code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from IPython.display import display_html
def dss(*args):
html_str=''
for df in args:
html_str+=df.to_html()
display_html(html_str.replace('table','table style="display:inline"'),raw=True)
df = pd.read_pickle('datasets/sky-final.p')
price_df = df.sort_values(by=['price', 'mins', 'stops'])
dss(df, price_df)
t_idx = list(df.index.values)
p_idx = list(price_df.index.values)
t_idx_diff = [abs(t_idx.index(i) - p_idx.index(i)) for i in t_idx]
df['diff'] = t_idx_diff
df
final_df = df.sort_values(by=['diff','price', 'mins', 'stops'])
dss(df, final_df)
| flight_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import pandas as pd
import numpy as np
import PIL
seed = 16
np.random.seed(seed)
from keras.utils.np_utils import to_categorical
from keras.preprocessing.image import ImageDataGenerator
# -
# Purpose of this notebook is to build/test working model using a smaller subset of classes and data to minimize iteration time and to test CovNets of varying size before running on broader dataset.
# +
#check using system GPU for processing
from tensorflow.python.client import device_lib
import tensorflow as tf
os.environ["CUDA_VISIBLE_DEVICES"]="0" #for training on gpu
print(device_lib.list_local_devices())
# +
# copied over the train, validate and test sets for 5 randomly selected breeds
# -
# Given the subset, I copied over their respesctive train/validate/test image folders from the broader image data set. I maintained the full size of each train, val and test set.
os.chdir('C:\\Users\\Abhi\Documents\\Capstone\\datasets_subset1')
# +
train_datagen = ImageDataGenerator(rotation_range=15, shear_range=0.1, channel_shift_range=20,
width_shift_range=0.1, height_shift_range=0.1, zoom_range=0.1, horizontal_flip=True,
fill_mode='nearest', rescale=1./255)
validation_datagen = ImageDataGenerator(rescale=1./255)
test_datagen = ImageDataGenerator(rescale=1./255)
batch_size = 25
train_generator = train_datagen.flow_from_directory('subset_train', target_size=(224,224), color_mode='rgb',
class_mode='categorical', shuffle=False, batch_size=batch_size)
validation_generator = validation_datagen.flow_from_directory('subset_val', target_size=(224,224), color_mode='rgb',
class_mode='categorical', shuffle=False, batch_size=batch_size)
test_generator = test_datagen.flow_from_directory('subset_test', target_size=(224,224), color_mode='rgb',
class_mode='categorical', shuffle=False, batch_size=batch_size)
# reminder to self... flow_from_directory infers the class labels
# +
# importing keras modules and setting up a few parameters, instantiating early stopping
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import Flatten
from keras.constraints import maxnorm
from keras.optimizers import SGD
from keras.optimizers import Adam
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
import keras.utils
from keras import backend as K
K.set_image_dim_ordering('tf')
from keras.callbacks import EarlyStopping
early_stopping = EarlyStopping(monitor='val_loss', patience=2)
# -
import tensorflow as tf
tf_config = tf.ConfigProto()
tf_config.gpu_options.per_process_gpu_memory_fraction = 0.99
# tf_config.gpu_options.allow_growth = True **this causes python to crash, error: las.cc:444] failed to create cublas handle: CUBLAS_STATUS_ALLOC_FAILED
sess = tf.Session(config=tf_config)
# +
input_shape = (224,224, 3)
num_classes = 5
# will create a few different models.... initial base model
base_model = Sequential()
base_model.add(Conv2D(64, (11, 11), strides=4, input_shape=input_shape, padding='valid', activation='relu', kernel_constraint=maxnorm(3)))
base_model.add(MaxPooling2D(pool_size=(2, 2)))
base_model.add(Conv2D(64, (4, 4), strides=2, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
base_model.add(MaxPooling2D(pool_size=(2, 2)))
base_model.add(Conv2D(64, (4, 4), strides=2, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
base_model.add(Flatten())
base_model.add(Dense(256, activation='relu', kernel_constraint=maxnorm(3)))
base_model.add(Dropout(0.2))
base_model.add(Dense(num_classes, activation='softmax'))
# Compile model
epochs = 10
lrate = 0.003
decay = lrate/epochs
sgd = SGD(lr=lrate, momentum=0.9, decay=decay, nesterov=False)
base_model.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['accuracy'])
print(base_model.summary())
# +
# train base_model
base_model.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=100, epochs=epochs, callbacks=[early_stopping])
# +
# same model, more epochs (10 -> 50) and fewere steps per epoch, prior model params saw train and validate accuracies double, expecting the model to be within 80% acc
base_model.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=50, epochs=25, callbacks=[early_stopping])
# -
# Looks like we are increasing in accuracy with each successive epoch, perhaps need to train for more epochs. Let's test a deeper network with the same amount of epochs and see if we can begin with a better first epoch accuracy of 15%.
# +
# taking the base model and adding more hidden layers
deep_model = Sequential()
deep_model = Sequential()
deep_model.add(Conv2D(64, (11, 11), strides=4, input_shape=input_shape, padding='valid', activation='relu', kernel_constraint=maxnorm(3)))
deep_model.add(MaxPooling2D(pool_size=(2, 2)))
deep_model.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model.add(MaxPooling2D(pool_size=(2, 2)))
deep_model.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model.add(MaxPooling2D(pool_size=(2, 2)))
deep_model.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model.add(Flatten())
deep_model.add(Dense(288, activation='relu', kernel_constraint=maxnorm(3)))
deep_model.add(Dropout(0.2))
deep_model.add(Dense(num_classes, activation='softmax'))
# Compile model
epochs = 10
lrate = 0.003
decay = lrate/epochs
sgd = SGD(lr=lrate, momentum=0.9, decay=decay, nesterov=False)
deep_model.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['accuracy'])
print(deep_model.summary())
# +
# train deeper model
deep_model.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=100, epochs=epochs, callbacks=[early_stopping])
# +
# looks like the deeper model overfits the training data, but performs better on the validation data... let's train for more epochs
deep_model.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=100, epochs=50, callbacks=[early_stopping])
# +
# deeper model ran into early stopping on the validation set
# lets try base model with more epochs
base_model.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=100, epochs=25, callbacks=[early_stopping])
# +
# deep model with Adam optimizer
# taking the base model and adding more hidden layers
deep_model_Adam = Sequential()
deep_model_Adam = Sequential()
deep_model_Adam.add(Conv2D(64, (11, 11), strides=4, input_shape=input_shape, padding='valid', activation='relu', kernel_constraint=maxnorm(3)))
deep_model_Adam.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_Adam.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_Adam.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_Adam.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_Adam.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_Adam.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_Adam.add(Flatten())
deep_model_Adam.add(Dense(288, activation='relu', kernel_constraint=maxnorm(3)))
deep_model_Adam.add(Dropout(0.2))
deep_model_Adam.add(Dense(num_classes, activation='softmax'))
# Compile model
adam_op = Adam(lr=0.0001, beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.0, amsgrad=False)
deep_model_Adam.compile(loss='categorical_crossentropy', optimizer=adam_op, metrics=['accuracy'])
print(deep_model_Adam.summary())
# -
deep_model_Adam.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=100, epochs=25, callbacks=[early_stopping])
# +
# tweaked deep model w/ Adam optimizer. Deeper network topology near the input (less convolution than prior models), more FC nodes
deep_model_Adam_2 = Sequential()
deep_model_Adam_2 = Sequential()
deep_model_Adam_2.add(Conv2D(64, (8, 8), strides=2, input_shape=input_shape, padding='valid', activation='relu', kernel_constraint=maxnorm(3)))
deep_model_Adam_2.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_Adam_2.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_Adam_2.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_Adam_2.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_Adam_2.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_Adam_2.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_Adam_2.add(Flatten())
deep_model_Adam_2.add(Dense(256, activation='relu', kernel_constraint=maxnorm(3)))
deep_model_Adam_2.add(Dropout(0.2))
deep_model_Adam_2.add(Dense(num_classes, activation='softmax'))
# Compile model
adam_op = Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.0, amsgrad=False)
deep_model_Adam_2.compile(loss='categorical_crossentropy', optimizer=adam_op, metrics=['accuracy'])
print(deep_model_Adam_2.summary())
# -
deep_model_Adam_2.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=50, epochs=50, callbacks=[early_stopping])
# Deeper topology not necessarily better and is over-fitting.
# +
# tweaked deep model w/ RMSProp optimizer again with Deeper network topology near the input (less convolution than prior models), more FC nodes
deep_model_RMS = Sequential()
deep_model_RMS = Sequential()
deep_model_RMS.add(Conv2D(64, (8, 8), strides=2, input_shape=input_shape, padding='valid', activation='relu', kernel_constraint=maxnorm(3)))
deep_model_RMS.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_RMS.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_RMS.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_RMS.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_RMS.add(MaxPooling2D(pool_size=(2, 2)))
deep_model_RMS.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
deep_model_RMS.add(Flatten())
deep_model_RMS.add(Dense(256, activation='relu', kernel_constraint=maxnorm(3)))
deep_model_RMS.add(Dropout(0.2))
deep_model_RMS.add(Dense(num_classes, activation='softmax'))
# Compile model
deep_model_RMS.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
print(deep_model_RMS.summary())
# -
deep_model_RMS.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=100, epochs=50, callbacks=[early_stopping])
# +
# so more layers doesn't work.... let us keep the standard 3 CONV layers and widen the toplogy
wide_model = Sequential()
wide_model.add(Conv2D(32, (3, 3), strides=1, input_shape=input_shape, padding='valid', activation='relu', kernel_constraint=maxnorm(3)))
wide_model.add(MaxPooling2D(pool_size=(2, 2)))
wide_model.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
wide_model.add(MaxPooling2D(pool_size=(2, 2)))
wide_model.add(Conv2D(32, (3, 3), strides=2, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
wide_model.add(MaxPooling2D(pool_size=(2, 2)))
wide_model.add(Flatten())
wide_model.add(Dense(256, activation='relu', kernel_constraint=maxnorm(3)))
wide_model.add(Dropout(0.2))
wide_model.add(Dense(num_classes, activation='softmax'))
# Compile model
epochs = 10
wide_model.compile(loss='categorical_crossentropy', optimizer=adam_op, metrics=['accuracy'])
print(wide_model.summary())
# -
wide_model.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=50, epochs=50, callbacks=[early_stopping])
# +
# wider doesn't necessarily work... however, slowing the learning rate seems to having a positive impact. same model as above, decrease LR
wide_model_slow_learn = Sequential()
wide_model_slow_learn.add(Conv2D(32, (3, 3), strides=1, input_shape=input_shape, padding='valid', activation='relu', kernel_constraint=maxnorm(3)))
wide_model_slow_learn.add(MaxPooling2D(pool_size=(2, 2)))
wide_model_slow_learn.add(Conv2D(32, (3, 3), strides=1, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
wide_model_slow_learn.add(MaxPooling2D(pool_size=(2, 2)))
wide_model_slow_learn.add(Conv2D(32, (3, 3), strides=2, activation='relu', padding='valid', kernel_constraint=maxnorm(3)))
wide_model_slow_learn.add(MaxPooling2D(pool_size=(2, 2)))
wide_model_slow_learn.add(Flatten())
wide_model_slow_learn.add(Dense(256, activation='relu', kernel_constraint=maxnorm(3)))
wide_model_slow_learn.add(Dropout(0.2))
wide_model_slow_learn.add(Dense(num_classes, activation='softmax'))
# Compile model
adam_op = Adam(lr=0.00001, beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.0, amsgrad=False)
wide_model_slow_learn.compile(loss='categorical_crossentropy', optimizer=adam_op, metrics=['accuracy'])
print(wide_model_slow_learn.summary())
# -
wide_model_slow_learn.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=50, epochs=50, callbacks=[early_stopping])
# +
# it appears a slower learning rate might be key in allowing prior models to train for more epochs...
# let's try a few earlier models with a decreased learning rate
# -
adam_op = Adam(lr=0.00001, beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.0, amsgrad=False)
deep_model_Adam.compile(loss='categorical_crossentropy', optimizer=adam_op, metrics=['accuracy'])
print(deep_model_Adam.summary())
deep_model_Adam.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=50, epochs=50, callbacks=[early_stopping])
# +
# let's try tye base model w/ decreased learning rate and Adam optimizer (vs. SGD)
base_model.compile(loss='categorical_crossentropy', optimizer=adam_op, metrics=['accuracy'])
print(base_model.summary())
# -
base_model.fit_generator(train_generator, validation_data=validation_generator,
steps_per_epoch=50, epochs=50, callbacks=[early_stopping])
# So far the wider model (with less stride, smaller Convolution filter) w/ more trainable parameters and the simple base model appear to perform the best on validation data.
# +
# let's test on these iterations of the base, deep and wide models
base_scores = base_model.evaluate_generator(test_generator, steps=25)
print("Accuracy: %.2f%%" % (base_scores[1]*100))
# -
deep_model_Adam_scores = deep_model_Adam.evaluate_generator(test_generator, steps=25)
print("Accuracy: %.2f%%" % (deep_model_Adam_scores[1]*100))
wide_model_slow_learn_scores = wide_model_slow_learn.evaluate_generator(test_generator, steps=25)
print("Accuracy: %.2f%%" % (wide_model_slow_learn_scores[1]*100))
# Next steps... pick top 2 or 3 models and test and note which performs best. use top 2-3 on broader image data set (simple_CNN notebook)
# saving models and weights just in case... will need to retrain on broader image sets anyways
base_model.save('subset_base_model.h5')
deep_model_Adam.save('subset_deep_model_Adam.h5')
wide_model_slow_learn.save('subset_wide_model_slow_learn.h5')
| Jupiter Notebook Files/simple_CNN_smaller_class_subset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cugraph_dev
# language: python
# name: cugraph_dev
# ---
# # K-Cores
#
#
# In this notebook, we will use cuGraph to identify the K-Cores clusters in a test graph
#
# Notebook Credits
# * Original Authors: <NAME>
# * Created: 10/28/2019
# * Last Edit: 03/03/2020
#
# RAPIDS Versions: 0.13
#
# Test Hardware
# * GV100 32G, CUDA 10.2
#
#
#
# ## Introduction
#
# Compute the k-core of the graph G based on the out degree of its nodes. A k-core of a graph is a maximal subgraph that contains nodes of degree k or more. This call does not support a graph with self-loops and parallel edges.
#
# For a detailed description of the algorithm see: https://en.wikipedia.org/wiki/Degeneracy_(graph_theory)
#
#
# To compute the K-Core cluster in cuGraph use: <br>
# * __gc = cugraph.k_core(G, G, k=None, core_number=None)__
# * G: A cugraph.Graph object
# * k: optional, The _k_ value to use
# * core_number: optional, Precomputed core number of the nodes of the graph G. The see noptebook on Core-Number
#
# Returns:
# * __gc__
# * A new graph the contains the K-Core
#
# All vertices with the same partition ID are in the same cluster
#
#
# ## cuGraph Notice
# The current version of cuGraph has some limitations:
#
# * Vertex IDs need to be 32-bit integers.
# * Vertex IDs are expected to be contiguous integers starting from 0.
#
# cuGraph provides the renumber function to mitigate this problem. Input vertex IDs for the renumber function can be either 32-bit or 64-bit integers, can be non-contiguous, and can start from an arbitrary number. The renumber function maps the provided input vertex IDs to 32-bit contiguous integers starting from 0. cuGraph still requires the renumbered vertex IDs to be representable in 32-bit integers. These limitations are being addressed and will be fixed soon.
# ### Test Data
# We will be using the Zachary Karate club dataset
# *<NAME>, An information flow model for conflict and fission in small groups, Journal of
# Anthropological Research 33, 452-473 (1977).*
#
#
# 
#
# ### Prep
# Import needed libraries
import cugraph
import cudf
# ### Read data using cuDF
# Test file
datafile='../data//karate-data.csv'
# read the data using cuDF
gdf = cudf.read_csv(datafile, delimiter='\t', names=['src', 'dst'], dtype=['int32', 'int32'] )
# create a Graph
G = cugraph.Graph()
G.from_cudf_edgelist(gdf, source='src', destination='dst')
print("Main Graph")
print("\tNumber of Vertices: " + str(G.number_of_vertices()))
print("\tNumber of Edges: " + str(G.number_of_edges()))
# ### Now run K-Cores
# Call k-cores on the graph
kcg = cugraph.k_core(G)
print("K-Core Graph")
print("\tNumber of Vertices: " + str(kcg.number_of_vertices()))
print("\tNumber of Edges: " + str(kcg.number_of_edges()))
# ### Let's looks at the results
# The results show that the roughly 2/3s of the edges have been removed.
# Let's look at the degrees of the vertices
d = kcg.degrees()
d.sort_values(by='out_degree', ascending=False)
# It is clear that only vertices with more than out_degree of 4 are connected.
# We could remove all the disconnected vertices, or do a subgraph extraction with vertices where out_degree != 0
# 
# We can also just get a list of all the remaining edges as COO
coo = kcg.view_edge_list()
# print out edge list
coo
# ### Just for fun
# Let's try specifying a K value. Looking at the original network picture, it is easy to see that most vertices has at least degree two.
# If we specify k = 2 then only one vertex should be dropped
# Call k-cores on the graph
kcg2 = cugraph.k_core(G, k=2)
print("K-Core Graph")
print("\tNumber of Vertices: " + str(kcg2.number_of_vertices()))
print("\tNumber of Edges: " + str(kcg2.number_of_edges()))
d2 = kcg2.degrees()
d2.query('out_degree == 0')
# ___
# Copyright (c) 2019, NVIDIA CORPORATION.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# ___
| notebooks/cores/kcore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## part 1 ##
ex = '''.#.
..#
###'''.strip().split('\n')
def process_input(lines):
z = 0
active = set()
for y, line in enumerate(lines):
for x, c in enumerate(line):
if c == '#':
active.add((x, y, z))
return active
exactive = process_input(ex)
exactive
def count_active_neighbors(pos, active):
x, y, z = pos
count = 0
for i in (x-1, x, x+1):
for j in (y-1, y, y+1):
for k in (z-1, z, z+1):
if (i,j,k) != (x,y,z):
if (i,j,k) in active:
count += 1
return count
count_active_neighbors((1,2,0), exactive)
def cycle(active):
minx = min(r[0] for r in active)
maxx = max(r[0] for r in active)
miny = min(r[1] for r in active)
maxy = max(r[1] for r in active)
minz = min(r[2] for r in active)
maxz = max(r[2] for r in active)
newactive = set()
for x in range(minx - 1, maxx + 2):
for y in range(miny - 1, maxy + 2):
for z in range(minz - 1, maxz + 2):
c = count_active_neighbors((x, y, z), active)
if (x, y, z) in active:
if (c == 2) or (c == 3):
newactive.add((x, y, z))
else:
if c == 3:
newactive.add((x, y, z))
return newactive
cycle(exactive)
excyc = exactive.copy()
for i in range(6):
excyc = cycle(excyc)
len(excyc)
data = '''....#...
.#..###.
.#.#.###
.#....#.
...#.#.#
#.......
##....#.
.##..#.#'''.strip().split('\n')
data_active = process_input(data)
len(data_active)
datacyc = data_active.copy()
for i in range(6):
datacyc = cycle(datacyc)
len(datacyc)
# ## part 2 ##
def process_input2(lines):
z = 0
w = 0
active = set()
for y, line in enumerate(lines):
for x, c in enumerate(line):
if c == '#':
active.add((x, y, z, w))
return active
def count_active_neighbors2(pos, active):
x, y, z, w = pos
count = 0
for i in (x-1, x, x+1):
for j in (y-1, y, y+1):
for k in (z-1, z, z+1):
for l in (w-1, w, w+1):
if (i,j,k,l) != (x,y,z,w):
if (i,j,k,l) in active:
count += 1
return count
def cycle2(active):
minx = min(r[0] for r in active)
maxx = max(r[0] for r in active)
miny = min(r[1] for r in active)
maxy = max(r[1] for r in active)
minz = min(r[2] for r in active)
maxz = max(r[2] for r in active)
minw = min(r[3] for r in active)
maxw = max(r[3] for r in active)
newactive = set()
for x in range(minx - 1, maxx + 2):
for y in range(miny - 1, maxy + 2):
for z in range(minz - 1, maxz + 2):
for w in range(minw - 1, maxw + 2):
c = count_active_neighbors2((x, y, z, w), active)
if (x, y, z, w) in active:
if (c == 2) or (c == 3):
newactive.add((x, y, z, w))
else:
if c == 3:
newactive.add((x, y, z, w))
return newactive
def process(lines):
active = process_input2(lines)
for i in range(6):
active = cycle2(active)
return len(active)
process(ex)
process(data)
| day17.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Building interactive plots using `bqplot` and `ipywidgets`
# + [markdown] deletable=true editable=true
# * `bqplot` is built on top of the `ipywidgets` framework
# * `ipwidgets` and `bqplot` widgets can be seamlessly integrated to build interactive plots
# * `bqplot` figure widgets can be stacked with UI controls available in `ipywidgets` by using `Layout` classes (Box, HBox, VBox) in `ipywidgets`
# (Note that *only* `Figure` objects (not `Mark` objects) inherit from `DOMWidget` class and can be combined with other widgets from `ipywidgets`)
# * Trait attributes of widgets can be linked using callbacks. Callbacks should be registered using the `observe` method
#
# Please follow these links for detailed documentation on:
# 1. [Layout and Styling of Jupyter Widgets](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20Styling.html)
# * [Linking Widgets](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20Events.html)
#
# <br>Let's look at examples of linking plots with UI controls
# + deletable=true editable=true
import numpy as np
# + deletable=true editable=true
from ipywidgets import Button, VBox
import bqplot.pyplot as plt
# + [markdown] deletable=true editable=true
# Update the plot on a button click
# + deletable=true editable=true
y = np.random.randn(100).cumsum() # simple random walk
# create a button
update_btn = Button(description='Update', button_style='success')
# create a figure widget
fig1 = plt.figure(animation_duration=750)
line = plt.plot(y)
# define an on_click function
def on_btn_click():
# update the y attribute of line mark
line.y = np.random.randn(100).cumsum() # another random walk
# register the on_click function
update_btn.on_click(lambda btn: on_btn_click())
# stack button and figure using VBox
VBox([fig1, update_btn])
# + [markdown] deletable=true editable=true
# Let's look at an example where we link a plot to a dropdown menu
# + deletable=true editable=true
import pandas as pd
# create a dummy time series for 5 dummy stock tickers
dates = pd.date_range(start='20180101', end='20181231')
n = len(dates)
tickers = list('ABCDE')
prices = pd.DataFrame(np.random.randn(n, 5).cumsum(axis=0), columns=tickers)
# + deletable=true editable=true
from ipywidgets import Dropdown
# create a dropdown menu for tickers
dropdown = Dropdown(description='Ticker', options=tickers)
# create figure for plotting time series
current_ticker = dropdown.value
fig_title_tmpl = '"{}" Time Series' # string template for title of the figure
fig2 = plt.figure(title=fig_title_tmpl.format(current_ticker))
fig2.layout.width = '900px'
time_series = plt.plot(dates, prices[current_ticker])
plt.xlabel('Date')
plt.ylabel('Price')
# 1. create a callback which updates the plot when dropdown item is selected
def update_plot(*args):
selected_ticker = dropdown.value
# update the y attribute of the mark by selecting
# the column from the price data frame
time_series.y = prices[selected_ticker]
# update the title of the figure
fig2.title = fig_title_tmpl.format(selected_ticker)
# 2. register the callback by using the 'observe' method
dropdown.observe(update_plot, 'value')
# stack the dropdown and fig widgets using VBox
VBox([dropdown, fig2])
# + [markdown] deletable=true editable=true
# Let's now create a scatter plot where we select X and Y data from the two dropdown menus
# + deletable=true editable=true
from ipywidgets import Dropdown, HBox
# create two dropdown menus for X and Y attributes of scatter
x_dropdown = Dropdown(description='X', options=tickers, value='A')
y_dropdown = Dropdown(description='Y', options=tickers, value='B')
# create figure for plotting the scatter
x_ticker = x_dropdown.value
y_ticker = y_dropdown.value
# set up fig_margin to allow space to display color bar
fig_margin = dict(top=20, bottom=40, left=60, right=80)
fig3 = plt.figure(animation_duration=1000, fig_margin=fig_margin)
# custom axis options for color data
axes_options = {'color': {'tick_format': '%m/%y',
'side': 'right',
'num_ticks': 5}}
scatter = plt.scatter(x=prices[x_ticker],
y=prices[y_ticker],
color=dates, # represent chronology using color scale
stroke='black',
colors=['red'],
default_size=32,
axes_options=axes_options)
plt.xlabel(x_ticker)
plt.ylabel(y_ticker)
# 1. create a callback which updates the plot when dropdown item is selected
def update_scatter(*args):
x_ticker = x_dropdown.value
y_ticker = y_dropdown.value
# update the x and y attributes of the mark by selecting
# the column from the price data frame
with scatter.hold_sync():
scatter.x = prices[x_ticker]
scatter.y = prices[y_ticker]
# update the title of the figure
plt.xlabel(x_ticker)
plt.ylabel(y_ticker)
# 2. register the callback by using the 'observe' method
x_dropdown.observe(update_scatter, 'value')
y_dropdown.observe(update_scatter, 'value')
# stack the dropdown and fig widgets using VBox
VBox([HBox([x_dropdown, y_dropdown]), fig3])
| examples/Tutorials/Linking Plots With Widgets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import numpy as np
# __we usually start by creating a figure and axes__
fig = plt.figure()
ax = plt.axes()
# __we can then use `ax.plot`__
# +
fig = plt.figure()
ax = plt.axes()
x = np.linspace(0,10,1000)
ax.plot(x, np.sin(x));
ax.plot(x, np.cos(x))
# -
# __alternatively using the pylab interface__
plt.plot(x, np.sin(x)); # convinience like qplot in ggplot
# multiple plot using the pylab interface
plt.plot(x, np.sin(x))
plt.plot(x, np.cos(x));
# ### Adjusting the Plot: Line Colors and Styles
# e.g
plt.plt(x, np.sin(x - 0), color='blue') # color by name
plt.plot(x, np.sin(x - 1), color='g') # short color code (rgbcmyk)
plt.plot(x, np.sin(x - 2), color='0.75') # Grayscale between 0 and 1
plt.plot(x, np.sin(x - 3), color='#FFDD44') # Hex code (RRGGBB from 00 to FF)
plt.plot(x, np.sin(x - 4), color=(1.0,0.2,0.3)) # RGB tuple, values 0 and 1
plt.plot(x, np.sin(x - 5), color='chartreuse'); # all HTML color names supported
# line style keyword
plt.plot(x, x + 0, linestyle='solid')
plt.plot(x, x + 1, linestyle='dashed')
plt.plot(x, x + 2, linestyle='dashdot')
plt.plot(x, x + 3, linestyle='dotted');
# For short, you can use the following codes:
plt.plot(x, x + 4, linestyle='-') # solid
plt.plot(x, x + 5, linestyle='--') # dashed
plt.plot(x, x + 6, linestyle='-.') # dashdot
plt.plot(x, x + 7, linestyle=':'); # dotted
# or combining line and color
plt.plot(x, x + 0, '-g') # solid green
plt.plot(x, x + 1, '--c') # dashed cyan
plt.plot(x, x + 2, '-.k') # dashdot black
plt.plot(x, x + 3, ':r'); # dotted red
# +
# plt.plot?
# -
# #### Adjusting the Plot: Axes Limits
plt.plot(x, np.sin(x))
plt.xlim(-1, 11)
plt.ylim(-1.5, 1.5);
# __if reverse is needed for some reason (lol)__
# +
plt.plot(x, np.sin(x))
plt.xlim(10, 0)
plt.ylim(1.2, -1.2)
# -
# `plt.axis` (note this is with the 'i'), makes it even easire
# +
plt.plot(x, np.sin(x))
plt.axis([-1, 11, -1.5, 1.5]) # specify limit as list
# +
# automatically tighten the bounds
plt.plot(x, np.sin(x))
plt.axis('tight');
# -
# aspect ratio equal
plt.plot(x, np.sin(x))
plt.axis('equal');
# #### Labelling Plots
plt.plot(x, np.sin(x))
plt.title("A sine curve")
plt.xlabel("x")
plt.ylabel("sin(x)");
# __legend by specifying label in plot command__
# +
plt.plot(x, np.sin(x), '-g', label='sin(x)') # specifying label
plt.plot(x, np.cos(x), ':b', label='cos(x)')
plt.axis('equal')
plt.legend();
# -
# For transitioning between MATLAB-style functions and object-oriented methods,
# make the following changes:
# - plt.xlabel() → ax.set_xlabel()
# - plt.ylabel() → ax.set_ylabel()
# - plt.xlim() → ax.set_xlim()
# - plt.ylim() → ax.set_ylim()
# - plt.title() → ax.set_title()
# __using the OOP plotting however:__
# +
ax = plt.axes()
ax.plot(x, np.sin(x))
# it's easier to just use the set method instead of calling each func as in other
ax.set(xlim=(0,10), ylim=(-2,2),
xlabel='x', ylabel='sin(x)',
title='A Simple Plot');
| Chapter4/Simple Line Plots.ipynb |
# # Set and get hyperparameters in scikit-learn
#
# This notebook shows how one can get and set the value of hyperparameter in
# a scikit-learn estimator. We recall that hyperparameters refer to the
# parameter that will control the learning process.
#
# They should not be confused with the fitted parameters, resulting from the
# training. These fitted parameters are recognizable in scikit-learn because
# they are spelled with a final underscore `_`, for instance `model.coef_`.
#
# We will start by loading the adult census dataset and only use the numerical
# feature.
# +
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census.csv")
target_name = "class"
numerical_columns = [
"age", "capital-gain", "capital-loss", "hours-per-week"]
target = adult_census[target_name]
data = adult_census[numerical_columns]
# -
# <div class="admonition caution alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;">Caution!</p>
# <p class="last">Here and later, we use the name <tt class="docutils literal">data</tt> and <tt class="docutils literal">target</tt> to be explicit. In
# scikit-learn, documentation <tt class="docutils literal">data</tt> is commonly named <tt class="docutils literal">X</tt> and <tt class="docutils literal">target</tt> is
# commonly called <tt class="docutils literal">y</tt>.</p>
# </div>
# Our data is only numerical.
data.head()
# Let's create a simple predictive model made of a scaler followed by a
# logistic regression classifier.
#
# As mentioned in previous notebooks, many models, including linear ones,
# work better if all features have a similar scaling. For this purpose,
# we use a `StandardScaler`, which transforms the data by rescaling features.
# +
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
model = Pipeline(steps=[
("preprocessor", StandardScaler()),
("classifier", LogisticRegression())
])
# -
# We can evaluate the statistical performance of the model via
# cross-validation.
# +
from sklearn.model_selection import cross_validate
cv_results = cross_validate(model, data, target)
scores = cv_results["test_score"]
print(f"Accuracy score via cross-validation:\n"
f"{scores.mean():.3f} +/- {scores.std():.3f}")
# -
# We created a model with the default `C` value that is equal to 1. We saw in
# the previous exercise that we will be interested to set the value of an
# hyperparameter. One possibility is to set the parameter when we create the
# model instance. However, we might be interested to set the value of the
# parameter after the instance is created.
#
# Actually scikit-learn estimators have a `set_params` method that allows you
# to change the parameter of a model after it has been created. For example, we
# can set `C=1e-3` and fit and evaluate the model:
model.set_params(classifier__C=1e-3)
cv_results = cross_validate(model, data, target)
scores = cv_results["test_score"]
print(f"Accuracy score via cross-validation:\n"
f"{scores.mean():.3f} +/- {scores.std():.3f}")
# When the model of interest is a `Pipeline`, the parameter names are of the
# form `<model_name>__<parameter_name>` (note the double underscore in the
# middle). In our case, `classifier` comes from the `Pipeline` definition and
# `C` is the parameter name of `LogisticRegression`.
#
# In general, you can use the `get_params` method on scikit-learn models to
# list all the parameters with their values. For example, if you want to
# get all the parameter names, you can use:
for parameter in model.get_params():
print(parameter)
# `.get_params()` returns a `dict` whose keys are the parameter names and whose
# values are the parameter values. If you want to get the value of a single
# parameter, for example `classifier__C`, you can use:
model.get_params()['classifier__C']
# We can vary systematically the value of C to see if there is an optimal
# value.
for C in [1e-3, 1e-2, 1e-1, 1, 10]:
model.set_params(classifier__C=C)
cv_results = cross_validate(model, data, target)
scores = cv_results["test_score"]
print(f"Accuracy score via cross-validation with C={C}:\n"
f"{scores.mean():.3f} +/- {scores.std():.3f}")
# We can see that as long as C is high enough, the model seems to perform
# well.
#
# What we did here is very manual: it involves scanning the values for C
# and picking the best one manually. In the next lesson, we will see how
# to do this automatically.
#
# <div class="admonition warning alert alert-danger">
# <p class="first admonition-title" style="font-weight: bold;">Warning</p>
# <p class="last">When we evaluate a family of models on test data and pick the best
# performer, we can not trust the corresponding prediction accuracy, and
# we need to apply the selected model to new data. Indeed, the test data
# has been used to select the model, and it is thus no longer independent
# from this model.</p>
# </div>
# In this notebook we have seen:
#
# - how to use `get_params` and `set_params` to get the parameters of a model
# and set them.
| notebooks/parameter_tuning_manual.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
N = 2**6+1
x = np.linspace(0.995,1,N)
plt.plot(x, (1-x)**6)
import sympy
X = sympy.var('x')
sympy.expand((1-X)**6)
x = np.linspace(0.995,1,N)
plt.plot(x, x**6 - 6*x**5 + 15*x**4 - 20*x**3 + 15*x**2 - 6*x + 1)
0.01 * 1e2 #NOT what the computer does
# +
#The computer knows how to do the sum between number which have the same powers:
#This is what the computer does:
1e-16 = 0.1 * 1e1 + 0.0000000000000001 * 1e1 = (0.1 + 0.0000000000000001) * 1e1
#if the condition number of the summation is larger that 10^15 the summation is not allowed.
# -
A=np.random.random((50,50))
A.shape
K = np.linalg.cond(A)
A.shape, K
# +
#stiamo cercando di capire cosa significa avere un bad conditions number o un buon conditions
#number per semplici operazioni con la moltiplicazione
#cioe` vogliamo esplorare qual`e` il rounding error( l'errore dovuto al fatto che e` un numero finito)
#rounding error=10^15
#condition number:something that rappresent the inverse of signal to input ratio
# +
#repeating this few times the max error should reach k, the condition number of A
#np.linalg.cond(A)
estimated_k = 0
n_realizations = 1000
for j in range(n_realizations):
x = np.random.random((50,))
norm_x = np.linalg.norm(x)
dx = np.random.random((50,))
norm_dx = np.linalg.norm(dx)
y = A.dot(x)
norm_y = np.linalg.norm(y)
dy = np.linalg.norm(A.dot(dx))
norm_dy = np.linalg.norm(dy)
kx = (norm_dy/norm_y) / (norm_dx/norm_x) #normalize perturb in output divided normalize perturb in input
estimated_k = max(kx, estimated_k)
estimated_k
# -
kx, norm_x, norm_y, norm_dx, norm_dy
# Now we try to estimate truncation error
def FD(f, x, dx): #valuta la funzione f nel punto x usando dx come approx step
return (f(x+dx)-f(x))/dx
N = 2**6+1
x = np.linspace(0,1,N)
# +
def myf(x):
return np.sin(np.pi * xi)
def supnorm(f_of_x):
return max(abs(f_of_x))
y = myf(x)
yprime = np.pi * np.cos(np.pi * x)
FDyprime = FD(myf, x, 2**(-3))
# +
#plot(x,yprime 'ro')
dxs = []
errors = []
for i in range(2,51): #{2, 3, 4, 5}
dx = 2**(-i)
dxs.append(dx)
FDprime = FD(myf, x, dx)
# plot(x,FDprime)
error = supnorm(yprime-FDprime)
errors.append(error)
loglog(dxs, errors)
#plot(x,FD(myf,x,2**(-i)))
#plot(x,FDyprime)
yprime.shape
# -
| notes/Notes_lesson3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
feature_data_file = "/home/jeffrey/repos/VSB_Power_Line_Fault_Detection/extracted_features/"+"train_features_thresh_5.0_db4.csv"
feature_data_file = "/home/jeffrey/repos/VSB_Power_Line_Fault_Detection/extracted_features/"+"train_features_noCancel_thresh_4.0_db4.csv"
df = pd.read_csv(feature_data_file)
df.head()
df.describe()
df["fault"].value_counts()/len(df) # imbalanced data set
# +
from sklearn.preprocessing import normalize
#feature_matrix_columns = ["entropy", "median", "mean", "std", "var", "rms", "no_zero_crossings", "no_mean_crossings", "fault"]
#feature_matrix_columns = ["median", "mean", "std", "var", "rms", "no_zero_crossings", "no_mean_crossings", "fault"]
#feature_matrix_columns = ["entropy", "fault"]
#features = df[["entropy", "n5", "n25", "n75", "n95", "median", "mean", "std", "var", "rms", "no_zero_crossings", "no_mean_crossings", "min_height", "max_height", "mean_height", "min_width", "max_width", "mean_width", "num_detect_peak", "num_true_peaks"]]
features = df[["entropy", "mean", "std", "var", "rms", "no_zero_crossings", "no_mean_crossings", "min_height", "max_height", "mean_height", "min_width", "max_width", "mean_width", "num_detect_peak", "num_true_peaks", "fault"]]
features = df[["entropy", "std", "max_width", "mean_width", "num_true_peaks", "fault"]]
#df2 = df[feature_matrix_columns]
#df_n = normalize(df2, axis=0)
features.head()
# +
sns.set_style("whitegrid")
#g = sns.PairGrid(df2) # Create a Pair Grid Instance
#g.map_upper(sns.regplot, color="#51ACC5")
#g.map_diag(sns.kdeplot, legend=False, shade=True)
#g.map_lower(sns.kdeplot, shade=False, cmap="Blues")
sns.pairplot(features, diag_kind="kde", kind="reg", hue="fault", palette="RdBu_r", height=4)
# -
feature_data_file = "/home/jeffrey/repos/VSB_Power_Line_Fault_Detection/"+"train_features_coif2.csv"
df = pd.read_csv(feature_data_file)
df.describe()
# +
feature_matrix_columns = ["entropy", "median", "mean", "std", "var", "rms", "no_zero_crossings", "no_mean_crossings", "fault"]
#feature_matrix_columns = ["median", "mean", "std", "var", "rms", "no_zero_crossings", "no_mean_crossings", "fault"]
#feature_matrix_columns = ["entropy", "fault"]
df2 = df[feature_matrix_columns]
sns.set_style("whitegrid")
#g = sns.PairGrid(df2) # Create a Pair Grid Instance
#g.map_upper(sns.regplot, color="#51ACC5")
#g.map_diag(sns.kdeplot, legend=False, shade=True)
#g.map_lower(sns.kdeplot, shade=False, cmap="Blues")
sns.pairplot(df2, diag_kind="kde", kind="reg", hue="fault", palette="husl", height=3)
# -
| research/VSB_2_Extracted_Feature_Data_Exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "MNIST Digit Classifier using PyTorch"
# > "A simple workflow on how to build a multilayer perceptron to classify MNIST handwritten digits using PyTorch. We define a custom Dataset class to load and preprocess the input data. The neural network architecture is built using a sequential layer, just like the Keras framework. We define the training and testing loop manually using Python for-loop. The final model is evaluated using a standard accuracy measure."
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>, <NAME>, <NAME>
# - categories: [python, classification, computer-vision, pytorch]
# - image: images/mnist-digit-pytorch.jpeg
# # Background
# + [markdown] cell_id="00000-d74ccc7c-e4f5-41ea-b2d0-fdb09ff53bd5" deepnote_cell_type="markdown" tags=[]
# Deep learning is part of a broader family of machine learning methods based on artificial neural networks with representation learning. These neural networks attempt to simulate the behavior of the human brain allowing it to “learn” from large amounts of data. Not only possible to apply in large amounts of data, but it also allows to deal with unstructured data such as image, text, and sound.
#
# You can try to implement a neural network from scratch. But, do you think this is a good idea when building deep learning models on a real-world dataset? It is definitely possible if you have days or weeks to spare waiting for the model to build. But, in any conditions we have many constrain to do it e.g time and cost.
#
# Here is the good news, now we can use deep learning frameworks that aim to simplify the implementation of complex deep learning models. Using these frameworks, we can implement complex models like convolutional neural networks in no time.
#
# A deep learning framework is a tool that allows us to build deep learning models more easily and quickly. They provide a clear and concise way for defining models using a collection of pre-built and optimized components. Instead of writing hundreds of lines of code, we can use a suitable framework to help us to build such a model quickly.
# + [markdown] cell_id="00002-f5f5f37e-a558-47d5-8fdd-8f88e7099889" deepnote_cell_type="markdown" tags=[]
# 
#
# |Keras | PyTorch|
# |---|---|
# |Keras was released in March 2015| While PyTorch was released in October 2016|
# |Keras has a high level API| While PyTorch has a low level API|
# |Keras is comparatively slower in speed | While PyTorch has a higher speed than Keras, suitable for high performance|
# |Keras has a simple architecture, making it more readable and easy to use | While PyTorch has very low readablility due to a complex architecture|
# |Keras has a smaller community support | While PyTorch has a stronger community support|
# |Keras is mostly used for small datasets due to its slow speed | While PyTorch is preferred for large datasets and high performance|
# |Debugging in Keras is difficult due to presence of computational junk | While debugging in PyTorch is easier and faster|
# |Keras provides static computation graphs | While PyTorch provides dynamic computation graphs|
# |Backend for Keras include:TensorFlow, Theano and Microsoft CNTK backend | While PyTorch has no backend implementation|
# -
# # Installation
# + [markdown] cell_id="00001-021932f4-ebaa-4c6c-ae2a-e05f27bdc395" deepnote_cell_type="markdown" deepnote_to_be_reexecuted=false execution_millis=9 execution_start=1629441374960 source_hash="28e118d0" tags=[]
# Please ensure you have installed the following packages to run sample notebook. Here is the short instruction on how to create a new conda environment with those package inside it.
#
# 1. Open the terminal or Anaconda command prompt.
#
# 
#
# 2. Create new conda environment by running the following command.
#
# `conda create -n <env_name> python=3.7`
#
# 3. Activate the conda environment by running the following command.
#
# `conda activate <env_name>`
#
# 4. Install additional packages such as `pandas`, `numpy`, `matplotlib`, `seaborn`, and `sklearn` inside the conda environment.
#
# `pip install pandas numpy matplotlib seaborn scikit-learn`
#
# 5. Install `torch` into the environment.
#
# `pip3 install torch torchvision torchaudio`
# -
# > Note: For more details, please visit [PyTorch setup](https://pytorch.org/get-started/locally/)
# + [markdown] cell_id="00004-12df2948-40d4-4cf2-b425-ce3dfaa00a4c" deepnote_cell_type="text-cell-h1" is_collapsed=false tags=[]
# # Libraries
# -
# After the required package is installed, load the package into your workspace using the `import`
# + cell_id="00001-32c246fa-8cb4-4171-b469-02ae701dee8a" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=15 execution_start=1629447748748 source_hash="a50c2f92" tags=[]
import pandas as pd # for read_csv
import numpy as np # for np.Inf
import matplotlib.pyplot as plt # visualization
import seaborn as sns # heatmap visualization
from sklearn.metrics import confusion_matrix, classification_report # evaluation metrics
import pickle # serialization
import torch
from torch.utils.data import Dataset, DataLoader, random_split
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
plt.style.use("seaborn")
torch.manual_seed(123)
# -
# # Workflow
#
# 
# + [markdown] cell_id="00005-d05d942a-df5a-4572-8094-287371bcc5ce" deepnote_cell_type="text-cell-h1" is_collapsed=false tags=[]
# # Load Data
# + [markdown] cell_id="00007-9e0d1160-7384-4207-97b6-ee9cbb07cb07" deepnote_cell_type="markdown" tags=[]
# ## Working with Images
# + [markdown] cell_id="00008-aba71842-888f-4bf5-830a-466cb08e910e" deepnote_cell_type="markdown" tags=[]
# Let's start implement our existing knowledge of neural network using `torch` in Python to solve an **image classification** problem. We'll use famous [**MNIST Handwritten Digits Data**](https://www.kaggle.com/c/digit-recognizer/data) as our training dataset. It consists of 28 by 28 pixels grayscale images of handwritten digits (0 to 9) and labels for each image indicating which digit it represents. There are two files of data to be downloaded: `train.csv` is pixel data with actual digit label, whereas `test.csv` is pixel data **without** the actual digit label. Here are some sample image from dataset:
#
# 
#
# It's evident that these images are relatively small in size, and recognizing the digits can sometimes be challenging even for the human eye. While it's useful to look at these images, there's just one problem here: PyTorch doesn't know how to work with images. We need to convert the images into tensors. We can do this by specifying a transform while creating our dataset.
#
# PyTorch `Dataset` allow us to specify one or more transformation functions that are applied to the images as they are loaded. We'll use the **`torch.tensor()`** to convert the pixel values into tensors.
# + cell_id="00003-96a15415-66f5-446a-8b29-74254031c1b5" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1629447748750 source_hash="442593e2" tags=[]
class MNISTDataset(Dataset):
# constructor
def __init__(self, file_path):
# read data
df = pd.read_csv(file_path)
target_col = "label"
self.label_exist = target_col in df.columns
if self.label_exist:
# split feature-target
X = df.drop(columns=target_col).values
y = df[target_col].values
# convert numpy array to tensor
self.y = torch.tensor(y)
else:
X = df.values
self.X = torch.tensor(X, dtype=torch.float32)
# scaling
self.X /= 255
# for iteration
def __getitem__(self, idx):
if self.label_exist:
return self.X[idx], self.y[idx]
else:
return self.X[idx]
# to check num of observations
def __len__(self):
return self.X.size()[0]
# + cell_id="00004-20a1be2d-cf77-4a39-976e-16655e40c6fc" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2509 execution_start=1629447748756 source_hash="9f99c286" tags=[]
# load dataset
mnist_data = MNISTDataset("data_input/mnist/train.csv")
print(mnist_data.X.size())
# + cell_id="00006-dc7a1cd4-c597-49d7-9dd0-48b4fe70ba7c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1629447751240 source_hash="df5b0b2c" tags=[]
# global variables
SPLIT_PROP = 0.8
BATCH_SIZE = 16
# + [markdown] cell_id="00013-45ac3714-1ee6-4915-916a-fdac4a9c45d6" deepnote_cell_type="markdown" tags=[]
# ## Training and Validation Datasets
# + [markdown] cell_id="00014-43a3824b-64ae-45bb-bb42-cf955786bc06" deepnote_cell_type="markdown" tags=[]
# While building real-world machine learning models, it is quite common to split the dataset into three parts:
#
# - **Training set** - used to train the model, i.e., compute the loss and adjust the model's weights using gradient descent.
# - **Validation set** - used to evaluate the model during training, adjust hyperparameters (learning rate, etc.), and pick the best version of the model.
# - **Test set** - used to compare different models or approaches and report the model's final accuracy.
#
# + [markdown] cell_id="00015-45d93ae1-2690-419f-be1a-8a79e806e548" deepnote_cell_type="markdown" tags=[]
# In the MNIST dataset, there are 42,000 images in `train.csv` and 784 images in `test.csv`. The test set is standardized so that different researchers can report their models' results against the same collection of images.
#
# Since there's no predefined validation set, we must manually split the 42,000 images into training, validation, and test datasets. Let's set aside 42,000 randomly chosen images for validation. We can do this using the **`random_split`** method from `torch`.
# + cell_id="00006-d6330cb0-5ab3-4c10-b622-b5fb57db857c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=45 execution_start=1629447751246 source_hash="f5632c55" tags=[]
train_val_size = int(SPLIT_PROP * len(mnist_data))
test_size = len(mnist_data) - train_val_size
train_size = int(SPLIT_PROP * train_val_size)
val_size = train_val_size - train_size
# train-val-test split
train_mnist, val_mnist, test_mnist = random_split(mnist_data, [train_size, val_size, test_size])
print(f"Training set: {len(train_mnist):,} images")
print(f"Validation set: {len(val_mnist):,} images")
print(f"Testing set: {len(test_mnist):,} images")
# + [markdown] cell_id="00017-4c738033-4ea3-492c-ad38-c7d782a12dfd" deepnote_cell_type="markdown" tags=[]
# It's essential to choose a random sample for creating a validation and test set. Training data is often sorted by the target labels, i.e., images of 0s, followed by 1s, followed by 2s, etc. If we create a validation set using the last 20% of images, it would only consist of 8s and 9s. In contrast, the training set would contain no 8s or 9s. Such a training-validation would make it impossible to train a useful model.
#
# We can now create `DataLoader` to help us load the data in batches. We'll use a batch size of 16. We set **`shuffle=True`** for the data loader to ensure that the batches generated in each epoch are different. This randomization helps generalize & speed up the training process.
# + cell_id="00005-434da867-d604-4523-a19c-928a7ed4b2bf" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=21 execution_start=1629447751275 source_hash="e2912be0" tags=[]
train_loader = DataLoader(train_mnist, batch_size=BATCH_SIZE, shuffle=True)
val_loader = DataLoader(val_mnist, batch_size=BATCH_SIZE, shuffle=True)
test_loader = DataLoader(test_mnist, batch_size=BATCH_SIZE, shuffle=True)
print(f"Training data loader: {len(train_loader):,} images per batch")
print(f"Validation data loader: {len(val_loader):,} images per batch")
print(f"Testing data loader: {len(test_loader):,} images per batch")
# + [markdown] cell_id="00010-b0290a59-c716-4427-8727-7b6693243814" deepnote_cell_type="markdown" tags=[]
# # Visualize Data
# + [markdown] cell_id="00021-43f16f43-fc6c-45f6-b978-d93428db0fcd" deepnote_cell_type="markdown" tags=[]
# Let's check the proportion of each class labels, since it is important to ensure that the model can learn each digit in a balanced and fair manner. We check the distribution of digits in training, validation, and also test set.
# + cell_id="00011-7ecb710a-d4cb-4f06-b92a-524cc5e8b79c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=589 execution_start=1629447751285 source_hash="6f8a9d2c" tags=[]
# collapse-hide
# get target variable proportion
train_prop = mnist_data.y[train_mnist.indices].bincount()
val_prop = mnist_data.y[val_mnist.indices].bincount()
test_prop = mnist_data.y[test_mnist.indices].bincount()
# visualization
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
for ax, prop, title, col in zip(axes, [train_prop, val_prop, test_prop], ['Train', 'Validation', 'Test'], 'bgr'):
class_target = range(0, 10)
ax.bar(class_target, prop, color=col)
ax.set_title(title)
ax.set_xticks(class_target)
ax.set_xlabel("Label")
axes[0].set_ylabel("Frequency")
plt.show()
# + [markdown] cell_id="00023-a854c3f6-3933-4f1b-a640-69f7a88b1d27" deepnote_cell_type="markdown" tags=[]
# The distribution of our class labels do seem to be spread out quite evenly, so there's no problem. Next, we visualize a couple of images from `train_loader`:
# + cell_id="00011-214f5f27-9340-4ea1-beff-ea4dc9a8698a" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1030 execution_start=1629447751863 source_hash="b7de16a2" tags=[]
# collapse-hide
class_label = ['zero', 'one', 'two', 'three',
'four', 'five', 'six', 'seven', 'eight', 'nine']
images, labels = next(iter(train_loader))
images = images.reshape((train_loader.batch_size, 28, 28))
fig, axes = plt.subplots(2, 8, figsize=(12, 4))
for ax, img, label in zip(axes.flat, images, labels):
ax.imshow(img, cmap="gray")
ax.axis("off")
ax.set_title(class_label[label])
plt.tight_layout()
# + [markdown] cell_id="00011-54b7f1a8-6eac-4c81-a3b4-0e783bd3d70f" deepnote_cell_type="markdown" tags=[]
# # Define Model Architecture
# + [markdown] cell_id="00030-1f14e51f-7d45-47f8-9c6a-a603150e9564" deepnote_cell_type="markdown" tags=[]
# We'll create a neural network with three layers: two hidden and one output layer. Additionally, we'll use ReLu activation function between each layer. Let's create a **`nn.Sequential`** object, which consists of linear fully-connected layer:
#
# - The input size is 784 nodes as the MNIST dataset consists of 28 by 28 pixels
# - The hidden sizes are 128 and 64 respectively, this number can be increased or decreased to change the learning capacity of the model
# - The output size is 10 as the MNIST dataset has 10 target classes (0 to 9)
# + cell_id="00015-dc239169-482e-4994-9380-5d00f58e8fa5" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=25 execution_start=1629447752898 source_hash="39ed5061" tags=[]
# Sequential, Keras style
input_size = 784
hidden_sizes = [128, 64]
output_size = 10
model = nn.Sequential(
nn.Linear(input_size, hidden_sizes[0]),
nn.ReLU(),
nn.Linear(hidden_sizes[0], hidden_sizes[1]),
nn.ReLU(),
nn.Linear(hidden_sizes[1], output_size) # logit score
)
model
# + [markdown] cell_id="00032-a93c3d43-97a3-49e5-aaf4-a795f2b842ed" deepnote_cell_type="markdown" tags=[]
# A little bit about the mathematical detail: The image vector of size 784 are transformed into intermediate output vector of length 128 then 64 by performing a matrix multiplication of inputs matrix. Thus, input and layer1 outputs have linear relationship, i.e., each element of layer outputs is a weighted sum of elements from inputs. Thus, even as we train the model and modify the weights, layer1 can only capture linear relationships between inputs and outputs.
# + [markdown] cell_id="00035-1a830b36-8ac0-4963-bd21-4ff5558eded7" deepnote_cell_type="markdown" tags=[]
# Activation function such as **Rectified Linear Unit (ReLU)** is used to introduce non-linearity to the model. It has the formula `relu(x) = max(0,x)` i.e. it simply replaces negative values in a given tensor with the value 0. We refer to ReLU as the activation function, because for each input certain outputs are activated (those with non-zero values) while others turned off (those with zero values). ReLU can be seen visually as follows:
#
# 
# + [markdown] cell_id="00038-35f8eb7b-4a25-47e2-a662-5199064a3327" deepnote_cell_type="markdown" tags=[]
# The output layer returns a batch of vectors of size 10. This predicted output is then being compared with actual label, quantified by using **`nn.CrossEntropyLoss()`**. It combines `LogSoftmax` and `NLLLoss` (Negative Log Likelihood Loss) in one single class.
#
# The loss value is used to update the parameter weights in the model. The parameter update algorithm can be implemented via an optimizer. In this case, we are using **`torch.optim.Adam()`** with learning rate (`lr=0.001`).
# + cell_id="00015-7fa1979b-f1f6-4127-9181-9339f1fe2647" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=476138288 execution_start=1629447752928 source_hash="c29ccb40" tags=[]
# specify loss function (cross-entropy)
criterion = nn.CrossEntropyLoss()
# specify optimizer (Adam) and learning rate = 0.001
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
# + [markdown] cell_id="00016-7486aca8-e1ab-485e-9643-888b4352c1a1" deepnote_cell_type="markdown" tags=[]
# # Train the Model
# + [markdown] cell_id="00039-f69be028-7bb3-429f-af30-c391b18e26c7" deepnote_cell_type="markdown" tags=[]
# Using `torch`, we have to manually loop the data per epoch to **train the model**. Here are the training loop for one epoch:
#
# 1. Clear gradients of all optimized variables from `optimizer`
# 2. Forward pass: compute predicted output by passing input data to the `model`
# 3. Calculate the loss based on specified `criterion`
# 4. Backward pass: compute gradient of the loss with respect to `model` parameters
# 5. Perform parameter update using `optimizer` algorithm
# 6. Accumulate training loss and accuracy
#
# Inside the training loop, we also **validate the model** by performing the following steps:
#
# 1. Forward pass: compute predicted output by passing input data to the `model`
# 2. Calculate the loss based on specified `criterion`
# 3. Accumulate validation loss and accuracy
#
# Special notes:
#
# - If you use `Dropout` or `BatchNorm` layer, don't forget to use `model.train()` when training the model and `model.eval()` when validating/testing the model so that the layer behaves accordingly.
# - Don't forget to set `torch.no_grad()` before validating the model to disable the gradient calculation since we are not updating the parameters.
# + [markdown] cell_id="00040-c51e2690-f67c-4be0-a1d0-1f6100238c15" deepnote_cell_type="markdown" tags=[]
# But first let's us define `evaluate_accuracy` to calculate accuracy given the predicted `logits` and `y_true` actual label.
# + cell_id="00019-d783395d-8dbe-4f98-b7ed-bf5498c390a3" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1629447752934 source_hash="e2088c9c" tags=[]
def evaluate_accuracy(logits, y_true):
# get index with the largest logit value PER OBSERVATION
_, y_pred = torch.max(logits, dim=1)
# calculate proportion of correct prediction
correct_pred = (y_pred == y_true).float()
acc = correct_pred.sum() / len(correct_pred)
return acc * 100
# + [markdown] cell_id="00042-197fb939-d890-45ec-aec4-7e42f9ffc10a" deepnote_cell_type="markdown" tags=[]
# Define training loop with the following parameters:
#
# - `model`: untrained model
# - `train_loader`: data train of a `DataLoader` object
# - `val_loader`: data validation of a `DataLoader` object
# - `criterion`: loss function to be optimized
# - `optimizer`: optimization algorithm to used on the `model` parameters
# - `n_epochs`: number of training epochs
# - `model_file_name`: file name of serialized model. During the training loop, model with the lowest validation loss will be saved as a serialized model with `.pt` extension.
# + cell_id="00020-2a5077d4-cd26-456a-a6ed-71d260ba1c41" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1628971614659 source_hash="f63f9075" tags=[]
def train(model, train_loader, val_loader, criterion, optimizer, n_epochs, model_file_name='model.pt'):
# initialize container variable for model performance results per epoch
history = {
'n_epochs': n_epochs,
'loss': {
'train': [],
'val': []
},
'acc': {
'train': [],
'val': []
}
}
# initialize tracker for minimum validation loss
val_loss_min = np.Inf
# loop per epoch
for epoch in range(n_epochs):
# initialize tracker for training performance
train_acc = 0
train_loss = 0
###################
# train the model #
###################
# prepare model for training
model.train()
# loop for each batch
for data, target in train_loader:
# STEP 1: clear gradients
optimizer.zero_grad()
# STEP 2: forward pass
output = model(data)
# STEP 3: calculate the loss
loss = criterion(output, target)
# STEP 4: backward pass
loss.backward()
# STEP 5: perform parameter update
optimizer.step()
# STEP 6: accumulate training loss and accuracy
train_loss += loss.item() * data.size(0)
acc = evaluate_accuracy(output, target)
train_acc += acc.item() * data.size(0)
######################
# validate the model #
######################
# disable gradient calculation
with torch.no_grad():
# initialize tracker for validation performance
val_acc = 0
val_loss = 0
# prepare model for evaluation
model.eval()
# loop for each batch
for data, target in val_loader:
# STEP 1: forward pass
output = model(data)
# STEP 2: calculate the loss
loss = criterion(output, target)
# STEP 3: accumulate validation loss and accuracy
val_loss += loss.item() * data.size(0)
acc = evaluate_accuracy(output, target)
val_acc += acc.item() * data.size(0)
####################
# model evaluation #
####################
# calculate average loss over an epoch
train_loss /= len(train_loader.sampler)
val_loss /= len(val_loader.sampler)
history['loss']['train'].append(train_loss)
history['loss']['val'].append(val_loss)
# calculate average accuracy over an epoch
train_acc /= len(train_loader.sampler)
val_acc /= len(val_loader.sampler)
history['acc']['train'].append(train_acc)
history['acc']['val'].append(val_acc)
# print training progress per epoch
print(f'Epoch {epoch+1:03} | Train Loss: {train_loss:.5f} | Val Loss: {val_loss:.5f} | Train Acc: {train_acc:.2f} | Val Acc: {val_acc:.2f}')
# save model if validation loss has decreased
if val_loss <= val_loss_min:
print(
f'Validation loss decreased ({val_loss_min:.5f} --> {val_loss:.5f}) Saving model to {model_file_name}...')
torch.save(model.state_dict(), model_file_name)
val_loss_min = val_loss
print()
# return model performance history
return history
# + cell_id="00020-11beb9c3-5bb3-47ee-be0e-e0943986fa72" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=93453 execution_start=1628971614705 source_hash="b6765ecd" tags=[]
history = train(
model, train_loader, val_loader, criterion, optimizer, n_epochs=20,
model_file_name='cache/model-mnist-digit.pt'
)
# + cell_id="00040-777bb825-1eee-4e1c-856e-c8c5e09d6680" deepnote_cell_type="code" tags=[]
# save history dictionary as pickle file
with open('cache/history-mnist-digit.pickle', 'wb') as f:
pickle.dump(history, f, protocol=pickle.HIGHEST_PROTOCOL)
# + [markdown] cell_id="00045-37ae4e07-9651-4fa8-a810-7350a9a437a7" deepnote_cell_type="markdown" tags=[]
# Visualize the loss and accuracy from `history` to see the model performance for each epoch.
# + cell_id="00023-5eadefac-e0c8-4e29-b7ea-9473e52bd06c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=498 execution_start=1629447758154 source_hash="19a34daf" tags=[]
# collapse-hide
# load previously saved history dictionary
with open('cache/history-mnist-digit.pickle', 'rb') as f:
history = pickle.load(f)
# visualization
epoch_list = range(1, history['n_epochs']+1)
fig, axes = plt.subplots(1, 2, figsize=(18, 6))
for ax, metric in zip(axes, ['loss', 'acc']):
ax.plot(epoch_list, history[metric]['train'], label=f"train_{metric}")
ax.plot(epoch_list, history[metric]['val'], label=f"val_{metric}")
ax.set_xlabel('epoch')
ax.set_ylabel(metric)
ax.legend()
# + [markdown] cell_id="00047-7c052b65-b0cd-4c4f-a6d3-ea949bfb825b" deepnote_cell_type="markdown" tags=[]
# From the visualization, we can deduct that the model is a good enough since the difference of model performance on training and validation is not too much different, and the accuracy is converging at 96-97%.
# + [markdown] cell_id="00024-b6e63891-c106-4408-a876-22634dbb785e" deepnote_cell_type="markdown" tags=[]
# # Test the Model
# + [markdown] cell_id="00049-d1add3e1-42f5-44cd-b100-a33087e5b6ef" deepnote_cell_type="markdown" tags=[]
# In this section, we are going to test the model performance by using confusion matrix. Here are the steps:
#
# 1. Forward pass: compute predicted output by passing input data to the **trained** `model`
# 2. Get predicted label by retrieve index with the largest logit value per observation
# 3. Append actual and predicted label to `y_test` and `y_pred` respectively
# + cell_id="00023-89c9586b-0f02-4ab9-94ba-581764f30d9e" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=24 execution_start=1629447763230 source_hash="6c1d125c" tags=[]
# load the best model
model.load_state_dict(torch.load('cache/model-mnist-digit.pt'))
# + cell_id="00025-579d9054-2159-4b3e-af99-663262c87612" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=443 execution_start=1629447765791 source_hash="fbbe2310" tags=[]
y_test = []
y_pred = []
# disable gradient calculation
with torch.no_grad():
# prepare model for evaluation
model.eval()
# loop for each data
for data, target in test_loader:
# STEP 1: forward pass
output = model(data)
# STEP 2: get predicted label
_, label = torch.max(output, dim=1)
# STEP 3: append actual and predicted label
y_test += target.numpy().tolist()
y_pred += label.numpy().tolist()
# + [markdown] cell_id="00052-003f3060-15b4-4bda-97fb-a8f7f9103377" deepnote_cell_type="markdown" tags=[]
# Create a confusion matrix heatmap using `seaborn` and classification report by using `sklearn` to know the final model performance on test data.
# + cell_id="00027-bca38ce6-df6e-4bc1-878c-d03a89ba8e0c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=877 execution_start=1629448146336 source_hash="faad80df" tags=[]
# collapse-hide
plt.subplots(figsize=(10, 8))
ax = sns.heatmap(confusion_matrix(y_test, y_pred), annot=True, fmt="d")
ax.set_xlabel("Predicted Label")
ax.set_ylabel("Actual Label")
plt.show()
# + cell_id="00027-3fa978ed-3aa5-4579-94dd-4e8ea0615f7e" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=46 execution_start=1629448153350 source_hash="13ae2281" tags=[]
print(classification_report(y_test, y_pred))
# + [markdown] cell_id="00027-5359ca02-ee03-4955-81e2-d6196e71a386" deepnote_cell_type="markdown" tags=[]
# Eventually, the model achieves 97% accuracy and other metrics reach >= 92% on unseen data.
# + [markdown] cell_id="00029-cb30984d-3c83-4888-a5fb-254df5fa29a0" deepnote_cell_type="markdown" tags=[]
# # Predict the Unlabeled Data
# + [markdown] cell_id="00057-d272f6fe-dcb4-4036-aa6f-05732d7bb894" deepnote_cell_type="markdown" tags=[]
# In this last section, we use the trained model to predict the unlabeled data from `test.csv`, which only consists 784 columns of pixel without the actual label.
# + cell_id="00030-cc7edc59-8d81-4129-b9ef-0c9bdc30b11e" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1663 execution_start=1629448277442 source_hash="1cee7086" tags=[]
# load test.csv
mnist_data_unlabeled = MNISTDataset("data_input/mnist/test.csv")
unlabeled_loader = DataLoader(mnist_data_unlabeled, batch_size=1, shuffle=True)
print(mnist_data_unlabeled.X.size())
# + [markdown] cell_id="00059-3d761a25-d23d-4528-b8bd-27ee4c2ef1ec" deepnote_cell_type="markdown" tags=[]
# Each image is feeded into the model and `F.softmax` activation function will be applied on the output value to convert **logit scores to probability**. The visualization tells us the predicted probability given an image, and then class with the highest probability will be the final predicted label of that image.
# + cell_id="00031-e6361aa6-2470-4b0a-8dd8-23867ef550ea" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=968 execution_start=1629448280606 source_hash="1d1de797" tags=[]
# loop data
for idx in range(3):
# get image from loader
image = next(iter(unlabeled_loader))
image = image.reshape((28, 28))
##############
# prediction #
##############
with torch.no_grad():
# forward pass
output = model(image.reshape(1, -1))
# calculate probability
prob = F.softmax(output, dim=1)
#################
# visualization #
#################
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
# show digit image
axes[0].imshow(image, cmap="gray")
axes[0].axis("off")
# predicted probability barplot
axes[1].barh(range(0, 10), prob.reshape(-1))
axes[1].invert_yaxis()
axes[1].set_yticks(range(0, 10))
axes[1].set_xlabel("Predicted Probability")
plt.tight_layout()
plt.show()
# + [markdown] cell_id="00056-04c8a3ee-04ae-4b89-be6a-45f8739429f4" deepnote_cell_type="markdown" tags=[]
# # References
# + [markdown] cell_id="00057-d962454a-8705-42d3-b368-84ad7ca49f79" deepnote_cell_type="markdown" tags=[]
# - [PyTorch Installation](https://pytorch.org/get-started/locally/)
# - [MNIST Digit Recognizer Dataset](https://www.kaggle.com/c/digit-recognizer/data)
# - [Writing Custom Datasets, DataLoaders, and Transforms](https://pytorch.org/tutorials/beginner/data_loading_tutorial.html)
# -
# # Further Readings
# There's a lot of scope to experiment here, and I encourage you to use the interactive nature of Jupyter to play around with the various parameters. Here are a few ideas:
#
# * Try changing the size of the hidden layer, or add more hidden layers and see if you can achieve a higher accuracy.
#
# * Try changing the batch size and learning rate to see if you can achieve the same accuracy in fewer epochs.
#
# * Compare the training times on a CPU vs. GPU. Do you see a significant difference. How does it vary with the size of the dataset and the size of the model (no. of weights and parameters)?
#
# * Try building a model for a different dataset, such as the [CIFAR10 or CIFAR100 datasets](https://www.cs.toronto.edu/~kriz/cifar.html).
#
#
# Here are some references for further reading:
#
# * [A visual proof that neural networks can compute any function](http://neuralnetworksanddeeplearning.com/chap4.html), also known as the Universal Approximation Theorem.
#
# * [But what *is* a neural network?](https://www.youtube.com/watch?v=aircAruvnKk) - A visual and intuitive introduction to what neural networks are and what the intermediate layers represent
#
# * [Stanford CS229 Lecture notes on Backpropagation](http://cs229.stanford.edu/notes/cs229-notes-backprop.pdf) - for a more mathematical treatment of how gradients are calculated and weights are updated for neural networks with multiple layers.
| _notebooks/2021-08-27-mnist-digit-pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AI-LAB SESSION 1: Uninformed search
#
# In this first session we will work on uninformed search
#
# ## Maze environments
#
# The environments used are **SmallMaze** (visible in the figure) and its variations
# 
# The agent starts in cell $(0, 2)$ and has to reach the treasure in $(4, 3)$
#
# ## Assignment 1
#
# Your first assignment is to implement the BFS algorithm on **SmallMaze**. In particular, you are required to implement both *tree_search* and *graph_search* versions of BFS that will be called by the generic *bfs*.
#
# The results returned by your *bfs* must be a tuple $(path, stats)$ in the following form:
# * *path* - tuple of state identifiers forming a path from the start state to the goal state. ``None`` if no solution is found
# * *stats* - tuple of:
# * *time* - time elapsed between the start and the end of the algorithm
# * *expc* - number of nodes explored. A node is considered as explored when removed from the fringe and analyzed
# * *maxnodes* - maximum number of nodes in memory at the same time (fringe + closed)
#
# After the correctness of your implementations have been assessed, you can run the algorithms on other two maze environments: **GrdMaze** and **BlockedMaze**.
#
# Function *build_path* can be used to return a tuple of states from the root node to another node by following *parent* links
# +
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
import gym
import envs
from timeit import default_timer as timer
from utils.fringe import FringeNode, QueueFringe
explored = 0 # var global
def build_path(node):
"""
Builds a path going backward from a node
Args:
node: node to start from
Returns:
path from root to ``node``
"""
path = []
while node.parent is not None:
path.append(node.state)
node = node.parent
return tuple(reversed(path))
# -
# The next two functions have to be implemented
def tree_search(environment, fringe):
"""
Tree search
Args:
environment: OpenAI Gym environment
fringe: instance of Fringe data structure
Returns:
(path, stats): solution as a path and stats.
The stats are a tuple of (expc, maxnodes): number of explored states, max nodes in memory
"""
maxNodes=0; explored=0;
root = FringeNode(environment.startstate)
fringe.add(root);
while True:
explored=explored+1
if fringe.is_empty():
return None, (explored,maxNodes);
nodo = fringe.remove();
if nodo.state==environment.goalstate:
return build_path(nodo),(explored,maxNodes);
#4 azioni possibili
for action in range(env.action_space.n):
#esploro i vicini
child=environment.sample(nodo.state,action);
child=FringeNode(child,nodo);
#if child.state not in fringe:
#N.B: commentando le 2 righe successive faccio il controllo
#del goal state dopo e così viene come quello nelle soluzioni
#if child.state == environment.goalstate:
# return build_path(child), (explored,maxNodes)
fringe.add(child)
maxNodes = max(maxNodes, len(fringe))
# +
def graph_search(environment, fringe):
"""
Graph search
Args:
environment: OpenAI Gym environment
fringe: instance of Fringe data structure
Returns:
(path, stats): solution as a path and stats.
The stats are a tuple of (expc, maxnodes): number of explored nodes, max nodes in memory
"""
maxNodes=0; explored=0;
root = FringeNode(environment.startstate)
fringe.add(root);
closed=set();
while True:
if fringe.is_empty():
return None, (explored,maxNodes);
nodo = fringe.remove();
explored=explored+1;
if environment.goalstate == nodo.state:
return build_path(nodo),(explored,maxNodes);
closed.add(nodo.state)
#4 azioni possibili
for action in range(env.action_space.n):
#esploro i vicini --sample() lavora solo sugli indici interi
child=environment.sample(nodo.state,action);
child=FringeNode(child,nodo);
if child.state not in fringe and child.state not in closed:
if child.state == environment.goalstate:
return build_path(child), (explored,maxNodes)
fringe.add(child)
maxNodes=max(maxNodes, (len(fringe)+len(closed)))
# -
def bfs(environment, search_type):
"""
Breadth-first search
Args:
environment: OpenAI Gym environment
search_type: type of search - tree_search or graph_search (function pointer)
Returns:
(path, stats): solution as a path and stats.
The stats are a tuple of (time, expc, maxnodes): elapsed time, number of explored nodes, max nodes in memory
"""
t = timer()
path, stats = search_type(environment, QueueFringe())
return path, (timer() - t, stats[0], stats[1])
# The following code calls your tree search version of BFS and prints the results
# +
envname = "SmallMaze-v0" # Other options are GrdMaze-v0 and BlockedMaze-v0
envname2 = "GrdMaze-v0"
envname3 = "BlockedMaze-v0"
print("\n----------------------------------------------------------------")
print("\tTREE SEARCH")
print("\tEnvironment: ", envname)
print("----------------------------------------------------------------\n")
# Create and render the environment
env = gym.make(envname)
env.render()
solution, stats = bfs(env, tree_search) # Perform BFS
if solution is not None:
solution = [env.state_to_pos(s) for s in solution]
# Print stats and path
print("\n\nExecution time: {0}s\nN° of nodes explored: {1}\nMax n° of nodes in memory: {2}\nSolution: {3}".format(
round(stats[0], 4), stats[1], stats[2], solution))
# -
# Correct results for BFS tree search can be found [here](files/results/bfs_tree_search_results.txt)
#
# The following code calls your graph search version of BFS and prints the results
# +
envname = "SmallMaze-v0" # Other options are GrdMaze-v0 and BlockedMaze-v0
envname2= "GrdMaze-v0"
envname3= "BlockedMaze-v0"
print("\n----------------------------------------------------------------")
print("\tGRAPH SEARCH")
print("\tEnvironment: ", envname)
print("----------------------------------------------------------------\n")
# Create and render the environment
env = gym.make(envname)
env.render()
solution, stats = bfs(env, graph_search) # Perform BFS
if solution is not None:
solution = [env.state_to_pos(s) for s in solution]
# Print stats and path
print("\n\nExecution time: {0}s\nN° of nodes explored: {1}\nMax n° of nodes in memory: {2}\nSolution: {3}".format(
round(stats[0], 4), stats[1], stats[2], solution))
# -
# Correct results for BFS graph search can be found [here](files/results/bfs_graph_search_results.txt)
#
# ## Assignment 2
#
# Your second assignment is to implement the IDS algorithm on **SmallMaze**. In particular, you are required to implement both *dls_ts* (depth-limited tree search) and *dls_gs* (depth-limited graph search) versions of IDS that will be called by the generic *ids*. The recursions must be implemented in *rdls_ts* (recursive depth-limited tree search) and *rdls_gs* (recursive depth-limited graph search) called by *dls_ts* and *dls_gs* respectively.
#
# Similarly to assignment 1, the results returned by your *ids* must be a tuple $(path, stats)$ in the following form:
# * *path* - tuple of state identifiers forming a path from the start state to the goal state. ``None`` if no solution is found
# * *stats* - tuple of:
# * *time* - time elapsed between the start and the end of the algorithm
# * *expc* - number of nodes explored. A node is considered as explored when removed from the fringe and analyzed
# * *maxnodes* - maximum number of nodes in memory at the same time (the depth of the recursion stack + closed)
#
# After the correctness of your implementations have been assessed, you can run the algorithms on other two maze environments: **GrdMaze** and **BlockedMaze**.
#
# **FringeNode** has a useful variable that can be set in the constructor and can be used to track the depth of a node in the path (and consequently of the recursion stack of IDS): *pathcost*. If the root node has a *pathcost* of 0, its children will have a *pathcost* increased by 1
start = env.startstate
root = FringeNode(start) # parent = None and pathcost = 0 as default
child = FringeNode(env.sample(start, 0), root, root.pathcost + 1) # pathcost is the third argument
print("Root pathcost: {}\tChild pathcost: {}".format(root.pathcost, child.pathcost))
# Here you can implement the various functions requested
def dls_ts(environment, limit):
"""
Depth-limited search (tree search)
Args:
environment: OpenAI Gym environment
limit: depth limit budget
Returns:
(path, cutoff, stats): solution as a path, cutoff flag and stats.
The stats are a tuple of (time, expc, maxnodes): elapsed time, number of explored nodes, max nodes in memory
"""
t = timer()
path, cutoff, expc, maxdepth = rdls_ts(environment, FringeNode(environment.startstate), limit)
return path, cutoff, (timer() - t, expc, maxdepth)
# +
def rdls_ts(environment, node, limit):
"""
Recursive depth-limited search (tree search version)
Args:
environment: OpenAI Gym environment
node: node to explore
limit: depth limit budget
Returns:
(path, cutoff, expc, maxdepth): path, cutoff flag, number of explored nodes, max nodes in memory
"""
global explored;
explored = explored + 1;
maxdepth = node.pathcost
if node.state == environment.goalstate:
return build_path(node), False, explored, maxdepth
if limit == 0:
cutoff = True
return None, cutoff, explored, maxdepth;
cutoff_occurred = False;
for action in range(environment.action_space.n):
child = FringeNode(environment.sample(node.state, action), node, node.pathcost+1);
path, cutoff, explored, new_depth = rdls_ts(environment, child, limit - 1);
maxdepth = max(maxdepth, new_depth)
if cutoff == True: #result == cutoff
cutoff_occurred = True
elif not(path == None and cutoff == False): #result != failure
return path, cutoff, explored, maxdepth;
if cutoff_occurred:
cutoff = True
return None, cutoff, explored, maxdepth
return None, False, explored, maxdepth #return failure, so cutoff is false and result is none
# -
def dls_gs(environment, limit):
"""
Depth-limited search (graph search)
Args:
environment: OpenAI Gym environment
limit: depth limit budget
Returns:
(path, cutoff, stats): solution as a path, cutoff flag and stats.
The stats are a tuple of (time, expc, maxnodes): elapsed time, number of explored nodes, max nodes in memory
"""
t = timer()
path, cutoff, expc, maxdepth = rdls_gs(environment, FringeNode(environment.startstate), limit, closed=set())
return path, cutoff, (timer() - t, expc, maxdepth);
def rdls_gs(environment, node, limit, closed):
"""
Recursive depth-limited search (graph search version)
Args:
environment: OpenAI Gym environment
node: node to explore
limit: depth limit budget
closed: completely explored nodes
Returns:
(path, cutoff, expc, maxdepth): path, cutoff flag, n° of nodes explored, max nodes in memory
"""
global explored;
maxdepth = node.pathcost;
if node.state == environment.goalstate:
explored += 1
return (build_path(node), False, explored, maxdepth)
if limit == 0:
cutoff = True
if node.state not in closed:
explored += 1
closed.add(node.state)
return None, cutoff, explored, maxdepth
cutoff_occurred = False
if node.state not in closed:
explored += 1
closed.add(node.state)
for action in range(environment.action_space.n):
child = FringeNode(environment.sample(node.state, action), node, node.pathcost+1)
path, cutoff, explored, new_depth = rdls_gs(environment, child, limit - 1, closed)
maxdepth = max(maxdepth, new_depth)
if cutoff == True: #result == cutoff
cutoff_occurred = True
elif not(path == None and cutoff == False): #result != failure
return path, cutoff, explored, maxdepth
if cutoff_occurred:
cutoff = True
return None, cutoff, explored, maxdepth
return None, False, explored, maxdepth #return failure, so cutoff is false and result is none
def ids(environment, search_type):
"""
Iterative deepening depth-first search
Args:
environment: OpenAI Gym environment
search_type: type of search (graph or tree) - dls_gs or dls_ts (function pointer)
Returns:
(path, stats): solution as a path and stats.
The stats are a tuple of (time, expc, maxnodes): elapsed time, number of explored nodes, max nodes in memory
"""
# from depth = 0 to infinity
# solution, cutoff, stats = stype(problem, depth)
# return solution, (timer() - t, expc, maxdepth)
global explored
explored = 0
depth = 0
t = timer()
while True:
solution, cutoff, stats = search_type(environment, depth)
if cutoff is not True:
return solution, (timer() - t, stats[1], stats[2] + 1)
depth = depth + 1;
# The following code calls your tree search version of IDS and prints the results
# +
envname = "SmallMaze-v0" # Other options are GrdMaze-v0 and BlockedMaze-v0
envname2 = "GrdMaze-v0"
envname3 = "BlockedMaze-v0"
print("\n----------------------------------------------------------------")
print("\tTREE SEARCH")
print("\tEnvironment: ", envname)
print("----------------------------------------------------------------\n")
# Create and render the environment
env = gym.make(envname)
env.render()
solution, stats = ids(env, dls_ts) # Perform
if solution is not None:
solution = [env.state_to_pos(s) for s in solution]
# Print stats and path
print("\n\nExecution time: {0}s\nN° of nodes explored: {1}\nMax n° of nodes in memory: {2}\nSolution: {3}".format(
round(stats[0], 4), stats[1], stats[2], solution))
# -
# Correct results for IDS tree search can be found [here](files/results/ids_tree_search_results.txt)
#
# The following code calls your graph search version of BFS and prints the results
# +
envname = "SmallMaze-v0" # Other options are GrdMaze-v0 and BlockedMaze-v0
envname2 = "GrdMaze-v0"
envname3 = "BlockedMaze-v0"
print("\n----------------------------------------------------------------")
print("\tGRAPH SEARCH")
print("\tEnvironment: ", envname)
print("----------------------------------------------------------------\n")
# Create and render the environment
env = gym.make(envname)
env.render()
solution, stats = ids(env, dls_gs) # Perform BFS
if solution is not None:
solution = [env.state_to_pos(s) for s in solution]
# Print stats and path
print("\n\nExecution time: {0}s\nN° of nodes explored: {1}\nMax n° of nodes in memory: {2}\nSolution: {3}".format(
round(stats[0], 4), stats[1], stats[2], solution))
# -
# Correct results for IDS graph search can be found [here](files/results/ids_graph_search_results.txt)
#
# ## Discussion
#
# Now that you have correctly implemented both BFS and IDS what can you say about the solutions they compute? Are there significant differences in the stats?
Metodologie graph search ottimizzano il numero di nodi esplorati
| ai-lab/session1/session1_us.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## What is it for?
# The Dataset API enables you to build complex input pipelines from simple, reusable pieces. For example, the pipeline for an image model might aggregate data from files in a distributed file system, apply random perturbations to each image, and merge randomly selected images into a batch for training. The pipeline for a text model might involve extracting symbols from raw text data, converting them to embedding identifiers with a lookup table, and batching together sequences of different lengths. The Dataset API makes it easy to deal with large amounts of data, different data formats, and complicated transformations.
# ## Data structure
# * tf.data.Dataset: A sequence of elements.
#
# * elements: each elements contains one or more Tenor objects.
#
# ## How to create Dataset
# ### From Numpy arrays
# +
import numpy as np
import tensorflow as tf
np_array_1 = np.array([1,2,3,4])
np_array_2 = np.array([5,6,7,8])
dataset = tf.data.Dataset.from_tensor_slices((np_array_1, np_array_2))
# -
dataset.output_shapes, dataset.output_types
# there is a simpler api for just 1 tensor
# +
import numpy as np
import tensorflow as tf
np_array_1 = np.array([1,2,3,4])
dataset = tf.data.Dataset.from_tensor_slices((np_array_1))
# -
dataset.output_shapes, dataset.output_types
# If all of your input data fit in memory, the simplest way to create a Dataset from them is to convert them to tf.Tensor objects and use Dataset.from_tensor_slices().
# ### From TFRecords data
# The Dataset API supports a variety of file formats so that you can process large datasets that do not fit in memory. For example, the TFRecord file format is a simple record-oriented binary format that many TensorFlow applications use for training data. The tf.data.TFRecordDataset class enables you to stream over the contents of one or more TFRecord files as part of an input pipeline.
# Creates a dataset that reads all of the examples from two files.
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)
# The filenames argument to the TFRecordDataset initializer can either be a string, a list of strings, or a **tf.Tensor of strings. Therefore if you have two sets of files for training and validation purposes, you can use a tf.placeholder(tf.string) to represent the filenames, and initialize an iterator from the appropriate filenames**.
#
#
# ### Consuming text data: tf.data.TextLineDataset
# Many datasets are distributed as one or more text files. The tf.data.TextLineDataset provides an easy way to extract lines from one or more text files. Given one or more filenames, a TextLineDataset will produce one string-valued element per line of those files. Like a TFRecordDataset, TextLineDataset accepts filenames as a tf.Tensor, so you can parameterize it by passing a tf.placeholder(tf.string).
# ## Pre-processing data with Dataset.map
# ### Parsing tf.Example protocol buffer messages
# Many input pipelines extract tf.train.Example protocol buffer messages from a TFRecord-format file (written, for example, using tf.python_io.TFRecordWriter). Each tf.train.Example record contains one or more "features", and the input pipeline typically converts these features into tensors.
# +
# Transforms a scalar string `example_proto` into a pair of a scalar string and
# a scalar integer, representing an image and its label, respectively.
def _parse_function(example_proto):
features = {"image": tf.FixedLenFeature((), tf.string, default_value=""),
"label": tf.FixedLenFeature((), tf.int32, default_value=0)}
parsed_features = tf.parse_single_example(example_proto, features)
return parsed_features["image"], parsed_features["label"]
# Creates a dataset that reads all of the examples from two files, and extracts
# the image and label features.
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)
dataset = dataset.map(_parse_function)
# -
# ### Applying arbitrary Python logic with tf.py_func()
# ## Batching dataset elements
# +
import tensorflow as tf
sess = tf.Session()
dataset = tf.data.Dataset.range(20)
data_batch = dataset.batch(4)
iterator = data_batch.make_one_shot_iterator()
next_element = iterator.get_next()
with sess.as_default() as sess:
# tf.train.start_queue_runners()
for i in range(5):
value = sess.run(next_element)
print(value)
# -
# use tf.train.shuffle_batch
# +
import tensorflow as tf
sess = tf.Session()
dataset = tf.data.Dataset.range(20)
iterator = dataset.make_one_shot_iterator()
next_element = iterator.get_next()
# next_element.set_shape([1])
batch_elment = tf.train.shuffle_batch([next_element], batch_size=4, capacity=64, min_after_dequeue=32)
with sess.as_default() as sess:
tf.train.start_queue_runners()
for i in range(5):
value = sess.run(batch_elment)
print(value)
# -
# ## Randomly shuffling input data
# +
import tensorflow as tf
sess = tf.Session()
dataset = tf.data.Dataset.range(80)
dataset = dataset.repeat(2)
dataset = dataset.shuffle(buffer_size=160)
dataset = dataset.batch(16)
iterator = dataset.make_one_shot_iterator()
next_element = iterator.get_next()
# next_element.set_shape([1])
with sess.as_default() as sess:
for i in range(10):
value = sess.run(next_element)
print(value)
# -
| tf_dataset/tf_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#import libraries
import os
import numpy as np
import pandas as pd
import geopandas as gpd
import seaborn as sns
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.animation as animation
# +
#read geometry data from csv
counties_shp = gpd.read_file('mapdata201907311006/COUNTY_MOI_1080726.shp',encoding='utf-8')
countynames = counties_shp['COUNTYNAME']
# +
#Find all files in certain category
path = "company"
files=[]
for r,d,f in os.walk(path):
for file in f:
files.append(os.path.join(r,file))
company_data = pd.DataFrame()
# -
list_of_years=[]
for f in files:
year_month = f[-10:-4]
list_of_years.append(year_month)
company_data = pd.read_csv(f)
company_data = company_data.drop(company_data.columns[[0,1,2,4]],axis=1)
company_data['公司所在地']=company_data['公司所在地'].apply(lambda t: t[:3])
MoneyDic={'County':[],year_month:[]}
for c in countynames:
count = 0.0 # in million
for index, row in company_data.iterrows():
if row[0] == c :
count += (float)(row[1])/1000000.0
MoneyDic['County'].append(c)
MoneyDic[year_month].append(round(count,2))
#print(MoneyDic)
money_merge = pd.DataFrame(MoneyDic)
counties_shp=counties_shp.merge(money_merge,left_on=('COUNTYNAME'),right_on=('County'))
print(year_month)
#counties_shp
# +
# save all the maps in the charts folder
output_path = 'charts'
# counter for the for loop
i = 0
# set the min and max range for the choropleth map
vmin, vmax = 0, 10000
print(list_of_years)
# -
# start the for loop to create one map per year
for year in list_of_years:
fig = counties_shp.plot(column=year, cmap='Blues', figsize=(10,10), linewidth=0.8, edgecolor='0.8', vmin=vmin, vmax=vmax, legend=True, norm=plt.Normalize(vmin=vmin, vmax=vmax))
ax = fig.axis((119.2,122.5,21.5,25.5))
# remove axis of chart
fig.axis('off')
# add a title
fig.set_title('New Company in Taiwan', \
fontdict={'fontsize': '25',
'fontweight' : '3'})
fig.annotate(year,
xy=(0.1, .225), xycoords='figure fraction',
horizontalalignment='left', verticalalignment='top',
fontsize=25)
# this will save the figure as a high-res png in the output path. you can also save as svg if you prefer.
filepath = os.path.join(output_path, year+'_company.png')
chart = fig.get_figure()
chart.savefig(filepath, dpi=300)
#import sys
# #!{sys.executable} -m pip install imageio
import imageio
path = "charts"
imagefile_list = []
for r,d,f in os.walk(path):
for file in f:
imagefile_list.append(os.path.join(r,file))
sorted_imagefile_list=sorted(imagefile_list)
images = []
for filename in sorted_imagefile_list:
#print(filename[-3:])
if filename[-3:] == "png":
images.append(imageio.imread(filename))
imageio.mimsave('company.gif', images)
| 01_python_crawler/04_new_company/Animated per year and month.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SMIB system as in Milano's book example 8.1 (AVR added)
# %matplotlib widget
import numpy as np
import matplotlib.pyplot as plt
# ## Import system module
from iso_milano_ex8p1_4ord_avr_gov import iso_milano_ex8p1_4ord_avr_gov_class,daesolver
# ## Instantiate system
syst = iso_milano_ex8p1_4ord_avr_gov_class()
syst.initialize([{}],
1.0);
syst.report_x()
syst.report_y()
syst.simulate([{'t_end':1.0, 'P_l':0.8, 'H':6.0,'P_cig':0.1, 'T_m':5, 'D':0},
{'t_end':60.0, 'P_l':0.9, 'P_cig':0.1}],1.0);
# +
fig, axes = plt.subplots(nrows=2, ncols=1, figsize=(5, 4))
axes[0].plot(syst.T, syst.get_values('P_t') , label='$\sf P_t$')
axes[0].plot(syst.T, syst.get_values('p_m') , label='$\sf p_m$')
axes[0].plot(syst.T, syst.get_values('P_l') , label='$\sf P_l$')
axes[0].plot(syst.T, syst.get_values('P_cig') , label='$\sf P_{cig}$')
axes[1].plot(syst.T, syst.get_values('omega') , label='$\sf \omega$')
for ax in axes.flatten():
ax.grid()
ax.legend()
# +
Dp = 0.1
Dt = 1.0
idx = np.argmax(syst.T>(1.0+Dt))
Dw = 1-syst.get_values('omega')[idx]
RoCoF = Dw/Dt
print(f'RoCoF = {RoCoF*50:0.2f} Hz/s')
H = Dp/(2*RoCoF)
print(f'H = {H:0.2f} s')
# -
nadir = np.min(syst.get_values('omega'))
nadir*50
print(f'nadir = {nadir*50:0.2f} Hz')
print(f'Δf(nadir) = {50-nadir*50:0.2f} Hz')
# +
f_end = syst.get_values('omega')[-1]
Dw_end = 1.0-f_end
print(f'nadir = {nadir*50:0.2f} Hz')
print(f'Δf(nadir) = {50-nadir*50:0.2f} Hz')
print(f'f(end) = {f_end*50:0.2f} Hz')
print(f'Δf(end) = {50-f_end*50:0.2f} Hz')
Droop = Dw_end/Dp
print(f'Droop = {Droop:0.2f}')
# -
syst.simulate([{'t_end':1.0, 'P_l':0.8, 'H':3.0,'P_cig':0.1, 'T_m':5, 'D':0},
{'t_end':60.0, 'P_l':0.9, 'P_cig':0.1}],1.0);
# +
fig, axes = plt.subplots(nrows=2, ncols=1, figsize=(5, 4))
axes[0].plot(syst.T, syst.get_values('P_t') , label='$\sf P_t$')
axes[0].plot(syst.T, syst.get_values('p_m') , label='$\sf p_m$')
axes[0].plot(syst.T, syst.get_values('P_l') , label='$\sf P_l$')
axes[0].plot(syst.T, syst.get_values('P_cig') , label='$\sf P_{cig}$')
axes[1].plot(syst.T, syst.get_values('omega') , label='$\sf \omega$')
for ax in axes.flatten():
ax.grid()
ax.legend()
# +
Dp = 0.1
Dt = 1.0
idx = np.argmax(syst.T>(1.0+Dt))
Dw = 1-syst.get_values('omega')[idx]
RoCoF = Dw/Dt
print(f'RoCoF = {RoCoF*50:0.2f} Hz/s')
H = Dp/(2*RoCoF)
print(f'H = {H:0.2f} s')
nadir = np.min(syst.get_values('omega'))
nadir*50
print(f'nadir = {nadir*50:0.2f} Hz')
print(f'Δf(nadir) = {50-nadir*50:0.2f} Hz')
f_end = syst.get_values('omega')[-1]
Dw_end = 1.0-f_end
print(f'nadir = {nadir*50:0.2f} Hz')
print(f'Δf(nadir) = {50-nadir*50:0.2f} Hz')
print(f'f(end) = {f_end*50:0.2f} Hz')
print(f'Δf(end) = {50-f_end*50:0.2f} Hz')
Droop = Dw_end/Dp
print(f'Droop = {Droop:0.2f}')
# -
import numba
print(numba.__version__)
| examples/machines/iso_milano_ex8p1/iso_milano_ex8p1_4ord_avr_gov.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumPy ( _Numerical Python_ )
# Libreria que permite manipular grandes cantidades de datos por medio del uso de objetos especiales conocidos como arreglos o <span class="mark">arrays</span>, los cuales tiene un gran parecido con el tipo de datos list, pero con un manejo mucho mas optimizado hacia la manipulacion de datos de varias dimensiones. Se utiliza ampliamente en el mundo de las ciencias de datos pues son el fundamento de los DataFrames, objetos que permiten estudiar de manera tabular y grafica las relaciones entre los datos.
#
# Las listas por definicion son mas felxibles, pero los arrays de numpy son mas eficientes para el almacenamiento y manipulacion de datos.
# importando numpy
import numpy
numpy.__version__
# importando numpy como debe ser
import numpy as np
np.<TAB>
# +
# np?
# -
# Mas ayuda en: http://www.numpy.org.
# ## Arreglos en python
# ##### Creando arreglos en python
# +
import array
l = [0, 1, 2, 3, 4, 5, 6]
A = array.array('i', l)
# -
print(A)
print(type(A))
# ##### Creando arreglos con numpy
# A partir de listas. No es conveniente agregar mas de un tipo de datos al arreglo. Por ejemplo, no es buena practica mezclar enteros con strings.
array1 = np.array([1, 'a', 3, 4, 5])
print(array1)
# +
# Ejemplo de upcasting
array2 = np.array([3.14, 4, 2, 3])
print(array2)
# +
# Tambien es posible predefinir el tipo de datos
array2 = np.array([3.14, 4, 2, 3], dtype = "float32")
print(array2)
# +
# Arreglos multidimensionales
arreglo = np.array([[1, 2, 3], [2, 4, 6], [3, 8, 9]])
print(arreglo)
# -
# ##### Creando arreglos desde cero
# <span class="mark">np.zeros</span>: Util para crear un arreglo de ceros.
np.zeros(100, dtype = "int")
np.zeros((3, 5), dtype = "int")
# <span class="mark">np.ones</span>: Util para crear arreglos de unos
np.ones(100)
np.ones((7, 5), dtype = "int")
# <span class="mark">np.full</span>: Util para crear un arreglo lleno con lo que se especifique.
print(np.full(15, "Hola"))
print(np.full((7, 8), 3.14))
# <span class="mark">np.arange</span>: Util para crear un arreglo de una secuencia desde un valor inicial hasta un valor final. Se puede especificar un tercer argumento para establecer un salto. Si se da solo un argumento, el arreglo comenzara desde cero.
np.arange(5)
np.arange(3, 15)
np.arange(3, 20, 2)
# <span class="mark">np.linspace</span>: Util para crear un arreglo de una cantidad de numeros entre dos valores establecidos. Los valores retornados estaran igualmente espaciados.
# 5 valores de 0 a dos
np.linspace(0, 2, 5, dtype = "float32")
# 30 valores de 0 a 1
np.linspace(0, 1, 30)
# Tambien se puede al reves
np.linspace(10, 2, 6)
# Y tambien con negativos
np.linspace(-5, 5, 10)
# <span class="mark">np.random.random</span>: Util para crear arreglos de numeros aleatorios.
# Un numero aleatorio
np.random.random(1)
# Varios numeros aleatorios
print(np.random.random((4, 7)))
# Varios numeros aleatorios
np.random.random((1, 7))
# <span class="mark">np.random.normal</span>: Util para crear un arreglo de numeros aleatorios de ciertas dimensiones a partir de una distribucion normal.
