO1 data")
data = data.describe("Dataset of 6 samples, 1 feature")
data_ptr = data.send(duet, pointable=True)
# -
duet.store.pandas
data
duet.requests.add_handler(
action="accept",
print_local=True, # print the result in your notebook
)
| examples/private-ai-series/duet_fl/Duet_FL_1_Data_Owner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
from torchvision import models
from torchvision.transforms import transforms
from torchvision.transforms.functional import crop
import cv2
import albumentations as A
from albumentations.pytorch import ToTensorV2
import numpy as np
import pandas as pd
from tqdm import tqdm
from dataclasses import dataclass
import time
import matplotlib.pyplot as plt
from IPython import display
# %matplotlib inline
# +
@dataclass
class State:
frame: torch.Tensor
sp: float # stamina points
zoom: int
@dataclass
class Action:
direction: int
time_steps: int
def find_state_with_stone(df: pd.DataFrame, max_attempt: int = 10) -> State:
while True:
index = np.random.randint(0, len(df) - 1)
frame = load_image(df["video"][index], df["frame"][index])
train_aug = A.Compose([A.Normalize(mean=(0.5,), std=(0.5,)),
ToTensorV2(transpose_mask=False),
])
frame = train_aug(image=frame)['image']
if env.check_frame(frame):
break
sp = df["sp"][index]
zoom = df["zoom"][index]
state = State(frame, sp, zoom)
return state
def see_plot(pict, size=(6, 6), title: str = None):
plt.figure(figsize=size)
plt.imshow(pict, cmap='gray')
if title is not None:
plt.title(title)
plt.show()
def load_image(video, frame):
path = '../surviv_rl_data/all_videoframes_rgb_96/{}/'.format(video)
p = cv2.imread(path + 'f_{}.jpg'.format(frame))
return p[:,:,::-1]
def convrelu(in_channels, out_channels, kernel, padding):
return nn.Sequential(
nn.Conv2d(in_channels, out_channels, kernel, padding=padding),
nn.BatchNorm2d(out_channels),
nn.ReLU(inplace=True),
)
class ResNetUNet_v2(nn.Module):
def __init__(self, n_class):
super().__init__()
self.base_model = models.resnet18(pretrained=True)
self.base_layers = list(self.base_model.children())
self.layer0 = nn.Sequential(*self.base_layers[:3]) # size=(N, 64, x.H/2, x.W/2)
self.layer0_1x1 = convrelu(64, 64, 1, 0)
self.layer1 = nn.Sequential(*self.base_layers[3:5]) # size=(N, 64, x.H/4, x.W/4)
self.layer1_1x1 = convrelu(64, 64, 1, 0)
self.layer2 = self.base_layers[5] # size=(N, 128, x.H/8, x.W/8)
self.layer2_1x1 = convrelu(128, 128, 1, 0)
self.layer3 = self.base_layers[6] # size=(N, 256, x.H/16, x.W/16)
self.layer3_1x1 = convrelu(256, 256, 1, 0)
self.layer4 = self.base_layers[7] # size=(N, 512, x.H/32, x.W/32)
self.layer4_1x1 = convrelu(512, 512, 1, 0)
self.upsample = nn.Upsample(scale_factor=2, mode='bilinear', align_corners=True)
self.conv_up3 = convrelu(256 + 512, 512, 3, 1)
self.conv_up2 = convrelu(128 + 512, 256, 3, 1)
self.conv_up1 = convrelu(64 + 256, 256, 3, 1)
self.conv_up0 = convrelu(64 + 256, 128, 3, 1)
self.conv_original_size0 = convrelu(3, 64, 3, 1)
self.conv_original_size1 = convrelu(64, 64, 3, 1)
self.conv_original_size2 = convrelu(64 + 128, 64, 3, 1)
self.dropout = nn.Dropout(0.5)
self.conv_last = nn.Conv2d(64, n_class, 1)
self.act_last = nn.Tanh()
self.support_conv1 = nn.Conv2d(11, 512, 1) # (bath,10+1) --> (batch,512)
def forward(self, inp):
x_original = self.conv_original_size0(inp[0])
x_original = self.conv_original_size1(x_original)
layer0 = self.layer0(inp[0])
layer1 = self.layer1(layer0)
layer2 = self.layer2(layer1)
layer3 = self.layer3(layer2)
layer4 = self.layer4(layer3)
cond = self.support_conv1(torch.unsqueeze(torch.unsqueeze(inp[1], 2), 2)) # ([8, 8]) --> Size([8, 512, 1, 1])
layer4 = self.layer4_1x1(layer4 + cond)
x = self.upsample(layer4)
layer3 = self.layer3_1x1(layer3)
x = torch.cat([x, layer3], dim=1)
x = self.conv_up3(x)
x = self.upsample(x)
layer2 = self.layer2_1x1(layer2)
x = torch.cat([x, layer2], dim=1)
x = self.conv_up2(x)
x = self.upsample(x)
layer1 = self.layer1_1x1(layer1)
x = torch.cat([x, layer1], dim=1)
x = self.conv_up1(x)
x = self.upsample(x)
layer0 = self.layer0_1x1(layer0)
x = torch.cat([x, layer0], dim=1)
x = self.conv_up0(x)
x = self.upsample(x)
x = torch.cat([x, x_original], dim=1)
x = self.conv_original_size2(x)
x = self.dropout(x)
out = self.conv_last(x)
out = self.act_last(out)
return out
#====================================================================
class StoneClassifier(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(3, 8, 3, 2, 1)
self.conv2 = nn.Conv2d(8, 16, 3, 2, 1)
self.conv3 = nn.Conv2d(16, 32, 3, 2, 1)
self.fc1 = nn.Linear(32 * 3 * 3, 128)
self.fc3 = nn.Linear(128, 2)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.relu(self.conv2(x))
x = F.relu(self.conv3(x))
x = torch.flatten(x, 1)
x = F.relu(self.fc1(x))
x = F.softmax(self.fc3(x), dim=1)
return x
class NeuralEnv:
def __init__(self,
env_model_path: str,
reward_model_path: str,
device: str,
batch_size = 16,
reward_confidence=0.5,
stone_frac=0.0,
step_size=4,
max_step=14):
'''
input params:
env_model_path [str] : path to model s_next=model(s_curr,action)
reward_model_path [str] : path to model reward=model(s_curr)
device [str] : one of {'cpu', 'cuda:0', 'cuda:1'}
batch_size [int] : len of batch
reward_confidence [flt] : classificator's confidence
stone_frac [flt] : part of the initial states with guaranteed stones
step_size [int] :
max_step [int] :
output params:
all output-variables will be torch.tensors in the selected DEVICE
all input-variables have to be torch.tensors in the selected DEVICE
'''
self.device = device
self.batch_size = batch_size
self.reward_confidence = reward_confidence
self.stone_frac = stone_frac
self.step_size = step_size
self.max_step = max_step
self.reward_frame_transform = transforms.Compose([transforms.CenterCrop(24)])
self.frame_transform = A.Compose([A.Normalize(mean=(0.5,), std=(0.5,)),
ToTensorV2(transpose_mask=False)])
self.model = ResNetUNet_v2(3)
self.model.load_state_dict(torch.load(env_model_path, map_location=self.device))
self.model = self.model.to(self.device)
self.model.eval()
self.stone_classifier = StoneClassifier()
self.stone_classifier.load_state_dict(torch.load(reward_model_path, map_location=self.device))
self.stone_classifier = self.stone_classifier.to(self.device)
self.stone_classifier.eval()
self.df = pd.read_csv('../surviv_rl_data/dataset_inventory_v2.csv')
self.df = self.df[self.df.zoom == 1].reset_index()
#----------------------------------------------------------------------------------------------------
def reset(self, specific_state = None):
'''
output params:
init_s [float torch tensor [-1...1]] : batch of initial states (batch,3,96,96)
init_supp [float torch tensor] : batch of initial support vector (batch,2)
'''
init_s = torch.zeros(self.batch_size,3,96,96).float()
init_supp = torch.zeros(self.batch_size,2).float()
for i in range(self.batch_size):
while True:
j = np.random.randint(len(self.df))
if specific_state is not None:
j = specific_state
frame = load_image(self.df["video"][j], self.df["frame"][j])
frame = self.frame_transform(image=frame)['image']
if self.check_frame(frame):
break
supp = torch.tensor([self.df["sp"][j]/100,self.df["zoom"][j]/15]).float()
#if check_frame(frame)==True:
# init_s[i] = frame
# init_supp[i] = supp
init_s[i] = frame
init_supp[i] = supp
return init_s.to(self.device),init_supp.to(self.device)
#----------------------------------------------------------------------------------------------------
def get_reward(self, state):
'''
input params:
state [float torch.tensor [-1...1]] : batch of states (batch,3,96,96)
output params:
r [float torch.tensor [0...1]] : batch of rewards (batch,1)
'''
state = self.reward_frame_transform(state)
with torch.no_grad():
r = self.stone_classifier(state)[:,1].unsqueeze(1)
r = (r>self.reward_confidence).float().detach()
return r
#----------------------------------------------------------------------------------------------------
def step(self, s_curr, supp_curr, action):
'''
input params:
s_curr [float torch.tensor [-1...1]] : batch of current states (batch,3,96,96)
supp_curr [float torch tensor] : batch of current support vector (batch,2)
action [int torch tensor {1,...,8}] : batch of chosen direction (batch,1)
output params:
s_next [float torch.tensor [-1...1]] : batch of next states (batch,3,96,96)
supp_next [float torch tensor] : batch of next support vector =supp_curr (batch,2)
reward [float torch.tensor [0...1]] : batch of rewards (batch,1)
'''
action_ohe = F.one_hot(action.squeeze()-1, num_classes=8).float() # (batch,8)
if len(action_ohe.shape) == 1:
action_ohe = action_ohe[None]
n = torch.tensor([self.step_size/self.max_step]*self.batch_size)
n = n.unsqueeze(1).float().to(self.device) # (batch,1)
v = torch.cat([action_ohe,supp_curr,n], dim=1) # (batch,8+2+1)
with torch.no_grad():
s_next = self.model((s_curr,v)).detach()
reward = self.get_reward(s_next)
return s_next, supp_curr, reward
# Check if frame has stone
def check_frame(self, frame) -> bool:
crops = []
for i in range(4):
for j in range(4):
cropped = crop(frame, i * 24, j * 24, 24, 24)[None]
crops.append(cropped)
expected_stone = self.stone_classifier(torch.cat(crops)).max(dim=0)[0][1].item()
return expected_stone > 0.5
# -
class DQN(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(3, 8, 3, 2, 1)
self.conv2 = nn.Conv2d(8, 16, 3, 2, 1)
self.conv3 = nn.Conv2d(16, 32, 3, 2, 1)
self.linear1 = nn.Linear(32 * 12 * 12, 256)
self.linear2 = nn.Linear(256, 8)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.relu(self.conv2(x))
x = F.relu(self.conv3(x))
x = F.relu(self.linear1(torch.flatten(x, 1)))
x = self.linear2(x)
x = F.softmax(x, dim=1)
return x
dqn = DQN()
dqn.load_state_dict(torch.load("../best_models/dqn_v1.pth"))
target_network = DQN()
target_network.load_state_dict(dqn.state_dict())
env = NeuralEnv("../best_models/resunet_v5.pth",
"../best_models/laggg_stone_classifier_v2.pth",
"cpu",
1,
reward_confidence=0.95)
device = torch.device("cpu")
# +
from replay_buffer import ReplayBuffer
EXP_REPLAY_SIZE = 1000
exp_replay = ReplayBuffer(EXP_REPLAY_SIZE)
for i in range(EXP_REPLAY_SIZE):
s, supp = env.reset()
a = np.random.randint(8)
next_s, _, r = env.step(s, supp, torch.LongTensor([a + 1]))
r = r[0][0].detach().cpu()
exp_replay.add(s[0].detach().cpu(), a, r, next_s[0].detach().cpu(), False)
# -
def get_action(state, epsilon=0):
state = torch.tensor(state)
q_values = dqn(state).detach().numpy()
if np.random.rand() < epsilon:
return int(np.random.randint(len(q_values[0])))
else:
return int(np.argmax(q_values, axis=1))
def compute_td_loss(states, actions, rewards, next_states, target_network, gamma=0.99, check_shapes=False):
states = torch.tensor(states, dtype=torch.float32)
actions = torch.tensor(actions, dtype=torch.long)
rewards = torch.tensor(rewards, dtype=torch.float32)
next_states = torch.tensor(next_states, dtype=torch.float32)
predicted_qvalues = dqn(states)
predicted_qvalues_for_actions = predicted_qvalues[
range(states.shape[0]), actions
]
predicted_next_qvalues = target_network(next_states)
next_state_values = predicted_next_qvalues.max(dim=1)[0]