# Matriz de 3x3 a partir de una distribucion normal estandar con media cero y desvest 1
np.random.normal(0, 1, (3, 3))
# <span class="mark">np.random.randint</span>: Lista de numeros enteros aleatorios en un intervalo dado
# Matriz 3x3 de valores enteros de 0 a 9
np.random.randint(0, 10, (3, 3))
# <span class="mark">np.eye</span>: Utli para crear matrices indentidad.
np.eye(4)
# <span class="mark">np.empty</span>: permite crear un array vacio, muy util para llenarlo despues de lo que queramos.
np.empty(10)
# ## Tipos de datos en NumPY
# Se habia mencionado que lo recomendado es crear arreglos con el mismo tipo de datos, buscando sobre todo la eficiencia en la gestion de la memoria y el manejo de las operaciones. A continuacion se comparte una tabla con los tipos de datos que se manejan con numpy y el prefijo dtype = ""
#
# Tabla tomada de: _Python Data Science Handbook, <NAME>, 2016, O'Reilly Media, Inc._
# 
# ## Atributos de los arrays
# +
import numpy as np
np.random.seed(0) # semilla para que se produzcan siempre los mismos resultados
x1 = np.random.randint(100, size = 6) # una dimension
x2 = np.random.randint(100, size = (3, 4)) # dos dimensiones
x3 = np.random.randint(100, size = (3, 4, 5)) # tres dimensiones
# -
print(x1, end = "\n"*2)
print(x2, end = "\n"*2)
print(x3, end = "\n"*2)
# <span class="mark">ndim</span>, <span class="mark">shape</span> y <span class="mark">size</span> son tres importantes atributos utlizados con frecuencia para obtener informacion de los arreglos. A continuacion se muestra su uso:
print("x3 ndim: ", x3.ndim) # Dimension del arreglo
print("x3 shape:", x3.shape) # Forma del arreglo
print("x3 size: ", x3.size) # Cantidad de elementos
# <span class="mark">dtype</span> es un atributo no tan comun pero bastante util a la hora de evaluar los tipos de datos almacenados:
#
print("dtype:", x3.dtype) # Tipo de datos almacenados
# ## Indexado en arreglos
# El indexado en los arreglo funciona de forma similar que en las listas, con la excepcion de la multidimesionalodad. A continuacion se ilustrara este hecho:
x1
print(x1[2])
print(x1[0])
print(x1[4])
print(x1[-1])
print(x1[-2])
print(x1[-4])
x2
print(x2[0, 1])
print(x2[1, 3])
print(x2[2, 3])
print(x2[0, -1])
print(x2[1, -3])
print(x2[-2, -3])
# ## Slicing en arreglos
# Este concepto tambien funciona de forma similar que en las listas, con la estructura base
#
# x[start:stop:step]
# ##### Arreglos unidimensionales
# Arreglo de ejemplo
x = np.arange(10)
print(x)
# Elementos del 1 al 3
print(x[1:4])
# Todos los elementos hasta el 5
x[:5]
# Arreglo al reves
x[::-1]
# Saltando de dos en dos
x[::2]
# ##### Arreglos multidimensionales
print(x2)
# Filas 0 y 1 y columnas 0, 1 y 2
x2[:2, :3]
# Filas 0, 1 y 2, y columnas de dos en dos
x2[:3, ::2]
# Dandole la vuelta a todo
x2[::-1, ::-1]
# Fila 2
x2[1,:]
# Columna 3
x2[:, 2]
# ##### El problema de la copia de arrays
x2
# Creando una submatriz a partir de una extraccion a x2
x2_sub = x2[:2, :2]
print(x2_sub)
# Modificando a x2_sub
x2_sub[0, 0] = 100
print(x2_sub)
# X2 fue afectado!
print(x2)
# Esto nos muestra que no se puede simplemente asignar una array a otro si lo que queremos es crear una copia. PAra ello debemos usar la instruccion <span class="mark">.copy()</span>, al igual que lo hicimos con las listas.
print(x2)
# Nueva extraccion de x2
x3_sub = x2[:2, :2].copy()
print(x3_sub)
# Modificando a x3_sub
x3_sub[0, 0] = 99
print(x3_sub)
# X2 no se vio afectado con el cambio a x3
print(x2)
# ## <span class="burk">MINIDESAFIO</span>
# **1.** Crea un array o arreglo unidimensional donde le indiques el tamaño por teclado, y ademas crea una función que rellene el array con numeros solicitados por teclado. Muestralos por pantalla.
#
# **Tip**: el metodo <span class="mark">.append()</span> que se utiliza para agregar elementos a una lista vacia tambien funciona, y de la misma manera, con arreglos
#
# +
dimension = int(input("Introduzca el tamanio del arreglo unidimensional: " ))
arreglo = np.ones(dimension, dtype = "int")
bandera = 0
while bandera < dimension:
valor = int(input("Introduzca el valor: "))
arreglo[bandera]*=valor
bandera += 1
print(arreglo)
# -
# **2.** Cree dos arreglos unidimensionales del mismo tamanio. El tamanio se debe pedir por teclado. En el primero almacene nombres de paises y en el segundo sus capitales.
# +
dimension = int(input("Introduzca el tamanio del arreglo unidimensional: " ))
arreglo1 = np.full(dimension, "*******************")
arreglo2 = np.full(dimension, "*******************")
for i in range(dimension):
pais = input("Introduzca el pais: ")
capital = input("Introduzca la capital: ")
arreglo1[i] = pais
arreglo2[i] = capital
print(arreglo1)
print(arreglo2)
# -
# **3.** Investigue que es la transpuesta de una matriz. Cree una funcion que tome una matriz cuadrada de cualquier tamanio y devuelva se transpuesta. Procure que la matriz sea cuadrada y de numeros aleatorios. Al final, compruebe que su resultado es igual que aplicar la operacion .T:
#
# matriz.T
matriz = np.array([[1, -7, 3], [2, 4, 6], [0, 3, 9]])
print(matriz, end= "\n"*4)
print(matriz.T)
# ## Redimensionando arreglos
# Muchas veces queremos redimensionar arreglos por diferentes motivos, por ejemplo, cuandon deseamos pasar un arreglo unidimensional a otro multidimensional. Para ello usaremos la instruccion <span class="mark">reshape()</span>:
uni = np.arange(9)
print(uni)
bi = uni.reshape((3, 3))
print(bi)
# Es importante tener en cuenta que los tamanios deben ser coherentes, de lo contrario se obtendra un error.
print(x3)
print(x3.size)
print(x3.reshape(60))
# ## Concatenacion de arreglos y particiones
# La concatenacion es la union de dos arreglos en uno solo. Esto se puede hacer por filas o por columnas mientras que las dimensiones encajen, de lo contrario puede dar lugar a errores. PAra esto usaremos la funcion <span class="mark">concatenate()</span>
a = np.array([1, 2, 3])
b = np.array([3, 2, 1])
print(a, b)
print(np.concatenate([a, b]))
print(x2)
# una matriz sobre la otra: axis = 0 es por filas
print(np.concatenate([x2, x2], axis = 0))
# una matriz al lado de la otra: axis = 1 es por filas
print(np.concatenate([x2, x2], axis = 1))
# Splitting o particionamiento es la operacion contraria, es decir, separar un array en dos o mas diferentes, con la condicion de que coincidan las dimensiones. Para particionar arreglos unidimensionales se usara la instruccion <span class="mark">.split()</span>:
x = [1, 2, 3, 99, 99, 3, 2, 1]
x1, x2, x3 = np.split(x, [3, 5])
print(x1, x2, x3)
x = [1, 2, 3, 99, 99, 3, 2, 1]
x1, x2 = np.split(x, [3])
print(x1, x2)
grid = np.arange(16).reshape((4, 4))
print(grid)
# Para particionar arreglos multidimensionales, es necesario especificar si se quiere hacer la particion por filas o por columnas, ademas de indicar por medio de una lista los lugares por donde se particionara. Para realizar una particion en columnas, es decir, una particion vertical, se usa <span class="mark">.vsplit()</span> y para las horizontales se usara <span class="mark">.hsplit()</span>
# USando vsplit para particionar arreglos multidimensionales
upper, lower = np.vsplit(grid, [2])
print(upper)
print(lower)
# Usando hsplit para particionar arreglos multidimensionales
left, right = np.hsplit(grid, [2])
print(left)
print(right)
# ## <span class="burk">MINIDESAFIO</span>
# **1.** Acceda al siguiente enlace y aprenda un poco sobre multiplicacion de matrices:
#
# https://www.problemasyecuaciones.com/matrices/multiplicar-matrices-producto-matricial-ejemplos-explicados-propiedades-matriz.html
#
# Con esa informacion clara, cree una funcion que reciba como argumentos dos matrices cuadradas y devuelva el producto de estas.
#
# Matrices:
# 
# **2.** Usando la funcion creada en el punto 1, intente hacer la operacion inversa de esas dos matrices. Seguramente necesitara redimensionar alguna o las dos matrices. Imprima el resultado en la pantalla.
# **Nota:** Es posible que haya tenido alguna dificultad para realizar las anteriores operaciones...a continuacion le muestro una alternativa que seguramente le gustara mas:
import numpy as np
A = np.array([[1, 2], [-2, 0]])
B = np.array([[1, 0, 2], [0, 2, 0]])
print(A)
print(B)
AB = np.dot(B.reshape(3,2), A)
print(AB)
# # Computacion en numpy: Funciones universales
# En esta seccion podremos comprobar de primera mano el porque de la importancia de numpy en el area de las ciencias de datos. Se aprendera sobre el concepto _vectorizacion_ , la cual sera una tecnica que nos permitira dejar atrs los lentos ciclos (no en todos los casos), y optimizar nuestros programas para que sean mucho mas rapidos, lo cual es escencial en el manejo de grandes cantidades de datos.
# ## Python es lento!!! (seccion basada en el libro _Python Data Science Handbook, <NAME>, 2016, O'Reilly Media, Inc._ )
# fijando la semilla de los numeros aleatorios
np.random.seed(7)
np.random.randint(1, 10)
# prueba de ello:
import numpy as np
np.random.seed(0)
def compute_reciprocals(values):
output = np.empty(len(values))
for i in range(len(values)):
output[i] = 1.0 / values[i]
return output
values = np.random.randint(1, 10, size = 5)
compute_reciprocals(values)
big_array = np.random.randint(1, 100, size=1000000)
# %timeit compute_reciprocals(big_array)
# ## Introduciendo Ufuncs
# Como se menciono anteriormente, python tiene ciertos problemas de eficiencia que se manifiestan cuando se manejan muchos datos, por ello se creo la posibilidad de vectorizar las operaciones, lo que permite que las operaciones que apliquemos sobre los arrays, terminen siendo aplicadas directamente a cada elemento del array.
print(compute_reciprocals(values))
print(1.0 / values)
# %timeit (1.0 / big_array)
# Operaciones entre arrays
print(np.arange(5)/np.arange(1,6))
# Inclusive para arreglos multidimensionales
arreglo = np.arange(9).reshape((3,3))
2**arreglo
# Las ufuncs presentan dos variadades, las ufuncs unarias, que operan sobre una sola entrada, y las ufuncs binarias, que operan sobre sobre dos entradas.
# +
# Ejemplos de broadcasting gracias a numpy y al uso de ufuncs
x = np.arange(9)
print("x: ", x)
print("x + 10: ", x + 10)
print("x - 10: ", x - 10)
print("x * 10: ", x * 10)
print("x / 10: ", x / 10)
print("x // 10: ", x // 10)
print("x % 10: ", x % 10)
print("-x: ", -x)
print("(5*x + 2)**2: ", (5*x + 2)**2)
# -
# <span class="mark">np.abs()</span>: Valor absoluto
y = np.arange(-10, 5)
print("y: ", y)
print("|y|: ", np.abs(y))
# <span class="mark">Funciones trigonometricas</span>:
theta = np.linspace(0, np.pi, 3)
print("theta: ", theta)
print("sin(theta): ", np.sin(theta))
print("cos(theta): ", np.cos(theta))
print("tan(theta): ", np.tan(theta))
# <span class="mark">Funciones trigonometricas inversas</span>:
x = [-1, 0, 1]
print("x: ", theta)
print("arcsin(theta): ", np.arcsin(x))
print("arccos(theta): ", np.arccos(x))
print("arctan(theta): ", np.arctan(x))
# <span class="mark">Exponentes y logaritmos</span>:
x = [1, 2, 3]
print("x: ", x)
print("e^x: ", np.exp(x))
print("2^x: ", np.exp2(x))
print("3^x: ", np.power(3, x))
print("ln(x): ", np.log(x))
print("log2(x): ", np.log2(x))
print("log10: ", np.log10(x))
# ##### Ufuncs especializadas
from scipy import special
# Funcion gamma y relacionadas
x = [1, 5, 10]
print("gamma(x): ", special.gamma(x))
print("ln|gamma(x)|", special.gammaln(x))
print("beta(x, 2): ", special.beta(x, 2))
# Funcion error, su complemento e inversa
x = np.array([0, 0.3, 0.7, 1.0])
print("erf(x) =", special.erf(x))
print("erfc(x) =", special.erfc(x))
print("erfinv(x) =", special.erfinv(x))
# ## <span class="burk">MINIDESAFIO</span>
# **1.** Cree un arreglo unidimensional con numero desde -100 hasta 100. Luego de esto calcule el valor obtenido de estos valores al evaluarlos en la siguiente funcion usando ciclos for y midiendo el tiempo de ejecucion con el metodo magico %timeit:
#
# $f(x) = \frac{\cos(x - 1) + 7x - 2}{sen(2x - 3) -7x^2 + 2}$
#
# **2.** Partiendo del punto 1, vuelva a evaluar la misma funcion pero utilizando ufuncs, y mida el tiempo nuevamente.
#
# **3.** Saque alguna conclusion de los puntos anteriores.
import math
arreglo = np.arange(-100, 100)
def funcion(array):
resultado = []
for i in array:
numerador = math.cos(i - 1) + 7*i - 2
denominador = math.sin(2*i - 3) - 7*i**2+2
func = numerador / denominador
resultado.append(func)
return resultado
# %timeit funcion(arreglo)
import numpy as np
arreglo = np.arange(-100, 100)
# %timeit numerador = (np.cos(arreglo - 1) + 7*arreglo - 2)/(np.sin(2*arreglo - 3) - 7*arreglo**2 + 2)
# ## Caracteristicas avanzadas de las ufuncs
# ### Especificando la salida
# Muchas veces es bastante util especificar la salida en donde los calculos se almacenaran, para luego darles uso. Cuando se maneja el operador de asignacion =, muchas veces se pueden obtener errore de copiado como se vio antes. Para esto numpy ofrece la posibilidad de especificar la salidad con el argumento <span class="mark">out</span>.
# usando el argumento out
x = np.arange(5)
y = np.empty(5)
np.multiply(x, 10, out=y) # np.multiply es equivalente a x*10
print(y)
# usando ademas slicing
y = np.zeros(10)
print(y)
np.power(2, x, out=y[::2]) # np.power es equivalente a 2**x
print(y)
# ### Agregados
# Funciones especiales que se aplican directamente sobre un solo objeto, tomando todos los elementos para el calculo.
# Usando reduce
x = np.arange(1, 10)
print(x)
print(np.add.reduce(x))
print(np.multiply.reduce(x))
# Usando acuumulate
print(x)
print(np.add.accumulate(x))
print(np.multiply.accumulate(x))
# ## Min, Max y todo lo demas
# +
# usando la version rapida de sum():
arreglo = np.random.random(100)
print(arreglo, end = "\n"*2)
print(np.sum(arreglo))
# -
arreglo2 = np.random.random((5, 5))
print(arreglo2, end = "\n"*2)
print(np.sum(arreglo2))
# Usando min y max en sus versiones rapidas
print(np.min(arreglo), np.max(arreglo))
print(np.min(arreglo2), np.max(arreglo2))
# Usando el argumento axis (0: columnas, 1: filas)
print(np.min(arreglo2, axis = 0), np.max(arreglo2, axis = 0), end = "\n"*3)
print(np.min(arreglo2, axis = 1), np.max(arreglo2, axis = 1))
print(np.sum(arreglo2, axis = 0), end = "\n"*3)
print(np.sum(arreglo2, axis = 1))
# Otros metodos a tener en cuenta:
# 
# ## <span class="burk">MINIDESAFIO</span>
# **1.** Calcular los valores de media, dessviacion estandar, maximo y minimo del siguiente conjunto de valores:
#
# [189 170 189 163 183 171 185 168 173 183 173 173 175 178 183 193 178 173 174 183 183 168 170 178 182 180 183 178 182 188 175 179 183 193 182 183 177 185 188 188 182 185]
#
# Imprimir sus valores en pantalla.
#
# **2.** Con los mismos datos anteriores, calcular el percentil 25, la mediana (la cual es igual al percentil 50) y el percentil 75.
datos = np.array([189, 170, 189, 163, 183, 171, 185, 168, 173, 183, 173, 173, 175, 178, 183, 193, 178, 173, 174,
183, 183, 168, 170, 178, 182, 180, 183, 178, 182, 188, 175, 179, 183, 193, 182, 183, 177, 185,
188, 188, 182, 185])
print(np.mean(datos))
print(np.std(datos))
print(np.max(datos))
print(np.min(datos))
print(np.percentile(datos, 25))
print(np.median(datos))
print(np.percentile(datos, 75))
# +
# Como se veria la distribucion de los datos
# %matplotlib notebook
import matplotlib.pyplot as plt
datos = np.array([189, 170, 189, 163, 183, 171, 185, 168, 173, 183, 173, 173, 175, 178, 183, 193, 178, 173, 174,
183, 183, 168, 170, 178, 182, 180, 183, 178, 182, 188, 175, 179, 183, 193, 182, 183, 177, 185,
188, 188, 182, 185])
plt.style.use("ggplot")
plt.hist(datos, edgecolor = "k")
plt.title('Height Distribution of US Presidents')
plt.xlabel('datos')
plt.ylabel('number');
# -
# ## Computacion sobre arreglos: Broadcasting
# Ya se ha hablado del bradcasting por medio de ejemplos en las secciones anteriores. El broadcasting se puede definir como una extension que realiza numpy sobre ciertos datos cuando estos se quieren operar con otros mas extensos o de mayor dimension. A continuacion veremos mas ejemplo de esta practica que permite vectorizar nuestras operaciones
"""Ejemplo con suma"""
a = np.array([0, 1, 2])
b = np.array([5, 5, 5])
print(a)
print(b, end = "\n"*2)
print(a + b)
print(a)
print(a + 5)
matriz = np.ones((3,3))
print(matriz)
print(matriz + np.arange(3))
"""Ejemplo con vectores fila y columna"""
a = np.arange(3)
b = np.arange(3)[:, np.newaxis]
print(a)
print(b)
print(a+b)
# 
# ### Reglas del broadcasting
# • **Regla 1:** Si las dos matrices difieren en su número de dimensiones, la forma de la que tiene menos dimensiones se rellena con unos en su lado principal (izquierdo).
#
# • **Regla 2:** Si la forma de las dos matrices no coincide en alguna dimensión, la matriz con forma igual a 1 en esa dimensión se estira para que coincida con la otra forma.
#
# • **Regla 3:** Si en alguna dimensión los tamaños no concuerdan y ninguno es igual a 1, se genera un error.
"""Ejemplo 1"""
M = np.ones((2, 3))
a = np.arange(3)
print(M, end = "\n"*2)
print(a, end = "\n"*2)
print(M + a)
# 
"""Ejemplo 2"""
a = np.arange(3).reshape((3, 1))
b = np.arange(3)
print(a, "\n"*2)
print(b, "\n"*2)
print(a + b)
# 
"""Ejemplo 3"""
M = np.ones((3, 2))
a = np.arange(3)
print(M, "\n"*2)
print(a, "\n"*2)
print(M + a)
# 
# ## Operadores logicos con ufuncs
x = np.random.randint(1, 50, 5)
print(x)
print(x < 30)
print(x != 41)
print(x == 38)
# Equivalencia de los operadores logicos
# 
matriz = np.random.randint(1, 100, 9).reshape((3,3))
print(matriz)
print(matriz < 50)
print(np.sum((matriz > 10) & (matriz < 50)))
# Equivalencia de los operadores logicos con ufuncs:
# 
print(np.sum((matriz > 10) | (matriz < 50)))
print(np.sum(matriz == 60))
"""Ejemplo de enmascaramiento"""
nueva = matriz[[matriz > 54]]
print(nueva)
# ## <span class="burk">MINIDESAFIO</span>
# **1.** Construir una funcion que calcule el producto de dos matrices usando numpy.
#
# **2.** Construir una funcion que calcule le producto punto de dos vectores usando numpy.
#
# **3.** Cree un array con elmentos aleatorios del -200 al 200. Desde ese array cree otros arrays que contengan a los positivos, a los negativos y a los mayores a 40.
#
# **4.** (Tomado de: http://www.denebola.org/japp/CC/numpy.html) Crea un array bidimensional 5x5 con todos los valores cero. Usando el indexado de arrays, asigna 1 a todos los elementos de la última fila y 5 a todos los elementos de la primera columna. Finalmente, asigna el valor 100 a todos los elementos del subarray central 3x3 de la matriz de 5x5.
vector1 = np.random.randint(1, 1000, 3)
vector2 = np.random.randint(1, 1000, 3)
print(vector1, vector2)
print(np.dot(vector1, vector2))
print(np.cross(vector1, vector2))
a.(bxc)
ax(b.c)
v1 = (a, b, 0)
v2 = (e, d, 0)
(b*f-d*c, e*c-a*f, a*d-b*e)
(v1[1]*v2[2] - v2[1]*v1[2],
| Semana-3/Numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Duplicate Data
#
# A data set might have duplicate data: in other words, the same record is represented multiple times. Sometimes, it's easy to find and eliminate duplicate data like when two records are exactly the same. At other times, like what was discussed in the video, duplicate data is hard to spot.
#
# # Exercise 1
#
# From the World Bank GDP data, count the number of countries that have had a project totalamt greater than 1 billion dollars (1,000,000,000). To get the count, you'll have to remove duplicate data rows.
# +
import pandas as pd
# read in the projects data set and do some basic wrangling
projects = pd.read_csv('../data/projects_data.csv', dtype=str)
projects.drop('Unnamed: 56', axis=1, inplace=True)
projects['totalamt'] = pd.to_numeric(projects['totalamt'].str.replace(',', ''))
projects['countryname'] = projects['countryname'].str.split(';', expand=True)[0]
projects['boardapprovaldate'] = pd.to_datetime(projects['boardapprovaldate'])
# TODO: filter the data frame for projects over 1 billion dollars
# TODO: count the number of unique countries in the results
# -
# # Exercise 2 (challenge)
#
# This exercise is more challenging. The projects data set contains data about Yugoslavia, which was an Eastern European country until 1992. Yugoslavia eventually broke up into 7 countries: Bosnia and Herzegovina, Croatia, Kosovo, Macedonia, Montenegro, Serbia, and Slovenia.
#
# But the projects dataset has some ambiguity in how it treats Yugoslavia and the 7 countries that came from Yugoslavia. Your task is to find Yugoslavia projects that are probably represented multiple times in the data set.
# +
# TODO: output all projects for the 'Socialist Federal Republic of Yugoslavia'
# HINT: You can use the exact country name or use the pandas str.contains() method to search for Yugoslavia
# -
# Yugoslavia officially ended on [April 27th, 1992](https://en.wikipedia.org/wiki/Yugoslavia).
#
# In the code cell below, filter for projects with a 'boardapprovaldate' prior to April 27th, 1992 **and** with 'countryname' Bosnia and Herzegovina, Croatia, Kosovo, Macedonia, Serbia **or** Slovenia. You'll see there are a total of 12 projects in the data set that match this criteria. Save the results in the republics variable
# +
import datetime
# TODO: filter the projects data set for project boardapprovaldate prior to April 27th, 1992 AND with countryname
# of either 'Bosnia and Herzegovina', 'Croatia', 'Kosovo', 'Macedonia', 'Serbia', or 'Sovenia'. Store the
# results in the republics variable
#
# TODO: so that it's easier to see all the data, keep only these columns:
# ['regionname', 'countryname', 'lendinginstr', 'totalamt', 'boardapprovaldate',
# 'location','GeoLocID', 'GeoLocName', 'Latitude','Longitude','Country', 'project_name']
# TODO: sort the results by boardapprovaldate
republics = None
# show the results
republics
# -
# Are these projects also represented in the data labeled Yugoslavia? In the code cell below, filter for Yugoslavia projects approved between February 1st, 1980 and May 23rd, 1989 which are the minimum and maximum dates in the results above. Store the results in the yugoslavia variable.
#
# The goal is to see if there are any projects represented more than once in the data set.
# +
# TODO: Filter the projects data for Yugoslavia projects between
# February 1st, 1980 and May 23rd, 1989. Store the results in the
# Yugoslavia variable. Keep the same columns as the previous code cell.
# Sort the values by boardapprovaldate
yugoslavia = None
# show the results
yugoslavia
# -
# And as a final step, try to see if there are any projects in the republics variable and yugoslavia variable that could be the same project.
#
# There are multiple ways to do that. As a suggestion, find unique dates in the republics variable. Then separately find unique dates in the yugoslavia variable. Concatenate (ie append) the results together. And then count the number of times each date occurs in this list. If a date occurs twice, that means the same boardapprovaldate appeared in both the Yugoslavia data as well as in the republics data.
#
# You'll should find that there are three suspicious cases:
#
# * July 26th, 1983
# * March 31st, 1987
# * October 13th, 1987
# * May 23rd, 1989
# +
import numpy as np
# TODO: find the unique dates in the republics variable
republic_unique_dates = None
# TODO: find the unique dates in the yugoslavia variable
yugoslavia_unique_dates = None
# TODO: make a list of the results appending one list to the other
dates = None
# TODO: print out the dates that appeared twice in the results
# -
# # Conclusion
#
# On July 26th, 1983, for example, projects were approved for Bosnia and Herzegovina, Croatia, Macedonia, Slovenia, and Yugoslavia. The code below shows the projects for that date. You'll notice that Yugoslavia had two projects, one of which was called "Power Transmission Project (03) Energy Managem...". The projects in the other countries were all called "POWER TRANS.III".
#
# This looks like a case of duplicate data. What you end up doing with this knowledge would depend on the context. For example, if you wanted to get a true count for the total number of projects in the data set, should all of these projects be counted as one project?
#
# Run the code cell below to see the projects in question.
# +
import datetime
# run this code cell to see the duplicate data
pd.concat([yugoslavia[yugoslavia['boardapprovaldate'] == datetime.date(1983, 7, 26)], republics[republics['boardapprovaldate'] == datetime.date(1983, 7, 26)]])
# -
| lessons/ETLPipelines/11_duplicatedata_exercise/.ipynb_checkpoints/11_duplicatedata_exercise-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Apply different Kernels
# #### In Covolutional Neural Network (CNN), the images pass through different type of filters, in purpose to extract features.
# #### In this simple notebook, I tried to apply different kind of kernels (filters), as an attempt to totally absorb the concept of convolution, and to know how CNN extracts its own features from images!
from PIL import Image
from numpy import array
import matplotlib.pyplot as plt
from scipy import signal
from scipy import misc
import numpy as np
import cv2
img = cv2.imread("images/cat.jpg",0)
img.shape
# #### This function takes the image's martix, and the kernel's as well, the convolve them using 'signal.convolve2d'
def DisplayImagesBeforeAfterConvolution(image, kernel):
plt.figure(figsize=(10,10))
plt.subplot(121),plt.imshow(image),plt.title('Original')
plt.xticks([]), plt.yticks([])
filtered_image1 = signal.convolve2d(image,kernel)
plt.subplot(122),plt.imshow(filtered_image1),plt.title('Filtered')
plt.xticks([]), plt.yticks([])
plt.show()
# ### 1- Horizontal edge detection kernel
# \begin{equation}
# \begin{bmatrix}
# -1 & -2 & -1 \\
# 0 & 0 & 0 \\
# 1 & 2 & 1
# \end{bmatrix}
# \label{eq:aeqn}
# \end{equation}
kernel1 = np.array([[-1,-2,-1],[0,0,0],[1,2,1]])
DisplayImagesBeforeAfterConvolution(img, kernel1)
# ### 2- Vertical edge detection kernel
# \begin{equation}
# \begin{bmatrix}
# -1 & 0 & 1 \\
# -2 & 0 & 2 \\
# -1 & 0 & 1
# \end{bmatrix}
# \label{eq:aeqn}
# \end{equation}
kernel2= np.array([[-1,0,1],[-2,0,2],[-1,0,1]])
DisplayImagesBeforeAfterConvolution(img, kernel2)
# ### 3- Inverse
# \begin{equation}
# \begin{bmatrix}
# -1 & -1 & -1 \\
# -1 & -1 & -1 \\
# -1 & -1 & -1
# \end{bmatrix}
# \label{eq:aeqn}
# \end{equation}
kernel4= np.array([[-1,-1,-1],[-1,-1,-1],[-1,-1,-1]])
DisplayImagesBeforeAfterConvolution(img, kernel4)
# ### 4- Sharpness kernel
# \begin{equation}
# \begin{bmatrix}
# 0 & -1 & 0 \\
# -1 & 5 & -1 \\
# 0 & -1 & 0
# \end{bmatrix}
# \label{eq:aeqn}
# \end{equation}
kernel4= np.array([[0,-1,0],[-1,5,-1],[0,-1,0]])
DisplayImagesBeforeAfterConvolution(img, kernel4)
# ### 5- Smoothness kernel
# \begin{equation}
# \begin{bmatrix}
# 1/9 & 1/9 & 1/9 \\
# 1/9 & 1/9 & 1/9 \\
# 1/9 & 1/9 & 1/9
# \end{bmatrix}
# \label{eq:aeqn}
# \end{equation}
kernel5= np.array([[1,1,1],[1,1,1],[1,1,1]])*1/9
DisplayImagesBeforeAfterConvolution(img, kernel5)
# ### 6- Random Karnel
kernel6= np.array([[-4,5,-6],[-2,-3,14],[-6,1,-8]])*1/9
DisplayImagesBeforeAfterConvolution(img, kernel)
| 1-Kernels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
'''snippet to filter out the hits coming from intergenic or other regions that
cerebra shouldnt be finding hits in '''
import pandas as pd
# +
readpath = '../funco_bench_basic/hap_map_bench_ensp.vcf'
writepath = '../funco_bench_basic/hap_map_bench_ensp_clean.vcf'
keep_lines = 0
not_keep_lines = 0
deplorables = ['|IGR|', '|FIVE_PRIME_UTR|', '|THREE_PRIME_UTR|', '|FIVE_PRIME_FLANK|', '|THREE_PRIME_FLANK|',
'|COULD_NOT_DETERMINE|', '|SILENT|'] # does cerebra keep silent?
with open(readpath, 'r') as rp:
with open(writepath, 'w') as wp:
lines = rp.readlines()
for line in lines:
if any(ele in line for ele in deplorables):
not_keep_lines += 1
else:
keep_lines += 1
wp.write(line)
#wp.write("\n") # not sure this is needed
wp.close()
rp.close()
# -
print(not_keep_lines)
print(keep_lines)
| python_notebooks/get_non_UTR_lines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#download data of bitmap images
# !wget https://storage.googleapis.com/quickdraw_dataset/full/numpy_bitmap/hand.npy
# +
#import modules
import numpy as np
from tqdm import tqdm, trange
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
# import tensorflow as tf
import pickle as pkl
import time
# -
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
data = np.load('hand.npy') #input is the file location stored as npy
data.shape
# +
#plot sample images
fig=plt.figure(figsize=(10, 10))
columns = 15
rows = 10
for i in range(1, columns*rows):
img = data[i+44].reshape((28,28))
fig.add_subplot(rows, columns, i)
plt.imshow(img)
plt.show()
# -
def model_inputs(real_dims, z_dims):
'''
real_dims : df dimensions
z_dims : placeholder dimensions
return input dims, input placeholder dims
'''
inputs_real = tf.placeholder(tf.float32, shape=(None, real_dims), name='input_real')
inputs_z = tf.placeholder(tf.float32, shape=(None, z_dims), name='input_z')
return inputs_real, inputs_z
def get_data(path):
try:
data = np.load(path)
Y = []
for i in trange(data.shape[0]):
Y.append([1,0])
Y = np.array(Y)
(x_train, y_train, x_test, y_test) = train_test_split(data, Y)
x_train = (x_train.astype(np.float32)) / 255
x_train = x_train.reshape(x_train.shape[0], 784)
return (x_train, y_train, x_test, y_test)
except Exception as e:
print (e)
x_train, y_train, x_test, y_test = get_data('hand.npy')
x_train.shape
def generator(z, out_dims, n_units=128, reuse=False, alpha=0.01):
with tf.variable_scope('generator', reuse=reuse):
#hidden layer
h1 = tf.layers.dense(z,n_units, activation=None,)
#leaky relu implementation
h1 = tf.maximum(alpha*h1, h1)
#tanh
logits = tf.layers.dense(h1, out_dims)
out = tf.tanh(logits)
return out
def discriminator(x, n_units=128, reuse=False, alpha=0.01):
with tf.variable_scope('discriminator', reuse=reuse):
#hidden layer
h1 = tf.layers.dense(x, n_units, activation=None)
#leaky_relu
h1 = tf.maximum(alpha*h1, h1)
#sigmoid
logits = tf.layers.dense(h1, 1, activation=None)
out = tf.sigmoid(logits)
return out, logits
# +
#hyperparams
r_size = 784
z_size = 100
g_units = 128
d_units = 128
alpha = 0.01
smooth = 0.1
# +
from tensorflow.python.framework import ops
ops.reset_default_graph()
inputs_real, inputs_z = model_inputs(r_size, z_size)
# -
g_out = generator(inputs_z, r_size, g_units)
# +
d_out_real, real_logit = discriminator(inputs_real,)
d_out_fake, fake_logits = discriminator(g_out, reuse=True)
# -
d_loss_real = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=real_logit, labels=tf.ones_like(real_logit)*(1-smooth)))
d_loss_fake = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=fake_logits, labels=tf.zeros_like(fake_logits)))
d_loss = d_loss_fake+d_loss_real
g_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=fake_logits, labels=tf.ones_like(fake_logits)))
lr = 0.002
tvar = tf.trainable_variables()
gvar = [var for var in tvar if var.name.startswith('generator')]
dvar = [var for var in tvar if var.name.startswith('discriminator')]
d_opt = tf.train.AdamOptimizer(learning_rate=lr).minimize(d_loss, var_list=dvar)
g_opt = tf.train.AdamOptimizer(lr).minimize(g_loss,var_list=gvar)
# +
start = time.time()
batch_size = 100000
epochs = 100
samples = []
losses = []
# Only save generator variables
saver = tf.train.Saver(var_list=gvar)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for e in range(epochs):
for ii in range(x_train.shape[0]//batch_size):
batch = x_train[np.random.randint(0, x_train.shape[0], size=batch_size)]
batch_images = batch*2 - 1
# Sample random noise for G
batch_z = np.random.uniform(-1, 1, size=(batch_size, z_size))
# Run optimizers
_ = sess.run(d_opt, feed_dict={inputs_real: batch_images, inputs_z: batch_z})
_ = sess.run(g_opt, feed_dict={inputs_z: batch_z})
# At the end of each epoch, get the losses and print them out
train_loss_d = sess.run(d_loss, {inputs_z: batch_z, inputs_real: batch_images})
train_loss_g = g_loss.eval({inputs_z: batch_z})
if e%10 == 0:
print("Epoch {}/{}...".format(e, epochs),
"Discriminator Loss: {:.4f}...".format(train_loss_d),
"Generator Loss: {:.4f}".format(train_loss_g))
# Save losses to view after training
losses.append((train_loss_d, train_loss_g))
# Sample from generator as we're training for viewing afterwards
sample_z = np.random.uniform(-1, 1, size=(16, z_size))
gen_samples = sess.run(
generator(inputs_z, r_size, n_units=g_units, reuse=True, alpha=alpha),
feed_dict={inputs_z: sample_z})
samples.append(gen_samples)
saver.save(sess, './checkpoints/generator.ckpt')
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
end = time.time()
print('Time elapsed : ', (end-start)/60, ' mins.')
# -
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator')
plt.plot(losses.T[1], label='Generator')
plt.title("Training Losses")
plt.legend()
plt.show()
def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(5,5), nrows=4, ncols=4, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((28,28)), cmap='Greys')
return fig, axes
# Load samples from generator taken while training
with open('train_samples.pkl', 'rb') as f:
samples = pkl.load(f)
_ = view_samples(80, samples)
# +
rows, cols = 10, 10
fig, axes = plt.subplots(figsize=(8,6), nrows=rows, ncols=cols, sharex=True, sharey=True)
for sample, ax_row in zip(samples[::int(len(samples)/rows)], axes):
for img, ax in zip(sample[::int(len(sample)/cols)], ax_row):
ax.imshow(img.reshape((28,28)), cmap='Greys')
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
# -
| notebooks/dooDleGan-Cloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Which Machine Learning Models Are Sensitive To Outliers?
# * Naivye Bayes Classifier ------ Not Sensitive To Outliers
# * SVM -------------------------- Not Sensitive To Outliers
# * Linear Regression------------- Sensitive To Outliers
# * Logistic Regression----------- Sensitive To Outliers
# * DecisionTree ----------------- Not Sensitive
# * Ensemble(RF,XGboost,GB)--- Not Sensitive
# * KNN--------------------------- Not Sensitive
# * Kmeans------------------------ Sensitive
# * Hierarichal--------------------- Sensitive
# * PCA--------------------------- Sensitive
# * Neural Networks------------- Sensitive
import pandas as pd
df = pd.read_csv('titanic.csv')
df.head()
df.Age.isnull().sum()
df.Age = df.Age.dropna()
df.Age.hist(bins=20)
df.Fare.hist(bins=20)
df.Age.describe()
# ## Gaussian Distributed
figure=df.Age.hist(bins=20)
figure.set_title('Age')
figure.set_xlabel('Age')
figure.set_ylabel('No of passenger')
import seaborn as sns
sns.boxplot(df['Age'])
# ### If the data is normally distributed use this method:
Lower_boundaary = df.Age.mean() - 3 * df.Age.std()
Upper_boundaary = df.Age.mean() + 3 * df.Age.std()
print(f"Upper Boundaary: {Upper_boundaary}")
print(f"Lower Boundaary: {Lower_boundaary}")
print(f"Mean of Age: {df.Age.mean()}")
# #### Replace outliers with upper boundary limit
df.loc[df['Age']>=73, 'Age']=73
df.Age.hist(bins=50)
# ## Skewed Distribution:
sns.distplot(df.Fare)
sns.boxplot(df.Fare)
df.Fare.describe()
# ### If the data is skewed distribution use this method
# Find IQR value
IQR = df.Fare.quantile(0.75)-df.Fare.quantile(0.25)
print(f"InterQuartile Range : {IQR}")
# Apply IQR value in the Formula
Lower_boundaary = df.Fare.quantile(0.25) - (1.5*IQR)
Upper_boundaary = df.Fare.quantile(0.75) + (1.5*IQR)
print(f"Upper Boundaary: {Upper_boundaary}")
print(f"Lower Boundaary: {Lower_boundaary}")
# #### Find Extreme Outlier
# Instead 1.5 value use 3 to find the extreme outliers
Lower_boundaary = df.Fare.quantile(0.25) - (3*IQR)
Upper_boundaary = df.Fare.quantile(0.75) + (3*IQR)
print(f"Upper Boundaary: {Upper_boundaary}")
print(f"Lower Boundaary: {Lower_boundaary}")
# #### Replace Outliers with Upper Boundary value
df.loc[df['Fare']>=100, 'Fare']=100
df.Fare.hist(bins=20)
# #### Splitting dataset for train_test_split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df[['Age', 'Fare']].fillna(0), df['Survived'], test_size=0.2, random_state=0)
# #### Using LogisticRegressor
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
y_pred_prob = classifier.predict_proba(X_test)
from sklearn.metrics import accuracy_score, roc_auc_score
print(f'Accuracy Score : {accuracy_score(y_test, y_pred)}')
print(f'Roc Accuracy Score : {roc_auc_score(y_test, y_pred_prob[:,1])}')
# #### Using Ensemble Techniques
from sklearn.ensemble import RandomForestClassifier
RFC = RandomForestClassifier(n_estimators=200,random_state=0)
RFC.fit(X_train, y_train)
y_pred = RFC.predict(X_test)
y_pred_prob = RFC.predict_proba(X_test)
print(f'Accuracy Score : {accuracy_score(y_test, y_pred)}')
print(f'Roc Accuracy Score : {roc_auc_score(y_test, y_pred_prob[:,1])}')
| Outliers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="VU4uue9rpPM-"
# ## Análise Exploratória
# + executionInfo={"elapsed": 970, "status": "ok", "timestamp": 1608010105240, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="3bjBybE__WrW"
import pandas as pd
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
sns.set_style('darkgrid')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 846, "status": "ok", "timestamp": 1608010107360, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="-1ALvVQiFCGE" outputId="b157b660-5849-4f35-f3b9-107e847f3206"
#df = pd.read_csv('/content/drive/MyDrive/Estudo data science/Machine learning/Aula Diego/Trabalho 1/Dados/Bruto/merged_2015.csv')
df = pd.read_csv('../../dados/brutos/merged_2015.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 333} executionInfo={"elapsed": 4055, "status": "ok", "timestamp": 1607996837073, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="XKFLz8N9Fbxl" outputId="4cf5c344-473d-41c9-e12e-d2386d3804f5"
df.describe()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 860, "status": "ok", "timestamp": 1607996840093, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="9VoVsPmzFpSX" outputId="3a0db69d-62e0-44b7-a5e5-781f5270c354"
df.isnull().sum()[df.isnull().sum() > 0].sort_values(ascending=False)
# -
# ## Distribuição das dependências administrativas
# + colab={"base_uri": "https://localhost:8080/", "height": 292} executionInfo={"elapsed": 950, "status": "ok", "timestamp": 1608000849363, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="AaGIRoyDs7n6" outputId="acd3b265-bc10-4fc1-d897-c0640b098044"
aux = df.copy()
tipo_map = {
1: 'Federal',
2: 'Estadual',
3: 'Municipal',
4: 'Privada'
}
aux['TP_DEPENDENCIA'] = aux['TP_DEPENDENCIA'].map(tipo_map)
ordem = [tipo_map[2], tipo_map[4], tipo_map[1], tipo_map[3]]
# plt.figure(figsize=(10,8))
sns.set(font_scale=1.2)
plt.tight_layout()
escolas_plt = sns.countplot(x='TP_DEPENDENCIA', data=aux, order=ordem, palette='YlGnBu_r')
escolas_plt.set(xlabel='Dependência administrativa', ylabel='Quantidade de escolas')
plt.savefig('../../graficos/analise/countplot_dist_dep.pdf')
# + colab={"base_uri": "https://localhost:8080/", "height": 250} executionInfo={"elapsed": 972, "status": "ok", "timestamp": 1607996849870, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="nifaeOvLd42F" outputId="b2ae6b1b-8044-4976-dfc4-ac0d1f6ce524"
cores = ['#225ea8', '#41b6c4', '#a1dab4', '#ffffcc']
plt.tight_layout()
sns.set(font_scale=1)
plt.pie(df['TP_DEPENDENCIA'].value_counts(), colors=cores,
autopct='%1.1f%%', pctdistance=1.2)
plt.legend(['Estadual', 'Privada', 'Federal', 'Municipal'], title='Dependência',
loc='lower right', bbox_to_anchor=(1, 0, 0.5, 1))
plt.subplots_adjust(left=0.0)
plt.savefig('../../graficos/analise/pizza_dist_dep.pdf')
# -
# ## Distribuição das notas do Enem
# - CN: Ciências da Natureza e suas Tecnologias
# - CH: Ciências Humanas e suas Tecnologias
# - LP: Linguagens, Códigos e suas Tecnologias
# - MT: Matemática e suas Tecnologias
# - RED: Redação
# + colab={"base_uri": "https://localhost:8080/", "height": 285} executionInfo={"elapsed": 1852, "status": "ok", "timestamp": 1607996858636, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="Kzn4gd2BfITX" outputId="68434e97-f725-4562-ea5d-b7890d4fb2bc"
fig = sns.violinplot(x='value', y='variable',
data=df[['NU_MEDIA_CN', 'NU_MEDIA_CH',
'NU_MEDIA_LP', 'NU_MEDIA_MT',
'NU_MEDIA_RED']].rename(columns={
'NU_MEDIA_CN':'CN',
'NU_MEDIA_CH':'CH',
'NU_MEDIA_LP':'LP',
'NU_MEDIA_MT':'MT',
'NU_MEDIA_RED':'RED'
}).melt(),
palette='YlGnBu_r')
sns.set(font_scale=1.2)
plt.tight_layout()
fig.set(xlabel='Nota média', ylabel='Prova')
plt.savefig('../../graficos/analise/violinplot_dist_notas.pdf')
# -
# Desvio padrão para cada matéria.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 933, "status": "ok", "timestamp": 1607996881946, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="dSUeC91gg8uX" outputId="21c0d19a-ff02-4a23-b4ae-8a840a6de6e5"
df[['NU_MEDIA_CN', 'NU_MEDIA_CH','NU_MEDIA_LP', 'NU_MEDIA_MT','NU_MEDIA_RED']].std()
# -
# ## Distribuição das notas de redação
# + colab={"base_uri": "https://localhost:8080/", "height": 382} executionInfo={"elapsed": 1756, "status": "ok", "timestamp": 1608005803605, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="lBv_QqBGdxBV" outputId="b0424510-8a05-4f26-da5e-d63ff0820c9f"
sns.set(font_scale=1)
plt.tight_layout()
fig = sns.displot(data=df, x='NU_MEDIA_RED', kde=True)
fig.set(xlabel='Nota média de redação', ylabel='Quantidade')
plt.savefig('../../graficos/analise/dist_red.pdf')
# -
# ## Correlação e mapa de calor para a nota de redação
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2457, "status": "ok", "timestamp": 1607996893865, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="Q1G3AOr-dytq" outputId="a89d0d4c-dcac-46cf-c012-5c40e755eb8d"
corr_cols = df.drop(['NU_MEDIA_MT',
'NU_MEDIA_LP',
'NU_MEDIA_CH',
'NU_MEDIA_CN',
'TP_DEPENDENCIA',
'TP_DEPENDENCIA_ADM_ESCOLA',
'CO_LINGUA_INDIGENA'], axis=1).corr()['NU_MEDIA_RED'].abs().sort_values(ascending = False)[:10].index
corr = df[corr_cols].corr()
corr['NU_MEDIA_RED']
# + colab={"base_uri": "https://localhost:8080/", "height": 285} executionInfo={"elapsed": 3346, "status": "ok", "timestamp": 1607996894764, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="dVhr-y5Xd2qU" outputId="cbb5d87c-307b-4718-cc4b-138ec46b278b"
sns.set(font_scale=0.8)
mask = np.zeros_like(corr)
mask[np.triu_indices_from(mask)] = True
sns.heatmap(corr, cmap='YlGnBu', mask=mask, linewidths=.5, annot=True, fmt=".2f")
sns.set(font_scale=1.3)
plt.tight_layout()
plt.savefig('../../graficos/analise/heatmap.pdf')
# -
# Gráficos da nota de redação com as variáveis contínuas de maior correlação:
# + colab={"base_uri": "https://localhost:8080/", "height": 291} executionInfo={"elapsed": 2183, "status": "ok", "timestamp": 1607998739555, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="MvYyUg4VFhNx" outputId="7b2a0ec2-22fa-42bb-9c91-f07b26c20df4"
sns.set(font_scale=1.2)
plt.tight_layout()
fig = sns.scatterplot(x='NU_TAXA_PARTICIPACAO', y='NU_MEDIA_RED', data=df, alpha=0.5)
fig.set(xlabel='Taxa de participação na prova', ylabel='Nota média de redação')
plt.savefig('../../graficos/analise/scatter_particip_red.pdf')
# + colab={"base_uri": "https://localhost:8080/", "height": 291} executionInfo={"elapsed": 1893, "status": "ok", "timestamp": 1607998751443, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="B7GYCOjyGcmw" outputId="4cded1c7-2856-4b26-c6ef-044c3da00456"
sns.set(font_scale=1.2)
plt.tight_layout()
fig = sns.scatterplot(x='NU_TAXA_ABANDONO', y='NU_MEDIA_RED', data=df, alpha=0.5)
fig.set(xlabel='Taxa de abandono da escola', ylabel='Nota média de redação')
plt.savefig('../../graficos/analise/scatter_abandon_red.pdf')
# -
# ## Distribuição da nota média de redação em categorias
# ### Dependência administrativa
# + colab={"base_uri": "https://localhost:8080/", "height": 289} executionInfo={"elapsed": 1223, "status": "ok", "timestamp": 1608001129233, "user": {"displayName": "Jo\u0<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="XWuaHYLKyefC" outputId="b53eb635-2e26-422a-cba3-23480f591d16"
aux = df.copy()
tipo_map = {
1: 'Federal',
2: 'Estadual',
3: 'Municipal',
4: 'Privada'
}
aux.loc[:,'TP_DEPENDENCIA'] = aux['TP_DEPENDENCIA'].map(tipo_map)
ordem = [tipo_map[2], tipo_map[4], tipo_map[1], tipo_map[3]]
# plt.figure(figsize=(10,8))
sns.set(font_scale=1.2)
plt.tight_layout()
fig = sns.boxplot(x='TP_DEPENDENCIA', y='NU_MEDIA_RED', data=aux, order=ordem, palette='YlGnBu_r')
fig.set(xlabel='Dependência administrativa', ylabel='Nota média de redação')
plt.savefig('../../graficos/analise/boxplot_dep_red.pdf')
# -
# ### Região
# + colab={"base_uri": "https://localhost:8080/", "height": 289} executionInfo={"elapsed": 1129, "status": "ok", "timestamp": 1608011940840, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="BbAwgOPLJtYz" outputId="21aafbb0-7190-45a2-e922-a45c79c80e51"
aux = df.copy()
regiao_map = {
1: 'Norte',
2: 'Nordeste',
3: 'Sudeste',
4: 'Sul',
5: 'Centro-Oeste'
}
aux.loc[:,'CO_REGIAO'] = aux['CO_REGIAO'].map(regiao_map)
sns.set(font_scale=1.2)
plt.tight_layout()
fig = sns.boxplot(data=aux, x='CO_REGIAO', y='NU_MEDIA_RED', palette='YlGnBu_r')
fig.set(xlabel='Região', ylabel='Nota média de redação')
plt.savefig('../../graficos/analise/boxplot_regiao_red.pdf')
# -
# ### Alimentação
# + colab={"base_uri": "https://localhost:8080/", "height": 365} executionInfo={"elapsed": 3258, "status": "ok", "timestamp": 1608012000828, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="svecCwngjTrx" outputId="26f92f78-03c4-4eaf-a505-b3617c68685b"
aux = df.copy()
alim_map = {
0: 'Não ({})'.format(df['IN_ALIMENTACAO'].value_counts()[0]),
1: 'Sim ({})'.format(df['IN_ALIMENTACAO'].value_counts()[1])
}
aux.loc[:,'IN_ALIMENTACAO'] = aux['IN_ALIMENTACAO'].map(alim_map)
sns.set(font_scale=1)
sns.set_style('darkgrid')
fig = sns.catplot(x='IN_ALIMENTACAO', y='NU_MEDIA_RED', data=aux, alpha=0.2, palette='Dark2')
fig.set(xlabel='Fornecimento de alimentação', ylabel='Nota média de redação')
fig.tight_layout()
plt.savefig('../../graficos/analise/catplot_alim_red.pdf')
# -
# ### Água
# + colab={"base_uri": "https://localhost:8080/", "height": 382} executionInfo={"elapsed": 3165, "status": "ok", "timestamp": 1608012032415, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="A-zhtSNHmsR2" outputId="8de36a15-df83-46b3-e704-ba2acd44f2b8"
aux = df.copy()
agua_map = {
0: 'Não ({})'.format(df['IN_AGUA_FILTRADA'].value_counts()[0]),
1: 'Sim ({})'.format(df['IN_AGUA_FILTRADA'].value_counts()[1])
}
aux.loc[:,'IN_AGUA_FILTRADA'] = aux['IN_AGUA_FILTRADA'].map(agua_map)
ordem = [agua_map[0], agua_map[1]]
# sns.set(font_scale=1)
plt.tight_layout()
fig = sns.catplot(x='IN_AGUA_FILTRADA', y='NU_MEDIA_RED', data=aux, alpha=0.2, order=ordem, palette='Dark2')
fig.set(xlabel='Fornecimento de água filtrada', ylabel='Nota média de redação')
fig.tight_layout()
plt.savefig('../../graficos/analise/catplot_agua_red.pdf')
# + colab={"base_uri": "https://localhost:8080/", "height": 410} executionInfo={"elapsed": 8290, "status": "ok", "timestamp": 1607996899766, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="I0OlmbteuY80" outputId="d7d54dd6-5402-442d-afcf-13225d95a54f"
aux = df.copy()
agua_map = {
1: 'Não ({})'.format(df['IN_AGUA_INEXISTENTE'].value_counts()[1]),
0: 'Sim ({})'.format(df['IN_AGUA_INEXISTENTE'].value_counts()[0])
}
aux.loc[:,'IN_AGUA_INEXISTENTE'] = aux['IN_AGUA_INEXISTENTE'].map(agua_map)
ordem = [agua_map[1], agua_map[0]]
sns.set(font_scale=1.3)
plt.tight_layout()
fig = sns.catplot(x='IN_AGUA_INEXISTENTE', y='NU_MEDIA_RED', data=aux, alpha=0.2, order=ordem, palette='Dark2')
fig.set(xlabel='Fornecimento de água', ylabel='Nota média de redação')
# -
# ### Laboratório de ciências
# + colab={"base_uri": "https://localhost:8080/", "height": 400} executionInfo={"elapsed": 3070, "status": "ok", "timestamp": 1608010209469, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="3__YFWJwnOtc" outputId="e88d0c48-c8d1-4e5b-e47c-ffaf808577e5"
aux = df.copy()
ciencia_map = {
0: 'Não ({})'.format(df['IN_LABORATORIO_CIENCIAS'].value_counts()[0]),
1: 'Sim ({})'.format(df['IN_LABORATORIO_CIENCIAS'].value_counts()[1])
}
aux.loc[:,'IN_LABORATORIO_CIENCIAS'] = aux['IN_LABORATORIO_CIENCIAS'].map(ciencia_map)
ordem = [ciencia_map[0], ciencia_map[1]]
# sns.set(font_scale=1)
plt.tight_layout()
fig = sns.catplot(x='IN_LABORATORIO_CIENCIAS', y='NU_MEDIA_RED', data=aux, alpha=0.2, order=ordem, palette='Dark2')
fig.set(xlabel='Existência de laboratório de ciências', ylabel='Nota média de redação')
plt.savefig('../../graficos/analise/catplot_ciencia_red.pdf')
# -
# ### Esgoto
# + colab={"base_uri": "https://localhost:8080/", "height": 382} executionInfo={"elapsed": 3453, "status": "ok", "timestamp": 1608012044070, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="K0SbKkk27wtu" outputId="b371ae4a-87e4-4048-94ac-ae2aa549ba50"
aux = df.copy()
esgoto_map = {
0: 'Não ({})'.format(df['IN_ESGOTO_REDE_PUBLICA'].value_counts()[0]),
1: 'Sim ({})'.format(df['IN_ESGOTO_REDE_PUBLICA'].value_counts()[1])
}
aux.loc[:,'IN_ESGOTO_REDE_PUBLICA'] = aux['IN_ESGOTO_REDE_PUBLICA'].map(esgoto_map)
ordem = [esgoto_map[0], esgoto_map[1]]
# sns.set(font_scale=1)
plt.tight_layout()
fig = sns.catplot(x='IN_ESGOTO_REDE_PUBLICA', y='NU_MEDIA_RED', data=aux, alpha=0.2, order=ordem, palette='Dark2')
fig.set(xlabel='Esgoto tratado pela rede pública', ylabel='Nota média de redação')
fig.tight_layout()
plt.savefig('../../graficos/analise/catplot_esgoto_red.pdf')
# + colab={"base_uri": "https://localhost:8080/", "height": 416} executionInfo={"elapsed": 1669, "status": "ok", "timestamp": 1608007382589, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjRIgV0DhdG_-UgNITfYFIhbxJ51p017AeuwJQ3=s64", "userId": "01802867954268273239"}, "user_tz": 180} id="giUdkZTw8AjE" outputId="5e4f5820-e356-48db-a086-9d61e50017ff"
aux = df.copy()
esgoto_map = {
1: 'Não ({})'.format(df['IN_ESGOTO_INEXISTENTE'].value_counts()[1]),
0: 'Sim ({})'.format(df['IN_ESGOTO_INEXISTENTE'].value_counts()[0])
}
aux.loc[:,'IN_ESGOTO_INEXISTENTE'] = aux['IN_ESGOTO_INEXISTENTE'].map(esgoto_map)
ordem = [esgoto_map[1], esgoto_map[0]]
sns.set(font_scale=1)
plt.tight_layout()
fig = sns.catplot(x='IN_ESGOTO_INEXISTENTE', y='NU_MEDIA_RED', data=aux, alpha=0.2, order=ordem, palette='Dark2')
fig.set(xlabel='Tratamento inexistente de esgoto', ylabel='Nota média de redação')
| notebooks/dados/analise_exploratoria.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ogvKVnCm0J6u"
import pandas as pd
# + id="TrfWNjiw5D4C"
import matplotlib.pyplot as plt
# + id="hWQ4BEqr5k8C"
df=pd.read_csv("HRDataset_v14.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 427} id="2msEvokl5uXL" outputId="5c18a1f4-7d1c-4ebf-f500-b66c1fa1b6d7"
df.head()
# + id="yx9AX_Hb5vgK"
# Determine the total number of people who are Sales Executive
# + colab={"base_uri": "https://localhost:8080/"} id="-Sof2Ef39L2R" outputId="e8812d8a-716b-4db3-8ec6-407f8a9668c2"
df.loc[df['Position'] == "Production Technician I"].count()
# + id="MBB8brZqBiPp"
#Determine the number of males.
# + colab={"base_uri": "https://localhost:8080/"} id="wXNUBrac9yjh" outputId="874d8718-7cc8-4c86-826e-f93aa040b9d5"
df.loc[df['Gender'] == "Male"].count()
# + colab={"base_uri": "https://localhost:8080/"} id="ZNNz2IrUBexD" outputId="afd85ad3-dad8-4a1a-d740-e357197ebc6c"
#Find persons who are employed under “<NAME>”.
df.loc[df['ManagerName'] == "<NAME>"].count()
# + colab={"base_uri": "https://localhost:8080/"} id="o0FSBNlYEw8P" outputId="22330447-e887-4063-e524-a66e9256e7d6"
#List active employee names.
df['Employee_Name'].loc[df['EmploymentStatus'] == "Active"]
# + colab={"base_uri": "https://localhost:8080/", "height": 748} id="x18lDHi-Fuxr" outputId="3aa284b7-9019-45fb-c99a-473da0bf354c"
#List active employee names who are married
df.loc[(df['EmploymentStatus'] == "Active" ) & (df['MaritalDesc'] == "Married") ]
# + colab={"base_uri": "https://localhost:8080/"} id="EvpoOJwmGB9-" outputId="47d0925f-1225-4ce3-98e3-cbd76eb1452b"
#Find a US citizen earning less than $60,000.
df['Salary'].loc[(df['EmploymentStatus'] == "Active" ) & (df['CitizenDesc'] == "US Citizen") ]>6000
# + colab={"base_uri": "https://localhost:8080/"} id="U36WdaL8JYB0" outputId="e57ab049-a86c-4506-90a7-bf625e81fe02"
#Find a active employees earning less than $60,000.
for k in df.index :
if (df.loc[k,'EmploymentStatus'] == "Active" ) & (df.loc[k,'Salary']>6000):
print(df.loc[k,'Employee_Name'])
# + colab={"base_uri": "https://localhost:8080/"} id="YZICSvpoOnjN" outputId="9885908a-d467-4147-992f-633822c52157"
#Find the best Production Technician II based on EngagementSurvey
df['EngagementSurvey'].max()
# + colab={"base_uri": "https://localhost:8080/"} id="Ri0ZmsKEKZOD" outputId="c675b238-99c7-4d20-c470-04a1fb844026"
#Find the best Production Technician II based on EngagementSurvey
for k in df.index :
if (df.loc[k,'Position'] == "Production Technician I" ) & (df.loc[k,'EngagementSurvey']== df['EngagementSurvey'].max()) :
print(df.loc[k,'Employee_Name'])
# + colab={"base_uri": "https://localhost:8080/"} id="i24S6gdgNOWE" outputId="b6d28927-c0fb-4e35-9dc5-dabaf0ad31b7"
#Find the positions which are drawing maximum Salary
for k in df.index :
if (df.loc[k,'Salary']== df['Salary'].max()) :
print(df.loc[k,'Position'])
# + colab={"base_uri": "https://localhost:8080/"} id="8nk0xZxnPPGI" outputId="368e62b9-f3a8-4d73-aae7-5f605e72e62c"
#Find the positions which are drawing lower than average salary
for k in df.index :
if (df.loc[k,'Salary'] < df['Salary'].mean()) :
print(df.loc[k,'Position'])
# + colab={"base_uri": "https://localhost:8080/"} id="GsMyy2CMPXha" outputId="2a98ca5f-10b8-42f5-a017-6733d123adb6"
#Find the persons which are drawing lower than average salary
for k in df.index :
if (df.loc[k,'Salary'] < df['Salary'].mean()) :
print(df.loc[k,'Employee_Name'])
# + colab={"base_uri": "https://localhost:8080/"} id="Z1DtvO5cPgN0" outputId="86201ccb-d873-4691-ce48-7d066364930d"
#Find the persons which are drawing lower than average salary and are not satisfactory
for k in df.index :
if (df.loc[k,'Salary'] < df['Salary'].mean()) & (df.loc[k,'JobSatisfaction'] < df['JobSatisfaction'].mean()) :
print(df.loc[k,'Employee_Name'])
# + id="UX5iaoMQRBjp"
for k in df.index :
if (df.loc[k,'Salary'] == df['Salary'].min()) & (df.loc[k,'JobSatisfaction'] == df['JobSatisfaction'].max()) :
print(df.loc[k,'Employee_Name'])
# + colab={"base_uri": "https://localhost:8080/"} id="PvwwDIeYRJTY" outputId="74119084-dd12-44a4-9a7f-66506cd9cb8a"
for k in df.index :
if (df.loc[k,'Salary'] == df['Salary'].min()) & (df.loc[k,'JobSatisfaction'] >= df['JobSatisfaction'].min()) :
print(df.loc[k,'Employee_Name'])
# + colab={"base_uri": "https://localhost:8080/"} id="8XwxglqMRbdR" outputId="3072cf9c-85e7-442e-a65e-695652b09d5d"
for k in df.index :
if (df.loc[k,'Salary'] < df['Salary'].mean()) & (df.loc[k,'JobSatisfaction'] >= 2) :
print(df.loc[k,'Employee_Name'])
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="NxR9P-k7SDRe" outputId="5860df75-c3f9-448c-9424-1889c3b665f2"
# Fine the relation between DistanceFromHome and Salary
plt.scatter(df['DistanceFromHome'], df['Salary'])
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="S2klFdNAT_e1" outputId="698a1d50-cbb8-4d3c-9a64-d840f4438533"
# Find the relation between DistanceFromHome and employee satisfaction
plt.scatter(df['MonthlyIncome'], df['JobSatisfaction'])
# + colab={"base_uri": "https://localhost:8080/", "height": 748} id="rKaHOsrzVYV2" outputId="a0e35b0c-d53e-4078-fe28-139e7946df91"
df1=df.sort_values(by=['EmpID'], ascending=False)
df1
# + colab={"base_uri": "https://localhost:8080/"} id="8loMrDmIa547" outputId="22e6c9ee-70ff-4e5a-e60d-46e30114384a"
print( df1['PerformanceRating'])
# + id="uZlLLI4-d95o"
| Data Analytics/02.HRA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Calibrating the flats
#
# Recall that the counts in an astronomical image include dark current, noise, and
# a near-constant offset, the bias. The individual flat frames need to have bias
# and dark removed from them. Depending on the exposure times of the images you
# have, you may or may not need to subtract dark and bias separately.
#
# If the combined dark frame needs to be scaled to a different exposure time, then
# bias and dark must be handled separately; otherwise, the dark and bias can be
# removed in a single step because dark frames also include bias.
#
# The potential reduction steps for flat frames are below:
#
# + Subtract overscan and trim, if necessary.
# + Subtract bias, if necessary.
# + Subtract dark current, scaling if necessary (scale down when possible).
#
# As in the chapters about bias and dark we will work through two examples. In
# Example 1 the darks are not scaled and the overscan region is used as part of
# the calibration. In Example 2 the darks are scaled and the overscan region is
# trimmed off without being used.
# ## Function definition
#
# The function below finds the nearest dark exposure time to the exposure time of
# a given image. An exception is raised if the difference in exposure time is
# larger than `tolerance`, unless `tolerance` is set to `None`. A small numerical
# tolerance is most useful if you anticipate not scaling the dark frames and
# finding a dark exposure time close to the time of the image. Disregarding the
# tolerance is useful if the intent is to scale the dark frames anyway.
def find_nearest_dark_exposure(image, dark_exposure_times, tolerance=0.5):
"""
Find the nearest exposure time of a dark frame to the exposure time of the image,
raising an error if the difference in exposure time is more than tolerance.
Parameters
----------
image : astropy.nddata.CCDData
Image for which a matching dark is needed.
dark_exposure_times : list
Exposure times for which there are darks.
tolerance : float or ``None``, optional
Maximum difference, in seconds, between the image and the closest dark. Set
to ``None`` to skip the tolerance test.
Returns
-------
float
Closest dark exposure time to the image.
"""
dark_exposures = np.array(list(dark_exposure_times))
idx = np.argmin(np.abs(dark_exposures - image.header['exptime']))
closest_dark_exposure = dark_exposures[idx]
if (tolerance is not None and
np.abs(image.header['exptime'] - closest_dark_exposure) > tolerance):
raise RuntimeError('Closest dark exposure time is {} for flat of exposure '
'time {}.'.format(closest_dark_exposure, a_flat.header['exptime']))
return closest_dark_exposure
# +
from pathlib import Path
from astropy import units as u
from astropy.nddata import CCDData
import ccdproc as ccdp
from matplotlib import pyplot as plt
import numpy as np
from convenience_functions import show_image
# -
# Use custom style for larger fonts and figures
plt.style.use('guide.mplstyle')
# ## Example 1: No scaling of dark frames
#
# The images for this example are from chip 0 of the Large Format Camera at
# Palomar Observatory. The raw images are [on Zenodo](https://doi.org/10.5281/zenodo.3254683), and this notebook assumes that
# you have worked through the notebooks on bias and dark so that there is a folder
# called `example1-reduced` in the same folder as this notebook.
#
# We'll go through this example twice: once with a single image to explain each
# step, and then again to process all of the flat frames in the directory of raw data.
#
# An image collection is defined below, along with a couple of settings useful for
# this example.
# +
reduced_path = Path('example1-reduced')
ifc_reduced = ccdp.ImageFileCollection(reduced_path)
combined_dark_files = ifc_reduced.files_filtered(imagetyp='dark', combined=True)
flat_image_type = 'FLATFIELD'
# -
# The raw data should be in the directory `example-cryo-LFC`.
# +
raw_data = Path('example-cryo-LFC')
ifc_raw = ccdp.ImageFileCollection(raw_data)
# -
# The cell below checks that the files needed are available.
# +
n_combined_dark = len(combined_dark_files)
expected_exposure_times = set([7, 70, 300])
if n_combined_dark < 3:
raise RuntimeError('One or more combined dark is missing. Please re-run the dark notebook.')
elif n_combined_dark > 3:
raise RuntimeError('There are more combined dark frames than expected.')
actual_exposure_times = set(h['exptime'] for h in ifc_reduced.headers(imagetyp='dark', combined=True))
if (expected_exposure_times - actual_exposure_times):
raise RuntimeError('Encountered unexpected exposure time in combined darks. '
'The unexpected times are {}'.format(actual_exposure_times - expected_exposure_times))
# -
# First, get one of the flat frames as a `CCDData` object and display it.
# +
a_flat = CCDData.read(ifc_raw.files_filtered(imagetyp='flatfield', include_path=True)[0], unit='adu')
show_image(a_flat, cmap='gray')
# -
# There is not a lot of variation in this. Note that the overscan region on the
# right stands out as a black bar, and there is apparently also an overscan
# region across the top of the chip. There appears to be a slight variation in
# pixel values from the bottom to the top of the image.
# ### Subtract overscan and trim, if necessary
#
# The overscan is useful for the LFC and needs to be subtracted and trimmed off.
# See [this example in the dark reduction notebook](03-05-Calibrate-dark-images.ipynb#Decide-which-calibration--steps-to-take) for a review of the overscan parameters.
# The overscan region is the Python slice `[:, 2055:]` while the region to be
# retained after trimming is the Python slice `[:, :2048]`.
# +
# Subtract the overscan
a_flat_reduced = ccdp.subtract_overscan(a_flat, overscan=a_flat[:, 2055:], median=True)
# Trim the overscan
a_flat_reduced = ccdp.trim_image(a_flat_reduced[:, :2048])
# Display the result so far
show_image(a_flat_reduced, cmap='gray')
plt.title('Single flat frame, overscan subtracted and trimmed')
# -
# Trimming off the overscan makes such a big difference primarily because the
# image stretch changed; the lowest pixel values are now around 18000 instead of
# 2000. With that change, the nonuniformity across the detector is much clearer.
# ### Subtracting bias is not necessary in this example
#
# For this particular set of images there are darks with exposure time 7, 70, and
# 300 sec. The flat images have the exposure times listed below:
set(ifc_raw.summary['exptime'][ifc_raw.summary['imagetyp'] == 'FLATFIELD'])
# These are close enough to the exposure time of the dark frames that there is no
# need to scale the darks by exposure time. If the darks are not going to be
# scaled then there is no need to subtract the bias.
# ### Subtract dark current, no scaling necessary in this example
#
# We need to subtract the dark without scaling it. Rather than manually figuring
# out which dark to subtract, we use the dark frame closest in exposure time to the
# flat, within a tolerance of 1 second to ensure that we do not end up using a
# dark *too* far off in exposure time from the flat.
#
# First, find the dark exposure time closest to the flat. We will need to do this
# again later in the notebook, so we define a function to do it.
closest_dark = find_nearest_dark_exposure(a_flat_reduced, actual_exposure_times)
# It will be convenient to be able to access the darks via a dictionary whose key
# is the exposure time, so we set that up below.
combined_darks = {ccd.header['exptime']: ccd for ccd in ifc_reduced.ccds(imagetyp='dark', combined=True)}
# Next, we subtract the dark from the flat and display the result.
a_flat_reduced = ccdp.subtract_dark(a_flat_reduced, combined_darks[closest_dark],
exposure_time='exptime', exposure_unit=u.second, scale=False)
show_image(a_flat_reduced, cmap='gray')
# There is not much change here; that is not surprising since the dark current in
# this camera is low.
# ### Calibrate all of the flats in the folder
# The cell below calibrates each of the flats in the folder, automatically
# grabbing the correct combined dark for each flat.
for ccd, file_name in ifc_raw.ccds(imagetyp='FLATFIELD', # Just get the bias frames
ccd_kwargs={'unit': 'adu'}, # CCDData requires a unit for the image if
# it is not in the header
return_fname=True # Provide the file name too.
):
# Subtract the overscan
ccd = ccdp.subtract_overscan(ccd, overscan=ccd[:, 2055:], median=True)
# Trim the overscan
ccd = ccdp.trim_image(ccd[:, :2048])
# Find the correct dark exposure
closest_dark = find_nearest_dark_exposure(ccd, actual_exposure_times)
# Subtract the dark current
ccd = ccdp.subtract_dark(ccd, combined_darks[closest_dark],
exposure_time='exptime', exposure_unit=u.second)
# Save the result; there are some duplicate file names so pre-pend "flat"
ccd.write(reduced_path / ('flat-' + file_name))
# ## Example 2: Dark frames are scaled
#
# The images in this example, like in the previous notebooks, is a
# thermoelectrically-cooled CCD described in more detail in the
# [overscan notebook](01-08-Overscan.ipynb#Case-2:-Thermo-electrically-cooled-Apogee-Aspen-CG16M).
#
# We'll go through this example twice: once with a single image to explain each
# step, and then again to process all of the flat frames in the directory of raw data.
#
# An image collection is defined below, along with a couple of settings useful for
# this example.
# +
reduced_path = Path('example2-reduced')
ifc_reduced = ccdp.ImageFileCollection(reduced_path)
combined_dark_files = ifc_reduced.files_filtered(imagetyp='dark', combined=True)
flat_image_type = 'FLAT'
# -
# The raw data should be in the directory `example-thermo-electric`.
# +
raw_data = Path('example-thermo-electric')
ifc_raw = ccdp.ImageFileCollection(raw_data)
# -
# The cell below checks that the files needed are available.
# +
n_combined_dark = len(combined_dark_files)
n_dark_expected = 1
expected_exposure_times = set([90])
if n_combined_dark < n_dark_expected:
raise RuntimeError('One or more combined dark is missing. Please re-run the dark notebook.')
elif n_combined_dark > n_dark_expected:
raise RuntimeError('There are more combined dark frames than expected.')
actual_exposure_times = set(h['exptime'] for h in ifc_reduced.headers(imagetyp='dark', combined=True))
if (expected_exposure_times - actual_exposure_times):
raise RuntimeError('Encountered unexpected exposure time in combined darks. '
'The unexpected times are {}'.format(actual_exposure_times - expected_exposure_times))
# -
# First, get one of the flat frames as a `CCDData` object and display it.
# +
a_flat = CCDData.read(ifc_raw.files_filtered(imagetyp='flat', include_path=True)[0], unit='adu')
show_image(a_flat, cmap='gray')
# -
# There is a much different pattern of variation across the sensor in this case
# than in Example 1. The multiple "donuts" in the image are pieces of dust and
# there is significant vignetting (darkening) in the top and bottom corners of the
# image on the right side.
# ### Subtract overscan and trim: only trim for this camera
#
# The overscan is not useful for this camera. The region to be retained after
# trimming is the Python slice `[:, :4096]`.
# +
# Trim the overscan
a_flat_reduced = ccdp.trim_image(a_flat[:, :4096])
# Display the result so far
show_image(a_flat_reduced, cmap='gray')
plt.title('Single flat frame, overscan trimmed')
# -
# Trimming off the overscan did not make a big difference primarily because the
# overscan region of this camera is not useful. A useful overscan would have had
# values around the bias level for this camera, about 1200 counts. The image
# stretch did change a bit; prior to trimming the lower end of the color scale was
# 38000 and now it is 40000.
# ### Subtracting bias is necessary
#
# For this particular set of images there is a combined dark with exposure time 90
# sec. The flat images have the exposure times listed below:
set(ifc_raw.summary['exptime'][ifc_raw.summary['imagetyp'] == 'FLAT'])
# These are quite different than the exposure time of the dark frames, so the dark
# will need to be scaled by exposure time, which means that the bias has been
# removed from the combined dark.
#
# Because of that, the bias needs to be removed from the flat before subtracting the
# dark.
combined_bias = list(ifc_reduced.ccds(combined=True, imagetyp='bias'))[0]
a_flat_reduced = ccdp.subtract_bias(a_flat_reduced, combined_bias)
# Display the result so far
show_image(a_flat_reduced, cmap='gray')
plt.title('Single flat frame, overscan trimmed and bias subtracted');
# Except for a change in the image scale shown on the color bar there isn't much
# visually different after subtracting the bias.
# ### Subtract dark current, scaling as needed
#
# Here we will need to scale the dark from the 90 sec exposure time of the dark
# frame to the exposure time of each flat image. The [ccdproc function
# `subtract_dark`](https://ccdproc.readthedocs.io/en/latest/api/ccdproc.subtract_dark.html#ccdproc.subtract_dark) provides keywords for doing this scaling automatically.
closest_dark = find_nearest_dark_exposure(a_flat_reduced, actual_exposure_times, tolerance=100)
# It will be convenient to be able to access the darks via a dictionary whose key
# is the exposure time, so we set that up below.
combined_darks = {ccd.header['exptime']: ccd for ccd in ifc_reduced.ccds(imagetyp='dark', combined=True)}
combined_darks
# Next, we subtract the dark from the flat and display the result.
a_flat_reduced = ccdp.subtract_dark(a_flat_reduced, combined_darks[closest_dark],
exposure_time='exptime', exposure_unit=u.second, scale=True)
show_image(a_flat_reduced, cmap='gray')
# There is not much change here; that is not surprising since the dark current in
# this camera is low.
# ### Calibrate all of the flats in the folder
# The cell below calibrates each of the flats in the folder, automatically
# grabbing the correct combined dark for each flat.
for ccd, file_name in ifc_raw.ccds(imagetyp='FLAT', # Just get the bias frames
return_fname=True # Provide the file name too.
):
# Trim the overscan
ccd = ccdp.trim_image(ccd[:, :4096])
# Find the correct dark exposure
closest_dark = find_nearest_dark_exposure(ccd, actual_exposure_times, tolerance=100)
# Subtract the dark current
ccd = ccdp.subtract_dark(ccd, combined_darks[closest_dark],
exposure_time='exptime', exposure_unit=u.second, scale=True)
# Save the result
ccd.write(reduced_path / file_name)
| notebooks/05-03-Calibrating-the-flats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../img/logo_white_bkg_small.png" align="left" />
#
# # Worksheet 6.0 Clustering
# This worksheet covers concepts relating to unsupervised learning. It should take no more than 20-30 minutes to complete. Please raise your hand if you get stuck.
#
# There are many ways to accomplish the tasks that you are presented with, however you will find that by using the techniques covered in class, the exercises should be relatively simple.
#
# ## Import the Libraries
# For this exercise, we will be using:
# * Pandas (http://pandas.pydata.org/pandas-docs/stable/)
# * Numpy (https://docs.scipy.org/doc/numpy/reference/)
# * Matplotlib (http://matplotlib.org/api/pyplot_api.html)
#
import pandas as pd
import numpy as np
from sklearn.cluster import KMeans, MiniBatchKMeans, DBSCAN
from sklearn.preprocessing import StandardScaler
from yellowbrick.cluster import KElbowVisualizer, SilhouetteVisualizer
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import style
from scipy.spatial.distance import cdist
style.use("ggplot")
# %matplotlib inline
# # K-Means Clustering
# In this example notebook, you will see how to implement K-Means Clustering in Python using Scikit-Learn and Pandas.
# Adapted from https://pythonprogramming.net/flat-clustering-machine-learning-python-scikit-learn/
#
# ## Step 1: Get Data:
# The first step is to prepare or generate the data. In this dataset, the observations only have two features, but K-Means can be used with any number of features. Since this is an unsupervised example, it is not necessary to have a "target" column.
data = pd.DataFrame([[1, 2],
[5, 8],
[1.5, 1.8],
[8, 8],
[1, 0.6],
[9, 11]], columns=['x','y'])
print( data )
# ## Step 2: Build the Model:
# Much like the supervised models, you first create the model then call the `.fit()` method using your data source. The model is now populated with both your centroids and labels. These can be accessed via the `.cluster_centers_` and `labels_` properties respectively.
#
# You can view the complete documentation here: http://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html
#
# K-Means also has a `.predict()` method which can be used to predict the label for an observation.
# +
kmeans = KMeans(n_clusters=2).fit(data)
centroids = kmeans.cluster_centers_
labels = kmeans.labels_
print(centroids)
print(labels)
# +
test = pd.DataFrame([[4,2]])
kmeans.predict(test)
print(test)
# -
# # Visualizing the Clusters
# The code below visualizes the clusters.
# +
data['labels'] = labels
#plt.plot(data, colors[data['labels'], markersize = 10)
group1 = data[data['labels']==1].plot( kind='scatter', x='x', y='y', s=100, color='DarkGreen', label="Group 1" )
group2 = data[data['labels']==0].plot( kind='scatter', x='x', y='y', s=100,color='Brown', ax=group1, label="Group 2" )
group1.legend(loc='upper center', bbox_to_anchor=(0.5, 1.05),
ncol=3, fancybox=True, shadow=True)
plt.scatter(centroids[:, 0],centroids[:, 1], marker = "x", s=150, linewidths = 5, zorder = 10)
plt.show()
# -
# # Clustering with Real Data
# Now that you've tried K-means on some generated data, let's try it on some real data and see what we can produce. As before the first step is to read in the data into a DataFrame.
#
# We will be using this data later, but the dataset consists of approximately 6000 domains--5000 of which were generated by various botnets and 1000 are from the Alexa 1 Million. The columns are:
#
# * `dsrc`: The source of the domain
# * `domain`: The actual domain
# * `length`: The length of the domain
# * `dicts`: Percentage containing dictionary words
# * `entropy`: The entropy of the domain
# * `numbers`: The number of digits in the domain
# * `ngram`: Different n-grams which appear in the domain (?)
data = pd.read_csv('../data/dga-full.csv')
data.sample(5)
# ## Scaling the Data
# Since clustering relies on measuring distances between objects it is important that all data points be on the same scale. There are various methods for doing this, which are beyond the scope of this class, however, for this example, we will use scikit-learn's `StandardScaler` to accomplish this. (http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html)
#
# The StandardScaler transforms each column by:
# * Subtracting from the element in each row the mean for each feature (column) and then taking this value and
# * Dividing by that feature's (column's) standard deviation.
#
# Scikit-learn has a transformer interface which is very similar to the other scikit-learn interfaces. The basic steps are:
# 1. Create the Scaler object
# 2. Using the feature matrix, call the `.fit()` method to "train" the Scaler
# 3. Use the `.transform()` method to scale the data.
#
# **NOTE**: When using a Scaler, it is important to train the scaler on your data, and use this trained scalers on any future predictions.
feature_columns = ['length', 'dicts','entropy','numbers','ngram']
scaled_feature_columns = ['scaled_length', 'scaled_dicts','scaled_entropy','scaled_numbers','scaled_ngram']
# +
#Step 1: Create the scaler
#Steps 2 & 3: Fit the scaler and transform this data
#Put the scaled data into a dataframe
# -
# Take a look at the data and you'll see that the data is now all scaled consistently.
# Finally for convenience, we're going to merge the scaled data with the non-scaled data.
final_data = pd.merge( data, scaled_df, left_index=True, right_index=True)
# ### Your Turn!
# Now that we have data that is suitable (maybe) for clustering, in the section below, perform K-Means clustering on this data set. Initially, start out with 2 clusters and assign the `cluster id` as a column in your DataFrame.
#
# Then do a `value_counts()` on the `dsrc` column for each cluster to see how the model divided the data. Try various values for `k` to see how it performed.
#
# Remember to use the **scaled features** for your clustering.
## Your code here...
# ## Food for thought:
# Now that you've done clustering with various numbers of clusters, it appears that the data acutally does break evenly into 2 clusters. Take a look at the original data and see if you can come up with a reason why that is the case.
# ## Visualizing Performance
# As we already know, it is difficult to measure the performance of clustering models since there usually is no known ground truth from which to evaluate your model. However, there are two techniques which
#
# The K-Elbow Visualizer implements the “elbow” method of selecting the optimal number of clusters for K-means clustering. K-means is a simple unsupervised machine learning algorithm that groups data into a specified number (k) of clusters. Because the user must specify in advance what k to choose, the algorithm is somewhat naive – it assigns all members to k clusters even if that is not the right k for the dataset.
#
# The elbow method runs k-means clustering on the dataset for a range of values for k (say from 1-10) and then for each value of k computes an average score for all clusters. By default, the distortion_score is computed, the sum of square distances from each point to its assigned center. Other metrics can also be used such as the silhouette_score, the mean silhouette coefficient for all samples or the calinski_harabaz_score, which computes the ratio of dispersion between and within clusters.
#
# When these overall metrics for each model are plotted, it is possible to visually determine the best value for K. If the line chart looks like an arm, then the “elbow” (the point of inflection on the curve) is the best value of k. The “arm” can be either up or down, but if there is a strong inflection point, it is a good indication that the underlying model fits best at that point. (http://www.scikit-yb.org/en/latest/api/cluster/elbow.html)
#
# In python there is a module called `YellowBrick` which facilitates visualizing the K-Elbow score. All of YellowBrick's visualizations follow essentually the same pattern:
#
# 1. Create the Visualizer Object
# 2. Call the `.fit()` method using the data
# 3. Call the `.poof()` method to render the visualization
#
# The snippet below demonstrates how to use the elbow method to visualize the clustering model's performance on this dataset.
# ```python
# visualizer = KElbowVisualizer(KMeans(), k=(4,12))
#
# visualizer.fit( feature_matrix )
# visualizer.poof()
# ```
#
# ### Your Turn!
# In the box below, create a visualization using the elbow method to see if there are any inflection points in the distortion score.
# Your code here...
# ## Another Way to Visualize Clustering Performance
# The Silhouette Coefficient is used when the ground-truth about the dataset is unknown and computes the density of clusters computed by the model. The score is computed by averaging the silhouette coefficient for each sample, computed as the difference between the average intra-cluster distance and the mean nearest-cluster distance for each sample, normalized by the maximum value. This produces a score between 1 and -1, where 1 is highly dense clusters and -1 is completely incorrect clustering. (http://www.scikit-yb.org/en/latest/api/cluster/silhouette.html)
#
#
# ### Your Turn!
# Using the YellowBrick `SilhouetteVisualizer`, try visualizing models with various values of `K`.
#
# **Note**: This visualization is quite expensive, so I recommend performing this using a sample o your original data.
# Your code here...
# ## DBSCAN
# Now that you've tried K-Means, let's try doing some clustering using DBSCAN (http://scikit-learn.org/stable/modules/generated/sklearn.cluster.DBSCAN.html). Remember that the main tuning parameters for DBSCAN are:
#
# * **epsilon (eps)**: The minimum distance between two samples
# * **min_samples**: The minimum number of samples needed to form a neighborhood
#
# By default epsilon is 0.5 and the min_samples is 5. First, try DBSCAN with the default options. If you use the `fit_predict()` function, you can save the results in a new column in your data.
#
# How did this compare with K-Means? Given that you actually know what the data really is, how did DBSCAN do in terms of identifing meaningful clusters? Look at the `dsrc` column and do `value_counts()` for the various neighhborhoods. What did you notice?
#
# Try again, but this time experiment with the values of epsilon and min_samples and see what DBSCAN comes up with.
# +
#Your code here...
| notebooks/Worksheet 6.0 - Clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 练习 1:写程序,可由键盘读入用户姓名例如<NAME>,让用户输入出生的月份与日期,判断用户星座,假设用户是金牛座,则输出,Mr. right,你是非常有性格的金牛座!。
# +
name = input('请输入你的姓名')
print('你好',name)
print('请输入出生的月份与日期')
month = int(input('月份:'))
date = int(input('日期:'))
if month == 4:
if date < 20:
print(name, '你是白羊座')
else:
print(name,'你是非常有性格的金牛座')
if month == 5:
if date < 21:
print(name, '你是非常有性格的金牛座')
else:
print(name,'你是双子座')
if month == 6:
if date < 22:
print(name, '你是双子座')
else:
print(name,'你是巨蟹座')
if month == 7:
if date < 23:
print(name, '你是巨蟹座')
else:
print(name,'你是狮子座')
if month == 8:
if date < 23:
print(name, '你是狮子座')
else:
print(name,'你是处女座')
if month == 9:
if date < 24:
print(name, '你是处女座')
else:
print(name,'你是天秤座')
if month == 10:
if date < 24:
print(name, '你是天秤座')
else:
print(name,'你是天蝎座')
if month == 11:
if date < 23:
print(name, '你是天蝎座')
else:
print(name,'你是射手座')
if month == 12:
if date < 22:
print(name, '你是射手座')
else:
print(name,'你是摩羯座')
if month == 1:
if date < 20:
print(name, '你是摩羯座')
else:
print(name,'你是水瓶座')
if month == 2:
if date < 19:
print(name, '你是水瓶座')
else:
print(name,'你是双鱼座')
if month == 3:
if date < 22:
print(name, '你是双鱼座')
else:
print(name,'你是白羊座')
# -
# ## 练习 2:写程序,可由键盘读入两个整数m与n(n不等于0),询问用户意图,如果要求和则计算从m到n的和输出,如果要乘积则计算从m到n的积并输出,如果要求余数则计算m除以n的余数的值并输出,否则则计算m整除n的值并输出。
# +
m = int(input('请输入一个整数,回车结束'))
n = int(input('请输入一个整数,不为零'))
intend = input('请输入计算意图,如 + * %')
if m<n:
min_number = m
else:
min_number = n
total = min_number
if intend == '+':
if m<n:
while m<n:
m = m + 1
total = total + m
print(total)
else:
while m > n:
n = n + 1
total = total + n
print(total)
elif intend == '*':
if m<n:
while m<n:
m = m + 1
total = total * m
print(total)
else:
while m > n:
n = n + 1
total = total * n
print(total)
elif intend == '%':
print(m % n)
else:
print(m // n)
# -
# ## 练习 3:写程序,能够根据北京雾霾PM2.5数值给出对应的防护建议。如当PM2.5数值大于500,则应该打开空气净化器,戴防雾霾口罩等。
# +
number = int(input('现在北京的PM2.5指数是多少?请输入整数'))
if number > 500:
print('应该打开空气净化器,戴防雾霾口罩')
elif 300 < number < 500:
print('尽量呆在室内不出门,出门佩戴防雾霾口罩')
elif 200 < number < 300:
print('尽量不要进行户外活动')
elif 100 < number < 200:
print('轻度污染,可进行户外活动,可不佩戴口罩')
else:
print('无须特别注意')
# -
# ## 尝试性练习:写程序,能够在屏幕上显示空行。
print('空行是我')
print('空行是我')
print('空行是我')
print( )
print('我是空行')
# ## 练习 4:英文单词单数转复数,要求输入一个英文动词(单数形式),能够得到其复数形式,或给出单数转复数形式的建议
# +
word = input('请输入一个单词,回车结束')
if word.endswith('s') or word.endswith('sh') or word.endswith('ch') or word.endswith('x'):
print(word,'es',sep = '')
elif word.endswith('y'):
if word.endswith('ay') or word.endswith('ey') or word.endswith('iy') or word.endswith('oy') or word.endswith('uy'):
print(word,'s',sep = '')
else:
word = word[:-1]
print(word,'ies',sep = '')
elif word.endswith('f'):
word = word[:-1]
print(word,'ves',sep = '')
elif word.endswith('fe'):
word = word[:-2]
print(word,'ves',sep = '')
elif word.endswith('o'):
print('词尾加s或者es')
else:
print(word,'s',sep = '')
# -
| chapter1/homework/localization/3-22/201611680049(3).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: jupyterlab
# language: python
# name: jupyterlab
# ---
# +
import os
import sys
sys.path.append('../')
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import src.io as sio
import src.preprocessing as spp
import src.fitting as sft
import ipympl
# -
DATA_FOLDER = "20210816_RTStageTesting_MW_MembraneSample"
DATA_FOLDERPATH = sio.get_qudiamond_folderpath(DATA_FOLDER)
FIGURE_FOLDERPATH = sio.get_figure_folderpath(DATA_FOLDER)
CONFOCAL_SUBFOLDER = r"2021\10\20211004\Confocal"
CONFOCAL_SUBFOLDERPATH = sio.get_qudi_data_path(os.path.join(DATA_FOLDER, CONFOCAL_SUBFOLDER))
# +
ODMR_SUBFOLDER1 = r"2021\10\20211020\ODMR"
ODMR_SUBFOLDERPATH1 = sio.get_qudi_data_path(os.path.join(DATA_FOLDER, ODMR_SUBFOLDER1))
ODMR_SUBFOLDER2 = r"2021\09\20210930\ODMR"
ODMR_SUBFOLDERPATH2 = sio.get_qudi_data_path(os.path.join(DATA_FOLDER, ODMR_SUBFOLDER2))
# -
# # CONFOCAL1
# +
fname = "20211004-1254-39_pillar1_confocal_xy_image_1.dat"
params = sio.read_qudi_parameters(fname, CONFOCAL_SUBFOLDERPATH)
data = np.genfromtxt(CONFOCAL_SUBFOLDERPATH + fname, dtype=None, delimiter='\t')
# +
# %matplotlib widget
extent = np.array([params['X image min (m)'], params['X image max (m)'], params['Y image min'], params['Y image max']]) * 1e6
fig, ax = plt.subplots()
img = ax.imshow(data/1e3, cmap="inferno", origin="lower", vmin=30, vmax=400, extent=extent)
cbar = fig.colorbar(img, ax=ax)
cbar.set_label("Fluorescence (Mc/s)")
ax.set_xlabel("X position (μm)")
ax.set_ylabel("Y position (μm)")
#sio.save_figures(fname, folder=os.path.join(FIGURE_FOLDERPATH, CONFOCAL_SUBFOLDER))
# -
# # CONFOCAL2
# +
fname = "20211004-1346-16_pillar1_confocal_xy_image_1.dat"
params = sio.read_qudi_parameters(fname, CONFOCAL_SUBFOLDERPATH)
data = np.genfromtxt(CONFOCAL_SUBFOLDERPATH + fname, dtype=None, delimiter='\t')
# +
# %matplotlib widget
extent = np.array([params['X image min (m)'], params['X image max (m)'], params['Y image min'], params['Y image max']]) * 1e6
fig, ax = plt.subplots()
img = ax.imshow(data/1e3, cmap="inferno", origin="lower", vmin=60, vmax=400, extent=extent)
cbar = fig.colorbar(img, ax=ax)
cbar.set_label("Fluorescence (Mc/s)")
ax.set_xlabel("X position (μm)")
ax.set_ylabel("Y position (μm)")
sio.save_figures(fname, folder=os.path.join(FIGURE_FOLDERPATH, CONFOCAL_SUBFOLDER))
# +
fig, (ax1, ax2) = plt.subplots(nrows=2)
fname = "20211020-1701-48_ODMR_data_ch0.dat"
params = sio.read_qudi_parameters(fname, ODMR_SUBFOLDERPATH1)
data = np.genfromtxt(ODMR_SUBFOLDERPATH1 + fname, dtype=np.float, delimiter='\t')
freq, counts = data[:, 0], data[:, 1]
a = 15
ax1.plot(freq[a:]/1e9, counts[a:]/1e3, "o--", label="NV1")
ax1.set_xlabel("Frequency (GHz)")
ax1.set_ylabel("Fluorescence (kc/s)")
fname = "20210930-1836-08_ODMR_data_ch0.dat"
params = sio.read_qudi_parameters(fname, ODMR_SUBFOLDERPATH2)
data = np.genfromtxt(ODMR_SUBFOLDERPATH2 + fname, dtype=np.float, delimiter='\t')
freq, counts = data[:, 0], data[:, 1]
a, b = 20, 55
ax2.plot(freq[a:b]/1e9, counts[a:b]/1e3, "o--", color="tab:orange", label="NV2")
ax2.set_xlabel("Frequency (GHz)")
ax2.set_ylabel("Fluorescence (kc/s)")
fig.legend()
sio.save_figures("Comparison_ODMR_at_two_NVs", folder=os.path.join(FIGURE_FOLDERPATH, ODMR_SUBFOLDER1))
# -
# # ODMR1
# +
fname = "20211020-1701-48_ODMR_data_ch0.dat"
params = sio.read_qudi_parameters(fname, ODMR_SUBFOLDERPATH1)
data = np.genfromtxt(ODMR_SUBFOLDERPATH1 + fname, dtype=np.float, delimiter='\t')
freq, counts = data[:, 0], data[:, 1]
# -
fig, ax = plt.subplots()
a = 15
ax.plot(freq[a:]/1e9, counts[a:]/1e3, "o--")
ax.set_xlabel("Frequency (GHz)")
ax.set_ylabel("Fluorescence (kc/s)")
sio.save_figures(fname, folder=os.path.join(FIGURE_FOLDERPATH, ODMR_SUBFOLDER1))
# # ODMR2
# +
fname = "20210930-1836-08_ODMR_data_ch0.dat"
params = sio.read_qudi_parameters(fname, ODMR_SUBFOLDERPATH2)
data = np.genfromtxt(ODMR_SUBFOLDERPATH2 + fname, dtype=np.float, delimiter='\t')
freq, counts = data[:, 0], data[:, 1]
# -
fig, ax = plt.subplots()
a, b = 20, 55
ax.plot(freq[a:b]/1e9, counts[a:b]/1e3, "o--")
ax.set_xlabel("Frequency (GHz)")
ax.set_ylabel("Fluorescence (kc/s)")
sio.save_figures(fname, folder=os.path.join(FIGURE_FOLDERPATH, ODMR_SUBFOLDER2))
# # MW transmission
| notebooks/20210816_RTStageTesting_MW_MembraneSample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Explore The Data: Explore Continuous Features
#
# Using the Titanic dataset from [this](https://www.kaggle.com/c/titanic/overview) Kaggle competition.
#
# This dataset contains information about 891 people who were on board the ship when departed on April 15th, 1912. As noted in the description on Kaggle's website, some people aboard the ship were more likely to survive the wreck than others. There were not enough lifeboats for everybody so women, children, and the upper-class were prioritized. Using the information about these 891 passengers, the challenge is to build a model to predict which people would survive based on the following fields:
#
# - **Name** (str) - Name of the passenger
# - **Pclass** (int) - Ticket class (1st, 2nd, or 3rd)
# - **Sex** (str) - Gender of the passenger
# - **Age** (float) - Age in years
# - **SibSp** (int) - Number of siblings and spouses aboard
# - **Parch** (int) - Number of parents and children aboard
# - **Ticket** (str) - Ticket number
# - **Fare** (float) - Passenger fare
# - **Cabin** (str) - Cabin number
# - **Embarked** (str) - Port of embarkation (C = Cherbourg, Q = Queenstown, S = Southampton)
#
# **This section focuses on exploring the `Pclass`, `Age`, `SibSp`, `Parch`, and `Fare` features.**
# ### Read In Data
# +
# Read in our data
import pandas as pd
from scipy import stats
titanic = pd.read_csv('../../../data/titanic.csv')
titanic.head()
# -
# Drop all categorical features
cat_feat = ['PassengerId', 'Name', 'Ticket', 'Sex', 'Cabin', 'Embarked']
titanic.drop(cat_feat, axis=1, inplace=True)
titanic.head()
# ### Explore Continuous Features
# Look at the general distribution of these features
titanic.describe()
# Look at the correlation matrix
titanic.corr()
# Look at fare by different passenger class levels
titanic.groupby('Pclass')['Fare'].describe()
# not much fare overlap between 3 classes -- correlated features may confuse models
# +
def describe_cont_feature(feature):
print('\n*** Results for {} ***'.format(feature))
print(titanic.groupby('Survived')[feature].describe())
print(ttest(feature))
def ttest(feature):
survived = titanic[titanic['Survived']==1][feature]
not_survived = titanic[titanic['Survived']==0][feature]
tstat, pval = stats.ttest_ind(survived, not_survived, equal_var=False)
print('t-statistic: {:.1f}, p-value: {:.3}'.format(tstat, pval))
# -
# Look at the distribution of each feature at each level of the target variable
for feature in ['Pclass', 'Age', 'SibSp', 'Parch', 'Fare']:
describe_cont_feature(feature)
# Pclass and Fare show stat sig results between survive/not
# Look at the average value of each feature based on whether Age is missing
titanic.groupby(titanic['Age'].isnull()).mean()
# missing age does not matter that much vs data w/ age reported
# need to find out if missing value has any meaningful patterns
| ml_feature/03_Explore_Data/03_02/End/03_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# Notebook to analyse the PyBossa taskruns from the Expert App Version 2.