# compute "target q-values" for loss - it's what's inside square parentheses in the above formula.
target_qvalues_for_actions = rewards + gamma * next_state_values
loss = torch.mean((predicted_qvalues_for_actions -
target_qvalues_for_actions.detach()) ** 2)
return loss
opt = torch.optim.Adam(dqn.parameters(), lr=1e-5)
def generate_session(env, t_max=3, epsilon=0, train=False):
total_reward = 0
s, supp = env.reset()
for t in range(t_max):
a = get_action(s, epsilon=epsilon)
next_s, next_supp, r = env.step(s, supp, torch.LongTensor([a + 1]))
r = r[0][0].detach().cpu()
exp_replay.add(s[0], a, r, next_s[0], False)
if train:
opt.zero_grad()
s_batch, a_batch, r_batch, next_s_batch, is_done_batch = exp_replay.sample(32)
loss = compute_td_loss(s_batch, a_batch, r_batch, next_s_batch, target_network, gamma=0.5)
loss.backward()
opt.step()
total_reward += r
s = next_s
return total_reward
epsilon = 0.5
for i in range(100):
session_rewards = [generate_session(env, epsilon=epsilon, train=True) for _ in range(100)]
print("epoch #{}\tmean reward = {:.3f}\tepsilon = {:.3f}".format(i, np.mean(session_rewards), epsilon))
if i % 10 == 0:
target_network.load_state_dict(dqn.state_dict())
epsilon *= 0.99
s_curr, supp_curr = env.reset()
cum_reward = 0
picts = []
for i in range(20):
chosen_action = get_action(s_curr, epsilon=0)
s_next, supp_next, reward = env.step(s_curr, supp_curr, torch.LongTensor([[chosen_action + 1]]).to(device))
cum_reward += reward[0][0].detach().cpu()
display.clear_output(wait=True)
image = s_next[0].permute(1, 2, 0).cpu().detach().numpy() / 2 + 0.5
see_plot(image, (5, 5), f"{cum_reward}")
picts.append(image)
time.sleep(0.1)
s_curr, supp_curr = s_next, supp_next
torch.save(dqn.state_dict(), "../best_models/dqn_v7.pth")
import imageio
with imageio.get_writer('test3.gif') as writer:
for pict in picts:
writer.append_data(pict)
| jupyter/rl/surviv_dqn_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cheminformatics using Python: Working with Chemical Compounds
# ## What is Cheminformatics?
# - **Cheminformatics** is the use of computer and informational techniques, applied to a range of problems in the field of chemistry.
# - It is also known as **chemoinformatics** and **chemical informatics**.
# - These in *silico* techniques are used in pharmaceutical companies in the process of drug discovery.
# ## Application of Cheminformatics
# - The primary application of **cheminformatics** is in the storage of information relating to compounds.
# - **Quantitative structure–activity relationship (QSAR)** analysis also forms a part of cheminformatics.
# ## Why Cheminformatics?
# - Chemoinformatics helps to reduce the time taken for identifying potential drug targets as well as to understand **physical**, **chemical** and **biological** properties of several chemical compounds.
# - Outputs of chemoinformatics may also direct the course of wet laboratory experiments.
# ## Cheminformatics Tasks
# - Similarity search of chemicals.
# - Prediction of affinities, physicochemical properties and pharmacokinetic properties
# - Prediction of molecular solubility.
# ## References
# - https://onlinelibrary.wiley.com/doi/abs/10.1002/9780470744307
# ## Important Terminology
# - **SMILES Notation:** SMILES (Simplified Molecular Input Line Entry System) is a chemical notation that allows a user to represent a chemical structure in a way that can be used by the computer. SMILES is an easily learned and flexible notation.
# ## Data Collection
import pandas as pd
from chembl_webresource_client.new_client import new_client
# ## Introduction to RDKit
# RDKit is a collection of cheminformatics and machine-learning software written in C++ and Python.
from rdkit import Chem
from rdkit.Chem import Draw
from rdkit.Chem.Draw import IPythonConsole
from rdkit.Chem import Descriptors
import numpy as np
# molecules from smiles
Chem.MolFromSmiles('CCC')
# molecules from smiles
mol = Chem.MolFromSmiles('CCC')
mol
asp = Chem.MolFromSmiles('CC(=O)OC1=CC=CC=C1C(=O)O')
asp
asp.GetNumAtoms()
asp.GetNumBonds()
asp.GetNumHeavyAtoms()
# molecule to smiles
Chem.MolToSmiles(asp)
asp
Descriptors.MolWt(asp)
Descriptors.NumRotatableBonds(asp)
Descriptors.MolLogP(asp)
smiles_list = [
'CC(=O)OC1=CC=CC=C1C(=O)O',
'C(C(=O)O)N',
'C1=C(NC=N1)CC(C(=O)O)N'
]
mols_list = []
for smiles in smiles_list:
mol = Chem.MolFromSmiles(smiles)
mols_list.append(mol)
mols_list
Draw.MolsToGridImage(mols_list)
| notebooks/Cheminformatics using Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## LIBRARY IMPORT
import pandas as pd
import numpy as np
import os
import re
# ## READING FILES
train_reviews = []
for line in open('../movie-sentiment-analysis/aclImdb/movie_data/full_train.txt','r',encoding='utf-8'):
train_reviews.append(line.strip())
test_reviews = []
for line in open('../movie-sentiment-analysis/aclImdb/movie_data/full_test.txt','r',encoding='utf-8'):
test_reviews.append(line.strip())
# ## CLEANING AND PROCESSING
no_space = re.compile("(\.)|(\;)|(\:)|(\!)|(\')|(\?)|(\,)|(\")|(\()|(\))|(\[)|(\])|(\d+)")
with_space = re.compile("(<br\s*/><br\s*/>)|(\-)|(\/)")
def cleaningandprocessing(reviews):
reviews = [no_space.sub("",line.lower()) for line in reviews]
reviews = [with_space.sub(" ",line) for line in reviews]
return reviews
clean_train_reviews = cleaningandprocessing(train_reviews)
clean_test_reviews = cleaningandprocessing(test_reviews)
# ## BASELINE OF LOGISTIC REGRESSION PERFORMED
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
from sklearn.linear_model import LogisticRegression
baseline_vectorizer = CountVectorizer(binary=True)
baseline_vectorizer.fit(clean_train_reviews)
X_train = baseline_vectorizer.transform(clean_train_reviews)
X_test = baseline_vectorizer.transform(clean_test_reviews)
target = [1 if i <12500 else 0 for i in range(25000)]
x_train, x_test, y_train, y_test = train_test_split(X_train, target, test_size=0.8, random_state=42)
for c in [0.01,0.05,0.1,0.5,1]:
lr = LogisticRegression(C=c)
lr.fit(x_train, y_train)
print("For: ", c)
print("\n")
print(accuracy_score(y_test, lr.predict(x_test)))
print("\n")
print(confusion_matrix(y_test, lr.predict(x_test)))
print("\n")
print(classification_report(y_test, lr.predict(x_test)))
print("\n\n")
final_model = LogisticRegression(C=0.1)
final_model.fit(X_train, target)
print(accuracy_score(target, final_model.predict(X_test)))
print("\n")
print(confusion_matrix(target, final_model.predict(X_test)))
print("\n")
print(classification_report(target,final_model.predict(X_test)))
# ### TILL NOW MAX ACCURACY ACHIEVED ON FINAL MODEL IS 87.93
def get_stemmed_text(corpus):
from nltk.stem.porter import PorterStemmer
stemmer = PorterStemmer()
return [' '.join([stemmer.stem(word) for word in review.split()]) for review in corpus]
stemmed_reviews_train = get_stemmed_text(clean_train_reviews)
stemmed_reviews_test = get_stemmed_text(clean_test_reviews)
| BaselineLogisticRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Design of Reinforced Concrete Sections as per IS 456:2000
# While reinforced concrete sections can have any shape, the most commonly used sections are the rectangular, flanged (T and L) and circular sections. The simplest of these is the rectangular section. IS 456 prescribes both the limit state method and working stress method. Of these, limit state method is more commonly used, however, certain types of structures are required to be design using working stress method. Some examples of such structures are liquid retaining structures and bridges. We will begin with the limit state method of design and first take up rectangular sections.
#
# The basic parameters required in the design of reinforced concrete sections are characteristic strengths of materials, partial safety factors for materials and the stress strain relations for the materials. All further derivations will be derived from these fundamental parameters. We will use the notations from IS 456:2000
#
# 1. Characteristic strength of concrete $f_{ck}$: Depends on the grade of concrete, which can be from M20 to M80. The corresponding characteristic strengths, in N/mm$^2$ are 20 and 80, respectively.
# 2. Characteristic strength of strength of steel $f_y$: Depends on the grade of steel, which can be one of Mild steel, Fe 415 and Fe 500. The corresponding characteristic strengths, in N/mm$^2$ are 250, 415 and 500 respectively.
# 3. The partial safety factor for material is 1.5 for concrete and 1.15 for steel.
# 4. For the limit state of collapse by flexure is defined in terms of the following parameters:
# 1. For design purposes, the compressive strength of concrete is obtained as $0.67 \frac{f_{ck}}{\gamma_m} = 0.67 \frac{f_{ck}}{1.5} = 0.446 f_{ck}$
# 2. The tensile strength of concrete is ignored
# 3. The stresses in the reinforcement are derived from representative stress-strain for the type of steel used. For design purposes, design strength of steel in tension and compression is assumed to be identical and is taken as $\frac{f_y}{\gamma_m} = \frac{f_y}{1.15} = 0.87 f_y$
# 4. The strain in tension reinforcement in the section at failure shall not be less than $\frac{f_y}{1.15 E_s} + 0.002 = 0.87 f_y/E_s + 0.002$
#
# ## Stress Strain Curve of Concrete
# The stress strain curve for concrete for the limit state of collapse in flexure is specified with the help of following parameters:
#
# 1. The yield strain of concrete is $\varepsilon_{cy} = 0.002$
# 2. The maximum strain in concrete is $\varepsilon_{cu} = 0.0035$
# 3. The stress strain relation is parabolic up to yield and subsequently it is constant up to the maximum strain of 0.0035
#
# ## Stress Strain Curve for Mild Steel
# The stress strain curve for mild steel is bilinear and is specified as follows:
#
# 1. From zero strain to yield strain $0.87 f_y / E_s$, stress varies linearly and the modulus of elasticity is $E_s = 2 \times 10^5$ N/mm$^2$
# 2. After yield, stress remains constant at $0.87 f_y$
# 3. Steel is assumed to be infinitely ductile and hence is assumed to be capable of undergoing infinite strains withour breaking
#
# ## Stress Strain Curve for Cold Worked Steels
# The stress strain curve for cold worked steels is defined in terms of a series of straight lines, with the modulus of elasticity at start same as that for mild steel, namely, $E_s = 2 \times 10^5$ N/mm$^2$. The code specifies stress for different levels of inelastic strain.
# +
# %matplotlib inline
import math
import numpy as np
import matplotlib.pyplot as plt
ecy = 0.002
ecu = 0.0035
Es = 2.0e5
gamma_c = 1.5
gamma_s = 1.15
def fc(e):
if e < 0:
return 0.0
elif e < ecy:
return 0.67 / 1.5 * (e / ecy) * (2.0 - (e / ecy))
elif e <= ecu:
return 0.67 / 1.5
else:
return 0.0
def interpolate(x, y, xx):
if xx < x[0]:
return None
elif xx > x[-1]:
return None
for i in range(len(x)-1):
x1 = x[i]
if xx == x1:
return (y[i])
elif xx == x[i+1]:
return y[i+1]
elif xx < x[i+1]:
y1 = y[i]
x2 = x[i+1]
y2 = y[i+1]
yy = y1 + (y2 - y1) / (x2 - x1) * (xx - x1)
return yy
def fs_cw_stress_strain(fy):
inelstrain = np.array([0.0, 0.0, 0.0001, 0.0003, 0.0007, 0.001, 0.002], dtype=float)
sigma = np.array([0.0, 0.8, 0.85, 0.90, 0.95, 0.975, 1.0, 1.0], dtype=float)
eps = np.zeros(sigma.shape, dtype=float)
for i in range(1,len(inelstrain)):
sigma[i] *= (fy / 1.15)
eps[i] = (sigma[i] / Es) + inelstrain[i]
sigma[-1] = sigma[-2]
eps[-1] = np.inf
return eps, sigma
def fs(e, fy, steel='mild steel'):
e = abs(e)
esy = fy / gamma_s / Es
if steel == 'mild steel':
if e < esy:
return e * Es
else:
return fy / gamma_s
elif steel == 'cold worked steel':
pass
return
if __name__ == '__main__':
print -0.01, fc(-0.01)
print 0.0, fc(0.0)
print 0.001, fc(0.001)
print 0.002, fc(0.002)
print 0.003, fc(0.003)
print 0.0035, fc(0.0035)
print 0.004, fc(0.004)
x = np.linspace(0, 0.0035, 201)
y = np.zeros(x.shape, dtype=float)
for i in range(len(x)):
y[i] = fc(x[i])
plt.plot(x, y)
plt.ylim(0.0, 0.5)
plt.grid()
plt.show()
x = np.linspace(0, 0.002, 1001)
y = np.zeros(x.shape, dtype=float)
for i in range(len(x)):
y[i] = fs(x[i], 250)
plt.plot(x, y)
plt.grid()
plt.show()
# +
eps, str = fs_cw_stress_strain(415)
print interpolate(eps, str, -0.01)
print interpolate(eps, str, 0)
print interpolate(eps, str, 0.001)
print interpolate(eps, str, 0.002)
print interpolate(eps, str, 0.005)
# -
# ## Rectangular Sections
#
# A rectangular section with reinforcement only on the tension side is the simplest possible section. Depending on the dimensions of the cross section, grades of steel and concrete used and the amount of reinforcement provided, the section can be classified as an under-reinforced, balanced or an over-reinforced section. If the area of reinforcement provided is such that steel reaches its specified minimum strain at the same time when strain in the extreme concrete fibre reaches its breaking strain, the section is said to be **balanced**.
#
# On the other hand, if when concrete reaches its breaking strain steel has not yet reached its minimum specified strain, section is said to be **over-reinforced**. This occurs when the area of tension reinforcement provided is more than what is required for a balanced section. Concrete fails before steel, and being a brittle material, section fails without giving sufficient advance warning.
#
# If the strain in steel is more than the specified minimum starin when concrete reaches its breaking strain, section is said to be **under-reinforced**. This occurs when the area of reinforcement provided is less than that required by a balanced section. In this case, steel fails first, but being a ductile material, it does not break immediately. It is capable of undergoing large deformations beyond yield without breaking, thereby giving concrete sufficient opportunity to reach its breaking strain. Failure of such a section is gradual and there is sufficient advance warning before the section collapses.
#
# It is a preferred method of design to provde a depth greater than the depth of a balanced section. Thus the capacity of the section is greater than the required design moment if the section is designed as a balanced section. It is thus possible to generate the required resistance equal to the design moment by providing area of reinforcement less than that for a balanced section, thereby making the section under-reinforced.
#
# ### Balanced Section
# A balanced section must have a specified depth if it is to generate a resisting moment exactly equal to the design moment and have the required strains in tension steel and concrete at the same instant of time. This can be obtained by equating the ratio of the sides of the similar triangles.
#
# $$\frac{x_{u, max}}{0.0035} = \frac{d - x_{u, max}}{0.002 + \frac{0.87 f_y}{E_s}}$$
#
# Solving for $x_{u, max}$ we get $\frac{x_{u, max}}{d} = \frac{0.0035}{0.0055 + \frac{0.87 f_y}{E_s}}$
#
# A rectangular section having this exact depth would be a balanced section when it is provided with the exact area of steel required by a balanced section, obtained by equating the magnitudes of the compressive force in concrete with the tensile force in steel.
#
# $$A_{st, lim} (0.87 f_y) = 0.36 f_{ck} b x_{u, max}$$
#
# Solving for $A_{st, lim}$, we get $A_{st, lim} = \frac{0.36 f_{ck} b x_{u, max}}{0.87 f_y}$. This can be written as
#
# $$p_{t, lim} = \frac{A_{st, lim}}{b d} \cdot 100 = \frac{0.36 f_{ck}}{0.87 f_y} \cdot \frac{x_{u, max}}{d} \cdot 100$$
#
# When the above exact area of steel is provided, the balanced section generates the maximum possible moment of resistance (without providing compression steel). This moment can be calculated by the moment due to the couple generated by the compressive force in concrete and tensile force in steel.
#
# $$M_{u, lim} = 0.87 f_y A_{st, lim} \left( d - 0.416 x_{u, max} \right) = 0.36 f_{ck} b x_{u, max} \left( d - 0.416 x_{u, max} \right) $$
#
# This can be rearranged as follows:
#
# $$\frac{M_{u, lim}}{f_{ck} b d^2} = 0.36 \frac{x_{u, max}}{d} \left( 1 - 0.416 \frac{x_{u, max}}{d} \right) $$
# +
def k1(fy):
return (0.0035 / (0.0055 + 0.87 * fy / 2e5))
def k2(fy):
kk1 = k1(fy)
return (0.36 * kk1 * (1 - 0.416 * kk1))
def k3(fy):
kk1 = k1(fy)
return (36 / 0.87 * kk1)
fy = [250, 415, 500]
for ffy in fy:
print "%6d %8.3f %8.3f %8.3f" % (ffy, k1(ffy), k2(ffy), k3(ffy))
# -
# ## Under-reinforced Sections
#
# However, it is not a good idea to design a section as balanced, for the following reasons:
#
# 1. It is never possible to provide the exact amout of steel that is required by a balanced section as bar diameters are discrete and providing the required area of steel may require providing fraction of a bar, which is impossible.
# 2. If we provide more steel than that required by a balanced section, section will become over-reinforced and while it can generate the required moment of resistance, its failure is brittle, which is undesirable.
# 2. If we provide less steel than that required by a balanced section, the maximum moment of resistance it can generate is less than the design moment and hence the design is unsafe.
#
# It is therefore a better approach to provde a depth more than that required by a balanced section and provide area of steel sufficient to generate the required moment of resistance. Hence the actual depth of the neutral axis provided is $x_u > x_{u, max}$. Equating the compressive and tensile forces, we get
#
# $$\frac{p_t}{100} \cdot b d (0.87 f_y) = 0.36 f_{ck} b x_u$$
#
# Therefore
#
# $$\frac{x_u}{d} = \frac{p_t}{100} \frac{0.87 f_y}{0.36 f_{ck}} = \frac{p_t}{100} 0.4138 \frac{f_y}{f_{ck}}$$
#
# $$M_u = 0.36 f_{ck} b x_u (d - 0.416 x_u)$$
#
# $$\frac{M_u}{f_{ck} b d^2} = 0.36 \frac{x_u}{d} \left( 1 - 0.416 \frac{x_u}{d} \right)$$
#
# Substituting $\frac{x_u}{d} = \frac{p_t}{100} 0.4138 \frac{f_y}{f_{ck}}$, we get
#
# $$\frac{M_u}{0.87 f_y b d^2} = 0.$$
#
# $$\left( \frac{p_t}{100} \right)^2 - 5.8092 \frac{f_{ck}}{f_y} \frac{p_t}{100} + 6.6788 \frac{f_{ck}}{f_y^2} \frac{M_u}{b d^2} = 0$$
#
# $$\frac{p_t}{100} = 2.9046 \frac{f_{ck}}{f_y} \pm \sqrt{\left( 2.9046 \frac{f_{ck}}{f_y} \right)^2 - 6.6788 \frac{f_{ck}}{f_y^2} \frac{M_u}{b d^2}}$$
# +
def pt(Mu, b, d, fck, fy):
a = 2.9046 * fck / fy
b = math.sqrt(a**2 - (6.6788*fck*Mu / (fy**2 * b * d**2)))
return a - b
Mu = 80e6
b = 230
d =415
fck = 20
fy = 415
p = pt(Mu, b, d, fck, fy)
print p, p/100 * b * d
# -
# # Area of Concrete Stress Block
#
# Strain distribution across the depth of a cross section is assumed tb vary linearly. At the limit state of collapse in flexure, strain in the extreme compression edge is $\epsilon_{cu}=0.0035$, assuming the section as under reinforced. Stress distribution in concrete varies parabolically from $0 \leq \epsilon_c \leq \epsilon_{cy}$, where $\epsilon_{cy}=0.002$. Stress distribution in concrete is constant from $\epsilon_{cy} \leq \epsilon_c \leq \epsilon_{cu}$, where $\epsilon_{cu}=0.0035$. The yield stress in concrete is $f_{cy}= \frac{0.67}{1.5} f_{ck} = 0.446 f_{ck}$. If $x_u$ is the depth of the neautral axis from the compression edge, then depth of parabolic portion of stress block is $\frac{4}{7} x_u$ and depth of the uniform portion of the stress block is $\frac{3}{7} x_u$. In order to compute the area and centroid of the stress block, let us consider the case when depth of neutral axis below the compression edge is given as $x_u$ and we wish to compute the area and centroid of the stress block up to a distance $x$ from the neutral axis.
#
# To simplify the derivation of the equations, let us use the following notations: $0 \leq x \leq R=\frac{4}{7} x_u$ is the parabolic portion of the stress block and $R \leq x \leq \frac{3}{4} R$ is the rectangular portion of the stress block. STress starts with zero, varies parabolicaly and reaches the maximum value $h = \frac{0.67}{1.5} f_{ck} = 0.446 f_{ck}$ at $x=R$. Aftre that stress remains constant over the rectangular portion.
#
# Let us derive expressions for area of stress block and distance of centroid of stress block from neutral axis for portion of the stress block starting from $x_0 \leq x \leq (R + \frac{3}{4}R)$, where $x$ is measured from the neutral axis.