# # Load Libraries and Data
from mod_finder_util import mod_finder_util
mod_finder_util.add_modules_origin_search_path()
# +
import pandas as pd
import seaborn as sns
import numpy as np
import modules.utils.firefox_dataset_p2 as fd
from sklearn.metrics import cohen_kappa_score
taskruns = fd.TaskRuns.read_expert_taskruns_df()
# -
# # Grouping Contributions By Time
# +
taskruns['created'] = pd.to_datetime(taskruns['created'], yearfirst=True)
taskruns['created_month'] = taskruns.apply(lambda row: row['created'].month, axis=1)
taskruns['created_day'] = taskruns.apply(lambda row: row['created'].day, axis=1)
taskruns['created_hour'] = taskruns.apply(lambda row: row['created'].hour, axis=1)
grouped_trs = taskruns.groupby(by=['created_month','created_day','created_hour']).count()
grouped_trs
# -
# # Count TaskRuns by Task
# +
cols = ['task_id','id']
df = taskruns[cols].groupby(by='task_id').count()
df.rename(columns={"id" :'count_trs'}, inplace=True)
display(df.tail(10))
print(df.shape)
# -
# # Check All Tasks Have At Least One Answer
print(set(df.index) - set(range(1928,2020+1,1)))
# # Analysis of Taskruns Infos
taskruns[['bug_id','answers']].head(10)
taskruns.dtypes
# # Compare Order of Taskruns
#
# The order of the taskruns must be the same, ordered by the finish time or by the bug_id fields.
# +
taskruns_ordered_by_finish_time = taskruns.sort_values(by='finish_time')
taskruns_ordered_by_bug_id = taskruns.copy() ## already ordered by bug_id
diffs = taskruns_ordered_by_finish_time.bug_id - taskruns_ordered_by_bug_id.bug_id
d = np.sum(diff)
assert d == 0
| preprocessing/trs_analysis/exp_trs_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Cleaning
# +
# import the library
# %matplotlib inline
import pandas as pd
import numpy as np
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
# convert scientific notation to decimals
pd.set_option('display.float_format', lambda x: '%.2f' % x)
# -
# # 1. Load Datasets
# +
#Source file: https://www.kaggle.com/usdot/flight-delays#flights.csv
#Main dataset
df_delayed_flights = pd.read_csv('../../Data/flights.csv', low_memory=False)
#Complementary datasets
df_airports = pd.read_csv('../../Data/airports.csv', low_memory=False)
df_airlines = pd.read_csv('../../Data/airlines.csv', low_memory=False)
# -
# # 2. Summarize the data
# +
print('------- Main Dataset, Flights -------')
print(df_delayed_flights.shape)
print(df_delayed_flights.columns)
print(df_delayed_flights.head())
print(df_delayed_flights.describe())
print('\n ------- Airports -------')
print(df_airports.shape)
print(df_airports.columns)
print(df_airports.head())
print(df_airports.describe())
print('\n ------- Airlines -------')
print(df_airlines.shape)
print(df_airlines.columns)
print(df_airlines.head())
print(df_airlines.describe())
# -
# # 3. Data Cleaning
# ## 3.1. Merge Columns - DATE
df_delayed_flights['DATE'] = pd.to_datetime(df_delayed_flights[['YEAR','MONTH', 'DAY']])
print(df_delayed_flights.columns)
# ## 3.2. Remove not relevant and duplicated columns
df_delayed_flights = df_delayed_flights.drop(['YEAR','DAY','DAY_OF_WEEK'], axis=1)
df_delayed_flights = df_delayed_flights.drop(['TAXI_OUT','TAXI_IN','WHEELS_OFF', 'WHEELS_ON','AIR_TIME','ELAPSED_TIME'], axis=1)
print(df_delayed_flights.columns)
# +
df_airports = df_airports[['IATA_CODE','AIRPORT']]
print(df_airports.columns)
# -
# ## 3.3. Rename Columns
df_airports.columns = ['AIRPORT_CODE','AIRPORT_NAME']
df_airlines.columns = ['AIRLINE_CODE','AIRLINE_NAME']
# ## 3.4. Change type
# +
# Function to convert from 'HHMM' string to datetime.time
# Reference: https://www.kaggle.com/fabiendaniel/predicting-flight-delays-tutorial
import datetime
def format_time(time_string):
if isinstance(time_string, datetime.time):
return time_string
if pd.isnull(time_string):
return np.nan
else:
if time_string == 2400: #12 midnight
time_string = 0
time_string = "{0:04d}".format(int(time_string)) #Format the string to be as 4 decimals (hh:mm)
formated_time = datetime.time(int(time_string[0:2]), int(time_string[2:4])) #Split the 4 digits into 2 parts
return formated_time
# +
# Call the Function:
df_delayed_flights['SCHEDULED_DEPARTURE'] = df_delayed_flights['SCHEDULED_DEPARTURE'].apply(format_time)
df_delayed_flights['DEPARTURE_TIME'] = df_delayed_flights['DEPARTURE_TIME'].apply(format_time)
df_delayed_flights['SCHEDULED_ARRIVAL'] =df_delayed_flights['SCHEDULED_ARRIVAL'].apply(format_time)
df_delayed_flights['ARRIVAL_TIME'] = df_delayed_flights['ARRIVAL_TIME'].apply(format_time)
#Print a sample..
df_delayed_flights.loc[:5, ['SCHEDULED_DEPARTURE', 'SCHEDULED_ARRIVAL', 'DEPARTURE_TIME',
'ARRIVAL_TIME', 'DEPARTURE_DELAY', 'ARRIVAL_DELAY']]
# -
# ## 3.5. Change Values
# +
#Replace cancellation reason with meaningful values
df_delayed_flights["CANCELLATION_REASON"].replace({'A':'Airline',
'B':'Weather',
'C':'National Air System',
'D':'Security'}, inplace=True)
df_delayed_flights["CANCELLATION_REASON"].value_counts()
# -
# ## 3.6. Missing values
print("Delayed Flights Missing Values:\n", df_delayed_flights.isnull().sum())
print("Airlines Missing Values:\n", df_airlines.isnull().sum())
print("Airports Missing Values:\n", df_airports.isnull().sum())
# ### CANCELLATION_REASON
print("Total number of delayed flights: ", len(df_delayed_flights))
print("Cancelled flights= ", sum(df_delayed_flights['CANCELLED']))
print(df_delayed_flights['CANCELLATION_REASON'].value_counts())
# #### Conclusion: Number of missing data under the variable "CANCELLATION_REASON" is large since when the flight was not cancelled, no reason code was assigned. We can replace no values with "Not Cancelled":
df_delayed_flights['CANCELLATION_REASON'] = df_delayed_flights['CANCELLATION_REASON'].fillna('Not_Cancelled')
df_delayed_flights['CANCELLATION_REASON'].isnull().sum()
print(df_delayed_flights['CANCELLATION_REASON'].value_counts())
# # 4. Remove Outliers
df_delayed_flights[["DEPARTURE_DELAY","ARRIVAL_DELAY"]].plot.box()
plt.show()
# +
plt.hist(df_delayed_flights['ARRIVAL_DELAY'], bins=150)
plt.title("Arrival Delays")
plt.show()
plt.hist(df_delayed_flights['DEPARTURE_DELAY'], bins=150)
plt.title("Departure Delays")
plt.show()
# +
#Determine Outliers
mean_arrival_delays = np.mean(df_delayed_flights['ARRIVAL_DELAY'] )
sd_arrival_delays = np.std(df_delayed_flights['ARRIVAL_DELAY'])
mean_departure_delays = np.mean(df_delayed_flights['DEPARTURE_DELAY'])
sd_departure_delays = np.std(df_delayed_flights['DEPARTURE_DELAY'])
print('Arrival Delays:\t \t Mean = {0} \t SD = {1}'.format(mean_arrival_delays, sd_arrival_delays))
print('Departure Delays:\t Mean = {0} \t SD = {1}'.format(mean_departure_delays, sd_departure_delays))
# +
#Arrrival_delay or Departure_delay != 0
arrival_delays = df_delayed_flights[df_delayed_flights['ARRIVAL_DELAY'] != 0.00]['ARRIVAL_DELAY']
departure_delays = df_delayed_flights[df_delayed_flights['DEPARTURE_DELAY'] != 0.00]['DEPARTURE_DELAY']
print(arrival_delays.shape)
mean_ad = np.mean(arrival_delays)
sd_ad = np.std(arrival_delays)
mean_dd = np.mean(departure_delays)
sd_dd = np.std(departure_delays)
print("With removing on-time flights:")
print('Arrival Delays:\t \t Mean = {0} \t SD = {1}'.format(mean_ad, sd_ad))
print('Departure Delays:\t Mean = {0} \t SD = {1}'.format(mean_dd, sd_dd))
# +
# Removing
flights_to_remove = []
# remove based on arrival and departure delays (normal distribution)
flights_to_remove = flights_to_remove + list(df_delayed_flights[df_delayed_flights['ARRIVAL_DELAY'] > mean_ad + 3.0* sd_ad].index)
flights_to_remove = flights_to_remove + list(df_delayed_flights[df_delayed_flights['ARRIVAL_DELAY'] < mean_ad - 3.0* sd_ad].index)
flights_to_remove = flights_to_remove + list(df_delayed_flights[df_delayed_flights['DEPARTURE_DELAY'] > mean_dd + 3.0* sd_dd].index)
flights_to_remove = flights_to_remove + list(df_delayed_flights[df_delayed_flights['DEPARTURE_DELAY'] < mean_dd - 3.0* sd_dd].index)
print('')
print('# Flights to remove', len(flights_to_remove))
# +
new_delayed_flights = df_delayed_flights[~df_delayed_flights.index.isin(flights_to_remove)]
print("Was: ", df_delayed_flights.shape, " Now: ", new_delayed_flights.shape)
# +
plt.hist(new_delayed_flights['ARRIVAL_DELAY'], bins=150)
plt.title("Arrival Delays")
plt.show()
plt.hist(new_delayed_flights['DEPARTURE_DELAY'], bins=150)
plt.title("Departure Delays")
plt.show()
# -
# ## 5. Merging datasets
# +
df_merge_v1 = new_delayed_flights.copy()
#Merge Airlines and Flights dfs
df_merge_v1 = pd.merge(new_delayed_flights, df_airlines, left_on='AIRLINE', right_on='AIRLINE_CODE', how='left')
#Merge Airports and Flights dfs on Origin_Airport and Airport_Code
df_merge_v1 = pd.merge(df_merge_v1, df_airports, left_on='ORIGIN_AIRPORT', right_on='AIRPORT_CODE', how='left')
df_merge_v1.rename(columns={'ORIGIN_AIRPORT':'ORIGIN_AC', #Origin Airport Code
'AIRPORT_NAME':'ORIGIN_AIRPORT', #Origin Airport Name
'DESTINATION_AIRPORT': 'DESTINATION_AC'}, inplace=True) #Dest Airport Code
df_merge_v1.drop(['AIRLINE','AIRPORT_CODE'], axis=1, inplace=True)
#Merge Airports and Flights dfs on Destination_Airport and Airport_Code
df_merge_v1 = pd.merge(df_merge_v1, df_airports, left_on='DESTINATION_AC', right_on='AIRPORT_CODE', how='left')
df_merge_v1.rename(columns={'AIRPORT_NAME':'DESTINATION_AIRPORT'}, inplace=True) #Dest Airport Name
df_merge_v1.drop('AIRPORT_CODE', axis=1, inplace=True)
# +
print("Merged Dataframe Columns: \n", df_merge_v1.columns)
df_merge_v1[['ORIGIN_AIRPORT', 'ORIGIN_AC','DESTINATION_AIRPORT', 'DESTINATION_AC']]
# -
# ## Save file
df_merge_v1.to_csv('../../Data/flightsmerged.csv', index=False)
| Code/Cleaning_and_Exploration/Data-Cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
#
# # Filtering the Pancealab Twitter Dataset
#
# The dataset available on the Pancealab [github repo](https://github.com/thepanacealab/covid19_twitter) is very large
# and analysing it in its entirety would be expensive, lengthy and out of scope for the project. Instead using this notebook,
# we can filter the files by selecting a number of tweets and a number of languages. It was decided that 1000 from 4 languages
# would suffice for this part of the project. Later on two more languages are added.
# + pycharm={"name": "#%%\n"}
import pandas as pd
from tqdm.notebook import tqdm
valid_languages = ['en', 'es', 'fr', 'de']
# + pycharm={"name": "#%%\n"}
import importlib
import matplotlib.pyplot as plt
import os
import sys
from multiplex import drawable
drawable = importlib.reload(drawable)
# reload the style
plt.style.use(os.path.join(sys.path[0], 'styles', "multiplex.style"))
def pie(data, labelname, figname):
figure = plt.figure(figsize=(5, 5))
viz = drawable.Drawable(figure)
temp = data[labelname].value_counts()
df = pd.DataFrame({'labels': [str(temp.values.tolist()[i]) + ' ' + l for i, l in enumerate(temp.index)],
'values': temp.values
})
labels = df['labels']
sizes = df['values']
patches, texts = viz.pie(sizes, shadow=True, startangle=90, pctdistance=1.1, labeldistance=1.2)
plt.legend(patches, labels, loc="right")
viz.axis('equal')
viz.set_title(figname, loc='left')
viz.savefig(fname='Output/'+figname,dpi=300, bbox_inches='tight')
# + [markdown] pycharm={"name": "#%% md\n"}
# In this cell we go over the files in the ProcessOrNot/Data/pancealab/ folder and extract 1000 tweets for each language
# chosen.
# + pycharm={"name": "#%%\n"}
files = ['Data/pancealab/2021-01-01_clean-dataset.tsv',
'Data/pancealab/2021-02-01_clean-dataset.tsv',
'Data/pancealab/2021-03-01_clean-dataset.tsv']
for i, f in enumerate(files):
data = pd.read_csv(f, sep='\t')
valid_data = pd.DataFrame(columns=data.columns)
lang_indices = {}
# add each index to its label
languages = data["lang"]
for j, lang in tqdm(enumerate(languages)):
if lang in valid_languages:
if lang not in lang_indices:
lang_indices[lang] = []
lang_indices[lang].append(j)
for lang in valid_languages:
valid_data = valid_data.append(data.iloc[lang_indices[lang][:1000]])
v_counts = valid_data['lang'].value_counts()
pie(valid_data, 'lang', '2021-0'+str(i)+'-01 Filtered Language Distribution')
valid_data.to_csv('Data/FilteredTwitterIDs/' + str(i) + '.csv')
| ProcessOrNot/Collect Tweet ID.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + kernel="SoS"
[100]
parameter: num = 2
input: for_each = {'i': range(num)}
output: f'test_{i}.txt'
task:
sh: expand=True
sleep {5*i}
echo test_{i} > test_{i}.txt
echo "Test_{i}.txt generated"
# + kernel="SoS"
# %sosrun --num 3
# -
| doc/examples/HomePage_Example_3B.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:notebook] *
# language: python
# name: conda-env-notebook-py
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import xarray as xr
import intake,yaml
import intake_esm
from scipy import special
import keras
from keras.models import Model
from keras.layers import Dense, Input
# ## Retrieve for the data (here example of IPSL-Historical)
#
# for the other models and scenarios, please change the variable "expname_filter" and "model_filter"
col_url = "https://cmip6-nc.s3.us-east-2.amazonaws.com/esgf-world.json"
col = intake.open_esm_datastore(col_url)
# %pwd
# +
#define location of weights file and image output here
import sys,os
wgtsdir = "../../ensembleMLP"
imgdir = "png_historical_IPSL"
if(os.path.exists(wgtsdir)):
print("weight dir exists")
else:
sys.exit("weights directory is invalid")
if(os.path.exists(wgtsdir+'/vanillamodel_0.h5')):
print("weight file exists")
else:
sys.exit("weights directory is invalid:"+wgtsdir+'/vanillamodel_0.h5')
if not os.path.exists(imgdir):
os.makedirs(imgdir)
# +
#Examples to just search for what we want from the catalog
expname_filter = ['historical']
table_id_filter = 'Omon'
model_filter = 'IPSL-CM6A-LR'
#variable_id_filter = "zos"
grid_label_filter = 'gn'
ens_filter = "r1i1p1f1"
#version_filter = "v20190726"
cat = col.search(experiment_id=expname_filter,
mip_table=table_id_filter,
model=model_filter,
variable=['zos'],
grid_label=grid_label_filter,
ensemble_member = ens_filter
)
col_tauuo = col.search(experiment_id=expname_filter,
model=model_filter,
mip_table=table_id_filter,
grid_label=grid_label_filter,
variable=['tauuo'],
ensemble_member = ens_filter
)
col_tauvo = col.search(experiment_id=expname_filter,
model=model_filter,
mip_table=table_id_filter,
grid_label=grid_label_filter,
variable=['tauvo'],
ensemble_member = ens_filter
)
col_bathm = col.search(experiment_id=['1pctCO2'],
model=model_filter,
mip_table='Ofx',
grid_label=grid_label_filter,
variable='deptho'
)
# -
cat.df
col_bathm.df
col_tauuo.df
col_tauvo.df
# ### Read data
dset_dict_zos = cat.to_dataset_dict(cdf_kwargs={'chunks': {'time': 20}}, storage_options={'anon':True})
dset_dict_tauuo = col_tauuo.to_dataset_dict(cdf_kwargs={'chunks': {'time': 20}}, storage_options={'anon':True})
dset_dict_tauvo = col_tauvo.to_dataset_dict(cdf_kwargs={'chunks': {'time': 20}}, storage_options={'anon':True})
dset_dict_bathm = col_bathm.to_dataset_dict(storage_options={'anon':True})
zos_xr = dset_dict_zos["CMIP6.IPSL.IPSL-CM6A-LR.historical.Omon"]
tauuo_xr = dset_dict_tauuo["CMIP6.IPSL.IPSL-CM6A-LR.historical.Omon"]
tauvo_xr = dset_dict_tauvo["CMIP6.IPSL.IPSL-CM6A-LR.historical.Omon"]
bathm_xr = dset_dict_bathm["CMIP6.IPSL.IPSL-CM6A-LR.1pctCO2.Ofx"]
# ### Some plotting
# +
plt.figure(figsize=(15,15))
plt.subplot(2,2,1)
zos_xr.zos.sel(time=slice("1992", "2011")).mean(axis=1).plot()
plt.subplot(2,2,2)
bathm_xr.deptho[0,:,:,:].max(axis=0).plot()
plt.subplot(2,2,3)
tauuo_xr.tauuo.sel(time=slice("1992", "2011")).mean(axis=1).plot()
plt.subplot(2,2,4)
tauvo_xr.tauvo.sel(time=slice("1992", "2011")).mean(axis=1).plot()
# -
# ### Preparing the features for the EnsembleMLP
def grad(d,y,x):
"Returns a three point derivative."
g0, g1 = np.gradient(d)
grady=g0/y
gradx=g1/x
return grady, gradx
Bathm = bathm_xr.deptho[0,:,:,:].max(axis=0) ###This is specific to IPSL bathymetry
####
lonRoll = np.roll(Bathm.nav_lon.values, axis=1, shift=-1)
Londiff = lonRoll - Bathm.nav_lon.values
latDiff=1.111774765625000e+05
latY=np.gradient(Bathm.nav_lat.values, axis=0)*latDiff
lonX=np.abs(np.cos(Bathm.nav_lat.values*np.pi/180))*latDiff*Londiff
####
Omega=7.2921e-5
f=(2*Omega*np.sin(Bathm.nav_lat.values*np.pi/180))
from pickle import load
#######!!!!!! it might ask you to use scikit-learn 0.22.2.post1 !!!!!! check if you get the good .mean_ and .scale_
#(array([-2.20681035e-11, 7.85894841e-03, 1.92149912e-08, 1.38805767e-07,
# 3.96657293e+03, -1.45941754e-04, 5.42086609e-04, -1.21754470e-05]),
# array([1.46399667e-10, 7.07289355e-01, 1.89893314e-07, 4.55081599e-07,
# 1.44475515e+03, 4.87804804e-03, 4.64697134e-03, 9.54868121e-05]))
scaler = load(open('scaler_v1.pkl', 'rb'))
scaler.mean_, scaler.scale_
listyears = ["1872","1891","1892","1911","1912","1931","1932","1951","1952","1971","1972","1991","1992","2011"]
# +
datapredicted_results = np.nan * np.zeros((7,)+Bathm.shape)
for i in range(7):
print(listyears[2*i]+listyears[2*i+1])
SSH20Mean = zos_xr.zos.isel(ensemble_member=0).sel(time=slice(listyears[2*i], listyears[2*i+1])).mean(axis=0).values
TauUOMean = tauuo_xr.tauuo.isel(ensemble_member=0).sel(time=slice(listyears[2*i], listyears[2*i+1])).mean(axis=0).values
TauVOMean = tauvo_xr.tauvo.isel(ensemble_member=0).sel(time=slice(listyears[2*i], listyears[2*i+1])).mean(axis=0).values
####
print('Calculate grads')
gradSSH_y, gradSSH_x = grad(SSH20Mean,latY,lonX)
gradBathm_y, gradBathm_x = grad(Bathm,latY,lonX)
gradUx_y, gradUx_x = grad(TauUOMean,latY,lonX)
gradUy_y, gradUy_x = grad(TauVOMean,latY,lonX)
####
print('Calculate CurlTau')
curlTau =(gradUy_x-gradUx_y)/1032
####
print('Calculate missindx')
missingdataindex = np.isnan(curlTau*SSH20Mean*
gradSSH_x*gradSSH_y*
Bathm*gradBathm_x*
gradBathm_y*f)
####
print('Creating TotalDataset')
TotalDataset = np.stack((curlTau[~missingdataindex],
SSH20Mean[~missingdataindex],
gradSSH_x[~missingdataindex],
gradSSH_y[~missingdataindex],
Bathm.values[~missingdataindex],
gradBathm_x[~missingdataindex],
gradBathm_y[~missingdataindex],
f[~missingdataindex]),1)
####
print('NN pred')
datapredicted_results_vanilla_ensemble = np.nan * np.zeros((50,TotalDataset.shape[0],6))
for j in range(50):
I1 = Input(shape=(8,))
h1 = Dense(24, activation='tanh')(I1)
h1 = Dense(24, activation='tanh')(h1)
h1 = Dense(16, activation='tanh')(h1)
h1 = Dense(16, activation='tanh')(h1)
Output = Dense(6, activation='softmax')(h1)
vanillamodel = Model(I1, Output)
vanillamodel.load_weights(wgtsdir+'/vanillamodel_'+str(j)+'.h5') ###path in github repo
datapredicted = vanillamodel.predict(scaler.transform(TotalDataset))#a1r
datapredicted_results_vanilla_ensemble[j,...] = datapredicted
tmpMean = np.mean(datapredicted_results_vanilla_ensemble,0)
tmpSoftmax = special.softmax(tmpMean,1)
datapredicted_results[i,:,:][~missingdataindex] = np.argmax(tmpSoftmax, 1)
####
#np.save('/home/jovyan/gfdl-THOR/Othermodels/IPSL-Historical-198701-201112-EnsembleMLP.npy',datapredicted_results)
# -
from matplotlib.colors import ListedColormap
mycmap = ListedColormap(['royalblue', 'cyan','yellow', 'orange', 'magenta', 'red'])
ecco_label = np.load('eccolabel.npy')
# +
plt.figure(figsize=(20,10))
plt.subplot(1,2,1)
plt.imshow(datapredicted_results[-1,:,:][::-1,:], cmap=mycmap)
plt.colorbar(fraction=0.024, pad=0.04)
plt.title('IPSL regimes 1992-2011 predicted by the Ensemble NN')
plt.subplot(1,2,2)
plt.imshow(ecco_label[::-1,:], cmap=mycmap)
plt.colorbar(fraction=0.024, pad=0.04)
plt.title('ECCO regimes 1992-2011')
# -
# # Save the images for the 20 years chunks
for i in range(7):
plt.figure(figsize=(20,10))
#plt.subplot(1,2,1)
plt.imshow(datapredicted_results[i,:,:][::-1,:], cmap=mycmap)
plt.colorbar(fraction=0.024, pad=0.04)
plt.title('historical '+str(listyears[2*i])+' '+str(listyears[2*i+1]), fontsize=50)
plt.savefig('png_historical_IPSL/'+str(listyears[2*i])+str(listyears[2*i+1])+'.png')
# ## create a gif
# +
import os
import imageio
png_dir = 'png_historical_IPSL/'
images = []
for file_name in sorted(os.listdir(png_dir)):
if file_name.endswith('.png'):
file_path = os.path.join(png_dir, file_name)
images.append(imageio.imread(file_path))
imageio.mimsave('png_historical_IPSL/IPSL-png_historical.gif', images, fps=1)
# -
| THOR/ApplicationOnCMIPModels/IPSL Model/.ipynb_checkpoints/IPSL-EnsemlbleMLP-Historical-Github-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from typing import List
import Item
# +
def get_distance(item1: Item, item2: Item):
cord1 = item1.coordinate
cord2 = item2.coordinate
tuple1 = (cord1['lat'], cord1['lon'])
tuple2 = (cord2['lat'], cord2['lon'])
return haversine(tuple1, tuple2)
# +
class Planner:
optimal_route = []
optimal_cost = 0
types = {'malls', 'restaurants', 'cafes', 'fast_food', 'architecture', 'cultural', 'sport', 'natural',
'marketplaces', 'hotel'}
constraints = {
'malls': 2,
'restaurants': 2,
'cafes': 1,
'fast_food': 1,
'architecture': 3,
'cultural': 3,
'sport': 3,
'natural': 2,
'marketplaces': 2,
'hotel': 1,
'historic': 2,
'shop': 2
}
def __init__(self, items: List):
self.graph = []
self.items = items
for i in range(len(items)):
cities_names.append(f"{items[i].item_id}, Type:{items[i].item_type}")
self.graph.append([])
for j in range(len(items)):
self.graph[i].append(floor(get_distance(items[i], items[j]) * 1000) / 1000)
def delta(self, n1, n2, n3, n4):
return self.graph[n1][n3] + self.graph[n2][n4] - self.graph[n1][n2] - self.graph[n3][n4]
def plan_two_opt(self, iterations=5):
i = 0
total_costs = []
while i < iterations:
i += 1
initial_route = [0] + random.sample(range(1, len(cities_names)), len(cities_names) - 1)
best_route = initial_route
improved = True
while improved:
improved = False
for i in range(1, len(self.graph) - 2):
for j in range(i + 1, len(self.graph)):
if j - i == 1:
continue
if self.delta(best_route[i - 1], best_route[i], best_route[j - 1], best_route[j]) < 0:
best_route[i:j] = best_route[j - 1:i - 1:-1]
improved = True
path = [self.items[i] for i in best_route]
cost = 0
for i in range(1, len(path) - 1):
cost += get_distance(path[i], path[i - 1])
total_costs.append((cost, best_route))
total_costs = sorted(total_costs, key=lambda x: x[0])
self.optimal_cost, self.optimal_route = total_costs[0]
return self.optimal_route, self.optimal_cost, path
def arrange_trip(self, path):
p_range = []
sub_region = []
cpath = path.copy()
types = ['hotel', 'malls', 'restaurants', 'cafes', 'fast_food', 'architecture', 'cultural', 'sport', 'natural',
'marketplaces']
food_types = ['restaurants', 'fast_food']
poi_types = ['cultural', 'architecture']
market_types = ['malls', 'cafes', 'marketplaces']
entertainment_types = ['sport', 'natural']
orders = [['hotel'], food_types, random.choice(poi_types), random.choice(food_types),
random.choice(market_types)]
type_index = 0
while cpath:
src = cpath[0]
order = orders[type_index]
for p in path:
if p in cpath:
dist = get_distance(src, p)
if dist < 50:
# p_range.append((p, dist))
p_range.append(p)
cpath.remove(p)
else:
if len(orders) > type_index + 1:
type_index += 1
sub_region.append(p_range)
p_range = []
days = []
for sub in sub_region:
a = []
if len(sub) > 4:
for s in sub:
a.append(s)
if len(a) >= 5:
days.append(Day(len(days), a))
a = []
else:
days.append(Day(len(days), sub))
return days
# -
import pickle
with open('../../../testing/samples/paris_london_trip_data.pkl', 'rb') as input:
m_trip = pickle.load(input)
types = ['hotel', 'hotel','malls', 'restaurants', 'food', 'food', 'cafes', 'fast_food', 'architecture', 'cultural', 'sport', 'natural',
'marketplaces', 'food', 'historic']
# +
constraints = {
}
# -
from ortools.linear_solver import pywraplp
# +
from ortools.sat.python import cp_model
def SimpleSatProgram():
"""Minimal CP-SAT example to showcase calling the solver."""
# Creates the model.
# [START model]
model = cp_model.CpModel()
# [END model]
# Creates the variables.
# [START variables]
num_vals = 4
x = model.NewIntVar(0, num_vals - 1, 'x')
y = model.NewIntVar(0, num_vals - 1, 'y')
z = model.NewIntVar(0, num_vals - 1, 'z')
# [END variables]
# Creates the constraints.
# [START constraints]
model.Add(x != y)
# [END constraints]
# Creates a solver and solves the model.
# [START solve]
solver = cp_model.CpSolver()
status = solver.Solve(model)
# [END solve]
if status == cp_model.OPTIMAL:
print('x = %i' % solver.Value(x))
print('y = %i' % solver.Value(y))
print('z = %i' % solver.Value(z))
SimpleSatProgram()
# [END program]
# -
model = pywraplp.Solver.CreateSolver('GLOP')
x_morning = model.NumVar(0,1,'x_mor')
x_after = model.NumVar(0,5,'x_aft')
y_morning = model. NumVar(0,1,'y_mor')
y_after = model.NumVar(0,1,'y_aft')
# +
model.Add(x_morning <=1)
model.Add(x_after <=5)
model.Add(y_morning <=5)
model.Add(y_after <=1)
# -
model.Maximize(x_morning + 2*x_after + 4 *y_morning + 5 * y_after)
model.Solve()
model.Objective().Value()
x_after.solution_value()
x_morning.solution_value()
y_after.solution_value()
y_morning.solution_value()
solver = pywraplp.Solver.CreateSolver('GLOP')
xi = np.array([solver.NumVar(0,1,str(i)) for i in range(10)])
ci = np.array([np.random.randint(0,10) for i in range(10)])
pop = [np.random.randint(0,10) for i in range(10)]
interest = [np.random.randint(0,10) for i in range(10)]
solver.Add(np.sum([xi[i] for i in range(10)]) == 1)
solver.Add(np.sum([ci[i] * xi[i] for i in range(10)]) <= 10)
m = 6
solver.Maximize(np.sum([xi[i] * (m * interest[i] + (1-m) *pop[i]) for i in range(10)]))
solver.Solve()
solver.Objective().Value()
[(xi[i].solution_value()) for i in range(10)]
# +
model = cp_model.CpModel()
# -
riks_meus_morning = model.NewIntVar(0,1,'a')
riks_meus_after = model.NewIntVar(0,1,'b')
vangoug_meus_morning = model.NewIntVar(0,1,'c')
vangoug_meus_after = model.NewIntVar(0,1,'d')
model.Add(riks_meus_morning + riks_meus_after <= 1)
model.Add(vangoug_meus_morning + vangoug_meus_after <=1)
model.Add(riks_meus_morning +vangoug_meus_morning <=1)
model.Add(riks_meus_after + vangoug_meus_after <=1)
# model.Add(3*riks_meus_morning + 7* riks_meus_after + 2* vangoug_meus_morning + 4* vangoug_meus_after <= 50)
model.Maximize(9 * riks_meus_morning + 6 * riks_meus_after + 9 * vangoug_meus_morning + 6 * vangoug_meus_after)
# +
solver = cp_model.CpSolver()
status = solver.Solve(model)
if status == cp_model.OPTIMAL:
print('Minimum of objective function: %i' % solver.ObjectiveValue())
print()
print('riks_meus_morning value: ', solver.Value(riks_meus_morning))
print('riks_meus_after value: ', solver.Value(riks_meus_after))
print('vangoug_meus_morning value: ', solver.Value(vangoug_meus_morning))
print('vangoug_meus_after value: ', solver.Value(vangoug_meus_after))
else:
print('no Optimal Solution')
# -
| Search Engine/Trips-planning-system-main/search_engine/trip_planner/trip_classes/Planner_ILP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Emoji Prediction using Transfer Learning
#
# Understanding the sentiment of the sentence and then predicting an Emoji for it.
# ## Importing Libraries
# +
import numpy as np
import pandas as pd
import emoji
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Input, Dropout, LSTM, Activation
from tensorflow.keras.layers import Embedding
from tensorflow.keras.callbacks import ReduceLROnPlateau, ModelCheckpoint
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_score, recall_score
from matplotlib import pyplot as plt
import seaborn as sns
print ("Imports ready!")
# -
# ## Goal of the project
# The task is to build an Emojifier by using word vector representations. The model will take an input sentence and find the most appropriate emoji to be used with this sentence - from an assortment of 5 emoji's at its disposal.
#
# - 😁
# - 😓
# - 🖤
# - 🍴
# - ⚾
# ## Loading the dataset
# +
train = pd.read_csv ('dataset/train_emoji.csv', usecols = [0, 1], header = None)
test = pd.read_csv ('dataset/test_emoji.csv', usecols = [0, 1], header = None)
print (f'Training data shape = {train.shape}, Validation data shape = {test.shape}')
train.head (3)
# -
train[1].value_counts()
test.head (3)
# ## Data Preparation
# +
'''
Mapping label to a emoji
'''
emoji_dictionary = {"0": "\u2764\uFE0F", # 🖤
"1": ":baseball:", # ⚾
"2": ":beaming_face_with_smiling_eyes:", # 😁
"3": ":downcast_face_with_sweat:", # 😓
"4": ":fork_and_knife:", # 🍴
}
# +
'''
Getting x_train, y_train from train.csv
x_test, y_test from test.csv
'''
x_train = train.values[:, 0]
y_train = to_categorical (train.values[:, 1])
x_test = test.values[:, 0]
y_test = to_categorical (test.values[:, 1])
# +
maxLen = 0 # Len of longest sentence (by number of words)
for sent in x_train:
maxLen = max (maxLen, len (sent.split (' ')))
for sent in x_test:
maxLen = max (maxLen, len (sent.split (' ')))
print (f"Length of longest sentence (by number of words) is : {maxLen}")
# +
'''
Frst 10 training points
'''
for i, sent in enumerate (x_train):
if i == 10:
break
label = str (np.argmax (y_train[i]))
print (f"Sentence : {sent} , Emoji : {emoji.emojize (emoji_dictionary[label])}")
# -
# ## Creating word embeddings for the words that are present in the dataset
# I'll be using word vector representations of the words in the sentence so I need word vector representations of the words in the sentences. I'll use the Glove vectors for this representation.
#
# Based on few iterations 100 d vectors seem to work best for this case.
# +
'''
embeddings dictionary:
key = word
value = embedding vector [100 dimension vector]
'''
embeddings = {}
with open ('glove.6B.100d.txt', encoding = 'utf-8') as f:
for line in f:
values = line.split () # splits the word and coeff
word = values[0] # word
coeffs = np.asarray (values[1 : ], dtype = 'float32') # makes a word vector of len 50 for each word
embeddings[word] = coeffs
# -
def getOutputEmbeddings(X):
embedding_matrix_output = np.zeros ((X.shape[0], 10, 100)) # X.shape (num_of_sentences, max_len, embedding_dim)
for ix in range (X.shape[0]): # go to every sentence
X[ix] = X[ix].split () # get a list of words of the sentence
for jx in range (len(X[ix])): # go to every word
embedding_matrix_output[ix][jx] = embeddings[X[ix][jx].lower ()]
return embedding_matrix_output
# +
emb_x_train = getOutputEmbeddings(x_train) # getting embeddings for train data
emb_x_test = getOutputEmbeddings(x_test) # getting embeddings for test data
emb_x_train.shape, emb_x_test.shape
# -
# ## Model
model = Sequential()
model.add (LSTM (128, input_shape = (10, 100),return_sequences=True)) # to create a stacked LSTM model
model.add (Dropout (0.5)) # regularisation
model.add (LSTM (128, input_shape = (10, 100))) # stacking LSTM layer
model.add (Dropout (0.5))
model.add (Dense (5))
model.add (Activation ('softmax'))
model.summary()
# +
'''Callbacks'''
reduce_lr = ReduceLROnPlateau(monitor = 'loss', factor = 0.3, patience = 3, min_lr = 0.0001, verbose = 1)
checkpoint = ModelCheckpoint ("best_model.h5", monitor = 'val_acc', verbose = True, save_best_only = True)
callbacks = [reduce_lr, checkpoint]
# -
model.compile (optimizer = 'adam', loss = 'categorical_crossentropy', metrics = ['acc'])
hist = model.fit (emb_x_train, y_train, epochs = 25, batch_size = 16, shuffle = True, validation_split = 0.3, callbacks = callbacks)
# ## Model Accuracy
model.load_weights ('best_model.h5')
model.evaluate (emb_x_test, y_test)
# ## Checking the predictions
pred = model.predict_classes(emb_x_test)
print("Sentence : Actual Prediction")
for i in range(10):
print(' '.join(x_test[i]), end = " : ")
print(emoji.emojize(emoji_dictionary[str(np.argmax(y_test[i]))]), end = " \t\t")
print(emoji.emojize(emoji_dictionary[str(pred[i])]))
# I am upset : ❤️ 😓
# We had such a lovely dinner tonight : ❤️ 😁
#
# for these sentences our predictions were also good.
# +
'''
Confusion Matrix
'''
Y_test = [np.argmax (i) for i in y_test]
cm = confusion_matrix(Y_test, pred)
# -
sns.heatmap(cm, annot = True)
plt.show ()
# ## Tasks for future
# - Get more data
# - Try different model architectures
# ________
| EmojiPred.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9 (tensorflow)
# language: python
# name: tensorflow
# ---
# <a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_02_4_pandas_functional.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # T81-558: Applications of Deep Neural Networks
# **Module 2: Python for Machine Learning**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Module 2 Material
#
# Main video lecture:
#
# * Part 2.1: Introduction to Pandas [[Video]](https://www.youtube.com/watch?v=bN4UuCBdpZc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_02_1_python_pandas.ipynb)
# * Part 2.2: Categorical Values [[Video]](https://www.youtube.com/watch?v=4a1odDpG0Ho&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_02_2_pandas_cat.ipynb)
# * Part 2.3: Grouping, Sorting, and Shuffling in Python Pandas [[Video]](https://www.youtube.com/watch?v=YS4wm5gD8DM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_02_3_pandas_grouping.ipynb)
# * **Part 2.4: Using Apply and Map in Pandas for Keras** [[Video]](https://www.youtube.com/watch?v=XNCEZ4WaPBY&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_4_pandas_functional.ipynb)
# * Part 2.5: Feature Engineering in Pandas for Deep Learning in Keras [[Video]](https://www.youtube.com/watch?v=BWPTj4_Mi9E&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_02_5_pandas_features.ipynb)
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow.
try:
# %tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
# # Part 2.4: Apply and Map
# If you've ever worked with Big Data or functional programming languages before, you've likely heard of map/reduce. Map and reduce are two functions that apply a task that you create to a data frame. Pandas supports functional programming techniques that allow you to use functions across en entire data frame. In addition to functions that you write, Pandas also provides several standard functions for use with data frames.
# ### Using Map with Dataframes
#
# The map function allows you to transform a column by mapping certain values in that column to other values. Consider the Auto MPG data set that contains a field **origin_name** that holds a value between one and three that indicates the geographic origin of each car. We can see how to use the map function to transform this numeric origin into the textual name of each origin.
#
# We will begin by loading the Auto MPG data set.
# +
import os
import pandas as pd
import numpy as np
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA', '?'])
pd.set_option('display.max_columns', 7)
pd.set_option('display.max_rows', 5)
display(df)
# -
# The **map** method in Pandas operates on a single column. You provide **map** with a dictionary of values to transform the target column. The map keys specify what values in the target column should be turned into values specified by those keys. The following code shows how the map function can transform the numeric values of 1, 2, and 3 into the string values of North America, Europe and Asia.
# +
# Apply the map
df['origin_name'] = df['origin'].map(
{1: 'North America', 2: 'Europe', 3: 'Asia'})
# Shuffle the data, so that we hopefully see
# more regions.
df = df.reindex(np.random.permutation(df.index))
# Display
pd.set_option('display.max_columns', 7)
pd.set_option('display.max_rows', 10)
display(df)
# -
# ### Using Apply with Dataframes
#
# The **apply** function of the data frame can run a function over the entire data frame. You can use either be a traditional named function or a lambda function. Python will execute the provided function against each of the rows or columns in the data frame. The **axis** parameter specifies of the function is run across rows or columns. For axis = 1, rows are used. The following code calculates a series called **efficiency** that is the **displacement** divided by **horsepower**.
efficiency = df.apply(lambda x: x['displacement']/x['horsepower'], axis=1)
display(efficiency[0:10])
# You can now insert this series into the data frame, either as a new column or to replace an existing column. The following code inserts this new series into the data frame.
df['efficiency'] = efficiency
# ### Feature Engineering with Apply and Map
# In this section, we will see how to calculate a complex feature using map, apply, and grouping. The data set is the following CSV:
#
# * https://www.irs.gov/pub/irs-soi/16zpallagi.csv
#
# This URL contains US Government public data for "SOI Tax Stats - Individual Income Tax Statistics." The entry point to the website is here:
#
# * https://www.irs.gov/statistics/soi-tax-stats-individual-income-tax-statistics-2016-zip-code-data-soi
#
# Documentation describing this data is at the above link.
#
# For this feature, we will attempt to estimate the adjusted gross income (AGI) for each of the zip codes. The data file contains many columns; however, you will only use the following:
#
# * **STATE** - The state (e.g., MO)
# * **zipcode** - The zipcode (e.g. 63017)
# * **agi_stub** - Six different brackets of annual income (1 through 6)
# * **N1** - The number of tax returns for each of the agi_stubs
#
# Note, the file will have six rows for each zip code, for each of the agi_stub brackets. You can skip zip codes with 0 or 99999.
#
# We will create an output CSV with these columns; however, only one row per zip code. Calculate a weighted average of the income brackets. For example, the following six rows are present for 63017:
#
#
# |zipcode |agi_stub | N1 |
# |--|--|-- |
# |63017 |1 | 4710 |
# |63017 |2 | 2780 |
# |63017 |3 | 2130 |
# |63017 |4 | 2010 |
# |63017 |5 | 5240 |
# |63017 |6 | 3510 |
#
#
# We must combine these six rows into one. For privacy reasons, AGI's are broken out into 6 buckets. We need to combine the buckets and estimate the actual AGI of a zipcode. To do this, consider the values for N1:
#
# * 1 = 1 to 25,000
# * 2 = 25,000 to 50,000
# * 3 = 50,000 to 75,000
# * 4 = 75,000 to 100,000
# * 5 = 100,000 to 200,000
# * 6 = 200,000 or more
#
# The median of each of these ranges is approximately:
#
# * 1 = 12,500
# * 2 = 37,500
# * 3 = 62,500
# * 4 = 87,500
# * 5 = 112,500
# * 6 = 212,500
#
# Using this you can estimate 63017's average AGI as:
#
# ```
# >>> totalCount = 4710 + 2780 + 2130 + 2010 + 5240 + 3510
# >>> totalAGI = 4710 * 12500 + 2780 * 37500 + 2130 * 62500
# + 2010 * 87500 + 5240 * 112500 + 3510 * 212500
# >>> print(totalAGI / totalCount)
#
# 88689.89205103042
# ```
#
# We begin by reading in the government data.
# +
import pandas as pd
df=pd.read_csv('https://www.irs.gov/pub/irs-soi/16zpallagi.csv')
# -
# First, we trim all zip codes that are either 0 or 99999. We also select the three fields that we need.
# +
df=df.loc[(df['zipcode']!=0) & (df['zipcode']!=99999),
['STATE','zipcode','agi_stub','N1']]
pd.set_option('display.max_columns', 0)
pd.set_option('display.max_rows', 10)
display(df)
# -
# We replace all of the **agi_stub** values with the correct median values with the **map** function.
# +
medians = {1:12500,2:37500,3:62500,4:87500,5:112500,6:212500}
df['agi_stub']=df.agi_stub.map(medians)
pd.set_option('display.max_columns', 0)
pd.set_option('display.max_rows', 10)
display(df)
# -
# Next, we group the data frame by zip code.
groups = df.groupby(by='zipcode')
# The program applies a lambda is applied across the groups, and then calculates the AGI estimate.
df = pd.DataFrame(groups.apply(
lambda x:sum(x['N1']*x['agi_stub'])/sum(x['N1']))) \
.reset_index()
# +
pd.set_option('display.max_columns', 0)
pd.set_option('display.max_rows', 10)
display(df)
# -
# We can now rename the new agi_estimate column.
df.columns = ['zipcode','agi_estimate']
# +
pd.set_option('display.max_columns', 0)
pd.set_option('display.max_rows', 10)
display(df)
# -
# Finally, we check to see that our zip code of 63017 got the correct value.
df[ df['zipcode']==63017 ]
| t81_558_class_02_4_pandas_functional.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Libraries
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import tensorflow as tf
SEED = 2018
np.random.seed(SEED)
tf.set_random_seed(SEED)
from tqdm import tqdm
tqdm.pandas()
import os
print(os.listdir("../input"))
# +
import zipfile
z= zipfile.ZipFile('../input/embeddings.zip')
z.extractall()
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
from tensorflow.keras.layers import *
from tensorflow.keras.models import *
from tensorflow.keras.optimizers import *
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau
from tensorflow.keras import backend as K
#from keras.engine.topology import Layer
from tensorflow.keras.layers import Layer
from tensorflow.keras import initializers, regularizers, constraints, optimizers, layers
import gc, re
from sklearn import metrics
from sklearn.model_selection import GridSearchCV, StratifiedKFold
from sklearn.metrics import f1_score, roc_auc_score
# -
# # Data Loading/ Cleaning
# + _uuid="90b5e8186e3c5dfaa16dbce540331c49dcc11801"
def clean_text(x):
x = str(x)
for punct in "/-'":
x = x.replace(punct, ' ')
for punct in '&':
x = x.replace(punct, f' {punct} ')
for punct in '?!.,"#$%\'()*+-/:;<=>@[\\]^_`{|}~' + '“”’':
x = x.replace(punct, '')
return x
def clean_numbers(x):
x = re.sub('[0-9]{5,}', '#####', x)
x = re.sub('[0-9]{4}', '####', x)
x = re.sub('[0-9]{3}', '###', x)
x = re.sub('[0-9]{2}', '##', x)
return x
def _get_mispell(mispell_dict):
mispell_re = re.compile('(%s)' % '|'.join(mispell_dict.keys()))
return mispell_dict, mispell_re
mispell_dict = {'colour':'color',
'centre':'center',
'didnt':'did not',
'doesnt':'does not',
'isnt':'is not',
'shouldnt':'should not',
'favourite':'favorite',
'travelling':'traveling',
'counselling':'counseling',
'theatre':'theater',
'cancelled':'canceled',
'labour':'labor',
'organisation':'organization',
'wwii':'world war 2',
'citicise':'criticize',
'instagram': 'social medium',
'whatsapp': 'social medium',
'snapchat': 'social medium'
}
mispellings, mispellings_re = _get_mispell(mispell_dict)
def replace_typical_misspell(text):
def replace(match):
return mispellings[match.group(0)]
return mispellings_re.sub(replace, text)
# -
# # Data Loading / Preprocessing
# + _uuid="417fc7b1456a0340dbbf7b5afe44432ed7a0f5d9"
def load_and_preprocess_data(max_features=50000, maxlen=70):
train_df = pd.read_csv("../input/train.csv")
test_df = pd.read_csv("../input/test.csv")
print("Train shape : ",train_df.shape)
print("Test shape : ",test_df.shape)
# Clean the text
train_df["question_text"] = train_df["question_text"].progress_apply(lambda x: clean_text(x))
test_df["question_text"] = test_df["question_text"].progress_apply(lambda x: clean_text(x))
# Clean numbers
train_df["question_text"] = train_df["question_text"].progress_apply(lambda x: clean_numbers(x))
test_df["question_text"] = test_df["question_text"].progress_apply(lambda x: clean_numbers(x))
# Clean speelings
train_df["question_text"] = train_df["question_text"].progress_apply(lambda x: replace_typical_misspell(x))
test_df["question_text"] = test_df["question_text"].progress_apply(lambda x: replace_typical_misspell(x))
## fill up the missing values
train_X = train_df["question_text"].fillna("_##_").values
test_X = test_df["question_text"].fillna("_##_").values
## Tokenize the sentences
tokenizer = Tokenizer(num_words=max_features)
tokenizer.fit_on_texts(list(train_X))
train_X = tokenizer.texts_to_sequences(train_X)
test_X = tokenizer.texts_to_sequences(test_X)
## Pad the sentences
train_X = pad_sequences(train_X, maxlen=maxlen)
test_X = pad_sequences(test_X, maxlen=maxlen)
## Get the target values
train_y = train_df['target'].values
#shuffling the data
np.random.seed(2018)
trn_idx = np.random.permutation(len(train_X))
train_X = train_X[trn_idx]
train_y = train_y[trn_idx]
return train_X, test_X, train_y, tokenizer.word_index
# + _uuid="63e672daf4c1baa189fb60d0585ddd55da210fc8"
def load_glove(word_index):
EMBEDDING_FILE = './glove.840B.300d/glove.840B.300d.txt'
def get_coefs(word,*arr): return word, np.asarray(arr, dtype='float32')
embeddings_index = dict(get_coefs(*o.split(" ")) for o in open(EMBEDDING_FILE))
all_embs = np.stack(embeddings_index.values())
emb_mean,emb_std = all_embs.mean(), all_embs.std()
embed_size = all_embs.shape[1]
# word_index = tokenizer.word_index
nb_words = min(max_features, len(word_index))
embedding_matrix = np.random.normal(emb_mean, emb_std, (nb_words, embed_size))
for word, i in word_index.items():
if i >= max_features: continue
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None: embedding_matrix[i] = embedding_vector
return embedding_matrix
def load_fasttext(word_index):
EMBEDDING_FILE = './wiki-news-300d-1M/wiki-news-300d-1M.vec'
def get_coefs(word,*arr): return word, np.asarray(arr, dtype='float32')
embeddings_index = dict(get_coefs(*o.split(" ")) for o in open(EMBEDDING_FILE) if len(o)>100)
all_embs = np.stack(embeddings_index.values())
emb_mean,emb_std = all_embs.mean(), all_embs.std()
embed_size = all_embs.shape[1]
# word_index = tokenizer.word_index
nb_words = min(max_features, len(word_index))
embedding_matrix = np.random.normal(emb_mean, emb_std, (nb_words, embed_size))
for word, i in word_index.items():
if i >= max_features: continue
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None: embedding_matrix[i] = embedding_vector
return embedding_matrix
def load_para(word_index):
EMBEDDING_FILE = './paragram_300_sl999/paragram_300_sl999.txt'
def get_coefs(word,*arr): return word, np.asarray(arr, dtype='float32')
embeddings_index = dict(get_coefs(*o.split(" ")) for o in open(EMBEDDING_FILE, encoding="utf8", errors='ignore') if len(o)>100)
all_embs = np.stack(embeddings_index.values())
emb_mean,emb_std = all_embs.mean(), all_embs.std()
embed_size = all_embs.shape[1]
# word_index = tokenizer.word_index
nb_words = min(max_features, len(word_index))
embedding_matrix = np.random.normal(emb_mean, emb_std, (nb_words, embed_size))
for word, i in word_index.items():
if i >= max_features: continue
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None: embedding_matrix[i] = embedding_vector
return embedding_matrix
# + _uuid="2aa88090981b6310b6644a0f3acc6dbd2b3d9741"
class Attention(Layer):
def __init__(self, step_dim,
W_regularizer=None, b_regularizer=None,
W_constraint=None, b_constraint=None,
bias=True, **kwargs):
self.supports_masking = True
self.init = initializers.get('glorot_uniform')
self.W_regularizer = regularizers.get(W_regularizer)
self.b_regularizer = regularizers.get(b_regularizer)
self.W_constraint = constraints.get(W_constraint)
self.b_constraint = constraints.get(b_constraint)
self.bias = bias
self.step_dim = step_dim
self.features_dim = 0
super(Attention, self).__init__(**kwargs)
def build(self, input_shape):
assert len(input_shape) == 3
self.W = self.add_weight('{}_W'.format(self.name), shape=(input_shape[-1].value,),
initializer=self.init,
regularizer=self.W_regularizer,
constraint=self.W_constraint)
self.features_dim = input_shape[-1]
if self.bias:
self.b = self.add_weight('{}_b'.format(self.name), shape=(input_shape[1].value,),
initializer='zero',
regularizer=self.b_regularizer,
constraint=self.b_constraint
)
else:
self.b = None
self.built = True
def compute_mask(self, input, input_mask=None):
return None
def call(self, x, mask=None):
features_dim = self.features_dim
step_dim = self.step_dim
eij = K.reshape(K.dot(K.reshape(x, (-1, features_dim)),
K.reshape(self.W, (features_dim, 1))), (-1, step_dim))
if self.bias:
eij += self.b
eij = K.tanh(eij)
a = K.exp(eij)
if mask is not None:
a *= K.cast(mask, K.floatx())
a /= K.cast(K.sum(a, axis=1, keepdims=True) + K.epsilon(), K.floatx())
a = K.expand_dims(a)
weighted_input = x * a
return K.sum(weighted_input, axis=1)
def compute_output_shape(self, input_shape):
return input_shape[0], self.features_dim
# + _uuid="79fcb1c003aca5f86fa0dbb10ce1d08c6506231f"
def model_lstm_attention(embedding_matrix, maxlen=70, max_features=50000, units=64):
inp = Input(shape=(maxlen,))
x = Embedding(max_features, embed_size, weights=[embedding_matrix], trainable=False)(inp)
x = Bidirectional(CuDNNLSTM(units*2, return_sequences=True))(x)
x = Bidirectional(CuDNNLSTM(units, return_sequences=True))(x)
x = Attention(maxlen)(x)
x = Dense(units, activation='relu')(x)
outp= Dense(1, activation='sigmoid')(x)
model = Model(inputs=inp, outputs=outp)
model.compile(loss='binary_crossentropy', optimizer=Adam(), metrics=['accuracy'])
return model
def model_gru_attention(embedding_matrix, maxlen=70, max_features=50000, units=64):
inp = Input(shape=(maxlen,))
x = Embedding(max_features, embed_size, weights=[embedding_matrix], trainable=False)(inp)
x = Bidirectional(CuDNNGRU(units*2, return_sequences=True))(x)
x = Bidirectional(CuDNNGRU(units, return_sequences=True))(x)
x = Attention(maxlen)(x)
x = Dense(units, activation='relu')(x)
outp= Dense(1, activation='sigmoid')(x)
model = Model(inputs=inp, outputs=outp)
model.compile(loss='binary_crossentropy', optimizer=Adam(), metrics=['accuracy'])
return model
def model_gru_attention_3(embedding_matrix, maxlen=70, max_features=50000, units=64):
inp = Input(shape=(maxlen,))
x = Embedding(max_features, embed_size, weights=[embedding_matrix], trainable=False)(inp)
x = Bidirectional(CuDNNGRU(units*2, kernel_initializer=initializers.glorot_uniform(seed = SEED), return_sequences=True))(x)
x = Bidirectional(CuDNNGRU(units, kernel_initializer=initializers.glorot_uniform(seed = SEED), return_sequences=True))(x)
x = Bidirectional(CuDNNGRU(units//2, kernel_initializer=initializers.glorot_uniform(seed = SEED), return_sequences=True))(x)
x = Attention(maxlen)(x)
outp= Dense(1, kernel_initializer=initializers.he_uniform(seed=SEED), activation='sigmoid')(x)
model = Model(inputs=inp, outputs=outp)
model.compile(loss='binary_crossentropy', optimizer=Adam(), metrics=['accuracy'])
return model
# + _uuid="65d008ef0b60ac644eb55cc0fc8a726bf50f5414"
def train_pred(model, train_X, train_y, val_X, val_y, epochs=2):
for e in range(epochs):
model.fit(train_X, train_y, batch_size=512, epochs=1, validation_data=(val_X, val_y))
pred_val_y = model.predict([val_X], batch_size=1024, verbose=0)
best_thresh = 0.5
best_score = 0.0
for thresh in np.arange(0.1, 0.501, 0.01):
thresh = np.round(thresh, 2)
score = metrics.f1_score(val_y, (pred_val_y > thresh).astype(int))
if score > best_score:
best_thresh = thresh
best_score = score
print("Val F1 Score: {:.4f}".format(best_score))
pred_test_y = model.predict([test_X], batch_size=1024, verbose=0)
print('='*100)
return pred_val_y, pred_test_y, best_score
# + _uuid="16377fced6017ddc0d40e12f71da9f37897161cb"
embed_size = 300
max_features = 100000
maxlen = 40
# + _uuid="9b370506e3512111916c3f7c012f1a6a145e6470"
import time
start_time = time.time()
train_X, test_X, train_y, word_index = load_and_preprocess_data(max_features=max_features, maxlen=maxlen)
total_time = (time.time() - start_time)/60.0
print("Took {0} minutes".format(total_time))
# + _uuid="58d920c9d05e253a473099a5a0ee4328f5b3dd71"
import time
start_time = time.time()
embedding_matrix_1 = load_glove(word_index)
#embedding_matrix_2 = load_fasttext(word_index)
embedding_matrix_3 = load_para(word_index)
total_time = (time.time() - start_time)/60.0
print("Took {0} minutes".format(total_time))
# + _uuid="730f4de3b8bc15b84adda90268eaeac44bac28c2"
embedding_matrix = np.mean([embedding_matrix_1, embedding_matrix_3], axis = 0)
print(np.shape(embedding_matrix))
# + _uuid="dab72672c52e429f4687c033cfd84d4ce7b1c525"
def threshold_search(y_true, y_proba):
best_threshold = 0
best_score = 0
for threshold in [i * 0.01 for i in range(100)]:
score = f1_score(y_true=y_true, y_pred=y_proba > threshold)
if score > best_score:
best_threshold = threshold
best_score = score
search_result = {'threshold': best_threshold, 'f1': best_score}
return search_result
# + _uuid="020dc129d1aa9013bdf6a3d499819190f3edba3d"
train_meta = np.zeros(train_y.shape)
test_meta = np.zeros(test_X.shape[0])
splits = list(StratifiedKFold(n_splits=4, shuffle=True, random_state=SEED).split(train_X, train_y))
for idx, (train_idx, valid_idx) in enumerate(splits):
X_train = train_X[train_idx]
y_train = train_y[train_idx]
X_val = train_X[valid_idx]
y_val = train_y[valid_idx]
tmp_result = []
model = model_gru_attention_3(embedding_matrix, maxlen=maxlen, max_features=max_features, units=64)
pred_val_y, pred_test_y, best_score = train_pred(model, X_train, y_train, X_val, y_val, epochs = 2)
tmp_result.append([pred_val_y, pred_test_y, best_score])
model = model_gru_attention(embedding_matrix, maxlen=maxlen, max_features=max_features, units=64)
pred_val_y, pred_test_y, best_score = train_pred(model, X_train, y_train, X_val, y_val, epochs = 2)
tmp_result.append([pred_val_y, pred_test_y, best_score])
model = model_lstm_attention(embedding_matrix_1, maxlen=maxlen, max_features=max_features, units=64)
pred_val_y, pred_test_y, best_score = train_pred(model, X_train, y_train, X_val, y_val, epochs = 2)
tmp_result.append([pred_val_y, pred_test_y, best_score])
pred_val_y, pred_test_y, best_score = np.mean(tmp_result, axis=0)
print("Mean Result: ", best_score)
train_meta[valid_idx] = pred_val_y.reshape(-1)
test_meta += pred_test_y.reshape(-1) / len(splits)
# + _uuid="7cced03c9fa80299dd7d2ca67bfe7966b210113b"
search_result = threshold_search(train_y, train_meta)
print(search_result)
sub = pd.read_csv('../input/sample_submission.csv')
sub.prediction = test_meta > search_result['threshold']
sub.to_csv("submission.csv", index=False)
| 28-12-Quora-Project/quora-gru (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from autumn.tools.runs import ManagedRun
mr = ManagedRun("sm_sir/philippines/1637190072/808413a")
mr.calibration.download_outputs()
do_df = mr.full_run.get_derived_outputs()
pbi = mr.powerbi.get_db()
pbi.get_uncertainty()['incidence'].plot()
from autumn.tools.runs.utils import collate_columns_to_urun
from numpy.random import choice
import pandas as pd
collated = collate_columns_to_urun(do_df)
uruns = choice(collated.urun.unique(), 50, False)
for urun in uruns:
filtered = collated[collated['urun']==urun]
s = pd.Series(filtered['incidence'].to_numpy(), index=filtered['times'])
s.plot()
| notebooks/user/dshipman/CalibrationPlotOneOff.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="iQ0SsHfUtj8d"
# <h1><strong>Análise de sentimentos em avaliações de e-commerce</strong></h1>
#
#
# **Objetivo**:
#
# Classificar as avaliações de e-commerce em positiva ou negativa.
#
# * Dataset: https://nijianmo.github.io/amazon/index.html
#
# ***Orientador:*** <NAME>
#
# ***Orientando:*** <NAME>
# + [markdown] id="3VDVe7I2ajbK"
# # Importando as bibliotecas
# + id="wfFEXZC0WS-V" executionInfo={"status": "ok", "timestamp": 1645490245076, "user_tz": 240, "elapsed": 2226, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
import numpy as np
import pandas as pd
import os
import joblib
import re
import nltk
from nltk.tokenize import sent_tokenize, word_tokenize
from nltk.stem.porter import PorterStemmer
import gzip
import json
# from textblob import TextBlob
# from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
# from flair.models import TextClassifier
# from flair.data import Sentence
# import autosklearn.classification
from tpot import TPOTClassifier
#import autogluon as ag
#from autogluon.core.utils.loaders.load_pd import load
#from autogluon.text import TextPredictor
#from autogluon.tabular import TabularDataset, TabularPredictor
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, accuracy_score, classification_report
# + [markdown] id="U6RfC6N1vqup"
# # Importando os *datasets*
# + id="do94YizuROsb" executionInfo={"status": "ok", "timestamp": 1645490245080, "user_tz": 240, "elapsed": 1, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
def parse(path):
g = gzip.open(path, 'rb')
for l in g:
yield json.loads(l)
def getDF(path):
i = 0
df = {}
for d in parse(path):
df[i] = d
i += 1
if i >= 20000:
break
return pd.DataFrame.from_dict(df, orient='index')
# + id="c7zwVW_Bvv91" executionInfo={"status": "ok", "timestamp": 1645490245128, "user_tz": 240, "elapsed": 47, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
datasets = {}
datasets['Amazon Fashion'] = getDF('/content/drive/MyDrive/<NAME>/dataset/AMAZON_FASHION_5.json.gz')
#datasets['All Beauty'] = getDF('/content/drive/MyDrive/<NAME>/dataset/All_Beauty_5.json.gz')
#datasets['Appliances'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Appliances_5.json.gz')
#datasets['Arts, Crafts and Sewing'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Arts_Crafts_and_Sewing_5.json.gz')
#datasets['Automotive'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Automotive_5.json.gz')
#datasets['Books'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Books_5.json.gz')
#datasets['CDs and Vinyl'] = getDF('/content/drive/MyDrive/<NAME>/dataset/CDs_and_Vinyl_5.json.gz')
#datasets['Cell Phones and Accessories'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Cell_Phones_and_Accessories_5.json.gz')
#datasets['Clothing, Shoes and Jewelry'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Clothing_Shoes_and_Jewelry_5.json.gz')
#datasets['Digital Music'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Digital_Music_5.json.gz')
#datasets['Electronics'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Electronics_5.json.gz')
#datasets['Gift Cards'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Gift_Cards_5.json.gz')
#datasets['Grocery and Gourmet Food'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Grocery_and_Gourmet_Food_5.json.gz')
#datasets['Home and Kitchen'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Home_and_Kitchen_5.json.gz')
#datasets['Industrial and Scientific'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Industrial_and_Scientific_5.json.gz')
#datasets['Kindle Store'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Kindle_Store_5.json.gz')
#datasets['Luxury Beauty'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Luxury_Beauty_5.json.gz')
#datasets['Magazine Subscriptions'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Magazine_Subscriptions_5.json.gz')
#datasets['Movies and TV'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Movies_and_TV_5.json.gz')
#datasets['Musical Instruments'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Musical_Instruments_5.json.gz')
#datasets['Office Products'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Office_Products_5.json.gz')
#datasets['Patio, Lawn and Garden'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Patio_Lawn_and_Garden_5.json.gz')
#datasets['Pet Supplies'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Pet_Supplies_5.json.gz')
#datasets['Prime Pantry'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Prime_Pantry_5.json.gz')
#datasets['Software'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Software_5.json.gz')
#datasets['Sports and Outdoors'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Sports_and_Outdoors_5.json.gz')
#datasets['Tools and Home Improvement'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Tools_and_Home_Improvement_5.json.gz')
#datasets['Toys and Games'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Toys_and_Games_5.json.gz')
#datasets['Video Games'] = getDF('/content/drive/MyDrive/<NAME>/dataset/Video_Games_5.json.gz')
# + [markdown] id="q6FrJ-kHTLGz"
# # Visualização dos *datasets*
# + id="XmJaNSzL49-l" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1645490245132, "user_tz": 240, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}} outputId="8cd637b3-cbe9-4367-b2fe-52bd053aaff1"
for key, dataset in datasets.items():
print(key)
dataset.info()
print('\n')
# + [markdown] id="TI5TF5Rzd8Rt"
# # Modelagem
# + [markdown] id="xu77a2Uw6PdL"
# ## Seleciona as colunas '*reviewText*' e '*overall*'
# + id="8D2AzJyAZibh" executionInfo={"status": "ok", "timestamp": 1645490245134, "user_tz": 240, "elapsed": 1, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
def select_columns(dataset):
dataset = dataset[['reviewText', 'overall']]
dataset = dataset.dropna()
dataset = dataset.reset_index(drop=True)
return dataset
# + id="704NLyW26PzS" executionInfo={"status": "ok", "timestamp": 1645490245144, "user_tz": 240, "elapsed": 9, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
for key, dataset in datasets.items():
dataset= select_columns(dataset)
datasets[key] = dataset
# + [markdown] id="HHSjAcip7b3J"
# ## Encoding
# + [markdown] id="GRcTs1OPA6ts"
# Seleciona o atributo *'overall'* e o converte da seguinte forma:
#
#
#
# * 1, 2 e 3: negativa
# * 4 e 5: positiva
# + id="4ASCe60r9_rt" executionInfo={"status": "ok", "timestamp": 1645490245146, "user_tz": 240, "elapsed": 1, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
for key, dataset in datasets.items():
dataset['overall'] = np.where(dataset['overall'] >= 4.0, 1, 0)
datasets[key] = dataset
# + [markdown] id="qaek5UNVu36o"
# ## Limpando as avaliações
# + id="zOLyUmteD2iq" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1645490245152, "user_tz": 240, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}} outputId="f9d4dc7d-c495-4d5d-a448-f75f131296ff"
english_stopwords = list()
with open('/content/drive/MyDrive/Edson Henrique/code/stopwords/english', 'r') as a_file:
for line in a_file:
stripped_line = line.strip()
english_stopwords.append(stripped_line)
excluded_words = ("not", "no", 'aren', "aren't", 'couldn', "couldn't", 'didn', "didn't", 'doesn', "doesn't", 'hadn', "hadn't", 'hasn', "hasn't", 'haven', "haven't", 'isn', "isn't", 'shouldn', "shouldn't", "won't", 'wouldn', "wouldn't")
english_stopwords = [x for x in english_stopwords if x not in excluded_words]
print(english_stopwords)
# + id="9yuwtrWIEnN0" executionInfo={"status": "ok", "timestamp": 1645490245159, "user_tz": 240, "elapsed": 1, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
def limpa_texto(dataset):
corpus = []
for i in range (0, dataset.shape[0]):
review = re.sub('[^a-zA-Z]', ' ', dataset['reviewText'][i])
review = review.lower()
review = review.split()
ps = PorterStemmer()
review = [ps.stem(word) for word in review if not word in set(english_stopwords)]
review = ' '.join(review)
corpus.append(review)
return corpus
# + [markdown] id="M1U7mvWrubZJ"
# ## Criando o modelo *bag of words*
# + id="9HM2b8eJuZH_" executionInfo={"status": "ok", "timestamp": 1645490245162, "user_tz": 240, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
def cria_bag_words(corpus):
cv = CountVectorizer(max_features = 1500)
X = cv.fit_transform(corpus).toarray()
return X
# + [markdown] id="VisX3nMolBln"
# ## Reduzindo o tamanho dos dados
# + id="d56-61ORlBLi" executionInfo={"status": "ok", "timestamp": 1645490245163, "user_tz": 240, "elapsed": 0, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
for key, dataset in datasets.items():
dataset['overall'] = dataset['overall'].astype(np.uint8)
# + colab={"base_uri": "https://localhost:8080/"} id="Flwt4GMPlbwN" executionInfo={"status": "ok", "timestamp": 1645490245168, "user_tz": 240, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}} outputId="f068ade5-e316-4d91-d35e-a8aaa3de7509"
for key, dataset in datasets.items():
print(key)
dataset.info()
print('\n')
# + [markdown] id="OmLuXzpgAUOB"
# # Treino
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Oi7t5MyaT34I" executionInfo={"status": "ok", "timestamp": 1645490245177, "user_tz": 240, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}} outputId="9bad4c9e-b512-4e43-ff4f-0d300ec5ccfc"
def ler_dataset_resultados():
dir = '/content/drive/MyDrive/Edson Henrique/code/'
filename = 'resultados.csv'
if True == os.path.isfile(dir + filename):
# load results dataset from disk
resultados_dataset = pd.read_csv(dir + filename, index_col=0)
else:
result_dict = {'TextBlob': np.nan, 'Vader' : np.nan, 'Flair' : np.nan, 'AutoGluon' : np.nan, 'Auto-Sklearn' : np.nan, 'TPOT' : np.nan}
resultados_dataset = pd.DataFrame(result_dict, index = datasets.keys())
# save results dataset to disk
resultados_dataset.to_csv(dir + filename, index=True)
return resultados_dataset
def salvar_dataset_resultados(dataset):
dir = '/content/drive/MyDrive/Edson Henrique/code/'
filename = 'resultados.csv'
resultados_dataset.to_csv(dir + filename, index=True)
resultados_dataset = ler_dataset_resultados()
resultados_dataset
# + [markdown] id="4CLDc9slhOZX"
# ## TextBlob
# + id="bVNZgcJWt2Xm" executionInfo={"status": "ok", "timestamp": 1645490245180, "user_tz": 240, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# f_categorizar_blob = (lambda x: 1 if x > 0 else 0)
# def classificar_blob(dataset):
# dataset['TextBlob'] = np.nan
# for i in range (0, dataset.shape[0]):
# blob = TextBlob(dataset['reviewText'].values[i])
# dataset.iloc[i,2] = f_categorizar_blob(blob.sentiment.polarity)
# return dataset
# + id="BSKPPN6OjDn3" executionInfo={"status": "ok", "timestamp": 1645490245183, "user_tz": 240, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# dir = '/content/drive/MyDrive/Edson Henrique/model_save/'
# for key, dataset in datasets.items():
# filename = key + '_TextBlob.csv'
# if True == os.path.isfile(dir + filename):
# # load dataset from disk
# dataset = pd.read_csv(dir + filename)
# else:
# dataset = classificar_blob(dataset)
# # save new dataset to disk
# dataset.to_csv(dir + filename)
# datasets[key] = dataset
# y_true = dataset['overall'].values
# y_pred = dataset['TextBlob'].values
# print('Classificador TextBlob')
# print('Dataset: ', key)
# print('Matriz de confusão')
# print(confusion_matrix(y_true, y_pred))
# print('\nMétricas')
# print(classification_report(y_true, y_pred, digits=3))
# print('\nScore')
# score = accuracy_score(y_true, y_pred)
# print(score)
# print('\n')
# if pd.isna(resultados_dataset.at[key, 'TextBlob']):
# resultados_dataset.loc[key]['TextBlob'] = score
# salvar_dataset_resultados(resultados_dataset)
# + [markdown] id="RdC4MihfhX28"
# ## vaderSentiment
# + id="TNNFGxmhxwEl" executionInfo={"status": "ok", "timestamp": 1645490245184, "user_tz": 240, "elapsed": 1, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# f_categorizar_vader = (lambda x: 1 if x > 0 else 0)
# def classificar_vaderSentiment(dataset):
# dataset['vaderSentiment'] = np.nan
# analyzer = SentimentIntensityAnalyzer()
# for i in range (0, dataset.shape[0]):
# vs = analyzer.polarity_scores(dataset['reviewText'].values[i])
# dataset.iloc[i,3] = f_categorizar_vader(vs['compound'])
# return dataset
# + id="wyPs3b4pxwEm" executionInfo={"status": "ok", "timestamp": 1645490245193, "user_tz": 240, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# dir = '/content/drive/MyDrive/<NAME>/model_save/'
# for key, dataset in datasets.items():
# filename = key + '_vaderSentiment.csv'
# if True == os.path.isfile(dir + filename):
# # load dataset from disk
# dataset = pd.read_csv(dir + filename)
# else:
# dataset = classificar_vaderSentiment(dataset)
# # save new dataset to disk
# dataset.to_csv(dir + filename)
# datasets[key] = dataset
# y_true = dataset['overall'].values
# y_pred = dataset['vaderSentiment'].values
# print('Classificador vaderSentiment')
# print('Dataset: ', key)
# print('Matriz de confusão')
# print(confusion_matrix(y_true, y_pred))
# print('\nMétricas')
# print(classification_report(y_true, y_pred, digits=3))
# print('\nScore')
# score = accuracy_score(y_true, y_pred)
# print(score)
# print('\n')
# if pd.isna(resultados_dataset.at[key, 'Vader']):
# resultados_dataset.loc[key]['Vader'] = score
# salvar_dataset_resultados(resultados_dataset)
# + [markdown] id="l5viuJ5ohb8o"
# ## Flair
# + id="C8s4kFsr5-1t" executionInfo={"status": "ok", "timestamp": 1645490245196, "user_tz": 240, "elapsed": 1, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# def flair_sentiment(texts, classifier):
# sentences = [Sentence(text) for text in texts]
# classifier.predict(sentences, mini_batch_size=32)
# return [
# (sent.labels[0].score, sent.labels[0].value)
# for sent in sentences
# ]
# def classificar_flair(dataset):
# dataset['Flair'] = np.nan
# classifier = TextClassifier.load('sentiment-fast')
# sentiments = flair_sentiment(dataset['reviewText'].values, classifier)
# dataset['Flair'] = [sent[1] for sent in sentiments]
# dataset['Flair'] = pd.Categorical(dataset['Flair'])
# dataset['Flair'] = dataset['Flair'].cat.rename_categories({'NEGATIVE': 0, 'POSITIVE': 1})
# return dataset
# + id="gsyX0sCx5-1u" executionInfo={"status": "ok", "timestamp": 1645490245202, "user_tz": 240, "elapsed": 5, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# dir = '/content/drive/MyDrive/<NAME>/model_save/'
# for key, dataset in datasets.items():
# filename = key + '_Flair.csv'
# if True == os.path.isfile(dir + filename):
# # load dataset from disk
# dataset = pd.read_csv(dir + filename)
# else:
# dataset = classificar_flair(dataset)
# # save new dataset to disk
# dataset.to_csv(dir + filename)
# datasets[key] = dataset
# y_true = dataset['overall'].values
# y_pred = dataset['Flair'].values
# print('Classificador Flair')
# print('Dataset: ', key)
# print('Matriz de confusão')
# print(confusion_matrix(y_true, y_pred))
# print('\nMétricas')
# print(classification_report(y_true, y_pred, digits=3))
# print('\nScore')
# score = accuracy_score(y_true, y_pred)
# print(score)
# print('\n')
# if pd.isna(resultados_dataset.at[key, 'Flair']):
# resultados_dataset.loc[key]['Flair'] = score
# salvar_dataset_resultados(resultados_dataset)
# + [markdown] id="t2fXNMK1UgWP"
# ## AutoGluon
# + id="suKn9XUAqorq" executionInfo={"status": "ok", "timestamp": 1645490245224, "user_tz": 240, "elapsed": 22, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# dir = '/content/drive/MyDrive/<NAME>/model_save/'
# def classificar_autogluon(dataset, modelname):
# #cls = TextPredictor(label='overall', eval_metric='acc', path=(dir + modelname))
# cls = TextPredictor(label='overall', eval_metric='acc')
# # Dividindo o dataset em treino e teste
# train_data, test_data = train_test_split(dataset, test_size = 0.2, random_state = 0)
# cls.fit(train_data, time_limit=60)
# y_pred = cls.predict(test_data)
# y_test = test_data.iloc[:, -1].values
# return y_pred, y_test
# + id="tGf-25P2qorr" executionInfo={"status": "ok", "timestamp": 1645490245226, "user_tz": 240, "elapsed": 1, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# for key, dataset in datasets.items():
# modelname = 'autogluon_' + key
# if True == os.path.isdir(dir + modelname):
# # load model from disk
# cls = TextPredictor.load(dir + modelname)
# corpus = limpa_texto(dataset) # todo
# X = cria_bag_words(corpus) # todo
# y = dataset.iloc[:, -1].values # todo
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# y_pred = cls.predict(X_test) # todo
# else:
# y_pred, y_test = classificar_autogluon(dataset, modelname)
# print('Classificador AutoGluon')
# print('Dataset: ', key)
# print('Matriz de confusão')
# print(confusion_matrix(y_test, y_pred))
# print('\nMétricas')
# print(classification_report(y_test, y_pred, digits=3))
# print('\nScore')
# score = accuracy_score(y_test, y_pred)
# print(score)
# print('\n')
# if pd.isna(resultados_dataset.at[key, 'AutoGluon']):
# resultados_dataset.loc[key]['AutoGluon'] = score
# salvar_dataset_resultados(resultados_dataset)
# + [markdown] id="59JOoET_UlO_"
# ## Auto-Sklearn
# + id="346A7hoWU1g7" executionInfo={"status": "ok", "timestamp": 1645490245229, "user_tz": 240, "elapsed": 2, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# def classificar_auto_sklearn(dataset):
# cls = autosklearn.classification.AutoSklearnClassifier(n_jobs=8, time_left_for_this_task=60)
# corpus = limpa_texto(dataset)
# X = cria_bag_words(corpus)
# y = dataset.iloc[:, -1].values
# # Dividindo o dataset em treino e teste
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# cls.fit(X_train, y_train)
# y_pred = cls.predict(X_test)
# return y_pred, y_test
# + id="9yRW6uUrU1g8" executionInfo={"status": "ok", "timestamp": 1645490245231, "user_tz": 240, "elapsed": 0, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
# dir = '/content/drive/MyDrive/<NAME>/model_save/'
# for key, dataset in datasets.items():
# filename = key + '_Auto_Sklearn.sav'
# if True == os.path.isfile(dir + filename):
# # load model from disk
# clf_auto_sklearn = joblib.load(dir + filename)
# corpus = limpa_texto(dataset) # todo
# X = cria_bag_words(corpus) # todo
# y = dataset.iloc[:, -1].values # todo
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# y_pred = clf_auto_sklearn.predict(X_test) # todo
# else:
# y_pred, y_test = classificar_auto_sklearn(dataset)
# print('Classificador Auto-Sklearn')
# print('Dataset: ', key)
# print('Matriz de confusão')
# print(confusion_matrix(y_test, y_pred))
# print('\nMétricas')
# print(classification_report(y_test, y_pred, digits=3))
# print('\nScore')
# score = accuracy_score(y_test, y_pred)
# print(score)
# print('\n')
# if pd.isna(resultados_dataset.at[key, 'Auto-Sklearn']):
# resultados_dataset.loc[key]['Auto-Sklearn'] = score
# salvar_dataset_resultados(resultados_dataset)
# + [markdown] id="-gVScdfJUp6-"
# ## TPOT
# + id="Q71DVIf2czWY" executionInfo={"status": "ok", "timestamp": 1645490272127, "user_tz": 240, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}}
def classificar_tpot(dataset):
tpot = TPOTClassifier(max_time_mins = 5, memory='auto', n_jobs=-2, use_dask = True, random_state=0)
corpus = limpa_texto(dataset)
X = cria_bag_words(corpus)
y = dataset.iloc[:, -1].values
# Dividindo o dataset em treino e teste
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
tpot.fit(X_train, y_train)
y_pred = tpot.predict(X_test)
return y_pred, y_test
# + id="ib_aTRaiczWZ"
for key, dataset in datasets.items():
y_pred, y_test = classificar_tpot(dataset)
print('Classificador TPOT')
print('Dataset: ', key)
print('Matriz de confusão')
print(confusion_matrix(y_test, y_pred))
print('\nMétricas')
print(classification_report(y_test, y_pred, digits=3))
print('\nScore')
score = accuracy_score(y_test, y_pred)
print(score)
print('\n')
if pd.isna(resultados_dataset.at[key, 'TPOT']):
resultados_dataset.loc[key]['TPOT'] = score
salvar_dataset_resultados(resultados_dataset)
# + [markdown] id="SmY50VpIsuRU"
# # Resultados
# + id="Bhpph3pD87g4" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1645490044634, "user_tz": 240, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgaKjqj1vGBoR4llyAObbSbLjfrdGVGCpihAwf_fQ=s64", "userId": "13763402595986886854"}} outputId="8a44afc9-8f46-4963-bb1e-7a8511e0debb"
resultados_dataset
| review_data_by_category.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/TattooeDeer/T3-ANN/blob/master/Tarea3.ipynb)
# + id="U33cHS9jqmBN" colab_type="code" colab={}
# + [markdown] id="4GkiOm_HqvUC" colab_type="text"
# # Tarea 3 Redes Neuronales
# ## <NAME>. 201273604-8
# ## <NAME>.
#
#
# + id="9Vtp45CXrMVy" colab_type="code" colab={}
| Tarea3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Exceptions
#
# When we learned about testing, we saw that Python complains when things go wrong by raising an "Exception" naming a type of error:
#
#
#
# + tags=["raises-exception"]
1 / 0
# -
# Exceptions are objects, forming a [class hierarchy](https://docs.python.org/3/library/exceptions.html#exception-hierarchy). We just raised an instance
# of the `ZeroDivisionError` class, making the program crash. If we want more
# information about where this class fits in the hierarchy, we can use [Python's
# `inspect` module](https://docs.python.org/3/library/inspect.html) to get a chain of classes, from `ZeroDivisionError` up to `object`:
# +
import inspect
inspect.getmro(ZeroDivisionError)
# -
#
#
# So we can see that a zero division error is a particular kind of Arithmetic Error.
#
#
#
# + tags=["raises-exception"]
x = 1
for y in x:
print(y)
# -
inspect.getmro(TypeError)
# ## Create your own Exception
# When we were looking at testing, we saw that it is important for code to crash with a meaningful exception type when something is wrong.
# We raise an Exception with `raise`. Often, we can look for an appropriate exception from the standard set to raise.
#
# However, we may want to define our own exceptions. Doing this is as simple as inheriting from Exception (or one of its subclasses):
# + tags=["raises-exception"]
class MyCustomErrorType(ArithmeticError):
pass
raise (MyCustomErrorType("Problem"))
# -
#
#
# You can add custom data to your exception:
#
#
#
# + tags=["raises-exception"]
class MyCustomErrorType(Exception):
def __init__(self, category=None):
self.category = category
def __str__(self):
return f"Error, category {self.category}"
raise (MyCustomErrorType(404))
# -
#
#
# The real power of exceptions comes, however, not in letting them crash the program, but in letting your program handle them. We say that an exception has been "thrown" and then "caught".
#
#
#
# +
import yaml
try:
config = yaml.load(open("datasource.yaml"))
user = config["userid"]
password = config["password"]
except FileNotFoundError:
print("No password file found, using anonymous user.")
user = "anonymous"
password = <PASSWORD>
print(user)
# -
#
#
# Note that we specify only the error we expect to happen and want to handle. Sometimes you see code that catches everything:
#
#
#
# +
try:
config = yaml.lod(open("datasource.yaml"))
user = config["userid"]
password = config["password"]
except:
user = "anonymous"
password = <PASSWORD>
print(user)
# -
# This can be dangerous and can make it hard to find errors! There was a mistyped function name there ('`lod`'), but we did not notice the error, as the generic except caught it.
# Therefore, we should be specific and catch only the type of error we want.
# ## Managing multiple exceptions
# Let's create two credential files to read
# +
with open("datasource2.yaml", "w") as outfile:
outfile.write("userid: eidle\n")
outfile.write("password: <PASSWORD>")
with open("datasource3.yaml", "w") as outfile:
outfile.write("user: eidle\n")
outfile.write("password: <PASSWORD>")
# -
# And create a function that reads credentials files and returns the username and password to use.
def read_credentials(source):
try:
datasource = open(source)
config = yaml.safe_load(datasource)
user = config["userid"]
password = config["password"]
datasource.close()
except FileNotFoundError:
print("Password file missing")
user = "anonymous"
password = None
except KeyError:
print("Expected keys not found in file")
user = "anonymous"
password = None
return user, password
print(read_credentials("datasource2.yaml"))
print(read_credentials("datasource.yaml"))
print(read_credentials("datasource3.yaml"))
# This last code has a flaw: the file was successfully opened, the missing key was noticed, but not explicitly closed. It's normally OK, as Python will close the file as soon as it notices there are no longer any references to datasource in memory, after the function exits. But this is not good practice, you should keep a file handle for as short a time as possible.
def read_credentials(source):
try:
datasource = open(source)
config = yaml.load(datasource)
user = config["userid"]
password = config["password"]
except FileNotFoundError:
user = "anonymous"
password = None
finally:
datasource.close()
return user, password
# The `finally` clause is executed whether or not an exception occurs.
#
# The last optional clause of a `try` statement, an `else` clause is called only if an exception is NOT raised. It can be a better place than the `try` clause to put code other than that which you expect to raise the error, and which you do not want to be executed if the error is raised. It is executed in the same circumstances as code put in the end of the `try` block, the only difference is that errors raised during the `else` clause are not caught. Don't worry if this seems useless to you; most languages' implementations of try/except don't support such a clause.
def read_credentials(source):
try:
datasource = open(source)
except FileNotFoundError:
user = "anonymous"
password = None
else:
config = yaml.load(datasource)
user = config["userid"]
password = config["password"]
finally:
datasource.close()
return user, password
#
#
# Exceptions do not have to be caught close to the part of the program calling
# them. They can be caught anywhere "above" the calling point in
# the call stack: control can jump arbitrarily far in the program: up to the `except` clause of the "highest" containing try statement.
#
#
#
def f4(x):
if x == 0:
return
if x == 1:
raise ArithmeticError()
if x == 2:
raise SyntaxError()
if x == 3:
raise TypeError()
def f3(x):
try:
print("F3Before")
f4(x)
print("F3After")
except ArithmeticError:
print("F3Except (💣)")
def f2(x):
try:
print("F2Before")
f3(x)
print("F2After")
except SyntaxError:
print("F2Except (💣)")
def f1(x):
try:
print("F1Before")
f2(x)
print("F1After")
except TypeError:
print("F1Except (💣)")
f1(0)
f1(1)
f1(2)
f1(3)
# ## Design with Exceptions
#
# Now we know how exceptions work, we need to think about the design implications... How best to use them.
#
# Traditional software design theory will tell you that they should only be used
# to describe and recover from **exceptional** conditions: things going wrong.
# Normal program flow shouldn't use them.
#
# Python's designers take a different view: use of exceptions in normal flow is
# considered OK. For example, all iterators raise a `StopIteration` exception to
# indicate the iteration is complete.
#
# A commonly recommended Python design pattern is to use exceptions to determine
# whether an object implements a protocol (concept/interface), rather than testing
# on type.
#
# For example, we might want a function which can be supplied *either* a data
# series *or* a path to a location on disk where data can be found. We can
# examine the type of the supplied content:
# +
import yaml
def analysis(source):
if type(source) == dict:
name = source["modelname"]
else:
content = open(source)
source = yaml.safe_load(content)
name = source["modelname"]
print(name)
# -
analysis({"modelname": "Super"})
with open("example.yaml", "w") as outfile:
outfile.write("modelname: brilliant\n")
analysis("example.yaml")
#
#
#
# However, we can also use the try-it-and-handle-exceptions approach to this.
#
#
#
# +
def analysis(source):
try:
name = source["modelname"]
except TypeError:
content = open(source)
source = yaml.safe_load(content)
name = source["modelname"]
print(name)
analysis("example.yaml")
# -
# This approach is more extensible, and **behaves properly if we give it some
# other data-source which responds like a dictionary or string.**
# +
def analysis(source):
try:
name = source["modelname"]
except TypeError:
# Source was not a dictionary-like object
# Maybe it is a file path
try:
content = open(source)
source = yaml.safe_load(content)
name = source["modelname"]
except IOError:
# Maybe it was already raw YAML content
source = yaml.safe_load(source)
name = source["modelname"]
print(name)
analysis("modelname: Amazing")
# -
# Sometimes we want to catch an error, partially handle it, perhaps add some
# extra data to the exception, and then re-raise to be caught again further up
# the call stack.
#
# The keyword "`raise`" with no argument in an `except:` clause will cause the
# caught error to be re-thrown. Doing this is the only circumstance where it is
# safe to do `except:` without catching a specific type of error.
try:
# Something
pass
except:
# Do this code here if anything goes wrong
raise
# If you want to be more explicit about where the error came from, you can use the `raise from` syntax, which will create a chain of exceptions:
# + tags=["raises-exception"]
def lower_function():
raise ValueError("Error in lower function!")
def higher_function():
try:
lower_function()
except ValueError as e:
raise RuntimeError("Error in higher function!") from e
higher_function()
# -
#
#
# It can be useful to catch and re-throw an error as you go up the chain, doing any clean-up needed for each layer of a program.
#
# The error will finally be caught and not re-thrown only at a higher program
# layer that knows how to recover. This is known as the "throw low catch high"
# principle.
#
#
#
| module08_advanced_programming_techniques/08_03_exceptions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
import ipyjana
w = ipyjana.HelloWorld()
assert w.value == 'Hello Worldooo'
w
import ipywidgets as widgets
w.value = "hohoho"
| examples/introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true id="5PuTC8MC0Hpn"
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"></ul></div>
# + colab={"base_uri": "https://localhost:8080/"} id="_2-verv20MUn" executionInfo={"status": "ok", "timestamp": 1612975252174, "user_tz": 300, "elapsed": 21990, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}} outputId="ba5a764b-1910-4294-8cf0-fef027cd86a9"
from google.colab import drive
drive.mount('/content/drive')
# + id="ylmXqkVxrj1j"
# + colab={"base_uri": "https://localhost:8080/"} id="1AMVu46hovJr" executionInfo={"status": "ok", "timestamp": 1612975467757, "user_tz": 300, "elapsed": 8202, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}} outputId="70ffac65-754e-43f7-dd02-2e33d4e7e484"
from os.path import expanduser
try:
import google.colab
IN_COLAB = True
# %tensorflow_version 1.x
import tensorflow as tf
print("Using tensorflow v" + str(tf.__version__))
from tensorflow.keras.callbacks import ReduceLROnPlateau, EarlyStopping
data_path = '/content/drive/MyDrive/GitHub IU/bam/deepposekit_clone/deepposekit_data_custom/'
source = 'hand'
HOME = data_path + f"{source}/"
print(HOME, source)
except:
import tensorflow as tf
print("Using tensorflow v" + str(tf.__version__))
from tensorflow.keras.callbacks import ReduceLROnPlateau, EarlyStopping
IN_COLAB = False
data_path = 'E:\\Work\\github repos\\deepposekit_data_custom\\'
source = 'hand'
HOME = data_path + f"{source}/"
print(HOME, source)
import numpy as np
import matplotlib.pyplot as plt
import sys
# !{sys.executable} -m pip install -U deepposekit pyrealsense2
# HOW TO USE THIS SCRIPT:
# 1. Update the source
# 2. Make sure annotator is uncommented
# 3. Maybe change the text scaling
from deepposekit import Annotator
from deepposekit.io import VideoReader, DataGenerator, initialize_dataset, TrainingGenerator, BaseGenerator
from deepposekit.io import ImageGenerator, VideoWriter
from deepposekit.io.utils import merge_new_images
from deepposekit.annotate import KMeansSampler
from deepposekit.augment import FlipAxis
from deepposekit.models import StackedDenseNet, DeepLabCut,StackedHourglass,LEAP
from deepposekit.models import load_model
from deepposekit.callbacks import Logger, ModelCheckpoint
from scipy.signal import find_peaks
#import pyrealsense2 as rs
import tqdm
import cv2
import imgaug.augmenters as iaa
import imgaug as ia
#source = 'chick-toy'
#HOME = f"{source}/"
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="ww-MxZIVqDU-" executionInfo={"status": "ok", "timestamp": 1612975470413, "user_tz": 300, "elapsed": 384, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}} outputId="76284ad1-73d4-4b24-b4ab-095a9613859b"
HOME + f'{source}_annotation_set.h5'
# + id="bdy91hM7ulwu" executionInfo={"status": "ok", "timestamp": 1612975470534, "user_tz": 300, "elapsed": 191, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}}
def resize_raw_video(resize_shape = (256, 256)):
cap = cv2.VideoCapture(HOME + f'{source}_raw.mp4')
# fourcc = cv2.VideoWriter_fourcc(*'XVID')
# out = cv2.VideoWriter(HOME + video_file_path + 'pose_human_test_resized.mp4',fourcc, 30, resize_shape)
out = VideoWriter(HOME + f'{source}.mp4', resize_shape, 'mp4v', 30.0, color=True)
while True:
ret, frame = cap.read()
if ret == True:
b = cv2.resize(frame,resize_shape,fx=0,fy=0, interpolation = cv2.INTER_CUBIC)
out.write(b)
else:
break
cap.release()
out.close()
cv2.destroyAllWindows()
# + id="iFFAC5NN0hBo" executionInfo={"status": "ok", "timestamp": 1612975471468, "user_tz": 300, "elapsed": 468, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}}
#resize_raw_video(resize_shape = (256, 256))
# + id="EgF0kAkHo4iA" executionInfo={"status": "ok", "timestamp": 1612975471469, "user_tz": 300, "elapsed": 281, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}}
def annotate_dataset(overwrite=False):
print("[INFO] Preparing Data")
# batch_size must be large else kmeans can't be performed
reader = VideoReader(HOME + f'{source}.mp4', batch_size=100, gray=True)
randomly_sampled_frames = []
for idx in tqdm.tqdm(range(len(reader)-1)):
batch = reader[idx]
random_sample = batch[np.random.choice(batch.shape[0], 10, replace=False)]
randomly_sampled_frames.append(random_sample)
reader.close()
randomly_sampled_frames = np.concatenate(randomly_sampled_frames)
kmeans = KMeansSampler(n_clusters=10, max_iter=100, n_init=20, batch_size=100, verbose=True)
kmeans.fit(randomly_sampled_frames)
kmeans_sampled_frames, kmeans_cluster_labels = kmeans.sample_data(randomly_sampled_frames, n_samples_per_label=50)
try:
initialize_dataset(
images=kmeans_sampled_frames,
datapath=HOME + f'{source}_annotation_set.h5',
skeleton=HOME + 'skeleton.csv',
overwrite=overwrite
)
except OSError:
print("[INFO] Dataset Exists - Passing.")