#
# \begin{align*}
# f_c(x) &=
# \begin{cases}
# h \left[ 2 \left( \frac{x}{R} \right) - \left( \frac{x}{R} \right)^2 \right] & \text{ if } 0 \leq x \leq R \\
# h & \text{ if } R \leq x \leq \frac{7}{4} R
# \end{cases}
# \end{align*}
#
# The area of the stress block for $0 \leq x \leq R$ can be written as
# $$
# A(x_0) = \int_{0}^{x_0} f_c(x) dx = h x_0 \left[\left(\frac{x_0}{R} \right) - \frac{1}{3} \left( \frac{x_0}{R} \right)^2 \right] \quad \text{ for } 0 \leq x_0 \leq R
# $$
# and first moment of area about the neutral axis can be written as
# $$
# A \bar{x} = \int_{0}^{x_0} f_c(x) x dx = h x_0^2 \left[ \frac{2}{3} \frac{x_0}{R} - \frac{1}{4} \left( \frac{x_0}{R} \right)^2 \right]
# $$
# Distance of the centroid of the parabolic portion of the stress block is given as
# $$
# \bar{x}(x_0) = \frac{A \bar{x}}{A} = x_0 \frac{\left[ \frac{2}{3} \frac{x_0}{R} - \frac{1}{4} \left( \frac{x_0}{R} \right)^2 \right]}{\left[ \frac{x_0}{R} - \frac{1}{3} \left( \frac{x_0}{R} \right)^2 \right]}
# $$
# For the particular case when $x_0 = R$, we have
# \begin{align*}
# A(R) &= \frac{2}{3} h R \\
# \bar{x}(R) &= \frac{5}{8} R
# \end{align*}
# Knowing $R=\frac{4}{7} x_u$, and $h = \frac{1}{1.5^2} f_{ck} = \left( \frac{2}{3} \right)^2 f_{ck}$, we can now find the area and centroid of the full stress block:
# \begin{align*}
# A &= \frac{3}{4} hR + \frac{2}{3} hR = \frac{17}{12} hR = \frac{68}{189} f_{ck} x_u \approx 0.36 f_{ck} x_u \\
# A \bar{x} &= \frac{3}{4} hR \left(R + \frac{1}{2} \frac{3}{4} R \right) + \frac{2}{3} hR \frac{5}{8} R = \frac{139}{96} hR^2 \\
# \bar{x} &= \frac{A \bar{x}} {A} = \frac{139}{96} \frac{12}{17} R = \frac{139}{136} R = \frac{139}{238} x_u \approx 0.584 x_u
# \end{align*}
# Distance of the centroid from the compression edge can be obtained as
# $$
# \bar{x} = x_u - \frac{139}{238} x_u = \frac{99}{238} x_u \approx 0.416 x_u
# $$
#
# Let us determine the area and centroid for the case when the a part of the stress block from the compression edge up to a distance $x_0$ from the neutral axis is considered. The equations are:
# \begin{align*}
# A &= \left( \frac{3}{4} h R \right) + \left( \frac{2}{3} hR \right) - hx_0 \left[ \left( \frac{x_0}{R} \right) - \frac{1}{3} \left( \frac{x_0}{R} \right)^2 \right] \\
# A \bar{x} &= \left( \frac{3}{4} h R \right) \left( \frac{11}{8} R \right) + \left( \frac{2}{3} hR \right) \left( \frac{5}{8} R \right) - hx_0^2 \left[ \left( \frac{x_0}{R} \right) - \frac{1}{3} \left( \frac{x_0}{R} \right)^2 \right] \frac{\left[ \frac{2}{3} \frac{x_0}{R} - \frac{1}{4} \left( \frac{x_0}{R} \right)^2 \right]}{\left[ \frac{x_0}{R} - \frac{1}{3} \left( \frac{x_0}{R} \right)^2 \right]}
# \end{align*}
#
# +
from __future__ import division, print_function
from math import pi
class ConcStressBlock(object):
def __init__(self, h, xu):
self.h = h
self.xu = xu
self.R = 4 / 7 * self.xu
def __str__(self):
s = "Concrete Stress Block, yield stress = %.3f, depth = %.2f" % (self.h, self.xu)
return s
def set_xu(self, xu):
self.xu = xu
self.R = 4.0/7.0*self.xu
return
def rect_area(self):
return 3/4*self.h*self.R
def rect_firstmom(self):
return 11/8*self.R
def par_area(self, x):
xr = x / self.R
return self.h * x * (xr - 1/3*xr**2)
def par_firstmom(self, x):
xr = x / self.R
return self.h * x * x * (2/3*xr - xr*xr/4)
def area(self, x, b=1.0):
if x > self.R:
A = self.h * (self.xu - x)
else:
A = 17/12*self.h*self.R - self.par_area(x)
return A * b
def firstmom(self, x, b=1.0):
if x > self.R:
Ax = self.h * (self.xu - x) * (self.R + (self.xu - x)/2.0)
else:
Ax = 139/96*self.h*self.R**2 - self.par_firstmom(x)
return Ax * b
def centroid_na(self, x):
A = self.area(x)
Ax = self.firstmom(x)
return Ax / A
def centroid(self, x):
return 1.0 - self.centroid_na(x)
def calc_mom_curv(b, d, dd, fck, fy, Asc, Ast, xu, ec=0.0035):
es = ec * (d - xu) / xu
esc = ec * (xu - dd) / xu
return esc, es
csb = ConcStressBlock(0.67/1.5, 1.0)
print(csb)
print(csb.par_area(4/7))
print(csb.par_firstmom(4/7))
print(csb.rect_area(), csb.rect_firstmom())
print(csb.area(0.0))
print(csb.firstmom(0))
print(csb.centroid(0), csb.centroid_na(0))
print(calc_mom_curv(230, 415, 40, 20, 415, 3*pi*16**2/4, 4*pi*20**2/4, 150, 0.0035))
# -
| rcdesign.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Vectorized String Operations
# One strength of Python is its relative ease in handling and manipulating string data.
# Pandas builds on this and provides a comprehensive set of *vectorized string operations* that become an essential piece of the type of munging required when working with (read: cleaning up) real-world data.
# In this section, we'll walk through some of the Pandas string operations, and then take a look at using them to partially clean up a very messy dataset of recipes collected from the Internet.
# ## Introducing Pandas String Operations
#
# We saw in previous sections how tools like NumPy and Pandas generalize arithmetic operations so that we can easily and quickly perform the same operation on many array elements. For example:
import numpy as np
x = np.array([2, 3, 5, 7, 11, 13])
x * 2
# This *vectorization* of operations simplifies the syntax of operating on arrays of data: we no longer have to worry about the size or shape of the array, but just about what operation we want done.
# For arrays of strings, NumPy does not provide such simple access, and thus you're stuck using a more verbose loop syntax:
data = ['peter', 'Paul', 'MARY', 'gUIDO']
[s.capitalize() for s in data]
# This is perhaps sufficient to work with some data, but it will break if there are any missing values.
# For example:
data = ['peter', 'Paul', None, 'MARY', 'gUIDO']
[s.capitalize() for s in data]
# Pandas includes features to address both this need for vectorized string operations and for correctly handling missing data via the ``str`` attribute of Pandas Series and Index objects containing strings.
# So, for example, suppose we create a Pandas Series with this data:
import pandas as pd
names = pd.Series(data)
names
# We can now call a single method that will capitalize all the entries, while skipping over any missing values:
names.str.capitalize()
# Using tab completion on this ``str`` attribute will list all the vectorized string methods available to Pandas.
# ## Tables of Pandas String Methods
#
# If you have a good understanding of string manipulation in Python, most of Pandas string syntax is intuitive enough that it's probably sufficient to just list a table of available methods; we will start with that here, before diving deeper into a few of the subtleties.
# The examples in this section use the following series of names:
monte = pd.Series(['<NAME>', '<NAME>', '<NAME>',
'<NAME>', '<NAME>', '<NAME>'])
# ### Methods similar to Python string methods
# Nearly all Python's built-in string methods are mirrored by a Pandas vectorized string method. Here is a list of Pandas ``str`` methods that mirror Python string methods:
#
# | | | | |
# |-------------|------------------|------------------|------------------|
# |``len()`` | ``lower()`` | ``translate()`` | ``islower()`` |
# |``ljust()`` | ``upper()`` | ``startswith()`` | ``isupper()`` |
# |``rjust()`` | ``find()`` | ``endswith()`` | ``isnumeric()`` |
# |``center()`` | ``rfind()`` | ``isalnum()`` | ``isdecimal()`` |
# |``zfill()`` | ``index()`` | ``isalpha()`` | ``split()`` |
# |``strip()`` | ``rindex()`` | ``isdigit()`` | ``rsplit()`` |
# |``rstrip()`` | ``capitalize()`` | ``isspace()`` | ``partition()`` |
# |``lstrip()`` | ``swapcase()`` | ``istitle()`` | ``rpartition()`` |
#
# Notice that these have various return values. Some, like ``lower()``, return a series of strings:
monte.str.lower()
# But some others return numbers:
monte.str.len()
# Or Boolean values:
monte.str.startswith('T')
# Still others return lists or other compound values for each element:
monte.str.split()
# We'll see further manipulations of this kind of series-of-lists object as we continue our discussion.
# ### Methods using regular expressions
#
# In addition, there are several methods that accept regular expressions to examine the content of each string element, and follow some of the API conventions of Python's built-in ``re`` module:
#
# | Method | Description |
# |--------|-------------|
# | ``match()`` | Call ``re.match()`` on each element, returning a boolean. |
# | ``extract()`` | Call ``re.match()`` on each element, returning matched groups as strings.|
# | ``findall()`` | Call ``re.findall()`` on each element |
# | ``replace()`` | Replace occurrences of pattern with some other string|
# | ``contains()`` | Call ``re.search()`` on each element, returning a boolean |
# | ``count()`` | Count occurrences of pattern|
# | ``split()`` | Equivalent to ``str.split()``, but accepts regexps |
# | ``rsplit()`` | Equivalent to ``str.rsplit()``, but accepts regexps |
# With these, you can do a wide range of interesting operations.
# For example, we can extract the first name from each by asking for a contiguous group of characters at the beginning of each element:
monte.str.extract('([A-Za-z]+)', expand=False)
# Or we can do something more complicated, like finding all names that start and end with a consonant, making use of the start-of-string (``^``) and end-of-string (``$``) regular expression characters:
monte.str.findall(r'^[^AEIOU].*[^aeiou]$')
# The ability to concisely apply regular expressions across ``Series`` or ``Dataframe`` entries opens up many possibilities for analysis and cleaning of data.
# ### Miscellaneous methods
# Finally, there are some miscellaneous methods that enable other convenient operations:
#
# | Method | Description |
# |--------|-------------|
# | ``get()`` | Index each element |
# | ``slice()`` | Slice each element|
# | ``slice_replace()`` | Replace slice in each element with passed value|
# | ``cat()`` | Concatenate strings|
# | ``repeat()`` | Repeat values |
# | ``normalize()`` | Return Unicode form of string |
# | ``pad()`` | Add whitespace to left, right, or both sides of strings|
# | ``wrap()`` | Split long strings into lines with length less than a given width|
# | ``join()`` | Join strings in each element of the Series with passed separator|
# | ``get_dummies()`` | extract dummy variables as a dataframe |
# #### Vectorized item access and slicing
#
# The ``get()`` and ``slice()`` operations, in particular, enable vectorized element access from each array.
# For example, we can get a slice of the first three characters of each array using ``str.slice(0, 3)``.
# Note that this behavior is also available through Python's normal indexing syntax–for example, ``df.str.slice(0, 3)`` is equivalent to ``df.str[0:3]``:
monte.str[0:3]
# Indexing via ``df.str.get(i)`` and ``df.str[i]`` is likewise similar.
#
# These ``get()`` and ``slice()`` methods also let you access elements of arrays returned by ``split()``.
# For example, to extract the last name of each entry, we can combine ``split()`` and ``get()``:
monte.str.split().str.get(-1)
# #### Indicator variables
#
# Another method that requires a bit of extra explanation is the ``get_dummies()`` method.
# This is useful when your data has a column containing some sort of coded indicator.
# For example, we might have a dataset that contains information in the form of codes, such as A="born in America," B="born in the United Kingdom," C="likes cheese," D="likes spam":
full_monte = pd.DataFrame({'name': monte,
'info': ['B|C|D', 'B|D', 'A|C',
'B|D', 'B|C', 'B|C|D']})
full_monte
# The ``get_dummies()`` routine lets you quickly split-out these indicator variables into a ``DataFrame``:
full_monte['info'].str.get_dummies('|')
# With these operations as building blocks, you can construct an endless range of string processing procedures when cleaning your data.
#
# We won't dive further into these methods here, but I encourage you to read through ["Working with Text Data"](http://pandas.pydata.org/pandas-docs/stable/text.html) in the Pandas online documentation.
# ## Example: Recipe Database
#
# These vectorized string operations become most useful in the process of cleaning up messy, real-world data.
# Here I'll walk through an example of that, using an open recipe database compiled from various sources on the Web.
# Our goal will be to parse the recipe data into ingredient lists, so we can quickly find a recipe based on some ingredients we have on hand.
#
# The scripts used to compile this can be found at https://github.com/fictivekin/openrecipes, and the link to the current version of the database is found there as well.
#
# As of Spring 2016, this database is about 30 MB, and can be downloaded and unzipped with these commands:
# !curl -O http://openrecipes.s3.amazonaws.com/recipeitems-latest.json.gz
# !gunzip recipeitems-latest.json.gz
# The database is in JSON format, so we will try ``pd.read_json`` to read it:
try:
recipes = pd.read_json('recipeitems-latest.json')
except ValueError as e:
print("ValueError:", e)
# Oops! We get a ``ValueError`` mentioning that there is "trailing data."
# Searching for the text of this error on the Internet, it seems that it's due to using a file in which *each line* is itself a valid JSON, but the full file is not.
# Let's check if this interpretation is true:
with open('recipeitems-latest.json') as f:
line = f.readline()
pd.read_json(line).shape
# Yes, apparently each line is a valid JSON, so we'll need to string them together.
# One way we can do this is to actually construct a string representation containing all these JSON entries, and then load the whole thing with ``pd.read_json``:
# read the entire file into a Python array
with open('recipeitems-latest.json', 'r') as f:
# Extract each line
data = (line.strip() for line in f)
# Reformat so each line is the element of a list
data_json = "[{0}]".format(','.join(data))
# read the result as a JSON
recipes = pd.read_json(data_json)
recipes.shape
# We see there are nearly 200,000 recipes, and 17 columns.
# Let's take a look at one row to see what we have:
recipes.iloc[0]
# There is a lot of information there, but much of it is in a very messy form, as is typical of data scraped from the Web.
# In particular, the ingredient list is in string format; we're going to have to carefully extract the information we're interested in.
# Let's start by taking a closer look at the ingredients:
recipes.ingredients.str.len().describe()
# The ingredient lists average 250 characters long, with a minimum of 0 and a maximum of nearly 10,000 characters!
#
# Just out of curiousity, let's see which recipe has the longest ingredient list:
recipes.name[np.argmax(recipes.ingredients.str.len())]
# That certainly looks like an involved recipe.
#
# We can do other aggregate explorations; for example, let's see how many of the recipes are for breakfast food:
recipes.description.str.contains('[Bb]reakfast').sum()
# Or how many of the recipes list cinnamon as an ingredient:
recipes.ingredients.str.contains('[Cc]innamon').sum()
# We could even look to see whether any recipes misspell the ingredient as "cinamon":
recipes.ingredients.str.contains('[Cc]inamon').sum()
# This is the type of essential data exploration that is possible with Pandas string tools.
# It is data munging like this that Python really excels at.
# ### A simple recipe recommender
#
# Let's go a bit further, and start working on a simple recipe recommendation system: given a list of ingredients, find a recipe that uses all those ingredients.
# While conceptually straightforward, the task is complicated by the heterogeneity of the data: there is no easy operation, for example, to extract a clean list of ingredients from each row.
# So we will cheat a bit: we'll start with a list of common ingredients, and simply search to see whether they are in each recipe's ingredient list.
# For simplicity, let's just stick with herbs and spices for the time being:
spice_list = ['salt', 'pepper', 'oregano', 'sage', 'parsley',
'rosemary', 'tarragon', 'thyme', 'paprika', 'cumin']
# We can then build a Boolean ``DataFrame`` consisting of True and False values, indicating whether this ingredient appears in the list:
import re
spice_df = pd.DataFrame(dict((spice, recipes.ingredients.str.contains(spice, re.IGNORECASE))
for spice in spice_list))
spice_df.head()
# Now, as an example, let's say we'd like to find a recipe that uses parsley, paprika, and tarragon.
# We can compute this very quickly using the ``query()`` method of ``DataFrame``s, discussed in *High-Performance Pandas: ``eval()`` and ``query()``*:
selection = spice_df.query('parsley & paprika & tarragon')
len(selection)
# We find only 10 recipes with this combination; let's use the index returned by this selection to discover the names of the recipes that have this combination:
recipes.name[selection.index]
# Now that we have narrowed down our recipe selection by a factor of almost 20,000, we are in a position to make a more informed decision about what we'd like to cook for dinner.
# ### Going further with recipes
#
# Hopefully this example has given you a bit of a flavor (ba-dum!) for the types of data cleaning operations that are efficiently enabled by Pandas string methods.
# Of course, building a very robust recipe recommendation system would require a *lot* more work!
# Extracting full ingredient lists from each recipe would be an important piece of the task; unfortunately, the wide variety of formats used makes this a relatively time-consuming process.
# This points to the truism that in data science, cleaning and munging of real-world data often comprises the majority of the work, and Pandas provides the tools that can help you do this efficiently.
| notebooks/Python-in-2-days/D1_L5_Pandas/10-Working-With-Strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from xmlr import xmlparse
from xmlr import xmliter
from xmlr import xmliter, XMLParsingMethods
import xml.etree.ElementTree
count=0
images=[]
for d in xmliter('/home/gnanesh/Downloads/theodore_plus_training.xml','image'):
for k, v in d.items():
if k == '@name':
images.append(v)
if k == 'box':
for x in v:
if x["@label"] == 'person':
print (v )
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
# -
| data_extra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
import pandas as pd
df = pd.read_csv('../datasets/UN.csv')
print('----')
# print the raw column information plus summary header
print(df)
print('----')
# look at the types of each column explicitly
print('Individual columns - Python data types')
[(x, type(df[x][0])) for x in df.columns]
| K-means/Exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pantelisziazo/various_projects/blob/main/Sentiment_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="PyVXrhRLqQby"
# # Introduction
#
# + [markdown] id="7X6OgsjYx6TE"
# ## Overview
# + [markdown] id="2mMQlobfqqXy"
# In this script I apply **Sentiment Analysis** over a specific (*non-public available*) dataset.
#
# The main purpose of this project is to make a classifer that **reads a tweet** and finds the **sentiment** of the it (e.g. if the tweet expresses something negative or has neutral sentiment).
#
# + [markdown] id="zMT3pLBvrkDp"
# ## More information
#
#
# + [markdown] id="zUrqwEBRyAyE"
# * My objective is to write a script that is **affordable to everyone** : runs quite fast, is easy to read and understandable. That means that I will make use of [Colab](https://colab.research.google.com/) platform, do not make extensive analysis of the data (but provide information about future improvements) and write as clear code as possible.
# * I will try to keep the *workload* needed for this task to less than the workload of a working day.
# * The dataset includes sensitive data at some cases. So the appearance of it is going to be *blurred*.
# * For any question, please contact me through email at **<EMAIL>** .
#
# + [markdown] id="-gob84PUxwtL"
# # Classification
# + [markdown] id="WOSm1neBstVD"
# ## Build the appropriate environment and take a first look at the data.
# + id="hY6FjouKUUcC"
# import all the necessary libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from collections import Counter
import seaborn as sns
import datetime
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, f1_score
from scipy import sparse
# + id="-k4yvkyHUoNw"
# read the .json file with the data
data = pd.read_json('/content/palo_assignment.json', lines =True)
# + colab={"base_uri": "https://localhost:8080/"} id="L5ryqRsdU60p" outputId="12b41ddd-2011-4763-cc7e-4c0857abe372"
# the command below is to make blurry the appearance of the pandas Dataframes
def blurry(s):
return 'color: transparent; text-shadow: 0 0 5px rgba(0,0,0,0.5)'
# data.head().style.applymap(blurry)
# unfortunately the github does not support this command
data.info()
# + [markdown] id="0NOVXgWDqVPq"
# The data is :
#
# * **id** : the index of the tweet. It does not provide a special information for the classification process that is why this column will be left out.
# * **channel** : the source of the text phrase. All the texts are tweets (come from Twitter) so this column does not hold special information (will be left out).
# * **text** : the text phrase of a tweet. The main "x-value" of our classification problem.
# * **createdate** : datetime variable at the format "Year-Month-Day Hour-Minutes-Seconds" .
# * **impact** : a numeric variable that takes values from 0 to 1 .
# * **query_name** : the bank name that each tweet refers to.
# * **sentiment** : the sentiment of the tweet, the "y-value" of the classification problem, the variable that we want to predict.
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="KV5rN5jSWLuf" outputId="aa1da52f-372e-412e-c255-ef859e2078b4"
# histogram of the possible sentiments (the y-value)
plt.figure(figsize=(6,4))
plt.hist(data['sentiment'])
plt.title('Distribution of Sentiment Labels')
plt.xlabel('Sentiments')
plt.ylabel('Number of examples per sentiment')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="t2FWAVpzYnu0" outputId="0908f2fc-8703-4719-cc7f-2e75809d93dc"
# get the exact number of each category
Counter(data['sentiment'])
# + [markdown] id="ebvy23VvY3Rv"
# As we see the sentiment variable of the dataset is quite unbalnced. Having only 42 examples of the category 'Positive' when the other 2 have over 2000 . That is a problem that needs special treatment.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="q7ilyQN2WU9h" outputId="7bbc81d2-b130-4b0c-8cda-65670a04a24d"
# histogram of the possible Banks the tweets are reffered to
plt.figure(figsize=(6,4))
plt.hist(data['query_name'])
plt.title('Distribution of Reffered Banks')
plt.xlabel('Bank Names')
plt.ylabel('Number of examples per Bank')
plt.show()
# + [markdown] id="fylqBRdww4Zp"
# ## Get an insight of the dataset
# + [markdown] id="55vF1WWfw83R"
# The dataset seems quite interesting and easy to extract information.
# Here I will try to answer some questions, like :
#
# * Which was the month with the most negative sentiment ?
# * Which was the bank with the most negative tweets at the above month ?
# * Any other correlation between the data ?
#
#
# + id="hOouRQNgXGF-" colab={"base_uri": "https://localhost:8080/", "height": 296} outputId="7a856595-7c88-436a-f1d0-cbb96912e062"
# read only the day , not the time of the 'createdate' column
dates = [date.split(" ")[0] for date in data['createdate']]
# parse the 'Month' of each tweet
months = [datetime.datetime.strptime(month, "%Y-%m-%d").month for month in dates]
# make a heatmap
sns.heatmap(pd.crosstab(pd.Series(months, name='Month Number'),
pd.Series(list(data['sentiment']), name='Sentiment')), annot=True,fmt=".1f")
# + colab={"base_uri": "https://localhost:8080/", "height": 350} id="RB3ky3F8nDpO" outputId="80b2ef74-66c2-4e35-e940-bf624fa012eb"
# append the Months to the dataframe
data['Month'] = months
# get the data that refer to month with the most negative sentiment (7)
most_negative_month_data = data[data['Month'] == 7]
bank_most_negative_month_data = most_negative_month_data[most_negative_month_data['sentiment'] == 'negative']
plt.figure(figsize=(7,5))
plt.hist(bank_most_negative_month_data['query_name'])
plt.title('Distribution of Reffered Banks at the Month with the most negative sentiment tweets')
plt.xlabel('Bank Names')
plt.ylabel('Number of examples per Bank')
plt.show()
plt.show()
# + id="iDofHLdtnHOq" colab={"base_uri": "https://localhost:8080/", "height": 296} outputId="cbfab534-2a98-4975-d95f-7c10607da92e"
# additional correlation betweenthe data
# correlation between the banks and the sentiment
sns.heatmap(pd.crosstab(pd.Series(list(data['query_name']), name='Banks'),
pd.Series(list(data['sentiment']), name='Sentiment')), annot=True,fmt=".1f")
# + colab={"base_uri": "https://localhost:8080/"} id="nVW3XUE92CaH" outputId="7d4e013f-08c5-4d3d-debe-60404ec20ff0"
# correlation of the banks with the impact variable
data['query_name'].astype('category').cat.codes.corr(data['impact'])
# almost 0 -> possibly does not hold a great insight if we wanted to predict the bank names
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="LPBd5O-40nDM" outputId="6c7067a5-5a9d-4026-e4af-c7b10ea450b5"
# correlation and scatter plot of the sentiment with the impact varible
print(data['sentiment'].astype('category').cat.codes.corr(data['impact']))
# different from zero -> the classification results may be better if we include this variable too
plt.scatter(x = data['sentiment'], y = data['impact'])
plt.show()
# + [markdown] id="kcdKfEFUIYVP"
# ## Final pre-processing of the data
# + id="cQ9REAIrpNab"
# convert the string variables to numerical in order to be inserted to the classifier
data['query_number'] = data['query_name'].astype('category')
data['query_number'] = data['query_number'].cat.codes
# + [markdown] id="9QREtRRW9Qd-"
# Since the dataset is unbalnced and I want to find a quick solution, I am going to **left out** the examples of the **'Positive' sentiment** category in order to **simplify the problem**.