# THIS CANNOT BE DONE FROM WITHIN GOOGLE COLAB. USE PYCHARM or an IDE
Annotator(datapath = HOME + f'{source}_annotation_set.h5',
dataset ='images',
skeleton = HOME + 'skeleton.csv',
shuffle_colors = False,
text_scale = 0.2).run()
# + id="bBquSTbtrA8v" executionInfo={"elapsed": 143, "status": "ok", "timestamp": 1612973077530, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}, "user_tz": 300} outputId="03b42e28-e958-4e9d-dad8-ac2c42e0b761"
#annotate_dataset(overwrite=True)
# + colab={"base_uri": "https://localhost:8080/"} id="blGahhS_rfE_" executionInfo={"status": "ok", "timestamp": 1612975475171, "user_tz": 300, "elapsed": 205, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}} outputId="48443ae0-80f1-49ac-b76e-98d2f3fd64d1"
import os
os.listdir(HOME)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="oOfG7rEFrbcy" executionInfo={"status": "ok", "timestamp": 1612975475639, "user_tz": 300, "elapsed": 320, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}} outputId="c4cbb745-d781-4bbe-e898-9f36e1efe5c4"
HOME + f'{source}_annotation_set.h5'
# + id="Qs2jbvr5rAQe" executionInfo={"status": "ok", "timestamp": 1612975476583, "user_tz": 300, "elapsed": 284, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}}
def prepare_model():
print("Loading Data Generator")
data_generator = DataGenerator(HOME + f'{source}_annotation_set.h5', mode="annotated")
print("Creating Data Augmenter")
augmenter = []
augmenter.append(FlipAxis(data_generator, axis=0)) # flip image up-down
augmenter.append(FlipAxis(data_generator, axis=1)) # flip image left-right
sometimes = []
sometimes.append(iaa.Affine(scale={"x": (0.95, 1.05), "y": (0.95, 1.05)},
translate_percent={'x': (-0.05, 0.05), 'y': (-0.05, 0.05)},
shear=(-8, 8),
order=ia.ALL,
cval=ia.ALL,
mode=ia.ALL)
)
sometimes.append(iaa.Affine(scale=(0.8, 1.2),
mode=ia.ALL,
order=ia.ALL,
cval=ia.ALL)
)
augmenter.append(iaa.Sometimes(0.75, sometimes))
augmenter.append(iaa.Affine(rotate=(-180, 180),
mode=ia.ALL,
order=ia.ALL,
cval=ia.ALL)
)
augmenter = iaa.Sequential(augmenter)
print("Creating Training Generator")
train_generator = TrainingGenerator(generator=data_generator,
downsample_factor=2,
augmenter=augmenter,
sigma=3,
validation_split=0.1,
use_graph=True,
random_seed=1,
graph_scale=1)
print(train_generator.get_config())
train_generator.on_epoch_end()
with tf.device("gpu:0"):
print("[INFO] Preparing Model")
# SELECT MODEL
# model = StackedDenseNet(train_generator, n_stacks=5, growth_rate=32, pretrained=True)
# model = DeepLabCut(train_generator, backbone="resnet50")
# model = DeepLabCut(train_generator, backbone="mobilenetv2", alpha=0.75) # Increase alpha to improve accuracy
model = DeepLabCut(train_generator, backbone="densenet121")
# model = LEAP(train_generator)
# model = StackedHourglass(train_generator)
model.get_config()
reduce_lr = ReduceLROnPlateau(monitor="loss", factor=0.2, verbose=1, patience=20)
model_checkpoint = ModelCheckpoint(
HOME + "best_model_densenet.h5",
monitor="loss",
# monitor="loss" # use if validation_split=0
verbose=1,
save_best_only=True,
)
early_stop = EarlyStopping(
monitor="loss",
# monitor="loss" # use if validation_split=0
min_delta=0.001,
patience=100,
verbose=1
)
print("Training model...")
callbacks = [early_stop, reduce_lr, model_checkpoint]
model.fit(
batch_size=1,
validation_batch_size=1,
callbacks=callbacks,
epochs=2,
steps_per_epoch=None,
)
model = load_model(
HOME + "best_model_densenet.h5",
augmenter=augmenter,
generator=data_generator,
)
model.fit(
batch_size=1,
validation_batch_size=1,
callbacks=callbacks,
epochs=500,
steps_per_epoch=None,
)
# + colab={"base_uri": "https://localhost:8080/"} id="2acH7dpDo6SN" executionInfo={"status": "ok", "timestamp": 1612978884467, "user_tz": 300, "elapsed": 3210591, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}} outputId="dddfee7a-5ef3-4557-b850-1adbc7de5791"
prepare_model()
# + id="c81LKvmgrDRe" executionInfo={"status": "ok", "timestamp": 1612978884949, "user_tz": 300, "elapsed": 191, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}}
def create_video():
print("[INFO] Creating Output Video")
with tf.device("gpu:0"):
model = load_model(HOME + 'best_model_densenet.h5')
model_size = tuple(model.input_shape[:2])
print(model_size, model_size[::-1])
model_size = model_size[::-1]
print("Reading Video...")
reader = VideoReader(HOME + f'{source}.mp4', batch_size=1, gray=True)
predictions = model.predict(reader, verbose=1)
np.save(HOME + 'predictions.npy', predictions)
#############################################
data_generator = DataGenerator(HOME + f'{source}_annotation_set.h5')
predictions = predictions[..., :2]
print(predictions.shape)
cmap = plt.cm.hsv(np.linspace(0, 1, data_generator.keypoints_shape[0]))[:, :3][:, ::-1] * 255
writer = VideoWriter(HOME + f'{source}_predicted.mp4', model_size, 'mp4v', 30.0, color=True)
for frame, keypoints in tqdm.tqdm(zip(reader, predictions)):
frame = frame[0]
frame = cv2.cvtColor(frame, cv2.COLOR_GRAY2RGB)
for i, node in enumerate(data_generator.graph):
if node >= 0:
pt1 = keypoints[i]
pt2 = keypoints[node]
cv2.line(frame, (pt1[0], pt1[1]), (pt2[0], pt2[1]), (0, 0, 255), 1, cv2.LINE_AA)
for i, keypoint in enumerate(keypoints):
keypoint = keypoint.astype(int)
cv2.circle(frame, (keypoint[0], keypoint[1]), 1, tuple(cmap[i]), -1, cv2.LINE_AA)
writer.write(frame)
writer.close()
reader.close()
# + colab={"base_uri": "https://localhost:8080/"} id="g_an9IySo7Z6" executionInfo={"status": "ok", "timestamp": 1612978986947, "user_tz": 300, "elapsed": 102169, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08904726702539992993"}} outputId="550c9826-f510-4aa1-e1f4-66057ad7c6a8"
create_video()
print("[INFO] Process Finished")
# + id="uAA015xfswNS"
| examples/DeepPoseKit main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="CU4uf3Zsd7XX"
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import OrdinalEncoder
import pickle
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="yltbS_J1U5ha" executionInfo={"status": "ok", "timestamp": 1621149596667, "user_tz": -120, "elapsed": 30132, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="4e64a9d3-7edc-4c7f-8483-f26c1fafa186"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="GDGSnD-TuOpZ"
# ## Classes Bonus Malus
# + id="ySjqhfUrfo24" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621149602386, "user_tz": -120, "elapsed": 5630, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="69e72e3e-ae16-4917-c721-0bf6f3a6076c"
classe= pd.read_csv('/content/drive/MyDrive/dataset/ClassesBonusMalus.csv', sep=';')
print(classe.shape)
pd.set_option("display.max_rows", 10)
# + [markdown] id="3C4vb7dWvben"
# **we fill the empty values with the most frequent values**
# + id="kQcrdzPWhU3d" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621149604437, "user_tz": -120, "elapsed": 802, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="cf540f99-0af2-4181-efcc-e9ece35cf89c"
print(classe.columns)
print(classe["CodeBonusMalus_id"].isna().sum())
print(classe["CodePolice_id"].isna().sum())
print(classe["CodeVehicule_id"].isna().sum())
print(classe["CodeAssure_id"].isna().sum())
print("CodeSouscripteur_id",classe["CodeSouscripteur_id"].isna().sum())
print("codeUsage",classe["codeUsage"].isna().sum())
print("ETAT_VEHICULE",classe["ETAT_VEHICULE"].isna().sum())
print("DATE_AFFECTATION",classe["DATE_AFFECTATION"].isna().sum())
print("classeBonusMalus",classe["classeBonusMalus"].isna().sum())
print("classeBonusMalusCompagnie",classe["classeBonusMalusCompagnie"].isna().sum())
# + id="oV25uedzjNuS"
classe["CodeSouscripteur_id"]=classe["CodeSouscripteur_id"].fillna(classe["CodeSouscripteur_id"].value_counts().idxmax())
classe["codeUsage"]=classe["codeUsage"].fillna(classe["codeUsage"].value_counts().idxmax())
classe["ETAT_VEHICULE"]=classe["ETAT_VEHICULE"].fillna(classe["ETAT_VEHICULE"].value_counts().idxmax())
classe["DATE_AFFECTATION"]=classe["DATE_AFFECTATION"].fillna(classe["DATE_AFFECTATION"].value_counts().idxmax())
classe["classeBonusMalus"]=classe["classeBonusMalus"].fillna(classe["classeBonusMalus"].value_counts().idxmax())
classe["classeBonusMalusCompagnie"]=classe["classeBonusMalusCompagnie"].fillna(classe["classeBonusMalusCompagnie"].value_counts().idxmax())
classe["coefBonusMalus"]=classe["coefBonusMalus"].fillna(classe["coefBonusMalus"].value_counts().idxmax())
# + id="xOiNDErpzPvJ"
classe["DATE_AFFECTATION"]=classe["DATE_AFFECTATION"].replace('2516-07-08 00:00:00.000000', '2016-07-08 00:00:00.000000')
classe["DATE_AFFECTATION"]=classe["DATE_AFFECTATION"].replace('2115-03-02 00:00:00.000000', '2015-03-02 00:00:00.000000')
# + id="rGVHqA0Cs-uh" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1620587053004, "user_tz": -120, "elapsed": 4745, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="984ca28a-7d78-4886-e901-b337dc4f911d"
classe
# + colab={"base_uri": "https://localhost:8080/"} id="TG9WqrBIDQ7d" executionInfo={"status": "ok", "timestamp": 1621149728432, "user_tz": -120, "elapsed": 724, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="7248221c-5988-4d15-f91d-50fd30d606aa"
pd.set_option("display.max_rows", 30)
classe.dtypes
# + colab={"base_uri": "https://localhost:8080/"} id="Ofji0X6eSx0J" executionInfo={"status": "ok", "timestamp": 1621153897266, "user_tz": -120, "elapsed": 710, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="95836637-b34f-4f3c-fcca-0b16458b2a6a"
print(classe['codeUsage'].value_counts())
# + id="_MYf5tI-HOxP"
classe.to_csv('/content/drive/MyDrive/dataset/new_ClassesBonusMalus2.csv')
# + [markdown] id="pbaZbBbXbA6n"
# ## Police
# + id="cAUqMA_PURmL" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621149993757, "user_tz": -120, "elapsed": 2979, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="c0b369d9-528a-468d-c275-d275bf91214a"
police = pd.read_csv('/content/drive/MyDrive/dataset/PoliceFinalCleaned.csv')
# + [markdown] id="_zD-z9FQ2ntK"
# **we discover the distribution of data values**
# + id="BdbnveHlbA6r" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621149994186, "user_tz": -120, "elapsed": 3325, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="20d92a80-b929-42e5-cf6f-77d9e69a3fc8"
police.isna().sum()
# + id="itMXziwWbA6t" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621149994198, "user_tz": -120, "elapsed": 3276, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="274c93bf-061f-4b99-b5f2-44029c05cdc3"
print('naturePolice \n',police['naturePolice'].value_counts())
print('typePolice \n',police['typePolice'].value_counts())
print('Etat_Police \n',police['Etat_Police'].value_counts())
#print('verouillageModifPolice \n',police['verouillageModifPolice'].value_counts())
# + id="WKLo4EeRyfxL" colab={"base_uri": "https://localhost:8080/", "height": 444} executionInfo={"status": "ok", "timestamp": 1621149995200, "user_tz": -120, "elapsed": 4203, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="4cf45c86-b78c-4035-c317-3cb70e0f9ed5"
police.dropna()
# + colab={"base_uri": "https://localhost:8080/"} id="YiYP3WBlt3u7" executionInfo={"status": "ok", "timestamp": 1621149995215, "user_tz": -120, "elapsed": 4151, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="27815744-5f81-436b-b9ce-c6a271bd366a"
police['dateEcheancePolice'].value_counts()
# + id="Ki5BeRwFw_ya"
police['dateEcheancePolice']=police['dateEcheancePolice']*0.01
# + id="ir-Yqwop4rJV" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621149995242, "user_tz": -120, "elapsed": 4039, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="dc38e5b7-590a-4d1a-eb11-5c785759ab07"
police['dateEcheancePolice'].isna().sum()
# + id="dWKh4FEo1Po0" colab={"base_uri": "https://localhost:8080/", "height": 53} executionInfo={"status": "ok", "timestamp": 1621149995257, "user_tz": -120, "elapsed": 3982, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="39ddfe0e-8831-4b40-ccd2-5bc066a8350b"
"""
police['dateEcheancePolice'].index
dateEcheancePolice = pd.DataFrame(rows,index=police['dateEcheancePolice'].index, columns=['dateEcheancePolice'])
dateEcheancePolice
"""
# + id="8qmXWN-p5NTG" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621149995269, "user_tz": -120, "elapsed": 3929, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="c6c5dc66-63aa-4cab-bd0b-7845f0f26134"
police['dateEcheancePolice'].isna().sum()
# + id="ehdeEW8P3M4k" colab={"base_uri": "https://localhost:8080/", "height": 444} executionInfo={"status": "ok", "timestamp": 1621149995757, "user_tz": -120, "elapsed": 4359, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="407f0b94-377e-4ce9-c8bb-89295e516ec8"
police.dropna()
# + id="9hVxsr2VHFYQ"
a=pd.unique(police['numPolice'])
np.savetxt("numPolice.csv",a,fmt = '%s', delimiter=";")
# + colab={"base_uri": "https://localhost:8080/"} id="cMKSSaSqD4_N" executionInfo={"status": "ok", "timestamp": 1621154179678, "user_tz": -120, "elapsed": 1057, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="32064eb4-fb7d-49f9-e245-2aa0883e4548"
pd.set_option("display.max_rows", 30)
print(police['typeIntermediaire'].value_counts())
print(police['naturePolice'].value_counts())
print(police['typePolice'].value_counts())
print(police['Etat_Police'].value_counts())
print(police['dateEcheancePolice'].value_counts())
print(police['DATE_RESILIATION'].value_counts())
print(police['dateEcheancePolice'].value_counts())
# + id="KdMITFR6qfz9"
police.to_csv('/content/drive/MyDrive/dataset/new_police.csv')
# + [markdown] id="uCB31Q9C1PYH"
# **we make the correspondence between police and class**
# + [markdown] id="w2OawLhLbA60"
# ## Vehicule
# + id="8wt_L1RCbA62" colab={"base_uri": "https://localhost:8080/", "height": 479} executionInfo={"status": "ok", "timestamp": 1621152701453, "user_tz": -120, "elapsed": 4179, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="0c7000bb-3d80-4fba-a0f3-038b1a924ddb"
Vehicule = pd.read_csv('/content/drive/MyDrive/dataset/Vehicule.csv',sep=";")
Vehicule
# + [markdown] id="gfTSaqFU2e8A"
# **we discover the distribution of data values**
# + id="91wINPL8bA64" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621152701805, "user_tz": -120, "elapsed": 4520, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="ea6ed032-16fc-4ff7-fab9-a2343e99149e"
Vehicule.isna().sum()
# + id="Iy7Cz10MbA66"
Vehicule.drop(columns=["numChassis","dateDerniereVisite","dateAjout","etatVehicule","dateMiseEpave","DATE_RETRAIT","dateMiseCirculation","dateMiseAjourVehicule" ], axis=1, inplace=True)
# + id="WX8yhUPvbA7h" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621152702077, "user_tz": -120, "elapsed": 4780, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="dac0fd2d-64ff-4d53-dbb3-094386ba3364"
Vehicule['energie'].value_counts()
# + id="cHZy3_jNbA78"
Vehicule['energie']=Vehicule['energie'].fillna('GASOIL')
# + id="4V5RSFTgbA7-" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621152702326, "user_tz": -120, "elapsed": 5020, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="b4ff4d8b-c3df-426b-885e-1e8c01a981f6"
Vehicule.isna().sum()
# + id="zKpLun7HbA7_"
Vehicule=Vehicule.dropna()
# + id="SvEBvhdibA8A" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621152852576, "user_tz": -120, "elapsed": 630, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="7a90d787-a799-44a4-9bdc-6b037f1444b2"
Vehicule.dtypes
# + colab={"base_uri": "https://localhost:8080/"} id="CvgyFzzTOVtc" executionInfo={"status": "ok", "timestamp": 1621153694039, "user_tz": -120, "elapsed": 725, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="aca83c9f-3134-4051-85de-d8931b9fbc11"
print(Vehicule['CodeVehicule_id'].value_counts())
print(Vehicule['puissanceFiscal'].value_counts())
print(Vehicule['typeImmatriculation'].value_counts())
print(Vehicule['dateInsertion'].value_counts())
print(Vehicule['codeMarque'].value_counts())
print(Vehicule['numImmatriculation'].value_counts())
# + id="c_E9MvnEbA8B" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1621152702847, "user_tz": -120, "elapsed": 5518, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="3b1bfb47-fc06-46e8-e847-1177c788c64d"
Vehicule.shape
# + id="ffXvOV1aqTGV"
Vehicule.to_csv('/content/drive/MyDrive/dataset/new_Vehicule.csv')
# + [markdown] id="_EOz6PshQSji"
# ## Merge data
# + [markdown] id="zJEqK6II1Aeb"
# **we make the correspondence between Vehicule and police_class**
# + id="QGnt5SBMbA8C"
police_class=pd.merge(police,classe,on='CodePolice_id')
Vehicule_police_class=pd.merge(police_class,Vehicule,on='CodeVehicule_id')
# + colab={"base_uri": "https://localhost:8080/"} id="x4Q5QJ8RUUJK" executionInfo={"status": "ok", "timestamp": 1620587077600, "user_tz": -120, "elapsed": 29237, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="a6a8a866-366a-47ed-b10d-c5d17e120d5e"
Vehicule_police_class.isna().sum().max()
# + id="4dweLVTV-rMP" colab={"base_uri": "https://localhost:8080/", "height": 617} executionInfo={"status": "ok", "timestamp": 1620587077602, "user_tz": -120, "elapsed": 29231, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiE49cF8HkA-A6gYj3IyadS-il3FO__p7maJTkKKA=s64", "userId": "14956717124952145367"}} outputId="dd42f8dd-3b37-4130-9bff-dba39ea961d2"
Vehicule_police_class
# + id="BRAKu2HB-tmT"
Vehicule_police_class.to_csv('/content/drive/MyDrive/dataset/new_date_Vehicule_police_class2.csv')
| DataCGA/Insurance_Data_cleaning_merge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MartyWeissman/PythonForMathematics/blob/main/Math152_Feb18_2021.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5f0GXobaNvLv"
# # Teaching notebook, Feb 18, 2021
#
# Introduction to Numpy!
#
# Outline:
#
# 1. Creating numpy ndarrays
# 2. Coercing types and overflow
# 3. Broadcasting math functions
# 4. Cumulative sum. Approximating $\gamma$
# 4. Plotting a function (line plots)
#
# + id="mDvKuTSGNpGa"
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="NWjciq0iSCBO" outputId="345bd269-a8d4-46f3-ec86-452059e90a82"
np.sqrt(1000)
# + colab={"base_uri": "https://localhost:8080/"} id="iJFE70ksSB4F" outputId="eb2d9d3a-70cd-46a6-bf6e-1aa14228047a"
np.random.random(5) # An array of random numbers between 0 and 1
# + [markdown] id="OEXF7AMhN9N0"
# Numpy ndarrays ("ndarray" is an "n-dimensional array")
# + id="IfHhtcdONzHB"
A = np.array([1,2,3,4,5])
# + colab={"base_uri": "https://localhost:8080/"} id="wfmoBO3vSY8K" outputId="cd51fb06-d98e-4d24-a79f-96a47b632b42"
print(A)
# + id="AmVJttRBSaoJ"
L = [1,2,3,4,5]
# + colab={"base_uri": "https://localhost:8080/"} id="kKD4CHFNScxh" outputId="642420ff-75ab-4510-ab2e-911402053fbc"
print(L)
# + colab={"base_uri": "https://localhost:8080/"} id="6Irwuh5tSdZY" outputId="115fb794-7cfe-4f28-d39f-fc6e5fad9f31"
type(L)
# + colab={"base_uri": "https://localhost:8080/"} id="caWqMBmdSfri" outputId="c0271a14-4812-43f2-a1eb-4e94a502302d"
type(A)
# + colab={"base_uri": "https://localhost:8080/"} id="CXBHrZctSguh" outputId="f42636b6-54a7-4a61-a2e1-692f794fd3eb"
A[2]
# + colab={"base_uri": "https://localhost:8080/"} id="MpvL-_7-Sjpp" outputId="ddf1ba01-f5a6-4225-c6c1-9819bd058e84"
type(A[2])
# + id="pq-iPFq1SkuI"
L = [1, 'S', 3.7]
# + colab={"base_uri": "https://localhost:8080/"} id="wefA_wSPSpxw" outputId="047101e8-d046-4fad-9758-4f96413563d7"
type(L)
# + colab={"base_uri": "https://localhost:8080/"} id="y00-XRO8Sre3" outputId="3ebdbe68-e18c-45ee-d845-ea009dd78f8e"
type(L[0])
# + colab={"base_uri": "https://localhost:8080/"} id="aElfGOCNSsqO" outputId="2f0afab7-f224-4b0c-ac40-1ffab38b996e"
type(L[1])
# + colab={"base_uri": "https://localhost:8080/"} id="otYIRpkOSueX" outputId="994e8733-8bbd-4d69-977e-5e4beab6a2af"
type(L[2])
# + id="57MI5MfySwHf"
A = np.array([1,'S',3.7])
# + colab={"base_uri": "https://localhost:8080/"} id="7HcdB0QzS0IG" outputId="da86a40a-a8d8-4194-c91b-a84e5d30432d"
print(A)
# + [markdown] id="kuSEZMXPS39D"
# Numpy arrays must have all terms of the same type!!
#
# It will try to force this, when possible.
# + id="ujerCqzRTI2k"
n = np.int64(37)
# + colab={"base_uri": "https://localhost:8080/"} id="tjZAX-nzTcVx" outputId="a318c1df-8696-483b-ddc1-6f5548dbc3a0"
n
# + colab={"base_uri": "https://localhost:8080/"} id="af1MuLsOTc35" outputId="0bfbbc10-11d7-4df6-cbf1-51ce79b886a4"
type(n)
# + colab={"base_uri": "https://localhost:8080/"} id="rJm54qjuTeQZ" outputId="740937b6-92fa-4ccb-9f50-1f762c55757b"
n**10
# + colab={"base_uri": "https://localhost:8080/"} id="X5PA4mL3TimI" outputId="b64e67bd-4836-4ac7-f2e4-a3a6c0abf620"
37**10
# + colab={"base_uri": "https://localhost:8080/"} id="aNCBgGDdTkVw" outputId="c1b4eda6-6edb-4f92-fd82-47f3e93f9c77"
n**100
# + colab={"base_uri": "https://localhost:8080/"} id="Dm5zxDfQTliI" outputId="5bfa7d62-3215-45c2-d33a-2361f5c60d9f"
37**100
# + [markdown] id="J6rFQVCCUBhs"
# ## Broadcasting functions!
# + id="idd5P_CETrx4"
A = np.array([1,2,3,4,5])
# + colab={"base_uri": "https://localhost:8080/"} id="dVP_0zXbUEpF" outputId="5bf07999-49af-401f-fcd0-9724cd23978c"
print(A)
# + colab={"base_uri": "https://localhost:8080/"} id="2HdYV34qUFX8" outputId="94ed2ba4-386c-49a9-a0ac-02c27bdb3d04"
type(A)
# + colab={"base_uri": "https://localhost:8080/"} id="eifFGlBuUGF0" outputId="8e248ca5-bfaa-4846-ff50-e7ccb020fc95"
2*[1,2,3,4,5]
# + colab={"base_uri": "https://localhost:8080/"} id="hIKjxdHJUPDT" outputId="28b3b35d-a66f-4a7a-bd17-a91c8ef34fe6"
2*A
# + colab={"base_uri": "https://localhost:8080/"} id="DgGiJDEFUX3K" outputId="99b738cb-d8b2-4a7c-b903-6851c07a921b"
np.sin(A)
# + colab={"base_uri": "https://localhost:8080/"} id="lmUjmIbOUbwB" outputId="10f420f1-69ca-4b7b-f021-e96d5d4bc153"
np.sin(A*A)
# + colab={"base_uri": "https://localhost:8080/"} id="zpO6ZdXKUg2J" outputId="19e293a5-a0ac-4353-ab81-226f4a359dce"
A*A
# + [markdown] id="b1RKDBy3Usen"
# The Euler-Mascheroni constant is
# $$\gamma = \lim_{N \rightarrow \infty} \left( \sum_{n=1}^N \frac{1}{n} - \log(N) \right).$$
# + id="Ftm0OhKgUiRY"
A = np.arange(1,1000000.0)
# + colab={"base_uri": "https://localhost:8080/"} id="yVpSMuUOVClr" outputId="cf30e1d1-c199-4b07-ec0a-e0c301a97796"
A[:10]
# + colab={"base_uri": "https://localhost:8080/"} id="LQeMLQnWVDU9" outputId="3ec02641-7fc9-4319-8d8e-35d40b4d8a39"
reciprocals = 1/A
print(reciprocals[:10])
# + colab={"base_uri": "https://localhost:8080/"} id="s_sH6a7_VRWK" outputId="512b3e61-7cd1-4f1a-d8ae-ae014a0d18e3"
logarithms = np.log(A)
print(logarithms[:10])
# + id="kbLkakEBVYmH"
rec_sums = np.cumsum(reciprocals)
# + colab={"base_uri": "https://localhost:8080/"} id="5mbgXPrnVgcG" outputId="aa873ee0-2ba7-43cd-e922-a0e7d423923a"
rec_sums[:10]
# + colab={"base_uri": "https://localhost:8080/"} id="bcuwDpf9VhYO" outputId="c14de3b0-231d-4881-8cab-1032819108c9"
gamma_approx = rec_sums - logarithms
print(gamma_approx[:10])
# + colab={"base_uri": "https://localhost:8080/"} id="nNOgtvH2VwR0" outputId="a039a550-e434-4060-b957-45a9ce2a6007"
gamma_approx[-1]
# + colab={"base_uri": "https://localhost:8080/"} id="1MMW8ePpV0xc" outputId="414a3ba4-7de1-4bb8-dcba-5a9c588b048c"
np.euler_gamma
# + id="IwMYb_P1V4Qs"
| Math152_Feb18_2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="XEv_uqBG9xfP" outputId="6ab4ce93-d7c7-4ca1-9bdf-6537c90bd191"
# !wget https://www.fjc.gov/sites/default/files/idb/textfiles/cv88on_0.zip --no-check-certificate
# !unzip cv88on_0.zip
# + colab={"base_uri": "https://localhost:8080/"} id="_1x3zvMv-hmq" outputId="d26ab665-34b8-472e-c601-d2b6b9c506a9"
# !ls
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="Xk2z05MgHN2l" outputId="2430c334-b090-4beb-d7c0-cc6b6ab0a4b6"
import warnings
warnings.filterwarnings('ignore')
# for some basic operations
import numpy as np
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="sxVRmw4R-tZH" outputId="d65ec1e1-3476-4e30-ec6a-53b5f17f0a50"
def convert(val):
if val == -8 or val == "-8":
return np.nan
return val
#conv_dict = {"":convert} converters=conv_dict
#req_col = []
#d_type=
to_drop = ["TITL","SECTION","FILEJUDG","FILEMAG","MDLDOCK","PLT","DEF","SUBSECT","DJOINED","PRETRIAL","TRIBEGAN","TRIALEND","TDATEUSE","FDATEUSE"]
df = pd.read_csv("cv88on.txt", sep='\t',encoding='cp1252').drop(to_drop,axis='columns')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="kkMTMU0QzJ1U" outputId="ff844817-da94-4052-f18f-b1cd8cb650b8"
df["NOS"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="mDx0qY_tH8fW" outputId="a0ca4060-9201-4ce5-a444-0bffa04392d1"
df.info()
# + id="ZBbHQpba6eqf"
all_columns = df.columns
for i in range(0,len(all_columns)):
(df[all_columns[i]]).replace("-8",np.nan,inplace=True)
for i in range(0,len(all_columns)):
(df[all_columns[i]]).replace(-8,np.nan,inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="gYvR-xyfE9Z4" outputId="88615323-2b4e-4eea-cb0d-7b6008929306"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="nKRyqnXFUVWS" outputId="f5bcb04b-5e13-47fd-afb4-0f03bed28bb2"
df["JUDGMENT"].replace(4,np.nan,inplace=True)
df["JUDGMENT"].replace(0,np.nan,inplace=True)
#df.drop(df[df["JUDGMENT"] == 4].index ,inplace=True)
#df.drop(df[df["JUDGMENT"] == 0].index ,inplace=True)
df["JUDGMENT"]=df["JUDGMENT"]*10
df["LABEL"] = df["NOJ"] + df["JUDGMENT"]
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="1AwGpjgUtn1u" outputId="1f7b9fc8-860a-44fe-f8c3-2d189252f5e8"
df["TERMDATE"]= pd.to_datetime(df["TERMDATE"])
df["FILEDATE"]= pd.to_datetime(df["FILEDATE"])
df["CASEDAYS"] = (df["TERMDATE"]-df["FILEDATE"]).dt.days
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="DfqKIy0o_HkZ" outputId="c52e1bff-5696-41a6-fe62-994863398aee"
missing_value_count = df.isnull().sum()
total_cells = np.product(df.shape)
total_missing = missing_value_count.sum()
percentage_missing = (total_missing/total_cells)*100
print("Missing Values Count:")
print(missing_value_count)
print("Percentage of missing data:",percentage_missing)
# + colab={"base_uri": "https://localhost:8080/"} id="7b4XY2GSC4aj" outputId="3a719c0f-fa5c-458f-dbc4-d7f4b3791f93"
df.dropna(subset=["LABEL","ORIGIN"],inplace=True)
threshold = len(df)*0.80
print("Old Shape:",df.shape)
df.dropna(axis=1,thresh=threshold,inplace=True)
print("New Shape:",df.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 289} id="IDqUKEgqjvwj" outputId="20e9bfee-f218-48ff-bfe4-5d9fe89db829"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="ffG9NJ-TLEya" outputId="3e79895d-c94c-493a-f8c1-73629a0dc5de"
#to_drop = ["TITL","SECTION","FILEJUDG","FILEMAG","MDLDOCK","PLT","DEF","SUBSECT","DJOINED","PRETRIAL","TRIBEGAN","TRIALEND","AMTREC","TDATEUSE","FDATEUSE"]
#print(to_drop)
#print(to_drop[0])
#for i in range(0,len(to_drop)):
# df.drop(to_drop[i],axis=1,inplace=True)
#df.drop(["TITL","SECTION","FILEJUDG","FILEMAG","MDLDOCK","PLT","DEF","SUBSECT","DJOINED","PRETRIAL","TRIBEGAN","TRIALEND","AMTREC","TDATEUSE","FDATEUSE"],axis=1,inplace=True)
#df.isnull().any()
# + colab={"base_uri": "https://localhost:8080/", "height": 683} id="8SOJzLuC4vV4" outputId="2da97283-0e69-494e-a011-ac429dfbacd3"
missing_value_count = df.isnull().sum()
total_cells = np.product(df.shape)
total_missing = missing_value_count.sum()
percentage_missing = (total_missing/total_cells)*100
print("Missing Values Count:")
print(missing_value_count)
print("Percentage of missing data:",percentage_missing)
df.tail()
# + id="qspMOaZXdkEE"
#Data type modification
df["DISTRICT"] = df["DISTRICT"].astype(str)
df["DISTRICT"].replace("3A","30",inplace=True)
df["DISTRICT"].replace("3C","31",inplace=True)
df["DISTRICT"].replace("3E","32",inplace=True)
df["DISTRICT"].replace("3G","33",inplace=True)
df["DISTRICT"].replace("3J","34",inplace=True)
df["DISTRICT"].replace("3L","35",inplace=True)
df["DISTRICT"].replace("3N","96",inplace=True)
df["DISTRICT"].replace("7","95",inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="nwHWqiGu4Yeq" outputId="7c4e55d3-5750-4f32-f9d9-586ab161e5a1"
df["PROSE"].fillna(0,inplace=True)
df["PROSE"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="wDM_Cq234WZ6" outputId="5c25065d-9239-4214-9d5f-c4a123c49ab0"
df["STATUSCD"].fillna("L",inplace=True)
df["STATUSCD"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="-bSwa86A3zV7" outputId="462fd3a1-3881-40eb-ac6c-7c8bf4ee1907"
df["DEMANDED"].fillna(0,inplace=True)
df["DEMANDED"].value_counts()
df["AMTREC"].fillna(0,inplace=True)
df["AMTREC"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="_pD80lXoENWr" outputId="e9c3f117-1cf2-4c7c-a1dd-fd68e7494687"
df["JURY"].fillna("N",inplace=True)
df["JURY"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="qxtAAgurFgOg" outputId="b4069484-6c50-4436-fdb3-48a771d7e057"
print("Unique values of target variable :-", df['LABEL'].unique())
# + colab={"base_uri": "https://localhost:8080/"} id="shfogZq6iBy8" outputId="250effd9-3a01-4646-b80a-bc67fd9a3d2b"
df["LABEL"].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="ofA6aU2fcXJz" outputId="69a9b6c3-abd4-4082-e4fe-4dfad64c7c97"
print("Unique values of target variable :-", df['DISTRICT'].unique())
# + colab={"base_uri": "https://localhost:8080/"} id="DJ2rc5xHHCcp" outputId="85c6a1f5-fe0e-4bc3-ad59-f573ddfcaa95"
print("Number of sample under each target value :- \n",df['LABEL'].value_counts())
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="YT2dAG3LX6GH" outputId="de187e31-1337-4433-c49d-a7bfaf2588aa"
noj_labels = 'No Montary Award','Monetery Award Only','Monetery Award and other','Injunction','Forefiet','Costs only','Costs and attorney fees'
#colours = ['silver']
plt.pie(df["LABEL"].value_counts(),startangle=90,autopct='%.1f%%')
plt.legend(noj_labels, loc="best")
plt.axis('equal')
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="1shMHWXR7JfR" outputId="198e5bd9-2771-41de-8fe6-9eda27c79320"
val = df["LABEL"].value_counts()
print(val)
# + colab={"base_uri": "https://localhost:8080/", "height": 483} id="KwWYCTqUbVMO" outputId="8970f3ec-368a-44b7-c5f3-97ab380879a9"
fig, ax = plt.subplots()
size = 1
vals = df["LABEL"]
cmap = plt.get_cmap("tab20c")
outer_colors = cmap(np.arange(3)*4)
inner_colors = cmap([1, 2, 3,4,5, 6, 7,8,9, 10])
inner_vals = []
for index,value in df["LABEL"].value_counts():
if index <= 10:
inner_vals[0]+=value
if index > 10 and index < 20:
inner_vals[1]+=value_counts
return inner_vals
ax.pie(vals.value_counts(), radius=2.5, colors=outer_colors,
wedgeprops=dict(width=size, edgecolor='w'))
ax.pie(vals.flatten(), radius=1-size, colors=inner_colors,
wedgeprops=dict(width=size, edgecolor='w'))
ax.set(aspect="equal", title='Pie plot with `ax.pie`')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 644} id="MUToiEuLn1X4" outputId="2f4cb0b6-db15-4860-ed57-edda10809aef"
plt.figure(figsize=(10,10))
sns.heatmap(data=df.corr(), cmap="seismic")
plt.show()
# + [markdown] id="U4WZYjBdyXJ_"
# Highest Correlation:
# PRO SE vs DOCKET
# PRO SE vs TAPEYEAR
#
# + id="pntjWWOGzWTz"
df.to_csv("Output.csv", index=False, header=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="SOExU5e31wuX" outputId="9de4c54b-df18-43dc-cf60-68bd1307f828"
from google.colab import files
files.download("Output.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 163} id="4YnhwmUnZ4Ah" outputId="45c7dd2f-2761-45bf-f10e-fee9687b1997"
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="Rc5HoUyb9WmR" outputId="0573ae14-31d4-4fca-c9a3-00e718ebe5c8"
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import feature_column
from tensorflow.keras import layers
from sklearn.model_selection import train_test_split
train, test = train_test_split(df, test_size=0.2)
train, val = train_test_split(train, test_size=0.2)
print(len(train), 'train examples')
print(len(val), 'validation examples')
print(len(test), 'test examples')
# + id="77Q4qq65g9tL"
def get_compiled_model():
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, activation='relu'),
tf.keras.layers.Dense(10, activation='relu'),
tf.keras.layers.Dense(1)
])
model.compile(optimizer='adam',
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
return model
# + id="S7_DLIdkhGd4"
model = get_compiled_model()
model.fit(train, epochs=5)
| Projects/ProjectLL/LL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src='./img/intel-logo.jpg' width=50%, Fig1>
#
# # OpenCV 기초강좌
#
# <font size=5><b>02. 이미지 채널<b></font>
#
# <div align='right'>성 민 석 (Minsuk Sung)</div>
# <div align='right'>류 회 성 (Hoesung Ryu)</div>
#
# <img src='./img/OpenCV_Logo_with_text.png' width=20%, Fig2>
#
#
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#들어가기-앞서" data-toc-modified-id="들어가기-앞서-1"><span class="toc-item-num">1 </span>들어가기 앞서</a></span></li><li><span><a href="#Color-space-변환" data-toc-modified-id="Color-space-변환-2"><span class="toc-item-num">2 </span>Color-space 변환</a></span><ul class="toc-item"><li><span><a href="#BGR--->-Grayscale" data-toc-modified-id="BGR--->-Grayscale-2.1"><span class="toc-item-num">2.1 </span>BGR --> Grayscale</a></span></li><li><span><a href="#BGR->HSV로-변환" data-toc-modified-id="BGR->HSV로-변환-2.2"><span class="toc-item-num">2.2 </span>BGR->HSV로 변환</a></span></li><li><span><a href="#수동으로-채널을-B,G,R-로-분리-한-후-다시-RGB로-합치기" data-toc-modified-id="수동으로-채널을-B,G,R-로-분리-한-후-다시-RGB로-합치기-2.3"><span class="toc-item-num">2.3 </span>수동으로 채널을 B,G,R 로 분리 한 후 다시 RGB로 합치기</a></span></li><li><span><a href="#BGR->HSV로-변환" data-toc-modified-id="BGR->HSV로-변환-2.4"><span class="toc-item-num">2.4 </span>BGR->HSV로 변환</a></span></li></ul></li></ul></div>
# -
# ## 들어가기 앞서
#
# 1. Grayscale Image
# <img src='./img/grayscale.png' width=90%>
# Pixel당 8bit, 즉 256단계의 명암(빛의 세기)을 표현할 수 있는 이미지입니다.
#
#
# 2. Color Image
#
# Color 이미지는 pixel의 색을 표현하기 위해서 pixel당 24bit를 사용합니다. 총 16,777,216 가지의 색을 표현할 수 있습니다. 이것을 일반적으로 True color image라고 합니다. pixel은 RGB 각각을 위해서 8bit를 사용하게 됩니다. OpenCV에서는 BGR로 표현을 하기 때문에 Blue->(255,0,0), Green->(0,255,0), Red->(0,0,255), White->(255,255,255), Black->(0,0,0)으로 표현할 수 있습니다.
#
# - RGB
# <img src='./img/rgb_image.png' width=100%>
# RGB 모델은 빛의 삼원색인 빨간색, 초록색, 파란색을 기본 색으로 사용을 합니다. 정육면체 모델 형태로 표현할 수 있습니다.<p>
#
# - HSV
# <img src='./img/HSV.jpeg' width=80%>
# 이미지 처리에서 가장 많이 사용되는 형태의 Color 모델입니다. 하나의 모델에서 색과 채도, 명도를 모두 알 수 있습니다.
#
# - H(ue) : 색상. 일반적인 색을 의미함. 원추모형에서 각도로 표현이 됨.(0: Red, 120도 : Green, 240: Blue)
# - S(aturation) : 채도. 색읜 순수성을 의미하며 일반적으로 짙다, 흐리다로 표현이 됨. 중심에서 바깥쪽으로 이동하면 채도가 높음.
# - V(alue) : 명도. 색의 밝고 어두운 정도. 수직축의 깊이로 표현. 어둡다 밝다로 표현이 됨.
# ## Color-space 변환
#
#
# `OpenCV` 에서는 이미지를 `BGR`로 읽으며, 150여가지 변환 방법이 있습니다. 그 중에 서 많이 사용되는 `BGR<->Gray`, `BGR<->RGB`, `BGR<->HSV` 에 대해서 알아 보겠습니다. 변환을 위해서 사용하는 함수는 `cv2.cvtColor()` 함수 입니다.
#
# ```
# cv2.cvtColor(src, code)
# Params src: image
# Params code: 변환 코드
# ```
# ### BGR --> Grayscale
#
# `cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)`
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import cv2
# 원본그대로 불러오기
image = cv2.imread("./img/toy.jpg")
# +
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
plt.title("gray_image")
plt.imshow(gray,cmap='gray')
plt.xticks([]) # x축 눈금 없애기
plt.yticks([]) # y축 눈금 없애기
plt.show()
# -
# ### BGR->HSV로 변환
# `cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)`
#
# +
rgb = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
plt.title("rgb_image")
plt.imshow(rgb)
plt.xticks([]) # x축 눈금 없애기
plt.yticks([]) # y축 눈금 없애기
plt.show()
# -
# ### 수동으로 채널을 B,G,R 로 분리 한 후 다시 RGB로 합치기
# +
# B,G,R 로 분리
b, g, r = cv2.split(image)
# RGB 로 다시 합치기
manual_image =cv2.merge([r,g,b])
plt.title("rgb_image")
plt.imshow(manual_image)
plt.xticks([]) # x축 눈금 없애기
plt.yticks([]) # y축 눈금 없애기
plt.show()
# -
# ### BGR->HSV로 변환
#
# `cv2.cvtColor(frame, cv2.COLOR_BGR2HSV)`
#
# +
hsv = cv2.cvtColor(image, cv2.COLOR_BGR2HSV)
plt.title("HSV_image")
plt.imshow(hsv)
plt.xticks([]) # x축 눈금 없애기
plt.yticks([]) # y축 눈금 없애기
plt.show()
| opencv/OpenCV Lecture02. Image-Channel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from scipy.optimize import fmin_tnc
# %matplotlib inline
from matplotlib import pyplot as plt
# ==================== Load Data ====================
data = pd.read_table('data\\week3\\ex2data1.txt', sep=',', header=None, names=['X_col1', 'X_col2', 'Y_col3'], dtype={'X_col1': np.float64, 'X_col2': np.float64, 'Y_col3': np.uint8})
data.head()
X = data.loc[:, ['X_col1', 'X_col2']].to_numpy()
y = data.loc[:, 'Y_col3'].to_numpy()
# ==================== Part 1: Plotting ====================
# +
fig, ax = plt.subplots()
ax.scatter(np.take(X[:, 0], np.argwhere(y==1)), np.take(X[:, 1], np.argwhere(y==1)), c='black', marker='+', label='Admitted')
ax.scatter(np.take(X[:, 0], np.argwhere(y==0)), np.take(X[:, 1], np.argwhere(y==0)), c='yellow', marker='o', facecolors='none', edgecolors='black', label='Not admitted')
ax.set_xlabel('Exam 1 score', size=10)
ax.set_ylabel('Exam 2 score', size=10)
ax.grid(True)
ax.legend(fontsize=10)
ax.set_xlim([30, 100])
ax.set_ylim([30, 100])
plt.show()
# -
# ============ Part 2: Compute Cost and Gradient ============
# 1 / (1 + exp(-h))
def sigmoid(z):
return 1.0 / (1.0 + np.exp(-z))
def costFunction(theta, X, y):
theta = theta.reshape(theta.size, 1)
hypo = sigmoid(np.matmul(X, theta))
J = (1.0 / y.size) * (np.matmul(-1.0 * y.T, np.log(hypo)) - np.matmul((1.0 - y.T), np.log(1.0 - hypo)))
grad = (1.0 / y.size) * np.matmul((hypo - y).T, X)
return J.item(), grad.T.flatten()
initial_theta = np.zeros((X.shape[1] + 1,), dtype=np.float64)
X1 = np.hstack((np.ones((X.shape[0], 1), dtype=np.float64), X))
cost, grad = costFunction(initial_theta, X1, y.reshape(y.size, 1))
print('Cost at initial theta (zeros): {:.6f}\nExpected cost (approx): 0.693'.format(cost))
print('Gradient at initial theta (zeros): \n {}'.format('\n '.join('{:.6f}'.format(item) for item in grad)))
print('Expected gradients (approx):\n -0.1000\n -12.0092\n -11.2628\n')
test_theta = np.array([-24, 0.2, 0.2], dtype=np.float64)
cost, grad = costFunction(test_theta, X1, y.reshape(y.size, 1))
print('Cost at test theta: {:.6f}\nExpected cost (approx): 0.218'.format(cost))
print('Gradient at test theta: \n {}'.format('\n '.join('{:.6f}'.format(item) for item in grad)))
print('Expected gradients (approx):\n 0.043\n 2.566\n 2.647\n')
# ============= Part 3: Optimizing using fminunc =============
minimization = fmin_tnc(func=costFunction, x0=initial_theta, args=(X1, y.reshape(y.size, 1)))
cost, grad = costFunction(minimization[0], X1, y.reshape(y.size, 1))
print('Cost at theta found by fmin_tnc: {:.6f}\nExpected cost (approx): 0.203'.format(cost))
print('theta: \n {}'.format('\n '.join('{:.6f}'.format(item) for item in minimization[0])))
print('Expected theta (approx):\n -25.161\n 0.206\n 0.201\n')
plot_X = np.array([np.min(X1[:, 1]) - 2, np.max(X1[:, 1]) + 2], dtype=np.float64)
plot_Y = (-1.0 / minimization[0][2]) * (minimization[0][1] * plot_X + minimization[0][0])
# +
fig, ax = plt.subplots()
ax.scatter(np.take(X1[:, 1], np.argwhere(y==1)), np.take(X1[:, 2], np.argwhere(y==1)), c='black', marker='+', label='Admitted')
ax.scatter(np.take(X1[:, 1], np.argwhere(y==0)), np.take(X1[:, 2], np.argwhere(y==0)), c='yellow', marker='o', facecolors='none', edgecolors='black', label='Not admitted')
ax.plot(plot_X, plot_Y, label='Decision Boundary')
ax.set_xlabel('Exam 1 score', size=10)
ax.set_ylabel('Exam 2 score', size=10)
ax.grid(True)
ax.legend(fontsize=10)
ax.set_xlim([30, 100])
ax.set_ylim([30, 100])
plt.show()
# -
# ============== Part 4: Predict and Accuracies ==============
prob = sigmoid(np.matmul(np.array([1, 45, 85], dtype=np.float64), minimization[0]))
print('For a student with scores 45 and 85, we predict an admission probability of {:.6f}\nExpected value: 0.775 +/- 0.002'.format(prob))
def predict(theta, X):
return sigmoid(np.matmul(X, theta)).round()
p = predict(minimization[0], X1)
print('Train Accuracy: {:.6f}\nExpected accuracy (approx): 89.0'.format(np.mean((p==y).astype(np.float64)) * 100))
| week3_ex2.ipynb |
# ---
# title: "Automate visuzliation with unique()"
# date: 2020-04-12T14:41:32+02:00
# author: "<NAME>"
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import pandas as pd
summer_2016_medals = pd.read_csv('summer2016.csv', index_col=0)
# -
# ### Unique values of a column
# +
# Extract the "Sport" column
sports_column = summer_2016_medals['Sport']
# Find the unique values of the "Sport" column
sports = sports_column.unique()
# Print out the unique sports values
print(sports)
# -
# ### Automate plotting
# +
fig, ax = plt.subplots()
# Loop over the different sports branches
for sport in sports:
# Extract the rows only for this sport
sport_df = summer_2016_medals[summer_2016_medals['Sport'] == sport]
# Add a bar for the "Weight" mean with std y error bar
ax.bar(sport, sport_df['Weight'].mean(), yerr=sport_df['Weight'].std())
ax.set_ylabel("Weight")
ax.set_xticklabels(sports, rotation=90)
# Save the figure to file
fig.savefig('sports_weights.png')
| courses/datacamp/notes/python/matplotlibTMP/automateplot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocessing (imputation, standardization, final clean) and modeling
# sklearn modeling the median imputed training data using min/max clinically guided aggregation.
# preprocessing includes variable formatting (categorical to factor variables in r, train/test split, and median imputation).
#
# +
import pandas as pd
import matplotlib.pyplot as plt
import os, sys
from pathlib import Path
import seaborn as sns
import numpy as np
import glob
from sklearn.metrics import confusion_matrix, classification_report, roc_curve, roc_auc_score, accuracy_score, auc, precision_recall_fscore_support, pairwise, f1_score, log_loss, make_scorer
from sklearn.metrics import precision_score, recall_score
from sklearn import metrics
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
from sklearn.externals.joblib import Memory
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder, Imputer
from sklearn.model_selection import StratifiedKFold, GridSearchCV, RandomizedSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.utils import validation
from scipy.sparse import issparse
from scipy.spatial import distance
from sklearn import svm
#importin xg boost and all needed otherstuff
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from xgboost import XGBClassifier #conda install -c conda-forge xgboost to install
##adding these, lets see if it helps with xgboost crash
os.environ['KMP_DUPLICATE_LIB_OK']='True'
#reducing warnings that are super common in my model
import warnings
from sklearn.exceptions import DataConversionWarning
warnings.simplefilter(action='ignore') #ignore all warnings
#memory = Memory(cachedir='/tmp', verbose=0)
#@memory.cache above any def fxn.
RANDOM_STATE = 15485867
# %matplotlib inline
plt.style.use('seaborn-white')
from notebook.services.config import ConfigManager
cm = ConfigManager()
cm.update('livereveal', {
'width': 1024,
'height': 768,
'scroll': True,
})
# %load_ext autotime
# -
# ## importing datasets and doing final formatting/processing
# +
#patients of interest from rotation_cohort_generation
from parameters import final_pt_df_v, date, repository_path, lower_window, upper_window, folder, date, time_col, time_var, patient_df, categorical, save_boolean
#patients of interest from rotation_cohort_generation
final_pt_df2 = final_pt_df_v #pd.read_csv('/Users/geickelb1/Documents/GitHub/mimiciii-antibiotics-modeling/data/raw/csv/%s_final_pt_df2.csv'%(most_updated_patient_df), index_col=0)
del(final_pt_df_v)
patients= list(final_pt_df2['subject_id'].unique())
hadm_id= list(final_pt_df2['hadm_id'].unique())
icustay_id= list(final_pt_df2['icustay_id'].unique())
icustay_id= [int(x) for x in icustay_id]
# -
len(patients)
len(icustay_id)
folder
# +
# ###########
# save_boolean=False
# ############
# -
address=str(repository_path)+'/data/cleaned_merged_agg/'+'%s/'%(folder)
train_data= pd.read_csv(Path(address+'%s_%s_cleaned_merged_agg.csv' %(date, 'train')), index_col=0)
test_data= pd.read_csv(Path(address+'%s_%s_cleaned_merged_agg.csv' %(date, 'test')), index_col=0)
### added 11/8/19: removed daily sofa score from prediction model, since the components used to calculate it are already in model
### 12/12/19: #removing o2_flow since it is mostly sparce and is not a helpful variable
train_data.drop('daily_sofa', axis=1, inplace=True)
test_data.drop('daily_sofa', axis=1, inplace=True)
train_data.drop('o2_flow', axis=1, inplace=True)
test_data.drop('o2_flow', axis=1, inplace=True)
categorical.remove('o2_flow')
train_data['pao2fio2ratio'].head()
# ## getting descriptive numbers for publication
final_pt_df2.head()#['icustay_id'].nunique()
final_pt_df2.loc[final_pt_df2['final_bin'].isin(['C_neg/A_partial','C_pos/A_full']),'subject_id'].nunique()
list(train_data)
full_demo=pd.concat([train_data[['icustay_id','yearsold','ethnicity','gender']],test_data[['icustay_id','yearsold','ethnicity','gender']]])
full_demo.describe()
full_demo.head()
full_demo.merge(final_pt_df2[['icustay_id','subject_id','final_bin']])['final_bin'].value_counts()
full_demo['yearsold'].min()
full_demo['ethnicity'].value_counts()
full_demo['ethnicity'].value_counts()
full_demo['gender'].value_counts()
test_data[['icustay_id','yearsold','ethnicity','gender']].head()
train_data.merge(final_pt_df2[['icustay_id','subject_id','final_bin']])['final_bin'].value_counts()
test_data.merge(final_pt_df2[['icustay_id','subject_id','final_bin']])['final_bin'].value_counts()
final_pt_df2['final_bin'].value_counts()
# ## back to the formatting and final preprocessing
def rename_fxn(data):
"""
renames data columns to be more readable.
"""
rename_dic={
"('max', 'sodium')": "maxSodium" ,
"('max', 'sodium')" : "maxSodium",
"('min', 'sodium')" : "minSodium",
"('max', 'calcium')" : "maxCalcium",
"('min', 'calcium')" : "minCalcium",
"('max', 'sodium')": "maxSodium",
"('min', 'sodium')": "minSodium",
"('max', 'wbc')": "maxWBC",
"('min', 'wbc')": "minWBC"#,
# "bands": "ibands",
# "pco2": "ipco2"
}
data=data.rename(rename_dic, axis='columns').copy()
return(data)
def last_minute_clean(data):
"some last minute data cleaning steps that were missed prior to this. eventually should be transitioned to earlier in pipeline"
data.loc[data['ethnicity'].isnull(), 'ethnicity']='unknown/other'
data.loc[data['yearsold']>90,"yearsold"]=90
return(data)
def standardize_fxn(data):
"""
function operations:
2) standardize last 2 columns to be standardized (weight and age)
"""
from sklearn.impute import SimpleImputer
### I chose to standardize the weight and age here (after the feature_aggregation)... I can't remember why though. either way, it doesn't affect anything, but is a bit clunky.
weight_median=np.log(data.loc[data['final_bin']=="C_neg/A_partial","weight"]+1).median()
weight_quant1=np.log(data.loc[data['final_bin']=="C_neg/A_partial","weight"]+1).quantile(0.25)#.between(train_data['col'].quantile(.25), df['col'].quantile(.75), inclusive=True)]
weight_quant3=np.log(data.loc[data['final_bin']=="C_neg/A_partial","weight"]+1).quantile(0.75)
weight_iqr=weight_quant3-weight_quant1
#print(weight_median,weight_quant3,weight_quant1, weight_iqr)
age_median=np.log(data.loc[data['final_bin']=="C_neg/A_partial","yearsold"]+1).median()
age_quant1=np.log(data.loc[data['final_bin']=="C_neg/A_partial","yearsold"]+1).quantile(0.25)
age_quant3=np.log(data.loc[data['final_bin']=="C_neg/A_partial","yearsold"]+1).quantile(0.75)
age_iqr=age_quant3-age_quant1
#converting to log scaled standardized data for age/weight
data['weight']=data['weight'].apply(lambda x: (np.log(x+1)-weight_median)/weight_iqr)
data['yearsold']=data['yearsold'].apply(lambda x: (np.log(x+1)-age_median)/age_iqr)
return(data)
def remap_categorical(data, categorical):
"""remapps the 0 and 1 back to interpretable values in the dataframe. useful to use before onehot encoding. also useful to fill in some missing values and nan's"""
remapping_dic={
'vent_recieved':{0:'None', 2:'Mech' , 1:'OxygenTherapy', np.nan:'None'},
'cancer_elix':{0:'no_cancer_cormorb', 1:'cancer_comorbidity'},
'gender':{0:'F', 1:'M', np.nan:'NA'},
'leukocyte':{0:'Neg/Not_tested' , 1:'pos'},
'nitrite':{0:'Neg/Not_tested' , 1:'pos'},
'rrt':{0:'not_recieved' , 1:'recieved'},
'phenylephrine':{0:'not_recieved' , 1:'recieved'},
'norepinephrine':{0:'not_recieved' , 1:'recieved'},
'vasopressin':{0:'not_recieved' , 1:'recieved'},
'dobutamine':{0:'not_recieved' , 1:'recieved'},
'dopamine':{0:'not_recieved' , 1:'recieved'},
'epinephrine':{0:'not_recieved' , 1:'recieved'},
'any_vasoactives':{0:'not_recieved' , 1:'recieved'},
'o2_flow':{0:'no_flow', 1:"recieved_O2"}
}
for col in list(data):
if col in ['ethnicity', 'bands','pco2']:
pass
elif col in categorical:
data[col]=data[col].replace(remapping_dic[col])
return(data)
def onehotencoder(data, categorical):
"""
function operations:
3) convert categorical columns to proper format
"""
data = pd.get_dummies(data, columns = categorical, drop_first=True)
return(data)
def median_imputer(data, categorical):
"""
function operations:
4) median impute
"""
from sklearn.impute import SimpleImputer
#pop outcome off
outcome= data.pop("final_bin").values
#impute
imp = SimpleImputer(missing_values=np.nan, strategy='median')
imp.fit(data[[x for x in list(data) if x not in categorical]])
data[[x for x in list(data) if x not in categorical]]= imp.transform(data[[x for x in list(data) if x not in categorical]])
#reattach outcome
data['final_bin']= outcome
return(data)
def preprocessing(data, encode=True):
"""
function operations: (each one of the below steps uses one of the functions defined above.)
1) rename columns
2) some last minute cleanup
3) standardize last 2 columns to be standardized
4) median impute
5) remap categorical values to be interpretable and encode them for modeling
6)
"""
from sklearn.impute import SimpleImputer
###rename
data=rename_fxn(data)
###some last minute data cleaning steps that were missed prior to this. eventually should be transitioned to earlier in pipeline
data=last_minute_clean(data)
##scale & standardize the remaining 2 columns (age and weight)
data=standardize_fxn(data)
### onehot encoding categorical var
#remapping the 0 and 1 back to categorical values for more interpreable onehotencoding column headings
data=remap_categorical(data, categorical)
#impute
data= median_imputer(data, categorical)
#onehotencoding
if encode==True:
data=onehotencoder(data, categorical)
#binarizing and poping outcome for training data
data.loc[data['final_bin']=="C_pos/A_full","final_bin"]=1
data.loc[data['final_bin']=="C_neg/A_partial","final_bin"]=0
data['final_bin']=pd.to_numeric(data['final_bin'])
## establishing training data and labels
x= data.copy()
z_icustay_id=x.pop('icustay_id')
y= x.pop("final_bin").values
return(x, y, z_icustay_id)
# # todo: get some statistics on missingness right before i perform imputation.
# +
x_train, y_train, z_icustay_id = preprocessing(pd.merge(train_data, final_pt_df2[['icustay_id','final_bin']]))
x_test, y_test, z_icustay_id_test= preprocessing(pd.merge(test_data, final_pt_df2[['icustay_id','final_bin']]))
z_subject_id=pd.merge(pd.DataFrame(z_icustay_id), final_pt_df2[['icustay_id','subject_id']], how='left')['subject_id'] #7205
# -
y.reshape(1,-1).ravel()
x_train2, y_train2, z_icustay_id2 = preprocessing(pd.merge(train_data, final_pt_df2[['icustay_id','final_bin']]),encode=False)
x_test2, y_test2, z_icustay_id_test2= preprocessing(pd.merge(test_data, final_pt_df2[['icustay_id','final_bin']]),encode=False)
###checking distribution of all categorical variables
full_set=pd.concat([x_train2,x_test2])
for element in categorical:
print("###{}###".format(element))
print(full_set[element].value_counts(), '\n')
len(full_set)
# #### quick qc to ensure the imputation and standardization of age/weight worked
train_data[['chloride','weight','yearsold']].head()
x_train[['chloride','weight','yearsold']].head()
len(list(x_train))
# ### getting table for paper
list(x_train)
pd.merge(train_data['icustay_id'], final_pt_df2[['icustay_id','final_bin']])['final_bin'].value_counts()
pd.merge(test_data['icustay_id'], final_pt_df2[['icustay_id','final_bin']])['final_bin'].value_counts()
# ## looking at correlation of all variables
# +
corr = x_train.corr().abs()
plt.figure(figsize=(25, 20))
ax = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=45,
horizontalalignment='right'
);
# -
sol = (corr.where(np.triu(np.ones(corr.shape), k=1).astype(np.bool)).stack().sort_values(ascending=False))
cor_df=pd.DataFrame(sol)#.sort_values(kind="quicksort") #[-10:0])
cor_df=cor_df.reset_index()
cor_df=cor_df.rename(columns={'level_0': 'corx', 'level_1': 'cory', 0:'corr'})
cor_df2=cor_df[(cor_df['corx']!=cor_df['cory']) & (cor_df['corr']>0.7)].sort_values('corr', ascending=False)
cor_df2.head()
# ### DROPING one of the 2 columns with correlation >0.7 (depreciated)
# +
# x_train.drop(columns=list(cor_df2['corx']), inplace=True, errors='raise')
# x_test.drop(columns=list(cor_df2['corx']), inplace=True, errors='raise')
# -
# ### formatting x and y for modleing
x=np.array(x_train.copy())
y=y_train.copy() #copy of y_train
y=y.astype('int')
#time_interval=4
print(len(x_train),len(x_test))
y_train
len(list(x_train))
def save_df(df, df_name='default', rel_path='/data/final/'):
"""
simple function for saving result table. uses the date and supplied df name and saves to the savepath specified above.
"""
global folder
save_path= str(repository_path)+rel_path
address=save_path+'{}_{}/'.format(date,folder)
if not os.path.exists(address):
print(address)
os.makedirs(address)
else:
print(address)
if address.endswith('/')==False:
address= address+'/'
if df_name == 'default':
df_name =[x for x in globals() if globals()[x] is df][0]
pd.DataFrame(df).to_csv(Path(address+'{}_{}.csv'.format(date, df_name)))
if save_boolean==True:
save_df(x_train, 'x_train_final', rel_path='/data/final/')
save_df(x_test, 'x_test_final', rel_path='/data/final/')
save_df(y_train, 'y_train_final', rel_path='/data/final/')
save_df(y_test, 'y_test_final', rel_path='/data/final/')
save_df(z_icustay_id, 'icustay_train', rel_path='/data/final/') #just added, need to rerun all models. 12-19-19
save_df(z_icustay_id_test, 'icustay_test', rel_path='/data/final/')
# # Modelbuilding
# ## test entire trainset and predict trainset.
# * step1) hypertune and evaluate models on 10fold cv.
# * step2) test entire train set and predict testset.
# * step3) generate figures
from parameters import nfolds, scoring, n_iter, gridsearch
##defaults
# nfolds=10
# scoring='roc_auc'
# n_iter=20 #for gridsearch
# gridsearch=False #gridsearch=False means it does triaged hyperparameter combinations based on some algorithm. True= tests all
# +
def evaluate(model, x, y):
"simple classification evaluation metrics and output used in my hypertuning functions"
from sklearn.metrics import log_loss
y_hat = model.predict(x)
y_hat_proba = model.predict_proba(x)[:, 1]
errors = abs(y_hat - y)
mape = 100 * np.mean(errors / y)
accuracy = 100 - mape
auc=roc_auc_score(y, y_hat_proba)
loss= log_loss(y, y_hat_proba)
print ('the AUC is: {:0.3f}'.format(auc))
print ('the logloss is: {:0.3f}'.format(loss))
print(confusion_matrix(y, y_hat))
print(classification_report(y,y_hat, digits=3))
if scoring=='neg_log_loss':
return_value=loss
elif scoring=='roc_auc':
return_value=auc
else:
raise ValueError
return (return_value)
# -
def hypertuning_fxn(X, y, nfolds, model , param_grid, scoring=scoring, gridsearch=True, n_iter=20, verbose=False):
from sklearn.model_selection import GroupKFold
np.random.seed(12345)
if gridsearch==True:
grid_search = GridSearchCV(estimator= model,
param_grid=param_grid,
cv=GroupKFold(nfolds),
scoring=scoring,
return_train_score=True,
n_jobs = -1)
else:
grid_search = RandomizedSearchCV(estimator= model,
param_distributions= param_grid,
n_iter=n_iter,
cv=GroupKFold(nfolds),
scoring=scoring,
return_train_score=True,
random_state=12345,
n_jobs = -1)
grid_search.fit(X, y, groups=z_subject_id)
print(" scorer function: {}".format(scoring))
print(" ##### CV performance: mean & sd scores #####")
means = grid_search.cv_results_['mean_test_score']
stds = grid_search.cv_results_['std_test_score']
print('best cv score: {:0.3f}'.format(grid_search.best_score_))
print('best cv params: ', grid_search.best_params_)
worst_index=np.argmin(grid_search.cv_results_['mean_test_score'])
print('worst cv score: {:0.3f}'.format(grid_search.cv_results_['mean_test_score'][worst_index]))
print('worst cv params: ', grid_search.cv_results_['params'][worst_index])
##
if verbose==True:
for mean, std, params in zip(means, stds, grid_search.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"% (mean, std * 2, params))
print('##### training set performance #####\n')
print(' best hypertuned model training set performance:')
best_random = grid_search.best_estimator_
best_random_auc = evaluate(best_random, x, y)
print(' worst hypertuned model training set performance:')
worst_params= grid_search.cv_results_['params'][worst_index]
worst_random=model.set_params(**worst_params)
worst_random.fit(x,y)
worst_random_auc = evaluate(worst_random, x, y)
print('relative scorer change of {:0.2f}%. between worst and best hyperparams on TRAINING set (may be overfit)'.format( 100 * (best_random_auc - worst_random_auc) / worst_random_auc))
return(grid_search)
def hypertuned_cv_fxn(x, y, model_in, nfolds):
"""
the goal of this function is to take the best hypertuned model and
generate average and std for F-1, precision, recall, npv, and AUC across each fold.
Ideally i could have generated this above in my hypertuning cv function,
but it actually took less computational time to just rerun cv on the best performing evaluator and collect all of the averaged performance metrics
"""
from sklearn.model_selection import GroupKFold
import sklearn.metrics as metrics
from sklearn.metrics import precision_score, roc_auc_score, f1_score, recall_score
from sklearn.base import clone
pos_label=1
model= clone(model_in, safe=True)
np.random.seed(12345)
group_kfold = GroupKFold(n_splits=nfolds)
group_kfold.get_n_splits(x, y, z_subject_id)
f1_y_cv=[]
auc_y_cv=[]
prec_y_cv=[]
recall_y_cv=[]
npv_y_cv=[]
for train_index, test_index in group_kfold.split(x, y, z_subject_id):
x_train_cv, x_test_cv = x[train_index], x[test_index]
y_train_cv, y_test_cv = y[train_index], y[test_index]
model.fit(x_train_cv, y_train_cv)
y_proba = model.predict_proba(x_test_cv)[:,1]
y_pred = model.predict(x_test_cv)
fpr, tpr, thresholds = metrics.roc_curve(y_test_cv, y_proba, pos_label=pos_label)
#gathering the optimal youden_index and df of tpr/fpr for auc and index of that optimal youden. idx is needed in the roc
youden_threshold, roc_df, idx= optimal_youden_index(fpr, tpr, thresholds,tp90=True)
y_pred_youden = [1 if y >= youden_threshold else 0 for y in y_proba]
npv_y=confusion_matrix(y_test_cv, y_pred_youden)[0,0]/sum(np.array(y_pred_youden)==0)
npv_y_cv.append(npv_y)
prec_y= precision_score(y_true=y_test_cv, y_pred= y_pred_youden, pos_label=pos_label)
prec_y_cv.append(prec_y)
recall_y= recall_score(y_true=y_test_cv, y_pred= y_pred_youden, pos_label=pos_label)
recall_y_cv.append(recall_y)
f1_y= f1_score(y_true=y_test_cv, y_pred= y_pred_youden, pos_label=pos_label)
f1_y_cv.append(f1_y)
###need to debug this.###
auc_y=roc_auc_score(y_true=y_test_cv, y_score= y_proba)
auc_y_cv.append(auc_y)
youden_dic_cv= {'model':type(model).__name__,
'auc':np.mean(auc_y_cv),
'auc_sd':np.std(auc_y_cv),
'precision':np.mean(prec_y_cv),
'precision_sd':np.std(prec_y_cv),
'recall':np.mean(recall_y_cv),
'recall_sd':np.std(recall_y_cv),
'f1':np.mean(f1_y_cv),
'f1_sd':np.std(f1_y_cv),
'npv':np.mean(npv_y_cv),
'npv_sd':np.std(npv_y_cv)}
return(youden_dic_cv)
# ## youden index and plotting functions
# +
def saveplot(plt, figure_name):
"""
simple function for saving plots
"""
address = str(repository_path)+'/figures/{}_{}'.format(date,folder)
print(address)
if not os.path.exists(address):
os.makedirs(address)
plt.savefig(address+"/{}.png".format(figure_name),bbox_inches='tight')
def optimal_youden_index(fpr, tpr, thresholds, tp90=True):
"""
inputs fpr, tpr, thresholds from metrics.roc(),
outputs the clasification threshold, roc dataframe, and the index of roc dataframe for optimal youden index
"""
#making dataframe out of the thresholds
roc_df= pd.DataFrame({"thresholds": thresholds,"fpr":fpr, "tpr": tpr})
roc_df.iloc[0,0] =1
roc_df['yuden']= roc_df['tpr']-roc_df['fpr']
if tp90==True:
idx= roc_df[roc_df['tpr']>=0.9]['yuden'].idxmax() #changed this so now finds optimial yuden threshold but tp>=90%
else:
idx=roc_df['yuden'].idxmax() #MAX INDEX
youden_threshold=roc_df.iloc[idx,0] #threshold for max youden
return(youden_threshold, roc_df, idx)
def plot_roc(fpr, tpr, roc_auc, roc_df, idx, save=save_boolean,model_name=None, folder_name=None, file_name=None):
plt.title('ROC with optimal Youden Index')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
#finding the point on the line given threshold 0.5 (finding closest row in roc_df)
og_idx=roc_df.iloc[(roc_df['thresholds']-0.5).abs().argsort()[:1]].index[0]
plt.plot(roc_df.iloc[og_idx,1], roc_df.iloc[og_idx,2],marker='o', markersize=5, color="g")
plt.annotate(s="P(>=0.5)",xy=(roc_df.iloc[og_idx,1]+0.02, roc_df.iloc[og_idx,2]-0.04),color='g') #textcoords
plt.plot(roc_df.iloc[idx,1], roc_df.iloc[idx,2],marker='o', markersize=5, color="r") ##
plt.annotate(s="TPR>=0.9",xy=(roc_df.iloc[idx,1]+0.02, roc_df.iloc[idx,2]-0.04),color='r' ) #textcoords
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.grid(color='grey', linestyle='-', linewidth=1, alpha=0.2)
if save==True:
saveplot(plt, figure_name="{}_roc".format(model_name))
else: pass
plt.show()
# -
def classifier_eval(model, x=x, y=y, proba_input=False,pos_label=1, print_default=True,model_name=None, folder_name=None, save=save_boolean):
import sklearn.metrics as metrics
from sklearn.metrics import precision_score, roc_auc_score, f1_score, recall_score
"""
classification evaluation function. able to print/save the following:
print/save the following:
ROC curve marked with threshold for optimal youden (maximizing tpr+fpr with constraint that tpr>0.9)
using 0.5 threshold:
confusion matrix
classification report
npv
accuracy
using optimal youden (maximizing tpr+fpr with constraint that tpr>0.9):
confusion matrix
classification report
npv
accuracy
output:
outputs modelname, auc, precision, recall, f1, and npv to a dictionary.
notes:
youden's J statistic:
J= sensitivity + specificity -1
(truepos/ truepos+falseneg) + (true neg/ trueneg + falsepos) -1.
NOTE: with tpr>0.9 turned on, the youden statistic is basically just the furthest point on the line away from the midline with tpr>=0.9
NOTE2: this function arguably does too much. in the future it may be better to seperate it out into more compartmental functions like with preprocessing().
"""
if proba_input==True:
y_proba= model
y_pred=[1 if y >= 0.5 else 0 for y in y_proba]
else:
model_name=type(model).__name__
y_pred = model.predict(x)
y_proba = model.predict_proba(x)[:,1]
fpr, tpr, thresholds = metrics.roc_curve(y, y_proba, pos_label=pos_label)
roc_auc = metrics.auc(fpr, tpr)
# print("AUROC:",roc_auc)
#gathering the optimal youden_index and df of tpr/fpr for auc and index of that optimal youden. idx is needed in the roc
youden_threshold, roc_df, idx= optimal_youden_index(fpr, tpr, thresholds,tp90=True)
#plotting roc
plot_roc(fpr, tpr, roc_auc, roc_df, idx, save=save, model_name=model_name,folder_name=folder)
plt.show(), plt.close()
#printing npv, recall, precision, accuracy
npv=confusion_matrix(y, y_pred)[0,0]/sum(np.array(y_pred)==0)
prec= precision_score(y_true=y, y_pred= y_pred, pos_label=pos_label)
recall= recall_score(y_true=y, y_pred= y_pred, pos_label=pos_label)
f1= f1_score(y_true=y, y_pred= y_pred, pos_label=pos_label)
if print_default==True: ###can opt to not print the 0.5 classification threshold classification report/conf matrix
#plotting confusion matrixs
print("\n******* Using 0.5 Classification Threshold *******\n")
print(confusion_matrix(y, y_pred))
print ('the Accuracy is: {:01.3f}'.format(accuracy_score(y, y_pred)))
print ("npv: {:01.3f}".format(npv))
print ('the classification_report:\n', classification_report(y,y_pred, digits=3))
else:
pass
#### YOUDEN ADJUSTMENT #####
print("\n******* Using Optimal Youden Classification Threshold *******\n")
print("\nthe Youden optimal index is : {:01.3f}".format(youden_threshold))
y_pred_youden = [1 if y >= youden_threshold else 0 for y in y_proba]
npv_y=confusion_matrix(y, y_pred_youden)[0,0]/sum(np.array(y_pred_youden)==0)
prec_y= precision_score(y_true=y, y_pred= y_pred_youden, pos_label=pos_label)
recall_y= recall_score(y_true=y, y_pred= y_pred_youden, pos_label=pos_label)
f1_y= f1_score(y_true=y, y_pred= y_pred_youden, pos_label=pos_label)
auc_y=roc_auc_score(y_true=y, y_score= y_proba)
##plotting and saving confusion matrix
confusion_youden=confusion_matrix(y, y_pred_youden)
#plotting confusion matrixs
print(confusion_matrix(y, y_pred_youden))
print ('the Accuracy is: {:01.3f}'.format(accuracy_score(y, y_pred_youden)))
print ("npv: {:01.3f}".format(npv_y))
print ('the classification_report:\n', classification_report(y,y_pred_youden, digits=3))
youden_dic= {'model':model_name, 'auc':auc_y, 'precision':prec_y, 'recall':recall_y, 'f1':f1_y, 'npv':npv_y}
return(youden_dic)
# ## step1) hypertuning and cv
# * xgboost
# * randomforest
# * support vector classifier
# * logistic regression (l1 and l2)
# * k-nearest neighbors
# #### xgboost hypertune
###xgboost
model= XGBClassifier(n_estimators=100, min_child_weight=2, #changed: GridSearchCV ->RandomizedSearchCV
gamma=0, subsample=0.8, colsample_bytree=0.8,
objective='binary:logistic', n_jobs=-1, seed=27)
scale_pos_weight = [1, 5, 10] #0.1
max_depth = [1, 2, 3, 4, 5]
learning_rate=[0.01, 0.1, 0.5, 1]
param_grid = {'scale_pos_weight': scale_pos_weight, 'max_depth' : max_depth, "learning_rate":learning_rate}
xgboost_hyper=hypertuning_fxn(x, y, nfolds=nfolds, model=model , param_grid=param_grid, scoring=scoring, n_iter=n_iter, gridsearch=gridsearch)
y.size
sum(y)
xgboost_cv= hypertuned_cv_fxn(x, y, xgboost_hyper.best_estimator_, nfolds=nfolds)
xgboost_cv
# #### RF hypertune
# +
###rf
#{'bootstrap': False, 'class_weight': None, 'max_depth': 25, 'max_features': 'auto', 'min_samples_leaf': 2, 'min_samples_split': 10, 'n_estimators': 200}
# Number of trees in random forest
n_estimators = [10,15, 25, 50, 150, 250] #[int(x) for x in np.linspace(start = 10, stop = 1000, num = 5)]
# Number of features to consider at every split
max_features = [3,10,20,'auto']
# Maximum number of levels in tree
max_depth = [5,10, 25]#[int(x) for x in np.linspace(5, 110, num = 5)]
#max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10]
# Minimum number of samples required at each leaf node
min_samples_leaf = [2, 5, 10]
# Method of selecting samples for training each tree. supposedly better with false when classes aren't perfectly ballanced
bootstrap = [True, False] #[True, False] #[False]
#### note about bootstrap= off
###Without bootstrapping, all of the data is used to fit the model, so there is not random variation between trees with respect to the selected examples at each stage.
###However, random forest has a second source of variation, which is the random subset of features to try at each split.
### The documentation states "The sub-sample size is always the same as the original input sample size but the samples are drawn with replacement if bootstrap=True (default),"
### which implies that bootstrap=False draws a sample of size equal to the number of training examples without replacement, i.e. the same training set is always used.
### we test this turned off because with unballanced classes turning this off can sometimes improve performance
#class_weight is either a dictionary of each class to a uniform weight for that class (e.g., {1:.9, 2:.5, 3:.01}), or is a string telling sklearn how to automatically determine this dictionary.
class_weight= [None, {0:(1/np.bincount(y))[0], 1:(1/np.bincount(y))[1]}]
param_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'class_weight': class_weight,
'bootstrap': bootstrap}
model= RandomForestClassifier(criterion='entropy', random_state=12345)
#rf_hyper=hypertuning_fxn(x, y, nfolds=nfolds, model=model , param_grid=param_grid, scoring=scoring,n_iter = n_iter, gridsearch=False)
rf_hyper=hypertuning_fxn(x, y, nfolds=nfolds, model=model , param_grid=param_grid, scoring=scoring,n_iter = n_iter, gridsearch=False)
# -
y
# +
# best cv score: 0.791
# best cv params: {'n_estimators': 50, 'min_samples_split': 10, 'min_samples_leaf': 5, 'max_features': 3, 'max_depth': 25, 'class_weight': {0: 0.000181422351233672, 1: 0.0005906674542232723}, 'bootstrap': False}
# #changing from niter=40 and bootstrap t/f:
# best cv score: 0.788
# best cv params: {'n_estimators': 250, 'min_samples_split': 2, 'min_samples_leaf': 2, 'max_features': 'auto', 'max_depth': 10, 'class_weight': None, 'bootstrap': True}
# -
rf_cv= hypertuned_cv_fxn(x, y, rf_hyper.best_estimator_, nfolds=nfolds)
# #### SVC hypertune
# +
model= svm.SVC(probability=True)
kernel = ['linear']#, 'rbf','poly']
#gamma = [0.1, 1, 'auto'] #Kernel coefficient for ‘rbf’, ‘poly’ and ‘sigmoid’. default=’auto’ uses 1 / n_features
C = [0.1, 1, 10, 100] #Penalty parameter C of the error term.
degree = [1,2]
class_weight=['balanced', None]
param_grid = {'kernel': kernel,
#'gamma': gamma,
'C': C,
'degree': degree,
'class_weight':class_weight}
svc_hyper=hypertuning_fxn(x, y, nfolds=nfolds, model=model , param_grid=param_grid,scoring=scoring, n_iter=5, gridsearch=False)
#svc_hyper=hypertuning_fxn(x, y, nfolds=4, model=model , param_grid=param_grid, base_model= base_model, scoring="neg_log_loss", n_iter=10, gridsearch=False)
# -
svc_cv= hypertuned_cv_fxn(x, y, svc_hyper.best_estimator_, nfolds=10)
# ### logistic regression hypertune
# +
###logreg
model= LogisticRegression(solver='lbfgs',random_state=12345)
#model = LogisticRegression(penalty='l2', dual=False, tol=0.0001, C=10, fit_intercept=True, intercept_scaling=1, class_weight='balanced', random_state=None)
penalty = ['l1','l2']
class_weight=['balanced',None]
param_grid = {'penalty': penalty,
'class_weight': class_weight}
logreg_hyper=hypertuning_fxn(x, y, nfolds=nfolds, model=model , param_grid=param_grid, scoring=scoring, n_iter=n_iter, gridsearch=True)
# -
logreg_cv= hypertuned_cv_fxn(x, y, logreg_hyper.best_estimator_, nfolds=10)
# ### KNN hypertune
# +
from sklearn.neighbors import KNeighborsClassifier
model= KNeighborsClassifier()
n_neighbors = [3,4,5, 8, 10, 25]
weights=['uniform']
p=[1,2] #1= mmanhattan, 2= euclidian
param_grid = {'n_neighbors': n_neighbors,
'weights': weights,
'p': p}
knn_hyper=hypertuning_fxn(x, y, nfolds=nfolds, model=model , param_grid=param_grid, scoring=scoring, n_iter=n_iter, gridsearch=True)
# -
knn_cv= hypertuned_cv_fxn(x, y, knn_hyper.best_estimator_, nfolds=10)
# ### Multilayer Perceptron (MLP) hypertune
# +
### MLP w/ standard scalar and early stopping (improved cv performance)
from sklearn.preprocessing import StandardScaler # doctest: +SKIP
# scaler = StandardScaler() # doctest: +SKIP
# scaler.fit(x) # doctest: +SKIP
# x_scalar = scaler.transform(x) # doctest: +SKIP
from sklearn.neural_network import MLPClassifier
#hidden_layer_sizes = [(50,), (100,), (150,), (250,)] #origional used parameters #(50,50), (100,100), (150,150),(300,)
hidden_layer_sizes =[(50), (50,50), (50,50,50), (50,100,50), (100,),(150)]
solver= ['sgd', 'adam']
# 'alpha': [0.0001, 0.05],
# 'learning_rate': ['constant','adaptive'],
activation= ['relu','tanh']#, 'logistic']
alpha= [0.001, 0.05] #L2 penalization degree #[0.001, 0.01]
learning_rate= ['adaptive']
learning_rate_init= [0.001, 0.01]
param_grid = {'hidden_layer_sizes': hidden_layer_sizes,
'solver':solver,
'activation':activation,
'alpha': alpha,
'learning_rate': learning_rate,
'learning_rate_init': learning_rate_init
}
model= MLPClassifier(early_stopping=True, random_state=12345)
#removed teh x_scalar
mlp_hyper=hypertuning_fxn(x, y, nfolds=nfolds, model=model , param_grid=param_grid, scoring=scoring,n_iter = 80, gridsearch=True) #gridsearch=False: testing with smaller, randomized grid
#gridsearch=False: testing with smaller, randomized grid. went from ~30 sec to 13min when ==True
# adding in solver: sgd and adam 1.5min ->8min
# -
mlp_cv= hypertuned_cv_fxn(x, y, mlp_hyper.best_estimator_, nfolds=10)
# ## idea: Rashamon effect: does ensembling models improve prediction performance?
# the ensemble is intresting if the ensemble predicts a different group of patients (or has different variable importance).
#
# rashamon effect where we don't gain from ensemble: identifying same patints in different ways
# rashamon effect where we do gain from ensemble: identifying DIFFERENT patients in the different ways
# +
# to test this, i first need to instantiate my other models, then ensemble them together.
# -
def reset_model(model_name, hardcode=False):
global xgboost_hyper, logreg_hyper, rf_hyper, knn_hyper, mlp_hyper, svc_hyper
if model_name== 'xgboost':
model = xgboost_hyper.best_estimator_
elif model_name== 'logreg':
model = logreg_hyper.best_estimator_
elif model_name== 'rf':
model = rf_hyper.best_estimator_
elif model_name== 'svc':
model = svc_hyper.best_estimator_
elif model_name== 'knn':
model = knn_hyper.best_estimator_
elif model_name== 'mlp':
model = mlp_hyper.best_estimator_
return(model)
# # test entire trainset and predict testset.
# *<del> step1) hypertune xgb via 10fold cv.
# * step2) test entire trainset and predict testset.
# * step3) generate figures
def get_auc_score(model,train_index, x=x,y=y):
y_pred_proba = model.predict_proba(x[train_index])[:, 1]
roc_score=roc_auc_score(y[train_index], y_pred_proba)
return(roc_score)
# # Ensemble CV and testing models
# ## test entire trainset and predict trainset.
# <del> * step1) hypertune xgb on 10fold cv.
#
# <del> * step2) test entire train set and predict testset.
# * step3) generate figures
#
#setting up test table
test_summary_df= pd.DataFrame({'model':[],'auc':[], 'precision':[], 'recall':[], 'f1':[], 'npv':[]})
test_summary_df
# ### model fitting
# +
xgboost = reset_model('xgboost')
xgboost.fit(x, y)
logreg = reset_model('logreg')
logreg.fit(x, y)
rf= reset_model('rf')
rf.fit(x,y)
svc= reset_model('svc')
svc.fit(x, y)
knn= reset_model('knn')
knn.fit(x,y)
mlp= reset_model('mlp')
mlp.fit(x,y)
from sklearn.ensemble import VotingClassifier
#create a dictionary of our models
estimators=[("xgboost", xgboost), ('rf', rf), ('log_reg', logreg), ('mlp',mlp), ('svc',svc)]
#create our voting classifier, inputting our models
ensemble = VotingClassifier(estimators, voting='soft', n_jobs=-1)
ensemble.fit(x,y)
# If ‘hard’, uses predicted class labels for majority rule voting.
# Else if ‘soft’, predicts the class label based on the argmax of the sums of the predicted probabilities,
# which is recommended for an ensemble of well-calibrated classifiers.
# from sklearn.naive_bayes import GaussianNB
# gnb =GaussianNB()
# nb_y_pred = gnb.fit(x, y)
# -
# ### wrapping up ensemble and CV
#running the ensemble in cv
ensemble_cv= hypertuned_cv_fxn(x, y, ensemble, nfolds=nfolds)
# +
#updating the cv table with ensemble data
cv_summary_df= pd.DataFrame([rf_cv,
logreg_cv,
xgboost_cv,
svc_cv,
knn_cv,
mlp_cv,
ensemble_cv])
cv_summary_df= cv_summary_df.set_index('model').round(decimals=3).sort_values('auc', ascending=False)
if save_boolean==True:
save_df(cv_summary_df, df_name='default', rel_path='/tables/')
cv_summary_df
# -
# ## global model test set evaluation
xgboost_eval= classifier_eval(xgboost, x=np.array(x_test), y=y_test, model_name='xgboost', folder_name=folder)
rf_eval= classifier_eval(rf, x=np.array(x_test), y=y_test, model_name='rf', folder_name=folder)
mlp_eval= classifier_eval(mlp, x=np.array(x_test), y=y_test, model_name='mlp', folder_name=folder)
svc_eval= classifier_eval(svc, x=np.array(x_test), y=y_test, model_name='svc', folder_name=folder)
# logreg_eval= classifier_eval(logreg, x=np.array(x_te
st), y=y_test)
logreg_eval= classifier_eval(logreg, x=np.array(x_test), y=y_test, model_name='logreg', folder_name=folder)
knn_eval= classifier_eval(knn, x=np.array(x_test), y=y_test, model_name='knn', folder_name=folder)
ensemble_eval= classifier_eval(ensemble,
x=np.array(x_test),
y=y_test,
model_name='ensemble',
folder_name=folder, save=True)
# looks like its the rashamon effect that doesn't benefit from ensembling.
#
# ie there are hidden variables that can discriminate the patients.
# # classification performance table
test_summary_df= pd.DataFrame([xgboost_eval, rf_eval, logreg_eval, mlp_eval, svc_eval, knn_eval, ensemble_eval])
test_summary_df=test_summary_df.set_index('model').round(decimals=3).sort_values('auc', ascending=False)
if save_boolean==True:
save_df(test_summary_df, df_name='default', rel_path='/tables/')
print(test_summary_df)
# # QC: checking performance on training set for xgboost and rf
xgboost_train= classifier_eval(xgboost, x=np.array(x_train), y=y_train, model_name='xgboost_train', folder_name=folder, save=False)
rf_train= classifier_eval(rf, x=np.array(x_train), y=y_train, model_name='rf_train', folder_name=folder, save=False)
# # publication figures
# making a model dictionary for ease of use in plotting
keys= ['rf', 'xgboost', 'logreg', 'mlp','svc', 'knn', 'ensemble']
model_list=[rf, xgboost, logreg, mlp, svc, knn, ensemble]
models_dic={}
for i in range(0,len(model_list)):
models_dic.update({keys[i] : model_list[i]} )
# ## stacked roc curve
def roc_publishing(model, x, y, proba_input=False,pos_label=1, print_default=True, model_name=None):
import sklearn.metrics as metrics
from sklearn.metrics import precision_score, roc_auc_score, f1_score, recall_score
model_name=type(model).__name__
y_proba = model.predict_proba(x)[:,1]
fpr, tpr, thresholds = metrics.roc_curve(y, y_proba, pos_label=pos_label)
roc_auc = metrics.auc(fpr, tpr)
#gathering the optimal youden_index and df of tpr/fpr for auc and index of that optimal youden. idx is needed in the roc
youden_threshold, roc_df, idx= optimal_youden_index(fpr, tpr, thresholds, tp90=True)
return(fpr, tpr, roc_auc, roc_df, idx)
# +
def stacked_roc(x_test, y_test, models_dic, first_bold=True):
"""
plotting function to plot a stacked ROC based on models in a dictionary.
first_bold=True means that the first model in the dic will stand out and be a solid line, while others are dotted
"""
global save_boolean
plt.style.use('seaborn-white')
plt.rcParams['figure.figsize'] = [7, 4]
if first_bold==True: i=0
else: i=1
# #####
# y_proba = model.predict_proba(x)[:,1]
# fpr, tpr, thresholds = metrics.roc_curve(y, y_proba, pos_label=pos_label)
# roc_auc = metrics.auc(fpr, tpr)
# #gathering the optimal youden_index and df of tpr/fpr for auc and index of that optimal youden. idx is needed in the roc
# youden_threshold, roc_df, idx= optimal_youden_index(fpr, tpr, thresholds,tp90=True)
# #plotting roc
# plot_roc(fpr, tpr, roc_auc, roc_df, idx, save=save, model_name=model_name,folder_name=folder)
# ######
for model_name in models_dic.keys():
#ax4= plt.plot(fpr, tpr, label = '%s AUC = %0.3f' % (model_name, roc_auc), linestyle='dotted')
if i==0:
model=models_dic[model_name]
fpr, tpr, roc_auc, roc_df, idx= roc_publishing(model, x=np.array(x_test), y=y_test, model_name=model_name)
print(model_name, roc_auc)
ax1= plt.plot(fpr, tpr, 'b', label = '%s AUC = %0.3f' % (model_name, roc_auc), linewidth=2)
og_idx=roc_df.iloc[(roc_df['thresholds']-0.5).abs().argsort()[:1]].index[0]
# plt.plot(roc_df.iloc[og_idx,1], roc_df.iloc[og_idx,2],marker='o', markersize=8, color="black")
# plt.plot(roc_df.iloc[idx,1], roc_df.iloc[idx,2],marker='o', markersize=6, color='r') ##
else:
model=models_dic[model_name]
fpr, tpr, roc_auc, roc_df, idx= roc_publishing(model, x=np.array(x_test), y=y_test, model_name=model_name)
print(model_name, roc_auc)
ax1= plt.plot(fpr, tpr, label = '%s AUC = %0.3f' % (model_name, roc_auc), linestyle='dotted')
og_idx=roc_df.iloc[(roc_df['thresholds']-0.5).abs().argsort()[:1]].index[0]
# plt.plot(roc_df.iloc[og_idx,1], roc_df.iloc[og_idx,2],marker='o', markersize=8, color="black")
# plt.plot(roc_df.iloc[idx,1], roc_df.iloc[idx,2],marker='o', markersize=6, color='r') ##
i+=1
###annotating the plot
plt.legend(loc = 'lower right')
# plt.annotate(s="P(0.5)",xy=(0.71, 0.50),color='black', size=10) #textcoords #alt: xy=(0.78, 0.345)
# plt.plot(0.68, 0.51, 'ro', color='black') #alt: (0.73, 0.36, 'ro', color='black')
# plt.annotate(s="P(tuned)",xy=(0.71, 0.56),color='black', size=10) #textcoords #alt: xy=(0.78, 0.405)
# plt.plot(0.68, 0.57, 'ro', color='r') #alt: (0.73, 0.42, 'ro', color='r')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate', size=14)
plt.xlabel('False Positive Rate', size=14)
if save_boolean==True:
saveplot(plt,'stacked_roc')
else: pass
plt.show()
# -
models_dic.keys()
stacked_roc(x_test, y_test, models_dic, first_bold=False)
# #
# # variable importance
# * first: find the top N (default=10) most important variables for each model
# * second: make a set of all unique variables present in the combined list of all models top N variables = varimp_set
# * find the variable importances for each model for varimp_set variables
# * scale the variable importance for each model relative to the MOST IMPORTANT VARIABLE:
# * ie: var1= 50 (max); var2= 25; var3= 5 --> varimp: var1=100, var2=50, var3=10
# * plot these as a line plot in descending order for the best performing model
from parameters import n_varimp
def find_N_varimp_set(x_train, models_dic):
"""
function that takes in a dictionary of models and the x_train dataframe and returns the set of variables present in the combined list of each model's top N most important variables.
1) find top N variables for each model
2) make list of all models top N
3) filter to only unique values in list = varimp_names
"""
global n_varimp
features_dic={}
top_set_dic={}
for model_name in models_dic.keys():
model= models_dic[model_name]
print(model_name)
if model_name in ['knn','ensemble', 'mlp']:
pass
elif model_name in ['logreg','svc']:
feature_importance = abs(model.coef_[0])
sorted_idx = np.argsort(feature_importance)[-n_varimp:]#[0]
features =list(np.array(x_train.columns)[sorted_idx][-n_varimp:])
features_dic.update( {model_name :features } )
else:
feat_importances = pd.Series(model.feature_importances_, index=x_train.columns)
features=feat_importances.nlargest(n_varimp).sort_values()
features=list(features.reset_index()['index'])
features_dic.update( {model_name :features } )
#######
set_features=[]
for features in features_dic.values():
set_features=set_features+features
set_features=set(set_features)
varimp_names=list(set_features)
return(varimp_names)
def topN_rel_imp(models_dic, varimp_names):
"""
input:dictionary of models and the top N set of important variables among models
output: relative variable importance for each model of all set(varimp_names) variables.
note: relative variable importance determined by dividing each variable importance by the value of the most important variable. this makes all values a comparison to the most important varaible:
ie 50 rel variable importance = half as important as the most important variable
"""
# finding the index of the set(varimp_names) in the dataframe.
#getting index of the set(top10) variables in x_train
xtrain_column_index_list=[]
for element in varimp_names:
variable_index=list(x_train).index(element)
xtrain_column_index_list.append(variable_index)
top_set_dic={} #instantiating dictionary
for model_name in models_dic.keys(): ##now that we have set of top N variables for each model. we can make relative importance for all unique variables in the set
model= models_dic[model_name]
if model_name in ['knn','ensemble', 'mlp']:
pass
elif model_name in ['logreg','svc']:
imp= abs(model.coef_[0])[xtrain_column_index_list]
rel_imp=100.0 * (imp / imp.max())
features =list(np.array(x_train.columns)[xtrain_column_index_list])#[-n_varimp:])
top_set= pd.Series(rel_imp,features).sort_values()
top_set_dic.update( {model_name :top_set } )
else:
imp=pd.Series(models_dic[model_name].feature_importances_, index=x_train.columns)[xtrain_column_index_list]
imp=imp.sort_values()
rel_imp=100.0 * (imp / imp.max())
features =list(np.array(x_train.columns)[xtrain_column_index_list])
top_set= rel_imp
top_set_dic.update( {model_name :top_set } )
return(top_set_dic)
def roc_name_adjust(varimp_names):
"""
cleans up the column names for the variable importance plot for publishing
"""
adjusted_names=[]
mapper={'vent_recieved_2.0': 'mechanical ventilation recieved',
'vent_recieved_1.0': 'oxygen ventilation recieved',
'vent_recieved_1.0': 'no ventilation recieved',
'pao2fio2ratio':'PaO2:FiO2',
# 'pao2fio2ratio_(475, 3000]': 'PaO2:FiO2 (475-3000)',
# 'pao2fio2ratio_(200, 333]': 'PaO2:FiO2 (200-333)',
# 'pao2fio2ratio_(333, 475]': 'PaO2:FiO2 (333-475)',
'ipco2_>50': 'pCO2 (>50)',
'ibands_>10': 'bands (>10)',
'ibands_absent': 'bands (missing)'}
for element in varimp_names:
if element in mapper.keys():
element= mapper[element]
adjusted_names.append(element)
elif "_1.0" in element:
element= element.strip("_1.0") + ' (Y/N)'
adjusted_names.append(element)
else:
adjusted_names.append(element)
return(adjusted_names)
def plot_topN_rel_imp(top_set_dic, varimp_names, xvar_rotation=80):
"""
plot the variable importance plots as a lineplot
rotation: the amount of xvar rotation
"""
global save_boolean
df_base=pd.DataFrame(index=varimp_names)
for model_name in top_set_dic.keys():
df_base[model_name]= top_set_dic[model_name]
adjusted_names=roc_name_adjust(varimp_names)
df_base.index=adjusted_names
df_base.sort_values('rf', ascending=False)
plt.style.use('seaborn-ticks')
plt.rcParams['figure.figsize'] = [10,10]#[7, 7]
plt.plot(df_base.sort_values('rf', ascending=True))
#plt.set_xticklabels(adjusted_names,rotation=30)
plt.xticks(rotation=xvar_rotation)#, ha='right')
plt.ylabel("Relative Variable Importance")
plt.legend(list(df_base))
if save_boolean==True:
saveplot(plt,'variable_importance')
return(df_base)
# +
#find set(topN) variables
varimp_names= find_N_varimp_set(x_train, models_dic)
#find rel importance of set(topN) variables for each model
top_set_dic= topN_rel_imp(models_dic, varimp_names)
#plot the variable importance plot
topN_rel_imp_df= plot_topN_rel_imp(top_set_dic, varimp_names, xvar_rotation=80)
# -
all_features=[
'bilirubin','bun','chloride',
'creatinine','glucose','heartrate',
'inr','lactate','potassium',
'ptt','resprate','sum_elix',
'temperature','bicarbonate','diasbp',
'hemoglobin','meanartpress','mingcs',
'pao2fio2ratio','ph','platelet',
'spo2','sysbp','maxCalcium',
'maxSodium','maxWBC','minCalcium',
'minSodium','minWBC','weight',
'yearsold','ethnicity_black','ethnicity_hispanic',
'ethnicity_unknown/other','ethnicity_white/nonhispanic','bands_>10',
'bands_absent','pco2_>50','pco2_absent',
'any_vasoactives_recieved','leukocyte_pos','nitrite_pos',
'vent_recieved_None','vent_recieved_Oxygen','dobutamine_recieved',
'gender_M','gender_NA','dopamine_recieved',
'epinephrine_recieved','norepinephrine_recieved','phenylephrine_recieved',
'rrt_recieved','vasopressin_recieved','cancer_elix_no_cancer_cormorb'
]
# # Model Saving
def model_save(model, model_name):
import pickle
modelpath=str(repository_path)+'/models/{}_{}'.format(date,folder)
if not os.path.exists(modelpath):
print(modelpath)
os.makedirs(modelpath)
filename = str(modelpath)+'/finalized_{}.sav'.format(model_name)
pickle.dump(model, open(filename, 'wb'))
model_save(xgboost,'xgboost')
model_save(rf,'rf')
model_save(logreg,'logreg')
model_save(svc,'svc')
model_save(knn,'knn')
model_save(mlp,'mlp')
model_save(ensemble,'ensemble')
folder
date
# cool model debugging code, useful to make validation curves across hyperparameters:
# +
# def val_curve_plot(train_scoreNum, test_scoreNum, param_range, param_name, rename_x_tick=False, param_range_name=['0','1']):
# """quick function to plot a validation curve across different hyperparameters"""
# plt.rcParams['figure.figsize'] = [7, 4]
# train_scores_mean = np.mean(train_scoreNum, axis=1)
# train_scores_std = np.std(train_scoreNum, axis=1)
# test_scores_mean = np.mean(test_scoreNum, axis=1)
# test_scores_std = np.std(test_scoreNum, axis=1)
# plt.title("Validation Curve with RF and parameter:{}".format(param_name))
# plt.xlabel(r"$\gamma$")
# plt.ylabel("Score")
# plt.ylim(0.6, 1.1)
# try:
# plt.xlim(0, max(param_range))
# except TypeError:
# pass
# lw = 2
# if rename_x_tick==False:
# param_range_name=param_range
# plt.errorbar(param_range_name,train_scores_mean, yerr=train_scores_std, alpha=0.6, color='darkorange', lw=lw, label='train')
# plt.errorbar(param_range_name,test_scores_mean, yerr=test_scores_std, alpha=0.6, color='blue', lw=lw, label='test')
# plt.legend(loc='best')
# plt.show()
# +
# from sklearn.model_selection import validation_curve
# param_range=[2, 10, 15, 25, 35, 50, 200]#[100, 300, 500, 750, 800, 1200]
# train_scoreNum, test_scoreNum = validation_curve(
# RandomForestClassifier(),
# X = x, y = y,
# param_name = 'n_estimators',
# param_range = param_range, cv = 5)
# val_curve_plot(train_scoreNum, test_scoreNum, param_range, param_name='n_estimators')
# +
#list(x_train2.loc[:,[x for x in list(x_train2) if x not in categorical]])
# -
# # exploring LIME visualization package (depreciated at current point)
# +
# x_train2=x_train.copy()
# +
# # trying to get LIME package working
# import lime
# import lime.lime_tabular
# +
# explainer = lime.lime_tabular.LimeTabularExplainer(np.array(x_train), feature_names=list(x_train), class_names=[0,1], discretize_continuous=True)
# explainer = lime.lime_tabular.LimeTabularExplainer(train ,feature_names = feature_names,class_names=class_names,
# categorical_features=categorical_features,
# categorical_names=categorical_names, kernel_width=3
# +
# i = np.random.randint(0, x_test.shape[0])
# exp = explainer.explain_instance(np.array(x_test)[i], rf.predict_proba, num_features=len(list(x_train)), top_labels=1)
# exp.show_in_notebook(show_table=True, show_all=False)
# +
# y_test[i]
# +
# np.array(x_test)[i]
# +
# x_test.iloc[i,:5]
# +
# exp = explainer.explain_instance(np.array(x_test)[i], xgboost.predict_proba, num_features=len(list(x_train)), top_labels=1)
# exp.show_in_notebook(show_table=True, show_all=False)
# -
# Now, there is a lot going on here. First, note that the row we are explained is displayed on the right side, in table format. Since we had the show_all parameter set to false, only the features used in the explanation are displayed.