#
# Someone else could apply oversampling over the 'Positive' category at the train subset only in order to increase the examples of the 'Positive' category.
#
# In the case that the dataset had more examples but with the same unbalance as here, someone could apply indersampling over the 2 other categories ('Negative' and 'Neutral') over the training subset only.
# + id="70egqWU59RHv"
# drop the unnecessary data
# if you still prefer to hold the 'positive' category just omit the command-line below
# (and also re-define the calculation of f1 score )
data.drop(data[data['sentiment'] == 'positive'].index, inplace=True)
# these 2 columns do not hold information
data = data.drop(labels=['id','channel'], axis=1)
# + id="O_9iqnyxuP96"
y_data = data['sentiment']
x_data = data.drop(['sentiment'], axis=1)
# split the dataset to train, development and test subsets
x_nontest, x_test, y_nontest, y_test = train_test_split(x_data, y_data, random_state= 3, test_size=0.15)
x_train, x_dev, y_train, y_dev = train_test_split(x_nontest, y_nontest, random_state= 3, test_size=0.2)
# + id="_K2o_OQTAnug"
# define a function that concatenates the tfidf matrix with another column of the dataset
def matrix_plus_variables(matrix,variable_data):
matrix_array = matrix.toarray()
output = []
for i in range(len(variable_data)):
output.append(np.append(matrix_array[i],variable_data[i]))
return sparse.csr_matrix(output)
# + [markdown] id="Ku6ylBOWIe9z"
# ## Hyper-parameter tuning
# + id="9SqlMMGju5xE"
# define the hyperparameter of the problem
ngram_range = [1,2]
max_features = [100,500]
max_tf = [0.75,0.9]
min_tf = [5,10]
regularization_strength = [1, 0.1, 0.01, 0.001]
# + id="rxT2Rnovu5z5"
accuracies, f_scores = [], []
best_ac , best_f1 = 0,0
for a in ngram_range:
for b in max_features:
for c in max_tf:
for d in min_tf:
# turn text to tfidf embeddings
vectorizer = TfidfVectorizer(ngram_range=(1, a), max_features = b, max_df= c, min_df= d)
xtrain_tfidf = vectorizer.fit_transform(x_train['text'])
xdev_tfidf = vectorizer.transform(x_dev['text'])
# concatenate the 'bank name' variable to the future input of the classifier
x_train_data = matrix_plus_variables(xtrain_tfidf,list(x_train['query_number']))
x_dev_data = matrix_plus_variables(xdev_tfidf,list(x_dev['query_number']))
# concatenate the 'impact' variable to the future input of the classifier
x_train_data = matrix_plus_variables(x_train_data,list(x_train['impact']))
x_dev_data = matrix_plus_variables(x_dev_data,list(x_dev['impact']))
# define the classifier
for e in regularization_strength:
logistic = LogisticRegression(C=e)
logistic.fit(x_train_data, y_train)
predictions = logistic.predict(x_dev_data)
# calculate the scores
ac = accuracy_score(y_dev, predictions)
accuracies.append(ac)
f1 = f1_score(y_dev,predictions, pos_label='negative')
f_scores.append(f1)
# hold the best scoring parameters
if ac > best_ac:
best_ac = ac
ac_ngram, ac_maxfeatures, ac_maxtf, ac_mintf, ac_regularization = a,b,c,d,e
if f1 > best_f1:
best_f1 = f1
f1_ngram, f1_maxfeatures, f1_maxtf, f1_mintf, f1_regularization = a,b,c,d,e
# + colab={"base_uri": "https://localhost:8080/"} id="XSLTeJ9mGL1t" outputId="9ce15a9a-3945-476d-ecb2-594ffab8dc05"
# print the highest scoring parameters
for i in [ac_ngram, ac_maxfeatures, ac_maxtf, ac_mintf, ac_regularization]:
print(i)
print("Best accuracy at hyperparameter tuning = ", best_ac)
print()
print()
for i in [f1_ngram, f1_maxfeatures, f1_maxtf, f1_mintf, f1_regularization]:
print(i)
print("Best F1 score at hyperparameter tuning = ", best_f1)
# + [markdown] id="--SXef8tIkT2"
# ## Final Model
# + id="3-W7ro5B-7Td" colab={"base_uri": "https://localhost:8080/"} outputId="72ee5baf-80ec-4cc7-844e-0a7373c40cc3"
# highest scoring accuracy model
# follow the same process as in the hyperparameter tuning
ac_vectorizer = TfidfVectorizer(ngram_range=(1, ac_ngram), max_features = ac_maxfeatures,
max_df= ac_maxtf, min_df= ac_mintf)
xnontest_tfidf = ac_vectorizer.fit_transform(x_nontest['text'])
xtest_tfidf = ac_vectorizer.transform(x_test['text'])
x_nontest_data = matrix_plus_variables(xnontest_tfidf,list(x_nontest['query_number']))
x_test_data = matrix_plus_variables(xtest_tfidf,list(x_test['query_number']))
x_nontest_data = matrix_plus_variables(x_nontest_data,list(x_nontest['impact']))
x_test_data = matrix_plus_variables(x_test_data,list(x_test['impact']))
ac_logistic = LogisticRegression(C = ac_regularization)
ac_logistic.fit(x_nontest_data, y_nontest)
ac_predictions = ac_logistic.predict(x_test_data)
final_ac = accuracy_score(y_test, ac_predictions)
print("Final Accuracy = ", final_ac)
final_f1 = f1_score(y_test, ac_predictions, pos_label="negative")
print("Final F1 score = ", final_f1)
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="V4R7WqNUroQc" outputId="f2a72450-1b43-4838-eb84-8eb77848a842"
# a heatmap of the true - predicted values
sns.heatmap(pd.crosstab(pd.Series(ac_predictions, name='Predictions'),
pd.Series(list(y_test), name='Truth')), annot=True,fmt=".1f")
# + [markdown] id="GgCuOUuVM_Km"
# # Additional information regarding TF-IDF scores of the dataset
# + [markdown] id="bSAsCww5tWru"
# Try to find the top N n-grams (unigrams, bigrams, trigrams) according to tf-idf scores
# + id="v3-HPsdHr4YV"
data = pd.read_json('/content/palo_assignment.json', lines =True)
def get_top_ngrams(ngram_range = 1 , top_n = 20, text = data['text']):
tfidf_vectorizer = TfidfVectorizer(ngram_range=(ngram_range, ngram_range), min_df=10) # keep fewer features
# get the ngrams to array matrix
ngrams = tfidf_vectorizer.fit_transform(text).toarray()
# get the name of the features
feature_names = tfidf_vectorizer.get_feature_names()
# make a dataframe with the tfidf score of each feature for each example
tfidf_data = pd.DataFrame(ngrams, columns = feature_names)
# sort the above dataframe
sorted_tfidf_dataframe = pd.DataFrame(np.sort(tfidf_data.values, axis=0), index=tfidf_data.index, columns=tfidf_data.columns)
# keep the highest scoring line
top_tfidf_score = sorted_tfidf_dataframe.tail(1)
# get the indices of the highest tfidf scoring words
top_n_indices = np.argsort(top_tfidf_score.values)[0]
top_n_values = []
# locate the words and the tfidf score
for i in top_n_indices[-top_n:]:
top_n_values.append(top_tfidf_score.iloc[:,i])
return top_n_values
# + colab={"base_uri": "https://localhost:8080/"} id="uCL6IitVndO-" outputId="b65f7437-24af-4e55-a579-7a46b4fea1e8"
# for 1-gram , top 20
answer1 = get_top_ngrams(ngram_range=1,top_n=20)
answer1.reverse()
for i in answer1:
print(i.values, "\t", i.name)
# + colab={"base_uri": "https://localhost:8080/"} id="wQjjVqneeOFH" outputId="e9d6a84e-e4b8-4b93-9a9a-f6b9cbf59db2"
# for 2-grams , top 10
answer2 = get_top_ngrams(ngram_range=2,top_n=10)
answer2.reverse()
for i in answer2:
print(i.values, "\t", i.name)
# + colab={"base_uri": "https://localhost:8080/"} id="SCFhiWK-oOIu" outputId="c93e57c6-dc79-41b3-af2a-3bac7535fcc0"
# for 3-grams , top 10
answer3 = get_top_ngrams(ngram_range=3,top_n=10)
answer3.reverse()
for i in answer3:
print(i.values, "\t", i.name)
# + [markdown] id="NoPj2GvLNOSB"
# # Conclusion
# + [markdown] id="uF4g2TQvq2tJ"
# ## Summary
# + [markdown] id="4jpIeTz4qr0H"
# In order to complete the task above I worked 5 hours, wanting to keep the workload at the levels of a single working day.
#
# The process I followed :
#
# * Take a look at the dataset and get a quick insight of it
# * Decide the variables I will include to the classification process
# * Build (tune and train) a classifier
#
#
# + [markdown] id="8ypCMeB7ripn"
# It is obvious that the results are promising and have margins for improvement.
# + [markdown] id="VhGSREJJrgIe"
# ## Future Work
#
# + [markdown] id="csbLRd5muyOo"
# The most promising ways to improve this classifier are:
#
# * Insert the datetime variable to the problem. The hour that each tweet is posted may hold information about the sentiment.