#
# The value column displays the original value for each feature.
#
# Note that LIME has discretized the features in the explanation. This is because we let discretize_continuous=True in the constructor (this is the default). Discretized features make for more intuitive explanations.
folder
| notebooks/outcomes_and_alternative_calcs/07-modeling-save_before_change.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# 
#
# ---
# # Read CSV File and Put in Pandas Data Frame
#
# **Author list:** <NAME>
#
# **References / Sources:**
#
#
# **License Agreement:** Feel free to do whatever you want with this code
#
# ___
# +
# -*- coding: utf-8 -*-
"""
Created on Wed Oct 14 03:32:39 2015
@author: ikhlaqsidhu
"""
# General syntax to import specific functions in a library:
##from (library) import (specific library function)
from pandas import DataFrame, read_csv
# General syntax to import a library but no functions:
##import (library) as (give the library a nickname/alias)
import matplotlib.pyplot as plt
import pandas as pd #this is how I usually import pandas
import sys #only needed to determine Python version number
# from StringIO import StringIO
# import pandas
df0 = pd.read_csv('d3.txt')
print df0
# -
| code_blocks/CSV-Read-to-Pandas/Read-CSV-to-Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Info on Assignments and Final Project
#
# Below is a little video that introduces the assignments and the final project.
#
# <mark style="background-color: lightcoral">**Note** We are going to make some changes to the final project this year, so have a look at that part of the video just to get some overview of how it will be. Because of this, you won't find its description on GitHub just yet. </mark>
# Video 2: The assignments
from IPython.display import YouTubeVideo
YouTubeVideo("sSQBcbQ3FL0",width=800, height=450)
| lectures/Assignments_And_Final_Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Check Resume
# +
import io
from pdfminer.converter import TextConverter
from pdfminer.pdfinterp import PDFPageInterpreter
from pdfminer.pdfinterp import PDFResourceManager
from pdfminer.pdfpage import PDFPage
# Docx resume
import docx2txt
# Wordcloud
import re
import operator
from nltk.tokenize import word_tokenize
from nltk.corpus import stopwords
set(stopwords.words('english'))
from wordcloud import WordCloud
from nltk.probability import FreqDist
import matplotlib.pyplot as plt
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics.pairwise import cosine_similarity
# -
with open("../static/data/keywords.json") as content:
terms = json.load(content)
def read_pdf_resume(pdf_doc):
resource_manager = PDFResourceManager()
fake_file_handle = io.StringIO()
converter = TextConverter(resource_manager, fake_file_handle)
page_interpreter = PDFPageInterpreter(resource_manager, converter)
with open(pdf_doc, 'rb') as fh:
for page in PDFPage.get_pages(fh, caching=True,check_extractable=True):
page_interpreter.process_page(page)
text = fake_file_handle.getvalue()
# close open handles
converter.close()
fake_file_handle.close()
if text:
return text
def read_word_resume(word_doc):
resume = docx2txt.process(word_doc)
resume = str(resume)
#print(resume)
text = ''.join(resume)
text = text.replace("\n", "")
if text:
return text
def clean_job_decsription(jd):
"""A function to create a word cloud based on the input text parameter.
"""
## Clean the Text
# Lower
clean_jd = jd.lower()
# remove punctuation
clean_jd = re.sub(r'[^\w\s]', '', clean_jd)
# remove trailing spaces
clean_jd = clean_jd.strip()
# remove numbers
clean_jd = re.sub('[0-9]+', '', clean_jd)
# tokenize
clean_jd = word_tokenize(clean_jd)
# remove stop words
stop = stopwords.words('english')
clean_jd = [w for w in clean_jd if not w in stop]
return(clean_jd)
def create_word_cloud(jd):
corpus = jd
fdist = FreqDist(corpus)
#print(fdist.most_common(100))
words = ' '.join(corpus)
words = words.split()
# create a empty dictionary
data = dict()
# Get frequency for each words where word is the key and the count is the value
for word in (words):
word = word.lower()
data[word] = data.get(word, 0) + 1
# Sort the dictionary in reverse order to print first the most used terms
dict(sorted(data.items(), key=operator.itemgetter(1),reverse=True))
word_cloud = WordCloud(width = 800, height = 800,
background_color ='white',max_words = 500)
word_cloud.generate_from_frequencies(data)
# plot the WordCloud image
plt.figure(figsize = (10, 8), edgecolor = 'k')
plt.imshow(word_cloud,interpolation = 'bilinear')
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show()
def get_resume_score(text):
cv = CountVectorizer(stop_words='english')
count_matrix = cv.fit_transform(text)
#Print the similarity scores
print("\nSimilarity Scores:")
#get the match percentage
matchPercentage = cosine_similarity(count_matrix)[0][1] * 100
matchPercentage = round(matchPercentage, 2) # round to two decimal
print("Your resume matches about "+ str(matchPercentage)+ "% of the job description.")
if __name__ == '__main__':
extn = input("Enter File Extension: ")
file_path = input("Enter File Path: ")
if extn == "pdf":
resume = read_pdf_resume(file_path)
else:
resume = read_word_resume(file_path)
job_description = input("\nEnter the Job Description: ")
## Get a Keywords Cloud
clean_jd = clean_job_decsription(job_description)
create_word_cloud(clean_jd)
text = [resume, job_description]
## Get a Match score
get_resume_score(text)
read_pdf_resume("../static/data/temp resume.pdf")
| scripts/resume_checker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + id="7LYQx0iEvQoR"
class Human:
def __init__(self, name):
self.name = name
self.energy = 10
self.friends = []
def __str__(self):
s = 'Hi, my name is {0}'.format(self.name)
return s
def move(self, steps=1):
if self.energy < steps:
print("I don't have the energy to walk that far")
else:
self.energy -= steps
def sleep(self):
self.energy = 10
def makefriend(self, friend):
self.friends.append(friend)
friend.friends.append(self)
# + id="9WOs0gifx_Mh"
a = Human('Alice')
b = Human('Bruno')
c = Human('Chantel')
# + colab={"base_uri": "https://localhost:8080/"} id="VR4eCp91yXCS" outputId="e269f42b-1020-4225-bc00-03d9fd61fb01"
print(a)
a.move(3)
print(a.energy)
# + colab={"base_uri": "https://localhost:8080/"} id="o75Dp-Xfyj3w" outputId="c024aee9-21a8-43bc-918e-aea7fd85fa0a"
a.makefriend(b)
a.makefriend(c)
print("Alice's friends: ")
for i in a.friends:
print(i.name)
print("\nBruno's friends: ")
for i in b.friends:
print(i.name)
# + id="ncPHMHjqytyP"
class Bakery:
def __init__(self, cookies, cost):
self.numcookies = cookies
self.cost = cost
def __str__(self):
s = 'This Bakery has {0} cookies at ${1} each'.format(self.numcookies,
self.cost)
return s
# + id="r3APw78OsBeA"
class Baker(Human):
def __init__(self, name):
super().__init__(name)
self.sales = 0
def make_cookies(self, bakery, n_cookies):
bakery.numcookies += n_cookies
def sell_cookies(self, customer, bakery, n_cookies):
if n_cookies > bakery.numcookies:
print("This bakery doesn't have {0} cookies to sell".format(n_cookies))
return
subtotal = n_cookies * bakery.cost
if customer.money < subtotal:
print("The customer can't afford these cookies.")
return
bakery.numcookies -= n_cookies
self.sales += subtotal
customer.money -= subtotal
customer.cookies += n_cookies
# + id="RQiDoyfCvHbN"
class Customer(Human):
def __init__(self, name, money):
super().__init__(name)
self.money = money
self.cookies = 0
def __str__(self):
s = super().__str__() + ' and I have {0} cookies'.format(self.cookies)
return s
# + colab={"base_uri": "https://localhost:8080/"} id="8GPXKfn1yOll" outputId="70d8fca8-5249-4a29-8551-c31f0ced8d77"
romina = Baker('Romina')
print(romina)
# + colab={"base_uri": "https://localhost:8080/"} id="aPM1XpuhyYUh" outputId="c8f4791b-fa31-4970-825b-5195d372818b"
kelsey = Customer('Kelsey', 10)
print(kelsey)
# + colab={"base_uri": "https://localhost:8080/"} id="6YGVzjflyfdu" outputId="d9dbb6cb-79d1-4bde-b97b-5f05815722b5"
my_bakery = Bakery(20, 1.25)
print(my_bakery)
# + colab={"base_uri": "https://localhost:8080/"} id="-dRiCA035pAn" outputId="785995d4-6b7f-4100-f58b-b48c4934def0"
romina.sell_cookies(kelsey, my_bakery, 5)
print(kelsey)
print(kelsey.money)
print(my_bakery)
# + id="36jC2zU26FWN"
| book/tutorials/ObjectOriented/OOP_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import re
def print_separator():
print()
print('Prargraph Analysis')
print('--------------------')
def get_file_names(dir_name):
return os.listdir(dir_name)
def print_len_counts(Letter_count, word_len, sentence_len):
print("Average Letter Count: {0:.2f}".format(Letter_count/word_len))
print("Average Sentence Length: {0:.1f}".format(word_len/(sentence_len-1)))
def print_info_for_file_at_path(path):
textpath = os.path.join(foldername, current_file)
with open(textpath, newline='') as textfile:
paragraph = ''.join(textfile.readlines())
print_separator()
word = re.split("\s|(?<!\d)[.,][,.](?!\d)", paragraph)
print("Approximate Word Count: {0}".format(len(word)))
sentence = re.split(r'[.!?]+', paragraph)
print("Approximate Sentence Count: {0}".format(len(sentence)-1))
letter_lst = []
for char in list(paragraph):
if char.isalpha():
letter_lst.append(char)
print_len_counts(len(letter_lst), len(word), len(sentence))
print()
foldername = 'raw_data'
file_names = get_file_names(foldername)
for current_file in file_names:
print_info_for_file_at_path(current_file)
| PyParagraph/NotebookVersion_forfun.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 圖表,plot(),屬性kind有hist,kde等,偏斜程度skew(),2個屬性的相關性plot.scatter()
# ,聚合agg()傳入字典類型,分組groupby
# %%writefile loadData.py
import pandas as pd
import numpy as np
import os
def loadDate():
raw_data_path = os.path.join(os.path.pardir,'data','raw')
train_file_path = os.path.join(os.path.join(raw_data_path,'train.csv'))
test_file_path = os.path.join(os.path.join(raw_data_path,'test.csv'))
train_df = pd.read_csv(train_file_path,index_col='PassengerId')
test_df = pd.read_csv(test_file_path,index_col='PassengerId')
df= pd.concat((train_df,test_df),axis=0)
return df
df.Age.plot(kind='hist',title='histogram for Age',color='c')
#設置更多bucket
df.Age.plot(kind='hist',color='c',bins=20);
#kde
df.Age.plot(kind='kde',color='c');
df.Fare.plot(kind='hist',title='histogram for Fare',color='c')
#比較年齡和票價的偏斜函數
print('skewness for age: {0:.02f}'.format(df.Age.skew()))
print('skewness for fare: {0:.02f}'.format(df.Fare.skew()))
df.plot.scatter(x='Age',y='Fare',color='c')
#年齡和票價沒有明顯的相關性
df.plot.scatter(x='Age',y='Fare',color='c',alpha=0.1)
#客艙和票價的關係
df.plot.scatter(x='Pclass',y='Fare',alpha=0.15)
df.groupby('Sex').Age.median()
df.groupby('Pclass').Fare.median()
df.groupby(['Pclass'])['Fare','Age'].median()
#聚合,獲得票價的的平均值年齡的中位數,使用agg傳入字典
df.groupby('Pclass').agg({'Fare':'mean','Age':'median'})
aggre={
'Fare':{
'mean',
'median',
max,
np.max
},
'Age': {
'median',
min,
max,
lambda x:max(x)-min(x)
}
}
df.groupby('Pclass').agg(aggre)
#雙重分祖
df.groupby(['Pclass','Embarked']).Fare.median()
| notebooks/2020-05-02-GroupingAndAgg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# 
# ___
# - Dimensionality Reduction
# ## Principal component analysis (PCA)
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pylab as plt
import seaborn as sb
from IPython.display import Image
from IPython.core.display import HTML
from pylab import rcParams
import sklearn
from sklearn import decomposition
from sklearn.decomposition import PCA
from sklearn import datasets
# -
# %matplotlib inline
rcParams['figure.figsize'] = 5, 4
sb.set_style('whitegrid')
# ### PCA on the iris dataset
iris = datasets.load_iris()
X = iris.data
variable_names = iris.feature_names
X[0:10,]
# +
pca = decomposition.PCA()
iris_pca = pca.fit_transform(X)
pca.explained_variance_ratio_
# -
pca.explained_variance_ratio_.sum()
comps = pd.DataFrame(pca.components_, columns=variable_names)
comps
sb.heatmap(comps)
| heatmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# <a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fcurriculum-notebooks&branch=master&subPath=Mathematics/StatisticsProject/AccessingData/movie-theatre-releases.ipynb&depth=1" target="_parent"><img src="https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true" width="123" height="24" alt="Open in Callysto"/></a>
# # Movie Theatre Release Numbers
#
# We can get data about theatrical release gross profits from [Box Office Mojo by IMDBPro](https://www.boxofficemojo.com).
year = 2019
url = 'https://www.boxofficemojo.com/year/world/'+str(year)
import pandas as pd
box_office = pd.read_html(url)[0]
box_office
# [](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| _build/jupyter_execute/curriculum-notebooks/Mathematics/StatisticsProject/AccessingData/movie-theatre-releases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example used in the launch video
# +
# %load_ext autoreload
# %autoreload 2
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict, Headings
# +
design = designs.DesignPlanar()
gui = MetalGUI(design)
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
TransmonPocket.get_template_options(design)
# -
# ### Default settings for trace width and trace gap
design.variables['cpw_width'] = '15 um'
design.variables['cpw_gap'] = '9 um'
# ### 4 transmons (3 pins each) + 4 CPWs
# +
# Allow running the same cell here multiple times to overwrite changes
design.overwrite_enabled = True
## Custom options for all the transmons
options = dict(
# Some options we want to modify from the defaults
# (see below for defaults)
pad_width = '425 um',
pocket_height = '650um',
# Adding 4 connectors (see below for defaults)
connection_pads=dict(
a = dict(loc_W=+1,loc_H=-1, pad_width='200um'),
b = dict(loc_W=-1,loc_H=+1, pad_height='30um'),
c = dict(loc_W=-1,loc_H=-1, pad_height='50um')
)
)
## Create 4 transmons
q1 = TransmonPocket(design, 'Q1', options = dict(
pos_x='+2.42251mm', pos_y='+0.0mm', **options))
q2 = TransmonPocket(design, 'Q2', options = dict(
pos_x='+0.0mm', pos_y='-0.95mm', orientation = '270', **options))
q3 = TransmonPocket(design, 'Q3', options = dict(
pos_x='-2.42251mm', pos_y='+0.0mm', orientation = '180', **options))
q4 = TransmonPocket(design, 'Q4', options = dict(
pos_x='+0.0mm', pos_y='+0.95mm', orientation = '90', **options))
from qiskit_metal.qlibrary.tlines.meandered import RouteMeander
RouteMeander.get_template_options(design)
options = Dict(
lead=Dict(
start_straight='0.2mm',
end_straight='0.2mm'),
trace_gap='9um',
trace_width='15um')
def connect(component_name: str, component1: str, pin1: str, component2: str, pin2: str,
length: str, asymmetry='0 um', flip=False, fillet='90um'):
"""Connect two pins with a CPW."""
myoptions = Dict(
fillet=fillet,
pin_inputs=Dict(
start_pin=Dict(
component=component1,
pin=pin1),
end_pin=Dict(
component=component2,
pin=pin2)),
total_length=length)
myoptions.update(options)
myoptions.meander.asymmetry = asymmetry
myoptions.meander.lead_direction_inverted = 'true' if flip else 'false'
return RouteMeander(design, component_name, myoptions)
asym = 140
cpw1 = connect('cpw1', 'Q1', 'c', 'Q2', 'b', '5.6 mm', f'+{asym}um')
cpw2 = connect('cpw2', 'Q3', 'b', 'Q2', 'c', '5.7 mm', f'-{asym}um', flip=True)
cpw3 = connect('cpw3', 'Q3', 'c', 'Q4', 'b', '5.6 mm', f'+{asym}um')
cpw4 = connect('cpw4', 'Q1', 'b', 'Q4', 'c', '5.7 mm', f'-{asym}um', flip=True)
gui.rebuild()
gui.autoscale()
# -
# ### 4 Open to Ground Pins
# +
from qiskit_metal.qlibrary.terminations.open_to_ground import OpenToGround
otg1 = OpenToGround(design, 'OTG1', options = dict(pos_x='2771um', pos_y='0', orientation='180', gap='9um', width='15um'))
otg2 = OpenToGround(design, 'OTG2', options = dict(pos_x='0um', pos_y='-1301um', orientation='90', gap='9um', width='15um'))
otg3 = OpenToGround(design, 'OTG3', options = dict(pos_x='-2771um', pos_y='0', orientation='0', gap='9um', width='15um'))
otg4 = OpenToGround(design, 'OTG4', options = dict(pos_x='0um', pos_y='1301um', orientation='270', gap='9um', width='15um'))
# -
# ### V1 Launchers
# +
from qiskit_metal.qlibrary.terminations.launchpad_wb import LaunchpadWirebond
p1_c = LaunchpadWirebond(design, 'P1_C', options = dict(pos_x='4000um', pos_y='2812um', orientation='270', lead_length='0um', cpw_gap='9um', cpw_width='15um'))
p2_c = LaunchpadWirebond(design, 'P2_C', options = dict(pos_x='4000um', pos_y='-2812um', orientation='90', lead_length='0um', cpw_gap='9um', cpw_width='15um'))
p3_c = LaunchpadWirebond(design, 'P3_C', options = dict(pos_x='-4000um', pos_y='-2812um', orientation='90', lead_length='0um', cpw_gap='9um', cpw_width='15um'))
p4_c = LaunchpadWirebond(design, 'P4_C', options = dict(pos_x='-4000um', pos_y='2812um', orientation='270', lead_length='0um', cpw_gap='9um', cpw_width='15um'))
# -
#
# ### V2 Launchers
# +
from qiskit_metal.qlibrary.terminations.launchpad_wb_coupled import LaunchpadWirebondCoupled
p1_q = LaunchpadWirebondCoupled(design, 'P1_Q', options = dict(pos_x='4020um', pos_y='0', orientation='180', lead_length='0um', cpw_gap='9um', cpw_width='15um'))
p2_q = LaunchpadWirebondCoupled(design, 'P2_Q', options = dict(pos_x='-990um', pos_y='-2812um', orientation='90', lead_length='0um', cpw_gap='9um', cpw_width='15um'))
p3_q = LaunchpadWirebondCoupled(design, 'P3_Q', options = dict(pos_x='-4020um', pos_y='0', orientation='0', lead_length='0um', cpw_gap='9um', cpw_width='15um'))
p4_q = LaunchpadWirebondCoupled(design, 'P4_Q', options = dict(pos_x='990um', pos_y='2812um', orientation='270', lead_length='0um', cpw_gap='9um', cpw_width='15um'))
gui.rebuild()
gui.autoscale()
# -
# ### Charge Lines to Corners
# +
import numpy as np
from collections import OrderedDict
from qiskit_metal.qlibrary.tlines.anchored_path import RouteAnchors
anchors1c = OrderedDict()
anchors1c[0] = np.array([3, 0])
anchors1c[1] = np.array([3, 2.5])
anchors1c[2] = np.array([4, 2.5])
options_line_cl1 = {'pin_inputs':
{'start_pin': {'component': 'OTG1', 'pin': 'open'},
'end_pin': {'component': 'P1_C', 'pin': 'tie'}},
'leadin': {'start': '0um', 'end': '0um'},
'anchors': anchors1c,
'trace_gap': '9um',
'trace_width': '15um',
'fillet': '90um'
}
cl1 = RouteAnchors(design, 'line_cl1', options_line_cl1)
anchors2c = OrderedDict()
anchors2c[0] = np.array([0, -1.5])
anchors2c[1] = np.array([2, -1.5])
anchors2c[2] = np.array([2, -2.5])
anchors2c[3] = np.array([4, -2.5])
options_line_cl2 = {'pin_inputs':
{'start_pin': {'component': 'OTG2', 'pin': 'open'},
'end_pin': {'component': 'P2_C', 'pin': 'tie'}},
'leadin': {'start': '0um', 'end': '0um'},
'anchors': anchors2c,
'fillet': '90um',
'trace_gap': '9um',
'trace_width': '15um'
}
cl2 = RouteAnchors(design, 'line_cl2', options_line_cl2)
anchors3c = OrderedDict()
anchors3c[0] = np.array([-3, 0])
anchors3c[1] = np.array([-3, -2.5])
anchors3c[2] = np.array([-4, -2.5])
options_line_cl3 = {'pin_inputs':
{'start_pin': {'component': 'OTG3', 'pin': 'open'},
'end_pin': {'component': 'P3_C', 'pin': 'tie'}},
'leadin': {'start': '0um', 'end': '0um'},
'anchors': anchors3c,
'fillet': '90um',
'trace_gap': '9um',
'trace_width': '15um'
}
cl3 = RouteAnchors(design, 'line_cl3', options_line_cl3)
anchors4c = OrderedDict()
anchors4c[0] = np.array([0, 1.5])
anchors4c[1] = np.array([-2, 1.5])
anchors4c[2] = np.array([-2, 2.5])
anchors4c[3] = np.array([-4, 2.5])
options_line_cl4 = {'pin_inputs':
{'start_pin': {'component': 'OTG4', 'pin': 'open'},
'end_pin': {'component': 'P4_C', 'pin': 'tie'}},
'leadin': {'start': '0um', 'end': '0um'},
'anchors': anchors4c,
'fillet': '90um',
'trace_gap': '9um',
'trace_width': '15um'
}
cl4 = RouteAnchors(design, 'line_cl4', options_line_cl4)
gui.rebuild()
gui.autoscale()
# -
# ### Exchange Coupler Lines
# +
options = Dict(
lead=Dict(
start_straight='0.35mm',
end_straight='0.15mm'),
trace_gap='9um',
trace_width='15um')
ol1 = connect('ol1', 'Q1', 'a', 'P1_Q', 'tie', '5.5 mm', f'+{asym}um', flip=True)
ol2 = connect('ol2', 'Q2', 'a', 'P2_Q', 'tie', '13.0 mm', f'+{asym}um', flip=True)
ol3 = connect('ol3', 'Q3', 'a', 'P3_Q', 'tie', '5.5 mm', f'+{asym}um', flip=True)
ol4 = connect('ol4', 'Q4', 'a', 'P4_Q', 'tie', '13.0 mm', f'+{asym}um', flip=True)
gui.rebuild()
gui.autoscale()
# -
# ### Export to GDS
# +
#QDesign enables GDS renderer during init.
a_gds = design.renderers.gds
a_gds.options
# -
#If you have a fillet_value and there are LineSegments that are shorter than 2*fillet_value,
#When true, the short segments will not be fillet'd.
a_gds.options['short_segments_to_not_fillet'] = 'True'
SCALE_FILLET = 2.0
a_gds.options['check_short_segments_by_scaling_fillet'] = SCALE_FILLET
a_gds.options['path_filename'] = '../resources/Fake_Junctions.GDS'
# +
# Export GDS file for all components in design.
# Please change the path where you want to write a GDS file.
#def export_to_gds(self, file_name: str, highlight_qcomponents: list = []) -> int:
a_gds.export_to_gds("Launch_Notebook.gds")
# -
gui.main_window.close()
# # END of first video.
#
# # Below, other launch video. requires Ansys previous setup
# +
# %load_ext autoreload
# %autoreload 2
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict, Headings
# +
from qiskit_metal.renderers.renderer_ansys.ansys_renderer import QAnsysRenderer
QAnsysRenderer.default_options
# +
design = designs.DesignPlanar()
gui = MetalGUI(design)
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
# -
design.variables['cpw_width'] = '15 um'
design.variables['cpw_gap'] = '9 um'
# +
# Allow running the same cell here multiple times to overwrite changes
design.overwrite_enabled = True
## Custom options for all the transmons
options = dict(
# Some options we want to modify from the defaults
# (see below for defaults)
pad_width = '425 um',
pocket_height = '650um',
# Adding 4 connectors (see below for defaults)
connection_pads=dict(
a = dict(loc_W=+1,loc_H=-1, pad_width='200um'),
b = dict(loc_W=-1,loc_H=+1, pad_height='30um'),
c = dict(loc_W=-1,loc_H=-1, pad_height='50um')
)
)
## Create 4 transmons
q1 = TransmonPocket(design, 'Q1', options = dict(
pos_x='+2.42251mm', pos_y='+0.0mm', **options))
q2 = TransmonPocket(design, 'Q2', options = dict(
pos_x='+0.0mm', pos_y='-0.95mm', orientation = '270', **options))
q3 = TransmonPocket(design, 'Q3', options = dict(
pos_x='-2.42251mm', pos_y='+0.0mm', orientation = '180', **options))
q4 = TransmonPocket(design, 'Q4', options = dict(
pos_x='+0.0mm', pos_y='+0.95mm', orientation = '90', **options))
from qiskit_metal.qlibrary.tlines.meandered import RouteMeander
RouteMeander.get_template_options(design)
options = Dict(
lead=Dict(
start_straight='0.2mm',
end_straight='0.2mm'),
trace_gap='9um',
trace_width='15um')
def connect(component_name: str, component1: str, pin1: str, component2: str, pin2: str,
length: str, asymmetry='0 um', flip=False, fillet='90um'):
"""Connect two pins with a CPW."""
myoptions = Dict(
fillet=fillet,
pin_inputs=Dict(
start_pin=Dict(
component=component1,
pin=pin1),
end_pin=Dict(
component=component2,
pin=pin2)),
total_length=length)
myoptions.update(options)
myoptions.meander.asymmetry = asymmetry
myoptions.meander.lead_direction_inverted = 'true' if flip else 'false'
return RouteMeander(design, component_name, myoptions)
asym = 140
cpw1 = connect('cpw1', 'Q1', 'c', 'Q2', 'b', '5.6 mm', f'+{asym}um')
cpw2 = connect('cpw2', 'Q3', 'b', 'Q2', 'c', '5.7 mm', f'-{asym}um', flip=True)
cpw3 = connect('cpw3', 'Q3', 'c', 'Q4', 'b', '5.6 mm', f'+{asym}um')
cpw4 = connect('cpw4', 'Q1', 'b', 'Q4', 'c', '5.7 mm', f'-{asym}um', flip=True)
gui.rebuild()
gui.autoscale()
# -
fourq = design.renderers.hfss
# this will error out if Ansys is not configured correctly. Also you will need to open Ansys before connecting.
fourq.connect_ansys()
fourq.add_eigenmode_design("FourQ")
fourq.render_design([], [])
fourq.disconnect_ansys()
| tutorials/Appendix A Full design flow examples/Example used in the launch video.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from tensorflow import keras
print(tf.__version__)
print(keras.__version__)
fashion_mnist = keras.datasets.fashion_mnist
(X_train_full, y_train_full), (X_test, y_test) = fashion_mnist.load_data()
print(X_train_full.shape)
print(X_train_full.dtype)
print(y_train_full.shape)
print(y_train_full.dtype)
print(X_test.shape)
print(X_test.dtype)
import numpy as np
from pprint import pprint
# +
A_indices = np.where(y_train_full<8)
B_indices = np.where(y_train_full>=8)
X_train_full_A = X_train_full[A_indices]
X_train_full_B = X_train_full[B_indices]
y_train_full_A = y_train_full[A_indices]
y_train_full_B = y_train_full[B_indices]
print("size A = {}".format(len(y_train_full_A)))
print("size B = {}".format(len(y_train_full_B)))
# +
A_indices_test = np.where(y_test<8)
B_indices_test = np.where(y_test>=8)
y_test_A = y_test[A_indices_test]
y_test_B = y_test[B_indices_test]
y_test_B = y_test_B - 8
X_test_A = X_test[A_indices_test] / 255.0
X_test_B = X_test[B_indices_test] / 255.0
pprint(y_test[:100])
pprint("len y_test_A = {}".format(len(y_test_A)))
pprint("len y_test_B = {}".format(len(y_test_B)))
pprint(y_test_A[:100])
pprint(y_test_B[:100])
# +
X_valid_A, X_train_A = X_train_full_A[:4000] / 255.0, X_train_full_A[4000:] / 255.0
y_valid_A, y_train_A = y_train_full_A[:4000], y_train_full_A[4000:]
X_valid_B, X_train_B = X_train_full_B[:1000] / 255.0, X_train_full_B[1000:] / 255.0
y_valid_B, y_train_B = y_train_full_B[:1000], y_train_full_B[1000:]
y_train_B = y_train_B - 8
y_valid_B = y_valid_B - 8
pprint(y_train_B[:100])
pprint(y_valid_B[:100])
# -
class_names_A = ["T-shirt/top", "Trouser", "Pullover", "Dress", "Coat",
"Sandal", "Shirt", "Sneaker"]
class_names_B = ["Bag", "Ankle boot"]
model_A = keras.models.Sequential()
model_A.add(keras.layers.Flatten(input_shape=[28, 28]))
model_A.add(keras.layers.Dense(300, activation="relu"))
model_A.add(keras.layers.Dense(100, activation="relu"))
model_A.add(keras.layers.Dense(8, activation="softmax"))
print(model_A.summary())
model_A.compile(loss="sparse_categorical_crossentropy", optimizer="Adam", metrics=["accuracy"])
history = model_A.fit(X_train_A, y_train_A, epochs=30, validation_data=(X_valid_A, y_valid_A))
model_A.evaluate(X_test_A, y_test_A)
# +
model_A_clone = keras.models.clone_model(model_A)
model_A_clone.set_weights(model_A.get_weights())
model_B_on_A = keras.models.Sequential(model_A.layers[:-1])
model_B_on_A.add(keras.layers.Dense(1, activation='sigmoid'))
for layer in model_B_on_A.layers[:-1]:
layer.trainable = False
model_B_on_A.compile(loss="binary_crossentropy", optimizer="Adam", metrics=["accuracy"])
# +
history = model_B_on_A.fit(X_train_B, y_train_B, epochs=4, validation_data=(X_valid_B, y_valid_B))
for layer in model_B_on_A.layers[:-1]:
layer.trainable = True
optimizer = keras.optimizers.Adam(lr=1e-4)
model_B_on_A.compile(loss='binary_crossentropy', optimizer=optimizer, metrics=["accuracy"])
history = model_B_on_A.fit(X_train_B, y_train_B, epochs=16, validation_data=(X_valid_B, y_valid_B))
# -
pprint(y_valid_B[:100])
| chapter11/transfer_learning_example.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# +
# 9.3 가솔린 소비량: 가솔린 자동차의 연비를 결정하는 요인들을 연구하기 위하여 30개 자동차 모형으로부터 데이터를 수집하였다.
# 반응변수 Y는 1갈론당 주행거리(마일)을 나타내는 가솔린 연료소모량이고, 이에 영향을 끼치는 11개 예측변수들은 자동차의 물리적인, 기계적인 성격과 관련된 것들이다.
# 표 9.16에 사용된 변수의 내역과 데이터가 주어져 있다. 표 9.17의 원데이터는 1975년도 자동차 관련 잡지인 Motor Trend로부터 얻어진 것이다.
#여기에서는 데이터에 공선성이 존재하는지를 파악하고자 한다.
setwd('H:/01.Korea/01.2019-1학기/05.회귀분석 방법론/Report/Chapter9')
p256_data <- read.table("P256.txt", header = TRUE)
str(p256_data)
# +
# Y : 주행거리 (마일/갈론)
# X1 : 배기량 (입방 인치)
# X2 : 마력 (피트/파운드)
# X3 : 염력 (피트/파운드)
# X4 : 압축비
# X5 : 후진축비
# X6 : 내연 기화기 (배럴)
# X7 : 변속기수
# X8 : 전체길이 (인치)
# X9 : 폭 (인치)
# X10 : 무게 (파운드)
# X11 : 변속기형태(1=자동식, 0=수동식)
# +
# (a) 예측변수 X1, ..., X11 의 상관계수 행렬을 계산하고, 대응하는 짝별 산점도를 그려라. 공선성의 증거를 찾아보아라.
# -
cor(p256_data)
p256_cor = cor(p256_data[,2:12]) #상관계수행렬
panel.cor <- function(x,y,digits=2, prefix="", cex.cor,...)
{
usr <- par("usr"); on.exit(par(usr))
par(usr=c(0,1,0,1))
r <- abs(cor(x,y))
txt <- format(c(r,0.123456789),digits=digits)[1]
txt <- paste0(prefix,txt)
if(missing(cex.cor)) cex.cor <- 1.5/strwidth(txt)
text(0.5,0.5,txt, cex=cex.cor*r)
}
pairs(p256_data ,
lower.panel=function(x,y){
points(x,y);
abline(0, 1, col='red')
},
upper.panel = panel.cor
)
#install.packages("corrplot")
library(corrplot)
corrplot(p256_cor
, method="number" # 색 입힌 사각형
, addshade="all" )# 상관관계 방향선 제시)
# (b) 상관계수로부터 고유값, 고유벡터, 상태수를 계산하여라. 이 데이터에는 다중공선성이 존해 하는가?
p256_cor_eigen <- eigen(p256_cor)$values
cindex_256 <- sqrt(p256_cor_eigen[1]/p256_cor_eigen);
p256_cor_eigen
cindex_256
| 1st semester/01.LinearRegression/Chapter9/HW(9.3).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# +
from io import BytesIO
import time
import sys
import json
import boto3
import PIL.Image
import sagemaker
import numpy
import gym
from gym import wrappers
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
# %matplotlib inline
from importlib import reload
from IPython import display
import ipywidgets as widgets
from IPython.display import display as i_display
from heuristics_utils import simulate
from battlesnake_gym.snake_gym import BattlesnakeGym
# -
# # Define the openAI gym
# Optionally, you can define the initial game state (the situation simulator) of the snakes and food.
# To use the initial state, set `USE_INITIAL_STATE = True` and enter the desired coordinates of the snake and food using the initial_state dictionary. The dictionary follows the same format as the battlesnake API.
# +
USE_INITIAL_STATE = False
# Sample initial state for the situation simulator
initial_state = {
"turn": 4,
"board": {
"height": 11,
"width": 11,
"food": [
{
"x": 1,
"y": 3
}
],
"snakes": [{
"health": 90,
"body": [{"x": 3, "y": 0}],
},
{
"health": 90,
"body": [{"x": 6, "y": 0}],
},
{
"health": 90,
"body": [{"x": 2, "y": 5}],
},
{
"health": 90,
"body": [{"x": 6, "y": 4}],
},
{
"health": 90,
"body": [{"x": 7, "y": 3}],
},
]
}
}
if USE_INITIAL_STATE == False:
initial_state = None
# -
# The parameters here must match the ones provided during training (except initial_state)
map_size = (11, 11)
number_of_snakes = 5
random_snake = False # If this is set to true, the simulator will not use the network but just output random actions
env = BattlesnakeGym(map_size=map_size, number_of_snakes=number_of_snakes, observation_type="max-bordered-51s",
initial_game_state=initial_state)
# # Load the trained model
# Load the RLlib models.
# %cd inference
# !mkdir output
# !tar -C output -xvf model.tar.gz
# %cd ..
model_filepath = "inference/output/1/"
if random_snake:
net = None
else:
imported = tf.saved_model.load(model_filepath)
net = imported.signatures["serving_default"]
# Clean up the model
# !rm -r inference/output
# # Simulation loop
#
# Run a simulation with the environment with the heuritics that you wrote.
# To edit the heuristics, edit the file `RLlibEnv/inference/src/battlesnake_heuristics`.
# Note that you can track the progress of your work with git.
# +
import inference.inference_src.battlesnake_heuristics
reload(inference.inference_src.battlesnake_heuristics)
from inference.inference_src.battlesnake_heuristics import MyBattlesnakeHeuristics
heuristics = MyBattlesnakeHeuristics()
infos, rgb_arrays, actions, heuristics_remarks, json_array = simulate(env, net, heuristics, number_of_snakes, random_snake)
# -
# # Playback the simulation
#
# Defines the user interface of the simulator.
# +
def get_env_json():
if slider.value < len(json_array):
return json_array[slider.value]
else:
return ""
def play_simulation(_):
for i in range(slider.value, len(rgb_arrays) - slider.value - 1):
slider.value = slider.value + 1
display_image(slider.value)
time.sleep(0.2)
def on_left_button_pressed(_):
if slider.value > 0:
slider.value = slider.value - 1
display_image(slider.value)
def on_right_button_pressed(_):
if slider.value < len(rgb_arrays):
slider.value = slider.value + 1
display_image(slider.value)
def display_image(index):
if index >= len(rgb_arrays):
return
info = infos[index]
action = actions[index]
heuristics = heuristics_remarks[index]
snake_colours = env.snakes.get_snake_colours()
line_0 = [widgets.Label("Turn count".format(info["current_turn"])),
widgets.Label("Snake")]
line_1 = [widgets.Label(""), widgets.Label("Health")]
line_2 = [widgets.Label("{}".format(info["current_turn"])),
widgets.Label("Action")]
line_3 = [widgets.Label(""), widgets.Label("Gym remarks")]
line_4 = [widgets.Label(""), widgets.Label("Heur. remarks")]
action_convertion_dict = {0: "Up", 1: "Down", 2: "Left", 3: "Right", 4: "None"}
for snake_id in range(number_of_snakes):
snake_health = "{}".format(info["snake_health"][snake_id])
snake_health_widget = widgets.Label(snake_health)
snake_action = "{}".format(action_convertion_dict[action[snake_id]])
snake_action_widget = widgets.Label(snake_action)
snake_colour = snake_colours[snake_id]
hex_colour = '#%02x%02x%02x' % (snake_colour[0], snake_colour[1], snake_colour[2])
snake_colour_widget = widgets.HTML(value = f"<b><font color="+hex_colour+">⬤</b>")
gym_remarks = ""
if snake_id in info["snake_info"]:
if info["snake_info"][snake_id] != "Did not colide":
gym_remarks = "{}".format(info["snake_info"][snake_id])
gym_remarks_widget = widgets.Label(gym_remarks)
heuris_remarks = "{}".format(heuristics[snake_id])
heuris_remarks_widget = widgets.Label(heuris_remarks)
line_0.append(snake_colour_widget)
line_1.append(snake_health_widget)
line_2.append(snake_action_widget)
line_3.append(gym_remarks_widget)
line_4.append(heuris_remarks_widget)
line_0_widget = widgets.VBox(line_0)
line_1_widget= widgets.VBox(line_1)
line_2_widget = widgets.VBox(line_2)
line_3_widget = widgets.VBox(line_3)
line_4_widget = widgets.VBox(line_4)
info_widget = widgets.HBox([line_0_widget, line_1_widget, line_2_widget, line_3_widget, line_4_widget])
image = PIL.Image.fromarray(rgb_arrays[index])
f = BytesIO()
image.save(f, "png")
states_widget = widgets.Image(value=f.getvalue(), width=500)
main_widgets_list = [states_widget, info_widget]
main_widget = widgets.HBox(main_widgets_list)
display.clear_output(wait=True)
i_display(navigator)
i_display(main_widget)
left_button = widgets.Button(description='◄')
left_button.on_click(on_left_button_pressed)
right_button = widgets.Button(description='►')
right_button.on_click(on_right_button_pressed)
slider = widgets.IntSlider(max=len(rgb_arrays) - 1)
play_button = widgets.Button(description='Play')
play_button.on_click(play_simulation)
navigator = widgets.HBox([left_button, right_button, slider, play_button])
display_image(index=0)
# -
# To get a JSON representation of the gym (environment), run the following function. You can also use output of the following function as an initial_state of the gym.
#
# *Please provide this json array if you are reporting bugs in the gym*
get_env_json()
# # Deploy the SageMaker endpoint
# This section will deploy your new heuristics into the SageMaker endpoint
# +
sage_session = sagemaker.session.Session()
s3_bucket = "SAGEMAKER_S3_BUCKET"
role = sagemaker.get_execution_role()
endpoint_instance_type = "SAGEMAKER_INFERENCE_INSTANCE_TYPE"
print("Your sagemaker s3_bucket is s3://{}".format(s3_bucket))
model_data = "s3://{}/battlesnake-aws/pretrainedmodels/model.tar.gz".format(s3_bucket)
# -
# ## Deploy your new heuristics
# Using the new heuristics you developed, a new SageMaker endpoint will be created.
#
# Firstly, delete the old endpoint, model and endpoint config.
sm_client = boto3.client(service_name='sagemaker')
sm_client.delete_endpoint(EndpointName='battlesnake-endpoint')
sm_client.delete_endpoint_config(EndpointConfigName='battlesnake-endpoint')
sm_client.delete_model(ModelName="battlesnake-rllib")
# Run the following cells to create a new model and endpoint with the new heuristics
# +
from sagemaker.tensorflow.serving import Model
model = Model(model_data=model_data,
role=role,
entry_point="inference.py",
source_dir='inference/inference_src',
framework_version='2.1.0',
name="battlesnake-rllib",
code_location='s3://{}//battlesnake-aws/code'.format(s3_bucket)
)
# Deploy an inference endpoint
predictor = model.deploy(initial_instance_count=1, instance_type=endpoint_instance_type, endpoint_name='battlesnake-endpoint')
# -
# ## Testing the new endpoint
# You should see `Action to take is X`
# +
import numpy as np
from time import time
state = np.zeros(shape=(1, 21, 21, 6), dtype=np.float32).tolist()
health_dict = {0: 50, 1: 50}
json = {"turn": 4,
"board": {
"height": 11,
"width": 11,
"food": [],
"snakes": []
},
"you": {
"id": "snake-id-string",
"name": "<NAME>",
"health": 90,
"body": [{"x": 1, "y": 3}]
}
}
before = time()
action_mask = np.array([1, 1, 1, 1]).tolist()
action = predictor.predict({"state": state, "action_mask": action_mask,
"prev_action": -1,
"prev_reward": -1, "seq_lens": -1,
"all_health": health_dict, "json": json})
elapsed = time() - before
action_to_take = action["outputs"]["heuristisc_action"]
print("Action to take {}".format(action_to_take))
print("Inference took %.2f ms" % (elapsed*1000))
| RLlibEnv/HeuristicsDeveloper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://landlab.github.io"><img style="float: left" src="../../landlab_header.png"></a>
# # Introduction to the FlowAccumulator
#
# Landlab directs flow and accumulates it using two types of components:
#
# **FlowDirectors** use the topography to determine how flow moves between adjacent nodes. For every node in the grid it determines the nodes to receive flow and the proportion of flow to send from one node to its receiver.
#
# The **FlowAccumulator** uses the direction and proportion of flow moving between each node and (optionally) water runoff to calculate drainage area and discharge.
#
# In this tutorial we will go over how to initialize and run the **FlowAccumulator**. For tutorials on how to initialize and run a **FlowDirector** and a brief comparison between the different flow direction algorithms or for more detailed examples that contrast the differences between each flow direction algorithm, refer to the other tutorials in this section.
#
# First, we import the necessary python modules and make a small plotting routine.
# %matplotlib inline
# +
# import plotting tools
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import matplotlib as mpl
# import numpy
import numpy as np
# import necessary landlab components
from landlab import RasterModelGrid, HexModelGrid
from landlab.components import FlowAccumulator
from landlab.components import (FlowDirectorD8, FlowDirectorDINF,
FlowDirectorMFD, FlowDirectorSteepest)
from landlab.components import DepressionFinderAndRouter
# import landlab plotting functionality
from landlab.plot.drainage_plot import drainage_plot
# create a plotting routine to make a 3d plot of our surface.
def surf_plot(mg,
surface='topographic__elevation',
title='Surface plot of topography'):
fig = plt.figure()
ax = fig.gca(projection='3d')
# Plot the surface.
Z = mg.at_node[surface].reshape(mg.shape)
color = cm.gray((Z - Z.min()) / (Z.max() - Z.min()))
surf = ax.plot_surface(mg.x_of_node.reshape(mg.shape),
mg.y_of_node.reshape(mg.shape),
Z,
rstride=1,
cstride=1,
facecolors=color,
linewidth=0.,
antialiased=False)
ax.view_init(elev=35, azim=-120)
ax.set_xlabel('X axis')
ax.set_ylabel('Y axis')
ax.set_zlabel('Elevation')
plt.title(title)
plt.show()
# -
# ## Topographic grids
#
# For this tutorial we will consider one topographic surface. Here it is plotted in three dimensions.
mg = RasterModelGrid((10, 10))
_ = mg.add_field('topographic__elevation',
3. * mg.x_of_node**2 + mg.y_of_node**2,
at='node')
surf_plot(mg, title='Grid 1')
# ## Initalizing and running the FlowAccumulator
#
# To instantiate the **FlowAccumulator**, you must pass it the minimum of a model grid that has a field called ```'topographic__elevation'```.
#
# Alternatively, you can pass it the name of another field name at node, or an array with length number of nodes. This is the surface over which flow is first directed and then accumulated.
#
# **FlowAccumulator** will create and use a **FlowDirector** to calculate flow directions. The default **FlowDirector** is **FlowDirectorSteepest**, which is the same as D4 in the special case of a raster grid. There are a few different ways to specify which **FlowDirector** you want **FlowAccumulator** to use. The next section will go over these options.
#
# **FlowAccumulator** can take a constant or spatially variable input called ```runoff_rate```, which it uses to calculate discharge. Alternatively, if there is an `at_node` field called ```water__unit_flux_in``` and no value is specified as the ```runoff_rate```, **FlowAccumulator** will use the values stored in ```water__unit_flux_in```.
#
# In addition to directing flow and accumulating it in one step, **FlowAccumulator** can also deal with depression finding internally. This can be done by passing a **DepressionFinder** to the keyword argument ```depression_finder```. The default behavior is to not deal with depressions internally.
#
# Finally, if the **FlowDirector** you are using takes any keyword arguments, those can be passed to the **FlowAccumulator**. For example, **FlowDirectorMFD** has to option to use diagonals in addition to links and to proportion flow based on either the slope or the the square root of slope.
# +
fa = FlowAccumulator(mg)
# this is the same as writing:
fa = FlowAccumulator(mg,
surface='topographic__elevation',
flow_director='FlowDirectorSteepest',
runoff_rate=None,
depression_finder=None)
# -
# The **FlowAccumulator** has two public methods: ```run_one_step()``` and ```accumulate_flow()```.
#
# Both use the values of the surface provided to identify flow directions (and in the case of directing to more than one receiver, proportions) and then calculate discharge and drainage area. Both store the same information about receivers, proportions, and other calculated values to the model grid as fields. The difference is that ```run_one_step()``` does not return any values, while ```accumulate_flow()``` returns the drainage area and discharge as variables.
# +
fa.run_one_step()
(da, q) = fa.accumulate_flow()
# -
# We can illustrate the receiver node **FlowDirectionSteepest** has assigned to each donor node using a plotting function in Landlab called ```drainage_plot```. We will see many of these plots in this tutorial so let's take a moment to walk through the plot and what it contains.
#
# - The background image (white to black) shows the values of topographic elevation of the underlying surface or any other at_node field we choose to plot.
#
# - The colors of the dots inside of each pixel show the locations of the nodes and the type of node.
#
# - The arrows show the direction of flow, and the color shows the proportion of flow that travels along that link.
#
# - An X on top of a node indicates that node is a local sink and flows to itself.
#
# Note that in Landlab *Boundary Nodes*, or nodes that are on the edge of a grid, do not have area and do not contribute flow to nodes. These nodes can either be *Fixed Gradient Nodes*, *Fixed Value Nodes*, or *Closed Nodes*. With the exception of *Closed Nodes* the boundary nodes can receive flow.
#
# An important step in all flow direction and accumulation is setting the proper boundary condition. Refer to the boundary condition tutorials for more information.
plt.figure()
drainage_plot(mg)
# In this drainage plot, we can see that all of the flow is routed down the steepest link. A plot of the drainage area would illustrate how the flow would move. Next let's make a similar plot except that instead of plotting the topographic elevation as the background, we will plot the drainage area.
plt.figure()
drainage_plot(mg, 'drainage_area')
# If we print out the drainage area, we can see that its maximum reaches 64, which is the total area of the interior of the grid.
print(mg.at_node['drainage_area'].reshape(mg.shape))
# This is the same number as the number of core nodes. This makes sense becaue these are the only nodes in Landlab that have area, and in our model grid they each have an area of one.
print(mg.number_of_core_nodes)
# We can rain on the surface, store that rain in the field ```water__unit_flux_in```, and then re-run the **FlowAccumulator**. As an example, we will 'rain' a uniformly distributed random number between 0 and 1 on every node.
#
# Since we already ran the **FlowAccumulator**, under the hood our grid already has a field called ```water__unit_flux_in``` and we need to set the ```clobber``` keyword to ```True```.
# +
rain = 1. + 5. * np.random.rand(mg.number_of_nodes)
plt.imshow(rain.reshape(mg.shape), origin='lower', cmap='PuBu', vmin=0)
plt.colorbar()
plt.show()
_ = mg.add_field('water__unit_flux_in', rain, at='node', clobber=True)
# -
# Next, we re-run the **FlowAccumulator** and plot the discharge.
fa.run_one_step()
plt.figure()
drainage_plot(mg, 'surface_water__discharge', title='Discharge')
# The basic pattern of drainage is the same but the values for the surface water discharge are different than for drainage area.
# ### Alternative ways to specify the FlowDirector
#
# **FlowAccumulator** allows the **FlowDirector** to be specified one of four ways:
# 1. As a string of the full name of the **FlowDirector** (e.g., ```'FlowDirectorSteepest'``` or ```'FlowDirectorD8'``` )
# 2. As a string of the short name of the **FlowDirector** method (e.g., ```'Steepest'``` or ```'D8'```)
# 3. As the class name for the desired **FlowDirector** component.
# 4. As an instantiated version of a **FlowDirector** component.
#
# Thus, the following four ways to instantiate a **FlowAccumulator** are equivalent.
# +
# option 1: Full name of FlowDirector
fa = FlowAccumulator(mg,
surface='topographic__elevation',
flow_director='FlowDirectorSteepest')
# option 2: Short name of FlowDirector
fa = FlowAccumulator(mg,
surface='topographic__elevation',
flow_director='Steepest')
# option 3: Uninstantiated FlowDirector Component
fa = FlowAccumulator(mg,
surface='topographic__elevation',
flow_director=FlowDirectorSteepest)
# option 4: Instantiated FlowDirector Component
fd = FlowDirectorSteepest(mg)
fa = FlowAccumulator(mg, surface='topographic__elevation', flow_director=fd)
# -
# ### Providing a DepressionFinder
#
# Just as with providing the **FlowDirector**, the **DepressionFinder** can be provided multiple ways. While there are presently four different **FlowDirectors** in Landlab, there is only one **DepressionFinder**.
#
# 1. As a string of the full name of the **DepressionFinder** (e.g., ```'DepressionFinderAndRouter'```)
# 2. As the class name of the **DepressionFinder** component.
# 3. As an instantiated version of a **DepressionFinder** component.
#
# NOTE: The current Landlab depression finder only works with **FlowDirectorSteepest** and **FlowDirectorD8** no matter how the depression finder is run. This is because the depression finder presently only works with route-to-one methods.
#
# Thus, the following three ways to instantiated a **DepressionFinder** are equivalent.
# +
# option 1: Full name of FlowDirector
fa = FlowAccumulator(mg,
surface='topographic__elevation',
flow_director='FlowDirectorD8',
depression_finder='DepressionFinderAndRouter')
# option 2: Uninstantiated FlowDirector Component
fa = FlowAccumulator(mg,
surface='topographic__elevation',
flow_director=FlowDirectorD8,
depression_finder='DepressionFinderAndRouter')
# option 3: Instantiated FlowDirector Component
fd = FlowDirectorD8(mg)
df = DepressionFinderAndRouter(mg)
fa = FlowAccumulator(mg,
surface='topographic__elevation',
flow_director=fd,
depression_finder=df)
# -
# Methods for specifying can be mixed, such that the following is permissible.
df = DepressionFinderAndRouter(mg)
fa = FlowAccumulator(mg,
surface='topographic__elevation',
flow_director='D8',
depression_finder=df)
# ### Using the DepressionFinder with FlowAccumulator
#
# To conclude this tutorial, we examine an example of a Hexagonal Model grid with a depression.
# +
hmg = HexModelGrid((9, 5))
_ = hmg.add_field('topographic__elevation',
hmg.x_of_node + hmg.y_of_node,
at='node')
fa = FlowAccumulator(hmg, flow_director='MFD')
fa.run_one_step()
plt.figure()
drainage_plot(hmg)
# -
plt.figure()
drainage_plot(hmg, 'drainage_area')
# We will put a depression in the middle of the topography, and then see what the drainage plot looks like.
# +
hmg_hole = HexModelGrid((9, 5))
z = hmg_hole.add_field('topographic__elevation',
hmg_hole.x_of_node + np.round(hmg_hole.y_of_node),
at='node')
hole_nodes = [21, 22, 23, 30, 31, 39, 40]
z[hole_nodes] = z[hole_nodes] * 0.1
fa = FlowAccumulator(hmg_hole, flow_director='Steepest')
fa.run_one_step()
plt.figure()
drainage_plot(hmg_hole)
# -
plt.figure()
drainage_plot(hmg_hole, 'drainage_area')
# As you can see, the flow gets stuck in the hole. We'd like the flow in the hole to move out and to the boundary.
#
# To route the flow out of the hole, we have two options.
# 1. Run the **FlowAccumulator** and then the **DepressionFinder**
# 2. Run them together in **FlowAccumulator**.
#
# The options look like the following and they are equivalent.
# +
# OPTION 1
fa = FlowAccumulator(hmg_hole, flow_director='Steepest')
fa.run_one_step()
df = DepressionFinderAndRouter(hmg_hole)
df.map_depressions()
# OPTION 2
fa = FlowAccumulator(hmg_hole,
flow_director='Steepest',
depression_finder='DepressionFinderAndRouter')
fa.run_one_step()
plt.figure()
drainage_plot(hmg_hole, 'drainage_area')
# -
# As you can see the flow is now routed out of the hole and down to a boundary.
#
# # Conclusion
#
# This tutorial went over how to run the **FlowAccumulator**. To learn more, consider one of two additional tutorials about directing and accumulating flow in Landlab:
#
# 1. **Introduction to FlowDirector**: A tutorial that goes over the different **FlowDirectors** present in Landlab and how to create and run a **FlowDirector**.
#
# 2. **Comparison of FlowDirectors**: A tutorial that constrasts the different methods in more detail and over surfaces that are more complicated than a simple sloping ramp.
#
#
| notebooks/tutorials/flow_direction_and_accumulation/the_FlowAccumulator.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// + [markdown] dotnet_interactive={"language": "csharp"}
// # Моделируем кубернетес кластер
// -
// > Этот ноутбук и соответствующую модель можно пощупать в [](https://mybinder.org/v2/gh/eosfor/scripting-notes/HEAD)
// ## Задача
//
// Довольно часто, при проектировании инфраструктуры под Kubernetes (AKS) в Microsoft Azure, у заказчиков возникают странные вопросы: какого размера будет все это добро, или, сколько все это будет нам стоить? Для того чтобы хоть как-то ответить на этот вопрос надо предположить какого размера виртуальные машины выбрать, сколько их должно быть, сколько они стоят в нужном регионе. Размер машин, в свою очередь, зависит от количества и прожорливости приложений, а также от количества экземпляров каждого из них. И так далее. А ведь нам хочется еще и подобрать все это по минимальной цене. В общем и целом, входных переменных довольно много и свести их в кучу, да еще и для каждого размера виртуальных машин отдельно, скажем прямо - не самая интересная работа в мире.
//
// В этом эксперименте мы попробуем построить модель, которая сможет оценить необходимый размер кластера, исходя из количества и размеров приложений. А также будет стараться уменьшить его стоимость.
// ## Формальные требования
//
// Начнем с виртуальных машин. Первое и, наверное, самое важное - ресурсов машины, должно хватить на все запущенные экземпляры приложения. Другими словами, в сумме, приложения должны сожрать меньше, чем доступно на ноде. Кроме того, здесь стоит учесть нагрузку, которую создает сам Kubernetes. Для этого имеет смысл зарезервировать некий процент ресурсов, и не отдавать его приложениям. С другой стороны, чтобы ресурсы не простаивали, нужно иметь некую нижнюю планку потребления ресурсов. Так же, было бы хорошо, чтобы суммарно, все приложения влезали в кластер, а для этого модель должна определять необходимое количество нод, держа в уме нагрузку на приложения.
// Переходим к приложениям. Положим, каждое приложение должно исполняться как минимум в двух экземплярах, на случай отказа одного из них. При этом эти экземпляры должны исполняться на разных нодах, по той же причине. Плюс к этому, было бы очень неплохо, если бы модель умела оценить количество нод, в зависимости от предполагаемой нагрузки на приложение.
// ## Построение модели
//
// В качестве инструмента для моделирования будем использовать minizinc.
// ### Моделирование приложений
//
// Начнем с описания приложений. Мы хотим запускать в кластере некое, заранее известное количество приложений, с известными максимальными значениями потребления ресурсов. Все эти значения - переменные, что дает нам возможность настраивать модель и менять ее поведение. Для моделирования "сложных" объектов в minizinc применяется следующий трюк. Задается некий ENUM тип, который используется как индекс в массивах со значениями атрибутов
//
// ```powershell
// enum appNames = {A, B, C, D, E};
// array[appNames] of int: appRAM = [2,4,6,8,4];
// array[appNames] of int: appCPU = [200,400,400,450,2250];
// array[appNames] of int: appConnectionsPerInstance = [5,2,2, 2,2];
// array[appNames] of int: appConnectionsFact = [20,4,3,4,10];
// ```
//
// Здесь `{A, B, C, D, E}` - имена приложений. Массивы `appRAM`, `appCPU`, `appConnectionsPerInstance`, `appConnectionsFact` объемы потребляемыых ресурсов: памяти, процессора, оценочое количество "подключений", которое может обслужить один экземпляр приложения и фактическое количество таких "подключений"
// ### Ноды кластера
//
// Нам нужно смоделировать объект ноды с его атрибутами, поэтому
//
// ```powershell
// enum nodeKinds = {D2sv5, D4sv5, D8sv5};
// array[nodeKinds] of int: clusterNodeRAM = [8,16,32];
// array[nodeKinds] of int: clusterNodeCPU = [2000,4000,8000];
// array[nodeKinds] of float: clusterNodePrice = [0.048, 0.096, 0.192];
// ```
//
// Тут я использую имена размеров виртуальных машин Microsoft Azure, их стоимость и параметры. Но в целом, все ровно то же самое, что и с приложениями
// ### Decision variables
//
// Стоит сделать отступление и упомянуть, что в minizinc есть два типа "переменных": обычные переменные, к которым мы все привыкли, и "decision variables". Последние представляют "решения", которые модель на minizinc должна принять, чтобы требования и ограничения модели были соблюдены. Иными словами, значения обычных переменных мы должны задать вручную, а значения "decision variables" minizinc должен найти сам. В этом, собственно, вся штука. Мы задаем ограничения, и просим модель подобрать такие значения для "decision variables", которые удовлетворяли бы всем требованиям
//
// В нашем случае подбиать подходящее количество нод в кластере мы будем руками, просто перезапуская модель. Поэтому просто зададим некий изначальный размер, который будет случить верхней границей размера.
//
// ```powershell
// int: maxClusterSize = 6;
// set of int: CLUSTER = 1..maxClusterSize;
// ```
//
// Теперь начинается самое интересное. На каждой ноде этого кластера мы хотим запусить одно или несколько приложений. Смоделируем это как некую матрицу, в которой измерениями будут ноды кластера и имена приложений. Другими словами мы построим матрицу А размера m x n, где строки m - означают ноды кластера, а столбцы n - приложения. Элемент матрицы a[i,j] - булевская "decision variable", которая показывает запущено ли на ноде i приложение j.
//
// ```powershell
// array[CLUSTER, appNames] of var bool: runningApps;
// ```
//
// В этом примере индексами массива служат списки `CLUSTER` и `appNames` а значениями - decision variables.
// ### Добавляем ограничения
//
// Мы хотим чтобы модель учитывала следующие основные требования:
//
// Суммарное потребление RAM всеми приложениями, исполняющимися на одной ноде должно лежать между некими минимальным и максимальным значениями. Максимальное значение задает ограничение сверху, например мы хотим зарезервировать некий процент памяти для самого кубернетиса. Нижняя граница в свою очередь требует, чтобы нода не была пустой.
//
// ```
// % sum of RAM of all apps on the node >=minNodeRAMConsumption and <= maxNodeRAMConsumption of total node RAM
// constraint forall (i in CLUSTER) (
// nodeRAMConsumption[i] >= (clusterNodeRAM[nodes[i]] * minNodeRAMConsumption) /\
// nodeRAMConsumption[i] <= (clusterNodeRAM[nodes[i]] * maxNodeRAMConsumption)
// );
// ```
//
// Ровно то же самое для CPU.
//
// ```
// % sum of CPU of all apps on the node >= minNodeCPUConsumption and <= maxNodeCPUConsumption of total node CPU
// constraint forall (i in CLUSTER) (
// nodeCPUConsumption[i] >= (clusterNodeCPU[nodes[i]] * minNodeCPUConsumption) /\
// nodeCPUConsumption[i] <= (clusterNodeCPU[nodes[i]] * maxNodeCPUConsumption)
// );
// ```
//
// Остался сущий пустяк, мы хотим, чтобы модель учитывала нагрузку на приложения. У нас есть максимальная нагрука на экземпляр - `appConnectionsPerInstance`, после которой необходимо добавить еще один, и текущая фактическая нагрузка - `appConnectionsFact`. На основе этих значений мы можем потребовать, чтобы модель посчитала количество экземпляров каждого приложения. И затем потребовать, чтобы суммарное количество экземпляров приложения в кластере совпадало с рассчетным.
//
// ```
// % assuming the load, calculate target number of pods
// array[appNames] of var int: appInstanceCount;
// constraint forall (j in appNames)(
// appInstanceCount[j] = ceil(appConnectionsFact[j]/appConnectionsPerInstance[j])
// );
//
// % assuming the load, use calculated number of pods
// constraint forall (j in appNames) (
// sum(i in CLUSTER) (runningApps[i,j]) = appInstanceCount[j]
// );
// ```
// ### Немного оптимизации
//
// Кроме всего прочего мы хотим, чтобы цена всего кластера была минимальной. Для этого мы задаем сумму как цель для оптимизации и просим модель минимизировать эту функцию
//
//
// ```
// var float: cost = (sum(i in nodes)(clusterNodePrice[i])) * 24 * 30 ; % target function to optimize
// solve minimize cost;
// ```
// ## Пощупаем модель
//
// Готовая модель хранится [вот тут](./kubernetes-model/model.mzn).
// Запустим ее. Для этого у вас должен быть установлен minizinc. Либо можно попробовать этот ноутбук в [](https://mybinder.org/v2/gh/eosfor/scripting-notes/HEAD). Контейнер с этим ноутбуком уже содержит minizinc.
// + dotnet_interactive={"language": "pwsh"}
minizinc ./kubernetes-model/model.mzn
// -
// Как это читать.
//
// ```
// D4sv5; CPU:2850/4000; RAM:12/16, A;C;E;
// ```
//
// - `D4sv5` размер виртуальной машины, которую выбрала модель
// - `CPU:2850/4000` текущая загрузка/максимальная загрузка ноды
// - `RAM:12/16` текущая загрузка/максимальная загрузка ноды
// - `A;C;E` приложения, которые модель расположила на этой ноде
//
// Кроме этого есть два специальных индикатора
//
// - `----------` так обозначаются решения, предлагаемые моделью. Их может быть больше одного
// - `==========` так отмечается оптимальное решение
//
// В нашем случае модель нашла оптимальное решение и показала его
//
| notebooks/ru/kubernetes-model-minizinc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python38-azureml
# kernelspec:
# display_name: Python 3.8 - AzureML
# language: python
# name: python38-azureml
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# # Hands on - Surfing Your Data using Azure SDK for Python
#
# __Notebook Version:__ 1.0<br>
# __Python Version:__ Python 3.8 - AzureML<br>
# __Required Packages:__ No<br>
# __Platforms Supported:__ Azure Machine Learning Notebooks
#
# __Data Source Required:__ Log Analytics tables
#
# ### Description
# This notebook will provide step-by-step instructions and sample code to guide you through Azure authentication, Sentinel log data discovery by using Azure SDK for Python and Kusto Query Language (KQL).<br>
# *** No need to download and install any other Python modules. ***<br>
# *** Please run the cells sequentially to avoid errors. *** <br>
# Need to know more about KQL? [Getting started with Kusto Query Language](https://docs.microsoft.com/en-us/azure/data-explorer/kusto/concepts/).
#
# ## Table of Contents
# 1. Warm-up
# 2. Azure Authentication
# 3. Log Analytics Data Queries
# 4. Bonus: Sentinel Watchlist Items Retrieval
# + [markdown] nteract={"transient": {"deleting": false}}
# ## 1. Warm-up
# + gather={"logged": 1605055050943} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# If you need to know what Python modules are available, you may run this:
# help("modules")
# + gather={"logged": 1627596157670} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Load Python libraries that will be used in this notebook
from azure.common.client_factory import get_client_from_cli_profile
from azure.common.credentials import get_azure_cli_credentials
from azure.loganalytics.models import QueryBody
from azure.mgmt.loganalytics import LogAnalyticsManagementClient
from azure.loganalytics import LogAnalyticsDataClient
from IPython.display import display, HTML, Markdown
import pandas as pd
import json
import ipywidgets
import matplotlib.pyplot as plt
# + gather={"logged": 1627596159719} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Functions will be used in this notebook
def read_config_values(file_path):
"This loads pre-generated parameters for Sentinel Workspace"
with open(file_path) as json_file:
if json_file:
json_config = json.load(json_file)
return (json_config["tenant_id"],
json_config["subscription_id"],
json_config["resource_group"],
json_config["workspace_id"],
json_config["workspace_name"],
json_config["user_alias"],
json_config["user_object_id"])
return None
def has_valid_token():
"Check to see if there is a valid AAD token"
try:
credentials, sub_id = get_azure_cli_credentials()
creds = credentials._get_cred(resource=None)
token = creds._token_retriever()[2]
print("Successfully signed in.")
return True
except Exception as ex:
if "Please run 'az login' to setup account" in str(ex):
print(str(ex))
return False
elif "AADSTS70043: The refresh token has expired" in str(ex):
message = "**The refresh token has expired. <br> Please continue your login process. Then: <br> 1. If you plan to run multiple notebooks on the same compute instance today, you may restart the compute instance by clicking 'Compute' on left menu, then select the instance, clicking 'Restart'; <br> 2. Otherwise, you may just restart the kernel from top menu. <br> Finally, close and re-load the notebook, then re-run cells one by one from the top.**"
display(Markdown(message))
return False
except:
print("Please restart the kernel, and run 'az login'.")
return False
def process_result(result):
"This function processes data returned from Azure LogAnalyticsDataClient, it returns pandas DataFrame."
json_result = result.as_dict()
cols = pd.json_normalize(json_result['tables'][0], 'columns')
final_result = pd.json_normalize(json_result['tables'][0], 'rows')
if final_result.shape[0] != 0:
final_result.columns = cols.name
return final_result
# + gather={"logged": 1627596164365} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Calling the above function to populate Sentinel workspace parameters
# The file, config.json, was generated by the system, however, you may modify the values, or manually set the variables
tenant_id, subscription_id, resource_group, workspace_id, workspace_name, user_alias, user_object_id = read_config_values('config.json');
# + [markdown] nteract={"transient": {"deleting": false}}
# ## 2. Azure Authentication
# + gather={"logged": 1627596167655}
# Azure CLI is used to get device code to login into Azure, you need to copy the code and open the DeviceLogin site.
# You may add [--tenant $tenant_id] to the command
if has_valid_token() == False:
# !az login --tenant $tenant_id --use-device-code
# Initialzie Azure LogAnalyticsDataClient, which is used to access Sentinel log data in Azure Log Analytics.
# You may need to change resource_uri for various cloud environments.
resource_uri = "https://api.loganalytics.io"
la_client = get_client_from_cli_profile(LogAnalyticsManagementClient, subscription_id = subscription_id)
creds, _ = get_azure_cli_credentials(resource=resource_uri)
la_data_client = LogAnalyticsDataClient(creds)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1627596170786}
# If you encounter error like: "got an unexpected keyword argument 'user_agent'" at the above cell, you may run the following command as a temporarily work-around to continue:
# Please uncomment the following line and run it:
# # !pip install --upgrade azure-cli
# Then re-run the cell above
# + [markdown] nteract={"transient": {"deleting": false}}
# ## 3. Log Analytics Data Queries
# + gather={"logged": 1627596175658} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Get all tables available using Kusto query language. If you need to know more about KQL, please check out the link provided at the introductory section.
tables_result = None
table_list = None
all_tables_query = "union withsource = SentinelTableName * | distinct SentinelTableName | sort by SentinelTableName asc"
if la_data_client != None:
tables_result = la_data_client.query(workspace_id, QueryBody(query=all_tables_query))
# + gather={"logged": 1627596178815} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Process the data using above function, then convert DataFrame to list
if tables_result != None:
table_list = process_result(tables_result)
tables = sorted(table_list.SentinelTableName.tolist())
table_dropdown = ipywidgets.Dropdown(options=tables, description='Tables:')
display(table_dropdown)
# + gather={"logged": 1627596186625} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# You may query the table based on your needs, here I use TimeGenerated column as an example, going back to 7 days, counting events per day
# Then process the data and display the result
# To look at the query, you may run: print(sample_query)
date_column_name = "TimeGenerated"
count_column_name = "Count"
if table_list.empty == False:
table_name = table_dropdown.value
sample_query = "{0} | where {1} >= ago(7d) | summarize {2}=count() by format_datetime({1}, 'yyyy-M-dd') | order by {1} asc".format(table_name, date_column_name, count_column_name)
print("Query:" + sample_query)
print("===================")
result_sample = la_data_client.query(workspace_id, QueryBody(query=sample_query))
sample_result = process_result(result_sample)
print(sample_result)
# + gather={"logged": 1627596191716} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Then plot a bar chart
if sample_result.empty == False:
plt.bar(sample_result[date_column_name], sample_result[count_column_name])
plt.rcParams['figure.figsize'] = [14,2.5]
# + [markdown] nteract={"transient": {"deleting": false}}
# ## 4. Bonus: Sentinel Watchlist Items Retrieval
#
# + gather={"logged": 1619562234520} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}} tags=["parameters"]
# Here we are going to use the Sentinel Watchlist name that you got in previous Hands-on notebook to get all Watchlist items
# First, please set the watchlist_name
watchlist_name = ipywidgets.Text(value='[[YOUR WATCHLIST NAME]]',description='watchlist_name: ')
display(watchlist_name)
# + gather={"logged": 1619562245589} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Using Aazure SDK for Python: LogAnalyticsDataClient to get items
watchlist_query = "_GetWatchlist('{0}')".format(watchlist_name.value)
result_watchlist = la_data_client.query(workspace_id, QueryBody(query=watchlist_query))
my_watchlist_items = process_result(result_watchlist)
print(my_watchlist_items)
| Hands-on 2. Surfing Data using Azure SDK.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit # import the curve fitting function
import pandas as pd
# %matplotlib inline
# ## Argon
# +
Argon = pd.read_table('Ar.txt',delimiter=', ',engine='python', header=None)
Amu = Argon[0] #These are the values of amu that the mass spec searches for
Argon = np.array([entry[:-1] for entry in Argon[1]],dtype='float')*1e6
# -
# ### Raw Argon Data
plt.figure(figsize=(9,4))
plt.scatter(Amu, Argon);
ax = plt.gca()
#ax.set_yscale('log')
plt.xlim(12,45);
plt.ylim(0,4)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.savefig('RawArgon.png')
np.argmax(Argon)
np.argmax(Argon[0:250])
Argon[191]
Argon[391]/Argon[191]
# ### Substract Argon Background
# +
Arbkd = pd.read_table('Background_Ar.txt',delimiter=', ',engine='python', header=None)
Arbkd = np.array([entry[:-1] for entry in Arbkd[1]],dtype='float')*1e6
# -
plt.figure(figsize=(9,4))
plt.scatter(Amu, Argon - Arbkd);
ax = plt.gca()
#ax.set_yscale('log')
plt.xlim(12,45);
plt.ylim(0,4)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.savefig('TrueArgon.png')
# Peak at 14 amu single Nitrogen, probably from Air. Peak at 20, 28, 32 are Neon, N2, and O2. 20 could be CH2 from methanol
# Peak at 40 is surely Argon atom
plt.figure(figsize=(9,4))
plt.scatter(Amu, Arbkd);
ax = plt.gca()
#ax.set_yscale('log')
plt.xlim(12,30);
plt.ylim(0,0.5)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.savefig('ArgonBackground.png')
# Background substraction has removed a single peak at 18 amu and 17 amu, which is the water peak and OH+ fragment.
# ### Argon Background
# ## Kyrpton
# +
Krypton = pd.read_table('Kr.txt',delimiter=', ',engine='python', header=None)
Krypton = np.array([entry[:-1] for entry in Krypton[1]],dtype='float')*1e6
Krbkd = pd.read_table('Background_Kr.txt',delimiter=', ',engine='python', header=None)
Krbkd = np.array([entry[:-1] for entry in Krbkd[1]],dtype='float')*1e6
# -
plt.figure(figsize=(9,4))
plt.scatter(Amu, Krypton - Krbkd);
ax = plt.gca()
plt.xlim(12,85);
plt.ylim(0,6)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.savefig('Krypton.png')
# Here, and for all subsequent measurements on this day, there is a slight peak at 40 amu, which is to be some residual from the Argon test.
# Too heavy to be measured, possible for it to have shown near 40.
# ## Neon
# +
Neon = pd.read_table('Ne.txt',delimiter=', ',engine='python', header=None)
Neon = np.array([entry[:-1] for entry in Neon[1]],dtype='float')*1e6
Nebkd = pd.read_table('Background_Ne.txt',delimiter=', ',engine='python', header=None)
Nebkd = np.array([entry[:-1] for entry in Nebkd[1]],dtype='float')*1e6
plt.figure(figsize=(9,4))
plt.scatter(Amu, Neon - Nebkd);
ax = plt.gca()
plt.xlim(12,35);
plt.ylim(0,3.2)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.savefig('Neon.png')
# -
# 20 is Neon. 22 is Unknown. Possibly doubly ionized Neon that grabbed two Hydrogen atoms? Seems far fetched.
# ## Air
# +
Air = pd.read_table('Air.txt',delimiter=', ',engine='python', header=None)
Air = np.array([entry[:-1] for entry in Air[1]],dtype='float')*1e6
plt.figure(figsize=(9,4))
plt.scatter(Amu, Air - Nebkd);
ax = plt.gca()
plt.xlim(12,45);
plt.ylim(0,3.2)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.savefig('Air.png')
# -
np.argmax(Air[250:289])
np.argmax(Air[289:])
np.argmax(Air[330:])
Air[250:289][21]/Air[289:][22]
Air[250:289][21]/Air[330:][61]
# Expected ratios are N2/O2 = 3.7 and N2/Ar = 89
# # Day 2
# ## Quick Exhale vs Hold Breath
# +
Quick = pd.read_table('QuickExhale.txt',delimiter=', ',engine='python', header=None)
Quick = np.array([entry[:-1] for entry in Quick[1]],dtype='float')*1e6
Quickbkd = pd.read_table('Background_Breath.txt',delimiter=', ',engine='python', header=None)
Quickbkd = np.array([entry[:-1] for entry in Quickbkd[1]],dtype='float')*1e6
Hold = pd.read_table('HoldBreath30s.txt',delimiter=', ',engine='python', header=None)
Hold = np.array([entry[:-1] for entry in Hold[1]],dtype='float')*1e6
plt.figure(figsize=(9,4))
plt.scatter(Amu, Quick - Quickbkd,color='blue',label='Quick Exhale');
plt.scatter(Amu, Hold - Quickbkd,color='red',label = 'Hold Breath');
ax = plt.gca()
plt.xlim(12,35);
plt.ylim(0,8.5)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.legend(loc='upper left')
plt.savefig('Breath.png')
# -
np.argmax(Quick[250:289])
Quick[250:290][22]
np.argmax(Quick[289:])
Quick[289:][23]
Quick[250:290][22]/Quick[289:][23]
np.argmax(Hold[250:289])
np.argmax(Hold[289:])
Hold[250:290][22]/Hold[289:][23]
# Note that we were unable to gaurentee that the leak valve was in the exact same position across these two measurements. Qualitatively, we can only extract that in the quick exhale situation the ratio of N2/O2 was 4.68, while in the holding of breath it was 4.51. Not quite as expected.
# ## Compressed Air Comparison
# +
Can1 = pd.read_table('CompressedAir_Tetrafluoroethane.txt',delimiter=', ',engine='python', header=None)
Can1 = np.array([entry[:-1] for entry in Can1[1]],dtype='float')*1e6
Can2 = pd.read_table('CompressedAir_Difluoroethane.txt',delimiter=', ',engine='python', header=None)
Can2 = np.array([entry[:-1] for entry in Can2[1]],dtype='float')*1e6
plt.figure(figsize=(9,4))
plt.scatter(Amu, Can1 - Quickbkd,color='blue',label='Tetrafluoroethane');
plt.scatter(Amu, Can2 - Quickbkd,color='red',label = 'Difluoroethane');
ax = plt.gca()
plt.xlim(10,65);
plt.ylim(0,8.5)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.legend(loc='upper right')
plt.savefig('CompressedAir.png')
# -
# Peak at 19 possibly fluorine
# +
Volcano = pd.read_table('Volcano.txt',delimiter=', ',engine='python', header=None)
Volcano = np.array([entry[:-1] for entry in Volcano[1]],dtype='float')*1e6
VolcanoBackground = pd.read_table('VolcanoBackground.txt',delimiter=', ',engine='python', header=None)
VolcanoBackground = np.array([entry[:-1] for entry in VolcanoBackground[1]],dtype='float')*1e6
plt.figure(figsize=(9,4))
plt.scatter(Amu, Volcano - VolcanoBackground);
ax = plt.gca()
plt.xlim(10,35);
plt.ylim(0,8.5)
plt.xlabel('Particle Mass [Amu]',size=18);
plt.ylabel('Pressure [Torr]$\cdot 10^{-6}$',size=18);
plt.xticks(size = 11);
plt.yticks(size = 11);
plt.savefig('Volcano.png')
# -
| MassSpectrometer/BackgroundSubstract.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pyttsx3
import wikipedia
import speech_recognition as sr
import webbrowser
import datetime
import os
import sys
import smtplib
from news import speak_news
from diction import translate
from loc import weather
from youtube import you
engine = pyttsx3.init()
voices = engine.getProperty('voices')
engine.setProperty('voice', voices[0].id)
# print(voices[0].id)
# -
def speak(audio):
engine.say(audio)
engine.runAndWait()
def takeCommand():
r = sr.Recognizer()
with sr.Microphone() as source:
print('Listening...')
r.pause_threshold = 1
r.energy_threshold = 494
r.adjust_for_ambient_noise(source, duration=1.5)
audio = r.listen(source)
try:
print('Recognizing..')
query = r.recognize_google(audio, language='en-in')
print(f'User said: {query}\n')
except Exception as e:
# print(e)
print('Say that again please...')
return 'None'
return query
def wishMe():
hour = int(datetime.datetime.now().hour)
if hour >= 0 and hour < 12:
speak("Good Morning SIR")
elif hour >= 12 and hour < 18:
speak("Good Afternoon SIR")
else:
speak('Good Evening SIR')
weather()
speak('I am JARVIS. Please tell me how can I help you SIR?')
def sendEmail(to, content):
server = smtplib.SMTP('smtp.gmail.com', 587)
server.ehlo()
server.starttls()
server.login('email', 'password')
server.sendmail('email', to, content)
server.close()
if __name__ == '__main__':
chrome_path = 'C:\\Program Files (x86)\\Google\\Chrome\\Application\\chrome.exe'
webbrowser.register('chrome', None, webbrowser.BackgroundBrowser(chrome_path))
wishMe()
while True:
query = takeCommand().lower()
if 'wikipedia' in query:
speak('Searching Wikipedia....')
query = query.replace('wikipedia', '')
results = wikipedia.summary(query, sentences=2)
speak('According to Wikipedia')
print(results)
speak(results)
elif 'open youtube' in query:
webbrowser.get('chrome').open_new_tab('https://youtube.com')
elif 'open google' in query:
webbrowser.get('chrome').open_new_tab('https://google.com')
elif 'open stackoverflow' in query:
webbrowser.get('chrome').open_new_tab('https://stackoverflow.com')
elif 'play music' in query:
os.startfile("D:\\RoiNa.mp3")
elif 'search youtube' in query:
speak('What you want to search on Youtube?')
you(takeCommand())
elif 'the time' in query:
strTime = datetime.datetime.now().strftime("%H:%M:%S")
speak(f'Sir, the time is {strTime}')
elif 'search' in query:
speak('What do you want to search for?')
search = takeCommand()
url = 'https://google.com/search?q=' + search
webbrowser.get('chrome').open_new_tab(url)
speak('Here is What I found for' + search)
elif 'location' in query:
speak('What is the location?')
location = takeCommand()
url = 'https://google.nl/maps/place/' + location + '/&'
webbrowser.get('chrome').open_new_tab(url)
speak('Here is the location ' + location)
elif 'your master' in query:
speak('Gaurav is my master. He created me couple of days ago')
elif 'your name' in query:
speak('My name is JARVIS')
elif 'stands for' in query:
speak('J.A.R.V.I.S stands for JUST A RATHER VERY INTELLIGENT SYSTEM')
elif 'open code' in query:
os.startfile("C:\\Users\\gs935\\AppData\\Local\\Programs\\Microsoft VS Code\\Code.exe")
elif 'shutdown' in query:
os.system('shutdown /p /f')
elif 'github' in query:
webbrowser.get('chrome').open_new_tab('https://github.com/gauravsingh9356')
elif 'sleep' in query:
sys.exit()
elif 'dictionary' in query:
speak('What you want to search in your intelligent dictionary?')
translate(takeCommand())
elif 'news' in query:
speak('Ofcourse sir..')
speak_news()
elif 'email to gaurav' in query:
try:
speak('What should I say?')
content = takeCommand()
to = 'email'
sendEmail(to, content)
speak('Email has been sent!')
except Exception as e:
speak('Sorry sir, Not able to send email at the moment')
| .ipynb_checkpoints/jarvis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Stack Overflow Questions by Library Data Collection
#
# The following code was used to collect the data from the Stack Overflow API's [/search](https://api.stackexchange.com/docs/search) endpoint. Be advised that due to throttling running this code yourself will take a long time to complete.
# +
import datetime as dt
import json
import re
import requests
import time
from IPython import display
libraries = [
'pandas',
'matplotlib',
'numpy',
'seaborn',
'geopandas',
'geoviews',
'altair',
'yellowbrick',
'vega',
'holoviews',
'hvplot',
'bokeh'
]
stack_overflow_tags = (
'https://api.stackexchange.com/'
'/2.3/search?page={}&pagesize=100&order=asc&sort=creation'
f'&tagged={";".join(libraries)}'
'&site=stackoverflow'
)
def throttle(wait):
resume_time = str(dt.datetime.now() + dt.timedelta(seconds=wait))
print(f'Waiting until {resume_time} before resuming...')
time.sleep(wait)
done = False
questions = []
page = 1
while not done:
display.clear_output(wait=True)
print(f'Requesting page {page} at {str(dt.datetime.now())}...')
response = requests.get(stack_overflow_tags.format(page))
if not response.ok:
if response.status_code == 400:
message = response.json()
if message['error_name'] == 'throttle_violation':
print(message['error_message'])
wait = int(re.search(r'\d+', message['error_message'])[0])
throttle(wait)
continue
print(f'Request failed with status {response.status_code}')
break
results = response.json()
done = not bool(results['has_more'])
page += 1
questions.extend([
{
'tags': question['tags'],
'creation_date': question['creation_date'],
'title': question['title']
}
for question in results['items']
])
latest_question_creation_date = str(dt.datetime.fromtimestamp(questions[-1]['creation_date']))
print(f'Latest question retrieved was created at {latest_question_creation_date}')
# make sure we don't go too fast
print(f'Remaining quota: {results["quota_remaining"]}')
throttle(results.get('backoff', 1 if page % 10 else 10))
display.clear_output(wait=True)
print('Done')
with open('../stackoverflow_questions.json', 'w') as file:
json.dump(questions, file)
# -
# Questions collected:
len(questions)
# ## Processing the data for files used in the workshop
# +
import pandas as pd
df = pd.read_json('../stackoverflow_questions.json').assign(
creation_date=lambda x: pd.to_datetime(x.creation_date, unit='s')
).set_index('creation_date')
df.head()
# -
# *Source: Stack Exchange Network*
#
# Create Boolean columns for each libary, indicating whether the question was tagged with that library:
df = df.assign(
**{
library: df.tags.apply(lambda y: f'{library}' in y)
for library in libraries
}
)
df.head()
# Pandas is the only library with more questions in this dataset tagged than not:
df.iloc[:,2:].describe()
# Save this to a CSV file:
df.to_csv('../stackoverflow.zip', compression=dict(method='zip', archive_name='stackoverflow.csv'))
# ##### Questions by tag
# We will look at questions tagged with the libraries above, but there are other tags in this dataset. Out of curiosity, what are the most common tags in this dataset overall?