# * Keep the 'positive' category and apply oversampling at the train set of the task.
# * Exclude stopwords of the text while applying the Tf-Idf vectorizer.
# * Make more extensive hyper-paramter tuning.
# * Try more complex classifiers, like a MLP.
#
#
# ---
#
# I hope you find this script understandable and funny to read.
#
#
| Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] inputHidden=false outputHidden=false
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# -
# #### Version Check
# Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
# + inputHidden=false outputHidden=false
import plotly
plotly.__version__
# -
# ### Basic Carpet Plot
# + inputHidden=false outputHidden=false
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [4, 4, 4, 4.5, 4.5, 4.5, 5, 5, 5, 6, 6, 6],
b = [1, 2, 3, 1, 2, 3, 1, 2, 3, 1, 2, 3],
y = [2, 3.5, 4, 3, 4.5, 5, 5.5, 6.5, 7.5, 8, 8.5, 10],
aaxis = dict(
tickprefix = 'a = ',
ticksuffix = 'm',
smoothing = 1,
minorgridcount = 9
),
baxis = dict(
tickprefix = 'b = ',
ticksuffix = 'Pa',
smoothing = 1,
minorgridcount = 9
)
)
data = [trace1]
fig = go.Figure(data = data)
py.iplot(fig, filename = "scattercarpet/basic")
# -
# ### Add Carpet Scatter Trace
# + inputHidden=false outputHidden=false
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [4, 4, 4, 4.5, 4.5, 4.5, 5, 5, 5, 6, 6, 6],
b = [1, 2, 3, 1, 2, 3, 1, 2, 3, 1, 2, 3],
y = [2, 3.5, 4, 3, 4.5, 5, 5.5, 6.5, 7.5, 8, 8.5, 10],
aaxis = dict(
tickprefix = 'a = ',
ticksuffix = 'm',
smoothing = 1,
minorgridcount = 9
),
baxis = dict(
tickprefix = 'b = ',
ticksuffix = 'Pa',
smoothing = 1,
minorgridcount = 9
)
)
trace2 = go.Scattercarpet(
a = [4, 4.5, 5, 6],
b = [2.5, 2.5, 2.5, 2.5],
line = dict(
shape = 'spline',
smoothing = 1,
color = 'blue'
)
)
data = [trace1,trace2]
fig = go.Figure(data = data)
py.iplot(fig, filename = "scattercarpet/add-scattercarpet")
# -
# ### Add Multiple Scatter Traces
# + inputHidden=false outputHidden=false
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [0.1,0.2,0.3],
b = [1,2,3],
y = [[1,2.2,3],[1.5,2.7,3.5],[1.7,2.9,3.7]],
cheaterslope = 1,
aaxis = dict(
title = "a",
tickmode = "linear",
dtick = 0.05
),
baxis = dict(
title = "b",
tickmode = "linear",
dtick = 0.05
)
)
trace2 = go.Scattercarpet(
name = "b = 1.5",
a = [0.05, 0.15, 0.25, 0.35],
b = [1.5, 1.5, 1.5, 1.5]
)
trace3 = go.Scattercarpet(
name = "b = 2",
a = [0.05, 0.15, 0.25, 0.35],
b = [2, 2, 2, 2]
)
trace4 = go.Scattercarpet(
name = "b = 2.5",
a = [0.05, 0.15, 0.25, 0.35],
b = [2.5, 2.5, 2.5, 2.5]
)
trace5 = go.Scattercarpet(
name = "a = 0.15",
a = [0.15, 0.15, 0.15, 0.15],
b = [0.5, 1.5, 2.5, 3.5],
line = dict(
smoothing = 1,
shape = "spline"
)
)
trace6 = go.Scattercarpet(
name = "a = 0.2",
a = [0.2, 0.2, 0.2, 0.2],
b = [0.5, 1.5, 2.5, 3.5],
line = dict(
smoothing = 1,
shape = "spline"
),
marker = dict(
size = [10, 20, 30, 40],
color = ["#000", "#f00", "#ff0", "#fff"]
)
)
trace7 = go.Scattercarpet(
name = "a = 0.25",
a = [0.25, 0.25, 0.25, 0.25],
b = [0.5, 1.5, 2.5, 3.5],
line = dict(
smoothing = 1,
shape = "spline"
)
)
layout = go.Layout(
title = "scattercarpet extrapolation, clipping, and smoothing",
hovermode = "closest"
)
data = [trace1,trace2,trace3,trace4,trace5,trace6,trace7]
fig = go.Figure(data = data, layout = layout)
py.iplot(fig, filename = "scattercarpet/multiple")
# -
# ### Reference
# See https://plotly.com/python/reference/#scattercarpet for more information and chart attribute options!
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'scattercarpet.ipynb', 'python/carpet-scatter/', 'Carpet Scatter Plot',
'How to make carpet scatter plots in Python with Plotly.',
title = 'Carpet Scatter Plots | Plotly',
has_thumbnail='true', thumbnail='thumbnail/scattercarpet.jpg',
language='python',
# page_type='example_index', // note this is only if you want the tutorial to appear on the main page: plot.ly/python
display_as='scientific', order=28,
ipynb= '~notebook_demo/146')
# -
| _posts/python-v3/scientific/carpet-scatter/scattercarpet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color= 'Green'>Optimal System Identification for LIGO</font>
# ### of linear, time-invariant (LTI) systems
# ***
# * the LIGO Control Systems Working Group wiki: https://wiki.ligo.org/CSWG/OptTF
# * Rana's public GitHub page on LIGO Controls problems: https://github.com/rxa254/LIGO-Controls-Problems
# ***
# This notebook is meant to give an introduction to a couple of kinds of sysID problems in LIGO. The goal is to generate a few specific strategies to do this better for important cases, and eventually to make a more general tool for this.
# # Overview
# ## The Identification Problem
# We would like to know what our physical plants (optics, suspensions, electronics, cavities) are doing. In nearly all cases, we need not consider the nonlinearity or time-dependence of the plant (notable exceptions due to thermal processes and slow drift and changes in laser power).
#
# We approach this problem by making Transfer Functions (TF) of the system that we are interested in.
#
# How to make a TF measurement:
# 1. with enough SNR at the frequencies of interest
# 1. without saturating the actuators too much
# 1. within a small enough amount of time (so that our precious commissioning / observation time is not squandered)
# + jupyter={"outputs_hidden": false}
# %matplotlib inline
# Import packages.
import matplotlib.pyplot as plt
import scipy.signal as sig
import numpy as np
# +
# Let's define the system to be 'identified'
fs = 1024
# pendulum
zz = []
f_p = 1
theta_p = 60
pp = [f_p * np.exp(1j * theta_p * np.pi/180)]
pp = [pp[0], np.conj(pp[0])]
pp = np.array(pp)
pend = sig.ZerosPolesGain(zz, pp, 1)
sos = sig.zpk2sos(zz, pp/(fs/2), 1)
# + jupyter={"outputs_hidden": false}
# filter some white noise using this filter
dur = 64
x = np.random.randn(fs*dur)
np.shape(x)
# -
# ## Conclusion
#
# * Need help in writing the code to do this
# ## References
#
# The Sys ID book by <NAME> Shoukens:
# https://books.google.com/books?id=3lGJWtjGDzsC
#
# SysID classroom exercises:
# https://caltech.tind.io/record/748967?ln=en
#
# How to take the frequency response measurement and find the plant parameters:
#
# ["Parameter Estimation and Model Order Identification of LTI Systems"](https://lmgtfy.com/?q=10.0.3.248%2Fj.ifacol.2016.07.333)
#
# How to estimate the covariance matrix:
#
# How to iterate the multi-sine excitation waveform based on the matrix:
| OptimalSysID/optimalTF_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # Poster popularity by country
#
# This notebook loads data of poster viewership at the SfN 2016 annual meeting, organized by the countries that were affiliated with each poster.
#
# We find that the poster popularity across countries is not significant compare to what is expected by chance.
# ### Import libraries and load data
# +
# %config InlineBackend.figure_format = 'retina'
# %matplotlib inline
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('white')
import pandas as pd
# -
# Load data
df = pd.DataFrame.from_csv('./posterviewers_by_country.csv')
key_N = 'Number of people'
# # 1. Summarize data by country
# 0. Count number of posters from each state
# Calculate mean poster popularity
states = df['Country'].unique()
dict_state_counts = {'Country':states,'count':np.zeros(len(states)),'popularity':np.zeros(len(states))}
for i, s in enumerate(states):
dict_state_counts['count'][i] = int(sum(df['Country']==s))
dict_state_counts['popularity'][i] = np.round(np.mean(df[df['Country']==s][key_N]),3)
df_counts = pd.DataFrame.from_dict(dict_state_counts)
# Visualize dataframe
# count = total number of posters counted affiliated with that country
# popularity = average number of viewers at a poster affiliated with that country
df_counts.head()
# # 2. Poster popularity vs. prevalence
# Across states in the United States, we found a positive correlation between the number of posters from a state and the popularity of those posters. We debatably see this again across countries to a trending level of significance (1-tailed p-value = 0.06)
print sp.stats.spearmanr(np.log10(df_counts['count']),df_counts['popularity'])
plt.figure(figsize=(3,3))
plt.semilogx(df_counts['count'],df_counts['popularity'],'k.')
plt.xlabel('Number of posters\nin the state')
plt.ylabel('Average number of viewers per poster')
plt.ylim((-.1,3.6))
plt.xlim((.9,1000))
# # 3. Permutation tests: difference in popularity across countries
# In this code, we test if the relative popularity / unpopularity observed for any country is outside what is expected by chance
#
# Here, the most popular and least popular countries are defined by a nonparametric statiscal test between the number of viewers at posters from their country, compared to posters from all other countries.
# Simulate randomized data
Nperm = 100
N_posters = len(df)
rand_statepop = np.zeros((Nperm,len(states)),dtype=np.ndarray)
rand_statepopmean = np.zeros((Nperm,len(states)))
for i in range(Nperm):
# Random permutation of posters, organized by state
randperm_viewers = np.random.permutation(df[key_N].values)
for j, s in enumerate(states):
rand_statepop[i,j] = randperm_viewers[np.where(df['Country']==s)[0]]
rand_statepopmean[i,j] = np.mean(randperm_viewers[np.where(df['Country']==s)[0]])
# +
# True data: Calculate all p-values for the difference between 1 state's popularity and the rest
min_N_posters = 10
states_big = states[np.where(df_counts['count']>=min_N_posters)[0]]
N_big = len(states_big)
t_true_all = np.zeros(N_big)
p_true_all = np.zeros(N_big)
for i, state in enumerate(states_big):
t_true_all[i], _ = sp.stats.ttest_ind(df[df['Country']==state][key_N],df[df['Country']!=state][key_N])
_, p_true_all[i] = sp.stats.mannwhitneyu(df[df['Country']==state][key_N],df[df['Country']!=state][key_N])
pmin_pop = np.min(p_true_all[np.where(t_true_all>0)[0]])
pmin_unpop = np.min(p_true_all[np.where(t_true_all<0)[0]])
print 'Most popular country: ', states_big[np.argmax(t_true_all)], '. p=', str(pmin_pop)
print 'Least popular country: ', states_big[np.argmin(t_true_all)], '. p=', str(pmin_unpop)
# +
# Calculate minimum p-values for each permutation
# Calculate all p and t values
t_rand_all = np.zeros((Nperm,N_big))
p_rand_all = np.zeros((Nperm,N_big))
pmin_pop_rand = np.zeros(Nperm)
pmin_unpop_rand = np.zeros(Nperm)
for i in range(Nperm):
for j, state in enumerate(states_big):
idx_use = range(len(states_big))
idx_use.pop(j)
t_rand_all[i,j], _ = sp.stats.ttest_ind(rand_statepop[i,j],np.hstack(rand_statepop[i,idx_use]))
_, p_rand_all[i,j] = sp.stats.mannwhitneyu(rand_statepop[i,j],np.hstack(rand_statepop[i,idx_use]))
# Identify the greatest significance of a state being more popular than the rest
pmin_pop_rand[i] = np.min(p_rand_all[i][np.where(t_rand_all[i]>0)[0]])
# Identify the greatest significance of a state being less popular than the rest
pmin_unpop_rand[i] = np.min(p_rand_all[i][np.where(t_rand_all[i]<0)[0]])
# -
# Test if most popular and least popular countries are outside of expectation
print 'Chance of a state being more distinctly popular than Canada: '
print sum(i < pmin_pop for i in pmin_pop_rand) / float(len(pmin_pop_rand))
print 'Chance of a state being less distinctly popular than US: '
print sum(i < pmin_unpop for i in pmin_unpop_rand) / float(len(pmin_unpop_rand))
| sfn/.ipynb_checkpoints/Poster viewer distribution by state-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ImageNet classification
# This notebook shows an example of ImageNet classification
# The network that is used for inference is a variant of DoReFaNet, whose topology is illustrated in the following picture.