# +
from collections import Counter
cntr = Counter()
for i in df.tags:
cntr.update(i)
cntr.most_common()[:10]
# -
# ##### Looking at co-occurrences of libraries within a single question
# How often are multiple libraries from our list tagged in a single question?
df.iloc[:,2:].assign(co_occurrence=lambda x: x.sum(axis=1) > 1).co_occurrence.value_counts(normalize=True)
# The previous result is a little misleading because a majority of the questions are about pandas. If we look at this per library, we will see that there are some libraries that are present with others most of the time:
co_occur = df.iloc[:,2:]\
.assign(co_occurrence=lambda x: x.sum(axis=1) > 1)\
.groupby('co_occurrence').sum().T
co_occur.apply(lambda x: x / co_occur.sum(axis=1))
# ##### Adjacency matrix of co-occurences
# We will now create an adjacency matrix out of this information. Some questions are tagged with more than 1 additional library, so we will share the credit between them to be able to make a stacked bar plot with this information as percentages:
# +
def get_co_occurrences(df, library, normalize=True):
div = df.query(f'{library} == 1').sum(axis=1)
data = df.query(f'{library} == 1').apply(
lambda x: x / (div - df.query(f'{library} == 1')[library]) if x.name != library else x
)
co_occur = pd.Series(data.sum().values, index=df.columns)
if normalize:
co_occur /= co_occur[library]
return co_occur
def get_adj_matrix(df, libraries, normalize=True, nullify_diagonal=True):
data = df[libraries].apply(lambda x: x.astype(int))
data.columns = data.columns.rename('library')
adj_matrix = data.apply(lambda x: get_co_occurrences(data, x.name, False))
sort_order = adj_matrix.sum().sort_values(ascending=False).index
adj_matrix.index = adj_matrix.index.rename('co-occurring library')
if normalize:
adj_matrix /= data.sum()
if nullify_diagonal:
for i in range(len(libraries)):
adj_matrix.iloc[i, i] = 0
sort_order = adj_matrix.sum().sort_values(ascending=False).index
adj_matrix = adj_matrix.reindex(columns=sort_order)
else:
for library in libraries:
if normalize:
adj_matrix.loc[library, library] = 2 - adj_matrix.loc[:,library].sum()
else:
adj_matrix.loc[library, library] = (data.query(f'{library} == 1').sum(axis=1) == 1).sum()
return adj_matrix.reindex(sort_order).reindex(columns=sort_order)
# -
# We will build the adjacency matrix for a subset of the libraries. Note that the diagonal is set to 0 since we are calculating co-occurences only:
subset = ['pandas', 'numpy', 'matplotlib', 'seaborn', 'bokeh', 'holoviews', 'geoviews', 'geopandas', 'hvplot']
co_occur_pct = get_adj_matrix(
df, subset, normalize=True, nullify_diagonal=True
)
co_occur_pct.index = co_occur_pct.index.rename('co-occurring library')
co_occur_pct.T.to_csv('../stackoverflow_tag_co_occurrences.csv')
co_occur_pct
# While we won't use Seaborn in this workshop, it's very helpful at visualizing data in this format. Here, we make a heatmap:
# +
import matplotlib.pyplot as plt
import seaborn as sns
fig, ax = plt.subplots(figsize=(10, 10))
data = get_adj_matrix(
df, subset, normalize=True, nullify_diagonal=False
).T
sns.heatmap(
data, mask=data.round(4) < 0.01, annot=True, fmt='.1%',
cmap='Blues', cbar_kws={'shrink': 0.75},
square=True, vmin=0, vmax=1, center=0.5, ax=ax
)
ax.set(
xlabel='co-occurring library', ylabel='library',
title=(
'Co-Occurrence Percentages\n(diagonal represents the '
'percentage of questions tagged only with that library)'
)
)
fig.tight_layout()
| data/collection/stackoverflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This module runs ANN on TFIDF
#make necessary imports
import numpy as np
import pandas as pd
import itertools
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.linear_model import LogisticRegression
#Read the data
df=pd.read_csv('../../datasets/liar_tweaked/trainvectordata.csv')
testdf=pd.read_csv('../../datasets/liar_tweaked/testvectordata.csv')
validdf=pd.read_csv('../../datasets/liar_tweaked/validvectordata.csv')
x_train,y_train=df['statement'],df['label']
x_test,y_test=testdf['statement'],testdf['label']
x_valid,y_valid=validdf['statement'],validdf['label']
#tfidf
tfidf_vectorizer=TfidfVectorizer(stop_words='english', max_df=0.7)
tfidf_train=tfidf_vectorizer.fit_transform(df['statement'])
tfidf_test=tfidf_vectorizer.transform(testdf['statement'])
tfidf_valid=tfidf_vectorizer.transform(validdf['statement'])
tfidf_train
#building the classifier
def build_classifier():
clf=Sequential()
clf.add(Dense(output_dim=500,init='uniform',activation='relu',input_dim=11915))
clf.add(Dense(output_dim=100,init='uniform',activation='relu'))
clf.add(Dense(output_dim=50,init='uniform',activation='relu'))
clf.add(Dense(output_dim=20,init='uniform',activation='relu'))
clf.add(Dense(output_dim=10,init='uniform',activation='relu'))
clf.add(Dense(output_dim=5,init='uniform',activation='relu'))
clf.add(Dense(output_dim=1,init='uniform',activation='sigmoid'))
clf.compile(optimizer='adam', loss='binary_crossentropy',metrics=['accuracy'])
return clf
# +
#make necessary imports
import keras
from keras.models import Sequential
from keras.layers import Dense
from keras.wrappers.scikit_learn import KerasClassifier
from sklearn.model_selection import cross_val_score
#build ANN, use k fold cross validation
clf=KerasClassifier(build_fn=build_classifier, batch_size=10, nb_epoch=100)
accuracies=cross_val_score(estimator=clf, X=tfidf_train,y=df['label'],cv=10,n_jobs=-1)
# -
#see accuracies
accuracies
#fit on training data and check accuracies on both test and valid data
clf.fit(tfidf_train,y_train, batch_size=10, nb_epoch=10)
y_test_pred = clf.predict(tfidf_test)
print('algorithm - test dataset accuracy - valid dataset accuracy')
print('ANNTFIDF - ' ,round(accuracy_score(y_test, y_test_pred),4), ' - ', end='')
y_test_pred = clf.predict(tfidf_valid)
print(round(accuracy_score(y_valid, y_test_pred),4))
| models/ANN/.ipynb_checkpoints/ANNtfidf-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Taken from https://github.com/alexjungaalto/ResearchPublic/blob/master/RoadMarkingHelsinki/RoadMarkingMonitoring.ipynb
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from PIL import Image
import datetime
import numpy as np
import cv2
import pandas as pd
import matplotlib.pyplot as plt
from tensorflow.keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from owslib.wms import WebMapService
from skimage.feature import hog
import io
wms = WebMapService('https://kartta.hel.fi/ws/geoserver/avoindata/wms', version='1.1.1')
# define the bounding box of the entire city area
xmin = 24.92
ymin = 60.15
xmax = 24.99
ymax = 60.20
# we divide the entire city area evenly into nr_patches_x x nr_patches_y square patches
nr_patches_x = 50
nr_patches_y = 50
patch_x = (xmax-xmin)/nr_patches_x # width of an individual patch
patch_y = (ymax-ymin)/nr_patches_y # height of a single patch
# +
# choose one particular patch
nr_x = 11
nr_y = 10
# determine the bounding box for this particular patch
patch_box = (xmin+nr_x*patch_x,ymin+nr_y*patch_y,xmin+(nr_x+1)*patch_x,ymin+(nr_y+1)*patch_y)
ortholayer = 'avoindata:Ortoilmakuva_2019_5cm' # choose layer with aerial photographs
cs = 'EPSG:4326'
resx=500 # choose number of pixels in one direction
res = (resx,resx)
fig, axes = plt.subplots(1, 2, figsize=(15, 15))
img = wms.getmap(layers=[ortholayer],srs=cs,bbox=patch_box,size=res,format='image/jpeg',transparent=True)
A_1 = np.asarray(Image.open(io.BytesIO(img.read())))
axes[0].imshow(A_1)
axes[0].set_title("Layer obtained from the aerial photographs.")
img = wms.getmap(layers=['Karttasarja'],srs=cs,bbox=patch_box,size=res,format='image/jpeg',transparent=True)
A_1_layer_1 = np.asarray(Image.open(io.BytesIO(img.read())))
axes[1].imshow(A_1_layer_1)
axes[1].set_title("Layer obtained from the building map.")
plt.show()
# -
# # Getting the data
# +
import geopandas
import fiona
import pandas as pd
import numpy as np
from owslib.wms import WebMapService
import os
file=geopandas.read_file("klinj_suojatie.gpkg")
df=pd.DataFrame(file)
df.geometry=df.geometry.astype("str")
df["geometry"]=df["geometry"].str.replace("LINESTRING ","").str.replace(",","").str.replace("(","").str.replace(")","")
df[["min_x","max_y","max_x","min_y","a","b"]]=df["geometry"].str.split(" ",expand=True)
df=df.drop(columns=["a","b","geometry"])
df["id"]=df["gml_id"].str.replace("klinj_suojatie.","")
df["min_x"]=df["min_x"].astype(float)
df["min_y"]=df["min_y"].astype(float)
df["max_x"]=df["max_x"].astype(float)
df["max_y"]=df["max_y"].astype(float)
# Fixing inconsistencies
df.min_x, df.max_x = np.where(df.min_x > df.max_x, [df.max_x, df.min_x], [df.min_x, df.max_x])
df.min_y, df.max_y = np.where(df.min_y > df.max_y, [df.max_y, df.min_y], [df.min_y, df.max_y])
resolution = 100
# choose layers which are used for the pedestrian crossing images
layer = "Ortoilmakuva_2019_5cm"
#layer = "Vaaravariortoilmakuva_2019_20cm"
folder_name ="PedestrianCrossingImg/resolution_"+str(resolution)+"_layer_"+layer+"/"
if not os.path.exists(folder_name):
os.makedirs(folder_name)
filelist=os.listdir(folder_name)
nrfiles = len(filelist)
if nrfiles < 100 :
wms = WebMapService('https://kartta.hel.fi/ws/geoserver/avoindata/wms', version='1.3.0')
for i in range(0,len(df)):
# determine bounding box of ith pedestrian crossing
minx = df.loc[i,"min_x"]
miny = df.loc[i,"min_y"]
maxx = df.loc[i,"max_x"]
maxy = df.loc[i,"max_y"]
deltax = (maxx-minx)
deltay = (maxy-miny)
roi = (minx-deltax,miny-deltay,maxx+deltax,maxy+deltay)
#read in image patch from server
img=wms.getmap(layers=[layer],srs="EPSG:3879",bbox=roi,size=(resolution,resolution),format='image/png',transparent=True)
filename =folder_name + str(df.loc[i,"id"])+".png"
#filename = "pedestrian_crossing" + str(i)+".png"
out = open(filename, 'wb')
out.write(img.read())
out.close()
| Data preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="UIbfV6292_nr"
import numpy as np
import pandas as pd
import seaborn as sns
import scipy
import tables as tb
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] colab_type="text" id="UlaZAr522_n1"
# # Load Datasets
# -
test_data = pd.read_csv("testing_data_transform.csv.gz", compression="gzip")
test_data.head()
training_data = pd.read_csv("training_data_transform.csv.gz", compression="gzip")
training_data.head()
# # Training Data
vars = list(set(test_data.columns) - {"data_ind", "index"})
vars
# + colab={} colab_type="code" id="Or0YWe0vM1jx"
from sklearn.model_selection import train_test_split
# + colab={} colab_type="code" id="qU2LTdaVM0_c"
X_train, X_valid, y_train, y_valid = train_test_split(train_data[varaibles],
train_data.signal,
test_size=0.10,
random_state=42)
# +
idx = ~X_train.TX_pair.isna()
X_train.loc[~idx, list(set(X_train.columns) - {"TX", "TY", "X", "Y", "Z"})] = np.random.normal(size=(27876, 10))
idx = ~X_valid.TX_pair.isna()
X_valid.loc[~idx, list(set(X_valid.columns) - {"TX", "TY", "X", "Y", "Z"})] = np.random.normal(size=(3208, 10))
del idx
# + [markdown] colab_type="text" id="JtD_F2p82_o6"
# # XGBoost Model
# + colab={} colab_type="code" id="uap8SSOi2_o-"
import xgboost as xg
from xgboost import XGBClassifier
from sklearn.model_selection import StratifiedKFold, GridSearchCV
# + colab={} colab_type="code" id="Fco13gac2_pC"
param_grid = {
'n_estimators':[10, 20],
'max_depth':[15],
}
class XGBClassifier_tmp(XGBClassifier):
def predict(self, X):
return XGBClassifier.predict_proba(self, X)[:, 1]
clf = GridSearchCV(XGBClassifier_tmp(learning_rate=0.05,
subsample=0.8,
colsample_bytree=0.8,
n_jobs=-1,
tree_method='gpu_hist',
gpu_id=0,
warm_start=True),
param_grid=param_grid,
n_jobs=-1,
scoring='roc_auc',
verbose=7)
# + colab={"base_uri": "https://localhost:8080/", "height": 452} colab_type="code" id="hnDYBGEO2_pG" outputId="c4b744e1-a50a-47e2-b255-f7fd57e49916"
# %%time
clf.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 150} colab_type="code" id="o6dkOioI2_pN" outputId="bb1a9a23-a20e-498a-b4f4-f29323cad379"
clf.best_estimator_
# + colab={} colab_type="code" id="VdVBQt312_pS"
xgb_class = XGBClassifier_tmp(base_score=0.5,
booster='gbtree',
tree_method='gpu_hist',
gpu_id=0,
colsample_bylevel=1,
colsample_bytree=0.8,
gamma=0,
learning_rate=0.05,
max_delta_step=0,
max_depth=15,
min_child_weight=1,
missing=None,
n_estimators=100,
nthread=None,
objective='binary:logistic',
random_state=0,
reg_alpha=0,
reg_lambda=1,
scale_pos_weight=1,
seed=None,
silent=True,
subsample=0.8,
n_jobs=-1)
# + [markdown] colab_type="text" id="A-RQoVNON4uo"
# ### Validation
# + colab={} colab_type="code" id="F5tB1WNyKVpO"
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score, precision_score, recall_score
from scikitplot.metrics import plot_confusion_matrix, plot_roc
def evaluate_performace(y_true, y_pred, threshold=0.5):
y_probs = np.hstack([1-y_pred.reshape(-1, 1), y_pred.reshape(-1, 1)])
y_pred = y_pred >= threshold
print("Accuracy Score: ", accuracy_score(y_true, y_pred))
print("Precision Score: ", precision_score(y_true, y_pred))
print("Recall Score: ", recall_score(y_true, y_pred))
plot_confusion_matrix(y_true, y_pred, normalize=True, cmap="winter")
plot_roc(y_true, y_probs, classes_to_plot=[0], plot_macro=False, plot_micro=False)
plt.show()
# + colab={} colab_type="code" id="femCHZNZJrsh"
y_pred = clf.predict(X_valid)
# + colab={"base_uri": "https://localhost:8080/", "height": 623} colab_type="code" id="YlsmQVgVYc5z" outputId="63be7970-2d80-4050-88b5-b77afcc6ba50"
evaluate_performace(y_valid, y_pred, threshold=0.5)
# + [markdown] colab_type="text" id="hAtP2vQl2_pX"
# # Final Prediction
# + colab={} colab_type="code" id="T5Gp7R2DFDgb"
clf.fit(X_valid, y_valid)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="3bxfS1Os2_pY" outputId="88b3cd28-3024-4349-fbe4-80028314419e"
X_test = test_data[vars]
# + colab={} colab_type="code" id="si12cboF2_ph"
idx = ~X_test.TX_pair.isna()
X_test.loc[~idx, list(set(X_test.columns) - {"TX", "TY", "X", "Y", "Z"})] = np.random.normal(size=(128639, 10))
# + colab={} colab_type="code" id="C-7-2VyuCvcd"
probas = clf.predict(X_test, batch_size=5000)
# + colab={} colab_type="code" id="pW30coRF2_pn"
df = pd.DataFrame({'id': test_data["index"], 'signal': probas.reshape(-1)}).groupby('id')
agg = df.aggregate(('mean')).loc[:, ['signal']]
# + colab={} colab_type="code" id="AMLvD6B82_p2"
agg.to_csv('submission.csv.gz', index=True, compression='gzip')
# -
agg
| Dark-Matter-Signature/XGBoosted Trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
import cPickle as pickle
import codecs
from operator import itemgetter
from matplotlib import pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn import metrics
from sklearn.cluster import KMeans
from sklearn.cluster.bicluster import SpectralCoclustering
from biclustering.biclustering import DeltaBiclustering
from sklearn.metrics.cluster import normalized_mutual_info_score
from sklearn.metrics.cluster import adjusted_rand_score
# -
# %matplotlib inline
sns.set_palette("deep", desat=.6)
sns.set_context(rc={"figure.figsize": (8, 4)})
arena_news_stem_df = pd.read_pickle('arena_news_stem_df.pkl')
sport_news_stem_df = pd.read_pickle('sport_news_stem_df.pkl')
jovem_news_stem_df = pd.read_pickle('jovem_news_stem_df.pkl')
arena_news_df = pd.read_pickle('arena_news_df.pkl')
sport_news_df = pd.read_pickle('sport_news_df.pkl')
jovem_news_df = pd.read_pickle('jovem_news_df.pkl')
labels = np.array(len(arena_news_df)*[1] + len(sport_news_df)*[0])
all_news_df = arena_news_df.append(sport_news_df)
all_news_df.reset_index(inplace=True)
all_news_df.drop('index', axis=1, inplace=True)
all_news_df
for news in arena_news_df['all']:
c = 0
if u'playstation' in news or u'Playstation' in news or u'PlayStation' in news:
c += 1
print news
print c
# +
count_vect = CountVectorizer(encoding='UTF-8',lowercase=False,min_df=2)
X = count_vect.fit_transform(arena_news_df['all'].tolist() + sport_news_df['all'].tolist())
X_train_norm_tfidf = TfidfTransformer(norm=u'l2', use_idf=True).fit_transform(X)
X_train_norm = TfidfTransformer(norm=u'l2', use_idf=False).fit_transform(X)
# -
def to_csv(some_list, file_name, header):
def to_str(st):
if isinstance(st, str) or isinstance(st, unicode):
return st
else:
return str(st)
with codecs.open(file_name, 'w', 'utf-8') as file_stream:
file_stream.write(u','.join(header) + '\n')
for item in some_list:
if isinstance(item, tuple) or isinstance(item, list):
line = u','.join(map(to_str, item)) + '\n'
else:
line = item + '\n'
file_stream.write(line)
file_stream.close()
# # All words top 20
# +
def top_k(arr, k, axis=0):
top_inds = np.argsort(arr, axis=axis)[-k:]
top_vals = np.sort(arr, axis=axis)[-k:]
return top_inds, top_vals
def reverse(arr):
return arr[::-1]
def top_k_words(vec, count_vect, k):
top_inds, top_vals = top_k(vec, k)
words = count_vect.get_feature_names()
top_words = [words[i] for i in top_inds]
top_pairs = reverse(zip(top_words, top_vals))
return top_pairs
sum_per_word = np.sum(X_train_norm.toarray(), axis=0)
top_pairs = top_k_words(sum_per_word, count_vect, 9999999)
for word, value in top_pairs[0:20]:
print '%s, %s' % (word, value)
to_csv(top_pairs, 'all_words_top.csv', ['word', 'tf norm value'])
# -
# # Arena words top 20
sum_per_word = np.sum(X_train_norm.toarray()[0:100, :], axis=0)
top_pairs = top_k_words(sum_per_word, count_vect, 9999999)
for word, value in top_pairs[0:20]:
print '%s, %s' % (word, value)
to_csv(top_pairs, 'arena_words_top.csv', ['word', 'tf norm value'])
# # Sports words top 20
sum_per_word = np.sum(X_train_norm[100:200, :], axis=0)
top_pairs = top_k_words(sum_per_word, count_vect, 9999999)
for word, value in top_pairs[0:20]:
print '%s, %s' % (word, value)
to_csv(top_pairs, 'sports_words_top.csv', ['word', 'tf norm value'])
| TopWords.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mypython3
# language: python
# name: mypython3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import xarray as xr
#import cf_xarray as cfxr
#import cftime as cfdt
#import pandas as pd
#from datetime import datetime as dt
#from datetime import timedelta as td
import datetime as dt
import pyremo as pr
import cordex as cx
#import xclim as xc
import glob
#import sys, os
#import numpy as np
#from urllib.request import urlopen
#import json
#import cmor
#import iteration_utilities
#from pyhomogenize.time_control import time_control
# -
from dask.distributed import Client
client = Client()
client
# ### Define important functions
def open_mfdataset(files,
use_cftime=True,
parallel=True,
data_vars='minimal',
chunks={'time':1},
coords='minimal',
compat='override',
drop=None,
preprocess=None,
**kwargs):
"""optimized function for opening large cf datasets.
based on https://github.com/pydata/xarray/issues/1385#issuecomment-561920115
"""
def drop_all_coords(ds):
if preprocess is not None:
ds = preprocess(ds, use_cftime=use_cftime)
return ds.reset_coords(drop=True)
ds = xr.open_mfdataset(files,
parallel=parallel,
decode_times=False,
combine='by_coords',
preprocess=drop_all_coords,
decode_cf=False,
chunks=chunks,
data_vars=data_vars,
coords='minimal',
compat='override',
**kwargs)
return xr.decode_cf(ds, use_cftime=use_cftime)
def select_domain(ds, domain):
domain = cx.cordex_domain(domain)
# the method=='nearest' approach does not work well with dask
return ds.sel(rlon=slice(domain.rlon.min(), domain.rlon.max()), rlat=slice(domain.rlat.min(), domain.rlat.max()))
# ### Read hourly REMO raw data
pattern = '/work/ch0636/g300046/remo_results_056000/1979/hourly/e056000e_c167_197901.nc'
filenames = glob.glob(pattern)
filenames
ds = open_mfdataset(filenames, chunks={}, parallel=True, preprocess=pr.preprocess)
ds
ds.TEMP2.isel(time=0).plot()
# #### Example 3-hourly data
# #### resample function (alread implemented in pyremo but not in __init__.py)
loffsets = {"3H": dt.timedelta(hours=1, minutes=30), "6H": dt.timedelta(hours=3)}
def _get_loffset(time):
return loffsets.get(time, None)
def _resample(
ds, time, time_cell_method="point", label="left", time_offset=True, **kwargs
):
"""Resample a REMO variable."""
if time_cell_method == "point":
return ds.resample(time=time, label=label, **kwargs).interpolate("nearest")
elif time_cell_method == "mean":
if time_offset is True:
loffset = _get_loffset(time)
else:
loffset = None
return ds.resample(time=time, label=label, loffset=loffset, **kwargs).mean()
else:
raise Exception("unknown time_cell_method: {}".format(time_cell_method))
ds_resampled = _resample(ds, time='3H', time_cell_method='point', label=None, time_offset=False)
ds_resampled
ds_resampled.TEMP2.isel(time=0).plot()
(ds_resampled.TEMP2.isel(time=1) - ds.TEMP2.isel(time=2)).plot()
# #### resample method nearest produces a Nan-array for 1979-01-01 00:00:00 (timestep not available in raw data)
# #### Clear those empty arrays
def _clear_time_axis(ds):
for data_var in ds.data_vars:
ds = ds.dropna(dim='time',how='all')
return ds
ds_cleared = _clear_time_axis(ds_resampled)
ds_cleared.time
ds_cleared.TEMP2.isel(time=0).plot()
(ds_cleared.TEMP2.isel(time=0) - ds.TEMP2.isel(time=2)).plot()
filename = pr.cmor.cmorize_variable(ds_cleared, 'tas', '3hr', cx.cordex_cmor_table('remo_example'),
CORDEX_domain='EUR-11', allow_units_convert=True)
filename
# #### Chunk the file
ds_grouped = ds_cleared.groupby('time.dayofyear', squeeze=False)
ds_grouped.groups
# #### add groups with time lenght equals 1 to previous or next group
groups_dict = {}
i=1
while i <= len(ds_grouped):
if len(ds_grouped[i].time) == 1:
if i-1 in groups.keys():
groups_dict[i-1] = xr.concat([groups_dict[i-1],ds_grouped[i]], dim='time')
else:
groups_dict[i+1] = ds_grouped[i]
elif i in groups_dict.keys():
groups_dict[i] = xr.concat([groups_dict[i], ds_grouped[i]], dim='time')
else:
groups_dict[i] = ds_grouped[i]
i += 1
groups_dict
for name, group in groups_dict.items():
filename = pr.cmor.cmorize_variable(group, 'tas', '3hr', cx.cordex_cmor_table('remo_example'),
CORDEX_domain='EUR-11', allow_units_convert=True)
print(filename)
| notebooks/remo-cmor-hourly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: clouds113_kernel
# language: python
# name: clouds113_kernel
# ---
# # Coarse-graining NARVAL Data
# **For Figure 1 of the paper**
# +
import os
import sys
import xarray as xr
import numpy as np
import pandas as pd
import importlib
import matplotlib
import matplotlib.pyplot as plt
# For psyplot
import psyplot.project as psy
import matplotlib as mpl
# # %matplotlib inline
# # %config InlineBackend.close_figures = False
psy.rcParams['plotter.maps.xgrid'] = False
psy.rcParams['plotter.maps.ygrid'] = False
mpl.rcParams['figure.figsize'] = [10., 8.]
# -
path = '/pf/b/b309170/my_work/NARVAL/'
file_cg = 'for_paraview/clc_R02B04_NARVALII_2016072800_cloud_DOM01_0017_boxed_scaled.nc'
file_orig = 'for_paraview/dei4_NARVALII_2016072800_cloud_DOM01_ML_0017_clc_scaled.nc'
# #### Question 1: Does the coarse-graining look right?
# Of horizontal coarse-graining (psyplot): <br>
#
# If you get the error 'ValueError: Can only plot 2-dimensional data!', then you need to use cdo setgrid on the file first.
# +
# # Note that the cloud cover scheme used was a 0-1 cloud cover scheme.
# maps = psy.plot.mapplot(os.path.join(path, file_orig), dims = {'name': 'ccl', 'height': 40},
# projection='robin', cmap='Blues_r', title='Cloud cover on 20041105 at 15:00 (on layer 40)')
# plt.savefig('original_cloud_cover_snapshot.pdf')
# Note that the cloud cover scheme used was a 0-1 cloud cover scheme.
maps = psy.plot.mapplot(os.path.join(path, file_orig), dims = {'name': 'clc', 'height': 40}, cticksize=34,
projection='robin', cmap='Blues_r')
# maps.update(lonlatbox=[-180, 180, -90, 90])
plt.savefig('original_cloud_cover_snapshot_untitled_narval.pdf')
# +
# I had to cdo sellonlatbox first. Zooming in via lonlatbox in maps update did not work!
maps = psy.plot.mapplot(os.path.join(path, file_cg), dims = {'name': 'clc', 'height': 40}, cticksize=34,
projection='robin', cmap='Blues_r', bounds=[0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1])
# maps.update(lonlatbox=[-68, 15, -10, 20]) #[lon.min(), lon.max(), lat.min(), lat.max()]
# maps.update(lonlatbox=[-180, 180, -90, 90])
# plt.savefig('horizontally_coarse_grained_cloud_cover_untitled_narval.pdf')
# plt.savefig('test_2.pdf', bbox_inches=Bbox([[4, 4], [6, 6]]))
# -
# Of vertical coarse-graining:
# +
## Load original data
# Load clc profile
DS = xr.open_dataset('/pf/b/b309170/my_work/NARVAL/data/clc/clc_R02B04_NARVALII_2016072800_cloud_DOM01_0017.nc')
da = DS.clc.values
print(da.shape)
# Extract all nan_fields
nan_fields = np.where(~np.isnan(da[0, -1, :]))[0]
# # Some arbitrary horizontal field
rand_field = np.random.randint(len(nan_fields))
rand_field = nan_fields[rand_field]
# rand_field=10084 # To reconstruct the profile from the paper
print(rand_field)
cl_hr = da[0, :, rand_field]
# Load zg profile
DS = xr.open_dataset('/pf/b/b309170/my_work/NARVAL/data/z_ifc/zf_R02B04_NARVALI_fg_DOM01.nc')
da = DS.zf.values
zg_hr = da[:, rand_field]
# zg_hr = zg_hr[-91:] # Need the 91 earth-bound layers
# -
## Load vertically coarse-grained data
# Load clc profile
DS = xr.open_dataset('/pf/b/b309170/my_work/NARVAL/data_var_vertinterp/clc/int_var_clc_R02B04_NARVALII_2016072800_cloud_DOM01_0017.nc')
da = DS.clc.values
not_nan = ~np.isnan(da[0,:,rand_field])
cl_lr = da[0, not_nan, rand_field]
# Load zg profile
DS = xr.open_dataset('/pf/b/b309170/my_work/NARVAL/data_var_vertinterp/zg/zg_icon-a_capped.nc')
da = DS.zg.values
zg_lr = da[not_nan, rand_field]
# Increase the general font size
size_plot_elements = 16
matplotlib.rcParams['legend.fontsize'] = size_plot_elements
matplotlib.rcParams['axes.labelsize'] = size_plot_elements # For an axes xlabel and ylabel
matplotlib.rcParams['xtick.labelsize'] = size_plot_elements
matplotlib.rcParams['ytick.labelsize'] = size_plot_elements
# +
fig = plt.figure(figsize=(2,4))
# # Units in kilometers
# zg_hr = zg_hr/1000
# zg_lr = zg_lr/1000
# ax = fig.add_subplot(211, title='High-res vertical cloud cover profile', ylim=(0, np.max(zg_lr)), xlim=(-0.05,1),
# xlabel='Cloud Cover Fraction', ylabel='Mean height of a vertical layer in km')
ax = fig.add_subplot(111, ylim=(0, np.max(zg_lr[4:])), xlim=(-0.05,1), ylabel='z [km]', xticks=[0,0.5,1])
ax.plot(cl_hr/100, zg_hr)
ax.plot(cl_hr/100, zg_hr, 'b.')
plt.savefig('vertical_coarse-graining_narval_example_v2_1.pdf', bbox_inches='tight')
# +
fig = plt.figure(figsize=(2,4))
# ax_2 = fig.add_subplot(212, title='Low-res vertical cloud cover profile', ylim=(0, np.max(zg_lr)), xlim=(-0.05,1),
# xlabel='Cloud Cover Fraction', ylabel='Mean height of a vertical layer in km')
ax_2 = fig.add_subplot(111, ylim=(0, np.max(zg_lr[4:])), xlim=(-0.05,1),
xlabel='Cloud Fraction', ylabel='z [km]', xticks=[0,0.5,1])
ax_2.plot(cl_lr/100, zg_lr)
ax_2.plot(cl_lr/100, zg_lr, 'b.')
plt.savefig('vertical_coarse-graining_narval_example_v2_2.pdf', bbox_inches='tight')
# +
fig = plt.figure(figsize=(10,7))
ax = fig.add_subplot(121, title='High-res vertical cloud cover profile')
ax.plot(cl_hr/100, zg_hr)
ax.plot(cl_hr/100, zg_hr, 'b.')
ax_2 = fig.add_subplot(122, title='Low-res vertical cloud cover profile')
ax_2.plot(cl_lr/100, zg_lr)
ax_2.plot(cl_lr/100, zg_lr, 'b.')
| additional_content/analyzing_the_data/analyzing_narval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# ## Simple data preprocessing
# +
# From https://github.com/kulraghav/MLPractice/blob/master/Notebooks/spam_filter.ipynb
# when encoding is unknown
import chardet
with open("spam.tsv", 'rb') as f:
result = chardet.detect(f.read()) # or readline if the file is large
print(result['encoding'])
df_raw = pd.read_csv('spam.tsv', encoding=result['encoding'], delimiter='\t')
# -
df_raw.head()
df_raw.describe()
df_raw.drop_duplicates(subset="SMS", inplace=True)
from sklearn.model_selection import train_test_split
# +
X = df_raw['SMS']
y = df_raw['isSpam']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=10)
# -
# ## Create corpus and Vectorizer
# +
print("Shape of X is {}".format(X.shape))
print("Shape of X_train is {} and shape of y_train is {}".format(X_train.shape, y_train.shape))
print("Shape of X_test is {} and shape of y_test is {}".format(X_test.shape, y_test.shape))
train_corpus = list(X_train)
# +
"""
Featurizer: Train the featurizer on train data.
"""
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer(max_features=5000)
vectorizer.fit(train_corpus)
print("Number of features = {}".format(len(vectorizer.vocabulary_)))
print("Number of omitted words = {}".format(len(vectorizer.stop_words_)))
X_train_text_features = vectorizer.transform(list(X_train))
X_test_text_features = vectorizer.transform(list(X_test))
print("Shape of X_train_text_features is {}".format(X_train_text_features.shape))
print("Shape of X_test_text_features is {}".format(X_test_text_features.shape))
# -
#
# ## K nearest neighbours
#
# KNN classifers are the most simple classifiers. For each data point to classify we consider the k nearest neighbours of the training data and choose the majority class among them
#
# ## Logisitic Regression
#
# Logistic regression works exactly like linear regression but we apply the logit (sigmoid $f(x) = \frac{1}{1 + e^{-x}}$) function on the output from the linear regression to get a probability of each class
#
# ## Random Forests
#
# Random forests contain multiple decision trees. Each decision tree is trained on a subset of the features and then a weighted sum of their predicted is used for the final prediction.
# +
"""
Evaluation within training data: k-fold cross validation
- randomly partition the training data into k parts
- train on k-1 parts and evaluate on the remaining part
"""
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import RandomForestClassifier
from sklearn import neighbors
from sklearn.metrics import accuracy_score, confusion_matrix, f1_score
from sklearn.metrics import precision_recall_curve
import matplotlib.pyplot as plt
from sklearn.linear_model import LogisticRegression
lr_model = LogisticRegression()
rf_model = RandomForestClassifier(n_estimators=100, max_depth=50,random_state=0)
knn_model = neighbors.KNeighborsClassifier(n_neighbors=25)
models = [('Logistic Regression', lr_model),
('Random Forests', rf_model),
('K nearest neighbours', knn_model)]
for name, model in models:
print(name)
print("====================")
model.fit(X_train_text_features, y_train)
y_test_predicted = model.predict(X_test_text_features)
probs = [p[1] for p in model.predict_proba(X_test_text_features)]
print("The fraction of correctly classified samples is {}".format(accuracy_score(y_test, y_test_predicted)))
print("The number of correctly classified samples is {}".format(accuracy_score(y_test,
y_test_predicted, normalize=False)))
print("F1 score = {}".format(f1_score(y_test, y_test_predicted,pos_label='spam')))
print(pd.DataFrame(confusion_matrix(y_test_predicted, y_test),
index={'true ham', 'true spam'},
columns={'pred ham', 'pred spam'}))
print()
precision, recall, _ = precision_recall_curve(y_test, probs, pos_label='spam')
plt.plot(precision, recall, marker='.')
plt.show()
print("{} cv scores {}".format(name, cross_val_score(model, X=X_train_text_features, y=y_train, cv=5, n_jobs=4)))
print("====================")
# -
#
| spam/SMS Spam Filter .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#logistic-regression" data-toc-modified-id="logistic-regression-1"><span class="toc-item-num">1 </span>logistic regression</a></span><ul class="toc-item"><li><span><a href="#data" data-toc-modified-id="data-1.1"><span class="toc-item-num">1.1 </span>data</a></span></li><li><span><a href="#model" data-toc-modified-id="model-1.2"><span class="toc-item-num">1.2 </span>model</a></span></li><li><span><a href="#predict" data-toc-modified-id="predict-1.3"><span class="toc-item-num">1.3 </span>predict</a></span></li><li><span><a href="#contour" data-toc-modified-id="contour-1.4"><span class="toc-item-num">1.4 </span>contour</a></span></li></ul></li><li><span><a href="#polynomial-logistic-regression" data-toc-modified-id="polynomial-logistic-regression-2"><span class="toc-item-num">2 </span>polynomial logistic regression</a></span><ul class="toc-item"><li><span><a href="#data" data-toc-modified-id="data-2.1"><span class="toc-item-num">2.1 </span>data</a></span></li><li><span><a href="#model" data-toc-modified-id="model-2.2"><span class="toc-item-num">2.2 </span>model</a></span></li><li><span><a href="#predict" data-toc-modified-id="predict-2.3"><span class="toc-item-num">2.3 </span>predict</a></span></li><li><span><a href="#contour" data-toc-modified-id="contour-2.4"><span class="toc-item-num">2.4 </span>contour</a></span></li></ul></li></ul></div>
# -
from keras.layers import *
from keras.models import *
from keras.optimizers import *
from keras.callbacks import *
import keras
from keras import backend as K
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import itertools
# %matplotlib inline
# # logistic regression
# ## data
X = np.random.rand(1000, 2)
Y = np.where(X[:, 0] * X[:, 1] > 0.16, 1, 0)[:, np.newaxis]
plt.scatter(X[:, 0], X[:, 1], c=Y[:, 0])
# ## model
model_x = Input((2, ))
model_y = Dense(1, activation='sigmoid')(model_x)
model = Model(model_x, model_y)
model.compile(
loss='binary_crossentropy', optimizer='sgd', metrics=['accuracy'])
hist = model.fit(X, Y, batch_size=50, epochs=500, verbose=0)
print(model.evaluate(X, Y, verbose=0))
# ## predict
pred = model.predict(X) > 0.5
Y_pred = np.where(pred, 1, 0)
cond1 = np.logical_and(Y == 1, Y != Y_pred).flatten()
cond0 = np.logical_and(Y == 0, Y != Y_pred).flatten()
plt.scatter(X[:, 0], X[:, 1], c=Y[:, 0], marker='.')
plt.scatter(X[cond1][:, 0], X[cond1][:, 1], c='r', marker='x')
plt.scatter(X[cond0][:, 0], X[cond0][:, 1], c='g', marker='x')
# ## contour
px, py = np.meshgrid(np.linspace(0, 1), np.linspace(0, 1))
pxy = np.vstack((px.flatten(), py.flatten())).T
pz = model.predict(pxy).reshape(50, 50)
# pz = np.where(pz > 0.5, 1, 0)
plt.contourf(px, py, pz, 1, cmap=plt.cm.binary_r)
# plt.pcolormesh(px, py, pz, cmap=plt.cm.binary_r)
plt.colorbar()
plt.contour(px, py, pz, [0.5], colors='k')
plt.scatter(X[:, 0], X[:, 1], c=Y[:, 0], marker='.')
plt.tricontourf(X[:,0], X[:,1], Y_pred[:,0], 1, cmap=plt.cm.binary_r)
plt.colorbar()
plt.tricontour(X[:,0], X[:,1], Y_pred[:,0], [0.5], colors='k')
plt.scatter(X[:, 0], X[:, 1], c=Y[:, 0], marker='.')
# # polynomial logistic regression
# ## data
X = np.random.rand(1000, 2)
Y = np.where((X[:, 0]-0.5)**2/9 + (X[:, 1]-0.5)**2/6 < 0.01 + np.random.randn(1000)/300, 1, 0)[:, np.newaxis]
plt.scatter(X[:, 0], X[:, 1], c=Y[:, 0])
# ## model
def to_polynomial(x, y, n):
l = []
for i in range(n+1):
for j in range(i+1):
if i==0:
continue
l.append(x**(i-j) * y**j)
return l
model_x = Input((2, ))
model_y = Lambda(lambda x: K.map_fn(lambda y: K.stack(to_polynomial(y[0], y[1], 6)), x))(model_x)
model_y = Dense(1, activation='sigmoid')(model_y)
model = Model(model_x, model_y)
model.compile(loss='binary_crossentropy', optimizer=Adam(), metrics=['accuracy'])
hist = model.fit(X, Y, batch_size=50, epochs=500, verbose=0)
# ## predict
pred = model.predict(X)
Y_pred = np.where(pred>0.5, 1, 0)
cond0 = np.logical_and(Y==0, Y!=Y_pred).flatten()
cond1 = np.logical_and(Y==1, Y!=Y_pred).flatten()
plt.scatter(X[:,0], X[:,1], c=Y[:,0])
plt.scatter(X[cond0][:,0], X[cond0][:,1], c='g', marker='x')
plt.scatter(X[cond1][:,0], X[cond1][:,1], c='r', marker='x')
# ## contour
px, py = np.meshgrid(np.linspace(0, 1), np.linspace(0, 1))
pxy = np.vstack([px.flatten(), py.flatten()]).T
pz = model.predict(pxy).reshape(50, 50)
plt.contourf(px, py, pz, cmap=plt.cm.binary_r)
plt.colorbar()
plt.contour(px, py, pz, [0.5], colors='k')
plt.scatter(X[:,0], X[:,1], c=Y[:,0], marker='.')
| testzie/keras_logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="rDADvJplusjo" executionInfo={"status": "ok", "timestamp": 1625305175708, "user_tz": -300, "elapsed": 398, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="75a003b6-7062-4f1d-8a07-a16d9430dd06"
# %cd 'drive/My Drive/LearningCorner/FreeCodeCamp/Data Analysis with Python'
# + colab={"base_uri": "https://localhost:8080/"} id="YwIIsWYsvbE6" executionInfo={"status": "ok", "timestamp": 1625305235339, "user_tz": -300, "elapsed": 396, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="74a06ed8-2e53-4a5a-ca0f-e60a721817cd"
# !ls
# + colab={"base_uri": "https://localhost:8080/", "height": 306} id="zj8zXNWUuuSx" executionInfo={"status": "ok", "timestamp": 1625306090755, "user_tz": -300, "elapsed": 650, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="19de8a5f-89a8-4a69-f826-d6769630b657"
import pandas as pd
df = pd.read_csv('adult.data.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="UoFRbE--vg-Z" executionInfo={"status": "ok", "timestamp": 1625306097169, "user_tz": -300, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="ea553377-2ffe-4857-bb98-7c73bc943e45"
race = df.groupby('race').size()
race
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="Ak_7wceUvyTk" executionInfo={"status": "ok", "timestamp": 1625306358799, "user_tz": -300, "elapsed": 429, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="d8030f79-e2c3-4b24-ba85-99e773083445"
df[['age', 'sex']].groupby('sex').mean()
# + colab={"base_uri": "https://localhost:8080/"} id="VgYucwNgzGy1" executionInfo={"status": "ok", "timestamp": 1625306363184, "user_tz": -300, "elapsed": 398, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="590a0421-4ed4-4994-aaa7-5b03d5c5d4f6"
df[df['sex']=='Male']['age'].mean()
# + colab={"base_uri": "https://localhost:8080/"} id="shfQ-52AzvFL" executionInfo={"status": "ok", "timestamp": 1625310767049, "user_tz": -300, "elapsed": 384, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="f44083ed-3799-4b5d-9cb1-b0eec06669f9"
df[df['education']=='Bachelors'].shape[0]/df.shape[0]*100
# + colab={"base_uri": "https://localhost:8080/"} id="fpUOUfbdKwdg" executionInfo={"status": "ok", "timestamp": 1625312485713, "user_tz": -300, "elapsed": 385, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="191c224b-9789-4a00-c857-586a555a8e6d"
df.groupby('education').size()
# + id="BEy3FOChMp3e" executionInfo={"status": "ok", "timestamp": 1625313513936, "user_tz": -300, "elapsed": 419, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}}
h_e = df[df.education.isin(['Bachelors', 'Masters', 'Doctorate'])]
l_e = df[~df.education.isin(['Bachelors', 'Masters', 'Doctorate'])]
# + id="xqycMFZ7NoYf" executionInfo={"status": "ok", "timestamp": 1625313561072, "user_tz": -300, "elapsed": 16, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}}
h_e_r = h_e[h_e['salary']=='>50K']
l_e_r = l_e[l_e['salary']=='>50K']
# + colab={"base_uri": "https://localhost:8080/"} id="P5wg84GUIwdw" executionInfo={"status": "ok", "timestamp": 1625313588120, "user_tz": -300, "elapsed": 607, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="6ece3b61-f4d1-453f-81ff-cb124b78dd08"
print(h_e.shape[0])
print(h_e_r.shape[0])
print(h_e_r.shape[0]/h_e.shape[0])
# + colab={"base_uri": "https://localhost:8080/"} id="Tbcmt56IKjOf" executionInfo={"status": "ok", "timestamp": 1625313588734, "user_tz": -300, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="400ecf85-1423-4206-f941-26b5964b679d"
print(l_e.shape[0])
print(l_e_r.shape[0])
print(l_e_r.shape[0]/l_e.shape[0])
# + id="MXAx_2Wr75N_" executionInfo={"status": "ok", "timestamp": 1625313747780, "user_tz": -300, "elapsed": 396, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}}
min_hrs = df['hours-per-week'].min()
# + colab={"base_uri": "https://localhost:8080/"} id="xEOeN5Vu8QeS" executionInfo={"status": "ok", "timestamp": 1625314475091, "user_tz": -300, "elapsed": 384, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="910027bb-b2fe-4bb0-9053-c7035a658542"
min_workers = df.loc[(df['hours-per-week']==min_hrs)]
min_workers.shape[0]
# + colab={"base_uri": "https://localhost:8080/"} id="zQrSxqVMSsk4" executionInfo={"status": "ok", "timestamp": 1625314751841, "user_tz": -300, "elapsed": 403, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="7fa777d4-cea9-40ef-878a-ce160690a9ca"
min_workers[min_workers['salary']=='>50K'].shape[0]/min_workers.shape[0]*100
# + colab={"base_uri": "https://localhost:8080/"} id="EuRMaWDPXiY7" executionInfo={"status": "ok", "timestamp": 1625315933202, "user_tz": -300, "elapsed": 371, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="8192576d-08f5-42b3-b745-fab92e500438"
a = df[df['salary']=='>50K'].groupby('native-country').size()
b = df[['native-country','salary']].groupby('native-country').size()
print(type(a))
print(type(b))
# + colab={"base_uri": "https://localhost:8080/"} id="1rf2rK7fYiYo" executionInfo={"status": "ok", "timestamp": 1625316015204, "user_tz": -300, "elapsed": 531, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="2fb132a1-f19c-4949-a041-34d75f8b0671"
a/b
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="Z5aR8d4S9Z4L" executionInfo={"status": "ok", "timestamp": 1625315954864, "user_tz": -300, "elapsed": 401, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="903020e3-0686-472f-9c37-149727e365af"
(a/b).idxmax()
# + colab={"base_uri": "https://localhost:8080/"} id="g0hEWYlk-J4O" executionInfo={"status": "ok", "timestamp": 1625316044268, "user_tz": -300, "elapsed": 405, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="cc2490f7-ee27-4631-c20f-022938fab6d8"
(a/b).max()*100
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="O0Omc_vA-2jO" executionInfo={"status": "ok", "timestamp": 1625316129028, "user_tz": -300, "elapsed": 414, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07119957509138433407"}} outputId="83f16b86-c2c8-4085-82de-e8938d1f5c04"
df.loc[(df['native-country']=='India') & (df['salary']=='>50K')].groupby(df['occupation']).size().idxmax()
# + id="whqyHzH0CHQj"
| Courses/FreeCodeCamp/Data Analysis with Python/2 - Demographic Data Analyzer/Demographic Data Analyzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="aO-7t1Y7-hV4"
# # Import Libraries
# + id="8kH16rnZ7wt_"
from __future__ import print_function
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torch.optim as optim
from torchvision import datasets, transforms
# + id="0qQWwdPg55LW"
# HYPERPARAMS
num_epochs = 15
batch_size = 128
lr = 0.01
# CONSTANTS
NUM_CLASSES = 10
DATA_MEAN = (0.1307,)
DATA_STD = (0.3081,)
# cuda availability
cuda_available = torch.cuda.is_available()
device = "cuda" if cuda_available else "cpu"
# + [markdown] id="ky3f_Odl-7um"
# ## Data Transformations
#
# We first start with defining our data transformations. We need to think what our data is and how can we augment it to correct represent images which it might not see otherwise.
#
# + id="YtssFUKb-jqx"
# Train Phase transformations
train_transforms = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(DATA_MEAN, DATA_STD) # The mean and std have to be sequences (e.g., tuples), therefore you should add a comma after the values.
# Note the difference between (0.1307) and (0.1307,)
])
# Test Phase transformations
test_transforms = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(DATA_MEAN, DATA_STD)
])
# + [markdown] id="oQciFYo2B1mO"
# # Dataset and Creating Train/Test Split
# + id="_4A84rlfDA23" colab={"base_uri": "https://localhost:8080/", "height": 819, "referenced_widgets": ["f9d2807145344c00aa3d9e067a5a005d", "5606d141eead449bb6778538b79aae38", "<KEY>", "b07bc02d820d4f8b9df8ef1a704b31d3", "4eaf6e3d6e5d4a098780dbc05a910811", "<KEY>", "eaa0b4fbbac2421590cacf3e7dcc9e4a", "<KEY>", "391514f24fbe4395a359a96d97cb3322", "0b71ba45d2c94c44ac5514d09bd6ea1b", "2691289b85eb4c0094de700ec9df3c0a", "acb377128d7041748b3ece17c51847a9", "<KEY>", "ec25b8c5200b43ea9ed63ff16a99333b", "1654539e0b114647adb47e4dd8e2650a", "1b593ddef72945df92d55c623b9ca1b7", "<KEY>", "3fb6182866f74461834b1801ef3f5ad6", "18d0add0a94145b4ac5abd293798f014", "84f0e28d863144e7a7990e372fbccc91", "89d13cc97fad42b584424d41e574c7a5", "<KEY>", "9533efab83444e25aedbf1a41380fa7d", "0d3d4ab2e61c459098653940945ad6ac", "<KEY>", "54ca35db21af4421a4d7922047841f54", "d7848006149c455ea83918d5e5157a64", "4bc8bcdacef94010b6a8d1f116809ffa", "2308b66aee104f49ac36a23b70dfb691", "28a88a15e4ad4c5ca2b72648e658eb15", "<KEY>", "e991091d78ed450c9f29b2d88bf203f1"]} outputId="e03e5710-605d-4408-8b05-7da288fb16d4"
train = datasets.MNIST('./data', train=True, download=True, transform=train_transforms)
test = datasets.MNIST('./data', train=False, download=True, transform=test_transforms)
# + [markdown] id="qgldp_3-Dn0c"
# # Dataloader Arguments & Test/Train Dataloaders
#
# + id="C8OLDR79DrHG" colab={"base_uri": "https://localhost:8080/"} outputId="4d772f20-82c2-4ffc-a38f-ea194de2a5e2"
SEED = 1
# # CUDA?
cuda = torch.cuda.is_available()
print("CUDA Available?", cuda)
# For reproducibility
torch.manual_seed(SEED)
if cuda:
torch.cuda.manual_seed(SEED)
# dataloader arguments - something you'll fetch these from cmdprmt
dataloader_args = dict(shuffle=True, batch_size=batch_size, num_workers=4, pin_memory=True) if cuda else dict(shuffle=True, batch_size=64)
# train dataloader
train_loader = torch.utils.data.DataLoader(train, **dataloader_args)
# test dataloader
test_loader = torch.utils.data.DataLoader(test, **dataloader_args)
# + id="PkfjLQmx5qWx"
import matplotlib.pyplot as plt
import numpy as np
def display_image(image, title: str="Class label"):
"""
This function essentially takes in normalized tensors and the
Un-normalize them and display the image as output.
Args:
----
image: Image which we want to plot.
title: Label for that image.
"""
image = image.numpy().transpose((1, 2, 0)) # (C, H, W) --> (H, W, C)
# Convert mean and std to numpy array
mean = np.asarray(DATA_MEAN)
std = np.asarray(DATA_STD)
# unnormalize the image
image = std * image + mean
image = np.clip(image, 0, 1)
print(title)
fig = plt.figure() # Create a new figure
fig.set_figheight(15)
fig.set_figwidth(15)
ax = fig.add_subplot(111)
ax.axis("off") # Sqitch off the axis
ax.imshow(image)
# + colab={"base_uri": "https://localhost:8080/", "height": 921} id="apyHFcg051SS" outputId="fb3055a2-02fe-4fd9-f6bb-ccc9b998a3f1"
# Iteratre over and get 1 batch of training data
data, targets = next(iter(train_loader))
# make_grid takes all tensors(batch) and joins them into a single big tensor image (almost)
batch_grid = torchvision.utils.make_grid(data)
display_image(batch_grid, title=[str(cls.item()) for cls in targets])
# + [markdown] id="ubQL3H6RJL3h"
# # The model
# Let's start with the model we first saw
# + id="7FXQlB9kH1ov"
# ------99.2 /8.8k
import torch.nn.functional as F
dropout_value = 0.0
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# Input Block
self.convblock1 = nn.Sequential(
nn.Conv2d(in_channels=1, out_channels=8, kernel_size=(3, 3), padding=1, bias=False),
nn.ReLU(),
nn.BatchNorm2d(8),
nn.Dropout(dropout_value)
) # output_size = 28
# CONVOLUTION BLOCK 1
self.convblock2 = nn.Sequential(
nn.Conv2d(in_channels=8, out_channels=16, kernel_size=(3, 3), padding=1, bias=False),
nn.ReLU(),
nn.BatchNorm2d(16),
nn.Dropout(dropout_value)
) # output_size = 24
# TRANSITION BLOCK 1
self.convblock3 = nn.Sequential(
nn.Conv2d(in_channels=16, out_channels=4, kernel_size=(1, 1), padding=0, bias=False),
) # output_size = 24
self.pool1 = nn.MaxPool2d(2, 2) # output_size = 12
# CONVOLUTION BLOCK 2
self.convblock4 = nn.Sequential(
nn.Conv2d(in_channels=4, out_channels=8, kernel_size=(3, 3), padding=0, bias=False),
nn.ReLU(),
nn.BatchNorm2d(8),
nn.Dropout(dropout_value)
) # output_size = 10
self.convblock5 = nn.Sequential(
nn.Conv2d(in_channels=8, out_channels=16, kernel_size=(3, 3), padding=0, bias=False),
nn.ReLU(),
nn.BatchNorm2d(16),
nn.Dropout(dropout_value)
) # output_size = 8
self.convblock6 = nn.Sequential(
nn.Conv2d(in_channels=16, out_channels=32, kernel_size=(3, 3), padding=0, bias=False),
nn.ReLU(),
nn.BatchNorm2d(32),
nn.Dropout(dropout_value)
) # output_size = 6
# OUTPUT BLOCK
self.gap = nn.Sequential(
nn.AvgPool2d(kernel_size=6)
) # output_size = 1
self.convblock8 = nn.Sequential(
nn.Conv2d(in_channels=32, out_channels=16, kernel_size=(1, 1), padding=0, bias=False),
)
self.classifier = nn.Linear(16, 10)
self.dropout = nn.Dropout(dropout_value)
def forward(self, x):
x = self.convblock1(x)
x = self.convblock2(x)
x = self.convblock3(x)
x = self.pool1(x)
x = self.convblock4(x)
x = self.convblock5(x)
x = self.convblock6(x)
x = self.gap(x)
x = self.convblock8(x)
x = x.view(-1, 16)
x = self.classifier(x)
return F.log_softmax(x, dim=-1)
# + [markdown] id="M3-vp8X9LCWo"
# # Model Params
# Can't emphasize on how important viewing Model Summary is.
# Unfortunately, there is no in-built model visualizer, so we have to take external help
# + id="5skB97zIJQQe" colab={"base_uri": "https://localhost:8080/"} outputId="d746ea34-d7ae-48dd-f759-08b5f222c690"
# # !pip install torchsummary
from torchsummary import summary
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
print(device)
model = Net().to(device)
summary(model, input_size=(1, 28, 28))
# + [markdown] id="1__x_SbrL7z3"
# # Training and Testing
#
# Looking at logs can be boring, so we'll introduce **tqdm** progressbar to get cooler logs.
#
# Let's write train and test functions
# + id="fbkF2nN_LYIb"
from tqdm import tqdm
train_losses = []
test_losses = []
train_acc = []
test_acc = []
def train(model, device, train_loader, optimizer, epoch):
model.train()
pbar = tqdm(train_loader)
correct = 0
processed = 0
for batch_idx, (data, target) in enumerate(pbar):
# get samples
data, target = data.to(device), target.to(device)
# Init
optimizer.zero_grad()
# In PyTorch, we need to set the gradients to zero before starting to do backpropragation because PyTorch accumulates the gradients on subsequent backward passes.
# Because of this, when you start your training loop, ideally you should zero out the gradients so that you do the parameter update correctly.
# Predict
y_pred = model(data)
# Calculate loss
loss = F.nll_loss(y_pred, target)
train_losses.append(loss)
# Backpropagation
loss.backward()
optimizer.step()
# Update pbar-tqdm
pred = y_pred.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
processed += len(data)
pbar.set_description(desc= f'Loss={loss.item()} Batch_id={batch_idx} Accuracy={100*correct/processed:0.2f}')
train_acc.append(100*correct/processed)
def test(model, device, test_loader):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
test_loss += F.nll_loss(output, target, reduction='sum').item() # sum up batch loss
pred = output.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
test_losses.append(test_loss)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.2f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
test_acc.append(100. * correct / len(test_loader.dataset))
# + colab={"base_uri": "https://localhost:8080/"} id="9T4BQ9C90YU5" outputId="294de1b0-5254-4a3c-fd67-c59ba5858a5d"
from torch.optim.lr_scheduler import StepLR
model = Net().to(device)
# optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
optimizer = optim.Adam(model.parameters(), lr=0.05)
scheduler = StepLR(optimizer, step_size=5, gamma=0.1)
EPOCHS = 15
for epoch in range(EPOCHS):
print("EPOCH:", epoch)
train(model, device, train_loader, optimizer, epoch)
scheduler.step()
test(model, device, test_loader)
# + [markdown] id="drokW8wWODKq"
# # Let's plot our train and test metrics
#
# + id="87RaqGSEOWDe" colab={"base_uri": "https://localhost:8080/", "height": 625} outputId="32be73bd-3c0f-4008-8872-78b1d266df0e"
# %matplotlib inline
import matplotlib.pyplot as plt
fig, axs = plt.subplots(2,2,figsize=(15,10))
axs[0, 0].plot(train_losses)
axs[0, 0].set_title("Training Loss")
axs[1, 0].plot(train_acc[4000:])
axs[1, 0].set_title("Training Accuracy")
axs[0, 1].plot(test_losses)
axs[0, 1].set_title("Test Loss")
axs[1, 1].plot(test_acc)
axs[1, 1].set_title("Test Accuracy")
# + id="wjO3RK9UEnvF"
| Assignment_5/Notebooks/MNIST_regularization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: Ang Pangunahing Mga Kasangkapan ng Pribadong Deep Learning
#
# Maligayang pagdating sa panimulang tutorial ng PySyft para sa pagpapanatili ng privacy, desentralisadong Deep Learning. Ang seryeng ito ng mga notebook ay isang gabay na sunud-sunod na gabay para sa iyo upang makilala ang mga bagong tool at pamamaraan na kinakailangan para sa paggawa ng malalim na pag-aaral(Deep Learning) sa lihim / pribadong data / modelo nang hindi nakakasalamuha ang mga ito sa ilalim ng isang awtoridad
#
# **Saklaw:** Tandaan na hindi lamang namin pinag-uusapan ang tungkol sa kung paano ma-desentralisado / i-encrypt ang data, ngunit tutugunan namin kung paano magagamit ang PySyft upang matukoy ang buong ecosystem sa paligid ng data, kabilang ang mga Databases kung saan ang data ay naka-imbak at queried, at ang mga modelo ng neural na ginagamit upang kunin ang impormasyon mula sa data. Tulad ng mga bagong extension sa PySyft ay nilikha, ang mga notebook na ito ay idinugtong sa mga bagong tutorial upang maipaliwanag ang bagong ginagawa ng PySyft.
#
# Authors:
# - <NAME> - Twitter: [@iamtrask](https://twitter.com/iamtrask)
#
# ## Outline:
#
# - Unang Bahagi: Ang Pangunahing Kasangkapan ng Pri<NAME> Pag-aaral(Deep Learning)
#
#
# ## Why Take This Tutorial?
#
# **1) A Competitive Career Advantage** - For the past 20 years, the digital revolution has made data more and more accessible in ever larger quantities as analog processes have become digitized. However, with new regulation such as [GDPR](https://eugdpr.org/), enterprises are under pressure to have less freedom with how they use - and more importantly how they analyze - personal information. **Bottom Line:** Data Scientists aren't going to have access to as much data with "old school" tools, but by learning the tools of Private Deep Learning, YOU can be ahead of this curve and have a competitive advantage in your career.
#
# **2) Entrepreneurial Opportunities** - There are a whole host of problems in society that Deep Learning can solve, but many of the most important haven't been explored because it would require access to incredibly sensitive information about people (consider using Deep Learning to help people with mental or relationship issues!). Thus, learning Private Deep Learning unlocks a whole host of new startup opportunities for you which were not previously available to others without these toolsets.
#
# **3) Social Good** - Deep Learning can be used to solve a wide variety of problems in the real world, but Deep Learning on *personal information* is Deep Learning about people, *for people*. Learning how to do Deep Learning on data you don't own represents more than a career or entrepreneurial opportunity, it is the opportunity to help solve some of the most personal and important problems in people's lives - and to do it at scale.
#
# ## How do I get extra credit?
#
# - Star PySyft on GitHub! - [https://github.com/OpenMined/PySyft](https://github.com/OpenMined/PySyft)
# - Make a Youtube video teaching this notebook!
#
#
# ... ok ... let's do this!
# # Part -1: Prerequisites
#
# - Know PyTorch - if not then take the http://fast.ai course and come back
# - Read the PySyft Framework Paper https://arxiv.org/pdf/1811.04017.pdf! This will give you a thorough background on how PySyft is constructed which will help things make more sense.
# # Part 0: Setup
#
# To begin, you'll need to make sure you have the right things installed. To do so, head on over to PySyft's readme and follow the setup instructions. TLDR for most folks is.
#
# - Install Python 3.6 or higher
# - Install PyTorch 1.3
# - Clone PySyft (git clone https://github.com/OpenMined/PySyft.git)
# - cd PySyft
# - pip install -r pip-dep/requirements.txt
# - pip install -r pip-dep/requirements_udacity.txt
# - python setup.py install udacity
# - python setup.py test
#
# If any part of this doesn't work for you (or any of the tests fail) - first check the [README](https://github.com/OpenMined/PySyft.git) for installation help and then open a GitHub Issue or ping the #beginner channel in our slack! [slack.openmined.org](http://slack.openmined.org/)
# +
# Run this cell to see if things work
import sys
import torch
from torch.nn import Parameter
import torch.nn as nn
import torch.nn.functional as F
import syft as sy
hook = sy.TorchHook(torch)
torch.tensor([1,2,3,4,5])
# -
# If this cell executed, then you're off to the races! Let's do this!
# # Part 1: The Basic Tools of Private, Decentralized Data Science
#
# So - the first question you may be wondering is - How in the world do we train a model on data we don't have access to?
#
# Well, the answer is surprisingly simple. If you're used to working in PyTorch, then you're used to working with torch.Tensor objects like these!
x = torch.tensor([1,2,3,4,5])
y = x + x
print(y)
# Obviously, using these super fancy (and powerful!) tensors is important, but also requires you to have the data on your local machine. This is where our journey begins.
#
# # Section 1.1 - Sending Tensors to Bob's Machine
#
# Whereas normally we would perform data science / deep learning on the machine which holds the data, now we want to perform this kind of computation on some **other** machine. More specifically, we can no longer simply assume that the data is on our local machine.
#
# Thus, instead of using Torch tensors, we're now going to work with **pointers** to tensors. Let me show you what I mean. First, let's create a "pretend" machine owned by a "pretend" person - we'll call him Bob.
bob = sy.VirtualWorker(hook, id="bob")
# Let's say Bob's machine is on another planet - perhaps on Mars! But, at the moment the machine is empty. Let's create some data so that we can send it to Bob and learn about pointers!
x = torch.tensor([1,2,3,4,5])
y = torch.tensor([1,1,1,1,1])
# And now - let's send our tensors to Bob!!
x_ptr = x.send(bob)
y_ptr = y.send(bob)
x_ptr
# BOOM! Now Bob has two tensors! Don't believe me? Have a look for yourself!
bob._objects
z = x_ptr + x_ptr
z
bob._objects
# Now notice something. When we called `x.send(bob)` it returned a new object that we called `x_ptr`. This is our first *pointer* to a tensor. Pointers to tensors do NOT actually hold data themselves. Instead, they simply contain metadata about a tensor (with data) stored on another machine. The purpose of these tensors is to give us an intuitive API to tell the other machine to compute functions using this tensor. Let's take a look at the metadata that pointers contain.
x_ptr
# Check out that metadata!
#
# There are two main attributes specific to pointers:
#
# - `x_ptr.location : bob`, the location, a reference to the location that the pointer is pointing to
# - `x_ptr.id_at_location : <random integer>`, the id where the tensor is stored at location
#
# They are printed in the format `<id_at_location>@<location>`
#
# There are also other more generic attributes:
# - `x_ptr.id : <random integer>`, the id of our pointer tensor, it was allocated randomly
# - `x_ptr.owner : "me"`, the worker which owns the pointer tensor, here it's the local worker, named "me"
#
x_ptr.location
bob
bob == x_ptr.location
x_ptr.id_at_location
x_ptr.owner
# You may wonder why the local worker which owns the pointer is also a VirtualWorker, although we didn't create it.
# Fun fact, just like we had a VirtualWorker object for Bob, we (by default) always have one for us as well. This worker is automatically created when we called `hook = sy.TorchHook()` and so you don't usually have to create it yourself.
me = sy.local_worker
me
me == x_ptr.owner
# And finally, just like we can call .send() on a tensor, we can call .get() on a pointer to a tensor to get it back!!!
x_ptr
x_ptr.get()
y_ptr
y_ptr.get()
z.get()
bob._objects
# And as you can see... Bob no longer has the tensors anymore!!! They've moved back to our machine!
# # Section 1.2 - Using Tensor Pointers
#
# So, sending and receiving tensors from Bob is great, but this is hardly Deep Learning! We want to be able to perform tensor _operations_ on remote tensors. Fortunately, tensor pointers make this quite easy! You can just use pointers like you would normal tensors!
x = torch.tensor([1,2,3,4,5]).send(bob)
y = torch.tensor([1,1,1,1,1]).send(bob)
z = x + y
z
# And voilà!
#
# Behind the scenes, something very powerful happened. Instead of x and y computing an addition locally, a command was serialized and sent to Bob, who performed the computation, created a tensor z, and then returned the pointer to z back to us!
#
# If we call .get() on the pointer, we will then receive the result back to our machine!
z.get()
# ### Torch Functions
#
# This API has been extended to all of Torch's operations!!!
x
y
z = torch.add(x,y)
z
z.get()
# ### Variables (including backpropagation!)
x = torch.tensor([1,2,3,4,5.], requires_grad=True).send(bob)
y = torch.tensor([1,1,1,1,1.], requires_grad=True).send(bob)
z = (x + y).sum()
z.backward()
x = x.get()
x
x.grad
# So as you can see, the API is really quite flexible and capable of performing nearly any operation you would normally perform in Torch on *remote data*. This lays the groundwork for our more advanced privacy preserving protocols such as Federated Learning, Secure Multi-Party Computation, and Differential Privacy !
# # Congratulations!!! - Time to Join the Community!
#
# Congratulations on completing this notebook tutorial! If you enjoyed this and would like to join the movement toward privacy preserving, decentralized ownership of AI and the AI supply chain (data), you can do so in the following ways!
#
# ### Star PySyft on GitHub
#
# The easiest way to help our community is just by starring the GitHub repos! This helps raise awareness of the cool tools we're building.
#
# - [Star PySyft](https://github.com/OpenMined/PySyft)
#
# ### Join our Slack!
#
# The best way to keep up to date on the latest advancements is to join our community! You can do so by filling out the form at [http://slack.openmined.org](http://slack.openmined.org)
#
# ### Join a Code Project!
#
# The best way to contribute to our community is to become a code contributor! At any time you can go to PySyft GitHub Issues page and filter for "Projects". This will show you all the top level Tickets giving an overview of what projects you can join! If you don't want to join a project, but you would like to do a bit of coding, you can also look for more "one off" mini-projects by searching for GitHub issues marked "good first issue".
#
# - [PySyft Projects](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3AProject)
# - [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
#
# ### Donate
#
# If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
#
# [OpenMined's Open Collective Page](https://opencollective.com/openmined)
| examples/tutorials/Part 01 - The Basic Tools of Private Deep Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["parameters"]
epochs = 15
# -
# # Tutorial 2 - MultiLayer Split Neural Network
#
# <b>Recap:</b> The previous tutorial looked at building a basic SplitNN, where an NN was split into two segments on two seperate hosts. However, there is a lot more that we can do with this technique. An NN can be split any number of times without affecting the accuracy of the model.
#
# <b>Description: </b>Here we define a class which can procees a SplitNN of any number of layers. All it needs is a list of distributed models and their optimizers.
#
#
#
# <img src="images/MultiSplitNN.png" width="20%">
#
#
# In this tutorial, we demonstrate the SplitNN class with a 3 segment distribution [[1](https://arxiv.org/abs/1812.00564)]. This time;
#
# - <b>Alice</b>
# - Has Model Segment 1
# - Has the handwritten images
# - <b>Bob</b>
# - Has model Segment 2
# - <b>Claire</b>
# - Has Model Segment 3
# - Has the image labels
#
# We use the exact same model as we used in the previous tutorial, only this time we are splitting over 3 hosts, not two. However, we see the same loss being reported as there is <b>no reduction in accuracy</b> when training in this way. While we only use 3 models this can be done for any arbitrary number of models.
#
# Author:
# - <NAME> - Twitter: [@AJH4LL](https://twitter.com/AJH4LL) · GitHub: [@H4LL](https://github.com/H4LL)
#
import numpy as np
import torch
import torchvision
import matplotlib.pyplot as plt
from time import time
from torchvision import datasets, transforms
from torch import nn, optim
import syft as sy
import time
hook = sy.TorchHook(torch)
class SplitNN(torch.nn.Module):
def __init__(self, models, optimizers):
self.models = models
self.optimizers = optimizers
self.outputs = [None]*len(self.models)
self.inputs = [None]*len(self.models)
super().__init__()
def forward(self, x):
self.inputs[0] = x
self.outputs[0] = self.models[0](self.inputs[0])
for i in range(1, len(self.models)):
self.inputs[i] = self.outputs[i-1].detach().requires_grad_()
if self.outputs[i-1].location != self.models[i].location:
self.inputs[i] = self.inputs[i].move(self.models[i].location).requires_grad_()
self.outputs[i] = self.models[i](self.inputs[i])
return self.outputs[-1]
def backward(self):
for i in range(len(self.models)-2, -1, -1):
grad_in = self.inputs[i+1].grad.copy()
if self.outputs[i].location != self.inputs[i+1].location:
grad_in = grad_in.move(self.outputs[i].location)
self.outputs[i].backward(grad_in)
def zero_grads(self):
for opt in self.optimizers:
opt.zero_grad()
def step(self):
for opt in self.optimizers:
opt.step()
def train(self):
for model in self.models:
model.train()
def eval(self):
for model in self.models:
model.eval()
@property
def location(self):
return self.models[0].location if self.models and len(self.models) else None
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
trainset = datasets.MNIST('mnist', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# +
torch.manual_seed(0)
# Define our model segments
input_size = 784
hidden_sizes = [128, 640]
output_size = 10
models = [
nn.Sequential(
nn.Linear(input_size, hidden_sizes[0]),
nn.ReLU(),
),
nn.Sequential(
nn.Linear(hidden_sizes[0], hidden_sizes[1]),
nn.ReLU(),
),
nn.Sequential(
nn.Linear(hidden_sizes[1], output_size),
nn.LogSoftmax(dim=1)
)
]
# Create optimisers for each segment and link to them
optimizers = [
optim.SGD(model.parameters(), lr=0.03,)
for model in models
]
# create some workers
alice = sy.VirtualWorker(hook, id="alice")
bob = sy.VirtualWorker(hook, id="bob")
claire = sy.VirtualWorker(hook, id="claire")
# Send Model Segments to model locations
model_locations = [alice, bob, claire]
for model, location in zip(models, model_locations):
model.send(location)
#Instantiate a SpliNN class with our distributed segments and their respective optimizers
splitNN = SplitNN(models, optimizers)
# -
def train(x, target, splitNN):
#1) Zero our grads
splitNN.zero_grads()
#2) Make a prediction
pred = splitNN.forward(x)
#3) Figure out how much we missed by
criterion = nn.NLLLoss()
loss = criterion(pred, target)
#4) Backprop the loss on the end layer
loss.backward()
#5) Feed Gradients backward through the nework
splitNN.backward()
#6) Change the weights
splitNN.step()
return loss
for i in range(epochs):
running_loss = 0
splitNN.train()
for images, labels in trainloader:
images = images.send(models[0].location)
images = images.view(images.shape[0], -1)
labels = labels.send(models[-1].location)
loss = train(images, labels, splitNN)
running_loss += loss.get()
else:
print("Epoch {} - Training loss: {}".format(i, running_loss/len(trainloader)))
def test(model, dataloader, dataset_name):
model.eval()
correct = 0
with torch.no_grad():
for data, target in dataloader:
data = data.view(data.shape[0], -1).send(model.location)
output = model(data).get()
pred = output.data.max(1, keepdim=True)[1]
correct += pred.eq(target.data.view_as(pred)).sum()
print("{}: Accuracy {}/{} ({:.0f}%)".format(dataset_name,
correct,
len(dataloader.dataset),
100. * correct / len(dataloader.dataset)))
testset = datasets.MNIST('mnist', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=False)
test(splitNN, testloader, "Test set")
test(splitNN, trainloader, "Train set")
| examples/tutorials/advanced/Split Neural Network/Tutorial 2 - MultiLayer Split Neural Network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''base'': conda)'
# name: python37464bitbaseconda0633cb87348e4e1f83d9c32f4c55bf11
# ---
# ## Superstore data analysis project using two types of Dataframes:
# 1. Pandas Dataframes.
# 2. Koalas Dataframes.
from pyspark.sql import SparkSession
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from databricks import koalas as ks
spark = SparkSession.builder.appName('superstore').getOrCreate()
df = spark.read.csv('../datasets/superstore_dataset.csv',inferSchema=True,header=True, encoding = "ISO-8859-1")
data = pd.read_csv('../datasets/superstore_dataset.csv', encoding = "ISO-8859-1")
kdf = ks.from_pandas(data)
kdf.head(3)
data.head(5)
kdf.shape
data.shape
kdf.columns
data.columns
# ****
# #### 1. Who are the top-20 most profitable customers. Show them through plots.
# ****
# +
result = data.groupby(["Customer Name"])['Profit'].aggregate(np.sum).reset_index().sort_values('Profit',ascending = False).head(20)
sns.barplot(x='Customer Name',y='Profit',data=result)
# -
fig = plt.figure(figsize = (8,8))
ax1 = fig.add_subplot(111)
sns.barplot(x = "Customer Name",y= "Profit",
data=result,
ax = ax1)
ax1.set_ylabel("Profit", fontname="Arial", fontsize=12)
ax1.set_title("Top 20 Customers", fontname='Comic Sans MS', fontsize=18)
# Set the font name for axis tick labels to be Comic Sans
for tick in ax1.get_xticklabels():
tick.set_fontname("Comic Sans MS")
tick.set_fontsize(12)
for tick in ax1.get_yticklabels():
tick.set_fontname("Comic Sans MS")
tick.set_fontsize(12)
# Rotate the labels as the Customer names overwrites on top of each other
ax1.set_xticklabels(ax1.get_xticklabels(), rotation = 45)
plt.show()
# #### Implement in koalas:
#
# result = kdf.groupby(["Customer Name"])['Profit'].sum().reset_index().sort_values('Profit',ascending = False).head(20)
#
# sns.barplot(x='Customer Name',y='Profit',data=result.to_pandas())
# ****
# #### 2. What is the distribution of our customer Segment
# ****
#
kdf['Segment'].value_counts()
sns.countplot("Segment", data = kdf.to_pandas())
# #### Implementation in koalas and pandas df is same in this case.
# ****
# #### 3. Who are our top-20 oldest customers
# ****
data['Order Date']=pd.to_datetime(data['Order Date'])
oldest = pd.DataFrame({'Counts':data.groupby(["Order Date","Customer Name"]).size()}).reset_index()
oldest.head(20)
# #### Koalas implementation here
kdf['Order Date']=ks.to_datetime(kdf['Order Date'])
# Using pandas data to use groupby
old_date = ks.DataFrame({'Counts':data.groupby(["Order Date","Customer Name"]).size()}).reset_index()
old_date.head(10)
# ****
# #### 4. Which customers have visited this store just once
# ****
customer_visit = pd.DataFrame({'Counts': data.groupby('Customer Name').size()}).reset_index()
customer_visit[customer_visit['Counts']==1]
# #### Koalas implementation:
# +
customer_visit_k = kdf.groupby('Customer Name').size().reset_index()
customer_visit_k[customer_visit_k['count']==1]
# -
# #### Since it returns an empty data frame, there are no customers that have visited this store just once
# #### 6. What is the distribution of customers Market wise?
customer_market = pd.DataFrame({'counts': data.groupby(['Market','Customer Name']).size()}).reset_index()
customer_market.shape
sns.barplot(x='Market',y='counts',data=customer_market)
# #### Koalas implementation
customer_market_k = kdf.groupby(['Market','Customer Name']).size().reset_index()
customer_market_k.shape
sns.barplot(x='Market',y='count',data=customer_market_k.to_pandas())
#7. What is the distribution of customers Market wise and Region wise
customer_market_region = pd.DataFrame({'Count':data.groupby(['Market','Region','Customer Name']).size()}).reset_index()
sns.barplot(x='Market',hue='Region',y='Count',data=customer_market_region)
# #### Koalas implementation will be same as above
# #### for APAC, the largest # of customers are basd out of Oceania, followed by Southeast Asia
# #### for US, the largest # of customers are based out of Western Region followed by East
# #### 8.Distribution of Customers by Country & State - top 15
customer_country = pd.DataFrame({'Count':data.groupby(['Country','State']).size()}).reset_index().sort_values('Count',ascending = False).head(15)
customer_country
sns.barplot(x='Country',y='Count',hue='State',data=customer_country.sort_values('Country'))
# #### Koalas implementation will be same as above
# #### Top 20 Cities by Sales Volume
sales_city = data.groupby('City')['Quantity'].aggregate(np.sum).reset_index().sort_values('Quantity',ascending=False).head(20)
sales_city
ax = plt.figure().add_subplot()
sns.barplot(x='City',y='Quantity',data=sales_city,ax=ax)
ax.set_xticklabels(ax.get_xticklabels(),rotation=90)
plt.show()
# #### Top 10 products
sales_product = data.groupby(['Product Name'])['Quantity'].aggregate(np.sum).reset_index().sort_values('Quantity', ascending=False).head(10)
sales_product
sns.barplot(x='Quantity',y='Product Name',data=sales_product)
# #### Add Handyspark implementation here
# #### Top selling products by country (in US)
top_selling = data.groupby(['Product Name','Country'])['Quantity'].sum().reset_index().sort_values('Quantity',ascending=False)
# Only keeping US data
top_selling = top_selling[top_selling['Country']=='United States'].sort_values('Quantity',ascending=False).head(10)
sns.barplot(x='Quantity',y='Product Name',data=top_selling)
# #### Add HandySpark implementation here
# #### Sales of product category and subcategory
sales_category = data.groupby(['Category','Sub-Category'])['Quantity'].sum().reset_index().sort_values('Quantity',ascending=False)
sales_category
sns.barplot(x='Quantity',y='Category',hue='Sub-Category',data=sales_category)
# #### Koalas implementation:
sales_category_k = kdf.groupby(['Category','Sub-Category'])['Quantity'].sum().reset_index().sort_values('Quantity',ascending=False)
sales_category_k.head(2)
sns.barplot(x='Quantity',y='Category',hue='Sub-Category',data=sales_category_k.to_pandas())
# ### We can see that office Supplies, Binders and Storage are the best sellers.
| Code/Day80-Superstore_data_project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %%time
#raw 데이터 파일 읽기
import numpy as np
import pandas as pd
import time
data_set = []
with open("sise2local.ALL_CMDT.0213", 'rt', encoding='ISO-8859-1') as f:
data=f.readlines()
for line in data:
data_set.append(np.array(line.split(',')))
# -
# #### wall time : 2.78 s
# +
#종목 코드 추출 함수인데 이번엔 종목이 1개라 사용하지 않았습니다.
def productID_Code_set(data):
product_ID = [ [array[0][0:17], array[0][(17+18):(17+30)], array[0][(17+36):(17+47)]] for array in data_set if
((array[0][17:(17+5)]=='A0016') & (array[0][(17+408):(17+411)] == 'BM3') & (array[0][(17+454):(17+457)] == '001'))]
# row[0][17:] for row in data_set]
return [ code[0] for code in product_ID] , [code[1] for code in product_ID], [code[2] for code in product_ID]
# -
time_list , code_list, id_list = productID_Code_set(data_set)
print( '시간', time_list)
print( '종목코드', code_list)
print( '상품ID', id_list)
set(code_list), set(id_list)
#KRDRVFUBM3 : KRX 파생 선물 신3년국채
# +
#필요한 데이터 추출
def ex_data(data):
#Product_ID = set(productIDset(data)set)
return_array = [ [array[0][1:16],array[0][17:22], array[0][(17+23):(17+31)],array[0][(17+31):(17+37)], array[0][(17+228):(17+234)], array[0][(17+234):(17+240)], array[0][(17+144):(17+152)], array[0][(17+152):(17+158)], array[0][(17+135)], \
array[0][(17+218):(17+225)], array[0][(17+136):(17+143)] ]
for array in data
if (((array[0][(5+17):(17+17)] == 'KR4165N30007') or (array[0][(5+17):(17+17)] == 'KR4465N3N6S2'))& ((array[0][17:(17+5)] == 'A3016') or
(array[0][17:(17+5)] == 'G7016') ))]
return return_array
# -
ex_data_set=ex_data(data_set)
df_ex=pd.DataFrame(ex_data_set[:])
column_series = pd.Series(['시간','구분','현재가','체결수량','매도1호가','매도1단계잔량','매수1호가','매수1단계잔량','최종매도매수','총잔량매도','총잔량매수'])
df_ex.columns = column_series
#df_ex['시간'] = df_ex['시간'][0:2]+ df_ex['시간'][3:5] + df_ex['시간'][6:8] + df_ex['시간'][9:]
df_ex['시간'].loc[0]
def ex_data_b6016(data):
return_array = [ [array[0][1:16], array[0][17:22], array[0][(17+114):(17+122)],array[0][(17+122):(17+128)], array[0][(17+22):(17+40)],array[0][(17+40):(17+46)], array[0][(17+247)], array[0][(17+247):(17+255)] ]
for array in data
if (((array[0][(5+17):(17+17)] == 'KR4165N30007') or (array[0][(5+17):(17+17)] == 'KR4465N3N6S2'))&
( array[0][17:(17+5)] == 'B6016' ))]
return return_array
def ex_data_h1016(data):
return_array = [ [array[0][1:16],array[0][17:22], array[0][(17+5):(17+13)],array[0][(17+13):(17+19)], array[0][(17+19):(17+21)], array[0][(17+21):(17+32)], array[0][(17+32):(17+36)], \
array[0][(17+36):(17+45)], array[0][(17+54):(17+72)], array[0][(17+45):(17+54)], array[0][(17+72):(17+90)] ]
for array in data
if ((array[0][(17+21):(17+32)] == 'KRDRVFUBM3 ' )&
( array[0][17:(17+5)] == 'H1016' ))]
return return_array
b6 1~5 매수1단계 가격 : 8:40 , 매수1단계잔량 : 6: 46, 매도1단계 가격 : 8: 122 , 매도1단계잔량 : 6:128 예상체결 가격 부호 : 247 예상체결 가격 : 8:255
h1 생성 일자 8:13, 생성시각 6:19, 데이터구분 2:21, 상품ID 11:32, 투자자유형 4:36,
매수약정 수량 9:45, 매수약정대금 18:72, 매도약정수량 9:54, 매도약정대금 18:90
# +
ex_data_set = ex_data(data_set)
ex_data_h1016 = ex_data_h1016(data_set)
ex_data_b6016 = ex_data_b6016(data_set)
df_ex=pd.DataFrame(ex_data_set[:])
df_h1016 = pd.DataFrame(ex_data_h1016[:])
df_b6016 = pd.DataFrame(ex_data_b6016[:])
# -
def dataframe(data):
# 데이터 추출, dataframe 만들기
ex_data_set = ex_data(data)
ex_h1016 = ex_data_h1016(data)
ex_b6016 = ex_data_b6016(data)
df_ex=pd.DataFrame(ex_data_set[:])
df_h1016 = pd.DataFrame(ex_h1016[:])
df_b6016 = pd.DataFrame(ex_b6016[:])
#칼럼 설정
column_series = pd.Series(['시간','구분','현재가','체결수량','매도1호가','매도1단계잔량','매수1호가','매수1단계잔량','최종매도매수','총잔량매도','총잔량매수'])
df_ex.columns = column_series
b6016_column = pd.Series(['시간','구분','매도1단계가격','매도1단계잔량','매수1단계가격', '매수1단계잔량','예상체결가격부호', '예상체결가격' ])
df_b6016.columns = b6016_column
h1016_column = pd.Series(['시간','구분','생성일자', '생성시각','데이터구분','상품ID','투자자유형','매수약정수량','매수약정대금','매도약정수량','매도약정대금'])
df_h1016.columns = h1016_column
#시간변경 속성
for i in range(len(df_ex)):
df_ex['시간'].loc[i] = df_ex['시간'].loc[i][0:2]+ df_ex['시간'].loc[i][3:5] + df_ex['시간'].loc[i][6:8] + df_ex['시간'].loc[i][9:]
for j in range(len(df_h1016)):
df_h1016['시간'].loc[j] = df_h1016['시간'].loc[j][0:2]+ df_h1016['시간'].loc[j][3:5] + df_h1016['시간'].loc[j][6:8] + df_h1016['시간'].loc[j][9:]
for k in range(len(df_b6016)):
df_b6016['시간'].loc[k] = df_b6016['시간'].loc[k][0:2]+ df_b6016['시간'].loc[k][3:5] + df_b6016['시간'].loc[k][6:8] + df_b6016['시간'].loc[k][9:]
df_ex.apply(pd.to_numeric, errors ='ignore')
df_h1016.apply(pd.to_numeric, errors ='ignore')
df_b6016.apply(pd.to_numeric, errors ='ignore')
return df_ex, df_h1016, df_b6016
g7, h1, b6 = dataframe(data_set)
df_h1016["매수약정수량"] = df_h1016['매수약정수량'].apply(lambda x: int(x))
df_h1016["매도약정수량"] = df_h1016['매도약정수량'].apply(lambda x: int(x))
# +
type_df = df_h1016.groupby("투자자유형").sum()['매수약정수량']
type_df2 = df_h1016.groupby("투자자유형").sum()['매도약정수량']
print(type_df, type_df2, type_df + type_df2)
# -
top3 9999, 1000, 9000
df_h1016
buy_df = pd.Series(0)
sell_df = pd.Series(0)
buy_df = pd.concat( [buy_df, df_h1016['매수약정수량'][:-1]], axis = 0)
sell_df = pd.concat( [sell_df , df_h1016['매도약정수량'][:-1]], axis = 0)
buy_df = buy_df.apply(lambda x: int(x))
sell_df = sell_df.apply(lambda x: int(x))
buy_df2 = buy_df.reset_index()
sell_df2 = sell_df.reset_index()
df_h1016['매수수량차이'] = df_h1016['매수약정수량'] - buy_df2[0]
df_h1016['매도수량차이'] = df_h1016['매도약정수량'] - sell_df2[0]
df_h1016
h1016_filtered = df_h1016[df_h1016['매수수량차이'] < df_h1016['매도수량차이']]
h1016_filtered
h1016_filtered['시간']
df_b6016['예상체결가격' > 0]
df_ex
# #### wall time: 2.44s
df_ex
# +
df_h1016
# -
df_ex.to_csv("data_transformed_m.csv", mode='w', encoding='cp949')
df_ex.drop(0, inplace=True)
| finanace/function1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Soap Color with Colorpy
# * requires colorpy
# +
import thinfilm
import illuminants
import matplotlib.pyplot as plt
import numpy as np
illuminant = illuminants.get_illuminant_D65()
illuminants.scale_illuminant (illuminant, 9.50)
n = 1000
nm = np.linspace(0, 1000, n)
colors = np.zeros((n, 1, 3))
for i in range(n):
colors[i] = thinfilm.thin_film(1.003, 1.44, 1.003, nm[i]).illuminated_color (illuminant)
#plt.figure(figsize=(8, 1), dpi=80)
colors = colors.transpose((1, 0, 2))
plt.imshow(colors, aspect='auto')
# -
plt.figure(figsize=(8, 0.02), dpi=80)
plt.axis('off')
plt.imshow(colors,aspect='auto')#aspect='auto'
plt.savefig("SoapColor.png")
| SoapColor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparison of `bowtie`, `bowtie2`, and `kallisto`
import pandas as pd
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# ## Data preparation
#
# We will use some of Ben's Sjorgrens data for this. We will generate a random sample of 1 million reads from the full data set.
#
# Prepare data with Snakemake
#
# ```bash
# snakemake -s aligners.snakefile
# ```
#
# It appears that kallisto needs at least 51 bases of the reference to successfully align most of the reads. Must be some kind of off-by-one issue with the data structures.
# Load alignments
names = ['QNAME', 'FLAG', 'RNAME', 'POS', 'MAPQ', 'CIGAR', 'RNEXT', 'PNEXT', 'TLEN', 'SEQ', 'QUAL']
bowtie_alns = pd.read_csv('alns/bowtie-51mer.aln', sep='\t', header=None, usecols=list(range(11)), names=names)
bowtie2_alns = pd.read_csv('alns/bowtie2-51mer.aln', sep='\t', header=None, usecols=list(range(11)), names=names)
kallisto_alns = pd.read_csv('alns/kallisto-51mer.sam', sep='\t', header=None, usecols=list(range(11)), names=names, comment='@')
(bowtie_alns.RNAME != '*').sum() / len(bowtie_alns)
(bowtie2_alns.RNAME != '*').sum() / len(bowtie2_alns)
(kallisto_alns.RNAME != '*').sum() / len(kallisto_alns)
# ## Bowtie2 vs kallisto
bt2_k_joined = pd.merge(bowtie2_alns, kallisto_alns, how='inner', on='QNAME', suffixes=['_bt2', '_k'])
# How many reads do bowtie2 and kallisto agree on?
(bt2_k_joined.RNAME_bt2 == bt2_k_joined.RNAME_k).sum()
For the minority of reads they disagree on, what do they look like?
# For the minority of reads they disagree on, what do they look like
bt2_k_joined[bt2_k_joined.RNAME_bt2 != bt2_k_joined.RNAME_k].RNAME_k
# Mostly lower sensitivity of kallisto due to indels in the read. Specifically, out of
(bt2_k_joined.RNAME_bt2 != bt2_k_joined.RNAME_k).sum()
# discordant reads, the number where kallisto failed to map is
(bt2_k_joined[bt2_k_joined.RNAME_bt2 != bt2_k_joined.RNAME_k].RNAME_k == '*').sum()
# or as a fraction
(bt2_k_joined[bt2_k_joined.RNAME_bt2 != bt2_k_joined.RNAME_k].RNAME_k == '*').sum() / (bt2_k_joined.RNAME_bt2 != bt2_k_joined.RNAME_k).sum()
# Are there any cases where bowtie2 fails to align
(bt2_k_joined[bt2_k_joined.RNAME_bt2 != bt2_k_joined.RNAME_k].RNAME_bt2 == '*').sum()
# Which means **there are no cases where bowtie and kallisto align to different peptides**.
((bt2_k_joined.RNAME_bt2 != bt2_k_joined.RNAME_k) & (bt2_k_joined.RNAME_bt2 != '*') & (bt2_k_joined.RNAME_k != '*')).sum()
# What do examples look like of kallisto aligning and bowtie2 not?
bt2_k_joined[(bt2_k_joined.RNAME_bt2 != bt2_k_joined.RNAME_k) & (bt2_k_joined.RNAME_bt2 == '*')]
# Looks like there is a perfect match to a prefix and the latter part of the read doesn't match
#
# ```
# read AAATCCACCATTGTGAAGCAGATGAAGATCATTCATGGTTACTCAGAGCA
# ref AAATCCACCATTGTGAAGCAGATGAAGATCATTCATAAAAATGGTTACTCA
#
# read GGTCCTCACGCCGCCCGCGTTCGCGGGTTGGCATTACAATCCGCTTTCCA
# ref GGTCCTCACGCCGCCCGCGTTCGCGGGTTGGCATTCCTCCCACACCAGACT
# ```
# ## Bowtie vs kallisto
bt_k_joined = pd.merge(bowtie_alns, kallisto_alns, how='inner', on='QNAME', suffixes=['_bt', '_k'])
# How many reads do bowtie and kallisto agree on?
(bt_k_joined.RNAME_bt == bt_k_joined.RNAME_k).sum()
# For the minority of reads they disagree on, what do they look like
bt_k_joined[bt_k_joined.RNAME_bt != bt_k_joined.RNAME_k][['RNAME_bt', 'RNAME_k']]
# Looks like many disagreeents, but probably still few disagreements on a positive mapping.
(bt_k_joined.RNAME_bt != bt_k_joined.RNAME_k).sum()
# discordant reads, the number where kallisto failed to map is
(bt_k_joined[bt_k_joined.RNAME_bt != bt_k_joined.RNAME_k].RNAME_k == '*').sum()
# and the number where bowtie failed is
(bt_k_joined[bt_k_joined.RNAME_bt != bt_k_joined.RNAME_k].RNAME_bt == '*').sum()
# which means **there are no disagreements on mapping**. kallisto appears to be somewhat higher sensitivity.
# ## Quantitation
bowtie_counts = pd.read_csv('counts/bowtie-51mer.tsv', sep='\t', header=0, names=['id', 'input', 'output'])
bowtie2_counts = pd.read_csv('counts/bowtie2-51mer.tsv', sep='\t', header=0, names=['id', 'input', 'output'])
kallisto_counts = pd.read_csv('counts/kallisto-51mer.tsv', sep='\t', header=0)
fig, ax = plt.subplots()
_ = ax.hist(bowtie_counts.output, bins=100, log=True)
_ = ax.set(title='bowtie')
fig, ax = plt.subplots()
_ = ax.hist(bowtie2_counts.output, bins=100, log=True)
_ = ax.set(title='bowtie2')
fig, ax = plt.subplots()
_ = ax.hist(kallisto_counts.est_counts, bins=100, log=True)
_ = ax.set(title='kallisto')
bt2_k_counts = pd.merge(bowtie2_counts, kallisto_counts, how='inner', left_on='id', right_on='target_id')
fig, ax = plt.subplots()
ax.scatter(bt2_k_counts.output, bt2_k_counts.est_counts)
sp.stats.pearsonr(bt2_k_counts.output, bt2_k_counts.est_counts)
sp.stats.spearmanr(bt2_k_counts.output, bt2_k_counts.est_counts)
# Otherwise, the kallisto index is about 3x bigger than the bowtie indices, but kallisto (5.7 s single-threaded) is about 3.5x faster than bowtie2 (20 s) and 7.3x faster than bowtie (42 s; though still appears to be using 2 threads).
# Note: it appears that kallisto needs a few extra bases on the reference to achieve its sensitivity. Performed an analysis like so:
#
# Looked at discordant cells according to Ben.
#
# ```python
# cpm = pd.read_csv('cpm.tsv', sep='\t', index_col=0, header=0)
# mlxp = pd.read_csv('mlxp.tsv', sep='\t', index_col=0, header=0)
# beadsonlycols = list(filter(lambda c: 'BEADS_ONLY' in c, mlxp.columns))
#
# samples = ['Sjogrens.serum.Sjogrens.FS08-01647.20A20G.1']
#
# oligo1 = list(filter(lambda c: 'hblOligo32108' in c, mlxp.index))[0] # hit for Ben
# oligo2 = list(filter(lambda c: 'hblOligo223219' in c, mlxp.index))[0] # null for Ben
# oligos = [oligo1, oligo2]
#
# print(cpm[beadsonlycols + samples].loc[oligos].to_csv(sep='\t'))
# print(mlxp[beadsonlycols + samples].loc[oligos].to_csv(sep='\t'))
# ```
#
# Built some indices of different sizes
#
# ```python
# from Bio import SeqIO
# k = 60
# output = f'reference{k}.fasta'
# with open(output, 'w') as op:
# for sr in SeqIO.parse('/Users/laserson/repos/phage_libraries_private/human90/human90-ref.fasta', 'fasta'):
# print(sr[:k].format('fasta'), end='', file=op)
# ```
#
# ```bash
# kallisto index -i human90-50.idx reference50.fasta
# kallisto index -i human90-51.idx reference51.fasta
# kallisto index -i human90-52.idx reference52.fasta
# kallisto index -i human90-55.idx reference55.fasta
# kallisto index -i human90-60.idx reference60.fasta
#
#
# kallisto quant -i human90-50.idx -o quant-50 --single -l 50 -s 0.1 --pseudobam Sjogrens.serum.Sjogrens.FS08-01647.20A20G.1.fastq.gz > aln-50.sam
# kallisto quant -i human90-51.idx -o quant-51 --single -l 50 -s 0.1 --pseudobam Sjogrens.serum.Sjogrens.FS08-01647.20A20G.1.fastq.gz > aln-51.sam
# kallisto quant -i human90-52.idx -o quant-52 --single -l 50 -s 0.1 --pseudobam Sjogrens.serum.Sjogrens.FS08-01647.20A20G.1.fastq.gz > aln-52.sam
# kallisto quant -i human90-55.idx -o quant-55 --single -l 50 -s 0.1 --pseudobam Sjogrens.serum.Sjogrens.FS08-01647.20A20G.1.fastq.gz > aln-55.sam
# kallisto quant -i human90-60.idx -o quant-60 --single -l 50 -s 0.1 --pseudobam Sjogrens.serum.Sjogrens.FS08-01647.20A20G.1.fastq.gz > aln-60.sam
# ```
#
# Generated the following numbers of alignments
#
# ```
# 6,369 reads pseudoaligned
# 1,419,515 reads pseudoaligned
# 1,477,736 reads pseudoaligned
# 1,490,788 reads pseudoaligned
# 1,498,420 reads pseudoaligned
# ```
#
# But looking at the results
#
# ```bash
# grep hblOligo32108 quant-50/abundance.tsv
# grep hblOligo32108 quant-51/abundance.tsv
# grep hblOligo32108 quant-52/abundance.tsv
# grep hblOligo32108 quant-55/abundance.tsv
# grep hblOligo32108 quant-60/abundance.tsv
# ```
#
# It was clear that at least 52 bases was necessary for the 50 base read to get max alignments for the peptides chosen.
| notebooks/aligners/Aligners.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 11
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
# -
# ### SIR implementation
#
# We'll use a `State` object to represent the number (or fraction) of people in each compartment.
init = State(S=89, I=1, R=0)
# To convert from number of people to fractions, we divide through by the total.
init /= sum(init)
# `make_system` creates a `System` object with the given parameters.
def make_system(beta, gamma):
"""Make a system object for the SIR model.
beta: contact rate in days
gamma: recovery rate in days
returns: System object
"""
init = State(S=89, I=1, R=0)
init /= sum(init)
t0 = 0
t_end = 7 * 14
return System(init=init, t0=t0, t_end=t_end,
beta=beta, gamma=gamma)
# Here's an example with hypothetical values for `beta` and `gamma`.
# +
tc = 3 # time between contacts in days
tr = 4 # recovery time in days
beta = 1 / tc # contact rate in per day
gamma = 1 / tr # recovery rate in per day
system = make_system(beta, gamma)
# -
# The update function takes the state during the current time step and returns the state during the next time step.
def update_func(state, t, system):
"""Update the SIR model.
state: State with variables S, I, R
t: time step
system: System with beta and gamma
returns: State object
"""
s, i, r = state
infected = system.beta * i * s
recovered = system.gamma * i
s -= infected
i += infected - recovered
r += recovered
return State(S=s, I=i, R=r)
# To run a single time step, we call it like this:
state = update_func(init, 0, system)
# Now we can run a simulation by calling the update function for each time step.
def run_simulation(system, update_func):
"""Runs a simulation of the system.
system: System object
update_func: function that updates state
returns: State object for final state
"""
state = system.init
for t in linrange(system.t0, system.t_end):
state = update_func(state, t, system)
return state
# The result is the state of the system at `t_end`
run_simulation(system, update_func)
# **Exercise** Suppose the time between contacts is 4 days and the recovery time is 5 days. After 14 weeks, how many students, total, have been infected?
#
# Hint: what is the change in `S` between the beginning and the end of the simulation?
# +
# Solution goes here
# -
# ### Using TimeSeries objects
# If we want to store the state of the system at each time step, we can use one `TimeSeries` object for each state variable.
def run_simulation(system, update_func):
"""Runs a simulation of the system.
Add three Series objects to the System: S, I, R
system: System object
update_func: function that updates state
"""
S = TimeSeries()
I = TimeSeries()
R = TimeSeries()
state = system.init
t0 = system.t0
S[t0], I[t0], R[t0] = state
for t in linrange(system.t0, system.t_end):
state = update_func(state, t, system)
S[t+1], I[t+1], R[t+1] = state
return S, I, R
# Here's how we call it.
# +
tc = 3 # time between contacts in days
tr = 4 # recovery time in days
beta = 1 / tc # contact rate in per day
gamma = 1 / tr # recovery rate in per day
system = make_system(beta, gamma)
S, I, R = run_simulation(system, update_func)
# -
# And then we can plot the results.
def plot_results(S, I, R):
"""Plot the results of a SIR model.
S: TimeSeries
I: TimeSeries
R: TimeSeries
"""
plot(S, '--', label='Susceptible')
plot(I, '-', label='Infected')
plot(R, ':', label='Recovered')
decorate(xlabel='Time (days)',
ylabel='Fraction of population')
# Here's what they look like.
plot_results(S, I, R)
savefig('figs/chap05-fig01.pdf')
# ### Using a DataFrame
# Instead of making three `TimeSeries` objects, we can use one `DataFrame`.
#
# We have to use `row` to selects rows, rather than columns. But then Pandas does the right thing, matching up the state variables with the columns of the `DataFrame`.
def run_simulation(system, update_func):
"""Runs a simulation of the system.
system: System object
update_func: function that updates state
returns: TimeFrame
"""
frame = TimeFrame(columns=system.init.index)
frame.row[system.t0] = system.init
for t in linrange(system.t0, system.t_end):
frame.row[t+1] = update_func(frame.row[t], t, system)
return frame
# Here's how we run it, and what the result looks like.
# +
tc = 3 # time between contacts in days
tr = 4 # recovery time in days
beta = 1 / tc # contact rate in per day
gamma = 1 / tr # recovery rate in per day
system = make_system(beta, gamma)
results = run_simulation(system, update_func)
results.head()
# -
# We can extract the results and plot them.
plot_results(results.S, results.I, results.R)
# ## Exercises
#
# **Exercise** Suppose the time between contacts is 4 days and the recovery time is 5 days. Simulate this scenario for 14 weeks and plot the results.
# +
# Solution goes here
| code/chap11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Italian Code For Identification Of Drugs
# ## Introduction
# The function `clean_it_aic()` cleans a column containing Italian code for identification of drug (AIC) strings, and standardizes them in a given format. The function `validate_it_aic()` validates either a single AIC strings, a column of AIC strings or a DataFrame of AIC strings, returning `True` if the value is valid, and `False` otherwise.
# AIC strings can be converted to the following formats via the `output_format` parameter:
#
# * `compact`: only number strings without any seperators or whitespace, like "000307052"
# * `standard`: AIC strings with proper whitespace in the proper places. Note that in the case of AIC, the compact format is the same as the standard one.
# * `base10`: convert a BASE32 representation to a BASE10 one, like "000307052".
# * `base32`: convert a BASE10 representation to a BASE32 one, like "009CVD". Note 'compact' may contain both BASE10 and BASE32 represatation.
#
# Invalid parsing is handled with the `errors` parameter:
#
# * `coerce` (default): invalid parsing will be set to NaN
# * `ignore`: invalid parsing will return the input
# * `raise`: invalid parsing will raise an exception
#
# The following sections demonstrate the functionality of `clean_it_aic()` and `validate_it_aic()`.
# ### An example dataset containing AIC strings
import pandas as pd
import numpy as np
df = pd.DataFrame(
{
"aic": [
'000307052',
'999999',
'7542011030',
'7552A10004',
'8019010008',
"hello",
np.nan,
"NULL",
],
"address": [
"123 Pine Ave.",
"main st",
"1234 west main heights 57033",
"apt 1 789 s maple rd manhattan",
"robie house, 789 north main street",
"1111 S Figueroa St, Los Angeles, CA 90015",
"(staples center) 1111 S Figueroa St, Los Angeles",
"hello",
]
}
)
df
# ## 1. Default `clean_it_aic`
#
# By default, `clean_it_aic` will clean aic strings and output them in the standard format with proper separators.
from dataprep.clean import clean_it_aic
clean_it_aic(df, column = "aic")
# ## 2. Output formats
# This section demonstrates the output parameter.
# ### `standard` (default)
clean_it_aic(df, column = "aic", output_format="standard")
# ### `compact`
clean_it_aic(df, column = "aic", output_format="compact")
# ### `base10`
clean_it_aic(df, column = "aic", output_format="base10")
# ### `base32`
clean_it_aic(df, column = "aic", output_format="base32")
# ## 3. `inplace` parameter
#
# This deletes the given column from the returned DataFrame.
# A new column containing cleaned AIC strings is added with a title in the format `"{original title}_clean"`.
clean_it_aic(df, column="aic", inplace=True)
# ## 4. `errors` parameter
# ### `coerce` (default)
clean_it_aic(df, "aic", errors="coerce")
# ### `ignore`
clean_it_aic(df, "aic", errors="ignore")
# ## 4. `validate_it_aic()`
# `validate_it_aic()` returns `True` when the input is a valid AIC. Otherwise it returns `False`.
#
# The input of `validate_it_aic()` can be a string, a Pandas DataSeries, a Dask DataSeries, a Pandas DataFrame and a dask DataFrame.
#
# When the input is a string, a Pandas DataSeries or a Dask DataSeries, user doesn't need to specify a column name to be validated.
#
# When the input is a Pandas DataFrame or a dask DataFrame, user can both specify or not specify a column name to be validated. If user specify the column name, `validate_it_aic()` only returns the validation result for the specified column. If user doesn't specify the column name, `validate_it_aic()` returns the validation result for the whole DataFrame.
from dataprep.clean import validate_it_aic
print(validate_it_aic('000307052'))
print(validate_it_aic('999999'))
print(validate_it_aic('7542011030'))
print(validate_it_aic('7552A10004'))
print(validate_it_aic('8019010008'))
print(validate_it_aic("hello"))
print(validate_it_aic(np.nan))
print(validate_it_aic("NULL"))
# ### Series
validate_it_aic(df["aic"])
# ### DataFrame + Specify Column
validate_it_aic(df, column="aic")
# ### Only DataFrame
validate_it_aic(df)
| docs/source/user_guide/clean/clean_it_aic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This notebook is aimed at solving classification problems using logistic regression with gradient descent
# +
from sklearn.metrics import log_loss
from sklearn.linear_model import LogisticRegression
from sklearn import datasets
iris = datasets.load_iris()
import matplotlib.pyplot as plt
import numpy as np
from sklearn.metrics import accuracy_score
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# -
# ### This section will be for binary logistic regression
# +
# Using two classes for logistic regression
X = iris.data[:, :2][0:100]
y = iris.target[0:100]
y_c = iris.target[0:100]
y.shape = (100,1)
assert y.shape == (100,1)
assert X.shape == (100,2)
assert y_c.shape == (100,)
# -
# This data seems to be linerly separable
plt.scatter(X[:,0][0:50], X[:,1][0:50], c="blue", label="Setosa")
plt.scatter(X[:,0][50:100], X[:,1][50:100], c="orange", label="Versicolour")
plt.xlabel("Variable 1")
plt.ylabel("Variable 2")
plt.title("Flower Type")
plt.legend()
plt.show()
# ### Sigmoid activation function
#
# $$ h_\theta(x) = \frac{1}{1+e^{-x}} $$
# Returns the sigmoid activation of input
def sigmoid(x):
return 1 / (1 + np.exp(-x))
# ### Logistic regression cost function (log loss)
#
# $$ J(\theta) = - \frac{1}{m}\sum_{i=1}^{m}y^{(i)}log(h_\theta(x^{(i)}))+
# (1-y^{(i)})log(1-h_\theta(x^{(i)})$$
# $$ note: h_\theta(x^{(i)}) = \frac{1}{1+e^{-(bias+X\theta.T)}}$$
# Log loss for logistic regression
def get_cost(X,y,weight,bias):
return -np.average(y*np.log(sigmoid(bias+np.dot(X,weight.T)))+(1-y)*np.log(1-(sigmoid(bias+np.dot(X,weight.T)))))
# +
# Testing cost function
weight = np.array([[0],[0]])
weight.shape = (1,2)
bias = 1
assert weight.shape == (1,2)
# -
# Ensuring our cost function is working properly using sklearn implementation
assert get_cost(X,y,weight,bias) == log_loss(y,sigmoid(bias+np.dot(X,weight.T)))
# ### Updating weight and bias terms
#
# $$ \begin{split} & f^\prime(W) = \frac{1}{m}\big[h_\theta(x^{(i)})-y^{(i)})x_j^{(i)}\big] \\
# & f^\prime(Bias) = \frac{1}{m}\big[h_\theta(x^{(i)})-y^{(i)})\big] \end{split} $$
# Updates the weights and bias terms based on gradient descent
def update_param(X,y,weight,bias,lr=0.01):
"""
X.shape = (100,2)
y.shape = (100,1)
weight.shape = (1,2)
"""
df_dm = (1/len(X)) * np.dot((sigmoid(np.dot(X,weight.T))-y).T,X)
df_db = np.average(sigmoid(np.dot(X,weight.T))-y)
weight = weight - (lr * df_dm)
bias = bias - (lr * df_db)
return weight,bias
def train_model(X,y,weight,bias,iters,lr=0.01):
costs = []
for i in range(iters):
weight,bias = update_param(X,y,weight,bias,lr)
cost = get_cost(X,y,weight,bias)
costs.append(cost)
return weight,bias,costs
# Returns the prediction for any given X variable
def predictions(X,weight,bias):
"""
X.shape = (100,2)
weight = (1,2)
"""
return sigmoid(bias + np.dot(X,weight.T))
# +
# Parameters for testing out my logistic regression model
model_weights = np.array([[0],[0]])
model_weights.shape = (1,2)
model_bias = 0
assert model_weights.shape == (1,2)
# -
# Trying out the logistic regression model
ret_weight,ret_bias,ret_costs = train_model(X,y,model_weights,model_bias,15000,0.01)
time_multi = [i for i in range(15000)]
# Graphing my cost function
plt.plot(time_multi, ret_costs)
plt.title("Cost vs Time Logistic Regression")
plt.xlabel("Step Number")
plt.ylabel("Cost")
plt.show()
# Getting predicted values
pred = predictions(X,ret_weight,ret_bias)
pred_2 = predictions(X,ret_weight,ret_bias)
pred.shape
# Turning predictions into binary classifications
pred[pred >= 0.5] = 1
pred[pred < 0.5] = 0
# #### Overall my model performed very similar to the sklearn model
# Comparing my coefficients and score to sklearn coefficients
accuracy = accuracy_score(y,pred)
print("Accuracy:", accuracy)
print("Weights:", ret_weight)
print("Bias:", ret_bias)
print("Final cost:", ret_costs[-1])
# +
# Using sklearn's logistic regression to determine the ideal params
lr = LogisticRegression()
lr.fit(X, y)
predictedVals = lr.predict(X)
predictedProb = lr.predict_proba(X)
print("Accuracy: " + str(lr.score(X, y)))
print("Coefs: " + str(lr.coef_))
print("Intercept: " + str(lr.intercept_))
print("Final cost:", log_loss(y, predictedProb))
# -
# ### I will attempt to use regularization to see if my model improves (lambda will be my regularization term)
#
# This involves updating the derivative term for weights:
#
# $$ f^\prime(W) = \frac{1}{m}\big[h_\theta(x^{(i)})-y^{(i)})x_j^{(i)} \big] + \frac{\lambda}{m}\theta_j $$
# $$ \theta_j := \theta_j(1-\alpha\frac{\lambda}{m}) - \alpha\frac{1}{m}\big[h_\theta(x^{(i)})-y^{(i)})x_j^{(i)} \big]$$
#
# This also involves updating the cost function to include regularization:
#
# $$ J(\theta) = - \frac{1}{m}\sum_{i=1}^{m}y^{(i)}log(h_\theta(x^{(i)}))+
# (1-y^{(i)})log(1-h_\theta(x^{(i)}) + \frac{\lambda}{2m}\sum_{j=1}^{n}\theta_j^2$$
# Log loss for logistic regression
def get_cost_reg(X,y,weight,bias,lmbda):
inner = (y*np.log(sigmoid(bias+np.dot(X,weight.T)))+(1-y)*np.log(1-(sigmoid(bias+np.dot(X,weight.T)))))
return -np.average(inner+((lmbda/(2*len(X))*np.sum(weight**2))))
# Updates the weights and bias terms based on gradient descent
def update_param_reg(X,y,weight,bias,lr,lmbda):
"""
X.shape = (100,2)
y.shape = (100,1)
weight.shape = (1,2)
"""
df_dm = (1/len(X)) * np.dot((sigmoid(np.dot(X,weight.T))-y).T,X) #may need a negative
df_db = np.average(sigmoid(np.dot(X,weight.T))-y)
weight = weight * (1-lr*(lmbda/len(X))) - (lr * df_dm)
bias = bias - (lr * df_db)
return weight,bias
def train_model_reg(X,y,weight,bias,iters,lmbda,lr=0.01):
costs = []
for i in range(iters):
weight,bias = update_param_reg(X,y,weight,bias,lr,lmbda)
cost = get_cost_reg(X,y,weight,bias,lmbda)
costs.append(cost)
return weight,bias,costs
# Initializing params to test my regularized model
reg_weight = np.array([[0,0]])
reg_bias = 0
assert reg_weight.shape == (1,2)
lmbdas = [0.1,0.5,1,3,5]
end_costs = []
for lmbda in lmbdas:
_,_,reg_costs = train_model_reg(X,y,reg_weight,reg_bias,10000,lmbda,0.01)
end_costs.append(reg_costs[-1])
print("lambda value:{}, end cost: {}".format(lmbda,reg_costs[-1]))
reg_weight = np.array([[0,0]])
reg_bias = 0
assert reg_weight.shape == (1,2)
# It seems the lambda value of 0.5 worked the best so I will test the accuracy of that model
# Note that the new final cost value is smaller than my model without regularization
reg_weight,reg_bias,reg_costs = train_model_reg(X,y,reg_weight,reg_bias,13000,0.5,0.01)
# Checking the accuracy score of my new model
""" Although the cost is lower, the accuracy is lower. This is not surprising given that increasing regularization
ultimately is meant to reduce overfitting and I havn't done any train test split and thus overfitting on the
training data in this case would lead to a higher accuracy score.
"""
reg_pred = predictions(X,reg_weight,reg_bias)
reg_pred[reg_pred >= 0.5] = 1
reg_pred[reg_pred < 0.5] = 0
accuracy = accuracy_score(y,reg_pred)
print("Accuracy:", accuracy)
time_multi = [i for i in range(13000)]
# Graphing my cost function
plt.plot(time_multi, reg_costs)
plt.title("Cost vs Time Logistic Regression, Regularized")
plt.xlabel("Step Number")
plt.ylabel("Cost")
plt.show()
| simple_models/Logistic_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.preprocessing.text import Tokenizer
samples = ['The cat sat on the mat.', 'The dog ate my homework.']
tokenizer = Tokenizer(num_words=1000)
tokenizer.fit_on_texts(samples)
sequences = tokenizer.texts_to_sequences(samples)
sequences
one_hot_results = tokenizer.texts_to_matrix(samples, mode='binary')
one_hot_results.shape
word_index = tokenizer.word_index
word_index
from keras.layers import Embedding
embedding_layer = Embedding(1000, 64)
embedding_layer
from keras.datasets import imdb
from keras import preprocessing
# +
max_features = 10000
max_len = 20
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
x_train = preprocessing.sequence.pad_sequences(x_train, maxlen=max_len)
x_test = preprocessing.sequence.pad_sequences(x_test, maxlen=max_len)
# -
x_train.shape
x_test.shape
# +
from keras.models import Sequential
from keras.layers import Flatten, Dense, Embedding
model = Sequential()
model.add(Embedding(10000, 8, input_length=max_len))
model.add(Flatten())
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
# -
model.summary()
history = model.fit(x_train, y_train, epochs=10, batch_size=32, validation_split=0.2)
# # IMDbの単語埋め込み
# +
import os
imdb_dir = './data/aclImdb'
train_dir = os.path.join(imdb_dir, 'train')
labels = []
texts = []
for label_type in ['neg', 'pos']:
dir_name = os.path.join(train_dir, label_type)
for fname in os.listdir(dir_name):
if fname[-4:] == '.txt':
f = open(os.path.join(dir_name, fname))
texts.append(f.read())
f.close()
if label_type == 'neg':
labels.append(0)
else:
labels.append(1)
# -
len(texts)
len(labels)
texts[0]
labels[0]
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
import numpy as np
max_len = 100
training_samples = 200
validation_samples = 10000
max_words = 10000
tokenizer = Tokenizer(num_words=max_words)
tokenizer.fit_on_texts(texts)
sequences = tokenizer.texts_to_sequences(texts)
len(sequences)
print(sequences[0])
word_index = tokenizer.word_index
print(len(word_index))
list(word_index.keys())[:10]
data = pad_sequences(sequences, maxlen=max_len)
print(data.shape)
labels = np.asarray(labels)
print(data.shape)
print(labels.shape)
indices = np.arange(data.shape[0])
np.random.shuffle(indices)
data = data[indices]
labels = labels[indices]
x_train = data[:training_samples]
y_train = labels[:training_samples]
x_val = data[training_samples: training_samples + validation_samples]
y_val = labels[training_samples: training_samples + validation_samples]
x_train.shape
y_train.shape
x_val.shape
y_val.shape
# +
glove_dir = './data/glove.6B'
embeddings_index = {}
f = open(os.path.join(glove_dir, 'glove.6B.100d.txt'))
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print(len(embeddings_index))
# +
embedding_dim = 100
embedding_matrix = np.zeros((max_words, embedding_dim))
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if i < max_words:
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
# +
from keras.models import Sequential
from keras.layers import Embedding, Flatten, Dense
model = Sequential()
model.add(Embedding(max_words, embedding_dim, input_length=max_len))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# -
embedding_matrix.shape
model.layers[0].set_weights([embedding_matrix])
model.layers[0].trainable = False
model.summary()
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
history = model.fit(x_train, y_train, epochs=10, batch_size=32, validation_data=(x_val, y_val))
model.save('models/pre_trained_glove_model.h5')
# +
import matplotlib.pyplot as plt
# %matplotlib inline
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# plot accuracy
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
# plot loss
plt.figure()
plt.plot(epochs, loss, 'ro', label='Training loss')
plt.plot(epochs, val_loss, 'r', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# +
test_dir = os.path.join(imdb_dir, 'test')
labels = []
texts = []
for label_type in ['neg', 'pos']:
dir_name = os.path.join(test_dir, label_type)
for fname in sorted(os.listdir(dir_name)):
if fname[-4:] == '.txt':
f = open(os.path.join(dir_name, fname))
texts.append(f.read())
f.close()
if label_type == 'neg':
labels.append(0)
else:
labels.append(1)
sequences = tokenizer.texts_to_sequences(texts)
x_test = pad_sequences(sequences, maxlen=max_len)
y_test = np.asarray(labels)
# -
model.load_weights('models/pre_trained_glove_model.h5')
model.evaluate(x_test, y_test)
# ## RNN
# +
import numpy as np
timesteps = 100
input_features = 32
output_features = 64
inputs = np.random.random((timesteps, input_features))
state_t = np.zeros((output_features, ))
W = np.random.random((output_features, input_features))
U = np.random.random((output_features, output_features))
b = np.random.random((output_features, ))
# -
successive_outputs = []
for input_t in inputs:
output_t = np.tanh(np.dot(W, input_t) + np.dot(U, state_t) + b)
successive_outputs.append(output_t)
state_t = output_t
len(successive_outputs)
len(successive_outputs[0])
final_output_sequence = np.stack(successive_outputs, axis=0)
final_output_sequence.shape
from keras.models import Sequential
from keras.layers import Embedding, SimpleRNN
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32))
model.summary()
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32, return_sequences=True))
model.summary()
model = Sequential()
model.add(Embedding(10000, 32))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32, return_sequences=True))
model.add(SimpleRNN(32))
model.summary()
# +
from keras.datasets import imdb
from keras.preprocessing import sequence
max_features = 10000
max_len = 500
batch_size = 32
(input_train, y_train), (input_test, y_test) = imdb.load_data(num_words=max_features)
print(len(input_train))
print(len(input_test))
# -
input_train = sequence.pad_sequences(input_train, maxlen=max_len)
input_test = sequence.pad_sequences(input_test, maxlen=max_len)
input_train.shape
input_test.shape
# +
from keras.models import Sequential
from keras.layers import Embedding, SimpleRNN, Dense
model = Sequential()
model.add(Embedding(max_features, 32))
model.add(SimpleRNN(32))
model.add(Dense(1, activation='sigmoid'))
# -
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(input_train, y_train, epochs=10, batch_size=128, validation_split=0.2)
# +
import matplotlib.pyplot as plt
# %matplotlib inline
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# plot accuracy
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
# plot loss
plt.figure()
plt.plot(epochs, loss, 'ro', label='Training loss')
plt.plot(epochs, val_loss, 'r', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# +
from keras.layers import LSTM
model = Sequential()
model.add(Embedding(max_features, 32))
model.add(LSTM(32))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
history = model.fit(input_train, y_train, epochs=10, batch_size=128, validation_split=0.2)
# +
import matplotlib.pyplot as plt
# %matplotlib inline
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# plot accuracy
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
# plot loss
plt.figure()
plt.plot(epochs, loss, 'ro', label='Training loss')
plt.plot(epochs, val_loss, 'r', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
# ## 時系列データ
# +
import os
data_dir = './data/jena_climate'
fname = os.path.join(data_dir, 'jena_climate_2009_2016.csv')
# -
f = open(fname)
data = f.read()
f.close()
lines = data.split('\n')
header = lines[0].split(',')
lines = lines[1:]
header
len(lines)
# +
import numpy as np
float_data = np.zeros((len(lines), len(header) - 1))
for i, line in enumerate(lines):
values = [float(x) for x in line.split(',')[1:]]
float_data[i, :] = values
# -
float_data.shape
# +
from matplotlib import pyplot as plt
# %matplotlib inline
temp = float_data[:, 1]
plt.plot(range(len(temp)), temp)
plt.show()
# -
plt.plot(range(1440), temp[:1440])
mean = float_data[:200000].mean(axis=0)
float_data -= mean
std = float_data[:200000].std(axis=0)
float_data /= std
float_data.shape
def generator(data, lookback, delay, min_index, max_index, shuffle=False, batch_size=128, step=6):
if max_index is None:
max_index = len(data) - delay - 1
i = min_index + lookback
while 1:
if shuffle:
rows = np.random.randint(min_index + lookback, max_index, size=batch_size)
else:
if i + batch_size >= max_index:
i = min_index + lookback
rows = np.arange(i, min(i + batch_size, max_index))
i += len(rows)
samples = np.zeros((len(rows), lookback // step, data.shape[-1]))
targets = np.zeros((len(rows), ))
for j, row in enumerate(rows):
indices = range(rows[j] - lookback, rows[j], step)
samples[j] = data[indices]
targets[j] = data[rows[j] + delay][1]
yield samples, targets
# +
lookback = 1440 # 1440 / 60 / 10 = 過去10日間のデータを使用
step = 6 # 6 * 10 = 60分間隔でデータを取得
delay = 144 # 目的値は24時間後 = 24 * 60 / 10 = 144
batch_size = 128
train_gen = generator(float_data,
lookback=lookback,
delay=delay,
min_index=0,
max_index=200000,
shuffle=True,
step=step,
batch_size=batch_size)
val_gen = generator(float_data,
lookback=lookback,
delay=delay,
min_index=200001,
max_index=300000,
step=step,
batch_size=batch_size)
test_gen = generator(float_data,
lookback=lookback,
delay=delay,
min_index=300001,
max_index=None,
step=step,
batch_size=batch_size)
# -
val_steps = (300000 - 200001 - lookback) // batch_size
val_steps
test_steps = (len(float_data) - 300001 - lookback) // batch_size
test_steps
def evaluate_naive_method():
batch_maes = []
for step in range(val_steps):
# samples = (32, 240, 14) # 240は1440/6、14は特徴量(1が気温)
samples, targets = next(val_gen)
preds = samples[:, -1, 1]
mae = np.mean(np.abs(preds - targets))
batch_maes.append(mae)
print(np.mean(batch_maes))
evaluate_naive_method()
celsius_mae = 0.28960 * std[1]
celsius_mae
from keras.models import Sequential
from keras import layers
from keras.optimizers import RMSprop
# +
model = Sequential()
model.add(layers.Flatten(input_shape=(lookback // step, float_data.shape[-1])))
model.add(layers.Dense(32, activation='relu'))
model.add(layers.Dense(1))
model.compile(optimizer=RMSprop(), loss='mae')
history = model.fit_generator(train_gen,
steps_per_epoch=500,
epochs=20,
validation_data=val_gen,
validation_steps=val_steps)
# -
model.summary()
from keras.models import Sequential
from keras import layers
from keras.optimizers import RMSprop
model = Sequential()
model.add(layers.GRU(32, input_shape=(None, float_data.shape[-1])))
model.add(layers.Dense(1))
model.compile(optimizer=RMSprop(), loss='mae')
history = model.fit_generator(train_gen,
steps_per_epoch=500,
epochs=20,
validation_data=val_gen,
validation_steps=val_steps)
# val_loss: 0.4387
# +
from keras.models import Sequential
from keras import layers
from keras.optimizers import RMSprop
model = Sequential()
model.add(layers.GRU(32,
dropout=0.2,
recurrent_dropout=0.1,
input_shape=(None, float_data.shape[-1])))
model.add(layers.Dense(1))
model.compile(optimizer=RMSprop(), loss='mae')
history = model.fit_generator(train_gen,
steps_per_epoch=500,
epochs=40,
validation_data=val_gen,
validation_steps=val_steps)
# -
from keras.models import Sequential
from keras import layers
from keras.optimizers import RMSprop
model = Sequential()
model.add(layers.GRU(32, dropout=0.2, recurrent_dropout=0.1, return_sequences=True, input_shape=(None, float_data.shape[-1])))
model.add(layers.GRU(64, activation='relu', dropout=0.2, recurrent_dropout=0.1))
model.add(layers.Dense(1))
model.summary()
model.compile(optimizer=RMSprop(), loss='mae')
history = model.fit_generator(train_gen, steps_per_epoch=500, epochs=40, validation_data=val_gen, validation_steps=val_steps)
model = Sequential()
model.add(layers.Bidirectional(layers.GRU(32), input_shape=(None, float_data.shape[-1])))
model.add(layers.Dense(1))
model.compile(optimizer=RMSprop(), loss='mae')
history = model.fit_generator(train_gen, steps_per_epoch=500, epochs=40, validation_data=val_gen, validation_steps=val_steps)
from keras.datasets import imdb
from keras.preprocessing import sequence
# +
max_features = 10000
max_len = 500
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
x_train = sequence.pad_sequences(x_train, maxlen=max_len)
x_test = sequence.pad_sequences(x_test, maxlen=max_len)
# -
# (batch, seq_len)
x_train.shape, x_test.shape
from keras.models import Sequential
from keras import layers
from keras.optimizers import RMSprop
model = Sequential()
model.add(layers.Embedding(max_features, 128, input_length=max_len))
model.add(layers.Conv1D(32, 7, activation='relu'))
model.add(layers.MaxPooling1D(5))
model.add(layers.Conv1D(32, 7, activation='relu'))
model.add(layers.GlobalMaxPool1D())
model.add(layers.Dense(1))
model.summary()
model.compile(optimizer=RMSprop(lr=1e-4),
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(x_train, y_train, epochs=10, batch_size=128, validation_split=0.2)
# Conv1D + RNN
model = Sequential()
model.add(layers.Conv1D(32, 5, activation='relu', input_shape=(280, float_data.shape[-1])))
model.add(layers.MaxPooling1D(3))
model.add(layers.Conv1D(32, 5, activation='relu'))
model.add(layers.GRU(32, dropout=0.1, recurrent_dropout=0.5))
model.add(layers.Dense(1))
model.summary()
| dl_with_python/chapter6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import cv2
import numpy as np
from keras.utils import Sequence
from transform.sequences import RandomRotationTransformer
"""First, let's create a simple Sequence that load an image and resize it."""
class SimpleSequence(Sequence):
def __init__(self, paths, shape=(200, 200)):
self.paths = paths
self.shape = shape
self.batch_size = 1
def __len__(self):
return len(self.paths) // self.batch_size
def __getitem__(self, index):
paths = self.paths[index * self.batch_size:(index + 1) * self.batch_size]
X = [cv2.resize(cv2.imread(p), self.shape) for p in paths]
y = [cv2.cvtColor(x, cv2.COLOR_BGR2GRAY) for x in X]
return np.array(X), np.array(y).reshape([self.batch_size, self.shape[0], self.shape[1], 1])
# -
"""Transformers are Sequence that takes a Sequence to modify it."""
from glob import glob
paths = glob('/data/images_folder/*.jpg')
seq = SimpleSequence(paths)
transformer = RandomRotationTransformer(seq,rg=35)
X,y = transformer[0]
# +
"""Canonical example where X is transformed but not y"""
import matplotlib.pyplot as plt
# %matplotlib inline
im = np.concatenate((X[0],cv2.cvtColor(y[0],cv2.COLOR_GRAY2BGR)),1)
plt.imshow(cv2.cvtColor(im,cv2.COLOR_BGR2RGB))
# -
"""Applying the SAME transformation to X and y is done by specifying a mask."""
transformer = RandomRotationTransformer(seq,35,mask=[True,True])
X,y = transformer[0]
im = np.concatenate((X[0],cv2.cvtColor(y[0],cv2.COLOR_GRAY2BGR)),1)
plt.imshow(cv2.cvtColor(im,cv2.COLOR_BGR2RGB))
"""You can also disable data augmentation entirely."""
transformer = RandomRotationTransformer(seq,35,mask=False)
X,y = transformer[0]
im = np.concatenate((X[0],cv2.cvtColor(y[0],cv2.COLOR_GRAY2BGR)),1)
plt.imshow(cv2.cvtColor(im,cv2.COLOR_BGR2RGB))
| examples/simple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8 - AzureML
# language: python
# name: python38-azureml
# ---
# # Migrating from `azureml-core` Tracking to `MLflow` Tracking APIs
# ## Goal
#
# The goals of this notebook are:
# 1. To provide a comparison of logging APIs in `azureml-core` and `MLflow`
# 2. To provide examples that can be executed in real-time.
# 3. Provide a way to migrate from `azureml-core`'s run management and tracking APIs to `MLflow` run management and tracking APIs
#
# ## Prerequisites
#
# Before proceeding, please make sure you have the following pip packages installed:
# 1. `azureml-core`
# 2. `mlflow`
# 3. `azureml-mlflow`
#
# Please make sure you also have an Azure Machine Learning workspace. You can create one using the following steps: https://docs.microsoft.com/en-us/azure/machine-learning/quickstart-create-resources
#
# ## Relevant Documentation
#
# If you would like a more detailed view of the logging APIs available in `MLflow`, please see [here](https://www.mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_artifact). For a general overview of `MLflow` tracking in general, please see [here](https://www.mlflow.org/docs/latest/tracking.html).
# ## Setup
#
# Run the following cells to retrieve your workspace and set the `MLflow` tracking URI to point at the AzureML backend. This step is required for `MLflow` metrics and artifacts to get logged properly to your workspace.
# %pip install -r requirements.txt
# +
from azureml.core import Workspace
import mlflow
ws = Workspace.from_config()
mlflow.set_tracking_uri(ws.get_mlflow_tracking_uri())
# -
# ## Creating and Managing Experiments with `MlFlow`
#
# You can create and manage experiments using `Mlflow` just as you can with `azureml-core`. You can find more information on `mlflow.create_experiment()` [here](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.create_experiment), `mlflow.get_experiment_by_name()` [here]("https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.get_experiment"), and `mlflow.list_experiments()` [here](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.list_experiments). You can also set the default experiment to use for all runs with `mlflow.set_experiment()`; more information can be found [here](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.set_experiment).
# +
from mlflow.tracking import MlflowClient
mlflow.create_experiment("create-experiment-mlflow")
print(mlflow.get_experiment_by_name("create-experiment-mlflow"))
print(mlflow.list_experiments())
# -
# ## Creating and Managing Runs with `MLflow`
# ### Interactive Runs
#
# In `azureml-core`, interactive runs are started using `experiment.start_logging()`. You can start interactive runs using `MLflow`and an `AzureML` backend by doing the following:
#
# 1. Follow the setup instructions above
# 1. If you want to set an experiment for the run, you can call `mlflow.set_experiment("<name of experiment>")`, which will automatically create any new runs under that experiment.
# 1. Start the run by either:
# 1. Using `mlflow.start_run()`. This method returns an [`mlflow.ActiveRun`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.ActiveRun) (clickable link).
# 2. You can call this method using the context manager paradigm: `with mlflow.start_run() as run`.
# 1. Calling any of `MLflow`'s logging APIs. A new run will be started if one does not exist. You can retrieve the current active run by calling `mlflow.active_run()`
# 1. Complete the run.
# 1. If you are using the run as a context manager, the run will automatically complete when the context manager exits.
# 1. Otherwise, you can end the currently active run by calling `mlflow.end_run()`
#
#
# **Note**: The `mlflow.ActiveRun` object returned by `mlflow.active_run()` **will not** contain items like parameters, metrics, etc. You may find more information [here](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.active_run). See the section `Viewing Run Metrics` to see the steps retrieving metrics for a run
# Starting an MLflow run using a context manager
mlflow.set_experiment("create-experiment-mlflow-context-manager")
with mlflow.start_run() as run:
# run is started when context manager is entered and ended when context manager exits
pass
# Starting an MLflow
mlflow.set_experiment("create-experiment-mlflow-manual")
interactive_mlflow_run = mlflow.start_run()
# End the MLflow run manually
mlflow.end_run()
# Finally, an MLflow run can be started by calling one of the logging APIs
mlflow.set_experiment("create-experiment-mlflow-logging")
mlflow.log_metric("sample_metric", 1)
# Note that the run object returned by `mlflow.active_run()` will not have data like metrics and parameters
run = mlflow.active_run()
print(run)
# End the MLflow run manually
mlflow.end_run()
# ### Remote Runs
#
# For remote training runs, the tracking URI and experiment for the run are already set by the AzureML backend. Instead of calling `azureml-core`'s `Run.get_context()` to retrieve the run inside of a training script, you can either:
# 1. Call `mlflow.start_run()` to get the `MLflow` run for that particular training job. As with an interactive local run, you can use this method with the context manager paradigm.
# 1. Or, use one of the `MLflow` logging APIs to start logging metrics and artifacts. Afterwards, you can call `mlflow.active_run()` to retrieve the current `MLflow` run.
#
# It is important to note that unless you do one of the above in your training script, `mlflow.active_run()` will return `None`.
# ## Logging API Comparison
#
# The following section outlines each of the logging APIs available in the `azureml-core` and presents how to use `MLflow` to either:
# 1. Reproduce the same behavior
# 2. Provide functionality similar to that of `azureml-core`
#
# The structure of the document will be as follows:
#
# ## AzureML Tracking API
#
# < Table describing AzureML API, accepted parameters, and available `MLflow` alternatives with notes. The `MLflow` alternatives will have clickable links to the official `MLflow` documentation for that method >
#
# < Code examples >
# ### Start the interactive `AzureML` Run
# +
from azureml.core import Experiment
# create an AzureML experiment and start a run
experiment = Experiment(ws, "logging-with-mlflow-and-azureml-core")
azureml_run = experiment.start_logging()
# -
# ### Start the interactive `MLflow` Run
# Set the MLflow experiment and start a run
mlflow.set_experiment("logging-with-mlflow-and-azureml-core")
mlflow_run = mlflow.start_run()
# ###
# ## `azureml.core.Run.log()`
#
# | azureml.core API | MLFlow API | Notes |
# |-------------------------------|-------------------------------------------|-----------------|
# | `Run.log(name: str, value: float, description="", step=None)` | [`mlflow.log_metric(key: str, value: float, step: Optional[int] = None)`](https://www.mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_metric) | |
# | `Run.log(self, name: str, value: bool, description="", step=None)` | [`mlflow.log_metric(key: str, value: float, step: Optional[int] = None)`](https://www.mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_metric), where `value` is 0 or 1 | |
# | `Run.log(self, name: str, value: str, description="", step=None)` | `mlflow.log_text(text: str, artifact_file: str) ` | `mlflow.log_text()` logs the string as an artifact and will not be considered a metric. The result of `log_text()` will be displayed in the `Outputs + logs` tab of the AzureML Studio UI |
# ### Code example: logging an integer or float metric
# +
# Using AzureML
azureml_run.log("sample_int_metric", 1)
# Using MLflow
mlflow.log_metric("sample_int_metric", 1)
# -
# ### Code example: logging a boolean metric
# +
# Using AzureML
azureml_run.log("sample_boolean_metric", True)
# Using MLflow
mlflow.log_metric("sample_boolean_metric", 1)
# -
# ### Code example: logging a string metric
# +
# Using AzureML
azureml_run.log("sample_string_metric", "a_metric")
# Using MLflow. Note that the string will get logged only as an artifact and will not be logged as a metric
mlflow.log_text("sample_string_text", "string.txt")
# -
# ## `azureml.core.Run.log_image()`
# | azureml.core API | MLFlow equivalent or proposed alternative | Notes |
# |-------------------------------|-------------------------------------------|-------|
# | `Run.log_image(name: str, plot: Optional[matplotlib.pyplot] = None)` | [`mlflow.log_figure(figure: Union[matplotlib.figure.Figure, plotly.graph_objects.Figure], artifact_file: str)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_figure) | <ul><li/>Appears in Images tab.<li/>Logged as an artifact<li/>Both appear in Studio UI<li/>`mlflow.log_figure` is **experimental**</ul>
# | `Run.log_image(name: str, path: Optional[Union[str, os.PathLike, pathlib.Path] = None)` | [`mlflow.log_artifact(local_path: str, artifact_path: Optional[str] = None)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_artifact) |<ul><li/>Image logged with `MLflow` appears in the Images tab.<li/>`MLflow` logs image as an artifact<li/>Works with png and jpeg.</ul> |
# ### Logging an image saved to a png or jpeg file
# +
# Using AzureML
azureml_run.log_image("sample_image", path="Azure.png")
# Using MLflow
mlflow.log_artifact("Azure.png")
# -
# ### Logging a matplotlib.pyplot
# +
import matplotlib.pyplot as plt
plt.plot([1, 2, 3])
# Using AzureML
azureml_run.log_image("sample_pyplot", plot=plt)
# Using MLflow
fig, ax = plt.subplots()
ax.plot([0, 1], [2, 3])
mlflow.log_figure(fig, "sample_pyplot.png")
# -
# ## `azureml.core.Run.log_list()`
# | azureml.core API | MLFlow equivalent or proposed alternative | Notes |
# |-------------------------------|-------------------------------------------|-----------------|
# | `Run.log_list(name, value, description="")` | [`MlFlowClient().log_batch(run_id: str, metrics: Sequence[Metric] = (), params: Sequence[Param] = (), tags: Sequence[RunTag] = ())`](https://mlflow.org/docs/latest/python_api/mlflow.tracking.html#mlflow.tracking.MlflowClient.log_batch) | <ul><li/>Metrics logged with `MLflow` in metrics tab.<li/>`MLflow` does not support logging text metrics</ul> |
# +
list_to_log = [1, 2, 3, 2, 1, 2, 3, 2, 1]
# Using AzureML
azureml_run.log_list("sample_list", list_to_log)
# Using MLflow
from mlflow.entities import Metric
from mlflow.tracking import MlflowClient
import time
metrics = [
Metric(key="sample_list", value=val, timestamp=int(time.time() * 1000), step=0)
for val in list_to_log
]
MlflowClient().log_batch(mlflow_run.info.run_id, metrics=metrics)
# -
# ## `azureml.core.Run.log_row()`
# | azureml.core API | MLFlow equivalent or proposed alternative | Notes |
# |-------------------------------|-------------------------------------------|-------|
# | `Run.log_row(self, name, description=None, **kwargs)` | [`mlflow.log_metrics(metrics: Dict[str, float], step: Optional[int] = None)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_metrics) | <ul><li/>Does not render in UI as a table.<li/>Does not support logging text values</li></ul> |
# +
# v1 SDK
azureml_run.log_row("sample_table", col1=5, col2=10)
# MLFlow
metrics = {"sample_table.col1": 5, "sample_table.col2": 10}
mlflow.log_metrics(metrics)
# -
# ## `azureml.core.Run.log_table()`
# | azureml.core API | MLFlow equivalent or proposed alternative | Notes |
# |-------------------------------|-------------------------------------------|-----------------|
# | `Run.log_table(name, value, description="")` | [`mlflow.log_metrics(metrics: Dict[str, float], step: Optional[int] = None)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_metrics) | <ul><li/>`MLflow`logs metrics as different metrics for each column<li/>Metrics logged with `MLflow` appear in the metrics tab but not as a table<li/>`MLflow` does not support logging text values.</ul> |
# | `Run.log_table(name, value, description="")` | [`mlflow.log_artifact(local_path: str, artifact_path: Optional[str] = None)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_artifact) | <ul><li/>Stored as an artifact<li>Does not appear in the metrics.</ul> |
# +
# Using AzureML
table = {"col1": [1, 2, 3], "col2": [4, 5, 6]}
azureml_run.log_table("table", table)
# Using mlflow.log_metrics
# Add a metric for each column prefixed by metric name. Similar to log_row
row1 = {"table.col1": 5, "table.col2": 10}
# To be done for each row in the table
mlflow.log_metrics(row1)
# Using mlflow.log_artifact
import json
with open("table.json", "w") as f:
json.dump(table, f)
mlflow.log_artifact("table.json")
# -
# ## `azureml.core.Run.log_accuracy_table()`
# | azureml.core API | MLFlow equivalent or proposed alternative | Notes |
# |-------------------------------|-------------------------------------------|-----------------|
# | `Run.log_accuracy_table(name, value, description="")` | [`mlflow.log_dict(dictionary: Any, artifact_file: str)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_dict)| <ul><li/>Dict logged with `MLflow` does not render in the Studio UI as accuracy table.<li/>Dict logged with `MLflow` does not appear in the metrics tab<li/>`MLflow` logs dict as an artifact<li/>`log_dict` is **experimental**</ul> |
# +
ACCURACY_TABLE = (
'{"schema_type": "accuracy_table", "schema_version": "v1", "data": {"probability_tables": '
+ "[[[114311, 385689, 0, 0], [0, 0, 385689, 114311]], [[67998, 432002, 0, 0], [0, 0, "
+ '432002, 67998]]], "percentile_tables": [[[114311, 385689, 0, 0], [1, 0, 385689, '
+ '114310]], [[67998, 432002, 0, 0], [1, 0, 432002, 67997]]], "class_labels": ["0", "1"], '
+ '"probability_thresholds": [0.52], "percentile_thresholds": [0.09]}}'
)
# Using AzureML
azureml_run.log_accuracy_table("v1_accuracy_table", ACCURACY_TABLE)
# Using MLflow
mlflow.log_dict(ACCURACY_TABLE, "mlflow_accuracy_table.json")
# -
# ## `azureml.core.Run.log_confusion_matrix()`
# | azureml.core API | MLFlow equivalent or proposed alternative | Notes |
# |-------------------------------|-------------------------------------------|-----------------|
# | `Run.log_confusion_matrix(name, value, description="")` | [`mlflow.log_dict(dictionary: Any, artifact_file: str)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_dict)| <ul><li/>Dict logged with `MLflow` does not render in the Studio UI as confusion matrix.<li/>Dict logged with `MLflow` does not appear in the metrics tab<li/>`MLflow` logs dict as an artifact<li/>`log_dict` is **experimental**</ul> |
# +
CONF_MATRIX = (
'{"schema_type": "confusion_matrix", "schema_version": "v1", "data": {"class_labels": '
+ '["0", "1", "2", "3"], "matrix": [[3, 0, 1, 0], [0, 1, 0, 1], [0, 0, 1, 0], [0, 0, 0, 1]]}}'
)
# v1 SDK
azureml_run.log_confusion_matrix("v1_confusion_matrix", json.loads(CONF_MATRIX))
# MLFlow
mlflow.log_dict(CONF_MATRIX, "mlflow_confusion_matrix.json")
# -
# ## `azureml.core.Run.log_predictions()`
# | azureml.core API | MLFlow equivalent or proposed alternative | Notes |
# |-------------------------------|-------------------------------------------|-----------------|
# | `Run.log_predictions(name, value, description="")` | [`mlflow.log_dict(dictionary: Any, artifact_file: str)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_dict)| <ul><li/>Dict logged with `MLflow` does not render in the Studio UI as predictions.<li/>Dict logged with `MLflow` does not appear in the metrics tab<li/>`MLflow` logs dict as an artifact<li/>`log_dict` is **experimental**</ul> |
# +
PREDICTIONS = (
'{"schema_type": "predictions", "schema_version": "v1", "data": {"bin_averages": [0.25,'
+ ' 0.75], "bin_errors": [0.013, 0.042], "bin_counts": [56, 34], "bin_edges": [0.0, 0.5, 1.0]}}'
)
# v1 SDK
azureml_run.log_predictions("test_predictions", json.loads(PREDICTIONS))
# MLFlow
mlflow.log_dict(PREDICTIONS, "mlflow_predictions.json")
# -
# ## `azureml.core.Run.log_residuals()`
# | azureml.core API | MLFlow equivalent or proposed alternative | Notes |
# |-------------------------------|-------------------------------------------|-----------------|
# | `Run.log_residuals(name, value, description="")` | [`mlflow.log_dict(dictionary: Any, artifact_file: str)`](https://mlflow.org/docs/latest/python_api/mlflow.html#mlflow.log_dict)| <ul><li/>Dict logged with `MLflow` does not render in the Studio UI as residuals.<li/>Dict logged with `MLflow` does not appear in the metrics tab<li/>`MLflow` logs dict as an artifact<li/>`log_dict` is **experimental**</ul> |
# +
RESIDUALS = (
'{"schema_type": "residuals", "schema_version": "v1", "data": {"bin_edges": [100, 200, 300], '
+ '"bin_counts": [0.88, 20, 30, 50.99]}}'
)
# v1 SDK
azureml_run.log_residuals("test_residuals", json.loads(RESIDUALS))
# MLFlow
mlflow.log_dict(RESIDUALS, "mlflow_residuals.json")
# -
# End the AzureML and MLflow runs
azureml_run.complete()
mlflow.end_run()
# ## Retreiving Run Info and Data with `MlFlow`
#
# You can access run information using `MLflow` through the run object's `data` and `info` properties. See [here](https://mlflow.org/docs/latest/python_api/mlflow.entities.html#mlflow.entities.Run) for more information on the `MLflow.entities.Run` object and the information it exposes.
#
# After run has completed, you can retrieve it using the [`MlFlowClient()`](https://mlflow.org/docs/latest/python_api/mlflow.tracking.html#mlflow.tracking.MlflowClient) (clickable link).
# +
from mlflow.tracking import MlflowClient
# Use MlFlow to retrieve the run that was just completed
client = MlflowClient()
finished_mlflow_run = MlflowClient().get_run(mlflow_run.info.run_id)
# -
# You can view the metrics, parameters, and tags for the run in the `data` field of the run object.
metrics = finished_mlflow_run.data.metrics
tags = finished_mlflow_run.data.tags
params = finished_mlflow_run.data.params
# **Note:** The metrics dictionary under `mlflow.entities.Run.data.metrics` will only have the **most recently logged value** for a given metric name. For example, if you log, in order, 1, then 2, then 3, then 4 to a metric called `sample_metric`, only 4 will be present in the metrics dictionary for the key `sample_metric`.
#
# To get all metrics logged for a particular metric name, you can use `MlFlowClient.get_metric_history()`.
# +
with mlflow.start_run() as multiple_metrics_run:
mlflow.log_metric("sample_metric", 1)
mlflow.log_metric("sample_metric", 2)
mlflow.log_metric("sample_metric", 3)
mlflow.log_metric("sample_metric", 4)
print(client.get_run(multiple_metrics_run.info.run_id).data.metrics)
print(client.get_metric_history(multiple_metrics_run.info.run_id, "sample_metric"))
# -
# You can view general information about the run, such as start time, run id, experiment id, etc. through the `info` field of the run object
run_start_time = finished_mlflow_run.info.start_time
run_experiment_id = finished_mlflow_run.info.experiment_id
run_id = finished_mlflow_run.info.run_id
# ## Retrieving Run Artifacts with `MLflow`
# To view the artifacts of a run, you can use [`MlFlowClient.list_artifacts()`](https://mlflow.org/docs/latest/python_api/mlflow.tracking.html#mlflow.tracking.MlflowClient.list_artifacts) (clickable link)
client.list_artifacts(finished_mlflow_run.info.run_id)
# To download an artifact, you can use [`MLFlowClient.download_artifacts()`](https://www.mlflow.org/docs/latest/python_api/mlflow.tracking.html#mlflow.tracking.MlflowClient.download_artifacts) (clickable link)
client.download_artifacts(finished_mlflow_run.info.run_id, "Azure.png")
# ## Searching Runs with `MLflow`
# You can use [`mlflow.search_runs()`](https://www.mlflow.org/docs/latest/python_api/mlflow.html#mlflow.search_runs) (clickable link) to query for runs programatically. The search API is a simplified
# version of the SQL WHERE clause and supports the following functionality:
# 1. query on metrics, params
# 1. query on tags
# 1. query on run metadata (for example, status)
# 1. query on runs of single experiment or multiple experiments
#
# The `search_runs()` call returns either:
# 1. By default, a `pandas.DataFrame`.
# 1. Optionally, a list. You can specify the list option by passing `"list"` as the value for the keyword argument `output_format`.
#
# +
from mlflow.entities import ViewType
## example: get list of runs in order of descending accuracy and return as a list, use:
runs = mlflow.search_runs(
experiment_ids="0",
filter_string="",
run_view_type=ViewType.ACTIVE_ONLY,
max_results=1,
order_by=["metrics.acc DESC"],
output_format="list",
)
# -
## example: get all active runs from experiments IDs 3, 4, and 17 that used a CNN model with 10 layers
query = "params.model = 'CNN' and params.layers = '10'"
runs = mlflow.search_runs(
experiment_ids=["3", "4", "17"],
filter_string=query,
run_view_type=ViewType.ACTIVE_ONLY,
)
| notebooks/mlflow/mlflow-v1-comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Eye-test visualization for the Carriean task
# +
# %matplotlib inline
import sys
import os.path
import numpy as np
import seaborn as sns
ROOT_DIR = '/home/areshytko/repos/interview/'
sys.path.append(ROOT_DIR)
from carribean.points_grid import PointsGridGraph, four_points_connectivity, eight_points_connectivity
from carribean.carribean import get_best_island, generate_problem, visualize
np.random.seed(100)
# -
H, W = 100, 50
problem = generate_problem(H, W)
# ## Input generated data:
_ = sns.heatmap(problem)
# ## Compute islands
connectivity = four_points_connectivity
graph = PointsGridGraph(input_map=problem, connectivity_strategy=connectivity)
components = graph.get_connected_components(min_component_size=20)
components.head()
# ## Visualize island scores:
_ = sns.heatmap(visualize(components, problem))
| carribean_eye_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # The Perceptron
#
# We just employed an optimization method - stochastic gradient descent, without really thinking twice about why it should work at all. It's probably worth while to pause and see whether we can gain some intuition about why this should actually work at all. We start with considering the E. Coli of machine learning algorithms - the Perceptron. After that, we'll give a simple convergence proof for SGD. This chapter is not really needed for practitioners but will help to understand why the algorithms that we use are working at all.
import mxnet as mx
from mxnet import nd, autograd
import matplotlib.pyplot as plt
import numpy as np
mx.random.seed(1)
# ## A Separable Classification Problem
#
# The Perceptron algorithm aims to solve the following problem: given some classification problem of data $x \in \mathbb{R}^d$ and labels $y \in \{\pm 1\}$, can we find a linear function $f(x) = w^\top x + b$ such that $f(x) > 0$ whenever $y = 1$ and $f(x) < 0$ for $y = -1$. Obviously not all classification problems fall into this category but it's a very good baseline for what can be solved easily. It's also the kind of problems computers could solve in the 1960s. The easiest way to ensure that we have such a problem is to fake it by generating such data. We are going to make the problem a bit more interesting by specifying how well the data is separated.
# +
# generate fake data that is linearly separable with a margin epsilon given the data
def getfake(samples, dimensions, epsilon):
wfake = nd.random_normal(shape=(dimensions)) # fake weight vector for separation
bfake = nd.random_normal(shape=(1)) # fake bias
wfake = wfake / nd.norm(wfake) # rescale to unit length
# making some linearly separable data, simply by chosing the labels accordingly
X = nd.zeros(shape=(samples, dimensions))
Y = nd.zeros(shape=(samples))
i = 0
while (i < samples):
tmp = nd.random_normal(shape=(1,dimensions))
margin = nd.dot(tmp, wfake) + bfake
if (nd.norm(tmp).asscalar() < 3) & (abs(margin.asscalar()) > epsilon):
X[i,:] = tmp[0]
Y[i] = 1 if margin.asscalar() > 0 else -1
i += 1
return X, Y
# plot the data with colors chosen according to the labels
def plotdata(X,Y):
for (x,y) in zip(X,Y):
if (y.asscalar() == 1):
plt.scatter(x[0].asscalar(), x[1].asscalar(), color='r')
else:
plt.scatter(x[0].asscalar(), x[1].asscalar(), color='b')
# plot contour plots on a [-3,3] x [-3,3] grid
def plotscore(w,d):
xgrid = np.arange(-3, 3, 0.02)
ygrid = np.arange(-3, 3, 0.02)
xx, yy = np.meshgrid(xgrid, ygrid)
zz = nd.zeros(shape=(xgrid.size, ygrid.size, 2))
zz[:,:,0] = nd.array(xx)
zz[:,:,1] = nd.array(yy)
vv = nd.dot(zz,w) + d
CS = plt.contour(xgrid,ygrid,vv.asnumpy())
plt.clabel(CS, inline=1, fontsize=10)
X, Y = getfake(50, 2, 0.3)
plotdata(X,Y)
plt.show()
# -
# Now we are going to use the simplest possible algorithm to learn parameters. It's inspired by the [Hebbian Learning Rule](https://en.wikipedia.org/wiki/Hebbian_theory) which suggests that positive events should be reinforced and negative ones diminished. The analysis of the algorithm is due to Rosenblatt and we will give a detailed proof of it after illustrating how it works. In a nutshell, after initializing parameters $w = 0$ and $b = 0$ it updates them by $y x$ and $y$ respectively to ensure that they are properly aligned with the data. Let's see how well it works:
# +
def perceptron(w,b,x,y):
if (y * (nd.dot(w,x) + b)).asscalar() <= 0:
w += y * x
b += y
return 1
else:
return 0
w = nd.zeros(shape=(2))
b = nd.zeros(shape=(1))
for (x,y) in zip(X,Y):
res = perceptron(w,b,x,y)
if (res == 1):
print('Encountered an error and updated parameters')
print('data {}, label {}'.format(x.asnumpy(),y.asscalar()))
print('weight {}, bias {}'.format(w.asnumpy(),b.asscalar()))
plotscore(w,b)
plotdata(X,Y)
plt.scatter(x[0].asscalar(), x[1].asscalar(), color='g')
plt.show()
# -
# As we can see, the model has learned something - all the red dots are positive and all the blue dots correspond to a negative value. Moreover, we saw that the values for $w^\top x + b$ became more extreme as values over the grid. Did we just get lucky in terms of classification or is there any math behind it? Obviously there is, and there's actually a nice theorem to go with this. It's the perceptron convergence theorem.
#
# ## The Perceptron Convergence Theorem
#
# **Theorem** Given data $x_i$ with $\|x_i\| \leq R$ and labels $y_i \in \{\pm 1\}$ for which there exists some pair of parameters $(w^*, b^*)$ such that $y_i((w^*)^\top x_i + b) \geq \epsilon$ for all data, and for which $\|w^*\| \leq 1$ and $b^2 \leq 1$, then the perceptron algorithm converges after at most $2 (R^2 + 1)/\epsilon^2$ iterations.
#
# The cool thing is that this theorem is *independent of the dimensionality of the data*. Moreover, it is *independent of the number of observations*. Lastly, looking at the algorithm itself, we see that we only need to store the mistakes that the algorithm made - for the data that was classified correctly no update on $(w,b)$ happened.
# As a first step, let's check how accurate the theorem is.
# +
Eps = np.arange(0.025, 0.45, 0.025)
Err = np.zeros(shape=(Eps.size))
for j in range(10):
for (i,epsilon) in enumerate(Eps):
X, Y = getfake(1000, 2, epsilon)
for (x,y) in zip(X,Y):
Err[i] += perceptron(w,b,x,y)
Err = Err / 10.0
plt.plot(Eps, Err, label='average number of updates for training')
plt.legend()
plt.show()
# -
# As we can see, the number of errors (and with it, updates) decreases inversely with the width of the margin. Let's see whether we can put this into equations. The first thing to consider is the size of the inner product between $(w,b)$ and $(w^*, b^*)$, the parameter that solves the classification problem with margin $\epsilon$. Note that we do not need explicit knowledge of $(w^*, b^*)$ for this, just know about its existence. For convenience, we will index $w$ and $b$ by $t$, the number of updates on the parameters. Moreover, whenever convenient we will treat $(w,b)$ as a new vector with an extra dimension and with the appropriate terms such as norms $\|(w,b)\|$ and inner products.
#
# First off, $w_0^\top w^* + b_0 b^* = 0$ by construction. Second, by the update rule we have that
#
# $$\begin{eqnarray}
# (w_{t+1}, b_{t+1})^\top (w^*, b^*) = & (w_t, b_t)^\top (w^*, b^*) + y_t \left(x_t^\top w^* + b^*\right)\\
# \geq & (w_t, b_t)^\top (w^*, b^*) + \epsilon \\
# \geq & (t+1) \epsilon
# \end{eqnarray}$$
#
# Here the first equality follows from the definition of the weight updates. The next inequality follows from the fact that $(w^*, b^*)$ separate the problem with margin at least $\epsilon$, and the last inequality is simply a consequence of iterating this inequality $t+1$ times. Growing alignment between the 'ideal' and the actual weight vectors is great, but only if the actual weight vectors don't grow too rapidly. So we need a bound on their length:
#
# $$\begin{eqnarray}
# \|(w_{t+1}, b_{t+1})\|^2 \leq & \|(w_t, b_t)\|^2 + 2 y_t x_t^\top w_t + 2 y_t b_t + \|(x_t, 1)\|^2 \\
# = & \|(w_t, b_t)\|^2 + 2 y_t \left(x_t^\top w_t + b_t\right) + \|(x_t, 1)\|^2 \\
# \leq & \|(w_t, b_t)\|^2 + R^2 + 1 \\
# \leq & (t+1) (R^2 + 1)
# \end{eqnarray}$$
#
# Now let's combine both inequalities. By Cauchy-Schwartz, i.e. $\|a\| \cdot \|b\| \geq a^\top b$ and the first inequality we have that $t \epsilon \leq (w_t, b_t)^\top (w^*, b^*) \leq \|(w_t, b_t)\| \sqrt{2}$. Using the second inequality we furthermore get $\|(w_t, b_t)\| \leq \sqrt{t (R^2 + 1)}$. Combined this yields
#
# $$t \epsilon \leq \sqrt{2 t (R^2 + 1)}$$
#
# This is a strange equation - we have a linear term on the left and a sublinear term on the right. So this inequality clearly cannot hold indefinitely for large $t$. The only logical conclusion is that there must never be updates beyond when the inequality is no longer satisfied. We have $t \leq 2 (R^2 + 1)/\epsilon^2$, which proves our claim.
#
# **Note** - sometimes the perceptron convergence theorem is written without bias $b$. In this case a lot of things get simplified both in the proof and in the bound, since we can do away with the constant terms. Without going through details, the theorem becomes $t \leq R^2/\epsilon^2$.
#
# **Note** - the perceptron convergence proof crucially relied on the fact that the data is actually separable. If this is not the case, the perceptron algorithm will diverge. It will simply keep on trying to get things right by updating $(w,b)$. And since it has no safeguard to keep the parameters bounded, the solution will get worse. This sounds like an 'academic' concern, alas it is not. The avatar in the computer game [Black and White](https://en.wikipedia.org/wiki/Black_%26_White_(video_game%29) used the perceptron to adjust the avatar. Due to the poorly implemented update rule the game quickly became unplayable after a few hours (as one of the authors can confirm).
# ## Stochastic Gradient Descent
#
# The perceptron algorithm also can be viewed as a stochastic gradient descent algorithm, albeit with a rather strange loss function: $\mathrm{max}(0, -y f(x))$. This is commonly called the hinge loss. As can be checked quite easily, its gradient is $0$ whenever $y f(x) > 0$, i.e. whenever $x$ is classified correctly, and gradient $-y$ for incorrect classification. For a linear function, this leads exactly to the updates that we have (with the minor difference that we consider $f(x) = 0$ as an example of incorrect classification). To get some intuition, let's plot the loss function.
f = np.arange(-5,5,0.1)
zero = np.zeros(shape=(f.size))
lplus = np.max(np.array([f,zero]), axis=0)
lminus = np.max(np.array([-f,zero]), axis=0)
plt.plot(f,lplus, label='max(0,f(x))')
plt.plot(f,lminus, label='max(0,-f(x))')
plt.legend()
plt.show()
# More generally, a stochastic gradient descent algorithm uses the following template:
# ```
# initialize w
# loop over data and labels (x,y):
# compute f(x)
# compute loss gradient g = partial_w l(y, f(x))
# w = w - eta g
# ```
# Here the learning rate $\eta$ may well change as we iterate over the data. Moreover, we may traverse the data in nonlinear order (e.g. we might reshuffle the data), depending on the specific choices of the algorithm. The issue is that as we go over the data, sometimes the gradient might point us into the right direction and sometimes it might not. Intuitively, on average things *should* get better. But to be really sure, there's only one way to find out - we need to prove it. We pick a simple and elegant (albeit a bit restrictive) proof of [Nesterov and Vial](http://dl.acm.org/citation.cfm?id=1377347).
#
# The situation we consider are *convex* losses. This is a bit restrictive in the age of deep networks but still quite instructive (in addition to that, nonconvex convergence proofs are a lot messier). For recap - a convex function $f(x)$ satisfies $f(\lambda x + (1-\lambda) x') \leq \lambda f(x) + (1-\lambda) f(x')$, that is, the linear interpolant between function values is *larger* than the function values in between. Likewise, a convex set $S$ is a set where for any points $x, x' \in S$ the line $[x,x']$ is in the set, i.e. $\lambda x + (1-\lambda) x' \in S$ for all $\lambda \in [0,1]$. Now assume that $w^*$ is the minimizer of the expected loss that we are trying to minimize, e.g.
#
# $$w^* = \mathrm{argmin}_w R(w) \text{ where } R(w) = \frac{1}{m} \sum_{i=1}^m l(y_i, f(x_i, w))$$
#
# Let's assume that we actually *know* that $w^*$ is contained in some convex set $S$, e.g. a ball of radius $R$ around the origin. This is convenient since we want to make sure that during optimization our parameter $w$ doesn't accidentally diverge. We can ensure that, e.g. by shrinking it back to such a ball whenever needed.
#
# Secondly, assume that we have an upper bound on the magnitude of the gradient $g_i := \partial_w l(y_i, f(x_i, w))$ for all $i$ by some constant $L$ (it's called so since this is often referred to as the Lipschitz constant). Again, this is super useful since we don't want $w$ to diverge while we're optimizing. In practice, many algorithms employ e.g. *gradient clipping* to force our gradients to be well behaved, by shrinking the gradients back to something tractable.
#
# Third, to get rid of variance in the parameter $w_t$ that is obtained during the optimization, we use the weighted average over the entire optimization process as our solution, i.e. we use $\bar{w} := \sum_t \eta_t w_t / \sum_t \eta_t$.
#
# Let's look at the distance $r_t := \|w_t - w^*\|$, i.e. the distance between the optimal solution vector $w^*$ and what we currently have. It is bounded as follows:
#
# $$\begin{eqnarray}
# \|w_{t+1} - w^*\|^2 = & \|w_t - w^*\|^2 + \eta_t^2 \|g_t\|^2 - 2 \eta_t g_t^\top (w_t - w^*) \\
# \leq & \|w_t - w^*\|^2 + \eta_t^2 L^2 - 2 \eta_t g_t^\top (w_t - w^*)
# \end{eqnarray}$$
#
# Next we use convexity of $R(w)$. We know that $R(w^*) \geq R(w_t) + \partial_w R(w_t)^\top (w^* - w_t)$ and moreover that the average of function values is larger than the function value of the average, i.e. $\sum_{t=1}^T \eta_t R(w_t) / \sum_t \eta_t \geq R(\bar{w})$. The first inequality allows us to bound the expected decrease in distance to optimality via
#
# $$\mathbf{E}[r_{t+1} - r_t] \leq \eta_t^2 L^2 - 2 \eta_t \mathbf{E}[g_t^\top (w_t - w^*)] \leq
# \eta_t^2 L^2 - 2 \eta_t \mathbf{E}[R[w_t] - R[w^*]]$$
#
# Summing over $t$ and using the facts that $r_T \geq 0$ and that $w$ is contained inside a ball of radius $R$ yields:
#
# $$-R^2 \leq L^2 \sum_{t=1}^T \eta_t^2 - 2 \sum_t \eta_t \mathbf{E}[R[w_t] - R[w^*]]$$
#
# Rearranging terms, using convexity of $R$ the second time, and dividing by $\sum_t \eta_t$ yields a bound on how far we are likely to stray from the best possible solution:
#
# $$\mathbf{E}[R[\bar{w}]] - R[w^*] \leq \frac{R^2 + L^2 \sum_{t=1}^T \eta_t^2}{2\sum_{t=1}^T \eta_t}$$
#
# Depending on how we choose $\eta_t$ we will get different bounds. For instance, if we make $\eta$ constant, i.e. if we use a constant learning rate, we get the bounds $(R^2 + L^2 \eta^2 T)/(2 \eta T)$. This is minimized for $\eta = R/L\sqrt{T}$, yielding a bound of $RL/\sqrt{T}$. A few things are interesting in this context:
#
# * If we are potentially far away from the optimal solution, we should use a large learning rate (the O(R) dependency).
# * If the gradients are potentially large, we should use a smaller learning rate (the O(1/L) dependency).
# * If we have a long time to converge, we should use a smaller learning rate, but not too small.
# * Large gradients and a large degree of uncertainty as to how far we are away from the optimal solution lead to poor convergence.
# * More optimization steps make things better.
#
# None of these insights are terribly surprising, albeit useful to keep in mind when we use SGD in the wild. And this was the very point of going through this somewhat tedious proof. Furthermore, if we use a decreasing learning rate, e.g. $\eta_t = O(1/\sqrt{t})$, then our bounds are somewhat less tight, and we get a bound of $O(\log T / \sqrt{T})$ bound on how far away from optimality we might be. The key difference is that for the decreasing learning rate we need not know when to stop. In other words, we get an anytime algorithm that provides a good result at any time, albeit not as good as what we could expect if we knew how much time to optimize we have right from the beginning.
#
# ## Next
# [Environment](../chapter02_supervised-learning/environment.ipynb)
# For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
| chapter02_supervised-learning/perceptron.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#导入math编程库(包)
import math
# 从math编程库(包)中调用exp函数。
math.exp(2.0)
# +
#从math编程库(包)里导入exp函数
from math import exp
# 直接使用函数名称调用exp,不需要声明math编程库(包)。
exp(2.0)
# +
#从math编程库(包)里导入exp函数,并且将exp重命名为ep
from math import exp as ep
ep(2.0)
# -
| Chapter_3/Section_3.8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 07. Ensemble Learning and Random Forests
# Aggregating the result of different predictors is called **Ensemble Learning**. Similarly to the phenomenon known as _wisdom of the crowd_, these aggregated models often turn out to be more effective than any individual one.
#
# **Note**: Ensemble methods work best when the predictors are as independent from one another as possible.
# ### Voting Classifiers
#
# A very simple way to create an even better classifier is to aggregate the predictions of each classifier and predict the class that gets the most votes (**hard voting**) or predict the class with the highest class probability, averaged over all the individual classifiers (**soft voting**) assuming all the classifiers outputs are probabilities.
# ### Bagging and Pasting
#
# The method we discussed above involves using different classifiers on the same dataset. Another approach is to use the same training algorithm for every
# predictor, but to train them on different random subsets of the training set.
#
# This second method has two variants:
# 1. **Bagging** (with replacement)
# 2. **Pasting** (without replacement)
#
# Generally, the net result is that the ensemble has a similar bias but a lower variance than a single predictor trained on the original training set.
# #### Out-of-bag Evaluation
#
# With bagging, some instances may be sampled several times for any given predictor, while others may not be sampled at all. Usually this means around 37% of instances never sampled (they are left **out-of-bag** so to speak). We can therefore use them to evaluate our predictors, since our predictor has never seen them in training.
#
# Simply add `oob_score=True` to the classifier.
# #### Random Patches and Random Subspaces
#
# Sometimes, especially when dealing with high dimensionality, it may be helpful to sample a subset of **features** rather than samples.
# * Sampling both training instances and features is called the **Random Patches** method
# * Keeping all training instances and sampling features is called the **Random Subspaces** method
# ### Random Forests
#
# A Random Forest is an ensemble of Decision Trees, generally trained using bagging and with `max_samples` = training set.
#
# The Random Forest algorithm introduces extra randomness when growing trees; instead of searching for the very best feature when splitting a node, it searches for the best feature among a random subset of features. This results in greater diversity > high bias / low variance.
# #### Extra-Trees
#
# It is possible to make trees even more random by also using random thresholds for each feature rather than searching for the best possible thresholds. We call this type of tree **Extremely Randomized Trees** (Extra).
#
# Once again, this trades more bias for a lower variance. It also makes Extra-Trees much faster to train than regular Random Forests since finding the best possible threshold for each feature at every node is one of the most time-consuming tasks of growing a tree.
# #### Feature Importance
#
# Another great property of Random Forest is that they make easy to measure the relative importance of each feature.
# This is very easy to see with an example:
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import load_iris
iris = load_iris()
rnd_clf = RandomForestClassifier(n_estimators=500, n_jobs=-1)
rnd_clf.fit(iris["data"], iris["target"])
for name, score in zip(iris["feature_names"], rnd_clf.feature_importances_):
print(name, score)
# -
# Therefore, Random Forests are very useful if we need a quick understanding of our features, potentially to perform further **feature selection**.
# ### Boosting
#
# The general idea of most boosting methods is to train predictors sequentially, each trying to correct its predecessor.
#
# We will cover two of them: **AdaBoost** (Adaptative Boosting) and **Gradient Boosting**.
# #### AdaBoost
#
# One way for a new predictor to correct its predecessor is to pay a bit more attention to the training instances that the predecessor underfitted.
#
# Similarly to Gradient Descent, this is an iterative process, with the main difference being that while with GD we had to tweak parameters of a single predictor here we add predictor to an ensemble, with a strong empashis on underfitting cases.
# More formally:
#
# 1. A first predictor is trained and its weighted error rate $r_1$ is computed on the training set
#
# $r_j = \displaystyle{\frac{\sum^{m}_{i=1} w^{(i)} \text{ for } \hat{y}_j^{(i)} \ne {y}_j^{(i)} }{\sum^{m}_{i=1} w^{(i)}}} $ where $\hat{y}_j^{(i)} $ is the $j^{th}$ predictor's prediction for the $i^{th}$ instance
#
# 2. Prediction weight: the higher the more accurate the predictor. 0 = random. Negative = less accurate than random.
#
# $a_j = \eta log \frac{1-r_j}{r_j}$
#
# 3. Weight update rule: the iterative process continues until the desired number of predictors is reached, or when a perfect predictor is found.
#
# $\begin{cases}
# w^{(i)} & \hat{y}_j^{(i)} = {y}_j^{(i)}\\
# w^{(i)} exp(\alpha_j) & \hat{y}_j^{(i)} \ne {y}_j^{(i)}
# \end{cases}$
# **Note**: the resulting $w^{(i)}$ will be normalized.
# 4. AdaBoost predictions
#
# $ \hat{y}(x) = argmax (k) \sum_{j=1}^{N} \alpha_j$
#
# $\text{ for } \hat{y}_j(x) = k$
# #### Gradient Boosting
#
# Instead of tweaking the instance weights at every iteration like AdaBoost does, this method tries to fit the new predictor to the **residual errors** made by the previous predictor.
#
# In order to find the optimal number of trees, you can use early stopping using `staged_predict()`. Here we train an ensemble of 120 trees, measure the validation error and pick the model with the optimal number of tree.
# +
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
X_train, X_val, y_train, y_val = train_test_split(X, y)
# gradient boosting regression trees
gbrt = GradientBoostingRegressor(max_depth=2, n_estimators=120)
gbrt.fit(X_train, y_train)
errors = [mean_squared_error(y_val, y_pred)
for y_pred in gbrt.staged_predict(X_val)]
bst_n_estimators = np.argmin(errors)
gbrt_best = GradientBoostingRegressor(max_depth=2,n_estimators=bst_n_estimators)
gbrt_best.fit(X_train, y_train)
# -
# Or we can stop if the validation error doesn't improve for X iterations in a row.
#
# Example when X = 5:
# +
gbrt = GradientBoostingRegressor(max_depth=2, warm_start=True)
min_val_error = float("inf")
error_going_up = 0
for n_estimators in range(1, 120):
gbrt.n_estimators = n_estimators
gbrt.fit(X_train, y_train)
y_pred = gbrt.predict(X_val)
val_error = mean_squared_error(y_val, y_pred)
if val_error < min_val_error:
min_val_error = val_error
error_going_up = 0
else:
error_going_up += 1
if error_going_up == 5:
break # early stopping
# -
# **Note**: a very optimized implementation of Gradient Boosting is available in the library `XGBoost`.
# ### Stacking
# Stacked Generalization (or Stacking) is based on the idea that we can train a model to perform the result aggregation (instead of simple function such as hard / soft voting).
#
# We call this meta-model a **blending predictor**. To train the blender, a common approach is to use a hold-out set and proceed as follows:
#
# 1. Train first layer predictors on first subset of training set
# 2. Use our X first-layer predictors to conduct predictions on the hold out set they have never seen before
# 3. This X predictions will form a new X-dimentional dataset that we will use to train our blender predictor
# 4. [Optional] Training different type of predictors (Lin Reg, Random Forests etc.)
| .ipynb_checkpoints/07.Ensemble_Learning_and_Random_Forests-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
df = pd.read_csv("weather_data.csv")
df
new_df = df.replace(-99999,np.NaN)
new_df
new_df = df.replace([-99999,-88888],np.NaN)
new_df
new_df = df.replace({
"temperature":-99999,
"windspeed":[-99999,-88888],
"event":"0"
},np.NaN)
new_df
new_df = df.replace({
-99999:np.NaN,
-88888:np.NaN,
"0":"Sunny"
})
new_df
new_df = df.replace({
"temperature":"[A-Za-z]",
"windspeed":"[A-Za-z]",
},"",regex=True)
new_df
df = pd.DataFrame({
'score': ['exceptional','average', 'good', 'poor', 'average', 'exceptional'],
'student': ['rob', 'maya', 'parthiv', 'tom', 'julian', 'erica']
})
df
new_df = df.replace(["exceptional","good","average","poor"],[4,3,2,1])
new_df
| path_of_ML/Pandas/Replace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Fingerprint bit coverage and variability
#
# We check the coverage and variability of fingerprint bit positions across all fingerprints in our dataset.
#
# - Investigate missing bits across fingerprints
# - Investigate bit variability across fingerprints (standard deviation)
# - Get top X bit positions with no/high standard deviation
# %load_ext autoreload
# %autoreload 2
# +
from pathlib import Path
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from kissim.encoding import FingerprintGenerator
from src.paths import PATH_RESULTS
# -
plt.style.use("seaborn")
HERE = Path(_dh[-1]) # noqa: F821
RESULTS = PATH_RESULTS / "all"
# ## Load fingerprints
#
# Let's load our pre-calculated fingerprint saved in a _json_ file as `FingerprintGenerator` object.
fingerprint_generator = FingerprintGenerator.from_json(RESULTS / "fingerprints_clean.json")
fingerprint_generator_normalized = FingerprintGenerator.from_json(
RESULTS / "fingerprints_normalized.json"
)
if len(fingerprint_generator.data) == len(fingerprint_generator_normalized.data):
n_fingerprints = len(fingerprint_generator_normalized.data)
else:
raise ValueError("Number of unnormalized/normalized fingerprints not the same.")
print(f"Number of fingerprints: {n_fingerprints}")
# ## Missing bits across fingerprints
def nan_bit_ratio(fingerprint_generator, feature_group):
if feature_group == "physicochemical":
features = fingerprint_generator.physicochemical_exploded
elif feature_group == "distances":
features = fingerprint_generator.distances_exploded
elif feature_group == "moments":
features = fingerprint_generator.moments_exploded
else:
raise KeyError("Feature group unknown.")
features = features.unstack()
features_nan = features.isnull().sum(axis=0) / len(features)
features_nan = features_nan.unstack(0)
return features_nan
# 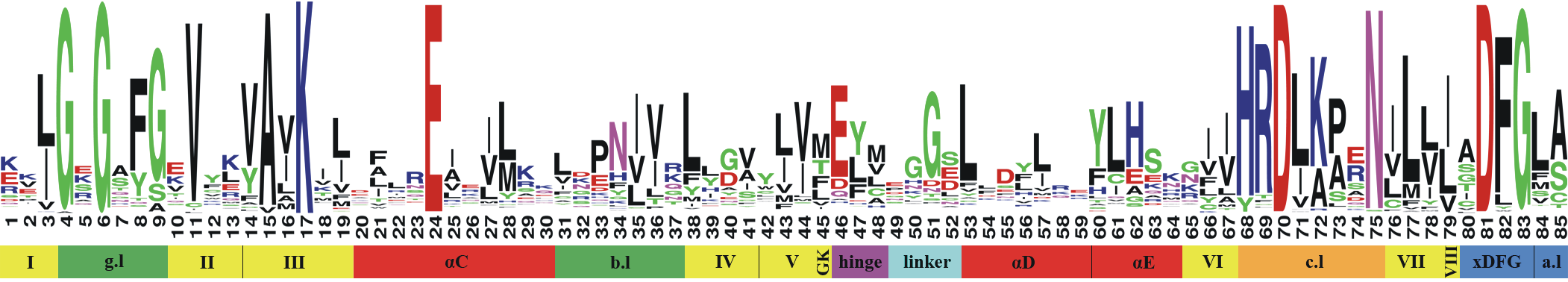
# ### Missing physicochemical bits
feature_group = "physicochemical"
features_nan = nan_bit_ratio(fingerprint_generator_normalized, feature_group)
features_nan
features_nan.describe()
# #### Heatmap (for an overview)
plt.figure(figsize=(20, 3))
sns.heatmap(
features_nan.transpose(),
cmap="Blues",
cbar_kws={"label": f"Ratio of NaN bits ({n_fingerprints} fingerprints)"},
vmin=0,
vmax=1,
)
plt.xlabel("KLIFS pocket residue index")
plt.ylabel("Feature");
# 
# #### Barplot (for concrete values)
ax = features_nan.plot(kind="bar", figsize=(22, 5), width=1, colormap="Set1")
ax.set_ylabel(f"Ratio of NaN bits ({n_fingerprints} fingerprints)");
# __Observations__
#
# - _Side chain orientation_: Gly-rich positions (e.g. 4, 6, 9, 83) have high ratios of missing side orientation values because Gly has no side chain.
# - _All feature types_: Higher ratios of missing bits for pocket positions with high ratio of missing residues (e.g. 50, 4-9, 83-58). Check out pattern resemblance with plot on "Missing residues in the kinase binding site" in [this notebook](https://github.com/volkamerlab/kissim_app/blob/master/notebooks/dataset/explore_dataset.ipynb).
# ### Missing distance bits
feature_group = "distances"
features_nan = nan_bit_ratio(fingerprint_generator_normalized, feature_group)
features_nan
features_nan.describe()
# #### Heatmap (for an overview)
plt.figure(figsize=(20, 2))
sns.heatmap(
features_nan.transpose(),
cmap="Blues",
cbar_kws={"label": f"Ratio of NaN bits ({n_fingerprints} fingerprints)"},
vmin=0,
vmax=1,
)
plt.xlabel("KLIFS pocket residue index")
plt.ylabel("Feature");
# #### Barplot (for concrete values)
ax = features_nan.plot(kind="bar", figsize=(22, 5), width=1, colormap="Set1")
ax.set_ylabel(f"Ratio of NaN bits ({n_fingerprints} fingerprints)");
# __Observations__
#
# - As before for the physicochemical bits, the pattern here resembles the missing residue pattern.
# ### Missing moments bits
feature_group = "moments"
features_nan = nan_bit_ratio(fingerprint_generator_normalized, feature_group)
features_nan
features = fingerprint_generator_normalized.moments_exploded
structures_nan = features[features.isna().any(axis=1)]
print(
f"Number of structures with at least one NaN moment: "
f"{len(structures_nan.index.get_level_values(0).unique())}"
)
structures_nan
# __Observations__
#
# - About 70 structures show at least one empty moment, matching well with the number of structures for which one or more subpocket centers could not be determined and thus, no distance distribution and no corresponding moments could be inferred. Check out "Number of structures that will have no subpocket centers" in [this notebook](https://github.com/volkamerlab/kissim_app/blob/master/notebooks/dataset/explore_dataset.ipynb).
# ## Bit variability across fingerprints
def bit_variability(fingerprint_generator, feature_group):
if feature_group == "physicochemical":
features = fingerprint_generator.physicochemical_exploded
elif feature_group == "distances":
features = fingerprint_generator.distances_exploded
elif feature_group == "moments":
features = fingerprint_generator.moments_exploded
else:
raise KeyError("Feature group unknown.")
features = features.unstack()
features_std = features.std()
features_std = features_std.unstack(0)
return features_std
# ### Physicochemical bit variability
features_std = bit_variability(fingerprint_generator_normalized, "physicochemical")
features_std
features_std.describe()
plt.figure(figsize=(20, 3))
sns.heatmap(
features_std.transpose(),
cmap="Blues",
cbar_kws={"label": f"Bit standard deviation ({n_fingerprints} fingerprints)"},
)
plt.xlabel("KLIFS pocket residue index")
plt.ylabel("Feature");
# __Observations__
#
# - Highest variability for HBD, lowest for aromatic and aliphatic
# - Conserved regions show as expected rigid size/pharmacophoric feature variability (e.g. 4, 6, 17, 24, 70, 75, 81-83).
#
# 
# ### Distances bit variability
features_std = bit_variability(fingerprint_generator_normalized, "distances")
features_std
features_std.describe()
plt.figure(figsize=(20, 3))
sns.heatmap(
features_std.transpose(),
cmap="Blues",
cbar_kws={"label": f"Bit standard deviation ({n_fingerprints} fingerprints)"},
)
plt.xlabel("KLIFS pocket residue index")
plt.ylabel("Feature");
# __Observations__
#
# - Higher variability for
# - residues 6-8 (part of the G-loop, known for loop flexibility)
# - residues 20-24 (part of aC-helix, known for conformational changes)
# - residues 82-85 (part of DFG-loop and activation loop, know for conformational changes)
# - residues 59-61 (part of aE-helix, ???)
# - residue 33 w.r.t. hinge region (???)
# 
# 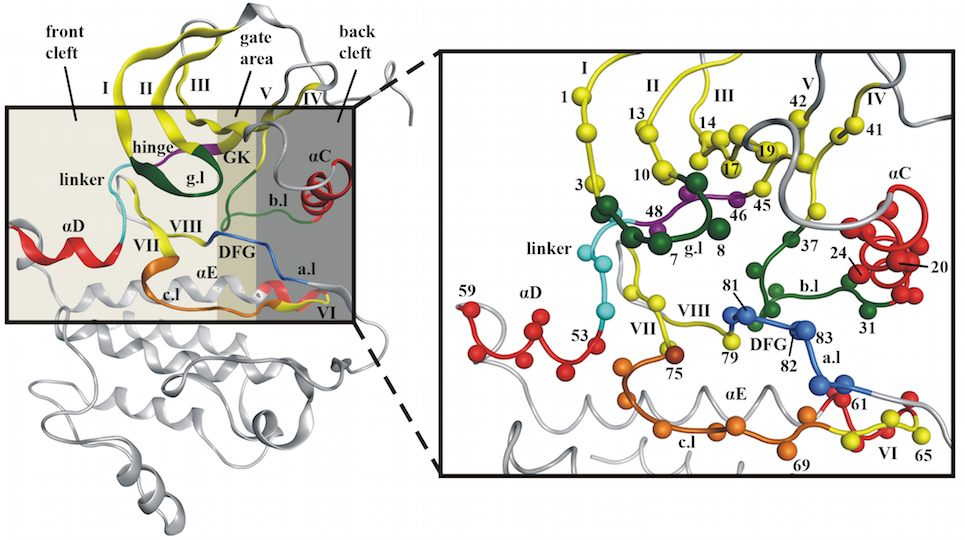
# ### Moments bit variability
features_std = bit_variability(fingerprint_generator_normalized, "moments")
features_std
plt.figure(figsize=(3, 3))
sns.heatmap(
features_std.transpose(),
cmap="Blues",
cbar_kws={"label": f"Bit standard deviation ({n_fingerprints} fingerprints)"},
)
plt.xlabel("KLIFS pocket residue index")
plt.ylabel("Feature");
# ## Top X bit with no/low/high variability
#
# Get per-bit standard deviation for all feature types and check for bits with no/low/high variability.
bits_std = []
for feature_group in ["physicochemical", "distances", "moments"]:
bits_std.append(bit_variability(fingerprint_generator_normalized, feature_group).unstack())
bits_std = pd.concat(bits_std)
bits_std.index.names = ["feature", "residue_ix"]
print(f"Number of bits with std: {len(bits_std)}")
# ### No variability
bits_std_no_variability = bits_std[bits_std == 0]
bits_std_no_variability.sort_index(level=1)
print(
f"Number of bits with no effect because they are always "
f"(or almost always) the same: {len(bits_std_no_variability)}"
)
# ### Low variability
bits_std_low_variability = bits_std[bits_std != 0].sort_values().head(30)
bits_std_low_variability.sort_index(level=1)
# ### High variability
# + tags=[]
bits_std_high_variability = bits_std[bits_std != 0].sort_values().tail(30)
bits_std_high_variability.sort_index(level=1)
| notebooks/004_fingerprints/005_fingerprint_bit_coverage_variability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Continuación Métodos Numéricos de Integración de Ecuaciones Diferenciales Ordinarias
# **Taller de Modelación Numérica** <br>
# Facultad de Ciencias, UNAM <br>
# Semestre 2022-2
#
# En este notebook veremos ejemplos de soluciones numércias a dos ecuaciones diferenciales ordinarias (EDO) el método Runge-Kutta orden 4 y utilizando la función `solve_ivp` del paquete scipy.
#
# Al final del notebook encontrarás el ejercicio de tarea correspondiente. Entrega tu solución en un notebook junto con los ejercicios del notebook 04_soluciones_EDOs.ipynb en la carpeta de Classroom con el nombre `apellido_nombre_tarea04_EDOs.ipynb`.
# ### Ejemplo 1: Runge-Kutta 4
#
# Resolvamos la ecuación
# $$y' = y - t^2 + 1$$
#
# con condición inicial $$y(0) = 0.5$$
from math import ceil
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ### Ejemplo 1 Métodos de Runge-Kutta
#
# Ahora resolvamos la misma ecuación que en el ejemplo 1 del método de Euler usando Runge-Kutta 4: $$ y'=y - t^2 + 1$$ con $$y(0) = 0.5$$
# +
h = 0.01 # Paso de tiempo
t = 0
y = 0.5
print('Paso 0: t = %6.3f, y = %18.15f\n' %(t, y))
TRK1 = np.empty(int(5/h))
YRK1 = np.empty(int(5/h))
for ii in range(int(5/h)):
k1 = h * (y - t**2 + 1) # h*f(t,y);
k2 = h * ((y+k1/2) - (t+h/2)**2 + 1) # h*f(t+h/2, y+k1/2);
k3 = h * ((y+k2/2) - (t+h/2)**2 + 1) # h*f(t+h/2, y+k2/2);
k4 = h * ((y+k3) - (t+h)**2 + 1) # h*f(t+h, y+k3);
y = y + (k1 + 2*k2 + 2*k3 + k4)/6
t = t + h
TRK1[ii] = t
YRK1[ii] = y
print('Paso %d: t = %6.3f, y = %18.15f\n' % (ii, t, y))
# -
# Sol. analítica
YARK1 = 2*TRK1 - np.exp(TRK1)/2 + TRK1**2 + 1
# +
fig,(ax0,ax1) = plt.subplots(1,2,figsize=(12,4))
ax0.plot(TRK1,YRK1,'-og', label='Runge-Kutta 4')
ax0.plot(TRK1,YARK1,'-*m',label='Analítica')
ax0.set_xlabel('t')
ax0.set_ylabel('y')
ax0.legend()
ax1.stem(TRK1,YARK1-YRK1,'b',use_line_collection=True)
ax1.set_xlabel('t')
ax1.set_ylabel(r'$y_a-y_{rk}$')
ax1.set_title('Diferencia entre RK 4 y solución analítica')
# -
# ### Ejemplo 2 Runge-Kutta
#
# Ahora resolvamos la misma ecuación que en el ejemplo 2 del método de Euler usando Runge-Kutta 4:
# $$y'= sin(t)$$
# $$y(0) = -1.$$
h=0.01; # este es el incremento de tiempo o del_t
t = 0; # tiempo inicial o t0
y = -1; # valor de y en t0
print('Paso 0: t = %6.3f, y = %18.15f\n' %(t, y))
def RK_f(a,b):
return(np.sin(a))
# +
TRK2 = np.empty(ceil(4*np.pi/h))
YRK2 = np.empty(ceil(4*np.pi/h))
for ii in range(ceil(4*np.pi/h)):
k1 = h * RK_f(t, y) # h*f(t,y);
k2 = h * RK_f(t+h/2, y+k1/2) # h*f(t+h/2, y+k1/2);
k3 = h * RK_f(t+h/2, y+k2/2) # h*f(t+h/2, y+k2/2);
k4 = h * RK_f(t+h, y+k3) # h*f(t+h, y+k3);
y = y + (k1 + 2*k2 + 2*k3 + k4)/6
t = t + h
TRK2[ii] = t
YRK2[ii] = y
print('Paso %d: t = %6.3f, y = %18.15f\n' %(ii, t, y))
# -
YARK2 = -np.cos(TRK2) # solución analítica
# +
fig,(ax0,ax1) = plt.subplots(1,2,figsize=(12,4))
ax0.plot(TRK2,YRK2,'-og', label='Runge-Kutta 4')
ax0.plot(TRK2,YRK2,'-*m',label='Analítica')
ax0.set_xlabel('t')
ax0.set_ylabel('y')
ax0.legend()
ax1.stem(TRK2,YARK2-YRK2,'b',use_line_collection=True)
ax1.set_xlabel('t')
ax1.set_ylabel(r'$y_a-y_{rk}$')
ax1.set_title('Diferencia entre RK4 y solución analítica')
# -
# ### Funciones de python para resolver Problemas de valores iniciales (IVP)
#
# EL paquete scipy de python contiene módulos para resolver EDOs con rutinas mucho más precisas que el método de Euler.
# La función recomendada para resolver problemas como los ejemplos anteriores (ojo, no es la única) es [`solve_ivp`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.integrate.solve_ivp.html?highlight=s).
#
# Como su nombre lo indica, esta función resuelve numéricamente el problema de valores iniciales para un sistema de EDOs de la forma
# $$y' = f(t, y),$$
# $$y(t_0) = y_0$$
#
# Donde $t$ es la variable independiente, $y(t)$ es un vector de N dimensiones y $f(t,y)$ es una función vector de N dimensiones que determina las ecuaciones diferenciales.
#
# El método que esta función usa por default es Runge-Kutta 4, pero se puede elegir de entre 5 métodos distintos. Consulta la [documentación](https://docs.scipy.org/doc/scipy/reference/generated/scipy.integrate.solve_ivp.html?highlight=s) para más detalles acerca de `solve_ivp`.
# +
# %% Importamos la función solve_ivp del módulo integrate del paquete scipy
from scipy.integrate import solve_ivp
# %% Definimos una función derivada (en este caso la misma que en el ejemplo 1)
def f(t, y):
dydt = y-t**2+1
return dydt
# %% Definimos el intervalo de tiempo y el valor inicial de la función y(t0).
t_intervalo = np.linspace(0, 5, 50) # un vector de dimensiones (1,50) que comienza en cero y va hasta 5
y_0 = np.array([0.5]) # 1D
# %% Resolvemos la ecuación diferencial
sol = solve_ivp(f, [t_intervalo[0], t_intervalo[-1]], y_0, t_eval=t_intervalo)
# -
# Comparemos los métodos:
fig,ax = plt.subplots(1,1, figsize=(5,4))
ax.plot(sol.t, sol.y[0],'b-',label='solve_ivp RK4',linewidth=3)
ax.plot(TRK1,YRK1,'r-', label='RK4')
ax.set_ylabel('y')
ax.set_xlabel('t')
ax.legend()
# ### Ejercicios de la Tarea 4 (parte 2)
#
# 1. Considera la ecuación diferencial $y’ = cos(t) + sin(t)$ con $y(t=0) = 1$ y $h=0.1$. Escribe un código para calcular la solución con el método de Runge-Kutta de orden 4 para el intervalo de 0 a $2\pi$.
#
# 2. Considera la ecuación diferencial $$y'=\frac{2y}{t}+t^2e^t$$ con $y(-2) = -10$..
#
# a) Calcula la solución analítica. Puedes utilizar algún programa en línea. <br>
# b) Escribe un código para calcular la solución con el método de Euler en el intervalo $\left[-2, 0\right)$.<br>
# c) Escribe un código para calcular la solución con el método de Runge-Kutta 2 en el intervalo $\left[-2, 0\right)$.<br>
# d) Calcula también la solución usando el método de Runge-Kutta 4 en el mismo intervalo.<br>
# e) En una figura muestra las soluciones analítica, de Euler y Runge-Kutta 2 y 4, con la leyenda correspondiente.<br>
# f) En otra figura grafica los errores de cada método numérico al compararlos con la solución analítica. <br>
# NOTA: Usar un paso de tiempo: h=0.1
| 02_sols_EDO/05_soluciones_EDOs_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# + [markdown] slideshow={"slide_type": "-"}
# # 2017-07-24 DeLong Econ 101b Spring 2018 Problem Sets Scratch...
# -
# ## Setting Up the Environment
# %matplotlib inline
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
from pandas import DataFrame, Series
from datetime import datetime
import pandas as pd
import plotly
plotly.tools.set_credentials_file(username='', api_key='')
import plotly.plotly as py
import plotly.graph_objs as go
from plotly.graph_objs import Scatter
sns.set(color_codes = True)
import statsmodels.formula.api as smf
# ## Chapter 1: Introduction to Macroeconomics
# #### Suppose a quantity grows at a steady proportional rate of 3% per year. How long will it take to double? Quadruple? Grow 1024-fold?
# #### Suppose we have a quantity x(t) that varies over time following the equation: dx(t)/dt = -(0.06)x + 0.36.
#
# Without integrating the equation
#
# 1. Tell me what the long-run steady-state value of x--that is, the limit of x as t approaches in infinity--is going to be.
# 2. Suppose that the value of x at time t=0, x(0), equals 12. Once again, without integrating the equation, tell me how long it will take x to close half the distance between its initial value of 12 and its steady-state value.
# 3. How long will it take to close 3/4 of the distance?
# 4. 7/8 of the distance?
# 5. 15/16 of the distance?
#
# Now you are allowed to integrate dx(t)/dt = -(0.06)x + 0.36.
#
# 1. Write down and solve the indefinite integral.
# 2. Write down and solve the definite integral for the initial condition x(0) = 12.
# 3. Write down and solve the definite integral for the initial condition x(0)=6.
# #### Suppose we have a quantity z = (x/y)<sup>β</sup>
#
# Suppose x is growing at 4% per year and that β=1/4:
#
# 1. How fast is z growing if y is growing at 0% per year?
# 2. If y is growing at 2% per year?
# 3. If y is growing at 4% per year?
# #### Rule of 72
#
# 1. If a quantity grows at about 3% per year, how long will it take to double?
# 2. If a quantity shrinks at about 4% per year, how long will it take it to halve itself?
# 3. If a quantity doubles five times, how large is it relative to its original value?
# 4. If a quantity halves itself three times, how large is it relative to its original value?
# #### Why do DeLong and Olney think that the interest rate and the level of the stock market are important macroeconomic variables?
# #### What are the principal flaws in using national product per worker as a measure of material welfare? Given these flaws, why do we use it anyway?
# #### What is the difference between the nominal interest rate and the real interest rate? Why do DeLong and Olney think that the real interest rate is more important?
# ## Chapter 2: Measuring the Macroeconomy
# #### National Income and Product Accounting
#
# Explain whether or not, why, and how the following items are included in the calculations of national product:
#
# 1. Increases in business inventories.
# 2. Fees earned by real estate agents on selling existing homes.
# 3. Social Security checks written by the government.
# 4. Building of a new dam by the Army Corps of Engineers.
# 5. Interest that your parents pay on the mortgage they have on their house.
# 6. Purchases of foreign-made trucks by American residents
# #### In or Out of National Product? And Why
#
# Explain whether or not, why, and how the following items are included in the calculation of national product:
#
# 1. The sale for $25,000 of an automobile that cost $20,000 to manufacture that had been produced here at home last year and carried over in inventory.
# 2. The sale for $35,000 of an automobile that cost $25,000 to manufacture newly- made at home this year.
# 3. The sale for $45,000 of an automobile that cost $30,000 to manufacture that was newly-made abroad this year and imported.
# 4. The sale for $25,000 of an automobile that cost $20,000 to manufacture that was made abroad and imported last year.
# #### In or Out of National Product? And Why II
#
# Explain whether or not, why, and how the following items are included in the calculation of GDP:
#
# 1. The purchase for $500 of a dishwasher produced here at home this year.
# 2. The purchase for $500 of a dishwasher made abroad this year.
# 3. The purchase for $500 of a used dishwasher.
# 4. The manufacture of a new dishwasher here at home for $500 of a dishwasher that
# then nobody wants to buy.
# #### Components of National Income and Product
#
# Suppose that the appliance store buys a refrigerator from the manufacturer on December 15, 2018 for $600, and that you then buy that refrigerator on January 15, 2019 for $1000:
#
# 1. What is the contribution to GDP in 2018?
# 2. How is the refrigerator accounted for in the NIPA in 2019?
# 3. What is the contribution to GDP in 2018?
# 4. How is the refrigerator accounted for in the NIPA in 2019?
# #### Estimating National Product
#
# The Bureau of Economic Analysis measures national product in two different ways: as total expenditure on the economy’s output of goods and services and as the total income of everyone in the economy. Since – as you learned in earlier courses – these two things are the same, the two approaches should give the same answer. But in practice they do not.
#
# For the period 2006:Q4–2012:Q4, get quarterly data on real GDP measured on the expenditure side (referred to in the National Income and Product Accounts as “Real Gross Domestic Product, chained dollars” – these are the headline GDP numbers you will see discussed in the news) and real GDP measured on the income side (referred to as “Real
# Gross Domestic Income, chained dollars”).
#
# 1. Describe where you found the data.
# 2. Compute the growth rate at an annual rate of each of the two series by quarter for
# 2007:Q1–2012:Q4.
# 3. Describe any two things you see when you compare the two series that you find
# interesting, and explain why you find them interesting.
# #### Calculating Real Magnitudes:
#
# 1. When you calculate real national product, do you do so by dividing nominal national product by the price level or by subtracting the price level from nominal national product?
# 2. When you calculate the real interest rate, do you do so by dividing the nominal interest rate by the price level or by subtracting the inflation rate from the nominal interest rate?
# 3. Are your answers to (a) and (b) the same? Why or why not?
# #### What, roughly, was the highest level the U.S. unemployment rate reached in:
#
# 1. the 20th century?
# 2. the past fifty years?
# 3. the twenty years before 2006?
# 4. Given your answers to (1) through (3), Do you think there is a connection between your answer to the qeustion above and the fact that Federal Reserve Chair <NAME> received a five-minute standing ovation at the end of the first of many events marking his retirement in 2005?
# #### The State of the Labor Market
#
# 1. About how many people lose or quit their jobs in an average month?
# 2. About how many people get jobs in an average month?
# 3. About how many people are unemployed in an average month?
# 4. About how many people are at work in an average month?
# 5. About how many people are unemployed now?
# #### National Income Accounting:
#
# 1. What was the level of real GDP in 2005 dollars in 1970?
# 2. What was the rate of inflation in the United States in 2000?
# 3. Explain whether or not, how, and why the following items are included in the calculation of GDP: (i) rent you pay on an apartment, (ii) purchase of a used textbook, (iii) purchase of a new tank by the Department of Defense, (iv) watching an advertisement on youtube.
# ## Chapter 3: Thinking Like an Economist
# #### In what sense can a line on a graph "be" an equation?
# ## Chapter 4: Theories of Economic Growth
# #### Production Functions
#
# 1. Suppose α = 1/2, E=1, L=100, and K=64; what is output per worker Y/L?
# 2. Suppose α = 1/2, E=3, L=196, and K=49; what is output per worker Y/L? € c. If both capital K and labor L triple, what happens to total output Y?
# 3. Holding E=1, suppose that capital per worker increases from 4 to 8 and then from 8 to 12. What happens to output per worker?
# #### Budget Deficits
#
# Consider an economy in which the depreciation rate is 2% per year, the rate of population increase is 1% per year, the rate of technological progress is 1% per year, and the private savings rate is 17% of GDP. Suppose that the government increases its budget deficit--which had been at 1% of GDP for a long time--to 5% of GDP and keeps it there indefinitely.
#
# 1. What is the effect of this shift on the economy's steady-state capital-output ratio? b. What is the effect of this shift on the economy's steady state growth path for output per worker?
# 2. Suppose that your forecast of output per worker 20 years in the future had been $120,000. What is your new forecast of output per worker twenty years hence?
# 3. Suppose that environmental regulations lead to a diversion of investment spending from investments that boost the capital stock to investments that replace polluting with less-polluting capital. In our standard growth model, what would be the consequences of such a diversion for the economy's capital-output ratio and for its balanced-growth path? Would it make sense to say that these environmental regulations diminished the economy's wealth?
# #### Savings Rates
#
# Consider an economy in which the depreciation rate is 4% per year, the rate of population increase is 1% per year, the rate of technological progress is 2% per year, and the national savings rate is 21% of GDP. Suppose that thrift-promoting policies raise the savings rate to 24.5% of GDP and keep it there indefinitely.