# The pink layers are executed in the Programmable Logic at reduced precision (1 bit for weights, 2 bit for activations) while the other layers are executed in python.
#
# This notebook shows how to classify an image choosen by the user, while [dorefanet-imagenet-samples](./dorefanet-imagenet-samples.ipynb) runs the classification on labeled images (extracted from the dataset)
#
# 
# +
import os, pickle, random
from datetime import datetime
from matplotlib import pyplot as plt
from PIL import Image
# %matplotlib inline
import numpy as np
import cv2
import qnn
from qnn import Dorefanet
from qnn import utils
# -
# ## 1. Instantiate a Classifier
# Creating a classifier will automatically download the bitstream onto the device, allocate memory buffers and load the network hyperparameters and weights.
# The neural network to be implemented is specified in a json file (*dorefanet-layers.json* in this example)
# The weights for the non-offloaded layers are also loaded in a numpy dictionary to be used for execution in python.
# +
classifier = Dorefanet()
classifier.init_accelerator()
net = classifier.load_network(json_layer="/opt/python3.6/lib/python3.6/site-packages/qnn/params/dorefanet-layers.json")
conv0_weights = np.load('/opt/python3.6/lib/python3.6/site-packages/qnn/params/dorefanet-conv0.npy', encoding="latin1").item()
fc_weights = np.load('/opt/python3.6/lib/python3.6/site-packages/qnn/params/dorefanet-fc-normalized.npy', encoding='latin1').item()
# -
# ## 2. Get ImageNet Classes information
# Load labels and *synsets* of the 1000 [ImageNet](www.image-net.org/) classes into dictionaries
with open("imagenet-classes.pkl", 'rb') as f:
classes = pickle.load(f)
names = dict((k, classes[k][1].split(',')[0]) for k in classes.keys())
synsets = dict((classes[k][0], classes[k][1].split(',')[0]) for k in classes.keys())
# ## 3. Open image to be classified
# The image to be run through object classification is loaded automatically from the path set in the *img_folder* variable. The original image is shown before processing and will be automatically selected as the last one downloaded (both variables *img_folder* and *image_name* can be modified to the desired path/image to be classified).
img_folder = "/home/xilinx/jupyter_notebooks/qnn/images/"
img_file = os.path.join(img_folder, max(os.listdir(img_folder), key=lambda f: os.path.getctime(os.path.join(img_folder, f))))
img, img_class = classifier.load_image(img_file)
im = Image.open(img_file)
im
# ## 4. Execute the first convolutional layer in Python
# The first layer of this neural network has not been quantized, thus will not be executed in the HW accelerator (which supports only quantized arithmetic).
# Python provides, with numpy, a backend to execute convolution and other matrix operations. For user convenience the most popular operations (convolutional layer, thresholding, relu and fully connected layer) are provided in the class.
# +
conv0_W = conv0_weights['conv0/W']
conv0_T = conv0_weights['conv0/T']
start = datetime.now()
# 1st convolutional layer execution, having as input the image and the trained parameters (weights)
conv0 = utils.conv_layer(img, conv0_W, stride=4)
# The result in then quantized to 2 bits representation for the subsequent HW offload
conv0 = utils.threshold(conv0, conv0_T)
# Allocate accelerator output buffer
end = datetime.now()
micros = int((end - start).total_seconds() * 1000000)
print("First layer SW implementation took {} microseconds".format(micros))
print(micros, file=open('timestamp.txt', 'w'))
# -
# ## 5. HW Offload of the quantized layers
# The core layers, which values have been quantized during training, are executed in the Programmable Logic. The hardware accelerator consists of a dataflow implementation of multiple layers (in this case, convolution+maxpool).
# The host code parses the network topology (specified in the json file) and manages the sequence of execution on the accelerator.
# +
# Compute offloaded convolutional layers
out_dim = net['merge4']['output_dim']
out_ch = net['merge4']['output_channels']
conv_output = classifier.get_accel_buffer(out_ch, out_dim);
conv_input = classifier.prepare_buffer(conv0)
start = datetime.now()
classifier.inference(conv_input, conv_output)
end = datetime.now()
micros = int((end - start).total_seconds() * 1000000)
print("HW implementation took {} microseconds".format(micros))
print(micros, file=open('timestamp.txt', 'a'))
conv_output = classifier.postprocess_buffer(conv_output)
# -
# ## 6. Fully connected layers in python
# The fully connected layers, are executed in the python backend and the classification finalized
# +
# Normalize results
fc_input = conv_output / np.max(conv_output)
start = datetime.now()
# FC Layer 0
fc0_W = fc_weights['fc0/Wn']
fc0_b = fc_weights['fc0/bn']
fc0_out = utils.fully_connected(fc_input, fc0_W, fc0_b)
fc0_out = utils.qrelu(fc0_out)
fc0_out = utils.quantize(fc0_out, 2)
# FC Layer 1
fc1_W = fc_weights['fc1/Wn']
fc1_b = fc_weights['fc1/bn']
fc1_out = utils.fully_connected(fc0_out, fc1_W, fc1_b)
fc1_out = utils.qrelu(fc1_out)
# FC Layer 2
fct_W = fc_weights['fct/W']
fct_b = np.zeros((fct_W.shape[1], ))
fct_out = utils.fully_connected(fc1_out, fct_W, fct_b)
end = datetime.now()
micros = int((end - start).total_seconds() * 1000000)
print("Fully-connected layers took {} microseconds".format(micros))
print(micros, file=open('timestamp.txt', 'a'))
# -
# ## 7. Classification Results
# The top-5 results of the inference are provided with the corresponding human readable labels.
# The final classification scores are computed by a SoftMax Operator, that gives the normalized probabilities for all the classes.
# +
# Softmax
out = utils.softmax(fct_out)
# Top-5 results
topn = utils.get_topn_indexes(out, 5)
for k in topn: print("class:{0:>20}\tprobability:{1:>8.2%}".format(names[k].lower(), out[k]))
x_pos = np.arange(len(topn))
plt.barh(x_pos, out[topn], height=0.4, color='g', zorder=3)
plt.yticks(x_pos, [names[k] for k in topn])
plt.gca().invert_yaxis()
plt.xlim([0,1])
plt.grid(zorder=0)
plt.show()
# -
# ## 8. Performance evaluation
# This part show the performance of both software and hardware execution in terms of execution time, number of operations and number of operations over time.
# The software execution includes the first convolutional layer and the fully connected layers, while the hardware execution includes all the offloaded convolutional layers
# +
array = np.loadtxt('timestamp.txt')
array = list(map(lambda x: x/1000000, array))
MOPS = [238.176256, 1073.856969]
TIME = [array[0] + array[2], array[1]]
MOPSS = [m / t for (m, t) in zip(MOPS ,TIME)]
LABELS = ['SW', 'HW']
f, ((ax1, ax2, ax3)) = plt.subplots(1, 3, sharex='col', sharey='row', figsize=(15,2))
x_pos = np.arange(len(LABELS))
plt.yticks(x_pos, LABELS)
ax1.barh(x_pos, TIME, height=0.6, color='r', zorder=3)
ax1.invert_yaxis()
ax1.set_xlabel("Execution Time [s]")
ax1.set_ylabel("Platform")
ax1.grid(zorder=0)
ax2.barh(x_pos, MOPS, height=0.6, color='g', zorder=3)
ax2.invert_yaxis()
ax2.set_xlabel("# of Operations [MOPS]")
ax2.grid(zorder=0)
ax3.barh(x_pos, MOPSS, height=0.6, color='b', zorder=3)
ax3.invert_yaxis()
ax3.set_xlabel("Performances [MOPS/s]")
ax3.grid(zorder=0)
plt.show()
# -
# ## 9. SW execution of the quantized layers
# The core layers, which values have been quantized during training, are now executed in SW (by executing the HLS C++ source code).
# The host code parses the network topology (specified in the json file) and manages the sequence of execution on the accelerator.
# +
classifier_sw = Dorefanet("python_sw")
classifier_sw.init_accelerator()
conv_output_sw = classifier_sw.get_accel_buffer(out_ch, out_dim);
start = datetime.now()
classifier_sw.inference(conv_input, conv_output_sw)
end = datetime.now()
micros = int((end - start).total_seconds() * 1000000)
print("HW implementation took {} microseconds".format(micros))
# -
# ## Reset the device
classifier.deinit_accelerator()
# +
from pynq import Xlnk
xlnk = Xlnk();
xlnk.xlnk_reset()
| notebooks/dorefanet-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
from avgn.utils.paths import DATA_DIR, most_recent_subdirectory, ensure_dir
from sklearn.preprocessing import scale
#from pyclustertend import hopkins, vat, ivat
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
from avgn.clusterability.hopkins import hopkins_statistic
datasets = list(DATA_DIR.glob('embeddings/**/*.pickle'))
len(datasets)
datasets = [i for i in datasets if i.parent.stem not in [
'buckeye',
'BIRD_DB_Vireo_cassinii',
'swamp_sparrow',
'batsong_segmented'
]]
len(datasets)
def remove_outliers(z, pct = 99.5):
""" GPU based UMAP algorithm produces some outliers that UMAP does not, but is much faster
this is a quick fix for that.
"""
_min = np.percentile(z, (100-pct), axis=0)
_max = np.percentile(z, pct, axis=0)
for col in range(np.shape(z)[1]):
mask = z[:,col] < _min[col]
z[mask,col] = _min[col]
mask = z[:,col] > _max[col]
z[mask,col] = _max[col]
return z
ensure_dir(DATA_DIR / 'clusterability' / 'convex_sample_indvs')
def gen_clusterability_df(dataset):
"""
m_prop_n is the number of samples over X to perform hopkins statistic on (.1 is reccomended)
"""
save_loc = DATA_DIR / 'clusterability' / 'convex_sample_indvs' / \
(dataset.parent.stem + '_ ' + dataset.stem + '.pickle')
#if save_loc.exists():
# return
ds = pd.read_pickle(dataset)
specs = np.stack(ds[['spectrogram']].spectrogram.values)
specs = specs.reshape(len(specs), -1)
specs = scale(specs)
umap_proj = np.vstack(ds[['umap']].umap.values)
umap_proj = remove_outliers(umap_proj, pct=99.5)
umap_proj = scale(umap_proj)
nex = len(umap_proj)
print((dataset, np.shape(umap_proj)))
fig, ax = plt.subplots()
ax.scatter(umap_proj[:, 0], umap_proj[:, 1], s=1, color='k', alpha=0.1)
plt.show()
hopkins_dict = {
'umap':
{
0.01: hopkins_statistic(umap_proj, m_prop_n=0.01, n_neighbors=1, distribution="uniform_convex_hull"),
0.1: hopkins_statistic(umap_proj, m_prop_n=0.1, n_neighbors=1, distribution="uniform_convex_hull"),
},
}
dsname = dataset.parent.parent.stem if dataset.parent.stem == 'indvs' else dataset.parent.stem
clusterability_df = pd.DataFrame([[dataset, dsname, dataset.stem,
hopkins_dict['umap'][0.01], hopkins_dict['umap'][0.1], nex]], columns=[
'df', 'dataset', 'indv',
'umap_hopkins_1', 'umap_hopkins_10', 'nex'
])
clusterability_df.to_pickle(save_loc)
ensure_dir(DATA_DIR / 'clusterability' / 'convex_sample')
from joblib import Parallel, delayed
clust_data = Parallel(n_jobs=-1, verbose=10)(
delayed(gen_clusterability_df)(dataset)
for dataset in tqdm(datasets)
)
| notebooks/09.0-clusterability/.ipynb_checkpoints/hopkins-metric-on-umap-convex-hull-final-w-indvs-checkpoint.ipynb |