#
# 1. Calculate—that is write an equation for it as a function of time—what the level of output per worker would have been had the savings rate remained at 21%, and had the economy initially been on its steady-state growth path.
# 2. Calculate—that is write an equation for it as a function of time—what the level of output per worker will have be now that the savings rate has increased.
# #### Population Growth
#
# Many project that by the middle of the twenty-first century the population of the United States will be stable. Using the Solow growth model, what would the qualitative effects of such a downward shift in the growth rate of the labor force be on the growth of output per worker and to the growth of total output? (Consider both the effect of zero population growth on the steady-state growth path, and the transition from the "old" positive population growth to the "new" zero population growth steady-state growth path.)
# #### Long Run Models
#
# We have cautioned you that the Solow growth model is—especially in its focus-on- the-steady-state-growth-path version—a long run model. What do we mean by that?
# #### Doubling Capital per Worker
#
# Consider an economy with the production function: Y=K^α(EL)^(1−α) on its steady-state balanced-growth path:
#
# 1. Suppose α = 1/2, E=1, L=100, and K=64; what is output per worker Y/L?
# 2. Suppose α = 1/2, E=3, L=196, and K=49; what is output per worker Y/L?
# 3. If both capital K and labor L triple, what happens to total output Y?
# 4. Holding E=1, suppose that capital per worker increases from 4 to 8 and then from
# 8 to 12. What happens to output per worker?
# #### Walking Up the Production Function
#
# Consider an economy with the production function: Y=K^α(EL)^(1−α) on its steady-state balanced-growth path:
#
# 1. Suppose α = 1/3, E=1, L=100, and K=64; what is output per worker Y/L?
# 2. Suppose α = 1/3, E=3, L=196, and K=49; what is output per worker Y/L?
# 3. If both capital K and labor L double, what happens to total output Y?
# 4. Holding E=1, suppose that capital per worker increases from 1 to 4 and then from
# 4 to 9. What happens to output per worker?
# #### Balanced Growth Path
#
# Suppose that an economy's production function is Y = K^0.5(EL)^0.5; suppose further that the savings rate s is 40 percent of GDP, that the depreciation rate δ is 4 percent per year, the population growth rate n is 0, and the rate of growth g of the efficiency of the labor force is 2 percent per year.
#
# 1. What is the steady-state balanced-growth capital-output ratio?
# 2. How fast does output per worker grow along the steady-state balanced-growth
# path?
# 3. How fast does total output grow along the steady-state balanced-growth path?
# 4. Suppose that all variables are the same save the production function, which instead is: K^0.8(EL)^0.2; how would your answers be different? Why would your answers are different?
# ## Chapter 5: Reality of Economic Growth: History
# #### Escape from the Malthusian Trap
#
# Suppose somebody who hasn't taken any economics courses were to ask you why humanity escaped from the Malthusian trap--of very low standards of living and slow population growth rates that nevertheless put pressure on available natural resources and kept output per worker from rising--in which humanity found itself between the year 8000 B.C.E. and 1800. What answer would you give? (One paragraph only, please!)
# #### Malthusian Economy
#
# Suppose—in the Malthusian model with natural resources—that population growth depends on the level of output per worker, so that:
#
# n = (.00002) x [(Y/L) - $400]
#
# That is, the population and labor force growth rate n is zero if output per worker equals $400, and that each $100 increase in output per worker raises the population growth rate by 0.2 percent per year. Suppose also that the natural-resources parameter β in the production function: Y = Kα (EL L)1−α−β (ER )β is 0.2. Suppose also that gL=0.
#
# 1. If gR=0.2% per year, what is the steady-state rate of population growth? If it is 0.4% per year? 1% per year?
# 2. How long does it take the population to double for each of the efficiency-of-using- resources growth rates above?
# 3. By what multiple does the population grow in a millennium for each of the efficiency-of-labor growth rates in (a) above?
# #### Malthus
#
# Write a paragraph explaining to somebody who hasn’t taken this course how human populations could increase from 170 million in year 1 to 500 million in year 1500 without there being any noticeable increases in life expectancy or median material standards of living.
# #### Economic History Facts
#
# 1. Roughly, how much larger was global world product in 1800 than it was in the year 1?
# 2. Roughly, how large is global world product today?
# 3. Roughly, how much larger is global world product today than it was in 2000?
# #### Argentina
#
# In 1960, Argentina had a level of output per worker of $14,000/year. In the 1940s and 1950s it had had a savings-investment share of 24% and a labor-force growth rate of 2% per year. Since 1960 Argentina has averaged a savings-investment share of 15% and a labor force growth rate of 1%/year. Assume that Argentina in the 1940s and 1950s had an efficiency of labor growth rate g of 1%/year. Assume α= 2⁄3. Assume the depreciation rate δ is 5%/year. Assume that Argentina in 1960 was on its 1940s and 1950s steady-state balanced-growth path, and that Argentina today is on its post-1960 balanced growth path.
#
# 1. What was Argentinaʼs capital-output ratio in 1960?
# 2. What was Argentinaʼs efficiency of labor E in 1960?
# 3. If the efficiency-of-labor growth rate had been 1%/year since 1960, what would Argentinaʼs efficiency of labor, capital-output ratio, and level of output per worker be today?
# 4. If the efficiency of labor growth rate had been 2%/year since 1960, what would Argentinaʼs efficiency of labor, capital-output ratio, and level of output per worker be today?
# 5. If the efficiency of labor growth rate had been 3%/year since 1960, what would Argentinaʼs efficiency of labor, capital-output ratio, and level of output per worker be today?
# 6. If the efficiency of labor growth rate had been 4%/year since 1960, what would Argentinaʼs efficiency of labor, capital-output ratio, and level of output per worker be today?
# 7. If the efficiency of labor growth rate had been 5%/year since 1960, what would Argentinaʼs efficiency of labor, capital-output ratio, and level of output per worker be today?
# 8. Argentinaʼs level of output per worker today is $27000/year. What do you guess its growth rate of the efficiency of labor has been on average since 1960?
# ## Chapter 6: Reality of Economic Growth: Present and Future
# #### Suppose somebody
#
# who hasn't taken any economics courses were to ask you why it is that some countries are so very, very much poorer than others in the world today. What answer would you give? (Two paragraphs only, please!)
# #### Constant Returns
#
# Suppose we have our standard growth model with s =15 percent, n = 1 percent, g = 1 percent, and δ = 3 percent. Suppose also that the current level of the efficiency of labor E is $5,000 per year and the current level of capital per worker is $25,000. Suppose further that the parameter α in the production function is equal to 1: α = 1.
#
# 1. What can you say about output per worker in this economy? What would you project output per worker to be at some point in the distant future?
# 2. Suppose the savings rate s is not 15 percent but 17.5 percent. How would that change your projections of the future growth of output per worker?
# 3. What effect does growth in the efficiency of labor have on output per worker when α = 1?
# 4. Why aren’t the normal Solow model tools of analysis and rules of thumb much use when α = 1?
# #### Computer Revolution
#
# Today it appears that because of the computer revolution the rate of growth of the efficiency of labor in the United States has more than doubled, from 1.0 percent per year before 1995 to about 3.0 percent per year since. Suppose this increase were to be permanent.Suppose the rate of labor force growth were to remain constant at 1 percent per year, the depreciation rate were to remain constant at 3 percent per year, and the American savings rate (plus foreign capital invested in America) were to remain constant at 20 percent per year. Assume that the efficiency of labor in the U.S. in 2006 is $18,500 per year, and that the diminishing-returns-to-capital parameter α is 1/2.
#
# 1. What is the change in the steady-state capital-output ratio because of this acceleration in efficiency-of-labor growth? What is the new steady-state capital- output ratio?
# 2. How would such a permanent acceleration in the rate of growth of the efficiency of labor change your forecast of the level of output per worker in 2040? c. How would your answers to (a) and (b) be different if α were 1/3? If it were 2/3?
# #### Capital:
#
# Suppose that you misestimate the capital stock—think that its rate of growth is lower than it actually is—but otherwise understand the parameters and the growth rates of the base-camp Solow growth model. How will your conclusions about the sources of economic growth be off?
# #### Prices of Capital Goods
#
# Let us suppose that the price of capital goods in terms of output-in-general P(k) varies, so that the law of motion for the capital stock in the Solow growth model is not:
#
# >dK/dt = sY − δK
#
# but is instead:
#
# >dKt = sY/P(k) − δK
#
# 1. Derive the steady-state balanced-growth capital-output ratio (K/Y)* for this model in which the price of capital goods varies.
# 2. Suppose that richer countries have lower prices of capital goods according to: Y−η PK =L Using your answer from (5), suppose that η=1/3. Now consider the three cases of α equal to 0.25, 0.5, and 0.75. Suppose that something happens in the economy to permanently increase the savings rate from 20% to 21%.
# 3. By what proportional fraction does steady-state output-per-worker increase relative to its previous steady-state balanced-growth path?
# #### Leninism and Stalinism
#
# Take a look at http://en.wikipedia.org/wiki/List_of_countries_by_GDP_%28PPP%29_per_capita_per_hour — the wikipedia page for “List of countries by GDP (PPP) per capita per hour.” Select some countries that were once ruled by Communist governments and compare them to neighboring countries that were lucky enough to escape Communist rule. How big a shadow do you guess having a Communist government imposes on a nation on average, even today?
# #### Long-Run American Growth
#
# Take a look at the 2006 Economic Report of the President at http://www.gpoaccess.gov/eop/index.html. Read pages 36-37 and 43-46. How would you translate the arguments made by <NAME> and his colleagues about the long-run growth rate of the American economy into concepts introduced in this course?
# #### International Disparities
#
# 1. Roughly, what is the gap between real per capita GDP in the U.S. today and real per capita GDP in China?
# 2. Roughly, what is the gap between real per capita GDP in China today and real per
# capita GDP in Mozambique?
# #### Botswana
#
# In the 1950s Botswanaʼs savings rate averaged 6 percent of GDP. In 1960 Botswanaʼs level of GDP per capita was 900 of todayʼs dollars per year. Since 1960 Botswanaʼs savings rate has averaged 30 percent of GDP. Today Botswanaʼs level of GDP per capita is 15000 per year.
#
# Assume that the diminishing-returns parameter α in our production function for Botswana is 0.5, that Botswanaʼs population growth rate n has been constant at 2 percent per year, and that its depreciation rate δ has been constant at 4 percent per year. Assume that Botswana was back in 1960 on its old steady- state balanced-growth path (for an s=0.06) and is now on its new steady-state balanced-growth path (for an s=0.30)
#
# 1. Suppose there had been no growth in the efficiency of labor in Botswana between 1960 and 2011, what do you predict that the level of GDP per capita would be in Botswana today?
# 2. How fast has the efficiency of labor grown in Botswana over the past 50 years?
# 3. What was the value of the efficiency of labor in Botswana in 1960?
# 4. What is the value of the efficiency of labor in Botswana today?
# #### Zambia
#
# In the 1950s Zambiaʼs savings rate averaged 24.5 percent of GDP. In 1960 Zambiaʼs level of GDP per capita was 1800 of todayʼs dollars per year. Since 1980 Zambiaʼs savings rate has averaged 24.5 percent of GDP. Today Zambiaʼs level of GDP per capita is 1300 per year.
#
# Assume that the diminishing-returns parameter α in our production function for Zambia is 0.5, that Zambiaʼs population growth rate n has been constant at 3% per year, and that its depreciation rate δ has been constant at 4.64% per year. Assume that Zambiaʼs was back in 1960 on its old steady-state balanced-growth path (for an s=0.3) and is now on its new steady-state balanced- growth path (for an s=0.2)
#
# 1. Suppose there had been no growth in the efficiency of labor in Zambia between 1960 and 2011, what do you predict that the level of GDP per capita would be in Zambia today?
# 2. How fast has the efficiency of labor grown in Zambia over the past 50 years?
# 3. What was the value of the efficiency of labor in Zambia in 1960?
# 4. What is the value of the efficiency of labor in Zambia today?
# #### France
#
# Since 1946 French population growth (including illegal immigration) has been constant at about 1 percent per year and France has had a savings share of 25 percent of GDP. Today France has a GDP per capita level of about 35,000 per year. The rate of growth of the efficiency of labor in France since the end of World War II in France has been constant at about 2 percent per year. Assume that France is today on its steady-state balanced-growth path.
#
# 1. If France remains on its current steady-state balanced-growth path, what will GDP per capita be in France in 2050?
# 2. If France remains on its current steady-state balanced-growth path, what will GDP per capita be in France in 2100?
# 3. What would Franceʼs level of GDP per capita have been back in 1946 if it had then been on todayʼs steady-state balanced-growth path?
# 4. In fact, Franceʼs level of GDP per capita back in 1946 was about 3,000 per year even though its efficiency of labor has grown at 2 percent per year since the end of World War II. Why do you think its level back then was so low
# #### Japan
#
# Japan has had a very high savings rate and a high growth rate of output per worker over the past half century, starting from an initial post-WWII very low level of capital per worker. What does the analysis suggest about Japan's ability to sustain a higher growth rate than other industrial countries?
# #### Italy
#
# Since 1946 Italian population growth (including illegal immigration) has been constant at about 1% per year and Italy has had a savings share of 25% of GDP. Today Italy has a GDP per capita level of about $25,000 per year. The rate of growth of the efficiency of labor in Italy since the end of World War II has been constant at about 2% per year. Assume that Italy is today on its steady-state balanced-growth path.
#
# 1. If Italy remains on its current steady-state balanced-growth path, what will GDP per capita be in Italy in 2050?
# 2. If Italy remains on its current steady-state balanced-growth path, what will GDP per capita be in France in 2100?
# 3. What would Italyʼs level of GDP per capita have been back in 1946 if it had then been on todayʼs steady-state balanced-growth path?
# 4. In fact, Italy level of GDP per capita back in 1946 was about $2,500 per year even though its efficiency of labor has grown at 2% per year since the end of World War II. Why do you think its level back then was so low?
# #### Environment
#
# Suppose that environmental regulations lead to a diversion of investment spending from investments that boost the capital stock to investments that replace polluting with less-polluting capital. In our standard growth model, what would be the consequences of such a diversion for the economy's capital-output ratio and for its balanced-growth path? Would it make sense to say that these environmental regulations diminished the economy's wealth?
# #### Korea
#
# In the 1950s, South Korea had a savings-investment share of GDP of 10%. In 1960, South Korea had a GDP per worker level of $2000 (at 2010 prices, in international dollars). Since 1960 South Koreaʼs savings-investment share of GDP has averaged 27.5%. Today South Korea has a GDP per worker level of $40,000. In the 1950s, South Koreaʼs population growth rate averaged 3% per year. Since 1960 South Koreaʼs population growth rate has averaged 1% per year. Assume that the depreciation rate on the capital stock has been constant at 5%. Assume that the diminishing-returns parameter in the production function α=0.5. Assume that the growth rate of South Koreaʼs efficiency of labor was 0 in the 1950s, and has been constant at some positive value g since. Assume that South Korea in 1960 was in its steady-state balanced-growth path, and is today on its steady-state balanced growth path.
#
# a. What was South Koreaʼs efficiency of labor E in 1960?
# b. Suppose the rate of growth of the efficiency of labor in South Korea since 1960
# has averaged 6% per year. What would be the efficiency of labor in South Korea
# today?
# c. Suppose the rate of growth of the efficiency of labor in South Korea since 1960
# has averaged 5% per year. What would be the capital-output ratio in South Korea
# today?
# d. Suppose the rate of growth of the efficiency of labor in South Korea since 1960
# has averaged 5% per year. What would be the level of output per worker in South
# Korea today?
# e. Do you think the average growth rate of the efficiency of labor in South Korea
# since 1960 has been faster or slower than 5%. Why?
# f. Suppose that we have α=2/3 rather than α=0.5. How would your answers be different?
# #### Korea II
#
# Since 1960 South Koreaʼs savings-investment share of GDP has averaged 27.5%. Since 1960 the United Statesʼs savings-investment share of GDP has averaged 20%. Today South Korea has a GDP per worker level of $40,000. Today the United States has a GDP per worker level of $70,000. Since 1960 South Koreaʼs and the United Statesʼs population growth rates have both averaged 1%/year. Assume that the depreciation rate on the capital stock has been constant at 5%/year and that the rate of improvement of the efficiency of labor in the United States has averaged 2% per year. Assume that both South Korea and the United States today are on their balanced growth paths.
#
# a. What is the efficiency of labor in South Korea today?
# b. What is the efficiency of labor in the United States today?
# c. If the efficiency of labor in the United States continues to grow at its long-run trend pace of 2% per year, what is your forecast of the level of output per worker in the United States in 2100?
# d. What is your forecast of output per worker in South Korea in 2100?
# #### Bangladesh
#
# Bangladesh: In 1960 annual output per worker in what was to become Bangladesh averaged 1200 dollars. Today annual output per worker in Bangladesh averages $3000. If output per worker in Bangladesh continues to grow at the average pace it has grown since 1960...
#
# a. How long will it take Bangladesh to achieve the productivity levels that South Korea has today?
# b. How long will it take Bangladesh to achieve the productivity levels that the U.S. has today?
# c. If output per worker in the U.S. continues to grow at its long-run historical average rate of 2 percent/year, what will output per worker in the U.S. be when Bangladesh becomes as prosperous as the United States is now?
# #### Comparisons
#
# 1. Roughly, what is the gap between real per capita GDP in Belgium today, real per capita in Indonesia, and real GDP per capita in Nigeria?
# 2. Roughly, what is the gap between real per capita GDP in Qatar today, real per capita in Venezuela, and real GDP per capita in Nigeria?
# #### Economic Growth
#
# With the ebbing of the computer revolution and the growing worry that an increasing share of economic activity in the future will be concentrated in high labor-cost low productivity-growth sectors, many economists fear that the rate of growth of the efficiency of labor in the United States will average 1.2 percent per year in the the future rather than the 2.0 percent per year that has been the average since 1900. Assume that the rate of labor force growth remains constant at 0.8 percent per year, that the depreciation rate were to remain constant at 3 percent per year, that the year-2010 efficiency of labor in the United States was $25,000 per year, that the diminishing-returns-to-capital parameter α in the production function is 1/2, and that the American savings rate (plus foreign capital invested in America) has averaged 20 percent per year.
#
# 1. What was the level of output per worker in 2010 if the United States was then on its steady-state balanced-growth path?
# 2. If these fears that productivity growth will fall to 1% per year are justified, what is your forecast of the efficiency of labor in the United States in 2050?
# 3. If these fears that productivity growth will fall to 1% per year are justified, what is your forecast of the level of GDP per worker in the United States in 2050 if the savings rate remains at 20 percent?
# 4. If these fears that productivity growth will fall to 1% per year are justified, what is your forecast of the level of GDP per worker in the United States in 2050 if tax cuts and large deficits lead the savings rate to average 15 percent?
# 5. If these fears that productivity growth will fall to 1% per year are justified, what is your forecast of the level of GDP per worker in the United States in 2050 if wise fiscal policies and government-subsidized savings plans lead the savings rate to average 25 percent?
# ## Chapter 7: Circular Flows and General Gluts
# #### Sources of Price Stickiness
#
# Think about the four possible source of price stickiness: money illusion, "fairness" considerations, misperceptions of price changes, and menu costs. What have you read or seen in the past two months that strike you as examples of any of these four phenomena? Which of the four strikes you as most likely to be the most important? Be brief!—one paragraph only.
# #### Institutional Sources of Price Stickiness
#
# What changes in the economy's institutions can you think of that would diminish price stickiness and increase price flexibility? What advantage in terms of the size of the business cycle would you expect to follow from such changes in institutions? What disadvantages do you think that such institutional changes might have? (One paragraph only!)
# #### <NAME>
#
# in 1803 claimed that because nobody makes anything without intending to use it or sell it, and nobody sells anything without intending to buy something else, that there could be no general shortage of demand in an economy-- that there could be a planned excess of supply of some commodities, but it would be balanced by a planned excess of demand of some other commodities. Was he wrong? Why was he wrong?
# ## Chapter 8: Building Blocks of the Flexible Price Model
# ## Chapter 9: Equilibrium in the Flexible Price Model
# #### Adding Up National Product
#
# In the simple income-expenditure model with real national product Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: where Y = C + O and C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose C = $10.25 trillion, I = $1.68 trillion, G = $2.97 trillion, GX = $1.66 trillion, the tax rate t=0, and IM = $2.10 trillion. What is national product Y?
# 2. Suppose C = $3.83 trillion, the tax rate t=0, I = $0.86 trillion, G = $1.18 trillion, GX = $.55 trillion, and Y = $5.80 trillion. What is gross imports IM?
# 3. Suppose Suppose IM = $0.29 trillion, the tax rate t=0, I = $0.48 trillion, G = $0.57 trillion, GX = $.28 trillion, and Y = $2.79 trillion. What is consumption spending C?
# 4. Suppose Suppose IM = $0.02 trillion, the tax rate t=0, I = $0.08 trillion, G = $0.11 trillion, GX = $0.03 trillion, and C = $0.33 trillion. What is national product Y?
# #### Adding Up National Product II
#
# In the simple income-expenditure model with real GDP Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose I = $1.7 trillion, G = $3 trillion, GX = $1.7 trillion, co = $3 trillion, cy = 0.5, the tax rate t=0, and imy = .15. What is national product Y?
# 2. Suppose I = $1.7 trillion, G = $3.5 trillion, GX = $1.7 trillion, co = $3 trillion, cy = 0.5, the tax rate t=0, and imy = .15. What is national product Y?
# 3. Suppose I = $1.7 trillion, G = $4 trillion, GX = $1.7 trillion, co = $3 trillion, cy = 0.5, the tax rate t=0, and imy = .15. What is national product Y?
# 4. Suppose I = $1.7 trillion, G = $2.5 trillion, GX = $1.7 trillion, co = $3 trillion, cy = 0.5, the tax rate t=0, and imy = .15. What is national product Y?
# #### Adding Up National Product III
#
# In the simple income-expenditure model with real GDP Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose I = $1.8 trillion, G = $3 trillion, GX = $1.7 trillion, co = $3 trillion, cy = 0.5, the tax rate t=0, and imy = .15. What is GDP Y?
# 2. Suppose I = $1.8 trillion, G = $3.5 trillion, GX = $1.7 trillion, co = $2 trillion, cy = 1.0, the tax rate t=0.85, and imy = .15. What is GDP Y?
# 3. Suppose I = $1.7 trillion, G = $2 trillion, GX = $1.8 trillion, co = $3 trillion, cy = 0.65, the tax rate t=0, and imy = .15. What is GDP Y?
# ## Chapter 10: Money, Prices, and Inflation
# #### Money Growth, Real Growth, and Inflation
#
# Suppose that the rate of labor force growth is 1% per year and that the rate of growth of the efficiency of labor is 2% per year. Suppose also that the rate of growth of the nominal money stock is 15% per year, and there are little or no changes in how the banking system works. Do you think that it is likely that the inflation rate is less than 5% per year? Why or why not?
# #### Open Market Operations
#
# What would the Federal Reserve have to do if it wanted to raise the monetary base today by $20 billion? What do you guess would happen to the interest rate on short-term government bonds if the Federal Reserve did this?
# #### "Money"
#
# Do you think that unspent balances on credit cards--the difference between what you currently owe on your credit card and the limit that the credit card company allows you-- should be counted as "money"? Why or why not? (One sentence only!)
# #### Quantity Theory of Money
#
# Suppose that the rate of labor force growth is 1% per year, the efficiency of labor is growing at 2% per year, and the economy is on its steady state growth path. Suppose also that the trend is that the velocity of money is growing at 1% per year.
#
# 1. How fast should the Federal Reserve seek to make the growth rate of the money stock if it its inflation target is price stability--a 0% per year rate of growth of the CPI?
# 2. How fast should the Federal Reserve seek to make the growth rate of the money stock if it its inflation target is a 2% per year rate of growth of the CPI?
# 3. How fast should the Federal Reserve seek to make the growth rate of the money stock if it its inflation target is a 5% per year rate of growth of the CPI?
# #### Quantity Theory of Money:
#
# 1. Suppose that the level of real potential output Y* is $14 trillion, the nominal money stock M is $2 trillion, and the velocity of money V is 7. What is the price level P?
# 2. Suppose that the level of real potential output Y* is $14 trillion, the nominal money stock M is $2.5 trillion, and the velocity of money V is 5. What is the price level P?
# 3. Suppose that the level of real potential output Y* is $14 trillion, the nominal money stock M is $4 trillion, and the velocity of money V is 7. What is the price level P?
# 4. Suppose that the level of real potential output Y* is $15 trillion, the nominal money stock M is $4 trillion, and the velocity of money V is 5. What is the price level P?
# ## Chapter 11: Consumption and the Multiplier
# #### The Keynesian Cross
#
# Consider an economy in which prices are sticky, the marginal propensity to consume out of disposable income Cy is 0.75, the tax rate t is 0.25, and the share of national income spent on imports IMy is 20 percent.
#
# 1. Suppose that total autonomous spending is $6 trillion. Graph planned expenditure as a function of total national income.
# 2. Determine the equilibrium level of national income and real product.
# 3. What is the value of the multiplier in this economy?
# 4. Suppose that total autonomous spending increases by $100 billion to $6.1 trillion. What happens to the equilibrium level of national income and real GDP, Y?
# #### Raising and Lowering Short-Run Real Product
#
# Classify the following set of changes into two groups: those that increase equilibrium real national product, and those that decrease real national product:
#
# 1. An increase in consumers' desire to spend today.
# 2. An increase in interest rates overseas.
# 3. A decline in foreign exchange speculators' confidence in the value of the home currency.
# 4. A fall in real GDP overseas.
# 5. An increase in government purchases.
# 6. An increase in managers' expectations of the future profitability of investments. g. An increase in the tax rate.
# #### Boosting Real Product via Government Purchases
#
# 1. Suppose that the government wishes (for good reasons) to increase the equilibrium level of real GDP by $800 billion. How would you suggest that the government go about figuring out how to accomplish this goal?
# 2. Suppose that the economy is short of its full-employment level of GDP, $12 trillion, by $1,000 billion, with the MPC out of disposable income Cy equal to 0.6, the import share IMy equal to 0.2, and the tax rate t equal to 25%.
# 3. Suppose the government wants to boost real GDP up to full employment by cutting taxes. How large a cut in the tax rate is required to boost real GDP to full employment? How large a cut in total tax collections is produced by this cut in the tax rate?
# 4. Suppose the government wants to boost real GDP up to full employment by increasing government spending. How large an increase in government spending is required to boost real GDP to full employment?
# 5. Can you account for any asymmetry between your answers?
# #### Shifting National Product
#
# In the simple income-expenditure model with real national product Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: where Y = C + O and C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose cy = 0.5, the tax rate t=0, imy = 0.5, and G increases by $100 billion. What happens to GDP Y?
# 2. Suppose cy = 0.5, the tax rate t=0, imy = 0.16667, and G increases by $100 billion. What happens to GDP Y?
# 3. Suppose cy = 0.9, the tax rate t=0, imy = 0.1, and G increases by $100 billion. What happens to GDP Y?
# 4. Suppose cy = 0.25, the tax rate t=0, imy = 0.25, and G increases by $100 billion. What happens to GDP Y?
# #### Shifting National Product II
#
# In the simple income-expenditure model with real GDP Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose cy = 0.5, the tax rate t=0, imy = 0.5, and I increases by $100 billion. What happens to GDP Y?
# 2. Suppose cy = 0.5, the tax rate t=0, imy = 0.16667, and GX increases by $100 billion. What happens to GDP Y?
# 3. Suppose cy = 0.8, the tax rate t=0, imy = 0.05, and I increases by $100 billion. What happens to GDP Y?
# 4. Suppose cy = 0.25, the tax rate t=0, imy = 0.25, and c0 increases by $100 billion. What happens to GDP Y?
# #### Shifting National Product III
#
# In the simple income-expenditure model with real GDP Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose cy = 0.6, the tax rate t=0, imy = 0.1, and I falls by $250 billion. What happens to national product Y?
# 2. Suppose cy = 0.6, the tax rate t=0.2, imy = 0.1, and I falls by $250 billion. What happens to national product GDP Y?
# 3. Suppose cy = 0.6, the tax rate t=0.4, imy = 0.1, and I falls by $250 billion. What happens to national product GDP Y?
# 4. Suppose cy = 0.6, the tax rate t=0.6, imy = 0.1, and I falls by $250 billion. What happens to national product GDP Y?
# 5. Since the 1930s, left-wing economists have argued that an economy with a higher share of government spending and taxes in GDP is less vulnerable to business cycle downturns. Why might this be so?
# #### Shifting National Product IV
#
# In the simple income-expenditure model with real GDP Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose cy = 0.6, the tax rate t=0.2, imy = 0.0, and G rises by $250 billion. What happens to GDP Y?
# 2. Suppose cy = 0.6, the tax rate t=0.2, imy = 0.1, and G rises by $250 billion. What happens to GDP Y?
# 3. Suppose cy = 0.6, the tax rate t=0.2, imy = 0.2, and G rises by $250 billion. What happens to GDP Y?
# 4. Suppose cy = 0.6, the tax rate t=0.2, imy = 0.3, and G rises by $250 billion. What happens to GDP Y?
# 5. Since the 1940s, economists have argued that as world trade and the propensity to import rise, government spending becomes less effective as a tool for boosting production and fighting depressions. Why might this be so?
# #### Shifting National Product V
#
# In the simple income-expenditure model with real GDP Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose cy = 0.4, the tax rate t=0.2, imy = 0.2, and I falls by $200 billion. What happens to national product Y?
# 2. Suppose cy = 0.6, the tax rate t=0.2, imy = 0.2, and I falls by $200 billion. What happens to national product Y?
# 3. Suppose cy = 0.8, the tax rate t=0.2, imy = 0.2, and I falls by $200 billion. What happens to national product Y?
# 4. Suppose cy = 1.0, the tax rate t=0.2, imy = 0.2, and I falls by $200 billion. What happens to national product Y?
# ## Chapter 12: The Investment and Savings Curve
# #### TFU
#
# Label each of the following statements as True, False, or Uncertain, and explain your answer briefly.
#
# 1. In the short run, an increase in consumer confidence raises the real interest rate, consumption, and real GDP.
# 2. The fact that the U.S. output growth was only slightly less volatile in the four decades or so after World War 2 than in the four decades or so before the Great Depression shows that fiscal and monetary policy do not have large short-run effects on output.
# 3. If consumers decide to save more at a given level of disposable income than before, investment will rise, and so output will rise.
# #### IS Equilibrium: Tax Increase
#
# Suppose there is a tax increase.
#
# 1. In the simple IS-MP model with a single real interest rate, does consumption rise, fall,
# stay the same, or is it impossible to tell? Does investment rise, fall, stay the same, or is it
# impossible to tell?
# 2. In the IS-MP model with an interest rate differential, does consumption rise, fall, stay
# the same, or is it impossible to tell? Does investment rise, fall, stay the same, or is it impossible to tell?
# #### True, False, or Uncertain, and explain your answer briefly:
#
# 1. “In the IS-MP model extended to include an interest rate differential, an increase in confidence in the soundness of the financial system increases output.”
# 2. “Introducing an interest rate differential into the IS-MP model makes the AD curve steeper than it otherwise would be.”
# 3. “If the fiscal policy multiplier is less than 1, then fiscal policy cannot help to end a recession.”
# #### Pick the Best Answer
#
# 1. Researchers who are trying to estimate the effects of changes in government purchases on real GDP often focus on military purchases because: a. The types of government purchases that policymakers are likely to use to combat a recession are similar in important ways to military purchases. b. Wars are often accompanied by price controls, which makes them particularly good times for isolating the effects of changes in government purchases. c. Changes in military purchases are caused mainly by geopolitical developments outside the United Sates, not by other factors affecting U.S. GDP. d. It is hard to obtain data on non-military purchases.
# 2. “asset price bubble” means: a. A large, rapid rise in asset prices. b. A rise in asset prices that is reversed.c. A rise in asset prices in response to reductions in interest rates. d. Asset prices being greater than their “fundamental” values.
# 3. Suppose a variable, y, is determined by a factor we can measure, x, and by other factors that we cannot. That is, y = a + bx + e, where a and b are parameters and e is the unobserved other factors. Estimates of b from a regression of y on x (estimated by “ordinary least squares”) will tend to overestimate the impact of x on y if: a. x and e are negatively correlated (that is, e tends to be low when x is high, and high when x is low). b. x and e are uncorrelated (that is, x has no consistent relationship with e). c. x and e are positively correlated (that is, e tends to be high when x is high, and low when x is low). d. a and x are positively correlated.
# 4. The fact that the bankruptcy of Lehman Brothers nearly caused the failure of institutions that had made loans to Lehman Brothers is an example of: a. “Confidence” contagion. b. “Coordination” or “Fire sale” contagion. c. “Counterparty” contagion. d. None of the above.
# 5. The “borrowing” interest rate, rb, is usually higher than the “saving” interest rate, rs, because: a. Financial intermediaries’ information production, liquidity transformation, and diversification provision are costly. b. Government regulations limit the interest rates that banks can pay. c. The interest rates that foreigners can get on their savings are generally lower than interest rates in the United States. d. The inflation rate faced by savers is usually lower than the inflation rate faced by borrowers.
# #### Income-Spending Shifts
#
# Consider ΔY = [ΔA0 + ΔG - (Ir + Xεεr)Δr]/(1-(1-t)cy + imy), the investment savings framework (with ΔA0 = Δc0 + ΔI0 -XεΔε0 + Xy*ΔY* + XεεrΔr). Suppose the multiplier 1/(1-(1-t)cy + imy) = 1.5 and the responsiveness of exports to the exchange rate Xε = 500...
#
# 1. What happens to Y if government purchases G goes up by $100 billion, and nothing else changes?
# 2. What happens to Y if baseline investment spending I0 goes up by $100 billion, and nothing else changes?
# 3. What happens to Y if baseline consumption spending c0 goes up by $100 billion, and nothing else changes?
# 4. What happens to Y if speculator confidence in the currency ε0 goes up by 20%— by 0.2—and nothing else changes?
# 5. Explain the similarities and the differences between your answers to (a)-(d).
# #### Income-Spending Shifts II
#
# Consider ΔY = [ΔA0 + ΔG - (Ir + Xεεr)Δr]/(1-(1-t)cy + imy), the investment savings framework (with ΔA0 = Δc0 + ΔI0 -XεΔε0 + Xy*ΔY* + XεεrΔr). Suppose the multiplier 1/(1-(1-t)cy + imy) = 1.5 and the responsiveness of exports to the exchange rate Xε = 500...
#
# 1. Suppose that capital controls keep the exchange rate from responding to changes in the interest rate—suppose that εr = 0—and suppose the sensitivity of investment spending to the interest rate Ir = 50. If the interest rate falls by 2%—by 200 basis points, or by 0.02—what happens to Y?
# 2. Suppose that the responsiveness of the exchange rate to changes in the interest εr = 1, and suppose the sensitivity of investment spending to the interest rate Ir = 50. If the interest rate falls by 2%—by 200 basis points, or by 0.02—what happens to Y?
# 3. Suppose that the responsiveness of the exchange rate to changes in the interest εr = 5, and suppose the sensitivity of investment spending to the interest rate Ir = 50. If the interest rate falls by 2%—by 200 basis points, or by 0.02—what happens to Y?
# 4. Suppose that the responsiveness of the exchange rate to changes in the interest εr = 20, and suppose the sensitivity of investment spending to the interest rate Ir = 50. If the interest rate falls by 2%—by 200 basis points, or by 0.02—what happens to Y?
# 5. Explain the similarities and the differences between your answers to (a)-(d). What features of the situation besides government controls on foreign investment might influence the value of εr?
# #### Income-Spending Shifts III
#
# Consider ΔY = [ΔA0 + ΔG - (Ir + Xεεr)Δr]/(1-(1-t)cy + imy), the investment savings framework (with ΔA0 = Δc0 + ΔI0 -XεΔε0 + Xy*ΔY* + XεεrΔr). Suppose the multiplier 1/(1-(1-t)cy + imy) = 1.5, the responsiveness of exports to the exchange rate Xε = 500, and the responsiveness of the exchange rate to changes in the interest εr = 10...
#
# 1. What happens to Y if the real interest rate r goes up by 1%—by 100 basis points or by 0.01?
# 2. What happens to Y if the real interest rate r goes up by 1%—by 100 basis points or by 0.01—and if baseline investment spending goes up by 50?
# 3. What happens to Y if the real interest rate r goes down by 1%—by 100 basis points or 0.01—and if speculator confidence in the currency goes up by 10%?
# 4. What happens to Y if the real interest rate r goes up by 1%—by 100 basis points or by 0.01— and if baseline consumption spending goes down by 50?
# #### Income-Spending Shifts IV
#
# Consider ΔY = [ΔA0 + ΔG - (Ir + Xεεr)Δr]/(1-(1-t)cy + imy), the investment savings framework (with ΔA0 = Δc0 + ΔI0 -XεΔε0 + Xy*ΔY* + XεεrΔr). Suppose the multiplier 1/(1-(1-t)cy + imy) = 1.5 and the responsiveness of exports to the exchange rate Xε = 500...
#
# 1. What happens to Y if the real interest rate r goes up by 2%—by 100 basis points or by 0.01—and if speculator confidence in the currency goes down by 20%?
# 2. What happens to Y if the real interest rate r goes up by 1%—by 100 basis points or by 0.01—and if baseline investment spending goes down by 200?
# 3. What happens to Y if the real interest rate r goes down by 1%—by 100 basis points or 0.01—and if speculator confidence in the currency goes up by 5%?
# 4. What happens to Y if the real interest rate r goes up by 1%—by 100 basis points or by 0.01— and if baseline consumption spending goes up by 100?
# 5. How would your answers to (8) be different if instead of 1/(1-(1-t)cy + imy) = 1.5, t=0.25, cy =0.8 imy=0.1?
# #### Income-Spending Shifts V
#
# Consider ΔY = [ΔA0 + ΔG - (Ir + Xεεr)Δr]/(1-(1-t)cy + imy), the investment savings framework (with ΔA0 = Δc0 + ΔI0 -XεΔε0 + Xy*ΔY* + XεεrΔr). Suppose the multiplier 1/(1-(1-t)cy + imy) = 1.5 and the responsiveness of exports to the exchange rate Xε = 500...
#
# 1. Why does a contractionary monetary policy abroad that raises interest rates abroad raise GDP at home?
# 2. Why does an outburst of enthusiasm among foreign exchange speculators that makes them more confident about the long-run value of the home currency reduce GDP at home?
# 3. When an outburst of enthusiasm among foreign exchange speculators that makes them more confident about the long-run value of the home currency reduces GDP at home, what components of GDP change and in which direction?
# 4. What would be the effects on GDP at home of a stimulative fiscal policy abroad that raised real GDP abroad?
# #### International Shocks
#
# Consider ΔY = [ΔA0 + ΔG - (Ir + Xεεr)Δr]/(1-(1-t)cy + imy), the investment savings framework (with ΔA0 = Δc0 + ΔI0 -XεΔε0 + XyfΔYf + XεεrΔrf). Suppose the responsiveness of exports to the exchange rate Xε = 500, the responsiveness of the exchange rate to interest rates εr=10, and the responsiveness of investment to the interest rate Ir = 1000. And suppose t=0.2, cy =0.8 imy=0.14:
#
# 1. What happens to Y if the real interest rate r goes up by 2%—by 0.02—and if speculator confidence in the currency goes down by 30%?
# 2. What happens to Y if the real interest rate r goes up by 1%—by 0.01—and if baseline investment spending goes down by 300?
# 3. What happens to Y if the real interest rate r goes down by 1%—by 0.01—and if speculator confidence in the currency goes up by 5
# #### Depression Economic
#
# In the simple income-expenditure model with real GDP Y equal to the sum of consumption spending by households C, investment spending by businesses I, government purchases G, and with net exports NX; with consumption spending C given by the equation: C = co + cyY(1-t); and with imports IM given by the equation: IM = imyY...
#
# 1. Suppose I = $1.7 trillion, G = $2 trillion, GX = $1.3 trillion, co = $3 trillion, cy = 0.85, the tax rate t=0, and imy = .10. What is GDP Y?
# 2. Suppose I = $1.7 trillion, G = $3.5 trillion, GX = $0.8 trillion, co = $3 trillion, cy = 0.65, the tax rate t=0, and imy = .15. What is GDP Y?
# 3. Suppose I = $2.3 trillion, G = $4 trillion, GX = $1.7 trillion, co = $3 trillion, cy = 0.6, the tax rate t=0.67, and imy = .2. What is GDP Y?
# 4. Suppose I = $1.5 trillion, G = $2.5 trillion, GX = $1.0 trillion, co = $3 trillion, cy = 0.67, the tax rate t=0, and imy=.0.WhatisGDPY?
# ## Chapter 13: Financial Crises
# #### Financial Crisis
#
# This problem asks you to perform an event study. In the extension of the IS-MP-IA model to include an interest rate differential, a financial crisis is assumed to raise the differential. This problem therefore asks you to investigate whether interest rate differentials (often referred to as interest rate spreads) rose at the time of some of the major events associated with the financial crisis in the fall of 2008.
#
# Specifically, consider the difference between the interest rates on BAA corporate bonds and on 30-year Treasury bonds. And consider any 3 of the following 4 events, all of which occurred in September 2008: (1) The government takeover of <NAME> and <NAME>; (2) Lehman Brothers’ announcement that it would file for bankruptcy; (3) The government takeover of AIG; (4) The House of Representatives’ rejection of the initial TARP legislation. (Note: no extra credit for considering all 4 events.)
# a.
# 1. For each event that you chose, find the day that it occurred. As far as you can tell, did it occur when financial markets were open (which, loosely speaking, corresponds to normal working hours in New York) or when they were closed? Describe how you found this information and how confident you are about it.
# 2. For each event, if it occurred when financial markets were open, find the change in the interest rate spread on the day of the event. If financial markets were closed, find the change in the spread on the next day of trading. Did these events lead to noticeable increases in the interest rate spread? Be sure to describe where you found your data.
# 3. What was the overall change in the spread from the end of August 2008 to the end of October 2008?
# 4. For many economic relationships (for example, between money and output), we would not expect a big effect in a day. Explain why one might or might not expect the worsening of a financial crisis to have a noticeable impact on interest rate spreads in a day.
# #### Risky Interest Rates
#
# Our model with an interest rate differential assumes that all investment depends on a single interest rate, rb. In truth, there are many interest rates, and they are relevant to many different kinds of investment.
#
# To partially capture this idea, suppose investment is the sum of two types of investment, a safer type I1 and a riskier type I2. I1 depends on rb1 and I2 depends on rb2: I1 = I1(rb1), I2 = I2(rb2), where both functions are decreasing. Suppose also that rb1 – rs = d1(Y) and rb2 – rs = d2(Y), again with both functions decreasing. To reflect the idea that Type 2 investment is riskier than Type 1 investment, assume that for a given Y, rb2 – rs is greater than rb1 – rs, and that when Y falls, rb2 – rs rises by more than rb1 – rs does.
#
# 1. In this model, suppose there is a fall in consumer confidence, so that C for a given Y – T is lower than before.
# i. What is the effect of this change on rs, Y, rb1 – rs, and rb2 – rs? ii. Is the change in rb1 – rs larger, smaller, or the same as the change in rb2 – rs, or is it not possible to tell?
# 2. One message we have stressed is that a modeling assumption is not inherently “good” or “bad”; the value of an assumption depends on the question we are trying to answer. Thus, give one example of a question that moving from a model with just one interest rate differential (rb – rs) to one with two differentials (rb1 – rs and rb2 – rs) would be helpful in answering. Give one example of a question where moving from a model with one differential to a model with two would complicate the analysis without generating any significant additional insights.
# #### Debt and Vulnerability
#
# Recent research stresses the role of high levels of debt for consumers and firms in causing or exacerbating short-run fluctuations.
#
# 1. What is some of the evidence that high levels of debt affect consumer or firm behavior?
# 2. If consumers and firms are trying to reduce their debt (deleverage), how, if at all, is this likely to show up in the IS-MP diagram? Why?
# 3. What policy actions would be most effective in combating a recession caused by deleveraging? Why?
# #### True, False, or Uncertain, and explain your answer briefly
#
# 1. “The bursting of an asset price bubble can lead to a financial crisis.”
# 2. “High income inequality can result in a long-lasting shortfall of aggregate demand and very prolonged high unemployment.”
#
# #### The Eurozone
#
# Both Iceland and Ireland suffered major banking crises starting around September 2008. One potentially important difference between the two countries is that Ireland is a member of the Eurozone while Iceland has its own currency:
#
# 1. By how much did the Icelandic Króna depreciate against the euro from August 2008 to its low point? How much lower is the Króna against the euro today than it was in August 2008?
# 2. By how much did the unemployment rate rise, in percentage points, in Iceland from its pre-crisis level to its peak? By how much did it rise in Ireland?
# 3. If the exchange rate was important to the different performances of the two countries, we would expect that to be reflected in the behavior of net exports. Is the behavior of net exports in the two countries consistent with the hypothesis that exchange rate depreciation helped to cushion the effects of the banking crisis in Iceland?
# Pick the best answer to each of questions 5–8. No explanations of your answers are needed.
# #### Financial Panics in the Great Depression
#
# In the paper by <NAME> Troost on banking panics in Mississippi, the authors find that the monetary policy beliefs and behavior of the Federal Reserve Banks of Atlanta and St. Louis were:
#
# 1. Fundamentally different before, during, and after the panics of 1930.
# 2. Different in the 1920s, but very similar starting in 1929.
# 3. Similar throughout the 1920s and 1930s.
# 4. Very different before and during the panic of 1930, but similar during later panics in
# the Depression.
# #### "Uncertainty"
#
# Uncertainty is likely to have an especially large negative short-run effect on spending by households and firms if:
#
# 1. They do not expect the uncertainty to be resolved soon.
# 2. There are large costs to reversing their spending decisions.
# 3. The real interest rate is high.
# 4. Inflation is high.
# 5. All of the above.
# 6. None of the above.
# #### Financial Sophistication
#
# In 1986 your lecturer <NAME> and <NAME> wrote a paper in which they argued that the coming of more sophisticated financial markets—in which households could borrow and repay easily and did not have to respond to a $1 decline in their incomes by cutting their consumption spending by a large fraction of $1—would be less vulnerable to business cycle downturns. It now looks as though we were catatrophically wrong. But why did we think this back then?
# ## Chapter 14: Liquidity and Money
# ## Chapter 15: The Phillips Curve, Expectations, and Monetary Policy
# #### Adaptive Expectations
#
# Suppose that the economy's Phillips curve is given by:
#
# >u = u' − β(π − πe)
#
# with β equal to 0.5 and the natural rate of unemployment u' equal to 5 percent. Suppose that the economy has for a long time had a constant inflation rate π equal to 2 percent per year. Suddenly the government announces a new policy: it will use fiscal policy to push the unemployment rate down by 2 percentage points--and promises it will keep that expanded fiscal policy in place indefinitely.
#
# Suppose, further that the dominant way of forming expectations in the economy is such that people have adaptive expectations of inflation--so that this year's expected inflation is equal to last year's actual inflation. What will be the course of inflation and unemployment in this economy in the years after the shift in fiscal policy? Track the economy out ten years, assuming that there are no additional shocks.
# #### Rational Expectations
#
# Suppose that the economy's Phillips curve is given by:
#
# >u = u' − β(π − πe)
#
# with β equal to 0.5 and the natural rate of unemployment u' equal to 5 percent. Suppose that the economy has for a long time had a constant inflation rate π equal to 2 percent per year. Suddenly the government announces a new policy: it will use fiscal policy to push the unemployment rate down by 2 percentage points--and promises it will keep that expanded fiscal policy in place indefinitely.
#
# Suppose further that agents in the economy have rational expectations of inflation--so that this year's expected inflation is what an economist knowing the structure of the economy and proposed economic policies would calculate actual inflation was likely to be. What will be the course of inflation and unemployment in this economy in the years after the shift in fiscal policy? Track the economy out twenty years, assuming that there are no additional shocks.
# #### The Sacrifice Ratio
#
# Suppose that an economy starts at an initial inflation level π0 and that its central bank seeks to reduce inflation down to some final level πT by pushing the unemployment rate u up above the natural rate of unemployment u'. Suppose that the relationship between inflation and unemployment is given by the adaptive-expectations Phillips Curve equation:
#
# >π = π−1+β(u'−u)
#
# where β is a known parameter. Let U stand for the total cumulative excess of € unemployment over the natural rate needed to accomplish this policy.
#
# Solve, algebraically, for the total cumulative excess unemployment U. In what units is U measured? Suppose that you are asked to calculate the sacrifice ratio S/( π0- πT). How is the sacrifice ratio related to the parameter β in the Phillips Curve? Write a paragraph explaining how you would go about deciding whether the policy of reducing inflation through higher-than-natural unemployment is a good one or a bad one.
# #### Quantity Theory of Money
#
# Suppose that the rate of labor force growth is 3% per year, the efficiency of labor is constant, and the economy is on its steady state growth path. Suppose also that the rate of growth of the nominal money stock is 10% per year. Do you think that it is likely that the inflation rate is less than 5% per year? Why or why not?
# #### Phillips Curve
#
# In the Phillips Curve framework in which π = E(π) + β(un - u)—the inflation rate π equals the previously-expected inflation rate E(π) plus the Phillips Curve slope parameter β times the difference between the economy's natural rate of unemployment un and the current rate of unemployment u...
#
# 1. If E(π) = 9% per year, u* = 6%, and u = 8%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 1/2?
# 2. If E(π) = 3% per year, u* = 4%, and u = 4%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 1/2?
# 3. If E(π) = 1% per year, u* = 7%, and u = 3%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 1/3?
# 4. If E(π) = 1% per year, u* = 7%, and u = 3%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 2/3?
# 5. If E(π) = 1% per year, u* = 7%, and u = 3%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 1?
# #### Monetary Policy
#
# Suppose we have an economy with a natural rate of unemployment of 6%, current expected inflation of 2%, and a Phillips Curve slope parameter of 1/2. Suppose that the Federal Reserve has a target ut for the unemployment rate and a target πT for the inflation rate, and suppose that for each percentage point inflation is above its target level the Federal Reserve raises unemployment by an extra percentage point above its target level.
#
# 1. If the target for the inflation rate is 2% and the target for the unemployment rate is 6%, what will inflation and unemployment be?
# 2. If the target for the inflation rate is 3% and the target for the unemployment rate is 4%, what will inflation and unemployment be?
# 3. If the target for the inflation rate is 6% and the target for the unemployment rate is 8%, what will inflation and unemployment be?
# 4. If the target for the inflation rate is 4% and the target for the unemployment rate is 4%, what will inflation and unemployment be?
# #### Monetary Policy II
#
# Suppose we have an economy with a natural rate of unemployment of 4%, current expected inflation of 15%, and a Phillips Curve slope parameter of 1/2. Suppose that the Federal Reserve has a target u0 for the unemployment rate and a target πt for the inflation rate, and suppose that for each percentage point inflation is above its target level the Federal Reserve raises unemployment by an extra two percentage points above its target level.
#
# 1. Suppose that from this year forward the Federal Reserve sets its target for the inflation rate at 3% and its target for the unemployment rate at 5%, what will inflation and unemployment be this year?
# 2. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be next year?
# 3. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be two years from now?
# 4. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be five years from now?
# 5. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be ten years from now?
# #### Monetary Policy III
#
# Suppose we have an economy with a natural rate of unemployment of 4%, and a Phillips Curve slope parameter of 1. Suppose that the Federal Reserve has a target u0 for the unemployment rate and a target πt for the inflation rate, and suppose that for each percentage point inflation is above its target level the Federal Reserve raises unemployment by an extra two percentage points above its target level.
#
# 1. If the Federal Reserve's target for the inflation rate is 2% and its target for the unemployment rate is 4%, what will the long run rate of inflation be?
# 2. If the Federal Reserve's target for the inflation rate is 2% and its target for the unemployment rate is 6%, what will the long run rate of inflation be?
# 3. If the Federal Reserve's target for the inflation rate is 4% and its target for the unemployment rate is 4%, what will the long run rate of inflation be?
# 4. If the Federal Reserve's target for the inflation rate is 4% and its target for the unemployment rate is 8%, what will the long run rate of inflation be?
# #### Phillips Curve
#
# In the Phillips Curve framework in which π = E(π) + β(u* - u)—the inflation rate π equals the previously-expected inflation rate E(π) plus the Phillips Curve slope parameter β times the difference between the economy's natural rate of unemployment u* and the current rate of unemployment u...
#
# 1. If E(π) = 2% per year, u* = 6%, and u = 10%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 1/2?
# 2. If E(π) = 3% per year, u* = 4%, and u = 6%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 1/2?
# 3. If E(π) = 6% per year, u* = 7%, and u = 3%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 1/3?
# 4. If E(π) = 1% per year, u* = 7%, and u = 9%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 2/3?
# 5. If E(π) = 4% per year, u* = 8%, and u = 5%, what is the inflation rate π going to be if the Phillips Curve slope parameter β = 1?
# #### Phillips Curve
#
# In the Phillips Curve framework in which π = E(π) + β(u* - u)— the inflation rate π equals the previously-expected inflation rate E(π) plus the Phillips Curve slope parameter β times the difference between the economy's natural rate of unemployment u* and the current rate of unemployment u...
#
# 1. IfE(π)=9%peryear,u*=6%,andu=8%,whatistheinflationrateπ going to be if the Phillips Curve slope parameter β = 1/2?
# 2. IfE(π)=3%peryear,u*=4%,andu=4%,whatistheinflationrateπ going to be if the Phillips Curve slope parameter β = 1/2?
# 3. IfE(π)=1%peryear,u*=7%,andu=3%,whatistheinflationrateπ going to be if the Phillips Curve slope parameter β = 1/3?
# 4. IfE(π)=1%peryear,u*=7%,andu=3%,whatistheinflationrateπ going to be if the Phillips Curve slope parameter β = 2/3?
# 5. IfE(π)=1%peryear,u*=7%,andu=3%,whatistheinflationrateπ going to be if the Phillips Curve slope parameter β = 1?
# #### Phillips Curve
#
# In the Phillips Curve framework in which π = E(π) + β(u* - u)— the inflation rate π equals the previously-expected inflation rate E(π) plus the Phillips Curve slope parameter β times the difference between the economy's natural rate of unemployment u* and the current rate of unemployment u...
#
# 1. Suppose that the economy starts out with an expected rate of inflation of 2%/year, a Phillips Curve slope parameter of 1/2, and a natural rate of unemployment of 5%. Suppose that the Federal Reserve decides to reduce the rate of unemployment to 3% through expansionary monetary policy and does so. What is the inflation rate?
# 2. Now look one year ahead to next year. Next year, expectations of inflation will be adaptive: that is, expectations of inflation next year will be equal to what inflation was this year. What will expectations of inflation be next year?
# 3. What will actual inflation be next year if the Federal Reserve continues to pursue expansionary monetary policies to keep the rate of unemployment at 3%?
# 4. If inflation expectations continue to be adaptive in this sense that each year's expected is the previous year's actual inflation, what will actual inflation be two years from now?
# 5. If inflation expectations continue to be adaptive and if the Federal Reserve continues to follow policies to keep the unemployment rate at 3%, what will actual inflation be five years from now?
# 6. If inflation expectations continue to be adaptive and if the Federal Reserve continues to follow policies to keep the unemployment rate at 3%, what will actual inflation be ten years from now?
# 7. If inflation expectations continue to be adaptive and if the Federal Reserve continues to follow policies to keep the unemployment rate at 3%, what will actual inflation be twenty years from now?
# 8. . Do you think that inflation expectations will continue to be adaptive in this sense as this process rolls forward over the next twenty years, or do you think the process by which expectations of inflation are formed will change? e. If you think it will change, what do you think it will change to?
#
# ## Chapter 16: Stabilization Policy
# #### Monetary vs. Fiscal Policy
#
# Why do economists today tend to believe that monetary policy is superior to discretionary fiscal policy as a stabilization policy tool? In what circumstances that you can imagine would this belief be reversed? (One paragraph only!)
# #### Raising Investment
#
# Suppose that the government and central bank together want to keep GDP constant but raise the rate of investment. What policies can they follow to achieve this? (One sentence only!)
# #### Credibility
#
# Former Federal Reserve Vice Chair <NAME> has remarked that <NAME>’s policies at fighting unemployment have been much more aggressive in attempting to reduce unemployment than any Federal Reserve Chair with less of a reputation as an inflation hawk would dare attempt. Can you make sense of this remark?
# #### Reputation
#
# Suppose that you believe that investors, businesses, and workers in your economy have rational expectations of inflation. Suppose that you have a choice between two candidates to head your central bank—one of whom believes that the central bank must always do whatever is necessary to keep inflation low, and the other of whom believes that if the central bank were to push unemployment above the natural rate to try to reduce inflation it would be making a serious mistake. Which candidate would you prefer to run your central bank, and why?
# #### Sticky Price Output and Interest Rates
#
# Describe how, if at all, each of the following developments affects the real interest rate and output in the short run (or whether it is not possible to tell). In parts (1) and (2), assume that the central bank is following an interest rate rule; in parts (3) and (4), use the information in the question to decide what assumption to make about how monetary policy is being conducted:
#
# 1. The government cuts taxes.
# 2. The government cuts taxes and government purchases by equal amounts.
# 3. The government cuts taxes and, at the same time, the central bank changes its monetary policy rule so that it sets a lower real interest rate at a given level of output than before.
# 4. The central bank lowers its target for the money stock.
# #### Monetary Policy Rules
#
# Suppose the central bank changes its policy rule to respond more to changes in output. Specifically, it decides not to change the real interest rate at the current level of output, but that it will increase it by more than before if output rises, and cut it by more than before if output falls:
#
# 1. Would you use the IS–MP or IS–LM model to analyze the effects of this development? Why?
# 2. How, if at all, would this development affect the two curves (IS and MP, or IS and LM)? Explain your reasoning.
# 3. Suppose that after this change in the monetary policy rule, firms become more optimistic about the profitability of investment projects, so investment demand at a given real interest rate is higher than before. How, if at all, does this change affect the real interest rate and output in the short run? For each variable (r and Y), is the effect larger than before the change in the rule, smaller, the same, or is it impossible to tell?
# #### Shocks Requriring Stabilization
#
# Describe how, if at all, each of the following developments affects real output, the exchange rate, and net exports in the short run. Assume the central bank is following an interest rate rule…
#
# 1. The discovery of new investment opportunities causes investment demand to be higher at a given interest rate than before.
# 2. The central bank changes its monetary policy rule so that it sets a lower real interest rate at a given level of output than before.
# 3. The demand for money increases (that is, consumers’ preferences change so that at a given level of i and Y they want to hold more real balances than before).
# #### Real Business Cycles
#
# Consider our usual IS-MP-IA model, starting in long-run equilibrium. Suppose that potential output falls because of reduced productivity.
# 1. How, if at all, would this change show up in the IS-MP diagram in the short run?
# 2. What will be the impact of the fall in on output and inflation in the long run?
# #### Econometrics
#
# Explain in a few sentences, without using any math, what is wrong with the following argument: “I obtained annual data on money growth and output growth, and then estimated the OLS regression Δ ln Y = a + b Δ ln M + u. After running the regression, I computed the correlation between money growth and the regression residual. It was exactly zero. Thus, since there is no correlation between the right-hand side variable and the residual, I know that my regression does not suffer from omitted-variable bias.
# #### Monetary Policy
#
# Suppose we have an economy with a natural rate of unemployment of 6%, current expected inflation of 4%, and a Phillips Curve slope parameter of 1/2. Suppose that the Federal Reserve has a target u0 for the unemployment rate and a target πt for the inflation rate.
#
# 1. If the target for the inflation rate is 4% and the target for the unemployment rate is 6%, what will inflation and unemployment be?
# 2. If the target for the inflation rate is 2% and the target for the unemployment rate is 6%, what will inflation and unemployment be if for each extra percentage point of inflation the Federal Reserve raises unemployment by an extra two percentage points?
# 3. If the target for the inflation rate is 2% and the target for the unemployment rate is 6%, what will inflation and unemployment be if for each extra percentage point of inflation the Federal Reserve raises unemployment by an extra half a percentage point?
# 4. If the target for the inflation rate is 2% and the target for the unemployment rate is 6%, what will inflation and unemployment be if for each extra percentage point of inflation the Federal Reserve raises unemployment by an extra percentage point?
# #### Monetary Policy
#
# Suppose we have an economy with a natural rate of unemployment of 6%, current expected inflation of 10%, and a Phillips Curve slope parameter of 1/2. Suppose that the Federal Reserve has a target u0 for the unemployment rate and a target πt for the inflation rate, and suppose that for each percentage point inflation is above its target level the Federal Reserve raises unemployment by an extra percentage point above its target level.
#
# 1. Suppose that from this year forward the Federal Reserve sets its target for the inflation rate at 3% and its target for the unemployment rate at 5%, what will inflation and unemployment be this year?
# 2. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be next year?
# 3. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be two years from now?
# 4. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be five years from now?
# 5. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be ten years from now?
# #### Monetary Policy
#
# Suppose we have an economy with a natural rate of unemployment of 6%, and a Phillips Curve slope parameter of 1/2. Suppose that the Federal Reserve has a target u0 for the unemployment rate and a target πt for the inflation rate, and suppose that for each percentage point inflation is above its target level the Federal Reserve raises unemployment by an extra two percentage points above its target level.
#
# 1. If the Federal Reserve's target for the inflation rate is 2% and its target for the unemployment rate is 4%, what will the long run rate of inflation be?
# 2. If the Federal Reserve's target for the inflation rate is 2% and its target for the unemployment rate is 6%, what will the long run rate of inflation be?
# 3. If the Federal Reserve's target for the inflation rate is 4% and its target for the unemployment rate is 4%, what will the long run rate of inflation be?
# 4. If the Federal Reserve's target for the inflation rate is 4% and its target for the unemployment rate is 8%, what will the long run rate of inflation be?
# #### Monetary Policy
#
# Suppose we have an economy with a natural rate of unemployment of 6%, current expected inflation of 2%, and a Phillips Curve slope parameter of 1/2. Suppose that the Federal Reserve has a target ut for the unemployment rate and a target πt for the inflation rate, and suppose that for each percentage point inflation is above its target level the Federal Reserve raises unemployment by an extra percentage point above its target level.
# 1. If the target for the inflation rate is 2% and the target for the unemployment rate is 6%, what will inflation and unemployment be?
# 2. If the target for the inflation rate is 3% and the target for the unemployment rate is 4%, what will inflation and unemployment be?
# 3. If the target for the inflation rate is 6% and the target for the unemployment rate is 8%, what will inflation and unemployment be? 4. If the target for the inflation rate is 4% and the target for the unemployment rate is 4%, what will inflation and unemployment be?
# #### Monetary Policy
#
# Suppose we have an economy with a natural rate of unemployment of 6%, current expected inflation of 10%, and a Phillips Curve slope parameter of 1/2. Suppose that the Federal Reserve has a target ut for the unemployment rate and a target πt for the inflation rate, and suppose that for each percentage point inflation is above its target level the Federal Reserve raises unemployment by an extra percentage point above its target level.
#
# 1. Suppose that from this year forward the Federal Reserve sets its target for the inflation rate at 2% and its target for the unemployment rate at 4%, what will inflation and unemployment be this year?
# 2. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be next year?
# 3. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be two years from now?
# 4. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be five years from now?
# 5. Suppose expected inflation is adaptive in that each year's expected inflation is the previous year's actual inflation. What will inflation and unemployment be ten years from now?
# ## Chapter 17: The Liquidity Trap
# #### DeLong and Summers (1992)
#
# The fact that the central bank cannot push the short-term nominal interest rate below zero raises the possibility of a liquidity trap—an inability of the central bank to push the long-term real interest rate down to a level that produces enough investment to get full employment. Back in 1992 <NAME> and <NAME> argued that this made it desirable for the Federal Reserve to have an inflation target of 4-5% per year rather than 1-2% per year: the higher inflation target, you see, would give the Federal Reserve more ability to reduce the long-term real interest rate when necessary. The Fed didn’t think much of this argument. What do you think of it?
# ## Chapter 18: The Government Debt
# ## Chapter 19: Secular Stagnation
# #### Secular Stagnation
#
# The three “coping mechanisms” that Reich argues middle class families used to deal with stagnating incomes included all of the following except:
#
# 1. Women move into paid work.
# 2. Everyone works longer hours.
# 3. Families stop investing in their children’s education. d. We draw down savings and borrow to the hilt.
# ## Chapter 20: The Short Run Shapes the Long Run: "Hysteresis"
# #### A Fluctuating Natural Rate of Unemployment
#
# Begin our very simple Phillips Curve:
#
# >π = πe +β(u' −u)
#
# with simple adaptive expectations:
#
# >πe =π−1
#
# But add a difference: the natural rate of unemployment depends on what unemployment was last year:
#
# >u' = (1−θ)u'−1 +θu−1
#
# for some parameter θ between zero and one.
#
# Suppose that the central bank induces a recession and raises the unemployment rate one percentage point above its natural rate for one year, and then lets unemployment fall back to its natural rate:
#
# 1. What is the time path of inflation as a result of this one-year shift in policy?
# 2. What is the time path of unemployment?
# 3. How does the Sacrifice Ratio—the amount of excess point-years of unemployment that must be run in order to permanently reduce the inflation rate by one percent—depend on the parameter θ?
# 4. What actions can you think of that the central bank might take that could reduce the value of θ?
# #### Supply Shocks
#
# Suppose that a supply shock hits the economy—that is, that for one year the Phillips curve is not:
#
# >π = πe + β(u'− u)
#
# But is instead:
#
# >π = πe +β(u' − u )+ s
#
# where st is some positive shock to inflation caused by, say, a spike in oil prices. And
# suppose that inflation expectations are adaptive:
#
# >πe = π−1
#
# 1. What happens to inflation over time if the central bank keeps unemployment at its natural rate always?
# 2. What happens to the unemployment rate over time if the central bank adjusts unemployment to keep inflation at its initial value π0 always?
# #### Permanent Unemployment Shocks
#
# Suppose we have our standard Phillips curve:
#
# >π = πe +β(u' − u)
#
# with our standard adaptive inflation expectations:
#
# >πe =π−1
#
# But that the natural rate of unemployment u' is subject to hysteresis: u' =u−1
#
# How is this situation is different from that of a constant natural rate of unemployment combined with static inflation expectations?
# #### The Computer Revolution and the Natural Rate of Unemployment
#
# In the late 1990s Alan Greenspan argued that the natural rate of unemployment in the U.S. had temporarily fallen because of the acceleration of productivity growth. Briefly, outline a model of worker behavior in which this claim of Alan Greenspan’s is true, and outline a model of worker behavior in which this claim of Alan Greenspan’s would have been false. Keep your answer to whatever equations you find necessary and to less than 300 words (and a couple of figures).
# #### European Unemployment
#
# Explain—to somebody who has never taken an economics course—the reasons you think that western European unemployment today is so much higher than western European unemployment was back in the 1960s.
# #### The Japanese Slowdown
#
# Explain—to somebody who has never taken an economics course—the reasons that you think economic growth in Japan slowed from 4.5% over 1973-1991 to less than 1.5% over 1991-present. Keep your answer to less than 300 words (and a couple of figures).
# #### Hysteresis
#
# When economists speak of hysteresis, they refer to the fact that:
#
# 1. Once a recession starts, it is likely to last for a while.
# 2. High unemployment tends to be followed by low unemployment.
# 3. Prolonged high unemployment can cause the natural rate of unemployment to rise.
# 4. Prolonged high unemployment will eventually reduce inflation.
# ## Chapter 21: International Economic Policy
# #### Speculative Attacks
#
# Explain—to somebody who has never taken an economics course—why a loss of confidence in the value of a country’s currency on the part of foreign exchange speculators is highly likely to lead to a fall in the rate of investment even if no financial crisis is created. Keep your answer to less than 150 words (and a figure or two).
# #### Foreign Monetary Policy
#
# Suppose that many foreign central banks tighten their monetary policies, so that real interest rates abroad rise.
#
# 1. How would you expect this change to affect the CF(r) function in the United States? Why?
# 2. How would the change affect output and the real interest rate in the short run? Explain.
# 3. How would the change affect net exports, and the real exchange rate in the short run? Explain.
# #### Flexible Price World Foreign Exchange Crises
#
# Start out with our consensus flexible-price model in “difference” form, the relevant parts of which are:
#
# >ΔY = ΔC + ΔI + ΔG + ΔX − ΔIM
# ΔC = Cy x (1−t)(ΔY)
# ΔI = ΔI0 − Ir(Δr)
# ΔX = Xε(Δε)
# Δε = Δε0 − εr(Δr)
#
# and let’s consider the effect of a collapse of confidence in the currency—a large sudden € rise in the Δε0 parameter that governs long-term expectations of the price of foreign currency.
#
# Suppose that there are no effects on the supply side—ΔY=0—no effects on investor confidence—ΔI0=0—no effects on imports—ΔIM=0—and that government spending is unchanged as well—ΔG=0.
#
# 1. In the flexible price model, solve for the equilibrium changes in the exchange rate ε, in the interest rate r, in investment I, and in exports X as a result of the rise in
# Δε0.
# 2. Suppose that the cause of the crisis is that government spending is rising rapidly and that investors have no confidence that the budget will ever be balanced, and thus that the government bonds they own will ever be repaid without inflation. Thus there is a relationship between government spending and the change in expectations of the exchange rate: Δε0 = θΔG for some value of the parameter θ. Now solve for the equilibrium changes in the exchange rate ε, in the interest rate r, in investment I, and in exports X as a result of the rise government spending G.
# €
# ΔY =ΔC+ΔI+ΔG+ΔX −ΔIM ΔC = Cy (1− t)ΔY
# ΔI=ΔI0 −IrΔr−Iε(Δε)2 ΔX = XεΔε
# ΔIM = −IMyΔY Δε=Δε0 −εrΔr
# #### Sticky Price Price World Foreign Exchange Crises
#
# Now let’s turn to the consensus sticky-price model, the relevant parts of which are:
#
# >ΔY = ΔC + ΔI + ΔG + ΔX − ΔIM
# ΔC = Cy x (1−t)(ΔY)
# ΔI = ΔI0 − Ir(Δr) - Iε(Δε)^2
# ΔX = Xε(Δε)
# ΔIM = −IMy(ΔY)
# Δε = Δε0 − εr(Δr)
#
# and once again consider the effect of a collapse of confidence in the currency—a large sudden rise in the Δε0 parameter that governs long-term expectations of the price of foreign currency—this time in the short run in which full employment is not guaranteed, and in which the real interest rate r is chosen by the central bank. And not the extra term in the investment equation: a large shift in the exchange rate causes massive bankruptcies, destabilizes the financial system, and causes investment I to collapse and output Y to fall.
#
# 1. Solve for the change in the exchange rate as a function of the change in the interest rate r, and then turn that around: if the change in the exchange rate is going to take on a certain value Δε, what must the change in the interest rate Δr be?
# 2. Now let’s view the central bank as choosing not the interest rate but the exchange rate. In this model, solve for the change in output ΔY as a function of the change in the exchange rate Δε (taking account of the fact that the central bank must choose the change in the interest rate Δr to make the change in the exchange rate equal to the chosen value.
# 3. What is the best that the central bank can do in the short run. Where should it choose to set the exchange rate and the interest rate (bearing in mind that they are linked) to make the effects of the crisis as small as possible?
# 4. Suppose that the IMF is on hand to help, and to give the central bank extra foreign exchange R that it can then spend to affect the exchange rate: Δε = Δε0 −εr(Δr) −εr(ΔR). Solve for how the best the central bank can do is affected by the addition of more help from the IMF.
# ## Chapter 22: Changes in the Macroeconomy and Changes in Macroeconomic Policy
# #### Pick the Best Answer
#
# 1. If the central bank is targeting the money supply, an increase in government purchases: a. Shifts the LM curve up. b. Shifts the LM curve down. c. Does not affect the LM curve. d. It is not possible to tell.
# 2. The IS curve slopes down because: a. As the real interest rate rises, the government increases taxes to finance the greater interest payments on its debt. b. As the real interest rate rises, the central bank tightens monetary policy. c. As the real interest rate rises, the government cuts back on its purchases. d. As the real interest rate rises, households invest less in the stock market. e. As the real interest rate rises, firms buy fewer machines and build fewer factories. f. (a) and (b). g. All of the above.
# 3. Saying that the regression yt = a + bxt + et suffers from omitted variable bias means that: a. There is correlation between the variables left out of the regression (which show up in e) and the variable whose effect on y we are trying to estimate (x). b. The variables left out of the regression (which show up in e) affect the variable whose behavior we are trying to understand (y). c. The researcher chose to omit some observations because they do not support his or her hypothesis. d. y is on average high when x is high, and low when x is low.
# 4. The following periods are listed in order from least to greatest macroeconomic volatility:va. 1886–1916, 1947–1985, 1985–2005. b. 1886–1916, 1985–2005, 1929–1941. c. 1985–2005, 1947–1985, 1929–1941. d. 1985–2005, 1929–1941, 1947–1985. e. None of the above.
# #### The Gold Standard
#
# Label the following statement as True, False, or Uncertain, and explain your answer briefly: “The gold standard played little role in the Great Depression.”
# #### Pick the Best Answer
#
# 1. Suppose that initially Y > Yf. As the economy moves to long-run equilibrium: a/ Inflation rises. b. Output falls. c. The real interest rate rises. d. (a) and (b). e. All of the above.
# 2. In the extension of the IS–MP–IA model to include the zero lower bound, the “kink” in the AD curve occurs at: a. The inflation rate given by the IA curve. b. The inflation rate that causes the IS and MP curves to intersect at the “kink” of the MP curve. c. Y = Yf. d. π=0. e. (c) and (d).
# 3. The fact that inflation has not fallen very much since 2007 despite the fact that unemployment has been very high over that period could be the result of any of the following except: a. Inflation expectations are “anchored.” b. The normal or natural rate of unemployment has risen substantially. c. The Federal Reserve is constrained by the zero lower bound. d. There have been inflation shocks acting to increase the inflation rate.
# ## Chapter 23: The Past, Present, and Future of Macroeconomics
# #### What (briefly!) does <NAME> think of <NAME>?
# #### What (briefly!) does <NAME> think of <NAME>?
# ## Auxiliary Readings
# #### Write one paragraph
#
# (one hundred to two hundred words) explaining to somebody who has not read the article what its main point is: <NAME>'s "Cartoon Model" http://krugman.blogs.nytimes.com/2008/03/10/a-cartoon-model-of-the-crisis-more-serious-wonkery/
# #### Write one paragraph
#
# (one hundred to two hundred words) explaining to somebody who has not read the article what its main point is: <NAME>, "Global Current Account Imbalances: Hard Landing or Soft Landing?" http://www.imf.org/external/np/speeches/2005/031505.htm
# #### Write one paragraph
#
# (one hundred to two hundred words) explaining to somebody who has not read the article what its main point is: <NAME>, "Global Imbalances" http://www.econ.berkeley.edu/~obstfeld/280c_sp07/adjustment.pdf
# #### Write one paragraph
#
# (one hundred to two hundred words) explaining to somebody who has not read the article what its main point is: <NAME>,"China's Exchange Rate Regime: The Long and Short of It" http://www.econ.berkeley.edu/~eichengr/research/short.pdf
# #### Write one paragraph
#
# (one hundred to two hundred words) explaining to somebody who has not read the article what its main point is: <NAME>, "The Dollar and the Renminbi" http://www.iie.com/publications/papers/paper.cfm?ResearchID=747
# Institute for International Economics, "China: The Balance Sheet" http://www.petersoninstitute.org/publications/chapters_preview/04648/0
# ## Essays
#
# 1. Why did America have a housing boom in the mid-2000s?
#
# 2. Why did the conditions that had been required for mortgage borrowers before 2000--20% down payment, evidence of a stable job, no more than a 33% ratio of housing expenses (including utilities and taxes) to income--disappear in the 2000s?
#
# 3. Why did the world economy fall into a very deep economic recession at the end of 2008?
#
# 4. Why is recovery from the current downturn in the United States likely to be partial and delayed?
#
# 5. Why are the economies of East and South Asia likely to grow much faster than the United States over the next half-decade or so?
#
# 6. Why is America’s health care spending per capita so much higher than health care spending in other industrialized countries?
#
# 7. What will happen if America never brings its government revenues up to balance with spending, but keeps running federal budget deficits into the future?
# ## Identifications
#
# 1. National product
# 2. Income-expenditure equation
# 3. Subprime mortgages
# 4. Unemployment rate
# 5. Inflation rate
# 6. Long-term real risky interest rate
# 7. Investment-savings curve
# 8. Investment-savings equation
# 9. Marginal propensity to consume
# 10. About how many people lose or quit their jobs in an average month?
# 11. About how many people get jobs in an average month?
# 12. About how many people are unemployed in an average month?
# ## Short Answers
#
# 1. Why did America have a housing boom in the mid-2000s?
# 2. Why did the conditions that had been required for mortgage borrowers before 2000--20% down payment, evidence of a stable job, no more than a 33% ratio of housing expenses (including utilities and taxes) to income-- disappear in the 2000s?
# 3. Why did the world economy fall into a very deep economic recession at the end of 2008?
# 4. What are the five (four positive, one negative) major components of national product on the expenditure side?
# 5. <NAME> in 1803 claimed that, because nobody makes anything without intending to use it or sell it, and nobody sells anything without intending to buy something else, that there could be no general shortage of demand in an economy--that there could be a planned excess of supply of some commodities, but it would be balanced by a planned excess of demand of some other commodities. Was he wrong? Why was he wrong?
| DeLong Problem Sets Original.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
random_scalar = tf.random_uniform(())
sess = tf.Session()
sess.run(random_scalar)
# +
double_random_scalar = 2 * random_scalar
double_random_scalar_gt_one = double_random_scalar > 1
sess.run(double_random_scalar_gt_one)
# -
np.random.rand(2,3)
random_tensor = tf.random_uniform((2,3))
double_random_tensor = 2 * random_tensor
double_random_tensor_gt_one = double_random_tensor > 1
sess.run(double_random_tensor_gt_one)
# +
def np_fn(x):
mu=0
sigma=1
exp=np.e ** ( -(((x-mu)**2) / (2*sigma**2)))
rslt=(1 / np.sqrt(2 * sigma**2 * np.pi)) * exp
print(rslt)
x = np.array([0, 1, 2, 3])
np_fn(x)
# +
import tensorflow as tf
import numpy as np
def tf_fn(x):
mu=0
sigma=1
x1 = tf.placeholder(tf.float32, [None])
exp=tf.exp(-((tf.subtract(x1,mu)**2) / (2*sigma**2)))
rslt = tf.rsqrt(2 * sigma**2 * np.pi)*exp
sess = tf.Session()
sess.run(tf.global_variables_initializer())
rslt2 = sess.run(rslt, {x1: x})
return rslt2
x = np.array([0, 1, 2, 3])
tf_fn(x)
| labs/lab1/Lab1 - first tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
# %load_ext autoreload
# %autoreload 2
from IPython.core.debugger import set_trace
import matplotlib.pyplot as plt
# %matplotlib notebook
from pathlib import Path
import glob
import regex
import dill
import gzip
import toolz, itertools, more_itertools
from collections import Counter, OrderedDict
import optuna
import sklearn
import math
import pandas as pd
import numpy as np
import seaborn as sns
from imblearn.combine import SMOTETomek
import statistics
import shap
### SEED ###
np.random.seed(42)
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.metrics import roc_curve, precision_recall_curve, precision_recall_fscore_support
from sklearn.metrics import average_precision_score
from matplotlib import pyplot
from sklearn.metrics import plot_roc_curve
from sklearn.metrics import classification_report
# ## DATA directory
DIR = Path(r'C:\Users\Abhij\OneDrive\Documents\GitHub\DNA-structure-prediction')
assert DIR.exists()
DATA = DIR/"data"
# ## Helper Functions to read pickled data
# +
try:
_file = DIR / "data"
os.mkdir(_file / "pkl")
except FileExistsError:
print("Dir exists")
def pkl_it(dataframe, filebase):
with open(DIR / "data" / "pkl" / str(filebase + ".pkl"), "wb") as fh:
dill.dump(dataframe, fh)
return
def unpkl_it(filebase):
with open(DIR / "data" / "pkl" / str(filebase + ".pkl"), "rb") as fh:
return dill.load(fh)
# -
# ## A flexible helper Class for running different ML algorithms
# It automatically chooses the best threshold for classification by locating the arg_max (index) of the best F-score
# +
import logging
from collections import defaultdict
logging.basicConfig(filename=DATA/'results.log', level=logging.INFO)
class Call_Plot():
def __init__(self, sklearn_model=True, model_name="SVM", repeated_k_fold=False):
plt.close()
self.model_name = model_name
self.fig, self.ax = plt.subplots()
self.ax.plot([0,1], [0,1], linestyle='--', label='Random choice')
self.ax.set_xlabel('False Positive Rate', fontsize=12)
self.ax.set_ylabel('True Positive Rate', fontsize=12)
self.fig2, self.ax2 = plt.subplots()
self.ax2.set_xlabel('Recall', fontsize=12)
self.ax2.set_ylabel('Precision', fontsize=12)
self.tprs = []
self.aucs = []
self.mean_fpr = np.linspace(0, 1, 100)
self.no_skill = []
self.sklearn_model = sklearn_model
self.results = defaultdict(list)
self.repeated_k_fold = repeated_k_fold
def Plot(self, data: dict, model, idx):
if self.sklearn_model:
y_pred_val = model.predict_proba(data["X_val"])[:,1]
else:
y_pred_val = model.predict(data["X_val"])
#Precision-Recall
precision, recall, thresholds = precision_recall_curve(data["y_val"], y_pred_val)
no_skill = len(data["y_val"][data["y_val"]==1]) / len(data["y_val"])
self.no_skill.append(no_skill)
avg_pr = average_precision_score(data["y_val"], y_pred_val)
auc_pr = sklearn.metrics.auc(recall, precision)
if self.repeated_k_fold:
self.ax2.plot(recall, precision, marker='.', label=f'Run {(idx)//5+1} Test Fold{(idx)%5+1}: AUC PR={auc_pr:.2f}')
else:
self.ax2.plot(recall, precision, marker='.', label=f'Test Fold{(idx)+1}: AUC PR={auc_pr:.2f}')
# convert to f score
fscore = (2 * precision * recall) / (precision + recall)
# locate the index of the largest f score
ix_pr = np.argmax(fscore)
self.ax2.scatter(recall[ix_pr], precision[ix_pr], marker='o', color='black')
Accuracy = sklearn.metrics.accuracy_score(data["y_val"], np.where(y_pred_val > thresholds[ix_pr], 1, 0))
target_names = ['B-DNA', 'A-DNA']
print(classification_report(data["y_val"], np.where(y_pred_val > thresholds[ix_pr], 1, 0), target_names=target_names))
F1 = sklearn.metrics.f1_score(data["y_val"], np.where(y_pred_val > thresholds[ix_pr], 1, 0))
MCC = sklearn.metrics.matthews_corrcoef(data["y_val"], np.where(y_pred_val > thresholds[ix_pr], 1, 0))
cohen_kappa_score = sklearn.metrics.cohen_kappa_score(data["y_val"], np.where(y_pred_val > thresholds[ix_pr], 1, 0))
logging.info(f'Fold {idx + 1}: Average PR: {avg_pr:.2f} ')
logging.info(f'Fold {idx + 1}: AUC PR: {auc_pr:.2f} ')
logging.info(f'Fold {idx + 1}: Best Threshold_f-score={thresholds[ix_pr]:.2f}, F-Score={fscore[ix_pr]}')
logging.info(f'Fold {idx + 1}: Accuracy: {Accuracy:.2f}')
logging.info(f'Fold {idx + 1}: F1: {F1:.2f}')
logging.info(f'Fold {idx + 1}: MCC: {MCC:.2f}')
#ROC-AUC
fpr, tpr, thresholds_auc = roc_curve(data["y_val"], y_pred_val)
# calculate the g-mean for each threshold
gmeans = np.sqrt(tpr * (1-fpr))
# locate the index of the largest g-mean
ix = np.argmax(gmeans)
if self.repeated_k_fold:
self.ax.plot(fpr, tpr, marker='.',
label=f'Run {(idx)//5+1} Test Fold{(idx)%5+1}: AUC={sklearn.metrics.auc(fpr, tpr):.2f}')
else:
self.ax.plot(fpr, tpr, marker='.',
label=f'Test Fold{(idx)+1}: AUC={sklearn.metrics.auc(fpr, tpr):.2f}')
self.ax.scatter(fpr[ix], tpr[ix], marker='o', color='black')
# axis labels
self.ax.legend(loc="lower left")
# Mean plot
interp_tpr = np.interp(self.mean_fpr, fpr, tpr)
interp_tpr[0] = 0.0
self.tprs.append(interp_tpr)
self.aucs.append(gmeans[ix])
logging.info(f'Fold {idx + 1}: ROC-AUC: {sklearn.metrics.auc(fpr, tpr):.2f}')
logging.info(f'Fold {idx + 1}: Best Threshold_ROC={thresholds_auc[ix]:.2f}, G-Mean_ROC={gmeans[ix]:.2f}')
print("Average PR: ", avg_pr )
print("AUC PR: ", auc_pr)
print('Best Threshold_f-score=%f, F-Score=%.3f' % (thresholds[ix_pr], fscore[ix_pr]))
print("AUC: ", sklearn.metrics.auc(fpr, tpr))
print('Best Threshold_ROC=%f, G-Mean_ROC=%.3f' % (thresholds_auc[ix], gmeans[ix]))
print("Accuracy: ", Accuracy )
print("F1: ", F1 )
print("MCC: ", MCC )
self.results["Average PR"].append(avg_pr)
self.results["AUC PR"].append(auc_pr)
self.results["ROC AUC"].append(sklearn.metrics.auc(fpr, tpr))
self.results["Accuracy"].append(Accuracy)
self.results["F1"].append(F1)
self.results["MCC"].append(MCC)
self.results["cohen_kappa_score"].append(cohen_kappa_score)
def post_Plot(self):
from sklearn.metrics import auc
mean_tpr = np.mean(self.tprs, axis=0)
mean_tpr[-1] = 1.0
mean_auc = auc(self.mean_fpr, mean_tpr)
std_auc = np.std(self.aucs)
self.ax.plot(self.mean_fpr, mean_tpr, color='b',
label=r'Mean ROC (AUC = %0.2f $\pm$ %0.2f)' % (mean_auc, std_auc),
lw=2, alpha=.8)
std_tpr = np.std(self.tprs, axis=0)
tprs_upper = np.minimum(mean_tpr + std_tpr, 1)
tprs_lower = np.maximum(mean_tpr - std_tpr, 0)
self.ax.fill_between(self.mean_fpr, tprs_lower, tprs_upper, color='grey', alpha=.2,
label=r'$\pm$ 1 std. dev.')
self.ax.legend(loc=(0.45, .05),fontsize='medium')
self.fig.savefig(DIR/"data"/"results"/f"{self.model_name}_AUC_ROC.png", dpi=600)
no_skill = np.mean(np.array(self.no_skill))
self.ax2.plot([0,1], [no_skill,no_skill], linestyle='--', label="Random")
self.ax2.legend(loc=(0.050, .08),fontsize='medium')
self.fig2.savefig(DIR/"data"/"results"/f"{self.model_name}_AUC_PR.png", dpi=600)
# -
# ## Read curated dataset
curated_data = unpkl_it("curated_dataset")
curated_data
#Check if any sequence has duplicate features
curated_data.drop_duplicates(subset=['AA/TT', 'GG/CC', 'AC/GT',
'CA/TG', 'AT/AT', 'TA/TA', 'AG/CT', 'GA/TC', 'CG/CG', 'GC/GC'], keep='last')
# ## Nested Cross-validation
from imblearn.combine import SMOTEENN, SMOTETomek
from sklearn.model_selection import RepeatedStratifiedKFold
# The function gen_data is a flexible generator that implements outer fold of Nested CV
# Here, we are using 5-fold stratified Nested cross validation (n_splits = 5)
# +
def gen_data(data: pd.DataFrame, RESAMPLING: bool=False):
X, y = data.drop(labels="target", axis=1), data["target"]
sss = RepeatedStratifiedKFold(n_splits=5, n_repeats=1, random_state=42)
#sss = StratifiedShuffleSplit(n_splits=5, test_size=0.20, random_state=42)
for train_index, val_index in sss.split(X, y):
smote_tomek = SMOTETomek(random_state=42)
X_resampled, y_resampled = smote_tomek.fit_resample(X.iloc[train_index,:], y.iloc[train_index])
yield {"X_train": X_resampled if RESAMPLING else X.iloc[train_index,:],
"y_train": y_resampled if RESAMPLING else y.iloc[train_index],
"X_val": X.iloc[val_index,:], "y_val": y.iloc[val_index]}
# -
# This generator implements inner fold of Nested CV, where we tune hyperparameters.
def gen_data_for_tuningHP(data: dict, RESAMPLING: bool=True):
X, y = data["X_train"], data["y_train"]
sss = StratifiedShuffleSplit(n_splits=3, test_size=0.3, random_state=42)
for train_index, val_index in sss.split(X, y):
smote_tomek = SMOTETomek(random_state=42)
X_resampled, y_resampled = smote_tomek.fit_resample(X.iloc[train_index,:], y.iloc[train_index])
yield {"X_train": X_resampled if RESAMPLING else X.iloc[train_index,:],
"y_train": y_resampled if RESAMPLING else y.iloc[train_index],
"X_val": X.iloc[val_index,:], "y_val": y.iloc[val_index]}
# ## Helper function: train_test_folds_reader
# This generator function reads the data from the "train_test_folds" folder and gives the same Train-Test splits
# used by us. At each iteration it yields a single split of the data
# Alternatively, you can run the `gen_data(curated_data.drop(labels=["item", "sequence"], axis=1), RESAMPLING=False)`, which gives the same split provided that you use the same seed.
#
# +
def train_test_folds_reader(*, folder) -> dict:
TRAIN_TEST = Path(folder)
for i in range(5):
yield {"X_train": pd.read_excel(folder/f"train_fold_{i+1}.xls", index_col=0).drop(labels=["target", "sequence", "item"], axis=1),
"y_train": pd.read_excel(folder/f"train_fold_{i+1}.xls", index_col=0)["target"],
"X_val": pd.read_excel(folder/f"test_fold_{i+1}.xls", index_col=0).drop(labels=["target", "sequence", "item"], axis=1),
"y_val": pd.read_excel(folder/f"test_fold_{i+1}.xls", index_col=0)["target"],}
# -
# ## Read the best SVM hyperparameters
best_svm_params = pd.read_csv(DATA/"tuned_hyperparameters"/"best_svm_params.csv", index_col=0)
best_svm_params
# ## Set up SVM training
#Class weight factor:
SCALE_POS_WEIGHT = lambda T,P: (100*(T-P)/T) / (100*P/T)
from sklearn.svm import SVC, NuSVC
from sklearn.model_selection import cross_val_score, cross_val_predict
import optuna
from optuna.pruners import HyperbandPruner
import copy
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
def trainer(data, param_updater):
train_x, train_y = data["X_train"], data["y_train"]
param = {
"probability": True,
"class_weight": "balanced",
"random_state": 42,
"verbose": False,
}
param.update(param_updater)
model = SVC(**param)
model = make_pipeline(StandardScaler(), model)
model.fit(train_x, train_y)
return model
# ## Train, Run and evaluate performance of SVM using tuned hyperparameters
# Here, we use the `train_test_folds_reader(folder=DATA/"train_test_folds")` to yield the same split of data as used us. We then use the tuned hyperparameters (best_svm_params) and convert it into a dictionary. We then train each model and evaluate the performance of each model on each test fold. Note that, alternatively we can also use the `gen_data(curated_data.drop(labels=["item", "sequence"], axis=1), RESAMPLING=False)` will give the same split of data if you use the same seed (42).
plt.close()
# *************OUTER*************
plot_Model_SVM_test = Call_Plot(repeated_k_fold=False)
for outer_idx, elem in enumerate(train_test_folds_reader(folder=DATA/"train_test_folds")):
# ***********Feed in the best hyperparams for each model************
model = trainer(elem, best_svm_params.T.to_dict()
[f"Model_{outer_idx + 1}"])
plot_Model_SVM_test.Plot(elem, model, outer_idx)
plot_Model_SVM_test.post_Plot()
pd.DataFrame(plot_Model_SVM_test.results)
pd.DataFrame(plot_Model_SVM_test.results).mean()
# ## Running from scratch - Run and evaluate performance of SVM under 5-fold stratified Nested CV
# +
import optuna
import copy
def objective(data, trial):
train_x, valid_x, train_y, valid_y = data["X_train"], data["X_val"], data["y_train"], data["y_val"]
param = {
"C": trial.suggest_discrete_uniform("C",0.1,1.0,0.1),
"kernel": trial.suggest_categorical("kernel",["rbf",]),
"gamma": trial.suggest_loguniform("gamma", 1e-3, 1e+3),
"probability": True,
"class_weight": "balanced",
"random_state": 42,
"verbose": False,
}
model = SVC(**param)
model = make_pipeline(StandardScaler(), model)
model.fit(train_x, train_y)
return sklearn.metrics.roc_auc_score(valid_y, model.predict(valid_x))
# -
import collections
Trial = collections.namedtuple("Trial",["value", "parameters"])
plt.close()
optuna.logging.set_verbosity(optuna.logging.WARNING)
results = []
outer_models = {}
best_models_svm = {}
hack_svm = {}
# *************OUTER*************
plot_Model_svm = Call_Plot(sklearn_model=True, model_name="SVM", repeated_k_fold=True)
for outer_idx, elem in enumerate(gen_data(curated_data.drop(labels=["item", "sequence"], axis=1), RESAMPLING = False)):
hack_svm[outer_idx] = elem
study_dict = {}
# ***********INNER************
for idx, data_in in enumerate(gen_data_for_tuningHP(elem, RESAMPLING=True)):
study = optuna.create_study(pruner=HyperbandPruner(max_resource="auto"),
direction="maximize")
study.optimize((toolz.curry(objective)(data_in)), n_trials=100)
#print("Number of finished trials: {}".format(len(study.trials)))
trial = study.best_trial
study_dict[idx] = Trial(trial.value, trial.params)
arg_max = max(study_dict, key=lambda d: study_dict[d].value) #max for AUC
best_models_svm[outer_idx] = trainer(elem, study_dict[arg_max].parameters)
plot_Model_svm.Plot(elem, best_models_svm[outer_idx], outer_idx)
plot_Model_svm.post_Plot()
pd.DataFrame(plot_Model_SVM_test.results)
pd.DataFrame(plot_Model_SVM_test.results).mean()
| src/.ipynb_checkpoints/SVM-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: U4-S1-NLP (Python 3)
# language: python
# name: u4-s1-nlp
# ---
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
#
# # Vector Representations
# ## *Data Science Unit 4 Sprint 2 Assignment 2*
# + colab={"base_uri": "https://localhost:8080/", "height": 159} colab_type="code" id="hyj-f9FDcVFp" outputId="5dd045fe-6e4c-458c-e2fc-253c3da9c805"
import re
import string
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import spacy
# + [markdown] colab_type="text" id="M7bcmqfGXrFG"
# ## 1) *Optional:* Scrape 100 Job Listings that contain the title "Data Scientist" from indeed.com
#
# At a minimum your final dataframe of job listings should contain
# - Job Title
# - Job Description
#
# If you choose to not to scrape the data, there is a CSV with outdated data in the directory. Remeber, if you scrape Indeed, you're helping yourself find a job. ;)
# + colab={} colab_type="code" id="KcYlc1URXhlC"
##### Your Code Here #####
raise Exception("\nThis task is not complete. \nReplace this line with your code for the task."
# + [markdown] colab_type="text" id="5C4xFZNtX1m2"
# ## 2) Use Spacy to tokenize / clean the listings
# + colab={} colab_type="code" id="dhUHuMr-X-II"
##### Your Code Here #####
raise Exception("\nThis task is not complete. \nReplace this line with your code for the task."
# + [markdown] colab_type="text" id="-lgCZNL_YycP"
# ## 3) Use Scikit-Learn's CountVectorizer to get word counts for each listing.
# + colab={} colab_type="code" id="X2PZ8Pj_YxcF"
##### Your Code Here #####
raise Exception("\nThis task is not complete. \nReplace this line with your code for the task."
# + [markdown] colab_type="text" id="Zo1iH_UeY7_n"
# ## 4) Visualize the most common word counts
# + colab={} colab_type="code" id="M5LB00uyZKV5"
##### Your Code Here #####
raise Exception("\nThis task is not complete. \nReplace this line with your code for the task."
# + [markdown] colab_type="text" id="bwFsTqrVZMYi"
# ## 5) Use Scikit-Learn's tfidfVectorizer to get a TF-IDF feature matrix
# + colab={} colab_type="code" id="-gx2gZCbl5Np"
##### Your Code Here #####
raise Exception("\nThis task is not complete. \nReplace this line with your code for the task."
# -
# ## 6) Create a NearestNeighbor Model. Write the description of your ideal datascience job and query your job listings.
# + inputHidden=false outputHidden=false
##### Your Code Here #####
raise Exception("\nThis task is not complete. \nReplace this line with your code for the task."
# + [markdown] colab_type="text" id="FiDfTWceoRkH"
# ## Stretch Goals
#
# - Try different visualizations for words and frequencies - what story do you want to tell with the data?
# - Scrape Job Listings for the job title "Data Analyst". How do these differ from Data Scientist Job Listings
# - Try and identify requirements for experience specific technologies that are asked for in the job listings. How are those distributed among the job listings?
# - Use a clustering algorithm to cluster documents by their most important terms. Do the clusters reveal any common themes?
# - **Hint:** K-means might not be the best algorithm for this. Do a little bit of research to see what might be good for this. Also, remember that algorithms that depend on Euclidean distance break down with high dimensional data.
# - Create a labeled dataset - which jobs will you apply for? Train a model to select the jobs you are most likely to apply for. :)
| module2-vector-representations/LS_DS_412_Vector_Representations_Assignment.ipynb |